code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Write a Python program to get the Fibonacci series between 0 to 50
# +
def f(n):
if n==0: return 0
elif n==1: return 1
else: return f(n-1)+f(n-2)
print(f(10))
# -
| condition and loop/fibonaci_generator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] nbsphinx="hidden"
# # Vitessce Widget Tutorial
# -
# # Visualization of single-cell RNA seq data
# ## 1. Import dependencies
#
# We need to import the classes and functions that we will be using from the corresponding packages.
# +
import os
from os.path import join
from urllib.request import urlretrieve
from anndata import read_h5ad
import scanpy as sc
from vitessce import (
VitessceConfig,
Component as cm,
CoordinationType as ct,
AnnDataWrapper,
)
# -
# ## 2. Download the data
#
# For this example, we need to download a dataset from the COVID-19 Cell Atlas https://www.covid19cellatlas.org/index.healthy.html#habib17.
os.makedirs("data", exist_ok=True)
adata_filepath = join("data", "habib17.processed.h5ad")
urlretrieve('https://covid19.cog.sanger.ac.uk/habib17.processed.h5ad', adata_filepath)
# ## 3. Load the data
#
# Note: this function may print a `FutureWarning`
adata = read_h5ad(adata_filepath)
# ## 3.1. Preprocess the Data For Visualization
#
# This dataset contains 25,587 genes. In order to visualize it efficiently, we convert it to CSC sparse format so that we can make fast requests for gene data. We also prepare to visualize the top 50 highly variable genes for the heatmap as ranked by dispersion norm, although one may use any boolean array filter for the heatmap.
top_dispersion = adata.var["dispersions_norm"][
sorted(
range(len(adata.var["dispersions_norm"])),
key=lambda k: adata.var["dispersions_norm"][k],
)[-51:][0]
]
adata.var["top_highly_variable"] = (
adata.var["dispersions_norm"] > top_dispersion
)
# ## 4. Create the Vitessce widget configuration
#
# Vitessce needs to know which pieces of data we are interested in visualizing, the visualization types we would like to use, and how we want to coordinate (or link) the views.
# ### 4.1. Instantiate a `VitessceConfig` object
#
# Use the `VitessceConfig(name, description)` constructor to create an instance.
vc = VitessceConfig(name='Habib et al', description='COVID-19 Healthy Donor Brain')
# ### 4.2. Add a dataset to the `VitessceConfig` instance
#
# In Vitessce, a dataset is a container for one file per data type. The `.add_dataset(name)` method on the `vc` instance sets up and returns a new dataset instance.
#
# Then, we can call the dataset's `.add_object(wrapper_object)` method to attach a "data wrapper" instance to our new dataset. For example, the `AnnDataWrapper` class knows how to convert AnnData objects to the corresponding Vitessce data types.
#
# Dataset wrapper classes may require additional parameters to resolve ambiguities. For instance, `AnnData` objects may store multiple clusterings or cell type annotation columns in the `adata.obs` dataframe. We can use the parameter `cell_set_obs_cols` to tell Vitessce which columns of the `obs` dataframe correspond to cell sets.
dataset = vc.add_dataset(name='Brain').add_object(AnnDataWrapper(
adata,
mappings_obsm=["X_umap"],
mappings_obsm_names=["UMAP"],
cell_set_obs=["CellType"],
cell_set_obs_names=["Cell Type"],
expression_matrix="X",
matrix_gene_var_filter="top_highly_variable"
)
)
# ### 4.3. Add visualizations to the `VitessceConfig` instance
#
# Now that we have added a dataset, we can configure visualizations. The `.add_view(dataset, component_type)` method adds a view (i.e. visualization or controller component) to the configuration.
#
# The `Component` enum class (which we have imported as `cm` here) can be used to fill in the `component_type` parameter.
#
# For convenience, the `SCATTERPLOT` component type takes the extra `mapping` keyword argument, which specifies which embedding should be used for mapping cells to (x,y) points on the plot.
scatterplot = vc.add_view(dataset, cm.SCATTERPLOT, mapping="UMAP")
cell_sets = vc.add_view(dataset, cm.CELL_SETS)
genes = vc.add_view(dataset, cm.GENES)
heatmap = vc.add_view(dataset, cm.HEATMAP)
# ### 4.4. Define the visualization layout
#
# The `vc.layout(view_concat)` method allows us to specify how our views will be arranged in the layout grid in the widget. The `|` and `/` characters are magic syntax for `hconcat(v1, v2)` and `vconcat(v1, v2)`, respectively.
vc.layout((scatterplot | cell_sets) / (heatmap | genes));
# ## 5. Create the widget
#
# The `vc.widget()` method returns the configured widget instance.
vw = vc.widget()
vw
| docs/notebooks/widget_brain.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import binascii
import ipaddress
# +
ADDRESSES = [
'10.9.0.6',
'fdfd:87b5:b475:5e3e:b1bc:e121:a8eb:14aa',
]
for ip in ADDRESSES:
addr = ipaddress.ip_address(ip)
print('{!r}'.format(addr))
print(' IP version:', addr.version)
print(' is private:', addr.is_private)
print(' packed form:', binascii.hexlify(addr.packed))
print(' integer:', int(addr))
print()
# +
NETWORKS = [
'10.9.0.0/24',
'fdfd:87b5:b475:5e3e::/64',
]
for n in NETWORKS:
net = ipaddress.ip_network(n)
print('{!r}'.format(net))
print(' is private:', net.is_private)
print(' broadcast:', net.broadcast_address)
print(' compressed:', net.compressed)
print(' with netmask:', net.with_netmask)
print(' with hostmask:', net.with_hostmask)
print(' num addresses:', net.num_addresses)
print()
# +
import ipaddress
NETWORKS = [
'10.9.0.0/24',
'fdfd:87b5:b475:5e3e::/64',
]
for n in NETWORKS:
net = ipaddress.ip_network(n)
print('{!r}'.format(net))
for i, ip in zip(range(3), net.hosts()):
print(ip)
print()
# +
NETWORKS = [
ipaddress.ip_network('10.9.0.0/24'),
ipaddress.ip_network('fdfd:87b5:b475:5e3e::/64'),
]
ADDRESSES = [
ipaddress.ip_address('10.9.0.6'),
ipaddress.ip_address('10.7.0.31'),
ipaddress.ip_address(
'fdfd:87b5:b475:5e3e:b1bc:e121:a8eb:14aa'
),
ipaddress.ip_address('fe80::3840:c439:b25e:63b0'),
]
for ip in ADDRESSES:
for net in NETWORKS:
if ip in net:
print('{}\nis on {}'.format(ip, net))
break
else:
print('{}\nis not on a known network'.format(ip))
print()
# +
import ipaddress
ADDRESSES = [
'10.9.0.6/24',
'fdfd:87b5:b475:5e3e:b1bc:e121:a8eb:14aa/64',
]
for ip in ADDRESSES:
iface = ipaddress.ip_interface(ip)
print('{!r}'.format(iface))
print('network:\n ', iface.network)
print('ip:\n ', iface.ip)
print('IP with prefixlen:\n ', iface.with_prefixlen)
print('netmask:\n ', iface.with_netmask)
print('hostmask:\n ', iface.with_hostmask)
print()
# -
| Python-Standard-Library/Network/ipaddress.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # JSON Path Expressions
#
# JSON documents have an inherent structure to them, similar to XML documents or a file system. Many of the JSON functions provided with Db2 need a method to navigate through a document to retrieve the object or item that the user wants. To illustrate how a JSON path expression points to a particular object, one of the customer documents will be used:
# ```json
# {
# "customerid": 100000,
# "identity": {
# "firstname": "Jacob",
# "lastname": "Hines",
# "birthdate": "1982-09-18"
# },
# "contact": {
# "street": "Main Street North",
# "city": "Amherst",
# "state": "OH",
# "zipcode": "44001",
# "email": "<EMAIL>",
# "phone": "813-689-8309"
# },
# "payment": {
# "card_type": "MCCD",
# "card_no": "4742-3005-2829-9227"
# },
# "purchases": [
# {
# "tx_date": "2018-02-14",
# "tx_no": 157972,
# "product_id": 1860,
# "product": "Ugliest Snow Blower",
# "quantity": 1,
# "item_cost": 51.86
# }, …additional purchases…
# ]
# }
# ```
# ### Document Structure
# Every JSON path expression begins with a dollar sign (`$`) to represent the root or top of the document structure. To traverse down the document, the dot/period (`.`) is used to move down one level. The "`$`" and "`.`" characters are reserved characters for the purposes of path expressions so care needs to be taken not to use them as a part of a key name in a key-value pair.
#
# In our customer document example, to refer to the value associated with the customerid key, the path expression would be:
# ```json
# $.customerid
# ```
# To retrieve the value associated with the identity key, the path expression would be:
# ```json
# $.identity
# ```
# The value referred to in this last example is the entire JSON object that is the value associated with identity so the following object would be returned:
# ```json
# {
# "firstname": "Jacob",
# "lastname" : "Hines",
# "birthdate": "1982-09-18"
# }
# ```
# If we needed to traverse the interior of the JSON OBJECT value associated with identity, for example to refer to the birthdate, then we would append the initial key name with a period and the internal key name for the value of interest:
# ```json
# $.identity.birthdate
# Result: "1982-09-18"
# ```
# Up to this point, we have only been referring to objects and individual key-value pairs within a JSON document. JSON allows for the use of simple arrays (such as phone numbers) or for arrays of objects (like purchases).
# A simple and complex array type are shown below:
# ```json
# {
# "employee": 10,
# "phoneno": ["592-533-9042","354-981-0032","919-778-1539"]
# }
#
# {
# "division": 5,
# "stores" :
# [
# {"id": 45, "city" : "Toronto"},
# {"id": 13, "city" : "Markham"},
# {"id": 93, "city" : "Schaumburg"}
# ]
# }
# ```
# To refer to an entire array you would just reference the object name.
# ```json
# $.phoneno
#
# Result: ["592-533-9042","354-981-0032","919-778-1539"]
# ```
# To reference the first element of an array, you would append an array specifier (square brackets `[]`) with the number representing the position of the element in the array inside the brackets. This number is also referred to as the array index value, or simply the index value. The first element of a JSON array always begins at index value zero.
# To refer to the first phone number from the above list, we would use this path:
# ```json
# $.phoneno[0]
#
# Result: "592-533-9042"
# ```
# An index value must be provided, otherwise the Db2 JSON functions will return a null (or an error depending on other settings which we have not yet discussed). The reverse situation, when you specify an index value of zero for a non-array field will cause an error in strict mode and be accepted under lax mode with the contents of the field will be returned. For instance, the following path expression is acceptable to Db2 JSON functions when using lax mode:
# ```json
# $.stores[0].city[0]
#
# Result: "Toronto"
# ```
# A good development practice would be to define how fields should be created in a document. If a field could potentially have multiple values, then single items should be inserted as arrays with a single value rather than as an atomic item. Examining the document will make it clear that an object could have more than one item in it such as in this example:
# ```json
# "phoneno": ["592-533-9042"]
# ```
# Dealing with arrays of objects is similar to simple objects – an index value is used before traversing down the document. To retrieve the city of the second store of the division would require the following path statement:
# ```json
# $stores[1].city
#
# Result: "Markham"
# ```
# Since stores is an array of objects, we must first select which object in the array needs to be retrieved. The `[1]` represents the second object in the array:
# ```json
# {"id": 13, "city" : "Markham"}
# ```
# We used the dot notation to traverse the contents of the object to refer to the city. Arrays can be nested to many levels and can make path expressions complex.
#
# The next document has two levels of arrays.
# ```json
# {
# "division": 5,
# "stores" :
# [
# {"id": 45, "phone": ["592-533-9042","354-981-0032"]},
# {"id": 13, "phone": ["634-231-9862"]},
# {"id": 93, "phone": ["883-687-1123","442-908-9435","331-991-2433"]}
# ]
# }
# ```
# To create a path to the second phone number of store 93, we would need to use two array specifications:
# ```json
# $.stores[2].phone[1]
# ```
# As the depth of the path expression increases, the potential for errors also becomes higher. One way of reducing the complexity of path expressions is to use two or more steps to traverse a document. For the example above, a user could use the following approach:
# * stores = using(document) find the contents of `$.stores[2]`
# * result = using(stores) find the contents of `$.phone[1]`
# ### Path Expression Summary
# The following table summarizes the examples of path expressions that have been covered so far (using the division/stores example above).
#
# |Pattern | Result
# |:-------|:------
# | `$.division` | `5`
# | `$.stores`| `[{"id": 45, "phone": ["592-533-9042","354-981-0032"]},`
# || `{"id": 13, "phone": ["634-231-9862"]},`
# || `{"id": 93, "phone": ["883-687-1123","442-908-9435","331-991-2433"]}]`
# | `$.stores[0]` | `{"id": 45, "phone": ["592-533-9042","354-981-0032"]}`
# | `$.stores[1].id` | `13`
# | `$.stores[2].phone` | `["883-687-1123","442-908-9435","331-991-2433"]`
# | `$.stores[2].phone[1]` | `"442-908-9435"`
# ### Simplifying JSON Path Expressions
# The previous section illustrated two shortcomings of JSON path expressions:
# * JSON path expressions can get complex, especially when dealing with arrays and objects within objects
# * Path expressions are limited to referencing only an individual object, array, or item
#
# When writing path expressions, the potential for spelling mistakes goes up as the path gets longer! If the field name is unique in a document, it can be referred to much more easily by using the asterisk (`*`) or wildcard character.
#
# The wildcard character (asterisk `*`) can be used to match any object in a level or an array. The asterisk does not match all levels in the document, just the immediate one.
#
# For instance, consider the following document:
# ```json
# {
# "employee": 10,
# "details" :
# {
# "name":
# {
# "first":"George",
# "last" :"Baklarz"
# },
# "phoneno": ["592-533-9042","354-981-0032","919-778-1539"]
# }
# }
# ```
# To refer to the last name of the individual in the document, we could write the following path expression:
# ```json
# $.details.name.last
#
# Result: Baklarz
# ```
# The asterisk can be used to match anything at the current level. The equivalent path expression is:
# ```json
# $.*.*.last
#
# Result: Baklarz
# ```
# This technique is useful when the key is unique but can cause problems when the key is duplicated throughout the document. The next section discusses the use of the wildcard character to retrieve multiple values and the pitfalls associated with it.
# ### Referring to Multiple Objects with JSON Path Expressions
# In some situations, a developer may want to retrieve all objects within a document that have the same key name. JSON path expressions include the option of using the asterisk character (`*`) to match any name at the current level. If there are multiple objects that match, then all the matches will be returned.
# The following document has multiple objects with the same name. From a development perspective, it doesn't make any sense to use different keywords for first and last names in a document, so this is a reasonable naming convention.
# ```json
# {
# "authors":
# {
# "primary" : {"first_name": "Paul", "last_name" : "Bird"},
# "secondary" : {"first_name": "George","last_name" : "Baklarz"}
# },
# "foreword":
# {
# "primary" : {"first_name": "Thomas","last_name" : "Hronis"},
# "formats": ["Hardcover","Paperback","eBook","PDF"]
# }
# }
# ```
# The following table summarizes to what an asterisk in the path expression is referring to.
#
# |Pattern |Path |Result
# |:----|:-----|:------
# |`$.authors.primary.last_name` |`$.authors.primary.last_name` |`Bird`
# |`$.*.primary.last_name`|`$.authors.primary.last_name`|`<NAME>`
# | |`$.foreword.primary.last_name`
# |`$.*.*.last_name`|`$.authors.primary.last_name` | `<NAME>`
# | | `$.authors.secondary.last_name`
# | | `$.foreword.primary.last_name`
# |`$.authors.*.last_name` |`$.authors.primary.last_name`|`<NAME>`
# | |`$.authors.secondary.last_name`
# As you can tell from the examples that there are drawbacks when using asterisk in a pattern:
# * The relationship of the item (last_name) within the document is unknown (i.e. what was the field part of?)
# * One or more last_name fields names could be returned, which means that the JSON function using this path needs to be able to handle more than one value (i.e. you can’t use a JSON scalar function, which expects to deal with one value, to handle multiple values. You will get an error!)
#
# A developer needs to be aware of these limitations when using wildcard expressions.
# The wildcard character can also be used in two other ways:
# * To refer to all elements in an array
# * To refer to all values in an object
#
# Using the wildcard character in an array specification allows the JSON path expression to retrieve the individual values that are in each array element. This is primarily used for arrays that contain objects rather than atomic values.
#
# The author example has been modified to make the author list into an array.
# ```json
# {
# "authors":
# [
# {"first_name": "Paul", "last_name" : "Bird"},
# {"first_name": "George","last_name" : "Baklarz"}
# ],
# "foreword":
# {
# "primary": {"first_name": "Thomas","last_name" : "Hronis"}
# },
# "formats": ["Hardcover","Paperback","eBook","PDF"]
# }
# ```
# Referring to all of the book formats in the document can be achieved using one of these two techniques:
# ```json
# $.formats
# $.formats.*
# ```
# In the first case, a single JSON array is returned which consists of an array of strings:
# ```json
# ["Hardcover","Paperback","eBook","PDF"]
# ```
# The second statement returns 4 individual values:
# ```json
# "Hardcover","Paperback","eBook","PDF"
# ```
# The wildcard character could also be placed at the end of a JSON path expression to retrieve all values in an object. The following path expression refers to all of the values in the foreword author.
# Note that the keys (first_name, last_name) are not retrieved.
# ```json
# $.foreword.primary.*
#
# Result: "Thomas","Hronis"
# ```
# If you wanted to retrieve all last_names in the authors array, you would use the following path expression:
# ```json
# $.authors[*].last_name
#
# Result: ["Bird","Baklarz"]
# ```
# The use of the wildcard character can be very powerful when dealing with JSON path expressions. The user must take care to ensure that the results being returned are from the appropriate level within the document since the path expression does not recurse into the document.
#
# The following table summarizes what an asterisk in an array specification and at the end of a JSON path expression would produce.
#
# | Pattern | Result
# |:--------|:----------
# |`$.authors` | `[{"first_name":"Paul", "last_name":"Bird"},`
# | | `{"first_name":"George","last_name":"Baklarz"}]`
# |`$.authors.*` | `"Paul","Bird","George","Baklarz"`
# |`$.authors[*].last_name` | `"Bird", "Baklarz"`
# |`$.foreword.primary.*` | `"Thomas", "Hronis"`
# | `$.formats` | `["Hardcover","Paperback","eBook","PDF"]`
# |`$.formats[*]` | `"Hardcover","Paperback","eBook","PDF"`
#
# In summary, a JSON path expression can be used to navigate to individual elements, objects, arrays, or allow for multiple matches within a document.
# ### Summary
# The following list summarizes how a JSON path expression is built.
# * The top of any path expression is the anchor symbol (`$`)
# * Traverse to specific objects by using the dot operator (`.`)
# * Use square brackets `[]` to refer to items in an array with the first item starting at zero
# * Use the backslash `\` as an escape character when key names include any of the JSON path characters `(.,*,$,[,])`
# * Use the asterisk (`*`) to match any object at the current level
# * Use the asterisk (`*`) to match all objects in an array or retrieve only the value fields from an object
# #### Copyright (C) IBM 2021, <NAME> [<EMAIL>]
| Db2_11.5_JSON_02_JSON_Path_Expressions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/davidguzmanr/Aprendizaje-Profundo/blob/main/Tareas/Tarea-1/Tarea_1_ejercicio_3.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="uq2Fk6KVnjX5"
# # Tarea 1: perceptrón y redes densas
#
# - **E. <NAME>**
# - **Introducción al Aprendizaje Profundo 2021-II**
# - **Licenciatura en Ciencia de Datos CU UNAM**
# + id="dn0AcOionkaF"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib.colors import ListedColormap
import seaborn as sns
import torch
import torch.nn as nn
import torch.optim as optim
from tqdm import trange, tqdm
sns.set_style('darkgrid')
# + [markdown] id="fwOzTqNYnxuM"
# ## Ejercicio 3
#
# Entrena una red completamente conectada para aproximar la compuerta XOR (puedes usar todas las herramientas de PyTorch). (3 puntos)
#
# + [markdown] id="RAZ3C9sjn0YA"
# **Solución.** Como vimos la compuerta XOR ($\oplus$) es no lineal, de modo que no la podemos aproximar con un solo perceptrón, pero podemos agregar neuronas y/o apilar varias capas.
#
# | $x_1$ | $x_2$ | $y$ |
# |-------|-------|-----|
# | 0 | 0 | 0 |
# | 0 | 1 | 1 |
# | 1 | 0 | 1 |
# | 1 | 1 | 0 |
# + id="TDcWlrCTn27j"
class XOR(nn.Module):
def __init__(self):
super(XOR, self).__init__()
self.fc = nn.Sequential(
nn.Linear(2, 4),
nn.ReLU(),
nn.Linear(4, 1),
nn.ReLU(),
)
def forward(self, x):
return self.fc(x)
# + id="Bx7Cww0OoAP4"
def train(model, X, y, lr=0.01, epochs=1000):
# Función de pérdida, error cuadrático medio (mean squared error),
# posteriormente usaré un umbral para hacer la clasificación en 0 o 1.
criterion = nn.MSELoss()
# Optimizador
opt = optim.SGD(model.parameters(), lr=lr)
losses = []
for epoch in trange(epochs):
loss_epoch = []
for x, y_true in zip(X, y):
y_pred = model(x) # predecimos
loss = criterion(y_pred, y_true) # calculamos el error
loss_epoch.append(loss.item())
opt.zero_grad() # vaciamos los gradientes
loss.backward() # retropropagamos
opt.step() # actualizamos parámetros
losses.append(np.mean(loss_epoch))
return losses
# + colab={"base_uri": "https://localhost:8080/"} id="QGWa2qUwnt9D" outputId="9730209b-7ece-4307-e7e0-e9abbdf0c5a2"
torch.manual_seed(0)
# Definimos nuestro modelo y probamos con unos datos sintéticos
model = XOR()
x = torch.zeros(1,2)
model(x)
# + colab={"base_uri": "https://localhost:8080/"} id="JXkTiy9nn-AD" outputId="db6f4e9e-3279-4516-80f7-27996b50ed19"
X_xor = torch.tensor([[0,0], [0,1], [1,0], [1,1]], dtype=torch.float32)
y_xor = torch.tensor([[0], [1], [1], [0]], dtype=torch.float32)
# Entrenamos
losses = train(model, X_xor, y_xor, epochs=1000, lr=0.01)
# + colab={"base_uri": "https://localhost:8080/", "height": 390} id="ZH8d4rdFoKL7" outputId="76440786-89c4-45b0-b0f7-8b1f6efbaa19"
plt.figure(figsize=(10,6))
plt.plot(losses)
plt.xlabel('Época', weight='bold')
plt.ylabel('Error (cuadrático medio)', weight='bold')
plt.show()
# + [markdown] id="oxTgi2BpxExz"
# Finalmente, aplicamos una función escalón que nos da el resultado que queremos.
# + id="1Qcv6RDsoR9U"
def step(z):
"""Computes step function."""
return 1.0 if z > 0.5 else 0.0
step_vectorize = np.vectorize(step)
# + colab={"base_uri": "https://localhost:8080/"} id="pcErDIJjovto" outputId="d23bb407-2781-4783-984a-46760daef8ec"
# predicción por cada ejemplo
print('---------------------')
print('x_1 \tx_2 \ty_hat')
print('---------------------')
for x in X_xor:
x1, x2 = x
y_hat = step(model(x))
print(f'{x1}\t{x2}\t{y_hat}')
# + [markdown] id="cfGt5jCpo3SP"
# Ahora veamos cómo está clasificando, vemos que en efecto es no lineal, pues no hay manera de obtener estas regiones usando únicamente una línea.
#
# <!-- ([Tensorflow playground](https://playground.tensorflow.org/)). -->
# + colab={"base_uri": "https://localhost:8080/", "height": 388} id="9G96JxqwoweD" outputId="c20733cb-3960-4671-f9fc-fdf1562f25a0"
X1, X2 = np.meshgrid(np.arange(start=-0.2, stop=1.2, step=0.01), np.arange(start=-0.2, stop=1.2, step=0.01))
Z = model(torch.tensor(np.array([X1.ravel(), X2.ravel()]).T, dtype=torch.float32)).reshape(X1.shape)
Z = step_vectorize(Z.detach().numpy())
plt.figure(figsize=(10,6))
plt.contourf(X1, X2, Z, alpha=0.2, cmap=ListedColormap(('orange', 'blue')))
plt.scatter([0,1], [0,1], color='orange', label='0')
plt.scatter([0,1], [1,0], color='blue', label='1')
plt.xlabel('X1', weight='bold')
plt.ylabel('X2', weight='bold')
plt.legend()
plt.show()
# + id="hcVgc0bKo1wr"
| Tareas/Tarea-1/Tarea_1_ejercicio_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import scipy
import matplotlib.pyplot as plt
# %matplotlib inline
import os, sys
import networkx as nx
# -
libpath = os.path.abspath('..')
if libpath not in sys.path:
sys.path.append(libpath)
from qubitrbm.qaoa import QAOA
from qubitrbm.optimize import Optimizer
from qubitrbm.rbm import RBM
from qubitrbm.utils import exact_fidelity
# ## The setup
# Define a graph to run QAOA on:
G = nx.random_regular_graph(d=3, n=12, seed=12345)
nx.draw_kamada_kawai(G, node_color='gold', node_size=500)
# For $p>1$, provided we have a small graph, we can find optimal angles exactly:
qaoa = QAOA(G, p=2)
# %%time
angles, costs = qaoa.optimize(init=[np.pi/8, np.pi/8, -np.pi/8, -np.pi/8], tol=1e-4)
fig, ax = plt.subplots(figsize=[8,5])
ax.plot(costs)
ax.set_xlabel('Iteration step', fontsize=20)
ax.set_ylabel(r'$\langle \mathcal{C} \rangle $', fontsize=30)
gammas, betas = np.split(angles, 2)
gammas[0] # \gamma _1
gammas[1] # \gamma _2
betas[0] # \beta _1
betas[1] # \beta _2
# Initialize an RBM ansatz with $N=12$ visible units, the same number as the underlying graph
logpsi = RBM(12)
# Exactly apply $U_C (\gamma _1) = \exp \left( -i \gamma _1 \sum _{\langle i, j \rangle } Z_i Z_j \right)$
logpsi.UC(G, gamma=gammas[0], mask=False)
# The process introduced a number of hidden units $n_h$ that's equal to the number of edges in the graph. (Plus 1 that was there by default when we initialized the RBM.)
#
# We can look at the numbers:
logpsi.nv, logpsi.nh
logpsi.alpha # = logpsi.nh / logpsi.nv
# ## The first optimization
# Now, initialize the optimizer and approximately apply $U_B (\beta _1) = \exp \left( -i \beta _1 \sum _i X_i \right)$
optim = Optimizer(logpsi, n_steps=800, n_chains=4, warmup=800, step=12)
# +
# %%time
for n in range(len(G)):
params, history = optim.sr_rx(n=n, beta=betas[0], resample_phi=3, verbose=True)
optim.machine.params = params
print(f'Done with qubit #{n+1}, reached fidelity {history[-1]}')
# -
logpsi.params = params
# It's a good check to compare exact fidelities at this point:
psi_exact = QAOA(G, p=1).simulate(gammas[0], betas[0]).final_state_vector
psi_rbm = logpsi.get_state_vector(normalized=True)
exact_fidelity(psi_exact, psi_rbm)
# Next, apply
#
# $$U_C (\gamma _2) = \exp \left( -i \gamma _2 \sum _{\langle i, j \rangle } Z_i Z_j \right)$$
logpsi.UC(G, gamma=gammas[1])
optim.machine = logpsi
# However, this doubled the number of hidden units:
logpsi.alpha
# ## The compression step
# We can keep the number of hidden units under control as we go to higher values of $p$ by performing a compression step, as described in the paper.
#
# Essentially, we define a smaller RBM with `RBM.alpha = 1.5` (the previous value or any we choose to compress to). Then, we optimize parameters of the new RBM to describe the same quantum state as the larger one, obtaining a compressed representaion of
#
# $$ \vert \psi \rangle = U_C (\gamma _2) \; U_B (\beta _1) \; U_C(\gamma _1) \; \vert + \rangle $$
# A heuristically good choice for initial RBM parameters are those values that exactly describe the following quantum state:
#
# $$ \vert \psi _\text{init} \rangle = U_C \left( \frac{\gamma_1 + \gamma _2}{2} \right) \; \vert + \rangle $$
aux = RBM(len(G))
aux.UC(G, (gammas[0] + gammas[1])/2)
init_params = aux.params
# Now, perform the compression:
# %%time
params, history = optim.sr_compress(init=init_params, resample_phi=2, verbose=True)
# Let's plot the fidelity as a function of compression optimizer step:
fig, ax = plt.subplots(figsize=[8,5])
ax.plot(history)
ax.set_xlabel('Iteration step', fontsize=30)
ax.set_ylabel('Fidelity', fontsize=30)
# Estimated fidelity reached:
history[-1]
logpsi = RBM(12, (len(params) - 12)//(12+1))
logpsi.params = params
logpsi.alpha
# Finally, we can apply $U_B (\beta _2) = \exp \left( -i \beta _2 \sum _i X_i \right)$
optim.machine = logpsi
# ## The second optimization
# +
# %%time
for n in range(len(G)):
params, history = optim.sr_rx(n=n, beta=betas[1], resample_phi=3, verbose=True)
optim.machine.params = params
print(f'Done with qubit #{n+1}, reached fidelity {history[-1]}')
# -
# And, compare the final output fidelity at $p=2$:
logpsi.params = params
psi_exact = QAOA(G, p=2).simulate(gammas, betas).final_state_vector
psi_rbm = logpsi.get_state_vector(normalized=True)
exact_fidelity(psi_exact, psi_rbm)
| examples/rbm_optimization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="j2KIji9Ts0Wn" colab_type="code" colab={}
'''
code by <NAME>(<NAME>) @graykode, modify by wmathor
Reference : https://github.com/jadore801120/attention-is-all-you-need-pytorch
https://github.com/JayParks/transformer, https://github.com/dhlee347/pytorchic-bert
'''
import re
import math
import torch
import numpy as np
from random import *
import torch.nn as nn
import torch.optim as optim
import torch.utils.data as Data
text = (
'Hello, how are you? I am Romeo.\n' # R
'Hello, Romeo My name is Juliet. Nice to meet you.\n' # J
'Nice meet you too. How are you today?\n' # R
'Great. My baseball team won the competition.\n' # J
'Oh Congratulations, Juliet\n' # R
'Thank you Romeo\n' # J
'Where are you going today?\n' # R
'I am going shopping. What about you?\n' # J
'I am going to visit my grandmother. she is not very well' # R
)
sentences = re.sub("[.,!?\\-]", '', text.lower()).split('\n') # filter '.', ',', '?', '!'
word_list = list(set(" ".join(sentences).split())) # ['hello', 'how', 'are', 'you',...]
word2idx = {'[PAD]' : 0, '[CLS]' : 1, '[SEP]' : 2, '[MASK]' : 3}
for i, w in enumerate(word_list):
word2idx[w] = i + 4
idx2word = {i: w for i, w in enumerate(word2idx)}
vocab_size = len(word2idx)
token_list = list()
for sentence in sentences:
arr = [word2idx[s] for s in sentence.split()]
token_list.append(arr)
# + id="-AuXO3rQJIUj" colab_type="code" colab={}
# BERT Parameters
maxlen = 30
batch_size = 6
max_pred = 5 # max tokens of prediction
n_layers = 6
n_heads = 12
d_model = 768
d_ff = 768*4 # 4*d_model, FeedForward dimension
d_k = d_v = 64 # dimension of K(=Q), V
n_segments = 2
# + id="bOXYOwFAJH93" colab_type="code" colab={}
# sample IsNext and NotNext to be same in small batch size
def make_data():
batch = []
positive = negative = 0
while positive != batch_size/2 or negative != batch_size/2:
tokens_a_index, tokens_b_index = randrange(len(sentences)), randrange(len(sentences)) # sample random index in sentences
tokens_a, tokens_b = token_list[tokens_a_index], token_list[tokens_b_index]
input_ids = [word2idx['[CLS]']] + tokens_a + [word2idx['[SEP]']] + tokens_b + [word2idx['[SEP]']]
segment_ids = [0] * (1 + len(tokens_a) + 1) + [1] * (len(tokens_b) + 1)
# MASK LM
n_pred = min(max_pred, max(1, int(len(input_ids) * 0.15))) # 15 % of tokens in one sentence
cand_maked_pos = [i for i, token in enumerate(input_ids)
if token != word2idx['[CLS]'] and token != word2idx['[SEP]']] # candidate masked position
shuffle(cand_maked_pos)
masked_tokens, masked_pos = [], []
for pos in cand_maked_pos[:n_pred]:
masked_pos.append(pos)
masked_tokens.append(input_ids[pos])
if random() < 0.8: # 80%
input_ids[pos] = word2idx['[MASK]'] # make mask
elif random() > 0.9: # 10%
index = randint(0, vocab_size - 1) # random index in vocabulary
while index < 4: # can't involve 'CLS', 'SEP', 'PAD'
index = randint(0, vocab_size - 1)
input_ids[pos] = index # replace
# Zero Paddings
n_pad = maxlen - len(input_ids)
input_ids.extend([0] * n_pad)
segment_ids.extend([0] * n_pad)
# Zero Padding (100% - 15%) tokens
if max_pred > n_pred:
n_pad = max_pred - n_pred
masked_tokens.extend([0] * n_pad)
masked_pos.extend([0] * n_pad)
if tokens_a_index + 1 == tokens_b_index and positive < batch_size/2:
batch.append([input_ids, segment_ids, masked_tokens, masked_pos, True]) # IsNext
positive += 1
elif tokens_a_index + 1 != tokens_b_index and negative < batch_size/2:
batch.append([input_ids, segment_ids, masked_tokens, masked_pos, False]) # NotNext
negative += 1
return batch
# Proprecessing Finished
# + id="bqxycRhzia7r" colab_type="code" colab={}
batch = make_data()
input_ids, segment_ids, masked_tokens, masked_pos, isNext = zip(*batch)
input_ids, segment_ids, masked_tokens, masked_pos, isNext = \
torch.LongTensor(input_ids), torch.LongTensor(segment_ids), torch.LongTensor(masked_tokens),\
torch.LongTensor(masked_pos), torch.LongTensor(isNext)
class MyDataSet(Data.Dataset):
def __init__(self, input_ids, segment_ids, masked_tokens, masked_pos, isNext):
self.input_ids = input_ids
self.segment_ids = segment_ids
self.masked_tokens = masked_tokens
self.masked_pos = masked_pos
self.isNext = isNext
def __len__(self):
return len(self.input_ids)
def __getitem__(self, idx):
return self.input_ids[idx], self.segment_ids[idx], self.masked_tokens[idx], self.masked_pos[idx], self.isNext[idx]
loader = Data.DataLoader(MyDataSet(input_ids, segment_ids, masked_tokens, masked_pos, isNext), batch_size, True)
# + id="6inMS744xRwh" colab_type="code" colab={}
def get_attn_pad_mask(seq_q, seq_k):
batch_size, seq_len = seq_q.size()
# eq(zero) is PAD token
pad_attn_mask = seq_q.data.eq(0).unsqueeze(1) # [batch_size, 1, seq_len]
return pad_attn_mask.expand(batch_size, seq_len, seq_len) # [batch_size, seq_len, seq_len]
def gelu(x):
"""
Implementation of the gelu activation function.
For information: OpenAI GPT's gelu is slightly different (and gives slightly different results):
0.5 * x * (1 + torch.tanh(math.sqrt(2 / math.pi) * (x + 0.044715 * torch.pow(x, 3))))
Also see https://arxiv.org/abs/1606.08415
"""
return x * 0.5 * (1.0 + torch.erf(x / math.sqrt(2.0)))
class Embedding(nn.Module):
def __init__(self):
super(Embedding, self).__init__()
self.tok_embed = nn.Embedding(vocab_size, d_model) # token embedding
self.pos_embed = nn.Embedding(maxlen, d_model) # position embedding
self.seg_embed = nn.Embedding(n_segments, d_model) # segment(token type) embedding
self.norm = nn.LayerNorm(d_model)
def forward(self, x, seg):
seq_len = x.size(1)
pos = torch.arange(seq_len, dtype=torch.long)
pos = pos.unsqueeze(0).expand_as(x) # [seq_len] -> [batch_size, seq_len]
embedding = self.tok_embed(x) + self.pos_embed(pos) + self.seg_embed(seg)
return self.norm(embedding)
class ScaledDotProductAttention(nn.Module):
def __init__(self):
super(ScaledDotProductAttention, self).__init__()
def forward(self, Q, K, V, attn_mask):
scores = torch.matmul(Q, K.transpose(-1, -2)) / np.sqrt(d_k) # scores : [batch_size, n_heads, seq_len, seq_len]
scores.masked_fill_(attn_mask, -1e9) # Fills elements of self tensor with value where mask is one.
attn = nn.Softmax(dim=-1)(scores)
context = torch.matmul(attn, V)
return context
class MultiHeadAttention(nn.Module):
def __init__(self):
super(MultiHeadAttention, self).__init__()
self.W_Q = nn.Linear(d_model, d_k * n_heads)
self.W_K = nn.Linear(d_model, d_k * n_heads)
self.W_V = nn.Linear(d_model, d_v * n_heads)
def forward(self, Q, K, V, attn_mask):
# q: [batch_size, seq_len, d_model], k: [batch_size, seq_len, d_model], v: [batch_size, seq_len, d_model]
residual, batch_size = Q, Q.size(0)
# (B, S, D) -proj-> (B, S, D) -split-> (B, S, H, W) -trans-> (B, H, S, W)
q_s = self.W_Q(Q).view(batch_size, -1, n_heads, d_k).transpose(1,2) # q_s: [batch_size, n_heads, seq_len, d_k]
k_s = self.W_K(K).view(batch_size, -1, n_heads, d_k).transpose(1,2) # k_s: [batch_size, n_heads, seq_len, d_k]
v_s = self.W_V(V).view(batch_size, -1, n_heads, d_v).transpose(1,2) # v_s: [batch_size, n_heads, seq_len, d_v]
attn_mask = attn_mask.unsqueeze(1).repeat(1, n_heads, 1, 1) # attn_mask : [batch_size, n_heads, seq_len, seq_len]
# context: [batch_size, n_heads, seq_len, d_v], attn: [batch_size, n_heads, seq_len, seq_len]
context = ScaledDotProductAttention()(q_s, k_s, v_s, attn_mask)
context = context.transpose(1, 2).contiguous().view(batch_size, -1, n_heads * d_v) # context: [batch_size, seq_len, n_heads, d_v]
output = nn.Linear(n_heads * d_v, d_model)(context)
return nn.LayerNorm(d_model)(output + residual) # output: [batch_size, seq_len, d_model]
class PoswiseFeedForwardNet(nn.Module):
def __init__(self):
super(PoswiseFeedForwardNet, self).__init__()
self.fc1 = nn.Linear(d_model, d_ff)
self.fc2 = nn.Linear(d_ff, d_model)
def forward(self, x):
# (batch_size, seq_len, d_model) -> (batch_size, seq_len, d_ff) -> (batch_size, seq_len, d_model)
return self.fc2(gelu(self.fc1(x)))
class EncoderLayer(nn.Module):
def __init__(self):
super(EncoderLayer, self).__init__()
self.enc_self_attn = MultiHeadAttention()
self.pos_ffn = PoswiseFeedForwardNet()
def forward(self, enc_inputs, enc_self_attn_mask):
enc_outputs = self.enc_self_attn(enc_inputs, enc_inputs, enc_inputs, enc_self_attn_mask) # enc_inputs to same Q,K,V
enc_outputs = self.pos_ffn(enc_outputs) # enc_outputs: [batch_size, seq_len, d_model]
return enc_outputs
class BERT(nn.Module):
def __init__(self):
super(BERT, self).__init__()
self.embedding = Embedding()
self.layers = nn.ModuleList([EncoderLayer() for _ in range(n_layers)])
self.fc = nn.Sequential(
nn.Linear(d_model, d_model),
nn.Dropout(0.5),
nn.Tanh(),
)
self.classifier = nn.Linear(d_model, 2)
self.linear = nn.Linear(d_model, d_model)
self.activ2 = gelu
# fc2 is shared with embedding layer
embed_weight = self.embedding.tok_embed.weight
self.fc2 = nn.Linear(d_model, vocab_size, bias=False)
self.fc2.weight = embed_weight
def forward(self, input_ids, segment_ids, masked_pos):
output = self.embedding(input_ids, segment_ids) # [bach_size, seq_len, d_model]
enc_self_attn_mask = get_attn_pad_mask(input_ids, input_ids) # [batch_size, maxlen, maxlen]
for layer in self.layers:
# output: [batch_size, max_len, d_model]
output = layer(output, enc_self_attn_mask)
# it will be decided by first token(CLS)
h_pooled = self.fc(output[:, 0]) # [batch_size, d_model]
logits_clsf = self.classifier(h_pooled) # [batch_size, 2] predict isNext
masked_pos = masked_pos[:, :, None].expand(-1, -1, d_model) # [batch_size, max_pred, d_model]
h_masked = torch.gather(output, 1, masked_pos) # masking position [batch_size, max_pred, d_model]
h_masked = self.activ2(self.linear(h_masked)) # [batch_size, max_pred, d_model]
logits_lm = self.fc2(h_masked) # [batch_size, max_pred, vocab_size]
return logits_lm, logits_clsf
model = BERT()
criterion = nn.CrossEntropyLoss()
optimizer = optim.Adadelta(model.parameters(), lr=0.001)
# + id="ShYHlLr-wA_Q" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 101} outputId="304b6e61-d3ea-4659-e1d3-8beca0976876"
for epoch in range(50):
for input_ids, segment_ids, masked_tokens, masked_pos, isNext in loader:
logits_lm, logits_clsf = model(input_ids, segment_ids, masked_pos)
loss_lm = criterion(logits_lm.view(-1, vocab_size), masked_tokens.view(-1)) # for masked LM
loss_lm = (loss_lm.float()).mean()
loss_clsf = criterion(logits_clsf, isNext) # for sentence classification
loss = loss_lm + loss_clsf
if (epoch + 1) % 10 == 0:
print('Epoch:', '%04d' % (epoch + 1), 'loss =', '{:.6f}'.format(loss))
optimizer.zero_grad()
loss.backward()
optimizer.step()
# + id="VMY0ypt8wC9H" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 269} outputId="2e4de07c-c757-473c-ca53-5bc983503feb"
# Predict mask tokens ans isNext
input_ids, segment_ids, masked_tokens, masked_pos, isNext = batch[1]
print(text)
print('================================')
print([idx2word[w] for w in input_ids if idx2word[w] != '[PAD]'])
logits_lm, logits_clsf = model(torch.LongTensor([input_ids]), \
torch.LongTensor([segment_ids]), torch.LongTensor([masked_pos]))
logits_lm = logits_lm.data.max(2)[1][0].data.numpy()
print('masked tokens list : ',[pos for pos in masked_tokens if pos != 0])
print('predict masked tokens list : ',[pos for pos in logits_lm if pos != 0])
logits_clsf = logits_clsf.data.max(1)[1].data.numpy()[0]
print('isNext : ', True if isNext else False)
print('predict isNext : ',True if logits_clsf else False)
# + id="cmlQcIJzUYVI" colab_type="code" colab={}
| 5-2.BERT/BERT-Torch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy
n = numpy.arange(27)
print(n)
n.reshape(3,9)
n.reshape(3,3,3)
import cv2
img = cv2.imread("smallgray.png", 0)
img
imgrgb = cv2.imread("smallgray.png", 1)
imgrgb
cv2.imwrite("newsmallgray.png", img)
img[0:2, 2:4]
img[2,4]
for i in img:
print(type(i))
print(i)
for i in img.flat:
print(i)
ims = numpy.hstack((img, img))
ims
ims = numpy.vstack((img, img))
ims
lst = numpy.hsplit(ims, 5)
lst
lst = numpy.vsplit(ims, 3)
lst
| jupyter/numpy/numpy-test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/RyanBrown55/tests/blob/main/AxioDataPull.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="Gf2z5ErU7XeS" outputId="a2a67d04-397d-49af-9242-e98beffab3f6"
from google.colab import drive
drive.mount("/content/gdrive")
# %cd /content/drive/My Drive/
# + colab={"base_uri": "https://localhost:8080/"} id="wjCPYL9ZtNtu" outputId="d37eda42-938d-4bd9-ea1b-8d673f0f4421"
# !pip install webdriver_manager
# !pip install selenium
# + colab={"base_uri": "https://localhost:8080/"} id="bCNKsiPJwUPc" outputId="56967dee-ccb9-498d-c152-2bd343646928"
# !apt install chromium-chromedriver
# + colab={"base_uri": "https://localhost:8080/"} id="Kb20IoTOtQsC" outputId="03cc1f06-025a-4046-af2b-dca033a94c59"
from webdriver_manager.chrome import ChromeDriverManager
from selenium.webdriver.common.action_chains import ActionChains
import pandas as pd
from selenium import webdriver
from selenium.webdriver.common.by import By
from selenium.common.exceptions import NoSuchElementException
from selenium.webdriver.common.desired_capabilities import DesiredCapabilities
from selenium.webdriver.support.ui import WebDriverWait
from selenium.webdriver.support import expected_conditions as ec
from selenium.webdriver.chrome.options import Options
from selenium.webdriver.support import expected_conditions as EC
from selenium.webdriver.common.action_chains import ActionChains
from selenium.webdriver.common.keys import Keys
import time
# + id="o12rr5wm3qTJ"
path = '/content/gdrive/My Drive/Axiodata'
# + id="H-Yle288XrS4"
import os
for filename in os.listdir('/content/gdrive/My Drive/Axiodata'):
if '.xlsx' in filename:
os.remove('/content/gdrive/My Drive/Axiodata/' + filename)
# + id="uuNaGlkSuLuE" colab={"base_uri": "https://localhost:8080/"} outputId="3c7883b1-030b-4276-cc30-fcb08882fbd7"
def get_axio_time():
drive.mount('/content/drive')
options = Options()
user_agent = 'Mozilla/5.0 (X11; Linux x86_64) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/60.0.3112.50 Safari/537.36'
options.add_argument('--no-sandbox')
options.add_argument('--disable-gpu')
options.add_argument('disable-infobars')
options.add_argument("--disable-extensions")
options.add_argument('--no-sandbox')
options.add_argument('--window-size=1920,1080')
options.add_argument("--headless") #Headless
options.add_argument('user-agent={0}'.format(user_agent))
prefs = {
"download.default_directory": path,
"safebrowsing.disable_download_protection": True,
"directory_upgrade": True}
options.add_experimental_option('prefs', prefs)
driver = webdriver.Chrome('chromedriver',options=options)
driver.command_executor._commands["send_command"] = ("POST", '/session/$sessionId/chromium/send_command')
params = {'cmd': 'Page.setDownloadBehavior', 'params': {'behavior': 'allow', 'downloadPath': path}}
command_result = driver.execute("send_command", params)
url = 'https://axio.realpage.com/Home'
driver.get(url)
user = WebDriverWait(driver, 10).until(EC.element_to_be_clickable((By.XPATH,"/html/body/div[1]/div[1]/div/div[1]/div[1]/div/div[2]/div/div[1]/input")))
user.send_keys('rb<PASSWORD> <EMAIL>')
passw = WebDriverWait(driver, 10).until(EC.element_to_be_clickable((By.XPATH,"/html/body/div[1]/div[1]/div/div[1]/div[1]/div/div[2]/div/div[2]/input")))
passw.send_keys('<PASSWORD>')
signIn = WebDriverWait(driver, 10).until(EC.element_to_be_clickable((By.XPATH,"/html/body/div[1]/div[1]/div/div[1]/div[1]/div/div[2]/div/div[4]/button")))
signIn.click()
pub = WebDriverWait(driver, 10).until(EC.element_to_be_clickable((By.XPATH,'/html/body/nav/div/div[2]/ul[1]/li[7]/a')))
pub.click()
time_df = WebDriverWait(driver, 10).until(EC.element_to_be_clickable((By.XPATH,'/html/body/div[1]/div[1]/div[1]/div/div[3]/ul/li[2]/ul/li[10]/ul/li[2]/div/span')))
time_df.click()
time.sleep(10)
download = WebDriverWait(driver, 10).until(EC.presence_of_element_located((By.XPATH,'/html/body/div[1]/div[1]/div[2]/div[5]/div/div/div[2]/table/tbody/tr[1]/td[3]/div[2]/img[1]')))
driver.execute_script("arguments[0].click();", download)
#make longer for national
#maybe adjust to wait for file in drive -- check if this works
time.sleep(60)
drive.flush_and_unmount()
get_axio_time()
| AxioDataPull.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="RvEEh_BPf3hG" colab_type="code" outputId="fc1ddf10-7c39-49e9-95e1-f72c27ba0453" colab={"base_uri": "https://localhost:8080/", "height": 34}
from google.colab import drive
from pathlib import Path
import tensorflow as tf
import keras
import os
if tf.test.is_gpu_available():
print('Using GPU')
else:
print('Not using GPU')
# + [markdown] id="hEPbm86PjxgT" colab_type="text"
# ### Connect to data
# + id="z4P2VuGVjzrH" colab_type="code" outputId="9b164473-cda7-4c9d-c400-227bcb04826c" colab={"base_uri": "https://localhost:8080/", "height": 34}
drive.mount('/content/gdrive', force_remount=True)
# + id="zTEmNaZuj9rz" colab_type="code" outputId="16d3da7a-39a2-4aaa-8fcc-9dd7e0cc163b" colab={"base_uri": "https://localhost:8080/", "height": 52}
root_path = Path('gdrive/My Drive/ADL/dataset')
print(os.listdir(root_path))
classes = ['smoking','not_smoking']
train_path = root_path/'train'
dirs = [ aDir.name for aDir in train_path.glob('*') if aDir.is_dir() ]
print(dirs)
# + id="lgD0CqCWjNJy" colab_type="code" colab={}
from tensorflow.python.keras.applications import ResNet50
from tensorflow.python.keras.models import Sequential
from tensorflow.python.keras.layers import Dense, Flatten, GlobalAveragePooling2D, BatchNormalization
from tensorflow.python.keras.applications.resnet50 import preprocess_input
from tensorflow.python.keras.preprocessing.image import ImageDataGenerator
from tensorflow.python.keras.preprocessing.image import load_img, img_to_array
# + id="_HDpguuWjjBU" colab_type="code" outputId="1c998da8-9f94-4667-9faa-107cc7727502" colab={"base_uri": "https://localhost:8080/", "height": 34}
data_generator = ImageDataGenerator(horizontal_flip=True,
width_shift_range = 0.4,
height_shift_range = 0.4,
zoom_range=0.3,
rotation_range=20,
)
image_size = 224
batch_size = 32
train_generator = data_generator.flow_from_directory(
root_path/'train',
target_size=(image_size, image_size),
batch_size=batch_size,
class_mode='categorical')
num_classes = len(train_generator.class_indices)
# + id="lMDaZfPrm3Yn" colab_type="code" outputId="da8c1f57-6225-49cc-eeed-c0d52ee782f1" colab={"base_uri": "https://localhost:8080/", "height": 141}
keras.applications.ResNet50(include_top=True, weights='imagenet', input_tensor=None, input_shape=None, pooling=None, classes=1000)
# + id="qDGfB32rjtBP" colab_type="code" colab={}
def resnet50_model(num_classes=2):
model = Sequential()
#keras.applications.resnet.ResNet50(include_top=True, weights='imagenet', input_tensor=None, input_shape=None, pooling=None, classes=1000)
model.add(ResNet50(include_top=False, pooling='avg', weights='imagenet'))
model.add(Flatten())
model.add(BatchNormalization())
model.add(Dense(2048, activation='relu'))
model.add(BatchNormalization())
model.add(Dense(1024, activation='relu'))
model.add(BatchNormalization())
model.add(Dense(num_classes, activation='softmax'))
model.layers[0].trainable = False
return model
model = resnet50_model(num_classes=num_classes)
# + id="DZArXBKEmLcW" colab_type="code" colab={}
#model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy','error'])
# + id="EoQl8AcznecY" colab_type="code" outputId="608806d5-601d-4cb4-ab49-c701c59455e1" colab={"base_uri": "https://localhost:8080/", "height": 34}
from keras import backend as K
K.tensorflow_backend._get_available_gpus()
#from tensorflow.python.client import device_lib
#print(device_lib.list_local_devices())
# + id="bqAShQvopQIA" colab_type="code" outputId="18cebf20-2652-4abf-dbb8-e68b186348db" colab={"base_uri": "https://localhost:8080/", "height": 382}
from keras.models import Sequential
from keras.layers import Dense, Activation
output_dim = nb_classes = 2
input_dim = 2048
# model = Sequential()
# model.add(Dense(output_dim, input_dim=input_dim, activation='softmax'))
# batch_size = 128
# nb_epoch = 20
model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
single_batch = next(iter(train_generator))
model.fit(
single_batch[0],
single_batch[1],
steps_per_epoch=1, # int(count/batch_size) + 1,
# batch_size=batch_size,
epochs=10)
# model.fit_generator(
# train_generator,
# steps_per_epoch=1, # int(count/batch_size) + 1,
# # batch_size=batch_size,
# epochs=10)
# + id="ObT_LAPznLg5" colab_type="code" outputId="7bf59edb-f77a-439c-e393-d79d4106adbe" colab={"base_uri": "https://localhost:8080/", "height": 443}
count = sum([len(files) for r, d, files in os.walk("../input/flowers-recognition/flowers/flowers/")])
model.fit_generator(
train_generator,
steps_per_epoch=int(count/batch_size) + 1,
epochs=10)
# + id="_GFtY_mInNyA" colab_type="code" colab={}
| notebooks/reference.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:anamic]
# language: python
# name: conda-env-anamic-py
# ---
# ### Field of View Simulator
#
# Show how to a microscope field of view with many microtubules.
# +
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
from pathlib import Path
import sys
sys.path.append("../")
import anamic
import numpy as np
import matplotlib.pyplot as plt
# +
# Common Parameters
pixel_size = 110 # nm/pixel
image_size_pixel = 512
# Per image parameters
image_parameters = {}
image_parameters['n_mt'] = {}
image_parameters['n_mt']['values'] = np.arange(80, 120)
image_parameters['n_mt']['prob'] = 'uniform'
image_parameters['signal_mean'] = {}
image_parameters['signal_mean']['values'] = {'loc': 700, 'scale': 10}
image_parameters['signal_mean']['prob'] = 'normal'
image_parameters['signal_std'] = {}
image_parameters['signal_std']['values'] = {'loc': 100, 'scale': 1}
image_parameters['signal_std']['prob'] = 'normal'
image_parameters['bg_mean'] = {}
image_parameters['bg_mean']['values'] = {'loc': 500, 'scale': 10}
image_parameters['bg_mean']['prob'] = 'normal'
image_parameters['bg_std'] = {}
image_parameters['bg_std']['values'] = {'loc': 24, 'scale': 1}
image_parameters['bg_std']['prob'] = 'normal'
image_parameters['noise_factor'] = {}
image_parameters['noise_factor']['values'] = {'loc': 1, 'scale': 0.1}
image_parameters['noise_factor']['prob'] = 'normal'
image_parameters['noise_factor']['values'] = [0.5]
image_parameters['noise_factor']['prob'] = [1]
image_parameters['mask_line_width'] = 4 # pixel
image_parameters['mask_backend'] = 'skimage'
# Per microtubule parameters.
microtubule_parameters = {}
microtubule_parameters['n_pf'] = {}
microtubule_parameters['n_pf']['values'] = [11, 12, 13, 14, 15]
microtubule_parameters['n_pf']['prob'] = [0.05, 0.05, 0.3, 0.1, 0.5]
microtubule_parameters['mt_length_nm'] = {}
microtubule_parameters['mt_length_nm']['values'] = np.arange(500, 10000)
microtubule_parameters['mt_length_nm']['prob'] = 'uniform'
microtubule_parameters['taper_length_nm'] = {}
microtubule_parameters['taper_length_nm']['values'] = np.arange(0, 1000)
microtubule_parameters['taper_length_nm']['prob'] = 'uniform'
microtubule_parameters['labeling_ratio'] = {}
microtubule_parameters['labeling_ratio']['values'] = [0.08, 0.09, 0.10, 0.11, 0.12, 0.13]
microtubule_parameters['labeling_ratio']['prob'] = 'uniform'
microtubule_parameters['pixel_size'] = pixel_size # nm/pixel
microtubule_parameters['x_offset'] = 2000 # nm
microtubule_parameters['y_offset'] = 2000 # nm
microtubule_parameters['psf_size'] = 135 # nm
image, masks, mts = anamic.simulator.create_fov(image_size_pixel, pixel_size, microtubule_parameters, image_parameters, return_positions=True)
# +
fig, axs = plt.subplots(ncols=2, figsize=(12, 6))
im = axs[0].imshow(image)
#fig.colorbar(im, ax=axs[0])
axs[1].imshow(masks.max(axis=0))
# -
| notebooks/Howto/2_Simulate_Many_Microtubules.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # COCO Keypoints
# Simple example on how to parse keypoints for the coco annotation format. For demonstration purposes we will be using the samples present on the repo instead of the full COCO dataset.
from icevision.all import *
data_dir = Path('/home/lgvaz/git/icevision/samples')
class_map = ClassMap(['person'])
parser = parsers.COCOKeyPointsParser(annotations_filepath=data_dir/'keypoints_annotations.json', img_dir=data_dir/'images')
records = parser.parse(data_splitter=SingleSplitSplitter())[0]
record = records[1]
show_record(record, figsize=(10,10), class_map=class_map)
test_tfms = tfms.A.Adapter([*tfms.A.resize_and_pad(512), tfms.A.Normalize()])
test_ds = Dataset(records, test_tfms)
show_sample(test_ds[0], figsize=(10,10), display_bbox=False)
| notebooks/coco_keypoints.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### <font color='green'>1. Description<font>
#
# Sentiment classification using Amazon review dataset (multi class classification).
# Dataset can be downloaded from https://s3.amazonaws.com/amazon-reviews-pds/tsv/amazon_reviews_us_Books_v1_02.tsv.gz
#
# The consumer reviews serve as feedback for businesses in terms of performance, product quality, and consumer service. An online review typically consists of free-form text and a star rating out of 5. The problem of predicting a user’s star rating for a product, given the user’s text review for that product is lately become a popular, albeit hard, problem in machine learning.
# Using this dataset, we train a classifier to predict product rating based on the review text.
#
# Predicting the ratings based on the text is particulary difficult tasks. The primary reason for the difficulty is that two person can provide different ratings for writing similar reviews. As the scale for ratings increases (scale of 5 to scale of 10), the tasks become increasingly difficult.
# ### <font color='green'>2. Data Preprocessing<font>
#
# For amazon review classification we will perform some data preparation and data cleaning steps. We will generate feature vectors using sklearn TF-IDF for review text.
import os
import pandas as pd
from collections import OrderedDict
def create_embed(x_train, x_test):
from sklearn.feature_extraction.text import CountVectorizer, TfidfTransformer
count_vect = CountVectorizer()
x_train_counts = count_vect.fit_transform(x_train)
x_test_counts = count_vect.transform(x_test)
tfidf_transformer = TfidfTransformer()
x_train_tfidf = tfidf_transformer.fit_transform(x_train_counts)
x_test_tfidf = tfidf_transformer.transform(x_test_counts)
return x_train_tfidf, x_test_tfidf
def preprocess_data(fname):
df = pd.read_csv(fname, sep='\t', error_bad_lines=False)
df = df[["review_body", "star_rating"]]
df = df.dropna().drop_duplicates().sample(frac=1) # why sampling?
print("Dataset contains {} reviews".format(df.shape[0]))
rating_categories = df["star_rating"].value_counts()
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(df["review_body"],
df["star_rating"],
random_state = 42)
x_train, x_test = create_embed(x_train, x_test)
return x_train, x_test, y_train, y_test, rating_categories
# +
#---- Data Preparation ----
# Please uncomment the below lines to download and unzip the dataset.
# #!wget -N https://s3.amazonaws.com/amazon-reviews-pds/tsv/amazon_reviews_us_Books_v1_02.tsv.gz
# #!gunzip amazon_reviews_us_Books_v1_02.tsv.gz
# #!mv amazon_reviews_us_Books_v1_02.tsv datasets
DATA_FILE = "datasets/amazon_reviews_us_Books_v1_02.tsv/amazon_reviews_us_Books_v1_02.tsv"
x_train, x_test, y_train, y_test, rating_categories = preprocess_data(DATA_FILE)
print("shape of train data: {}".format(x_train.shape))
print("shape of test data: {}".format(x_test.shape))
# -
# Label distribution summary
ax = rating_categories.plot(kind='bar', title='Label Distribution').\
set(xlabel="Rating Id's", ylabel="No. of reviewes")
# ### <font color='green'> 3. Algorithm Evaluation<font>
import time
from sklearn import metrics
train_time = []
test_time = []
accuracy = []
precision = []
recall = []
f1 = []
estimator_name = []
def evaluate(estimator, estimator_nm,
x_train, y_train,
x_test, y_test):
estimator_name.append(estimator_nm)
start_time = time.time()
estimator.fit(x_train, y_train)
train_time.append(round(time.time() - start_time, 4))
start_time = time.time()
pred_y = estimator.predict(x_test)
test_time.append(round(time.time() - start_time, 4))
accuracy.append(metrics.accuracy_score(y_test, pred_y))
precision.append(metrics.precision_score(y_test, pred_y, average='macro'))
recall.append(metrics.recall_score(y_test, pred_y, average='macro'))
f1.append(metrics.f1_score(y_test, pred_y, average='macro'))
target_names = ['rating 1.0', 'rating 2.0', 'rating 3.0', 'rating 4.0', 'rating 5.0']
return metrics.classification_report(y_test, pred_y, target_names=target_names)
# #### 3.1 Multinomial LogisticRegression
# +
#1. Demo: Multinomial LogisticRegression
import frovedis
TARGET = "multinomial_logistic_regression"
from frovedis.exrpc.server import FrovedisServer
FrovedisServer.initialize("mpirun -np 8 " + os.environ["FROVEDIS_SERVER"])
from frovedis.mllib.linear_model import LogisticRegression as frovLogisticRegression
f_est = frovLogisticRegression(max_iter=3100, penalty='none', \
lr_rate=0.001, tol=1e-8)
E_NM = TARGET + "_frovedis_" + frovedis.__version__
f_report = evaluate(f_est, E_NM, \
x_train, y_train, x_test, y_test)
f_est.release()
FrovedisServer.shut_down()
import sklearn
from sklearn.linear_model import LogisticRegression as skLogisticRegression
s_est = skLogisticRegression(max_iter = 3100, penalty='none', \
tol = 1e-8, n_jobs = 12)
E_NM = TARGET + "_sklearn_" + sklearn.__version__
s_report = evaluate(s_est, E_NM, \
x_train, y_train, x_test, y_test)
# LogisticRegression: Precision, Recall and F1 score for each class
print("Frovedis LogisticRegression metrices: ")
print(f_report)
print("Sklearn LogisticRegression metrices: ")
print(s_report)
# -
# #### 3.2 MultinomialNB
# +
#2. Demo: MultinomialNB
import frovedis
TARGET = "multinomial_naive_bayes"
from frovedis.exrpc.server import FrovedisServer
FrovedisServer.initialize("mpirun -np 8 " + os.environ["FROVEDIS_SERVER"])
from frovedis.mllib.naive_bayes import MultinomialNB as fMNB
f_est = fMNB()
E_NM = TARGET + "_frovedis_" + frovedis.__version__
f_report = evaluate(f_est, E_NM, \
x_train, y_train, x_test, y_test)
f_est.release()
FrovedisServer.shut_down()
import sklearn
from sklearn.naive_bayes import MultinomialNB as sMNB
s_est = sMNB()
E_NM = TARGET + "_sklearn_" + sklearn.__version__
s_report = evaluate(s_est, E_NM, \
x_train, y_train, x_test, y_test)
# MultinomialNB: Precision, Recall and F1 score for each class
print("Frovedis MultinomialNB metrices: ")
print(f_report)
print("Sklearn MultinomialNB metrices: ")
print(s_report)
# -
# #### 3.3 Bernoulli Naive Bayes
# +
# Demo: Bernoulli Naive Bayes
import frovedis
TARGET = "bernoulli_naive_bayes"
from frovedis.exrpc.server import FrovedisServer
FrovedisServer.initialize("mpirun -np 8 " + os.environ["FROVEDIS_SERVER"])
from frovedis.mllib.naive_bayes import BernoulliNB as frovNB
f_est = frovNB(alpha=1.0)
E_NM = TARGET + "_frovedis_" + frovedis.__version__
f_report = evaluate(f_est, E_NM, \
x_train, y_train, x_test, y_test)
f_est.release()
FrovedisServer.shut_down()
import sklearn
from sklearn.naive_bayes import BernoulliNB as skNB
s_est = skNB(alpha=1.0)
E_NM = TARGET + "_sklearn_" + sklearn.__version__
s_report = evaluate(s_est, E_NM, \
x_train, y_train, x_test, y_test)
# Precision, Recall and F1 score for each class
print("Frovedis Bernoulli Naive Bayes metrices: ")
print(f_report)
print("Sklearn Bernoulli Naive Bayes metrices: ")
print(s_report)
# -
# ### <font color='green'> 4. Performance summary<font>
summary = pd.DataFrame(OrderedDict({ "estimator": estimator_name,
"train time": train_time,
"test time": test_time,
"accuracy": accuracy,
"precision": precision,
"recall": recall,
"f1-score": f1
}))
summary
| doc/notebook/02_1_amazonreview_multiclass_classification_sparse.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 创建实验环境
# ## DataFrame中的同名列
# 大部分情况下,DataFrame都不鼓励DataFrame中出现同名列,单并不是完全禁止。
# ### merge的情况
from pandas import DataFrame, Series
import pandas as pd
df_1 = DataFrame({'A':[1,2,3],'B':[1,2,3]})
df_1
df_2 = DataFrame({'A':[1,2,3],'B':[1,2,3]})
df_2
df_1.merge(df_2,on='A')
# 可以看到merge函数会自动给两张表的同名列加上前缀以区分
# 如果强行去掉前缀, 那么会报错
df_1.merge(df_2, on = 'A', suffixes = ['',''])
# reset_index的时候,如果index的列名和已有的列名重复,那么会报错
try:
df = DataFrame({'A':[1,2,3]})
df.index.name = 'A'
df.reset_index()
except Exception as e:
print(e)
df = DataFrame({'A':[1,2,3,4],'B':[1,2,3,4]})
df
# pd.concat则不会检查是否存在重名列
df = pd.concat([df_1, df_2],axis=1)
df
# renname函数也不会去检查列名是否一致
df = DataFrame({'A':[1,2,3],'B':[1,2,3]})
df.rename(columns = {'B':'A'})
# insert函数则会提供allow_duplicates参数,让用户去决定是否允许重名列存在。
df_1.insert(0,'B',1,allow_duplicates=True)
df_1
# 如果同名列存在, 那么用列名去选取列的是否, 会同时把所有的列名都选择出来
# ### 总结:
# DataFrame中对于列名是否一致, 不同的函数表现并不一致。 有些允许, 有些禁止, 有些交由用户自己决定。 实践中, 我并没有遇到什么场景需要让两列的列名相同,建议避免出现这种情况。
# +
user = 'test'
password = '<PASSWORD>'
dbname = 'test'
port = '5432'
schema = 'test'
url = 'postgresql://{user}:{password}@localhost:{port}/{dbname}'.format(**locals())
# +
from sqlalchemy import create_engine, join, select
from sqlalchemy import Table, Column, Integer, String, MetaData, ForeignKey
from pandas import DataFrame, Series
drop_sql = 'drop table table_one'
engine = create_engine(url)
metadata = MetaData()
table_one = Table('table_one', metadata,
Column('A', String),
Column('B', Integer),
Column('C', Integer),
schema = 'test'
)
with engine.connect() as conn:
try:
conn.execute(drop_sql)
except:
pass
metadata.create_all(engine)
# -
# 如果创表的时候用了大写, 时候, 只能用'A'来选择
# +
s = "select 'A' from test.table_one"
with engine.connect() as conn:
print(conn.execute(s).fetchall())
# -
# sqlalchemy的话可以用A选择, 自动转成了'A'
# +
t = Table('table_one',metadata,schema = 'test')
s = select(
[t.c.A]
)
with engine.connect() as conn:
print(conn.execute(s).fetchall())
# -
print(s.compile())
# 如果建表的时候用小写, 那么怎么选都可以
# +
from sqlalchemy import create_engine, join, select
from sqlalchemy import Table, Column, Integer, String, MetaData, ForeignKey
from pandas import DataFrame, Series
drop_sql = 'drop table table_one'
engine = create_engine(url)
metadata = MetaData()
table_one = Table('table_one', metadata,
Column('a', String),
Column('b', Integer),
Column('c', Integer),
schema = 'test'
)
with engine.connect() as conn:
try:
conn.execute(drop_sql)
except:
pass
metadata.create_all(engine)
# +
s = "select 'a' from test.table_one"
with engine.connect() as conn:
print(conn.execute(s).fetchall())
# +
t = Table('table_one',metadata,schema = 'test')
s = select(
[t.c.a]
)
with engine.connect() as conn:
print(conn.execute(s).fetchall())
# -
# 最简单的方法就是sqlalchemy中一律使用小写建表
# +
如果直接用SQL建表
sql = '''
creat test.table(
A INTEGER,
B INTEGER
)
'''
# -
with engine.connect() as conn:
try:
conn.execute(drop_sql)
except:
pass
conn.execute(sql)
# +
user = 'test'
password = '<PASSWORD>'
dbname = 'test'
port = '5432'
schema = 'test'
url = 'postgresql://{user}:{password}@localhost:{port}/{dbname}'.format(**locals())
create_schema_sql = 'CREATE SCHEMA IF NOT EXISTS {schema_name}'.format(schema_name=schema)
engine = create_engine(url)
metadata = MetaData()
table_one = Table('table_one', metadata,
Column('A', String),
Column('B', Integer),
Column('C', Integer),
schema = schema
)
table_two = Table('table_two', metadata,
Column('A', String),
Column('B', Integer),
Column('D', Integer),
schema = schema
)
with engine.connect() as conn:
conn.execute(create_schema_sql)
metadata.create_all(engine)
df_one = DataFrame.from_records(
[
('a',1,1,),
('a',2,2,),
('b',1,3,),
('b',2,4,),
]
,columns = ['a','b','c']
)
df_one
df_two = DataFrame.from_records(
[
('a',1,11,),
('a',2,22,),
('b',1,33,),
('b',2,44,)
]
,columns = ['a','b','d']
)
df_two
with engine.connect() as conn:
df_one.to_sql('table_one', engine, if_exists = 'replace', schema = schema,index = False)
df_two.to_sql('table_two', engine, if_exists = 'replace', schema = schema, index = False)
# -
select
t.a
from
(
select
*
from
table_one
full join
table_two
on table_one.a=table_two.a and table_one.b=table_two.b
) t;
# +
# astype(object)
# -
import numpy as np
import pandas as pd
# +
from sqlalchemy import case, literal, cast, literal_column, String, create_engine, MetaData, Table, select, and_, or_, func
import pandas as pd
import os
os.environ['NLS_LANG'] = 'SIMPLIFIED CHINESE_CHINA.UTF8'
import warnings
warnings.filterwarnings('ignore')
from datetime import datetime, date
# -
# # PITFALL_1
# ## np.nan vs None
# ### 往数据库中写入时np.nan不可处理,需转换成None
# 在pandas.DataFrame中,np.nan和None是等价的。
#
# 但是,column.type是float64时,是以np.nan形式存在的;column.type是object时,是以None存在的。
#
# 而往数据库里insert(dataframe.to_records('dict'))时,存在np.nan会报错。必须转化成None。
# to_sql中的表现
df = pd.DataFrame({'A':[np.nan, 1, 2], 'B':[1,2,None]})
print(df)
for column in df:
print(df.loc[df[column].isnull(),column])
type(df['A'].iloc[0])
type(df['B'].iloc[2])
df = pd.DataFrame({'A':[np.nan, 1, 2], 'B':[1,2,None]})
print(df)
for column in df:
print(df.loc[pd.isnull(df[column]),column])
# +
df = pd.DataFrame({'A':[np.nan, '1', '2'], 'B':['1','2',None]})
print(df)
for column in df:
df.loc[pd.isnull(df[column]),column] = None
df[column] = df[column].astype(float)
df
# +
df = pd.DataFrame({'A':[np.nan, '1', '2'], 'B':['1','2',None]})
print(df)
for column in df:
df[column] = df[column].astype(float)
df.loc[pd.isnull(df[column]),column] = None
df
# +
df = pd.DataFrame({'A':[np.nan, '1', '2'], 'B':['1','2',None]})
print(df)
for column in df:
df.loc[pd.isnull(df[column]),column] = None
df[column] = df[column].astype(object)
df
# -
# # PITFALL_2
# ## 判断两个dataframe是否相同
# __注意行和列的顺序都要一致,且数据类型要一致。__
# * assert df1.equals(df2)
# * np.testing.assert_equal(df1.values, df2.values)
#
# pandas中默认空值=空值(np.nan 和 None等价),np认为它们不一样
#
# df1.values == df2.values 无法处理空值的情况,慎用!
df1 = pd.DataFrame({'A':[1,3,2],'B':[4,6,None]})
df2 = pd.DataFrame({'A':[1,2,3],'B':[4,None,6]})
df2.dtypes
df1
df2
df2.values
df1.sort_values('A').values == df2.values
df2.equals(df1) #错误原因:顺序不对
df2.equals(df1.sort_values('A')) #顺序对了,但是index不一致
new_df1 = df1.sort_values('A').reset_index().drop('index', axis=1)
new_df2 = df2.sort_values('A').reset_index().drop('index', axis=1)
new_df2.equals(new_df1)#此方法是先排序再删掉index再重构index
np.testing.assert_equal(df1.sort_values('A').values, df2.values)
#此方法只要顺序对了就好,index不影响,但是存在下面的问题
# ### 类型一致时:pandas中默认空值=空值(np.nan 和 None等价),np认为它们不一样
df4 = df2.copy()
df4['B']=df4['B'].astype(object)
df4.dtypes
df4
df3=df4.copy()
df3.loc[1,'B']=None
df3
df3.dtypes
df3.equals(df4)
np.testing.assert_equal(df3.values, df4.values)
# # PIDFALL_3
# ## 列名重复问题
#
# 在panda中,表格merge之后,不同表中的相同名字的列会被以_x,_y结尾保留下来,
#
# 但在sqlalchemy中,列名还维持在原来的状态,编写sql语句时不会报错,但是拉取数据结果时,会报错
df5 = pd.DataFrame({'A':['a','b','c'],'B':['x','y','x'],'C':[4,6,7]})
df6 = pd.DataFrame({'B':['x','y','x'],'C':[4,6,8]})
df5
df6
df5.merge(df6,on='B',how='left')
engine = create_engine('postgresql://datascience:Jrsjkxzbbd2333@localhost/etl')
df5.to_sql('df5',engine,index=False,if_exists='replace')
df6.to_sql('df6',engine,index=False,if_exists='replace')
metadata=MetaData(engine)
table5 = Table('df5',metadata,autoload=True)
table6 = Table('df6',metadata,autoload=True)
s = select([table5, table6]).where(table5.c.B == table6.c.B).alias()
print(s.c.keys()) # 是否有columns的attribute
# +
# 在数据库中直接跑sql的情况
# -
pd.read_sql(s,engine)
# 什么鬼?两个B,而且C不会改名字!!!我要把B拉出来看看~
pd.read_sql(select([s.c.B]),engine)#B拉不出来,因为有两个B,不知道拉哪个,尽管两个B的值是一样的
print(sql.compile(compile_kwargs={"literal_binds": True}))
# # PIDFALL_4
df7 = pd.DataFrame({'A':['a','b','c'],'B':['x','y',None],'C':[4,6,7]})
df8 = pd.DataFrame({'A':['a','b','c'],'B':['x','y',None],'D':[1,2,3]})
df7
df8
df7.merge(df8,on=['A','B'],how='left')
# +
df7.to_sql('df7',engine,index=False,if_exists='replace')
df8.to_sql('df8',engine,index=False,if_exists='replace')
table7 = Table('df7',metadata,autoload=True)
table8 = Table('df8',metadata,autoload=True)
# -
s = select(
[
table7,
table8.c.D
]
).where(
and_(
table7.c.B == table8.c.B,
table7.c.A == table8.c.A
)
).alias()
pd.read_sql(s,engine)
s2 = select(
[
table7,
table8.c.D
]
).select_from(
table7.outerjoin(
table8,
and_(
table7.c.B == table8.c.B,
table7.c.A == table8.c.A
)
)
).alias()
pd.read_sql(s2,engine)
s3 = select(
[
table7,
table8.c.D
]
).select_from(
table7.outerjoin(
table8,
and_(
table7.c.B == table8.c.B,
table7.c.A == table8.c.A
),full=True
)
).alias()
pd.read_sql(s3,engine)
# 若想要pandas里面的效果,即None=None,
s4 = select(
[
table7,
table8.c.D
]
).select_from(
table7.outerjoin(
table8,
and_(
table7.c.A == table8.c.A,
or_(
table7.c.B == table8.c.B,
and_(
table7.c.B ==None,
table8.c.B ==None
)
)
)
)
).alias()
pd.read_sql(s4,engine)
metadata.drop_all(engine, checkfirst=True)
from pandas import Series
from numpy import NaN
s = Series([None, NaN, 'a'])
s
s == None
s == NaN
s.isnull()
s is None
s.apply(lambda s:s is None)
# +
a = [1,2]
b = [1,2]
# -
a==b
a is b
# +
a = [1,2]
b = a
a is b
# -
from pandas import DataFrame
a=DataFrame(index=[1,2,3,4,5])
b=DataFrame(index=['1','2','3','4','5'])
a
b
# +
and &
a={'c':1233}
a=None
if (a is not None) & a['c'] =1233 :
if pd.notnull(a) and a['c'] =1233 :
# -
s1 = Series([1,2,3])
s2 = Series([])
s2 and s1
bool([1,2,3] and [])
[1,2,3] or []
# +
x = [1,2,3,4]
x or [1,1,1]
# -
s1 = Series([True, False])
s1
s2 = Series([True, True])
s2
s1&s2
| 0007_pandas_vs_sql/0008_DataFrame_vs_SQL_ch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="LaTSGa8RDC5H"
# In this module, we fit a linear model with positive constraints on the regression coefficients and compare the estimated coefficients to a classic linear regression.
# + id="pPDZ_HkHI3Zj" executionInfo={"status": "ok", "timestamp": 1636401648071, "user_tz": -480, "elapsed": 3879, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gict81qvzCzYor7OD2HAqOMscBAGuRrSJShYazDZg=s64", "userId": "02100760507760735849"}} outputId="2c117dce-4f0f-4697-ede1-46ee39e3bd25" colab={"base_uri": "https://localhost:8080/"}
# !pip install -U scikit-learn
# + id="z66zVnMLC_he" executionInfo={"status": "ok", "timestamp": 1636401648072, "user_tz": -480, "elapsed": 16, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gict81qvzCzYor7OD2HAqOMscBAGuRrSJShYazDZg=s64", "userId": "02100760507760735849"}}
import numpy as np
import matplotlib.pyplot as plt
from sklearn.metrics import r2_score
# + [markdown] id="7akUZZ6QDIMP"
# Generate some random data
# + id="_afHUDeqDHVm" executionInfo={"status": "ok", "timestamp": 1636401648072, "user_tz": -480, "elapsed": 15, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gict81qvzCzYor7OD2HAqOMscBAGuRrSJShYazDZg=s64", "userId": "02100760507760735849"}}
np.random.seed(42)
n_samples, n_features = 200, 50
X = np.random.randn(n_samples, n_features)
true_coef = 3 * np.random.randn(n_features)
# Threshold coefficients to render them non-negative
true_coef[true_coef < 0] = 0
y = np.dot(X, true_coef)
# Add some noise
y += 5 * np.random.normal(size=(n_samples,))
# + [markdown] id="P0EZySD6ESx1"
# Split the data in train set and test set
# + id="pUrHb2xbESJf" executionInfo={"status": "ok", "timestamp": 1636401648073, "user_tz": -480, "elapsed": 16, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gict81qvzCzYor7OD2HAqOMscBAGuRrSJShYazDZg=s64", "userId": "02100760507760735849"}}
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.5)
# + [markdown] id="OLuBbX5NEg6h"
# Fit the Non-Negative least squares.
# + colab={"base_uri": "https://localhost:8080/"} id="dQ13PiQNFHgZ" executionInfo={"status": "ok", "timestamp": 1636401648073, "user_tz": -480, "elapsed": 16, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gict81qvzCzYor7OD2HAqOMscBAGuRrSJShYazDZg=s64", "userId": "02100760507760735849"}} outputId="ab4dd328-5463-4e61-a858-7f0b98408d66"
from sklearn.linear_model import LinearRegression
reg_nnls = LinearRegression(positive=True)
y_pred_nnls = reg_nnls.fit(X_train, y_train).predict(X_test)
r2_score_nnls = r2_score(y_test, y_pred_nnls)
print("NNLS R2 score", r2_score_nnls)
# + [markdown] id="zshz57htF59q"
# Fit an OLS.
# + colab={"base_uri": "https://localhost:8080/"} id="RVE6O69bFmSc" executionInfo={"status": "ok", "timestamp": 1636401648073, "user_tz": -480, "elapsed": 14, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gict81qvzCzYor7OD2HAqOMscBAGuRrSJShYazDZg=s64", "userId": "02100760507760735849"}} outputId="72c71aea-17a0-4e38-a6b4-371b212a5c61"
reg_ols = LinearRegression()
y_pred_ols = reg_ols.fit(X_train, y_train).predict(X_test)
r2_score_ols = r2_score(y_test, y_pred_ols)
print("OLS R2 score", r2_score_ols)
# + [markdown] id="NWt6wbk0HbBf"
# Comparing the regression coefficients between OLS and NNLS, we can observe they are highly correlated (the dashed line is the identity relation), but the non-negative constraint shrinks some to 0. The Non-Negative Least squares inherently yield sparse results.
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="7240s1VSF9Nf" executionInfo={"status": "ok", "timestamp": 1636401648074, "user_tz": -480, "elapsed": 14, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gict81qvzCzYor7OD2HAqOMscBAGuRrSJShYazDZg=s64", "userId": "02100760507760735849"}} outputId="a26700d9-aa91-4057-d23b-a6a62b9dfd7e"
fig, ax = plt.subplots()
ax.plot(reg_ols.coef_, reg_nnls.coef_, linewidth=0, marker=".")
low_x, high_x = ax.get_xlim()
low_y, high_y = ax.get_ylim()
low = max(low_x, low_y)
high = min(high_x, high_y)
ax.plot([low, high], [low, high], ls="--", c=".3", alpha=0.5)
ax.set_xlabel("OLS regression coefficients", fontweight="bold")
ax.set_ylabel("NNLS regression coefficients", fontweight="bold")
| Week 2/Non-negative least squares.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# Azure ML & Azure Databricks notebooks by <NAME>.
#
# Copyright (c) Microsoft Corporation. All rights reserved.
#
# Licensed under the MIT License.
# 
# This notebook uses image from ACI notebook for deploying to AKS.
# +
import azureml.core
# Check core SDK version number
print("SDK version:", azureml.core.VERSION)
# -
# Set auth to be used by workspace related APIs.
# For automation or CI/CD ServicePrincipalAuthentication can be used.
# https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.authentication.serviceprincipalauthentication?view=azure-ml-py
auth = None
# +
from azureml.core import Workspace
ws = Workspace.from_config(auth = auth)
print('Workspace name: ' + ws.name,
'Azure region: ' + ws.location,
'Subscription id: ' + ws.subscription_id,
'Resource group: ' + ws.resource_group, sep = '\n')
# +
# List images by ws
from azureml.core.image import ContainerImage
for i in ContainerImage.list(workspace = ws):
print('{}(v.{} [{}]) stored at {} with build log {}'.format(i.name, i.version, i.creation_state, i.image_location, i.image_build_log_uri))
# -
from azureml.core.image import Image
myimage = Image(workspace=ws, name="aciws")
# +
#create AKS compute
#it may take 20-25 minutes to create a new cluster
from azureml.core.compute import AksCompute, ComputeTarget
# Use the default configuration (can also provide parameters to customize)
prov_config = AksCompute.provisioning_configuration()
aks_name = 'ps-aks-demo2'
# Create the cluster
aks_target = ComputeTarget.create(workspace = ws,
name = aks_name,
provisioning_configuration = prov_config)
aks_target.wait_for_completion(show_output = True)
print(aks_target.provisioning_state)
print(aks_target.provisioning_errors)
# -
from azureml.core.webservice import Webservice
help( Webservice.deploy_from_image)
# +
from azureml.core.webservice import Webservice, AksWebservice
from azureml.core.image import ContainerImage
#Set the web service configuration (using default here with app insights)
aks_config = AksWebservice.deploy_configuration(enable_app_insights=True)
#unique service name
service_name ='ps-aks-service'
# Webservice creation using single command, there is a variant to use image directly as well.
aks_service = Webservice.deploy_from_image(
workspace=ws,
name=service_name,
deployment_config = aks_config,
image = myimage,
deployment_target = aks_target
)
aks_service.wait_for_deployment(show_output=True)
# -
aks_service.deployment_status
#for using the Web HTTP API
print(aks_service.scoring_uri)
print(aks_service.get_keys())
# +
import json
#get the some sample data
test_data_path = "AdultCensusIncomeTest"
test = spark.read.parquet(test_data_path).limit(5)
test_json = json.dumps(test.toJSON().collect())
print(test_json)
# -
#using data defined above predict if income is >50K (1) or <=50K (0)
aks_service.run(input_data=test_json)
#comment to not delete the web service
aks_service.delete()
#image.delete()
#model.delete()
aks_target.delete()
# 
| how-to-use-azureml/azure-databricks/amlsdk/deploy-to-aks-existingimage-05.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#Libería para Cargar datos
from xlrd import open_workbook
book = open_workbook('../data/raw/refugios_nayarit.xlsx',on_demand=True)
for sheet in book.sheet_names():
print(sheet)
sheet = book.sheet_by_name("1-20")
sheet.row(6)
sheet.row(-2)
sheet_2 = book.sheet_by_name("21-40")
sheet_2.row(6)
sheet_2.row(-2)
sheet_3 = book.sheet_by_name("41-60")
sheet_3.row(6)
sheet_3.row(-2)
sheet_3.row(-2)[0].value
sheet_4 = book.sheet_by_name("406-425")
sheet_4.row(6)
sheet_4.row(-2)
# El dataset está aparentemente estructurado.
for sheet in book.
len(sheet_4.row(-2))
output_list = []
col_names = ["no", "refugio", "municipio","direccion","uso","servicios","capacidad", "latitud", "longitud","altitud","responsable","telefono" ]
for name in book.sheet_names():
output_dict = {}
sheet = book.sheet_by_name(name)
for i in range(6,sheet.nrows-1,1):
output_dict = { col_names[j]: sheet.row(i)[j].value for j in range(0,12,1)}
output_list.append(output_dict)
len(output_list)
import pandas as pd
data_df = pd.DataFrame.from_dict(output_list)
data_df.to_csv("../data/interim/excel_loaded_data.csv", index=False)
data_df[data_df["latitud"]==""]
def normalize_data(s):
s = s.lower()
return ''.join(c for c in unicodedata.normalize('NFD', s)
if unicodedata.category(c) != 'Mn')
a = normalize_data("ESCUELA PREPARATORIA PUENTE DE CAMOTLAN LA YESCA, NAYARIT")
import geocoder
g = geocoder.google(a)
g.latlng
blanks = data_df[data_df["latitud"]==""]
for blank in blanks.iterrows():
a = blank
print(a)
a[1]
coords = []
for blank in blanks.iterrows():
data = blank[1]
address = normalize_data(data["refugio"] + " "+ data["direccion"]+" " + data["municipio"]+ ", nayarit")
g = geocoder.google(address)
coords.append(g.latlng)
coords
def coords_converter(string):
string = string.strip()
degrees, minutes, seconds = coords_splitter(string)
decimal_coords = degrees + minutes/60.0 + seconds/3600
return decimal_coords
coords_splitter("21º09\'43.32")
def coords_splitter(string):
string_array = string.split("º")
secondpart = string_array[1].split("\'")
degrees = float(string_array[0])
minutes = float(secondpart[0])
seconds = float(secondpart[1].replace('"',''))
return [degrees, minutes, seconds]
coords_converter("21º09\'43.32\"")
# Verificar contenido de la base de datos hasta el momento.
def is_field_congruent(field):
expectedstrings = ["º", "\'", '"']
results = [constant in field for constant in expectedstrings]
if False in results:
return False
else:
return True
expectedstrings = ["º", "\'", '"']
string = "21º09\'43.32\""
results = [constant in string for constant in expectedstrings]
if False in results:
print(False)
data_df_valued = data_df
data_df_valued["longitud_check"] = data_df_valued["longitud"].apply(is_field_congruent)
data_df_valued["latitud_check"] = data_df_valued["latitud"].apply(is_field_congruent)
data_df_valued["longitud_check"].value_counts()
data_df_valued["latitud_check"].value_counts()
data_df_valued[data_df_valued["latitud_check"]== False]
# Al parecer el metodo de ir por caracteres específicos no está tan chido. Lets go for a go for all the numbahs approach
import re
p = re.findall('[0-9]+',"21º09\'43.32\"")
#segunda version de congruencia. A ver si encontramos 4 componentes, veda.
def is_field_congruent(field):
p = re.findall('[0-9]+',field)
if len(p) == 4:
return True
else:
return False
data_df_valued = data_df
data_df_valued["longitud_check"] = data_df_valued["longitud"].apply(is_field_congruent)
data_df_valued["latitud_check"] = data_df_valued["latitud"].apply(is_field_congruent)
data_df_valued["longitud_check"].value_counts()
data_df_valued["latitud_check"].value_counts()
# Right on, 8 matches en False, Tenemos 7 detectados como blank. Oh lord. Veamos cual es. Acompañenme a ver esta triste historia.
data_df_valued[data_df_valued["latitud_check"] == False]
data_df_valued[data_df_valued["longitud_check"] == False]
# Ya salió el peine. en realidad tenemos 9 registros malos. Parece ser que hay un par de datos de coordenadas que no vienen bien. Se les tratará como NA y se obtendrán sus coordenadas con Google maps.
def clean_row(row):
row["latitud_nueva"], row["longitud_nueva"] = clean_coordinates(row)
return row
def clean_coordinates(row):
matches_latitude = re.findall('[0-9]+',row["latitud"])
matches_longitude = re.findall('[0-9]+',row["longitud"])
if len(matches_longitude)!=4 or len(matches_latitude)!=4:
new_latitude, new_longitude = get_geo_code(row)
else:
new_latitude = coords_converter(matches_latitude)
new_longitude = coords_converter(matches_longitude)
if new_longitude < new_latitude:
switcharoo = new_latitude
new_latitude = new_longitude
new_longitude = switcharoo
new_longitude = - new_longitude # - por estar en el oeste.
return new_latitude, new_longitude
def get_geo_code(row):
address = normalize_data(row["refugio"] + " "+ row["direccion"]+" " + row["municipio"]+ ", nayarit")
g = geocoder.google(address)
latitude, longitude = g.latlng[0], g.latlng[1]
return longitude, latitude
# Nos quedamos solamente con la parte de la operación
def coords_converter(coordinate_list):
degrees = float(coordinate_list[0])
minutes = float(coordinate_list[1])
seconds = float(coordinate_list[2]+"."+coordinate_list[3])
decimal_coords = degrees + minutes/60.0 + seconds/3600
return decimal_coords
def try_address(address):
g = geocoder.google(address)
latitude, longitude = g.latlng[0], g.latlng[1]
return latitude, longitude
def get_geo_code(row):
address = normalize_data(row["refugio"] + ", "+ row["direccion"]+", " + row["municipio"]+ ", nayarit, mexico")
try:
latitude, longitude = try_address(address)
except:
try:
address = normalize_data(row["refugio"] + ", " + row["municipio"]+ ", nayarit, mexico")
latitude, longitude = try_address(address)
except:
try:
address = normalize_data( row["direccion"]+", " + row["municipio"]+ ", nayarit, mexico")
latitude, longitude = try_address(address)
except:
address = normalize_data(row["municipio"]+ ", nayarit, mexico")
latitude, longitude = try_address(address)
print("Llené con municipio")
return(latitude, longitude)
play_data = data_df_valued[data_df_valued["longitud_check"] == False]
data_df_valued[data_df_valued["longitud_check"] == False]
play_data = play_data.apply(clean_row, axis=1)
play_data
play_data_2 = data_df
play_data_2 = play_data_2.apply(clean_row, axis=1)
play_data_2
# %matplotlib inline
play_data_2["latitud_nueva"].hist()
play_data_2["longitud_nueva"].hist()
play_data_2[play_data_2["latitud_nueva"]< -21]
play_data_2 = data_df
play_data_2 = play_data_2.apply(clean_row, axis=1)
play_data_2["longitud_nueva"].hist()
play_data_2["latitud_nueva"].hist()
play_data_2[play_data_2["latitud_nueva"] > 30]
play_data_2[play_data_2["longitud_nueva"] > -50]
play_data_3 = data_df
play_data_3 = play_data_2.apply(clean_row, axis=1)
play_data_3["longitud_nueva"].hist()
play_data_3["latitud_nueva"].hist()
| notebooks/1-0.1-rdat-exploracionexcel-LimpiezaDatos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import cv2
ajedrez = cv2.imread('tablero-ajedrez.png')
plt.imshow(ajedrez)
# BGR A B/N
ajedrez_gris = cv2.cvtColor(ajedrez, cv2.COLOR_BGR2GRAY)
plt.imshow(ajedrez_gris, cmap='gray')
# +
#cv2.goodFeaturesToTrack(matrix, FEATURE_DETECT_MAX_CORNERS, FEATURE_DETECT_QUALITY_LEVEL,
# FEATURE_DETECT_MIN_DISTANCE)
esquinas = cv2.goodFeaturesToTrack(ajedrez_gris, 100, 0.01, 4)
# -
esquinas = np.int0(esquinas)
esquinas
for i in esquinas:
x, y = i.ravel()
#circulo cv2.circle(image, center_coordinates, radius, color, thickness)
#
cv2.circle(ajedrez, (x,y), 4, color=(0,255,0), thickness=16)
plt.imshow(ajedrez)
| ClaseJueves15102020/Esquinas/Ajedrez.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# nuclio: ignore
import nuclio
# +
# %nuclio config kind="nuclio:serving"
# %nuclio env MODEL_CLASS=ChurnModel
# %nuclio config spec.build.baseImage = "mlrun/ml-models"
# -
# ## Function Code
# +
import os
import json
import numpy as np
from cloudpickle import load
### Model Serving Class
import mlrun
class ChurnModel(mlrun.runtimes.MLModelServer):
def load(self):
"""
load multiple models in nested folders, churn model only
"""
clf_model_file, extra_data = self.get_model(".pkl")
self.model = load(open(str(clf_model_file), "rb"))
if "cox" in extra_data.keys():
cox_model_file = extra_data["cox"]
self.cox_model = load(open(str(cox_model_file), "rb"))
if "cox/km" in extra_data.keys():
km_model_file = extra_data["cox/km"]
self.km_model = load(open(str(km_model_file), "rb"))
return
def predict(self, body):
try:
# we have potentially 3 models to work with:
#if hasattr(self, "cox_model") and hasattr(self, "km_model"):
# hack for now, just predict using one:
feats = np.asarray(body["instances"], dtype=np.float32).reshape(-1, 23)
result = self.model.predict(feats, validate_features=False)
return result.tolist()
#else:
# raise Exception("models not found")
except Exception as e:
raise Exception("Failed to predict %s" % e)
# +
# nuclio: end-code
# -
# ### mlconfig
from mlrun import mlconf
import os
mlconf.dbpath = mlconf.dbpath or "http://mlrun-api:8080"
mlconf.artifact_path = mlconf.artifact_path or f"{os.environ['HOME']}/artifacts"
# <a id="test-locally"></a>
# ## Test the function locally
# +
model_dir = os.path.join(mlconf.artifact_path, "churn", "models")
my_server = ChurnModel("my-model", model_dir=model_dir)
my_server.load()
# -
DATA_URL = f"https://raw.githubusercontent.com/yjb-ds/testdata/master/data/churn-tests.csv"
import pandas as pd
xtest = pd.read_csv(DATA_URL)
# We can use the `.predict(body)` method to test the model.
# +
import json, numpy as np
# this should fail if the churn model hasn't been saved properly
preds = my_server.predict({"instances":xtest.values[:10,:-1].tolist()})
# -
print("predicted class:", preds)
# <a id="deploy"></a>
# ### **deploy our serving class using as a serverless function**
# in the following section we create a new model serving function which wraps our class , and specify model and other resources.
#
# the `models` dict store model names and the assosiated model **dir** URL (the URL can start with `S3://` and other blob store options), the faster way is to use a shared file volume, we use `.apply(mount_v3io())` to attach a v3io (iguazio data fabric) volume to our function. By default v3io will mount the current user home into the `\User` function path.
#
# **verify the model dir does contain a valid `model.bst` file**
from mlrun import new_model_server, mount_v3io
import requests
# +
fn = new_model_server("churn-test",
model_class="ChurnModel",
models={"churn_server_v1": f"{model_dir}"})
fn.spec.description = "churn classification and predictor"
fn.metadata.categories = ["serving", "ml"]
fn.metadata.labels = {"author": "yashab", "framework": "churn"}
fn.export("function.yaml")
# -
# ## tests
if "V3IO_HOME" in list(os.environ):
from mlrun import mount_v3io
fn.apply(mount_v3io())
else:
# is you set up mlrun using the instructions at
# https://github.com/mlrun/mlrun/blob/master/hack/local/README.md
from mlrun.platforms import mount_pvc
fn.apply(mount_pvc("nfsvol", "nfsvol", "/home/jovyan/data"))
#addr = fn.deploy(dashboard="http://172.17.0.66:8070", project="churn-project")
addr = fn.deploy(project="churn-project")
addr
# <a id="test-model-server"></a>
# ### **test our model server using HTTP request**
#
#
# We invoke our model serving function using test data, the data vector is specified in the `instances` attribute.
# KFServing protocol event
event_data = {"instances": xtest.values[:10,:-1].tolist()}
# +
import json
resp = requests.put(addr + "/churn_server_v1/predict", json=json.dumps(event_data))
# mlutils function for this?
tl = resp.text.replace("[","").replace("]","").split(",")
assert preds == [int(i) for i in np.asarray(tl)]
# -
# **[back to top](#top)**
| churn_server/churn_server.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
df=pd.read_csv("FitBit data.csv")
df.head()
# !pip install pandas_profiling
import pandas_profiling
df.shape
df.isnull().sum()
df.head(10)
df1=df.copy()
df1['ActivityDate'].nunique()
pd.DatetimeIndex(df['ActivityDate']).year
df1['year']=pd.DatetimeIndex(df1['ActivityDate']).year
df1['month']=pd.DatetimeIndex(df1['ActivityDate']).month
df1['day']=pd.DatetimeIndex(df1['ActivityDate']).day
df1.info()
df1=df1.drop(['TrackerDistance'],axis=1)
df1
df1.columns
df1.Calories
# +
plt.figure(figsize=(15,8))
# Usual boxplot
ax = sns.boxplot(x='day', y='Calories', data=df1)
# Add jitter with the swarmplot function.
ax = sns.swarmplot(x='day', y='Calories', data=df1, color="grey")
ax.set_title('Box plot of Calories with Jitter bu day of the month')
# -
pd.to_datetime(df1['ActivityDate']).dt.week
pd.DatetimeIndex(df1['ActivityDate']).month
df1['Week']=pd.to_datetime(df1['ActivityDate']).dt.week
df1['Year']=pd.to_datetime(df1['ActivityDate']).dt.year
df1.Year
df1.head(10)
df1.ActivityDate.dtype
df1['ActivityDate']=pd.to_datetime(df['ActivityDate'])
df1.ActivityDate.dtype
df1['weekday'] = df1['ActivityDate'].dt.weekday
df1['weekday']
df1.head()
plt.figure(figsize=(15,7))
ax=sns.barplot(x='weekday',y='Calories',data=df1)
ax.set_title('Barplot of calories by the day of the week')
plt.figure(figsize=(15,8))
ax=sns.scatterplot(x='Calories',y='SedentaryMinutes',data=df1)
ax.set_title('Scatter plot of Calories')
# +
# figure size
plt.figure(figsize=(15,8))
# Simple scatterplot
ax = sns.scatterplot(x='Calories', y='LightlyActiveMinutes', data=df1)
ax.set_title('Scatterplot of calories and intense_activities')
# +
# figure size
plt.figure(figsize=(15,8))
# Simple scatterplot between calories burnt in the moderately active minutes
ax = sns.scatterplot(x='Calories', y='FairlyActiveMinutes', data=df1)
ax.set_title('Scatterplot of calories vs Fairly Active Minutes')
# -
plt.figure(figsize=(15,7))
ax=sns.scatterplot(x='Calories',y='VeryActiveMinutes',data=df1)
ax.set_title('Scatter plot')
# +
col_select=['Calories','VeryActiveMinutes','FairlyActiveMinutes','LightlyActiveMinutes','SedentaryMinutes']
wide_df=df1[col_select]
plt.figure(figsize=(15,8))
ax=sns.lineplot(data=wide_df)
ax.set_title('Un-normalized value of calories and different activities based on activity minutes')
# -
plt.figure(figsize=(15,8))
ax=sns.scatterplot(x='Calories',y='TotalDistance',data=df1)
ax.set_title('Scatterplot of calories and intense_activities')
# +
## plot the raw values
rol_select = ['TotalDistance','LoggedActivitiesDistance','VeryActiveDistance','ModeratelyActiveDistance', 'LightActiveDistance']
wide_df1 = df1[rol_select]
# figure size
plt.figure(figsize=(15,8))
# timeseries plot using lineplot
ax = sns.lineplot(data=wide_df1)
ax.set_title('Un-normalized value of calories and different activities based on distance')
# -
| Fitbit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Lambda School Data Science
#
# *Unit 2, Sprint 1, Module 4*
#
# ---
# + [markdown] colab_type="text" id="7IXUfiQ2UKj6"
# # Logistic Regression
#
#
# ## Assignment 🌯
#
# You'll use a [**dataset of 400+ burrito reviews**](https://srcole.github.io/100burritos/). How accurately can you predict whether a burrito is rated 'Great'?
#
# > We have developed a 10-dimensional system for rating the burritos in San Diego. ... Generate models for what makes a burrito great and investigate correlations in its dimensions.
#
# - [ ] Do train/validate/test split. Train on reviews from 2016 & earlier. Validate on 2017. Test on 2018 & later.
# - [ ] Begin with baselines for classification.
# - [ ] Use scikit-learn for logistic regression.
# - [ ] Get your model's validation accuracy. (Multiple times if you try multiple iterations.)
# - [ ] Get your model's test accuracy. (One time, at the end.)
# - [ ] Commit your notebook to your fork of the GitHub repo.
#
#
# ## Stretch Goals
#
# - [ ] Add your own stretch goal(s) !
# - [ ] Make exploratory visualizations.
# - [ ] Do one-hot encoding.
# - [ ] Do [feature scaling](https://scikit-learn.org/stable/modules/preprocessing.html).
# - [ ] Get and plot your coefficients.
# - [ ] Try [scikit-learn pipelines](https://scikit-learn.org/stable/modules/compose.html).
# + colab={} colab_type="code" id="o9eSnDYhUGD7"
# %%capture
import sys
# If you're on Colab:
if 'google.colab' in sys.modules:
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Linear-Models/master/data/'
# !pip install category_encoders==2.*
# If you're working locally:
else:
DATA_PATH = '../data/'
# -
# Load data downloaded from https://srcole.github.io/100burritos/
import pandas as pd
pd.set_option('display.max_columns', 999)
df = pd.read_csv(DATA_PATH+'burritos/burritos.csv')
# Derive binary classification target:
# We define a 'Great' burrito as having an
# overall rating of 4 or higher, on a 5 point scale.
# Drop unrated burritos.
df = df.dropna(subset=['overall'])
df['Great'] = df['overall'] >= 4
# +
# Clean/combine the Burrito categories
df['Burrito'] = df['Burrito'].str.lower()
california = df['Burrito'].str.contains('california')
asada = df['Burrito'].str.contains('asada')
surf = df['Burrito'].str.contains('surf')
carnitas = df['Burrito'].str.contains('carnitas')
df.loc[california, 'Burrito'] = 'California'
df.loc[asada, 'Burrito'] = 'Asada'
df.loc[surf, 'Burrito'] = 'Surf & Turf'
df.loc[carnitas, 'Burrito'] = 'Carnitas'
df.loc[~california & ~asada & ~surf & ~carnitas, 'Burrito'] = 'Other'
# -
# Drop some high cardinality categoricals
df = df.drop(columns=['Notes', 'Location', 'Reviewer', 'Address', 'URL', 'Neighborhood'])
# Drop some columns to prevent "leakage"
df = df.drop(columns=['Rec', 'overall'])
df['Date'] = pd.to_datetime(df['Date'])
from IPython.display import display
display(df.columns)
display(df)
train = df[df['Date'].apply(lambda x : x.year < 2017)]
val = df[df['Date'].apply(lambda x : x.year == 2017)]
test = df[df['Date'].apply(lambda x : x.year > 2017)]
df.shape, train.shape, val.shape, test.shape
for c in ['Burrito', 'Date', 'Chips', 'Unreliable', 'NonSD', 'Beef', 'Pico',
'Guac', 'Cheese', 'Fries', 'Sour cream', 'Pork', 'Chicken', 'Shrimp',
'Fish', 'Rice', 'Beans', 'Lettuce', 'Tomato', 'Bell peper', 'Carrots',
'Cabbage', 'Sauce', 'Salsa.1', 'Cilantro', 'Onion', 'Taquito',
'Pineapple', 'Ham', 'Chile relleno', 'Nopales', 'Lobster', 'Egg',
'Mushroom', 'Bacon', 'Sushi', 'Avocado', 'Corn', 'Zucchini', 'Great']:
display(train[c].value_counts(dropna=False))
import numpy as np
mp = {'x': True, 'X': True, np.nan: False, 'Yes': True, 'No': False}
for c in train:
train[c] = train[c].map(lambda x : mp[x] if x in mp else x)
train
for c in train:
train[c].sum()
train = train.drop(['Mass (g)', 'Density (g/mL)', 'Queso'], axis=1)
for c in train:
display(train[c].value_counts())
| module4-logistic-regression/LS_DS_214_assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
from tensorflow.keras.datasets import mnist
import numpy as np
import h5py
from sklearn.model_selection import train_test_split
from os import walk
# ## Acquire The Data
batch_size = 128
img_rows, img_cols = 28, 28 # image dims
#load npy arrays
data_path = "data_files/" # folder for image files
for (dirpath, dirnames, filenames) in walk(data_path):
pass # file names accumulate in list 'filenames'
print(filenames)
num_images = 1000000 ### was 100000, reduce this number if memory issues.
num_files = len(filenames) # *** we have 10 files ***
images_per_category = num_images//num_files
seed = np.random.randint(1, 10e7)
i=0
print(images_per_category)
for file in filenames:
file_path = data_path + file
x = np.load(file_path)
x = x.astype('float32') ##normalise images
x /= 255.0
y = [i] * len(x) # create numeric label for this image
x = x[:images_per_category] # get our sample of images
y = y[:images_per_category] # get our sample of labels
if i == 0:
x_all = x
y_all = y
else:
x_all = np.concatenate((x,x_all), axis=0)
y_all = np.concatenate((y,y_all), axis=0)
i += 1
#split data arrays into train and test segments
x_train, x_test, y_train, y_test = train_test_split(x_all, y_all, test_size=0.2, random_state=42)
x_train = x_train.reshape(x_train.shape[0], img_rows, img_cols, 1)
x_test = x_test.reshape(x_test.shape[0], img_rows, img_cols, 1)
input_shape = (img_rows, img_cols, 1)
y_train = tf.keras.utils.to_categorical(y_train, num_files)
y_test = tf.keras.utils.to_categorical(y_test, num_files)
# +
print('x_train shape:', x_train.shape)
print(x_train.shape[0], 'train samples')
print(x_test.shape[0], 'test samples')
x_train, x_valid, y_train, y_valid = train_test_split(x_train, y_train, test_size=0.1, random_state=42)
# -
# ## Create the model
# +
model = tf.keras.Sequential()
model.add(tf.keras.layers.Conv2D(32, kernel_size=(3, 3), activation='relu', input_shape=input_shape))
model.add(tf.keras.layers.MaxPooling2D(pool_size=(2, 2)))
model.add(tf.keras.layers.Dropout(0.25))
model.add(tf.keras.layers.Conv2D(64, (3, 3), activation='relu'))
model.add(tf.keras.layers.MaxPooling2D(pool_size=(2, 2)))
model.add(tf.keras.layers.Dropout(0.25))
model.add(tf.keras.layers.Flatten())
model.add(tf.keras.layers.Dense(128, activation='relu'))
model.add(tf.keras.layers.Dropout(0.5))
model.add(tf.keras.layers.Dense(num_files, activation='softmax'))
print("Compiling...........")
# -
model.compile(loss=tf.keras.losses.categorical_crossentropy,
optimizer=tf.keras.optimizers.Adadelta(),
metrics=['accuracy'])
# ## Train the model
epochs=1 # for testing, for training use 25
callbacks=[tf.keras.callbacks.TensorBoard(log_dir = "./tb_log_dir", histogram_freq = 0)]
model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
callbacks=callbacks,
verbose=1,
validation_data=(x_valid, y_valid))
# +
score = model.evaluate(x_test, y_test, verbose=1)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
# -
# ## Test The Model
# +
#_test
import os
labels = [os.path.splitext(file)[0] for file in filenames]
print(labels)
print("\nFor each pair in the following, the first label is predicted, second is actual\n")
for i in range(20):
t = np.random.randint(len(x_test) )
x1= x_test[t]
x1 = x1.reshape(1,28,28,1)
p = model.predict(x1)
print("-------------------------")
print(labels[np.argmax(p)])
print(labels[np.argmax(y_test[t])])
print("-------------------------")
# -
# ## Save, Reload and Retest the Model
model.save("./QDrawModel.h5")
del model
from tensorflow.keras.models import load_model
import numpy as np
model = load_model('./QDrawModel.h5')
model.summary()
print("For each pair, first is predicted, second is actual")
for i in range(20):
t = np.random.randint(len(x_test))
x1= x_test[t]
x1 = x1.reshape(1,28,28,1)
p = model.predict(x1)
print("-------------------------")
print(labels[np.argmax(p)])
print(labels[np.argmax(y_test[t])])
print("-------------------------")
| Chapter06/CHapter6_QDraw_TF2_alpha.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from __future__ import print_function
from __future__ import division
import numpy as np
import torch
import numpy
from numpy import cov
from numpy import trace
from numpy import iscomplexobj
from numpy import asarray
from numpy.random import randint
from scipy.linalg import sqrtm
from skimage.transform import resize
import torch
import torch.nn as nn
import torch.optim as optim
import torchvision
from torchvision import datasets, models, transforms
import matplotlib.pyplot as plt
import time
import os
import copy
os.chdir('/Users/xxx/Documents/GANs_Research/my_imps/research_models/v3/evaluation')
# <h1>cvpr Models</h1>
# FID scores for test results generated from four different favtGAN architectures, where all images were rescaled, aligned, and properly formatted.
# <h2>Eurecom</h2>
#
# - EA_sensor_OG
# - EA_sensor_V3
# - EA_sensor_V4
# - EA_sensor_V5
#
#
#
# - EAI_sensor_OG
# - EAI_sensor_V3
# - EAI_sensor_V4
# - EAI_sensor_V5
#EA_sensor_08
# ! python -m pytorch_fid Eurecom/EA_sensor_08/real_B Eurecom/EA_sensor_08/fake_B
#EA_sensor_V4
# ! python -m pytorch_fid Eurecom/EA_sensor_V4/real_B Eurecom/EA_sensor_V4/fake_B
#EAI_sensor_V4
# ! python -m pytorch_fid Eurecom/EAI_sensor_V4/real_B Eurecom/EAI_sensor_V4/fake_B
#EAI_sensor_OG
# ! python -m pytorch_fid Eurecom/EAI_sensor_OG/real_B Eurecom/EAI_sensor_OG/fake_B
#EAI_sensor_V3
# ! python -m pytorch_fid Eurecom/EAI_sensor_V3/real_B Eurecom/EAI_sensor_V3/fake_B
# Eurecom pix2pix
# ! python -m pytorch_fid Eurecom/eurecom_pix2pix/real_B Eurecom/eurecom_pix2pix/fake_B
#EA_sensor_OG
# ! python -m pytorch_fid Eurecom/EA_sensor_OG/real_B Eurecom/EA_sensor_OG/fake_B
#EI_sensor_OG
# ! python -m pytorch_fid Eurecom/EI_sensor_OG/real_B Eurecom/EI_sensor_OG/fake_B
#EI_sensor_V3
# ! python -m pytorch_fid Eurecom/EI_sensor_V3/real_B Eurecom/EI_sensor_V3/fake_B
#EI_sensor_V4
# ! python -m pytorch_fid Eurecom/EI_sensor_V4/real_B Eurecom/EI_sensor_V4/fake_B
#EA
# ! python -m pytorch_fid Eurecom/EA_pix2pix/real_B Eurecom/EA_pix2pix/fake_B
#EAI
# ! python -m pytorch_fid Eurecom/EAI_pix2pix/real_B Eurecom/EAI_pix2pix/fake_B
# +
#EI pix2pix
# ! python -m pytorch_fid Eurecom/EI_pix2pix/real_B Eurecom/EI_pix2pix/fake_B
# -
#EI_sensor_V5
# ! python -m pytorch_fid Eurecom/EI_sensor_V5/real_B Eurecom/EI_sensor_V5/fake_B
#EAI sensor V5
# ! python -m pytorch_fid Eurecom/EAI_sensor_V5/real_B Eurecom/EAI_sensor_V5/fake_B
#Eio SENSOR V4
# ! python -m pytorch_fid Eurecom/EIO_sensor_V4/real_B Eurecom/EIO_sensor_V4/fake_B
# <h2>Iris</h2>
#EAI_sensor_V4
# ! python -m pytorch_fid Iris/EAI_sensor_V4/real_B Iris/EAI_sensor_V4/fake_B
#EAI_sensor_OG
# ! python -m pytorch_fid Iris/EAI_sensor_OG/real_B Iris/EAI_sensor_OG/fake_B
#EAI_sensor_V3
# ! python -m pytorch_fid Iris/EAI_sensor_V3/real_B Iris/EAI_sensor_V3/fake_B
#Iris pix2pix
# ! python -m pytorch_fid Iris/iris_pix2pix/real_B Iris/iris_pix2pix/fake_B
#EI_sensor_OG
# ! python -m pytorch_fid Iris/EI_sensor_OG/real_B Iris/EI_sensor_OG/fake_B
#EI_sensor_V3
# ! python -m pytorch_fid Iris/EI_sensor_V3/real_B Iris/EI_sensor_V3/fake_B
#EI_sensor_V4
# ! python -m pytorch_fid Iris/EI_sensor_V4/real_B Iris/EI_sensor_V4/fake_B
#EAI
# ! python -m pytorch_fid Iris/EAI_pix2pix/real_B Iris/EAI_pix2pix/fake_B
#EI pix2pix
# ! python -m pytorch_fid Iris/EI_pix2pix/real_B Iris/EI_pix2pix/fake_B
#EI sensor V5
# ! python -m pytorch_fid Iris/EI_sensor_V5/real_B Iris/EI_sensor_V5/fake_B
#EAI sensor V5
# ! python -m pytorch_fid Iris/EAI_sensor_V5/real_B Iris/EAI_sensor_V5/fake_B
#EIO SENSOR V4
# ! python -m pytorch_fid Iris/EIO_sensor_V4/real_B Iris/EIO_sensor_V4/fake_B
#IO sensor V4
# ! python -m pytorch_fid Iris/IO_sensor_V4/real_B Iris/IO_sensor_V4/fake_B
| quant_eval/notebooks/fid_cvpr.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: polarice
# language: python
# name: polarice
# ---
# # Examples
#
# Here we provide a few examples of tweets with certain morals.
# Due to privacy and ToS reasons, we cannot release real example, but created artificial one's inspired by tweets (i.e., altered tweets) in the dataset.
# %run frame_axis.py
fs_en = FrameSystem.load("moral.pkl")
fs_de = FrameSystem.load("moral_de.pkl")
fs_en.frame_axes, fs_de.frame_axes
from gensim.models import KeyedVectors
model_en = KeyedVectors.load_word2vec_format("cc.en.300.vec.gz", binary=False, limit=100_000)
model_de = KeyedVectors.load_word2vec_format("cc.de.300.vec.gz", binary=False, limit=100_000)
examples_de = [
"es benötigt eine sanierung des corona gesetzes",
"diese #CoronaWarnApp ist sinnlos",
"sorgenfrei durch corona",
]
for example_de in examples_de:
biases = []
for axis_name in fs_de.frame_axes.keys():
bias = compute_bias(example_de, fs_de.frame_axes[axis_name].axis, model_de)
biases.append(bias)
print(f"{example_de}: {biases}")
examples_en = [
"it is a priority to increase the number of healthcare providers",
"the stories of victims of terrible crimes should not be forgotten",
"your statement is ignorant and despicable low",
"this comment is spiteful",
]
for example_en in examples_en:
biases = []
for axis_name in fs_en.frame_axes.keys():
bias = compute_bias(example_en, fs_en.frame_axes[axis_name].axis, model_en)
biases.append(bias)
print(f"{example_en}: {biases}")
# Observation for German:
#
# The first sentence is an example of high authority due to focus on law (although it wants to change it).
# The second sentence does not see the benefit of the corona app, and thus might outline why caring for others is less important.
# The third lies in between and is somewhat ambigous.
#
# Observation for English:
#
# The first sentence is suggests that not enough care is given.
# The second sentence outlines the unfairness of crime victims, also emphasizes the degradation.
# The third and fourth refer to other statements and express their disgust towards them.
# ### Remarks
#
# This is just a showcase, for the real implementation (e.g., including preprocessing) refer to the other notebooks.
#
# Note that:
# - biases are very small unscale
# - some biases seemingly point in the wrong direction, as whole topics themselves can be biased (i.e., baseline bias)
# - thus only the combination of bias and intensity is beneficial
# - still, here we only report biases, as intensities would require a reference corpus
# - while we see that the approach works good for certain examples, it does less so on others, that is why it is important to consider the corpus as a whole
# - also, as reported in the paper, for some words it is not clear to which pole they belong (i.e., words such as wounds)
#
# Although, none of those tweets is real, all are an adapted form of real tweets (thus not completely artificial).
# + active=""
# # Code to get the extreme bias sentences in other scripts
#
# for bias in ["care_bias", "fair_bias", "auth_bias", "sanc_bias", "loya_bias"]:
# min_text = trans_df.iloc[trans_df[bias].argmin()]["full_text"]
# min_affil = trans_df.iloc[trans_df[bias].argmin()]["party"]
# print(f"Min {bias} by {min_affil}: {min_text}")
#
# max_text = trans_df.iloc[trans_df[bias].argmax()]["full_text"]
# max_affil = trans_df.iloc[trans_df[bias].argmax()]["party"]
# print(f"Max {bias} by {max_affil}: {max_text}")
#
# for bias in ["care_bias", "fair_bias", "auth_bias", "sanc_bias", "loya_bias"]:
# for party in ["D", "R"]:
# party_df = trans_df[trans_df["party"] == party]
#
# min_text = party_df.iloc[party_df[bias].argmin()]["full_text"]
# min_affil = party_df.iloc[party_df[bias].argmin()]["party"]
# print(f"Min {bias} by {party}: {min_text}")
#
# max_text = party_df.iloc[party_df[bias].argmax()]["full_text"]
# max_affil = party_df.iloc[party_df[bias].argmax()]["party"]
# print(f"Max {bias} by {party}: {max_text}")
# -
| examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# # Exploring Foreign Languages
#
# So far, we have been learning about general ways to explore texts through manipulating strings and regular expressions. Today, we will be focusing on what we can do when texts are in languages other than English. This will just be an introduction to some of the many different modules that can be used for these tasks. The goal is to learn some tools, including Polyglot and translation, that can be jumping off points to see what you may or may not need going forward.
#
# ### Lesson Outline:
# - Q&A about what we've gone over so far
# - Examples (with Sara's data)
# - Practice!
# ## Installations
# Uncomment and run the cell below!
# +
# #!pip install translation
# #!pip install py-translate
# #!pip install morfessor
# #!pip install polyglot
# #!pip install pycld2
# #!brew install intltool icu4c gettext
# #!brew link icu4c gettext --force
# #!CFLAGS=-I/usr/local/opt/icu4c/include LDFLAGS=-L/usr/local/opt/icu4c/lib pip3 install pyicu
# -
# ## Importing Text
import codecs
with codecs.open('Skyggebilleder af en Reise til Harzen.txt', 'r', encoding='utf-8', errors='ignore') as f:
read_text = f.read()
read_text
# pulling out a subsection of text for our examples
text_snippet = read_text[20000:23000]
# ## Translating Text
#
# There are many different ways that you could go about translating text within Python, but one of the easiest is the package `translation`. `translation` makes use of existing online translators. The module used to include a method for Google Translate, but the site no longer allows easy access. Bing is probably the most useful method for it.
#
# **Pros:**
# * Easy to set up
# * Runs quickly
#
# **Cons:**
# * Not always accurate
# * Internet connection needed
# * Language limitations
#
# The documentation (or lack there of): https://pypi.python.org/pypi/translation
import translation
translation.bing(text_snippet, dst = 'en')
# Other alternatives for translating your text include:
# * `py-translate`
# * Makes use of Google Translate
# * Often return errors / gets blocked
# * Can be used from the command line
# * Documentation: https://pypi.python.org/pypi/py-translate
#
#
# * API calls to Google Translate
# * Takes a little more set-up
# * Can be customized a little bit more
# * Can translate a LOT of text
# +
# using py-translate
from translate import translator
# calling tranlator function, telling it that the
translator('da', 'en',text_snippet[:200])
# -
# ## Polyglot
#
# Polyglot is "a natural language pipeline that supports massive multilingual applications," in other words, it does a lot of stuff. It is a sort of one-stop-shop for many different functions that you may want to apply to you text, and supports many different languages. We are going to run through some of its functionalities.
#
# Docs: http://polyglot.readthedocs.io/en/latest/
# #### Language Detection
# +
from polyglot.detect import Detector
# create a detector object that contains read_text
# and assigning it to DETECTED
detected = Detector(read_text)
# the .language method will return the language the most of
# the text is made up of and the system is confident about
print(detected.language)
# -
# sometimes there will be multiple languages within
# the text, and you will want to see all of them
for language in detected.languages:
print(language)
# if you try to pass in a string that is too short
# for the system to get a good read on, it will throw
# an error, alerting you to this fact
Detector("4")
# we can override that with the optional argument 'quiet=True'
print(Detector("4", quiet=True))
# here are all of the languages supported for language detection
from polyglot.utils import pretty_list
print(pretty_list(Detector.supported_languages()))
# #### Tokenization
#
# Similar to what we saw with NLTK, Polyglot can break our text up into words and sentences. Polyglot has the advantage of spanning multiple languages, and thus is more likely to identify proper breakpoint in languages other than English.
# +
from polyglot.text import Text
# creating a Text object that analyzes our text_snippet
text = Text(text_snippet)
# +
# Text also has a language instance variable
print(text.language)
# here, we are looking at text_snippet tokenized into words
text.words
# -
# now we are looking at text_snippet broken down into sentences
text.sentences
# #### Side Notes: Important Package Information
#
# Not all of the packages are downloaded for all functionalities for all languages in Polyglot. Instead of forcing you to download a lot of files in the beginning, the creators decided that it would be better for language extensions to be downloaded on an 'as-necessary' basis. You will occassionaly be told that you're lacking a package, and you will need to download it. You can either do that with the built-in downloader, or from the command line.
# staying within python
from polyglot.downloader import downloader
downloader.download("embeddings2.en")
# alternate command line method
# !polyglot download embeddings2.da pos2.da
# Also, if you're working with a language and want to know what Polyglot lets you do with a language, it provides a `supported_tasks` method.
# tasks available for english
downloader.supported_tasks(lang="en")
# tasks available for danish
downloader.supported_tasks(lang="da")
# #### Part of Speech Tagging
#
# Polyglot supports POS tagging for several languages.
# languages that polyglot supports for part of speech tagging
print(downloader.supported_languages_table("pos2"))
text.pos_tags
# #### Named Entity Recognition
#
# Polyglot can tag names and groups them into three main categories:
# * Locations (Tag: I-LOC): cities, countries, regions, continents, neighborhoods, administrative divisions ...
# * Organizations (Tag: I-ORG): sports teams, newspapers, banks, universities, schools, non-profits, companies, ...
# * Persons (Tag: I-PER): politicians, scientists, artists, atheletes ...
# languages that polyglot supports for part of speech tagging
print(downloader.supported_languages_table("ner2", 3))
# #!polyglot download ner2.da
text.entities
# #### Other Features of Polyglot
# * Nearest Neighbors -- http://polyglot.readthedocs.io/en/latest/Embeddings.html
# * Morpheme Generation -- http://polyglot.readthedocs.io/en/latest/MorphologicalAnalysis.html
# * Sentiment Analysis -- http://polyglot.readthedocs.io/en/latest/Sentiment.html
# * Transliteration -- http://polyglot.readthedocs.io/en/latest/Transliteration.html
# ## Code Summary:
#
# #### Translation:
# * `translation.bing(your_string, dst = 'en')`
#
# #### Polyglot:
# * `<Detector>.language`
# * `<Detector>.languages`
# * `<Text>.language`
# * `<Text>.words`
# * `<Text>.sentences`
# * `<Text>.pos_tags`
# * `<Text>.entities`
# ### Extra
# importing some more packages
from datascience import *
# %matplotlib inline
import seaborn as sns
# analyzing our text with a Polyglot Text object
whole_text = Text(read_text)
# the language of our text
print(whole_text.language)
# getting the part of speech tags for our corpus
print(whole_text.pos_tags)
words_and_poss = list(whole_text.pos_tags)
# putting those word / part of speech pairs into a table
wrd = Table(['Word', 'Part of Speech']).with_rows(words_and_poss)
# grouping those by part of speech to get the most commonly occuring parts of speech
df = wrd.group('Part of Speech').sort('count', descending=True).to_df()
df
# plotting the counts for each part of speech using seaborn
sns.barplot(x='Part of Speech', y='count', data=df)
# getting the most popular word for each part of speech type
wrd_counts = wrd.group('Word').join('Word', wrd).sort('count', descending=True)
wrd_counts.group(2, lambda x: x.item(0)).show(16)
# thats not very informative, so lets pull out the stop words
# using a list from http://snowball.tartarus.org/algorithms/danish/stop.txt
danish_stop_words = """og,
i,
jeg,
det,
at,
en,
den,
til,
er,
som,
på,
de,
med,
han,
af,
for,
ikke,
der,
var,
mig,
sig,
men,
et,
har,
om,
vi,
min,
havde,
ham,
hun,
nu,
over,
da,
fra,
du,
ud,
sin,
dem,
os,
op,
man,
hans,
hvor,
eller,
hvad,
skal,
selv,
her,
alle,
vil,
blev,
kunne,
ind,
når,
være,
dog,
noget,
ville,
jo,
deres,
efter,
ned,
skulle,
denne,
end,
dette,
mit,
også,
under,
have,
dig,
anden,
hende,
mine,
alt,
meget,
sit,
sine,
vor,
mod,
disse,
hvis,
din,
nogle,
hos,
blive,
mange,
ad,
bliver,
hendes,
været,
thi,
jer,
sådan"""
splt = danish_stop_words.split(',\n')
print(splt)
# determining which rows we need to change
not_in_stop_words = [x not in danish_stop_words for x in wrd_counts['Word']]
# most common words for each part of speech no longer including the stop words
wrd_counts.where(not_in_stop_words).group(2, lambda x: x.item(0)).show(16)
# retrieving all of the named entities that Polyglot detected
ner = str(whole_text.entities).split('I-')[1:]
ner[:5]
# splitting up the type and the name
split_type = [x.split('([') for x in ner]
split_type[:5]
# making a table out of that
entities = Table(['Type', 'Name']).with_rows(split_type)
entities
# how many of each type of entity there are
entities.group('Type')
# finding the most commonly occuring entities
entities.group('Name').sort('count', descending=True)
# possibly the most common names of people
entities.where('Type', 'PER').group('Name').sort('count', True)
| foreign_languages/Sara_danish.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using `salter` to examine the transit residuals
# +
# %matplotlib inline
# %config InlineBackend.figure_format = "retina"
from salter import LightCurve
import matplotlib.pyplot as plt
# +
# Specify a KIC number to get the light curve
# Here are some numbers to try: 4157325, 9651668, 9705459, 10386922
kic_number = 10815677#9651668
whole_lc = LightCurve.from_hdf5(kic_number)
# Plot the light curve (raw SAP flux)
whole_lc.plot()
# -
# Here's how you can quickly see what the transit parameters are:
for attr in dir(whole_lc.params):
if not attr.startswith('_'):
print("{0}: {1}".format(attr, getattr(whole_lc.params, attr)))
# Mask the out-of-transit portions of the light curve, normalize each transit with the "subtract-add-divide" method.
# +
from salter import subtract_add_divide
extra_oot_time = 2.0 # [durations]; Extra transit durations to keep before ingress/after egress
# Mask out-of-transit portions of light curves, chop into individual transits
near_transit = LightCurve(**whole_lc.mask_out_of_transit(oot_duration_fraction=extra_oot_time))
transits = near_transit.get_transit_light_curves()
# Normalize all transits with the subtract-add-divide method,
# using a second order polynomial
subtract_add_divide(whole_lc, transits)
# -
10815677
whole_lc.params.duration
# Plot the residuals from the transit model given the parameters from the KOI catalog.
# +
from salter import concatenate_transit_light_curves
normed_transits = concatenate_transit_light_curves(transits)
plt.plot(normed_transits.phases(),
normed_transits.fluxes - normed_transits.transit_model(),
'k.', alpha=0.3)
plt.xlabel('Orbital phase')
plt.ylabel('Residuals')
# -
# Solve for better $R_p/R_\star$, $u_1$, $u_2$ parameters:
# +
from salter import concatenate_transit_light_curves
normed_transits = concatenate_transit_light_curves(transits)
# Solve for better parameters for: Rp/Rs, u1, u2
normed_transits.fit_lc_3param()
# -
# Store residuals in a `Residuals` object, which has handy methods for statistical analysis.
# +
from salter import Residuals
r = Residuals(normed_transits, normed_transits.params, buffer_duration=0.3)
r.plot()
# -
# ### Out-of-transit: before vs after transit
# +
# Two sample KS test: are the distributions of the two samples the same?
print(r.ks(['out_of_transit', 'before_midtransit'],
['out_of_transit', 'after_midtransit']))
# k-sample Anderson test: are the distributions of the two samples the same?
print(r.anderson(['out_of_transit', 'before_midtransit'],
['out_of_transit', 'after_midtransit']))
# Independent sample T-test: are the means the two samples the same?
print(r.ttest(['out_of_transit', 'before_midtransit'],
['out_of_transit', 'after_midtransit']))
# -
# ### In-transit: before vs after mid-transit
# +
# Two sample KS test: are the distributions of the two samples the same?
print(r.ks(['in_transit', 'before_midtransit'],
['in_transit', 'after_midtransit']))
# k-sample Anderson test: are the distributions of the two samples the same?
print(r.anderson(['in_transit', 'before_midtransit'],
['in_transit', 'after_midtransit']))
# Independent sample T-test: are the means the two samples the same?
print(r.ttest(['in_transit', 'before_midtransit'],
['in_transit', 'after_midtransit']))
# -
# ### In-transit vs out-of-transit
# +
# Two sample KS test: are the distributions of the two samples the same?
print(r.ks('in_transit', 'out_of_transit'))
# k-sample Anderson test: are the distributions of the two samples the same?
print(r.anderson('in_transit', 'out_of_transit'))
# Independent sample T-test: are the means the two samples the same?
print(r.ttest('in_transit', 'out_of_transit'))
# -
| show_lc.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="SNkZODYR-AMY"
# ## Importing Important Libraries
# + colab={"base_uri": "https://localhost:8080/"} id="luoWWKmxuUNZ" outputId="25310c35-c803-4a7a-a818-a3747cafd815"
# !pip install ktrain
# + id="fva9JlZ_8z4r"
import numpy as np
import pandas as pd
import ktrain
from ktrain import text
import tensorflow as tf
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="i_7x3uum_uOR" outputId="d753ca8b-b554-4194-9e0f-de3f5ce73145"
#Let's check version of TensorFlow, so that when later we will reload the model, we can use same version of TensorFlow:
tf.version.VERSION
# + [markdown] id="-eVUFvXNAKDW"
# ## Reading The Data and preparing our train and test data sets
# + id="7pbqpj9XADLo"
train = pd.read_csv('/content/train (1).txt', header =None, sep =';', names = ['Input','Sentiment'], encoding='utf-8')
test = pd.read_csv('/content/test (1).txt', header = None, sep =';', names = ['Input','Sentiment'],encoding='utf-8')
# + id="PkIYZDE-A7GR" colab={"base_uri": "https://localhost:8080/", "height": 357} outputId="b68388a7-4448-4338-c75b-c3722eb7e686"
#Let's check few rows of our train dataset
train.head(10)
# + colab={"base_uri": "https://localhost:8080/"} id="rIevKaSsIrzt" outputId="41f3caa3-1fc1-4295-9e92-40f700d9e11e"
# let's check category wise distribution of our train data
train.Sentiment.value_counts()
# + id="QwLNI2gN6zoV"
X_test = test.Input.tolist()
X_train = train.Input.tolist()
y_test = test.Sentiment.tolist()
y_train = train.Sentiment.tolist()
# + colab={"base_uri": "https://localhost:8080/"} id="3NzpgYAD99qn" outputId="a1913233-ee6f-4d6a-ea23-bf751a0075ca"
# Let's check size of our test and train datasets:
print(len(X_test),len(X_train),len(y_test),len(y_train))
# + id="daH4bTbg-jbR"
#These are the factors in our datasets
factors = ['sadness','joy','anger', 'fear', 'love','surprise']
# + id="xCkvGRCo_HCF"
#Let's encode categories into numeric values
encoding = { 'sadness': 0,'joy':1,'anger':2,'fear':3,'love':4,'surprise':5}
# + id="vYM_AW1iAfTp"
y_train = [encoding[key] for key in y_train]
y_test = [encoding[key] for key in y_test]
# + [markdown] id="kfRoWx9gApCy"
# Model Building
# + id="CZqjWxbGAkla" colab={"base_uri": "https://localhost:8080/", "height": 49, "referenced_widgets": ["f3c5ecd3c75e420ca31b13e774dfb271", "da0430973a7d49d69172eab566504b3b", "2b26eb13a9184168ba7b985b019bb380", "962e14f2dbd442d681edf5382c7cceee", "1f6007e1e45e45e295e2925d203b9c7c", "6ccdf62daa3549d1be57c0ad3216627e", "7e3a41a5d1154fc9aeccdd0d0b9972a3", "3ee952379f734d3bb94cffb49509a22a", "7dcc630c6b124dac8c863a710fe29143", "8e542bb86a194c478d7c380314b8f344", "dd911949dea74846908610df16396d7b"]} outputId="61b1aa3e-682c-4d9a-cef3-be1942f0fad5"
#Building the model using transformer
# BERT based uncased model is being used. So,I can choose any other model.
#let's select maxlen of tokenization as 512 (as it's max for BERT).
Model ='bert-base-uncased'
MAXLEN = 512
trans = text.Transformer(Model, maxlen=MAXLEN, class_names= factors)
# + colab={"base_uri": "https://localhost:8080/", "height": 336, "referenced_widgets": ["8e920048f8ae466293cbf3a2c1f1322c", "f21e6757de05414099070e7180fc7d5c", "294abede940f47dc8edacc8c05e70461", "e684ec08f3394187840743e7a9f0beb2", "15715fbe15e344fda2f25ccef33c4c1a", "<KEY>", "<KEY>", "<KEY>", "0effb2a310e04f108f82e789e4fa3479", "<KEY>", "20e8897b1a974a9d955c024c124c05ce"]} id="6pZu9Sz-Bw02" outputId="bd0f4fef-515a-4168-d602-558e1ffd0352"
#preprocessing out test and train data sets
test_data = trans.preprocess_test(X_test,y_test)
train_data = trans.preprocess_test(X_train,y_train)
# + colab={"base_uri": "https://localhost:8080/", "height": 102, "referenced_widgets": ["50001c48132a4f338d40c40e501765af", "06581b4ab1be42b2a4ecd2e3b49a3af1", "<KEY>", "6f509b1c6e2546cc8d72d6e8c6e8327b", "659c61524cbe4bf2a9e17b40b5ed1b7e", "<KEY>", "2f851f4d32b0470b840ef67f22efbfb2", "8387765f9c8a480698e68366510dba25", "<KEY>", "8f5f123de2ac43b5a6641d3d6a292224", "5407e73deb4e47dd810f6215f9bb32b0"]} id="tNdKusfLCl-q" outputId="0cd62437-ace0-4d71-952f-ad35ca80daa7"
model = trans.get_classifier()
# + id="_kEjxkCaDPcy"
learner = ktrain.get_learner(model, train_data=train_data, val_data=test_data, batch_size=10)
# + colab={"base_uri": "https://localhost:8080/", "height": 434} id="80SwvxO_DneY" outputId="a5ccd36e-6481-464b-a07d-b01ce97a421a"
learner.lr_find(show_plot=True, max_epochs=2)
# + colab={"base_uri": "https://localhost:8080/"} id="DKYfaYdc6Abe" outputId="8611a8f8-0e9c-4072-9dbc-44574ceb42a3"
learner.fit_onecycle(3e-2, 2)
# + [markdown] id="ot-P3zeYGUL1"
# Confusion Matrix
# + colab={"base_uri": "https://localhost:8080/"} id="4krSuotlGEWZ" outputId="14dedffd-11ef-49ee-c63d-f176a5681f06"
learner.validate(val_data=test_data, class_names=factors)
# + [markdown] id="pBLowPx9GgvB"
# Checking the top 5 data points which are not performing good:
# + id="WGSDy9A6GEd5" colab={"base_uri": "https://localhost:8080/"} outputId="8ded8b4d-a53a-4f40-bafe-0928a6bf6141"
learner.view_top_losses(n=5, preproc=trans)
# + id="CUdszTlrGEhl" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="3b596e6d-90f0-44b0-d9e9-e1ac59afa726"
X_test[152]
# + [markdown] id="oTmo_xdFG6BN"
# We can see that the above data our model is predicting as joy but label is mentioned as surprise.
# + [markdown] id="W-vCyRDZHNoE"
# Data Prediction
# + id="q9xJWoGvGEng"
predictor = ktrain.get_predictor(learner.model, preproc=trans)
# + id="pc2jY4YmGEqZ"
input = 'I am very happy with this new kind of front camera.'
# + colab={"base_uri": "https://localhost:8080/", "height": 89} id="xvH-DctA5e8K" outputId="14cdc410-e7a1-47bb-92d7-afb9281bdd98"
predictor.predict(input)
# + id="d1_UL8enPYIw"
submission = pd.DataFrame({ 'Input': train.Input.values, 'Predicted': predictor })
submission.to_csv("my_submission.csv", index=False)
# + id="JnBTUtyAPth7"
submission_test = pd.DataFrame({ 'Input': test.Input.values, 'Predicted': predictor })
submission.to_csv("submission.csv", index=False)
# + id="fd09eSUVUM_A"
| Sentiment_Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import random
parents = 100
daughters = 0
times = 0
while (parents > 0):
for _ in range(parents):
direction = random.choice([0,1,2,3])
if (direction == 0):
daughters += 1
parents -= 1
times += 1
print(parents, daughters)
print("Times:", times)
| ScienceLab1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
import pickle
import json
import re
import numpy as np
from scipy.sparse import csr_matrix
from sklearn.svm import LinearSVC
from sklearn.metrics import classification_report
def feature_transform(train_file, words):
words_len = len(words)
row_idx = []
column_idx = []
data = []
y = []
ctr = 0
feature_length = words_len # Per head
f = open(train_file)
for line in f:
ctr += 1
if(line.rstrip()):
line = re.sub("\s+"," ",line)
line1 = line.split(";")
a1 = line1[0].split(" ")
a2 = line1[1].split(" ")
a3 = line1[2].split(" ")
if(a1[0] == "H"):
column_idx.append(words.index(a1[1]))
elif(a1[0] == "ROOT"):
column_idx.append(words.index("ROOT"))
row_idx += [ctr-1]*2
data += [1] *2
column_idx.append(feature_length + words.index(a2[2]))
y.append(a3[1])
f.close()
X = csr_matrix((data, (row_idx, column_idx)), shape=(ctr,2*(words_len)))
return X, y
# +
listfile = "data_tokens.json"
f = open(listfile)
data = json.load(f)
f.close()
words = data["words"]
train_file = 'training_data.txt'
test_file = "testing_data.txt"
X_train, y_train = feature_transform(train_file, words)
X_test, y_test = feature_transform(test_file, words)
model = LinearSVC()
model.fit(X_train, y_train)
pred_train = model.predict(X_train)
pred_test = model.predict(X_test)
# -
print(classification_report(y_train, pred_train))
print(classification_report(y_test, pred_test))
| src/word_LRU.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Install Party
# ### Install Apps
#
# Make sure you have all of the following programs installed.
# **NOTE** Uninstall Anaconda 2 if you have it installed.
#
# - [Anaconda 3 (Python 3.5)](https://www.continuum.io/downloads)
# - [Sublime Text 3](https://www.sublimetext.com/3)
# - [Slack](https://slack.com/downloads) - the desktop app, not the website!
# - XCode Command Line Tools (Mac Only) Run: `xcode-select --install`
# - [Git (Windows Only)](https://git-scm.com/downloads)
# - [Homebrew (Mac Only)](http://brew.sh/)
# - [iTerm2 (Mac Only)](https://www.iterm2.com/)
# ### Install Extra Packages
# Run this in your terminal:
#
# `pip install version_information arrow seaborn ujson`
# ### Check Package Versions
# +
import numpy
import scipy
import matplotlib
import pandas
import statsmodels
import seaborn
import sklearn
import nltk
print("numpy:", numpy.__version__)
print("scipy:", scipy.__version__)
print("matplotlib:", matplotlib.__version__)
print("statsmodels:", statsmodels.__version__)
print("pandas:", pandas.__version__)
print("seaborn:", seaborn.__version__)
print("sklearn:", sklearn.__version__)
print("nltk:", nltk.__version__)
# -
# ### Download NLTK Data
# Run this in terminal:
#
# `python -m nltk.downloader -d ~/nltk_data stopwords brown`
#
# If you have extra space on your computer download all of the nltk data:
#
# `python -m nltk.downloader -d ~/nltk_data all`
# %reload_ext version_information
# %version_information
| notebooks/install-party.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a name="pagetop"></a>
# <div style="width:1000 px">
#
# <div style="float:right; width:98 px; height:98px;"><img src="https://pbs.twimg.com/profile_images/1187259618/unidata_logo_rgb_sm_400x400.png" alt="Unidata Logo" style="height: 98px;"></div>
#
# <h1>Plotting Satellite Data</h1>
# <h3>Unidata Python Workshop</h3>
#
# <div style="clear:both"></div>
# </div>
#
# <hr style="height:2px;">
#
# <div style="float:right; width:250 px"><img src="https://unidata.github.io/MetPy/latest/_images/sphx_glr_GINI_Water_Vapor_001.png" alt="Example Satellite Image" style="height: 500px;"></div>
#
#
# ## Overview:
#
# * **Teaching:** 60 minutes
# * **Exercises:** 30 minutes
#
# ### Questions
# 1. Where are current GOES data available?
# 1. How can satellite data be obtained with Siphon?
# 1. How can MetPy simplify metadata parsing?
# 1. How can maps of satellite data be made?
#
# ### Table of Contents
# 1. <a href="#findingdata">Finding GOES data</a>
# 1. <a href="#dataaccess">Accessing data with Siphon</a>
# 1. <a href="#parse">Digging into and parsing the data</a>
# 1. <a href="#plotting">Plotting the data</a>
# 1. <a href="#animation">Bonus: Animations</a>
# <hr style="height:2px;">
#
# <a name="findingdata"></a>
# ## Finding GOES Data
# The first step is to find the satellite data. Normally, we would browse over to http://thredds.ucar.edu/thredds/ and find the top-level [THREDDS Data Server (TDS)](https://www.unidata.ucar.edu/software/thredds/current/tds/TDS.html) catalog. From there we can drill down to find satellite data products.
#
# For current data, you could navigate to the `Test Datasets` directory, then `GOES Products` and `GOES-16`. There are subfolders for the CONUS, full disk, mesoscale sector images, and other products. In each of these is a folder for each [channel of the ABI](http://www.goes-r.gov/education/ABI-bands-quick-info.html). In each channel there is a folder for every day in the approximately month-long rolling archive. As you can see, there are a massive amount of data coming down from GOES-16!
#
# In the next section we'll be downloading the data in a pythonic way, so our first task is to build a URL that matches the URL we manually navigated to in the web browser. To make it as flexible as possible, we'll want to use variables for the sector name (CONUS, full-disk, mesoscale-1, etc.), the date, and the ABI channel number.
# ### Exercise
#
# * Create variables named `image_date`, `region`, and `channel`. Assign them to today's date, the mesoscale-1 region, and ABI channel 8.
# * Construct a string `data_url` from these variables and the URL we navigated to above.
# * Verify that following your link will take you where you think it should.
# * Change the extension from `catalog.html` to `catalog.xml`. What do you see?
# +
from datetime import datetime
# Create variables for URL generation
# YOUR CODE GOES HERE
# Construct the data_url string
# YOUR CODE GOES HERE
# Print out your URL and verify it works!
# YOUR CODE GOES HERE
# -
# #### Solution
# # %load solutions/data_url.py
# <a href="#pagetop">Top</a>
# <hr style="height:2px;">
# <a name="dataaccess"></a>
# ## Accessing data with Siphon
# We could download the files to our computers from the THREDDS web interface, but that can become tedious for downloading many files, requires us to store them on our computer, and does not lend itself to automation.
#
# We can use [Siphon](https://github.com/Unidata/siphon) parse the catalog from the TDS. This provides us a nice programmatic way of accessing the data. We start by importing the `TDSCatalog` class from siphon and giving it the URL to the catalog we just surfed to manually. **Note:** Instead of giving it the link to the HTML catalog, we change the extension to XML, which asks the TDS for the XML version of the catalog. This is much better to work with in code. If you forget, the extension will be changed for you with a warning being issued from siphon.
#
# We want to create a `TDSCatalog` object called `cat` that we can examine and use to get handles to work with the data.
from siphon.catalog import TDSCatalog
cat = TDSCatalog(data_url)
# To find the latest file, we can look at the `cat.datasets` attribute. Let’s look at the last five datasets:
cat.datasets[-5:]
# We'll get the next to most recent dataset (sometimes the most recent will not have received all tiles yet) and store it as variable `dataset`. Note that we haven't actually downloaded or transferred any real data yet, just bits of metadata have been received from THREDDS and parsed by siphon.
dataset = cat.datasets[-2]
print(dataset)
# We're finally ready to get the actual data. We could download the file, then open that, but there is no need! We can use siphon to help us only get what we need and hold it in memory. Notice that we're using the XArray accessor which will make life much nicer that dealing with the raw netCDF (like we used to back in the days of early 2018).
ds = dataset.remote_access(use_xarray=True)
# <a href="#pagetop">Top</a>
# <hr style="height:2px;">
# <a name="parse"></a>
# ## Digging into and parsing the data
# Now that we've got some data - let's see what we actually got our hands on.
ds
# Great, so we have an XArray Dataset object, something we've dealt with before! We also see that we have the coordinates `time`, `y`, and `x` as well as the data variables of `Sectorized_CMI` and the projection information.
#
# The first thing we can do is get an XArray DataArray with the CF compliant metadata parsed out to do things like automatically generate our cartopy crs for us!
import metpy # This gives us the metpy data accessor
dat = ds.metpy.parse_cf('Sectorized_CMI')
dat
# We can see some useful metadata are attached to this DataArray - including a coordinate refernce system. Let's look at what it is:
dat.metpy.crs
# Ok - so we have as `CFProjection` object - but we really want a cartopy compatible crs. Metpy provides a cartopy_crs accessor to do the translation for you:
proj = dat.metpy.cartopy_crs
proj
# Before we get to plotting, let's go ahead and get a handle to the x and y coordinates that we'll need. This is easy to do with the accessor on our data variable DataArray.
x = dat.metpy.x
y = dat.metpy.y
x
# Metadata travels with these coordinates!
# <a href="#pagetop">Top</a>
# <hr style="height:2px;">
# <a name="plotting"></a>
# ## Plotting the Data
# To plot our data we'll use the matplotlib method `imshow`. This function maps a 2D array of values to a 2D visual representation of those values, commonly known as a picture. It can even handle arrays that are MxNx3 or MxNx4 for RGB pictures (think digital pictures) or RGBA data. Let's look at a simple example:
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
a = np.array([[1, 0.5, 0.25, 0, 0, 0],
[0.5, 1, 0.5, 0.25, 0, 0],
[0.25, 0.5, 1, 0.5, 0.25, 0],
[0, 0.25, 0.5, 1, 0.5, 0.25],
[0, 0, 0.25, 0.5, 1, 0.5],
[0, 0, 0, 0.25, 0.5, 1]])
fig, ax = plt.subplots(1, 1, figsize=(10, 8))
ax.imshow(a)
lats = np.array([50, 48, 46, 44, 42, 40])
lons = np.array([-136, -134, -132, -130, -128, -126])
import cartopy.crs as ccrs
fig = plt.figure(figsize=(10, 8))
ax = fig.add_subplot(1, 1, 1, projection=ccrs.PlateCarree())
ax.imshow(a)
import cartopy.crs as ccrs
fig = plt.figure(figsize=(10, 8))
ax = fig.add_subplot(1, 1, 1, projection=ccrs.PlateCarree())
ax.imshow(a, extent=(lons[0], lons[-1], lats[0], lats[-1]),
origin='upper')
# You can see this is really a visual representation of the array data - it's also a handy way to quickly look at data, especially things like resolution matricies, etc.
# ### Exercise
#
# * Using what you've learned about `imshow` plot the data array dat with it.
# * Add the cartopy states and borders features to your map
# * BONUS: Use the metpy `add_timestamp` method from `metpy.plots` to add a timestamp to the plot.
# * DAILY DOUBLE: Using the `start_date_time` attribute on the dataset `ds`, change the call to `add_timestamp` to use that date and time and the pretext to say `GOES 16 Channel X`.
# +
import cartopy.feature as cfeature
from metpy.plots import add_timestamp
fig = plt.figure(figsize=(10, 8))
ax = fig.add_subplot(1, 1, 1, projection=proj)
# Plot the data using imshow
# YOUR CODE GOES HERE
# Add country borders and states (use your favorite linestyle!)
# YOUR CODE GOES HERE
# Bonus/Daily Double
# YOUR CODE GOES HERE
# -
# #### Solution
# # %load solutions/sat_map.py
# ### Using Colortables
# The map is much improved now, but it would look much better with a different color scheme.
#
# Colormapping in matplotlib (which backs CartoPy) is handled through two pieces:
#
# - The norm (normalization) controls how data values are converted to floating point values in the range [0, 1]
# - The colormap controls how values are converted from floating point values in the range [0, 1] to colors (think colortable)
#
# Let's start by setting the colormap to be black and white and normalizing the data to get the best contrast. We'll make a histogram to see the distribution of values in the data, then clip that range down to enhance contrast in the data visualization. **Note:** `cmap` and `norm` can also be set during the `imshow` call as keyword arguments.
#
# We use `compressed` to remove any masked elements before making our histogram.
plt.hist(dat.to_masked_array().compressed(), bins=255);
# +
fig = plt.figure(figsize=(10, 8))
ax = fig.add_subplot(1, 1, 1, projection=proj)
# Plot the data using imshow
im = ax.imshow(dat, origin='upper',
extent=(x.min(), x.max(), y.min(), y.max()))
# Add country borders and states (use your favorite linestyle!)
ax.add_feature(cfeature.BORDERS, linewidth=2, edgecolor='black')
ax.add_feature(cfeature.STATES, linestyle=':', edgecolor='black')
# Bonus/Daily Double
timestamp = datetime.strptime(ds.start_date_time, '%Y%j%H%M%S')
add_timestamp(ax, timestamp, pretext='GOES-16 Ch.{} '.format(channel),
high_contrast=True, fontsize=16, y=0.01)
# +
# Set colormap
im.set_cmap('Greys')
# Set norm
im.set_norm(plt.Normalize(200,255))
# Show figure again
fig
# -
# In meteorology, we have many ‘standard’ colortables that have been used for certain types of data. We have included these in Metpy in the `metpy.plots.ctables` module. By importing the `ColortableRegistry` we gain access to the colortables, as well as handy normalizations to go with them. We can see the colortables available by looking at the dictionary keys. The colortables ending in `_r` are the reversed versions of the named colortable.
#
# Let’s use the `WVCIMSS` colormap, a direct conversion of the GEMPAK colormap. The code below asks for the colormap, as well as a normalization that covers the range 195 to 265. This was empirically determined to closely match other displays of water vapor data. We then apply it to the existing image we have been working with.
from metpy.plots import colortables
wv_norm, wv_cmap = colortables.get_with_range('WVCIMSS_r', 195, 265)
im.set_cmap(wv_cmap)
im.set_norm(wv_norm)
fig
# <a href="#pagetop">Top</a>
# <hr style="height:2px;">
# <a name="animation"></a>
# ## Bonus: Animations
# **NOTE:**
# This is just a quick taste of producing an animation using matplotlib. The animation support in matplotlib is robust, but sometimes installation of the underlying tool (ffmpeg) can be a little tricky. In order to make sure we get don't get bogged down, this is really more of a demo than something expected to work out of the box.
#
# Conda-forge has packages, so it may be as easy as:
# +
# #!conda install -y -n unidata-workshop -c http://conda.anaconda.org/conda-forge ffmpeg
# -
# First, we'll import the animation support from matplotlib. We also tell it that we want it to render the animations to HTML using the HTML5 video tag:
import os.path
import sys
from IPython.display import HTML
from matplotlib.animation import ArtistAnimation
# We create the base figure, then we loop over a bunch of the datasets to create an animation. For each one we pull out the data and plot both the timestamp and the image. The `ArtistAnimation` class takes the `Figure` instance and a list as required arguments. The contents of this list are a collection of matplotlib artists for each frame of the animation. In the loop below, we populate this list with the `Text` instance created when adding the timestamp as well as the image that results from plotting the data.
# +
import matplotlib as mpl
mpl.rcParams['animation.embed_limit'] = 50
# List used to store the contents of all frames. Each item in the list is a tuple of
# (image, text)
artists = []
# Get the IRMA case study catalog
cat = TDSCatalog('http://thredds.ucar.edu/thredds/catalog/casestudies/irma'
'/goes16/Mesoscale-1/Channel{:02d}/{:%Y%m%d}/'
'catalog.xml'.format(channel, datetime(2017, 9, 9)))
datasets = cat.datasets.filter_time_range(datetime(2017, 9, 9), datetime(2017, 9, 9, 6))
# Grab the first dataset and make the figure using its projection information
ds = datasets[0]
ds = ds.remote_access(use_xarray=True)
dat = ds.metpy.parse_cf('Sectorized_CMI')
proj = dat.metpy.cartopy_crs
fig = plt.figure(figsize=(10,8))
ax = fig.add_subplot(1, 1, 1, projection=proj)
plt.subplots_adjust(left=0.005, bottom=0.005, right=0.995, top=0.995, wspace=0, hspace=0)
ax.coastlines(resolution='50m', color='black')
ax.add_feature(cfeature.BORDERS, linewidth=2)
# Loop over the datasets and make the animation
for ds in datasets[::6]:
# Open the data
ds = ds.remote_access(service='OPENDAP', use_xarray=True)
dat = ds.metpy.parse_cf('Sectorized_CMI')
# Pull out the image data, x and y coordinates, and the time. Also go ahead and
# convert the time to a python datetime
x = dat['x']
y = dat['y']
timestamp = datetime.strptime(ds.start_date_time, '%Y%j%H%M%S')
img_data = ds['Sectorized_CMI']
# Plot the image and the timestamp. We save the results of these plotting functions
# so that we can tell the animation that these two things should be drawn as one
# frame in the animation
im = ax.imshow(dat, extent=(x.min(), x.max(), y.min(), y.max()), origin='upper',
cmap=wv_cmap, norm=wv_norm)
text_time = add_timestamp(ax, timestamp, pretext='GOES-16 Ch.{} '.format(channel),
high_contrast=True, fontsize=16, y=0.01)
# Stuff them in a tuple and add to the list of things to animate
artists.append((im, text_time))
# Create the animation--in addition to the required args, we also state that each
# frame should last 200 milliseconds
anim = ArtistAnimation(fig, artists, interval=200., blit=False)
anim.save('GOES_Animation.mp4')
HTML(anim.to_jshtml())
# -
# <a href="#pagetop">Top</a>
# <hr style="height:2px;">
| notebooks/Satellite_Data/PlottingSatelliteData.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### NLP | How tokenizing text, sentence, words works
# ### Text into sentences tokenization
# ### Sentences into words tokenization
# ### Sentences using regular expressions tokenization
# import dataset
from nltk.tokenize import sent_tokenize, word_tokenize,TreebankWordTokenizer, WordPunctTokenizer, RegexpTokenizer, regexp_tokenize
# # Sentence Tokenization – Splitting sentences in the paragraph
# +
text = "Hello everyone. Welcome to Government MCA College. You are studying NLP article"
print("Original Text :- {}".format(text))
print("Sentence Tokenization :- {}".format(sent_tokenize(text)))
# -
# # Word Tokenization – Splitting words in a sentence.
print("Word Tokenization :-\n {}".format(word_tokenize(text)))
# # Using TreebankWordTokenizer
tokenizer = TreebankWordTokenizer()
print("Using TreebankWordTokenizer :-\n {}".format(tokenizer.tokenize(text)))
# # WordPunctTokenizer – It seperates the punctuation from the words.
tokenizer = WordPunctTokenizer()
print("WordPunctTokenizer :- {}".format(tokenizer.tokenize("Let's see how it's working.")))
# # Using Regular Expression
# +
tokenizer = RegexpTokenizer("[\w']+")
text1 = "Let's see how it's working."
print("Using Regular Expression using RegexpTokenizer :- \n{}".format(tokenizer.tokenize(text1)))
text2 = "Let's see how it's working."
print("Using Regular Expression using regexp_tokenize funcation :- \n{}".format(regexp_tokenize(text, "[\w']+")))
| Semester V/Big Data Analytics (BDA) (4659306)/25/BDA_PR_25.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Extracting the SI prefix
#
# `ubermagutil` provides convenience methods for determining the prefix of a value, such as pico, micro, mega, etc. The prefixes are:
# +
import ubermagutil.units as uu
uu.si_prefixes
# -
# From a single value, prefix can be extracted using `si_multiplier`:
uu.si_multiplier(5e-12)
uu.si_multiplier(-3e6)
# Similarly, for a list of values, the largest prefix can be determined using `si_max_multiplier`:
uu.si_max_multiplier([2e-3, 2e-6, 6e-9])
| docs/si-prefix.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + deletable=false editable=false
# Initialize OK
from client.api.notebook import Notebook
ok = Notebook('hw11.ok')
# -
# # Homework 11: Classification
# **Reading**:
#
# * [Classification](https://www.inferentialthinking.com/chapters/17/classification.html)
# Please complete this notebook by filling in the cells provided. Before you begin, execute the following cell to load the provided tests. Each time you start your server, you will need to execute this cell again to load the tests.
#
# Directly sharing answers is not okay, but discussing problems with the course staff or with other students is encouraged.
#
# For all problems that you must write our explanations and sentences for, you **must** provide your answer in the designated space. Moreover, throughout this homework and all future ones, please be sure to not re-assign variables throughout the notebook! For example, if you use `max_temperature` in your answer to one question, do not reassign it later on.
# +
# Don't change this cell; just run it.
import numpy as np
from datascience import *
# These lines do some fancy plotting magic.
import matplotlib
# %matplotlib inline
import matplotlib.pyplot as plt
plt.style.use('fivethirtyeight')
import warnings
warnings.simplefilter('ignore', FutureWarning)
from matplotlib import patches
from ipywidgets import interact, interactive, fixed
import ipywidgets as widgets
from client.api.notebook import Notebook
ok = Notebook('hw11.ok')
# -
# ## 1. Reading Sign Language with Classification
#
# Brazilian Sign Language is a visual language used primarily by Brazilians who are deaf. It is more commonly called Libras. People who communicate with visual language are called *signers*. Here is a video of someone signing in Libras:
from IPython.lib.display import YouTubeVideo
YouTubeVideo("JD1AwcdkUUs")
# Programs like Siri or Google Now begin the process of understanding human speech by classifying short clips of raw sound into basic categories called *phones*. For example, the recorded sound of someone saying the word "robot" might be broken down into several phones: "rrr", "oh", "buh", "aah", and "tuh". Phones are then grouped together into further categories like words ("robot") and sentences ("I, for one, welcome our new robot overlords") that carry more meaning.
#
# A visual language like Libras has an analogous structure. Instead of phones, each word is made up of several *hand movements*. As a first step in interpreting Libras, we can break down a video clip into small segments, each containing a single hand movement. The task is then to figure out what hand movement each segment represents.
#
# We can do that with classification!
#
# The [data](https://archive.ics.uci.edu/ml/machine-learning-databases/libras/movement_libras.names) in this exercise come from Dias, Peres, and Biscaro, researchers at the University of Sao Paulo in Brazil. They identified 15 distinct hand movements in Libras (probably an oversimplification, but a useful one) and captured short videos of signers making those hand movements. (You can read more about their work [here](http://ieeexplore.ieee.org/Xplore/login.jsp?url=http%3A%2F%2Fieeexplore.ieee.org%2Fiel5%2F5161636%2F5178557%2F05178917.pdf&authDecision=-203). The paper is gated, so you will need to use your institution's Wi-Fi or VPN to access it.)
#
# For each video, they chose 45 still frames from the video and identified the location (in horizontal and vertical coordinates) of the signer's hand in each frame. Since there are two coordinates for each frame, this gives us a total of 90 numbers summarizing how a hand moved in each video. Those 90 numbers will be our *attributes*.
#
# Each video is *labeled* with the kind of hand movement the signer was making in it. Each label is one of 15 strings like "horizontal swing" or "vertical zigzag".
#
# For simplicity, we're going to focus on distinguishing between just two kinds of movements: "horizontal straight-line" and "vertical straight-line". We took the Sao Paulo researchers' original dataset, which was quite small, and used some simple techniques to create a much larger synthetic dataset.
#
# These data are in the file `movements.csv`. Run the next cell to load it.
# + deletable=false
movements = Table.read_table("movements.csv")
movements.take(np.arange(5))
# -
# The cell below displays movements graphically. Run it and use the slider to answer the next question.
# + deletable=false
# Just run this cell and use the slider it produces.
def display_whole_movement(row_idx):
num_frames = int((movements.num_columns-1)/2)
row = np.array(movements.drop("Movement type").row(row_idx))
xs = row[np.arange(0, 2*num_frames, 2)]
ys = row[np.arange(1, 2*num_frames, 2)]
plt.figure(figsize=(5,5))
plt.plot(xs, ys, c="gold")
plt.xlabel("x")
plt.ylabel("y")
plt.xlim(-.5, 1.5)
plt.ylim(-.5, 1.5)
plt.gca().set_aspect('equal', adjustable='box')
def display_hand(example, frame, display_truth):
time_idx = frame-1
display_whole_movement(example)
x = movements.column(2*time_idx).item(example)
y = movements.column(2*time_idx+1).item(example)
plt.annotate(
"frame {:d}".format(frame),
xy=(x, y), xytext=(-20, 20),
textcoords = 'offset points', ha = 'right', va = 'bottom',
color='white',
bbox = {'boxstyle': 'round,pad=0.5', 'fc': 'black', 'alpha':.4},
arrowprops = {'arrowstyle': '->', 'connectionstyle':'arc3,rad=0', 'color': 'black'})
plt.scatter(x, y, c="black", zorder=10)
plt.title("Hand positions for movement {:d}{}".format(example, "\n(True class: {})".format(movements.column("Movement type").item(example)) if display_truth else ""))
def animate_movement():
interact(
display_hand,
example=widgets.BoundedIntText(min=0, max=movements.num_rows-1, value=0, msg_throttle=1),
frame=widgets.IntSlider(min=1, max=int((movements.num_columns-1)/2), step=1, value=1, msg_throttle=1),
display_truth=fixed(False))
animate_movement()
# + [markdown] deletable=false editable=false
# #### Question 1
#
# Before we move on, check your understanding of the dataset. Judging by the plot, is the first movement example a vertical motion, or a horizontal motion? If it is hard to tell, does it seem more likely to be vertical or horizontal? This is the kind of question a classifier has to answer. Find out the right answer by looking at the `Movement type` column.
#
# Assign `first_movement` to `1` if the movement was vertical, or `2` if the movement was horizontal.
#
# <!--
# BEGIN QUESTION
# name: q1_1
# manual: false
# -->
# + deletable=false
first_movement = ...
# + deletable=false editable=false
ok.grade("q1_1");
# -
# ### Splitting the dataset
# We'll do 2 different kinds of things with the `movements` dataset:
# 1. We'll build a classifier that uses the movements with known labels as examples to classify similar movements. This is called *training*.
# 2. We'll evaluate or *test* the accuracy of the classifier we build.
#
# For reasons discussed in lecture and the textbook, we want to use separate datasets for these two purposes. So we split up our one dataset into two.
# + [markdown] deletable=false editable=false
# #### Question 2
#
# Create a table called `train_movements` and another table called `test_movements`. `train_movements` should include the first $\frac{11}{16}$th of the rows in `movements` (rounded to the nearest integer), and `test_movements` should include the remaining $\frac{5}{16}$th.
#
# Note that we do **not** mean the first 11 rows for the training test and rows 12-16 for the test set. We mean the first $\frac{11}{16} = 68.75$% of the table should be for the the trianing set, and the rest should be for the test set.
#
# *Hint:* Use the table method `take`.
#
# <!--
# BEGIN QUESTION
# name: q1_2
# manual: false
# -->
# + for_assignment_type="solution"
training_proportion = 11/16
num_movements = movements.num_rows
num_train = int(round(num_movements * training_proportion))
train_movements = ...
test_movements = ...
print("Training set:\t", train_movements.num_rows, "examples")
print("Test set:\t", test_movements.num_rows, "examples")
# + deletable=false editable=false
ok.grade("q1_2");
# -
# ### Using only 2 features
# First let's see how well we can distinguish two movements (a vertical line and a horizontal line) using the hand position from just a single frame (without the other 44).
# + [markdown] deletable=false editable=false
# #### Question 3
#
# Make a table called `train_two_features` with only 3 columns: the first frame’s x coordinate and first frame’s y coordinate (which are our chosen features), as well as the movement type. Use only the examples in train_movements.
#
# <!--
# BEGIN QUESTION
# name: q1_3
# manual: false
# -->
# + deletable=false
train_two_features = ...
train_two_features
# + deletable=false editable=false
ok.grade("q1_3");
# -
# Now we want to make a scatter plot of the frame coordinates, where the dots for horizontal straight-line movements have one color and the dots for vertical straight-line movements have another color. Here is a scatter plot without colors:
# + deletable=false
train_two_features.scatter("Frame 1 x", "Frame 1 y")
# -
# This isn't useful because we don't know which dots are which movement type. We need to tell Python how to color the dots. Let's use gold for vertical and blue for horizontal movements.
#
# `scatter` takes an extra argument called `colors` that's the name of an extra column in the table that contains colors (strings like "red" or "orange") for each row. So we need to create a table like this:
#
# |Frame 1 x|Frame 1 y|Movement type|Color|
# |-|-|-|-|
# |0.522768|0.769731|vertical straight-line|gold|
# |0.179546|0.658986|horizontal straight-line|blue|
# |...|...|...|...|
# + [markdown] deletable=false editable=false
# <div class="hide">\pagebreak</div>
#
# #### Question 4
#
# In the cell below, create a table named `with_colors`. It should have the same columns as the example table above, but with a row for each row in `train_two_features`. Then, create a scatter plot of your data.
#
# <!--
# BEGIN QUESTION
# name: q1_4
# manual: true
# image: true
# -->
# <!-- EXPORT TO PDF -->
# + deletable=false export_pdf=true manual_grade=true manual_problem_id="sign_lang_2"
# You should find the following table useful.
type_to_color = Table().with_columns(
"Movement type", make_array("vertical straight-line", "horizontal straight-line"),
"Color", make_array("gold", "blue"))
with_colors = ...
with_colors.scatter("Frame 1 x", "Frame 1 y", group="Color")
# + [markdown] deletable=false editable=false
# <div class="hide">\pagebreak</div>
#
# #### Question 5
#
# Based on the scatter plot, how well will a nearest-neighbor classifier based on only these 2 features (the x- and y-coordinates of the hand position in the first frame) work? Will it:
#
# 1. distinguish almost perfectly between vertical and horizontal movements;
# 2. distinguish somewhat well between vertical and horizontal movements, getting some correct but missing a substantial proportion; or
# 3. be basically useless in distinguishing between vertical and horizontal movements?
#
# # Why?
#
# <!--
# BEGIN QUESTION
# name: q1_5
# manual: true
# -->
# <!-- EXPORT TO PDF -->
# + [markdown] deletable=false export_pdf=true manual_grade=true manual_problem_id="sign_lang_3"
# *Write your answer here, replacing this text.*
# -
# ## 2. Classification Potpourri
#
# Throughout this question, we will aim to discuss some conceptual nuances of classification that often get overlooked when we're focused only on improving our accuracy and building the best classifier possible.
# + [markdown] deletable=false editable=false
# #### Question 1
#
# What is the point of a test-set? Should we use our test set to find the best possible number of neighbors for a k-NN classifer? Explain.
#
# <!--
# BEGIN QUESTION
# name: q2_1
# manual: true
# -->
# <!-- EXPORT TO PDF -->
# + [markdown] deletable=false export_pdf=true manual_grade=true manual_problem_id="potpourri_1"
# *Write your answer here, replacing this text.*
# + [markdown] deletable=false editable=false
# #### Question 2
# You have a large dataset which contains three columns. The first two are attributes of the person that might be predictive of whether or not someone has breast-cancer, and the third column indicates whether they have it or not. 99% of the table contains examples of people who do not have breast cancer.
#
# Imagine you are trying to use a k-NN classifier to use the first two columns to predict whether or not someone has breast cancer. You split your training and test set up as necessary, you develop a 7-NN classifier, and you notice your classifier predicts every point in the test set to be a person who does not have breast cancer. Is there a problem with your classifier? Explain this phenomenon.
#
# <!--
# BEGIN QUESTION
# name: q2_2
# manual: true
# -->
# <!-- EXPORT TO PDF -->
# + [markdown] deletable=false export_pdf=true manual_grade=true manual_problem_id="potpourri_2"
# *Write your answer here, replacing this text.*
# + [markdown] deletable=false editable=false
# #### Question 3
# You have a training set with data on the characteristics of 35 examples of fruit. 25 of the data points are apples, and the remaining 10 are oranges.
#
# You decide to make a k-NN classifier. Assign `k_upper_bound` to the smallest possible k such that the classifier will predict Apple for every point, regardless of how the data is spread out.
#
# Imagine that ties are broken at random for even values of k, so there is no guarantee of what will be picked if there is a tie.
#
# <!--
# BEGIN QUESTION
# name: q2_3
# manual: false
# -->
#
# + deletable=false
k_upper_bound = ...
# + deletable=false editable=false
ok.grade("q2_3");
# -
# ## 3. Submission
#
# Once you're finished, select "Save and Checkpoint" in the File menu, download this file in .ipynb format, and upload to Gradescope.
_ = ok.submit()
# For your convenience, you can run this cell to run all the tests at once!
import os
print("Running all tests...")
_ = [ok.grade(q[:-3]) for q in os.listdir("tests") if q.startswith('q') and len(q) <= 10]
print("Finished running all tests.")
| hw/hw11/hw11.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 38
# language: python
# name: python38
# ---
# + language="html"
# <!--Script block to left align Markdown Tables-->
# <style>
# table {margin-left: 0 !important;}
# </style>
# -
# Preamble script block to identify host, user, and kernel
import sys
# ! hostname
# ! whoami
print(sys.executable)
print(sys.version)
print(sys.version_info)
# ## Full name:
# ## R#:
# ## HEX:
# ## Title of the notebook
# ## Date:
# # Pandas
#
# A data table is called a `DataFrame` in pandas (and other programming environments too).
#
# The figure below from https://pandas.pydata.org/docs/getting_started/index.html illustrates a dataframe model:
#
# 
#
# Each column and each row in a dataframe is called a series, the header row, and index column are special.
#
# To use pandas, we need to import the module, generally pandas has numpy as a dependency so it also must be imported
#
import numpy as np #Importing NumPy library as "np"
import pandas as pd #Importing Pandas library as "pd"
# # Dataframe-structure using primative python
#
# First lets construct a dataframe like object using python primatives.
# We will construct 3 lists, one for row names, one for column names, and one for the content.
mytabular = np.random.randint(1,100,(5,4))
myrowname = ['A','B','C','D','E']
mycolname = ['W','X','Y','Z']
mytable = [['' for jcol in range(len(mycolname)+1)] for irow in range(len(myrowname)+1)] #non-null destination matrix, note the implied loop construction
# The above builds a placeholder named `mytable` for the psuedo-dataframe.
# Next we populate the table, using a for loop to write the column names in the first row, row names in the first column, and the table fill for the rest of the table.
for irow in range(1,len(myrowname)+1): # write the row names
mytable[irow][0]=myrowname[irow-1]
for jcol in range(1,len(mycolname)+1): # write the column names
mytable[0][jcol]=mycolname[jcol-1]
for irow in range(1,len(myrowname)+1): # fill the table (note the nested loop)
for jcol in range(1,len(mycolname)+1):
mytable[irow][jcol]=mytabular[irow-1][jcol-1]
# Now lets print the table out by row and we see we have a very dataframe-like structure
for irow in range(0,len(myrowname)+1):
print(mytable[irow][0:len(mycolname)+1])
# We can also query by row
print(mytable[3][0:len(mycolname)+1])
# Or by column
for irow in range(0,len(myrowname)+1): #cannot use implied loop in a column slice
print(mytable[irow][2])
# Or by row+column index; sort of looks like a spreadsheet syntax.
print(' ',mytable[0][3])
print(mytable[3][0],mytable[3][3])
# # Create a proper dataframe
# We will now do the same using pandas
df = pd.DataFrame(np.random.randint(1,100,(5,4)), ['A','B','C','D','E'], ['W','X','Y','Z'])
df
# We can also turn our table into a dataframe, notice how the constructor adds header row and index column
df1 = pd.DataFrame(mytable)
df1
# To get proper behavior, we can just reuse our original objects
df2 = pd.DataFrame(mytabular,myrowname,mycolname)
df2
# ### Getting the shape of dataframes
#
# The shape method will return the row and column rank (count) of a dataframe.
#
df.shape
df1.shape
df2.shape
# ### Appending new columns
# To append a column simply assign a value to a new column name to the dataframe
df['new']= None
df
# ## Appending new rows
# A bit trickier but we can create a copy of a row and concatenate it back into the dataframe.
newrow = df.loc[['E']].rename(index={"E": "X"}) # create a single row, rename the index
newtable = pd.concat([df,newrow]) # concatenate the row to bottom of df - note the syntax
newtable
# ### Removing Rows and Columns
#
# To remove a column is straightforward, we use the drop method
newtable.drop('new', axis=1, inplace = True)
newtable
# To remove a row, you really got to want to, easiest is probablty to create a new dataframe with the row removed
newtable = newtable.loc[['A','B','D','E','X']] # select all rows except C
newtable
# # Indexing
# We have already been indexing, but a few examples follow:
newtable['X'] #Selecing a single column
newtable[['X','W']] #Selecing a multiple columns
newtable.loc['E'] #Selecing rows based on label via loc[ ] indexer
newtable.loc[['E','X','B']] #Selecing multiple rows based on label via loc[ ] indexer
newtable.loc[['B','E','D'],['X','Y']] #Selecting elemens via both rows and columns via loc[ ] indexer
# # Conditional Selection
df = pd.DataFrame({'col1':[1,2,3,4,5,6,7,8],
'col2':[444,555,666,444,666,111,222,222],
'col3':['orange','apple','grape','mango','jackfruit','watermelon','banana','peach']})
df
# +
#What fruit corresponds to the number 555 in ‘col2’?
df[df['col2']==555]['col3']
# +
#What fruit corresponds to the minimum number in ‘col2’?
df[df['col2']==df['col2'].min()]['col3']
# -
# # Descriptor Functions
# +
#Creating a dataframe from a dictionary
df = pd.DataFrame({'col1':[1,2,3,4,5,6,7,8],
'col2':[444,555,666,444,666,111,222,222],
'col3':['orange','apple','grape','mango','jackfruit','watermelon','banana','peach']})
df
# -
# ### `head` method
#
# Returns the first few rows, useful to infer structure
# +
#Returns only the first five rows
df.head()
# -
# ### `info` method
#
# Returns the data model (data column count, names, data types)
# +
#Info about the dataframe
df.info()
# -
# ### `describe` method
#
# Returns summary statistics of each numeric column.
# Also returns the minimum and maximum value in each column, and the IQR (Interquartile Range).
# Again useful to understand structure of the columns.
# +
#Statistics of the dataframe
df.describe()
# -
# ### Counting and Sum methods
#
# There are also methods for counts and sums by specific columns
df['col2'].sum() #Sum of a specified column
# The `unique` method returns a list of unique values (filters out duplicates in the list, underlying dataframe is preserved)
df['col2'].unique() #Returns the list of unique values along the indexed column
# The `nunique` method returns a count of unique values
df['col2'].nunique() #Returns the total number of unique values along the indexed column
# The `value_counts()` method returns the count of each unique value (kind of like a histogram, but each value is the bin)
df['col2'].value_counts() #Returns the number of occurences of each unique value
# ## Using functions in dataframes - symbolic apply
#
# The power of pandas is an ability to apply a function to each element of a dataframe series (or a whole frame) by a technique called symbolic (or synthetic programming) application of the function.
#
# Its pretty complicated but quite handy, best shown by an example
# +
def times2(x): # A prototype function to scalar multiply an object x by 2
return(x*2)
print(df)
print('Apply the times2 function to col2')
df['col2'].apply(times2) #Symbolic apply the function to each element of column col2, result is another dataframe
# -
# ## Sorts
df.sort_values('col2', ascending = True) #Sorting based on columns
# ## Exercise 1
# Create a prototype function to compute the cube root of a numeric object (literally two lines to define the function), recall exponentation is available in primative python.
#
# Apply your function to column **'X'** of dataframe **`newtable`** created above
# +
# Define your function here:
# Symbolic apply here:
# -
# # Aggregating (Grouping Values) dataframe contents
#
# +
#Creating a dataframe from a dictionary
data = {
'key' : ['A', 'B', 'C', 'A', 'B', 'C'],
'data1' : [1, 2, 3, 4, 5, 6],
'data2' : [10, 11, 12, 13, 14, 15],
'data3' : [20, 21, 22, 13, 24, 25]
}
df1 = pd.DataFrame(data)
df1
# +
# Grouping and summing values in all the columns based on the column 'key'
df1.groupby('key').sum()
# +
# Grouping and summing values in the selected columns based on the column 'key'
df1.groupby('key')[['data1', 'data2']].sum()
# -
# # Filtering out missing values
# +
#Creating a dataframe from a dictionary
df = pd.DataFrame({'col1':[1,2,3,4,None,6,7,None],
'col2':[444,555,None,444,666,111,None,222],
'col3':['orange','apple','grape','mango','jackfruit','watermelon','banana','peach']})
df
# -
# Below we drop any row that contains a `NaN` code.
df_dropped = df.dropna()
df_dropped
# Below we replace `NaN` codes with some value, in this case 0
df_filled1 = df.fillna(0)
df_filled1
# Below we replace `NaN` codes with some value, in this case the mean value of of the column in which the missing value code resides.
df_filled2 = df.fillna(df.mean())
df_filled2
# ## Exercise 2
# Replace the **'NaN'** codes with the string 'missing' in dataframe **'df'**
# +
# Replace the NaN with the string 'missing' here:
# -
# # Reading a File into a Dataframe
#
# Pandas has methods to read common file types, such as `csv`,`xlsx`, and `json`. Ordinary text files are also quite manageable.
#
# On a machine you control you can write script to retrieve files from the internet and process them.
#
# On `CoCalc` you have to manually upload the target file to the directory where the script resides. The system commands `wget` and `curl` are blocked in the free accounts.
readfilecsv = pd.read_csv('CSV_ReadingFile.csv') #Reading a .csv file
print(readfilecsv)
# Similar to reading and writing .csv files, you can also read and write .xslx files as below (useful to know this)
readfileexcel = pd.read_excel('Excel_ReadingFile.xlsx', sheet_name='Sheet1') #Reading a .xlsx file
print(readfileexcel)
# # Writing a dataframe to file
#Creating and writing to a .csv file
readfilecsv = pd.read_csv('CSV_ReadingFile.csv')
readfilecsv.to_csv('CSV_WritingFile1.csv')
readfilecsv = pd.read_csv('CSV_WritingFile1.csv')
print(readfilecsv)
#Creating and writing to a .csv file by excluding row labels
readfilecsv = pd.read_csv('CSV_ReadingFile.csv')
readfilecsv.to_csv('CSV_WritingFile2.csv', index = False)
readfilecsv = pd.read_csv('CSV_WritingFile2.csv')
print(readfilecsv)
#Creating and writing to a .xlsx file
readfileexcel = pd.read_excel('Excel_ReadingFile.xlsx', sheet_name='Sheet1', index = False)
readfileexcel.to_excel('Excel_WritingFile.xlsx', sheet_name='MySheet', index = False)
readfileexcel = pd.read_excel('Excel_WritingFile.xlsx', sheet_name='MySheet', index = False)
print(readfileexcel)
# ## Exercise 3
# Download the file named `concreteData.xls` if you have not done so, then upload to your CoCalc environment.
#
# Read the file into a dataframe named **'concreteData'**
#
# Then perform the following activities.
#
# 1. Examine the first few rows of the dataframe and describe the structure (using words) in a markdown cell just after you run the descriptor method
#
# 2. Simplify the column names to "Cement", "BlastFurnaceSlag", "FlyAsh", "Water", "Superplasticizer", "CoarseAggregate", "FineAggregate", "Age", "CC_Strength"
#
# 3. Determine and report summary statistics for each of the columns.
#
# 4. Then insert and run the script below into your notebook (after the summary statistics), describe the output (using words) in a markdown cell.
#
# import matplotlib.pyplot as plt
# import seaborn as sns
# # %matplotlib inline
# sns.pairplot(concreteData)
# plt.show()
# Read the file in this cell
# Use head method to show first few rows, describe the data model
# Rename columns in this cell (Hard, but use example above, there is a hint in a text file named hint.txt)
# Use describe method to find summary statistics for each column
# Insert and run the required script, be sure the dataframe is named correctly
# You will get a pretty cool plot if all goes well
# The script takes awhile (about a minute on my server)
| 1-Lessons/Lesson08/Lab8/src/Lab8-FullNarrative.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # A self-inflicted tutorial on sampling schemes
# Following notes available at:
# * [Tutorial by Andrieu et. al. ](http://www.cs.ubc.ca/~arnaud/andrieu_defreitas_doucet_jordan_intromontecarlomachinelearning.pdf 'An introduction to MCMC for Machine Learning')
#
from __future__ import print_function
import torch
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
from scipy.stats import multivariate_normal as mv
from scipy.stats import norm
# %matplotlib inline
import matplotlib.mlab as mlab
import matplotlib.cm as cm
from sklearn import mixture
# +
# Calculate pdf (normalized) for x, mean m and stdev s
# If x is an 1-dim array, should use 1D
# Bimodal uses scipy and is normalized
gauss_1D = lambda x,m,s: norm.pdf(x,loc=m,scale=s)
# np.exp(-(x - m)**2/(2 * s**2)) / (s*np.sqrt(np.pi*2))
# Uncomment above for un-normalized
gauss_2D = lambda x,m,sigma: mv.pdf(x,m,sigma)
# -
# Test 1D
n = np.linspace(-5,5,100)
res = gauss_1D(n,1,0.3)
plt.plot(n,res);
# Test 2D
n = np.linspace(0,30,100)
m = np.linspace(0,30,100)
x,y = np.meshgrid(n,m)
# Scipy needs a third axis, hence the use of dstack
pos = np.dstack((x, y))
cov_indepedent = [[4,0],[0,9]]
cov_mixed = [[4,2],[2,9]]
#plot the pdf for independent normals
res2 = gauss_2D(pos,[10,15],cov_mixed)
plt.contourf(x,y,res2);
# ## Gaussian Mixture Models
# - 1D
n = np.arange(-50,50)
res_mixed= 2* gauss_1D(n,10,2) + 3*gauss_1D(n,20,2)
plt.plot(n,res_mixed);
# The workflow for **2D** is the following:
# - Generate a gaussian mixture model by calculating pdfs
# - Sample from the gmm using your *favourite* sampling scheme
# - Plot the gmm together with the samples
# +
def gmm(x,y,mu1,mu2,cov1,cov2):
pos = np.dstack((x,y))
g1 = gauss_2D(pos,mu1,cov1)
g2 = gauss_2D(pos,mu2,cov2)
return (2*g1 + 3*g2)/5
# Covariances and means
mu1 = [-1,-1]
mu2 = [1,2]
cov1 = [[1,-0.8],[-0.8,1]]
cov2 = [[1.5,0.6],[0.6,0.8]]
# -
# # Metropolis Hastings MCMC
# The goal is to sample correctly from the target distribution $p(x)$ defined as GMM here. Ingredients:
# * A proposal $q(x^*|x)$ here taken to be $x^* \sim N(x,\sigma)$
# * Accept $x^*$ with $min\{1,\frac{p(x^*)q(x|x^*)}{p(x)q(x^*|x)}\}$
# * More specifically, if $(U(0,1) < acc)$ then $x = x^*$
# Just fooling around
samp = []
for i in range(1000):
samp.append(np.random.normal())
plt.hist(samp, bins=100, normed=True);
def metropolis_hastings(N,burnin):
# Initial sample
x = np.zeros(2)
r = np.zeros(2)
# Compute p(x_initial) = p(x)
p = gmm(x[0],x[1],mu1,mu2,cov1,cov2)
p1 = gmm(x[0],x[1],mu1,mu2,cov1,cov2)
samples = []
samples1 = []
acc = 0
acc_ratio = []
# Collect every 10th sample
for i in range(N):
# Propose x^* ~ q(x*|x) = N(x,1) = x + N(0,1) * 1
#x_star = np.random.normal(loc = x,scale =1,size=2)
x_star = x + np.random.normal(size=2)
rn = r + np.random.normal(size=2)
# Compute p(x^*)
pn = gmm(rn[0],rn[1],mu1,mu2,cov1,cov2)
p_star = gmm(x_star[0],x_star[1],mu1,mu2,cov1,cov2)
# Compute pdf q(x|x*).pdf
q = gauss_2D(x,x_star,1)
# Compute q(x^*|x).pdf
q_star = gauss_2D(x_star,x,1)
# Accept or reject using U(0,1)
u = np.random.rand()
if pn >=p1:
p1 = pn
r = rn
elif u < pn/p1:
p1 = pn
r = rn
ratio = (p_star * q)/(p * q_star)
if u < min(1,ratio):
x = x_star
p = p_star
acc +=1
acc_ratio.append(acc/(i+1))
# keep every 10th sample
if i % 10==0:
samples.append(x)
samples1.append(r)
return [samples, samples1, acc_ratio]
# +
# Sample from the gmm using
'''Metropolis Hastings'''
[samples,samples1,acc_ratio] = metropolis_hastings(N = 100000, burnin = 10)
n = np.linspace(np.min(samples),np.max(samples),1000)
m = np.linspace(np.min(samples),np.max(samples),1000)
x,y = np.meshgrid(n,m)
fig, ax = plt.subplots(1, 4, figsize=(25,12))
# Compute PDFs
z = gmm(x,y,mu1,mu2,cov1,cov2)
# Plot target distribution (gmm) and MH samples together
samples = np.array(samples)
samples1 = np.array(samples1)
acc_ratio = np.array(acc_ratio)
ax[1].scatter(samples[:, 0], samples[:, 1], alpha=0.5, s=1)
ax[1].set_title('MH samples for GMM')
ax[3].plot(acc_ratio)
ax[3].set_title('Acceptance ratio for MH')
ax[2].scatter(samples1[:,0],samples1[:,1],alpha=0.5, s=1)
ax[2].set_title('MH samples without proposal')
CS = ax[0].contour(x,y,z);
ax[0].clabel(CS, inline=1, fontsize=10)
ax[0].set_title('GMM pdfs');
# -
# **After fitting the GMM, my samples are marginally better**
#
# ** For finite model selection you can use Bayesian or Akaike information criterion - BIC, AIC**
#
# **To do**: Autocorrelation - but you need a different example, won't work for this one.
#
# +
mix = mixture.GaussianMixture(n_components=2, covariance_type='full')
mix.fit(samples)
mix1 = mixture.GaussianMixture(n_components=2, covariance_type='full')
mix1.fit(samples1)
print('MH samples mean and covariance using proposal:\n {0}\n {1}'.format(mix.means_, mix.covariances_))
print('MH samples mean and covariance with symmetric proposal:\n {0}\n {1}'.format(mix1.means_, mix1.covariances_))
# -
# # Rejection sampling
# * Main and **very crucial** question: **How do you choose M and the proposal q?**
# - Especially since you don't know how to bound p (q has to essentially bound p)
# * Main condition: For $u \sim U(0,1)$ and $M$ a scaling constant, the proposal distribution satifies:
# * $ u * M * q(x^{i}) < p(x^i)$
# * We'll say $q = N(0,1)$ and $M = 2$ and stop when we have 1000 accepted samples. We'll plot the ratio as before.
# * It suffers from a main limitation, that in high dimensional space, the acceptance probability scales inversely with M - which means if M is large, you accept very little samples
def rejection_sampling(N, M):
acc = 0
samples = []
iterations = 0
acc_ratio = []
# Collect all samples that are accepted, no burn-in here
while acc<N:
# Sample from N(0,1) - could try N(x_previous,1) or something totally different
x = np.random.normal(loc=0,scale=2,size=2)
# Sample u ~ U(0,1)
u = np.random.rand()
# Compute p(x) and q(x,0,1)
p = gmm(x[0],x[1],mu1,mu2,cov1,cov2)
q = gauss_2D(x,[0,0],2)
#print(x,p,q)
if u < p/(M * q):
samples.append(x)
acc+=1
if acc%1000== 0:
print('{0} samples accepted'.format(acc))
iterations+=1
acc_ratio.append(acc/iterations)
return [samples, acc_ratio]
# +
# Sample from the gmm using
'''Rejection sampling'''
[samples,acc_ratio] = rejection_sampling(N = 10000, M = 2)
n = np.linspace(np.min(samples),np.max(samples),1000)
m = np.linspace(np.min(samples),np.max(samples),1000)
x,y = np.meshgrid(n,m)
fig, ax = plt.subplots(1, 3, figsize=(25,12))
# Compute PDFs
z = gmm(x,y,mu1,mu2,cov1,cov2)
# Plot target distribution (gmm) and rejection samples together
samples = np.array(samples)
acc_ratio = np.array(acc_ratio)
CS = ax[0].contour(x,y,z);
ax[0].clabel(CS, inline=1, fontsize=10)
ax[0].set_title('GMM pdfs');
ax[1].scatter(samples[:, 0], samples[:, 1], alpha=0.5, s=1);
ax[1].set_title('Rejection samples for GMM')
ax[2].plot(acc_ratio)
ax[2].set_title('Acceptance ratio for Rejection sampling');
# -
# # Importance sampling - Buggy
# # Sampling importance re-sampling
# * Simple idea, put more importance in sampling from regions of high density. Sounds ideal.
# * Rewrite $p(x) = w(x) * q(x)$. ** Again choice of q is done such that variance of the estimator is minimized. (what about bias?)**
# * Minimize $\Sigma_q(x)[f^2(x)w^2(x)]$ and after applying Jensen's inequality you get that the optimal q is lower bounded by:
# * $q^*(x) \propto |f(x)| p(x)$
# * This is all in the context of calculating expectations for $f(x)$ of course. It tells you that "efficient sampling occurs when you focus on sampling from $p(x)$ in the important regions where $|f(x)| p(x)$ is high". Turns out you can have situations where sampling from $q^*(x)$ can be more beneficial than sampling from $p(x)$ directly. Where you need to calculate expectations/integrals w.r.t $p(x)$
# * Note that AIS (Adaptive importance sampling) performs well in the case of the Boltzmann Machines when it comes to evaluating the partition function
# * **Update**: Two days later and 10 tutorials that have no examples, <NAME> enlightens me once again.." However, we do not directly get samples from p(x). To get samples from p(x), we must sample from the weighted sample from our importance
# sampler. This process is called Sampling Importance Re-sampling (SIR)"
# ** Because for some fucking reason everyone wants to calculate integrals. NO, it shouldn't be the case, an introductory thing should cover just the sampling part even if that's more complicated than the integral!!!!" **
a = np.ones(100,)
b = np.zeros((2,100))
np.dot(b,a)
def importance_sampling(N):
i = 1
samples = []
w = 0
normalized_w = 0
q = np.zeros(100)
# Collect all samples that are accepted, no burn-in here
while i<=N:
# Sample from N(0,1) - could try N(x_previous,1) or something totally different
x = np.random.normal(loc=0,scale=2,size=(2,100))
# Compute p(x) and q(x,0,1)
p = gmm(x[0,:],x[1,:],mu1,mu2,cov1,cov2)
for j in range(100):
q[j] = gauss_2D(x[:,j],[0,0],2)
# use w(x_i) as estimate for p(x_i) = w(x_i) * q(x_i)
# Re-sample x with prob proportional to normalized_w
w = p/q
#print(x.shape)
#print(w.shape)
val = np.dot(x,w)
normalized_w = val/100
#u = np.random.rand()
#if (u < w):
samples.append(normalized_w)
i+=1
return samples
# +
# Sample from the gmm using
'''Importance sampling'''
samples = importance_sampling(N = 500)
n = np.linspace(np.min(samples),np.max(samples),1000)
m = np.linspace(np.min(samples),np.max(samples),1000)
x,y = np.meshgrid(n,m)
fig, ax = plt.subplots(1, 2, figsize=(25,12))
# Compute PDFs
z = gmm(x,y,mu1,mu2,cov1,cov2)
# Plot target distribution (gmm) and importance samples together
samples = np.array(samples)
CS = ax[0].contour(x,y,z);
ax[0].clabel(CS, inline=1, fontsize=10)
ax[0].set_title('GMM pdfs');
ax[1].scatter(samples[:, 0], samples[:, 1], alpha=0.5, s=1);
ax[1].set_title('Importance samples for GMM');
#ax[2].plot(acc_ratio)
#ax[2].set_title('Acceptance ratio for Importance sampling');
# -
# # Hamiltonian Monte Carlo
# # Gibbs sampling
# See RBM
| sampling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Upload to flickr
# +
import os.path
import re
import sys
import tarfile
import time
import json
# import multiprocessing as mp
import itertools
from collections import Counter
# import tensorflow.python.platform
# from six.moves import urllib
import numpy as np
# import tensorflow as tf
# import h5py
import glob
import cPickle as pickle
# import matplotlib as mpl
# import matplotlib.pyplot as plt
# import matplotlib.image as mpimg
#from tensorflow.python.platform import gfile
import collections
#from run_inference import predict_star, predict
# import pandas as pd
# import seaborn as sns
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
# +
# need to map the 20 nearest neighbors also
# -
# ## Login and example of using Flickr API
import flickrapi
# +
api_key = u'<KEY>'
api_secret = u'<KEY>'
flickr = flickrapi.FlickrAPI(api_key, api_secret, format='json')
if not flickr.token_valid(perms=u'write'):
# Get a request token
flickr.get_request_token(oauth_callback='oob')
# Open a browser at the authentication URL. Do this however
# you want, as long as the user visits that URL.
authorize_url = flickr.auth_url(perms=u'write')
print authorize_url
# Get the verifier code from the user. Do this however you
# want, as long as the user gives the application the code.
verifier = unicode(raw_input('Verifier code: '))
# Trade the request token for an access token
flickr.get_access_token(verifier)
# -
flickr.photos.addTags(photo_id=11303139385, tags="metalworking")
flickr.photos.removeTag(photo_id=11303139385, tag_id="12383156-11303139385-335238")
old_tags = json.loads(flickr.tags.getListPhoto(photo_id=11303139385))["photo"]["tags"]["tag"]
for tag in old_tags:
print tag["_content"], tag["id"]
# ## Adding tags
# mapping from index to category
(image_metadata, book_metadata, image_to_idx) = pickle.load(open("/data/all_metadata_1M_tags.pkl", 'r'))
# mapping from index to URL
(idx_to_url, blid_to_url) = pickle.load(open("/data/image_to_url_mappings.pkl"))
len(idx_to_url.keys())
tag_files = glob.glob("/data/nearest_neighbor_tagging/tags/*_final.pkl")
for fn in tag_files:
print fn
tag_files = glob.glob("/data/nearest_neighbor_tagging/tags/*_final.pkl")
for fn in tag_files:
category = os.path.basename(fn)[:-10]
print category
if category == "people" or category == "animals":
category = "organism"
if category == "diagrams" or category == "architecture": continue
img_to_tag = pickle.load(open(fn, 'r'))
ctr = 0
for img in img_to_tag:
if ctr % 2000 == 0: print ctr
ctr += 1
flickr_id = int(idx_to_url[img])
if category == "organism" and ctr < 1000:
old_tags = json.loads(flickr.tags.getListPhoto(photo_id=flickr_id))["photo"]["tags"]["tag"]
for tag in old_tags:
if tag["_content"] == "sherlocknet:category=mammal":
flickr.photos.removeTag(photo_id=flickr_id, tag_id=tag["id"])
our_tags = ', '.join(["sherlocknet:tag=\"{}\"".format(tag[0]) for tag in img_to_tag[img]])
our_tags = our_tags + ', sherlocknet:category=\"{}\"'.format(category)
flickr.photos.addTags(photo_id=flickr_id, tags=our_tags)
verifier = str(raw_input('Verifier code: '))
print verifier
if category == "organism" and ctr < 1000:
old_tags = json.loads(flickr.tags.getListPhoto(photo_id=flickr_id))["photo"]["tags"]["tag"]
for tag in old_tags:
if tag["_content"] == "sherlocknet:category=mammal":
flickr.photos.removeTag(photo_id=flickr_id, tag_id=tag["id"])
our_tags = ', '.join(["sherlocknet:tag=\"{}\"".format(tag[0]) for tag in img_to_tag[img]])
our_tags = our_tags + ', sherlocknet:category=\"{}\"'.format(category)
flickr.photos.addTags(photo_id=flickr_id, tags=our_tags)
| scripts/.ipynb_checkpoints/upload_to_flickr-2-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="IdQi30u3pg-m"
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import Sequential
from tensorflow.keras.layers import Flatten, Dense, Dropout, BatchNormalization
from tensorflow.keras.layers import Conv1D, MaxPool1D
from tensorflow.keras.optimizers import Adam
print(tf.__version__)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="28brD1iNphJv" outputId="6373a68f-4121-49c1-b279-8b598464e654"
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
# + colab={} colab_type="code" id="YAnPhCEEphSi"
from sklearn import datasets, metrics
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
# -
cancer = datasets.load_breast_cancer()
print(cancer.DESCR)
X = pd.DataFrame(data=cancer.data, columns=cancer.feature_names)
X.head()
y = cancer.target
y
cancer.target_names
X.shape
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=0, stratify=y)
X_train.shape
X_test.shape
scaler = StandardScaler()
X_train = scaler.fit_transform(X_train)
X_test = scaler.transform(X_test)
X_train = X_train.reshape(455,30,1)
X_test = X_test.reshape(114,30,1)
# +
epochs = 50
model = Sequential()
model.add(Conv1D(filters=32,
kernel_size=2,
activation='relu',
input_shape=(30,1)))
model.add(BatchNormalization())
model.add(Dropout(0.2))
model.add(Conv1D(filters=64,
kernel_size=2,
activation='relu'))
model.add(BatchNormalization())
model.add(Dropout(0.5))
model.add(Flatten())
model.add(Dense(64, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(1, activation='sigmoid'))
# -
model.summary()
model.compile(optimizer=Adam(lr=0.00005), loss = 'binary_crossentropy', metrics=['accuracy'])
history = model.fit(X_train,
y_train,
epochs=epochs,
validation_data=(X_test, y_test),
verbose=1)
def plot_learningCurve(history, epoch):
# Plot training & validation accuracy values
epoch_range = range(1, epoch+1)
plt.plot(epoch_range, history.history['accuracy'])
plt.plot(epoch_range, history.history['val_accuracy'])
plt.title('Model accuracy')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(['Train', 'Val'], loc='upper left')
plt.show()
# Plot training & validation loss values
plt.plot(epoch_range, history.history['loss'])
plt.plot(epoch_range, history.history['val_loss'])
plt.title('Model loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train', 'Val'], loc='upper left')
plt.show()
history.history
plot_learningCurve(history, epochs)
| jupyter/TensorFlow/Deep Learning with TensorFlow 2.0 - Breast Cancer Detection Using CNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: haystack
# language: python
# name: haystack
# ---
# ## Task: Question Answering for Game of Thrones
#
# <img style="float: right;" src="https://upload.wikimedia.org/wikipedia/en/d/d8/Game_of_Thrones_title_card.jpg">
#
# Question Answering can be used in a variety of use cases. A very common one: Using it to navigate through complex knowledge bases or long documents ("search setting").
#
# A "knowledge base" could for example be your website, an internal wiki or a collection of financial reports.
# In this tutorial we will work on a slightly different domain: "Game of Thrones".
#
# Let's see how we can use a bunch of wikipedia articles to answer a variety of questions about the
# marvellous seven kingdoms...
#
# Let's start by adjust the working directory so that it is the root of the repository
# This should be run just once.
import os
os.chdir('../')
print("Current working directory is {}".format(os.getcwd()))
# + pycharm={"is_executing": false}
from haystack import Finder
from haystack.database.sql import SQLDocumentStore
from haystack.indexing.cleaning import clean_wiki_text
from haystack.indexing.io import write_documents_to_db, fetch_archive_from_http
from haystack.reader.farm import FARMReader
from haystack.reader.transformers import TransformersReader
from haystack.retriever.tfidf import TfidfRetriever
from haystack.utils import print_answers
# -
# ## Indexing & cleaning documents
# + pycharm={"is_executing": false}
# Let's first get some documents that we want to query
# Here: 517 Wikipedia articles for Game of Thrones
doc_dir = "data/article_txt_got"
s3_url = "https://s3.eu-central-1.amazonaws.com/deepset.ai-farm-qa/datasets/documents/wiki_gameofthrones_txt.zip"
fetch_archive_from_http(url=s3_url, output_dir=doc_dir)
# The documents can be stored in different types of "DocumentStores".
# For dev we suggest a light-weight SQL DB
# For production we suggest elasticsearch
document_store = SQLDocumentStore(url="sqlite:///qa.db")
# Now, let's write the docs to our DB.
# You can optionally supply a cleaning function that is applied to each doc (e.g. to remove footers)
# It must take a str as input, and return a str.
write_documents_to_db(document_store=document_store, document_dir=doc_dir, clean_func=clean_wiki_text, only_empty_db=True)
# -
# ## Initalize Reader, Retriever & Finder
# + pycharm={"is_executing": false, "name": "#%%\n"}
# A retriever identifies the k most promising chunks of text that might contain the answer for our question
# Retrievers use some simple but fast algorithm, here: TF-IDF
retriever = TfidfRetriever(document_store=document_store)
# + pycharm={"is_executing": false}
# A reader scans the text chunks in detail and extracts the k best answers
# Reader use more powerful but slower deep learning models
# You can select a local model or any of the QA models published on huggingface's model hub (https://huggingface.co/models)
# here: a medium sized BERT QA model trained via FARM on Squad 2.0
reader = FARMReader(model_name_or_path="deepset/bert-base-cased-squad2", use_gpu=False)
# OR: use alternatively a reader from huggingface's transformers package (https://github.com/huggingface/transformers)
# reader = TransformersReader(model="distilbert-base-uncased-distilled-squad", tokenizer="distilbert-base-uncased", use_gpu=-1)
# + pycharm={"is_executing": false}
# The Finder sticks together reader and retriever in a pipeline to answer our actual questions
finder = Finder(reader, retriever)
# -
# ## Voilà! Ask a question!
# + pycharm={"is_executing": false}
# You can configure how many candidates the reader and retriever shall return
# The higher top_k_retriever, the better (but also the slower) your answers.
prediction = finder.get_answers(question="Who is the father of <NAME>?", top_k_retriever=10, top_k_reader=5)
# +
#prediction = finder.get_answers(question="Who created the Dothraki vocabulary?", top_k_reader=5)
#prediction = finder.get_answers(question="Who is the sister of Sansa?", top_k_reader=5)
# + pycharm={"is_executing": false, "name": "#%%\n"}
print_answers(prediction, details="minimal")
# -
| tutorials/Tutorial1_Basic_QA_Pipeline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# #!/usr/bin/env python
# coding: utf-8
import rosbag, os, matplotlib, pickle
from matplotlib import pyplot as plt
from scipy.interpolate import interp1d
from qsrlib.qsrlib import QSRlib, QSRlib_Request_Message
from qsrlib_io.world_trace import Object_State, World_Trace
from qsrlib.qsrlib import QSR_QTC_BC_Simplified
import numpy as np
import pandas as pd
import datetime as dt
os.chdir("/home/loz/QTC_Trajectory_HMMs/from_bags/")
# +
# In[2]:
lab_bags = [os.path.join(dp, f) for dp, dn, fn in os.walk(os.path.expanduser("~/QTC_Trajectory_HMMs/from_bags/study_HRSI_rosbags")) for f in fn]
lab_bags
# +
# In[60]:
r_positions = []
h_positions = []
r_state_seqs = []
h_state_seqs = []
qsrlib = QSRlib()
for bag_path in lab_bags:
bag = rosbag.Bag(bag_path)
r_xs = []
r_ys = []
r_state_seq = []
for topic, msg, t in bag.read_messages(topics=['/robot5/control/odom']):
t = t.to_sec()
x = msg.pose.pose.position.x
y = msg.pose.pose.position.y
r_xs.append(x)
r_ys.append(y)
r_state_seq.append(Object_State(name="robot", timestamp=t, x=x, y=y))
r_state_seqs.append(r_state_seq)
h_xs = []
h_ys = []
h_state_seq = []
for topic, msg, t in bag.read_messages(topics=['/robot5/people_tracker_filtered/positions']):
t = t.to_sec()
try:
x = msg.poses[0].position.x
y = msg.poses[0].position.y
h_xs.append(x)
h_ys.append(y)
h_state_seq.append(Object_State(name="human", timestamp=t, x=x, y=y))
except:
pass
h_state_seqs.append(h_state_seq)
bag.close()
r_positions.append([r_xs, r_ys])
h_positions.append([h_xs, h_ys])
# In[61]:
# +
# Test getting QTC_C sequence
bag_no = 0
quantisation_factor = 0.01
world = World_Trace()
h_x = [h_state_seqs[bag_no][i].x for i in range(len(h_state_seqs[bag_no]))]
h_y = [h_state_seqs[bag_no][i].y for i in range(len(h_state_seqs[bag_no]))]
r_x = [r_state_seqs[bag_no][i].x for i in range(len(r_state_seqs[bag_no]))]
r_y = [r_state_seqs[bag_no][i].y for i in range(len(r_state_seqs[bag_no]))]
# +
# In[62]:
# Downsample state series' to 200kHz frequency
h_state_series = pd.DataFrame({"x": h_x, "y": h_y},
index=[pd.to_datetime(h_state_seqs[bag_no][i].timestamp, unit="s") for i in range(len(h_state_seqs[bag_no]))])
h_state_series = h_state_series.resample("200ms").mean()
# +
# In[63]:
r_state_series = pd.DataFrame({"x": r_x, "y": r_y},
index=[pd.to_datetime(r_state_seqs[bag_no][i].timestamp, unit="s") for i in range(len(r_state_seqs[bag_no]))])
r_state_series = r_state_series.resample("200ms").mean()
# +
# In[64]:
# Create world_trace state series from downsampled human position data
h_state_seq = []
for index, row in h_state_series.iterrows():
x = row['x']
y = row['y']
t = (pd.to_datetime(index) - dt.datetime(1970,1,1)).total_seconds()
h_state_seq.append(Object_State(name="human", timestamp=t, x=x, y=y))
# +
# In[65]:
# Create world_trace state series from downsampled robot position data
r_state_seq = []
for index, row in r_state_series.iterrows():
x = row['x']
y = row['y']
t = (pd.to_datetime(index) - dt.datetime(1970,1,1)).total_seconds()
r_state_seq.append(Object_State(name="robot", timestamp=t, x=x, y=y))
# +
# In[83]:
world.add_object_state_series(h_state_seq)
world.add_object_state_series(r_state_seq)
# make a QSRlib request message
dynamic_args = {"qtccs": {"no_collapse": True, "quantisation_factor": quantisation_factor,
"validate": False, "qsrs_for": [("human", "robot")]}}
qsrlib_request_message = QSRlib_Request_Message(
'qtccs', world, dynamic_args)
# request your QSRs
qsrlib_response_message = qsrlib.request_qsrs(req_msg=qsrlib_request_message)
qsrlib_response_message
# -
# Get QSR at each timestamp
timestamps = qsrlib_response_message.qsrs.get_sorted_timestamps()
for t in timestamps:
for val in qsrlib_response_message.qsrs.trace[t].qsrs.values():
print(val.qsr['qtccs'].replace(",",""))
# In[9]:
bag_path = lab_bags[bag_no]
# bag_path[67:].replace("/", "_")[:-4]
"_".join(bag_path[59:].replace("/", "_")[:-4].split("_")[:2])
# # Build dict of bags and their QTC_C sequences
# +
quantisation_factor = 0.01
qtc_seqs = {}
for bag_no in range(len(lab_bags)):
qtc_seq = []
bag_path = lab_bags[bag_no]
sit_code = "_".join(bag_path[59:].replace("/", "_")[:-4].split("_")[:2])
initial_bag_nos = range(1,6)
print(sit_code)
world = World_Trace()
h_x = [h_state_seqs[bag_no][i].x for i in range(len(h_state_seqs[bag_no]))]
h_y = [h_state_seqs[bag_no][i].y for i in range(len(h_state_seqs[bag_no]))]
r_x = [r_state_seqs[bag_no][i].x for i in range(len(r_state_seqs[bag_no]))]
r_y = [r_state_seqs[bag_no][i].y for i in range(len(r_state_seqs[bag_no]))]
# Downsample state series' to 200kHz frequency
h_state_series = pd.DataFrame({"x": h_x, "y": h_y},
index=[pd.to_datetime(h_state_seqs[bag_no][i].timestamp, unit="s") for i in range(len(h_state_seqs[bag_no]))])
h_state_series = h_state_series.resample("200ms").mean()
r_state_series = pd.DataFrame({"x": r_x, "y": r_y},
index=[pd.to_datetime(r_state_seqs[bag_no][i].timestamp, unit="s") for i in range(len(r_state_seqs[bag_no]))])
r_state_series = r_state_series.resample("200ms").mean()
with open("lab_sit_starts_ends.pickle", "r") as f:
starts_ends_ts = pickle.load(f)
# if int(sit_code.split("_")[-1]) in initial_bag_nos:
# h_state_series = h_state_series.loc[starts_ends_ts[sit_code][0]:starts_ends_ts[sit_code][1]]
# r_state_series = r_state_series.loc[starts_ends_ts[sit_code][0]:starts_ends_ts[sit_code][1]]
start = max(r_state_series.index.min(), h_state_series.index.min())
end = min(r_state_series.index.max(), h_state_series.index.max())
h_state_series = h_state_series.resample("200ms").interpolate()
r_state_series = r_state_series.resample("200ms").interpolate()
r_state_series = r_state_series.loc[start:end]
h_state_series = h_state_series.loc[start:end]
plt.plot(r_state_series.x.values, r_state_series.y.values)
plt.plot(h_state_series.x.values, h_state_series.y.values)
plt.show()
raw_input()
plt.close()
# Create world_trace state series from downsampled human position data
h_state_seq = []
for index, row in h_state_series.iterrows():
x = row['x']
y = row['y']
t = (pd.to_datetime(index) - dt.datetime(1970,1,1)).total_seconds()
h_state_seq.append(Object_State(name="human", timestamp=t, x=x, y=y))
# Create world_trace state series from downsampled robot position data
r_state_seq = []
for index, row in r_state_series.iterrows():
x = row['x']
y = row['y']
t = (pd.to_datetime(index) - dt.datetime(1970,1,1)).total_seconds()
r_state_seq.append(Object_State(name="robot", timestamp=t, x=x, y=y))
# Add human and robot trajectories to world
world.add_object_state_series(h_state_seq)
world.add_object_state_series(r_state_seq)
# make a QSRlib request message
dynamic_args = {"qtccs": {"no_collapse": False, "quantisation_factor": quantisation_factor,
"validate": False, "qsrs_for": [("robot", "human")]}}
qsrlib_request_message = QSRlib_Request_Message(
'qtccs', world, dynamic_args)
# request your QSRs
qsrlib_response_message = qsrlib.request_qsrs(req_msg=qsrlib_request_message)
qsrlib_response_message
# Get QSR at each timestamp
timestamps = qsrlib_response_message.qsrs.get_sorted_timestamps()
for t in timestamps:
for val in qsrlib_response_message.qsrs.trace[t].qsrs.values():
qtc_seq.append(val.qsr['qtccs'].replace(",",""))
qtc_seqs[sit_code] = qtc_seq
# -
| from_bags/qtcs_from_bags_py2/plot_study_bags.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
age = input("How old are you? ")
age = int(age)
if age > 65:
print("Enjoy your retirement!")
print("You handsome!")
elif age > 25:
print("I hope you have a job :D")
elif age > 6:
print("Study hard, get good grades, listen to your mom")
else:
print("You're so biiig and cuuute")
age > 1
age < 1
age == 1
age >= 1
age <= 1
1 <= age < 100
1 <= age and age < 100
1 < age or age < 100
number_of_times_to_say_hello = 10
number_of_times_i_said_hello = 0
while number_of_times_i_said_hello < number_of_times_to_say_hello:
print("hello there")
number_of_times_i_said_hello = number_of_times_i_said_hello + 1
| Lesson 4 - Control Flow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Thyroid disease prediction simple ann
from PIL import Image
import cv2
image=Image.open("C:/Users/U/Downloads/thyroid prediction in simple ann/thyroid.jpg")
fig=plt.imshow(image)
fig.axes.get_xaxis().set_visible(False)
fig.axes.get_yaxis().set_visible(False)
print("The most common thyroid disorder is hypothyroidism. Hypo- means deficient or under(active), so hypothyroidism is a condition in which the thyroid gland is underperforming or producing too little thyroid hormone.. Recognizing the symptoms of hypothyroidism is extremely important.")
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
data=pd.read_csv("hypothyroid.csv")
data.head()
data.tail()
data.shape
data.describe(include="all")
data.nunique()
data.columns
data.binaryClass.value_counts()
data.binaryClass=data.binaryClass.map({"P":1,"N":0}).astype(int)
data.sex.value_counts()
data=data.replace({"?":np.NAN})
data.sex.value_counts()
data.pregnant.value_counts()
data.pregnant=data.pregnant.replace({"f":0,"t":1})
data.isnull().sum()
data.dtypes
data=data.replace({"t":1,"f":0})
data.head()
data["TBG"].value_counts()
del data["TBG"]
data["referral source"].value_counts()
del data["referral source"]
data.info()
data["T3 measured"].value_counts()
data["TBG measured"].value_counts()
data["on thyroxine"].value_counts()
data.dtypes
data.isnull().sum()
data.head()
colums=data.columns[data.dtypes.eq("object")]
data[columns]=data[columns].apply(pd.to_numeric,errors="coerce")
data.dtypes
data.isnull().sum()
data.fillna(data.mean())
data.dtypes
data.isnull().sum()
data.sex.value_counts()
data.isna().sum()
data["age"].fillna(data["age"].mean(),inplace=True)
data.sex=data.sex.replace({"F":1,"M":0})
data["sex"].fillna(data["sex"].mode(),inplace=True)
from sklearn.impute import SimpleImputer
imputer=SimpleImputer(strategy="mean")
data["TSH"]=imputer.fit_transform(data[["TSH"]])
data["T3"]=imputer.fit_transform(data[["T3"]])
data["TT4"]=imputer.fit_transform(data[["TT4"]])
data["T4U"]=imputer.fit_transform(data[["T4U"]])
data["FTI"]=imputer.fit_transform(data[["FTI"]])
data.isnull().sum()
data.sex=data.sex.fillna(0)
data2.to_csv("thyroid.csv")
sns.heatmap(data.isnull())
plt.hist(data.sex)
plt.plot(data["on thyroxine"],data["pregnant"])
sns.set(rc={"figure.figsize":[8,8]},font_scale=1.2)
sns.distplot(data["age"])
sns.distplot(data.sex)
sns.distplot(data.T3)
sns.distplot(data['FTI'])
sns.jointplot(x="pregnant",y="age",data=data,kind="scatter")
sns.jointplot(x="age",y="pregnant",data=data,kind="reg")
sns.jointplot(x="age",y="TT4",data=data,kind="scatter")
sns.countplot(x="binaryClass",data=data)
sns.countplot(x="binaryClass",hue="sex",data=data)
sns.boxplot(x="binaryClass",y="age",data=data)
data1=data[data.age<=(data.age.mean()+3*data.age.std())]
data1.shape
data1.to_csv("thyroid.csv")
sns.boxplot(x="binaryClass",y="age",data=data1)
data1.corr()
sns.heatmap(data1.corr())
x=data1.drop("binaryClass",axis=1)
y=data1.binaryClass
x
x.info()
x.isna().sum()
y
import statsmodels.api as sm
x=sm.add_constant(x)
result=sm.OLS(y,x).fit()
result.summary()
# # THIS IS ALL ABOUT EDA....MODEL BUILDING IS DONE VIA ANOTHER .py NAMED (thyroid_prediction.py) FILE GO CHECK IT OUT
| notebooks/thyroid disease prediction simple ann.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import pandas as pd
from sklearn import datasets
from sklearn import linear_model
from sklearn.cross_validation import train_test_split
import matplotlib.pyplot as plt
from sklearn.model_selection import KFold
import sklearn.metrics
import math
df=pd.read_csv("train_day.csv", header=0)
data = pd.DataFrame(df, columns = ['date','store_nbr','item_nbr','onpromotion','family','class','perishable','transactions','city','state','type','cluster','transferred','dcoilwtico','yea','mon','day']).as_matrix()
target = pd.DataFrame(df,columns=['unit_sales']).as_matrix()
model = linear_model.PassiveAggressiveRegressor()
# +
#X_train, X_test, y_train, y_test = train_test_split(data, target, test_size=0.2, random_state=42)
kf = KFold(n_splits=5)
X=data
y=target
def NWRMSLE(y, pred, w):
return sklearn.metrics.mean_squared_error(y_test, pred,sample_weight=w)**0.5
error=0
for train_index, test_index in kf.split(X):
X_train, X_test = X[train_index], X[test_index]
y_train, y_test = y[train_index], y[test_index]
mymodel = model.fit(X_train, y_train)
pred=mymodel.predict(X_test)
error+=NWRMSLE(y_test,pred,X_test[:,10])
error/=5
print error
# -
fig, ax = plt.subplots()
x_=y_test
y_=pred
ax.scatter(x_,y_,color='red')
ax.plot([0,y_.max()], [0,y_.max()],linewidth=3,color='black')
ax.set_xlabel('Data')
ax.set_ylabel('Predicted')
plt.title('Passive Aggressive Regressor')
plt.show()
# +
extra1=0
extra2=0
sum=0
for i in range(len(x_)):
sum+=x_[i]
avg=sum/len(x_)
print "Total units: ",sum
print "Average units: ",avg
for i in range(len(x_)):
extra1+=abs(x_[i]-avg)
extra2+=abs(x_[i]-y_[i])
print extra1,extra2
| PassiveAggressive.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="SB93Ge748VQs"
# ##### Copyright 2019 The TensorFlow Authors.
# + cellView="form" colab={} colab_type="code" id="0sK8X2O9bTlz"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="HEYuO5NFwDK9"
# # Get started with TensorBoard
#
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/tensorboard/get_started"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/tensorboard/blob/master/docs/get_started.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/tensorboard/blob/master/docs/get_started.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="56V5oun18ZdZ"
# In machine learning, to improve something you often need to be able to measure it. TensorBoard is a tool for providing the measurements and visualizations needed during the machine learning workflow. It enables tracking experiment metrics like loss and accuracy, visualizing the model graph, projecting embeddings to a lower dimensional space, and much more.
#
# This quickstart will show how to quickly get started with TensorBoard. The remaining guides in this website provide more details on specific capabilities, many of which are not included here.
# + colab={"base_uri": "https://localhost:8080/", "height": 85} colab_type="code" id="6B95Hb6YVgPZ" outputId="9f27f106-41e8-40bd-f73d-6abbb1314643"
try:
# # %tensorflow_version only exists in Colab.
# %tensorflow_version 2.x
except Exception:
pass
# Load the TensorBoard notebook extension
# %load_ext tensorboard
# + colab={} colab_type="code" id="_wqSAZExy6xV"
import tensorflow as tf
import datetime
# + colab={} colab_type="code" id="Ao7fJW1Pyiza"
# Clear any logs from previous runs
# !rm -rf ./logs/
# + [markdown] colab_type="text" id="z5pr9vuHVgXY"
# Using the [MNIST](https://en.wikipedia.org/wiki/MNIST_database) dataset as the example, normalize the data and write a function that creates a simple Keras model for classifying the images into 10 classes.
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="j-DHsby18cot" outputId="4c910ebd-2c93-4138-fec7-0a36ae3d90bf"
mnist = tf.keras.datasets.mnist
(x_train, y_train),(x_test, y_test) = mnist.load_data()
x_train, x_test = x_train / 255.0, x_test / 255.0
def create_model():
return tf.keras.models.Sequential([
tf.keras.layers.Flatten(input_shape=(28, 28)),
tf.keras.layers.Dense(512, activation='relu'),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(10, activation='softmax')
])
# + [markdown] colab_type="text" id="XKUjdIoV87um"
# ## Using TensorBoard with Keras Model.fit()
# + [markdown] colab_type="text" id="8CL_lxdn8-Sv"
# When training with Keras's [Model.fit()](https://www.tensorflow.org/api_docs/python/tf/keras/models/Model#fit), adding the `tf.keras.callbacks.TensorBoard` callback ensures that logs are created and stored. Additionally, enable histogram computation every epoch with `histogram_freq=1` (this is off by default)
#
# Place the logs in a timestamped subdirectory to allow easy selection of different training runs.
# + colab={"base_uri": "https://localhost:8080/", "height": 221} colab_type="code" id="WAQThq539CEJ" outputId="73a348aa-504c-48a7-f720-5dd0478e2aa9"
model = create_model()
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
log_dir = "logs/fit/" + datetime.datetime.now().strftime("%Y%m%d-%H%M%S")
tensorboard_callback = tf.keras.callbacks.TensorBoard(log_dir=log_dir, histogram_freq=1)
model.fit(x=x_train,
y=y_train,
epochs=5,
validation_data=(x_test, y_test),
callbacks=[tensorboard_callback])
# + [markdown] colab_type="text" id="asjGpmD09dRl"
# Start TensorBoard through the command line or within a notebook experience. The two interfaces are generally the same. In notebooks, use the `%tensorboard` line magic. On the command line, run the same command without "%".
# + colab={} colab_type="code" id="A4UKgTLb9fKI"
# %tensorboard --logdir logs/fit
# + [markdown] colab_type="text" id="MCsoUNb6YhGc"
# <img class="tfo-display-only-on-site" src="https://github.com/tensorflow/tensorboard/blob/master/docs/images/quickstart_model_fit.png?raw=1"/>
# + [markdown] colab_type="text" id="Gi4PaRm39of2"
# A brief overview of the dashboards shown (tabs in top navigation bar):
#
# * The **Scalars** dashboard shows how the loss and metrics change with every epoch. You can use it to also track training speed, learning rate, and other scalar values.
# * The **Graphs** dashboard helps you visualize your model. In this case, the Keras graph of layers is shown which can help you ensure it is built correctly.
# * The **Distributions** and **Histograms** dashboards show the distribution of a Tensor over time. This can be useful to visualize weights and biases and verify that they are changing in an expected way.
#
# Additional TensorBoard plugins are automatically enabled when you log other types of data. For example, the Keras TensorBoard callback lets you log images and embeddings as well. You can see what other plugins are available in TensorBoard by clicking on the "inactive" dropdown towards the top right.
# + [markdown] colab_type="text" id="nB718NOH95yG"
# ## Using TensorBoard with other methods
# + [markdown] colab_type="text" id="IKNt0nWs-Ekt"
# When training with methods such as [`tf.GradientTape()`](https://www.tensorflow.org/api_docs/python/tf/GradientTape), use `tf.summary` to log the required information.
#
# Use the same dataset as above, but convert it to `tf.data.Dataset` to take advantage of batching capabilities:
# + colab={} colab_type="code" id="nnHx4DsMezy1"
train_dataset = tf.data.Dataset.from_tensor_slices((x_train, y_train))
test_dataset = tf.data.Dataset.from_tensor_slices((x_test, y_test))
train_dataset = train_dataset.shuffle(60000).batch(64)
test_dataset = test_dataset.batch(64)
# + [markdown] colab_type="text" id="SzpmTmJafJ10"
# The training code follows the [advanced quickstart](https://www.tensorflow.org/tutorials/quickstart/advanced) tutorial, but shows how to log metrics to TensorBoard. Choose loss and optimizer:
# + colab={} colab_type="code" id="H2Y5-aPbAANs"
loss_object = tf.keras.losses.SparseCategoricalCrossentropy()
optimizer = tf.keras.optimizers.Adam()
# + [markdown] colab_type="text" id="cKhIIDj9Hbfy"
# Create stateful metrics that can be used to accumulate values during training and logged at any point:
# + colab={} colab_type="code" id="jD0tEWrgH0TL"
# Define our metrics
train_loss = tf.keras.metrics.Mean('train_loss', dtype=tf.float32)
train_accuracy = tf.keras.metrics.SparseCategoricalAccuracy('train_accuracy')
test_loss = tf.keras.metrics.Mean('test_loss', dtype=tf.float32)
test_accuracy = tf.keras.metrics.SparseCategoricalAccuracy('test_accuracy')
# + [markdown] colab_type="text" id="szw_KrgOg-OT"
# Define the training and test functions:
# + colab={} colab_type="code" id="TTWcJO35IJgK"
def train_step(model, optimizer, x_train, y_train):
with tf.GradientTape() as tape:
predictions = model(x_train, training=True)
loss = loss_object(y_train, predictions)
grads = tape.gradient(loss, model.trainable_variables)
optimizer.apply_gradients(zip(grads, model.trainable_variables))
train_loss(loss)
train_accuracy(y_train, predictions)
def test_step(model, x_test, y_test):
predictions = model(x_test)
loss = loss_object(y_test, predictions)
test_loss(loss)
test_accuracy(y_test, predictions)
# + [markdown] colab_type="text" id="nucPZBKPJR3A"
# Set up summary writers to write the summaries to disk in a different logs directory:
# + colab={} colab_type="code" id="3Qp-exmbWf4w"
current_time = datetime.datetime.now().strftime("%Y%m%d-%H%M%S")
train_log_dir = 'logs/gradient_tape/' + current_time + '/train'
test_log_dir = 'logs/gradient_tape/' + current_time + '/test'
train_summary_writer = tf.summary.create_file_writer(train_log_dir)
test_summary_writer = tf.summary.create_file_writer(test_log_dir)
# + [markdown] colab_type="text" id="qgUJgDdKWUKF"
# Start training. Use `tf.summary.scalar()` to log metrics (loss and accuracy) during training/testing within the scope of the summary writers to write the summaries to disk. You have control over which metrics to log and how often to do it. Other `tf.summary` functions enable logging other types of data.
# + colab={"base_uri": "https://localhost:8080/", "height": 102} colab_type="code" id="odWvHPpKJvb_" outputId="cff49dc7-2c87-4ea0-f36f-f0af948d7a65"
model = create_model() # reset our model
EPOCHS = 5
for epoch in range(EPOCHS):
for (x_train, y_train) in train_dataset:
train_step(model, optimizer, x_train, y_train)
with train_summary_writer.as_default():
tf.summary.scalar('loss', train_loss.result(), step=epoch)
tf.summary.scalar('accuracy', train_accuracy.result(), step=epoch)
for (x_test, y_test) in test_dataset:
test_step(model, x_test, y_test)
with test_summary_writer.as_default():
tf.summary.scalar('loss', test_loss.result(), step=epoch)
tf.summary.scalar('accuracy', test_accuracy.result(), step=epoch)
template = 'Epoch {}, Loss: {}, Accuracy: {}, Test Loss: {}, Test Accuracy: {}'
print (template.format(epoch+1,
train_loss.result(),
train_accuracy.result()*100,
test_loss.result(),
test_accuracy.result()*100))
# Reset metrics every epoch
train_loss.reset_states()
test_loss.reset_states()
train_accuracy.reset_states()
test_accuracy.reset_states()
# + [markdown] colab_type="text" id="JikosQ84fzcA"
# Open TensorBoard again, this time pointing it at the new log directory. We could have also started TensorBoard to monitor training while it progresses.
# + colab={} colab_type="code" id="-Iue509kgOyE"
# %tensorboard --logdir logs/gradient_tape
# + [markdown] colab_type="text" id="NVpnilhEgQXk"
# <img class="tfo-display-only-on-site" src="https://github.com/tensorflow/tensorboard/blob/master/docs/images/quickstart_gradient_tape.png?raw=1"/>
# + [markdown] colab_type="text" id="ozbwXgPIkCKV"
# That's it! You have now seen how to use TensorBoard both through the Keras callback and through `tf.summary` for more custom scenarios.
| docs/get_started.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from fastai.vision.all import *
import kornia
#export
def get_aug_pipe(size, min_scale=0.2, max_scale=1., stats=imagenet_stats, s=.6,
color=True, xtra_tfms=[]):
"SimCLR augmentations"
tfms = []
tfms += [kornia.augmentation.RandomResizedCrop((size, size),
scale=(min_scale, max_scale),
ratio=(3/4, 4/3))]
tfms += [kornia.augmentation.RandomHorizontalFlip()]
if color: tfms += [kornia.augmentation.ColorJitter(0.8*s, 0.8*s, 0.8*s, 0.2*s)]
if color: tfms += [kornia.augmentation.RandomGrayscale(p=0.2)]
if stats is not None: tfms += [Normalize.from_stats(*stats)]
tfms += xtra_tfms
pipe = Pipeline(tfms)
pipe.split_idx = 0
return pipe
#export
def create_encoder(arch, n_in=3, pretrained=True, cut=None, concat_pool=True):
"Create encoder from a given arch backbone"
encoder = create_body(arch, n_in, pretrained, cut)
pool = AdaptiveConcatPool2d() if concat_pool else nn.AdaptiveAvgPool2d(1)
return nn.Sequential(*encoder, pool, Flatten())
#export
class MLP(Module):
"MLP module as described in paper"
def __init__(self, dim, projection_size=256, hidden_size=2048):
self.net = nn.Sequential(
nn.Linear(dim, hidden_size),
nn.BatchNorm1d(hidden_size),
nn.ReLU(inplace=True),
nn.Linear(hidden_size, projection_size)
)
def forward(self, x):
return self.net(x)
#export
class SwAVModel(Module):
def __init__(self,encoder,projector,prototypes):
self.encoder,self.projector,self.prototypes = encoder,projector,prototypes
def forward(self, inputs):
if not isinstance(inputs, list): inputs = [inputs]
crop_idxs = torch.cumsum(torch.unique_consecutive(
torch.tensor([inp.shape[-1] for inp in inputs]),
return_counts=True)[1], 0)
start_idx = 0
for idx in crop_idxs:
_z = self.encoder(torch.cat(inputs[start_idx: idx]))
if not start_idx: z = _z
else: z = torch.cat((z, _z))
start_idx = idx
z = F.normalize(self.projector(z))
return z, self.prototypes(z)
#export
def create_swav_model(arch=resnet50, n_in=3, pretrained=True, cut=None, concat_pool=True,
hidden_size=256, projection_size=128, n_protos=3000):
"Create SwAV from a given arch"
encoder = create_encoder(arch, n_in, pretrained, cut, concat_pool)
with torch.no_grad(): representation = encoder(torch.randn((2,n_in,128,128)))
projector = MLP(representation.size(1), projection_size, hidden_size=hidden_size)
prototypes = nn.Linear(projection_size, n_protos, bias=False)
apply_init(projector)
with torch.no_grad():
w = prototypes.weight.data.clone()
prototypes.weight.copy_(F.normalize(w))
return SwAVModel(encoder, projector, prototypes)
#export
def sinkhorn_knopp(Q, nmb_iters, device=default_device):
"https://en.wikipedia.org/wiki/Sinkhorn%27s_theorem#Sinkhorn-Knopp_algorithm"
with torch.no_grad():
sum_Q = torch.sum(Q)
Q /= sum_Q
r = (torch.ones(Q.shape[0]) / Q.shape[0]).to(device)
c = (torch.ones(Q.shape[1]) / Q.shape[1]).to(device)
curr_sum = torch.sum(Q, dim=1)
for it in range(nmb_iters):
u = curr_sum
Q *= (r / u).unsqueeze(1)
Q *= (c / torch.sum(Q, dim=0)).unsqueeze(0)
curr_sum = torch.sum(Q, dim=1)
return (Q / torch.sum(Q, dim=0, keepdim=True)).t().float()
class SWAVLoss(Module):
def __init__(self): pass
def forward(self,log_ps,qs):
"Multi crop loss"
loss = 0
for i in range(len(qs)):
l = 0
q = qs[i]
for p in (log_ps[:i] + log_ps[i+1:]):
l -= torch.mean(torch.sum(q*p, dim=1))/(len(log_ps)-1)
loss += l/len(qs)
return loss
#export
class SWAV(Callback):
def __init__(self, crop_sizes=[224,96],
num_crops=[2,6],
min_scales=[0.25,0.05],
max_scales=[1.,0.14],
crop_assgn_ids=[0,1],
eps=0.05,
n_sinkh_iter=3,
temp=0.1,
**aug_kwargs):
store_attr('num_crops,crop_assgn_ids,temp,eps,n_sinkh_iter')
self.augs = []
for nc, size, mins, maxs in zip(num_crops, crop_sizes, min_scales, max_scales):
self.augs += [get_aug_pipe(size, mins, maxs, **aug_kwargs) for i in range(nc)]
def before_batch(self):
"Compute multi crop inputs"
self.bs = self.x.size(0)
self.learn.xb = ([aug(self.x) for aug in self.augs],)
def after_pred(self):
"Compute ps and qs"
embedding, output = self.pred
with torch.no_grad():
qs = []
for i in self.crop_assgn_ids:
# TODO: Store previous batch embeddings
# to be used in Q calculation
# Store approx num_proto//bs batches
# output.size(1)//self.bs
target_b = output[self.bs*i:self.bs*(i+1)]
q = torch.exp(target_b/self.eps).t()
q = sinkhorn_knopp(q, self.n_sinkh_iter, q.device)
qs.append(q)
log_ps = []
for v in np.arange(np.sum(self.num_crops)):
log_p = F.log_softmax(output[self.bs*v:self.bs*(v+1)] / self.temp, dim=1)
log_ps.append(log_p)
self.learn.pred, self.learn.yb = log_ps, (qs,)
def after_batch(self):
with torch.no_grad():
w = learn.model.prototypes.weight.data.clone()
learn.model.prototypes.weight.data.copy_(F.normalize(w))
def show_one(self):
xb = self.learn.xb[0]
i = np.random.choice(self.bs)
images = [aug.normalize.decode(b.to('cpu').clone()).clamp(0.1)[i]
for b, aug in zip(xb, self.augs)]
show_images(images)
# ## Pretext Training
sqrmom=0.99
mom=0.95
beta=0.
eps=1e-4
opt_func = partial(ranger, mom=mom, sqr_mom=sqrmom, eps=eps, beta=beta)
def get_dls(size, bs, workers=None):
path = URLs.IMAGEWANG_160 if size <= 160 else URLs.IMAGEWANG
source = untar_data(path)
files = get_image_files(source)
tfms = [[PILImage.create, ToTensor, RandomResizedCrop(size, min_scale=0.9)],
[parent_label, Categorize()]]
dsets = Datasets(files, tfms=tfms, splits=RandomSplitter(valid_pct=0.1)(files))
batch_tfms = [IntToFloatTensor]
dls = dsets.dataloaders(bs=bs, num_workers=workers, after_batch=batch_tfms)
return dls
bs=64
resize, size = 256, 224
dls = get_dls(resize, bs)
model = create_swav_model(arch=xresnet34, n_in=3, pretrained=False)
learn = Learner(dls, model, SWAVLoss(),
cbs=[SWAV(crop_sizes=[size,96],
num_crops=[2,6],
min_scales=[0.25,0.2],
max_scales=[1.0,0.35]),
TerminateOnNaNCallback()])
b = dls.one_batch()
learn._split(b)
learn('before_batch')
learn.swav.show_one()
# +
# learn.to_fp16();
# -
learn.lr_find()
lr=1e-2
wd=1e-2
epochs=100
learn.unfreeze()
learn.fit_flat_cos(epochs, lr, wd=wd, pct_start=0.5)
save_name = f'swav_iwang_sz{size}_epc100'; save_name
learn.save(save_name)
torch.save(learn.model.encoder.state_dict(), learn.path/learn.model_dir/f'{save_name}_encoder.pth')
learn.recorder.losses
# ## Downstream Task
def get_dls(size, bs, workers=None):
path = URLs.IMAGEWANG_160 if size <= 160 else URLs.IMAGEWANG
source = untar_data(path)
files = get_image_files(source, folders=['train', 'val'])
splits = GrandparentSplitter(valid_name='val')(files)
item_aug = [RandomResizedCrop(size, min_scale=0.35), FlipItem(0.5)]
tfms = [[PILImage.create, ToTensor, *item_aug],
[parent_label, Categorize()]]
dsets = Datasets(files, tfms=tfms, splits=splits)
batch_tfms = [IntToFloatTensor, Normalize.from_stats(*imagenet_stats)]
dls = dsets.dataloaders(bs=bs, num_workers=workers, after_batch=batch_tfms)
return dls
def do_train(epochs=5, runs=5, size=size, bs=bs, lr=lr, save_name=None):
dls = get_dls(size, bs)
for run in range(runs):
print(f'Run: {run}')
learn = cnn_learner(dls, xresnet34, opt_func=opt_func, normalize=False,
metrics=[accuracy,top_k_accuracy], loss_func=LabelSmoothingCrossEntropy(),
pretrained=False)
if save_name is not None:
state_dict = torch.load(learn.path/learn.model_dir/f'{save_name}_encoder.pth')
learn.model[0].load_state_dict(state_dict)
print("Model loaded...")
learn.unfreeze()
learn.fit_flat_cos(epochs, lr, wd=wd)
# ### 5 epochs
lr = 1e-2
epochs = 5
runs = 5
do_train(epochs, runs, lr=lr, save_name=save_name)
np.mean([0.737592, 0.727412, 0.720540,0.737592,0.735556])
# ### 20 epochs
epochs = 20
runs = 3
do_train(epochs, runs, save_name=save_name)
np.mean([0.763553, 0.768134, 0.763299])
# ### 80 epochs
epochs = 80
runs = 1
do_train(epochs, runs, save_name=save_name)
# ### 200 epochs
epochs = 200
runs = 1
do_train(epochs, runs, save_name=save_name)
| nbs/examples/swav_iwang_224.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/naagarjunsa/data-science-portfolio/blob/main/deep-learning/binary_classification_imdb.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="sjqkcXxwvRAl" outputId="9364efd2-2d4e-471b-adbe-2a7b3f47d213"
from keras.datasets import imdb
(train_data, train_labels), (test_data, test_labels) = imdb.load_data(num_words=1000)
# + colab={"base_uri": "https://localhost:8080/"} id="cjy0UJoCLlOY" outputId="8c396392-aba6-475a-fdf4-86130c62b706"
train_data[0]
# + colab={"base_uri": "https://localhost:8080/"} id="DsADl9EOLxLt" outputId="697ee5b7-4105-47b4-bb31-0b37010414c2"
train_labels[0]
# + id="EcQdntqIL0K9"
word_index = imdb.get_word_index()
reverse_word_index = dict ([(value, key) for (key, value) in word_index.items()])
decoded_review = ' '.join([reverse_word_index.get(i-3,'?') for i in train_data[0]])
# + colab={"base_uri": "https://localhost:8080/", "height": 188} id="UZyz98OeMP8N" outputId="dad92278-fe9e-47c0-ae9b-fd8bc606a81b"
decoded_review
# + colab={"base_uri": "https://localhost:8080/"} id="4cUGWEEeMT49" outputId="2ab21feb-7d2c-4855-8f31-3174ce382a64"
import numpy as np
def vectorize_sequences(sequences, dimension=10000):
results = np.zeros((len(sequences), dimension))
for i, sequence in enumerate(sequences):
results[i, sequence] = 1.
return results
x_train = vectorize_sequences(train_data)
x_test = vectorize_sequences(test_data)
x_train[0]
# + colab={"base_uri": "https://localhost:8080/"} id="W5uIhlwxNTbW" outputId="51cee939-f677-4dbf-e2c3-306af41a39b8"
y_train = np.asarray(train_labels).astype('float32')
y_test = np.asarray(test_labels).astype('float32')
y_train[0]
# + id="YZ8v47D5SQug"
from keras import models
from keras import layers
model = models.Sequential()
model.add(layers.Dense(16, activation='relu', input_shape=(10000,)))
model.add(layers.Dense(16, activation='relu'))
model.add(layers.Dense(1, activation='sigmoid'))
# + id="QYSHR7vnTDxd"
from keras import optimizers
from keras import losses
from keras import metrics
model.compile(optimizer=optimizers.RMSprop(lr=0.01),
loss= losses.binary_crossentropy,
metrics=[metrics.binary_accuracy])
# + id="D1VYUI6jTdpV"
x_val = x_train[:10000]
partial_x_train = x_train[10000:]
y_val = y_train[:10000]
partial_y_train = y_train[10000:]
# + colab={"base_uri": "https://localhost:8080/"} id="yegBlFQKUMPI" outputId="3b9984e9-4aee-4403-93fd-cc4bcf2c35ae"
history = model.fit(partial_x_train,
partial_y_train,
batch_size = 256,
epochs=4,
verbose=1,
validation_data=(x_val, y_val))
# + colab={"base_uri": "https://localhost:8080/"} id="7UhMoDwhUlr9" outputId="f173547f-bcac-481e-84bb-8dfb61f1641c"
history_dict = history.history
history_dict.keys()
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="orMm9WZvU0St" outputId="9c08a4c7-93b4-45c6-faef-5713dfef9232"
import matplotlib.pyplot as plt
loss_values = history_dict['loss']
val_loss_values = history_dict['val_loss']
acc = history_dict['binary_accuracy']
val_acc = history_dict['val_binary_accuracy']
epochs = range(1, len(acc) + 1)
plt.plot(epochs, loss_values, 'bo', label = 'Training Loss')
plt.plot(epochs, val_loss_values, 'b', label = 'Validation Loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="XzaThbe-VTGG" outputId="023cb775-377b-4bf7-dcb4-81cc1ee790c1"
model = models.Sequential()
model.add(layers.Dense(16, activation='relu', input_shape=(10000,)))
model.add(layers.Dense(16, activation='relu'))
model.add(layers.Dense(1, activation='sigmoid'))
model.compile(optimizer='rmsprop',
loss='binary_crossentropy',
metrics=['accuracy'])
model.fit(x_train, y_train, epochs=4, batch_size=512, verbose=1)
results = model.evaluate(x_test, y_test)
# + colab={"base_uri": "https://localhost:8080/"} id="98PrVFwrW1Bb" outputId="87ce5a26-c917-41b3-d754-f14aad332ed8"
results
# + id="TVMUihhpW1qW"
| deep-learning/binary_classification_imdb.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import speech_recognition as sr
import webbrowser
r1=sr.Recognizer()
r2=sr.Recognizer()
r3 = sr.Recognizer()
with sr.Microphone() as source:
print('[search edureka:search youtube]')
print('speak now')
audio=r3.listen(source)
if 'edureka' in r2.recognize_google(audio):
r2=sr.Recognizer()
url='https://www.edureka.co/'
with sr.Microphone() as source:
print('Search your Query')
audio =r2.listen(source)
try:
get=r2.recognize_google(audio)
print(get)
wb.get().open_new(url+get)
except sr.UnknownValueError:
print('Error')
except sr.RequestError as e:
print('Failed'.format(e))
# -
| Exercise py/Untitled5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Batch culture in COMETS
#
# One of the most common experimental designs in microbial ecology and evolution consists on periodically transfering a small amount of a grown bacterial culture to fresh media, which is known as "batch culture".
#
# In `COMETS`, this is easily done using three parameters
# * `batchDilution` is either True or False; specifies whether periodic dilutions are performed.
# * `dilTime` is the periodicity at which dilutions are performed, in hr.
# * `dilFactor` is the dilution factor applied. If the value passed here is larger than one, that will be the dilution factor; if it is smaller than one, it will be the applied dilution properly. For instance, a 1:100 dilution can be specified as either 100 or 0.01.
#
# Let's first do a batch culture with no oxygen in the environment.
# +
import comets as c
import os
import pandas as pd
import cobra as cb
pd.options.display.max_rows = 10
# Set relevant parameters
b_params = c.params()
b_params.all_params['batchDilution'] = True
b_params.all_params['dilTime'] = 24
b_params.all_params['dilFactor'] = 100
b_params.all_params['timeStep'] = .2
b_params.all_params['maxCycles'] = 400
# Set layout
b_layout = c.layout('test_batch/batch_layout')
b_layout.media.loc[b_layout.media['metabolite'] == 'glc__D_e', 'init_amount'] = 20
b_layout.media.loc[b_layout.media['metabolite'] == 'o2_e', 'init_amount'] = 0
b_layout.media
# -
# Now, run simulation and draw the biomass curve.
# +
# create comets object from the loaded parameters and layout
batch_anaerob = c.comets(b_layout, b_params)
# run comets simulation
batch_anaerob.run()
# -
batch_anaerob.total_biomass.plot(x = 'cycle', y = 'iJO1366')
# Here, we see that the population is unable to reach enough density in a single cycle to maintain a stable population, an is therefore going to extinction.
#
# Lets now add oxygen to the environment, and redo the simulation.
b_layout.media.loc[b_layout.media['metabolite'] == 'o2_e', 'init_amount'] = 1000
# +
# create comets object from the loaded parameters and layout
batch_aerob = c.comets(b_layout, b_params)
# run comets simulation
batch_aerob.run()
# -
batch_aerob.total_biomass.plot(x = 'cycle', y = 'iJO1366')
# Now, with oxygen in the environment enabling respiration, the growth rate is much higher, and the population soon reaches stationnary phase, maintaining the population through the successive dilutions.
| .ipynb_checkpoints/serial_transfers-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import gym
gym.version.VERSION
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
from gridworld_env import GridworldEnv
env = GridworldEnv(4) # Number of plan
env.render()
env.observation_space
print(env.reset())
print(env.action_space.sample())
env.step(env.action_space.sample())
env.verbose = True
print( env._get_agent_start_target_state())
env.grid_map_shape
env.observation_space
env.action_space
# Try random policy
env.reset()
moves = [0,2,2,3,2,4]
env.render()
print('State \t\t\t\t - Reward')
for i in range(len(moves)):
move = env.action_space.sample()
obs , reward, _, _ = env.step(moves[i])
print(obs,'\t\t\t',reward)
env.render()
print('Total episode reward: ', env.episode_total_reward)
# ## Q-learning Example
# +
# Q learning params
ALPHA = 0.1 # learning rate
GAMMA = 0.95 # reward discount
LEARNING_COUNT = 1000
TEST_COUNT = 100
TURN_LIMIT = 1000
IS_MONITOR = True
class Agent:
def __init__(self, env):
self.env = env
self.episode_reward = 0.0
self.q_val = np.zeros(64 * 5).reshape(64, 5).astype(np.float32)
def learn(self):
# one episode learning
state = self.env.reset()
#self.env.render()
for t in range(TURN_LIMIT):
act = self.env.action_space.sample() # random
next_state, reward, done, info = self.env.step(act)
q_next_max = np.max(self.q_val[int(64.*(next_state[0]+1.)/2.)])
# Q <- Q + a(Q' - Q)
# <=> Q <- (1-a)Q + a(Q')
self.q_val[int(64.*(state[0]+1.)/2.)][act] = (1 - ALPHA) * self.q_val[int(64.*(state[0]+1.)/2.)][act]\
+ ALPHA * (reward + GAMMA * q_next_max)
self.episode_reward += reward
#self.env.render()
if done:
return self.env.episode_total_reward
else:
state = next_state
return 0.0 # over limit
def test(self):
state = self.env.reset()
for t in range(TURN_LIMIT):
act = np.argmax(self.q_val[int(64.*(state[0]+1.)/2.)])
next_state, reward, done, info = self.env.step(act)
if done:
return self.env.episode_total_reward
else:
state = next_state
return 0.0 # over limit
env = GridworldEnv(1)
env.reset()
agent = Agent(env)
print("###### LEARNING #####")
reward_total = 0.0
for i in range(LEARNING_COUNT):
reward_total += agent.learn()
print("episodes : {}".format(LEARNING_COUNT))
print("total reward : {}".format(reward_total))
print("average reward: {:.2f}".format(reward_total / LEARNING_COUNT))
print("Q Value :{}".format(agent.q_val))
print("###### TEST #####")
reward_total = 0.0
for i in range(TEST_COUNT):
reward_total += agent.test()
print("episodes : {}".format(TEST_COUNT))
print("total reward : {}".format(reward_total))
print("average reward: {:.2f}".format(reward_total / TEST_COUNT))
# -
Q = agent.q_val
policy_function = np.argmax( Q , axis = 1).reshape(8,8)
def plot_policy( policy_function ):
plt.figure()
plt.imshow( policy_function , interpolation='none' )
plt.colorbar()
for row in range( policy_function.shape[0] ):
for col in range( policy_function.shape[1] ):
if policy_function[row][col] == 0:
continue
if policy_function[row][col] == 1:
dx = 0; dy = .5
if policy_function[row][col] == 2:
dx = 0; dy = -.5
if policy_function[row][col] == 3:
dx = -.5; dy = 0
if policy_function[row][col] == 4:
dx = .5; dy = 0
plt.arrow( col , row , dx , dy , shape='full', fc='w' , ec='w' , lw=3, length_includes_head=True, head_width=.2 )
plt.title( 'Policy' )
plt.show()
plot_policy(policy_function)
env.reset()
env.render()
| code/envs/GridWorld-Test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="VfWx3rqSen-p" executionInfo={"status": "ok", "timestamp": 1629682464026, "user_tz": 240, "elapsed": 5757, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13953621006807715822"}} outputId="5216386b-bfba-4d20-cd9f-84a863beb26e"
import pandas as pd
import numpy as np
import os
import random
import progressbar
import pickle
import matplotlib.pyplot as plt
from sklearn.linear_model import (GammaRegressor, PoissonRegressor,
TweedieRegressor)
from sklearn.model_selection import cross_validate
# !pip install -U imbalanced-learn
from imblearn.over_sampling import (ADASYN, BorderlineSMOTE, KMeansSMOTE,
RandomOverSampler, SMOTE, SMOTEN, SMOTENC,
SVMSMOTE)
from imblearn.under_sampling import (AllKNN, ClusterCentroids,
CondensedNearestNeighbour,
EditedNearestNeighbours,
InstanceHardnessThreshold,
NearMiss, NeighbourhoodCleaningRule,
OneSidedSelection, RandomUnderSampler,
RepeatedEditedNearestNeighbours,
TomekLinks)
from imblearn.combine import SMOTEENN, SMOTETomek
import seaborn as sb
from google.colab import drive
drive.mount('/content/gdrive')
from sklearn.model_selection import cross_validate
# !pip install -U imbalanced-learn
from imblearn.over_sampling import (ADASYN, BorderlineSMOTE, KMeansSMOTE,
RandomOverSampler, SMOTE, SMOTEN, SMOTENC,
SVMSMOTE)
from imblearn.under_sampling import (AllKNN, ClusterCentroids,
CondensedNearestNeighbour,
EditedNearestNeighbours,
InstanceHardnessThreshold,
NearMiss, NeighbourhoodCleaningRule,
OneSidedSelection, RandomUnderSampler,
RepeatedEditedNearestNeighbours,
TomekLinks)
from imblearn.combine import SMOTEENN, SMOTETomek
import seaborn as sb
from google.colab import drive
drive.mount('/content/gdrive')
# + [markdown] id="zUeNOIWTr7rK"
# Total number of data
# + id="QjeAshw_r99D" executionInfo={"status": "ok", "timestamp": 1629682046945, "user_tz": 240, "elapsed": 51, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13953621006807715822"}}
data_per_csv = 512
def total():
total_files = 0
for files in os.listdir('gdrive/My Drive/Summer Research/HRV/Outlier Free/All/'):
total_files += 1
return total_files
# + [markdown] id="e8NL-4nNI2u8"
# Load data
# + id="WzdEjxhlbWcL" executionInfo={"status": "ok", "timestamp": 1629682046946, "user_tz": 240, "elapsed": 48, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13953621006807715822"}}
def loadHRVData(c):
hrv_and_labels = list()
if c == 'wt':
with open('gdrive/My Drive/Summer Research/Variables/wt_pseudoimage_hrv_and_labels.pkl', 'rb') as file:
#load data from file
hrv_and_labels = pickle.load(file)
elif c == 'wt denoised':
with open('gdrive/My Drive/Summer Research/Variables/wt_denoised_pseudoimage_hrv_and_labels.pkl', 'rb') as file:
#load data from file
hrv_and_labels = pickle.load(file)
elif c == 'normal':
size = (163, 223, 4)
with open('gdrive/My Drive/Summer Research/Variables/normal_hrv_and_labels.pkl', 'rb') as file:
#load data from file
hrv_and_labels = pickle.load(file)
elif c == 'array':
with open('gdrive/My Drive/Summer Research/Variables/array_hrv_and_labels.pkl', 'rb') as file:
#load data from file
hrv_and_labels = pickle.load(file)
elif c == 'wt a1d1d2d3 coords':
with open('gdrive/My Drive/Summer Research/Variables/wt_a1d1d2d3_coords_hrv_and_labels.pkl', 'rb') as file:
#save data to a file
hrv_and_labels = pickle.load(file)
elif c == 'wt a1d1d2d3 denoised coords':
with open('gdrive/My Drive/Summer Research/Variables/wt_a1d1d2d3_denoised_coords_hrv_and_labels.pkl', 'rb') as file:
#save data to a file
hrv_and_labels = pickle.load(file)
return hrv_and_labels
# + [markdown] id="ktj6NuDRUX_H"
# Confusion matrix (tp+fp+tn+fn set to equal 1)
# + id="v66E98unUZFH" executionInfo={"status": "ok", "timestamp": 1629682046947, "user_tz": 240, "elapsed": 46, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13953621006807715822"}}
def calculate_average_confusion_matrix(recall, precision, accuracy, method):
fp_list = list()
tp_list = list()
if method == 'each':
# tp/(tp+fp)=Precision, tp/(tp+fn)=Recall, (tp+tn)/(tp+fn+fp+tn)=Accuracy
tp_avg = fp_avg = tn_avg = fn_avg = 0
for i in range(len(recall)):
fn = (1 / recall[i]) - 1
fp = (1 / precision[i]) - 1
tp = 1
tn = (1 - accuracy[i] - accuracy[i] * fn - accuracy[i] * fp) / (accuracy[i] - 1)
fp_list.append(fp/(fp+tn))
tp_list.append(tp/(tp+fn))
tp_avg += tp
fp_avg += fp
tn_avg += tn
fn_avg += fn
sum = fn_avg + fp_avg + tp_avg + tn_avg
fn_avg, fp_avg, tp_avg, tn_avg = fn_avg/sum, fp_avg/sum, tp_avg/sum, tn_avg/sum
elif method == 'avg':
r_avg = recall.sum()/len(recall)
p_avg = precision.sum()/len(precision)
a_avg = accuracy.sum()/len(accuracy)
fn = (1 / r_avg) - 1
fp = (1 / p_avg) - 1
tp = 1
tn = (1 - a_avg - a_avg * fn - a_avg * fp) / (a_avg - 1)
sum = fn + fp + tp + tn
fn_avg, fp_avg, tp_avg, tn_avg = fn/sum, fp/sum, tp/sum, tn/sum
fp_list.append(fp_avg)
tp_list.append(tp_avg)
tpr = tp/(tp+fn)
tnr = tn/(tn+fp)
return (tpr+tnr)/2, np.array([[tp_avg, fp_avg], [fn_avg, tn_avg]]), fp_list, tp_list
# + [markdown] id="3WugkQ_xbiRz"
# Oversampling and undersampling
# + id="fOo8Yclhbker" executionInfo={"status": "ok", "timestamp": 1629682046948, "user_tz": 240, "elapsed": 46, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13953621006807715822"}}
def resampling(args):
if args == 'SMOTEENN':
resampler = SMOTEENN(n_jobs=-1)
elif args == 'SMOTETomek':
resampler = SMOTETomek(n_jobs=-1)
return resampler
# + [markdown] id="XTrcf7W2JzW_"
# GLM model
# + id="9x-N1YJTJ0rv" executionInfo={"status": "ok", "timestamp": 1629682641659, "user_tz": 240, "elapsed": 159, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13953621006807715822"}}
def GLMModel(X, y):
scores = []
avg_acc = []
#params = [10]
model = GammaRegressor()
#K-fold Cross Validation
scores = cross_validate(model, X, y, cv=10, scoring=('accuracy', 'neg_root_mean_squared_error', 'precision', 'recall', 'roc_auc'), n_jobs=-1, verbose=0, return_train_score=False)
return scores
# + id="u4CZGRm7J_9h" executionInfo={"status": "ok", "timestamp": 1629682046950, "user_tz": 240, "elapsed": 43, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13953621006807715822"}}
def metrics(scores, resampling_method, data_choice):
which_metric = 'test'
acc = scores[which_metric+'_accuracy']
rmse = scores[which_metric+'_neg_root_mean_squared_error']
precision = scores[which_metric+'_precision']
recall = scores[which_metric+'_recall']
auc = scores[which_metric+'_roc_auc']
bAcc, matrix, fp, tp = calculate_average_confusion_matrix(recall, precision, acc, 'each')
dir = 'gdrive/My Drive/Summer Research/Figures/GLM/'
file_name = resampling_method+'-resampled '+data_choice
with open(dir+file_name+'.txt', 'w') as file:
file.write("Accuracy:"+str(acc))
file.write("\nBalanced accuracy: "+str(bAcc))
file.write("\nRMSE:"+str(rmse))
file.write("\nPrecision:"+str(precision))
file.write("\nRecall:"+str(recall))
file.write("\nROC AUC:"+str(auc))
#TODO: generate PR, ROC, PROC graphs
plt.figure(figsize=(12,8))
plt.scatter(fp, tp, label="HRV, auc="+str(auc))
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.xlim((0,1))
plt.ylim((0,1))
plt.show()
plt.figure(figsize=(12,8))
plt.scatter(precision, recall)
plt.xlabel('Precision')
plt.ylabel('Recall')
plt.title('Precision v. Recall')
plt.xlim((0,1))
plt.ylim((0,1))
plt.show()
plt.figure(figsize=(12,8))
g = sb.heatmap(matrix, annot=True, fmt='.2%', cmap='Greens', xticklabels=['Predicted Healthy', 'Predicted Diabetes'], yticklabels=['Healthy', 'Diabetes'])
g.set_yticklabels(labels=g.get_yticklabels(), va='center')
g.set_title('Confusion Matrix')
plt.savefig(dir+file_name)
# + id="aPOxnhd_XOIO" colab={"base_uri": "https://localhost:8080/", "height": 330} executionInfo={"status": "error", "timestamp": 1629682648988, "user_tz": 240, "elapsed": 5547, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13953621006807715822"}} outputId="1d698976-af5e-41f9-fa32-a22dd75ad7e3"
data_choices = {
1:'wt',
2:'wt denoised',
3:'array',
4:'wt a1d1d2d3 coords',
5:'wt a1d1d2d3 denoised coords'
}
widgets = [' [',
progressbar.Timer(format= 'elapsed time: %(elapsed)s'),
'] ',
progressbar.Bar('#'),' (',
progressbar.ETA(), ') ',
]
bar = progressbar.ProgressBar(max_value=10, widgets=widgets).start()
count = 0
for r in ['SMOTETomek', 'SMOTEENN']:
resampling_method = r
for i in range(len(data_choices)):
count += 1
bar.update(count)
data_choice = data_choices[1]
hrv_and_labels = loadHRVData(data_choice)
random.shuffle(hrv_and_labels)
X = np.array([item[0] for item in hrv_and_labels]).reshape(total(),-1)
y = np.array([item[1] for item in hrv_and_labels])
X_resampled, y_resampled = resampling(resampling_method).fit_resample(X, y)
scores = GLMModel(X_resampled, y_resampled)
metrics(scores, resampling_method, data_choice)
| Code/HRV Classification/GLM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="TjPTaRB4mpCd" colab_type="text"
# # Colab FAQ
#
# For some basic overview and features offered in Colab notebooks, check out: [Overview of Colaboratory Features](https://colab.research.google.com/notebooks/basic_features_overview.ipynb)
#
# You need to use the colab GPU for this assignmentby selecting:
#
# > **Runtime** → **Change runtime type** → **Hardware Accelerator: GPU**
# + [markdown] id="s9IS9B9-yUU5" colab_type="text"
# ## Setup PyTorch
# All files are stored at /content/csc421/a3/ folder
#
# + [markdown] id="axbuunY8UdTB" colab_type="text"
#
# + id="Z-6MQhMOlHXD" colab_type="code" outputId="1975e2a4-5c73-4262-f208-3642b7609b82" colab={"base_uri": "https://localhost:8080/", "height": 674}
######################################################################
# Setup python environment and change the current working directory
######################################################################
# !pip install torch torchvision
# !pip install Pillow==4.0.0
# %mkdir -p /My Drive/University/CSC413/P3/
# %cd /My Drive/University/CSC413/P3
# + [markdown] id="9DaTdRNuUra7" colab_type="text"
# # Helper code
# + [markdown] id="4BIpGwANoQOg" colab_type="text"
# ## Utility functions
# + id="D-UJHBYZkh7f" colab_type="code" colab={}
import os
import pdb
import argparse
import pickle as pkl
from collections import defaultdict
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
from torch.autograd import Variable
from six.moves.urllib.request import urlretrieve
import tarfile
import pickle
import sys
def get_file(fname,
origin,
untar=False,
extract=False,
archive_format='auto',
cache_dir='data'):
datadir = os.path.join(cache_dir)
if not os.path.exists(datadir):
os.makedirs(datadir)
if untar:
untar_fpath = os.path.join(datadir, fname)
fpath = untar_fpath + '.tar.gz'
else:
fpath = os.path.join(datadir, fname)
print(fpath)
if not os.path.exists(fpath):
print('Downloading data from', origin)
error_msg = 'URL fetch failure on {}: {} -- {}'
try:
try:
urlretrieve(origin, fpath)
except URLError as e:
raise Exception(error_msg.format(origin, e.errno, e.reason))
except HTTPError as e:
raise Exception(error_msg.format(origin, e.code, e.msg))
except (Exception, KeyboardInterrupt) as e:
if os.path.exists(fpath):
os.remove(fpath)
raise
if untar:
if not os.path.exists(untar_fpath):
print('Extracting file.')
with tarfile.open(fpath) as archive:
archive.extractall(datadir)
return untar_fpath
if extract:
_extract_archive(fpath, datadir, archive_format)
return fpath
class AttrDict(dict):
def __init__(self, *args, **kwargs):
super(AttrDict, self).__init__(*args, **kwargs)
self.__dict__ = self
def to_var(tensor, cuda):
"""Wraps a Tensor in a Variable, optionally placing it on the GPU.
Arguments:
tensor: A Tensor object.
cuda: A boolean flag indicating whether to use the GPU.
Returns:
A Variable object, on the GPU if cuda==True.
"""
if cuda:
return Variable(tensor.cuda())
else:
return Variable(tensor)
def create_dir_if_not_exists(directory):
"""Creates a directory if it doesn't already exist.
"""
if not os.path.exists(directory):
os.makedirs(directory)
def save_loss_plot(train_losses, val_losses, opts):
"""Saves a plot of the training and validation loss curves.
"""
plt.figure()
plt.plot(range(len(train_losses)), train_losses)
plt.plot(range(len(val_losses)), val_losses)
plt.title('BS={}, nhid={}'.format(opts.batch_size, opts.hidden_size), fontsize=20)
plt.xlabel('Epochs', fontsize=16)
plt.ylabel('Loss', fontsize=16)
plt.xticks(fontsize=14)
plt.yticks(fontsize=14)
plt.tight_layout()
plt.savefig(os.path.join(opts.checkpoint_path, 'loss_plot.pdf'))
plt.close()
def checkpoint(encoder, decoder, idx_dict, opts):
"""Saves the current encoder and decoder models, along with idx_dict, which
contains the char_to_index and index_to_char mappings, and the start_token
and end_token values.
"""
with open(os.path.join(opts.checkpoint_path, 'encoder.pt'), 'wb') as f:
torch.save(encoder, f)
with open(os.path.join(opts.checkpoint_path, 'decoder.pt'), 'wb') as f:
torch.save(decoder, f)
with open(os.path.join(opts.checkpoint_path, 'idx_dict.pkl'), 'wb') as f:
pkl.dump(idx_dict, f)
# + [markdown] id="pbvpn4MaV0I1" colab_type="text"
# ## Data loader
# + id="XVT4TNTOV3Eg" colab_type="code" colab={}
def read_lines(filename):
"""Read a file and split it into lines.
"""
lines = open(filename).read().strip().lower().split('\n')
return lines
def read_pairs(filename):
"""Reads lines that consist of two words, separated by a space.
Returns:
source_words: A list of the first word in each line of the file.
target_words: A list of the second word in each line of the file.
"""
lines = read_lines(filename)
source_words, target_words = [], []
for line in lines:
line = line.strip()
if line:
source, target = line.split()
source_words.append(source)
target_words.append(target)
return source_words, target_words
def all_alpha_or_dash(s):
"""Helper function to check whether a string is alphabetic, allowing dashes '-'.
"""
return all(c.isalpha() or c == '-' for c in s)
def filter_lines(lines):
"""Filters lines to consist of only alphabetic characters or dashes "-".
"""
return [line for line in lines if all_alpha_or_dash(line)]
def load_data():
"""Loads (English, Pig-Latin) word pairs, and creates mappings from characters to indexes.
"""
source_lines, target_lines = read_pairs('data/pig_latin_data.txt')
# Filter lines
source_lines = filter_lines(source_lines)
target_lines = filter_lines(target_lines)
all_characters = set(''.join(source_lines)) | set(''.join(target_lines))
# Create a dictionary mapping each character to a unique index
char_to_index = { char: index for (index, char) in enumerate(sorted(list(all_characters))) }
# Add start and end tokens to the dictionary
start_token = len(char_to_index)
end_token = len(char_to_index) + 1
char_to_index['SOS'] = start_token
char_to_index['EOS'] = end_token
# Create the inverse mapping, from indexes to characters (used to decode the model's predictions)
index_to_char = { index: char for (char, index) in char_to_index.items() }
# Store the final size of the vocabulary
vocab_size = len(char_to_index)
line_pairs = list(set(zip(source_lines, target_lines))) # Python 3
idx_dict = { 'char_to_index': char_to_index,
'index_to_char': index_to_char,
'start_token': start_token,
'end_token': end_token }
return line_pairs, vocab_size, idx_dict
def create_dict(pairs):
"""Creates a mapping { (source_length, target_length): [list of (source, target) pairs]
This is used to make batches: each batch consists of two parallel tensors, one containing
all source indexes and the other containing all corresponding target indexes.
Within a batch, all the source words are the same length, and all the target words are
the same length.
"""
unique_pairs = list(set(pairs)) # Find all unique (source, target) pairs
d = defaultdict(list)
for (s,t) in unique_pairs:
d[(len(s), len(t))].append((s,t))
return d
# + [markdown] id="bRWfRdmVVjUl" colab_type="text"
# ## Training and evaluation code
# + id="wa5-onJhoSeM" colab_type="code" colab={}
def string_to_index_list(s, char_to_index, end_token):
"""Converts a sentence into a list of indexes (for each character).
"""
return [char_to_index[char] for char in s] + [end_token] # Adds the end token to each index list
def translate_sentence(sentence, encoder, decoder, idx_dict, opts):
"""Translates a sentence from English to Pig-Latin, by splitting the sentence into
words (whitespace-separated), running the encoder-decoder model to translate each
word independently, and then stitching the words back together with spaces between them.
"""
if idx_dict is None:
line_pairs, vocab_size, idx_dict = load_data()
return ' '.join([translate(word, encoder, decoder, idx_dict, opts) for word in sentence.split()])
def translate(input_string, encoder, decoder, idx_dict, opts):
"""Translates a given string from English to Pig-Latin.
"""
char_to_index = idx_dict['char_to_index']
index_to_char = idx_dict['index_to_char']
start_token = idx_dict['start_token']
end_token = idx_dict['end_token']
max_generated_chars = 20
gen_string = ''
indexes = string_to_index_list(input_string, char_to_index, end_token)
indexes = to_var(torch.LongTensor(indexes).unsqueeze(0), opts.cuda) # Unsqueeze to make it like BS = 1
encoder_annotations, encoder_last_hidden = encoder(indexes)
decoder_hidden = encoder_last_hidden
decoder_input = to_var(torch.LongTensor([[start_token]]), opts.cuda) # For BS = 1
decoder_inputs = decoder_input
for i in range(max_generated_chars):
## slow decoding, recompute everything at each time
decoder_outputs, attention_weights = decoder(decoder_inputs, encoder_annotations, decoder_hidden)
generated_words = F.softmax(decoder_outputs, dim=2).max(2)[1]
ni = generated_words.cpu().numpy().reshape(-1) # LongTensor of size 1
ni = ni[-1] #latest output token
decoder_inputs = torch.cat([decoder_input, generated_words], dim=1)
if ni == end_token:
break
else:
gen_string = "".join(
[index_to_char[int(item)]
for item in generated_words.cpu().numpy().reshape(-1)])
return gen_string
def visualize_attention(input_string, encoder, decoder, idx_dict, opts):
"""Generates a heatmap to show where attention is focused in each decoder step.
"""
if idx_dict is None:
line_pairs, vocab_size, idx_dict = load_data()
char_to_index = idx_dict['char_to_index']
index_to_char = idx_dict['index_to_char']
start_token = idx_dict['start_token']
end_token = idx_dict['end_token']
max_generated_chars = 20
gen_string = ''
indexes = string_to_index_list(input_string, char_to_index, end_token)
indexes = to_var(torch.LongTensor(indexes).unsqueeze(0), opts.cuda) # Unsqueeze to make it like BS = 1
encoder_annotations, encoder_hidden = encoder(indexes)
decoder_hidden = encoder_hidden
decoder_input = to_var(torch.LongTensor([[start_token]]), opts.cuda) # For BS = 1
decoder_inputs = decoder_input
produced_end_token = False
for i in range(max_generated_chars):
## slow decoding, recompute everything at each time
decoder_outputs, attention_weights = decoder(decoder_inputs, encoder_annotations, decoder_hidden)
generated_words = F.softmax(decoder_outputs, dim=2).max(2)[1]
ni = generated_words.cpu().numpy().reshape(-1) # LongTensor of size 1
ni = ni[-1] #latest output token
decoder_inputs = torch.cat([decoder_input, generated_words], dim=1)
if ni == end_token:
break
else:
gen_string = "".join(
[index_to_char[int(item)]
for item in generated_words.cpu().numpy().reshape(-1)])
if isinstance(attention_weights, tuple):
## transformer's attention mweights
attention_weights, self_attention_weights = attention_weights
all_attention_weights = attention_weights.data.cpu().numpy()
for i in range(len(all_attention_weights)):
attention_weights_matrix = all_attention_weights[i].squeeze()
fig = plt.figure()
ax = fig.add_subplot(111)
cax = ax.matshow(attention_weights_matrix, cmap='bone')
fig.colorbar(cax)
# Set up axes
ax.set_yticklabels([''] + list(input_string) + ['EOS'], rotation=90)
ax.set_xticklabels([''] + list(gen_string) + (['EOS'] if produced_end_token else []))
# Show label at every tick
ax.xaxis.set_major_locator(ticker.MultipleLocator(1))
ax.yaxis.set_major_locator(ticker.MultipleLocator(1))
# Add title
plt.xlabel('Attention weights to the source sentence in layer {}'.format(i+1))
plt.tight_layout()
plt.grid('off')
plt.show()
#plt.savefig(save)
#plt.close(fig)
return gen_string
def compute_loss(data_dict, encoder, decoder, idx_dict, criterion, optimizer, opts):
"""Train/Evaluate the model on a dataset.
Arguments:
data_dict: The validation/test word pairs, organized by source and target lengths.
encoder: An encoder model to produce annotations for each step of the input sequence.
decoder: A decoder model (with or without attention) to generate output tokens.
idx_dict: Contains char-to-index and index-to-char mappings, and start & end token indexes.
criterion: Used to compute the CrossEntropyLoss for each decoder output.
optimizer: Train the weights if an optimizer is given. None if only evaluate the model.
opts: The command-line arguments.
Returns:
mean_loss: The average loss over all batches from data_dict.
"""
start_token = idx_dict['start_token']
end_token = idx_dict['end_token']
char_to_index = idx_dict['char_to_index']
losses = []
for key in data_dict:
input_strings, target_strings = zip(*data_dict[key])
input_tensors = [torch.LongTensor(string_to_index_list(s, char_to_index, end_token)) for s in input_strings]
target_tensors = [torch.LongTensor(string_to_index_list(s, char_to_index, end_token)) for s in target_strings]
num_tensors = len(input_tensors)
num_batches = int(np.ceil(num_tensors / float(opts.batch_size)))
for i in range(num_batches):
start = i * opts.batch_size
end = start + opts.batch_size
inputs = to_var(torch.stack(input_tensors[start:end]), opts.cuda)
targets = to_var(torch.stack(target_tensors[start:end]), opts.cuda)
# The batch size may be different in each epoch
BS = inputs.size(0)
encoder_annotations, encoder_hidden = encoder(inputs)
# The last hidden state of the encoder becomes the first hidden state of the decoder
decoder_hidden = encoder_hidden
start_vector = torch.ones(BS).long().unsqueeze(1) * start_token # BS x 1 --> 16x1 CHECKED
decoder_input = to_var(start_vector, opts.cuda) # BS x 1 --> 16x1 CHECKED
loss = 0.0
seq_len = targets.size(1) # Gets seq_len from BS x seq_len
decoder_inputs = torch.cat([decoder_input, targets[:, 0:-1]], dim=1) # Gets decoder inputs by shifting the targets to the right
decoder_outputs, attention_weights = decoder(decoder_inputs, encoder_annotations, encoder_hidden)
decoder_outputs_flatten = decoder_outputs.view(-1, decoder_outputs.size(2))
targets_flatten = targets.view(-1)
loss = criterion(decoder_outputs_flatten, targets_flatten)
losses.append(loss.item())
## training if an optimizer is provided
if optimizer:
# Zero gradients
optimizer.zero_grad()
# Compute gradients
loss.backward()
# Update the parameters of the encoder and decoder
optimizer.step()
mean_loss = np.mean(losses)
return mean_loss
def training_loop(train_dict, val_dict, idx_dict, encoder, decoder, criterion, optimizer, opts):
"""Runs the main training loop; evaluates the model on the val set every epoch.
* Prints training and val loss each epoch.
* Prints qualitative translation results each epoch using TEST_SENTENCE
* Saves an attention map for TEST_WORD_ATTN each epoch
Arguments:
train_dict: The training word pairs, organized by source and target lengths.
val_dict: The validation word pairs, organized by source and target lengths.
idx_dict: Contains char-to-index and index-to-char mappings, and start & end token indexes.
encoder: An encoder model to produce annotations for each step of the input sequence.
decoder: A decoder model (with or without attention) to generate output tokens.
criterion: Used to compute the CrossEntropyLoss for each decoder output.
optimizer: Implements a step rule to update the parameters of the encoder and decoder.
opts: The command-line arguments.
"""
start_token = idx_dict['start_token']
end_token = idx_dict['end_token']
char_to_index = idx_dict['char_to_index']
loss_log = open(os.path.join(opts.checkpoint_path, 'loss_log.txt'), 'w')
best_val_loss = 1e6
train_losses = []
val_losses = []
for epoch in range(opts.nepochs):
optimizer.param_groups[0]['lr'] *= opts.lr_decay
train_loss = compute_loss(train_dict, encoder, decoder, idx_dict, criterion, optimizer, opts)
val_loss = compute_loss(val_dict, encoder, decoder, idx_dict, criterion, None, opts)
if val_loss < best_val_loss:
checkpoint(encoder, decoder, idx_dict, opts)
gen_string = translate_sentence(TEST_SENTENCE, encoder, decoder, idx_dict, opts)
print("Epoch: {:3d} | Train loss: {:.3f} | Val loss: {:.3f} | Gen: {:20s}".format(epoch, train_loss, val_loss, gen_string))
loss_log.write('{} {} {}\n'.format(epoch, train_loss, val_loss))
loss_log.flush()
train_losses.append(train_loss)
val_losses.append(val_loss)
save_loss_plot(train_losses, val_losses, opts)
def print_data_stats(line_pairs, vocab_size, idx_dict):
"""Prints example word pairs, the number of data points, and the vocabulary.
"""
print('=' * 80)
print('Data Stats'.center(80))
print('-' * 80)
for pair in line_pairs[:5]:
print(pair)
print('Num unique word pairs: {}'.format(len(line_pairs)))
print('Vocabulary: {}'.format(idx_dict['char_to_index'].keys()))
print('Vocab size: {}'.format(vocab_size))
print('=' * 80)
def train(opts):
line_pairs, vocab_size, idx_dict = load_data()
print_data_stats(line_pairs, vocab_size, idx_dict)
# Split the line pairs into an 80% train and 20% val split
num_lines = len(line_pairs)
num_train = int(0.8 * num_lines)
train_pairs, val_pairs = line_pairs[:num_train], line_pairs[num_train:]
# Group the data by the lengths of the source and target words, to form batches
train_dict = create_dict(train_pairs)
val_dict = create_dict(val_pairs)
##########################################################################
### Setup: Create Encoder, Decoder, Learning Criterion, and Optimizers ###
##########################################################################
if opts.encoder_type == "rnn":
encoder = GRUEncoder(vocab_size=vocab_size,
hidden_size=opts.hidden_size,
opts=opts)
elif opts.encoder_type == "transformer":
encoder = TransformerEncoder(vocab_size=vocab_size,
hidden_size=opts.hidden_size,
num_layers=opts.num_transformer_layers,
opts=opts)
else:
print("why is it here")
raise NotImplementedError
if opts.decoder_type == 'rnn':
decoder = RNNDecoder(vocab_size=vocab_size,
hidden_size=opts.hidden_size)
elif opts.decoder_type == 'rnn_attention':
decoder = RNNAttentionDecoder(vocab_size=vocab_size,
hidden_size=opts.hidden_size,
attention_type=opts.attention_type)
elif opts.decoder_type == 'transformer':
decoder = TransformerDecoder(vocab_size=vocab_size,
hidden_size=opts.hidden_size,
num_layers=opts.num_transformer_layers)
else:
print("why is it here")
raise NotImplementedError
#### setup checkpoint path
model_name = 'h{}-bs{}-{}'.format(opts.hidden_size,
opts.batch_size,
opts.decoder_type)
opts.checkpoint_path = model_name
create_dir_if_not_exists(opts.checkpoint_path)
####
if opts.cuda:
encoder.cuda()
decoder.cuda()
print("Moved models to GPU!")
criterion = nn.CrossEntropyLoss()
optimizer = optim.Adam(list(encoder.parameters()) + list(decoder.parameters()), lr=opts.learning_rate)
try:
training_loop(train_dict, val_dict, idx_dict, encoder, decoder, criterion, optimizer, opts)
except KeyboardInterrupt:
print('Exiting early from training.')
return encoder, decoder
return encoder, decoder
def print_opts(opts):
"""Prints the values of all command-line arguments.
"""
print('=' * 80)
print('Opts'.center(80))
print('-' * 80)
for key in opts.__dict__:
print('{:>30}: {:<30}'.format(key, opts.__dict__[key]).center(80))
print('=' * 80)
# + [markdown] id="0yh08KhgnA30" colab_type="text"
# ## Download dataset
# + id="aROU2xZanDKq" colab_type="code" outputId="a12adfe1-5abd-47e4-97dc-c880b8ec3b1d" colab={"base_uri": "https://localhost:8080/", "height": 34}
######################################################################
# Download Translation datasets
######################################################################
data_fpath = get_file(fname='pig_latin_data.txt',
origin='http://www.cs.toronto.edu/~jba/pig_latin_data.txt',
untar=False)
# + [markdown] id="YDYMr7NclZdw" colab_type="text"
# # Part 1: Gated Recurrent Unit (GRU
# + [markdown] id="dCae1mOUlZrC" colab_type="text"
# ## Step 1: GRU Cell
# Please implement the Gated Recurent Unit class defined in the next cell.
# + id="3HMO7FD6l5RU" colab_type="code" colab={}
class MyGRUCell(nn.Module):
def __init__(self, input_size, hidden_size):
super(MyGRUCell, self).__init__()
self.input_size = input_size
self.hidden_size = hidden_size
# ------------
# FILL THIS IN
# ------------
## Input linear layers
self.Wiz = nn.Linear(input_size, hidden_size)
self.Wir = nn.Linear(input_size, hidden_size)
self.Wih = nn.Linear(input_size, hidden_size)
## Hidden linear layers
self.Whz = nn.Linear(hidden_size, hidden_size)
self.Whr = nn.Linear(hidden_size, hidden_size)
self.Whh = nn.Linear(hidden_size, hidden_size)
def forward(self, x, h_prev):
"""Forward pass of the GRU computation for one time step.
Arguments
x: batch_size x input_size
h_prev: batch_size x hidden_size
Returns:
h_new: batch_size x hidden_size
"""
# ------------
# FILL THIS IN
# ------------
z = torch.sigmoid(self.Wiz(x) + self.Whz(h_prev))
r = torch.sigmoid(self.Wir(x) + self.Whr(h_prev))
g = torch.tanh(self.Wih(x) * self.Whh(h_prev))
h_new = (1-z) * g + z * h_prev
return h_new
# + [markdown] id="ecEq4TP2lZ4Z" colab_type="text"
# ## Step 2: GRU Encoder
# Please inspect the following recurrent encoder/decoder implementations. Make sure to run the cells before proceeding.
# + id="8jDNim2fmVJV" colab_type="code" colab={}
class GRUEncoder(nn.Module):
def __init__(self, vocab_size, hidden_size, opts):
super(GRUEncoder, self).__init__()
self.vocab_size = vocab_size
self.hidden_size = hidden_size
self.opts = opts
self.embedding = nn.Embedding(vocab_size, hidden_size)
self.gru = MyGRUCell(hidden_size, hidden_size)
def forward(self, inputs):
"""Forward pass of the encoder RNN.
Arguments:
inputs: Input token indexes across a batch for all time steps in the sequence. (batch_size x seq_len)
Returns:
annotations: The hidden states computed at each step of the input sequence. (batch_size x seq_len x hidden_size)
hidden: The final hidden state of the encoder, for each sequence in a batch. (batch_size x hidden_size)
"""
batch_size, seq_len = inputs.size()
hidden = self.init_hidden(batch_size)
encoded = self.embedding(inputs) # batch_size x seq_len x hidden_size
annotations = []
for i in range(seq_len):
x = encoded[:,i,:] # Get the current time step, across the whole batch
hidden = self.gru(x, hidden)
annotations.append(hidden)
annotations = torch.stack(annotations, dim=1)
return annotations, hidden
def init_hidden(self, bs):
"""Creates a tensor of zeros to represent the initial hidden states
of a batch of sequences.
Arguments:
bs: The batch size for the initial hidden state.
Returns:
hidden: An initial hidden state of all zeros. (batch_size x hidden_size)
"""
return to_var(torch.zeros(bs, self.hidden_size), self.opts.cuda)
# + id="HvwizYM9ma4p" colab_type="code" colab={}
class RNNDecoder(nn.Module):
def __init__(self, vocab_size, hidden_size):
super(RNNDecoder, self).__init__()
self.vocab_size = vocab_size
self.hidden_size = hidden_size
self.embedding = nn.Embedding(vocab_size, hidden_size)
self.rnn = MyGRUCell(input_size=hidden_size, hidden_size=hidden_size)
self.out = nn.Linear(hidden_size, vocab_size)
def forward(self, inputs, annotations, hidden_init):
"""Forward pass of the non-attentional decoder RNN.
Arguments:
inputs: Input token indexes across a batch. (batch_size x seq_len)
annotations: This is not used here. It just maintains consistency with the
interface used by the AttentionDecoder class.
hidden_init: The hidden states from the last step of encoder, across a batch. (batch_size x hidden_size)
Returns:
output: Un-normalized scores for each token in the vocabulary, across a batch for all the decoding time steps. (batch_size x decoder_seq_len x vocab_size)
None
"""
batch_size, seq_len = inputs.size()
embed = self.embedding(inputs) # batch_size x seq_len x hidden_size
hiddens = []
h_prev = hidden_init
for i in range(seq_len):
x = embed[:,i,:] # Get the current time step input tokens, across the whole batch
h_prev = self.rnn(x, h_prev) # batch_size x hidden_size
hiddens.append(h_prev)
hiddens = torch.stack(hiddens, dim=1) # batch_size x seq_len x hidden_size
output = self.out(hiddens) # batch_size x seq_len x vocab_size
return output, None
# + [markdown] id="TSDTbsydlaGI" colab_type="text"
# ## Step 3: Training and Analysis
# Train the following language model comprised of recurrent encoder and decoders.
# + id="Gv25zCMQGVzD" colab_type="code" colab={}
import warnings
warnings.filterwarnings("ignore")
# + id="H3YLrAjsmx_W" colab_type="code" outputId="340fa1ec-debc-46a4-bb25-f34ffd72590a" colab={"base_uri": "https://localhost:8080/", "height": 1000}
TEST_SENTENCE = 'the air conditioning is working'
args = AttrDict()
args_dict = {
'cuda':True,
'nepochs':100,
'checkpoint_dir':"checkpoints",
'learning_rate':0.005,
'lr_decay':0.99,
'batch_size':64,
'hidden_size':20,
'encoder_type': 'rnn', # options: rnn / transformer
'decoder_type': 'rnn', # options: rnn / rnn_attention / transformer
'attention_type': '', # options: additive / scaled_dot
}
args.update(args_dict)
print_opts(args)
rnn_encoder, rnn_decoder = train(args)
translated = translate_sentence(TEST_SENTENCE, rnn_encoder, rnn_decoder, None, args)
print("source:\t\t{} \ntranslated:\t{}".format(TEST_SENTENCE, translated))
# + [markdown] id="cE4ijaCzneAt" colab_type="text"
# Try translating different sentences by changing the variable TEST_SENTENCE. Identify two distinct failure modes and briefly describe them.
# + id="WrNnz8W1nULf" colab_type="code" outputId="d41b7472-9c8c-4814-9717-49f327d6f167" colab={"base_uri": "https://localhost:8080/", "height": 87}
TEST_SENTENCE = 'tea team tight'
translated = translate_sentence(TEST_SENTENCE, rnn_encoder, rnn_decoder, None, args)
print("source:\t\t{} \ntranslated:\t{}".format(TEST_SENTENCE, translated))
TEST_SENTENCE = 'shopping fighting running'
translated = translate_sentence(TEST_SENTENCE, rnn_encoder, rnn_decoder, None, args)
print("source:\t\t{} \ntranslated:\t{}".format(TEST_SENTENCE, translated))
# + [markdown] id="RWwA6OGqlaTq" colab_type="text"
# # Part 2: Additive Attention
# + [markdown] id="AJSafHSAmu_w" colab_type="text"
# ## Step 1: Additive Attention
# Already implemented the additive attention mechanism. Write down the mathematical expression for $\tilde{\alpha}_i^{(t)}, \alpha_i^{(t)}, c_t$ as a function of $W_1, W_2, b_1, b_2, Q_t, K_i$.
# + id="AdewEVSMo5jJ" colab_type="code" colab={}
class AdditiveAttention(nn.Module):
def __init__(self, hidden_size):
super(AdditiveAttention, self).__init__()
self.hidden_size = hidden_size
# A two layer fully-connected network
# hidden_size*2 --> hidden_size, ReLU, hidden_size --> 1
self.attention_network = nn.Sequential(
nn.Linear(hidden_size*2, hidden_size),
nn.ReLU(),
nn.Linear(hidden_size, 1)
)
self.softmax = nn.Softmax(dim=1)
def forward(self, queries, keys, values):
"""The forward pass of the additive attention mechanism.
Arguments:
queries: The current decoder hidden state. (batch_size x hidden_size)
keys: The encoder hidden states for each step of the input sequence. (batch_size x seq_len x hidden_size)
values: The encoder hidden states for each step of the input sequence. (batch_size x seq_len x hidden_size)
Returns:
context: weighted average of the values (batch_size x 1 x hidden_size)
attention_weights: Normalized attention weights for each encoder hidden state. (batch_size x seq_len x 1)
The attention_weights must be a softmax weighting over the seq_len annotations.
"""
batch_size = keys.size(0)
expanded_queries = queries.view(batch_size, -1, self.hidden_size).expand_as(keys)
concat_inputs = torch.cat([expanded_queries, keys], dim=2)
unnormalized_attention = self.attention_network(concat_inputs)
attention_weights = self.softmax(unnormalized_attention)
context = torch.bmm(attention_weights.transpose(2,1), values)
return context, attention_weights
# + [markdown] id="73_p8d5EmvOJ" colab_type="text"
# ## Step 2: RNN Additive Attention Decoder
# We will now implement a recurrent decoder that makes use of the additive attention mechanism. Read the description in the assignment worksheet and complete the following implementation.
# + id="RJaABkXrpJSw" colab_type="code" colab={}
class RNNAttentionDecoder(nn.Module):
def __init__(self, vocab_size, hidden_size, attention_type='scaled_dot'):
super(RNNAttentionDecoder, self).__init__()
self.vocab_size = vocab_size
self.hidden_size = hidden_size
self.embedding = nn.Embedding(vocab_size, hidden_size)
self.rnn = MyGRUCell(input_size=hidden_size*2, hidden_size=hidden_size)
if attention_type == 'additive':
self.attention = AdditiveAttention(hidden_size=hidden_size)
elif attention_type == 'scaled_dot':
self.attention = ScaledDotAttention(hidden_size=hidden_size)
self.out = nn.Linear(hidden_size, vocab_size)
def forward(self, inputs, annotations, hidden_init):
"""Forward pass of the attention-based decoder RNN.
Arguments:
inputs: Input token indexes across a batch for all the time step. (batch_size x decoder_seq_len)
annotations: The encoder hidden states for each step of the input.
sequence. (batch_size x seq_len x hidden_size)
hidden_init: The final hidden states from the encoder, across a batch. (batch_size x hidden_size)
Returns:
output: Un-normalized scores for each token in the vocabulary, across a batch for all the decoding time steps. (batch_size x decoder_seq_len x vocab_size)
attentions: The stacked attention weights applied to the encoder annotations (batch_size x encoder_seq_len x decoder_seq_len)
"""
batch_size, seq_len = inputs.size()
embed = self.embedding(inputs) # batch_size x seq_len x hidden_size
hiddens = []
attentions = []
h_prev = hidden_init
for i in range(seq_len):
# ------------
# FILL THIS IN - START
# ------------
embed_current = embed[:,i,:] # Get the current time step, across the whole batch (batch size x hidden size)
context, attention_weights = self.attention(embed_current, annotations, annotations) # batch_size x 1 x hidden_size
embed_and_context = torch.cat([embed_current, torch.squeeze(context, dim=1)], dim=1) # batch_size x (2*hidden_size)
h_prev = self.rnn(embed_and_context, h_prev) # batch_size x hidden_size
# ------------
# FILL THIS IN - END
# ------------
hiddens.append(h_prev)
attentions.append(attention_weights)
hiddens = torch.stack(hiddens, dim=1) # batch_size x seq_len x hidden_size
attentions = torch.cat(attentions, dim=2) # batch_size x seq_len x seq_len
output = self.out(hiddens) # batch_size x seq_len x vocab_size
return output, attentions
# + [markdown] id="vYPae08Io1Fi" colab_type="text"
# ## Step 3: Training and Analysis
# Train the following language model that uses a recurrent encoder, and a recurrent decoder that has an additive attention component.
# + id="o3-FuzY1pepu" colab_type="code" outputId="12f2df4c-6c17-42b2-a61d-542ed4d5ed2a" colab={"base_uri": "https://localhost:8080/", "height": 1000}
TEST_SENTENCE = 'the air conditioning is working'
args = AttrDict()
args_dict = {
'cuda':True,
'nepochs':100,
'checkpoint_dir':"checkpoints",
'learning_rate':0.005,
'lr_decay':0.99,
'batch_size':64,
'hidden_size':20,
'encoder_type': 'rnn', # options: rnn / transformer
'decoder_type': 'rnn_attention', # options: rnn / rnn_attention / transformer
'attention_type': 'additive', # options: additive / scaled_dot
}
args.update(args_dict)
print_opts(args)
rnn_attn_encoder, rnn_attn_decoder = train(args)
translated = translate_sentence(TEST_SENTENCE, rnn_attn_encoder, rnn_attn_decoder, None, args)
print("source:\t\t{} \ntranslated:\t{}".format(TEST_SENTENCE, translated))
# + id="VNVKbLc0ACj_" colab_type="code" outputId="d0dfdc06-1adf-4157-acb3-d1312f49dbfa" colab={"base_uri": "https://localhost:8080/", "height": 51}
TEST_SENTENCE = 'the air conditioning is working'
translated = translate_sentence(TEST_SENTENCE, rnn_attn_encoder, rnn_attn_decoder, None, args)
print("source:\t\t{} \ntranslated:\t{}".format(TEST_SENTENCE, translated))
# + [markdown] id="kw_GOIvzo1ix" colab_type="text"
# # Part 3: Scaled Dot Product Attention
# + [markdown] id="xq7nhsEio1w-" colab_type="text"
# ## Step 1: Implement Dot-Product Attention
# Implement the scaled dot product attention module described in the assignment worksheet.
# + id="d_j3oY3hqsJQ" colab_type="code" colab={}
class ScaledDotAttention(nn.Module):
def __init__(self, hidden_size):
super(ScaledDotAttention, self).__init__()
self.hidden_size = hidden_size
self.Q = nn.Linear(hidden_size, hidden_size)
self.K = nn.Linear(hidden_size, hidden_size)
self.V = nn.Linear(hidden_size, hidden_size)
self.softmax = nn.Softmax(dim=1)
self.scaling_factor = torch.rsqrt(torch.tensor(self.hidden_size, dtype= torch.float))
def forward(self, queries, keys, values):
"""The forward pass of the scaled dot attention mechanism.
Arguments:
queries: The current decoder hidden state, 2D or 3D tensor. (batch_size x (k) x hidden_size)
keys: The encoder hidden states for each step of the input sequence. (batch_size x seq_len x hidden_size)
values: The encoder hidden states for each step of the input sequence. (batch_size x seq_len x hidden_size)
Returns:
context: weighted average of the values (batch_size x k x hidden_size)
attention_weights: Normalized attention weights for each encoder hidden state. (batch_size x seq_len x k)
The output must be a softmax weighting over the seq_len annotations.
"""
# ------------
# FILL THIS IN
# ------------
batch_size = keys.size(0)
q = self.Q(queries)
k = self.K(keys)
v = self.V(values)
unnormalized_attention = (k @ q.transpose(1,2))/ self.scaling_factor
attention_weights = self.softmax(unnormalized_attention)
context = attention_weights.transpose(1,2) @ v
return context, attention_weights
# + [markdown] id="unReAOrjo113" colab_type="text"
# ## Step 2: Implement Causal Dot-Product Attention
# Now implement the scaled causal dot product described in the assignment worksheet.
# + id="ovigzQffrKqj" colab_type="code" colab={}
class CausalScaledDotAttention(nn.Module):
def __init__(self, hidden_size):
super(CausalScaledDotAttention, self).__init__()
self.hidden_size = hidden_size
self.neg_inf = torch.tensor(-1e7)
self.Q = nn.Linear(hidden_size, hidden_size)
self.K = nn.Linear(hidden_size, hidden_size)
self.V = nn.Linear(hidden_size, hidden_size)
self.softmax = nn.Softmax(dim=1)
self.scaling_factor = torch.rsqrt(torch.tensor(self.hidden_size, dtype= torch.float))
def forward(self, queries, keys, values):
"""The forward pass of the scaled dot attention mechanism.
Arguments:
queries: The current decoder hidden state, 2D or 3D tensor. (batch_size x (k) x hidden_size)
keys: The encoder hidden states for each step of the input sequence. (batch_size x seq_len x hidden_size)
values: The encoder hidden states for each step of the input sequence. (batch_size x seq_len x hidden_size)
Returns:
context: weighted average of the values (batch_size x k x hidden_size)
attention_weights: Normalized attention weights for each encoder hidden state. (batch_size x seq_len x k)
The output must be a softmax weighting over the seq_len annotations.
"""
# ------------
# FILL THIS IN
# ------------
batch_size = keys.size(0)
q = self.Q(queries)
k = self.K(keys)
v = self.V(values)
unnormalized_attention = (k @ q.transpose(1,2))/ self.scaling_factor
mask = torch.tril(unnormalized_attention)
mask[mask == 0] = self.neg_inf
attention_weights = self.softmax(mask)
context = attention_weights.transpose(1,2) @ v
return context, attention_weights
# + [markdown] id="9tcpUFKqo2Oi" colab_type="text"
# ## Step 3: Transformer Encoder
# Complete the following transformer encoder implementation.
# + id="N3B-fWsarlVk" colab_type="code" colab={}
class TransformerEncoder(nn.Module):
def __init__(self, vocab_size, hidden_size, num_layers, opts):
super(TransformerEncoder, self).__init__()
self.vocab_size = vocab_size
self.hidden_size = hidden_size
self.num_layers = num_layers
self.opts = opts
self.embedding = nn.Embedding(vocab_size, hidden_size)
# IMPORTANT CORRECTION: NON-CAUSAL ATTENTION SHOULD HAVE BEEN
# USED IN THE TRANSFORMER ENCODER.
# NEW VERSION:
self.self_attentions = nn.ModuleList([ScaledDotAttention(
hidden_size=hidden_size,
) for i in range(self.num_layers)])
# PREVIONS VERSION:
# self.self_attentions = nn.ModuleList([CausalScaledDotAttention(
# hidden_size=hidden_size,
# ) for i in range(self.num_layers)])
self.attention_mlps = nn.ModuleList([nn.Sequential(
nn.Linear(hidden_size, hidden_size),
nn.ReLU(),
) for i in range(self.num_layers)])
self.positional_encodings = self.create_positional_encodings()
def forward(self, inputs):
"""Forward pass of the encoder RNN.
Arguments:
inputs: Input token indexes across a batch for all time steps in the sequence. (batch_size x seq_len)
Returns:
annotations: The hidden states computed at each step of the input sequence. (batch_size x seq_len x hidden_size)
hidden: The final hidden state of the encoder, for each sequence in a batch. (batch_size x hidden_size)
"""
batch_size, seq_len = inputs.size()
# ------------
# FILL THIS IN - START
# ------------
encoded = self.embedding(inputs) # batch_size x seq_len x hidden_size
# Add positinal embeddings from self.create_positional_encodings. (a'la https://arxiv.org/pdf/1706.03762.pdf, section 3.5)
encoded += self.positional_encodings[:seq_len]
annotations = encoded
for i in range(self.num_layers):
new_annotations, self_attention_weights = self.self_attentions[i](encoded, annotations, annotations) # batch_size x seq_len x hidden_size
residual_annotations = annotations + new_annotations
new_annotations = self.attention_mlps[i](residual_annotations)
annotations = residual_annotations + new_annotations
# ------------
# FILL THIS IN - END
# ------------
# Transformer encoder does not have a last hidden layer.
return annotations, None
def create_positional_encodings(self, max_seq_len=1000):
"""Creates positional encodings for the inputs.
Arguments:
max_seq_len: a number larger than the maximum string length we expect to encounter during training
Returns:
pos_encodings: (max_seq_len, hidden_dim) Positional encodings for a sequence with length max_seq_len.
"""
pos_indices = torch.arange(max_seq_len)[..., None]
dim_indices = torch.arange(self.hidden_size//2)[None, ...]
exponents = (2*dim_indices).float()/(self.hidden_size)
trig_args = pos_indices / (10000**exponents)
sin_terms = torch.sin(trig_args)
cos_terms = torch.cos(trig_args)
pos_encodings = torch.zeros((max_seq_len, self.hidden_size))
pos_encodings[:, 0::2] = sin_terms
pos_encodings[:, 1::2] = cos_terms
if self.opts.cuda:
pos_encodings = pos_encodings.cuda()
return pos_encodings
# + [markdown] id="z1hDi020rT36" colab_type="text"
# ## Step 4: Transformer Decoder
# Complete the following transformer decoder implementation.
# + id="nyvTZFxtrvc6" colab_type="code" colab={}
class TransformerDecoder(nn.Module):
def __init__(self, vocab_size, hidden_size, num_layers):
super(TransformerDecoder, self).__init__()
self.vocab_size = vocab_size
self.hidden_size = hidden_size
self.embedding = nn.Embedding(vocab_size, hidden_size)
self.num_layers = num_layers
self.self_attentions = nn.ModuleList([ScaledDotAttention(
hidden_size=hidden_size,
) for i in range(self.num_layers)])
self.encoder_attentions = nn.ModuleList([ScaledDotAttention(
hidden_size=hidden_size,
) for i in range(self.num_layers)])
self.attention_mlps = nn.ModuleList([nn.Sequential(
nn.Linear(hidden_size, hidden_size),
nn.ReLU(),
) for i in range(self.num_layers)])
self.out = nn.Linear(hidden_size, vocab_size)
self.positional_encodings = self.create_positional_encodings()
def forward(self, inputs, annotations, hidden_init):
"""Forward pass of the attention-based decoder RNN.
Arguments:
inputs: Input token indexes across a batch for all the time step. (batch_size x decoder_seq_len)
annotations: The encoder hidden states for each step of the input.
sequence. (batch_size x seq_len x hidden_size)
hidden_init: Not used in the transformer decoder
Returns:
output: Un-normalized scores for each token in the vocabulary, across a batch for all the decoding time steps. (batch_size x decoder_seq_len x vocab_size)
attentions: The stacked attention weights applied to the encoder annotations (batch_size x encoder_seq_len x decoder_seq_len)
"""
batch_size, seq_len = inputs.size()
embed = self.embedding(inputs) # batch_size x seq_len x hidden_size
# THIS LINE WAS ADDED AS A CORRECTION.
embed = embed + self.positional_encodings[:seq_len]
encoder_attention_weights_list = []
self_attention_weights_list = []
contexts = embed
for i in range(self.num_layers):
# ------------
# FILL THIS IN - START
# ------------
new_contexts, self_attention_weights = self.self_attentions[i](contexts, annotations, annotations) # batch_size x seq_len x hidden_size
residual_contexts = contexts + new_contexts
new_contexts, encoder_attention_weights = self.encoder_attentions[i](residual_contexts, annotations, annotations) # batch_size x seq_len x hidden_size
residual_contexts = residual_contexts + new_contexts
new_contexts = self.attention_mlps[i](residual_contexts)
contexts = residual_contexts + new_contexts
# ------------
# FILL THIS IN - END
# ------------
encoder_attention_weights_list.append(encoder_attention_weights)
self_attention_weights_list.append(self_attention_weights)
output = self.out(contexts)
encoder_attention_weights = torch.stack(encoder_attention_weights_list)
self_attention_weights = torch.stack(self_attention_weights_list)
return output, (encoder_attention_weights, self_attention_weights)
def create_positional_encodings(self, max_seq_len=1000):
"""Creates positional encodings for the inputs.
Arguments:
max_seq_len: a number larger than the maximum string length we expect to encounter during training
Returns:
pos_encodings: (max_seq_len, hidden_dim) Positional encodings for a sequence with length max_seq_len.
"""
pos_indices = torch.arange(max_seq_len)[..., None]
dim_indices = torch.arange(self.hidden_size//2)[None, ...]
exponents = (2*dim_indices).float()/(self.hidden_size)
trig_args = pos_indices / (10000**exponents)
sin_terms = torch.sin(trig_args)
cos_terms = torch.cos(trig_args)
pos_encodings = torch.zeros((max_seq_len, self.hidden_size))
pos_encodings[:, 0::2] = sin_terms
pos_encodings[:, 1::2] = cos_terms
pos_encodings = pos_encodings.cuda()
return pos_encodings
# + [markdown] id="29ZjkXTNrUKb" colab_type="text"
#
# ## Step 5: Training and analysis
# Now, train the following language model that's comprised of a (simplified) transformer encoder and transformer decoder.
# + id="SmoTgrDcr_dw" colab_type="code" outputId="d6cf0dac-34f5-4848-d54a-698df062b2d6" colab={"base_uri": "https://localhost:8080/", "height": 1000}
TEST_SENTENCE = 'the air conditioning is working'
args = AttrDict()
args_dict = {
'cuda':True,
'nepochs':100,
'checkpoint_dir':"checkpoints",
'learning_rate':0.0005, ## INCREASE BY AN ORDER OF MAGNITUDE
'lr_decay':0.99,
'batch_size':64,
'hidden_size':20,
'encoder_type': 'transformer',
'decoder_type': 'transformer', # options: rnn / rnn_attention / transformer
'num_transformer_layers': 3,
}
args.update(args_dict)
print_opts(args)
transformer_encoder, transformer_decoder = train(args)
translated = translate_sentence(TEST_SENTENCE, transformer_encoder, transformer_decoder, None, args)
print("source:\t\t{} \ntranslated:\t{}".format(TEST_SENTENCE, translated))
# + id="R18s80gzC6A8" colab_type="code" outputId="1f405b47-239f-4b08-f693-9690142e8a76" colab={"base_uri": "https://localhost:8080/", "height": 51}
TEST_SENTENCE = 'the air conditioning is working'
translated = translate_sentence(TEST_SENTENCE, transformer_encoder, transformer_decoder, None, args)
print("source:\t\t{} \ntranslated:\t{}".format(TEST_SENTENCE, translated))
# + [markdown] id="MBnBXRG8mvcn" colab_type="text"
# # Optional: Attention Visualizations
#
# One of the benefits of using attention is that it allows us to gain insight into the inner workings of the model.
#
# By visualizing the attention weights generated for the input tokens in each decoder step, we can see where the model focuses while producing each output token.
#
# The code in this section loads the model you trained from the previous section and uses it to translate a given set of words: it prints the translations and display heatmaps to show how attention is used at each step.
# + [markdown] id="JqEC0vN9mvpV" colab_type="text"
# ## Step 1: Visualize Attention Masks
# Play around with visualizing attention maps generated by the previous two models you've trained. Inspect visualizations in one success and one failure case for both models.
# + id="Dkfz-u-MtudL" colab_type="code" colab={}
TEST_WORD_ATTN = 'street'
visualize_attention(TEST_WORD_ATTN, rnn_attn_encoder, rnn_attn_decoder, None, args)
# + id="Ssa7g35zt2yj" colab_type="code" colab={}
TEST_WORD_ATTN = 'street'
visualize_attention(TEST_WORD_ATTN, transformer_encoder, transformer_decoder, None, args, )
# + id="owstslMF-wdN" colab_type="code" colab={}
| Programming Assignment/PA3/nmt.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="8XZPNwc4gdSZ" colab_type="text"
# Note: The inspiration for the structural model of code has been taken from https://github.com/ananyahjha93/cycle-consistent-vae. Although all the models, dataloaders and analysis based code are purely original.
# + [markdown] id="QFnTELfByKPm" colab_type="text"
# All the console outputs are logged into 'Log' folder as 'txt' files.
#
# + id="g2YoVQRCWSMa" colab_type="code" colab={}
import os
from google.colab import drive
# + id="fq9PddBuXJwT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="82e8eec8-fff4-4152-cc0f-f9b2ef683356" executionInfo={"status": "ok", "timestamp": 1589632142494, "user_tz": -330, "elapsed": 1135, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "17863093696646821487"}}
drive.mount('/content/drive/')
os.chdir('/content/drive/My Drive/DL_Group/Assignments/Assignment_3')
# + [markdown] id="GHH4c6jMLUzD" colab_type="text"
# #IMPORT LIBS
#
# + id="af7ZzByQ7lL5" colab_type="code" colab={}
import h5py
import numpy as np
import numpy as np
import matplotlib.pyplot as plt
import os
import scipy.io
from mpl_toolkits.axes_grid1 import ImageGrid
import random
import torch
from torch.utils.data import DataLoader,TensorDataset
from torch.autograd import Variable
import torch.nn as nn
import torch.nn.functional as F
from itertools import cycle
import h5py
import time
from matplotlib import gridspec
from sklearn.utils import shuffle
import matplotlib.gridspec as gridspec
device=torch.device('cuda' if torch.cuda.is_available() else 'cpu')
# + [markdown] id="GrwS-22yLaHf" colab_type="text"
# Dataset Processing
#
# + id="w5zXxpIaXKrg" colab_type="code" colab={}
def load_data(data_path):
data=h5py.File(data_path,'r')
x=data['x']
y=data['y']
x=np.array(x,dtype=np.float16)
y=np.array(y)
return (x,y)
# + id="PwESM3MuIhVY" colab_type="code" colab={}
traina,trainb=load_data('Datasets/Q2/train_data.h5')
# + id="pCwv3OCqiSF2" colab_type="code" colab={}
a,b=load_data('Datasets/Q2/val_data.h5')
# + id="kRurZHb3IhtT" colab_type="code" colab={}
testa,testb=load_data('Datasets/Q2/test_data.h5')
# + [markdown] id="o6obkwtgaz6y" colab_type="text"
# #HELPER FUNCTIONS
#
#
#
#
# + id="t93eZqvwdSya" colab_type="code" colab={}
import torch
import torch.nn as nn
import matplotlib.pyplot as plt
from torch.autograd import Variable
from mpl_toolkits.axes_grid1 import ImageGrid
from torchvision.transforms import Compose, ToTensor
# from PIL import Image
# compose a transform configuration
transform_config = Compose([ToTensor()])
def mse_loss(input, target):
return torch.sum((input - target).pow(2)) / input.data.nelement()
def l1_loss(input, target):
return torch.sum(torch.abs(input - target)) / input.data.nelement()
def l2_loss(pred,target):
loss=torch.sum((pred-target).pow(2))/pred.data.nelement()
return loss
def reparameterize(training, mu, logvar):
if training:
std = logvar.mul(0.5).exp_()
eps = Variable(std.data.new(std.size()).normal_())
return eps.mul(std).add_(mu)
else:
return mu
def weights_init(layer):
if isinstance(layer, nn.Conv2d):
layer.weight.data.normal_(0.0, 0.05)
layer.bias.data.zero_()
elif isinstance(layer, nn.BatchNorm2d):
layer.weight.data.normal_(1.0, 0.02)
layer.bias.data.zero_()
elif isinstance(layer, nn.Linear):
layer.weight.data.normal_(0.0, 0.05)
layer.bias.data.zero_()
def accuracy(pred,y):
count=0
for i in range(len(pred)):
idx=torch.argmax(pred[i])
idx_class=torch.argmax(y[i])
if idx.item()==idx_class.item():
count+=1
return count/len(y)
def imshow_grid(images, shape=[2, 8], name='default', save=False):
"""
Plot images in a grid of a given shape.
Initial code from: https://github.com/pumpikano/tf-dann/blob/master/utils.py
"""
fig = plt.figure(1)
grid = ImageGrid(fig, 111, nrows_ncols=shape, axes_pad=0.05)
size = shape[0] * shape[1]
for i in range(size):
grid[i].axis('off')
# print(images[i])
grid[i].imshow(images[i]) # The AxesGrid object work as a list of axes.
if save:
plt.savefig('reconstructed_images/' + str(name) + '.png')
plt.clf()
else:
plt.show()
# + [markdown] id="Qgl8fISOJUqB" colab_type="text"
# DATA LOADING FOR THE MODEL
#
#
# + id="XEc8bsPqcvXr" colab_type="code" colab={}
import random
import numpy as np
from itertools import cycle
from torchvision import datasets,transforms
from torch.utils.data import Dataset, DataLoader
class ToTensor(object):
"""Convert ndarrays in sample to Tensors."""
def __call__(self, sample):
image, landmarks = sample[0], sample[1]
# swap color axis because
# numpy image: H x W x C
# torch image: C X H X W
image = image.transpose((2, 0, 1))
return (torch.from_numpy(image),torch.from_numpy(landmarks))
class MNIST_Paired():
def __init__(self, x,y=[],train=True, transform=None):
self.dat=x
self.data_dict = {}
for i in range(self.__len__()):
image,label = self.dat[i]
try:
self.data_dict[label.item()]
except KeyError:
self.data_dict[label.item()] = []
self.data_dict[label.item()].append(image)
def __len__(self):
return len(self.dat)
def __getitem__(self, index):
image= self.dat[index][0]
label=self.dat[index][1]
# return another image of the same class randomly selected from the data dictionary
# this is done to simulate pair-wise labeling of data
return image, random.SystemRandom().choice(self.data_dict[label.item()]), label
# + [markdown] id="hpXMEbSBa75B" colab_type="text"
# MODEL DEFINITION
# + id="kpYufqtzAz76" colab_type="code" colab={}
import torch
import torch.nn as nn
import torch.nn.functional as F
from torchvision import datasets
from torch.autograd import Variable
from torch.utils.data import DataLoader
from itertools import cycle
from collections import OrderedDict
# from utils import reparameterize, transform_config
class Encoder(nn.Module):
def __init__(self, style_dim, class_dim):
super(Encoder, self).__init__()
self.conv_model = nn.Sequential(OrderedDict([
('convolution_1',
nn.Conv2d(in_channels=3, out_channels=16, kernel_size=5, stride=2, padding=1, bias=True)),
('convolution_1_in', nn.InstanceNorm2d(num_features=16, track_running_stats=True)),
('ReLU_1', nn.ReLU(inplace=True)),
('convolution_2',
nn.Conv2d(in_channels=16, out_channels=32, kernel_size=5, stride=2, padding=1, bias=True)),
('convolution_2_in', nn.InstanceNorm2d(num_features=32, track_running_stats=True)),
('ReLU_2', nn.ReLU(inplace=True)),
('convolution_3',
nn.Conv2d(in_channels=32, out_channels=64, kernel_size=5, stride=2, padding=1, bias=True)),
('convolution_3_in', nn.InstanceNorm2d(num_features=64, track_running_stats=True)),
('ReLU_3', nn.ReLU(inplace=True)),
('convolution_4',
nn.Conv2d(in_channels=64, out_channels=128, kernel_size=5, stride=2, padding=1, bias=True)),
('convolution_4_in', nn.InstanceNorm2d(num_features=128, track_running_stats=True)),
('ReLU_4', nn.ReLU(inplace=True))
]))
# Style embeddings
self.style_mu = nn.Linear(in_features=512, out_features=style_dim, bias=True)
self.style_logvar = nn.Linear(in_features=512, out_features=style_dim, bias=True)
# Class embeddings
self.class_output = nn.Linear(in_features=512, out_features=class_dim, bias=True)
def forward(self, x):
x = self.conv_model(x)
x = x.view(x.size(0), x.size(1) * x.size(2) * x.size(3))
style_embeddings_mu = self.style_mu(x)
style_embeddings_logvar = self.style_logvar(x)
class_embeddings = self.class_output(x)
return style_embeddings_mu, style_embeddings_logvar, class_embeddings
class Decoder(nn.Module):
def __init__(self, style_dim, class_dim):
super(Decoder, self).__init__()
# Style embeddings input
self.style_input = nn.Linear(in_features=style_dim, out_features=512, bias=True)
# Class embeddings input
self.class_input = nn.Linear(in_features=class_dim, out_features=512, bias=True)
self.deconv_model = nn.Sequential(OrderedDict([
('deconvolution_1',
nn.ConvTranspose2d(in_channels=256, out_channels=64, kernel_size=4, stride=2, padding=0, bias=True)),
('deconvolution_1_in', nn.InstanceNorm2d(num_features=64, track_running_stats=True)),
('LeakyReLU_1', nn.LeakyReLU(negative_slope=0.2, inplace=True)),
('deconvolution_2',
nn.ConvTranspose2d(in_channels=64, out_channels=32, kernel_size=4, stride=2, padding=0, bias=True)),
('deconvolution_2_in', nn.InstanceNorm2d(num_features=32, track_running_stats=True)),
('LeakyReLU_2', nn.LeakyReLU(negative_slope=0.2, inplace=True)),
('deconvolution_3',
nn.ConvTranspose2d(in_channels=32, out_channels=16, kernel_size=4, stride=2, padding=0, bias=True)),
('deconvolution_3_in', nn.InstanceNorm2d(num_features=16, track_running_stats=True)),
('LeakyReLU_3', nn.LeakyReLU(negative_slope=0.2, inplace=True)),
('deconvolution_4',
nn.ConvTranspose2d(in_channels=16, out_channels=3, kernel_size=4, stride=2, padding=1, bias=True)),
('sigmoid_final', nn.Sigmoid())
]))
def forward(self, style_embeddings, class_embeddings):
style_embeddings = F.leaky_relu_(self.style_input(style_embeddings), negative_slope=0.2)
class_embeddings = F.leaky_relu_(self.class_input(class_embeddings), negative_slope=0.2)
x = torch.cat((style_embeddings, class_embeddings), dim=1)
x = x.view(x.size(0), 256, 2, 2)
x = self.deconv_model(x)
return x
class Classifier(nn.Module):
def __init__(self, z_dim, num_classes):
super(Classifier, self).__init__()
self.fc_model = nn.Sequential(OrderedDict( [
('fc_1', nn.Linear(in_features=z_dim, out_features=256, bias=True)),
('fc_1_bn', nn.BatchNorm1d(num_features=256)),
('LeakyRelu_1', nn.LeakyReLU(negative_slope=0.2, inplace=True)),
('fc_2', nn.Linear(in_features=256, out_features=256, bias=True)),
('fc_2_bn', nn.BatchNorm1d(num_features=256)),
('LeakyRelu_2', nn.LeakyReLU(negative_slope=0.2, inplace=True)),
('fc_3', nn.Linear(in_features=256, out_features=num_classes, bias=True))
]))
def forward(self, z):
x = self.fc_model(z)
return x
# + [markdown] id="j6NzeTBXXaBE" colab_type="text"
# #**TRAINING ONLY**
#
#
#
# + id="g80vaBv30vZJ" colab_type="code" outputId="02213d75-98a4-4678-832b-7089b476d381" executionInfo={"status": "ok", "timestamp": 1589607213404, "user_tz": -330, "elapsed": 88299, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "17863093696646821487"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
def cycle(iterable):
iterator = iter(iterable)
while True:
try:
yield next(iterator)
except StopIteration:
iterator = iter(iterable)
a=np.moveaxis(a,0,-1)
a=a/255.0
a=np.moveaxis(a,-1,0)
print(a.shape)
# val_Data = MNIST_Paired(a,b,transform=transforms.Compose([ToTensor()]))
val_f=TensorDataset(torch.from_numpy(a).to(device),torch.from_numpy(b).to(device))
val_Data=MNIST_Paired(val_f)
val_loader = cycle(DataLoader(val_Data, batch_size=64, shuffle=False, num_workers=0, drop_last=True))
# + id="9D7wQmEcIcvt" colab_type="code" outputId="a8bafbdf-ec77-4168-cfd8-4e99c420e868" executionInfo={"status": "ok", "timestamp": 1589607238232, "user_tz": -330, "elapsed": 112569, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "17863093696646821487"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
traina=traina/255.0
traina=traina.astype('float')
print(traina.shape)
# train_Data = MNIST_Paired(traina,trainb,transform=transforms.Compose([ToTensor()]))
# train_loader = cycle(DataLoader(train_Data, batch_size=64, shuffle=False,drop_last=True))
train_df=TensorDataset(torch.from_numpy(traina),torch.from_numpy(trainb))
train_Data = MNIST_Paired(train_df)
train_loader = cycle(DataLoader(train_Data, batch_size=64, shuffle=False,drop_last=True))
# + id="JXP9VrQkIc7y" colab_type="code" colab={}
# testa=testa/255.0
# testa=testa.astype('float')
# print(testa.shape)
# test_Data = MNIST_Paired(testa,testb,transform=transforms.Compose([ToTensor()]))
# test_loader = cycle(DataLoader(test_Data, batch_size=64, shuffle=False, num_workers=0, drop_last=True))
# + id="L4kPaZp0Fw_5" colab_type="code" colab={}
batch_size=64
train_batches=len(train_Data)//batch_size
# + id="Jd1W6leuIXGd" colab_type="code" colab={}
val_batches=len(val_Data)//batch_size
# + id="bRkJgXDRHD68" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 302} outputId="0f5eba67-bac7-4157-fb2b-31f12cda21e0" executionInfo={"status": "ok", "timestamp": 1589607238254, "user_tz": -330, "elapsed": 103619, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "17863093696646821487"}}
import os
import numpy as np
from itertools import cycle
import torch
import torch.nn as nn
import torch.optim as optim
from torch.autograd import Variable
from torch.utils.data import DataLoader
# from utils import imshow_grid, mse_loss, reparameterize, l1_loss
# FLAGS={}
image_size=60
num_channels=3 #RGB
initial_learning_rate=0.0001
style_dim=512
class_dim=512
num_classes=672
reconstruction_coef=2
reverse_cycle_coef=10
kl_divergence_coef=3
beta_1=0.9
beta_2=0.999
encoder_save='encoderwts'
decoder_save='decoderwts'
log_file='Outputs/Q2/logs.txt'
load_saved=False
# print(FLAGS)
epochs=100
encoder = Encoder(style_dim,class_dim).to(device)
encoder.apply(weights_init)
decoder = Decoder(style_dim,class_dim).to(device)
decoder.apply(weights_init)
########### if saved and want to finetune
# if load_saved:
# encoder.load_state_dict(torch.load(os.path.join('Outputs/Q2/checkpoints', encoder_save)))
# decoder.load_state_dict(torch.load(os.path.join('Outputs/Q2/checkpoints', decoder_save)))
# + id="aoLRPjTxIvOE" colab_type="code" colab={}
reconstruction_loss_list,kl_div_loss_list,reverse_cycle_loss_list=[],[],[]
x1 = torch.FloatTensor(batch_size, num_channels, image_size, image_size).to(device)
x2 = torch.FloatTensor(batch_size, num_channels, image_size, image_size).to(device)
x3 = torch.FloatTensor(batch_size, num_channels, image_size, image_size).to(device)
style_latent_space = torch.FloatTensor(batch_size, style_dim).to(device)
forward_optimizer = optim.Adam(list(encoder.parameters()) + list(decoder.parameters()),lr=initial_learning_rate)
reverse_optimizer = optim.Adam(list(encoder.parameters()),lr=initial_learning_rate)
forward_optim_scheduler = optim.lr_scheduler.StepLR(forward_optimizer, step_size=80, gamma=0.1)
reverse_optim_scheduler = optim.lr_scheduler.StepLR(reverse_optimizer, step_size=80, gamma=0.1)
# load_saved is false when training is started from 0th iteration
# if not FLAGS.load_saved:
# with open(FLAGS.log_file, 'w') as log:
# log.write('Epoch\tIteration\tReconstruction_loss\tKL_divergence_loss\tReverse_cycle_loss\n')
# + id="8hA9WYiDm_eZ" colab_type="code" colab={}
if not os.path.exists('Outputs/Q2/checkpoints/encoder_weights_new'):
os.makedirs('Outputs/Q2/checkpoints/encoder_weights_new')
if not os.path.exists('Outputs/Q2/checkpoints/decoder_weights_new'):
os.makedirs('Outputs/Q2/checkpoints/decoder_weights_new')
if not os.path.exists('Outputs/Q2/reconstructed_images'):
os.makedirs('Outputs/Q2/reconstructed_images')
# + id="hT23ntioHVuC" colab_type="code" colab={}
#disconnected colab at 97th epoch running from last checkpoint
# encoder.load_state_dict(torch.load('Outputs/Q2/checkpoints/encoder_weights_new/encoder99.pt'))
# decoder.load_state_dict(torch.load('Outputs/Q2/checkpoints/decoder_weights_new/decoder99.pt'))
# epochs=100
# + id="f0p_0DmMTw2-" colab_type="code" outputId="63455a27-6dcf-4714-9cb6-70e61eee9346" colab={"base_uri": "https://localhost:8080/", "height": 1000} executionInfo={"status": "ok", "timestamp": 1589621724262, "user_tz": -330, "elapsed": 6150342, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "17863093696646821487"}}
for epoch in range(epochs):
print('')
start=time.time()
print('Epoch #' + str(epoch+100) + '..........................................................................')
loss1=0
loss2=0
loss3=0
# update the learning rate scheduler
forward_optim_scheduler.step()
reverse_optim_scheduler.step()
for iteration in range(train_batches):
forward_optimizer.zero_grad()
image_batch_1, image_batch_2, _ = next(train_loader)
# forward go
x1.copy_(image_batch_1)
x2.copy_(image_batch_2)
style_mu_1,style_logvar_1,class_latent_space_1=encoder(Variable(x1))
style_latent_space_1=reparameterize(training=False,mu=style_mu_1,logvar=style_logvar_1)
kl_divergence_loss_1 = kl_divergence_coef * (-0.5*torch.sum(1+style_logvar_1-style_mu_1.pow(2)-style_logvar_1.exp()))/(batch_size*num_channels *image_size * image_size)
kl_divergence_loss_1.backward(retain_graph=True)
style_mu_2,style_logvar_2,class_latent_space_2=encoder(Variable(x2))
style_latent_space_2=reparameterize(training=False,mu=style_mu_2,logvar=style_logvar_2)
kl_divergence_loss_2 = kl_divergence_coef *(- 0.5 * torch.sum(1 + style_logvar_2 - style_mu_2.pow(2) - style_logvar_2.exp()))/(batch_size * num_channels * image_size * image_size)
kl_divergence_loss_2.backward(retain_graph=True)
reconstructed_x1=decoder(style_latent_space_1, class_latent_space_2)
reconstructed_x2=decoder(style_latent_space_2, class_latent_space_1)
reconstruction_error_1=reconstruction_coef*l2_loss(reconstructed_x1,Variable(x1))
reconstruction_error_1.backward(retain_graph=True)
reconstruction_error_2=reconstruction_coef*l2_loss(reconstructed_x2,Variable(x2))
reconstruction_error_2.backward()
reconstruction_error = (reconstruction_error_1+reconstruction_error_2)
reconstruction_error/=reconstruction_coef
kl_divergence_error=(kl_divergence_loss_1 + kl_divergence_loss_2)
kl_divergence_error/=kl_divergence_coef
forward_optimizer.step()
# reverse cycle
reverse_optimizer.zero_grad()
image_batch_1, _, __=next(train_loader)
image_batch_2, _, __=next(train_loader)
style_latent_space.normal_(0., 1.)
x1.copy_(image_batch_1)
x2.copy_(image_batch_2)
_, __, class_latent_space_1=encoder(Variable(x1))
_, __, class_latent_space_2=encoder(Variable(x2))
reconstructed_x2=decoder(Variable(style_latent_space),class_latent_space_2.detach())
style_mu_2, style_logvar_2,_=encoder(reconstructed_x2)
style_latent_space_2=reparameterize(training=False, mu=style_mu_2, logvar=style_logvar_2)
reconstructed_x1=decoder(Variable(style_latent_space),class_latent_space_1.detach())
style_mu_1, style_logvar_1,_=encoder(reconstructed_x1)
style_latent_space_1=reparameterize(training=False, mu=style_mu_1, logvar=style_logvar_1)
reverse_cycle_loss=reverse_cycle_coef*l1_loss(style_latent_space_1,style_latent_space_2)
reverse_cycle_loss.backward()
reverse_cycle_loss/=reverse_cycle_coef
reverse_optimizer.step()
loss1+=reconstruction_error.data.storage().tolist()[0]
loss2+=kl_divergence_error.data.storage().tolist()[0]
loss3+=reverse_cycle_loss.data.storage().tolist()[0]
reverse_cycle_loss_list.append(loss3/train_batches)
kl_div_loss_list.append(loss2/train_batches)
reconstruction_loss_list.append(loss1/train_batches)
# save model after every 5 epochs
if (epoch + 1) % 5 == 0 or (epoch + 1) == epochs:
torch.save(encoder.state_dict(), 'Outputs/Q2/checkpoints/encoder_weights_new/encoder'+str(epoch+100)+'.pt')
torch.save(decoder.state_dict(), 'Outputs/Q2/checkpoints/decoder_weights_new/decoder'+str(epoch+100)+'.pt')
print('Epoch ',epoch+1+100,'/',200,' epoch_duration:',str(time.time()-start),'s',' reconstruction_loss:',str(loss1/train_batches),' kl_div_loss:',str(loss2/train_batches),' reverse_cycle_loss:',str(loss3/train_batches))
# + id="XKE21_SyN2P_" colab_type="code" colab={}
## OUTPUT OF THE TRAIN LOGGED AT log.txt
# + id="9OHExbjL68Mi" colab_type="code" colab={}
reconstruction_loss_list=['8.686999438088488e-05', '4.269123199251782e-05', '2.1179927396116924e-05', '1.0709047545061765e-05', '5.6343378006374265e-06', '3.188040495646576e-06', '2.02006064131533e-06', '1.463808805608927e-06', '1.1958591330243926e-06', '1.0627330056990196e-06', '9.940594822977892e-07', '9.575120591405642e-07', '9.374113942318454e-07', '9.258824681258132e-07', '9.189525180380174e-07', '9.144328097993317e-07', '9.112714108779472e-07', '9.08837065538091e-07', '9.068687751773363e-07', '9.052816743297895e-07', '9.039488737625165e-07', '9.027927261014555e-07', '9.018257205000429e-07', '9.009862176724369e-07', '9.002361025931932e-07', '8.996191514015401e-07', '8.990166013469245e-07', '8.984561278951025e-07', '8.978820072789865e-07', '8.97442961920644e-07', '8.970996717959397e-07', '8.96716516072963e-07', '8.963562669800562e-07', '8.960511599702153e-07', '8.9576371821828e-07', '8.954733684551828e-07', '8.952041250624539e-07', '8.950332957849515e-07', '8.948145313520039e-07', '8.945997608030967e-07', '8.944161137026921e-07', '8.942379519513132e-07', '8.940749799473955e-07', '8.939443301338058e-07', '8.937733140091231e-07', '8.936592314757748e-07', '8.935118760765892e-07', '8.933755950840232e-07', '8.933095619654818e-07', '8.931808614313189e-07', '8.930786031060902e-07', '8.929968376921463e-07', '8.928584272387839e-07', '8.927694156384552e-07', '8.926644725453908e-07', '8.925860888241344e-07', '8.924989106080895e-07', '8.924600879843166e-07', '8.92386941300403e-07', '8.922977178254317e-07', '8.92263050782734e-07', '8.921467239975991e-07', '8.920679044222721e-07', '8.919822654360418e-07', '8.919161582983722e-07', '8.918503328014405e-07', '8.918063267121803e-07', '8.917286277292013e-07', '8.916866463289108e-07', '8.916302924995539e-07', '8.915765838521124e-07', '8.915492647005351e-07', '8.877864256434065e-07', '8.876439422159518e-07', '8.876295590725145e-07', '8.87617662644627e-07', '8.876070513278457e-07', '8.875973987836941e-07', '8.875795577201355e-07', '8.875717077257799e-07', '8.875638150375184e-07', '8.875565515527407e-07', '8.875495381381115e-07', '8.875427159872737e-07', '8.875364376521077e-07', '8.875309498085149e-07', '8.87523862783703e-07', '8.875185907815236e-07', '8.875121722189597e-07', '8.875064426068113e-07', '8.87501758177468e-07', '8.87495939987733e-07', '8.874913145692196e-07', '8.874865324428062e-07', '8.874822563863987e-07', '8.874764373378783e-07']
# + id="rT_QShFMOPJp" colab_type="code" colab={}
reverse_cycle_loss_list=['4.707272468811739e-06', '2.6501810468367996e-06', '1.4749440365269182e-06', '8.054847523172291e-07', '4.525515653627938e-07', '2.708283743800984e-07', '1.5085091391858323e-07', '9.641950230398645e-08', '8.847828442342381e-08', '6.55811960554832e-08', '5.6773074667348214e-08', '5.21840590371819e-08', '4.990178399229123e-08', '4.802152098448937e-08', '6.681784010295122e-07', '1.9332751661228265e-07', '1.265028668510106e-07', '1.0344338351908701e-07', '9.71053237857658e-08', '9.157948052078734e-08', '8.11206840891579e-08', '8.538353690522095e-08', '7.310827568972763e-08', '6.748690528044083e-08', '6.622117333273086e-08', '5.9342840532517484e-08', '5.617290101524464e-08', '5.5106348085178397e-08', '5.0582306895548196e-08', '4.822717490971362e-08', '4.623837549710803e-08', '4.466634141945522e-08', '4.296147278821389e-08', '4.094390084975962e-08', '3.921948738338346e-08', '3.777341974812349e-08', '3.605224368825328e-08', '3.4731797741584295e-08', '3.3302909738940894e-08', '3.2038269880080337e-08', '3.0755434332938336e-08', '2.946532622626333e-08', '2.8067926966488104e-08', '2.6615413253024544e-08', '2.5198999341845146e-08', '2.3909380318604162e-08', '2.253214889042234e-08', '2.1176340311666186e-08', '1.979399154064392e-08', '1.8455478375909585e-08', '1.71285889529525e-08', '1.58348342221078e-08', '1.4578857620409244e-08', '1.339639200274127e-08', '1.2247314902603596e-08', '1.1129319790927762e-08', '1.0060414945875806e-08', '9.078821535964052e-09', '8.135074069857204e-09', '7.290312463706712e-09', '6.52487921950112e-09', '5.82191203032594e-09', '5.201716388081392e-09', '4.644014358679404e-09', '4.152162998399623e-09', '3.734565138989804e-09', '3.3612490646932165e-09', '3.035969489875024e-09', '2.7653941772800417e-09', '2.533018675385785e-09', '2.3524644492991937e-09', '2.193701717800627e-09', '2.0699441898340583e-09', '2.0522661517439184e-09', '2.038994847943803e-09', '2.0282522028010513e-09', '2.015443019350916e-09', '2.0033219567640286e-09', '1.979950767048299e-09', '1.9706624846748515e-09', '1.9592423376725297e-09', '1.9501554764330503e-09', '1.9385394277026357e-09', '1.930051341449584e-09', '1.9201938518048913e-09', '1.9119587187657706e-09', '1.902277260972044e-09', '1.891916719690234e-09', '1.881838760359868e-09', '1.871549126147243e-09', '1.8645449718962498e-09', '1.856300657712076e-09', '1.8485684437222863e-09', '1.842696810566582e-09', '1.8354789843897124e-09', '1.8267566922765197e-09']
# + id="bBTAXypDOXN0" colab_type="code" colab={}
kl_div_loss_list=['7.443934558857023e-07', '5.076582914030336e-07', '3.63301266769432e-07', '2.76420196690805e-07', '2.4651075964831383e-07', '2.7875008207598617e-07', '3.0789541179653737e-07', '3.605410653756853e-07', '4.143392753079876e-07', '4.2704175105389945e-07', '4.992279627299958e-07', '5.345434403283853e-07', '5.002465674643684e-07', '4.000425379159561e-07', '4.873048310617659e-07', '1.6925274307960607e-07', '1.236494621353051e-07', '9.752275688775926e-08', '9.381564365522493e-08', '1.1349518633684586e-07', '7.371429386254192e-08', '2.7326041259535935e-07', '1.722071698017392e-07', '2.056276956413918e-07', '2.6753034815579737e-07', '1.0328117336522489e-07', '1.0582008432103923e-07', '1.4445032183410233e-07', '8.421658749936956e-08', '6.987070748163788e-08', '6.414850094185842e-08', '6.031600562187777e-08', '5.925637496016557e-08', '5.2439107407774566e-08', '4.860964064840964e-08', '5.316329406565787e-08', '4.5238488610470834e-08', '4.432438050551325e-08', '4.2539275495899184e-08', '4.177832016837214e-08', '4.1438524122235575e-08', '4.2460512776770647e-08', '4.0021676621521e-08', '3.9396676864376795e-08', '3.8663511865161697e-08', '3.8517822744921455e-08', '3.743467412312788e-08', '3.666546753629273e-08', '3.6273676055819174e-08', '3.584918586126549e-08', '3.5263152221744006e-08', '3.477396607723413e-08', '3.433776850900083e-08', '3.4844314588902395e-08', '3.4051052193083756e-08', '3.3744188335657844e-08', '3.349724857269544e-08', '3.32443361996367e-08', '3.3005249549239006e-08', '3.2896913697678356e-08', '3.272922082098732e-08', '3.24403577279896e-08', '3.246739855767692e-08', '3.224058896243111e-08', '3.21558567923099e-08', '3.199498556549853e-08', '3.1933765778464996e-08', '3.1815430512658084e-08', '3.179021454585347e-08', '3.1592735423749827e-08', '3.153815368925399e-08', '3.14687145598275e-08', '1.900816982391334e-09', '1.9260805082749227e-09', '2.0739896987659045e-09', '2.1303050314987283e-09', '2.1279555076458765e-09', '2.114566139135075e-09', '2.0854353239489425e-09', '2.0739751827196656e-09', '2.063673061053923e-09', '2.051614444678307e-09', '2.0418082654683323e-09', '2.0313635152876756e-09', '2.0220653042456294e-09', '2.013666918102428e-09', '2.0048476226885594e-09', '1.9964421435780572e-09', '1.9877647494430854e-09', '1.979247102343857e-09', '1.9705849113154444e-09', '1.963677817313977e-09', '1.955609707344891e-09', '1.9484894812734767e-09', '1.9409863518207987e-09', '1.9333334337932912e-09']
# + id="hLNa3wKQOx0P" colab_type="code" colab={}
for i in range(96):
reconstruction_loss_list[i]=float(reconstruction_loss_list[i])
reverse_cycle_loss_list[i]=float(reverse_cycle_loss_list[i])
kl_div_loss_list[i]=float(kl_div_loss_list[i])
# + id="FQ8tSnLxUAzd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 868} outputId="e8e52fe8-0ed9-401f-efc8-a00a6ea42437" executionInfo={"status": "ok", "timestamp": 1589623360720, "user_tz": -330, "elapsed": 1886, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "17863093696646821487"}}
plt.figure()
plt.title('reverse_cycle_loss')
plt.xlabel('epochs')
plt.ylabel('loss')
plt.plot(reverse_cycle_loss_list)
plt.figure()
plt.title('reconstruction_loss')
plt.xlabel('epochs')
plt.ylabel('loss')
plt.plot(reconstruction_loss_list)
plt.figure()
plt.title('kl_div_loss')
plt.xlabel('epochs')
plt.ylabel('loss')
plt.plot(kl_div_loss_list)
# + [markdown] id="OmWXJWs6wt5l" colab_type="text"
# #QUESTION 5 - UNSPECIFIED TO SPECIFIED
# + [markdown] id="xG0NK5pKbApT" colab_type="text"
# THIS LOADING IS FOR REST OF THE Qs
#
# + id="VFgBKBCJtkgK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="794dea51-9b4f-4ce9-aa31-2750b7d37a6e" executionInfo={"status": "ok", "timestamp": 1589636050204, "user_tz": -330, "elapsed": 29192, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "17863093696646821487"}}
a=a/255.0
a=a.astype('float')
print(a.shape)
val_d=TensorDataset(torch.from_numpy(a),torch.from_numpy(b))
val_Data = MNIST_Paired(val_d)
val_loader = (DataLoader(val_Data, batch_size=64, shuffle=True, num_workers=0, drop_last=True))
traina=traina/255.0
traina=traina.astype('float')
print(traina.shape)
# train_Data = MNIST_Paired(traina,trainb,transform=transforms.Compose([ToTensor()]))
# train_loader = cycle(DataLoader(train_Data, batch_size=64, shuffle=False,drop_last=True))
train_df=TensorDataset(torch.from_numpy(traina),torch.from_numpy(trainb))
train_Data = MNIST_Paired(train_df)
train_loader = (DataLoader(train_Data, batch_size=64, shuffle=True,drop_last=True))
# testa=testa/255.0
# testa=testa.astype('float')
# print(testa.shape)
# test_d=TensorDataset(torch.from_numpy(testa),torch.from_numpy(testb))
# test_Data = MNIST_Paired(test_d)
# test_loader = (DataLoader(test_Data, batch_size=64, shuffle=True, num_workers=0, drop_last=True))
# + [markdown] id="HuWZSjLrcoes" colab_type="text"
# bELOW Loader is req in Q2
# + id="H3Dj3bXDcl7d" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="dc63ce82-9de8-48d8-d604-0fda06668d19" executionInfo={"status": "ok", "timestamp": 1589630854116, "user_tz": -330, "elapsed": 97652, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "17863093696646821487"}}
testa=testa/255.0
testa=testa.astype('float')
print(testa.shape)
test_df=TensorDataset(torch.from_numpy(testa),torch.from_numpy(testb))
test_Data = MNIST_Paired(test_df)
test_loader = (DataLoader(test_Data, batch_size=64, shuffle=True, num_workers=0, drop_last=True))
# + id="ng29cY9jxMkm" colab_type="code" colab={}
batch_size=64
train_batches=len(train_Data)//batch_size
# + id="nxgI5TZ6xUBT" colab_type="code" colab={}
val_batches=len(val_Data)//batch_size
# + id="NgHX8dvhUVcD" colab_type="code" colab={}
class Predictor(nn.Module):
def __init__(self,in_dim,out_dim):
super(Predictor,self).__init__()
self.f1=nn.Linear(in_dim,256)
self.batch_norm1=nn.BatchNorm1d(num_features=256)
self.f2=nn.Linear(256,256)
self.batch_norm2=nn.BatchNorm1d(num_features=256)
self.f3=nn.Linear(256,out_dim)
def forward(self,x):
x=self.f1(x)
x=self.batch_norm1(x)
x=F.relu(x)
x=self.f2(x)
x=self.batch_norm2(x)
x=F.relu(x)
x=self.f3(x)
return x
# + id="NFQtLybctSJm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 134} outputId="86c5ab9b-c77d-45e8-f016-18a6fb925ead" executionInfo={"status": "ok", "timestamp": 1589625829859, "user_tz": -330, "elapsed": 1155, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "17863093696646821487"}}
image_size=60
num_channels=3 #RGB
initial_learning_rate=0.0001
style_dim=512
class_dim=512
num_classes=672
reconstruction_coef=2.
reverse_cycle_coef=10.
kl_divergence_coef=3.
beta_1=0.9
beta_2=0.999
log_file='Outputs/Q2/logs.txt'
load_saved=False
# print(FLAGS)
epochs=20
ztospredictor=Predictor(style_dim,class_dim).to(device)
encoder=Encoder(style_dim,class_dim).to(device)
decoder=Decoder(style_dim,class_dim).to(device)
encoder.load_state_dict(torch.load('Outputs/Q2/checkpoints/encoder_weights_new/encoder199.pt'))
decoder.load_state_dict(torch.load('Outputs/Q2/checkpoints/decoder_weights_new/decoder199.pt'))
print(ztospredictor)
# + id="NkoMRbVIu1b_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 857} outputId="7a27f0f3-5006-4a56-be92-2e149a2a05b4" executionInfo={"status": "ok", "timestamp": 1589627603117, "user_tz": -330, "elapsed": 882000, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "17863093696646821487"}}
epochs=50
train_loss_list=[]
val_loss_list=[]
criterion=nn.MSELoss()
optim=torch.optim.Adam(ztospredictor.parameters(),lr=0.01)
x1=torch.FloatTensor(batch_size,num_channels,image_size,image_size).to(device)
for epoch in range(epochs):
train_loss=0
train_acc=0
ztospredictor.train()
val_iterator=iter(val_loader)
train_iterator=iter(train_loader)
for i,bat in enumerate(train_iterator):
x=bat[0]
x1.copy_(x)
optim.zero_grad()
with torch.no_grad():
style_mu,style_logvar,class_latent=encoder(Variable(x1))
s_pred=ztospredictor(style_mu)
loss=criterion(s_pred,class_latent)
loss.backward()
optim.step()
train_loss+=loss.item()
ztospredictor.eval()
validation_loss=0
with torch.no_grad():
for i,bat in enumerate(val_iterator):
x=bat[0]
x1.copy_(x)
style_mu,style_logvar,class_latent=encoder(Variable(x1))
s_pred=ztospredictor(style_mu)
loss=criterion(s_pred,class_latent)
validation_loss+=loss.item()
print('Epoch: '+str(epoch+1)+'/'+str(epochs)+' loss: '+str(train_loss/train_batches)+' val_loss: '+str(validation_loss/val_batches))
train_loss_list.append(train_loss/train_batches)
val_loss_list.append(validation_loss/val_batches)
torch.save(ztospredictor.state_dict(),os.getcwd()+'/Outputs/Q2/checkpoints/predictor/ztospredictor.pt')
# + id="L3LnRLKjzApt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 312} outputId="8d94af1d-7d66-4cc5-8743-f516b71aeeaf" executionInfo={"status": "ok", "timestamp": 1589626203369, "user_tz": -330, "elapsed": 1702, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "17863093696646821487"}}
plt.figure()
plt.title('losses vs epochs')
plt.plot(val_loss_list,label='validation')
plt.plot(train_loss_list,label='train')
plt.xlabel('epochs')
plt.ylabel('losses')
plt.legend(loc='upper right')
# + id="0KkKrCu54zuQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="75673134-7e3f-499c-f9d1-322b1a949fd2" executionInfo={"status": "ok", "timestamp": 1589626585020, "user_tz": -330, "elapsed": 2236, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "17863093696646821487"}}
testa=testa/255.0
testa=testa.astype('float')
print(testa.shape)
test_d=TensorDataset(torch.from_numpy(testa),torch.from_numpy(testb))
test_Data = MNIST_Paired(test_d)
test_loader = (DataLoader(test_Data, batch_size=64, shuffle=True, num_workers=0, drop_last=True))
test_iterator=iter(test_loader)
# + id="sRBxXUpc1zI5" colab_type="code" colab={}
image_count=0
encoder.load_state_dict(torch.load('Outputs/Q2/checkpoints/encoder_weights_new/encoder199.pt'))
decoder.load_state_dict(torch.load('Outputs/Q2/checkpoints/decoder_weights_new/decoder199.pt'))
ztospredictor=Predictor(style_dim,class_dim).to(device)
ztospredictor.load_state_dict(torch.load('Outputs/Q2/checkpoints/predictor/ztospredictor.pt'))
image_batch=next(test_iterator)[0]
x1.copy_(image_batch)
style_mu,style_logvar,class_latent=encoder(Variable(x1))
s_pred=ztospredictor(style_mu)
reconstructed_img_batch_s=decoder(style_mu,class_latent)
reconstructed_img_batch_s_pred=decoder(style_mu,s_pred)
reconstruction_err=reconstruction_coef*l2_loss(reconstructed_img_batch_s,reconstructed_img_batch_s_pred)
gs=gridspec.GridSpec(8,8,width_ratios=[1,1,1,1,1,1,1,1],height_ratios=[1,1,1,1,1,1,1,1],wspace=0,hspace=0)
reconstructed_img=np.transpose(reconstructed_img_batch_s.cpu().data.numpy(),(0,2,3,1))
fig1=plt.figure(figsize=(8,8))
# fig1.suptitle('Image Reconstructions with encoder generated class-latent space')
for i in range(8):
for j in range(8):
if image_count<batch_size:
ax=plt.subplot(gs[i,j])
ax.axis('off')
ax.imshow(reconstructed_img[image_count])
image_count+=1
image_count=0
reconstructed_img=np.transpose(reconstructed_img_batch_s_pred.cpu().data.numpy(),(0,2,3,1))
fig2=plt.figure(figsize=(8,8))
# fig2.suptitle('Image Reconstructions with network generated class-latent space')
for i in range(8):
for j in range(8):
if image_count<batch_size:
ax=plt.subplot(gs[i,j])
ax.axis('off')
ax.imshow(reconstructed_img[image_count])
image_count+=1
print('Difference in reconstruction error: '+str(reconstruction_err.data.storage().tolist()[0]))
# + [markdown] id="3kNbusySDYQs" colab_type="text"
# # QUESTION 5 SPECIFIED TO UNSPECIFIED
# + id="e4c_nV1bAv0J" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 134} outputId="9d434581-50e9-420f-b1e1-92ecbd59f370" executionInfo={"status": "ok", "timestamp": 1589636050205, "user_tz": -330, "elapsed": 11984, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "17863093696646821487"}}
image_size=60
num_channels=3 #RGB
initial_learning_rate=0.0001
style_dim=512
class_dim=512
num_classes=672
reconstruction_coef=2.
reverse_cycle_coef=10.
kl_divergence_coef=3.
beta_1=0.9
beta_2=0.999
log_file='Outputs/Q2/logs.txt'
load_saved=False
# print(FLAGS)
epochs=20
stozpredictor=Predictor(class_dim,style_dim).to(device)
encoder=Encoder(style_dim,class_dim).to(device)
decoder=Decoder(style_dim,class_dim).to(device)
encoder.load_state_dict(torch.load('Outputs/Q2/checkpoints/encoder_weights_new/encoder199.pt'))
decoder.load_state_dict(torch.load('Outputs/Q2/checkpoints/decoder_weights_new/decoder199.pt'))
print(stozpredictor)
# + id="CLsYOl7hDzpT" colab_type="code" colab={}
train_loss_list=[]
val_loss_list=[]
criterion=nn.MSELoss()
optim=torch.optim.Adam(stozpredictor.parameters(),lr=0.01)
x1=torch.FloatTensor(batch_size,num_channels,image_size,image_size).to(device)
for epoch in range(epochs):
train_loss=0
stozpredictor.train()
val_iterator=iter(val_loader)
train_iterator=iter(train_loader)
for i,bat in enumerate(train_iterator):
x=bat[0]
x1.copy_(x)
optim.zero_grad()
with torch.no_grad():
style_mu,style_logvar,class_latent=encoder(Variable(x1))
z_pred=stozpredictor(class_latent)
loss=criterion(z_pred,style_mu)
loss.backward()
optim.step()
train_loss+=loss.item()
stozpredictor.eval()
validation_loss=0
with torch.no_grad():
for i,bat in enumerate(val_iterator):
x=bat[0]
x1.copy_(x)
style_mu,style_logvar,class_latent=encoder(Variable(x1))
z_pred=stozpredictor(class_latent)
loss=criterion(z_pred,style_mu)
validation_loss+=loss.item()
print('Epoch: '+str(epoch+1)+'/'+str(epochs)+' loss: '+str(train_loss/train_batches)+' val_loss: '+str(validation_loss/val_batches))
train_loss_list.append(train_loss/train_batches)
val_loss_list.append(validation_loss/val_batches)
torch.save(stozpredictor.state_dict(),os.getcwd()+'/Outputs/Q2/checkpoints/predictor/stozpredictor.pt')
# + id="eGwhe9TRHuXl" colab_type="code" colab={}
plt.figure()
plt.title('losses vs epochs')
plt.plot(val_loss_list,label='validation')
plt.plot(train_loss_list,label='train')
plt.xlabel('epochs')
plt.ylabel('losses')
plt.legend(loc='upper right')
# + id="fWedRCDBHzUN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="7f2c9a01-888a-4b66-f5f3-d947644b34a5" executionInfo={"status": "ok", "timestamp": 1589546804973, "user_tz": -330, "elapsed": 6017, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "17863093696646821487"}}
testa=testa/255.0
testa=testa.astype('float')
print(testa.shape)
test_d=TensorDataset(torch.from_numpy(testa),torch.from_numpy(testb))
test_Data = MNIST_Paired(test_d)
test_loader = (DataLoader(test_Data, batch_size=64, shuffle=True, num_workers=0, drop_last=True))
test_iterator=iter(test_loader)
# + id="LxcjabRJH34n" colab_type="code" colab={}
image_count=0
stozpredictor=Predictor(style_dim,class_dim).to(device)
stozpredictor.load_state_dict(torch.load('Outputs/Q2/checkpoints/predictor/stozpredictor.pt'))
image_batch=next(test_iterator)[0]
x1.copy_(image_batch)
style_mu,style_logvar,class_latent=encoder(Variable(x1))
z_pred=stozpredictor(class_latent)
reconstructed_img_batch_s=decoder(style_mu,class_latent)
reconstructed_img_batch_s_pred=decoder(z_pred,class_latent)
reconstruction_err=reconstruction_coef*l2_loss(reconstructed_img_batch_s,reconstructed_img_batch_s_pred)
gs=gridspec.GridSpec(8,8,width_ratios=[1,1,1,1,1,1,1,1],height_ratios=[1,1,1,1,1,1,1,1],wspace=0,hspace=0)
reconstructed_img=np.transpose(reconstructed_img_batch_s.cpu().data.numpy(),(0,2,3,1))
fig1=plt.figure(figsize=(8,8))
# fig1.suptitle('Image Reconstructions with encoder generated class-latent space')
for i in range(8):
for j in range(8):
if image_count<batch_size:
ax=plt.subplot(gs[i,j])
ax.axis('off')
ax.imshow(reconstructed_img[image_count])
image_count+=1
image_count=0
reconstructed_img=np.transpose(reconstructed_img_batch_s_pred.cpu().data.numpy(),(0,2,3,1))
fig2=plt.figure(figsize=(8,8))
# fig2.suptitle('Image Reconstructions with network generated class-latent space')
for i in range(8):
for j in range(8):
if image_count<batch_size:
ax=plt.subplot(gs[i,j])
ax.axis('off')
ax.imshow(reconstructed_img[image_count])
image_count+=1
print('Reconstruction error: '+str(reconstruction_err.data.storage().tolist()[0]))
# + [markdown] id="bofYoTDcJYH6" colab_type="text"
# # QUESTION 4 SPECIFIED PARTITION OF LATENT SPACE
# + id="6jkKYYqZJCEc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 235} outputId="0575105c-249d-4dae-92cd-5d631475dc05" executionInfo={"status": "ok", "timestamp": 1589633816473, "user_tz": -330, "elapsed": 24457, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "17863093696646821487"}}
traina=np.moveaxis(traina,0,-1)
traina=traina/255.0
traina=np.moveaxis(traina,-1,0)
print(traina.shape)
# train_Data = MNIST_Paired(traina,trainb,transform=transforms.Compose([ToTensor()]))
# train_loader = cycle(DataLoader(train_Data, batch_size=64, shuffle=False,drop_last=True))
train_df=TensorDataset(torch.from_numpy(traina).to(device),torch.from_numpy(trainb).to(device))
train_Data = MNIST_Paired(train_df)
batch_size=64
image_size=60
num_channels=3 #RGB
initial_learning_rate=0.0001
style_dim=512
class_dim=512
num_classes=672
reconstruction_coef=2.
reverse_cycle_coef=10.
kl_divergence_coef=3.
beta_1=0.9
beta_2=0.999
log_file='Outputs/Q2/logs.txt'
load_saved=False
# print(FLAGS)
epochs=10
sclassifier=Classifier(class_dim,num_classes).to(device)
sclassifier.apply(weights_init)
criterion=nn.BCELoss()
optimiser=torch.optim.Adam(sclassifier.parameters())
total_params=sum(p.numel() for p in sclassifier.parameters() if p.requires_grad)
print('total_params:'+str(total_params))
print(sclassifier)
datax=[]
datay=[]
train_loss_list=[]
train_acc_list=[]
val_loss_list=[]
val_acc_list=[]
encoder=Encoder(style_dim,class_dim).to(device)
encoder.load_state_dict(torch.load('Outputs/Q2/checkpoints/encoder_weights_new/encoder199.pt'))
idx_list=list(range(0,len(train_df)))
random.shuffle(idx_list)
x,y=train_df.tensors
x=x.cpu().numpy()
y=y.cpu().numpy()
for i in range(len(idx_list)):
datax.append(x[idx_list[i]])
datay.append(y[idx_list[i]])
datax=np.array(datax)
datay=np.array(datay)
x_val=datax[int(0.8*len(datax)):]
x_train=datax[0:int(0.8*len(datax))]
y_train=datay[0:int(0.8*len(datay))]
y_val=datay[int(0.8*len(datay)):]
train_d_s=TensorDataset(torch.from_numpy(x_train).to(device),torch.from_numpy(y_train).to(device))
val_d_s=TensorDataset(torch.from_numpy(x_val).to(device),torch.from_numpy(y_val).to(device))
train_loader=DataLoader(train_d_s,shuffle=False,batch_size=batch_size,drop_last=True)
val_loader=DataLoader(val_d_s,batch_size=batch_size,shuffle=False,drop_last=True)
# + id="MwkNmLn_MTsJ" colab_type="code" colab={}
# import tensorflow
x1=torch.FloatTensor(batch_size,num_channels,image_size,image_size).to(device)
criterion=nn.BCELoss()
for epoch in range(epochs):
train_iterator=iter(train_loader)
val_iterator=iter(val_loader)
epoch_loss=0
epoch_acc=0
sclassifier.train()
for i,bat in enumerate(train_iterator):
optimiser.zero_grad()
x=bat[0]
x1.copy_(x)
y=bat[1]
with torch.no_grad():
style_mu,style_logvar,class_latent=encoder(Variable(x1))
predicted=sclassifier(class_latent)
y=y.cpu().detach().numpy()
y=np.eye(num_classes)[y]
y=torch.from_numpy(y).float().cuda()
loss=criterion(predicted,y)
epoch_loss+=loss.item()
acc=accuracy(predicted,y)
epoch_acc+=acc
loss.backward()
optimiser.step()
# print(y.sum())
# print(loss.item())
# print(epoch_loss)
train_loss=epoch_loss/len(train_iterator)
train_acc=epoch_acc/len(train_iterator)
train_loss_list.append(train_loss)
train_acc_list.append(train_acc)
epoch_loss=0
epoch_acc=0
sclassifier.eval()
for i,bat in enumerate(val_iterator):
x=bat[0]
x1.copy_(x)
y=bat[1]
with torch.no_grad():
style_mu,style_logvar,class_latent=encoder(Variable(x1))
predicted=sclassifier(class_latent)
y=y.cpu().detach().numpy()
y=np.eye(num_classes)[y]
y=torch.from_numpy(y).float().cuda()
loss=criterion(predicted,y)
epoch_loss+=loss.item()
acc=accuracy(predicted,y)
epoch_acc+=acc
val_loss=epoch_loss/len(val_iterator)
val_acc=epoch_acc/len(val_iterator)
val_loss_list.append(val_loss)
val_acc_list.append(val_acc)
print('Epoch ',epoch+1,'/',epochs,' loss:',train_loss,' acc:',train_acc,' val_loss:',val_loss,' val_acc:',val_acc)
torch.save(sclassifier.state_dict(),os.getcwd()+'/Outputs/Q2/checkpoints/predictor/sclassifier.pt')
# + id="vGjE4rfzO5jV" colab_type="code" colab={}
plt.figure()
plt.title('acc vs epochs')
plt.plot(train_acc_list,label='train')
plt.plot(val_acc_list,label='validation')
plt.xlabel('epochs')
plt.ylabel('acc')
plt.legend(loc='upper left')
plt.figure()
plt.title('loss vs epochs')
plt.plot(train_loss_list,label='train')
plt.plot(val_loss_list,label='validation')
plt.xlabel('epochs')
plt.ylabel('loss')
plt.legend(loc='upper right')
# + id="fybrihnlSiZ1" colab_type="code" colab={}
sclassifier=Classifier(class_dim,num_classes).to(device)
sclassifier.load_state_dict(torch.load(os.getcwd()+'/Outputs/Q2/checkpoints/predictor/sclassifier.pt'))
sclassifier.eval()
test_acc=0
val_iterator=iter(val_loader)
for i,bat in enumerate(val_iterator):
x=bat[0]
x1.copy_(x)
y=bat[1]
with torch.no_grad():
style_mu,style_logvar,class_latent=encoder(Variable(x1))
pred=sclassifier(class_latent)
y=y.cpu().detach().numpy()
y=np.eye(num_classes)[y]
y=torch.from_numpy(y).float().cuda()
loss=criterion(pred,y)
acc=accuracy(pred,y)
test_acc+=acc
print('Test accuracy: '+str((test_acc/len(val_iterator))*100)+'%')
# + [markdown] id="IZg_kIFsU4dN" colab_type="text"
# #QUESTION 4 UNSPECIFIED PARTITION OF LATENT SPACE
# + id="MH7nQ_-NU9cq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 202} outputId="42a563ea-0aa7-4b9e-9f12-a069d36dea0a"
image_size=60
num_channels=3 #RGB
initial_learning_rate=0.0001
style_dim=512
class_dim=512
num_classes=672
reconstruction_coef=2.
reverse_cycle_coef=10.
kl_divergence_coef=3.
beta_1=0.9
beta_2=0.999
log_file='Outputs/Q2/logs.txt'
load_saved=False
# print(FLAGS)
epochs=10
zclassifier=Classifier(style_dim,num_classes).to(device)
zclassifier.apply(weights_init)
criterion=nn.BCELoss()
optimiser=torch.optim.Adam(zclassifier.parameters())
print(zclassifier)
datax=[]
datay=[]
train_loss_list=[]
train_acc_list=[]
val_loss_list=[]
val_acc_list=[]
encoder=Encoder(style_dim,class_dim).to(device)
encoder.load_state_dict(torch.load('Outputs/Q2/checkpoints/encoder_weights/encoder98.pt'))
idx_list=list(range(0,len(train_df)))
random.shuffle(idx_list)
x,y=train_df.tensors
x=x.cpu().numpy()
y=y.cpu().numpy()
for i in range(len(idx_list)):
datax.append(x[idx_list[i]])
datay.append(y[idx_list[i]])
datax=np.array(datax)
datay=np.array(datay)
x_val=datax[int(0.7*len(datax)):]
x_train=datax[0:int(0.7*len(datax))]
y_train=datay[0:int(0.7*len(datay))]
y_val=datay[int(0.7*len(datay)):]
train_d_s=TensorDataset(torch.from_numpy(x_train).to(device),torch.from_numpy(y_train).to(device))
val_d_s=TensorDataset(torch.from_numpy(x_val).to(device),torch.from_numpy(y_val).to(device))
train_loader=DataLoader(train_d_s,shuffle=False,batch_size=batch_size,drop_last=True)
val_loader=DataLoader(val_d_s,batch_size=batch_size,shuffle=False,drop_last=True)
# + id="fvqSOW0OVk9S" colab_type="code" colab={}
x1=torch.FloatTensor(batch_size,num_channels,image_size,image_size).to(device)
for epoch in range(epochs):
train_iterator=iter(train_loader)
val_iterator=iter(val_loader)
epoch_loss=0
epoch_acc=0
zclassifier.train()
for i,bat in enumerate(train_iterator):
x=bat[0]
x1.copy_(x)
y=bat[1]
optimiser.zero_grad()
with torch.no_grad():
style_mu,style_logvar,class_latent=encoder(Variable(x1))
z_latent_space=reparameterize(training=True,mu=style_mu,logvar=style_logvar)
predicted=zclassifier(z_latent_space)
y=y.cpu().detach().numpy()
y=np.eye(num_classes)[y]
y=torch.from_numpy(y).float().cuda()
loss=criterion(predicted,y)
loss.backward()
optimiser.step()
epoch_loss+=loss.item()
# print(epoch_loss)
acc=accuracy(predicted,y)
epoch_acc+=acc
train_loss=epoch_loss/len(train_iterator)
train_acc=epoch_acc/len(train_iterator)
train_loss_list.append(train_loss)
train_acc_list.append(train_acc)
epoch_loss=0
epoch_acc=0
zclassifier.eval()
for i,bat in enumerate(val_iterator):
x=bat[0]
x1.copy_(x)
y=bat[1]
with torch.no_grad():
style_mu,style_logvar,class_latent=encoder(Variable(x1))
z_latent_space=reparameterize(training=True,mu=style_mu,logvar=style_logvar)
predicted=zclassifier(z_latent_space)
y=y.cpu().detach().numpy()
y=np.eye(num_classes)[y]
y=torch.from_numpy(y).float().cuda()
loss=criterion(predicted,y)
epoch_loss+=loss.item()
acc=accuracy(predicted,y)
epoch_acc+=acc
val_loss=epoch_loss/len(val_iterator)
val_acc=epoch_acc/len(val_iterator)
val_loss_list.append(val_loss)
val_acc_list.append(val_acc)
print('Epoch ',epoch+1,'/',epochs,' loss:',train_loss,' acc:',train_acc,' val_loss:',val_loss,' val_acc:',val_acc)
torch.save(zclassifier.state_dict(),os.getcwd()+'/Outputs/Q2/checkpoints/predictor/zclassifier.pt')
# + id="ZC_StGREW97G" colab_type="code" colab={}
plt.figure()
plt.title('acc vs epochs')
plt.plot(train_acc_list,label='train')
plt.plot(val_acc_list,label='validation')
plt.xlabel('epochs')
plt.ylabel('acc')
plt.legend(loc='upper left')
plt.figure()
plt.title('loss vs epochs')
plt.plot(train_loss_list,label='train')
plt.plot(val_loss_list,label='validation')
plt.xlabel('epochs')
plt.ylabel('loss')
plt.legend(loc='upper right')
# + id="rSIgsypsWw0d" colab_type="code" colab={}
zclassifier=Classifier(class_dim,num_classes).to(device)
zclassifier.load_state_dict(torch.load(os.getcwd()+'/Outputs/Q2/checkpoints/predictor/zclassifier.pt'))
zclassifier.eval()
test_acc=0
val_iterator=iter(val_loader)
for i,bat in enumerate(val_iterator):
x=bat[0]
x1.copy_(x)
y=bat[1]
with torch.no_grad():
style_mu,style_logvar,class_latent=encoder(Variable(x1))
pred=zclassifier(style_mu)
y=y.cpu().detach().numpy()
y=np.eye(num_classes)[y]
y=torch.from_numpy(y).float().cuda()
loss=criterion(pred,y)
acc=accuracy(pred,y)
test_acc+=acc
print('Test accuracy: '+str((test_acc/len(val_iterator))*100)+'%')
# + [markdown] id="e7xV-3SsX2_S" colab_type="text"
# #QUESTION 2
#
# + id="8GXOFXNCXORd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 983} outputId="9c0ccd3b-34c3-448c-e683-41bde83d6424" executionInfo={"status": "ok", "timestamp": 1589628034607, "user_tz": -330, "elapsed": 5666, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "17863093696646821487"}}
col=[]
row=[]
x1=torch.FloatTensor(1,num_channels,image_size,image_size).to(device)
x2=torch.FloatTensor(1,num_channels,image_size,image_size).to(device)
## model style grid transfer
for i in range(8):
image,_=test_df[random.randint(0,len(test_df)-1)]
col.append(image)
row.append(image)
encoder=Encoder(style_dim,class_dim).to(device)
decoder=Decoder(style_dim,class_dim).to(device)
encoder.load_state_dict(torch.load('Outputs/Q2/checkpoints/encoder_weights_new/encoder199.pt'))
decoder.load_state_dict(torch.load('Outputs/Q2/checkpoints/decoder_weights_new/decoder199.pt'))
## complete grid
gs1=gridspec.GridSpec(8,8,width_ratios=[1,1,1,1,1,1,1,1],height_ratios=[1,1,1,1,1,1,1,1],wspace=0,hspace=0)
fig1=plt.figure(figsize=(8,8))
for i in range(len(row)):
x1.copy_(row[i])
style_mu,style_logvar,class_latent=encoder(Variable(x1))
for j in range(len(col)):
x2.copy_(col[j])
style_mu2,style_logvar2,class_latent2=encoder(Variable(x2))
reconstructed_img=decoder(style_mu,class_latent2)
reconstructed_img=reconstructed_img.squeeze(0)
reconstructed_img=np.transpose(reconstructed_img.cpu().data.numpy(),(1,2,0))
ax=plt.subplot(gs1[i,j])
ax.axis('off')
ax.imshow(reconstructed_img)
## row print
gs2=gridspec.GridSpec(1,8,width_ratios=[1,1,1,1,1,1,1,1],wspace=0,hspace=0)
fig2=plt.figure(figsize=(8,1))
for i in range(8):
image=row[i]
image=image.squeeze(0)
image=np.transpose(image.cpu().data.numpy(),(1,2,0))
image=image.astype('float')
ax=plt.subplot(gs2[i])
ax.axis('off')
ax.imshow(image)
##column
fig3=plt.figure(figsize=(1,8))
gs3=gridspec.GridSpec(8,1,height_ratios=[1,1,1,1,1,1,1,1],wspace=0,hspace=0)
for i in range(8):
image=col[i]
image=image.squeeze(0)
image=np.transpose(image.cpu().data.numpy(),(1,2,0))
image=image.astype('float')
ax=plt.subplot(gs3[i])
ax.axis('off')
ax.imshow(image)
# + [markdown] id="kAhpezmHg4xR" colab_type="text"
# #QUESTION 3
#
# + id="Ms1YQPiJg67B" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 466} outputId="e2096cd0-4149-4b80-9dd7-084d1b7733e8" executionInfo={"status": "ok", "timestamp": 1589628255008, "user_tz": -330, "elapsed": 7315, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "17863093696646821487"}}
encoder=Encoder(style_dim,class_dim).to(device)
decoder=Decoder(style_dim,class_dim).to(device)
encoder.load_state_dict(torch.load('Outputs/Q2/checkpoints/encoder_weights_new/encoder199.pt'))
decoder.load_state_dict(torch.load('Outputs/Q2/checkpoints/decoder_weights_new/decoder199.pt'))
## complete grid
gs1=gridspec.GridSpec(10,10,width_ratios=[1,1,1,1,1,1,1,1,1,1],height_ratios=[1,1,1,1,1,1,1,1,1,1],wspace=0,hspace=0)
fig1=plt.figure(figsize=(8,8))
x1=torch.FloatTensor(1,num_channels,image_size,image_size).to(device)
x2=torch.FloatTensor(1,num_channels,image_size,image_size).to(device)
image1,_=test_df[random.randint(0,len(test_df)-1)]
image2,_=test_df[random.randint(0,len(test_df)-1)]
x1.copy_(image1)
x2.copy_(image2)
style_mu1,style_logvar1,class_latent1=encoder(Variable(x1))
style_mu2,style_logvar2,class_latent2=encoder(Variable(x2))
diff_style=style_mu2-style_mu1
diff_class=class_latent2-class_latent1
n=10
inter_style=torch.zeros((n,1,diff_style.shape[1])).to(device)
inter_class=torch.zeros((n,1,diff_class.shape[1])).to(device)
for i in range(n):
inter_style[i]=style_mu1+(i/(n-1))*diff_style
inter_class[i]=class_latent1+(i/(n-1))*diff_class
for i in range(10):
for j in range(10):
reconstructed_img=decoder(inter_style[i],inter_class[j])
reconstructed_img=reconstructed_img.squeeze(0)
reconstructed_img=np.transpose(reconstructed_img.cpu().data.numpy(),(1,2,0))
ax=plt.subplot(gs1[i,j])
ax.axis('off')
ax.imshow(reconstructed_img)
plt.savefig('q1_inter.png',dpi=300)
# + id="3ytZTU0FuNkD" colab_type="code" colab={}
encoder=Encoder(style_dim,class_dim).to(device)
decoder=Decoder(style_dim,class_dim).to(device)
encoder.load_state_dict(torch.load('Outputs/Q2/checkpoints/encoder_weights_new/encoder199.pt'))
decoder.load_state_dict(torch.load('Outputs/Q2/checkpoints/decoder_weights_new/decoder199.pt'))
## complete grid
gs1=gridspec.GridSpec(10,10,width_ratios=[1,1,1,1,1,1,1,1,1,1],height_ratios=[1,1,1,1,1,1,1,1,1,1],wspace=0,hspace=0)
fig1=plt.figure(figsize=(8,8))
x1=torch.FloatTensor(1,num_channels,image_size,image_size).to(device)
x2=torch.FloatTensor(1,num_channels,image_size,image_size).to(device)
image1,_=test_df[random.randint(0,len(test_df)-1)]
image2,_=test_df[random.randint(0,len(test_df)-1)]
x1.copy_(image1)
x2.copy_(image2)
style_mu1,style_logvar1,class_latent1=encoder(Variable(x1))
style_mu2,style_logvar2,class_latent2=encoder(Variable(x2))
diff_style=style_mu2-style_mu1
diff_class=class_latent2-class_latent1
n=10
inter_style=torch.zeros((n,1,diff_style.shape[1])).to(device)
inter_class=torch.zeros((n,1,diff_class.shape[1])).to(device)
for i in range(n):
inter_style[i]=style_mu1+(i/(n-1))*diff_style
inter_class[i]=class_latent1+(i/(n-1))*diff_class
for i in range(10):
for j in range(10):
reconstructed_img=decoder(inter_style[i],inter_class[j])
reconstructed_img=reconstructed_img.squeeze(0)
reconstructed_img=np.transpose(reconstructed_img.cpu().data.numpy(),(1,2,0))
ax=plt.subplot(gs1[i,j])
ax.axis('off')
ax.imshow(reconstructed_img)
plt.savefig('q2_inter.png',dpi=300)
# + id="V4taVXRWuOc-" colab_type="code" colab={}
encoder=Encoder(style_dim,class_dim).to(device)
decoder=Decoder(style_dim,class_dim).to(device)
encoder.load_state_dict(torch.load('Outputs/Q2/checkpoints/encoder_weights_new/encoder199.pt'))
decoder.load_state_dict(torch.load('Outputs/Q2/checkpoints/decoder_weights_new/decoder199.pt'))
## complete grid
gs1=gridspec.GridSpec(10,10,width_ratios=[1,1,1,1,1,1,1,1,1,1],height_ratios=[1,1,1,1,1,1,1,1,1,1],wspace=0,hspace=0)
fig1=plt.figure(figsize=(8,8))
x1=torch.FloatTensor(1,num_channels,image_size,image_size).to(device)
x2=torch.FloatTensor(1,num_channels,image_size,image_size).to(device)
image1,_=test_df[random.randint(0,len(test_df)-1)]
image2,_=test_df[random.randint(0,len(test_df)-1)]
x1.copy_(image1)
x2.copy_(image2)
style_mu1,style_logvar1,class_latent1=encoder(Variable(x1))
style_mu2,style_logvar2,class_latent2=encoder(Variable(x2))
diff_style=style_mu2-style_mu1
diff_class=class_latent2-class_latent1
n=10
inter_style=torch.zeros((n,1,diff_style.shape[1])).to(device)
inter_class=torch.zeros((n,1,diff_class.shape[1])).to(device)
for i in range(n):
inter_style[i]=style_mu1+(i/(n-1))*diff_style
inter_class[i]=class_latent1+(i/(n-1))*diff_class
for i in range(10):
for j in range(10):
reconstructed_img=decoder(inter_style[i],inter_class[j])
reconstructed_img=reconstructed_img.squeeze(0)
reconstructed_img=np.transpose(reconstructed_img.cpu().data.numpy(),(1,2,0))
ax=plt.subplot(gs1[i,j])
ax.axis('off')
ax.imshow(reconstructed_img)
plt.savefig('q3_inter.png',dpi=300)
| main.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .cpp
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: C++17
// language: C++17
// name: xcpp17
// ---
// # C++ Dev Environment
//
// - the following tools are recommended for this course
// 1. Visual Studio (VS) Code editor
// - light weight cross-platform editor for many programming languages; has rich extensions
// 2. git client for version control
// - Note: VS Code provides GUI-based git
// 3. g++ compiler
//
// - follow the instructions from https://github.com/rambasnet/DevEnvSetup to setup Jupyter Notebook on various platforms
// ## Using g++ compiler on Windows WSL, Mac and Linux
// - the steps provided here assumes that you're using the recommended C++ dev environment above
// - open a Terminal program
// - be familiar with the terminal and some [basic bash commands](https://sites.tufts.edu/cbi/files/2013/01/linux_cheat_sheet.pdf)
// - change current working directory to where the right folder where the .cpp file is
// - use `ls` commad to see all the contents of the directory
// - use `cd <dir_name>` command to change directory to the given dir_name
// - make sure the current working directory is where your `.cpp` file is
// - use `pwd` command on a `*nix` terminal to know the current working directory
// - compile using g++
// - run the executable
// - the following sequence of commands are worth remembering
// - can use these commands on repl.it cloud-based IDE as well
//
// ```bash
// $ cd projectFolder # change working directory to the project folder
// $ pwd # print current working directory
// $ ls # list contents of current directory
// $ g++ -std=c++17 -o outputProgram inputFile.cpp # compile inputFile.cpp to outputPrgram
// $ ./outputProgram # run output program
// ```
//
// ## Using Make program
// - a great way to compile, build, run, test and deploy C/C++ program
// - create a file named `Makefile` inside the project folder
// - see a quick tutorial on Makefile [https://makefiletutorial.com/](https://makefiletutorial.com/)
// - see [makefile_demos](makefile_demos) for various Makefile examples
// - use Makefile template provided in [makefile_demos/Makefile_template](./makefile_demos/Makefile_template)
// - run the following commands from inside the project folder on a Terminal
//
// ```bash
// $ cd projectFolder # change current working director - folder with c++ file(s)
// $ make # build program
// $ ls # see the name of your executable in the current directory
// $ ./programName # run the program by it's name
// $ make clean # run clean rule; usually deletes all object/exe files
// ```
| DevEnvironmentSetup.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### 2. Criando um Projeto
# **2.1. Comando para criar um projeto (django-admin)**
# <br/>
# ```
# $ django-admin startproject siteblog
# ```
# **2.1.1. Manipulando informações sensíveis (arquivo .env)**
# <br/>
# Fonte: https://ecossistemagis.com/geodjango-variaveis-de-ambiente-em-arquivo-env/
# <br/>
# - Com o Ambiente Virtual ativado, instale o pacote "dotenv".
# > `
# $ pip install python-dotenv
# `
# - Na mesma pasta em que está seu arquivo "settings.py", crie um novo arquivo chamado ".env"
# - Crie o arquivo .env e acrescente o seguinte:
# > ```
# DJANGO_SECRET_KEY='<put here your secret key>'
# DB_USER=myuser
# DB_PASSWORD=<PASSWORD>
# DB_HOST=host
# DB_PORT=port
# ```
#
# - No arquivo "settings.py", acrescente o seguinte código (importando o pacote):
# > ```
# import os
# from pathlib import Path
# from dotenv import load_dotenv
# load_dotenv() # permite acessar as variáveis definidas em .env
# ```
#
# - E um pouco abaixo, ainda no arquivo "settings.py"... (carregando a variável):
# > `
# SECRET_KEY = os.environ.get('DJANGO_SECRET_KEY')
# `
#
# **2.2. Criando uma base de dados local (SQLite)**
# - Execute o comando:
#
# >`
# $ python manage.py migrate
# `
#
# <br/>
# Desse modo, as tabelas padrões do Django serão criadas.
# <br/>
# <br/>
#
# **IMPORTANTE:**
#
# Às vezes, ocorre um erro na configuração do banco de dados SQLite:
# > ```
# DATABASES = {
# 'default': {
# 'ENGINE': 'django.db.backends.sqlite3',
# # 'NAME': BASE_DIR / 'db.sqlite3',# <===ANTES
# 'NAME': str(os.path.join(BASE_DIR, "db.sqlite3")) # <=== ALTERAÇÃO PARA CORRIGIR
# }
# }
# ```
# **2.3. Gere e guarde as dependências usadas até o momento:**
# >`pip freeze > requirements.txt`
# **2.4. Execute o Servidor e veja a sua aplicação funcionando:**
# >`$ python manage.py runserver`
#
# **OBS:** *É possível parametrizar esse comando:*
# >`$ python manage.py runserver 127.0.0.1:8001`
#
# **2.5. Configurações Adicionais importantes e necessárias: (arquivo: settings.py)**
# > ```
# DEBUG = True (Para ambientes de produção deve ser False. Pode ser criado um arquivo dev.py e outro prd.py)
# ALLOWED_HOSTS = ['127.0.0.1', '.pythonanywhere.com']
# TIME_ZONE = 'America/Sao_Paulo'
# LANGUAGE_CODE = 'pt-BR'
# STATIC_URL = '/static/'
# STATIC_ROOT = os.path.join(BASE_DIR, 'static')
# ```
| notebooks/2.Criando-Projeto.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] _cell_guid="dafef955-4c2c-a871-f1d8-3e0d306393b0" papermill={"duration": 0.029137, "end_time": "2021-11-25T08:05:50.561192", "exception": false, "start_time": "2021-11-25T08:05:50.532055", "status": "completed"} tags=[]
# # Using the Wisconsin breast cancer diagnostic data set for predictive analysis
# Attribute Information:
#
# - 1) ID number
# - 2) Diagnosis (M = malignant, B = benign)
#
# -3-32.Ten real-valued features are computed for each cell nucleus:
#
# - a) radius (mean of distances from center to points on the perimeter)
# - b) texture (standard deviation of gray-scale values)
# - c) perimeter
# - d) area
# - e) smoothness (local variation in radius lengths)
# - f) compactness (perimeter^2 / area - 1.0)
# - g). concavity (severity of concave portions of the contour)
# - h). concave points (number of concave portions of the contour)
# - i). symmetry
# - j). fractal dimension ("coastline approximation" - 1)
#
# The mean, standard error and "worst" or largest (mean of the three largest values) of these features were computed for each image, resulting in 30 features. For instance, field 3 is Mean Radius, field 13 is Radius SE, field 23 is Worst Radius.
#
#
# For this analysis, as a guide to predictive analysis I followed the instructions and discussion on "A Complete Tutorial on Tree Based Modeling from Scratch (in R & Python)" at Analytics Vidhya.
# + [markdown] _cell_guid="5e26372e-f1bd-b50f-0c1c-33a44306d1f7" papermill={"duration": 0.025366, "end_time": "2021-11-25T08:05:50.612294", "exception": false, "start_time": "2021-11-25T08:05:50.586928", "status": "completed"} tags=[]
# #Load Libraries
# + _cell_guid="2768ce80-1a7d-ca31-a35f-29cf0ef7fb15" papermill={"duration": 1.041675, "end_time": "2021-11-25T08:05:51.679431", "exception": false, "start_time": "2021-11-25T08:05:50.637756", "status": "completed"} tags=[]
import numpy as np
import pandas as pd
# %matplotlib inline
import matplotlib.pyplot as plt
import matplotlib.gridspec as gridspec
import mpld3 as mpl
from sklearn.model_selection import train_test_split
from sklearn.model_selection import KFold
from sklearn import metrics
# + [markdown] _cell_guid="09b9d090-2cba-ad5a-58ce-84208f95dba4" papermill={"duration": 0.025544, "end_time": "2021-11-25T08:05:51.730413", "exception": false, "start_time": "2021-11-25T08:05:51.704869", "status": "completed"} tags=[]
# # Load the data
# + _cell_guid="9180cb22-53d2-6bf2-3a29-99448ab808fb" papermill={"duration": 0.107613, "end_time": "2021-11-25T08:05:51.863755", "exception": false, "start_time": "2021-11-25T08:05:51.756142", "status": "completed"} tags=[]
df = pd.read_csv("../input/data.csv",header = 0)
df.head()
# + [markdown] _cell_guid="e382010d-1d71-b8d6-4a6e-a0abc9e42372" papermill={"duration": 0.026219, "end_time": "2021-11-25T08:05:51.916566", "exception": false, "start_time": "2021-11-25T08:05:51.890347", "status": "completed"} tags=[]
# # Clean and prepare data
# + _cell_guid="f9fd3701-af9d-8d8c-5d0e-e2673d7977fe" papermill={"duration": 0.042167, "end_time": "2021-11-25T08:05:51.984588", "exception": false, "start_time": "2021-11-25T08:05:51.942421", "status": "completed"} tags=[]
df.drop('id',axis=1,inplace=True)
df.drop('Unnamed: 32',axis=1,inplace=True)
len(df)
# + _cell_guid="083fe464-8dac-713e-d0a1-46435c0d93fa" papermill={"duration": 0.037072, "end_time": "2021-11-25T08:05:52.049374", "exception": false, "start_time": "2021-11-25T08:05:52.012302", "status": "completed"} tags=[]
df.diagnosis.unique()
# + _cell_guid="0882e4c2-3d4d-d4d9-5f49-f36c1b248b93" papermill={"duration": 0.058435, "end_time": "2021-11-25T08:05:52.134664", "exception": false, "start_time": "2021-11-25T08:05:52.076229", "status": "completed"} tags=[]
#Convert
df['diagnosis'] = df['diagnosis'].map({'M':1,'B':0})
df.head()
# Explore data
# + _cell_guid="cfd882cd-1719-4093-934a-539faf665353" papermill={"duration": 0.116726, "end_time": "2021-11-25T08:05:52.279608", "exception": false, "start_time": "2021-11-25T08:05:52.162882", "status": "completed"} tags=[]
df.describe()
# + _cell_guid="aa80be8a-4022-038b-d7b7-0789df4ef973" papermill={"duration": 0.445746, "end_time": "2021-11-25T08:05:52.753778", "exception": false, "start_time": "2021-11-25T08:05:52.308032", "status": "completed"} tags=[]
df.describe()
plt.hist(df['diagnosis'])
plt.title('Diagnosis (M=1 , B=0)')
plt.show()
# + [markdown] _cell_guid="56b72979-5155-2a99-1b6e-a55cbf72d2a3" papermill={"duration": 0.028712, "end_time": "2021-11-25T08:05:52.812326", "exception": false, "start_time": "2021-11-25T08:05:52.783614", "status": "completed"} tags=[]
# ### nucleus features vs diagnosis
# + _cell_guid="bc36c937-c5d8-8635-480b-777a94571310" papermill={"duration": 0.040922, "end_time": "2021-11-25T08:05:52.883830", "exception": false, "start_time": "2021-11-25T08:05:52.842908", "status": "completed"} tags=[]
features_mean=list(df.columns[1:11])
# split dataframe into two based on diagnosis
dfM=df[df['diagnosis'] ==1]
dfB=df[df['diagnosis'] ==0]
# + _cell_guid="3f3b5e1b-605d-51b4-28c7-c551b5d13a48" papermill={"duration": 3.920665, "end_time": "2021-11-25T08:05:56.833955", "exception": false, "start_time": "2021-11-25T08:05:52.913290", "status": "completed"} tags=[]
plt.rcParams.update({'font.size': 8})
fig, axes = plt.subplots(nrows=5, ncols=2, figsize=(8,10))
axes = axes.ravel()
for idx,ax in enumerate(axes):
ax.figure
binwidth= (max(df[features_mean[idx]]) - min(df[features_mean[idx]]))/50
ax.hist([dfM[features_mean[idx]],dfB[features_mean[idx]]], alpha=0.5,stacked=True, label=['M','B'],color=['r','g'],bins=np.arange(min(df[features_mean[idx]]), max(df[features_mean[idx]]) + binwidth, binwidth) , density = True,)
ax.legend(loc='upper right')
ax.set_title(features_mean[idx])
plt.tight_layout()
plt.show()
# + [markdown] _cell_guid="4b8d6133-427b-1ecf-0e24-9ec2afea0a0e" papermill={"duration": 0.030816, "end_time": "2021-11-25T08:05:56.895903", "exception": false, "start_time": "2021-11-25T08:05:56.865087", "status": "completed"} tags=[]
# ### Observations
#
# 1. mean values of cell radius, perimeter, area, compactness, concavity and concave points can be used in classification of the cancer. Larger values of these parameters tends to show a correlation with malignant tumors.
# 2. mean values of texture, smoothness, symmetry or fractual dimension does not show a particular preference of one diagnosis over the other. In any of the histograms there are no noticeable large outliers that warrants further cleanup.
# + [markdown] _cell_guid="ac11039f-0418-3553-9412-ae3d50bef4e4" papermill={"duration": 0.030468, "end_time": "2021-11-25T08:05:56.957557", "exception": false, "start_time": "2021-11-25T08:05:56.927089", "status": "completed"} tags=[]
# ## Creating a test set and a training set
# Since this data set is not ordered, I am going to do a simple 70:30 split to create a training data set and a test data set.
# + _cell_guid="1390898b-a338-7395-635f-6e1c216861e6" papermill={"duration": 0.040684, "end_time": "2021-11-25T08:05:57.028656", "exception": false, "start_time": "2021-11-25T08:05:56.987972", "status": "completed"} tags=[]
traindf, testdf = train_test_split(df, test_size = 0.3)
# + [markdown] _cell_guid="45dac047-52fb-b847-a521-fa6883ebd5f6" papermill={"duration": 0.029978, "end_time": "2021-11-25T08:05:57.089625", "exception": false, "start_time": "2021-11-25T08:05:57.059647", "status": "completed"} tags=[]
# ## Model Building
#
# Here we are going to build a classification model and evaluate its performance using the training set.
#
#
# + [markdown] papermill={"duration": 0.029945, "end_time": "2021-11-25T08:05:57.150680", "exception": false, "start_time": "2021-11-25T08:05:57.120735", "status": "completed"} tags=[]
# # Naive Bayes model
# + papermill={"duration": 0.04056, "end_time": "2021-11-25T08:05:57.221667", "exception": false, "start_time": "2021-11-25T08:05:57.181107", "status": "completed"} tags=[]
from sklearn.naive_bayes import GaussianNB
model=GaussianNB()
predictor_var = ['radius_mean','perimeter_mean','area_mean','compactness_mean','concave points_mean']
outcome_var='diagnosis'
# + papermill={"duration": 0.048417, "end_time": "2021-11-25T08:05:57.301871", "exception": false, "start_time": "2021-11-25T08:05:57.253454", "status": "completed"} tags=[]
model.fit(traindf[predictor_var],traindf[outcome_var])
predictions = model.predict(traindf[predictor_var])
accuracy = metrics.accuracy_score(predictions,traindf[outcome_var])
print("Accuracy : %s" % "{0:.3%}".format(accuracy))
# + papermill={"duration": 0.422861, "end_time": "2021-11-25T08:05:57.757672", "exception": false, "start_time": "2021-11-25T08:05:57.334811", "status": "completed"} tags=[]
import seaborn as sns
sns.heatmap(metrics.confusion_matrix(predictions,traindf[outcome_var]),annot=True)
# + papermill={"duration": 0.073701, "end_time": "2021-11-25T08:05:57.864024", "exception": false, "start_time": "2021-11-25T08:05:57.790323", "status": "completed"} tags=[]
from sklearn.model_selection import cross_val_score
from statistics import mean
print(mean(cross_val_score(model, traindf[predictor_var],traindf[outcome_var], cv=5))*100)
# + [markdown] papermill={"duration": 0.032521, "end_time": "2021-11-25T08:05:57.928943", "exception": false, "start_time": "2021-11-25T08:05:57.896422", "status": "completed"} tags=[]
# # KNN Model
# + papermill={"duration": 0.186638, "end_time": "2021-11-25T08:05:58.148025", "exception": false, "start_time": "2021-11-25T08:05:57.961387", "status": "completed"} tags=[]
from sklearn.neighbors import KNeighborsClassifier
model=KNeighborsClassifier(n_neighbors=4)
predictor_var = ['radius_mean','perimeter_mean','area_mean','compactness_mean','concave points_mean']
outcome_var='diagnosis'
# + papermill={"duration": 0.067455, "end_time": "2021-11-25T08:05:58.247898", "exception": false, "start_time": "2021-11-25T08:05:58.180443", "status": "completed"} tags=[]
model.fit(traindf[predictor_var],traindf[outcome_var])
predictions = model.predict(traindf[predictor_var])
accuracy = metrics.accuracy_score(predictions,traindf[outcome_var])
print("Accuracy : %s" % "{0:.3%}".format(accuracy))
# + papermill={"duration": 0.086077, "end_time": "2021-11-25T08:05:58.369076", "exception": false, "start_time": "2021-11-25T08:05:58.282999", "status": "completed"} tags=[]
from sklearn.model_selection import cross_val_score
from statistics import mean
print(mean(cross_val_score(model, traindf[predictor_var],traindf[outcome_var], cv=5))*100)
# + papermill={"duration": 1.498948, "end_time": "2021-11-25T08:05:59.901562", "exception": false, "start_time": "2021-11-25T08:05:58.402614", "status": "completed"} tags=[]
import numpy as np
x_train=traindf[predictor_var]
y_train=traindf[outcome_var]
x_test=testdf[predictor_var]
y_test=testdf[outcome_var]
trainAccuracy=[]
testAccuracy=[]
errorRate=[]
for k in range(1,40):
model=KNeighborsClassifier(n_neighbors=k)
model.fit(x_train,y_train)
pred_i = model.predict(x_test)
errorRate.append(np.mean(pred_i != y_test))
trainAccuracy.append(model.score(x_train,y_train))
testAccuracy.append(model.score(x_test,y_test))
# + papermill={"duration": 0.292232, "end_time": "2021-11-25T08:06:00.226818", "exception": false, "start_time": "2021-11-25T08:05:59.934586", "status": "completed"} tags=[]
plt.figure(figsize=(10,6))
plt.plot(range(1,40),errorRate,color='blue', linestyle='dashed',
marker='o',markerfacecolor='red', markersize=10)
plt.title('Error Rate vs. K Value')
plt.xlabel('K')
plt.ylabel('Error Rate')
print("Minimum error:-",min(errorRate),"at K =",errorRate.index(min(errorRate))+1)
# + papermill={"duration": 0.302149, "end_time": "2021-11-25T08:06:00.562904", "exception": false, "start_time": "2021-11-25T08:06:00.260755", "status": "completed"} tags=[]
from matplotlib import pyplot as plt,style
plt.figure(figsize=(12,6))
plt.plot(range(1,40),trainAccuracy,label="Train Score",marker="o",markerfacecolor="teal",color="blue",linestyle="dashed")
plt.plot(range(1,40),testAccuracy,label="Test Score",marker="o",markerfacecolor="red",color="black",linestyle="dashed")
plt.legend()
plt.xlabel("Number of Neighbors")
plt.ylabel("Score")
plt.title("Nbd Vs Score")
plt.show()
# + [markdown] papermill={"duration": 0.035819, "end_time": "2021-11-25T08:06:00.634531", "exception": false, "start_time": "2021-11-25T08:06:00.598712", "status": "completed"} tags=[]
# Testing with new K Value= 30
#
# + papermill={"duration": 0.043365, "end_time": "2021-11-25T08:06:00.712988", "exception": false, "start_time": "2021-11-25T08:06:00.669623", "status": "completed"} tags=[]
from sklearn.neighbors import KNeighborsClassifier
model=KNeighborsClassifier(n_neighbors=31)
predictor_var = ['radius_mean','perimeter_mean','area_mean','compactness_mean','concave points_mean']
outcome_var='diagnosis'
# + papermill={"duration": 0.066677, "end_time": "2021-11-25T08:06:00.815531", "exception": false, "start_time": "2021-11-25T08:06:00.748854", "status": "completed"} tags=[]
model.fit(traindf[predictor_var],traindf[outcome_var])
predictions = model.predict(traindf[predictor_var])
accuracy = metrics.accuracy_score(predictions,traindf[outcome_var])
print("Accuracy : %s" % "{0:.3%}".format(accuracy))
# + papermill={"duration": 0.110472, "end_time": "2021-11-25T08:06:00.972381", "exception": false, "start_time": "2021-11-25T08:06:00.861909", "status": "completed"} tags=[]
from sklearn.model_selection import cross_val_score
from statistics import mean
print(mean(cross_val_score(model, traindf[predictor_var],traindf[outcome_var], cv=5))*100)
# + [markdown] papermill={"duration": 0.036383, "end_time": "2021-11-25T08:06:01.046268", "exception": false, "start_time": "2021-11-25T08:06:01.009885", "status": "completed"} tags=[]
# We can see after using K values of 31 our cross-validation and accuracy scores are very close to each other compared to our initial assumption of 4 where cross validation scores were lower and accuracy scores were lot higher on train dataset
# + papermill={"duration": 0.036314, "end_time": "2021-11-25T08:06:01.119614", "exception": false, "start_time": "2021-11-25T08:06:01.083300", "status": "completed"} tags=[]
| breast-cancer-prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# #!/usr/bin/python3
# coding: utf-8
# MHLW
# -
from datetime import datetime as dt
from datetime import timedelta as td
import matplotlib.pyplot as plt
import numpy as np
import os
import pandas as pd
import sys
import plotly
import plotly.express as px
import plotly.tools as tls
import plotly.graph_objects as go
import plotly.io as pio
import plotly.offline as offline
from plotly.subplots import make_subplots
if "ipy" in sys.argv[0]:
offline.init_notebook_mode()
from cov19utils import create_basic_plot_figure, \
show_and_clear, moving_average, \
blank2zero, csv2array, \
get_twitter, tweet_with_image, \
get_gpr_predict, FONT_NAME, DT_OFFSET, \
download_if_needed, show_and_save_plotly, \
make_exp_fit_graph
from scipy.optimize import curve_fit
from scipy.special import factorial
from fftdenoise import fft_denoise
if dt.now().weekday() != 5:
print("Today is not Saturday.")
if not "ipy" in sys.argv[0]:
sys.exit()
if dt.now().hour < 18:
print("before 6 pm.")
if not "ipy" in sys.argv[0]:
sys.exit()
today_str = dt.now().isoformat()[:16].replace('T', ' ')
# +
# 厚労省の OpenData を参照する
base_uri = "https://www.mhlw.go.jp/content/"
raws = dict(
posis = "pcr_positive_daily.csv",
# 日別PCR検査人数よりも検査機関別の数値を使用すべき
tests = "pcr_tested_daily.csv",
cases = "cases_total.csv",
recov = "recovery_total.csv",
death = "death_total.csv",
pcr = "pcr_case_daily.csv")
offsets = dict(
dates = 0, # 日付オフセット
cases = 1, # 入院治療を要する者(Total)
death = 2, # 死亡者数(Total)
pcr = 3, # PCR検査件数 3:感染研 4:検疫 5:保健所 6:民間 7:大学 8:医療機関 9:民間自費
pcrs = 10, # 上記の合算
posis = 11, # 陽性者数(Daily)
tests = 12, # PCR検査 人数(Daily)
recov = 13, # 退院(Total)
ratio = 14, # 陽性率(Daily) = 陽性者数 / 検査人数
total = 15, # 陽性者数(Total)
) #
# 集計期間
dt_range = (dt.today() - dt.strptime(DT_OFFSET, "%Y/%m/%d")).days
# 配列初期化
all_data_arr = []
for i in np.arange(dt_range):
all_data_arr.append([i, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0])
# データの取得
for k, v in raws.items():
download_if_needed(base_uri, v)
# データの集計
for k, v in raws.items():
if v != 0:
csv2array(all_data_arr, k, v, offsets[k])
# 陽性率等の計算
for i in np.arange(dt_range):
div = all_data_arr[i][offsets['pcrs']]
if div != 0:
all_data_arr[i][offsets['ratio']] = max(0, min(100, (all_data_arr[i][offsets['posis']] / div) * 100))
if i == 0:
all_data_arr[i][offsets['total']] = all_data_arr[i][offsets['posis']]
else:
all_data_arr[i][offsets['total']] = all_data_arr[i][offsets['posis']] + all_data_arr[i-1][offsets['total']]
all_data_np = np.array(all_data_arr)
# +
#for line in all_data_arr:
# print(line)
# +
updated = (dt.strptime(DT_OFFSET, "%Y/%m/%d") + td(days=int(all_data_np[-1][0]))).isoformat()[:10]
with open("mhlw.prev.tmp", "rt") as f:
prev = f.read().rstrip()
print("updated: {}, prev: {}".format(updated, prev))
if prev == updated:
print("maybe the same data, nothing to do.")
if "ipy" in sys.argv[0]:
pass#exit()
else:
sys.exit()
with open("mhlw.prev.tmp", "wt") as f:
f.write(updated)
# -
from_date = dt.strptime(DT_OFFSET, "%Y/%m/%d")
xbins = [from_date + td(days=i) for i in range(dt_range)]
days2pred = 4 * 7
xbins_pred = [from_date + td(days=i) for i in range(dt_range + days2pred)]
ave_mov_days = 7
# 移動平均を算出する
posis_mov_mean = moving_average(all_data_np[:, offsets['posis']])
ratio_mov_mean = moving_average(all_data_np[:, offsets['ratio']])
print("陽性者数(移動平均): {}".format(posis_mov_mean[-1]))
print(" 陽性率(移動平均): {}".format(ratio_mov_mean[-1]))
# +
#all_data_np
# -
X = np.arange(0, len(posis_mov_mean))[:, np.newaxis]
X_pred = np.arange(0, len(xbins_pred))[:, np.newaxis]
y_posis = get_gpr_predict(X, all_data_np[:, offsets['posis']], X_pred, 80, 10, 200)
y_ratio = get_gpr_predict(X, all_data_np[:, offsets['ratio']], X_pred, 80, 10, 200)
fig = make_subplots(specs=[[{"secondary_y": True}]])
fig.add_trace(go.Scatter(x=xbins, y=all_data_np[:, offsets['posis']], mode='markers', name='陽性者数', marker=dict(size=4)), secondary_y=False)
fig.add_trace(go.Scatter(x=xbins_pred, y=y_posis, mode='lines', name='予測値', line=dict(width=1)), secondary_y=False)
fig.add_trace(go.Bar(x=xbins, y=posis_mov_mean, name='移動平均', opacity=0.5), secondary_y=False)
fig.add_trace(go.Scatter(x=xbins, y=all_data_np[:, offsets['ratio']], mode='markers', name='陽性率[%]', marker=dict(size=4)), secondary_y=True)
fig.add_trace(go.Scatter(x=xbins_pred, y=y_ratio, mode='lines', name='予測値', line=dict(width=1)), secondary_y=True)
fig.add_trace(go.Bar(x=xbins, y=ratio_mov_mean, name='移動平均', opacity=0.8, marker_color='yellow'), secondary_y=True)
fig.update_layout(
barmode='overlay',
xaxis=dict(title='日付', type='date', dtick=1209600000.0, tickformat="%_m/%-d",
range=[xbins[30], xbins_pred[-1]]),
yaxis=dict(title='人数', type='log'),
yaxis2=dict(title='陽性率[%]', range=[0,50]),
title='全国 新型コロナ 陽性者数/陽性率 ({})'.format(today_str),
)
show_and_save_plotly(fig, "mhlw-posis.jpg", js=False)
# +
#fft_denoise(xbins[200:], all_data_np[200:, offsets['posis']], freq_int=0.15, freq_th=0.07, freq_min_A=0.01)
# -
y_tests = get_gpr_predict(X, all_data_np[:, offsets['pcrs']], X_pred, 1, 1, 5)
# 移動平均を算出する
tests_mov_mean = moving_average(all_data_np[:, offsets['pcrs']])
print("検査人数(移動平均): {}".format(tests_mov_mean[-1]))
fig = make_subplots(specs=[[{"secondary_y": True}]])
fig.add_trace(go.Scatter(x=xbins, y=all_data_np[:, offsets['pcrs']], mode='markers', name='検査人数', marker=dict(size=4)), secondary_y=False)
fig.add_trace(go.Scatter(x=xbins_pred, y=y_tests, mode='lines', name='予測値', line=dict(width=1)), secondary_y=False)
fig.add_trace(go.Bar(x=xbins, y=tests_mov_mean, name='移動平均', opacity=0.5), secondary_y=False)
fig.add_trace(go.Scatter(x=xbins, y=all_data_np[:, offsets['ratio']], mode='markers', name='陽性率[%]', marker=dict(size=4)), secondary_y=True)
fig.add_trace(go.Scatter(x=xbins_pred, y=y_ratio, mode='lines', name='予測値', line=dict(width=1)), secondary_y=True)
fig.add_trace(go.Bar(x=xbins, y=ratio_mov_mean, name='移動平均', opacity=0.8, marker_color='yellow'), secondary_y=True)
fig.update_layout(
barmode='overlay',
xaxis=dict(title='日付', type='date', dtick=1209600000.0, tickformat="%_m/%-d",
range=[xbins[30], xbins_pred[-1]]),
yaxis=dict(title='人数'),#, range=[0, np.max(y_tests)]),
yaxis2=dict(title='陽性率[%]', range=[0,50]),
title='全国 新型コロナ 検査人数/陽性率 ({})'.format(today_str),
)
show_and_save_plotly(fig, "mhlw-tests.jpg", js=False)
fig = make_subplots(specs=[[{"secondary_y": True}]])
fig.add_trace(go.Bar(x=xbins, y=all_data_np[:, offsets['total']],
name='陽性者', opacity=0.8, marker_color='#c08080'), secondary_y=False)
fig.add_trace(go.Bar(x=xbins, y=all_data_np[:, offsets['recov']],
name='退院者', opacity=0.8, marker_color='#00c000'), secondary_y=False)
fig.add_trace(go.Bar(x=xbins, y=all_data_np[:, offsets['cases']],
name='入院中', opacity=0.8, marker_color='yellow'), secondary_y=False)
deads = all_data_np[:, offsets['death']]
deads_os = 0
for i in range(7):
if deads[-(1+i)] == 0:
deads_os = i + 1
print("deads offset: {}".format(deads_os))
if deads_os != 0:
fig.add_trace(go.Scatter(x=xbins[:-deads_os], y=deads[:-deads_os], name="死者",
line=dict(width=1, color='magenta')), secondary_y=True)
else:
fig.add_trace(go.Scatter(x=xbins, y=deads, name="死者",
line=dict(width=1, color='magenta')), secondary_y=True)
fig.update_layout(
barmode='overlay',
xaxis=dict(title='日付', type='date', dtick=1209600000.0, tickformat="%_m/%-d",
range=[xbins[40], xbins[-1]]),
yaxis=dict(title='人数'),
yaxis2=dict(range=[0, np.max(all_data_np[:, offsets['death']])+10]),
title='全国 新型コロナ 陽性者/退院者/入院中/死者 ({})'.format(today_str),
)
show_and_save_plotly(fig, "mhlw-total.jpg", js=False)
tw_body_total = "全国 新型コロナ 累計陽性者/退院者/死者(" + today_str + ") "
tw_body_total += " https://geneasyura.github.io/cov19-hm/mhlw.html "
tw_body_tests = "全国 新型コロナ 検査人数/陽性率(" + today_str + ") "
tw_body_tests += " https://geneasyura.github.io/cov19-hm/mhlw.html "
tw_body_posis = "全国 新型コロナ 陽性者/陽性率(" + today_str + ") "
tw_body_posis += " https://geneasyura.github.io/cov19-hm/mhlw.html "
tw = get_twitter()
tweet_with_image(tw, "docs/images/mhlw-posis.jpg", tw_body_posis)
tweet_with_image(tw, "docs/images/mhlw-tests.jpg", tw_body_tests)
tweet_with_image(tw, "docs/images/mhlw-total.jpg", tw_body_total)
# 実効再生産数
ogiwara_uri = "https://toyokeizai.net/sp/visual/tko/covid19/csv/"
ern_file = "effective_reproduction_number.csv"
download_if_needed(ogiwara_uri, ern_file)
ern_data_arr = []
for i in np.arange(dt_range):
ern_data_arr.append([i, 0, 0, 0])
csv2array(ern_data_arr, 'ern', ern_file, 1)
ern_data_np = np.array(ern_data_arr)
#print(ern_data_np[:,1])
y_ern = get_gpr_predict(X, ern_data_np[:, 1], X_pred, 80, 10, 200)
fig = go.Figure()
fig.add_trace(go.Bar(x=xbins, y=ern_data_np[:, 1], name="実効再生産数", opacity=0.5))
fig.add_trace(go.Scatter(x=xbins_pred, y=y_ern, mode='lines', name='予測値', line=dict(width=1)))
fig.update_layout(
xaxis=dict(title='日付', type='date', dtick=1209600000.0, tickformat="%_m/%-d",
range=[xbins[44], xbins_pred[-1]]),
yaxis=dict(title='実効再生産'),
title='全国 新型コロナ 実効再生産数 ({})'.format(today_str),
)
show_and_save_plotly(fig, "ogiwara-ern.jpg", js=False)
tw_body_ern = "全国 新型コロナ 実効再生産数 ({})".format(today_str)
tw_body_ern += " https://geneasyura.github.io/cov19-hm/mhlw.html "
tweet_with_image(tw, "docs/images/ogiwara-ern.jpg", tw_body_ern)
title = '全国 新型コロナ 新規陽性者移動平均/指数近似 (' + today_str + ')'
xdata = np.array(xbins)
#ydata = all_data_np[:, offsets['posis']]
ydata = posis_mov_mean
xos = 310
make_exp_fit_graph(tw,
xdata[xos:], ydata[xos:],
title, "mhlw-fit.jpg",
"mhlw-doubling-time.html", "mhlw.html", needs_tw=False)
np_influ = np.loadtxt("csv/influ.csv", skiprows=1, delimiter=',', dtype=float)
np_influ
# 0-max normalization
for j in np.arange(1, np_influ.shape[1]):
peak = 1#np_influ[:, j].max()
np_influ[:, j] = np_influ[:, j] / peak
mena_influ = np_influ[:, 1:].mean(axis=1)
# W36 (2020/8/31) から
os_w36 = (dt(2020, 8, 31) - from_date).days
#print(from_date, w36of2020, os_w36)
xbins = all_data_np[os_w36:, offsets['dates']]
ybins = all_data_np[os_w36:, offsets['posis']]
y_by_week = []
w_by_week = []
for i in np.arange(0, len(xbins), 7):
w_by_week.append(int(36 + (i / 7)))
y_by_week.append(int(ybins[i:i + 7].sum()))
print(y_by_week)
print(w_by_week)
def poisson_func(x, a, b):
return b * ((a**x * np.exp(-a)) / factorial(x))
y_values = np.array(y_by_week) - min(y_by_week)
y_values[:4] = 0
(a, b), p0 = curve_fit(
poisson_func, np.arange(len(w_by_week)), y_values, maxfev=1000
)
print("a:{}, b:{}, p0:{}".format(a, b, p0))
xhat = np.arange(40)# + w_by_week[0]
yhat = poisson_func(xhat, a, b) + min(y_by_week)
fig = make_subplots(specs=[[{"secondary_y": True}]])
fig.add_trace(go.Scatter(
x=w_by_week, y=y_by_week,
mode='lines+markers', name='COVID-19',
line=dict(width=.5),
marker=dict(size=5)), secondary_y=False)
#fig.add_trace(go.Scatter(
# x=xhat+w_by_week[0], y=yhat,
# mode='lines', name='ポワソン分布予測',
# line=dict(width=.5)),
# secondary_y=False)
fig.add_trace(go.Scatter(
x=np_influ[:, 0], y=mena_influ,
line=dict(width=.7),
name='インフル平均', opacity=0.5), secondary_y=True)
for j in np.arange(1, np_influ.shape[1]):
fig.add_trace(go.Scatter(
x=np_influ[:, 0], y=np_influ[:, j],
line=dict(width=.7),
name='インフル{}'.format(2020-j), opacity=0.5), secondary_y=True)
fig.update_layout(
xaxis=dict(title='週数'),
yaxis=dict(title='COVID-19 感染者数'),
yaxis2=dict(title='インフルエンザ 感染者数'),
title='全国 新型コロナ インフルとの比較 ({})'.format(today_str),
)
show_and_save_plotly(fig, "mhlw-influ.jpg", js=False)
tw_body_influ = "全国 新型コロナ インフルとの比較 ({})".format(today_str)
tw_body_influ += " https://geneasyura.github.io/cov19-hm/influ.html "
tweet_with_image(tw, "docs/images/mhlw-influ.jpg", tw_body_influ)
| mhlw.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="Yln-wzDzE2XB"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import math
from sklearn.ensemble import RandomForestClassifier
from sklearn import datasets
from sklearn import metrics
from sklearn.model_selection import train_test_split
from ipywidgets import IntProgress
from IPython.display import display
# + colab={"base_uri": "https://localhost:8080/"} id="kUIfTBwC7a81" outputId="aad68592-8dee-4b8a-cd44-555bcc07a7c7"
# !gdown --id '1FXlXCQI2PlBDasMxLsNZvAarq9_DHomU' --output taxa-bar-plots-left-trim-level6.csv
# + colab={"base_uri": "https://localhost:8080/", "height": 299} id="d6PvPMcktCEk" outputId="9d28dc9e-223d-4c09-9584-71661d903769"
train = pd.read_csv("taxa-bar-plots-left-trim-level6.csv",index_col='number')
train.head()
# + colab={"base_uri": "https://localhost:8080/"} id="nBHSTsyTedOr" outputId="da49b9e1-5051-4898-c0d1-e29d15889273"
train.info()
# + id="ef5oh2zU6cHw"
CAT_COL = ["index", "sex", "smoker", "Clinical"]
NUM_COL=[]
for i in range(len(train.columns)):
NUM_COL.append(train.columns[i])
NUM_COL.remove('index')
NUM_COL.remove('Clinical')
NUM_COL.remove('sex')
NUM_COL.remove('smoker')
cat_col = []
num_col = []
for col in train:
if col in CAT_COL:
cat_col.append(col)
elif col in NUM_COL:
num_col.append(col)
for col in cat_col:
train[col] = train[col].astype(str)
df_cat = train.loc[:,cat_col] # take all the categorical columns
df_cat = pd.get_dummies(df_cat) # one hot encoding
df_num = train.loc[:,num_col] # take all the numerical columns
df_final = pd.concat([df_cat, df_num], axis=1) # concat categorical/numerical data
# + colab={"base_uri": "https://localhost:8080/", "height": 299} id="4rsZbO2Dbb31" outputId="f4e8005d-655d-4f56-eefa-a2f598904b37"
df_final.head()
# + colab={"base_uri": "https://localhost:8080/"} id="ClBkPVtMq9hp" outputId="86af1256-5b12-403c-e264-6f533fd56ceb"
not_select = ["index", "Clinical", "input", "filtered", "denoised", "non-chimeric"]
train_select = train.drop(not_select,axis=1)
train_select.info()
# + id="Dv4LE3iTv2oJ"
cat_col = []
num_col = []
for col in train_select:
if col in CAT_COL:
cat_col.append(col)
elif col in NUM_COL:
num_col.append(col)
for col in cat_col:
if train_select[col].dtype != "O":
# print(col)
train_select[col] = train_select[col].astype(str)
df_cat_select = train_select.loc[:,cat_col] # take all the categorical columns
df_cat_select = pd.get_dummies(df_cat_select) # one hot encoding
df_num_select = train_select.loc[:,num_col] # take all the numerical columns
df_final_select = pd.concat([df_cat_select, df_num_select], axis=1) # concat categorical/numerical data
# + colab={"base_uri": "https://localhost:8080/", "height": 299} id="6uX8WzkkbOKO" outputId="43411911-535b-482c-e00a-33aa07fbee8b"
df_final_select.head()
# + [markdown] id="WlXuzo6BHkIq"
# #Random Forest Classifier
# + colab={"base_uri": "https://localhost:8080/"} id="oT7bA9UMwVDo" outputId="989890ad-680d-42a1-c0b9-01b1a0119fe8"
#Use RandomForestClassifier to predict Clinical
x = df_final_select
y = train["Clinical"]
# y = np.array(y,dtype=int)
X_train,X_test,y_train,y_test = train_test_split(x,y,test_size=0.1,random_state=0)
#RandomForest
rfc = RandomForestClassifier()
#rfc=RandomForestClassifier(n_estimators=100,n_jobs = -1,random_state =50, min_samples_leaf = 10)
rfc.fit(X_train,y_train)
y_predict = rfc.predict(X_train)
score_rfc = rfc.score(X_test,y_test)
print("Random Forest Accuracy = ",score_rfc*100," %")
# + colab={"base_uri": "https://localhost:8080/"} id="PwMtRn5DGUTc" outputId="21e3814c-cfae-4e74-b743-2ecaeceefa8c"
from sklearn.model_selection import KFold
x = df_final_select
y = train["Clinical"]
kf = KFold(n_splits=5)
best_accuracy = 0
for train_index , test_index in kf.split(x):
X_train, X_test, y_train, y_test = x.iloc[train_index], x.iloc[test_index], y.iloc[train_index], y.iloc[test_index]
rfc = RandomForestClassifier()
rfc.fit(X_train,y_train)
y_predict = rfc.predict(X_train)
accuracy = rfc.score(X_test,y_test)
print("Random Forest Accuracy = ",accuracy*100,"%")
if accuracy > best_accuracy:
best_accuracy = accuracy
best_rfc = rfc
print("Best Accuracy = ",best_accuracy*100,"%")
# + colab={"base_uri": "https://localhost:8080/"} id="lVsvVd54GmW6" outputId="fcb6e29d-06b9-48a4-ba28-2a02f4d5d18d"
#random forest final accuracy
x_train_new = x
y_train_new = train["Clinical"]
y_train_new = y_train_new.reset_index(drop=True)
y_pred = best_rfc.predict(x_train_new)
count = 0
for i in range(y_pred.shape[0]):
if y_pred[i] != y_train_new[i]:
count += 1
# print(y_pred[i],y_train_new[i])
rfc_accuracy = 1-count/y_pred.shape[0]
print("Random Forest Accuracy = ",rfc_accuracy*100,"%")
# + [markdown] id="9c86bBBnHotC"
# #SVM
# + colab={"base_uri": "https://localhost:8080/"} id="-Qzl52nywn9w" outputId="2d77d240-c150-4eb1-d8dc-acd8d9cbc1fa"
from sklearn import svm
#Use SVM to predict Clinical
x = df_final_select
y = train["Clinical"]
# y = np.array(y,dtype=int)
X_train,X_test,y_train,y_test = train_test_split(x,y,test_size=0.1,random_state=0)
clf = svm.SVC()
clf.fit(X_train,y_train)
y_predict = clf.predict(X_train)
score_clf = clf.score(X_test,y_test)
print("SVM Accuracy = ",score_clf*100," %")
# + colab={"base_uri": "https://localhost:8080/"} id="d2IT2vHgHa8j" outputId="040f80a5-2f29-4ca4-b6bb-5bb48d20f89d"
from sklearn.model_selection import KFold
x = df_final_select
y = train["Clinical"]
kf = KFold(n_splits=5)
best_accuracy = 0
for train_index , test_index in kf.split(x):
X_train, X_test, y_train, y_test = x.iloc[train_index], x.iloc[test_index], y.iloc[train_index], y.iloc[test_index]
clf = svm.SVC()
clf.fit(X_train,y_train)
y_predict = clf.predict(X_train)
accuracy = clf.score(X_test,y_test)
print("SVM Accuracy = ",accuracy*100,"%")
if accuracy > best_accuracy:
best_accuracy = accuracy
best_clf = clf
print("Best Accuracy = ",best_accuracy*100,"%")
# + colab={"base_uri": "https://localhost:8080/"} id="FAjipyAMHbwi" outputId="cb6128de-1997-4600-81a8-6d83096bb41b"
#SVM final accuracy
x_train_new = x
y_train_new = train["Clinical"]
y_train_new = y_train_new.reset_index(drop=True)
y_pred = best_clf.predict(x_train_new)
count = 0
for i in range(y_pred.shape[0]):
# print(y_pred[i],y_train_new[i])
if y_pred[i] != y_train_new[i]:
count += 1
clf_accuracy = 1-count/y_pred.shape[0]
print("SVM Accuracy = ",clf_accuracy*100,"%")
# + [markdown] id="lARu7KV5HrH0"
# #Neural network MLPClassifier
# + colab={"base_uri": "https://localhost:8080/"} id="8vWodBUQwvoX" outputId="393bd750-8409-44da-8c42-7c722b407c31"
from sklearn.neural_network import MLPClassifier
#Use Neural Network MLPClassifier to predict Clinical
x = df_final_select
y = train["Clinical"]
# y = np.array(y,dtype=int)
X_train,X_test,y_train,y_test = train_test_split(x,y,test_size=0.1,random_state=0)
nnclf = MLPClassifier(solver='adam', alpha=1e-5, hidden_layer_sizes=(50, 30), random_state=1, max_iter=2000)
nnclf.fit(X_train,y_train)
y_predict = nnclf.predict(X_train)
score_nnclf = nnclf.score(X_test,y_test)
print("Neural Network Accuracy = ",score_nnclf*100," %")
# + colab={"base_uri": "https://localhost:8080/"} id="CoT7SNAQIww6" outputId="3cc91398-334c-4bf8-e289-e2468bdf2922"
from sklearn.model_selection import KFold
x = df_final_select
y = train["Clinical"]
kf = KFold(n_splits=5)
best_accuracy = 0
for train_index , test_index in kf.split(x):
X_train, X_test, y_train, y_test = x.iloc[train_index], x.iloc[test_index], y.iloc[train_index], y.iloc[test_index]
nnclf = MLPClassifier(solver='adam', alpha=1e-5, hidden_layer_sizes=(50, 30), random_state=1, max_iter=2000)
nnclf.fit(X_train,y_train)
y_predict = nnclf.predict(X_train)
accuracy = nnclf.score(X_test,y_test)
print("NN Accuracy = ",accuracy*100,"%")
if accuracy > best_accuracy:
best_accuracy = accuracy
best_nnclf = nnclf
print("NN Accuracy = ",best_accuracy*100,"%")
# + colab={"base_uri": "https://localhost:8080/"} id="ZbVGlnT_I7D5" outputId="a6a6c66c-7ae1-4d4f-c146-7a7013791c93"
#NN final accuracy
x_train_new = x
y_train_new = train["Clinical"]
y_train_new = y_train_new.reset_index(drop=True)
y_pred = best_nnclf.predict(x_train_new)
count = 0
for i in range(y_pred.shape[0]):
# print(y_pred[i],y_train_new[i])
if y_pred[i] != y_train_new[i]:
count += 1
nnclf_accuracy = 1-count/y_pred.shape[0]
print("NN Accuracy = ",nnclf_accuracy*100,"%")
# + [markdown] id="dG7iRLaQHxpa"
# #Logistic Regression
# + colab={"base_uri": "https://localhost:8080/"} id="2a8ZSpJW06uP" outputId="feff014c-a816-44bf-801c-8f27f08e31a2"
from sklearn.linear_model import LogisticRegression
#Use Logistic Regression to predict Clinical
x = df_final_select
y = train["Clinical"]
# y = np.array(y,dtype=int)
X_train,X_test,y_train,y_test = train_test_split(x,y,test_size=0.1,random_state=0)
logclf = LogisticRegression(random_state=0).fit(X_train,y_train)
logclf.predict(X_train)
logclf.predict_proba(X_train)
score_logclf = logclf.score(X_test,y_test)
print("Logistic Regression Accuracy = ",score_logclf*100," %")
# + colab={"base_uri": "https://localhost:8080/"} id="vSWmvYzoJQwz" outputId="9946746c-60fa-483f-f610-1ad203016bf9"
from sklearn.model_selection import KFold
x = df_final_select
y = train["Clinical"]
kf = KFold(n_splits=5)
best_accuracy = 0
for train_index , test_index in kf.split(x):
X_train, X_test, y_train, y_test = x.iloc[train_index], x.iloc[test_index], y.iloc[train_index], y.iloc[test_index]
logclf = LogisticRegression(random_state=0).fit(X_train,y_train)
y_predict = logclf.predict(X_train)
accuracy = logclf.score(X_test,y_test)
print("Logistic Regression Accuracy = ",accuracy*100,"%")
if accuracy > best_accuracy:
best_accuracy = accuracy
best_logclf = logclf
print("Logistic Regression Accuracy = ",best_accuracy*100,"%")
# + colab={"base_uri": "https://localhost:8080/"} id="aMrM4R1NJTIj" outputId="adc3d41a-aede-4ba0-f050-0113a6ceb4ac"
#Logistic final accuracy
x_train_new = x
y_train_new = train["Clinical"]
y_train_new = y_train_new.reset_index(drop=True)
y_pred = best_logclf.predict(x_train_new)
count = 0
for i in range(y_pred.shape[0]):
# print(y_pred[i],y_train_new[i])
if y_pred[i] != y_train_new[i]:
count += 1
logclf_accuracy = 1-count/y_pred.shape[0]
print("Logistic Regression Accuracy = ",logclf_accuracy*100,"%")
| Qiime2_Taxonomy_Train_Allsamples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/zarrinan/DS-Sprint-02-Storytelling-With-Data/blob/master/module3-make-explanatory-visualizations/LS_DS_123_Make_explanatory_visualizations_LESSON.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="-8-trVo__vRE" colab_type="text"
# _Lambda School Data Science_
#
# # Choose appropriate visualizations
#
#
# Recreate this [example by FiveThirtyEight:](https://fivethirtyeight.com/features/al-gores-new-movie-exposes-the-big-flaw-in-online-movie-ratings/)
# + id="ya_w5WORGs-n" colab_type="code" outputId="abef29d7-ff23-409e-b8ce-8bca459102d6" colab={"base_uri": "https://localhost:8080/", "height": 628}
from IPython.display import display, Image
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# !pip install --upgrade seaborn
import seaborn as sns
url = 'https://fivethirtyeight.com/wp-content/uploads/2017/09/mehtahickey-inconvenient-0830-1.png'
example = Image(url=url, width=500)
display(example)
# + [markdown] id="HioPkYtUG03B" colab_type="text"
# Using this data:
#
# https://github.com/fivethirtyeight/data/tree/master/inconvenient-sequel
#
# ### Stretch goals
#
# Recreate more examples from [FiveThityEight's shared data repository](https://data.fivethirtyeight.com/).
#
# For example:
# - [thanksgiving-2015](https://fivethirtyeight.com/features/heres-what-your-part-of-america-eats-on-thanksgiving/) ([`altair`](https://altair-viz.github.io/gallery/index.html#maps))
# - [candy-power-ranking](https://fivethirtyeight.com/features/the-ultimate-halloween-candy-power-ranking/) ([`statsmodels`](https://www.statsmodels.org/stable/index.html))
# + id="7NNwyEb0iOcS" colab_type="code" colab={}
# + id="P_3fLWPpiRrZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 480} outputId="ce8b3941-a8b1-454d-89b5-2a961f04d262"
fake = pd.Series([2,3,4,6,14,2,3])
plt.style.use('fivethirtyeight')
ax = fake.plot.bar(color='#EC713B', width=0.9)
ax.set(xlabel='Ratings', ylabel='Percent of total votes',yticks=range(0,50, 10))
ax.text(x=-2, y=50, s ="'An Inconvinient Sequel: Truth To Power' is divisible", fontsize=16, fontweight='bold')
ax.text(x=-2, y=46, s ='IMDb ratings for the film as of Aug. 29', fontsize=16)
ax.tick_params(labelrotation=0)
# + id="N89pO3kX_-7p" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 355} outputId="4e69af83-a21d-4ea0-f7d7-107b7af015ba"
df = pd.read_csv('https://raw.githubusercontent.com/fivethirtyeight/data/master/inconvenient-sequel/ratings.csv')
print(df.shape)
df.head()
# + id="Vb-z_JzAauC4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 886} outputId="e5effd31-3f4b-4956-8598-731cf2cf02e2"
df.sample(1).T
# + id="FrL7sPmOc4sY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 136} outputId="21c914ec-06c7-4c10-cced-d532b3a0c485"
df.timestamp = pd.to_datetime(df.timestamp)
df.timestamp.describe()
# + id="hPFlSZb0dF_T" colab_type="code" colab={}
df.set_index(df.timestamp, inplace=True)
# + id="-1-3twtqdPJd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 369} outputId="a187f880-81f5-40f5-fc60-73d7188f8334"
df.head()
# + id="tgyzGpthdUsN" colab_type="code" colab={}
df['2017-08-29']
# + id="v2rZep2Edl5M" colab_type="code" colab={}
df.category.value_counts()
# + id="iHWPIFtUd7YR" colab_type="code" colab={}
df.link.value_counts()
# + id="AS6-DGMPeYMj" colab_type="code" colab={}
df[df.category=='IMDb users']
# + id="frSTKYZ4e8rM" colab_type="code" colab={}
lastday = df['2017-08-29']
lastday[lastday.category=='IMDb users'].respondents.plot();
# + id="B2pqvB78f8yC" colab_type="code" colab={}
final = df.tail(1)
columns = ['1_pct', '2_pct','3_pct','4_pct','5_pct','6_pct','7_pct','8_pct','9_pct','10_pct']
data = final[columns].T
# + id="ekemI3swghFu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 111} outputId="29173c39-5641-4876-832a-39208a07460d"
data.T
# + id="_dpRDZbtgkWY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 407} outputId="4d881d44-e330-45d5-8e3f-fab8adf42de3"
data.plot.bar()
#columns = ['{}_pct'.format(k) for k in range(1,11)]
#ax_real.get_legend().remove()
# + id="V9rTQJ4phZcv" colab_type="code" colab={}
data.index = range(1,11)
# + id="KX55L3Whh6g5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="30b58eed-ba63-4cb8-ff0b-a907e43d2aba"
data.head()
# + id="fJnOYW3oiBCX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 388} outputId="e1410e38-a444-4e49-e3c7-a7605210b0d9"
data.plot.bar()
# + id="6Zx2HrJljdzR" colab_type="code" colab={}
colors = ['#333333'] *10
colors[0] = '#ec713b'
# + id="Cq0Y2WfKjaFy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 542} outputId="094c7212-13c8-41f4-dcc3-01c24f7ef750"
plt.style.use('fivethirtyeight')
ax_real = data.plot.bar(color=colors, width=0.9, legend=False);
ax_real.set(xlabel='Rating',
ylabel='Percent of total votes',
yticks=range(0,50, 10))
ax_real.text(x=-2, y=50, s ="'An Inconvinient Sequel: Truth To Power' is divisible",
fontsize=16, fontweight='bold')
ax_real.text(x=-2, y=47, s ='IMDb ratings for the film as of Aug. 29', fontsize=16, fontweight='normal')
ax_real.tick_params(labelrotation=0)
# + id="b8lLOJS5ASJ9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 438} outputId="a9a428c9-73ea-46f1-904c-8679273a7ae0"
display(example)
# + id="aQiHvxopIn9Q" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 136} outputId="4800fa28-ada3-44f7-80e2-423b9c005ddd"
df.timestamp = pd.to_datetime(df.timestamp)
df.timestamp.describe()
# + id="ZoNOue36zS12" colab_type="code" colab={}
df.set_index(df.timestamp, inplace=True)
# + id="E4gLbrwjzcD1" colab_type="code" colab={}
# + id="L7X4XcubgWx7" colab_type="code" colab={}
| module3-make-explanatory-visualizations/LS_DS_123_Make_explanatory_visualizations_LESSON.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # PROBLEMAS DIVERSOS
# <h3>1.</h3>
# Realizar una función que permita la carga de n alumnos. Por cada alumno se deberá preguntar el nombre completo y permitir el ingreso de 3 notas. Las notas deben estar comprendidas entre 0 y 10. Devolver el listado de alumnos.
# +
nombres=[]
notas=[]
for x in range(3):
nom=input("Ingrese el nombre del alumno:")
nombres.append(nom)
no1=int(input("Ingrese la primer nota:"))
no2=int(input("Ingrese la segunda nota:"))
notas.append([no1,no2])
for x in range(3):
print(nombres[x],notas[x][0],notas[x][1])
# -
# ### 2.
# Definir una función que dado un listado de alumnos evalúe cuántos aprobaron y cuántos desaprobaron, teniendo en cuenta que se aprueba con 4. La nota será el promedio de las 3 notas para cada alumno.
# ### 3.
# Informar el promedio de nota del curso total.
# +
n == int(input("Ingrese la cantidad de notas a promediar: "))
suma==0
i=1
while(1 <=n)
print("Ingrese la nota numero: ",i)
nota == float(input())
suma == suma+nota
i+=1
prom == suma / n
print("El promedio de notas es: ",prom)
# -
# ### 4.
# Realizar una función que indique quién tuvo el promedio más alto y quién tuvo la nota promedio más baja.
# +
def find_lowest_grade(students):
grades = [s[1] for s in students]
grades = sorted(grades)
min_grade = grades[0]
for i in range(len(grades)):
if grades[i] > min_grade:
return grades[i]
return min_grade
def find_students_by_grade(students, grade):
return [s[0] for s in students if s[1] == grade]
if __name__ == '__main__':
students = []
for _ in range(int(input())):
name = input()
score = float(input())
students.append([name, score])
lowest_grade = find_second_lowest_grade(students)
students_by_grade = find_students_by_grade(students, second_lowest_grade)
students_by_grade = sorted(students_by_grade)
for s in students_by_grade:
print(s)
# -
6
# ### 5.
# Realizar una función que permita buscar un alumno por nombre, siendo el nombre completo o parcial, y devuelva una lista con los n alumnos que concuerden con ese nombre junto con todos sus datos, incluido el promedio de sus notas.
| Ejercicios_Modulo2_resueltos/Ejercicios_Jupiter_modulo2_resuelto/Problemas Diversos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
from sklearn.model_selection import ShuffleSplit, KFold
from sklearn.ensemble import RandomForestClassifier, ExtraTreesClassifier, GradientBoostingClassifier
from xgboost import XGBClassifier
from sklearn.metrics import roc_auc_score
class blending_layer1(object):
def __init__(self, train_x, train_y, val_x, val_y, test_x):
self.train_x = train_x
self.train_y = train_y
self.val_x = val_x
self.val_y = val_y
self.test_x = test_x
def shuffle_data(self, model=None):
ss = ShuffleSplit(n_splits=4, random_state=0, test_size=.25)
for n_fold, (trn_idx, val_idx) in enumerate(ss.split(self.train_x)):
if n_fold == 0:
rf_train_x = self.train_x[val_idx]
rf_train_y = self.train_y[val_idx]
elif n_fold == 1:
et_train_x = self.train_x[val_idx]
et_train_y = self.train_y[val_idx]
elif n_fold == 2:
gbdt_train_x = self.train_x[val_idx]
gbdt_train_y = self.train_y[val_idx]
else:
xgb_train_x = self.train_x[val_idx]
xgb_train_y = self.train_y[val_idx]
if model == 'rf':
return rf_train_x, rf_train_y
elif model == 'et':
return et_train_x, et_train_y
elif model == 'gbdt':
return gbdt_train_x, gbdt_train_y
elif model == 'xgb':
return xgb_train_x, xgb_train_y
else:
return ss
def rf_model(self, parameters=None):
rf_train_x, rf_train_y = self.shuffle_data('rf')
kf = KFold(n_splits=5, shuffle=True, random_state=0)
rf_val_pred = np.zeros((rf_train_x.shape[0],))
rf_off_val_pred = np.zeros((self.val_x.shape[0],))
rf_test_pred = np.zeros((self.test_x.shape[0],))
for n_fold, (trn_idx, val_idx) in enumerate(kf.split(rf_train_x)):
rf = RandomForestClassifier(**parameters)
rf.fit(rf_train_x[trn_idx], rf_train_y[trn_idx])
rf_val_pred[val_idx] = rf.predict_proba(rf_train_x[val_idx])[:, 1]
rf_off_val_pred += rf.predict_proba(self.val_x)[:, 1] / kf.n_splits
rf_test_pred += rf.predict_proba(self.test_x)[:, 1] / kf.n_splits
print('Fold %d auc score: %.6f'%(n_fold+1, roc_auc_score(rf_train_y[val_idx], rf_val_pred[val_idx])))
print('Validate auc score:', roc_auc_score(rf_train_y, rf_val_pred))
print('Off auc score:', roc_auc_score(self.val_y, rf_off_val_pred))
return rf_val_pred, rf_off_val_pred, rf_test_pred
def et_model(self, parameters=None):
et_train_x, et_train_y = self.shuffle_data('et')
kf = KFold(n_splits=5, shuffle=True, random_state=0)
et_val_pred = np.zeros((et_train_x.shape[0],))
et_off_val_pred = np.zeros((self.val_x.shape[0],))
et_test_pred = np.zeros((self.test_x.shape[0],))
for n_fold, (trn_idx, val_idx) in enumerate(kf.split(et_train_x)):
et = ExtraTreesClassifier(**parameters)
et.fit(et_train_x[trn_idx], et_train_y[trn_idx])
et_val_pred[val_idx] = et.predict_proba(et_train_x[val_idx])[:, 1]
et_off_val_pred += et.predict_proba(self.val_x)[:, 1] / kf.n_splits
et_test_pred += et.predict_proba(self.test_x)[:, 1] / kf.n_splits
print('Fold %d auc score: %.6f'%(n_fold+1, roc_auc_score(et_train_y[val_idx], et_val_pred[val_idx])))
print('Validate auc score:', roc_auc_score(et_train_y, et_val_pred))
print('Off auc score:', roc_auc_score(self.val_y, et_off_val_pred))
return et_val_pred, et_off_val_pred, et_test_pred
def gbdt_model(self, parameters=None):
gbdt_train_x, gbdt_train_y = self.shuffle_data('gbdt')
kf = KFold(n_splits=5, shuffle=True, random_state=0)
gbdt_val_pred = np.zeros((gbdt_train_x.shape[0],))
gbdt_off_val_pred = np.zeros((self.val_x.shape[0],))
gbdt_test_pred = np.zeros((self.test_x.shape[0],))
for n_fold, (trn_idx, val_idx) in enumerate(kf.split(gbdt_train_x)):
gbdt = GradientBoostingClassifier(**parameters)
gbdt.fit(gbdt_train_x[trn_idx], gbdt_train_y[trn_idx])
gbdt_val_pred[val_idx] = gbdt.predict_proba(gbdt_train_x[val_idx])[:, 1]
gbdt_off_val_pred += gbdt.predict_proba(self.val_x)[:, 1] / kf.n_splits
gbdt_test_pred += gbdt.predict_proba(self.test_x)[:, 1] / kf.n_splits
print('Fold %d auc score: %.6f'%(n_fold+1, roc_auc_score(gbdt_train_y[val_idx], gbdt_val_pred[val_idx])))
print('Validate auc score:', roc_auc_score(gbdt_train_y, gbdt_val_pred))
print('Off auc score:', roc_auc_score(self.val_y, gbdt_off_val_pred))
return gbdt_val_pred, gbdt_off_val_pred, gbdt_test_pred
def xgb_model(self, parameters=None):
xgb_train_x, xgb_train_y = self.shuffle_data('xgb')
kf = KFold(n_splits=5, shuffle=True, random_state=0)
xgb_val_pred = np.zeros((xgb_train_x.shape[0],))
xgb_off_val_pred = np.zeros((self.val_x.shape[0],))
xgb_test_pred = np.zeros((self.test_x.shape[0],))
for n_fold, (trn_idx, val_idx) in enumerate(kf.split(xgb_train_x)):
xgb = XGBClassifier(**parameters)
xgb.fit(xgb_train_x[trn_idx], xgb_train_y[trn_idx])
xgb_val_pred[val_idx] = xgb.predict_proba(xgb_train_x[val_idx])[:, 1]
xgb_off_val_pred += xgb.predict_proba(self.val_x)[:, 1] / kf.n_splits
xgb_test_pred += xgb.predict_proba(self.test_x)[:, 1] / kf.n_splits
print('Fold %d auc score: %.6f'%(n_fold+1, roc_auc_score(xgb_train_y[val_idx], xgb_val_pred[val_idx])))
print('Validate auc score:', roc_auc_score(xgb_train_y, xgb_val_pred))
print('Off auc score:', roc_auc_score(self.val_y, xgb_off_val_pred))
return xgb_val_pred, xgb_off_val_pred, xgb_test_pred
def merge_data(self):
rf_val_pred, rf_off_val_pred, rf_test_pred = self.rf_model(parameters={"n_jobs": -1})
et_val_pred, et_off_val_pred, et_test_pred = self.et_model(parameters={"n_jobs": -1})
gbdt_val_pred, gbdt_off_val_pred, gbdt_test_pred = self.gbdt_model(parameters={"learning_rate": 0.3})
xgb_val_pred, xgb_off_val_pred, xgb_test_pred = self.xgb_model(parameters={"n_jobs": -1})
val_pred = np.zeros((self.train_x.shape[0],))
ss = self.shuffle_data()
for n_fold, (trn_idx, val_idx) in enumerate(ss.split(self.train_x)):
if n_fold == 0:
val_pred[val_idx] = rf_val_pred
elif n_fold == 1:
val_pred[val_idx] = et_val_pred
elif n_fold == 2:
val_pred[val_idx] = gbdt_val_pred
else:
val_pred[val_idx] = xgb_val_pred
off_val_pred = (rf_off_val_pred + et_off_val_pred + gbdt_off_val_pred + xgb_off_val_pred) / 4
test_pred = (rf_test_pred + et_test_pred + gbdt_test_pred + xgb_test_pred) / 4
return val_pred, off_val_pred, test_pred
train_x, train_y, val_x, val_y, test_x = np.random.random((7000,10)), np.random.randint(0,2,size=(7000,)), np.random.random((3000,10)), np.random.randint(0,2,size=(3000,)), np.random.random((3000,10))
bld_layer = blending_layer1(train_x, train_y, val_x, val_y, test_x)
val_pred, off_val_pred, test_pred = bld_layer.merge_data()
# -
print(val_pred.shape, off_val_pred.shape, test_pred.shape)
train_x = pd.read_csv('../data/train_.csv')
test_x = pd.read_csv('../data/test_.csv')
train_y = pd.read_csv('../data/y_.csv')
val_x = pd.read_csv('../data/val_x_.csv')
val_y = pd.read_csv('../data/val_y_.csv')
val_x = val_x[train_x.columns]
test_x = test_x[train_x.columns]
train_x, train_y, val_x, val_y, test_x = train_x.values, train_y.values, val_x.values, val_y.values, test_x.values
bld_layer = blending_layer1(train_x, train_y, val_x, val_y, test_x)
val_pred, off_val_pred, test_pred = bld_layer.merge_data()
print(val_pred.shape, off_val_pred.shape, test_pred.shape)
val_pred, off_val_pred, test_pred = pd.DataFrame(val_pred), pd.DataFrame(off_val_pred), pd.DataFrame(test_pred)
val_pred.head()
val_pred.to_csv('../data/val_pred.csv', index=False)
off_val_pred.to_csv('../data/off_val_pred.csv', index=False)
test_pred.to_csv('../data/test_pred.csv',index=False)
# +
#print(train_x.shape, train_y.shape, val_x.shape, val_y.shape,test_x.shape)
# +
#train_x.head()
# +
#test_x.head()
# +
#val_x.head()
# -
| model/blending_layer1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
from tkinter import *
from sklearn.cluster import KMeans
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier
data=pd.read_csv('food.csv')
Breakfastdata=data['Breakfast']
BreakfastdataNumpy=Breakfastdata.to_numpy()
Lunchdata=data['Lunch']
LunchdataNumpy=Lunchdata.to_numpy()
Dinnerdata=data['Dinner']
DinnerdataNumpy=Dinnerdata.to_numpy()
Food_itemsdata=data['Food_items']
def show_entry_fields():
print("\n Age: %s\n Veg-NonVeg: %s\n Weight: %s kg\n Hight: %s cm\n" % (e1.get(), e2.get(),e3.get(), e4.get()))
def Weight_Loss():
show_entry_fields()
breakfastfoodseparated=[]
Lunchfoodseparated=[]
Dinnerfoodseparated=[]
breakfastfoodseparatedID=[]
LunchfoodseparatedID=[]
DinnerfoodseparatedID=[]
for i in range(len(Breakfastdata)):
if BreakfastdataNumpy[i]==1:
breakfastfoodseparated.append( Food_itemsdata[i] )
breakfastfoodseparatedID.append(i)
if LunchdataNumpy[i]==1:
Lunchfoodseparated.append(Food_itemsdata[i])
LunchfoodseparatedID.append(i)
if DinnerdataNumpy[i]==1:
Dinnerfoodseparated.append(Food_itemsdata[i])
DinnerfoodseparatedID.append(i)
# retrieving Lunch data rows by loc method |
LunchfoodseparatedIDdata = data.iloc[LunchfoodseparatedID]
LunchfoodseparatedIDdata=LunchfoodseparatedIDdata.T
#print(LunchfoodseparatedIDdata)
val=list(np.arange(5,15))
Valapnd=[0]+val
LunchfoodseparatedIDdata=LunchfoodseparatedIDdata.iloc[Valapnd]
LunchfoodseparatedIDdata=LunchfoodseparatedIDdata.T
#print(LunchfoodseparatedIDdata)
# retrieving Breafast data rows by loc method
breakfastfoodseparatedIDdata = data.iloc[breakfastfoodseparatedID]
breakfastfoodseparatedIDdata=breakfastfoodseparatedIDdata.T
val=list(np.arange(5,15))
Valapnd=[0]+val
breakfastfoodseparatedIDdata=breakfastfoodseparatedIDdata.iloc[Valapnd]
breakfastfoodseparatedIDdata=breakfastfoodseparatedIDdata.T
# retrieving Dinner Data rows by loc method
DinnerfoodseparatedIDdata = data.iloc[DinnerfoodseparatedID]
DinnerfoodseparatedIDdata=DinnerfoodseparatedIDdata.T
val=list(np.arange(5,15))
Valapnd=[0]+val
DinnerfoodseparatedIDdata=DinnerfoodseparatedIDdata.iloc[Valapnd]
DinnerfoodseparatedIDdata=DinnerfoodseparatedIDdata.T
#calculating BMI
age=int(e1.get())
veg=float(e2.get())
weight=float(e3.get())
height=float(e4.get())
bmi = weight/((height/100)**2)
agewiseinp=0
for lp in range (0,80,20):
test_list=np.arange(lp,lp+20)
for i in test_list:
if(i == age):
tr=round(lp/20)
agecl=round(lp/20)
#conditions
print("Your body mass index is: ", bmi)
if ( bmi < 16):
print("Acoording to your BMI, you are Severely Underweight")
clbmi=4
elif ( bmi >= 16 and bmi < 18.5):
print("Acoording to your BMI, you are Underweight")
clbmi=3
elif ( bmi >= 18.5 and bmi < 25):
print("Acoording to your BMI, you are Healthy")
clbmi=2
elif ( bmi >= 25 and bmi < 30):
print("Acoording to your BMI, you are Overweight")
clbmi=1
elif ( bmi >=30):
print("Acoording to your BMI, you are Severely Overweight")
clbmi=0
#converting into numpy array
DinnerfoodseparatedIDdata=DinnerfoodseparatedIDdata.to_numpy()
LunchfoodseparatedIDdata=LunchfoodseparatedIDdata.to_numpy()
breakfastfoodseparatedIDdata=breakfastfoodseparatedIDdata.to_numpy()
ti=(clbmi+agecl)/2
## K-Means Based Dinner Food
Datacalorie=DinnerfoodseparatedIDdata[1:,1:len(DinnerfoodseparatedIDdata)]
X = np.array(Datacalorie)
kmeans = KMeans(n_clusters=3, random_state=0).fit(X)
XValu=np.arange(0,len(kmeans.labels_))
# retrieving the labels for dinner food
dnrlbl=kmeans.labels_
## K-Means Based lunch Food
Datacalorie=LunchfoodseparatedIDdata[1:,1:len(LunchfoodseparatedIDdata)]
X = np.array(Datacalorie)
kmeans = KMeans(n_clusters=3, random_state=0).fit(X)
XValu=np.arange(0,len(kmeans.labels_))
# retrieving the labels for lunch food
lnchlbl=kmeans.labels_
## K-Means Based lunch Food
Datacalorie=breakfastfoodseparatedIDdata[1:,1:len(breakfastfoodseparatedIDdata)]
X = np.array(Datacalorie)
kmeans = KMeans(n_clusters=3, random_state=0).fit(X)
XValu=np.arange(0,len(kmeans.labels_))
# retrieving the labels for breakfast food
brklbl=kmeans.labels_
inp=[]
## Reading of the Dataet
datafin=pd.read_csv('nutrition_distriution.csv')
## train set
dataTog=datafin.T
bmicls=[0,1,2,3,4]
agecls=[0,1,2,3,4]
weightlosscat = dataTog.iloc[[1,2,7,8]]
weightlosscat=weightlosscat.T
weightgaincat= dataTog.iloc[[0,1,2,3,4,7,9,10]]
weightgaincat=weightgaincat.T
healthycat = dataTog.iloc[[1,2,3,4,6,7,9]]
healthycat=healthycat.T
weightlosscatDdata=weightlosscat.to_numpy()
weightgaincatDdata=weightgaincat.to_numpy()
healthycatDdata=healthycat.to_numpy()
weightlosscat=weightlosscatDdata[1:,0:len(weightlosscatDdata)]
weightgaincat=weightgaincatDdata[1:,0:len(weightgaincatDdata)]
healthycat=healthycatDdata[1:,0:len(healthycatDdata)]
weightlossfin=np.zeros((len(weightlosscat)*5,6),dtype=np.float32)
weightgainfin=np.zeros((len(weightgaincat)*5,10),dtype=np.float32)
healthycatfin=np.zeros((len(healthycat)*5,9),dtype=np.float32)
t=0
r=0
s=0
yt=[]
yr=[]
ys=[]
for zz in range(5):
for jj in range(len(weightlosscat)):
valloc=list(weightlosscat[jj])
valloc.append(bmicls[zz])
valloc.append(agecls[zz])
weightlossfin[t]=np.array(valloc)
yt.append(brklbl[jj])
t+=1
for jj in range(len(weightgaincat)):
valloc=list(weightgaincat[jj])
valloc.append(bmicls[zz])
valloc.append(agecls[zz])
weightgainfin[r]=np.array(valloc)
yr.append(lnchlbl[jj])
r+=1
for jj in range(len(healthycat)):
valloc=list(healthycat[jj])
valloc.append(bmicls[zz])
valloc.append(agecls[zz])
healthycatfin[s]=np.array(valloc)
ys.append(dnrlbl[jj])
s+=1
X_test=np.zeros((len(weightlosscat),6),dtype=np.float32)
print('####################')
#randomforest
for jj in range(len(weightlosscat)):
valloc=list(weightlosscat[jj])
valloc.append(agecl)
valloc.append(clbmi)
X_test[jj]=np.array(valloc)*ti
X_train=weightlossfin# Features
y_train=yt # Labels
#Create a Gaussian Classifier
clf=RandomForestClassifier(n_estimators=100)
#Train the model using the training sets y_pred=clf.predict(X_test)
clf.fit(X_train,y_train)
#print (X_test[1])
X_test2=X_test
y_pred=clf.predict(X_test)
print ('SUGGESTED FOOD ITEMS ::')
for ii in range(len(y_pred)):
if y_pred[ii]==2: #weightloss
print (Food_itemsdata[ii])
findata=Food_itemsdata[ii]
if int(veg)==1:
datanv=['Chicken Burger']
for it in range(len(datanv)):
if findata==datanv[it]:
print('VegNovVeg')
print('\n Thank You for taking our recommendations. :)')
def Weight_Gain():
show_entry_fields()
breakfastfoodseparated=[]
Lunchfoodseparated=[]
Dinnerfoodseparated=[]
breakfastfoodseparatedID=[]
LunchfoodseparatedID=[]
DinnerfoodseparatedID=[]
for i in range(len(Breakfastdata)):
if BreakfastdataNumpy[i]==1:
breakfastfoodseparated.append( Food_itemsdata[i] )
breakfastfoodseparatedID.append(i)
if LunchdataNumpy[i]==1:
Lunchfoodseparated.append(Food_itemsdata[i])
LunchfoodseparatedID.append(i)
if DinnerdataNumpy[i]==1:
Dinnerfoodseparated.append(Food_itemsdata[i])
DinnerfoodseparatedID.append(i)
# retrieving rows by loc method |
LunchfoodseparatedIDdata = data.iloc[LunchfoodseparatedID]
LunchfoodseparatedIDdata=LunchfoodseparatedIDdata.T
val=list(np.arange(5,15))
Valapnd=[0]+val
LunchfoodseparatedIDdata=LunchfoodseparatedIDdata.iloc[Valapnd]
LunchfoodseparatedIDdata=LunchfoodseparatedIDdata.T
# retrieving rows by loc method
breakfastfoodseparatedIDdata = data.iloc[breakfastfoodseparatedID]
breakfastfoodseparatedIDdata=breakfastfoodseparatedIDdata.T
val=list(np.arange(5,15))
Valapnd=[0]+val
breakfastfoodseparatedIDdata=breakfastfoodseparatedIDdata.iloc[Valapnd]
breakfastfoodseparatedIDdata=breakfastfoodseparatedIDdata.T
# retrieving rows by loc method
DinnerfoodseparatedIDdata = data.iloc[DinnerfoodseparatedID]
DinnerfoodseparatedIDdata=DinnerfoodseparatedIDdata.T
val=list(np.arange(5,15))
Valapnd=[0]+val
DinnerfoodseparatedIDdata=DinnerfoodseparatedIDdata.iloc[Valapnd]
DinnerfoodseparatedIDdata=DinnerfoodseparatedIDdata.T
#claculating BMI
age=int(e1.get())
veg=float(e2.get())
weight=float(e3.get())
height=float(e4.get())
bmi = weight/((height/100)**2)
for lp in range (0,80,20):
test_list=np.arange(lp,lp+20)
for i in test_list:
if(i == age):
tr=round(lp/20)
agecl=round(lp/20)
print("Your body mass index is: ", bmi)
if ( bmi < 16):
print("Acoording to your BMI, you are Severely Underweight")
clbmi=4
elif ( bmi >= 16 and bmi < 18.5):
print("Acoording to your BMI, you are Underweight")
clbmi=3
elif ( bmi >= 18.5 and bmi < 25):
print("Acoording to your BMI, you are Healthy")
clbmi=2
elif ( bmi >= 25 and bmi < 30):
print("Acoording to your BMI, you are Overweight")
clbmi=1
elif ( bmi >=30):
print("Acoording to your BMI, you are Severely Overweight")
clbmi=0
DinnerfoodseparatedIDdata=DinnerfoodseparatedIDdata.to_numpy()
LunchfoodseparatedIDdata=LunchfoodseparatedIDdata.to_numpy()
breakfastfoodseparatedIDdata=breakfastfoodseparatedIDdata.to_numpy()
ti=(bmi+agecl)/2
## K-Means Based Dinner Food
Datacalorie=DinnerfoodseparatedIDdata[1:,1:len(DinnerfoodseparatedIDdata)]
X = np.array(Datacalorie)
kmeans = KMeans(n_clusters=3, random_state=0).fit(X)
XValu=np.arange(0,len(kmeans.labels_))
# plt.bar(XValu,kmeans.labels_)
dnrlbl=kmeans.labels_
# plt.title("Predicted Low-High Weigted Calorie Foods")
## K-Means Based lunch Food
Datacalorie=LunchfoodseparatedIDdata[1:,1:len(LunchfoodseparatedIDdata)]
X = np.array(Datacalorie)
kmeans = KMeans(n_clusters=3, random_state=0).fit(X)
XValu=np.arange(0,len(kmeans.labels_))
# fig,axs=plt.subplots(1,1,figsize=(15,5))
# plt.bar(XValu,kmeans.labels_)
lnchlbl=kmeans.labels_
# plt.title("Predicted Low-High Weigted Calorie Foods")
## K-Means Based lunch Food
Datacalorie=breakfastfoodseparatedIDdata[1:,1:len(breakfastfoodseparatedIDdata)]
X = np.array(Datacalorie)
kmeans = KMeans(n_clusters=3, random_state=0).fit(X)
XValu=np.arange(0,len(kmeans.labels_))
# fig,axs=plt.subplots(1,1,figsize=(15,5))
# plt.bar(XValu,kmeans.labels_)
brklbl=kmeans.labels_
# plt.title("Predicted Low-High Weigted Calorie Foods")
inp=[]
## Reading of the Dataet
datafin=pd.read_csv('nutrition_distriution.csv')
datafin.head(5)
dataTog=datafin.T
bmicls=[0,1,2,3,4]
agecls=[0,1,2,3,4]
weightlosscat = dataTog.iloc[[1,2,7,8]]
weightlosscat=weightlosscat.T
weightgaincat= dataTog.iloc[[0,1,2,3,4,7,9,10]]
weightgaincat=weightgaincat.T
healthycat = dataTog.iloc[[1,2,3,4,6,7,9]]
healthycat=healthycat.T
weightlosscatDdata=weightlosscat.to_numpy()
weightgaincatDdata=weightgaincat.to_numpy()
healthycatDdata=healthycat.to_numpy()
weightlosscat=weightlosscatDdata[1:,0:len(weightlosscatDdata)]
weightgaincat=weightgaincatDdata[1:,0:len(weightgaincatDdata)]
healthycat=healthycatDdata[1:,0:len(healthycatDdata)]
weightlossfin=np.zeros((len(weightlosscat)*5,6),dtype=np.float32)
weightgainfin=np.zeros((len(weightgaincat)*5,10),dtype=np.float32)
healthycatfin=np.zeros((len(healthycat)*5,9),dtype=np.float32)
t=0
r=0
s=0
yt=[]
yr=[]
ys=[]
for zz in range(5):
for jj in range(len(weightlosscat)):
valloc=list(weightlosscat[jj])
valloc.append(bmicls[zz])
valloc.append(agecls[zz])
weightlossfin[t]=np.array(valloc)
yt.append(brklbl[jj])
t+=1
for jj in range(len(weightgaincat)):
valloc=list(weightgaincat[jj])
#print (valloc)
valloc.append(bmicls[zz])
valloc.append(agecls[zz])
weightgainfin[r]=np.array(valloc)
yr.append(lnchlbl[jj])
r+=1
for jj in range(len(healthycat)):
valloc=list(healthycat[jj])
valloc.append(bmicls[zz])
valloc.append(agecls[zz])
healthycatfin[s]=np.array(valloc)
ys.append(dnrlbl[jj])
s+=1
X_test=np.zeros((len(weightgaincat),10),dtype=np.float32)
print('####################')
# In[287]:
for jj in range(len(weightgaincat)):
valloc=list(weightgaincat[jj])
valloc.append(agecl)
valloc.append(clbmi)
X_test[jj]=np.array(valloc)*ti
X_train=weightgainfin# Features
y_train=yr # Labels
#Create a Gaussian Classifier
clf=RandomForestClassifier(n_estimators=100)
#Train the model using the training sets y_pred=clf.predict(X_test)
clf.fit(X_train,y_train)
X_test2=X_test
y_pred=clf.predict(X_test)
print ('SUGGESTED FOOD ITEMS ::')
for ii in range(len(y_pred)):
if y_pred[ii]==2:
print (Food_itemsdata[ii])
findata=Food_itemsdata[ii]
if int(veg)==1:
datanv=['Chicken Burger']
for it in range(len(datanv)):
if findata==datanv[it]:
print('VegNovVeg')
print('\n Thank You for taking our recommendations. :)')
def Healthy():
show_entry_fields()
breakfastfoodseparated=[]
Lunchfoodseparated=[]
Dinnerfoodseparated=[]
breakfastfoodseparatedID=[]
LunchfoodseparatedID=[]
DinnerfoodseparatedID=[]
for i in range(len(Breakfastdata)):
if BreakfastdataNumpy[i]==1:
breakfastfoodseparated.append( Food_itemsdata[i] )
breakfastfoodseparatedID.append(i)
if LunchdataNumpy[i]==1:
Lunchfoodseparated.append(Food_itemsdata[i])
LunchfoodseparatedID.append(i)
if DinnerdataNumpy[i]==1:
Dinnerfoodseparated.append(Food_itemsdata[i])
DinnerfoodseparatedID.append(i)
# retrieving rows by loc method |
LunchfoodseparatedIDdata = data.iloc[LunchfoodseparatedID]
LunchfoodseparatedIDdata=LunchfoodseparatedIDdata.T
val=list(np.arange(5,15))
Valapnd=[0]+val
LunchfoodseparatedIDdata=LunchfoodseparatedIDdata.iloc[Valapnd]
LunchfoodseparatedIDdata=LunchfoodseparatedIDdata.T
# retrieving rows by loc method
breakfastfoodseparatedIDdata = data.iloc[breakfastfoodseparatedID]
breakfastfoodseparatedIDdata=breakfastfoodseparatedIDdata.T
val=list(np.arange(5,15))
Valapnd=[0]+val
breakfastfoodseparatedIDdata=breakfastfoodseparatedIDdata.iloc[Valapnd]
breakfastfoodseparatedIDdata=breakfastfoodseparatedIDdata.T
# retrieving rows by loc method
DinnerfoodseparatedIDdata = data.iloc[DinnerfoodseparatedID]
DinnerfoodseparatedIDdata=DinnerfoodseparatedIDdata.T
val=list(np.arange(5,15))
Valapnd=[0]+val
DinnerfoodseparatedIDdata=DinnerfoodseparatedIDdata.iloc[Valapnd]
DinnerfoodseparatedIDdata=DinnerfoodseparatedIDdata.T
age=int(e1.get())
veg=float(e2.get())
weight=float(e3.get())
height=float(e4.get())
bmi = weight/((height/100)**2)
agewiseinp=0
for lp in range (0,80,20):
test_list=np.arange(lp,lp+20)
for i in test_list:
if(i == age):
tr=round(lp/20)
agecl=round(lp/20)
#conditions
print("Your body mass index is: ", bmi)
if ( bmi < 16):
print("Acoording to your BMI, you are Severely Underweight")
clbmi=4
elif ( bmi >= 16 and bmi < 18.5):
print("Acoording to your BMI, you are Underweight")
clbmi=3
elif ( bmi >= 18.5 and bmi < 25):
print("Acoording to your BMI, you are Healthy")
clbmi=2
elif ( bmi >= 25 and bmi < 30):
print("Acoording to your BMI, you are Overweight")
clbmi=1
elif ( bmi >=30):
print("Acoording to your BMI, you are Severely Overweight")
clbmi=0
DinnerfoodseparatedIDdata=DinnerfoodseparatedIDdata.to_numpy()
LunchfoodseparatedIDdata=LunchfoodseparatedIDdata.to_numpy()
breakfastfoodseparatedIDdata=breakfastfoodseparatedIDdata.to_numpy()
ti=(bmi+agecl)/2
Datacalorie=DinnerfoodseparatedIDdata[1:,1:len(DinnerfoodseparatedIDdata)]
X = np.array(Datacalorie)
kmeans = KMeans(n_clusters=3, random_state=0).fit(X)
XValu=np.arange(0,len(kmeans.labels_))
# fig,axs=plt.subplots(1,1,figsize=(15,5))
# plt.bar(XValu,kmeans.labels_)
dnrlbl=kmeans.labels_
# plt.title("Predicted Low-High Weigted Calorie Foods")
Datacalorie=LunchfoodseparatedIDdata[1:,1:len(LunchfoodseparatedIDdata)]
X = np.array(Datacalorie)
kmeans = KMeans(n_clusters=3, random_state=0).fit(X)
#print ('## Prediction Result ##')
#print(kmeans.labels_)
XValu=np.arange(0,len(kmeans.labels_))
# fig,axs=plt.subplots(1,1,figsize=(15,5))
# plt.bar(XValu,kmeans.labels_)
lnchlbl=kmeans.labels_
# plt.title("Predicted Low-High Weigted Calorie Foods")
Datacalorie=breakfastfoodseparatedIDdata[1:,1:len(breakfastfoodseparatedIDdata)]
X = np.array(Datacalorie)
kmeans = KMeans(n_clusters=3, random_state=0).fit(X)
XValu=np.arange(0,len(kmeans.labels_))
# fig,axs=plt.subplots(1,1,figsize=(15,5))
# plt.bar(XValu,kmeans.labels_)
brklbl=kmeans.labels_
# print (len(brklbl))
# plt.title("Predicted Low-High Weigted Calorie Foods")
inp=[]
## Reading of the Dataet
datafin=pd.read_csv('nutrition_distriution.csv')
datafin.head(5)
dataTog=datafin.T
bmicls=[0,1,2,3,4]
agecls=[0,1,2,3,4]
weightlosscat = dataTog.iloc[[1,2,7,8]]
weightlosscat=weightlosscat.T
weightgaincat= dataTog.iloc[[0,1,2,3,4,7,9,10]]
weightgaincat=weightgaincat.T
healthycat = dataTog.iloc[[1,2,3,4,6,7,9]]
healthycat=healthycat.T
weightlosscatDdata=weightlosscat.to_numpy()
weightgaincatDdata=weightgaincat.to_numpy()
healthycatDdata=healthycat.to_numpy()
weightlosscat=weightlosscatDdata[1:,0:len(weightlosscatDdata)]
weightgaincat=weightgaincatDdata[1:,0:len(weightgaincatDdata)]
healthycat=healthycatDdata[1:,0:len(healthycatDdata)]
weightlossfin=np.zeros((len(weightlosscat)*5,6),dtype=np.float32)
weightgainfin=np.zeros((len(weightgaincat)*5,10),dtype=np.float32)
healthycatfin=np.zeros((len(healthycat)*5,9),dtype=np.float32)
t=0
r=0
s=0
yt=[]
yr=[]
ys=[]
for zz in range(5):
for jj in range(len(weightlosscat)):
valloc=list(weightlosscat[jj])
valloc.append(bmicls[zz])
valloc.append(agecls[zz])
weightlossfin[t]=np.array(valloc)
yt.append(brklbl[jj])
t+=1
for jj in range(len(weightgaincat)):
valloc=list(weightgaincat[jj])
#print (valloc)
valloc.append(bmicls[zz])
valloc.append(agecls[zz])
weightgainfin[r]=np.array(valloc)
yr.append(lnchlbl[jj])
r+=1
for jj in range(len(healthycat)):
valloc=list(healthycat[jj])
valloc.append(bmicls[zz])
valloc.append(agecls[zz])
healthycatfin[s]=np.array(valloc)
ys.append(dnrlbl[jj])
s+=1
X_test=np.zeros((len(healthycat)*5,9),dtype=np.float32)
for jj in range(len(healthycat)):
valloc=list(healthycat[jj])
valloc.append(agecl)
valloc.append(clbmi)
X_test[jj]=np.array(valloc)*ti
X_train=healthycatfin# Features
y_train=ys # Labels
#Create a Gaussian Classifier
clf=RandomForestClassifier(n_estimators=100)
#Train the model using the training sets y_pred=clf.predict(X_test)
clf.fit(X_train,y_train)
X_test2=X_test
y_pred=clf.predict(X_test)
print ('SUGGESTED FOOD ITEMS ::')
for ii in range(len(y_pred)):
if y_pred[ii]==2:
print (Food_itemsdata[ii])
findata=Food_itemsdata[ii]
if int(veg)==1:
datanv=['Chicken Burger']
print('\n Thank You for taking our recommendations. :)')
if __name__ == '__main__':
main_win = Tk()
Label(main_win,text="Age").grid(row=0,column=0,sticky=W,pady=4)
Label(main_win,text="veg/Non veg (1/0)").grid(row=1,column=0,sticky=W,pady=4)
Label(main_win,text="Weight (in kg)").grid(row=2,column=0,sticky=W,pady=4)
Label(main_win,text="Height (in cm)").grid(row=3,column=0,sticky=W,pady=4)
e1 = Entry(main_win)
e2 = Entry(main_win)
e3 = Entry(main_win)
e4 = Entry(main_win)
e1.grid(row=0, column=1)
e2.grid(row=1, column=1)
e3.grid(row=2, column=1)
e4.grid(row=3, column=1)
Button(main_win,text='Quit',command=main_win.quit).grid(row=5,column=0,sticky=W,pady=4)
Button(main_win,text='Weight Loss',command=Weight_Loss).grid(row=1,column=4,sticky=W,pady=4)
Button(main_win,text='Weight Gain',command=Weight_Gain).grid(row=2,column=4,sticky=W,pady=4)
Button(main_win,text='Healthy',command=Healthy).grid(row=3,column=4,sticky=W,pady=4)
main_win.geometry("400x200")
main_win.wm_title("DIET RECOMMENDATION SYSTEM")
main_win.mainloop()
# +
from tkinter import *
win = Tk()
win.geometry("400x200")
win.wm_title("xcv")
Button(win,text='Quit',command=win.quit).grid(row=5,column=0,sticky=W,pady=4)
win.mainloop()
# -
| Diet Recommendation System.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sympy as sym
import numpy as np
# +
def rotationGlobalX(alpha):
return np.array([[1,0,0],[0,np.cos(alpha),-np.sin(alpha)],[0,np.sin(alpha),np.cos(alpha)]])
def rotationGlobalY(beta):
return np.array([[np.cos(beta),0,np.sin(beta)], [0,1,0],[-np.sin(beta),0,np.cos(beta)]])
def rotationGlobalZ(gamma):
return np.array([[np.cos(gamma),-np.sin(gamma),0],[np.sin(gamma),np.cos(gamma),0],[0,0,1]])
def rotationLocalX(alpha):
return np.array([[1,0,0],[0,np.cos(alpha),np.sin(alpha)],[0,-np.sin(alpha),np.cos(alpha)]])
def rotationLocalY(beta):
return np.array([[np.cos(beta),0,-np.sin(beta)], [0,1,0],[np.sin(beta),0,np.cos(beta)]])
def rotationLocalZ(gamma):
return np.array([[np.cos(gamma),np.sin(gamma),0],[-np.sin(gamma),np.cos(gamma),0],[0,0,1]])
# +
from mpl_toolkits.mplot3d import Axes3D
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib notebook
plt.rcParams['figure.figsize']=10,10
coefs = (1, 3, 15) # Coefficients in a0/c x**2 + a1/c y**2 + a2/c z**2 = 1
# Radii corresponding to the coefficients:
rx, ry, rz = 1/np.sqrt(coefs)
# Set of all spherical angles:
u = np.linspace(0, 2 * np.pi, 30)
v = np.linspace(0, np.pi, 30)
# Cartesian coordinates that correspond to the spherical angles:
# (this is the equation of an ellipsoid):
x = rx * np.outer(np.cos(u), np.sin(v))
y = ry * np.outer(np.sin(u), np.sin(v))
z = rz * np.outer(np.ones_like(u), np.cos(v))
fig = plt.figure(figsize=plt.figaspect(1)) # Square figure
ax = fig.add_subplot(111, projection='3d')
xr = np.reshape(x, (1,-1))
yr = np.reshape(y, (1,-1))
zr = np.reshape(z, (1,-1))
RX = rotationGlobalX(np.pi/3)
RY = rotationGlobalY(np.pi/3)
RZ = rotationGlobalZ(np.pi/3)
Rx = rotationLocalX(np.pi/3)
Ry = rotationLocalY(np.pi/3)
Rz = rotationLocalZ(np.pi/3)
rRotx = RZ@RY@RX@np.vstack((xr,yr,zr))
print(np.shape(rRotx))
# Plot:
ax.plot_surface(np.reshape(rRotx[0,:],(30,30)), np.reshape(rRotx[1,:],(30,30)),
np.reshape(rRotx[2,:],(30,30)), rstride=4, cstride=4, color='b')
# Adjustment of the axes, so that they all have the same span:
max_radius = max(rx, ry, rz)
for axis in 'xyz':
getattr(ax, 'set_{}lim'.format(axis))((-max_radius, max_radius))
plt.show()
# +
coefs = (1, 3, 15) # Coefficients in a0/c x**2 + a1/c y**2 + a2/c z**2 = 1
# Radii corresponding to the coefficients:
rx, ry, rz = 1/np.sqrt(coefs)
# Set of all spherical angles:
u = np.linspace(0, 2 * np.pi, 30)
v = np.linspace(0, np.pi, 30)
# Cartesian coordinates that correspond to the spherical angles:
# (this is the equation of an ellipsoid):
x = rx * np.outer(np.cos(u), np.sin(v))
y = ry * np.outer(np.sin(u), np.sin(v))
z = rz * np.outer(np.ones_like(u), np.cos(v))
fig = plt.figure(figsize=plt.figaspect(1)) # Square figure
ax = fig.add_subplot(111, projection='3d')
xr = np.reshape(x, (1,-1))
yr = np.reshape(y, (1,-1))
zr = np.reshape(z, (1,-1))
RX = rotationGlobalX(np.pi/3)
RY = rotationGlobalY(np.pi/3)
RZ = rotationGlobalZ(np.pi/3)
Rx = rotationLocalX(np.pi/3)
Ry = rotationLocalY(np.pi/3)
Rz = rotationLocalZ(np.pi/3)
rRotx = RY@RX@np.vstack((xr,yr,zr))
print(np.shape(rRotx))
# Plot:
ax.plot_surface(np.reshape(rRotx[0,:],(30,30)), np.reshape(rRotx[1,:],(30,30)),
np.reshape(rRotx[2,:],(30,30)), rstride=4, cstride=4, color='b')
# Adjustment of the axes, so that they all have the same span:
max_radius = max(rx, ry, rz)
for axis in 'xyz':
getattr(ax, 'set_{}lim'.format(axis))((-max_radius, max_radius))
plt.show()
# -
np.sin(np.arccos(0.7))
print(RZ@RY@RX)
import sympy as sym
sym.init_printing()
a,b,g = sym.symbols('alpha, beta, gamma')
RX = sym.Matrix([[1,0,0],[0,sym.cos(a),-sym.sin(a)],[0,sym.sin(a),sym.cos(a)]])
RY = sym.Matrix([[sym.cos(b),0,sym.sin(b)],[0,1,0],[-sym.sin(b),0,sym.cos(b)]])
RZ = sym.Matrix([[sym.cos(g),-sym.sin(g),0],[sym.sin(g),sym.cos(g),0],[0,0,1]])
RX,RY,RZ
R = RZ@RY@RX
R
mm = np.array([2.71, 10.22, 26.52])
lm = np.array([2.92, 10.10, 18.85])
fh = np.array([5.05, 41.90, 15.41])
mc = np.array([8.29, 41.88, 26.52])
ajc = (mm + lm)/2
kjc = (fh + mc)/2
i = np.array([1,0,0])
j = np.array([0,1,0])
k = np.array([0,0,1])
v1 = kjc - ajc
v1 = v1 / np.sqrt(v1[0]**2+v1[1]**2+v1[2]**2)
v2 = (mm-lm) - ((mm-lm)@v1)*v1
v2 = v2/ np.sqrt(v2[0]**2+v2[1]**2+v2[2]**2)
v3 = k - (k@v1)*v1 - (k@v2)*v2
v3 = v3/ np.sqrt(v3[0]**2+v3[1]**2+v3[2]**2)
v1
R = np.array([v1,v2,v3])
RGlobal = R.T
RGlobal
alpha = np.arctan2(RGlobal[2,1],RGlobal[2,2])*180/np.pi
alpha
beta = np.arctan2(-RGlobal[2,0],np.sqrt(RGlobal[2,1]**2+RGlobal[2,2]**2))*180/np.pi
beta
gamma = np.arctan2(RGlobal[1,0],RGlobal[0,0])*180/np.pi
gamma
R2 = np.array([[0, 0.71, 0.7],[0,0.7,-0.71],[-1,0,0]])
R2
alpha = np.arctan2(R[2,1],R[2,2])*180/np.pi
alpha
gamma = np.arctan2(R[1,0],R[0,0])*180/np.pi
gamma
beta = np.arctan2(-R[2,0],np.sqrt(R[2,1]**2+R[2,2]**2))*180/np.pi
beta
R = RY@RZ@RX
R
alpha = np.arctan2(-R2[1,2],R2[1,1])*180/np.pi
alpha
gamma = 0
beta = 90
import sympy as sym
sym.init_printing()
a,b,g = sym.symbols('alpha, beta, gamma')
RX = sym.Matrix([[1,0,0],[0,sym.cos(a), -sym.sin(a)],[0,sym.sin(a), sym.cos(a)]])
RY = sym.Matrix([[sym.cos(b),0, sym.sin(b)],[0,1,0],[-sym.sin(b),0, sym.cos(b)]])
RZ = sym.Matrix([[sym.cos(g), -sym.sin(g), 0],[sym.sin(g), sym.cos(g),0],[0,0,1]])
RX,RY,RZ
RXYZ = RZ*RY*RX
RXYZ
RZXY = RZ*RX*RY
RZXY
| notebooks/elipsoid3DRotMatrix1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/MichaelColcol/CPEN-21A-CPE-1-1/blob/main/Demo1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="UoMyd29wmDOc" outputId="2508dc92-0051-4288-c878-ce5fff80792f"
b = "sally"
print(b)
# + colab={"base_uri": "https://localhost:8080/"} id="FMRd-xRcnHWu" outputId="a24209bd-6390-42b2-c215-adfa002a0db4"
a = 'Sally'
A = 'John'
print(a)
print (A)
# + colab={"base_uri": "https://localhost:8080/"} id="prIlytTVmmeu" outputId="011e7dce-b141-404d-fd35-73121b636440"
b = "sally"
print (type(b))
# + colab={"base_uri": "https://localhost:8080/"} id="z5-CP57ImM22" outputId="7112fb86-ed2c-4a9e-82e9-3d3ac4b9b466"
a, b, c = 0, 1, 2
print(a)
print(b)
print(c)
# + colab={"base_uri": "https://localhost:8080/"} id="CeL_XNosn3P3" outputId="47f9a0f1-3ee2-4389-fd28-662b5bc70e79"
a, b, c = 0, 1, 2
print(a) # This is a program using type function
print(b)
print(c)
# + colab={"base_uri": "https://localhost:8080/"} id="ALkUNxQfmbte" outputId="1c257df6-a288-415d-b98d-db414a154870"
a = float(4)
print(a)
# + colab={"base_uri": "https://localhost:8080/"} id="csZti8qQmzNu" outputId="12f6770d-207b-4526-ecc7-7a16c5dab291"
a = 4.50
print(type(a))
# + colab={"base_uri": "https://localhost:8080/"} id="LqNNn1Snih1Y" outputId="51a314ba-c67f-4f08-e53b-0f9d993dcdb4"
a,b,c, = 0,1,2
print(type(a))
# + colab={"base_uri": "https://localhost:8080/"} id="ktzZDWwDjHX2" outputId="2cad6d80-ace2-41df-b8b9-87c22228e52e"
x = y = z="four"
print(x)
print(y)
print(z)
# + colab={"base_uri": "https://localhost:8080/"} id="n-O_6QnJjVoO" outputId="b96ea881-d853-408b-b334-d9278d3d5b75"
x= "enjoying"
print('Python programming is '+ x)
# + colab={"base_uri": "https://localhost:8080/"} id="9CEjiDrdjYaG" outputId="d4f1bdba-5d5c-450b-acca-483fcffa4566"
x = 4
y = 5
print(x+y)
print(x-y)
# + colab={"base_uri": "https://localhost:8080/"} id="WiTeKY7IpqPP" outputId="87d415e5-007f-4160-9694-1372cce731ca"
x<y and x==x
# + colab={"base_uri": "https://localhost:8080/"} id="zhgfFxaNp4pg" outputId="1b7cbdf8-6ab8-4235-a4e2-9cd0ba14ca1f"
x>y or x==x
# + colab={"base_uri": "https://localhost:8080/"} id="2fr_iaZip-aQ" outputId="31833a91-a923-46b6-a9a7-c9ce75891078"
not(x>y or x==x)
| Demo1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import requests
import numpy as np
import pandas as pd
from pandas.io.json import json_normalize
url = "https://www.fema.gov/api/open/v1/DisasterDeclarationsSummaries?$top=10&$orderby=disasterNumber&$format=json"
apiResponse = requests.get(url)
responseJson = apiResponse.json()
print(responseJson)
# +
import requests
import numpy as np
import pandas as pd
from pandas.io.json import json_normalize
url = "https://www.fema.gov/api/open/v1/DisasterDeclarationsSummaries?$top=10&$orderby=disasterNumber&$format=json"
apiResponse = requests.get(url)
responseJson = apiResponse.json()
df = json_normalize(responseJson, 'DisasterDeclarationsSummaries')
print(df)
# +
import requests
import numpy as np
import pandas as pd
from pandas.io.json import json_normalize
url = "https://www.fema.gov/api/open/v1/DisasterDeclarationsSummaries?$top=10&$orderby=disasterNumber&$format=json"
url2 = "https://www.fema.gov/api/open/v1/IndividualAssistanceHousingRegistrantsLargeDisasters?$top=10&$orderby=disasterNumber&$format=json"
apiResponse = requests.get(url)
apiResponse2 = requests.get(url2)
responseJson = apiResponse.json()
responseJson2 = apiResponse2.json()
df = json_normalize(responseJson, 'DisasterDeclarationsSummaries')
df2 = json_normalize(responseJson2, 'IndividualAssistanceHousingRegistrantsLargeDisasters')
print(df2)
# +
import requests
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from pandas.io.json import json_normalize
url = "https://www.fema.gov/api/open/v1/DisasterDeclarationsSummaries?$top=1000&$orderby=disasterNumber&$format=json"
url2 = "https://www.fema.gov/api/open/v1/IndividualAssistanceHousingRegistrantsLargeDisasters?$top=1000&$orderby=disasterNumber&$format=json"
apiResponse = requests.get(url)
apiResponse2 = requests.get(url2)
responseJson = apiResponse.json()
responseJson2 = apiResponse2.json()
df = json_normalize(responseJson, 'DisasterDeclarationsSummaries')
df2 = json_normalize(responseJson2, 'IndividualAssistanceHousingRegistrantsLargeDisasters')
frames = [df, df2]
pd.concat(frames, join='outer', sort=False)
# +
import requests
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from pandas.io.json import json_normalize
url = "https://www.fema.gov/api/open/v1/DisasterDeclarationsSummaries?$top=1000&$orderby=disasterNumber&$format=json"
url2 = "https://www.fema.gov/api/open/v1/IndividualAssistanceHousingRegistrantsLargeDisasters?$top=1000&$orderby=disasterNumber&$format=json"
apiResponse = requests.get(url)
apiResponse2 = requests.get(url2)
responseJson = apiResponse.json()
responseJson2 = apiResponse2.json()
df = json_normalize(responseJson, 'DisasterDeclarationsSummaries')
df2 = json_normalize(responseJson2, 'IndividualAssistanceHousingRegistrantsLargeDisasters')
frames = [df, df2]
dfcat = pd.concat(frames, join='outer')
df = df.cumsum()
plt.figure()
dfcat.plot()
# -
| openfema.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Introduction
#
# This notebook shows how to plot an XRD plot for the two polymorphs of CsCl ($Pm\overline{3}m$ and $Fm\overline{3}m$). You can also use matgenie.py's diffraction command to plot an XRD pattern from a structure file.
# +
# Set up some imports that we will need
from pymatgen import Lattice, Structure
from pymatgen.analysis.diffraction.xrd import XRDCalculator
from IPython.display import Image, display
# %matplotlib inline
# -
# # $\alpha$-CsCl ($Pm\overline{3}m$)
#
# Let's start with the typical $\alpha$ form of CsCl.
# +
# Create CsCl structure
a = 4.209 #Angstrom
latt = Lattice.cubic(a)
structure = Structure(latt, ["Cs", "Cl"], [[0, 0, 0], [0.5, 0.5, 0.5]])
c = XRDCalculator()
c.show_xrd_plot(structure)
# -
# Compare it with the experimental XRD pattern below.
display(Image(filename=('./PDF - alpha CsCl.png')))
# # $\beta$-CsCl ($Fm\overline{3}m$)
#
# Let's now look at the $\beta$ (high-temperature) form of CsCl.
# +
# Create CsCl structure
a = 6.923 #Angstrom
latt = Lattice.cubic(a)
structure = Structure(latt, ["Cs", "Cs", "Cs", "Cs", "Cl", "Cl", "Cl", "Cl"],
[[0, 0, 0], [0.5, 0.5, 0], [0, 0.5, 0.5], [0.5, 0, 0.5],
[0.5, 0.5, 0.5], [0, 0, 0.5], [0, 0.5, 0], [0.5, 0, 0]])
c.show_xrd_plot(structure)
# -
# Compare it with the experimental XRD pattern below.
display(Image(filename=('./PDF - beta CsCl.png')))
| examples/Calculating XRD patterns.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import gym
import math
import random
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
from collections import namedtuple
from itertools import count
from PIL import Image
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
import torchvision.transforms as T
# -
import gym
env = gym.make("CartPole-v1")
observation = env.reset()
for _ in range(1000):
env.render()
action = env.action_space.sample() # your agent here (this takes random actions)
observation, reward, done, info = env.step(action)
if done:
observation = env.reset()
env.close()
env = gym.make("CartPole-v1")
# # Hill Climb Test
import gym
import numpy as np
import matplotlib.pyplot as plt
from gym import wrappers
def run_episode(env, parameters):
observation = env.reset()
totalreward = 0
counter = 0
for _ in range(200):
env.render()
action = 0 if np.matmul(parameters, observation) < 0 else 1
observation, reward, done, info = env.step(action)
totalreward += reward
counter+= 1
if done:
break
return totalreward
# +
def train(submit):
env = gym.make('CartPole-v0')
if submit:
env = wrappers.Monitor(env, '/tmp/CartPole-v0-hill-climbing', None, True)
episodes_per_update = 5
noise_scaling = 0.1
parameters = np.random.rand(4) * 2 - 1 # random weights between [-1, 1]
bestreward = 0
counter = 0
for episode in range(2000):
counter += 1
newparams = parameters + (np.random.rand(4) * 2 - 1) * noise_scaling
print(episode)
reward = run_episode(env, newparams)
if reward > bestreward:
bestreward = reward
parameters = newparams
if reward == 200:
print('Yay')
break
return counter
# + jupyter={"outputs_hidden": true}
train(True)
# -
# Because it's hill climbing, not surprised it sucks. Your parameters are set
# # Q-Learning
# I follow this: https://dev.to/n1try/cartpole-with-q-learning---first-experiences-with-openai-gym
#
# Q-learning makes a Q-table with discrete actions and state pairs. Since the observation_space is a 4 tuple of floats, we will need to discretize it. But how mnay states should we discretize it to?
#
# Goal: Stay alive for 200 time steps
#
# Well, we take out x and x' because the cart probably won't leave the screen in 200 time steps.
#
# Now we are only left with theta(angle) and theta' (angle velocity) to worry about. Theta is [-0.42, .42] while theta' is [-3.4*10<sup>38</sup>, 3.4*10<sup>38</sup>]
#
# Q-learning uses one function to fetch the best action from the q-table and another function to update the q-table based on the last action. Rewards are 1 for every time step alive.
#
# Interestingly, the hyperparameters: alpha (learning rate), epsilon (exploration rate) and gamma (discount factor) are interesting to choose.
import gym
import numpy as np
import matplotlib.pyplot as plt
from gym import wrappers
from gym import ObservationWrapper
from gym import spaces
import math
# Helper code to discretize observation space:
#
# Copied from:
# https://github.com/ngc92/space-wrappers/blob/master/space_wrappers/observation_wrappers.py
from space_wrappers import observation_wrappers as ow
# Q-learning algorithm following pseudocode from: https://towardsdatascience.com/introduction-to-various-reinforcement-learning-algorithms-i-q-learning-sarsa-dqn-ddpg-72a5e0cb6287
# and mainly this dude's: https://dev.to/n1try/cartpole-with-q-learning---first-experiences-with-openai-gym
#
# Here's his github: https://gist.github.com/n1try/af0b8476ae4106ec098fea1dfe57f578 <br>
# Here's the reasoning he followed: https://medium.com/@tuzzer/cart-pole-balancing-with-q-learning-b54c6068d947
def Qlearning():
discount = 1.0 # You don't want to discount since your goal is to survive as long as possible
num_episodes = 1000
buckets=(1, 1, 6, 12,)
def discretize(obs):
upper_bounds = [env.observation_space.high[0], 0.5, env.observation_space.high[2], math.radians(50)]
lower_bounds = [env.observation_space.low[0], -0.5, env.observation_space.low[2], -math.radians(50)]
ratios = [(obs[i] + abs(lower_bounds[i])) / (upper_bounds[i] - lower_bounds[i]) for i in range(len(obs))]
new_obs = [int(round((buckets[i] - 1) * ratios[i])) for i in range(len(obs))]
new_obs = [min(buckets[i] - 1, max(0, new_obs[i])) for i in range(len(obs))]
return tuple(new_obs)
env = gym.make('CartPole-v0')
# Initialize a Q-table
num_actions = 2
qtable = np.zeros(buckets + (num_actions,))
# Loop for every episode
for ep in range(num_episodes):
# Optimized epsilon
epsilon = max(0.1, min(1, 1.0 - math.log10((ep + 1) / 25)))
alpha = max(0.1, min(1.0, 1.0 - math.log10((ep + 1) / 25)))
state = discretize(env.reset())
done = False
score = 0
# Loop for each step of episode
while not done:
if ep % 100 == 0:
env.render()
# Select action using epsilon Greedy policy: Either folo policy or pick a random action
action = np.random.choice([np.argmax(qtable[state]), env.action_space.sample()], 1, p=[1-epsilon, epsilon])[0]
# Do the new action
observation, reward, done, info = env.step(action)
new_state = discretize(observation)
# Update Q Table
qtable[state][action] = qtable[state][action] + alpha * (reward + discount * np.max(qtable[new_state]) - qtable[state][action])
score += reward
state = new_state
print("Episode {}, Score: {}".format(ep, score))
env.close()
Qlearning()
# +
# don't forget to do plots of the logistics and such
| CartPole/.ipynb_checkpoints/Q-learning-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# # read_excel
import pandas as pd
df = pd.read_excel('./data/usa_email_sample_db.xlsx')
df
# # Read tab separated
import pandas as pd
df = pd.read_csv('./data/usa_email_sample_db.txt', sep="\t")
df
# # Read csv (with commas in the column)
# - Try opening this in excel (everything will be read in one column)
# - This will load fine with this data
import pandas as pd
df = pd.read_csv('./data/usa_email_sample_db.csv')
df
# Read from http, https, ftp, S3
import pandas as pd
df = pd.read_csv('https://vincentarelbundock.github.io/Rdatasets/csv/boot/acme.csv')
df
# # Read parquet
# ! pip3 install python-snappy==0.5.4 --user # a requisite for snappy compression with fastparquet
df.to_parquet('./data/usa_email_sample_db.parquet')
df = pd.read_parquet('./data/usa_email_sample_db.parquet')
df
# # Read from clipboard
import pandas as pd
df = pd.read_clipboard(sep='\s+') # ‘s+’ denotes one or more whitespace characters.
df
# # Read Pickled objects
# - Python pickle module is used for serializing and de-serializing a Python object structure. ... Pickling is a way to convert a python object (list, dict, etc.) into a character stream. The idea is that this character stream contains all the information necessary to reconstruct the object in another python script.
import pandas as pd
df = pd.DataFrame({"foo": range(5), "bar": range(5, 10)})
pd.to_pickle(df, "./data/dummy.pkl")
unpickled_df = pd.read_pickle("./data/dummy.pkl")
unpickled_df
# # Read json
# - Orient parameter is key [Read more](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.read_json.html)
import pandas as pd
df = pd.read_json('./data/test.json', orient='records')
df
# # Read clipboard
import pandas as pd
df = pd.read_csv('./data/error_tab_line.txt', sep="\t")
df
# # How do we address load issues
# - Get a delimiter that is not part of the string in any of the columns (psv is one extensively used by db guys)
# - Manually Clean up the file before reading (How if there are millions of rows?)
# - Use parameter "error_bad_lines" and ignore the bad lines and move on (Clean it up later)
import pandas as pd
df = pd.read_csv('./data/error_tab_line.txt', sep="\t", error_bad_lines=False)
df
# # For relational db, see relational_database_basic notebook
| class1_explore/load_various_formats.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Where is the tracer on the shelf?
#
# This notebook explores the effects of changing the vertical diffusivity (constant, 3D), changing the isopycnal diffusivity in GMREDI and having a canyon vs a flat shelf on the distribution of tracer over the shelf.
# +
#import gsw as sw # Gibbs seawater package
from math import *
import matplotlib.pyplot as plt
import matplotlib.colors as mcolors
# %matplotlib inline
from MITgcmutils import rdmds
from netCDF4 import Dataset
import numpy as np
import os
import pylab as pl
import scipy.io
import scipy as spy
import seaborn as sns
import sys
# +
lib_path = os.path.abspath('../../Building_canyon/BuildCanyon/PythonModulesMITgcm') # Add absolute path to my python scripts
sys.path.append(lib_path)
import ReadOutTools_MITgcm as rout
# -
sns.set()
sns.set_style('darkgrid')
sns.set_context('notebook')
# +
#Varying-K_iso runs:
CanyonGrid='/ocean/kramosmu/MITgcm/CanyonUpwelling/360x360x90_8Tr_LinProfiles_BarkleyHyd_GMREDI/run13/grid.glob.nc'
CanyonGridOut = Dataset(CanyonGrid)
CNTrun13 = '/ocean/kramosmu/MITgcm/CanyonUpwelling/360x360x90_8Tr_LinProfiles_BarkleyHyd_GMREDI/run13/state.0000000000.glob.nc'
StateOut13 = Dataset(CNTrun13)
CNTrun12Tr = '/ocean/kramosmu/MITgcm/CanyonUpwelling/360x360x90_8Tr_LinProfiles_BarkleyHyd_GMREDI/run12/ptracers.0000000000.glob.nc'
CNTrun13Tr = '/ocean/kramosmu/MITgcm/CanyonUpwelling/360x360x90_8Tr_LinProfiles_BarkleyHyd_GMREDI/run13/ptracers.0000000000.glob.nc'
CNTrun14Tr = '/ocean/kramosmu/MITgcm/CanyonUpwelling/360x360x90_8Tr_LinProfiles_BarkleyHyd_GMREDI/run14/ptracers.0000000000.glob.nc'
CNTrun19Tr = '/ocean/kramosmu/MITgcm/CanyonUpwelling/360x360x90_8Tr_LinProfiles_BarkleyHyd_GMREDI/run19/ptracersGlob.nc'
#Varying-K_v 3D runs
Kv3Drun01Tr = '/ocean/kramosmu/MITgcm/CanyonUpwelling/360x360x90_3Tr_LinProfiles_BarkleyHyd_3DdiffKz/run01/ptracersGlob.nc'
Kv3Drun02Tr = '/ocean/kramosmu/MITgcm/CanyonUpwelling/360x360x90_3Tr_LinProfiles_BarkleyHyd_3DdiffKz/run02/ptracersGlob.nc'
Kv3Drun03Tr = '/ocean/kramosmu/MITgcm/CanyonUpwelling/360x360x90_3Tr_LinProfiles_BarkleyHyd_3DdiffKz/run03/ptracersGlob.nc'
#No Canyon run
NoCrun17Tr = '/ocean/kramosmu/MITgcm/CanyonUpwelling/360x360x90_8Tr_LinProfiles_BarkleyHyd_GMREDI/run17/ptracersGlob.nc'
NoCGrid = '/ocean/kramosmu/MITgcm/CanyonUpwelling/360x360x90_8Tr_LinProfiles_BarkleyHyd_GMREDI/run17/gridGlob.nc'
NoCGridOut = Dataset(NoCGrid)
#No GMREDI runs
NoREDI02Tr = '/ocean/kramosmu/MITgcm/CanyonUpwelling/360x360x90_3Tr_Linprofiles_BarkleyHyd/run02/ptracersGlob.nc'
NoREDI03Tr = '/ocean/kramosmu/MITgcm/CanyonUpwelling/360x360x90_3Tr_Linprofiles_BarkleyHyd/run03/ptracersGlob.nc'
NoREDINoCTr = '/ocean/kramosmu/MITgcm/CanyonUpwelling/360x360x90_3Tr_Linprofiles_BarkleyHyd/run04/ptracersGlob.nc'
# +
# General input
nx = 360
ny = 360
nz = 90
nt = 19 # t dimension size
z = StateOut13.variables['Z']
#print(z[10])
Time = StateOut13.variables['T']
#print(Time[:])
xc = rout.getField(CanyonGrid, 'XC') # x coords tracer cells
yc = rout.getField(CanyonGrid, 'YC') # y coords tracer cells
drF = CanyonGridOut.variables['drF'] # vertical distance between faces
dxG = rout.getField(CanyonGrid,'dxG')
bathy = rout.getField(CanyonGrid, 'Depth')
rA = rout.getField(CanyonGrid, 'rA') # area of cells (x-y)
hFacC = rout.getField(CanyonGrid, 'HFacC')
MaskC = rout.getMask(CanyonGrid,'HFacC') # same for both runs
MaskNoC = rout.getMask(NoCGrid,'HFacC')
hFacCNoC = rout.getField(NoCGrid,'HFacC')
rANoC = rout.getField(NoCGrid,'rA')
drFNoC= NoCGridOut.variables['drF']
# +
# Load tracers variable K_iso
Tr1Iso100 = rout.getField(CNTrun14Tr,'Tr1') # Tracer 1 CNT run19 , Kz = E-5
Tr2Iso100 = rout.getField(CNTrun14Tr,'Tr3') # Tracer 3 CNT run19 , Kz = E-3
Tr1Iso10 = rout.getField(CNTrun12Tr,'Tr1') # Tracer 1 CNT run12 , Kz = E-5
Tr2Iso10 = rout.getField(CNTrun12Tr,'Tr2') # Tracer 2 CNT run12 , Kz = E-3
Tr1Iso1 = rout.getField(CNTrun13Tr,'Tr1') # Tracer 1 CNT run13 , Kz = E-5
Tr2Iso1 = rout.getField(CNTrun13Tr,'Tr2') # Tracer 2 CNT run13 , Kz = E-3
Tr1Iso01 = rout.getField(CNTrun14Tr,'Tr1') # Tracer 1 CNT run14 , Kz = E-5
Tr2Iso01 = rout.getField(CNTrun14Tr,'Tr2') # Tracer 2 CNT run14 , Kz = E-3
# -
# Load tracers variable K_v
Tr13D = rout.getField(Kv3Drun01Tr,'Tr1') # Tracer 1 3D run01 , Kz = E-7 out, E-3 in
Tr23D = rout.getField(Kv3Drun02Tr,'Tr1') # Tracer 1 3D run02 , Kz = E-7 out, E-4 in
Tr33D = rout.getField(Kv3Drun03Tr,'Tr1') # Tracer 1 3D run03 , Kz = E-5 out, E-3 in
# Load tracers of no canyon run
Tr1NoC = rout.getField(NoCrun17Tr,'Tr1') # Tracer 1 NoC run17CNT , Kz = E-5
Tr2NoC = rout.getField(NoCrun17Tr,'Tr2') # Tracer 2 NoC run17CNT , Kz = E-3
# +
# Load tracers of no REDI runs
Tr1NoREDI02 = rout.getField(NoREDI02Tr,'Tr1') # Tracer 1 NoREDI run02 , Kz = E-5
Tr2NoREDI02 = rout.getField(NoREDI02Tr,'Tr2') # Tracer 2 NoREDI run02 , Kz = E-4
Tr3NoREDI02 = rout.getField(NoREDI02Tr,'Tr3') # Tracer 3 NoREDI run02 , Kz = E-3
Tr1NoREDI03 = rout.getField(NoREDI03Tr,'Tr1') # Tracer 1 NoREDI run03 , Kz = E-5
Tr2NoREDI03 = rout.getField(NoREDI03Tr,'Tr2') # Tracer 2 NoREDI run03 , Kz = E-4
Tr3NoREDI03 = rout.getField(NoREDI03Tr,'Tr3') # Tracer 3 NoREDI run03 , Kz = E-3
Tr1NoREDINoC = rout.getField(NoREDINoCTr,'Tr1') # Tracer 1 NoREDI run04 , Kz = E-5
Tr2NoREDINoC = rout.getField(NoREDINoCTr,'Tr2') # Tracer 2 NoREDI run04 , Kz = E-4
Tr3NoREDINoC = rout.getField(NoREDINoCTr,'Tr3') # Tracer 3 NoREDI run04 , Kz = E-3
# -
# ### How much water with concentration higher than a limit is there along the shelf? How much tracer mass along the shelf?
def HowMuchWaterX(Tr,MaskC,nzlim,rA,hFacC,drF,tt,nx,dx):
'''
INPUT----------------------------------------------------------------------------------------------------------------
Tr : Array with concentration values for a tracer. Until this function is more general, this should be size 19x90x360x360
MaskC : Land mask for tracer
nzlim : The nz index under which to look for water properties
rA : Area of cell faces at C points (360x360)
fFacC : Fraction of open cell (90x360x360)
drF : Distance between cell faces (90)
tt : Time slice to calculate. Int 0<=tt<19
nx : x dimension (along shelf)
dx :
OUTPUT----------------------------------------------------------------------------------------------------------------
WaterX = (360) Arrays with the volume of water at each x-position over the shelf [tt,:28,:197,xx]
TrX = (360) Arrays with the mass of tracer (umol) at each x-position over the shelf [tt,:28,:197,xx].
Total mass of tracer at xx on the shelf.
-----------------------------------------------------------------------------------------------------------------------
'''
WaterX= np.zeros(nx)
TrX= np.zeros(nx)
TrMask0=np.ma.array(Tr[0,:,:,:],mask=MaskC[:,:,:])
trlim = TrMask0[nzlim,50,180]
hFacCSwap = np.swapaxes(hFacC, 0, 2)
#print('tracer limit is: ',trlim)
TrMask=np.ma.array(Tr[tt,:,:,:],mask=MaskC[:,:,:])
for ii,trac in np.ndenumerate(TrMask[:28,197:,:]) :
if trac >= trlim:
WaterX[ii[2]] = WaterX[ii[2]] + hFacC[ii]*drF[ii[0]]*rA[ii[1],ii[2]]/dx[ii[1],ii[2]]
VolX = (np.swapaxes(hFacCSwap[:,197:,:28]*drF[:28],0,2))*rA[197:,:]
TrX[:] = np.sum(np.sum((VolX*TrMask[:28,197:,:]*1000.0),axis=0),axis=0)/dx[0,:] #[1 umol/l=1000 umol/m^3]
return(WaterX,TrX)
# ### Case 1: Changing $K_{iso}$ in GMREDI
# +
fig45=plt.figure(figsize=(18,12))
sns.set(context='paper', style='whitegrid', font='sans-serif', font_scale=1.3, rc={"lines.linewidth": 1.5})
time = 6
(WaterXIso100, Tr1XIso100) = HowMuchWaterX(Tr1Iso100,MaskNoC,30,rA,hFacCNoC,drFNoC,time,nx,dxG)
(WaterXIso10, Tr1XIso10) = HowMuchWaterX(Tr1Iso10,MaskNoC,30,rA,hFacCNoC,drFNoC,time,nx,dxG)
(WaterXIso1, Tr1XIso1) = HowMuchWaterX(Tr1Iso1,MaskNoC,30,rA,hFacCNoC,drFNoC,time,nx,dxG)
(WaterXIso01, Tr1XIso01) = HowMuchWaterX(Tr1Iso01,MaskNoC,30,rANoC,hFacCNoC,drFNoC,time,nx,dxG)
(WaterXNoREDI02, Tr1XNoREDI02) = HowMuchWaterX(Tr1NoREDI02,MaskNoC,30,rANoC,hFacCNoC,drFNoC,time,nx,dxG)
(WaterXNoREDI03, Tr1XNoREDI03) = HowMuchWaterX(Tr1NoREDI03,MaskNoC,30,rANoC,hFacCNoC,drFNoC,time,nx,dxG)
ax1 = plt.subplot(2,3,1)
ax1.plot(xc[0,:],(WaterXIso100)*1000.0,'-',label=('$k_{Iso}=100m^2s^{-1}$,$k_{v}=10^{-5}m^2s^{-1}$ day %d' %(time/2.0))) # 1000m/km
ax1.plot(xc[0,:],(WaterXIso10)*1000.0,'-',label=('$k_{Iso}=10m^2s^{-1}$ ')) # 1000m/km
ax1.plot(xc[0,:],(WaterXIso1)*1000.0,'-',label=('$k_{Iso}=1m^2s^{-1}$'))
ax1.plot(xc[0,:],(WaterXIso01)*1000.0,'-',label=('$k_{Iso}=0.1m^2s^{-1}$ '))
ax1.plot(xc[0,:],(WaterXNoREDI02)*1000.0,'-',label=('$k_{h}=10^{-5}m^2s^{-1}$ '))
ax1.plot(xc[0,:],(WaterXNoREDI03)*1000.0,'-',label=('$k_{h}=10^{-7}m^2s^{-1}$ '))
plt.ylabel('Water over shelf C > 7.40 $umol$ $l^{-1}$ ($m^3 km^{-1}$)')
plt.xlabel('Along-shore distance ($km$)')
labels = [10,20,30,40, 50, 60, 70, 80,90,100,110,120]
plt.xticks([10000,20000,30000,40000,50000,60000,70000,80000,90000,100000,110000,120000], labels)
#ax2.title = ('day %d' %(time/2.0))
plt.legend(loc=2)
ax4 = plt.subplot(2,3,4)
ax4.plot(xc[0,:],(Tr1XIso100)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1XIso100)*dxG[0,:]*1.E-6)))) # 1000m/km
ax4.plot(xc[0,:],(Tr1XIso10)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1XIso10)*dxG[0,:]*1.E-6)))) # 1000m/km
ax4.plot(xc[0,:],(Tr1XIso1)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1XIso1)*dxG[0,:]*1.E-6))))
ax4.plot(xc[0,:],(Tr1XIso01)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1XIso01)*dxG[0,:]*1.E-6))))
ax4.plot(xc[0,:],(Tr1XNoREDI02)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1XNoREDI02)*dxG[0,:]*1.E-6))))
ax4.plot(xc[0,:],(Tr1XNoREDI03)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1XNoREDI03)*dxG[0,:]*1.E-6))))
plt.ylabel('Tracer mass per km over shelf ($mol$ $km^{-1}$)')
plt.xlabel('Along-shore distance ($km$)')
labels = [10,20,30,40, 50, 60, 70, 80,90,100,110,120]
plt.xticks([10000,20000,30000,40000,50000,60000,70000,80000,90000,100000,110000,120000], labels)
#ax2.title = ('day %d' %(time/2.0))
plt.legend(loc=0)
time = 10
(WaterXIso100, Tr1XIso100) = HowMuchWaterX(Tr1Iso100,MaskNoC,30,rA,hFacCNoC,drFNoC,time,nx,dxG)
(WaterXIso10, Tr1XIso10) = HowMuchWaterX(Tr1Iso10,MaskNoC,30,rA,hFacCNoC,drFNoC,time,nx,dxG)
(WaterXIso1, Tr1XIso1) = HowMuchWaterX(Tr1Iso1,MaskNoC,30,rA,hFacCNoC,drFNoC,time,nx,dxG)
(WaterXIso01, Tr1XIso01) = HowMuchWaterX(Tr1Iso01,MaskNoC,30,rANoC,hFacCNoC,drFNoC,time,nx,dxG)
(WaterXNoREDI02, Tr1XNoREDI02) = HowMuchWaterX(Tr1NoREDI02,MaskNoC,30,rANoC,hFacCNoC,drFNoC,time,nx,dxG)
(WaterXNoREDI03, Tr1XNoREDI03) = HowMuchWaterX(Tr1NoREDI03,MaskNoC,30,rANoC,hFacCNoC,drFNoC,time,nx,dxG)
ax2 = plt.subplot(2,3,2)
ax2.plot(xc[0,:],(WaterXIso100)*1000.0,'-',label=('$k_{Iso}=100m^2s^{-1}$,$k_{v}=10^{-5}m^2s^{-1}$ day %d' %(time/2.0))) # 1000m/km
ax2.plot(xc[0,:],(WaterXIso10)*1000.0,'-',label=('$k_{Iso}=10m^2s^{-1}$')) # 1000m/km
ax2.plot(xc[0,:],(WaterXIso1)*1000.0,'-',label=('$k_{Iso}=1m^2s^{-1}$'))
ax2.plot(xc[0,:],(WaterXIso01)*1000.0,'-',label=('$k_{Iso}=0.1m^2s^{-1}$'))
ax2.plot(xc[0,:],(WaterXNoREDI02)*1000.0,'-',label=('$k_{h}=10^{-5}m^2s^{-1}$ '))
ax2.plot(xc[0,:],(WaterXNoREDI03)*1000.0,'-',label=('$k_{h}=10^{-7}m^2s^{-1}$ '))
plt.ylabel('Water over shelf C > 7.40 $umol$ $l^{-1}$ ($m^3 km^{-1}$)')
plt.xlabel('Along-shore distance ($km$)')
labels = [10,20,30,40, 50, 60, 70, 80,90,100,110,120]
plt.xticks([10000,20000,30000,40000,50000,60000,70000,80000,90000,100000,110000,120000], labels)
#plt.title = ('day %d' %(time/2.0))
plt.legend(loc=2)
ax5 = plt.subplot(2,3,5)
ax5.plot(xc[0,:],(Tr1XIso100)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1XIso100)*dxG[0,:]*1.E-6)))) # 1000m/km
ax5.plot(xc[0,:],(Tr1XIso10)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1XIso10)*dxG[0,:]*1.E-6)))) # 1000m/km
ax5.plot(xc[0,:],(Tr1XIso1)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1XIso1)*dxG[0,:]*1.E-6))))
ax5.plot(xc[0,:],(Tr1XIso01)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1XIso01)*dxG[0,:]*1.E-6))))
ax5.plot(xc[0,:],(Tr1XNoREDI02)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1XNoREDI02)*dxG[0,:]*1.E-6))))
ax5.plot(xc[0,:],(Tr1XNoREDI03)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1XNoREDI03)*dxG[0,:]*1.E-6))))
plt.ylabel('Tracer mass per km over shelf ($mol$ $km^{-1}$)')
plt.xlabel('Along-shore distance ($km$)')
labels = [10,20,30,40, 50, 60, 70, 80,90,100,110,120]
plt.xticks([10000,20000,30000,40000,50000,60000,70000,80000,90000,100000,110000,120000], labels)
#ax2.title = ('day %d' %(time/2.0))
plt.legend(loc=0)
time = 16
(WaterXIso100, Tr1XIso100) = HowMuchWaterX(Tr1Iso100,MaskNoC,30,rA,hFacCNoC,drFNoC,time,nx,dxG)
(WaterXIso10, Tr1XIso10) = HowMuchWaterX(Tr1Iso10,MaskNoC,30,rA,hFacCNoC,drFNoC,time,nx,dxG)
(WaterXIso1, Tr1XIso1) = HowMuchWaterX(Tr1Iso1,MaskNoC,30,rA,hFacCNoC,drFNoC,time,nx,dxG)
(WaterXIso01, Tr1XIso01) = HowMuchWaterX(Tr1Iso01,MaskNoC,30,rANoC,hFacCNoC,drFNoC,time,nx,dxG)
(WaterXNoREDI02, Tr1XNoREDI02) = HowMuchWaterX(Tr1NoREDI02,MaskNoC,30,rANoC,hFacCNoC,drFNoC,time,nx,dxG)
(WaterXNoREDI03, Tr1XNoREDI03) = HowMuchWaterX(Tr1NoREDI03,MaskNoC,30,rANoC,hFacCNoC,drFNoC,time,nx,dxG)
ax3 = plt.subplot(2,3,3)
ax3.plot(xc[0,:],(WaterXIso100)*1000.0,'-',label=('$k_{Iso}=100m^2s^{-1}$,$k_{v}=10^{-5}m^2s^{-1}$ day %d' %(time/2.0))) # 1000m/km
ax3.plot(xc[0,:],(WaterXIso10)*1000.0,'-',label=('$k_{Iso}=10m^2s^{-1}$')) # 1000m/km
ax3.plot(xc[0,:],(WaterXIso1)*1000.0,'-',label=('$k_{Iso}=1m^2s^{-1}$'))
ax3.plot(xc[0,:],(WaterXIso01)*1000.0,'-',label=('$k_{Iso}=0.1m^2s^{-1}$'))
ax3.plot(xc[0,:],(WaterXNoREDI02)*1000.0,'-',label=('$k_{h}=10^{-5}m^2s^{-1}$ '))
ax3.plot(xc[0,:],(WaterXNoREDI03)*1000.0,'-',label=('$k_{h}=10^{-7}m^2s^{-1}$ '))
plt.ylabel('Water over shelf C > 7.40 $umol$ $l^{-1}$ ($m^3 km^{-1}$)')
plt.xlabel('Along-shore distance ($km$)')
labels = [10,20,30,40, 50, 60, 70, 80,90,100,110,120]
plt.xticks([10000,20000,30000,40000,50000,60000,70000,80000,90000,100000,110000,120000], labels)
#plt.title = ('day %d' %(time/2.0))
plt.legend(loc=2)
ax6 = plt.subplot(2,3,6)
ax6.plot(xc[0,:],(Tr1XIso100)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1XIso100)*dxG[0,:]*1.E-6)))) # 1000m/km
ax6.plot(xc[0,:],(Tr1XIso10)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1XIso10)*dxG[0,:]*1.E-6)))) # 1000m/km
ax6.plot(xc[0,:],(Tr1XIso1)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1XIso1)*dxG[0,:]*1.E-6))))
ax6.plot(xc[0,:],(Tr1XIso01)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1XIso01)*dxG[0,:]*1.E-6))))
ax6.plot(xc[0,:],(Tr1XNoREDI02)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1XNoREDI02)*dxG[0,:]*1.E-6))))
ax6.plot(xc[0,:],(Tr1XNoREDI03)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1XNoREDI03)*dxG[0,:]*1.E-6))))
plt.ylabel('Tracer mass per km over shelf ($mol$ $km^{-1}$)')
plt.xlabel('Along-shore distance ($km$)')
labels = [10,20,30,40, 50, 60, 70, 80,90,100,110,120]
plt.xticks([10000,20000,30000,40000,50000,60000,70000,80000,90000,100000,110000,120000], labels)
#ax2.title = ('day %d' %(time/2.0))
plt.legend(loc=0)
#fig45.savefig('/ocean/kramosmu/Figures/WaterVolumeOverShelf/H20TrPerKm3DCNT1-NoC1.eps', format='eps', dpi=1000,bbox_extra_artists=(leg,), bbox_inches='tight')
# -
# ### Case 2: Enhanced mixing inside the canyon (3D vertical diffusivity)
#
# +
fig45=plt.figure(figsize=(18,12))
sns.set(context='paper', style='whitegrid', font='sans-serif', font_scale=1.3, rc={"lines.linewidth": 1.5})
time = 6
(WaterX3D1, Tr1X3D) = HowMuchWaterX(Tr13D,MaskNoC,30,rA,hFacCNoC,drFNoC,time,nx,dxG)
(WaterX3D2, Tr2X3D) = HowMuchWaterX(Tr23D,MaskNoC,30,rA,hFacCNoC,drFNoC,time,nx,dxG)
(WaterX3D3, Tr3X3D) = HowMuchWaterX(Tr33D,MaskNoC,30,rANoC,hFacCNoC,drFNoC,time,nx,dxG)
(WaterX1Iso1, Tr1XIso1) = HowMuchWaterX(Tr1Iso1,MaskNoC,30,rANoC,hFacCNoC,drFNoC,time,nx,dxG)
ax1 = plt.subplot(2,3,1)
ax1.plot(xc[0,:],(WaterX3D1)*1000.0,'-',label=('Tr1 3D day %d' %(time/2.0))) # 1000m/km
ax1.plot(xc[0,:],(WaterX3D2)*1000.0,'-',label=('Tr2 3D day %d' %(time/2.0)))
ax1.plot(xc[0,:],(WaterX3D3)*1000.0,'-',label=('Tr3 3D day %d' %(time/2.0)))
ax1.plot(xc[0,:],(WaterX1Iso1)*1000.0,'-',label=('CNT day %d' %(time/2.0)))
plt.ylabel('Water over shelf C > 7.40 $umol$ $l^{-1}$ ($m^3 km^{-1}$)')
plt.xlabel('Along-shore distance ($km$)')
labels = [10,20,30,40, 50, 60, 70, 80,90,100,110,120]
plt.xticks([10000,20000,30000,40000,50000,60000,70000,80000,90000,100000,110000,120000], labels)
#ax2.title = ('day %d' %(time/2.0))
plt.legend(loc=0)
ax4 = plt.subplot(2,3,4)
ax4.plot(xc[0,:],(Tr1X3D)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1X3D)*dxG[0,:]*1.E-6)))) # 1000m/km
ax4.plot(xc[0,:],(Tr2X3D)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr2X3D)*dxG[0,:]*1.E-6))))
ax4.plot(xc[0,:],(Tr3X3D)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr3X3D)*dxG[0,:]*1.E-6))))
ax4.plot(xc[0,:],(Tr1XIso1)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1XIso1)*dxG[0,:]*1.E-6))))
plt.ylabel('Tracer mass per km over shelf ($mol$ $km^{-1}$)')
plt.xlabel('Along-shore distance ($km$)')
labels = [10,20,30,40, 50, 60, 70, 80,90,100,110,120]
plt.xticks([10000,20000,30000,40000,50000,60000,70000,80000,90000,100000,110000,120000], labels)
#ax2.title = ('day %d' %(time/2.0))
plt.legend(loc=0)
time = 10
(WaterX3D1, Tr1X3D) = HowMuchWaterX(Tr13D,MaskNoC,30,rA,hFacCNoC,drFNoC,time,nx,dxG)
(WaterX3D2, Tr2X3D) = HowMuchWaterX(Tr23D,MaskNoC,30,rA,hFacCNoC,drFNoC,time,nx,dxG)
(WaterX3D3, Tr3X3D) = HowMuchWaterX(Tr33D,MaskNoC,30,rANoC,hFacCNoC,drFNoC,time,nx,dxG)
(WaterX1Iso1, Tr1XIso1) = HowMuchWaterX(Tr1Iso1,MaskNoC,30,rANoC,hFacCNoC,drFNoC,time,nx,dxG)
ax2 = plt.subplot(2,3,2)
ax2.plot(xc[0,:],(WaterX3D1)*1000.0,'-',label=('Tr1 3D day %d' %(time/2.0))) # 1000m/km
ax2.plot(xc[0,:],(WaterX3D2)*1000.0,'-',label=('Tr2 3D day %d' %(time/2.0)))
ax2.plot(xc[0,:],(WaterX3D3)*1000.0,'-',label=('Tr3 3D day %d' %(time/2.0)))
ax2.plot(xc[0,:],(WaterX1Iso1)*1000.0,'-',label=('CNT day %d' %(time/2.0)))
plt.ylabel('Water over shelf C > 7.40 $umol$ $l^{-1}$ ($m^3 km^{-1}$)')
plt.xlabel('Along-shore distance ($km$)')
labels = [10,20,30,40, 50, 60, 70, 80,90,100,110,120]
plt.xticks([10000,20000,30000,40000,50000,60000,70000,80000,90000,100000,110000,120000], labels)
#plt.title = ('day %d' %(time/2.0))
plt.legend(loc=0)
ax5 = plt.subplot(2,3,5)
ax5.plot(xc[0,:],(Tr1X3D)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1X3D)*dxG[0,:]*1.E-6)))) # 1000m/km
ax5.plot(xc[0,:],(Tr2X3D)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr2X3D)*dxG[0,:]*1.E-6))))
ax5.plot(xc[0,:],(Tr3X3D)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr3X3D)*dxG[0,:]*1.E-6))))
ax5.plot(xc[0,:],(Tr1XIso1)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1XIso1)*dxG[0,:]*1.E-6))))
plt.ylabel('Tracer mass per km over shelf ($mol$ $km^{-1}$)')
plt.xlabel('Along-shore distance ($km$)')
labels = [10,20,30,40, 50, 60, 70, 80,90,100,110,120]
plt.xticks([10000,20000,30000,40000,50000,60000,70000,80000,90000,100000,110000,120000], labels)
#ax2.title = ('day %d' %(time/2.0))
plt.legend(loc=0)
time = 16
(WaterX3D1, Tr1X3D) = HowMuchWaterX(Tr13D,MaskNoC,30,rA,hFacCNoC,drFNoC,time,nx,dxG)
(WaterX3D2, Tr2X3D) = HowMuchWaterX(Tr23D,MaskNoC,30,rA,hFacCNoC,drFNoC,time,nx,dxG)
(WaterX3D3, Tr3X3D) = HowMuchWaterX(Tr33D,MaskNoC,30,rANoC,hFacCNoC,drFNoC,time,nx,dxG)
(WaterX1Iso1, Tr1XIso1) = HowMuchWaterX(Tr1Iso1,MaskNoC,30,rANoC,hFacCNoC,drFNoC,time,nx,dxG)
ax3 = plt.subplot(2,3,3)
ax3.plot(xc[0,:],(WaterX3D1)*1000.0,'-',label=('Tr1 3D day %d' %(time/2.0))) # 1000m/km
ax3.plot(xc[0,:],(WaterX3D2)*1000.0,'-',label=('Tr2 3D day %d' %(time/2.0)))
ax3.plot(xc[0,:],(WaterX3D3)*1000.0,'-',label=('Tr3 3D day %d' %(time/2.0)))
ax3.plot(xc[0,:],(WaterX1Iso1)*1000.0,'-',label=('CNT day %d' %(time/2.0)))
plt.ylabel('Water over shelf C > 7.40 $umol$ $l^{-1}$ ($m^3 km^{-1}$)')
plt.xlabel('Along-shore distance ($km$)')
labels = [10,20,30,40, 50, 60, 70, 80,90,100,110,120]
plt.xticks([10000,20000,30000,40000,50000,60000,70000,80000,90000,100000,110000,120000], labels)
#plt.title = ('day %d' %(time/2.0))
plt.legend(loc=0)
ax6 = plt.subplot(2,3,6)
ax6.plot(xc[0,:],(Tr1X3D)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1X3D)*dxG[0,:]*1.E-6)))) # 1000m/km
ax6.plot(xc[0,:],(Tr2X3D)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr2X3D)*dxG[0,:]*1.E-6))))
ax6.plot(xc[0,:],(Tr3X3D)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr3X3D)*dxG[0,:]*1.E-6))))
ax6.plot(xc[0,:],(Tr1XIso1)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1XIso1)*dxG[0,:]*1.E-6))))
plt.ylabel('Tracer mass per km over shelf ($mol$ $km^{-1}$)')
plt.xlabel('Along-shore distance ($km$)')
labels = [10,20,30,40, 50, 60, 70, 80,90,100,110,120]
plt.xticks([10000,20000,30000,40000,50000,60000,70000,80000,90000,100000,110000,120000], labels)
#ax2.title = ('day %d' %(time/2.0))
plt.legend(loc=0)
#fig45.savefig('/ocean/kramosmu/Figures/WaterVolumeOverShelf/H20TrPerKm3DCNT1-NoC1.eps', format='eps', dpi=1000,bbox_extra_artists=(leg,), bbox_inches='tight')
# -
# ### Case 3: Varying Kv and flat shelf
# +
fig45=plt.figure(figsize=(18,12))
sns.set(context='paper', style='whitegrid', font='sans-serif', font_scale=1.3, rc={"lines.linewidth": 1.5})
time = 6
(WaterX1Iso1, Tr1XIso1) = HowMuchWaterX(Tr1Iso1,MaskNoC,30,rA,hFacCNoC,drFNoC,time,nx,dxG)
(WaterX2Iso1, Tr2XIso1) = HowMuchWaterX(Tr2Iso1,MaskNoC,30,rA,hFacCNoC,drFNoC,time,nx,dxG)
(WaterX1NoC, Tr1XNoC) = HowMuchWaterX(Tr1NoC,MaskNoC,30,rANoC,hFacCNoC,drFNoC,time,nx,dxG)
(WaterX2NoC, Tr2XNoC) = HowMuchWaterX(Tr2NoC,MaskNoC,30,rANoC,hFacCNoC,drFNoC,time,nx,dxG)
ax1 = plt.subplot(2,3,1)
ax1.plot(xc[0,:],(WaterX1Iso1)*1000.0,'-',label=('Tr1 Cny day %d' %(time/2.0))) # 1000m/km
ax1.plot(xc[0,:],(WaterX2Iso1)*1000.0,'-',label=('Tr2 Cny day %d' %(time/2.0)))
ax1.plot(xc[0,:],(WaterX1NoC)*1000.0,'-',label=('Tr1 NoC day %d' %(time/2.0)))
ax1.plot(xc[0,:],(WaterX2NoC)*1000.0,'-',label=('Tr2 NoC %d' %(time/2.0)))
plt.ylabel('Water over shelf C > 7.40 $umol$ $l^{-1}$ ($m^3 km^{-1}$)')
plt.xlabel('Along-shore distance ($km$)')
labels = [10,20,30,40, 50, 60, 70, 80,90,100,110,120]
plt.xticks([10000,20000,30000,40000,50000,60000,70000,80000,90000,100000,110000,120000], labels)
#ax2.title = ('day %d' %(time/2.0))
plt.legend(loc=0)
ax4 = plt.subplot(2,3,4)
ax4.plot(xc[0,:],(Tr1XIso1)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1XIso1)*dxG[0,:]*1.E-6)))) # 1000m/km
ax4.plot(xc[0,:],(Tr2XIso1)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr2XIso1)*dxG[0,:]*1.E-6))))
ax4.plot(xc[0,:],(Tr1XNoC)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1XNoC)*dxG[0,:]*1.E-6))))
ax4.plot(xc[0,:],(Tr2XNoC)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr2XNoC)*dxG[0,:]*1.E-6))))
plt.ylabel('Tracer mass per km over shelf ($mol$ $km^{-1}$)')
plt.xlabel('Along-shore distance ($km$)')
labels = [10,20,30,40, 50, 60, 70, 80,90,100,110,120]
plt.xticks([10000,20000,30000,40000,50000,60000,70000,80000,90000,100000,110000,120000], labels)
#ax2.title = ('day %d' %(time/2.0))
plt.legend(loc=0)
time = 10
(WaterX1Iso1, Tr1XIso1) = HowMuchWaterX(Tr1Iso1,MaskNoC,30,rA,hFacCNoC,drFNoC,time,nx,dxG)
(WaterX2Iso1, Tr2XIso1) = HowMuchWaterX(Tr2Iso1,MaskNoC,30,rA,hFacCNoC,drFNoC,time,nx,dxG)
(WaterX1NoC, Tr1XNoC) = HowMuchWaterX(Tr1NoC,MaskNoC,30,rANoC,hFacCNoC,drFNoC,time,nx,dxG)
(WaterX2NoC, Tr2XNoC) = HowMuchWaterX(Tr2NoC,MaskNoC,30,rANoC,hFacCNoC,drFNoC,time,nx,dxG)
ax2 = plt.subplot(2,3,2)
ax2.plot(xc[0,:],(WaterX1Iso1)*1000.0,'-',label=('Tr1 Cny day %d' %(time/2.0))) # 1000m/km
ax2.plot(xc[0,:],(WaterX2Iso1)*1000.0,'-',label=('Tr2 Cny day %d' %(time/2.0)))
ax2.plot(xc[0,:],(WaterX1NoC)*1000.0,'-',label=('Tr1 NoC day %d' %(time/2.0)))
ax2.plot(xc[0,:],(WaterX2NoC)*1000.0,'-',label=('Tr2 NoC %d' %(time/2.0)))
plt.ylabel('Water over shelf C > 7.40 $umol$ $l^{-1}$ ($m^3 km^{-1}$)')
plt.xlabel('Along-shore distance ($km$)')
labels = [10,20,30,40, 50, 60, 70, 80,90,100,110,120]
plt.xticks([10000,20000,30000,40000,50000,60000,70000,80000,90000,100000,110000,120000], labels)
#plt.title = ('day %d' %(time/2.0))
plt.legend(loc=0)
ax5 = plt.subplot(2,3,5)
ax5.plot(xc[0,:],(Tr1XIso1)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1XIso1)*dxG[0,:]*1.E-6)))) # 1000m/km
ax5.plot(xc[0,:],(Tr2XIso1)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr2XIso1)*dxG[0,:]*1.E-6))))
ax5.plot(xc[0,:],(Tr1XNoC)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1XNoC)*dxG[0,:]*1.E-6))))
ax5.plot(xc[0,:],(Tr2XNoC)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr2XNoC)*dxG[0,:]*1.E-6))))
plt.ylabel('Tracer mass per km over shelf ($mol$ $km^{-1}$)')
plt.xlabel('Along-shore distance ($km$)')
labels = [10,20,30,40, 50, 60, 70, 80,90,100,110,120]
plt.xticks([10000,20000,30000,40000,50000,60000,70000,80000,90000,100000,110000,120000], labels)
#ax2.title = ('day %d' %(time/2.0))
plt.legend(loc=0)
time = 16
(WaterX1Iso1, Tr1XIso1) = HowMuchWaterX(Tr1Iso1,MaskNoC,30,rA,hFacCNoC,drFNoC,time,nx,dxG)
(WaterX2Iso1, Tr2XIso1) = HowMuchWaterX(Tr2Iso1,MaskNoC,30,rA,hFacCNoC,drFNoC,time,nx,dxG)
(WaterX1NoC, Tr1XNoC) = HowMuchWaterX(Tr1NoC,MaskNoC,30,rANoC,hFacCNoC,drFNoC,time,nx,dxG)
(WaterX2NoC, Tr2XNoC) = HowMuchWaterX(Tr2NoC,MaskNoC,30,rANoC,hFacCNoC,drFNoC,time,nx,dxG)
ax3 = plt.subplot(2,3,3)
ax3.plot(xc[0,:],(WaterX1Iso1)*1000.0,'-',label=('Tr1 Cny day %d' %(time/2.0))) # 1000m/km
ax3.plot(xc[0,:],(WaterX2Iso1)*1000.0,'-',label=('Tr2 Cny day %d' %(time/2.0)))
ax3.plot(xc[0,:],(WaterX1NoC)*1000.0,'-',label=('Tr1 NoC day %d' %(time/2.0)))
ax3.plot(xc[0,:],(WaterX2NoC)*1000.0,'-',label=('Tr2 NoC %d' %(time/2.0)))
plt.ylabel('Water over shelf C > 7.40 $umol$ $l^{-1}$ ($m^3 km^{-1}$)')
plt.xlabel('Along-shore distance ($km$)')
labels = [10,20,30,40, 50, 60, 70, 80,90,100,110,120]
plt.xticks([10000,20000,30000,40000,50000,60000,70000,80000,90000,100000,110000,120000], labels)
#plt.title = ('day %d' %(time/2.0))
plt.legend(loc=0)
ax6 = plt.subplot(2,3,6)
ax6.plot(xc[0,:],(Tr1XIso1)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1XIso1)*dxG[0,:]*1.E-6)))) # 1000m/km
ax6.plot(xc[0,:],(Tr2XIso1)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr2XIso1)*dxG[0,:]*1.E-6))))
ax6.plot(xc[0,:],(Tr1XNoC)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1XNoC)*dxG[0,:]*1.E-6))))
ax6.plot(xc[0,:],(Tr2XNoC)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr2XNoC)*dxG[0,:]*1.E-6))))
plt.ylabel('Tracer mass per km over shelf ($mol$ $km^{-1}$)')
plt.xlabel('Along-shore distance ($km$)')
labels = [10,20,30,40, 50, 60, 70, 80,90,100,110,120]
plt.xticks([10000,20000,30000,40000,50000,60000,70000,80000,90000,100000,110000,120000], labels)
#ax2.title = ('day %d' %(time/2.0))
plt.legend(loc=0)
#fig45.savefig('/ocean/kramosmu/Figures/WaterVolumeOverShelf/H20TrPerKm3DCNT1-NoC1.eps', format='eps', dpi=1000,bbox_extra_artists=(leg,), bbox_inches='tight')
# +
fig45=plt.figure(figsize=(18,12))
sns.set(context='paper', style='whitegrid', font='sans-serif', font_scale=1.3, rc={"lines.linewidth": 1.5})
time = 6
(WaterX1NoC, Tr1XNoC) = HowMuchWaterX(Tr1NoC,MaskNoC,30,rANoC,hFacCNoC,drFNoC,time,nx,dxG)
(WaterX1NoCNoR, Tr1XNoCNoR) = HowMuchWaterX(Tr1NoREDINoC,MaskNoC,30,rANoC,hFacCNoC,drFNoC,time,nx,dxG)
ax1 = plt.subplot(2,3,1)
ax1.plot(xc[0,:],(WaterX1NoC)*1000.0,'-',label=('NoC day %d' %(time/2.0)))
ax1.plot(xc[0,:],(WaterX1NoCNoR)*1000.0,'-',label=('NoC NoREDI day %d' %(time/2.0)))
plt.ylabel('Water over shelf C > 7.40 $umol$ $l^{-1}$ ($m^3 km^{-1}$)')
plt.xlabel('Along-shore distance ($km$)')
labels = [10,20,30,40, 50, 60, 70, 80,90,100,110,120]
plt.xticks([10000,20000,30000,40000,50000,60000,70000,80000,90000,100000,110000,120000], labels)
#ax2.title = ('day %d' %(time/2.0))
plt.legend(loc=0)
ax4 = plt.subplot(2,3,4)
ax4.plot(xc[0,:],(Tr1XNoC)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1XNoC)*dxG[0,:]*1.E-6))))
ax4.plot(xc[0,:],(Tr1XNoCNoR)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1XNoCNoR)*dxG[0,:]*1.E-6))))
plt.ylabel('Tracer mass per km over shelf ($mol$ $km^{-1}$)')
plt.xlabel('Along-shore distance ($km$)')
labels = [10,20,30,40, 50, 60, 70, 80,90,100,110,120]
plt.xticks([10000,20000,30000,40000,50000,60000,70000,80000,90000,100000,110000,120000], labels)
#ax2.title = ('day %d' %(time/2.0))
plt.legend(loc=0)
time = 10
(WaterX1NoC, Tr1XNoC) = HowMuchWaterX(Tr1NoC,MaskNoC,30,rANoC,hFacCNoC,drFNoC,time,nx,dxG)
(WaterX1NoCNoR, Tr1XNoCNoR) = HowMuchWaterX(Tr1NoREDINoC,MaskNoC,30,rANoC,hFacCNoC,drFNoC,time,nx,dxG)
ax2 = plt.subplot(2,3,2)
ax2.plot(xc[0,:],(WaterX1NoC)*1000.0,'-',label=('NoC day %d' %(time/2.0)))
ax2.plot(xc[0,:],(WaterX1NoCNoR)*1000.0,'-',label=('NoC NoREDI day %d' %(time/2.0)))
plt.ylabel('Water over shelf C > 7.40 $umol$ $l^{-1}$ ($m^3 km^{-1}$)')
plt.xlabel('Along-shore distance ($km$)')
labels = [10,20,30,40, 50, 60, 70, 80,90,100,110,120]
plt.xticks([10000,20000,30000,40000,50000,60000,70000,80000,90000,100000,110000,120000], labels)
#plt.title = ('day %d' %(time/2.0))
plt.legend(loc=0)
ax5 = plt.subplot(2,3,5)
ax5.plot(xc[0,:],(Tr1XNoC)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1XNoC)*dxG[0,:]*1.E-6))))
ax5.plot(xc[0,:],(Tr1XNoCNoR)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1XNoCNoR)*dxG[0,:]*1.E-6))))
plt.ylabel('Tracer mass per km over shelf ($mol$ $km^{-1}$)')
plt.xlabel('Along-shore distance ($km$)')
labels = [10,20,30,40, 50, 60, 70, 80,90,100,110,120]
plt.xticks([10000,20000,30000,40000,50000,60000,70000,80000,90000,100000,110000,120000], labels)
#ax2.title = ('day %d' %(time/2.0))
plt.legend(loc=0)
time = 16
(WaterX1NoC, Tr1XNoC) = HowMuchWaterX(Tr1NoC,MaskNoC,30,rANoC,hFacCNoC,drFNoC,time,nx,dxG)
(WaterX1NoCNoR, Tr1XNoCNoR) = HowMuchWaterX(Tr1NoREDINoC,MaskNoC,30,rANoC,hFacCNoC,drFNoC,time,nx,dxG)
ax3 = plt.subplot(2,3,3)
ax3.plot(xc[0,:],(WaterX1NoC)*1000.0,'-',label=(' NoC day %d' %(time/2.0))) # 1000m/km
ax3.plot(xc[0,:],(WaterX1NoCNoR)*1000.0,'-',label=('NoC NoREDI day %d' %(time/2.0)))
plt.ylabel('Water over shelf C > 7.40 $umol$ $l^{-1}$ ($m^3 km^{-1}$)')
plt.xlabel('Along-shore distance ($km$)')
labels = [10,20,30,40, 50, 60, 70, 80,90,100,110,120]
plt.xticks([10000,20000,30000,40000,50000,60000,70000,80000,90000,100000,110000,120000], labels)
#plt.title = ('day %d' %(time/2.0))
plt.legend(loc=0)
ax6 = plt.subplot(2,3,6)
ax6.plot(xc[0,:],(Tr1XNoC)*1.E-3,'-',label=('%.3e mol' %(np.sum(( Tr1XNoC)*dxG[0,:]*1.E-6))))
ax6.plot(xc[0,:],(Tr1XNoCNoR)*1.E-3,'-',label=('%.3e mol' %(np.sum(( Tr1XNoCNoR)*dxG[0,:]*1.E-6))))
plt.ylabel('Tracer mass per km over shelf ($mol$ $km^{-1}$)')
plt.xlabel('Along-shore distance ($km$)')
labels = [10,20,30,40, 50, 60, 70, 80,90,100,110,120]
plt.xticks([10000,20000,30000,40000,50000,60000,70000,80000,90000,100000,110000,120000], labels)
#ax2.title = ('day %d' %(time/2.0))
plt.legend(loc=0)
#fig45.savefig('/ocean/kramosmu/Figures/WaterVolumeOverShelf/H20TrPerKm3DCNT1-NoC1.eps', format='eps', dpi=1000,bbox_extra_artists=(leg,), bbox_inches='tight')
# +
fig45=plt.figure(figsize=(18,12))
sns.set(context='paper', style='whitegrid', font='sans-serif', font_scale=1.3, rc={"lines.linewidth": 1.5})
time = 6
(WaterX1Iso1, Tr1XIso1) = HowMuchWaterX(Tr1Iso1,MaskNoC,30,rA,hFacCNoC,drFNoC,time,nx,dxG)
(WaterX33D, Tr3X3D) = HowMuchWaterX(Tr33D,MaskNoC,30,rA,hFacCNoC,drFNoC,time,nx,dxG)
(WaterX1NoC, Tr1XNoC) = HowMuchWaterX(Tr1NoC,MaskNoC,30,rANoC,hFacCNoC,drFNoC,time,nx,dxG)
(WaterX1NoREDI02, Tr1XNoREDI02) = HowMuchWaterX(Tr1NoREDI02,MaskNoC,30,rA,hFacCNoC,drFNoC,time,nx,dxG)
(WaterX1NoCNoR, Tr1XNoCNoR) = HowMuchWaterX(Tr1NoREDINoC,MaskNoC,30,rANoC,hFacCNoC,drFNoC,time,nx,dxG)
ax1 = plt.subplot(2,3,1)
ax1.plot(xc[0,:],(WaterX1Iso1-WaterX1NoC)*1000.0,'-',label=('CNT Tr1 - NoC day %d' %(time/2.0))) # 1000m/km
ax1.plot(xc[0,:],(WaterX33D-WaterX1NoC)*1000.0,'-',label=('3D Tr3 - NoC day %d' %(time/2.0)))
ax1.plot(xc[0,:],(WaterX1NoREDI02-WaterX1NoCNoR)*1000.0,'-',label=('NoR Tr1 - NoCNoR day %d' %(time/2.0)))
plt.ylabel('Water over shelf C > 7.40 $umol$ $l^{-1}$ ($m^3 km^{-1}$)')
plt.xlabel('Along-shore distance ($km$)')
labels = [10,20,30,40, 50, 60, 70, 80,90,100,110,120]
plt.xticks([10000,20000,30000,40000,50000,60000,70000,80000,90000,100000,110000,120000], labels)
#ax2.title = ('day %d' %(time/2.0))
plt.legend(loc=0)
ax4 = plt.subplot(2,3,4)
ax4.plot(xc[0,:],(Tr1XIso1-Tr1XNoC)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1XIso1-Tr1XNoC)*dxG[0,:]*1.E-6)))) # 1000m/km
ax4.plot(xc[0,:],(Tr3X3D - Tr1XNoC)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr3X3D - Tr1XNoC)*dxG[0,:]*1.E-6))))
ax4.plot(xc[0,:],(Tr1XNoREDI02 - Tr1XNoCNoR)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1NoREDI02 - Tr1XNoCNoR)*dxG[0,:]*1.E-6))))
plt.ylabel('Tracer mass per km over shelf ($mol$ $km^{-1}$)')
plt.xlabel('Along-shore distance ($km$)')
labels = [10,20,30,40, 50, 60, 70, 80,90,100,110,120]
plt.xticks([10000,20000,30000,40000,50000,60000,70000,80000,90000,100000,110000,120000], labels)
#ax2.title = ('day %d' %(time/2.0))
plt.legend(loc=0)
time = 10
(WaterX1Iso1, Tr1XIso1) = HowMuchWaterX(Tr1Iso1,MaskNoC,30,rA,hFacCNoC,drFNoC,time,nx,dxG)
(WaterX33D, Tr3X3D) = HowMuchWaterX(Tr33D,MaskNoC,30,rA,hFacCNoC,drFNoC,time,nx,dxG)
(WaterX1NoC, Tr1XNoC) = HowMuchWaterX(Tr1NoC,MaskNoC,30,rANoC,hFacCNoC,drFNoC,time,nx,dxG)
(WaterX1NoREDI02, Tr1XNoREDI02) = HowMuchWaterX(Tr1NoREDI02,MaskNoC,30,rA,hFacCNoC,drFNoC,time,nx,dxG)
(WaterX1NoCNoR, Tr1XNoCNoR) = HowMuchWaterX(Tr1NoREDINoC,MaskNoC,30,rANoC,hFacCNoC,drFNoC,time,nx,dxG)
ax2 = plt.subplot(2,3,2)
ax2.plot(xc[0,:],(WaterX1Iso1-WaterX1NoC)*1000.0,'-',label=('CNT Tr1 - NoC day %d' %(time/2.0))) # 1000m/km
ax2.plot(xc[0,:],(WaterX33D-WaterX1NoC)*1000.0,'-',label=('3D Tr3 - NoC day %d' %(time/2.0)))
ax2.plot(xc[0,:],(WaterX1NoREDI02-WaterX1NoCNoR)*1000.0,'-',label=('NoR Tr1 - NoCNoR day %d' %(time/2.0)))
plt.ylabel('Water over shelf C > 7.40 $umol$ $l^{-1}$ ($m^3 km^{-1}$)')
plt.xlabel('Along-shore distance ($km$)')
labels = [10,20,30,40, 50, 60, 70, 80,90,100,110,120]
plt.xticks([10000,20000,30000,40000,50000,60000,70000,80000,90000,100000,110000,120000], labels)
#plt.title = ('day %d' %(time/2.0))
plt.legend(loc=0)
ax5 = plt.subplot(2,3,5)
ax5.plot(xc[0,:],(Tr1XIso1-Tr1XNoC)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1XIso1-Tr1XNoC)*dxG[0,:]*1.E-6)))) # 1000m/km
ax5.plot(xc[0,:],(Tr3X3D - Tr1XNoC)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr3X3D - Tr1XNoC)*dxG[0,:]*1.E-6))))
ax5.plot(xc[0,:],(Tr1XNoREDI02 - Tr1XNoCNoR)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1NoREDI02 - Tr1XNoCNoR)*dxG[0,:]*1.E-6))))
plt.ylabel('Tracer mass per km over shelf ($mol$ $km^{-1}$)')
plt.xlabel('Along-shore distance ($km$)')
labels = [10,20,30,40, 50, 60, 70, 80,90,100,110,120]
plt.xticks([10000,20000,30000,40000,50000,60000,70000,80000,90000,100000,110000,120000], labels)
#ax2.title = ('day %d' %(time/2.0))
plt.legend(loc=0)
time = 16
(WaterX1Iso1, Tr1XIso1) = HowMuchWaterX(Tr1Iso1,MaskNoC,30,rA,hFacCNoC,drFNoC,time,nx,dxG)
(WaterX33D, Tr3X3D) = HowMuchWaterX(Tr33D,MaskNoC,30,rA,hFacCNoC,drFNoC,time,nx,dxG)
(WaterX1NoC, Tr1XNoC) = HowMuchWaterX(Tr1NoC,MaskNoC,30,rANoC,hFacCNoC,drFNoC,time,nx,dxG)
(WaterX1NoREDI02, Tr1XNoREDI02) = HowMuchWaterX(Tr1NoREDI02,MaskNoC,30,rA,hFacCNoC,drFNoC,time,nx,dxG)
(WaterX1NoCNoR, Tr1XNoCNoR) = HowMuchWaterX(Tr1NoREDINoC,MaskNoC,30,rANoC,hFacCNoC,drFNoC,time,nx,dxG)
ax3 = plt.subplot(2,3,3)
ax3.plot(xc[0,:],(WaterX1Iso1-WaterX1NoC)*1000.0,'-',label=('CNT Tr1 - NoC day %d' %(time/2.0))) # 1000m/km
ax3.plot(xc[0,:],(WaterX33D-WaterX1NoC)*1000.0,'-',label=('3D Tr3 - NoC day %d' %(time/2.0)))
ax3.plot(xc[0,:],(WaterXNoREDI02-WaterX1NoCNoR)*1000.0,'-',label=('NoR Tr1 - NoCNoR %d' %(time/2.0)))
plt.ylabel('Water over shelf C > 7.40 $umol$ $l^{-1}$ ($m^3 km^{-1}$)')
plt.xlabel('Along-shore distance ($km$)')
labels = [10,20,30,40, 50, 60, 70, 80,90,100,110,120]
plt.xticks([10000,20000,30000,40000,50000,60000,70000,80000,90000,100000,110000,120000], labels)
#plt.title = ('day %d' %(time/2.0))
plt.legend(loc=0)
ax6 = plt.subplot(2,3,6)
ax6.plot(xc[0,:],(Tr1XIso1-Tr1XNoC)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1XIso1-Tr1XNoC)*dxG[0,:]*1.E-6)))) # 1000m/km
ax6.plot(xc[0,:],(Tr3X3D - Tr1XNoC)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr3X3D - Tr1XNoC)*dxG[0,:]*1.E-6))))
ax6.plot(xc[0,:],(Tr1XNoREDI02 - Tr1XNoCNoR)*1.E-3,'-',label=('%.3e mol' %(np.sum((Tr1NoREDI02 - Tr1XNoCNoR)*dxG[0,:]*1.E-6))))
plt.ylabel('Tracer mass per km over shelf ($mol$ $km^{-1}$)')
plt.xlabel('Along-shore distance ($km$)')
labels = [10,20,30,40, 50, 60, 70, 80,90,100,110,120]
plt.xticks([10000,20000,30000,40000,50000,60000,70000,80000,90000,100000,110000,120000], labels)
#ax2.title = ('day %d' %(time/2.0))
plt.legend(loc=0)
#fig45.savefig('/ocean/kramosmu/Figures/WaterVolumeOverShelf/H20TrPerKm3DCNT1-NoC1.eps', format='eps', dpi=1000,bbox_extra_artists=(leg,), bbox_inches='tight')
# -
print(np.shape(Tr1NoREDI02 ))
print(np.shape(Tr1XNoCNoR))
| TotalTracerAlongShelfCases.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <font color='blue'>Data Science Academy</font>
# # <font color='blue'>Big Data Real-Time Analytics com Python e Spark</font>
#
# # <font color='blue'>Capítulo 6</font>
# # Machine Learning em Python - Parte 2 - Regressão
from IPython.display import Image
Image(url = 'images/processo.png')
import sklearn as sl
import warnings
warnings.filterwarnings("ignore")
sl.__version__
# ## Definição do Problema de Negócio
# Vamos criar um modelo preditivo que seja capaz de prever o preço de casas com base em uma série de variáveis (características) sobre diversas casas em um bairro de Boston, cidade dos EUA.
#
# Dataset: https://www.cs.toronto.edu/~delve/data/boston/bostonDetail.html
# ## Avaliando a Performance
# https://scikit-learn.org/stable/modules/model_evaluation.html
# As métricas que você escolhe para avaliar a performance do modelo vão influenciar a forma como a performance é medida e comparada com modelos criados com outros algoritmos.
# ### Métricas para Algoritmos de Regressão
# Métricas Para Avaliar Modelos de Regressão
#
# - Mean Squared Error (MSE)
# - Root Mean Squared Error (RMSE)
# - Mean Absolute Error (MAE)
# - R Squared (R²)
# - Adjusted R Squared (R²)
# - Mean Square Percentage Error (MSPE)
# - Mean Absolute Percentage Error (MAPE)
# - Root Mean Squared Logarithmic Error (RMSLE)
#
from IPython.display import Image
Image(url = 'images/mse.png')
from IPython.display import Image
Image(url = 'images/rmse.png')
from IPython.display import Image
Image(url = 'images/mae.png')
from IPython.display import Image
Image(url = 'images/r2.png')
# Como vamos agora estudar as métricas para regressão, usaremos outro dataset, o Boston Houses.
# #### MSE
#
# É talvez a métrica mais simples e comum para a avaliação de regressão, mas também provavelmente a menos útil. O MSE basicamente mede o erro quadrado médio de nossas previsões. Para cada ponto, calcula a diferença quadrada entre as previsões e o valor real da variável alvo e, em seguida, calcula a média desses valores.
#
# Quanto maior esse valor, pior é o modelo. Esse valor nunca será negativo, já que estamos elevando ao quadrado os erros individuais de previsão, mas seria zero para um modelo perfeito.
# +
# MSE - Mean Squared Error
# Similar ao MAE, fornece a magnitude do erro do modelo.
# Quanto maior, pior é o modelo!
# Ao extrairmos a raiz quadrada do MSE convertemos as unidades de volta ao original,
# o que pode ser útil para descrição e apresentação. Isso é chamado RMSE (Root Mean Squared Error)
# Import dos módulos
from pandas import read_csv
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error
from sklearn.linear_model import LinearRegression
# Carregando os dados
arquivo = 'data/boston-houses.csv'
colunas = ['CRIM', 'ZN', 'INDUS', 'CHAS', 'NOX', 'RM', 'AGE', 'DIS', 'RAD', 'TAX', 'PTRATIO','B', 'LSTAT', 'MEDV']
dados = read_csv(arquivo, delim_whitespace = True, names = colunas)
array = dados.values
# Separando o array em componentes de input e output
X = array[:,0:13]
Y = array[:,13]
# Divide os dados em treino e teste
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size = 0.33, random_state = 5)
# Criando o modelo
modelo = LinearRegression()
# Treinando o modelo
modelo.fit(X_train, Y_train)
# Fazendo previsões
Y_pred = modelo.predict(X_test)
# Resultado
mse = mean_squared_error(Y_test, Y_pred)
print("O MSE do modelo é:", mse)
# -
# #### MAE
# +
# MAE
# Mean Absolute Error
# É a soma da diferença absoluta entre previsões e valores reais.
# Fornece uma ideia de quão erradas estão nossas previsões.
# Valor igual a 0 indica que não há erro, sendo a previsão perfeita.
# Import dos módulos
from pandas import read_csv
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_absolute_error
from sklearn.linear_model import LinearRegression
# Carregando os dados
arquivo = 'data/boston-houses.csv'
colunas = ['CRIM', 'ZN', 'INDUS', 'CHAS', 'NOX', 'RM', 'AGE', 'DIS', 'RAD', 'TAX', 'PTRATIO','B', 'LSTAT', 'MEDV']
dados = read_csv(arquivo, delim_whitespace = True, names = colunas)
array = dados.values
# Separando o array em componentes de input e output
X = array[:,0:13]
Y = array[:,13]
# Divide os dados em treino e teste
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size = 0.33, random_state = 5)
# Criando o modelo
modelo = LinearRegression()
# Treinando o modelo
modelo.fit(X_train, Y_train)
# Fazendo previsões
Y_pred = modelo.predict(X_test)
# Resultado
mae = mean_absolute_error(Y_test, Y_pred)
print("O MAE do modelo é:", mae)
# -
# ### R^2
# +
# R^2
# Essa métrica fornece uma indicação do nível de precisão das previsões em relação aos valores observados.
# Também chamado de coeficiente de determinação.
# Valores entre 0 e 1, sendo 0 o valor ideal.
# Import dos módulos
from pandas import read_csv
from sklearn.model_selection import train_test_split
from sklearn.metrics import r2_score
from sklearn.linear_model import LinearRegression
# Carregando os dados
arquivo = 'data/boston-houses.csv'
colunas = ['CRIM', 'ZN', 'INDUS', 'CHAS', 'NOX', 'RM', 'AGE', 'DIS', 'RAD', 'TAX', 'PTRATIO','B', 'LSTAT', 'MEDV']
dados = read_csv(arquivo, delim_whitespace = True, names = colunas)
array = dados.values
# Separando o array em componentes de input e output
X = array[:,0:13]
Y = array[:,13]
# Divide os dados em treino e teste
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size = 0.33, random_state = 5)
# Criando o modelo
modelo = LinearRegression()
# Treinando o modelo
modelo.fit(X_train, Y_train)
# Fazendo previsões
Y_pred = modelo.predict(X_test)
# Resultado
r2 = r2_score(Y_test, Y_pred)
print("O R2 do modelo é:", r2)
# -
# # Algoritmos de Regressão
# ## Regressão Linear
# Assume que os dados estão em Distribuição Normal e também assume que as variáveis são relevantes para a construção do modelo e que não sejam colineares, ou seja, variáveis com alta correlação (cabe a você, Cientista de Dados, entregar ao algoritmo as variáveis realmente relevantes).
# +
# Import dos módulos
from pandas import read_csv
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error
from sklearn.linear_model import LinearRegression
# Carregando os dados
arquivo = 'data/boston-houses.csv'
colunas = ['CRIM', 'ZN', 'INDUS', 'CHAS', 'NOX', 'RM', 'AGE', 'DIS', 'RAD', 'TAX', 'PTRATIO','B', 'LSTAT', 'MEDV']
dados = read_csv(arquivo, delim_whitespace = True, names = colunas)
array = dados.values
# Separando o array em componentes de input e output
X = array[:,0:13]
Y = array[:,13]
# Divide os dados em treino e teste
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size = 0.33, random_state = 5)
# Criando o modelo
modelo = LinearRegression()
# Treinando o modelo
modelo.fit(X_train, Y_train)
# Fazendo previsões
Y_pred = modelo.predict(X_test)
# Resultado
mse = mean_squared_error(Y_test, Y_pred)
print("O MSE do modelo é:", mse)
# -
# ## Ridge Regression
# Extensão para a regressão linear onde a loss function é modificada para minimizar a complexidade do modelo.
# +
# Import dos módulos
from pandas import read_csv
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error
from sklearn.linear_model import Ridge
# Carregando os dados
arquivo = 'data/boston-houses.csv'
colunas = ['CRIM', 'ZN', 'INDUS', 'CHAS', 'NOX', 'RM', 'AGE', 'DIS', 'RAD', 'TAX', 'PTRATIO','B', 'LSTAT', 'MEDV']
dados = read_csv(arquivo, delim_whitespace = True, names = colunas)
array = dados.values
# Separando o array em componentes de input e output
X = array[:,0:13]
Y = array[:,13]
# Divide os dados em treino e teste
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size = 0.33, random_state = 5)
# Criando o modelo
modelo = Ridge()
# Treinando o modelo
modelo.fit(X_train, Y_train)
# Fazendo previsões
Y_pred = modelo.predict(X_test)
# Resultado
mse = mean_squared_error(Y_test, Y_pred)
print("O MSE do modelo é:", mse)
# -
# ## Lasso Regression
# Lasso (Least Absolute Shrinkage and Selection Operator) Regression é uma modificação da regressão linear e assim como a Ridge Regression, a loss function é modificada para minimizar a complexidade do modelo.
# +
# Import dos módulos
from pandas import read_csv
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error
from sklearn.linear_model import Lasso
# Carregando os dados
arquivo = 'data/boston-houses.csv'
colunas = ['CRIM', 'ZN', 'INDUS', 'CHAS', 'NOX', 'RM', 'AGE', 'DIS', 'RAD', 'TAX', 'PTRATIO','B', 'LSTAT', 'MEDV']
dados = read_csv(arquivo, delim_whitespace = True, names = colunas)
array = dados.values
# Separando o array em componentes de input e output
X = array[:,0:13]
Y = array[:,13]
# Divide os dados em treino e teste
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size = 0.33, random_state = 5)
# Criando o modelo
modelo = Lasso()
# Treinando o modelo
modelo.fit(X_train, Y_train)
# Fazendo previsões
Y_pred = modelo.predict(X_test)
# Resultado
mse = mean_squared_error(Y_test, Y_pred)
print("O MSE do modelo é:", mse)
# -
# ## ElasticNet Regression
# ElasticNet é uma forma de regularização da regressão que combina as propriedades da regressão Ridge e LASSO. O objetivo é minimizar a complexidade do modelo, penalizando o modelo usando a soma dos quadrados dos coeficientes.
# +
# Import dos módulos
from pandas import read_csv
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error
from sklearn.linear_model import ElasticNet
# Carregando os dados
arquivo = 'data/boston-houses.csv'
colunas = ['CRIM', 'ZN', 'INDUS', 'CHAS', 'NOX', 'RM', 'AGE', 'DIS', 'RAD', 'TAX', 'PTRATIO','B', 'LSTAT', 'MEDV']
dados = read_csv(arquivo, delim_whitespace = True, names = colunas)
array = dados.values
# Separando o array em componentes de input e output
X = array[:,0:13]
Y = array[:,13]
# Divide os dados em treino e teste
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size = 0.33, random_state = 5)
# Criando o modelo
modelo = ElasticNet()
# Treinando o modelo
modelo.fit(X_train, Y_train)
# Fazendo previsões
Y_pred = modelo.predict(X_test)
# Resultado
mse = mean_squared_error(Y_test, Y_pred)
print("O MSE do modelo é:", mse)
# -
# ## KNN
# +
# Import dos módulos
from pandas import read_csv
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error
from sklearn.neighbors import KNeighborsRegressor
# Carregando os dados
arquivo = 'data/boston-houses.csv'
colunas = ['CRIM', 'ZN', 'INDUS', 'CHAS', 'NOX', 'RM', 'AGE', 'DIS', 'RAD', 'TAX', 'PTRATIO','B', 'LSTAT', 'MEDV']
dados = read_csv(arquivo, delim_whitespace = True, names = colunas)
array = dados.values
# Separando o array em componentes de input e output
X = array[:,0:13]
Y = array[:,13]
# Divide os dados em treino e teste
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size = 0.33, random_state = 5)
# Criando o modelo
modelo = KNeighborsRegressor()
# Treinando o modelo
modelo.fit(X_train, Y_train)
# Fazendo previsões
Y_pred = modelo.predict(X_test)
# Resultado
mse = mean_squared_error(Y_test, Y_pred)
print("O MSE do modelo é:", mse)
# -
# ## CART
# +
# Import dos módulos
from pandas import read_csv
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error
from sklearn.tree import DecisionTreeRegressor
# Carregando os dados
arquivo = 'data/boston-houses.csv'
colunas = ['CRIM', 'ZN', 'INDUS', 'CHAS', 'NOX', 'RM', 'AGE', 'DIS', 'RAD', 'TAX', 'PTRATIO','B', 'LSTAT', 'MEDV']
dados = read_csv(arquivo, delim_whitespace = True, names = colunas)
array = dados.values
# Separando o array em componentes de input e output
X = array[:,0:13]
Y = array[:,13]
# Divide os dados em treino e teste
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size = 0.33, random_state = 5)
# Criando o modelo
modelo = DecisionTreeRegressor()
# Treinando o modelo
modelo.fit(X_train, Y_train)
# Fazendo previsões
Y_pred = modelo.predict(X_test)
# Resultado
mse = mean_squared_error(Y_test, Y_pred)
print("O MSE do modelo é:", mse)
# -
# ## SVM
# +
# Import dos módulos
from pandas import read_csv
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error
from sklearn.svm import SVR
# Carregando os dados
arquivo = 'data/boston-houses.csv'
colunas = ['CRIM', 'ZN', 'INDUS', 'CHAS', 'NOX', 'RM', 'AGE', 'DIS', 'RAD', 'TAX', 'PTRATIO','B', 'LSTAT', 'MEDV']
dados = read_csv(arquivo, delim_whitespace = True, names = colunas)
array = dados.values
# Separando o array em componentes de input e output
X = array[:,0:13]
Y = array[:,13]
# Divide os dados em treino e teste
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size = 0.33, random_state = 5)
# Criando o modelo
modelo = SVR()
# Treinando o modelo
modelo.fit(X_train, Y_train)
# Fazendo previsões
Y_pred = modelo.predict(X_test)
# Resultado
mse = mean_squared_error(Y_test, Y_pred)
print("O MSE do modelo é:", mse)
# -
# ## Otimização do Modelo - Ajuste de Parâmetros
# Todos os algoritmos de Machine Learning são parametrizados, o que significa que você pode ajustar a performance do seu modelo preditivo, através do tuning (ajuste fino) dos parâmetros. Seu trabalho é encontrar a melhor combinação entre os parâmetros em cada algoritmo de Machine Learning. Esse processo também é chamado de Otimização Hyperparâmetro. O scikit-learn oferece dois métodos para otimização automática dos parâmetros: Grid Search Parameter Tuning e Random Search Parameter Tuning.
# ### Grid Search Parameter Tuning
# Este método realiza metodicamente combinações entre todos os parâmetros do algoritmo, criando um grid. Vamos experimentar este método utilizando o algoritmo de Regressão Ridge. No exemplo abaixo veremos que o valor 1 para o parâmetro alpha atingiu a melhor performance.
# +
# Import dos módulos
from pandas import read_csv
import numpy as np
from sklearn.model_selection import GridSearchCV
from sklearn.linear_model import Ridge
# Carregando os dados
arquivo = 'data/boston-houses.csv'
colunas = ['CRIM', 'ZN', 'INDUS', 'CHAS', 'NOX', 'RM', 'AGE', 'DIS', 'RAD', 'TAX', 'PTRATIO','B', 'LSTAT', 'MEDV']
dados = read_csv(arquivo, delim_whitespace = True, names = colunas)
array = dados.values
# Separando o array em componentes de input e output
X = array[:,0:8]
Y = array[:,8]
# Definindo os valores que serão testados
valores_alphas = np.array([1,0.1,0.01,0.001,0.0001,0])
valores_grid = dict(alpha = valores_alphas)
# Criando o modelo
modelo = Ridge()
# Criando o grid
grid = GridSearchCV(estimator = modelo, param_grid = valores_grid)
grid.fit(X, Y)
# Print do resultado
print("Melhores Parâmetros do Modelo:\n", grid.best_estimator_)
# -
# ### Random Search Parameter Tuning
# Este método gera amostras dos parâmetros dos algoritmos a partir de uma distribuição randômica uniforme para um número fixo de interações. Um modelo é construído e testado para cada combinação de parâmetros. Neste exemplo veremos que o valor muito próximo de 1 para o parâmetro alpha é o que vai apresentar os melhores resultados.
# +
# Import dos módulos
from pandas import read_csv
import numpy as np
from scipy.stats import uniform
from sklearn.linear_model import Ridge
from sklearn.model_selection import RandomizedSearchCV
# Carregando os dados
arquivo = 'data/boston-houses.csv'
colunas = ['CRIM', 'ZN', 'INDUS', 'CHAS', 'NOX', 'RM', 'AGE', 'DIS', 'RAD', 'TAX', 'PTRATIO','B', 'LSTAT', 'MEDV']
dados = read_csv(arquivo, delim_whitespace = True, names = colunas)
array = dados.values
# Separando o array em componentes de input e output
X = array[:,0:8]
Y = array[:,8]
# Definindo os valores que serão testados
valores_grid = {'alpha': uniform()}
seed = 7
# Criando o modelo
modelo = Ridge()
iterations = 100
rsearch = RandomizedSearchCV(estimator = modelo,
param_distributions = valores_grid,
n_iter = iterations,
random_state = seed)
rsearch.fit(X, Y)
# Print do resultado
print("Melhores Parâmetros do Modelo:\n", rsearch.best_estimator_)
# -
# # Salvando o resultado do seu trabalho
# +
# Import dos módulos
from pandas import read_csv
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error
from sklearn.linear_model import Ridge
import pickle
# Carregando os dados
arquivo = 'data/boston-houses.csv'
colunas = ['CRIM', 'ZN', 'INDUS', 'CHAS', 'NOX', 'RM', 'AGE', 'DIS', 'RAD', 'TAX', 'PTRATIO','B', 'LSTAT', 'MEDV']
dados = read_csv(arquivo, delim_whitespace = True, names = colunas)
array = dados.values
# Separando o array em componentes de input e output
X = array[:,0:13]
Y = array[:,13]
# Definindo os valores para o número de folds
teste_size = 0.35
seed = 7
# Criando o dataset de treino e de teste
X_treino, X_teste, Y_treino, Y_teste = train_test_split(X, Y, test_size = teste_size, random_state = seed)
# Criando o modelo
modelo = Ridge()
# Treinando o modelo
modelo.fit(X_treino, Y_treino)
# Salvando o modelo
arquivo = 'modelos/modelo_regressor_final.sav'
pickle.dump(modelo, open(arquivo, 'wb'))
print("Modelo salvo!")
# Carregando o arquivo
modelo_regressor_final = pickle.load(open(arquivo, 'rb'))
print("Modelo carregado!")
# Print do resultado
# Fazendo previsões
Y_pred = modelo_regressor_final.predict(X_test)
# Resultado
mse = mean_squared_error(Y_test, Y_pred)
print("O MSE do modelo é:", mse)
# -
# # Fim
# ### Obrigado - Data Science Academy - <a href="http://facebook.com/dsacademybr">facebook.com/dsacademybr</a>
| Cap06 - Machine Learning em Python/Machine Learning em Python - Regressão/Processo-Machine-Learning-Parte2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# # Sample grouping
# We are going to linger into the concept of sample groups. As in the previous
# section, we will give an example to highlight some surprising results. This
# time, we will use the handwritten digits dataset.
# +
from sklearn.datasets import load_digits
digits = load_digits()
data, target = digits.data, digits.target
# -
# We will recreate the same model used in the previous exercise:
# a logistic regression classifier with preprocessor to scale the data.
# +
from sklearn.preprocessing import MinMaxScaler
from sklearn.linear_model import LogisticRegression
from sklearn.pipeline import make_pipeline
model = make_pipeline(MinMaxScaler(), LogisticRegression(max_iter=1_000))
# -
# We will use the same baseline model. We will use a `KFold` cross-validation
# without shuffling the data at first.
# +
from sklearn.model_selection import cross_val_score, KFold
cv = KFold(shuffle=False)
test_score_no_shuffling = cross_val_score(model, data, target, cv=cv,
n_jobs=2)
print(f"The average accuracy is "
f"{test_score_no_shuffling.mean():.3f} +/- "
f"{test_score_no_shuffling.std():.3f}")
# -
# Now, let's repeat the experiment by shuffling the data within the
# cross-validation.
cv = KFold(shuffle=True)
test_score_with_shuffling = cross_val_score(model, data, target, cv=cv,
n_jobs=2)
print(f"The average accuracy is "
f"{test_score_with_shuffling.mean():.3f} +/- "
f"{test_score_with_shuffling.std():.3f}")
# We observe that shuffling the data improves the mean accuracy.
# We could go a little further and plot the distribution of the testing
# score. We can first concatenate the test scores.
# +
import pandas as pd
all_scores = pd.DataFrame(
[test_score_no_shuffling, test_score_with_shuffling],
index=["KFold without shuffling", "KFold with shuffling"],
).T
# -
# Let's plot the distribution now.
# +
import matplotlib.pyplot as plt
all_scores.plot.hist(bins=10, edgecolor="black", alpha=0.7)
plt.xlim([0.8, 1.0])
plt.xlabel("Accuracy score")
plt.legend(bbox_to_anchor=(1.05, 0.8), loc="upper left")
_ = plt.title("Distribution of the test scores")
# -
# The cross-validation testing error that uses the shuffling has less
# variance than the one that does not impose any shuffling. It means that some
# specific fold leads to a low score in this case.
print(test_score_no_shuffling)
# Thus, there is an underlying structure in the data that shuffling will break
# and get better results. To get a better understanding, we should read the
# documentation shipped with the dataset.
print(digits.DESCR)
# If we read carefully, 13 writers wrote the digits of our dataset, accounting
# for a total amount of 1797 samples. Thus, a writer wrote several times the
# same numbers. Let's suppose that the writer samples are grouped.
# Subsequently, not shuffling the data will keep all writer samples together
# either in the training or the testing sets. Mixing the data will break this
# structure, and therefore digits written by the same writer will be available
# in both the training and testing sets.
#
# Besides, a writer will usually tend to write digits in the same manner. Thus,
# our model will learn to identify a writer's pattern for each digit instead of
# recognizing the digit itself.
#
# We can solve this problem by ensuring that the data associated with a writer
# should either belong to the training or the testing set. Thus, we want to
# group samples for each writer.
#
# Indeed, we can recover the groups by looking at the target variable.
target[:200]
#
# It might not be obvious at first, but there is a structure in the target:
# there is a repetitive pattern that always starts by some series of ordered
# digits from 0 to 9 followed by random digits at a certain point. If we look
# in details, we see that there is 14 such patterns, always with around 130
# samples each.
#
# Even if it is not exactly corresponding to the 13 writers in the
# documentation (maybe one writer wrote two series of digits), we can
# make the hypothesis that each of these patterns corresponds to a different
# writer and thus a different group.
# +
from itertools import count
import numpy as np
# defines the lower and upper bounds of sample indices
# for each writer
writer_boundaries = [0, 130, 256, 386, 516, 646, 776, 915, 1029,
1157, 1287, 1415, 1545, 1667, 1797]
groups = np.zeros_like(target)
lower_bounds = writer_boundaries[:-1]
upper_bounds = writer_boundaries[1:]
for group_id, lb, up in zip(count(), lower_bounds, upper_bounds):
groups[lb:up] = group_id
# -
# We can check the grouping by plotting the indices linked to writer ids.
plt.plot(groups)
plt.yticks(np.unique(groups))
plt.xticks(writer_boundaries, rotation=90)
plt.xlabel("Target index")
plt.ylabel("Writer index")
_ = plt.title("Underlying writer groups existing in the target")
# Once we group the digits by writer, we can use cross-validation to take this
# information into account: the class containing `Group` should be used.
# +
from sklearn.model_selection import GroupKFold
cv = GroupKFold()
test_score = cross_val_score(model, data, target, groups=groups, cv=cv,
n_jobs=2)
print(f"The average accuracy is "
f"{test_score.mean():.3f} +/- "
f"{test_score.std():.3f}")
# -
# We see that this strategy is less optimistic regarding the model generalization
# performance. However, this is the most reliable if our goal is to make
# handwritten digits recognition writers independent. Besides, we can as well
# see that the standard deviation was reduced.
all_scores = pd.DataFrame(
[test_score_no_shuffling, test_score_with_shuffling, test_score],
index=["KFold without shuffling", "KFold with shuffling",
"KFold with groups"],
).T
all_scores.plot.hist(bins=10, edgecolor="black", alpha=0.7)
plt.xlim([0.8, 1.0])
plt.xlabel("Accuracy score")
plt.legend(bbox_to_anchor=(1.05, 0.8), loc="upper left")
_ = plt.title("Distribution of the test scores")
# As a conclusion, it is really important to take any sample grouping pattern
# into account when evaluating a model. Otherwise, the results obtained will
# be over-optimistic in regards with reality.
| notebooks/80 - cross_validation_grouping.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Una vez finalizado el curso, por favor ayúdanos completando la siguiente **Encuesta de FINAL del curso**.
#
# [](https://forms.office.com/Pages/ResponsePage.aspx?id=r4yvt9iDREaFrjF8VFIjwUHkKiCq1wxFstxAwkoFiilUOExRVkVMWlZERVcyWlpUU1EyTFg4T1Q3WC4u)
#
# ## 11. Ecuaciones Diferenciales
#
# [Playlist de Ciencia de Datos en castellano](https://www.youtube.com/playlist?list=PLjyvn6Y1kpbEmRY4-ELeRA80ZywV7Xd67)
# [](https://www.youtube.com/watch?v=HReAo38LoM4&list=PLLBUgWXdTBDg1Qgmwt4jKtVn9BWh5-zgy "Python Data Science")
#
# Algunas ecuaciones con términos diferenciales surgen de relaciones fundamentales como conservación de masa, energía y momento. Por ejemplo, la aculumación de masa $\frac{dm}{dt}$ en un volumen de control es igual a la masa que entra $\dot m_{in}$ menos la masa que sale $\dot m_{out}$ de ese volumen.
#
# $\frac{dm}{dt} = \dot m_{in} - \dot m_{out}$
#
# Se puede desarrollar un modelo dinámico mediante regresión de datos o con relaciones fundamentales sin necesidad de datos. Incluso las relaciones fundamentales pueden tener parámetros desconocidos o inciertos. Un enfoque para el modelado dinámico es combinar relaciones físicas fundamentales con Ciencia de Datos. Este enfoque usa lo mejor de ambos métodos porque crea un modelo que se alinea con los valores medidos y puede extrapolarse a regiones donde los datos son limitados o inexistentes.
#
# 
#
# En este primer ejercicio para [resolver ecuaciones diferenciales](https://www.youtube.com/watch?v=v9fGOHQMeIA) vamos a utilizar `odeint`. Los mismos ejemplos también serán [resueltos con Gekko](https://apmonitor.com/pdc/index.php/Main/PythonDifferentialEquations). Ambos alcanzan resultados equivalentes de simulación. Sin embargo, Gekko está diseñado para usar ecuaciones diferenciales en optimización o combinarse con aprendizaje automático (machine learning). La función `odeint` tiene como propósito principal resolver ecuaciones diferenciales ordinarias (EDO), y requiere tres entradas (inputs).
#
# y = odeint(model, y0, t)
#
# 1. `model` Nombre de la Función que devuelve la derivada para un par de valores solicitados `y`, `t`, de la forma `dydt = model(y,t)`.
# 2. `y0` Condiciones iniciales.
# 3. `t` Puntos de tiempo donde se reporta la solución.
#
# 
#
# ### Resolver Ecuaciones Diferenciales
#
# Resolveremos la ecuación diferencial con la condición inicial $y(0) = 5$:
#
# $ k \, \frac{dy}{dt} = -y$
#
# Donde $k=10$. La solución para `y` se reporta desde un tiempo inicial `0` hasta un tiempo final `20`. También se grafica el resultado para $y(t)$ vs. $t$. Notemos cómo se establece la ecuación para obtener la derivada como `dydt = -(1.0/k) * y` a partir de la función.
# +
import numpy as np
from scipy.integrate import odeint
# función que devuelve dy/dt
def model(y,t):
k = 10.0
dydt = -(1.0/k) * y
return dydt
y0 = 5 # condición inicial
t = np.linspace(0,20) # puntos de tiempo
y = odeint(model,y0,t) # resolución de la ODE
import matplotlib.pyplot as plt
# %matplotlib inline
plt.plot(t,y)
plt.xlabel('Tiempo'); plt.ylabel('y(t)')
plt.show()
# -
# 
#
# ### Resolver Ecuaciones Diferenciales con Gekko
#
# [Python Gekko](https://gekko.readthedocs.io/en/latest/) resuelve la misma ecuación diferencial. Está diseñado para problemas a gran escala. El [tutorial de Gekko en inglés](https://apmonitor.com/wiki/index.php/Main/GekkoPythonOptimization) nos muestra cómo resolver otro tipo de problemas con ecuaciones y optimización.
# +
from gekko import GEKKO
m = GEKKO(remote=False) # modelo GEKKO
m.time = np.linspace(0,20) # puntos de tiempo
y = m.Var(5.0); k = 10.0 # variables y constantes GEKKO
m.Equation(k*y.dt()+y==0) # Ecuación GEKKO
m.options.IMODE = 4 # Simulación dinámica
m.solve(disp=False) # Resolución
plt.plot(m.time,y)
plt.xlabel('Tiempo'); plt.ylabel('y(t)')
plt.show()
# -
# 
#
# ### Actividad sobre Ecuaciones Diferenciales
#
# Resuelve la ecuación diferencial con condición inicial $y(0) = 10$:
#
# $ k \, \frac{dy}{dt} = -y$
#
# Compara las primeras cinco soluciones de `y` entre los tiempos `0` y `20` con `k=[1,2,5,10,20]`.
# 
#
# ### Solución Simbólica
#
# Los problemas con ecuaciones diferenciales que tienen solución analítica pueden expresarse simbólicamente. Una librería con símbolos matemáticos en Python es `sympy`. Sympy determina la solución analítica como $y(x)=C_1 \, \exp{\left(-\frac{x}{k}\right)}$. Con la condición inicial $y(0)=5$, y la constante $C_1$ igual a 5.
from IPython.display import display
import sympy as sym
from sympy.abc import x, k
y = sym.Function('y')
ans = sym.dsolve(sym.Derivative(y(x), x) + y(x)/k, y(x))
display(ans)
# 
#
# ### Resolver Ecuaciones Diferenciales con Entradas (Input) `u`
#
# Las ecuaciones diferenciales también pueden tener una entrada (atributo) que cambie desde una fuente externa (entrada exógena). Por ejemplo, cambios interactivos debido a medidas de un sensor, a personas (manualmente) o seleccionados por un computador.
#
# 
#
# Calcula la respuesta `y(t)` cuando la entrada `u` cambia desde `0` a `2` en `t = 5`.
#
# $2 \frac{dy(t)}{dt} + y(t) = u(t)$
#
# La condición inicial es `y(0)=1` y la solución puede calcularse hasta `t=15`. **Ayuda**: La expresión `y(t)` no es equivalente a `y` multiplicado por `t`. Esta indica que `y` cambia con el tiempo y se escribe como una función del tiempo. Hay ejemplos adicionales para [odeint en inglés](https://apmonitor.com/pdc/index.php/Main/SolveDifferentialEquations) y [Gekko en inglés](https://apmonitor.com/pdc/index.php/Main/PythonDifferentialEquations) por si necesitas ayuda.
# ### Actividad con el TCLab
#
# 
#
# ### Recolección de Datos
#
# 
#
# Enciende el calentador 1 al 100% y guarda el valor de $T_1$ cada 5 segundos durante 3 minutos. Los datos deben incluir un total de 37 puntos para cada sensor de temperatura.
import numpy as np
import pandas as pd
import tclab
import time
# Recolectar datos por 3 minutos, cada 5 segundos
n = 37
tm = np.linspace(0,180,n)
t1s = np.empty(n); t2s = np.empty(n)
with tclab.TCLab() as lab:
lab.Q1(100); lab.Q2(0)
print('Tiempo T1 T2')
for i in range(n):
t1s[i] = lab.T1; t2s[i] = lab.T2
print(tm[i],t1s[i],t2s[i])
time.sleep(5.0)
# Colocar en un Dataframe
data = pd.DataFrame(np.column_stack((tm,t1s,t2s)),\
columns=['Tiempo','T1','T2'])
data.to_csv('11-data.csv',index=False)
# 
#
# ### Resolver Ecuaciones Diferenciales
#
# Usa los parámetros `a`, `b` y `c` del módulo [10. Resolver Ecuaciones](https://github.com/APMonitor/data_science/blob/master/10.%20Solve_Equations.ipynb) o utiliza los siguientes valores:
#
# | Parámetro | Valor |
# |------|------|
# | a | 78.6 |
# | b | -50.3 |
# | c | -0.003677 |
#
# Resuelve la ecuación diferencial ordinaria (ODE en inglés) con estos valores.
#
# $\frac{dT_1}{dt} = c (T_1-a)$
#
# La condición inicial para $T_1$ es $a + b$. Muestra la solución para la ODE en el intervalo de tiempo desde `0` hasta `180` segundos. Grafica el valor medido de $T_1$ en la misma figura que muestra la predicción de la temperatura por la ODE. Añade las etiquetas necesarias en el gráfico.
| 11. Ecuaciones_diferenciales.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ### Basic Introduction
#
# LeNet-5, from the paper Gradient-Based Learning Applied to Document Recognition, is a very efficient convolutional neural network for handwritten character recognition.
#
#
# <a href="http://yann.lecun.com/exdb/publis/pdf/lecun-98.pdf" target="_blank">Paper: <u>Gradient-Based Learning Applied to Document Recognition</u></a>
#
# **Authors**: <NAME>, <NAME>, <NAME>, and <NAME>
#
# **Published in**: Proceedings of the IEEE (1998)
#
# ### Structure of the LeNet network
#
# LeNet5 is a small network, it contains the basic modules of deep learning: convolutional layer, pooling layer, and full link layer. It is the basis of other deep learning models. Here we analyze LeNet5 in depth. At the same time, through example analysis, deepen the understanding of the convolutional layer and pooling layer.
#
# 
#
#
# LeNet-5 Total seven layer , does not comprise an input, each containing a trainable parameters; each layer has a plurality of the Map the Feature , a characteristic of each of the input FeatureMap extracted by means of a convolution filter, and then each FeatureMap There are multiple neurons.
#
# 
#
# Detailed explanation of each layer parameter:
#
# #### **INPUT Layer**
#
# The first is the data INPUT layer. The size of the input image is uniformly normalized to 32 * 32.
#
# > Note: This layer does not count as the network structure of LeNet-5. Traditionally, the input layer is not considered as one of the network hierarchy.
#
#
# #### **C1 layer-convolutional layer**
#
# >**Input picture**: 32 * 32
#
# >**Convolution kernel size**: 5 * 5
#
# >**Convolution kernel types**: 6
#
# >**Output featuremap size**: 28 * 28 (32-5 + 1) = 28
#
# >**Number of neurons**: 28 * 28 * 6
#
# >**Trainable parameters**: (5 * 5 + 1) * 6 (5 * 5 = 25 unit parameters and one bias parameter per filter, a total of 6 filters)
#
# >**Number of connections**: (5 * 5 + 1) * 6 * 28 * 28 = 122304
# **Detailed description:**
#
# 1. The first convolution operation is performed on the input image (using 6 convolution kernels of size 5 * 5) to obtain 6 C1 feature maps (6 feature maps of size 28 * 28, 32-5 + 1 = 28).
#
# 2. Let's take a look at how many parameters are needed. The size of the convolution kernel is 5 * 5, and there are 6 * (5 * 5 + 1) = 156 parameters in total, where +1 indicates that a kernel has a bias.
#
# 3. For the convolutional layer C1, each pixel in C1 is connected to 5 * 5 pixels and 1 bias in the input image, so there are 156 * 28 * 28 = 122304 connections in total. There are 122,304 connections, but we only need to learn 156 parameters, mainly through weight sharing.
#
# #### **S2 layer-pooling layer (downsampling layer)**
#
# >**Input**: 28 * 28
#
# >**Sampling area**: 2 * 2
#
# >**Sampling method**: 4 inputs are added, multiplied by a trainable parameter, plus a trainable offset. Results via sigmoid
#
# >**Sampling type**: 6
#
# >**Output featureMap size**: 14 * 14 (28/2)
#
# >**Number of neurons**: 14 * 14 * 6
#
# >**Trainable parameters**: 2 * 6 (the weight of the sum + the offset)
#
# >**Number of connections**: (2 * 2 + 1) * 6 * 14 * 14
#
# >The size of each feature map in S2 is 1/4 of the size of the feature map in C1.
#
# **Detailed description:**
#
# The pooling operation is followed immediately after the first convolution. Pooling is performed using 2 * 2 kernels, and S2, 6 feature maps of 14 * 14 (28/2 = 14) are obtained.
#
# The pooling layer of S2 is the sum of the pixels in the 2 * 2 area in C1 multiplied by a weight coefficient plus an offset, and then the result is mapped again.
#
# So each pooling core has two training parameters, so there are 2x6 = 12 training parameters, but there are 5x14x14x6 = 5880 connections.
#
# #### **C3 layer-convolutional layer**
#
# >**Input**: all 6 or several feature map combinations in S2
#
# >**Convolution kernel size**: 5 * 5
#
# >**Convolution kernel type**: 16
#
# >**Output featureMap size**: 10 * 10 (14-5 + 1) = 10
#
# >Each feature map in C3 is connected to all 6 or several feature maps in S2, indicating that the feature map of this layer is a different combination of the feature maps extracted from the previous layer.
#
# >One way is that the first 6 feature maps of C3 take 3 adjacent feature map subsets in S2 as input. The next 6 feature maps take 4 subsets of neighboring feature maps in S2 as input. The next three take the non-adjacent 4 feature map subsets as input. The last one takes all the feature maps in S2 as input.
#
# >**The trainable parameters are**: 6 * (3 * 5 * 5 + 1) + 6 * (4 * 5 * 5 + 1) + 3 * (4 * 5 * 5 + 1) + 1 * (6 * 5 * 5 +1) = 1516
#
# >**Number of connections**: 10 * 10 * 1516 = 151600
#
# **Detailed description:**
#
# After the first pooling, the second convolution, the output of the second convolution is C3, 16 10x10 feature maps, and the size of the convolution kernel is 5 * 5. We know that S2 has 6 14 * 14 feature maps, how to get 16 feature maps from 6 feature maps? Here are the 16 feature maps calculated by the special combination of the feature maps of S2. details as follows:
#
#
#
#
# The first 6 feature maps of C3 (corresponding to the 6th column of the first red box in the figure above) are connected to the 3 feature maps connected to the S2 layer (the first red box in the above figure), and the next 6 feature maps are connected to the S2 layer The 4 feature maps are connected (the second red box in the figure above), the next 3 feature maps are connected with the 4 feature maps that are not connected at the S2 layer, and the last is connected with all the feature maps at the S2 layer. The convolution kernel size is still 5 * 5, so there are 6 * (3 * 5 * 5 + 1) + 6 * (4 * 5 * 5 + 1) + 3 * (4 * 5 * 5 + 1) +1 * (6 * 5 * 5 + 1) = 1516 parameters. The image size is 10 * 10, so there are 151600 connections.
#
# 
#
#
# The convolution structure of C3 and the first 3 graphs in S2 is shown below:
#
# 
# #### **S4 layer-pooling layer (downsampling layer)**
#
# >**Input**: 10 * 10
#
# >**Sampling area**: 2 * 2
#
# >**Sampling method**: 4 inputs are added, multiplied by a trainable parameter, plus a trainable offset. Results via sigmoid
#
# >**Sampling type**: 16
#
# >**Output featureMap size**: 5 * 5 (10/2)
#
# >**Number of neurons**: 5 * 5 * 16 = 400
#
# >**Trainable parameters**: 2 * 16 = 32 (the weight of the sum + the offset)
#
# >**Number of connections**: 16 * (2 * 2 + 1) * 5 * 5 = 2000
#
# >The size of each feature map in S4 is 1/4 of the size of the feature map in C3
#
# **Detailed description:**
#
# S4 is the pooling layer, the window size is still 2 * 2, a total of 16 feature maps, and the 16 10x10 maps of the C3 layer are pooled in units of 2x2 to obtain 16 5x5 feature maps. This layer has a total of 32 training parameters of 2x16, 5x5x5x16 = 2000 connections.
#
# *The connection is similar to the S2 layer.*
#
# #### **C5 layer-convolution layer**
#
# >**Input**: All 16 unit feature maps of the S4 layer (all connected to s4)
#
# >**Convolution kernel size**: 5 * 5
#
# >**Convolution kernel type**: 120
#
# >**Output featureMap size**: 1 * 1 (5-5 + 1)
#
# >**Trainable parameters / connection**: 120 * (16 * 5 * 5 + 1) = 48120
#
# **Detailed description:**
#
#
# The C5 layer is a convolutional layer. Since the size of the 16 images of the S4 layer is 5x5, which is the same as the size of the convolution kernel, the size of the image formed after convolution is 1x1. This results in 120 convolution results. Each is connected to the 16 maps on the previous level. So there are (5x5x16 + 1) x120 = 48120 parameters, and there are also 48120 connections. The network structure of the C5 layer is as follows:
#
# 
# #### **F6 layer-fully connected layer**
#
# >**Input**: c5 120-dimensional vector
#
# >**Calculation method**: calculate the dot product between the input vector and the weight vector, plus an offset, and the result is output through the sigmoid function.
#
# >**Trainable parameters**: 84 * (120 + 1) = 10164
#
# **Detailed description:**
#
# Layer 6 is a fully connected layer. The F6 layer has 84 nodes, corresponding to a 7x12 bitmap, -1 means white, 1 means black, so the black and white of the bitmap of each symbol corresponds to a code. The training parameters and number of connections for this layer are (120 + 1) x84 = 10164. The ASCII encoding diagram is as follows:
#
# 
#
# The connection method of the F6 layer is as follows:
#
# 
#
#
# #### **Output layer-fully connected layer**
#
# The output layer is also a fully connected layer, with a total of 10 nodes, which respectively represent the numbers 0 to 9, and if the value of node i is 0, the result of network recognition is the number i. A radial basis function (RBF) network connection is used. Assuming x is the input of the previous layer and y is the output of the RBF, the calculation of the RBF output is:
#
# 
#
# The value of the above formula w_ij is determined by the bitmap encoding of i, where i ranges from 0 to 9, and j ranges from 0 to 7 * 12-1. The closer the value of the RBF output is to 0, the closer it is to i, that is, the closer to the ASCII encoding figure of i, it means that the recognition result input by the current network is the character i. This layer has 84x10 = 840 parameters and connections.
#
# 
#
#
# **Summary**
#
#
# * LeNet-5 is a very efficient convolutional neural network for handwritten character recognition.
# * Convolutional neural networks can make good use of the structural information of images.
# * The convolutional layer has fewer parameters, which is also determined by the main characteristics of the convolutional layer, that is, local connection and shared weights.
#
# ### Code Implementation
# +
import tensorflow as tf
from keras.datasets import mnist
from keras.layers import Conv2D, MaxPooling2D
from keras.layers import Dense, Flatten
from keras.models import Sequential
from tensorflow.keras.utils import to_categorical
# Loading the dataset and perform splitting
(x_train, y_train), (x_test, y_test) = mnist.load_data()
# Peforming reshaping operation
x_train = x_train.reshape(x_train.shape[0], 28, 28, 1)
x_test = x_test.reshape(x_test.shape[0], 28, 28, 1)
# Normalization
x_train = x_train / 255
x_test = x_test / 255
# One Hot Encoding
y_train = tf.keras.utils.to_categorical(y_train, 10)
y_test = tf.keras.utils.to_categorical(y_test, 10)
# Building the Model Architecture
model = Sequential()
# Select 6 feature convolution kernels with a size of 5 * 5 (without offset), and get 66 feature maps. The size of each feature map is 32−5 + 1 = 2832−5 + 1 = 28.
# That is, the number of neurons has been reduced from 10241024 to 28 ∗ 28 = 784 28 ∗ 28 = 784.
# Parameters between input layer and C1 layer: 6 ∗ (5 ∗ 5 + 1)
model.add(Conv2D(6, kernel_size=(5, 5), activation='relu', input_shape=(28, 28, 1)))
# The input of this layer is the output of the first layer, which is a 28 * 28 * 6 node matrix.
# The size of the filter used in this layer is 2 * 2, and the step length and width are both 2, so the output matrix size of this layer is 14 * 14 * 6.
model.add(MaxPooling2D(pool_size=(2, 2)))
# The input matrix size of this layer is 14 * 14 * 6, the filter size used is 5 * 5, and the depth is 16. This layer does not use all 0 padding, and the step size is 1.
# The output matrix size of this layer is 10 * 10 * 16. This layer has 5 * 5 * 6 * 16 + 16 = 2416 parameters
model.add(Conv2D(16, kernel_size=(5, 5), activation='relu'))
# The input matrix size of this layer is 10 * 10 * 16. The size of the filter used in this layer is 2 * 2, and the length and width steps are both 2, so the output matrix size of this layer is 5 * 5 * 16.
model.add(MaxPooling2D(pool_size=(2, 2)))
# The input matrix size of this layer is 5 * 5 * 16. This layer is called a convolution layer in the LeNet-5 paper, but because the size of the filter is 5 * 5, #
# So it is not different from the fully connected layer. If the nodes in the 5 * 5 * 16 matrix are pulled into a vector, then this layer is the same as the fully connected layer.
# The number of output nodes in this layer is 120, with a total of 5 * 5 * 16 * 120 + 120 = 48120 parameters.
model.add(Flatten())
model.add(Dense(120, activation='relu'))
# The number of input nodes in this layer is 120 and the number of output nodes is 84. The total parameter is 120 * 84 + 84 = 10164 (w + b)
model.add(Dense(84, activation='relu'))
# The number of input nodes in this layer is 84 and the number of output nodes is 10. The total parameter is 84 * 10 + 10 = 850
model.add(Dense(10, activation='softmax'))
model.summary()
model.compile(loss=tf.keras.metrics.categorical_crossentropy, optimizer=tf.keras.optimizers.Adam(), metrics=['accuracy'])
model.fit(x_train, y_train, batch_size=128, epochs=20, verbose=1, validation_data=(x_test, y_test))
score = model.evaluate(x_test, y_test)
print('Test Loss:', score[0])
print('Test accuracy:', score[1])
# -
| CNN/CNN-Architecture/LeNet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 逻辑回归
# 1、逻辑回归与线性回归的联系与区别
# 2、逻辑回归的原理
# 3、逻辑回归损失函数推导及优化
# 4、正则化与模型评估指标
# 5、逻辑回归的优缺点
# 6、样本不均衡问题解决办法
# 7、sklearn参数
# 8、代码实现
# ## 1、逻辑回归与线性回归的联系与区别
# 线性回归解决的是连续变量问题,那么在分类任务中可以用线性回归吗?例如判断是良性肿瘤还是恶性肿瘤,判断是垃圾邮件还是正常邮件,等等……
# 答案是也可以,但是效果不好,见下图:
# <img src = "https://img-blog.csdnimg.cn/20190830143654227.png?x-oss-process=image/watermark,type_ZmFuZ3poZW5naGVpdGk,shadow_10,text_aHR0cHM6Ly9ibG9nLmNzZG4ubmV0L2RwZW5nd2FuZw==,size_16,color_FFFFFF,t_70">
# 图显示了是否购买玩具和年龄之间的关系,可以用线性回归拟合成一条直线,将购买标注为1,不购买标注为0,拟合后取当0.5值为阈值来划分类别。
#
# $$\hat y = \begin{cases}
# 1,& f(x)>0.5\\
# 0,& f(x)<0.5
# \end{cases}$$
# 可以看到,在途中,年龄的区分点约为19岁。
# 但当数据点不平衡时,很容易影响到阈值,见以下图:
# <img src = "https://img-blog.csdnimg.cn/20190830143716479.png?x-oss-process=image/watermark,type_ZmFuZ3poZW5naGVpdGk,shadow_10,text_aHR0cHM6Ly9ibG9nLmNzZG4ubmV0L2RwZW5nd2FuZw==,size_16,color_FFFFFF,t_70">
# 可以看到,0值样本的年龄段往高年龄端偏移后,真实的阈值依然是19岁左右,但拟合出来的曲线的阈值往后边偏移了。可以想想,负样本越多,年龄大的人越多,偏移越严重。
# 可是这符合实际情况吗?实际情况是60岁的老人和80岁的老人都不会购买玩具,增加几位80岁的老人,并不会影响20岁以下人群购买玩具的概率。但因为拟合曲线原本的值域为$(-\infty \ \infty)$而转换后的值域为[0,1],阈值对变量偏移很敏感。
# ## 2、逻辑回归的原理
# 因此理想的替代函数应当预测分类为0或1的概率,当为1的概率大于0.5时,判断为1,当为1的概率小于0.5时,判断为0。因概率的值域为$[0,1]$,这样的设定比线性回归合理很多。
# 常用的替代函数为Sigmoid函数,即:
# $$
# h(z) = \frac{1}{1+e^{-z}}
# $$
# 其中,$z = \theta^T x$
# 我们可以看到,当z大于0时,函数大于0.5;当函数等于0时,函数等于0.5;函数小于0时,函数小于0.5。如果用函数表示目标分到某一类的概率,我们可以采用以下“单位阶跃函数”来判断数据的类别:
# $$h(z) = \left\{
# \begin{aligned}
# 0,& & z<0 \\
# 0.5, & & z=0 \\
# 1, & & z>0
# \end{aligned}
# \right.$$
# 若Z大于0,则判断为正例;若小于0,判断为反例;若等于0,可任意判别。由于Sigmoid函数单调且可导,且fu
# ## 3、逻辑回归损失函数推导及优化
# $$
# P(y=1|x;\theta) = h_\theta (x) \\
# P(y=0|x;\theta) = 1-h_\theta (x)
# $$
# 可以写作一般公式,
# $$P(y|x;\theta)= h(x)^y (1-h(x))^{(1-y)}$$
# 极大似然函数为,
# $$
# L(\theta) = \prod^{m}_{i=1}h_\theta (x^{(i)})^{y^{(i)}} (1-h_\theta (x^{(i)})^{(1-y^{(i)})}
# $$
# 对数极大似然函数为,
# $$l(\theta) = log L(\theta) = \sum^{m}_{i=1} y^{(i)}log h_\theta (x^{(i)}) + (1-y^{(i)})log (1-h_\theta (x^{(i)}))
# $$
# 损失函数为,
# $$
# J(\theta) = -\frac{1}{m}l(\theta) = -\frac{1}{m}\sum^{m}_{i=1} y^{(i)}h_\theta (x^{(i)}) + (1-y^{(i)})(1-h_\theta (x^{(i)}))
# $$
# 损失函数表示了预测值和真实值之间的差异程度,预测值和真实值越接近,则损失函数越小。
# 我们用梯度下降法求解,
# $$\theta:=\theta-\alpha\Delta_\theta J(\theta) = \theta + \frac{\alpha}{m}\Delta_\theta l(\theta)$$
# 由于$g'_\theta(z) = g_\theta (z)(1-g_\theta(z))$
# <td bgcolor=#87CEEB>小练习:试证明,当$g(z) = \frac{1}{1+e^{-z}}$</td>
# <td bgcolor=#87CEEB>$g'(z) = g(z)(1-g(z))$</td>
# 证明:
# \begin{align*}
# g'(z)
# &= \frac{-1}{(1+e^{-z})^2}(-e^{-z}) \\
# &= \frac{e^{-z}}{(1+e^{-z})^2} \\
# &= \frac{1}{1+e^{-z}} \cdot \frac{e^{-z}}{1+e^{-z}} \\
# &= g(z)(1-g(z))
# \end{align*}
# 因此可以求得,
# \begin{align*}
# \frac{\partial J(\theta)}{\partial \theta_i}
# &= \frac{1}{m}\sum^m_{i=1} y^{(i)}\frac{1}{h_\theta (x^{(i)})} h_\theta (x^{(i)})(1-h_\theta (x^{(i)}))x^{(i)} + (1-y^{(i)})\frac{1}{1-h_\theta (x^{(i)})}h_\theta (x^{(i)})(h_\theta (x^{(i)})-1)\\
# &= \frac{1}{m}\sum^m_{i=1}y^{(i)}(1-h_\theta (x^{(i)}))x^{(i)}+(y^{(i)}-1)h_\theta (x^{(i)})x^{(i)} \\
# & = \frac{1}{m}\sum^m_{i=1}(y^{(i)} - h_\theta (x^{(i)}))x^{(i)}
# \end{align*}
# ## 4、正则化与模型评估指标
# #### 正则化
# 我们可以在损失函数后面,加上正则化函数,即$\theta$的惩罚项,来抑制过拟合问题。
# 我们考虑L2正则,在损失函数后面加一项和$\theta$相关的函数,令
# $$
# J(\theta) =\frac{1}{m}\sum^{m}_{i=1} y^{(i)}h_\theta (x^{(i)}) + (1-y^{(i)})(1-h_\theta (x^{(i)})) + \frac{\lambda}{2m}\sum^m_{i=1}\theta_i^2
# $$
# $$
# \Delta_{\theta_i} l(\theta) = \frac{1}{m}\sum^m_{i=1}(y^{(i)} - h_\theta (x^{(i)}))x^{(i)} + \frac{\lambda}{m}\theta_i
# $$
# 用正则化后的函数替代原先的函数进行梯度法迭代获得最优化条件下的参数
# #### 逻辑回归的评价指标
# 由于逻辑回归模型属于分类模型,不能用线性回归的评价指标。
# 二元分类的评价指标基本都适用于逻辑回归。
# 观察以下混淆矩阵,
# 
# 我们可以用查准率和查全率来评价预测结果:
# * 查准率 $P =\frac{TP}{TP+FP}$
# * 查全率 $R =\frac{TP}{TP+FN}$
# 我们可以用P-R曲线表示查准率和查全率之间的关系:
# 查准率和查全率经常相互矛盾,一般查准率高时查全率低,查全率高时查准率低。我们经常针对具体的应用场景决定更看重哪一个指标。
# <td bgcolor=#87CEEB>小练习:试举例说明,在什么场景下,更看重查准率;在什么情况下,更看重查全率,并说明原因。</td>
# <img src = "https://ask.qcloudimg.com/http-save/developer-news/4b1eek0aoz.jpeg?imageView2/2/w/1620"></img>
# P-R曲线越靠外,则预测效果越好,如果两条P-R曲线相交,则难说孰优孰率,可以计算曲线包住的面积。
# 我们可以用$F_\beta$表达对查准率/查全率的不同偏好,定义为:
# $$F_\beta = \frac{(1+\beta^2) \cdot P \cdot R}{(\beta^2 \cdot P)+R}$$
# $\beta$大于1时,查全率有更大影响;$\beta$小于1时,查准率有更大影响;$\beta$等于0时,即标准的F1度量。
# 但是,当正负样本分布发生变化时,P-R曲线会受到较大影响。试想负样本的数量扩大10倍,FP和TN都会成倍增加,会影响到查准率和查全率。这时候我们可以用ROC曲线来评价模型。
#
#
# 定义:
# * $TPR = \frac{TP}{TP+FN}$
# * $FPR = \frac{FP}{TN+FP}$
# 绘图的过程可以将样本的预测概率排序,将分类阈值从最小向最大移动,则每个阈值会得到一组TPR和FPR的位置点,将所有的点连结,则绘制成ROC曲线,同时,计算曲线下的面积,即AUC值。AUC值越大,则说明预测的效果越好。<br>
# <img src = "https://upload-images.jianshu.io/upload_images/11525720-dd2545eaaaa7c2ba.png?imageMogr2/auto-orient/strip|imageView2/2/w/1200/format/webp" width = 500></img>
# ## 5、逻辑回归的优缺点
# * 优点:从上面介绍已经可以额看到,逻辑回归的思想简洁,可以很好的解决二问题。
# * 缺点:
#
# 观察下图
# <img src = "https://img-blog.csdnimg.cn/2019083014370448.png?x-oss-process=image/watermark,type_ZmFuZ3poZW5naGVpdGk,shadow_10,text_aHR0cHM6Ly9ibG9nLmNzZG4ubmV0L2RwZW5nd2FuZw==,size_16,color_FFFFFF,t_70"></img>
# 因为预测结果呈Z字型(或反Z字型),因此当数据集中在中间区域时,对概率的变化会很敏感,可能使得预测结果缺乏区分度。
#
# 第二,由于逻辑回归依然是线性划分,对于非线性的数据集适应性弱。
#
# 第三,当特征空间很大,性能欠佳。
#
# 第四,只能处理两分类问题。
#
# ## 6、样本不均衡问题
# 每200封邮件里有1封垃圾邮件,当碰到一封新邮件时,我们只需将其预测为正常邮件,可以达到99.5%的准确率。但是这样的学习器毫无价值,因为他无法判断出任意一封垃圾邮件。
# 我们预测时,实际上是用预测出的概率值与一个阈值进行比较,例如当y > 0.5时,判断为正例。$\frac{y}{1-y}$表示了正例可能性和反例可能性的比值。阈值为0.5时,即$\frac{y}{1-y}>1$时,预测为正例。
# 如果令$m_+$为样本正例数,$m_-$为样本负例数,随机的观测几率为$\frac{m_+}{m_-}$。只要分类器的预测概率高于观测几率,应判定为正例,即
# $$
# \frac{y}{1-y}>\frac{m_+}{m_-},预测为正例
# $$
# 这时候,需要对预测值进行调整,使得$\frac {y'}{1-y'}=\frac{y}{1-y} \cdot\frac{m_+}{m_-}$,那么,0.5的阈值依然是合理的分类决策。
# 这就是类别不平衡的一个基本策略——“再缩放”。
# “再缩放”的三类做法:
# * 欠采样:去除一些反例使得正反例数目接近。
# * 过采样:增加一些正例使得正反例数目接近。
# * 阈值移动:基于原始集学习,当在预测是,将决策阈值改为符合样本正负样本比例的值。
# 可以想到,过采样因为增加了数据,时间开销大于欠采样法。但欠采样法由于随机丢弃反例,可能丢失一些重要信息。这时候可以将反例划分成若干个集合分别学习,从全局来看并非丢失重要信息。
# ## 7、sklearn参数
# Logistics Regression参数名称
#
# 函数调用形式
#
# LogisticRegression(penalty='l2',dual=False,tol=1e-4,C=1.0,fit_intercept=True,intercept_scaling=1,class_weight=None,random_state=None,solver='liblinear',max_iter=100,multi_class='ovr',verbose=0,warm_start=False, n_jobs=1)
#
# #### penalty
#
# 字符串型,’l1’ or ‘l2’,默认:’l2’;正则化类型。
#
# #### dual
#
# 布尔型,默认:False。当样本数>特征数时,令dual=False;用于liblinear解决器中L2正则化。
#
# #### tol
#
# 浮点型,默认:1e-4;迭代终止判断的误差范围。
#
# #### C
#
# 浮点型,默认:1.0;其值等于正则化强度的倒数,为正的浮点数。数值越小表示正则化越强。
#
# #### fit_intercept
#
# 布尔型,默认:True;指定是否应该向决策函数添加常量(即偏差或截距)。
#
# #### intercept_scaling
#
# 浮点型,默认为1;仅仅当solver是”liblinear”时有用。
#
# #### class_weight
#
# 默认为None;与“{class_label: weight}”形式中的类相关联的权重。如果不给,则所有的类的权重都应该是1。
#
# #### random_state
#
# 整型,默认None;当“solver”==“sag”或“liblinear”时使用。在变换数据时使用的伪随机数生成器的种子。如果是整数, random_state为随机数生成器使用的种子;若为RandomState实例,则random_state为随机数生成器;如果没有,随机数生成器就是' np.random '使用的RandomState实例。
#
# #### solver
#
# {'newton-cg', 'lbfgs', 'liblinear', 'sag', 'saga'},默认: 'liblinear';用于优化问题的算法。
#
# 对于小数据集来说,“liblinear”是个不错的选择,而“sag”和'saga'对于大型数据集会更快。
#
# 对于多类问题,只有'newton-cg', 'sag', 'saga'和'lbfgs'可以处理多项损失;“liblinear”仅限于“one-versus-rest”分类。
#
# #### max_iter
#
# 最大迭代次数,整型,默认是100;
#
# #### multi_class
#
# 字符串型,{ovr', 'multinomial'},默认:'ovr';如果选择的选项是“ovr”,那么一个二进制问题适合于每个标签,否则损失最小化就是整个概率分布的多项式损失。对liblinear solver无效。
#
# #### verbose
#
# 整型,默认是0;对于liblinear和lbfgs solver,verbose可以设为任意正数。
#
# #### warm_start
#
# 布尔型,默认为False;当设置为True时,重用前一个调用的解决方案以适合初始化。否则,只擦除前一个解决方案。对liblinear解码器无效。
#
# #### n_jobs
#
# 整型,默认是1;如果multi_class='ovr' ,则为在类上并行时使用的CPU核数。无论是否指定了multi_class,当将' solver ' '设置为'liblinear'时,将忽略此参数。如果给定值为-1,则使用所有核。
#
# 原文链接:https://blog.csdn.net/qq_38683692/article/details/82533460
# ## 8、代码实现
# #### 1、先尝试调用sklearn的线性回归模型训练数据,尝试以下代码,画图查看分类的结果
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# df_X = pd.read_csv('http://cs229.stanford.edu/ps/ps1/logistic_x.txt', sep='\ +', header=None, engine='python')
# ys = pd.read_csv('http://cs229.stanford.edu/ps/ps1/logistic_y.txt', sep='\ +', header=None, engine='python')
df_X = pd.read_csv('./logistic_x.txt', sep='\ +',header=None, engine='python')
ys = pd.read_csv('./logistic_y.txt', sep='\ +',header=None, engine='python')
ys = ys.astype(int)
df_X['label'] = ys[0].values
# +
ax = plt.axes()
df_X.query('label == 0').plot.scatter(x=0, y=1, ax=ax, color='blue')
df_X.query('label == 1').plot.scatter(x=0, y=1, ax=ax, color='red')
# -
Xs = df_X[[0, 1]].values
Xs = np.hstack([np.ones((Xs.shape[0], 1)), Xs])
ys = df_X['label'].values
# +
from __future__ import print_function
import numpy as np
from sklearn.linear_model import LogisticRegression
import mlflow
import mlflow.sklearn
lr = LogisticRegression(fit_intercept=False)
lr.fit(Xs, ys)
score = lr.score(Xs, ys)
print("Coefficient: %s" % lr.coef_)
print("Score: %s" % score)
mlflow.log_metric("score", score)
mlflow.sklearn.log_model(lr, "model")
print("Model saved in run %s" % mlflow.active_run().info.run_uuid)
# +
ax = plt.axes()
df_X.query('label == 0').plot.scatter(x=0, y=1, ax=ax, color='blue')
df_X.query('label == 1').plot.scatter(x=0, y=1, ax=ax, color='red')
_xs = np.array([np.min(Xs[:,1]), np.max(Xs[:,1])])
# for k, theta in enumerate(all_thetas):
# _ys = (theta[0] + theta[1] * _xs) / (- theta[2])
# plt.plot(_xs, _ys, label='iter {0}'.format(k + 1), lw=0.5)
_ys = (lr.coef_[0][0] + lr.coef_[0][1] * _xs) / (- lr.coef_[0][2])
plt.plot(_xs, _ys, lw=1)
plt.legend(bbox_to_anchor=(1.04,1), loc="upper left")
# -
# #### 2 用梯度下降法将相同的数据分类,画图和sklearn的结果相比较
# +
class LGR_GD():
def __init__(self):
self.w = None
def fit(self,X,y,alpha=0.001,loss = 1e-10): # 设定步长为0.001,判断是否收敛的条件为1e-10
y = y.reshape(-1,1) #重塑y值的维度以便矩阵运算
[m,d] = np.shape(X) #自变量的维度
self.w = np.zeros((1,d)) #将参数的初始值定为0
tol = 1e5
n_iters =0
while tol > loss:
zs = X.dot(self.w.T)
h_f = 1 / (1 + np.exp(-zs))
theta = self.w + alpha *np.sum(X*(y - h_f),axis=0) #计算迭代的参数值
tol = np.sum(np.abs(theta - self.w))
self.w = theta
n_iters += 1
self.w = theta
def predict(self, X):
# 用已经拟合的参数值预测新自变量
y_pred = X.dot(self.w)
return y_pred
if __name__ == "__main__":
lgr_gd = LGR_GD()
lgr_gd.fit(Xs,ys)
# +
ax = plt.axes()
df_X.query('label == 0').plot.scatter(x=0, y=1, ax=ax, color='blue')
df_X.query('label == 1').plot.scatter(x=0, y=1, ax=ax, color='red')
_xs = np.array([np.min(Xs[:,1]), np.max(Xs[:,1])])
_ys = (lr_gd.w[0][0] + lr_gd.w[0][1] * _xs) / (- lr_gd.w[0][2])
plt.plot(_xs, _ys, lw=1)
plt.legend(bbox_to_anchor=(1.04,1), loc="upper left")
# -
# ## 参考:
# 吴恩达 CS229课程
# 周志华 《机器学习》
# https://blog.csdn.net/dpengwang/article/details/100159369
# https://blog.csdn.net/u014106644/article/details/83660226
# https://blog.csdn.net/abcjennifer/article/details/7716281
# https://blog.csdn.net/portfloat/article/details/79200695
#
# https://cloud.tencent.com/developer/news/319664
# https://www.jianshu.com/p/2ca96fce7e81
# https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LogisticRegression.html
# https://blog.csdn.net/yoggieCDA/article/details/88953206
# https://blog.csdn.net/ustbclearwang/article/details/81235892
# https://blog.csdn.net/qq_38683692/article/details/82533460
| origin_data/Task2_logistic_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
def suffixArray(s):
""" Given T return suffix array SA(T). We use Python's sorted
function here for simplicity, but we can do better. """
satups = sorted([(s[i:], i) for i in range(len(s))])
# Extract and return just the offsets
# print(satups)
return map(lambda x: x[1], satups)
def bwtViaSa(t):
""" Given T, returns BWT(T) by way of the suffix array. """
bw = []
for si in suffixArray(t):
if si == 0: bw.append('$')
else: bw.append(t[si-1])
return ''.join(bw) # return string-ized version of list bw
list(suffixArray('ABAABA'))
bwtViaSa('ABAABA$')
bwtViaSa('CTGCTGCTGCTGAACTG$')
| notebooks/05_Burrows_Wheeler_with_SA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import sys
# add the 'src' directory as one where we can import modules
# os.pardir refers to parent directory.
# src_dir = os.path.join(os.getcwd(), '../src') => works the same
src_dir = os.path.join(os.getcwd(), os.pardir, 'src')
sys.path.append(src_dir)
# Load the "autoreload" extension
# %load_ext autoreload
# always reload modules marked with "%aimport"
# %autoreload 2
# +
# Experiment 1
# %run ../src/models/predict_model.py --data ../data/test/ndr_blog.tsv.gz --output ../data/results/propap_1.tsv --feat tfidf --vtr ../models/propap_vtr_1 --clf ../models/propap_clf_1
# -
# Experiment 2
# %run ../src/models/train_model.py --data ../data/train/partisan-gov-large.2-label.tsv.gz --preprocess True --feat tfidf --model nb --model_params '{"alpha": 0.001}' --vtr ../models/propap_vtr_2 --clf ../models/propap_clf_2
# Experiment 3
# %run ../src/models/train_model.py --data ../data/train/partisan-gov-large.2-label.tsv.gz --preprocess True --stop_words True --feat tfidf --model nb --model_params '{"alpha": 0.001}' --vtr ../models/propap_vtr_3 --clf ../models/propap_clf_3
# +
# Experiment 4
# %run ../src/models/train_model.py --data ../data/train/partisan-gov-large.2-label.tsv.gz --feat count --model nb --model_params '{"alpha":0.009}' --vtr ../models/propap_vtr_4 --clf ../models/propap_clf_4
# -
# Experiment 5
# %run ../src/models/train_model.py --data ../data/train/partisan-gov-large.2-label.tsv.gz --preprocess True --feat count --model nb --model_params '{"alpha":0.009}' --vtr ../models/propap_vtr_5 --clf ../models/propap_clf_5
# Experiment 6
# %run ../src/models/train_model.py --data ../data/train/partisan-gov-large.2-label.tsv.gz --preprocess True --stop_words True --feat count --model nb --model_params '{"alpha":0.009}' --vtr ../models/propap_vtr_6 --clf ../models/propap_clf_6
# +
# Experiment 7
# %run ../src/models/train_model.py --data ../data/train/partisan-gov-large.2-label.tsv.gz --feat count --model lr --model_params '{"C": 1, "penalty":"l2"}' --vtr ../models/propap_vtr_7 --clf ../models/propap_clf_7
# -
# Experiment 8
# %run ../src/models/train_model.py --data ../data/train/partisan-gov-large.2-label.tsv.gz --feat count --preprocess True --model lr --model_params '{"C": 1, "penalty":"l2"}' --vtr ../models/propap_vtr_8 --clf ../models/propap_clf_8
# Experiment 9
# %run ../src/models/train_model.py --data ../data/train/partisan-gov-large.2-label.tsv.gz --preprocess True --stop_words True --feat count --model lr --model_params '{"C": 1, "penalty":"l2"}' --vtr ../models/propap_vtr_9 --clf ../models/propap_clf_9
# +
# Experiment 10
# %run ../src/models/train_model.py --data ../data/train/partisan-gov-large.2-label.tsv.gz --feat tfidf --model lr --model_params '{"C": 1, "penalty":"l2"}' --vtr ../models/propap_vtr_10 --clf ../models/propap_clf_10
# -
# Experiment 11
# %run ../src/models/train_model.py --data ../data/train/partisan-gov-large.2-label.tsv.gz --preprocess True --feat tfidf --model lr --model_params '{"C": 1, "penalty":"l2"}' --vtr ../models/propap_vtr_11 --clf ../models/propap_clf_11
# Experiment 12
# %run ../src/models/train_model.py --data ../data/train/partisan-gov-large.2-label.tsv.gz --preprocess True --stop_words True --feat tfidf --model lr --model_params '{"C": 1, "penalty":"l2"}' --vtr ../models/propap_vtr_12 --clf ../models/propap_clf_12
# Experiment 13
# %run ../src/models/train_model.py --data ../data/train/partisan-gov-large.2-label.tsv.gz --preprocess True --feat dict --emotion_words ../resources/emotion_words --lsd_words ../resources/LSD2011_ALL.txt --word_feature True --ngram 14 --fiveWoneH 111111 --pos True --model nb --model_params '{"alpha":0.001}' --num_topk_features 1000 --vtr ../models/propap_vtr_13 --clf ../models/propap_clf_13
# +
# Experiment 14
# %run ../src/models/train_model.py --data ../data/train/partisan-gov-large.2-label.tsv.gz --preprocess True --feat dict --emotion_words ../resources/emotion_words --lsd_words ../resources/LSD2011_ALL.txt --word_feature True --ngram 14 --fiveWoneH 111111 --pos True --model lr --model_params '{"C":1,"penalty":"l1"}' --num_topk_features 1000 --vtr ../models/propap_vtr_14 --clf ../models/propap_clf_14
# +
# Experiment 15
# %run ../src/models/train_model.py --data ../data/train/partisan-gov-large.2-label.tsv.gz --preprocess True --feat dict --model lr --model_params '{"C":1,"penalty":"l1"}' --ngrams 13 --vtr ../models/propap_vtr_15 --clf ../models/propap_clf_15
# -
# Experiment 16
# %run ../src/models/train_model.py --data ../data/train/partisan-gov-large.2-label.tsv.gz --preprocess True --feat dict --model nb --model_params '{"alpha":0.001}' --ngrams 13 --vtr ../models/propap_vtr_16 --clf ../models/propap_clf_16
# Experiment 17
# %run ../src/models/train_model.py --data ../data/train/partisan-gov-large.2-label.tsv.gz --preprocess True --feat dict --model svc --model_params '{"kernel":"linear", "C":1}' --ngrams 13 --vtr ../models/propap_vtr_17 --clf ../models/propap_clf_17
# Experiment 18
# %run ../src/models/train_model.py --data ../data/train/partisan-gov-large.2-label.tsv.gz --preprocess True --feat dict --emotion_words ../resources/emotion_words --lsd_words ../resources/LSD2011_ALL.txt --word_feature True --ngram 13 --fiveWoneH 111111 --pos True --model svc --model_params '{"C": 1, "kernel":"linear"}' --num_topk_features 1500 --vtr ../models/propap_vtr_18 --clf ../models/propap_clf_18
| notebooks/propap_pred.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Visualising Data
# ================
#
# The purpose of scientific computation is insight not numbers: To understand the meaning of the (many) numbers we compute, we often need postprocessing, statistical analysis and graphical visualisation of our data. The following sections describe
#
# - Matplotlib/Pylab — which allows us to generate high quality graphs of the type *y* = *f*(*x*) (and a bit more)
#
# - Visual Python — which is a very handy tool to quickly generate animations of time dependent processes taking place in 3d space.
#
#
# ## What is matplotlib?
#
# Matplotlib is the most popular and mature library for plotting data using
# Python. It has all of the functionality you would expect, including the ability to control
# the formatting of plots and figures at a very fine level.
#
# The official matplotlib documentation is at http://matplotlib.org/
# The matplotlib gallery is at http://matplotlib.org/gallery.html
# ### Importing Matplotlib
#
# Just as we use the ``np`` shorthand for NumPy and the ``pd`` shorthand for Pandas, we will use some standard shorthands for Matplotlib imports:
import matplotlib as mpl
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
# The ``plt`` interface is what we will use most often, as we shall see throughout this chapter.
# ### First Plot
plt.plot(np.random.normal(size=200), np.random.normal(size=200), 'o')
# ### ``show()`` or No ``show()``? How to Display Your Plots
# A visualization you can't see won't be of much use, but just how you view your Matplotlib plots depends on the context.
#
# #### Plotting from an IPython notebook
#
# Plotting interactively within an IPython notebook can be done with the ``%matplotlib`` command, and works in a similar way to the IPython shell.
# In the IPython notebook, you also have the option of embedding graphics directly in the notebook, with two possible options:
#
# - ``%matplotlib notebook`` will lead to *interactive* plots embedded within the notebook
# - ``%matplotlib inline`` will lead to *static* images of your plot embedded in the notebook
#
# For this book, we will generally opt for ``%matplotlib inline``:
# %matplotlib inline
# After running this command (it needs to be done only once per kernel/session), any cell within the notebook that creates a plot will embed a PNG image of the resulting graphic:
plt.plot(np.random.normal(size=200), np.random.normal(size=200), 'd')
# ## Plotting in Pandas
#
# On the other hand, Pandas includes methods for DataFrame and Series objects that are relatively high-level, and that make reasonable assumptions about how the plot should look.
normals = pd.Series(np.random.normal(size=30))
print(normals)
normals.plot()
# Notice that by default a line plot is drawn, and a light grid is included. All of this can be changed, however:
normals.cumsum().plot(grid=True)
# Similarly, for a DataFrame:
# +
variables = pd.DataFrame({'normal': np.random.normal(size=10),
'gamma': np.random.gamma(1, size=10),
'poisson': np.random.poisson(size=10)})
print(variables)
variables.cumsum(0).plot()
# -
# As an illustration of the high-level nature of Pandas plots, we can split multiple series into subplots with single argument per `plot`:
variables.cumsum(0).plot(subplots=True)
# Or, we may want to have some series displayed on the secondary y-axis, which can allow for greater detail and less empty space:
variables.cumsum(0).plot(secondary_y='normal')
# If we would prefer slightly more control, we can use matplotlib's `subplots` function directly, and manually assign plots to its axes:
fig, axes = plt.subplots(nrows=3, ncols=1, figsize=(4, 12))
for i,var in enumerate(['normal','gamma','poisson']):
variables[var].cumsum(0).plot(ax=axes[i], title=var)
axes[0].set_ylabel('cumulative sum')
# ### Saving Figures to File
#
# One pleasing feature of Matplotlib is the ability to save figures in a wide variety of formats.
# Saving a figure can be done using the ``savefig()`` command.
# For example, to save the previous figure as a PNG file, you can run this:
fig.savefig('figure.png')
# We now have a file called ``my_figure.png`` in the current working directory:
# !ls -lh figure.png
# To confirm that it contains what we think it contains, let's use the IPython ``Image`` object to display the contents of this file:
from IPython.display import Image
Image('figure.png')
# In ``savefig()``, the file format is inferred from the extension of the given filename.
# Depending on what you have installed in the backend, various distinctive file formats are available.
# The list of supported file types can be found for your system by using the following method of the figure canvas object:
fig.canvas.get_supported_filetypes()
# Note that when saving your figure, it's not necessary to use ``plt.show()`` or related commands discussed earlier.
# #### Object-oriented interface
#
# The object-oriented interface is available for these complicated situations, and whenever there is needa of slightly more control over your figure.
# Rather than depending on some notion of an "active" figure or axes, in the object-oriented interface the plotting functions are *methods* of explicit ``Figure`` and ``Axes`` objects.
# # Simple Line Plots
# Perhaps the simplest of all plots is the visualization of a single function $y = f(x)$.
# Here we will take a first look at creating a simple plot of this type.
# +
fig = plt.figure()
ax = plt.axes()
s = np.linspace(0, 20, 100)
print(s)
ax.plot(s, np.sin(s));
# -
# Alternatively, we can use the pylab interface and let the figure and axes be created for us in the background:
plt.plot(s, np.sin(s))
# That's all there is to plotting simple functions in Matplotlib!
# We'll now dive into some more details about how to control the appearance of the axes and lines.
# ## Line Colors and Styles
# The first adjustment you might wish to make to a plot is to control the line colors and styles.
# The ``plt.plot()`` function takes additional arguments that can be used to specify these.
# To adjust the color, you can use the ``color`` keyword, which accepts a string argument representing virtually any imaginable color.
# The color can be specified in a variety of ways:
plt.plot(s, np.sin(s - 0), color='red')
plt.plot(s, np.sin(s - 1), color='y')
plt.plot(s, np.sin(s - 2), color='0.55')
# If no color is specified, Matplotlib will automatically cycle through a set of default colors for multiple lines.
#
# Similarly, the line style can be adjusted using the ``linestyle`` keyword:
plt.plot(s, np.sin(s - 0), linestyle='-')
plt.plot(s, np.sin(s - 1), linestyle=':')
plt.plot(s, np.sin(s - 2), linestyle='-.')
# ## Adjusting the Plot: Axes Limits
#
# Matplotlib does a decent job of choosing default axes limits for your plot, but sometimes it's advised to have finer control.
# The most basic way to adjust axis limits is to use the ``plt.xlim()`` and ``plt.ylim()`` methods:
# +
plt.plot(s, np.sin(s - 0))
plt.xlim(-1, 15)
plt.ylim(-2, 2);
# -
# For more information on axis limits and other capabilities of the ``plt.axis`` method, refer to the ``plt.axis`` docstring.
# ## Labeling Plots
#
# As the last piece of this section, we'll briefly look at the labelling of plots: titles, axis labels, and simple legends.
#
# Titles and axis labels are the simplest such labels—there are methods that can be used to quickly set them:
plt.plot(s, np.cos(s))
plt.title("Cosine Curve")
plt.xlabel("x")
plt.ylabel("cos(x)")
# The position, size, and style of these labels can be adjusted using optional arguments to the function.
# For more information, see the Matplotlib documentation and the docstrings of each of these functions.
# # Simple Scatter Plots
# Another commonly used plot type is the simple scatter plot, a close cousin of the line plot.
# Instead of points being joined by line segments, here the points are represented individually with a dot, circle, or other shape.
# We’ll start by setting up the notebook for plotting and importing the functions that we use:
plt.style.use('seaborn-whitegrid')
# ## Scatter Plots with ``plt.plot``
#
# In the previous section we looked at ``plt.plot``/``ax.plot`` to produce line plots.
# It turns out that this same function can produce scatter plots as well:
# +
x = np.linspace(1, 10, 50)
y = np.log(x)
plt.plot(x, y, 'o-', color='green')
# -
# For even more possibilities, these character codes can be used together with line and color codes to plot points along with a line connecting them:
plt.plot(x, y, '-or');
# Additional keyword arguments to ``plt.plot`` specify a wide range of properties of the lines and markers:
# ## Scatter Plots with ``plt.scatter``
#
# A second, more powerful method of creating scatter plots is the ``plt.scatter`` function, which can be used very similarly to the ``plt.plot`` function:
plt.scatter(x, y, marker='o')
# The primary difference of ``plt.scatter`` from ``plt.plot`` is that it can be used to create scatter plots where the properties of each individual point (size, face color, edge color, etc.) can be individually controlled or mapped to data.
#
# Let's show this by creating a random scatter plot with points of many colors and sizes.
# In order to better see the overlapping results, we'll also use the ``alpha`` keyword to adjust the transparency level:
# +
rng = np.random.RandomState(7)
x = rng.randn(100)
y = rng.randn(100)
colors = rng.rand(100)
sizes = 1000 * rng.rand(100)
plt.scatter(x, y, c=colors, s=sizes, alpha=0.5,
cmap='Reds')
plt.colorbar() # show color scale
# -
# Notice that the color argument is automatically mapped to a color scale (shown here by the ``colorbar()`` command), and that the size argument is given in pixels.
# This way, the color and size of points can be used to convey information through visualization, in order to visualize multidimensional data.
#
# For example, we might use the Iris data from Scikit-Learn, where each sample is one of three types of flowers that has had the size of its petals and sepals carefully measured:
# ## ``plot`` Versus ``scatter``: A Note on Efficiency
#
# Apart from the different features available in ``plt.plot`` and ``plt.scatter``, why one might you choose to use one over the other? While it doesn't matter much for small amounts of data, as datasets get larger than a few thousand points, ``plt.plot`` can be noticeably more efficient than ``plt.scatter``.
# The reason is that ``plt.scatter`` has the capability to render different size and/or color for each point, so the renderer must do the extra work of constructing each point individually.
# In ``plt.plot``, on the other hand, the points are always essentially clones of each other, so the work of determining the appearance of the points is done only once for the entire set of data.
# For large datasets, the difference between these two can lead to vastly different performance, and for this reason, ``plt.plot`` should be preferred over ``plt.scatter`` for large datasets.
# # Visualizing Errors
# For any scientific measurement, accurate accounting of errors is nearly as important, if not more important, than accurate reporting of the number itself.
# For example, imagine that I am using some astrophysical observations to estimate the Hubble Constant, the local measurement of the expansion rate of the Universe.
# I know that the current literature suggests a value of around 71 (km/s)/Mpc, and I measure a value of 74 (km/s)/Mpc with my method. Are the values consistent? The only correct answer, given in this information, is this: there is no way to know.
#
# Suppose I augment this information with reported uncertainties: the current literature suggests a value of around 71 $\pm$ 2.5 (km/s)/Mpc, and my method has measured a value of 74 $\pm$ 5 (km/s)/Mpc. Now are the values consistent? That is a question that can be quantitatively answered.
#
# In visualization of data and results, showing these errors effectively can make a plot convey much more complete information.
# ## Errorbars
#
# A basic errorbar can be created with a single Matplotlib function call:
# +
x = np.linspace(0, 10, 100)
dy = 0.1
y = np.sin(x) + dy * np.random.randn(100)
plt.errorbar(x, y, yerr=dy, fmt='.b');
# -
# In addition to these basic options, the ``errorbar`` function has many options to fine-tune the outputs.
# Using these additional options you can easily customize the aesthetics of your errorbar plot.
# I often find it helpful, especially in crowded plots, to make the errorbars lighter than the points themselves:
# ## Histograms
# A simple histogram can be a great first step in understanding a dataset.
# +
plt.style.use('seaborn-white')
data = np.random.randn(10000)
print(data)
# -
plt.hist(data)
# The ``hist()`` function has many options to tune both the calculation and the display;
# Here's an example of a more customized histogram:
plt.hist(data, bins=5, normed=True, alpha=0.6,
histtype='stepfilled')
# The ``plt.hist`` docstring has more information on other customization options available.
# I find this combination of ``histtype='stepfilled'`` along with some transparency ``alpha`` to be very useful when comparing histograms of several distributions:
# +
x = np.random.normal(0, 0.8, 1000)
y = np.random.normal(-2, 1, 1000)
kwargs = dict(histtype='stepfilled', alpha=0.5, normed=True, bins=50)
plt.hist(x, **kwargs)
plt.hist(y, **kwargs)
# -
# If you would like to simply compute the histogram (that is, count the number of points in a given bin) and not display it, the ``np.histogram()`` function is available:
counts, bin_edges = np.histogram(data, bins=5)
print(counts)
# ## Two-Dimensional Histograms
#
# Just as we create histograms in one dimension by dividing the number-line into bins, we can also create histograms in two-dimensions by dividing points among two-dimensional bins.
# We'll take a brief look at several ways to do this here.
# We'll start by defining some data—an ``x`` and ``y`` array drawn from a multivariate Gaussian distribution:
mean = [5, 2]
cov = [[3, 1], [2, 0.5]]
x, y = np.random.multivariate_normal(mean, cov, 10000)
# ### ``plt.hist2d``: Two-dimensional histogram
#
# One straightforward way to plot a two-dimensional histogram is to use Matplotlib's ``plt.hist2d`` function:
plt.hist2d(x, y, bins=20, cmap='Reds')
cb = plt.colorbar()
# ### ``plt.hexbin``: Hexagonal binnings
#
# The two-dimensional histogram creates a tesselation of squares across the axes.
# Another natural shape for such a tesselation is the regular hexagon.
# For this purpose, Matplotlib provides the ``plt.hexbin`` routine, which will represents a two-dimensional dataset binned within a grid of hexagons:
# +
plt.hexbin(x, y, gridsize=20, cmap='Reds')
cb = plt.colorbar()
# -
# ``plt.hexbin`` has a number of interesting options, including the ability to specify weights for each point, and to change the output in each bin to any NumPy aggregate (mean of weights, standard deviation of weights, etc.).
| CODE_FILES/AiRobosoft/Data Analysis/Matplotlib.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <center><h1> Predicting Wine Quality by using Watson Machine Learning</h1></center>
#
# ### <NAME>
#
# <p>This notebook contains steps and code to create a predictive model to predict Wine Quality and then deploy that model to Watson Machine Learning so it can be used in an application.</p>
#
# ## Learning Goals
# The learning goals of this notebook are:
# * Load a CSV file into the Object Storage Service linked to my Data Science Experience
# * Create an Pandas machine learning model
# * Train and evaluate a model
# * Persist a model in a Watson Machine Learning repository
#
# ## 1. Setup
#
# Before you use the sample code in this notebook, you must perform the following setup tasks:
# * Create a Watson Machine Learning Service instance (a free plan is offered) and associate it with your project
# * Upload wine quality data to the Object Store service that is part of your data Science Experience trial
#
# ## 2. Load and explore data
# <p>In this section load the data as Pandas DataFrame and perform a basic exploration.</p>
#
# <p>Load the data to the Pandas DataFrame from the associated Object Storage instance.</p>
# +
import types
import pandas as pd
from ibm_botocore.client import Config
import ibm_boto3
def __iter__(self): return 0
# @hidden_cell
# The following code accesses a file in your IBM Cloud Object Storage. It includes your credentials.
# You might want to remove those credentials before you share your notebook.
client_8c14de1d42f947c2812d9d92c44fc593 = ibm_boto3.client(service_name='s3',
ibm_api_key_id='<KEY>',
ibm_auth_endpoint="https://iam.bluemix.net/oidc/token",
config=Config(signature_version='oauth'),
endpoint_url='https://s3-api.us-geo.objectstorage.service.networklayer.com')
body = client_8c14de1d42f947c2812d9d92c44fc593.get_object(Bucket='default-donotdelete-pr-ijbvuypsfmigjt',Key='winequality-red.csv')['Body']
# add missing __iter__ method, so pandas accepts body as file-like object
if not hasattr(body, "__iter__"): body.__iter__ = types.MethodType( __iter__, body )
red = pd.read_csv(body)
red.head()
body = client_8c14de1d42f947c2812d9d92c44fc593.get_object(Bucket='default-donotdelete-pr-ijbvuypsfmigjt',Key='winequality-white.csv')['Body']
# add missing __iter__ method, so pandas accepts body as file-like object
if not hasattr(body, "__iter__"): body.__iter__ = types.MethodType( __iter__, body )
white = pd.read_csv(body)
white.head()
# -
# Explore the loaded data by using the following Pandas DataFrame methods:
# * print dataframe info
# * print head records
# * count tail records
# * print sample records
# * describe dataframe
# * check isnull values in dataframe
# +
# Print info on white wine
print(white.info())
# Print info on red wine
print(red.info())
# -
# First rows of `red`
red.head()
# Last rows of `white`
white.tail()
# Take a sample of 5 rows of `red`
red.sample(5)
# Describe `white`
white.describe()
# Double check for null values in `red`
pd.isnull(red)
# ## 3 Interactive Visualizations with Matplotlib and Numpy
#
# ### 3.1: Visualize Alcohol vs Frequency
# Distribution of Alcohol in % Vol for red and white wines.
# +
import matplotlib.pyplot as plt
fig, ax = plt.subplots(1, 2)
ax[0].hist(red.alcohol, 10, facecolor='red', alpha=0.5, label="Red wine")
ax[1].hist(white.alcohol, 10, facecolor='white', ec="black", lw=0.5, alpha=0.5, label="White wine")
fig.subplots_adjust(left=0, right=1, bottom=0, top=0.5, hspace=0.05, wspace=1)
ax[0].set_ylim([0, 1000])
ax[0].set_xlabel("Alcohol in % Vol")
ax[0].set_ylabel("Frequency")
ax[1].set_xlabel("Alcohol in % Vol")
ax[1].set_ylabel("Frequency")
fig.suptitle("Distribution of Alcohol in % Vol")
plt.show()
# -
# ### 3.2: Print histograms of alcohol using Numpy
# Histogram of alcohol for red and white wines.
import numpy as np
print(np.histogram(red.alcohol, bins=[7,8,9,10,11,12,13,14,15]))
print(np.histogram(white.alcohol, bins=[7,8,9,10,11,12,13,14,15]))
# ### 3.3: Visualize Quality vs Sulphates
# Quality vs Sulphates for red and white wines.
# +
import matplotlib.pyplot as plt
fig, ax = plt.subplots(1, 2, figsize=(8, 4))
ax[0].scatter(red['quality'], red["sulphates"], color="red")
ax[1].scatter(white['quality'], white['sulphates'], color="white", edgecolors="black", lw=0.5)
ax[0].set_title("Red Wine")
ax[1].set_title("White Wine")
ax[0].set_xlabel("Quality")
ax[1].set_xlabel("Quality")
ax[0].set_ylabel("Sulphates")
ax[1].set_ylabel("Sulphates")
ax[0].set_xlim([0,10])
ax[1].set_xlim([0,10])
ax[0].set_ylim([0,2.5])
ax[1].set_ylim([0,2.5])
fig.subplots_adjust(wspace=0.5)
fig.suptitle("Wine Quality by Amount of Sulphates")
plt.show()
# -
# ### 3.4: Visualize Quality vs Volatile Acidity vs Alcohol
# Quality vs volatile acidity vs alcohol for red and white wines.
# +
import matplotlib.pyplot as plt
import numpy as np
np.random.seed(570)
redlabels = np.unique(red['quality'])
whitelabels = np.unique(white['quality'])
import matplotlib.pyplot as plt
fig, ax = plt.subplots(1, 2, figsize=(8, 4))
redcolors = np.random.rand(6,4)
whitecolors = np.append(redcolors, np.random.rand(1,4), axis=0)
for i in range(len(redcolors)):
redy = red['alcohol'][red.quality == redlabels[i]]
redx = red['volatile acidity'][red.quality == redlabels[i]]
ax[0].scatter(redx, redy, c=redcolors[i])
for i in range(len(whitecolors)):
whitey = white['alcohol'][white.quality == whitelabels[i]]
whitex = white['volatile acidity'][white.quality == whitelabels[i]]
ax[1].scatter(whitex, whitey, c=whitecolors[i])
ax[0].set_title("Red Wine")
ax[1].set_title("White Wine")
ax[0].set_xlim([0,1.7])
ax[1].set_xlim([0,1.7])
ax[0].set_ylim([5,15.5])
ax[1].set_ylim([5,15.5])
ax[0].set_xlabel("Volatile Acidity")
ax[0].set_ylabel("Alcohol")
ax[1].set_xlabel("Volatile Acidity")
ax[1].set_ylabel("Alcohol")
#ax[0].legend(redlabels, loc='best', bbox_to_anchor=(1.3, 1))
ax[1].legend(whitelabels, loc='best', bbox_to_anchor=(1.3, 1))
#fig.suptitle("Alcohol - Volatile Acidity")
fig.subplots_adjust(top=0.85, wspace=0.7)
plt.show()
# -
# ## 4. Create Pandas machine learning model
# In this section I prepare data, create and train Pandas machine learning model.
#
# ### 4.1: Prepare data
# In this subsection data is joined and prepared: labels are separated from the features.
# Append `white` to `red`
wines = red.append(white, ignore_index=True)
wines.shape
# +
# Isolate target labels
y = wines.quality
# Isolate data
X = wines.drop('quality', axis=1)
# -
# ### 4.2: Visualize data using Seaborn heatmap
# Heatmap
import seaborn as sns
corr = wines.corr()
sns.heatmap(corr,
xticklabels=corr.columns.values,
yticklabels=corr.columns.values,
cmap="YlGnBu")
# ### 4.3: Preprocess Data
# Standardize features by removing the mean and scaling to unit variance
# +
# Import `StandardScaler` from `sklearn.preprocessing`
from sklearn.preprocessing import StandardScaler
import numpy as np
# Scale the data with `StandardScaler`
X = StandardScaler().fit_transform(X)
# -
# ### 4.4: Creating model
# Creating model using K fold validation partitions
# +
import numpy as np
from sklearn.model_selection import StratifiedKFold
from keras.models import Sequential
from keras.layers import Dense
from keras.optimizers import SGD, RMSprop
seed = 7
np.random.seed(seed)
kfold = StratifiedKFold(n_splits=5, shuffle=True, random_state=seed)
for train, test in kfold.split(X, y):
model = Sequential()
model.add(Dense(128, input_dim=11, activation='relu'))
model.add(Dense(1))
# rmsprop = RMSprop(lr=0.001)
sgd=SGD(lr=0.01)
model.compile(optimizer=sgd, loss='mse', metrics=['mae'])
# model.compile(optimizer='rmsprop', loss='mse', metrics=['mae'])
model.fit(X[train], y[train], epochs=10, verbose=1)
# -
# ### 4.5: Evaluate model
# Evaluate model by checking Mean Squared Error (MSE) and the Mean Absolute Error (MAE) and R2 score or the regression score function
# +
mse_value, mae_value = model.evaluate(X[test], y[test], verbose=0)
print(mse_value)
print(mae_value)
# -
from sklearn.metrics import r2_score
y_pred = model.predict(X[test])
r2_score(y[test], y_pred)
y_pred
# Model output shape
model.summary()
# ### 4.6: Compare the results
# Check the predictions
y_pred = y_pred.astype(int)
predictions = np.column_stack((y[test], y_pred));
print(predictions[:15])
# ## 5. Persist model
# Import client libraries.
wml_credentials={
"url": "https://ibm-watson-ml.mybluemix.net",
"username": "0faaa0df-0f3a-4aa7-835d-8929d6cda36e",
"password": "<PASSWORD>",
"instance_id": "2074d379-2141-42dc-9340-a78bf626de46"
}
from watson_machine_learning_client import \
WatsonMachineLearningAPIClient
client = WatsonMachineLearningAPIClient(wml_credentials)
h5file = 'winemodel-keras.h5'
gzfile = 'winemodel-keras.tar.gz'
model.save(h5file)
import tarfile
with tarfile.open(gzfile, 'w:gz') as tf:
tf.add(h5file)
metadata = {
client.repository.ModelMetaNames.NAME: 'Wine Quality Prediction Model',
client.repository.ModelMetaNames.FRAMEWORK_NAME: 'tensorflow',
client.repository.ModelMetaNames.FRAMEWORK_VERSION: '1.3',
client.repository.ModelMetaNames.RUNTIME_NAME: 'python',
client.repository.ModelMetaNames.RUNTIME_VERSION: '3.5',
client.repository.ModelMetaNames.FRAMEWORK_LIBRARIES: [{'name':'keras', 'version': '2.1.3'}]
}
published_model = client.repository.store_model(model=gzfile, meta_props=metadata)
published_model
# ## 6. Load model to verify that it was saved correctly
# You can load your model to make sure that it was saved correctly.
# +
import json
import requests
from base64 import b64encode
token_url = service_path + "/v3/identity/token"
# NOTE: for python 2.x, uncomment below, and comment out the next line of code:
#userAndPass = b64encode(bytes(username + ':' + password)).decode("ascii")
# Use below for python 3.x, comment below out for python 2.x
userAndPass = b64encode(bytes(username + ':' + password, "utf-8")).decode("ascii")
headers = { 'Authorization' : 'Basic %s' % userAndPass }
response = requests.request("GET", token_url, headers=headers)
watson_ml_token = json.loads(response.text)['token']
print(watson_ml_token)
# -
# ### 6.2 Preview currenly published models
# +
model_url = service_path + "/v3/wml_instances/" + instance_id + "/published_models"
headers = {'authorization': 'Bearer ' + watson_ml_token }
response = requests.request("GET", model_url, headers=headers)
published_models = json.loads(response.text)
print(json.dumps(published_models, indent=2))
# -
# Read the details of any returned models
print('{} model(s) are available in your Watson ML Service'.format(len(published_models['resources'])))
for model in published_models['resources']:
print('\t- name: {}'.format(model['entity']['name']))
print('\t model_id: {}'.format(model['metadata']['guid']))
print('\t deployments: {}'.format(model['entity']['deployments']['count']))
# Create a new deployment of the Model
# +
# Update this `model_id` with the model_id from model that you wish to deploy listed above.
model_id = '213f75a7-406f-4c9d-b1ca-6f07edc7bbca'
deployment_url = service_path + "/v3/wml_instances/" + instance_id + "/published_models/" + model_id + "/deployments"
payload = "{\"name\": \"Wine Quality Prediction Model Deployment\", \"description\": \"First deployment of Wine Quality Prediction Model\", \"type\": \"online\"}"
headers = {'authorization': 'Bearer ' + watson_ml_token, 'content-type': "application/json" }
response = requests.request("POST", deployment_url, data=payload, headers=headers)
print(response.text)
# +
deployment = json.loads(response.text)
print('Model {} deployed.'.format(model_id))
print('\tname: {}'.format(deployment['entity']['name']))
print('\tdeployment_id: {}'.format(deployment['metadata']['guid']))
print('\tstatus: {}'.format(deployment['entity']['status']))
print('\tscoring_url: {}'.format(deployment['entity']['scoring_url']))
# -
# ### Monitor the status of deployment
# +
# Update this `deployment_id` from the newly deployed model from above.
deployment_id = "7de2f5a3-44a1-439d-ada8-ce9e4c73eb60"
deployment_details_url = service_path + "/v3/wml_instances/" + instance_id + "/published_models/" + model_id + "/deployments/" + deployment_id
headers = {'authorization': 'Bearer ' + watson_ml_token, 'content-type': "application/json" }
response = requests.request("GET", deployment_url, headers=headers)
print(response.text)
# +
deployment_details = json.loads(response.text)
for resources in deployment_details['resources']:
print('name: {}'.format(resources['entity']['name']))
print('status: {}'.format(resources['entity']['status']))
print('scoring url: {}'.format(resources['entity']['scoring_url']))
# -
# ## 6.3 Invoke prediction model deployment
# Define a method to call scoring url. Replace the **scoring_url** in the method below with the scoring_url returned from above.
def get_prediction_ml(fa, va, ca, rs, ch, fsd, tsd, d, p, s, a):
scoring_url = 'https://ibm-watson-ml.mybluemix.net/v3/wml_instances/2074d379-2141-42dc-9340-a78bf626de46/deployments/7de2f5a3-44a1-439d-ada8-ce9e4c73eb60/online'
scoring_payload = { "fields":["fixed acidity", "volatile acidity", "citric acid", "residual sugar", "chlorides", "free sulfur dioxide", "total sulfur dioxide", "density", "pH", "sulphates", "alcohol"],"values":[[fa, va, ca, rs, ch, fsd, tsd, d, p, s, a]]}
header = {'authorization': 'Bearer ' + watson_ml_token, 'content-type': "application/json" }
scoring_response = requests.post(scoring_url, json=scoring_payload, headers=header)
print(scoring_response.text)
return (json.loads(scoring_response.text).get("values")[0][0])
# +
data = np.array([7, 0.27, 0.36 ,20.7 ,0.045 ,45 ,170 ,1.001 ,3 ,0.45 ,8.8]).reshape(-1, 1)
data = StandardScaler().fit_transform(data)
data.T
# print('What is quality of wine with such characteristics?: {}'.format(get_prediction_ml(0.45, 3.28, -2.19, -0.59, 1.19, -0.31, -0.86, 0.7, -0.11, 0.99, -0.58)))
# -
| WinePredictKeras.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Problem 2
#
# Now that we have built a dataframe of the postal code of each neighborhood along with the borough name and neighborhood name, in order to utilize the Foursquare location data, we need to get the latitude and the longitude coordinates of each neighborhood.
# +
# Import libraries
import pandas as pd
from bs4 import BeautifulSoup
import requests
# Website url
url = 'https://en.wikipedia.org/wiki/List_of_postal_codes_of_Canada:_M'
# Scrapping data from thw website
website_script = requests.get(url) # Website script (download the HTML content)
website_content = website_script.content # Website content (HTML content)
# Make HTML look Beautiful
website_soup = BeautifulSoup(website_content, 'html.parser')
# Get Toronto neighborhood dataframe
def get_toronto_neighborhood_df(soup, table_class):
# Table data
table = soup.find_all('table', class_=table_class)
# Table dataframe
df = pd.read_html(str(table))[0]
# Remove rows where Borough is 'Not assigned'
df = df[df['Borough'] != 'Not assigned']
# Sort ascending values
df.sort_values(by=['Postal Code'], ascending=True, inplace=True)
# Return dataframe
return df
# Get result dataframe
def get_result_df(neighborhood_df):
# Latitude and Longitude dataframe
lat_lng_coords_df = pd.read_csv('https://cocl.us/Geospatial_data')
# The result of both dataframe
result_df = pd.merge(neighborhood_df, lat_lng_coords_df, on="Postal Code")
# Rename Postal Code and Neighborhood column
result_df.rename(columns={"Neighbourhood": "Neighborhood", "Postal Code": "PostalCode"}, inplace=True)
# Reset index
result_df.reset_index(drop=True, inplace=True)
# Return result dataframe
return result_df
# Dataframe
toronto_neighborhood_df = get_toronto_neighborhood_df(website_soup, 'wikitable sortable') # Toronto neighborhood Dataframe
result_df = get_result_df(toronto_neighborhood_df) # Result dataframe
# -
# Dataframe output
print(result_df.head(12))
| toronto_neighborhood_geographical_coordinates.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Fiddling about a bit
# +
# imports
from pkg_resources import resource_filename
from astropy.table import Table
from astropy.coordinates import SkyCoord
from astropy import units as u
# -
# ## Load up
DM_file = resource_filename('pulsars', 'data/atnf_cat/DM_cat_v1.56.dat')
DMs = Table.read(DM_file, format='ascii')
DMs
# ### Coords
coords = SkyCoord(ra=DMs['RAJ'], dec=DMs['DECJ'], unit=(u.hourangle, u.deg))
# ## Clouds
# ### Manchester+06
mfl = DMs['Pref'] == 'mfl+06'
DMs[mfl]
# ### LMC coords
lmc_distance = 50 * u.kpc
lmc_coord = SkyCoord('J052334.6-694522', unit=(u.hourangle, u.deg),
distance=lmc_distance)
lmc_coord.separation(coords[mfl]).to('deg').value
# ### Others
close_to_lmc = lmc_coord.separation(coords) < 3*u.deg
DMs[close_to_lmc]
| doc/nb/Fiddling_about.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # WeatherPy
# ----
#
# #### Note
# * Instructions have been included for each segment. You do not have to follow them exactly, but they are included to help you think through the steps.
# +
import gmaps
import json
import requests
from api_keys import api_key
from api_keys import g_key
import random
import pandas as pd
import numpy as np
import json
import matplotlib.pyplot as plt
from scipy.stats import linregress
# Access maps with unique API key
gmaps.configure(api_key=g_key)
# -
# ## Generate Cities List
# +
# Import cities file as DataFrame
#cities_pd = pd.read_csv("worldcities.csv")
cities_pd = pd.read_csv("cities.csv")
cities_pd.head(100)
# -
# ### Perform API Calls
# * Perform a weather check on each city using a series of successive API calls.
# * Include a print log of each city as it'sbeing processed (with the city number and city name).
#
# +
url = "http://api.openweathermap.org/data/2.5/weather?"
#cities = cities_pd["city_ascii"]
#api.openweathermap.org/data/2.5/weather?lat={lat}&lon={lon}&appid={your api key}
cities = cities_pd["City"]
cntry = cities_pd["Country"]
lat = cities_pd["Lat"]
lng = cities_pd["Lng"]
temper = cities_pd["Max Temp"]
hum = cities_pd["Humidity"]
cloud = cities_pd["Cloudiness"]
speed = cities_pd["Wind Speed"]
nor_lat = []
nor_hum = []
nor_temper = []
nor_cloud = []
nor_speed = []
sou_lat = []
sou_hum = []
sou_temper = []
sou_cloud = []
sou_speed = []
units = "metric"
impl = "imperial"
query_url = f"{url}appid={api_key}&units={impl}&q="
# -
# ### Convert Raw Data to DataFrame
# * Export the city data into a .csv.
# * Display the DataFrame
# +
# Get the indices of cities that have humidity over 100%.
# Make a new DataFrame equal to the city data to drop all humidity outliers by index.
# Passing "inplace=False" will make a copy of the city_data DataFrame, which we call "clean_city_data".
#by default all humidity are less than 100
for index, row in cities_pd.iterrows():
try:
if (row["Lat"] >= 0 ):
nor_lat.append(row['Lat'])
nor_temper.append(row['Max Temp'])
nor_hum.append(row['Humidity'])
nor_speed.append(row['Wind Speed'])
nor_cloud.append(row['Cloudiness'])
else:
sou_lat.append(row['Lat'])
sou_temper.append(row['Max Temp'])
sou_hum.append(row['Humidity'])
sou_speed.append(row['Wind Speed'])
sou_cloud.append(row['Cloudiness'])
except:
pass
# -
cities_pd.head(100)
# ## Inspect the data and remove the cities where the humidity > 100%.
# ----
# Skip this step if there are no cities that have humidity > 100%.
complete_wea_dict = {
"lat": lat,
"lng": lng,
"temper": temper,
"hum": hum,
"cloud": cloud,
"speed": speed
}
complete_wea_dict_data = pd.DataFrame(complete_wea_dict)
complete_wea_dict_data
complete_nor_wea_dict = {
"nor_lat": nor_lat,
"nor_hum": nor_hum,
"nor_temper": nor_temper,
"nor_cloud": nor_cloud,
"nor_speed": nor_speed
}
complete_nor_wea_dict_data = pd.DataFrame(complete_nor_wea_dict)
complete_nor_wea_dict_data
complete_sou_wea_dict = {
"sou_lat": sou_lat,
"sou_hum": sou_hum,
"sou_temper": sou_temper,
"sou_cloud": sou_cloud,
"sou_speed": sou_speed
}
complete_sou_wea_dict_data = pd.DataFrame(complete_sou_wea_dict)
complete_sou_wea_dict_data
# Get the indices of cities that have humidity over 100%.
humd_over_more = complete_wea_dict_data.loc[complete_wea_dict_data["hum"] >100]
humd_over_more
# Make a new DataFrame equal to the city data to drop all humidity outliers by index.
# Passing "inplace=False" will make a copy of the city_data DataFrame, which we call "clean_city_data".
complete_wea_dict_data
# ## Plotting the Data
# * Use proper labeling of the plots using plot titles (including date of analysis) and axes labels.
# * Save the plotted figures as .pngs.
# ## Latitude vs. Temperature Plot
#This is for temperature vs latitude
x_limit = 100
x_axis = lat
data = temper
plt.scatter(x_axis, data, marker="o", facecolors="red", edgecolors="black",
s=x_axis, alpha=0.75)
plt.ylim(30, 100)
plt.xlim(20,80)
plt.xlabel(" Latitude ")
plt.ylabel(" Temperature in ")
plt.savefig("../output_img/temp_vs_latitude.png")
plt.show()
# ## Latitude vs. Humidity Plot
# +
#This is for humidity vs latitude
x_limit = 100
x_axis = lat
data = hum
plt.scatter(x_axis, data, marker="o", facecolors="blue", edgecolors="black",
s=x_axis, alpha=0.75)
plt.ylim(0, 110)
plt.xlim(20,80)
plt.xlabel(" Latitude ")
plt.ylabel(" Humidity in degree")
plt.savefig("../output_img/humidity_vs_latitude.png")
plt.show()
# -
# ## Latitude vs. Cloudiness Plot
# +
#This is for cloudiness vs latitude
x_limit = 100
x_axis = lat
data = cloud
plt.scatter(x_axis, data, marker="o", facecolors="blue", edgecolors="black",
s=x_axis, alpha=0.75)
plt.ylim(-1, 80)
plt.xlim(0,110)
plt.xlabel(" latitude ")
plt.ylabel(" cloudiness in degree")
plt.savefig("../output_img/cloudiness_vs_latitude.png")
plt.show()
# -
# ## Latitude vs. Wind Speed Plot
# +
#This is for windspeed(mph) vs latitude
x_limit = 100
x_axis = lat
data = speed
plt.scatter(x_axis, data, marker="o", facecolors="yellow", edgecolors="black",
s=x_axis, alpha=0.75)
plt.ylim(0, 50)
plt.xlim(0,80)
plt.xlabel(" Latitude in degree ")
plt.ylabel(" Windspeed in mph")
plt.savefig("../output_img/windspeed_vs_latitude.png")
plt.show()
# -
# ## Linear Regression
# +
weather_dict = {
"lat": nor_lat,
"temp": nor_temper
}
weather_data = pd.DataFrame(weather_dict)
max_temp = weather_data["temp"]
lat_data = weather_data["lat"]
x_values = lat_data
y_values = max_temp
(wea_slope, wea_intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
wea_regress_values = x_values* wea_slope + wea_intercept
line_eq = "y = " + str(round(wea_slope,2)) + "x + " + str(round(wea_intercept,2))
plt.scatter(x_values,y_values)
plt.plot(x_values,wea_regress_values,"r-")
plt.annotate(line_eq,(6,10),fontsize=15,color="red")
plt.xlabel('The Latitude')
plt.ylabel('Max(Temperature) in Fahrenheit')
plt.savefig("../output_img/northern_max_temp_vs_latitude.png")
plt.show()
print(f' The linear regression is {line_eq}')
# -
#This is for temperature vs latitude
x_limit = 100
x_axis = nor_lat
data = nor_temper
plt.scatter(x_axis, data, marker="o", facecolors="red", edgecolors="black",
s=x_axis, alpha=0.75)
plt.ylim(30, 100)
plt.xlim(0,70)
plt.xlabel(" Latitude ")
plt.ylabel(" Humidity in ")
plt.savefig("../output_img/northern_temp_vs_latitude.png")
plt.show()
# +
# max temperature vs latitude
x_limit = 100
x_axis = lat_data
data = max_temp
plt.scatter(x_axis, data, marker="o", facecolors="blue", edgecolors="black",
s=x_axis, alpha=0.75)
plt.ylim(0, 110)
plt.xlim(-10,80)
plt.xlabel(" latitude ")
plt.ylabel("max temperature in farhenheit")
plt.show()
# -
# #### Northern Hemisphere - Max Temp vs. Latitude Linear Regression
#Max temperature vs Latitude, the linear regression is provided as print statement in Northen hemisphere
weather_dict = {
"lat": nor_lat,
"temp": nor_temper
}
weather_data = pd.DataFrame(weather_dict)
max_temp = weather_data["temp"]
lat_data = weather_data["lat"]
x_values = lat_data
y_values = max_temp
(wea_slope, wea_intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
wea_regress_values = x_values* wea_slope + wea_intercept
line_eq = "y = " + str(round(wea_slope,2)) + "x + " + str(round(wea_intercept,2))
plt.scatter(x_values,y_values)
plt.plot(x_values,wea_regress_values,"r-")
plt.annotate(line_eq,(6,10),fontsize=15,color="red")
plt.xlabel('The Latitude')
plt.ylabel('Max(Temperature) in Fahrenheit')
plt.savefig("../output_img/northern_max_temp_vs_latitude.png")
plt.show()
print(f"The r-squared is: {rvalue**2}")
print(f' The linear regression is {line_eq}')
# #### Southern Hemisphere - Max Temp vs. Latitude Linear Regression
# +
#Max temperature vs Latitude, the linear regression is provided as print statement for southern hemisphere
weather_dict = {
"lat": sou_lat,
"temp": sou_temper
}
weather_data = pd.DataFrame(weather_dict)
max_temp = weather_data["temp"]
lat_data = weather_data["lat"]
x_values = lat_data
y_values = max_temp
(wea_slope, wea_intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
wea_regress_values = x_values* wea_slope + wea_intercept
line_eq = "y = " + str(round(wea_slope,2)) + "x + " + str(round(wea_intercept,2))
plt.scatter(x_values,y_values)
plt.plot(x_values,wea_regress_values,"r-")
plt.annotate(line_eq,(6,10),fontsize=15,color="red")
plt.xlabel('The Latitude')
plt.ylabel('Max(Temperature) in Fahrenheit')
plt.savefig("../output_img/southern_max_temp_vs_latitude.png")
plt.show()
print(f"The r-squared is: {rvalue**2}")
print(f' The linear regression is {line_eq}')
# -
# #### Northern Hemisphere - Humidity (%) vs. Latitude Linear Regression
#Max humidty vs Latitude, the linear regression is provided as print statement in Northen hemisphere
weather_dict = {
"lat": nor_lat,
"temp": nor_hum
}
weather_data = pd.DataFrame(weather_dict)
max_temp = weather_data["temp"]
lat_data = weather_data["lat"]
x_values = lat_data
y_values = max_temp
(wea_slope, wea_intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
wea_regress_values = x_values* wea_slope + wea_intercept
line_eq = "y = " + str(round(wea_slope,2)) + "x + " + str(round(wea_intercept,2))
plt.scatter(x_values,y_values)
plt.plot(x_values,wea_regress_values,"r-")
plt.annotate(line_eq,(6,10),fontsize=15,color="red")
plt.xlabel('The Latitude')
plt.ylabel('Max(Humidity) ')
plt.savefig("../output_img/northern_max_humidity_vs_latitude.png")
plt.show()
print(f"The r-squared is: {rvalue**2}")
print(f' The linear regression is {line_eq}')
# #### Southern Hemisphere - Humidity (%) vs. Latitude Linear Regression
#Max humidty vs Latitude, the linear regression is provided as print statement in Southern hemisphere
weather_dict = {
"lat": sou_lat,
"temp": sou_hum
}
weather_data = pd.DataFrame(weather_dict)
max_temp = weather_data["temp"]
lat_data = weather_data["lat"]
x_values = lat_data
y_values = max_temp
(wea_slope, wea_intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
wea_regress_values = x_values* wea_slope + wea_intercept
line_eq = "y = " + str(round(wea_slope,2)) + "x + " + str(round(wea_intercept,2))
plt.scatter(x_values,y_values)
plt.plot(x_values,wea_regress_values,"r-")
plt.annotate(line_eq,(6,10),fontsize=15,color="red")
plt.xlabel('The Latitude')
plt.ylabel('Max(Humidity) ')
plt.savefig("../output_img/southern_max_humidity_vs_latitude.png")
plt.show()
print(f"The r-squared is: {rvalue**2}")
print(f' The linear regression is {line_eq}')
# #### Northern Hemisphere - Cloudiness (%) vs. Latitude Linear Regression
#Max Cloudiness vs Latitude, the linear regression is provided as print statement in Northen hemisphere
weather_dict = {
"lat": nor_lat,
"temp": nor_cloud
}
weather_data = pd.DataFrame(weather_dict)
max_temp = weather_data["temp"]
lat_data = weather_data["lat"]
x_values = lat_data
y_values = max_temp
(wea_slope, wea_intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
wea_regress_values = x_values* wea_slope + wea_intercept
line_eq = "y = " + str(round(wea_slope,2)) + "x + " + str(round(wea_intercept,2))
plt.scatter(x_values,y_values)
plt.plot(x_values,wea_regress_values,"r-")
plt.annotate(line_eq,(6,10),fontsize=15,color="red")
plt.xlabel('The Latitude')
plt.ylabel('Max(Cloudiness) ')
plt.savefig("../output_img/northern_max_cloudiness_vs_latitude.png")
plt.show()
print(f"The r-squared is: {rvalue**2}")
print(f' The linear regression is {line_eq}')
# #### Southern Hemisphere - Cloudiness (%) vs. Latitude Linear Regression
#Max Cloudiness vs Latitude, the linear regression is provided as print statement in Southern hemisphere
weather_dict = {
"lat": sou_lat,
"temp": sou_cloud
}
weather_data = pd.DataFrame(weather_dict)
max_temp = weather_data["temp"]
lat_data = weather_data["lat"]
x_values = lat_data
y_values = max_temp
(wea_slope, wea_intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
wea_regress_values = x_values* wea_slope + wea_intercept
line_eq = "y = " + str(round(wea_slope,2)) + "x + " + str(round(wea_intercept,2))
plt.scatter(x_values,y_values)
plt.plot(x_values,wea_regress_values,"r-")
plt.annotate(line_eq,(6,10),fontsize=15,color="red")
plt.xlabel('The Latitude')
plt.ylabel('Max(Cloudiness) ')
plt.savefig("../output_img/southern_max_cloudiness_vs_latitude.png")
plt.show()
print(f"The r-squared is: {rvalue**2}")
print(f' The linear regression is {line_eq}')
# #### Northern Hemisphere - Wind Speed (mph) vs. Latitude Linear Regression
#Max wind speed vs Latitude, the linear regression is provided as print statement in Northen hemisphere
weather_dict = {
"lat": nor_lat,
"temp": nor_speed
}
weather_data = pd.DataFrame(weather_dict)
max_temp = weather_data["temp"]
lat_data = weather_data["lat"]
x_values = lat_data
y_values = max_temp
(wea_slope, wea_intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
wea_regress_values = x_values* wea_slope + wea_intercept
line_eq = "y = " + str(round(wea_slope,2)) + "x + " + str(round(wea_intercept,2))
plt.scatter(x_values,y_values)
plt.plot(x_values,wea_regress_values,"r-")
plt.annotate(line_eq,(6,10),fontsize=15,color="red")
plt.xlabel('The Latitude')
plt.ylabel('Max(wind speed) ')
plt.savefig("../output_img/northern_max_windspeed_vs_latitude.png")
plt.show()
print(f"The r-squared is: {rvalue**2}")
print(f' The linear regression is {line_eq}')
# #### Southern Hemisphere - Wind Speed (mph) vs. Latitude Linear Regression
#Max wind speed vs Latitude, the linear regression is provided as print statement in Southern hemisphere
weather_dict = {
"lat": sou_lat,
"temp": sou_speed
}
weather_data = pd.DataFrame(weather_dict)
max_temp = weather_data["temp"]
lat_data = weather_data["lat"]
x_values = lat_data
y_values = max_temp
(wea_slope, wea_intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
wea_regress_values = x_values* wea_slope + wea_intercept
line_eq = "y = " + str(round(wea_slope,2)) + "x + " + str(round(wea_intercept,2))
plt.scatter(x_values,y_values)
plt.plot(x_values,wea_regress_values,"r-")
plt.annotate(line_eq,(6,10),fontsize=15,color="red")
plt.xlabel('The Latitude')
plt.ylabel('Max(wind speed) ')
plt.savefig("../output_img/southern_max_windspeed_vs_latitude.png")
plt.show()
print(f"The r-squared is: {rvalue**2}")
print(f' The linear regression is {line_eq}')
| WeatherPy/WeatherPy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/devpatelio/dr.cnn/blob/main/TransferLearningCNN.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="p8El4NHrVgHc"
import torch
import torchvision
import torch.nn as nn
import torch.nn.functional as F
import torchvision.transforms as transforms
from __future__ import print_function, division
import os
import pandas as pd
from skimage import io, transform
import numpy as np
import matplotlib.pyplot as plt
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
# Ignore warnings
import warnings
warnings.filterwarnings("ignore")
plt.ion() # interactive mode
# + colab={"base_uri": "https://localhost:8080/"} id="YLTVQ_pGVpUe" outputId="dedb159b-cfd4-4b87-d024-27e1e7f510fe"
from google.colab import drive
drive.mount('/content/drive')
# + id="PQCDELvh5-Qj"
trainer_names_csv = pd.read_csv('/content/drive/MyDrive/pyTorch/diabetic-retinopathy-224x224-gaussian-filtered/train.csv')
n = len(trainer_names_csv)
# data_labels = pd.DataFrame(data = trainer_names_csv)
img_names = trainer_names_csv.iloc[:n, 0]
label = trainer_names_csv.iloc[:n, 1]
# + id="iExvWEjk6Cvu"
from torch.utils.data import Dataset
import pandas as pd
import os
from PIL import Image
import torch
class DRDataset(Dataset):
def __init__ (self, csv_file, root_dir, transform=None):
"""
csv_file = labels
root_dir = images
"""
self.train_data_csv = pd.read_csv(csv_file)
self.root_dir = root_dir
self.transform = transform
def __len__(self):
return len(self.train_data_csv)
def __getitem__ (self, idx):
name = self.train_data_csv.iloc[idx, 0]
img_name = os.path.join(self.root_dir, name +'.png') ##fix logic
image = Image.open(img_name).convert("RGB")
y_labels = torch.tensor(float(self.train_data_csv.iloc[idx, 1:]))
if self.transform is not None:
image = self.transform(image)
return (image, y_labels)
# + colab={"base_uri": "https://localhost:8080/"} id="yH5Pj5uB6QZu" outputId="65a07f11-1592-417b-ddf4-513564721a70"
from tqdm import tqdm
import random
transform = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225]),
])
num_epochs = 10
learning_rate = 0.01
train_CNN = True
batch_size = 10
shuffle = True
pin_memory = True
num_workers = 0
train_size = int(0.7 * n)
validation_size = n - train_size
# image_dataset = DRDataset(csv_file='/content/drive/MyDrive/pyTorch/diabetic-retinopathy-224x224-gaussian-filtered/train.csv', root_dir='/content/drive/MyDrive/pyTorch/diabetic-retinopathy-224x224-gaussian-filtered/gaussian_filtered_images/gaussian_filtered_images/')
dataset = DRDataset(csv_file='/content/drive/MyDrive/pyTorch/diabetic-retinopathy-224x224-gaussian-filtered/train.csv', root_dir='/content/drive/MyDrive/pyTorch/diabetic-retinopathy-224x224-gaussian-filtered/gaussian_filtered_images/gaussian_filtered_images/',
transform=transform)
train_set, validation_set = torch.utils.data.random_split(dataset, [train_size, validation_size])
train_loader = torch.utils.data.DataLoader(dataset=train_set, shuffle=shuffle, batch_size=batch_size,num_workers=num_workers, pin_memory=pin_memory)
validation_loader = torch.utils.data.DataLoader(dataset=validation_set, shuffle=shuffle, batch_size=batch_size,num_workers=num_workers, pin_memory=pin_memory)
dataiter = iter(train_loader)
image, label = dataiter.next()
# print(data[0].shape)
# print(data[1].shape)
# print(data[1])
print(image.shape)
print(label.shape)
# + colab={"base_uri": "https://localhost:8080/", "height": 104, "referenced_widgets": ["dcfaa053d71e46ff95d22b4d4bb854a8", "c1106998bf7846ebbf8c253414cae418", "2c87f5499d5b407ea3235599f781a209", "cd5fcc833fc646178d3795470ce774a5", "956d7f7cef45406a9658d6b327f778ad", "140bddef59a540bbbd0a657e9cc170ec", "821e16a3a7e64f6ba73c8fe982dd05f5", "ffb3775c17484af993b382cb01a80a35"]} id="TWjwoykl7AI3" outputId="f0a3d410-19ae-4834-d375-7311f507005d"
import torchvision.models as models
from torch.autograd import Variable
import time
model_vgg = models.vgg16(pretrained=True)
for param in model_vgg.parameters():
param.requires_grad = False
def preconvfeat(dataset):
conv_features = []
labels_list = []
for i, data in enumerate(dataset, 0):
inputs, labels = data
inputs , labels = Variable(inputs),Variable(labels)
x = model_vgg.features(inputs)
conv_features.extend(x.data.cpu().numpy())
labels_list.extend(labels.data.cpu().numpy())
conv_features = np.concatenate([[feat] for feat in conv_features])
return (conv_features,labels_list)
# + colab={"base_uri": "https://localhost:8080/", "height": 240} id="GC-ts2sA68kE" outputId="315c9f86-7815-4039-e04c-0e2913be6c4a"
from __future__ import print_function, division
import torch
import torch.nn as nn
import torch.optim as optim
from torch.optim import lr_scheduler
import numpy as np
import torchvision
from torchvision import datasets, models, transforms
import time
import os
import copy
import cv2
plt.ion()
mean = [0.485, 0.456, 0.406]
std = [0.229, 0.224, 0.225]
data_transforms = {
'train': transforms.Compose([
transforms.RandomResizedCrop(224),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
transforms.Normalize(mean, std)
]),
'val': transforms.Compose([
transforms.Resize(256),
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize(mean, std)
]),
}
train_set, validation_set = torch.utils.data.random_split(dataset, [train_size, validation_size])
image_dataset = {'train': train_set, 'val': validation_set}
dataloaders = {x: torch.utils.data.DataLoader(image_dataset[x], batch_size=batch_size, shuffle=True) for x in ['train', 'val']}
dataset_sizes = {x: len(image_dataset[x]) for x in ['train', 'val']}
device = torch.device("cpu")
def imshow(inp, title):
imshow_mean = np.array(mean)
imshow_std = np.array(std)
inp = (inp.numpy().transpose((1, 2, 0))) * imshow_std + imshow_mean
inp = np.clip(inp, 0, 1)
inp = cv2.resize(inp, (2520, 1420))
plt.imshow(inp)
plt.pause(0.001)
dataiters = iter(dataloaders['train'])
images, labels = dataiters.next()
out = torchvision.utils.make_grid(images, 5) # represents no. of images per row displayed
imshow(out, title=f'{labels}')
def train_model(model, criterion, optimizer, scheduler, num_epochs):
since = time.time()
best_models_wts = copy.deepcopy(model.state_dict())
best_acc = 0.0
for epoch in range(num_epochs):
print(f'Epoch {epoch+1}/{num_epochs}')
print('-' * 10)
for session_type in ['train', 'val']:
if session_type == 'train':
model.train()
elif session_type == 'val':
model.eval()
run_loss, run_corrects = 0.0, 0
for inputs, labels in dataloaders[session_type]:
inputs = inputs.to(device)
labels = labels.to(device)
optimizer.zero_grad()
with torch.requires_grad(True, session_type == 'train'):
outputs = model(inputs)
tensor_pred, long_pred = torch.max(outputs, 1)
loss = criterion(outputs, labels.type(torch.LongTensor))
if session_type == 'train':
loss.backward()
optimizer.step()
run_loss =+ loss.item() * inputs.size(0)
if preds == labels.data:
run_corrects += torch.sum(keepdim=True)
else:
run_corrects += torch.sum(keepdim=False)
if session_type == 'train':
scheduler.step() ## changes the learning rate as per the gradient
epoch_loss = run_loss/dataset_sizes[session_type] #as a loss item
epoch_acc = run_corrects.double()/dataset_sizes[session_type] #as a percent of total correct preds
print('{} Loss: {:.4f} Acc: {:.4f}'.format(session_type, epoch_loss, epoch_acc))
print('{} Learning Rate {:.4f}'.format(session_type, optimizer.param_groups[0]['lr'])) ##returns change in learning rate over each epoch
if session_type == 'val' and epoch_acc > best_acc: #during validation, rewrites best_accuracy with current epoch_accuracy
best_acc = epoch_acc
best_model_wts = copy.deepcopy(model.state_dict()) #copies model and reruns next epoch with new model
time_elapsed = time.time() - since
print('Training complete in {:.0f}m {:.0f}s'.format(time_elapsed//60, time_elapsed % 60))
print('Best val Acc: {:4f}'.format(best_acc))
model.load_state_dict(best_model_wts)
return model
# + id="iIIUmegj7jRP"
def visualize_model(model, num_images=20):
was_training = model.training
model.eval()
images_so_far = 0
fig = plt.figure()
with torch.no_grad():
for i, (inputs, labels) in enumerate(dataloaders['val']):
inputs = inputs.to(device)
labels = labels.to(device)
outputs = model(inputs)
_, preds = torch.max(outputs, 1)
for j in range(inputs.size()[0]):
images_so_far += 1
ax = plt.subplot(num_images//2, 2, images_so_far)
ax.axis('off')
ax.set_title('predicted: {}'.format(class_names[preds[j]]))
imshow(inputs.cpu().data[j])
if images_so_far == num_images:
model.train(mode=was_training)
return
model.train(mode=was_training)
# + colab={"background_save": true, "base_uri": "https://localhost:8080/", "referenced_widgets": ["17e727f8b7b84a5981686631c332bb42", "4851ad1dbb9444c3a3bc6c9f7f6e8f41", "e15dd8e7e22f49da99991060b1ae65f7", "381cbb47bdc94f219c1e277d2df6d1af", "411c2d17a74c41b19b9a36d909515505", "18d70a43f4ce4537a6988e693321fa7b", "14dceb3c55204dbb98f980693bd56bb0", "0d185b8f058b4aa79c7c7e037d5bf480"]} id="Hb2VRI1N7kOX" outputId="3b951764-2f16-42ab-c8a6-bb35fa9ce2c5"
model_ft = models.resnet18(pretrained=True)
num_ftrs = model_ft.fc.in_features
model_ft.fc = nn.Linear(num_ftrs, 5) #output of layer is going to be 5, input is num_ftrs
#in the train_model function, we pre-established the dataset being used -> you can change that easily for new datasets if you wanted
model_ft = model_ft.to(device)
criterion = nn.CrossEntropyLoss()
optimizer_ft = optim.SGD(model_ft.parameters(), lr=0.001, momentum=0.9)
exp_lr_scheduler = lr_scheduler.StepLR(optimizer_ft, step_size=7, gamma=0.1) #reduces lr by 0.1 ever 7 epochs
model_ft = train_model(model_ft, criterion, optimizer_ft, exp_lr_scheduler, num_epochs=25)
# + id="uao_9uUXHa3c"
model = models.resnet18(pretrained=True)
class SaveOutput:
def __init__(self):
self.outputs = []
def __call__(self, module, module_in, module_out):
self.outputs.append(module_out)
def clear(self):
self.outputs = []
save_output = SaveOutput()
hook_handles = []
for layer in model.modules():
if isinstance(layer, torch.nn.modules.conv.Conv2d):
handle = layer.register_forward_hook(save_output.__call__)
hook_handles.append(handle)
image = Image.open('/content/drive/MyDrive/pyTorch/diabetic-retinopathy-224x224-gaussian-filtered/gaussian_filtered_images/gaussian_filtered_images/000c1434d8d7.png')
transform = transforms.Compose([transforms.Resize((224, 224)), transforms.ToTensor()])
X = transform(image).unsqueeze(dim=0).to(device)
out = model(X)
# + id="OJx48h548Jqf" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="0fdb7d2e-f55e-483f-be7b-9f0bcc9d454e"
np_image = save_output.outputs[0].detach().numpy()
with plt.style.context("seaborn-white"):
plt.figure(figsize=(20, 20), frameon=False)
for i in range(16):
plt.subplot(4, 4, i+1)
plt.imshow(np_image[0, idx])
plt.setp(plt.gcf().get_axes(), xticks=[], yticks=[]);
# + id="6gRze4Dhq0jQ"
| TransferLearningCNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 366098, "status": "ok", "timestamp": 1611381721020, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416044610109286808"}, "user_tz": -480} id="2usGK3TYMbsr" outputId="43958b8e-6f82-402f-d49a-9af83b851e59"
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="iqI5L27rHjAg"
# ##Import Packages
# + executionInfo={"elapsed": 3193, "status": "ok", "timestamp": 1611382921252, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416044610109286808"}, "user_tz": -480} id="FpUe6694fq3h"
import time
import numpy as np
import os
import cv2
import random
import matplotlib.pyplot as plt
import pickle
from tqdm.notebook import tqdm
from sklearn.model_selection import train_test_split
from sklearn.decomposition import PCA
import sklearn.metrics as metrics
from sklearn.metrics import plot_confusion_matrix
from sklearn.ensemble import RandomForestClassifier
# + [markdown] id="ciqkz1NoIE9L"
# ##Load Dataset
# + executionInfo={"elapsed": 1519, "status": "ok", "timestamp": 1611382921254, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416044610109286808"}, "user_tz": -480} id="UvxIGvY_5OwA"
# Define labels of the classes and subclasses
CLASSES = ["healthy", "ill"]
SUBCLASSES = ["fever", "sore throat", "running nose"]
# + colab={"base_uri": "https://localhost:8080/", "height": 210, "referenced_widgets": ["3300a2f2c8d6443fbdd4eca227e8360f", "b81fee72fa964c77aef7876478e8948b", "1696019ab3614bea9f42aaecd66a4d0d", "0ce2a67ae1fb43fab0e0c6bc50cf0477", "<KEY>", "9e292248651d473c935145ad1d6f35a8", "ec7589917fef4217ba67c2dd22138915", "0086a79d3296433f8842cca15a6172ba", "c4a5062ab8d54d31a9f376fd4e767316", "99d0c4a7d7bb4c0396470cfded512db2", "306e92b78fde45e297d68932a544cce4", "1b1275b07ff14609a696c00b0d2fb3b9", "eec9511a2a9c4eb1be0f1f7402f9e8f0", "<KEY>", "012178aad1994faf8c0d19b6e71294c1", "<KEY>", "0e280e8faef946579abec72a7d9d90e5", "c0449e58955e4ff7ae47d84fb0e66e4e", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "7f42b636687045be9a9f482abb8672a8", "<KEY>", "<KEY>", "238362c0450949e182ca557d62ce0063", "6208119de5884983af49c944740195aa", "<KEY>", "b448e69a864e42a898c6d14262862169"]} executionInfo={"elapsed": 210031, "status": "ok", "timestamp": 1611383130611, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416044610109286808"}, "user_tz": -480} id="azh7MJDPcS9u" outputId="06e89ed1-35e6-4741-d43f-9d614c1d6262"
folder = "/content/drive/MyDrive/FYP/dataset"
img_array = []
class_num_list1 = []
subcls_num_list1 = ["None"]*500 # fill the list with 500 "None's" first to represent "healthy" samples
for c in CLASSES:
if (c == "healthy"):
path = os.path.join(folder,c) # create path to dataset folder
class_num = 0 # get the CLASSES index, 0 = healthy, 1 = ill
for img in tqdm(os.listdir(path)): # iterate over each image per folder
try:
img = cv2.imread(os.path.join(path,img)) # read image into a list
img_array.append(img)
class_num_list1.append(class_num) # save a list of the class numbers
except Exception as e:
print("error")
elif (c == "ill"):
for s in SUBCLASSES:
path = os.path.join(folder,c,s) # create path to ill subfolders
class_num = 1
subclass_num = SUBCLASSES.index(s)
for img in tqdm(os.listdir(path)): # iterate over each image per folder
try:
img = cv2.imread(os.path.join(path,img)) # read image into a list
img_array.append(img)
class_num_list1.append(class_num)
subcls_num_list1.append(subclass_num) # save a list of the subclass numbers; 0 = fever, 1 = sore throat, 2 = running nose
except Exception as e:
print("error")
# + colab={"base_uri": "https://localhost:8080/", "height": 210, "referenced_widgets": ["828f9fc45d11493fba65c3e63e70f1a1", "<KEY>", "<KEY>", "b5d7c5a061ac41449a063f3b2a2252cd", "f5486e707a024330bac8e7ff3f609d79", "f296fefceaa94bd68ee8b4c6f0721903", "073a6c0760ac4f9abfdaef307e135212", "ec063ba9138c470d82b6eaca8a2d05c3", "9006b8fe9e8d4d39be75a229d2a54507", "72225f7e7d7240feb839e6440137a8fb", "fbe9664fcd874247a105d238142ef10f", "6e69d6ce7ced4f138164df88cf9d5e18", "<KEY>", "1332dd54f4574873b44e214ca5392fcd", "<KEY>", "<KEY>", "a0a4746513d54f288a7c6de91523ebbe", "79b5dbac33e84fe9b4c2a0764db23775", "aa543bbe7ba0426c8f70bb719545a9da", "<KEY>", "7355f5bed7ba43acaeac5e01e7716806", "bb17425d3e5341228451d33baa4b3992", "0a8d44a8fe1f416eaac8799d53d68389", "9d0be38181f246f593a7825e48e1efc8", "bd01e9d0ef0147e99f77181c2de9155b", "9d8b903df5914ffea5a5e9d47f213261", "<KEY>", "<KEY>", "f4dc43df114e4c438374b5a34d8a73bd", "9f6b578f9cc74d009f275555493597d9", "517e2a2c4ae64d139c7e06ca7594c48e", "844ded6dcdd54c3cb5d20be290fdd947"]} executionInfo={"elapsed": 422390, "status": "ok", "timestamp": 1611383345226, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416044610109286808"}, "user_tz": -480} id="Auf0PEu2cS9x" outputId="f6d71d2e-cf49-4a8f-e465-70c729c063b4"
folder_aug = "/content/drive/MyDrive/FYP/augmented images"
aug_array = []
class_num_list2 = []
subcls_num_list2 = ["None"]*500 # fill the list with 500 "None's" first to represent "healthy" samples
for c in CLASSES:
if (c == "healthy"):
path = os.path.join(folder_aug,c) # create path to dataset folder_aug
class_num = 0 # get the CLASSES index, 0 = healthy, 1 = ill
for img in tqdm(os.listdir(path)): # iterate over each image per folder_aug
try:https://forms.gle/wu1oZvrwYKGUpYM18
img = cv2.imread(os.path.join(path,img)) # read image into a list
aug_array.append(img)
class_num_list2.append(class_num) # save a list of the class numbers
except Exception as e:
print("error")
elif (c == "ill"):
for s in SUBCLASSES:
path = os.path.join(folder_aug,c,s) # create path to ill subfolder_augs
class_num = 1
subclass_num = SUBCLASSES.index(s)
for img in tqdm(os.listdir(path)): # iterate over each image per folder_aug
try:
img = cv2.imread(os.path.join(path,img)) # read image into a list
aug_array.append(img)
class_num_list2.append(class_num)
subcls_num_list2.append(subclass_num) # save a list of the subclass numbers; 0 = fever, 1 = sore throat, 2 = running nose
except Exception as e:
print("error")
# + [markdown] id="2kLF31p6HGit"
# ## First-Level Classification
# > ### *Classify samples into healthy (0) and ill (1) classes*
#
# + executionInfo={"elapsed": 1674, "status": "ok", "timestamp": 1611383464830, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416044610109286808"}, "user_tz": -480} id="MEuw9582cS91"
# Create partial dataset of original images (will add in the augmented set after spliting)
dataset = []
for i, img in enumerate(img_array):
try:
img_gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
dataset.append([img_gray, class_num_list1[i], subcls_num_list1[i]])
except Exception as e:
print("error")
# + executionInfo={"elapsed": 2182, "status": "ok", "timestamp": 1611383465347, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416044610109286808"}, "user_tz": -480} id="_XSiQmhxcS93"
# Create the augmented images' dataset
aug_dataset = []
for i, aug in enumerate(aug_array):
try:
aug_gray = cv2.cvtColor(aug, cv2.COLOR_BGR2GRAY)
aug_dataset.append([aug_gray, class_num_list2[i], subcls_num_list2[i]])
except Exception as e:
print("error")
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 2147, "status": "ok", "timestamp": 1611383465352, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416044610109286808"}, "user_tz": -480} id="58eiMLPFcS94" outputId="e50418fc-61f7-474e-9103-70c29d20bcf2"
# Split the original dataset
train_set_before, test_set = train_test_split(dataset, test_size=0.2, shuffle=True)
print('Train set length before data augmentation: ', len(train_set_before))
print('Test set length: ', len(test_set))
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 2131, "status": "ok", "timestamp": 1611383465354, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416044610109286808"}, "user_tz": -480} id="x3rHcqT8cS95" outputId="712004d1-ba25-45c0-f9d9-4d9bb629f259"
# Add the augmented dataset into the training set, then shuffle
train_set = []
train_set = train_set_before.copy()
for aug_set in aug_dataset:
if (len(train_set) < 1319):
train_set.append(aug_set)
random.shuffle(train_set)
len(train_set)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 1208, "status": "ok", "timestamp": 1611384169968, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416044610109286808"}, "user_tz": -480} id="O9Z3yfSY5OwN" outputId="b6b7b939-c6e7-4b1c-a8f2-1f8e856cb7fd"
# Create training set
X_train = []
y_train = []
gray_img1 = []
subcls_num = []
for features, label, subcls in train_set:
X_train.append(features)
y_train.append(label)
subcls_num.append(subcls)
gray_img1.append(features)
#X_train = np.array(X_train).reshape(-1, 200, 200, 1)
X_train = np.array(X_train).reshape(len(y_train), -1)
y_train = np.array(y_train)
print(X_train.shape)
print(y_train.shape)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 1192, "status": "ok", "timestamp": 1611384169971, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416044610109286808"}, "user_tz": -480} id="m8kNRZrX5OwO" outputId="e3202970-1d1d-4715-d067-49c5caefa2e3"
# Create testing set
X_test = []
y_test = []
gray_img2 = []
for features, label, subcls in test_set:
X_test.append(features)
y_test.append(label)
gray_img2.append(features)
#X_test = np.array(X_test).reshape(-1, 200, 200, 1)
X_test = np.array(X_test).reshape(len(y_test), -1)
y_test = np.array(y_test)
print(X_test.shape)
print(y_test.shape)
# + [markdown] id="-IK1KAsRF0ou"
# ### Feature Extraction
# + executionInfo={"elapsed": 8928, "status": "ok", "timestamp": 1611384177720, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416044610109286808"}, "user_tz": -480} id="wXb4-XExdd8H"
# Apply PCA to extract features
n_components = 50
pca = PCA(n_components=n_components, svd_solver='randomized',
whiten=True).fit(X_train)
X_train = pca.transform(X_train)
X_test = pca.transform(X_test)
# + [markdown] id="84UEBtn7idcv"
# ### Train Model
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 15693, "status": "ok", "timestamp": 1611384184494, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416044610109286808"}, "user_tz": -480} id="yw-8CAJS5OwW" outputId="3e14bb7d-3ff0-4bda-d4cb-00706385d047"
# Train the classifier
rf = RandomForestClassifier(n_estimators=1000, n_jobs=-1)
start_train = time.time() # to record the training time
rf.fit(X_train, y_train)
end_train = time.time()
print(end_train - start_train, "seconds")
# + [markdown] id="EN0feoe9o5yx"
# #### Training Performance
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 16371, "status": "ok", "timestamp": 1611384185188, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416044610109286808"}, "user_tz": -480} id="5flMCsPGcS-C" outputId="0ffe1995-a109-4daa-e854-ec3f9b713739"
y_pred = rf.predict(X_train)
ill_pred = y_pred
print("Classification report for - \n{}:\n{}\n".format(
rf, metrics.classification_report(y_train, y_pred)))
train_acc = metrics.accuracy_score(y_train, y_pred)*100
print("Training accuracy: " + str(train_acc))
# + colab={"base_uri": "https://localhost:8080/", "height": 295} executionInfo={"elapsed": 16728, "status": "ok", "timestamp": 1611384185567, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416044610109286808"}, "user_tz": -480} id="zBZGN1w9cS-D" outputId="3edc543e-a7ce-4d6c-de3d-8ea8ee197c12"
plot_confusion_matrix(rf, X_train, y_train, cmap='plasma')
# + [markdown] id="d5qV4luLo5yz"
# #### Testing Performance
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 17059, "status": "ok", "timestamp": 1611384185917, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416044610109286808"}, "user_tz": -480} id="5AzTPuIncS-E" outputId="14d0ab55-e644-4564-8174-1cc3e2014bb4"
y_pred = rf.predict(X_test)
print("Classification report for - \n{}:\n{}\n".format(
rf, metrics.classification_report(y_test, y_pred)))
test_acc = metrics.accuracy_score(y_test, y_pred)*100
print("Testing accuracy: " + str(test_acc))
# + colab={"base_uri": "https://localhost:8080/", "height": 295} executionInfo={"elapsed": 17665, "status": "ok", "timestamp": 1611384186543, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416044610109286808"}, "user_tz": -480} id="x0VzOzcRcS-F" outputId="af6300b7-6956-422e-84d5-b5433603d6a9"
plot_confusion_matrix(rf, X_test, y_test, cmap='plasma')
# + executionInfo={"elapsed": 17651, "status": "ok", "timestamp": 1611384186545, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416044610109286808"}, "user_tz": -480} id="5Pbb-Mm9dd8L"
def get_wrong_case(pred_result, test_result):
#get wrongly classified samples
for i in range(len(pred_result)):
predicted = pred_result[i]
actual = test_result[i]
if(actual != predicted):
wrong_case.append([i,predicted,actual])
# + executionInfo={"elapsed": 727, "status": "ok", "timestamp": 1611386796860, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416044610109286808"}, "user_tz": -480} id="ZYpCERhsdd8L"
def plot(wrong_case, row, col, gray, labels):
if (labels == SUBCLASSES):
fsize = 18
top = 5
bottom = 4
else:
fsize = 20
top = 3
bottom = 2.3
plt.figure(figsize=(40, 40))
plt.subplots_adjust(top=top, bottom=bottom)
for i in range(len(wrong_case)):
plt.subplot(row, col, i+1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
title = str(i+1) + '. Predicted: ' + labels[wrong_case[i][1]] + '\nActual: ' + labels[wrong_case[i][2]]
plt.title(title, fontdict = {'fontsize' : fsize})
plt.imshow(gray[wrong_case[i][0]], cmap='gray')
plt.show()
print("The number of misclassified images: ")
print(len(wrong_case))
# + executionInfo={"elapsed": 17634, "status": "ok", "timestamp": 1611384186549, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416044610109286808"}, "user_tz": -480} id="QljJDCD1dd8M"
wrong_case = []
get_wrong_case(y_pred, y_test)
# + [markdown] id="v8np4hKTo5y4"
# ## Second-Level Classification
# > ### *Classify 'ill' cases into fever (0), sore throat (1) and running nose (2) subclasses*
#
#
# + executionInfo={"elapsed": 20179, "status": "ok", "timestamp": 1611384189122, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416044610109286808"}, "user_tz": -480} id="nrBA0NQzdd8O"
ill_case_index = []
ill_dataset = []
# + executionInfo={"elapsed": 20170, "status": "ok", "timestamp": 1611384189124, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416044610109286808"}, "user_tz": -480} id="j1WjQ8N4dd8P"
# Get the indexes of samples correctly classified as "ill"
for i, a in enumerate(ill_pred):
if((ill_pred[i] == 1) and (ill_pred[i] == y_train[i]) and (subcls_num[i] != "None")): # 1 = ill, filter out the cases with "healthy" true labels
ill_case_index.append(i) # save the indexes of ill cases
# + executionInfo={"elapsed": 20163, "status": "ok", "timestamp": 1611384189126, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416044610109286808"}, "user_tz": -480} id="1eV_TObZdd8Q"
# Create dataset for second-level classification
for a in ill_case_index:
ill_dataset.append([X_train[a], subcls_num[a], gray_img1[a]]) # subcls_num[] contains subclass labels (e.g 0, 1, 2)
ill_dataset = np.array(ill_dataset, dtype=object)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 20156, "status": "ok", "timestamp": 1611384189128, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416044610109286808"}, "user_tz": -480} id="M6m_e-cAdd8U" outputId="79c7b855-2ee6-412e-ae7a-a1a72d05179a"
train_ill_set, test_ill_set = train_test_split(ill_dataset, test_size=0.3, shuffle=True)
print('Train set length: ', len(train_ill_set))
print('Test set length: ', len(test_ill_set))
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 20147, "status": "ok", "timestamp": 1611384189132, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416044610109286808"}, "user_tz": -480} id="TUfvP3iMdd8V" outputId="9669655f-f20e-4800-847d-afcf4b38ac05"
X_train_ill = []
y_train_ill = []
gray_img_ill1 = []
for features, label, gray in train_ill_set:
X_train_ill.append(features)
y_train_ill.append(label)
gray_img_ill1.append(gray)
X_train_ill = np.array(X_train_ill)
y_train_ill = np.array(y_train_ill)
print(X_train_ill.shape)
print(y_train_ill.shape)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 20139, "status": "ok", "timestamp": 1611384189138, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416044610109286808"}, "user_tz": -480} id="d2AdlGZMdd8W" outputId="1f7bcbdf-0583-49e1-c853-74c7db36fbb3"
X_test_ill = []
y_test_ill = []
gray_img_ill2 = []
for features, label, gray in test_ill_set:
X_test_ill.append(features)
y_test_ill.append(label)
gray_img_ill2.append(gray)
X_test_ill = np.array(X_test_ill)
y_test_ill = np.array(y_test_ill)
print(X_test_ill.shape)
print(y_test_ill.shape)
# + [markdown] id="HZwp1-eSi04s"
# ### Train Model
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 22441, "status": "ok", "timestamp": 1611384191453, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416044610109286808"}, "user_tz": -480} id="sO_ygpoBdd8Y" outputId="153de337-7e42-41a0-f282-59b51374821d"
rf = RandomForestClassifier(n_estimators=1200, n_jobs=-1)
start_train_ill = time.time()
rf.fit(X_train_ill, y_train_ill)
end_train_ill = time.time()
print(end_train_ill - start_train_ill, "seconds")
# + [markdown] id="iflaNAaZIRLQ"
# #### Training Performance
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 22767, "status": "ok", "timestamp": 1611384191793, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416044610109286808"}, "user_tz": -480} id="tUXmJWoRcS-M" outputId="4ec2637a-489d-4ae3-b17f-cd31a1687ce6"
y_pred_ill = rf.predict(X_train_ill)
print("Classification report for - \n{}:\n{}\n".format(
rf, metrics.classification_report(y_train_ill, y_pred_ill)))
train_ill_acc = metrics.accuracy_score(y_train_ill, y_pred_ill)*100
print("Training accuracy: " + str(train_ill_acc))
# + colab={"base_uri": "https://localhost:8080/", "height": 295} executionInfo={"elapsed": 23591, "status": "ok", "timestamp": 1611384192633, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416044610109286808"}, "user_tz": -480} id="xnyct16DcS-M" outputId="0a79c119-c26f-4bc4-ca70-3584b74ca60b"
plot_confusion_matrix(rf, X_train_ill, y_train_ill, cmap='viridis')
# + [markdown] id="P9WK2WiqIXdl"
# #### Testing Performance
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 23907, "status": "ok", "timestamp": 1611384192964, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416044610109286808"}, "user_tz": -480} id="bzoPpMUMcS-N" outputId="dea7bb86-c847-4f64-e451-b272b8a55c25"
y_pred_ill = rf.predict(X_test_ill)
print("Classification report for - \n{}:\n{}\n".format(
rf, metrics.classification_report(y_test_ill, y_pred_ill)))
test_ill_acc = metrics.accuracy_score(y_test_ill, y_pred_ill)*100
print("Testing accuracy: " + str(test_ill_acc))
# + colab={"base_uri": "https://localhost:8080/", "height": 295} executionInfo={"elapsed": 24980, "status": "ok", "timestamp": 1611384194059, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416044610109286808"}, "user_tz": -480} id="XPfD2ihBcS-O" outputId="b3e7a3e7-e146-4278-e104-ccaf0ccac83e"
plot_confusion_matrix(rf, X_test_ill, y_test_ill, cmap='viridis')
# + executionInfo={"elapsed": 24961, "status": "ok", "timestamp": 1611384194062, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416044610109286808"}, "user_tz": -480} id="ICP4o56RcS-O"
wrong_case = []
get_wrong_case(y_pred_ill, y_test_ill)
| PCA_RF_classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Problem Understanding
# The Internet has profoundly changed the way we buy things, but the online shopping of today is likely not the end of that change; after each purchase we still need to wait multiple days for physical goods to be carried to our doorstep. This is where drones come in autonomous, electric vehicles delivering online purchases. Flying, so never stuck in traffic. As drone technology improves every year, there remains a major issue: how do we manage and coordinate all those drones?
#
# ## Task
# Given a hypothetical fleet of drones, a list of customer orders and availability of the individual products in warehouses, your task is to schedule the drone operations so that the orders are completed as soon as possible. You will need to handle the complications of multiple drones, customer orders, product types and weights, warehouses, and delivery destinations.
import seaborn as sns
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from scipy.spatial import distance_matrix
# # Import dataset
with open('../data/busy_day.in') as file:
data_list = file.read().splitlines()
print(' Rows of grid,columns of grid,drones,turns, maxpay load in units(u):',data_list[0],
'\n Different product types:',data_list[1],
'\n product types weigh:',data_list[2],
'\n warehouses:',data_list[3],
'\n First warehouse location at first warehouse (row, column):',data_list[4],
'\n Inventory of products:',data_list[5],
'\n second warehouse location (row, column) :',data_list[6],
'\n Inventory of products at second ware house:',data_list[7],
'\n Number of orders:',data_list[24],
'\n First order to be delivered to:',data_list[25],
'\n Number of items in first order:',data_list[26],
'\n Items of product types:',data_list[27] )
# # Examine data
weight = [int(i) for i in data_list[2].split(" ")]
ax = sns.distplot(weight)
ax.set_title('Product Weight Distribution')
ax.set_xlabel("Weight")
plt.show()
warehouse = {}
for i in range(10):
warehouse[i] = [int(i) for i in data_list[5+2*i].split(" ")]
df = pd.DataFrame(warehouse).T
df = df.add_prefix("prd_")
ax = sns.distplot(df.sum())
ax.set_title("Product count distribution")
ax.set_xlabel("Item count")
plt.show()
print("There are on average",df.sum().mean(),"items for each product across all warehouses")
data = df.sum(axis=1).to_frame()
ax = sns.barplot(data = data, x = data.index, y = data[0])
ax.set_title("Products at each warehouse")
ax.set_xlabel("Warehouse")
ax.set_ylabel("Product count")
plt.show()
# # Show Location
warehouse_location = {}
for i in range(10):
warehouse_location[i] = [int(i) for i in data_list[4+2*i].split(" ")]
df_wh_coor = pd.DataFrame(warehouse_location).T
df_wh_coor.columns = ["X-C","Y-C"]
df_wh_coor["type"] = "warehouse"
order_location = {}
for i in range(1250) :
order_location[i] = [int(i) for i in data_list[25+3*i].split(" ")]
df_order_coor = pd.DataFrame(order_location).T
df_order_coor.columns = ["X-C","Y-C"]
df_order_coor["type"] = "order"
plane = pd.concat([df_wh_coor, df_order_coor])
plt.figure(figsize=(15,8))
sns.scatterplot(x = plane["X-C"],y= plane["Y-C"], hue=plane["type"])
plt.show()
# # Orders
orders = {}
for i in range(1250):
orders[i] = [int(i) for i in data_list[27+3*i].split(" ")]
# # Distances
temp = plane[["X-C","Y-C"]]
distances = distance_matrix(temp.values.tolist(),temp.values.tolist())
distances
# # Optimization
from ortools.graph import pywrapgraph
from ortools.constraint_solver import routing_enums_pb2
from ortools.constraint_solver import pywrapcp
start_node = np.repeat(np.arange(0,10), 1250).tolist()
end_node = np.tile(np.arange(10,1250), 10).tolist()
def create_data_model():
"""Stores the data for the problem."""
data = {}
data['distance_matrix'] = distances
data['num_vehicles'] = 30
data['depot'] = 0
return data
# +
def distance_callback(from_index, to_index):
"""Returns the distance between the two nodes."""
# Convert from routing variable Index to distance matrix NodeIndex.
from_node = manager.IndexToNode(from_index)
to_node = manager.IndexToNode(to_index)
return data['distance_matrix'][from_node][to_node]
transit_callback_index = routing.RegisterTransitCallback(distance_callback)
# -
dimension_name = 'Distance'
routing.AddDimension(
transit_callback_index,
0, # no slack
3000, # vehicle maximum travel distance
True, # start cumul to zero
dimension_name)
distance_dimension = routing.GetDimensionOrDie(dimension_name)
distance_dimension.SetGlobalSpanCostCoefficient(100)
def print_solution(data, manager, routing, solution):
"""Prints solution on console."""
max_route_distance = 0
for vehicle_id in range(data['num_vehicles']):
index = routing.Start(vehicle_id)
plan_output = 'Route for vehicle {}:\n'.format(vehicle_id)
route_distance = 0
while not routing.IsEnd(index):
plan_output += ' {} -> '.format(manager.IndexToNode(index))
previous_index = index
index = solution.Value(routing.NextVar(index))
route_distance += routing.GetArcCostForVehicle(
previous_index, index, vehicle_id)
plan_output += '{}\n'.format(manager.IndexToNode(index))
plan_output += 'Distance of the route: {}m\n'.format(route_distance)
print(plan_output)
max_route_distance = max(route_distance, max_route_distance)
print('Maximum of the route distances: {}m'.format(max_route_distance))
# +
def main():
"""Solve the CVRP problem."""
# Instantiate the data problem.
data = create_data_model()
# Create the routing index manager.
manager = pywrapcp.RoutingIndexManager(len(data['distance_matrix']),
data['num_vehicles'], data['depot'])
# Create Routing Model.
routing = pywrapcp.RoutingModel(manager)
# Define cost of each arc.
routing.SetArcCostEvaluatorOfAllVehicles(transit_callback_index)
# Add Distance constraint.
dimension_name = 'Distance'
routing.AddDimension(
transit_callback_index,
0, # no slack
3000, # vehicle maximum travel distance
True, # start cumul to zero
dimension_name)
distance_dimension = routing.GetDimensionOrDie(dimension_name)
distance_dimension.SetGlobalSpanCostCoefficient(100)
# Setting first solution heuristic.
search_parameters = pywrapcp.DefaultRoutingSearchParameters()
search_parameters.first_solution_strategy = (
routing_enums_pb2.FirstSolutionStrategy.PATH_CHEAPEST_ARC)
# Solve the problem.
solution = routing.SolveWithParameters(search_parameters)
# Print solution on console.
if solution:
print_solution(data, manager, routing, solution)
if __name__ == '__main__':
main()
# -
| Notebook/.ipynb_checkpoints/Drone-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# formats: ipynb,.md//md
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from nbev3devsim.load_nbev3devwidget import roboSim, eds
# %load_ext nbev3devsim
# %load_ext nbtutor
# + [markdown] activity=true
# # 7 Optional extra - handling images
#
#
# One of the advantages of using the Python `PIL` package is that a range of *methods* (that is, *functions*) are defined on each image object that allow us to manipulate it *as an image*. (We can then also access the data defining the transformed image *as data* if we need it in that format.)
#
# We can preview the area in our sampled image by cropping the image to the area of interest:
#
# ```python
# # The crop area is (x, y, x + side, y + side)
# cropped_image = img.crop((3, 3, 17, 17));
#
# display(cropped_image.size)
# zoom_img(cropped_image)
# ```
#
# In order to present this image as a test image to the trained MLP, we need to resize it so that it is the same size as the training images:
#
# ```python
# from PIL import Image
#
# # TO DO - should we add crop and resize as parameters to generate_image?
# resized_cropped_image = cropped_image.resize((28, 28), Image.LANCZOS)
# zoom_img(resized_cropped_image)
# ```
#
# ```python
# generate_image(roboSim.image_data(), index,
# crop=(3, 3, 17, 17),
# resize=(28, 28))
# ```
#
# ```python
# image_class_predictor(MLP, resized_cropped_image);
# ```
#
# ```python
# The `sensor_data.sensor_image_focus()` function will take an image, crop it to the central area, a
# ```
#
#
# ## Playing with images
#
# If we create a blank image of size 28x28 pixels with a grey background, we can paste a copy of our cropped image into it.
#
# ```python
# _grey_background = 200
#
# _image_size = (28, 28)
# _image_mode = 'L' #greyscale image mode
#
# shift_image = Image.new(_image_mode, _image_size, _grey_background)
#
# # Set an offset for where to paste the image
# _xy_offset = (2, 6)
#
# shift_image.paste(cropped_image, _xy_offset)
# zoom_img(shift_image)
# ```
#
# Alternatively, we might zoom the cropped image back to the original image size. (The `Image.LANCZOS` setting defines a filter that is used to interpolate new pixel values in the scaled image based on the pixel values in the cropped image. As such, it may introduce digital artefacts of its own into the scaled image.)
#
# ```python
# resized_image = cropped_image.resize(image_image.size, Image.LANCZOS)
#
# display( resized_image.size)
# zoom_img(resized_image)
# ```
#
#
# Note that other image handling tools are available to us within the `PIL` package that allow us to perform other "native" image manipulating functions, such as cropping an image:
#
# ```python
# # Define the limits of the crop operation
#
# # Cut 6 pixel columns on lefthand edge
# x0 = 6
# # Cut 10 pixel columns on righthand edge
# x1 = image_image.size[1] -10
#
# # Leave the rows alone
# y0 = 0
# y1 = image_image.size[1]
#
# crop_image = image_image.crop((x0, y0, x1, y1))
# zoom_img(crop_image)
# ```
| content/08. Remote services and multi-agent systems/Section_00_07.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Dataset Report Example
#
# DatasetReportBuilder instance can analyze the dataset and generate a report in HTML format.
#
# +
import hana_ml
from hana_ml import dataframe
from data_load_utils import DataSets, Settings
url, port, user, pwd = Settings.load_config("../../config/e2edata.ini")
connection_context = dataframe.ConnectionContext(url, port, user, pwd)
# -
# ## Create DatasetReportBuilder Instance
from hana_ml.visualizers.dataset_report import DatasetReportBuilder
datasetReportBuilder = DatasetReportBuilder()
# ## Prepare Diabetes Dataset
# Original data comes from National Institute of Diabetes and Digestive and Kidney Diseases. The collected dataset is aiming at, based on certain diagnostic measurements, diagnostically predicting whether or not a patient has diabetes. In particular, patients contained in the dataset are females of Pima Indian heritage, all above the age of 20. Dataset is form Kaggle, for tutorials use only.
#
# The dataset contains the following diagnositic <b>attributes</b>:<br>
# $\rhd$ "PREGNANCIES" - Number of times pregnant,<br>
# $\rhd$ "GLUCOSE" - Plasma glucose concentration a 2 hours in an oral glucose tolerance test,<br>
# $\rhd$ "BLOODPRESSURE" - Diastolic blood pressure (mm Hg),<br>
# $\rhd$ "SKINTHICKNESS" - Triceps skin fold thickness (mm),<br>
# $\rhd$ "INSULIN" - 2-Hour serum insulin (mu U/ml),<br>
# $\rhd$ "BMI" - Body mass index $(\text{weight in kg})/(\text{height in m})^2$,<br>
# $\rhd$ "PEDIGREE" - Diabetes pedigree function,<br>
# $\rhd$ "AGE" - Age (years),<br>
# $\rhd$ "CLASS" - Class variable (0 or 1) 268 of 768 are 1(diabetes), the others are 0(non-diabetes).
full_tbl, training_valid_tbl, test_tbl, _ = DataSets.load_diabetes_data(connection_context)
diabetes_dataset = connection_context.table(training_valid_tbl)
# ## Analyze Diabetes Dataset
datasetReportBuilder.build(diabetes_dataset,key="ID")
# ## Display Report
datasetReportBuilder.generate_notebook_iframe_report()
# ## Generate HTML Report
# +
# datasetReportBuilder.generate_html_report('diabetes')
# -
# ## Prepare Iris Dataset
# A data set that identifies different types of Iris's is used to demonstrate the use of multi layer perceptron in SAP HANA. This data set is also used in a clustering example where the objective was to cluster the flowers into three clusters and the intuition was that the three clusters would correspond to the three types of Iris's in the data set. Since we know the labels (i.e. the types of Iris's), we can use classification to create a model to predict the type of flower based on features or characteristics that are explained below.
#
# ## Iris Data Set
# The data set used is from University of California, Irvine (https://archive.ics.uci.edu/ml/datasets/iris). For tutorials use only. This data set contains attributes of a plant iris. There are three species of Iris plants.
# <table>
# <tr><td>Iris Setosa</td><td><img src="images/Iris_setosa.jpg" title="Iris Sertosa" style="float:left;" width="300" height="50" /></td>
# <td>Iris Versicolor</td><td><img src="images/Iris_versicolor.jpg" title="Iris Versicolor" style="float:left;" width="300" height="50" /></td>
# <td>Iris Virginica</td><td><img src="images/Iris_virginica.jpg" title="Iris Virginica" style="float:left;" width="300" height="50" /></td></tr>
# </table>
#
# The data contains the following attributes for various flowers:
# <table align="left"><tr><td>
# <li align="top">sepal length in cm</li>
# <li align="left">sepal width in cm</li>
# <li align="left">petal length in cm</li>
# <li align="left">petal width in cm</li>
# </td><td><img src="images/sepal_petal.jpg" style="float:left;" width="200" height="40" /></td></tr></table>
#
# Although the flower is identified in the data set, we will cluster the data set into 3 clusters since we know there are three different flowers. The hope is that the cluster will correspond to each of the flowers.
#
# A different notebook will use a classification algorithm to predict the type of flower based on the sepal and petal dimensions.
full_tbl, training_tbl, validation_tbl, test_tbl = DataSets.load_iris_data(connection_context)
iris_dataset = connection_context.table(training_tbl)
# ## Analyze Iris Dataset
datasetReportBuilder.build(iris_dataset,key="ID")
# ## Display Report
datasetReportBuilder.generate_notebook_iframe_report()
# ## Generate HTML Report
# +
# datasetReportBuilder.generate_html_report('iris')
# -
| Python-API/pal/notebooks/DatasetReport.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # MadMiner particle physics tutorial
#
# # Part 4c: Information Geometry
#
# <NAME>, <NAME>, <NAME>, and <NAME> 2018-2019
# ## 0. Preparations
# +
import sys
import os
madminer_src_path = "/Users/felixkling/Documents/GitHub/madminer"
sys.path.append(madminer_src_path)
from __future__ import absolute_import, division, print_function, unicode_literals
import six
import logging
import numpy as np
import matplotlib
from matplotlib import pyplot as plt
# %matplotlib inline
from madminer.fisherinformation import FisherInformation, InformationGeometry
from madminer.plotting import plot_fisher_information_contours_2d
# +
# MadMiner output
logging.basicConfig(
format='%(asctime)-5.5s %(name)-20.20s %(levelname)-7.7s %(message)s',
datefmt='%H:%M',
level=logging.INFO
)
# Output of all other modules (e.g. matplotlib)
for key in logging.Logger.manager.loggerDict:
if "madminer" not in key:
logging.getLogger(key).setLevel(logging.WARNING)
# -
# Let's look at a simple example to unserstand what happens in information geometry. At first we note that the Fisher Information is a symmetric positive definite rank two tensor, and therefore can be seen as a Riemanian metric. It can therefore be used to calculate distances between points in parameter space.
#
# Previously, in tutorial 4b, we have considered the **local distance** $d_{local}(\theta,\theta_0)$ between two points $\theta$ and $\theta_0$. It is defined in the tangent space of $\theta_0$, where the metric is constant and hence flat, and can simply be calculated as $d_{local}(\theta,\theta_0) = I_{ij}(\theta_0) \times (\theta-\theta_0)^i (\theta-\theta_0)^j$.
#
# Going beyond this local approximation, we can calculate a **global distance** $d_{global}(\theta,\theta_0)$ which takes into account the fact that the information is not constant throughout the parameter space. Using our knowledge from general relativity, this distance is defined as
# \begin{equation}
# d(\theta,\theta_0)= \text{min} \int_{\theta_0}^{\theta} ds \sqrt{I_{ij}(\theta(s)) \frac{d\theta^i}{ds}\frac{d\theta^j}{ds}}
# \end{equation}
# where $\theta(s)$ is the geodesic (the shortest path) connecting $\theta_0$ and $\theta$. This path is follows the geodesic equation
# \begin{equation}
# \frac{d^2\theta^i}{ds^2} = - \Gamma^i_{jk} \frac{d\theta^j}{ds}\frac{d\theta^k}{ds} \quad \text{with} \quad
# \Gamma^i_{jk} = \frac{1}{2} I^{im} \Big(\frac{\partial I_{mk}}{\partial \theta^j} + \frac{\partial I_{mj}}{\partial \theta^k} - \frac{\partial I_{jk}}{\partial \theta^m}\Big) \quad \text{and} \quad I^{im} I_{mj} = \delta^i_j \ .
# \end{equation}
# In practice, we obtain the geodesics by numerically integrating the geodesic equation, starting at a parameter point $\theta_0$ with a velocity $\theta'_0=(\theta/ds)_0$
# ## 1. Stand Alone Example
# In the following, we consider a sample geometry with Fisher Information $I_{ij}(\theta)= (( 1+\theta_1/4 , 1 ),( 1 , 2-\theta_2/2))$ and determine the geodesics and distance contours for illistration. At first, we initialize a new class `InformationGeometry` and define the Fisher Information via the function `information_from_formula()`.
# +
formula="np.array([[1 + 0.25*theta[0] ,1],[1, 2 - 0.5*theta[1] ]])"
infogeo=InformationGeometry()
infogeo.information_from_formula(formula=formula,dimension=2)
# -
# Now we obtain one particular geodesic path staring at $\theta_0$ in the direction of $\Delta \theta_0$ using the function `find_trajectory()`.
thetas,distances=infogeo.find_trajectory(
theta0=np.array([0.,0.]),
dtheta0=np.array([1.,1.]),
limits=np.array([[-1.,1.],[-1.,1.]]),
stepsize=0.025,
)
# For comparisson, let's do the same for a constant Fisher Information $I_{ij}(\theta)=I_{ij}(\theta_0)=((1,1),(1,2))$.
formula_lin="np.array([[1 ,1],[1, 2 ]])"
infogeo_lin=InformationGeometry()
infogeo_lin.information_from_formula(formula=formula_lin,dimension=2)
thetas_lin,distances_lin=infogeo_lin.find_trajectory(
theta0=np.array([0.,0.]),
dtheta0=np.array([1.,1.]),
limits=np.array([[-1.,1.],[-1.,1.]]),
stepsize=0.025,
)
# and plot the results
# +
cmin, cmax = 0., 2
fig = plt.figure(figsize=(6,5))
plt.scatter(
thetas_lin.T[0],thetas_lin.T[1],c=distances_lin,
s=10., cmap='viridis',marker='o',vmin=cmin, vmax=cmax,
)
sc = plt.scatter(
thetas.T[0],thetas.T[1],c=distances,
s=10., cmap='viridis',marker='o',vmin=cmin, vmax=cmax,
)
plt.scatter( [0],[0],c='k')
cb = plt.colorbar(sc)
cb.set_label(r'Distance $d(\theta,\theta_0)$')
plt.xlabel(r'$\theta_1$')
plt.ylabel(r'$\theta_2$')
plt.tight_layout()
plt.show()
# -
# We can see that the geodesic trajectory is curved. The colorbar denotes the distance from the origin.
# Let us now see how we can construct the distance contours using the function `distance_contours`.
# +
grid_ranges = [(-1, 1.), (-1, 1.)]
grid_resolutions = [25, 25]
theta_grid,p_values,distance_grid,(thetas,distances)=infogeo.distance_contours(
np.array([0.,0.]),
grid_ranges=grid_ranges,
grid_resolutions=grid_resolutions,
stepsize=0.08,
ntrajectories=30,
continous_sampling=True,
return_trajectories=True,
)
# -
# and plot the results
# +
#Prepare Plot
cmin, cmax = 0., 2
fig = plt.figure(figsize=(15.0, 4.0 ))
bin_size = (grid_ranges[0][1] - grid_ranges[0][0])/(grid_resolutions[0] - 1)
edges = np.linspace(grid_ranges[0][0] - bin_size/2, grid_ranges[0][1] + bin_size/2, grid_resolutions[0] + 1)
centers = np.linspace(grid_ranges[0][0], grid_ranges[0][1], grid_resolutions[0])
#Plot
ax = plt.subplot(1,3,1)
sc = ax.scatter(thetas.T[0],thetas.T[1],c=distances,vmin=cmin, vmax=cmax,)
cb = plt.colorbar(sc,ax=ax, extend='both')
cb.set_label(r'Distance $d(\theta,\theta_0)$')
ax.set_xlabel(r'$\theta_1$')
ax.set_ylabel(r'$\theta_2$')
ax = plt.subplot(1,3,2)
cm = ax.pcolormesh(
edges, edges, distance_grid.reshape((grid_resolutions[0], grid_resolutions[1])).T,
vmin=cmin, vmax=cmax,
cmap='viridis'
)
cb = plt.colorbar(cm, ax=ax, extend='both')
cb.set_label(r'Distance $d(\theta,\theta_0)$')
ax.set_xlabel(r'$\theta_1$')
ax.set_ylabel(r'$\theta_2$')
ax = plt.subplot(1,3,3)
cm = ax.pcolormesh(
edges, edges, p_values.reshape((grid_resolutions[0], grid_resolutions[1])).T,
norm=matplotlib.colors.LogNorm(vmin=0.1, vmax=1),
cmap='viridis'
)
cb = plt.colorbar(cm, ax=ax, extend='both')
cb.set_label('Expected p-value')
ax.set_xlabel(r'$\theta_1$')
ax.set_ylabel(r'$\theta_2$')
plt.tight_layout()
plt.show()
# -
# The left plot shows the distance values along generated geodesics. These values are interpolated into a continuous function shown in the middle plot. In the right plot we convert the distances into expected p-values.
# ## 2. Information Geometry Bounds for Example Process
# Now that we understand how Information Geometry works in principle, let's apply it to our example process. Let's first create a grid of theta values
# +
def make_theta_grid(theta_ranges, resolutions):
theta_each = []
for resolution, (theta_min, theta_max) in zip(resolutions, theta_ranges):
theta_each.append(np.linspace(theta_min, theta_max, resolution))
theta_grid_each = np.meshgrid(*theta_each, indexing="ij")
theta_grid_each = [theta.flatten() for theta in theta_grid_each]
theta_grid = np.vstack(theta_grid_each).T
return theta_grid
grid_ranges = [(-1, 1.), (-1, 1.)]
grid_resolutions = [25, 25]
theta_grid = make_theta_grid(grid_ranges,grid_resolutions)
# -
# Now we create a grid of Fisher Informations. Since this might take some time, we already prepared the results, which can be loaded directly.
model='alices'
calculate_fisher_grid=False
if calculate_fisher_grid:
fisher = FisherInformation('data/lhe_data_shuffled.h5')
fisher_grid=[]
for theta in theta_grid:
fisher_info, _ = fisher.full_information(
theta=theta,
model_file='models/'+model,
luminosity=300.*1000.,
include_xsec_info=False,
)
fisher_grid.append(fisher_info)
np.save("limits/infogeo_thetagrid_"+model+".npy", theta_grid)
np.save("limits/infogeo_fishergrid_"+model+".npy", fisher_grid)
else:
theta_grid=np.load("limits/infogeo_thetagrid_"+model+".npy")
fisher_grid=np.load("limits/infogeo_fishergrid_"+model+".npy")
# In the next step, we initialize the `InformationGeoemtry` class using this input data. Using the function `information_from_grid()`, the provided grid is interpolated using a piecewise linear function and the information can be calculated at every point.
infogeo=InformationGeometry()
infogeo.information_from_grid(
theta_grid="limits/infogeo_thetagrid_"+model+".npy",
fisherinformation_grid="limits/infogeo_fishergrid_"+model+".npy",
)
# As before, we can now obtain the p-values using the `distance_contours()` function
theta_grid,p_values_infogeo,distance_grid,(thetas,distances)=infogeo.distance_contours(
np.array([0.,0.]),
grid_ranges=grid_ranges,
grid_resolutions=grid_resolutions,
stepsize=0.05,
ntrajectories=300,
return_trajectories=True,
)
# and plot it again
# +
#Prepare Plot
cmin, cmax = 0., 6
fig = plt.figure(figsize=(15.0, 4.0 ))
bin_size = (grid_ranges[0][1] - grid_ranges[0][0])/(grid_resolutions[0] - 1)
edges = np.linspace(grid_ranges[0][0] - bin_size/2, grid_ranges[0][1] + bin_size/2, grid_resolutions[0] + 1)
centers = np.linspace(grid_ranges[0][0], grid_ranges[0][1], grid_resolutions[0])
#Plot
ax = plt.subplot(1,3,1)
sc = ax.scatter(thetas.T[0],thetas.T[1],c=distances,vmin=cmin, vmax=cmax,s=10,)
cb = plt.colorbar(sc,ax=ax, extend='both')
cb.set_label(r'Distance $d(\theta,\theta_0)$')
ax.set_xlabel(r'$\theta_1$')
ax.set_ylabel(r'$\theta_2$')
ax = plt.subplot(1,3,2)
cm = ax.pcolormesh(
edges, edges, distance_grid.reshape((grid_resolutions[0], grid_resolutions[1])).T,
vmin=cmin, vmax=cmax,
cmap='viridis'
)
cb = plt.colorbar(cm, ax=ax, extend='both')
cb.set_label(r'Distance $d(\theta,\theta_0)$')
ax.set_xlabel(r'$\theta_1$')
ax.set_ylabel(r'$\theta_2$')
ax = plt.subplot(1,3,3)
cm = ax.pcolormesh(
edges, edges, p_values.reshape((grid_resolutions[0], grid_resolutions[1])).T,
norm=matplotlib.colors.LogNorm(vmin=0.01, vmax=1),
cmap='viridis'
)
cb = plt.colorbar(cm, ax=ax, extend='both')
cb.set_label('Expected p-value')
ax.set_xlabel(r'$\theta_1$')
ax.set_ylabel(r'$\theta_2$')
plt.tight_layout()
plt.show()
# -
# ## 3. Compare to other results
# Load previous results and add Information Geometry results
# +
[p_values,mle]=np.load("limits/limits.npy")
p_values["InfoGeo"] = p_values_infogeo.flatten()
mle["InfoGeo"] = 312
# -
# and plot them together with the obtained Information Geometry results
# +
show = "InfoGeo"
bin_size = (grid_ranges[0][1] - grid_ranges[0][0])/(grid_resolutions[0] - 1)
edges = np.linspace(grid_ranges[0][0] - bin_size/2, grid_ranges[0][1] + bin_size/2, grid_resolutions[0] + 1)
centers = np.linspace(grid_ranges[0][0], grid_ranges[0][1], grid_resolutions[0])
fig = plt.figure(figsize=(6,5))
ax = plt.gca()
cmin, cmax = 1.e-2, 1.
pcm = ax.pcolormesh(
edges, edges, p_values[show].reshape((grid_resolutions[0], grid_resolutions[1])).T,
norm=matplotlib.colors.LogNorm(vmin=cmin, vmax=cmax),
cmap='Greys_r'
)
cbar = fig.colorbar(pcm, ax=ax, extend='both')
for i, (label, p_value) in enumerate(six.iteritems(p_values)):
plt.contour(
centers, centers, p_value.reshape((grid_resolutions[0], grid_resolutions[1])).T,
levels=[0.32],
linestyles='-', colors='C{}'.format(i)
)
plt.scatter(
theta_grid[mle[label]][0], theta_grid[mle[label]][1],
s=80., color='C{}'.format(i), marker='*',
label=label
)
plt.legend()
plt.xlabel(r'$\theta_0$')
plt.ylabel(r'$\theta_1$')
cbar.set_label('Expected p-value ({})'.format(show))
plt.tight_layout()
plt.show()
# -
# Finally, we compare the obtained distance $d(\theta,\theta_0)$ with the expected log-likelihood ratio $q(\theta,\theta_0) = E[-2 \log r(x|\theta,\theta_0)|\theta_0]$. We can see that there is an approximately linear relationship.
# +
from scipy.stats.distributions import chi2
#Prepare Plot
cmin, cmax = 0., 6
fig = plt.figure(figsize=(5.0, 5.0 ))
#Plot
ax = plt.subplot(1,1,1)
ax.scatter(chi2.ppf(1-p_values["ALICES"], df=2),distance_grid.flatten()**2,c="red",)
ax.set_xlabel(r'$q(\theta,\theta_0)$ (ALICES)')
ax.set_ylabel(r'$d^2(\theta,\theta_0)$ ')
ax.set_xlim(0,20)
ax.set_ylim(0,20)
plt.tight_layout()
plt.show()
# -
| examples/tutorial_particle_physics/4c_information_geometry.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Deep Neural Network with PyTorch
#
# Credits: \
# https://jovian.ai/aakashns/04-feedforward-nn \
# https://pytorch.org/docs/stable/data.html
import torch
import torchvision
import numpy as np
import matplotlib.pyplot as plt
import torch.nn as nn
import torch.nn.functional as F
from torchvision.datasets import MNIST
from torchvision.transforms import ToTensor
from torchvision.utils import make_grid
from torch.utils.data.dataloader import DataLoader
from torch.utils.data import random_split
# %matplotlib inline
dataset = MNIST(root='data/', download=True, transform=ToTensor())
# +
val_size = 10000
train_size = len(dataset) - val_size
train_ds, val_ds = random_split(dataset, [train_size, val_size])
len(train_ds), len(val_ds)
# -
batch_size=128
train_loader = DataLoader(train_ds, batch_size, shuffle=True, num_workers=4, pin_memory=True)
val_loader = DataLoader(val_ds, batch_size*2, num_workers=4, pin_memory=True)
for images, _ in train_loader:
print('images.shape:', images.shape)
plt.figure(figsize=(16,8))
plt.axis('off')
plt.imshow(make_grid(images, nrow=16).permute((1, 2, 0)))
break
# ### Model
# We'll create a neural network with one hidden layer.
def accuracy(outputs, labels):
_, preds = torch.max(outputs, dim=1)
return torch.tensor(torch.sum(preds == labels).item() / len(preds))
class MnistModel(nn.Module):
"""Feedfoward neural network with 1 hidden layer"""
def __init__(self, in_size, hidden_size, out_size):
super().__init__()
# hidden layer
self.linear1 = nn.Linear(in_size, hidden_size)
# output layer
self.linear2 = nn.Linear(hidden_size, out_size)
def forward(self, xb):
# Flatten the image tensors
xb = xb.view(xb.size(0), -1)
# Get intermediate outputs using hidden layer
out = self.linear1(xb)
# Apply activation function
out = F.relu(out)
# Get predictions using output layer
out = self.linear2(out)
return out
def training_step(self, batch):
images, labels = batch
out = self(images) # Generate predictions
loss = F.cross_entropy(out, labels) # Calculate loss
return loss
def validation_step(self, batch):
images, labels = batch
out = self(images) # Generate predictions
loss = F.cross_entropy(out, labels) # Calculate loss
acc = accuracy(out, labels) # Calculate accuracy
return {'val_loss': loss, 'val_acc': acc}
def validation_epoch_end(self, outputs):
batch_losses = [x['val_loss'] for x in outputs]
epoch_loss = torch.stack(batch_losses).mean() # Combine losses
batch_accs = [x['val_acc'] for x in outputs]
epoch_acc = torch.stack(batch_accs).mean() # Combine accuracies
return {'val_loss': epoch_loss.item(), 'val_acc': epoch_acc.item()}
def epoch_end(self, epoch, result):
print("Epoch [{}], val_loss: {:.4f}, val_acc: {:.4f}".format(epoch, \
result['val_loss'], result['val_acc']))
input_size = 784
hidden_size = 32
num_classes = 10
model = MnistModel(input_size, hidden_size=32, out_size=num_classes)
for t in model.parameters():
print(t.shape)
# +
for images, labels in train_loader:
outputs = model(images)
loss = F.cross_entropy(outputs, labels)
print('Loss:', loss.item())
break
print('outputs.shape : ', outputs.shape)
print('Sample outputs :\n', outputs[:2].data)
# -
# ### Using a GPU
# Check if a GPU is available and the required NVIDIA CUDA drivers are installed:
torch.cuda.is_available()
def get_default_device():
"""Pick GPU if available, else CPU"""
if torch.cuda.is_available():
return torch.device('cuda')
else:
return torch.device('cpu')
device = get_default_device()
device
# Define a function that can move data and model to a chosen device:
def to_device(data, device):
"""Move tensor(s) to chosen device"""
if isinstance(data, (list,tuple)):
return [to_device(x, device) for x in data]
return data.to(device, non_blocking=True)
for images, labels in train_loader:
print(images.shape)
images = to_device(images, device)
print(images.device)
break
# Define a `DeviceDataLoader` class to wrap our existing data loaders and move data to the selected device, as a batches are accessed. Interestingly, we don't need to extend an existing class to create a PyTorch dataloader. All we need is an `__iter__` method to retrieve batches of data, and an `__len__` method to get the number of batches.
class DeviceDataLoader():
"""Wrap a dataloader to move data to a device"""
def __init__(self, dl, device):
self.dl = dl
self.device = device
def __iter__(self):
"""Yield a batch of data after moving it to device"""
for b in self.dl:
yield to_device(b, self.device)
def __len__(self):
"""Number of batches"""
return len(self.dl)
train_loader = DeviceDataLoader(train_loader, device)
val_loader = DeviceDataLoader(val_loader, device)
# Tensors that have been moved to the GPU's RAM have a `device` property which includes the word `cuda`. Let's verify this by looking at a batch of data from `valid_dl`.
for xb, yb in val_loader:
print('xb.device:', xb.device)
print('yb:', yb)
break
# ### Training the model
# +
def evaluate(model, val_loader):
outputs = [model.validation_step(batch) for batch in val_loader]
return model.validation_epoch_end(outputs)
def fit(epochs, lr, model, train_loader, val_loader, opt_func=torch.optim.SGD):
history = []
optimizer = opt_func(model.parameters(), lr)
for epoch in range(epochs):
# Training Phase
for batch in train_loader:
loss = model.training_step(batch)
loss.backward()
optimizer.step()
optimizer.zero_grad()
# Validation phase
result = evaluate(model, val_loader)
model.epoch_end(epoch, result)
history.append(result)
return history
# -
# Before we train the model, we need to ensure that the data and the model's parameters (weights and biases) are on the same device (CPU or GPU). We can reuse the `to_device` function to move the model's parameters to the right device.
model = MnistModel(input_size, hidden_size=hidden_size, out_size=num_classes)
to_device(model, device)
history = [evaluate(model, val_loader)]
history
history += fit(5, 0.5, model, train_loader, val_loader)
history += fit(5, 0.1, model, train_loader, val_loader)
losses = [x['val_loss'] for x in history]
plt.plot(losses, '-x')
plt.xlabel('epoch')
plt.ylabel('loss')
plt.title('Loss vs. No. of epochs')
accuracies = [x['val_acc'] for x in history]
plt.plot(accuracies, '-x')
plt.xlabel('epoch')
plt.ylabel('accuracy')
plt.title('Accuracy vs. No. of epochs')
| MLP/pytorch-deep-neural-network.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lesson 2 - Image Classification Models from Scratch
# ## Lesson Video:
#hide_input
from IPython.lib.display import YouTubeVideo
YouTubeVideo('_SKqrTlXNt8')
#hide
#Run once per session
# !pip install fastai wwf -q --upgrade
#hide_input
from wwf.utils import state_versions
state_versions(['fastai', 'fastcore', 'wwf'])
# Grab our vision related libraries
from fastai.vision.all import *
# Below you will find the exact imports for everything we use today
# +
from torch import nn
from fastai.callback.hook import summary
from fastai.callback.schedule import fit_one_cycle, lr_find
from fastai.callback.progress import ProgressCallback
from fastai.data.core import Datasets, DataLoaders, show_at
from fastai.data.external import untar_data, URLs
from fastai.data.transforms import Categorize, GrandparentSplitter, parent_label, ToTensor, IntToFloatTensor, Normalize
from fastai.layers import Flatten
from fastai.learner import Learner
from fastai.metrics import accuracy, CrossEntropyLossFlat
from fastai.vision.augment import CropPad, RandomCrop, PadMode
from fastai.vision.core import PILImageBW
from fastai.vision.utils import get_image_files
# -
# And our data
path = untar_data(URLs.MNIST)
# ## Working with the data
items = get_image_files(path)
items[0]
# Create an image object. Done automatically with `ImageBlock`.
im = PILImageBW.create(items[0])
im.show()
# Split our data with `GrandparentSplitter`, which will make use of a `train` and `valid` folder.
splits = GrandparentSplitter(train_name='training', valid_name='testing')
items[:3]
# Splits need to be applied to some items
splits = splits(items)
splits[0][:5], splits[1][:5]
# * Make a `Datasets`
#
# * Expects items, transforms for describing our problem, and a splitting method
dsrc = Datasets(items, tfms=[[PILImageBW.create], [parent_label, Categorize]],
splits=splits)
# We can look at an item in our `Datasets` with `show_at`
show_at(dsrc.train, 3)
# We can see that it's a `PILImage` of a three, along with a label of `3`
# Next we need to give ourselves some transforms on the data! These will need to:
# 1. Ensure our images are all the same size
# 2. Make sure our output are the `tensor` our models are wanting
# 3. Give some image augmentation
tfms = [ToTensor(), CropPad(size=34, pad_mode=PadMode.Zeros), RandomCrop(size=28)]
# * `ToTensor`: Converts to tensor
# * `CropPad` and `RandomCrop`: Resizing transforms
# * Applied on the `CPU` via `after_item`
gpu_tfms = [IntToFloatTensor(), Normalize()]
# * `IntToFloatTensor`: Converts to a float
# * `Normalize`: Normalizes data
dls = dsrc.dataloaders(bs=128, after_item=tfms, after_batch=gpu_tfms)
# And show a batch
dls.show_batch()
# From here we need to see what our model will expect
xb, yb = dls.one_batch()
# And now the shapes:
xb.shape, yb.shape
dls.c
# So our input shape will be a [128 x 1 x 28 x 28] and our output shape will be a [128] tensor that we need to condense into 10 classes
# ## The Model
#
# Our models are made up of **layers**, and each layer represents a matrix multiplication to end up with our final `y`. For this image problem, we will use a **Convolutional layer**, a **Batch Normalization layer**, an **Activation Function**, and a **Flattening layer**
# ### Convolutional Layer
#
# These are always the first layer in our network. I will be borrowing an analogy from [here](https://adeshpande3.github.io/A-Beginner%27s-Guide-To-Understanding-Convolutional-Neural-Networks/) by <NAME>.
#
# Our example Convolutional layer will be 5x5x1
#
# Imagine a flashlight that is shining over the top left of an image, which covers a 5x5 section of pixels at one given moment. This flashlight then slides crosses our pixels at all areas in the picture. This flashlight is called a **filter**, which can also be called a **neuron** or **kernel**. The region it is currently looking over is called a **receptive field**. This filter is also an array of numbers called **weights** (or **parameters**). The depth of this filter **must** be the same as the depth of our input. In our case it is 1 (in a color image this is 3). Now once this filter begins moving (or **convolving**) around the image, it is multiplying the values inside this filter with the original pixel value of our image (also called **element wise multiplications**). These are then summed up (in our case this is just one multiplication of 28x28) to an individual value, which is a representation of **just** the top left of our image. Now repeat this until every unique location has a number and we will get what is called an **activation** or **feature map**. This feature map will be 784 different locations, which turns into a 28x28 array
#
# In our model the Convolutional layer will 3x3 instead (kernel_size = 3) and it will move 2 pixels instead of 1 during each step (stride = 2) resulting in 14x14 feature map (as can be seen in `learner.summary()` below. To fully understand convolution layers and their parameters, there is an excellent tutorial [here](https://arxiv.org/pdf/1603.07285.pdf).
def conv(ni, nf): return nn.Conv2d(ni, nf, kernel_size=3, stride=2, padding=1, bias=False)
# Here we can see our `ni` is equivalent to the depth of the filter, and `nf` is equivalent to how many filters we will be using. (Fun fact this always has to be divisible by the size of our image).
# ### Batch Normalization
#
# As we send our tensors through our model, it is important to normalize our data throughout the network. Doing so can allow for a much larger improvement in training speed, along with allowing each layer to learn independantly (as each layer is then re-normalized according to it's outputs)
def bn(nf): return nn.BatchNorm2d(nf)
# `nf` will be the same as the filter output from our previous convolutional layer
# ### Activation functions
#
# They give our models non-linearity and work with the `weights` we mentioned earlier along with a `bias` through a process called **back-propagation**. These allow our models to learn and perform more complex tasks because they can choose to fire or activate one of those neurons mentioned earlier. On a simple sense, let's look at the `ReLU` activation function. It operates by turning any negative values to zero, as visualized below:
#
# 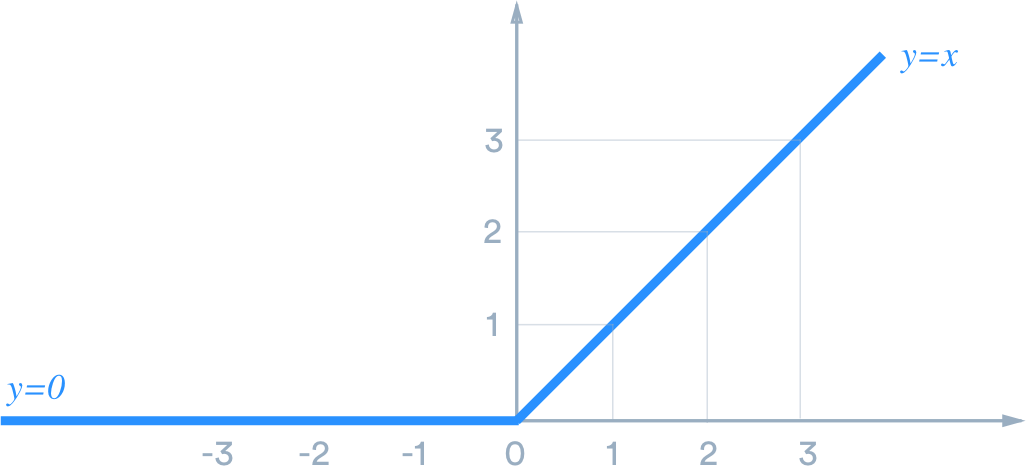
# From "A Practical Guide to ReLU by <NAME>u [URL](https://medium.com/@danqing/a-practical-guide-to-relu-b83ca804f1f7).
def ReLU(): return nn.ReLU(inplace=False)
# ### Flattening
#
# The last bit we need to do is take all these activations and this outcoming matrix and flatten it into a single dimention of predictions. We do this with a `Flatten()` module
# +
# Flatten??
# -
# ## Making a Model
#
# * Five convolutional layers
# * `nn.Sequential`
# * 1 -> 32 -> 10
model = nn.Sequential(
conv(1, 8),
bn(8),
ReLU(),
conv(8, 16),
bn(16),
ReLU(),
conv(16,32),
bn(32),
ReLU(),
conv(32, 16),
bn(16),
ReLU(),
conv(16, 10),
bn(10),
Flatten()
)
# Now let's make our `Learner`
learn = Learner(dls, model, loss_func=CrossEntropyLossFlat(), metrics=accuracy)
# We can then also call `learn.summary` to take a look at all the sizes with their **exact** output shapes
learn.summary()
# `learn.summary` also tells us:
# * Total parameters
# * Trainable parameters
# * Optimizer
# * Loss function
# * Applied `Callbacks`
learn.lr_find()
# Let's use a learning rate around 1e-1 (0.1)
learn.fit_one_cycle(3, lr_max=1e-1)
# ## Simplify it
# * Try to make it more like `ResNet`.
# * `ConvLayer` contains a `Conv2d`, `BatchNorm2d`, and an activation function
def conv2(ni, nf): return ConvLayer(ni, nf, stride=2)
# And make a new model
net = nn.Sequential(
conv2(1,8),
conv2(8,16),
conv2(16,32),
conv2(32,16),
conv2(16,10),
Flatten()
)
# Great! That looks much better to read! Let's make sure we get (roughly) the same results with it.
learn = Learner(dls, net, loss_func=CrossEntropyLossFlat(), metrics=accuracy)
learn.fit_one_cycle(3, lr_max=1e-1)
# Almost the exact same! Perfect! Now let's get a bit more advanced
# ## ResNet (kinda)
#
# The ResNet architecture is built with what are known as ResBlocks. Each of these blocks consist of two `ConvLayers` that we made before, where the number of filters do not change. Let's generate these layers.
class ResBlock(Module):
def __init__(self, nf):
self.conv1 = ConvLayer(nf, nf)
self.conv2 = ConvLayer(nf, nf)
def forward(self, x): return x + self.conv2(self.conv1(x))
# * Class notation
# * `__init__`
# * `foward`
# Let's add these in between each of our `conv2` layers of that last model.
net = nn.Sequential(
conv2(1,8),
ResBlock(8),
conv2(8,16),
ResBlock(16),
conv2(16,32),
ResBlock(32),
conv2(32,16),
ResBlock(16),
conv2(16,10),
Flatten()
)
net
# Awesome! We're building a pretty substantial model here. Let's try to make it **even simpler**. We know we call a convolutional layer before each `ResBlock` and they all have the same filters, so let's make that layer!
def conv_and_res(ni, nf): return nn.Sequential(conv2(ni, nf), ResBlock(nf))
net = nn.Sequential(
conv_and_res(1,8),
conv_and_res(8,16),
conv_and_res(16,32),
conv_and_res(32,16),
conv2(16,10),
Flatten()
)
# And now we have something that resembles a ResNet! Let's see how it performs
learn = Learner(dls, net, loss_func=CrossEntropyLossFlat(), metrics=accuracy)
learn.lr_find()
# Let's do 1e-1 again
learn.fit_one_cycle(3, lr_max=1e-1)
| nbs/course2020/vision/02_MNIST.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# #!/usr/bin/env python3
import pandas
import lz4.frame
import gzip
import io
import pyarrow.parquet as pq
import pyarrow as pa
import numpy as np
import datetime
import matplotlib.pyplot as plt
from glob import glob
from plumbum.cmd import rm
# +
def load_data(filename, filter_initial=True):
df = pq.read_table(filename).to_pandas()
if filter_initial:
df = df[df['Event Type'] != 'Initial']
return df
def get_second_data(df, current_second):
time = sec2string(current_second)
return df.loc[df['Event Time'].values == time]
def get_minute_data(df, current_minute):
time = min2string(current_minute)
next_time = min2string(current_minute + 1)
return df.loc[(df['Event Time'].values >= time) & (df['Event Time'].values < next_time)]
# +
def sec2string(sec):
m, s = divmod(sec, 60)
h, m = divmod(m, 60)
return "%02d:%02d:%02d" %(h, m, s)
def min2string(minute):
h, m = divmod(minute, 60)
return "%02d:%02d:00" %(h, m)
# +
def get_avg_price(df_chunk, percent_change, prev_price, when):
df_chunk = filter_df(df_chunk, event_type='Fill')
if len(df_chunk) == 0:
current_avg_price = prev_price
else:
if when == 'start':
current_avg_price = df_chunk.iloc[0, -1]
elif when == 'end':
current_avg_price = df_chunk.iloc[-1, -1]
return current_avg_price
def calc_percent_change(current_price, prev_price):
try:
percent_change = (current_price - prev_price) / prev_price
except:
percent_change = 0.0
return percent_change
# +
def filter_df(df_chunk, side=None, event_type=None, order_type=None):
if side is not None:
df_chunk = df_chunk.loc[df_chunk['Side'].values == side]
if event_type is not None:
df_chunk = df_chunk.loc[df_chunk['Event Type'].values == event_type]
if order_type is not None:
df_chunk = df_chunk.loc[df_chunk['Order Type'].values == order_type]
return df_chunk
def get_frequency(df_chunk):
return len(df_chunk)
def get_volume(df_chunk, volume_type=None):
if volume_type=='filled':
return sum(df_chunk['Fill Price (USD)'] * df_chunk['Fill Quantity (BTC)'])
if volume_type=='unfilled':
return sum(df_chunk['Limit Price (USD)'] * df_chunk['Original Quantity (BTC)'])
def calculate_percentage(value1, value2):
if value1 == 0.0 and value2 == 0.0:
percentage = 0.5
else:
try:
percentage = value1 / (value1 + value2 + 1e-6)
except:
percentage = None
return percentage
# +
def one_hot(index, length):
onehot = [0.]*length
onehot[index] = 1.
return onehot
def extract_temporal_features(df_chunk):
year, month, day = df_chunk['Event Date'].values[0].split('-')
day_of_week = int(datetime.datetime(int(year), int(month), int(day)).weekday())
hour = int(df_chunk['Event Time'].values[0][0:2])
month = int(month) - 1
return one_hot(month, 12), one_hot(day_of_week, 7), one_hot(hour, 24)
# +
def vol_freq(df_chunk, volume_type):
return get_volume(df_chunk, volume_type=volume_type), get_frequency(df_chunk)
def get_raw_features(df_chunk, side=None):
x = {}
x['vol_markets'], x['freq_markets'] = vol_freq(filter_df(df_chunk, side=side, event_type='Fill', order_type='market'), volume_type='filled')
x['vol_filled_limits'], x['freq_filled_limits'] = vol_freq(filter_df(df_chunk, side=side, event_type='Fill', order_type='limit'), volume_type='filled')
x['vol_placed_limits'], x['freq_placed_limits'] = vol_freq(filter_df(df_chunk, side=side, event_type='Place', order_type='limit'), volume_type='unfilled')
x['vol_cancelled_limits'], x['freq_cancelled_limits'] = vol_freq(filter_df(df_chunk, side=side, event_type='Cancel', order_type='limit'), volume_type='unfilled')
return x
def compute_features(x_buy, x_sell):
# Buys:
# -Volume of Filled Markets vs Filled Limits
# -Volume of Placed Limits vs Filled Limits
# -Frequency of Filled Markets vs Filled Limits
# -Frequency of Placed Limits vs Filled Limits
# Sells:
# -Volume of Filled Markets vs Filled Limits
# -Volume of Placed Limited vs Filled Limits
# -Frequency of Filled Markets vs Filled Limits
# -Frequency of Placed Limits vs Filled Limits
# Buys vs Sells:
# -Volume of Filled Market Sells vs Volume of Filled Market Buys
# -Volume of Placed Limit Sells vs Volume of Placed Limit Buys
# -Volume of Cancelled Limit Sells vs Volume Cancelled Limit Buys
# -Frequency of Filled Market Sells vs Frequency of Filled Market Buys
# -Frequency of Placed Limit Sells vs Frequency of Placed Limit Buys
# -Frequency of Cancelled Limit Sells vs Frequency of Cancelled Limit Buys
features = []
# Buys:
features.append(calculate_percentage(x_buy['vol_markets'], x_buy['vol_filled_limits']))
features.append(calculate_percentage(x_buy['vol_placed_limits'], x_buy['vol_filled_limits']))
features.append(calculate_percentage(x_buy['freq_markets'], x_buy['freq_filled_limits']))
features.append(calculate_percentage(x_buy['freq_placed_limits'], x_buy['freq_filled_limits']))
# Sells:
features.append(calculate_percentage(x_sell['vol_markets'], x_sell['vol_filled_limits']))
features.append(calculate_percentage(x_sell['vol_placed_limits'], x_sell['vol_filled_limits']))
features.append(calculate_percentage(x_sell['freq_markets'], x_sell['freq_filled_limits']))
features.append(calculate_percentage(x_sell['freq_placed_limits'], x_sell['freq_filled_limits']))
# Buys vs Sells:
features.append(calculate_percentage(x_sell['vol_markets'], x_buy['vol_markets']))
features.append(calculate_percentage(x_sell['vol_placed_limits'], x_buy['vol_placed_limits']))
features.append(calculate_percentage(x_sell['vol_cancelled_limits'], x_buy['vol_cancelled_limits']))
features.append(calculate_percentage(x_sell['freq_markets'], x_buy['freq_markets']))
features.append(calculate_percentage(x_sell['freq_placed_limits'], x_buy['freq_placed_limits']))
features.append(calculate_percentage(x_sell['freq_cancelled_limits'], x_buy['freq_cancelled_limits']))
return features
# -
def get_all_features(df_chunk, percent_change, prev_price):
# Current price, percent change
current_price = get_avg_price(df_chunk, percent_change, prev_price, when='end')
percent_change = calc_percent_change(current_price, prev_price)
feature_vec = [current_price, percent_change]
# Order book features
x_buy = get_raw_features(df_chunk, side='buy')
x_sell = get_raw_features(df_chunk, side='sell')
feature_vec.extend(compute_features(x_buy, x_sell))
# Temporal features
month_vec, day_vec, hour_vec = extract_temporal_features(df_chunk)
feature_vec.extend(month_vec)
feature_vec.extend(day_vec)
feature_vec.extend(hour_vec)
return feature_vec
def write_tmp_parquet(df, outfile):
outfile = outfile.replace('cboe/parquet_BTCUSD/', 'cboe/parquet_preprocessed_BTCUSD/')
pq.write_table(pa.Table.from_pandas(df), outfile, compression='snappy')
def preprocess_day(filename, visualize=True, write_parquet=False, verbose=True):
print(filename)
df = load_data(filename, filter_initial=True)
# Initialize previous price
percent_change=0.0
prev_price = get_avg_price(df, percent_change=None, prev_price=None, when='start')
# Compute feature vector for each minute of the day
all_X = []
for minute in range(24*60):
if verbose:
if minute%100 == 0:
print('Minutes:', minute)
# Select one minute of data from order book
df_chunk = get_minute_data(df, minute)
if len(df_chunk) == 0: # skip minutes with no data
continue
# Extract features, X
X = get_all_features(df_chunk, percent_change, prev_price)
prev_price = X[0]
percent_change = X[1]
#all_X.append(X[1:])
all_X.append(X)
#columns = ['current_price','percent_change',
columns = ['current_price', 'percent_change',
'buy_vol_mark_vs_fillLim','buy_vol_placeLim_vs_fillLim','buy_freq_mark_vs_fillLim','buy_freq_placeLim_vs_fillLim',
'sell_vol_mark_vs_fillLim','sell_vol_placeLim_vs_fillLim','sell_freq_mark_vs_fillLim','sell_freq_placeLim_vs_fillLim',
'vol_markSells_vs_markBuys','vol_placeLimSells_vs_placeLimBuys','vol_CancelLimSells_vs_CancelLimBuys',
'freq_markSells_vs_markBuys','freq_placeLimSells_vs_placeLimBuys','freq_CancelLimSells_vs_CancelLimBuys',
'm0','m1','m2','m3','m4','m5','m6','m7','m8','m9','m10','m11',
'd0','d1','d2','d3','d4','d5','d6',
'h0','h1','h2','h3','h4','h5','h6','h7','h8','h9','h10','h11',
'h12','h13','h14','h15','h16','h17','h18','h19','h20','h21','h22','h23']
# Convert to pandas DF
new_df = pandas.DataFrame.from_records(all_X, columns=columns)
# Compute labels, Y
new_df = calculate_y(new_df)
# Write DF to tmp file to later be concatenated with all others
if write_parquet:
write_tmp_parquet(new_df, filename)
# Visualize
if visualize:
visualize_features(new_df)
return new_df
def check_number_of_events(df, timesteps, resolution='minute', event_type=None, order_type=None):
# Filter data
if event_type is not None:
df = df.loc[df['Event Type'].values == event_type]
if order_type is not None:
df = df.loc[df['Order Type'].values == order_type]
chunk_lengths = []
# Minute resolution
if resolution == 'minute':
for minute in range(timesteps):
chunk_lengths.append(len(get_minute_data(df, minute)))
if minute%200 == 0:
print('Minute:', minute)
# Second resolution
elif resolution == 'second':
for sec in range(timesteps):
chunk_lengths.append(len(get_second_data(df, sec)))
if sec%100 == 0:
print('Second', sec)
# Visualize
plt.figure(figsize=(20,2));
plt.plot(chunk_lengths);
plt.figure(figsize=(20,5));
plt.hist(chunk_lengths, bins=40);
return chunk_lengths
def calculate_y(new_df):
new_df['y_percent_change'] = new_df.iloc[:,1]
new_df['y_percent_change'] = new_df['y_percent_change'].shift(-1)
new_df = new_df[:-1]
return new_df
def visualize_features(df):
for column_idx in range(16):
title = df.columns[column_idx]
df.plot(y = column_idx, figsize=(20,2), title=title)
# +
# # Visualize events
# event_type = "Fill"
# order_type = None
# resolution = 'minute'
# timesteps = 24*60
# chunk_lengths = check_number_of_events(df, timesteps=timesteps, resolution=resolution,
# event_type=event_type, order_type=order_type)
# -
# # Preprocess all files
count = 0
filenames = sorted(glob('cboe/parquet_BTCUSD/*.parquet'))
filenames.reverse()
for day in range(len(filenames)):
filename = filenames[day]
new_df = preprocess_day(filename, write_parquet=True, visualize=False, verbose=False)
count += 1
print(count, '/', len(filenames))
| preprocessing_final.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="v8KR9V4Hy-vw"
# # Imports
# + cellView="both" colab={} colab_type="code" id="idfu7sA0vExR"
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import sys
assert sys.version_info.major == 3
import os
add_paths = True
if add_paths:
sys.path.insert(0, os.path.join(os.path.abspath(os.getcwd()), '..', '..'))
sys.path.insert(
0,
os.path.join(os.path.abspath(os.getcwd()), '..', '..', 'build', 'python'))
import pyspiel
from google3.third_party.open_spiel.python.algorithms import cfr
from google3.third_party.open_spiel.python.algorithms import exploitability
from google3.third_party.open_spiel.python.algorithms import expected_game_score
from google3.third_party.open_spiel.python.bots import uniform_random
from google3.third_party.open_spiel.python.visualizations import treeviz
# + colab={} colab_type="code" id="HLXNc0ZCvExt"
games_list = pyspiel.registered_names()
print("Registered games:")
print(games_list)
game = pyspiel.load_game("universal_poker")
# + colab={} colab_type="code" id="vqyfMHs2vEx7"
"""Test that Python and C++ bots can be called by a C++ algorithm."""
from absl.testing import absltest
import numpy as np
from google3.third_party.open_spiel.python.bots import uniform_random
game = pyspiel.load_game("leduc_poker")
bots = [
pyspiel.make_uniform_random_bot(0, 1234),
uniform_random.UniformRandomBot(1, np.random.RandomState(4321)),
]
results = np.array([
pyspiel.evaluate_bots(game.new_initial_state(), bots, iteration)
for iteration in range(10000)
])
leduc_average_results = np.mean(results, axis=0)
print(leduc_average_results)
game = pyspiel.load_game("universal_poker")
bots = [
pyspiel.make_uniform_random_bot(0, 1234),
uniform_random.UniformRandomBot(1, np.random.RandomState(4321)),
]
results = np.array([
pyspiel.evaluate_bots(game.new_initial_state(), bots, iteration)
for iteration in range(10000)
])
universal_poker_average_results = np.mean(results, axis=0)
print(universal_poker_average_results)
#np.testing.assert_allclose(universal_poker_average_results, leduc_average_results, atol=0.1)
# + colab={} colab_type="code" id="RhI6kVnkvEyE"
universal_poker_kuhn_limit_3p = """\
GAMEDEF
limit
numPlayers = 3
numRounds = 1
blind = 1 1 1
raiseSize = 1
firstPlayer = 1
maxRaises = 1
numSuits = 1
numRanks = 4
numHoleCards = 1
numBoardCards = 0
END GAMEDEF
"""
game = pyspiel.load_game(
"universal_poker",
{"gamedef": pyspiel.GameParameter(universal_poker_kuhn_limit_3p)})
str(game)
# + colab={} colab_type="code" id="lpLJhzBEvEyM"
# Compare exloitability for two games
players = 2
iterations = 10
print_freq = 1
def compare_exploitability(game_1, game_2):
cfr_solver_1 = cfr.CFRSolver(game_1)
cfr_solver_2 = cfr.CFRSolver(game_2)
for i in range(iterations):
cfr_solver_1.evaluate_and_update_policy()
cfr_solver_2.evaluate_and_update_policy()
if i % print_freq == 0:
conv_1 = exploitability.exploitability(game_1,
cfr_solver_1.average_policy())
conv_2 = exploitability.exploitability(game_2,
cfr_solver_2.average_policy())
print("Iteration {} exploitability of the {} vs: {}".format(
i, conv_1, conv_2))
print("Final exploitability is {} vs {}".format(conv_1, conv_2))
game_1 = pyspiel.load_game("kuhn_poker",
{"players": pyspiel.GameParameter(2)})
universal_poker_kuhn_limit_2p = """\
GAMEDEF
limit
numPlayers = 2
numRounds = 1
blind = 1 1
raiseSize = 1
firstPlayer = 1
maxRaises = 1
numSuits = 1
numRanks = 3
numHoleCards = 1
numBoardCards = 0
END GAMEDEF
"""
game_2 = pyspiel.load_game(
"universal_poker",
{"gamedef": pyspiel.GameParameter(universal_poker_kuhn_limit_2p)})
compare_exploitability(game_1, game_2)
# + colab={} colab_type="code" id="0Zltqy5PNM8P"
game_1 = pyspiel.load_game("leduc_poker",
{"players": pyspiel.GameParameter(2)})
# Taken verbatim from the linked paper above: "In Leduc hold'em, the deck
# consists of two suits with three cards in each suit. There are two rounds.
# In the first round a single private card is dealt to each player. In the
# second round a single board card is revealed. There is a two-bet maximum,
# with raise amounts of 2 and 4 in the first and second round, respectively.
# Both players start the first round with 1 already in the pot.
universal_poker_leduc_limit_2p = """\
GAMEDEF
limit
numPlayers = 2
numRounds = 2
blind = 1 1
raiseSize = 1 1
firstPlayer = 1 1
maxRaises = 2 2
raiseSize = 2 4
numSuits = 2
numRanks = 3
numHoleCards = 1 0
numBoardCards = 0 1
END GAMEDEF
"""
game_2 = pyspiel.load_game(
"universal_poker",
{"gamedef": pyspiel.GameParameter(universal_poker_leduc_limit_2p)})
compare_exploitability(game_1, game_2)
# + colab={} colab_type="code" id="zk4rz8mvvEyb"
game = "universal_poker"
out = "/tmp/gametree.png"
prog = "dot"
group_infosets = False
group_terminal = False
verbose = False
def _zero_sum_node_decorator(state):
"""Custom node decorator that only shows the return of the first player."""
attrs = treeviz.default_node_decorator(state) # get default attributes
if state.is_terminal():
attrs["label"] = str(int(state.returns()[0]))
return attrs
game = pyspiel.load_game(
game, {"gamedef": pyspiel.GameParameter(universal_poker_kuhn_limit_2p)})
game_type = game.get_type()
if game_type.dynamics != pyspiel.GameType.Dynamics.SEQUENTIAL:
raise ValueError("Game must be sequential, not {}".format(game_type.dynamics))
if (game_type.utility == pyspiel.GameType.Utility.ZERO_SUM and
game.num_players() == 2):
gametree = treeviz.GameTree(
game,
node_decorator=_zero_sum_node_decorator,
group_infosets=group_infosets,
group_terminal=group_terminal)
else:
gametree = treeviz.GameTree(game) # use default decorators
if verbose:
logging.info("Game tree:\n%s", gametree.to_string())
gametree.draw(out, prog=prog)
# + colab={} colab_type="code" id="4rvvGu65M1jk"
| third_party/open_spiel/colabs/test_universal_poker.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Check if notebook-minio integration is working
# !pip install awscli s3fs pandas -q
# !aws --endpoint-url $MINIO_ENDPOINT_URL s3 ls
# !aws --endpoint-url $MINIO_ENDPOINT_URL s3 mb s3://bpk-nb-minio
# !aws --endpoint-url $MINIO_ENDPOINT_URL s3 ls
# !aws --endpoint-url $MINIO_ENDPOINT_URL s3 ls s3://bpk-nb-minio/
# !aws --endpoint-url $MINIO_ENDPOINT_URL s3 cp sample.txt s3://bpk-nb-minio/uploaded-sample.txt
# !aws --endpoint-url $MINIO_ENDPOINT_URL s3 ls s3://bpk-nb-minio/
# !aws --endpoint-url $MINIO_ENDPOINT_URL s3 cp s3://bpk-nb-minio/uploaded-sample.txt downloaded-sample.txt
# Now, go to the MinIO console or use CLI to check if values are in expected MinIO object storage.
# !ls
# # Check if pandas can download data from MinIO
# !aws --endpoint-url $MINIO_ENDPOINT_URL s3 cp sample.csv s3://bpk-nb-minio/uploaded.csv
import pandas as pd
import os
pd.read_csv("s3://bpk-nb-minio/uploaded.csv", delimiter=";",storage_options={
'key': os.environ['AWS_ACCESS_KEY_ID'],
'secret': os.environ['AWS_SECRET_ACCESS_KEY'],
'client_kwargs':{
'endpoint_url': os.environ['MINIO_ENDPOINT_URL']
}
})
# # Clean up
# !aws --endpoint-url $MINIO_ENDPOINT_URL s3 rm s3://bpk-nb-minio/uploaded-sample.txt
# !aws --endpoint-url $MINIO_ENDPOINT_URL s3 rm s3://bpk-nb-minio/uploaded.csv
# !aws --endpoint-url $MINIO_ENDPOINT_URL s3 rb s3://bpk-nb-minio
# rm downloaded-sample.txt
| notebook-integrations/minio-integration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Second Assignment
# #### 1) Create a function called **"even_squared"** that receives an integer value **N**, and returns a list containing, in ascending order, the square of each of the even values, from 1 to N, including N if applicable.
# +
def even_squared(N):
'''Receives a positive integer N and returns a list of the even squares from 1 to N'''
return [x**2 for x in range(2,N+1,2)]
print(even_squared(15))
# -
# #### 2) Using a while loop and the **input()** function, read an indefinite amount of **integers** until the number read is **-1**. After this process, print two lists on the screen: The first containing the even integers, and the second containing the odd integers. Both must be in ascending order.
even, odd = [], []
while 1:
number_read = int(input("Insert an integer: "))
if number_read == -1:
break
if number_read%2:
odd.append(number_read)
else:
even.append(number_read)
print(sorted(even))
print(sorted(odd))
# #### 3) Create a function called **"even_account"** that receives a list of integers, counts the number of existing even elements, and returns this count.
# +
def even_account(l):
'''Receives a list of integers and returns the number of even numbers in it'''
return len(list(filter(lambda x:x%2==0,l)))
print(even_account([1,5,2,6,8,13,27,154,20,22,25,17]))
# -
# #### 4) Create a function called **"squared_list"** that receives a list of integers and returns another list whose elements are the squares of the elements of the first.
# +
def squared_list(l):
'''Receive a list of integers and returns another list with the squared elements from the original list'''
return list(map(lambda x: x**2, l))
# return [element ** 2 for element in l]
print(squared_list([1,3,4,2,6]))
# -
# #### 5) Create a function called **"descending"** that receives two lists of integers and returns a single list, which contains all the elements in descending order, and may include repeated elements.
# +
def descending(A,B):
'''Receives two lists of integers and returns a joint list of elements in descending order'''
C = A + B
C.sort(reverse=True)
return C
print(descending([1,3,2,4,7],[0,6,5,2,1,3]))
# -
# #### 6) Create a function called **"adding"** that receives a list **A**, and an arbitrary number of integers as input. Return a new list containing the elements of **A** plus the integers passed as input, in the order in which they were given. Here is an example:
#
# >```python
# >>>> A = [10,20,30]
# >>>> adding(A, 4, 10, 50, 1)
# > [10, 20, 30, 4, 10, 50, 1]
# ```
# +
def adding(A,*integers):
'''Receives a list of integers and an arbitray amount of integers and returns
a new list with the original list extended with the given integers'''
A.extend(integers)
return A
print(adding([5,10,15,20,11],25,30,35,40,45,127,12))
# -
# #### 7) Create a function called **"intersection"** that receives two input lists and returns another list with the values that belong to the two lists simultaneously (intersection) without repetition of values and in ascending order. Use only lists (do not use sets); loops and conditionals. See the example:
#
# >```python
# >>>> A = [-2, 0, 1, 2, 3]
# >>>> B = [-1, 2, 3, 6, 8]
# >>>> intersection(A,B)
# > [2, 3]
# ```
# +
def intersection(A,B):
'''Receives two lists and returns a new list
with the common elements in ascending order'''
C = [value for value in A if value in B]
res = []
[res.append(x) for x in C if x not in res]
return sorted(res)
print(intersection([1,3,2,2,7,4],[2,4,8,16]))
# -
# #### 8) Create a function called **"union"** that receives two input lists and returns another list with the union of the elements of the two received, without repetition of elements and in ascending order. Use only lists (do not use sets); loops and conditionals. See the example:
#
# >```python
# >>>> A = [-2, 0, 1, 2]
# >>>> B = [-1, 1, 2, 10]
# >>>> union(A,B)
# > [-2, ,-1, 0, 1, 2, 10]
# ```
# +
def union(A,B):
'''Receives two lists of integers and returns a new list with all
elements from both, in ascending order, without repetitions'''
C = A + B
res = []
[res.append(x) for x in C if x not in res]
return sorted(res)
print(union([1,3,2],[4,6,5,3,2]))
# -
# #### 9) Generalize the **"intersection"** function so that it receives an indefinite number of lists and returns the intersection of all of them. Call the new function **intersection2**.
# +
import functools
def intersection2(*lists):
'''Receives an arbitrary number of lists of integers and returns
a new list with the elements common to all of them, without repetitions
and in ascending order'''
#return functools.reduce(intercept,lists) #one way to do
l1 = lists[0] #another way to do
for l2 in lists:
l1 = list(set(l1) & set(l2))
return l1
print(intersection2([1,2,3,4,5,6],[1,2,4,8],[2,4,6,8],[2,3,4,5,6]))
# -
# ## Challenge
#
# #### 10) Create a function named **"matrix"** that implements matrix multiplication:
#
# Given the matrices:
#
# $A_{m\times n}=
# \left[\begin{matrix}
# a_{11}&a_{12}&...&a_{1n}\\
# a_{21}&a_{22}&...&a_{2n}\\
# \vdots &\vdots &&\vdots\\
# a_{m1}&a_{m2}&...&a_{mn}\\
# \end{matrix}\right]$
#
# We will represent then as a list of lists.
#
# $A = [[a_{11},a_{12},...,a_{1n}],[a_{21},a_{22},...,a_{2n}], . . . ,[a_{m1},a_{m2},...,a_{mn}]]$
#
# The **"matrix"** funtion must receive two matrices $A$ e $B$ in the specified format and return $A\times B$
# +
def matrix(A, B):
'''Receives two matrices (lists of lists) and returns the
matrix multiplication, also as a list of lists '''
if len(A)!=len(B[0]) or len(A[0])!=len(B):
raise ValueError('Incompatible dimensions')
m, n , lst, C, element= len(B[0]), len(A), [], [], 0
for i in range(n):
for j in range(m):
for k in range(len(B)):
element+=A[i][k]*B[k][j]
lst.append(element)
element = 0
C.append(lst)
lst= []
return C
X=[[1,0],[0,1],[3,0]]
Y=[[5,2,1],[2,1,1]]
print(matrix(X,Y))
# -
matrix([[1,2],[4,5]],[[1,2],[2,3],[3,4]])
# ## Unit Tests for solutions
# +
import unittest
class TestLista02(unittest.TestCase):
def test_even_squared(self):
self.assertEqual(even_squared(25),[4, 16, 36, 64, 100, 144, 196, 256, 324, 400, 484, 576])
self.assertEqual(even_squared(1),[])
self.assertEqual(even_squared(123),[4, 16, 36, 64, 100, 144, 196, 256, 324, 400, 484, 576,\
676, 784, 900, 1024, 1156, 1296, 1444, 1600, 1764, 1936,\
2116, 2304, 2500, 2704, 2916, 3136, 3364, 3600, 3844, 4096,\
4356, 4624, 4900, 5184, 5476, 5776, 6084, 6400, 6724, 7056,\
7396, 7744, 8100, 8464, 8836, 9216, 9604, 10000, 10404, 10816,\
11236, 11664, 12100, 12544, 12996, 13456, 13924, 14400, 14884])
'''Question 2 manually corrected'''
def test_even_account(self):
self.assertEqual(even_account([1,2,3,8,46,12,0,13,27,135,26,168,1649]),7)
self.assertEqual(even_account([827, 647, 664, 270, 561, 665, 781, 69, 245, 324, 957, 911, 213, 9, 93]),3)
self.assertEqual(even_account([422, 11, 469, 340, 717, 310, 956, 220, 557, 804, 751, 908, 917, 524, 234, 308]),10)
def test_squared_list(self):
self.assertEqual(squared_list([697, 983, 600, 651, 747, 884, 769]),\
[485809, 966289, 360000, 423801, 558009, 781456, 591361])
self.assertEqual(squared_list([96, 89, 35, 38, 19, 12, 63, 24, 81]),\
[9216, 7921, 1225, 1444, 361, 144, 3969, 576, 6561])
self.assertEqual(squared_list([4, 52, 95, 71, 64, 42, 44, 9, 77, 41]),\
[16, 2704, 9025, 5041, 4096, 1764, 1936, 81, 5929, 1681])
def test_descending(self):
self.assertEqual(descending([54, 23, 9, 63, 26, 60, 85, 84, 62, 42, 28, 23, 64, 2, 73],\
[46, 94, 89, 14, 2, 40, 1, 3, 10, 7]),\
[94, 89, 85, 84, 73, 64, 63, 62, 60, 54, 46, 42, 40, 28, 26, 23, 23, 14, 10, 9, 7, 3, 2, 2, 1])
self.assertEqual(descending([69, 52, 4, 21, 51],\
[71, 7, 86, 100, 18, 90]),\
[100, 90, 86, 71, 69, 52, 51, 21, 18, 7, 4])
self.assertEqual(descending([74, 22, 32, 52, 45, 29, 99, 54, 3, 21, 54, 5, 26],\
[58, 72, 16, 69, 56, 49, 6, 52]),\
[99, 74, 72, 69, 58, 56, 54, 54, 52, 52, 49, 45, 32, 29, 26, 22, 21, 16, 6, 5, 3])
def test_adding(self):
self.assertEqual(adding([95, 23],51, 51, 96, 47, 45, 23, 64, 2, 10, 75, 49),\
[95, 23, 51, 51, 96, 47, 45, 23, 64, 2, 10, 75, 49])
self.assertEqual(adding([47, 99, 22, 45, 18, 90, 81, 27, 79, 49, 52, 33, 95],33, 88, 89, 93, 9, 29, 95, 94, 12, 36, 6, 0),\
[47, 99, 22, 45, 18, 90, 81, 27, 79, 49, 52, 33, 95, 33, 88, 89, 93, 9, 29, 95, 94, 12, 36, 6, 0])
self.assertEqual(adding([56, 96, 21, 57, 70],29, 13, 100, 18, 96, 73, 25, 50, 39),\
[56, 96, 21, 57, 70, 29, 13, 100, 18, 96, 73, 25, 50, 39])
def test_intersection(self):
self.assertEqual(intersection([37, 94, 61, 12, 50, 37, 10],[36, 2, 76, 31, 67, 2, 40]),[])
self.assertEqual(intersection([67, 19, 76, 31, 55, 53],[76, 7, 24, 48, 8]),[76])
self.assertEqual(intersection([7, 3, 9, 3, 3, 5, 7, 1],[4, 2, 4, 4, 7, 5, 1, 9]),[1, 5, 7, 9])
def test_union(self):
self.assertEqual(union([1, 3, 42, 23, 14, 28, 86, 86, 0, 68],[52, 18, 11, 14, 27, 38, 79, 51, 38]),\
[0, 1, 3, 11, 14, 18, 23, 27, 28, 38, 42, 51, 52, 68, 79, 86])
self.assertEqual(union([4, 5, 9, 3, 9, 6, 9, 10, 3, 6],[1, 8, 3, 1, 8, 10, 3, 6, 8, 9, 5, 10, 6]),\
[1, 3, 4, 5, 6, 8, 9, 10])
self.assertEqual(union([6, 0, 2, 4, 4, 0, 7, 5, 1, 5],[2, 1, 7]),\
[0, 1, 2, 4, 5, 6, 7])
def test_intersection2(self):
self.assertEqual(intersection2([2, 4, 5, 0, 4, 3, 5, 2, 2, 0, 0, 3],\
[1, 5, 4, 5, 4, 3, 4, 3, 5, 4, 2, 0, 0, 0],\
[0, 0, 5, 1, 4, 4, 4, 3, 4, 2, 1, 5, 3, 3],\
[4, 3, 4, 2, 5, 0, 3, 1, 0, 4, 4],\
[3, 5, 0, 5, 0, 0, 5, 4, 0, 4, 0, 2, 3, 4],\
[2, 0, 1, 0, 5, 2, 3, 2, 4, 5, 2, 4, 2],\
[1, 4, 5, 3, 1, 5, 2, 2, 5, 1, 1, 5, 1, 0, 5],\
[3, 2, 1, 5, 1, 0, 5, 5, 2, 2, 2, 5, 0, 4, 5],\
[4, 5, 3, 3, 4, 1, 1, 1, 0, 5],\
[2, 2, 1, 2, 4, 3, 1, 2, 5, 3, 4, 4, 3]),\
[3, 4, 5])
self.assertEqual(intersection2([4, 4, 5, 1, 0, 3, 0, 4, 5, 2, 5, 2, 3],\
[3, 2, 2, 2, 5, 3, 3, 2, 0, 1, 4],\
[4, 1, 2, 3, 0, 0, 2, 3, 3, 2, 3, 3, 4],\
[1, 3, 1, 4, 4, 5, 4, 4, 5, 2, 5],\
[3, 4, 3, 1, 3, 2, 3, 2, 5, 1, 1],\
[1, 0, 0, 3, 1, 2, 4, 0, 0, 4, 2, 4, 2, 0]),\
[1, 2, 3, 4])
self.assertEqual(intersection2([5, 4, 0, 3, 5, 1, 0, 3, 1, 3],\
[1, 5, 4, 2, 1, 0, 4, 3, 4, 3],\
[4, 5, 3, 3, 5, 2, 2, 3, 0, 0, 4, 4, 5],\
[2, 2, 4, 2, 5, 0, 0, 5, 4, 4, 1, 3, 0, 4],\
[1, 4, 3, 5, 3, 0, 3, 0, 3, 1],\
[3, 1, 3, 4, 2, 2, 4, 2, 0, 0, 3, 3, 5, 3, 3]),\
[0, 3, 4, 5])
def test_matrix(self):
self.assertEqual(matrix([[97, 22, 30, 3, 49, 16, 10, 55, 53], [83, 50, 62, 13, 51, 18, 28, 89, 17],\
[12, 4, 99, 90, 70, 76, 84, 59, 96], [59, 7, 81, 91, 81, 94, 45, 91, 57]],\
[[27, 94, 22, 22], [33, 14, 81, 91], [75, 28, 13, 51], [79, 70, 53, 78],\
[19, 29, 93, 99], [18, 8, 50, 3], [31, 92, 63, 66], [81, 49, 53, 75], [34, 2, 64, 49]]),\
[[13618, 15746, 16759, 18181], [19516, 19742, 20583, 25011], [28336, 23705, 31518, 34528],\
[29023, 26096, 31280, 34053]])
self.assertEqual(matrix([[7, 45, 49, 39, 4], [0, 26, 88, 44, 51], [32, 69, 16, 99, 10], [17, 15, 37, 19, 20]],\
[[56, 77, 33, 39], [40, 94, 1, 95], [20, 55, 55, 72], [40, 75, 86, 13], [20, 29, 82, 25]]),\
[[4812, 10505, 6653, 8683], [5580, 12063, 12832, 10653], [9032, 17545, 11339, 10492], [3452, 6759, 5885, 5499]])
self.assertEqual(matrix([[80, 56], [44, 22], [8, 63]],[[88, 18, 100], [94, 37, 95]]),\
[[12304, 3512, 13320], [5940, 1606, 6490], [6626, 2475, 6785]])
unittest.main(argv=['first-arg-is-ignored'], exit=False)
| Assigments/Assignment_2_solutions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:.conda-py36]
# language: python
# name: conda-env-.conda-py36-py
# ---
# # 00__make_files
#
# in this notebook, i make the files necessary for finding CAGE reads that intersect our regions of interest (orthologous TSSs between human and mouse). final files are BED files with a 50 bp buffer surrounding the TSS (in both human and mouse).
# +
import warnings
warnings.filterwarnings('ignore')
import pandas as pd
import math
import matplotlib.pyplot as plt
import numpy as np
import re
import seaborn as sns
import sys
from scipy.stats import spearmanr
# import utils
sys.path.append("../../../utils")
from plotting_utils import *
# %matplotlib inline
# %config InlineBackend.figure_format = 'svg'
mpl.rcParams['figure.autolayout'] = False
# -
# ## variables
human_master_f = "../../../data/01__design/00__genome_list/hg19.master_list.txt.gz"
mouse_master_f = "../../../data/01__design/00__genome_list/mm9.master_list.txt.gz"
# ## 1. import data
human_master = pd.read_table(human_master_f, sep="\t")
human_master.head()
mouse_master = pd.read_table(mouse_master_f, sep="\t")
mouse_master.head()
# ## 2. filter to seq orths only
human_master_filt = human_master[human_master["seq_orth"]]
len(human_master_filt)
mouse_master_filt = mouse_master[mouse_master["seq_orth"]]
len(mouse_master_filt)
# ## 3. find TSS coords for human/mouse paired regions
# do it for both the "human" file (started from human) and the "mouse" file (started from mouse)
human_bed_hg19 = human_master_filt[["chr_tss_hg19", "start_tss_hg19", "end_tss_hg19", "cage_id_hg19",
"score_tss_hg19", "strand_tss_hg19"]].drop_duplicates()
print(len(human_bed_hg19))
human_bed_hg19.head()
human_bed_mm9 = human_master_filt[["chr_tss_mm9", "start_tss_mm9", "end_tss_mm9", "cage_id_hg19",
"score_tss_hg19", "strand_tss_mm9"]].drop_duplicates()
print(len(human_bed_mm9))
human_bed_mm9.head()
human_bed_mm9[human_bed_mm9["cage_id_hg19"] == "chr1:203273760..203273784,-"]
mouse_bed_mm9 = mouse_master_filt[["chr_tss_mm9", "start_tss_mm9", "end_tss_mm9", "cage_id_mm9",
"score_tss_mm9", "strand_tss_mm9"]].drop_duplicates()
print(len(mouse_bed_mm9))
mouse_bed_mm9.head()
mouse_bed_hg19 = mouse_master_filt[["chr_tss_hg19", "start_tss_hg19", "end_tss_hg19", "cage_id_mm9",
"score_tss_mm9", "strand_tss_hg19"]].drop_duplicates()
print(len(mouse_bed_hg19))
mouse_bed_hg19.head()
# ## 4. group hg19/mm9 files together for bed intersect
human_bed_hg19["cage_id"] = "HUMAN_CAGE_ID__" + human_bed_hg19["cage_id_hg19"]
mouse_bed_hg19["cage_id"] = "MOUSE_CAGE_ID__" + mouse_bed_hg19["cage_id_mm9"]
human_bed_hg19["score"] = "HUMAN_SCORE__" + human_bed_hg19["score_tss_hg19"].astype(str)
mouse_bed_hg19["score"] = "MOUSE_SCORE__" + mouse_bed_hg19["score_tss_mm9"].astype(str)
human_bed_hg19.head()
human_bed_mm9["cage_id"] = "HUMAN_CAGE_ID__" + human_bed_mm9["cage_id_hg19"]
mouse_bed_mm9["cage_id"] = "MOUSE_CAGE_ID__" + mouse_bed_mm9["cage_id_mm9"]
human_bed_mm9["score"] = "HUMAN_SCORE__" + human_bed_hg19["score_tss_hg19"].astype(str)
mouse_bed_mm9["score"] = "MOUSE_SCORE__" + mouse_bed_hg19["score_tss_mm9"].astype(str)
human_bed_mm9.head()
hg19_bed = human_bed_hg19[["chr_tss_hg19", "start_tss_hg19", "end_tss_hg19", "cage_id", "score", "strand_tss_hg19"]]
hg19_bed = hg19_bed.append(mouse_bed_hg19[["chr_tss_hg19", "start_tss_hg19", "end_tss_hg19", "cage_id", "score", "strand_tss_hg19"]])
hg19_bed.drop_duplicates(inplace=True)
print(len(hg19_bed))
hg19_bed.sample(5)
mm9_bed = human_bed_mm9[["chr_tss_mm9", "start_tss_mm9", "end_tss_mm9", "cage_id", "score", "strand_tss_mm9"]]
mm9_bed = mm9_bed.append(mouse_bed_mm9[["chr_tss_mm9", "start_tss_mm9", "end_tss_mm9", "cage_id", "score", "strand_tss_mm9"]])
mm9_bed.drop_duplicates(inplace=True)
print(len(mm9_bed))
mm9_bed.sample(5)
# ## 5. add buffer of +/- 50 bp
hg19_bed["start_tss_hg19"] = hg19_bed["start_tss_hg19"].astype(int) - 49
hg19_bed["end_tss_hg19"] = hg19_bed["end_tss_hg19"].astype(int) + 50
hg19_bed["score"] = 0
hg19_bed.head()
mm9_bed["start_tss_mm9"] = mm9_bed["start_tss_mm9"].astype(int) - 49
mm9_bed["end_tss_mm9"] = mm9_bed["end_tss_mm9"].astype(int) + 50
mm9_bed["score"] = 0
mm9_bed.head()
# ## 6. write files
hg19_bed.to_csv("../../../data/01__design/00__genome_list/hg19_master.50buff.bed", header=False, index=False, sep="\t")
mm9_bed.to_csv("../../../data/01__design/00__genome_list/mm9_master.50buff.bed", header=False, index=False, sep="\t")
| analysis/00__design/00__remap_cage/00__make_files.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
data = pd.read_csv('C:/Users/anura/Google Drive/TF_2_Notebooks_and_Data/heart.csv')
data.head()
data.info()
data.describe().transpose()
import seaborn as sns
import matplotlib.pyplot as plt
sns.countplot(x='output',data = data)
sns.heatmap(data.corr())
sns.pairplot(data=data)
data.corr().transpose()
data.corr()['output'].sort_values()
data.corr()['output'].sort_values().plot(kind='bar')
X = data.drop('output',axis = 1).values
y = data['output'].values
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X,y,test_size=0.25,random_state=101)
X_train
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler()
scaler.fit(X_train)
X_train = scaler.transform(X_train)
X_test = scaler.transform(X_test)
from tensorflow import keras
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense,Activation,Dropout
X_train.shape
# +
model = Sequential()
model.add(Dense(units=13,activation='relu'))
model.add(Dense(units=26,activation='relu'))
model.add(Dense(units=13,activation='relu'))
model.add(Dense(units=13,activation='relu'))
model.add(Dense(units=1,activation='sigmoid'))
# For a binary classification problem
model.compile(loss='binary_crossentropy', optimizer='adam')
# -
model.fit(x=X_train,
y=y_train,
epochs=600,
validation_data=(X_test, y_test), verbose=1
)
model_loss = pd.DataFrame(model.history.history)
model_loss.plot()
from tensorflow.keras.callbacks import EarlyStopping
early_stop = EarlyStopping(monitor='val_loss', mode='min', verbose=1, patience=25)
model.fit(x=X_train,
y=y_train,
epochs=600,
validation_data=(X_test, y_test), verbose=1,
callbacks=[early_stop]
)
model_loss = pd.DataFrame(model.history.history)
model_loss.plot()
predictions = model.predict_classes(X_test)
from sklearn.metrics import classification_report,confusion_matrix
print(classification_report(y_test,predictions))
| Heart Attack Prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This notebook was prepared by [<NAME>](https://github.com/donnemartin). Source and license info is on [GitHub](https://github.com/donnemartin/interactive-coding-challenges).
# # Solution Notebook
# ## Problem: Given two strings, find the longest common subsequence.
#
# * [Constraints](#Constraints)
# * [Test Cases](#Test-Cases)
# * [Algorithm](#Algorithm)
# * [Code](#Code)
# * [Unit Test](#Unit-Test)
# ## Constraints
#
# * Can we assume the inputs are valid?
# * No
# * Can we assume the strings are ASCII?
# * Yes
# * Is this case sensitive?
# * Yes
# * Is a subsequence a non-contiguous block of chars?
# * Yes
# * Do we expect a string as a result?
# * Yes
# * Can we assume this fits memory?
# * Yes
# ## Test Cases
#
# * str0 or str1 is None -> Exception
# * str0 or str1 equals 0 -> ''
# * General case
#
# str0 = 'ABCDEFGHIJ'
# str1 = 'FOOBCDBCDE'
#
# result: 'BCDE'
# ## Algorithm
#
# We'll use bottom up dynamic programming to build a table.
#
# <pre>
#
# The rows (i) represent str0.
# The columns (j) represent str1.
#
# str1
# -------------------------------------------------
# | | | A | B | C | D | E | F | G | H | I | J |
# -------------------------------------------------
# | | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
# | F | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 1 | 1 | 1 |
# | O | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 1 | 1 | 1 |
# s | O | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 1 | 1 | 1 |
# t | B | 0 | 0 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
# r | C | 0 | 0 | 1 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 |
# 0 | D | 0 | 0 | 1 | 2 | 3 | 3 | 3 | 3 | 3 | 3 | 3 |
# | B | 0 | 0 | 1 | 2 | 3 | 3 | 3 | 3 | 3 | 3 | 3 |
# | C | 0 | 0 | 1 | 2 | 3 | 3 | 3 | 3 | 3 | 3 | 3 |
# | D | 0 | 0 | 1 | 2 | 3 | 3 | 3 | 3 | 3 | 3 | 3 |
# | E | 0 | 0 | 1 | 2 | 3 | 4 | 4 | 4 | 4 | 4 | 4 |
# -------------------------------------------------
#
# if str1[j] != str0[i]:
# T[i][j] = max(
# T[i][j - 1],
# T[i - 1][j])
# else:
# T[i][j] = T[i - 1][j - 1] + 1
# </pre>
#
# Complexity:
# * Time: O(m * n), where m is the length of str0 and n is the length of str1
# * Space: O(m * n), where m is the length of str0 and n is the length of str1
# ## Code
class StringCompare(object):
def longest_common_subseq(self, str0, str1):
if str0 is None or str1 is None:
raise TypeError('str input cannot be None')
# Add one to number of rows and cols for the dp table's
# first row of 0's and first col of 0's
num_rows = len(str0) + 1
num_cols = len(str1) + 1
T = [[None] * num_cols for _ in range(num_rows)]
for i in range(num_rows):
for j in range(num_cols):
if i == 0 or j == 0:
T[i][j] = 0
elif str0[j - 1] != str1[i - 1]:
T[i][j] = max(T[i][j - 1],
T[i - 1][j])
else:
T[i][j] = T[i - 1][j - 1] + 1
results = ''
i = num_rows - 1
j = num_cols - 1
# Walk backwards to determine the subsequence
while T[i][j]:
if T[i][j] == T[i][j - 1]:
j -= 1
elif T[i][j] == T[i - 1][j]:
i -= 1
elif T[i][j] == T[i - 1][j - 1] + 1:
results += str1[i - 1]
i -= 1
j -= 1
else:
raise Exception('Error constructing table')
# Walking backwards results in a string in reverse order
return results[::-1]
# ## Unit Test
# +
# %%writefile test_longest_common_subseq.py
from nose.tools import assert_equal, assert_raises
class TestLongestCommonSubseq(object):
def test_longest_common_subseq(self):
str_comp = StringCompare()
assert_raises(TypeError, str_comp.longest_common_subseq, None, None)
assert_equal(str_comp.longest_common_subseq('', ''), '')
str0 = 'ABCDEFGHIJ'
str1 = 'FOOBCDBCDE'
expected = 'BCDE'
assert_equal(str_comp.longest_common_subseq(str0, str1), expected)
print('Success: test_longest_common_subseq')
def main():
test = TestLongestCommonSubseq()
test.test_longest_common_subseq()
if __name__ == '__main__':
main()
# -
# %run -i test_longest_common_subseq.py
| recursion_dynamic/longest_common_subsequence/longest_common_subseq_solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import cv2
import numpy as np
import matplotlib.image as mpimg
import matplotlib.pyplot as plt
# %matplotlib inline
# +
# Load our image - this should be a new frame since last time!
binary_warped = mpimg.imread('warped_example.jpg')
# Polynomial fit values from the previous frame
# Make sure to grab the actual values from the previous step in your project!
left_fit = np.array([ 2.13935315e-04, -3.77507980e-01, 4.76902175e+02])
right_fit = np.array([4.17622148e-04, -4.93848953e-01, 1.11806170e+03])
def fit_poly(img_shape, leftx, lefty, rightx, righty):
### TO-DO: Fit a second order polynomial to each with np.polyfit() ###
left_fit = np.polyfit(lefty, leftx, 2)
right_fit = np.polyfit(righty, rightx, 2)
# Generate x and y values for plotting
ploty = np.linspace(0, img_shape[0]-1, img_shape[0])
### TO-DO: Calc both polynomials using ploty, left_fit and right_fit ###
left_fitx = left_fit[0]*ploty**2 + left_fit[1]*ploty + left_fit[2]
right_fitx = right_fit[0]*ploty**2 + right_fit[1]*ploty + right_fit[2]
return left_fitx, right_fitx, ploty
def search_around_poly(binary_warped):
# HYPERPARAMETER
# Choose the width of the margin around the previous polynomial to search
# The quiz grader expects 100 here, but feel free to tune on your own!
margin = 100
# Grab activated pixels
nonzero = binary_warped.nonzero()
nonzeroy = np.array(nonzero[0])
nonzerox = np.array(nonzero[1])
### TO-DO: Set the area of search based on activated x-values ###
### within the +/- margin of our polynomial function ###
### Hint: consider the window areas for the similarly named variables ###
### in the previous quiz, but change the windows to our new search area ###
left_lane_inds = ((nonzerox > (left_fit[0]*(nonzeroy**2) + left_fit[1]*nonzeroy +
left_fit[2] - margin)) & (nonzerox < (left_fit[0]*(nonzeroy**2) +
left_fit[1]*nonzeroy + left_fit[2] + margin)))
right_lane_inds = ((nonzerox > (right_fit[0]*(nonzeroy**2) + right_fit[1]*nonzeroy +
right_fit[2] - margin)) & (nonzerox < (right_fit[0]*(nonzeroy**2) +
right_fit[1]*nonzeroy + right_fit[2] + margin)))
# Again, extract left and right line pixel positions
leftx = nonzerox[left_lane_inds]
lefty = nonzeroy[left_lane_inds]
rightx = nonzerox[right_lane_inds]
righty = nonzeroy[right_lane_inds]
# Fit new polynomials
left_fitx, right_fitx, ploty = fit_poly(binary_warped.shape, leftx, lefty, rightx, righty)
## Visualization ##
# Create an image to draw on and an image to show the selection window
out_img = np.dstack((binary_warped, binary_warped, binary_warped))*255
window_img = np.zeros_like(out_img)
# Color in left and right line pixels
out_img[nonzeroy[left_lane_inds], nonzerox[left_lane_inds]] = [255, 0, 0]
out_img[nonzeroy[right_lane_inds], nonzerox[right_lane_inds]] = [0, 0, 255]
# Generate a polygon to illustrate the search window area
# And recast the x and y points into usable format for cv2.fillPoly()
left_line_window1 = np.array([np.transpose(np.vstack([left_fitx-margin, ploty]))])
left_line_window2 = np.array([np.flipud(np.transpose(np.vstack([left_fitx+margin,
ploty])))])
left_line_pts = np.hstack((left_line_window1, left_line_window2))
right_line_window1 = np.array([np.transpose(np.vstack([right_fitx-margin, ploty]))])
right_line_window2 = np.array([np.flipud(np.transpose(np.vstack([right_fitx+margin,
ploty])))])
right_line_pts = np.hstack((right_line_window1, right_line_window2))
# Draw the lane onto the warped blank image
cv2.fillPoly(window_img, np.int_([left_line_pts]), (0,255, 0))
cv2.fillPoly(window_img, np.int_([right_line_pts]), (0,255, 0))
result = cv2.addWeighted(out_img, 1, window_img, 0.3, 0)
# Plot the polynomial lines onto the image
plt.plot(left_fitx, ploty, color='yellow')
plt.plot(right_fitx, ploty, color='yellow')
## End visualization steps ##
return result
# Run image through the pipeline
# Note that in your project, you'll also want to feed in the previous fits
result = search_around_poly(binary_warped)
# View your output
plt.imshow(result)
# -
| test_code/11-SearchFromPrior.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .scala
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Scala 2.11 with Spark 2.1
// language: scala
// name: scala-spark21
// ---
// +
// The code was removed by Watson Studio for sharing.
// -
dfData.createOrReplaceTempView("dfData")
dfData.count
dfData.printSchema
val results = spark.sql("SELECT dfData.doc.properties.naturecode FROM dfData")
results.collect
val results2 = spark.sql("SELECT * FROM dfData WHERE dfData.doc.properties.naturecode='LARCEN'")
results2.collect
results2.count
| Build SQL queries.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %reset -f
import os
from cfg import Settings
import torch
import torchvision.datasets as datasets
import torchvision
import torch.nn as nn
import torch.optim as optim
from torch.nn import functional as F
from tensorboardX import SummaryWriter
from sklearn.metrics import roc_auc_score
import matplotlib.pyplot as plt
# -
# # Constants
DATA_DIR = '../data/prepared/'
settings = Settings()
BATCH_SIZE, DEVICE_NAME = settings.get_params()
NUM_WORKERS = 0
#L_R = 1e-3
device = torch.device(DEVICE_NAME)
# # Functions
# +
def pt_loader(path):
return torch.load(path)
def get_dataset(data_dir):
dataset = datasets.DatasetFolder(data_dir, pt_loader, ['pt'])
return dataset
def get_loader(name, data_dir, batch_size, num_workers=0):
if name == 'train' or name == 'val':
dataset = get_dataset(data_dir=os.path.join(data_dir, name))
loader = torch.utils.data.DataLoader(
dataset,
batch_size=batch_size,
num_workers=num_workers,
shuffle=True, pin_memory=True, sampler=None, drop_last=True
)
return loader
else:
raise NotImplemented
# -
# # Net
# +
class SpoofNet(nn.Module):
def __init__(self):
super(SpoofNet, self).__init__()
self.resnet = torchvision.models.resnet18(pretrained=True)
self.fc0 = nn.Linear(1000,500)
self.fc1 = nn.Linear(500,250)
self.fc2 = nn.Linear(250,125)
self.fc3 = nn.Linear(125,50)
self.fc4 = nn.Linear(50,25)
self.fc5 = nn.Linear(25,1)
def fully_conected(self ,x):
x = F.relu(self.fc0(x))
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = F.relu(self.fc3(x))
x = F.relu(self.fc4(x))
x = torch.sigmoid(self.fc5(x))
return x
def forward(self, x):
x = self.resnet(x)
out = self.fully_conected(x)
return out
# Convolutional neural network
class ConvNet1(nn.Module):
def __init__(self):
super(ConvNet1, self).__init__()
self.layer1 = nn.Sequential(
nn.Conv2d(3, 16, kernel_size=5, stride=1, padding=2),
nn.BatchNorm2d(16),
nn.ReLU(),
nn.MaxPool2d(kernel_size=2, stride=2))
self.layer2 = nn.Sequential(
nn.Conv2d(16, 32, kernel_size=5, stride=1, padding=2),
nn.BatchNorm2d(32),
nn.ReLU(),
nn.MaxPool2d(kernel_size=2, stride=2))
self.fc = nn.Linear(56*56*32, 1)
def forward(self, x):
out = self.layer1(x)
out = self.layer2(out)
out = out.reshape(out.size(0), -1)
out = self.fc(out)
return out
class ConvNet2(nn.Module):
def __init__(self):
super(ConvNet2, self).__init__()
self.layer1 = nn.Sequential(
nn.Conv2d(3, 5, kernel_size=3, stride=2, padding=1),
#nn.BatchNorm2d(5),
nn.ReLU())
self.layer2 = nn.Sequential(
nn.Conv2d(5, 7, kernel_size=5, stride=2, padding=1),
#nn.BatchNorm2d(7),
nn.ReLU())
self.layer3 = nn.Sequential(
nn.Conv2d(7, 9, kernel_size=5, stride=2, padding=2),
#nn.BatchNorm2d(9),
nn.ReLU())
self.layer4 = nn.Sequential(
nn.Conv2d(9, 11, kernel_size=5, stride=2, padding=2),
#nn.BatchNorm2d(11),
nn.ReLU())
self.layer5 = nn.Sequential(
nn.Conv2d(11, 13, kernel_size=5, stride=2, padding=2),
#nn.BatchNorm2d(13),
nn.ReLU())
self.fc = nn.Linear(7*7*13, 1)
def forward(self, x):
out = self.layer1(x)
out = self.layer2(out)
out = self.layer3(out)
out = self.layer4(out)
out = self.layer5(out)
out = out.reshape(out.size(0), -1)
out = torch.sigmoid(self.fc(out))
return out
# -
# # Define data loaders
train_loader = get_loader('train', DATA_DIR, BATCH_SIZE, NUM_WORKERS)
val_loader = get_loader('val', DATA_DIR, BATCH_SIZE, NUM_WORKERS)
# # Define model
model = ConvNet2()
#model = torch.load('checkpoints/ResNet50-epoch-11.pt')
criterion = nn.BCELoss()
optimizer = torch.optim.Adam(model.parameters(),lr=1e-3)
torch.cuda.empty_cache()
model = model.to(device)
log_writer = SummaryWriter(log_dir='logs/7-200epoch/')
# # Training
# +
itter = 0
for epoch in range(201):
created = False
for x,y in train_loader:
x , y = x.to(device) , y.to(device)
output = model(x)
loss = criterion(output,y.float())
if not created:
outputs = output
trues = y.float()
created = True
else:
outputs = torch.cat((outputs,output))
trues = torch.cat((trues,y.float()))
optimizer.zero_grad()
loss.backward()
optimizer.step()
log_writer.add_scalar('train/loss', loss.item(), itter)
itter +=1
predictions = (outputs.cpu().data.numpy() > 0.5).astype(int)
train_accuracy = (predictions.T == trues.cpu().data.numpy().astype(int)).mean()
log_writer.add_scalar('train/accuracy', train_accuracy, itter)
with torch.no_grad():
created = False
for x_val,y_val in val_loader:
x_val, y_val = x_val.to(device) , y_val.to(device)
output = model(x_val)
if not created:
outputs = output
trues = y_val.float()
created = True
else:
outputs = torch.cat((outputs,output))
trues = torch.cat((trues,y_val.float()))
val_loss = criterion(outputs,trues)
#val_metrics = roc_auc_score(trues.int().cpu().data.numpy(),outputs.cpu().data.numpy())
predictions = (outputs.cpu().data.numpy() > 0.5).astype(int)
val_accuracy = (predictions.T == trues.cpu().data.numpy().astype(int)).mean()
log_writer.add_scalar('val/loss', val_loss.item(), itter)
#log_writer.add_scalar('val/metrics', val_metrics, itter)
log_writer.add_scalar('val/accuracy', val_accuracy, itter)
print('epoch',epoch,'finished')
if (epoch) % 10 == 0:torch.save(model, os.path.join('checkpoints', 'MyConvNet-epoch-'+str(epoch+1)+'.pt'))
log_writer.close()
# -
# # Try single images
image_name = 'val/spoof/429'
image = torch.load('../data/prepared/' + image_name + '.pt').to(device)
plt.imshow(image.permute(1,2,0))
image = image.reshape(1, image.shape[0], image.shape[1], image.shape[2])
pred = model.forward(image)
print('spoof probability:', pred.cpu().data.numpy()[0,0])
| models/convnet_training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Linear Regression
from si.data import Dataset, summary,StandardScaler
from si.supervised import LinearRegression,LinearRegressionReg
import numpy as np
import os
DIR = os.path.dirname(os.path.realpath('.'))
filename = os.path.join(DIR, 'datasets/lr-example1.data')
dataset = Dataset.from_data(filename, labeled=True)
StandardScaler().fit_transform(dataset,inline=True)
summary(dataset)
import matplotlib.pyplot as plt
# %matplotlib inline
if dataset.X.shape[1]==1:
plt.scatter(dataset.X, dataset.y)
plt.show()
# ## Linear Regression using closed form
lr = LinearRegression()
lr.fit(dataset)
print('Theta = ', lr.theta)
idx = 10
x = dataset.X[idx]
print("x = ",x)
y = dataset.y[idx]
y_pred = lr.predict(x)
print("y_pred = ",y_pred)
print("y_true = ", y)
lr.cost()
if dataset.X.shape[1] == 1:
plt.scatter(dataset.X, dataset.y)
plt.plot(lr.X[:,1], np.dot(lr.X, lr.theta), '-', color='red')
plt.show()
# ## Linear Regression using gradient descent
lr = LinearRegression(gd=True,epochs=50000)
lr.fit(dataset)
print('Theta = ', lr.theta)
plt.plot(list(lr.history.keys()), [ y[1] for y in lr.history.values()], '-', color='red')
plt.title('Cost')
plt.show()
# # Linear Regression with Regularization
lr = LinearRegressionReg()
lr.fit(dataset)
print('Theta = ', lr.theta)
idx = 10
x = dataset.X[idx]
print("x = ", x)
y = dataset.y[idx]
y_pred = lr.predict(x)
print("y_pred = ", y_pred)
print("y_true = ", y)
# # Logistic Regression
from si.supervised import LogisticRegression, LogisticRegressionReg
import pandas as pd
filename = os.path.join(DIR, 'datasets/iris.data')
df = pd.read_csv(filename)
iris = Dataset.from_dataframe(df,ylabel="class")
y = [int(x != 'Iris-setosa') for x in iris.y]
dataset = Dataset(iris.X[:,:2],np.array(y))
summary(dataset)
plt.scatter(dataset.X[:,0], dataset.X[:,1],c=dataset.y)
plt.show()
logreg = LogisticRegression(epochs=20000)
logreg.fit(dataset)
logreg.theta
plt.scatter(dataset.X[:,0], dataset.X[:,1],c=dataset.y)
_x = np.linspace(min(dataset.X[:,0]),max(dataset.X[:,0]),2)
_y = [(-logreg.theta[0]-logreg.theta[1]*x)/logreg.theta[2] for x in _x]
plt.plot(_x, _y, '-', color='red')
plt.show()
plt.plot(list(logreg.history.keys()), [ y[1] for y in logreg.history.values()], '-', color='red')
plt.title('Cost')
plt.show()
ex = np.array([5.5, 2])
print("Pred. example:", logreg.predict(ex))
# # Logistic Regression with L2 regularization
logreg = LogisticRegressionReg()
logreg.fit(dataset)
logreg.theta
plt.scatter(dataset.X[:,0], dataset.X[:,1],c=dataset.y)
_x = np.linspace(min(dataset.X[:,0]),max(dataset.X[:,0]),2)
_y = [(-logreg.theta[0]-logreg.theta[1]*x)/logreg.theta[2] for x in _x]
plt.plot(_x, _y, '-', color='red')
plt.show()
ex = np.array([5.5, 2])
print("Pred. example:", logreg.predict(ex))
# # Cross-validation
from si.util import CrossValidationScore
logreg = LogisticRegression(epochs=1000)
cv = CrossValidationScore(logreg,dataset,cv=5)
cv.run()
cv.toDataframe()
logreg = LogisticRegressionReg(epochs=500, lbd=0.5)
cv = CrossValidationScore(logreg, dataset, cv=4)
cv.run()
cv.toDataframe()
# # Grid Search with Cross-Validation
from si.util import GridSearchCV
parameters ={'epochs':[100,200,400,800,1000],'lbd':[0,0.2,0.4,0.6]}
gs = GridSearchCV(logreg, dataset, parameters, cv=3, split=0.8)
gs.run()
df = gs.toDataframe()
df
| scripts/eval2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Performance optimization overview
#
# The purpose of this tutorial is two-fold
#
# * Illustrate the performance optimizations applied to the code generated by an `Operator`.
# * Describe the options Devito provides to users to steer the optimization process.
#
# As we shall see, most optimizations are automatically applied as they're known to systematically improve performance.
#
# An Operator has several preset **optimization levels**; the fundamental ones are `noop` and `advanced`. With `noop`, no performance optimizations are introduced. With `advanced`, several flop-reducing and data locality optimization passes are applied. Examples of flop-reducing optimization passes are common sub-expressions elimination and factorization, while examples of data locality optimization passes are loop fusion and cache blocking. Optimization levels in Devito are conceptually akin to the `-O2, -O3, ...` flags in classic C/C++/Fortran compilers.
#
# An optimization pass may provide knobs, or **options**, for fine-grained tuning. As explained in the next sections, some of these options are given at compile-time, others at run-time.
#
# **\*\* Remark \*\***
#
# Parallelism -- both shared-memory (e.g., OpenMP) and distributed-memory (MPI) -- is _by default disabled_ and is _not_ controlled via the optimization level. In this tutorial we will also show how to enable OpenMP parallelism (you'll see it's trivial!). Another mini-guide about parallelism in Devito and related aspects is available [here](https://github.com/devitocodes/devito/tree/master/benchmarks/user#a-step-back-configuring-your-machine-for-reliable-benchmarking).
#
# **\*\*\*\***
# ## Outline
#
# * [API](#API)
# * [Default values](#Default-values)
# * [Running example](#Running-example)
# * [OpenMP parallelism](#OpenMP-parallelism)
# * [The `advanced` mode](#The-advanced-mode)
# * [The `advanced-fsg` mode](#The-advanced-fsg-mode)
# ## API
#
# The optimization level may be changed in various ways:
#
# * globally, through the `DEVITO_OPT` environment variable. For example, to disable all optimizations on all `Operator`'s, one could run with
#
# ```
# DEVITO_OPT=noop python ...
# ```
#
# * programmatically, adding the following lines to a program
#
# ```
# from devito import configuration
# configuration['opt'] = 'noop'
# ```
#
# * locally, as an `Operator` argument
#
# ```
# Operator(..., opt='noop')
# ```
#
# Local takes precedence over programmatic, and programmatic takes precedence over global.
#
# The optimization options, instead, may only be changed locally. The syntax to specify an option is
#
# ```
# Operator(..., opt=('advanced', {<optimization options>})
# ```
#
# A concrete example (you can ignore the meaning for now) is
#
# ```
# Operator(..., opt=('advanced', {'blocklevels': 2})
# ```
#
# That is, options are to be specified _together with_ the optimization level (`advanced`).
# ## Default values
#
# By default, all `Operator`s are run with the optimization level set to `advanced`. So this
#
# ```
# Operator(Eq(...))
# ```
#
# is equivalent to
#
# ```
# Operator(Eq(...), opt='advanced')
# ```
#
# and obviously also to
#
# ```
# Operator(Eq(...), opt=('advanced', {}))
# ```
#
# In virtually all scenarios, regardless of application and underlying architecture, this ensures very good performance -- but not necessarily the very best.
# ## Misc
#
# The following functions will be used throughout the notebook for printing generated code.
from examples.performance import unidiff_output, print_kernel
# The following cell is only needed for Continuous Integration. But actually it's also an example of how "programmatic takes precedence over global" (see [API](#API) section).
from devito import configuration
configuration['language'] = 'C'
configuration['platform'] = 'bdw' # Optimize for an Intel Broadwell
configuration['opt-options']['par-collapse-ncores'] = 1 # Maximize use loop collapsing
# ## Running example
#
# Throughout the notebook we will generate `Operator`'s for the following time-marching `Eq`.
# +
from devito import Eq, Grid, Operator, Function, TimeFunction, sin
grid = Grid(shape=(80, 80, 80))
f = Function(name='f', grid=grid)
u = TimeFunction(name='u', grid=grid, space_order=4)
eq = Eq(u.forward, f**2*sin(f)*u.dy.dy)
# -
# Despite its simplicity, this `Eq` is all we need to showcase the key components of the Devito optimization engine.
# ## OpenMP parallelism
#
# There are several ways to enable OpenMP parallelism. The one we use here consists of supplying an _option_ to an `Operator`. The next cell illustrates the difference between two `Operator`'s generated with the `noop` optimization level, but with OpenMP enabled on the latter one.
# +
op0 = Operator(eq, opt=('noop'))
op0_omp = Operator(eq, opt=('noop', {'openmp': True}))
# print(op0)
# print(unidiff_output(str(op0), str(op0_omp))) # Uncomment to print out the diff only
print_kernel(op0_omp)
# -
# The OpenMP-ized `op0_omp` `Operator` includes:
#
# - the header file `"omp.h"`
# - a `#pragma omp parallel num_threads(nthreads)` directive
# - a `#pragma omp for collapse(...) schedule(dynamic,1)` directive
#
# More complex `Operator`'s will have more directives, more types of directives, different iteration scheduling strategies based on heuristics and empirical tuning (e.g., `static` instead of `dynamic`), etc.
#
# The reason for `collapse(1)`, rather than `collapse(3)`, boils down to using `opt=('noop', ...)`; if the default `advanced` mode had been used instead, we would see the latter clause.
#
# We note how the OpenMP pass introduces a new symbol, `nthreads`. This allows users to explicitly control the number of threads with which an `Operator` is run.
#
# ```
# op0_omp.apply(time_M=0) # Picks up `nthreads` from the standard environment variable OMP_NUM_THREADS
# op0_omp.apply(time_M=0, nthreads=2) # Runs with 2 threads per parallel loop
# ```
# A few optimization options are available for this pass (but not on all platforms, see [here](https://github.com/devitocodes/devito/blob/master/examples/performance/README.md)), though in our experience the default values do a fine job:
#
# * `par-collapse-ncores`: use a collapse clause only if the number of available physical cores is greater than this value (default=4).
# * `par-collapse-work`: use a collapse clause only if the trip count of the collapsable loops is statically known to exceed this value (default=100).
# * `par-chunk-nonaffine`: a coefficient to adjust the chunk size in non-affine parallel loops. The larger the coefficient, the smaller the chunk size (default=3).
# * `par-dynamic-work`: use dynamic scheduling if the operation count per iteration exceeds this value. Otherwise, use static scheduling (default=10).
# * `par-nested`: use nested parallelism if the number of hyperthreads per core is greater than this value (default=2).
#
# So, for instance, we could switch to static scheduling by constructing the following `Operator`
op0_b0_omp = Operator(eq, opt=('noop', {'openmp': True, 'par-dynamic-work': 100}))
print_kernel(op0_b0_omp)
# ## The `advanced` mode
#
# The default optimization level in Devito is `advanced`. This mode performs several compilation passes to optimize the `Operator` for computation (number of flops), working set size, and data locality. In the next paragraphs we'll dissect the `advanced` mode to analyze, one by one, some of its key passes.
# ### Loop blocking
#
# The next cell creates a new `Operator` that adds loop blocking to what we had in `op0_omp`.
op1_omp = Operator(eq, opt=('blocking', {'openmp': True}))
# print(op1_omp) # Uncomment to see the *whole* generated code
print_kernel(op1_omp)
# **\*\* Remark \*\***
#
# `'blocking'` is **not** an optimization level -- it rather identifies a specific compilation pass. In other words, the `advanced` mode defines an ordered sequence of passes, and `blocking` is one such pass.
#
# **\*\*\*\***
# The `blocking` pass creates additional loops over blocks. In this simple `Operator` there's just one loop nest, so only a pair of additional loops are created. In more complex `Operator`'s, several loop nests may individually be blocked, whereas others may be left unblocked -- this is decided by the Devito compiler according to certain heuristics. The size of a block is represented by the symbols `x0_blk0_size` and `y0_blk0_size`, which are runtime parameters akin to `nthreads`.
#
# By default, Devito applies 2D blocking and sets the default block shape to 8x8. There are two ways to set a different block shape:
#
# * passing an explicit value. For instance, below we run with a 24x8 block shape
#
# ```
# op1_omp.apply(..., x0_blk0_size=24)
# ```
#
# * letting the autotuner pick up a better block shape for us. There are several autotuning modes. A short summary is available [here](https://github.com/devitocodes/devito/wiki/FAQ#devito_autotuning)
#
# ```
# op1_omp.apply(..., autotune='aggressive')
# ```
#
# Loop blocking also provides two optimization options:
#
# * `blockinner={False, True}` -- to enable 3D (or any nD, n>2) blocking
# * `blocklevels={int}` -- to enable hierarchical blocking, to exploit multiple levels of the cache hierarchy
#
# In the example below, we construct an `Operator` with six-dimensional loop blocking: the first three loops represent outer blocks, whereas the second three loops represent inner blocks within an outer block.
op1_omp_6D = Operator(eq, opt=('blocking', {'blockinner': True, 'blocklevels': 2, 'openmp': True}))
# print(op1_omp_6D) # Uncomment to see the *whole* generated code
print_kernel(op1_omp_6D)
# ### SIMD vectorization
#
# Devito enforces SIMD vectorization through OpenMP pragmas.
op2_omp = Operator(eq, opt=('blocking', 'simd', {'openmp': True}))
# print(op2_omp) # Uncomment to see the generated code
# print(unidiff_output(str(op1_omp), str(op2_omp))) # Uncomment to print out the diff only
# ### Code motion
#
# The `advanced` mode has a code motion pass. In explicit PDE solvers, this is most commonly used to lift expensive time-invariant sub-expressions out of the inner loops. The pass is however quite general in that it is not restricted to the concept of time-invariance -- any sub-expression invariant with respect to a subset of `Dimension`s is a code motion candidate. In our running example, `sin(f)` gets hoisted out of the inner loops since it is determined to be an expensive invariant sub-expression. In other words, the compiler trades the redundant computation of `sin(f)` for additional storage (the `r0[...]` array).
op3_omp = Operator(eq, opt=('lift', {'openmp': True}))
print_kernel(op3_omp)
# ### Basic flop-reducing transformations
#
# Among the simpler flop-reducing transformations applied by the `advanced` mode we find:
#
# * "classic" common sub-expressions elimination (CSE),
# * factorization,
# * optimization of powers
#
# The cell below shows how the computation of `u` changes by incrementally applying these three passes. First of all, we observe how the reciprocal of the symbolic spacing `h_y` gets assigned to a temporary, `r0`, as it appears in several sub-expressions. This is the effect of CSE.
op4_omp = Operator(eq, opt=('cse', {'openmp': True}))
print(unidiff_output(str(op0_omp), str(op4_omp)))
# The factorization pass makes sure `r0` is collected to reduce the number of multiplications.
op5_omp = Operator(eq, opt=('cse', 'factorize', {'openmp': True}))
print(unidiff_output(str(op4_omp), str(op5_omp)))
# Finally, `opt-pows` turns costly `pow` calls into multiplications.
op6_omp = Operator(eq, opt=('cse', 'factorize', 'opt-pows', {'openmp': True}))
print(unidiff_output(str(op5_omp), str(op6_omp)))
# ### Cross-iteration redundancy elimination (CIRE)
#
# This is perhaps the most advanced among the optimization passes in Devito. CIRE [1] searches for redundancies across consecutive loop iterations. These are often induced by a mixture of nested, high-order, partial derivatives. The underlying idea is very simple. Consider:
#
# ```
# r0 = a[i-1] + a[i] + a[i+1]
# ```
#
# at `i=1`, we have
#
# ```
# r0 = a[0] + a[1] + a[2]
# ```
#
# at `i=2`, we have
#
# ```
# r0 = a[1] + a[2] + a[3]
# ```
#
# So the sub-expression `a[1] + a[2]` is computed twice, by two consecutive iterations.
# What makes CIRE complicated is the generalization to arbitrary expressions, the presence of multiple dimensions, the scheduling strategy due to the trade-off between redundant compute and working set, and the co-existance with other optimizations (e.g., blocking, vectorization). All these aspects won't be treated here. What instead we will show is the effect of CIRE in our running example and the optimization options at our disposal to drive the detection and scheduling of the captured redundancies.
#
# In our running example, some cross-iteration redundancies are induced by the nested first-order derivatives along `y`. As we see below, these redundancies are captured and assigned to the two-dimensional temporary `r0`.
#
# Note: the name `cire-sops` means "Apply CIRE to sum-of-product expressions". A sum-of-product is what a finite difference derivative produces.
op7_omp = Operator(eq, opt=('cire-sops', {'openmp': True}))
print_kernel(op7_omp)
# print(unidiff_output(str(op7_omp), str(op0_omp))) # Uncomment to print out the diff only
# We note that since there are no redundancies along `x`, the compiler is smart to figure out that `r0` and `u` can safely be computed within the same `x` loop. This is nice -- not only is the reuse distance decreased, but also a grid-sized temporary avoided.
#
# #### The `min-storage` option
#
# Let's now consider a variant of our running example
eq = Eq(u.forward, f**2*sin(f)*(u.dy.dy + u.dx.dx))
op7_b0_omp = Operator(eq, opt=('cire-sops', {'openmp': True}))
print_kernel(op7_b0_omp)
# As expected, there are now two temporaries, one stemming from `u.dy.dy` and the other from `u.dx.dx`. A key difference with respect to `op7_omp` here is that both are grid-size temporaries. This might seem odd at first -- why should the `u.dy.dy` temporary, that is `r1`, now be a three-dimensional temporary when we know already, from what seen in the previous cell, it could be a two-dimensional temporary? This is merely a compiler heuristic: by adding an extra dimension to `r1`, both temporaries can be scheduled within the same loop nest, thus augmenting data reuse and potentially enabling further cross-expression optimizations. We can disable this heuristic through the `min-storage` option.
op7_b1_omp = Operator(eq, opt=('cire-sops', {'openmp': True, 'min-storage': True}))
print_kernel(op7_b1_omp)
# #### The `cire-mingain` option
#
# So far so good -- we've seen that Devito can capture and schedule cross-iteration redundancies. But what if, for example, we actually do _not_ want certain redundancies to be captured? There are a few reasons we may like that way, for example we're allocating too much extra memory for the tensor temporaries, and we rather prefer to avoid that. For this, the user can tell Devito what the *minimum gain*, or simply `mingain`, which stems from optimizing a CIRE candidate must be in order to trigger the transformation. Such `mingain` is an integer number reflecting the number and type of arithmetic operations in a CIRE candidate. The Devito compiler computes the `gain` of a CIRE candidate, compares it to the `mingain`, and acts as follows:
#
# * if `gain < mingain`, then the CIRE candidate is discarded;
# * if `mingain <= gain <= mingain*3`, then the CIRE candidate is optimized if and only if it doesn't lead to an increase in working set size;
# * if `gain > mingain*3`, then the CIRE candidate is optimized.
#
# The default `mingain` is set to 10, a value that was determined empirically. The coefficient `3` in the relationals above was also determined empirically. The user can change the `mingain`'s default value via the `cire-mingain` optimization option.
#
# To compute the `gain` of a CIRE candidate, Devito applies the following rules:
#
# * Any integer arithmetic operation (e.g. for array indexing) has a cost of 0.
# * A basic arithmetic operation such as `+` and `*` has a cost of 1.
# * A `/` has a cost of 5.
# * A power with non-integer exponent has a cost of 50.
# * A power with non-negative integer exponent `n` has a cost of `n-1` (i.e., the number of `*` it will be converted into).
# * A trascendental function (`sin`, `cos`, etc.) has a cost of 100.
#
# Further, if a CIRE candidate is invariant with respect to one or more dimensions, then the `gain` is proportionately scaled up.
#
# Let's now take a look at `cire-mingain` in action.
eq = Eq(u.forward, f**2*sin(f)*u.dy.dy) # Back to original running example
op8_omp = Operator(eq, opt=('cire-sops', {'openmp': True, 'cire-mingain': 31}))
print_kernel(op8_omp)
# We observe how setting `cire-mingain=31` makes the tensor temporary produced by `op7_omp` disappear. Any `cire-mingain` greater than 31 will have the same effect. Let's see why. The CIRE candidate in question is `u.dy`, which expands to:
#
# ```
# 8.33333333e-2F*u[t0][x + 4][y + 2][z + 4]/h_y - 6.66666667e-1F*u[t0][x + 4][y + 3][z + 4]/h_y + 6.66666667e-1F*u[t0][x + 4][y + 5][z + 4]/h_y - 8.33333333e-2F*u[t0][x + 4][y + 6][z + 4]/h_y
# ```
#
# * Three `+` (or `-`) => 3*1 = 3
# * Four `/` by `h_y` => 4*5 = 20
# * Four two-way `*` with operands `(1/h_y, u)` => 8
#
# So we have a cost of 31. Since the CIRE candidate stems from `.dy`, and since `u`'s space order is 4, the CIRE candidates appears 4 times in total, with shifted indices along `y`. It follows that the gain due to optimizing it away through a tensor temporary would be 31\*(4 - 1) = 93. However, `mingain*3 = 31*3 = 93`, so `mingain < gain <= mingain*3`, and since there would be an increase in working set size (the new tensor temporary to be allocated), then the CIRE candidate is discarded.
#
# As shown below, any `cire-mingain` smaller than 31 will lead to capturing the tensor temporary.
op8_b1_omp = Operator(eq, opt=('cire-sops', {'openmp': True, 'cire-mingain': 30}))
print_kernel(op8_b1_omp)
# #### Dimension-invariant sub-expressions
#
# Cross-iteration redundancies may also be searched across dimension-invariant sub-expressions, typically time-invariants.
eq = Eq(u.forward, f**2*sin(f)*u.dy.dy) # Back to original running example
op11_omp = Operator(eq, opt=('lift', {'openmp': True}))
# print_kernel(op11_omp) # Uncomment to see the generated code
# The `lift` pass triggers CIRE for dimension-invariant sub-expressions. As seen before, this leads to producing one tensor temporary. By setting `cire-mingain` to a larger value, we can avoid a grid-size temporary to be allocated, in exchange for a trascendental function (`sin(...)`) to be computed at each iteration.
op12_omp = Operator(eq, opt=('lift', {'openmp': True, 'cire-mingain': 34}))
print_kernel(op12_omp)
# #### The `cire-maxpar` option
#
# Sometimes it's possible to trade storage for parallelism (i.e., for more parallel dimensions). For this, Devito provides the `cire-maxpar` option which is by default set to:
#
# * False on CPU backends
# * True on GPU backends
#
# Let's see what happens when we switch it on
op13_omp = Operator(eq, opt=('cire-sops', {'openmp': True, 'cire-maxpar': True}))
print_kernel(op13_omp)
# The generated code uses a three-dimensional temporary that gets written and subsequently read in two separate `x-y-z` loop nests. Now, both loops can safely be openmp-collapsed, which is vital when running on GPUs.
# #### Impact of CIRE in the `advanced` mode
#
# The `advanced` mode triggers all of the passes we've seen so far... and in fact, many more! Some of them, however, aren't visible in our running example (e.g., all of the MPI-related optimizations). These will be treated in a future notebook.
#
# Obviously, all of the optimization options (e.g. `cire-mingain`, `blocklevels`) are applicable and composable in any arbitrary way.
op14_omp = Operator(eq, openmp=True)
print(op14_omp)
# op14_b0_omp = Operator(eq, opt=('advanced', {'min-storage': True}))
# A crucial observation here is that CIRE is applied on top of loop blocking -- the `r2` temporary is computed within the same block as `u`, which in turn requires an additional iteration at the block edge along `y` to be performed (the first `y` loop starts at `y0_blk0 - 2`, while the second one at `y0_blk0`). Further, the size of `r2` is now a function of the block shape. Altogether, this implements what in the literature is often referred to as "overlapped tiling" (or "overlapped blocking"): data reuse across consecutive loop nests is obtained by cross-loop blocking, which in turn requires a certain degree of redundant computation at the block edge. Clearly, there's a tension between the block shape and the amount of redundant computation. For example, a small block shape guarantees a small(er) working set, and thus better data reuse, but also requires more redundant computation.
# #### The `cire-rotate` option
#
# So far we've seen two ways to compute the tensor temporaries:
#
# * The temporary dimensions span the whole grid;
# * The temporary dimensions span a block created by the loop blocking pass.
#
# There are a few other ways, and in particular there's a third way supported in Devito, enabled through the `cire-rotate` option:
#
# * The temporary outermost-dimension is a function of the stencil radius; all other temporary dimensions are a function of the loop blocking shape.
#
# Let's jump straight into an example
op15_omp = Operator(eq, opt=('advanced', {'openmp': True, 'cire-rotate': True}))
print_kernel(op15_omp)
# There are two key things to notice here:
#
# * The `r2` temporary is a pointer to a two-dimensional array of size `[2][z_size]`. It's obtained via casting of `pr2[tid]`, which in turn is defined as
print(op15_omp.body.allocs[4])
# * Within the `y` loop there are several iteration variables, some of which (`yr0`, `yr1`, ...) employ modulo increment to cyclically produce the indices 0 and 1.
#
# In essence, with `cire-rotate`, instead of computing an entire slice of `y` values, at each `y` iteration we only keep track of the values that are strictly necessary to evaluate `u` at `y` -- only two values in this case. This results in a working set reduction, at the price of turning one parallel loop (`y`) into a sequential one.
# ## The `advanced-fsg` mode
#
# The alternative `advanced-fsg` optimization level applies the same passes as `advanced`, but in a different order. The key difference is that `-fsg` does not generate overlapped blocking code across CIRE-generated loop nests.
eq = Eq(u.forward, f**2*sin(f)*u.dy.dy) # Back to original running example
op17_omp = Operator(eq, opt=('advanced-fsg', {'openmp': True}))
print(op17_omp)
# The `x` loop here is still shared by the two loop nests, but the `y` one isn't. Analogously, if we consider the alternative `eq` already used in `op7_b0_omp`, we get two completely separate, and therefore individually blocked, loop nests.
eq = Eq(u.forward, f**2*sin(f)*(u.dy.dy + u.dx.dx))
op17_b0 = Operator(eq, opt=('advanced-fsg', {'openmp': True}))
print(op17_b0)
# # References
#
# [1] <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, and <NAME>. 2020. Architecture and Performance of Devito, a System for Automated Stencil Computation. ACM Trans. Math. Softw. 46, 1, Article 6 (April 2020), 28 pages. DOI:https://doi.org/10.1145/3374916
| examples/performance/00_overview.ipynb |