code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# ### 网络科学理论
# ***
# ***
# # 网络科学简介
# ***
# ***
#
# 王成军
#
# <EMAIL>
#
# 计算传播网 http://computational-communication.com
# + [markdown] slideshow={"slide_type": "slide"}
# # FROM <NAME> TO NETWORK THEORY
# ### A SIMPLE STORY (1) The fate of Saddam and network science
#
# - <NAME>: the fifth President of **Iraq**, serving in this capacity from 16 July 1979 until 9 April **2003**
# - Invasion that started in March 19, 2003. Many of the regime's high ranking officials, including <NAME>, avoided capture.
# - Hussein was last spotted kissing a baby in Baghdad in April 2003, and then his trace went cold.
#
# - Designed a deck of cards, each card engraved with the images of the 55 most wanted.
# - It worked: by May 1, 2003, 15 men on the cards were captured, and by the end of the month another 12 were under custody.
# - Yet, the ace of spades, i.e. Hussein himself, remained at large.
#
#
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# <img src = './img/saddam.png' width = 500>
# + [markdown] slideshow={"slide_type": "slide"}
# # The capture of <NAME>
#
# - shows the strong **predictive power** of networks.
#
# - underlies the need to obtain **accurate maps of the networks** we aim to study;
# - and the often heroic **difficulties of the mapping** process.
#
# - demonstrates the remarkable **stability of these networks**
# - The capture of Hussein was not based on fresh intelligence
# - but rather on his **pre-invasion social links**, unearthed from old photos stacked in his family album.
#
# - shows that the choice of network we focus on makes a huge difference:
# - the **hierarchical tree** captured the official organization of the Iraqi government,
# - was of no use when it came to <NAME>'s whereabouts.
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ## How about <NAME>?
#
# - the founder of al-Qaeda, the organization that claimed responsibility for the September 11 attacks on the United States.
# + [markdown] slideshow={"slide_type": "slide"}
# # A SIMPLE STORY (2): August 15, 2003 blackout.
#
# <img src='./img/blackout.png' width = 800>
#
# + [markdown] slideshow={"slide_type": "slide"}
# # VULNERABILITY
# DUE TO INTERCONNECTIVITY
#
# - The 2003 blackout is a typical example of a cascading failure.
# - 1997, when the International Monetary Fund pressured the central banks of several Pacific nations to limit their credit.
# - 2009-2011 financial melt-down
# + [markdown] slideshow={"slide_type": "slide"}
# An important theme of this class:
#
# - we must understand **how network structure affects the robustness of a complex system**.
#
# - develop quantitative tools to assess the interplay between network structure and the dynamical processes on the networks, and their impact on failures.
#
# - We will learn that failures reality failures follow reproducible laws, that can be quantified and even predicted using the tools of network science.
#
# + [markdown] slideshow={"slide_type": "slide"}
# NETWORKS AT THE HEART OF
# # COMPLEX SYSTEMS
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Complex
#
# [adj., v. kuh m-pleks, kom-pleks; n. kom-pleks]
# –adjective
# - composed of many interconnected parts; compound; composite: a complex highway system.
# - characterized by a very complicated or involved arrangement of parts, units, etc.: complex machinery.
# - so complicated or intricate as to be hard to understand or deal with: a complex problem.
# Source: Dictionary.com
#
# + [markdown] slideshow={"slide_type": "slide"}
#
# # Complexity
#
# a scientific theory which asserts that some systems display behavioral phenomena that are completely inexplicable by any conventional analysis of the systems’ constituent parts. These phenomena, commonly referred to as emergent behaviour, seem to occur in many complex systems involving living organisms, such as a stock market or the human brain.
#
# Source: <NAME>, Encyclopædia Britannica
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# # COMPLEX SYSTEMS
#
# - society
# - brain
# - market
# - cell
#
# ## <NAME>: I think the next century will be the century of complexity.
# + [markdown] slideshow={"slide_type": "slide"}
# # Behind each complex system there is a network, that defines the interactions between the component.
#
# + [markdown] slideshow={"slide_type": "slide"}
# <img src = './img/facebook.png' width = 800>
# + [markdown] slideshow={"slide_type": "slide"}
# - Social graph
# - Organization
# - Brain
# - finantial network
# - business
# - Internet
# - Genes
# + [markdown] slideshow={"slide_type": "slide"}
# Behind each system studied in complexity there is an intricate wiring diagram, or a network, that defines the interactions between the component.
#
# # We will never understand complex system unless we map out and understand the networks behind them.
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# # TWO FORCES HELPED THE EMERGENCE OF NETWORK SCIENCE
#
# + [markdown] slideshow={"slide_type": "slide"}
# # THE HISTORY OF NETWORK ANALYSIS
#
# - Graph theory: 1735, Euler
#
# - Social Network Research: 1930s, Moreno
#
# - Communication networks/internet: 1960s
#
# - Ecological Networks: May, 1979.
#
# + [markdown] slideshow={"slide_type": "slide"}
#
# While the study of networks has a long history from graph theory to sociology, **the modern chapter of network science emerged only during the first decade of the 21st century, following the publication of two seminal papers in 1998 and 1999**.
#
# The explosive interest in network science is well documented by the citation pattern of two classic network papers, the 1959 paper by <NAME> and <NAME> that marks the beginning of the study of random networks in graph theory [4] and the 1973 paper by <NAME>, the most cited social network paper [5].
#
# Both papers were hardly or only moderately cited before 2000. The explosive growth of citations to these papers in the 21st century documents the emergence of network science, drawing a new, interdisciplinary audience to these classic publications.
#
# + [markdown] slideshow={"slide_type": "slide"}
# <img src = './img/citation.png' width = 500>
# + [markdown] slideshow={"slide_type": "slide"}
# # THE EMERGENCE OF NETWORK SCIENCE
# - Movie Actor Network, 1998;
# - World Wide Web, 1999.
# - C elegans neural wiring diagram 1990
# - Citation Network, 1998
# - Metabolic Network, 2000;
# - PPI network, 2001
#
# + [markdown] slideshow={"slide_type": "slide"}
# # The universality of network characteristics:
# The architecture of networks emerging in various domains of science, nature, and technology are more similar to each other than one would have expected.
#
# + [markdown] slideshow={"slide_type": "slide"}
# # THE CHARACTERISTICS OF NETWORK SCIENCE
# - Interdisciplinary
# - Empirical
# - Quantitative and Mathematical
# - Computational
#
# + [markdown] slideshow={"slide_type": "slide"}
# # THE IMPACT OF NETWORK SCIENCE
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Google
# Market Cap(2010 Jan 1):
# $189 billion
#
# # Cisco Systems
# networking gear Market cap (Jan 1, 2919):
# $112 billion
#
# # Facebook
# market cap:
# $50 billion
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Health: From drug design to metabolic engineering.
# The human genome project, completed in 2001, offered the first comprehensive list of all human genes.
#
# - Yet, to fully understand how our cells function, and the origin of disease,
# - we need accurate maps that tell us how these genes and other cellular components interact with each other.
# + [markdown] slideshow={"slide_type": "slide"}
# # Security: Fighting Terrorism.
# Terrorism is one of the maladies of the 21st century, absorbing significant resources to combat it worldwide.
#
# - **Network thinking** is increasingly present in the arsenal of various law enforcement agencies in charge of limiting terrorist activities.
# - To disrupt the financial network of terrorist organizations
# - to map terrorist networks
# - to uncover the role of their members and their capabilities.
#
# - Using social networks to capture <NAME>
# - Capturing of the individuals behind the March 11, 2004 Madrid train bombings **through the examination of the mobile call network**.
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Epidemics: From forecasting to halting deadly viruses.
#
# While the **H1N1 pandemic** was not as devastating as it was feared at the beginning of the outbreak in 2009, it gained a special role in the history of epidemics: it was **the first pandemic whose course and time evolution was accurately predicted months before the pandemic reached its peak**.
#
# - Before 2000 epidemic modeling was dominated by **compartment models**, assuming that everyone can infect everyone else one word the same socio-physical compartment.
# - The emergence of a network-based framework has fundamentally changed this, offering a new level of predictability in epidemic phenomena.
#
# ### In January 2010 network science tools have predicted the conditions necessary for the emergence of viruses spreading through mobile phones.
#
# ### The first major mobile epidemic outbreak
# in the fall of 2010 in China, infecting over 300,000 phones each day, closely followed the predicted scenario.
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Brain Research: Mapping neural network.
# The human brain, consisting of hundreds of billions of interlinked neurons, is one of the least understood networks from the perspective of network science.
#
# The reason is simple:
# - we lack maps telling us which neurons link to each other.
# - The only fully mapped neural map available for research is that of the C.Elegans worm, with only 300 neurons.
#
# Driven by the potential impact of such maps, in 2010 the **National Institutes of Health** has initiated the Connectome project, aimed at developing the technologies that could provide an accurate neuron-level map of mammalian brains.
#
# + [markdown] slideshow={"slide_type": "slide"}
# # The Bridges of Konigsberg
#
# + [markdown] slideshow={"slide_type": "slide"}
# <img src = './img/konigsberg.png' width = 500>
# + [markdown] slideshow={"slide_type": "fragment"}
# # Can one walk across the seven bridges and never cross the same bridge twice and get back to the starting place?
#
# + [markdown] slideshow={"slide_type": "subslide"}
# Can one walk across the seven bridges and never cross the same bridge twice and get back to the starting place?
#
# <img src ='./img/euler.png' width = 300>
# + [markdown] slideshow={"slide_type": "slide"}
# # Euler’s theorem (1735):
#
# - If a graph has more than two nodes of odd degree, there is no path.
# - If a graph is connected and has no odd degree nodes, it has at least one path.
#
# + [markdown] slideshow={"slide_type": "slide"}
# COMPONENTS OF A COMPLEX SYSTEM
#
# # Networks and graphs
# - components: nodes, vertices N
# - interactions: links, edges L
# - system: network, graph (N,L)
#
# + [markdown] slideshow={"slide_type": "subslide"}
# network often refers to real systems
# - www,
# - social network
# - metabolic network.
#
# Language: (Network, node, link)
#
# + [markdown] slideshow={"slide_type": "subslide"}
# graph: mathematical representation of a network
# - web graph,
# - social graph (a Facebook term)
#
# Language: (Graph, vertex, edge)
#
#
# # G(N, L)
# + [markdown] slideshow={"slide_type": "subslide"}
# <img src = './img/net.png' width = 800>
# + [markdown] slideshow={"slide_type": "subslide"}
# # CHOOSING A PROPER REPRESENTATION
#
# The choice of the proper network representation determines our ability to use network theory successfully.
#
# In some cases there is a unique, unambiguous representation.
# In other cases, the representation is by no means unique.
#
# For example, the way we assign the links between a group of individuals will determine the nature of the question we can study.
#
# + [markdown] slideshow={"slide_type": "slide"}
# If you connect individuals that work with each other, you will explore the professional network.
#
# http://www.theyrule.net
# + [markdown] slideshow={"slide_type": "subslide"}
# If you connect those that have a romantic and sexual relationship, you will be exploring the sexual networks.
#
# + [markdown] slideshow={"slide_type": "subslide"}
# If you connect individuals based on their first name (all Peters connected to each other), you will be exploring what?
#
# # It is a network, nevertheless.
#
# + [markdown] slideshow={"slide_type": "subslide"}
# # UNDIRECTED VS. DIRECTED NETWORKS
#
# + [markdown] slideshow={"slide_type": "subslide"}
# # Undirected
# Links: undirected
# - co-authorship
# - actor network
# - protein interactions
# + slideshow={"slide_type": "subslide"}
# %matplotlib inline
import matplotlib.pyplot as plt
import networkx as nx
Gu = nx.Graph()
for i, j in [(1, 2), (1, 4), (4, 2), (4, 3)]:
Gu.add_edge(i,j)
nx.draw(Gu, with_labels = True)
# + [markdown] slideshow={"slide_type": "subslide"}
# # Directed
# Links: directed
# - urls on the www
# - phone calls
# - metabolic reactions
# + slideshow={"slide_type": "subslide"}
import networkx as nx
Gd = nx.DiGraph()
for i, j in [(1, 2), (1, 4), (4, 2), (4, 3)]:
Gd.add_edge(i,j)
nx.draw(Gd, with_labels = True)
# + [markdown] slideshow={"slide_type": "subslide"}
# <img src = './img/networks.png' width = 1000>
# + [markdown] slideshow={"slide_type": "subslide"}
# # Degree, Average Degree and Degree Distribution
#
# + slideshow={"slide_type": "subslide"}
nx.draw(Gu, with_labels = True)
# + [markdown] slideshow={"slide_type": "subslide"}
# # Undirected network:
# Node degree: the number of links connected to the node.
# ## $k_1 = k_2 = 2, k_3 = 3, k_4 = 1$
# + slideshow={"slide_type": "subslide"}
nx.draw(Gd, with_labels = True)
# + [markdown] slideshow={"slide_type": "subslide"}
# # Directed network
# In directed networks we can define an in-degree and out-degree. The (total) degree is the sum of in-and out-degree.
#
# ## $k_3^{in} = 2, k_3^{out} = 1, k_3 = 3$
#
# Source: a node with $k^{in}= 0$; Sink: a node with $k^{out}= 0$.
#
# + [markdown] slideshow={"slide_type": "subslide"}
# For a sample of N values: $x_1, x_2, ..., x_N$:
#
# # Average(mean):
#
# ## $<x> = \frac{x_1 +x_2 + ...+x_N}{N} = \frac{1}{N}\sum_{i = 1}^{N} x_i$
# + [markdown] slideshow={"slide_type": "subslide"}
# For a sample of N values: $x_1, x_2, ..., x_N$:
#
# # The nth moment:
#
# ## $<x^n> = \frac{x_1^n +x_2^n + ...+x_N^n}{N} = \frac{1}{N}\sum_{i = 1}^{N} x_i^n$
# + [markdown] slideshow={"slide_type": "subslide"}
# For a sample of N values: $x_1, x_2, ..., x_N$:
#
# # Standard deviation:
#
# ## $\sigma_x = \sqrt{\frac{1}{N}\sum_{i = 1}^{N} (x_i - <x>)^2}$
# + slideshow={"slide_type": "subslide"}
import numpy as np
x = [1, 1, 1, 2, 2, 3]
np.mean(x), np.sum(x), np.std(x)
# + [markdown] slideshow={"slide_type": "subslide"}
# For a sample of N values: $x_1, x_2, ..., x_N$:
#
# # Distribution of x:
#
# ## $p_x = \frac{The \: frequency \: of \: x}{The\: Number \:of\: Observations}$
#
# 其中,$p_x 满足 \sum_i p_x = 1$
# + slideshow={"slide_type": "subslide"}
# 直方图
plt.hist(x)
plt.show()
# + slideshow={"slide_type": "subslide"}
from collections import defaultdict, Counter
freq = defaultdict(int)
for i in x:
freq[i] +=1
freq
# + slideshow={"slide_type": "subslide"}
freq_sum = np.sum(freq.values())
freq_sum
# + slideshow={"slide_type": "subslide"}
px = [float(i)/freq_sum for i in freq.values()]
px
# + slideshow={"slide_type": "subslide"}
plt.plot(freq.keys(), px, 'r-o')
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# # Average Degree
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Undirected
#
# # $<k> = \frac{1}{N} \sum_{i = 1}^{N} k_i = \frac{2L}{N}$
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Directed
# # $<k^{in}> = \frac{1}{N} \sum_{i=1}^N k_i^{in}= <k^{out}> = \frac{1}{N} \sum_{i=1}^N k_i^{out} = \frac{L}{N}$
# + [markdown] slideshow={"slide_type": "slide"}
# # Degree distribution
# P(k): probability that a randomly selected node has degree k
#
#
# $N_k = The \:number\: of \:nodes\:with \:degree\: k$
#
# ## $P(k) = \frac{N_k}{N}$
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Adjacency matrix
# $A_{ij} =1$ if there is a link between node i and j
#
# $A_{ij} =0$ if there is no link between node i and j
# + slideshow={"slide_type": "slide"}
plt.figure(1)
plt.subplot(121)
pos = nx.spring_layout(Gu) #定义一个布局,此处采用了spring布局方式
nx.draw(Gu, pos, with_labels = True)
plt.subplot(122)
nx.draw(Gd, pos, with_labels = True)
# + [markdown] slideshow={"slide_type": "subslide"}
# # Undirected
# $A_{ij} =1$ if there is a link between node i and j
#
# $A_{ij} =0$ if there is no link between node i and j
#
# ## $A_{ij}=\begin{bmatrix} 0&1 &0 &1 \\ 1&0 &0 &1 \\ 0 &0 &0 &1 \\ 1&1 &1 & 0 \end{bmatrix}$
# + [markdown] slideshow={"slide_type": "subslide"}
# # Undirected
#
# 无向网络的矩阵是对称的。
#
# ## $A_{ij} = A_{ji} , \: A_{ii} = 0$
#
# ## $k_i = \sum_{j=1}^N A_{ij}, \: k_j = \sum_{i=1}^N A_{ij} $
#
# 网络中的链接数量$L$可以表达为:
#
# ## $ L = \frac{1}{2}\sum_{i=1}^N k_i = \frac{1}{2}\sum_{ij}^N A_{ij} $
# + [markdown] slideshow={"slide_type": "subslide"}
# # Directed
# $A_{ij} =1$ if there is a link between node i and j
#
# $A_{ij} =0$ if there is no link between node i and j
#
# ## $A_{ij}=\begin{bmatrix} 0&0 &0 &0 \\ 1&0 &0 &1 \\ 0 &0 &0 &1 \\ 1&0 &0 & 0 \end{bmatrix}$
#
# Note that for a directed graph the matrix is not symmetric.
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Directed
# ## $A_{ij} \neq A_{ji}, \: A_{ii} = 0$
# ## $k_i^{in} = \sum_{j=1}^N A_{ij}, \: k_j^{out} = \sum_{i=1}^N A_{ij} $
# ## $ L = \sum_{i=1}^N k_i^{in} = \sum_{j=1}^N k_j^{out}= \frac{1}{2}\sum_{i,j}^N A_{ij} $
# + [markdown] slideshow={"slide_type": "slide"}
# # WEIGHTED AND UNWEIGHTED NETWORKS
#
# ## $A_{ij} = W_{ij}$
# + [markdown] slideshow={"slide_type": "slide"}
# # BIPARTITE NETWORKS
#
# + [markdown] slideshow={"slide_type": "slide"}
# bipartite graph (or bigraph) is a graph whose nodes can be divided into two disjoint sets U and V such that every link connects a node in U to one in V; that is, U and V are independent sets.
#
# - Hits algorithm
# - recommendation system
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Ingredient-Flavor Bipartite Network
#
# <img src = './img/bipartite.png' width = 800>
# + [markdown] slideshow={"slide_type": "slide"}
# # Path 路径
# A path is a sequence of nodes in which each node is adjacent to the next one
# - In a directed network, the path can follow only the direction of an arrow.
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Distance 距离
#
# The distance (shortest path, geodesic path) between two nodes is defined as the number of edges along the shortest path connecting them.
#
# > If the two nodes are disconnected, the distance is **infinity**.
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Diameter 直径
#
# **Diameter $d_{max}$** is the maximum distance between any pair of nodes in the graph.
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Shortest Path 最短路径
# The path with the shortest length between two nodes (distance).
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Average path length/distance, $<d>$ 平均路径长度
#
#
# The average of the shortest paths for all pairs of nodes.
#
#
# - for a **directed graph**: where $d_{ij}$ is the distance from node i to node j
#
# ## $<d> = \frac{1}{2 L }\sum_{i, j \neq i} d_{ij}$
#
# > 有向网络当中的$d_{ij}$数量是链接数量L的2倍
#
# - In an **undirected** graph $d_{ij} =d_{ji}$ , so we only need to count them once
#
# > 无向网络当中的$d_{ij}$数量是链接数量L
#
#
# ## $<d> = \frac{1}{L }\sum_{i, j > i} d_{ij}$
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Cycle 环
# A path with the same start and end node.
#
# + [markdown] slideshow={"slide_type": "slide"}
# # CONNECTEDNESS
#
# + [markdown] slideshow={"slide_type": "subslide"}
# # Connected (undirected) graph
#
# > In a connected **undirected** graph, any two vertices can be joined by a path.
#
# > A disconnected graph is made up by two or more connected components.
#
# - Largest Component: Giant Component
# - The rest: Isolates
#
# ## Bridge 桥
# if we erase it, the graph becomes disconnected.
# + [markdown] slideshow={"slide_type": "subslide"}
# ## The adjacency matrix of a network with several components can be written in a block-diagonal form, so that nonzero elements are confined to squares, with all other elements being zero:
#
# <img src = './img/block.png' width = 600>
# + [markdown] slideshow={"slide_type": "subslide"}
# # Strongly connected *directed* graph 强连通有向图
#
# has a path from each node to every other node and vice versa (e.g. AB path and BA path).
#
# # Weakly connected directed graph 弱连接有向图
# it is connected if we disregard the edge directions.
#
# Strongly connected components can be identified, but not every node is part of a nontrivial strongly connected component.
#
# + [markdown] slideshow={"slide_type": "subslide"}
# # In-component -> SCC ->Out-component
#
# - In-component: nodes that can reach the **scc** (strongly connected component 强连通分量或强连通子图)
# - Out-component: nodes that can be reached from the scc.
#
# > 万维网的蝴蝶结模型🎀 bowtie model
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# # Clustering coefficient 聚集系数
#
# + [markdown] slideshow={"slide_type": "subslide"}
# # Clustering coefficient 聚集系数
# what fraction of your neighbors are connected? Watts & Strogatz, Nature 1998.
#
# # 节点$i$的朋友之间是否也是朋友?
#
# ## Node i with degree $k_i$ 节点i有k个朋友
#
# > ## $e_i$ represents the number of links between the $k_i$ neighbors of node i.
#
# > ## 节点i的k个朋友之间全部是朋友的数量 $\frac{k_i(k_i -1)}{2}$
#
#
# # $C_i = \frac{2e_i}{k_i(k_i -1)}$
#
# $C_i$ in [0,1]
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# # 节点的**聚集系数**
#
# <img src = './img/cc.png' width = 500>
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# # Global Clustering Coefficient 全局聚集系数(i.e., Transtivity 传递性)
#
# > triangles 三角形
# > triplets 三元组
#
# - A triplet consists of three connected nodes.
# - A triangle therefore includes three closed triplets
# - A triangle forms three **connected triplets**
# - **A connected triplet** is defined to be a connected subgraph consisting of three vertices and **two edges**.
#
# ## $C = \frac{\mbox{number of closed triplets}}{\mbox{number of connected triplets of vertices}}$
#
# ## $C = \frac{3 \times \mbox{number of triangles}}{\mbox{number of connected triplets of vertices}}$
#
# + slideshow={"slide_type": "subslide"}
G1 = nx.complete_graph(4)
pos = nx.spring_layout(G1) #定义一个布局,此处采用了spring布局方式
nx.draw(G1, pos = pos, with_labels = True)
# + slideshow={"slide_type": "fragment"}
print(nx.transitivity(G1))
# + slideshow={"slide_type": "subslide"}
G2 = nx.Graph()
for i, j in [(1, 2), (1, 3), (1, 0), (3, 0)]:
G2.add_edge(i,j)
nx.draw(G2,pos = pos, with_labels = True)
# + slideshow={"slide_type": "fragment"}
print(nx.transitivity(G2))
# 开放三元组有5个,闭合三元组有3个
# + slideshow={"slide_type": "subslide"}
G3 = nx.Graph()
for i, j in [(1, 2), (1, 3), (1, 0)]:
G3.add_edge(i,j)
nx.draw(G3, pos =pos, with_labels = True)
# + slideshow={"slide_type": "fragment"}
print(nx.transitivity(G3))
# 开放三元组有3个,闭合三元组有0个
# + [markdown] slideshow={"slide_type": "slide"}
# THREE CENTRAL QUANTITIES IN NETWORK SCIENCE
# - A. Degree distribution: $p_k$
# - B. Path length: $<d>$
# - C. Clustering coefficient: $C_i$
#
# + [markdown] slideshow={"slide_type": "subslide"}
# # Typical Network Science Research
#
# - Discovering, Modeling, Verification
# - WATTSDJ,STROGATZSH.Collective dynamics of‘small-world’ networks. Nature, 1998, 393(6684): 440–442.
# - <NAME>, <NAME>. Emergence of scaling in random networks. Science, 1999, 286(5439): 509-512.
# + [markdown] slideshow={"slide_type": "subslide"}
# # Typical Math Style
# <NAME> & <NAME>, The average distance in random graphs with given expected degree,. PNAS, 19, 15879-15882 (2002).
# + [markdown] slideshow={"slide_type": "subslide"}
# # Typical Physical Style
# A.-L.Barabási,R.Albert,H.Jeong Mean-field theory for scale-free random networks. Physica A 272, 173–187 (1999).
# + [markdown] slideshow={"slide_type": "subslide"}
# # Typical Computer Science Style
#
# - Community detection
# - Link prediction
# - Recommendation algorithms
# + [markdown] slideshow={"slide_type": "subslide"}
# # Typical control sytle
# Controllability of Complex Networks
#
# <NAME>, <NAME>, Barabási A L. Nature, 2011, 473(7346): 167-173.
# + [markdown] slideshow={"slide_type": "slide"}
# # 阅读材料
# - Barabasi 2016 Network Science. Cambridge
# - 汪小帆、李翔、陈关荣 2012 网络科学导论. 高等教育出版社
# - 梅拉妮·米歇尔 2011 复杂,湖南科学技术出版社
# - 菲利普-鲍尔 2004 预知社会:群体行为的内在法则,当代中国出版社
# - 巴拉巴西 2007 链接:网络新科学 湖南科技出版社
# -
| code/15.network_science_intro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Index of Runs and Variables
# These notebooks document model runs that are avaiable for analysis.
# +
# %matplotlib inline
import cosima_cookbook as cc
import dataset
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
from ipywidgets import widgets, interact, fixed, interactive
from IPython.display import display, HTML
import tqdm
# -
# Currently, the cookbook searches for NetCDF4 files in the following directories
cc.netcdf_index.directoriesToSearch
# We first generate a database of all variables in all netCDF4 files found within these directories. Note that this needs only be called once as the database will persist between sessions. If new output files are added to the data directories, build_index() will only import the new files.
#
# If this database ever becomes corrupted, it can be be safely deleted and will be recreated whenever build_index() is next called.
cc.build_index()
# This index of all variables is stored in a SQL database. If needed, it can be accessed directly using the `dataset` module.
cc.netcdf_index.database_file
db = dataset.connect(cc.netcdf_index.database_url)
rows = db.query('select * from ncfiles where variable = "tau_x" and experiment = "025deg_jra55_ryf_spinup1" limit 5')
for row in rows:
print(row)
# Let's bring this database into memory as a Pandas DataFrame for further analysis.
data = []
for row in tqdm.tqdm_notebook(db['ncfiles'].all(), total = 1180000):
data.append(row)
df = pd.DataFrame(data)
df
# Many of the output files have names of the form `output__123_45.nc`. Here we constuct a more generalized name for this output file, namely the regular expression `output__\d+_\d+.nc`.
# [To do: add basename_pattern to build_index() ]
# +
pat = '(?P<root>[^\d]+)(?P<index>__\d+_\d+)?(?P<indexice>.\d+\-\d+)?(?P<ext>\.nc)'
repl = lambda m: m.group('root') + ('__\d+_\d+' if m.group('index') else '') + ('.\d+-\d+' if m.group('indexice') else '')+ m.group('ext')
df['basename_pattern'] = df.basename.str.replace(pat, repl)
# -
display(df)
# ### Number of runs per configuration/experiment
# The data directory contains several model __configurations__ (e.g. mom01v5 or mom025)
#
# Each configuration contains a number of __experiments__ (e.g. KDS75 or KDS75_wind)
#
# The output is a set of several __runs__ (e.g. output266)
table = pd.pivot_table(df, index=["rootdir", "configuration",],
values=['experiment'],
aggfunc=lambda x: len(x.unique()))
table
# This table shows the number of experiments that have been performed for each configuration.
# ## MOM-SIS 0.1$^\circ$
#
# | Experiment Name | Description |
# |-----------------|-----------------|
# |GFDL50 | Original simulation with 50 vertical levels. Ran from Levitus for about 60 years, but data output only saved from about year 40.|
# |KDS75 | Branched from GFDL50 at year 45 (re-zeroed), but with Kial Stewart's 75 level scheme. Has now run for 103 years. Years 90-100 have 5-daily output.|
# | KDS75_wind | Short (5-year) Antarctic wind perturbation case, branched from KDS75 at year 40.|
# | KDS75_PI | <NAME>'s Poleward Intensification wind experiment. Branched from KDS75 at year 70, will run until year 100 with 5-daily output for the last decade|
# | KDS75_UP | <NAME>'s Increased winds case. Branched from KDS75 at year 70, will run until year 100 with 5-daily output for the last decade. (In Progress) |
# +
table = pd.pivot_table(df, index=["configuration", "experiment",],
values=['run'],
aggfunc=lambda x: len(x.unique()))
table.query('configuration == "mom01v5"')
# +
table = pd.pivot_table(df, index=["experiment", "basename_pattern", "variable"],
values=['ncfile'],
aggfunc=lambda x: len(x.unique()))
with pd.option_context('display.max_rows', None):
display(table.query('experiment == "KDS75"'))
# -
# ### Output frequency over time
# +
import re, dask
import xarray as xr
pat = '(?P<root>[^\d]+)(?P<index>__\d+_\d+)?(?P<indexice>\.\d+\-\d+)?(?P<ext>\.nc)'
regex = re.compile(pat)
variable = 'average_T1'
rows = db.query("SELECT ncfile, basename from ncfiles where experiment = 'KDS75_PI' and variable = '{}' "
"order by ncfile".format(variable))
def handle(row):
m = regex.match(row['basename'])
basename_pattern = m.group('root') + ('__\d+_\d+' if m.group('index') else '') + ('.\d+-\d+' if m.group('indexice') else '')+ m.group('ext')
print(row['ncfile'], row['basename'], basename_pattern)
dsx = xr.open_dataset(row['ncfile'], decode_times=False)
return (basename_pattern, dsx.average_T1.values, dsx.average_DT.values)
rows = list(rows)
bag = dask.bag.from_sequence(rows)
bag = bag.map(handle)
# -
from distributed import Client
from distributed.diagnostics.progressbar import progress
client = Client()
futures = client.compute(bag)
display(progress(futures))
# +
result = futures.result()
l = []
for basename, T1, DT in result:
for t1, dt in zip(T1, DT):
l.append([basename, t1, dt])
df = pd.DataFrame(l, columns = ['basename', 'T1', 'DT'])
df = df.sort_values(['basename', 'T1'])
# +
plt.figure(figsize=(24,4))
labels = []
for n, g in enumerate(df.groupby('basename')):
labels.append(g[0])
T1 = g[1].T1
DT = g[1].DT
plt.barh(n*np.ones_like(DT[::2]), DT[::2], height=1, left=T1[::2])
plt.barh(n*np.ones_like(DT[1::2]), DT[1::2], height=1, left=T1[1::2])
plt.yticks(range(len(labels)), labels)
#plt.xlim(34000, 34300)
plt.show()
# -
# ## MOM-SIS 0.25$^\circ$ Diagnostics
#
# | Experiment Name | Description |
# |-----------------|-----------------|
# |mom025_nyf | Original simulation, rerun from WOA13 initial conditions.|
# |mom025_nyf_salt | As above, with new ew salt restoring file from WOA13 surface data. (Not running yet)|
# ## ACCESS-OM2-025 Preliminary Analysis
#
# |** Run Name** | **Forcing** | ** Run ** | ** Status **|
# |--------------|---------|-------------------------------------------------|-------------|
# |025deg_jra55_ryf_spinup1 | JRA55 RYF9091| This is our initial 0.25° test. Ran for a decade before sea ice build-up overwhelmed us! | Aborted after 10 years.|
# |025deg_jra55_ryf_spinup2 | JRA55 RYF9091| This is our initial 0.25° test with the sea ice parameter fixed. Less sea ice buildup, but there seems to be a problem with salinity conservation. It seems we are not doing runoff properly ... | Stopped at 50 years.|
# |025deg_jra55_ryf_spinup3 | JRA55 RYF9091| Third attempt at 0.25° test. This run is very unstable, and we think it might be something to do with runoff. Will try to fix this and start again. | Up to 8 years.|
# |025deg_jra55_ryf | JRA55 RYF9091| Latest attempt at 0.25° test. | Started 5/8/17|
#
#
# ## ACCESS-OM2 Preliminary Analysis
#
# |** Run Name** | **Forcing** | ** Run ** | ** Status **|
# |--------------|---------|-------------------------------------------------|-------------|
# |1deg_jra55_ryf_spinup1 | JRA55 RYF9091| A short 10 year spinup with first pre-release code. Had bugs in runoff and salt fluxes.| Aborted after 10 years.|
# |1deg_jra55_ryf | JRA55 RYF9091| second attempt at 1° test. | Up to 50 years.|
#
#
| configurations/MOM_Run_Summary.ipynb |
# # Test-Driven Development in ACL2
#
# Professional programmers know a simple fact: If your code isn't **extensively** tested, it's not working. In fact, this is the biggest reason why code reuse is so important. It's not that writing a linked list is hard. You learned how to do that in your first or second programming class. The reason why we use `List<Integer>` instead of writing our own linked list of integers in Java is that `List<Integer>` has been tested extensively, so we can be confident that it works.
#
# Let me be very clear. Testing is not a thing that you do in class because your professors make you. **Testing is something that you should always do as a a programmer.** In fact, learning how to to test is an essential part of becoming a professional. It's what separates us from the folks who write code like this:
#
# <img src="https://miro.medium.com/max/500/0*vsvLVt-w4WivlTOn.jpg">
#
# To illustrate how to test functions in ACL2, let's begin with the definition of two functions we defined in a previous tutorial:
# +
(defsnapshot triangular-definition)
(definec triangular (n :nat) :nat
(if (zp n)
0
(+ (triangular (- n 1)) n)))
(defsnapshot zeta2-definition)
(definec zeta2 (n :nat) :rational
(if (zp n)
0
(+ (zeta2 (- n 1))
(/ 1 (* n n)))))
# -
# ## Unit Tests
#
# At the very least, you should always write several **unit tests** for each of your functions. Professional programmers routinely write dozens of unit tests per function, but I will only ask you to write five for each.
#
# A **unit test** simply checks that the function returns the right value for some given inputs. I usually test that the function does the right thing for a few inputs that I compute by hand. Later, if I discover a bug in the code, I add that particular input to my collection of tests, so the collection of tests is always growing.
#
# Here is a simple unit test for `triangular`. We know that `(triangular 3) = 1+2+3 = 6`. In ACL2, you can perform this unit test using the function `check-expect`, as follows:
(check-expect (triangular 3) 6)
# The output should look like this:
#
# ACL2S !>>(CHECK-EXPECT (TRIANGULAR 3) 6)
# :PASSED
#
# The `:PASSED` is the indication that the test passes, as expected.
#
# To see what happens when ACL2 discovers an error, you can change the value "6" in that `check-expect`:
(check-expect (triangular 3) -9999999)
# It can sometimes be more convenient to leave the expected value as an expression without fully computing it, as in the following:
(check-expect (triangular 5) (+ 1 2 3 4 5))
# ## Properties and Randomized Testing
#
# Every programming language comes with a unit-testing framework, e.g., JUnit for Java. These are great for checking what your code does for specific inputs, but sometimes programmers we often have more general beliefs about our programs that should also be checked. For example, you may believe that no matter what the input to your function, it will **never** return null. To test properties like this, you need to test with a large number of inputs, and this is where testing with **random inputs** makes sense.
#
# For instance, you may remember that $1+2+\cdots+n=\frac{n(n+1)}{2}$. ACL2 provides the command `test?` to test properties like this. What ACL2 will do is to try many different random values of $n$ and see if the property holds for each value. Let's see how it works!
#
(test? (equal (triangular n)
(/ (* n (+ n 1)) 2)))
# The result probably looks something like this:
#
# **Summary of Cgen/testing**
# We tested 223 examples across 1 subgoals, of which 222 (222 unique)
# satisfied the hypotheses, and found 3 counterexamples and 219 witnesses.
#
# We falsified the conjecture. Here are counterexamples:
# [found in : "top"]
# -- ((N 8/49))
# -- ((N #C(2/3 1)))
# -- ((N -45))
#
# Cases in which the conjecture is true include:
# [found in : "top"]
# -- ((N '((T) T)))
# -- ((N '(A)))
# -- ((N '(1 . T))
#
# Test? found a counterexample.
#
# This is not what we expected! ACL2 discovered that our property is not true. (Actually, a cynic would say that **is** as as expected. Programmers are often mistaken in their beliefs about their programs.)
#
# ACL2 proactively helps us debug the program by giving us random inputs where the program failed to satisfy the property. In particular, we can see above that the program does not work correctly when `n` is `8/49`, `#c(2/3 1)` (which is a complex number), or `-45`. This should give you a very good idea of what went wrong. We had intended `n` to be a natural number, but ACL2 is finding that our property is not true in some cases where `n` is a rational, or a complex number, or a negative integer. What we can see is that our property is buggy, though if we're lucky the program is correct.
#
# Let's fix the property. We believe that $1+ 2+\cdots+n=\frac{n(n+1)}{2}$, but only when $n$ is a natural number. This is where logic comes in. We can express this property using logical implication. Oh, one more thing: The ACL2 built-in `(natp n)` is true precisely when `n` is a natural number.
(test? (implies (natp n)
(equal (triangular n)
(/ (* n (+ n 1)) 2))))
# When you submit that test to ACL2, you should see something like the following:
#
# **Summary of Cgen/testing**
# We tested 3000 examples across 3 subgoals, of which 2939 (2939 unique)
# satisfied the hypotheses, and found 0 counterexamples and 2939 witnesses.
#
# Cases in which the conjecture is true include:
# [found in : "top"]
# -- ((N 732))
# -- ((N 14))
# -- ((N 767))
#
# Test? succeeded. No counterexamples were found.
#
# The important line is the last one. It says that no counterexamples were found, which means that all random values of `n` satisfied the property; they all passed! In the first paragraph, ACL2 tells us that it tried 3,000 random values of `n`, of which 2,939 were actually different values of `n`. Among those 2,939 values were 732, 14, and 767. They all passed, so that's 2,939 out of 2,939. You should probably feel pretty confident that the property is right. It's still possible that it fails in some strange cases, but the odds are in your favor.
# We can do the same thing with `(zeta2 n)`. Euler discovered that $\zeta(2) = \pi^2/6$. $\zeta(2)$ is an infinite sum, and `(zeta2 n)` is an approximation to $\zeta(2)$ by adding up some of those terms. So we have that
#
# $$(zeta2\, n) \le \zeta(2) = \frac{\pi^2}/6 \le \frac{3.1416^2}{6}$$
#
# We can test this with ACL2:
(test? (<= (zeta2 n) (/ (expt 3.1416 2) 6)))
# This is a mixed success. ACL2 reports that
#
# **Summary of Cgen/testing**
# We tested 3000 examples across 3 subgoals, of which 2762 (2762 unique)
# satisfied the hypotheses, and found 0 counterexamples and 2762 witnesses.
#
# Cases in which the conjecture is true include:
# [found in : "top"]
# -- ((N '(-9/14 1/2 -1)))
# -- ((N '(2 . T)))
# -- ((N NIL))
#
# Test? succeeded. No counterexamples were found.
#
# On the one hand, ACL2 did not find any counterexamples, so all random values of `n` succeeded. On the other hand, when you look at some of those random values of `n`, it's clear that ACL2 is **not** testing with appropriate values. After all, we intend that `n` is a natural number, so why are we testing when `n=NIL`?
#
# Again, we missed a hypothesis in our property. Let's fix it.
(test? (implies (natp n)
(<= (zeta2 n) (/ (expt 3.1416 2) 6))))
# That's more like it! This time ACL2 tested our property with 2,882 values of `n`, and the property passed all those tests. And, each one of those random values of `n` was a natural number, like 86, 30, and 3. It's still possible that the property fails for some other values of `n`, but this does make me confident that the property is true.
#
# **Beware of overconfidence!** We sais above that "this does make me confident that the property is true," but we did not say "this does make confident that the program works." The property we mentioned, namely that `(zeta2 n)` is less than or equal to $\pi^2/6$ is not enough by itself to guarantee that the program works. For example, it could be the case that the function `(zeta2 n)` always returns 0, and this property would be true! In the testing community, these properties are known as **little theories**. They can help you gain confidence in your program, even though they do give you total guarantee of success. Such properties are very common in practice, but you should always remember their limitations. It's possible to give more reassuring properties. For example, if we could show that both of these properties are true, I would feel very confident in the program:
#
# * $\zeta_2(n) \le \pi^2/6$
# * $\epsilon > 0 \rightarrow [(\exists N) (n>N \rightarrow \pi^2/6 - \epsilon \le \zeta_2(n)]$
#
# When it really matters that your program works correctly, you may need to test both of the properties above, but testing the second property is very much harder than the first property, because of the $\exists N$. (You may recall how much easier it is to prove properties with $\forall x$ than with $\exists x$, since for a $\exists x$ you actually have to *find* an $x$ with the right property.)
# ## Properties and Proofs
#
# When it really matters that your program works correctly, randomized testing is better than just unit tests, but even randomized testing may not be enough. This is when you consider going the extra mile and **proving** that your property is true.
#
# ACL2 is more than a programming language; it is a state-of-the-art theorem prover. In industry, ACL2 is used in various settings where correctness is essential, e.g., in the design of hardware chips or mission-critical software. We'll use ACL2 to (try to) prove the properties we tested above.
#
# To prove a property in ACL2, you have two choices
#
# * `(thm ...property...)`
# * `(defthm name ...property...)`
#
# The first form tries to prove the property and then essentially forgets about it. The second form gives the property a name, and remembers the property. If ACL2 remembers a property, it is able to use later when proving other properties. This is the way that ACL2 is used in industry. It is almost never the case that ACL2 discovers the proof of your property automatically. So what you have to do is prove a sequence of theorems that lead up to the property you want. Industrial proof efforts will often require hundreds (or thousands) of these intermediate theorems!
#
# But let's just try to prove the first property and see what ACL2 can do.
(thm (implies (natp n)
(equal (triangular n)
(/ (* n (+ n 1)) 2))))
# Success! The last lines of the output look like this
#
# **Summary of Cgen/testing**
# We tested 2000 examples across 2 subgoals, of which 1930 (1930 unique)
# satisfied the hypotheses, and found 0 counterexamples and 1930 witnesses.
#
# Cases in which the conjecture is true include:
# [found in : "Goal"]
# -- ((N 667))
# -- ((N 7))
# -- ((N 1))
#
# Proof succeeded.
#
# The last line is the most important one; it says that the proof succeeded, which means that our conjecture about `(triangular n)` is true. As you can see from the output, ACL2 first used randomized testing to check the property. After all, if testing revealed that the property is false, there's no point in trying to find a proof.
#
# The output from ACL2 is actually quite verbose. Before getting this summary at the end, ACL2 does two things. First, it describes the proof that it found. In this particular case, ACL2 used mathematical induction on the variable `n`. Second, it list all the facts that it used in the proof. For example, this proof used the fact called `|(expt (+ x y) 2)|`, which is the familiar result from algebra that $(x+y)^2 = x^2 + 2xy + y^2$. That's what we meant earlier when we said that ACL2 uses previously-proved properties as it's trying to prove a new property.
#
# If you use `thm` to prove a property property, ACL2 will essentially forget it after it's done with the proof, so it will not be able to use this fact later, as it tries to prove something else. If we want ACL2 to remember the property so it can be used later, you use `defthm` instead of `thm`. Let 's do that!
(defthm triangular-formula
(implies (natp n)
(equal (triangular n)
(/ (* n (+ n 1)) 2))))
# Again, the proof succeeds. The output is similar to what we saw with `thm` (ACL2 did find the same proof, after all), but instead of finishing with "Proof succeeded", ACL2 finished with the name of the theorem. That's the name that it will use to remember this fact, `TRIANGULAR-FORMULA` in this case.
# That worked very well! Let's do it again, this time with the property about `zeta2`:
(thm (implies (natp n)
(<= (zeta2 n) (/ (expt 3.1416 2) 6))))
# We were not as lucky this time! The proof
#
# ******** FAILED ********
#
# as ACL2 puts it. I.e., ACL2 was unable to find a proof of our property. However, you can see from the output that ACL2 tested the property 4,000 times, and each time the property worked. What we see here is that randomized testing hasn't found a bug, but ACL2 cannot find a proof that there aren't any bugs.
#
# This is very common. The truth is we got quite lucky with the proof of `TRIANGULAR-FORMULA` above. As we mentioned earlier, this is where you would have to come up with some intermediate properties that ACL2 can use to find the proof of the property we want. The process requires you to look at the proof attempt, try to understand why it failed, and use that to discover a key lemma that may be useful. Often, the "Key Checkpoints" that ACL2 mentions at the end of the failed proof attempt are good places to start. The tutorial on "The Method" describes this process in more detail. But for this specific property, finding a proof in ACL2 would actually be quite difficult, so we will simply move on. But if you're really interested in this, a proof of this fact (called the Basel Problem) would make a wonderful Master's thesis!
| context/acl2-notebooks/programming-tutorial/.ipynb_checkpoints/06 - Test-Driven Development-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# # Effect of September-11 Terrorist Attack on Hate Crimes Against Muslims in the United Stateds
# #### Course: Data Bootcamp
# #### Report Author: <NAME>
# #### Report Date: May 12, 2017
# #### New York University
# #### Leonard N. Stern School of Business
#
# 
#
# Image Source = [http://www.huffingtonpost.com/entry/hate-crimes-muslims-since-911_us_57e00644e4b04a1497b59970](http://www.huffingtonpost.com/entry/hate-crimes-muslims-since-911_us_57e00644e4b04a1497b59970)
# # Research Question:
#
# ### Main Research Question:
# #### Q: Did September-11 terrorist attack have an impact on hate crimes against Muslims? How much and what impact did the incident have?
#
# ### Analysis Questions: I will be using the data to answer the following questions. Answers to the following questions will be used to answer the main research question.
#
# 1. How have the hate crimes against Muslims changed in terms of number of incidents per year?
# 1. Did September-11 Terrorist Attack had an impact on the hate crimes against Muslims? If so, how much impact did September-11 Terrorist Attack had?
# 2. How have the hate crimes against All religion changed in terms of number of incidents per year?
# 3. What percentage of hate crimes motivated by religion identity target Muslims every year?
# 3. On average what percentages of attacks motivated by religion targetted Muslims, before and after the September-11 Terrorist Attack?
# # Data Source
#
# The project focuses on the affect of 9/11 incident on the change in the hate crimes against Muslims in the United States. The data used for the project has been collected as [Hate Crime Statistics](https://ucr.fbi.gov/hate-crime) through the [Uniform Crime Reporting (UCR) Program](https://ucr.fbi.gov/) of [Federal Bureau of Investigation (FBI)](https://www.fbi.gov/). The data is avialable on from FBI's website, where the data is reported on an yearly basis.
#
# ###### The data is available for years [1995 to 2015](https://ucr.fbi.gov/hate-crime), except 2009.
#
# #### For each year: The data has been divided into different tables, based on the following aspects:
# * Incidents and Offenses
# * Victims
# * Offenders
# * Location Type
# * Hate Crime by Jurisdication
#
# ###### The projects uses the data categorised based on Incidents and Offenses, since the data is categorised into different types of hate crimes including Anti-Religion. The project utelises the data from year 1995-2015.
#
# #### Limitations of the Data:
# * The data does not include the statistics for Hawaii. The data is not reported for Hawaii in the records.
# * The FBI collects data from independent law-enforcing agencies in different towns, cities, counties, metropolitan areas and university areas. Therefore, the data is contingent upon their reporting.
# * There is no data available for [2009](https://ucr.fbi.gov/hate-crime/2009).
# # Preliminaries
# +
import sys # system module
import pandas as pd # data package
import matplotlib.pyplot as plt # graphics module
import datetime as dt # date and time module
import numpy as np # foundation for pandas
import requests
from bs4 import BeautifulSoup
# %matplotlib inline
# check versions (overkill, but why not?)
print('Python version: ', sys.version)
print('Pandas version: ', pd.__version__)
print('Today: ', dt.date.today())
plt.style.use('ggplot')
# -
# # Data Import (2005 - 2015)
#
# Files for years 2005-2015 are available in [excel format](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.read_excel.html). The files are downloaded into the working directory, and then imported using [pandas library](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.read_excel.html). For the above-mentioned years,the data is available in [different tables](https://ucr.fbi.gov/hate-crime/2015/topic-pages/incidentsandoffenses_final) and these tables categorise the data differently. We have used Table1, since it categorises the hate criimes against religion into different religions, which is most appropriate for this project. For example, for year 2015, we will get Table 1 from the [source](https://ucr.fbi.gov/hate-crime/2015/tables-and-data-declarations/1tabledatadecpdf).
# # 2015 Data
# Download Source = [https://ucr.fbi.gov/hate-crime/2015/tables-and-data-declarations/1tabledatadecpdf](https://ucr.fbi.gov/hate-crime/2015/tables-and-data-declarations/1tabledatadecpdf)
#
# The file is saved locally as "table1_2015.xls"
url = 'table1_2015.xls'
headers = ['Incidents','Offenses','Victims1','Known offenders2']
data_2015 = pd.read_excel(url, skiprows=3, skipfooter=3, parse_cols="A,B,C,D,E", headers = None, names = ["Motivation","Incidents","Offenses","Victims","Known Offenders"])
religion_2015 = data_2015[12:27]
#columns = ['Incidents','Offenses', 'Victims','Known Offenders']
#original_2015 = religion_2015.copy(deep=True)
#for col in columns:
# new_val = religion_2015.iloc[5][col] + religion_2015[col][7:14].sum()
#print(new_val)
#religion_2015 = religion_2015.set_value('Anti-Other Religion',col,new_val)
#religion_2015.ix['Anti-Other Religion',col] = new_val
religion_2015
# # 2014 Data
# Download Source = [https://ucr.fbi.gov/hate-crime/2014/tables/table-1](https://ucr.fbi.gov/hate-crime/2014/tables/table-1)
#
# The file is saved locally as "table1_2014.xls"
url = 'table1_2014.xls'
headers = ['Incidents','Offenses','Victims1','Known offenders2']
data_2014 = pd.read_excel(url, skiprows=3, skipfooter=3, parse_cols="A,B,C,D,E",headers = None, names = ["Motivation","Incidents","Offenses","Victims","Known Offenders"])
religion_2014 = data_2014[9:17]
# # 2013 Data
# Download Source = [https://ucr.fbi.gov/hate-crime/2013/tables/1tabledatadecpdf/table_1_incidents_offenses_victims_and_known_offenders_by_bias_motivation_2013.xls](https://ucr.fbi.gov/hate-crime/2013/tables/1tabledatadecpdf/table_1_incidents_offenses_victims_and_known_offenders_by_bias_motivation_2013.xls)
#
# The file is saved locally as "table1_2013.xls"
url = 'table1_2013.xls'
headers = ['Incidents','Offenses','Victims1','Known offenders2']
data_2013 = pd.read_excel(url, skiprows=3, skipfooter=3, parse_cols="A,B,C,D,E",headers = None, names = ["Motivation","Incidents","Offenses","Victims","Known Offenders"])
religion_2013 = data_2013[9:17]
# # 2012 Data
# Download Source = [https://ucr.fbi.gov/hate-crime/2012/tables-and-data-declarations/1tabledatadecpdf/table_1_incidents_offenses_victims_and_known_offenders_by_bias_motivation_2012.xls](https://ucr.fbi.gov/hate-crime/2012/tables-and-data-declarations/1tabledatadecpdf/table_1_incidents_offenses_victims_and_known_offenders_by_bias_motivation_2012.xls)
#
# The file is saved locally as "table1_2012.xls"
url = 'table1_2012.xls'
headers = ['Incidents','Offenses','Victims1','Known offenders2']
data_2012 = pd.read_excel(url, skiprows=3, skipfooter=3, parse_cols="A,B,C,D,E",headers = None, names = ["Motivation","Incidents","Offenses","Victims","Known Offenders"])
religion_2012 = data_2012[8:16]
# # 2011 Data
# Download Source = [https://ucr.fbi.gov/hate-crime/2011/tables/table-1](https://ucr.fbi.gov/hate-crime/2011/tables/table-1)
#
# The file is saved locally as "table1_2011.xls"
url = 'table1_2011.xls'
headers = ['Incidents','Offenses','Victims1','Known offenders2']
data_2011 = pd.read_excel(url, skiprows=3, skipfooter=3, parse_cols="A,B,C,D,E",headers = None, names = ["Motivation","Incidents","Offenses","Victims","Known Offenders"])
religion_2011 = data_2011[8:16]
# # 2010 Data
# Download Source = [https://ucr.fbi.gov/hate-crime/2010/tables/table-1-incidents-offenses-victims-and-known-offenders-by-bias-motivation-2010.xls](https://ucr.fbi.gov/hate-crime/2010/tables/table-1-incidents-offenses-victims-and-known-offenders-by-bias-motivation-2010.xls)
#
# The file is saved locally as "table1_2010.xls"
url = 'table1_2010.xls'
headers = ['Incidents','Offenses','Victims1','Known offenders2']
data_2010 = pd.read_excel(url, skiprows=3, skipfooter=3, parse_cols="A,B,C,D,E",headers = None, names = ["Motivation","Incidents","Offenses","Victims","Known Offenders"])
religion_2010 = data_2010[7:15]
# # 2009 Data
# Download Source = [https://ucr.fbi.gov/hate-crime/2010/tables/table-1-incidents-offenses-victims-and-known-offenders-by-bias-motivation-2010.xls](https://ucr.fbi.gov/hate-crime/2010/tables/table-1-incidents-offenses-victims-and-known-offenders-by-bias-motivation-2010.xls)
#
# The file is saved locally as "table1_2010.xls"
# # 2008 Data
# Download Source = [https://ucr.fbi.gov/hate-crime/2008](https://ucr.fbi.gov/hate-crime/2008)
#
# The file is saved locally as "table1_2008.xls"
url = 'table1_2008.xls'
headers = ['Incidents','Offenses','Victims1','Known offenders2']
data_2008 = pd.read_excel(url, skiprows=3, skipfooter=3, parse_cols="A,B,C,D,E",headers = None, names = ["Motivation","Incidents","Offenses","Victims","Known Offenders"])
religion_2008 = data_2008[7:15]
# # 2007 Data
# Download Source = [https://ucr.fbi.gov/hate-crime/2007](https://ucr.fbi.gov/hate-crime/2007)
#
# The file is saved locally as "table1_2007.xls"
url = 'table1_2007.xls'
headers = ['Incidents','Offenses','Victims1','Known offenders2']
data_2007 = pd.read_excel(url, skiprows=3, skipfooter=3, parse_cols="A,B,C,D,E",headers = None, names = ["Motivation","Incidents","Offenses","Victims","Known Offenders"])
religion_2007 = data_2007[7:15]
# # 2006 Data
# Download Source = [https://ucr.fbi.gov/hate-crime/2006](https://ucr.fbi.gov/hate-crime/2006)
#
# The file is saved locally as "table1_2006.xls"
url = 'table1_2006.xls'
headers = ['Incidents','Offenses','Victims1','Known offenders2']
data_2006 = pd.read_excel(url, skiprows=3, skipfooter=3, parse_cols="A,B,C,D,E",headers = None, names = ["Motivation","Incidents","Offenses","Victims","Known Offenders"])
religion_2006 = data_2006[7:15]
# # 2005 Data
# Download Source = [https://ucr.fbi.gov/hate-crime/2005](https://ucr.fbi.gov/hate-crime/2005)
#
# The file is saved locally as "table1_2005.xls"
url = 'table1_2005.xls'
headers = ['Incidents','Offenses','Victims1','Known offenders2']
data_2005 = pd.read_excel(url, skiprows=3, skipfooter=3, parse_cols="A,B,C,D,E",headers = None, names = ["Motivation","Incidents","Offenses","Victims","Known Offenders"])
religion_2005 = data_2005[7:15]
# # Data Web Scraping: 2004
#
# Download Source = [https://www2.fbi.gov/ucr/hc2004/hctable1.htm](https://www2.fbi.gov/ucr/hc2004/hctable1.htm)
#
# The data for 2004 is available as a HTML table on the website.
#
# ##### Following steps are followed in data collection for 2004:
#
# 1. Request the content of the source page using python [request libray](http://docs.python-requests.org/en/master/).
# 2. Format the import webpage contect using python [BeautifulSoup libray's](https://www.crummy.com/software/BeautifulSoup/bs4/doc/) 'html.parser'.
# 3. Since the relevant data is available in html table, loop through the relevant table rows, and extract the data values.
# 4. For each row, write the extracted data values into a local file, named 'source_2004.txt'
target = 'source_2004.txt'
target = open(target, "w")
url = "https://www2.fbi.gov/ucr/hc2004/hctable1.htm"
data_2004 = requests.get(url)
data_2004_soup = BeautifulSoup(data_2004.content, 'html.parser')
data_2004_soup
religion_part = data_2004_soup.find_all('tr')
for row_number in range(9,17):
row = religion_part[row_number]
tmp_string = ''
table_header = row.find('th')
table_values = row.find_all('td')
tmp_string += table_header.text + ' '
for tb in table_values:
tmp_string += tb.text + ' '
tmp_string = tmp_string[:-1].replace('\n','') + '\n'
target.write(tmp_string)
target.close()
# # Data Import 1996 - 2004
#
# Files for yeas 1995-2003, Table 1s are available within pdf reports, and will require a seperate importation technique as compared to the excel files. The relevant data is manually copied from each pdf file seperate, and stored into seperte local files.
#
# ##### Steps for Data Collection using example of 2003:
#
# 1. [The pdf data report](https://ucr.fbi.gov/hate-crime/2003) is available on FBI's website.
# 2. Copy the relevant table rows representing the religion section from Table 1 in the report.
# 3. Paste the copied data into a local file named 'source_2003.txt'
# 4. Save the file.
# 5. Repeat the same steps for year 1996-2002 too.
#
# Once the Data has been saved locally, it is cleaned and converted into .csv format, such that it can be directly imported into DataFrames afterwards.
#
# ##### Steps for cleaing and converting the files to .csv format, and loading them in pandas DataFrames, using year 2003 as example:
# 1. Open the source file e.g source_2003
# 2. Open the target file e.g table1_2003.csv
# 3. Loop through each line in the source file:
# * Remove the endline character i.e '\n'
# * Remove all the commas ',' from the line.
# * Split the line into an array, using empty space as split character.
# * Check if the number of array elements are greater than. If so, array[:-4] are part of the index in the table: join these elements into one element.
# * join the resulting elemets into a string using ',' as join character, and ending the string with newline character '\n'.
# * write the resutling string to target file.
# 4. Close the target and source files.
# 5. Use pandas_readcsv(filename) method to read the .csv file into DataFrames. Set DataFrame headers to ["Motivation","Incidents","Offenses","Victims","Known Offenders"]. Name the returned DataFrame as religion_2003.
# 6. Save religion_2003 to all_years array of DataFrames.
# 5. Loop through the years 1996 to 2003 and repeat the same steps.
# Global Variables
all_years = [] # list of all the DataFrames.
sourcenames = ["source_"+str(year)+".txt" for year in range(1996,2005)] # list of source files names for 1996-2003, to be converted to .csv
targetnames = ["table1_"+str(year)+".csv" for year in range(1996,2005)] # List of name of all .csv files, to be imported in DataFrames
datanames = ["religion_"+str(year) for year in range(1996,2005)] # List of name of all dataframes, to be created e.g religion_1998,religion1999
'''
Steps for cleaing and converting the files to .csv format,
and loading them in pandas DataFrames, using year 2003 as example:
'''
# Loop through the years 1996 to 2003 and repeat the same steps.
for i in range(9):
source = sourcenames[i]
target = targetnames[i]
try:
#Open the source file e.g source_2003
source = open(source,"r",)
except:
print("Could not open the source file")
else:
# Open the target file e.g table1_2003.csv
target = open(target, "w")
lines = source.readlines();
rows = len(lines)
cols = 5
# Loop through each line in the source file:
for line in lines:
# Remove the endline character i.e '\n'
line = line.replace('\n','')
# Remove all the commas ',' from the line.
line = line.replace(",","")
# Split the line into an array, using empty space as split character
line_elements= line.split(' ')
# Check if the number of array elements are greater than. If so, array[:-4] are part of the index in the table: join these elements into one element.
if len(line_elements) > 5:
# join the resulting elemets into a string using ',' as join character, and ending the string with newline character '\n'.
new_line = " ".join(line_elements[:-4]) + ',' + ','.join(line_elements[-4:]) + '\n'
else:
# join the resulting elemets into a string using ',' as join character, and ending the string with newline character '\n'.
new_line = ','.join(line_elements) + '\n'
# write the resutling string to target file.
target.write(new_line)
# Close the target and source files.
source.close()
target.close()
url = targetnames[i]
# Use pandas_readcsv(filename) method to read the .csv file into DataFrames. Set DataFrame headers to ["Motivation","Incidents","Offenses","Victims","Known Offenders"]. Name the returned DataFrame as religion_2003.
exec('%s = pd.read_csv(url, engine = "python", names = ["Motivation","Incidents","Offenses","Victims","Known Offenders"])' % (datanames[i]))
# Save religion_2003 to all_years array of DataFrames.
exec('all_years.append(%s)' % (datanames[i]))
# adding DataFrames for years 2005-2015 excluding 2009 into the all_years list
all_years.extend([religion_2005,religion_2006,religion_2007,religion_2008,religion_2010,religion_2011,religion_2012,religion_2013,religion_2014])
print('Variable dtypes:\n', religion_2000.dtypes, sep='')
religion_1996
rel = religion_1996['Motivation']
rel
# # DataFrame Description for a particular year
# #### Headers:
# 1. Motivation: The Motivation behind the hate crime. Anti-Islamic means hate crimes motivated by sentiment againsts Islam/Muslims(followers of Islam)
# 2. Incidents: Total Number of reported incidents of hate crimes, for a particular motivation
# 3. Offenses: Total Number of reported offenses of hate crimes, for a particular motivation
# 4. Victims : Total Number of reported victims of hate crimes, for a particular motivation
# 5. Known Offender: Total Number of reported known offenses of hate crimes, for a particular motivation
#
# #### Indexes : Motivation (Following motivations have been recorded)
# 1. Religion: (Total Number for All Religions)
# 2. Anti-Jewish
# 3. Anti-Catholic
# 4. Anti-Protestant
# 5. Anti-Islamic
# 6. Anti-Other Religious Group (Total Number for Other religius groups)
# 7. Anti-Multi-Religious Group (Total Number for crimes which targetted multiple religions together)
# 8. Anti-Atheism/Agnosticism/etc.
#
# #### Example for Year 2003 is shown below
religion_2003
# # Combining DataFrames for all years into one DataFrame
#
# all_years is the list of the DataFrames for all years.
# We want to combine the data for all the years into one DataFrame so that it can be used for analysis.
#
# ### Folloing Steps Are taken for Combining the Data:
#
# 1. Combine 8 Motivations with the different data values' headers:
# * Use the 8 motivations : ['All Religion','Anti-Jewish','Anti-Catholic','Anti-Protestants','Anti-Islamic','Anti-Other Religion','Anti-Multiple Religion,Group','Anti-Atheism/Agnosticism/etc.']
# * Use the 4 Data Values headers = ['Incidents','Offenses', 'Victims','Known Offenders']
# * Create 32 headers such that for each motivation, there are 4 different headers for the different data values.
# * E.g for 'Anti-Jewish' motivation, the resulting headers will be Anti-Jewish: Incidents,Anti-Jewish: Offenses,Anti-Jewish: Victims', and Anti-Jewish: Known Offenders.
# * all_years_headers is the list of all the generated headers.
#
# 2. Generate a list called all_years_keys, which will correspond to the indices of the new DataFrame.
# all_years_keys = ['1996',
# '1997',
# '1998',
# '1999',
# '2000',
# '2001',
# '2002',
# '2003',
# '2004',
# '2005',
# '2006',
# '2007',
# '2008',
# '2010',
# '2011',
# '2012',
# '2013',
# '2014']
#
# 3. Create the combined DataFrame:
# * Loop through all_year - the list of the DataFrames representing each year.
# * Within each DataFrameLoop through all rows within each DataFrame:
# * Within each row, loop through all column values
# * add the column values into a temporary list
# * Add the temporary list cosisting of all the data values of the data frame into all_years_list. all_years_list is the double-nested list of data values for all years.
#
# 4. Create the DataFrame using all_years_list as data, all_years_keys as indices, all_years_headers as headers. Name this DataFrame hc, representing hate crimes
#Variables and Description
# List of Indices (Motivation) in a DataFrame for a particular yaer
header_rows = ['All Religion','Anti-Jewish','Anti-Catholic','Anti-Protestants','Anti-Islamic','Anti-Other Religion','Anti-Multiple Religion,Group','Anti-Atheism/Agnosticism/etc.']
# List of headers in a DataFrame for particular yaer
columns = ['Incidents','Offenses', 'Victims','Known Offenders']
# List of headers for the new DataFrame
all_years_headers = []
#List of list of all values in the DataFrames for all years
all_years_list=[]
# List of the new indices, representing all reported years, for the new DataFrams.
all_years_keys = []
# +
'''
Folloing Steps Are taken for Combining the Data:
'''
'''
Combine 8 Motivations with the different data values' headers:
* Use the 8 motivations : ['All Religion','Anti-Jewish','Anti-Catholic','Anti-Protestants','Anti-Islamic','
Anti-Other Religion','Anti-Multiple Religion,Group','Anti-Atheism/Agnosticism/etc.']
* Use the 4 Data Values headers = ['Incidents','Offenses', 'Victims','Known Offenders']
* Create 32 headers such that for each motivation, there are 4 different headers for the different data values.
* E.g for 'Anti-Jewish' motivation, the resulting headers will be Anti-Jewish: Incidents,Anti-Jewish: Offenses,
Anti-Jewish: Victims', and Anti-Jewish: Known Offenders.
* all_years_headers is the list of all the generated headers.
'''
for row in header_rows:
for col in columns:
header_val = row + ': ' + str(col)
all_years_headers.append(header_val)
'''
Generate a list called all_years_keys, which will correspond to the indices of the new DataFrame.
'''
for i in list(range(1996,2009)) + list(range(2010, 2015)):
all_years_keys.append(str(i))
count = 0
'''
Create the combined DataFrame:
'''
# Loop through all_year - the list of the DataFrames representing each year *
for single_year in all_years:
tmp_list =[]
# Within each DataFrameLoop through all rows :
for row in range(8):
current_row = single_year.iloc[row]
# Within each row, loop through all column values
for col in columns:
# add the column values into a temporary list
tmp_list.append(current_row[col])
# Add the temporary list cosisting of all the data values of the data frame into all_years_list.
all_years_list.append(tmp_list)
count+=1
'''
Create the DataFrame using all_years_list as data, all_years_keys as indices, all_years_headers as headers.
Name this DataFrame hc, representing hate crimes
'''
hc = pd.DataFrame(all_years_list, columns= all_years_headers, index = all_years_keys)
# -
hc
# ### Q: 1. How have the hate crimes against Muslims changed in terms of number of incidents per year?
anti_islam = hc['Anti-Islamic: Incidents']
anti_islam.plot(kind='line',
grid = True,
title = 'Anti-Islam Hate Crimes',
sharey = True,
sharex = True,
use_index = True,
legend = True,
fontsize = 10
)
# ### Answer:
# The number of hate crime incidents against Muslims have fluctuated a lot over the years. The most striking number of incidents took place in 2011 as shown by the graph above. Before 2011, the maximum number of incidents were 32, and the minimum were 211. In 2011, the number of incidents were 481. After 2011, the max number of incidents were 156, and the minimum were 105. In recent years, the hate crimes against Muslims have started to rise again.From 1995 to 2014, the number of incidents changed as shown below:
print(anti_islam)
# ### Q : Did September-11 Terrorist Attack had an impact on the hate crimes against Muslims? If so, how much impact did September-11 Terrorist Attack had?
# +
anti_islam_2011 = anti_islam[5]
anti_islam_2010 = anti_islam[4]
anti_islam_2012 = anti_islam[6]
percentage_change_2011 = (((anti_islam_2011 - anti_islam_2010)/anti_islam_2010)*100)
percentage_change_2012 = (((anti_islam_2012 - anti_islam_2011)/anti_islam_2011)*100)
print("Hate Crimes against Muslims growth in 2011 from 2010: ", percentage_change_2011, '%')
print("Hate Crimes against Muslims growth in 2010 from 2011: ", percentage_change_2012, '%')
anti_islam_before_2011 = anti_islam[:5].mean()
anti_islam_after_2011 = anti_islam[6:].mean()
print('Average hate crimes against Muslims before 2011: ', anti_islam_before_2011)
print('Average hate crimes against Muslims before 2011: ', anti_islam_after_2011)
avg = (((anti_islam_after_2011 - anti_islam_before_2011)/anti_islam_before_2011)*100)
print('Percentage increased in the average number of hate crimes against Muslims after 2011: ', avg)
# -
# #### Answer:
# September-11 Terrorist Attack had a huge impact on the number of hate-crimes against Muslims. The incident took place in 2011, where there were 481 hate crimes against Muslims, as opposed to 28 in 2010. The number of hate crimes against Muslims increased by more than 16 times (1672%) in 2011 as compared to 2010. In the following year (2012), the number of hate crimes against Muslims decreased by almost 68%. The average number of hate crimes against Muslims were 27 before 2011, and after 2011, they have increased to an average of 142. The average number of hate crimes against Muslims increased by more than 4 times (421%).
# ### Q: How have the hate crimes against All religion changed in terms of number of incidents per year?
anti_religion = hc['All Religion: Incidents']
anti_religion.plot(kind='line',
title = 'Hate Crimes Against All Religion',
sharey = True,
sharex = True,
use_index = True,
legend = True)
anti_religion_2011 = anti_religion[5]
anti_religion_2010 = anti_religion[4]
anti_religion_2012 = anti_religion[6]
avg_before_2011 = anti_religion[:5].mean()
avg_after_2011 = anti_religion[6:].mean()
avg_after_2008 = anti_religion[13:].mean()
print('Average Number of Crimes before 2011 : ', avg_before_2011)
print('Avearage Number of Crimes after 2011 : ', avg_after_2011)
print('Avearage Number of Crimes after 2008 : ', avg_after_2008)
print('Hate Crimes in 2011 : ', anti_religion_2011)
# #### Answer:
# As shown in the graph above, the number of hate_crimes against all religion fluctuated by going up and down in between 1996 and 2008, with a very high peak in 2011. Since 2008, the number has seen a consistent and steady decrease. It is the same year, <NAME> got elected as the President of the United States. The average number of crimes before 2011 were 1412, and after 2011 they were 1288. In 2011, there were 1828, and most of the stark increase can be contributed towards the stark increase in the hate crimes against Muslims.
# ### Q: What percentage of hate crimes motivated by religion identity target Muslims every year?
#
anti_muslim_percentage= (hc['Anti-Islamic: Incidents']/hc['All Religion: Incidents'])*100
anti_muslim_percentage.plot(kind = 'line',
title = 'Percentage of Hate Crimes Against Muslims Among All Religion',
sharey = True,
sharex = True,
use_index = True)
# #### Answer
# The ratio being discussed is shown above. Hate crimes targetting Muslims as a ratio of the hate-crimes motivated by religion has increased a lot in 2011 because of September-11 terrorist attack.Before, 2011, it was below 3% consistently, and in 2011 it went beyond 25%. After 2011, it never went down to its pre-2011 number. This also shows that the September-11 incident has increased the general sentiment against Muslims, and even after over a decade, the effect of September-11 on hate crimes against Muslims is clearly evident. Moreover, we can also see that the ratio has been rising in recent years showing that even though the number of hate-crimes against religions as a whole are decreasing, among those numbers, the ratio of attacks on Muslims in increaseing.
# ### Q: On average what percentages of attacks motivated by religion targetted Muslims, before and after the September-11 Terrorist Attack?
avg_before_2011 = something[:5].mean()
#not including 2011 in either average before or after 2011
avg_after_2011 = something[6:].mean()
perc_increase = (((avg_after_2011 - avg_before_2011)/avg_before_2011)*100)
print(avg_before_2011, avg_after_2011, perc_increase)
growth_list = []
# #### Answer
# Mean of Percentage of Attacks on Muslims among All Hate Crimes motivated by religion, before 2011 : 1.9259431800868643 %
#
# Mean of Percentage of Attacks on Muslims among All Hate Crimes motivated by religion, after 2011 : 11.20777275833266 %
#
# Percentage Chaneg among the two means : 481.93683355846264 %
# # Answer to Main Research Question:
# #### Q: Did September-11 terrorist attack have an impact on hate crimes against Muslims? How much and what impact did the incident have?
# # Answer:
# The number of hate crime incidents against Muslims have fluctuated a lot over the years. The most striking number of incidents took place in 2011 as shown by the graph above. Before 2011, the maximum number of incidents were 32, and the minimum were 211. In 2011, the number of incidents were 481. After 2011, the max number of incidents were 156, and the minimum were 105. In recent years, the hate crimes against Muslims have started to rise again.From 1995 to 2014, the number of incidents changed as shown above.
# September-11 Terrorist Attack had a huge impact on the number of hate-crimes against Muslims. The incident took place in 2011, where there were 481 hate crimes against Muslims, as opposed to 28 in 2010. The number of hate crimes against Muslims increased by more than 16 times (1672%) in 2011 as compared to 2010. In the following year (2012), the number of hate crimes against Muslims decreased by almost 68%. The average number of hate crimes against Muslims were 27 before 2011, and after 2011, they have increased to an average of 142. The average number of hate crimes against Muslims increased by more than 4 times (421%).
# As shown in the 3rd graph above, the number of hate_crimes against all religion fluctuated by going up and down in between 1996 and 2008, with a very high peak in 2011. Since 2008, the number has seen a consistent and steady decrease. It is the same year, <NAME> got elected as the President of the United States. The average number of crimes before 2011 were 1412, and after 2011 they were 1288. In 2011, there were 1828, and most of the stark increase can be contributed towards the stark increase in the hate crimes against Muslims.
# Hate crimes targetting Muslims as a ratio of the hate-crimes motivated by religion increased a lot in 2011 because of September-11 terrorist attack.Before, 2011, it was below 3% consistently, and in 2011 it went beyond 25%. After 2011, it never went down to its pre-2011 number. This also shows that the September-11 incident has increased the general sentiment against Muslims, and even after over a decade, the effect of September-11 on hate crimes against Muslims is clearly evident. Moreover, we can also see that the ratio has been rising in recent years showing that even though the number of hate-crimes against religions as a whole are decreasing, among those numbers, the ratio of attacks on Muslims in increaseing.
# The Mean of Percentage of Attacks on Muslims among All Hate Crimes motivated by religion, before 2011 is 1.9259431800868643 %. The Mean of Percentage of Attacks on Muslims among All Hate Crimes motivated by religion, after 2011 is 11.20777275833266 %. Percentage Chaneg among the two means is 481.93683355846264 %.
#
# # Conclusion:
# September-11 Terrorist attack had a pivotal impact on the number of hate crimes against Muslims, and the affect of the in
#
| UG_S17/Ahmad-HateCrimesAgainstMuslims.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tass
#
# +
import requests
import numpy as np
import datetime
import re
from bs4 import BeautifulSoup
from fake_useragent import UserAgent
import time
import datetime
import pandas as pd
import pickle
import time
from tqdm import tqdm
from joblib import Parallel, delayed
from tqdm import tqdm
# +
from multiprocessing.pool import ThreadPool
def Separator(vect, part):
n = len(vect)
vec_parts = [round(n/part)*i for i in range(part)]
vec_parts.append(n)
out = [vect[vec_parts[i]:vec_parts[i+1]] for i in range(part)]
return(out)
def Map(vect, parser_function):
out = [parser_function(item) for item in vect]
return(out)
def Reduce(l):
ll = [ ]
for item in l:
ll.extend(item)
return(ll)
def MRDownloader(what, parts, parser_function):
separatorlist = Separator(what, parts)
def Mp(what):
return Map(what, parser_function=parser_function)
pool = ThreadPool(parts)
l = pool.map(Mp, separatorlist)
itog = Reduce(l)
return(itog)
# -
# # 1. Download hrefs
def tass_lenta(before,limit=200):
mainpage = 'http://tass.ru/api/news/lenta?limit='+str(limit)+'&before='+str(before)
response = requests.get(mainpage)
dic = response.json()
cur_news = [ ]
df = dic['articles']
for item in df:
try:
new = { }
new['title'] = item['title']
new['category'] = item['section']['title']
new['href'] = item['url']
new['date'] = datetime.datetime.fromtimestamp(int(item['time'])).strftime('%Y-%m-%d %H:%M:%S')
new['uci_time'] = int(item['time'])
cur_news.append(new)
except:
print('Error! \n',item, '\n')
return cur_news
'1514764790' # 31 декабря 2017 года 23:59:59
'1483228799' # 2016
'1451606399' # 2015
'1420070399' # 2014
'1388534399' # 2013
'1356998399' # 2012
'1325375999' # 2011
'1293839999' # 2010
'1262303999' # 2009
'1230767999' # 2008
# +
# tass_news = [ ]
# before = 999999999999
for i in tqdm(range(100000)):
current_news = tass_lenta(before)
before = current_news[-1]['uci_time']
tass_news.extend(current_news)
if before < 1514674664:
break
# -
before
len(tass_news)
tass_news2 = [item for item in tass_news if item['date'].split('-')[0] == '2020']
len(tass_news2)
tass_news3 = [item for item in tass_news2 if item['date'].split('-')[1] in ['06', '07', '08', '09', '10', '11']]
len(tass_news3)
tass_news = tass_news3
# +
print('В списке сейчас: ', len(tass_news))
df = pd.DataFrame(tass_news)
print(df.shape)
print('Без дубликатов: ', len(set(df.href)))
print(df.drop_duplicates().shape)
with open('news_data/tass_titles_new.pickle', 'wb') as f:
pickle.dump(tass_news, f)
df.tail()
# -
# ```
# Что ещё можно достать нахаляву из df
# {'audio': False, 'color': '2', 'flash': False, 'id': '4070830',
# 'is_breaking_news': False, 'is_online': False, 'live_text': False,
# 'marked': False, 'photos': False, 'search_queries': None,
# 'section': {'id': '25', 'title': 'Экономика и бизнес', 'url': '/ekonomika'},
# 'show_at_common_feed': True, 'show_at_section_feed': True, 'slideshow': False
# 'time': '1488624568', 'title': 'Глава ВТБ: курс доллара к концу года может достигнуть 61-62 рублей',
# 'topics': None, 'url': '/ekonomika/4070830', 'video': False},
# ```
# # 2. Download news
# +
with open('news_data/tass_titles_new.pickle', 'rb') as f:
tass_titles = pickle.load(f)
len(tass_titles)
# +
def page_content(url):
response = requests.get(url)
html = response.content
soup = BeautifulSoup(html,"lxml")
vvv = soup.findAll("div", { "class" : "news" })[0]
snippet = vvv.find_all("div", {"class": "news-header__lead"})[0].text
text = vvv.find_all("div", {"class": "text-content"})[0].text
return text, snippet
def get_tass_news(item_from_vect):
url = "http://tass.ru" + item_from_vect['href']
try:
text, snippet = page_content(url)
item_from_vect['text'] = text.strip()
item_from_vect['snippet'] = snippet.strip()
return item_from_vect
except:
print(url)
return{ }
get_tass_news(tass_titles[0])
# -
# itog_titles = [item for item in tass_titles if item['date'][:4] == "2020"]
# len(itog_titles)
itog_titles = tass_titles
def Separator(vect, part):
n = len(vect)
vec_parts = [round(n/part)*i for i in range(part)]
vec_parts.append(n)
out = [vect[vec_parts[i]:vec_parts[i+1]] for i in range(part)]
return(out)
# +
i = 0
x_batch = Separator(itog_titles, 500)
# -
result = [ ]
for batch in tqdm(x_batch[(198+186):]):
n_jobs = -1
result_cur = Parallel(n_jobs=n_jobs)(delayed(get_tass_news)(
text) for text in batch)
print('Скачал батч номер ' + str(i))
i += 1
result.extend(result_cur)
# +
# # %%time
# itog_news = MRDownloader(itog_titles, 10, parser_function=get_tass_news)
# len(itog_news)
# -
len(result)
# хули так мало, по ходу парсер не докачивает какие-то случаи
itog_news = [itog for itog in result if len(itog.keys()) != 0 ]
len(itog_news)
pd.DataFrame(itog_news).head()
with open('news_data/tass_news_new_part3.pickle', 'wb') as f:
pickle.dump(itog_news, f)
#
#
| 01.news_parser/4.tass.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_tensorflow_p36
# language: python
# name: conda_tensorflow_p36
# ---
# !pip install imgaug
# !pip install keras==2.1.3
# +
import argparse
import logging
import os
import time
import keras
import numpy as np
import cv2
import matplotlib.pyplot as plt
from tqdm.notebook import tqdm
from semantic_segmentation.data_generators import BatchGenerator
from semantic_segmentation.model_runner import ModelRunner
from semantic_segmentation.net import NetManager, NetConfig
# -
net_manager = NetManager("./logs/2020-11-13_11.28.53")
net_config = net_manager.load_model()
net_config = NetConfig.from_others(net_config)
model = net_manager.get_keras_model()
input_image_shape = (1, net_config.get_max_side(), net_config.get_max_side(), 1)
model.predict(np.zeros(input_image_shape))
t = time.time()
model.predict(np.zeros(input_image_shape))
t = time.time() - t
# +
def evaluate_write_to_disk(model):
base_dir = "/home/ec2-user/SageMaker/benchmarks/dataset/ynet/"
all_bbx = []
for i in tqdm(range(90000, 100000)):
test_image_path = "X/train_{}.png".format(i)
img = cv2.imread(base_dir + test_image_path)
if len(boxes) == 0:
print("OOPS")
all_bbx.append(boxes)
with open("yolo-prediction.pickle", "wb") as f:
pickle.dump(all_bbx, f)
print("done")
evaluate_write_to_disk(model)
# -
input_image_shape
# +
base_dir = "/home/ec2-user/SageMaker/benchmarks/dataset/ynet/"
i = 90000
test_image_path = "X/train_{}.png".format(i)
img = cv2.imread(base_dir + test_image_path)
gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
result = model.predict(gray.reshape(1, 400, 400, 1)).squeeze()
# -
plt.imshow(result, cmap='gray')
plt.imshow(gray)
import tensorflow as tf
tf.__version__
import h5py
h5py.__version__
| Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
## Data Preprocessing
import pandas as pd
import numpy as np
## Data Visualization
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
## Modeling
from sklearn.tree import DecisionTreeRegressor
from sklearn import ensemble
from sklearn.model_selection import train_test_split, \
cross_val_score, \
GridSearchCV
## Find out execution time
from datetime import datetime
## Graph visualization
from sklearn.tree import export_graphviz
from sklearn.externals.six import StringIO
from IPython.display import Image
import pydotplus
## Suppress warnings
import warnings
warnings.filterwarnings("ignore")
# -
df = pd.read_csv('_inputs/A_exam.csv')
df.shape
df.head()
df = pd.get_dummies(df, columns=['D', 'E'], drop_first = True)
X = df.drop(columns=['Response'])
Y = df[['Response']]
X.head()
Y.head()
X_train, X_test, y_train, y_test = train_test_split(X,Y,test_size = 0.3)
#
# # Decision Tree
#
np.linspace(1, 20, 20, endpoint=True)
# +
dt = DecisionTreeRegressor(random_state=42)
startTime = datetime.now()
param_grid = {
'max_features': list(range(1,X_train.shape[1])),
'max_depth' : np.linspace(1, 20, 20, endpoint=True),
# 'min_samples_split': np.linspace(0.1, 1.0, 5, endpoint=True),
# 'min_samples_leaf': np.linspace(0.1, 0.5, 5, endpoint=True)
}
CV_dt = GridSearchCV(estimator = dt, param_grid = param_grid, cv = 5)
CV_dt.fit(X_train, y_train)
print(CV_dt.best_params_)
print(datetime.now() - startTime)
# +
dt = DecisionTreeRegressor(
max_features = 7,
max_depth = 2,
# min_samples_leaf = 0.1,
# min_samples_split = 0.1
)
# Train Decision Tree Classifer
dt = dt.fit(X_train,y_train)
#Predict the response for test dataset
pred = dt.predict(X_test)
dt_score = dt.score(X_test,y_test)
print('Score of decision tree classifier: ', dt_score)
# +
dot_data = StringIO()
export_graphviz(dt,
out_file = dot_data,
filled = True,
rounded = True,
special_characters = True,
feature_names = X_train.columns.values
)
graph = pydotplus.graph_from_dot_data(dot_data.getvalue())
Image(graph.create_png())
# -
print(dict(zip(X.columns, dt.feature_importances_)))
#
# # AdaBoost
#
# +
score = np.array([])
startTime = datetime.now()
for i in [100, 150, 200, 250, 300]:
for j in [0.001, 0.01, 0.1]:
bc = ensemble.AdaBoostRegressor(DecisionTreeRegressor(), n_estimators = i, learning_rate=j)
mod_cv = cross_val_score(bc, X_train, y_train, cv = 5)
score = np.append(score, np.mean(mod_cv))
print("n_estimators->", i, "learning_rate->", j, "Mean->", np.mean(mod_cv))
# +
Ada = ensemble.AdaBoostRegressor(DecisionTreeRegressor(), n_estimators = 100, learning_rate=0.001)
Ada = Ada.fit(X_train,y_train)
pred = Ada.predict(X_test)
Ada_score = Ada.score(X_test,y_test)
print('Score of AdaBoost classifier: ', Ada_score)
# +
feature_importances = np.mean([
tree.feature_importances_ for tree in Ada.estimators_
], axis=0)
print(dict(zip(X.columns, feature_importances)))
# -
#
# # Best Model Selection
#
print('Score of decision tree classifier: ', dt_score)
print('Score of AdaBoost classifier: ', Ada_score)
| Exams/Part 2/Midterm_1/Final_Tree_Regression_LKH.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + dc={"key": "4"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 1. Introduction
# <p><img src="https://assets.datacamp.com/production/project_1197/img/google_play_store.png" alt="Google Play logo"></p>
# <p>Mobile apps are everywhere. They are easy to create and can be very lucrative from the business standpoint. Specifically, Android is expanding as an operating system and has captured more than 74% of the total market<sup><a href="https://www.statista.com/statistics/272698/global-market-share-held-by-mobile-operating-systems-since-2009">[1]</a></sup>. </p>
# <p>The Google Play Store apps data has enormous potential to facilitate data-driven decisions and insights for businesses. In this notebook, we will analyze the Android app market by comparing ~10k apps in Google Play across different categories. We will also use the user reviews to draw a qualitative comparision between the apps.</p>
# <p>The dataset you will use here was scraped from Google Play Store in September 2018 and was published on <a href="https://www.kaggle.com/lava18/google-play-store-apps">Kaggle</a>. Here are the details: <br>
# <br></p>
# <div style="background-color: #efebe4; color: #05192d; text-align:left; vertical-align: middle; padding: 15px 25px 15px 25px; line-height: 1.6;">
# <div style="font-size:20px"><b>datasets/apps.csv</b></div>
# This file contains all the details of the apps on Google Play. There are 9 features that describe a given app.
# <ul>
# <li><b>App:</b> Name of the app</li>
# <li><b>Category:</b> Category of the app. Some examples are: ART_AND_DESIGN, FINANCE, COMICS, BEAUTY etc.</li>
# <li><b>Rating:</b> The current average rating (out of 5) of the app on Google Play</li>
# <li><b>Reviews:</b> Number of user reviews given on the app</li>
# <li><b>Size:</b> Size of the app in MB (megabytes)</li>
# <li><b>Installs:</b> Number of times the app was downloaded from Google Play</li>
# <li><b>Type:</b> Whether the app is paid or free</li>
# <li><b>Price:</b> Price of the app in US$</li>
# <li><b>Last Updated:</b> Date on which the app was last updated on Google Play </li>
#
# </ul>
# </div>
# <div style="background-color: #efebe4; color: #05192d; text-align:left; vertical-align: middle; padding: 15px 25px 15px 25px; line-height: 1.6;">
# <div style="font-size:20px"><b>datasets/user_reviews.csv</b></div>
# This file contains a random sample of 100 <i>[most helpful first](https://www.androidpolice.com/2019/01/21/google-play-stores-redesigned-ratings-and-reviews-section-lets-you-easily-filter-by-star-rating/)</i> user reviews for each app. The text in each review has been pre-processed and passed through a sentiment analyzer.
# <ul>
# <li><b>App:</b> Name of the app on which the user review was provided. Matches the `App` column of the `apps.csv` file</li>
# <li><b>Review:</b> The pre-processed user review text</li>
# <li><b>Sentiment Category:</b> Sentiment category of the user review - Positive, Negative or Neutral</li>
# <li><b>Sentiment Score:</b> Sentiment score of the user review. It lies between [-1,1]. A higher score denotes a more positive sentiment.</li>
#
# </ul>
# </div>
# <p>From here on, it will be your task to explore and manipulate the data until you are able to answer the three questions described in the instructions panel.<br></p>
# + [markdown] dc={"key": "4"}
# ## 1. Read the `apps.csv` file and clean the `Installs` column to convert it into integer data type
# + dc={"key": "4"}
import pandas as pd
# + dc={"key": "4"}
apps = pd.read_csv('apps2.csv')
apps.info()
apps.head()
# + dc={"key": "4"}
chars_to_remove = [',', '+']
for char in chars_to_remove:
apps['Installs'] = apps['Installs'].apply(lambda x: x.replace(char, ''))
apps.info()
apps.head()
# + dc={"key": "4"}
apps['Installs'] = apps['Installs'].astype(int)
apps.info()
# + [markdown] dc={"key": "4"}
# ## 2. Find the number of apps in each category, the average price, and the average rating.
# + dc={"key": "4"}
app_category_info = apps.groupby('Category').agg({'App': 'count', 'Price': 'mean', 'Rating': 'mean'})
app_category_info
# + dc={"key": "4"}
app_category_info = app_category_info.rename(columns={'App': 'Number of apps', 'Price': 'Average price', 'Rating': 'Average rating'})
app_category_info
# + [markdown] dc={"key": "4"}
# ## 3. Find the top 10 free `FINANCE` apps having the highest average sentiment score.
# + dc={"key": "4"}
reviews = pd.read_csv('user_reviews2.csv')
reviews.info()
reviews.head()
# + dc={"key": "4"}
finance_apps = apps[apps['Category'] == 'FINANCE']
finance_apps.head()
# + dc={"key": "4"}
free_finance_apps = finance_apps[finance_apps['Type'] == 'Free']
free_finance_apps.head()
# + dc={"key": "4"}
merged_df = pd.merge(free_finance_apps, reviews, on='App')
merged_df.head()
# + dc={"key": "4"}
app_sentiment_score = merged_df.groupby('App').agg({'Sentiment Score': 'mean'})
app_sentiment_score.head()
# + dc={"key": "4"}
user_feedback = app_sentiment_score.sort_values(by='Sentiment Score', ascending=False)
user_feedback.head()
# + dc={"key": "4"}
top_10_user_feedback = user_feedback[:10]
top_10_user_feedback
| the_android_app_market_on_google_play/the_android_app_market_on_google_play_unguided.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Opengameart.org CC0 Image Download
#
# This notebook downloads and cleans data from opengameart.org.
# #### Step 1: Get Collection Links
# #### Step 2: Get File Links
# #### Step 3: Download Files
# #### Step 4: Copy and Rename Files
import requests, bs4, os, json, zipfile, io, cv2, numpy as np
from statistics import mean
from collections import Counter
from datetime import datetime
from distutils.dir_util import copy_tree
def est_time_remaining(i, total, times, last_time, a='Page'):
"""
Calculates the time remaining on a function given time and iteration count.
Parameters:
i: iteration number
total: total iterations
times: list of run times
last_time: last datetime
a: things iterated over
Returns:
times, now
"""
now = datetime.now()
times.append((now-last_time).microseconds/1000000)
avg_time = round(mean(times), 2)
ETA = round(avg_time*(total-i))
print("{}/{} {}(s) Scraped. Avg. Time: {}. Est. Remaining: {}".format(i, total, a, avg_time, ETA), end='\r')
return times, now
# ## Step 1: Get Collection Links
def get_links(dim, pages=2):
"""
Gets every collection link from opengameart.org
parameters:
dim: dimension integer
pages: number of pages o scrape
Returns:
list of links
"""
times = []; last_time = datetime.now()
if 'data_1.json' not in os.listdir():
if dim == 2:
base = "https://opengameart.org/art-search-advanced?keys=&title=&field_art_tags_tid_op=or&field_art_tags_tid=&name=&field_art_type_tid%5B%5D=9&field_art_licenses_tid%5B%5D=4&sort_by=count&sort_order=DESC&items_per_page=144&Collection=&page={}"
if dim == 3:
base = "https://opengameart.org/art-search-advanced?keys=&title=&field_art_tags_tid_op=or&field_art_tags_tid=&name=&field_art_type_tid%5B%5D=10&field_art_licenses_tid%5B%5D=4&sort_by=count&sort_order=DESC&items_per_page=144&Collection=&page={}"
links_all = []
for page in list(range(pages)):
r = requests.get(base.format(str(page)))
if r.status_code==200:
soup = bs4.BeautifulSoup(r.content, 'lxml')
links = []
for s in soup.find_all('div', {'class':'field-item even'}):
try:
href = s.find('a')['href']
if '/content' in href:
links.append(href)
except:
pass
links_all+=links; links_all=list(set(links_all))
times, last_time = est_time_remaining(page+1, pages, times, last_time)
return links_all
links_all = get_links(2)
len(links_all)
# ## Step 2: Get File Links
def get_file_links(links_all):
"""For each collection, gets the links of the files to download
Parameters:
links_all: list of collection links
Returns:
list of file links
"""
files = []; i = 1
total = len(links_all); times = []; last_time=datetime.now()
for link in links_all:
base='https://opengameart.org'
try:
r=requests.get(base+link)
if r.status_code == 200:
soup = bs4.BeautifulSoup(r.content, 'lxml')
try:
file_path = soup.find('span', {'class':'file'}).find('a')['href']
files.append([link, file_path])
except:
pass
try:
for div in soup.find('div', {'class':'group-right right-column'}).find('div', {'class':'field-items'}).find_all('div'):
files.append([link,div.find('a')['href']])
except:
pass
except:
pass
times, last_time = est_time_remaining(i, total, times, last_time); i+=1
return files
file_links = get_file_links(links_all[:3])
len(file_links)
# ## Step 3: Download Files
def download_files(files):
"""Downloads every file to the local directory (requires about 10gb storage for all collections)
Parameters:
files: list of files to download
Returns:
None
"""
try:
os.makedirs('SpriteFiles')
except:
return "Directory Exists"
len(links_all); times = []; last_time=datetime.now()
i=1; l=len(files)
for file_pair in files:
if 'zip' in file_pair[-1]:
try:
os.makedirs('SpriteFiles/'+file_pair[0].split('/')[-1])
except:
pass
try:
r = requests.get(file_pair[-1])
z = zipfile.ZipFile(io.BytesIO(r.content))
z.extractall('SpriteFiles/'+file_pair[0].split('/')[-1])
except:
print(file_pair[-1])
if 'png' in file_pair[-1]:
try:
os.makedirs('SpriteFiles/'+file_pair[0].split('/')[-1])
except:
pass
try:
r = requests.get(file_pair[-1], stream=True)
if r.status_code == 200:
with open('SpriteFiles/'+file_pair[0].split('/')[-1]+'/test.png', 'wb') as f:
for chunk in r:
f.write(chunk)
except:
pass
times, last_time = est_time_remaining(i, l, times, last_time, a='Files'); i+=1
download_files(file_links[:2])
# ## Step 4: Copy and Rename Files to be Sorted
# note your mappings may differ due to newly created collections.
# use the mapping file for categorizing the sprites (if desired)
def copy_rename():
"""This function takes the downloaded raw sprite files and moves them into another folder.
Then, it renames the files based on an index (lots of duplicates like sword).
Only use this function before sorting into categories for the first time.
Parameters:
None
Returns:
None
"""
try:
os.makedirs('SpriteFiles2D')
fromDirectory = "SpriteFiles2DBase"
toDirectory = "SpriteFiles2D"
copy_tree(fromDirectory, toDirectory)
ind = 0; key = {}
for root_folder in os.listdir('SpriteFiles2D'):
os.makedirs('RenamedSprites\\'+root_folder)
for r, d, filenames in os.walk('SpriteFiles2D\\'+root_folder):
for filename in filenames:
old_path = os.path.join(r,filename)
if '.png' in filename:
new_path = "RenamedSprites\\"+root_folder+"\\"+f"{ind:05d}"+".png"
os.rename(old_path, new_path)
key[old_path] = f"{ind:05d}"
ind+=1
else:
os.remove(old_path)
inv_key = {v: k for k, v in key.items()}
mappings = {'filepath_id':key, 'id_filepath':inv_key}
with open('mappings.json', 'w') as outfile:
json.dump(mappings, outfile)
except:
pass
| Code/opengameartDownloadCC0.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "-"}
# # OuterSPACE spMspM accelerator
#
# First, include some libraries
# +
# Run boilerplate code to set up environment
# %run ../prelude.py
# -
# ## Control panel to control display behavior manually
# +
# Uncomment the appropriate lines to change the default behavior
# FTD.showAnimations(True) # Turn on animations
# FTD.showAnimations(False) # Turn off animations
# FTD.setStyle(matrix=True) # Show tensor as a matrix
# FTD.setStyle(matrix=False) # Show tensor as a fiber tree
# -
# ## Read matrices
#
# +
a = Tensor.fromYAMLfile(os.path.join(data_dir, "sparse-matrix-a.yaml"))
b = Tensor.fromYAMLfile(os.path.join(data_dir, "sparse-matrix-b.yaml"))
# Transpose the "a" matrix as desired by the outer product traveral order
at = Tensor.fromFiber(["K", "M"], a.getRoot().swapRanks())
print("Input A")
displayTensor(a.setColor("blue"))
print("Input A - transposed")
displayTensor(at.setColor("blue"))
print("Input B")
displayTensor(b.setColor("green"))
z_verify = Tensor.fromYAMLfile(os.path.join(data_dir, "sparse-matrix-a_times_b.yaml"))
print("Result Z (precomputed)")
displayTensor(z_verify)
# -
# ## Outer Product - Naive
#
# +
z = Tensor(rank_ids=["M", "N"])
canvas = createCanvas(at, b, z)
at_k = at.getRoot()
b_k = b.getRoot()
z_m = z.getRoot()
for k, (at_m, b_n) in at_k & b_k:
#print(f" {k}, ({a_m}, {b_n})")
for m, (z_n_ref, at_val) in z_m << at_m:
#print(f" {m}, ({z_n_ref}, {at_val})")
for n, (z_ref, b_val) in z_n_ref << b_n:
#print(f" {n}, ({z_ref}, {b_val})")
z_ref += at_val * b_val
canvas.addFrame((k, m), (k, n), (m, n))
print("Result Z (computed)")
displayTensor(z)
displayCanvas(canvas)
# -
# ## Check result
z_verify == z
# # OuterSPACE - Step 1
# +
t = Tensor(rank_ids=["M", "K", "N"])
canvas =createCanvas(at, b, t)
at_k = at.getRoot()
b_k = b.getRoot()
t_m = t.getRoot()
for k, (at_m, b_n) in at_k & b_k: # Parallelize here
#print(f" {k}, ({a_m}, {b_n})")
for m, at_val in at_m:
#print(f" {m}, {at_val})")
temp_n = t_m.getPayloadRef(m, k)
for n, (t_ref, b_val) in temp_n << b_n:
#print(f" {n}, ({z_ref}, {b_val})")
t_ref += at_val * b_val
canvas.addFrame((k, m), (k, n), (m, k, n))
print("Intermediate Output - T")
displayTensor(t)
displayCanvas(canvas)
# -
# # Outerspace - Step 2 - Serial
# +
# t = Tensor(rank_ids=["M", "K", "N"]) - created in step 1 above
t.setColor("blue")
z = Tensor(rank_ids=["M", "N"])
print("Input Intermediate - T")
displayTensor(t)
canvas = createCanvas(t, z)
t_m = t.getRoot()
z_m = z.getRoot()
for m, (z_n, t_k) in z_m << t_m:
for k, t_n in t_k: # Coords are all ordered
for n, (z_ref, t_val) in z_n << t_n:
z_ref += t_val
canvas.addFrame((m, k, n), (m, n))
print("Tensor z")
displayTensor(z)
displayCanvas(canvas)
# -
# ## Check result
z_verify == z
# # Outerspace - Step 2 - Wide merge - Parallel (opportunity)
# +
# t = Tensor(rank_ids=["M", "K", "N"]) - created in step 1 above
t.setColor("blue")
z = Tensor(rank_ids=["M", "N"])
print("Input Intermediate - T")
displayTensor(t)
canvas = createCanvas(t, z)
t_m = t.getRoot()
z_m = z.getRoot()
for m, (z_n, t_k) in z_m << t_m:
print(f"\n\nt_k for coord m = {m}")
displayTensor(t_k)
tp_n = t_k.swapRanks() # The merge!
print(f"Transposed nt_k for coord m = {m} -> tp_n")
displayTensor(Tensor.fromFiber(["N", "K"], tp_n))
for n, (z_ref, tp_k) in z_n << tp_n: # Ordered coords
print(f"Sum tp_k = {tp_k}")
shards = []
for k, tp_val in tp_k: # Parallel!
z_ref += tp_val # Spatial reduction
shards.append((m, k, n))
#displayTensor(t, shards)
canvas.addFrame(shards, (m, n))
print(f"\nTensor z (for tp_n at coord m = {m} after all tp_k reductions)")
displayTensor(z)
print("Final Result - Z")
displayTensor(z)
displayCanvas(canvas)
# -
# ## Check result
z_verify == z
# # Outerspace - Step 2 - Narrrow merge - Parallel (opportunity)
#
# Implementation incomplete...
# +
# t = Tensor(rank_ids=["M", "K", "N"]) - created in step 1 above
z = Tensor(rank_ids=["M", "N"])
displayTensor(t)
t_m = t.getRoot()
z_m = z.getRoot()
# Want to do logrithmic merge!!!
for m, (z_n, t_k) in z_m << t_m:
displayTensor(t_k)
t_k1 = t_k.splitEqual(2)
displayTensor(t_k1)
for k1, t_k0 in t_k1:
print(f"\n\nt_k at coord m = {m} for split {k1}")
displayTensor(t_k)
tp_n = t_k0.swapRanks() # The merge!
print(f"Transposed = tp_n")
displayTensor(Tensor.fromFiber(["N", "K"], tp_n))
for n, (z_ref, tp_k) in z_n << tp_n: # Ordered coords
print(f"tp_k = {tp_k}")
for k, tp_val in tp_k: # Parallel!
z_ref += tp_val # Spatial reduction
print(f"\nTensor z (for tp_n for coord m = {m} after split {k1} tp_k reductions)")
displayTensor(z)
print("Final tensor z")
displayTensor(z)
# -
# ## Check result
z_verify == z
# ## Testing area
#
# For running alternative algorithms
displayTensor(a)
asplit = a.getRoot().splitEqual(1)
displayTensor(asplit)
| notebooks/sparse-gemm/outer-space.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import os
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import matplotlib.gridspec as gridspec
from scipy.stats import spearmanr
import numpy as np
data_dir = "/Users/amlalejini/DataPlayground/GPTP2018-MAPE-PROJ"
signalgp_data_fpath = os.path.join(data_dir, "signalgp_pop_data.csv")
scopegp_data_fpath = os.path.join(data_dir, "scopegp_trait_data_filtered.csv")
# Load data
signalgp_data = pd.read_csv(signalgp_data_fpath)
scopegp_data = pd.read_csv(scopegp_data_fpath)
# Filter data by problem
scopegp_LOGIC_ALL = scopegp_data[scopegp_data["problem"] == "LOGIC"]
scopegp_SQUARES_ALL = scopegp_data[scopegp_data["problem"] == "SQUARES"]
scopegp_SMALLEST_ALL = scopegp_data[scopegp_data["problem"] == "SMALLEST"]
scopegp_SUM_ALL = scopegp_data[scopegp_data["problem"] == "SUM"]
scopegp_COLLATZ_ALL = scopegp_data[scopegp_data["problem"] == "COLLATZ"]
scopegp_SYMREG_ALL = scopegp_data[scopegp_data["problem"] == "SYMREG"]
UPDATE = 50000
scopegp_LOGIC = scopegp_LOGIC_ALL[scopegp_LOGIC_ALL["update"] == UPDATE]
scopegp_SQUARES = scopegp_SQUARES_ALL[scopegp_SQUARES_ALL["update"] == UPDATE]
scopegp_SMALLEST = scopegp_SMALLEST_ALL[scopegp_SMALLEST_ALL["update"] == UPDATE]
scopegp_SUM = scopegp_SUM_ALL[scopegp_SUM_ALL["update"] == UPDATE]
scopegp_COLLATZ = scopegp_COLLATZ_ALL[scopegp_COLLATZ_ALL["update"] == UPDATE]
scopegp_SYMREG = scopegp_SYMREG_ALL[scopegp_SYMREG_ALL["update"] == UPDATE]
# +
scope_min_inst_ent = 0
scope_max_inst_ent = 5
min_scope_cnt = 0
max_scope_cnt = 17
# -
xy_label_fs = 18
xy_tick_fs = 14
cmap = sns.cubehelix_palette(as_cmap=True)
# ## Logic Problem
# +
# LOGIC problem
min_fitness = 10 # You must have at least 10 fitness to not be filtered.
# - RANDOM (all), TOURNAMENT (FILTERED), MAPE (FILTERED), LEX (FILTERED)
scopegp_LOGIC_RAND = scopegp_LOGIC[scopegp_LOGIC["selection_method"] == "RAND"]
scopegp_LOGIC_TOURN = scopegp_LOGIC[scopegp_LOGIC["selection_method"] == "TOURN"]
scopegp_LOGIC_MAPE = scopegp_LOGIC[scopegp_LOGIC["selection_method"] == "MAPE"]
scopegp_LOGIC_LEX = scopegp_LOGIC[scopegp_LOGIC["selection_method"] == "LEX"]
# Filter to only successful organisms
scopegp_LOGIC_TOURN = scopegp_LOGIC_TOURN[scopegp_LOGIC_TOURN["fitness"]>= min_fitness]
scopegp_LOGIC_MAPE = scopegp_LOGIC_MAPE[scopegp_LOGIC_MAPE["fitness"] >= min_fitness]
scopegp_LOGIC_LEX = scopegp_LOGIC_LEX[scopegp_LOGIC_LEX["fitness"] >= min_fitness]
# +
fig = plt.figure(1)
gridspec.GridSpec(1,16)
fig.set_size_inches(21, 7)
with sns.axes_style("darkgrid"):
ax1 = plt.subplot2grid((1,12), (0,0), colspan=3)
sns.boxplot(x="selection_method", y="fitness", data=scopegp_LOGIC, ax=ax1)
# sns.swarmplot(x="selection_method", y="fitness", data=scopegp_LOGIC, ax=ax1, color=".1")
ax1.set_xlabel("FITNESS")
ax1.set_ylabel("Fitness")
# ax1.set_ylim(min_fitness, max_fitness + 10)
# for tick in ax1.get_yticklabels():
# tick.set_fontsize(y_tick_fs)
# for tick in ax1.get_xticklabels():
# tick.set_fontsize(x_tick_fs)
# ax1.yaxis.label.set_fontsize(y_label_fs)
# ax1.xaxis.label.set_fontsize(x_label_fs)
ax1.xaxis.set_label_position('top')
ax2 = plt.subplot2grid((1,12), (0,4), colspan=3)
p2 = sns.boxplot(x="selection_method", y="instruction_entropy", data=scopegp_LOGIC, ax=ax2)
# sns.swarmplot(x="sim_thresh", y="fitness", data=evo_df_DS0[evo_df_DS0["update"] == 10000], ax=ax2, color=".1")
# ax2.set_xticklabels(labels)
ax2.set_xlabel("INSTRUCTION ENTROPY")
# ax2.set_ylabel("")
# plt.setp(ax2.get_yticklabels(), visible = False)
# ax2.set_ylim(min_fitness, max_fitness + 10)
# for tick in ax2.get_yticklabels():
# tick.set_fontsize(y_tick_fs)
# for tick in ax2.get_xticklabels():
# tick.set_fontsize(x_tick_fs)
# ax2.yaxis.label.set_fontsize(y_label_fs)
# ax2.xaxis.label.set_fontsize(x_label_fs)
ax2.xaxis.set_label_position('top')
ax3 = plt.subplot2grid((1,12), (0,8), colspan=3)
p3 = sns.boxplot(x="selection_method", y="scope_count", data=scopegp_LOGIC, ax=ax3)
# sns.swarmplot(x="sim_thresh", y="fitness", data=evo_df_DS0[evo_df_DS0["update"] == 10000], ax=ax2, color=".1")
# ax2.set_xticklabels(labels)
ax3.set_xlabel("SCOPE COUNT")
ax3.xaxis.set_label_position('top')
# -
# ### Logic Problem - Tournament
# +
fig = plt.figure(1)
fig.set_size_inches(7,7)
with sns.axes_style("white"):
g = sns.jointplot(data=scopegp_LOGIC_TOURN, x="instruction_entropy", y="scope_count", kind="kde",
xlim=(scope_min_inst_ent, scope_max_inst_ent), ylim=(min_scope_cnt, max_scope_cnt),
stat_func=None, shade=True, cmap=cmap, color="Grey", shade_lowest=False)
g.set_axis_labels("Instruction Entropy", "Scopes Used")
ax = g.ax_joint
ax.xaxis.label.set_fontsize(xy_label_fs)
ax.yaxis.label.set_fontsize(xy_label_fs)
for tick in ax.get_xticklabels():
tick.set_fontsize(xy_tick_fs)
for tick in ax.get_yticklabels():
tick.set_fontsize(xy_tick_fs)
plt.title('Logic Problem - Tournament ', y=1.2, x=-3)
# -
# ### Logic Problem - Map-Elites
fig = plt.figure(1)
fig.set_size_inches(7,7)
with sns.axes_style("white"):
g = sns.jointplot(data=scopegp_LOGIC_MAPE, x="instruction_entropy", y="scope_count", kind="kde",
xlim=(scope_min_inst_ent, scope_max_inst_ent), ylim=(min_scope_cnt, max_scope_cnt),
stat_func=None, shade=True, cmap=cmap, shade_lowest=False, color="Grey")
g.set_axis_labels("Instruction Entropy", "Scopes Used")
ax = g.ax_joint
ax.xaxis.label.set_fontsize(xy_label_fs)
ax.yaxis.label.set_fontsize(xy_label_fs)
for tick in ax.get_xticklabels():
tick.set_fontsize(xy_tick_fs)
for tick in ax.get_yticklabels():
tick.set_fontsize(xy_tick_fs)
plt.title('Logic Problem - MAPE ', y=1.2, x=-3)
# ### Logic Problem - Lexicase
fig = plt.figure(1)
fig.set_size_inches(7,7)
with sns.axes_style("white"):
g = sns.jointplot(data=scopegp_LOGIC_LEX, x="instruction_entropy", y="scope_count", kind="kde",
xlim=(scope_min_inst_ent, scope_max_inst_ent), ylim=(min_scope_cnt, max_scope_cnt),
stat_func=None, shade=True, cmap=cmap, shade_lowest=False, color="Grey")
g.set_axis_labels("Instruction Entropy", "Scopes Used")
ax = g.ax_joint
ax.xaxis.label.set_fontsize(xy_label_fs)
ax.yaxis.label.set_fontsize(xy_label_fs)
for tick in ax.get_xticklabels():
tick.set_fontsize(xy_tick_fs)
for tick in ax.get_yticklabels():
tick.set_fontsize(xy_tick_fs)
plt.title('Logic Problem - Lexicase ', y=1.2, x=-3)
# ### Logic Problem - Random drift
fig = plt.figure(1)
fig.set_size_inches(7,7)
with sns.axes_style("white"):
g = sns.jointplot(data=scopegp_LOGIC_RAND, x="instruction_entropy", y="scope_count", kind="kde",
xlim=(scope_min_inst_ent, scope_max_inst_ent), ylim=(min_scope_cnt, max_scope_cnt),
stat_func=None, shade=True, cmap=cmap, shade_lowest=False, color="Grey")
g.set_axis_labels("Instruction Entropy", "Scopes Used")
ax = g.ax_joint
ax.xaxis.label.set_fontsize(xy_label_fs)
ax.yaxis.label.set_fontsize(xy_label_fs)
for tick in ax.get_xticklabels():
tick.set_fontsize(xy_tick_fs)
for tick in ax.get_yticklabels():
tick.set_fontsize(xy_tick_fs)
plt.title('Logic Problem - Random Drift ', y=1.2, x=-3)
# ---
# ## Squares Problem
# +
# LOGIC problem
min_fitness = 10000 # You must have at least 10 fitness to not be filtered.
# - RANDOM (all), TOURNAMENT (FILTERED), MAPE (FILTERED), LEX (FILTERED)
scopegp_SQUARES_RAND = scopegp_SQUARES[scopegp_SQUARES["selection_method"] == "RAND"]
scopegp_SQUARES_TOURN = scopegp_SQUARES[scopegp_SQUARES["selection_method"] == "TOURN"]
scopegp_SQUARES_MAPE = scopegp_SQUARES[scopegp_SQUARES["selection_method"] == "MAPE"]
scopegp_SQUARES_LEX = scopegp_SQUARES[scopegp_SQUARES["selection_method"] == "LEX"]
# Filter to only successful organisms
scopegp_SQUARES_TOURN = scopegp_SQUARES_TOURN[scopegp_SQUARES_TOURN["fitness"]>= min_fitness]
scopegp_SQUARES_MAPE = scopegp_SQUARES_MAPE[scopegp_SQUARES_MAPE["fitness"] >= min_fitness]
scopegp_SQUARES_LEX = scopegp_SQUARES_LEX[scopegp_SQUARES_LEX["fitness"] >= min_fitness]
# +
fig = plt.figure(1)
gridspec.GridSpec(1,16)
fig.set_size_inches(21, 7)
with sns.axes_style("darkgrid"):
ax1 = plt.subplot2grid((1,12), (0,0), colspan=3)
sns.boxplot(x="selection_method", y="fitness", data=scopegp_SQUARES, ax=ax1)
ax1.set_xlabel("FITNESS")
ax1.set_ylabel("Fitness")
ax1.xaxis.set_label_position('top')
ax2 = plt.subplot2grid((1,12), (0,4), colspan=3)
p2 = sns.boxplot(x="selection_method", y="instruction_entropy", data=scopegp_SQUARES, ax=ax2)
ax2.set_xlabel("INSTRUCTION ENTROPY")
ax2.xaxis.set_label_position('top')
ax3 = plt.subplot2grid((1,12), (0,8), colspan=3)
p3 = sns.boxplot(x="selection_method", y="scope_count", data=scopegp_SQUARES, ax=ax3)
ax3.set_xlabel("SCOPE COUNT")
ax3.xaxis.set_label_position('top')
# -
# ### Squares Problem - Tournament
# +
fig = plt.figure(1)
fig.set_size_inches(7,7)
with sns.axes_style("white"):
g = sns.jointplot(data=scopegp_SQUARES_TOURN, x="instruction_entropy", y="scope_count", kind="kde",
xlim=(scope_min_inst_ent, scope_max_inst_ent), ylim=(min_scope_cnt, max_scope_cnt),
stat_func=None, shade=True, cmap=cmap, color="Grey", shade_lowest=False)
g.set_axis_labels("Instruction Entropy", "Scopes Used")
ax = g.ax_joint
ax.xaxis.label.set_fontsize(xy_label_fs)
ax.yaxis.label.set_fontsize(xy_label_fs)
for tick in ax.get_xticklabels():
tick.set_fontsize(xy_tick_fs)
for tick in ax.get_yticklabels():
tick.set_fontsize(xy_tick_fs)
plt.title('Squares Problem - Tournament ', y=1.2, x=-3)
# -
# ### Squares Problem - Map-Elites
fig = plt.figure(1)
fig.set_size_inches(7,7)
with sns.axes_style("white"):
g = sns.jointplot(data=scopegp_SQUARES_MAPE, x="instruction_entropy", y="scope_count", kind="kde",
xlim=(scope_min_inst_ent, scope_max_inst_ent), ylim=(min_scope_cnt, max_scope_cnt),
stat_func=None, shade=True, cmap=cmap, shade_lowest=False, color="Grey")
g.set_axis_labels("Instruction Entropy", "Scopes Used")
ax = g.ax_joint
ax.xaxis.label.set_fontsize(xy_label_fs)
ax.yaxis.label.set_fontsize(xy_label_fs)
for tick in ax.get_xticklabels():
tick.set_fontsize(xy_tick_fs)
for tick in ax.get_yticklabels():
tick.set_fontsize(xy_tick_fs)
plt.title('Squares Problem - MAPE ', y=1.2, x=-3)
# ### Squares Problem - Lexicase
fig = plt.figure(1)
fig.set_size_inches(7,7)
with sns.axes_style("white"):
g = sns.jointplot(data=scopegp_SQUARES_LEX, x="instruction_entropy", y="scope_count", kind="kde",
xlim=(scope_min_inst_ent, scope_max_inst_ent), ylim=(min_scope_cnt, max_scope_cnt),
stat_func=None, shade=True, cmap=cmap, shade_lowest=False, color="Grey")
g.set_axis_labels("Instruction Entropy", "Scopes Used")
ax = g.ax_joint
ax.xaxis.label.set_fontsize(xy_label_fs)
ax.yaxis.label.set_fontsize(xy_label_fs)
for tick in ax.get_xticklabels():
tick.set_fontsize(xy_tick_fs)
for tick in ax.get_yticklabels():
tick.set_fontsize(xy_tick_fs)
plt.title('Squares Problem - Lexicase ', y=1.2, x=-3)
# ### Squares Problem - Random drift
fig = plt.figure(1)
fig.set_size_inches(7,7)
with sns.axes_style("white"):
g = sns.jointplot(data=scopegp_SQUARES_RAND, x="instruction_entropy", y="scope_count", kind="kde",
xlim=(scope_min_inst_ent, scope_max_inst_ent), ylim=(min_scope_cnt, max_scope_cnt),
stat_func=None, shade=True, cmap=cmap, shade_lowest=False, color="Grey")
g.set_axis_labels("Instruction Entropy", "Scopes Used")
ax = g.ax_joint
ax.xaxis.label.set_fontsize(xy_label_fs)
ax.yaxis.label.set_fontsize(xy_label_fs)
for tick in ax.get_xticklabels():
tick.set_fontsize(xy_tick_fs)
for tick in ax.get_yticklabels():
tick.set_fontsize(xy_tick_fs)
plt.title('Squares Problem - Random Drift ', y=1.2, x=-3)
# ---
# ## Smallest Problem
# +
# SMALLEST problem
min_fitness = 200000 # You must have at least 10 fitness to not be filtered.
# - RANDOM (all), TOURNAMENT (FILTERED), MAPE (FILTERED), LEX (FILTERED)
scopegp_SMALLEST_RAND = scopegp_SMALLEST[scopegp_SMALLEST["selection_method"] == "RAND"]
scopegp_SMALLEST_TOURN = scopegp_SMALLEST[scopegp_SMALLEST["selection_method"] == "TOURN"]
scopegp_SMALLEST_MAPE = scopegp_SMALLEST[scopegp_SMALLEST["selection_method"] == "MAPE"]
scopegp_SMALLEST_LEX = scopegp_SMALLEST[scopegp_SMALLEST["selection_method"] == "LEX"]
# Filter to only successful organisms
scopegp_SMALLEST_TOURN = scopegp_SMALLEST_TOURN[scopegp_SMALLEST_TOURN["fitness"]>= min_fitness]
scopegp_SMALLEST_MAPE = scopegp_SMALLEST_MAPE[scopegp_SMALLEST_MAPE["fitness"] >= min_fitness]
scopegp_SMALLEST_LEX = scopegp_SMALLEST_LEX[scopegp_SMALLEST_LEX["fitness"] >= min_fitness]
# +
fig = plt.figure(1)
gridspec.GridSpec(1,16)
fig.set_size_inches(21, 7)
with sns.axes_style("darkgrid"):
ax1 = plt.subplot2grid((1,12), (0,0), colspan=3)
sns.boxplot(x="selection_method", y="fitness", data=scopegp_SMALLEST, ax=ax1)
ax1.set_xlabel("FITNESS")
ax1.set_ylabel("Fitness")
ax1.xaxis.set_label_position('top')
ax2 = plt.subplot2grid((1,12), (0,4), colspan=3)
p2 = sns.boxplot(x="selection_method", y="instruction_entropy", data=scopegp_SMALLEST, ax=ax2)
ax2.set_xlabel("INSTRUCTION ENTROPY")
ax2.xaxis.set_label_position('top')
ax3 = plt.subplot2grid((1,12), (0,8), colspan=3)
p3 = sns.boxplot(x="selection_method", y="scope_count", data=scopegp_SMALLEST, ax=ax3)
ax3.set_xlabel("SCOPE COUNT")
ax3.xaxis.set_label_position('top')
# -
# ### Smallest Problem - Tournament
# +
fig = plt.figure(1)
fig.set_size_inches(7,7)
with sns.axes_style("white"):
g = sns.jointplot(data=scopegp_SMALLEST_TOURN, x="instruction_entropy", y="scope_count", kind="kde",
xlim=(scope_min_inst_ent, scope_max_inst_ent), ylim=(min_scope_cnt, max_scope_cnt),
stat_func=None, shade=True, cmap=cmap, color="Grey", shade_lowest=False)
g.set_axis_labels("Instruction Entropy", "Scopes Used")
ax = g.ax_joint
ax.xaxis.label.set_fontsize(xy_label_fs)
ax.yaxis.label.set_fontsize(xy_label_fs)
for tick in ax.get_xticklabels():
tick.set_fontsize(xy_tick_fs)
for tick in ax.get_yticklabels():
tick.set_fontsize(xy_tick_fs)
plt.title('SMALLEST Problem - Tournament ', y=1.2, x=-3)
# -
# ### SMALLEST Problem - Map-Elites
fig = plt.figure(1)
fig.set_size_inches(7,7)
with sns.axes_style("white"):
g = sns.jointplot(data=scopegp_SMALLEST_MAPE, x="instruction_entropy", y="scope_count", kind="kde",
xlim=(scope_min_inst_ent, scope_max_inst_ent), ylim=(min_scope_cnt, max_scope_cnt),
stat_func=None, shade=True, cmap=cmap, shade_lowest=False, color="Grey")
g.set_axis_labels("Instruction Entropy", "Scopes Used")
ax = g.ax_joint
ax.xaxis.label.set_fontsize(xy_label_fs)
ax.yaxis.label.set_fontsize(xy_label_fs)
for tick in ax.get_xticklabels():
tick.set_fontsize(xy_tick_fs)
for tick in ax.get_yticklabels():
tick.set_fontsize(xy_tick_fs)
plt.title('SMALLEST Problem - MAPE ', y=1.2, x=-3)
# ### SMALLEST Problem - Lexicase
fig = plt.figure(1)
fig.set_size_inches(7,7)
with sns.axes_style("white"):
g = sns.jointplot(data=scopegp_SMALLEST_LEX, x="instruction_entropy", y="scope_count", kind="kde",
xlim=(scope_min_inst_ent, scope_max_inst_ent), ylim=(min_scope_cnt, max_scope_cnt),
stat_func=None, shade=True, cmap=cmap, shade_lowest=False, color="Grey")
g.set_axis_labels("Instruction Entropy", "Scopes Used")
ax = g.ax_joint
ax.xaxis.label.set_fontsize(xy_label_fs)
ax.yaxis.label.set_fontsize(xy_label_fs)
for tick in ax.get_xticklabels():
tick.set_fontsize(xy_tick_fs)
for tick in ax.get_yticklabels():
tick.set_fontsize(xy_tick_fs)
plt.title('SMALLEST Problem - Lexicase ', y=1.2, x=-3)
# ### SMALLEST Problem - Random drift
fig = plt.figure(1)
fig.set_size_inches(7,7)
with sns.axes_style("white"):
g = sns.jointplot(data=scopegp_SMALLEST_RAND, x="instruction_entropy", y="scope_count", kind="kde",
xlim=(scope_min_inst_ent, scope_max_inst_ent), ylim=(min_scope_cnt, max_scope_cnt),
stat_func=None, shade=True, cmap=cmap, shade_lowest=False, color="Grey")
g.set_axis_labels("Instruction Entropy", "Scopes Used")
ax = g.ax_joint
ax.xaxis.label.set_fontsize(xy_label_fs)
ax.yaxis.label.set_fontsize(xy_label_fs)
for tick in ax.get_xticklabels():
tick.set_fontsize(xy_tick_fs)
for tick in ax.get_yticklabels():
tick.set_fontsize(xy_tick_fs)
plt.title('SMALLEST Problem - Random Drift ', y=1.2, x=-3)
# ---
# ## SUM PROBLEM
# +
# SUM problem
min_fitness = 200000 # You must have at least 10 fitness to not be filtered.
# - RANDOM (all), TOURNAMENT (FILTERED), MAPE (FILTERED), LEX (FILTERED)
scopegp_SUM_RAND = scopegp_SUM[scopegp_SUM["selection_method"] == "RAND"]
scopegp_SUM_TOURN = scopegp_SUM[scopegp_SUM["selection_method"] == "TOURN"]
scopegp_SUM_MAPE = scopegp_SUM[scopegp_SUM["selection_method"] == "MAPE"]
scopegp_SUM_LEX = scopegp_SUM[scopegp_SUM["selection_method"] == "LEX"]
# Filter to only successful organisms
scopegp_SUM_TOURN = scopegp_SUM_TOURN[scopegp_SUM_TOURN["fitness"]>= min_fitness]
scopegp_SUM_MAPE = scopegp_SUM_MAPE[scopegp_SUM_MAPE["fitness"] >= min_fitness]
scopegp_SUM_LEX = scopegp_SUM_LEX[scopegp_SUM_LEX["fitness"] >= min_fitness]
# +
fig = plt.figure(1)
gridspec.GridSpec(1,16)
fig.set_size_inches(21, 7)
with sns.axes_style("darkgrid"):
ax1 = plt.subplot2grid((1,12), (0,0), colspan=3)
sns.boxplot(x="selection_method", y="fitness", data=scopegp_SUM, ax=ax1)
ax1.set_xlabel("FITNESS")
ax1.set_ylabel("Fitness")
ax1.xaxis.set_label_position('top')
ax2 = plt.subplot2grid((1,12), (0,4), colspan=3)
p2 = sns.boxplot(x="selection_method", y="instruction_entropy", data=scopegp_SUM, ax=ax2)
ax2.set_xlabel("INSTRUCTION ENTROPY")
ax2.xaxis.set_label_position('top')
ax3 = plt.subplot2grid((1,12), (0,8), colspan=3)
p3 = sns.boxplot(x="selection_method", y="scope_count", data=scopegp_SUM, ax=ax3)
ax3.set_xlabel("SCOPE COUNT")
ax3.xaxis.set_label_position('top')
# -
# ### SUM Problem - Tournament
# +
fig = plt.figure(1)
fig.set_size_inches(7,7)
with sns.axes_style("white"):
g = sns.jointplot(data=scopegp_SUM_TOURN, x="instruction_entropy", y="scope_count", kind="kde",
xlim=(scope_min_inst_ent, scope_max_inst_ent), ylim=(min_scope_cnt, max_scope_cnt),
stat_func=None, shade=True, cmap=cmap, color="Grey", shade_lowest=False)
g.set_axis_labels("Instruction Entropy", "Scopes Used")
ax = g.ax_joint
ax.xaxis.label.set_fontsize(xy_label_fs)
ax.yaxis.label.set_fontsize(xy_label_fs)
for tick in ax.get_xticklabels():
tick.set_fontsize(xy_tick_fs)
for tick in ax.get_yticklabels():
tick.set_fontsize(xy_tick_fs)
plt.title('SUM Problem - Tournament ', y=1.2, x=-3)
# -
# ### SUM Problem - Map-Elites
fig = plt.figure(1)
fig.set_size_inches(7,7)
with sns.axes_style("white"):
g = sns.jointplot(data=scopegp_SUM_MAPE, x="instruction_entropy", y="scope_count", kind="kde",
xlim=(scope_min_inst_ent, scope_max_inst_ent), ylim=(min_scope_cnt, max_scope_cnt),
stat_func=None, shade=True, cmap=cmap, shade_lowest=False, color="Grey")
g.set_axis_labels("Instruction Entropy", "Scopes Used")
ax = g.ax_joint
ax.xaxis.label.set_fontsize(xy_label_fs)
ax.yaxis.label.set_fontsize(xy_label_fs)
for tick in ax.get_xticklabels():
tick.set_fontsize(xy_tick_fs)
for tick in ax.get_yticklabels():
tick.set_fontsize(xy_tick_fs)
plt.title('SUM Problem - MAPE ', y=1.2, x=-3)
# ### SUM Problem - Lexicase
fig = plt.figure(1)
fig.set_size_inches(7,7)
with sns.axes_style("white"):
g = sns.jointplot(data=scopegp_SUM_LEX, x="instruction_entropy", y="scope_count", kind="kde",
xlim=(scope_min_inst_ent, scope_max_inst_ent), ylim=(min_scope_cnt, max_scope_cnt),
stat_func=None, shade=True, cmap=cmap, shade_lowest=False, color="Grey")
g.set_axis_labels("Instruction Entropy", "Scopes Used")
ax = g.ax_joint
ax.xaxis.label.set_fontsize(xy_label_fs)
ax.yaxis.label.set_fontsize(xy_label_fs)
for tick in ax.get_xticklabels():
tick.set_fontsize(xy_tick_fs)
for tick in ax.get_yticklabels():
tick.set_fontsize(xy_tick_fs)
plt.title('SUM Problem - Lexicase ', y=1.2, x=-3)
# ### SUM Problem - Random drift
fig = plt.figure(1)
fig.set_size_inches(7,7)
with sns.axes_style("white"):
g = sns.jointplot(data=scopegp_SUM_RAND, x="instruction_entropy", y="scope_count", kind="kde",
xlim=(scope_min_inst_ent, scope_max_inst_ent), ylim=(min_scope_cnt, max_scope_cnt),
stat_func=None, shade=True, cmap=cmap, shade_lowest=False, color="Grey")
g.set_axis_labels("Instruction Entropy", "Scopes Used")
ax = g.ax_joint
ax.xaxis.label.set_fontsize(xy_label_fs)
ax.yaxis.label.set_fontsize(xy_label_fs)
for tick in ax.get_xticklabels():
tick.set_fontsize(xy_tick_fs)
for tick in ax.get_yticklabels():
tick.set_fontsize(xy_tick_fs)
plt.title('SUM Problem - Random Drift ', y=1.2, x=-3)
# ----
#
# ## COLLATZ PROBLEM
# +
# COLLATZ problem
min_fitness = 175000 # You must have at least 10 fitness to not be filtered.
# - RANDOM (all), TOURNAMENT (FILTERED), MAPE (FILTERED), LEX (FILTERED)
scopegp_COLLATZ_RAND = scopegp_COLLATZ[scopegp_COLLATZ["selection_method"] == "RAND"]
scopegp_COLLATZ_TOURN = scopegp_COLLATZ[scopegp_COLLATZ["selection_method"] == "TOURN"]
scopegp_COLLATZ_MAPE = scopegp_COLLATZ[scopegp_COLLATZ["selection_method"] == "MAPE"]
scopegp_COLLATZ_LEX = scopegp_COLLATZ[scopegp_COLLATZ["selection_method"] == "LEX"]
# Filter to only successful organisms
scopegp_COLLATZ_TOURN = scopegp_COLLATZ_TOURN[scopegp_COLLATZ_TOURN["fitness"]>= min_fitness]
scopegp_COLLATZ_MAPE = scopegp_COLLATZ_MAPE[scopegp_COLLATZ_MAPE["fitness"] >= min_fitness]
scopegp_COLLATZ_LEX = scopegp_COLLATZ_LEX[scopegp_COLLATZ_LEX["fitness"] >= min_fitness]
# +
fig = plt.figure(1)
gridspec.GridSpec(1,16)
fig.set_size_inches(21, 7)
with sns.axes_style("darkgrid"):
ax1 = plt.subplot2grid((1,12), (0,0), colspan=3)
sns.boxplot(x="selection_method", y="fitness", data=scopegp_COLLATZ, ax=ax1)
ax1.set_xlabel("FITNESS")
ax1.set_ylabel("Fitness")
ax1.xaxis.set_label_position('top')
ax2 = plt.subplot2grid((1,12), (0,4), colspan=3)
p2 = sns.boxplot(x="selection_method", y="instruction_entropy", data=scopegp_COLLATZ, ax=ax2)
ax2.set_xlabel("INSTRUCTION ENTROPY")
ax2.xaxis.set_label_position('top')
ax3 = plt.subplot2grid((1,12), (0,8), colspan=3)
p3 = sns.boxplot(x="selection_method", y="scope_count", data=scopegp_COLLATZ, ax=ax3)
ax3.set_xlabel("SCOPE COUNT")
ax3.xaxis.set_label_position('top')
# -
# ### COLLATZ Problem - Tournament
# +
fig = plt.figure(1)
fig.set_size_inches(7,7)
with sns.axes_style("white"):
g = sns.jointplot(data=scopegp_COLLATZ_TOURN, x="instruction_entropy", y="scope_count", kind="kde",
xlim=(scope_min_inst_ent, scope_max_inst_ent), ylim=(min_scope_cnt, max_scope_cnt),
stat_func=None, shade=True, cmap=cmap, color="Grey", shade_lowest=False)
g.set_axis_labels("Instruction Entropy", "Scopes Used")
ax = g.ax_joint
ax.xaxis.label.set_fontsize(xy_label_fs)
ax.yaxis.label.set_fontsize(xy_label_fs)
for tick in ax.get_xticklabels():
tick.set_fontsize(xy_tick_fs)
for tick in ax.get_yticklabels():
tick.set_fontsize(xy_tick_fs)
plt.title('COLLATZ Problem - Tournament ', y=1.2, x=-3)
# -
# ### COLLATZ Problem - Map-Elites
fig = plt.figure(1)
fig.set_size_inches(7,7)
with sns.axes_style("white"):
g = sns.jointplot(data=scopegp_COLLATZ_MAPE, x="instruction_entropy", y="scope_count", kind="kde",
xlim=(scope_min_inst_ent, scope_max_inst_ent), ylim=(min_scope_cnt, max_scope_cnt),
stat_func=None, shade=True, cmap=cmap, shade_lowest=False, color="Grey")
g.set_axis_labels("Instruction Entropy", "Scopes Used")
ax = g.ax_joint
ax.xaxis.label.set_fontsize(xy_label_fs)
ax.yaxis.label.set_fontsize(xy_label_fs)
for tick in ax.get_xticklabels():
tick.set_fontsize(xy_tick_fs)
for tick in ax.get_yticklabels():
tick.set_fontsize(xy_tick_fs)
plt.title('COLLATZ Problem - MAPE ', y=1.2, x=-3)
# ### COLLATZ Problem - Lexicase
fig = plt.figure(1)
fig.set_size_inches(7,7)
with sns.axes_style("white"):
g = sns.jointplot(data=scopegp_COLLATZ_LEX, x="instruction_entropy", y="scope_count", kind="kde",
xlim=(scope_min_inst_ent, scope_max_inst_ent), ylim=(min_scope_cnt, max_scope_cnt),
stat_func=None, shade=True, cmap=cmap, shade_lowest=False, color="Grey")
g.set_axis_labels("Instruction Entropy", "Scopes Used")
ax = g.ax_joint
ax.xaxis.label.set_fontsize(xy_label_fs)
ax.yaxis.label.set_fontsize(xy_label_fs)
for tick in ax.get_xticklabels():
tick.set_fontsize(xy_tick_fs)
for tick in ax.get_yticklabels():
tick.set_fontsize(xy_tick_fs)
plt.title('COLLATZ Problem - Lexicase ', y=1.2, x=-3)
# ### COLLATZ Problem - Random drift
fig = plt.figure(1)
fig.set_size_inches(7,7)
with sns.axes_style("white"):
g = sns.jointplot(data=scopegp_COLLATZ_RAND, x="instruction_entropy", y="scope_count", kind="kde",
xlim=(scope_min_inst_ent, scope_max_inst_ent), ylim=(min_scope_cnt, max_scope_cnt),
stat_func=None, shade=True, cmap=cmap, shade_lowest=False, color="Grey")
g.set_axis_labels("Instruction Entropy", "Scopes Used")
ax = g.ax_joint
ax.xaxis.label.set_fontsize(xy_label_fs)
ax.yaxis.label.set_fontsize(xy_label_fs)
for tick in ax.get_xticklabels():
tick.set_fontsize(xy_tick_fs)
for tick in ax.get_yticklabels():
tick.set_fontsize(xy_tick_fs)
plt.title('COLLATZ Problem - Random Drift ', y=1.2, x=-3)
# ---
#
# ## SYMREG PROBLEM
# +
# SYMREG problem
min_fitness = 200000 # You must have at least 10 fitness to not be filtered.
# - RANDOM (all), TOURNAMENT (FILTERED), MAPE (FILTERED), LEX (FILTERED)
scopegp_SYMREG_RAND = scopegp_SYMREG[scopegp_SYMREG["selection_method"] == "RAND"]
scopegp_SYMREG_TOURN = scopegp_SYMREG[scopegp_SYMREG["selection_method"] == "TOURN"]
scopegp_SYMREG_MAPE = scopegp_SYMREG[scopegp_SYMREG["selection_method"] == "MAPE"]
scopegp_SYMREG_LEX = scopegp_SYMREG[scopegp_SYMREG["selection_method"] == "LEX"]
# Filter to only successful organisms
scopegp_SYMREG_TOURN = scopegp_SYMREG_TOURN[scopegp_SYMREG_TOURN["fitness"]>= min_fitness]
scopegp_SYMREG_MAPE = scopegp_SYMREG_MAPE[scopegp_SYMREG_MAPE["fitness"] >= min_fitness]
scopegp_SYMREG_LEX = scopegp_SYMREG_LEX[scopegp_SYMREG_LEX["fitness"] >= min_fitness]
# +
fig = plt.figure(1)
gridspec.GridSpec(1,16)
fig.set_size_inches(21, 7)
with sns.axes_style("darkgrid"):
ax1 = plt.subplot2grid((1,12), (0,0), colspan=3)
sns.boxplot(x="selection_method", y="fitness", data=scopegp_SYMREG, ax=ax1)
ax1.set_xlabel("FITNESS")
ax1.set_ylabel("Fitness")
ax1.xaxis.set_label_position('top')
ax2 = plt.subplot2grid((1,12), (0,4), colspan=3)
p2 = sns.boxplot(x="selection_method", y="instruction_entropy", data=scopegp_SYMREG, ax=ax2)
ax2.set_xlabel("INSTRUCTION ENTROPY")
ax2.xaxis.set_label_position('top')
ax3 = plt.subplot2grid((1,12), (0,8), colspan=3)
p3 = sns.boxplot(x="selection_method", y="scope_count", data=scopegp_SYMREG, ax=ax3)
ax3.set_xlabel("SCOPE COUNT")
ax3.xaxis.set_label_position('top')
# -
# ### SYMREG Problem - Tournament
# +
fig = plt.figure(1)
fig.set_size_inches(7,7)
with sns.axes_style("white"):
g = sns.jointplot(data=scopegp_SYMREG_TOURN, x="instruction_entropy", y="scope_count", kind="kde",
xlim=(scope_min_inst_ent, scope_max_inst_ent), ylim=(min_scope_cnt, max_scope_cnt),
stat_func=None, shade=True, cmap=cmap, color="Grey", shade_lowest=False)
g.set_axis_labels("Instruction Entropy", "Scopes Used")
ax = g.ax_joint
ax.xaxis.label.set_fontsize(xy_label_fs)
ax.yaxis.label.set_fontsize(xy_label_fs)
for tick in ax.get_xticklabels():
tick.set_fontsize(xy_tick_fs)
for tick in ax.get_yticklabels():
tick.set_fontsize(xy_tick_fs)
plt.title('SYMREG Problem - Tournament ', y=1.2, x=-3)
# -
# ### SYMREG Problem - Map-Elites
fig = plt.figure(1)
fig.set_size_inches(7,7)
with sns.axes_style("white"):
g = sns.jointplot(data=scopegp_SYMREG_MAPE, x="instruction_entropy", y="scope_count", kind="kde",
xlim=(scope_min_inst_ent, scope_max_inst_ent), ylim=(min_scope_cnt, max_scope_cnt),
stat_func=None, shade=True, cmap=cmap, shade_lowest=False, color="Grey")
g.set_axis_labels("Instruction Entropy", "Scopes Used")
ax = g.ax_joint
ax.xaxis.label.set_fontsize(xy_label_fs)
ax.yaxis.label.set_fontsize(xy_label_fs)
for tick in ax.get_xticklabels():
tick.set_fontsize(xy_tick_fs)
for tick in ax.get_yticklabels():
tick.set_fontsize(xy_tick_fs)
plt.title('SYMREG Problem - MAPE ', y=1.2, x=-3)
# ### SYMREG Problem - Lexicase
fig = plt.figure(1)
fig.set_size_inches(7,7)
with sns.axes_style("white"):
g = sns.jointplot(data=scopegp_SYMREG_LEX, x="instruction_entropy", y="scope_count", kind="kde",
xlim=(scope_min_inst_ent, scope_max_inst_ent), ylim=(min_scope_cnt, max_scope_cnt),
stat_func=None, shade=True, cmap=cmap, shade_lowest=False, color="Grey")
g.set_axis_labels("Instruction Entropy", "Scopes Used")
ax = g.ax_joint
ax.xaxis.label.set_fontsize(xy_label_fs)
ax.yaxis.label.set_fontsize(xy_label_fs)
for tick in ax.get_xticklabels():
tick.set_fontsize(xy_tick_fs)
for tick in ax.get_yticklabels():
tick.set_fontsize(xy_tick_fs)
plt.title('SYMREG Problem - Lexicase ', y=1.2, x=-3)
# ### SYMREG Problem - Random drift
fig = plt.figure(1)
fig.set_size_inches(7,7)
with sns.axes_style("white"):
g = sns.jointplot(data=scopegp_SYMREG_RAND, x="instruction_entropy", y="scope_count", kind="kde",
xlim=(scope_min_inst_ent, scope_max_inst_ent), ylim=(min_scope_cnt, max_scope_cnt),
stat_func=None, shade=True, cmap=cmap, shade_lowest=False, color="Grey")
g.set_axis_labels("Instruction Entropy", "Scopes Used")
ax = g.ax_joint
ax.xaxis.label.set_fontsize(xy_label_fs)
ax.yaxis.label.set_fontsize(xy_label_fs)
for tick in ax.get_xticklabels():
tick.set_fontsize(xy_tick_fs)
for tick in ax.get_yticklabels():
tick.set_fontsize(xy_tick_fs)
plt.title('SYMREG Problem - Random Drift ', y=1.2, x=-3)
| analysis/data_exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:mvdev] *
# language: python
# name: conda-env-mvdev-py
# ---
# # Assessing the Conditional Independence Views Requirement of Multi-view Spectral Clustering
import numpy as np
from numpy.random import multivariate_normal
import scipy as scp
from mvlearn.cluster.mv_spectral import MultiviewSpectralClustering
from sklearn.cluster import SpectralClustering
from sklearn.metrics import normalized_mutual_info_score as nmi_score
from sklearn.datasets import fetch_covtype
import matplotlib.pyplot as plt
# %matplotlib inline
from sklearn.manifold import TSNE
import warnings
warnings.filterwarnings("ignore")
RANDOM_SEED=10
# ##### Creating an artificial dataset where the conditional independence assumption between views holds
#
# Here, we create an artificial dataset where the conditional independence assumption between
# views, given the true labels, is enforced. Our artificial dataset is derived from the forest
# covertypes dataset from the scikit-learn package. This dataset is comprised of 7 different classes, with
# with 54 different numerical features per sample. To create our artificial data, we will select 500 samples from
# each of the first 6 classes in the dataset, and from these, construct 3 artificial classes with
# 2 views each. <br>
def get_ci_data(num_samples=500):
#Load in the vectorized news group data from scikit-learn package
cov = fetch_covtype()
all_data = np.array(cov.data)
all_targets = np.array(cov.target)
#Set class pairings as described in the multiview clustering paper
view1_classes = [1, 2, 3]
view2_classes = [4, 5, 6]
#Create lists to hold data and labels for each of the classes across 2 different views
labels = [num for num in range(len(view1_classes)) for _ in range(num_samples)]
labels = np.array(labels)
view1_data = list()
view2_data = list()
#Randomly sample items from each of the selected classes in view1
for class_num in view1_classes:
class_data = all_data[(all_targets == class_num)]
indices = np.random.choice(class_data.shape[0], num_samples)
view1_data.append(class_data[indices])
view1_data = np.concatenate(view1_data)
#Randomly sample items from each of the selected classes in view2
for class_num in view2_classes:
class_data = all_data[(all_targets == class_num)]
indices = np.random.choice(class_data.shape[0], num_samples)
view2_data.append(class_data[indices])
view2_data = np.concatenate(view2_data)
#Shuffle and normalize vectors
shuffled_inds = np.random.permutation(num_samples * len(view1_classes))
view1_data = np.vstack(view1_data)
view2_data = np.vstack(view2_data)
view1_data = view1_data[shuffled_inds]
view2_data = view2_data[shuffled_inds]
magnitudes1 = np.linalg.norm(view1_data, axis=0)
magnitudes2 = np.linalg.norm(view2_data, axis=0)
magnitudes1[magnitudes1 == 0] = 1
magnitudes2[magnitudes2 == 0] = 1
magnitudes1 = magnitudes1.reshape((1, -1))
magnitudes2 = magnitudes2.reshape((1, -1))
view1_data /= magnitudes1
view2_data /= magnitudes2
labels = labels[shuffled_inds]
return [view1_data, view2_data], labels
# ##### Creating a function to perform both single-view and multi-view spectral clustering
#
# In the following function, we will perform single-view spectral clustering on the two views separately and on them concatenated together. We also perform multi-view clustering using the multi-view algorithm. We will also compare the performance of multi-view and single-view versions of spectral clustering. We will evaluate the purity of the resulting clusters from each algorithm with respect to the class labels using the normalized mutual information metric.
def perform_clustering(seed, m_data, labels, n_clusters):
#################Single-view spectral clustering#####################
# Cluster each view separately
s_spectral = SpectralClustering(n_clusters=n_clusters, random_state=RANDOM_SEED, n_init=100)
s_clusters_v1 = s_spectral.fit_predict(m_data[0])
s_clusters_v2 = s_spectral.fit_predict(m_data[1])
# Concatenate the multiple views into a single view
s_data = np.hstack(m_data)
s_clusters = s_spectral.fit_predict(s_data)
# Compute nmi between true class labels and single-view cluster labels
s_nmi_v1 = nmi_score(labels, s_clusters_v1)
s_nmi_v2 = nmi_score(labels, s_clusters_v2)
s_nmi = nmi_score(labels, s_clusters)
print('Single-view View 1 NMI Score: {0:.3f}\n'.format(s_nmi_v1))
print('Single-view View 2 NMI Score: {0:.3f}\n'.format(s_nmi_v2))
print('Single-view Concatenated NMI Score: {0:.3f}\n'.format(s_nmi))
#################Multi-view spectral clustering######################
# Use the MultiviewSpectralClustering instance to cluster the data
m_spectral = MultiviewSpectralClustering(n_clusters=n_clusters, random_state=RANDOM_SEED, n_init=100)
m_clusters = m_spectral.fit_predict(m_data)
# Compute nmi between true class labels and multi-view cluster labels
m_nmi = nmi_score(labels, m_clusters)
print('Multi-view Concatenated NMI Score: {0:.3f}\n'.format(m_nmi))
return m_clusters
# ##### Creating a function to display data and the results of clustering
# The following function plots both views of data given a dataset and corresponding labels.
def display_plots(pre_title, data, labels):
# plot the views
plt.figure()
fig, ax = plt.subplots(1,2, figsize=(14,5))
dot_size=10
ax[0].scatter(new_data[0][:, 0], new_data[0][:, 1],c=labels,s=dot_size)
ax[0].set_title(pre_title + ' View 1')
ax[0].axes.get_xaxis().set_visible(False)
ax[0].axes.get_yaxis().set_visible(False)
ax[1].scatter(new_data[1][:, 0], new_data[1][:, 1],c=labels,s=dot_size)
ax[1].set_title(pre_title + ' View 2')
ax[1].axes.get_xaxis().set_visible(False)
ax[1].axes.get_yaxis().set_visible(False)
plt.show()
# ## Comparing multi-view and single-view spectral clustering on our data set with conditionally independent views
#
# The co-training framework relies on the fundamental assumption that data views are conditionally independent. If all views are informative and conditionally independent, then Multi-view Spectral Clustering is expected to produce higher quality clusters than Single-view Spectral Clustering, for either view or for both views concatenated together. Here, we will evaluate the quality of clusters by using the normalized mutual information metric, which is essentially a measure of the purity of clusters with respect to the true underlying class labels. <br>
#
# As we see below, Multi-view Spectral Clustering produces clusters with lower purity than those produced by Single-view Spectral clustering on the concatenated views, which is surprising.
# +
data, labels = get_ci_data()
m_clusters = perform_clustering(RANDOM_SEED, data, labels, 3)
# Running TSNE to display clustering results via low dimensional embedding
tsne = TSNE()
new_data = list()
new_data.append(tsne.fit_transform(data[0]))
new_data.append(tsne.fit_transform(data[1]))
display_plots('True Labels', new_data, labels)
display_plots('Multi-view Clustering Results', new_data, m_clusters)
# -
# ##### Creating an artificial dataset where the conditional independence assumption between views does not hold
#
# Here, we create an artificial dataset where the conditional independence assumption between
# views, given the true labels, is violated. We again derive our dataset from the forest covertypes
# dataset from sklearn. However, this time, we use only the first 3 classes of the dataset, which will
# correspond to the 3 clusters for view 1. To produce view 2, we will apply a simple nonlinear transformation to view 1
# using the logistic function, and we will apply a negligible amount of noise to the second view to avoid convergence
# issues. This will result in a dataset where the correspondance between views is very high.
def get_cd_data(num_samples=500):
#Load in the vectorized news group data from scikit-learn package
cov = fetch_covtype()
all_data = np.array(cov.data)
all_targets = np.array(cov.target)
#Set class pairings as described in the multiview clustering paper
view1_classes = [1, 2, 3]
view2_classes = [4, 5, 6]
#Create lists to hold data and labels for each of the classes across 2 different views
labels = [num for num in range(len(view1_classes)) for _ in range(num_samples)]
labels = np.array(labels)
view1_data = list()
view2_data = list()
#Randomly sample 500 items from each of the selected classes in view1
for class_num in view1_classes:
class_data = all_data[(all_targets == class_num)]
indices = np.random.choice(class_data.shape[0], num_samples)
view1_data.append(class_data[indices])
view1_data = np.concatenate(view1_data)
#Construct view 2 by applying a nonlinear transformation
#to data from view 1 comprised of a linear transformation
#and a logistic nonlinearity
t_mat = np.random.random((view1_data.shape[1], 50))
noise = 0.005 - 0.01*np.random.random((view1_data.shape[1], 50))
t_mat *= noise
transformed = view1_data @ t_mat
view2_data = scp.special.expit(transformed)
#Shuffle and normalize vectors
shuffled_inds = np.random.permutation(num_samples * len(view1_classes))
view1_data = np.vstack(view1_data)
view2_data = np.vstack(view2_data)
view1_data = view1_data[shuffled_inds]
view2_data = view2_data[shuffled_inds]
magnitudes1 = np.linalg.norm(view1_data, axis=0)
magnitudes2 = np.linalg.norm(view2_data, axis=0)
magnitudes1[magnitudes1 == 0] = 1
magnitudes2[magnitudes2 == 0] = 1
magnitudes1 = magnitudes1.reshape((1, -1))
magnitudes2 = magnitudes2.reshape((1, -1))
view1_data /= magnitudes1
view2_data /= magnitudes2
labels = labels[shuffled_inds]
return [view1_data, view2_data], labels
# ## Comparing multi-view and single-view spectral clustering on our data set with conditionally dependent views
#
# As mentioned before, the co-training framework relies on the fundamental assumption that data views are conditionally independent. Here, we will again compare the performance of single-view and multi-view spectral clustering using the same methods as before, but on our conditionally dependent dataset. <br>
#
# As we see below, Multi-view Spectral Clustering does not beat the best Single-view spectral clustering performance with respect to purity, since that the views are conditionally dependent.
data, labels = get_cd_data()
m_clusters = perform_clustering(RANDOM_SEED, data, labels, 3)
| docs/tutorials/cluster/MVSpectralClustering/MultiviewSpectralValidation_ComplexData.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Nipype Showcase
#
# What's all the hype about Nipype? Is it really that good? Short answer: Yes!
#
# Long answer: ... well, let's consider a very simple fMRI preprocessing workflow that just performs:
# 1. slice time correction
# 2. motion correction
# 3. smoothing
# # Preparing the preprocessing workflow
# First, we need to import the main Nipype tools: `Node` and `Workflow`
from nipype import Node, Workflow
# Now, we can import the interfaces that we want to use for the preprocessing.
from nipype.interfaces.fsl import SliceTimer, MCFLIRT, Smooth
# Next, we will put the three interfaces into a node and define the specific input parameters.
# Initiate a node to correct for slice wise acquisition
slicetimer = Node(SliceTimer(index_dir=False,
interleaved=True,
time_repetition=2.5),
name="slicetimer")
# Initiate a node to correct for motion
mcflirt = Node(MCFLIRT(mean_vol=True,
save_plots=True),
name="mcflirt")
# Initiate a node to smooth functional images
smooth = Node(Smooth(fwhm=4), name="smooth")
# After creating the nodes, we can now create the preprocessing workflow.
preproc01 = Workflow(name='preproc01', base_dir='.')
# Now, we can put all the nodes into this preprocessing workflow. We specify the data flow / execution flow of the workflow by connecting the corresponding nodes to each other.
preproc01.connect([(slicetimer, mcflirt, [('slice_time_corrected_file', 'in_file')]),
(mcflirt, smooth, [('out_file', 'in_file')])])
# To better understand what we did we can write out the workflow graph and visualize it directly in this notebook.
preproc01.write_graph(graph2use='orig')
# Visualize graph
from IPython.display import Image
Image(filename="preproc01/graph_detailed.png")
# # Run the workflow on one functional image
#
# Now, that we've created a workflow, let's run it on a functional image.
#
# For this, we first need to specify the input file of the very first node, i.e. the `slicetimer` node.
slicetimer.inputs.in_file = '/data/ds000114/sub-01/ses-test/func/sub-01_ses-test_task-fingerfootlips_bold.nii.gz'
# To show off Nipype's parallelization power, let's run the workflow in parallel, on 5 processors and let's stop the execution time:
# %time preproc01.run('MultiProc', plugin_args={'n_procs': 5})
# ## Conclusion
#
# Nice, the whole execution took ~2min. But wait... The parallelization didn't really help.
#
# That's true, but because there was no possibility to run the workflow in parallel. Each node depends on the output of the previous node.
# # Results of `preproc01`
#
# So, what did we get? Let's look at the output folder `preproc01`:
# !tree preproc01 -I '*js|*json|*pklz|_report|*.dot|*html'
# # Rerunning of a workflow
# Now, for fun. Let's run the workflow again, but let's change the `fwhm` value of the Gaussian smoothing kernel to `2`.
smooth.inputs.fwhm = 2
# And let's run the workflow again.
# %time preproc01.run('MultiProc', plugin_args={'n_procs': 5})
# ## Conclusion
#
# Interesting, now it only took ~15s to execute the whole workflow again. **What happened?**
#
# As you can see from the log above, Nipype didn't execute the two nodes `slicetimer` and `mclfirt` again. This, because their input values didn't change from the last execution. The `preproc01` workflow therefore only had to rerun the node `smooth`.
# # Running a workflow in parallel
# Ok, ok... Rerunning a workflow again is faster. That's nice and all, but I want more. **You spoke of parallel execution!**
#
# We saw that the `preproc01` workflow takes about ~2min to execute completely. So, if we would run the workflow on five functional images, it should take about ~10min total. This, of course, assuming the execution will be done sequentially. Now, let's see how long it takes if we run it in parallel.
# First, let's copy/clone 'preproc01'
preproc02 = preproc01.clone('preproc02')
preproc03 = preproc01.clone('preproc03')
preproc04 = preproc01.clone('preproc04')
preproc05 = preproc01.clone('preproc05')
# We now have five different preprocessing workflows. If we want to run them in parallel, we can put them all in another workflow.
metaflow = Workflow(name='metaflow', base_dir='.')
# Now we can add the five preproc workflows to the bigger metaflow
metaflow.add_nodes([preproc01, preproc02, preproc03,
preproc04, preproc05])
# **Note:** We now have a workflow (`metaflow`), that contains five other workflows (`preproc0?`), each of them containing three nodes.
#
# To better understand this, let's visualize this `metaflow`.
# As before, let's write the graph of the workflow
metaflow.write_graph(graph2use='flat')
# And visualize the graph
from IPython.display import Image
Image(filename="metaflow/graph_detailed.png")
# Ah... so now we can see that the `metaflow` has potential for parallelization. So let's put it to test
# %time metaflow.run('MultiProc', plugin_args={'n_procs': 5})
# This time we can see that Nipype uses all available processors.
#
# And if all went well, the total execution time should still be around ~2min.
#
# That's why Nipype is so amazing. The days of opening multiple SPMs, FSLs, AFNIs etc. are past!
# # Results of `metaflow`
# !tree metaflow -I '*js|*json|*pklz|_report|*.dot|*html'
| notebooks/introduction_showcase.ipynb |
# ---
# title: "Hyperparameter tunning for trees with GridSearch for Tree"
# date: 2020-04-12T14:41:32+02:00
# author: "<NAME>"
# type: technical_note
# draft: false
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# - Problem: search for a set of optimal hyperparameters for a learning algorithm.
# - Solution: find a set of optimal hyperparameters that results in an optimal model.
# - Optimal model: yields an optimal score.
# - Score: in sklearn defaults to accuracy (classication) and $R^2$ (regression).
# - Cross validation is used to estimate the generalization performance.
# ### Import modules
# +
# Manipulation
import numpy as np
import pandas as pd
# Visualization
import seaborn as sns
import matplotlib.pyplot as plt
# Selection
from sklearn.model_selection import train_test_split
from sklearn.model_selection import GridSearchCV
# Metrics
from sklearn.metrics import mean_squared_error as MSE
from sklearn.metrics import accuracy_score
from sklearn.metrics import roc_auc_score
# Models
from sklearn.tree import DecisionTreeClassifier
# -
# ### Load data
# +
# Set seed for reproducibility
SEED=1
# read csv into df
df = pd.read_csv('liver/indian_liver_patient_preprocessed.csv')
# Get features of interest and target labels
X = df.iloc[:,:-1]
y = (df.iloc[:,-1]).astype(int)
# Split dataset into 80% train, 20% test
X_train, X_test, y_train, y_test= train_test_split(X, y,
test_size=0.2,
stratify=y,
random_state=SEED)
# -
# ### Evaluate performance of an untuned dt
dt = DecisionTreeClassifier(random_state=SEED)
dt.fit(X_train, y_train)
y_pred_proba = dt.predict_proba(X_test)[:,1]
test_roc_auc = roc_auc_score(y_test, y_pred_proba)
print('Test set ROC AUC score: {:.3f}'.format(test_roc_auc))
# ### Get all hyperparameters of dt
# Instantiate dt
dt = DecisionTreeClassifier(random_state=SEED)
print(dt.get_params())
# ### Define search grid
# Define params_dt
params_dt = {'max_depth': [2,3,4],
'min_samples_leaf': [0.12,0.14,0.16,0.18]
}
# ### Search for optimical tree
# `GridSearchCV` with the `estimator` option becomes the new classifer, as if I have instantiated the estimator with the correct set of hyperparameters.
# +
# Instantiate grid_dt
grid_dt = GridSearchCV(estimator=dt,
param_grid=params_dt,
scoring='roc_auc',
cv=5,
n_jobs=-1)
# Fit 'grid_dt' to the training data
grid_dt.fit(X_train, y_train)
# Extract best hyperparameters from 'grid_dt'
best_hyperparams = grid_dt.best_params_
print('Best hyerparameters:\n', best_hyperparams)
# Extract best CV score from 'grid_dt'
best_CV_score = grid_dt.best_score_
print('Best CV accuracy: {:.3f}'.format(best_CV_score))
# Extract the best estimator
best_model = grid_dt.best_estimator_
# Predict the test set probabilities of the positive class
y_pred_proba = grid_dt.predict_proba(X_test)[:,1]
# Compute test_roc_auc
test_roc_auc = roc_auc_score(y_test, y_pred_proba)
# Print test_roc_auc
print('Test set ROC AUC score: {:.3f}'.format(test_roc_auc))
# -
# ### Evaluate the optimical tree
dt = DecisionTreeClassifier(max_depth=3, min_samples_leaf=0.12, random_state=SEED)
dt.fit(X_train, y_train)
y_pred_proba = dt.predict_proba(X_test)[:,1]
test_roc_auc = roc_auc_score(y_test, y_pred_proba)
print('Test set ROC AUC score: {:.3f}'.format(test_roc_auc))
# Good improvement upon an untuned classification-tree would achieve a ROC AUC score of 0.54
| courses/datacamp/notes/python/sklearn/tree_hyperparameters.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/KevinTheRainmaker/Recommendation_Algorithms/blob/main/Internship_Model_01.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="PQQbDaiX7Ymg" outputId="12ef23ce-b4b0-4fc3-f186-bc995e81c4d8"
# !pip install -q implicit
# + [markdown] id="o9e02rsw-2F8"
# ## Packages
# + id="iZ66JezW-1k4"
import os
import pandas as pd
import tqdm.notebook as tqdm
from collections import Counter
from google.colab import drive
import scipy.sparse as sps
import implicit
# + [markdown] id="FZjyCi6K78l8"
# ## Data Loader
# + colab={"base_uri": "https://localhost:8080/"} id="tiA8IPEZ7wTF" outputId="d0950114-192b-44ba-c93f-4941f39da129"
drive.mount('/content/drive')
path = '/content/drive/MyDrive/data'
# + id="nws0olIH_S_d"
sps_mat_1 = pd.read_csv(os.path.join(path, 'implicit_trial.csv'), encoding='utf-8')
sps_mat_1.fillna(0, inplace=True)
sps_mat_2 = pd.read_csv(os.path.join(path, 'implicit_trial.csv'), encoding='utf-8')
sps_mat_2.fillna(0, inplace=True)
sps_mat_3 = pd.read_csv(os.path.join(path, 'implicit_trial.csv'), encoding='utf-8')
sps_mat_3.fillna(0, inplace=True)
sps_mat_4 = pd.read_csv(os.path.join(path, 'implicit_trial.csv'), encoding='utf-8')
sps_mat_4.fillna(0, inplace=True)
# + [markdown] id="G7PsMxEG9cp6"
# ## Processor
# + id="efjobhA-9QFY" colab={"base_uri": "https://localhost:8080/"} outputId="6ad6b14b-120e-4d37-aee4-688e0a3b83a6"
sps_mat_pro_1 = sps.csr_matrix(sps_mat_1)
sps_mat_pro_2 = sps.csr_matrix(sps_mat_2)
sps_mat_pro_3 = sps.csr_matrix(sps_mat_3)
sps_mat_pro_4 = sps.csr_matrix(sps_mat_4)
# + [markdown] id="f91G5pjX9u-z"
# ## Model
# + id="KAbg1Mt--Pm_"
# configuration
n_factors = [x for x in range(50,350,50)]
lrs = [1/pow(10,x) for x in range(1,4)]
# model name
collector = [sps_mat_pro_1, sps_mat_pro_2, sps_mat_pro_3, sps_mat_pro_4]
# + colab={"base_uri": "https://localhost:8080/", "height": 145, "referenced_widgets": ["<KEY>", "<KEY>", "<KEY>", "adb9dea0bcc64ad49f84027fb2c8061e", "c09e06a4e95d41de99747e9876870558", "1f556db1c64c4a868183fd8ede850eda", "5ecb69e170ad4946a2c00a7937556562", "<KEY>", "6d24e69d68a74420be5c7c3931faa03b", "b84f0f1a3c6e4d1faaf6e1d8cecc1060", "<KEY>", "<KEY>", "<KEY>", "8c2882a2e0e845d695cc919e168eb255", "a8f30f35c8b848a99ffaf74c55ca22c3", "<KEY>", "55feb5f72849496e93fb41e6add5e2dc", "91590004314649d891662dedc517b690", "<KEY>", "615e5d66b224417ea8d2c5de42e61ea5", "<KEY>", "7483314e0da34fedbb50973fa815f732", "03e582353e2245c3ae825465f1b61233", "435173fe5d3a46c1ba374af4b28e4577", "45d68036a8f8483d8ce4be03ecc42867", "bde4fbb660ea4b65899ecbb4e41fa331", "faabc93ff56a4c9bbe8e5567c56ce6c6", "d546a52394d14b859ba1d25c9623ac7a", "3c49e4e4a3e844e7a498ee6ded3f437d", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "40266e93ddae4c6893f35f668093aaed", "8027d9a30bd34057992350aca36ba851", "262c04af2780499c8b31b0ae6e8a8d0b", "ddb028fa40a34c42901164e02085815f", "f310b9ac7b4345f0a924ced227fcee00", "<KEY>", "95a72d8904704f1bbcccc41e6d8cf19f", "<KEY>", "2d2da20df80342248f7f2af2977ec2e8"]} id="4KwgCjJV9qyA" outputId="98cabf3c-b206-4fec-868a-171067994d90"
for data in collector:
name = f'model[{data}]'
name = implicit.cpu.bpr.BayesianPersonalizedRanking(factors=100, learning_rate=0.01)
name.fit(item_users=data, show_progress=True)
# + [markdown] id="JFo7k_HgfgbV"
# ## Recommend
# + id="C3FqB7zD-KqH"
# sps_mat_1['movieId']
movie_ids = [int(x) for x in sps_mat_1['movieId']]
user_ids = [int(x) for x in sps_mat_1.columns[1:]]
# + colab={"base_uri": "https://localhost:8080/"} id="YboK3kqDf2We" outputId="90646632-4f03-44c2-e656-df99330b33db"
for user_id in user_ids:
user_items = sps_mat_processed.tocsr()
temp = Counter(dict(model.recommend(user_id,user_items, N=15)))
temp
# + colab={"base_uri": "https://localhost:8080/"} id="L6ZUcnXYsvNP" outputId="6fe57002-cc11-4c53-f192-5e81033984db"
(result_1+result_2).columns
# + colab={"base_uri": "https://localhost:8080/", "height": 677} id="8-MsKk0olB8S" outputId="c1b89391-9979-4a66-c57f-20ea62daa4fa"
(result_1+result_2).sort_values(by=[('user_610', 'score')], axis=0)
# + id="JHHHsZqQpoJZ"
| Internship_Model_01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Problem Set 2 - Arctic ice maps
# The purpose of this problem set is to become familiar with working with image data, plotting it, and combining it in various ways for analysis.
#
# The data used in this problem set was collected by the AMSR-E instrument [Aqua](http://en.wikipedia.org/wiki/Aqua_%28satellite%29) satellite. The data consists of maps of the concentration of ice in the Arctic collected between 2006 and 2011. I have downloaded and extracted the maps from [here](http://www.iup.uni-bremen.de/seaice/amsr/) and have put them in a format that you can more easily used (if you are interested in using the raw data directly, let me know and I can show you).
#
# The data you should is in the [data/ice_data.tgz](data/ice_data.tgz) file (on Windows you can use the [data/ice_data.zip](data/ice_data.zip) file). This is actually a subset of the data, with only two ice maps every month.
#
# The data is in 'Numpy' format, which means that you can read it as a Numpy array using:
#
# >>> import numpy as np
# >>> data = np.load('ice_data/20080415.npy')
#
# which will give you a 2-d array. Just for information, this was created with:
#
# >>> np.save('ice_data/20080415.npy', data)
# ## Part 1 - examining a single map (6 points)
# **Start off by reading in one of the maps as shown above, and plot it with Matplotlib**. Note that to get the correct orientation, you will need to call the ``imshow`` command with the ``origin='lower'`` option, which ensures that the (0,0) pixels is on the bottom left, not the top left. You can try and use different colormaps if you like (set by the ``cmap`` option) - see [here](http://matplotlib.org/examples/color/colormaps_reference.html) for information on the available colormaps. You can specify a colormap to use with e.g. ``cmap=plt.cm.jet`` (i.e. ``cmap=plt.cm.`` followed by the name of the colormap). Note that you can make figures larger by specifying e.g.
#
# >>> plt.figure(figsize=(8,8))
#
# where the size is given in inches. Try and find a way to plot a colorbar on the side, to show what color corresponds to what value. Remember that you can always look at the examples in the [Matplotlib Gallery](http://matplotlib.org/gallery.html) to find examples. You can also try and remove the tick labels (``100``, ``200``, etc.) since they are not useful - but don't worry if you can't figure out how.
# ## Part 2 - reading in multiple maps (10 points)
# We now want to make a plot of the ice concentration over time. Reading in a single map is easy, but since we have 137 maps, we do not want to read them all in individually by hand. **Write a loop over all the available files, and inside the loop, read in the data to a variable (e.g. ``data``), and also extract the year, month, and day as integer values (e.g. ``year``, ``month``, and ``day``)**. Then, also inside the loop, **compute a variable ``time``** which is essentially the fractional time in years (so 1st July 2012 is 2012.5). You can assume for simplicity that each month has 30 days - this will not affect the results later. Finally, also **compute for each file the total number of pixels that have a value above 50%**. After the loop, **make a plot of the number of pixels with a concentration above 50% against time**.
#
# You will likely notice that the ticks are in a strange format, where they are given in years since 2006, but you can change this with the following code:
#
# >>> from matplotlib.ticker import ScalarFormatter
# >>> plt.gca().xaxis.set_major_formatter(ScalarFormatter(useOffset=False))
#
# **Describe what you see in the plot**.
#
# We now want something a little more quantitative than just the number of pixels, so we will try and compute the area where the ice concentration is above a given threshold. However, we first need to know the area of the pixels in the image, and since we are looking at a projection of a spherical surface, each pixel will be a different area. The areas (in km^2) are contained inside the file named ``ice_data_area.npy``. **Read in the areas and make a plot (with colorbar) to see how the pixel area is changing over the image.**
#
# Now, loop over the files again as before, but this time, for each file, **compute the total area where the concentration of ice is 99% or above. Make a new plot showing the area of >99% ice concentration against time.**
#
# **Describe what you see - how does the minimum change over time?**
# ## Part 3 - visualizing changes over time (10 points)
# **Find the date at which the area of the region where the ice concentration is above 99% is the smallest**. What is the value of the minimum area?
#
# Next, **read in the map for this minimum, and the map for the same day and month but from 2006**. **Make a side-by-side plot showing the 2006 and the 2011 data**.
#
# **Compute the difference between the two maps** so that a loss in ice over time will correspond to a negative value, and a gain in ice will correspond to a positive value. **Make a plot** of the difference, and use the ``RdBu`` colormap to highlight the changes (include a colorbar).
# ## Part 4 - yearly averages (4 points)
# **Compute average ice concentration maps for 2006 and 2011, and plot them side by side.**
# ## Epilogue
# The data that we have here only cover five years, so we cannot reliably extract information about long term trends. However, it is worth noting that the minimum ice coverage you found here was a record minimum - never before (in recorded history) had the size of the ice shelf been so small. This is part of a long term trend due to global warming. In 2012, the record was again beaten, and most scientists believe that by ~2050, the Arctic will be completely ice-free for at least part of the summer.
| Python4Scientitsts/3 wed.pdf/Problem Set 2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.12 64-bit (''base'': conda)'
# language: python
# name: python3
# ---
# # [01/09/21] Weight Distributions
import sys
sys.path = ['/home/mansheej/open_lth'] + sys.path
from foundations import hparams, step
import matplotlib.pyplot as plt
import models.registry
import numpy as np
import pandas as pd
from pathlib import Path
from pruning.mask import Mask
import seaborn as sns
import torch
plt.style.use('default')
sns.set_theme(
style='ticks',
font_scale=1.2,
rc={
'axes.linewidth': '0.8',
'axes.grid': True,
'figure.constrained_layout.use': True,
'grid.linewidth': '0.8',
'legend.edgecolor': '1.0',
'legend.fontsize': 'small',
'legend.title_fontsize': 'small',
'xtick.major.width': '0.8',
'ytick.major.width': '0.8'
},
)
# +
exp_meta_paths = [
Path(f'/home/mansheej/open_lth_data/lottery_78a119e24960764e0de0964887d2597f/'), # 50k x 2ep @ 0.4
Path(f'/home/mansheej/open_lth_data/lottery_acab023c0d3d306bb3ac850a9a4eeaf0/'), # 50k x 1ep @ 0.2
# Path(f'/home/mansheej/open_lth_data/lottery_b279562b990bac9b852b17b287fca1ef/'), # 50k x 1 @ 0.4
# Path(f'/home/mansheej/open_lth_data/lottery_148749c4bcfd56a8458e542c11095ca7/'), # 128 R x 400it @ 0.4
Path(f'/home/mansheej/open_lth_data/lottery_867a92df0db62b580aa466941524b0af/'), # 256 R x 400it @ 0.4
# Path(f'/home/mansheej/open_lth_data/lottery_32864827ed21bf58b724036a033774bf/'), # 512 R x 400it @ 0.4
# Path(f'/home/mansheej/open_lth_data/lottery_a3f64ad3b4f6746c59fbc42e3e849d9f/'), # 1024 R x 400it @ 0.4
Path(f'/home/mansheej/open_lth_data/lottery_8d9ced57b5dd8c36aefe60d19f5a48d4/'), # 2048 R x 400it @ 0.4
]
exp_paths = [[emp / f'replicate_{i}' for i in range(1, 5)] for emp in exp_meta_paths]
plt.figure(figsize=(8.4, 4.8))
ls = []
for i, eps in enumerate(exp_paths):
num_levels = 15
acc_run_level = []
for p in eps:
acc_level = []
for l in range(num_levels + 1):
df = pd.read_csv(p / f'level_{l}/main/logger', header=None)
acc_level.append(df[2].iloc[-2])
acc_run_level.append(acc_level)
acc_run_level = np.array(acc_run_level)
x = np.arange(16)
ys = acc_run_level
y_mean, y_std = ys.mean(0), ys.std(0)
c = f'C{i}'
l = plt.plot(x, y_mean, c=c, alpha=0.8, linewidth=2)
ls.append(l[0])
plt.fill_between(x, y_mean + y_std, y_mean - y_std, color=c, alpha=0.2)
plt.legend(
ls,
[
'2 Passes, All 50000 Examples, 782 Steps',
'1 Pass, All 50000 Examples, 391 Steps',
# '400 Passes, 128 Random Exampes, 400 Steps',
'200 Passes, 256 Random Exampes, 400 Steps',
# '100 Passes, 512 Random Exampes, 400 Steps',
'25 Passes, 2048 Random Exampes, 400 Steps',
],
)
plt.xlim(0, 15)
plt.ylim(0.845, 0.925)
plt.xticks(np.arange(0, 16, 2), [f'{f*100:.1f}' for f in 0.8**np.arange(0, 16, 2)])
plt.xlabel('% Weights Remaining')
plt.ylabel('Test Accuracy')
plt.title('CIFAR10 ResNet20: Pretrain 400 Steps with Random Data')
sns.despine()
# plt.savefig('/home/mansheej/open_lth/figs/0023.svg')
plt.show()
# -
data_sizes = [256, 2048]
exp_hashes = ['lottery_867a92df0db62b580aa466941524b0af', 'lottery_8d9ced57b5dd8c36aefe60d19f5a48d4']
levels = 15
replicates = [1, 2, 3, 4]
iteration = 400
batch_size = 128
model_name='cifar_resnet_20'
model_init='kaiming_normal'
batchnorm_init='uniform'
outputs = 10
weights = {}
for data_size, exp_hash in zip(data_sizes, exp_hashes):
weights[data_size] = {}
for replicate in replicates:
# pretrained model
path = f'/home/mansheej/open_lth_data/{exp_hash}/replicate_{replicate}/level_pretrain/main'
model_hparams = hparams.ModelHparams(model_name, model_init, batchnorm_init)
model = models.registry.load(
save_location=path,
save_step=step.Step(iteration, data_size // batch_size),
model_hparams=model_hparams,
outputs=10
)
with torch.no_grad():
prunable_params = []
for name, params in model.named_parameters():
if name in model.prunable_layer_names:
prunable_params.append(params.flatten())
prunable_params = torch.cat(prunable_params).numpy()
weights[data_size][replicate] = prunable_params
data_size = 50000
weights[data_size] = {}
for replicate in replicates:
path = f'/home/mansheej/open_lth_data/lottery_acab023c0d3d306bb3ac850a9a4eeaf0/replicate_{replicate}/level_pretrain/main'
model_hparams = hparams.ModelHparams(model_name, model_init, batchnorm_init)
model = models.registry.load(
save_location=path,
save_step=step.Step(391, 391),
model_hparams=model_hparams,
outputs=10
)
with torch.no_grad():
prunable_params = []
for name, params in model.named_parameters():
if name in model.prunable_layer_names:
prunable_params.append(params.flatten())
prunable_params = torch.cat(prunable_params).numpy()
weights[data_size][replicate] = prunable_params
masks = {}
for data_size, exp_hash in zip(data_sizes, exp_hashes):
masks[data_size] = {}
for replicate in replicates:
masks[data_size][replicate] = {}
for level in range(levels + 1):
path = f'/home/mansheej/open_lth_data/{exp_hash}/replicate_{replicate}/level_{level}/main'
mask = Mask.load(path)
mask = torch.cat([mask[name].flatten() for name in model.prunable_layer_names]).numpy().astype(bool)
masks[data_size][replicate][level] = mask
data_size = 50000
masks[data_size] = {}
for replicate in replicates:
masks[data_size][replicate] = {}
for level in range(levels + 1):
path = f'/home/mansheej/open_lth_data/lottery_acab023c0d3d306bb3ac850a9a4eeaf0/replicate_{replicate}/level_{level}/main'
mask = Mask.load(path)
mask = torch.cat([mask[name].flatten() for name in model.prunable_layer_names]).numpy().astype(bool)
masks[data_size][replicate][level] = mask
agg_ws = {k: np.abs(np.concatenate([v[r] for r in replicates])) for k, v in weights.items()}
agg_ms = {k: [np.abs(np.concatenate([v[r][l] for r in replicates])) for l in range(levels + 1)] for k, v in masks.items()}
# +
fig, axes = plt.subplots(4, 4, sharex=True, figsize=(12.8, 9.6))
for level in range(levels + 1):
i, j = level//4, level%4
ax = axes[i, j]
# ax.hist(agg_ws[256][agg_ms[256][level]], bins=100, range=(0,2.5), density=True, histtype='step', lw=1.5, label='data size 256')
# ax.hist(agg_ws[2048][agg_ms[2048][level]], bins=100, range=(0,2.5), density=True, histtype='step', lw=1.5, alpha=0.8, label='data size 2048')
ax.hist(agg_ws[256][agg_ms[256][level]], bins=100, range=(0,2.5), histtype='step', lw=1.5, label='data size 256')
ax.hist(agg_ws[2048][agg_ms[2048][level]], bins=100, range=(0,2.5), histtype='step', lw=1.5, alpha=0.8, label='data size 2048')
ax.hist(agg_ws[50000][agg_ms[50000][level]], bins=100, range=(0,2.5), histtype='step', lw=1.5, alpha=0.8, label='data size 50000')
ax.legend()
# ax.set_xlim(1.5, 2.5)
ax.set_ylim(0, 2000)
ax.set_title(f'Level {level}', fontsize='small')
sns.despine()
# fig.savefig('w_dist_2.svg')
# -
| nbs/22_01_09__Weight_Distributions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
# ## Facies classification using Random forest and engineered features
#
#
# #### Contest entry by: <a href="https://github.com/mycarta"><NAME></a>, <a href="https://github.com/dahlmb"><NAME></a>, with a contribution by <NAME>.
#
# #### [Original contest notebook](https://github.com/seg/2016-ml-contest/blob/master/Facies_classification.ipynb) by <NAME>, [Enthought](https://www.enthought.com/)
#
#
# <a rel="license" href="http://creativecommons.org/licenses/by/4.0/"><img alt="Creative Commons License" style="border-width:0" src="https://i.creativecommons.org/l/by/4.0/88x31.png" /></a><br /><span xmlns:dct="http://purl.org/dc/terms/" property="dct:title">The code and ideas in this notebook,</span> by <span xmlns:cc="http://creativecommons.org/ns#" property="cc:attributionName"><NAME> and <NAME>, </span> are licensed under a <a rel="license" href="http://creativecommons.org/licenses/by/4.0/">Creative Commons Attribution 4.0 International License</a>.
# ### Loading the dataset with selected set of top 70 engineered features.
#
# - We first created a large set of moments and GLCM features. The workflow is described in the 03_Facies_classification_MandMs_feature_engineering_commented.ipynb notebook (with huge thanks go to <NAME> for his critically needed Pandas magic, and useful suggestions).
# - We then selected 70 using a Sequential (Forward) Feature Selector form <NAME>'s [mlxtend](http://rasbt.github.io/mlxtend/user_guide/feature_selection/SequentialFeatureSelector/) library. Details in the 03_Facies_classification-MandMs_SFS_feature_selection.ipynb notebook.
#
# +
# %matplotlib inline
import numpy as np
import scipy as sp
from scipy.stats import randint as sp_randint
from scipy.signal import argrelextrema
import matplotlib as mpl
import matplotlib.pyplot as plt
import pandas as pd
from sklearn import preprocessing
from sklearn.metrics import f1_score, make_scorer
from sklearn.model_selection import LeaveOneGroupOut, validation_curve
# -
filename = 'SFS_top70_selected_engineered_features.csv'
training_data = pd.read_csv(filename)
training_data.describe()
training_data['Well Name'] = training_data['Well Name'].astype('category')
training_data['Formation'] = training_data['Formation'].astype('category')
training_data['Well Name'].unique()
# Now we extract just the feature variables we need to perform the classification. The predictor variables are the five log values and two geologic constraining variables, **and we are also using depth**. We also get a vector of the facies labels that correspond to each feature vector.
y = training_data['Facies'].values
print y[25:40]
print np.shape(y)
X = training_data.drop(['Formation', 'Well Name','Facies'], axis=1)
print np.shape(X)
X.describe(percentiles=[.05, .25, .50, .75, .95])
# ### Preprocessing data with standard scaler
scaler = preprocessing.StandardScaler().fit(X)
X = scaler.transform(X)
# ### Make F1 performance scorers
Fscorer = make_scorer(f1_score, average = 'micro')
# ### Parameter tuning ( maximum number of features and number of estimators): validation curves combined with leave one well out cross validation
wells = training_data["Well Name"].values
logo = LeaveOneGroupOut()
# ### Random forest classifier
#
# In Random Forest classifiers serveral decision trees (often hundreds - a forest of trees) are created and trained on a random subsets of samples (drawn with replacement) and features (drawn without replacement); the decision trees work together to make a more accurate classification (description from <NAME>'s <a href="http://nbviewer.jupyter.org/github/rhiever/Data-Analysis-and-Machine-Learning-Projects/blob/master/example-data-science-notebook/Example%20Machine%20Learning%20Notebook.ipynb"> excellent notebook</a>).
#
# +
from sklearn.ensemble import RandomForestClassifier
RF_clf100 = RandomForestClassifier (n_estimators=100, n_jobs=-1, random_state = 49)
RF_clf200 = RandomForestClassifier (n_estimators=200, n_jobs=-1, random_state = 49)
RF_clf300 = RandomForestClassifier (n_estimators=300, n_jobs=-1, random_state = 49)
RF_clf400 = RandomForestClassifier (n_estimators=400, n_jobs=-1, random_state = 49)
RF_clf500 = RandomForestClassifier (n_estimators=500, n_jobs=-1, random_state = 49)
RF_clf600 = RandomForestClassifier (n_estimators=600, n_jobs=-1, random_state = 49)
param_name = "max_features"
#param_range = [5, 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60]
param_range = [9, 12, 15, 18, 21, 24, 27, 30, 33, 36, 39, 42, 45, 48, 51]
plt.figure()
plt.suptitle('n_estimators = 100', fontsize=14, fontweight='bold')
_, test_scores = validation_curve(RF_clf100, X, y, cv=logo.split(X, y, groups=wells),
param_name=param_name, param_range=param_range,
scoring=Fscorer, n_jobs=-1)
test_scores_mean = np.mean(test_scores, axis=1)
plt.plot(param_range, test_scores_mean)
plt.xlabel(param_name)
plt.xlim(min(param_range), max(param_range))
plt.ylabel("F1")
plt.ylim(0.47, 0.57)
plt.show()
#print max(test_scores_mean[argrelextrema(test_scores_mean, np.greater)])
print np.amax(test_scores_mean)
print np.array(param_range)[test_scores_mean.argmax(axis=0)]
plt.figure()
plt.suptitle('n_estimators = 200', fontsize=14, fontweight='bold')
_, test_scores = validation_curve(RF_clf200, X, y, cv=logo.split(X, y, groups=wells),
param_name=param_name, param_range=param_range,
scoring=Fscorer, n_jobs=-1)
test_scores_mean = np.mean(test_scores, axis=1)
plt.plot(param_range, test_scores_mean)
plt.xlabel(param_name)
plt.xlim(min(param_range), max(param_range))
plt.ylabel("F1")
plt.ylim(0.47, 0.57)
plt.show()
#print max(test_scores_mean[argrelextrema(test_scores_mean, np.greater)])
print np.amax(test_scores_mean)
print np.array(param_range)[test_scores_mean.argmax(axis=0)]
plt.figure()
plt.suptitle('n_estimators = 300', fontsize=14, fontweight='bold')
_, test_scores = validation_curve(RF_clf300, X, y, cv=logo.split(X, y, groups=wells),
param_name=param_name, param_range=param_range,
scoring=Fscorer, n_jobs=-1)
test_scores_mean = np.mean(test_scores, axis=1)
plt.plot(param_range, test_scores_mean)
plt.xlabel(param_name)
plt.xlim(min(param_range), max(param_range))
plt.ylabel("F1")
plt.ylim(0.47, 0.57)
plt.show()
#print max(test_scores_mean[argrelextrema(test_scores_mean, np.greater)])
print np.amax(test_scores_mean)
print np.array(param_range)[test_scores_mean.argmax(axis=0)]
plt.figure()
plt.suptitle('n_estimators = 400', fontsize=14, fontweight='bold')
_, test_scores = validation_curve(RF_clf400, X, y, cv=logo.split(X, y, groups=wells),
param_name=param_name, param_range=param_range,
scoring=Fscorer, n_jobs=-1)
test_scores_mean = np.mean(test_scores, axis=1)
plt.plot(param_range, test_scores_mean)
plt.xlabel(param_name)
plt.xlim(min(param_range), max(param_range))
plt.ylabel("F1")
plt.ylim(0.47, 0.57)
plt.show()
#print max(test_scores_mean[argrelextrema(test_scores_mean, np.greater)])
print np.amax(test_scores_mean)
print np.array(param_range)[test_scores_mean.argmax(axis=0)]
plt.figure()
plt.suptitle('n_estimators = 500', fontsize=14, fontweight='bold')
_, test_scores = validation_curve(RF_clf500, X, y, cv=logo.split(X, y, groups=wells),
param_name=param_name, param_range=param_range,
scoring=Fscorer, n_jobs=-1)
test_scores_mean = np.mean(test_scores, axis=1)
plt.plot(param_range, test_scores_mean)
plt.xlabel(param_name)
plt.xlim(min(param_range), max(param_range))
plt.ylabel("F1")
plt.ylim(0.47, 0.57)
plt.show()
#print max(test_scores_mean[argrelextrema(test_scores_mean, np.greater)])
print np.amax(test_scores_mean)
print np.array(param_range)[test_scores_mean.argmax(axis=0)]
plt.figure()
plt.suptitle('n_estimators = 600', fontsize=14, fontweight='bold')
_, test_scores = validation_curve(RF_clf600, X, y, cv=logo.split(X, y, groups=wells),
param_name=param_name, param_range=param_range,
scoring=Fscorer, n_jobs=-1)
test_scores_mean = np.mean(test_scores, axis=1)
plt.plot(param_range, test_scores_mean)
plt.xlabel(param_name)
plt.xlim(min(param_range), max(param_range))
plt.ylabel("F1")
plt.ylim(0.47, 0.57)
plt.show()
#print max(test_scores_mean[argrelextrema(test_scores_mean, np.greater)])
print np.amax(test_scores_mean)
print np.array(param_range)[test_scores_mean.argmax(axis=0)]
# -
# ### Average test F1 score with leave one well out
# +
RF_clf_f1 = RandomForestClassifier (n_estimators=600, max_features = 21,
n_jobs=-1, random_state = 49)
f1_RF = []
for train, test in logo.split(X, y, groups=wells):
well_name = wells[test[0]]
RF_clf_f1.fit(X[train], y[train])
pred = RF_clf_f1.predict(X[test])
sc = f1_score(y[test], pred, labels = np.arange(10), average = 'micro')
print("{:>20s} {:.3f}".format(well_name, sc))
f1_RF.append(sc)
print "-Average leave-one-well-out F1 Score: %6f" % (sum(f1_RF)/(1.0*(len(f1_RF))))
# -
# ### Predicting and saving facies for blind wells
RF_clf_b = RandomForestClassifier (n_estimators=600, max_features = 21,
n_jobs=-1, random_state = 49)
blind = pd.read_csv('engineered_features_validation_set_top70.csv')
X_blind = np.array(blind.drop(['Formation', 'Well Name'], axis=1))
scaler1 = preprocessing.StandardScaler().fit(X_blind)
X_blind = scaler1.transform(X_blind)
y_pred = RF_clf_b.fit(X, y).predict(X_blind)
#blind['Facies'] = y_pred
np.save('ypred_RF_SFS_VC.npy', y_pred)
| MandMs/03_Facies_classification-MandMs_RandomForest_EngineeredFeatures_SFSelection_ValidationCurves.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + editable=false jupyter={"outputs_hidden": false}
from sklearn import datasets
from sklearn.naive_bayes import GaussianNB, MultinomialNB
from sklearn.model_selection import train_test_split
import numpy as np
import pandas as pd
data = pd.read_csv('wine_original.csv')
labels = data['class']
del data['class']
X_train, X_test, y_train, y_test = train_test_split(data, labels, test_size=0.2, random_state=5)
# + editable=false jupyter={"outputs_hidden": false}
X_train
# + editable=false jupyter={"outputs_hidden": false}
gnb = GaussianNB()
gnb.fit(X_train, y_train)
y_pred = gnb.predict(X_test)
y_train_pred = gnb.predict(X_train)
print('Training accuracy = ' + str(np.sum(y_train_pred == y_train) / len(y_train)))
print('Test accuracy = ' + str(np.sum(y_pred == y_test) / len(y_test)))
# + editable=false jupyter={"outputs_hidden": false}
X_train, X_valid, y_train, y_valid = train_test_split(data, labels, test_size=0.2, random_state=5)
alphas = [0.1, 0.5, 1, 2, 3, 4, 5, 10, 100]
best_alpha = 0.1
best_acc = 0.0
for alpha in alphas:
clf = MultinomialNB(alpha=alpha)
clf.fit(X_train, y_train)
y_pred = clf.predict(X_valid)
accuracy = np.sum(y_pred == y_valid) / len(y_valid)
print('Validation accuracy = ' + str(accuracy) + ' at alpha = ' + str(alpha))
if accuracy > best_acc:
best_acc = accuracy
best_alpha = alpha
print('Best alpha = ' + str(best_alpha))
# + editable=false jupyter={"outputs_hidden": false}
X_train = np.concatenate((X_train, X_valid))
y_train = np.concatenate((y_train, y_valid))
clf = MultinomialNB(alpha=best_alpha)
clf.fit(X_train, y_train)
y_pred = clf.predict(X_test)
y_train_pred = clf.predict(X_train)
print('Training accuracy = ' + str(np.sum(y_train_pred == y_train) / len(y_train)))
print('Test accuracy = ' + str(np.sum(y_pred == y_test) / len(y_test)))
# + editable=false jupyter={"outputs_hidden": false}
# + editable=false jupyter={"outputs_hidden": false}
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
X_train, X_test, y_train, y_test = train_test_split(data, labels, test_size=0.2, random_state=5)
clf = LinearDiscriminantAnalysis()
clf.fit(X_train, y_train)
y_pred = clf.predict(X_test)
y_train_pred = clf.predict(X_train)
print('Training accuracy = ' + str(np.sum(y_train_pred == y_train) / len(y_train)))
print('Test accuracy = ' + str(np.sum(y_pred == y_test) / len(y_test)))
# + editable=false jupyter={"outputs_hidden": false}
# + editable=false jupyter={"outputs_hidden": false}
| jupyter_experiment/test/Task3/task3_notebook_B.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.6 64-bit
# language: python
# name: python37664bit910bb1eb151e492faa58b1db9f13ff98
# ---
# ### Alternating Characters
# You are given a string containing characters A and B only. Your task is to change it into a string such that there are no matching adjacent characters. To do this, you are allowed to delete zero or more characters in the string.
# Your task is to find the minimum number of required deletions.
# For example, given the string s = AABAAB, remove an A at position ) 0 and 3 to make s = ABAB in 2 deletions.
#
# #### Function Description
# Complete the alternating Characters function in the editor below. It must return an integer representing the minimum number of deletions to make the alternating string.
# alternatingCharacters has the following parameters(s):
# * s: a string
#
# #### Input Format
# The first line contains an integer 1, the number of queries.
# The next q lines each contain a string s
#
# #### Constraints
# * 1<= q <= 10
# * 1 <=|s|<=10^5
string = "AABAAB"
# +
def alternatingCharacters(s):
deletions = 0
for i in range(len(string)-1):
if string[i] == string[i+1]:
deletions+=1
return deletions
print(alternatingCharacters(string))
# -
| Python/Alternating Characters/Alternating Characters.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Product Matching based on text attributes
# * Author - <NAME>
# * Date - January 16, 2019
# ## Table of Contents
# * [Executive Summary](#exec-sum)
# * [Data Wrangling](#data-wrangle)
# * [Loading and parsing data](#load-parse)
# * [Translate to English](#translate)
# * [String preprocessing](#string-prep)
# * [Exploratory Analysis](#ea)
# * [Matching Products](#match-prod)
# * [Method 1 - MinHash + LSH](#m1)
# * [Training](#m1_train)
# * [Validation](#m1_val)
# * [Method 2 - Levenshtein distance + Thresholding](#m1)
# * [Training](#m2_train)
# * [Validation](#m2_val)
#
#
# ## Executive summary <a class="anchor" id="exec-sum"></a>
#
# * The MinHash + LSH approach seems to be able to find matches, but requires improvement to reach an acceptable F1-score [This was not feasible in the time frame]:
# * PRECISION - **37.4 %**
# * RECALL - **52.1 %**
# * F1-SCORE - **43.5 %**
# * The Levenshtein distance method is a naive one, and was used simply to gain some intuition on the data
# * Other variants of LSH methods, as well as methods listed in future scope can be tried to improve accuracy
# * Blocking can be carried out by BRAND to improve matching. Exploratory analysis shows that products are very unevenly distributed across brands
# ## Data wrangling <a class="anchor" id="data-wrangle"></a>
# ### Load and parse data <a class="anchor" id="load-parse"></a>
import pandas as pd
import json
import fileinput
# +
def read_json_by_line(line):
"""
Takes in a line of a file, and parses it to JSON
Args:
line - Line from a file
Returns:
Parsed JSON for that line
"""
line.strip()
try:
js = json.loads(line)
except:
print("Error :" + line)
return(js)
n = 10
with fileinput.input('../data/dataset.jsonl') as file:
json_list = [read_json_by_line(line) for line in file]
df = pd.concat(list(map(lambda js : pd.io.json.json_normalize(js),
json_list)), sort = False)
# -
df.to_csv("../data/raw_data.csv")
del df
# +
def drop_col_by_pattern(df, col_patterns):
"""
Takes in list of patterns for column names to drop from the dataset
and returns a the dataset with columns dropped
Args:
df - The Pandas DataFrame from which columns need to be dropped
col_patterns - List of patterns to check
Returns:
A Pandas DataFrame with columns dropped
"""
for p in col_patterns:
df = df.drop(df.filter(regex = p).columns, axis = 1)
return(df)
# -
cols = ['NAME', 'DESCRIPTION', 'DALTIX_ID', 'BRAND']
final_df = pd.read_csv("../data/raw_data.csv", skipinitialspace=True, usecols=cols)
#final_df = drop_col_by_pattern(df, col_patterns)
final_df.count()
# ### Translate all to English <a class="anchor" id="translate"></a>
# +
import re
from googletrans import Translator
def clean_and_translate(t):
"""
Takes in a text and translates it to English. Additionally the text
is cleaned of special characters except space
Args:
t - Text to translate
Returns:
Translated text
"""
translator = Translator()
t = re.sub(r"[^a-zA-Z0-9]+", ' ', t)
#try:
#translator.translate(t).text
#except:
# print("Error in translation API")
return(t)
def translate_cols(df, cols):
"""
Translates all the rows of the specified columns of a data frame
Args:
df - A Pandas DataFrame containing the columns to translate
cols - A list of the names of the columns to be translated
Returns:
A Pandas DataFrame with specified columns translated
"""
for c in cols:
df[c] = df[c].apply(lambda t: clean_and_translate(t) if t is not None else t)
return(df)
cols = ['NAME', 'DESCRIPTION']
#final_df = translate_cols(final_df, cols)
# -
final_df['DESCRIPTION'].head()
final_df.to_csv("../data/translated_data.csv")
final_df = pd.read_csv("../data/translated_data.csv")
# ### String preprocessing <a class="anchor" id="string-prep"></a>
import nltk
#nltk.download('stopwords')
#nltk.download('punkt')
#nltk.download('wordnet')
final_df.head()
# +
from nltk.corpus import stopwords
from nltk.tokenize import word_tokenize
from nltk.stem import PorterStemmer
from nltk.stem import WordNetLemmatizer
def remove_stop_words(string):
"""
Removes stopwords from a given string, based on the NLTK English dictionary
of stopwords
Args:
string - The string on which stopwords are to be removed
Returns:
A string with stopwords removed
"""
stop_words = set(stopwords.words('english'))
words = string.split()
cleaned_string = ""
for r in words:
if not r in stop_words:
cleaned_string = cleaned_string + r + " "
return(cleaned_string)
#remove_stop_words("The quick brown fox went to a school")
def stem_words(string):
"""
Stems words in a given string, using the Porter Stemmer
for English
Args:
string - The string to be stemmed
Returns:
A string with its words stemmed
"""
stemmer = PorterStemmer()
words = word_tokenize(string)
stemmed_string = ""
for w in words:
sw = stemmer.stem(w)
stemmed_string = stemmed_string + sw + " "
return(stemmed_string)
#stem_words("The world is a troubling place where I am annoyed")
def lemmatize_words(string):
"""
Lemmatizes words a given string, based on the WordNet lemmatizer
for English
Args:
string - The string to be lemmatized
Returns:
A string with its words lemmatized
"""
wordnet_lemmatizer = WordNetLemmatizer()
words = word_tokenize(string)
lemmatized_string = ""
for w in words:
lw = wordnet_lemmatizer.lemmatize(w)
lemmatized_string = lemmatized_string + lw + " "
return(lemmatized_string)
#lemmatize_words("The world is a troubling place where I am annoyed")
def clean_string_in_cols(df, cols, operations = ['stem', 'lemmatize', 'stop-word']):
"""
Preprocesses string columns of a data frame with the ability to optionally select
various operations such as stemming, lemmatization and stop word removal.
Args:
df - The data frame containing the string columns to be processed
cols - The string columns which should be processed
operations - A list of operations to be executed. Options are stem, lemmatize,
and stop-word
Returns:
A data frame with the string columns processed
"""
for c in cols:
if 'stop-word' in operations:
df[c] = df[c].apply(lambda t: remove_stop_words(t) if t is not None else t)
if 'stem' in operations:
df[c] = df[c].apply(lambda t: stem_words(t) if t is not None else t)
if 'lemmatize' in operations:
df[c] = df[c].apply(lambda t: lemmatize_words(t) if t is not None else t)
return(df)
cols = ['NAME', 'DESCRIPTION']
operations = ['stop-word']
final_df = clean_string_in_cols(final_df, cols, operations)
# -
final_df.head()
final_df.head()
final_df.to_csv('../data/trans_processed_data.csv')
del final_df
# ## Exploratory analysis <a class="anchor" id="ea"></a>
res = final_df.groupby("BRAND").agg('count')
res
# +
from wordcloud import WordCloud
comment_words = ' '
for val in final_df.NAME:
val = str(val)
tokens = val.split()
for i in range(len(tokens)):
tokens[i] = tokens[i].lower()
for words in tokens:
comment_words = comment_words + words + ' '
wordcloud = WordCloud(width = 800, height = 800,
background_color ='white',
min_font_size = 10).generate(comment_words)
# +
import matplotlib.pyplot as plt
plt.figure(figsize = (8, 8), facecolor = None)
plt.imshow(wordcloud)
plt.axis("off")
plt.tight_layout(pad = 0)
plt.show()
# -
# ## Matching Products <a class="anchor" id="match-prod"></a>
# ### Method 1 - MinHash + LSH <a class="anchor" id="m1"></a>
# #### Training <a class="anchor" id="m1_train"></a>
from __future__ import division
import os
import re
import random
import time
import binascii
import base64
from bisect import bisect_right
from heapq import heappop, heappush
minhash_df = pd.read_csv('../data/trans_processed_data.csv')
minhash_df = drop_col_by_pattern(minhash_df, "Unnamed")
minhash_df.head()
# +
#minhash_df = final_df.loc[:, ['DALTIX_ID', 'NAME', 'DESCRIPTION', 'BRAND']]
def concat_cols(row, cols):
concat_op = ''
for c in cols:
concat_op = concat_op + row[c] + " | "
return concat_op
feature_cols = ['NAME', 'DESCRIPTION', 'BRAND']
minhash_df['TEXT'] = minhash_df.loc[:, feature_cols].astype(str).apply(lambda r:
concat_cols(r, feature_cols)
if all(v is not None for v in r)
else r, axis = 1)
minhash_df_final = minhash_df.loc[:, ['DALTIX_ID','TEXT']]
minhash_df_final.head()
# -
del minhash_df
minhash_df_final.to_csv('../data/minhash_df.csv')
minhash_df_final = pd.read_csv('../data/minhash_df.csv')
# +
import binascii
def get_shingles_in_text(words):
"""
Construct shingles of 3 words from a list of words, and convert them to hashes
Args:
words - A list of words
Returns:
A list of hashes of shingles
"""
text_shingles = set()
for i in range(1, len(words) - 2):
shingle = words[i] + " " + words[i + 1] + words[i + 2]
hash_val = binascii.crc32(shingle.encode()) & 0xffffffff
text_shingles.add(hash_val)
return(text_shingles)
def get_shingles_in_df(df, id_col, text_col):
"""
Construct shingles for each row of a column in a data frame and create a key-value
pair of id and shingle hash
Args:
df - The data frame containing the string column to be processed
id_col - The column name of the id variable
text_col - The name of the string column which should be processed
Returns:
A set of key-value pairs
"""
text_as_shingle_sets = set()
minhash_df_final[text_col].str.split(" ")
text_as_shingle_sets[df[id_col]] = df[text_col].map(lambda x : get_shingles_in_text(x))
return(text_as_shingle_sets)
text_as_shingle_sets = minhash_df_final['TEXT'].map(lambda x : get_shingles_in_text(x))
# +
# data sketch
from datasketch import MinHash, MinHashLSH
def get_minhash(text):
"""
Generate a MinHash for a text
Args:
text - text for which MinHash is to be generated
Returns:
MinHash object
"""
text = f'{text}'
words = set(text.split())
m = MinHash(num_perm = 64)
for w in words:
m.update(w.encode('utf8'))
return(m)
def minhash_lsh_sim(df, id_col, text_col,
jaccard_sim_threshold = 0.5):
"""
Generates a set of MinHashes for the all the text values,
and and LSH object defined with a particular threshold of Jaccard similarity
which can be queried for matches
Args:
df - The data frame containing the string column
id_col - The column name of the id variable
text_col - The name of the string column
Returns:
A set of MinHashes and an LSH object to be queried for matches
"""
minhashes = {}
lsh = MinHashLSH(jaccard_sim_threshold, num_perm = 64)
for index, row in df.iterrows():
id_val = row[id_col]
text = row[text_col]
mh = get_minhash(text)
minhashes[id_val] = mh
lsh.insert(id_val, mh)
return minhashes, lsh
minhashes, lsh = minhash_lsh_sim(minhash_df_final, 'DALTIX_ID', 'TEXT', 0.999)
# +
import random
m1 = minhashes[minhash_df_final['DALTIX_ID'].iloc[random.randint(0,99999)]]
m2 = minhashes[minhash_df_final['DALTIX_ID'].iloc[random.randint(0,99999)]]
lsh.query(m1)
#m1.jaccard(m2)
# +
def get_submission(df, id_col, text_col, minhashes, lsh):
"""
Construcs a list of matches for submission
Args:
df - The data frame containing the string column
id_col - The column name of the id variable
text_col - The name of the string column,
minhashes - Set of MinHashes
lsh - LSH object to be queried
Returns:
A list for of matched pairs for submission
"""
submission_list = []
for index, row in df.iterrows():
id_val = row[id_col]
text = row[text_col]
mh = minhashes.get(id_val)
matches = lsh.query(mh)
for m in matches:
submission_list.append((id_val, m))
return(submission_list)
submission = get_submission(minhash_df_final, 'DALTIX_ID', 'TEXT', minhashes, lsh)
sub_df = pd.DataFrame(submission, columns = ['daltix_id_1', 'daltix_id_2'])
# -
sub_df = sub_df[sub_df['daltix_id_1'] != sub_df['daltix_id_2']]
len(sub_df.index)
# +
#del minhash_df_final
# -
sub_df.to_csv("../submission/submission.csv")
# #### Validation <a class="anchor" id="m1_val"></a>
eval_data = pd.read_csv("../data/y_true.csv")
# +
eval_data.columns = ['daltix_id_1', 'daltix_id_2']
eval_data_rev = eval_data
eval_data_rev.columns = ['daltix_id_2', 'daltix_id_1']
#eval_data_rev.head()
tp = len(pd.concat([sub_df.merge(eval_data, on = ['daltix_id_1', 'daltix_id_2']),
sub_df.merge(eval_data_rev, on = ['daltix_id_1', 'daltix_id_2'])]).index)
fp = len(sub_df.index) - tp
precision = tp/len(sub_df.index)
print("PRECISION: ", precision*100, "%")
recall = tp/len(eval_data.index)
print("RECALL: ", recall*100, "%")
f1 = 2/((1/recall) + (1/precision))
print("F1-SCORE: ", f1*100, "%")
# -
# ### Method 2 - String similarity and thresholding <a class="anchor" id="m2"></a>
# #### Training <a class="anchor" id="m2_train"></a>
# +
final_df = pd.read_csv('../data/trans_processed_data.csv')
final_df = drop_col_by_pattern(final_df, 'Unnamed')
cross_df = final_df.assign(tmp = 1).merge(final_df.assign(tmp = 1), on = 'tmp').drop('tmp', 1)
cross_df.head()
# +
from nltk.metrics import edit_distance
# Calculating the Levenstein similarity per column
def calc_lev_dist(cross_df, edit_distance_col_list):
"""
Calculates the Levenshtein Edit similarity between string columns
Args:
cross_df - A data frame containing a cross product of records
edit_distance_col_list - Names of columns to be considered for calculating the similarity
Returns:
An updated data frame containing column-wise and total similarity for each record
"""
for col in edit_distance_col_list:
cross_df[col + '_DIST'] = cross_df.loc[:, [col + '_x', col + '_y']].dropna().apply(lambda x :
edit_distance(*x)
#if all(v is not None for v in x)
#else 0.00
, axis = 1)
max_val = max(cross_df.loc[:, col + '_DIST'])
cross_df[col + '_SIM'] = cross_df.loc[:, col + "_DIST"].apply(lambda x : 1 - x/max_val)
return(cross_df)
edit_distance_col_list = ['NAME', 'DESCRIPTION']
cross_df = calc_lev_dist(cross_df, edit_distance_col_list)
# +
# Calculating overall Levenshtein similarity across columns
cross_df['TOTAL_SIM'] = cross_df.loc[:,
cross_df.filter(regex = '_SIM').columns].apply(lambda x : sum(x)/len(x),
axis = 1)
cross_df = cross_df[cross_df.TOTAL_SIM != 1]
# -
cross_df.loc[:, ['BRAND_x', 'BRAND_y']]
# +
threshold = 0.3
# Similarity on NAME, DESCRIPTION
result = cross_df[
(cross_df.NAME_SIM > threshold) &
#(cross_df.BRAND_x == cross_df.BRAND_y) &
(cross_df.SHOP_x != cross_df.SHOP_y)
].loc[: , ['DALTIX_ID_x', 'DALTIX_ID_y']]
result.to_csv('../submission/submissions_lev.csv')
# -
# #### Validation <a class="anchor" id="m2_val"></a>
# +
eval_data = pd.read_csv("../data/y_true.csv")
sub_lev_df = pd.read_csv('../submission/archive/submissions_lev.csv')
sub_lev_df = drop_col_by_pattern(sub_lev_df, 'Unnamed')
sub_lev_df.columns = ['daltix_id_1', 'daltix_id_2']
# +
eval_data.columns = ['daltix_id_1', 'daltix_id_2']
eval_data_rev = eval_data
eval_data_rev.columns = ['daltix_id_2', 'daltix_id_1']
tp = len(pd.concat([sub_df.merge(eval_data, on = ['daltix_id_1', 'daltix_id_2']),
sub_df.merge(eval_data_rev, on = ['daltix_id_1', 'daltix_id_2'])]).index)
fp = len(sub_df.index) - tp
precision = tp/len(sub_df.index)
print("PRECISION: ", precision*100, "%")
recall = tp/len(eval_data.index)
print("RECALL: ", recall*100, "%")
f1 = 2/((1/recall) + (1/precision))
print("F1-SCORE: ", f1*100, "%")
| code/Product_Matching_Based_On_Text_Attributes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="fhjkJlpnhWeb"
import pandas as pd
df = pd.read_pickle("202012-202103_processed.pkl")
# + colab={"base_uri": "https://localhost:8080/", "height": 589} id="1Vnz0ysgQ4Ae" outputId="cbf6ed71-dc64-4701-e723-31e8bbb51e32"
df
# + id="_-YgvR6jk4Wo"
content=df.content_for_lda.to_list()
# + id="r4dSPNo1k7ef"
content=[x.lower() for x in content]
# + colab={"base_uri": "https://localhost:8080/"} id="BpU2Yr-cwv62" outputId="5dbc4391-af33-4eab-92b1-88a1e37f6f1f"
content
# -
remove_list = ['-pron-','not','say']
corpus = []
for index in range(len(content)):
temp = content[index].split(" ")
#temp = [x.lower() for x in temp]
sen = [word for word in temp if not (len(word) < 3 or word in remove_list)] #去掉小于3个字符或在remove_list中的词
corpus.append(sen)
# + colab={"base_uri": "https://localhost:8080/"} id="hIaWEq5YlHgD" outputId="4b235d05-87d2-4294-e896-13dfd127908f"
len(corpus)
# -
del content
# + id="1ZJzeCjChzPG"
import numpy as np
import time
from gensim import corpora
t1=time.time()
dictionary = corpora.Dictionary(corpus,prune_at=3000000)
t2=time.time()
print('生成词典耗时',t2-t1,'s')
# + id="rxOOeQs0iy4w"
dictionary.filter_extremes(no_below=75,no_above=0.9) # 删掉只在不超过75个文本中出现过的词,删掉在90%及以上的文本都出现了的词
#dictionary.filter_n_most_frequent(10)
dictionary.compactify() # 去掉因删除词汇而出现的空白
# + colab={"base_uri": "https://localhost:8080/"} id="XuDCA7t5iEto" outputId="e97a572d-0d62-4345-cc60-7ff37c2807a8"
print(dictionary)
# + id="XevoowyvlYrD"
t1=time.time()
vec = [dictionary.doc2bow(s) for s in corpus]
t2=time.time()
print('生成词袋模型耗时',t2-t1,'s')
# -
del corpus
vec[0]
# + colab={"base_uri": "https://localhost:8080/"} id="xO-rQJeClmt_" outputId="f43eca21-e2e2-4dd2-8d71-d2bd917ae216"
vec[3482316]
# +
import logging
if len(vec) == len(df):
print('文档数量正常')
else:
logging.critical('需检查文档索引')
# -
dictionary.save('dict.dict')
corpora.MmCorpus.serialize('bow.mm',vec)
| gensim_lda.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Analysing FiveThirtyEight's 2014 Star Wars Survey
#
# In this project we'll be exploring, cleaning and analysing a data set abou Star Wars from the [FiveThirtyEight website](https://fivethirtyeight.com/). The main goal of this project to answer some questions about the popularity of each Star Wars episode.
#
# While waiting for Star Wars: The Force Awakens to come out, the team at FiveThirtyEight surveyed Star Wars fans using the online tool SurveyMonkey. They received 835 total responses, which you download from their [GitHub repository](https://github.com/fivethirtyeight/data/tree/master/star-wars-survey).The data has several columns, including:
#
# * RespondentID - An anonymized ID for the respondent (person taking the survey)
# * Gender - The respondent's gender
# * Age - The respondent's age
# * Household Income - The respondent's income
# * Education - The respondent's education level
# * Location (Census Region) - The respondent's location
# * Have you seen any of the 6 films in the Star Wars franchise? - Has a Yes or No response
# * Do you consider yourself to be a fan of the Star Wars film franchise? - Has a Yes or No response
# * Which character shot first?
# * Are you familiar with the Expanded Universe?
#
#
#
# ## Exploring the Data
#
# Let's begin by importing the libraries we'll use and reading the `.csv` file into a DataFrame.
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
star_wars = pd.read_csv('datasets/star_wars.csv', encoding='ISO-8859-1')
# -
# Let's take a look at the first five rows of the `star_wars` DataFrame and then we'll print the name of all its columns.
star_wars.head()
star_wars.columns
# We can notice three main points here:
#
# * The `RespondentID` column is supposed to be a unique iD for each respondent, but as we can see it contains some null values, which does not make much sense. We'll fix this by removing these those rows.
#
# * Some columns, such as the second and third ones, only contain values fo 'Yes' and 'No'. We can transform these values into booleans so it's easier to work with.
#
# * For some questions, the respondent had to check one or more boxes and this type of data is difficult to represent in columnar format. For instance, the respondent checked off a series of boxes in response to the question, `Which of the following Star Wars films have you seen? Please select all that apply.` The columns for this question are:
#
# * Which of the following Star Wars films have you seen? Please select all that apply. - Whether or not the respondent saw Star Wars: Episode I The Phantom Menace.
# * Unnamed: 4 - Whether or not the respondent saw Star Wars: Episode II Attack of the Clones.
# * Unnamed: 5 - Whether or not the respondent saw Star Wars: Episode III Revenge of the Sith.
# * Unnamed: 6 - Whether or not the respondent saw Star Wars: Episode IV A New Hope.
# * Unnamed: 7 - Whether or not the respondent saw Star Wars: Episode V The Empire Strikes Back.
# * Unnamed: 8 - Whether or not the respondent saw Star Wars: Episode VI Return of the Jedi.
#
# For columns like these, we'll also tranform the data into boolean values. We'll rename the columns so their name becomes more intuitive too.
# ## Cleaning the Data
#
# First, let's exclude the rows where the `RespondentID` column is null.
star_wars = star_wars[star_wars['RespondentID'].notnull()]
star_wars['RespondentID'].isnull().sum()
# Now, we'll use the `Series.map()` method to solve the second point mentioned above, transforming the values in the second and third columns to boolean values.
# +
yes_no = {'Yes' : True, 'No' : False}
star_wars['Have you seen any of the 6 films in the Star Wars franchise?'] = star_wars['Have you seen any of the 6 films in the Star Wars franchise?'].map(yes_no)
star_wars['Do you consider yourself to be a fan of the Star Wars film franchise?'] = star_wars['Do you consider yourself to be a fan of the Star Wars film franchise?'].map(yes_no)
# -
# Let's check the result.
star_wars.head()
# We'll also use the `Series.map()` method to deal with the third point we listed, transforming the values in the next six columns to booleans.
# +
films_seen = {
'Star Wars: Episode I The Phantom Menace': True, np.nan: False,
'Star Wars: Episode II Attack of the Clones' : True,
'Star Wars: Episode III Revenge of the Sith' : True,
'Star Wars: Episode IV A New Hope' : True,
'Star Wars: Episode V The Empire Strikes Back' : True,
'Star Wars: Episode VI Return of the Jedi' : True
}
for col in star_wars.columns[3:9]:
star_wars[col] = star_wars[col].map(films_seen)
star_wars.head()
# -
# As these columns represent if each respondent saw each movie, we'll rename them so they become more intuitive.
star_wars = star_wars.rename(columns = {
'Which of the following Star Wars films have you seen? Please select all that apply.' : 'seen_1',
'Unnamed: 4' : 'seen_2',
'Unnamed: 5' : 'seen_3',
'Unnamed: 6' : 'seen_4',
'Unnamed: 7' : 'seen_5',
'Unnamed: 8' : 'seen_6'
})
star_wars.columns
# As we move on to next columns, we can see that they show the same pattern. The difference is that these columns contain values from 1 to 6 that represent the ranking of preffered films for each respondent.
#
# * Please rank the Star Wars films in order of preference with 1 being your favorite film in the franchise and 6 being your least favorite film. - The respondent' ranking for Star Wars: Episode I The Phantom Menace
# * Unnamed: 10 - The respondent' ranking for Star Wars: Episode II Attack of the Clones
# * Unnamed: 11 - The respondent' ranking for Star Wars: Episode III Revenge of the Sith
# * Unnamed: 12 - The respondent' ranking for Star Wars: Episode IV A New Hope
# * Unnamed: 13 - The respondent' ranking for Star Wars: Episode V The Empire Strikes Back
# * Unnamed: 14 - The respondent' ranking for Star Wars: Episode VI Return of the Jedi
#
# We'll convert these values to a numeric type and then rename the columns, just like we did for the past six columns.
star_wars[star_wars.columns[9:15]] = star_wars[star_wars.columns[9:15]].astype(float)
star_wars = star_wars.rename(columns = {
'Please rank the Star Wars films in order of preference with 1 being your favorite film in the franchise and 6 being your least favorite film.' : 'ranking_1',
'Unnamed: 10' : 'ranking_2',
'Unnamed: 11' : 'ranking_3',
'Unnamed: 12' : 'ranking_4',
'Unnamed: 13' : 'ranking_5',
'Unnamed: 14' : 'ranking_6'
})
star_wars.columns
# ## Analysis
#
# As these columns now contain numeric values, we can calculate the average ranking for each movie.
ranking = star_wars[star_wars.columns[9:15]].mean()
ranking.sort_values(ascending=False)
# We can also plot this in a bar chart. Note that we first created two lists that are used to set the labels and colors in the charts.
# +
films = ['Episode 1', 'Episode 2', 'Episode 3', 'Episode 4', 'Episode 5', 'Episode 6']
colors = [(0/255,107/255,164/255), (255/255, 128/255, 14/255), (44/255, 160/255, 44/255), (214/255, 39/255, 40/255), (188/255, 189/255, 34/255), (148/255, 103/255, 89/255)]
fig, ax = plt.subplots()
ax.bar(range(6), ranking, align='center', color=colors, edgecolor=colors)
ax.tick_params(bottom=False, top=False, left=False, right=False)
plt.xticks(np.arange(6), films, rotation=45)
ax.set_ylim(0,6)
ax.set_ylabel('Average Ranking')
for kew, spine in ax.spines.items():
spine.set_visible(False)
plt.show()
# -
# Episode 3 has the highest average ranking and episode 5 has the lowest one.
# We'll repeat the process to find out the most seen movie. But now we'll transform the values to percentages.
most_seen = star_wars[star_wars.columns[3:9]].sum() / star_wars.shape[0] * 100
most_seen.sort_values(ascending=False)
# +
fig, ax = plt.subplots()
ax.bar(range(6), most_seen, align='center', color=colors, edgecolor=colors)
ax.tick_params(bottom=False, top=False, left=False, right=False)
plt.xticks(np.arange(6), films, rotation=45)
ax.set_ylabel('% that watched')
for kew, spine in ax.spines.items():
spine.set_visible(False)
plt.show()
# -
# Episodes 5 and 6 are the most seen while Episode 3 is the less seen. It's interesting to note that the most seen episode has the lowest ranking while the less seen episode has the highest ranking.
# ### Gender
#
# We'll now split the DataFrame by genders and then we'll see the ranking and the most watched episodes by each gender.
# +
males = star_wars[star_wars['Gender'] == 'Male']
females = star_wars[star_wars['Gender'] == 'Female']
ranking_m = males[males.columns[9:15]].mean()
ranking_f = females[females.columns[9:15]].mean()
print('Male ranking')
print(ranking_m.sort_values(ascending=False))
print('\n')
print('Female ranking')
print(ranking_f.sort_values(ascending=False))
# +
fig = plt.figure(figsize=(10, 4))
for i in range(0,2):
ax = fig.add_subplot(1, 2, i+1)
chart = ranking_f
title = 'Female Ranking'
ax.tick_params(labelleft=False)
if i == 0:
chart = ranking_m
title = 'Male Ranking'
ax.set_ylabel('Average Ranking')
ax.tick_params(labelleft=True)
ax.bar(range(6), chart, align='center', color=colors, edgecolor=colors)
ax.tick_params(bottom=False, top=False, left=False, right=False)
plt.xticks(np.arange(6), films, rotation=45)
ax.set_title(title)
ax.set_ylim(0,6)
for kew, spine in ax.spines.items():
spine.set_visible(False)
plt.show()
# -
# The charts are pretty much the same.
#
# Now let's see the most seen movies for each gender.
# +
seen_m = males[males.columns[3:9]].sum()/ males.shape[0] * 100
seen_f = females[females.columns[3:9]].sum()/ females.shape[0] * 100
print('Male ranking')
print(seen_m.sort_values(ascending=False))
print('\n')
print('Female ranking')
print(seen_f.sort_values(ascending=False))
# +
fig = plt.figure(figsize=(12, 5))
for i in range(0,2):
ax = fig.add_subplot(1, 2, i+1)
chart = seen_f
title = 'Female Most Seen'
ax.tick_params(labelleft=False)
if i == 0:
chart = seen_m
title = 'Male Most Seen'
ax.set_ylabel('% that watched')
ax.tick_params(labelleft=True)
ax.bar(range(6), chart, align='center', color=colors, edgecolor=colors)
ax.tick_params(bottom=False, top=False, left=False, right=False)
plt.xticks(np.arange(6), films, rotation=45)
ax.set_title(title)
ax.set_ylim(0,100)
for kew, spine in ax.spines.items():
spine.set_visible(False)
plt.show()
# -
# The order of the most watched episodes is the same. Less women watched the movies, though.
# ### Education
# We'll now explore the `Education` column as we create a ranking for each distinct level of edcuation in the survey.
# +
unique_edu = star_wars['Education'].dropna().unique()
rankings_edu = []
for c in unique_edu:
df = star_wars[star_wars['Education'] == c]
ranking_c = df[df.columns[9:15]].mean()
rankings_edu.append(ranking_c)
print(c)
print(ranking_c.sort_values(ascending=False))
print('\n')
# +
fig = plt.figure(figsize=(16,12))
for i in range (0,5):
ax = fig.add_subplot(2, 3, i+1)
ax.bar(range(6), rankings_edu[i], align='center', color=colors, edgecolor=colors)
ax.set_title(unique_edu[i])
ax.tick_params(bottom=False, top=False, left=False, right=False, labelbottom=False, labelleft=False)
ax.set_ylim(0,6)
plt.xticks(np.arange(6), films, rotation=45)
if i >= 2:
ax.tick_params(labelbottom=True)
if i ==0 or i == 3:
ax.tick_params(labelleft=True)
for kew, spine in ax.spines.items():
spine.set_visible(False)
plt.show()
# -
# Episode 5 has the lowest ranking in every level of education, while Episode 3 has the highest ranking in all levels of education but 'Less than high school degree'. The ranking for peolple with less than high school degree also is the most volatile one, the reason for this might be that it is the category with the smallest sample, as we can see below.
star_wars['Education'].value_counts()
# ### Who Shot First?
#
# As we approch the end of this projecct, we'll investigate the relation between the answer in the 'Do you consider yourself to be a fan of the Star Wars film franchise?' column and the answer in the 'Which character shot first?' column. That is a [controversial topic](https://en.wikipedia.org/wiki/Han_shot_first) for Star Wars fans and we expect to find that the answer for this question differs if the respondent considers himself a Star Wars fan or not.
# +
fan = star_wars['Do you consider yourself to be a fan of the Star Wars film franchise?'].dropna().unique()
fan_shot = []
for i in fan:
df = star_wars[star_wars['Do you consider yourself to be a fan of the Star Wars film franchise?'] == i]
shot = pd.Series(df.iloc[:,29].value_counts().sort_index())
fan_shot.append(shot)
print('Fan: {}'.format(i))
print(shot.sort_values(ascending=False))
print('\n)')
# -
# Now let's plot the charts.
# +
fig = plt.figure(figsize=(12,6))
ax = fig.add_subplot(1, 2, 1)
ax.bar(range(3), fan_shot[0], align='center', color=colors, edgecolor=colors)
ax.set_title('Fan')
ax.tick_params(bottom=False, top=False, left=False, right=False, labelbottom=False)
ax.set_ylim(0,300)
ax.set_ylabel('Amount of votes')
for kew, spine in ax.spines.items():
spine.set_visible(False)
ax = fig.add_subplot(1, 2, 2)
ax.bar(range(2,3), fan_shot[1][2], align='center', color=(44/255, 160/255, 44/255), edgecolor=(44/255, 160/255, 44/255), label="I don't understand")
ax.bar(range(1,2), fan_shot[1][1], align='center', color=(255/255, 128/255, 14/255), edgecolor=(255/255, 128/255, 14/255), label='Han')
ax.bar(range(1), fan_shot[1][0], align='center', color=(0/255,107/255,164/255), edgecolor=(0/255,107/255,164/255), label='Greedo')
ax.set_title('Not a Fan')
ax.tick_params(bottom=False, top=False, left=False, right=False, labelleft=False, labelbottom=False)
ax.set_ylim(0,300)
ax.legend()
for kew, spine in ax.spines.items():
spine.set_visible(False)
plt.show()
# -
# As expected, the answers were very different. Most of the fans think Han Solo shot firts and the others are pretty divided between Greedo shot first and those that did not understand the question. For the no fans, however, the absolutely marjority did not understand the question. It is also import no note that the number of fans the reponded the survey is greater than the number of not fans.
| Analysing FiveThirtyEight's 2014 Star Wars Survey.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Dummy Variables & One Hot Encoding
# <img src="ct_vb.png" height=600 width=600>
# <img src="1hot&encode.png" height=600 width=600>
# +
import pandas as pd
df = pd.read_csv("homeprices.csv")
df
# -
# ### Using pandas to create dummy variables
dummies = pd.get_dummies(df.town) # Dummies of town column
dummies
merged_df = pd.concat([df,dummies],axis='columns')
merged_df
final_df = merged_df.drop(['town'], axis='columns') # No longer we need town column
final_df
# # Dummy Variable Trap
#
# When deriving one variable from other variables, they are known to be multi-colinear. Here, known values of california and georgia can easily infer value of new jersey state, i.e. california=0 and georgia=0. There for these state variables are called to be multi-colinear. In this situation linear regression won't work as expected. Hence, we need to drop one column.
#
# *NOTE: sklearn library takes care of dummy variable trap hence even if we don't drop one of the state columns it is going to work, however we should make a habit of taking care of dummy variable trap ourselves just in case library that we are using is not handling this for us*
final_df = final_df.drop(['west windsor'], axis='columns')
final_df # Avoiding dummy variables trap
X = final_df.drop('price', axis='columns') # Since price is our dependent variable
X
y = final_df.price
y
# +
from sklearn.linear_model import LinearRegression
model = LinearRegression()
model.fit(X,y)
# -
model.predict(X) # Predicting prices for all X
# #### Checking Accuracy
model.score(X,y)
# #### Prediction
model.predict([[3400,0,0]]) # 3400 sqr ft home in west windsor
model.predict([[3400,0,0]]) # 3400 sqr ft home in west windsor
# ### Using sklearn OneHotEncoder
# First step is to use label encoder to convert town names into numbers
from sklearn.preprocessing import LabelEncoder # 1. Using Label Encoder
lbl_e = LabelEncoder()
df_lbl_e = df
df_lbl_e.town = lbl_e.fit_transform(df.town)
df_lbl_e
X = df_lbl_e[['town','area']].values # Converting to 2D arrays
X
y = df_lbl_e.price.values # Price array
y
# ### Now using one hot encoder to create dummy variables for each of the town
from sklearn.preprocessing import OneHotEncoder # 2. OneHotEncoder
from sklearn.compose import ColumnTransformer # 3. ColumnTransformer
ct = ColumnTransformer([('town', OneHotEncoder(), [0])], remainder = 'passthrough')
# Identifying town 0th column
X = ct.fit_transform(X)
X
X = X[:,1:] # X=X[Taking all the rows, Skipinng zero'th column] for avoiding dummy variable trap
# X=X[row,column]
X
model.fit(X,y)
model.predict([[0,1,3400]]) # 3400 sqr ft home in west windsor
model.predict([[1,0,2800]]) # 2800 sqr ft home in robbinsville
# ## Exercise
# At the same level as this notebook on github, it contains carprices.csv. This file has car sell prices for 3 different models. First plot data points on a scatter plot chart to see if linear regression model can be applied. If yes, then build a model that can answer following questions,
#
# 1) Predict price of a mercedez benz that is 4 yr old with mileage 45000
#
# 2) Predict price of a BMW X5 that is 7 yr old with mileage 86000
#
# 3) Tell me the score (accuracy) of your model. (Hint: use LinearRegression().score()))
| Machine Learning/5. Dummy Variables & One Hot Encoding.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pickle
import numpy as np
import matplotlib
matplotlib.use("Qt5Agg")
import matplotlib.pyplot as plt
#from galaxymodule.galaxy import Galaxy
import tree
import collections
#from galaxymodule import galaxy
import load
from analysis.cal_lambda import *
from temp.makegal import *
# +
nout=782
Mcut = 1e10
cal_lambda_params = dict(npix_per_reff=5,
rscale=3.0,
method='ellip',
n_pseudo=1,
verbose=False,
voronoi=None,#voronoi_dict,
mge_interpol = True)
mk_gal_params = dict()
# Load data
import tree
s = load.sim.Sim(nout=nout)
# Check for tree file before reading.
# Or, the fortran routine crashes causing the kernel crash without any error message.
# -
#no, ns, zr, ae =
nnza = np.genfromtxt("./nout_nstep_zred_aexp.txt",
dtype=[("nout", int),
("nstep", int),
("zred", float),
("aexp", float)])
all_gcats=[]
for nout in nnza["nout"]:
all_gcats.append(tree.halomodule.Halo(nout=nout, is_gal=True))
gcat = tree.halomodule.Halo(nout=nout, is_gal=True)
from galaxymodule import mk_gal_params as mgp
mgp.HAGN["verbose"] = False
tt = tree.tmtree.Tree(is_gal=True)
tnow = tt.tree[tt.tree["nstep"]==max(tt.tree["nstep"])]
large_last = tnow[(tnow["m"] > 1) * (tnow["m"] < 2)]
print(large_last["idx"])
final_idxs = large_last["idx"][large_last["id"] % 10 == 5]
final_ids = large_last["id"][large_last["id"] % 10 == 5]
for aexp in tt.aexps:
print(nnza["nout"][np.abs(aexp - nnza["aexp"]).argmin()])
merger_dist_all=[]
for fid,fidx in zip(final_ids, final_idxs):
maintree, idx_prgs_alltime = tt.extract_direct_full_tree(fidx)
all_direct_prgs = tt.get_all_trees(idx_prgs_alltime)
all_step=[]
for this_step in all_direct_prgs[1:2]:
step_now = 782
gg = load.rd_GM.Gal(nout=step_now,
catalog=np.copy(gcat.data[fid-1]),
info=s.info)
gg.debug=False
#gg.mk_gal(**mgp.HAGN)
mk_gal(gg,**mgp.HAGN)
gg.cal_norm_vec()
# Make sure that the catalog is not being modified.
all_gal = []
for this_gal in this_step:
this_gal_relang=[]
#this_gal_relang.append(0)
if this_gal is None:
continue
for gal in this_gal[0:1]:
#only at the last moment
#gal = this_gal[0]
print(gal["nstep"])
rel_pos = (gg.header["xg"] - gal["xp"])*1e3 # in kpc
rel_vel = gg.header["vg"] - gal["vp"]
jx,jy,jz = np.cross(rel_pos, rel_vel)
j_orbital=(jx,jy,jz)/np.sqrt(jx**2 + jy**2 + jz**2)
# rotation axis
relang = 180 / np.pi * np.arccos(np.dot(gg.meta.nvec, j_orbital))
this_gal_relang.append(relang)
all_gal.append(this_gal_relang)
#print("---")
all_step.append(all_gal)
merger_dist_all.append(all_step)
fig, ax = plt.subplots()
for all_step, idx in zip(merger_dist_all[2:3],final_idxs[2:3]):
for i, this_step in enumerate(all_step):
for all_gal in this_step:
ax.scatter(t_lookback[i],all_gal, c=cm.hot(i*4))
#for this_gal in all_gal:
#print(this_gal)
#print("--")
ax.set_ylim([0,180])
ax.set_title("Merger angle")
ax.set_xlabel("lookback time")
ax.set_ylabel("degree")
plt.show()#savefig("Merger_angle_dist_"+str(idx).zfill(8)+".png")
#plt.cla()
t_lookback = tt.age_univs[-1] - np.array(tt.age_univs[::-1])
# +
from matplotlib import cm
fig, ax = plt.subplots()
for all_step, idx in zip(merger_dist_all,final_idxs):
for i, this_step in enumerate(all_step):
for all_gal in this_step:
ax.scatter(t_lookback[i],all_gal, c=cm.hot(i*4))
#for this_gal in all_gal:
#print(this_gal)
#print("--")
ax.set_ylim([0,180])
ax.set_title("Merger angle")
ax.set_xlabel("lookback time")
ax.set_ylabel("degree")
plt.savefig("Merger_angle_dist_"+str(idx).zfill(8)+".png")
plt.cla()
# -
l = [item for sublist in merger_dist_all for item in sublist]
ll = [item for sublist in l for item in sublist]
lll = np.array(ll).ravel()
fig, ax = plt.subplots()
ax.hist(lll)
ax.set_title("relaive angle distribution (183 galaxies, 1~2e11M*, z=0)")
ax.set_xlabel("degree")
ax.set_ylabel("#")
plt.show()
# # B/T
gg.cal_b2t()
all_step=[]
for this_step in all_data[1:]:
all_gal = []
for this_gal in this_step:
this_gal_relang=[]
#this_gal_relang.append(0)
if this_gal is None:
continue
for gal in this_gal[0:1]: #only at the last moment
#gal = this_gal[0]
#print(gal)
rel_pos = (gg.header["xg"] - gal["xp"])*1e3 # in kpc
rel_vel = gg.header["vg"] - gal["vp"]
jx,jy,jz = np.cross(rel_pos, rel_vel)
j_orbital=(jx,jy,jz)/np.sqrt(jx**2 + jy**2 + jz**2)
# rotation axis
relang = 180 / np.pi * np.arccos(np.dot(gg.meta.nvec, j_orbital))
this_gal_relang.append(relang)
all_gal.append(this_gal_relang)
#print("---")
all_step.append(all_gal)
| scripts/scripts_ipynb/Merger_orbit_dist.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction to Programming and Numerical Analysis: Model Project
# +
# We import all the necessary packets at the beginning of our code:
import numpy as np
from scipy import linalg
from scipy import optimize
import sympy as sm
sm.init_printing(use_unicode=True)
# %matplotlib inline
import matplotlib.pyplot as plt
plt.style.use('seaborn-whitegrid')
from matplotlib import cm
from mpl_toolkits.mplot3d import Axes3D
import ipywidgets as widgets
# -
# ## **Basic OLG**
# ### **Introduction & household optimization**
#
# The Overlapping Generations Model (OLG) is a class of models in order to study the development of an economy, mostly economic growth. OLG models are characterized by discrete time periods such that in every period at least two different generations live. In the case we study here, there are two generations per period; a young generation that works and an old generation that is retired (meaning that the individuals themselves also only live for two periods, however they overlap with each others life cycles). They all consume in both periods but they only save in their life-time period 1 as they die at the end of period 2 and use all their resources before doing so. Hence, they have a labor income in their 1st period and a capital income in their 2nd period.
#
# - **Beginning of period t:** generation t born
# - **Period t:** generation t works
# - **End of period t:** genertaion t consumes and saves
# - **Period t+1:** generation t lends
# - **End of period t+1:** generation t consumes and dies
#
# We will first solve the general case of the model and then complement the model by including the government, which raises taxes or issues debt-bonds. By that, we show that the model is applicable in a much broader setting and therefore also of use to answer more sophisticated questions.
# First of all, we have the following CRRA life-time utility function:
#
# \begin{eqnarray*}
# U_t=\frac{C_{1t}^{1-\theta}}{1-\theta} + \frac{1}{(1+\rho)}\frac{C_{2t+1}^{1-\theta}}{1-\theta}
# \end{eqnarray*}
#
# However, due to simpliticity, but also as theta is not significantly different from 1 in many studies, we take
#
# \begin{eqnarray*}
# \theta=1
# \end{eqnarray*}
#
# Consequently, we need then to logarithmize and get:
#
# \begin{eqnarray*}
# U_t=ln(C_{1t})+ \frac{1}{(1+\rho)}ln(C_{2t+1})
# \end{eqnarray*}
#
# Furthermore, households have the following budget constraint:
#
# \begin{eqnarray*}
# A_tw_t=C_{1t}+ \frac{C_{2t+1}}{1+r_{t+1}}
# \end{eqnarray*}
#
# which comes from the single period constraints:
#
# \begin{eqnarray*}
# C_{1t}=A_tw_t-S_t
# \end{eqnarray*}
#
# and
#
# \begin{eqnarray*}
# C_{2t+1}=(1+r_{t+1} )S_t
# \end{eqnarray*}
#
# - $U_t$ = life-time utility
# - $C_1$ = consumption as young (in period t)
# - $C_2$ = consumption as old (in period t+1)
# - $\rho$ = discount rate on consumption (>-1)
# - $\theta$ = risk aversion parameter
# - $A_t$ = total factor productivity in period t
# - $w_t$ = wage rate
# - $S_t$ = savings in period t
# - $r_{t+1}$ = interest rate on savings (technicially paid at the end of period t+1)
# <p> <br>
# +
# Define the first sympy variables and parameters:
Ut = sm.symbols('U_t')
C1t = sm.symbols('C_1t')
C2t1 = sm.symbols('C_2t+1')
alpha = sm.symbols('alpha')
rho = sm.symbols('rho')
rt1 = sm.symbols('r_t+1')
wt = sm.symbols('w_t')
At = sm.symbols('A_t')
St = sm.symbols('S_t') # total savings
s = sm.symbols('s(r_t+1)') # saving rate
# +
# Define the utiliy function as sympy:
utility_function = sm.ln(C1t)+(1/(1+rho))*sm.ln(C2t1)
print('Utility Function:')
display(sm.Eq(Ut,utility_function))
# +
# Define the budget constraints as sympy:
budget_constraint_p1 = sm.Eq(C1t,wt*At-St)
budget_constraint_p2 = sm.Eq(C2t1,(1+rt1)*St)
St_from_bcp2 = sm.solve(budget_constraint_p2,St)
budget_constraint_by_sympy = budget_constraint_p1.subs(St,St_from_bcp2[0])
# Define budget constraint as in the outline:
budget_constraint = sm.Eq(C1t+(C2t1/(1+rt1)),wt*At)
# Showing the results:
print('Budget Constraints 1 & 2 as defined above:')
display(budget_constraint_p1)
display(budget_constraint_p2)
print('Combine the constraints:')
display(budget_constraint_by_sympy)
print('Reformulate:')
display(budget_constraint)
# +
# Optimum value for C1t by inserting budget constraint into the utility function and differentiating:
C2t1_from_con = sm.solve(budget_constraint,C2t1)
objective_subs = utility_function.subs(C2t1,C2t1_from_con[0])
foc1 = sm.diff(objective_subs,C1t)
sol1 = sm.solve(sm.Eq(foc1,0),C1t)
opt_C1t = sm.Eq(C1t,sol1[0])
# Optimum value for C2t1 by inserting budget constraint into the utility function and differentiating:
C1t_from_con = sm.solve(budget_constraint,C1t)
objective_subs = utility_function.subs(C1t,C1t_from_con[0])
foc2 = sm.diff(objective_subs,C2t1)
sol2 = sm.solve(sm.Eq(foc2,0),C2t1)
opt_C2t1 = sm.Eq(C2t1,sol2[0])
# We can then straightforward derive the Euler-equation (C2t1/C1t) by taking together the two latter results.
# The Euler-equation has the advantage that it shows the relative change in consumption and not the level.
euler = sol2[0]/sol1[0]
euler_equation=sm.Eq(C2t1/C1t,euler)
# Showing the results:
print('optimal Consumption in period 1:')
display(opt_C1t)
print('optimal Consumption in period 2:')
display(opt_C2t1)
print('Euler Equation:')
display(euler_equation)
# +
# In order to get total savings, we only have to rearrange the budget constraint from period 1 and insert optimal consumption we just derived:
savings_period1 = sm.solve(budget_constraint_p1,St)
savings_period1_fullequation = sm.Eq(St,savings_period1[0])
savings_formula = savings_period1_fullequation.subs(C1t,sol1[0])
savings_formula_simplified=sm.simplify(savings_formula)
# We print all the steps and the result:
print('Saving Equation:')
display(savings_period1_fullequation)
print('Insert optimal first period consumption:')
display(savings_formula)
print('Simplify:')
display(savings_formula_simplified)
# +
# Having calculated total savings, we can also calculate the saving rate:
total_savings=sm.Eq(St ,s*wt*At)
saving_r = sm.solve(total_savings,s)
saving_r2 = sm.Eq(s ,saving_r[0])
saving_rate_equation = saving_r2.subs(St,sm.simplify(savings_period1[0].subs(C1t,sol1[0])))
print('Total savings:')
display(total_savings)
print('Reformulate:')
display(saving_r2)
print('Insert Saving Equation and simplify:')
display(saving_rate_equation)
# +
# For later, we define the saving rate
# as saving_rate_equation shows the whole equation for illustration reasons, but we only need the right-hand term).
saving_rate = 1/(2+rho)
print('Saving Rate: s=')
display(saving_rate)
# -
# ### **Production**
# <p> <br>
# So far we have just introduced the households. We consider the case where firms have a standart neoclssical Cobb-Douglas production function:
#
# \begin{eqnarray*}
# Y_t=F(K_t,A_tL_t)=(K_t)^\alpha(A_tL_t)^{1-\alpha}
# \end{eqnarray*}
#
# where total factor productivity as well as population grow with a certain rate:
#
# \begin{eqnarray*}
# A_t=(1+g)A_{t-1}
# \end{eqnarray*}
#
# \begin{eqnarray*}
# L_t=(1+n)L_{t-1}
# \end{eqnarray*}
#
# - $K_t$ = capital stock in period t
# - $A_t$ = total factor productivity in t
# - $L_t$ = amount of labour in t
# - $\alpha$ = capital share in production
# - $g$ = growth rate of total factor productivity
# - $n$ = growth rate of population
#
# As we use the terms of effectice labor, it is however much easier to exert the intensive form of production. Hence we define:
#
# \begin{eqnarray*}
# \frac{1}{A_tL_t}F(K_t,A_tL_t)=F(\frac{K_t}{A_tL_t},1)
# \end{eqnarray*}
#
# and in this particular case
#
# \begin{eqnarray*}
# y=f(k_t)=k_t^\alpha
# \end{eqnarray*}
#
# with
#
# \begin{eqnarray*}
# k_t=\frac{K_t}{A_tL_t}, y_t=\frac{Y_t}{A_tL_t}, f(k_t)=F(k_t,1)
# \end{eqnarray*}
#
# We further assume that factor markets are perfektly competitive, i.e.:
#
# \begin{eqnarray*}
# r_t=f'(k_t)
# \end{eqnarray*}
#
# \begin{eqnarray*}
# w_t=f(k_t)-f'(k_t)k_t
# \end{eqnarray*}
# <p> <br>
# +
# We define the missing corresponding sympy variables:
yt = sm.symbols('y_t')
kt = sm.symbols('k_t')
rt = sm.symbols('r_t')
# +
# Define the production function:
f = kt**alpha
print('Production Function:')
display(sm.Eq(yt,f))
# +
# Calculate the real interest rate:
real_interest_rate = sm.diff(f,kt)
real_interest_rate = sm.simplify(real_interest_rate)
print('Real Interest Rate:')
display(sm.Eq(rt,real_interest_rate))
# +
# Calculate the wage rate per unit of effective labor:
wage_rate = f-sm.diff(f,kt)*kt
wage_rate = sm.simplify(wage_rate)
print('Wage Rate per Unit of Effective Labor:')
display(sm.Eq(wt,wage_rate))
# -
# ### **Balanced growth path & steady state**
# <p> <br>
# We are now able to set up the transition to the balanced growth path:
#
# \begin{eqnarray*}
# K_{t+1}=s(r_{t+1})w_tA_tL_t
# \end{eqnarray*}
#
# and therefore
#
# \begin{eqnarray*}
# k_{t+1}=s(r_{t+1})w_t\frac{A_tL_t}{A_{t+1}L_{t+1}}=s(r_{t+1})w_t\frac{A_tL_t}{A_t(1+g)L_t(1+n)}=s(r_{t+1})w_t\frac{1}{(1+g)(1+n)}
# \end{eqnarray*}
# <p> <br>
# +
# Define again the necessary sympy variables:
kt1 = sm.symbols('k_t+1')
g = sm.symbols('g')
n = sm.symbols('n')
# +
# We also define the transition function as sympy:
transition_int = sm.Eq(kt1,s*wt*(1/((1+g)*(1+n))))
print('Transition Equation:')
display(transition_int)
# +
# And we can plug in the saving rate s as well as the wage rate wt:
transition_int2 = transition_int.subs(s,saving_rate)
transition_int2 = transition_int2.subs(wt,wage_rate)
print('Transition Equation with Saving and Wage rates plugged-in:')
display(transition_int2)
# +
# We then simply replace k_t+1 with k_t:
ss = transition_int2.subs(kt1,kt)
print('In SS k is constant:')
display(ss)
# +
# And are the able to calculate the steady state. The numerator looks complicated is however only the product of (1+g)(1+n)(2+p):
kss = sm.solve(ss,kt)[0]
print('Steady-State k:')
display(kss)
# -
# ### **Steady-state calculation**
#
# In this part we will now set up functions in order to calculate the steady-state.
# +
# It is then straightforward to set up a steady-sate function:
ss_func = sm.lambdify((alpha,g,n,rho),kss)
# we define a function with which we can calcluate the steady state with flexible parameteres:
def ss_calc(alpha,g,n,rho):
result = ss_func(alpha,g,n,rho)
print(f'The steady state solved by the standard python function is: {result}')
# -
# with this widget one can choose the parameter and it yields the steady state capital stock
widgets.interact(
ss_calc,
alpha = widgets.FloatSlider(description="$\\alpha$", min=0.01, max=0.99, step=0.01, continuous_update=False,),
rho = widgets.FloatSlider(description="$\\rho$", min=0, max=0.99, step=0.01, continuous_update=False,),
g = widgets.FloatSlider(description="g", min=-0.99, max=0.99, step=0.01, continuous_update=False,),
n = widgets.FloatSlider(description="n", min=-0.99, max=0.99, step=0.01, continuous_update=False,),
)
# +
# we call the steady-state function with certain values as input as comparison:
# we choose parameters, which we think are reasonable today
result = ss_func(1/3,0.03,0.02,0.05)
print(f'The steady state solved by a standart python function is: {result}')
# +
# We can now solve this the problem numerically with optimize.root_scalar.
# However we have to make sure that we do not overwrite the already defined variables:
alpha_2 = 1/3
g_2 = 0.03
n_2 = 0.02
rho_2 =0.05
obj_kss = lambda kss: kss - ((kss**alpha_2*(1-alpha_2))/((1+g_2)*(1+n_2)*(2+rho_2)))
result2 = optimize.root_scalar(obj_kss,bracket=[0.1,100],method='brentq')
print(f'The steady state solved numerically with optimize.root_scalar is: {result2.root}')
print(f'Details are as followed:')
result2
# +
# We test if the two solutions are the same:
np.allclose(result, result2.root)
# -
# ### **Steady-state illustration**
#
# We now set up a plot, where the balanced growth path as well as the steady-state are shown. With several sliders one then can choose values for the different variabels and parameters.
# +
# We have to define first an equation corresponding to transition_int2 i.e. the corresponding path:
def kt1_func(kt, alpha, rho, n, g):
return ((1-alpha)*kt**alpha)/((2+rho)*(1+n)*(1+g))
kt = np.linspace(0,1,500)
# +
# And the plot function:
def plot_OLG(alpha,rho,g,n):
plt.figure(dpi=150)
plt.plot(kt,kt1_func(kt, alpha, rho, n, g), color = 'green')
plt.plot(range(10),range(10), color = 'blue')
plt.plot(ss_func(alpha,g,n,rho),ss_func(alpha,g,n,rho), marker='*',color='red')
plt.title('OLG model: Basic version')
plt.xlabel('$ k_t $')
plt.ylabel( '$ k_ {t + 1} $')
plt.xlim([0,1])
plt.ylim([0,1])
plt.legend(('BGP', '45\N{DEGREE SIGN}-line', 'steady state'), loc='upper left')
plt.grid()
sol = np.round(ss_func(alpha,g,n,rho),3)
text = 'Steady state at: ' + str(sol)
plt.text(0.64,0.95,text, fontsize='small', c='red', bbox=dict(facecolor='none', edgecolor='red'))
text = 'Here you can choose all variables and parameters of the basic OLG model!'
plt.text(0, 1.1, text, fontsize='x-small', c='black')
# Plotting plots, with interactive slider
widgets.interact(
plot_OLG,
alpha = widgets.FloatSlider(description="$\\alpha$", min=0.01, max=0.99, step=0.01, value=alpha_2, continuous_update=False,),
rho = widgets.FloatSlider(description="$\\rho$", min=-0.99, max=0.99, step=0.01, value=rho_2, continuous_update=False,),
g = widgets.FloatSlider(description="g", min=-0.99, max=0.99, step=0.01, value=g_2, continuous_update=False,),
n = widgets.FloatSlider(description="n", min=-0.99, max=0.99, step=0.01, value=n_2, continuous_update=False,),
)
# -
# ### Comparison of the steeady state depending on the parameters
# In the steady state the capital per unit of effective labor is constant and depends only on four parameters. In the this section, we compare some output variables like the consumption, utility and output depending on these input parameters.
def parameter_comparison(alpha, rho, n, g, alpha2, rho2, n2, g2):
# steady-state k
result = ss_func(alpha, rho, n, g)
result2 = ss_func(alpha2, rho2, n2, g2)
fig, axs = plt.subplots(4, 2, figsize=(15,15))
fig.suptitle('Steady State Comparison Depending on Different Parameters', fontsize=20)
# Technology
At = A*(1+g)**t
At2 = A*(1+g2)**t
axs[0, 0].plot(t, At, c='blue')
axs[0, 0].plot(t, At2, c='green')
axs[0, 0].set_title('Technology A')
axs[0, 0].grid()
axs[0, 0].legend(('Parameters 1', 'Parameters 2'), loc='upper left')
# Labor
Lt = L*(1+n)**t
Lt2 = L*(1+n2)**t
axs[0, 1].plot(t, Lt, c='blue')
axs[0, 1].plot(t, Lt2, c='green')
axs[0, 1].set_title('Labor L')
axs[0, 1].grid()
axs[0, 1].legend(('Parameters 1', 'Parameters 2'), loc='upper left')
# wage as defined above
w = np.empty(t.size)
w2 = np.empty(t.size)
for i in range(t.size):
w[i] = result**alpha *(1-alpha)
w2[i] = result2**alpha2 *(1-alpha2)
axs[1, 0].plot(t, w, c='blue')
axs[1, 0].plot(t, w2, c='green')
axs[1, 0].set_title('Wage Rate per Unit of Effective Labor')
axs[1, 0].grid()
axs[1, 0].legend(('Parameters 1', 'Parameters 2'), loc='center left')
# real interest rate as defined above
r = np.empty(t.size)
r2 = np.empty(t.size)
for i in range(t.size):
r[i] = result**(alpha-1) *(alpha)
r2[i] = result2**(alpha2-1) *(alpha2)
axs[1, 1].plot(t, r, c='blue')
axs[1, 1].plot(t, r2, c='green')
axs[1, 1].set_title('Real Interest Rate')
axs[1, 1].grid()
axs[1, 1].legend(('Parameters 1', 'Parameters 2'), loc='center left')
# consumption period 1 as defined above
C_t1 = A*(1+g)**t*w*(rho+1)/(rho+2)
C2_t1 = A*(1+g2)**t*w2*(rho2+1)/(rho2+2)
axs[2, 0].plot(t, C_t1, c='blue')
axs[2, 0].plot(t, C2_t1, c='green')
axs[2, 0].set_title('HH Consumption in Period 1')
axs[2, 0].grid()
axs[2, 0].legend(('Parameters 1', 'Parameters 2'), loc='upper left')
# consumption period 2 as defined above
C_t2 = A*(1+g)**t*w*(1+r)/(rho+2)
C2_t2 = A*(1+g2)**t*w2*(1+r2)/(rho2+2)
axs[2, 1].plot(t, C_t2, c='blue')
axs[2, 1].plot(t, C2_t2, c='green')
axs[2, 1].set_title('HH Consumption in Period 2')
axs[2, 1].grid()
axs[2, 1].legend(('Parameters 1', 'Parameters 2'), loc='upper left')
# utility
U = np.log(C_t1) + np.log(C_t2)/(rho+1)
U2 = np.log(C2_t1) + np.log(C2_t2)/(rho2+1)
axs[3, 0].plot(t, U, c='blue')
axs[3, 0].plot(t, U2, c='green')
axs[3, 0].set_title('Lifetime Utility of a HH')
axs[3, 0].grid()
axs[3, 0].legend(('Parameters 1', 'Parameters 2'), loc='upper left')
# production
Y = result**alpha*(A*(1+g)**t *(L*(1+n)**t))
Y2 = result2**alpha2*(A*(1+g2)**t *(L*(1+n2)**t))
axs[3, 1].plot(t, Y, c='blue')
axs[3, 1].plot(t, Y2, c='green')
axs[3, 1].set_title('Production of the Whole Economy Y')
axs[3, 1].grid()
axs[3, 1].legend(('Parameters 1', 'Parameters 2'), loc='upper left')
fig.text(0.5, 0.10, 'Time', ha='center', va='center', fontsize=16)
fig.text(0.08, 0.5, 'Variable of Interest', ha='center', va='center', rotation='vertical', fontsize=16)
A = 1
L = 1
t = np.linspace(0,100,100)
widgets.interact(
parameter_comparison,
alpha = widgets.FloatSlider(description="$\\alpha_1$", min=0.01, max=0.99, step=0.01, value=alpha_2, continuous_update=False,),
rho = widgets.FloatSlider(description="$\\rho_1$", min=-0.99, max=0.99, step=0.01, value= rho_2, continuous_update=False,),
g = widgets.FloatSlider(description="$g_1$", min=-0.99, max=0.99, step=0.01, continuous_update=False,),
n = widgets.FloatSlider(description="$n_1$", min=-0.99, max=0.99, step=0.01, continuous_update=False,),
alpha2 = widgets.FloatSlider(description="$\\alpha_2$", min=0.01, max=0.99, step=0.01, value=alpha_2, continuous_update=False,),
rho2 = widgets.FloatSlider(description="$\\rho_2$", min=-0.99, max=0.99, step=0.01, value=rho_2, continuous_update=False,),
g2 = widgets.FloatSlider(description="$g_2$", min=-0.99, max=0.99, step=0.01, value=g_2, continuous_update=False,),
n2 = widgets.FloatSlider(description="$n_2$", min=-0.99, max=0.99, step=0.01, value=n_2, continuous_update=False,),
)
# The figure shows that:
# - in the steady state the wage per unit of effective labor is constant, this statement is true for all variables measured in effective labor units
# - the real interest rate is constant, because it depends on the steady state capital stock per unit of effective labor, which is constant
# - the houshold consumption and lifetime utility depends crucially on the technology growth, g
# - the whole production of the economy grows with g and n
# ## **OLG with a government**
# <p> <br>
# The basic OLG from the first part is not that realistic. What is missing for example is goverment spending, which of course has to be financed somehow. The two basic means to finance government spending is either through taxes or otherwise by issuing debt-bonds. Both of them reduce the available income of some generations, and it is therefore interesting to study what effect such an implementation of a goverment has for the balanced-growth-path as well as the steady-state. As before, we do not change the setting of log-utility and Cobb-Douglas productivity. Furthermore, goverment spending G_t is also measured as spending per unit of effective labor (as well as taxes and bonds). What is obvious is that the government spending must be balanced, i.e.
#
# \begin{eqnarray*}
# G_t=T_t\\ G_t=b_t
# \end{eqnarray*}
#
# We first consider a labor tax, i.e.
#
# \begin{eqnarray*}
# C_{1t}=A_t(w_t-T_t)-S_t
# \end{eqnarray*}
#
# Savings are then:
#
# \begin{eqnarray*}
# S_t=\frac{1}{2+\rho}A_t(w_t-T_t)
# \end{eqnarray*}
#
# Ant the balanced growth path becomes:
#
# \begin{eqnarray*}
# k_{t+1}=\frac{(1-\alpha)k_t^\alpha-T_t}{(1+g)(1+n)(2+\rho)}
# \end{eqnarray*}
#
# With debt issuing, it is much more straightforward, as individuals buy government bonds instead of saving and the balanced growth path becomes therefore:
#
# \begin{eqnarray*}
# k_{t+1}=\frac{(1-\alpha)k_t^\alpha}{(1+g)(1+n)(2+\rho)}-b_{t+1}
# \end{eqnarray*}
#
# Let's analyse the effect with a nice plot:
# +
# We redefine the corresponding path steady-state functions:
kt = sm.symbols('k_t')
kt1 = sm.symbols('k_t+1')
alpha = sm.symbols('alpha')
T = sm.symbols('T_t')
bt1 = sm.symbols('b_t+1')
g = sm.symbols('g')
n = sm.symbols('n')
rho = sm.symbols('rho')
tax_transition = sm.Eq(kt1,((1-alpha)*kt**alpha-T)/((1+g)*(1+n)*(2+rho)))
bond_transition = sm.Eq(kt1,(((1-alpha)*kt**alpha)/((1+g)*(1+n)*(2+rho)))-bt1)
ss_tax = tax_transition.subs(kt1,kt)
# kss = sm.solve(ss_tax,kt)[0] # Try this command; we are not able to calculate the kss-formula as before!!!
# This means that we are not able to plot stars for the new steady states, which is no that bad as
# steady states are obvious when plotting a 45°-line.
# +
# We also redefine the corresponding path equations:
def kt1_no_gov_func(kt, alpha, rho, n, g):
return ((1-alpha)*kt**alpha)/((2+rho)*(1+n)*(1+g))
def kt1_with_tax_func(kt, alpha, rho, n, g, G): # With G as variable instead of T_t
return ((1-alpha)*kt**alpha-G)/((2+rho)*(1+n)*(1+g))
def kt1_with_bonds_func(kt, alpha, rho, n, g, G): # With G as variable instead of b_t+1
return ((1-alpha)*kt**alpha)/((2+rho)*(1+n)*(1+g))-G
kt = np.linspace(0,1,500)
# +
# And again the plot function:
def plot_OLG_with_G(alpha,rho,g,n,G):
plt.figure(dpi=150)
plt.plot(kt,kt1_no_gov_func(kt, alpha, rho, n, g))
plt.plot(kt,kt1_with_tax_func(kt, alpha, rho, n, g, G))
plt.plot(kt,kt1_with_bonds_func(kt, alpha, rho, n, g, G))
plt.plot(range(10),range(10),color='black')
plt.title('OLG model with government G = ' + str(G))
plt.xlabel('$ k_t $')
plt.ylabel('$ k_ {t + 1} $')
plt.xlim([0,1])
plt.ylim([0,1])
plt.legend(('G=0','Tax','Bonds','45\N{DEGREE SIGN}-line'), loc='upper left')
plt.grid()
text = 'Here you can also choose the size of government!'
plt.text(0, 1.1, text, fontsize='x-small', c='black')
# -
widgets.interact(
plot_OLG_with_G,
G = widgets.FloatSlider(description="G_size", min=0, max=0.25, step=0.01, value=0.1, continuous_update=False),
alpha = widgets.FloatSlider(description="$\\alpha$", min=0.01, max=0.99, step=0.01, value=alpha_2, continuous_update=False,),
rho = widgets.FloatSlider(description="$\\rho$", min=-0.99, max=0.99, step=0.01, value=rho_2, continuous_update=False,),
g = widgets.FloatSlider(description="g", min=-0.99, max=0.99, step=0.01, value=g_2, continuous_update=False,),
n = widgets.FloatSlider(description="n", min=-0.99, max=0.99, step=0.01, value=n_2, continuous_update=False,),
)
# As a result, we see that goverment spending i.e. its financing reduces the steady-state capital stock, however the effect of taxes is smaller than the effect of debt-issuing.
# +
# As above we can now solve this the problem numerically with optimize.root_scalar.
alpha = 1/3
g = 0.03
n = 0.02
rho =0.05
# As can be seen in the figure, the government spending can not be really big otherwise the model does not converge to a steady state.
G= 0.04
obj_kss_ = lambda kss: kss - ((1-alpha)*kss**alpha)/((2+rho)*(1+n)*(1+g))
obj_kss_tax = lambda kss: kss - ((1-alpha)*kss**alpha-G)/((2+rho)*(1+n)*(1+g)) # With G as variable instead of T_t
obj_kss_bond = lambda kss: kss - (((1-alpha)*kss**alpha)/((2+rho)*(1+n)*(1+g))-G) # With G as variable instead of b_t+1
result_ = optimize.root_scalar(obj_kss_,bracket=[0.1,100],method='brentq')
result_t = optimize.root_scalar(obj_kss_tax,bracket=[0.1,100],method='brentq')
result_b = optimize.root_scalar(obj_kss_bond,bracket=[0.1,100],method='brentq')
print(f'The steady state solved numerically with optimize.root_scalar is without government: {result_.root}')
print(f'The steady state solved numerically with optimize.root_scalar is with taxes: {result_t.root}')
print(f'The steady state solved numerically with optimize.root_scalar is with bonds: {result_b.root}')
# -
# ## Conclusion
#
# In our model project we analyzed a standard overlapping generations model. First, we solved the household optimization using sympy. Similarly, we solved the production side. Combining these two optimal behaviors, we are able to calculate the steady state. We then visualize the steady state and the evolution of the key variables in this steady state. Moreover, we make this visualization interactive, so we can directly observe the effect of the model parameter. In the last step, we introduce a government and analyze the steady state in this case. Unfortunately, we are unable to calculate a closed form solution for the steady state. But we can show that the effect of taxes is smaller than the effect of bonds.
#
| modelproject/OLG_model_swiss_python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PyCharm (oxford_uni)
# language: python
# name: pycharm-b1674269
# ---
# + pycharm={"name": "#%%\n"}
#
# *
# * *
# * *
# * *
# * *
# * *
# *
#
# March Twenty-Fifth, Two Thousand Twenty-One
# Red Queen
# <EMAIL>
#
# The digital humanities only looks like it's computational. Actually, it's data cleaning. Forever.
#
# Use me for cleaning Gutentag XML exports to get down to the nested paragraphs and return pandas dataframes
# that project found here: https://gutentag.sdsu.edu/
# + pycharm={"name": "#%%\n"}
from xml.etree.ElementTree import ElementTree
import pandas as pd
from html import unescape
from xml.etree import ElementTree
from nltk import sent_tokenize, re
import xmltodict
# -
# Read in XML file from Gutentag for paragraphs. Shelley and Dickens are sampled
# + pycharm={"name": "#%%\n"}
def xml_to_dataframe(f_path):
root = ElementTree.parse(f_path).getroot()
elements = root.getchildren()[0][-1] #You may have to play with this to isolate the element
f_dict = {'paragraph': []}
for b in elements:
if b.attrib.get('type', None) == 'chapter':
for t in b.getchildren():
try:
if len(t.text.strip()) > 30:
f_dict['paragraph'].append(t.text)
except:
continue
else:
try:
if len(b.text.strip()) > 30:
f_dict['paragraph'].append(b.text)
except:
continue #Extremely broad exceptions that may skip over large text portions; YMMV
df = pd.DataFrame.from_dict(f_dict)
return df
# -
# The odd case of parsing letters as XML. Austen letters are sampled
# + pycharm={"name": "#%%\n"}
def letters_to_dataframe(f_path):
elements = xmltodict.parse(open(f_path).read())
paragraph_groups = [t['body']['p'] for t in elements['TEI']['text']['group']['text']]
paragraph_objs = [item for sublist in paragraph_groups for item in sublist]
paragraphs = []
for item in paragraph_objs:
try:
if len(item['#text']) > 30:
paragraphs.append(item['#text'])
except:
continue
f_dict = {'paragraph': paragraphs}
df = pd.DataFrame.from_dict(f_dict)
return df
# -
# Define a function to feed in a text file and get back a dataframe; sampled with Lovelace's
# letters to <NAME>
# + pycharm={"name": "#%%\n"}
def text_to_dataframe(f_path):
file = open(f_path, 'rt', encoding='utf8')
text = file.read()
file.close()
sent = sent_tokenize(text)
df = pd.DataFrame(sent)
df.rename(columns={0:"paragraph"}, inplace=True)
return df
# + pycharm={"name": "#%%\n"}
#Get some dataframes
frankenstein_df = xml_to_dataframe('Frankenstein.xml')
print(frankenstein_df)
pickwick_df = xml_to_dataframe('Pickwick.xml')
#print(pickwick_df)
austen_df = letters_to_dataframe('AustenLetters.xml')
#print(austen_df)
lovelace_df = text_to_dataframe('Lovelace.txt')
#print(lovelace_df)
# + pycharm={"name": "#%%\n"}
# Define function to clean text
def clean(text):
text = unescape(text)
text = re.sub('[^a-zA-Z]', ' ', text)
text = ' '.join([w.lower() for w in text.split()])
return text
# -
# Clean up the dataframes and remove cruft.
# + pycharm={"name": "#%%\n"}
# Create column with cleaned text
frankenstein_df['cleaned_text'] = frankenstein_df.paragraph.map(lambda x: clean(x))
frankenstein_df = frankenstein_df['cleaned_text']
pickwick_df['cleaned_text'] = pickwick_df.paragraph.map(lambda x: clean(x))
pickwick_df = pickwick_df['cleaned_text']
austen_df['cleaned_text'] = austen_df.paragraph.map(lambda x: clean(x))
austen_df = austen_df['cleaned_text']
lovelace_df['cleaned_text'] = lovelace_df.paragraph.map(lambda x: clean(x))
lovelace_df = lovelace_df['cleaned_text']
# -
# How many paragraphs are in our dataframes?
# + pycharm={"name": "#%%\n"}
print(len(frankenstein_df))
print(len(pickwick_df))
print(len(austen_df))
print(len(lovelace_df))
| gutentag_xml_cleaner.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3-azureml
# kernelspec:
# display_name: Python 3.6 - AzureML
# language: python
# name: python3-azureml
# ---
# + [markdown] nteract={"transient": {"deleting": false}}
# # Preprocess the data with RAPIDS using GPUs
#
# In this notebook, you'll be using a subset of high-dimensional airline data: the Airline Service Quality Performance dataset, distributed by the U.S. Bureau of Transportation Statistics. 1987-2021. https://www.bts.dot.gov/browse-statistical-products-and-data/bts-publications/airline-service-quality-performance-234-time)
#
# This dataset is open source and provided on an ongoing basis by the U.S. Bureau of Transportation Statistics.
#
# Each month, the Bureau publishes a new csv file containing all flight information for the prior month. To train a robust machine learning model, you'd want to combine data over multiple years to use as a training dataset. In this exercise, you'll use data of only 10 days for illustration purposes. However, even when working with large amounts of data, the script should execute quickly as it uses cuDF to load and preprocess the data.
#
# In addition to the flight data, you'll also be downloading a file containing metadata and geo-coordinates of each airport and a file containing the code mappings for each airline. Airlines and airports rarely change, and as such, these files are static and do not change on a monthly basis. They do, however, contain information that we will later need to be mapped to the full airline dataset. (<NAME>. "airports.csv", distributed by OurAirports. August 2, 2021. https://ourairports.com/data/airports.csv)
# + [markdown] nteract={"transient": {"deleting": false}}
# ## Get environment variables
#
# Before you can submit the job, you have to get all necessary environment variables such as the workspace and environment.
# + gather={"logged": 1641821027915}
from azureml.core import Workspace
ws = Workspace.from_config()
# + gather={"logged": 1641821029553} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
from azureml.core import Environment
from azureml.core.runconfig import DockerConfiguration
rapidsai_env = Environment.get(workspace=ws, name="rapids-mlflow")
d_config = DockerConfiguration(arguments=['-it'])
# + [markdown] nteract={"transient": {"deleting": false}}
# ## Define the configuration and submit the run
#
# Now that you have defined all necessary variables, you can define the script run configuration and submit the run.
#
# **Warning!** Change the value of the compute_target variable to your compute cluster name before running the code below!
# + gather={"logged": 1641821043046} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
from azureml.core import ScriptRunConfig
src = ScriptRunConfig(source_directory='script',
compute_target="<your-compute-cluster>",
environment=rapidsai_env,
docker_runtime_config=d_config)
# + [markdown] nteract={"transient": {"deleting": false}}
# To learn what is done during preprocessing, explore the script `preprocess-rapids.py` in the `script` folder.
#
# The following cell will initiate the run. Note that first, the compute cluster has to scale up from 0 nodes. Once a node is available, it will execute the script. The execution of the script should be fast and you can see the execution time in the **Details** tab of the **Experiment** run afterwards.
# + gather={"logged": 1641821575025} jupyter={"outputs_hidden": true, "source_hidden": false} nteract={"transient": {"deleting": false}}
from azureml.core import Experiment
run = Experiment(ws,'preprocess-data').submit(src)
run.wait_for_completion(show_output=True)
# + [markdown] nteract={"transient": {"deleting": false}}
# You should get a notification in the Studio that a new run has started and is running.
#
# You can also navigate to the **Experiments** tab, and find the experiment `preprocess-data` there.
#
# Once it has finished running, have a look at the **Metrics** tab to learn how much data was processed. In the **Details** tab, you can see how long it took to run. You'll also find the processed data in the **Outputs+logs** in the **outputs** folder.
| Allfiles/Labs/01-preprocess-data/01-process-data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="W_tvPdyfA-BL"
# ##### Copyright 2019 The TensorFlow Authors.
# + cellView="form" colab={} colab_type="code" id="0O_LFhwSBCjm"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="9-3Pry4jh1-E"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/examples/blob/master/courses/udacity_intro_to_tensorflow_for_deep_learning/l06c01_tensorflow_hub_and_transfer_learning.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/examples/blob/master/courses/udacity_intro_to_tensorflow_for_deep_learning/l06c01_tensorflow_hub_and_transfer_learning.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="NxjpzKTvg_dd"
# # TensorFlow Hub and Transfer Learning
# + [markdown] colab_type="text" id="crU-iluJIEzw"
# [TensorFlow Hub](http://tensorflow.org/hub) is an online repository of already trained TensorFlow models that you can use.
# These models can either be used as is, or they can be used for Transfer Learning.
#
# Transfer learning is a process where you take an existing trained model, and extend it to do additional work. This involves leaving the bulk of the model unchanged, while adding and retraining the final layers, in order to get a different set of possible outputs.
#
# In this Colab we will do both.
#
# Here, you can see all the models available in [TensorFlow Module Hub](https://tfhub.dev/).
#
# ## Concepts that will be covered in this Colab
#
# 1. Use a TensorFlow Hub model for prediction.
# 2. Use a TensorFlow Hub model for Dogs vs. Cats dataset.
# 3. Do simple transfer learning with TensorFlow Hub.
#
# Before starting this Colab, you should reset the Colab environment by selecting `Runtime -> Reset all runtimes...` from menu above.
# + [markdown] colab_type="text" id="7RVsYZLEpEWs"
# # Imports
#
#
# + [markdown] colab_type="text" id="fL7DqCwbmfwi"
# This Colab will require us to use some things which are not yet in official releases of TensorFlow. So below, we're first installing a nightly version of TensorFlow as well as TensorFlow Hub.
#
# This will switch your installation of TensorFlow in Colab to this TensorFlow version. Once you are finished with this Colab, you should switch batch to the latest stable release of TensorFlow by doing selecting `Runtime -> Reset all runtimes...` in the menus above. This will reset the Colab environment to its original state.
# + colab={} colab_type="code" id="e3BXzUGabcI9"
# !pip install tf-nightly-gpu
# !pip install "tensorflow_hub==0.4.0"
# !pip install -U tensorflow_datasets
# + [markdown] colab_type="text" id="ZUCEcRdhnyWn"
# Some normal imports we've seen before. The new one is importing tensorflow_hub which was installed above, and which this Colab will make heavy use of.
# + colab={} colab_type="code" id="OGNpmn43C0O6"
from __future__ import absolute_import, division, print_function, unicode_literals
import matplotlib.pylab as plt
import tensorflow as tf
tf.logging.set_verbosity(tf.logging.ERROR)
tf.enable_eager_execution()
import tensorflow_hub as hub
import tensorflow_datasets as tfds
from tensorflow.keras import layers
# + [markdown] colab_type="text" id="s4YuF5HvpM1W"
# # Part 1: Use a TensorFlow Hub MobileNet for prediction
# + [markdown] colab_type="text" id="4Sh2sPc10V0b"
# In this part of the Colab, we'll take a trained model, load it into to Keras, and try it out.
#
# The model that we'll use is MobileNet v2 (but any model from [tf2 compatible image classifier url from tfhub.dev](https://tfhub.dev/s?q=tf2&module-type=image-classification) would work).
# + [markdown] colab_type="text" id="xEY_Ow5loN6q"
# ## Download the classifier
#
# Download the MobileNet model and create a Keras model from it.
# MobileNet is expecting images of 224 $\times$ 224 pixels, in 3 color channels (RGB).
# + colab={} colab_type="code" id="y_6bGjoPtzau"
CLASSIFIER_URL ="https://tfhub.dev/google/tf2-preview/mobilenet_v2/classification/2"
IMAGE_RES = 224
model = tf.keras.Sequential([
hub.KerasLayer(CLASSIFIER_URL, input_shape=(IMAGE_RES, IMAGE_RES, 3))
])
# + [markdown] colab_type="text" id="pwZXaoV0uXp2"
# ## Run it on a single image
# + [markdown] colab_type="text" id="TQItP1i55-di"
# MobileNet has been trained on the ImageNet dataset. ImageNet has 1000 different output classes, and one of them is military uniforms.
# Let's get an image containing a military uniform that is not part of ImageNet, and see if our model can predict that it is a military uniform.
# + colab={} colab_type="code" id="w5wDjXNjuXGD"
import numpy as np
import PIL.Image as Image
grace_hopper = tf.keras.utils.get_file('image.jpg','https://storage.googleapis.com/download.tensorflow.org/example_images/grace_hopper.jpg')
grace_hopper = Image.open(grace_hopper).resize((IMAGE_RES, IMAGE_RES))
grace_hopper
# + colab={} colab_type="code" id="BEmmBnGbLxPp"
grace_hopper = np.array(grace_hopper)/255.0
grace_hopper.shape
# + [markdown] colab_type="text" id="0Ic8OEEo2b73"
# Remember, models always want a batch of images to process. So here, we add a batch dimension, and pass the image to the model for prediction.
# + colab={} colab_type="code" id="EMquyn29v8q3"
result = model.predict(grace_hopper[np.newaxis, ...])
result.shape
# + [markdown] colab_type="text" id="NKzjqENF6jDF"
# The result is a 1001 element vector of logits, rating the probability of each class for the image.
#
# So the top class ID can be found with argmax. But how can we know what class this actually is and in particular if that class ID in the ImageNet dataset denotes a military uniform or something else?
# + colab={} colab_type="code" id="rgXb44vt6goJ"
predicted_class = np.argmax(result[0], axis=-1)
predicted_class
# + [markdown] colab_type="text" id="YrxLMajMoxkf"
# ## Decode the predictions
#
# To see what our predicted_class is in the ImageNet dataset, download the ImageNet labels and fetch the row that the model predicted.
# + colab={} colab_type="code" id="ij6SrDxcxzry"
labels_path = tf.keras.utils.get_file('ImageNetLabels.txt','https://storage.googleapis.com/download.tensorflow.org/data/ImageNetLabels.txt')
imagenet_labels = np.array(open(labels_path).read().splitlines())
plt.imshow(grace_hopper)
plt.axis('off')
predicted_class_name = imagenet_labels[predicted_class]
_ = plt.title("Prediction: " + predicted_class_name.title())
# + [markdown] colab_type="text" id="a6TNYYAM4u2-"
# Bingo. Our model correctly predicted miliatry uniform!
# + [markdown] colab_type="text" id="amfzqn1Oo7Om"
# # Part 2: Use a TensorFlow Hub models for the Cats vs. Dogs dataset
# + [markdown] colab_type="text" id="K-nIpVJ94xrw"
# Now we'll use the full MobileNet model and see how it can perform on the Dogs vs. Cats dataset.
# + [markdown] colab_type="text" id="Z93vvAdGxDMD"
# ## Dataset
#
# We can use TensorFlow Datasets to load the Dogs vs Cats dataset.
# + colab={} colab_type="code" id="DrIUV3V0xDL_"
splits = tfds.Split.ALL.subsplit(weighted=(80, 20))
splits, info = tfds.load('cats_vs_dogs', with_info=True, as_supervised=True, split = splits)
(train_examples, validation_examples) = splits
num_examples = info.splits['train'].num_examples
num_classes = info.features['label'].num_classes
# + [markdown] colab_type="text" id="UlFZ_hwjCLgS"
# The images in the Dogs vs. Cats dataset are not all the same size.
# + colab={} colab_type="code" id="W4lDPkn2cpWZ"
for i, example_image in enumerate(train_examples.take(3)):
print("Image {} shape: {}".format(i+1, example_image[0].shape))
# + [markdown] colab_type="text" id="mbgpD3E6gM2P"
# So we need to reformat all images to the resolution expected by MobileNet (224, 224)
#
# The `.repeat()` and `steps_per_epoch` here is not required, but saves ~15s per epoch, since the shuffle-buffer only has to cold-start once.
# + colab={} colab_type="code" id="we_ftzQxNf7e"
def format_image(image, label):
image = tf.image.resize(image, (IMAGE_RES, IMAGE_RES))/255.0
return image, label
BATCH_SIZE = 32
train_batches = train_examples.shuffle(num_examples//4).map(format_image).batch(BATCH_SIZE).prefetch(1)
validation_batches = validation_examples.map(format_image).batch(BATCH_SIZE).prefetch(1)
# + [markdown] colab_type="text" id="0gTN7M_GxDLx"
# ## Run the classifier on a batch of images
# + [markdown] colab_type="text" id="O3fvrZR8xDLv"
# Remember our `model` object is still the full MobileNet model trained on ImageNet, so it has 1000 possible output classes.
# ImageNet has a lot of dogs and cats in it, so let's see if it can predict the images in our Dogs vs. Cats dataset.
#
# + colab={} colab_type="code" id="kii_jWZYOn0B"
image_batch, label_batch = next(iter(train_batches.take(1)))
image_batch = image_batch.numpy()
label_batch = label_batch.numpy()
result_batch = model.predict(image_batch)
predicted_class_names = imagenet_labels[np.argmax(result_batch, axis=-1)]
predicted_class_names
# + [markdown] colab_type="text" id="QmvSWg9nxDLa"
# The labels seem to match names of Dogs and Cats. Let's now plot the images from our Dogs vs Cats dataset and put the ImageNet label next to them.
# + colab={} colab_type="code" id="IXTB22SpxDLP"
plt.figure(figsize=(10,9))
for n in range(30):
plt.subplot(6,5,n+1)
plt.imshow(image_batch[n])
plt.title(predicted_class_names[n])
plt.axis('off')
_ = plt.suptitle("ImageNet predictions")
# + [markdown] colab_type="text" id="JzV457OXreQP"
# # Part 3: Do simple transfer learning with TensorFlow Hub
#
# Let's now use TensorFlow Hub to do Transfer Learning.
#
# With transfer learning we reuse parts of an already trained model and change the final layer, or several layers, of the model, and then retrain those layers on our own dataset.
#
# In addition to complete models, TensorFlow Hub also distributes models without the last classification layer. These can be used to easily do transfer learning. We will continue using MobileNet v2 because in later parts of this course, we will take this model and deploy on a mobile device using [TensorFlow Lite](https://www.tensorflow.org/lite). Any [image feature vector url from tfhub.dev](https://tfhub.dev/s?module-type=image-feature-vector&q=tf2) would work here.
#
# We'll also continue to use the Dogs vs Cats dataset, so we will be able to compare the performance of this model against the ones we created from scratch earlier.
#
# Note that we're calling the partial model from TensorFlow Hub (without the final classification layer) a `feature_extractor`. The reasoning for this term is that it will take the input all the way to a layer containing a number of features. So it has done the bulk of the work in identifying the content of an image, except for creating the final probability distribution. That is, it has extracted the features of the image.
# + colab={} colab_type="code" id="5wB030nezBwI"
URL = "https://tfhub.dev/google/tf2-preview/mobilenet_v2/feature_vector/2"
feature_extractor = hub.KerasLayer(URL,
input_shape=(IMAGE_RES, IMAGE_RES,3))
# + [markdown] colab_type="text" id="pkSvAPvKOWg2"
# Let's run a batch of images through this, and see the final shape. 32 is the number of images, and 1280 is the number of neurons in the last layer of the partial model from TensorFlow Hub.
# + colab={} colab_type="code" id="Of7i-35F09ls"
feature_batch = feature_extractor(image_batch)
print(feature_batch.shape)
# + [markdown] colab_type="text" id="CtFmF7A5E4tk"
# Freeze the variables in the feature extractor layer, so that the training only modifies the final classifier layer.
# + colab={} colab_type="code" id="Jg5ar6rcE4H-"
feature_extractor.trainable = False
# + [markdown] colab_type="text" id="RPVeouTksO9q"
# ## Attach a classification head
#
# Now wrap the hub layer in a `tf.keras.Sequential` model, and add a new classification layer.
# + colab={} colab_type="code" id="mGcY27fY1q3Q"
model = tf.keras.Sequential([
feature_extractor,
layers.Dense(2, activation='softmax')
])
model.summary()
# + [markdown] colab_type="text" id="OHbXQqIquFxQ"
# ## Train the model
#
# We now train this model like any other, by first calling `compile` followed by `fit`.
# + colab={} colab_type="code" id="3n0Wb9ylKd8R"
model.compile(
optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
EPOCHS = 6
history = model.fit(train_batches,
epochs=EPOCHS,
validation_data=validation_batches)
# + [markdown] colab_type="text" id="76as-K8-vFQJ"
# You can see we get ~97% validation accuracy, which is absolutely awesome. This is a huge improvement over the model we created in the previous lesson, where we were able to get ~83% accuracy. The reason for this difference is that MobileNet was carefully designed over a long time by experts, then trained on a massive dataset (ImageNet).
#
# Although not equivalent to TensorFlow Hub, you can check out how to create MobileNet in Keras [here](https://github.com/keras-team/keras-applications/blob/master/keras_applications/mobilenet.py).
#
# Let's plot the training and validation accuracy/loss graphs.
# + colab={} colab_type="code" id="d28dhbFpr98b"
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs_range = range(EPOCHS)
plt.figure(figsize=(8, 8))
plt.subplot(1, 2, 1)
plt.plot(epochs_range, acc, label='Training Accuracy')
plt.plot(epochs_range, val_acc, label='Validation Accuracy')
plt.legend(loc='lower right')
plt.title('Training and Validation Accuracy')
plt.subplot(1, 2, 2)
plt.plot(epochs_range, loss, label='Training Loss')
plt.plot(epochs_range, val_loss, label='Validation Loss')
plt.legend(loc='upper right')
plt.title('Training and Validation Loss')
plt.show()
# + [markdown] colab_type="text" id="5zmoDisGvNye"
# What is a bit curious here is that validation performance is better than training performance, right from the start to the end of execution.
#
# One reason for this is that validation performance is measured at the end of the epoch, but training performance is the average values across the epoch.
#
# The bigger reason though is that we're reusing a large part of MobileNet which is already trained on Dogs and Cats images. While doing training, the network is still performing image augmentation on the training images, but not on the validation dataset. This means the training images may be harder to classify compared to the normal images in the validation dataset.
# + [markdown] colab_type="text" id="kb__ZN8uFn-D"
# ## Check the predictions
#
# To redo the plot from before, first get the ordered list of class names.
# + colab={} colab_type="code" id="W_Zvg2i0fzJu"
class_names = np.array(info.features['label'].names)
class_names
# + [markdown] colab_type="text" id="4Olg6MsNGJTL"
# Run the image batch through the model and comvert the indices to class names.
# + colab={} colab_type="code" id="fCLVCpEjJ_VP"
predicted_batch = model.predict(image_batch)
predicted_batch = tf.squeeze(predicted_batch).numpy()
predicted_ids = np.argmax(predicted_batch, axis=-1)
predicted_class_names = class_names[predicted_ids]
predicted_class_names
# + [markdown] colab_type="text" id="CkGbZxl9GZs-"
# Let's look at the true labels and predicted ones.
# + colab={} colab_type="code" id="nL9IhOmGI5dJ"
print("Labels: ", label_batch)
print("Predicted labels: ", predicted_ids)
# + colab={} colab_type="code" id="wC_AYRJU9NQe"
plt.figure(figsize=(10,9))
for n in range(30):
plt.subplot(6,5,n+1)
plt.imshow(image_batch[n])
color = "blue" if predicted_ids[n] == label_batch[n] else "red"
plt.title(predicted_class_names[n].title(), color=color)
plt.axis('off')
_ = plt.suptitle("Model predictions (blue: correct, red: incorrect)")
| courses/udacity_intro_to_tensorflow_for_deep_learning/l06c01_tensorflow_hub_and_transfer_learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ```
# Copyright 2021 IBM Corporation
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ```
# # Logistic Regression on Epsilon Dataset
#
# ## Background
#
# This is a synthetic dataset from the [PASCAL Large Scale Learning Challenge](https://www.k4all.org/project/large-scale-learning-challenge/). This challenge is concerned with the scalability and efficiency of existing ML approaches with respect to computational, memory or communication resources, e.g. resulting from a high algorithmic complexity, from the size or dimensionality of the data set, and from the trade-off between distributed resolution and communication costs.
#
# ## Source
#
# In this example, we download the dataset from the [UCI Machine Learning Repository](https://archive.ics.uci.edu/ml/datasets.php).
#
# ## Goal
# The goal of this notebook is to illustrate how Snap ML can accelerate training of a logistic regression model on this dataset.
#
# ## Code
# cd ../../
CACHE_DIR='cache-dir'
import numpy as np
import time
from datasets import Epsilon
from sklearn.linear_model import LogisticRegression
from snapml import LogisticRegression as SnapLogisticRegression
from sklearn.metrics import roc_auc_score as score
dataset = Epsilon(cache_dir=CACHE_DIR)
X_train, X_test, y_train, y_test = dataset.get_train_test_split()
print("Number of examples: %d" % (X_train.shape[0]))
print("Number of features: %d" % (X_train.shape[1]))
print("Number of classes: %d" % (len(np.unique(y_train))))
model = LogisticRegression(fit_intercept=False, n_jobs=4)
t0 = time.time()
model.fit(X_train, y_train)
t_fit_sklearn = time.time()-t0
score_sklearn = score(y_test, model.predict_proba(X_test)[:,1])
print("Training time (sklearn): %6.2f seconds" % (t_fit_sklearn))
print("ROC AUC score (sklearn): %.4f" % (score_sklearn))
model = SnapLogisticRegression(fit_intercept=False, n_jobs=4)
t0 = time.time()
model.fit(X_train, y_train)
t_fit_snapml = time.time()-t0
score_snapml = score(y_test, model.predict_proba(X_test)[:,1])
print("Training time (snapml): %6.2f seconds" % (t_fit_snapml))
print("ROC AUC score (snapml): %.4f" % (score_snapml))
speed_up = t_fit_sklearn/t_fit_snapml
score_diff = (score_snapml-score_sklearn)/score_sklearn
print("Speed-up: %.1f x" % (speed_up))
print("Relative diff. in score: %.4f" % (score_diff))
# ## Disclaimer
#
# Performance results always depend on the hardware and software environment.
#
# Information regarding the environment that was used to run this notebook are provided below:
import utils
environment = utils.get_environment()
for k,v in environment.items():
print("%15s: %s" % (k, v))
# ## Record Statistics
#
# Finally, we record the enviroment and performance statistics for analysis outside of this standalone notebook.
import scrapbook as sb
sb.glue("result", {
'dataset': dataset.name,
'n_examples_train': X_train.shape[0],
'n_examples_test': X_test.shape[0],
'n_features': X_train.shape[1],
'n_classes': len(np.unique(y_train)),
'model': type(model).__name__,
'score': score.__name__,
't_fit_sklearn': t_fit_sklearn,
'score_sklearn': score_sklearn,
't_fit_snapml': t_fit_snapml,
'score_snapml': score_snapml,
'score_diff': score_diff,
'speed_up': speed_up,
**environment,
})
| examples/training/logistic_regression/example_epsilon.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.1 64-bit
# language: python
# name: python38164bit9a8e3b63a08644a087459c5617ed8408
# ---
# # Predict CO2 Emissions
#
# <hr>
#
# ### About
#
# > Predicting CO2 emission of a car can be useful in building models of cars that will produce less CO2 which will then reduce the harmful effects of excessive CO2 in the environment.
# #### Importing Module
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn import linear_model, preprocessing
from sklearn.preprocessing import StandardScaler, LabelEncoder
from sklearn.metrics import mean_squared_error, r2_score
from sklearn.model_selection import KFold, learning_curve, cross_val_score
# #### Loading the dataset
df = pd.read_csv('data.csv')
df.head()
df.describe()
# <hr>
#
# ## Feature Selection
# #### Pearson Correlation
plt.figure(figsize=(12,10))
cor = df.corr()
sns.heatmap(cor, annot=True, cmap=plt.cm.YlGn)
plt.show()
df.corr()['CO2EMISSIONS'].sort_values()
# #### Categorical Data
#
# Performing One Hot Encoding
# +
# Select only categorical variables
category_df = df.select_dtypes('object')
# One hot encode the variables
dummy_df = pd.get_dummies(category_df)
# Put the grade back in the dataframe
# dummy_df['Average Grades'] = df['Average Grades']
dummy_df['CO2EMISSIONS'] = df['CO2EMISSIONS']
# Find correlations with grade
# dummy_df.corr()['Average Grades'].sort_values()
dummy_df.corr()['CO2EMISSIONS'].sort_values()
# -
# #### ENGINESIZE
sns.scatterplot(x='ENGINESIZE', y='CO2EMISSIONS', data=df)
# #### CYLINDERS
sns.scatterplot(x='CYLINDERS', y='CO2EMISSIONS', data=df)
# #### FUELCONSUMPTION_CITY
sns.scatterplot(x='FUELCONSUMPTION_CITY', y='CO2EMISSIONS', data=df)
# #### FUELCONSUMPTION_HWY
sns.scatterplot(x='FUELCONSUMPTION_HWY', y='CO2EMISSIONS', data=df)
# #### FUELCONSUMPTION_COMB
sns.scatterplot(x='FUELCONSUMPTION_COMB', y='CO2EMISSIONS', data=df)
# #### Pearson Correlation
plt.figure(figsize=(12,10))
cor = df.corr()
sns.heatmap(cor, annot=True, cmap=plt.cm.YlOrRd)
plt.show()
# Since 'FUELCONSUMPTION_CITY', 'FUELCONSUMPTION_HWY', 'FUELCONSUMPTION_COMB' are highly correlated so we will use only one of these features
# #### Final Features
df = df[['ENGINESIZE','CYLINDERS','FUELCONSUMPTION_COMB','CO2EMISSIONS']]
df.head()
# <hr>
#
# ## Creating the model
kf = KFold(n_splits=10, shuffle=True)
# #### Shuffling the dataframe
_df = df.sample(frac=1, random_state=999)
# #### Splitting the dataset
# +
features = ['ENGINESIZE', 'CYLINDERS', 'FUELCONSUMPTION_COMB']
target = ['CO2EMISSIONS']
train_dataset = _df[:900]
X_train = train_dataset[features]
y_train = train_dataset[target]
test_dataset = _df[901:1067]
X_test = test_dataset[features]
y_test = test_dataset[target]
# -
# #### Feature Scaling
# +
scaler = StandardScaler()
X_train = scaler.fit_transform(np.asanyarray(X_train))
y_train = np.asanyarray(y_train)
X_test = scaler.fit_transform(np.asanyarray(X_test))
y_test = np.asanyarray(y_test)
# -
# #### Cross Validation
# +
scoring = 'r2'
score = cross_val_score(linear_model.LinearRegression(), X_train, y_train, cv=kf, scoring=scoring)
score.mean()
# -
# #### Learning Curve
# +
_sizes = [i for i in range(1, 720, 20)]
train_sizes = np.array([_sizes]) # Relative sizes
scoring = 'neg_mean_squared_error'
lr = linear_model.LinearRegression()
train_sizes_abs, train_scores, cv_scores = learning_curve(lr, X_train, y_train, train_sizes=train_sizes, cv=kf, scoring=scoring)
# +
train_scores_mean = []
for row in train_scores:
_mean = row.mean()
train_scores_mean.append(_mean)
cv_scores_mean = []
for row in cv_scores:
_mean = row.mean()
cv_scores_mean.append(_mean)
train_scores_mean = -np.array(train_scores_mean)
cv_scores_mean = -np.array(cv_scores_mean)
print(train_scores_mean)
print()
print(cv_scores_mean)
# +
plt.plot(train_sizes_abs, train_scores_mean, label='Train')
plt.plot(train_sizes_abs, cv_scores_mean, label='Cross Validation')
plt.legend()
# -
# #### Fitting the model
model = lr.fit(X_train, y_train)
# #### Optimal Parameters
# +
coefficient = model.coef_
intercept = model.intercept_
print("Coefficient: ", coefficient)
print("Intercept: ", model.intercept_)
# -
# #### Predictions
y_test_pred = model.predict(X_test)
y_test_pred.astype('int64')
y_test
# #### Evaluation
# +
rms_error = mean_squared_error(y_test, y_test_pred)
r2_score_value = r2_score(y_test, y_test_pred)
print(f"Root mean squared error: {rms_error}")
print(f"R2-score: {r2_score_value}")
# -
# #### Pipeline
from sklearn.pipeline import Pipeline
# +
scaling = ('scale', StandardScaler())
model = ('model', linear_model.LinearRegression())
# Steps in the pipeline
steps = [scaling, model]
pipe = Pipeline(steps=steps)
# Fiitting the model
model = pipe.fit(X_train, y_train)
# Out-Of-Sample Forecast
y_test_pred = model.predict(X_test)
# Evaluation
rms_error = mean_squared_error(y_test, y_test_pred, squared=False)
r2_score_value = r2_score(y_test, y_test_pred)
print(f"Root mean squared error: {rms_error}")
print(f"R2-score: {r2_score_value}")
# -
# #### Saving the model
# +
from joblib import dump
dump(model, 'model.joblib')
# -
# #### Visualizing our prediction against actual values
# +
f, (ax1, ax2) = plt.subplots(nrows=1, ncols=2, figsize=(13, 5))
ax1.plot(np.arange(len(y_test)), y_test, label='Actual')
ax2.plot(np.arange(len(y_test_pred)), y_test_pred, label='Prediction')
ax1.legend()
ax2.legend()
f, ax3 = plt.subplots(nrows=1, ncols=1, figsize=(13, 5))
ax3.plot(np.arange(len(y_test)), y_test, label='Actual')
ax3.plot(np.arange(len(y_test_pred)), y_test_pred, label='Prediction')
ax3.legend()
# -
# <hr>
| venv/src/notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Chapter 7: List
numbers = [1, 2, 3, 4, 3, 44, 3, 3, 5]
names = ['Amin', 'Sahar', 'Armin', 'Narges', 'Raana', 'Milad']
lst = [20, 'Mahdi', [20, 'Mahdi']]
empty = []
s = names[0]
s
s[1]
lst[2][1][0]
# lists are mutable
names[0] = 'Reza'
names
for name in enumerate(names):
print(name)
lst1 = ['A','B','C']
lst2 = ['a','b','c']
for name in zip(lst2,lst1):
print(name)
zip(lst2,lst1)
lst = list(range(10))
lst
lst / 2
# +
new_lst = []
for item in lst:
new_lst.append(item/2)
new_lst
# -
lst
# +
for i in range(len(lst)):
lst[i] = lst[i]/2
lst
# -
# repeat it 2 times
i = 0
for number in numbers:
numbers[i] = number * 2
i += 1
lst3 = [number / 2 for number in numbers]
lst3
numbers
new_nums = numbers / 2
# instead you can use:
new_numbers = [numbers[i]/2 for i in range(len(numbers))]
new_numbers
# ### List operations
first = [10, 20, 30]
second = [40, 50, 60]
first + second
first * 3
# ### List Slices
numbers[0:3], numbers[-1], numbers[:4], numbers[4:]
# ### List Methods
names
dir(names)
lst = list(range(10))
lst1 = [item/2 for item in lst if item%2==0]
lst, lst1
lst.append(lst1)
lst
help(names.sort())
lst2 = [2, 25, 3, 55, 71, 1]
lst2.sort()
lst2
names, names.sort(), names
# #### append()
names.append('Amin')
names
# #### sort()
names.sort(reverse = True)
names
# #### extend()
t1 = ['a', 'b', 'c']
t2 = ['d', 'e']
t1.extend(t2)
print(t1)
# #### pop()
t = ['a', 'b', 'c']
x = t.pop(0)
print("t is: ", t)
print("x is: ", x)
# #### del()
t = ['a', 'b', 'c']
del t[1]
print("t is: ", t)
t = ['a', 'b', 'c', 'd', 'e', 'f']
del t[1:5]
print('t is: ', t)
t = ['a', 'b', 'c']
t.remove('b')
print('t is: ', t)
# #### Some Functions
nums = [3, 41, 12, 9, 74, 15]
print('length of nums is = ', len(nums))
print('maximum of nums is = ', max(nums))
print('minimum of nums is = ', min(nums))
print('summation of nums is = ', sum(nums))
print('average of nums is = ', sum(nums)/len(nums))
# EXE
total = 0
count = 0
average = None
while (True):
inp = input('Enter a number: ')
if inp == 'done':
break
value = float(inp)
total = total + value
count = count + 1
average = total / count
print('Average:', average)
numlist = list()
average = None
while (True):
inp = input('Enter a number: ')
if inp == 'done': break
value = float(inp)
numlist.append(value)
average = sum(numlist) / len(numlist)
print('Average:', average)
# ### Lists and Strings
s = 'spam'
t = list(s)
print(t)
s = 'This is <NAME>, from Prata Technology.'
t = s.split()
print(t)
delimiter = ','
s.split(delimiter)
delimiter = '$'
s1 = 'This$is$Amin$ Oroji$,$ from $Prata$ Technology'
s1.split(delimiter)
t = ['This', 'is', 'Amin', 'Oroji']
delimiter = ' '
delimiter.join(t)
t = ['This', 'is', 'Amin', 'Oroji']
delimiter = ' * '
delimiter.join(t)
s = ' Amin Oroji '
t = s.split()
print(t)
delimiter = ' '
delimiter.join(t)
while True:
inp = input()
if inp == 'done':
break
t = inp.split()
print(' '.join(t))
# ### Objects and Values
a = 'banana'
b = 'banana'
a is b
a = [1, 2, 3]
b = [1, 2, 3]
a is b
type(a)
# two lists are ***equivalent***, because they have the same elements, but ***not identical***, because they are **not** the ***same object***.
# ### Aliasing
a = [1, 2, 3]
b = a
a is b
# An object with more than one reference has more than one name, so we say that the object is aliased.
a[0] = 25
a
b
# If the aliased object is mutable, changes made with one alias affect the other:
b[0] = 17
print(a)
# #### copy()
c = a.copy()
c
a is c
c[0] = 12
print('a = ', a)
print('c = ', c)
| Python_CH7.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Question1
#
# Create a function that takes a string and returns a string in which each character is repeated once. Examples
#
# double_char("String") ➞ "SSttrriinngg"
#
# double_char("Hello World!") ➞ "HHeelllloo WWoorrlldd!!"
#
# double_char("1234!_ ") ➞ "11223344!!__ "
#
def double_char(str):
return ''.join([i*2 for i in str])
print(double_char("String"))
print(double_char("Hello World!"))
print(double_char("1234!_ "))
# ### Question2
#
# Create a function that reverses a boolean value and returns the string "boolean expected" if another variable type is given. Examples
#
# reverse(True) ➞ False
#
# reverse(False) ➞ True
#
# reverse(0) ➞ "boolean expected"
#
# reverse(None) ➞ "boolean expected"
#
# +
def reverse(arg=None):
return not arg if type(arg) == bool else "boolean expected"
print(reverse(True)) # False
print(reverse(False)) # True
print(reverse(0)) # "boolean expected"
print(reverse(None)) # "boolean expected"
# -
# ### Question3
#
# Create a function that returns the thickness (in meters) of a piece of paper after folding it n number of times. The paper starts off with a thickness of 0.5mm. Examples
#
# num_layers(1) ➞ "0.001m"
#
# Paper folded once is 1mm (equal to 0.001m)
#
# num_layers(4) ➞ "0.008m"
#
# Paper folded 4 times is 8mm (equal to 0.008m)
#
# num_layers(21) ➞ "1048.576m"
#
# Paper folded 21 times is 1048576mm (equal to 1048.576m)
# +
def num_layers(n):
sum_mm, sum_m = 0.5, 0.0005
for i in range(n):
sum_mm *= 2
sum_m *= 2
print(f"Paper folded {n} times is {str(round(sum_mm))+'mm'} (equal to {str(sum_m)+'m'})")
num_layers(21)
# -
# ### Question4
#
# Create a function that takes a single string as argument and returns an ordered list containing the indices of all capital letters in the string. Examples
#
# index_of_caps("eDaBiT") ➞ [1, 3, 5]
#
# index_of_caps("eQuINoX") ➞ [1, 3, 4, 6]
#
# index_of_caps("determine") ➞ []
#
# index_of_caps("STRIKE") ➞ [0, 1, 2, 3, 4, 5]
#
# index_of_caps("sUn") ➞ [1]
# +
def index_of_caps(str):
indexes = []
for i in range(len(str)):
if str[i].isupper():
indexes.append(i)
return indexes
print(index_of_caps('eDaBiT'))
print(index_of_caps('eQuINoX'))
print(index_of_caps('determine'))
print(index_of_caps('STRIKE'))
print(index_of_caps('sUn'))
# -
# ### Question5
#
# Using list comprehensions, create a function that finds all even numbers from 1 to the given number. Examples
#
# find_even_nums(8) ➞ [2, 4, 6, 8]
#
# find_even_nums(4) ➞ [2, 4]
#
# find_even_nums(2) ➞ [2]
# +
def find_even_nums(n):
even = [i for i in range(1,n+1) if i%2 == 0]
return even
print(find_even_nums(8))
print(find_even_nums(4))
print(find_even_nums(2))
# -
| Programming_Assingment19.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
#
# # Linear Equations
#
#
# ## Outline
#
# 1. Setup
# 2. Special cases
# 3. L-U Factorization
# 4. Pivoting
# 5. Benchmarking
# 6. Ill conditioning
# 7. Iterative methods
# 8. Resources
#
#
# ## Setup
# The linear equation is the most elementary problem that arises in computational economic analysis. In a linear equation, an $n \times n$ matrix $A$ and an n-vector $b$ are given, and one must compute the $n$-vector $x$ that satisfies
# $Ax = b$.
#
# Linear equations arise naturally in many economic applications such as
#
# - Linear multicommodity market equilibrium models
# - Finite-state financial market models
# - Markov chain models
# - Ordinary least squares
#
# They more commonly arise indirectly from numerical solution to nonlinear and functional equations:
#
# - Nonlinear multicommodity market models
# - Multiperson static game models
# - Dynamic optimization models
# - Rational expectations models
#
# Applications often require the repeated solution of very large linear equation systems. In these situations, issues regarding speed, storage requirements, and preciseness of the solution of such equations can arise.
# +
import pandas as pd
import numpy as np
from linear_algorithms import backward_substitution
from linear_algorithms import forward_substitution
from linear_algorithms import gauss_seidel
from linear_algorithms import solve
from linear_plots import plot_operation_count
from linear_problems import get_ill_problem_1
from linear_problems import get_inverse_demand_problem
# -
#
# ## Special cases
#
#
# We can start with some special cases to develop a basic understanding for the core building blocks for more complicated settings. Let's start with the case of a lower triangular matrix $A$, where we can solve the linear equation by a simple backward or forward substitution. Let's consider the following setup.
#
# $$
# A = \begin{bmatrix}
# a_{11} & 0 & 0 \\
# a_{21} & a_{22} & 0 \\
# a_{31} & a_{32} & a_{33} \\
# \end{bmatrix}
# $$
#
# Consider an algorithmic implementation of forward-substitution as an example.
#
# $$
# x_i = \left ( b_i - \sum_{j=1}^{i-1} a_{ij}x_j \right )/a_{ii}
# $$
??forward_substitution
# +
def test_problem():
A = np.tril(np.random.normal(size=(3, 3)))
x = np.random.normal(size=3)
b = np.matmul(A, x)
return A, b, x
for _ in range(10):
A, b, x_true = test_problem()
x_solve = forward_substitution(A, b)
np.testing.assert_almost_equal(x_solve, x_true)
# -
# ### _Questions_
#
# * How can we make the test code more generic and sample test problems of different dimensions?
# * Is there a way to control the randomness in the test function?
# * Is there software out there that allows to automate parts of the testing?
#
# If we have an upper triangular matrix, we can use backward substitution to solve the linear system
??backward_substitution
# ### _Exercise_
#
#
# * Implement the same testing setup as above the backward-substitution function.
# We can now build on these two functions to tackle more complex tasks. This is a good example on how to develop scientific software step-by-step ensuring that each component is well tested before integrating into more involved settings.
#
# ## L-U Factorization
#
#
# Most linear equations encountered in practice, however, do not have a triangular $A$ matrix. Doolittle and Crout have shown that any matrix $A$ can be decomposed into the product of a (row-permuted) lower and upper triangular matrix $L$ and $U$, respectively $A=L \times U$ using **Gaussian elimination**. We will not look into the Gaussian elimination algorithm, but there is an example application in our textbook where you can follow along step by step. The L-U algorithm is designed to decompose the $A$ matrix into the product of lower and upper triangular matrices, allowing the linear equation to be solved using a combination of backward and forward substitution.
#
# Here are the two core steps:
#
# * Factorization phase
#
# \begin{align*}
# A = LU
# \end{align*}
#
# * Solution phase:
#
# \begin{align*}
# Ax = (LU)x=L(Ux) = b,
# \end{align*}
#
# where we solve $Ly = b$ using forward-substitution and $Ux=y$ using backward-substitution.
#
# Adding to this the two building blocks we developed earlier `forward_substitution` and `backward_substitution`, we can now write a quite generic function to solve systems of linear equations.
??solve
# Let's see if this is actually working.
A = np.array([[3, 1], [1, 2]])
x_true = np.array([9, 8])
b = A @ x_true
x_solve = solve(A, b)
np.testing.assert_almost_equal(x_true, x_solve)
# ## Pivoting
#
#
# Rounding error can cause serious error when solving linear equations. Let's consider the following example, where $\epsilon$ is a tiny number.
#
# $$
# \begin{bmatrix} \epsilon & 1\\ 1 & 1 \end{bmatrix} \times \left[ \begin{array}{c} x_1 \\ x_2 \end{array} \right] = \left[\begin{array}{c} 1 \\ 2 \end{array} \right]
# $$
#
#
# It is easy to verify that the right solution is
#
# $$
# \begin{aligned}
# x_1 & = \frac{1}{1 - \epsilon} \\
# x_2 & = \frac{1 - 2 \epsilon}{1 - \epsilon}
# \end{aligned}
# $$
#
# and thus $x_1$ is slightly more than one and $x_2$ is slightly less than one. To solve the system using Gaussian elimination we need to add $-1/\epsilon$ times the first row to the second row. We end up with
#
#
# $$
# \begin{bmatrix} \epsilon & 1 \\ 0 & 1 - \frac{1}{\epsilon} \end{bmatrix} \times \left[ \begin{array}{c} x_1 \\ x_2 \end{array} \right] = \left[ \begin{array}{c} 1 \\ 2 - \frac{1}{\epsilon} \end{array} \right],
# $$
#
# which we can then solve recursively.
#
# $$
# \begin{aligned}
# x_2 & = \frac{2 - 1/\epsilon}{1 - 1/\epsilon} \\
# x_1 & = \frac{1 - x_2}{\epsilon}
# \end{aligned}
# $$
# Let's translate this into code.
eps = 1e-17
A = np.array([[eps, 1], [1, 1]])
b = np.array([1, 2])
# We can now use our solution algorithm.
solve(A, b)
# But we have to realize that the results are grossly inaccurate. What happened? Is there any hope to apply `numpy`'s routine?
np.linalg.solve(A, b)
# This algorithm does automatically check whether such rounding errors can be avoided by simply changing the order of rows. This is called **pivoting** and changes the recursive solution to
#
# $$
# \begin{aligned}
# x_2 & = \frac{1 - 2\epsilon}{1 - \epsilon} \\
# x_1 & = 2 - x_2
# \end{aligned}
# $$
#
# which can be solved more accurately. Our implementation also solves the modified problem well.
A = np.array([[1, 1], [eps, 1]])
b = np.array([2, 1])
solve(A, b)
# Building your own numerical routines is usually the only way to really understand the algorithms and learn about all the potential pitfalls. However, the default should be to rely on battle-tested production code. For linear algebra there are numerous well established libraries available. Building your own numerical routines is usually the only way to really understand the algorithms and learn about all the potential pitfalls. However, the default should be to rely on battle-tested production code. For linear algebra there are numerous well established libraries available.
#
# ## Benchmarking
#
#
# How does solving a system of linear equations by an $L-U$ decomposition compare to other alternatives of solving the system of linear equations.
plot_operation_count()
# The right setup for your numerical needs depends on your particular problem. For example, this trade-off looks very different if you have to solve numerous linear equations that only differ in $b$ but not $A$. In this case you only need to compute the inverse once.
# ### _Exercise_
#
#
# * Set up a benchmarking exercise that compares the time to solution for the two approaches for $m=\{1, 100\}$ and $n = \{50, 100\}$, where $n$ denotes the number of linear equations and $m$ the number of repeated solutions.
#
# ## Ill Conditioning
#
# Some linear equations are inherently difficult to solve accurately on a computer. This difficulty occurs when the A matrix is structured in such a way that a small perturbation $\delta b$ in the data vector $b$ induces a large change $\delta x$ in the solution vector $x$. In such cases the linear equation or, more generally, the $A$ matrix is said to be **ill conditioned**.
#
# One measure of ill conditioning in a linear equation Ax = b is the “elasticity” of the solution vector $x$ with respect to the data vector $b$
#
# $$
# \epsilon = \sup_{||\delta b|| > 0} \frac{||\delta x|| / ||x||}{||\delta b|| / ||b||}
# $$
#
# The elasticity gives the maximum percentage change in the size of the solution vector $x$ induced by a $1$ percent change in the size of the data vector $b$. If the elasticity is large, then small errors in the computer representation of the data vector $b$ can produce large errors in the computed solution vector x. Equivalently, the computed solution $x$ will have far fewer significant digits than the data vector $b$.
#
# In practice, the elasticity is estimated using the condition number of the matrix $A$, which for invertible $A$ is defined by $\kappa \equiv ||A|| \cdot ||A^{-1} ||$. The condition number is always greater than or equal to one. Numerical analysts often use the rough rule of thumb that for each power of $10$ in the condition number, one significant digit is lost in the computed solution vector $x$. Thus, if $A$ has a condition number of $1,000$, the computed solution vector $x$ will have about three fewer significant digits than the data vector $b$.
#
# Let's look at an example, where the solution vector is all ones but but the linear equation is notoriously ill-conditioned.
??get_ill_problem_1
# How does the solution error depend on the condition number in this setting.
# +
rslt = dict((("Condition", []), ("Error", []), ("Dimension", [])))
grid = [5, 10, 15, 25]
for n in grid:
A, b, x_true = get_ill_problem_1(n)
x_solve = np.linalg.solve(A, b)
rslt["Condition"].append(np.linalg.cond(A))
rslt["Error"].append(np.linalg.norm(x_solve - x_true, 1))
rslt["Dimension"].append(n)
# -
pd.DataFrame.from_dict(rslt)
# There is a more general lesson here. Always be skeptical about the quality of your numerical results. See these two papers for an exploratory analysis of econometric software packages and nonliner optimization. Yes, they are rather old and much progress has been made, but the general points remain valid.
#
# * <NAME>., and <NAME>. (2003). [Verifying the Solution from a Nonlinear Solver: A Case Study](https://www.aeaweb.org/articles?id=10.1257/000282803322157133). *American Economic Review*, 93 (3): 873-892.
#
# * <NAME>., and <NAME>. (1999). [The Numerical Reliability of Econometric Software](https://www.aeaweb.org/articles?id=10.1257/jel.37.2.633). *Journal of Economic Literature*, 37 (2): 633-665.
# ### _Exercise_
#
# Let's consider the following example as well, which is taken from Johansson (2015, p.131).
#
# $$
# \left[\begin{array}{c} 1 \\ 2 \end{array}\right] = \begin{bmatrix} 1 & \sqrt{p}\\ 1 & \frac{1}{\sqrt{p}} \end{bmatrix} \times \left[ \begin{array}{c} x_1 \\ x_2 \end{array} \right]
# $$
#
# This system is singular for $p=1$ and for $p$ in the vicinity of one is ill-conditioned.
#
# * Create two plots that show the condition number and the error between the analytic and numerical solution.
# +
el = np.linspace(0.8,1.2, 20)
rslt = dict((("Condition", []), ("Error", []), ("P", [])))
for p in el:
A = np.array([[1, np.sqrt(p)], [1, 1/np.sqrt(p)]])
b = np.array([1,2])
x_solve = np.linalg.solve(A, b)
rslt["Condition"].append(np.linalg.cond(A))
rslt["Error"].append(np.linalg.norm(x_solve - x_true, 1)) #determine x_solve?
rslt["Dimension"].append(p)
fig, ax =plt.subplots
# -
def test_2():
"""Solution to exercise 2."""
def benchmarking_alternatives():
def tic():
return time.time()
def toc(t):
return time.time() - t
print(
"{:^5} {:^5} {:^11} {:^11} \n{}".format(
"m", "n", "np.solve(A,b)", "dot(inv(A), b)", "-" * 40
)
)
for m, n in product([1, 100], [50, 500]):
a = np.random.rand(n, n)
b = np.random.rand(n, 1)
tt = tic()
[np.linalg.solve(a, b) for _ in range(m)]
f1 = 100 * toc(tt)
tt = tic()
a_inv = np.linalg.inv(a)
[np.dot(a_inv, b) for _ in range(m)]
f2 = 100 * toc(tt)
print(f" {m:3} {n:3} {f1:11.2f} {f2:11.2f}")
benchmarking_alternatives()
#
# ## Iterative methods
#
# Algorithms based on Gaussian elimination are called exact or, more properly, direct methods because they would generate exact solutions for the linear equation $Ax = b$ after a finite number of operations, if not for rounding error. Such methods are ideal for moderately sized linear equations but may be impractical for large ones. Other methods, called iterative methods, can often be used to solve large linear equations more efficiently if the $A$ matrix is sparse, that is, if $A$ is composed mostly of zero entries. Iterative methods are designed to generate a sequence of increasingly accurate approximations to the solution of a linear equation, but they generally do not yield an exact solution after a prescribed number of steps, even in theory.
#
#
# The most widely used iterative methods for solving a linear equation $Ax = b$ are developed by choosing an easily invertible matrix $Q$ and writing the linear equation in the equivalent form
#
# $$
# Qx = b + (Q - A)x
# $$
#
# or
#
# $$
# x = Q^{-1} b + (I - Q^{-1} A)x
# $$
#
# This form of the linear equation suggests the iteration rule
#
# $$
# x^{k+1}\leftarrow Q^{-1} b + (I - Q^{-1} A)x^{k}
# $$
#
# which, if convergent, must converge to a solution of the linear equation. Ideally, the so-called splitting matrix $Q$ will satisfy two criteria. First, $Q^{-1}b$ and $Q^{-1} A$ should be relatively easy to compute. This criterion is met if $Q$ is either diagonal or triangular. There are two popular approaches:
#
# * The **Gauss-Seidel** method sets $Q$ equal to the upper triangular matrix formed from the upper triangular elements of $A$.
#
# * The **Gauss-Jacobi** method sets $Q$ equal to the diagonal matrix formed from the diagonal entries of $A$.
??gauss_seidel
# ### _Exercise_
#
# * Implement the Gauss-Jacobi method.
from linear_solutions_tests import gauss_jacobi # noqa: E402
# Let's conclude with an economic application as outlined in Judd (1998). Suppose we have the following inverse demand function $p = 10 - q$ and the following supply curve $q = p / 2 +1$. Equilibrium is where supply equals demand and thus we need to solve the following linear system.
#
# $$
# \left[\begin{array}{c} 10 \\ -2 \end{array}\right] = \begin{bmatrix} 1 & 1\\ 1 & -2\end{bmatrix} \times \left[ \begin{array}{c} p \\ q \end{array} \right]
# $$
A, b, x_true = get_inverse_demand_problem()
# Now we can compre the two solution approaches and make sure that they in fact give the same result.
x_seidel = gauss_seidel(A, b)
x_jacobi = gauss_jacobi(A, b)
np.testing.assert_almost_equal(x_seidel, x_jacobi)
#
# ## Resources
#
# #### Software
#
# * **The PARDISO Solver Project**: https://www.pardiso-project.org
#
# * **LAPACK — Linear Algebra PACKage**: http://www.netlib.org/lapack
#
# #### References
#
# - <NAME>. *Numerical Python: scientific computing and data science applications with NumPy, SciPy and Matplotlib*. Apress, 2018.
#
# - <NAME>, <NAME>, <NAME>, and <NAME>. *Numerical recipes: The art of scientific computing*. Cambridge University Press, 1986.
#
| labs/linear_equations/notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import psi4
import pdft
from cubeprop import Cube
import pandas as pd
import qcelemental as qc
import numpy as np
import matplotlib.pyplot as plt
# -
# # PDFT Examples when target $n(r)$ is available
# --------------------------------------------------------------------------------------
# # <center>$Be_2$ Closed-Shell</center>
# --------------------------------------------------------------------------------------
# +
#First Fragment
f1_geometry = psi4.geometry("""
0 1
@Be 0.0 0.0 0.0
Be 0.0 0.0 4.52
symmetry c1
unit bohr
""")
f1_geometry.set_name("Beryllium 1")
#Second Fragment
f2_geometry = psi4.geometry("""
0 1
Be 0.0 0.0 0.0
@Be 0.0 0.0 4.52
symmetry c1
unit bohr
""")
f2_geometry.set_name("Beryllium 2")
#Full Molecule
mol_geometry = psi4.geometry("""
0 1
Be 0.0 0.0 0.0
Be 0.0 0.0 4.52
symmetry c1
unit bohr
""")
mol_geometry.set_name("Be2")
#Psi4 Options:
psi4.set_options({'DFT_SPHERICAL_POINTS': 110,
'DFT_RADIAL_POINTS': 5,
'REFERENCE' : 'RKS'})
psi4.set_options({'cubeprop_tasks' : ['density'],
'cubic_grid_spacing' : [0.1, 0.1, 0.1]})
energy, wfn = psi4.energy("SVWN/aug-cc-pVDZ", molecule=mol_geometry, return_wfn=True)
#Make fragment calculations:
f1 = pdft.Molecule(f1_geometry, "aug-cc-pVDZ", "SVWN")
f2 = pdft.Molecule(f2_geometry, "aug-cc-pVDZ", "SVWN")
mol = pdft.Molecule(mol_geometry, "aug-cc-pVDZ", "SVWN")
#Start a pdft systemm, and perform calculation to find vp
be2 = pdft.Embedding([f1, f2], mol)
vp = be2.find_vp_response(0.001, maxiter=40, atol=4e-5)
# -
vp_plot = Cube(wfn)
vp_plot.plot_matrix(vp,1,35)
mol.get_plot()
be2.get_energies()
# --------------------------------------------------------------------------------------
# # <center> $Li_2$ Open-Shell</center>
# --------------------------------------------------------------------------------------
# +
#First Fragment
f1_geometry = psi4.geometry("""
0 2
@Li 0.0 0.0 0.0
Li 0.0 0.0 4.52
symmetry c1
unit bohr
""")
f1_geometry.set_name("Lithium 1")
#Second Fragment
f2_geometry = psi4.geometry("""
0 2
Li 0.0 0.0 0.0
@Li 0.0 0.0 4.52
symmetry c1
unit bohr
""")
f2_geometry.set_name("Lithium 2")
#Full Molecule
mol_geometry = psi4.geometry("""
0 1
Li 0.0 0.0 0.0
Li 0.0 0.0 4.52
symmetry c1
unit bohr
""")
mol_geometry.set_name("Li2")
#Psi4 Options:
psi4.set_options({'DFT_SPHERICAL_POINTS': 110,
'DFT_RADIAL_POINTS': 5,
'REFERENCE' : 'UKS'})
psi4.set_options({'cubeprop_tasks' : ['density'],
'cubic_grid_spacing' : [0.1, 0.1, 0.1]})
energy_2, wfn_2 = psi4.energy("SVWN/cc-pVDZ", molecule=mol_geometry, return_wfn=True)
#Make fragment calculations:
f1 = pdft.U_Molecule(f1_geometry, "cc-pVDZ", "SVWN")
f2 = pdft.U_Molecule(f2_geometry, "cc-pVDZ", "SVWN")
mol = pdft.U_Molecule(mol_geometry, "cc-pVDZ", "SVWN")
energy_2, wfn_2 = psi4.energy("SVWN/cc-pVDZ", molecule=mol_geometry, return_wfn=True)
#Start a pdft systemm, and perform calculation to find vp
li2 = pdft.U_Embedding([f1, f2], mol)
vp = li2.find_vp(0.001, maxiter=9, atol=1e-5)
# -
vp_plot = Cube(wfn_2)
vp_plot.plot_matrix(vp[2],1,35)
li2.get_energies()
# --------------------------------------------------------------------------------------
# # <center> Water Dimer </center>
# --------------------------------------------------------------------------------------
# +
#First Fragment
f1_geometry = psi4.geometry("""
0 1
@O -1.464 0.099 0.300
@H -1.956 0.624 -0.340
@H -1.797 -0.799 0.206
O 1.369 0.146 -0.395
H 1.894 0.486 0.335
H 0.451 0.165 -0.083
symmetry c1
unit bohr
""")
f1_geometry.set_name("H20_1")
#Second Fragment
f2_geometry = psi4.geometry("""
0 1
O -1.464 0.099 0.300
H -1.956 0.624 -0.340
H -1.797 -0.799 0.206
@O 1.369 0.146 -0.395
@H 1.894 0.486 0.335
@H 0.451 0.165 -0.083
symmetry c1
unit bohr
""")
f2_geometry.set_name("H20_2")
#Full Molecule
mol_geometry = psi4.geometry("""
0 1
O -1.464 0.099 0.300
H -1.956 0.624 -0.340
H -1.797 -0.799 0.206
O 1.369 0.146 -0.395
H 1.894 0.486 0.335
H 0.451 0.165 -0.083
symmetry c1
unit bohr
""")
mol_geometry.set_name("Water Dimer")
#Psi4 Options:
psi4.set_options({'DFT_SPHERICAL_POINTS': 110,
'DFT_RADIAL_POINTS': 5,
'REFERENCE' : 'RKS'})
psi4.set_options({'cubeprop_tasks' : ['density'],
'cubic_grid_spacing' : [0.1, 0.1, 0.1]})
energy_3, wfn_3 = psi4.energy("SVWN/cc-pVDZ", molecule=mol_geometry, return_wfn=True)
#Make fragment calculations:
f1 = pdft.Molecule(f1_geometry, "cc-pVDZ", "SVWN")
f2 = pdft.Molecule(f2_geometry, "cc-pVDZ", "SVWN")
mol = pdft.Molecule(mol_geometry, "cc-pVDZ", "SVWN")
#Start a pdft systemm, and perform calculation to find vp
h20_2 = pdft.Embedding([f1, f2], mol)
vp = h20_2.find_vp(0.001, maxiter=5, atol=1e-5)
# -
h20_2.get_energies()
vp_plot = Cube(wfn_3)
vp_plot.plot_matrix(vp, 2,60)
| pdft/pdft_examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:xtalmd] *
# language: python
# name: conda-env-xtalmd-py
# ---
# ## Analysis xtal minimizations
#
# Description: Brief analysis of results from minimizing xtal structures with two different force fields (Sage, Parsley) and three different minimization strategies. The three minimization strategies were:
#
# 1.) Minimize box paramters and position vectors together ("All-in-one")
#
# 2.) Minimize box parameters and positions vectors in alternating fashion
#
# 3.) First do 1.), then do 2.)
# +
MAIN="/nfs/data_onsager/projects/xtalmd/xtalmd-scripts/examples/OFFBenchmark"
success_code_list = list()
with open(f"{MAIN}/success_xml.txt", "r") as fopen:
for line in fopen:
line = line.replace("\n", "")
line = line.replace("CIF/", "")
line = line.replace(".cif", "")
if len(line) > 0:
success_code_list.append(line)
success_xml = 0
failed_xml = 0
success_pdb = 0
failed_pdb = 0
with open(f"{MAIN}/success_xml.txt", "r") as fopen:
success_xml = len(fopen.readlines())-1
with open(f"{MAIN}/failed_xml.txt", "r") as fopen:
failed_xml = len(fopen.readlines())-1
with open(f"{MAIN}/success_pdb.txt", "r") as fopen:
success_pdb = len(fopen.readlines())-1
with open(f"{MAIN}/failed_pdb.txt", "r") as fopen:
failed_pdb = len(fopen.readlines())-1
print(f"Successful xml: {success_xml}")
print(f"Failed xml: {failed_xml}")
print(f"Successful pdb: {success_pdb}")
print(f"Failed pdb: {failed_pdb}")
# -
# From 293 CIF files, 232 xtal structures were successfully build and an RDKIT object was generated for molecule in the xtal structure. For 178 of these 232 xtal structures, sage ff parameters were obtained. Failing to obtain ff parameters were due to various reasons, for instance no ff parameter found, no hydrogen present, stereochemistry not clear. However these failures were not well documented at this point.
# +
import os
import numpy as np
from openmm import app
def get_final_ene(path):
if not os.path.exists(path):
return None, None
data=np.loadtxt(path, delimiter=',')
if data.ndim == 1:
final_ene = data[1]
start_ene = data[1]
else:
final_ene = data[-1,1]
start_ene = data[0,1]
return final_ene, start_ene
def get_rmsd(pdb_path_1, pdb_path_2):
if not os.path.exists(pdb_path_1):
return None
if not os.path.exists(pdb_path_2):
return None
pos1 = app.PDBFile(pdb_path_1).getPositions(asNumpy=True)
pos2 = app.PDBFile(pdb_path_2).getPositions(asNumpy=True)
topology = app.PDBFile(pdb_path_1).getTopology()
non_H_idxs = list()
for atom_idx, atom in enumerate(topology.atoms()):
if atom.element.atomic_number != 1:
non_H_idxs.append(atom_idx)
diff = np.linalg.norm(
pos1[non_H_idxs]-pos2[non_H_idxs],
axis=1
)
rmsd = np.sqrt(
np.mean(
diff**2
)
)
return rmsd
def retrieve_data(ff_name, query_code_list):
code_list = list()
ene_list = list()
rmsd_list = list()
for code in query_code_list:
### ================================= ###
### Simultaneous minimization pos/box ###
### ================================= ###
ene_min, ene_start = get_final_ene(
f"{MAIN}/{ff_name}_min/{code}.csv"
)
rmsd_min = get_rmsd(
f"{MAIN}/{ff_name}_xml/{code}.pdb",
f"{MAIN}/{ff_name}_min/{code}.pdb"
)
### ================================ ###
### Alternating minimization pos/box ###
### ================================ ###
ene_min_alternating, _ = get_final_ene(
f"{MAIN}/{ff_name}_min_alternating/{code}.csv"
)
rmsd_min_alternating = get_rmsd(
f"{MAIN}/{ff_name}_xml/{code}.pdb",
f"{MAIN}/{ff_name}_min_alternating/{code}.pdb",
)
### ================================================ ###
### Subsequent simultaneous/alternating minimization ###
### ================================================ ###
ene_min_combo, _ = get_final_ene(
f"{MAIN}/{ff_name}_min_combo/{code}.csv"
)
rmsd_min_combo = get_rmsd(
f"{MAIN}/{ff_name}_xml/{code}.pdb",
f"{MAIN}/{ff_name}_min_combo/{code}.pdb",
)
if any(
[
ene_min == None,
ene_min_alternating == None,
ene_min_combo == None,
]
):
print(f"Could not find {ff_name} minimum energy for {code}")
continue
code_list.append(code)
ene_list.append(
[
ene_min - ene_start,
ene_min_alternating - ene_start,
ene_min_combo - ene_start
]
)
rmsd_list.append(
[
rmsd_min,
rmsd_min_alternating,
rmsd_min_combo
]
)
return code_list, ene_list, rmsd_list
# -
parsley_code_list, parsley_ene_list, parsley_rmsd_list = retrieve_data("parsley", success_code_list)
sage_code_list, sage_ene_list, sage_rmsd_list = retrieve_data("sage", success_code_list)
print(f"{len(parsley_code_list)} succesful minimizations for Parsley.")
print(f"{len(sage_code_list)} succesful minimizations for Sage.")
# From the 178 xtal structures with ff parameters found, 165 xtal structures could be successfully minimized using Parsley and 168 were successfully minimized using Sage.
# +
import matplotlib.pyplot as plt
parsley_ene_list = np.array(parsley_ene_list)
parsley_rmsd_list = np.array(parsley_rmsd_list)
plt.hist(
parsley_rmsd_list[:,0],
density=False,
bins=20
)
plt.xlim(0,0.25)
plt.ylim(0,60)
plt.xlabel("RMSD [nm]")
plt.ylabel("Counts")
plt.title("PARSLEY - All-in-one minimization")
plt.show()
plt.hist(
parsley_rmsd_list[:,1],
density=False,
bins=20
)
plt.xlim(0,0.25)
plt.ylim(0,60)
plt.xlabel("RMSD [nm]")
plt.ylabel("Counts")
plt.title("PARSLEY - Alternating minimization")
plt.show()
plt.hist(
parsley_rmsd_list[:,2],
density=False,
bins=20
)
plt.xlim(0,0.25)
plt.ylim(0,60)
plt.xlabel("RMSD [nm]")
plt.ylabel("Counts")
plt.title("PARSLEY - All-in-one, then alternating minimization")
plt.show()
# +
import matplotlib.pyplot as plt
sage_ene_list = np.array(sage_ene_list)
sage_rmsd_list = np.array(sage_rmsd_list)
plt.hist(
sage_rmsd_list[:,0],
density=False,
bins=20
)
plt.xlim(0,0.25)
plt.ylim(0,60)
plt.xlabel("RMSD [nm]")
plt.ylabel("Counts")
plt.title("SAGE - All-in-one minimization")
plt.show()
plt.hist(
sage_rmsd_list[:,1],
density=False,
bins=20
)
plt.xlim(0,0.25)
plt.ylim(0,60)
plt.xlabel("RMSD [nm]")
plt.ylabel("Counts")
plt.title("SAGE - Alternating minimization")
plt.show()
plt.hist(
sage_rmsd_list[:,2],
density=False,
bins=20
)
plt.xlim(0,0.25)
plt.ylim(0,60)
plt.xlabel("RMSD [nm]")
plt.ylabel("Counts")
plt.title("SAGE - All-in-one, then alternating minimization")
plt.show()
# -
# Overall it seems that "All-in-one" minimization did not perform as well as the other two. In many cases (not documented how many) the BFGS minimizer stopped after only few iterations with the warning "Desired error not necessarily achieved due to precision loss.". This potentially solvable by carefully playing with the parameters of the algorithm.
#
# The alternate routes involving the alternating minimization worked better here.
| examples/OFFBenchmark/OFFBenchmark-analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <table width = "100%">
# <tr style="background-color:white;">
# <!-- QWorld Logo -->
# <td style="text-align:left;width:200px;">
# <a href="https://qworld.net/" target="_blank"><img src="../images/QWorld.png"> </a></td>
# <td style="text-align:right;vertical-align:bottom;font-size:16px;">
# Prepared by <a href="https://gitlab.com/AkashNarayanan" target="_blank"> <NAME></a></td>
# </table>
# <hr>
# So far we have learned about how to use the Ocean SDK tools. Now let's use those tools to formulate Binary Quadratic Models for some combinatorial optimization problems. In this notebook, we will learn how to formulate BQM for the Maximum Cut problem.
#
# # BQM Formulation of the Maximum Cut Problem
#
# To briefly recall, the goal of the maximum cut problem is to partition a set of vertices of a graph into two disjoint sets such that the number of edges that are cut by the partition is maximized.
#
# The QUBO objective function for a graph with edge set E is
#
# $$\min \sum_{(i, j) \in E} (-x_i - x_j + 2x_ix_j)$$
#
# The Ising objective function for a graph with edge set E is
#
# $$\min \sum_{(i, j) \in E} s_i s_j$$
#
# `dwave-networkx` package in the Ocean SDK has implementations of graph-theory algorithms for some combinatorial optimization problems like Maximum Cut, Graph Colouring, Traveling Salesman, etc. You can find out more details about the available algorithms [here](https://docs.ocean.dwavesys.com/en/stable/docs_dnx/reference/algorithms/index.html).
#
# There are two implemented algorithms `maximum_cut` and `weighted_maximum_cut` for solving the Maximum Cut problem.
#
# We can either use these already implemented algorithms or solve by formulating the problem as a QUBO or Ising from scratch. We will look at both the ways this problem can be solved.
# # Built-in Algorithm
# +
import networkx as nx
G = nx.Graph([(1,2),(1,3),(2,4),(3,4),(3,5),(4,5)])
nx.draw(G, with_labels=True, font_weight='bold')
# +
import dimod
from dimod.reference.samplers import ExactSolver
import dwave_networkx as dnx
sampler = ExactSolver()
cut = dnx.maximum_cut(G, sampler)
print(cut)
# -
# # Formulating QUBO from Scratch
# <div class="alert alert-block alert-danger">
# The below code snippet is from D-Wave's GitHub repository. This is put here for the sake of discussion during the meeting. It will be replaced with some other example later on.
# </div>
# +
from collections import defaultdict
import networkx as nx
from dimod.reference.samplers import ExactSolver
import matplotlib
matplotlib.use("agg")
from matplotlib import pyplot as plt
# ------- Set up our graph -------
# Create empty graph
G = nx.Graph()
# Add edges to the graph (also adds nodes)
G.add_edges_from([(1,2),(1,3),(2,4),(3,4),(3,5),(4,5)])
# ------- Set up our QUBO dictionary -------
# Initialize our Q matrix
Q = defaultdict(int)
# Update Q matrix for every edge in the graph
for i, j in G.edges:
Q[(i,i)]+= -1
Q[(j,j)]+= -1
Q[(i,j)]+= 2
sampler = ExactSolver()
response = sampler.sample_qubo(Q)
lut = response.first.sample
# Interpret best result in terms of nodes and edges
S0 = [node for node in G.nodes if not lut[node]]
S1 = [node for node in G.nodes if lut[node]]
cut_edges = [(u, v) for u, v in G.edges if lut[u]!=lut[v]]
uncut_edges = [(u, v) for u, v in G.edges if lut[u]==lut[v]]
# Display best result
pos = nx.spring_layout(G)
nx.draw_networkx_nodes(G, pos, nodelist=S0, node_color='r')
nx.draw_networkx_nodes(G, pos, nodelist=S1, node_color='c')
nx.draw_networkx_edges(G, pos, edgelist=cut_edges, style='dashdot', alpha=0.5, width=3)
nx.draw_networkx_edges(G, pos, edgelist=uncut_edges, style='solid', width=3)
nx.draw_networkx_labels(G, pos)
| notebooks/BQM_Maximum_Cut.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + raw_mimetype="text/restructuredtext" active=""
# .. _nb_pso:
# -
# .. meta::
# :description: An implementation of the famous Particle Swarm Optimization (PSO) algorithm which is inspired by the behavior of the movement of particles represented by their position and velocity. Each particle is updated considering the cognitive and social behavior in a swarm.
# .. meta::
# :keywords: Particle Swarm Optimization, Nature-inspired Algorithm, Single-objective Optimization, Python
# # Particle Swarm Optimization (PSO)
# Particle Swarm Optimization was proposed in 1995 by <NAME> Eberhart <cite data-cite="pso"></cite> based on the simulating of social behavior. The algorithm uses a *swarm* of particles to guide its search. Each particle has a velocity and is influenced by locally and globally best-found solutions. Many different implementations have been proposed in the past and, therefore, it is quite difficult to refer to THE correct implementation of PSO. However, the general concepts shall be explained in the following.
#
# Given the following variables:
#
# - $X_{d}^{(i)}$ d-th coordinate of i-th particle's position
# - $V_{d}^{(i)}$ d-th coordinate of i-th particle's velocity
# - $\omega$ Inertia weight
# - $P_{d}^{(i)}$ d-th coordinate of i-th particle's *personal* best
# - $G_{d}^{(i)}$ d-th coordinate of the globally (sometimes also only locally) best solution found
# - $c_1$ and $c_2$ Two weight values to balance exploiting the particle's best $P_{d}^{(i)}$ and swarm's best $G_{d}^{(i)}$
# - $r_1$ and $r_2$ Two random values being create for the velocity update
#
# The velocity update is given by:
# \begin{equation}
# V_{d}^{(i)} = \omega \, V_{d}^{(i)} \;+\; c_1 \, r_1 \, \left(P_{d}^{(i)} - X_{d}^{(i)}\right) \;+\; c_2 \, r_2 \, \left(G_{d}^{(i)} - X_{d}^{(i)}\right)
# \end{equation}
# The corresponding position value is then updated by:
# \begin{equation}
# X_{d}^{(i)} = X_{d}^{(i)} + V_{d}^{(i)}
# \end{equation}
# The social behavior is incorporated by using the *globally* (or locally) best-found solution in the swarm for the velocity update. Besides the social behavior, the swarm's cognitive behavior is determined by the particle's *personal* best solution found.
# The cognitive and social components need to be well balanced to ensure that the algorithm performs well on a variety of optimization problems.
# Thus, some effort has been made to determine suitable values for $c_1$ and $c_2$. In **pymoo** both values are updated as proposed in <cite data-cite="pso_adapative"></cite>. Our implementation deviates in some implementation details (e.g. fuzzy state change) but follows the general principles proposed in the paper.
# +
from pymoo.algorithms.so_pso import PSO, PSOAnimation
from pymoo.factory import Ackley
from pymoo.optimize import minimize
problem = Ackley()
algorithm = PSO(max_velocity_rate=0.025)
res = minimize(problem,
algorithm,
callback=PSOAnimation(fname="pso.mp4"),
seed=1,
save_history=True,
verbose=False)
print("Best solution found: \nX = %s\nF = %s" % (res.X, res.F))
# -
# Assuming you have our third-party library `pyrecorder` installed you can create an animation for a two-dimensional problem easily. Below we provide the code to observe the behavior of the swarm. We have reduced to set the maximum velocity to `max_velocity_rate=0.025` for illustration purposes. Otherwise, the algorithm converges even quicker to the global optimum of the `Ackley` function with two variables.
# <div style='text-align:center'>
# <video width="600" height="450" controls>
# <source src="http://pymoo.org/animations/pso.mp4" type="video/mp4">
# </video>
# </div>
# In general, the PSO algorithm can be used by execute the following code. For the available parameters please see the API description below.
# + code="algorithms/usage_pso.py"
from pymoo.algorithms.so_pso import PSO
from pymoo.factory import Rastrigin
from pymoo.optimize import minimize
problem = Rastrigin()
algorithm = PSO()
res = minimize(problem,
algorithm,
seed=1,
verbose=False)
print("Best solution found: \nX = %s\nF = %s" % (res.X, res.F))
# -
# ### API
# + raw_mimetype="text/restructuredtext" active=""
# .. autoclass:: pymoo.algorithms.so_pso.PSO
# :noindex:
# :no-undoc-members:
| doc/source/algorithms/pso.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Py3-basic
# language: python
# name: python3
# ---
# # Active features in each voxel
# * First, we cluster each feature using K-means clustering to 2 clusters, to find out where the feature is active in.
# * Then, we get the ratio of active features in each voxel.
import numpy as np
# +
'''Clustering each feature to 2 clusters to find the active regions'''
def cluster_festures_separately(features_arr):
'''
This function clusters each feature of the input features array (samples * features) separately
using K-means clustering and returns an array of labeled features (samples * features).
Args:
features_arr: numpy array
The numpy array of features (samples * features).
Returns:
labels_arr: numpy array
The numpy array of labeled features (samples * features).
'''
# Making an array to store all labels
labels_arr = np.zeros_like(features_arr)
for feature_n in range(features_arr.shape[1]):
# Selecting the feature
feature = features_arr[:, feature_n].reshape(-1,1)
# Running clustering
n_clusters = 2
kmeans = KMeans(n_clusters = n_clusters, random_state = 0)
kmeans.fit_predict(feature)
labels_arr[:, feature_n] = kmeans.labels_
return labels_arr
# +
# Loading the features
features_arr = np.load('files/SFT_100features.npy')
n_features = features_arr.shape[1]
# Clustering the features separately
labels_arr = cluster_festures_separately(features_arr)
# List of counts
counts_list = []
for i in range(labels_arr.shape[0]):
ratio = np.sum(labels_arr[i,:]) / n_features
counts_list.append(ratio)
| Notebooks/02_analyses/Fig4_Active_Featues.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # A concrete example of the case for nested functions: Function scoping
#
#
# Calling a function from within another function is really useful.
#
# In cases where the calling function simply imlements a more general from of the called function,
# defining the called function within the calling function allows for cleaner more robust codebase
# and API.
# ## The gcd example
# Consider a function for computing the greatest common divisor of an abitrary number of integers.
# A quick online search will reveal several algos for computing the gcd of 2 ints.
#
# One implementation may involve defining a function for computing the gcd of 2 ints "t_gcd" and another
# "gcd" that successievly applies the 2 int solution on an abitrary number of ints.
#
# t_gcd(a, b) -> n
# gcd(a, b, c, ..., j) -> t_gcd(...(t_gcd(t_gcd(a, b), c), ...), j) -> n
#
# It is perfectly possible and reasonable to define t_gcd and gcd seperately and call t_gcd from inside gcd.
# However, since gcd will work for 2 or more numbers, allowing t_gcd to exisit within the global scope or
# anywhere outside the scope of gcd can lead to confusion for users of the code and even the developer.
#
# Binding t_gcd within the scope of gcd means it is not possible to acess it from any where else, ergo there'd
# be one and only one way to compute greatest common divisors in the program. This will make the program easier
# to document and maintain.
#
# This is the essence of a closure. It binds the scope an object (function, variable, e.t.c) within a function
#
def gcd(*args):
"""(int, ..., int)->int
return the highest common factor of an abitrary
number of arguments "*args"
Preconditon: every argument must be an integer
>>> gcd(4, 8, 32)
4
"""
from functools import reduce
def t_gcd(a, b):
"""(int, int)->int
Return the greatest common divisor of a and b
came up with this from thought experiments
while driving. Should confirm against prexisting
solutions.
>>> t_gcd(6,9)
3
"""
assert isinstance(a, int) and isinstance(b, int), \
"Both a and b must be integers"
a, b = abs(a), abs(b)
if a == 0 or b == 0:
return max(a, b)
else:
while a != b:
diff = abs(a - b)
a, b = min(a, b), diff
return a
return reduce(t_gcd, args)
print(gcd(4, 8, 32))
print(gcd(6, 9))
print(t_gcd(6, 9))
# **The verdict:**
# As you can see, gcd works perfectly for 2 or more inputs and the user cannot access t_gcd.
#
# ## The C case.
# Nested fuctions are not available in standard C however GCC provides extensions to allow nested functions.
# These extentions are enabled by default.
#
# Visual studio doesnt have this capability (they dont even pretend to support C).
# You can achieve similar in C++ 11 and above using lambdas.
#
| Nested Functions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h2 align='center' style='color:purple'>Finding best model and hyper parameter tunning using GridSearchCV</h2>
# **For iris flower dataset in sklearn library, we are going to find out best model and best hyper parameters using GridSearchCV**
# <img src='iris_petal_sepal.png' height=300 width=300 />
# **Load iris flower dataset**
from sklearn import svm, datasets
iris = datasets.load_iris()
import pandas as pd
df = pd.DataFrame(iris.data,columns=iris.feature_names)
df['flower'] = iris.target
df['flower'] = df['flower'].apply(lambda x: iris.target_names[x])
df[47:150]
# <h3 style='color:blue'>Approach 1: Use train_test_split and manually tune parameters by trial and error</h3>
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(iris.data, iris.target, test_size=0.3)
model = svm.SVC(kernel='rbf',C=30,gamma='auto')
model.fit(X_train,y_train)
model.score(X_test, y_test)
# <h3 style='color:blue'>Approach 2: Use K Fold Cross validation</h3>
# **Manually try suppling models with different parameters to cross_val_score function with 5 fold cross validation**
cross_val_score(svm.SVC(kernel='linear',C=10,gamma='auto'),iris.data, iris.target, cv=5)
cross_val_score(svm.SVC(kernel='rbf',C=10,gamma='auto'),iris.data, iris.target, cv=5)
cross_val_score(svm.SVC(kernel='rbf',C=20,gamma='auto'),iris.data, iris.target, cv=5)
# **Above approach is tiresome and very manual. We can use for loop as an alternative**
# +
kernels = ['rbf', 'linear']
C = [1,10,20]
avg_scores = {}
for kval in kernels:
for cval in C:
cv_scores = cross_val_score(svm.SVC(kernel=kval,C=cval,gamma='auto'),iris.data, iris.target, cv=5)
avg_scores[kval + '_' + str(cval)] = np.average(cv_scores)
avg_scores
# -
# **From above results we can say that rbf with C=1 or 10 or linear with C=1 will give best performance**
# <h3 style='color:blue'>Approach 3: Use GridSearchCV</h3>
# **GridSearchCV does exactly same thing as for loop above but in a single line of code**
from sklearn.model_selection import GridSearchCV
clf = GridSearchCV(svm.SVC(gamma='auto'), {
'C': [1,10,20],
'kernel': ['rbf','linear']
}, cv=5, return_train_score=False)
clf.fit(iris.data, iris.target)
clf.cv_results_
df = pd.DataFrame(clf.cv_results_)
df
df[['param_C','param_kernel','mean_test_score']]
clf.best_params_
clf.best_score_
dir(clf)
# **Use RandomizedSearchCV to reduce number of iterations and with random combination of parameters. This is useful when you have too many parameters to try and your training time is longer. It helps reduce the cost of computation**
from sklearn.model_selection import RandomizedSearchCV
rs = RandomizedSearchCV(svm.SVC(gamma='auto'), {
'C': [1,10,20],
'kernel': ['rbf','linear']
},
cv=5,
return_train_score=False,
n_iter=2
)
rs.fit(iris.data, iris.target)
pd.DataFrame(rs.cv_results_)[['param_C','param_kernel','mean_test_score']]
# **How about different models with different hyperparameters?**
# +
from sklearn import svm
from sklearn.ensemble import RandomForestClassifier
from sklearn.linear_model import LogisticRegression
model_params = {
'svm': {
'model': svm.SVC(gamma='auto'),
'params' : {
'C': [1,10,20],
'kernel': ['rbf','linear']
}
},
'random_forest': {
'model': RandomForestClassifier(),
'params' : {
'n_estimators': [1,5,10]
}
},
'logistic_regression' : {
'model': LogisticRegression(solver='liblinear',multi_class='auto'),
'params': {
'C': [1,5,10]
}
}
}
# +
scores = []
for model_name, mp in model_params.items():
clf = GridSearchCV(mp['model'], mp['params'], cv=5, return_train_score=False)
clf.fit(iris.data, iris.target)
scores.append({
'model': model_name,
'best_score': clf.best_score_,
'best_params': clf.best_params_
})
df = pd.DataFrame(scores,columns=['model','best_score','best_params'])
df
# -
# **Based on above, I can conclude that SVM with C=1 and kernel='rbf' is the best model for solving my problem of iris flower classification**
| Program's_Contributed_By_Contributors/AI-Summer-Course/py-master/ML/15_gridsearch/15_grid_search.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# header files
import numpy as np
import torch
import torch.nn as nn
import torchvision
from google.colab import drive
drive.mount('/content/drive')
np.random.seed(1234)
torch.manual_seed(1234)
torch.cuda.manual_seed(1234)
# define transforms
train_transforms = torchvision.transforms.Compose([torchvision.transforms.RandomRotation(30),
torchvision.transforms.Resize((224, 224)),
torchvision.transforms.RandomHorizontalFlip(),
torchvision.transforms.ToTensor(),
torchvision.transforms.Normalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225])])
# datasets
train_data = torchvision.datasets.ImageFolder("/content/drive/My Drive/train_images/", transform=train_transforms)
val_data = torchvision.datasets.ImageFolder("/content/drive/My Drive/val_images/", transform=train_transforms)
print(len(train_data))
print(len(val_data))
# load the data
train_loader = torch.utils.data.DataLoader(train_data, batch_size=32, shuffle=True, num_workers=16, pin_memory=True)
val_loader = torch.utils.data.DataLoader(val_data, batch_size=32, shuffle=True, num_workers=16, pin_memory=True)
# model class
class VGG19_CBAM(torch.nn.Module):
# init function
def __init__(self, model, num_classes=2):
super().__init__()
# pool layer
self.pool = torch.nn.Sequential(torch.nn.MaxPool2d(kernel_size=2, stride=2))
# spatial attention
self.spatial_attention = torch.nn.Sequential(
torch.nn.Conv2d(2, 1, kernel_size=7, padding=3, stride=1),
torch.nn.BatchNorm2d(1),
torch.nn.Sigmoid()
)
# channel attention
self.max_pool_1 = torch.nn.Sequential(torch.nn.MaxPool2d(kernel_size=224, stride=224))
self.max_pool_2 = torch.nn.Sequential(torch.nn.MaxPool2d(kernel_size=112, stride=112))
self.max_pool_3 = torch.nn.Sequential(torch.nn.MaxPool2d(kernel_size=56, stride=56))
self.max_pool_4 = torch.nn.Sequential(torch.nn.MaxPool2d(kernel_size=28, stride=28))
self.max_pool_5 = torch.nn.Sequential(torch.nn.MaxPool2d(kernel_size=14, stride=14))
self.avg_pool_1 = torch.nn.Sequential(torch.nn.AvgPool2d(kernel_size=224, stride=224))
self.avg_pool_2 = torch.nn.Sequential(torch.nn.AvgPool2d(kernel_size=112, stride=112))
self.avg_pool_3 = torch.nn.Sequential(torch.nn.AvgPool2d(kernel_size=56, stride=56))
self.avg_pool_4 = torch.nn.Sequential(torch.nn.AvgPool2d(kernel_size=28, stride=28))
self.avg_pool_5 = torch.nn.Sequential(torch.nn.AvgPool2d(kernel_size=14, stride=14))
# features
self.features_1 = torch.nn.Sequential(*list(model.features.children())[:3])
self.features_2 = torch.nn.Sequential(*list(model.features.children())[3:6])
self.features_3 = torch.nn.Sequential(*list(model.features.children())[7:10])
self.features_4 = torch.nn.Sequential(*list(model.features.children())[10:13])
self.features_5 = torch.nn.Sequential(*list(model.features.children())[14:17])
self.features_6 = torch.nn.Sequential(*list(model.features.children())[17:20])
self.features_7 = torch.nn.Sequential(*list(model.features.children())[20:23])
self.features_8 = torch.nn.Sequential(*list(model.features.children())[23:26])
self.features_9 = torch.nn.Sequential(*list(model.features.children())[27:30])
self.features_10 = torch.nn.Sequential(*list(model.features.children())[30:33])
self.features_11 = torch.nn.Sequential(*list(model.features.children())[33:36])
self.features_12 = torch.nn.Sequential(*list(model.features.children())[36:39])
self.features_13 = torch.nn.Sequential(*list(model.features.children())[40:43])
self.features_14 = torch.nn.Sequential(*list(model.features.children())[43:46])
self.features_15 = torch.nn.Sequential(*list(model.features.children())[46:49])
self.features_16 = torch.nn.Sequential(*list(model.features.children())[49:52])
self.avgpool = nn.AdaptiveAvgPool2d(7)
# classifier
self.classifier = torch.nn.Sequential(
torch.nn.Linear(25088, 4096),
torch.nn.ReLU(inplace=True),
torch.nn.Dropout(),
torch.nn.Linear(4096, 4096),
torch.nn.ReLU(inplace=True),
torch.nn.Dropout(),
torch.nn.Linear(4096, 2)
)
# forward
def forward(self, x):
x = self.features_1(x)
scale = torch.nn.functional.sigmoid(self.max_pool_1(x) + self.avg_pool_1(x)).expand_as(x)
x = x * scale
scale = torch.cat((torch.max(x, 1)[0].unsqueeze(1), torch.mean(x, 1).unsqueeze(1)), dim=1)
scale = self.spatial_attention(scale)
x = x * scale
x = self.features_2(x)
scale = torch.nn.functional.sigmoid(self.max_pool_1(x) + self.avg_pool_1(x)).expand_as(x)
x = x * scale
scale = torch.cat((torch.max(x, 1)[0].unsqueeze(1), torch.mean(x, 1).unsqueeze(1)), dim=1)
scale = self.spatial_attention(scale)
x = x * scale
x = self.pool(x)
x = self.features_3(x)
scale = torch.nn.functional.sigmoid(self.max_pool_2(x) + self.avg_pool_2(x)).expand_as(x)
x = x * scale
scale = torch.cat((torch.max(x, 1)[0].unsqueeze(1), torch.mean(x, 1).unsqueeze(1)), dim=1)
scale = self.spatial_attention(scale)
x = x * scale
x = self.features_4(x)
scale = torch.nn.functional.sigmoid(self.max_pool_2(x) + self.avg_pool_2(x)).expand_as(x)
x = x * scale
scale = torch.cat((torch.max(x, 1)[0].unsqueeze(1), torch.mean(x, 1).unsqueeze(1)), dim=1)
scale = self.spatial_attention(scale)
x = x * scale
x = self.pool(x)
x = self.features_5(x)
scale = torch.nn.functional.sigmoid(self.max_pool_3(x) + self.avg_pool_3(x)).expand_as(x)
x = x * scale
scale = torch.cat((torch.max(x, 1)[0].unsqueeze(1), torch.mean(x, 1).unsqueeze(1)), dim=1)
scale = self.spatial_attention(scale)
x = x * scale
x = self.features_6(x)
scale = torch.nn.functional.sigmoid(self.max_pool_3(x) + self.avg_pool_3(x)).expand_as(x)
x = x * scale
scale = torch.cat((torch.max(x, 1)[0].unsqueeze(1), torch.mean(x, 1).unsqueeze(1)), dim=1)
scale = self.spatial_attention(scale)
x = x * scale
x = self.features_7(x)
scale = torch.nn.functional.sigmoid(self.max_pool_3(x) + self.avg_pool_3(x)).expand_as(x)
x = x * scale
scale = torch.cat((torch.max(x, 1)[0].unsqueeze(1), torch.mean(x, 1).unsqueeze(1)), dim=1)
scale = self.spatial_attention(scale)
x = x * scale
x = self.features_8(x)
scale = torch.nn.functional.sigmoid(self.max_pool_3(x) + self.avg_pool_3(x)).expand_as(x)
x = x * scale
scale = torch.cat((torch.max(x, 1)[0].unsqueeze(1), torch.mean(x, 1).unsqueeze(1)), dim=1)
scale = self.spatial_attention(scale)
x = x * scale
x = self.pool(x)
x = self.features_9(x)
scale = torch.nn.functional.sigmoid(self.max_pool_4(x) + self.avg_pool_4(x)).expand_as(x)
x = x * scale
scale = torch.cat((torch.max(x, 1)[0].unsqueeze(1), torch.mean(x, 1).unsqueeze(1)), dim=1)
scale = self.spatial_attention(scale)
x = x * scale
x = self.features_10(x)
scale = torch.nn.functional.sigmoid(self.max_pool_4(x) + self.avg_pool_4(x)).expand_as(x)
x = x * scale
scale = torch.cat((torch.max(x, 1)[0].unsqueeze(1), torch.mean(x, 1).unsqueeze(1)), dim=1)
scale = self.spatial_attention(scale)
x = x * scale
x = self.features_11(x)
scale = torch.nn.functional.sigmoid(self.max_pool_4(x) + self.avg_pool_4(x)).expand_as(x)
x = x * scale
scale = torch.cat((torch.max(x, 1)[0].unsqueeze(1), torch.mean(x, 1).unsqueeze(1)), dim=1)
scale = self.spatial_attention(scale)
x = x * scale
x = self.features_12(x)
scale = torch.nn.functional.sigmoid(self.max_pool_4(x) + self.avg_pool_4(x)).expand_as(x)
x = x * scale
scale = torch.cat((torch.max(x, 1)[0].unsqueeze(1), torch.mean(x, 1).unsqueeze(1)), dim=1)
scale = self.spatial_attention(scale)
x = x * scale
x = self.pool(x)
x = self.features_13(x)
scale = torch.nn.functional.sigmoid(self.max_pool_5(x) + self.avg_pool_5(x)).expand_as(x)
x = x * scale
scale = torch.cat((torch.max(x, 1)[0].unsqueeze(1), torch.mean(x, 1).unsqueeze(1)), dim=1)
scale = self.spatial_attention(scale)
x = x * scale
x = self.features_14(x)
scale = torch.nn.functional.sigmoid(self.max_pool_5(x) + self.avg_pool_5(x)).expand_as(x)
x = x * scale
scale = torch.cat((torch.max(x, 1)[0].unsqueeze(1), torch.mean(x, 1).unsqueeze(1)), dim=1)
scale = self.spatial_attention(scale)
x = x * scale
x = self.features_15(x)
scale = torch.nn.functional.sigmoid(self.max_pool_5(x) + self.avg_pool_5(x)).expand_as(x)
x = x * scale
scale = torch.cat((torch.max(x, 1)[0].unsqueeze(1), torch.mean(x, 1).unsqueeze(1)), dim=1)
scale = self.spatial_attention(scale)
x = x * scale
x = self.features_16(x)
scale = torch.nn.functional.sigmoid(self.max_pool_5(x) + self.avg_pool_5(x)).expand_as(x)
x = x * scale
scale = torch.cat((torch.max(x, 1)[0].unsqueeze(1), torch.mean(x, 1).unsqueeze(1)), dim=1)
scale = self.spatial_attention(scale)
x = x * scale
x = self.pool(x)
x = self.avgpool(x)
x = x.view(x.shape[0], -1)
x = self.classifier(x)
return x
# loss
criterion = torch.nn.CrossEntropyLoss()
# +
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
pretrained_model = torchvision.models.vgg19_bn(pretrained=True)
model = VGG19_CBAM(pretrained_model, 2)
model.to(device)
print(model)
# -
# optimizer to be used
optimizer = torch.optim.SGD(model.parameters(), lr=1e-3, momentum=0.9, weight_decay=0.001)
# +
train_losses = []
train_acc = []
val_losses = []
val_acc = []
best_metric = -1
best_metric_epoch = -1
# train and validate
for epoch in range(0, 30):
# train
model.train()
training_loss = 0.0
total = 0
correct = 0
for i, (input, target) in enumerate(train_loader):
input = input.to(device)
target = target.to(device)
optimizer.zero_grad()
output = model(input)
loss = criterion(output, target)
loss.backward()
optimizer.step()
training_loss = training_loss + loss.item()
_, predicted = output.max(1)
total += target.size(0)
correct += predicted.eq(target).sum().item()
training_loss = training_loss / float(len(train_loader))
training_accuracy = str(100.0 * (float(correct) / float(total)))
train_losses.append(training_loss)
train_acc.append(training_accuracy)
# validate
model.eval()
valid_loss = 0.0
total = 0
correct = 0
for i, (input, target) in enumerate(val_loader):
with torch.no_grad():
input = input.to(device)
target = target.to(device)
output = model(input)
loss = criterion(output, target)
_, predicted = output.max(1)
total += target.size(0)
correct += predicted.eq(target).sum().item()
valid_loss = valid_loss + loss.item()
valid_loss = valid_loss / float(len(val_loader))
valid_accuracy = str(100.0 * (float(correct) / float(total)))
val_losses.append(valid_loss)
val_acc.append(valid_accuracy)
# store best model
if(float(valid_accuracy) > best_metric and epoch >= 10):
best_metric = float(valid_accuracy)
best_metric_epoch = epoch
torch.save(model.state_dict(), "best_model.pth")
print()
print("Epoch" + str(epoch) + ":")
print("Training Accuracy: " + str(training_accuracy) + " Validation Accuracy: " + str(valid_accuracy))
print("Training Loss: " + str(training_loss) + " Validation Loss: " + str(valid_loss))
print()
# +
import matplotlib.pyplot as plt
e = []
for index in range(0, 30):
e.append(index)
plt.plot(e, train_losses)
plt.show()
# -
plt.plot(e, val_losses)
plt.show()
| notebooks/.ipynb_checkpoints/vgg19_pretrained_cbam-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
os.environ['CUDA_VISIBLE_DEVICES']='0,1,2,3'
# %run -p ../latent_ode_infocnf.py --adjoint False --visualize True --niters 1200 --monitor_freq 200 --lr 0.001 --save experiments_lr_0_001_1200/cnf --gpu 3
| examples/main_latent_ode-Copy3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import rebound
import reboundx
import pandas as pd
import numpy as np
from numpy import genfromtxt
import csv
import scipy
from scipy import signal
from IPython.display import display, clear_output
import matplotlib.pyplot as plt
from matplotlib import colors
# %matplotlib inline
# +
radeg = np.pi/180
########################
# Define functions for adding a number of generated Trojan asteroids at ~L4/L5
########################
def add_L4(sim, number):
a_rand = np.random.normal(20,2,size=number)
a_rand = a_rand/100 + 5
e_rand = np.random.normal(9,2,size=number)
e_rand = e_rand/100
w_rand = np.random.normal(170,4,size=number)*radeg
half = int(number/2)
i_rand1 = np.random.normal(9,4,size=half+1)*radeg
i_rand2 = np.random.normal(-9,4,size=half)*radeg
i_rand = np.concatenate((i_rand1,i_rand2))
for i in range(number):
sem = a_rand[i]
ecc = e_rand[i]
icl = i_rand[i]
Ome = w_rand[i]
has = 'L4 {0}'.format(i)
sim.add(m=0, primary=sim.particles['Sun'], a=sem, e=ecc, inc=icl, Omega=Ome, hash=has)
return
def add_L5(sim, number):
a_rand = np.random.normal(20,2,size=number)
a_rand = a_rand/100 + 5
e_rand = np.random.normal(9,2,size=number)
e_rand = e_rand/100
w_rand = np.random.normal(60,4,size=number)*radeg
half = int(number/2)
i_rand1 = np.random.normal(9,4,size=half+1)*radeg
i_rand2 = np.random.normal(-9,4,size=half)*radeg
i_rand = np.concatenate((i_rand1,i_rand2))
for i in range(number):
sem = a_rand[i]
ecc = e_rand[i]
icl = i_rand[i]
Ome = w_rand[i]
has = 'L5 {0}'.format(i)
sim.add(m=0, primary=sim.particles['Sun'], a=sem, e=ecc, inc=icl, Omega=Ome, hash=has)
return
# +
def masses(x):
# for input array of time values, approximate M_star (in M_sol) at those times in its life
y = np.zeros_like(x)
for i, time in enumerate(x):
if (time <= 1.132e10):
y[i] = 1
elif (1.132e10 < time <= 1.1336e10):
y[i] = 0.05 * (708.5 - time/(1.6e7))**(1/3) + .95
elif (1.1336e10 < time <= 1.1463e10):
y[i] = -8**((time - 1.1463e10)/574511)/2.4 + .95
elif (1.1463e10 < time):
y[i] = 0.54
return y
def lums_array(x):
# for input array of time values, approximate log(L_star) (in log(L_sol)) at those times
y = np.zeros_like(x)
for i, time in enumerate(x):
if (time <= 1.113e10):
y[i] = 1.05
elif (1.113e10 < time <= 1.1225e10):
y[i] = 1.45 + ((1.45 - 1.1)/(1.1225e10 - 1.1135e10))*(time - 1.1225e10)
elif (1.1225e10 < time <= 1.125e10):
y[i] = 1.45
elif (1.125 < time <= 1.1336e10):
y[i] = 1.35 + .1*1.002**((time - 1.125e10)/58000)
elif (1.1336e10 < time <= 1.142e10):
y[i] = 1.673
elif (1.142e10 < time <= 1.14397e10):
y[i] = 3.198e-9*time - 34.85
elif (1.14397e10 < time <= 1.14479e10):
y[i] = 1.736 + 0.032*1.5**((time - 1.14455e10)/360000)
elif (1.14479e10 < time <= 1.1462e10):
y[i] = 2.15 + 0.00021*1.5**((time - 1.1444e10)/870000)
elif (1.1462e10 < time <= 1.14632e10):
y[i] = 3.5 + (.43/0.0001e10)*(time - 1.1463e10)
elif (1.14632e10 < time <= 1.14636e10):
y[i] = 2.3*((time - 1.1463e10)/45000)**(-0.3)
elif (1.14636e10 < time <= 1.14654715e10):
y[i] = .2 + ((.2 - 1.05)/(1.14654715e10 - 1.14636e10))*(time - 1.14654715e10)
elif (1.14654715e10 < time):
y[i] = .2
return y
def inst_lum(x):
# for a single time input, output log(L_star) (in log(L_sol)) at that time
time = x
if (time <= 1.113e10):
y = 1.05
elif (1.113e10 < time <= 1.1225e10):
y = 1.45 + ((1.45 - 1.1)/(1.1225e10 - 1.1135e10))*(time - 1.1225e10)
elif (1.1225e10 < time <= 1.125e10):
y = 1.45
elif (1.125 < time <= 1.1336e10):
y = 1.35 + .1*1.002**((time - 1.125e10)/58000)
elif (1.1336e10 < time <= 1.142e10):
y = 1.673
elif (1.142e10 < time <= 1.14397e10):
y = 3.198e-9*time - 34.85
elif (1.14397e10 < time <= 1.14479e10):
y = 1.736 + 0.032*1.5**((time - 1.14455e10)/360000)
elif (1.14479e10 < time <= 1.1462e10):
y = 2.15 + 0.00021*1.5**((time - 1.1444e10)/870000)
elif (1.1462e10 < time <= 1.14632e10):
y = 3.5 + (.43/0.0001e10)*(time - 1.1463e10)
elif (1.14632e10 < time <= 1.14636e10):
y = 2.3*((time - 1.1463e10)/45000)**(-0.3)
elif (1.14636e10 < time <= 1.14654715e10):
y = .2 + ((.2 - 1.05)/(1.14654715e10 - 1.14636e10))*(time - 1.14654715e10)
elif (1.14654715e10 < time):
y = .2
return y
###############################
# Define our all-important custom force, derived from Veras et al. 2019 eq. 23,
# using their Model A to encapsulate edge case Yarkovsky physics
###############################
def yark(simp, rebx_force, particles, N):
sim = simp.contents
part = sim.particles
current_time = sim.t + T0
L_sol = np.exp(inst_lum(current_time))
sim.move_to_hel()
for troj in range(num_tr):
i = troj + 1
x = part[i].x ; y = part[i].y ; z = part[i].z
vx = part[i].vx; vy = part[i].vy; vz = part[i].vz
R = troj_radii[i-1]
m_ast = troj_masses[i-1]
c = 63197.8 # speed of light in au/yr
r = np.sqrt(x**2 + y**2 + z**2)
A = (R**2 * L_sol)/(4*m_ast*r**2)
D = (c - vx*x - vy*y - vz*z)/r
part[i].ax += A/c**2 * (D*x - vx)
part[i].ay += A/c**2 * (D*0.25*x + D*y - vx - vy)
part[i].az += A/c**2 * (D*z - vz)
return
# +
######################
# Get an array of masses and luminosities over the period of interest.
# This cell also plots those masses and luminosities.
######################
N_times = 1000
T0 = 1.14625e10
t_tot = 750000
ts = np.linspace(0, t_tot, N_times)
mtimes = masses(ts + T0)
lumins = lums_array(ts + T0)
plt.plot(ts + T0, mtimes)
plt.plot(ts + T0, lumins)
plt.legend([r"$M_{star}$ / $M_\odot$", r"log($L_{star}$) - log($L_\odot$)"])
plt.xlabel("Time/yr")
plt.show()
# -
# # Trojan analysis
# +
sim = rebound.Simulation()
M0 = mtimes[0]
print("Star initial mass:", M0, "M_sol")
print("Star initial age: ", T0, "yrs")
radeg = np.pi/180
Om_jup = 100.556*radeg
om_jup = 14.753*radeg
num_L4 = 20
num_L5 = 20
num_tr = num_L4 + num_L5
#############
# Add sun, trojans, Jupiter, and Saturn
#############
sim.add(m=M0,x=0, y=0, z=0, vx=0, vy=0, vz=0, hash='Sun')
add_L4(sim, num_L4)
add_L5(sim, num_L5)
sim.add(m=9.543e-4, a=5.2, e=.04839, inc=.022689, Omega=Om_jup, omega=om_jup, hash='jupiter')
sim.add(m=2.8575e-4, primary=sim.particles['Sun'], a=9.537, e=0.05415, inc=0.043284, Omega=1.9844, omega=1.61324, hash='Saturn')
#############
#set simulation parameters
#############
sim.dt = 0.5
sim.move_to_com()
ps = sim.particles
fig, ax = rebound.OrbitPlot(sim)
# +
rad_ast = 10 # radius in km
troj_radii = np.full(num_tr, rad_ast/1.496e+8) # gives each asteroid a radius in AU
mass_typic = 3*(4/3)*np.pi*(rad_ast*100000)**3 # gives typical mass @ this radius, w/ density = 3 g cm^-3
troj_masses = np.random.normal(mass_typic, .3*mass_typic, num_tr)
# gives array of values around that mass
troj_masses /= 1.9891e33 # divides each mass by M_sol to get masses in M_sol
print("Typical asteroid mass:", mass_typic, "g")
print("Average asteroid mass:", np.mean(troj_masses), "M_sol")
# -
Nout = 100000
times = np.linspace(0,t_tot,Nout)
mstar = np.zeros(Nout)
# +
rebx = reboundx.Extras(sim)
starmass = reboundx.Interpolator(rebx, ts, mtimes, 'spline')
yrkv = rebx.create_force("yarkovsky")
yrkv.force_type = "vel"
yrkv.update_accelerations = yark
rebx.add_force(yrkv)
#gh = rebx.load_force("gravitational_harmonics")
#rebx.add_force(gh)
#mof = rebx.load_force("modify_orbits_forces")
#rebx.add_force(mof)
J2 = 14736e-6
J2prime = 0.045020
R_jup = 0.000477895
#ps['jupiter'].params["J2"] = J2prime
#ps['jupiter'].params["R_eq"] = R_jup
# +
# initialize arrays for tracking progression of bodies over integration
mass = np.zeros(Nout)
x_sol = np.zeros(Nout); y_sol = np.zeros(Nout)
x_sol[0] = ps['Sun'].x
y_sol[0] = ps['Sun'].y
x_jup = np.zeros(Nout); y_jup = np.zeros(Nout)
x_jup[0] = ps['jupiter'].x
y_jup[0] = ps['jupiter'].y
a_jup = np.zeros(Nout)
e_jup = np.zeros(Nout)
i_jup = np.zeros(Nout)
pmjup = np.zeros(Nout)
lmjup = np.zeros(Nout)
a_jup[0] = ps['jupiter'].a
e_jup[0] = ps['jupiter'].e
i_jup[0] = ps['jupiter'].inc
pmjup[0] = ps['jupiter'].pomega
lmjup[0] = ps['jupiter'].l
a_vals = np.zeros((num_tr, Nout))
e_vals = np.zeros((num_tr, Nout))
i_vals = np.zeros((num_tr, Nout))
pmvals = np.zeros((num_tr, Nout))
lmvals = np.zeros((num_tr, Nout))
x_vals = np.zeros((num_tr, Nout))
y_vals = np.zeros((num_tr, Nout))
for moon in range(num_L4):
a_vals[moon,0] = ps['L4 {0}'.format(moon)].a
e_vals[moon,0] = ps['L4 {0}'.format(moon)].e
i_vals[moon,0] = ps['L4 {0}'.format(moon)].inc
lmvals[moon,0] = ps['L4 {0}'.format(moon)].l
pmvals[moon,0] = ps['L4 {0}'.format(moon)].pomega
x_vals[moon,0] = ps['L4 {0}'.format(moon)].x
y_vals[moon,0] = ps['L4 {0}'.format(moon)].y
for moon in range(num_L5):
a_vals[moon + num_L4,0] = ps['L5 {0}'.format(moon)].a
e_vals[moon + num_L4,0] = ps['L5 {0}'.format(moon)].e
i_vals[moon + num_L4,0] = ps['L5 {0}'.format(moon)].inc
lmvals[moon + num_L4,0] = ps['L5 {0}'.format(moon)].l
pmvals[moon + num_L4,0] = ps['L5 {0}'.format(moon)].pomega
x_vals[moon + num_L4,0] = ps['L5 {0}'.format(moon)].x
y_vals[moon + num_L4,0] = ps['L5 {0}'.format(moon)].y
# +
# %%time
for i, time in enumerate(times):
sim.integrate(time)
ps[0].m = starmass.interpolate(rebx, t=sim.t)
sim.move_to_com()
mass[i] = ps['Sun'].m
a_jup[i] = ps['jupiter'].a
e_jup[i] = ps['jupiter'].e
i_jup[i] = ps['jupiter'].inc
pmjup[i] = ps['jupiter'].pomega
lmjup[i] = ps['jupiter'].l
if i == 0:
fig = rebound.OrbitPlot(sim, figsize=(4,4), periastron=True)
if (i+1)%(Nout/4) == 0:
fig = rebound.OrbitPlot(sim, figsize=(4,4), periastron=True)
for moon in range(num_L4):
a_vals[moon,i] = ps['L4 {0}'.format(moon)].a
e_vals[moon,i] = ps['L4 {0}'.format(moon)].e
i_vals[moon,i] = ps['L4 {0}'.format(moon)].inc
lmvals[moon,i] = ps['L4 {0}'.format(moon)].l
pmvals[moon,i] = ps['L4 {0}'.format(moon)].pomega
x_vals[moon,i] = ps['L4 {0}'.format(moon)].x
y_vals[moon,i] = ps['L4 {0}'.format(moon)].y
for moon in range(num_L5):
a_vals[moon + num_L4,i] = ps['L5 {0}'.format(moon)].a
e_vals[moon + num_L4,i] = ps['L5 {0}'.format(moon)].e
i_vals[moon + num_L4,i] = ps['L5 {0}'.format(moon)].inc
lmvals[moon + num_L4,i] = ps['L5 {0}'.format(moon)].l
pmvals[moon + num_L4,i] = ps['L5 {0}'.format(moon)].pomega
x_vals[moon + num_L4,i] = ps['L5 {0}'.format(moon)].x
y_vals[moon + num_L4,i] = ps['L5 {0}'.format(moon)].y
# +
i_vals /= radeg
fig, ax = plt.subplots(2,1,figsize=(15,5), sharex=True)
end = 71600
ax[0].plot(times[:end],mass[:end], label='simulation')
ax[0].set_ylabel(r"$M_\star$ / $M_\odot$", fontsize=16)
ax[1].plot(times[:end],a_jup[:end])
ax[1].set_ylabel(r"$a_{Jup}$ / AU", fontsize=16)
fig.tight_layout()
trojs, axes = plt.subplots(4,1,figsize=(15,10))
axes[0].plot(times[:end], a_vals.T[:end, num_L4:], 'b')
axes[0].plot(times[:end], a_vals.T[:end, :num_L4], 'r')
axes[0].plot(times[:end], a_jup[:end], "k")
axes[0].set_ylabel(r"$a_{troj}$ / AU", fontsize=16)
axes[1].plot(times[:end], e_vals.T[:end, num_L4:], 'b')
axes[1].plot(times[:end], e_vals.T[:end, :num_L4], 'r')
axes[1].set_ylabel(r"$e_{troj}$", fontsize=16)
axes[2].plot(times[:end], (lmvals.T[:end,10]), 'b')
axes[2].plot(times[:end], (lmvals.T[:end,0]), 'r')
#axes[2].plot(times[:], signal.medfilt(Omvals.T[:, 0:5],[499,1]), 'r')
#axes[2].plot(times, change(Omvals.T)[:,1], 'k', a=3)
axes[2].set_ylabel(r"$\lambda_{troj}$", fontsize=16)
axes[3].plot(times[:end], i_vals.T[:end, num_L4:], 'b')
axes[3].plot(times[:end], i_vals.T[:end, :num_L4], 'r')
axes[3].set_ylabel(r"$i$ / degrees", fontsize=16)
plt.show()
# -
# +
fft_4lambda1 = scipy.fft.rfft(np.sin(lmvals.T[end-7500:end,:num_L4]), axis=0)
fft_4lambda2 = scipy.fft.rfft(np.sin(lmvals.T[0:7500,:num_L4]), axis=0)
fft_5lambda1 = scipy.fft.rfft(np.sin(lmvals.T[end-7500:end,num_L4:]), axis=0)
fft_5lambda2 = scipy.fft.rfft(np.sin(lmvals.T[0:7500,num_L4:]), axis=0)
freq = scipy.fft.rfftfreq(times[0:7500].shape[-1])
fig, ax = plt.subplots(1,2, figsize=(15,5), sharey=True)
ax[0].plot(freq,np.abs(np.mean(fft_4lambda2, axis=1)), c="r", ls=":", alpha=.5)
ax[0].plot(freq,np.abs(np.mean(fft_4lambda1, axis=1)), c="r")
ax[1].plot(freq,np.abs(np.mean(fft_5lambda2, axis=1)), c="b", ls=":", alpha=.5)
ax[1].plot(freq,np.abs(np.mean(fft_5lambda1, axis=1)), c="b")
for i in range(2):
ax[i].set_yscale("log")
ax[i].set_xlim(0,0.17)
fig.suptitle(r"FFT of sin($\lambda$)", fontsize=14)
ax[0].legend(["start of sim", "end of sim"])
ax[0].set_title(r"Average of $L_4$ data")
ax[1].set_title(r"Average of $L_5$ data")
ax[0].set_ylabel("Power", fontsize=12)
fig.tight_layout()
fig.show()
# -
plt.plot(freq,np.abs(np.mean(fft_4lambda1, axis=1)), "r")
plt.plot(freq,np.abs(np.mean(fft_5lambda1, axis=1)), "b")
plt.yscale("log")
plt.xlim(-0.01,0.17)
plt.ylabel("Power")
plt.legend([r"Average $L_4$",r"Average $L_5$"])
plt.title(r"FFT of sin($\lambda$) at end of simulation")
plt.show()
# +
fft_4lambda1 = scipy.fft.rfft(a_vals.T[end-7500:end,:num_L4], axis=0)
fft_4lambda2 = scipy.fft.rfft(a_vals.T[0:7500,:num_L4], axis=0)
fft_5lambda1 = scipy.fft.rfft(a_vals.T[end-7500:end,num_L4:], axis=0)
fft_5lambda2 = scipy.fft.rfft(a_vals.T[0:7500,num_L4:], axis=0)
freq = scipy.fft.rfftfreq(times[0:7500].shape[-1])
fig, ax = plt.subplots(1,2, figsize=(15,5), sharey=True)
ax[0].plot(freq,np.abs(np.mean(fft_4lambda2, axis=1)), c="r", ls=":", alpha=.5)
ax[0].plot(freq,np.abs(np.mean(fft_4lambda1, axis=1)), c="r")
ax[1].plot(freq,np.abs(np.mean(fft_5lambda2, axis=1)), c="b", ls=":", alpha=.5)
ax[1].plot(freq,np.abs(np.mean(fft_5lambda1, axis=1)), c="b")
for i in range(2):
ax[i].set_yscale("log")
ax[i].set_xlim(-0.01,0.2)
fig.suptitle(r"FFT of $a$", fontsize=14)
ax[0].legend(["start of sim", "end of sim"])
ax[0].set_title(r"Average of $L_4$ data")
ax[1].set_title(r"Average of $L_5$ data")
ax[0].set_ylabel("Power", fontsize=12)
fig.tight_layout()
fig.show()
# +
xsep = np.abs(ps['Sun'].x - ps['jupiter'].x)
ysep = np.abs(ps['Sun'].y - ps['jupiter'].y)
r_jupsol = np.sqrt(xsep**2 + ysep**2)
x_L4 = r_jupsol*np.cos(np.pi/3)
x_L5 = x_L4
y_L4 = r_jupsol*np.sin(np.pi/3)
y_L5 = -y_L4
fig, ax = plt.subplots(figsize=(10,10))
ax.scatter(y_vals.T[end,:],x_vals.T[end,:])
ax.scatter(ps[0].y,ps[0].x)
ax.scatter(ps['jupiter'].y,ps['jupiter'].x)
#ax.scatter([x_L4,x_L5],[y_L4,y_L5], c='k', s=60)
#ax.axvline(x_L4,ymin=-10,ymax=10, c='k', alpha=.4)
#ax.axhline(0,xmin=-10,xmax=10,c='k',alpha=.4)
ax.set_aspect("equal")
#ax.set_ylim(-10,10)
#ax.set_xlim(-10,10)
fig.show()
# +
a_inits_L4 = a_vals.T[0,:num_L4] ; a_inits_L5 = a_vals.T[0,num_L4:]
a_final_L4 = a_vals.T[Nout-1,:num_L4] ; a_final_L5 = a_vals.T[Nout-1,num_L4:]
plt.scatter(np.power(a_inits_L4,1.5)/a_jup[0]**1.5, (np.fabs(a_final_L4-a_inits_L4)+1.0e-16)/a_inits_L4,marker=".",s=36,c="b")
plt.scatter(np.power(a_inits_L5,1.5)/a_jup[0]**1.5, (np.fabs(a_final_L5-a_inits_L5)+1.0e-16)/a_inits_L5,marker=".",s=36,c="r")
plt.xlabel(r"Initial Period ratio $P_{ast}/P_{Jup}$")
plt.ylabel(r"log($\Delta a/a_{initial}$)")
plt.yscale("log")
plt.show()
# -
# Idea for this plot came from https://rebound.readthedocs.io/en/latest/ipython/Testparticles.html -- doesn't look quite as cool as the one there, perhaps I need more particles
| 9-Archive/Old_Sims/.ipynb_checkpoints/Trojs-Sim-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="m-kPGh3nymNf" colab_type="code" outputId="53901d2b-ae48-4901-ceaf-6e869b9fb896" executionInfo={"status": "ok", "timestamp": 1583448838320, "user_tz": -60, "elapsed": 521, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14989719145641357731"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# cd "/content/drive/My Drive/Colab Notebooks/dw_matrix/matrix_two/dw_matrix_car"
# + id="uxpMn2gn59lR" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="6323bb2d-aa3b-4c47-c712-b85acc28ec60" executionInfo={"status": "ok", "timestamp": 1583448858298, "user_tz": -60, "elapsed": 496, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14989719145641357731"}}
pwd
# + id="UlpjxBvOyqro" colab_type="code" outputId="10feee64-59ad-43a1-e1d8-aa7ce3761161" executionInfo={"status": "ok", "timestamp": 1583447059127, "user_tz": -60, "elapsed": 4820, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14989719145641357731"}} colab={"base_uri": "https://localhost:8080/", "height": 168}
import pandas as pd
import numpy as np
from sklearn.dummy import DummyRegressor
from sklearn.tree import DecisionTreeRegressor
from sklearn.ensemble import RandomForestRegressor
import xgboost as xgb
from sklearn.metrics import mean_absolute_error as mae
from sklearn.model_selection import cross_val_score, KFold
import eli5
from eli5.sklearn import PermutationImportance
# + id="Ie3JlYO5zDo6" colab_type="code" outputId="5b6c02aa-5283-4e60-fdf9-b8548a745efa" executionInfo={"status": "ok", "timestamp": 1583450364074, "user_tz": -60, "elapsed": 2480, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14989719145641357731"}} colab={"base_uri": "https://localhost:8080/", "height": 644}
df = pd.read_hdf('data/car.h5')
df.shape
df.head(5)
# + id="newT5rOdzQLU" colab_type="code" colab={}
SUFFIX_CAT = '__cat'
for feat in df.columns:
if isinstance(df[feat][0], list): continue
factorized_values = df[feat].factorize()[0]
if SUFFIX_CAT in feat:
df[feat] = factorized_values
else:
df[feat + SUFFIX_CAT] = factorized_values
# + id="H4SVop_tzTUW" colab_type="code" outputId="b4d939e3-f5d9-414d-d4af-589ef1e3e5ae" executionInfo={"status": "ok", "timestamp": 1583447107243, "user_tz": -60, "elapsed": 515, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14989719145641357731"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
cat_feats = [x for x in df.columns if SUFFIX_CAT in x]
cat_feats = [x for x in cat_feats if 'price' not in x]
len(cat_feats)
# + id="XzkL2aC4zWRj" colab_type="code" outputId="fac25c68-506c-45c2-bced-9336aedf9cdd" executionInfo={"status": "ok", "timestamp": 1583447122540, "user_tz": -60, "elapsed": 5005, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14989719145641357731"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
X = df[cat_feats].values
y = df['price_value'].values
model = DecisionTreeRegressor(max_depth=5)
scores = cross_val_score(model, X, y, cv=3, scoring='neg_mean_absolute_error')
np.mean(scores), np.std(scores)
# + id="B_JJFn_vzY6U" colab_type="code" colab={}
def run_model(model, feats):
X = df[feats].values
y = df['price_value'].values
scores = cross_val_score(model, X, y, cv=3, scoring='neg_mean_absolute_error')
return np.mean(scores), np.std(scores)
# + id="wDEcZEywzb8s" colab_type="code" outputId="dd9181ec-9964-4c65-d2d4-f04e4ed452d4" executionInfo={"status": "ok", "timestamp": 1583447143932, "user_tz": -60, "elapsed": 3632, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14989719145641357731"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
model = DecisionTreeRegressor(max_depth=5)
run_model(model, cat_feats)
# + id="KyFI1vcIzed_" colab_type="code" outputId="40ee98b1-af05-47c0-aea7-5e256ce6a8e6" executionInfo={"status": "ok", "timestamp": 1583447235949, "user_tz": -60, "elapsed": 85278, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14989719145641357731"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
model = RandomForestRegressor(max_depth=5, n_estimators=50, random_state=0)
run_model(model, cat_feats)
# + id="2qPMrSFLzg_9" colab_type="code" outputId="6a956971-418c-4815-b2f3-80fab066d633" executionInfo={"status": "ok", "timestamp": 1583447294918, "user_tz": -60, "elapsed": 132284, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14989719145641357731"}} colab={"base_uri": "https://localhost:8080/", "height": 88}
xgb_params = {
'max_depth': 5,
'n_estimators': 50,
'learning_rate': 0.1,
'seed': 0
}
model = run_model(xgb.XGBRegressor(**xgb_params), cat_feats)
# + id="Vk-qCfZmzj61" colab_type="code" outputId="5df65165-3ea3-403b-8fc8-c931402ac285" executionInfo={"status": "ok", "timestamp": 1583447295728, "user_tz": -60, "elapsed": 807, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14989719145641357731"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
model
# + id="Cs09hM0lzndE" colab_type="code" outputId="4ab75d75-3ddb-42a6-ab42-f9cc980f8e8f" executionInfo={"status": "ok", "timestamp": 1583447648895, "user_tz": -60, "elapsed": 353971, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14989719145641357731"}} colab={"base_uri": "https://localhost:8080/", "height": 428}
m = xgb.XGBRegressor(max_depth=5, n_estimators=50, learning_rate=0.1, seed=0)
m.fit(X, y)
imp = PermutationImportance(m, random_state=0).fit(X, y)
eli5.show_weights(imp, feature_names=cat_feats)
# + id="kCxgZzgGzoEN" colab_type="code" outputId="ee404829-34c3-4c9b-ee31-d1b65c021611" executionInfo={"status": "ok", "timestamp": 1583447648898, "user_tz": -60, "elapsed": 353967, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14989719145641357731"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
feats=['param_napęd__cat','param_rok-produkcji__cat','param_stan__cat','param_skrzynia-biegów__cat','param_faktura-vat__cat','param_moc__cat','param_marka-pojazdu__cat','feature_kamera-cofania__cat','param_typ__cat','param_pojemność-skokowa__cat','seller_name__cat','feature_wspomaganie-kierownicy__cat','param_model-pojazdu__cat','param_wersja__cat','param_kod-silnika__cat','feature_system-start-stop__cat','feature_asystent-pasa-ruchu__cat','feature_czujniki-parkowania-przednie__cat','feature_łopatki-zmiany-biegów__cat','feature_regulowane-zawieszenie__cat']
len(feats)
# + id="pWpvb1ZXzwA_" colab_type="code" outputId="970b620b-3f4b-4bef-d657-b155a23b3af7" executionInfo={"status": "ok", "timestamp": 1583447661404, "user_tz": -60, "elapsed": 366472, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14989719145641357731"}} colab={"base_uri": "https://localhost:8080/", "height": 88}
model = run_model(xgb.XGBRegressor(**xgb_params), feats)
# + id="6NJhanUdzzxf" colab_type="code" outputId="d7bc9698-67d3-4fde-ed70-536a4801367a" executionInfo={"status": "ok", "timestamp": 1583447661405, "user_tz": -60, "elapsed": 366469, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14989719145641357731"}} colab={"base_uri": "https://localhost:8080/", "height": 68}
df['param_napęd'].unique()
# + id="d-mLmfkNz23f" colab_type="code" colab={}
df['param_rok-produkcji'] = df['param_rok-produkcji'].map(lambda x: -1 if str(x) == 'None' else int(x))
# + id="tGvBVzeEz6en" colab_type="code" outputId="fdf985c2-241c-42f6-eeb0-2694437dba7e" executionInfo={"status": "ok", "timestamp": 1583447673969, "user_tz": -60, "elapsed": 379027, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14989719145641357731"}} colab={"base_uri": "https://localhost:8080/", "height": 105}
feats=['param_napęd__cat','param_rok-produkcji','param_stan__cat','param_skrzynia-biegów__cat','param_faktura-vat__cat','param_moc__cat','param_marka-pojazdu__cat','feature_kamera-cofania__cat','param_typ__cat','param_pojemność-skokowa__cat','seller_name__cat','feature_wspomaganie-kierownicy__cat','param_model-pojazdu__cat','param_wersja__cat','param_kod-silnika__cat','feature_system-start-stop__cat','feature_asystent-pasa-ruchu__cat','feature_czujniki-parkowania-przednie__cat','feature_łopatki-zmiany-biegów__cat','feature_regulowane-zawieszenie__cat']
len(feats)
model = run_model(xgb.XGBRegressor(**xgb_params), feats)
model
# + id="LiRyGZh6z9DJ" colab_type="code" outputId="9eca6584-e5ea-4f8f-9046-83f51e4bb0b5" executionInfo={"status": "ok", "timestamp": 1583447673971, "user_tz": -60, "elapsed": 379026, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14989719145641357731"}} colab={"base_uri": "https://localhost:8080/", "height": 1000}
df['param_moc'].unique()
# + id="_SJzsQ72z_yP" colab_type="code" colab={}
df['param_moc'] = df['param_moc'].map(lambda x: -1 if str(x) == 'None' else int(x.split(' ')[0]))
# + id="K6LkJT330Dne" colab_type="code" outputId="8288bba0-4dab-4132-e57d-1a6ba1b973e7" executionInfo={"status": "ok", "timestamp": 1583447674198, "user_tz": -60, "elapsed": 372555, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14989719145641357731"}} colab={"base_uri": "https://localhost:8080/", "height": 680}
df['param_moc'].unique()
# + id="N5PfIGiJ0F2w" colab_type="code" outputId="bb592279-be76-48a1-b30e-0554f6d55cd2" executionInfo={"status": "ok", "timestamp": 1583447686914, "user_tz": -60, "elapsed": 368154, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14989719145641357731"}} colab={"base_uri": "https://localhost:8080/", "height": 88}
feats=['param_napęd__cat','param_rok-produkcji','param_stan__cat','param_skrzynia-biegów__cat','param_faktura-vat__cat','param_moc','param_marka-pojazdu__cat','feature_kamera-cofania__cat','param_typ__cat','param_pojemność-skokowa__cat','seller_name__cat','feature_wspomaganie-kierownicy__cat','param_model-pojazdu__cat','param_wersja__cat','param_kod-silnika__cat','feature_system-start-stop__cat','feature_asystent-pasa-ruchu__cat','feature_czujniki-parkowania-przednie__cat','feature_łopatki-zmiany-biegów__cat','feature_regulowane-zawieszenie__cat']
len(feats)
model = run_model(xgb.XGBRegressor(**xgb_params), feats)
# + id="z8bHPqiL0KCA" colab_type="code" outputId="f738a6a1-f71b-49c3-d188-4f5fb8dd6282" executionInfo={"status": "ok", "timestamp": 1583447687785, "user_tz": -60, "elapsed": 861, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14989719145641357731"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
model
# + id="lDd1qoRM0RYg" colab_type="code" colab={}
df['param_pojemność-skokowa'] = df['param_pojemność-skokowa'].map(lambda x: -1 if str(x) == 'None' else int(str(x).split('cm')[0].replace(' ', '')))
# + id="hvxvpHBI0R4o" colab_type="code" outputId="e07c7239-f2aa-4f1e-9ae5-5a1d9817fb49" executionInfo={"status": "ok", "timestamp": 1583447688145, "user_tz": -60, "elapsed": 1214, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14989719145641357731"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
df['param_pojemność-skokowa'].unique()
# + id="ndHHweXw0UjA" colab_type="code" outputId="454bf344-55bb-4fec-bf53-a14180ceb52c" executionInfo={"status": "ok", "timestamp": 1583447700089, "user_tz": -60, "elapsed": 13155, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14989719145641357731"}} colab={"base_uri": "https://localhost:8080/", "height": 105}
feats=['param_napęd__cat','param_rok-produkcji','param_stan__cat','param_skrzynia-biegów__cat','param_faktura-vat__cat','param_moc','param_marka-pojazdu__cat','feature_kamera-cofania__cat','param_typ__cat','param_pojemność-skokowa','seller_name__cat','feature_wspomaganie-kierownicy__cat','param_model-pojazdu__cat','param_wersja__cat','param_kod-silnika__cat','feature_system-start-stop__cat','feature_asystent-pasa-ruchu__cat','feature_czujniki-parkowania-przednie__cat','feature_łopatki-zmiany-biegów__cat','feature_regulowane-zawieszenie__cat']
len(feats)
run_model(xgb.XGBRegressor(**xgb_params), feats)
# + id="qnOY6k8j0XNo" colab_type="code" colab={}
| day4_xgboost.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Running and fitting EZmock
#
# -----
# This jupyter notebook demonstrates the process of generating EZmock catalogues with user-supplied parameters and comparing the clustering measurements with references.
import os, sys
pyez_path = os.getcwd() + '/python'
if not os.path.isfile(pyez_path + '/pyEZmock.py'):
pyez_path='/global/u2/z/zhaoc/work/pyEZmock/python'
sys.path.append(pyez_path)
from pyEZmock import pyEZmock
# %matplotlib inline
# -----
# The `pyEZmock` class should be initialised with the working directory, and optionally the locations of the EZmock and clustering codes, e.g.
#
# ```python
# ez = pyEZmock(workdir, exe='/path/to/EZmock', pk_exe='/path/to/powspec',
# xi_exe='/path/to/FCFC', bk_exe='/path/to/bispec')
# ```
#
# The `set_param` and `set_clustering` functions initialise parameters for EZmock construction and clustering measurements.
# For parameters that do not vary during the fit, it is recommended to set them at initialisation (via the `set_param` function).
# The clustering statistics to be computed, as well as the references for the fit, are defined by the `set_clustering` function. In particular, both real- and redshift-space clustering measurements are supported.
#
# Please consult `help(ez.set_param)` and `help(ez.set_clustering)` for detailed explanations of the parameters.
workdir = os.environ['SCRATCH'] + '/EZmock'
ez = pyEZmock(workdir)
ez.set_param(boxsize=1000, num_grid=256, redshift=0.9873, num_tracer=1000000, omega_m=0.3089,
init_pk='data/Planck15.UNIT.loguniform.pk')
ez.set_clustering(pk='both', pk_grid=512, xi='redshift', xi_z_ref_col=[1,2], bk='redshift', bk_grid=512,
pk_r_ref='test/PK_UNIT_DESI_Shadab_HOD_snap97_ELG_v1_4col_real.dat',
pk_z_ref='test/PK_UNIT_DESI_Shadab_HOD_snap97_ELG_v1_4col.dat',
xi_z_ref='test/2PCF_UNIT_DESI_Shadab_HOD_snap97_ELG_v1_4col.dat',
bk_z_ref='test/BK_UNIT_DESI_Shadab_HOD_snap97_ELG_v1_4col.dat')
# -----
# Free parameters for the fit can be set via the `run` function, which generates the job script for running EZmocks and clustering measurements. The job script can be either submitted to a slurm queue, or run manually when `queue` is not provided. And the typical 4 free parameters for EZmocks are listed below:
#
# | Parameter | Description |
# |---------------|------------------------------------------|
# | `pdf_base` | Base number for PDF mapping |
# | `dens_scat` | Density scattering scaling parameter |
# | `rand_motion` | Scatter of peculiar velocity |
# | `dens_cut` | Critical density for structure formation |
#
# For physical explanations of the parameters, see [arXiv:2007.08997](https://arxiv.org/abs/2007.08997). And the impacts of the parameters on clustering statistics can be found in the [\[cheatsheet\]](doc/EZmock_params_cheatsheet.pdf).
#
# In practice, the function `run` is expected to be called many times for the fit. However, it may take long before the job enters the queue. It is thus highly recommended to reserve an interactive node, and run the job manually there, by e.g.
#
# ```
# salloc -N 1 -q interactive -L SCRATCH -C haswell -t 4:00:00
# ```
# +
# Submit the job to the debug queue (not recommended for fit).
ez.run(nthreads=64, queue='debug', pdf_base=0.5, dens_scat=1, rand_motion=100, dens_cut=1.02)
# Provide a command for running the job manually (recommended with interactive nodes).
#ez.run(nthreads=64, pdf_base=0.5, dens_scat=1, rand_motion=100, dens_cut=0)
# -
# -----
# Plot clustering measurements set via the `set_clustering` function, obtained from the following sources in sequence:
#
# 1. previous runs;
# 2. the references;
# 3. the current run.
ez.plot()
# -----
# Continue running EZmocks and making plots after parameter revision.
ez.run(nthreads=64, pdf_base=0.2, dens_scat=1, rand_motion=100, dens_cut=1.02)
ez.plot()
ez.run(nthreads=64, pdf_base=0.2, dens_scat=0.5, rand_motion=100, dens_cut=1.02)
ez.plot()
ez.run(nthreads=64, pdf_base=0.2, dens_scat=0.5, rand_motion=200, dens_cut=1.02)
ez.plot()
# -----
# Generate the job for the mass production of mocks, with different random seeds. Note that it is possible to change the configurations for clustering measurements using `ez.set_clustering` before the mass production.
#
# If the `queue` argument is missing, a job list file for the [jobfork](https://github.com/cheng-zhao/jobfork) tool is generated. The MPI scheduler of this tool runs the list of jobs in parallel.
#
# If `queue` is provided, the job can be submitted by `sbatch <job_script>`, where `job_script` is the return value of `massive_jobs`. This job script runs multiple jobs in parallel, see the [NERSC docs](https://docs.nersc.gov/jobs/examples/#multiple-parallel-jobs-simultaneously).
# +
# Generate a job script for all realisations.
ez.massive_jobs(nthreads=32, seeds=range(4), queue='regular', clustering=False)
# Provide a job list file for the jobfork tool.
#ez.massive_jobs(nthreads=32, seeds=range(4), clustering=False)
# -
# -----
# Check parameters of the current (`params` function) and previous (`history` function) runs. The indices of the histories are consistent with the ones showing in the plots above.
ez.params()
print('----------')
ez.history()
# -----
# Clear the histories by slices.
ez.clear(slice(0,2)) # same as removing history[0:2]
#ez.clear(slice(None)) # remove all histories
ez.history()
# -----
# To recover histories, one could call the `restore` function, or reinitialise the `pyEZmock` class with the `restore` argument.
ez.restore()
#ez = pyEZmock(workdir, restore=True)
ez.history()
| example_pyez.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7
# language: python
# name: python3
# ---
# # Association of Research Libraries Data
# ### Introduction
# The Association of Research Libraries (ARL) is a voluntary professional organization of research intensive libraries across North America. These libraries can be public libraries (NYPL) and private libraries (the Smithsonian Institute) but the majority are at universities and colleges. Many libraries use the statistics generated by the ARL for benchmarking purposes, but often only use geography as the determining factor in comparisons. This can lead to very unfair and unreasonable comparisons between institutions, such as comparing University of Toronto with Queen's University because they are both in Ontario, despite the fact that U of T is more than 4 times the size. Because libraries represent a significant investment in research for an institution, for anyone who works in research directly, or who's business serves researchers, having a more accurate comparison would be helpful.
# This capstone project will take data from ARL to find university libraries with similar research profiles and map them so we can see which universities are similar. This could be used by people looking for employment in universities, or businesses who want to know if their company might be successful in a similar environment elsewhere.
# ### Data
# The data from ARL includes details like the number of students, both full time and part time, as well as the number of undergraduate and graduate students. It includes the number of full time faculty and whether the school has a medical or legal program. Other information includes the budgets of the university libraries, amount spent on renewable resources (eg. journal subscriptions) as compared to books and other one-time purchases, the staffing levels of the libraries, and what percentage are professional staff, support staff and student assistants.
# ### Methodology
# First we import the libraries we need:
import numpy as np
import pandas as pd
import matplotlib as mpl
import matplotlib.pyplot as plt
# Then we get the data:
# +
import types
from botocore.client import Config
import ibm_boto3
def __iter__(self): return 0
# @hidden_cell
# The following code accesses a file in your IBM Cloud Object Storage. It includes your credentials.
# You might want to remove those credentials before you share the notebook.
client_6202ed263f8f4f51983b845ba3b9469f = ibm_boto3.client(service_name='s3',
ibm_api_key_id='<KEY>',
ibm_auth_endpoint="https://iam.cloud.ibm.com/oidc/token",
config=Config(signature_version='oauth'),
endpoint_url='https://s3-api.us-geo.objectstorage.service.networklayer.com')
body = client_6202ed263f8f4f51983b845ba3b9469f.get_object(Bucket='arlstatistics-donotdelete-pr-kbcmhh3usmtlbb',Key='arl_statistics_data_download.csv')['Body']
# add missing __iter__ method, so pandas accepts body as file-like object
if not hasattr(body, "__iter__"): body.__iter__ = types.MethodType( __iter__, body )
# -
df_data_1 = pd.read_csv(body)
df_data_1.head()
# Great! Now looking at the year column, I can see quite a bit of this data is out of date. Let's just strip it to the most recent year available, which is 2019.
# +
df_data_1 = df_data_1.drop(df_data_1[df_data_1.year < 2019].index)
# -
df_data_1.head()
# Wow! Alabama has doubled its student population (totstu column) in the last 20 years. This is why we need to always be changing our comparators.
# As we also mentioned, not every library in the ARL is a university library. Because we're interested in making comparisons by student population size, let's get rid of institutions with no students
df_data_1.tail(10)
df_data_1.drop(df_data_1.tail(8).index,inplace=True) # drop last n rows
df_data_1.tail(10)
from sklearn.cluster import KMeans
# OK, there's a lot of columns we're not going to use here, so let's get rid of them. In fact, for this exercise, the only thing I'm really interested in is the number of faculty and students in each category. So everything else can go.
df_data_1.drop(columns =['year','instno', 'type', 'region', 'membyr', 'exch', 'rank2', 'in_index', 'rank1', 'index', 'volsadg', 'volsadn', 'volswdn', 'mono', 'serpur','sernpur', 'currser', 'microf', 'govdocs', 'mss', 'maps', 'graphic', 'audio', 'video', 'compfil', 'totcirc', 'expmono','expser', 'expoth', 'expmisc', 'expbnd', 'vols', 'illtot', 'ilbtot', 'grppres', 'presptcp', 'reftrans', 'initcirc', 'totstfx', 'explm', 'salprf', 'salnprf', 'salstud', 'totsal', 'opexp', 'totexp', 'phdawd', 'phdfld'
])
# Apparently a lot of these columns aren't integers, so let's change that.
df_data_1['totstu'] = df_data_1['totstu'].astype(int)
df_data_1['totpt'] = df_data_1['totpt'].astype(int)
df_data_1['gradstu'] = df_data_1['gradstu'].astype(int)
df_data_1['gradpt'] = df_data_1['gradpt'].astype(int)
df_data_1['fac'] = df_data_1['fac'].astype(int)
df_data_1.columns[1:]
df_data_1.columns = list(map(str, df_data_1.columns))
df_data_1.set_index('inam', inplace=True)
# OK, that's better. Now we can create a scatter plot that shows the ratio of students to faculty in ARL institutions. Note it isn't completely accurate. For some reason, ARL only asks for the number of full-time, tenure track faculty. If you've been following higher education for the last couple of decades, you'll know that there has been a remarkable increase in using adjunct faculty to do the work. It also doesn't include clinical faculty, which means if your school has a medical, nursing, dental or vetrinary school, it will be significantly underreporting the number of faculty. However, from the perspective of the library, this is significant because we serve all faculty, regardless of whether they are tenure track or adjuncts the same.
# +
df_data_1.plot(kind='scatter', x='totstu', y='fac', figsize=(10, 6), color='darkblue')
plt.title('Student Faculty Ratios')
plt.xlabel('Students')
plt.ylabel('Faculty')
plt.show()
# -
# Isn't that scatter plot a thing of beauty! Wow! There's some real outliers at the top end, places like U of T (80,000 students). You can see why even though they are also in Ontario, it doesn't make sense to compare ourselves to them.
#
# So now let's get down to the business of clustering our schools. We're going to use machine learning to classify schools based on similar characteristics - the number of students and faculty reported.
# +
from sklearn.cluster import DBSCAN
from sklearn.preprocessing import StandardScaler
from collections import Counter
from pylab import rcParams
rcParams['figure.figsize'] = 14, 6
# %matplotlib inline
# -
df_data_1.info()
# We need to turn all this data into a numpy array using integers, so we can normalize the data.
# Prepare data for model
dbscan_data = df_data_1[['totstu', 'fac']]
dbscan_data = dbscan_data.values.astype('int',)
dbscan_data
# Now let's normalize the data, so we aren't overcounting students and undercounting faculty. Again, we're still undercounting, because ARL doesn't look at adjuncts, but we can only go with the data we have.
# Normalize data
dbscan_data_scaler = StandardScaler().fit(dbscan_data)
dbscan_data = dbscan_data_scaler.transform(dbscan_data)
dbscan_data
# With DBSCAN, you need to guesstimate your epsilon based on your knowledge of the data. This was tricky, because we have a lot of outliers at the far reaches, but the rest of the schools are clustered pretty close together already. I kept the standard number of minimum samples at 4, but played a but with the epsilon until I got a number of clusters that seemed reasonable.
# +
# Construct model
'''
-- min_samples :: requires a minimum 20 data points in a neighborhood
-- eps :: in radius 0.02
'''
model = DBSCAN(eps = 0.18, min_samples = 4, metric='euclidean').\
fit(dbscan_data)
model
# +
# Visualize the results
# separate outliers from clustered data
outliers_df = dbscan_data[model.labels_ == -1]
clusters_df = dbscan_data[model.labels_ != -1]
colors = model.labels_
colors_clusters = colors[colors != -1]
color_outliers = 'black'
# Get info about the clusters
clusters = Counter(model.labels_)
print(clusters)
print('Number of clusters = {}'.format(len(clusters)-1))
# -
# Now let's take those clusters of universities and plot them on a scatter plot. I'll make the plot and dots a little bigger so we can see it better.
plt.scatter(dbscan_data[:,0], dbscan_data[:,1], s=100, c=model.labels_)
plt.gcf().set_size_inches((12, 12))
plt.show()
# There! Now we have 5 groups of universities that are similar. The teal ones, which are the smallest. The purple, which includes Queen's, which are medium sized and includes the largest number of similar universities. The yellow, which are 5 larger universities, and two groups of light and dark green clusters, which are larger still. The black dots are the outliers, which don't fit into any cluster.
# ### Results
# Now let's look at which schools are in which clusters. First, the thirteen teal schools:
# +
df_data_1["Clus_Db"]=model.labels_
df_data_1[df_data_1["Clus_Db"]==0]
df_data_1[df_data_1["Clus_Db"]==1]
# -
# Now the 35 purple schools:
# +
df_data_1["Clus_Db"]=model.labels_
df_data_1[df_data_1["Clus_Db"]==-1]
df_data_1[df_data_1["Clus_Db"]==0]
# -
# Now the four dark green schools:
# +
df_data_1["Clus_Db"]=model.labels_
df_data_1[df_data_1["Clus_Db"]==1]
df_data_1[df_data_1["Clus_Db"]==2]
# -
# The five yellow schools:
# +
df_data_1["Clus_Db"]=model.labels_
df_data_1[df_data_1["Clus_Db"]==2]
df_data_1[df_data_1["Clus_Db"]==3]
# -
# And the five light green schools:
# +
df_data_1["Clus_Db"]=model.labels_
df_data_1[df_data_1["Clus_Db"]==3]
df_data_1[df_data_1["Clus_Db"]==4]
# -
# # FourSquare data
# Now I'm going to try to use FourSquare data to compare the four dark green schools: University of Arizona, UCLA, University of Illinois at Urbana and USC. These four schools form the smallest cluster and might be a good opportunity to expand your business with a similar client base.
# +
import requests # library to handle requests
import pandas as pd # library for data analsysis
import numpy as np # library to handle data in a vectorized manner
import random # library for random number generation
# !pip install geopy
from geopy.geocoders import Nominatim # module to convert an address into latitude and longitude values
# libraries for displaying images
from IPython.display import Image
from IPython.core.display import HTML
# tranforming json file into a pandas dataframe library
from pandas.io.json import json_normalize
# ! pip install folium==0.5.0
import folium # plotting library
print('Folium installed')
print('Libraries imported.')
# -
CLIENT_ID = 'J3MOVTWUGSMGZC5KLYX4VKGUKQN3USXSKLBZBEVN2RWTDOUH' # your Foursquare ID
CLIENT_SECRET = '<KEY>' # your Foursquare Secret
ACCESS_TOKEN = 'ECOL5AE1ID2ZGTSRVR0D2FALGEN1EER225COPYCIJV4HO<PASSWORD>' # your FourSquare Access Token
VERSION = '20180604'
LIMIT = 30
print('Your credentails:')
print('CLIENT_ID: ' + CLIENT_ID)
print('CLIENT_SECRET:' + CLIENT_SECRET)
venue_id = '4b1f5dfbf964a520dd2524e3' # fIND THE VENUE IDS OF THE DIFFERENT COLLEGES
url = 'https://api.foursquare.com/v2/venues/{}?client_id={}&client_secret={}&oauth_token={}&v={}'.format(venue_id, CLIENT_ID, CLIENT_SECRET,ACCESS_TOKEN, VERSION)
url
| ARL Library comparisons.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/nforesperance/TensorFlow/blob/master/retrain_model.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="HWYjYJC4_KnK" colab_type="code" colab={}
try:
# # %tensorflow_version only exists in Colab.
# %tensorflow_version 2.x
except Exception:
pass
# !pip install pyyaml h5py # Required to save models in HDF5 format
# + id="W346i8kUCCxc" colab_type="code" outputId="d666935b-937e-480c-b4ca-7cea4ee524c5" colab={"base_uri": "https://localhost:8080/", "height": 34}
from __future__ import absolute_import, division, print_function, unicode_literals
import os
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras.layers import Conv2D,MaxPool2D,Flatten,Dense
print(tf.version.VERSION)
# + id="tPXqImHxCKWz" colab_type="code" colab={}
(train_images, train_labels), (test_images, test_labels) = tf.keras.datasets.mnist.load_data()
train_images = (train_images.reshape((train_images.shape[0], 28, 28, 1)))/ 255.0
test_images = (test_images.reshape((test_images.shape[0], 28, 28, 1))) / 255.0
test_images = test_images / 255.0
train_images = train_images / 255.0
# + id="0rpmwtRCCOIV" colab_type="code" colab={}
# Define a simple sequential model
# Define a simple sequential model
def create_model():
model = tf.keras.models.Sequential()
model.add(Conv2D(32, (3, 3), activation='relu', kernel_initializer='he_uniform', input_shape=(28, 28, 1)))
model.add(MaxPool2D((2, 2)))
model.add(Flatten())
model.add(Dense(100, activation='relu', kernel_initializer='he_uniform'))
model.add(Dense(10, activation='softmax'))
model.compile(optimizer='adam',
loss=tf.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
return model
# Create a basic model instance
model = create_model()
# + id="oENY5t5YzMHG" colab_type="code" colab={}
checkpoint_path = "trained0/cp.ckpt"
checkpoint_dir = os.path.dirname(checkpoint_path)
save_path = "trained0"
# + [markdown] id="d-g-PkXDyw_6" colab_type="text"
# ## N:B
# ###The following cell should be ran only once or model will retrain completely
# #### checkpoint is necessary if to be able to load best weights if there was an interuption before restore_best_weight
# + id="QHFyWizFCXMI" colab_type="code" colab={}
#N:B The Followind Code is ran only once
# Create a callback that saves the model's weights
easystopping = tf.keras.callbacks.EarlyStopping(monitor='val_loss', min_delta=0, patience=0, verbose=1, mode='min', baseline=None, restore_best_weights=True)
checkpoint = tf.keras.callbacks.ModelCheckpoint(filepath=checkpoint_path,
monitor='val_loss',
verbose=1,
save_weights_only=True,
save_best_only=True,
mode='min')
# Train the model with the new callback
model.fit(train_images, train_labels, validation_data=(train_images,
train_labels),
epochs=20,
batch_size=50,
verbose = 2,
callbacks=[checkpoint,easystopping])
# model.load_weights(checkpoint_path)
# model.save("trained1/best.h5")
# + id="nFJF-P-Xq_XX" colab_type="code" colab={}
model.save(save_path+"/best.h5")
# + [markdown] id="jYuhoskJxpHH" colab_type="text"
# ## Retraining
# ### The following is used to load the trained model and retrain it
#
# + id="sXyc1YCuChGH" colab_type="code" colab={}
for i in range (2):
del model
model = tf.keras.models.load_model(save_path+'/best.h5')
easystopping = tf.keras.callbacks.EarlyStopping(monitor='val_loss', min_delta=0, patience=0, verbose=1, mode='min', baseline=None, restore_best_weights=True)
checkpoint = tf.keras.callbacks.ModelCheckpoint(filepath=checkpoint_path,
monitor='val_loss',
verbose=1,
save_weights_only=True,
save_best_only=True,
mode='min')
# Train the model with the new callback
model.fit(train_images, train_labels, validation_data=(train_images,
train_labels),
epochs=20,
batch_size=50,
verbose = 2,
callbacks=[checkpoint,easystopping])
model.save(save_path+"/best.h5")
# + [markdown] id="Ad27yGP5x_eM" colab_type="text"
# ## Testing
# + id="H4-2gyiZEUdG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 84} outputId="05113f26-b5b2-4039-b14e-0b256f28b506"
# model.save("trained/best.h5")
del model
model = tf.keras.models.load_model(save_path+'/best.h5')
# Re-evaluate the model
loss,acc1 = model.evaluate(test_images, test_labels, verbose=2)
print("Restored model, accuracy: {:5.2f}%".format(100*acc1))
del model
model = create_model()
model.load_weights(checkpoint_path)
# Re-evaluate the model
loss,acc2 = model.evaluate(test_images, test_labels, verbose=2)
print("Restored model, accuracy: {:5.2f}%".format(100*acc2))
if acc2 > acc1:
model.save(save_path+"/best.h5")
print("Checkpoint Model Used")
| retrain_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img align="centre" src="../../figs/Github_banner.jpg" width="100%">
#
# # Southern Africa Cropland Mask
# ## Background
#
# The notebooks in this folder provide the means for generating a cropland mask (crop/non-crop) for the Southern Africa study region (Figure 1), for the year 2019 at 10m resolution. To obtain classifications a Random Forest algorithm is trained using training data in the `data/` folder (`Southern_training_data_<YYYYMMDD>.geojson`). The entire algorithm is summarised in figure 2.
#
# The definition of cropping used to collect the training data is:
#
# “A piece of land of minimum 0.16 ha that is sowed/planted and harvest-able at least once within the 12 months after the sowing/planting date.”
#
# This definition will exclude non-planted grazing lands and perennial crops which can be difficult for satellite imagery to differentiate from natural vegetation.
#
# _Figure 1: Study area for the notebooks in this workflow_
#
# <img align="center" src="../../figs/study_area_southern.png" width="700">
#
#
# _Figure 2: The algorithm used to generate the cropland mask for Southern Africa_
#
# <img align="center" src="../../figs/cropmask_algo_eastern.PNG" width="900">
#
# ---
# ## Getting Started
#
# There are six notebooks in this collection which, if run sequentially, will reproduce Digital Earth Africa's cropmask for the Southern region of Africa.
# To begin working through the notebooks in this `Southern Africa Cropland Mask` workflow, go to the first notebook `Extract_training_data.ipynb`.
#
# 1. [Extract_training_data](1_Extract_training_data.ipynb)
# 2. [Inspect_training_data](2_Inspect_training_data.ipynb)
# 3. [Train_fit_evaluate_classifier](3_Train_fit_evaluate_classifier.ipynb)
# 4. [Predict](4_Predict.ipynb)
# 5. [Accuracy_assessment](5_Accuracy_assessment.ipynb)
#
# ***
#
# ## Additional information
#
# **License:** The code in this notebook is licensed under the [Apache License, Version 2.0](https://www.apache.org/licenses/LICENSE-2.0).
# Digital Earth Africa data is licensed under the [Creative Commons by Attribution 4.0](https://creativecommons.org/licenses/by/4.0/) license.
#
# **Contact:** If you need assistance, please post a question on the [Open Data Cube Slack channel](http://slack.opendatacube.org/) or on the [GIS Stack Exchange](https://gis.stackexchange.com/questions/ask?tags=open-data-cube) using the `open-data-cube` tag (you can view previously asked questions [here](https://gis.stackexchange.com/questions/tagged/open-data-cube)).
# If you would like to report an issue with this notebook, you can file one on [Github](https://github.com/digitalearthafrica/deafrica-sandbox-notebooks).
#
| testing/southern_cropmask/0_README.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: text_analytics
# language: python
# name: text_analytics
# ---
# # Class of Week 7
# Content:
# 1. TF-IDF
# 2. Singular Value Decomposition
# 3. Latent Semantic Indexing
# 4. Latent Dirichlet Allocation
# 5. Open Questions and Project Support
# ## TF-IDF
# Term-Frequency-Inverse-Document Frequency.
# Bag-of-words approach that gives more weight to 'important' words.
# Useful resources:
# https://www.youtube.com/watch?v=4vT4fzjkGCQ
# https://www.youtube.com/watch?v=hXNbFNCgPfY
#
# ### Basic TF-IDF
# +
# Create two documents.
docs = ['the dog sits on the table',
'the cat sits on the sofa']
docs
# +
import collections
# Convert documents to counter.
docs_counter = [collections.Counter(doc.split()) for doc in docs]
docs_counter
# +
import itertools
# Create unique term set.
terms = set(itertools.chain.from_iterable(docs_counter))
terms
# +
# Create term-frequency function.
def tf(t, d):
"""Calculates term-frequency for term t in document d."""
# If term in document, return frequency. Else, return null:
if t in d.keys():
return d[t]
else:
return 0
print(tf('cat', docs_counter[0]))
print(tf('cat', docs_counter[1]))
# +
# Calculate term-frequency matrix.
tf_matrix = [{t:tf(t, d) for t in terms} for d in docs_counter]
tf_matrix
# +
import math
# Create inverse document-frequency function.
def idf(t, D):
"""Calculates inverse document-frequency for term t in documents D."""
return math.log(len(D) / len([d for d in D if t in d.keys()]),2)
print(idf('the', docs_counter))
print(idf('cat', docs_counter))
# +
# Calculate inverse document-frequency vector.
idf_vector = {t: idf(t, docs_counter) for t in terms}
idf_vector
# +
# Calculate term-frequency inverse document-frequency matrix.
tfidf_matrix = [{t: tf_vector[t]*idf_vector[t] for t in terms} for tf_vector in tf_matrix]
tfidf_matrix
# +
import pandas
# Label term-frequency columns.
tf_cols = ['tf_' + str(i + 1) for i in range(len(tf_matrix))]
# Labels term-frequency inverse-document-frequency columns.
tfidf_cols = ['tfidf_' + str(i + 1) for i in range(len(tfidf_matrix))]
# Create function to build pandas data frame.
def create_df(tf_matrix, idf_vector, tfidf_matrix, tf_cols, tfidf_cols):
# Create data frame dictionary.
df_dict = {}
# Fill data frame dictionary.
for tf_col, tf_vector in zip(tf_cols, tf_matrix): df_dict[tf_col] = tf_vector
df_dict['idf'] = idf_vector
for tfidf_col, tfidf_vector in zip(tfidf_cols, tfidf_matrix): df_dict[tfidf_col] = tfidf_vector
# Create column order.
col_order = []
col_order.extend(tf_cols)
col_order.append('idf')
col_order.extend(tfidf_cols)
# Create data frame and order by column order.
df = pandas.DataFrame.from_dict(df_dict)
df = df[col_order]
return df
# -
# Print data frame.
print('Standard TF-IDF\n')
print(create_df(tf_matrix, idf_vector, tfidf_matrix, tf_cols, tfidf_cols))
# ### Advanced TF-IDF
# Sub-linear term-frequency.
def tf(t, d, sub_linear=False):
"""Calculates term-frequency for term t in document d."""
# If term in document, return frequency. Else, return null:
if t in d.keys():
# If sub_linear, return log of tf.
if sub_linear:
return math.log(d[t]) + 1
else:
return d[t]
else:
return 0
# +
tf_matrix = [{t:tf(t, d, sub_linear=True) for t in terms} for d in docs_counter]
tfidf_matrix = [{t: tf_vector[t] * idf_vector[t] for t in terms} for tf_vector in tf_matrix]
print('Sub-Linear TF-IDF\n')
print(create_df(tf_matrix, idf_vector, tfidf_matrix, tf_cols, tfidf_cols))
# -
# Smoother for inverse document-frequency.
def idf(t, D, smoother=False):
"""Calculates inverse document-frequency for term t in documents D."""
val = len(D) / len([d for d in D if t in d.keys()])
# If smoother, add 1 to val
if smoother:
val += 1
return math.log(val, 2)
# +
idf_vector = {t: idf(t, docs_counter, True) for t in terms}
tfidf_matrix = [{t: tf_vector[t] * idf_vector[t] for t in terms} for tf_vector in tf_matrix]
print('Smoother TF-IDF\n')
print(create_df(tf_matrix, idf_vector, tfidf_matrix, tf_cols, tfidf_cols))
# +
import numpy
# Normalizing term-frequency.
def tf(t, d, sub_linear=False, normalization=None):
"""Calculates term-frequency for term t in document d."""
# If normalization is in ['l1', 'l2'], apply normalization.
if normalization in ['l1', 'l2']:
# If normalization is 'l1', apply l1 normalization.
if normalization == 'l1':
normalizer = numpy.sum(numpy.abs(list(d.values())))
# If normalization is 'l2', apply l2 normalization.
if normalization == 'l2':
normalizer = numpy.sqrt(numpy.sum(numpy.square(list(d.values()))))
d_norm = {word: d[word] / normalizer for word in d.keys()}
else:
d_norm = d
# If term in document, return frequency. Else, return null:
if t in d_norm.keys():
# If sub_linear, return log of tf.
if sub_linear:
return math.log(d_norm[t])
else:
return d_norm[t]
else:
return 0
# +
tf_matrix = [{t:tf(t, d, False, 'l1') for t in terms} for d in docs_counter]
tfidf_matrix = [{t: tf_vector[t] * idf_vector[t] for t in terms} for tf_vector in tf_matrix]
print('L1 TF-IDF\n')
print(create_df(tf_matrix, idf_vector, tfidf_matrix, tf_cols, tfidf_cols))
# +
tf_matrix = [{t:tf(t, d, False, 'l2') for t in terms} for d in docs_counter]
tfidf_matrix = [{t: tf_vector[t] * idf_vector[t] for t in terms} for tf_vector in tf_matrix]
print('L2 TF-IDF\n')
print(create_df(tf_matrix, idf_vector, tfidf_matrix, tf_cols, tfidf_cols))
# -
# ### TF-IDF with Gensim
# +
from gensim.models import TfidfModel
from gensim.corpora import Dictionary
docs_tokenized = [d.split() for d in docs]
dct = Dictionary(docs_tokenized) # fit dictionary
corpus = [dct.doc2bow(line) for line in docs_tokenized] # convert dataset to BoW format
model = TfidfModel(corpus) # fit model
vector = model[corpus[0]] # apply model
vector
# -
# ## Singular Value Decomposition
# Useful resources:
# https://www.youtube.com/watch?v=P5mlg91as1c
# +
movie_dict = {'matrix': [1, 3, 4, 5, 0, 0, 0],
'alien': [1, 3, 4, 5, 2, 0, 1],
'serenity': [1, 3, 4, 5, 0, 0, 0],
'casablanca': [0, 0, 0, 0, 4, 5, 2],
'amelie': [0, 0, 0, 0, 4, 5, 2]}
movie_matrix = pandas.DataFrame.from_dict(movie_dict)
print('\n\nOriginal Matrix\n')
print(movie_matrix)
# +
from sklearn.decomposition import TruncatedSVD
svd_model = TruncatedSVD(n_components=2)
svd_features = svd_model.fit_transform(movie_matrix)
print('SVD Features\n')
print(pandas.DataFrame(svd_features))
print('\n\nSVD Singular Values\n')
print(pandas.DataFrame(svd_model.singular_values_))
print('\n\nSVD Components\n')
print(pandas.DataFrame(svd_model.components_))
# -
# ## Latent Semantic Indexing
# Useful resources:
# https://www.youtube.com/watch?v=BJ0MnawUpaU
# +
from gensim import corpora, models
dictionary = corpora.Dictionary(docs_tokenized)
# print(dictionary.token2id)
corpus = [dictionary.doc2bow(text) for text in docs_tokenized]
# print(corpus)
tfidf = models.TfidfModel(corpus)
corpus_tfidf = tfidf[corpus]
total_topics = 2
lsi = models.LsiModel(corpus_tfidf,
id2word=dictionary,
num_topics=total_topics)
pandas.DataFrame(lsi.print_topics(total_topics))
df = pandas.DataFrame(lsi.projection.u)
df['term'] = lsi.id2word.id2token.values()
df = df.set_index('term')
print(df)
# +
from sklearn.decomposition import LatentDirichletAllocation
lda_model = LatentDirichletAllocation(n_components=2, doc_topic_prior=0.9, topic_word_prior=0.9)
lda_features = lda_model.fit_transform(movie_matrix)
print('\n\nLDA Features\n')
print(pandas.DataFrame(lda_features))
print('\n\nLDA Components\n')
print(pandas.DataFrame(lda_model.exp_dirichlet_component_))
# -
| notebooks/week_7/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Example block-wise adaptation
# ### Important: this notebook has been run by using data from Subject S9 of Dataset-1. Note that Table I of the manuscript show the average of these results overall subjects of Dataset-1.
# +
import numpy as np
from numpy import unravel_index
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.model_selection import train_test_split
from sklearn import preprocessing
import ot
import scipy.io
import mne
from mne.decoding import CSP
mne.set_log_level(verbose='warning') #to avoid info at terminal
import matplotlib.pyplot as pl
from random import seed
seed(30)
from MIOTDAfunctions import*
# get the functions from RPA package
import rpa.transfer_learning as TL
from pyriemann.classification import MDM
from pyriemann.estimation import Covariances
from pyriemann.utils.base import invsqrtm
import timeit
#ignore warning
from warnings import simplefilter
# ignore all future warnings
simplefilter(action='ignore', category=FutureWarning)
simplefilter(action='ignore', category=UserWarning)
# +
results_acc=[]
results_all=[]
results_all_inv=[]
rango_cl = [0.1, 0.5, 1, 2, 5, 10, 20]
rango_e = [0.1, 0.5, 1, 2, 5, 10, 20]
metric = 'sqeuclidean'
outerkfold = 10 # for faster online computation select a lower value
innerkfold = dict(nfold=10, train_size=0.8)
# -
# ## Methods definition
def SC(Gte, Yte, lda):
start = timeit.default_timer()
acc = lda.score(Gte, Yte)
stop = timeit.default_timer()
time = stop - start
return acc, time
def SR(Data_S2, Labels_S2, re, Xtr, Ytr, Xte, Yte):
start = timeit.default_timer()
#Get Data
Xtr2add = Data_S2[0:20*re+20]
Ytr2add = Labels_S2[0:20*re+20]
Xtr2 = np.vstack(((Xtr, Xtr2add)))
Ytr2 = np.hstack(((Ytr, Ytr2add)))
Ytr2 = Ytr2[len(Ytr2add):]
Xtr2 = Xtr2[len(Ytr2add):]
# Create a new CSP
csp = CSP(n_components=6, reg='empirical', log=True, norm_trace=False, cov_est='epoch')
#learn new csp filters
Gtr = csp.fit_transform(Xtr2,Ytr2)
#learn new lda
lda = LinearDiscriminantAnalysis()
lda.fit(Gtr, Ytr2)
# Apply on new test data
Gte = csp.transform(Xte)
#ldatest
acc = lda.score(Gte, Yte)
# time
stop = timeit.default_timer()
time = stop - start
return acc, time
def Sinkhorn_Transport(Gtr, Ytr, Gval, Yval, Gte, Yte, rango_e, metric, outerkfold, innerkfold, M):
lda = LinearDiscriminantAnalysis()
# Subset selection
G_FOTDAs_, Y_FOTDAs_, regu_FOTDAs_=\
SelectSubsetTraining_OTDAs(Gtr, Ytr, Gval, Yval, rango_e, lda, metric, outerkfold, innerkfold, M)
#time
start = timeit.default_timer()
Gtr_daot = G_FOTDAs_
Ytr_daot = Y_FOTDAs_
otda = ot.da.SinkhornTransport(metric=metric, reg_e=regu_FOTDAs_)
#learn the map
otda.fit(Xs=Gtr_daot, ys=Ytr_daot, Xt=Gval)
#apply the mapping over source data
transp_Xs = otda.transform(Xs=Gtr)
# train a new classifier bases upon the transform source data
lda.fit(transp_Xs, Ytr)
# Compute acc
yt_predict = lda.predict(Gte)
acc = accuracy_score(Yte, yt_predict)
# time
stop = timeit.default_timer()
time = stop - start
return acc, time
def GroupLasso_Transport(Gtr, Ytr, Gval, Yval, Gte, Yte, rango_e, rango_cl, metric, outerkfold, innerkfold, M):
lda = LinearDiscriminantAnalysis()
# Subset selection
G_FOTDAl1l2_, Y_FOTDAl1l2_, regu_FOTDAl1l2_=\
SelectSubsetTraining_OTDAl1l2(Gtr, Ytr, Gval, Yval, rango_e, rango_cl, lda, metric, outerkfold, innerkfold, M)
#time
start = timeit.default_timer()
Gtr_daot = G_FOTDAl1l2_
Ytr_daot = Y_FOTDAl1l2_
otda = ot.da.SinkhornL1l2Transport(metric = metric ,reg_e = regu_FOTDAl1l2_[0], reg_cl = regu_FOTDAl1l2_[1])
otda.fit(Xs=Gtr_daot, ys=Ytr_daot, Xt=Gval)
#transport taget samples onto source samples
transp_Xs = otda.transform(Xs=Gtr)
# train a new classifier bases upon the transform source data
lda.fit(transp_Xs,Ytr)
# Compute acc
yt_predict = lda.predict(Gte)
acc = accuracy_score(Yte, yt_predict)
# time
stop = timeit.default_timer()
time = stop - start
return acc, time
def Backward_Sinkhorn_Transport(Gtr, Ytr, Gval, Yval, Gte, Yte, rango_e, lda, metric, outerkfold, innerkfold, M):
# the classifier already trained is an input of the function
# Subset selection
G_BOTDAs_, Y_BOTDAs_, regu_BOTDAs_=\
SelectSubsetTraining_BOTDAs(Gtr, Ytr, Gval, Yval, rango_e, lda, metric, outerkfold, innerkfold, M)
# time
start = timeit.default_timer()
Gtr_botda = G_BOTDAs_
Ytr_botda = Y_BOTDAs_
# Transport plan
botda = ot.da.SinkhornTransport(metric=metric, reg_e=regu_BOTDAs_)
botda.fit(Xs=Gval, ys=Yval, Xt=Gtr_botda)
#transport testing samples
transp_Xt_backward = botda.transform(Xs=Gte)
# Compute accuracy without retraining
yt_predict = lda.predict(transp_Xt_backward)
acc = accuracy_score(Yte, yt_predict)
# time
stop = timeit.default_timer()
time = stop - start
return acc, time
def Backward_GroupLasso_Transport(Gtr, Ytr, Gval, Yval, Gte, Yte, rango_e, rango_cl, lda, metric, outerkfold, innerkfold, M):
# the classifier already trained is an input of the function
# Subset selection
G_BOTDAl1l2_, Y_BOTDAl1l2_, regu_BOTDAl1l2_=\
SelectSubsetTraining_BOTDAl1l2(Gtr, Ytr, Gval, Yval, rango_e, rango_cl, lda, metric, outerkfold, innerkfold, M)
#time
start = timeit.default_timer()
Gtr_botda = G_BOTDAl1l2_
Ytr_botda = Y_BOTDAl1l2_
botda = ot.da.SinkhornL1l2Transport(metric=metric, reg_e=regu_BOTDAl1l2_[0], reg_cl=regu_BOTDAl1l2_[1])
botda.fit(Xs=Gval, ys=Yval, Xt=Gtr_botda)
#transport testing samples
transp_Xt_backward=botda.transform(Xs=Gte)
# Compute accuracy without retraining
yt_predict = lda.predict(transp_Xt_backward)
acc = accuracy_score(Yte, yt_predict)
# time
stop = timeit.default_timer()
time = stop - start
return acc, time
def RPA(Xtr,Xval,Xte,Ytr,Yval,Yte):
# time
start = timeit.default_timer()
# cov matrix estimation
cov_tr = Covariances().transform(Xtr)
cov_val= Covariances().transform(Xval)
cov_te = Covariances().transform(Xte)
clf = MDM()
source={'covs':cov_tr, 'labels': Ytr}
target_org_train={'covs':cov_val, 'labels': Yval}
target_org_test={'covs':cov_te, 'labels': Yte}
# re-centered matrices
source_rct, target_rct_train, target_rct_test = TL.RPA_recenter(source, target_org_train, target_org_test)
# stretched the re-centered matrices
source_rcs, target_rcs_train, target_rcs_test = TL.RPA_stretch(source_rct, target_rct_train, target_rct_test)
# rotate the re-centered-stretched matrices using information from classes
source_rpa, target_rpa_train, target_rpa_test = TL.RPA_rotate(source_rcs, target_rcs_train, target_rcs_test)
# get data
covs_source, y_source = source_rpa['covs'], source_rpa['labels']
covs_target_train, y_target_train = target_rpa_train['covs'], target_rpa_train['labels']
covs_target_test, y_target_test = target_rpa_test['covs'], target_rpa_test['labels']
# append train and validation data
covs_train = np.concatenate([covs_source, covs_target_train])
y_train = np.concatenate([y_source, y_target_train])
# train
clf.fit(covs_train, y_train)
# test
covs_test = covs_target_test
y_test = y_target_test
y_pred = clf.predict(covs_test)
#acc
acc = accuracy_score(Yte, y_pred)
# time
stop = timeit.default_timer()
time = stop - start
return acc, time
def EU(Xtr,Xval,Xte,Ytr,Yval,Yte):
# time
start = timeit.default_timer()
# Estimate single trial covariance
cov_tr = Covariances().transform(Xtr)
cov_val= Covariances().transform(Xval)
Ctr = cov_tr.mean(0)
Cval = cov_val.mean(0)
# aligment
Xtr_eu = np.asarray([np.dot(invsqrtm(Ctr), epoch) for epoch in Xtr])
Xval_eu = np.asarray([np.dot(invsqrtm(Cval), epoch) for epoch in Xval])
Xte_eu = np.asarray([np.dot(invsqrtm(Cval), epoch) for epoch in Xte])
# append train and validation data
x_train = np.concatenate([Xtr_eu, Xval_eu])
y_train = np.concatenate([Ytr, Yval])
# train new csp+lda
csp = CSP(n_components=6, reg='empirical', log=True, norm_trace=False, cov_est='epoch')
# learn csp filters
Gtr = csp.fit_transform(x_train,y_train)
# learn lda
lda = LinearDiscriminantAnalysis()
lda.fit(Gtr,y_train)
# test
Gte = csp.transform(Xte_eu)
# acc
acc = lda.score(Gte, Yte)
# time
stop = timeit.default_timer()
time = stop - start
return acc, time
# ## Load and filter data
# +
fName = 'Data/DataSession1_S9.mat'
s = scipy.io.loadmat(fName)
Data_S1=s["X"]
Labels_S1=s["y"]
Labels_S1=np.squeeze(Labels_S1)
#filterting with mne
[nt, nc, ns]=np.shape(Data_S1)
Data_S1=np.reshape(Data_S1, [nt, nc*ns])
Data_S1=mne.filter.filter_data(Data_S1, 128, 8, 30)
Data_S1=np.reshape(Data_S1, [nt,nc,ns])
fName = 'Data/DataSession2_S9.mat'
s2 = scipy.io.loadmat(fName)
Data_S2=s2["X"]
Labels_S2=s2["y"]
Labels_S2=np.squeeze(Labels_S2)
#filterting with mne
[nt, nc, ns]=np.shape(Data_S2)
Data_S2=np.reshape(Data_S2, [nt, nc*ns])
Data_S2=mne.filter.filter_data(Data_S2, 128, 8, 30)
Data_S2=np.reshape(Data_S2, [nt,nc,ns])
# -
# ### Learn CSP+LDA from source data (Data_S1)
Xtr = Data_S1
Ytr = Labels_S1
csp = CSP(n_components=6, reg='empirical', log=True, norm_trace=False, cov_est='epoch')
#learn csp filters
Gtr = csp.fit_transform(Xtr, Ytr)
#learn lda
lda = LinearDiscriminantAnalysis()
lda.fit(Gtr,Ytr)
# ### For each run of 20 trials each, make the data adaptation
# #### (Be patient, running this cell can take around 30 min)
for re in range(0,7):
print('Running testing RUN={:1.0f}'.format(re))
#testing run
Xte = Data_S2[0+20*(re+1):20*(re+1)+20]
Yte = Labels_S2[0+20*(re+1):20*(re+1)+20]
#transportation set-prior data
Xval = Data_S2[0:20*re+20]
Yval = Labels_S2[0:20*re+20]
#feature computation
Gval = csp.transform(Xval)
Gte = csp.transform(Xte)
M = len(Yval) #for the source subset selection
# SC
acc_sc, time_sc = SC(Gte, Yte, lda)
# SR
acc_sr, time_sr = SR(Data_S2, Labels_S2, re, Xtr, Ytr, Xte, Yte)
#%% # Sinkhorn Transport
acc_fotdas, time_fs = Sinkhorn_Transport(Gtr, Ytr, Gval, Yval, Gte, Yte, rango_e, metric, outerkfold, innerkfold, M)
#%% # Group-Lasso Transport
acc_fotdal1l2, time_fg = GroupLasso_Transport(Gtr, Ytr, Gval, Yval, Gte, Yte, rango_e, rango_cl, metric, outerkfold, innerkfold, M)
#%% # Backward Sinkhorn Transport
acc_botdas, time_bs = Backward_Sinkhorn_Transport(Gtr, Ytr, Gval, Yval, Gte, Yte, rango_e, lda, metric, outerkfold, innerkfold, M)
#%% # Backward Group-Lasso Transport
acc_botdal1l2, time_bg = Backward_GroupLasso_Transport(Gtr, Ytr, Gval, Yval, Gte, Yte, rango_e, rango_cl, lda, metric, outerkfold, innerkfold, M)
# Riemann
acc_rpa, time_rpa = RPA(Xtr, Xval, Xte, Ytr, Yval, Yte)
# Euclidean
acc_eu, time_eu = EU(Xtr, Xval, Xte, Ytr, Yval, Yte)
# print results
# accuracy
acc = {}
acc["sc"] = acc_sc
acc["sr"] = acc_sr
acc["rpa"] = acc_rpa
acc["ea"] = acc_eu
acc["fotda_s"] = acc_fotdas
acc["fotda_l1l2"] = acc_fotdal1l2
acc["botda_s"] = acc_botdas
acc["botda_l1l2"] = acc_botdal1l2
# computing time
time = {}
time["sr"] = round(time_sr,3)
time["rpa"] = round(time_rpa,3)
time["eu"] = round(time_eu,3)
time["fotda_s"] = round(time_fs,3)
time["fotda_l1l2"] = round(time_fg,3)
time["botda_s"] = round(time_bs,3)
time["botda_l1l2"] = round(time_bg,3)
row_format ="{:>12}" * (len(acc.keys()) + 1)
values = [ '%.2f' % elem for elem in list(acc.values())]
print("ACCURACY")
print(row_format.format("", *acc.keys()))
print(row_format.format("", *values))
row_format ="{:>15}" * (len(time.keys()) + 1)
values = [ '%.2f' % elem for elem in list(time.values())]
print("ADAPTIVE TIME")
print(row_format.format(" ", *time.keys()))
print(row_format.format("", *values))
| paper_example_blockwise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:dankd]
# language: python
# name: conda-env-dankd-py
# ---
# +
import re
import multiprocessing as mul
from multiprocessing.dummy import Pool
import numpy as np
import pandas as pd
from tqdm import tqdm
from dankypipe import pipe
tqdm.pandas()
def isfloat(x):
try:
float(x)
return True
except:
return False
def isint(x):
try:
int(x)
return True
except:
return False
# +
train = pd.read_pickle('train.pickle').sort_values(by='AvSigVersion')
test = pd.read_pickle('test.pickle').sort_values(by='AvSigVersion')
train = train.rename(columns={'HasDetections':'Target'})
# + code_folding=[2]
def transform(df):
df = df.copy()
df['MajorEngineVersion'] = df.EngineVersion.apply(lambda x: int(x.split('.')[2]))
df['MinorEngineVersion'] = df.EngineVersion.apply(lambda x: int(x.split('.')[3]))
df['EngineVersion_float'] = df.EngineVersion.apply(lambda x: float('.'.join(x.split('.')[2:])))
df['MajorAppVersion'] = df.AppVersion.apply(lambda x: int(x.split('.')[1]))
df['MinorAppVersion'] = df.AppVersion.apply(lambda x: x.split('.')[2])
mlen = np.max([len(v) for v in df['MinorAppVersion']])
df['MinorAppVersion'] = df.MinorAppVersion.apply(lambda x: int(f'1{x.zfill(mlen)}'))
df['FinestAppVersion'] = df.AppVersion.apply(lambda x: x.split('.')[3])
mlen = np.max([len(v) for v in df['FinestAppVersion']])
df['FinestAppVersion'] = df.FinestAppVersion.apply(lambda x: int(f'1{x.zfill(mlen)}'))
df['AppVersion_float'] = [
float(f'{t[0]}.{t[1]}{t[2]}') for t in df[
['MajorAppVersion', 'MinorAppVersion', 'FinestAppVersion']
].itertuples()
]
def intx(x, i):
x = x.split('.')[i]
return int(x) if isint(x) else np.nan
def floatx(x, i, j=None):
if j is not None:
x = '.'.join(x.split('.')[i:j])
else:
x = '.'.join(x.split('.')[i:])
return float(x) if isfloat(x) else np.nan
df['MajorAvSigVersion'] = df.AvSigVersion.apply(intx, i=1)
df['MinorAvSigVersion'] = df.AvSigVersion.apply(intx, i=2)
df['AvSigVersion_float'] = df.AvSigVersion.apply(floatx, i=1, j=3)
df['Census_MajorOSVersion'] = df.Census_OSVersion.apply(intx, i=2)
df['Census_MinorOSVersion'] = df.Census_OSVersion.apply(intx, i=3)
df['Census_OSVersion_float'] = df.Census_OSVersion.apply(floatx, i=2)
return df
# -
cols = ['EngineVersion', 'AppVersion', 'AvSigVersion', 'Census_OSVersion', 'MachineIdentifier']
train = train[cols+['Target']]
test = test[cols]
train = transform(train)
test = transform(test)
train.drop(columns=['EngineVersion', 'AppVersion', 'Census_OSVersion', 'AvSigVersion'], inplace=True, errors='ignore')
test.drop(columns=['EngineVersion', 'AppVersion', 'Census_OSVersion', 'AvSigVersion'], inplace=True, errors='ignore')
train = train.sort_values(by='AvSigVersion_float')
test = test.sort_values(by='AvSigVersion_float')
train.head()
test.head()
val_idx = int(len(train)*.7)
val_idx
# +
pbar = tqdm(total=len(train.columns)-2)
#for c in train.columns:
def fx(c):
if c == 'MachineIdentifier' or c == 'Target':
return
train_ = train[['MachineIdentifier', c]].iloc[:val_idx, :]
val_ = train[['MachineIdentifier', c]].iloc[val_idx:, :]
test_ = test[['MachineIdentifier', c]]
try:
pipe.upload_feature(c, (train_, val_, test_), overwrite=False)
except ValueError:
pass
pbar.update(1)
pool = Pool(mul.cpu_count())
pool.map(fx, train.columns.tolist())
pool.close()
pool.join()
| notebooks/feature_engineering/Versions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <!--BOOK_INFORMATION-->
# <img align="left" style="padding-right:10px;" src="figures/PDSH-cover-small.png">
# *This notebook contains an excerpt from the [Python Data Science Handbook](http://shop.oreilly.com/product/0636920034919.do) by <NAME>; the content is available [on GitHub](https://github.com/jakevdp/PythonDataScienceHandbook).*
#
# *The text is released under the [CC-BY-NC-ND license](https://creativecommons.org/licenses/by-nc-nd/3.0/us/legalcode), and code is released under the [MIT license](https://opensource.org/licenses/MIT). If you find this content useful, please consider supporting the work by [buying the book](http://shop.oreilly.com/product/0636920034919.do)!*
# # Data Manipulation with Pandas
# In the previous chapter, we dove into detail on NumPy and its ``ndarray`` object, which provides efficient storage and manipulation of dense typed arrays in Python.
# Here we'll build on this knowledge by looking in detail at the data structures provided by the Pandas library.
# Pandas is a newer package built on top of NumPy, and provides an efficient implementation of a ``DataFrame``.
# ``DataFrame``s are essentially multidimensional arrays with attached row and column labels, and often with heterogeneous types and/or missing data.
# As well as offering a convenient storage interface for labeled data, Pandas implements a number of powerful data operations familiar to users of both database frameworks and spreadsheet programs.
#
# As we saw, NumPy's ``ndarray`` data structure provides essential features for the type of clean, well-organized data typically seen in numerical computing tasks.
# While it serves this purpose very well, its limitations become clear when we need more flexibility (e.g., attaching labels to data, working with missing data, etc.) and when attempting operations that do not map well to element-wise broadcasting (e.g., groupings, pivots, etc.), each of which is an important piece of analyzing the less structured data available in many forms in the world around us.
# Pandas, and in particular its ``Series`` and ``DataFrame`` objects, builds on the NumPy array structure and provides efficient access to these sorts of "data munging" tasks that occupy much of a data scientist's time.
#
# In this chapter, we will focus on the mechanics of using ``Series``, ``DataFrame``, and related structures effectively.
# We will use examples drawn from real datasets where appropriate, but these examples are not necessarily the focus.
# ## Reminder about Built-In Documentation
#
# As you read through this chapter, don't forget that IPython gives you the ability to quickly explore the contents of a package (by using the tab-completion feature) as well as the documentation of various functions (using the ``?`` character). (Refer back to [Help and Documentation in IPython](01.01-Help-And-Documentation.ipynb) if you need a refresher on this.)
#
# For example, to display all the contents of the pandas namespace, you can type
#
# ```ipython
# In [3]: pd.<TAB>
# ```
#
# And to display Pandas's built-in documentation, you can use this:
#
# ```ipython
# In [4]: pd?
# ```
#
# More detailed documentation, along with tutorials and other resources, can be found at http://pandas.pydata.org/.
# # Introducing Pandas Objects
# At the very basic level, Pandas objects can be thought of as enhanced versions of NumPy structured arrays in which the rows and columns are identified with labels rather than simple integer indices.
# As we will see during the course of this chapter, Pandas provides a host of useful tools, methods, and functionality on top of the basic data structures, but nearly everything that follows will require an understanding of what these structures are.
# Thus, before we go any further, let's introduce these three fundamental Pandas data structures: the ``Series``, ``DataFrame``, and ``Index``.
#
# We will start our code sessions with the standard NumPy and Pandas imports:
import numpy as np
import pandas as pd
# ## The Pandas Series Object
#
# A Pandas ``Series`` is a one-dimensional array of indexed data.
# It can be created from a list or array as follows:
data = pd.Series([0.25, 0.5, 0.75, 1.0])
data
# As we see in the output, the ``Series`` wraps both a sequence of values and a sequence of indices, which we can access with the ``values`` and ``index`` attributes.
# The ``values`` are simply a familiar NumPy array:
data.values
# The ``index`` is an array-like object of type ``pd.Index``, which we'll discuss in more detail momentarily.
data.index
# Like with a NumPy array, data can be accessed by the associated index via the familiar Python square-bracket notation:
data[1]
data[1:3]
# As we will see, though, the Pandas ``Series`` is much more general and flexible than the one-dimensional NumPy array that it emulates.
# ### ``Series`` as generalized NumPy array
# From what we've seen so far, it may look like the ``Series`` object is basically interchangeable with a one-dimensional NumPy array.
# The essential difference is the presence of the index: while the Numpy Array has an *implicitly defined* integer index used to access the values, the Pandas ``Series`` has an *explicitly defined* index associated with the values.
#
# This explicit index definition gives the ``Series`` object additional capabilities. For example, the index need not be an integer, but can consist of values of any desired type.
# For example, if we wish, we can use strings as an index:
data = pd.Series([0.25, 0.5, 0.75, 1.0],
index=['a', 'b', 'c', 'd'])
data
# And the item access works as expected:
data['b']
# We can even use non-contiguous or non-sequential indices:
data = pd.Series([0.25, 0.5, 0.75, 1.0],
index=[2, 5, 3, 7])
data
data[5]
# ### Series as specialized dictionary
#
# In this way, you can think of a Pandas ``Series`` a bit like a specialization of a Python dictionary.
# A dictionary is a structure that maps arbitrary keys to a set of arbitrary values, and a ``Series`` is a structure which maps typed keys to a set of typed values.
# This typing is important: just as the type-specific compiled code behind a NumPy array makes it more efficient than a Python list for certain operations, the type information of a Pandas ``Series`` makes it much more efficient than Python dictionaries for certain operations.
#
# The ``Series``-as-dictionary analogy can be made even more clear by constructing a ``Series`` object directly from a Python dictionary:
population_dict = {'California': 38332521,
'Texas': 26448193,
'New York': 19651127,
'Florida': 19552860,
'Illinois': 12882135}
population = pd.Series(population_dict)
population
# By default, a ``Series`` will be created where the index is drawn from the sorted keys.
# From here, typical dictionary-style item access can be performed:
population['California']
# Unlike a dictionary, though, the ``Series`` also supports array-style operations such as slicing:
population['California':'Illinois']
# We'll discuss some of the quirks of Pandas indexing and slicing in [Data Indexing and Selection](03.02-Data-Indexing-and-Selection.ipynb).
# ### Constructing Series objects
#
# We've already seen a few ways of constructing a Pandas ``Series`` from scratch; all of them are some version of the following:
#
# ```python
# >>> pd.Series(data, index=index)
# ```
#
# where ``index`` is an optional argument, and ``data`` can be one of many entities.
#
# For example, ``data`` can be a list or NumPy array, in which case ``index`` defaults to an integer sequence:
pd.Series([2, 4, 6])
# ``data`` can be a scalar, which is repeated to fill the specified index:
pd.Series(5, index=[100, 200, 300])
# ``data`` can be a dictionary, in which ``index`` defaults to the sorted dictionary keys:
pd.Series({2:'a', 1:'b', 3:'c'})
# In each case, the index can be explicitly set if a different result is preferred:
pd.Series({2:'a', 1:'b', 3:'c'}, index=[3, 2])
# Notice that in this case, the ``Series`` is populated only with the explicitly identified keys.
# ## The Pandas DataFrame Object
#
# The next fundamental structure in Pandas is the ``DataFrame``.
# Like the ``Series`` object discussed in the previous section, the ``DataFrame`` can be thought of either as a generalization of a NumPy array, or as a specialization of a Python dictionary.
# We'll now take a look at each of these perspectives.
# ### DataFrame as a generalized NumPy array
# If a ``Series`` is an analog of a one-dimensional array with flexible indices, a ``DataFrame`` is an analog of a two-dimensional array with both flexible row indices and flexible column names.
# Just as you might think of a two-dimensional array as an ordered sequence of aligned one-dimensional columns, you can think of a ``DataFrame`` as a sequence of aligned ``Series`` objects.
# Here, by "aligned" we mean that they share the same index.
#
# To demonstrate this, let's first construct a new ``Series`` listing the area of each of the five states discussed in the previous section:
area_dict = {'California': 423967, 'Texas': 695662, 'New York': 141297,
'Florida': 170312, 'Illinois': 149995}
area = pd.Series(area_dict)
area
# Now that we have this along with the ``population`` Series from before, we can use a dictionary to construct a single two-dimensional object containing this information:
states = pd.DataFrame({'population': population,
'area': area})
states
# Like the ``Series`` object, the ``DataFrame`` has an ``index`` attribute that gives access to the index labels:
states.index
# Additionally, the ``DataFrame`` has a ``columns`` attribute, which is an ``Index`` object holding the column labels:
states.columns
# Thus the ``DataFrame`` can be thought of as a generalization of a two-dimensional NumPy array, where both the rows and columns have a generalized index for accessing the data.
# ### DataFrame as specialized dictionary
#
# Similarly, we can also think of a ``DataFrame`` as a specialization of a dictionary.
# Where a dictionary maps a key to a value, a ``DataFrame`` maps a column name to a ``Series`` of column data.
# For example, asking for the ``'area'`` attribute returns the ``Series`` object containing the areas we saw earlier:
states['area']
# Notice the potential point of confusion here: in a two-dimesnional NumPy array, ``data[0]`` will return the first *row*. For a ``DataFrame``, ``data['col0']`` will return the first *column*.
# Because of this, it is probably better to think about ``DataFrame``s as generalized dictionaries rather than generalized arrays, though both ways of looking at the situation can be useful.
# We'll explore more flexible means of indexing ``DataFrame``s in [Data Indexing and Selection](03.02-Data-Indexing-and-Selection.ipynb).
# ### Constructing DataFrame objects
#
# A Pandas ``DataFrame`` can be constructed in a variety of ways.
# Here we'll give several examples.
# #### From a single Series object
#
# A ``DataFrame`` is a collection of ``Series`` objects, and a single-column ``DataFrame`` can be constructed from a single ``Series``:
pd.DataFrame(population, columns=['population'])
# #### From a list of dicts
#
# Any list of dictionaries can be made into a ``DataFrame``.
# We'll use a simple list comprehension to create some data:
data = [{'a': i, 'b': 2 * i}
for i in range(3)]
pd.DataFrame(data)
# Even if some keys in the dictionary are missing, Pandas will fill them in with ``NaN`` (i.e., "not a number") values:
pd.DataFrame([{'a': 1, 'b': 2}, {'b': 3, 'c': 4}])
# #### From a dictionary of Series objects
#
# As we saw before, a ``DataFrame`` can be constructed from a dictionary of ``Series`` objects as well:
pd.DataFrame({'population': population,
'area': area})
# #### From a two-dimensional NumPy array
#
# Given a two-dimensional array of data, we can create a ``DataFrame`` with any specified column and index names.
# If omitted, an integer index will be used for each:
pd.DataFrame(np.random.rand(3, 2),
columns=['foo', 'bar'],
index=['a', 'b', 'c'])
# #### From a NumPy structured array
#
# We covered structured arrays in [Structured Data: NumPy's Structured Arrays](02.09-Structured-Data-NumPy.ipynb).
# A Pandas ``DataFrame`` operates much like a structured array, and can be created directly from one:
A = np.zeros(3, dtype=[('A', 'i8'), ('B', 'f8')])
A
pd.DataFrame(A)
# ## The Pandas Index Object
#
# We have seen here that both the ``Series`` and ``DataFrame`` objects contain an explicit *index* that lets you reference and modify data.
# This ``Index`` object is an interesting structure in itself, and it can be thought of either as an *immutable array* or as an *ordered set* (technically a multi-set, as ``Index`` objects may contain repeated values).
# Those views have some interesting consequences in the operations available on ``Index`` objects.
# As a simple example, let's construct an ``Index`` from a list of integers:
ind = pd.Index([2, 3, 5, 7, 11])
ind
# ### Index as immutable array
#
# The ``Index`` in many ways operates like an array.
# For example, we can use standard Python indexing notation to retrieve values or slices:
ind[1]
ind[::2]
# ``Index`` objects also have many of the attributes familiar from NumPy arrays:
print(ind.size, ind.shape, ind.ndim, ind.dtype)
# One difference between ``Index`` objects and NumPy arrays is that indices are immutable–that is, they cannot be modified via the normal means:
ind[1] = 0
# This immutability makes it safer to share indices between multiple ``DataFrame``s and arrays, without the potential for side effects from inadvertent index modification.
# ### Index as ordered set
#
# Pandas objects are designed to facilitate operations such as joins across datasets, which depend on many aspects of set arithmetic.
# The ``Index`` object follows many of the conventions used by Python's built-in ``set`` data structure, so that unions, intersections, differences, and other combinations can be computed in a familiar way:
indA = pd.Index([1, 3, 5, 7, 9])
indB = pd.Index([2, 3, 5, 7, 11])
indA & indB # intersection
indA | indB # union
indA ^ indB # symmetric difference
# These operations may also be accessed via object methods, for example ``indA.intersection(indB)``.
# # Data Indexing and Selection
# In [Chapter 2](02.00-Introduction-to-NumPy.ipynb), we looked in detail at methods and tools to access, set, and modify values in NumPy arrays.
# These included indexing (e.g., ``arr[2, 1]``), slicing (e.g., ``arr[:, 1:5]``), masking (e.g., ``arr[arr > 0]``), fancy indexing (e.g., ``arr[0, [1, 5]]``), and combinations thereof (e.g., ``arr[:, [1, 5]]``).
# Here we'll look at similar means of accessing and modifying values in Pandas ``Series`` and ``DataFrame`` objects.
# If you have used the NumPy patterns, the corresponding patterns in Pandas will feel very familiar, though there are a few quirks to be aware of.
#
# We'll start with the simple case of the one-dimensional ``Series`` object, and then move on to the more complicated two-dimesnional ``DataFrame`` object.
# ## Data Selection in Series
#
# As we saw in the previous section, a ``Series`` object acts in many ways like a one-dimensional NumPy array, and in many ways like a standard Python dictionary.
# If we keep these two overlapping analogies in mind, it will help us to understand the patterns of data indexing and selection in these arrays.
# ### Series as dictionary
#
# Like a dictionary, the ``Series`` object provides a mapping from a collection of keys to a collection of values:
import pandas as pd
data = pd.Series([0.25, 0.5, 0.75, 1.0],
index=['a', 'b', 'c', 'd'])
data
data['b']
# We can also use dictionary-like Python expressions and methods to examine the keys/indices and values:
'a' in data
data.keys()
list(data.items())
# ``Series`` objects can even be modified with a dictionary-like syntax.
# Just as you can extend a dictionary by assigning to a new key, you can extend a ``Series`` by assigning to a new index value:
data['e'] = 1.25
data
# This easy mutability of the objects is a convenient feature: under the hood, Pandas is making decisions about memory layout and data copying that might need to take place; the user generally does not need to worry about these issues.
# ### Series as one-dimensional array
# A ``Series`` builds on this dictionary-like interface and provides array-style item selection via the same basic mechanisms as NumPy arrays – that is, *slices*, *masking*, and *fancy indexing*.
# Examples of these are as follows:
# slicing by explicit index
data['a':'c']
# slicing by implicit integer index
data[0:2]
# masking
data[(data > 0.3) & (data < 0.8)]
# fancy indexing
data[['a', 'e']]
# Among these, slicing may be the source of the most confusion.
# Notice that when slicing with an explicit index (i.e., ``data['a':'c']``), the final index is *included* in the slice, while when slicing with an implicit index (i.e., ``data[0:2]``), the final index is *excluded* from the slice.
# ### Indexers: loc, iloc, and ix
#
# These slicing and indexing conventions can be a source of confusion.
# For example, if your ``Series`` has an explicit integer index, an indexing operation such as ``data[1]`` will use the explicit indices, while a slicing operation like ``data[1:3]`` will use the implicit Python-style index.
data = pd.Series(['a', 'b', 'c'], index=[1, 3, 5])
data
# explicit index when indexing
data[1]
# implicit index when slicing
data[1:3]
# Because of this potential confusion in the case of integer indexes, Pandas provides some special *indexer* attributes that explicitly expose certain indexing schemes.
# These are not functional methods, but attributes that expose a particular slicing interface to the data in the ``Series``.
#
# First, the ``loc`` attribute allows indexing and slicing that always references the explicit index:
data.loc[1]
data.loc[1:3]
# The ``iloc`` attribute allows indexing and slicing that always references the implicit Python-style index:
data.iloc[1]
data.iloc[1:3]
# A third indexing attribute, ``ix``, is a hybrid of the two, and for ``Series`` objects is equivalent to standard ``[]``-based indexing.
# The purpose of the ``ix`` indexer will become more apparent in the context of ``DataFrame`` objects, which we will discuss in a moment.
#
# One guiding principle of Python code is that "explicit is better than implicit."
# The explicit nature of ``loc`` and ``iloc`` make them very useful in maintaining clean and readable code; especially in the case of integer indexes, I recommend using these both to make code easier to read and understand, and to prevent subtle bugs due to the mixed indexing/slicing convention.
# ## Data Selection in DataFrame
#
# Recall that a ``DataFrame`` acts in many ways like a two-dimensional or structured array, and in other ways like a dictionary of ``Series`` structures sharing the same index.
# These analogies can be helpful to keep in mind as we explore data selection within this structure.
# ### DataFrame as a dictionary
#
# The first analogy we will consider is the ``DataFrame`` as a dictionary of related ``Series`` objects.
# Let's return to our example of areas and populations of states:
area = pd.Series({'California': 423967, 'Texas': 695662,
'New York': 141297, 'Florida': 170312,
'Illinois': 149995})
pop = pd.Series({'California': 38332521, 'Texas': 26448193,
'New York': 19651127, 'Florida': 19552860,
'Illinois': 12882135})
data = pd.DataFrame({'area':area, 'pop':pop})
data
# The individual ``Series`` that make up the columns of the ``DataFrame`` can be accessed via dictionary-style indexing of the column name:
data['area']
# Equivalently, we can use attribute-style access with column names that are strings:
data.area
# This attribute-style column access actually accesses the exact same object as the dictionary-style access:
data.area is data['area']
# Though this is a useful shorthand, keep in mind that it does not work for all cases!
# For example, if the column names are not strings, or if the column names conflict with methods of the ``DataFrame``, this attribute-style access is not possible.
# For example, the ``DataFrame`` has a ``pop()`` method, so ``data.pop`` will point to this rather than the ``"pop"`` column:
data.pop is data['pop']
# In particular, you should avoid the temptation to try column assignment via attribute (i.e., use ``data['pop'] = z`` rather than ``data.pop = z``).
#
# Like with the ``Series`` objects discussed earlier, this dictionary-style syntax can also be used to modify the object, in this case adding a new column:
data['density'] = data['pop'] / data['area']
data
# This shows a preview of the straightforward syntax of element-by-element arithmetic between ``Series`` objects; we'll dig into this further in [Operating on Data in Pandas](03.03-Operations-in-Pandas.ipynb).
# ### DataFrame as two-dimensional array
#
# As mentioned previously, we can also view the ``DataFrame`` as an enhanced two-dimensional array.
# We can examine the raw underlying data array using the ``values`` attribute:
data.values
# With this picture in mind, many familiar array-like observations can be done on the ``DataFrame`` itself.
# For example, we can transpose the full ``DataFrame`` to swap rows and columns:
data.T
# When it comes to indexing of ``DataFrame`` objects, however, it is clear that the dictionary-style indexing of columns precludes our ability to simply treat it as a NumPy array.
# In particular, passing a single index to an array accesses a row:
data.values[0]
# and passing a single "index" to a ``DataFrame`` accesses a column:
data['area']
# Thus for array-style indexing, we need another convention.
# Here Pandas again uses the ``loc``, ``iloc``, and ``ix`` indexers mentioned earlier.
# Using the ``iloc`` indexer, we can index the underlying array as if it is a simple NumPy array (using the implicit Python-style index), but the ``DataFrame`` index and column labels are maintained in the result:
data.iloc[:3, :2]
# Similarly, using the ``loc`` indexer we can index the underlying data in an array-like style but using the explicit index and column names:
data.loc[:'Illinois', :'pop']
# The ``ix`` indexer allows a hybrid of these two approaches:
data.ix[:3, :'pop']
# Keep in mind that for integer indices, the ``ix`` indexer is subject to the same potential sources of confusion as discussed for integer-indexed ``Series`` objects.
#
# Any of the familiar NumPy-style data access patterns can be used within these indexers.
# For example, in the ``loc`` indexer we can combine masking and fancy indexing as in the following:
data.loc[data.density > 100, ['pop', 'density']]
# Any of these indexing conventions may also be used to set or modify values; this is done in the standard way that you might be accustomed to from working with NumPy:
data.iloc[0, 2] = 90
data
# To build up your fluency in Pandas data manipulation, I suggest spending some time with a simple ``DataFrame`` and exploring the types of indexing, slicing, masking, and fancy indexing that are allowed by these various indexing approaches.
# ### Additional indexing conventions
#
# There are a couple extra indexing conventions that might seem at odds with the preceding discussion, but nevertheless can be very useful in practice.
# First, while *indexing* refers to columns, *slicing* refers to rows:
data['Florida':'Illinois']
# Such slices can also refer to rows by number rather than by index:
data[1:3]
# Similarly, direct masking operations are also interpreted row-wise rather than column-wise:
data[data.density > 100]
# These two conventions are syntactically similar to those on a NumPy array, and while these may not precisely fit the mold of the Pandas conventions, they are nevertheless quite useful in practice.
# # Operating on Data in Pandas
# One of the essential pieces of NumPy is the ability to perform quick element-wise operations, both with basic arithmetic (addition, subtraction, multiplication, etc.) and with more sophisticated operations (trigonometric functions, exponential and logarithmic functions, etc.).
# Pandas inherits much of this functionality from NumPy, and the ufuncs that we introduced in [Computation on NumPy Arrays: Universal Functions](02.03-Computation-on-arrays-ufuncs.ipynb) are key to this.
#
# Pandas includes a couple useful twists, however: for unary operations like negation and trigonometric functions, these ufuncs will *preserve index and column labels* in the output, and for binary operations such as addition and multiplication, Pandas will automatically *align indices* when passing the objects to the ufunc.
# This means that keeping the context of data and combining data from different sources–both potentially error-prone tasks with raw NumPy arrays–become essentially foolproof ones with Pandas.
# We will additionally see that there are well-defined operations between one-dimensional ``Series`` structures and two-dimensional ``DataFrame`` structures.
# ## Ufuncs: Index Preservation
#
# Because Pandas is designed to work with NumPy, any NumPy ufunc will work on Pandas ``Series`` and ``DataFrame`` objects.
# Let's start by defining a simple ``Series`` and ``DataFrame`` on which to demonstrate this:
import pandas as pd
import numpy as np
rng = np.random.RandomState(42)
ser = pd.Series(rng.randint(0, 10, 4))
ser
df = pd.DataFrame(rng.randint(0, 10, (3, 4)),
columns=['A', 'B', 'C', 'D'])
df
# If we apply a NumPy ufunc on either of these objects, the result will be another Pandas object *with the indices preserved:*
np.exp(ser)
# Or, for a slightly more complex calculation:
np.sin(df * np.pi / 4)
# Any of the ufuncs discussed in [Computation on NumPy Arrays: Universal Functions](02.03-Computation-on-arrays-ufuncs.ipynb) can be used in a similar manner.
# ## UFuncs: Index Alignment
#
# For binary operations on two ``Series`` or ``DataFrame`` objects, Pandas will align indices in the process of performing the operation.
# This is very convenient when working with incomplete data, as we'll see in some of the examples that follow.
# ### Index alignment in Series
#
# As an example, suppose we are combining two different data sources, and find only the top three US states by *area* and the top three US states by *population*:
area = pd.Series({'Alaska': 1723337, 'Texas': 695662,
'California': 423967}, name='area')
population = pd.Series({'California': 38332521, 'Texas': 26448193,
'New York': 19651127}, name='population')
# Let's see what happens when we divide these to compute the population density:
population / area
# The resulting array contains the *union* of indices of the two input arrays, which could be determined using standard Python set arithmetic on these indices:
area.index | population.index
# Any item for which one or the other does not have an entry is marked with ``NaN``, or "Not a Number," which is how Pandas marks missing data (see further discussion of missing data in [Handling Missing Data](03.04-Missing-Values.ipynb)).
# This index matching is implemented this way for any of Python's built-in arithmetic expressions; any missing values are filled in with NaN by default:
A = pd.Series([2, 4, 6], index=[0, 1, 2])
B = pd.Series([1, 3, 5], index=[1, 2, 3])
A + B
# If using NaN values is not the desired behavior, the fill value can be modified using appropriate object methods in place of the operators.
# For example, calling ``A.add(B)`` is equivalent to calling ``A + B``, but allows optional explicit specification of the fill value for any elements in ``A`` or ``B`` that might be missing:
A.add(B, fill_value=0)
# ### Index alignment in DataFrame
#
# A similar type of alignment takes place for *both* columns and indices when performing operations on ``DataFrame``s:
A = pd.DataFrame(rng.randint(0, 20, (2, 2)),
columns=list('AB'))
A
B = pd.DataFrame(rng.randint(0, 10, (3, 3)),
columns=list('BAC'))
B
A + B
# Notice that indices are aligned correctly irrespective of their order in the two objects, and indices in the result are sorted.
# As was the case with ``Series``, we can use the associated object's arithmetic method and pass any desired ``fill_value`` to be used in place of missing entries.
# Here we'll fill with the mean of all values in ``A`` (computed by first stacking the rows of ``A``):
fill = A.stack().mean()
A.add(B, fill_value=fill)
# The following table lists Python operators and their equivalent Pandas object methods:
#
# | Python Operator | Pandas Method(s) |
# |-----------------|---------------------------------------|
# | ``+`` | ``add()`` |
# | ``-`` | ``sub()``, ``subtract()`` |
# | ``*`` | ``mul()``, ``multiply()`` |
# | ``/`` | ``truediv()``, ``div()``, ``divide()``|
# | ``//`` | ``floordiv()`` |
# | ``%`` | ``mod()`` |
# | ``**`` | ``pow()`` |
#
# ## Ufuncs: Operations Between DataFrame and Series
#
# When performing operations between a ``DataFrame`` and a ``Series``, the index and column alignment is similarly maintained.
# Operations between a ``DataFrame`` and a ``Series`` are similar to operations between a two-dimensional and one-dimensional NumPy array.
# Consider one common operation, where we find the difference of a two-dimensional array and one of its rows:
A = rng.randint(10, size=(3, 4))
A
A - A[0]
# According to NumPy's broadcasting rules (see [Computation on Arrays: Broadcasting](02.05-Computation-on-arrays-broadcasting.ipynb)), subtraction between a two-dimensional array and one of its rows is applied row-wise.
#
# In Pandas, the convention similarly operates row-wise by default:
df = pd.DataFrame(A, columns=list('QRST'))
df - df.iloc[0]
# If you would instead like to operate column-wise, you can use the object methods mentioned earlier, while specifying the ``axis`` keyword:
df.subtract(df['R'], axis=0)
# Note that these ``DataFrame``/``Series`` operations, like the operations discussed above, will automatically align indices between the two elements:
halfrow = df.iloc[0, ::2]
halfrow
df - halfrow
# This preservation and alignment of indices and columns means that operations on data in Pandas will always maintain the data context, which prevents the types of silly errors that might come up when working with heterogeneous and/or misaligned data in raw NumPy arrays.
| 04-Linear-Regression-with-Python/related-tutorials/01-C-introduction-pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#Libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
#Data
path='C:\\Users\\sagi\\Desktop\\Learning\\ML\\Datasets\\petrol_consumption.csv'
dataset = pd.read_csv(path)
#Explore the data
dataset.shape
dataset.describe()
dataset.head()
#Preprocess the data
X = dataset[['Petrol_tax', 'Average_income', 'Paved_Highways',
'Population_Driver_licence(%)']]
y = dataset['Petrol_Consumption']
#Split
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=0)
#Train
from sklearn.linear_model import LinearRegression
regressor = LinearRegression()
regressor.fit(X_train, y_train)
coeff_df = pd.DataFrame(regressor.coef_, X.columns, columns=['Coefficient'])
print(coeff_df)
# +
#Making prediction
y_pred = regressor.predict(X_test)
df = pd.DataFrame({'Actual': y_test, 'Predicted': y_pred})
print(df)
# +
#Evaluation
from sklearn import metrics
print('Mean Absolute Error:', metrics.mean_absolute_error(y_test, y_pred))
print('Mean Squared Error:', metrics.mean_squared_error(y_test, y_pred))
print('Root Mean Squared Error:', np.sqrt(metrics.mean_squared_error(y_test, y_pred)))
#RMSE is slightly greater than 10% of the mean value of the gas consumption in all states
#So the model is not very accurate, but it can still be useful to make predictions
| Multiple-Linear-Regression-petrol_consumption.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %gui qt
import os
import sys
sys.path.insert(0, '/Volumes/group/awagner/sgagnon/scripts/lyman-tools/viz/')
from plot_surface import *
from nilearn import plotting as nip
# %gui qt
# %matplotlib inline
# %gui qt
mni_template = '/Volumes/group/awagner/sgagnon/AP/data/avg152T1_brain.nii.gz'
def plot_snapviews(snapshots, snap_views, hemi='lh'):
n_panels = len(snap_views)
f, axes = plt.subplots(1,n_panels, figsize=(5*n_panels,10))
for ax, view in zip(axes.flat, snap_views):
ax.set_axis_off()
ax.imshow(snapshots[view], rasterized=True)
f.savefig('/Volumes/group/awagner/sgagnon/AP/results/control-stress_safe_sourcehit-cr_{hemi}_p005.png'.format(hemi=hemi),
dpi=600)
print 'running'
# ## Plot frontoparietal networks
# +
hemi = 'lh'
subid='fsaverage'
brain = Brain(subid, hemi, "semi7", background='white', size=(800,800), cortex=("gray", -2, 7, True))
brain.add_label('frontoparietal', hemi=hemi, color=(204./255, 121./255, 167./255), alpha = 0.7)
brain.add_label('frontoparietal', hemi=hemi, color=(204./255, 121./255, 167./255), borders=True)
brain.add_label('dorsalattn', hemi=hemi, color=(0./255, 158./255, 115./255), alpha = 0.7)
brain.add_label('dorsalattn', hemi=hemi, color=(0./255, 158./255, 115./255), borders=True)
brain.save_imageset('/Users/steph-backup/Dropbox/Stanford/Papers/Dissertation/Figures_defense/AP_frontoparietal_{hemi}_colorblind'.format(hemi=hemi),
['lat'], filetype='tiff')
# +
hemi = 'lh'
subid='fsaverage'
brain = Brain(subid, hemi, "semi7", background='white', size=(800,800), cortex='low_contrast')
brain.add_label('frontoparietal', hemi=hemi, color=(112./255, 201./255, 231./255), alpha = 0.7)
brain.add_label('frontoparietal', hemi=hemi, color=(112./255, 201./255, 231./255), borders=True)
brain.add_label('dorsalattn', hemi=hemi, color=(202./255, 152./255, 235./255), alpha = 0.7)
brain.add_label('dorsalattn', hemi=hemi, color=(202./255, 152./255, 235./255), borders=True)
brain.save_imageset('/Users/steph-backup/Dropbox/Stanford/Papers/Dissertation/Figures_defense/AP_frontoparietal_{hemi}'.format(hemi=hemi),
['lat', 'fro'], filetype='tiff')
# -
# ## localizer coefficients on example brain
# ### Just mask:
# +
subid = 'ap164'
place_coefs = '/Volumes/group/awagner/sgagnon/AP/analysis/mvpa_raw/'+subid+'/importance_maps/bilat-parahipp_fusi_inftemp_nohipp_coef_place.nii.gz'
mask = '/Volumes/group/awagner/sgagnon/AP/data/'+subid+'/masks/bilat-parahipp_fusi_inftemp_nohipp.nii.gz'
for hemi in ['lh', 'rh']:
brain = Brain(subid, hemi, "inflated", background='white', size=(850,800))
maskmap = project_volume_data(mask, hemi, subject_id=subid, smooth_fwhm=0).astype(bool)
maskmap = ~maskmap
brain.add_data(maskmap, min=0, max=10, thresh=.5, hemi=hemi,
colormap="bone", alpha=.6, colorbar=False)
brain.show_view("ven")
brain.save_imageset('/Users/steph-backup/Dropbox/Stanford/Papers/Dissertation/Figures/AP_VTCmask_{hemi}_ap164'.format(hemi=hemi),
['ven'], filetype='tiff')
brain.save_imageset('/Volumes/group/awagner/sgagnon/AP/results/AP_VTCmask_{hemi}_ap164'.format(hemi=hemi),
['ven'], filetype='tiff')
# -
brain.show_vi
brain.show_view({'distance': 600}, roll=0)
brain.show_view({'azimuth': -177, 'elevation': 88}, roll=91)
brain.save_image('/Volumes/group/awagner/sgagnon/AP/results/AP_VTCmask_ap164_')
brain.show_view()
# +
subid = 'ap164'
place_coefs = '/Volumes/group/awagner/sgagnon/AP/analysis/mvpa_raw/'+subid+'/importance_maps/bilat-parahipp_fusi_inftemp_nohipp_coef_place.nii.gz'
mask = '/Volumes/group/awagner/sgagnon/AP/data/'+subid+'/masks/bilat-parahipp_fusi_inftemp_nohipp.nii.gz'
for hemi in ['lh', 'rh']:
brain = Brain(subid, hemi, "inflated", background='white', size=(850,800))
coefmap = project_volume_data(place_coefs, hemi, subject_id=subid, smooth_fwhm=0)
maskmap = project_volume_data(mask, hemi, subject_id=subid, smooth_fwhm=0).astype(bool)
brain.add_data(coefmap, min=-0.09, max=0.09, colormap="PiYG", hemi=hemi, alpha=.8, colorbar=True)
maskmap = ~maskmap
brain.add_data(maskmap, min=0, max=10, thresh=.5, hemi=hemi,
colormap="bone", alpha=.6, colorbar=False)
brain.show_view("ven")
brain.save_imageset('/Users/steph-backup/Dropbox/Stanford/Papers/Dissertation/Figures/AP_localizercoef_green_cmap_{hemi}_ap164'.format(hemi=hemi),
['ven'], filetype='tiff')
# +
subid = 'ap164'
place_coefs = '/Volumes/group/awagner/sgagnon/AP/analysis/mvpa_raw/'+subid+'/importance_maps/bilat-parahipp_fusi_inftemp_nohipp_coef_place.nii.gz'
mask = '/Volumes/group/awagner/sgagnon/AP/data/'+subid+'/masks/bilat-parahipp_fusi_inftemp_nohipp.nii.gz'
for hemi in ['lh', 'rh']:
brain = Brain(subid, hemi, "inflated", background='white', size=(850,800))
coefmap = project_volume_data(place_coefs, hemi, subject_id=subid, smooth_fwhm=0)
maskmap = project_volume_data(mask, hemi, subject_id=subid, smooth_fwhm=0).astype(bool)
brain.add_data(coefmap, min = 0, thresh=0.000000000000000000000000001, colormap="Greens", hemi=hemi, alpha=.8, colorbar=False)
brain.add_data(-coefmap, min = 0, thresh=0.000000000000000000000000001, colormap="PuRd", hemi=hemi, alpha=.8, colorbar=False)
maskmap = ~maskmap
brain.add_data(maskmap, min=0, max=10, thresh=.5, hemi=hemi,
colormap="bone", alpha=.6, colorbar=False)
brain.show_view("ven")
brain.save_imageset('/Users/steph-backup/Dropbox/Stanford/Papers/Dissertation/Figures/AP_localizercoef_{hemi}_ap164'.format(hemi=hemi),
['ven'], filetype='tiff')
# -
# ## Sphere from SH > CR by group:
# +
z_thresh = .5
stat_map = op.join('/Volumes/group/awagner/sgagnon/AP/analysis/ap_memory_raw/group_control-stress/mni/sourcehit-cr/zstat1_peak1_5mm_sphere_masked.nii.gz')
fig = plt.figure(dpi=600, figsize=(10,5))
display = nip.plot_stat_map(stat_map, bg_img=mni_template,
threshold=z_thresh,
black_bg=False,
cut_coords=[22,-36, -2], draw_cross=False,
cmap='PuBu', colorbar=False, annotate=False, figure=fig)
display.annotate(size=20)
fig.savefig('/Users/steph-backup/Dropbox/Stanford/Papers/Dissertation/Figures/DPI150_jpg/AP_sphere_sh-cr_ctrl-str.jpeg', dpi=150)
# -
# ## Assoc hit > CR
# #### Group interaction (p < 0.05, corrected):
# +
exp = 'ap_memory_raw'
group = 'group_control-stress'
contrast = 'sourcehit-cr'
z_thresh = 2.3
stat_map = op.join('/Volumes/group/awagner/sgagnon/AP/analysis',
exp, group, 'mni', contrast, 'zstat1_threshold.nii.gz')
fig = plt.figure(dpi=600, figsize=(10,5))
display = nip.plot_stat_map(stat_map, bg_img=mni_template,
threshold=z_thresh, display_mode='yx',
black_bg=False,
cut_coords=[28, -34],
cmap='PuBu', colorbar=True, annotate=True, figure=fig)
display.annotate(size=20)
fig.savefig('/Volumes/group/awagner/sgagnon/AP/results/ap_memory_raw_control-stress_sourcehit-cr.png')
# +
exp = 'ap_memory_raw'
group = 'group_control-stress'
contrast = 'sourcehit-cr'
z_thresh = 2.3
stat_map = op.join('/Volumes/group/awagner/sgagnon/AP/analysis',
exp, group, 'mni', contrast, 'zstat1_threshold.nii.gz')
fig = plt.figure(dpi=600, figsize=(10,5))
display = nip.plot_stat_map(stat_map, bg_img=mni_template,
threshold=z_thresh, display_mode='yx',
black_bg=False,
cut_coords=[28, -34], draw_cross=False,
cmap='PuBu', colorbar=True, annotate=False, figure=fig)
fig.savefig('/Volumes/group/awagner/sgagnon/AP/results/ap_memory_raw_control-stress_sourcehit-cr_nocrossbar.png')
# +
exp = 'ap_memory_raw'
group = 'group_control-stress'
contrast = 'sourcehit-cr'
z_thresh = 2.58
stat_map = op.join('/Volumes/group/awagner/sgagnon/AP/analysis',
exp, group, 'mni', contrast, 'zstat1.nii.gz')
fig = plt.figure(dpi=600, figsize=(10,5))
display= nip.plot_stat_map(stat_map, bg_img=mni_template,
threshold=z_thresh, display_mode='yz',
black_bg=False,
cut_coords=[52, 20],
cmap='PuBu', colorbar=True, annotate=True, figure=fig)
display.annotate(size=20)
fig.savefig('/Volumes/group/awagner/sgagnon/AP/results/ap_memory_raw_control-stress_sourcehit-cr_fpc_p005uncorr.tiff')
# -
# ### Conjunction
# +
subj = 'fsaverage'
hemis = ['lh', 'rh']
exp = ['ap_memory_raw']
z_thresh = 2.3 # p < 0.01
contrast = 'sourcehit-CR'
colors = ['Blues_r', 'Oranges_r']
for hemi in hemis:
plot_groups(subj, hemi, exp, contrast, colors, z_thresh,
'/Volumes/group/awagner/sgagnon/AP/results/controlstress_conj_sourcehit-CR/map_nocontour_{hemi}_p05corr.png'.format(hemi=hemi),
save_views =['lat', 'med', 'fro'],
alpha=.8,
save_file=True, colorbar=True,
base_exp='/Volumes/group/awagner/sgagnon/AP',
sig_to_z=False, regspace='mni',
corrected=True,
plot_conjunction=True,
add_border=False)
# +
# add contour for interaction
subj = 'fsaverage'
hemis = ['lh', 'rh']
exp = ['ap_memory_raw']
z_thresh = 2.3 # p < 0.01
contrast = 'sourcehit-CR'
colors = ['Blues_r', 'Oranges_r']
for hemi in hemis:
plot_groups(subj, hemi, exp, contrast, colors, z_thresh,
'/Volumes/group/awagner/sgagnon/AP/results/controlstress_conj_sourcehit-CR/map_{hemi}_p05corr.png'.format(hemi=hemi),
save_views =['lat', 'med', 'fro'],
alpha=.8,
save_file=True,
base_exp='/Volumes/group/awagner/sgagnon/AP',
sig_to_z=False, regspace='mni',
corrected=True,
plot_conjunction=True,
add_border='/Volumes/group/awagner/sgagnon/AP/analysis/ap_memory_raw/group_control-stress/mni/sourcehit-cr/zstat1_threshold.nii.gz',
border_max=True)
# +
subj = 'fsaverage'
hemis = ['lh', 'rh']
exp = ['ap_memory_raw']
z_thresh = 2.3 # p < 0.01
contrasts = ['sourcehit-CR']
colors = ['Reds_r']
alpha = 1
snap_views = ['lat', 'med']
plot_conjunction=False
for hemi in hemis:
b, snapshots = plot_contrasts(subj, hemi, exp, contrasts, colors, z_thresh, '',
base_exp='/Volumes/group/awagner/sgagnon/AP',
group='group_control',
snap_views=snap_views, sig_to_z=False, regspace='mni',
corrected=True,
save_file=False, plot_conjunction=plot_conjunction)
plot_snapviews(snapshots, snap_views, hemi=hemi)
# -
# ### Uncorrected p < 0.005; control > stress (assoc hit > CR)
# +
subj = 'fsaverage'
hemis = ['lh', 'rh']
exp = ['ap_memory_raw']
z_thresh = 2.58 # p < 0.005
contrasts = ['sourcehit-CR']
colors = ['Blues_r']
for hemi in hemis:
plot_contrasts(subj, hemi, exp, contrasts, colors, z_thresh,
'/Volumes/group/awagner/sgagnon/AP/results/control-stress_sourcehit-CR/map_{hemi}_p005uncorr.png'.format(hemi=hemi),
save_views =['lat', 'med', 'fro'],
base_exp='/Volumes/group/awagner/sgagnon/AP',
alpha = 0.8,
group='group_control-stress',
colorbar=True,
sig_to_z=False, regspace='mni',
corrected=False)
# +
# Visualize overlaid with Yeo CCN network (maroon / left lateral IPS)
b, snapshots = plot_contrasts(subj, 'lh', exp, contrasts, colors, z_thresh, '',
base_exp='/Volumes/group/awagner/sgagnon/AP',
group='group_control-stress',
snap_views=snap_views, sig_to_z=False, regspace='mni',
corrected=False,
save_file=False, plot_conjunction=plot_conjunction)
b.add_label('17Networks_13', borders=True, color='maroon')
# -
# ### Uncorrected p < 0.005; safe runs, control > stress (assoc hit > CR)
# +
subj = 'fsaverage'
hemis = ['lh', 'rh']
exp = ['ap_memory_raw-byshockCond']
z_thresh = 2.5758 # p < 0.005
contrasts = ['safe_sourcehit-CR']
colors = ['Reds_r']
alpha = 1
snap_views = ['fro', 'lat']
plot_conjunction=False
for hemi in hemis:
b, snapshots = plot_contrasts(subj, hemi, exp, contrasts, colors, z_thresh, '',
base_exp='/Volumes/group/awagner/sgagnon/AP',
group='group_control-stress',
snap_views=snap_views, sig_to_z=False, regspace='mni',
corrected=False,
save_file=False, plot_conjunction=plot_conjunction)
plot_snapviews(snapshots, snap_views, hemi=hemi)
# -
# # By source accuracy
# +
subj = 'fsaverage'
hemis = ['lh', 'rh']
exp = ['ap_memory_raw']
z_thresh = 2.3 # p < 0.01
contrasts = ['sourcehit-cr']
colors = ['PuBu_r']
alpha = 1
save_views = ['lat', 'med', 'fro']
plot_conjunction=False
# os.mkdir('/Volumes/group/awagner/sgagnon/AP/results/cov_sourceAcc_sh-cr')
for hemi in hemis:
plot_contrasts(subj, hemi, exp, contrasts, colors, z_thresh,
'/Volumes/group/awagner/sgagnon/AP/results/cov_sourceAcc_sh-cr/map_{hemi}_p05corr.png'.format(hemi=hemi),
base_exp='/Volumes/group/awagner/sgagnon/AP',
group='group_cov_sourceAcc',
sig_to_z=False,
alpha=.9,
regspace='mni',
save_views=save_views,
skip_reg_dir=False,
corrected=True, colorbar=True,
save_file=True, plot_conjunction=plot_conjunction)
# +
exp = 'ap_memory_raw'
group = 'group_cov_sourceAcc'
contrast = 'sourcehit-cr'
z_thresh = 2.3
stat_map = op.join('/Volumes/group/awagner/sgagnon/AP/analysis',
exp, group, 'mni', contrast, 'zstat1_threshold.nii.gz')
fig = plt.figure(dpi=600, figsize=(15,5))
display = nip.plot_stat_map(stat_map, bg_img=mni_template,
threshold=z_thresh, display_mode='ortho',
black_bg=False,
cut_coords=[24, -36, -8],
cmap='PuBu', colorbar=True, annotate=True, figure=fig)
display.annotate(size=20)
fig.savefig('/Volumes/group/awagner/sgagnon/AP/results/ap_memory_raw_covSourceAcc_sourcehit-cr.png')
# -
# # Plot searchlight localizer
# +
subj = 'fsaverage'
mask_file = '/Volumes/group/awagner/sgagnon/AP/analysis/ap_memory_raw/group_control-stress/mni/sourcehit-cr/{hemi}.group_mask.mgz'
stat_temp = '/Volumes/group/awagner/sgagnon/AP/analysis/mvpa_raw/searchlight/localizer_acc_t_tstat1_mask_p05corr.nii.gz'
sign = 'pos'
save_name = '/Volumes/group/awagner/sgagnon/AP/results/searchlight_localizer/{hemi}_localizer'
# os.mkdir('/Volumes/group/awagner/sgagnon/AP/results/searchlight_localizer')
z_thresh = 1.7
for hemi in ['lh', 'rh']:
b = Brain(subj, hemi, 'semi7', background="white")
add_mask_overlay(b, mask_file.format(hemi=hemi))
z_max = calculate_sat_point(stat_temp, '', sign, subj=subj, sig_to_z=False)
add_stat_overlay(b, stat_temp, z_thresh, z_max, sign,
hemi=hemi, sig_to_z=False, color='Greens_r', alpha=.9,
output=True, colorbar=True)
b.save_imageset(save_name.format(hemi=hemi), ['lat', 'fro', 'med', 'ven'])
# -
# # Plot searchlight reinstatement
# +
subj = 'fsaverage'
mask_file = '/Volumes/group/awagner/sgagnon/AP/analysis/ap_memory_raw/group_control-stress/mni/sourcehit-cr/{hemi}.group_mask.mgz'
stat_temp = '/Volumes/group/awagner/sgagnon/AP/analysis/mvpa_raw/searchlight_test/sourcehit_time{time}_acc_t_tstat1_mask_p05corr.nii.gz'
sign = 'pos'
save_name = '/Volumes/group/awagner/sgagnon/AP/results/searchlight_test/{hemi}_test_{time}'
# os.mkdir('/Volumes/group/awagner/sgagnon/AP/results/searchlight_test')
z_thresh = 1.7
for time in [0,2,4,6,8,10,12]:
for hemi in ['lh', 'rh']:
b = Brain(subj, hemi, 'semi7', background="white")
add_mask_overlay(b, mask_file.format(hemi=hemi))
add_stat_overlay(b, stat_temp.format(time=str(time)), z_thresh, 10, sign,
hemi=hemi, sig_to_z=False, color='Greens_r', alpha=.9,
output=True, colorbar=True)
b.save_imageset(save_name.format(hemi=hemi, time=str(time)), ['lat', 'fro', 'med', 'ven'])
# -
| SNI/scripts/notebooks/AP Manuscript Figures.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="WxppwDLKB1gj"
# 
# <center><b>Complejidad Algorítmica – CC41</b></center> <br>
# <center><b>Trabajo Parcial</b></center> <br>
# <center> Carrera de Ingeniería de Software y Ciencias de la computación</center>
# <br>
#
#
#
# <center>u201711783 - <NAME></center>
# <center>u201914955 - <NAME></center>
# <center>u201917621 - <NAME></center>
# + [markdown] id="FOSydxfmdW3l"
# # Link de la documentación del código
# https://colab.research.google.com/drive/17izFcQetKJ6KNvODLTDSKguBx0DBjsg8?usp=sharing
#
# #Link de github
# https://github.com/lucas1619/tf-complejidad
# + [markdown] id="oth65QPxudan"
# # Introducción
#
# El encontrar un camino más corto para llegar de un lugar a otro es algo muy importante para las empresas de delivery ahora, mientras más rápido llegas, más clientes obtienes; asimismo, en los juegos como los laberintos, el encontrar el camino hacia el final es muy importante por si estás jugando con otros compañeros y el que encuentra el camino gana, nuestro presente trabajo se trata del juego Quoridor, un juego que tiene como objetivo llegar primero hasta la base del rival. Además, en cada turno se debe escoger entre colocar dificultades como muros o avanzar. Dicho juego lo vamos a tratar como un problema de "Path Finding" donde se busca encontrar la ruta más corta entre dos nodos de un grafo con algoritmos eficientes considerando el tiempo de ejecución, el espacio en disco y la complejidad del mismo.
# + [markdown] id="F6Cov2sYuLka"
# # <NAME>
#
# ## Complejidad Algorítmica
#
# La complejidad algorítmica es una métrica que nos ayuda a describir el comportamiento de un algoritmo en términos de tiempo de ejecución y la memoria que requiere. Un problema puede tener distintas soluciones, pero más importante que hallar una solución es que esta sea viable y eficaz, el medir la complejidad de los algoritmos nos ayuda a saber cuál es el que debemos implementar.
# Al tiempo de ejecución se le llama complejidad temporal y a la memoria requerida para solucionar un problema se le llama complejidad espacial, ambos dependen del tamaño del problema. La complejidad algorítmica no depende siempre del tiempo de ejecución, la cantidad de memoria consumida, el sistema en donde se corra el algoritmo y el estilo de programación implementado, sino del número de instrucciones necesarias para resolver el problema. Sin embargo, esto no siempre es así, los algoritmos de búsqueda y ordenamiento son una prueba de ello pues estos iteran hasta llegar a la solución deseada, estos algoritmos presentan el mejor y peor caso, por ejemplo: el mejor caso para el algoritmo de búsqueda sería cuando el número a buscar sea el primero del arreglo y el peor cuando este sea el último del arreglo. En la notación, se suele emplear la **contracción**, por ejemplo un algoritmo de complejidad Θ(3n² + 5n + 9) se reduce a Θ(n²), a continuación una tabla que muestra los órdenes de complejidad más comunes:
#
# 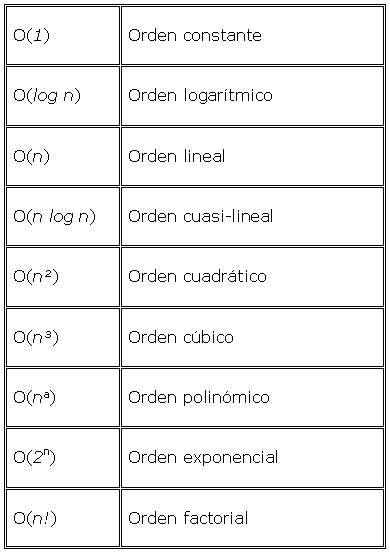
#
#
# ## Pathfinding
# Se le denomina de esta manera al área de inteligencia artificial que tiene como objetivo encontrar el mejor camino de un
# punto a otro en mapas representados digitalmente . Estos algoritmos tienen que definir por dónde dirigirse, basados en la información
# que se tiene del ambiente. Existen diversos tipos de algoritmos como A*, ACO, JPS, etc que buscan reducir el tiempo de búsqueda y los
# recursos utilizados, con el objetivo de hacer más eficiente la tarea. En este proyecto usaremos algunos de ellos y compararemos su eficiencia.
#
# 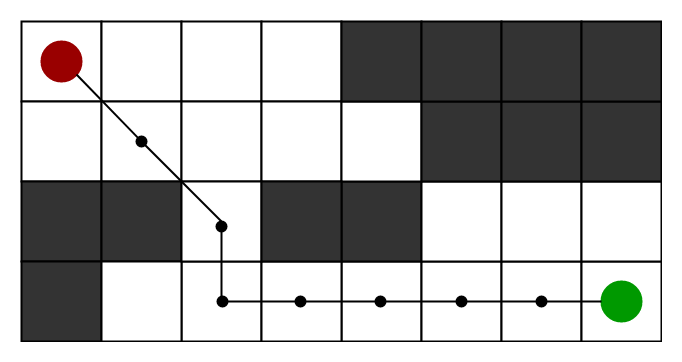
# + [markdown] id="GkWghGf1ul-Y"
# # Estado del arte
#
# ## Algoritmos a utilizar
#
# - Algoritmo A Star
# - Algoritmo Dijkstra
# - Bellman Ford
#
# ### Algoritmo de Dijkstra
#
# El algoritmo de Dijkstra (también llamado Búsqueda de costo uniforme) nos permite priorizar qué caminos explorar. En lugar de explorar todas las rutas posibles por igual, favorece las rutas de menor costo. Por lo que se le pueden asignar rutas a los vertices. Por ello la idea del algoritmo consiste en ir explorando todos los caminos más cortos que parten del vértice origen y que llevan a todos los demás vértices; cuando se obtiene el camino más corto desde el vértice origen hasta el resto de los vértices que componen el grafo, el algoritmo se detiene.
#
# #### Pseudocodigo
#
# 
#
# #### **Complejidad del algoritmo**
#
# El algoritmo consiste en n-1 iteraciones, como máximo. En cada iteración, se añade un vértice al conjunto distinguido.
# En cada iteración, se identifica el vértice con la menor etiqueta entre los que no están en Sk. El número de estas operaciones está acotado por n-1.
# Además, se realizan una suma y una comparación para actualizar la etiqueta de cada uno de los vértices que no están en Sk.
# Luego, en cada iteración se realizan a lo sumo 2(n-1) operaciones.
#
# Entonces:
#
# Teorema: El algoritmo de Dijkstra realiza O(n2) operaciones (sumas y comparaciones) para determinar la longitud del camino más corto entre dos vértices de un grafo ponderado simple, conexo y no dirigido con n vértices.
#
# En conclusión
#
# Tiempo de ejecución = O(|A|.𝑻_𝒅𝒌+|v|.𝑻_𝒅𝒎)
# donde:
# |A|: Número de aristas
# 𝑻_𝒅𝒌: Complejidad de disminuir clave
# |V|: Número de vértices
# 𝑻_𝒅𝒎: Complejidad de extraer mínimo
#
#
#
#
#
# ### Algoritmo Bellman Ford
#
# El algoritmo de Bellman-Ford es un algoritmo que calcula las rutas más cortas desde un único vértice de origen a todos los demás vértices en un dígrafo ponderado de igual manera trabaja con pesos. Su funcionamiento consiste en partir de un vértice origen que será ingresado luego Bellman-Ford relaja todas las aristas y lo hace |V| – 1 veces, siendo |V| el número de vértices del grafo. Las repeticiones permiten a las distancias mínimas recorrer el árbol, ya que en la ausencia de ciclos negativos, el camino más corto solo visita cada vértice una vez. A diferencia de la solución voraz, la cual depende de la suposición de que los pesos sean positivos, esta solución se aproxima más al caso general.
#
# #### Pseudocodigo
#
# 
#
#
# **Complejidad del algoritmo**
#
# La idea fuerte del algoritmo es que si el grafo tiene V nodos, alcanza con recorrer las aristas V−1 veces. Se ve entonces que la complejidad del algoritmo es **O(V.E)** donde V es la cantidad de vértices y E la de aristas.
#
#
# ### Algoritmo A Star
# Es una de las mejores técnicas para búsqueda de rutas y recorridos de grafos. A diferencia de otros, este es un algoritmo inteligente, muchos juegos utilizan este algoritmo para encontrar el camino más corto de forma __eficiente__ , cualidad que es altamente importante para este trabajo. Usaremos este algoritmo para poder encontrar el camino más corto a la meta y lo modificaremos de acuerdo a las condiciones que considera el juego **Quoridor**, de acuerdo al juego, el tablero se representará por un grafo.
#
#
# **A continuación la explicación de este algoritmo:**
#
# Tenemos un grafo con múltiples nodos, y queremos llegar a un nodo específico desde el primer nodo lo más rápido posible.
#
# Lo que hace el Algoritmo A* es que en cada paso elige un nodo de acuerdo a un valor __f__ que es la suma de __g__ y __h__, en cada paso elige el nodo con el menor valor de __f__ y continua el camino:
#
# 
#
# * n : nodo previo en el camino
#
# * g(n) : costo del camino del nodo del inicio hasta n
#
# * h(n) : una heurística que estima el costo del camino más corto desde n hasta el nodo deseado
#
# Consideremos que el grafo es este:
#
# 
#
# Los números escritos en azul en las aristas representan la distancia entre los nodos y los números escritos con rojo representan el valor de la heurística
#
# El algoritmo A* usa está formula  para encontrar el camino más corto.
#
#
#
# **Ejemplo: Queremos encontrar el camino más corto entre A y J**
#
#
#
# **Nodo de inicio es A**
#
# A se relaciona con B y F
#
# Calcularemos el F(B) y el F(F)
#
# F(B) = 6 + 8 = 14
#
# F(F) = 3 + 6 = 9
#
# El menor es F(F), F será nuestro nuevo nodo de inicio
#
#
#
#
# **Nodo de inicio es F**
#
# F se relaciona con G y H
#
# Calcularemos el F(G) y el F(H)
#
# F(G) = 4 + 5 = 9 ----> 4 = 3 + 1 ---> COSTO DEL CAMINO HASTA AHORA
#
# F(H) = 10 + 3 = 13 ----> 10 = 3 + 7 ---> COSTO DEL CAMINO HASTA AHORA
#
# El menor es F(G), G será nuestro nuevo nodo de inicio
#
#
#
#
# **Nodo de inicio es G**
#
# G solo se relaciona con I
#
# Calcularemos el F(I)
#
# F(I) = 7 + 1 = 8 ----> 7 = 3 + 1 + 3 ---> COSTO DEL CAMINO HASTA AHORA
#
# I será el nuevo nodo de inicio
#
#
#
#
# **Nodo de inicio es I**
#
# I se relaciona con E, H y J
#
# Calcularemos el F(E), el F(H) y el F(J)
#
# F(E) = 12 + 3 = 15 ----> 12 = 3 + 1 + 3 + 5 ---> COSTO DEL CAMINO HASTA AHORA
#
# F(H) = 9 + 3 = 12 ----> 10 = 3 + 1 + 3 + 2 ---> COSTO DEL CAMINO HASTA AHORA
#
# F(J) = 10 + 0 = 10 ----> 10 = 3 + 1 + 3 + 3 ---> COSTO DEL CAMINO HASTA AHORA
#
# El menor es F(J), J será nuestro nuevo nodo de inicio
#
#
#
#
# **Como queriamos llegar al nodo J paramos ahí.**
#
# Así quedaría el recorrido:
#
# 
# El camino fue **A F G I J**
#
# ##Pseudocódigo
#
# 
#
#
# **Complejidad del algoritmo:**
#
# Puede llevarnos viajar por todo el borde del grafo desde el nodo origen hasta el nodo que deseemos. Entonces, en el peor de los casos su complejidad es
#
# 
#
# E: Número de aristas en el gráfico
#
#
# ## Espacio de búsqueda de cada algoritmo:
#
#
# ### <NAME>
#
# El espacio de busqueda de <NAME> debido a su naturaleza voraz encuentra el camino en un tiempo menor a comparación de los otros. Debido a que al seleccionar vorazmente el nodo de peso mínimo aun sin procesar simplemente relaja todas las aristas, y lo hace |V|-1 veces, siendo |V| el número de vértices en el grafo.
#
# ### Algoritmo Dijkstra
#
# El espacio de búsqueda de este algoritmo funciona bien para encontrar el camino más corto, pero pierde tiempo explorando en direcciones que no son prometedoras.
# Debido a que trabaja por etapas, y toma en cada etapa la mejor solución sin considerar consecuencias futuras. Sin embargo el óptimo encontrado en una etapa puede modificarse posteriormente si surge una solución mejor.
#
# 
# ### Algoritmo A Star
#
# El espacio de búsqueda de este algoritmo es menor que el de Dijkstra, pues mientras Dijkstra considera caminos que no son prometedores, A Star al usar una cola de prioridad, ordena a los nodos de acuerdo al menor valor obtenido de la heurística, la cual es la distancia estimada del nodo en el que nos encontramos hasta el nodo al que queremos llegar. De tal manera que obtenemos y exploramos el nodo más cercano al nodo final. El valor obtenido de la heuristica es sumado a la distancia entre nodo concurrente y el nodo, lo cual nos permite obtener el camino más corto y más rápido.
#
# Este sería gráficamente el espacio de búsqueda que ocupa el algoritmo A Star en un tablero 15 x 15 :
#
# 
#
#
#
#
#
#
#
#
# + [markdown] id="tiGFqHVguuYY"
# # Metodología
# La metodología que usamos para resolver este problema se divide en tres partes:
#
# 1. Investigación
#
# 2. Desarrollo
#
# 3. Tests y Experimentos
#
# ### 1. Investigación
# Para dar inicio a la investigación, primero indagamos acerca del problema, debido a ello pudimos observar que debíamos hacer uso de algoritmos relacionados a la busqueda del camino más corto. Usaremos estos algoritmos para el movimiento de los peones a través de los obstáculos o ,en otras palabras, Pathfinding. Asimismo, buscamos algoritmos que podrían ser útiles para nuestro trabajo, los que encontramos fueron A*, Dijkstra y Bellman Ford. Además, se hizo uso de la libreria pygames, para la parte gráfica del juego (representación del tablero y peones).
#
#
# ### 2. Desarrollo
#
# En primer lugar, implementamos una lista doblemente enlazada, lo siguiente fue implementar un grafo, el cual era la combinación de un vector de listas doblemente enlazadas y dentro de ello, una función que permitía que las conexiones entre nodos se almacenen como listas de adyacencia. Elegimos implementarlo de esta manera, debido a que deseamos un acceso constante a los nodos, eso lo obtenemos del vector de listas doblemente enlazadas y además, un almacenado dinámico a sus adyacentes y la lista doblemente enlazada permite ello. De tal manera, que las operaciones de acceso e insertado en nuestro juego sean de O(1)
#
# En segundo lugar, nuestro objetivo fue implementar los algoritmos de path-finding. Con el proposito de avanzar el trabajo de manera más rápida, al ser un grupo de 3 integrantes, cada integrante implemento uno de los algoritmos de path-finding de acuerdo al grafo construido previamente. Al testear que los algoritmos funcionaban y encuentraban las rutas adecuadas pasamos al siguiente paso.
#
# En tercer lugar, separamos el código de manera estructurada y ordenada haciendo uso del paradigma de la Programación Orientada a Objetos, debido a que cada uno trabajo por su cuenta, el código estaba desordenado y decidimos ordenarlo en clases, las clases que identificamos fueron:
#
# - Clase Lista Doblemente Enlazada (clase que contiene la implementación de la lista)
# - Clase Tablero (clase que contiene la implementación del grafo entre otras funcionalidades del mismo)
# - Clase Jugador (clase que contiene todas las funcionalidades del peon del juego)
# - Clase Pensamiento (clase que contiene los algoritmos que seguirán los peones)
# - Clase Quoridor (clase que contiene todo el juego)
#
# En cuarto lugar, implementamos el pensamiento del jugador, ejemplo: cuando saltar, cuando cambiar de ruta, cuando retroceder, cuando seguir si ve que todo está tranquilo, y también implementamos la función mover, la cual después de pensar moverá el circulo arriba, abajo, a la izquierda o derecha.
#
# En quinto lugar, implementamos la parte gráfica con la ayuda de la librería pygames, definiendo a cada nodo como un cuadrado del tablero y al peón como un circulo dentro de este.
#
# En último lugar, testeamos el juego de distintas maneras, lo cual nos confirmo que los algoritmos funcionaban de manera correcta y nos detallo el tiempo que demoraba cada uno de acuerdo al tablero dado. Además, los algoritmos que eran más eficientes para el caso. Debido a que no contamos con paredes, el desenvolvimiento de los algoritmos fueron muy similares, pues no hay barreras y tienen el camino libre, además de una distancia de 1 entre cada nodo. En la entrega final se podrá apreciar de mejor manera el desenvolvimiento de los tres algoritmos implementados en este trabajo.
#
# ### 3. Tests y Experimentos
#
# La forma en la cual haremos los test es dandole como entrada el tamaño del tablero y midiendo el tiempo de ejecucion.
#
# Para esta tarea, primero necesitamos una manera en la cual medir el tiempo de cada juego, por ello se diseño el siguiente método dentro de la clase Quoridor
#
#
#
# + id="rFQmD4iihKk3"
def test(self, alg):
print("Comenzo")
start_time = time.time()
self.ganador = False
self.turno = 0
for i in range(len(self.lista_de_jugadores)):
self.lista_de_jugadores[i].pensamiento.indica = alg
while self.ganador == False:
self.lista_de_jugadores[self.turno].piensa(
self.lista_de_jugadores[(self.turno + 1) % len(self.lista_de_jugadores)], self.tablero)
self.ganador = self.lista_de_jugadores[self.turno].mueve(
self.lista_de_jugadores[(self.turno + 1) % len(self.lista_de_jugadores)],
self.tablero)
self.turno = (self.turno + 1) % len(self.lista_de_jugadores)
elapsed_time = time.time() - start_time
return elapsed_time*1000.0
# + [markdown] id="O29RHG6AhV8z"
# Despues necesitabamos una forma en la cual recopilar el tiempo que nos retorna el método test, para ello se diseño el siguiente algoritmo:
# + id="kZ2MfsJ1noV2"
from Quoridor import Quoridor
import matplotlib.pyplot as plt
def main():
algoritmos = [1,2,3]
titulo = ["B<NAME>", "Dijikstra", "A Star"]
tiempos = [] #guarda los tiempos
tam = [] #guarda los tamaños
for i,algoritmo in enumerate(algoritmos):
size = 5
while(size <= 5000):
game = Quoridor(size, 2)
tiempos.append(game.test(algoritmo))
tam.append(size)
if(tiempos[len(tiempos) - 1] > 60000):
break
if size < 10:
size += 5
elif size < 100:
size +=10
elif size < 1000:
size += 100
else:
size += 500
#hacer para que matplotlib genere el grafico
plt.plot(tam, tiempos)
plt.title(titulo[i], fontsize=22)
plt.xlabel("Tamaño del tablero", fontsize=18)
plt.ylabel("Tiempo en ms", fontsize=18)
plt.show()
#limpiamos
tiempos.clear()
tam.clear()
# + id="zPNeA8LcoAD0" outputId="74ff655f-9c25-4670-de84-62ccf26bf98e" colab={"base_uri": "https://localhost:8080/", "height": 893}
main()
# + id="c8tuTvDs6vQ_"
| Informes/Informes TP/TP_Informe.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] _uuid="fb9097ae88608c29c159c1443947bc699486827c"
# <h1><center><font size="6">Santander Customer Transaction Prediction</font></center></h1>
# <h1><center><font size="5">Can you identify who will make a transaction?</font></center></h1>
#
# <img src="https://upload.wikimedia.org/wikipedia/commons/thumb/4/4a/Another_new_Santander_bank_-_geograph.org.uk_-_1710962.jpg/640px-Another_new_Santander_bank_-_geograph.org.uk_-_1710962.jpg" width="500"></img>
#
# <br>
# <b>
#
# Our data science team is continually challenging our machine learning algorithms, working with the global data science community to make sure we can more accurately identify new ways to solve our most common challenge, binary classification problems such as: is a customer satisfied? Will a customer buy this product? Can a customer pay this loan?
#
# In this challenge, we invite Kagglers to help us identify which customers will make a specific transaction in the future, irrespective of the amount of money transacted. The data provided for this competition has the same structure as the real data we have available to solve this problem.
#
# The data is anonimyzed, each row containing 200 numerical values identified just with a number.</b>
#
#
# + [markdown] _uuid="0f36c1c80b51c9e42180e18be0c8492f8733dec1"
# <a id=1><pre><b>Load Packages</b></pre></a>
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _kg_hide-input=true _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load in
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
from numba import jit
import lightgbm as lgb
from sklearn.metrics import roc_auc_score
from sklearn.model_selection import StratifiedKFold, KFold
import seaborn as sns
import matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
# Input data files are available in the "../input/" directory.
# For example, running this (by clicking run or pressing Shift+Enter) will list the files in the input directory
import os
import warnings
warnings.filterwarnings('ignore')
print(os.listdir("../input"))
# Any results you write to the current directory are saved as output.
# + [markdown] _uuid="a1f421494beba2a81bdf8bb126396cf3bd44089d"
# <a id=1><pre><b>Import the Data</b></pre></a>
# + _kg_hide-output=true _uuid="06cd64c0f3127126bdc26a2e6c58c776241a17d8"
train = pd.read_csv('../input/train.csv')
test = pd.read_csv('../input/test.csv')
features = [c for c in train.columns if c not in ['ID_code', 'target']]
target = train['target']
print ("Data is ready!")
# + _kg_hide-input=true _uuid="073d6a5c3be3de1954c5f50b42de85ffd9dce70e"
print ("Test ",test.shape)
print ("Train ",train.shape)
# + _uuid="913b6742afe21df29fb375b827b13fd0fe216206"
train.head(15)
# + [markdown] _uuid="c981dcd6094bf77b2430027d080e88a170c589b1"
# **Target = 0 or Target = 1, binary classification**
# + _uuid="617fb878c3c21297ee8eeafced14e6da724d8db5"
train.describe()
# + [markdown] _uuid="8a0ed560d6a2221493e969d2f0df0210901c8c4d"
# Let's see basic stats on the 2 different groups.
# + _uuid="8d1bd4c8d56f56d6682544a2fcf241b04312fb1d"
train[train['target']==0].describe()
# + _uuid="be00303167aad2db33cb086d7c01cc478e762ef8"
train[train['target']==1].describe()
# + [markdown] _uuid="d3c39873cca10b2277a97e604400edb2ecb0d5cc"
# **Missing data**
# + _uuid="5d810d2b297aa690f35302648983fd32468549c0"
print ("Missing data at training")
train.isnull().values.any()
# + _uuid="76198ed9effbb06e7066212155217650d086de68"
print ("Missing data at test")
test.isnull().values.any()
# + [markdown] _uuid="ec6500b9583ddb330b5c4ad481f926c70b2d4d71"
# **There is no missing data**
# + _uuid="8b8ab029992e553f4bbaae2e50b006db467447d7"
train = train.drop(["ID_code", "target"], axis=1)
# + [markdown] _uuid="acb08624d674014def07cb2a3930a791efcdc6f1"
# ### Check for Class Imbalance
# + _kg_hide-input=true _uuid="9228adbcbbcbbe4ad5c9d32179a8b66b35c78883"
sns.set_style('whitegrid')
sns.countplot(target)
sns.set_style('whitegrid')
# + [markdown] _uuid="6aff340a7d088efbd96e49bc4d68439e53322ed6"
# <a id=1><pre><b>Classification augment</b></pre></a>
# + _kg_hide-input=false _uuid="9f40ca4b8947c5be86b38141ab71ce68805d5ec6"
@jit
def augment(x,y,t=2):
xs,xn = [],[]
for i in range(t):
mask = y>0
x1 = x[mask].copy()
ids = np.arange(x1.shape[0])
for c in range(x1.shape[1]):
np.random.shuffle(ids)
x1[:,c] = x1[ids][:,c]
xs.append(x1)
for i in range(t//2):
mask = y==0
x1 = x[mask].copy()
ids = np.arange(x1.shape[0])
for c in range(x1.shape[1]):
np.random.shuffle(ids)
x1[:,c] = x1[ids][:,c]
xn.append(x1)
xs = np.vstack(xs)
xn = np.vstack(xn)
ys = np.ones(xs.shape[0])
yn = np.zeros(xn.shape[0])
x = np.vstack([x,xs,xn])
y = np.concatenate([y,ys,yn])
return x,y
# + [markdown] _uuid="50977096e5862aae1e81ee95213a6a5795e33a01"
# # Build the Light GBM Model
# + [markdown] _uuid="df044938b1cc6deb0b505274df0fc3cdb54ffdb4"
# <a id=1><pre><b>Parameters</b></pre></a>
# + _kg_hide-input=false _uuid="ce2a155cd34809f36d665ce38bb3ec632ca62746"
param = {
'bagging_freq': 5,
'bagging_fraction': 0.335,
'boost_from_average':'false',
'boost': 'gbdt',
'feature_fraction': 0.041,
'learning_rate': 0.0083,
'max_depth': -1,
'metric':'auc',
'min_data_in_leaf': 80,
'min_sum_hessian_in_leaf': 10.0,
'num_leaves': 13,
'num_threads': 8,
'tree_learner': 'serial',
'objective': 'binary',
'verbosity': -1
}
# + _uuid="8cf186957ca8ba2c018701178284190727edde87"
#kfold = 15
#folds = StratifiedKFold(n_splits=kfold, shuffle=False, random_state=44000)
num_folds = 11
features = [c for c in train.columns if c not in ['ID_code', 'target']]
folds = KFold(n_splits=num_folds, random_state=2319)
oof = np.zeros(len(train))
getVal = np.zeros(len(train))
predictions = np.zeros(len(target))
feature_importance_df = pd.DataFrame()
# + [markdown] _uuid="3ad089fe785780964b537d7b477a51eb42501e69"
# <a id=1><pre><b>Run LGBM model</b></pre></a>
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _kg_hide-input=true _kg_hide-output=true _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
for fold_, (trn_idx, val_idx) in enumerate(folds.split(train.values, target.values)):
X_train, y_train = train.iloc[trn_idx][features], target.iloc[trn_idx]
X_valid, y_valid = train.iloc[val_idx][features], target.iloc[val_idx]
X_tr, y_tr = augment(X_train.values, y_train.values)
X_tr = pd.DataFrame(X_tr)
print("Fold idx:{}".format(fold_ + 1))
trn_data = lgb.Dataset(X_tr, label=y_tr)
val_data = lgb.Dataset(X_valid, label=y_valid)
clf = lgb.train(param, trn_data, 1000000, valid_sets = [trn_data, val_data], verbose_eval=5000, early_stopping_rounds = 4000)
oof[val_idx] = clf.predict(train.iloc[val_idx][features], num_iteration=clf.best_iteration)
getVal[val_idx]+= clf.predict(train.iloc[val_idx][features], num_iteration=clf.best_iteration) / folds.n_splits
fold_importance_df = pd.DataFrame()
fold_importance_df["feature"] = features
fold_importance_df["importance"] = clf.feature_importance()
fold_importance_df["fold"] = fold_ + 1
feature_importance_df = pd.concat([feature_importance_df, fold_importance_df], axis=0)
predictions += clf.predict(test[features], num_iteration=clf.best_iteration) / folds.n_splits
# + _kg_hide-input=true _uuid="3e93e480b7f8404f0a460d4663696ab0b142b1dc"
print("\n >> CV score: {:<8.5f}".format(roc_auc_score(target, oof)))
# + _kg_hide-input=true _uuid="c65e08b0176a7b4f91ef95ea8e238f48934ba9cf"
cols = (feature_importance_df[["feature", "importance"]]
.groupby("feature")
.mean()
.sort_values(by="importance", ascending=False)[:1000].index)
best_features = feature_importance_df.loc[feature_importance_df.feature.isin(cols)]
plt.figure(figsize=(14,26))
sns.barplot(x="importance", y="feature", data=best_features.sort_values(by="importance",ascending=False))
plt.title('LightGBM Features (averaged over folds)')
plt.tight_layout()
plt.savefig('lgbm_importances.png')
# + [markdown] _uuid="757de0c47f13334bde694a06b3b4eba4c3ae9ad9"
# # Submission
# + _uuid="b2c5fc1d50c619bc36c1041e63c7a8802ffd8ac0"
submission = pd.DataFrame({"ID_code": test.ID_code.values})
submission["target"] = predictions
submission.to_csv("submission.csv", index=False)
# + _uuid="5fa54f6ffd8cbff04259aa8f56147655a49414bc"
submission.head()
| 12 customer prediction/santander-magic-lgb-0-901.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# # Conjugador de verbos en mapudungun
# ## Proponemos dos funciones principales
# ## (1) dado un verbo en infinitivo, conjugamos el verbo en presente (o pasado) en todas las personas gramaticales
# ## (2) dado un verbo conjugado, identificamos la persona y el tiempo verbal
#
# ### Para lograr esto, disponemos de un diccionario de verbos, lo que nos entrega el significado en espanol. En el caso de palabras OOV, nos basamos unicamente en la morfologia.
# # (1) verbo en espanol a verbo conjugado en mapudungun
import pandas as pd
import ipywidgets as widgets
widgets.interact_manual.opts['manual_name'] = 'mapudungun mew!'
## verbo en espanol a conjugacion en mapudungun
verbos=pd.read_csv('verbs.csv',header=0,sep='\t')
verbos = verbos.sort_values(['esp', 'mapu'], ascending=[1, 0])
verbos_esp=[verbo for verbo in verbos.esp]
verbos_mapu=[verbo for verbo in verbos.mapu]
personas={'singular':{'primera':'iñche','segunda':'eymi','tercera':'fey'},'dual':{'primera':'iñchiw','segunda':'eymu','tercera':'feyengu'},'plural':{'primera':'iñchiñ','segunda':'eymün','tercera':'feyengün'}}
consonantes=['n','w']
# -
## primera función
@widgets.interact_manual(verb_esp=verbos_esp, numero=['singular','dual','plural'], persona=['primera','segunda','tercera'],polaridad=['positiva','negativa'],tiempo=['futuro','no-futuro'])
def verb_to_mapudungun(verb_esp,numero,persona,polaridad,tiempo):
verbos={esp:mapu for (esp,mapu) in zip(verbos_esp,verbos_mapu)}
base=verbos[verb_esp]
conjugacion={'singular':{'primera':'(yo) iñche','segunda':'(tú) eymi','tercera':'(ella/él) fey'},'dual':{'primera':'(nosotras/nosotros dos) iñchiw','segunda':'(ustedes dos) eymu','tercera':'(ellas/ellos dos) feyengu'},'plural':{'primera':'(nosotras/nosotros) iñchiñ','segunda':'(ustedes) eymün','tercera':'(ellas/ellos) feyengün'}}
if polaridad=='positiva':## persona gramatical + base + futuro + polaridad
if tiempo=='futuro':
traduccion=conjugacion[numero][persona]+' '+base+'a'
else:
traduccion=conjugacion[numero][persona]+' '+base
elif polaridad=='negativa':
traduccion=conjugacion[numero][persona]+' '+base+'la'
if base[-1] in consonantes: ## terminan en consonante
if numero=='singular':
if persona=='primera':
traduccion=traduccion+'ün'
elif persona=='segunda':
traduccion=traduccion+'imi'
else:
traduccion=traduccion+'i'
elif numero=='dual':
if persona=='primera':
traduccion=traduccion+'iyu'
elif persona=='segunda':
traduccion=traduccion+'imu'
else:
traduccion=traduccion+'ingu'
else:
if persona=='primera':
traduccion=traduccion+'iyiñ'
elif persona=='segunda':
traduccion=traduccion+'imün'
else:
traduccion=traduccion+'ingün'
elif base[-1]=='i': ## termina en i
if numero=='singular':
if persona=='primera':
traduccion=traduccion+'n'
elif persona=='segunda':
traduccion=traduccion+'mi'
else:
traduccion=traduccion
elif numero=='dual':
if persona=='primera':
traduccion=traduccion+'yu'
elif persona=='segunda':
traduccion=traduccion+'mu'
else:
traduccion=traduccion+'ngu'
else:
if persona=='primera':
traduccion=traduccion+'iñ'
elif persona=='segunda':
traduccion=traduccion+'mün'
else:
traduccion=traduccion+'ngün'
else: ## en otro caso
if numero=='singular':
if persona=='primera':
traduccion=traduccion+'n'
elif persona=='segunda':
traduccion=traduccion+'ymi'
else:
traduccion=traduccion+'y'
elif numero=='dual':
if persona=='primera':
traduccion=traduccion+'yu'
elif persona=='segunda':
traduccion=traduccion+'ymu'
else:
traduccion=traduccion+'yngu'
else:
if persona=='primera':
traduccion=traduccion+'iñ'
elif persona=='segunda':
traduccion=traduccion+'ymün'
else:
traduccion=traduccion+'yngün'
return traduccion
| verb_morphology.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import numpy as np
from scipy import stats
import matplotlib.pyplot as plt
# -
# # Cellular automata & fractal dimension
# ## Define the automaton
# +
class Cellular1D:
def __init__(self, init_state, rule):
self.init_state = np.array(init_state)
self.rule = np.array(rule)
def run(self, n_timesteps, window=[4, 2, 1]): # Window [4, 2, 1] interprets the neighborhood as a binary number
"""Runs the automaton for `n_timesteps` steps.
Window 4, 2, 1 inspired by Downey's Think Complexity'"""
self.grid = np.zeros([n_timesteps + 1, self.init_state.shape[0]])
self.grid[0] = self.init_state
for i in range(1, self.grid.shape[0]):
c_i = np.correlate(self.grid[i - 1], window, mode='same').astype('int') # Cast to int for later indexing
self.grid[i] = self.rule[::-1][c_i]
return self.grid
def get_rule(rule):
"""Takes a decimal rule name and unpacks it to its binary representation"""
rule = np.array([rule], dtype='uint8')
return np.unpackbits(rule)
# -
def get_dimension(grid):
"""Computes fractal dimension of a 1D cellular automaton"""
cells = np.cumsum(grid.sum(axis=1))
steps = np.arange(grid.shape[0]) + 1
return stats.linregress(np.log(steps), np.log(cells))[0]
# ## Initialize and plot
# Define hyperparams
RULE = 18
STEPS = 500
WIDTH = 1001
P = .5
# +
# Define params
init_state = np.zeros(WIDTH)
# init_state = np.random.binomial(1, P, WIDTH)
# Set the middle point to 1
init_state[WIDTH // 2] = 1
rule = get_rule(RULE)
# Initialize the automaton
c1 = Cellular1D(init_state, rule)
grid = c1.run(STEPS)
# Get dimension
dimension = get_dimension(grid)
# Plot
plt.figure(figsize=(15, 200))
plt.imshow(grid, interpolation=None)
plt.axis('off')
plt.title(f'Dimension = ${dimension}$')
plt.show()
# -
| cellular-automata/02__1D_cellular_automaton_fractal_dimension.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import pickle
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score, confusion_matrix, classification_report
from yellowbrick.classifier import ConfusionMatrix
from sklearn.preprocessing import LabelEncoder
from sklearn.preprocessing import OneHotEncoder
from sklearn.compose import ColumnTransformer
from sklearn.impute import SimpleImputer
from sklearn.naive_bayes import GaussianNB
from sklearn.tree import DecisionTreeClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.svm import SVC
from sklearn.neural_network import MLPClassifier
with open('data/census.pkl', 'rb') as f:
x_census_treinamento, x_census_teste, y_census_treinamento, y_census_teste = pickle.load(f)
rede_neural_census = MLPClassifier(verbose=True, max_iter=1000, tol=0.000010, hidden_layer_sizes=(55,55))
rede_neural_census.fit(x_census_treinamento, y_census_treinamento)
previsoes = rede_neural_census.predict(x_census_teste)
previsoes
accuracy_score(y_census_teste, previsoes)
| estudo/aprendizagem-redes-neurais/rede-neural-censo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import seaborn as sns
import pandas as pd
import numpy as np
import re
pd.set_option('display.max_columns', 500)
#open csv, drop unnecessary columns
nba = pd.read_csv("../nba.csv")
nba = nba.drop(columns = ["Unnamed: 6", "Start (ET)","Notes","Unnamed: 7","Attend."])
#clean date and turn to date_time type
nba.Date = nba.Date.str.replace("^[A-z]{3}","-")
nba.Date.str.lstrip("- ")
nba.Date = nba.Date.str.replace(" ","-").str.lstrip("-")
nba.Date = pd.to_datetime(nba.Date)
#renaming columns
nba = nba.rename(columns = {"PTS":"AwayPTS", "PTS.1":"HomePTS", "Visitor/Neutral":"Away","Home/Neutral":"Home", "Attend.":"Attend"})
#
nba.head()
#creating homewins columns
nba["HomeWin"] = np.where(nba["HomePTS"] > nba["AwayPTS"], 1,0)
# +
# timeawaytest = nba.drop(columns = ["Date","AwayPTS","HomePTS"])
# timeawaytest["TimeAway"] = nbatest["Timeaway"]
# timeawaytest["HomeWinStreak"] = nbatest["HomeWinStreak"]
# timeawaytest["AwayWinStreak"] = nbatest["AwayWinStreak"]
# # timeawaytest[["AwayWinStreak","HomeLoseStreak","AwayLoseStreak","HomeCoachSavage","AwayCoachSavage"]]=nbatest[["AwayWinStreak","HomeLoseStreak","AwayLoseStreak","HomeCoachSavage","AwayCoachSavage"]]
# # timeawaytest[["HomeAllStars","AwayAllStars"]] = nbatest[["HomeAllstars","AwayAllstars"]]
# timeawaytest
# -
nbatest = nba[["Away","Home","HomeWin"]]
from sklearn import preprocessing
dummies = pd.get_dummies(nbatest[["Away","Home"]])
nbatest[dummies.columns] = dummies
nbatest = nbatest.drop(columns = ["Home","Away"])
nbatest
# +
#defining functions that check the accuracy score of our predictions using different models
#Random Forests
def RFscore(nbatest):
X_train = nbatest[:984].drop(columns = "HomeWin")
y_train = nbatest["HomeWin"][:984]
X_test = nbatest[984:].drop(columns = "HomeWin")
y_test = nbatest["HomeWin"][984:]
from sklearn.ensemble import RandomForestClassifier
from sklearn.datasets import make_classification
mod = RandomForestClassifier(n_estimators = 100)
mod.fit(X_train, y_train)
y_pred = mod.predict(X_test)
from sklearn.metrics import accuracy_score
return accuracy_score(y_test,y_pred)
#SVC
def SVCscore(nbatest):
#train data/ test data
X_train = nbatest[:984].drop(columns = "HomeWin")
y_train = nbatest["HomeWin"][:984]
X_test = nbatest[984:].drop(columns = "HomeWin")
y_test = nbatest["HomeWin"][984:]
#fit model
from sklearn.svm import SVC
svc = SVC(gamma='auto')
svc.fit(X_train, y_train)
y_pred = svc.predict(X_test)
from sklearn.metrics import accuracy_score
return accuracy_score(y_test,y_pred)
#LogisticRegression
def LRscore(nbatest):
X_train = nbatest[:984].drop(columns = "HomeWin")
y_train = nbatest["HomeWin"][:984]
X_test = nbatest[984:].drop(columns = "HomeWin")
y_test = nbatest["HomeWin"][984:]
from sklearn.linear_model import LogisticRegression
mod = LogisticRegression()
mod.fit(X_train,y_train)
y_pred = mod.predict(X_test)
from sklearn.metrics import accuracy_score
return accuracy_score(y_test,y_pred)
# -
print("Logistic Regression Accuracy Score: ",LRscore(nbatest),"\nSVC Accuracy score:",SVCscore(nbatest),"\nRandom Forests Accuracy score:",RFscore(nbatest))
# +
#Adding "time on the road" feature
nbatest["Timeaway"] = 0
away_counts = {}
for i in nba["Away"].unique():
away_counts[i] = 0
for i in range(len(nba.Away)):
away_counts[nba.Away[i]] +=1
away_counts[nba.Home[i]] = 0
nbatest.iloc[i,-1] = away_counts[nba.Away[i]]
print("Logistic Regression Accuracy Score: ",LRscore(nbatest),"\nSVC Accuracy score:",SVCscore(nbatest),"\nRandom Forests Accuracy score:",RFscore(nbatest))
# -
from sklearn import preprocessing
scaler = preprocessing.StandardScaler().fit(nbatest[["Timeaway"]])
scaled = scaler.transform(nbatest[["Timeaway"]])
nbatest[["Timeaway"]]= scaled
print("Logistic Regression Accuracy Score: ",LRscore(nbatest),"\nSVC Accuracy score:",SVCscore(nbatest),"\nRandom Forests Accuracy score:",RFscore(nbatest))
# +
#adding Home win streak and Away win streak features
nbatest["HomeWinStreak"] = 0
nbatest["AwayWinStreak"] = 0
win_counts = {}
for i in nba["Away"].unique():
win_counts[i] = 0
for row in range(len(nba)):
nbatest.iloc[row,-2] = win_counts[nba["Home"][row]]
nbatest.iloc[row,-1] = win_counts[nba["Away"][row]]
if nbatest["HomeWin"][row] == 1:
win_counts[nba["Home"][row]] +=1
win_counts[nba["Away"][row]] == 0
else:
win_counts[nba["Away"][row]] +=1
win_counts[nba["Home"][row]] == 0
print("Logistic Regression Accuracy Score: ",LRscore(nbatest),"\nSVC Accuracy score:",SVCscore(nbatest),"\nRandom Forests Accuracy score:",RFscore(nbatest))
# -
from sklearn import preprocessing
scaler = preprocessing.StandardScaler().fit(nbatest[["HomeWinStreak","AwayWinStreak"]])
scaled = scaler.transform(nbatest[["HomeWinStreak","AwayWinStreak"]])
nbatest[["HomeWinStreak","AwayWinStreak"]]= scaled
print("Logistic Regression Accuracy Score: ",LRscore(nbatest),"\nSVC Accuracy score:",SVCscore(nbatest),"\nRandom Forests Accuracy score:",RFscore(nbatest))
def SVCrandscore(nbatest):
#train data/ test data
from sklearn.model_selection import train_test_split
X = nbatest.drop(columns = "HomeWin")
y = nbatest["HomeWin"]
X_train,X_test,y_train,y_test = train_test_split(X,y,train_size=.8)
#fit model
from sklearn.svm import SVC
svc = SVC(gamma='auto')
svc.fit(X_train, y_train)
y_pred = svc.predict(X_test)
from sklearn.metrics import accuracy_score
return accuracy_score(y_test,y_pred)
# +
#adding Home lose streak and Away lose streak features
nbatest["HomeLoseStreak"] = 0
nbatest["AwayLoseStreak"] = 0
loss_counts = {}
for i in nba["Away"].unique():
loss_counts[i] = 0
for row in range(len(nba)):
if nbatest["HomeWin"][row] == 1:
loss_counts[nba["Away"][row]] +=1
loss_counts[nba["Home"][row]] == 0
else:
loss_counts[nba["Home"][row]] +=1
loss_counts[nba["Away"][row]] == 0
nbatest.iloc[row,-2] = loss_counts[nba["Home"][row]]
nbatest.iloc[row,-1] = loss_counts[nba["Away"][row]]
print("Logistic Regression Accuracy Score: ",LRscore(nbatest),"\nSVC Accuracy score:",SVCscore(nbatest),"\nRandom Forests Accuracy score:",RFscore(nbatest))
# -
from sklearn import preprocessing
scaler = preprocessing.StandardScaler().fit(nbatest[["HomeLoseStreak","AwayLoseStreak"]])
scaled = scaler.transform(nbatest[["HomeLoseStreak","AwayLoseStreak"]])
nbatest[["HomeLoseStreak","AwayLoseStreak"]]= scaled
print("Logistic Regression Accuracy Score: ",LRscore(nbatest),"\nSVC Accuracy score:",SVCscore(nbatest),"\nRandom Forests Accuracy score:",RFscore(nbatest))
# +
nbatest["HomeCoachSavage"] = 0
nbatest["AwayCoachSavage"] = 0
nbatest.loc[nba["Home"]=="San Antonio Spurs","HomeCoachSavage"] = 6
nbatest.loc[nba["Home"]=="Boston Celtics","HomeCoachSavage"] = 5
nbatest.loc[nba["Home"]=="Golden State Warriors","HomeCoachSavage"] = 4
nbatest.loc[nba["Home"]=="Utah Jazz","HomeCoachSavage"] = 3
nbatest.loc[nba["Home"]=="Houston Rockets","HomeCoachSavage"] = 2
nbatest.loc[nba["Home"]=="Toronto Raptors","HomeCoachSavage"] = 1
nbatest.loc[nba["Away"]=="San Antonio Spurs","AwayCoachSavage"] = 6
nbatest.loc[nba["Away"]=="Boston Celtics","AwayCoachSavage"] = 5
nbatest.loc[nba["Away"]=="Golden State Warriors","AwayCoachSavage"] = 4
nbatest.loc[nba["Away"]=="Utah Jazz","AwayCoachSavage"] = 3
nbatest.loc[nba["Away"]=="Houston Rockets","AwayCoachSavage"] = 2
nbatest.loc[nba["Away"]=="Toronto Raptors","AwayCoachSavage"] = 1
print("Logistic Regression Accuracy Score: ",LRscore(nbatest),"\nSVC Accuracy score:",SVCscore(nbatest),"\nRandom Forests Accuracy score:",RFscore(nbatest))
# -
from sklearn import preprocessing
scaler = preprocessing.StandardScaler().fit(nbatest[["HomeCoachSavage","AwayCoachSavage"]])
scaled = scaler.transform(nbatest[["HomeCoachSavage","AwayCoachSavage"]])
nbatest[["HomeCoachSavage","AwayCoachSavage"]]= scaled
print("Logistic Regression Accuracy Score: ",LRscore(nbatest),"\nSVC Accuracy score:",SVCscore(nbatest),"\nRandom Forests Accuracy score:",RFscore(nbatest))
allstar_count = {}
for i in nba.Home.unique():
allstar_count[i] = 0
allstar_count['Philadelphia 76ers'] = 2
allstar_count['Milwaukee Bucks'] = 2
allstar_count['Oklahoma City Thunder']=2
allstar_count['Golden State Warriors'] = 3
allstar_count['Denver Nuggets'] = 1
allstar_count['Detroit Pistons'] = 1
allstar_count['Brooklyn Nets'] = 1
allstar_count['Orlando Magic'] = 1
allstar_count['Toronto Raptors'] = 2
allstar_count['Dallas Mavericks'] = 1
allstar_count['Los Angeles Lakers'] = 1
allstar_count['Houston Rockets'] = 1
allstar_count['Orlando Magic'] = 1
allstar_count['Boston Celtics'] = 1
allstar_count['New Orleans Pelicans'] = 1
allstar_count[ 'Portland Trail Blazers'] = 1
allstar_count['San Antonio Spurs'] = 1
allstar_count['Minnesota Timberwolves'] = 1
allstar_count['Washington Wizards'] = 1
allstar_count['Miami Heat'] = 1
nbatest["HomeAllstars"] = 0
nbatest["AwayAllstars"] = 0
for i in range(len(nba)):
nbatest.iloc[i,-1] = allstar_count[nba["Away"][i]]
nbatest.iloc[i,-2] = allstar_count[nba["Home"][i]]
print("Logistic Regression Accuracy Score: ",LRscore(nbatest),"\nSVC Accuracy score:",SVCscore(nbatest),"\nRandom Forests Accuracy score:",RFscore(nbatest))
from sklearn import preprocessing
scaler = preprocessing.StandardScaler().fit(nbatest[["HomeAllstars","AwayAllstars"]])
scaled = scaler.transform(nbatest[["HomeAllstars","AwayAllstars"]])
nbatest[["HomeAllstars","AwayAllstars"]]= scaled
print("Logistic Regression Accuracy Score: ",LRscore(nbatest),"\nSVC Accuracy score:",SVCscore(nbatest),"\nRandom Forests Accuracy score:",RFscore(nbatest))
# +
depth = pd.read_csv("depth.csv")
depth["avg"] = (depth["bench1"]+depth["bench2"]+depth["bench3"]+depth["bench4"]+depth["bench5"])/5
# depth
# -
depth.sort_values("avg")
depth_dict ={}
for i in depth.team.unique():
depth_dict[i] = float(depth[depth["team"] == i]["avg"])
nba.head()
print(depth_dict, nba.head())
nbatest["HomeBenchRating"] = 0
nbatest["AwayBenchRating"] = 0
for i in range(len(nba)):
nbatest.iloc[i,-1] = depth_dict[nba.Away[i]]
nbatest.iloc[i,-2] = depth_dict[nba.Home[i]]
print("Logistic Regression Accuracy Score: ",LRscore(nbatest),"\nSVC Accuracy score:",SVCscore(nbatest),"\nRandom Forests Accuracy score:",RFscore(nbatest))
nbatest.head(10)
from sklearn import preprocessing
scaler = preprocessing.StandardScaler().fit(nbatest[["AwayBenchRating","HomeBenchRating"]])
scaled = scaler.transform(nbatest[["AwayBenchRating","HomeBenchRating"]])
nbatest[["AwayBenchRating","HomeBenchRating"]]= scaled
print("Logistic Regression Accuracy Score: ",LRscore(nbatest),"\nSVC Accuracy score:",SVCscore(nbatest),"\nRandom Forests Accuracy score:",RFscore(nbatest))
score_dict = {}
for i in nba.Home.unique():
score_dict[i] = {"counter" : 0, "totalscore" : 0}
score_dict["Miami Heat"]
nbatest["AwayAvgScore"] = 0
nbatest["HomeAvgScore"] = 0
for i in range(len(nba)):
#Home team counter and total score
score_dict[nba.Home[i]]["counter"] +=1
score_dict[nba.Home[i]]["totalscore"] += nba.HomePTS[i]
#Away team counter and score
score_dict[nba.Away[i]]["counter"]+= 1
score_dict[nba.Away[i]]["totalscore"] += nba.AwayPTS[i]
nbatest.iloc[i,-1] = score_dict[nba.Home[i]]["totalscore"] / score_dict[nba.Home[i]]["counter"]
nbatest.iloc[i,-2] = score_dict[nba.Away[i]]["totalscore"] / score_dict[nba.Away[i]]["counter"]
print("Logistic Regression Accuracy Score: ",LRscore(nbatest),"\nSVC Accuracy score:",SVCscore(nbatest),"\nRandom Forests Accuracy score:",RFscore(nbatest))
from sklearn import preprocessing
scaler = preprocessing.StandardScaler().fit(nbatest[["AwayAvgScore","HomeAvgScore"]])
scaled = scaler.transform(nbatest[["AwayAvgScore","HomeAvgScore"]])
nbatest[["AwayAvgScore","HomeAvgScore"]]= scaled
print("Logistic Regression Accuracy Score: ",LRscore(nbatest),"\nSVC Accuracy score:",SVCscore(nbatest),"\nRandom Forests Accuracy score:",RFscore(nbatest))
sns.heatmap(nbatest[['Timeaway', 'HomeWinStreak', 'AwayWinStreak',
'HomeLoseStreak', 'AwayLoseStreak', 'HomeCoachSavage',
'AwayCoachSavage', 'HomeAllstars', 'AwayAllstars',"HomeBenchRating","AwayBenchRating"]].corr())
#RANKINGS
#Creates dictionary with team and their rankings
feb_ranks = pd.read_csv("feb_ranks.csv")
feb_ranks["Team"] = feb_ranks["Western Conference"]
rankdict = {}
for i in range(len(feb_ranks.Team)):
rankdict[feb_ranks.Team[i]] = feb_ranks.Rk[i]
#Creates HomeRank and AwayRank for each matchup in nba dataset
nbatest["HomeRank"] = 0
nbatest["AwayRank"] = 0
for i in range(len(nbatest.HomeRank)):
#Setting AwayRank for row i
nbatest.iloc[i,-1] = rankdict[nba.Away[i]]
#Setting HomeRank for row i
nbatest.iloc[i,-2] = rankdict[nba.Home[i]]
from sklearn import preprocessing
scaler = preprocessing.StandardScaler().fit(nbatest[["HomeRank","AwayRank"]])
scaled = scaler.transform(nbatest[["HomeRank","AwayRank"]])
nbatest[["HomeRank","AwayRank"]]= scaled
print("Logistic Regression Accuracy Score: ",LRscore(nbatest),"\nSVC Accuracy score:",SVCscore(nbatest),"\nRandom Forests Accuracy score:",RFscore(nbatest))
# +
#creates a single column that checks if the home team is ranked higher than away team
rankspread = nbatest["HomeRank"]-nbatest["AwayRank"]
nbatest["HomeRanksHigher"]= 0
#if rankspread is positive, home is better
#if rankspread is negative, away is better
for i in range(len(nbatest)):
if rankspread[i] > 0:
nbatest.iloc[i,-1] = 1
else:
nbatest.iloc[i,-1] = 0
print("Logistic Regression Accuracy Score: ",LRscore(nbatest),"\nSVC Accuracy score:",SVCscore(nbatest),"\nRandom Forests Accuracy score:",RFscore(nbatest))
# -
# +
#creates a single column that checks if the home team is significantly better than the away team
rankspread = nbatest["HomeRank"]-nbatest["AwayRank"]
nbatest["HomeisFav"]= 0
for i in range(len(nbatest)):
if rankspread[i] > 0 and rankspread[i] >5:
nbatest.iloc[i,-1] = 1
elif rankspread[i]<0 and abs(rankspread[i])>5:
nbatest.iloc[i,-1] = 0
else:
nbatest.iloc[i,-1] = 0
nbatest.drop(columns = ["HomeRank","AwayRank"], inplace = True)
print("Logistic Regression Accuracy Score: ",LRscore(nbatest),"\nSVC Accuracy score:",SVCscore(nbatest),"\nRandom Forests Accuracy score:",RFscore(nbatest))
# +
nbatest[["HomeWinStreak","AwayWinStreak"]]
# -
| nba-winloss-predictor/final/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Model 1
# +
import re
from sklearn.model_selection import train_test_split
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.pipeline import make_pipeline
from sklearn.linear_model import LogisticRegression
neu = '0'
pos = '0'
neg = '1'
title='aif'
f = open('/home/ydw/capston/python/data/sum(fromPH.D+twitter)/text.txt', 'r', encoding='utf-8')
text = f.read().splitlines()
f.close()
f = open('/home/ydw/capston/python/data/sum(fromPH.D+twitter)/result.txt', 'r', encoding='utf-8')
r_y = f.read().splitlines()
f.close()
y = []
for i in r_y:
if(i == '2'):
y.append(neu)
elif(i == '0'):
y.append(pos)
else:
y.append(neg)
f = open('/home/ydw/capston/python/data/twitter/positive.txt', 'r', encoding='utf-8')
p_text = f.read().splitlines()
f.close()
f = open('/home/ydw/capston/python/data/twitter/negative.txt', 'r', encoding='utf-8')
n_text = f.read().splitlines()
f.close()
p_text = p_text[35000:35000+3000]
n_text = n_text[35000:35000+3000]
p_text = [' '.join(re.sub("(RT )|(@\S+)|(\w+:\/\/\S+)|", "", doc).split()) for doc in p_text]
n_text = [' '.join(re.sub("(RT )|(@\S+)|(\w+:\/\/\S+)|", "", doc).split()) for doc in n_text]
for i in p_text:
text.append(i)
y.append(pos)
for i in n_text:
text.append(i)
y.append(neg)
#One-hot encoding Model
text_train, text_test, y_train, y_test = train_test_split(text, y, test_size=0.2, random_state=1)
pipe_b = make_pipeline(CountVectorizer(ngram_range=(1,3), min_df=0), LogisticRegression())
pipe_b.fit(text_train, y_train)
f = open('/home/ydw/capston/python/data/application/reply_{}_100.txt'.format(title), 'r', encoding='utf-8')
text = f.read().splitlines()
f.close()
f = open('/home/ydw/capston/python/data/application/{}.txt'.format(title), 'r', encoding='utf-8')
r_y = f.read().splitlines()
f.close()
y = []
for i in r_y:
if(i == 'l'):
y.append(neu)
elif(i == 'p'):
y.append(pos)
elif(i == 'n'):
y.append(neg)
else:
y.append('NA')
pre_y = []
for doc in text:
rdoc = [doc]
pre_y.append(pipe_b.predict(rdoc)[0])
count = 0
for i in range(len(y)):
if(y[i] == pre_y[i]):
count = count+1
a = pre_y
count/len(y), pre_y.count('0'), y.count('0')
# -
# # Model 2
# +
import re
import numpy as np
import fasttext
neu = '0'
pos = '1'
neg = '0'
title='aif'
f = open('/home/ydw/capston/python/data/expert/neutral.txt', 'r', encoding='utf-8')
text = f.read().splitlines()
f.close()
y = []
for i in range(len(text)):
y.append(neu)
f = open('/home/ydw/capston/python/data/twitter/positive.txt', 'r', encoding='utf-8')
p_text = f.read().splitlines()
f.close()
f = open('/home/ydw/capston/python/data/twitter/negative.txt', 'r', encoding='utf-8')
n_text = f.read().splitlines()
f.close()
p_text = p_text[35000:35000+3000]
n_text = n_text[35000:35000+3000]
p_text = [' '.join(re.sub("(RT )|(@\S+)|(\w+:\/\/\S+)|", "", doc).split()) for doc in p_text]
n_text = [' '.join(re.sub("(RT )|(@\S+)|(\w+:\/\/\S+)|", "", doc).split()) for doc in n_text]
for i in p_text:
text.append(i)
y.append(pos)
for i in n_text:
text.append(i)
y.append(neg)
#fasttext Model
for i in range(len(text)):
text[i] = '__label__'+y[i]+' '+text[i]+'\n'
fasttext_train = open("/home/ydw/capston/python/data/sum(fromPH.D+twitter)/fasttext_data/fasttext_train.txt", "w", encoding='utf-8')
fasttext_test = open("/home/ydw/capston/python/data/sum(fromPH.D+twitter)/fasttext_data/fasttext_test.txt", "w", encoding='utf-8')
for i in text:
if(np.random.uniform() < 0.2):
fasttext_test.write(i)
else:
fasttext_train.write(i)
classifier = fasttext.supervised('/home/ydw/capston/python/data/sum(fromPH.D+twitter)/fasttext_data/fasttext_train.txt',
'/home/ydw/capston/python/data/sum(fromPH.D+twitter)/fasttext_data/classifier',
dim=20, loss='softmax', epoch=10, label_prefix='__label__')
f = open('/home/ydw/capston/python/data/application/reply_{}_100.txt'.format(title), 'r', encoding='utf-8')
text = f.read().splitlines()
f.close()
f = open('/home/ydw/capston/python/data/application/{}.txt'.format(title), 'r', encoding='utf-8')
r_y = f.read().splitlines()
f.close()
y = []
for i in r_y:
if(i == 'l'):
y.append(neu)
elif(i == 'p'):
y.append(pos)
elif(i == 'n'):
y.append(neg)
else:
y.append('NA')
pre_y = []
for doc in text:
rdoc = [doc]
pre_y.append(classifier.predict(rdoc)[0][0])
count = 0
for i in range(len(y)):
if(y[i] == pre_y[i]):
count = count+1
b = pre_y
count/len(y), pre_y.count('0'), y.count('0')
# -
# # Cross
index = []
re_text = []
for i in range(len(a)):
if(a[i]=='1' and b[i]=='1'):
index.append(i)
re_text.append(text[i])
a[i] = '3'
elif(a[i]=='0' and b[i]=='1'):
a[i] = '0'
elif(a[i]=='0' and b[i]=='0'):
a[i] = '2'
# +
f = open('/home/ydw/capston/python/data/twitter/positive.txt', 'r', encoding='utf-8')
p_text = f.read().splitlines()
f.close()
f = open('/home/ydw/capston/python/data/twitter/negative.txt', 'r', encoding='utf-8')
n_text = f.read().splitlines()
f.close()
p_text = p_text[:20000]
n_text = n_text[:20000]
p_text = [' '.join(re.sub("(RT )|(@\S+)|(\w+:\/\/\S+)|", "", doc).split()) for doc in p_text]
n_text = [' '.join(re.sub("(RT )|(@\S+)|(\w+:\/\/\S+)|", "", doc).split()) for doc in n_text]
text = []
y = []
for i in p_text:
text.append(i)
y.append('0')
for i in n_text:
text.append(i)
y.append('1')
text_train, text_test, y_train, y_test = train_test_split(text, y, test_size=0.2, random_state=1)
pipe_n = make_pipeline(CountVectorizer(ngram_range=(1,3), min_df=0), LogisticRegression())
pipe_n.fit(text_train, y_train)
# +
pre_re_y = []
for doc in re_text:
rdoc = [doc]
pre_re_y.append(pipe_n.predict(rdoc)[0])
k = 0
for i in index:
a[i] = pre_re_y[k]
k=k+1
# -
# # Test
# +
f = open('/home/ydw/capston/python/data/application/{}.txt'.format(title), 'r', encoding='utf-8')
r_y = f.read().splitlines()
f.close()
y = []
for i in r_y:
if(i == 'l'):
y.append('2')
elif(i == 'p'):
y.append('0')
elif(i == 'n'):
y.append('1')
else:
y.append('NA')
count = 0
for i in range(len(y)):
if(y[i] == a[i]):
count = count+1
print(count/len(y))
y.count('0'), a.count('0'),y.count('1'), a.count('1'),y.count('2'), a.count('2')
| 03_10_Two_label cross.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="scikit.png" align="center">
#
# <h2 align="center">Simple Linear Regression</h2>
# - Introduction
# - Data Set
# - Loading the Data and Importing Libraries
# - Gathering Data
# - Exploratory Data Analysis
# - Creating Simple Linear Regression
# - Multiple Linear Regression Model
# ## <font color='brown'>Introduction</font>
# Linear Regression is a useful tool for predicting a quantitative response.
# We have an input vector $X^T = (X_1, X_2,...,X_p)$, and want to predict a real-valued output $Y$. The linear regression model has the form
# <h4 align="center"> $f(x) = \beta_0 + \sum_{j=1}^p X_j \beta_j$. </h4>
# The linear model either assumes that the regression function $E(Y|X)$ is linear, or that the linear model is a reasonable approximation.Here the $\beta_j$'s are unknown parameters or coefficients, and the variables $X_j$ can come from different sources. No matter the source of $X_j$, the model is linear in the parameters.
# **Simple Linear Regression**: <h5 align=center>$$Y = \beta_0 + \beta_1 X + \epsilon$$</h5>
# **Multiple Linear Regression**: <h5 align=center>$$Y = \beta_0 + \beta_1 X_1 + \beta_2 X_2 +...+ \beta_p X_p + \epsilon$$ </h5>
# <h5 align=center> $$sales = \beta_0 + \beta_1 \times TV + \beta_2 \times radio + \beta_3 \times newspaper + \epsilon$$ </h5>
# - $sales: $ predictor or feature
# - $\beta_0: $ slope coefficient
# - $\beta_1, \beta_2, \beta_3: $ intercept terms
# - TV, radio, newspaper: response or target variables
# ## <font color='brown'>Data Set</font>
# The adverstiting dataset captures sales revenue generated with respect to advertisement spends across multiple channles like radio, tv and newspaper. [Source](http://faculty.marshall.usc.edu/gareth-james/ISL/data.html)
# ## <font color='brown'>Loading the Data and Importing Libraries</font>
import pandas as pd
import numpy as np
import seaborn as sns
from scipy.stats import skew
from sklearn.linear_model import LinearRegression
from sklearn.metrics import r2_score
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error
# !pip install yellowbrick
from yellowbrick.regressor import PredictionError, ResidualsPlot
from sklearn import metrics
# %matplotlib inline
import matplotlib.pyplot as plt
plt.style.use("ggplot")
plt.rcParams['figure.figsize'] = (12, 8)
print("libraries imported..")
# ## <font color='brown'>Software Needed</font>
# Software: Python and Jupyter Notebook
#
# The following packages (libraries) need to be installed:
#
# 1. pandas
# 2. NumPy
# 3. scikit Learn
# 4. Yellow Brick
#
# ## <font color='brown'>**Gathering Data**</font>
df = pd.read_csv("data/Advertising.csv")
df.head()
df.shape
# We can see that ther are no missing value.
#finding null value
df.isnull().sum()
#Let's find duplicate data set
df.duplicated().sum()
df.describe()
df.info()
#lets drop Unnamed:0 Column from data frame using index base method
df.drop(df.columns[[0]], axis = 1, inplace = True)
df.info()
# ## <font color='brown'>**Exploratory Data Analysis**</font>
#
# +
sns.distplot(df.sales)
# -
sns.distplot(df.newspaper)
sns.distplot(df.radio)
# ### <font color='green'>**Exploring Relationships between Predictors and Response**</font>
#
sns.pairplot(df, x_vars=['TV','radio','newspaper'], y_vars='sales', height=7,
aspect =0.7, kind='reg')
sns.pairplot(df, x_vars=['TV','radio','newspaper'], y_vars='sales', height=7, aspect=0.7);
df.TV.corr(df.sales)
df.corr()
sns.heatmap(df.corr(),annot=True)
# Tv is hihgly corr with sales
# ## <font color='brown'>**Creating the Simple Linear Regression Model**</font>
#
X = df[['TV']]
X.head()
y = df.sales
type(y)
# Since its panad series we can use scitkit function
#
X_train,X_test, y_train, y_test = train_test_split(X,y, random_state=1)
print(X_train.shape)
print(X_test.shape)
print(y_train.shape)
print(y_test.shape)
lm0 = LinearRegression()
lm0.fit(X_train, y_train)
# ### <font color='blue'>Interpreting Model Coefficients</font>
#
print(lm0.intercept_)
print(lm0.coef_)
# B0 = 6.91, B1 = 0.48 coef associate with spending on tv ads. For given amount of radio and news paper ads spending a unit increased in Tv ads spending is associated with a 0.048 unit increased in the sales revune. for a given amount of radio and newspaper ads spending an additional 1000 USD spend on TV ads is associated with an increase in sales of 48 items.
# This is the statement associate with corr not causation.If increase in tv ads spending was associated with decreasing in sales then B1 i.e coef would be negative.
# ### <font color='blue'>Making Predictions with our Model</font>
#
#making prediction on test set
y_pred = lm0.predict(X_test) #this is going to make pred on 25% of test set data
y_pred[:5] #since it is numpy array we use this method
# This are first 5 values of the predicted sales reveune on test set.
#
# Now we need to compare our predicted value with actual value, hence evaluation metrics comes into play.
# ## <font color='brown'>**Multiple Linear Regression Model**</font>
#
# +
# create X and y
feature_cols = ['TV', 'radio', 'newspaper']
X = df[feature_cols]
y = df.sales
# instantiate and fit
lm1 = LinearRegression() #model 1m1 refer to first model
lm1.fit(X, y)
# print the coefficients
print(lm1.intercept_)
print(lm1.coef_)
# -
# for given amount of Tv and news paper ads spending an additional
# usd 1000 on Tv ads, radio ads leads to an increase in sales by apporx 46 and 189 units respectively.coef of news paper was significanlty non zero or close to zero.
# pair the feature names with the coefficients
list(zip(feature_cols, lm1.coef_))
sns.heatmap(df.corr(), annot=True)
# we can see that more is spend on newspaper compared to radio, but news paper ads has no direct impact on sales. The market where more money is spend on radio ads leads to more sale compared to newspaper.
# ### <font color='blue'> Feature Selection</font>
#
# How well does the model fit the data? what response value should predict and how accurate is our prediction?
# Which predictor are associated with response inorder to git a single model ivolving all those predictor this process is known as feautre selection or varaible selection.
# +
lm2 = LinearRegression().fit(X[['TV', 'radio']], y) #lm2 second model with Tv, Radio only
lm2_preds = lm2.predict(X[['TV', 'radio']])
print("R^2: ", r2_score(y, lm2_preds))
# +
lm3 = LinearRegression().fit(X[['TV', 'radio', 'newspaper']], y) #lm3 third model with newspaper
lm3_preds = lm3.predict(X[['TV', 'radio', 'newspaper']])
print("R^2: ", r2_score(y, lm3_preds))
# -
# we can see here that model that uses all three advs media to predice sales
# and that those use only two Tv and Radio has similar R square value meaning newspaper does not have impact on our data so we can drop this newspaper column.
# ### <font color='blue'>Model Evaluation Using Train/Test Split and Metrics</font>
#
# **Mean Absolute Error** (MAE) is the mean of the absolute value of the errors: <h5 align=center>$$\frac{1}{n}\sum_{i=1}^{n} \left |y_i - \hat{y_i} \right |$$</h5>
# **Mean Squared Error** (MSE) is the mean of the squared errors: <h5 align=center>$$\frac{1}{n}\sum_{i=1}^{n} (y_i - \hat{y_i})^2$$</h5>
# **Root Mean Squared Error** (RMSE) is the mean of the squared errors: <h5 align=center>$$\sqrt{\frac{1}{n}\sum_{i=1}^{n} (y_i - \hat{y_i})^2}$$</h5>
# Let's use train/test split with RMSE to see whether newspaper should be kept in the model:
# RMSE calcauset diff btn acutal value and predicated value of the response variable. The lesser the value of RMSE the better is the model
# +
X = df[['TV', 'radio', 'newspaper']]
y = df.sales
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state = 1)
lm4 = LinearRegression() #l4 indicates fourth model
lm4.fit(X_train, y_train)
lm4_preds = lm4.predict(X_test)
print("RMSE :", np.sqrt(mean_squared_error(y_test, lm4_preds)))
print("R^2: ", r2_score(y_test, lm4_preds))
# +
X = df[['TV', 'radio']]
y = df.sales
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state = 1)
lm5 = LinearRegression() #lm5 fifth model without newspaper
lm5.fit(X_train, y_train)
lm5_preds = lm5.predict(X_test)
print("RMSE :", np.sqrt(mean_squared_error(y_test, lm5_preds)))
print("R^2: ", r2_score(y_test, lm5_preds))
# -
# we can see when we omitted newspaper oyr RMSE decread and our R square increased so this is better model for us.
# +
#Data visualization and model diagnostic library called yellow brick
visualizer = PredictionError(lm5)
visualizer.fit(X_train, y_train) # Fit the training data to the visualizer
visualizer.score(X_test, y_test) # Evaluate the model on the test data
visualizer.poof()
# -
visualizer = ResidualsPlot(lm5)
visualizer.fit(X_train, y_train)
visualizer.score(X_test, y_test)
visualizer.poof()
# here we can see pred erorr plot for mult lin reg model and we can see here our line of identity and line of best fit. although our R square is high there are still some unexpaleind varaiblity in our data , model is unable to caputre and this is due to interaction effect or better known as Synergy.
# ### <font color='blue'>Interaction Effect (Synergy)</font>
#
#
# one way of extending this model to allow for interaction effect is to include third predictor lets call it interaction term whihc is constructed by computing the product of x1,x2, and so on that is mult of value corresponding to Tv and radio column.
df['interaction'] = df['TV'] * df['radio']
# +
X = df[['TV', 'radio', 'interaction']]
y = df.sales
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state = 1)
lm6 = LinearRegression() #our 6th model
lm6.fit(X_train, y_train)
lm6_preds = lm6.predict(X_test)
print("RMSE :", np.sqrt(mean_squared_error(y_test, lm6_preds)))
print("R^2: ", r2_score(y_test, lm6_preds))
# -
# r square is 97% and dramtic decrease of RMSE which is very good sing for us.
# +
visualizer = PredictionError(lm6)
visualizer.fit(X_train, y_train) # Fit the training data to the visualizer
visualizer.score(X_test, y_test) # Evaluate the model on the test data
visualizer.poof()
# -
# here we can see that this model interation term fitted nicely with our data plot. It explain that about 97% of the variablity in the data.So we can conclude that lm6 this model is best model than other model above.
| Scikit_learn_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=true editable=true
# # Illegal Fishing Data Exploration
# + deletable=true editable=true
import tensorflow as tf
import numpy as np
import scipy as sp
# + deletable=true editable=true
import os
os.chdir("/Users/jonathangessert/Dev/illegal-fishing/")
print(os.getcwd())
# + deletable=true editable=true
data = np.load('/Users/jonathangessert/Dev/illegal-fishing/training-data/data/tracks/100043174358994.npz')
# + deletable=true editable=true
| .ipynb_checkpoints/notebook-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.5 64-bit (conda)
# metadata:
# interpreter:
# hash: b3ba2566441a7c06988d0923437866b63cedc61552a5af99d1f4fb67d367b25f
# name: python3
# ---
# +
from azureml.core import Workspace, Datastore, Dataset
# List workspace
ws = Workspace.from_config()
ws_list = Workspace.list(subscription_id="123456")
print(ws_list['dp-100'])
# -
# Display default Datastore
default_ds = ws.get_default_datastore()
print(default_ds)
# List Datastores
store_list = list(ws.datastores)
print(store_list)
# +
# List Datasets
dataset_list = list(ws.datasets)
print(dataset_list)
# Get dataset by name
loan_ds = Dataset.get_by_name(ws, "Loan Applications Using SDK")
print(loan_ds.name)
# Get dataset by keys
ds_list = list(ws.datasets.keys())
for items in ds_list:
print(items)
| Basic/list_workspace_objects.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# +
import numpy as np
from sklearn.model_selection import train_test_split
import torch
from torch.autograd import Variable
from torch import nn
from torch import optim
from torch.utils.data import TensorDataset, DataLoader
import torch.nn.functional as F
from livelossplot import PlotLosses
# +
base_string = "()()" + (32 - 4) * " "
def shuffle_string(s):
indices = np.arange(len(s), dtype='uint8')
np.random.shuffle(indices)
return "".join(base_string[i] for i in indices)
def is_correct(seq):
open_brackets = 0
val = {"(": 1, " ": 0, ")": -1}
for c in seq:
open_brackets += val[c]
if open_brackets < 0:
return False
return open_brackets == 0
char2id = {" ": 0, "(": 1, ")": 2}
def generate_pairs(size):
X = np.zeros((size, 3, len(base_string)), dtype='float32')
Y = np.zeros((size), dtype='int64')
for i in range(size):
s = shuffle_string(base_string)
Y[i] = int(is_correct(s))
for j, c in enumerate(s):
X[i, char2id[c], j] = 1.
return X, Y
def generate_train_test_pairs(size):
X, Y = generate_pairs(size)
return train_test_split(X, Y, test_size=0.25, random_state=42)
# -
x = shuffle_string(base_string)
print(x, "- correct" if is_correct(x) else "- incorrect")
# +
X_train, X_test, Y_train, Y_test = generate_train_test_pairs(1000)
trainloader = DataLoader(TensorDataset(torch.from_numpy(X_train), torch.from_numpy(Y_train)),
batch_size=32, shuffle=True)
testloader = DataLoader(TensorDataset(torch.from_numpy(X_test), torch.from_numpy(Y_test)),
batch_size=32, shuffle=False)
# -
class Recurrent(nn.Module):
def __init__(self, rnn_size):
super(Recurrent, self).__init__()
self.lstm = nn.GRU(input_size=3, hidden_size=rnn_size)
self.fc = nn.Linear(rnn_size, 2)
def forward(self, x):
x = x.permute(2, 0, 1)
output, _ = self.lstm(x)
last_output = output[-1, :, :]
res = self.fc(last_output)
return res
net = Recurrent(8)
criterion = nn.modules.CrossEntropyLoss(size_average=False)
optimizer = optim.Adam(net.parameters(), lr=0.003)
# +
liveloss = PlotLosses()
for epoch in range(30):
epoch_loss = 0.0
epoch_correct = 0
epoch_loss_val = 0.0
epoch_correct_val = 0
net.train()
for inputs, labels in trainloader:
inputs, labels = Variable(inputs), Variable(labels)
optimizer.zero_grad()
outputs = net(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
epoch_loss += loss.data[0]
epoch_correct += (outputs.max(1)[1] == labels).sum().data[0]
avg_loss = epoch_loss / len(trainloader.dataset)
avg_accuracy = epoch_correct / len(trainloader.dataset)
net.eval()
for inputs, labels in testloader:
inputs, labels = Variable(inputs), Variable(labels)
outputs = net(inputs)
loss = criterion(outputs, labels)
epoch_loss_val += loss.data[0]
epoch_correct_val += (outputs.max(1)[1] == labels).sum().data[0]
avg_loss_val = epoch_loss_val / len(testloader.dataset)
avg_accuracy_val = epoch_correct_val / len(testloader.dataset)
liveloss.update({
'log loss': avg_loss,
'val_log loss': avg_loss_val,
'accuracy': avg_accuracy,
'val_accuracy': avg_accuracy_val
})
liveloss.draw()
# -
| examples/pytorch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 算法 10.1(观测序列的生成)
# - 使用p175,10.2.1 直接计算法
# - 测试数据1:测试数据:p177,例10.2(主要的测试数据)
# - 测试数据2:测试数据:p173,例10.1(盒子和球模型)(用来与前向和后向算法做对照可以相互印证,结果是否正确。另外,由于p177,例10.2中,a的状态的数量、b的状态的数量和观测序列的长度是对齐的,都是3,难以辨别对应的参数是否用对了,所以p173,例10.1没有对齐的数据可以用来检验一下。)
# - 测试结果1:与书中结果一致:0.13018四舍五入为0.13022
# +
import math
class MarkovDirectly(object):
def probability(self, a, b, pi, output):
an = len(a[0]) # a的状态的数量
bn = len(b[0]) # b的状态的数量
m = len(output) # 观测序列的长度
inputs = []
total_count = int(math.pow(an, m))
print('total_count: %s' % total_count)
result = 0
# 【p175,公式10.13】整个for循环就是实现这一个公式
for i in range(total_count): # i表示总的循环数
index_li = MarkovDirectly.get_sequence_index(i, an, m)
print('q_t的集合:%s' % index_li, end='。') # 即本次循环的状态集合,即本次循环的选中的盒子的集合
prob = pi[index_li[0]] # pi(i1)
for k in range(m - 1):
prob *= b[index_li[k]][output[k]] # b_i1(o1), b_i2(o2)
print('i%s->i%s' % (k, k+1), end=' ')
prob *= a[index_li[k]][index_li[k+1]] # a_i1_i2, a_i2_i3
prob *= b[index_li[m - 1]][output[m - 1]] # b_i3(o3)
print(' %.5f' % prob)
result += prob
return result
@staticmethod
def get_sequence_index(num, an, m):
if num:
index_li = []
while num:
quotient, remainder = divmod(num, an)
num = quotient
index_li.append(remainder)
if len(index_li) < m:
index_li.extend([0]*(m - len(index_li)))
return index_li
else:
return [0]*m
# +
# 测试数据1:测试数据:p177,例10.2(主要的测试数据)
a = [
[0.5, 0.2, 0.3],
[0.3, 0.5, 0.2],
[0.2, 0.3, 0.5]
]
b = [
[0.5, 0.5],
[0.4, 0.6],
[0.7, 0.3]
]
pi = [0.2, 0.4, 0.4]
output = [0, 1, 0]
mf = MarkovDirectly()
print(mf.probability(a, b, pi, output))
# +
# 测试数据2:测试数据:p173,例10.1(盒子和球模型)
a = [
[0, 1, 0, 0],
[0.4, 0, 0.6, 0],
[0, 0.4, 0, 0.6],
[0, 0, 0.5, 0.5]
]
b = [
[0.5, 0.5],
[0.3, 0.7],
[0.6, 0.4],
[0.8, 0.2]
]
pi = (0.25, 0.25, 0.25, 0.25)
output = [0, 0, 1, 1, 0]
mf = MarkovDirectly()
print(mf.probability(a, b, pi, output))
# -
| statistical_learning_method/algorithm 10.1(generation of observation sequence).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 101 NumPy Exercises for Data Analysis (Python)
# #### Questions by <NAME>
# The goal of the numpy exercises is to serve as a reference as well as to get you to apply numpy beyond the basics. The questions are of 4 levels of difficulties with L1 being the easiest to L4 being the hardest.
# ### Difficulty Level: L1
# Q.1 Import numpy as np and print the version number.
import numpy as np
np.__version__
# +
# Q.2 Create a 1D array of numbers from 0 to 9
np.arange(0,10,1)
# ans np.arange(10)
# -
# Q.3 Create a 3×3 numpy array of all True’s
np.full((3,3), True, dtype=bool)
# +
# Q.4 Extract all odd numbers from arr
arr = np.array([0, 1, 2, 3, 4, 5, 6, 7, 8, 9])
arr[arr % 2 == 1]
# +
# Q.5 Replace all odd numbers in arr with -1
arr = np.array([0, 1, 2, 3, 4, 5, 6, 7, 8, 9])
arr[arr % 2 == 1 ] = -1
arr
# +
# Q.6 Replace all odd numbers in arr with -1 without changing arr
arr = np.array([0, 1, 2, 3, 4, 5, 6, 7, 8, 9])
out = np.where((arr % 2 == 1),-1,arr)
print(out)
arr
# +
# Q.7 Convert a 1D array to a 2D array with 2 rows
np.arange(10).reshape(2,-1) # Setting to -1 automatically decides the number of cols
# -
# ### Difficulty Level: L2
# Q. Stack the arrays a and b horizontally.
a = np.arange(10).reshape(2,-1)
b = np.repeat(1, 10).reshape(2,-1)
np.hstack([a,b])
# +
#Q. Create the following pattern without hardcoding. Use only numpy functions and the below input array a.
a = np.array([1,2,3])
np.r_[np.repeat(a, 3), np.tile(a, 3)]
# np.r_ (is used to concatenate any number of array slice along row (first) axis)
# +
#Q. Get the common items between a and b
a = np.array([1,2,3,2,3,4,3,4,5,6])
b = np.array([7,2,10,2,7,4,9,4,9,8])
np.intersect1d(a,b)
# +
# Q. From array a remove all items present in array b
a = np.array([1,2,3,4,5])
b = np.array([5,6,7,8,9])
np.setdiff1d(a,b) # np.setdiff1d From 'a' remove all of 'b'
# +
# Q. Get the positions where elements of a and b match
a = np.array([1,2,3,2,3,4,3,4,5,6])
b = np.array([7,2,10,2,7,4,9,4,9,8])
np.where(a == b)
# +
# Q. Get all items between 5 and 10 from a.
a = np.array([2, 6, 1, 9, 10, 3, 27])
index = np.where((a >=5) & (a <= 10))
print(a[index])
# or
index2 = np.where(np.logical_and(a>=5, a<=10))
print(a[index2])
# or
a[(a >=5) & (a <= 10)]
# +
# Q.15 Convert the function maxx that works on two scalars, to work on two arrays.
# def maxx(x, y):
# """Get the maximum of two items"""
# if [x] >= [y]:
# return [x]
# else:
# return [y]
# maxx([1, 5],[2, 6])
# -
#Q.16 Swap columns 1 and 2 in the array arr.
arr = np.arange(9).reshape(3,3)
print(arr)
arr[:, [1,0,2]]
# +
# Q.17 Swap rows 1 and 2 in the array arr:
arr = np.arange(9).reshape(3,3)
print(arr)
print(arr[[1,0,2],:])
# +
# Q.18 Reverse the rows of a 2D array arr.
arr = np.arange(9).reshape(3,3)
print(arr)
print(arr[::-1])
# -
#Q.19 Reverse the columns of a 2D array arr.
arr = np.arange(9).reshape(3,3)
print(arr)
print(arr[:,::-1])
# +
#Q.20 Create a 2D array of shape 5x3 to contain random decimal numbers between 5 and 10.
np.random.uniform(5,10, size=(5,3))
# -
# ### Difficulty Level: L1
#Q.21 Print or show only 3 decimal places of the numpy array rand_arr.
rand_arr = np.random.random((5,3))
np.set_printoptions(precision=3)
print(rand_arr[:4])
# OR
np.round(rand_arr,3)
# +
# Q.22 Pretty print rand_arr by suppressing the scientific notation (like 1e10)
# Create the random array
np.random.seed(100)
rand_arr = np.random.random([3,3])/1e3
# rand_arr
np.set_printoptions(suppress=True, precision=6)
rand_arr
# -
#Q.23 Limit the number of items printed in python numpy array a to a maximum of 6 elements.
a = np.arange(15)
np.set_printoptions(threshold=6)
a
#Q.24 Print the full numpy array a without truncating.
np.set_printoptions(threshold=6)
a = np.arange(15)
a
np.set_printoptions(threshold=np.sys.maxsize)
a
# ### Difficulty Level: L2
# +
# Q.25 Import the iris dataset keeping the text intact.
iris = np.genfromtxt('iris.data', delimiter=',', dtype='object')
names = ('sepallength', 'sepalwidth', 'petallength', 'petalwidth', 'species')
iris[:3]
# +
# Q.26 Extract the text column species from the 1D iris imported in previous question.
iris_1d = np.genfromtxt('iris.data', delimiter=',', dtype=object)
print(iris_1d.shape)
species = np.array([row[4] for row in iris_1d])
species[:5]
# -
#Q.27 Convert the 1D iris to 2D array iris_2d by omitting the species text field.
iris_2d = np.genfromtxt('iris.data', delimiter=',', dtype='float', usecols=(0,1,2,3))
iris_2d[:5]
# +
# Q.28 Find the mean, median, standard deviation of iris's sepallength (1st column)
sepallength = np.genfromtxt('iris.data', delimiter=',', dtype='float', usecols=[0])
mean = np.mean(sepallength)
median = np.median(sepallength)
std = np.std(sepallength)
print(mean)
print(median)
print(std)
# -
# Q.29 Create a normalized form of iris's sepallength whose values range exactly between
# 0 and 1 so that the minimum has value 0 and maximum has value 1.
Smax, Smin = sepallength.max(), sepallength.min()
S = (sepallength - Smin)/(Smax - Smin)
print(S)
# OR
s = (sepallength - Smin)/sepallength.ptp()
print(s)
# # Difficulty Level: L3
# Q.30 Compute the softmax score of sepallength.
| 101 NumPy Exercises for Data Analysis (Python).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
import os
from tqdm import tqdm
import urllib.request as req
SOURCE_URL = "https://download.microsoft.com/download/3/E/1/3E1C3F21-ECDB-4869-8368-6DEBA77B919F/kagglecatsanddogs_3367a.zip"
# +
DESTINATION = "data"
os.makedirs(DESTINATION, exist_ok=True)
data_file = "data.zip"
DESTINATION_ZIP_PATH = os.path.join(DESTINATION, data_file)
# -
filename, headers = req.urlretrieve(SOURCE_URL, DESTINATION_ZIP_PATH)
print(headers)
| research_env/CNN-Example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="zX4Kg8DUTKWO"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="Gqi4N0TcSLf8"
# **Note:** This notebook can run using TensorFlow 2.5.0
# + id="z1zuX6UDSLf9"
# #!pip install tensorflow==2.5.0
# + id="P-AhVYeBWgQ3" outputId="5ea3155a-f2f5-4d45-f58e-08ec8c0a3680" colab={"base_uri": "https://localhost:8080/"}
import tensorflow as tf
print(tf.__version__)
# # !pip install -q tensorflow-datasets
# + id="_IoM4VFxWpMR" outputId="6d37bac9-4c48-4c70-9d30-4eded3769ee1" colab={"base_uri": "https://localhost:8080/", "height": 343, "referenced_widgets": ["838e78f879b74e9cb0537bbeaf1639f1", "9b75b4863a1d43c4a7633af9e2d80226", "<KEY>", "<KEY>", "88fa9c52274c45cf8d333b4666a1fe96", "c089a196e06d4a698b85481aedd402b6", "e630949e1d894d43954711ec5894cb47", "<KEY>", "<KEY>", "<KEY>", "3315b1d33087438e843dd43788fb4e6e", "<KEY>", "0fe4e8744a2148298133704a50250ee7", "<KEY>", "<KEY>", "62184302384d49629c6a53e737d230d1", "<KEY>", "077d53f5dde24027b56c41a996a436c0", "<KEY>", "b273e57c0a5740c0af91a9bf7900017c", "4dbcdd61291848b98ea9b40e096db1bf", "77e3fc8d71934b05ac1894b2e89a48ad", "98fb2e607d034adb868a92b9249de036", "cd5a15cddbea453fa85affcb5a57120b", "<KEY>", "<KEY>", "<KEY>", "ba624adad3664e09b4020526b18b40b1", "284e9965220745499d7f976fe4dc438f", "<KEY>", "<KEY>", "ad25952cff434d2d82a8021483ce656a", "<KEY>", "470dbe3654bc4a879fcbebee6145029a", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "ad45d13fe77540f29e244862de9abc1a", "<KEY>", "97f10da6607d4866949b4b96da597860", "69e0345582f4434ebef29c1a63a5f60c", "<KEY>", "<KEY>", "<KEY>", "e8a8c758507c4562a09059d2e74e62d9", "<KEY>", "2eff4557f4214b7ea4eb6d02d349a707", "fba4f4e65d564a8f92024af2dc9e1aa1", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "eccbe5ce50b54b799ae674df2c95c84b", "437bd212b381402285a9a887c0d4e2db", "<KEY>", "b6dccd1cea014dc5ac55f79209ad6849", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "0d890ea282c746609fafd30a744455fc", "d2ab795608f548b3926ea73b30eae720", "<KEY>", "7cfd0ff4605d4bafaf96f2fd12306712", "<KEY>", "<KEY>", "c85e331a45b54561a83df64e700ed5fa", "e44963318db44384a8a952791eac6112", "467b546afd8d419a84c1e3eccf7a1285", "71fac5318edf4c3981eeb1d130b0aa4e", "5506c5a73d334385aad1e84fa6ad3e11", "d1fce9c64c6241ac982123098bd06dc6", "cf59fdac580640e2968e61255176a74d", "7b7e2adef63549b792928e0d774def90", "14ffe3760d93409bb27d172544fc97eb", "763f3723f2ee4eeca77c221e8cbca2fc", "<KEY>", "<KEY>", "<KEY>", "e4c7f447547b4d66a5753cadd78f4b83", "54734e8617c64959b2241e6f057afd9c", "62a5f10447a94125b74feea0bff08a6c", "cfb4053245f64eaca34c40573e8a09b3", "<KEY>", "<KEY>", "<KEY>", "<KEY>"]}
import tensorflow_datasets as tfds
imdb, info = tfds.load("imdb_reviews", with_info=True, as_supervised=True)
# + id="wHQ2Ko0zl7M4"
import numpy as np
train_data, test_data = imdb['train'], imdb['test']
training_sentences = []
training_labels = []
testing_sentences = []
testing_labels = []
# str(s.tonumpy()) is needed in Python3 instead of just s.numpy()
for s,l in train_data:
training_sentences.append(s.numpy().decode('utf8'))
training_labels.append(l.numpy())
for s,l in test_data:
testing_sentences.append(s.numpy().decode('utf8'))
testing_labels.append(l.numpy())
training_labels_final = np.array(training_labels)
testing_labels_final = np.array(testing_labels)
# + id="7n15yyMdmoH1"
vocab_size = 10000
embedding_dim = 16
max_length = 120
trunc_type='post'
oov_tok = "<OOV>"
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
tokenizer = Tokenizer(num_words = vocab_size, oov_token=oov_tok)
tokenizer.fit_on_texts(training_sentences)
word_index = tokenizer.word_index
sequences = tokenizer.texts_to_sequences(training_sentences)
padded = pad_sequences(sequences,maxlen=max_length, truncating=trunc_type)
testing_sequences = tokenizer.texts_to_sequences(testing_sentences)
testing_padded = pad_sequences(testing_sequences,maxlen=max_length)
# + id="9axf0uIXVMhO" outputId="0df29bfd-057b-487c-8c15-40d2175af6d4" colab={"base_uri": "https://localhost:8080/"}
reverse_word_index = dict([(value, key) for (key, value) in word_index.items()])
def decode_review(text):
return ' '.join([reverse_word_index.get(i, '?') for i in text])
print(decode_review(padded[3]))
print(training_sentences[3])
# + id="5NEpdhb8AxID" outputId="62400e65-a7f9-4163-9f92-072af1b85c5f" colab={"base_uri": "https://localhost:8080/"}
model = tf.keras.Sequential([
tf.keras.layers.Embedding(vocab_size, embedding_dim, input_length=max_length),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(6, activation='relu'),
tf.keras.layers.Dense(1, activation='sigmoid')
])
model.compile(loss='binary_crossentropy',optimizer='adam',metrics=['accuracy'])
model.summary()
# + id="V5LLrXC-uNX6" outputId="b70f9390-a00c-4316-c438-1d9ca56884f2" colab={"base_uri": "https://localhost:8080/"}
num_epochs = 10
model.fit(padded, training_labels_final, epochs=num_epochs, validation_data=(testing_padded, testing_labels_final))
# + id="yAmjJqEyCOF_" outputId="21767665-6b24-4cc2-cb8c-c817c130d3cc" colab={"base_uri": "https://localhost:8080/"}
e = model.layers[0]
weights = e.get_weights()[0]
print(weights.shape) # shape: (vocab_size, embedding_dim)
# + id="jmB0Uxk0ycP6"
import io
out_v = io.open('vecs.tsv', 'w', encoding='utf-8')
out_m = io.open('meta.tsv', 'w', encoding='utf-8')
for word_num in range(1, vocab_size):
word = reverse_word_index[word_num]
embeddings = weights[word_num]
out_m.write(word + "\n")
out_v.write('\t'.join([str(x) for x in embeddings]) + "\n")
out_v.close()
out_m.close()
# + id="VDeqpOCVydtq" outputId="d6d61247-306e-4894-d211-877ea89c39fd" colab={"base_uri": "https://localhost:8080/", "height": 17}
try:
from google.colab import files
except ImportError:
pass
else:
files.download('vecs.tsv')
files.download('meta.tsv')
# + id="YRxoxc2apscY"
sentence = "I really think this is amazing. honest."
sequence = tokenizer.texts_to_sequences([sentence])
print(sequence)
# + id="lDHphRogSLgB"
| C3/W2/ungraded_labs/C3_W2_Lab_1_imdb.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
import numpy as np
import pandas as pd
import os
import matplotlib.pyplot as plt
import seaborn as sns
import tensorflow as tf
# -
# # 1. Upload Data
# +
data = pd.read_csv("fer2013.csv")
data.head()
# -
# # 2.Let's examine the dataset
data.shape
data['Usage'].value_counts()
# +
data_list = data['Usage'].value_counts()
groups = ['Training', 'PublicTest', 'PrivateTest']
colors = sns.color_palette('pastel')[0:5]
plt.pie(data_list, labels = groups, colors = colors, autopct='%.0f%%')
plt.show()
# +
# Emotion Counts
training = data.loc[data["Usage"] == "Training"]
emotion_map = {0:"Angry", 1:"Disgust", 2:"Fear", 3:"Happy", 4:"Sad", 5:"Surprise", 6:"Neutral"}
emotion_counts = training['emotion'].value_counts(sort=False).reset_index()
emotion_counts.columns = ['emotion', 'number']
emotion_counts['emotion'] = emotion_counts['emotion'].map(emotion_map)
plt.figure(figsize=(6,4))
sns.barplot(emotion_counts.emotion, emotion_counts.number)
plt.title('Class distribution')
plt.ylabel('Number', fontsize=12)
plt.xlabel('Emotions', fontsize=12)
plt.show()
# +
public_test = data.loc[data["Usage"] == "PublicTest"]
emotion_map = {0:"Angry", 1:"Disgust", 2:"Fear", 3:"Happy", 4:"Sad", 5:"Surprise", 6:"Neutral"}
emotion_counts = public_test['emotion'].value_counts(sort=False).reset_index()
emotion_counts.columns = ['emotion', 'number']
emotion_counts['emotion'] = emotion_counts['emotion'].map(emotion_map)
plt.figure(figsize=(6,4))
sns.barplot(emotion_counts.emotion, emotion_counts.number)
plt.title('Class distribution')
plt.ylabel('Number', fontsize=12)
plt.xlabel('Emotions', fontsize=12)
plt.show()
# +
emotion_map = {0:"Angry", 1:"Disgust", 2:"Fear", 3:"Happy", 4:"Sad", 5:"Surprise", 6:"Neutral"}
emotion_counts = training['emotion'].value_counts(sort=False).reset_index()
emotion_counts.columns = ['emotion', 'number']
emotion_counts['emotion'] = emotion_counts['emotion'].map(emotion_map)
plt.figure(figsize=(6,4))
sns.barplot(emotion_counts.emotion, emotion_counts.number)
plt.title('Class distribution')
plt.ylabel('Number', fontsize=12)
plt.xlabel('Emotions', fontsize=12)
plt.show()
# +
# Let's look at the first 4 pictures
from PIL import Image
from IPython.display import display
height = int(np.sqrt(len(data.pixels[0].split())))
width = int(height)
depth = 1
labels = ["Angry", "Disgust", "Fear", "Happy", "Sad", "Surprise", "Neutral"]
for i in range(0, 4):
matrix_array = np.mat(data.pixels[i]).reshape(height, width)
img = Image.fromarray(matrix_array.astype(np.uint8))
display(img)
print(labels[data.emotion[i]])
# -
# # 3. Data processing
# +
from tensorflow.keras.utils import to_categorical
training = data.loc[data["Usage"] == "Training"]
public_test = data.loc[data["Usage"] == "PublicTest"]
private_test = data.loc[data["Usage"] == "PrivateTest"]
# Converts a label vector (integers) to binary class matrix.
train_labels = training["emotion"]
train_labels = to_categorical(train_labels)
# train_labels.shape: (28709, 7)
train_pixels = training["pixels"].str.split(" ").tolist()
train_pixels = np.uint8(train_pixels)
# train_pixels.shape (28709, 48*48
train_pixels = train_pixels.reshape((28709, 48, 48, 1))
# normalization
train_pixels = train_pixels.astype("float32") / 255
private_labels = private_test["emotion"]
private_labels = to_categorical(private_labels)
private_pixels = private_test["pixels"].str.split(" ").tolist()
private_pixels = np.uint8(private_pixels)
private_pixels = private_pixels.reshape((3589, 48, 48, 1))
private_pixels = private_pixels.astype("float32") / 255
public_labels = public_test["emotion"]
public_labels = to_categorical(public_labels)
public_pixels = public_test["pixels"].str.split(" ").tolist()
public_pixels = np.uint8(public_pixels)
public_pixels = public_pixels.reshape((3589, 48, 48, 1))
public_pixels = public_pixels.astype("float32") / 255
# -
# # 4. Create Model
model = tf.keras.models.Sequential([
tf.keras.layers.Conv2D(32, (3,3), activation='relu', input_shape=(48,48,1)),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.Conv2D(32, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D(2, 2),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Conv2D(64, (3,3), activation='relu'),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.MaxPooling2D(2, 2),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Conv2D(128, (3,3), activation='relu'),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.MaxPooling2D(2, 2),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(256, activation='relu'),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(7, activation='softmax'),
])
model.summary()
model.compile(optimizer="Adam", loss="categorical_crossentropy", metrics="accuracy")
history = model.fit(train_pixels, train_labels, validation_data=(private_pixels, private_labels),
epochs=20, batch_size = 128)
# # 4. Model Evaluation
# +
train_acc = history.history["accuracy"]
validation_acc = history.history["val_accuracy"]
loss = history.history["loss"]
val_loss = history.history["val_loss"]
epochs = range(len(train_acc))
plt.plot(epochs, train_acc, 'bo', label="Training Accuracy")
plt.plot(epochs, validation_acc, 'b', label="Validation Accuracy")
plt.title("Train and Validation Accuracy")
plt.legend()
plt.figure()
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
| facial-expression-identification-using-cnn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: nlp
# language: python
# name: nlp
# ---
# # News Headlines Sentiment
#
# Use the news api to pull the latest news articles for bitcoin and ethereum and create a DataFrame of sentiment scores for each coin.
#
# Use descriptive statistics to answer the following questions:
# 1. Which coin had the highest mean positive score?
# 2. Which coin had the highest negative score?
# 3. Which coin had the highest positive score?
# +
# Initial imports
import os
import pandas as pd
from dotenv import load_dotenv
from newsapi import NewsApiClient
#import nltk
#nltk.downloader.download('vader_lexicon')
from nltk.sentiment.vader import SentimentIntensityAnalyzer
analyzer = SentimentIntensityAnalyzer()
# %matplotlib inline
# -
# Read your api key environment variable
# YOUR CODE HERE!
news_api_key = os.getenv("NEWS_API_KEY")
# Create a newsapi client
# YOUR CODE HERE!
newsapi = NewsApiClient(api_key=api_key)
# +
# Fetch the Bitcoin news articles
# YOUR CODE HERE!
bitcoin_headlines = newsapi.get_top_headlines(q="bitcoin", language="en", country="us")
print(f"Total articles about Bitcoin: {bitcoin_headlines['totalResults']}")
# +
# Fetch the Ethereum news articles
# YOUR CODE HERE!
eth_headlines = newsapi.get_top_headlines(q="ethereum", language="en", country="us")
print(f"Total articles about Ethereum: {eth_headlines['totalResults']}")
# +
# Create the Bitcoin sentiment scores DataFrame
# YOUR CODE HERE!
Bitcoin_sentiments = []
for article in bitcoin_headlines["articles"]:
try:
text = article["content"]
sentiment = analyzer.polarity_scores(text)
compound = sentiment["compound"]
pos = sentiment["pos"]
neu = sentiment["neu"]
neg = sentiment["neg"]
Bitcoin_sentiments.append({
"Compound": compound,
"Positive": pos,
"Negative": neg,
"Neutral": neu,
"Text": text
})
except AttributeError:
pass
# Create DataFrame
Bitcoin_df = pd.DataFrame(Bitcoin_sentiments)
# Reorder DataFrame columns
cols = ["Compound", "Positive", "Negative", "Neutral","Text"]
Bitcoin_df = Bitcoin_df[cols]
Bitcoin_df.head()
# +
# Create the ethereum sentiment scores DataFrame
# YOUR CODE HERE!
Eth_sentiments = []
for article in eth_headlines["articles"]:
try:
text = article["content"]
sentiment = analyzer.polarity_scores(text)
compound = sentiment["compound"]
pos = sentiment["pos"]
neu = sentiment["neu"]
neg = sentiment["neg"]
Ethereum_sentiments.append({
"Compound": compound,
"Positive": pos,
"Negative": neg,
"Neutral": neu,
"Text": text
})
except AttributeError:
pass
# Create DataFrame
Ethereum_df = pd.DataFrame(Ethereum_sentiments)
# Reorder DataFrame columns
cols = ["Compound", "Positive", "Negative", "Neutral","Text"]
Eth_df = Ethereum_df[cols]
Eth_df.head()
# +
# Describe the Bitcoin Sentiment
# YOUR CODE HERE!
Bitcoin_df.describe()
# +
# Describe the Ethereum Sentiment
# YOUR CODE HERE!
Eth_df.describe()
# -
# ### Questions:
#
# Q: Which coin had the highest mean positive score?
#
# A:
#
# Q: Which coin had the highest compound score?
#
# A:
#
# Q. Which coin had the highest positive score?
#
# A:
# ---
# # Tokenizer
#
# In this section, you will use NLTK and Python to tokenize the text for each coin. Be sure to:
# 1. Lowercase each word
# 2. Remove Punctuation
# 3. Remove Stopwords
from nltk.tokenize import word_tokenize, sent_tokenize
from nltk.corpus import stopwords
from nltk.stem import WordNetLemmatizer, PorterStemmer
from string import punctuation
import re
lemmatizer = WordNetLemmatizer()
# +
# Expand the default stopwords list if necessary
# YOUR CODE HERE!
# +
# Complete the tokenizer function
def tokenizer(text):
"""Tokenizes text."""
sw = set(stopwords.words('english'))
regex = re.compile("[^a-zA-Z ]")
re_clean = regex.sub('', article)
words = word_tokenize(re_clean)
lem = [lemmatizer.lemmatize(word) for word in words]
output = [word.lower() for word in lem if word.lower() not in sw]
return tokens
# +
# Create a new tokens column for bitcoin
# YOUR CODE HERE!
Bitcoin_df["tokens"] = Bitcoin_df.Text.apply(tokenizer)
Bitcoin_df.head()
# +
# Create a new tokens column for ethereum
# YOUR CODE HERE!
Eth_df["tokens"] = Eth_df.Text.apply(tokenizer)
Eth_df.head()
# -
# ---
# # NGrams and Frequency Analysis
#
# In this section you will look at the ngrams and word frequency for each coin.
#
# 1. Use NLTK to produce the n-grams for N = 2.
# 2. List the top 10 words for each coin.
from collections import Counter
from nltk import ngrams
#tokenized Bitcoin articles
bitcoin_p = tokenizer(Bitcoin_df.Text.str.cat())
bitcoin_p
# +
eth_p= tokenizer(Eth_df.Text.str.cat())
eth_p
# -
# Generate the Bitcoin N-grams where N=2
# YOUR CODE HERE!
N=2
bigram_counts_B = Counter(ngrams(bitcoin_p, N))
print(dict(bigram_counts_B))
# +
# Generate the Ethereum N-grams where N=2
# YOUR CODE HERE!
N= 2
bigram_counts_E = Counter(ngrams(Eth_p, N))
print(dict(bigram_counts_E))
# -
# Use the token_count function to generate the top 10 words from each coin
def token_count(tokens, N=10):
"""Returns the top N tokens from the frequency count"""
return Counter(tokens).most_common(N)
# +
# Get the top 10 words for Bitcoin
# YOUR CODE HERE!
bitcoin_top10= token_count(bitcoin_p, 10)
bitcoin_top10
# +
# Get the top 10 words for Ethereum
# YOUR CODE HERE!
eth_top10= token_count(Eth_p,10)
ethereum_top10
# -
# # Word Clouds
#
# In this section, you will generate word clouds for each coin to summarize the news for each coin
from wordcloud import WordCloud
import matplotlib.pyplot as plt
plt.style.use('seaborn-whitegrid')
import matplotlib as mpl
mpl.rcParams['figure.figsize'] = [20.0, 10.0]
# +
# Generate the Bitcoin word cloud
# YOUR CODE HERE!
cloud1 = WordCloud(background_color='blue').generate(Bitcoin_df.Text.str.cat())
cloud1
plt.axis("off")
plt.imshow(cloud1)
# +
# Generate the Ethereum word cloud
# YOUR CODE HERE!
cloud2 = WordCloud(background_color='white').generate(Eth_df.Text.str.cat())
cloud2
plt.axis("off")
plt.imshow(cloud2)
# -
# # Named Entity Recognition
#
# In this section, you will build a named entity recognition model for both coins and visualize the tags using SpaCy.
import spacy
from spacy import displacy
# +
# Optional - download a language model for SpaCy
# # !python -m spacy download en_core_web_sm
# -
# Load the spaCy model
nlp = spacy.load('en_core_web_sm')
# ## Bitcoin NER
# +
# Concatenate all of the bitcoin text together
# YOUR CODE HERE!
all_concat_bitcoin = Bitcoin_df.Text.str.cat()
all_concat_bitcoin
# +
# Run the NER processor on all of the text
# YOUR CODE HERE!
bitcoin_doc = nlp(all_concat_bitcoin)
bitcoin_doc
# Add a title to the document
bitcoin_doc.user_data["Title"] = "Bitcoin NER"
# +
# Render the visualization
# YOUR CODE HERE!
displacy.render(bitcoin_doc, style = 'ent')
# +
# List all Entities
# YOUR CODE HERE!
for i in bitcoin_doc.ents:
print(i.text, i.label_)
# -
# ---
# ## Ethereum NER
# +
# Concatenate all of the bitcoin text together
# YOUR CODE HERE!
all_concat_ethereum = Eth_df.Text.str.cat()
all_concat_ethereum
# +
# Run the NER processor on all of the text
# YOUR CODE HERE!
# Add a title to the document
ethereum_doc = nlp(all_concat_ethereum)
ethereum_doc
ethereum_doc.user_data["Title"] = "Ethereum NER"
# +
# Render the visualization
# YOUR CODE HERE!
displacy.render(ethereum_doc, style = 'ent')
# +
# List all Entities
# YOUR CODE HERE!
for i in ethereum_doc.ents:
print(i.text, i.label_)
| crypto_sentiment-Submission.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using PhysicalQuantites in Python
# %precision 2
from PhysicalQuantities import q
# ## The convenience class 'q' can be used to specify units:
q.mm, q.m, q.km
2 * q.mm**2
v = 30 * q.m / q.s
v.to('km/h')
p = 20*q.dBm + 3*q.dB
p, p.W
# ## List of defined units
q.table
| examples/pq-q-units.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## Number of synapses per connection
# In this notebook you will analyze and validate one of parameter of the connectome: the number of synapses per connection.
#
# Cell A is connected to cell B if the axon of one cell makes at least one synapse on the other cell (we do not consider gap junction).
#
# A connection has a direction, so between A and B we can identify two pathways. A->B means that axon from A creates the synapse(s) on B, while B->A means that axon from B creates the synapse(s) on A.
#
# Another important point is that a connection may have one or multiple synapses, which has an impact on the anatomy and physiology of the network.
#
# In this notebook, you will analyze the number of synapses per connection in all the possible pathways.
#
# ---
# Import some python packages.
# +
import itertools
import numpy as np
import pandas as pd
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn
from bluepysnap import Circuit
from bluepysnap.bbp import Cell
from pathlib import Path
# -
# Set the random seed
np.random.seed(42)
# Reading and preparing the data.
CIRCUIT_BASE = Path('/mnt/user/shared/Palermo workshop 2022 edition/circuit-O1')
circuit_path = CIRCUIT_BASE / 'circuit_config.json'
circuit = Circuit(circuit_path)
cells = circuit.nodes["hippocampus_neurons"]
conn = circuit.edges["hippocampus_neurons__hippocampus_neurons__chemical"]
# ### Analysis
#
# Initialize where to store the results.
#
# Since all the possible pathways form a 2D matrix, it is convenient to have one matrix where to store means and another one where to store standard deviations.
#
# Furthermore, you are going to compare the result with values extracted from literature. Among those values, you have the number of synapses per connection in parvalbumin positive (PV+) cells, a group that includes SP_PVBC, SP_BS, and SP_AA. The group PV is already defined in the circuit, but you have to include it in the matrices.
mtypes = cells.property_values(Cell.MTYPE)
model_mean = pd.DataFrame(index=mtypes, columns=mtypes.union({u'PV'}), dtype=float)
model_std = pd.DataFrame(index=mtypes, columns=mtypes.union({u'PV'}), dtype=float)
# The analysis could be quite expensive, so better to reduce the number of samples.
#
# Furthermore, since you will repeat the same analysis many times, it is convenient to create a helper function.
nsample = 100
def sample_nsyn(pre, post):
it = conn.iter_connections(pre, post, return_edge_count=True)
return np.array([p[2] for p in itertools.islice(it, nsample)])
# Here, you run the analysis.
#
# Note that the function sample_nsyn returns a 2D matrix with all the same connections for a given pathway.
#
# From this matrix, you will calculate the mean and standard deviation that end up in the result matrices. The current position in the analysis loop is indicated by the processed m-type.
for pre_mtype in mtypes:
for post_mtype in mtypes:
data = sample_nsyn(
pre={Cell.MTYPE: pre_mtype, Cell.REGION: {'$regex': 'mc2.*'}},
post={Cell.MTYPE: post_mtype}
)
if len(data) != 0:
model_mean[post_mtype][pre_mtype] = data.mean()
model_std[post_mtype][pre_mtype] = data.std()
print(pre_mtype)
model_mean
# Plot the result using an heatmap.
#
# Note that when you have a white cell, it means that the two cell types are not connected (at least in the sample tested).
# +
ax = seaborn.heatmap(model_mean)
fig = plt.gcf()
fig.suptitle('Number of synapses per connection', )
ax.set_xlabel('presynaptic mtype')
ax.set_ylabel('postsynaptic mtype')
ax.collections[0].colorbar.set_label("# synapses")
fig.show()
# -
# ### Validation
#
# After having analyzing the circuit, you can compare the model with experimental data extracted from literature.
#
# The next cell loads the experimental data and puts it in a pandas dataframe.
bio_path = CIRCUIT_BASE / 'bioname' / 'nsyn_per_connection_20190131.tsv'
df = pd.read_csv(bio_path, skiprows=1, names=['pre', 'post', 'bio_mean', 'bio_std'], usecols=[0, 1, 2, 3], delim_whitespace=True)
df.head()
# As you can see, there are only a limited number of pathways.
#
# Extract from the result matrices only the pathways for which you have experimental data.
df['mod_mean'] = np.NAN
df['mod_std'] = np.NAN
for idx in df.index:
pre = df.loc[idx, 'pre']
post = df.loc[idx, 'post']
df.loc[idx, 'mod_mean'] = model_mean[post][pre]
df.loc[idx, 'mod_std'] = model_std[post][pre]
# Now we plot the results. The more points are lying on the diagonal, the more the model is close to experimental values.
# +
plt.clf
x = df['mod_mean'].values
y = df['bio_mean'].values
# remove nan value from the array
l = np.linspace(0, max(x[~np.isnan(x)].max(), y.max()), 50)
# l = np.linspace(0, max(x.max(), y.max()), 50)
fig, ax = plt.subplots()
fig.suptitle('synapses per connection')
ax.plot(x, y, 'o')
ax.errorbar(x, y, xerr=df['mod_std'].values, yerr=df['bio_std'].values, fmt='o', ecolor='g', capthick=2)
ax.plot(l, l, 'k--')
ax.set_xlabel('Model (#)')
ax.set_ylabel('Experiment (#)')
fig.show()
# -
# ### Exercise #1
# Calculate the average number of synapses per connection in the four classes of connections (EE, EI, IE, II). Put the answers in a list called _ans\_1_ in the order (EE, EI, IE, II).
#
# ### Exercise #2
# Calculate the distribution of number of synapses from SP_PVBC to SP_PC. Provide a list for _ans\_2_, with the first element being the mean value and the second element being the standard deviation of the number.
# +
# Work here
| 4_nsyns_per_conn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Homework 05
# ### Exercise 1 - Terminology
#
# Describe the following terms with your own words:
#
# ***boolean array:*** An array only containing true/false or 0/1 data
#
# ***shape:*** shape tell about the dimensions and their length of an array
#
# ***axis:*** axis is another term for dimensions of an array. For instance, a point has zero dimensions/axis, a line has 1 dimension/axis and a matrix is described using 2 dimensions/axis
# Answer the following questions:
#
# ***Which ways are there to select one or more elements from a Numpy array?*** Indexing or fancy indexing. E.g. m = np.array([[1,2,3],
# [10,0,5]])
# m[1,1] gives then back the element 0
#
# ***What is the difference between Numpy and Scipy?*** Numpy is used for used for various operations of the data. Wheras Scipy has more advanced functions which are built on Numpy
#
# ### Exercise 2 - Download data from entsoe-e for Lecture 6
# For lecture 6, we need to download data from the Entso-e transparency platform: Entso-e provides (almost) real-time data on European electricity systems. We will download hourly load data (i.e. electricity demand) for all systems in Europe. First, you need to get a user account at Entsoe-e here.
#
# We are going to use the S-FTP server of Entso-e. To use S-FTP in Python, you have to install the package pysftp. You can do so here in the notebook by executing the following command (please be aware that this may take some time):
#
# !conda install -c conda-forge pysftp --yes
# Now we are ready to download the data. In principle, you simply have to fill out your account information (by setting USER and PWD), decide where to put the data locally by assigning a path to a DOWNLOAD_DIR and run the 4 cells below. If the download directory does not exist, it will be created. The download will take some time, so you may want to run the script overnight.
#
# If the download fails at some point, you can restart it by simply executing the cell again. Files which are already downloaded will not be downloaded again. Hint: I had problems downloading to a directoy which was on a google drive - so if you run into an error message, which says OSError: size mismatch in get! you may want to choose a directory which is not on a google drive or possibly a dropbox. Also, this error may occur if your disk is full.
#
# +
import os
import pysftp
# if you want, you can modify this too, per default it will create a folder
# in the parant folder of the homework repository:
DOWNLOAD_DIR = '../../entsoe-data'
CATEGORIES = [
'ActualTotalLoad'
]
# -
# To avoid storing the user credentials in the public Github repository,
# these commands will ask you to enter them interactively:
from getpass import getpass
user = getpass('User for ENTSO-E API:')
pwd = getpass('Password for ENTSO-E API:')
def download_entsoe_data(user, pwd, category, output_dir, server_uri='sftp-transparency.entsoe.eu'):
"""Download a dataset from ENTSO-E's transparency data sftp server.
Contact ENTSO-E to receive login credentials:
https://transparency.entsoe.eu/usrm/user/createPublicUser
:param user: user name required for connecting with sftp server
:param pwd: password required for connecting with sftp server
:param category: ENTSO-E data category to be downloaded
:param output_dir: directory where downloaded data is saved to, a separate
subdirectory is created for each category.
:param server_uri: URI of ENTSO-E transparency server (default last updated on 2020-05-01)
"""
abspath = os.path.abspath(output_dir)
# check if local_dir exists and create if it doesn't
if not os.path.exists(abspath):
os.mkdir(abspath)
print (f'Successfully created the directory {abspath} and using it for download')
else:
print (f'{abspath} exists and will be used for download')
print("\nCopy this path for other notebooks, e.g. the next lecture or homework:\n"
f"DOWNLOAD_DIR = '{abspath}'\n")
cnopts = pysftp.CnOpts()
cnopts.hostkeys = None
# connect to entsoe server via sFTP
entsoe_dir = f'/TP_export/{category}'
with pysftp.Connection(server_uri, username=user, password=<PASSWORD>, cnopts=cnopts) as sftp:
sftp.chdir(entsoe_dir)
files_entsoe = sftp.listdir()
to_download = list(files_entsoe)
print(f'In total, {len(to_download)} files are going to be downloaded')
# download files not on disk
for file in to_download:
print(f'Downloading file {file}...')
dest_file = os.path.join(abspath, file)
if not os.path.exists(dest_file):
temp_file = os.path.join(abspath, f'{file}.partial')
sftp.get(f'{entsoe_dir}/{file}', temp_file)
os.rename(temp_file, dest_file)
print(f'{file} downloaded successfully.')
else:
print(f'{file} already present locally, skipping download.')
sftp.close()
print("All downloads completed")
# download data...
for category in CATEGORIES:
download_entsoe_data(user, pwd, category, DOWNLOAD_DIR)
# **Privacy note:** If you don't want to publish the path to your repository on Github (it may contain your Windows user name for example), clear the output of the cell above before saving the Notebook! (In the menu via Cell -> Current outputs -> Clear.)
# ### Exercise 3 - Create a diagonal matrix
#
# Create a matrix `m` with shape `(4, 4)` by using `np.zeros()` and set the 4 diagonal elements to `1` by using indexing using `np.arange()`. Do not use more two assign statements in total for this exercise!
#
# Bonus: Find multiple ways to avoid calling `np.arange()` twice and analyze which is the best regarding readability, performance and memory usage!
#
# Note: Normally you would use `np.diag()` to do this. You can also have a look into the code using `np.diag??`, but it's probably easier to write your own implementation (which might be less generic and slower, but way simpler).
# +
import numpy as np
m = np.zeros((4,4))
print(f'The original matrix m = \n{m}')
m[np.arange(4),np.arange(4)]=1
print(f'\nThe matrix m with 1 = \n{m}')
##avoid calling np.arange twice
m2 = np.zeros((4,4))
i = np.arange(4)
m2[i,i] = 1
print(f'\nThe matrix m with 1 = \n{m}')
m3 = np.zeros((4,4))
d_ind = np.diag_indices(len(m3),ndim=2)
m3[d_ind] = 1
print("Diagonal matrix with np.diag = \n", m3)
# -
# ### Exercise 4 - Invasion
#
# Create a canvas using `np.zeros()` of shape `(8, 11)`. Then set the following elements to one using fancy slicing techniques:
#
# - Rows 4 and 5 completely.
# - In row 3 all elements except the first one.
# - In row 2 all elements except the first two ones.
# - The two elements defined by: `row_idcs, column_idcs = [0, 1], [2, 3]`
# - In row 6 the elements in column 0 and 2.
# - In row 7 all elements except the first three and the last three.
#
# And then afterwards the following elements to zero:
# - The three elements defined by: `row_idcs, column_idcs = [3, 5, 7], [3, 1, 5]`
#
# As a last step, set assign the content of the first five columns to the last five columns in reversed order. This can be done by using a `step=-1` and starting with 4, i.e. the first five columns in reversed order are indexed by `canvas[:, 4::-1]`.
#
# Then plot the canvas using `plt.imshow()` with the parameter `cmap='gray'`!
#
# **Hint:** it helps a lot to have all commands in one cell (including the `imshow()` command) and execute the cell often, to check the result.
#
# **Note:** When ever the instruction says "first element" it is something like `x[0]`, because it refers to the first one in the array. If it is column 1 or row 1 it is `x[1]`, because it refers then to the index of the column/row.
#
# **Note:** It is `canvas[row_index, column_index]`, so if you are thinking in x/y coordinates, it is `canvas[y, x]` and the y axis goes downwards.
# +
import matplotlib
import matplotlib.pyplot as plt
canvas = np.zeros((8,11))
canvas[[4,5],:] = 1
canvas[3,1:] = 1
canvas[2,2:] = 1
row_idcs, column_idcs = [0, 1], [2, 3]
canvas[row_idcs,column_idcs] = 1
canvas[6,[0,2]] = 1
canvas[7,3:8] = 1
print(canvas)
row_idcs, column_idcs = [3, 5, 7], [3, 1, 5]
canvas[row_idcs,column_idcs] = 0
print(f'\n {canvas}')
canvas[:,6:] = canvas[:, 4::-1]
print(f'\n {canvas}')
plt.imshow(canvas, cmap='gray')
# -
# ### Exercise 5 - Draw a circle
#
# Draw a full circle: first define a resolution e.g. $N=50$. Then define coordinates $x$ and $y$ using `np.linspace()` and pass the resolution as parameter `num=N`. Use `np.meshgrid()` to define a grid `xx` and `yy`. Define a canvas of shape `(N, N)` using `np.zeros()`. Then use the circle formula $x^2 + y^2 < r^2$ to define all circle points on the grid (use $r=2$). Then use the boolean 2D expression to set the inside of the circle to 1. Finally plot the canvas using `imshow()`.
# +
N= 5000 #the higer the N, the "finer" the circle
x = np.linspace(-5,5,N)
y = np.linspace(-5,5,N)
#print(x, y)
xx, yy = np.meshgrid(x,y)
print(xx[:3,:3])
print(yy[:3,:3])
canvas = np.zeros((N,N))
#define all circle points on the grid with r= 2 and circle formula: 𝑥²+𝑦² < 𝑟
all_points = xx**2 + yy**2
print(all_points[:3,:3])
is_on_circle = all_points < 2
canvas[is_on_circle] = 1
plt.imshow(canvas)
plt.xlabel("x")
plt.ylabel("y")
# -
# ### Exercise 6 - Frequency of shades of gray
#
# Convert the picture `numpy-meme.png` to gray scale and plot a histogram!
#
# **Instructions:** Load the image by using `plt.imread()`. This will return a three dimensional array (width, height and colors) with values between zero and one. Using the formula `gray = red * 0.2125 + green * 0.7154 + blue * 0.0721`, convert the picture to shades of gray. Look at the shape of the image and pick the right axis by looking at the length of the array in this axis! You can first calculate a weighted version of the array by multiplying with a vector of length 3 (and the three weights) and then sum along the right axis. Check the shape of the gray image afterwards and plot it using `plt.imshow()` with the parameter `cmap='gray'`. It should be only two dimensional now. Use `image_gray.flatten()` to get all pixels as one-dimensional vector and pass this to the function `plt.hist()` with the parameter `bins=50` to get 50 bins with different gray values.
# +
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
matplotlib.rc('figure', figsize=(15, 10))
im = plt.imread('numpy-meme.png')
print(im.shape)
#the third axis contains the colours
weight = [0.2125, 0.7154, 0.0721]
im_grey = np.sum(im * weight, axis = 2)
print(im_grey.shape)
plt.imshow(im_grey, cmap="gray")
# +
im_grey_flat = im_grey.flatten()
plt.hist(im_grey_flat, bins=50)
# -
# ### Exercise 7 - Count colors (optional)
#
# Calculate the number of colors used in the picture `numpy-meme.png` and the percentage of the color space (3 x 8bit, i.e. 256 values per color) used!
#
# **Instructions:** Load the image by using `plt.imread()`. This will return a three dimensional array (width, height and colors) with values between zero and one. Multiplying the array with 255 will restore the original 8bit values (integer values between 0 and 255). After multiplying by 255 use `image = image.astype(int)` to convert the image to integer type. Plot the `image` using `plt.imshow()` to see the image and guess the result. Check the shape of the array. One of the axes is of length three - this is the color axis (red, green and blue). We want to map all colors to unique integers. This can be done by defining `colors = red + green * 256 + blue * 256**2`. This is a unique mapping between the triples `(red, green, blue)` and the integers `color` similar to decimal digits (three values between 0 and 9 e.g. `(3, 5, 1)` can be mapped to a three digit number `3 + 5 * 10 + 1 * 100 = 153`). Then use `np.unique()` to get an array with unique colors (in the mapped form as in `color`). This can be used to determine the number of unique colors in the image. This value can also be used to calculate the percentage of the color space used.
#
# <small>Image source: https://me.me/i/1-import-numpy-1-import-numpy-as-np-there-is-e4a6fb9cf75b413dbb3154794fd3d603</small>
# Inspired by [this exercise](https://github.com/rougier/numpy-100/blob/master/100_Numpy_exercises_with_solutions.md#66-considering-a-wh3-image-of-dtypeubyte-compute-the-number-of-unique-colors-) (MIT licensed, [DOI](https://zenodo.org/badge/latestdoi/10173/rougier/numpy-100))
# +
im2 = im * 255
im2 = im2.astype(int)
plt.imshow(im2)
print("im2[:3,:3,:]",im2[:3,:3,:])
#guess 12%
print(f'\nim2.shape = {im2.shape}')
#colors = red + green * 256 + blue * 256**2
colors = [1,256,256**2]
im2_colors = np.sum(im2 * colors, axis = 2)
print(f'\n im2_colors.shape = {im2_colors.shape}')
print(f'\n im2_colors[:3,:3] = \n {im2_colors[:3,:3]}')
# #np.unique?
col_count = np.unique(im2_colors)
print(f'\n col_count.shape = {col_count.shape}')
print(f'\n col_count[:10] = {col_count[:10]}')
max_c = 256**3###I am not sure about the maximum numbers of colors that can appear in colorspace
print(f' \n max numbers of colors = {max_c}')
##calculate % of colorspace used
perc_colspace = len(col_count) / max_c
print(f' % of colourspace used: {round(perc_colspace*100,4)}')
| homework05-numpy-scipy/homework05-karnerk.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] _uuid="bab978a4d9bc483f0697223cca084e52213fdf00"
# # Case 2 - MLFLOW - Tunning Hiper-Parametros do Modelo Original
# ## <NAME>
# + [markdown] _uuid="9e67cd962fd4fb67f0daf0b1db26a91d22f788bc"
# <img style="float: left;" src="https://guardian.ng/wp-content/uploads/2016/08/Heart-diseases.jpg" width="350px"/>
# -
import warnings
warnings.filterwarnings('ignore')
# +
# #!pip install mlflow
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
import mlflow
import mlflow.pyfunc
import mlflow.sklearn
import numpy as np
import sklearn
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import roc_auc_score
from mlflow.models.signature import infer_signature
from mlflow.utils.environment import _mlflow_conda_env
from sklearn.model_selection import train_test_split
import cloudpickle
import time
np.random.seed(123) #ensure reproducibility
from sklearn.model_selection import GridSearchCV
# + [markdown] _uuid="87aec28b7dd55601a7363cb7b613907e98f24518"
# <a id='section2'></a>
# + [markdown] _uuid="78d63e79cfb6f48e78dab7c785e8e952a08d518c"
# # The Data
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
dt = pd.read_csv("../dados/heart.csv")
# + _uuid="f3caf3de0a7e6d4602b26a1e72bf42d42ef0aac0"
dt.columns = ['age', 'sex', 'chest_pain_type', 'resting_blood_pressure', 'cholesterol', 'fasting_blood_sugar', 'rest_ecg', 'max_heart_rate_achieved',
'exercise_induced_angina', 'st_depression', 'st_slope', 'num_major_vessels', 'thalassemia', 'target']
# + _uuid="755235c8db67e5d76ee2fdc5cd55390e60e61ee9"
dt['sex'][dt['sex'] == 0] = 'female'
dt['sex'][dt['sex'] == 1] = 'male'
dt['chest_pain_type'][dt['chest_pain_type'] == 1] = 'typical angina'
dt['chest_pain_type'][dt['chest_pain_type'] == 2] = 'atypical angina'
dt['chest_pain_type'][dt['chest_pain_type'] == 3] = 'non-anginal pain'
dt['chest_pain_type'][dt['chest_pain_type'] == 4] = 'asymptomatic'
dt['fasting_blood_sugar'][dt['fasting_blood_sugar'] == 0] = 'lower than 120mg/ml'
dt['fasting_blood_sugar'][dt['fasting_blood_sugar'] == 1] = 'greater than 120mg/ml'
dt['rest_ecg'][dt['rest_ecg'] == 0] = 'normal'
dt['rest_ecg'][dt['rest_ecg'] == 1] = 'ST-T wave abnormality'
dt['rest_ecg'][dt['rest_ecg'] == 2] = 'left ventricular hypertrophy'
dt['exercise_induced_angina'][dt['exercise_induced_angina'] == 0] = 'no'
dt['exercise_induced_angina'][dt['exercise_induced_angina'] == 1] = 'yes'
dt['st_slope'][dt['st_slope'] == 1] = 'upsloping'
dt['st_slope'][dt['st_slope'] == 2] = 'flat'
dt['st_slope'][dt['st_slope'] == 3] = 'downsloping'
dt['thalassemia'][dt['thalassemia'] == 1] = 'normal'
dt['thalassemia'][dt['thalassemia'] == 2] = 'fixed defect'
dt['thalassemia'][dt['thalassemia'] == 3] = 'reversable defect'
# + _uuid="12edd841e271a4f7c8c039aa73412c0d6d7e5dad"
dt['sex'] = dt['sex'].astype('object')
dt['chest_pain_type'] = dt['chest_pain_type'].astype('object')
dt['fasting_blood_sugar'] = dt['fasting_blood_sugar'].astype('object')
dt['rest_ecg'] = dt['rest_ecg'].astype('object')
dt['exercise_induced_angina'] = dt['exercise_induced_angina'].astype('object')
dt['st_slope'] = dt['st_slope'].astype('object')
dt['thalassemia'] = dt['thalassemia'].astype('object')
# + _uuid="b6ec4deb644854301fa463758df32d6171f1c615"
dt = pd.get_dummies(dt, drop_first=True)
# + [markdown] _uuid="58c7f30375a2ffb7e02763e249e441a12cd437f1"
# # Registro do Modelo em MLFLOW
# ## Tunning Model - Version 2 - Modelo Escolhido no HyperTunning
# + _uuid="315ebc70bfe105f4b224974415db867d3d1e6b66"
X_train, X_test, y_train, y_test = train_test_split(dt.drop('target', 1), dt['target'], test_size = .2, random_state=10)
# + [markdown] _uuid="24d613abb60bb713089e3474e23323260e70b64b"
# <a id='section4'></a>
# -
def rodarTunning(X_train, y_train, X_test, y_test, rf_classifier):
mlflow.sklearn.autolog()
param_grid = {'n_estimators': [50, 75, 100, 125, 150, 175],
'min_samples_split':[2,4,6,8,10],
'min_samples_leaf': [1, 2, 3, 4],
'max_depth': [5, 10, 15, 20, 25]}
metrics = ['f1', 'recall', 'precision', 'roc_auc', 'neg_log_loss', 'neg_brier_score',
'average_precision', 'balanced_accuracy']
grid_obj = GridSearchCV(rf_classifier,
return_train_score=True,
param_grid=param_grid,
scoring=metrics,
cv=10,
refit='f1')
grid_fit = grid_obj.fit(X_train, y_train)
rf_opt = grid_fit.best_estimator_
mlflow.sklearn.log_model(grid_obj.best_estimator_, "best model")
mlflow.log_metric('best score', grid_obj.best_score_)
for k in grid_obj.best_params_.keys():
mlflow.log_param(k, grid_obj.best_params_[k])
print("Model saved in run %s" % mlflow.active_run().info.run_uuid)
print('='*20)
print("best params: " + str(grid_obj.best_estimator_))
print("best params: " + str(grid_obj.best_params_))
print('best score:', grid_obj.best_score_)
print('='*20)
print(classification_report(y_test, rf_opt.predict(X_test)))
print('New Accuracy of Model on train set: {:.2f}'.format(rf_opt.score(X_train, y_train)*100))
print('New Accuracy of Model on test set: {:.2f}'.format(rf_opt.score(X_test, y_test)*100))
return rf_opt
rf_classifier = RandomForestClassifier(class_weight = "balanced", random_state=7)
rf_opt = rodarTunning(X_train, y_train, X_test, y_test, rf_classifier)
| Geral/fonte_tunning/Tunning_Version2_mlflow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.5 64-bit
# name: python3
# ---
# Teste função - variável local e global
def teste(v, i):
valor = v
incremento = i
resultado = valor + incremento
return resultado
# instanciando a função
a = teste(10,1)
#
a
# Esta linha dará erro - a variável "resultado" não está definida globalmente
resultado
# Classes e métodos - funções são métodos dentro das classes
# Podemos ter vários métodos dentro das classes
class EntendendoClasses:
def incrementa(self, v, i):
valor = v
incremento = i
resultado = valor + incremento
return resultado
# Instanciando a classe no objeto "a" ~ variável.
a = EntendendoClasses()
a
# Chamando a função(método) dentro das classes
# As variáveis dentro das funções são atributos dentro de uma função(método).
b = a.incrementa(10,1)
# Aplicação do método incrementa da classe EntendendoClasses
b
a = EntendendoClasses().incrementa(10,1)
a
# self ~ dele mesmo - cada variável associa a si mesmo a cada saída da classe através do objeto
class EntendendoClasses:
def incrementa(self, v, i):
self.valor = v
self.incremento = i
self.resultado = self.valor + self.incremento
return self.resultado
a = EntendendoClasses()
b = a.incrementa(10,1)
# valor também recebe os atributos de incementa
a.valor
a.incrementa(11,1)
a.valor
# instanciamento de classe
b = EntendendoClasses()
b
# Instanciamento da classe pelo objeto b e seus argumentos
b.incrementa(10,3)
b.valor
# Classe com o método (função) construtor - constroi, define os valores inicias dos atributos
class EntendendoClasses:
def __init__(self, v:int, i:int):
self.valor = v
self.incremento = i
def incrementa(self):
self.valor = self.valor + self.incremento # self.valor += selfincremento
a = EntendendoClasses(10, 1)
a.incrementa()
a.valor
a.incrementa()
a.valor
b = EntendendoClasses(10, 1)
b.incrementa()
b.valor
# Classe com o método (função) construtor - com valores padrão
class EntendendoClasses:
def __init__(self, v=10, i=1):
self.valor = v
self.incremento = i
def incrementa(self):
self.valor = self.valor + self.incremento
a = EntendendoClasses()
a.incrementa()
a.valor
# +
# Depois de instanciarmos a = EntendendoClasses(), quando executarmos a.incrementa, o que o inrerpretador faz é
# EntendendoClasses().incrementa(a, 10, 1)
# +
# Objeto -
# Métodos(comportamento): compartilhado com todos os objetos criados a partir da mesma classe
# Atributos(estado): Não compartilhados com outros objetos criados a partir da mesma classe
# -
# Classe com mais funções
class EntendendoClasses:
def __init__(self, v=10, i=1):
self.valor = v
self.incremento = i
self.valor_exponencial = v
def incrementa(self):
self.valor = self.valor + self.incremento
def verifica(self):
if self.valor > 12:
print('Ultrapassou 12')
else:
print('Não ultrapassou 12')
def exponencial(self, e):
self.valor_exponencial = self.valor**e
def incrementa_quadrado(self):
self.incrementa()
self.exponencial(2)
a = EntendendoClasses()
a.incrementa()
a.valor
a.verifica
a.exponencial(3)
a.valor_exponencial
a.incrementa_quadrado()
a.valor
a.valor_exponencial
b = EntendendoClasses(50, 5)
b.incrementa()
b.valor
# Herança
class Calculos(EntendendoClasses):
pass
c = Calculos()
c.incrementa_quadrado()
c.valor_exponencial
class Calculos(EntendendoClasses):
def decrementa(self):
self.valor = self.valor - self.incremento
c = Calculos()
c.incrementa()
c.valor
c.decrementa()
c.valor
class Calculos(EntendendoClasses):
def __init__(self, d=5):
supper().__init__(v=10, i=1)
self.divisor = d
def decrementa(self):
self.valor = self.valor - self.incremento
def divide(self):
self.valor = self.valor/self.divisor
c = Calculos()
c.incrementa()
c.valor
c.decrementa()
c.divide()
| python/projetos_dev/class_python/classesMetodosAtributos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
os.environ['CUDA_VISIBLE_DEVICES'] = '1'
# +
import sys
SOURCE_DIR = os.path.dirname(os.path.dirname(os.path.abspath(__name__)))
sys.path.insert(0, SOURCE_DIR)
# -
import tensorflow as tf
import malaya_speech
import malaya_speech.train
from malaya_speech.train.model import pix2pix
import numpy as np
inputs = tf.placeholder(tf.float32, (None, 256, 256, 1))
targets = tf.placeholder(tf.float32, (None, 256, 256, 1))
def define_generator(inputs):
return pix2pix.generator.get_generator(inputs, 1)
model = pix2pix.Model(define_generator, inputs, targets)
sess = model.sess
fetches = model.fetches
writer = tf.summary.FileWriter('./out')
saver = tf.train.Saver()
# +
import numpy as np
test_mel = np.random.uniform(size = (1, 256, 256, 1))
# -
for i in range(5):
results = sess.run(fetches, feed_dict = {inputs: test_mel, targets: test_mel})
print(results)
writer.add_summary(results['summary'], results['global_step'])
saver.save(sess, './out/model.ckpt', global_step = results['global_step'])
| test/test-pix2pix.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:universe]
# language: python
# name: conda-env-universe-py
# ---
# # TensorBoard
#
# <font size =3.0> TensorBoard is the tensorflow's visualization tool which can be used to visualize the
# computation graph. It can also be used to plot various quantitative metrics and results of
# several intermediate calculations. Using tensorboard, we can easily visualize complex
# models which would be useful for debugging and also sharing.
# Now let us build a basic computation graph and visualize that in tensorboard.
# <font size=3.0> First, let us import the library
import tensorflow as tf
#
# <font size=3.0> Next, we initialize the variables
a = tf.constant(5)
b = tf.constant(4)
c = tf.multiply(a,b)
d = tf.constant(2)
e = tf.constant(3)
f = tf.multiply(d,e)
g = tf.add(c,f)
#
# <font size=3.0> Now, we will create a tensorflow session, we will write the results of our graph to file
# called event file using tf.summary.FileWriter()
with tf.Session() as sess:
writer = tf.summary.FileWriter("logs", sess.graph)
print(sess.run(g))
writer.close()
# <font size=3.0> In order to run the tensorboard, go to your terminal, locate the working directory and
# type
#
# tensorboard --logdir=logs --port=6003
# # Adding Scope
#
# <font size=3.0> Scoping is used to reduce complexity and helps to better understand the model by
# grouping the related nodes together, For instance, in the above example, we can break
# down our graph into two different groups called computation and result. If you look at the
# previous example we can see that nodes, a to e perform the computation and node g
# calculate the result. So we can group them separately using the scope for easy
# understanding. Scoping can be created using tf.name_scope() function.
with tf.name_scope("Computation"):
a = tf.constant(5)
b = tf.constant(4)
c = tf.multiply(a,b)
d = tf.constant(2)
e = tf.constant(3)
f = tf.multiply(d,e)
# +
with tf.name_scope("Result"):
g = tf.add(c,f)
# -
# <font size=3.0>
# If you see the computation scope, we can further break down in to separate parts for even
# more good understanding. Say we can create scope as part 1 which has nodes a to c and
# scope as part 2 which has nodes d to e since part 1 and 2 are independent of each other.
with tf.name_scope("Computation"):
with tf.name_scope("Part1"):
a = tf.constant(5)
b = tf.constant(4)
c = tf.multiply(a,b)
with tf.name_scope("Part2"):
d = tf.constant(2)
e = tf.constant(3)
f = tf.multiply(d,e)
#
# <font size=3.0>
# Scoping can be better understood by visualizing them in the tensorboard. The complete
# code looks like as follows,
with tf.name_scope("Computation"):
with tf.name_scope("Part1"):
a = tf.constant(5)
b = tf.constant(4)
c = tf.multiply(a,b)
with tf.name_scope("Part2"):
d = tf.constant(2)
e = tf.constant(3)
f = tf.multiply(d,e)
with tf.name_scope("Result"):
g = tf.add(c,f)
with tf.Session() as sess:
writer = tf.summary.FileWriter("logs", sess.graph)
print(sess.run(g))
writer.close()
# <font size=3.0> In order to run the tensorboard, go to your terminal, locate the working directory and
# type
#
# tensorboard --logdir=logs --port=6003
# <font size=3.5> If you look at the TensorBoard you can easily understand how scoping helps us to reduce
# complexity in understanding by grouping the similar nodes together. Scoping is widely
# used while working on a complex projects to better understand the functionality and
# dependencies of nodes.
| Chapter02/.ipynb_checkpoints/TensorBoard-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ASEM000/Physics-informed-neural-network-in-JAX/blob/main/%5B4%5D_ODE_Supervised_and_PINN.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="8mIeFcVvykBV"
# Credits : <NAME> @Asem000 Septemeber 2021
#
# Problem motivation credits : [<NAME>](https://chrisrackauckas.com) from [MIT 18.837](https://mitmath.github.io/18337/lecture3/sciml.html)
#
# + [markdown] id="FCZXu1J9u60x"
# ### Supervised learning with PINN
#
# the aim of this notebook is to demonstrated how can we use assumed/approximated physics loss along with supervised loss to better predict an arbitrary model given limited measurements.
#
# We measure the force at locations x_i of of a **real** spring system and use the x_i->F(x_i) pairs for supervised training . We later **assume** that the real spring follows hookes law and encode the **ideal** spring law in the physics loss formulation with the superivised loss .
#
# We compare the real spring (x,F(x)) against both strictly supervised loss and supervised loss with ideal spring physics loss
#
# <img src="https://i.imgur.com/A2fwUNg.png" width=50%>
#
# <img src="https://i.imgur.com/wwpNTTi.png" width=50%>
# + colab={"base_uri": "https://localhost:8080/"} id="vAR0swbLX_ZI" outputId="00ddea2c-105c-49e8-ec30-b22b11b99f5a"
#Imports
import jax
import jax.numpy as jnp
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import cm
import matplotlib as mpl
# !pip install optax
import optax
# !pip install numba
import numba
import sympy as sp
# + [markdown] id="7bg4nSbsXVwD"
# ### Mass-Spring-Damper Problem
# + [markdown] id="DGSu3EJwxDf7"
# We construct the following mass spring damper problem . Then we select about 7 points as our measurements points .
#
# + id="P9664e-mVMTN" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="3d21926f-572f-4a79-ad7b-bdbe3334094a"
t,k,m,c= sp.symbols('t k m c')
x = sp.Function('x')
diffeq = sp.Eq(m*x(t).diff(t,t) +k*x(t) + c* x(t).diff(t) ,0)
diffeq
# + [markdown] id="kChagUv8JtIv"
# $\text{State space representation}$
#
# $\frac{d}{dt}$
# $\begin{bmatrix}
# x \\
# \dot{x}
# \end{bmatrix}$
# $=$
# $
# \begin{bmatrix}
# 0 & 1 \\
# \frac{-k}{m} & \frac{-c}{m}
# \end{bmatrix}
# $
# $\begin{bmatrix}
# x \\
# \dot{x}
# \end{bmatrix}$
#
# $\frac{dx}{dt}=\dot{x}$
#
# $\frac{d^2x}{dt^2} =\frac{-k}{m} \ x +\frac{-c}{m} \ \dot{x}$
#
# + [markdown] id="CXelBMmysTw2"
# #### Problem configuration
# + colab={"base_uri": "https://localhost:8080/", "height": 54} id="ospW1dBky9gx" outputId="e6dfde7b-daf9-4e00-ac22-ea36539d523f"
kv = 1 #spring constant
mv = 1 # mass
cv = 0.5 # damping coefficient
diffeq = diffeq.subs({k:kv,c:cv,m:mv})
diffeq
# + colab={"base_uri": "https://localhost:8080/", "height": 38} id="nOl3m8RqsPN9" outputId="045dd056-fe89-487a-fa84-0e76114a0fb6"
# inital conditon
sp.Eq(x(t).subs(t,0),0)
# + colab={"base_uri": "https://localhost:8080/", "height": 60} id="T5VcnYbDsJYB" outputId="7d94d12d-0472-4ad3-b3ff-8af3bec3c9ea"
# inital condition
sp.Eq(x(t).diff(t).subs(t,0),1)
# + id="r9KVq1yjYfld"
ics={
x(0):0,
x(t).diff(t).subs(t,0):1,
}
# + id="_nI9-fj7wmKA"
#Displacement
D = sp.simplify(sp.dsolve(diffeq,ics=ics).rhs)
#Velocity
V = D.diff(t)
#Force
F = -kv * D - cv* D.diff(t)
# + colab={"base_uri": "https://localhost:8080/", "height": 312} id="jFA7s9u4z4Uk" outputId="c565564d-5413-4a3a-beca-8bfed7f7ed2e"
D_func = sp.lambdify(t,D)
V_func = sp.lambdify(t,V)
F_func = sp.lambdify(t,F)
# Full solution
dT = 1e-3
T_full = np.arange(0,3*np.pi+dT,dT)
D_full = D_func(T_full)
V_full = V_func(T_full)
F_full = F_func(T_full)
# Measurement solution
dT = np.pi/2
T_part = np.arange(0,3*np.pi+dT,dT)
D_part = D_func(T_part)
V_part = V_func(T_part)
F_part = F_func(T_part)
plt.figure(1);
plt.plot(T_full,F_full,'k',label='True spring model');
plt.scatter(T_part,F_part,label='Force measurements');
plt.legend();
plt.xlabel('t');
plt.ylabel('F')
plt.title('Force measurements')
# + colab={"base_uri": "https://localhost:8080/"} id="figsqCK4igpx" outputId="84220b63-1bbc-4cda-cdad-86e381343c84"
# time position
X = T_part.reshape(-1,1)
# force position
Y = F_part.reshape(-1,1)
conds = jnp.concatenate([X,Y],axis=1)
#collocation points
colloc = jnp.linspace(0,3*np.pi,1_000).reshape(-1,1)
# + [markdown] id="NQ61lEQeXgrc"
# ### Constructing the MLP
# + id="Lml6PGLPZgmr"
def ODE_loss(t,x):
x_t=lambda t:jax.grad(lambda t:jnp.sum(x(t)))(t)
x_tt=lambda t:jax.grad(lambda t:jnp.sum(x_t(t)))(t)
return x_tt(t) + 1.0*x(t)
# + id="KoZZJl2TbI_n"
def init_params(layers):
keys = jax.random.split(jax.random.PRNGKey(0),len(layers)-1)
params = list()
for key,n_in,n_out in zip(keys,layers[:-1],layers[1:]):
lb, ub = -(1 / jnp.sqrt(n_in)), (1 / jnp.sqrt(n_in)) # xavier initialization lower and upper bound
W = lb + (ub-lb) * jax.random.uniform(key,shape=(n_in,n_out))
B = jax.random.uniform(key,shape=(n_out,))
params.append({'W':W,'B':B})
return params
def fwd(params,t):
X = jnp.concatenate([t],axis=1)
*hidden,last = params
for layer in hidden :
X = jax.nn.tanh(X@layer['W']+layer['B'])
return X@last['W'] + last['B']
@jax.jit
def MSE(true,pred):
return jnp.mean((true-pred)**2)
def loss_fun(params,colloc,conds,Alpha=1.0,Beta= 0.0):
'''
Conds => Function satisfied at measured points by supervised loss and physics loss
colloc=> Function satisfied at the collocation points by physics loss
'''
X,Y = conds[:,[0]],conds[:,[1]]
t_c = colloc #collocation point
loss = 0.
supervised_loss = Alpha * jnp.mean((fwd(params,X) - Y)**2)
loss += supervised_loss
xfunc = lambda t : fwd(params,t)
physics_loss = Beta * jnp.mean(ODE_loss(t_c,xfunc)**2)
loss += physics_loss
return loss
@jax.jit
def update(opt_state,params,colloc,conds,Alpha=1.0,Beta=0.0):
# Get the gradient w.r.t to MLP params
grads=jax.jit(jax.grad(loss_fun,0))(params,colloc,conds,Alpha,Beta)
#Update params
updates, opt_state = optimizer.update(grads, opt_state)
params = optax.apply_updates(params, updates)
return opt_state,params
# + [markdown] id="OGrOF0D3fR-D"
# ### Supervised only
#
# + [markdown] id="Uf8_lWUOxb-Y"
# We try to map the x->F(x) with strictly supervised loss formulation
# + id="jySmbUwic5yk" colab={"base_uri": "https://localhost:8080/"} outputId="9e9b7db8-5cb9-4e64-cc2d-c1f03d979b76"
# %%time
Alpha,Beta = 1.,0.
# construct the MLP of 6 hidden layers of 8 neurons for each layer
params = init_params([1] + [8]*6+[1])
optimizer = optax.adam(1e-3)
opt_state = optimizer.init(params)
epochs = 10_000
for _ in range(epochs):
opt_state,params = update(opt_state,params,colloc,conds,Alpha,Beta)
# print loss and epoch info
if _ %(1000) ==0:
print(f'Epoch={_}\tloss={loss_fun(params,colloc,conds,Alpha,Beta):.3e}')
# + colab={"base_uri": "https://localhost:8080/", "height": 312} id="XafxrUPtc0k6" outputId="a27ddc9b-5db0-4c16-8e10-b2e52b308797"
pred = fwd(params,X)
plt.figure(3);
plt.plot(T_full,F_full,'k',label='True force');
plt.scatter(T_part,F_part,c='k',label='Measurement points force');
plt.plot(T_full.reshape(-1,1),fwd(params,T_full.reshape(-1,1)),'r',label='Predicted force')
plt.scatter(T_part.reshape(-1,1),fwd(params,T_part.reshape(-1,1)),c='r',label='Predicted points force')
plt.legend();
plt.xlabel('t');
plt.ylabel('F')
plt.title('Supervised training only')
# + [markdown] id="_dzrCiiigBnc"
# ### Physics loss with Supervised loss
# + [markdown] id="-Q2yEr_gxk29"
# In here , **we use hookes law approximation** for the physics loss as we observe that the model at hand is ideally a spring
#
# $Loss \ function = \alpha . SupervisedLoss + \beta .PhysicsLoss$
#
# $ \alpha = 1 \ , \ \beta = 0.1$
# + colab={"base_uri": "https://localhost:8080/"} id="FDZVtsrgfwKd" outputId="b5f76586-66c9-4199-b9dc-598c2670dff6"
# %%time
Alpha,Beta = 1.,0.1
# construct the MLP of 6 hidden layers of 8 neurons for each layer
params = init_params([1] + [8]*6+[1])
optimizer = optax.adam(1e-3)
opt_state = optimizer.init(params)
epochs = 10_000
for _ in range(epochs):
opt_state,params = update(opt_state,params,colloc,conds,Alpha,Beta)
# print loss and epoch info
if _ %(1000) ==0:
print(f'Epoch={_}\tloss={loss_fun(params,colloc,conds,Alpha,Beta):.3e}')
# + colab={"base_uri": "https://localhost:8080/", "height": 312} id="RR5R0TM0gJmq" outputId="f4c92859-fcb1-4543-8bd4-525d279642ed"
pred = fwd(params,X)
plt.figure(3);
plt.plot(T_full,F_full,'k',label='True force');
plt.scatter(T_part,F_part,c='k',label='Measurement points force');
plt.plot(T_full.reshape(-1,1),fwd(params,T_full.reshape(-1,1)),'--r',label='Predicted force')
plt.scatter(T_part.reshape(-1,1),fwd(params,T_part.reshape(-1,1)),c='r',label='Predicted points force')
plt.legend();
plt.xlabel('t');
plt.ylabel('F')
plt.title('Supervised loss + PINN loss training')
| [4]_ODE_Supervised_and_PINN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
# !{sys.executable} -m pip install python-dotenv
# +
import boto3
from boto3 import s3
import pandas as pd
import numpy as np
from dotenv import find_dotenv, load_dotenv
import os
import s3fs
import io
from pathlib import Path
from sklearn.model_selection import train_test_split
from src.features.dates import convert_to_date
from src.data.sets import split_sets_random
from src.data.sets import save_sets
from src.models.null import NullModel
# -
# %load_ext autoreload
# %autoreload 2
load_dotenv(find_dotenv())
aws_access_key_id = os.environ.get('aws_access_key_id')
aws_secret_access_key = os.environ.get('aws_secret_access_key')
# # Set up directories
project_dir = Path.cwd().parent
data_dir = project_dir / 'data'
raw_data_dir = data_dir / 'raw'
interim_data_dir = data_dir / 'interim'
processed_data_dir = data_dir / 'processed'
# Where is boto3 looking for my credentials?
boto3.set_stream_logger('botocore', level='DEBUG')
def list_bucket_contents(bucket,
aws_access_key_id,
aws_secret_access_key,
match=''):
s3 = boto3.resource('s3',
aws_access_key_id=aws_access_key_id,
aws_secret_access_key=aws_secret_access_key)
bucket_resource = s3.Bucket(bucket)
# bucket_resource = s3_resource.Bucket(bucket)
for key in bucket_resource.objects.all():
if match in key.key:
print(key.key)
list_bucket_contents(bucket='nyc-tlc',
aws_access_key_id=aws_access_key_id,
aws_secret_access_key=aws_secret_access_key,
match='2020')
s3 = boto3.client('s3',
aws_access_key_id=aws_access_key_id,
aws_secret_access_key=aws_secret_access_key)
obj = s3.get_object(Bucket='nyc-tlc', Key='trip data/yellow_tripdata_2020-04.csv')
df = pd.read_csv(io.BytesIO(obj['Body'].read()))
df.head()
df.shape
df.info()
df.describe()
path = raw_data_dir / 'df.csv'
df.to_csv(path, index=False)
# # Cleaning the Data
df_cleaned = df.copy(deep=True)
date_cols = df_cleaned.filter(like='date').columns
df_cleaned.loc[:, date_cols] = df_cleaned[date_cols].apply(convert_to_date)
# ## Add `trip_duration`
df_cleaned.loc[:, 'trip_duration'] = (df_cleaned.tpep_dropoff_datetime - df_cleaned.tpep_pickup_datetime).dt.seconds
# ## Binning `trip_duration`
df_cleaned.loc[:, 'trip_duration'] = pd.cut(df_cleaned['trip_duration'],
bins=[0, 300, 600, 1800, 100000],
labels=['x<5min', 'x<10min', 'x<30min', 'x>=30min'])
# ## Extract date features
df_cleaned.loc[:, 'tpep_pickup_month'] = df_cleaned.tpep_pickup_datetime.dt.month
df_cleaned.loc[:, 'tpep_pickup_hourofday'] = df_cleaned.tpep_pickup_datetime.dt.hour
df_cleaned.loc[:, 'tpep_pickup_dayofweek'] = df_cleaned.tpep_pickup_datetime.dt.day
# ## One-hot encoding
dummy_cols = ['VendorID', 'RatecodeID', 'store_and_fwd_flag']
df_cleaned = pd.get_dummies(df_cleaned, columns=dummy_cols)
# ## Drop columns
drop_cols = ['tpep_pickup_datetime',
'tpep_dropoff_datetime',
'PULocationID',
'DOLocationID']
df_cleaned.drop(columns=drop_cols, inplace=True)
# ## Save data
path = interim_data_dir / 'df_cleaned.csv'
df_cleaned.to_csv(path, index=False)
# # Split Data
X_train, X_test, y_train, y_test = split_sets_random(df_cleaned,
target_col='passenger_count',
to_numpy=True)
# ## Save the data sets
save_sets(X_train, X_test, y_train, y_test, processed_data_dir)
df_cleaned
# # Baseline model
base_model = NullModel(target_type='classification')
| notebooks/1_data_prep.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:metis] *
# language: python
# name: conda-env-metis-py
# ---
# # Scraping Job Postings from LinkedIn
# This code is adapted and modified from the following article: https://maoviola.medium.com/a-complete-guide-to-web-scraping-linkedin-job-postings-ad290fcaa97f and Cohort 2's work.
# ### Data Source
# LinkedIn job post board. This data collection is focusing on job posts near Rancho Cardova, California in the past months. <br>
# This code will focus on **date, job titles, company names, job descriptions, and job criteria**.
# ### Import Libraries
# +
#Import packages
import time, os
import pandas as pd
from selenium import webdriver
from selenium.webdriver.common.keys import Keys
from selenium.webdriver.support.ui import Select
from bs4 import BeautifulSoup as bs
import requests
import re
import pickle
#Hide Warnings
import warnings
warnings.filterwarnings('ignore')
# -
# ### Initialize Chromedriver for Selenium
# The chromedriver is being used in this case. If you have not previously installed it, you can do so at:
# https://chromedriver.chromium.org/downloads. <br>
# Be sure to move the chromedriver to the **Application Folder** for the code below to work. <br>
# <br>
# The Selenium functions here are tasked to: <br>
# - Get the location set to Rancho Cordova
# - Select 'Past Month' in "Date Posted"
# - Select 'within 10 miles' in "Distanct"
# +
# Set up chromedirver
chromedriver = "/Applications/chromedriver"
os.environ["webdriver.chrome.driver"] = chromedriver
driver = webdriver.Chrome(chromedriver)
source = 'https://www.linkedin.com/jobs'
driver.get(source)
# Get Location to Rancho Cordova
location_box_clear = driver.find_element_by_xpath('//*[@id="JOBS"]/section[2]/button')
location_box_clear.click()
location_box = driver.find_element_by_xpath('//*[@id="JOBS"]/section[2]/input')
location_box.click()
location_box.send_keys("Rancho Cordova, California, United States")
location_box.send_keys(Keys.RETURN)
# +
# Select past month
time_dropdown = driver.find_element_by_xpath('//*[@id="jserp-filters"]/ul/li[1]/div/div/button')
time_dropdown.click()
past_month_button = driver.find_element_by_xpath('//*[@id="jserp-filters"]/ul/li[1]/div/div/div/fieldset/div/div[3]/label')
past_month_button.click()
time_done_button = driver.find_element_by_xpath('//*[@id="jserp-filters"]/ul/li[1]/div/div/div/button')
time_done_button.click()
# Pause in action or linkedin will jump to sign in page
time.sleep(5)
# within 10 miles
distance_dropdown = driver.find_element_by_xpath('//*[@id="jserp-filters"]/ul/li[2]/div/div/button')
distance_dropdown.click()
filter_10mi = driver.find_element_by_xpath('//*[@id="jserp-filters"]/ul/li[2]/div/div/div/fieldset/div/div[1]/label')
filter_10mi.click()
distance_done_button = driver.find_element_by_xpath('//*[@id="jserp-filters"]/ul/li[2]/div/div/div/button')
distance_done_button.click()
# -
# Additional filters can be set with **Company, Salary, Location, Job Type, Experience Level, On-site/Remote**.
# ### How many job posts are associated with the job seach?
# +
#How many jobs are curently available within 10 miles of Rancho Cordova on LinkedIn
no_of_jobs = driver.find_element_by_css_selector('h1>span').get_attribute('innerText')
print('There are', no_of_jobs, 'jobs available within 10 miles of Rancho Cordova on LinkedIn over the past month.')
# -
# ### Show all the jobs
# The following segment of code will scroll and click "Show more job" until all available job post are showing.
# +
#Browse all jobs for the search.
# Set pause time
SCROLL_PAUSE_TIME = 10
last_height = driver.execute_script("return document.body.scrollHeight")
while True:
#Scroll until hit the see more jobs button.
driver.execute_script('window.scrollTo(0, document.body.scrollHeight);')
# Wait to load page
time.sleep(SCROLL_PAUSE_TIME)
try:
#Click the see more jobs button and then keep scrolling.
driver.find_element_by_xpath('//*[@id="main-content"]/section/button').click()
time.sleep(15)
print("clicked loading button")
except:
pass
time.sleep(15)
print("no loading button")
# Calculate new scroll height and compare with last scroll height
new_height = driver.execute_script("return document.body.scrollHeight")
# Wait to load page
time.sleep(SCROLL_PAUSE_TIME)
# Stop the scrolling and button clicking if the page isn't loading more jobs
if new_height == last_height:
print("loading button stopped working")
break
last_height = new_height
# This can take awhile
# -
# ***No more loading with clicking on the button on the webdriver***
# ### Create a list of all jobs in the search
#Create a list of the jobs.
job_lists = driver.find_element_by_class_name('jobs-search__results-list')
jobs = job_lists.find_elements_by_tag_name('li')
#Test that it collected all jobs.
#If it significantly dropped, the sleep.time time may need to be increased to allow:
#More loading time or
#Not set off restrictions for the site.
print(len(jobs), 'were collected from the search')
# It seems like LinkedIn only lets you get to that amount of job listings (close to the number from previous cohort) <br>
# Another option is adding more filter for the search to narrow down the search.
# ### Getting Job basic information
# The following steps can be done together, but there is higher risk of the tasks being canceled due to interactions with the the website. Overall, there are fewer errors and everything is completed in less time by breaking it all up.
# +
#Pull basic information from each job.
job_title = []
company_name = []
date = []
job_link = []
for job in jobs:
job_title0 = job.find_element_by_css_selector('h3').get_attribute('innerText')
job_title.append(job_title0)
company_name0 = job.find_element_by_css_selector('h4').get_attribute('innerText')
company_name.append(company_name0)
date0 = job.find_element_by_css_selector('div>div>time').get_attribute('datetime')
date.append(date0)
job_link0 = job.find_element_by_css_selector('a').get_attribute('href')
job_link.append(job_link0)
# -
#See first 5 of each for verification.
print('Job Titles:',job_title[:5])
print(' ')
print('Company Names:',company_name[:5])
print(' ')
print('Date:', date[:5])
# Create and save a dataframe of the collected data.
job_post_data = pd.DataFrame({'Date': date,
'Company': company_name,
'Title': job_title,
'Job Link': job_link})
job_post_data.head()
# ### Getting more job details
# **Note**: The following code will have longer times in scraping due to time.sleep(), but it helps working around StaleElementReferenceException.
# +
#Inital job description and criteria list
jd = []
cl = []
#Get job descriptions and criteria list
for job in jobs:
job.click()
jd_path = 'show-more-less-html__markup'
detail_path = 'description__job-criteria-list'
try:
jd0 = driver.find_element_by_class_name(jd_path).get_attribute('innerText')
jd.append(jd0)
details = driver.find_element_by_class_name(detail_path).get_attribute('innerText')
cl.append(details)
time.sleep(20)
except: # working around StaleElementReferenceException
time.sleep(15)
jd0 = driver.find_element_by_class_name(jd_path).get_attribute('innerText')
jd.append(jd0)
details = driver.find_element_by_class_name(detail_path).get_attribute('innerText')
cl.append(details)
time.sleep(20)
# -
# Verify description is correct
print(jd[0])
print(cl[0])
# +
# Initial criteria list
# cl = []
# #Get job criteria.
# # batchsize = 5
# # for i in range(0, len(jobs), batchsize):
# # batch = jobs[i:i+batchsize]
# for job in batch:
# job.click()
# detail_path = 'description__job-criteria-list'
# try:
# details = driver.find_element_by_class_name(detail_path).get_attribute('innerText')
# cl.append(details)
# time.sleep(20)
# except:
# time.sleep(15)
# details = driver.find_element_by_class_name(detail_path).get_attribute('innerText')
# cl.append(details)
# time.sleep(20)
# +
# Verify criteria is correct
# print(cl[0])
# -
# Verify that new lists are the same length as the df
print(len(cl))
print(len(jd))
job_post_data["job_description"] = jb
job_post_data["criteria"] = cl
job_post_data.head()
job_post_data.to_csv('../Data/LinkedIn_Job_Postings_notcleaned.csv', index = False)
# Another notebook will detail the cleaning process for the text columns.
| Notebooks/.ipynb_checkpoints/webscrapping_linkedin_remoteonly-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import plotly.plotly as py
import plotly.graph_objs as go
from plotly.offline import download_plotlyjs, init_notebook_mode, plot, iplot
import pandas as pd
import re
from datetime import datetime, timedelta
from dateutil import parser
# Plotly inline
init_notebook_mode(connected=True)
# -
# Generate graphs
def parse_logs(filename_pairs_arr, hours=1):
""""Function to generate plotly graph of CellRanger Monitor Logs with CellRanger std_out Annotations
Inputs:
filename_pairs_arr: Array of usage monitoring logs and matching annotation logs in this format-
[["monitroig_log.log","corresponding_std_out.txt"], ["monitroig_log2.log","corresponding_std_out2.txt"]]
hours: default length of x axis in hours
Outpus:
Plotly Graph of Maps: The Legend items are grouped by usage type, disk, core and mem. The title is the VM stats of the run.
The CellRanger task names are outputted in the middle of the graph vertically at the time they occured.
The xaxis starts at the time of the first task started, and is defaulted to have the range of hours input.
"""
# Collection Arrays
# Data: Plotly traces of line graphs of usages
data = []
# Start times of every task
earlys = []
# Cellranger Tasks
events = []
# Iterate through filename pairs and populate collection arrays
for filename_pair in filename_pairs_arr:
# monitoring log is first file
monitor = filename_pair[0]
# std out is second file
std = filename_pair[1]
# read std out file
with open(std) as b:
lines = b.readlines()
# parse through the std out file to get major task names
for line in lines:
# theres a bunch of meta output that we ignore like copyright info
# lines that matter have "[runtime]" in them, so get them
if "[runtime]" in line:
# split the line into components we want
# EXAMPLE LINE WE WANT
""""
2018-07-16 21:13:45 [runtime] (ready) ID.HJCVJBGX5.MAKE_FASTQS_CS.MAKE_FASTQS.PREPARE_SAMPLESHEET
"""
line_arr = line.strip().split()
# important tasks have '(ready)' after '[runtime]'
if line_arr[3] == '(ready)':
# parse date
dt = parser.parse(' '.join(line_arr[0:2]))
# Get the event name, can be different formats, this should generally get all of them into right format though
event_name = ''.join(line_arr[4:]).split(".")[-1]
# add formatted event [datetime, event_name] to collection
events = events + [[dt, event_name]]
# Open the monitor log and parse it
with open(monitor) as f:
lines = f.readlines()
# Read Caps of Stats we monitor, each one is parsed and converted to float
cpu_cap = int(re.sub("[^$0-9.]","", [line for line in lines if 'CPU:' in line][0]))
mem_cap = float(re.sub("[^$0-9.]","", [line for line in lines if 'Total Memory:' in line][0]))
# check if the last line ends in T, if it does use terabytes as unit for memory
memory_tb = [line for line in lines if 'Total Memory:' in line][0].strip()[-1] == 'T'
memory_unit = "memory usage (TB)" if memory_tb else "memory usage (GB)"
disk_cap = float(re.sub("[^$0-9.]","", [line for line in lines if 'Total Disk space:' in line][0]))
# check if the last line ends in T, if it does use terabytes as unit for disk space
disk_tb = [line for line in lines if 'Total Disk space:' in line][0].strip()[-1] == 'T'
disk_unit = 'disk usage (TB)' if disk_tb else "disk usage (GB)"
# Collections for plotly trace
# cpu usages
cpu = []
# mem usages
mem = []
# disk usages
disk = []
# measurement times
time = []
# parse through each line and add it to collection
# EXAMPLE MONITORING LINE GROUP
""""
[Tue Jul 17 04:14:59 UTC 2018]
* CPU usage: 8.1%
* Memory usage: 6%
* Disk usage: 57%
"""
for line in lines:
if 'CPU usage:' in line:
# if it's a cpu line parse to float and add to cpu collector
cpu = cpu + [float(re.sub("[^$0-9.]","", line))]
elif 'Memory usage:' in line:
# if it's a mem line parse to float and add to mem collector
mem = mem + [float(re.sub("[^$0-9.]","", line ))]
elif 'Disk usage:' in line:
# if it's a disk line parse to float and add to disk collector
disk = disk + [float(re.sub("[^$0-9.]","", line))]
elif '[' in line:
# if it's a time line parse to datetime object and add to time collector
time = time + [parser.parse(line.replace("[", '').replace("]", '').strip())]
# create a dataframe to make adding to plotly easier
df = pd.DataFrame(data={'cores': cpu, memory_unit: mem, disk_unit: disk, 'time':time})
# create the title- eg "Usage: 64.0 Cores, 236.0 G Memory, 394.0 G Disk Space"
title_mem = " TB Memory, " if memory_tb else " GB Memory, "
title_disk = " TB Disk Space" if disk_tb else " GB Disk Space"
title = "Usage: " + str(cpu_cap) + " Cores, " + str(mem_cap) + title_mem + str(disk_cap) + title_disk
# CPU usage trace (Note the legend group)
trace1 = go.Scatter(
x = df.time,
y = df['cores'],
name = monitor + ' core usage',
legendgroup = 'core usage',
)
# Memory Usage trace (Note the legend group)
trace2 = go.Scatter(
x = df.time,
y = df[memory_unit],
name = monitor + ' ' + memory_unit,
legendgroup = memory_unit,
)
# Disk Usage trace (Note the legend group)
trace3 = go.Scatter(
x = df.time,
y = df[disk_unit],
name = monitor + ' ' + disk_unit,
legendgroup = disk_unit,
)
# Add to data collector for plotly
data = data + [trace1, trace2, trace3]
# get the earliest time (aka start of task) and add it to early collector
earlys = earlys + [min(time)]
# TODO figure out a better way to make sure these annotations, which can be long, do not overlap
def height(num, x =40, maxh =60, by =10):
"""Generator to alternate heights of annotation labels
Inputs:
num: number of heights we want to output
minh: min height of annotation
maxh: max height of annotation
by: amount to increase height every time
Outputs:
Yields varying heights from minh, minh + by... maxh, minh, minh + by...
"""
n = 0
minh = 40
while n < num:
if minh == maxh:
yield x
minh = 40
n +=1
yield minh
minh += by
n += 1
# get heights of annotations
heights = height(len(events))
# create annotations layout list
annotations = []
for event in sorted(events, key=lambda x : x[0]):
annotation = dict(x = event[0], y = next(heights),
xref = 'x', yref = 'y', text = event[1],
textangle = -45, showarrow = False)
annotations = annotations + [annotation]
# generate layout using collectors
layout = dict(title = title,
xaxis = dict(title = 'Time', range = [min(earlys), min(earlys)+ timedelta(hours=hours)]),
yaxis = dict(title = '% Usage', range = [0,100]),
annotations = annotations)
# plot
iplot({'data':data, 'layout':layout})
| scripts/benchmarks/cellranger_orchestra_pipeline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tarea
# Calcular el valor presente de los siguientes flujos:
flujos = [1, 1, 1, 1, 1, 1, 1, 1, 1, 101]
# Los plazos de los flujos en días son:
plazos = [182, 365, 547, 730, 912, 1095, 1277, 1460, 1642, 1825]
# Para calcular el valor presente usar una tasa del 2.00% compuesta act/365, la misma convención de la TIR de mercado que usa la Bolsa de Comercio.
# **Pista:**
for p in plazos:
print(p)
for f in flujos:
print(f)
# Investigar el `zip` de 2 o más `list` en Python, esto permite iterar en paralelo con varias `list`.
| 01-01_c_intro-python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="f89E_hzQw6HO"
# # Práctico 2: Recomendación de videojuegos
#
# En este práctico trabajaremos con un subconjunto de datos sobre [videojuegos de Steam](http://cseweb.ucsd.edu/~jmcauley/datasets.html#steam_data). Para facilitar un poco el práctico, se les dará el conjunto de datos previamente procesado. En este mismo notebook mostraremos el proceso de limpieza, para que quede registro del proceso (de todas maneras, por el tamaño de los datos no recomendamos que pierdan tiempo en el proceso salvo que lo consideren útil a fines personales).
#
# El conjunto de datos se basa en dos partes: lista de juegos (items), y lista de reviews de usuarios sobre distintos juegos. Este último, en su versión original es muy grande, (pesa 1.3GB), por lo que será solo una muestra del mismo sobre la que trabajarán.
#
# A diferencia del conjunto de datos de LastFM utilizados en el [Práctico 1](./practico1.ipynb), en este caso los datos no están particularmente pensados para un sistema de recomendación, por lo que requerirá de un poco más de trabajo general sobre el dataset.
#
# La idea es que, de manera similar al práctico anterior, realicen un sistema de recomendación. A diferencia del práctico anterior, este será un poco más completo y deberán hacer dos sistemas, uno que, dado un nombre de usuario le recomiende una lista de juegos, y otro que dado el título de un juego, recomiende una lista de juegos similares. Además, en este caso se requiere que el segundo sistema (el que recomienda juegos basado en el nombre de un juego en particular) haga uso de la información de contenido (i.e. o bien harán un filtrado basado en contenido o algo híbrido).
# + [markdown] colab_type="text" id="5Qfs_yKqw6HQ"
# ## Obtención y limpieza del conjunto de datos
#
# El conjunto de datos originalmente se encuentra en archivos que deberían ser de formato "JSON". Sin embargo, en realidad es un archivo donde cada línea es un objeto de JSON. Hay un problema no obstante y es que las líneas están mal formateadas, dado que no respetan el estándar JSON de utilizar comillas dobles (**"**) y en su lugar utilizan comillas simples (**'**). Afortunadamente, se pueden evaluar como diccionarios de Python, lo cuál permite trabajarlos directamente.
# + colab={"base_uri": "https://localhost:8080/", "height": 106} colab_type="code" id="cvfAiYezAbVv" outputId="f231ba7d-d0c5-46e2-c8d4-afde76689a48"
# !pip install scikit-surprise
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="sDKq68unAePC" outputId="3ce54c20-c8de-4de3-92fd-fa3068d63440"
import surprise
surprise.__version__
# + colab={} colab_type="code" id="MLN3S_cJw6Hi"
import pandas as pd
import io # needed because of weird encoding of u.item file
import numpy as np
from surprise import KNNBaseline
from surprise import Dataset
from surprise import get_dataset_dir
from surprise import Reader, Dataset
from surprise.model_selection import cross_validate
from surprise import NormalPredictor
from surprise import KNNBasic
from surprise import KNNWithMeans
from surprise import KNNWithZScore
from surprise import KNNBaseline
from surprise import SVD
from surprise import BaselineOnly
from surprise import SVDpp
from surprise import NMF
from surprise import SlopeOne
from surprise import CoClustering
from surprise.accuracy import rmse
from surprise import accuracy
from surprise.model_selection import train_test_split
from surprise.model_selection import cross_validate
from surprise import NormalPredictor
from surprise import KNNBasic
from surprise import KNNWithMeans
from surprise import KNNWithZScore
from surprise import KNNBaseline
from surprise import SVD
from surprise import BaselineOnly
from surprise import SVDpp
from surprise import NMF
from surprise import SlopeOne
from surprise import CoClustering
from surprise.accuracy import rmse
from surprise import accuracy
from surprise.model_selection import train_test_split
from sklearn.feature_extraction.text import TfidfVectorizer, CountVectorizer
from sklearn.metrics.pairwise import linear_kernel, cosine_similarity
from nltk.stem.snowball import SnowballStemmer
from nltk.stem.wordnet import WordNetLemmatizer
from nltk.corpus import wordnet
# + [markdown] colab_type="text" id="qbYFVB4ww6Hy"
# ## Conjunto de datos limpio
#
# Para descargar el conjunto de datos que se utilizará en el práctico, basta con ejecutar la siguiente celda.
# + colab={"base_uri": "https://localhost:8080/", "height": 124} colab_type="code" id="99eMOQ6fw6Hz" outputId="e3b53637-3a4e-4de9-e632-63861519aef2" language="bash"
#
# mkdir -p data/steam/
# curl -L -o data/steam/games.json.gz https://cs.famaf.unc.edu.ar/\~ccardellino/diplomatura/games.json.gz
# curl -L -o data/steam/reviews.json.gz https://cs.famaf.unc.edu.ar/\~ccardellino/diplomatura/reviews.json.gz
# + [markdown] colab_type="text" id="R3YSNgk1w6H2"
# ## Ejercicio 1: Análisis Exploratorio de Datos
#
# Ya teniendo los datos, podemos cargarlos y empezar con el práctico. Antes que nada vamos a hacer una exploración de los datos. Lo principal a tener en cuenta para este caso es que debemos identificar las variables con las que vamos a trabajar. A diferencia del práctico anterior, este conjunto de datos no está documentado, por lo que la exploración es necesaria para poder entender que cosas van a definir nuestro sistema de recomendación.
# + colab={} colab_type="code" id="4U90qeGLw6H3"
import pandas as pd
# + [markdown] colab_type="text" id="ow_8uKHYw6H5"
# ### Características del conjunto de datos sobre videojuegos
#
# Las características del conjunto de datos de videojuegos tienen la información necesaria para hacer el "vector de contenido" utilizado en el segundo sistema de recomendación. Su tarea es hacer un análisis sobre dicho conjunto de datos y descartar aquella información redundante.
# + colab={"base_uri": "https://localhost:8080/", "height": 602} colab_type="code" id="-kLuoSNww6H6" outputId="500acf7c-de72-4895-d4a1-e5756756ab2c"
games = pd.read_json("./data/steam/games.json.gz")
games.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="XGuUlwm9NR0x" outputId="76089e40-4cc3-43c4-d83a-24678662d161"
games[games['app_name'].str.find('Age of')>-1]
# + [markdown] colab_type="text" id="uqRzwVUlw6H9"
# #### Valores Nulos en el Dataset
# + colab={"base_uri": "https://localhost:8080/", "height": 284} colab_type="code" id="hRDo3bpfw6H9" outputId="264c729d-f0ad-4826-c36e-5f24aa12fa78"
pd.isna(games).sum()
# + [markdown] colab_type="text" id="Ga99ebmTw6IA"
# #### Verificar app_name nula
# + colab={"base_uri": "https://localhost:8080/", "height": 123} colab_type="code" id="d9XGpqwAw6IB" outputId="0fc93544-5ce3-441e-d097-ca3fc34e793b"
games[games['app_name'].isnull()]
# + [markdown] colab_type="text" id="I9o7QAoXw6IF"
# - El primer caso no se pudo encontrar los datos correspondientes al juego por lo que se elimina.
# - Se buscó en la página https://store.steampowered.com/app/317160/_/ el nombre del juego con id 317160 y es DUET
#
# + colab={} colab_type="code" id="xB6Lo6kvw6IG"
games.loc[games.id == 317160.0, 'app_name'] = 'DUET'
# + colab={} colab_type="code" id="Kg94tMDPw6IJ"
games = games.drop(games[games['id'].isnull() & games['app_name'].isnull()].index)
# + [markdown] colab_type="text" id="9iKJ3tPPw6IP"
# #### Verificar los dos que tienen id nulo
# + colab={"base_uri": "https://localhost:8080/", "height": 198} colab_type="code" id="zpG26Bwzw6IQ" outputId="145a6e74-a583-4624-d7c8-821546e0cac2"
games[games['id'].isnull()]
# + [markdown] colab_type="text" id="F_-mBeFlw6IT"
# - En steam sale que el ID de este juego es 200260, por lo que hacemos verificacion para asegurar que no esta duplicado
# + colab={"base_uri": "https://localhost:8080/", "height": 198} colab_type="code" id="0Im1uh0zw6IT" outputId="f60c20d2-79de-45cb-cd0a-2d2fbc2aece6"
games[games['id']==200260]
# + [markdown] colab_type="text" id="IKFSrhyjw6IW"
# - Efecticamente está duplicado por lo que despues se hará verificación de la unicidad de los elementos.
# + [markdown] colab_type="text" id="esooUNXjw6IX"
# #### Verificar tags y genres
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="sn0MwZmhw6IX" outputId="e0b1a396-7e38-4139-cfd9-2966e8fbe7e0"
games[games['tags'].isnull() & games['genres'].isnull()]
# + [markdown] colab_type="text" id="2JiOuht9w6Ia"
# import requests
# import re
# from simplejson import JSONDecodeError
# import json
# for index, records in games[games['tags'].isnull() & games['genres'].isnull()][:1].iterrows():
#
# response = requests.get(url = 'https://store.steampowered.com/app/'+str(int(215914)))
# print('https://store.steampowered.com/app/'+str(int(215914)))
# recomendation = re.search("GStoreItemData.AddStoreItemDataSet\(([^\)]+)\)", response.text, re.IGNORECASE)
# try:
# if recomendation:
# titles = recomendation.group(1)
# try:
# y = json.loads(titles)
# except JSONDecodeError as error:
# break
# print(y['rgApps'].keys())
# if len(y['rgApps']) != 0:
# for rec_id in list(y['rgApps'].keys()):
# print(rec_id)
# response = requests.get(url = 'https://store.steampowered.com/app/'+str(rec_id))
# soup = BeautifulSoup(response.text, "html.parser")
# try:
#
# font = soup.find_all("a", href=re.compile("genre"),attrs={'class': None})
# genre_list = []
# for each_a in font:
# genre_list.append(each_a.text)
# print(genre_list)
# break
# except AttributeError as error:
# print("No reco")
# except AttributeError as error:
# print("No <a> found")
#
#
# + [markdown] colab_type="text" id="SUegFfj-Xrbs"
# #### Verificar Duplicidad
# + colab={} colab_type="code" id="LrjvDGD8XuuC"
duplicated = games[games.duplicated('app_name',keep=False)].sort_values(by='app_name')
# + colab={"base_uri": "https://localhost:8080/", "height": 55} colab_type="code" id="BjqGYv9QdZND" outputId="2f354ae6-b596-45a0-8b47-e74bbe153343"
# creo un diccionarios de app_name con los id duplicados.
diccionario_dupli = {}
lista_dupli = []
for index,duplicated_game in duplicated.iterrows():
if duplicated_game[2] in diccionario_dupli:
diccionario_dupli[duplicated_game[2]].append(duplicated_game[10])
else:
diccionario_dupli[duplicated_game[2]] = [duplicated_game[10]]
lista_dupli.append(duplicated_game[10])
print(diccionario_dupli)
# + [markdown] colab_type="text" id="M6CArcddw6Ib"
# ### Características del conjunto de datos de reviews
#
# Este será el conjunto de datos a utilizar para obtener información sobre los usuarios y su interacción con videojuegos. Como se puede observar no hay un rating explícito, sino uno implícito a calcular, que será parte de su trabajo (deberán descubrir que característica les puede dar información que puede ser equivalente a un rating).
# + colab={"base_uri": "https://localhost:8080/", "height": 278} colab_type="code" id="fGPxnSUUw6Ib" outputId="290a3fad-5e93-4b15-d2e7-14ac9e28ede4"
reviews = pd.read_json("./data/steam/reviews.json.gz")
reviews.head()
# + colab={} colab_type="code" id="YlmR9g5SNLV4"
# + [markdown] colab_type="text" id="CcBxFCe0yRaH"
# - De este dataset vamos a considerar ***hours*** como el rating implicito del juego. Observamos que hay un total de 2442 usuarios que no ingresaron la cantidad de horas que jugaron el juego.
# + colab={"base_uri": "https://localhost:8080/", "height": 568} colab_type="code" id="cHtsbqfww6Id" outputId="107d544a-5a87-42c3-c5f2-4ffd84c3aa38"
reviews[reviews.hours.isnull()]
# + [markdown] colab_type="text" id="O76Od7Gdy3G_"
# - Elegimos un usuario que no haya ingresado las horas de un juego, en este caso product_id==228260. Observamos la cantidad de horas que los otros usuarios jugaron.
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 282} colab_type="code" id="4Aap9ntKyAVZ" outputId="700f1c1b-6e43-4051-cd64-91a952b54bee"
reviews[reviews.product_id==228260].describe()
# + [markdown] colab_type="text" id="2euQGv_uGJhH"
# ### Distribucion de horas jugadas
# + colab={"base_uri": "https://localhost:8080/", "height": 283} colab_type="code" id="NMoWAGQFFtxX" outputId="c189b83d-8667-4b99-b6b1-e2e7bd329da8"
reviews = pd.read_json("./data/steam/reviews.json.gz")
reviews['hours'].value_counts().sort_index(ascending=False).plot(kind='kde')
# + [markdown] colab_type="text" id="9Z6eBj0JGTz3"
# ### Distribucion de Juegos jugados
# + colab={"base_uri": "https://localhost:8080/", "height": 283} colab_type="code" id="qc0PHRNMGaVj" outputId="23c6f7bf-9b45-4c17-a6e9-511f99f10ef6"
reviews.groupby('product_id')['hours'].count().plot(kind='kde')
# + colab={"base_uri": "https://localhost:8080/", "height": 340} colab_type="code" id="-53pSlBkH4AO" outputId="e7ba0ac9-45bc-4524-caf1-057be4a3d504"
reviews.groupby('product_id')['hours'].count().sort_values(ascending =False).reset_index()[:10]
# + colab={"base_uri": "https://localhost:8080/", "height": 358} colab_type="code" id="gEJQlJwJGqo2" outputId="20d54b95-1fe1-4404-fbe5-f8f57912b1de"
mas_jugados = list(reviews.groupby('product_id')['hours'].count().sort_values(ascending =False).reset_index()[:10]['product_id'])
print(mas_jugados)
games[games['id'].isin(mas_jugados)][['app_name','id','genres']]
# + colab={"base_uri": "https://localhost:8080/", "height": 52} colab_type="code" id="w5PoofkPfa_4" outputId="74712b7d-bda4-41d2-98e9-a073a704f1ba"
n_users = reviews.username.unique().shape[0]
n_items = reviews.product_id.unique().shape[0]
print(str(n_users) + ' users')
print(str(n_items) + ' items')
# + colab={} colab_type="code" id="cXIVNWnBXU6k"
from scipy.sparse import csr_matrix
def create_X(df):
"""
Generates a sparse matrix from ratings dataframe.
Args:
df: pandas dataframe containing 3 columns (userId, movieId, rating)
Returns:
X: sparse matrix
user_mapper: dict that maps user id's to user indices
user_inv_mapper: dict that maps user indices to user id's
movie_mapper: dict that maps movie id's to movie indices
movie_inv_mapper: dict that maps movie indices to movie id's
"""
M = df['username'].nunique()
N = df['product_id'].nunique()
user_mapper = dict(zip(np.unique(df["username"]), list(range(M))))
artist_mapper = dict(zip(np.unique(df["product_id"]), list(range(N))))
#user_inv_mapper = dict(zip(list(range(M)), np.unique(df["username"])))
#artist_inv_mapper = dict(zip(list(range(N)), np.unique(df["product_id"])))
user_index = [user_mapper[i] for i in df['username']]
item_index = [artist_mapper[i] for i in df['product_id']]
X = csr_matrix((df["hours"], (user_index,item_index)), shape=(M,N))
return X
reviews = pd.read_json("./data/steam/reviews.json.gz")
X = create_X(reviews)
# + colab={"base_uri": "https://localhost:8080/", "height": 52} colab_type="code" id="mAhY3BLnYXob" outputId="865d9dbe-374d-4f1e-aa96-ea8c9fdcffa1"
n_total = X.shape[0]*X.shape[1]
n_ratings = X.nnz
sparsity = n_ratings/n_total
print(n_total)
print(f"Matrix sparsity: {round(sparsity*100,2)}%")
# + colab={} colab_type="code" id="07QstMwB34ML"
def print_pivot_table(ds):
return pd.pivot_table(reviews[:100],values='hours',index='product_id',columns='username')
# + [markdown] colab_type="text" id="7beS4FWGDLVr"
# ### Additional methods
#
# + colab={} colab_type="code" id="scX2hBG2w6Ii"
def clean_game_df(games):
games.loc[games.id == 317160.0, 'app_name'] = 'DUET'
games = games.drop(games[games['id'].isnull() & games['app_name'].isnull()].index)
return games
# + colab={} colab_type="code" id="EzwQslSLDlbH"
def normalizar(reviews):
reviews['hours'] = reviews.apply(lambda row: row['hours']/reviews.groupby('username')['hours'].mean()[row['username']], axis = 1)
return reviews
# + [markdown] colab_type="text" id="FcZ2RHHUw6If"
# ## Ejercicio 2 - Sistema de Recomendación Basado en Usuarios
#
# Este sistema de recomendación deberá entrenar un algoritmo y desarrollar una interfaz que, dado un usuario, le devuelva una lista con los juegos más recomendados.
# + [markdown] colab_type="text" id="kRCpxUZjn6uM"
# ### User-Based Collaborative Filtering
# + colab={"base_uri": "https://localhost:8080/", "height": 346} colab_type="code" id="gvpXk8IzCnMn" outputId="05915bb5-366d-4ab6-b00e-aa96649c2942"
from collections import defaultdict
from surprise import SVD
from surprise import Dataset
def read_item_names():
games = pd.read_json("./data/steam/games.json.gz")
games = clean_game_df(games)
rid_to_name = {}
name_to_rid = {}
for index, line in games.iterrows():
rid_to_name[line['id']] = line['app_name']
return rid_to_name
def get_top_n(predictions, n=10):
'''Return the top-N recommendation for each user from a set of predictions.
Args:
predictions(list of Prediction objects): The list of predictions, as
returned by the test method of an algorithm.
n(int): The number of recommendation to output for each user. Default
is 10.
Returns:
A dict where keys are user (raw) ids and values are lists of tuples:
[(raw item id, rating estimation), ...] of size n.
'''
# First map the predictions to each user.
top_n = defaultdict(list)
for uid, iid, true_r, est, _ in predictions:
top_n[uid].append((iid, est))
# Then sort the predictions for each user and retrieve the k highest ones.
for uid, user_ratings in top_n.items():
user_ratings.sort(key=lambda x: x[1], reverse=True)
top_n[uid] = user_ratings[:n]
return top_n
# First train an SVD algorithm on the movielens dataset.
reviews = pd.read_json("./data/steam/reviews.json.gz")
reviews = reviews[['username','product_id','hours']][:10000]
reviews = normalizar(reviews)
reviews['hours'] = pd.to_numeric(reviews['hours'], errors='coerce')
reviews = reviews.dropna(subset=['hours'])
reader = Reader(rating_scale=(reviews.hours.min(), reviews.hours.max()))
data = Dataset.load_from_df(reviews, reader)
trainset = data.build_full_trainset()
sim_options = {'name': 'pearson_baseline', 'user_based': True}
print("Algo KNNBasic")
algo = KNNBasic(sim_options=sim_options, verbose=True)
algo.fit(trainset)
# Than predict ratings for all pairs (u, i) that are NOT in the training set.
testset = trainset.build_anti_testset()
predictions = algo.test(testset)
top_n = get_top_n(predictions, n=10)
read_id = read_item_names()
# Print the recommended items for each user
# Print the recommended items for each user
# -
# Games recommended for user 'Spodermen'
for (iid, _) in top_n['Spodermen']:
print('\t'+(read_id[iid]))
# + [markdown] colab_type="text" id="xz0wnpt4Evzu"
# #### Se redujo el dataset porque me quedo sin memoria en COLAB
# + [markdown] colab_type="text" id="HNuHo_yln6uQ"
# ### Item-Based Collaborative Filtering
# + colab={} colab_type="code" id="J7JCUqITw6Il"
import io # needed because of weird encoding of u.item file
from surprise import KNNBaseline
from surprise import Dataset
from surprise import get_dataset_dir
from surprise import Reader, Dataset
def read_item_names():
"""Read the u.item file from MovieLens 100-k dataset and return two
mappings to convert raw ids into movie names and movie names into raw ids.
"""
games = pd.read_json("./data/steam/games.json.gz")
games = clean_game_df(games)
rid_to_name = {}
name_to_rid = {}
for index, line in games.iterrows():
rid_to_name[line['id']] = line['app_name']
name_to_rid[line['app_name']] = line['id']
return rid_to_name, name_to_rid
def get_top_n(game_user_name , k = 10, item=True):
print("GET TOP N")
# First, train the algortihm to compute the similarities between items
print("Reading reviews")
reviews = pd.read_json("./data/steam/reviews.json.gz")
reviews = reviews[['username','product_id','hours']][:20000]
reviews = normalizar(reviews)
reviews['hours'] = pd.to_numeric(reviews['hours'], errors='coerce')
reviews = reviews.dropna(subset=['hours'])
reader = Reader(rating_scale=(reviews.hours.min(), reviews.hours.max()))
data = Dataset.load_from_df(reviews, reader)
trainset = data.build_full_trainset()
sim_options = {'name': 'pearson_baseline', 'user_based': False}
print("Creting KNNWithMeans")
algo = KNNBasic(sim_options=sim_options, verbose=True)
print(cross_validate(algo, data, measures=[ 'RMSE'], cv=3, verbose=True))
print("Trainning")
algo.fit(trainset)
# Read the mappings raw id <-> movie name
rid_to_name, name_to_rid = read_item_names()
if item:
artist_raw_id = name_to_rid[game_user_name]
artist_inner_id = algo.trainset.to_inner_iid(artist_raw_id)
print("Get neighbors")
artist_neighbors = algo.get_neighbors(artist_inner_id, k=10)
# Convert inner ids of the neighbors into names.
artist_neighbors = (algo.trainset.to_raw_iid(inner_id)
for inner_id in artist_neighbors)
artist_neighbors = (rid_to_name[rid]
for rid in artist_neighbors)
print()
print('The 10 nearest neighbors of {} are:', game_user_name)
for artit in artist_neighbors:
print(artit)
else:
inner_uid = algo.trainset.to_inner_uid(game_user_name)
print("Get neighbors")
games_neighbors = algo.get_neighbors(inner_uid, k=10)
# Convert inner ids of the neighbors into names.
artist_neighbors = (algo.trainset.to_raw_iid(inner_id)
for inner_id in artist_neighbors)
artist_neighbors = (rid_to_name[rid]
for rid in artist_neighbors)
print()
print('The 10 nearest neighbors of {} are:', artist_name)
for artit in artist_neighbors:
print(artit)
# + colab={"base_uri": "https://localhost:8080/", "height": 748} colab_type="code" id="sBh_8ZzBLgBV" outputId="7321620a-736f-41b6-f164-dfc6e95409e6"
get_top_n('Age of Empires II HD')
# + colab={} colab_type="code" id="u6MVLNYoOySx"
# + [markdown] colab_type="text" id="MOHmlnY-KWy6"
# ### Look for best model
# + colab={"base_uri": "https://localhost:8080/", "height": 248} colab_type="code" id="d1ephepSGz53" outputId="ae99d48f-160e-428a-afe8-5cbbc4794cbf"
reviews = pd.read_json("./data/steam/reviews.json.gz")
reviews = reviews[['username','product_id','hours']][:20000]
reviews = normalizar(reviews)
reader = Reader(rating_scale=(reviews.hours.min(), reviews.hours.max()))
reviews.info()
reviews['hours'] = pd.to_numeric(reviews['hours'], errors='coerce')
reviews = reviews.dropna(subset=['hours'])
print(reviews[reviews['hours'].isnull()])
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="WUyzNfHaLneY" outputId="b9b08180-1dc2-4171-80bd-b086237c01fb"
import gc
data = Dataset.load_from_df(reviews, reader)
benchmark = []
# Iterate over all algorithms
for algorithm in [ KNNBaseline(), KNNBasic(), KNNWithMeans(), KNNWithZScore(), BaselineOnly()]:
# Perform cross validation
results = cross_validate(algorithm, data, measures=['RMSE'], cv=3, verbose=1)
# Get results & append algorithm name
tmp = pd.DataFrame.from_dict(results).mean(axis=0)
tmp = tmp.append(pd.Series([str(algorithm).split(' ')[0].split('.')[-1]], index=['Algorithm']))
benchmark.append(tmp)
gc.collect()
pd.DataFrame(benchmark).set_index('Algorithm').sort_values('test_rmse')
# + [markdown] colab_type="text" id="SteiZzTuBnKk"
# Best Algorithm KNNBasic
# + [markdown] colab_type="text" id="AKqSxHl8w6Ii"
# ## Ejercicio 3 - Sistema de Recomendación Basado en Juegos
#
# Similar al caso anterior, con la diferencia de que este sistema espera como entrada el nombre de un juego y devuelve una lista de juegos similares. El sistema deberá estar programado en base a información de contenido de los juegos (i.e. filtrado basado en contenido o sistema híbrido).
# -
# Basado en https://github.com/rounakbanik/movies/blob/master/movies_recommender.ipynb
# + [markdown] colab_type="text" id="HX-KqnUSp7Sd"
# ### Games Description Based Recommender
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="TD1dlGyufxpR" outputId="6327283e-9be1-4364-f949-d2e9b20b4325"
reviews = pd.read_json("./data/steam/reviews.json.gz").head(25601)
games = pd.read_json("./data/steam/games.json.gz")
reviews.shape
# + colab={} colab_type="code" id="S55nxAj86dGH"
reviews = reviews.merge(games, left_on='product_id', right_on='id', )
# -
reviews['app_name'].unique()
# + colab={"base_uri": "https://localhost:8080/", "height": 568} colab_type="code" id="meAOZuwNqXmu" outputId="3cbd7048-6309-4a5c-963d-9ce431ebd8c0"
reviews.info()
# + colab={} colab_type="code" id="1MNMkngMqwCR"
reviews['text'] = reviews['text'].fillna('')
# + colab={} colab_type="code" id="urSJ4wo6rCiI"
tf = TfidfVectorizer(analyzer='word',ngram_range=(1, 2),min_df=0, stop_words='english')
tfidf_matrix = tf.fit_transform(reviews['text'])
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="ajqc_V0vrmWy" outputId="42c89c23-ae79-409c-b6d3-ea655176c5ac"
tfidf_matrix.shape
# + [markdown] colab_type="text" id="PPpMbfwDrsr4"
# #### Cosine Similarity
# I will be using the Cosine Similarity to calculate a numeric quantity that denotes the similarity between two games. Mathematically, it is defined as follows:
#
# $cosine(x,y) = \frac{x. y^\intercal}{||x||.||y||} $
#
# Since we have used the TF-IDF Vectorizer, calculating the Dot Product will directly give us the Cosine Similarity Score. Therefore, we will use sklearn's linear_kernel instead of cosine_similarities since it is much faster.
# + colab={} colab_type="code" id="25LoZpl-ryvp"
cosine_sim = linear_kernel(tfidf_matrix, tfidf_matrix)
# + colab={} colab_type="code" id="jtaMqC-AsHjR"
cosine_sim[0]
# + [markdown] colab_type="text" id="TGSttzWpsnsY"
# We now have a pairwise cosine similarity matrix for all the movies in our dataset.
# + colab={} colab_type="code" id="uN3ZHtRZtFwB"
reviews = reviews.reset_index()
product_ids = reviews['app_name']
indices = pd.Series(reviews.index, index=reviews['app_name'])
# + colab={} colab_type="code" id="9B2k4aOwuKEG"
def get_recommendations(id):
idx = indices[id]
sim_scores = list(enumerate(cosine_sim[idx]))
sim_scores = sorted(sim_scores, key=lambda x: x[1], reverse=True)
sim_scores = sim_scores[1:31]
product_indices = [i[0] for i in sim_scores]
return list(product_ids.iloc[product_indices].head(10))
# + colab={"base_uri": "https://localhost:8080/", "height": 292} colab_type="code" id="XWh6pk_jutJj" outputId="5dfd24b6-119c-4ad2-e8e3-724f2170baa6"
get_recommendations('Need For Speed: Hot Pursuit')
# + [markdown] colab_type="text" id="coY9f_TswaJK"
# ### Metadata Based Recommender
# + [markdown] colab_type="text" id="20NX256-to8j"
# Dado que el recomendador basado en los comentarios no es bueno, realizamos otro basado en el genero, tags, publisher y developer. Como menciona el autor, esta es una solución bastante pobre pero en los resultados se ve que funciona bastante bien.
# La idea es implementar un analisis de texto, y el texto es el resultado de la concatenacion del género, publisher, developer y tags.
# + colab={"base_uri": "https://localhost:8080/", "height": 602} colab_type="code" id="CsnX-wlwweVz" outputId="eaf8f91f-6dd9-4591-bd81-88101ae3a9f1"
games = pd.read_json("./data/steam/games.json.gz")
games.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 355} colab_type="code" id="KTXgDe8Oh6nq" outputId="cde616f1-43d0-4411-b2bf-cc1bd6bfefbb"
games.info()
# + colab={} colab_type="code" id="4pJw-bSJiA-H"
from ast import literal_eval
smd = pd.DataFrame()
smd['title'] = games['title']
smd['publisher'] = games['publisher'].astype(str).fillna('').apply(lambda x: str.lower(x.replace(" ", ""))).apply(lambda x: str.lower(x.replace("none", "")))
smd['genres'] = games['genres'].fillna('[]').astype(str)
smd['tags'] = games['tags'].fillna('[]').astype(str)
smd['developer'] = games['developer'].astype(str).fillna('').apply(lambda x: str.lower(x.replace(" ", ""))).apply(lambda x: str.lower(x.replace("none", "")))
# + colab={"base_uri": "https://localhost:8080/", "height": 399} colab_type="code" id="97MOB-sQj6Y_" outputId="9d37b35a-484b-49a2-df14-ebcfd44cbd50"
smd
# + colab={} colab_type="code" id="GIvWNfArpl1m"
smd['soup'] = smd['developer'] + smd['tags'] + smd['genres'] + smd['publisher']
smd['soup'] = smd['soup'].apply(lambda x: ''.join(x))
# + colab={"base_uri": "https://localhost:8080/", "height": 230} colab_type="code" id="BVELQTGoqyi6" outputId="9b765fda-10da-4241-b2b2-98332d71de86"
smd['soup']
# + colab={} colab_type="code" id="Trhv4aiTrk-3"
count = CountVectorizer(analyzer='word',ngram_range=(1, 2),min_df=0, stop_words='english')
count_matrix = count.fit_transform(smd['soup'])
# + colab={} colab_type="code" id="EQDNGGZUrnyP"
cosine_sim = cosine_similarity(count_matrix, count_matrix)
# + colab={} colab_type="code" id="WEx4D3-psDKh"
smd = smd.reset_index()
titles = smd['title']
indices = pd.Series(smd.index, index=smd['title'])
# + colab={} colab_type="code" id="-4cPiZsrvLSR"
def get_recommendations(id, titles, indices):
print(id)
idx = indices[id]
print(idx)
sim_scores = list(enumerate(cosine_sim[idx]))
sim_scores = sorted(sim_scores, key=lambda x: x[1], reverse=True)
sim_scores = sim_scores[1:31]
product_indices = [i[0] for i in sim_scores]
games_id_list = list(titles.iloc[product_indices].head(10))
return games_id_list
# + colab={"base_uri": "https://localhost:8080/", "height": 230} colab_type="code" id="dC-CODL7sCe3" outputId="cc60c520-a3e4-4929-8ccc-61739929acc0"
get_recommendations('Counter-Strike', titles, indices)
# + colab={"base_uri": "https://localhost:8080/", "height": 230} colab_type="code" id="w83uERhnblCO" outputId="cbf2c463-2fe7-43af-e851-194d99eebe0f"
get_recommendations('Need For Speed: Hot Pursuit', titles, indices)
# + colab={} colab_type="code" id="xu3pe4HLxLiG"
| SteamRecommendationSystem.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/probml/pyprobml/blob/master/notebooks/intro/jax.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="E4bE-S8yDALH" colab_type="text"
# # JAX <a class="anchor" id="jax"></a>
#
# [JAX](https://github.com/google/jax) is a version of Numpy that runs fast on CPU, GPU and TPU.
# In addition to having a fast backend, JAX supports
# several useful python-level program transformations:
#
# * [vmap](#vmap), vectorized map operator for automatic vectorization or batching.
# * [autograd](#AD), for automatic differentiation.
# * [jit](#jit), just in time compiler for speeding up your code (even on a CPU!).
#
# We illustrate these below.
#
# More details can be found at the
# * [official JAX quickstart page](https://github.com/google/jax#quickstart-colab-in-the-cloud).
# * ["You don't know jax"](https://colinraffel.com/blog/you-don-t-know-jax.html), blog post by <NAME>
#
#
# Other relevant PyProbML notebooks:
# * [Autodiff in JAX](https://github.com/probml/pyprobml/blob/master/notebooks/opt/opt.ipynb#AD-jax)
# * [SGD using JAX](https://github.com/probml/pyprobml/blob/master/notebooks/opt/opt.ipynb#SGD-jax)
# * [Linear algebra](https://github.com/probml/pyprobml/blob/master/notebooks/linalg/linalg.ipynb)
#
# Various libraries build on top of JAX, including the following:
# * [Stax](https://github.com/google/jax/blob/master/jax/experimental/stax.py), a DSL for DNNs, which provides an API which is similar to Keras or PyTorch.
# * [Trax](https://github.com/tensorflow/tensor2tensor/tree/master/tensor2tensor/trax), similar to Stax.
# * [Optim](https://github.com/google/jax/blob/master/jax/experimental/optimizers.py), a small library of fast first-order optimization methods available
# * [NumPyro](https://github.com/pyro-ppl/numpyro), a JAX backend for [Pyro](https://github.com/pyro-ppl/pyro) probabilistic modeling framework.
#
#
#
#
# **Make sure you select 'GPU' from the 'Runtime' tab at the top of this page.**
#
# + id="eCI0G3tfDFSs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 291} outputId="4598fd06-7ac2-4913-873d-525d13e0bf3e"
# Standard Python libraries
from __future__ import absolute_import, division, print_function, unicode_literals
import os
import time
import numpy as np
np.set_printoptions(precision=3)
import glob
import matplotlib.pyplot as plt
import PIL
import imageio
from IPython import display
# %matplotlib inline
import sklearn
import seaborn as sns;
sns.set(style="ticks", color_codes=True)
import pandas as pd
pd.set_option('precision', 2) # 2 decimal places
pd.set_option('display.max_rows', 20)
pd.set_option('display.max_columns', 30)
pd.set_option('display.width', 100) # wide windows
# Check we can plot stuff
plt.figure()
plt.plot(range(10))
# + id="BlTTyQuJGtmf" colab_type="code" colab={}
# Install
# !pip install --upgrade -q https://storage.googleapis.com/jax-releases/cuda$(echo $CUDA_VERSION | sed -e 's/\.//' -e 's/\..*//')/jaxlib-$(pip search jaxlib | grep -oP '[0-9\.]+' | head -n 1)-cp36-none-linux_x86_64.whl
# !pip install --upgrade -q jax
# + id="Z9kAsUWYDIOk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 69} outputId="d53d1eaa-7e3d-400d-9905-5f0f447e9eb0"
# Load libraries
import jax
import jax.numpy as np
print(np.zeros((3,3))) # make sure it runs
import numpy as onp # original numpy
from jax import grad, hessian, jit, vmap
# + [markdown] id="0Zsh5DdOF4R1" colab_type="text"
# ### Vmap <a class="anchor" id="vmap"></a>
#
#
# To illustrate vmap, consider a logistic regression model.
# + id="5XAMcxMsF0-Q" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="5744be81-df29-4d85-ac35-3b1f5adbff10"
def sigmoid(x): return 0.5 * (np.tanh(x / 2.) + 1)
def predict_single(w, x):
return sigmoid(np.dot(w, x)) # <(D) , (D)> = (1) # inner product
def predict_batch(w, X):
return sigmoid(np.dot(X, w)) # (N,D) * (D,1) = (N,1) # matrix-vector multiply
D = 2
N = 3
onp.random.seed(42)
w = onp.random.randn(D)
X = onp.random.randn(N, D)
y = onp.random.randint(0, 2, N)
# We can apply predict_batch to a matrix of data, but we cannot apply predict_single in this way
# because the order of the arguments to np.dot is incorrect.
p1 = predict_batch(w, X)
try:
p2 = predict_single(w, X)
except:
print('cannot apply to batch')
# + [markdown] id="S-qWaSNBGIqg" colab_type="text"
# To avoid having to think about batch shape, it is often easier to write a function that works on single
# input vectors. We can then apply this in a loop.
# + id="VFqzy2ZFF7Fc" colab_type="code" colab={}
p3 = [predict_single(w, x) for x in X]
assert np.allclose(p1, p3)
# + [markdown] id="ZZSksj-1GZcU" colab_type="text"
# Unfortunately, mapping down a list is slow.
# Fortunately, JAX provides `vmap`, which has the same effect, but can be parallelized.
#
# We first apply the `predict_single` function to its first arugment, w, to get a function that only
# depends on x. We then vectorize this, and map the resulting modified function along rows (dimension 0)
# of the data matrix.
# + id="EMNkPb9GGZ1O" colab_type="code" colab={}
from functools import partial
predict_single_w = partial(predict_single, w)
predict_batch_w = vmap(predict_single_w)
p4 = predict_batch_w(X)
p5 = vmap(predict_single, in_axes=(None, 0))(w, X)
assert np.allclose(p1, p4)
assert np.allclose(p1, p5)
# + [markdown] id="qA2RYhnNG9hh" colab_type="text"
# ### Autograd <a class="anchor" id="AD"></a>
#
# In this section, we illustrate automatic differentiation by using it to compute the gradient of the negative log likelihood of a logistic regression model for each example.
# + id="Isql2l4MGfIt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 87} outputId="618c710e-857b-4d02-f544-427e66724dd6"
def predict(weights, inputs):
return sigmoid(np.dot(inputs, weights))
def loss(weights, inputs, targets):
preds = predict(weights, inputs)
logprobs = np.log(preds) * targets + np.log(1 - preds) * (1 - targets)
return -np.sum(logprobs)
def loss2(params, data):
return loss(params, data[0], data[1])
print(loss(w, X, y))
assert np.isclose(loss(w, X, y), loss2(w, (X,y)))
# Gradient function
grad_fun = grad(loss)
# Gradient of each example in the batch
grads = vmap(partial(grad_fun, w))(X,y)
print(grads)
assert grads.shape == (N,D)
grads2 = vmap(grad_fun, in_axes=(None, 0, 0))(w, X, y)
assert np.allclose(grads, grads2)
grads3 = vmap(grad(loss2), in_axes=(None, 0))(w, (X, y))
assert np.allclose(grads, grads3)
# Gradient for entire batch
grad_sum = np.sum(grads, axis=0)
assert grad_sum.shape == (D,)
# + id="CGxDFho3H5ou" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 69} outputId="a8b08446-2a77-4dbc-bbb9-14f1c3fdbad0"
# We can We can also compute Hessians, as we illustrate below.
from jax import hessian
H0 = hessian(loss)(w, X[0,:], y[0])
print(H0)
hessian_fun = hessian(loss)
H = vmap(hessian_fun, in_axes=(None, 0, 0))(w, X, y)
print(H.shape)
# + [markdown] id="GnOIJRGxJigp" colab_type="text"
#
# ### JIT (just in time compilation) <a class="anchor" id="JIT"></a>
#
# In this section, we illustrate how to use the Jax JIT compiler to make code go faster (even on a CPU). However, it does not work on arbitrary Python code, as we explain below.
#
#
#
# + id="qsaHkNovICfd" colab_type="code" colab={}
grad_fun_jit = jit(grad_fun) # speedup gradient function
grads_jit = vmap(partial(grad_fun_jit, w))(X,y)
assert np.allclose(grads, grads_jit)
# + id="GGZUbPDLKIkv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 69} outputId="4192101c-2c6a-4019-d939-4d358fa34507"
# We can apply JIT to non ML applications as well.
def slow_f(x):
# Element-wise ops see a large benefit from fusion
return x * x + x * 2.0
x = np.ones((5000, 5000))
fast_f = jit(slow_f)
# %timeit -n10 -r3 fast_f(x)
# %timeit -n10 -r3 slow_f(x)
# + [markdown] id="77-33YadKNni" colab_type="text"
# We can also add the `%jit` decorator in front of a function.
#
# Note that JIT compilation requires that the control flow through the function can be determined by the shape (but not concrete value) of its inputs. The function below violates this, since when x<3, it takes one branch, whereas when x>= 3, it takes the other.
# + id="6Ps1W8LhKKj9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="560b98e7-ff00-43af-c7f8-9ea28c91fb47"
@jit
def f(x):
if x < 3:
return 3. * x ** 2
else:
return -4 * x
# This will fail!
try:
print(f(2))
except Exception as e:
print("ERROR:", e)
# + [markdown] id="8tHh4tcXKTRf" colab_type="text"
# We can fix this by telling JAX to trace the control flow through the function using concrete values of some of its arguments. JAX will then compile different versions, depending on the input values. See below for an example.
#
# + id="DMaRplccKRHc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="7e1150b3-a530-4dcc-cc5c-418b4792219d"
def f(x):
if x < 3:
return 3. * x ** 2
else:
return -4 * x
f = jit(f, static_argnums=(0,))
print(f(2.))
# + [markdown] id="Q1KDjxB7KXwi" colab_type="text"
# One solution to this is to use `lax.fori_loop`, which can compile just a single version of the function. Unfortunately, you cannot apply `grad` to code that uses this construct; instead, it is typically used inside an optimization loop, where the inner part contains the gradient, and the outer (non differentiable) part just iterates until convergence.
# + id="pDIOlLcBKVVE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="04998507-1985-4488-e053-63de1cca0f46"
from jax import lax
init_val = 0
start = 0
stop = 10
body_fun = lambda i,x: x+i
y = lax.fori_loop(start, stop, body_fun, init_val)
print(y)
# + [markdown] id="w2zrYp3-KcMc" colab_type="text"
# There are a few other subtleties. If your function has global side-effects, JAX's tracer can cause weird things to happen. A common gotcha is trying to print arrays inside jit'd functions:
# + id="ZG5HQ2dnKZ67" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 122} outputId="e4399277-07a7-49cf-eae2-465f737cd5ba"
def f(x):
print(x)
y = 2 * x
print(y)
return y
y1 = f(2)
@jit
def f(x):
print(x)
y = 2 * x
print(y)
return y
y2 = f(2)
print(y1)
print(y2)
# + [markdown] id="UZ0T4PxnKgBN" colab_type="text"
# ## A few differences from Numpy
#
# Below we list a few items where Jax differs from Numpy.
# See also the official [list of common gotchas](https://colab.research.google.com/github/google/jax/blob/master/notebooks/Common_Gotchas_in_JAX.ipynb).
# + [markdown] id="MUOZdeYBKjWc" colab_type="text"
# ### Random number generation
#
# The API for Jax is basically identical to Numpy, except for pseudo random number
# generation (PRNG).
# This is because Jax does not maintain any global state, i.e., it is purely functional.
# This design "provides reproducible results invariant to compilation boundaries and backends,
# while also maximizing performance by enabling vectorized generation and parallelization across random calls"
# (to quote [the official page](https://github.com/google/jax#a-brief-tour)).
#
# Thus, whenever we do anything stochastic, we need to give it a fresh RNG key. We can do this by splitting the existing key into pieces. We can do this indefinitely, as shown below.
# + id="dcTYfznjKeHC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 104} outputId="018242a2-ce96-45a9-9078-0a33dcc07017"
import jax.random as random
key = random.PRNGKey(0)
print(random.normal(key, shape=(3,))) # [ 1.81608593 -0.48262325 0.33988902]
print(random.normal(key, shape=(3,))) # [ 1.81608593 -0.48262325 0.33988902] ## identical results
# To make a new key, we split the current key into two pieces.
key, subkey = random.split(key)
print(random.normal(subkey, shape=(3,))) # [ 1.1378783 -1.22095478 -0.59153646]
# We can continue to split off new pieces from the global key.
key, subkey = random.split(key)
print(random.normal(subkey, shape=(3,))) # [-0.06607265 0.16676566 1.17800343]
# We can always use original numpy if we like (although this may interfere with the deterministic behavior of jax)
onp.random.seed(42)
print(onp.random.randn(3))
# + [markdown] id="xwmI9DH2K_nl" colab_type="text"
# ### Implicitly casting lists to vectors
#
# You cannot treat a list of numbers as a vector. Instead you must explicitly create the vector using the np.array() constructor.
#
# + id="bUURw01jKnXC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 315} outputId="ebddc278-19e6-4287-8aea-c65a80e3fb71"
# You cannot treat a list of numbers as a vector.
S = np.diag([1.0, 2.0, 3.0])
# + id="2mMgaegpLCYw" colab_type="code" colab={}
# Instead you should explicitly construct the vector.
S = np.diag(np.array([1.0, 2.0, 3.0]))
# + [markdown] id="AieugSOZLGi5" colab_type="text"
# ### Mutation of arrays
#
# Since JAX is functional, you cannot mutate arrays in place,
# since this makes program analysis and transformation very difficult. JAX requires a pure functional expression of a numerical program.
# Instead, JAX offers the functional update functions: `index_update`, `index_add`, `index_min`, `index_max`, and the `index` helper. These are illustrated below.
#
# Note: If the input values of `index_update` aren't reused, jit-compiled code will perform these operations in-place, rather than making a copy.
#
# + id="GEKfhvrTLEBf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 346} outputId="7694ce2b-8813-4c60-ea28-ca0e5bf6e844"
# You cannot assign directly to elements of an array.
jax_array = np.zeros((3,3), dtype=np.float32)
# In place update of JAX's array will yield an error!
jax_array[1, :] = 1.0
# + id="CBi15EAcLIru" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 297} outputId="0546aab4-12bb-4aa9-d983-8744079b78ca"
from jax.ops import index, index_add, index_update
jax_array = np.zeros((3, 3))
print("original array:")
print(jax_array)
new_jax_array = index_update(jax_array, index[1, :], 1.)
new_jax_array2 = index_add(new_jax_array, index[:, 2], 7.)
print("new array post update")
print(new_jax_array)
print("new array post add")
print(new_jax_array2)
print("old array unchanged:")
print(jax_array)
# + id="q7U9Ee15LLC-" colab_type="code" colab={}
| notebooks/intro/jax.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # The relationship between Zn and diabetes
# - Author: <NAME>
# - Date: 7 Oct
#
# ### Key variable description
#
# #### Response
# **DIABBC** is a `categorial variable`, the value has different meaning:
# - 1: Ever told has diabetes mellitus, still current and long term
# - 3: Ever told has diabetes mellitus, not current
# - 5: Never told has diabetes mellitus
#
#
# #### Predictor
# - **ZINCT1** Zinc (total) Day 1 mg
# - (for all person)
# - **ZINCT2** Zinc (total) Day 2 mg
# - (for all persons who completed second (CATI) nutrition interview)
# - **SEX**
# - 1: Male
# - 2: Female
#
# ## Findings:
# 1. **The mean of zn in the diabetes group (DIABBC = 1) is lower than that in the health group (DIABBC = 1)**, Confirm by:
# - Distrubtion visualization
# - Hypothesis testing on the group mean with 5% confidence level
#
# 2. No clear difference between Zn and sex/gender
#
# ## Questions:
# See below.
#
#
import pandas as pd
import pyspark
from datetime import datetime
import os.path
import datetime
pd.set_option('display.max_columns', 500)
import seaborn as sns
import numpy as np
import calendar
import matplotlib
import matplotlib.pyplot as plt
import warnings
warnings.filterwarnings('ignore')
from sklearn.ensemble import RandomForestClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.svm import SVC, SVR
from sklearn.model_selection import cross_val_predict
from sklearn.metrics import confusion_matrix
from sklearn.pipeline import Pipeline
from sklearn.model_selection import train_test_split
from sklearn import metrics
from sklearn.externals import joblib
from scipy.stats import norm, skew
import statsmodels.api as sm
blue = sns.color_palette('Blues')[-2]
color = sns.color_palette()
sns.set_style('darkgrid')
pd.set_option('display.float_format', lambda x: '{:.3f}'.format(x)) #Limiting floats output to 3 decimal points
import nltk
from sklearn.model_selection import RandomizedSearchCV, GridSearchCV, StratifiedKFold
# #### Load and join dataset
merged_food_nutr_binom = pd.read_csv('data/merged_food_nutr_binom.csv')
merged_food_nutr_binom.head()
# ### Relationship between ZINCT1 and ZINCT2
selected_col = ["DIABBC", "ZINCT1","ZINCT2", "AGEC","SEX", "INCDEC"]
data = merged_food_nutr_binom
### heatmap on selected columns
corr = data[selected_col].corr()
fig, ax = plt.subplots(figsize=(10,5))
sns.heatmap(corr, annot=True,
xticklabels=corr.columns.values,
yticklabels=corr.columns.values,ax=ax)
# ### Distribution of Zn conditional on Diabbc
ax = sns.displot(merged_food_nutr_binom, x="ZINCT1", hue="DIABBC")
ax.fig.suptitle('Distribution of Zn1 conditional on Diabbc', fontsize=20)
# From plot above, we can see that the dist of ZINCT1 are right-skewed, thus we do transformation
ax = sns.displot(merged_food_nutr_binom, x="ZINCT2", hue="DIABBC")
ax.fig.suptitle('Distribution of Zn2 conditional on Diabbc', fontsize=20)
# From the plot above we can see that, ZINCT2 has lots of missing value (0) as not all people attend the day 2 testing, thus we create another group removing rows containing ZINCT2 = 0
new_data_ZINCT2 = merged_food_nutr_binom[merged_food_nutr_binom['ZINCT2'] > 0] # remove the rows with missing value
new_data_ZINCT1 = merged_food_nutr_binom[merged_food_nutr_binom['ZINCT1'] > 0] # remove the rows with missing value
new_data_ZINCT3 = new_data_ZINCT1[new_data_ZINCT1['ZINCT2'] > 0] # remove the rows with missing value
# +
merged_food_nutr_binom['log_ZINCT1'] = np.log(merged_food_nutr_binom['ZINCT1']+1)
merged_food_nutr_binom['log_ZINCT2'] = np.log(merged_food_nutr_binom['ZINCT2']+1)
new_data_ZINCT1['log_ZINCT1'] = np.log(new_data_ZINCT1['ZINCT1'])
#new_data_ZINCT1['log_ZINCT2'] = np.log(new_data_ZINCT1['ZINCT2'])
#new_data_ZINCT2['log_ZINCT1'] = np.log(new_data_ZINCT2['ZINCT1'])
new_data_ZINCT2['log_ZINCT2'] = np.log(new_data_ZINCT2['ZINCT2'])
new_data_ZINCT3['log_ZINCT1'] = np.log(new_data_ZINCT3['ZINCT1'])
new_data_ZINCT3['log_ZINCT2'] = np.log(new_data_ZINCT3['ZINCT2'])
# -
ax = sns.displot(merged_food_nutr_binom, x="log_ZINCT1", hue="DIABBC")
ax.fig.suptitle('Distribution of LOG_Zn conditional on Diabbc', fontsize=20)
ax = sns.displot(new_data_ZINCT2, x="log_ZINCT2", hue="DIABBC")
ax.fig.suptitle('Distribution of LOG_Zn conditional on Diabbc', fontsize=20)
# From plot above, we can see that the distribution of zn in the healthy group (Diabbc = 5) looks quite different from that of diabete gruop (Diabbc = 1), mainly since that the we have much more observations of group (Diabbc = 5) than that of group (Diabbc = 1).
#
# So it is hard to fingure out is the mean of zn in these groups are different. So let's try hypothesis testing!
# ### T test on the mean of Zn (healthy group (Diabbc = 5) VS diabete gruop (Diabbc = 1))
#
# > T- Test :- A t-test is a type of inferential statistic which is used to determine if there is a significant difference between the means of two groups which may be related in certain features.
#
# > Two sampled T-test :-The Independent Samples t Test or 2-sample t-test compares the means of two independent groups in order to determine whether there is statistical evidence that the associated population means are significantly different.
#
# - H0: The mean of Zn in healthy group (Diabbc = 5) and diabete gruop (Diabbc = 1) are the same
# - H1: The mean of Zn in two groups are different
# - **Our testing is based on the data removing the missing value**
# - If the p_value < 0.05, it means that the mean of two groups are different
# - From the values below, we can see that the mean of two groups are statistically different since (p_value < 0.05)
# +
from scipy.stats import ttest_ind
diabetes_group = new_data_ZINCT1[new_data_ZINCT1['DIABBC']==1]
healthy_group = new_data_ZINCT1[new_data_ZINCT1['DIABBC']==5]
diabetes_group2 = new_data_ZINCT2[new_data_ZINCT2['DIABBC']==1]
healthy_group2 = new_data_ZINCT2[new_data_ZINCT2['DIABBC']==5]
diabetes_group3 = new_data_ZINCT3[new_data_ZINCT3['DIABBC']==1]
healthy_group3 = new_data_ZINCT3[new_data_ZINCT3['DIABBC']==5]
print("Mean of healthy group of ZINCT1:", np.mean(healthy_group['ZINCT1']))
print("Mean of diabetes group of ZINCT1:", np.mean(diabetes_group['ZINCT1']))
print("T test on the ZINCT1:")
print(ttest_ind(diabetes_group['ZINCT1'], healthy_group['ZINCT1']))
print(ttest_ind(diabetes_group3['ZINCT1'], healthy_group3['ZINCT1']))
print("\nMean of healthy group of ZINCT2:", np.mean(healthy_group2['ZINCT2']))
print("Mean of diabetes group of ZINCT2:", np.mean(diabetes_group2['ZINCT2']))
print("T test on the ZINCT2:")
print(ttest_ind(diabetes_group2['ZINCT2'], healthy_group2['ZINCT2']))
print(ttest_ind(diabetes_group3['ZINCT2'], healthy_group3['ZINCT2']))
# -
# ### But questions:
# #### Q1: should the mean of Zn in healthy group larger than the mean of Zn in diabetes group?
# in the data.
# Mean of healthy group of ZINCT1 (12.978)< Mean of diabetes group of ZINCT1 (13.050)
# but
# Mean of healthy group of ZINCT2 (12.102)> Mean of diabetes group of ZINCT2 (11.766)
# **Which variable (ZINCT1 or ZINCT2) is realiable? or both correct?**
#
# #### Q2: the difference of the group mean above looks not large, is it significant in the real world?
#
# ### Distribution of Zn conditional on Diabbc and Sex
#
# **From the plot below, no clear difference between zn and sex**
g = sns.FacetGrid(new_data_ZINCT1, col="DIABBC", row="SEX")
g.map_dataframe(sns.histplot, x="ZINCT1")
g = sns.FacetGrid(new_data_ZINCT1, col='DIABBC', row="SEX")
g.map_dataframe(sns.histplot, x="log_ZINCT1")
g = sns.FacetGrid(new_data_ZINCT2, col="DIABBC", row="SEX")
g.map_dataframe(sns.histplot, x="ZINCT1")
# ### Relationship between ZINCT1 and ZINCT2
# the plot below are the one remove missing values
g = sns.FacetGrid(new_data_ZINCT2, col='DIABBC', hue="SEX")
g.map_dataframe(sns.scatterplot, x="ZINCT1", y="ZINCT2")
g.set_axis_labels(" Zinc (total) Day 1 mg", " Zinc (total) Day 2 mg")
g.add_legend()
# From the plot above, we can see some outliers in the female group (sex = 2)
#
# ### Question:
# Are the outliers in the data are mis-recorded?? (See some points are moe than 50 mg)
# ### Combine ZINCT1 and ZINCT2
new_data_ZINCT3['mean_ZINCT'] = (new_data_ZINCT3['ZINCT1'] + new_data_ZINCT3['ZINCT2'])/2
merged_food_nutr_binom.to_csv('data/merged_food_nutr_binom_7Oct.csv', index=False)
| Zn_stat_check_7Oct.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 对应剑指offer第三章-高质量的代码
# 1. 规范性
# >* 书写清晰
# >* 布局合理
# >* 命名合理
# 2. 完整性
# >* 完成基本功能
# >* 考虑边界条件
# >* 做好错误处理
# 1. 鲁棒性
# >* 采取防御性编程
# >* 处理无效输入
# # 16 数值的整数次方
# 1. 位运算的适用范围仅仅是整数
# 2. 递归算法适用负数
# +
# 位运算
int pow_test(int m, int n)
{
int sum = 1;
int temp = m;
while (n)
{
if (n & 1 == 1)
sum *= temp;
n=n >> 1;
temp *= temp;
}
return sum ;
}
# 递归
double pow_try(double m, int n)
{
if (n == 0)
return 1.0;
if (n == 1)
return m;
double base = pow_try(m,n>>1);
base *= base;
if (n & 1 == 1)
base *= m;
return base;
}
# -
# # 17 打印 1-最大的n位数
# 难点:大数--需要将数转化为**字符串**或者**数组**
# # 18 删除连续的重复数-链表
void deleteDuplication(ListNode** pHead)
{
if (pHead == nullptr || *pHead == nullptr)
return;
ListNode* pPre = nullptr;
ListNode* pCur = *pHead;
while (pCur != nullptr)
{
ListNode* pNex = pCur->n_next;
bool needDelete = false;
if (pNex != nullptr&&pCur->n_value == pNex->n_value)
needDelete = true;
if (!needDelete)
{
pPre = pCur;
pCur = pNex;
}
else
{
int value = pCur->n_value;
ListNode* pTobeDel = pCur;
while (pTobeDel != nullptr&&pTobeDel->n_value == value)
{
pNex = pTobeDel->n_next;
delete pTobeDel;
pTobeDel = nullptr;
pTobeDel = pNex;
}
// 若指针头就是重复,且被删除,后面无后续数字,则
if (pPre == nullptr)
*pHead = pNex;
else
pPre->n_next = pNex;
pCur = pNex;
}
}
}
# # 21 调整数组,使奇数在前,偶数在后
# * 借鉴快排的方法
# * 可以将__(a[i] & 1) == 1__转换成一个函数__fun(n)__,增强其泛化能力
void changearray(vector<int> &a)
{
//int tmp = a[0];
int i = 0;
int j = a.size() - 1;
while (i <= j)
{
while ((a[i] & 1) == 1) { i++; }
while ((a[j] & 1) == 0) { j--; }
if (i <= j)
{
swap(a[i], a[j]);
cout << "i is " << a[i] << ' ' << "j is " << a[j] << "\n";
i++;
j--;
}
}
}
# # 22 链表中倒数第K个节点
# * 双指针
# * 第一个指针到(k-1)时,第二个与第一个开始同步前进
# * 增加测试样例,提高鲁棒性
ListNode* FindKthNode(ListNode*pListHead, int k)
{
if (pListHead == NULL || k == 0)
return NULL;
ListNode* ANode = pListHead;
ListNode* BNode = ANode;
for (int i = 0;i < k - 1;i++)
ANode = ANode->n_next;
while (ANode->n_next)
{
ANode = ANode->n_next;
BNode = BNode->n_next;
}
return BNode;
}
# # 23 链表中*环*的入口节点
# 1. 先判断是否有环:双指针,一快一慢,快的追上慢的,则有环
# 2. 找入口:双指针,第一个比第二个多环的节点数n,从链表头部开始循环
# # 24 反转链表
ListNode* ReverseList(ListNode* pHead)
{
// 定义三个循环节点
ListNode* pre;
ListNode* curr;
ListNode* follow;
// 初始化
pre = pHead;
curr = pHead->n_next;
follow = curr->n_next;
// 循环,每次改变一个节点的指针方向
pHead->n_next = NULL;
while (follow!=NULL)
{
curr->n_next = pre;
pre = curr;
curr = follow;
follow = follow->n_next;
}
curr->n_next = pre;
}
# # 25 合并两个排序好的链表
ListNode*Merge(ListNode* pHead1, ListNode* pHead2)
{
if (pHead1 == NULL)
return pHead2;
if (pHead2 == NULL)
return pHead1;
ListNode* MergedHead = NULL;
if (pHead1->n_next < pHead2->n_next)
{
MergedHead = pHead1;
MergedHead->n_next = Merge(pHead1->n_next,pHead2);
}
else
{
MergedHead = pHead2;
MergedHead->n_next = Merge(pHead1,pHead2->n_next);
}
return MergedHead;
}
# # 26 判断B是否是A的子结构-树
# * 子树不单单只有两层
# * 若树的值不是__int__型,则不能直接用等号判断大小
# +
bool a_include_b(BinaryTree* father, BinaryTree* son)
{
if (son == nullptr)
return true;
if (father == nullptr)
return false;
if (father->p_value != son->p_value)
return false;
return a_include_b(father->p_left, son->p_left) && a_include_b(father->p_right, son->p_right);
}
bool SonTree(BinaryTree* father, BinaryTree* son)
{
bool result = false;
if (father != nullptr &&son != nullptr)
{
if (father->p_value == son->p_value)
result = a_include_b(father, son);
if (!result)
result = SonTree(father->p_left, son);
if(!result)
result = SonTree(father->p_right, son);
}
return result;
}
| Lead_To_Offer_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !pip install simpletransformers
# +
import random
import json
#eng = '/home/jovyan/data/sbersquad_train_clean_final_translated.json'
eng = '/home/jovyan/data/squad/train-v2.0.json'
rus = '/home/jovyan/data/sberquad/sbersquad_train_clean_final.json'
end_data = json.load(open(eng, 'r'))
rus_data = json.load(open(rus, 'r'))
# -
rus_data['paragraphs'][0]
len(end_data['data'])
len(end_data['data'][0])
end_data = [end_data['data'][i]['paragraphs'] for i in range(len(end_data['data']))]
end_data = [item for sublist in end_data for item in sublist]
len(end_data)
"""cross_ling = []
for i in range(len(end_data['paragraphs'])):
cross_ling.append(end_data['paragraphs'][i])
new = end_data['paragraphs'][i]
new['qas'] = rus_data['paragraphs'][i]['qas']
#cross_ling.append(new)
for i in range(len(rus_data['paragraphs'])):
cross_ling.append(rus_data['paragraphs'][i])
new = rus_data['paragraphs'][i]
new['qas'] = end_data['paragraphs'][i]['qas']
#cross_ling.append(new)
random.shuffle(cross_ling)"""
cross_ling = rus_data['paragraphs']+end_data
random.shuffle(cross_ling)
cross_ling[0]
len(cross_ling)
len(new_cross_ling)
from simpletransformers.question_answering import QuestionAnsweringModel
# !mkdir /home/jovyan/data/huawei/simple
# +
train_args = {
"reprocess_input_data": True,
"overwrite_output_dir": True,
#"special_tokens_list":['<s>', '</s>', 'lang:', 'question:', 'answer:'],
"output_dir" : '/home/jovyan/data/huawei/simple'
}
model = QuestionAnsweringModel(
"bert",
"DeepPavlov/rubert-base-cased",
args=train_args,
)
model.train_model(cross_ling[:15000], eval_data=cross_ling[15000:])
# -
# Evaluate the model
result, texts = model.eval_model(cross_ling[15000:])
text = '''Как передает RegioNews, об этом сообщил первый заместитель директора Одесского припортового завода <NAME>.
«Завод уже в процессе остановки. На сегодня причины две — переполненный состав карбамида и заоблачная цена сырья (природного газа)», — сообщил он.
По его словам, «окно возможностей» для стабилизации работы завода закрылось. Как отмечает Щуриков, пока остается лишь шанс провести приватизацию завода.
Напомним, Одесский припортовый завод выпускал свою продукцию с 1978 года. Специализация предприятия — производство аммиака, карбамида и другой химической продукции. Предприятие возобновило свою работу два года назад после длительного простоя.
'''
test = [{'context':text, 'qas':[{'id':0, "question":'What are two main reasons of the plant shutdown?'}]}]
answers, probabilities = model.predict(test)
answers
import pandas as pd
test_en_en = pd.read_csv('./data/huawei/answers/sbersquad_dev_en_en.csv', sep='\t')
test_en_en.head()
test_en_en_predict = []
j = 0
for i in range(len(test_en_en)):
dic = {'context':test_en_en.iloc[i]['text+question'].split('question:')[0], 'qas':[{"id":j, "question":test_en_en.iloc[i]['text+question'].split('question:')[1]}]}
test_en_en_predict.append(dic)
j+=1
len(test_en_en)
answers, probabilities = model.predict(test_en_en_predict)
def gen_answer(answ):
a = answ['answer'][0]
if a in ['', ' ']:
a = answ['answer'][1]
return a
# +
test_en_en['generated_answers'] = [gen_answer(i) for i in answers]
# -
test_en_en.head()
# !mkdir ./data/huawei/answers_rubert
test_en_en.to_csv('./data/huawei/answers_rubert/squad_dev_en_en.csv', sep='\t')
answers[0]
test_en_rus = pd.read_csv('./data/huawei/answers/sbersquad_dev_en_rus.csv', sep='\t')
test_en_rus_predict = []
j = 0
for i in range(len(test_en_rus)):
#dic = {context:test_en_rus.iloc[i][], 'qas':[{"id":0, "question":test_en_rus.iloc[i][]}]}
dic = {'context':test_en_rus.iloc[i]['text+question'].split('question:')[0], 'qas':[{"id":j, "question":test_en_rus.iloc[i]['text+question'].split('question:')[1]}]}
test_en_rus_predict.append(dic)
j+=1
answers, probabilities = model.predict(test_en_rus_predict)
test_en_rus['generated_answers'] = [gen_answer(i) for i in answers]
test_en_rus.head()
test_en_rus.to_csv('./data/huawei/answers_rubert/squad_dev_en_rus.csv', sep='\t')
test_rus_rus = pd.read_csv('./data/huawei/answers/sbersquad_dev_rus_rus.csv', sep='\t')
test_rus_rus_predict = []
j = 0
for i in range(len(test_rus_rus)):
#dic = {context:test_rus_rus.iloc[i][], 'qas':[{"id":0, "question":test_rus_rus.iloc[i][]}]}
dic = {'context':test_rus_rus.iloc[i]['text+question'].split('question:')[0], 'qas':[{"id":j, "question":test_rus_rus.iloc[i]['text+question'].split('question:')[1]}]}
test_rus_rus_predict.append(dic)
j+=1
answers, probabilities = model.predict(test_rus_rus_predict)
test_rus_rus['generated_answers'] = [gen_answer(i) for i in answers]
test_rus_rus['generated_answers'] = [gen_answer(i) for i in answers]
test_rus_rus.head()
test_rus_rus.to_csv('./data/huawei/answers_rubert/squad_dev_rus_rus.csv', sep='\t')
test_rus_en = pd.read_csv('./data/huawei/answers/sbersquad_dev_rus_en.csv', sep='\t')
test_rus_en_predict = []
j = 0
for i in range(len(test_rus_en)):
#dic = {context:test_rus_en.iloc[i][], 'qas':[{"id":0, "question":test_rus_en.iloc[i][]}]}
dic = {'context':test_rus_en.iloc[i]['text+question'].split('question:')[0], 'qas':[{"id":j, "question":test_rus_en.iloc[i]['text+question'].split('question:')[1]}]}
test_rus_en_predict.append(dic)
j+=1
answers, probabilities = model.predict(test_rus_en_predict)
test_rus_en['generated_answers'] = [gen_answer(i) for i in answers]
test_rus_en.head()
test_rus_en.to_csv('./data/huawei/answers_rubert/squad_dev_rus_en.csv', sep='\t')
# +
eng = '/home/jovyan/data/sbersquad_train_clean_final_translated.json'
#eng = '/home/jovyan/data/squad/train-v2.0.json'
rus = '/home/jovyan/data/sberquad/sbersquad_train_clean_final.json'
end_data = json.load(open(eng, 'r'))
rus_data = json.load(open(rus, 'r'))
# -
cross_ling = []
for i in range(len(end_data['paragraphs'])):
cross_ling.append(end_data['paragraphs'][i])
new = end_data['paragraphs'][i]
new['qas'] = rus_data['paragraphs'][i]['qas']
cross_ling.append(new)
for i in range(len(rus_data['paragraphs'])):
cross_ling.append(rus_data['paragraphs'][i])
new = rus_data['paragraphs'][i]
new['qas'] = end_data['paragraphs'][i]['qas']
cross_ling.append(new)
random.shuffle(cross_ling)
new_cross_ling = []
for i in cross_ling:
qas = i['qas']
con = i['context']
checked = []
for j in qas:
if j['answers'][0]['answer_start']<len(con):
checked.append(j)
else:
print(j['id'])
new_cross_ling.append({"context":i['context'], "qas":checked})
len(new_cross_ling)
# +
train_args = {
"reprocess_input_data": True,
"overwrite_output_dir": True,
#"special_tokens_list":['<s>', '</s>', 'lang:', 'question:', 'answer:'],
"output_dir" : '/home/jovyan/data/huawei/simple'
}
model = QuestionAnsweringModel(
"bert",
"bert-base-multilingual-cased",
args=train_args,
)
model.train_model(new_cross_ling[:16000])#, eval_data=new_cross_ling[30000:])
# -
test_en_en = pd.read_csv('./data/huawei/answers/sbersquad_dev_en_en.csv', sep='\t')
test_en_en.head()
| notebooks/rubert_tune_crosslingual_QA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/FairozaAmira/AI-programming-1-a/blob/master/Lecture08.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="BCXsi3NgIvdo" colab_type="text"
# # 第8回目の講義中の練習問題回答
#
# ## `for` ループ
#
# 1. `for`ループを使って、1 から 5 まで表示しなさい。
# + id="ODSHRdygIstJ" colab_type="code" outputId="805a98ca-ec69-48b2-c7de-9543d3f55d5f" colab={"base_uri": "https://localhost:8080/", "height": 100}
for N in [1, 2, 3, 4, 5]:
print(N)
# + [markdown] id="TjD-HxsyKk84" colab_type="text"
# 2. 1番目で書いたコードを一行で表示するように修正しなさい。
# + id="nlZmxX08K4_4" colab_type="code" outputId="85820ae5-288e-476a-b322-6781ec03c3ea" colab={"base_uri": "https://localhost:8080/", "height": 33}
for N in [1,2,3,4,5]:
print(N, end=' ')
# + [markdown] id="gGkZVYinK8eP" colab_type="text"
# 3. `range`を使って、1から5まで一行に表示しなさい。
# + id="ppSTR3UbLFVR" colab_type="code" outputId="77585293-e808-471c-9779-38472e20f1d9" colab={"base_uri": "https://localhost:8080/", "height": 33}
for N in range(1,6):
print(N, end=" ")
# + [markdown] id="_6u_e_8zLOYM" colab_type="text"
# `range`は普通0から始まる。`in range(start,end+1,step)`という意味。
# + [markdown] id="LjLaHmaZLbjz" colab_type="text"
# 4. 1 から 5 までの一つのリストを作りなさい。
# + id="XHApM3W5LmN8" colab_type="code" outputId="37c157be-6622-424a-ba98-81695ca368f2" colab={"base_uri": "https://localhost:8080/", "height": 33}
list(range(1,6))
# + [markdown] id="5x38uWcoLuUe" colab_type="text"
# 5. 3の倍数のNループを作りなさい。但し、Nは30より小さいまたは等しいとする。
# + id="385reuFlMAOE" colab_type="code" outputId="58f32c5e-71f5-4430-e47d-f8da4c43c02c" colab={"base_uri": "https://localhost:8080/", "height": 184}
for N in range(3,31,3):
print(N)
# + [markdown] id="o6aMTrSvMDdJ" colab_type="text"
# 6. 1から5までの合計を計算しなさい。
# + id="a9GqTNp_MMMS" colab_type="code" outputId="11eeb057-1c34-4b86-8250-9dea1658a796" colab={"base_uri": "https://localhost:8080/", "height": 33}
#totalを初期化する
total = 0
for N in range(1,6):
total += N #totalを毎回Nと加算させる
print(total)
# + [markdown] id="QKlfoScGMW4F" colab_type="text"
# 7. `places`リストを作り、`for`ループを使って、スペースを”-”と代入して、最初の文字が大文字にした`new_places`リストを作りなさい。
#
# `places = ["kamigyo ku","nakagyo ku", "sakyo ku"]`を<br/>
# `new_places=['Kamigyo-ku', 'Nakagyo-ku', 'Sakyo-ku']`にしなさい。
#
# + id="i-s7kMRaPate" colab_type="code" outputId="87100e7a-b5c0-4a1c-faeb-2f9dd914fe9c" colab={"base_uri": "https://localhost:8080/", "height": 33}
places = ["kamigyo ku","nakagyo ku", "sakyo ku"]
new_places = []
for place in places:
new_places.append(place.capitalize().replace(" ","-"))
print(new_places)
# + id="fK-FRbYNPdfq" colab_type="code" outputId="0aa69ae8-ab2b-4465-e2ba-84612dd09191" colab={"base_uri": "https://localhost:8080/", "height": 33}
new_places = ["kamigyo ku","nakagyo ku", "sakyo ku"]
for i in range(len(new_places)):
new_places[i] = new_places[i].capitalize().replace(" ", "-")
print(new_places)
# + [markdown] id="oLDdnhp4PgvF" colab_type="text"
# 8. かごの中にある果物の数を数えましょう。
#
# かごの辞書は下記通り。<br/>
# `basket_items = {'apples': 4, 'oranges': 19, 'kites': 3, 'sandwiches': 8}`
# 果物のリストは下記通り。<br/>
# `fruits = ['apples', 'oranges', 'pears', 'peaches', 'grapes', 'bananas']`
#
# + id="MwEUrD0HQEOX" colab_type="code" outputId="a004b606-8ab7-4cec-d3da-8ea9cd8cd0d5" colab={"base_uri": "https://localhost:8080/", "height": 33}
total = 0
basket_items = {'apples': 4, 'oranges': 19, 'kites': 3, 'sandwiches': 8}
fruits = ['apples', 'oranges', 'pears', 'peaches', 'grapes', 'bananas']
for object, calculate in basket_items.items():
if object in fruits:
total += calculate
print("かごの中には{}個の果物が入っている.".format(total))
# + [markdown] id="Hz2QBDAXQUnb" colab_type="text"
# 9. かごの中には果物ではないものを数えなさい。
# + id="S-WPOJkMQbM-" colab_type="code" outputId="004e704e-7090-4fa9-b347-b5a24b45439f" colab={"base_uri": "https://localhost:8080/", "height": 33}
total = 0
for object, calculate in basket_items.items():
if object not in fruits:
total += calculate
print("かごの中には{}個のものが果物ではない.".format(total))
# + [markdown] id="0dq7mp5wRejT" colab_type="text"
# ## `while` ループ
#
# 1. 1から10まで`while`ループを使って、表示しなさい。同じく、`for`ループを使って表示しなさい。比較してみましょう。
# + id="EeXA6aeBR9MS" colab_type="code" outputId="2c709978-71fc-46be-99ea-286df9e4bb2a" colab={"base_uri": "https://localhost:8080/", "height": 184}
i = 1 #初期化
while i < 11: #最後の値
print(i)
i += 1 #ステップ
# + id="XDSoFkYRSHnX" colab_type="code" outputId="404daa6c-cbdc-4a8e-8ff7-ecc6f324af64" colab={"base_uri": "https://localhost:8080/", "height": 184}
for i in range(1,11):
print(i)
# + [markdown] id="yImR7WqiSKKJ" colab_type="text"
# 2. 1から10までの合計を`while`と`for`ループを使って、計算しなさい。
# + id="0lwr_twQSUn7" colab_type="code" outputId="7e63759a-e9b8-4dcf-ed54-b50be9c1c999" colab={"base_uri": "https://localhost:8080/", "height": 35}
i = 1
total = 0
while i <= 10:
total += i
i += 1
print(total)
# + id="2_Up3tcDSYRW" colab_type="code" outputId="83c0d2a9-98d7-4867-de4c-91915d6eae86" colab={"base_uri": "https://localhost:8080/", "height": 33}
total = 0
for i in range(1,11):
total += i
print(total)
# + [markdown] id="vJXy3WEpSauq" colab_type="text"
# 3. 50の一番近い2乗を`while`ループを使って、表示しなさい。答えは`49`。
# + id="JAUQmz6BS3rp" colab_type="code" outputId="82a1b5c3-d5af-4c1d-e649-8ad65598bc2d" colab={"base_uri": "https://localhost:8080/", "height": 33}
limit = 50
num = 0
while (num+1)**2 < limit:
num += 1
nearest_square = num ** 2
print(nearest_square)
# + [markdown] id="nkW9ALUqTSUB" colab_type="text"
# ## `break`と`continue`
#
# 1. 1から20の奇数を`continue`を使って、表示しなさい。
# + id="okkAhb6ITjRZ" colab_type="code" outputId="4b514369-c327-463f-a06d-04f26de1f22b" colab={"base_uri": "https://localhost:8080/", "height": 33}
for n in range(20):
#もし計算のあまりが0の場合、ループをスキップ
if n % 2 == 0:
continue
print(n, end=' ')
# + [markdown] id="mAI0uxlxTs2V" colab_type="text"
# 2. `break`を使って、下記の文章を100文字まで表示しなさい。
#
# `機械学習では、センサやデータベースなどに由来するサンプルデータを入力して解析を行い、そのデータから有用な規則、ルール、知識表現、判断基準などを抽出し、アルゴリズムを発展させる。そのアルゴリズムは、まずそのデータ(訓練例と呼ぶ)を生成した潜在的機構の特徴(確率分布)を捉え、複雑な関係を識別・定量化する。次に学習・識別したパターンを用いて新たなデータについて予測・決定を行う。データは、観測された変数群のとる関係の具体例と見ることができる`
# + id="qvBRg27AUHCH" colab_type="code" outputId="4cc73d16-1e8d-4e99-bf1e-fdeb3fe55855" colab={"base_uri": "https://localhost:8080/", "height": 53}
paragraph = "機械学習では、センサやデータベースなどに由来するサンプルデータを入力して解析を行い、そのデータから有用な規則、ルール、知識表現、判断基準などを抽出し、アルゴリズムを発展させる。そのアルゴリズムは、まずそのデータ(訓練例と呼ぶ)を生成した潜在的機構の特徴(確率分布)を捉え、複雑な関係を識別・定量化する。次に学習・識別したパターンを用いて新たなデータについて予測・決定を行う。データは、観測された変数群のとる関係の具体例と見ることができる"
for p in paragraph:
if len(paragraph) >= 100:
sentence = paragraph[:100]
break
print(sentence)
# + [markdown] id="Q-TWq9kMUvmg" colab_type="text"
# ## 生入力スクリプト
#
# 1. ユーザーを自分の名前を入力させてください。
# + id="3fMM_APlU-K4" colab_type="code" outputId="a283ed4f-1970-4831-96b6-169dcff1d9eb" colab={"base_uri": "https://localhost:8080/", "height": 50}
name = input("名前: ")
print(name)
# + [markdown] id="r2lsLHrpVD9q" colab_type="text"
# 2.ユーザーに`こんにちは`とあいさつしましょう。
# + id="Ed2hcKIUVKyk" colab_type="code" outputId="1b74c12e-475f-49d2-93c4-826d3b6f012e" colab={"base_uri": "https://localhost:8080/", "height": 33}
print("こんにちは {}!".format(name))
# + [markdown] id="7RoHWT4tVP2h" colab_type="text"
# 3. ユーザーの年齢を聞きましょう。
# + id="jjseAqpXVVSN" colab_type="code" outputId="c55b9ca4-b6cc-482b-c52d-91b9854b1cc8" colab={"base_uri": "https://localhost:8080/", "height": 50}
age = input("年齢: ")
print(age)
# + [markdown] id="-3OSugxpVdXk" colab_type="text"
# 4. ユーザーの5年後の年齢を表示しなさい。
# + id="psxpvkYeVkXH" colab_type="code" outputId="61c47cd2-3360-4d75-aae8-63bdd3184950" colab={"base_uri": "https://localhost:8080/", "height": 33}
age = int(age)
age_5_years_later = age + 5
print(age_5_years_later)
# + [markdown] id="DnN2M0PfVoNs" colab_type="text"
# 5. 下記のように出力しましょう。
#
# `こんにちは <name>さん!
# <name>さんは今<age>歳ですが、5年後は<age_5_years_later>歳になりますよ。`
# + id="fEVmPOS0V9eR" colab_type="code" outputId="c55f5996-d47a-4441-e648-6aceaf9789fd" colab={"base_uri": "https://localhost:8080/", "height": 33}
print("こんにちは{}さん! {}さんは今{}歳ですが、5年後は{}歳になりますよ".format(name, name, age, age_5_years_later))
# + [markdown] id="6rtloOijXEEu" colab_type="text"
# 6. もしたくさんのユーザーがいて、`for`ループを使って、どうやって簡単に表示できますか?
# + id="v_-VlrHaXTfG" colab_type="code" outputId="f611bae9-cd56-473b-edb7-1b78469929a0" colab={"base_uri": "https://localhost:8080/", "height": 84}
names = input("コマで区切って名前を入力して: ").title().split(",")
ages = input("コマで区切って年齢を入力して: ").split(",")
message = "こんにちは{}さん! {}さんは今{}歳ですが、5年後は{}歳になりますよ"
for name, age in zip(names, ages):
print(message.format(name, name, age, int(age)+5))
# + [markdown] id="OBh4zpKkZHho" colab_type="text"
# ## エラーと例外
#
# 1. 変数を設定せずに、表示しようとしたらどうなりますか?
# + id="1Jk1gjacZR4B" colab_type="code" outputId="1f0a8ac6-71eb-4aed-eb7d-323fd5213490" colab={"base_uri": "https://localhost:8080/", "height": 161}
print(x)
# + [markdown] id="a4ygLBWPZTqY" colab_type="text"
# 2. 文字列と整数を算術しようとしたら、どうなりますか?
#
# + id="f9hYDpDsZe2W" colab_type="code" outputId="8991f40b-1b28-4b03-94de-a96f210b5978" colab={"base_uri": "https://localhost:8080/", "height": 161}
1 + 'abc'
# + [markdown] id="Htv4yeciZqeS" colab_type="text"
# 3. 存在しないリストのインデックスをアクセスしようとしたら、どうなりますか?
# + id="gikc5c0nZzeb" colab_type="code" outputId="af4166fd-b1eb-4e9e-8273-63abe522b4f8" colab={"base_uri": "https://localhost:8080/", "height": 178}
L = ["a", "b", "c"]
L[1000]
# + id="7g7eN9HqZ42T" colab_type="code" outputId="46c81502-10d4-41f1-c4da-25e260ecdd79" colab={"base_uri": "https://localhost:8080/", "height": 178}
L = ["a", "b", "c"]
L["d"]
# + [markdown] id="xA1ECEdHZ28L" colab_type="text"
# 4. `try`と`except`の関数を使って、エラーを見つかりましょう。
#
# 下記の文を実行してみてください。
#
# ```
# try:
# print("this gets executed first")
# except:
# print("this gets executed only if there is an error")
# ```
#
#
# + id="sXcwqd8LaPqJ" colab_type="code" outputId="8c2415a7-6349-4f0a-b461-f3948c6cce35" colab={"base_uri": "https://localhost:8080/", "height": 33}
try:
print("this gets executed first")
except:
print("this gets executed only if there is an error")
# + [markdown] id="KQWJXqmkbpCz" colab_type="text"
# 5. `x=1/0`を`try`のところに足してみたら?
# + id="clO4CuwTbuJt" colab_type="code" outputId="a917b77c-a815-4ecd-b652-2f1cab75c9bb" colab={"base_uri": "https://localhost:8080/", "height": 50}
try:
print("let's try something:")
x = 1 / 0 # ZeroDivisionError
except:
print("something bad happened!")
# + [markdown] id="QLXziXiAb0Qp" colab_type="text"
# 6. 例外を`raise`しましょう。
# + id="ZDH2yvxMb56P" colab_type="code" outputId="7acac87c-fe45-4532-ab92-18828da0a8e2" colab={"base_uri": "https://localhost:8080/", "height": 161}
raise RuntimeError("my error message")
# + [markdown] id="VbbSLgzSb8ue" colab_type="text"
# 7. `x=10`とすると、`x`は5以上の場合、例外を`raise`してください。
# + id="DtuqWftzcI0H" colab_type="code" outputId="7968ac01-118d-46fc-de83-2331b9463cdd" colab={"base_uri": "https://localhost:8080/", "height": 215}
x = 10
if x > 5:
raise Exception('x should not exceed 5. The value of x was {}'. format(x))
# + [markdown] id="_bJBvSwTcluS" colab_type="text"
# 8. `try`と`except`の他に、`else`と`finally`は例外を実行させる。`finally`なら、どんなことがあっても、実行させる。
#
# + id="nTVdOk5udBQC" colab_type="code" outputId="ca8467a9-f5c8-4123-f88f-2356be4cb63d" colab={"base_uri": "https://localhost:8080/", "height": 67}
try:
print("try something here")
except:
print("this happens only if it fails")
else:
print("this happens only if it succeeds")
finally:
print("this happens no matter what")
| Lecture08.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **잡히지 않은 중복값들을 마저 제거해준 노트북입니다.**
#
# 1. 영어의 대소문자 여부가 다르게 들어간 것
# 2. 띄어쓰기가 다르게 들어간 것
# 3. 피처링이 다른 버전
import pandas as pd
# 곡 정보 파일 불러오기
df = pd.read_csv('../Data/song_data_yewon_ver05.csv')
df
# 제목의 공백(띄어쓰기)를 모두 제거한다
df['title'] = df['title'].str.replace(r' ', '')
# 제목의 영어 부분을 전부 소문자로 바꿔준다
df['title'] = df['title'].str.lower()
# 그리고 다시 중복값을 제거해준다.
df = df.drop_duplicates(['artist', 'title'], keep='last')
# 중복 값을 찍어보니 잘 지워졌다! (띄어쓰기 제거 테스트)
df[df['title'] == '결혼 하지마']
df[df['title'] == '결혼하지마']
# 중복 값을 찍어보니 잘 지워졌다! (영어 대->소문자 변환 테스트)
df[df['title'] == '어이\(UH-EE\)']
df[df['title'].str.contains('어이\(uh-ee\)')]
# 제목 열을 새로 만들어서
df['t'] = df['title']
# 괄호 안의 부분을 없앤다.
df.t = df.t.str.replace(r'\(.*?\)','')
# 새로 만든 열의 중복값을 제거한다.
df = df.drop_duplicates(['artist', 't'], keep='last')
# 새로 만든 열을 다시 지워준다.
df = df.drop('t', axis = 1)
# 중복 값을 찍어보니 잘 지워졌다! (하나만 남음) (피처링 다른 버전 제거 테스트)
df[df['title'].str.contains('highwaystar')]
df.shape
df.to_csv('../Data/song_data_yoon_ver01.csv', index=False)
| SongTidy/song_tidy_yoon_ver01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Here we recreate column "3" of Table 1 of the Li and Ding paper.
import numpy as np
from cibin import tau_twosided_ci
# +
alpha = .05
cases = [(1,1,1,13),
(2,6,8,0),
(6,0,11,3),
(6,4,4,6),
(1,1,3,19),
(8,4,5,7)]
print(f"n\t\t\t3")
print(f"-----\t\t\t-----")
for n in cases:
N = sum(n)
n11, n10, n01, n00 = n
ci, _, _ = tau_twosided_ci(n11, n10, n01, n00, alpha, exact=(N<20))
print(f"{n}\t\t{N*np.array(ci)}")
# -
tau_twosided_ci(2, 6, 8, 0, 0.05)
# The final row is approximate, so it does not exactly match with the paper.
| cibin-demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:seq]
# language: python
# name: conda-env-seq-py
# ---
# ### Load Model
# +
import sys
sys.path.insert(0, '../')
import aux
from aux import *
# #%load_ext autoreload
# #%autoreload 2
##Clear Memory
tf.reset_default_graph()
tf.keras.backend.clear_session()
gc.collect()
##
NUM_GPU = len(get_available_gpus())
if(NUM_GPU>0) :
config = tf.ConfigProto()
config.gpu_options.allow_growth = True # dynamically grow the memory used on the GPU
print(tf.__version__)
print(keras.__version__)
#tpu_grpc_url = TPUClusterResolver(tpu=['edv-tpu2'] , zone='us-central1-a').get_master()
### Load the Model in a separate graph here as there are two models in this figure.
fitness_function_graph = tf.Graph()
with fitness_function_graph.as_default():
model_conditions='Glu'
model, scaler,batch_size = load_model(model_conditions)
plotting_alpha=0.2
# -
# ### Load Data
#
# <li>Y_pred contains the predictions
# <li>X is the format of one-hot encoded input expected
# +
def read_hq_testdata(filename) :
with open(filename) as f:
reader = csv.reader(f, delimiter="\t")
d = list(reader)
sequences = [di[0] for di in d]
for i in tqdm(range(0,len(sequences))) :
if (len(sequences[i]) > 110) :
sequences[i] = sequences[i][-110:]
if (len(sequences[i]) < 110) :
while (len(sequences[i]) < 110) :
sequences[i] = 'N'+sequences[i]
A_onehot = np.array([1,0,0,0] , dtype=np.bool)
C_onehot = np.array([0,1,0,0] , dtype=np.bool)
G_onehot = np.array([0,0,1,0] , dtype=np.bool)
T_onehot = np.array([0,0,0,1] , dtype=np.bool)
N_onehot = np.array([0,0,0,0] , dtype=np.bool)
mapper = {'A':A_onehot,'C':C_onehot,'G':G_onehot,'T':T_onehot,'N':N_onehot}
worddim = len(mapper['A'])
seqdata = np.asarray(sequences)
seqdata_transformed = seq2feature(seqdata)
print(seqdata_transformed.shape)
expressions = [di[1] for di in d]
expdata = np.asarray(expressions)
expdata = expdata.astype('float')
return np.squeeze(seqdata_transformed),expdata
X,Y = read_hq_testdata(os.path.join('..','..','data','Glu','HQ_testdata.txt'))
Y = [float(x) for x in Y]
Y_pred = evaluate_model(X, model, scaler, batch_size , fitness_function_graph)
#plt.scatter(Y_pred,Y)
plt.xlabel('Predicted')
plt.ylabel('Measured')
#print('MSE',sklearn.metrics.mean_squared_error(Y_pred,Y))
# -
# ### Evaluate Model
# <li> Here, we have the measured expression values Y corresponding to the sequences X
# +
fig=plt.figure(figsize=(4,4), dpi= 200, facecolor='w', edgecolor='k')
sns.regplot(Y_pred,Y , s=5 , linewidth=0.25)
plt.title(scipy.stats.spearmanr(Y_pred,Y)[0])
plt.xlabel('Predicted')
plt.ylabel('Measured')
ax = plt.gca()
plt.xticks([])
plt.yticks([])
ax.autoscale(enable=True, axis='x', tight=True)
ax.autoscale(enable=True, axis='y', tight=True)
# -
| manuscript_code/model/tpu_model/use_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import numpy as np
from tensorflow.keras import layers
from tensorflow.keras import Model
"""!wget --no-check-certificate \
https://storage.googleapis.com/mledu-datasets/inception_v3_weights_tf_dim_ordering_tf_kernels_notop.h5 \
-O /home/soundarzozm/Desktop/mask_detector/inception_v3_weights_tf_dim_ordering_tf_kernels_notop.h5"""
from tensorflow.keras.applications.inception_v3 import InceptionV3
local_weights_file = '/home/soundarzozm/Desktop/mask_detector/inception_v3_weights_tf_dim_ordering_tf_kernels_notop.h5'
pre_trained_model = InceptionV3(input_shape = (256, 256, 3),
include_top = False,
weights = None)
pre_trained_model.load_weights(local_weights_file)
for layer in pre_trained_model.layers:
layer.trainable = False
pre_trained_model.summary()
last_layer = pre_trained_model.get_layer('mixed7')
print('last layer output shape: ', last_layer.output_shape)
last_output = last_layer.output
# +
from tensorflow.keras.optimizers import RMSprop
x = layers.Flatten()(last_output)
x = layers.Dense(1024, activation='relu')(x)
x = layers.Dropout(0.2)(x)
x = layers.Dense (1, activation='sigmoid')(x)
model = Model(pre_trained_model.input, x)
model.compile(optimizer = RMSprop(lr=0.0001),
loss = 'binary_crossentropy',
metrics = ['accuracy'])
# +
from tensorflow.keras.preprocessing.image import ImageDataGenerator
base_dir = '/home/soundarzozm/Desktop/mask_detector'
train_dir = os.path.join(base_dir, 'train')
validation_dir = os.path.join(base_dir, 'validation')
test_dir = os.path.join(base_dir, 'test')
train_mask_dir = os.path.join(train_dir, 'with_mask')
train_no_mask_dir = os.path.join(train_dir, 'without_mask')
validation_mask_dir = os.path.join(validation_dir, 'with_mask')
validation_no_mask_dir = os.path.join(validation_dir, 'without_mask')
train_mask_fnames = os.listdir(train_mask_dir)
train_no_mask_fnames = os.listdir(train_no_mask_dir)
train_datagen = ImageDataGenerator(rescale = 1./255.,
rotation_range = 40,
width_shift_range = 0.2,
height_shift_range = 0.2,
shear_range = 0.2,
zoom_range = 0.2,
horizontal_flip = True)
test_datagen = ImageDataGenerator(rescale = 1.0/255.)
train_generator = train_datagen.flow_from_directory(train_dir,
batch_size = 20,
class_mode = 'binary',
target_size = (256, 256))
validation_generator = test_datagen.flow_from_directory(validation_dir,
batch_size = 20,
class_mode = 'binary',
target_size = (256, 256))
test_generator = test_datagen.flow_from_directory(test_dir,
batch_size = 1,
shuffle=False,
class_mode='binary',
target_size = (256, 256))
# -
history = model.fit(
train_generator,
validation_data = validation_generator,
steps_per_epoch = 100,
epochs = 4,
validation_steps = 50,
verbose = 1)
# +
import matplotlib.pyplot as plt
acc = history.history['accuracy']
val_acc = history.history['val_accuracy']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(acc))
plt.plot(epochs, acc, 'r', label='Training accuracy')
plt.plot(epochs, val_acc, 'b', label='Validation accuracy')
plt.plot(epochs, loss, 'g', label='Training loss')
plt.plot(epochs, val_loss, 'y', label='Validation loss')
plt.title('Training and validation accuracy and loss')
plt.legend(loc=0)
plt.figure()
plt.show()
# +
filenames = test_generator.filenames
nb_samples = len(filenames)
predict = model.predict(test_generator, steps = nb_samples)
# -
model.save("model.h5")
print(predict)
| training_notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: torch_env
# language: python
# name: torch_env
# ---
# # 2장에서 다루는 내용
#
# - 2장에서는 자연어의 의미를 임베딩에 어떻게 녹여낼 수 있는지를 주로 살핀다.
# ### 2장 벡터가 어떻게 의미를 가지게 되는가
# 2.1 자연어 계산과 이해
# 2.2 어떤 단어가 많이 쓰였는가
# 2.2.1 백오브워즈 가정
# 2.2.2 TF-IDF
# 2.2.3 Deep Averaging Network
# 2.3 단어가 어떤 순서로 쓰였는가
# 2.3.1 통계 기반 언어 모델
# 2.3.2 뉴럴 네트워크 기반 언어 모델
# 2.4 어떤 단어가 같이 쓰였는가
# 2.4.1 분포 가정
# 2.4.2 분포와 의미 (1): 형태소
# 2.4.3 분포의 의미 (2): 품사
# 2.4.4 점별 상호 정보량
# 2.4.5 Word2Vec
# 2.5 이 장의 요약
# 2.6 참고문헌
# # 2.1 자연어 계산과 이해
# 컴퓨터는 자연어를 사람처럼 이해할 수 없는 계산기일 뿐이지만, 임베딩을 활용하면 자연어를 계산하는 것이 가능해진다.
#
# 임베딩: 컴퓨터가 처리할 수 있는 숫자들의 나열인 벡터
#
# 어떻게? 자연어의 통계적 패턴 정보를 통째로 임베딩에 넣는다.
#
# 임베딩 구축 시 사용하는 통계 정보 3가지
# 1. 문장에 어떤 단어가 (많이) 쓰였는지 [빈도]
# 2. 단어가 어떤 (순서)로 등장하는지 [순서]
# 3. 문장에 어떤 단어가 (같이) 나타났는지 [동시출현]
#
# |구분 | 백오브워즈 가정 | 언어 모델 | 분포 가정 |
# |--------|-------------------|----------------|-------|
# |내용 | 어떤 단어가 (많이) 쓰였는가 | 단어가 어떤 순서로 쓰였는가 | 어떤 단어가 같이 쓰였는가|
# |대표 통계량| TF-IDF | - | PMI |
# |대표 모델 | Deep Averaging Network | ELMo, GPT | Word2Vec |
# ## Bag Of Words (BOW) = 어떤 단어가 (많이) 쓰였는지!
#
# - 저자의 의도는 단어 사용 여부나 그 빈도에서 드러난다.
# - 순서 정보는 무시.
# - 가장 많이 사용하는 것은 TF-IDF / Deep Learning Version = Deep Averagin Network
#
# ## Language Model = 단어 시퀀스가 얼마나 자연스러운지에 대한 확률
#
# - ELMo, GPT 등과 같은 뉴럴 네트워크 기반의 언어모델
#
# ## Distributional Hypothesis = 문장에서 어떤 단어가 같이 쓰였는지를 고려
#
# - 단어의 의미는 그 주변 문맥을 통해 유추 가능
# - 대표통계량은 점별 상호 정보량 (PMI, Pointwise Mutual Information)
# - 대표모델은 Word2Vec
#
# 백오브 워즈 가정, 언어 모델, 분포 가정은 말뭉치의 통계적 패턴을 서로 다른 각도에서 분석하는 것이며, 상호 보완적이다.
# # 2.2 어떤 단어가 많이 쓰였는가
#
# ### 2.2.1 Bag Of Words
#
# > Bag이란 중복 원소를 허용한 집합(Multiset), 순서는 고려하지 않아!
#
# {a,a,b,c,c,c} = {c,a,b,c,a,c}, {c,a,c,b,a,c}
#
# <b>백오브워즈</b>는 문장을 단어들로 나두고 이들을 중복 집합에 넣어 임베딩으로 활용하는 것이라고 보면 된다.
#
# ##### 가정: 저자가 생각한 주제가 문서에서의 단어 사용에 녹아있다!
#
# ### 2.2.2 TF-IDF
#
# 단순하게 단어 빈도만을 임베딩으로 활용하는 것에는 단점이 있다.
#
# '을/를'과 같은 조사와 같은 경우는 어떠한 문서에도 등장하지만, 문서에 대하여 어떠한 내용도 담고 있지 않다.
# 이를 위해 <b>Term Frequency-Inverse Document Frequency</b>가 필요하다.
#
#
#
# # 2.3 단어가 어떤 순서로 쓰였는가
# ### 2.3.1 통계 기반 언어 모델
# > 언어 모델이란 단어 시퀀스에 확률을 부여하는 모델
#
# BOW는 순서를 무시하지만 언어모델은 시퀀스 정보를 명시적으로 학습.
# 단어가 n개라면 P(w1, w2, ...,wn)을 반환한다.
#
# 잘 학습된 언어 모델이라면 어떤 문장이 그럴듯한지(확률값이 높은지), 주어진 단어 시퀀스 다음 단어는 무엇이 올지 파악.
#
# ex)
# > 누명을 쓰다 (높은 확률)
# 누명을 당하다 (낮은 확률)
#
# * n-gram: n개 단어를 뜻하는 용어
# bi-gram: (난폭, 운전) / (눈, 뜨다)
# tri-gram: (누명,을,쓰다) / (초코, 칩, 쿠키)
# ...
#
# n-gram은 n-gram에 기반한 언어 모델을 의미하기도 하여, 단어들을 n개씩 묶어서 빈도를 학습했다는 뜻이기도 하다.
#
# * n-gram 허점:
# "
# # 2.4 어떤 단어가 같이 쓰였는가
# ### 2.4.4 점별 상호 정보량(PMI, Pointwise Mutual Information)
#
# 점별 상호 정보량은 두 확률변수 사이의 상관성을 계량화하는 단위다. (완전 독립 시, 0)
# 독립이라 함은 한 단어의 등장이 다른 단어의 등장에 전혀 영향을 주지 않는 다는 뜻이다.
# 만일 영향을 준다면 PMI값이 커지게 된다.
#
# <img src="image/PMI.png" width=300 height=300 />
#
# PMI는 분포 가정에 따른 단어 가중치 할당 기법이기에, PMI 행렬의 행벡터 자체를 해당 단어의 임베딩으로 사용 가능하다.
#
# <img src="image/word_context_matrix.png" width=300 height=300 />
#
#
| 2_Meaning_of_Vector.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lab 1: Welcome to Python!
# ## Goals
# Welcome to your first lab day! Labs in CS41 are designed to be your opportunity to experiment with Python and gain hands-on experience with the language.
#
# The primary goal of this lab is to ensure that your Python installation process went smoothly, and that there are no lingering Python 2/3 bugs floating around.
#
# This lab also gives you the chance to write what might be your first programs in Python and allows you to experiment with both scripts and with the interactive interpreter!
#
# These problems are not intended to be algorithmically challenging - just ways to flex your new Python 3 muscles. Even if the problems seem simple, work through them quickly, and then you're free to go.
#
# As always, have fun, and enjoy the (remainder of the) class period!
# ## Zen of Python
#
# Run the following code cell by selecting the cell and pressing Shift+Enter.
import this
# ## Hello World
#
# Edit the following cell so that it prints `"Hello, world!"` when executed.
# +
# Edit me so that I print out "Hello, world!" when run!
# -
# ### Fizz, Buzz, FizzBuzz!
# If we list all of the natural numbers under 41 that are a multiple of 3 or 5, we get
#
# ```
# 3, 5, 6, 9, 10, 12, 15,
# 18, 20, 21, 24, 25, 27, 30,
# 33, 35, 36, 39, 40
# ```
#
# The sum of these numbers is 408.
#
# Find the sum of all the multiples of 3 or 5 below 1001.
# +
def fizzbuzz(n):
"""Returns the sum of all numbers < `n` divisible by 3 or 5."""
pass
fizzbuzz(1001)
# -
# ### Collatz Sequence
# Depending on who you took CS106A from, you may have seen this problem before.
#
# The *Collatz sequence* is an iterative sequence defined on the positive integers by:
#
# ```
# n -> n / 2 if n is even
# n -> 3n + 1 if n is odd
# ```
#
# For example, using the rule above and starting with 13 yields the sequence:
#
# ```
# 13 -> 40 -> 20 -> 10 -> 5 -> 16 -> 8 -> 4 -> 2 -> 1
# ```
#
# It can be seen that this sequence (starting at 13 and finishing at 1) contains 10 terms. Although unproven, it it hypothesized that all starting numbers finish at 1.
#
# What is the length of the longest chain which has a starting number under 1000?
#
# *NOTE: Once the chain starts the terms are allowed to go above one thousand.*
#
# Challenge: Same question, but for any starting number under 1,000,000 (you may need to implement a cleverer-than-naive algorithm)
# +
def collatz_len(n):
"""Computes the length of the Collatz sequence starting at `n`."""
pass
def max_collatz_len(n):
"""Computes the longest Collatz sequence length for starting numbers < `n`"""
# -
# ### Fahrenheit-to-Celsius converter
# Write a program to convert degrees Fahrenheit to degrees Celcius by (1) asking the user for a number (not necessarily integral) representing the current temperature in degrees Fahrenheit, (2) converting that value into the equivalent degrees Celsius, and (3) printing the final equivalent value.
#
# For example, your program should be able to emulate the following three sample runs:
#
# ```
# Temperature F? 212
# It is 100.0 degrees Celsius.
#
# Temperature F? 98.6
# It is 37.0 degrees Celsius.
#
# Temperature F? 10
# It is -12.222222222222221 degrees Celsius.
# ```
#
# Want to be fancy (challenge)? Try to print the final temperature to two decimal places. *Hint: Take a look at the [`round()`](https://docs.python.org/3.4/library/functions.html#round) function. Isn't Python great?*
# +
def convert_fahr_to_cels(deg_fahr):
pass
def convert():
pass
# -
# ## Bonus Challenges
#
# Don't worry about getting to these bonus problems. In most cases, bonus questions ask you to think more critically or use more advanced algorithms.
# ### Zen Printing
#
# Write a program using `print()` that, when run, prints out a tic-tac-toe board.
#
# ```
# X | . | .
# -----------
# . | O | .
# -----------
# . | O | X
# ```
#
# You may find the optional arguments to `print()` useful, which you can read about [here](https://docs.python.org/3/library/functions.html#print). In no more than five minutes, try to use these optional arguments to print out this particular tic-tac-toe board.
# +
# Print a tic-tac-toe board using optional arguments.
# -
# Maybe you were able to print out the tic-tac-toe board. Maybe not. In the five minutes you've been working on that, I've gotten bored with normal tic-tac-toe (too many ties!) so now, I want to play SUPER tic-tac-toe.
#
# Write a program that prints out a SUPER tic-tac-toe board.
#
# ```
# | | H | | H | |
# --+--+--H--+--+--H--+--+--
# | | H | | H | |
# --+--+--H--+--+--H--+--+--
# | | H | | H | |
# ========+========+========
# | | H | | H | |
# --+--+--H--+--+--H--+--+--
# | | H | | H | |
# --+--+--H--+--+--H--+--+--
# | | H | | H | |
# ========+========+========
# | | H | | H | |
# --+--+--H--+--+--H--+--+--
# | | H | | H | |
# --+--+--H--+--+--H--+--+--
# | | H | | H | |
# ```
#
# You'll find that there might be many ways to solve this problem. Which do you think is the most 'pythonic?' Talk to someone next to you about your approach to this problem. Remember the Zen of Python!
# ## Done Early?
#
# Read [Python’s Style Guide](https://www.python.org/dev/peps/pep-0008/), keeping the Zen of Python in mind. In what ways do you notice that Python's style guidelines are influence by Python's core philosophy? Some portions of the style guide cover language features that we haven't yet touched on in class - feel free to skip over those sections for now.
# ## Submitting Labs
#
# Alright, you did it! There's nothing to submit for this lab. You're free to leave as soon as you've finished this lab.
# *Credit to ProjectEuler and InterviewCake for some problem ideas.*
#
# > With <3 by @sredmond
| notebooks/lab1-warmup-notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Advanced Lane Finding Project
#
# The goals / steps of this project are the following:
#
# * Compute the camera calibration matrix and distortion coefficients given a set of chessboard images.
# * Apply a distortion correction to raw images.
# * Use color transforms, gradients, etc., to create a thresholded binary image.
# * Apply a perspective transform to rectify binary image ("birds-eye view").
# * Detect lane pixels and fit to find the lane boundary.
# * Determine the curvature of the lane and vehicle position with respect to center.
# * Warp the detected lane boundaries back onto the original image.
# * Output visual display of the lane boundaries and numerical estimation of lane curvature and vehicle position.
#
# ---
# ## First, I'll compute the camera calibration using chessboard images, and extract the object points and images points
# ## Compute camera calibration and distortion correction for chessboard images
# +
import numpy as np
import cv2
import glob
import matplotlib.pyplot as plt
# %matplotlib inline
# prepare object points, like (0,0,0), (1,0,0), (2,0,0) ....,(6,5,0)
nx=9
ny=6
objp = np.zeros((nx*ny,3), np.float32)
objp[:,:2] = np.mgrid[0:nx,0:ny].T.reshape(-1,2)
# Arrays to store object points and image points from all the images.
objpoints = [] # 3d points in real world space
imgpoints = [] # 2d points in image plane.
# Make a list of calibration images
images = glob.glob('../camera_cal/calibration*.jpg')
# Step through the list and search for chessboard corners
for fname in images:
img = cv2.imread(fname)
gray = cv2.cvtColor(img,cv2.COLOR_BGR2GRAY)
# Find the chessboard corners
ret, corners = cv2.findChessboardCorners(gray, (9,6),None)
# If found, add object points, image points
if ret == True:
objpoints.append(objp)
imgpoints.append(corners)
#ret, mtx, dist, rvecs, tvecs = cv2.calibrateCamera(objpoints, imgpoints, gray.shape[::-1], None, None)
# Draw and display the corners
img = cv2.drawChessboardCorners(img, (9,6), corners, ret)
cv2.imshow('img',img)
cv2.waitKey(500)
cv2.destroyAllWindows()
# -
# ## Undistort and unwarp the images using camera calibration and perspective transform for Chessboards
# +
import numpy as np
import cv2
import glob
import matplotlib.pyplot as plt
# %matplotlib inline
# prepare object points, like (0,0,0), (1,0,0), (2,0,0) ....,(6,5,0)
nx=9
ny=6
objp = np.zeros((nx*ny,3), np.float32)
objp[:,:2] = np.mgrid[0:nx,0:ny].T.reshape(-1,2)
# Arrays to store object points and image points from all the images.
objpoints = [] # 3d points in real world space
imgpoints = [] # 2d points in image plane.
# Make a list of calibration images
images = glob.glob('../camera_cal/calibration*.jpg')
img=cv2.imread(images[19])
img=np.array(img)
## Define the function for image unwarp
def Unwarp_img(img, nx, ny):
# Find the chessboard corners
gray=cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
ret, corners=cv2.findChessboardCorners(gray, (nx, ny), None)
if ret == True:
objpoints.append(objp)
imgpoints.append(corners)
ret, mtx, dist, rvecs, tvecs = cv2.calibrateCamera(objpoints, imgpoints, gray.shape[::-1], None, None)
undist = cv2.undistort(img, mtx, dist, None, mtx)
offset=100
img_size=(gray.shape[1],gray.shape[0])
src=np.float32([corners[0,:,:], corners[nx-1,:,:],corners[-1,:,:],corners[-nx,:,:]])
dst = np.float32([[offset, offset], [img_size[0]-offset, offset],
[img_size[0]-offset, img_size[1]-offset],
[offset, img_size[1]-offset]])
# Given src and dst points, calculate the perspective transform matrix
M = cv2.getPerspectiveTransform(src, dst)
Minv = cv2.getPerspectiveTransform(dst, src)
# Warp the image using OpenCV warpPerspective()
warped = cv2.warpPerspective(undist, M, img_size)
return warped, M
warped_img, perspective_M=Unwarp_img(img, nx, ny)
f, (ax1, ax2) = plt.subplots(1, 2, figsize=(24, 9))
f.tight_layout()
ax1.imshow(img)
ax1.set_title('Original Image', fontsize=30)
ax2.imshow(warped_img)
ax2.set_title('Undistorted and Warped Image', fontsize=30)
plt.savefig('Undistorted_Warped_Image.png')
# -
# # Pipeline for Lane detection on images
# ### Define all functions
#
#
# ## Apply Camera calibration, distortion correction, color&gradients threshold and perspective transforms to the raw testing images
# +
import numpy as np
import cv2
import glob
import matplotlib.pyplot as plt
# %matplotlib inline
## Apply Camera calibration, distortion correction
def camera_calibration():
nx=9
ny=6
objp = np.zeros((nx*ny,3), np.float32)
objp[:,:2] = np.mgrid[0:nx,0:ny].T.reshape(-1,2)
# Arrays to store object points and image points from all the images.
objpoints = [] # 3d points in real world space
imgpoints = [] # 2d points in image plane.
# Make a list of calibration images
images = glob.glob('../camera_cal/calibration*.jpg')
# Step through the list and search for chessboard corners
for fname in images:
img_chess = cv2.imread(fname)
gray_chess = cv2.cvtColor(img_chess,cv2.COLOR_BGR2GRAY)
# Find the chessboard corners
ret, corners = cv2.findChessboardCorners(gray_chess, (9,6),None)
# If found, add object points, image points
if ret == True:
objpoints.append(objp)
imgpoints.append(corners)
shape=gray_chess.shape[::-1]
np.save('objpoints', objpoints)
np.save('imgpoints', imgpoints)
np.save('shape', shape)
return None
## Apply Camera calibration, distortion correction
def undistortion(img):
# Arrays to store object points and image points from all the images.
objpoints = np.load('objpoints.npy') # 3d points in real world space
imgpoints = np.load('imgpoints.npy') # 2d points in image plane.
shape=tuple(np.load('shape.npy'))
ret, mtx, dist, rvecs, tvecs = cv2.calibrateCamera(objpoints, imgpoints,shape,None, None)
undist=cv2.undistort(img, mtx, dist, None, mtx)
return undist
# read an image from a folder
#img=mpimg.imread('solidYellowCurve2.jpg')
img=cv2.imread('../test_images/test6.jpg')
b,g,r=cv2.split(img)
img=cv2.merge([r,g,b])
# print out some info about this image
#img=cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
print ('This image is:', type(img),'with dimensions of',img.shape)
# transform from color to gray scale
img_gray=cv2.cvtColor(img,cv2.COLOR_RGB2GRAY) #mpimg loads image as RGB
dst=undistortion(img)
f, (ax1, ax2)=plt.subplots(1,2, figsize=(24,9))
f.tight_layout()
ax1.imshow(img)
ax1.set_title('Original Image', fontsize=30)
ax2.imshow(dst)
# +
thresh_grad_min = 20
thresh_grad_max = 100
color_thre_min, color_thre_max=170, 255
## Define Color & gradients function
def gradient_threshold(img, thresh_grad_min, thresh_grad_max, color_thre_min, color_thre_max):
hls=cv2.cvtColor(img, cv2.COLOR_RGB2HLS)
s_channel=hls[:,:,2]
gray=cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
## Sobelx
sobelx = cv2.Sobel(gray, cv2.CV_64F, 1, 0) # Take the derivative in x
abs_sobelx = np.absolute(sobelx) # Absolute x derivative to accentuate lines away from horizontal
scaled_sobel = np.uint8(255*abs_sobelx/np.max(abs_sobelx))
# Threshold x gradient
sxbinary = np.zeros_like(scaled_sobel)
sxbinary[(scaled_sobel >= thresh_grad_min) & (scaled_sobel <= thresh_grad_max)] = 1
# Color threshold
s_binary=np.zeros_like(s_channel)
s_binary[(s_channel>=color_thre_min)&(s_channel<color_thre_max)]=1
# stack each channel to an image to see the contribution of individual channels
color_binary = np.dstack(( np.zeros_like(sxbinary), sxbinary, s_binary)) * 255
# combine two binary thresholds
combined_binary=np.zeros_like(s_binary)
combined_binary[(sxbinary==1)|(s_binary==1)]=1
return combined_binary
# +
## Perspective transform
## Define the function for image unwarp
def warped_img(img):
gray=cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
img_size=(gray.shape[1],gray.shape[0])
#src=np.float32([[500,500],[780,500],[img.shape[1],img.shape[0]],[150,img.shape[0]]])
src=np.float32([[580,460],[700,460],[1040,680],[260,680]])
dst = np.array([[200, 0], [1000, 0], [1000, img.shape[0]], [200, img.shape[0]]], np.float32)
# Given src and dst points, calculate the perspective transform matrix
M = cv2.getPerspectiveTransform(src, dst)
Minv = cv2.getPerspectiveTransform(dst, src)
# Warp the image using OpenCV warpPerspective()
warped = cv2.warpPerspective(img, M, img_size,flags=cv2.INTER_LINEAR)
return warped
# +
## Define the function of detecting lane lines
def find_lines(binary):
# calculate the current base leftx and rightx pixels location
image_height=binary.shape[0]
image_width=binary.shape[1]
histogram=np.sum(binary[binary.shape[0]//2:,:], axis=0)
midpoint=np.int(binary.shape[1]//2)
leftx_base=np.argmax(histogram[:midpoint]) # define the base left lanes location
rightx_base=np.argmax(histogram[midpoint:])+midpoint # define the base right lanes location
leftx_current=leftx_base
rightx_current=rightx_base
out_img=np.dstack((binary, binary, binary))
# nonzero x & y locations
nonzerox=np.array(binary.nonzero()[1])
nonzeroy=np.array(binary.nonzero()[0])
# left and right lane lines indices
leftx_ind, rightx_ind=[],[]
# Hyper parameters for windows
nwindow=9
margin=100
minpxl=50
for window in range(nwindow):
# define the parameters of windwos on image
win_height=np.int(image_height//nwindow)
win_y_low=image_height-(window+1)*win_height
win_y_high=image_height-window*win_height
win_left_low=leftx_current-margin
win_left_high=leftx_current+margin
win_right_low=rightx_current-margin
win_right_high=rightx_current+margin
# identify the nonzero pixels within the window
detected_leftx_ind=((nonzeroy >= win_y_low) & (nonzeroy < win_y_high) &
(nonzerox >= win_left_low) & (nonzerox < win_left_high)).nonzero()[0]
detected_rightx_ind=((nonzeroy >= win_y_low) & (nonzeroy < win_y_high) &
(nonzerox >= win_right_low) & (nonzerox < win_right_high)).nonzero()[0]
if len(detected_leftx_ind)>minpxl:
leftx_current=np.int(np.mean(nonzerox[detected_leftx_ind]))
if len(detected_rightx_ind)>minpxl:
rightx_current=np.int(np.mean(nonzerox[detected_rightx_ind]))
leftx_ind.append(detected_leftx_ind)
rightx_ind.append(detected_rightx_ind)
try:
leftx_ind=np.concatenate(leftx_ind)
rightx_ind=np.concatenate(rightx_ind)
except ValueErrors:
pass
leftx=nonzerox[leftx_ind]
rightx=nonzerox[rightx_ind]
lefty=nonzeroy[leftx_ind]
righty=nonzeroy[rightx_ind]
return leftx, lefty, rightx, righty, out_img
# define polynomial function of lane lines
def polynomial_fit(binary):
# Find our lane pixels first
leftx, lefty, rightx, righty, out_img = find_lines(binary)
# Fit a second order polynomial to each using `np.polyfit`
left_fit = np.polyfit(lefty, leftx, 2)
right_fit = np.polyfit(righty, rightx, 2)
# Generate x and y values for plotting
ploty = np.linspace(0, binary.shape[0]-1, binary.shape[0] )
try:
left_fitx = left_fit[0]*ploty**2 + left_fit[1]*ploty + left_fit[2]
right_fitx = right_fit[0]*ploty**2 + right_fit[1]*ploty + right_fit[2]
except TypeError:
# Avoids an error if `left` and `right_fit` are still none or incorrect
print('The function failed to fit a line!')
left_fitx = 1*ploty**2 + 1*ploty
right_fitx = 1*ploty**2 + 1*ploty
## Visualization ##
# Colors in the left and right lane regions
out_img[lefty, leftx] = [255, 0, 0]
out_img[righty, rightx] = [0, 0, 255]
# Plots the left and right polynomials on the lane lines
plt.plot(left_fitx, ploty, color='yellow')
plt.plot(right_fitx, ploty, color='yellow')
return out_img
# +
def curvature_cal(binary):
# Find our lane pixels first
leftx, lefty, rightx, righty, out_img = find_lines(binary)
# covert x, y from pixels to meters
ym_per_pix = 30/720 # meters per pixel in y dimension
xm_per_pix = 3.7/700 # meters per pixel in x dimension
# Fit a second order polynomial to each using `np.polyfit`
left_fit_cr = np.polyfit(lefty*ym_per_pix, leftx*xm_per_pix, 2)
right_fit_cr = np.polyfit(righty*ym_per_pix, rightx*xm_per_pix, 2)
# Generate x and y values for plotting
ploty = np.linspace(0, binary.shape[0]-1, binary.shape[0] )
y_eval=np.max(ploty)
left_curverad = ((1 + (2*left_fit_cr[0]*y_eval*ym_per_pix + left_fit_cr[1])**2)**1.5) / np.absolute(2*left_fit_cr[0])
right_curverad = ((1 + (2*right_fit_cr[0]*y_eval*ym_per_pix + right_fit_cr[1])**2)**1.5) / np.absolute(2*right_fit_cr[0])
return left_curverad, right_curverad
# -
# ## Pipeline (Video)
# +
# importing some useful packages
#from moviepy.editor import VideoFileClip
import imageio
imageio.plugins.ffmpeg.download()
import matplotlib.pyplot as plt
import numpy as np
import matplotlib.image as mpimg
import cv2
import os
from moviepy.editor import VideoFileClip
import glob
## Apply Camera calibration, distortion correction
def undistortion(img):
# Arrays to store object points and image points from all the images.
objpoints = np.load('objpoints.npy') # 3d points in real world space
imgpoints = np.load('imgpoints.npy') # 2d points in image plane.
shape=tuple(np.load('shape.npy'))
ret, mtx, dist, rvecs, tvecs = cv2.calibrateCamera(objpoints, imgpoints,shape,None, None)
undist=cv2.undistort(img, mtx, dist, None, mtx)
return undist
thresh_grad_min = 20
thresh_grad_max = 100
color_thre_min, color_thre_max=170, 255
## Define Color & gradients function
def gradient_threshold(img, thresh_grad_min, thresh_grad_max, color_thre_min, color_thre_max):
hls=cv2.cvtColor(img, cv2.COLOR_RGB2HLS)
s_channel=hls[:,:,2]
gray=cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
## Sobelx
sobelx = cv2.Sobel(gray, cv2.CV_64F, 1, 0) # Take the derivative in x
abs_sobelx = np.absolute(sobelx) # Absolute x derivative to accentuate lines away from horizontal
scaled_sobel = np.uint8(255*abs_sobelx/np.max(abs_sobelx))
# Threshold x gradient
sxbinary = np.zeros_like(scaled_sobel)
sxbinary[(scaled_sobel >= thresh_grad_min) & (scaled_sobel <= thresh_grad_max)] = 1
# Color threshold
s_binary=np.zeros_like(s_channel)
s_binary[(s_channel>=color_thre_min)&(s_channel<color_thre_max)]=1
# stack each channel to an image to see the contribution of individual channels
color_binary = np.dstack(( np.zeros_like(sxbinary), sxbinary, s_binary)) * 255
# combine two binary thresholds
combined_binary=np.zeros_like(s_binary)
combined_binary[(sxbinary==1)|(s_binary==1)]=1
return combined_binary
## Perspective transform
## Define the function for image unwarp
def warped_img(img):
gray=cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
img_size=(gray.shape[1],gray.shape[0])
#src=np.float32([[500,500],[780,500],[img.shape[1],img.shape[0]],[150,img.shape[0]]])
src=np.float32([[580,460],[700,460],[1040,680],[260,680]])
dst = np.array([[200, 0], [1000, 0], [1000, img.shape[0]], [200, img.shape[0]]], np.float32)
# Given src and dst points, calculate the perspective transform matrix
M = cv2.getPerspectiveTransform(src, dst)
Minv = cv2.getPerspectiveTransform(dst, src)
# Warp the image using OpenCV warpPerspective()
warped = cv2.warpPerspective(img, M, img_size)
return warped, M, Minv
## Define the function of detecting lane lines
def find_lines(binary):
# calculate the current base leftx and rightx pixels location
image_height=binary.shape[0]
image_width=binary.shape[1]
histogram=np.sum(binary[binary.shape[0]//2:,:], axis=0)
midpoint=np.int(binary.shape[1]//2)
leftx_base=np.argmax(histogram[:midpoint]) # define the base left lanes location
rightx_base=np.argmax(histogram[midpoint:])+midpoint # define the base right lanes location
leftx_current=leftx_base
rightx_current=rightx_base
out_img=np.dstack((binary, binary, binary))
# nonzero x & y locations
nonzerox=np.array(binary.nonzero()[1])
nonzeroy=np.array(binary.nonzero()[0])
# left and right lane lines indices
leftx_ind, rightx_ind=[],[]
# Hyper parameters for windows
nwindow=9
margin=100
minpxl=50
for window in range(nwindow):
# define the parameters of windwos on image
win_height=np.int(image_height//nwindow)
win_y_low=image_height-(window+1)*win_height
win_y_high=image_height-window*win_height
win_left_low=leftx_current-margin
win_left_high=leftx_current+margin
win_right_low=rightx_current-margin
win_right_high=rightx_current+margin
# identify the nonzero pixels within the window
detected_leftx_ind=((nonzeroy >= win_y_low) & (nonzeroy < win_y_high) &
(nonzerox >= win_left_low) & (nonzerox < win_left_high)).nonzero()[0]
detected_rightx_ind=((nonzeroy >= win_y_low) & (nonzeroy < win_y_high) &
(nonzerox >= win_right_low) & (nonzerox < win_right_high)).nonzero()[0]
if len(detected_leftx_ind)>minpxl:
leftx_current=np.int(np.mean(nonzerox[detected_leftx_ind]))
if len(detected_rightx_ind)>minpxl:
rightx_current=np.int(np.mean(nonzerox[detected_rightx_ind]))
leftx_ind.append(detected_leftx_ind)
rightx_ind.append(detected_rightx_ind)
try:
leftx_ind=np.concatenate(leftx_ind)
rightx_ind=np.concatenate(rightx_ind)
except ValueErrors:
pass
leftx=nonzerox[leftx_ind]
rightx=nonzerox[rightx_ind]
lefty=nonzeroy[leftx_ind]
righty=nonzeroy[rightx_ind]
return leftx, lefty, rightx, righty, out_img
# define polynomial function of lane lines
def polynomial_fit(binary, Minv):
# Find our lane pixels first
leftx, lefty, rightx, righty, out_img = find_lines(binary)
# Fit a second order polynomial to each using `np.polyfit`
left_fit = np.polyfit(lefty, leftx, 2)
right_fit = np.polyfit(righty, rightx, 2)
# Generate x and y values for plotting
ploty = np.linspace(0, binary.shape[0]-1, binary.shape[0] )
try:
left_fitx = left_fit[0]*ploty**2 + left_fit[1]*ploty + left_fit[2]
right_fitx = right_fit[0]*ploty**2 + right_fit[1]*ploty + right_fit[2]
except TypeError:
# Avoids an error if `left` and `right_fit` are still none or incorrect
print('The function failed to fit a line!')
left_fitx = 1*ploty**2 + 1*ploty
right_fitx = 1*ploty**2 + 1*ploty
# Create an image to draw the lines on
warp_zero = np.zeros_like(binary).astype(np.uint8)
color_warp = np.dstack((warp_zero, warp_zero, warp_zero))
# Recast the x and y points into usable format for cv2.fillPoly()
pts_left = np.array([np.transpose(np.vstack([left_fitx, ploty]))])
pts_right = np.array([np.flipud(np.transpose(np.vstack([right_fitx, ploty])))])
pts = np.hstack((pts_left, pts_right))
# Draw the lane onto the warped blank image
cv2.fillPoly(color_warp, np.int_([pts]), (0,255, 0))
# Warp the blank back to original image space using inverse perspective matrix (Minv)
newwarp = cv2.warpPerspective(color_warp, Minv, (binary.shape[1], binary.shape[0]))
## Visualization ##
# Colors in the left and right lane regions
out_img[lefty, leftx] = [255, 0, 0]
out_img[righty, rightx] = [0, 0, 255]
# Plots the left and right polynomials on the lane lines
#plt.plot(left_fitx, ploty, color='yellow')
#plt.plot(right_fitx, ploty, color='yellow')
return out_img, newwarp
def curvature_cal(binary):
# Find our lane pixels first
leftx, lefty, rightx, righty, out_img = find_lines(binary)
# covert x, y from pixels to meters
ym_per_pix = 30/720 # meters per pixel in y dimension
xm_per_pix = 3.7/700 # meters per pixel in x dimension
image_height=binary.shape[0]
image_width=binary.shape[1]
histogram=np.sum(binary[binary.shape[0]//2:,:], axis=0)
midpoint=np.int(binary.shape[1]//2)
leftx_base=np.argmax(histogram[:midpoint]) # define the base left lanes location
rightx_base=np.argmax(histogram[midpoint:])+midpoint # define the base right lanes location
lane_center=np.int((leftx_base+rightx_base)//2)
img_center=np.int(image_width//2)
Veh_pos=np.float32((img_center-lane_center)*xm_per_pix) # calculate the vehicel position, left '-', right off center '+'
# Fit a second order polynomial to each using `np.polyfit`
left_fit_cr = np.polyfit(lefty*ym_per_pix, leftx*xm_per_pix, 2)
right_fit_cr = np.polyfit(righty*ym_per_pix, rightx*xm_per_pix, 2)
# Generate x and y values for plotting
ploty = np.linspace(0, binary.shape[0]-1, binary.shape[0] )
y_eval=np.max(ploty)
left_curverad = ((1 + (2*left_fit_cr[0]*y_eval*ym_per_pix + left_fit_cr[1])**2)**1.5) / np.absolute(2*left_fit_cr[0])
right_curverad = ((1 + (2*right_fit_cr[0]*y_eval*ym_per_pix + right_fit_cr[1])**2)**1.5) / np.absolute(2*right_fit_cr[0])
lane_curverad=(left_curverad+right_curverad)/2
return lane_curverad, Veh_pos
def process_image(img):
# print out some info about this image
#img=cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
# transform from color to gray scale
#b,g,r=cv2.split(img)
#img=cv2.merge([r,g,b])
img_gray=cv2.cvtColor(img,cv2.COLOR_RGB2GRAY) #mpimg loads image as RGB
# Apply Gaussian smoothing
#kernel_size=5
#blur_gray=cv2.GaussianBlur(img_gray,(kernel_size,kernel_size),0,0)
# Undistort the image
undistort_img=undistortion(img)
#color & gradients threshold
img_warped, perspective_M, Minv=warped_img(undistort_img)
warped_binary=gradient_threshold(img_warped, thresh_grad_min, thresh_grad_max, color_thre_min, color_thre_max)
out_img, newwarp=polynomial_fit(warped_binary,Minv)
lane_curverad, Veh_pos=curvature_cal(warped_binary)
# Combine the result with the original image
result = cv2.addWeighted(undistort_img, 1, newwarp, 0.3, 0)
if Veh_pos > 0:
car_pos_text = "{:04.2f}m right off center".format(Veh_pos)
else:
car_pos_text = '{:04.3f}m left off center'.format(abs(Veh_pos))
cv2.putText(result, "Lane curve: {:04.2f}m".format(lane_curverad), (10, 50), cv2.FONT_HERSHEY_SIMPLEX, 1.5,
color=(255, 255, 255), thickness=2)
cv2.putText(result, "Car is {}".format(car_pos_text), (10, 100), cv2.FONT_HERSHEY_SIMPLEX, 1.5, color=(255, 255, 255),
thickness=2)
return result
# +
from IPython.display import HTML
white_output = 'final_result.mp4'
## To speed up the testing process you may want to try your pipeline on a shorter subclip of the video
## To do so add .subclip(start_second,end_second) to the end of the line below
## Where start_second and end_second are integer values representing the start and end of the subclip
## You may also uncomment the following line for a subclip of the first 5 seconds
##clip1 = VideoFileClip("test_videos/solidWhiteRight.mp4").subclip(0,5)
clip1 = VideoFileClip("../project_video.mp4")
white_clip = clip1.fl_image(process_image) #NOTE: this function expects color images!!
# %time white_clip.write_videofile(white_output, audio=False)
HTML("""
<video width="960" height="540" controls>
<source src="{0}">
</video>
""".format(white_output))
# -
| examples/.ipynb_checkpoints/example-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/BaishaliChetia/CapsNet-Keras/blob/master/capsFashionMnistipynb.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="3qZ5-69C1ggq"
import numpy as np
import tensorflow as tf
import pandas as pd
import tensorflow.keras as K
#import tensorflow_model_optimization as tfmot
# + colab={"base_uri": "https://localhost:8080/"} id="bOK2RyTe4v8Q" outputId="835a23b1-fe4d-4f59-de90-99077fd440a9"
pip install -q tensorflow-model-optimization
# + id="wIMpFAPJ40X5"
import tensorflow_model_optimization as tfmot
# + id="LUjiHNeK4WAv"
caps1_n_maps = 32
caps1_n_caps = caps1_n_maps * 6 * 6 # 1152 primary capsules
caps1_n_dims = 8
caps2_n_caps = 10
caps2_n_dims = 16
tf.random.set_seed(500000)
# + id="zUmUW99P4YQt"
# + id="sukwGEY4MlPV"
#class SquashLayer(K.layers.Layer, tfmot.sparsity.keras.PrunableLayer):
class SquashLayer(K.layers.Layer):
def __init__(self, axis=-1, **kwargs):
super(SquashLayer, self).__init__(**kwargs)
self.axis = axis
def build(self, input_shapes):
pass
"""
def get_prunable_weights(self):
return []
"""
def call(self, inputs):
EPSILON = 1.0e-9
squared_norm = tf.compat.v1.reduce_sum(tf.square(inputs),\
axis=self.axis,\
keepdims=True)
safe_norm = tf.sqrt(squared_norm + EPSILON)
squash_factor = squared_norm / (1. + squared_norm)
unit_vector = inputs / safe_norm
return squash_factor * unit_vector
def get_config(self):
config = super(SquashLayer, self).get_config()
config.update({"axis": self.axis})
return config
#class SafeNorm(K.layers.Layer, tfmot.sparsity.keras.PrunableLayer):
class SafeNorm(K.layers.Layer):
def __init__(self, axis=-1, keep_dims = False, **kwargs):
super(SafeNorm, self).__init__(**kwargs)
self.axis = axis
self.keep_dims = keep_dims
def build(self, input_shapes):
pass
"""
def get_prunable_weights(self):
return []
"""
def call(self, input):
EPSILON = 1.0e-9
squared_norm = tf.compat.v1.reduce_sum(tf.square(inputs),\
axis=self.axis,\
keepdims= self.keep_dims)
safe_norm = tf.sqrt(squared_norm + EPSILON)
return safe_norm
def get_config(self):
config = super(SafeNorm, self).get_config()
config.update({"axis": self.axis, "keep_dims": self.keep_dims})
return config
# This should be the part where the digit layer, and where we tile things
# This is incomplete, and work in progress
# TODO: Complete this
class MyDigitCapsLayer(K.layers.Layer, tfmot.sparsity.keras.PrunableLayer):
def __init__(self, **kwargs):
super(MyDigitCapsLayer, self).__init__(**kwargs)
def get_config(self):
config = super(MyDigitCapsLayer, self).get_config()
return config
def build(self, input_shapes):
init_sigma = 0.1 # TODO: use
self.kernel = self.add_weight(\
"kernel",\
(caps1_n_caps, caps2_n_caps, caps2_n_dims, caps1_n_dims),\
initializer="random_normal",\
dtype=tf.float32)
# To debug this function, I used prints to print the shape
# expand_dims just adds an exis, so if you say expand_dims(inshape=(5, 3), -1),
# you get the output shape (5, 3, 1), it just adds an axis at the end
# Then tile just multiplies one of the dimensions (that is it stacks along that direction N times)
# so tile(inshape=(5, 3, 1), [1, 1, 1000]) will yield a shape (5, 3, 1000)
#
# Notice I didn't tile in build, but in call, Most probaly this is the right thing to do
# but we'll only figure out when we actually train
def get_prunable_weights(self):
return [self.kernel]
def call(self, inputs):
# Add a dimension at the end
exp1 = tf.expand_dims(inputs, -1, name="caps1_output_expanded")
# add a dimension along 3rd axis
exp1 = tf.expand_dims(exp1, 2, name="caps2_output_espanced")
# tile along 3rd axis
tile = tf.tile(exp1, [1, 1, caps2_n_caps, 1, 1], name="caps1_output_tiled")
caps2_predicted = tf.matmul(self.kernel, tile, name="caps2_predicted")
return caps2_predicted
# https://www.tensorflow.org/api_docs/python/tf/keras/losses/Loss
class MarginLoss(K.losses.Loss):
def __init__(self, **kwargs):
super(MarginLoss, self).__init__(**kwargs)
def get_config(self):
config = super(MarginLoss, self).get_config()
return config
def safe_norm(self, input, axis=-2, epsilon=1e-5, keep_dims=False, name=None):
squared_norm = tf.reduce_sum(tf.square(input), axis=axis,
keepdims=keep_dims)
return tf.sqrt(squared_norm + epsilon)
"""
def get_prunable_weights(self):
return []
"""
def call(self,y_true, input):
# print(f"y_true.shape = {y_true.shape}, y_pred.shape = {y_pred.shape}")
# return K.losses.MeanSquaredError()(y_true, y_pred)
#y_true = K.Input(shape=[], dtype=tf.int64, name="y")
m_plus = 0.9
m_minus = 0.1
lambda_ = 0.5
#y_true one hot encode y_train
T = tf.one_hot(y_true, depth=caps2_n_caps, name="T")
caps2_output_norm = self.safe_norm(input, keep_dims = True)
present_error_raw = tf.square(\
tf.maximum(0., m_plus - caps2_output_norm),
name="present_error_raw")
present_error = tf.reshape(\
present_error_raw, shape=(-1, 10),
name="present_error")
absent_error_raw = tf.square(\
tf.maximum(0., caps2_output_norm - m_minus),
name="absent_error_raw")
absent_error = tf.reshape(\
absent_error_raw, shape=(-1, 10),
name="absent_error")
L = tf.add(\
T * present_error,\
lambda_ * (1.0 - T) * absent_error,
name="L")
marginLoss = tf.reduce_mean(\
tf.reduce_sum(L, axis=1),\
name="margin_loss")
return marginLoss
#class RoutingByAgreement(K.layers.Layer, tfmot.sparsity.keras.PrunableLayer):
class RoutingByAgreement(K.layers.Layer):
def __init__(self, round_number=-1, **kwargs):
super(RoutingByAgreement, self).__init__(**kwargs)
self.round_number = round_number
def get_config(self):
config = super(RoutingByAgreement, self).get_config()
config.update({"round_number": self.round_number})
return config
def build(self, input_shapes):
self.raw_weights_1 = self.add_weight("raw_weights", \
(caps1_n_caps, caps2_n_caps, 1, 1), \
initializer = "zeros", \
dtype=tf.float32,)
#print("Routing layer: self.raw_weights = ", self.raw_weights.shape, "input_shape = ", input_shapes)
def get_prunable_weights(self):
return [self.raw_weights_1]
@staticmethod
def squash(inputs):
EPSILON = 1.0e-5
squared_norm = tf.compat.v1.reduce_sum(tf.square(inputs),\
keepdims=True)
safe_norm = tf.sqrt(squared_norm + EPSILON)
squash_factor = squared_norm / (1. + squared_norm)
unit_vector = inputs / safe_norm
return squash_factor * unit_vector
def single_round_routing(self, inputs, raw_weights, agreement):
raw_weights = tf.add(raw_weights, agreement)
routing_wt = tf.nn.softmax(raw_weights, axis=2)
wt_predictions = tf.multiply(routing_wt, inputs)
wt_sum = tf.reduce_sum(wt_predictions, axis=1, keepdims=True)
return wt_sum
def call(self, inputs):
agreement = tf.zeros(shape=self.raw_weights_1.shape)
sqsh_wt_sum = None
x = inputs
for i in range(2):
wt_sum = self.single_round_routing(inputs, self.raw_weights_1, agreement)
sqsh_wt_sum = RoutingByAgreement.squash(wt_sum)
sqsh_wt_sum_tiled = tf.tile(\
sqsh_wt_sum ,\
[1, caps1_n_caps, 1, 1, 1],\
name="caps2_output_round_1_tiled")
agreement = tf.matmul(\
x, \
sqsh_wt_sum_tiled,\
transpose_a=True,\
name="agreement")
return sqsh_wt_sum
class MyAccuracy(K.metrics.Metric):
def __init__(self, **kwargs):
super(MyAccuracy, self).__init__(**kwargs)
self.acc_obj = K.metrics.Accuracy()
self.state = 0
def get_config(self):
config = super(MyAccuracy, self).get_config()
config.update({"acc_obj": None, "state": self.state})
return config
def safe_norm(self, input, axis=-2, epsilon=1e-5, keep_dims=True, name=None):
squared_norm = tf.reduce_sum(tf.square(input), axis=axis,
keepdims=keep_dims)
return tf.sqrt(squared_norm + epsilon)
def update_state(self, y_true, input, sample_weight=None):
if self.acc_obj is None:
self.acc_obj = K.metrics.Accuracy()
y_proba = self.safe_norm(input, axis=-2)
y_proba_argmax = tf.argmax(y_proba, axis=2)
y_pred = tf.squeeze(y_proba_argmax, axis=[1,2], name="y_pred")
#y_true = tf.reshape(y_true, (y_true.shape[0], ))
y_true = tf.cast(y_true, dtype=tf.int64)
self.acc_obj.update_state(y_true, y_pred, sample_weight)
def reset_state(self):
self.acc_obj.reset_state()
def result(self):
return self.acc_obj.result()
class MyReshapeLayer(K.layers.Layer):
def __init__(self, axis=-1, keep_dims = False, **kwargs):
super(MyReshapeLayer, self).__init__(**kwargs)
def build(self, input_shapes):
pass
def safe_norm(self, input, axis=-2, epsilon=1e-5, keep_dims=True, name=None):
squared_norm = tf.reduce_sum(tf.square(input), axis=axis,
keepdims=keep_dims)
return tf.sqrt(squared_norm + epsilon)
def call(self, input):
print('printing shapes ------------------- ')
EPSILON = 1.0e-9
print(input)
y_proba = self.safe_norm(input, axis=-2)
print(y_proba)
y_proba_argmax = tf.argmax(y_proba, axis=2)
print(y_proba_argmax)
y_pred = tf.squeeze(y_proba_argmax, axis=[1,2], name="y_pred")
print(y_pred)
return tf.cast(y_pred, tf.int64)
def get_config(self):
config = super(MyReshapeLayer, self).get_config()
return config
# + id="-HEFxQ-Z4eD7"
# + colab={"base_uri": "https://localhost:8080/"} id="aSEe-231jn49" outputId="c9711cb3-69a9-4f4c-ab09-372f01cd6cc3"
(x_train, y_train,), (x_test, y_test) = K.datasets.fashion_mnist.load_data()
#print(x_train.shape, x_test.shape)
x_train = x_train/255.0
x_test = x_test/255.0
# + id="42mw5bOB5OXs"
# + colab={"base_uri": "https://localhost:8080/"} id="UnmSudqTMlPX" outputId="1dfa624a-6d38-4b22-a329-56f044bd4212"
class Model:
@staticmethod
def build(inshape=(28, 28, 1)):
inp = K.Input(shape=inshape, dtype=tf.float32, name='input')
# Primary capsules
# For each digit in the batch
# 32 maps, each 6x6 grid of 8 dimensional vectors
# First Conv layer
conv1_params = \
{
"filters": 256,
"kernel_size": 9,
"strides": 1,
"padding": "valid",
"activation": tf.nn.relu,
}
x = K.layers.Conv2D(**conv1_params, name="conv_layer_1")(inp)
# Second conv layer
conv2_params = \
{
"filters": caps1_n_maps * caps1_n_dims, # 256 convolutional filters
"kernel_size": 9,
"strides": 2,
"padding": "valid",
"activation": tf.nn.relu
}
x = K.layers.Conv2D(**conv2_params, name="conv_layer_2")(x)
# Reshape
x = K.layers.Reshape(\
(caps1_n_caps, caps1_n_dims),\
name="reshape_layer_1")(x)
x = SquashLayer(name="caps1_output_layer")(x)
x = MyDigitCapsLayer(name = "caps2_predicted")(x)
caps2_predicted = x # Save this value for later
#routing by agreement (2 rounds)
x = RoutingByAgreement(name="routing1", round_number=2)(x)
return K.Model(inputs=inp, outputs=x, name='my_model')
m = Model.build()
print(m.summary())
# + id="-Kdi6KMY5dw6"
# + id="ohNmlixkrAek"
from keras.callbacks import ModelCheckpoint, CSVLogger
comparison_metric = MyAccuracy()
#checkpoint_filepath = "/content/drive/MyDrive/Weights/weights-improvement-{epoch:02d}-{val_my_accuracy:.2f}.hdf5"
model_checkpoint_callback = tf.keras.callbacks.ModelCheckpoint(
filepath = "/content/drive/MyDrive/FashionMnistResults/best_weights1.hdf5",
save_weights_only=True,
#monitor=f"val_{comparison_metric.name}",
monitor="val_my_accuracy",
mode='max',
save_best_only=True)
model_checkpoint_callback2 = tf.keras.callbacks.ModelCheckpoint(
filepath = "/content/drive/MyDrive/FashionMnistResults/latest_weights1.hdf5",
save_weights_only=True,
monitor=f"val_{comparison_metric.name}",
mode='max',
save_best_only=False)
log_csv = CSVLogger("/content/drive/MyDrive/FashionMnistResults/mylogs1.csv", separator = ",", append = False)
callback_list = [model_checkpoint_callback, model_checkpoint_callback2, log_csv]
# + id="LYB8WPX95xWp"
# + colab={"base_uri": "https://localhost:8080/"} id="cahMgtOqMlPa" outputId="3fed3c7a-6aa3-43a6-b16c-e66515aa1a6f"
m.compile(optimizer='adam', loss=MarginLoss(), metrics=[MyAccuracy()])
history = m.fit(x_train, y_train, batch_size=32, epochs=70, verbose= 1, validation_split=0.2, callbacks = callback_list)
# + colab={"base_uri": "https://localhost:8080/"} id="GksD0OMC3hjU" outputId="42d346db-c62b-41f9-f0b6-943a107f4931"
print(f'Best Validation Accuracy = {np.max(history.history["val_my_accuracy_2"])}')
print(f'Best Training Accuracy = {np.max(history.history["my_accuracy_2"])}')
# + colab={"base_uri": "https://localhost:8080/", "height": 648} id="r7XqyAli6Dza" outputId="a16b2f28-cb49-422a-d8bc-82278503b8df"
import matplotlib.pyplot as plt
plt.rcParams["figure.figsize"] = (30, 10)
plt.rcParams["font.size"] = 20
fig, ax = plt.subplots(1, 2)
ax[0].plot(history.history['my_accuracy_2'])
ax[0].plot(history.history['val_my_accuracy_2'])
ax[0].set_title('Model Accuracy')
ax[0].set_ylabel('Accuracy')
ax[0].set_xlabel('Epoch')
ax[0].legend(['Training Accuracy', 'Validation Accuracy'], loc='best')
ax[1].plot(history.history['loss'])
ax[1].plot(history.history['val_loss'])
ax[1].set_title('Model Loss')
ax[1].set_ylabel('Loss')
ax[1].set_xlabel('Epoch')
ax[1].legend(['Training Loss', 'Validation Loss'], loc='best')
plt.show()
| capsFashionMnistipynb.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="pr1eEWIUjiBc" colab_type="text"
# # Earth Engine REST API Quickstart
#
# This is a demonstration notebook for using the Earth Engine REST API. See the complete guide for more information: https://developers.google.com/earth-engine/reference/Quickstart.
#
# + [markdown] id="OfMAA6YhPuFl" colab_type="text"
# ## Authentication
#
# The first step is to choose a project and login to Google Cloud.
# + id="FRm2HczTIlKe" colab_type="code" colab={}
# INSERT YOUR PROJECT HERE
PROJECT = 'your-project'
# !gcloud auth login --project {PROJECT}
# + [markdown] id="hnufOtSfP0jX" colab_type="text"
# ## Define service account credentials
# + id="tLxOnKL2Nk5p" colab_type="code" colab={}
# INSERT YOUR SERVICE ACCOUNT HERE
SERVICE_ACCOUNT='your-service-account@your-project.iam.gserviceaccount.com'
KEY = 'private-key.json'
# !gcloud iam service-accounts keys create {KEY} --iam-account {SERVICE_ACCOUNT}
# + [markdown] id="6QksNfvaY5em" colab_type="text"
# ## Create an authorized session to make HTTP requests
# + id="h2MHcyeqLufx" colab_type="code" colab={}
from google.auth.transport.requests import AuthorizedSession
from google.oauth2 import service_account
credentials = service_account.Credentials.from_service_account_file(KEY)
scoped_credentials = credentials.with_scopes(
['https://www.googleapis.com/auth/cloud-platform'])
session = AuthorizedSession(scoped_credentials)
url = 'https://earthengine.googleapis.com/v1alpha/projects/earthengine-public/assets/LANDSAT'
response = session.get(url)
from pprint import pprint
import json
pprint(json.loads(response.content))
# + [markdown] id="_KjWa7KJY_7m" colab_type="text"
# ## Get a list of images at a point
# + [markdown] id="5kKbIDctpZeH" colab_type="text"
# Query for Sentinel-2 images at a specific location, in a specific time range and with estimated cloud cover less than 10%.
# + id="0bENTPjMQr5h" colab_type="code" colab={}
import urllib
coords = [-122.085, 37.422]
project = 'projects/earthengine-public'
asset_id = 'COPERNICUS/S2'
name = '{}/assets/{}'.format(project, asset_id)
url = 'https://earthengine.googleapis.com/v1alpha/{}:listImages?{}'.format(
name, urllib.parse.urlencode({
'startTime': '2017-04-01T00:00:00.000Z',
'endTime': '2017-05-01T00:00:00.000Z',
'region': '{"type":"Point", "coordinates":' + str(coords) + '}',
'filter': 'CLOUDY_PIXEL_PERCENTAGE < 10',
}))
response = session.get(url)
content = response.content
for asset in json.loads(content)['images']:
id = asset['id']
cloud_cover = asset['properties']['CLOUDY_PIXEL_PERCENTAGE']
print('%s : %s' % (id, cloud_cover))
# + [markdown] id="DY0MfkjiAAW_" colab_type="text"
# ## Inspect an image
#
# Get the asset name from the previous output and request its metadata.
# + id="ddzrXIl4ADLk" colab_type="code" colab={}
asset_id = 'COPERNICUS/S2/20170430T190351_20170430T190351_T10SEG'
name = '{}/assets/{}'.format(project, asset_id)
url = 'https://earthengine.googleapis.com/v1alpha/{}'.format(name)
response = session.get(url)
content = response.content
asset = json.loads(content)
print('Band Names: %s' % ','.join(band['id'] for band in asset['bands']))
print('First Band: %s' % json.dumps(asset['bands'][0], indent=2, sort_keys=True))
# + [markdown] id="O5I63cC6ZDQn" colab_type="text"
# ## Get pixels from one of the images
# + id="xJhv6bfEZHa2" colab_type="code" colab={}
import numpy
import io
name = '{}/assets/{}'.format(project, asset_id)
url = 'https://earthengine.googleapis.com/v1alpha/{}:getPixels'.format(name)
body = json.dumps({
'fileFormat': 'NPY',
'bandIds': ['B2', 'B3', 'B4', 'B8'],
'grid': {
'affineTransform': {
'scaleX': 10,
'scaleY': -10,
'translateX': 499980,
'translateY': 4200000,
},
'dimensions': {'width': 256, 'height': 256},
},
})
pixels_response = session.post(url, body)
pixels_content = pixels_response.content
array = numpy.load(io.BytesIO(pixels_content))
print('Shape: %s' % (array.shape,))
print('Data:')
print(array)
# + [markdown] id="jcwE2W8kFojo" colab_type="text"
# ## Get a thumbnail of an image
#
# Note that `name` and `asset` are already defined from the request to get the asset metadata.
# + id="xs0ZHHKmFovV" colab_type="code" colab={}
url = 'https://earthengine.googleapis.com/v1alpha/{}:getPixels'.format(name)
body = json.dumps({
'fileFormat': 'PNG',
'bandIds': ['B4', 'B3', 'B2'],
'region': asset['geometry'],
'grid': {
'dimensions': {'width': 256, 'height': 256},
},
'visualizationOptions': {
'ranges': [{'min': 0, 'max': 3000}],
},
})
image_response = session.post(url, body)
image_content = image_response.content
# Import the Image function from the IPython.display module.
from IPython.display import Image
Image(image_content)
| python/examples/ipynb/Earth_Engine_REST_API_Quickstart.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
# %config InlineBackend.figure_format = "retina"
import matplotlib.pyplot as plt
#import seaborn as sns
#sns.set_style('ticks')
import numpy as np
import scipy.integrate as integ
from astropy.io import ascii
#from scipy import interpolate
#import scipy.stats as stats
#from astropy.table import Table, Column
#import readsnap as rs
#reload(rs)
plt.rcParams['figure.figsize'] = (8,5)
plt.rcParams['legend.frameon'] = False
plt.rcParams['legend.fontsize'] = 15
plt.rcParams['legend.borderpad'] = 0.1
plt.rcParams['legend.labelspacing'] = 0.1
plt.rcParams['legend.handletextpad'] = 0.1
plt.rcParams['font.family'] = 'stixgeneral'
plt.rcParams['font.size'] = 15
plt.rcParams['lines.linewidth'] = 2
plt.rcParams['axes.labelsize'] = 15
# CGS
G = 6.6726e-08 # [G]=cm^3 g^-1 s^-2
Ro = 6.96e10
Mo = 1.99e33
c = 2.9979e+10
day = 60*60*24
# -
def readDataHeger(filename):
# Alejandro - 16/06/2020
# Values taken from Aldo's script. Double check.
data = np.genfromtxt(filename)
m = data[:,1] # cell outer total mass
r = data[:,2] # cell outer radius
v = data[:,3] # cell outer velocity
rho = data[:,4] # cell density
Omega = data[:,9] #5*s26_data[:,9] # cell specific angular momentum
jprofile = (2./3.)*Omega*r**2
T = data[:,5] # cell temperature
p = data[:,6] # cell pressure
e = data[:,7] # cell specific energy
S = data[:,8] # cell specific entropy
return m,r,v,rho,Omega,jprofile,T,p,e,S
def calculateBindingEnergy(m,r):
Min = m[::-1]
rin = r[::-1]
E_bind = integ.cumtrapz(-G*Min/rin,Min)
E_bind = E_bind[::-1]
E_bind = np.append(E_bind,E_bind[-1])
return E_bind
# Load Alex's models
mHE16C,rHE16C,vHE16C,rhoHE16C,OmegaHE16C,jprofileHE16C,THE16C,pHE16C,eHE16C,SHE16C = readDataHeger('../stellarProfiles/35OC@presn') # Temp hack
m12SH,r12SH,v12SH,rho12SH,Omega12SH,jprofile12SH,T12SH,p12SH,e12SH,S12SH = readDataHeger('../stellarProfiles/12SH@presn')
m12SF,r12SF,v12SF,rho12SF,Omega12SF,jprofile12SF,T12SF,p12SF,e12SF,S12SF = readDataHeger('../stellarProfiles/12SF@presn')
plt.plot(rHE16C/Ro,mHE16C/Mo)
plt.plot(r12SH/Ro,m12SH/Mo)
plt.plot(r12SF/Ro,m12SF/Mo)
plt.xlabel("Radial coordinate [Rsol]")
plt.ylabel("Mass coordinate [Msol]")
plt.legend(("HE16C","12SH","12SF"))
#plt.semilogx()
# +
plt.plot(mHE16C/Mo,((rHE16C/Ro)**3)*rhoHE16C)
plt.axvline(x=1.0,color='r',label='1 Msol')
plt.axvline(x=1.33,color='g',label='1.33 Msol')
plt.axvline(x=2.0,color='k',label='2 Msol')
plt.ylabel("Density*R^3 [g]")
plt.xlabel("Mass coordinate [Msol]")
plt.semilogy()
plt.legend()
plt.figure(2)
plt.plot(mHE16C/Mo,rHE16C/Ro)
plt.axvline(x=1.0,color='r',label='1 Msol')
plt.axvline(x=1.3,color='g',label='1.3 Msol')
# plt.axvline(x=2.0,color='k',label='2 Msol')
# plt.plot(m12SH/Mo,r12SH/Ro)
# plt.plot(r12SF/Ro,m12SF/Mo)
plt.ylabel("Radial coordinate [Rsol]")
plt.xlabel("Mass coordinate [Msol]")
plt.semilogy()
plt.legend(("HE16C"))
# -
plt.plot(rHE16C/Ro,rhoHE16C,'--')
plt.ylabel("Density [g cm^-3]")
plt.xlabel("Radial coordinate [Rsol]")
plt.semilogy()
E_bindHE16C = calculateBindingEnergy(mHE16C,rHE16C)
E_bind12SH = calculateBindingEnergy(m12SH,r12SH)
E_bind12SF = calculateBindingEnergy(m12SF,r12SF)
plt.plot(mHE16C/Mo,E_bindHE16C)
# plt.plot(m12SH/Mo,E_bind12SH)
# plt.plot(m12SF/Mo,E_bind12SF)
plt.ylabel("E_bind [ergs?]")
plt.xlabel("Mass coordinate [Msol]")
plt.legend(("HE16C","12SH","12SF"))
plt.semilogy()
# plt.ylim(1e49,1e52)
pwd
a = np.asarray([ rHE16C/Ro, mHE16C/Mo, rhoHE16C, eHE16C])
np.savetxt("mHE16C.csv", a, delimiter=",")
| scripts/postProcessing/PlottingHegerModels.ipynb |