code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %matplotlib inline
#
# # Use source space morphing
#
#
# This example shows how to use source space morphing (as opposed to
# SourceEstimate morphing) to create data that can be compared between
# subjects.
#
# <div class="alert alert-danger"><h4>Warning</h4><p>Source space morphing will likely lead to source spaces that are
# less evenly sampled than source spaces created for individual
# subjects. Use with caution and check effects on localization
# before use.</p></div>
#
#
# +
# Authors: <NAME> <<EMAIL>>
# <NAME> <<EMAIL>>
#
# License: BSD (3-clause)
import os.path as op
import mne
data_path = mne.datasets.sample.data_path()
subjects_dir = op.join(data_path, 'subjects')
fname_trans = op.join(data_path, 'MEG', 'sample',
'sample_audvis_raw-trans.fif')
fname_bem = op.join(subjects_dir, 'sample', 'bem',
'sample-5120-bem-sol.fif')
fname_src_fs = op.join(subjects_dir, 'fsaverage', 'bem',
'fsaverage-ico-5-src.fif')
raw_fname = op.join(data_path, 'MEG', 'sample', 'sample_audvis_raw.fif')
# Get relevant channel information
info = mne.io.read_info(raw_fname)
info = mne.pick_info(info, mne.pick_types(info, meg=True, eeg=False,
exclude=[]))
# Morph fsaverage's source space to sample
src_fs = mne.read_source_spaces(fname_src_fs)
src_morph = mne.morph_source_spaces(src_fs, subject_to='sample',
subjects_dir=subjects_dir)
# Compute the forward with our morphed source space
fwd = mne.make_forward_solution(info, trans=fname_trans,
src=src_morph, bem=fname_bem)
# fwd = mne.convert_forward_solution(fwd, surf_ori=True, force_fixed=True)
mag_map = mne.sensitivity_map(fwd, ch_type='mag')
# Return this SourceEstimate (on sample's surfaces) to fsaverage's surfaces
mag_map_fs = mag_map.to_original_src(src_fs, subjects_dir=subjects_dir)
# Plot the result, which tracks the sulcal-gyral folding
# outliers may occur, we'll place the cutoff at 99 percent.
kwargs = dict(clim=dict(kind='percent', lims=[0, 50, 99]),
# no smoothing, let's see the dipoles on the cortex.
smoothing_steps=1, hemi='rh', views=['lat'])
# Now note that the dipoles on fsaverage are almost equidistant while
# morphing will distribute the dipoles unevenly across the given subject's
# cortical surface to achieve the closest approximation to the average brain.
# Our testing code suggests a correlation of higher than 0.99.
brain_subject = mag_map.plot( # plot forward in subject source space (morphed)
time_label=None, subjects_dir=subjects_dir, **kwargs)
brain_fs = mag_map_fs.plot( # plot forward in original source space (remapped)
time_label=None, subjects_dir=subjects_dir, **kwargs)
| 0.14/_downloads/plot_source_space_morphing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: da-conda
# language: python
# name: auto_da-conda
# ---
# %matplotlib inline
import matplotlib.pyplot as plt
import itertools
import pandas as pd
from os.path import join
import skbio
import numpy as np
import seaborn as sns
from skbio.stats.distance import permanova
# ##Define a function to take the intersect of a distance matrix and map
def filter_dm_and_map(dm, map_df):
ids_to_exclude = set(dm.ids) - set(map_df.index.values)
ids_to_keep = set(dm.ids) - ids_to_exclude
filtered_dm = dm.filter(ids_to_keep)
filtered_map = map_df.loc[ids_to_keep]
return filtered_dm, filtered_map
# ##Load distance matrices
home = '/home/office-microbe-files'
map_fp = join(home, 'master_map_150908.txt')
uw_dm = skbio.DistanceMatrix.read(join(home, 'core_div_out/bdiv_even1000/unweighted_unifrac_dm.txt'))
w_dm = skbio.DistanceMatrix.read(join(home, 'core_div_out/bdiv_even1000/weighted_unifrac_dm.txt'))
# Load the mapping file
# --------------------
# The mapping file will be filtered to only include 16S, row 3 Flagstaff period 2 and 3 samples with no duplicates or replicates. The following is intentionally verbose to improve readability
city_of_interest = 'flagstaff'
# +
sample_md = pd.read_csv(map_fp, sep='\t', index_col=0, dtype=str)
sample_md = sample_md[sample_md['16SITS'] == '16S']
sample_md = sample_md[sample_md['Row'] != '2']
sample_md = sample_md[sample_md['OfficeSample'] == 'yes']
sample_md = sample_md[sample_md['City'] == city_of_interest]
sample_md = sample_md[(sample_md['Period'] == '2') | (sample_md['Period'] == '3')]
# -
replicate_ids = '''F2F.2.Ce.021
F2F.2.Ce.022
F2F.3.Ce.021
F2F.3.Ce.022
F2W.2.Ca.021
F2W.2.Ca.022
F2W.2.Ce.021
F2W.2.Ce.022
F3W.2.Ce.021
F3W.2.Ce.022
F1F.3.Ca.021
F1F.3.Ca.022
F1C.3.Ca.021
F1C.3.Ca.022
F1W.2.Ce.021
F1W.2.Ce.022
F1W.3.Dr.021
F1W.3.Dr.022
F1C.3.Dr.021
F1C.3.Dr.022
F2W.3.Dr.059
F3F.2.Ce.078'''.split('\n')
reps = sample_md[sample_md['Description'].isin(replicate_ids)]
reps = reps.drop(reps.drop_duplicates('Description').index).index
sample_md.drop(reps, inplace=True)
# ###Filter map and matrices
uw_dm, sample_md_uw = filter_dm_and_map(uw_dm, sample_md)
w_dm, sample_md_w = filter_dm_and_map(w_dm, sample_md)
# Retrieve all distances between materials
# ---------------------
ids = {}
for location in ['floor', 'ceiling', 'wall']:
for material in ['carpet', 'drywall', 'ceiling']:
key = '{0}_{1}'.format(location, material)
ids[key] = sample_md_w[(sample_md_w['PlateLocation'] == location) &
(sample_md_w['Material'] == material)].index
# Weighted unifrac
# -----------------
# +
dist_df = pd.DataFrame(data=np.nan, index=sorted(ids.keys()), columns=sorted(ids.keys()))
for key_pairs in itertools.product(ids.keys(), repeat=2):
dists = [w_dm[i] for i in itertools.product(ids[key_pairs[0]], ids[key_pairs[1]])]
dists = np.array(dists)
dists = dists[dists != 0]
dist_df[key_pairs[0]][key_pairs[1]] = np.median(dists)
# +
mask = np.zeros_like(dist_df.values)
mask[np.triu_indices_from(mask)] = True
np.fill_diagonal(mask, 0)
plt.figure(figsize=(12,10))
ticks = ['Ceiling/Carpet', 'Ceiling/Ceiling Tile', 'Ceiling/Drywall',
'Floor/Carpet', 'Floor/Ceiling Tile', 'Floor/Drywall',
'Wall/Carpet', 'Wall/Ceiling Tile', 'Wall/Drywall']
with plt.rc_context(dict(sns.axes_style("dark"),
**sns.plotting_context("notebook", font_scale=2))):
ax = sns.heatmap(dist_df, mask=mask, square=True, cmap="YlGnBu")
ax.set_yticklabels(ticks[::-1])
ax.set_title('Weighted UniFrac')
plt.xticks(np.arange(9) -.2, ticks, rotation=45)
plt.savefig('figure-2-1.svg', dpi=300)
# -
# Unweighted Unifrac
# ------------
ids = {}
for location in ['floor', 'ceiling', 'wall']:
for material in ['carpet', 'drywall', 'ceiling']:
key = '{0}_{1}'.format(location, material)
ids[key] = sample_md_uw[(sample_md_uw['PlateLocation'] == location) &
(sample_md_uw['Material'] == material)].index
dist_df = pd.DataFrame(data=np.nan, index=sorted(ids.keys()), columns=sorted(ids.keys()))
for key_pairs in itertools.product(ids.keys(), repeat=2):
dists = [uw_dm[i] for i in itertools.product(ids[key_pairs[0]], ids[key_pairs[1]])]
dists = np.array(dists)
dists = dists[dists != 0]
dist_df[key_pairs[0]][key_pairs[1]] = np.median(dists)
# +
mask = np.zeros_like(dist_df.values)
mask[np.triu_indices_from(mask)] = True
np.fill_diagonal(mask, 0)
plt.figure(figsize=(12,10))
ticks = ['Ceiling/Carpet', 'Ceiling/Ceiling Tile', 'Ceiling/Drywall',
'Floor/Carpet', 'Floor/Ceiling Tile', 'Floor/Drywall',
'Wall/Carpet', 'Wall/Ceiling Tile', 'Wall/Drywall']
with plt.rc_context(dict(sns.axes_style("dark"),
**sns.plotting_context("notebook", font_scale=2))):
ax = sns.heatmap(dist_df, mask=mask, square=True, cmap="YlGnBu")
ax.set_yticklabels(ticks[::-1])
ax.set_title('Unweighted UniFrac')
plt.xticks(np.arange(9) -.2, ticks, rotation=45)
plt.savefig('figure-2-2.svg', dpi=300)
# -
permanova(w_dm, sample_md, column='PlateLocation', permutations=9999)
permanova(uw_dm, sample_md, column='PlateLocation', permutations=9999)
permanova(w_dm, sample_md, column='Material', permutations=9999)
permanova(uw_dm, sample_md, column='Material', permutations=9999)
| Final/Figure-2/figure-2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import torch
import torch.optim as optim
import torchvision
from torchvision import transforms, datasets, models
import torch.nn as nn
import torch.nn.functional as F
import torch.utils.data as data_utils
from torch.autograd import Variable
import pandas as pd
import numpy as np
import os
from tqdm import tqdm
from skimage import io as skio
from skimage import transform as sktr
from skimage import color as skco
from IPython.display import clear_output
import matplotlib.pyplot as plt
# -
cuda_device=4
batch_size=50
train = pd.read_csv('train_ready.csv')
val = pd.read_csv('valid_ready.csv')
test = pd.read_csv('test_ready.csv')
net = models.resnet50(pretrained=True)
for param in net.parameters():
param.requires_grad = False
num_ftrs = net.fc.in_features
net.fc = torch.nn.Sequential(torch.nn.Linear(num_ftrs, 100),
torch.nn.Dropout(),
torch.nn.LeakyReLU(),
torch.nn.Linear(100, 2))
net = net.cuda(cuda_device)
opt = optim.Adam(net.fc.parameters())
X_train = []
X_val = []
X_test = []
y_train = []
y_val = []
y_test = []
files = set(os.listdir('data'))
def get_image(path):
return skco.gray2rgb(sktr.resize(skio.imread(path), (224, 224)))
# +
for i in tqdm(range(train.shape[0])):
if train.loc[i, 'filename'].split('\\')[-1] in files:
X_train.append(get_image('data/' + train.loc[i, 'filename'].split('\\')[-1]))
y_train.append(train.loc[i, 'label'])
for i in tqdm(range(val.shape[0])):
if val.loc[i, 'filename'].split('\\')[-1] in files:
X_val.append(get_image('data/' + val.loc[i, 'filename'].split('\\')[-1]))
y_val.append(train.loc[i, 'label'])
for i in tqdm(range(test.shape[0])):
if test.loc[i, 'filename'].split('\\')[-1] in files:
X_test.append(get_image('data/' + test.loc[i, 'filename'].split('\\')[-1]))
y_test.append(test.loc[i, 'label'])
# -
X_train = np.array(X_train).transpose([0, 3, 1, 2])
X_val = np.array(X_val).transpose([0, 3, 1, 2])
X_test = np.array(X_test).transpose([0, 3, 1, 2])
y_train = np.array(y_train)
y_val = np.array(y_val)
y_test = np.array(y_test)
# +
train_d = data_utils.TensorDataset(torch.Tensor(X_train), torch.LongTensor(y_train))
train_loader = data_utils.DataLoader(train_d, batch_size=batch_size, shuffle=True)
val_d = data_utils.TensorDataset(torch.Tensor(X_val), torch.LongTensor(y_val))
val_loader = data_utils.DataLoader(val_d, batch_size=batch_size, shuffle=True)
test_d = data_utils.TensorDataset(torch.Tensor(X_test), torch.LongTensor(y_test))
test_loader = data_utils.DataLoader(test_d, batch_size=batch_size, shuffle=True)
# -
n_epoch = 10
val_scores = []
train_scores = []
for epoch in tqdm(range(n_epoch)):
val_loss = 0
train_loss = 0
i = 0
net.train()
for (x_batch, y_batch) in train_loader:
x_batch = Variable(x_batch).cuda(cuda_device)
y_batch = Variable(y_batch).cuda(cuda_device)
loss = F.cross_entropy(net.forward(x_batch), y_batch).cuda(cuda_device)
opt.zero_grad()
loss.backward()
opt.step()
train_loss += loss.cpu().data.numpy()
i += 1
train_scores.append(train_loss / i)
i = 0
net.eval()
for (x_batch, y_batch) in val_loader:
x_batch = Variable(x_batch).cuda(cuda_device)
y_batch = Variable(y_batch).cuda(cuda_device)
loss = F.cross_entropy(net.forward(x_batch), y_batch).cuda(cuda_device)
val_loss += loss.cpu().data.numpy()
i += 1
val_scores.append(val_loss / i)
clear_output()
plt.plot(val_scores, label='validate')
plt.plot(train_scores, label='train')
plt.legend()
plt.xlabel='epoch'
plt.ylabel='entropy score'
plt.show()
loss = 0
for (x_batch, y_batch) in test_loader:
x_batch = Variable(x_batch).cuda(cuda_device)
y_batch = Variable(y_batch).cuda(cuda_device)
pred = net.forward(x_batch).max(dim=1)[1]
loss += (y_batch == pred).sum().cpu().data.numpy()
print(loss / X_test.shape[0])
| 6_sem/image_analysis/lab8/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="a8xO3SKOmBgS"
# # Notebook [1]: First steps with cdQA
# + [markdown] colab_type="text" id="v874L1S8mBgV"
# This notebook shows how to use the `cdQA` pipeline to perform question answering on a custom dataset.
# + [markdown] colab_type="text" id="93FkcDXTmBgW"
# ***Note:*** *If you are using colab, you will need to install `cdQA` by executing `!pip install cdqa` in a cell.*
# + colab={"base_uri": "https://localhost:8080/", "height": 666} colab_type="code" id="iXhvKJckmQ0u" outputId="fbec9d01-043e-4dd2-e5c6-4560afcb148d"
# !pip install cdqa
# + colab={"base_uri": "https://localhost:8080/", "height": 117} colab_type="code" id="HY_Lt_u7mBgX" outputId="0d337e2d-595b-4831-bdfa-ee3879982675"
import os
import pandas as pd
from ast import literal_eval
from cdqa.utils.filters import filter_paragraphs
from cdqa.pipeline import QAPipeline
# + [markdown] colab_type="text" id="uZxVq9ZSmBgb"
# ### Download pre-trained reader model and example dataset
# + colab={"base_uri": "https://localhost:8080/", "height": 85} colab_type="code" id="PfP-VDQlmBgc" outputId="c87c6935-0b2a-4be3-c5bc-9d0fa5cefd05"
from cdqa.utils.download import download_model, download_bnpp_data
download_bnpp_data(dir='./data/bnpp_newsroom_v1.1/')
download_model(model='bert-squad_1.1', dir='./models')
# + [markdown] colab_type="text" id="HPx_reWKmBge"
# ### Visualize the dataset
# + colab={"base_uri": "https://localhost:8080/", "height": 289} colab_type="code" id="h5HtFQijmBgf" outputId="0939cb6b-f06f-461e-d4aa-dbb656479283"
df = pd.read_csv('./data/bnpp_newsroom_v1.1/bnpp_newsroom-v1.1.csv', converters={'paragraphs': literal_eval})
df = filter_paragraphs(df)
df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 139} colab_type="code" id="O5ZeuM9jsz4p" outputId="3fbacb26-6d8d-4020-8800-1dfb4a0a1a39"
df.loc[0].paragraphs
# + colab={"base_uri": "https://localhost:8080/", "height": 54} colab_type="code" id="qHZTSHHRsjeg" outputId="b9a2fc0d-ab99-441b-8a9f-a9b9def43e8c"
''.join(df.loc[0].paragraphs)
# + [markdown] colab_type="text" id="Dfe9D4GcmBgi"
# ### Instantiate the cdQA pipeline from a pre-trained reader model
# + colab={"base_uri": "https://localhost:8080/", "height": 323} colab_type="code" id="ohXSOwiemBgi" outputId="cc35fe37-14c6-4d4b-f008-e5eb7a0d7b18"
cdqa_pipeline = QAPipeline(reader='./models/bert_qa.joblib')
cdqa_pipeline.fit_retriever(df=df)
# + [markdown] colab_type="text" id="36aw9kYEmBgk"
# ### Execute a query
# + colab={} colab_type="code" id="jcBCtCpYmBgl"
query = 'Since when does the Excellence Program of BNP Paribas exist?'
prediction = cdqa_pipeline.predict(query)
# + [markdown] colab_type="text" id="Gy_6eNDymBgn"
# ### Explore predictions
# + colab={"base_uri": "https://localhost:8080/", "height": 105} colab_type="code" id="FdvFtKDqmBgo" outputId="6f43283d-10fa-40a8-df10-b11f8804be59"
print('query: {}'.format(query))
print('answer: {}'.format(prediction[0]))
print('title: {}'.format(prediction[1]))
print('paragraph: {}'.format(prediction[2]))
# + colab={} colab_type="code" id="bHP9QfYbs_j8"
query2 = "Who's BNP Paribas' CEO?"
prediction2 = cdqa_pipeline.predict(query2)
# + colab={"base_uri": "https://localhost:8080/", "height": 105} colab_type="code" id="ARsfjoNktJQW" outputId="5ef147ef-d2c7-4119-e5b0-063dbd518666"
print('query: {}'.format(query2))
print('answer: {}'.format(prediction2[0]))
print('title: {}'.format(prediction2[1]))
print('paragraph: {}'.format(prediction2[2]))
# + colab={} colab_type="code" id="eGhKKM7BtS0K"
| examples/tutorial-first-steps-cdqa-rh.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# HIDDEN
from datascience import *
from prob140 import *
import numpy as np
import matplotlib.pyplot as plt
plt.style.use('fivethirtyeight')
# %matplotlib inline
import math
from scipy import stats
from scipy import misc
# ### Occupation Times Within a Block ###
# Suppose the chain starts at $i$. Then it is intuitively plausible that once it returns to $i$, it "starts all over again" as though it were starting at $i$ at time 0. This hand-wavy statement can be turned into mathematics, but in this course we'll just accept it without doing the math.
#
# Start the chain at $i$, that is, with the initial distribution $P(X_0 = i) = 1$. Define an *$i$-block* to be the chain till the step before it returns to $i$:
#
# - Under the initial condition that $X_0 = i$, an $i$-block is $X_0, X_1, X_2, \ldots, X_{W_i - 1}$.
#
# Here $W_i$ is the hitting time of $i$ as defined in the previous section:
#
# $$
# W_i = \inf \{n \ge 1: X_n = i \}
# $$
#
# Notice that in an $i$-block, the chain is only at $i$ once, at time 0.
#
# Notice also that the length of the $i$-block is $W_i$. You can see this by counting indices in the definition of the $i$-block. But the $i$-block doesn't end at time $W_i$; it ends one step earlier. Its length is $W_i$ because it includes time 0, whereas $W_i$ starts counting time at 1. Informally, the $i$-block includes the $i$ at the start of the block but not the $i$ directly following the block, whereas $W_i$ doesn't include time 0 but does include the moment when $i$ appears at the end of the block.
#
# This will become clear when you look at the graph below, in which $i = 3$. The blue dots are the $i$-block. There are 8 dots in it, corresponding to times 0 through 7. The red dot immediately after the $i$-block shows the return to $i$. It's at time 8.
# HIDDEN
tosses = make_array(1, 1, 1, -1, -1, 1, -1, -1)
fortune = np.cumsum(np.append(3, tosses))
plt.scatter(np.arange(9), fortune, color='darkblue')
plt.scatter(8, 3, color='r')
plt.ylim(-0.5, 7.5)
plt.xlim(-0.2, 8.5)
plt.xlabel('$n$')
plt.ylabel('$X_n$', rotation=0)
plt.xticks(np.arange(0, 9, 1))
plt.yticks(np.arange(0, 8, 1))
plt.title('$i$-Block with $i=3$ and $W_i$ = 8');
# We have been careful not to include $X_{W_i}$ in the $i$-block. At time $W_i$ the chain returns to $i$, and we will think of that as the start of the next $i$-block. Because the chain "starts over" at $i$, we can imagine the entire chain as identically distributed $i$-blocks strung together one after another. Therefore there are close relations between long run properties of the chain and short run properties on an $i$-block. Let's look at one of these relations.
#
# Let $X_0 = i$ and let $N(j)$ be the number of times the chain is at $j$ in the $i$-block. We will call $N(j)$ the *number of visits to $j$ in an $i$-block*.
#
# As we have already observed, $N(i) = 1$ with probability 1. By partitioning the $i$-block according to visits to the different states, the length of the $i$-block can be written as a sum:
#
# $$
# \text{length of } i\text{-block} = W_i = 1 + \sum_{j \ne i} N(j)
# $$
#
# You can check this in the graph above where $i=3$. The graph shows a path for which $N(3) = 1$, $N(4) = 3 = N(5)$, $N(6) = 1$, and $N(j) = 0$ for all other states $j$. The sum of all these occupation times is 8, which is also the value of $W_i$.
#
# So
# $$
# \frac{1}{\pi(i)} = E(W_i \mid X_0 = i) = 1 + \sum_{j \ne i} E(N(j) \mid X_0 = i)
# $$
#
# ### Expected Occupation Times in an $i$-Block ###
# Fix a state $j$. The expected number of visits to $j$ in an $i$-block is
#
# $$
# E(N(j) \mid X_0 = i) = \frac{\pi(j)}{\pi(i)}
# $$
#
# A formal proof requires a little care; we won't go through it. Rather, we will show why the formula is consistent with our previous calculations and with intuition.
#
# - The formula is correct for $j = i$, because $N(i) = 1$ with probability 1.
# - The terms add up to $\frac{1}{\pi(i)}$ as we have shown they must.
#
# $$
# 1 + \sum_{j \ne i} \frac{\pi(j)}{\pi(i)}
# ~ = ~ \frac{\pi(i)}{\pi(i)} + \sum_{j \ne i} \frac{\pi(j)}{\pi(i)}
# ~ = ~ \frac{1}{\pi(i)} \big{(} \sum_{\text{all }j} \pi(j) \big{)}
# ~ = ~ \frac{1}{\pi(i)}
# $$
#
# because $\pi$ is a probability distribution and hence sums to 1.
#
# - Think of the chain as a string of $i$-blocks. You know that overall, the expected proportion of times that the chain spends at $j$ is $\pi(j)$. Since the $i$-blocks are identically distributed copies of each other, it makes sense that the chain is expected to spend the same proportion $\pi(j)$ of time in $j$ in each $i$-block. Since the length of an $i$-block is expected to be $1/\pi(i)$, the expected number of times the chain is at $j$ in an $i$-block is $\pi(j)/\pi(i)$.
# ### Example: Ehrenfest Model ####
# In the example of the Ehrenfest model we worked with in the previous section, the stationary distribution is binomial $(N, 1/2)$. Suppose the chain starts at $X_n = 0$, that is, with Container 1 empty. Then for every $k$, the expected number of times Container 1 has $k$ particles before it is once again empty is
#
# $$
# \frac{\binom{N}{k}(1/2)^N}{\binom{N}{0}(1/2)^N} = \binom{N}{k}
# $$
# ### Example: Uniform Stationary Distribution ###
# Consider any chain that has a stationary distribution that is uniform. You have seen in exercises that chains with doubly stochastic transition matrices fall into this category. Suppose such a chain starts at state $i$. Then for any other state $j$, the expected number of times the chain visits $j$ before returning to $i$ is 1.
| miscellaneous_notebooks/Hitting_and_Occupation_Times/Occupation_Times_Within_a_Block.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: analysis
# language: python
# name: analysis
# ---
# # Multivariate Outliers Detection
#
# https://scikit-learn.org/stable/modules/outlier_detection.html
# https://scikit-learn.org/stable/modules/generated/sklearn.neighbors.LocalOutlierFactor.html
# + active=""
# !pip install scikit-learn==0.22
# -
# %matplotlib inline
import warnings
warnings.filterwarnings('ignore')
import pandas as pd
import numpy as np
# ## load data
from sklearn.datasets import load_iris
iris = load_iris()
X = iris.data
target = iris.target
names = iris.target_names
data = pd.DataFrame(X, columns=iris.feature_names)
data['species'] = iris.target
data['species'] = data['species'].replace(to_replace= [0, 1, 2], value = ['setosa', 'versicolor', 'virginica'])
data.shape
# # Multivariate OUTLIERS detection for a df
# columns selection
col_x1 = ['petal length (cm)', 'petal width (cm)']
col_x2 = ['sepal length (cm)', 'sepal width (cm)']
# plot
import matplotlib.pyplot as plt
fig, (ax1, ax2) = plt.subplots(ncols = 2, figsize = (12, 6))
data[col_x1].plot(kind = 'scatter', x = col_x1[0], y = col_x1[1], ax = ax1)
data[col_x2].plot(kind = 'scatter', x = col_x2[0], y = col_x2[1], ax = ax2)
plt.show()
# https://scikit-learn.org/stable/modules/generated/sklearn.neighbors.LocalOutlierFactor.html
# https://stackoverflow.com/questions/34643548/how-to-use-mahalanobis-distance-in-sklearn-distancemetrics
# +
## detect multivariate outliers using Local Outlier Factor
def detect_outliers_LOF(X:np.array, n_neighbors:int = 25, n_jobs:int = 2, verbose:bool = False)->np.array:
"""
Detect multivariate outliers using Local Outlier Factor.
X -- array of values to be analyzed.
n_neighbors -- number of neighbors to be used by the algorithm (default, 25).
n_jobs -- number of jobs to be used (default, 2).
verbose -- display extra information (default, False).
return -- array of tagged samples (-1: it is outlier, 1: it is not).
"""
from sklearn.neighbors import LocalOutlierFactor
# initialize
clf = LocalOutlierFactor(n_neighbors=n_neighbors,
algorithm = "auto",
metric = 'minkowski',
p = 2,
contamination="auto",
novelty=False,
n_jobs = n_jobs
)
# estimate
try:
y_pred = clf.fit(X).predict(X)
except:
y_pred = clf.fit_predict(X)
# display
if verbose:
print(f'There are {y_pred[y_pred == -1].shape[0]} outliers from {y_pred.shape[0]}.')
# return
return y_pred
# -
## multivariante outliers detection for a given column of a df
def multivariate_outliers_detection(data:pd.DataFrame,
col_names: list,
is_remove:bool = True,
methodology:'function' = detect_outliers_LOF,
verbose:bool = False)->pd.DataFrame:
"""
Multivariante outliers detection for a given column of a df.
data -- dataframe to be analyzed.
col_names -- columns to be used.
is_remove -- if removing outliers or just detect (default, True).
methodology -- function of method to be used to remove / detect outliers (default, detect_outliers_LOF()).
verbose -- display extra information (default, False).
return -- df of values without outliers or a mask with detected outliers.
"""
# validate
for col in col_names:
assert col in data.columns.tolist()
# initialize if just detection
if not is_remove:
df_mask = pd.DataFrame(np.zeros(data[col_names].shape, dtype=bool), columns = col_names)
# outliers detection
y_pred = methodology(data[col_names].values, n_neighbors = 25, verbose = verbose)
# number of outliers
num_outliers = len(y_pred[y_pred == -1])
# validate
if num_outliers > 0:
# if removing
if is_remove:
# add labels
data['label'] = y_pred
# filter
ni = len(data)
data = data[data.label == 1]
nf = len(data)
# display
if verbose:
print(f'It was removed {ni - nf} records.')
# remove unnecessary column
data.drop('label', axis = 1, inplace = True)
# if just detection
else:
# loop of columns
for col in col_names:
# inpute True if it is an outlier
df_mask[col] = [True if v == -1 else False for v in y_pred]
# return
if is_remove:
return data
else:
return df_mask
col_x = col_x1 + col_x2
dfr = multivariate_outliers_detection(data, col_x, is_remove = True, verbose = True)
import matplotlib.pyplot as plt
fig, ax = plt.subplots()
col_x = col_x1
data.plot(kind='scatter', x = col_x[0], y = col_x[1], color = 'red', ax = ax)
dfr.plot(kind='scatter', x = col_x[0], y = col_x[1], color = 'blue', ax = ax)
plt.show()
# +
import matplotlib
import matplotlib.pyplot as plt
from sklearn import svm
from sklearn.datasets import make_moons, make_blobs
matplotlib.rcParams["contour.negative_linestyle"] = "solid"
# Example settings
n_samples = 300
outliers_fraction = 0.15
n_outliers = int(outliers_fraction * n_samples)
n_inliers = n_samples - n_outliers
# Define datasets
blobs_params = dict(random_state=0, n_samples=n_inliers, n_features=2)
datasets = [
make_blobs(centers=[[0, 0], [0, 0]], cluster_std=0.5, **blobs_params)[0],
make_blobs(centers=[[2, 2], [-2, -2]], cluster_std=[0.5, 0.5], **blobs_params)[0],
make_blobs(centers=[[2, 2], [-2, -2]], cluster_std=[1.5, 0.3], **blobs_params)[0],
4.0
* (
make_moons(n_samples=n_samples, noise=0.05, random_state=0)[0]
- np.array([0.5, 0.25])
),
14.0 * (np.random.RandomState(42).rand(n_samples, 2) - 0.5),
]
rng = np.random.RandomState(42)
for i_dataset, X in enumerate(datasets[:]):
# Add outliers
X = np.concatenate([X, rng.uniform(low=-6, high=6, size=(n_outliers, 2))], axis=0)
fig, ax = plt.subplots()
y_pred = detect_outliers_LOF(X)
colors = np.array(["#377eb8", "#ff7f00"])
plt.scatter(X[:, 0], X[:, 1], s=10, color=colors[(y_pred + 1) // 2])
plt.xlim(-7, 7)
plt.ylim(-7, 7)
plt.xticks(())
plt.yticks(())
plot_num += 1
plt.show()
# -
| notebooks/analysis/analysis_anomalies/notebook-multivariate_outliers_detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Demo notebook for accessing MTBS data on Azure
#
# This notebook provides an example of accessing Monitoring Trends in Burn Severity (MTBS) Mosiacs for CONUS and Alaska from blob storage on Azure. The data is stored in annual cloud optimized GeoTIFF files.
#
# MTBS data are stored in the West Europe Azure region, so this notebook will run most efficiently on Azure compute located in West Europe. We recommend that substantial computation depending on MTBS data also be situated in West Europe. You don't want to download hundreds of terabytes to your laptop! If you are using MTBS data for environmental science applications, consider applying for an [AI for Earth grant](http://aka.ms/ai4egrants) to support your compute requirements.
# ### Imports and constants
# +
import xarray as xr
from adlfs import AzureBlobFileSystem
# Year to investigate and plot
year = 2018
# Storage resources
storage_account_name = 'cpdataeuwest'
region = 'conus' # 'conus' or 'ak'
folder = f'cpdata/raw/mtbs/{region}/30m/severity'
# -
# ### List the data files
#
# The MTBS data on Azure is available for either the CONUS or Alaska region.
#
# We can use `adlfs` to list the files in either region:
fs = AzureBlobFileSystem(account_name=storage_account_name)
mosaic_files = fs.glob(folder + '/*.tif')
print('Found {} mosaic files:'.format(len(mosaic_files)))
for k in range(0,10):
print(mosaic_files[k])
print('...')
# ### Open one data file with xarray
url = 'https://' + storage_account_name + '.blob.core.windows.net/' + \
folder + '/' + str(year) + '.tif'
print('Reading data from {}'.format(url))
da = xr.open_rasterio(url,chunks={'x': 2560, 'y': 2560})
da
# ### Plot one fire event
#
# We can select a subset of the data to make a plot over a single fire event.
da.sel(band=1, y=slice(2.16e6, 2.08e6),
x=slice(None, -2.23e6)).plot.contourf(cmap='Set1_r')
| data/mtbs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: torch
# language: python
# name: torch
# ---
# +
import torch
import numpy as np
import matplotlib.pyplot as pl
from ipywidgets import interact, interactive, fixed, interact_manual
import ipywidgets as widgets
# -
# Minimal class
# +
class WeibullCDF_IOFunc:
'''Implements the Weibull CDF function
f(I) = 1-exp(- ((I-I0)/scale)^k )
'''
def __init__(self, I0=0., scale=40., k=10., requires_grad=False, mmax=1., constrained_at_Iref=False, Iref=-20):
'''
Args:
theta: localization parameter (max intensity associated with 0% masking)
scale: scale parameter (63% masking intensity reached at I0+scale)
k: shape parameter
mmax: maximum masking
constrained_at_Iref: if True, constrains the function to equal 1 at Iref. (in this case, mmax is superfluous)
Iref: Iref in dB in the case of 'constrained_at_Iref
'''
self.I0=I0
self.scale=scale
self.k=k
self.constrained_at_Iref=constrained_at_Iref
self._Iref=Iref
self.mmax=mmax
def __call__(self, I):
Delta_I=np.maximum((I-self.I0), 0.)
if self.constrained_at_Iref:
Delta_I_ref=np.maximum((self._Iref-self.I0), 0.)
return (1-np.exp( -(Delta_I/self.scale)**self.k))/(1-np.exp( -(Delta_I_ref/self.scale)**self.k))
else:
return self.mmax*(1-np.exp( -(Delta_I/self.scale)**self.k))
class LogLinearSuppression:
def __init__(self, a, I0_supp, requires_grad=False):
'''
Args:
a: suppression amount dB/dB
'''
self.a=a
self.I0_supp=I0_supp
def __call__(self, I_supp):
return self.a*(I_supp-self.I0_supp)
# -
#Iref = float(100 +10 - 32)
fs=48828
#Iref = 10*np.log10( 10**(Iref/10)/(fs/2) ) #spectral density
#Iref-=20
#Iref=21
Iref=12
#NB: remove 'manual' for continuous update
@interact(I0=widgets.FloatSlider(value=-20, min=-40, max=0, step=1),
scale=widgets.FloatSlider(value=30, min=1, max=100, step=1),
k=widgets.FloatSlider(value=10, min=1, max=20, step=0.5))
def plotwbcdf(I0, scale, k):
wb_cdf=WeibullCDF_IOFunc(I0=I0,
scale=scale,
k=k,
requires_grad=False,
constrained_at_Iref=True,
Iref=Iref)
wb_cdf2=WeibullCDF_IOFunc(I0=-31., #-15
scale=37.,
k=3.5,
mmax=1.,
requires_grad=False,
constrained_at_Iref=True,
Iref=Iref)
I=np.linspace(-30, 30)
pl.figure()
pl.plot(I, wb_cdf(I))
pl.plot(I, wb_cdf2(I))
pl.xlim([-30, 30])
pl.title('Masking IO Function')
pl.xlabel('Power spectral density (dB)')
pl.show()
# Weibull CDF simulation 'suppression effect'
supp=LogLinearSuppression(0.6, -30)
#NB: remove 'manual' for continuous update
@interact(I0=widgets.FloatSlider(value=-20, min=-40, max=0, step=1),
scale=widgets.FloatSlider(value=30, min=1, max=100, step=1),
k=widgets.FloatSlider(value=10, min=1, max=20, step=0.5))
def plotwbcdf(I0, scale, k):
wb_cdf=WeibullCDF_IOFunc(I0=I0,
scale=scale,
k=k,
requires_grad=False,
constrained_at_Iref=True,
Iref=Iref)
wb_cdf2=WeibullCDF_IOFunc(I0=-31., #-15
scale=37.,
k=3.5,
mmax=1.,
requires_grad=False,
constrained_at_Iref=True,
Iref=Iref)
I=np.linspace(-30, 30)
pl.figure()
pl.plot(I, wb_cdf(I), color='C0')
#pl.plot(I, wb_cdf(I - supp(I) ) , color='C0', linestyle='--')
pl.plot(I, wb_cdf(I - supp(I) )/wb_cdf(Iref - supp(Iref)) , color='C0', linestyle='--')
pl.plot(I, wb_cdf2(I), color='C1')
#pl.plot(I, wb_cdf2(I - supp(I) ), color='C1', linestyle='--')
pl.xlim([-30, 30])
pl.title('Masking IO Function')
pl.xlabel('Power spectral density (dB)')
pl.show()
# Sigmoid
#NB: remove 'manual' for continuous update
@interact(mu=widgets.FloatSlider(value=0, min=-20, max=20, step=1),
a=widgets.FloatSlider(value=0.25, min=0.01, max=2, step=0.1))
def plotsigm(mu, a):
def sigm(I):
return 1/(1+np.exp(- a*(I-mu)))
I=np.linspace(-30, 20)
pl.figure()
pl.plot(I, sigm(I))
pl.xlim([-20, 20])
pl.title('Masking IO Function')
pl.xlabel('Power spectral density (dB)')
pl.show()
Delta_I_ref
| utils/test weibull.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
import kwat
# ## print_stack_state
# +
def fu1(ar, ke="Local"):
lo1 = "Local"
kwat.python.print_stack_state()
fu1("Local")
# +
def fu2(ar):
lo2 = "Not local"
fu1(ar)
fu2("Local")
# -
# ## cast
import numpy as np
for an in [None, "None", False, "False", True, "True", 0, 0.0, "0", np.nan]:
print(kwat.python.cast(an))
| nb/python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import itertools
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.ticker import NullFormatter
import pandas as pd
import numpy as np
import matplotlib.ticker as ticker
from sklearn import preprocessing
# %matplotlib inline
url = "https://en.wikipedia.org/wiki/List_of_postal_codes_of_Canada:_M"
df_list = pd.read_html(url)
len(df_list), type(df_list)
df_list
df = df_list[0]
df.head
df.head(10)
type(df)
df = df[df.Borough != 'Not assigned']
df.head(10)
len(df)
df.reset_index(drop=True)
df.count()
df = df.replace(['Not assigned'], np.nan)
df = df.reset_index(drop=True)
df
# Cria um index dos valores que são Nan
nan_index = df[df.isnull()].index
# Para todos os Nans
for i in range(df.isnull().sum()):
# Extrai os valores da outra coluna que você quer procurar
dado_nan = df[['coluna']][df.isnull()].iloc[i].values()
# Substitui com as médias dos valores dentro da faixa desejada
df['novaColuna'][nan_index[i]] = df[abs(df.coluna - dado_nan[0]) < 2.5].mean()
for i in df.index:
dado_NA = df["Borough"].iloc[i].values()
df.at[i,"Borough"]
df['Neighbourhood'] = df['Neighbourhood'].str.replace('Not assigned', dado_NA))
# Cria um index dos valores que são Nan
nan_index = df[df.isnull()].index
# Para todos os Nans
for i in range(df.isnull().sum()):
# Extrai os valores da outra coluna que você quer procurar
dado_nan = df[['coluna']][df.isnull()].iloc[i].values()
# Substitui com as médias dos valores dentro da faixa desejada
df['novaColuna'][nan_index[i]] = df[abs(df.coluna - dado_nan[0]) < 2.5].mean()
def brg()
df.dtype
| Diversos/Capstone IBM Data Science.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Chi-Square
#
# In this Statistics Appendix Lecture, we'll go over the Chi-Square Distribution and the Chi-Square Test.
#
# Let's start by introducing the general idea of observed and theoretical frequencies, then later we'll approach the idea of the Chi-Sqaure Distribution and its definition. After that we'll do a qcuik example with Scipy on using the Chi-Square Test.
#
#
# Supppose that you tossed a coin 100 times. Theoretically you would expect 50 tails and 50 heads, however it is pretty unlikely you get that result exactly. Then a question arises... how far off from you expected/theoretical frequency would you have to be in order to conclude that the observed result is statistically significant and is not just due to random variations.
#
# We can begin to think about this question by defining an example set of possible events. We'll call them Events 1 through *k*. Each of these events has an expected (theoretical) frequency and an observed frequency. We can display this as a table:
# <table>
# <tr>
# <th>Event</th>
# <th>Event 1</th>
# <th>Event 2</th>
# <th>Event 3</th>
# <th>...</th>
# <th>Event k</th>
# </tr>
# <tr>
# <td>Observed Frequency</td>
# <td>$$o_1$$</td>
# <td>$$o_2$$</td>
# <td>$$o_3$$</td>
# <td>...</td>
# <td>$$o_k$$</td>
# </tr>
# <tr>
# <td>Expected Frequency</td>
# <td>$$e_1$$</td>
# <td>$$e_2$$</td>
# <td>$$e_3$$</td>
# <td>...</td>
# <td>$$e_k$$</td>
# </tr>
# </table>
# Since we wanted to know whether observed frequencies differ significantly from the expected frequencies we'll have to define a term for a measure of discrepency.
#
# We'll define this measure as Chi-Square, which will be the sum of the squared difference between the observed and expected frequency divided by the expected frequency for all events. To show this more clearly, this is mathematically written as:
# $$ \chi ^2 = \frac{(o_1 - e_1)^2}{e_1}+\frac{(o_2 - e_2)^2}{e_2}+...+\frac{(o_k - e_k)^2}{e_k} $$
# Which is the same as:
# $$\chi ^2 = \sum^{k}_{j=1} \frac{(o_j - e_j)^2}{e_j} $$
# If the total frequency is N
# $$ \sum o_j = \sum e_j = N $$
# Then we could rewrite the Chi-Square Formula to be:
# $$ \chi ^2 = \sum \frac{o_j ^2}{e_j ^2} - N$$
# We can now see that if the Chi Square value is equal to zero, then the observed and theoretical frequencies agree exactly. While if the Chi square value is greater than zero, they do not agree.
#
# The sampling distribution of Chi Square is approximated very closely by the *Chi-Square distribution*
# ### The Chi Square Test for Goodness of Fit
#
# We can now use the [Chi-Square test](http://stattrek.com/chi-square-test/goodness-of-fit.aspx?Tutorial=AP) can be used to determine how well a theoretical distribution fits an observed empirical distribution.
#
# Scipy will basically be constructing and looking up this table for us:
#
# 
#
# Let's go ahead and do an example problem. Say you are at a casino and are in charge of monitoring a [craps](http://en.wikipedia.org/wiki/Craps)(a dice game where two dice are rolled). You are suspcious that a player may have switched out the casino's dice for their own. How do we use the Chi-Square test to check whether or not this player is cheating?
#
# You will need some observations in order to begin. You begin to keep track of this player's roll outcomes.You record the next 500 rolls taking note of the sum of the dice roll result and the number of times it occurs.
#
# You record the following:
# <table>
# <td>Sum of Dice Roll</td>
# <td>2</td>
# <td>3</td>
# <td>4</td>
# <td>5</td>
# <td>6</td>
# <td>7</td>
# <td>8</td>
# <td>9</td>
# <td>10</td>
# <td>11</td>
# <td>12</td>
# <tr>
# <td>Number of Times Observed</td>
# <td>8</td>
# <td>32</td>
# <td>48</td>
# <td>59</td>
# <td>67</td>
# <td>84</td>
# <td>76</td>
# <td>57</td>
# <td>34</td>
# <td>28</td>
# <td>7</td>
# </tr>
# </table>
# Now we also know the espected frequency of these sums for a fair dice. That frequency distribution looks like this:
# <table>
# <td>Sum of Dice Roll</td>
# <td>2</td>
# <td>3</td>
# <td>4</td>
# <td>5</td>
# <td>6</td>
# <td>7</td>
# <td>8</td>
# <td>9</td>
# <td>10</td>
# <td>11</td>
# <td>12</td>
# </tr>
# <tr>
# <td>Expected Frequency</td>
# <td>1/36</td>
# <td>2/36</td>
# <td>3/36</td>
# <td>4/36</td>
# <td>5/36</td>
# <td>6/36</td>
# <td>5/36</td>
# <td>4/36</td>
# <td>3/36</td>
# <td>2/36</td>
# <td>1/36</td>
# </tr>
# </table>
# Now we can calculated the expected number of rolls by multiplying the expected frequency with the total sum of the rolls (500 rolls).
# Check sum of the rolls
observed = [8,32,48,59,67,84,76,57,34,28,7]
roll_sum = sum(observed)
roll_sum
# +
# The expected frequency
freq = [1,2,3,4,5,6,5,4,3,2,1]
# Note use of float for python 2.7
possible_rolls = 1.0/36
freq = [possible_rolls*dice for dice in freq]
#Check
freq
# -
# Excellent, now let's multiply our frequency by the sum to get the expected number of rolls for each frequency.
expected = [roll_sum*f for f in freq]
expected
# We can now use Scipy to perform the [Chi Square Test](http://docs.scipy.org/doc/scipy-0.15.1/reference/generated/scipy.stats.chisquare.html) by using chisquare.
# +
from scipy import stats
chisq,p = stats.chisquare(observed,expected)
# -
chisq# chi-squared value
p#value
| DataScience365/Day-19/Chi_Square.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # The Annotated Transformer
# ---
# from http://nlp.seas.harvard.edu/2018/04/03/attention
#
# 
#
# > The Transformer from “Attention is All You Need” has been on a lot of people’s minds over the last year. Besides producing major improvements in translation quality, it provides a new architecture for many other NLP tasks.
#
# > The goal of reducing sequential computation also forms the foundation of the Extended Neural GPU, ByteNet and ConvS2S, all of which use convolutional neural networks as basic building block, computing hidden representations in parallel for all input and output positions. In these models, the number of operations required to relate signals from two arbitrary input or output positions grows in the distance between positions, linearly for ConvS2S and logarithmically for ByteNet. This makes it more difficult to learn dependencies between distant positions. In the Transformer this is reduced to a constant number of operations, albeit at the cost of reduced effective resolution due to averaging attention-weighted positions, an effect we counteract with Multi-Head Attention.
#
# > Self-attention, sometimes called intra-attention is an attention mechanism relating different positions of a single sequence in order to compute a representation of the sequence. Self-attention has been used successfully in a variety of tasks including reading comprehension, abstractive summarization, textual entailment and learning task-independent sentence representations. End- to-end memory networks are based on a recurrent attention mechanism instead of sequencealigned recurrence and have been shown to perform well on simple- language question answering and language modeling tasks.
#
# > To the best of our knowledge, however, the Transformer is the first transduction model relying entirely on self-attention to compute representations of its input and output without using sequence aligned RNNs or convolution.
# ## Module Imports
# +
import math
import copy
import time
import numpy as np
import torch
import torch.nn as nn
import torch.nn.functional as F
import matplotlib.pyplot as plt
import seaborn
seaborn.set_context(context="talk")
# %matplotlib inline
# -
# ## Model Architecture
# ---
# > Most competitive neural sequence transduction models have an encoder-decoder structure ([cite](https://arxiv.org/abs/1409.0473)). Here, the encoder maps an input sequence of symbol representations $(x_1,…,x_n)$ to a sequence of continuous representations $z=(z_1,…,z_n)$. Given z, the decoder then generates an output sequence $(y_1,…,y_m)$ of symbols one element at a time. At each step the model is auto-regressive ([cite](https://arxiv.org/abs/1308.0850)), consuming the previously generated symbols as additional input when generating the next.
#
# > The Transformer follows this overall architecture using stacked self-attention and point-wise, fully connected layers for both the encoder and decoder, shown in the left and right halves of Figure 1, respectively.
#
# 
#
# * [Embedding](#embedding)
# * [Positional Encoding](#positional_encoding)
# * [Encoder](#encoder)
# * [Encoder Layer](#encoder_layer)
# * [Sublayer Connections](#sublayer_connections):
# * [Multi-head Attention](#multi-head_attention)
# * [Position-wise Feed-Forward](#position-wise_feed-forward)
# * [Decoder](#decoder)
# * [Decoder Layer](#decoder_layer)
# * [Sublayer Connections](#sublayer_connections):
# * [Mask](#mask)
# * [Multi-head Attention](#multi-head_attention)
# * [Position-wise Feed-Forward](#position-wise_feed-forward)
# * [Generator](#generator)
class EncoderDecoder(nn.Module):
"""
A standard Encoder-Decoder architecture. Base for this and many
other models.
"""
def __init__(self, encoder, decoder, src_embed, tgt_embed, generator):
super(EncoderDecoder, self).__init__()
self.encoder = encoder
self.decoder = decoder
self.src_embed = src_embed
self.tgt_embed = tgt_embed
self.generator = generator
def forward(self, src, tgt, src_mask, tgt_mask):
"Take in and process masked src and target sequences."
return self.decode(self.encode(src, src_mask), src_mask,
tgt, tgt_mask)
def encode(self, src, src_mask):
return self.encoder(self.src_embed(src), src_mask)
def decode(self, memory, src_mask, tgt, tgt_mask):
return self.decoder(self.tgt_embed(tgt), memory, src_mask, tgt_mask)
# <a id='generator'></a>
# The generator part
#
# 
class Generator(nn.Module):
"Define standard linear + softmax generation step."
def __init__(self, d_model, vocab):
super(Generator, self).__init__()
self.proj = nn.Linear(d_model, vocab)
def forward(self, x):
return F.log_softmax(self.proj(x), dim=-1)
# ## Encoder and Decoder Stacks
# ### Encoder
# > The encoder is composed of a stack of $N=6$ identical layers.
def clones(module, N):
"Produce N identical layers."
return nn.ModuleList([copy.deepcopy(module) for _ in range(N)])
# <a id='encoder'></a>
# $N$ encoder layers stacked together to form the encoder module
#
# 
class Encoder(nn.Module):
"Core encoder is a stack of N layers"
def __init__(self, layer, N):
super(Encoder, self).__init__()
self.layers = clones(layer, N)
self.norm = nn.LayerNorm(layer.size)
def forward(self, x, mask):
"Pass the input (and mask) through each layer in turn."
for layer in self.layers:
x = layer(x, mask)
return self.norm(x)
# > We employ a residual connection ([cite](https://arxiv.org/abs/1512.03385)) around each of the two sub-layers, followed by layer normalization ([cite](https://arxiv.org/abs/1607.06450)).
#
# > That is, the output of each sub-layer is LayerNorm(x+Sublayer(x)), where Sublayer(x) is the function implemented by the sub-layer itself. We apply dropout (cite) to the output of each sub-layer, before it is added to the sub-layer input and normalized.
#
# > To facilitate these residual connections, all sub-layers in the model, as well as the embedding layers, produce outputs of dimension dmodel=512.
# <a id='sublayer_connection'></a>
# Sublayer Connection
#
# 
class SublayerConnection(nn.Module):
"""
A residual connection followed by a layer norm.
Note for code simplicity the norm is first as opposed to last.
"""
def __init__(self, size, dropout):
super(SublayerConnection, self).__init__()
self.norm = nn.LayerNorm(size)
self.dropout = nn.Dropout(dropout)
def forward(self, x, sublayer):
"Apply residual connection to any sublayer with the same size."
return x + self.dropout(sublayer(self.norm(x)))
# > Each layer has two sub-layers. The first is a multi-head self-attention mechanism, and the second is a simple, position-wise fully connected feed- forward network.
# <a id='encoder_layer'></a>
# Individual encoder layer
#
# !!! The implementation here is slightly different from how it appears in this image !!!
#
# According to the `forward()` method of `SublayerConnection` above, it **first applies normalization to the input**, performs either attention or feed-forward operation, applies dropout, and then adds the original input (residual connection). **No normalization is performed after the process**.
#
# 
class EncoderLayer(nn.Module):
"Encoder is made up of self-attn and feed forward (defined below)"
def __init__(self, size, self_attn, feed_forward, dropout):
super(EncoderLayer, self).__init__()
self.self_attn = self_attn
self.feed_forward = feed_forward
self.sublayer = clones(SublayerConnection(size, dropout), 2)
self.size = size
def forward(self, x, mask):
"Follow Figure 1 (left) for connections."
x = self.sublayer[0](x, lambda x: self.self_attn(x, x, x, mask))
return self.sublayer[1](x, self.feed_forward)
# ### Decoder
# > The decoder is also composed of a stack of $N=6$ identical layers.
# <a id='decoder'></a>
# $N$ decoder layers stacked together to form the decoder module
#
# 
class Decoder(nn.Module):
"Generic N layer decoder with masking."
def __init__(self, layer, N):
super(Decoder, self).__init__()
self.layers = clones(layer, N)
self.norm = LayerNorm(layer.size)
def forward(self, x, memory, src_mask, tgt_mask):
for layer in self.layers:
x = layer(x, memory, src_mask, tgt_mask)
return self.norm(x)
# > In addition to the two sub-layers in each encoder layer, the decoder inserts a third sub-layer, which performs multi-head attention over the output of the encoder stack. Similar to the encoder, we employ residual connections around each of the sub-layers, followed by layer normalization.
# <a id='decoder_layer'></a>
# Individual decoder layer
#
# !!! Similar to the encoder layer, the implementation here is slightly different from how it appears in this image !!!
#
# Again, **normalization is applied to the input** but **not to the output after the residual connection**.
#
# 
class DecoderLayer(nn.Module):
"Decoder is made of self-attn, src-attn, and feed forward (defined below)"
def __init__(self, size, self_attn, src_attn, feed_forward, dropout):
super(DecoderLayer, self).__init__()
self.size = size
self.self_attn = self_attn
self.src_attn = src_attn
self.feed_forward = feed_forward
self.sublayer = clones(SublayerConnection(size, dropout), 3)
def forward(self, x, memory, src_mask, tgt_mask):
"Follow Figure 1 (right) for connections."
m = memory
x = self.sublayer[0](x, lambda x: self.self_attn(x, x, x, tgt_mask))
x = self.sublayer[1](x, lambda x: self.src_attn(x, m, m, src_mask))
return self.sublayer[2](x, self.feed_forward)
# > We also modify the self-attention sub-layer in the decoder stack to prevent positions from attending to subsequent positions. This masking, combined with fact that the output embeddings are offset by one position, ensures that the predictions for position $i$ can depend only on the known outputs at positions less than $i$.
# <a id='mask'></a>
# Mask is used in the first multi-head attention layers in the decoder
#
# 
def subsequent_mask(size):
"Mask out subsequent positions."
# attn_shape = (1, size, size)
# subsequent_mask = np.triu(np.ones(attn_shape), k=1).astype('uint8')
# return torch.from_numpy(subsequent_mask) == 0
return torch.tril(torch.ones(1, size, size))
# Below the attention mask shows the position each tgt word (row) is allowed to look at (column). Words are blocked for attending to future words during training.
plt.figure(figsize=(5,5))
plt.imshow(subsequent_mask(20)[0]);
# ## Attention
# ---
# > An attention function can be described as mapping a query and a set of key-value pairs to an output, where the query, keys, values, and output are all vectors. The output is computed as a weighted sum of the values, where the weight assigned to each value is computed by a compatibility function of the query with the corresponding key.
#
# > We call our particular attention “Scaled Dot-Product Attention”. The input consists of queries and keys of dimension $d_k$, and values of dimension $d_v$. We compute the dot products of the query with all keys, divide each by $\sqrt{d_k}$, and apply a softmax function to obtain the weights on the values.
#
# 
#
# > In practice, we compute the attention function on a set of queries simultaneously, packed together into a matrix $Q$. The keys and values are also packed together into matrices $K$ and $V$. We compute the matrix of outputs as:
# $Attention(Q, K, V) = softmax(\dfrac{QK^T}{\sqrt{d_k}})V$
def attention(query, key, value, mask=None, dropout=None):
"Compute 'Scaled Dot Product Attention'"
d_k = query.size(-1)
scores = torch.matmul(query, key.transpose(-2, -1)) \
/ math.sqrt(d_k)
if mask is not None:
scores = scores.masked_fill(mask == 0, -1e9)
p_attn = F.softmax(scores, dim=-1)
if dropout is not None:
p_attn = dropout(p_attn)
return torch.matmul(p_attn, value), p_attn
# > The two most commonly used attention functions are additive attention ([cite](https://arxiv.org/abs/1409.0473)), and dot-product (multiplicative) attention. Dot-product attention is identical to our algorithm, except for the scaling factor of $\dfrac{1}{\sqrt{d_k}}$. Additive attention computes the compatibility function using a feed-forward network with a single hidden layer. While the two are similar in theoretical complexity, dot-product attention is much faster and more space-efficient in practice, since it can be implemented using highly optimized matrix multiplication code.
#
# > While for small values of $d_k$ the two mechanisms perform similarly, additive attention outperforms dot product attention without scaling for larger values of $d_k$ ([cite](https://arxiv.org/abs/1703.03906)). We suspect that for large values of $d_k$, the dot products grow large in magnitude, pushing the softmax function into regions where it has extremely small gradients (To illustrate why the dot products get large, assume that the components of $q$ and $k$ are independent random variables with mean $0$ and variance $1$. Then their dot product, $q \cdot k = \sum^{d_k}_{i=1} q_i k_i$, has mean $0$ and variance $d_k$.). To counteract this effect, we scale the dot products by $\dfrac{1}{\sqrt{d_k}}$.
#
# 
#
# > Multi-head attention allows the model to jointly attend to information from different representation subspaces at different positions. With a single attention head, averaging inhibits this.
#
# > $MultiHead(Q,K,V) = Concat(head_1,...,head_h)W^O \\ where \; head_i = Attention(QW^Q_i,KW^K_i,VW^V_i)$
#
# > Where the projections are parameter matrices $W^Q_i \in \mathbb{R}^{d_{model} \times d_k}$, $W^K_i \in \mathbb{R}^{d_{model} \times d_k}$, $W^V_i \in \mathbb{R}^{d_{model} \times d_v}$ and $W^O \in \mathbb{R}^{hd_v \times d_{model}}$. In this work we employ $h=8$ parallel attention layers, or heads. For each of these we use $d_k = d_v = \dfrac{d_{model}}{h} = 64$. Due to the reduced dimension of each head, the total computational cost is similar to that of single-head attention with full dimensionality.
# <a id='multi-head_attention'></a>
# Multi-head Attention
#
# 
class MultiHeadedAttention(nn.Module):
def __init__(self, h, d_model, dropout=0.1):
"Take in model size and number of heads."
super(MultiHeadedAttention, self).__init__()
assert d_model % h == 0
# We assume d_v always equals d_k
self.d_k = d_model // h
self.h = h
self.linears = clones(nn.Linear(d_model, d_model), 4)
self.attn = None
self.dropout = nn.Dropout(p=dropout)
def forward(self, query, key, value, mask=None):
"Implements Figure 2"
if mask is not None:
# Same mask applied to all h heads.
mask = mask.unsqueeze(1)
nbatches = query.size(0)
# 1) Do all the linear projections in batch from d_model => h x d_k
query, key, value = \
[l(x).view(nbatches, -1, self.h, self.d_k).transpose(1, 2)
for l, x in zip(self.linears, (query, key, value))]
# 2) Apply attention on all the projected vectors in batch.
x, self.attn = attention(query, key, value, mask=mask,
dropout=self.dropout)
# 3) "Concat" using a view and apply a final linear.
x = x.transpose(1, 2).contiguous() \
.view(nbatches, -1, self.h * self.d_k)
return self.linears[-1](x)
# ### Applications of Attention in our Model
#
# > The Transformer uses multi-head attention in three different ways:
#
# > 1) In “encoder-decoder attention” layers, the queries come from the previous decoder layer, and the memory keys and values come from the output of the encoder. This allows every position in the decoder to attend over all positions in the input sequence. This mimics the typical encoder-decoder attention mechanisms in sequence-to-sequence models such as ([cite](https://arxiv.org/abs/1609.08144)).
#
# > 2) The encoder contains self-attention layers. In a self-attention layer all of the keys, values and queries come from the same place, in this case, the output of the previous layer in the encoder. Each position in the encoder can attend to all positions in the previous layer of the encoder.
#
# > 3) Similarly, self-attention layers in the decoder allow each position in the decoder to attend to all positions in the decoder up to and including that position. We need to prevent leftward information flow in the decoder to preserve the auto-regressive property. We implement this inside of scaled dot- product attention by masking out (setting to $−\infty$) all values in the input of the softmax which correspond to illegal connections.
# ## Position-wise Feed-Forward Networks
# ---
# > In addition to attention sub-layers, each of the layers in our encoder and decoder contains a fully connected feed-forward network, which is applied to each position **separately** and **identically**. This consists of two linear transformations with a ReLU activation in between.
#
# > $FFN(x) = \max(0,xW_1+b_1)W_2 + b_2$
#
# > While the linear transformations are the same across different positions, they use different parameters from layer to layer. Another way of describing this is as two convolutions with kernel size $1$. The dimensionality of input and output is $d_{model} = 512$, and the inner-layer has dimensionality $d_{ff} = 2048$.
# <a id='position-wise_feed-forward'></a>
# Position-wise Feed-Forward
#
# 
class PositionwiseFeedForward(nn.Module):
"Implements FFN equation."
def __init__(self, d_model, d_ff, dropout=0.1):
super(PositionwiseFeedForward, self).__init__()
self.w_1 = nn.Linear(d_model, d_ff)
self.w_2 = nn.Linear(d_ff, d_model)
self.dropout = nn.Dropout(dropout)
def forward(self, x):
return self.w_2(self.dropout(F.relu(self.w_1(x))))
# ## Embeddings and Softmax
# ---
# > Similarly to other sequence transduction models, we use learned embeddings to convert the input tokens and output tokens to vectors of dimension $d_{model}$. We also use the usual learned linear transformation and softmax function to convert the decoder output to predicted next-token probabilities. In our model, we share the same weight matrix between the two embedding layers and the pre-softmax linear transformation, similar to ([cite](https://arxiv.org/abs/1608.05859)). In the embedding layers, we multiply those weights by $\sqrt{d_{model}}$.
# <a id='embedding'></a>
# Embedding Layers
#
# 
class Embeddings(nn.Module):
def __init__(self, d_model, vocab):
super(Embeddings, self).__init__()
self.lut = nn.Embedding(vocab, d_model)
self.d_model = d_model
def forward(self, x):
return self.lut(x) * math.sqrt(self.d_model)
# ## Positional Encoding
# ---
# > Since our model contains no recurrence and no convolution, in order for the model to make use of the order of the sequence, we must inject some information about the relative or absolute position of the tokens in the sequence. To this end, we add “positional encodings” to the input embeddings at the bottoms of the encoder and decoder stacks. The positional encodings have the same dimension $d_{model}$ as the embeddings, so that the two can be summed. There are many choices of positional encodings, learned and fixed ([cite](https://arxiv.org/pdf/1705.03122.pdf)).
#
# > In this work, we use sine and cosine functions of different frequencies:
#
# > $PE_{(pos,2i)} = \sin(\dfrac{pos}{10000^{2i/d_{model}}})$
#
# > $PE_{(pos,2i+1)} = \cos(\dfrac{pos}{10000^{2i/d_{model}}})$
#
# > where $pos$ is the position and $i$ is the dimension. That is, each dimension of the positional encoding corresponds to a sinusoid. The wavelengths form a geometric progression from $2\pi$ to $10000 \cdot 2\pi$. We chose this function because we hypothesized it would allow the model to easily learn to attend by relative positions, since for any fixed offset $k$, $PE_{pos+k}$ can be represented as a linear function of $PE_{pos}$.
#
# > In addition, we apply dropout to the sums of the embeddings and the positional encodings in both the encoder and decoder stacks. For the base model, we use a rate of $P_{drop} = 0.1$.
# <a id='positional_encoding'></a>
# Positional Encoding
#
# 
class PositionalEncoding(nn.Module):
"Implement the PE function."
def __init__(self, d_model, dropout, max_len=5000):
super(PositionalEncoding, self).__init__()
self.dropout = nn.Dropout(p=dropout)
# Compute the positional encodings once in log space.
pe = torch.zeros(max_len, d_model)
position = torch.arange(0, max_len, dtype=torch.float).unsqueeze(1)
div_term = torch.exp(torch.arange(0, d_model, 2, dtype=torch.float) *
-(math.log(10000.0) / d_model))
pe[:, 0::2] = torch.sin(position * div_term)
pe[:, 1::2] = torch.cos(position * div_term)
pe = pe.unsqueeze(0)
self.register_buffer('pe', pe)
def forward(self, x):
x = x + self.pe[:, :x.size(1)]
return self.dropout(x)
# Below the positional encoding will add in a sine wave based on position. The frequency and offset of the wave is different for each dimension.
plt.figure(figsize=(15, 5))
pe = PositionalEncoding(20, 0)
y = pe.forward(torch.zeros(1, 100, 20))
plt.plot(np.arange(100), y[0, :, 4:8].data.numpy())
plt.grid(True)
plt.legend(["dim %d"%p for p in [4,5,6,7]]);
# > We also experimented with using learned positional embeddings ([cite](https://arxiv.org/pdf/1705.03122.pdf)) instead, and found that the two versions produced nearly identical results. We chose the sinusoidal version because it may allow the model to extrapolate to sequence lengths longer than the ones encountered during training.
# ## Full Model
# ---
# Here we define a function that takes in hyperparameters and produces a full model.
def make_model(src_vocab, tgt_vocab, N=6,
d_model=512, d_ff=2048, h=8, dropout=0.1):
"Helper: Construct a model from hyperparameters."
c = copy.deepcopy
attn = MultiHeadedAttention(h, d_model)
ff = PositionwiseFeedForward(d_model, d_ff, dropout)
position_encoding = PositionalEncoding(d_model, dropout)
model = EncoderDecoder(
Encoder(layer=EncoderLayer(size=d_model, self_attn=c(attn), feed_forward=c(ff), dropout=dropout),
N=N),
Decoder(layer=DecoderLayer(size=d_model, self_attn=c(attn), src_attn=c(attn), feed_forward=c(ff), dropout=dropout),
N=N),
nn.Sequential(Embeddings(d_model, src_vocab), # src_embed
c(position_encoding)),
nn.Sequential(Embeddings(d_model, tgt_vocab), # tgt_embed
c(position_encoding)),
Generator(d_model, tgt_vocab)
)
# This was important from their code.
# Initialize parameters with Glorot / fan_avg.
for p in model.parameters():
if p.dim() > 1:
nn.init.xavier_uniform_(p)
return model
# Small example model.
tmp_model = make_model(src_vocab=10, tgt_vocab=10, N=2);
# ## Training
# ---
# > This section describes the training regime for our models.
#
# We stop for a quick interlude to introduce some of the tools needed to train a standard encoder decoder model. First we define a batch object that holds the src and target sentences for training, as well as constructing the masks.
# ### Batches and Masking
class Batch:
"Object for holding a batch of data with mask during training."
def __init__(self, src, trg=None, pad=0):
self.src = src
self.src_mask = (src != pad).unsqueeze(-2)
if trg is not None:
self.trg = trg[:, :-1]
self.trg_y = trg[:, 1:]
self.trg_mask = \
self.make_std_mask(self.trg, pad)
self.ntokens = (self.trg_y != pad).data.sum().item()
@staticmethod
def make_std_mask(tgt, pad):
"Create a mask to hide padding and future words."
tgt_mask = (tgt != pad).unsqueeze(-2)
tgt_mask = tgt_mask & subsequent_mask(tgt.size(-1)).type_as(tgt_mask.data)
return tgt_mask
# Next we create a generic training and scoring function to keep track of loss. We pass in a generic loss compute function that also handles parameter updates.
# ### Training Loop
def run_epoch(data_iter, model, loss_compute):
"Standard Training and Logging Function"
start = time.time()
total_tokens = 0
total_loss = 0
tokens = 0
for i, batch in enumerate(data_iter):
out = model.forward(batch.src, batch.trg,
batch.src_mask, batch.trg_mask)
loss = loss_compute(out, batch.trg_y, batch.ntokens)
total_loss += loss
total_tokens += batch.ntokens
tokens += batch.ntokens
elapsed = time.time() - start
print("Epoch Step: {:d} | Loss: {:f} | Running Loss: {:f} | Tokens per Sec: {:f} ".format(i, loss / batch.ntokens, total_loss / total_tokens, tokens / elapsed), end='\r')
start = time.time()
tokens = 0
print()
return total_loss / total_tokens
# ### Training Data and Batching
#
# > We trained on the standard WMT 2014 English-German dataset consisting of about 4.5 million sentence pairs. Sentences were encoded using byte-pair encoding, which has a shared source-target vocabulary of about 37000 tokens. For English- French, we used the significantly larger WMT 2014 English-French dataset consisting of 36M sentences and split tokens into a 32000 word-piece vocabulary.
#
# > Sentence pairs were batched together by approximate sequence length. Each training batch contained a set of sentence pairs containing approximately 25000 source tokens and 25000 target tokens.
#
# We will use torch text for batching. This is discussed in more detail below. Here we create batches in a torchtext function that ensures our batch size padded to the maximum batchsize does not surpass a threshold (25000 if we have 8 gpus).
global max_src_in_batch, max_tgt_in_batch
def batch_size_fn(new, count, sofar):
"Keep augmenting batch and calculate total number of tokens + padding."
global max_src_in_batch, max_tgt_in_batch
if count == 1:
max_src_in_batch = 0
max_tgt_in_batch = 0
max_src_in_batch = max(max_src_in_batch, len(new.src))
max_tgt_in_batch = max(max_tgt_in_batch, len(new.trg) + 2)
src_elements = count * max_src_in_batch
tgt_elements = count * max_tgt_in_batch
return max(src_elements, tgt_elements)
# ## Hardware and Schedule
# ---
# We trained our models on one machine with 8 NVIDIA P100 GPUs. For our base models using the hyperparameters described throughout the paper, each training step took about 0.4 seconds. We trained the base models for a total of 100,000 steps or 12 hours. For our big models, step time was 1.0 seconds. The big models were trained for 300,000 steps (3.5 days).
# ## Optimizer
# ---
# > We used the Adam optimizer ([cite](https://arxiv.org/abs/1412.6980)) with $\beta_1 = 0.9$, $\beta_2 = 0.98$ and $\epsilon = 10^{−9}$. We varied the learning rate over the course of training, according to the formula: $l_{rate} = d^{−0.5}_{model} \cdot \min(step\_num^{−0.5},step\_num \cdot warmup\_steps^{−1.5})$ This corresponds to increasing the learning rate linearly for the first warmupsteps training steps, and decreasing it thereafter proportionally to the inverse square root of the step number. We used $warmup\_steps = 4000$.
#
# Note: This part is very important. Need to train with this setup of the model.
# +
class NoamOpt:
"Optim wrapper that implements rate."
def __init__(self, model_size, factor, warmup, optimizer):
self.optimizer = optimizer
self._step = 0
self.warmup = warmup
self.factor = factor
self.model_size = model_size
self._rate = 0
def step(self):
"Update parameters and rate"
self._step += 1
rate = self.rate()
for p in self.optimizer.param_groups:
p['lr'] = rate
self._rate = rate
self.optimizer.step()
def rate(self, step = None):
"Implement `lrate` above"
if step is None:
step = self._step
return self.factor * \
(self.model_size ** (-0.5) *
min(step ** (-0.5), step * self.warmup ** (-1.5)))
def get_std_opt(model):
return NoamOpt(model.src_embed[0].d_model, 2, 4000,
torch.optim.Adam(model.parameters(), lr=0, betas=(0.9, 0.98), eps=1e-9))
# -
# Example of the curves of this model for different model sizes and for optimization hyperparameters.
# Three settings of the lrate hyperparameters.
opts = [NoamOpt(512, 1, 4000, None),
NoamOpt(512, 1, 8000, None),
NoamOpt(256, 1, 4000, None)]
plt.plot(np.arange(1, 20000), [[opt.rate(i) for opt in opts] for i in range(1, 20000)])
plt.legend(["512:4000", "512:8000", "256:4000"]);
# ## Regularization
# ---
# ### Label Smoothing
#
# > During training, we employed label smoothing of value $\epsilon_{ls} = 0.1$ ([cite](https://arxiv.org/abs/1512.00567)). This hurts perplexity, as the model learns to be more unsure, but improves accuracy and BLEU score.
#
# We implement label smoothing using the KL div loss. Instead of using a one-hot target distribution, we create a distribution that has confidence of the correct word and the rest of the smoothing mass distributed throughout the vocabulary.
class LabelSmoothing(nn.Module):
"Implement label smoothing."
def __init__(self, size, padding_idx, smoothing=0.0):
super(LabelSmoothing, self).__init__()
self.criterion = nn.KLDivLoss(reduction='sum')
self.padding_idx = padding_idx
self.confidence = 1.0 - smoothing
self.smoothing = smoothing
self.size = size
self.true_dist = None
def forward(self, x, target):
assert x.size(1) == self.size
true_dist = x.data.clone()
true_dist.fill_(self.smoothing / (self.size - 2))
true_dist.scatter_(1, target.data.unsqueeze(1), self.confidence)
true_dist[:, self.padding_idx] = 0
mask = torch.nonzero(target.data == self.padding_idx)
if mask.dim() > 0:
true_dist.index_fill_(0, mask.squeeze(), 0.0)
self.true_dist = true_dist
return self.criterion(x, true_dist)
# Here we can see an example of how the mass is distributed to the words based on confidence.
# +
# Example of label smoothing.
crit = LabelSmoothing(5, 0, 0.4)
predict = torch.FloatTensor([[0, 0.2, 0.7, 0.1, 0],
[0, 0.2, 0.7, 0.1, 0],
[0, 0.2, 0.7, 0.1, 0]])
v = crit(predict.log(),
torch.LongTensor([2, 1, 0]))
# Show the target distributions expected by the system.
plt.imshow(crit.true_dist);
# -
# Label smoothing actually starts to penalize the model if it gets very confident about a given choice.
crit = LabelSmoothing(5, 0, 0.1)
def loss(x):
d = x + 3 * 1
predict = torch.FloatTensor([[0, x / d, 1 / d, 1 / d, 1 / d],
])
#print(predict)
return crit(predict.log(),
torch.LongTensor([1])).item()
plt.plot(np.arange(1, 100), [loss(x) for x in range(1, 100)]);
# ## A First Example
# ---
# We can begin by trying out a simple copy-task. Given a random set of input symbols from a small vocabulary, the goal is to generate back those same symbols.
loss(1)
# ### Synthetic Data
def data_gen(V, batch, nbatches):
"Generate random data for a src-tgt copy task."
for i in range(nbatches):
data = torch.randint(1, V, size=(batch, 10), dtype=torch.long)
data[:, 0] = 1
src = data
tgt = data
yield Batch(src, tgt, 0)
# ### Loss Computation
class SimpleLossCompute:
"A simple loss compute and train function."
def __init__(self, generator, criterion, opt=None):
self.generator = generator
self.criterion = criterion
self.opt = opt
def __call__(self, x, y, norm):
x = self.generator(x)
loss = self.criterion(x.contiguous().view(-1, x.size(-1)),
y.contiguous().view(-1)) / norm
loss.backward()
if self.opt is not None:
self.opt.step()
self.opt.optimizer.zero_grad()
return loss.item() * norm
# ### Greedy Decoding
# +
# Train the simple copy task.
V = 11
criterion = LabelSmoothing(size=V, padding_idx=0, smoothing=0.0)
model = make_model(V, V, N=2)
model_opt = NoamOpt(model.src_embed[0].d_model, 1, 400,
torch.optim.Adam(model.parameters(), lr=0, betas=(0.9, 0.98), eps=1e-9))
for epoch in range(10):
model.train()
print("--- Epoch", epoch, "---")
print("Training")
run_epoch(data_gen(V, 30, 20), model,
SimpleLossCompute(model.generator, criterion, model_opt))
model.eval()
print("Evaluation")
run_epoch(data_gen(V, 30, 5), model,
SimpleLossCompute(model.generator, criterion, None))
# -
# This code predicts a translation using greedy decoding for simplicity.
# +
def greedy_decode(model, src, src_mask, max_len, start_symbol):
memory = model.encode(src, src_mask)
ys = torch.ones(1, 1).fill_(start_symbol).type_as(src.data)
for i in range(max_len-1):
out = model.decode(memory, src_mask,
ys,
subsequent_mask(ys.size(1)))
prob = model.generator(out[:, -1])
_, next_word = torch.max(prob, dim = 1)
next_word = next_word.data[0]
ys = torch.cat([ys,
torch.ones(1, 1).type_as(src.data).fill_(next_word)], dim=1)
return ys
model.eval()
src = torch.LongTensor([[1,2,3,4,5,6,7,8,9,10]])
src_mask = torch.ones(1, 1, 10)
print(greedy_decode(model, src, src_mask, max_len=10, start_symbol=1))
# -
# ## Results
# ---
# > On the WMT 2014 English-to-German translation task, the big transformer model (Transformer (big) in Table 2) outperforms the best previously reported models (including ensembles) by more than 2.0 BLEU, establishing a new state-of-the-art BLEU score of 28.4. The configuration of this model is listed in the bottom line of Table 3. Training took 3.5 days on 8 P100 GPUs. Even our base model surpasses all previously published models and ensembles, at a fraction of the training cost of any of the competitive models.
#
# > On the WMT 2014 English-to-French translation task, our big model achieves a BLEU score of 41.0, outperforming all of the previously published single models, at less than 1/4 the training cost of the previous state-of-the-art model. The Transformer (big) model trained for English-to-French used dropout rate Pdrop = 0.1, instead of 0.3.
#
# 
#
# The code we have written here is a version of the base model. There are fully trained version of this system available here [(Example Models)](http://opennmt.net/Models-py/).
#
# With the addtional extensions in the last section, the OpenNMT-py replication gets to 26.9 on EN-DE WMT. Here I have loaded in those parameters to our reimplemenation.
# !wget https://s3.amazonaws.com/opennmt-models/en-de-model.pt
model, SRC, TGT = torch.load("en-de-model.pt")
model.eval()
sent = "▁The ▁log ▁file ▁can ▁be ▁sent ▁secret ly ▁with ▁email ▁or ▁FTP ▁to ▁a ▁specified ▁receiver".split()
src = torch.LongTensor([[SRC.stoi[w] for w in sent]])
src = src
src_mask = (src != SRC.stoi["<blank>"]).unsqueeze(-2)
out = greedy_decode(model, src, src_mask,
max_len=60, start_symbol=TGT.stoi["<s>"])
print("Translation:", end="\t")
trans = "<s> "
for i in range(1, out.size(1)):
sym = TGT.itos[out[0, i]]
if sym == "</s>": break
trans += sym + " "
print(trans)
# ## Attention Visualization
# ---
# Even with a greedy decoder the translation looks pretty good. We can further visualize it to see what is happening at each layer of the attention
# +
tgt_sent = trans.split()
def draw(data, x, y, ax):
seaborn.heatmap(data,
xticklabels=x, square=True, yticklabels=y, vmin=0.0, vmax=1.0,
cbar=False, ax=ax)
for layer in range(1, 6, 2):
fig, axs = plt.subplots(1,4, figsize=(20, 10))
print("Encoder Layer", layer+1)
for h in range(4):
draw(model.encoder.layers[layer].self_attn.attn[0, h].data,
sent, sent if h ==0 else [], ax=axs[h])
plt.show()
for layer in range(1, 6, 2):
fig, axs = plt.subplots(1,4, figsize=(20, 10))
print("Decoder Self Layer", layer+1)
for h in range(4):
draw(model.decoder.layers[layer].self_attn.attn[0, h].data[:len(tgt_sent), :len(tgt_sent)],
tgt_sent, tgt_sent if h ==0 else [], ax=axs[h])
plt.show()
print("Decoder Src Layer", layer+1)
fig, axs = plt.subplots(1,4, figsize=(20, 10))
for h in range(4):
draw(model.decoder.layers[layer].self_attn.attn[0, h].data[:len(tgt_sent), :len(sent)],
sent, tgt_sent if h ==0 else [], ax=axs[h])
plt.show()
# -
# ## Conclusion
# ---
# Hopefully this code is useful for future research. Please reach out if you have any issues. If you find this code helpful, also check out our other OpenNMT tools.
# @inproceedings{opennmt,
# author = {<NAME> and
# <NAME> and
# <NAME> and
# <NAME> and
# <NAME>},
# title = {OpenNMT: Open-Source Toolkit for Neural Machine Translation},
# booktitle = {Proc. ACL},
# year = {2017},
# url = {https://doi.org/10.18653/v1/P17-4012},
# doi = {10.18653/v1/P17-4012}
# }
# Cheers, srush
# ---
| Transformer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/MrLeeking21/CPEN-21A-ECE-2-3/blob/main/Welcome_To_Colaboratory.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="ApLYkqw-A1z-"
# # **Final Exam**
# + [markdown] id="6JprrpdUAvdn"
# ##**Problem Statement 1.**
# Create a python program that will produce an output of sum of 10 numbers less than 5 using for loop statement.
# + id="j45haQ9SBk1M" outputId="3a75d0f9-bd9d-4796-dd23-7d52b0695247" colab={"base_uri": "https://localhost:8080/"}
sum=0
number=(-5,-4,-3,-2,-1,0,1,2,3,4)
for x in number:
sum=sum+x
print("Sum of 10 numbers less than 5 is:",sum)
# + [markdown] id="idmfBGNICCai"
# ##**Problem Statement 2.**
# Create a python program that will produce accept five numbers and determine the sum of first and last number among the five number entered using while loop
# + colab={"base_uri": "https://localhost:8080/"} id="SnwPOghpms5I" outputId="22687cab-2c13-42f0-a9ab-6288499c3906"
w=int(input())
y=w+35
g=w+28
h=w
while w<y:
print(w)
w+=7
print("The sum of first and last numver is",g+h)
# + [markdown] id="iZH8xzJjDtRl"
# ##**Problem Statement 3.**
# Create a Python program to calculate student grades. It accepts a numerical grade as input
# and it will display the character grade as output based on the given scale: (Use Nested-IF-Else statement).
# + colab={"base_uri": "https://localhost:8080/"} id="qe8-AsynpNYm" outputId="a65d0d2b-12bc-4e99-d40a-f9d1d19363be"
grade=float(input("Enter your grade "))
if grade>=90:
print("A")
elif 80<=grade<90:
print("B")
elif 70<=grade<80:
print("C")
elif 60<=grade<70:
print("D")
else:
print("E")
# + id="nuWHFSbXchvx" colab={"base_uri": "https://localhost:8080/"} outputId="f07e5ab7-e1e3-4562-c7fe-b8b7625cc2e9"
grade=int(input("Enter your grade"))
if grade>=90:
print("A")
elif 80<=grade<90:
print("B")
elif 70<=grade<80:
print("C")
elif 60<=grade<70:
print("D")
else:
print("E")
| Welcome_To_Colaboratory.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="xLXWx4uhqtYJ" colab_type="text"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/PreferredAI/tutorials/blob/master/recommender-systems/07_explanations.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/PreferredAI/tutorials/blob/master/recommender-systems/07_explanations.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# </table>
# + [markdown] id="wzPrmZOMtXt1" colab_type="text"
# # Explainable Recommendations
#
# + [markdown] id="cBBggZxMmFG3" colab_type="text"
# While the main objective of a recommender system is to identify the items to be recommended to a user, providing explanations to accompany the recommendations would be more persuasive as well as engender trust and transparency. There are different types of explanations. In this tutorial, we explore explainable recommendation approaches that rely on user product aspect-level sentiment for modeling explanations.
# + [markdown] id="MqktDa7H2hKz" colab_type="text"
# ## 1. Setup
# + id="41oWCMUG2eC_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="8562fb87-9b3e-4737-d119-213eef286bda"
# !pip install --quiet cornac==1.6.1
# + id="YqKrDcGH2k7E" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="60c18f42-8542-46a0-93dd-697b12834779"
import os
import sys
from collections import defaultdict
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
import cornac
from cornac.utils import cache
from cornac.datasets import amazon_toy
from cornac.eval_methods import RatioSplit
from cornac.data import Reader, SentimentModality
from cornac.models import EFM, MTER, NMF, BPR
print(f"System version: {sys.version}")
print(f"Cornac version: {cornac.__version__}")
SEED = 42
VERBOSE = False
# + [markdown] id="a1Ay68vEfD8y" colab_type="text"
# ## 2. Aspect-Level Sentiments
# To model fine-grained product aspect-ratings. Several works rely on sentiment analysis to extract aspect sentiment from product reviews. In other words, each review is now a list of aspect sentiments. Along with product rating, we also have aspect sentiments expressed in users' reviews. Here, we work with Toys and Games dataset, a sub-category of [Amazon reviews](http://jmcauley.ucsd.edu/data/amazon/).
#
# Below are some examples of aspect-level sentiments that have been extracted from users' reviews of items.
# + id="XvS6J8lm_a6M" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 359} outputId="5ce78506-ed70-4365-b63a-1dc39df6c577"
sentiment = amazon_toy.load_sentiment()
samples = sentiment[:10]
pd.DataFrame.from_dict({
"user": [tup[0] for tup in samples],
"item": [tup[1] for tup in samples],
"aspect-level sentiment": [tup[2] for tup in samples]
})
# + id="1cZ0lBVVdlVs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 391} outputId="1e399c24-4552-4161-dc5e-6b3a3c732d0d"
# Load rating and sentiment information
reader = Reader(min_item_freq=20)
rating = amazon_toy.load_feedback(reader)
# Use Sentiment Modality for aspect-level sentiment data
sentiment_modality = SentimentModality(data=sentiment)
rs = RatioSplit(
data=rating,
test_size=0.2,
exclude_unknowns=True,
sentiment=sentiment_modality,
verbose=VERBOSE,
seed=SEED,
)
print("Total number of aspects:", rs.sentiment.num_aspects)
print("Total number of opinions:", rs.sentiment.num_opinions)
id_aspect_map = {v:k for k, v in rs.sentiment.aspect_id_map.items()}
id_opinion_map = {v:k for k, v in rs.sentiment.opinion_id_map.items()}
# + [markdown] id="9vQgJDRUeqNL" colab_type="text"
# ## 3. Explicit Factor Model (EFM)
#
# EFM model extends Non-negative Matrix Factorization (NMF) with the additional information from **aspect-level sentiments**. The objective is to learn user, item, and aspect latent factors to explain user-item ratings, users' interest in certain aspects of the items, as well as the quality of items according to those aspects. In a nutshell, EFM factorizes three matrices: *rating matrix*, *user-aspect attention matrix*, and *item-aspect quality matrix*. Let's take a look at what the later two matrices are.
# + id="vHDD8hUEa9Ja" colab_type="code" colab={}
efm = EFM()
efm.train_set = rs.train_set
_, X, Y = efm._build_matrices(rs.train_set)
# + [markdown] id="NGO3fneebwnf" colab_type="text"
# ### User-Aspect Attention Matrix
#
# Let $\mathcal{F} = \{f_1, f_2, \dots, f_F\}$ be the set of aspects (e.g., screen, earphone).
#
# Let $\mathbf{X} \in \mathbb{R}^{N \times F}$ be a sparse aspect matrix for $N$ users and $F$ aspects, whereby each element $x_{if} \in \mathbf{X}$ indicates the degree of **attention** by user $i$ on aspect $f$, defined as follows:
#
# \begin{equation}
# x_{if} = \
# \begin{cases}
# 0, & \text{if user $i$ never mentions aspect $f$} \\
# 1 + (N-1)\left(\frac{2}{1+\exp(-t_{if})}-1\right), & \text{otherwise}
# \end{cases}
# \end{equation}
#
# where $N=5$ is the highest rating score, $t_{if}$ is the frequency of user $i$ mentions aspect $f$ across all her reviews.
#
# For illustration purpose, we show a small matrix $\mathbf{X}$ of 5 users and 5 aspects below.
# + id="BzLjB2CBbJTj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="d8c95260-9724-44e9-eeca-446ade10676c"
n_users = 5
n_aspects = 5
pd.DataFrame(
data=X[:n_users, :n_aspects].A,
index=[f"User {u + 1}" for u in np.arange(n_users)],
columns=[f"{id_aspect_map[i]}" for i in np.arange(n_aspects)]
)
# + [markdown] id="avkcE4jtfzfO" colab_type="text"
# In the example below, we can see that *User 4* finds the aspect *game* important, whereas *User 3* is concerned with *game* as well as *price*.
# + [markdown] id="PWvgl1rFb1gI" colab_type="text"
# ### Item-Aspect Quality Matrix
#
#
# Let $\mathbf{Y} \in \mathbb{R}^{M \times F}$ be a sparse aspect matrix for $M$ items and $F$ aspects, whereby $y_{jf} \in \mathbf{Y}$ indicates the **quality** of item $j$ on aspect $f$, defined as follows:
#
# \begin{equation}
# y_{jf} = \
# \begin{cases}
# 0, & \text{if item $j$ was never reviewed on aspect $f$} \\
# 1 + (N - 1) \left( \frac{1}{1+\exp(-s_{jf})} \right), & \text{otherwise}
# \end{cases}
# \end{equation}
#
# where $s_{jf}$ is the sum of sentiment values with which item $j$ has been mentioned with regards to aspect $f$ across all its reviews.
#
# We show a small matrix $Y$ of 5 items and 5 aspects below:
# + id="2UxisKhrb7Py" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="3294aae1-9125-4700-8aaf-6d8e9ac66cad"
n_items = 5
n_aspects = 5
pd.DataFrame(
data=Y[:n_items, :n_aspects].A,
index=[f"Item {u + 1}" for u in np.arange(n_items)],
columns=[f"{id_aspect_map[i]}" for i in np.arange(n_aspects)]
)
# + [markdown] id="Wx7p01aAgRPG" colab_type="text"
# We see from the example above that *Item 3* has a positive quality in the aspect *game*, whereas *Item 5* has positive quality on the other 4 aspects.
# + [markdown] id="iXoJMNqVc-1C" colab_type="text"
# ### Optimization
#
# As these matrices are sparse, for prediction, EFM jointly factorizes $X$ and $Y$ matrices along with rating matrix $R$. Learning the latent factors can be done via minimizing the following loss function:
#
# \begin{align}
# &\mathcal{L}(\mathbf{U_1, U_2, V, H_1, H_2} | \lambda_x, \lambda_y, \lambda_u, \lambda_h, \lambda_v) = ||\mathbf{U_1} \mathbf{U_2}^T + \mathbf{H_1} \mathbf{H_2}^T - \mathbf{R}||_F^2 + \lambda_x ||\mathbf{U_1} \mathbf{V}^T - \mathbf{X}||_F^2 + \lambda_y ||\mathbf{U_2} \mathbf{V}^T - \mathbf{Y}||_F^2 + \lambda_u(||\mathbf{U_1}||_F^2+||\mathbf{U_2}||_F^2) + \lambda_h(||\mathbf{H_1}||_F^2+||\mathbf{H_2}||_F^2) + \lambda_v ||\mathbf{V}||_F^2 \\
# &\text{such that: } \forall_{i, k} u_{ik} \ge 0, \forall_{j, k} v_{jk} \ge 0
# \end{align}
#
# The can be solved as a constrained optimization problem.
#
#
# Let's conduct an experiment with EFM model and compare with NMF as a baseline.
# + id="juZD_AzeftIi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 153} outputId="82afa406-ff3d-4013-fb30-c0a3f6d9fe1e"
efm = EFM(
num_explicit_factors=40,
num_latent_factors=60,
num_most_cared_aspects=15,
rating_scale=5.0,
alpha=0.85,
lambda_x=1,
lambda_y=1,
lambda_u=0.01,
lambda_h=0.01,
lambda_v=0.01,
max_iter=100,
verbose=VERBOSE,
seed=SEED,
)
# compare to baseline NMF
nmf = NMF(k=100, max_iter=100, verbose=VERBOSE, seed=SEED)
eval_metrics = [
cornac.metrics.RMSE(),
cornac.metrics.NDCG(k=50),
cornac.metrics.AUC()
]
cornac.Experiment(
eval_method=rs, models=[nmf, efm], metrics=eval_metrics
).run()
# + [markdown] id="rpAMFXCaDUpt" colab_type="text"
# ### Refining Ranking Prediction
#
# With EFM model, you can refine the recommendation after training by experimenting with different values of:
# * `num_most_cared_aspects` ($k$): integer, value range $\in[0, 429]$ as we have $429$ aspects in total
# * `alpha` $\in [0,1]$
#
# These parameters will affect ranking performance of the EFM model, as the ranking score is predicted as follow:
#
# $$
# ranking\_score = \alpha \cdot \frac{\sum_{c \in C_i}{\hat{x}_{if}\cdot\hat{y}_{jf}}}{k \cdot N} + (1-\alpha)\cdot\hat{r}_{ij}
# $$
# + id="6ax9icw4Ddb6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 136} outputId="50e84fb1-4570-440e-d26e-4d565d3ab348"
alpha = 0.9 # alpha value in range [0,1]
num_most_cared_aspects = 100
eval_metrics = [
cornac.metrics.NDCG(k=50),
cornac.metrics.AUC()
]
cornac.Experiment(
eval_method=rs,
models=[
EFM(
alpha=alpha,
num_most_cared_aspects=num_most_cared_aspects,
init_params={'U1': efm.U1, 'U2': efm.U2, 'H1': efm.H1, 'H2': efm.H2, 'V': efm.V},
trainable=False,
verbose=VERBOSE,
seed=SEED
)
],
metrics=eval_metrics
).run()
# + [markdown] id="oc7l9Mo8mHDN" colab_type="text"
# ### Recommendation Explanation with EFM
#
# Given a user and an item, EFM model is able of predicting **user's attention scores** as well as **item's quality scores** regarding the aspects. Those scores with the corresponding aspects will be the explanation on why a user *likes* or *dislikes* an item.
#
# Let's take a look at an example below. Feel free to explore other users and items!
# + id="Tat5X8bELfln" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 359} outputId="1e12d6f3-7232-4c20-c20f-8c9ab2812b45"
UIDX = 1
IIDX = 4
num_top_cared_aspects = 10
id_aspect_map = {v:k for k, v in rs.sentiment.aspect_id_map.items()}
predicted_user_aspect_scores = np.dot(efm.U1[UIDX], efm.V.T)
predicted_item_aspect_scores = np.dot(efm.U2[IIDX], efm.V.T)
top_cared_aspect_ids = (-predicted_user_aspect_scores).argsort()[:num_top_cared_aspects]
top_cared_aspects = [id_aspect_map[aid] for aid in top_cared_aspect_ids]
pd.DataFrame.from_dict({
"aspect": top_cared_aspects,
"user_aspect_attention_score": predicted_user_aspect_scores[top_cared_aspect_ids],
"item_aspect_quality_score": predicted_item_aspect_scores[top_cared_aspect_ids]
})
# + [markdown] id="v4H_VJIZIMiQ" colab_type="text"
# EFM takes an aspect with the **highest score** in `item_aspect_quality_score` as the well-performing aspect, and an aspect with the **lowest score** in `item_aspect_quality_score` as the poorly-performing aspect. See example explanations in their templates below.
# + id="m4gEW7XvA3wy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="e7e6034a-e869-4a3e-b4d4-75107bda250f"
perform_well_aspect = top_cared_aspects[predicted_item_aspect_scores[top_cared_aspect_ids].argmax()]
perform_poorly_aspect = top_cared_aspects[predicted_item_aspect_scores[top_cared_aspect_ids].argmin()]
explanation = \
f"You might interested in [{perform_well_aspect}], on which this product perform well. \n\
You might interested in [{perform_poorly_aspect}], on which this product perform poorly."
print("EFM explanation:")
print(explanation)
# + [markdown] id="X1vcUpSdnVWA" colab_type="text"
# ## 4. Multi-Task Explainable Recommendation (MTER)
#
# MTER model extends the concept of exploiting information from *Aspect-Level Sentiments* with tensor factorization (using Tucker Decomposition). The model takes in the input of three tensors. Let's go through each of them and see how they are constructed.
#
#
# + [markdown] id="zgLEBlP8JNPT" colab_type="text"
# ### Tensor\#1: User by Item by Aspect ($\mathbf{X}$)
#
# Let $\mathbf{R} \in \mathbb{R}^{N \times M}$ be a sparse rating matrix of $N$ users and $M$ items.
#
# Let $\mathbf{X} \in \mathbb{R}_{+}^{N \times M \times F}$ be a 3-dimensional tensor, each element $x_{ijf}$ indicates a relationship between user $i$, item $j$, and aspect $f$:
#
# \begin{equation}
# x_{ijf} = \
# \begin{cases}
# 0, & \text{if aspect $f$ has not been mentioned by user $i$ about item $j$} \\
# 1 + (N-1)\left(\frac{1}{1+\exp(-s_{ijf})}\right), & \text{otherwise}
# \end{cases}
# \end{equation}
#
# where $s_{ijf}$ is the sum of sentiment values with which item $j$ has been mentioned by user $i$ with regards to aspect $f$.
#
# We can extend $\mathbf{X}$ into $\mathbf{\tilde{X}}$ with the rating matrix $\mathbf{R}$ as the last slice or the $(F + 1)^{\mathrm{th}}$ aspect (i.e., $\tilde{x}_{ij(F+1)} = r_{ij}$).
# + [markdown] id="RzJuiT0TJZGV" colab_type="text"
# ### Tensor\#2: User by Aspect by Opinion ($\mathbf{Y}^{U}$)
#
# Let $\mathbf{Y}^{U} \in \mathbb{R}_{+}^{N \times F \times O}$ be a 3-dimensional tensor, each element $y^U_{ifo}$ indicates a relationship between user $i$, aspect $f$, and opinion $o$:
#
# \begin{equation}
# y^U_{ifo} = \
# \begin{cases}
# 0, & \text{if user $i$ has not been used opinion $o$ to describe aspect $f$ positively} \\
# 1 + (N-1)\left(\frac{1}{1+\exp(-t_{ifo})}\right), & \text{otherwise}
# \end{cases}
# \end{equation}
#
# where $t_{ifo}$ is the frequency with which user $i$ employs opinion $o$ to describe aspect $f$ positively across all her reviews.
#
# + [markdown] id="CLnMbMtTJZnr" colab_type="text"
# ### Tensor\#3: Item by Aspect by Opinion ($\mathbf{Y}^{I}$)
#
# Let $\mathbf{Y}^{I} \in \mathbb{R}_{+}^{M \times F \times O}$ be a 3-dimensional tensor, each element $y^I_{jfo}$ indicates a relationship between item $j$, aspect $f$, and opinion $o$:
#
# \begin{equation}
# y^I_{jfo} = \
# \begin{cases}
# 0, & \text{if item $j$ has not been described positively with opinion $o$ on aspect $f$} \\
# 1 + (N-1)\left(\frac{1}{1+\exp(-t_{jfo})}\right), & \text{otherwise}
# \end{cases}
# \end{equation}
#
# where $t_{jfo}$ is the frequency with which item $j$ has been described positively with opinion $o$ on aspect $f$ positively across all its reviews.
# + [markdown] id="ny5xlRKDJq8n" colab_type="text"
# ### Optimization
#
# MTER employs Tucker Decomposition to jointly factorize three tensors $\mathbf{\tilde{X}}$, $\mathbf{Y}^U$, and $\mathbf{Y}^I$. In addition, MTER also optimizes for a ranking objective akin to BPR where:
# * Positive triples $\mathbf{T} = \{ j >_{i} j' | x_{ij(F+1)} \in \mathbf{R}^+ \land x_{ij'(F+1)} \in \mathbf{R}^- \}$
# * For aspect (F + 1), which is the overall rating, user $i$ prefers item $j$ to item $j'$
#
# Learning the latent factors can be done via minimizing the following loss function:
#
# \begin{align}
# &\mathcal{L}(\mathbf{U, V, Z, W, C_1, C_2, C_3} | \lambda_B, \lambda) = ||\mathbf{\tilde{X}} - \mathbf{\hat{X}}||_F^2 + ||\mathbf{Y}^U - \hat{\mathbf{Y}}^U||_F^2 + ||\mathbf{Y}^I - \hat{\mathbf{Y}}^I||_F^2 - \lambda_B \sum_{j >_i j'} \ln(1 + \exp{(-(\hat{x}_{ij(F+1)} - \hat{x}_{ij'(F+1)}))}) + \lambda(||\mathbf{U}||_F^2+||\mathbf{V}||_F^2+||\mathbf{Z}||_F^2+||\mathbf{W}||_F^2 +||\mathbf{C_1}||_F^2 +||\mathbf{C_2}||_F^2 +||\mathbf{C_3}||_F^2) \\
# &\text{such that: } \mathbf{U} \ge 0, \mathbf{V} \ge 0, \mathbf{Z} \ge 0, \mathbf{W} \ge 0, \mathbf{C_1} \ge 0, \mathbf{C_2} \ge 0, \mathbf{C_3} \ge 0
# \end{align}
#
#
# The can be solved as a constrained optimization problem.
#
#
# Let's conduct an experiment with MTER model and compare with the BPR baseline.
# + id="BBsqIaj8nUyp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 153} outputId="41f625e1-9a87-4743-d4cf-ee7890bf2f29"
mter = MTER(
n_user_factors=10,
n_item_factors=10,
n_aspect_factors=10,
n_opinion_factors=10,
n_bpr_samples=1000,
n_element_samples=50,
lambda_reg=0.1,
lambda_bpr=10,
max_iter=3000,
lr=0.5,
verbose=VERBOSE,
seed=SEED,
)
# compare to baseline BPR
bpr = BPR(k=10, verbose=VERBOSE, seed=SEED)
eval_metrics = [
cornac.metrics.NDCG(k=50),
cornac.metrics.AUC()
]
# Instantiate and run an experiment
cornac.Experiment(
eval_method=rs, models=[bpr, mter], metrics=eval_metrics,
).run()
# + [markdown] id="KAe3fYPLFSPW" colab_type="text"
# ### Recommendation Explanation with MTER
#
# * To provide recommendation to user $i$, we rank items $j$ in terms of the predicted rating scores: $\hat{x}_{ij(F+1)}$
#
# * To determine which aspect $f$ of product $j$ a user $i$ cares about, we rank aspects $f$ in terms of: $\hat{x}_{ijf}$
#
# * To determine which opinion phrases $o$ to use when describing aspect $f$ while recommending item $j$ to user $i$, we rank phrases in terms of: $\hat{y}^U_{ifo} \times \hat{y}^I_{jfo}$
#
# Let's explore an example below on how we can generate explanations for recommendation by MTER model.
# + id="a5Tw_xf-FRaz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 111} outputId="d1043d57-5915-4b36-cd40-3db162e865c5"
UIDX = 10
IIDX = 10
num_top_aspects = 2
num_top_opinions = 3
item_aspect_ids = np.array(list(set([
tup[0]
for idx in rs.sentiment.item_sentiment[IIDX].values()
for tup in rs.sentiment.sentiment[idx]
])))
item_opinion_ids = np.array(list(set([
tup[1]
for idx in rs.sentiment.item_sentiment[IIDX].values()
for tup in rs.sentiment.sentiment[idx]
])))
item_aspects = [id_aspect_map[idx] for idx in item_aspect_ids]
ts1 = np.einsum("abc,a->bc", mter.G1, mter.U[UIDX])
ts2 = np.einsum("bc,b->c", ts1, mter.I[IIDX])
predicted_aspect_scores = np.einsum("c,Mc->M", ts2, mter.A)
top_aspect_ids = item_aspect_ids[(-predicted_aspect_scores[item_aspect_ids]).argsort()[:num_top_aspects]]
top_aspects = [id_aspect_map[idx] for idx in top_aspect_ids]
top_aspect_opinions = []
mter_explanations = []
for top_aspect_id, top_aspect in zip(top_aspect_ids, top_aspects):
ts1_G2 = np.einsum("abc,a->bc", mter.G2, mter.U[UIDX])
ts2_G2 = np.einsum("bc,b->c", ts1_G2, mter.A[top_aspect_id])
predicted_user_aspect_opinion_scores = np.einsum("c,Mc->M", ts2_G2, mter.O)
ts1_G3 = np.einsum("abc,a->bc", mter.G3, mter.I[IIDX])
ts2_G3 = np.einsum("bc,b->c", ts1_G3, mter.A[top_aspect_id])
predicted_item_aspect_opinion_scores = np.einsum("c,Mc->M", ts2_G3, mter.O)
predicted_aspect_opinion_scores = np.multiply(predicted_user_aspect_opinion_scores, predicted_item_aspect_opinion_scores)
top_opinion_ids = item_opinion_ids[(-predicted_aspect_opinion_scores[item_opinion_ids]).argsort()[:num_top_opinions]]
top_opinions = [id_opinion_map[idx] for idx in top_opinion_ids]
top_aspect_opinions.append(top_opinions)
# Generate explanation for top-1 aspect
mter_explanations.append(f"Its {top_aspect} is [{'] ['.join(top_opinions)}].")
pd.DataFrame.from_dict({"aspect": top_aspects, "top_opinions": top_aspect_opinions, "explanation": mter_explanations})
# + [markdown] id="b7t7OUvQ1IF1" colab_type="text"
# ## References
#
# 1. <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., & <NAME>. (2014). Explicit factor models for explainable recommendation based on phrase-level sentiment analysis. In SIGIR (pp. 83-92).
# 2. <NAME>., <NAME>., <NAME>., & <NAME>. (2018). Explainable recommendation via multi-task learning in opinionated text data. In SIGIR (pp. 165-174).
# 4. Cornac - A Comparative Framework for Multimodal Recommender Systems (https://cornac.preferred.ai/)
#
| _source/raw/preferredai_07_explanations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + tags=["remove_input"]
from datascience import *
path_data = '../../../../data/'
import numpy as np
import matplotlib
matplotlib.use('Agg', warn=False)
# %matplotlib inline
import matplotlib.pyplot as plots
plots.style.use('fivethirtyeight')
np.set_printoptions(legacy='1.13')
# + tags=["remove_input"]
def standard_units(any_numbers):
"Convert any array of numbers to standard units."
return (any_numbers - np.mean(any_numbers))/np.std(any_numbers)
def correlation(t, x, y):
return np.mean(standard_units(t.column(x))*standard_units(t.column(y)))
def slope(table, x, y):
r = correlation(table, x, y)
return r * np.std(table.column(y))/np.std(table.column(x))
def intercept(table, x, y):
a = slope(table, x, y)
return np.mean(table.column(y)) - a * np.mean(table.column(x))
def fit(table, x, y):
a = slope(table, x, y)
b = intercept(table, x, y)
return a * table.column(x) + b
def residual(table, x, y):
return table.column(y) - fit(table, x, y)
def scatter_fit(table, x, y):
plots.scatter(table.column(x), table.column(y), s=20)
plots.plot(table.column(x), fit(table, x, y), lw=2, color='gold')
plots.xlabel(x)
plots.ylabel(y)
# + tags=["remove_input"]
def draw_and_compare(true_slope, true_int, sample_size):
x = np.random.normal(50, 5, sample_size)
xlims = np.array([np.min(x), np.max(x)])
eps = np.random.normal(0, 6, sample_size)
y = (true_slope*x + true_int) + eps
tyche = Table().with_columns(
'x', x,
'y', y
)
plots.figure(figsize=(6, 16))
plots.subplot(4, 1, 1)
plots.scatter(tyche['x'], tyche['y'], s=20)
plots.plot(xlims, true_slope*xlims + true_int, lw=2, color='green')
plots.title('True Line, and Points Created')
plots.subplot(4, 1, 2)
plots.scatter(tyche['x'],tyche['y'], s=20)
plots.title('What We Get to See')
plots.subplot(4, 1, 3)
scatter_fit(tyche, 'x', 'y')
plots.xlabel("")
plots.ylabel("")
plots.title('Regression Line: Estimate of True Line')
plots.subplot(4, 1, 4)
scatter_fit(tyche, 'x', 'y')
plots.ylabel("")
xlims = np.array([np.min(tyche['x']), np.max(tyche['x'])])
plots.plot(xlims, true_slope*xlims + true_int, lw=2, color='green')
plots.title("Regression Line and True Line")
# + tags=["remove_input"]
baby = Table.read_table(path_data + 'baby.csv')
# -
# ### Inference for the True Slope ###
#
# Our simulations show that if the regression model holds and the sample size is large, then the regression line is likely to be close to the true line. This allows us to estimate the slope of the true line.
#
# We will use our familiar sample of mothers and their newborn babies to develop a method of estimating the slope of the true line. First, let's see if we believe that the regression model is an appropriate set of assumptions for describing the relation between birth weight and the number of gestational days.
scatter_fit(baby, 'Gestational Days', 'Birth Weight')
correlation(baby, 'Gestational Days', 'Birth Weight')
# By and large, the scatter looks fairly evenly distributed around the line, though there are some points that are scattered on the outskirts of the main cloud. The correlation is 0.4 and the regression line has a positive slope.
#
# Does this reflect the fact that the true line has a positive slope? To answer this question, let us see if we can estimate the true slope. We certainly have one estimate of it: the slope of our regression line. That's about 0.47 ounces per day.
slope(baby, 'Gestational Days', 'Birth Weight')
# But had the scatter plot come out differently, the regression line would have been different and might have had a different slope. How do we figure out how different the slope might have been?
#
# We need another sample of points, so that we can draw the regression line through the new scatter plot and find its slope. But from where will get another sample?
#
# You have guessed it – we will *bootstrap our original sample*. That will give us a bootstrapped scatter plot, through which we can draw a regression line.
# ### Bootstrapping the Scatter Plot ###
# We can simulate new samples by random sampling with replacement from the original sample, as many times as the original sample size. Each of these new samples will give us a scatter plot. We will call that a *bootstrapped scatter plot*, and for short, we will call the entire process *bootstrapping the scatter plot*.
#
# Here is the original scatter diagram from the sample, and four replications of the bootstrap resampling procedure. Notice how the resampled scatter plots are in general a little more sparse than the original. That is because some of the original points do not get selected in the samples.
# + tags=["remove_input"]
plots.figure(figsize=(8, 18))
plots.subplot(5, 1, 1)
plots.scatter(baby[1], baby[0], s=10)
plots.xlim([150, 400])
plots.title('Original sample')
for i in np.arange(1, 5, 1):
plots.subplot(5,1,i+1)
rep = baby.sample(with_replacement=True)
plots.scatter(rep[1], rep[0], s=10)
plots.xlim([150, 400])
plots.title('Bootstrap sample '+str(i))
# -
# ### Estimating the True Slope ###
#
# We can bootstrap the scatter plot a large number of times, and draw a regression line through each bootstrapped plot. Each of those lines has a slope. We can simply collect all the slopes and draw their empirical histogram. Recall that by default, the `sample` method draws at random with replacement, the same number of times as the number of rows in the table. That is, `sample` generates a bootstrap sample by default.
slopes = make_array()
for i in np.arange(5000):
bootstrap_sample = baby.sample()
bootstrap_slope = slope(bootstrap_sample, 'Gestational Days', 'Birth Weight')
slopes = np.append(slopes, bootstrap_slope)
Table().with_column('Bootstrap Slopes', slopes).hist(bins=20)
# We can then construct an approximate 95% confidence interval for the slope of the true line, using the bootstrap percentile method. The confidence interval extends from the 2.5th percentile to the 97.5th percentile of the 5000 bootstrapped slopes.
left = percentile(2.5, slopes)
right = percentile(97.5, slopes)
left, right
# An approximate 95% confidence interval for the true slope extends from about 0.38 ounces per day to about 0.56 ounces per day.
# ### A Function to Bootstrap the Slope ###
#
# Let us collect all the steps of our method of estimating the slope and define a function `bootstrap_slope` that carries them out. Its arguments are the name of the table and the labels of the predictor and response variables, and the desired number of bootstrap replications. In each replication, the function bootstraps the original scatter plot and calculates the slope of the resulting regression line. It then draws the histogram of all the generated slopes and prints the interval consisting of the "middle 95%" of the slopes.
def bootstrap_slope(table, x, y, repetitions):
# For each repetition:
# Bootstrap the scatter, get the slope of the regression line,
# augment the list of generated slopes
slopes = make_array()
for i in np.arange(repetitions):
bootstrap_sample = table.sample()
bootstrap_slope = slope(bootstrap_sample, x, y)
slopes = np.append(slopes, bootstrap_slope)
# Find the endpoints of the 95% confidence interval for the true slope
left = percentile(2.5, slopes)
right = percentile(97.5, slopes)
# Slope of the regression line from the original sample
observed_slope = slope(table, x, y)
# Display results
Table().with_column('Bootstrap Slopes', slopes).hist(bins=20)
plots.plot(make_array(left, right), make_array(0, 0), color='yellow', lw=8);
print('Slope of regression line:', observed_slope)
print('Approximate 95%-confidence interval for the true slope:')
print(left, right)
# When we call `bootstrap_slope` to find a confidence interval for the true slope when the response variable is birth weight and the predictor is gestational days, we get an interval very close to the one we obtained earlier: approximately 0.38 ounces per day to 0.56 ounces per day.
bootstrap_slope(baby, 'Gestational Days', 'Birth Weight', 5000)
# Now that we have a function that automates our process of estimating the slope of the true line in a regression model, we can use it on other variables as well.
#
# For example, let's examine the relation between birth weight and the mother's height. Do taller women tend to have heavier babies?
#
# The regression model seems reasonable, based on the scatter plot, but the correlation is not high. It's just about 0.2.
scatter_fit(baby, 'Maternal Height', 'Birth Weight')
correlation(baby, 'Maternal Height', 'Birth Weight')
# As before, we can use `bootstrap_slope` to estimate the slope of the true line in the regression model.
bootstrap_slope(baby, 'Maternal Height', 'Birth Weight', 5000)
# A 95% confidence interval for the true slope extends from about 1 ounce per inch to about 1.9 ounces per inch.
# ### Could the True Slope Be 0? ###
#
# Suppose we believe that our data follow the regression model, and we fit the regression line to estimate the true line. If the regression line isn't perfectly flat, as is almost invariably the case, we will be observing some linear association in the scatter plot.
#
# But what if that observation is spurious? In other words, what if the true line was flat – that is, there was no linear relation between the two variables – and the association that we observed was just due to randomness in generating the points that form our sample?
#
# Here is a simulation that illustrates why this question arises. We will once again call the function ``draw_and_compare``, this time requiring the true line to have slope 0. Our goal is to see whether our regression line shows a slope that is not 0.
#
# Remember that the arguments to the function ``draw_and_compare`` are the slope and the intercept of the true line, and the number of points to be generated.
draw_and_compare(0, 10, 25)
# Run the simulation a few times, keeping the slope of the true line 0 each time. You will notice that while the slope of the true line is 0, the slope of the regression line is typically not 0. The regression line sometimes slopes upwards, and sometimes downwards, each time giving us a false impression that the two variables are correlated.
# To decide whether or not the slope that we are seeing is real, we would like to test the following hypotheses:
#
# **Null Hypothesis.** The slope of the true line is 0.
#
# **Alternative Hypothesis.** The slope of the true line is not 0.
#
# We are well positioned to do this. Since we can construct a 95% confidence interval for the true slope, all we have to do is see whether the interval contains 0.
#
# If it doesn't, then we can reject the null hypothesis (with the 5% cutoff for the P-value).
#
# If the confidence interval for the true slope does contain 0, then we don't have enough evidence to reject the null hypothesis. Perhaps the slope that we are seeing is spurious.
# Let's use this method in an example. Suppose we try to estimate the birth weight of the baby based on the mother's age. Based on the sample, the slope of the regression line for estimating birth weight based on maternal age is positive, about 0.08 ounces per year.
slope(baby, 'Maternal Age', 'Birth Weight')
# Though the slope is positive, it's pretty small. The regression line is so close to flat that it raises the question of whether the true line is flat.
scatter_fit(baby, 'Maternal Age', 'Birth Weight')
# We can use `bootstrap_slope` to estimate the slope of the true line. The calculation shows that an approximate 95% bootstrap confidence interval for the true slope has a negative left end point and a positive right end point – in other words, the interval contains 0.
bootstrap_slope(baby, 'Maternal Age', 'Birth Weight', 5000)
# Because the interval contains 0, we cannot reject the null hypothesis that the slope of the true linear relation between maternal age and baby's birth weight is 0. Based on this analysis, it would be unwise to predict birth weight based on the regression model with maternal age as the predictor.
| interactivecontent/simulate-the-distribution-of-regression-coefficients/inference-for-the-true-slope.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ChristianPaul1127/CPEN-21A-ECE2-1/blob/main/Lab1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="CGdMpKBdqgR_"
# #Laboratory 1
# + colab={"base_uri": "https://localhost:8080/"} id="mP6C47y1qtRC" outputId="ea0b10f2-7fc7-4171-e10d-947e1c2c1059"
#Code for displaying “Welcome to Python Programming”
print("Welcome to Python Programming")
# + colab={"base_uri": "https://localhost:8080/"} id="EvflDNCxsIx_" outputId="85e9f608-42b2-4bd7-d31d-d6e87716e6e5"
x="Christian "
y="Paul "
z="Garcia"
Name=x+y+z
print("Name: "+Name)
A=int(19)
print("Age:", A, "years old")
Address="87 Don Placido Campos Avenue Dasmarinas City Cavite"
print("Address:",Address)
| Lab1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# https://reshamas.github.io/deploying-deep-learning-models-on-web-and-mobile/
# heroku account: hotmail, pw:D..homeps
# clone this repo:https://github.com/npatta01/web-deep-learning-classifier
#
#
# # on google cloud platform
# this is one app I got runnning, meant to do the bear detector, but maybe not working, trying to figure out
#
# http://beardetector-249718.appspot.com/
#
# you can open the google cloud console, see files in your home dir, /home/zszong73/
#
# the main.py file in /home/zszong73/gcpapp1, run the flask app? it renders index.html file? the index.html and other web page files are in the templates sub directories.
#
# update you deployment app: gcloud deployment-manager deployments update beardetector
# # heroku
# I got bear classifier model served on heroku on Nov 12, 2019, exciting moment. thanks to this link:
# https://reshamas.github.io/deploying-deep-learning-models-on-web-and-mobile/. It worked. But I still need to understand why and how. I basically used all the directory structure, scripts. I just replaced the model.pkl, classes.txt files with mine. and specified the fastai version i used in the requirements.txt file. New version fastai loads learner a little differently. load_learn(path=, file=) instead of fname=. the pickle file contianing all the model weights needs to be consitent with the file name you used in the app.py script when you load the model.
#
# need to understand the webpage component and how it rendered and how the url or the upload image is passed to the model for prediction.
# when deploy models to production, make sure you use the same version of libraries as you trained your model. if the model.pkl if too big, you can save to git release, or save to google drive and copy to the local repo, need to push it to the heroku master branch in order to serve the web app.
# * build dock file,
# * push app to github
# * log in to herocu cli: heroku login -i, enter email and password, <EMAIL>, D..S..20
# * log in to container: heroku container:login
# * cd to the project directory,
# * heroku create --app stuart-bear-classifier # this create an app, add heroku remote to the git repo
# * heroku container:push web --app stuart-bear-classifier # sometimes this command needs to be rerun.
# * heroku container:release web --app stuart-bear-classifier # optional add " --app appname"
# * heroku open # this will open the app in a web browser
#
# # update after deloyment
# * make the change in local repo
# * commit changes to heroku, git add then commit, then git push heroku master
# * repush and release the app using heroku container:push/release web --app appname
# * heroku open to launch the app in web browser.
# * added a heroku.yml file
# * git add/commit heroku.yml
# * heroku stack:set container # not sure what this does
# * git push heroku master
| nbs/dl1/delopy_deep_learning_model_on_web_20191108.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Homework 5: Mini Project
#
# <br>
#
# **CS 412 Introduction to Machine Learning, Fall 2018**
#
# **University of Illinois at Chicago**
#
# <br>
#
# *Due December 7, 2018, 11:59 PM*
# <hr>
#
# ## Selected option
#
# **Option 1:**
#
# You are working for a non-profit that is recruiting student volunteers to help with Alzheimer's patients. You have been tasked with predicting how suitable a person is for this task by predicting how empathetic he or she is. Using the Young People Survey dataset (https://www.kaggle.com/miroslavsabo/young-people-survey/), predict a person's "Empathy" as either "very empathetic" (answers 4 and 5) or "not very empathetic" (answers 1, 2, and 3). You can use any of the other attributes in the dataset to make this prediction; however, you should not handpick the predictive features but let an algorithm select them.
# <hr>
# <br>
#
# ## Step 1: Import required Python files
from main import main
# ## Step 2: Train models
#
# You may skip this step if you have a testing set file, and a pre-trained model dump file generated by the program with you.
# +
# TRAIN MODE
# ==========
mode = 'train'
dataset = 'data/responses.csv'
modelDumpFile = 'data/bestModel.pkl'
main(mode, dataset, modelDumpFile)
# -
# ## Step 3: Test models
#
# **Note:** In order to run the program in `test` mode, you will need to have a CSV file with the testing data present, along with a dump of the trained models generated from this program previously. If you don't have these files, simply run the program in `train` mode first.
# +
# TEST MODE
# =========
mode = 'test'
dataset = 'data/testSet.csv'
modelDumpFile = 'data/bestModel.pkl'
main(mode, dataset, modelDumpFile)
| hw5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Kontrollstrukturen
#
# ## Codeblöcke
#
# In Python werden Blöcke von Anweisungen durch Einrücken gebildet und nicht wie beispielsweise in Java oder JavaScript mit geschweiften Klammern.. Die letzte Codezeile vor dem Block endet mit einem Doppelpunkt `:`. Auf diese Weise erzwingt Python, dass ein Block auch visuell klar zum Ausdruck kommt.
#
# Das Einrücken kann Leerzeichen oder dem Tabulator erzeugt werden. Wichtig ist, dass innerhalb eines Blocks immer das gleiche Zeichen verwendet wird. Eine gute Konvention ist, dass innerhalb des Programms und über die Programme hinaus immer der gleiche Stil verwendet wird. Typisch sind *vier Leerzeichen* für das Einrücken.
# ## Kommentare
#
# Ein Kommentar (einzeilig) wird mit der Raute `#` eingeleitet. Dieses Zeichen kann an einer beliebigen Stelle auf einer Zeile stehen, üblicherweise hinter dem Code der betreffenden Zeile.
#
# Ein mehrzeiliger Kommentar (auch *docstring* genannt), wird durch drei `"""` eröffnet und abgeschlossen. Diese Form sollte vor allem zu Dokumentation von Funktionen und Klassen verwendet werden.
# ## Verzweigungen
#
# Für bedingte Verzweigungen gibt es in Python `if condition: ... elif condition2: ... else: ...`.
temperature = 15
if temperature <= 10:
print("kalt")
elif temperature > 10 and temperature <= 25:
print("angenehm")
else:
print("heiss")
# Übungen zu *if-else*: https://www.w3schools.com/python/exercise.asp?filename=exercise_ifelse1
# Anspruchsvolle Übungen zu *if-else*: https://realpython.com/quizzes/python-conditional-statements/viewer/
# ## Schleifen
#
# Es gibt bedingte Schleifen (`while`) und Iterationen (`for`).
#
# Mit `while` wird ein Block ausgeführt, bis eine Bedinung nicht mehr erfüllt ist:
i = 4
while (i < 9):
i = i+2
print(i)
# Übungen zu *while*-Schleifen: https://www.w3schools.com/python/exercise.asp?filename=exercise_while_loops1
#
# Mit *for*-Schleifen können wir über Listen iterieren.
# Beipiel: finde alle Primzahlen:
for n in range(2, 20):
for x in range(2, n):
if n % x == 0:
print(f"{n} ist gleich {x}*{n//x}")
break
else:
print(f"{n} ist einer Primzahl")
# *Hinweis 1*: in diesem Beispiel werden zwei Schleifen verschachtelt.
#
# *Hinweis 2*: dieses wie auch die anderen Beispiele zeigen, dass mit *f-String* gut lesbarer Code erzeugt werden kann. Wenn in den geschweiften Klammer Operationen ausgeführt werden, hat das allerdings den Nachteil, dass eine Fehlersuche schwierig werden kann.
# **Schlüsselwörter** `break` und `continue`.
#
# Mit `break` wird die innere Schleife verlassen, wenn festgestellt wird, dass die aktuell getestete Zahl keine Primzahl ist.
#
# Das Schlüsselwort `continue` bewirkt, dass die restlichen Anweisungen im Block nicht ausgeübt werden. Stattdessen wird die nächste Iteration aufgerufen.
for i in range(10):
print(f"Durchlauf {i}...")
continue
print("Das wird nicht ausgegeben!")
# Übungen zu *for*-Schleifen: https://www.w3schools.com/python/exercise.asp?filename=exercise_for_loops1
# ## Funktionen
#
# Eine Funktion in Python hat folgendes Aussehen: `def func_name(parameters): ... return value`.
#
# *Beispiel*: Währungskonverter:
# +
def euro_to_chf(euro_amount):
return euro_amount * 1.08015
print(f"50 Euro sind Fr. {euro_to_chf(50)}.")
# -
# Es gibt verschiedene Arten von Parametern und Argumenten bei Python-Funktionen:
# - optionales Argument: ein Parameter wird mit einem Standardwert ausgestattet
# - zwingendes Argument: ein Parameter ohne Standartwert
# - positionales Argument: Zuordnung des Übergabewerts (Arguments) zum Parameter auf Grund der Position
# - Schlüsselwort-Argument: Zuordnung des Übergabewerts zum Parameter auf Grund des Namens
# - multiple Argumente: einem Parameter werden eine Liste von Argumenten übergeben
# - multiple Schlüsselwort-Argumente: einem Parameter wird ein Argumente-Dictionary übergeben
# +
def concat_names(first='', middle='', last=''):
if middle.strip():
middle = middle[0] + '. '
print(f"{first} {middle}{last}")
# alle Argumente sind optional
print("vor Aufruf")
concat_names()
print("nach Aufruf")
# positional
concat_names("Adam", "Maria", "Riese")
# besser mit Schlüsselwort-Argumenten
concat_names(first="Adam", last="Riese")
concat_names(last="Riese", first="Adam")
concat_names(last="Riese", first="Adam", middle="Maria")
# -
# Beispiel für beliebig viele Argumente:
# +
def arithmetic_mean(x, *other):
sum = x
for i in other:
sum += i
return sum / (1.0 + len(other))
print(f"Mittelwert von [4, 5] ist {arithmetic_mean(4, 5)}!")
print(f"Mittelwert von [4, 9, 3, 33, 20] ist {arithmetic_mean(4, 9, 3, 33, 20)}!")
# -
# Beispiel für beliebig viele Schlüsselwort-Argumente:
# +
def kv_ex(**args):
for k,v in args.items():
print(f"{k}={v}")
print("1 =============")
kv_ex()
print("2 =============")
kv_ex(de="German",en="English",fr="French")
print("3 =============")
# -
# Übungen zu Funktionen: https://www.w3schools.com/python/exercise.asp?filename=exercise_functions1
#
#
# ## Fehlerbehandlung
#
# Mit einem `try: ... except: ...`-Block können Fehler in Python sauber abgefangen werden.
# Der `finally`-Block wird in jedem Fall durchlaufen. Das kann hilfreich sein, um einen Zustand aufzuräumen.
# +
def temp_convert(var):
try:
return int(var)
except ValueError as Argument:
print("Das Argument ist keine Zahl: ", Argument)
finally:
print("...")
temp_convert("xyz")
temp_convert(123)
# -
# ## *with*-Statement
#
# Bei einer Interaktion über die Systemgrenzen hinaus kann vieles schief gehen. Innerhalb des Systems können wir die Strukturen kontrollieren, ausserhalb des Systems ist das meistens nicht möglich. Ein typischer Fall ist die Interaktion mit dem Filesystem (Lesen/Schreiben von Files): Ist das File vorhanden? Ist der Inhalt kompatibel? Hat es Platz zum Schreiben?
#
# In solchen Fällen ist das `with`-Statement hilfreich. Es sorgt dafür, dass beim Verlassen des Blocks sauber aufgeräumt wird, z.B. dass ein offener File-Zugriff automatisch geschlossen wird. Dies erübrigt einen `finally`-Block und macht den Code besser lesbar.
# +
# Fall mit Fehlerbehandlung
file_name = "non_existing.txt"
try:
with open(file_name) as file:
print(f"Anzahl Zeichen in '{file_name}': {len(file.read())}")
except FileNotFoundError:
print(f"Fehler: das File '{file_name}' kann nicht gefunden werden!")
# korrekter Fall
file_name = "unicode.txt"
try:
with open(file_name) as file:
print(f"Anzahl Zeichen in '{file_name}': {len(file.read())}")
except FileNotFoundError:
print(f"Fehler: das File '{file_name}' kann nicht gefunden werden!")
# -
# ## *raise*-Statemen
#
# Mit *raise* kann eine Fehlerbehandlung ausgelöst werden:
# +
x = -1
if x < 0:
raise Exception("Sorry, keine negativen Zahlen erlaubt!")
# -
# ## Übungen: Kontrollstrukturen
#
# Fibonacci-Reihe:
# 1. Berechne die 100. Fibnoacci-Zahl und teile diese durch die vorhergehende.
# 1. Berechne alle Fibnoacci-Zahlen bis 50 und schreibe für jeden Schritt den Wert *fibo(n)/fibo(n-1)* hinaus.
| ws1/Kontrollstrukturen.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:autobfe]
# language: python
# name: conda-env-autobfe-py
# ---
# +
# %load_ext autoreload
# %autoreload 2
from matplotlib.path import Path
import numpy as np
import matplotlib.pyplot as plt
import shapely
import cv2
from PIL import Image
import argparse
import os, sys
sys.path.append(os.path.dirname(os.getcwd()))
import polygon_primitives.file_writer as fw
from image_processing import extract_window_wall_ratio, utils, contour_extraction
from scipy.spatial import Delaunay
from common.colors import continuous_palette_for_color
# -
# %pwd
# First, we set the image and parameter directories, as well as the merged polygons file path. We load the merged polygons, as we also initialize a dictionary for the Cameras. The Camera class stores all information related to the camera, i.e. intrinsic and extrinsic camera parameters.
# +
#Example file
mapped_files = ["DJI_0047.JPG", "DJI_0026.JPG", "DJI_0055.JPG", "DJI_0131.JPG", "DJI_0012.JPG"]
# filename = "DJI_0026.JPG" ## good
filename = "DJI_0047.JPG" ## good
# filename = "DJI_0027.JPG"
directory = "/Users/akprakash/Programming/autobfe/blume/"
facade_file = directory+ "blume_merged_polygons.txt"
image_dir = directory + "rgb/"
param_dir = directory + "params/"
predictions_dir = directory + "probs/"
predictions_dir2 = directory + "probs2/"
offset = np.loadtxt(directory + "blume_polygon_offset.txt",usecols=range(3), delimiter=',')
#Initializes a dictionary of Camera classes. See utils.py for more information.
camera_dict = utils.create_camera_dict(param_dir, filename='merged_blume2_calibrated_camera_parameters.txt', offset=offset)
#Loads pmatrices and image filenamees
p_matrices = np.loadtxt(param_dir + 'merged_blume2_pmatrix.txt', usecols=range(1,13))
#Loads the merged polygons, as well as a list of facade types (i.e. roof, wall, or floor)
merged_polygons, facade_type_list, file_format = fw.load_merged_polygon_facades(filename=facade_file)
#Offset adjustment parameter
height_adj = np.array([0.0, 0.0, 108])
offset = offset + height_adj
# -
# Next, we extract the contours for the window predictions, by taking the window prediction points and using them to create a shapely polygon.
# +
image_file = image_dir + filename
image = cv2.imread(image_file)
plt.imshow(image)
plt.show()
# +
window_file = predictions_dir + filename.split(".")[0] +".png"
print("Window predictions: ")
pred_image = cv2.imread(window_file)
plt.imshow(pred_image)
plt.show()
print(pred_image.shape)
# -
new_pred_image = []
for x in pred_image:
new_y = []
for y in x:
if not np.array_equal(y, np.array([84, 1, 68])):
new_y.append(np.array([89, 0, 237]))
else:
new_y.append(np.array([236, 237, 237]))
new_pred_image.append(np.array(new_y))
# +
pred_image2 = np.array(new_pred_image, dtype='uint8')
from PIL import Image
im = Image.fromarray(pred_image2)
plt.imshow(im)
im.save(predictions_dir2+filename.split(".")[0] +".png")
# +
window_file = predictions_dir2 + filename.split(".")[0] +".png"
print("Window predictions: ")
image = cv2.imread(window_file)
plt.imshow(image)
plt.show()
#Extract the contours of the window file
contours = contour_extraction.extract_contours(window_file)
#Create polygons from the window contours
window_polygons = utils.convert_polygons_shapely(contours)
def plot_shapely_polys(image_file, polys):
for poly in polys:
s = poly
s = poly.simplify(0.1, preserve_topology=True)
x,y = s.exterior.xy
plt.plot(x,y)
plt.show()
print("Extracted contours: ")
plt.imshow(image)
plot_shapely_polys(window_file, window_polygons)
# -
# Finally, for each window point, we obtain its 3D coordinates and use them to calculate the window to wall ratio.
# +
camera = camera_dict[filename]
pmatrix = camera.calc_pmatrix()
image_file = utils.load_image(image_dir + filename)
#Projects the merged polygon facades onto the camera image
projected_facades, projective_distances = extract_window_wall_ratio.project_merged_polygons(
merged_polygons, offset, pmatrix)
#Creates a dictionary mapping the facade to the windows contained within them, keyed by facade index
facade_window_map = extract_window_wall_ratio.get_facade_window_map(
window_polygons, projected_facades, projective_distances)
#Creates a list of all the facades in the merged polygon
facades = []
for poly in merged_polygons:
facades = facades + poly
facade_indices = list(facade_window_map.keys())
for i in facade_indices:
#Computes window to wall ratio
win_wall_ratio = extract_window_wall_ratio.get_window_wall_ratio(
projected_facades[i], facades[i], facade_window_map[i])
#Output printing:
print("Facade index: " + str(i))
print("Window-to-wall ratio: " + str(win_wall_ratio))
#Uncomment this line to plot the windows and facades on the image
extract_window_wall_ratio.plot_windows_facade(projected_facades[i], facade_window_map[i], image_file)
# -
window_total_area = 0
for window in window_polygons:
window_total_area+=window.area
facade_total_area = 0
for facade in projected_facades:
facade_total_area+=facade.area
print(window_total_area)
print(facade_total_area)
print("total wwr = {}".format(window_total_area/facade_total_area))
window_polygons
projected_facades
for facade in projected_facades:
print(shapely.geometry.mapping(facade))
# print(facade.area)
# +
# shapely.geometry.mapping(window)
# -
for facade_idx in facade_window_map:
facade = projected_facades[facade_idx]
windows = facade_window_map.get(facade_idx)
facade_window_area = 0
for window in windows:
facade_window_area+=window.area
print(facade.area, facade_window_area, facade_window_area/facade.area)
# +
for facade in projected_facades:
# plt.figure()
x,y = facade.exterior.xy
plt.plot(x,y)
# plt.plot(projected_facades[0].exterior.xy)
# -
facade_coordinates = []
for facade in projected_facades:
facade_coordinates.append(shapely.geometry.mapping(facade).get('coordinates')[0][:-1])
print(shapely.geometry.mapping(facade).get('coordinates')[0][:-1])
print(facade_coordinates)
len(facade_coordinates)
projected_facades
for polygon in facade_coordinates:
tup = coordinate[0]
found = False
for tup in polygon:
print("searching for "+str(tup))
connected_polys = []
for polygon2 in facade_coordinates:
tup2 = polygon2[0]
if polygon != polygon2 and tup == tup2:
print(polygon, polygon2)
connected_polys.append(polygon2)
print()
| Example_Notebooks/window-blume.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## POS tagging using modified Viterbi
# **Problem statement:**
# <n>Modify the Viterbi algorithm to solve the problem of unknown words using at least two techniques
#
# **Goals:**
# 1. **Write** the **vanilla Viterbi** algorithm for assigning POS tags.
# 2. **Solve** the problem of unknown words **using** at least **two techniques.**
# 3. **Compare** the tagging **accuracy** after making these modifications with the vanilla Viterbi algorithm.
# 4. **List down** at least **3 cases** from the sample test file which were **incorrectly tagged** by the original Viterbi.
# ### Data Preparation
# +
#Importing libraries
import nltk, pprint, re
from nltk.tokenize import word_tokenize
from sklearn.model_selection import train_test_split
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import time
import random
import warnings
warnings.filterwarnings('ignore')
# -
# reading the Treebank tagged sentences
nltk_data = list(nltk.corpus.treebank.tagged_sents(tagset='universal'))
# samples: Each sentence is a list of (word, pos) tuples
nltk_data[:3]
#Test-Train split test - 5%, train - 95% as given in problem statement
train_set, test_set = train_test_split(nltk_data,test_size=0.05,random_state=100)
print(len(train_set))
print(len(test_set))
# converting the list of sents to a list of (word, pos tag) tuples in train_set
tagged_words_train = [w for s in train_set for w in s]
print(len(tagged_words_train))
tagged_words_train[:10]
# converting the list of sents to a list of (word, pos tag) tuples in train_set
tagged_words_test = [w for s in test_set for w in s]
print(len(tagged_words_test))
tagged_words_test[:10]
#separating Words and Tags
words = [pair[0] for pair in tagged_words_train]
w = set(words)
tags = [pair[1] for pair in tagged_words_train]
t = set(tags)
print(len(w))
print(len(t))
print(t)
# ### Build the vanilla Viterbi based POS tagger
# #### Emission Probabilities
#the probability of the word for a given tag
# we will calculate number of times the word occuring, #tags occuring and # word given the tag occuring
def Emission(word, tag, train_bag = tagged_words_train):
words = [pair for pair in train_bag if pair[0]==word]
count_word = len(words)
tags = [pair for pair in train_bag if pair[1]==tag]
count_tag = len(tags)
words_given_tag = [pair[0] for pair in tags if pair[0]==word]
count_words_given_tag = len(words_given_tag)
(w,t,w_t) = (count_word, count_tag,count_words_given_tag)
print("count of " + str(word) + " = " + str(w))
print("count of " + str(tag) + " = " + str(t))
print("number of times " + str(word) + " is tagged as " +str(tag)+ " = " + str(w_t))
return(w,t,w_t)
def emission(word, tag, train_bag = tagged_words_train):
tags = [pair for pair in train_bag if pair[1]==tag]
count_tag = len(tags)
words_given_tag = [pair[0] for pair in tags if pair[0]==word]
count_words_given_tag = len(words_given_tag)
return(count_tag, count_words_given_tag)
Emission('years', 'NOUN')
# #### Transition Probabilities
#the probability of tag1 followed by tag2
# we will calculate number of times tag1 occuring tag2 occuring and tag1 followed by tag2
def Transition(t1, t2, train_bag = tagged_words_train):
tag1 = [pair[1] for pair in train_bag]
count_t1 = len([t for t in tags if t==t1])
tag2 = [pair[1] for pair in train_bag]
count_t2 = len([t for t in tags if t==t2])
count_t2_t1 = 0
for index in range(len(tag1)-1):
if tag1[index]==t1 and tag1[index+1] == t2:
count_t2_t1 += 1
(x,y,t1_t2) = (count_t1,count_t2,count_t2_t1)
print("count of " + str(t1) + " = " + str(x)),
print("count of " + str(t2) + " = " + str(y)),
print("number of times " + str(t1) + " is followed by " +str(t2)+ " = " + str(t1_t2))
return (x,y,t1_t2)
def transition(t1, t2, train_bag = tagged_words_train):
tag1 = [pair[1] for pair in train_bag]
count_t1 = len([t for t in tags if t==t1])
count_t2_t1 = 0
for index in range(len(tag1)-1):
if tag1[index]==t1 and tag1[index+1] == t2:
count_t2_t1 += 1
return(count_t1,count_t2_t1)
Transition("ADJ",'NOUN')
Transition('DET','.')
#Creating a matrix for all the transitions possible
tags_matrix = np.zeros((len(t), len(t)), dtype='float32')
for i, t1 in enumerate(list(t)):
for j, t2 in enumerate(list(t)):
tags_matrix[i, j] = transition (t1, t2)[1]/transition (t1, t2)[0]
#represting in dataframe form
tags_df = pd.DataFrame(tags_matrix, columns = list(t), index=list(t))
tags_df
#heatmap
plt.figure(figsize=(18, 12))
sns.heatmap(tags_df)
plt.show()
# frequent tags
# filter the df to get P(t2, t1) > 0.5
tags_frequent = tags_df[tags_df>0.5]
plt.figure(figsize=(18, 12))
sns.heatmap(tags_frequent)
plt.show()
# now viterbi algorithm
# Viterbi Heuristic
def vanillaviterbi(words, train_bag = tagged_words_train):
state = []
T = list(set([pair[1] for pair in train_bag]))
for key, word in enumerate(words):
#initialise list of probability column for a given observation
p = []
for tag in T:
if key == 0:
transition_p = tags_df.loc['.', tag]
else:
transition_p = tags_df.loc[state[-1], tag]
# compute emission and state probabilities
emission_p = emission(words[key], tag)[1]/emission(words[key], tag)[0]
state_probability = emission_p * transition_p
p.append(state_probability)
pmax = max(p)
# getting state for which probability is maximum
state_max = T[p.index(pmax)]
state.append(state_max)
return list(zip(words, state))
Testword = [word[0] for sentence in test_set for word in sentence]
start = time.time()
tagged_seq = vanillaviterbi(Testword)
end = time.time()
difference = end-start
# accuracy
check = [i for i, j in zip(tagged_seq, tagged_words_test) if i == j]
accuracy = len(check)/len(tagged_seq)
print(accuracy*100)
print(difference)
incorrect_tagged_cases = [[tagged_words_test[i-1],j] for i, j in enumerate(zip(tagged_seq, tagged_words_test)) if j[0]!=j[1]]
incorrect_tagged_cases
# ### Solve the problem of unknown words
# #### Modification 1
# __** problem statement: Which tag class do you think most unknown words belong to? Can you identify rules (e.g. based on morphological cues) that can be used to tag unknown words? **__
# so lets look at most commonly occuring tag and use it for the unknown words
def tagcount(t1, train_bag = tagged_words_train):
tag = [pair[1] for pair in train_bag]
count_t1 = len([t for t in tag if t==t1])
return count_t1
tagcount("NOUN")
x = list(t)
temp = [tagcount(i) for i in x]
temp
temp_df = pd.DataFrame(temp, columns = ["count"],index=[list(t)])
temp_df
# **Noun** is the most commonly occuring tag so lets add a function to tag all the unkown words as nouns
def viterbi_modf1(x, train_bag = tagged_words_train):
tags_seq = vanillaviterbi(x, train_bag)
a = [pair[0] for pair in tags_seq]
b = [pair[1] for pair in tags_seq]
for key, word in enumerate(a):
if word not in w:
b[key] = 'NOUN'
return list(zip(a, b))
start = time.time()
seq2 = viterbi_modf1(Testword)
end = time.time()
difference = end-start
seq2
# accuracy
check = [i for i, j in zip(seq2, tagged_words_test) if i == j]
accuracy = len(check)/len(seq2)
print(accuracy*100)
print(difference)
incorrect_tagged_cases = [[tagged_words_test[i-1],j] for i, j in enumerate(zip(seq2, tagged_words_test)) if j[0]!=j[1]]
incorrect_tagged_cases
# #### Modification 2
# The idea for modificaation 2 is we will define a set of rules for unknown words using regex tagger
# and combine it with unigram and bigram taggers
# #### Rulebased or Regex tagger
# specify patterns for tagging
# example from the NLTK book
patterns = [
(r'.*(ed|ing|es)$', 'VERB'), # any word ending with ed,es and ing is tagged as verb
(r'.*ly$', 'ADV'), # Adverb ending with 'ly'.
(r'^([0-9]|[aA-zZ])+\-[aA-zZ]*$','ADJ'), # Alpha Numeric - ADJ
(r'.*able$', 'ADJ'), # Adjective ending with 'able'.
(r'.*ful$', 'ADJ'), # Adjective ending with 'ful'.
(r'.*ous$', 'ADJ'), # Adjective ending with 'ous'.
(r'^[aA-zZ].*[0-9]+','NOUN'), # Alpha Numeric.
(r'.*ness$', 'NOUN'), # Nouns - words ending with 'ness'.
(r'.*\'s$', 'NOUN'), # possessive nouns - words ending with 's
(r'.*s$', 'NOUN'), # plural nouns.
(r'.*ers$', 'NOUN'), # Nouns ending with 'ers'.
(r'.*ment$', 'NOUN'), # Nouns - words ending with 'ment'.
(r'.*town$', 'NOUN'), # Nouns - words ending with 'town'.
(r'^(0|([*|-|$].*))','X'), # Any special character combination
(r'.*ould$', 'X'),
(r'(The|the|A|a|An|an|That|that|This|this|Those|those|These|these)$', 'DET'), # That/this/these/those belong to the category of Demonstrative determiners
(r'[0-9].?[,\/]?[0-9]*','NUM'), # Numbers
(r'.*', 'NOUN') # nouns
]
regexp_tagger = nltk.RegexpTagger(patterns)
# #### Unigram tagger
# Lexicon (or unigram tagger) backedoff by regex tagger
unigram_tagger = nltk.UnigramTagger(train_set,backoff = regexp_tagger)
unigram_tagger.evaluate(test_set)
# #### Bigram tagger
# bigram tagger backed up by both unigram and regex tagger as mentioned above
def POS_tagger(word, words_set = train_set):
bigram_tagger = nltk.BigramTagger(train_set, backoff = unigram_tagger)
return bigram_tagger.tag_sents([[(word)]])
POS_tagger('choked')
# #### Veterbi modified by the POS_tagger
def viterbi_ver2(x, train_bag = tagged_words_train):
tags_seq = vanillaviterbi(x, train_bag)
a = [pair[0] for pair in tags_seq]
b = [pair[1] for pair in tags_seq]
for key, word in enumerate(a):
if word not in w:
unknown_word_tag = POS_tagger(word)
for sent in unknown_word_tag:
for tup in sent:
b[key] = tup[1]
return list(zip(a, b))
def viterbi_modf2(sample, train_bag = tagged_words_train):
state = []
T = list(set([pair[1] for pair in train_bag]))
for key, word in enumerate(sample):
if word not in w:
unknown_word_tag = POS_tagger(word)
for sent in unknown_word_tag:
for tup in sent:
state.append(tup[1])
else:
p = []
for tag in T:
if key == 0:
transition_p = tags_df.loc['.', tag]
else:
transition_p = tags_df.loc[state[-1], tag]
# compute emission and state probabilities
emission_p = emission(words[key], tag)[1]/emission(words[key], tag)[0]
state_probability = emission_p * transition_p
p.append(state_probability)
pmax = max(p)
# getting state for which probability is maximum
state_max = T[p.index(pmax)]
state.append(state_max)
return list(zip(words, state))
start = time.time()
seq3 = viterbi_ver2(Testword)
end = time.time()
difference = end-start
seq3
# accuracy
check = [i for i, j in zip(seq3, tagged_words_test) if i == j]
accuracy = len(check)/len(seq3)
print(accuracy*100)
print(difference)
# ### List down cases which were incorrectly tagged by vanilaviterbi and got corrected by modifications
def comparing(x,y,wordsTag) :
return [(original,venila,updated) for original,venila,updated in zip(wordsTag,x,y) if (venila != original) & (updated == original) ]
# compared to vanila
comparing_df = pd.DataFrame(set(comparing(tagged_seq,seq3,tagged_words_test)),columns=['Actual Tag','Vanilla','Modified Viterbi'])
comparing_df
# ### original tags that which donot match with modified vitrebi
def mismatch(x,wordsTag) :
return [(original,updated) for original,updated in zip(wordsTag,x) if (updated != original) ]
mismatch_df = pd.DataFrame(set(mismatch(seq3,tagged_words_test)),columns=['Actual Tag','Modified Viterbi'])
mismatch_df
# So there are **196** mismatches, but if you observe in the original tags itself there are some wrong tags, such as, "the" is tagged as "NUM" which is not correct
# ### Accuary comparision
#
#
# ||<NAME>|Modified Viterbi|Viterbi with POS_tagger|
# | --- | --- | --- | --- |
# |Accuracy|90.35%|93.5%|95.17%|
#
# ## Testing on the sample test sentences
# +
sample_test_sent = 'Android is a mobile operating system developed by Google. Android has been the best-selling OS worldwide on smartphones since 2011 and on tablets since 2013. Google and Twitter made a deal in 2015 that gave Google access to Twitter\'s firehose. Twitter is an online news and social networking service on which users post and interact with messages known as tweets. Before entering politics, <NAME> was a domineering businessman and a television personality. The 2018 FIFA World Cup is the 21st FIFA World Cup, an international football tournament contested once every four years. This is the first World Cup to be held in Eastern Europe and the 11th time that it has been held in Europe. Show me the cheapest round trips from Dallas to Atlanta. I would like to see flights from Denver to Philadelphia. Show me the price of the flights leaving Atlanta at about 3 in the afternoon and arriving in San Francisco. NASA invited social media users to experience the launch of ICESAT-2 Satellite.'
test_words = word_tokenize(sample_test_sent)
start = time.time()
tagged_seq = viterbi_ver2(test_words)
end = time.time()
difference = end-start
# -
tagged_seq
| POS_tagging.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
fn = "../input/forbes_celebrity_list.csv"
df = pd.read_csv(fn)
# ?pd.read_csv
df # this is a Pandas dataframe
# Note that every row has an index & every column has a name
df.head()
df.head(10)
df.tail(10)
len(df)
df.columns
df["recipient"] # this is a Pandas series (every value has an index)
df["recipient"].head(10) # this is the same procedure as for the dataframe earlier
df[["recipient", "career", "rank"]].head()
df.loc[4]
df.loc[4, "recipient"]
df["career"].value_counts() # this is sorted from highest to lowest count
df["country"].value_counts()
df[df["country"] == "Philippines"]
df[(df["rank"] <= 5) & (df["year"] == 2015)] # In Python, AND is '&' and OR is '/'
recipients = pd.unique(df["recipient"])
len(recipients)
recipients
df.sort_values(by="recipient") # this is arranged alphabetically
df.sort_values(by="recipient", ascending=False)
df.sort_values(by=["rank", "year"], ascending=[True, False])
df.loc[df["career"] == "Sportsperson (Boxing)", "recipient"]
# this returns a Pandas Series
df.loc[df["career"] == "Sportsperson (Boxing)", ["recipient", "year", "rank"]]
# this returns another Pandas Dataframe
df["recipient"].value_counts()
df[df["recipient"] == "<NAME>"]
df_top = df.loc[(df["recipient"] == "<NAME>") & (df["rank"] == 1)]
df_top
len(df_top)
# Bonus exercise: get the top 5 per year arranged by year and rank
df[df["rank"] <= 5].sort_values(by=["year", "rank"], ascending=[False, True])
df[df["country"].str.contains("united", case = False)]
df[(df["rank"] >= 6) & (df["year"] % 2 != 0)]
| session-4/scripts/2_Explore_Forbes_Celebrity_List.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PySpark
# language: ''
# name: pysparkkernel
# ---
# ## Architecture of a Spark Application
#
# ### Big picture
#
# You will type your commands iin a local Spark session, and the SparkContext will take care of running your instructions distributed across the workers (executors) on a cluster. Each executor can have 1 or more CPU cores, its own memory cahe, and is responsible for handling its own distributed tasks. Communicaiton between local and workers and between worker and worker is handled by a cluster manager.
#
# 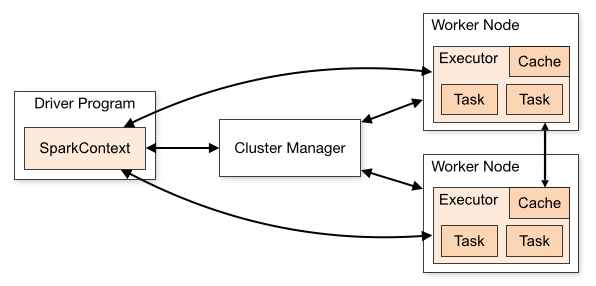
#
# Source: http://spark.apache.org/docs/latest/img/cluster-overview.png
#
# ### Organizaiton of Spark tasks
#
# Spark organizes tasks that can be performed without exchanging data across partitions into stages. The sequecne of tasks to be perfomed are laid out as a Directed Acyclic Graph (DAG). Tasks are differenitated into transforms (lazy evalutation - just add to DAG) and actions (eager evaluation - execute the specified path in the DAG). Note that calculations are not cached unless requested. Hence if you have triggered the action after RDD3 in the figure, then trigger the aciton after RDD6, RDD2 will be re-generated from RDD1 twice. We can avoid the re-calculation by persisting or cacheing RDD2.
#
# 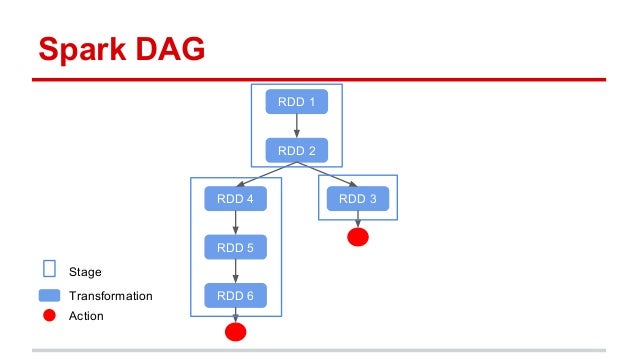
#
# Source: https://image.slidesharecdn.com/mapreducevsspark-150512052504-lva1-app6891/95/map-reduce-vs-spark-16-638.jpg?cb=1431408380
# - [PySpark API](https://spark.apache.org/docs/latest/api/python/pyspark.html)
# ## SparkContext
#
# A SparkContext represents the connection to a Spark cluster, and can be used to create RDDs, accumulators and broadcast variables on that cluster. Here we set it up to use local nodes - the argument `locals[*]` means to use the local machine as the cluster, using as many worker threads as there are cores. You can also explicitly set the number of cores with `locals[k]` where `k` is an integer.
#
# With Saprk 2.0 onwards, there is also a SparkSession that manages DataFrames, which is the preferred abstraction for working in Spark. However DataFrames are composed of RDDs, and it is still necesaary to understand how to use and mainpulate RDDs for low level operations.
# Depending on your setup, you many have to import SparkContext. This is not necessary in our Docker containers as we will be using `livy`.
#
# ```python
# from pyspark import SparkContext
# sc = SparkContext(master = 'local[*]')
# ```
# Start spark
# + language="spark"
# -
import numpy as np
# Version
spark.version
# Number of workers
sc.defaultParallelism
# Data in an RDD is distributed across partitions. It is most efficient if data does not have to be transferred across partitions. We can see the default minimumn number of partitions, and the actual number in an RDD later.
sc.defaultMinPartitions
# ## Resilient Distributed Datasets (RDD)
#
#
# ### Creating an RDD
#
# The RDD (Resilient Distributed Dataset) is a data storage abstraction - you can work with it as though it were single unit, while it may actually be distributed over many nodes in the computing cluster.
# #### A first example
# Distribute the data set to the workers
xs = sc.parallelize(range(10))
xs
xs.getNumPartitions()
# Return all data within each partition as a list. Note that the glom() operation operates on the distributed workers without centralizing the data.
xs.glom().collect()
# Only keep even numbers
xs = xs.filter(lambda x: x % 2 == 0)
xs
# Square all elements
xs = xs.map(lambda x: x**2)
xs
# Execute the code and return the final dataset
xs.collect()
# Reduce also triggers a calculation
xs.reduce(lambda x, y: x+y)
# #### A common Spark idiom chains mutiple functions together
(
sc.parallelize(range(10))
.filter(lambda x: x % 2 == 0)
.map(lambda x: x**2)
.collect()
)
# Actions and transforms
# ----
# A **transform** maps an RDD to another RDD - it is a lazy operation. To actually perform any work, we need to apply an **action**.
# ### Actions
x = sc.parallelize(np.random.randint(1, 6, 10))
x.collect()
x.take(5)
x.first()
x.top(5)
x.takeSample(True, 15)
x.count()
x.distinct().collect()
x.countByValue()
x.sum()
x.max()
x.mean()
x.stats()
# #### Reduce, fold and aggregate actions
# **From the API**:
#
# - reduce(f)
#
# > Reduces the elements of this RDD using the specified commutative and associative binary operator. Currently reduces partitions locally.
#
# - fold(zeroValue, op)
#
# > Aggregate the elements of each partition, and then the results for all the partitions, using a given associative function and a neutral “zero value.”
#
# > The function op(t1, t2) is allowed to modify t1 and return it as its result value to avoid object allocation; however, it should not modify t2.
#
# > This behaves somewhat differently from fold operations implemented for non-distributed collections in functional languages like Scala. This fold operation may be applied to partitions individually, and then fold those results into the final result, rather than apply the fold to each element sequentially in some defined ordering. For functions that are not commutative, the result may differ from that of a fold applied to a non-distributed collection.
#
# - aggregate(zeroValue, seqOp, combOp)
#
# > Aggregate the elements of each partition, and then the results for all the partitions, using a given combine functions and a neutral “zero value.”
#
# > The functions op(t1, t2) is allowed to modify t1 and return it as its result value to avoid object allocation; however, it should not modify t2.
#
# > The first function (seqOp) can return a different result type, U, than the type of this RDD. Thus, we need one operation for merging a T into an U and one operation for merging two U
#
# Notes:
#
# - All 3 operations take a binary op with signature op(accumulator, operand)
x = sc.parallelize(np.random.randint(1, 10, 12))
x.collect()
# **max** using reduce
x.reduce(lambda x, y: x if x > y else y)
# **sum** using `reduce`
x.reduce(lambda x, y: x+y)
# **sum** using fold
x.fold(0, lambda x, y: x+y)
# **prod** using reduce
x.reduce(lambda x, y: x*y)
# **prod** using fold
x.fold(1, lambda x, y: x*y)
# **sum** using aggregate
x.aggregate(0, lambda x, y: x + y, lambda x, y: x + y)
# **count** using aggregate
x.aggregate(0, lambda acc, _: acc + 1, lambda x, y: x+y)
# **mean** using aggregate
sum_count = x.aggregate([0,0],
lambda acc, x: (acc[0]+x, acc[1]+1),
lambda acc1, acc2: (acc1[0] + acc2[0], acc1[1]+ acc2[1]))
sum_count[0]/sum_count[1]
# **Warning**: Be very careful wiht fold and aggregate - the zero value must be "neutral". The behhavior can be different from Python's reduce with an initial value.
xs = x.collect()
xs = np.array(xs)
# **Exercise**: Explain the results shown below:
from functools import reduce
reduce(lambda x, y: x + y, xs, 1)
x.fold(1, lambda acc, val: acc + val)
x.aggregate(1, lambda x, y: x + y, lambda x, y: x + y)
# **Exercise**: Explain the results shown below:
reduce(lambda x, y: x + y**2, xs, 0)
np.sum(xs**2)
x.fold(0, lambda x, y: x + y**2)
x.aggregate(0, lambda x, y: x + y**2, lambda x, y: x + y)
# **Exercise**: Explain the results shown belwo:
x.fold([], lambda acc, val: acc + [val])
seqOp = lambda acc, val: acc + [val]
combOp = lambda acc, val: acc + val
x.aggregate([], seqOp, combOp)
# ### Transforms
x = sc.parallelize([1,2,3,4])
y = sc.parallelize([3,3,4,6])
x.map(lambda x: x + 1).collect()
x.filter(lambda x: x%3 == 0).collect()
# #### Think of flatMap as a map followed by a flatten operation
x.flatMap(lambda x: range(x-2, x)).collect()
x.sample(False, 0.5).collect()
# #### Set-like transformss
y.distinct().collect()
x.union(y).collect()
x.intersection(y).collect()
x.subtract(y).collect()
x.cartesian(y).collect()
# Note that flatmap gets rid of empty lists, and is a good way to ignore "missing" or "malformed" entires.
def conv(x):
try:
return [float(x)]
except:
return []
# +
s = "Thee square root of 3 is less than 3.14 unless you divide by 0".split()
x = sc.parallelize(s)
x.collect()
# -
x.map(conv).collect()
x.flatMap(conv).collect()
# Working with key-value pairs
# ----
#
# RDDs consissting of key-value pairs are required for many Spark operatinos. They can be created by using a function that returns an RDD composed of tuples.
data = [('ann', 'spring', 'math', 98),
('ann', 'fall', 'bio', 50),
('bob', 'spring', 'stats', 100),
('bob', 'fall', 'stats', 92),
('bob', 'summer', 'stats', 100),
('charles', 'spring', 'stats', 88),
('charles', 'fall', 'bio', 100)
]
rdd = sc.parallelize(data)
rdd.keys().collect()
rdd.collect()
# #### Functions `ByKey`
# Sum values by key
(
rdd.
map(lambda x: (x[0], x[3])).
reduceByKey(lambda x, y: x + y).
collect()
)
# Running list of values by key
(
rdd.
map(lambda x: ((x[0], x[3]))).
aggregateByKey([], lambda x, y: x + [y], lambda x, y: x + y).
collect()
)
# Average by key
(
rdd.
map(lambda x: ((x[0], x[3]))).
aggregateByKey([], lambda x, y: x + [y], lambda x, y: x + y).
map(lambda x: (x[0], sum(x[1])/len(x[1]))).
collect()
)
# Using a different key
(
rdd.
map(lambda x: ((x[2], x[3]))).
aggregateByKey([], lambda x, y: x + [y], lambda x, y: x + y).
map(lambda x: (x[0], sum(x[1])/len(x[1]))).
collect()
)
# ### Using key-value pairs to find most frequent words in Ulysses
# +
hadoop = sc._jvm.org.apache.hadoop
fs = hadoop.fs.FileSystem
conf = hadoop.conf.Configuration()
path = hadoop.fs.Path('/data/texts')
for f in fs.get(conf).listStatus(path):
print f.getPath()
# -
ulysses = sc.textFile('/data/texts/Portrait.txt')
ulysses.take(10)
import string
def tokenize(line):
table = dict.fromkeys(map(ord, string.punctuation))
return line.translate(table).lower().split()
words = ulysses.flatMap(lambda line: tokenize(line))
words.take(10)
words = words.map(lambda x: (x, 1))
words.take(10)
counts = words.reduceByKey(lambda x, y: x+y)
counts.take(10)
counts.takeOrdered(10, key=lambda x: -x[1])
# ### Word count chained version
(
ulysses.flatMap(lambda line: tokenize(line))
.map(lambda word: (word, 1))
.reduceByKey(lambda x, y: x + y)
.takeOrdered(10, key=lambda x: -x[1])
)
# ### Avoiding slow Python UDF tokenize
#
# We will see how to to this in the DataFrames notebook.
# ### CountByValue Action
# If you are sure that the results will fit into memory, you can get a dacitonary of counts more easily.
wc = (
ulysses.
flatMap(lambda line: tokenize(line)).
countByValue()
)
wc['the']
# Persisting data
# ----
#
# The `top_word` program will repeat ALL the computations each time we take an action such as `takeOrdered`. We need to `persist` or `cache` the results - they are similar except that `persist` gives more control over how the data is retained.
counts.is_cached
counts.persist()
counts.is_cached
counts.takeOrdered(5, lambda x: -x[1])
counts.take(5)
counts.takeOrdered(5, lambda x: x[0])
counts.keys().take(5)
counts.values().take(5)
count_dict = counts.collectAsMap()
count_dict['circle']
# #### Using cache instead of persist
counts.unpersist()
counts.is_cached
counts.cache()
counts.is_cached
# ### Merging key, value datasets
#
# We will build a second counts key: value RDD from another of Joyce's works - Portrait of the Artist as a Young Man.
portrait = sc.textFile('/data/texts/Portrait.txt')
counts1 = (
portrait.flatMap(lambda line: tokenize(line))
.map(lambda x: (x, 1))
.reduceByKey(lambda x,y: x+y)
)
counts1.persist()
# #### Combine counts for words found in both books
joined = counts.join(counts1)
joined.take(5)
# #### sum counts over words
s = joined.mapValues(lambda x: x[0] + x[1])
s.take(5)
# #### average counts across books
avg = joined.mapValues(lambda x: np.mean(x))
avg.take(5)
| notebooks/S15B_Working_With_RDDs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # "Floating point numbers"
# > "Understand computer representation of real numbers"
#
# - toc:true
# - branch: master
# - badges: true
# - comments: true
# - author: <NAME>
# - categories: [floating, point]
# + [markdown] id="lqBaWMY3txYs"
# Background
# ---
#
# Computer memory can store only discrete values, more specifically a predefined number of 0's and 1's for each data type. This also means, all the numbers on a continuous real number line cannot be be represented using predefined size for storage space and we will have to truncate/round extra digits that cannot be fit in the limited space. More specifically, this forces us to limit the range (minimum and maximum values) and density/packing (amount of numbers represented in a given interval) of representable real numbers.
#
# **Fixed point notation**
#
# One way we could represent real numbers is by using a fixed number of bits for the value before the decimal point and for the value after the decimal point. The problem is that we could be wasting space, and will not able to represent very small or very large numbers. For example consider 2 places for values before decimal point and 2 places for values after decimal point in a decimal number system (ignoring for now that computer actually uses binary number system and each place correspond to one bit of space). You can represent 00.01 to 99.99 nonzero values. The location of decimal point is fixed.
#
# **Scientific notation to the rescue**
#
# Could we optimize further? For example using scientific notation with the same number of places, we can store the exponent (in a chosen, implicitly assumed number base) and mantissa. Since we are storing exponent (as opposed to the integer part before the decimal point), we can cover a wider range of values. Also, let's allocate one bit for the sign (0 => positive number, 1 => negative number). Concretely,
#
# $(-1)^{sign} * mantissa * {\beta}^{e}$
#
# Here `mantissa` has a decimal point and we use $\beta$ as the base and $e$ is the exponent.
#
# Taking above example now you can represent $0.0 * 10^{00}$ to $9.9 * 10^{99}$ (for brevity, ignoring the negative numbers, and assuming we keep one place value before the decimal point and another for the digit after the decimal point. And two digits for the exponent. A total of 4 digits). By allowing negative values for the exponent, you can store very small and large numbers. In above example, smallest nonzero number $0.1 * 10^{-49}$ to $9.9 * 10^{50}$. You could use 2's complement or come up with a biased representation convention where, for example, you subtract 49 from exponent positions to get actual exponent so that you can represent negative exponents as non-negative numbers for ease. So, 0 to 48 represent negative exponents, 49 is exponent 0 and 50 to 99 represent positive exponents. Also, notice also that depending on how many places we are using after the decimal point, we loose precision.
#
# To reiterate, with fixed point, you know implicitly where the decimal point is - with floating point, decimal point location is encoded and dynamic (based on the value of exponent). And, very small and large numbers can also be represented.
#
# In both representations, if a number to be represented is not rational like $\sqrt 2$ or that do not have limited digits representation in chosen base (for example 0.1 has recurring digit pattern in binary representation), or has more digits than space allocated for `mantissa`, not all digits after the decimal point can be represented. So we have to truncate or round off leading to loss `precision`. More on this later.
#
# Also, notice that since we are using exponent, the gap between the consecutive representable numbers on the real line won't increase linearly, but exponentially or log-linearly as the exponent increases.
#
# **Normalization**
#
# Unfortunately, above representation is not unique as we could represent the same number in multiple ways, for example:
#
# $(-1)^{sign} * (mantissa * \beta) * {\beta}^{e-1}$, here ${mantissa}_{new} = {mantissa}$ * $\beta$ and $e_{new}$ = $e - 1$.
#
# How to deal with this non-unique representations? Normalize! Notice that if we make sure that `mantissa` is within the range defined by the following relation:
#
# $1 \le mantissa \lt \beta$
#
# This guarantees that the representation is unique. Why $1 \ge$, you may ask. If not, again representation is not unique. For example $0.01 * 10^2 = 0.001 * 10^3$. You already saw an example on why `mantissa` has to be less than $\beta$.
#
# So, in a normalized floating-point number, the mantissa has one non-zero digit to the left of the decimal point (another way to do this is have a non-zero first digit after the decimal point and only zero before the decimal point). The number zero has no normalized representation as it has no non-zero digit to put just to the left of the decimal point. Any floating-point number that can't be represented this way is said to be denormalized (for example the exponent becomes smaller than the smallest allowed value after normalizing).
#
# In case where $\beta=2$, the only possible value for the digit left of decimal in mantissa is `1` (for decimal system, the digits are `1` through `9`), so it can be effectively *implicitly* assumed and leave one extra bit available to increase the precision.
#
# Let $E_{min}$ = minimum value representable using given number of slots/bits for exponent. Likewise, for $E_{max}$.
#
# Let p = number of slots/digits in `mantissa` including the single digit before the decimal point.
#
# Let x be an arbitrary real number and fl(x) its floating point representation. So, if $x = (d_0.d_1 \dots d_{p-1} d_p d_{p+1} \dots)_\beta * {\beta}^e$, then $fl(x) = (d_0.d_1 \dots d_{p-1})_\beta * {\beta}^e$. In the rest of the document, the base is not shown for brevity.
#
# **Then, how do we represent 0?**
#
# We can either (a) say that all zeros in `mantissa` means it is 0 - but then we cannot assume implicit digit. So this reduces the precision (b) use one of the exponent (like $E_{min}$) to indicate 0, reducing the space of available exponents by 1 and keep higher precision. This also preserves that the numerical ordering of nonnegative real numbers corresponds to the lexicographic ordering of their floating-point representations. This option also allows us to compare two numbers by just doing integer comparison of sign bit and exponent bits in integer unit of computer instead of floating point unit. Integer units are faster and cheaper to build them. In the rest of the document, this second convention is adopted as this is the same strategy used by IEEE-754 standard.
#
# **What about bias?**
#
# Another complication is that the exponent needs to be negative to represent very small numbers. So, it could be represented using 2's complement. But comparison will be slower. By using a biased representation, it can be made faster. You saw an example above where we subtracted 49. In the rest of the document, this biased representation convention is adopted as this is the same strategy used by IEEE-754 standard.
#
# **Subnormal numbers**
#
# So, if we represent 0 with all zeros in `exponent`, we have a decision about the values in `mantissa`? We can say (1) all zeros in `mantissa` means the we are representing 0 (2) non zero `mantissa` means, it is denormalized number (do not assume *implicit* 1 in case of binary). This second option is useful. Why?
#
# The gap between 0 and the smallest normalized number is ($\beta^{E_{min}+1}$) greater than the gap between smallest normalized number and the next bigger floating point number ($\beta^{E_{min}+1-p+1}$). So, in general, even though the gap is non-increasing as we go from higher numbers to lower numbers, this rule is violated between the smallest normalized number and 0. So, this requires special case handling in proofs for this range or exclude certain floating point numbers from the proofs. Also, if the result of an operation (like subtracting two very small numbers whose exponent is already minimum allowed) cannot be normalized, it would have to be flushed to zero, an abrupt change. What can we do about this? If we allow denormalized numbers, we can assert that if the difference between $x_0$ and its nearest floating point is, say $u$, then, for any $x \le x_0$, the difference between x and its nearest floating point is at most $u$. Using this fact in proofs, we can conclude that the error produced by some sequence of computations is at most some value. Also, they help to underflow gradually. See slide 25 of http://www.cas.mcmaster.ca/~qiao/courses/cas708/slides/ch01.pdf for a picture.
#
#
# **Underflow and overflow**
#
# Given that we are using a fixed number of bits/places for exponent, you have a $E_{min}$ and a $E_{max}$ value that exponent can take. Underflow is the situation where numbers whose exponent is below the $E_{min}+1$ (remember, $E_{min}$ is already used to represent 0) exponent and hence cannot be represented (mostly as a result of an operation like subtracting two close numbers). Overflow is about the numbers whose exponent is above the $E_{max}-1$ (remember, $E_{max}$ is already used for $+-\infty$) exponent value and hence cannot be represented as well. Note that each kind of numbers can be positive or negative depending on the sign bit/place.
#
# Subnormal numbers can help with gradual underflow. Consider for example, $x = 1.10 * 10^{E_{min}+1}$, and $y = 1.00 * 10^{E_{min}+1}$. Then $x-y = 0.1 * 10^{E_{min}+1} = 1.0 * 10^{E_{min}}$. The exponent is smaller than smallest representable value, hence result will be $0$. So, even though $x \ne y$, $x-y = 0$. To handle these cases, we could use denormalized/subnormal numbers to represent the underflow numbers (above will be $0.1 * 10^{E_{min}}$ i.e., do not constrain that digit before the decimal point > 1). This also means we do not assume *implicit* 1, in case of binary when exponent = ${E_{min}}$. Since there could be many zeros after the decimal point, we lose precision (we have to truncate/round the small number to fit in the bits/place of `mantissa` size). Another example $a = 3.0 * 2^{-64}$ and $a * a$ is too small to be represented in 32 bit float format (IEEE-754).
#
# Also, notice that when we subtract two nearly equal quantities (and hence have matching digits for a great number of positions in `mantissa`), there will be significant loss of precision due to these large number of zeros in the fraction/mantissa.
#
# **How to represent special numbers**
#
# $\infty$, $-\infty$ ? We can have a convention that all 1's (in binary, or 9's in decimal) in exponent represent $\infty$. The content of `mantissa` in this case is 0. This is because, by convention we say that max value represented is $\beta^{E_{max}}$ and non-zero `mantissa` makes the number value greater than this (i.e., $\infty$) which is meaningless.
#
# We still need to address another aspect: Any number, with the exception of zero, divided by zero yields respectively $\infty$ or $-\infty$. Dividing zero with zero results in the special NaN, the Not a Number value. NaN is useful to represent situations like addition of two infinite numbers. So, how do we represent these? Notice that when representing $\infty$ we assumed all 0's for `mantissa`. So, we can use any non-zero as meaning it is a NaN.
# + [markdown] id="RDPFEh-enByj"
# An example
# ---
#
# Now background is out of our way, let's see an example of how floating points are represented in computer to get a concrete idea.
#
# We first pick how many bits we want to use to represent decimal numbers, say 8 bits.
#
# First bit (MSB) can be used for sign: 0(+ve), 1(-ve)
# Next 3 bits can be used for exponent.
# Remaining 4 bits can be used for mantissa.
#
# Let's use biased representation, so exponent value stored ranges from 0 to 7 (unsigned +ve number) with bias 3. So, actual values for exponent ranges from $E_{min} = -3$ to $E_{max} = 4$.
#
# Let's also say that exponent = 000 with mantissa = 0000 is reserved for 0.
#
# Let's say exponent = 111 mantissa = 0000 reserved for INFINITY.
#
# This will rule out -3 and 4 from being used as exponents for normal numbers. So, actual represented range is from 1 to 6 (biased representation) or as values from -2 to 3.
#
# So, we now have special cases:
#
# exponent = 000 with mantissa != 0000 => subnormal numbers
#
# exponent = 111 with mantissa != 0000 => NaN (Any number, with the exception of zero, divided by zero yields respectively ∞ or -∞. Dividing zero with zero results in the special NaN, the Not a Number value). NaN is useful to represent situations like addition of two infinites, $\frac{0}{0}$.
#
#
# max +ve value:
#
# 0 110 1111
#
#
# max -ve value:
#
# 1 110 1111
#
# Smallest +ve number that can be represented:
#
# 0 000 0001 (subnormal, note that mantissa cannot be 0000 as then it becomes 0).
#
# Smallest -ve number:
#
# 1 000 0001 (subnormal)
#
#
# Smallest normal +ve number:
#
# 0 001 0000
#
# Smallest normal -ve number:
#
# 1 001 0000
#
# Gap between 0 and smallest normal +ve number = $2^{1-3} = 0.25$.
# Gap between the smallest normal +ve number and the number next = <0 001 0001> - $2^{1-3}$ = $1.0001 x 2^{1-3} - 1.0000 x 2^{1-3} = 0.0001 x 2^{1-3} = 2^{1-3-4} = 0.015625$.
#
# So, we notice that floating point representation is discrete (unlike real line), not equally spaced throughout, and finite.
# + [markdown] id="Qs3JFwThmTeU"
# Measuring rounding errors
# ---
#
# Note that we are representing a real number using a fixed width storage (sign+mantissa+exponent) and hence we have a rounding error. This is measured using ULP standing for units in the last place. For a real number x, when represented using a floating point number, say, fl(x), with exponent e, a least possible change in the mantissa modifies the represented value fl(x) by ${\beta}^e * {\beta}^{-p+1} = {\beta}^{e-p+1}$. This is characterized as 1 ULP. So, all floating point numbers with the same exponent, $e$ have the same ULP. So, in a given representation (given radix, precision, base), 1 ULP is different for each value of the exponent and not constant throughout the representable range.
#
# How do we round? there are four types of rounding we can do. Here they are with some examples (where we assume to keep 5 digits after decimal point):
#
# * Round to nearest number representable: e.g. -0.001497 becomes -0.00150.
# * Round to zero/truncate: e.g. -0.001498 becomes -0.00149.
# * Round to +infinity (round up): e.g. -0.001498 becomes -0.00149.
# * Round to –infinity (round down): e.g. -0.001498 becomes -0.00150.
#
#
# How can we ensure the rounding is done correctly? particularly given that we cannot store digits after the lowest possible slot/digit? Use an extra digit(s) called guard digit(s). Then calculations are performed at slightly greater precision, and then stored in standard IEEE floating-point numbers after normalizing and rounding as above. Usually three extra bits are enough to ensure correctness.
#
# ULP is good for measuring rounding error. Relative error is good for measuring rounding error due to various formula (add/subtract etc).
#
# Rounding to the nearest floating-point number corresponds to an error of less than or equal to 0.5 ULP. However, when analyzing the rounding error caused by various formulas, relative error is a better measure as it is not affected by exponent.
# + [markdown] id="ppx6eczBaWAJ"
# Some proofs
# ---
#
# **Rounding error introduced by nearest rounding**
#
# $\beta = radix$ as above.
#
# let $m = \beta - 1$
#
# let $h = \frac{\beta}{2}$
#
# Then, $0.h = \frac{\beta}{2} {\beta}^{-1} = \frac{1}{2}$
#
# $x = d_0.d_1 \dots d_{p-1} d_p d_{p+1} \dots {\beta}^e$
#
# say $d_p \lt h$, then we just truncate
#
# $fl(x) = d_0.d_1 \dots d_{p-1} {\beta}^e$
#
#
# $x - fl(x) = 0. \dots d_{p} d_{p+1} \dots {\beta}^e$
#
# $x - fl(x) = 0.d_{p} d_{p+1} \dots {\beta}^{-p+1} {\beta}^e$
#
#
# $x - fl(x) \le 0.h m m m \dots {\beta}^{-p+1} {\beta}^e$
#
# $x - fl(x) \le 0.h {\beta}^{-p+1} {\beta}^e$
#
# $x - fl(x) \le \frac{1}{2} {\beta}^{-p+1+e}$
#
# Let $d_p \ge h$, then we add $ \frac{\beta}{2} \beta^{e-p} = \frac{1}{2} \beta^{e-p+1}$ and truncate.
#
# $fl(x) = d_0.d_1 \dots d_{p-1} \beta^e + \frac{1}{2} \beta^{e-p+1}$
#
# $x - fl(x) = 0.0 \dots d_{p} d_{p+1} \dots {\beta}^e - \frac{1}{2} \beta^{e-p+1}$
#
#
# $x - fl(x) = 0.d_{p} d_{p+1} \dots {\beta}^{e-p+1} - \frac{1}{2} \beta^{e-p+1}$
#
# $x - fl(x) \le 0.m m \dots {\beta}^{e-p+1} - \frac{1}{2} \beta^{e-p+1}$
#
#
# $x - fl(x) \lt 1.0 {\beta}^{e-p+1} - \frac{1}{2} \beta^{e-p+1}$
#
# $x - fl(x) \lt \frac{1}{2} {\beta}^{-p+1+e}$
#
# Also, ${\beta}^{-p+1+e}$ = ulp(x), so rounding to nearest floating point keeps absolute error within half-ulp.
#
#
# Consider relative error,
#
# $\frac{|x-fl(x)|}{|fl(x)|} \lt \frac{\frac{1}{2} {\beta}^{-p+1+e}}{\beta^e} = \frac{1}{2} \beta^{-p+1}$
#
# (here note that $|fl(x)| ≥ 1.0 × \beta^e$}
#
# Relative error is indpendent of exponent, hence the same for all numbers.
#
#
# **Rounding error introduced by pure truncation is twice as much. Why?**
#
#
# $fl(x) = d_0.d_1 \dots d_{p-1} {\beta}^e$
#
# $x - fl(x) = 0. \dots d_{p} d_{p+1} \dots {\beta}^e$
#
# $x - fl(x) \le 0.m m \dots {\beta}^{e-p+1}$
#
# $x - fl(x) \lt 1.0 \dots {\beta}^{e-p+1}$
#
#
# Operations (subtraction etc.,) using fixed bits in hardware lead to high rounding errors (the ulp between actual result vs. computed is high). Using a guard digit helps. But there is also problem of cancellation. And sometimes exact rounding (i.e,, assume infinite precision bits while operating, and round the result afterwards) is needed.
# + [markdown] id="jDvPio_pmu8n"
#
# Arithmetic exceptions
# ---
#
# Overflow condition: ±$\infty$, f = $1111 \dots$
#
# Underflow condition: flush to 0, ±$2^{-bias}$, [denormalized]
#
# Divide by zero: ±$\infty$
#
# Invalid numbers: NaN
#
# Inexact value due to rounding/truncation: arithmetic operations
#
# Obviously inexact would occur very often and is usually ignored. So is the case with underflow. We may want to catch the remaining exceptions.
# + [markdown] id="PJ4XChwWdLSa"
# Sample code to demonstrate some of the concepts
# ---
# + id="swnGRZugauXf"
from decimal import *
getcontext().prec = 28
# A class for creating floating points with given number of bits for exponent and precision
class MyFloat:
def __init__(self, exp_bits, precision):
self.exp_bits = exp_bits
self.mantissa_bits = precision - 1 # implicit 1
self.bias = (2**(self.exp_bits - 1)) - 1
self.max_exp_value = (2**self.exp_bits) - 1
def print_float(f):
print(bin(f))
def create(self, s, e, m):
return (s << (self.exp_bits + self.mantissa_bits) | (e << self.mantissa_bits) | m)
def sign(self, f):
return f >> (self.exp_bits + self.mantissa_bits)
def exp(self, f):
return ((f >> self.mantissa_bits) & (2**self.exp_bits -1))
def mantissa(self, f):
return f & ((2**self.mantissa_bits) - 1)
def fraction(self, f):
fracs = []
for i in range(0, self.mantissa_bits):
e = 2**(self.mantissa_bits-i)
fracs.append( (f & 1)/e )
f = f >> 1
return sum(fracs)
def _epsilon(self):
return -self.mantissa_bits
def ulp(self, n):
e = self.exp(n)
s = self.sign(n)
m = self.mantissa(n)
r = self._special(e, s, m)
if r is not None:
return r
e = e - self.bias
return "2**" + str(self._epsilon() + e)
def as_binary(self, f):
# TODO: handle subnormal and special numbers
s = 0
if f < 0:
s = 1
f = -f
e = 0
while f < 1:
f *= 2
e -= 1
while f > 2:
f = f/2
e += 1
e = e + self.bias
f -= 1 # implicit 1
mbits = []
if (f != 0):
while True:
f *= 2
if f > 1:
f -= 1
mbits.append("1")
elif f < 1:
mbits.append("0")
else:
mbits.append("1")
break
if (len(mbits) == self.mantissa_bits):
break
txt = "{0} " + "{0:b}".format(e).zfill(self.exp_bits) + " " + "".join(mbits).zfill(self.mantissa_bits)
return txt.format(s)
def _special(self, e, s, m):
if (e == self.max_exp_value) and (m == 0) and (s == 0):
return "+Infinity"
if (e == self.max_exp_value) and (m == 0) and (s == 1):
return "-Infinity"
if (e == self.max_exp_value and s == 0):
return "NaN"
if (e == self.max_exp_value and s == 1):
return "-NaN"
return None
def as_decimal(self, n):
e = self.exp(n)
s = self.sign(n)
m = self.mantissa(n)
r = self._special(e, s, m)
if r is not None:
return r
frac = self.fraction(m)
if e > 0 and e < self.max_exp_value:
# normalized numbers, add implicit 1
frac += 1.0
e -= self.bias
if e == -self.bias:
e += 1 # subnormal numbers, representable numbers which are immediately close to smallest normal number
# additionally, if we keep e=-3, then numbers smaller than smallest are represented causing confusion
# in this case we also cannot use assumed b0 = 1 as that leads to duplicate numbers, for example then one with when e=-2 already
return Decimal((-1)**self.sign(n)) * Decimal(2**e) * Decimal(frac)
# + colab={"base_uri": "https://localhost:8080/"} id="p4LEm1dhg43-" outputId="83543bbd-078c-4774-e736-c7358fc307a1"
# Create IEEE 754 style single float and print details to test the MyFloat class above
myFloat32 = MyFloat(8, 24)
def print_details(n):
print(bin(myFloat32.sign(n)))
print(bin(myFloat32.exp(n)))
print(bin(myFloat32.mantissa(n)))
print(myFloat32.mantissa_bits)
print(myFloat32.as_decimal(n))
print(myFloat32.ulp(n))
#print_details(0x3E800000)
#print_details(0x3E1BA5E3) # 1.5199999511241912841796875E-1
#print_details(0x3F7F7CEE) # 9.9800002574920654296875E-1
print_details(0x4480C000) # 1030.000000000
print("0.25 => " + myFloat32.ulp(0x3E800000))
print("2 => " + myFloat32.ulp(0x40000000))
print("3 => " + myFloat32.ulp(0x40400000))
print("4 => " + myFloat32.ulp(0x40800000))
print("10 => " + myFloat32.ulp(0x41200000))
print("100 => " + myFloat32.ulp(0x42C80000))
print("1030 => " + myFloat32.ulp(0x4480C000))
print("1030 => " + myFloat32.ulp(0x4480C000))
#verified above are correct Ulp by testing through https://docs.oracle.com/javase/1.5.0/docs/api/java/lang/Math.html#ulp(float)
print("Bits for 0.25 =", myFloat32.as_binary(0.25))
print("Bits for 0.1 =", myFloat32.as_binary(0.1))
# + [markdown] id="hUi6Epo2qBr_"
# Sample floating point number system
# ---
#
# Create a floating point number type with 1 bit for sign, 8 bits for exponent, and 5 bits (1 implicit and 4 explicit) for mantissa
#
# + colab={"base_uri": "https://localhost:8080/"} id="fw_5QHoFR-Ce" outputId="99726a87-fdef-4238-e404-36d3c4131aae"
# Create a floating point number type with 1 bit for sign, 8 bits for exponent, and 5 bits (1 implicit and 4 explicit) for mantissa
myFloat = MyFloat(3, 5)
print("Epsilon" + str(myFloat._epsilon()))
xs = []
ys = []
for i in range(0, 256):
xs.append(i)
print("N =", i, "-> ", end='')
y = myFloat.as_decimal(i)
print(y)
if (i > 111 and i < 128) or (i > 239 and i < 256):
ys.append(0)
else:
ys.append(y)
# + colab={"base_uri": "https://localhost:8080/", "height": 251} id="RyWrslXOKo2S" outputId="cf2cba07-d6b9-4906-e5a7-875708e7d37f"
# Density plot of the representable numbers for above example
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.pyplot import figure
figure(figsize=(16, 2), dpi=120)
values = ys[0:112]
plt.scatter(values, np.zeros_like(values), cmap="hot_r", vmin=-2)
plt.xticks(np.linspace(0, 16, 20))
plt.yticks([])
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="RsQSd-W9Uoki" outputId="6181b4dd-a6f9-4cb0-8dc7-8cdb5a37a50e"
# density histogram of floating point numbers for above example
import numpy as np
import matplotlib.pyplot as plt
lys = []
for i in range(16, 112):
ly = myFloat.as_decimal(i)
lys.append(ly)
plt.hist(lys, 'auto', facecolor='g', alpha=0.75)
plt.xlabel('floating point range')
plt.ylabel('count')
plt.title('Histogram showing floating point packing')
plt.grid(True)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="kqPaTAzctdrZ" outputId="401b7e23-f57a-4e92-c97f-fd99deacedbb"
# density histogram of log of floating point numbers for above example
import numpy as np
import matplotlib.pyplot as plt
plt.hist([np.log(float(x)) for x in lys], 'auto', facecolor='g', alpha=0.75)
plt.xlabel('log-floating point range')
plt.ylabel('count')
plt.title('Histogram showing log-linear packing of floating point numbers')
plt.grid(True)
plt.show()
# + [markdown] id="Os9XjfKW5OnO"
# Exercises
# ---
#
# * Understand half precision float vs bfloat16 representation
# * Demonstrate how the knowledge of floating point representation is useful:
# * Convert from float to bfloat16.
# * Execute an example arithmetic expression using both half-float and bfloat16 and see the difference in results and explain.
# * Fast square root by halving exponent directly
# * Machine epsilon: when two successive iterates differ by less than $|\epsilon|$, we may assume that the iteration has converged and stop the process. Provide a demonstration of this.
# + [markdown] id="Hny6vhED4sL0"
# References
# ---
#
# * https://docs.oracle.com/cd/E19957-01/806-3568/ncg_goldberg.html
# * https://www.exploringbinary.com/the-spacing-of-binary-floating-point-numbers/
# * https://softwareengineering.stackexchange.com/questions/215065/can-anyone-explain-representation-of-float-in-memory
# * https://stackoverflow.com/questions/55253233/convert-fp32-to-bfloat16-in-c/55254307#55254307
# * https://software.intel.com/content/www/us/en/develop/articles/intel-deep-learning-boost-new-instruction-bfloat16.html
# * http://www.binaryconvert.com/result_float.html?decimal=048046049053054050053
# * http://www.cs.utep.edu/interval-comp/hayes.pdf
# * https://stackoverflow.com/questions/40082459/what-is-overflow-and-underflow-in-floating-point
# * Floating point vs. bigint: https://stackoverflow.com/a/6320218
# * http://www.cs.jhu.edu/~jorgev/cs333/readings/8-Bit_Floating_Point.pdf
# * https://babbage.cs.qc.cuny.edu/IEEE-754.old/Decimal.html
# * https://www.youtube.com/watch?v=p8u_k2LIZyo
# * https://www-ljk.imag.fr/membres/Carine.Lucas/TPScilab/JMMuller/ulp-toms.pdf
# * https://matthew-brett.github.io/teaching/floating_error.html
# * http://www.math.pitt.edu/~trenchea/math1070/MATH1070_2_Error_and_Computer_Arithmetic.pdf
# * http://home.iitk.ac.in/~pranab/ESO208/rajesh/03-04/Errors.pdf
# * https://stackoverflow.com/a/7524916
# * http://www.cas.mcmaster.ca/~qiao/courses/cas708/slides/ch01.pdf
# * https://stackoverflow.com/questions/43965347/ulp-unit-of-least-precision
# * Algorithms for standard operations: https://www.rfwireless-world.com/Tutorials/floating-point-tutorial.html
# * https://www.sciencedirect.com/topics/computer-science/fixed-point-number
# * https://stackoverflow.com/questions/51170944/understanding-the-usefulness-of-denormalized-floating-point-numbers
# * https://www.ias.ac.in/public/Volumes/reso/021/01/0011-0030.pdf
#
| _notebooks/2021-07-27-FloatingPointNumbers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pet feeder - Live Demo
#
# The Jetson will in this demo see if the bowl is empty or full. If empty it will loose the grip (on a feeding bag), and when the bowl is full it will tighten the grip again
# ### Loading the model
# +
import torch
import torchvision
from torch2trt import TRTModule
device = torch.device('cuda')
model_trt = TRTModule()
model_trt.load_state_dict(torch.load('model_trt.pth'))
# -
# ### Creating a preprocessing function
#
# To match the format of the trained model to the camera, a preprocessing function is necessary. This will convert HWC layout to CHW layout, normalize, transfer data from CPU to GPU memory and add a batch dimension.
# +
import torchvision.transforms as transforms
import torch.nn.functional as F
import cv2
import PIL.Image
import numpy as np
mean = torch.Tensor([0.485, 0.456, 0.406]).cuda().half()
std = torch.Tensor([0.229, 0.224, 0.225]).cuda().half()
normalize = torchvision.transforms.Normalize(mean, std)
def preprocess(image):
image = PIL.Image.fromarray(image)
image = transforms.functional.to_tensor(image).to(device).half()
image.sub_(mean[:, None, None]).div_(std[:, None, None])
return image[None, ...]
# -
# ### Creating camera and widgets to show prediction of empty
# +
import traitlets
from IPython.display import display
import ipywidgets.widgets as widgets
from jetbot import Camera, bgr8_to_jpeg
camera = Camera.instance(width=224, height=224)
image = widgets.Image(format='jpeg', width=224, height=224)
empty_slider = widgets.FloatSlider(description='empty', min=0.0, max=1.0, orientation='vertical')
full_slider = widgets.FloatSlider(description='full', min=0.0, max=1.0, orientation='vertical')
camera_link = traitlets.dlink((camera, 'value'), (image, 'value'), transform=bgr8_to_jpeg)
# -
# ### Functions to grab or loose the grip, depending on the prediction
# +
import torch.nn.functional as F
import time
from jetbot import Robot
from SCSCtrl import TTLServo
# Creating robot to enable grabber
robot = Robot()
def grab():
TTLServo.servoAngleCtrl(4, -90, 1, 150)
def loose():
TTLServo.servoAngleCtrl(4, -10, 1, 150)
def update(change):
global empty_slider, full_slider, robot
new_image = change['new']
new_image = preprocess(new_image)
trt_image = model_trt(new_image)
# normalizing the output vector by using softmax
trt_image = F.softmax(trt_image, dim=1)
prob_empty = float(trt_image.flatten()[0])
prob_full = float(trt_image.flatten()[1])
empty_slider.value = prob_empty
full_slider.value = prob_full
# If the probability of an empty bowl is more than 80% then loose the grip
if prob_empty > 0.8:
loose()
else:
grab()
update({'new': camera.value})
# -
# ### Attaching the execution to the camera using observe
# +
camera.observe(update, names='value')
# Displaying image and sliders
display(widgets.VBox([widgets.HBox([image, empty_slider, full_slider])]))
# -
# ### To stop the robot and turn off the camera
# +
import time
camera.unobserve(update, names='value')
time.sleep(0.1) # add a small sleep to make sure frames have finished processing
robot.stop()
camera.stop()
# -
| live_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# orphan: true
# ---
# # Experimental Features
#
# - [Determining Variable Shapes at Runtime](dyn_shapes.ipynb)
# - [Simultaneous Coloring of Approximated Derivatives](approx_coloring.ipynb)
# - [Working with Plugins](plugins)
| openmdao/docs/openmdao_book/features/experimental/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Overview
#
# We're now switching focus away from the Network Science (for a little bit), beginning to think about _Natural Language Processing_ instead. In other words, today will be all about teaching your computer to "understand" text. This ties in nicely with our work on wikipedia, since wikipedia is a network of connected pieces of text. We've looked at the network so far - now, let's see if we can include the text. Today is about
#
# * Installing the _natural language toolkit_ (NLTK) package and learning the basics of how it works (Chapter 1)
# * Figuring out how to make NLTK to work with other types of text (Chapter 2).
# > **_Video Lecture_**. Today is all about working with NLTK, so not much lecturing - you can get my perspective and a little pep-talk
from IPython.display import YouTubeVideo
YouTubeVideo("Ph0EHmFT3n4",width=800, height=450)
# # Installing and the basics
#
# > _Reading_
# > The reading for today is Natural Language Processing with Python (NLPP) Chapter 1, Sections 1.1, 1.2, 1.3\. [It's free online](http://www.nltk.org/book/).
#
# > *Exercises*: NLPP Chapter 1\.
# >
# > * First, install `nltk` if it isn't installed already (there are some tips below that I recommend checking out before doing installing)
# > * Second, work through chapter 1. The book is set up as a kind of tutorial with lots of examples for you to work through. I recommend you read the text with an open IPython Notebook and type out the examples that you see. ***It becomes much more fun if you to add a few variations and see what happens***. Some of those examples might very well be due as assignments (see below the install tips), so those ones should definitely be in a `notebook`.
#
# ### NLTK Install tips
#
# Check to see if `nltk` is installed on your system by typing `import nltk` in a `notebook`. If it's not already installed, install it as part of _Anaconda_ by typing
#
# conda install nltk
#
# at the command prompt. If you don't have them, you can download the various corpora using a command-line version of the downloader that runs in Python notebooks: In the iPython notebook, run the code
#
# import nltk
# nltk.download()
#
# Now you can hit `d` to download, then type "book" to fetch the collection needed today's `nltk` session. Now that everything is up and running, let's get to the actual exercises.
# > *Exercises: NLPP Chapter 1 (the stuff that might be due in an upcoming assignment).
# >
# > The following exercises from Chapter 1 are what might be due in an assignment later on.
# >
# > * Try out the `concordance` method, using another text and a word of your own choosing.
# > * Also try out the `similar` and `common_context` methods for a few of your own examples.
# > * Create your own version of a dispersion plot ("your own version" means another text and different word).
# > * Explain in your own words what aspect of language _lexical diversity_ describes.
# > * Create frequency distributions for `text2`, including the cumulative frequency plot for the 75 most common words.
# > * What is a bigram? How does it relate to `collocations`. Explain in your own words.
# > * Work through ex 2-12 in NLPP's section 1.8\.
# > * Work through exercise 15, 17, 19, 22, 23, 26, 27, 28 in section 1.8\.
# # Working with NLTK and other types of text
#
# So far, we've worked with text from Wikipedia. But that's not the only source of text in the universe. In fact, it's far from it. Chapter 2 in NLPP1e is all about getting access to nicely curated texts that you can find built into NLTK.
# >
# > Reading: NLPP Chapter 2.1 - 2.4\.
# >
# > *Exercises*: NLPP Chapter 2\.
# >
# > * Solve exercise 4, 8, 11, 15, 16, 17, 18 in NLPP1e, section 2.8\. As always, I recommend you write up your solutions nicely in a `notebook`.
# > * Work through exercise 2.8.23 on Zipf's law. [Zipf's law](https://en.wikipedia.org/wiki/Zipf%27s_law) connects to a property of the Barabasi-Albert networks. Which one? Take a look at [this article](http://www.hpl.hp.com/research/idl/papers/ranking/adamicglottometrics.pdf) and write a paragraph or two describing other important instances of power-laws found on the internet.
# >
| lectures/Week6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
# %load_ext Cython
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
from scipy import stats
from scipy.optimize import curve_fit
from lmfit.models import *
from pyarrow import csv
import warnings
warnings.filterwarnings('ignore')
from nicks_plot_utils import Hist1D, Hist2D, Scatter
MP = 0.93827208816
E0 = 4.81726
ME = 0.00051099895
COLOR_BLUE=plt.rcParams['axes.prop_cycle'].by_key()['color'][0]
COLOR_RED=plt.rcParams['axes.prop_cycle'].by_key()['color'][1]
COLOR_DGREEN=plt.rcParams['axes.prop_cycle'].by_key()['color'][3]
# + magic_args="--annotate" language="cython"
# import numpy as np
# cimport numpy as np
# from libc.math cimport sin, cos, sqrt
# from scipy import stats
# cimport cython
#
# cdef float MP = 0.93827208816
# cdef float E0 = 4.81726
# #cdef float E0 = 4.8056
# cdef float ME = 0.00051099895
#
# cdef float p_targ_px = 0.0
# cdef float p_targ_py = 0.0
# cdef float p_targ_pz = 0.0
# cdef float p_targ_E = MP
#
# cdef float e_beam_px = 0.0
# cdef float e_beam_py = 0.0
# cdef float e_beam_pz = sqrt(E0**2-ME**2)
# cdef float e_beam_E = E0
#
# @np.vectorize
# def calc_W(float e_p, float e_theta, float e_phi):
# cdef float e_prime_px = e_p*sin(e_theta)*cos(e_phi)
# cdef float e_prime_py = e_p*sin(e_theta)*sin(e_phi)
# cdef float e_prime_pz = e_p*cos(e_theta)
# cdef float e_prime_E = sqrt(e_prime_px**2 + e_prime_py**2 + e_prime_pz**2 - ME**2)
#
# cdef float temp_px = e_beam_px - e_prime_px + p_targ_px
# cdef float temp_py = e_beam_py - e_prime_py + p_targ_py
# cdef float temp_pz = e_beam_pz - e_prime_pz + p_targ_pz
# cdef float temp_E = e_beam_E - e_prime_E + p_targ_E
#
#
# cdef float temp2 = temp_px**2+temp_py**2+temp_pz**2-temp_E**2
# cdef float temp3 = sqrt(-temp2)
#
#
# return temp3
#
#
# @np.vectorize
# def calc_q2(float e_p, float e_theta, float e_phi):
# cdef float e_prime_px = e_p*sin(e_theta)*cos(e_phi)
# cdef float e_prime_py = e_p*sin(e_theta)*sin(e_phi)
# cdef float e_prime_pz = e_p*cos(e_theta)
# cdef float e_prime_E = sqrt(e_prime_px**2 + e_prime_py**2 + e_prime_pz**2 - ME**2)
#
# cdef float temp_px = e_beam_px - e_prime_px
# cdef float temp_py = e_beam_py - e_prime_py
# cdef float temp_pz = e_beam_pz - e_prime_pz
# cdef float temp_E = e_beam_E - e_prime_E
#
# cdef float temp2 = temp_px**2+temp_py**2+temp_pz**2-temp_E**2
#
# return temp2
# +
@np.vectorize
def center_phi(phi, sec):
sector = {
1: 90,
2: 30,
3: -30,
4: -90,
5: -150,
6: 150
}
return phi-sector[sec]
@np.vectorize
def Theta_e_calc(theta_p):
return 2 * np.arctan(MP/((E0+MP)*np.tan(theta_p)))
@np.vectorize
def momentum_e_calc(e_theta):
return E0/(1 + (2*E0*np.sin(e_theta/2.0)**2)/MP)
def gauss(x, a, x0, sigma):
return a*np.exp(-(x-x0)**2/(2*sigma**2))
def FitFunc2(phi_e, theta_e,
alpha_A, beta_A, gamma_A,
alpha_B, beta_B, gamma_B,
alpha_C, beta_C, gamma_C,
alpha_D, beta_D, gamma_D):
"""
Equations 5.20 - 5.22 in KPark thesis (p. 71)
"""
A = (alpha_A * theta_e**2 + beta_A * theta_e + gamma_A) * phi_e**3
B = (alpha_B * theta_e**2 + beta_B * theta_e + gamma_B) * phi_e**2
C = (alpha_C * theta_e**2 + beta_C * theta_e + gamma_C) * phi_e
D = (alpha_D * theta_e**2 + beta_D * theta_e + gamma_D)
return A + B + C + D
def Dtheta(phi_e, theta_e, A, B, C, D):
"""
Mom Corrections for e6 (CLAS-NOTE 2003-005)
"""
first = (A+B*phi_e)*(np.cos(theta_e)/np.cos(phi_e))
second = (C+D*phi_e)*np.sin(theta_e)
return first + second
def Dpp(phi_e, theta_e, p, Bt, E, F, G, H):
first = (E+F*phi_e)*(np.cos(theta_e)/np.cos(phi_e))
second = (G+H*phi_e)*np.sin(theta_e)
return (first + second)*(p/Bt)
# +
#df = pd.read_csv("/Users/tylern/Data/momCorr.csv")
file_name = "/Users/tylern/Data/momCorr.csv"
pyTable = csv.read_csv(
file_name,
read_options=csv.ReadOptions(use_threads=True)
)
all_data = pyTable.to_pandas(strings_to_categorical=True)
channel = all_data[all_data.type == "channel"].copy()
df = all_data[all_data.type == "elastic"].copy()
df = df.drop(['type'], axis=1)
df = df[(df.W_uncorr > 0.5) & (df.W_uncorr < 1.5)]
df['e_phi_center'] = center_phi(np.rad2deg(df.e_phi), df.sector)
df['e_theta_calc'] = Theta_e_calc(df.p_theta)
df['delta_theta'] = df['e_theta_calc']-df['e_theta']
# df['p_p_calc'] = Theta_e_calc(df.e_theta_calc)
# df['mom_e_calc'] = momentum_e_calc(df.e_theta)
# df['delta_p'] = df.e_p/df.mom_e_calc
df = df[np.abs(df.delta_theta) < 0.005]
#df['w_corr'] = calc_W(df.e_p_corr, df.e_theta, df.e_phi)
#df['q2_corr'] = q2_calc(df.e_p_corr, df.e_theta, df.e_phi)
df.dropna(inplace=True)
# -
df.head()
# +
df['mom_e_calc'] = momentum_e_calc(df.e_theta)
df['delta_p'] = df.mom_e_calc/df.e_p
for sec in range(1,7):
data = df[df.sector == sec].dropna()
fig = plt.figure(figsize=(16,9))
y, x= np.histogram(data.delta_p, bins=500, range=(0.85,1.1))
y = y/np.max(y)
x = (x[:-1]+x[1:])/2.0
xs=np.linspace(0.85,1.1, 500)
mod = SkewedVoigtModel()
pars = mod.guess(y, x=x)
out = mod.fit(y, pars, x=x, nan_policy='omit')
ebar = plt.errorbar(x,y, yerr=stats.sem(y),fmt='.',alpha=0.4,
label=f'Sector: {sec}, {out.params["center"].value:0.2f}, \
{out.params["sigma"].value:0.2f}')
plt.plot(xs, out.eval(x=xs), '--', c=ebar[0].get_color(), lw=2)
# +
num_points = 12
grr_p = []
for sec in range(1,7):
for deg in range(10, 20):
df2 = df[(df.sector == sec) & (np.rad2deg(df.e_theta) >= deg) & (np.rad2deg(df.e_theta) < deg+1)]
phis = np.linspace(np.min(df2.e_phi_center), np.max(df2.e_phi_center), num_points+1)
for phi in range(0, num_points):
phi_min = phis[phi]
phi_max = phis[phi+1]
data = df2[(df2.e_phi_center > phi_min) & (df2.e_phi_center <= phi_max)]
y, x = np.histogram(data['delta_p'], bins=100, range=(0.85,1.1))
y = y/np.max(y)
x = (x[:-1]+x[1:])/2.0
if not np.any(np.isnan(y)) and len(y) >= 5:
try:
mod = GaussianModel()
pars = mod.guess(y, x=x)
out = mod.fit(y, pars, x=x)
if out.params['center'] < 1.01:
grr_p.append([((phi_min+phi_max) / 2.0),
out.params['center'],
out.params['fwhm'],
sec,
np.deg2rad(deg)])
except RuntimeError:
pass
grr_p = np.array(grr_p)
dfgrr_p = pd.DataFrame(data=grr_p, columns=["phi", "dP", "sigma", "sec", "min_deg"])
# +
final_fit_parameters_P = {}
#for sec in range(1,7):
for sec in [1]:
print(sec)
xs = np.linspace(-20,20,1000)
d = dfgrr_p[dfgrr_p.sec == sec]
fig = plt.figure(figsize=(12,8))
scatter = plt.scatter(d.phi, d.dP, c=np.rad2deg(d.min_deg))
sec_values = []
for i, deg in enumerate(np.unique(d.min_deg)):
d2 = dfgrr_p[(dfgrr_p.sec == sec) & (dfgrr_p.min_deg == deg)]
z = np.polyfit(d2.phi, d2.dP, 3)
p = np.poly1d(z)
sec_values.append(z)
#mod = PolynomialModel(3)
#pars = mod.guess(d2.dP, x=d2.phi)
#out = mod.fit(d2.dP, pars, x=d2.phi)
try:
plt.plot(xs, p(xs), alpha=0.8, c=scatter.legend_elements()[0][i].get_color())
except IndexError:
plt.plot(xs, p(xs), alpha=0.8)
plt.xlabel("$\phi_e$")
plt.ylabel("$\delta p_e$")
# print(out.params)
legend = plt.legend(*scatter.legend_elements(), title="$\\theta_e$", loc="upper right")
plt.ylim(0.8, 1.2)
plt.title("$\delta \\theta_e$ vs $\phi_e$ for slices in $\\theta_e$")
plt.show()
fig = plt.figure(figsize=(12,8))
sec_values = np.array(sec_values)
label="ABCD"
for i, letter in enumerate(label):
scalling = 10**(3-i)
if letter == "D":
scalling = 10**2
x = np.unique(np.rad2deg(dfgrr_p.min_deg))
y = sec_values.T[i]
if len(x) != len(y):
continue
sca = plt.errorbar(x, y*scalling, fmt='.', label=f'{letter} $\\times 10^{(5-i)}$')
z = np.polyfit(x, y, 3)
p = np.poly1d(z)
xx = np.linspace(12, 20, 500)
plt.plot(xx, p(xx)*scalling, alpha=0.5, c=sca[0].get_color())
mod = PolynomialModel(2)
pars = mod.guess(y, x=x)
#pars['c0'].set(value=0, min=-1E-4, max=1E-4)
#pars['c1'].set(value=0, min=-1E-4, max=1E-4)
#pars['c2'].set(value=0, min=-1E-15)
out = mod.fit(y, pars, x=x)
#plt.plot(xx, out.eval(params=out.params, x=xx), alpha=0.8, c=sca[0].get_color())
for j, abc in enumerate(['alpha','beta','gamma']):
final_fit_parameters_P[f'sec_{sec}_{letter}_{abc}'] = z[j]
#final_fit_parameters[f'sec_{sec}_{letter}_{abc}'] = out.params[f'c{j}'].value
plt.ylim(-0.5, 0.5)
plt.title("Fits of parameter functions A,B,C,D,E")
plt.ylabel("$\delta \\theta$")
plt.xlabel("$\\theta_e$")
plt.legend(loc="upper right")
plt.show()
print(final_fit_parameters_P)
# +
final_fit_parameters_P = {}
for sec in range(1,7):
for i, letter in enumerate("ABCD"):
for j, abc in enumerate(['alpha','beta','gamma']):
final_fit_parameters_P[f'sec_{sec}_{letter}_{abc}'] = 1
for sec in range(1,7):
print(sec)
xs = np.linspace(-20,20,1000)
mu = np.mean(dfgrr_p.dP)
sig = np.std(dfgrr_p.dP)
nsig = 3
d = dfgrr_p[(dfgrr_p.sec == sec) & (np.abs(dfgrr_p.dP) < mu+nsig*sig)]
fig, ax = plt.subplots(nrows=1, ncols=2, figsize=(16,9))
scatter = ax[0].scatter(d.phi, d.dP, c=np.rad2deg(d.min_deg))
sec_values = []
for deg in np.unique(dfgrr_p.min_deg):
d2 = dfgrr_p[(dfgrr_p.sec == sec) & (dfgrr_p.min_deg == deg)]
if len(d2) < 4:
continue
z = np.polyfit(d2.phi, d2.dP, 4)
p = np.poly1d(z)
ax[0].plot(xs, p(xs), label=z, alpha=0.8)
sec_values.append([deg, z])
legend = ax[0].legend(*scatter.legend_elements(), title="deg", loc='upper right')
ax[0].set_ylim(0.97, 1.03)
sec_values = np.array(sec_values)
print(sec_values)
label="ABCD"
for i, letter in enumerate("ABCD"):
ys = sec_values.T[i]
min_deg = sec_values.T[0][i] #np.unique(dfgrr_p.min_deg)
print(min_deg, ys)
#min_deg = min_deg[ np.abs(ys*10**(4-i)) < 0.2 ]
#ys = ys[ np.abs(ys*10**(4-i)) < 0.2 ]
# sca = ax[1].errorbar(min_deg, ys*10**(4-i), fmt='.', label=letter)
z = np.polyfit(min_deg, ys, 2)
p = np.poly1d(z)
xx = np.linspace(0.2, 0.5, 500)
ax[1].plot(xx, p(xx)*10**(4-i), alpha=0.5, c=sca[0].get_color())
mod = PolynomialModel(2)
pars = mod.guess(y, x=x)
pars['c0'].set(value=0, min=-1E-4, max=1E-4)
pars['c1'].set(value=0, min=-1E-4, max=1E-4)
pars['c2'].set(value=0, min=0)
out = mod.fit(y, pars, x=x)
#plt.plot(xx, out.eval(params=out.params, x=xx)*10**(5-i), alpha=0.8, c=sca[0].get_color())
for j, abc in enumerate(['alpha','beta','gamma']):
final_fit_parameters_P[f'sec_{sec}_{letter}_{abc}'] = z[j]
# final_fit_parameters_P[f'sec_{sec}_{letter}_{abc}'] = out.params[f'c{j}'].value
ax[1].set_ylim(-0.5, 0.5)
ax[1].legend(loc='upper right')
plt.show()
print(final_fit_parameters_P)
# +
df['p_corr'] = df['e_p']
#for sec in [1,5,6]:
for sec in [1,2,4,5,6]:
mask = (df.sector == sec)
valid = df[mask].copy()
if(sec == 4):
sec = 1
correction = FitFunc2(valid.e_phi, valid.e_theta,
final_fit_parameters_P[f'sec_{sec}_A_alpha'], final_fit_parameters_P[f'sec_{sec}_A_beta'], final_fit_parameters_P[f'sec_{sec}_A_gamma'],
final_fit_parameters_P[f'sec_{sec}_B_alpha'], final_fit_parameters_P[f'sec_{sec}_B_beta'], final_fit_parameters_P[f'sec_{sec}_B_gamma'],
final_fit_parameters_P[f'sec_{sec}_C_alpha'], final_fit_parameters_P[f'sec_{sec}_C_beta'], final_fit_parameters_P[f'sec_{sec}_C_gamma'],
final_fit_parameters_P[f'sec_{sec}_D_alpha'], final_fit_parameters_P[f'sec_{sec}_D_beta'], final_fit_parameters_P[f'sec_{sec}_D_gamma'],)
df.loc[mask, 'p_corr'] = valid.e_p * correction
df['W_corr'] = calc_W(df.p_corr, df.e_theta, df.e_phi)
df['Q2_corr'] = calc_q2(df.p_corr, df.e_theta, df.e_phi)
for sec in range(1,7):
fig = plt.figure(figsize=(16,9))
plt.axvline(MP, c='r')
xs=np.linspace(0.5, 1.5, 500)
data = df[df.sector == sec]
for W in ['W_uncorr','W_corr']:
y, x= np.histogram(data[W], bins=500, range=(0.8,1.2))
y = y/np.max(y)
x = (x[:-1]+x[1:])/2.0
mod = SkewedVoigtModel()
pars = mod.guess(y, x=x)
out = mod.fit(y, pars, x=x, nan_policy='omit')
ebar = plt.errorbar(x,y, yerr=stats.sem(y),fmt='.', alpha=0.4, label=f'Sector: {sec} {W}, {out.params["center"].value:0.5f}, {out.params["sigma"].value:0.5f}')
plt.plot(xs, out.eval(x=xs), '--', c=ebar[0].get_color(), lw=2)
plt.legend()
plt.show()
for sec in range(1,7):
fig, ax = plt.subplots(nrows=1, ncols=2, figsize=(16,9))
data = df[df.sector == sec]
for i, W in enumerate(['uncorr','corr']):
ax[i].axvline(MP, c=COLOR_RED)
ax[i].hist2d(data[f'W_{W}'],data[f'Q2_{W}'], bins=150, range=[[0.6,1.4],[0,5]])
plt.legend()
plt.show()
# +
all_data['p_corr'] = all_data['e_p']
#for sec in [1,5,6]:
for sec in range(1,7):
mask = (all_data.sector == sec)
valid = all_data[mask]
if(sec == 4):
sec = 1
correction = FitFunc(valid.e_phi, valid.e_theta,
final_fit_parameters_P[f'sec_{sec}_A_alpha'], final_fit_parameters_P[f'sec_{sec}_A_beta'], final_fit_parameters_P[f'sec_{sec}_A_gamma'],
final_fit_parameters_P[f'sec_{sec}_B_alpha'], final_fit_parameters_P[f'sec_{sec}_B_beta'], final_fit_parameters_P[f'sec_{sec}_B_gamma'],
final_fit_parameters_P[f'sec_{sec}_C_alpha'], final_fit_parameters_P[f'sec_{sec}_C_beta'], final_fit_parameters_P[f'sec_{sec}_C_gamma'],
final_fit_parameters_P[f'sec_{sec}_D_alpha'], final_fit_parameters_P[f'sec_{sec}_D_beta'], final_fit_parameters_P[f'sec_{sec}_D_gamma'],
final_fit_parameters_P[f'sec_{sec}_E_alpha'], final_fit_parameters_P[f'sec_{sec}_E_beta'], final_fit_parameters_P[f'sec_{sec}_E_gamma'],)
all_data.loc[mask, 'p_corr'] = valid.e_p * correction
all_data['W_corr'] = calc_W(all_data.p_corr, all_data.e_theta, all_data.e_phi)
all_data['Q2_corr'] = calc_q2(all_data.p_corr, all_data.e_theta, all_data.e_phi)
for sec in range(1, 7):
fig = plt.figure(figsize=(16,9))
xs=np.linspace(0.5, 4.5, 500)
for W in ['W_uncorr','W_corr']:
y, x= np.histogram(all_data[all_data.sector == sec][W], bins=500, range=(0.5,3.0))
y = y/np.max(y)
x = (x[:-1]+x[1:])/2.0
ebar = plt.errorbar(x,y, yerr=stats.sem(y),fmt='.',alpha=0.4,
label=f'Sector: {sec} {W}')
plt.legend()
plt.show()
# -
for sec in range(1,7):
fig, ax = plt.subplots(nrows=1, ncols=2, figsize=(16,9))
data = all_data[all_data.sector == sec]
for i, W in enumerate(['uncorr','corr']):
ax[i].axvline(MP, c=COLOR_RED)
ax[i].hist2d(data[f'W_{W}'],data[f'Q2_{W}'], bins=150, range=[[0.6,3.0],[0,5]])
plt.legend()
plt.show()
for sec in range(1,7):
print('{{',final_fit_parameters_P[f'sec_{sec}_A_alpha'],',', final_fit_parameters_P[f'sec_{sec}_A_beta'],',', final_fit_parameters_P[f'sec_{sec}_A_gamma'],'},\n',
'{',final_fit_parameters_P[f'sec_{sec}_B_alpha'],',', final_fit_parameters_P[f'sec_{sec}_B_beta'],',', final_fit_parameters_P[f'sec_{sec}_B_gamma'],'},\n',
'{',final_fit_parameters_P[f'sec_{sec}_C_alpha'],',', final_fit_parameters_P[f'sec_{sec}_C_beta'],',', final_fit_parameters_P[f'sec_{sec}_C_gamma'],'},\n',
'{',final_fit_parameters_P[f'sec_{sec}_D_alpha'],',', final_fit_parameters_P[f'sec_{sec}_D_beta'],',', final_fit_parameters_P[f'sec_{sec}_D_gamma'],'},\n',
'{',final_fit_parameters_P[f'sec_{sec}_E_alpha'],',', final_fit_parameters_P[f'sec_{sec}_E_beta'],',', final_fit_parameters_P[f'sec_{sec}_E_gamma'],'}},\n')
| momentumCorrections/MomentumCorrElectronPart2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# # Facebook Page Analysis Using Pandas
#
#
# + active=""
# import pickle
# import pandas as pd
# import string
# from operator import itemgetter
#
# -
# ### Load the data
# +
loaded_data= pickle.load(file=open("steam_data.pkl"))
# -
loaded_data[0]
# ### Normalizing the data
# +
df2 = pd.io.json.json_normalize(data=loaded_data)
# +
df2.head(10)
# -
df2.columns[0]
# ### Tailoring the data
df2.columns[0]
df2.drop('comments.data' , 1 , inplace=True)
df2.drop( df2.columns[[0,1,5,6,7]] , 1 , inplace=True)
df2.rename( columns= { "comments.summary.total_count" : "total_comments"} , inplace=True)
df2.rename( columns= { "likes.summary.total_count" : "total_likes"} , inplace=True)
# ### Cleaning null values
df2.fillna("",inplace=True)
# +
df2.head(10)
# -
# ### Working with Pandas
steam_df= df2
steam_df[ steam_df['total_comments'] > 100 ]
# ### Most Commented on Posts
top_comments=steam_df.sort_values('total_comments',ascending=False)
top_comments= top_comments.head(10)
top_comments
top_comments_id= top_comments['id']
top_comments_id
for i,x in top_comments_id.iteritems():
print ('https://www.facebook.com/'+x )
# ### Most Liked Posts
# +
df2.sort_values('total_likes',inplace=True,ascending=False)
top_likes = df2.head(10)['id']
for i,x in top_likes.iteritems():
print ('https://www.facebook.com/'+x +" "+ str(i))
# +
df2.head(10)
# +
df2.sort_values('total_likes',inplace=True,ascending=False)
top_likes = df2.head(10)['id']
for i,x in top_likes.iteritems():
print ('https://www.facebook.com/'+x +" "+ str(i))
# -
mapping = dict.fromkeys(map(ord, string.punctuation))
# ### Counting top words
# +
def count_words(df):
stop_words=["a","about","above","after","again","against","all","am","an","and","any","are","now",
"aren't","as","at","be","because","been","before","being","below","between","both","but","by",
"can't","cannot","could","couldn't","did","didn't","do","does","doesn't","doing","don't","down",
"during","each","few","for","from","further","had","hadn't","has","hasn't","have","haven't","having",
"he","he'd","he'll","he's","her","here","here's","hers","herself","him","himself","his","how","how's",
"i","i'd","i'll","i'm","i've","if","in","into","is","isn't","it","it's","its","itself","let's","me","more",
"most","mustn't","my","myself","no","nor","not","of",
"off","on","once","only","or","other","ought","our",
"ours ourselves","out","over","own","same","shan't",
"she","she'd","she'll","she's","should","shouldn't","so",
"some","such","than","that","that's","the","their","theirs",
"them","themselves","then","there","there's","these","they",
"they'd","they'll","they're","they've","this","those","through",
"to","too","under","until","up","very","was","wasn't","we","we'd",
"we'll","we're","we've","were","weren't","what","what's","when","when's",
"where","where's","which","while","who","who's","whom","why","why's","with",
"won't","would","wouldn't","you","you'd","you'll","you're","you've","your",'guys','just','day',
"yours","yourself","yourselves","will","ana","isa","can","hwa","wala","msh","wla","hya","will",'enta','7aga','mesh','dah','bas','elly','b2a','3ala','alf','enty','3al',]
mapping = dict.fromkeys(map(ord, string.punctuation))
words={}
for m in df2['message']:
try:
seperated_words=m.split(' ')
except:
pass
# print "ERROR"+str(m)
for word in seperated_words:
word=word.strip()
word=word.lower()
word=word.translate(mapping) # Removes any punctation
if word in stop_words:
continue
if len(word) < 3:
continue
if word in words:
words[word]+=1
else:
words[word]=1
print("done counting words")
return words
# -
words=count_words(steam_df)
words_df= pd.DataFrame(words.items(), columns=['Word' , 'count'])
words_df
words_df.sort_values('count' , ascending=False,inplace=True)
words_df
# %matplotlib inline
words_df[words_df['count'] > 1000 ].plot(x='Word',kind='bar')
#
# +
months=["january",
"february",
"march",
"april",
"May",
"june",
"july",
"august",
"september",
"october",
"november",
"december"]
print words_df[ words_df['Word'].isin(months) ]
# -
# ### Couning Posts the word "Free" and "Discount"
steam_df['free']= steam_df.message.str.contains('free')
steam_df.head()
steam_df.groupby(by= steam_df['free']).count()['id']
| Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# TensorFlow ≥2.0 is required
import tensorflow as tf
from tensorflow import keras
assert tf.__version__ >= "2.0"
# Python ≥3.5 is required
import sys
assert sys.version_info >= (3, 5)
# Scikit-Learn ≥0.20 is required
import sklearn
assert sklearn.__version__ >= "0.20"
# Common imports
import numpy as np
import os
# to make this notebook's output stable across runs
np.random.seed(42)
tf.random.set_seed(42)
# To plot pretty figures
# %matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
# -
from sklearn.datasets import load_sample_image
# Load sample images
china = load_sample_image ( "china.jpg" ) / 255
flower = load_sample_image ( "flower.jpg" ) / 255
images = np . array ([ china , flower ])
batch_size , height , width , channels = images . shape
# Create 2 filters
filters = np . zeros ( shape = ( 7 , 7 , channels , 2 ), dtype = np . float32 )
filters [:, 3 , :, 0 ] = 1 # vertical line
filters [ 3 , :, :, 1 ] = 1 # horizontal line
outputs = tf . nn . conv2d ( images , filters , strides = 1 , padding = "SAME" )
plt . imshow ( outputs [ 0 , :, :, 1 ], cmap = "gray" )
plt . show ()
# ### Notes
# - when valid padding is used: the sides are not padded, some rows may get ignored in case of stride not equal to 1
# - when same paddding is used, left & right may be padded so that no rows get ignored
# Started with chapter 10. Some references were there in chapter 14
# **Perceptron**
# 
# +
import numpy as np
from sklearn.datasets import load_iris
from sklearn.linear_model import Perceptron
iris = load_iris()
# selected column 2 & 3
X = iris.data[:, (2, 3)] # petal length, petal width
y = (iris.target == 0).astype(np.int) # Iris setosa?
per_clf = Perceptron()
per_clf.fit(X, y)
y_pred = per_clf.predict([[2, 0.5]])
# -
print(per_clf.predict([[4, 2.5]]))
iris.target
iris.data[:,0:3]
# The most common step function used in Perceptrons is the Heaviside step function
# 
| A14_deep_computer_vision_with_cnns.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import batoid
import numpy as np
import matplotlib.pyplot as plt
from ipywidgets import interact
import ipywidgets as widgets
# %matplotlib inline
# +
# Let's model my 12-inch f/5 Newtonian reflector.
# 12-inch is approx 0.3 m
# f/5 then means the focal length is 1.5 m
# Which then means the parabolic equation is:
# z = (x^2+y^2) / (4*1.5)
# In the other notebook, we just used a Paraboloid
# Let's try an Asphere here though, to see what happens
# When the conic constant is not exactly -1, and when
# the higher order aspheric constants are non-zero.
D = 0.3
fn = 5
f = D * fn # focal length
R = 2*f
def makeTelescope(defocus=0.0): # defocus in mm
telescope = batoid.CompoundOptic(
items = [
batoid.Mirror(
batoid.Asphere(R, -0.95, [1e-6, 1e-12]),
name="Mirror"
),
batoid.Detector(
batoid.Plane(),
name="detector",
coordSys=batoid.CoordSys(origin=[0,0,f+0.001*defocus])
)
]
)
return telescope
plate_scale = 1./f # radians / m approximate
# -
@interact(theta_x=widgets.FloatSlider(min=-1,max=1,step=0.01,value=0.1),
theta_y=widgets.FloatSlider(min=-1,max=1,step=0.01,value=-0.3),
focus=widgets.FloatSlider(min=-0.5, max=0.5, step=0.01,value=0.1))
def spot(theta_x, theta_y, focus):
"""Display a spot diagram for a Newtonian telescope.
@param theta_x Field angle in degrees
@param theta_y Field angle in degrees
@param focus Defocus distance in mm
"""
telescope = makeTelescope(focus)
rays = batoid.RayVector.asPolar(
backDist=1.0, nrad=40, naz=160, wavelength=500e-9,
theta_x=np.deg2rad(theta_x), theta_y=np.deg2rad(theta_y),
outer=D/2
)
telescope.trace(rays)
w = ~rays.vignetted
x, y = np.array(rays.x[w]), np.array(rays.y[w])
x -= np.mean(x)
y -= np.mean(y)
x *= plate_scale*206265
y *= plate_scale*206265
plt.figure(figsize=(4.5,4))
plt.scatter(x, y, s=1, alpha=0.5)
plt.xlim(-10, 10)
plt.ylim(-10, 10)
plt.title(r"$\theta_x = {:4.2f}\,,\theta_y = {:4.2f}\,, f={:4.2f}$".format(theta_x, theta_y, focus))
plt.xlabel("arcseconds")
plt.ylabel("arcseconds")
# +
| notebook/Aspheric Newtonian Telescope.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ### Importing necessary dependencies
# +
from glob import glob
import os
import cv2
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
# %matplotlib inline
# -
# ### Data directory
DATA_DIR = 'data'
# ## Extending Data Set
# +
for sample_dir in ['rosbag', 'rosbag1', 'rosbag2']:
for color, color_dir in enumerate(['green', 'yellow', 'red', 'none']):
data_dir = os.path.join(DATA_DIR, sample_dir, color_dir)
img_paths = glob(os.path.join(data_dir, '*.jpg'))
print('Processing: {}, number of images {}'.format(data_dir, len(img_paths)))
for img_path in img_paths:
file_name = img_path.split('/')[-1]
img = cv2.imread(img_path)
cv2.imwrite(os.path.join(DATA_DIR, 'ext_' + sample_dir, color_dir, file_name), cv2.flip(img, 1))
# -
# ## Verifying extension
for sample_dir in ['rosbag', 'ext_rosbag', 'rosbag1', 'ext_rosbag1', 'rosbag2', 'ext_rosbag2']:
for color, color_dir in enumerate(['green', 'yellow', 'red', 'none']):
data_dir = os.path.join(DATA_DIR, sample_dir, color_dir)
img_paths = glob(os.path.join(data_dir, '*.jpg'))
print('Dir: {}, number of images {}'.format(data_dir, len(img_paths)))
| Data_Extension.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # PDE-Learning of non-interacting fermions dynamics: Learning continuity and Euler Equations
# +
# %pylab inline
# %load_ext autoreload
# %autoreload 2
import sys
import time
import numpy as np
sys.path.insert(0, "../")
from PDE_search import *
from utils import save_dict, load_dict
from utils import generalized_euler_solver, get_euler_term_from_descr
import matplotlib.pyplot as plt
# -
# # Here we consider fermion dynamics in 1D tight-binding model with the Hamiltonian:
# $$ H = -J \sum_i (c^{+}_i c_{i+1} + c^+_{i+1} c_i),$$
# where $J$ is the hopping coefficient, $c^+_i$ ($c_i$) is the creation (annihilation) fermion operator.
#
# The physical observables of interest are the particle density $\rho(t,i) = \langle c^+_ic_i\rangle$ and velocity $v(t,i) = 2J\Im \langle c^+_{i+1} c_i\rangle/\rho(t,i)$.
# ## First, let's have a look at the dataset
# +
#-------------------------------------------------------
# Import data
fname ='./data/free_fermions_J1=-0.5_J2=0.0_L=1000.npy'
data_dict = load_dict(fname)
#-------------------------------------------------------
# Specifying time range
start, stop = 0, 1500
# Density
rho = data_dict['n']
# Velocity
v = data_dict['v']
# Temporal and Spatial points
t = data_dict['t']
x = data_dict['x']
# Plot data
xx, tt = np.meshgrid(x, t)
fig, ax = plt.subplots(1, 2, figsize = (12, 5));
plt_rho = ax[0].pcolormesh(xx, tt, rho, cmap='coolwarm')
ax[0].set_xlabel(r'$x$')
ax[0].set_ylabel(r'$t$')
ax[0].set_title(r'density, $\rho(t, x)$')
plt_v = ax[1].pcolormesh(xx, tt, v, cmap='coolwarm')
ax[1].set_xlabel(r'$x$')
ax[1].set_ylabel(r'$t$')
ax[1].set_title(r'velocity, $v(t, x)$')
fig.colorbar(plt_v, ax=ax[1])
fig.tight_layout(pad=2)
# -
dt = t[1]-t[0]
dx = x[1]-x[0]
m, n = rho.shape
print('dt: ', dt, 'dx: ', dx)
print('Dataset dimensions: ', rho.shape)
print('Xmax, Tmax: ', x[-1], t[-1])
# ## Preparing library of candidate terms:
# +
#-----------------------------------------------------------------------------
# Define symbolic expressions for the candidate terms in v_t=G(...)
#-----------------------------------------------------------------------------
v_descr = [
'rho_x',
'v*v_x',
'v^2*rho_x',
'v^2*rho*rho_x',
'v^2*rho^2*rho_x',
'v^2*rho^3*rho_x',
'v^2*rho^4*rho_x',
'v^2*rho^5*rho_x',
'v^2*rho_x*1/rho',
'rho*v*v_x',
'rho^2*v*v_x',
'rho^3*v*v_x',
'rho^4*v*v_x',
'rho^5*v*v_x',
'rho*rho_x',
'rho^2*rho_x',
'rho^3*rho_x',
'rho^4*rho_x',
'rho^5*rho_x',]
#-----------------------------------------------------------------------------
# Define boundary conditions, needed for evaluation of spatial derivatives
bc = "periodic"
#-----------------------------------------------------------------------------
# Compute temporal and spatial derivatives and build Theta(U) matrix
rhot = TotalFiniteDiff_t(rho, dt).reshape((n*m,1))
vt = TotalFiniteDiff_t(v, dt).reshape((n*m,1))
#-----------------------------------------------------------------------------
v_data = []
# Stack columns with the vectorized candidate terms to form matrix Theta(U)
for term in v_descr:
v_data.append(get_euler_term_from_descr(term, rho, v, x, bc).reshape((n*m,1)))
v_data = np.hstack(v_data)
v_Theta, v_descr = build_custom_Theta(v_data, v_descr)
# +
#-----------------------------------------------------------------------------
# Define symbolic expressions for the candidate terms in rho_t=F(...)
#-----------------------------------------------------------------------------
rho_descr = [ 'rho',
'v',
'rho_x',
'v_x',
'v*v_x',
'rho*rho_x',
'v*rho_x',
'rho*v_x',
'rho*v*rho_x',
'rho^2*v_x',
'rho^2*v*rho_x',
'rho^3*v_x',
'rho^3*v*rho_x',
'rho^4*v_x',
]
rho_data = []
# Stack columns with the vectorized candidate terms to form matrix Theta(U)
for term in rho_descr:
rho_data.append(get_euler_term_from_descr(term, rho, v, x, bc).reshape((n*m,1)))
rho_data = np.hstack(rho_data)
rho_Theta, rho_descr = build_custom_Theta(rho_data, rho_descr)
# -
print(r"Total number of terms for $rho_t=F(.)$:", len(rho_descr))
print(r"Total number of terms for $v_t=G(.)$:", len(v_descr))
# ## Search PDE for density: $\rho_t = F(...)$
xi_rho, _ = BruteForceL0(rho_Theta, rhot, rho_descr, l0_penalty=1e-3, lam_l2=0,
verbose=False, lhs_descr='rho_t')
# We receovered the continuity equation!
# ## Search PDE for velocity: $v_t = G(...)$
# Computation may take a while (~10 mins)
xi_v, _ = BruteForceL0(v_Theta, vt, v_descr, l0_penalty=1e-3, lam_l2=0, verbose=False, lhs_descr='v_t')
# The discovered system of equations is very close to the semiclassical equations of motion for the ideal Fermi gas:
# $$\begin{cases}
# \rho_t+(\rho v)_x = 0,\\
# v_t+v v_x + \frac{\pi^2}{m^2}\rho \rho_x = 0,
# \end{cases}
# $$
# where $m=1/(2J)=1$ is the fermion mass.
# ## Compare data and the solution of the inferred hydrodynamic PDE
rho_ev, v_ev = generalized_euler_solver(v_descr, xi_v.real,
rho[0], v[0], t, x,
num_integrator_steps=50,
fix_vvx_term=False)
# +
time_points = [0, 250, 500, -1]
for i, indx in enumerate(time_points):
if i == 0:
label1, label2 = 'data', 'PDE'
else:
label1, label2 = '', ''
ax[0].plot(x, rho[indx, :],
c='royalblue', label=label1)
ax[0].plot(x, rho_ev[indx,:],
ls='--', c='red', dashes=(4,3), label=label2)
ax[1].plot(x, v[indx, :],
c='royalblue', label=label1)
ax[1].plot(x, v_ev[indx,:],
ls='--', c='red', dashes=(4,3), label=label2)
ax[0].legend(loc="center left", bbox_to_anchor=(1, 0.5))
ax[0].set_xlabel(r'$x$')
ax[0].set_title(r'$\rho(x, t)$')
ax[1].legend(loc="center left", bbox_to_anchor=(1, 0.5))
ax[1].set_xlabel(r'$x$')
ax[1].set_title(r'$v(x, t)$')
fig.tight_layout(pad=3)
# -
| free_fermions/.ipynb_checkpoints/free_fermions_quadratic-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ################################################################# #<br>
# Basic plot for two-strain SIR model:<br>
# Bifurcation diagram for one parameter<br>
# ##################################################################
import sys
import numpy as np
import matplotlib as mpl
from matplotlib.font_manager import FontProperties
import matplotlib.pyplot as plt
from two_strain import *
# Run parameters
run_num = 1 # sys.argv[1]
end_time = 1000*365
output_interval = 365.0 # if not 365., need to adjust strobe interval
step_size = 1.0
sweep_par = "beta[0]" # e.g., "beta[0]", "a[1]", "alpha[0]"
par_min = 1.0/7.0
par_max = 7.0/7.0
n_points = 40 # number of points in parameter range
n_strobes = 50 # number of years to sample
# Strain parameters, including initial conditions
beta = np.array([5, 5])/7.0
epsilon = 0.1
gamma = np.array([1, 1])/7.0
mu = 1/(10*365.0)
alpha = np.array([1., 1.])
a = np.array([1., 1.])
omega = 2*np.pi/365.
obs_sd = 0.01
NSS = 0.2
NIS = 1e-3
NRS = 0.02
NRI = 0.0
NSI = 1e-3
NSR = 0.02
NIR = 0.0
# Organize and run simulations
SI = np.array([NSS, NIS, NRS, NRI, NSI, NSR, NIR], dtype="object")
ic = np.array([NSS, NIS, NRS, NRI, NSI, NSR, NIR, 1-np.sum(SI)], dtype="object")
par_vals = np.linspace(par_min, par_max, n_points)
bif_vals = np.zeros((len(par_vals), n_strobes))
for i in range(len(par_vals)):
print('Running value %d of %d' % (i+1,len(par_vals)))
exec(sweep_par + " = par_vals[i]")
params = np.array([gamma, mu, alpha, a, omega, beta, epsilon], dtype="object")
output = run_two_strain(end_time, output_interval, step_size, params, ic)
I = output[:, 1] + output[:, 6] # NIS + NIR
bif_vals[i, :] = I[-n_strobes:len(I)]
# Plot output
plt.plot(par_vals, bif_vals, '.k')
plt.xlabel(sweep_par)
plt.ylabel("NIS + NIR")
plt.xlim([par_min, par_max])
plt.show()
plt.savefig("bifurcation_" + sweep_par + ".png")
plt.close()
| sismid-SISMID2021/models/exercise/plot_two_strain_bd.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + deletable=true editable=true
import numpy as np
import multiprocessing as mp
from functools import partial
from matplotlib import pyplot as plt
# + deletable=true editable=true
"""
Gera uma reta aleatória
"""
def generate_line():
# Dimensões do poblema
d = 2
# Gera 2 pontos aleatórios entre 0 e 1 e os converte para
# o intervalo entre -1 e 1
p1 = np.random.rand(d) * 2 - 1
p2 = np.random.rand(d) * 2 - 1
# Calcula a reta na forma `y=ax + b` entre os pontos
a = (p2[1] - p1[1]) / (p2[0] - p1[0])
b = p1[1] - a*p1[0]
return np.array([b, a, -1])
"""
Gera os dados baseados em uma reta
"""
def generate_data(line, n):
# Gera 'n' pontos (x, y) aleatoriamente entre 0 e 1
# e os converte para o intervalo -1 e 1
points = (np.random.rand(n, 2) * 2) - 1
return points
"""
Calcula os valores de saída da função ideal
"""
def calc_y(line, data):
y = line[0] + np.dot(data, line[1:])
# Transforma os valores em -1 ou 1
# Valores em cima da reta são considerados -1
y = np.where(y > 0, 1, -1)
return y
# + deletable=true editable=true
"""
Executa o perceptron para um conjunto de dados e de pesos iniciais
Caso os pesos não sejam especificados, eles serão o vetor nulo
"""
def perceptron(data, y, weights=None):
if weights is None:
# Cria um vetor de zeros do tamanho da quantidade de dimensões + viés
weights = np.zeros(data.shape[1]+1 if len(data.shape) > 1 else 1)
# Contador de iterações
it = 0
# Adiciona a coluna do viés ao X
m_data = np.concatenate((np.ones((data.shape[0], 1)), data), axis=1)
# Vetor que indica quais pontos estão classificados erroneamente
# Inicialmente começa com 1(True) para todos os valores,
# pois nenhum dado foi classificado
miscl_mask = np.ones(m_data.shape[0], dtype=bool)
# Filtra os dados para que não foram classificados
miscl_data = m_data[miscl_mask]
# Executar até que não haja mais pontos classificados erroneamente
while miscl_data.shape[0] > 0:
# Filtra os dados de saída não classificados
miscl_y = y[miscl_mask]
# Seleciona um ponto aleatório dentro o conjunto dos não classificados
i = np.random.randint(miscl_data.shape[0])
# Atualiza os pesos
weights += miscl_y[i] * miscl_data[i]
# Calcula as saídas com os novos pesos
temp = np.sign(np.dot(m_data, weights))
# Atualiza os pontos não classificados
# Caso os valores calculados sejam diferentes dos corretos
# a subtração vai dar um valor diferente de 0
# que quando convertido para um tipo 'bool' valerá 'True'
# ou seja, classificado erroneamente
miscl_mask = np.array( temp - y, dtype=bool)
# Atualiza os dados classificados erroneamente
miscl_data = m_data[miscl_mask]
# Atualiza a iteração do algoritmo
it += 1
# Retorna os pesos finais e o número de iterações
return weights, it
# + deletable=true editable=true
def lin_regression(x, y):
m_x = np.concatenate((np.ones((x.shape[0], 1)), x), axis=1)
x_dagger = np.dot( np.linalg.inv( np.dot(m_x.T , m_x) ), m_x.T)
w_lin = np.dot(x_dagger, y)
return w_lin
# + deletable=true editable=true
def calc_error(line, w):
# Gera mil pontos para serem avaliados
ev_data = generate_data(line, 1000)
# Saída ideal
ev_f_y = calc_y(line, ev_data)
# Saída gerada pela g(x)
ev_g_y = calc_y(w, ev_data)
# Conta todos os pontos em que as saídas não foram iguais
misclassified = np.count_nonzero(ev_f_y - ev_g_y)
# Calcula a porcentagem dos pontos classificados erroneamente
return misclassified / ev_data.shape[0]
def experiment(N):
np.random.seed()
line = generate_line()
x = generate_data(line, N)
y = calc_y(line, x)
w_lin = lin_regression(x, y)
y_g = calc_y(w_lin, x)
e_in = np.count_nonzero( y - y_g ) / y.shape[0]
e_out = calc_error(line, w_lin)
w_perc, it = perceptron(x, y, weights=w_lin)
return [e_in, e_out, it]
# + deletable=true editable=true
"""
Executa um certo número de experimentos paralelamente
Caso o número de processos não seja espeficidado,
o multiprocessing utiliza o valor padrão,
que costuma ser o número de processadores
"""
def run_experiment(N, num_exp, processes=None):
pool = mp.Pool(processes)
# Executa os experimentos 'num_exp' vezes, passando como
# parâmetro para cada um, o número de dados N a serem gerados
results = np.array(pool.map(experiment, [N] * num_exp))
pool.close()
# Calcula a média dos resuldados por coluna (iterações, erro)
return np.mean(results, axis=0)
# + deletable=true editable=true
result_1, result_2 = run_experiment(100, 1000)[:2]
result_3 = run_experiment(10, 1000)[2]
print('Question 1:', result_1)
print('Question 2:', result_2)
print('Question 2:', result_3)
| trabalhos/trabalho2/trabalho2_linear.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python for Performance
#
# Python is not a fast language. Compared to other mainstream programming languages like Java, C#, Go, Javascript, C++, etc, Python is 2-10x slower on executing comparable tasks. If that's the case, why is Python so popular in applications with heavy number-crunching aspects like scientific computing, data science, and machine learning?
#
# The answer, of course, is that Python is fun and easy to program in and requires little boilerplate code compared to these other languages. But, more importantly, **it doesn't matter that Python is slow** because it can be extended using _"Native Modules"_ which are written in these other fast languages and can be used as though they were pure Python.
#
# Some of you may be using these modules already! NumPy and Pandas are among the most popular Python packages, and both rely on C and even FORTRAN to speed up the most peformance-critical paths of their libraries. Machine learning libraries like Tensorflow are also largely written in C and C++ for speed, but also to access low-level features in the CPU and GPU. Even Python's comprehensive standard library implements many functions in C. Native modules are an important part of how Python has remained relevant in this brave new big-data world.
#
#
# ## Why is Python so Slow?
#
# Python is an interpreted language, which means the code is read directly and freely, without first compiling and optimizing programs into machine instructions. In addition to that, Python is dynamically typed and garbage collected, which means Python figures out what kind of data type variables are and manages memory for the user. All these features make things easier for the programmer, but require processing power that would otherwise get used to complete the task.
#
#
# ## Demo Time!
#
# As an example of Python's speed, let's calculate a large dot product to show how dramatic the difference can be.
# +
import time
import numpy as np
# One Hundred MILLION dimensions...muahahahaha
v1 = np.random.rand(1, 100000000)
v2 = np.random.rand(1, 100000000)
start_time = time.time()
dot = 0
for x, y in zip(np.nditer(v1), np.nditer(v2)):
dot = dot + (x*y)
end_time = time.time()
python_time = end_time - start_time
print(f"That took {python_time} seconds")
# +
start_time = time.time()
better_dot = sum(x * y for x, y in zip(np.nditer(v1), np.nditer(v2)))
end_time = time.time()
better_python_time = end_time - start_time
print(f"That took {better_python_time} seconds")
std_multiple = python_time/better_python_time
print(f"\nThe standard lib code was {std_multiple} times faster than naive Python code")
# +
start_time = time.time()
best_dot = np.dot(v1, v2.T)
end_time = time.time()
best_python_time = end_time - start_time
print(best_python_time)
multiple = min(python_time, better_python_time)/best_python_time
print(f"\nThe numpy code was {multiple} times faster than pure Python code")
print("\n\n\n░░░░░░░░░▄▄▄▄▄\n░░░░░░░░▀▀▀██████▄▄▄\n░░░░░░▄▄▄▄▄░░█████████▄\n░░░░░▀▀▀▀█████▌░▀▐▄░▀▐█\n░░░▀▀█████▄▄░▀██████▄██ <GOTTA GO FAST!\n░░░▀▄▄▄▄▄░░▀▀█▄▀█════█▀\n░░░░░░░░▀▀▀▄░░▀▀███░▀░░░░░░▄▄\n░░░░░▄███▀▀██▄████████▄░▄▀▀▀██▌\n░░░██▀▄▄▄██▀▄███▀░▀▀████░░░░░▀█▄\n▄▀▀▀▄██▄▀▀▌████▒▒▒▒▒▒███░░░░▌▄▄▀\n▌░░░░▐▀████▐███▒▒▒▒▒▐██▌\n▀▄░░▄▀░░░▀▀████▒▒▒▒▄██▀\n░░▀▀░░░░░░▀▀█████████▀\n░░░░░░░░▄▄██▀██████▀█\n░░░░░░▄██▀░░░░░▀▀▀░░█\n░░░░░▄█░░░░░░░░░░░░░▐▌\n░▄▄▄▄█▌░░░░░░░░░░░░░░▀█▄▄▄▄▀▀▄\n▌░░░░░▐░░░░░░░░░░░░░░░░▀▀▄▄▄▀")
# -
# ## Woah...What Happened?
#
# So by now you're probably saying "Wow, Numpy is a crazy fast! ...what gives?" You're probably going to guess what comes next: It's a native module! According to the GitHub repo half of it is written in C! Moreover, that C code uses optimizations that are only available in programming languages that are close to the metal, such as densely packed homogenously-typed arrays and CPU-level hardware-accelerated SIMD instructions such as AVX vector operations which Python doesn't have much use for in its day-to-day life as a programming language.
#
# However, peformance was never necessarily a key requirement of Python's original design! When Python was first hatched it was a bash shell script stand-in for sysadmin tasks on a niche operating system called *Amoeba*, so it was intended more as a tool to describe jobs, organize information, and glue together specialized programs. In a way, Python is still doing the same thing when it calls on native modules like Numpy--it describes work for faster code!
#
#
# ## How to Write Fast Python
#
# All this implies that perhaps the way to write performant Python is to write as little Python as possible. As mentioned before, Python is designed to offload the number crunching to programs and applications that are better suited for it. For some advanced users, this might mean [making your own native modules](https://docs.python.org/3/extending/building.html) (which we can discuss in a future lesson if there is popular demand) but for most people trying to get performance out of Python, it's best to look up performance-oriented libraries and standard lib functions.
#
# Either way, the pattern will remain the same: Get your tasks onto fast, optimized code as soon as possible and keep the data there for as long as you can.
# # A Short Tour of Some Fast Modules
#
# To get you started, we'll show some examples of popular packages that make use of native modules to perform common Python tasks.
#
#
# ## Numpy
#
# `pip install numpy`
#
# As demonstrated above, Numpy is fast...really fast. It's the flagship library in the NumFocus ecosystem, which works on keeping Python relevant in the scientific computing space, which means crunching lots of numbers. Many performance-oriented Python libraries are built on Numpy.
#
# The key to Numpy's speed is vectorized data and linear algebra, which are easy to accelerate in hardware. That means that there is a bit of a learning curve to get the most of this library, but it also means you can operate on huge amounts of data hundreds of times faster than you could in pure Python.
#
#
# ## Pandas
#
# `pip install pandas`
#
# Pandas brings R dataframes to Python--or basically creates a database-like structure in memory that can be queried based on rows and column projections. Pandas is a particularly great tool for importing and cleaning data from csv, json objects, and other sources before feeding them into other libraries.
#
# Though the GitHub repo says Pandas is 93% Python, don't be fooled! Pandas is built on top of Numpy and benefits from a lot of its speed optimizations, along with writing its own critical code paths in C and C++.
#
#
# ## SciPy
#
# `python -m pip install --user numpy scipy matplotlib ipython jupyter pandas sympy nose`
#
# SciPy is the last NumFocus library we'll discuss here. While Numpy and Pandas are more to contain and wrangle data, SciPy is a library for doing analysis. Cumulatively, about half of it is written in low-level languages, includling about 23% in Fortran, which is the grandpappy of all scientific programming languages and is still used often in the scientific community due to its high performance and great math algorithms. If you're looking for things like fourier transforms, this is one of your best bets.
#
# ## Tensorflow
#
# `pip install tensorflow`
# `pip install tensorflow-gpu`
#
# Tensorflow is probably one of the most famous machine learning frameworks out there, and was one of the tools that put Python on the map as a premiere tool for deep learning practitioners. Deep learning in particular is a famously compute-intensive technique which was only recently made feasible in the mainstream due to the advancements in GPU technology driven by the video games industry. Basically, deep learning algorithms typically use vectors, and GPUs are basically big vector accelerators.
#
# Tensorflow is 61% C++, but of particular interest is its use of the CUDA API, which allows it to offload computation to the GPU. Similar to how you want your data to spend as little time as possible in Python for speed, Tensorflow tries to let the data spend as much time as possible on the GPU for learning, since the jump between hardware components is time-intensive on the scale of computation. All this gives Python deep learning superpowers that the language couldn't have on its own.
# # How to Evaluate Packages for Speed
#
# A good rule of thumb is to use GitHub's language percentage feature to see if a library is wired for speed. While this isn't 100% reliable as native code can still be written badly--and as seen in Pandas, a language breakdown can be decieving--seeing low-level languages designed for speed in a Python package is a good indicator the authors care about performance and have done some work to increase their package's speed. You can also check dependences to see if any of those are fast, such as is the case with Pandas, which largely uses Numpy, which itself is very fast.
#
# You can see the language breakdown of a GitHub repo by clicking the colored bar below the top-level statistics, which reveals language percentages.
#
# 
#
# In general you will be looking for compiled languages, the most popular of which include C and C++, which have bindings for Python native modules. Rust is another up-and-coming fast language, and Fortran comes up occasionally due to its history as a language for scientific computing. Beyond that, there are some fast languages that work on the JVM such as Java and Scala, though those are less common in the Python ecosystem.
| Python for Performance.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# デバイス毎に音楽の再生音量の最大値は異なる.騒音の大きさから,再生音量を実数値( $0 \leq x \leq 1.0$ )で求める関数があると便利.
# デバッグ用 SC-04J 上でAudioRecoderで計測した音量の値(騒音)と,その時に設定した音楽の再生音量を記録.
# 適当な関数にフィットさせて実数を返す関数を作る.
# +
# 実際の測定結果を元に,擬似データを作成する関数
def labelDispenser(x: int) -> int:
if x < 0:
raise ValueError("負値")
if x < 750:
return 0
elif x < 3750:
return 1
elif x < 9750:
return 2
elif x < 20000:
return 3
elif x < 30000:
return 4
elif x <= 32767:
return 5
else:
raise ValueError("大きすぎ")
# 再生音量は SC-04J では15段階中での整数値であり,これを 0 から1.0 までの実数値に射影する
x = np.arange(0,32767,10)
y = np.frompyfunc(labelDispenser,1,1)(x)
df = pd.DataFrame({"騒音":x, "再生音量":y})
df["再生音量(実数値)"] = df["再生音量"] / 15
# -
# 騒音値が10^3オーダ,再生音量が10^-1オーダなので,オーダを再生音量側に揃える
# プロット
x = df.loc[:,"騒音"] / 100000
y = df.loc[:,"再生音量(実数値)"]
plt.scatter(x, y)
plt.xlabel("env_noise")
plt.ylabel("play_level")
print("騒音とそのときの再生音量の対応イメージ")
# この段々になっているグラフを滑らかにしたような関数が欲しい
# +
# 再生音量の変化位置(境界)に注目
x = np.array([750, 3750, 9750, 20000, 30000]) / 100000
y = np.array([0.5, 1.5, 2.5, 3.5, 4.5]) / 15
# a + bx にフィット
from sklearn.linear_model import LinearRegression
lr = LinearRegression().fit(np.array(x).reshape(-1, 1), y)
print(lr.score(np.array(x).reshape(-1, 1),y), lr.intercept_, lr.coef_)
# a + bx + cx^2 にフィット
lr2 = LinearRegression().fit(np.dstack((np.power(np.array(x),2),np.array(x)))[0], y)
print(lr2.score(np.dstack((np.power(np.array(x),2),np.array(x)))[0],y), lr2.intercept_, lr2.coef_)
# a + b log(x) にフィット
log_r = LinearRegression().fit(np.array(np.log(x[1:])).reshape(-1, 1), y[1:])
print(log_r.score(np.array(np.log(x[1:])).reshape(-1, 1),y[1:]), log_r.intercept_, log_r.coef_)
# a + b sqrt(x) にフィット
lr3 = LinearRegression().fit(np.array(np.sqrt(x)).reshape(-1, 1), y)
print(lr3.score(np.array(np.sqrt(x)).reshape(-1, 1),y), lr3.intercept_, lr3.coef_)
# -
# グラフ描写
RNG = np.linspace(0, 0.32, 100)
plt.figure(figsize=(18,9))
plt.scatter(x, y, label="data")
plt.xlabel("env_noise")
plt.ylabel("play_level")
plt.plot(RNG, lr.intercept_ + lr.coef_ * RNG, label="a+bx", color="green")
plt.plot(RNG, lr2.intercept_ + lr2.coef_[1] * RNG + lr2.coef_[0] * RNG * RNG, label="a+bx+cx^2", color="red")
plt.plot(RNG, lr3.intercept_ + lr3.coef_ * np.sqrt(RNG), label="a+ b sqrt(x)", color="purple")
plt.plot(RNG, log_r.intercept_ + log_r.coef_ * np.log(RNG), label="a+ b log(x)", color="cyan")
plt.legend(loc='upper left')
# +
# 騒音から再生音量を求める関数を用いて,実際にデバイスの再生音量を決定してみる
# 関数の値を四捨五入して整数化
RNG = np.linspace(0.001, 0.32, 500)
DIV_NUM = 15 # デバイス毎に異なる,再生音量の最大値
plt.figure(figsize=(18,12))
plt.scatter(df["騒音"]/100000, df["再生音量"], label="data")
plt.plot(RNG, np.round(DIV_NUM * (lr.intercept_ + lr.coef_[0] * RNG)), label="a+bx round", color="green")
plt.plot(RNG, np.round(DIV_NUM * (lr2.intercept_ + lr2.coef_[1] * RNG + lr2.coef_[0] * RNG * RNG)), label="a+bx+cx^2 round", color="red")
plt.plot(RNG, np.round(DIV_NUM * (lr3.intercept_ + lr3.coef_ * np.sqrt(RNG))), label="a+ b sqrt(x)", color="purple")
plt.plot(RNG, np.round(DIV_NUM * (log_r.intercept_ + log_r.coef_ * np.log(RNG))), label="a+ b log(x)", color="cyan")
plt.legend(loc='upper left')
# -
# イメージとのギャップが最も小さい $ y = a + b \sqrt{x} $ を実装する
| notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Sobloo Sentinel-1 L1 SLC (SAFE)
# Sentinel-1 L1 SLC (SAFE) provides full scenes of Sentinel-1 (A/B)'s C-band synthetic aperture radar sensor in processing level L-1C Single Look Complex (SLC) in SAFE folder structure.
#
# In the example, the workflow, the area of interest and the workflow parameters are defined. After running the job, the results are downloaded and visualized. For more information, refer to the block's [UP42 Marketplace page](https://marketplace.up42.com/block/222c9742-ed19-4a2e-aa87-d6b58193cd31) and [Documentation](https://docs.up42.com/up42-blocks/data/sobloo-s1-slc.html).
import up42
# +
# add credentials and authenticate
up42.authenticate(project_id="12345", project_api_key="12345")
project = up42.initialize_project()
# Construct workflow
workflow = project.create_workflow(name="sobloo-s1-l1-workflow", use_existing=True)
# -
input_tasks = ["sobloo-s1-slc-fullscene"]
workflow.add_workflow_tasks(input_tasks)
# Define the aoi
barcelona_example_aoi = {"type":"Feature","properties":{},
"geometry":{"type": "Polygon","coordinates": [[[2.109314,41.394249],
[2.241211,41.391234],
[2.241211,41.327326],
[2.083707,41.326001],
[2.109314,41.394249]]]}}
# Other options to define the aoi include up42.draw_aoi(), up42.read_vector_file(), FeatureCollection, GeoDataFrame etc.
# Get input parameters
workflow.get_parameters_info()
# Define input parameters of the workflow to run it
input_parameters = workflow.construct_parameters(geometry=barcelona_example_aoi,
geometry_operation="intersects",
start_date="2018-01-01",
end_date="2021-12-31",
limit=1)
# In the above code cell, we added the basic input parameters. However, the block accepts additional parameters as per workflow.get_parameters_info(). You can add the parameters to our parameter configuration in the following steps:
input_parameters["sobloo-s1-slc-fullscene:1"].update({"mission_code": None,
"orbit_direction": None,
"acquisition_mode": None,
"orbit_relative_number": None})
# Check if everything is as expected
print(input_parameters)
# Price estimation
workflow.estimate_job(input_parameters)
# Run a test job to query data availability and check the configuration
test_job = workflow.test_job(input_parameters, track_status=True)
# Run the actual job
job = workflow.run_job(input_parameters, track_status=True)
# Please note that this job may take up to 1 hour to finish
# Download and plot results
job.download_results()
# + tags=[]
job.plot_quicklooks(filepaths="project_810e8039-cb5c-445b-b08d-53b0dde1bde0/job_02f371a3-6b74-4a00-b8d1-61332896d06e/679b09a8-94da-430a-ab96-57f110bd1252/S1B_IW_SLC__1SDV_20210305T174607_20210305T174634_025883_031642_A629.SAFE/preview/quick-look.png")
| examples/data-block-examples/sobloo-sentinel1-l1-example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Tulasi-ummadipolu/LetsUpgrade-Python-B7/blob/master/Day3Assignment2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="bY_YwH13GK-1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 799} outputId="8eaade74-d019-48dd-d6ca-fc678a285d3f"
for num in range(1,201):
if (num>1) :
for i in range(2,num):
if (num%i)==0:
break
else:
print(num)
| Day3Assignment2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
def create_slice(index_names,*args,**kwargs):
return tuple(map(lambda indx:kwargs.get(indx,slice(None)) ,index_names))
index_names = ["index_1","index_2"]
create_slice(index_names,index_1="3")
import pandas as pd
@pd.api.extensions.register_dataframe_accessor("fidx")
class FastIndex(object):
def __init__(self, df):
self._validate(pandas_obj)
self._df = df
@staticmethod
def _validate(obj):
if 'lat' not in obj.columns or 'lon' not in obj.columns:
raise AttributeError("Must have 'lat' and 'lon'.")
@property
def f_slice(self,**kwargs):
# return the geographic center point of this DataFrame
return create_slice(df.index.names,**kwargs)
def plot(self):
# plot this array's data on a map, e.g., using Cartopy
pass
| .ipynb_checkpoints/building the api-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Networkx example
#
# This Example will show you step by step, how to use networkx graphs with a PHOTONAI Graph pipeline.
#
# First: Necessary imports
# + pycharm={"is_executing": true}
from networkx.generators import barabasi_albert_graph, erdos_renyi_graph
from photonai.base import Hyperpipe, PipelineElement
from sklearn.model_selection import KFold
import networkx as nx
import numpy as np
# -
# Next: Generate graphs and labels
# + pycharm={"name": "#%%\n"}
# create dataset of 1000 graphs
rr_graphs = [erdos_renyi_graph(20, .25) for _ in range(500)]
er_graphs = [erdos_renyi_graph(20, .2) for _ in range(500)]
graphs = rr_graphs + er_graphs
# we have to transform the networkx graphs into numpy graphs before passing them to photon
graphs = np.array([nx.to_numpy_array(g) for g in graphs])
graphs = np.expand_dims(graphs, axis=-1)
# create labels
rr_labels = [0] * 500
er_labels = [1] * 500
labels = rr_labels + er_labels
# -
# Setup Hyperpipe
# + pycharm={"name": "#%%\n"}
# Design your Pipeline
my_pipe = Hyperpipe('networkx_example_pipe',
inner_cv=KFold(n_splits=2),
optimizer='sk_opt',
optimizer_params={'n_configurations': 25},
metrics=['accuracy', 'balanced_accuracy', 'recall', 'precision'],
best_config_metric='accuracy')
my_pipe.add(PipelineElement('GCNClassifier', feature_axis=0))
# -
# Final step: Train your Hyperpipe
# + pycharm={"name": "#%%\n"}
my_pipe.fit(graphs, labels)
| documentation/docs/examples/networkx_pipeline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# This small example shows you how to access JS-based requests via Selenium
# Like this, one can access raw data for scraping,
# for example on many JS-intensive/React-based websites
#
import time
from selenium import webdriver
from selenium.webdriver import DesiredCapabilities
from selenium.webdriver.support.ui import WebDriverWait
from selenium.webdriver.support import expected_conditions as EC
from selenium.webdriver.common.by import By
from selenium.webdriver.firefox.options import Options
from selenium.common.exceptions import ElementClickInterceptedException
from selenium.common.exceptions import ElementNotInteractableException
from selenium.common.exceptions import WebDriverException
import json
from datetime import datetime
import pandas as pd
# +
def process_browser_log_entry(entry):
response = json.loads(entry['message'])['message']
return response
def log_filter(log_):
return (
# is an actual response
log_["method"] == "Network.responseReceived"
# and json
and "json" in log_["params"]["response"]["mimeType"]
)
# -
def init_page():
#fetch a site that does xhr requests
driver.get("https://www.youtube.com/watch?v=DWcJFNfaw9c")
main_content_wait = WebDriverWait(driver, 20).until(
EC.presence_of_element_located((By.XPATH, '//iframe[@id="chatframe"]'))
)
time.sleep(3)
video_box = driver.find_element_by_xpath('//div[@id="movie_player"]')
video_box.click()
frame = driver.find_elements_by_xpath('//iframe[@id="chatframe"]')
# switch the webdriver object to the iframe.
driver.switch_to.frame(frame[0])
try:
#enable 'all' livechat
try:
driver.find_element_by_xpath('//div[@id="label-text"][@class="style-scope yt-dropdown-menu"]').click()
except ElementNotInteractableException:
init_page()
time.sleep(2.1)
driver.find_element_by_xpath('//a[@class="yt-simple-endpoint style-scope yt-dropdown-menu"][@tabindex="-1"]').click()
except ElementClickInterceptedException:
print('let\'s try again...')
init_page()
# make chrome log requests
capabilities = DesiredCapabilities.CHROME
capabilities["goog:loggingPrefs"] = {"performance": "ALL"} # newer: goog:loggingPrefs
driver = webdriver.Chrome(
desired_capabilities=capabilities
)
init_page()
iter_num = 0
while True:
iter_num += 1
if iter_num >= 100:
iter_num = 0
init_page()
# extract requests from logs
logs_raw = driver.get_log("performance")
logs = [json.loads(lr["message"])["message"] for lr in logs_raw]
json_list = []
for log in filter(log_filter, logs):
request_id = log["params"]["requestId"]
resp_url = log["params"]["response"]["url"]
#print(f"Caught {resp_url}")
try:
if 'https://www.youtube.com/youtubei/v1/live_chat/get_live_chat?key=' in resp_url:
body = driver.execute_cdp_cmd("Network.getResponseBody", {"requestId": request_id})
json_list.append(body)
except WebDriverException:
print('web driver exception!!!')
continue
'''
with open('look.txt', 'a', encoding='utf-8') as text_file:
body = driver.execute_cdp_cmd("Network.getResponseBody", {"requestId": request_id})
text_file.write(str(body))
json_list.append(body)
'''
#print(len(json_list))
message_list = []
self_message_list = []
for i in range(len(json_list)):
json_data = json.loads(json_list[i]['body'].replace('\n','').strip())
try:
actions = (json_data['continuationContents']['liveChatContinuation']['actions'])
except:
continue
for j in range(len(actions)):
try:
item = actions[j]['addChatItemAction']['item']['liveChatTextMessageRenderer']
author_channel_id = item['authorExternalChannelId']
author_name = item['authorName']['simpleText']
text = item['message']['runs'][0]['text']
post_time = item['timestampUsec']
post_time = post_time[0:10]
post_time = int(post_time)
author_photo = item['authorPhoto']['thumbnails'][0]['url']
post_time = datetime.utcfromtimestamp(post_time)
post_item = {
"Author" : author_name,
"Message" : text,
"Date" : post_time,
"Channel ID" : author_channel_id,
"Channel" : f'https://youtube.com/channel/{author_channel_id}'
}
message_list.append(post_item)
if 'biss' in text.lower():
self_message_list.append(post_item)
#print(post_item)
except Exception as e:
print(str(e))
continue
#message_list = list(set(message_list))
df = pd.DataFrame(message_list)
df = df.drop_duplicates()
#print(df)
df.to_csv('./data/youtube_lofi/test_run.csv', index=False, mode='a')
reply_df = pd.DataFrame(self_message_list)
reply_df = reply_df.drop_duplicates()
if len(self_message_list) > 0 :
reply_df.to_csv('./data/youtube_lofi/reply_runs_cumulative.csv', index=False, mode='a')
reply_df.to_csv('./data/youtube_lofi/reply_runs.csv', index=False, mode='a')
if len(message_list) < 1:
print('The world is ending!')
time.sleep(30)
| message_reader.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h2>In-class transcript from Lecture 10, February 13, 2019</h2>
#
# # Imports and defs for lecture
# +
# These are the standard imports for CS 111.
# This list may change as the quarter goes on.
import os
import math
import time
import struct
import json
import pandas as pd
import networkx as nx
import numpy as np
import numpy.linalg as npla
import scipy
import scipy.sparse.linalg as spla
from scipy import sparse
from scipy import linalg
import matplotlib.pyplot as plt
from matplotlib import cm
from mpl_toolkits.mplot3d import axes3d
# %matplotlib tk
# -
def make_M_from_E(E):
"""Make the PageRank matrix from the adjacency matrix of a graph
"""
n = E.shape[0]
outdegree = np.sum(E,0)
for j in range(n):
if outdegree[j] == 0:
E[:,j] = np.ones(n)
E[j,j] = 0
A = E / np.sum(E,0)
S = np.ones((n,n)) / n
m = 0.15
M = (1 - m) * A + m * S
return M
np.set_printoptions(precision = 4)
# # Lecture starts here
E = np.load('PageRankEG1.npy')
E
indegree = np.sum(E,0)
indegree
outdegree = np.sum(E,1)
outdegree
n = E.shape[0]
n
d, V = linalg.eig(E)
d
E
sum(E,0)
A = E / np.sum(E,0)
A
d, V = linalg.eig(A)
d
v = V[:,0].real
v
eig_perm = np.argsort(v)[::-1]
eig_perm
A @ v
np.sum(A,0)
A.T
np.sum(A.T,0)
A.T @ np.array([1,1,1,1])
E
E[0,2] = 0
E
A = E / np.sum(E,0)
A
linalg.eig(A)
E
E[:,2] = np.ones(n)
E[2,2] = 0
E
A = E / np.sum(E,0)
A
d, V = linalg.eig(A)
d
v = V[:,0].real
eig_perm = np.argsort(v)[::-1]
eig_perm
E2 = np.load('PageRankEG2.npy')
E2
A = E2 / np.sum(E2,0)
A
n = A.shape[0]
d,V = linalg.eig(A)
d
m = .15
M = (1-m) * A + m * np.ones((n,n))/n
M
d,V = linalg.eig(M)
d
make_M_from_E(E2)
E3 = np.load('PageRankEG3.npy')
E3.shape
# %matplotlib inline
plt.spy(E3)
with open('PageRankEG3.nodelabels') as f:
labels = f.read().splitlines()
for i in range(10):
print(i, labels[i])
M = make_M_from_E(E3)
M.shape
d, V = linalg.eig(M)
d[0]
v = V[:,0]
perm = np.argsort(v)[::-1]
perm[:10]
for i in range(10):
print(i, labels[perm[i]])
E = sparse.load_npz('webGoogle.npz')
E.shape
d, V = spla.eigs(E)
d
E = np.load('PageRankEG1.npy')
E
M = make_M_from_E(E)
M
x = np.ones(4)/4
x
# Power Method
for i in range(100):
x = M @ x
x = x / npla.norm(x)
x
d, V = linalg.eig(M)
V[:,0].real
| 02.13/Class_transcript_02_13_pagerank.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
#
# 1. Orthogonal Projections
# Recall that for projection of a vector $\boldsymbol x$ onto a 1-dimensional subspace $U$ with basis vector $\boldsymbol b$ we have
#
# $${\pi_U}(\boldsymbol x) = \frac{\boldsymbol b\boldsymbol b^T}{{\lVert\boldsymbol b \rVert}^2}\boldsymbol x $$
#
# And for the general projection onto an M-dimensional subspace $U$ with basis vectors $\boldsymbol b_1,\dotsc, \boldsymbol b_M$ we have
#
# $${\pi_U}(\boldsymbol x) = \boldsymbol B(\boldsymbol B^T\boldsymbol B)^{-1}\boldsymbol B^T\boldsymbol x $$
#
# where
#
# $$\boldsymbol B = [\boldsymbol b_1,...,\boldsymbol b_M]$$
#
#
# Your task is to implement orthogonal projections. We can split this into two steps
# 1. Find the projection matrix $\boldsymbol P$ that projects any $\boldsymbol x$ onto $U$.
# 2. The projected vector $\pi_U(\boldsymbol x)$ of $\boldsymbol x$ can then be written as $\pi_U(\boldsymbol x) = \boldsymbol P\boldsymbol x$.
#
# To perform step 1, you need to complete the function `projection_matrix_1d` and `projection_matrix_general`. To perform step 2, complete `project_1d` and `project_general`.
# ### Projection (1d)
import numpy as np
# +
def projection_matrix_1d(b):
"""Compute the projection matrix onto the space spanned by `b`
Args:
b: ndarray of dimension (D,), the basis for the subspace
Returns:
P: the projection matrix
"""
P = np.outer(b, b) / (np.linalg.norm(b)**2)
return P
def project_1d(x, b):
"""Compute the projection matrix onto the space spanned by `b`
Args:
x: the vector to be projected
b: ndarray of dimension (D,), the basis for the subspace
Returns:
y: ndarray of shape (D,) projection of x in space spanned by b
"""
p = projection_matrix_1d(b=b) @ x
return p
# +
# testing
from numpy.testing import assert_allclose
assert_allclose(
projection_matrix_1d(np.array([1, 2, 2])),
np.array([[1, 2, 2],
[2, 4, 4],
[2, 4, 4]]) / 9
)
assert_allclose(
project_1d(np.ones(3), np.array([1, 2, 2])),
np.array([5, 10, 10]) / 9
)
# -
# ### Projection (ND)
# +
def projection_matrix_general(B):
"""Compute the projection matrix onto the space spanned by the columns of `B`
Args:
B: ndarray of dimension (D, M), the basis for the subspace
Returns:
P: the projection matrix
"""
P = (B@(np.linalg.inv(B.T@B))@B.T)
return P
def project_general(x, B):
"""Compute the projection matrix onto the space spanned by the columns of `B`
Args:
x: ndarray of dimension (D, 1), the vector to be projected
B: ndarray of dimension (D, M), the basis for the subspace
Returns:
p: projection of x onto the subspac spanned by the columns of B; size (D, 1)
"""
p = projection_matrix_general(B) @ x
return p
# +
# testing
from numpy.testing import assert_allclose
B = np.array([[1, 0],
[1, 1],
[1, 2]])
assert_allclose(
projection_matrix_general(B),
np.array([[5, 2, -1],
[2, 2, 2],
[-1, 2, 5]]) / 6
)
assert_allclose(
project_general(np.array([6, 0, 0]).reshape(-1,1), B),
np.array([5, 2, -1]).reshape(-1,1)
)
| math4ml/33_projections.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from IPython.display import Image
from IPython.core.display import HTML
from sympy import *; x,h = symbols("x h")
Image(url= "https://i.imgur.com/a9ZXtwh.png")
expr = sqrt(3*x**2+4*x+8)
dexpr = (diff(expr)) #getting derivative of our expression
print(dexpr)
print(dexpr.subs(x,5))
Image(url= "https://i.imgur.com/JpePLE7.png")
| Calculus_Homework/WWB09.5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Problem set 6: Solving the Solow model
# [<img src="https://mybinder.org/badge_logo.svg">](https://mybinder.org/v2/gh/NumEconCopenhagen/exercises-2020/master?urlpath=lab/tree/PS6/problem_set_6.ipynb)
import numpy as np
from scipy import linalg
from scipy import optimize
import sympy as sm
# %load_ext autoreload
# %autoreload 2
# # Tasks
# ## Solving matrix equations I
np.random.seed(1900)
n = 5
A = np.random.uniform(size=(n,n))
b = np.random.uniform(size=n)
c = np.random.uniform(size=n)
d = np.random.uniform(size=n)
# **Question A:** Find the determinant of $[A \cdot A]^{-1}$
linalg.det(linalg.inv(np.dot(A,A)))
# **Answer:**
# + tags=[]
X = linalg.det(linalg.inv(A@A))
print(X)
# -
A
b
linalg.solve(A,b)
# **Question B:** Solve the following equation systems directly using **scipy**.
#
# $$
# \begin{aligned}
# Ax &= b \\
# Ax &= c \\
# Ax &= d
# \end{aligned}
# $$
for vec in [b,c,d]:
print(linalg.solve(A,vec))
# **Answer:**
# + tags=[]
xb = linalg.solve(A,b)
xc = linalg.solve(A,c)
xd = linalg.solve(A,d)
print('b:',xb)
print('c:',xc)
print('d:',xd)
# -
# **Question C:** Solve the same equation systems as above using `linalg.lu_factor()` and `linalg.lu_solve()`. What is the benefit of this approach?
LU,piv = linalg.lu_factor(A)
xb = linalg.lu_solve((LU,piv),b)
xc = linalg.lu_solve((LU,piv),c)
xd = linalg.lu_solve((LU,piv),d)
for vec in [xb,xc,xd]:
print(vec)
# **Answer:**
# + tags=[]
LU,piv = linalg.lu_factor(A) # only done once
xb = linalg.lu_solve((LU,piv),b) # much faster than regular solve
xc = linalg.lu_solve((LU,piv),c)
xd = linalg.lu_solve((LU,piv),d)
print('b:',xb)
print('c:',xc)
print('d:',xd)
# -
# ## Solving matrix equations II
F = np.array([[2.0, 1.0, -1.0], [-3.0, -1.0, 2], [-2.0, 1.0, 2.0]])
e = np.array([[8.0, -11.0, -3.0]])
print(e.shape)
F
e
# **Question:** Use the function `gauss_jordan()` in the `numecon_linalg` module located in this folder to solve
#
# $$
# Fx = e
# $$
import numecon_linalg
np.column_stack((F,e))
## Alternative stacking methods:
np.concatenate((F,e[:, np.newaxis]),axis=1)
np.append(F,e[:, np.newaxis],axis=1)
# It's actually easier if e is intitiated as a 2-D matrix of 3x1
# Using [[ ]]:
e2 = np.array([[8.0, -11.0, -3.0]])
print(e2.shape)
test = np.concatenate((F,e2.T),axis=1)
test
# +
Y = np.concatenate((F,e[:, np.newaxis]),axis=1)
numecon_linalg.gauss_jordan(Y)
Y
# -
# **Answer:**
# + tags=[]
import numecon_linalg
Y = np.column_stack((F,e))
numecon_linalg.gauss_jordan(Y)
print('solution',Y[:,-1])
assert np.allclose(F@Y[:,-1],e)
# -
# ## Symbolic
# **Question A:** Find
#
# $$
# \lim_{x \rightarrow 0} \frac{\sin(x)}{x}
# $$
#
# and
#
# $$
# \frac{\partial\sin(2x)}{\partial x}
# $$
x = sm.symbols('x')
sm.limit(sm.sin(x)/x,x,0)
sm.diff(sm.sin(2*x),x)
# **Answer:**
# + jupyter={"source_hidden": true} tags=[]
print('the limit is:')
x = sm.symbols('x')
sm.limit(sm.sin(x)/x,x,0)
# + jupyter={"source_hidden": true}
print('the derivative is')
x = sm.symbols('x')
sm.diff(sm.sin(2*x),x)
# -
# **Question B:** Solve the equation
#
# $$
# \frac{\sin(x)}{x} = 0
# $$
sm.solve(sm.sin(x)/x,x)
# **Answer:**
sm.solve(sm.sin(x)/x)
# # Problem: Solve the Solow model
# ## Introduction
# Consider the **standard Solow-model** where:
#
# 1. $K_t$ is capital2
# 2. $L_t$ is labor (growing with a constant rate of $n$)
# 3. $A_t$ is technology (growing with a constant rate of $g$)
# 4. $Y_t = F(K_t,A_tL_t)$ is GDP
#
# **Saving** is a constant fraction of GDP
#
# $$
# S_t = sY_t,\,s\in(0,1)
# $$
#
# such that **capital accumulates** according to
#
# $$
# K_{t+1}=S_{t}+(1-\delta)K_{t}=sF(K_{t},A_{t}L_{t})+(1-\delta)K_{t}, \delta \in (0,1)
# $$
#
# The **production function** has **constant-return to scale** such that
#
# $$
# \frac{Y_{t}}{A_{t}L_{t}}=\frac{F(K_{t},A_{t}L_{t})}{A_{t}L_{t}}=F(\tilde{k}_{t},1)\equiv f(\tilde{k}_{t})
# $$
#
# where $\tilde{k}_t = \frac{K_t}{A_{t}L_{t}}$ is the technology adjusted capital-labor ratio.
#
# The **transition equation** then becomes
#
# $$
# \tilde{k}_{t+1}= \frac{1}{(1+n)(1+g)}[sf(\tilde{k}_{t})+(1-\delta)\tilde{k}_{t}]
# $$
#
# If the **production function** is **Cobb-Douglas** then
#
# $$
# F(K_{t},A_{t}L_{t})=K_{t}^{\alpha}(A_{t}L_{t})^{1-\alpha}\Rightarrow f(\tilde{k}_{t})=\tilde{k}_{t}^{\alpha}
# $$
#
# If it is **CES** (with $\beta < 1, \beta \neq 0$) then
#
# $$
# F(K_{t},A_{t}L_{t})=(\alpha K_{t}^{\beta}+(1-\alpha)(A_{t}L_{t})^{\beta})^{\frac{1}{\beta}}\Rightarrow f(\tilde{k}_{t})=(\alpha\tilde{k}_{t}^{\beta}+(1-\alpha))^{\frac{1}{\beta}}
# $$
# ## Steady state
# Assume the production function is **Cobb-Douglas**.
# **Question A:** Use **sympy** to find an analytical expression for the steady state, i.e. solve
#
# $$
# \tilde{k}^{\ast}= \frac{1}{(1+n)(1+g)}[sf(\tilde{k}^{\ast})+(1-\delta)\tilde{k}^{\ast}]
# $$
k = sm.symbols('k')
alpha = sm.symbols('alpha')
delta = sm.symbols('delta')
s = sm.symbols('s')
g = sm.symbols('g')
n = sm.symbols('n')
k, alpha, delta, s , g, n = sm.symbols('k alpha delta s g n')
#f = lambda k : k**alpha
f = k**alpha
1/((1+n)*(1+g))*(s*f+(1-delta)*k)-k
k_star = sm.solve(1/((1+n)*(1+g))*(s*f+(1-delta)*k)-k,k)[0]
k_star
# **Answer:**
# + tags=[]
f = k**alpha
ss = sm.Eq(k,(s*f+(1-delta)*k)/((1+n)*(1+g)))
kss = sm.solve(ss,k)[0]
kss
# -
from IPython.display import display
# +
### CES Preferences, Sympy can get overwhelmed by difficult problems
beta = sm.symbols('beta',negative=True)
f_CES = (alpha *k**(beta)+(1-alpha))**(1/beta)
ss_CES = sm.Eq(k,(s*f_CES+(1-delta)*k)/((1+n)*(1+g)))
kss_CES = sm.solve(ss_CES,k)
for eq in kss_CES:
display(eq)
# -
# We store the solution and test them later when we have a numerical solution. We will find that only one is right (the second). <br>
# A further drawback of this solution is that I could not make it work when assuming $k\in\left(0,1\right)$, even though the solution is still correct in the interval.
ss_CES_func = [sm.lambdify((s,g,n,delta,alpha,beta),sol) for sol in kss_CES]
# **Question B:** Turn you solution into a Python function called as `ss_func(s,g,n,delta,alpha)`.
k_star
ss_func = sm.lambdify((s,g,n,delta,alpha),k_star)
# **Answer:**
# + jupyter={"source_hidden": true} tags=[]
ss_func = sm.lambdify((s,g,n,delta,alpha),kss)
# -
ss_func(0.2,0.02,0.01,0.1,1/3)
# **Question C**: Find the steady state numerically using root-finding with `optimize.root_scalar`.
# +
s = 0.2
g = 0.02
n = 0.01
alpha = 1/3
delta = 0.1
f = lambda k : k**alpha
solveit = lambda k: 1/((1+n)*(1+g))*(s*f(k)+(1-delta)*k)-k
# -
# Brentq::
optimize.root_scalar(solveit,bracket=[0.1,100],method='brentq')
# Bisect:
optimize.root_scalar(solveit,bracket=[0.1,100],method='bisect')
# secant, no brackets (bounds) needed, however two guesses are:
optimize.root_scalar(solveit,x0=50,x1=100,method='secant')
# +
## Finding the first order condition,
k_, alpha_, delta_, s_ , g_, n_ = sm.symbols('k alpha delta s g n')
solveit_ = 1/((1+n_)*(1+g_))*(s_*k_**alpha_+(1-delta_)*k_)-k_
solveit_diff_ = sm.diff(solveit_,k)
solveit_diff_l = sm.lambdify((k_, alpha_, delta_, s_ , g_, n_),solveit_diff_)
solveit_diff_
# +
# Newton, no brackets (bounds) needed, however the FOC is.
#solveit_diff = lambda k: 1/((1+n)*(1+g))*(s*alpha*k**(alpha-1)+(1-delta))-1
solveit_diff = lambda k: solveit_diff_l(k, alpha, delta, s , g, n)
optimize.root_scalar(solveit,x0=50,method='newton',fprime=solveit_diff)
# +
# Using root
obj_kss = lambda k : solveit(k[0])
optimize.root(obj_kss,x0=[2])
# +
# Using root
jac = lambda k : [solveit_diff(k[0])] ## Output needs to be array-like because root expects jacobian
optimize.root(obj_kss,jac=jac,x0=[2], method='hybr')
# -
import matplotlib.pyplot as plt
# +
s = 0.2
g = 0.02
n = 0.01
alpha = 1/3
delta = 0.1
params = {'s':0.2,'g':0.02,'n':0.01,'alpha':1/3,'delta':0.1}
transitioneq = lambda k,parms: (params['s']*k**params['alpha']+(1-params['delta'])*k)/((1+params['n'])*(1+params['g']))
def ss_num(transitioneq,params,guess=2,rtol=1.0000000000000001e-010,
atol=1e-12,max_iter=2000, printit=True):
'''
Finds the steady state(ss) for the transitionseq by simulating the model until equlibrium is reached
args:
transitioneq (function) : Transition equation to find ss for
params (dict) : A dictionary with parameters values for the transition equation
guees (float) : Inital guess
rtol,atol (float) : Relative and absolut tolerance for np.isclose
jump (float) : relative size of each jump when making a new guess
max_iter (int) : Maximum number of iterations
prinit (Bool) : Whether or not to print the results
'''
k1= transitioneq(guess,params)
cond = np.isclose(k1,guess,rtol=rtol,atol=atol)
i = 0
while not cond:
i+=1
if i>max_iter:
print('max iterations reached')
break
return None
# Make new guess by guessing the way of the evolution of k, so if kt+1>kt the new guess of k becomes larger
guess = k1
k1= transitioneq(guess,params)
cond = np.isclose(k1,guess,rtol=rtol,atol=atol) # Check if kt+1=kt
if printit:
print(f'K_star is {guess:.20f}')
print(f'Check: {k1:.20f}')
return guess
kstar = ss_num(transitioneq,params)
# +
#plot solution
ks = np.linspace(0,4,10000)
ks1 = [transitioneq(k,params) for k in ks]
fig = plt.figure()
ax = fig.add_subplot(1,1,1)
ax.plot(ks,ks1,label='$k_{t+1}(k_{t})$')
ax.plot(ks,ks,ls='--',color='black',label='$k_{t}=k_{t+1}$')
ax.plot(kstar,transitioneq(kstar,params),ls='',marker='o',color='blue',label='$k^{*}$')
ax.grid(True)
plt.legend(loc="upper left")
fig.tight_layout()
# -
# **Answer:**
# + tags=[]
f = lambda k: k**alpha
obj_kss = lambda kss: kss - (s*f(kss) + (1-delta)*kss)/((1+g)*(1+n))
result = optimize.root_scalar(obj_kss,bracket=[0.1,100],method='brentq')
print('the steady state for k is',result.root)
# -
# **Question D:** Now assume the production function is CES. Find the steady state for $k$ for the various values of $\beta$ shown below.
# +
betas = [-0.5,-0.25,-0.1,-0.05,0.05,0.1,0.25,0.5]
f = lambda k,params: (params['alpha']*k**params['beta']+(1-params['alpha']))**(1/params['beta'])
trans_eq = lambda kss,params:(params['s']*f(kss,params) + (1-params['delta'])*kss)/((1+params['g'])*(1+params['n']))
k_stars = np.empty(len(betas))
for i,beta in enumerate(betas):
params['beta']=beta
k_stars[i] = ss_num(trans_eq,params, printit=False)
print(f'When beta= {beta:5.2f} the steady state for k is {k_stars[i]:.15f}')
# +
for beta,k_star in zip(betas,k_stars):
params['beta']=beta
ks = np.linspace(1,3,10000)
ks1 = [trans_eq(k,params) for k in ks]
fig = plt.figure()
ax = fig.add_subplot(1,1,1)
ax.plot(ks,ks1,label='$k_{t+1}(k_{t})$')
ax.plot(ks,ks,ls='--',color='black',label='$k_{t}=k_{t+1}$')
ax.plot(k_star,transitioneq(k_star,params),ls='',marker='o',color='blue',label='$k^{*}$')
ax.grid(True)
plt.title(r'$\beta=$'+f'{beta}')
plt.legend(loc="upper left")
fig.tight_layout()
# + tags=[]
for beta in betas:
f = lambda k: (alpha*k**beta + (1-alpha))**(1/beta)
obj_kss = lambda kss: kss - (s*f(kss) + (1-delta)*kss)/((1+g)*(1+n))
result = optimize.root_scalar(obj_kss,bracket=[0.1,100],method='brentq')
print(f'for beta = {beta:.3f} the steady state for k is',result.root)
# -
# Test the symbolic solutions found earlier:
for beta in betas:
sol = [fun(s,g,n,delta,alpha,beta) for fun in ss_CES_func]
print(f'For beta = {beta:.3f} first solution is {sol[0]:.4f} and second is {sol[1]:.4f}')
| PS6/problem_set_6-sol.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/elysabethe/JornadaPython/blob/main/Faixapreta.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="vMeIWT4Twwwz"
# # Introdução a Python
#
# + id="xaHDQTWPwoVf"
# Escopo pai
#Escopo filho
# Escopo Neto
# + [markdown] id="nWrr52WKw5j_"
#
# #Variáveis
# + id="iAyUxU-NxDB1"
nome = 'elysa'
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="nX0bIshkxi6u" outputId="ebf24eb0-5dc2-4fa1-f5d3-31321116e261"
nome
# + id="vuFFHHvOyEHA"
n1 = 8
# + id="G3QGw0HUyIaQ"
n2 = 7
# + id="g9ylE8viyK4D"
soma=0
# + id="NNB5IWbU0IB2"
soma=n1+n2
# + colab={"base_uri": "https://localhost:8080/"} id="GJbMyHQq0Ki0" outputId="d8004f97-9005-4087-db96-a9684e5e6976"
soma
# + [markdown] id="043jqbWfw3sf"
# ### Operações Matemáticas
# + id="lEHP0JId02Ze"
*soma usamos +
*subtracao usamos -
*multiplicacao usamos *
*divisao usamos + /
# + id="tTCSkI9O1Oag"
soma = 10+15
# + colab={"base_uri": "https://localhost:8080/"} id="9LK1N-cX1T6Y" outputId="81a389ff-7097-471b-f1fe-47f98db559f4"
soma
# + id="pgH32eLu1Vym"
subtracao = 50-12
# + id="amT_ECmE1ZKt"
subtracao
# + id="PiBc0cxd1hkq"
multiplicacao = 4*9
# + id="dBkEv57K1kX8"
multiplicacao
# + id="ixR4s0yJ1lwn"
divisao = 5/5
# + id="qGHf07st1pc7"
divisao
# + [markdown] id="X6CUFXMe2fOW"
# # Métodos de Entrada
# formas de receber dados do usuário ou seja enviar valores para o programa
#
# No python usamos _input()_
# passamos o texto por parametro
# + colab={"base_uri": "https://localhost:8080/", "height": 129} id="qUO8BZ7K4DQd" outputId="5c7c118c-1fb4-401b-bed1-bf00b9b0f422"
nome input('Qual é o seu nome?')
# + colab={"base_uri": "https://localhost:8080/"} id="qNUzygFC5VZI" outputId="6948a682-4a7e-4d1b-98d4-1dcfdd7b3055"
nome = input('oiii ')
# + id="DX6pKB1c451l"
#conversaodedados de string para int
# + colab={"base_uri": "https://localhost:8080/"} id="TTF6KSN84-nC" outputId="9a429bae-c60d-48c1-d16a-ccbead914aa2"
bool(0)
# + id="QlzUqC0B4dUJ"
# + [markdown] id="546EHv0_90-L"
# ## Concatenação
# + id="GV01HhVY9_Bt"
nome = '<NAME>'
# + colab={"base_uri": "https://localhost:8080/"} id="lL2kKjqt-FgH" outputId="7d759e75-2813-4189-ebfa-05785241dad5"
print('Olá, eu me chamo', nome)
# + [markdown] id="8itIIo3w_VdU"
# # Média
#
# #vamos calcular a média bb
#
# + id="gWSP23-y_bp7"
# + id="DGfavted_h-u"
# + [markdown] id="il3sEji0_iVj"
# # Listas
#
# na pegada dos arrays meu amorzinho s2
# lembre-se que é o índice que determina o rolê
#
# + id="o9SRnWiAA3Jb"
lista=['cachorro', 'gato']
# + id="H0Wr8qBXBLlD"
mel = lista[0]
# + id="_K1_vIIVBRvf"
pepa = lista[1]
# + colab={"base_uri": "https://localhost:8080/"} id="rmEU-RYmBVXo" outputId="94d99938-789e-42e4-c2ac-f2d8fc2169ca"
print(lista)
# + colab={"base_uri": "https://localhost:8080/"} id="loVSSI2CBaBZ" outputId="2700e74f-83ad-4cf4-cfe7-1a32629c89b4"
print(lista[0])
# + colab={"base_uri": "https://localhost:8080/"} id="s1Se97BdBe5g" outputId="b6f55f17-b186-45c9-8b8a-93e33c89a690"
print('Lista dos animais', lista)
# + colab={"base_uri": "https://localhost:8080/"} id="MVL75MqABvcT" outputId="fa01f2b8-803b-45a0-e821-17126a128135"
#todos os valores em um array
type(lista)
# + id="3ee0uc96B4tK"
lista_dados = ['1', '2'], ['3','4']
# + colab={"base_uri": "https://localhost:8080/"} id="vea9XfZjDN0B" outputId="9f1070a5-dd87-4713-d7e5-0f15779b2930"
print(f'A matriz de leves {lista_dados[0][1]}')
# + colab={"base_uri": "https://localhost:8080/"} id="_KZEQSMAE2P1" outputId="0e467aee-10d7-4a91-9033-0a9f5d613b93"
len(lista_dados)
# + [markdown] id="qq5XZ8RiFLbd"
# # Condicionais
# if =>se
# elif => senao, se
# else => senao
#
# _lembre de colocar os:_
#
# ex: if nome =='Felipe':
# else:
# print('a')
#
#
#
# + [markdown] id="R8bkiuD850ch"
# # Calculadora
# + colab={"base_uri": "https://localhost:8080/", "height": 129} id="Th8mlESS56Y6" outputId="11865250-4b37-45ed-a80b-63c1737cfedf"
print('Informe a operação desejada:')
print(' 1. + ')
print(' 2. - ')
print(' 3. * ')
print(' 4. / ')
op = input()
if op == 1 :
print(n1+n2)
elif op=='2':
resultado = n1-n2
n1 = float(input('Informe o primeiro valor: '))
n2 = float(input('Informe o segundo valor: '))
print('A operação ',op,'gerou o resultado: ', resultado)
| Faixapreta.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: aquarium_kernel
# language: python
# name: aquarium_kernel
# ---
# +
import platform
import psutil
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
plt.rcParams['figure.facecolor']='white'
plt.rcParams['font.size']=16
import bioframe
import pyranges
# -
print(f"Bioframe v.{bioframe.__version__}")
print(f"PyRanges v.{pyranges.__version__}")
print(f"System Platform: {platform.platform()}")
print(f"{psutil.cpu_count()} CPUs at {psutil.cpu_freq().current:.0f} GHz")
def make_random_intervals(
n=1e5, n_chroms=1, max_coord=None, max_length=10, categorical_chroms=False,
sort=False
):
n = int(n)
n_chroms = int(n_chroms)
max_coord = (n // n_chroms) if max_coord is None else int(max_coord)
max_length = int(max_length)
chroms = np.array(['chr'+str(i+1) for i in range(n_chroms)])[
np.random.randint(0, n_chroms, n)]
starts = np.random.randint(0, max_coord, n)
ends = starts + np.random.randint(0, max_length, n)
df = pd.DataFrame({
'chrom':chroms,
'start':starts,
'end':ends
})
if categorical_chroms:
df['chrom'] = df['chrom'].astype('category')
if sort:
df = df.sort_values(['chrom','start','end']).reset_index()
return df
# # Overlap
# ### vs number of intervals
timings = {}
for n in [1e2, 1e3, 1e4, 1e5, 1e6]:
df = make_random_intervals(n=n, n_chroms=1)
df2 = make_random_intervals(n=n, n_chroms=1)
timings[n] = %timeit -o -r 1 bioframe.overlap(df, df2)
plt.loglog(
list(timings.keys()),
list([r.average for r in timings.values()]),
'o-',
)
plt.xlabel('N intervals')
plt.ylabel('time, ms')
plt.gca().set_aspect(1.0)
plt.grid()
# ### vs total number of intersections
timings = {}
n_intersections = {}
n = 1e4
for avg_interval_len in [3, 1e1, 3e1, 1e2, 3e2]:
df = make_random_intervals(n=n, n_chroms=1, max_length=avg_interval_len*2)
df2 = make_random_intervals(n=n, n_chroms=1, max_length=avg_interval_len*2)
timings[avg_interval_len] = %timeit -o -r 1 bioframe.overlap(df, df2)
n_intersections[avg_interval_len] = bioframe.overlap(df, df2).shape[0]
plt.loglog(
list(n_intersections.values()),
list([r.average for r in timings.values()]),
'o-',
)
plt.xlabel('N intersections')
plt.ylabel('time, ms')
plt.gca().set_aspect(1.0)
plt.grid()
# ### vs number of chromosomes
timings = {}
n_intersections = {}
n = 1e5
for n_chroms in [1, 3, 10, 30, 100, 300, 1000]:
df = make_random_intervals(n, n_chroms)
df2 = make_random_intervals(n, n_chroms)
timings[n_chroms] = %timeit -o -r 1 bioframe.overlap(df, df2)
n_intersections[n_chroms] = bioframe.overlap(df, df2).shape[0]
n_intersections
plt.loglog(
list(timings.keys()),
list([r.average for r in timings.values()]),
'o-',
)
plt.xlabel('# chromosomes')
plt.ylabel('time, ms')
# plt.gca().set_aspect(1.0)
plt.grid()
# ### vs other parameters: join type, sorted or categorical inputs
# +
df = make_random_intervals()
df2 = make_random_intervals()
# %timeit -r 1 bioframe.overlap(df, df2)
# %timeit -r 1 bioframe.overlap(df, df2, how='inner')
# %timeit -r 1 bioframe.overlap(df, df2, how='left')
# +
df = make_random_intervals()
df2 = make_random_intervals()
# %timeit -r 1 bioframe.overlap(df, df2)
# %timeit -r 1 bioframe.overlap(df, df2, return_overlap=True)
# %timeit -r 1 bioframe.overlap(df, df2, keep_order=True)
# +
df = make_random_intervals()
df2 = make_random_intervals()
# %timeit -r 1 bioframe.overlap(df, df2)
# %timeit -r 1 bioframe.overlap(df, df2, how='inner')
# %timeit -r 1 bioframe.ops._overlap_intidxs(df, df2)
# %timeit -r 1 bioframe.ops._overlap_intidxs(df, df2, how='inner')
# +
print('Default inputs (outer/inner joins):')
df = make_random_intervals()
df2 = make_random_intervals()
# %timeit -r 1 bioframe.overlap(df, df2)
# %timeit -r 1 bioframe.overlap(df, df2, how='inner')
print('Sorted inputs (outer/inner joins):')
df_sorted = make_random_intervals(sort=True)
df2_sorted = make_random_intervals(sort=True)
# %timeit -r 1 bioframe.overlap(df_sorted, df2_sorted)
# %timeit -r 1 bioframe.overlap(df_sorted, df2_sorted, how='inner')
print('Categorical chromosomes (outer/inner joins):')
df_cat = make_random_intervals(categorical_chroms=True)
df2_cat = make_random_intervals(categorical_chroms=True)
# %timeit -r 1 bioframe.overlap(df_cat, df2_cat)
# %timeit -r 1 bioframe.overlap(df_cat, df2_cat, how='inner')
# -
# # Vs Pyranges
# ### Default arguments
def df2pr(df):
return pyranges.PyRanges(
chromosomes=df.chrom,
starts=df.start,
ends=df.end,
)
timings_bf = {}
timings_pr = {}
for n in [1e2, 1e3, 1e4, 1e5, 1e6]:
df = make_random_intervals(n=n, n_chroms=1)
df2 = make_random_intervals(n=n, n_chroms=1)
pr = df2pr(df)
pr2 = df2pr(df2)
timings_bf[n] = %timeit -o -r 1 bioframe.overlap(df, df2)
timings_pr[n] = %timeit -o -r 1 pr.intersect(pr2)
plt.loglog(
list(timings_bf.keys()),
list([r.average for r in timings_bf.values()]),
'o-',
label='bioframe'
)
plt.loglog(
list(timings_pr.keys()),
list([r.average for r in timings_pr.values()]),
'o-',
label='pyranges'
)
plt.xlabel('N intervals')
plt.ylabel('time, ms')
plt.gca().set_aspect(1.0)
plt.grid()
plt.legend()
# ### +conversion from/into dataframes +"inner" join type
timings_bf = {}
timings_pr = {}
for n in [1e2, 1e3, 1e4, 1e5, 1e6, 3e6]:
df = make_random_intervals(n=n, n_chroms=1)
df2 = make_random_intervals(n=n, n_chroms=1)
timings_bf[n] = %timeit -o -r 1 bioframe.overlap(df, df2, how='inner')
timings_pr[n] = %timeit -o -r 1 df2pr(df).intersect(df2pr(df2)).as_df()
plt.loglog(
list(timings_bf.keys()),
list([r.average for r in timings_bf.values()]),
'o-',
label='bioframe'
)
plt.loglog(
list(timings_pr.keys()),
list([r.average for r in timings_pr.values()]),
'o-',
label='pyranges'
)
plt.xlabel('N intervals')
plt.ylabel('time, ms')
plt.gca().set_aspect(1.0)
plt.grid()
plt.legend()
# # Slicing
timings_bf = {}
timings_pr = {}
for n in [1e2, 1e3, 1e4, 1e5, 1e6, 3e6]:
df = make_random_intervals(n=n, n_chroms=1)
timings_bf[n] = %timeit -o -r 1 bioframe.select(df, ('chr1', n//2, n//4*3))
pr = df2pr(df)
timings_pr[n] = %timeit -o -r 1 pr['chr1', n//2:n//4*3]
# +
plt.loglog(
list(timings_bf.keys()),
list([r.average for r in timings_bf.values()]),
'o-',
label='bioframe'
)
plt.loglog(
list(timings_pr.keys()),
list([r.average for r in timings_pr.values()]),
'o-',
label='pyranges'
)
plt.xlabel('N intervals')
plt.ylabel('time, ms')
plt.gca().set_aspect(1.0)
plt.grid()
plt.legend()
# +
# # %timeit intarr=pd.IntervalIndex.from_arrays(starts1, ends1)
# # %timeit intarr=pd.IntervalIndex.from_arrays(starts1, ends1)
# # %timeit intarr.overlaps(pd.Interval(500000,600000))
# # %timeit (starts1<=600000)&(ends1>=500000)
| docs/notebooks/performance.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.4.1
# language: julia
# name: julia-1.4
# ---
# # Electronic Structure and Sommerfeld Factor
#
# This notebook shows how to calculate the sommerfeld factor for a charged defect
#
# See Landsberg, Recombination in Semiconductors (Cambridge University Press, 2009) for the detail.
# ## Bohr radius
# $a_{B}=\frac{4\pi\epsilon_{0}\hbar^{2}}{m_{e}e^{2}} = 5.291772083\times10^{-11} \mathrm{m} =
# 0.53 Å$
# $a_{B}^{*}=\frac{\epsilon}{m_{e}^{*}/m_{e}}a_{B}$
BohrRadius = 5.291772083e-11; # m
Boltzmann = 1.3806504e-23; # kg m^2 / K s^2
ElectronVolt = 1.602176487e-19; # kg m^2 / s^2
PlancksConstantHbar = 1.05457162825e-34; # kg m^2 / s
PlancksConstantH = 6.626070040e-34;
ElectronCharge = 1.602176487e-19; # A s
VacuumPermittivity = 8.854187817e-12; # A^2 s^4 / kg m^3
MassElectron = 9.10938188e-31; # kg
pi = 3.141592
# ## Effective masses of CZTS and CZTSe
# +
m_e_CZTS = 0.18
m_h_CZTS = 0.40
m_e_CZTSe = 0.10
m_h_CZTSe = 0.23
ϵ_0_CZTS = 9.9
ϵ_0_CZTSe = 11.4
a_B_e_CZTS = ϵ_0_CZTS/m_e_CZTS*BohrRadius
a_B_h_CZTS = ϵ_0_CZTS/m_h_CZTS*BohrRadius
a_B_e_CZTSe = ϵ_0_CZTSe/m_e_CZTSe*BohrRadius
a_B_h_CZTSe = ϵ_0_CZTSe/m_h_CZTSe*BohrRadius
println(a_B_e_CZTS)
println(a_B_h_CZTS)
println(a_B_e_CZTSe)
println(a_B_h_CZTSe)
# -
T = 330
NC=2*(2*pi*m_e_CZTS*MassElectron*Boltzmann*T/PlancksConstantH^2)^(3/2.)
NV=2*(2*pi*m_h_CZTS*MassElectron*Boltzmann*T/PlancksConstantH^2)^(3/2.)
println("effective electron dos: $(NC)")
println("effective hole dos: $(NV)")
# ## Thermal velocity
# $v_{t}=\sqrt(3k_{B}T/m^{*})$
T = 300
println("v_n-CZTS: $(sqrt(3*Boltzmann*T/(m_e_CZTS*MassElectron))*100) cm/s")
println("v_p-CZTS: $(sqrt(3*Boltzmann*T/(m_h_CZTS*MassElectron))*100) cm/s")
println("v_n-GaAs(0.063): $(sqrt(3*Boltzmann*T/(0.063*MassElectron))*100) cm/s")
println("v_p-GaAs(0.45): $(sqrt(3*Boltzmann*T/(0.45*MassElectron))*100) cm/s")
# ## Sommerfeld factor
#
# see Landsberg, Recombination in Semiconductors (Cambridge University Press, 2009)
# +
# Electron capture in CZTS
Z = 1
ER=13.6*ElectronCharge * (m_e_CZTS/ϵ_0_CZTS^2) # Effective Rydberg
s=4*Z*(pi*ER/Boltzmann/T)^0.5
println("Attractive Center: $(s)")
K=(pi^2*Z^2*ER/Boltzmann/T)
s=8/sqrt(3)*K^(2/3.)*exp(-3*K^(1/3.))
println("Repulsive Center: $(s)")
# +
# Hole capture in CZTS
Z = 1
ER=13.6*ElectronCharge * (m_h_CZTS/ϵ_0_CZTS^2) # Effective Rydberg
s=4*Z*(pi*ER/Boltzmann/T)^0.5
println("Attractive Center: $(s)")
K=(pi^2*Z^2*ER/Boltzmann/T)
s=8/sqrt(3)*K^(2/3.)*exp(-3*K^(1/3.))
println("Repulsive Center: $(s)")
# -
# ## Reading list
# 1. <NAME>, Phys. Status Solidi B 76, 647 (1976)
#
# 2. <NAME>, Phys. Status Solidi B 78, 625 (1976)
| example/notebook/Sommerfeld-factor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
from data_tools.generate_testing_dataset import call_df
df = call_df()
len(df.loc[df['condition'] == 'fair'])
df = df.loc[df['condition'] == 'fair']
df = df.loc[df['manufacturer']=='toyota']
df.isnull()
| webapp/datah/data_creation_experiments.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: p6
# language: python
# name: other-env
# ---
"""setup"""
try:
from ..__init__ import Import, Notebook, from_file, reload, update_hooks
except:
from aye import Import, Notebook, from_file, reload, update_hooks
import sys
"""test_resting_state"""
try:
try:
from . import test_basics
except:
import test_basics
# Throw an exception til here
assert False, "test_basics is in the sys.path as <{}>".format(test_basics.__file__)
except ModuleNotFoundError:
assert 'test_basics' not in globals()
# +
"""test_import_context_manager"""
with Import(Notebook):
try:
from . import test_basics
except:
import test_basics
finally:
assert test_basics
try:
try:
from . import test_from_file
except:
import test_from_file
# assert False, "test_from_file is in the sys.path as <{}>".format(test_from_file.__file__)
except ModuleNotFoundError:
assert 'test_from_file' not in globals()
update_hooks(Notebook)
try:
from . import test_from_file
except:
import test_from_file
assert test_from_file
update_hooks()
# -
| aye/tests/test_ordering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import logging
logging.basicConfig(filename = 'opSavingJohn.txt', filemode='w', level=logging.INFO)
import numpy as np
import os
from collections import defaultdict
from keras.optimizers import RMSprop
from keras.utils import plot_model
from pyfiction.agents.ssaqn_agent import SSAQNAgent
from pyfiction.agents.ssaqn_agent import load_embeddings
from pyfiction.simulators.games.savingjohn_simulator import SavingJohnSimulator
from pyfiction.simulators.text_games.simulators.MySimulator import StoryNode
logger = logging.getLogger(__name__)
"""
An example SSAQN agent for Saving John that uses online learning and prioritized sampling
"""
# Create the agent and specify maximum lengths of descriptions (in words)
agent = SSAQNAgent(train_simulators=SavingJohnSimulator(), mode = 'LSTM')
# Learn the vocabulary (the function samples the game using a random policy)
agent.initialize_tokens('vocabulary.txt')
optimizer = RMSprop(lr=0.0001)
embedding_dimensions = 300
lstm_dimensions = 32
dense_dimensions = 8
agent.create_model(embedding_dimensions=embedding_dimensions,
lstm_dimensions=lstm_dimensions,
dense_dimensions=dense_dimensions,
optimizer=optimizer,
embeddings='glove.840B.300d.txt',
embeddings_trainable=False)
# Visualize the model
try:
plot_model(agent.model, to_file='model.png', show_shapes=True)
except ImportError as e:
logger.warning("Couldn't print the model image: {}".format(e))
# Iteratively train the agent on a batch of previously seen examples while continuously expanding the experience buffer
# This example seems to converge to the optimal reward of 19.4
epochs = 1
for i in range(epochs):
logger.info('Epoch %s', i)
agent.train_online(episodes=256, batch_size=64, gamma=0.95, epsilon_decay=0.99, prioritized_fraction=0.25)
# inspect model weights:
# for layer in agent.model.layers:
# print('layer', layer.name)
# print(layer.get_weights())
# -
| pyfiction/examples/savingjohn/Pretrained embeddings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.7.1
# language: julia
# name: julia-1.7
# ---
# # Gaussian Elimination and LU Decomposition
versioninfo()
# * Goal: solve linear equation
# $$
# \mathbf{A} \mathbf{x} = \mathbf{b}.
# $$
# For simplicity we consider a square matrix $\mathbf{A} \in \mathbb{R}^{n \times n}$.
#
# * History: Chinese mathematical text [The Nine Chapters on the Mathematical Art](https://en.wikipedia.org/wiki/The_Nine_Chapters_on_the_Mathematical_Art), <NAME> and <NAME>.
#
# <img src="./google_doodle_gauss.png">
#
# * A [toy example](https://en.wikipedia.org/wiki/Gaussian_elimination#Example_of_the_algorithm).
A = [2.0 1.0 -1.0; -3.0 -1.0 2.0; -2.0 1.0 2.0]
b = [8.0, -11.0, -3.0]
# Julia way to solve linear equation
# equivalent to `solve(A, b)` in R
A \ b
# What happens when we call `A \ b` to solve a linear equation?
# ## Elementary operator matrix
#
# * **Elementary operator matrix** is the identity matrix with the 0 in position $(j,k)$ replaced by $c$
# $$
# \mathbf{E}_{jk}(c) = \begin{pmatrix}
# 1 & & \\
# & \ddots & \\
# & & 1 & \\
# & & & \ddots & \\
# & & c & & 1 & \\
# & & & & & \ddots \\
# & & & & & & 1
# \end{pmatrix} = \mathbf{I} + c \mathbf{e}_j \mathbf{e}_k^T.
# $$
#
# * $\mathbf{E}_{jk}(c)$ is unit triangular, full rank, and its inverse is
# $$
# \mathbf{E}_{jk}^{-1}(c) = \mathbf{E}_{jk}(-c).
# $$
#
# * $\mathbf{E}_{jk}(c)$ left-multiplies an $n \times m$ matrix $\mathbf{X}$ effectively replace its $j$-th row $\mathbf{x}_{j\cdot}$ by $c \mathbf{x}_{k \cdot} + \mathbf{x}_{j \cdot}$
# $$
# \mathbf{E}_{jk}(c) \times \mathbf{X} = \mathbf{E}_{jk}(c) \times \begin{pmatrix}
# & & \\
# \cdots & \mathbf{x}_{k\cdot} & \cdots \\
# & & \\
# \cdots & \mathbf{x}_{j\cdot} & \cdots \\
# & &
# \end{pmatrix} = \begin{pmatrix}
# & & \\
# \cdots & \mathbf{x}_{k\cdot} & \cdots \\
# & & \\
# \cdots & c \mathbf{x}_{k\cdot} + \mathbf{x}_{j\cdot} & \cdots \\
# & &
# \end{pmatrix}.
# $$
# $2m$ flops.
#
# * Gaussian elimination applies a sequence of elementary operator matrices to transform the linear system $\mathbf{A} \mathbf{x} = \mathbf{b}$ to an upper triangular system
# $$
# \begin{eqnarray*}
# \mathbf{E}_{n,n-1}(c_{n,n-1}) \cdots \mathbf{E}_{21}(c_{21}) \mathbf{A} \mathbf{x} &=& \mathbf{E}_{n,n-1}(c_{n,n-1}) \cdots \mathbf{E}_{21}(c_{21}) \mathbf{b} \\
# \mathbf{U} \mathbf{x} &=& \mathbf{b}_{\text{new}}.
# \end{eqnarray*}
# $$
#
# Column 1:
E21 = [1.0 0.0 0.0; 1.5 1.0 0.0; 0.0 0.0 1.0]
# zero (2, 1) entry
E21 * A
E31 = [1.0 0.0 0.0; 0.0 1.0 0.0; 1.0 0.0 1.0]
# zero (3, 1) entry
E31 * E21 * A
# Column 2:
E32 = [1.0 0.0 0.0; 0.0 1.0 0.0; 0.0 -4.0 1.0]
# zero (3, 2) entry
E32 * E31 * E21 * A
# ## Gauss transformations
#
# * For the first column,
# $$
# \mathbf{M}_1 = \mathbf{E}_{n1}(c_{n1}) \cdots \mathbf{E}_{31}(c_{31}) \mathbf{E}_{21}(c_{21}) = \begin{pmatrix}
# 1 & \\
# c_{21} & \\
# & \ddots & \\
# c_{n1} & & 1
# \end{pmatrix}
# $$
# For the $k$-th column,
# $$
# \mathbf{M}_k = \mathbf{E}_{nk}(c_{nk}) \cdots \mathbf{E}_{k+1,k}(c_{k+1,k}) = \begin{pmatrix}
# 1 & \\
# & \ddots \\
# & & 1 & \\
# & & c_{k+1,k} & 1\\
# & & \vdots & & \ddots \\
# & & c_{n,k} & & & 1
# \end{pmatrix}.
# $$
#
# * $\mathbf{M}_1, \ldots, \mathbf{M}_{n-1}$ are called the **Gauss transformations**.
M1 = E31 * E21
M2 = E32
# * Gauss transformations $\mathbf{M}_k$ are unit triangular, full rank, with inverse
# $$
# \mathbf{M}_k^{-1} = \mathbf{E}_{k+1,k}^{-1}(c_{k+1,k}) \cdots \mathbf{E}_{nk}^{-1}(c_{nk}) = \begin{pmatrix}
# 1 & \\
# & \ddots \\
# & & 1 & \\
# & & - c_{k+1,k}\\
# & & \vdots & & \ddots \\
# & & - c_{n,k} & & & 1
# \end{pmatrix}.
# $$
inv(M1)
inv(M2)
# ## LU decomposition
#
# Gaussian elimination does
# $$
# \mathbf{M}_{n-1} \cdots \mathbf{M}_1 \mathbf{A} = \mathbf{U}.
# $$
# Let
# \begin{equation*}
# \mathbf{L} = \mathbf{M}_1^{-1} \cdots \mathbf{M}_{n-1}^{-1} = \begin{pmatrix}
# 1 & & & & \\
# \,- c_{21} & \ddots & & & \\
# & & 1 & & \\
# \, - c_{k+1,1} & & - c_{k+1,k} & & \\
# & & \vdots & & \ddots \\
# \,- c_{n1} & & - c_{nk} & & & 1
# \end{pmatrix}.
# \end{equation*}
# Thus we have the **LU decomposition**
# $$
# \mathbf{A} = \mathbf{L} \mathbf{U},
# $$
# where $\mathbf{L}$ is unit lower triangular and $\mathbf{U}$ is upper triangular.
# collect negative multipliers into a unit lower triangular matrix
L = [1 0 0; -3/2 1 0; -1 4 1]
# upper triangular matrix after Gaussian elimination
U = [2 1 -1; 0 1/2 1/2; 0 0 -1]
# recovers original matrix
L * U
# * The whole LU algorithm is done in place, i.e., $\mathbf{A}$ is overwritten by $\mathbf{L}$ and $\mathbf{U}$.
#
# * LU decomposition exists if the principal sub-matrix $\mathbf{A}[1:k, 1:k]$ is non-singular for $k=1,\ldots,n-1$.
#
# * If the LU decomposition exists and $\mathbf{A}$ is non-singular, then the LU decmpositon is unique and
# $$
# \det(\mathbf{A}) = \det(\mathbf{L}) \det(\mathbf{U}) = \prod_{k=1}^n u_{kk}.
# $$
#
# * The LU decomposition costs
# $$
# 2(n-1)^2 + 2(n-2)^2 + \cdots + 2 \cdot 1^2 \approx \frac 23 n^3 \quad \text{flops}.
# $$
#
# <img src="http://www.netlib.org/utk/papers/factor/_25826_figure159.gif" width="500" align="center"/>
#
# * Actual implementations can differ: outer product LU ($kij$ loop), block outer product LU (higher level-3 fraction), Crout's algorithm ($jki$ loop).
#
# * Given LU decomposition $\mathbf{A} = \mathbf{L} \mathbf{U}$, solving $(\mathbf{L} \mathbf{U}) \mathbf{x} = \mathbf{b}$ for one right hand side costs $2n^2$ flops:
# - One forward substitution ($n^2$ flops) to solve
# $$
# \mathbf{L} \mathbf{y} = \mathbf{b}
# $$
# - One back substitution ($n^2$ flops) to solve
# $$
# \mathbf{U} \mathbf{x} = \mathbf{y}
# $$
#
# * Therefore to solve $\mathbf{A} \mathbf{x} = \mathbf{b}$ via LU, we need a total of
# $$
# \frac 23 n^3 + 2n^2 \quad \text{flops}.
# $$
#
# * If there are multiple right hand sides, LU only needs to be done once.
# ## Matrix inversion
#
# * For matrix inversion, there are $n$ right hand sides $\mathbf{e}_1, \ldots, \mathbf{e}_n$. Taking advantage of 0s reduces $2n^3$ flops to $\frac 43 n^3$ flops. So **matrix inversion via LU** costs
# $$
# \frac 23 n^3 + \frac 43 n^3 = 2n^3 \quad \text{flops}.
# $$
#
# * **No inversion mentality**:
# > **Whenever we see matrix inverse, we should think in terms of solving linear equations.**
#
# We do not compute matrix inverse unless
# 1. it is necessary to compute standard errors
# 2. number of right hand sides is much larger than $n$
# 3. $n$ is small
# ## Pivoting
#
# * Let
# $$
# \mathbf{A} = \begin{pmatrix}
# 0 & 1 \\
# 1 & 0 \\
# \end{pmatrix}.
# $$
# Is there a solution to $\mathbf{A} \mathbf{x} = \mathbf{b}$ for an arbitrary $\mathbf{b}$? Does GE/LU work for $\mathbf{A}$?
#
# * What if, during LU procedure, the **pivot** $a_{kk}$ is 0 or nearly 0 due to underflow?
# Solution: pivoting.
#
# * **Partial pivoting**: before zeroing the $k$-th column, the row with $\max_{i=k}^n |a_{ik}|$ is moved into the $k$-th row.
#
# * LU with partial pivoting yields
# $$
# \mathbf{P} \mathbf{A} = \mathbf{L} \mathbf{U},
# $$
# where $\mathbf{P}$ is a permutation matrix, $\mathbf{L}$ is unit lower triangular with $|\ell_{ij}| \le 1$, and $\mathbf{U}$ is upper triangular.
#
# * Complete pivoting: Do both row and column interchanges so that the largest entry in the sub matrix `A[k:n, k:n]` is permuted to the $(k,k)$-th entry. This yields the decomposition
# $$
# \mathbf{P} \mathbf{A} \mathbf{Q} = \mathbf{L} \mathbf{U},
# $$
# where $|\ell_{ij}| \le 1$.
#
# * LU decomposition with partial pivoting is the most commonly used methods for solving **general** linear systems. Complete pivoting is the most stable but costs more computation. Partial pivoting is stable most of times.
# ## LAPACK and Julia implementation
#
# * LAPACK: [?getrf](http://www.netlib.org/lapack/explore-html/dd/d9a/group__double_g_ecomputational_ga0019443faea08275ca60a734d0593e60.html#ga0019443faea08275ca60a734d0593e60) does $\mathbf{P} \mathbf{A} = \mathbf{L} \mathbf{U}$ (LU decomposition with partial pivoting) in place.
#
# * R: `solve()` implicitly performs LU decomposition (wrapper of LAPACK routine `dgesv`). `solve()` allows specifying a single or multiple right hand sides. If none, it computes the matrix inverse. The `matrix` package contains `lu()` function that outputs `L`, `U`, and `pivot`.
#
# * Julia:
# - [LinearAlgebra.lu](https://docs.julialang.org/en/v1/stdlib/LinearAlgebra/#LinearAlgebra.lu).
# - [LinearAlgebra.lu!](https://docs.julialang.org/en/v1/stdlib/LinearAlgebra/#LinearAlgebra.lu!). In-place version. Input matrix gets overwritten with L and U.
# - Or call LAPACK wrapper function [getrf!](https://docs.julialang.org/en/v1/stdlib/LinearAlgebra/#LinearAlgebra.LAPACK.getrf!) directly.
# - Other LU-related LAPACK wrapper functions: [gesv](https://docs.julialang.org/en/v1/stdlib/LinearAlgebra/#LinearAlgebra.LAPACK.gesv!), [gesvx](https://docs.julialang.org/en/v1/stdlib/LinearAlgebra/#LinearAlgebra.LAPACK.gesvx!), [trtri](https://docs.julialang.org/en/v1/stdlib/LinearAlgebra/#LinearAlgebra.LAPACK.trtri!) (inverse of triangular matrix), [trtrs](https://docs.julialang.org/en/v1/stdlib/LinearAlgebra/#LinearAlgebra.LAPACK.trtrs!).
A
# +
using LinearAlgebra
# second argument indicates partial pivoting (default) or not
alu = lu(A)
typeof(alu)
# -
dump(alu)
alu.L
alu.U
alu.p
alu.P
alu.L * alu.U
A[alu.p, :]
# this is doing two triangular solves, 2n^2 flops
alu \ b
det(A) # this does LU! O(n^3)
det(alu) # this is cheap O(n)
inv(A) # this does LU! O(n^3)
inv(alu) # this is cheap O(n^2)
# ## Further reading
#
# * Sections II.5.2 and II.5.3 of [Computational Statistics](http://ucla.worldcat.org/title/computational-statistics/oclc/437345409&referer=brief_results) by <NAME> (2010).
#
# * A definite reference is Chapter 3 of the book [Matrix Computation](http://catalog.library.ucla.edu/vwebv/holdingsInfo?bibId=7122088) by <NAME> and <NAME>.
#
# <img src="https://images-na.ssl-images-amazon.com/images/I/41Cs04RRiTL._SX309_BO1,204,203,200_.jpg" width="250" align="center"/>
#
# <img src="https://images-na.ssl-images-amazon.com/images/I/41f5vxegABL._SY344_BO1,204,203,200_.jpg" width="250" align="center"/>
| slides/11-gelu/gelu.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Import libraries
import pandas as pd
# Get data
file_name = 'https://raw.githubusercontent.com/rajeevratan84/datascienceforbusiness/master/adult.data'
census = pd.read_csv(file_name)
census.head()
# +
# Create dataframe with modified columnames
col_names = ['age', 'workclass', 'fnlwgt', 'education', 'education_num',
'marital_status', 'occupation', 'relationship', 'race', 'sex',
'capital_gain', 'capital_loss', 'hours_per_week', 'native_country', 'income']
census = pd.DataFrame(census.values, columns = col_names)
census.head()
# -
# Basic stats
print(f'Dataframe shape: {census.shape}')
print(f'Missing Values:\n {census.isnull().values.sum()}')
print(f'Unique Values:\n{census.nunique()}')
# Infer objects
census.info()
census = census.infer_objects()
census.info()
census.info()
# +
# Explore Income
census['income'] = census['income'].str.strip()
n = census.shape[0]
n_greater_50k = census.loc[census['income'] == '>50K'].shape[0]
n_less_equal_50k = census.loc[census['income'] == '<=50K'].shape[0]
greater_percent = (n_greater_50k / n) * 100
print(f'Total number of records: {n}')
print(f'Making more than 50K: {n_greater_50k}')
print(f'Making less or equal to 50K: {n_less_equal_50k}')
print(f'Percentage making more than 50K: {round(greater_percent, 2)}')
# +
import seaborn as sns
import matplotlib.pyplot as plt
sns.set(style='whitegrid', color_codes=True)
sns.catplot('sex', col='education', data=census, hue='income', kind='count', col_wrap=4)
# -
# Age histogram
census.hist('age')
# +
# Capital gain loss histogram comparisons
import numpy as np
fig, axes = plt.subplots(1, 2)
census.hist('capital_gain', bins=20, ax=axes[0])
census.hist('capital_loss', bins=20, ax=axes[1])
plt.tight_layout()
# +
# Apply log to capital gains losses
skewed = ['capital_gain', 'capital_loss']
census[skewed] = census[skewed].apply(lambda x: np.log(x + 1))
fig, axes = plt.subplots(1, 2)
census.hist('capital_gain', bins=20, ax=axes[0])
census.hist('capital_loss', bins=20, ax=axes[1])
plt.tight_layout()
# -
census.isin([' ?']).any(axis='rows')
census = census[census['workclass'] != ' ?']
census = census[census['occupation'] != ' ?']
census = census[census['native_country'] != ' ?']
# +
# Prepare data for modelling
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler()
numerical = ['age', 'education_num', 'capital_gain', 'capital_loss', 'hours_per_week']
census_minmax = pd.DataFrame(data=census)
census_minmax[numerical] = scaler.fit_transform(census_minmax[numerical])
census_minmax.head()
# +
# Set income as target and label 1 and 0 for <=50K
from sklearn.preprocessing import LabelEncoder
encoder = LabelEncoder()
income = census_minmax['income'].apply(lambda x: 0 if x == '<=50K' else 1)
target = pd.Series(encoder.fit_transform(income))
# -
# Set features
features = pd.get_dummies(census_minmax.drop('income', axis=1))
features_list = list(features.columns)
print(f'{len(features_list)} total features after get dummies')
# +
# Split data
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(features, target, test_size=0.33)
print(f'X_train shape: {X_train.shape}')
print(f'X_test shape: {X_test.shape}')
# +
from sklearn.metrics import fbeta_score, accuracy_score
from time import time
def train_predict(learner, sample_size, X_train, y_train, X_test, y_test):
'''
inputs:
- learner: the learning algorithm
- sample_size: training set sample size
- X_train: training features
- y_train: training targets
- X_test: testing features
- y_test: testing targets
'''
results = {}
# Fit the training data
start = time()
learner = learner.fit(X_train[:sample_size], y_train[:sample_size])
end = time()
results['training_time'] = end - start
# Get predictions
start = time()
preds_train = learner.predict(X_train[:300])
preds_test = learner.predict(X_test)
end = time()
results['prediction_time'] = end - start
results['training_accuracy'] = accuracy_score(y_train[:300], preds_train)
results['testing_accuracy'] = accuracy_score(y_test, preds_test)
results['training_fscore'] = fbeta_score(y_train[:300], preds_train, 0.5)
results['testing_fscore'] = fbeta_score(y_test, preds_test, 0.5)
# Print algorithm used and sample size
print(f'{learner.__class__.__name__} trained on {sample_size} samples')
return results
# +
# Compare DecisionTree, SVC, and AdaBoost
from sklearn.tree import DecisionTreeClassifier
from sklearn.svm import SVC
from sklearn.ensemble import AdaBoostClassifier
tree_clf = DecisionTreeClassifier()
svc_clf = SVC(gamma='auto')
adaboost_clf = AdaBoostClassifier()
# Set sample sizes for 1%, 10%, and 100%
one_percent = int(round(len(X_train) / 100))
ten_percent = int(round(len(X_train) / 10))
hundred_percent = int(round(len(X_train)))
# Get results
results = {}
for clf in [tree_clf, svc_clf, adaboost_clf]:
clf_name = clf.__class__.__name__
results[clf_name] = {}
for i, samples in enumerate([one_percent, ten_percent, hundred_percent]):
results[clf_name][i] = train_predict(clf, samples, X_train, y_train, X_test, y_test)
# -
for i in results.items():
print(i[0])
display(pd.DataFrame(i[1]).rename(columns={0:'1%', 1:'10%', 2:'100%'}))
# +
# Compare confusion matrix models
from sklearn.metrics import confusion_matrix
plt.figure()
for i, clf in enumerate([tree_clf, svc_clf, adaboost_clf]):
cm = confusion_matrix(y_test, clf.predict(X_test))
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
plt.figure(i)
sns.heatmap(cm, annot=True)
plt.ylabel('Actual')
plt.xlabel('Predicted')
plt.title(clf.__class__.__name__)
plt.show()
# +
# Optimize AdaBoostClassifier
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import make_scorer
clf = AdaBoostClassifier(base_estimator=DecisionTreeClassifier())
# Provide hyperparameters
h_params = {'n_estimators':[50,120],
'learning_rate':[0.1,0.5,1.],
'base_estimator__min_samples_split':np.arange(2,8,2),
'base_estimator__max_depth':np.arange(1,4,1)}
scorer = make_scorer(fbeta_score,beta=0.5)
grid_obj = GridSearchCV(clf, h_params, scorer)
grid_fit = grid_obj.fit(X_train, y_train)
best_clf = grid_fit.best_estimator_
predictions = (clf.fit(X_train, y_train)).predict(X_test)
best_predictions = best_clf.predict(X_test)
print('Unoptimized model\n-----------------')
print(f'Accuracy: {accuracy_score(y_test, predictions):.4f}')
print(f'F-score: {fbeta_score(y_test, predictions, 0.5):.4f}')
print('\nOptimized model\n---------------')
print(f'Accuracy: {accuracy_score(y_test, best_predictions):.4f}')
print(f'F-score: {fbeta_score(y_test, best_predictions, 0.5):.4f}')
print('\nModel\n-----')
print(best_clf)
| CensusPredictions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="nqKIbbV2a9Pf"
# ### Install required libraries
# + id="QorYZPeuLbHO" colab={"base_uri": "https://localhost:8080/"} outputId="65e2d33a-4388-4c3c-b74a-facfaecf1ef0"
# !pip install keras-tuner
# + [markdown] id="360EVSDRbH-k"
# ### Download Dataset
# + id="FxgoXpboha1e" colab={"base_uri": "https://localhost:8080/"} outputId="c4ba9f80-ad34-40e9-bbfa-aa3373f511ee"
# !wget 'https://s3.amazonaws.com/isic-challenge-data/2019/ISIC_2019_Training_GroundTruth.csv'
# + id="-KeFB_PQhUSG" colab={"base_uri": "https://localhost:8080/"} outputId="b48d3e16-4136-431e-b17a-536748e036e5"
# !wget 'https://s3.amazonaws.com/isic-challenge-data/2019/ISIC_2019_Training_Metadata.csv'
# + id="ULKVZWzj56QQ" colab={"base_uri": "https://localhost:8080/"} outputId="719d72f5-0374-4219-a2d1-cac1676dc2a7"
# !wget 'https://s3.amazonaws.com/isic-challenge-data/2019/ISIC_2019_Training_Input.zip'
# + id="Mx9kopXtf47r" colab={"base_uri": "https://localhost:8080/"} outputId="c8b8f1a9-6b7e-420b-c37e-99a3d709045b"
# !unzip 'ISIC_2019_Training_Input.zip'
# + id="E-CRfWJ0Vrjn"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import keras
# + id="UysT9yEJVroV"
labels = pd.read_csv('ISIC_2019_Training_GroundTruth.csv')
label_df = pd.DataFrame(labels)
# + id="rjeXCMc1Vrrc"
info = pd.read_csv('ISIC_2019_Training_Metadata.csv')
info_df = pd.DataFrame(info)
# + id="xqoWqXhgVruc" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="72c43117-ca54-4d05-f1df-0ea607bf68bc"
label_df.tail()
# + id="OuNLr8rPh3DO" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="3b133df0-2606-414c-ae35-c9cb8e804c70"
info_df.head()
# + id="A-FY9P28YbvI"
info_df = info_df.drop('image',axis=1)
# + id="K3wIS9vPVr8t" colab={"base_uri": "https://localhost:8080/", "height": 402} outputId="e6a0817d-3e9d-4421-f47b-367aa4e9c082"
info_df
# + id="wK4KHiVxXsem"
data = pd.concat([label_df, info_df], axis=1)
# + id="PHy1xSaXXsgy" colab={"base_uri": "https://localhost:8080/", "height": 343} outputId="d40d4baf-7086-4770-f076-54829c48b815"
data.head(10)
# + id="iNgexiXPZGMa"
data['lesion_id'].fillna(method ='bfill', inplace=True)
data["age_approx"].fillna(30.0, inplace = True)
data["sex"].fillna("male", inplace = True)
data["anatom_site_general"].fillna( method ='ffill', inplace = True)
# + id="T9ZSCfoNLS9r" colab={"base_uri": "https://localhost:8080/", "height": 639} outputId="fdd8486a-28bf-4957-d660-ad4ca9708e10"
data.head(20)
# + id="2Cst5RU5Jo-z" colab={"base_uri": "https://localhost:8080/"} outputId="32775e4a-db45-455a-80b1-2bda7d5b0048"
rows_with_nan = []
for index, row in data.iterrows():
is_nan_series = row.isnull()
if is_nan_series.any():
rows_with_nan.append(index)
print(rows_with_nan)
# + id="PYPY2064dd5C" colab={"base_uri": "https://localhost:8080/"} outputId="2c8221d1-ceb4-468f-e5be-801ef9da414a"
data['anatom_site_general'].unique()
# + id="19ZwrjLReOya"
anatom_site_general = {'anterior torso': 1,'upper extremity': 2,'posterior torso':3,'lower extremity':4, 'lateral torso':5,'head/neck':6,'palms/soles':7,'oral/genital':8}
data['anatom_site_general'] = [anatom_site_general[item] for item in data['anatom_site_general']]
# + id="6sP8dOKPD4qM"
sex = {'male': 0,'female':1}
data['sex'] = [sex[item] for item in data['sex']]
# + id="8j6kl9A3vxl9" colab={"base_uri": "https://localhost:8080/"} outputId="fe3dc35e-531b-41c3-c584-7804521b8138"
len(data['lesion_id'].unique())
# + id="omH03AXID0cz" colab={"base_uri": "https://localhost:8080/", "height": 225} outputId="164848b2-cfec-4f24-c868-96f1d26d3aa8"
data.head(6)
# + id="_aBWlkGIvw4C"
data = data.drop(['lesion_id'],axis=1)
# + id="rpYN7xLBZIeh"
target = data[['MEL']].values
# + id="__NAZhrsYV_a"
data = data.drop(['image','MEL','NV'],axis=1)
# + id="LqvPEh_HY7-z" colab={"base_uri": "https://localhost:8080/", "height": 402} outputId="728ef7cf-f26d-4597-817c-87716ca742f9"
data
# + id="bV667Gp4Y8EQ"
label = target
# + id="yW7RZQdSY8Gw" colab={"base_uri": "https://localhost:8080/"} outputId="7a4157a1-9bbb-469d-e750-5a10e6df9faa"
list0 = [data, label]
list1 = ['x_train','y_train']
for i in range(2):
print('The shape of the {} is {}'.format(list1[i],list0[i].shape))
# + id="LvqWm02BY8JX" colab={"base_uri": "https://localhost:8080/"} outputId="16638fc2-ae2b-4700-804d-6562ea6f0253"
_,D = data.shape
print(D)
# + id="p6zw95gai0um"
from google.colab import files
import cv2
# + [markdown] id="_FymV5RtPtAG"
# **I use this part to upload the downloaded images instead of download them in the colab.**
# + id="uAjeudjQY8ke" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="c9718323-e30e-4862-e24d-171ab776f4cb"
'''uploaded = files.upload()
train_image = []
for i in uploaded.keys():
train_image.append(cv2.resize(cv2.cvtColor(cv2.imread(i), cv2.COLOR_BGR2RGB), (32,32)))'''
# + id="3ayVcghzFwPx"
# Upload the images from folder
import os
def load_images_from_folder(folder):
train_image = []
for filename in os.listdir(folder):
img = cv2.imread(os.path.join(folder,filename))
if img is not None:
train_image.append(cv2.resize(cv2.cvtColor(img, cv2.COLOR_BGR2RGB), (32,32)))
return train_image
# + id="ABlV0xUvnIbC"
images = load_images_from_folder('ISIC_2019_Training_Input')
# + id="fwDcrSq89bpP"
from sklearn.preprocessing import StandardScaler
# + id="vy9NF6QV9bs0"
train_image = images[:20264]
test_image = images[20264:]
x_train = data[:20264]
x_test = data[20264:]
y_train = label[:20264]
y_test = label[20264:]
# + id="4zzRYUtzBrLF"
scaler = StandardScaler()
x_train = scaler.fit_transform(x_train)
x_test = scaler.fit_transform(x_test)
# + id="ccQ04O45BVNe" colab={"base_uri": "https://localhost:8080/", "height": 269} outputId="7a62f99f-a6eb-46ae-b4ec-eb5eef11c073"
i = 2
plt.imshow(train_image[i])
plt.xticks([])
plt.yticks([])
plt.show()
print(y_train[i])
# + id="sT03PmLl8ck0" colab={"base_uri": "https://localhost:8080/", "height": 269} outputId="432f269a-d304-4086-a8f3-a23ab6847c84"
i = 0
plt.imshow(test_image[i])
plt.xticks([])
plt.yticks([])
plt.show()
print(y_test[i])
# + id="cBmD6_MlkD7a"
train_image = np.asarray(train_image)
test_image = np.asarray(test_image)
# + id="xXMiu6AiY8nV"
train_image = train_image.astype('float32')
test_image = test_image.astype('float32')
# + id="tzHAOr2gY8v-"
mean = np.mean(train_image,axis=(0,1,2,3))
std = np.std(train_image,axis=(0,1,2,3))
train_image = (train_image-mean)/(std+1e-7)
test_image = (test_image-mean)/(std+1e-7)
# + id="dAF_-z0yY80v"
from keras.utils import np_utils
nClasses = 2
y_train = np_utils.to_categorical(y_train, nClasses)
y_test = np_utils.to_categorical(y_test, nClasses)
# + id="dyqmNDSdY8zI" colab={"base_uri": "https://localhost:8080/"} outputId="ddfe83d4-2b64-4b6f-9c0c-1cac1034c694"
print(test_image.shape)
print(y_train.shape)
print(y_test.shape)
# + id="5AbBn4b8Y8il"
input_shape = (32,32,3)
# + id="z52pZFwIjXm0"
from keras import layers
from keras.models import Model
from keras.models import Sequential
from keras.layers import Dense, Dropout , Input , Flatten , Conv2D , MaxPooling2D
from keras.layers.merge import concatenate
from keras.callbacks import EarlyStopping , ModelCheckpoint
from keras.optimizers import Adam, SGD, RMSprop
from keras import regularizers
from kerastuner.tuners import RandomSearch
# + id="ME-mhpTpllH4"
# + id="omX51bV-lx7k"
def build_model(hp):
# model_1
model1_in = keras.Input(shape=(32,32,3))
x = layers.Conv2D(64,(2,2),padding='same', activation='relu')(model1_in)
x = layers.Conv2D(64,(2,2), activation='relu')(x)
x = layers.MaxPooling2D(pool_size=(2, 2))(x)
x = layers.Conv2D(128,(2,2),padding='same', activation='relu')(x)
x = layers.Conv2D(128,(2,2), activation='relu')(x)
x = layers.MaxPooling2D(pool_size=(2,2))(x)
x = layers.Conv2D(256,(2,2),padding='same', activation='relu')(x)
x = layers.Conv2D(256,(2,2), activation='relu')(x)
x = layers.MaxPooling2D(pool_size=(2,2))(x)
x = layers.Conv2D(512,(2,2),padding='same', activation='relu')(x)
x = layers.Conv2D(512,(2,2), activation='relu')(x)
x = layers.MaxPooling2D(pool_size=(2,2))(x)
x = layers.Flatten()(x)
x = layers.Dense(256, activation='relu')(x)
x = layers.Dense(512, activation='relu')(x)
x = layers.Dense(1024, activation='relu')(x)
model1_out = layers.Dense(2, activation='sigmoid')(x)
model1 = keras.Model(model1_in, model1_out)
# model_2
model2_in = keras.Input(shape=(D,))
x = layers.Dense(16384, kernel_initializer='normal')(model2_in)
x = layers.BatchNormalization()(x)
x = layers.Activation('relu')(x)
x = layers.Dropout(0.1)(x)
x = layers.Dense(8192, kernel_initializer='normal')(x)
x = layers.BatchNormalization()(x)
x = layers.Activation('relu')(x)
x = layers.Dropout(0.1)(x)
x = layers.Dense(4096, kernel_initializer='normal')(x)
x = layers.BatchNormalization()(x)
x = layers.Activation('relu')(x)
x = layers.Dropout(0.1)(x)
x = layers.Dense(2048, kernel_initializer='normal')(x)
x = layers.BatchNormalization()(x)
x = layers.Activation('relu')(x)
x = layers.Dropout(0.1)(x)
x = layers.Dense(1024, kernel_initializer='normal')(x)
x = layers.BatchNormalization()(x)
x = layers.Activation('relu')(x)
x = layers.Dropout(0.1)(x)
x = layers.Dense(512, kernel_initializer='normal')(x)
x = layers.BatchNormalization()(x)
x = layers.Activation('relu')(x)
x = layers.Dropout(0.1)(x)
x = layers.Dense(128, kernel_initializer='normal')(x)
x = layers.BatchNormalization()(x)
x = layers.Activation('relu')(x)
x = layers.Dropout(0.1)(x)
x = layers.Dense(64, kernel_initializer='normal')(x)
x = layers.BatchNormalization()(x)
x = layers.Activation('relu')(x)
x = layers.Dropout(0.1)(x)
x = layers.Dense(32, kernel_initializer='normal')(x)
x = layers.BatchNormalization()(x)
x = layers.Activation('relu')(x)
x = layers.Dropout(0.1)(x)
x = layers.Dense(16, kernel_initializer='normal')(x)
x = layers.BatchNormalization()(x)
x = layers.Activation('relu')(x)
x = layers.Dropout(0.1)(x)
x = layers.Dense(4, kernel_initializer='normal')(x)
x = layers.BatchNormalization()(x)
x = layers.Activation('relu')(x)
x = layers.Dropout(0.1)(x)
model2_out = layers.Dense(2, kernel_initializer='normal')(x)
model2 = keras.Model(model2_in, model2_out)
concatenated = concatenate([model1_out, model2_out])
x = layers.Dense(units=hp.Int('units', min_value=32, max_value=512, step=32), activation='relu')(concatenated)
out = Dense(2, activation='sigmoid', name='output_layer',kernel_regularizer=regularizers.l1_l2(l1=1e-5, l2=1e-4))(x)
merged_model = Model([model1_in, model2_in], out)
merged_model.compile(optimizer=keras.optimizers.Adam(hp.Choice('learning_rate',values=[1e-2, 1e-3, 1e-4])),loss='binary_crossentropy',metrics=['accuracy'])
return merged_model
tuner = RandomSearch(build_model,objective='val_accuracy',max_trials=5,executions_per_trial=3,directory='my_dir',project_name='helloworld')
# + id="MwdRo9TlL2ES" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="7ea7d78b-ecef-4c77-c080-0fe9b61cfba6"
tuner.search_space_summary()
# + id="5p14DcsQL54s" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="f254f2cf-760b-4a18-b948-9a690d259572"
final=tuner.search([train_image, x_train], y=y_train, batch_size=32, epochs=5,verbose=1,validation_data=([test_image,x_test],y_test))
# + id="aENSTsO3VwXi"
best_model = tuner.get_best_models()[0]
# + id="Rvccq4XsWWFF" colab={"base_uri": "https://localhost:8080/", "height": 896} outputId="411c5646-7cae-4bfb-984b-0657f6e4fc01"
tuner.results_summary()
# + id="fiWDD39Lpon6" colab={"base_uri": "https://localhost:8080/"} outputId="d89b57fc-5410-498a-d31a-31f00164ef96"
best_model.evaluate([test_image,x_test],y_test)
| project_melanomia_keras_tuner.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Developing an AI application
#
# Going forward, AI algorithms will be incorporated into more and more everyday applications. For example, you might want to include an image classifier in a smart phone app. To do this, you'd use a deep learning model trained on hundreds of thousands of images as part of the overall application architecture. A large part of software development in the future will be using these types of models as common parts of applications.
#
# In this project, you'll train an image classifier to recognize different species of flowers. You can imagine using something like this in a phone app that tells you the name of the flower your camera is looking at. In practice you'd train this classifier, then export it for use in your application. We'll be using [this dataset](http://www.robots.ox.ac.uk/~vgg/data/flowers/102/index.html) of 102 flower categories, you can see a few examples below.
#
# <img src='assets/Flowers.png' width=500px>
#
# The project is broken down into multiple steps:
#
# * Load and preprocess the image dataset
# * Train the image classifier on your dataset
# * Use the trained classifier to predict image content
#
# We'll lead you through each part which you'll implement in Python.
#
# When you've completed this project, you'll have an application that can be trained on any set of labeled images. Here your network will be learning about flowers and end up as a command line application. But, what you do with your new skills depends on your imagination and effort in building a dataset. For example, imagine an app where you take a picture of a car, it tells you what the make and model is, then looks up information about it. Go build your own dataset and make something new.
#
# First up is importing the packages you'll need. It's good practice to keep all the imports at the beginning of your code. As you work through this notebook and find you need to import a package, make sure to add the import up here.
# +
# Imports here
import torch
import torchvision
from torch import nn, optim
from torchvision import transforms
from torchvision.datasets import ImageFolder
from torch.utils import data
from PIL import Image
import numpy as np
import os, random
import json
import signal
from contextlib import contextmanager
import requests
import matplotlib
import matplotlib.pyplot as plt
# -
# ## Load the data
#
# Here you'll use `torchvision` to load the data ([documentation](http://pytorch.org/docs/0.3.0/torchvision/index.html)). The data should be included alongside this notebook, otherwise you can [download it here](https://s3.amazonaws.com/content.udacity-data.com/nd089/flower_data.tar.gz). The dataset is split into three parts, training, validation, and testing. For the training, you'll want to apply transformations such as random scaling, cropping, and flipping. This will help the network generalize leading to better performance. You'll also need to make sure the input data is resized to 224x224 pixels as required by the pre-trained networks.
#
# The validation and testing sets are used to measure the model's performance on data it hasn't seen yet. For this you don't want any scaling or rotation transformations, but you'll need to resize then crop the images to the appropriate size.
#
# The pre-trained networks you'll use were trained on the ImageNet dataset where each color channel was normalized separately. For all three sets you'll need to normalize the means and standard deviations of the images to what the network expects. For the means, it's `[0.485, 0.456, 0.406]` and for the standard deviations `[0.229, 0.224, 0.225]`, calculated from the ImageNet images. These values will shift each color channel to be centered at 0 and range from -1 to 1.
#
data_dir = 'flowers'
train_dir = data_dir + '/train'
valid_dir = data_dir + '/valid'
test_dir = data_dir + '/test'
# +
# TODO: Define your transforms for the training, validation, and testing sets
# TODO: Define your transforms for the training, validation, and testing sets
train_trsfs = transforms.Compose([transforms.RandomRotation(30),
transforms.RandomResizedCrop(224),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225])
])
valid_test_trsfs = transforms.Compose([transforms.Resize(255),
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225])
])
# TODO: Load the datasets with ImageFolder
train_data = ImageFolder(root=train_dir, transform=train_trsfs)
valid_data = ImageFolder(root=valid_dir, transform=valid_test_trsfs)
test_data = ImageFolder(root=test_dir, transform=valid_test_trsfs)
# TODO: Using the image datasets and the trainforms, define the dataloaders
train_data_loader = data.DataLoader(train_data, batch_size=64, shuffle=True)
valid_data_loader = data.DataLoader(valid_data, batch_size=64)
test_data_loader = data.DataLoader(test_data, batch_size=64)
# -
# ### Label mapping
#
# You'll also need to load in a mapping from category label to category name. You can find this in the file `cat_to_name.json`. It's a JSON object which you can read in with the [`json` module](https://docs.python.org/2/library/json.html). This will give you a dictionary mapping the integer encoded categories to the actual names of the flowers.
# +
import json
with open('cat_to_name.json', 'r') as f:
cat_to_name = json.load(f)
# -
# # Building and training the classifier
#
# Now that the data is ready, it's time to build and train the classifier. As usual, you should use one of the pretrained models from `torchvision.models` to get the image features. Build and train a new feed-forward classifier using those features.
#
# We're going to leave this part up to you. Refer to [the rubric](https://review.udacity.com/#!/rubrics/1663/view) for guidance on successfully completing this section. Things you'll need to do:
#
# * Load a [pre-trained network](http://pytorch.org/docs/master/torchvision/models.html) (If you need a starting point, the VGG networks work great and are straightforward to use)
# * Define a new, untrained feed-forward network as a classifier, using ReLU activations and dropout
# * Train the classifier layers using backpropagation using the pre-trained network to get the features
# * Track the loss and accuracy on the validation set to determine the best hyperparameters
#
# We've left a cell open for you below, but use as many as you need. Our advice is to break the problem up into smaller parts you can run separately. Check that each part is doing what you expect, then move on to the next. You'll likely find that as you work through each part, you'll need to go back and modify your previous code. This is totally normal!
#
# When training make sure you're updating only the weights of the feed-forward network. You should be able to get the validation accuracy above 70% if you build everything right. Make sure to try different hyperparameters (learning rate, units in the classifier, epochs, etc) to find the best model. Save those hyperparameters to use as default values in the next part of the project.
#
# One last important tip if you're using the workspace to run your code: To avoid having your workspace disconnect during the long-running tasks in this notebook, please read in the earlier page in this lesson called Intro to
# GPU Workspaces about Keeping Your Session Active. You'll want to include code from the workspace_utils.py module.
#
# **Note for Workspace users:** If your network is over 1 GB when saved as a checkpoint, there might be issues with saving backups in your workspace. Typically this happens with wide dense layers after the convolutional layers. If your saved checkpoint is larger than 1 GB (you can open a terminal and check with `ls -lh`), you should reduce the size of your hidden layers and train again.
# +
# TODO: Build and train your network
# Load pretrained vgg16 Model
#vgg16 have several advantages: It has so many weight parameters, the models are very heavy, 550 MB + of weight size,
#Which also means long inference time
#source (https://forums.fast.ai/t/vgg-strength-and-limitations/1218)
model = models.vgg16(pretrained=True)
#When a model is loaded in PyTorch, all its parameters have their ‘requires_grad‘ field set to true by default.
#That means each and every change to the parameter values will be stored in order to be used in the back propagation graph
#used for training. This increases memory requirements. So,
#since most of the parameters in our pre-trained model are already trained for us, we reset the requires_grad field to false.
#to freeze part of the convolutional part of vgg16 model and train the rest
for param in model.parameters():
param.requires_grad = False
#Then we replace the final layer of the vgg16 model by a small set of Sequential layers.
#The inputs to the last fully connected layer of vgg16 is fed to a Linear layer,
#which are then fed into ReLU (rectified linear unit) and Dropout layers.
#Used OrderedDict to preserve the order in which the keys are inserted
from collections import OrderedDict
classifier = nn.Sequential(OrderedDict([
('inputs', nn.Linear(25088, 120)), #hidden layer 1 sets output to 120
('relu1', nn.ReLU()),
('dropout',nn.Dropout(0.5)), #could use a different droupout probability,but 0.5 usually works well
('hidden_layer1', nn.Linear(120, 90)), #hidden layer 2 output to 90
('relu2',nn.ReLU()),
('hidden_layer2',nn.Linear(90,70)), #hidden layer 3 output to 70
('relu3',nn.ReLU()),
('hidden_layer3',nn.Linear(70,102)),#output size = 102
('output', nn.LogSoftmax(dim=1))]))# For using NLLLoss()
model.classifier = classifier
#Since we will be doing the training on a GPU, we get the model ready for GPU.
# Convert model to be used on GPU
if torch.cuda.is_available():
model.cuda()
#define the criterion and the optimizer to be used for training.
#Negative Loss Likelihood function(NLLLoss()) as it can be used for classifying multiple classes.
#Adam is one the most popular optimizers because it can adapt the learning rate for each parameter individually.
criterion = nn.NLLLoss()
optimizer = optim.Adam(model.classifier.parameters(), lr=0.001)
# +
#Training is carried out for a fixed set of epochs, processing each image once in a single epoch.
#The training data loader loads data in batches.
#In our case, we have given a batch size of 64, which means each batch can have at most 64 images.
#with transfer learning, we need to compute gradients only for a small set of parameters
#that belong to the few newly added layers towards the end of the model.
#Gradient computation is done using the autograd and back propagation.
#PyTorch accumulates all the gradients in the backward pass.
# So it is essential to zero them out at the beginning of the training loop.
#This is achieved using the optimizer’s zero_grad function.
#Finally, after the gradients are computed in the backward pass, the parameters are updated using the optimizer’s step function.
#Total loss and accuracy is computed for the whole batch,
#which is then averaged over all the batches to get the loss and accuracy values for the whole epoch.
epochs=18
for epoch in range(epochs):
print("Epoch: {}/{}".format(epoch+1, epochs))
# tell the model that you are training . So effectively layers like dropout, batchnorm etc,
#which behave different on the train and test procedures know what is going on and hence can behave accordingly.
# Set to training mode
# Set to training mode
model.train()
# Loss and Accuracy within the epoch
train_loss = 0.0
train_acc = 0.0
valid_loss = 0.0
valid_acc = 0.0
for i, (inputs, labels) in enumerate(train_loader):
inputs = inputs.to('cuda')
labels = labels.to('cuda')
# Clean existing gradients
optimizer.zero_grad()
# Forward pass - compute outputs on input data using the model
outputs = model(inputs)
# Compute loss
loss = criterion(outputs, labels)
# Backpropagate the gradients
loss.backward()
# Update the parameters
optimizer.step()
# Compute the total loss for the batch and add it to train_loss
train_loss += loss.item() * inputs.size(0)
# Compute the accuracy
ret, predictions = torch.max(outputs.data, 1)
correct_counts = predictions.eq(labels.data.view_as(predictions))
# Convert correct_counts to float and then compute the mean
acc = torch.mean(correct_counts.type(torch.FloatTensor))
# Compute total accuracy in the whole batch and add to train_acc
train_acc += acc.item() * inputs.size(0)
print("Batch no: {:03d}, Loss on trainig: {:.4f}, Accuracy: {:.4f}".format(i, loss.item(), acc.item()))
# +
# TODO: Do validation on the test set
#Maintaining a separate validation set is important,
#so that we can stop the training at the right point and prevent overfitting.
#Validation is carried out in each epoch immediately after the training loop.
#Since we do not need any gradient computation in the validation process, it is done with torch.no_grad().
#For each validation batch, the inputs and labels are transferred to the GPU ( if cuda is available, else cpu).
#The inputs go through the forwards pass,
#followed by the loss and accuracy computations for the batch and at the end of the loop, for the whole epoch.
# Validation - No gradient tracking needed
with torch.no_grad():
# Set to evaluation mode
model.eval()
# Validation loop
for j, (inputs, labels) in enumerate(validation_loader):
inputs = inputs.to('cuda')
labels = labels.to('cuda')
# Forward pass - compute outputs on input data using the model
outputs = model(inputs)
# Compute loss
loss = criterion(outputs, labels)
# Compute the total loss for the batch and add it to valid_loss
valid_loss += loss.item() * inputs.size(0)
# Calculate validation accuracy
ret, predictions = torch.max(outputs.data, 1)
correct_counts = predictions.eq(labels.data.view_as(predictions))
# Convert correct_counts to float and then compute the mean
acc = torch.mean(correct_counts.type(torch.FloatTensor))
# Compute total accuracy in the whole batch and add to valid_acc
valid_acc += acc.item() * inputs.size(0)
print("Validation Batch number: {:03d}, Validation: Loss: {:.4f}, Accuracy: {:.4f}".format(j, loss.item(), acc.item()))
# -
# ## Testing your network
#
# It's good practice to test your trained network on test data, images the network has never seen either in training or validation. This will give you a good estimate for the model's performance on completely new images. Run the test images through the network and measure the accuracy, the same way you did validation. You should be able to reach around 70% accuracy on the test set if the model has been trained well.
# +
# Do validation on the test set
correct,total = 0,0
with torch.no_grad():
model.eval()
for data in train_loader:
images, labels = data
images, labels = images.to('cuda'), labels.to('cuda')
outputs = model(images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy on test images is: %d%%' % (100 * correct / total))
# -
# ## Save the checkpoint
#
# Now that your network is trained, save the model so you can load it later for making predictions. You probably want to save other things such as the mapping of classes to indices which you get from one of the image datasets: `image_datasets['train'].class_to_idx`. You can attach this to the model as an attribute which makes inference easier later on.
#
# ```model.class_to_idx = image_datasets['train'].class_to_idx```
#
# Remember that you'll want to completely rebuild the model later so you can use it for inference. Make sure to include any information you need in the checkpoint. If you want to load the model and keep training, you'll want to save the number of epochs as well as the optimizer state, `optimizer.state_dict`. You'll likely want to use this trained model in the next part of the project, so best to save it now.
# +
# TODO: Save the checkpoint
#The class ImageFolder has an attribute class_to_idx which is a dictionary mapping the name of the class to the index (label).
#So, you can access the classes with data.classes and for each class get the label with data.class_to_idx.
#torch.save: Saves a serialized object to disk. This function uses Python’s pickle utility for serialization.
#Models, tensors, and dictionaries of all kinds of objects can be saved using this function.
#torch.save(model, PATH)
#reference:https://pytorch.org/tutorials/beginner/saving_loading_models.html
model.class_to_idx = image_datasets['train'].class_to_idx
torch.save({'structure' :'alexnet',
'hidden_layer1':120,
'droupout':0.5,
'epochs':12,
'state_dict':model.state_dict(),
'class_to_idx':model.class_to_idx,
'optimizer_dict':optimizer.state_dict()},
'checkpoint.pth')
# -
# ## Loading the checkpoint
#
# At this point it's good to write a function that can load a checkpoint and rebuild the model. That way you can come back to this project and keep working on it without having to retrain the network.
# +
# TODO: Write a function that loads a checkpoint and rebuilds the model
#torch.load: Uses pickle’s unpickling facilities to deserialize pickled object files to memory.
#This function also facilitates the device to load the data into
#When you call torch.load() on a file which contains GPU tensors,
#those tensors will be loaded to GPU by default. You can call torch.load(.., map_location='cpu')
# and then load_state_dict() to avoid GPU RAM surge when loading a model checkpoint.
##reference:https://pytorch.org/tutorials/beginner/saving_loading_models.html
def loading_the_checkpoint(path='checkpoint.pth'):
#load the saved file
checkpoint = torch.load('checkpoint.pth')
#download pretrained model
model=models.vgg16(pretrained=True)
#to freeze parameters
for param in model.parameters():
param.requires_grad = False
#load from checkpoint
model.class_to_idx = checkpoint['class_to_idx']
model.load_state_dict(checkpoint['state_dict'])
return model
# -
# # Inference for classification
#
# Now you'll write a function to use a trained network for inference. That is, you'll pass an image into the network and predict the class of the flower in the image. Write a function called `predict` that takes an image and a model, then returns the top $K$ most likely classes along with the probabilities. It should look like
#
# ```python
# probs, classes = predict(image_path, model)
# print(probs)
# print(classes)
# > [ 0.01558163 0.01541934 0.01452626 0.01443549 0.01407339]
# > ['70', '3', '45', '62', '55']
# ```
#
# First you'll need to handle processing the input image such that it can be used in your network.
#
# ## Image Preprocessing
#
# You'll want to use `PIL` to load the image ([documentation](https://pillow.readthedocs.io/en/latest/reference/Image.html)). It's best to write a function that preprocesses the image so it can be used as input for the model. This function should process the images in the same manner used for training.
#
# First, resize the images where the shortest side is 256 pixels, keeping the aspect ratio. This can be done with the [`thumbnail`](http://pillow.readthedocs.io/en/3.1.x/reference/Image.html#PIL.Image.Image.thumbnail) or [`resize`](http://pillow.readthedocs.io/en/3.1.x/reference/Image.html#PIL.Image.Image.thumbnail) methods. Then you'll need to crop out the center 224x224 portion of the image.
#
# Color channels of images are typically encoded as integers 0-255, but the model expected floats 0-1. You'll need to convert the values. It's easiest with a Numpy array, which you can get from a PIL image like so `np_image = np.array(pil_image)`.
#
# As before, the network expects the images to be normalized in a specific way. For the means, it's `[0.485, 0.456, 0.406]` and for the standard deviations `[0.229, 0.224, 0.225]`. You'll want to subtract the means from each color channel, then divide by the standard deviation.
#
# And finally, PyTorch expects the color channel to be the first dimension but it's the third dimension in the PIL image and Numpy array. You can reorder dimensions using [`ndarray.transpose`](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.ndarray.transpose.html). The color channel needs to be first and retain the order of the other two dimensions.
# TODO: Process a PIL image for use in a PyTorch model
#Image.thumbnail(size, resample=3)
#https://pillow.readthedocs.io/en/3.1.x/reference/Image.html#PIL.Image.Image.thumbnail
# Crop the center of the image-->.crop((left, top, right, bottom))
#a.transpose().shape = (i[n-1], i[n-2], ... i[1], i[0])
#Refrence on processing the image-->https://stackoverflow.com/questions/55678459/resize-image-in-python
def process_image(image):
''' Scales, crops, and normalizes a PIL image for a PyTorch model,
returns an Numpy array
'''
img = PIL.Image.open(image)
# Get original dimensions
original_width, original_height = img.size
# Find shorter size and create settings to crop shortest side to 256
if original_width < original_height:
size=[256, 256**600]
else:
size=[256**600, 256]
img.thumbnail(size)
center = original_width/4, original_height/4
left, top, right, bottom = center[0]-(244/2), center[1]-(244/2), center[0]+(244/2), center[1]+(244/2)
img = img.crop((left, top, right, bottom))
numpy_img = np.array(img)/255
# Normalize each color channel
mean = [0.485, 0.456, 0.406]
std = [0.229, 0.224, 0.225]
numpy_img = (numpy_img-mean)/std
# Set the color to the first channel
numpy_img = numpy_img.transpose(2, 0, 1)
return numpy_img
# To check your work, the function below converts a PyTorch tensor and displays it in the notebook. If your `process_image` function works, running the output through this function should return the original image (except for the cropped out portions).
# +
def imshow(image, ax=None, title=None):
#img for tensor
if ax is None:
fig, ax = plt.subplots()
# PyTorch tensors assume the color channel is the first dimension
# but matplotlib assumes is the third dimension
image = image.transpose((1, 2, 0))
# Undo preprocessing
mean = np.array([0.485, 0.456, 0.406])
std = np.array([0.229, 0.224, 0.225])
image = std * image + mean
# Image needs to be clipped between 0 and 1 or it looks like noise when displayed
image = np.clip(image, 0, 1)
ax.imshow(image)
return ax
imshow(process_image("flowers/test/9/image_06413.jpg"))
imshow(process_image("flowers/test/37/image_03734.jpg"))
imshow(process_image("flowers/test/10/image_07090.jpg"))
# -
# ## Class Prediction
#
# Once you can get images in the correct format, it's time to write a function for making predictions with your model. A common practice is to predict the top 5 or so (usually called top-$K$) most probable classes. You'll want to calculate the class probabilities then find the $K$ largest values.
#
# To get the top $K$ largest values in a tensor use [`x.topk(k)`](http://pytorch.org/docs/master/torch.html#torch.topk). This method returns both the highest `k` probabilities and the indices of those probabilities corresponding to the classes. You need to convert from these indices to the actual class labels using `class_to_idx` which hopefully you added to the model or from an `ImageFolder` you used to load the data ([see here](#Save-the-checkpoint)). Make sure to invert the dictionary so you get a mapping from index to class as well.
#
# Again, this method should take a path to an image and a model checkpoint, then return the probabilities and classes.
#
# ```python
# probs, classes = predict(image_path, model)
# print(probs)
# print(classes)
# > [ 0.01558163 0.01541934 0.01452626 0.01443549 0.01407339]
# > ['70', '3', '45', '62', '55']
# ```
def predict(image_path, model, top_k=5):
''' Predict the class (or classes) of an image using a trained deep learning model.
image_path: string. Path to image, directly to image and not to folder.
model: pytorch neural network.
top_k: integer. The top K classes to be calculated
returns top_probabilities(k), top_labels
'''
# No need for GPU on this part (just causes problems)
model.to("cpu")
# Set model to evaluate
model.eval();
# Convert image from numpy to torch
torch_image = torch.from_numpy(np.expand_dims(process_image(image_path),
axis=0)).type(torch.FloatTensor).to("cpu")
# Find probabilities (results) by passing through the function (note the log softmax means that its on a log scale)
log_probs = model.forward(torch_image)
# Convert to linear scale
linear_probs = torch.exp(log_probs)
# Find the top 5 results
top_probs, top_labels = linear_probs.topk(top_k)
# Detatch all of the details
top_probs = np.array(top_probs.detach())[0] # This is not the correct way to do it but the correct way isnt working thanks to cpu/gpu issues so I don't care.
top_labels = np.array(top_labels.detach())[0]
# Convert to classes
idx_to_class = {val: key for key, val in
model.class_to_idx.items()}
top_labels = [idx_to_class[lab] for lab in top_labels]
top_flowers = [cat_to_name[lab] for lab in top_labels]
return top_probs, top_labels, top_flowers
# ## Sanity Checking
#
# Now that you can use a trained model for predictions, check to make sure it makes sense. Even if the testing accuracy is high, it's always good to check that there aren't obvious bugs. Use `matplotlib` to plot the probabilities for the top 5 classes as a bar graph, along with the input image. It should look like this:
#
# <img src='assets/inference_example.png' width=300px>
#
# You can convert from the class integer encoding to actual flower names with the `cat_to_name.json` file (should have been loaded earlier in the notebook). To show a PyTorch tensor as an image, use the `imshow` function defined above.
# +
# TODO: Display an image along with the top 5 classes
# Define image path
image_path = "flowers/test/37/image_03734.jpg"
# Set up plot
plt.figure(figsize = (6,10))
ax = plt.subplot(2,1,1)
# Set up title
flower_num = image_path.split('/')[2]
title_ = cat_to_name[flower_num]
# Plot flower
img = process_image(image_path)
imshow(img, ax, title = title_);
# Make prediction
probs, labs, flowers = predict(image_path, model)
# Plot bar chart
plt.subplot(2,1,2)
sns.barplot(x=probs, y=flowers, color=sns.color_palette()[0]);
plt.show()
# -
image_path = "flowers/test/9/image_06413.jpg"
plt.figure(figsize = (6,10))
ax = plt.subplot(2,1,1)
flower_num = image_path.split('/')[2]
title_ = cat_to_name[flower_num]
img = process_image(image_path)
imshow(img, ax, title = title_);
probs, labs, flowers = predict(image_path, model)
plt.subplot(2,1,2)
sns.barplot(x=probs, y=flowers, color=sns.color_palette()[0]);
plt.show()
| Image Classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.7 64-bit (''tf2'': conda)'
# metadata:
# interpreter:
# hash: 6217ec1083430fbb69f0f529f559f9d6a9398dfc34178f068ec9884c7c99515e
# name: 'Python 3.7.7 64-bit (''tf2'': conda)'
# ---
# ## TensorFlow Keras Layers
#
# 2020.9.12
#
# ### Reference
#
# https://www.tensorflow.org/api_docs/python/tf/keras/layers
import numpy as np
import tensorflow as tf
from tensorflow import keras
from tqdm import tqdm
import matplotlib.pyplot as plt
# ## Import the MNIST dataset
mnist = keras.datasets.mnist
(train_images, train_labels), (test_images, test_labels) = mnist.load_data()
train_images = np.expand_dims(train_images, -1)
# + tags=[]
print(f"train_images.shape:{train_images.shape}")
print(f"train_labels.shape:{train_labels.shape}")
# -
# ## tf.keras.Conv{x}D layers
#
# 卷积系列层
# + tags=[]
input_shape = train_images.shape[1:] # (28, 28, 1)
# make single sample x to form a batch, by expanding dimention
x = np.expand_dims(train_images[0], 0).astype('float32') # uint8 to float32
print(x.shape)
print(x.dtype)
m = tf.keras.layers.Conv2D(1, kernel_size=1, input_shape=input_shape)
y = m(x)
print(y.shape)
print(y.dtype)
# + tags=[]
print(np.max(x))
print(np.min(x))
print(np.max(y))
print(np.min(y))
# -
m.kernel
x0 = x[0,:,:,0]
x0.shape
# print(x0)
# + tags=[]
print(x.shape)
plt.figure()
plt.imshow(x[0,:,:,0], vmin=0, vmax=255, cmap="gray")
plt.title("origin image")
plt.figure()
plt.imshow(y[0,:,:,0], vmin=0, vmax=255, cmap="gray")
plt.title("image after 1x1 kernel convolution")
# -
# ## tf.keras.Conv2Dx layers
#
# Pooling 系列层
# ### tf.keras.layers.AveragePooling2D
input_shape = (2, 4, 5, 3) # (2, 4, 5, 3)
x = tf.random.normal(input_shape)
y = tf.keras.layers.AveragePooling2D(pool_size=(3, 3))(x)
y.shape
# ### tf.keras.layers.GlobalAveragePooling2D
m = tf.keras.layers.GlobalAveragePooling2D()
y = m(x)
np.average(x)
print(x[0].shape)
print(y)
# ## tf.pad
# t = tf.constant([[1, 2, 3], [4, 5, 6]])
t = np.asarray([[1,2,3],[4,5,6]])
paddings = tf.constant([[1, 1],[1, 1]])
# 'constant_values' is 0.
# rank of 't' is 2.
tf.pad(t, paddings, "CONSTANT")
t
| keras_basics/tf.keras.layers_test.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: julia-nteract-1.5
# kernelspec:
# argv:
# - C:\Users\snikula\AppData\Local\JuliaPro-1.5.4-1\Julia-1.5.4\bin\julia.exe
# - -i
# - --color=yes
# - C:\Users\snikula\.julia\packages\IJulia\e8kqU\src\kernel.jl
# - '{connection_file}'
# display_name: Julia nteract 1.5.4
# env: {}
# interrupt_mode: message
# language: julia
# name: julia-nteract-1.5
# ---
# + [markdown] nteract={"transient": {"deleting": false}}
# # Ionisoiva säteily
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
using Printf
using Dates
b=2.897768e-3 # Wienin siirtymälain vakio
c=2.99792e8 # Valon nopeus
h=6.6260693e-34 # Plancin vakio
eV=1.6021766e-19 # Elektorinivoltti
sigma=5.670374419e-8 # Stefan-Boltzman
@printf "%s\n" Dates.now();
versioninfo(verbose=false);
# + [markdown] nteract={"transient": {"deleting": false}}
# 13-6
#
# Kirjan vastaus: 300 h
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
# t*5µSv/h=0.5*3.2 mSv
t=0.5*3.2e-3/5e-6
# + [markdown] nteract={"transient": {"deleting": false}}
# 13-9
#
# Kirjan vastaus: 76 %
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
# I=I0*exp(-µx)
µ=46#1/m
t=6e-3#m
exp(-µ*t)
# + [markdown] nteract={"transient": {"deleting": false}}
# 13-10
#
# Kirjan vastaus: a) 3.0 1/m b) 0.24 %
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
k=0.5
t=0.23
µ=log(1/k)/t
t=2
@printf("Heikennyskerroin on %.1f 1/m.\n",µ)
@printf("%.2f %% säteilystä läpäisee 2 metrin vesikerroksen.\n",
exp(-µ*t)*100)
# + [markdown] nteract={"transient": {"deleting": false}}
# 13-11
#
# Kirjan vastaus: 1) noin 1/3
#
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
k=0.5
t=0.05
µ=log(1/k)/t
t=0.08
@printf("%.2f %% säteilystä läpäisee 8 cm pehmytkudoskerroksen.\n",
exp(-µ*t)*100)
# + [markdown] nteract={"transient": {"deleting": false}}
# 13-12
#
# Kirjan vastaus: 4.0 cm
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
k=22738/38791
t=0.03
µ=log(1/k)/t
k=0.5
t=log(1/k)/µ
# + [markdown] nteract={"transient": {"deleting": false}}
# 13-13
#
# Kirjan vastaus: 110 1/m
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
k=210/1205
t=0.016
µ=log(1/k)/t
# + [markdown] nteract={"transient": {"deleting": false}}
# 13-14
#
# Kirjan vastaus:<br>
# a) $\substack{222 \\ 86}Rn \rightarrow \substack{218 \\ 84}Po + \substack{4 \\ 2}He$<br><br>
# c) kyllä
| 7/13.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import cv2
import numpy as np
import os
face_cascade = cv2.CascadeClassifier(cv2.data.haarcascades + 'haarcascade_frontalface_default.xml')
eye_cascade = cv2.CascadeClassifier(cv2.data.haarcascades+'haarcascade_eye.xml')
# +
directory = 'C:\\Users\\<NAME>\\Desktop\\ML\\OpenCV\\Face_recognition\\FaceData\\'
filename = input('Enter your name')
cap = cv2.VideoCapture(0)
skip=0
face_data=[]
while True:
ret,img=cap.read()
if ret==False:
continue
faces = face_cascade.detectMultiScale(img2,1.3,5)
#To get the biggest face on the screen
sorted(faces,key=lambda f:f[2]*f[3])
#for face in faces[-1:]:
if len(faces)==0:
continue
x,y,w,h=faces[-1]
cv2.rectangle(img,(x,y),(x+w,y+h),(0,255,0),thickness=2)
#cropping the face
offset=10
face_section=img[y-offset:y+h+offset,x-offset:x+w+offset]
face_section=cv2.resize(face_section,(100,100))
skip+=1
if skip%10==0:
face_data.append(face_section)
print(len(face_data))
cv2.imshow('faces',img)
key_pressed = cv2.waitKey(1) &0xFF
if key_pressed==ord('q'):
break
cap.release()
cv2.destroyAllWindows()
#converting into array
face_data=np.asarray(face_data)
face_data=face_data.reshape((face_data.shape[0],-1))
np.save(directory+filename+'.npy',face_data)
# +
face_data=[]
face_label=[]
face_dic={}
label=0
for fx in os.listdir('C:\\Users\\<NAME>\\Desktop\\ML\\OpenCV\\Face_recognition\\FaceData'):
if fx.endswith('.npy'):
df1 = np.load('C:\\Users\\<NAME>\\Desktop\\ML\\OpenCV\\Face_recognition\\FaceData\\'+fx)
df1.reshape(-1,df1.shape[0])
face_data.append(df1)
df2 = label*np.ones(df1.shape[0])
face_dic[label]=fx[0:len(fx)-4]
label+=1
face_label.append(df2)
face_data=np.concatenate(face_data,axis=0)
face_label=np.concatenate(face_label,axis=0).reshape((-1,1))
trainset = np.concatenate((face_data,face_label),axis=1)
# +
###distance
def distance(x1,x2):
return np.sqrt(sum((x1-x2)**2))
###KNN
def knn(trainset,test,k=5):
vals=[]
X = trainset[:,0:-1]
Y= trainset[:,-1]
for i in range(len(X)):
d1=distance(test,X[i])
vals.append((d1,Y[i]))
vals=sorted(vals)
vals=np.array(vals)
vals=vals[0:k]
new_vals = np.unique(vals[:,1],return_counts=True)
index = new_vals[1].argmax()
pred = new_vals[0][index]
return pred
# +
####test data
cap = cv2.VideoCapture(0)
while True:
ret,img = cap.read()
if ret==False:
continue
faces = face_cascade.detectMultiScale(img,1.3,5)
sorted(faces,key=lambda f:f[2]*f[3])
#for face in faces[-1:]:
if len(faces)==0:
continue
x,y,w,h=faces[-1]
#cropping the face
offset=10
face_section=img[y-offset:y+h+offset,x-offset:x+w+offset]
face_section=cv2.resize(face_section,(100,100))
ans=int(knn(trainset,face_section.flatten()))
ans_name = face_dic[ans]
cv2.putText(img,ans_name,(x-10,y-10),cv2.FONT_HERSHEY_SIMPLEX,1,(255,0,255),2,cv2.LINE_AA)
cv2.rectangle(img,(x,y),(x+w,y+h),(0,255,0),thickness=2)
cv2.imshow('faces',img)
key_pressed = cv2.waitKey(1) &0xFF
if key_pressed==ord('q'):
break
cap.release()
cv2.destroyAllWindows()
# -
#
# +
# -
import os
os.listdir('C:\\Users\\<NAME>\\Desktop\\ML\\OpenCV\\Face_recognition\\FaceData')
# +
# -
img.shape
df1[300]
df2[30]
| Face-Detection/Face-Detection-using-KNN/Face_Recognition.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="95UzhBOzHUjx"
# # Code Objective:
#
# * Ensemble from BERT and XLMRoberta Prediction Probabilities to Enhance Performance of Coarse Grained Evaluation
#
# # Code Result:
# * (Previous) Accuracy - BERT Model for Coarse Grained Evaluation = 91.63 %
# * (Previous) Accuracy - XLMRoBerta for Coarse Grained Evaluation = 89.76 %
# * Accuracy - Ensemble (BERT+Roberta) for Coarse Grained Evaluation = 92.60 %
#
#
#
#
# + [markdown] id="hsXFEqjPINxA"
# # Importing Libraries
# + id="vlW5QKbi2_IN"
import os
import itertools
import numpy as np
import pandas as pd
import tensorflow as tf
from google.colab import drive
import matplotlib.pyplot as plt
from sklearn.metrics import confusion_matrix
from sklearn.metrics import classification_report
from sklearn.model_selection import train_test_split
from tensorflow.keras.models import Model, Sequential
from tensorflow.keras.layers import Dropout, Dense, Add, Multiply, Average, Concatenate, Input, Subtract
# + [markdown] id="jqHmJSIwIRQy"
# # Data Loading
# + id="NmvhuUbo5_cv"
path = '/content/'
# Probability Data Loading
x_train_1 = np.load(path + 'Train_Probs_Bert.npy', allow_pickle = True)
x_train_1 = x_train_1[:,:2]
x_test_1 = np.load(path + 'Test_Probs_Bert.npy', allow_pickle = True)
x_test_1 = x_test_1[:,:2]
x_train_2 = np.load(path + 'Train_Probs_XLMR.npy', allow_pickle = True)
x_test_2 = np.load(path + 'Test_Probs_XLMR.npy', allow_pickle = True)
y_train = np.load(path + 'Train_Labels.npy', allow_pickle = True)
y_test = np.load(path + 'Test_Labels.npy', allow_pickle = True)
print("X Train Prob BERT Size = {}".format(x_train_1.shape))
print("X Test Prob BERT Size = {}".format(x_test_1.shape))
print("X Train Prob XLMR Size = {}".format(x_train_2.shape))
print("X Train Prob XLMR Size = {}".format(x_test_2.shape))
print("Y Train Label Size = {}".format(y_train.shape))
print("Y Test Label Size = {}".format(y_test.shape))
# + [markdown] id="BhX_VueOIaBw"
# # Ensemble Architecture (BERT and XLMRoberta)
# + id="jCJN_30h54zt"
# Input Placeholders
input_1 = Input(shape = (2,))
input_2 = Input(shape = (2,))
# Ensemble Model Archirecture
def Classifier_Top(input_1,input_2):
activation = 'tanh'
z1 = Dense(units = 50, activation = activation)(input_1)
z1 = Dropout(0.2)(z1)
z2 = Dense(units = 50, activation = activation)(input_2)
z2 = Dropout(0.2)(z2)
z = Concatenate()([z1,z2])
z = Dense(units = 30, activation = activation)(z)
z = Dropout(0.2)(z)
z = Dense(units = 20, activation = activation)(z)
z = Dropout(0.2)(z)
z = Dense(units = 10, activation = activation)(z)
z = Dropout(0.2)(z)
z = Dense(units = 5, activation = activation)(z)
z = Dropout(0.2)(z)
output = Dense(units = 1, activation = 'sigmoid')(z)
model = Model(inputs = [input_1,input_2], outputs = output)
model.summary()
return model
# Compile and Train Model
def compile_and_train(model, num_epochs):
model.compile(optimizer= 'adam', loss= 'binary_crossentropy', metrics=['acc'])
history = model.fit([x_train_1,x_train_2], y_train, batch_size=32, epochs=num_epochs, validation_split=0.2)
return history
# + [markdown] id="HOvD3MJ_I0lw"
# # Training and Evaluation
# + id="TRMty4OQ8FZM"
Epochs = 10 # Training Iteration Number
Classifier = Classifier_Top(input_1,input_2)
history = compile_and_train(Classifier,Epochs)
# + id="2tBNo_rdCU9U"
# For Loading Already Finetuned Model
Classifier = Classifier_Top(input_1,input_2)
Classifier.compile(optimizer= 'adam', loss= 'binary_crossentropy', metrics=['acc'])
Classifier.load_weights('content/Task_1_Best.h5') # Load the Weight into colab local directory first
# + [markdown] id="mSpLGt7zJSzv"
# # Evaluation
# + id="KUZO_TTd-ybs"
test_scores = Classifier.evaluate([x_test_1,x_test_2], y_test, verbose=2)
print("Test loss:", test_scores[0])
print("Test accuracy:", test_scores[1])
# + [markdown] id="5aty4RWmJatg"
# # Model Save
# + id="2o3oOP_mz4yy"
Classifier.save_weights("Task_1_Best.h5")
# + [markdown] id="wIR3Dpm9JmYA"
# # Visualization of Test Result (Coarse Grained Evaluation)
# + id="QKtzfRfD0buz"
# Code for Changing
def pred_to_lab(y_pred_probs, mode):
labels = []
for i in y_pred_probs:
if i<0.5:
if mode == 'Num':
labels.append(0)
elif mode == 'Text':
labels.append('non-hostile')
else:
if mode == 'Num':
labels.append(1)
elif mode == 'Text':
labels.append('hostile')
if mode == 'Num':
return np.array(labels)
elif mode == 'Text':
return labels
y_pred_probs = Classifier.predict([x_test_1,x_test_2])
y_pred = pred_to_lab(y_pred_probs, mode = 'Num')
print(y_pred)
# + colab={"base_uri": "https://localhost:8080/", "height": 620} id="sZzKemQJ0xYA" outputId="3ba6fcff-f394-4c2f-a4f6-584112c2712d"
def plot_confusion_matrix(cm, target_names, title='Confusion matrix', cmap=None, normalize=False):
"""
given a sklearn confusion matrix (cm), make a nice plot
Arguments
---------
cm: confusion matrix from sklearn.metrics.confusion_matrix
target_names: given classification classes such as [0, 1, 2]
the class names, for example: ['high', 'medium', 'low']
title: the text to display at the top of the matrix
cmap: the gradient of the values displayed from matplotlib.pyplot.cm
see http://matplotlib.org/examples/color/colormaps_reference.html
plt.get_cmap('jet') or plt.cm.Blues
normalize: If False, plot the raw numbers
If True, plot the proportions
"""
accuracy = np.trace(cm) / np.sum(cm).astype('float')
misclass = 1 - accuracy
if cmap is None:
cmap = plt.get_cmap('Blues')
plt.figure(figsize=(8, 6))
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
if target_names is not None:
tick_marks = np.arange(len(target_names))
plt.xticks(tick_marks, target_names, rotation=45)
plt.yticks(tick_marks, target_names)
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
thresh = cm.max() / 1.5 if normalize else cm.max() / 2
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
if normalize:
plt.text(j, i, "{:0.4f}".format(cm[i, j]),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
else:
plt.text(j, i, "{:,}".format(cm[i, j]),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label\naccuracy={:0.4f}; misclass={:0.4f}'.format(accuracy, misclass))
plt.show()
print(classification_report(y_test,y_pred))
target_names = ['Non Hostile','Hostile']
cm = confusion_matrix(y_test, y_pred)
plot_confusion_matrix(cm, target_names)
# + id="zN5jxsNR1Wpg"
# Saving Predicted Labels of Coarse Grained Evaluation
np.save("Task_1_Pred_Labels.npy",y_pred)
| Fine-Tuning/Coarse_Grained_Ensemble_Finetune.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # NumPy is faster than list:
#
# - Fixed Type
# - Faster to read less bytes of memory
# - No type checking
# - Contiguous Memory
# - SIMD Vector Processing
# - Effective Cache Utilization
| FreeCodeCamp/Data Analysis with Python/Numpy/1 - What is NumPy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 学习内容
#
# 1. 相关概念(无监督学习、聚类的定义)
# 2. 性能度量(外部指标、内部指标)
# 3. 距离计算
# 4. 原型聚类
# K均值
# LVQ
# 高斯混合聚类
# 5. 层次聚类
# AGNES
# 自顶而下
# 6. 密度聚类
# DBSCAN
# 其他密度聚类算法
# 7. 优缺点
# 8. sklearn参数详解
# ### 1.相关概念
# * 无监督学习:
# 无监督学习是机器学习的一种方法,没有给定事先标记过的训练示例,自动对输入的数据进行分类或分群。无监督学习的主要运用包含:聚类分析、关系规则、维度缩减。它是监督式学习和强化学习等策略之外的一种选择。 一个常见的无监督学习是数据聚类。在人工神经网络中,生成对抗网络、自组织映射和适应性共振理论则是最常用的非监督式学习。
# * 聚类:
# 聚类是一种无监督学习。聚类是把相似的对象通过静态分类的方法分成不同的组别或者更多的子集,这样让在同一个子集中的成员对象都有相似的一些属性,常见的包括在坐标系中更加短的空间距离等。
# +
# 通过简单的例子来直接查看K均值聚类的效果
from sklearn.cluster import KMeans
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
# 聚类前
X = np.random.rand(100, 2)
plt.scatter(X[:, 0], X[:, 1], marker='o')
# +
#聚类后
kmeans = KMeans(n_clusters=2).fit(X)
label_pred = kmeans.labels_
plt.scatter(X[:, 0], X[:, 1], c=label_pred)
plt.show()
# -
#聚类后
kmeans = KMeans(n_clusters=2).fit(X)
label_pred = kmeans.labels_
plt.scatter(X[:, 0], X[:, 1], c=label_pred)
plt.show()
# ### 2.性能度量
# 在机器学习中我们都需要对任务进行评价以便于进行下一步的优化,聚类的性能度量主要有一下两种。
# * 外部指标:是指把算法得到的划分结果跟某个外部的“参考模型”(如专家给出的划分结果)比较
# * 内部指标:是指直接考察聚类结果,不利用任何参考模型的指标。
# ### 3.距离计算
# 在机器学习和数据挖掘中,我们经常需要知道个体间差异的大小,进而评价个体的相似性和类别。
# * 欧式距离(2-norm距离)
# * 曼哈顿距离(Manhattan distance, 1-norm距离)
# * 切比雪夫距离
# * 闵可夫斯基距离
# * 余弦相似性
# * 马氏距离
# 欧式距离:欧氏距离是最易于理解的一种距离计算方法,源自欧氏空间中两点间的距离公式。
# $$d(x,y)=\sqrt{\Sigma_{k=1}^n (x_k-y_k)^2}$$
# 曼哈顿距离:
# 曼哈顿距离也称为街区距离,计算公式如下:
# $$d(x,y)=\Sigma_{k=1}^n \left|x_k-y_k\right|$$
# 切比雪夫距离:$$d(x,y) = \lim_{n\rightarrow \infty} (\Sigma_{k=1}^n (\left|x_k-y_k\right|)^r)^\dfrac{1}{r} = max_k (\left|x_k-y_k\right|)$$
# 闵可夫斯基距离:
# $$d(x,y)=(\Sigma_{k=1}^n (\left|x_k-y_k\right|)^r)^\dfrac{1}{r}$$
# 式中,r是一个可变参数,根据参数r取值的不同,闵可夫斯基距离可以表示一类距离
# r = 1时,为曼哈顿距离
# r = 2时,为欧式距离
# r →∞时,为切比雪夫距离
# 闵可夫斯基距离包括欧式距离、曼哈顿距离、切比雪夫距离都假设数据各维属性的量纲和分布(期望、方差)相同,因此适用于度量独立同分布的数据对象。
# 余弦距离:
# 余弦相似度公式定义如下:
# $$cos(x,y)=\dfrac{xy}{\left|x\right|\left|y\right|} = \dfrac{\Sigma_{k=1}^n x_k y_k}{\sqrt{\Sigma_{k=1}^n x_k^2} \sqrt{\Sigma_{k=1}^n y_k^2}}$$
#
# 余弦相似度实际上是向量xx和yy夹角的余弦度量,可用来衡量两个向量方向的差异。如果余弦相似度为11,则xx和yy之间夹角为0°0°,两向量除模外可认为是相同的;如果预先相似度为00,则xx和yy之间夹角为90°90°,则认为两向量完全不同。在计算余弦距离时,将向量均规范化成具有长度11,因此不用考虑两个数据对象的量值。
# 余弦相似度常用来度量文本之间的相似性。文档可以用向量表示,向量的每个属性代表一个特定的词或术语在文档中出现的频率,尽管文档具有大量的属性,但每个文档向量都是稀疏的,具有相对较少的非零属性值。
# 马氏距离:
# $$mahalanobis(x,y)=(x-y)\Sigma^{-1}(x-y)^T$$
# 式中,Σ−1Σ−1是数据协方差矩阵的逆。
# 前面的距离度量方法大都假设样本独立同分布、数据属性之间不相关。马氏距离考虑了数据属性之间的相关性,排除了属性间相关性的干扰,而且与量纲无关。若协方差矩阵是对角阵,则马氏距离变成了标准欧式距离;若协方差矩阵是单位矩阵,各个样本向量之间独立同分布,则变成欧式距离。
# ### 4.原型聚类
# 原型聚类亦称"基于原型的聚类" (prototype-based clustering),此类算法假设聚类结构能通过一组原型刻画,在现实聚类任务中极为常用.通常情形下,算法先对原型进行初始化,然后对原型进行迭代更新求解.采用不同的原型表示、不同的求解方式,将产生不同的算法.
# * K均值
# * LVQ
# * 高斯混合聚类
# k均值聚类算法(k-means clustering algorithm)是一种迭代求解的聚类分析算法,其步骤是
# 创建 k 个点作为起始质心(通常是随机选择)
# 当任意一个点的簇分配结果发生改变时(不改变时算法结束)
# 对数据集中的每个数据点
# 对每个质心
# 计算质心与数据点之间的距离
# 将数据点分配到距其最近的簇
# 对每一个簇, 计算簇中所有点的均值并将均值作为质心
# 聚类中心以及分配给它们的对象就代表一个聚类。
# +
def distEclud(vecA, vecB):
'''
欧氏距离计算函数
:param vecA:
:param vecB:
:return:
'''
return np.sqrt(np.sum(np.power(vecA - vecB, 2)))
def randCent(dataMat, k):
'''
为给定数据集构建一个包含K个随机质心的集合,
随机质心必须要在整个数据集的边界之内,这可以通过找到数据集每一维的最小和最大值来完成
然后生成0到1.0之间的随机数并通过取值范围和最小值,以便确保随机点在数据的边界之内
:param dataMat:
:param k:
:return:
'''
# 获取样本数与特征值
m, n = np.shape(dataMat)
# 初始化质心,创建(k,n)个以零填充的矩阵
centroids = np.mat(np.zeros((k, n)))
print(centroids)
# 循环遍历特征值
for j in range(n):
# 计算每一列的最小值
minJ = np.min(dataMat[:, j])
# 计算每一列的范围值
rangeJ = float(np.max(dataMat[:, j]) - minJ)
# 计算每一列的质心,并将值赋给centroids
centroids[:, j] = np.mat(minJ + rangeJ * np.random.rand(k, 1))
# 返回质心
return centroids.A
def kMeans(dataMat, k, distMeas=distEclud):
'''
创建K个质心,然后将每个店分配到最近的质心,再重新计算质心。
这个过程重复数次,直到数据点的簇分配结果不再改变为止
:param dataMat: 数据集
:param k: 簇的数目
:param distMeans: 计算距离
:return:
'''
# 获取样本数和特征数
m, n = np.shape(dataMat)
# 初始化一个矩阵来存储每个点的簇分配结果
# clusterAssment包含两个列:一列记录簇索引值,第二列存储误差(误差是指当前点到簇质心的距离,后面会使用该误差来评价聚类的效果)
clusterAssment = np.mat(np.zeros((m, 2)))
# 创建质心,随机K个质心
centroids = randCent(dataMat, k)
# 初始化标志变量,用于判断迭代是否继续,如果True,则继续迭代
clusterChanged = True
while clusterChanged:
clusterChanged = False
# 遍历所有数据找到距离每个点最近的质心,
# 可以通过对每个点遍历所有质心并计算点到每个质心的距离来完成
for i in range(m):
minDist = float("inf")
minIndex = -1
for j in range(k):
# 计算数据点到质心的距离
# 计算距离是使用distMeas参数给出的距离公式,默认距离函数是distEclud
distJI = distMeas(centroids[j, :], dataMat[i, :])
# 如果距离比minDist(最小距离)还小,更新minDist(最小距离)和最小质心的index(索引)
if distJI < minDist:
minDist = distJI
minIndex = j
# 如果任一点的簇分配结果发生改变,则更新clusterChanged标志
if clusterAssment[i, 0] != minIndex: clusterChanged = True
# 更新簇分配结果为最小质心的index(索引),minDist(最小距离)的平方
clusterAssment[i, :] = minIndex, minDist ** 2
# print(centroids)
# 遍历所有质心并更新它们的取值
for cent in range(k):
# 通过数据过滤来获得给定簇的所有点
ptsInClust = dataMat[np.nonzero(clusterAssment[:, 0].A == cent)[0]]
# 计算所有点的均值,axis=0表示沿矩阵的列方向进行均值计算
centroids[cent, :] = np.mean(ptsInClust, axis=0)
# 返回所有的类质心与点分配结果
return centroids, clusterAssment
# +
# 运行Kmeans,假设有两聚类中心
center,label_pred = kMeans(X, k=2)
# 将标签转化成易绘图的形式
label = label_pred[:, 0].A.reshape(-1)
# 将结果可视化
plt.scatter(X[:, 0], X[:, 1], c=label)
plt.scatter(center[0, 0], center[0, 1], marker="*", s = 100)
plt.scatter(center[1, 0], center[1, 1], marker="*", s = 100)
plt.show()
# -
# 学习向量量化(Learning Vector Quantization,简称LVQ)属于原型聚类,即试图找到一组原型向量来聚类,每个原型向量代表一个簇,将空间划分为若干个簇,从而对于任意的样本,可以将它划入到它距离最近的簇中,不同的是LVQ假设数据样本带有类别标记,因此可以利用这些类别标记来辅助聚类。
def LVQ(X, y, k = 2, max_time=10000, ita=0.01):
# 初始化一组原型向量
init_index = np.random.choice(len(y), k)
px = X[init_index]
py = y[init_index]
print(px,py)
for n in range(max_time):
# 从样本集随机选择样本
j = np.random.choice(len(y), 1)
xj, yj = X[j], y[j]
# 计算并找出与样本最接近的原型向量
i = np.argmin([np.linalg.norm(xj - pi) for pi in px])
pyi = py[i]
# 更新原型向量
if pyi == yj:
px[i] = px[i] + ita * (xj - px[i])
else:
px[i] = px[i] - ita * (xj - px[i])
if n % 1000 == 0:
plt.scatter(px[:, 0], px[:, 1], marker=".", s = 100)
return px
plt.scatter(X[:, 0], X[:, 1])
P = LVQ(X, np.random.randint(0, 2, len(X)))
plt.scatter(P[:, 0], P[:, 1], marker="*", s = 100)
plt.show()
# 高斯混合聚类:高斯混合聚类与k均值、LVQ用原型向量来刻画聚类结构不同,高斯混合聚类采用概率模型来表达聚类原型。相对于k均值聚类算法使用 k 个原型向量来表达聚类结构,高斯混合聚类使用 k 个高斯概率密度函数混合来表达聚类结构
#
# 于是迭代更新 k 个簇原型向量的工作转换为了迭代更新 k 个高斯概率密度函数的任务。每个高斯概率密度函数代表一个簇,当一个新的样本进来时,我们可以通过这 k 的函数的值来为新样本分类
# ### 5.层次聚类
# 层次聚类(hierarchical clustering)基于簇间的相似度在不同层次上分析数据,从而形成树形的聚类结构,层次聚类一般有两种划分策略:自底向上的聚合(agglomerative)策略和自顶向下的分拆(divisive)策略
# * AGNES
# * 自顶而下
# AGNES算法是自底向上的层次聚类算法。开始时将数据集中的每个样本初始化为一个簇,然后找到距离最近的两个簇,将他们合并,不断重复这个过程,直达到到预设的聚类数目为止。
#
# 簇间距离的计算可以有三种形式:
# 最小距离:$d_{min}(C_i,C_j)=\min_{p\in C_i,q\in C_j}|p-q|.$
# 最大距离:$d_{max}(C_i,C_j)=\max_{p\in C_i,q\in C_j}|p-q|.$
# 平均距离:$d_{avg}(C_i,C_j)=\frac{1}{|C_i||C_j|}\sum_{p\in C_i}\sum_{q\in C_j}|p-q|.$
# ```
# 输入:样本集D={x1,x2,...,xm}D={x1,x2,...,xm}
# 聚类簇距离度量函数dd;
# 聚类簇数kk
# 过程:
# 1. for j=1,2,...,mj=1,2,...,m do
# 2. Cj={xj}Cj={xj}
# 3. end for
# 4. for i=1,2,...,mi=1,2,...,m do
# 5. for i=1,2,...,mi=1,2,...,m do
# 6. M(i,j)=d(Ci,Cj)M(i,j)=d(Ci,Cj);
# 7. M(j,i)=M(i,j)M(j,i)=M(i,j);
# 8. end for
# 9. end for
# 10. 设置当前聚类簇个数:q=mq=m;
# 11. while q>kq>k do
# 12. 找出距离最近的两个聚类簇Ci∗Ci∗和Cj∗Cj∗;
# 13. 合并Ci∗Ci∗和Cj∗Cj∗:Ci∗=Ci∗⋃Cj∗Ci∗=Ci∗⋃Cj∗;
# 14. for j=j∗+1,j∗+2,..,qj=j∗+1,j∗+2,..,q do
# 15. 将聚类簇CjCj重新编号为CjCj
# 16. end for
# 17. 删除距离矩阵MM的第j∗j∗行和第j∗j∗列;
# 18. for j=1,2,...,q−1j=1,2,...,q−1 do
# 19. M(i,j)=d(Ci,Cj)M(i,j)=d(Ci,Cj);
# 20. M(j,i)=M(i,j)M(j,i)=M(i,j);
# 21. end for
# 22. q=q−1q=q−1
# 23. end while
# 输出:簇划分:C={C1,C2,...,Ck}
# ```
#dist_min
def dist_min(Ci, Cj):
return np.min(distEclud(i, j) for i in Ci for j in Cj)
#dist_max
def dist_max(Ci, Cj):
return np.max(dist(i, j) for i in Ci for j in Cj)
#dist_avg
def dist_avg(Ci, Cj):
return sum(dist(i, j) for i in Ci for j in Cj)/(len(Ci)*len(Cj))
#找到距离最小的下标
def find_Min(M):
min_i = 1000
x = 0; y = 0
for i in range(len(M)):
for j in range(len(M[i])):
print(M[i][j], min_i)
if (i != j) and (
M[i][j] < min_i):
min_i = M[i][j];x = i; y = j
return (x, y, min_i)
def AGNES(dataset, dist, k):
#初始化C和M
C = [];M = []
# 初始化单样本聚类簇
for i in dataset:
Ci = []
Ci.append(i)
C.append(Ci)
# 初始化聚类簇距离矩阵
print(C)
for i in C:
Mi = []
for j in C:
Mi.append(dist(i, j))
print("Mi:", Mi)
M.append(Mi)
# 设置当前聚类簇个数
print(M)
q = len(dataset)
#合并更新
while q > k:
# 找出距离最近的两个聚类簇
x, y, min_i = find_Min(M)
# print(x, y, min)
# 合并两个聚类簇
#print(C[x])
C[x].extend(C[y])
#print(C[x])
#print("C:",C)
# 移除被合并的聚类簇
#print("1",np.delete(C, C[y]))
C = np.delete(C, C[y])
#print("c",C)
M = []
for i in C:
Mi = []
for j in C:
Mi.append(dist(i, j))
M.append(Mi)
q -= 1
return C
P = AGNES(dataset=X[:10], dist = dist_min, k=2)
P
# 自顶而下:把整个数据集视作一个簇,然后把一个簇分成几个簇,接着再分别把每一个簇分成更小的簇,如此反复下去,直到满足要求为止。
# ### 6.密度聚类
# 密度聚类假设聚类结构通过样本分布的紧密程度。此算法是基于密度的角度来考察样本之间的连接性,并基于连接性不断扩展聚类簇最后获得最终的结果。通过判断样本在区域空间内是否大于某个阈值来决定是否将其放到与之相近的样本中。
# * DBSCAN
# * 其他密度聚类算法
# DBSCAN
#
# e-邻域:对xj∈D,其∈邻域包含样本集D中与xj的距离不大于e的样本,即N(xj)= {xi∈D | dist(xi,xj)≤e};
# 核心对象(core object): 若xj的E-邻域至少包含MinPts个样本,即|Ne(xj)|≥MinPts,则xj是-一个核心对象;
# 密度直达(directly density- reachable):若xj位于xi的e-邻域中,且xi是核心对象,则称x;由xi密度直达;
# 密度可达(density. reachable): 对xi与xj,若存在样本序列P1,P2,... ,Pn,其中p1=xi,Pn=xj且pi+1由pi密度直达,则称xj由xi密度可达;
# 密度相连(density-conected): 对xi与xj,若存在xk使得xi与xj均由xk密度可达,则称xi与xj密度相连.
# ```
# 首先将数据集D中的所有对象标记为未处理状态
# for(数据集D中每个对象p) do
# if (p已经归入某个簇或标记为噪声) then
# continue;
# else
# 检查对象p的Eps邻域 NEps(p) ;
# if (NEps(p)包含的对象数小于MinPts) then
# 标记对象p为边界点或噪声点;
# else
# 标记对象p为核心点,并建立新簇C, 并将p邻域内所有点加入C
# for (NEps(p)中所有尚未被处理的对象q) do
# 检查其Eps邻域NEps(q),若NEps(q)包含至少MinPts个对象,则将NEps(q)中未归入任何一个簇的对象加入C;
# end for
# end if
# end if
# end for
# ```
# 优点
# ```
# 相比 K-平均算法,DBSCAN 不需要预先声明聚类数量。
# DBSCAN 可以找出任何形状的聚类,甚至能找出一个聚类,它包围但不连接另一个聚类,另外,由于 MinPts 参数,single-link effect (不同聚类以一点或极幼的线相连而被当成一个聚类)能有效地被避免。
# DBSCAN 能分辨噪音(局外点)。
# DBSCAN 只需两个参数,且对数据库内的点的次序几乎不敏感(两个聚类之间边缘的点有机会受次序的影响被分到不同的聚类,另外聚类的次序会受点的次序的影响)。
# DBSCAN 被设计成能配合可加速范围访问的数据库结构,例如 R*树。
# 如果对资料有足够的了解,可以选择适当的参数以获得最佳的分类。
# ```
# 缺点
# ```
# DBSCAN 不是完全决定性的:在两个聚类交界边缘的点会视乎它在数据库的次序决定加入哪个聚类,幸运地,这种情况并不常见,而且对整体的聚类结果影响不大——DBSCAN 对核心点和噪音都是决定性的。DBSCAN* 是一种变化了的算法,把交界点视为噪音,达到完全决定性的结果。
# DBSCAN 聚类分析的质素受函数 regionQuery(P,ε) 里所使用的度量影响,最常用的度量是欧几里得距离,尤其在高维度资料中,由于受所谓“维数灾难”影响,很难找出一个合适的 ε ,但事实上所有使用欧几里得距离的算法都受维数灾难影响。
# 如果数据库里的点有不同的密度,而该差异很大,DBSCAN 将不能提供一个好的聚类结果,因为不能选择一个适用于所有聚类的 minPts-ε 参数组合。
# 如果没有对资料和比例的足够理解,将很难选择适合的 ε 参数。
# ```
# +
def distance(data):
'''计算样本点之间的距离
:param data(mat):样本
:return:dis(mat):样本点之间的距离
'''
m, n = np.shape(data)
dis = np.mat(np.zeros((m, m)))
for i in range(m):
for j in range(i, m):
# 计算i和j之间的欧式距离
tmp = 0
for k in range(n):
tmp += (data[i, k] - data[j, k]) * (data[i, k] - data[j, k])
dis[i, j] = np.sqrt(tmp)
dis[j, i] = dis[i, j]
return dis
def find_eps(distance_D, eps):
'''找到距离≤eps的样本的索引
:param distance_D(mat):样本i与其他样本之间的距离
:param eps(float):半径的大小
:return: ind(list):与样本i之间的距离≤eps的样本的索引
'''
ind = []
n = np.shape(distance_D)[1]
for j in range(n):
if distance_D[0, j] <= eps:
ind.append(j)
return ind
def dbscan(data, eps, MinPts):
'''DBSCAN算法
:param data(mat):需要聚类的数据集
:param eps(float):半径
:param MinPts(int):半径内最少的数据点数
:return:
types(mat):每个样本的类型:核心点、边界点、噪音点
sub_class(mat):每个样本所属的类别
'''
m = np.shape(data)[0]
# 在types中,1为核心点,0为边界点,-1为噪音点
types = np.mat(np.zeros((1, m)))
sub_class = np.mat(np.zeros((1, m)))
# 用于判断该点是否处理过,0表示未处理过
dealt = np.mat(np.zeros((m, 1)))
# 计算每个数据点之间的距离
dis = distance(data)
# 用于标记类别
number = 1
# 对每一个点进行处理
for i in range(m):
# 找到未处理的点
if dealt[i, 0] == 0:
# 找到第i个点到其他所有点的距离
D = dis[i,]
# 找到半径eps内的所有点
ind = find_eps(D, eps)
# 区分点的类型
# 边界点
if len(ind) > 1 and len(ind) < MinPts + 1:
types[0, i] = 0
sub_class[0, i] = 0
# 噪音点
if len(ind) == 1:
types[0, i] = -1
sub_class[0, i] = -1
dealt[i, 0] = 1
# 核心点
if len(ind) >= MinPts + 1:
types[0, i] = 1
for x in ind:
sub_class[0, x] = number
# 判断核心点是否密度可达
while len(ind) > 0:
dealt[ind[0], 0] = 1
D = dis[ind[0],]
tmp = ind[0]
del ind[0]
ind_1 = find_eps(D, eps)
if len(ind_1) > 1: # 处理非噪音点
for x1 in ind_1:
sub_class[0, x1] = number
if len(ind_1) >= MinPts + 1:
types[0, tmp] = 1
else:
types[0, tmp] = 0
for j in range(len(ind_1)):
if dealt[ind_1[j], 0] == 0:
dealt[ind_1[j], 0] = 1
ind.append(ind_1[j])
sub_class[0, ind_1[j]] = number
number += 1
# 最后处理所有未分类的点为噪音点
ind_2 = ((sub_class == 0).nonzero())[1]
for x in ind_2:
sub_class[0, x] = -1
types[0, x] = -1
return types, sub_class
# -
types, P = dbscan(X, 0.1, 4)
# ### 7.优缺点
# 
# ### 8.sklearn参数详解
# 接下来就通过查看sklearn的参数去完成自己的例子
# https://sklearn.apachecn.org/docs/0.21.3/22.html
# 参考文献:
#
| origin_data/Task4_cluster.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + code_folding=[]
#matplotlib inline
from __future__ import division
import numpy as np
from numpy.random import rand
from numpy import linalg as LA
import matplotlib
import matplotlib.pyplot as plt
from scipy import interpolate
from matplotlib.patches import Arrow, Circle, Rectangle, Ellipse
from matplotlib.patches import ConnectionPatch, Polygon
from matplotlib import rc
rc('font',**{'family':'sans-serif', 'size' : 19}) #, 'sans-serif':['Arial']})
## for Palatino and other serif fonts use:
#rc('font',**{'family':'serif','serif':['Palatino']})
rc('text', usetex=True)
# %matplotlib inline
#info on phase diagram
#black dot -> Q=1/3 vortices unbind
#red dot -> Q=1 vortices unbind
#green triangles -> cv max
#list of tcs at L=40
list_of_everything = np.loadtxt('tcs.data')
lambda3=2.1
#fraction=j2/j6
#temperature range
Tmax = 1.6
Tmax_plot = 1.6
Tmin = 0.6
fig, ax = plt.subplots(figsize = (10, 10) )
#lambda = 0 KT points
tkt = 0.89
#plotting the two bare KT transitions
"""
plt.plot([0,2],[2*tkt,0], '--', color="Blue");
plt.plot([0,2],[0,2*tkt], '--', color="Blue");
"""
#all_cross = [[stiff_cross_j2, '*', 'black'], [sp_heat_cross_j2, '*', 'blue'], [binder_potts_j2, 'o', 'blue']]
#plot the black dotted box of the inside part
#plt.plot([0.5, 1.5], [Tmin, Tmin], color = 'black', linestyle = '--')
#plt.plot([0.5, 1.5], [Tmax, Tmax], color = 'black', linestyle = '--')
patches_stiff = []
patches_cv = []
patches_stiff2 = []
patches_cv2 = []
range_J2 = []
ixB = []
iyB = []
ixC = []
iyC = []
fP = []
fP_x = []
fKT1 = []
fKT1_x = []
fKT2 = []
fKT2_x = []
radius = 0.015
for i in range(len(list_of_everything)):
vals = list_of_everything[i]
if vals[3] == 0:
col = 'mediumpurple'
else:
col = 'teal'
#patches_stiff.append(Circle((vals[0], vals[2]), radius=0.005, facecolor=col, edgecolor = 'black', zorder =5))
#patches_stiff2.append(Circle((vals[0], vals[2]), radius=0.005, facecolor=col, edgecolor = 'black', zorder = 5))
patches_stiff2.append(Ellipse((vals[0], vals[2]), width=radius/2, height = radius, facecolor=col, edgecolor = 'black', zorder = 5))
range_J2.append(vals[0])
if 0.85 <= vals[0] <= 1.15:
ixB.append(vals[0])
ixC.append(vals[0])
iyB.append(vals[2])
if vals[0] <= 1.15:
fP_x.append(vals[0])
if vals[0] <= 0.85:
fKT1.append(vals[2])
fKT1_x.append(vals[0])
if 0.85 <= vals[0]:
fKT2.append(vals[2])
fKT2_x.append(vals[0])
range_J2 = np.array(range_J2)
N_cp = 40
Kc = 0.0
range_T = np.linspace(Tmin + 0.0001, Tmax, 60)
#print(range_T)
initial_cv_val = np.loadtxt('CV_data_pd.txt')
gridplot_cv = np.zeros((len(range_T), len(range_J2)))
for j in range(len(range_J2)):
#cv
#gridplot_cv[:,j] = (final_cv_val)
#log of cv
gridplot_cv[:,j] = np.log(initial_cv_val[:,j])
#get cv_max for that size
initial_cv_val_here = initial_cv_val[:,j]
maxcv = range_T[np.where(initial_cv_val_here == np.max(initial_cv_val_here))[0][0]]
#print(maxcv)
if range_J2[j] > 1.2:
maxcv = list_of_everything[j][1]
if range_J2[j] <= 1.15:
patches_cv2.append(Ellipse((range_J2[j], maxcv), width=radius/2, height = radius, facecolor='red', edgecolor = 'black', zorder = 6))
#patches_cv2.append(Circle((range_J2[j], maxcv), radius=0.005, facecolor='red', edgecolor = 'black', zorder = 6))
else:
#not used here
patches_cv.append(Rectangle((range_J2[j]- 0.01, maxcv - 0.01), 0.01, 0.001, facecolor='red', edgecolor = 'black', zorder = 6))
patches_cv2.append(Rectangle((range_J2[j] - 0.01, maxcv - 0.01), 0.01, 0.01, facecolor='red', edgecolor = 'black', zorder = 6))
if 0.85 <= range_J2[j] <= 1.15:
iyC.append(maxcv)
if range_J2[j] <= 1.15:
fP.append(maxcv)
ixB = np.array(ixB)[::-1]
ixC = np.array(ixC)
iyB = np.array(iyB)[::-1]
iyC = np.array(iyC)
im = ax.imshow(gridplot_cv, interpolation='spline16', cmap='YlGn',origin='lower',\
extent = [0.5 - 0.025, 1.5 + 0.025, 0.6 - 1/(2*59), 1.6 + 1/(2*59)])
#clb = plt.colorbar(im, shrink=0.5)
#clb.ax.tick_params(labelsize=12)
#clb.ax.set_title(r'$C_v/N$', fontsize = 12)
#clb.ax.set_title(r'$\log \; C_v$', fontsize = 12)
x1, x2, y1, y2 = 0.8, 1.15, 1.05, 1.3
ax.set_xlim(x1, x2)
ax.set_ylim(y1, y2)
plt.xlabel('$\Delta$', fontsize=20);
plt.ylabel('$\mathrm{Temperature} \; T$', fontsize=20)
#ticks
major_ticks_x = np.arange(0.8, 1.15 + 0.01, 0.05)
minor_ticks_x = np.arange(0.8, 1.15 + 0.01, 0.025)
major_ticks_y = np.arange(1.05, 1.3 + 0.01, 0.025)
minor_ticks_y = np.arange(1.05, 1.3 + 0.01, 0.025/2)
tick_print_x = []
for elem in major_ticks_x:
tick_print_x.append('${:.2f}$'.format(elem))
tick_print_y = []
for elem in major_ticks_y:
tick_print_y.append('${:.2f}$'.format(elem))
ax.set_xticks(major_ticks_x)
ax.set_yticks(major_ticks_y)
#ax.set_xticklabels(tick_print_x, fontsize = 16, rotation = 310)
ax.set_xticklabels(tick_print_x, fontsize = 16)
ax.set_yticklabels(tick_print_y, fontsize = 16)
ax.set_xticks(minor_ticks_x, minor=True)
ax.set_yticks(minor_ticks_y, minor=True)
#ax.set_xticklabels(tick_print, rotation=315)
ax.grid(which='minor', alpha=0.3)
ax.grid(which='major', alpha=0.6)
#ax.set_xlim([0,2])
#ax.set_ylim([0,Tmax_plot])
#ax.xaxis.set_label_coords(1.08, -0.03)
"""
textstr = r'III'
ax.text(0.8, 0.2, textstr, transform=ax.transAxes, fontsize=20,
verticalalignment='top', bbox= dict(boxstyle='square', fc="none", ec="k"))
textstr = r'I'
ax.text(0.1, 0.8, textstr, transform=ax.transAxes, fontsize=20,
verticalalignment='top', bbox= dict(boxstyle='square', fc="none", ec="k"))
textstr = r'IV'
ax.text(0.39, 0.59, textstr, transform=ax.transAxes, fontsize=20,
verticalalignment='top', bbox= dict(boxstyle='square', fc="none", ec="k"))
"""
#insert a shaded region
verts = [*zip(ixC, iyC), *zip(ixB, iyB)]
poly = Polygon(verts, facecolor='crimson', edgecolor='none', alpha = 0.6)
ax.add_patch(poly)
ax.plot(fP_x, fP, color = 'red')
ax.plot(fKT1_x, fKT1, color = 'mediumpurple')
ax.plot(fKT2_x, fKT2, color = 'teal')
for p in patches_stiff2:
ax.add_patch(p)
for ps in patches_cv2:
ax.add_patch(ps)
###########################
#####inset
###########################
#ax.set_ylim([0.6, 1.6])
#ax.set_ylim([0,Tmax_plot])
#ax.indicate_inset_zoom(axins)
ax.set_aspect(0.5)
plt.tight_layout()
plt.savefig('./fig-phasediagram-inset-cv.png', format='png',dpi = 100, bbox_inches='tight')
plt.show()
# -
| .ipynb_checkpoints/Phase Diagram-only inset-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: STRIFE_clone
# language: python
# name: strife_clone
# ---
# # Import STRIFE code and default arguments
# +
from STRIFE import STRIFE #STRIFE module
from parse_args import parse_args #Get all of the default arguments for STRIFE
from rdkit import Chem
from rdkit.Chem.Draw import IPythonConsole
# -
# %config Completer.use_jedi = False
import warnings
warnings.filterwarnings("ignore", category=DeprecationWarning)
args = parse_args()
# # Set arguments to run STRIFE
#
# To run STRIFE, we need to specify a few key arguments. You can find a full list of the arguments (and a brief comment on what each of them does) either by opening the ```parse_args.py``` file, or by running ```python STRIFE.py --help``` in the command line.
#
# In this notebook, we're going to run STRIFE on the default setting - i.e. the way in which the algorithm is outlined in the paper. For the sake of brevity, we've already computed the Fragment Hotspot Map and provide it to STRIFE as an input - alternatively, we could have specified a directory where we would like to save a Fragment Hotspot Map and STRIFE would calculate it and save it there for us by using the ```calculate_hotspots``` argument.
#
# ### Arguments we need to specify:
#
# ```protein```: The path to a pdb file which contains our protein of interest - the PDB file needs to prepared so that it can be used by GOLD to dock ligands. More information can be found [here](https://www.ccdc.cam.ac.uk/support-and-resources/ccdcresources/GOLD_User_Guide.pdf), but generally, you need to remove any ligands and waters and ensure that the protein has been protonated.
#
#
# Specifying the fragment of interest:
#
# There are two ways to tell STRIFE which fragment to elaborate (and which exit vector you want to make elaborations from).
#
# * Use the ```fragment_SDF``` argument to specify the structure of the fragment you want to elaborate. This must be a bound fragment that fits in the ```protein``` binding site. We also have to specify an ```exit_vector_idx``` - this is the index of the atom that the elaborations will be generated from. We have written a script ```specifyExitVector.py``` (see the README for more info) that you can use to help you identify the index of the atom you want to elaborate from
#
# * Alternatively, we can specify a ```fragment_SDF``` and ```fragment_smiles```. ```fragment_smiles``` is a SMILES string of the desired fragment, where the exit vector is denoted by a dummy atom (again ```specifyExitVector.py``` can help you obtain this SMILES string). You can provide either the raw string as an argument, or a file in which the SMILES string is saved.
#
# Storing the output:
#
# * Specify the directory you would like to store the output in as ```output_directory``` - if the directory doesn't already exist then it will be created.
#
args.protein = 'example/1q8t_protein.pdb' #PDB file to be the protein - ADAPT THIS LINE AS NEEDED FOR YOUR OWN RUNS
args.fragment_sdf = 'example/1q8t_frag.sdf'
args.fragment_smiles = 'example/1q8t_frag_smiles.smi'
args.output_directory = 'example/STRIFE_1Q8T'
args.hotspots_output = 'example/hotspotsOut/out.zip'
#args.calculate_hotspots = 'example/hotspotsOut' #This line is just to illustrate how to ask STRIFE how to precalculate an FHM if you haven't already
#^^ Don't uncomment if you're using args.hotspots_output
args.num_cpu_cores = 7
args.write_elaborations_dataset = True
# # Running STRIFE
#
# A common error with the below cell is something along the lines that 'NoneType' has no attribute 'GetNumHeavyAtoms()'. This is typically caused by providing empty acceptorHotspot.sdf or donorHotspot.sdf files (if you're not providing any Acceptor pharmacophoric points, simply omit that file from the output directory, and the same for Donor points).
#
# When selecting your pharmacophoric points in PyMol, make sure that when you save them, you use the state=0 argument. E.g.:
#
# (in PyMol command line)
# ```save path/to/output/directory/acceptorHotspot.sdf, HBA_selection, state = 0```
#
#
#
#Create the STRIFE class
STRIFE_model = STRIFE(args)
#Run STRIFE
STRIFE_model.run(args)
# # View the generated elaborations
ranked_elabs = STRIFE_model.pharmElabsTestLigEff
ranked_elabs.head(10)
# ### Now let's visualise the most highly ranked elaborations
Chem.Draw.MolsToGridImage([Chem.MolFromSmiles(s) for s in ranked_elabs['smiles'].drop_duplicates().head(16)], molsPerRow = 4)
# If we want to, we can view the quasi actives which determined the pharmacophoric profiles used in the refinement phase:
# +
#We can view basic information about each pharmacophoric point
for k in STRIFE_model.hSingles.keys():
print(f'****Pharmacophoric point: {k}****')
for kk in STRIFE_model.hSingles[k].keys():
print(f'{kk} : {STRIFE_model.hSingles[k][kk]}')
print('\n')
# +
#There is a set of quasi-actives for each pharmacophoric point
for k in STRIFE_model.singleQuasiActives.keys():
print(f'Quasi-Actives for pharmacophoric point {k}:')
print(STRIFE_model.singleQuasiActives[k])
print('\n')
# +
all_quasi_actives = []
for k in STRIFE_model.singleQuasiActives.keys():
all_quasi_actives = all_quasi_actives + list(STRIFE_model.singleQuasiActives[k]['smiles'])
all_quasi_actives = list(set(all_quasi_actives))
Chem.Draw.MolsToGridImage([Chem.MolFromSmiles(s) for s in all_quasi_actives])
# -
# ### Accessing the docked poses
#
# STRIFE saves the docked poses in the ```output_directory``` under the name ```pharmsElabsTestDocked.sdf```. You can view them in the binding pocket using a molecule viewer such as PyMol
docked_mols = Chem.SDMolSupplier(f'{args.output_directory}/pharmsElabsTestDocked.sdf')
docked_mols[0] #A 2D depiction of one of the docked mols
| STRIFE_notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import sys
import gin
import numpy as np
import pandas as pd
module_path = os.path.abspath(os.path.join('..'))
if module_path not in sys.path:
sys.path.append(module_path)
from IPython.core.display import clear_output, display
from ariadne.graph_net.graph_utils.graph_prepare_utils import to_pandas_graph_from_df, get_pd_line_graph, \
apply_nodes_restrictions, apply_edge_restriction, construct_output_graph
from ariadne.transformations import Compose, ConstraintsNormalize, ToCylindrical
import matplotlib.pyplot as plt
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
# +
from prepare import parse
csv_params = {
"sep": '\s+',
"nrows": 15000,
"encoding": 'utf-8',
"names": ['event', 'x', 'y', 'z', 'station', 'track', 'px', 'py', 'pz', 'X0', 'Y0', 'Z0']
}
suff_df = ('_p', '_c')
gin.bind_parameter('get_pd_line_graph.restrictions_0',(-0.2, 0.2))
gin.bind_parameter('get_pd_line_graph.restrictions_1', (-0.28, 0.28))
gin.bind_parameter('get_pd_line_graph.suffix_c', '_c')
gin.bind_parameter('get_pd_line_graph.suffix_p','_p')
gin.bind_parameter('get_pd_line_graph.spec_kwargs', {'suffix_c': '_c',
'suffix_p':'_p',
'axes':['r', 'phi', 'z']} )
#gin.bind_parameter('GraphNet_Processor.get_supernodes_df.suffix_c', '_c')
#gin.bind_parameter('GraphNet_Processor.get_supernodes_df.suffix_p', '_p')
#gin.bind_parameter('GraphNet_Processor.get_supernodes_df.axes', ['r', 'phi', 'z'])
#gin.bind_parameter('GraphNet_Processor.apply_edge_restriction.edge_restriction', 0.093)
_edge_restriction = 0.093
stations_constraints = {
0: {'x': [-166.6, 166.6], 'y': [-166.6, 166.6], 'z': [-423.5, 423.5]},
1: {'x': [-166.6, 166.6], 'y': [-166.6, 166.6], 'z': [-423.5, 423.5]},
2: {'x': [-166.6, 166.6], 'y': [-166.6, 166.6], 'z': [-423.5, 423.5]},
}
# -
events = parse("/zfs/hybrilit.jinr.ru/user/p/pgonchar/data/bes3/events/3.txt", csv_params=csv_params, events_quantity=':')
events = next(events)[0]
n_events = 312
events_to_analyze = events[events.event < n_events]
events_to_analyze
# +
from collections import namedtuple
GraphWithIndices = namedtuple('Graph', ['X', 'Ri', 'Ro', 'y', 'v', 'v_from_to','e_from_to', 'ev_id' ])
def construct_graph_with_indices(graph, v_inds, v_from_to, e_from_to, ev_id):
return GraphWithIndices(graph.X, graph.Ri, graph.Ro, graph.y, v_inds, v_from_to, e_from_to, ev_id)
# +
nodes = pd.DataFrame()
edges = pd.DataFrame()
graphs = {}
for idx, event in events_to_analyze.groupby('event'):
clear_output(wait=True)
display("Event #%09d" % idx)
transformer = Compose([
ConstraintsNormalize(
use_global_constraints=False,
constraints=stations_constraints
),
ToCylindrical(drop_old=True)
])
try:
event = transformer(event)
except AssertionError as err:
print("ASS error %r" % err)
continue
G = to_pandas_graph_from_df(event, suffixes=suff_df, compute_is_true_track=True)
nodes_t, edges_t = get_pd_line_graph(G, apply_nodes_restrictions)
edges_filtered = apply_edge_restriction(edges_t, edge_restriction=_edge_restriction)
graph = construct_output_graph(nodes_t, edges_filtered, ['y_p', 'y_c', 'z_p', 'z_c', 'z'],
[np.pi, np.pi, 1., 1., 1.], 'edge_index_p', 'edge_index_c')
ev_id = event.event.values[0]
graph_with_inds = construct_graph_with_indices(graph,
nodes_t.index.values,
nodes_t[['from_ind','to_ind']].values,
edges_filtered[['edge_index_p','edge_index_c']].values,
ev_id)
np.testing.assert_allclose(graph_with_inds.X,
nodes_t[['y_p', 'y_c', 'z_p', 'z_c', 'z']].values/[np.pi, np.pi, 1., 1., 1.])
assert ev_id not in graphs
graphs[ev_id] = graph_with_inds
# +
from ariadne.graph_net.model import GraphNet_v1
import torch
gin.bind_parameter('GraphNet_v1.input_dim', 5)
gin.bind_parameter('GraphNet_v1.hidden_dim', 128)
gin.bind_parameter('GraphNet_v1.n_iters', 1)
def weights_update(model, checkpoint):
model_dict = model.state_dict()
pretrained_dict = checkpoint['state_dict']
real_dict = {}
for (k,v) in model_dict.items():
needed_key = None
for pretr_key in pretrained_dict:
if k in pretr_key:
needed_key = pretr_key
break
assert needed_key is not None, "key %s not in pretrained_dict %r!" % (k, pretrained_dict.keys())
real_dict[k] = pretrained_dict[needed_key]
model.load_state_dict(real_dict)
model.eval()
return model
path = '/zfs/hybrilit.jinr.ru/user/g/gooldan/bes/ariadne/lightning_logs/version_63115/checkpoints/epoch=49.ckpt'
checkpoint = torch.load(path) if torch.cuda.is_available() else torch.load(path, map_location=torch.device('cpu'))
model = weights_update(model=GraphNet_v1(),
checkpoint=checkpoint)
# +
from ariadne.graph_net.dataset import GraphDatasetFromMemory, collate_fn
from ariadne.graph_net.data_loader import GraphDataLoader
graphs_list = list(graphs.values())
gin.bind_parameter('GraphDatasetFromMemory.input_graphs', graphs_list)
data_loader = GraphDataLoader(
batch_size=1,
dataset=GraphDatasetFromMemory,
collate_fn=collate_fn,
n_train=0,
n_valid=len(graphs_list)-1
).get_val_dataloader()
# -
a, b, c = list(iter(data_loader))[0][0]['inputs']
a
with torch.no_grad():
test_outputs = [(model(batch_input['inputs']).flatten(), batch_target.flatten())
for (batch_input, batch_target) in data_loader]
test_pred, test_target = zip(*test_outputs)
test_pred = np.concatenate(test_pred)
test_target = np.concatenate(test_target)
import sklearn.metrics
thresh = 0.5
y_pred, y_true = (test_pred > thresh), (test_target > thresh)
print('Test set results with threshold of', thresh)
print('Accuracy: %.6f' % sklearn.metrics.accuracy_score(y_true, y_pred))
print('Precision: %.6f' % sklearn.metrics.precision_score(y_true, y_pred))
print('Recall: %.6f' % sklearn.metrics.recall_score(y_true, y_pred))
# +
fpr, tpr, _ = sklearn.metrics.roc_curve(y_true, test_pred)
fig, (ax0, ax1) = plt.subplots(ncols=2, figsize=(12,5))
# Plot the model outputs
binning=dict(bins=50, range=(0,1), histtype='bar', log=True)
ax0.hist(test_pred[y_true==False], label='fake', **binning)
ax0.hist(test_pred[y_true], label='true', **binning)
ax0.set_xlabel('Model output')
ax0.legend(loc=0)
# Plot the ROC curve
auc = sklearn.metrics.auc(fpr, tpr)
ax1.plot(fpr, tpr)
ax1.plot([0, 1], [0, 1], '--')
ax1.set_xlabel('False positive rate')
ax1.set_ylabel('True positive rate')
ax1.set_title('ROC curve, AUC = %.3f' % auc)
plt.tight_layout()
# +
from itertools import chain
import datetime
total_events = 0
total_tracks = 0
total_tracks_true = 0
total_events_true = 0
total_hits = 0
total_hits_true = 0
all_tracks_df = pd.DataFrame(columns=['event', 'track', 'hit_0_id', 'hit_1_id', 'hit_2_id', 'px', 'py', 'pz', 'pred'])
all_events_df = pd.DataFrame(columns=['event','multiplicity','pred', 'time'])
for idx, event in events_to_analyze.groupby('event'):
ev_id_real = event.event.values[0]
clear_output(wait=True)
display("Event #%09d" % idx)
hits_in_event = set()
tracks_in_event = event[event.track != -1].track.nunique()
for tr_id, track in event.groupby('track'):
if tr_id != -1:
hits_in_event.update(track.index.values)
all_tracks_df = all_tracks_df.append({
'event':int(ev_id_real),
'track':int(tr_id),
'hit_0_id':int(track.index.values[0]),
'hit_1_id':int(track.index.values[1] if len(track.index.values) > 1 else -1),
'hit_2_id':int(track.index.values[2] if len(track.index.values) > 2 else -1),
'px':track.px.values[0],
'py':track.py.values[0],
'pz':track.pz.values[0],
'pred':int(0),
'multiplicity':int(tracks_in_event)
}, ignore_index=True)
#total_hits_inds.extend(hits_in_event)
total_tracks += tracks_in_event
total_hits += len(hits_in_event)
total_events += 1
if ev_id_real not in graphs:
continue
graphed_event = event[['event', 'x', 'y','z','track', 'station']]
time_start = datetime.datetime.now()
G = to_pandas_graph_from_df(graphed_event, suffixes=suff_df, compute_is_true_track=True)
tgt_graph = graphs[ev_id_real]
batch_input, batch_target = collate_fn([tgt_graph])
with torch.no_grad():
y_pred = model(batch_input['inputs']).numpy().flatten() > 0.5
time_end = datetime.datetime.now()
time_diff = (time_end - time_start).microseconds
superedge_pred = tgt_graph.e_from_to[y_pred]
edges_pred = [G.loc[pred]
for pred in superedge_pred]
reco_tracks=0
edge_ind_pred = [(edgs.iloc[0].index_old_p,
edgs.iloc[0].index_old_c,
edgs.iloc[1].index_old_c) for edgs in edges_pred]
for track_cand in edge_ind_pred:
atdv = all_tracks_df
match = ((atdv.hit_0_id == track_cand[0]) &
(atdv.hit_1_id == track_cand[1]) &
(atdv.hit_2_id == track_cand[2]))
matched_track = all_tracks_df[match]
assert len(matched_track) < 2
if matched_track.empty:
all_tracks_df = all_tracks_df.append({
'event':int(ev_id_real),
'track':-1,
'hit_0_id':int(track_cand[0]),
'hit_1_id':int(track_cand[1]),
'hit_2_id':int(track_cand[2]),
'px':np.inf,
'py':np.inf,
'pz':np.inf,
'pred':-1,
'multiplicity':int(tracks_in_event)
}, ignore_index=True)
else:
all_tracks_df.loc[match, 'pred'] = 1
total_tracks_true += 1
reco_tracks += 1
hits_pred = set(chain.from_iterable(edge_ind_pred))
found_hits = hits_pred.intersection(hits_in_event)
total_hits_true += len(found_hits)
preddd=0
if reco_tracks == tracks_in_event:
total_events_true += 1
preddd=1
all_events_df = all_events_df.append({
'event':int(ev_id_real),
'multiplicity':int(tracks_in_event),
'pred':preddd,
'time':time_diff
}, ignore_index=True)
all_tracks_df = all_tracks_df.astype({
'event': 'int32',
'track': 'int32',
'hit_0_id': 'int32',
'hit_1_id': 'int32',
'hit_2_id': 'int32',
'px': 'float32',
'py': 'float32',
'pz': 'float32',
'pred':'int32',
'multiplicity':'int32'
})
all_events_df = all_events_df.astype({
'event': 'int32',
'pred':'int32',
'multiplicity':'int32',
'time':'int32'
})
# -
print("Total track efficiency: %.04f" % (total_tracks_true / total_tracks))
print("Total hit efficiency: %.04f" % (total_hits_true / total_hits))
print("Total event efficiency: %.04f" % (total_events_true / total_events))
print("Mean event processing speed: %.02f" % (all_events_df.time.mean()))
all_events_df.pred.unique()
# +
# shortcut
from numpy import linalg as LA
all_tracks_df['pt'] = LA.norm(all_tracks_df[['px','py']].values, axis=1)
all_tracks_df['cos_t'] = (all_tracks_df[['pz']].values/ LA.norm(all_tracks_df[['px','py','pz']].values, axis=1, keepdims=True))
all_tracks_df['a_phi'] = np.arctan2(all_tracks_df[['px']].values, all_tracks_df[['py']].values)
# +
def get_diagram_arr_linspace(all_real_hits, found_hits, start, end, num, col):
spac = np.linspace(start, end, num=num)
step = (spac[1] - spac[0]) / 2
arr = []
spac_ret = []
for i in range(len(spac)-1):
beg = spac[i]
end = spac[i+1]
elems_real = all_real_hits[(all_real_hits[col] >= beg) & (all_real_hits[col] < end)]
elems_pred = found_hits[(found_hits[col] >= beg) & (found_hits[col] < end)]
#print(beg,end)
#print(len(elems_pred), len(elems_real))
if elems_real.empty:
# print("emp", beg, end)
continue
arr.append(len(elems_pred) / len(elems_real))
spac_ret.append(spac[i] + step)
return np.array(arr), np.array(spac_ret)
from scipy import stats
def get_diagram_for_boxplot(all_real_hits, found_hits, start, end, num, col, bin_array, int_mode):
y_vals, x_vals = get_diagram_arr_linspace(all_real_hits, found_hits, start, end, num, col)
np_y_vals = np.array(y_vals)
bin_array = bin_array
res_x_array = []
res_box_data = []
mean_box_array = []
for i in range(len(bin_array)-1):
beg = bin_array[i]
end = bin_array[i+1]
y_ind = np.where((x_vals>=beg) & (x_vals<end))
y_vals_corr = np_y_vals[y_ind]
y_vals_corr = y_vals_corr[~np.isnan(y_vals_corr)]
if len(y_vals_corr) == 0:
continue
#print(beg, end, i, stats.describe(y_vals_corr))
#print(y_vals_corr)
# if len(np.where(y_vals_corr < 1)) > 0:
# print(y_vals_corr)
# print(y_vals)
# print(beg,end)
res_box_data.append(y_vals_corr)
delta = 0 if int_mode else (end-beg)/2
res_x_array.append(beg + delta)
mean_box_array.append(np.mean(y_vals_corr))
return res_box_data, np.array(res_x_array), np.array(mean_box_array)
#plt.boxplot(res_box_data, positions=bin_array)
# +
from scipy.interpolate import make_interp_spline, BSpline
def boxplot_style(bp):
for box in bp['boxes']:
# change outline color
#box.set( color='#7570b3', linewidth=2)
# change fill color
box.set( facecolor = 'silver' )
## change color and linewidth of the whiskers
#for whisker in bp['whiskers']:
# whisker.set(color='#7570b3', linewidth=2)
#
### change color and linewidth of the caps
#for cap in bp['caps']:
# cap.set(color='#7570b3', linewidth=2)
#
### change color and linewidth of the medians
for median in bp['medians']:
median.set(color='tab:cyan', linewidth=3, alpha=0)
for median in bp['means']:
median.set(color='tab:green', linewidth=4, ls='-', zorder=5)
#
### change the style of fliers and their fill
#for flier in bp['fliers']:
# flier.set(marker='o', color='#e7298a', alpha=0.5)
def draw_for_col(tracks_real, tracks_pred_true,
col, col_pretty, total_events, n_ticks=150,
n_avg_ticks=-1, x_ticks=8,
custom_title=None,
ticks_custom=True,
with_boxplot=False,
int_mode=False, save_disk=True,
custom_draw_funcs=[],
diagram_func=None,
color_ax_ticks=False
):
color_ax_ticks = len(custom_draw_funcs) > 0 and color_ax_ticks
n_avg_ticks = n_ticks // 5 if n_avg_ticks < 0 else n_avg_ticks
delta = 1e-4 if not int_mode else 1
start = tracks_real[tracks_real[col] > -np.inf][col].min()
end = tracks_real[tracks_real[col] < np.inf][col].max()+delta
initial, spac = get_diagram_arr_linspace(tracks_real, tracks_pred_true, start, end, n_ticks, col)
maxX = int(end) if int_mode else end
plt.figure(figsize=(8,7))
ax = plt.subplot(111)
#plt.ylabel('Track efficiency', fontsize=12)
plt.xlabel(col_pretty, fontsize=12)
#plt.axis([0, maxX, 0, 1.005])
plt.plot(spac, initial, alpha=0.0, lw=1)
title = custom_title if custom_title else 'GraphNet_V1 track efficiency vs %s (%d events)' % (col_pretty, total_events)
plt.title(title, fontsize=14)
if not int_mode and ticks_custom:
plt.xticks(np.linspace(start, maxX, x_ticks))
else:
plt.locator_params(axis='x', nbins=x_ticks)
if diagram_func is None:
plt.yticks(np.round(np.linspace(0, 1, 11), decimals=2))
x_ticks = ax.get_xticks()
if with_boxplot:
old_ticks = x_ticks
delta_x = (x_ticks[1] - x_ticks[0])/2
diagram_func = get_diagram_for_boxplot if diagram_func is None else diagram_func
box_data, ticks_x, mean_data = diagram_func(tracks_real, tracks_pred_true, start, end,
n_ticks, col, x_ticks, int_mode)
bp = plt.boxplot(box_data, positions=ticks_x,
manage_ticks=False, meanline=True, showmeans=True,
widths=delta_x,patch_artist=True, sym='',zorder=3)
boxplot_style(bp)
# mean line
xnew = np.linspace(ticks_x.min(), ticks_x.max(), 500)
spl = make_interp_spline(ticks_x, mean_data, k=3) # type: BSpline
power_smooth = spl(xnew)
plt.plot(xnew, power_smooth, ls='--', color='tab:orange', label='mean efficiency', lw=3, zorder=4)
plt.xticks(old_ticks)
if diagram_func is None:
ax.set_ylim((-0.05, 1.05))
for draw_f in custom_draw_funcs:
draw_f(ax)
#plt.locator_params(axis='y', nbins=16)
ax.legend(loc=0)
if color_ax_ticks:
ax.tick_params(axis='y', labelcolor='tab:green')
ax.grid()
plt.tight_layout()
plt.rcParams['savefig.facecolor']='white'
os.makedirs('../output', exist_ok=True)
plt.savefig('../output/new_img_track_eff_%s_ev%r_t%d.png'%(col, total_events, n_ticks), dpi=300)
plt.show()
#from scipy.interpolate import make_interp_spline, BSpline
#second, spac2 = get_diagram_arr_linspace(tracks_real, tracks_pred_true, start, end, 25, 'pt')
#
#xnew = np.linspace(spac2.min(), spac2.max(), 30)
#
#spl = make_interp_spline(spac2, second, k=3) # type: BSpline
#power_smooth = spl(xnew)
# -
tracks_real = all_tracks_df[all_tracks_df.pred != -1]
tracks_pred_true = all_tracks_df[all_tracks_df.pred == 1]
draw_for_col(tracks_real, tracks_pred_true, 'pt', '$pt$', n_events, 80,
n_avg_ticks=48, x_ticks=14, ticks_custom=False, with_boxplot=True)
draw_for_col(tracks_real, tracks_pred_true, 'a_phi', '$a_\phi$',n_events, 350, n_avg_ticks=60, x_ticks=12, with_boxplot=True)
draw_for_col(tracks_real, tracks_pred_true, 'cos_t', '$cos_t$',20000, 350, n_avg_ticks=60,x_ticks=12, ticks_custom=False, with_boxplot=True)
events_real = all_events_df[all_events_df.pred != -1]
events_pred_true = all_events_df[all_events_df.pred == 1]
# +
def get_diagram_arr_linspaceer(all_real_hits, found_hits, start, end, num, col):
spac = np.linspace(start, end, num=num)
step = (spac[1] - spac[0]) / 2
arr = []
spac_ret = []
for i in range(len(spac)-1):
beg = spac[i]
end = spac[i+1]
elems_real = all_real_hits[(all_real_hits[col] >= beg) & (all_real_hits[col] < end)]
elems_pred = found_hits[(found_hits[col] >= beg) & (found_hits[col] < end)]
#print(beg,end)
#print(len(elems_pred), len(elems_real))
if elems_real.empty:
# print("emp", beg, end)
continue
arr.append(len(elems_real))
spac_ret.append(spac[i] + step)
return np.array(arr), np.array(spac_ret)
def custom_draw_dist(ax):
res_y, res_x = get_diagram_arr_linspaceer(events_real, events_real, 1, 13, 13, 'multiplicity')
res_x = res_x - 0.5
res_y = res_y
ax2 = ax.twinx() # instantiate a second axes that shares the same x-axis
color = 'tab:blue'
ax2.set_ylabel('amount', color=color) # we already handled the x-label with ax1
ax2.plot(res_x, res_y, color=color, label='events with such multiplicity')
ax2.tick_params(axis='y', labelcolor=color)
ax2.legend(loc=2)
#ax.set_yticks(np.round(np.linspace(0, 1, 11), decimals=2))
#plt.grid()
pass
draw_for_col(events_real, events_pred_true, 'multiplicity', 'multiplicity',20000, 13,
custom_title='GraphNet_V1 fully reconstructed events vs multiplicity (total %d events)' % (20000),
custom_draw_funcs=[custom_draw_dist],
n_avg_ticks=11,x_ticks=13, int_mode=True, ticks_custom=False, with_boxplot=True, color_ax_ticks=True)
# +
def get_diagram_arr_linspacer(all_real_hits, found_hits, start, end, num, col):
spac = np.linspace(start, end, num=num)
step = (spac[1] - spac[0]) / 2
arr = []
spac_ret = []
for i in range(len(spac)-1):
beg = spac[i]
end = spac[i+1]
elems_real = all_real_hits[(all_real_hits[col] >= beg) & (all_real_hits[col] < end)]
elems_pred = found_hits[(found_hits[col] >= beg) & (found_hits[col] < end)]
#print(beg,end)
#print(len(elems_pred), len(elems_real))
if elems_real.empty:
# print("emp", beg, end)
continue
arr.append(elems_real.time.values * 1e-3)
spac_ret.append(spac[i] + step)
return np.array(arr), np.array(spac_ret)
def get_diagram_for_boxplot_ex(all_real_hits, found_hits, start, end, num, col, bin_array, int_mode):
y_vals, x_vals = get_diagram_arr_linspacer(all_real_hits, found_hits, start, end, num, col)
np_y_vals = np.array(y_vals)
bin_array = bin_array
res_x_array = []
res_box_data = []
mean_box_array = []
for i in range(len(bin_array)-1):
beg = bin_array[i]
end = bin_array[i+1]
y_ind = np.where((x_vals>=beg) & (x_vals<end))
y_vals_corr = np_y_vals[y_ind]
if len(y_vals_corr) == 0:
continue
y_vals_corr = np.concatenate(y_vals_corr, axis=-1)
y_vals_corr = y_vals_corr[~np.isnan(y_vals_corr)]
if len(y_vals_corr) == 0:
continue
#print(beg, end, i, stats.describe(y_vals_corr))
#print(y_vals_corr)
# if len(np.where(y_vals_corr < 1)) > 0:
# print(y_vals_corr)
# print(y_vals)
# print(beg,end)
res_box_data.append(y_vals_corr)
delta = 0 if int_mode else (end-beg)/2
res_x_array.append(beg + delta)
mean_box_array.append(np.mean(y_vals_corr))
return res_box_data, np.array(res_x_array), np.array(mean_box_array)
draw_for_col(events_real, events_pred_true, 'multiplicity', 'multiplicity',20000, 13,
custom_title='GraphNet_V1 processing speed vs multiplicity (total %d events)' % (20000),
n_avg_ticks=11,x_ticks=13, int_mode=True, ticks_custom=False,
custom_draw_funcs=[
lambda ax:
ax.set_ylabel('time,\n $m sec$', fontsize=12,rotation=0, ha='right') #and
#ax.set_ylim((0,0.02))
],
with_boxplot=True, diagram_func=get_diagram_for_boxplot_ex,color_ax_ticks=False)
| notebooks/graphnet_evaluation_200621.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Analysis of inequality and energy-related CO2 emissions in Latin America and the Caribbean applying a Kaya factors decomposition methodology
#
# ### <NAME>
#
# IDA-LMDI method by Kaya factors
# +
# Cargar los datos
import pandas as pd
import numpy as np
import math as mt
df = pd.read_excel("dataset2.xlsx")
# +
# Calcular intensidad de CO2 para combustibles fosiles
def co2int_ff(fila):
resultado = (fila["emissions_fossils"]/fila["output_fossils"])
return resultado
df["co2int-ff"] = df.apply(co2int_ff, axis=1)
# +
# Calcular consumo de energia para combustibles fosiles
def fossils_tes(fila):
resultado = fila["output_fossils"]/(fila["tes"]*0.00027777777777778)
return resultado
df["fossils-tes"] = df.apply(fossils_tes, axis=1)
# +
# Calcular intensidad de energía
def tes_gdp(fila):
resultado = (fila["tes"]*0.00027777777777778)/(fila["gdp_ppp"]*1000000)
return resultado
df["tes-gdp"] = df.apply(tes_gdp, axis=1)
# +
# Calculo de pib per capita
def gdp_pop(fila):
resultado = (fila["gdp_ppp"]*1000000)/fila["pop"]
return resultado
df["gdp-pop"] = df.apply(gdp_pop, axis=1)
# +
#Calculo emisiones totales por año
total = df.groupby("year").sum()
# +
#IDA-LMDI decomposition analysis: 1990 - 2018
def l_a_b(a,b):
resultado = (a-b)/(np.log(a) - np.log(b))
return resultado
l_ctf_ct0_prom = l_a_b(total.loc[2018, "emissions_fossils"], total.loc[1990, "emissions_fossils"])
data1990 = df[df["year"]==1990].reset_index()
data2018 = df[df["year"]==2018].reset_index()
def l_a_b_fossils(a,b):
resultado = (a-b)/(np.log(a) - np.log(b))
return resultado
l_a_b_fossils = l_a_b_fossils(data2018["emissions_fossils"],data1990["emissions_fossils"])
l_a_b_fossils = l_a_b_fossils/l_ctf_ct0_prom
def ln_per_country(a,x,y):
resultado = a*(np.log(x/y))
return resultado
vw_per_country = ln_per_country(l_a_b_fossils,data2018["co2int-ff"],data1990["co2int-ff"])
vu_per_country = ln_per_country(l_a_b_fossils,data2018["fossils-tes"],data1990["fossils-tes"])
vh_per_country = ln_per_country(l_a_b_fossils,data2018["tes-gdp"],data1990["tes-gdp"])
vg_per_country = ln_per_country(l_a_b_fossils,data2018["gdp-pop"],data1990["gdp-pop"])
vp_per_country = ln_per_country(l_a_b_fossils,data2018["pop"],data1990["pop"])
vw_1990_2018 = mt.exp(vw_per_country.sum())
vu_1990_2018 = mt.exp(vu_per_country.sum())
vh_1990_2018 = mt.exp(vh_per_country.sum())
vg_1990_2018 = mt.exp(vg_per_country.sum())
vp_1990_2018 = mt.exp(vp_per_country.sum())
Vtot_1990_2018 = vw_1990_2018 * vu_1990_2018 * vh_1990_2018 * vg_1990_2018 * vp_1990_2018
V_1990_2018 = pd.DataFrame([(vw_1990_2018, vu_1990_2018, vh_1990_2018, vg_1990_2018, vp_1990_2018, Vtot_1990_2018)],
columns =('Vw_1990_2018', 'Vu_1990_2018', 'Vh_1990_2018', 'Vg_1990_2018', 'Vp_1990_2018', 'Vtot_1990_2018'))
V_1990_2018
# +
#IDA-LMDI decomposition analysis: 1990 - 1995
def l_a_b(a,b):
resultado = (a-b)/(np.log(a) - np.log(b))
return resultado
l_ctf_ct0_prom = l_a_b(total.loc[1995, "emissions_fossils"], total.loc[1990, "emissions_fossils"])
data1990 = df[df["year"]==1990].reset_index()
data1995 = df[df["year"]==1995].reset_index()
def l_a_b_fossils(a,b):
resultado = (a-b)/(np.log(a) - np.log(b))
return resultado
l_a_b_fossils = l_a_b_fossils(data1995["emissions_fossils"],data1990["emissions_fossils"])
l_a_b_fossils = l_a_b_fossils/l_ctf_ct0_prom
def ln_per_country(a,x,y):
resultado = a*(np.log(x/y))
return resultado
vw_per_country = ln_per_country(l_a_b_fossils,data1995["co2int-ff"],data1990["co2int-ff"])
vu_per_country = ln_per_country(l_a_b_fossils,data1995["fossils-tes"],data1990["fossils-tes"])
vh_per_country = ln_per_country(l_a_b_fossils,data1995["tes-gdp"],data1990["tes-gdp"])
vg_per_country = ln_per_country(l_a_b_fossils,data1995["gdp-pop"],data1990["gdp-pop"])
vp_per_country = ln_per_country(l_a_b_fossils,data1995["pop"],data1990["pop"])
vw_1990_1995 = mt.exp(vw_per_country.sum())
vu_1990_1995 = mt.exp(vu_per_country.sum())
vh_1990_1995 = mt.exp(vh_per_country.sum())
vg_1990_1995 = mt.exp(vg_per_country.sum())
vp_1990_1995 = mt.exp(vp_per_country.sum())
Vtot_1990_1995 = vw_1990_1995 * vu_1990_1995 * vh_1990_1995 * vg_1990_1995 * vp_1990_1995
V_1990_1995 = pd.DataFrame([(vw_1990_1995, vu_1990_1995, vh_1990_1995, vg_1990_1995, vp_1990_1995, Vtot_1990_1995)],
columns =('Vw_1990_1995', 'Vu_1990_1995', 'Vh_1990_1995', 'Vg_1990_1995', 'Vp_1990_1995', 'Vtot_1990_1995'))
V_1990_1995
# +
#IDA-LMDI decomposition analysis: 1995 - 2000
def l_a_b(a,b):
resultado = (a-b)/(np.log(a) - np.log(b))
return resultado
l_ctf_ct0_prom = l_a_b(total.loc[2000, "emissions_fossils"], total.loc[1995, "emissions_fossils"])
data1995 = df[df["year"]==1995].reset_index()
data2000 = df[df["year"]==2000].reset_index()
def l_a_b_fossils(a,b):
resultado = (a-b)/(np.log(a) - np.log(b))
return resultado
l_a_b_fossils = l_a_b_fossils(data2000["emissions_fossils"],data1995["emissions_fossils"])
l_a_b_fossils = l_a_b_fossils/l_ctf_ct0_prom
def ln_per_country(a,x,y):
resultado = a*(np.log(x/y))
return resultado
vw_per_country = ln_per_country(l_a_b_fossils,data2000["co2int-ff"],data1995["co2int-ff"])
vu_per_country = ln_per_country(l_a_b_fossils,data2000["fossils-tes"],data1995["fossils-tes"])
vh_per_country = ln_per_country(l_a_b_fossils,data2000["tes-gdp"],data1995["tes-gdp"])
vg_per_country = ln_per_country(l_a_b_fossils,data2000["gdp-pop"],data1995["gdp-pop"])
vp_per_country = ln_per_country(l_a_b_fossils,data2000["pop"],data1995["pop"])
vw_1995_2000 = mt.exp(vw_per_country.sum())
vu_1995_2000 = mt.exp(vu_per_country.sum())
vh_1995_2000 = mt.exp(vh_per_country.sum())
vg_1995_2000 = mt.exp(vg_per_country.sum())
vp_1995_2000 = mt.exp(vp_per_country.sum())
Vtot_1995_2000 = vw_1995_2000 * vu_1995_2000 * vh_1995_2000 * vg_1995_2000 * vp_1995_2000
V_1995_2000 = pd.DataFrame([(vw_1995_2000, vu_1995_2000, vh_1995_2000, vg_1995_2000, vp_1995_2000, Vtot_1995_2000)],
columns =('Vw_1995_2000', 'Vu_1995_2000', 'Vh_1995_2000', 'Vg_1995_2000', 'Vp_1995_2000', 'Vtot_1995_2000'))
V_1995_2000
# +
#IDA-LMDI decomposition analysis: 2000 - 2005
def l_a_b(a,b):
resultado = (a-b)/(np.log(a) - np.log(b))
return resultado
l_ctf_ct0_prom = l_a_b(total.loc[2005, "emissions_fossils"], total.loc[2000, "emissions_fossils"])
data2000 = df[df["year"]==2000].reset_index()
data2005 = df[df["year"]==2005].reset_index()
def l_a_b_fossils(a,b):
resultado = (a-b)/(np.log(a) - np.log(b))
return resultado
l_a_b_fossils = l_a_b_fossils(data2005["emissions_fossils"],data2000["emissions_fossils"])
l_a_b_fossils = l_a_b_fossils/l_ctf_ct0_prom
def ln_per_country(a,x,y):
resultado = a*(np.log(x/y))
return resultado
vw_per_country = ln_per_country(l_a_b_fossils,data2005["co2int-ff"],data2000["co2int-ff"])
vu_per_country = ln_per_country(l_a_b_fossils,data2005["fossils-tes"],data2000["fossils-tes"])
vh_per_country = ln_per_country(l_a_b_fossils,data2005["tes-gdp"],data2000["tes-gdp"])
vg_per_country = ln_per_country(l_a_b_fossils,data2005["gdp-pop"],data2000["gdp-pop"])
vp_per_country = ln_per_country(l_a_b_fossils,data2005["pop"],data2000["pop"])
vw_2000_2005 = mt.exp(vw_per_country.sum())
vu_2000_2005 = mt.exp(vu_per_country.sum())
vh_2000_2005 = mt.exp(vh_per_country.sum())
vg_2000_2005 = mt.exp(vg_per_country.sum())
vp_2000_2005 = mt.exp(vp_per_country.sum())
Vtot_2000_2005 = vw_2000_2005 * vu_2000_2005 * vh_2000_2005 * vg_2000_2005 * vp_2000_2005
V_2000_2005 = pd.DataFrame([(vw_2000_2005, vu_2000_2005, vh_2000_2005, vg_2000_2005, vp_2000_2005, Vtot_2000_2005)],
columns =('Vw_2000_2005', 'Vu_2000_2005', 'Vh_2000_2005', 'Vg_2000_2005', 'Vp_2000_2005', 'Vtot_2000_2005'))
V_2000_2005
# +
#IDA-LMDI decomposition analysis: 2005 - 2010
def l_a_b(a,b):
resultado = (a-b)/(np.log(a) - np.log(b))
return resultado
l_ctf_ct0_prom = l_a_b(total.loc[2010, "emissions_fossils"], total.loc[2005, "emissions_fossils"])
data2005 = df[df["year"]==2005].reset_index()
data2010 = df[df["year"]==2010].reset_index()
def l_a_b_fossils(a,b):
resultado = (a-b)/(np.log(a) - np.log(b))
return resultado
l_a_b_fossils = l_a_b_fossils(data2010["emissions_fossils"],data2005["emissions_fossils"])
l_a_b_fossils = l_a_b_fossils/l_ctf_ct0_prom
def ln_per_country(a,x,y):
resultado = a*(np.log(x/y))
return resultado
vw_per_country = ln_per_country(l_a_b_fossils,data2010["co2int-ff"],data2005["co2int-ff"])
vu_per_country = ln_per_country(l_a_b_fossils,data2010["fossils-tes"],data2005["fossils-tes"])
vh_per_country = ln_per_country(l_a_b_fossils,data2010["tes-gdp"],data2005["tes-gdp"])
vg_per_country = ln_per_country(l_a_b_fossils,data2010["gdp-pop"],data2005["gdp-pop"])
vp_per_country = ln_per_country(l_a_b_fossils,data2010["pop"],data2005["pop"])
vw_2005_2010 = mt.exp(vw_per_country.sum())
vu_2005_2010 = mt.exp(vu_per_country.sum())
vh_2005_2010 = mt.exp(vh_per_country.sum())
vg_2005_2010 = mt.exp(vg_per_country.sum())
vp_2005_2010 = mt.exp(vp_per_country.sum())
Vtot_2005_2010 = vw_2005_2010 * vu_2005_2010 * vh_2005_2010 * vg_2005_2010 * vp_2005_2010
V_2005_2010 = pd.DataFrame([(vw_2005_2010, vu_2005_2010, vh_2005_2010, vg_2005_2010, vp_2005_2010, Vtot_2005_2010)],
columns =('Vw_2005_2010', 'Vu_2005_2010', 'Vh_2005_2010', 'Vg_2005_2010', 'Vp_2005_2010', 'Vtot_2005_2010'))
V_2005_2010
# +
#IDA-LMDI decomposition analysis: 2010 - 2015
def l_a_b(a,b):
resultado = (a-b)/(np.log(a) - np.log(b))
return resultado
l_ctf_ct0_prom = l_a_b(total.loc[2015, "emissions_fossils"], total.loc[2010, "emissions_fossils"])
data2010 = df[df["year"]==2010].reset_index()
data2015 = df[df["year"]==2015].reset_index()
def l_a_b_fossils(a,b):
resultado = (a-b)/(np.log(a) - np.log(b))
return resultado
l_a_b_fossils = l_a_b_fossils(data2015["emissions_fossils"],data2010["emissions_fossils"])
l_a_b_fossils = l_a_b_fossils/l_ctf_ct0_prom
def ln_per_country(a,x,y):
resultado = a*(np.log(x/y))
return resultado
vw_per_country = ln_per_country(l_a_b_fossils,data2015["co2int-ff"],data2010["co2int-ff"])
vu_per_country = ln_per_country(l_a_b_fossils,data2015["fossils-tes"],data2010["fossils-tes"])
vh_per_country = ln_per_country(l_a_b_fossils,data2015["tes-gdp"],data2010["tes-gdp"])
vg_per_country = ln_per_country(l_a_b_fossils,data2015["gdp-pop"],data2010["gdp-pop"])
vp_per_country = ln_per_country(l_a_b_fossils,data2015["pop"],data2010["pop"])
vw_2010_2015 = mt.exp(vw_per_country.sum())
vu_2010_2015 = mt.exp(vu_per_country.sum())
vh_2010_2015 = mt.exp(vh_per_country.sum())
vg_2010_2015 = mt.exp(vg_per_country.sum())
vp_2010_2015 = mt.exp(vp_per_country.sum())
Vtot_2010_2015 = vw_2010_2015 * vu_2010_2015 * vh_2010_2015 * vg_2010_2015 * vp_2010_2015
V_2010_2015 = pd.DataFrame([(vw_2010_2015, vu_2010_2015, vh_2010_2015, vg_2010_2015, vp_2010_2015, Vtot_2010_2015)],
columns =('Vw_2010_2015', 'Vu_2010_2015', 'Vh_2010_2015', 'Vg_2010_2015', 'Vp_2010_2015', 'Vtot_2010_2015'))
V_2010_2015
# +
#IDA-LMDI decomposition analysis: 2015 - 2018
def l_a_b(a,b):
resultado = (a-b)/(np.log(a) - np.log(b))
return resultado
l_ctf_ct0_prom = l_a_b(total.loc[2018, "emissions_fossils"], total.loc[2015, "emissions_fossils"])
data2015 = df[df["year"]==2015].reset_index()
data2018 = df[df["year"]==2018].reset_index()
def l_a_b_fossils(a,b):
resultado = (a-b)/(np.log(a) - np.log(b))
return resultado
l_a_b_fossils = l_a_b_fossils(data2018["emissions_fossils"],data2015["emissions_fossils"])
l_a_b_fossils = l_a_b_fossils/l_ctf_ct0_prom
def ln_per_country(a,x,y):
resultado = a*(np.log(x/y))
return resultado
vw_per_country = ln_per_country(l_a_b_fossils,data2018["co2int-ff"],data2015["co2int-ff"])
vu_per_country = ln_per_country(l_a_b_fossils,data2018["fossils-tes"],data2015["fossils-tes"])
vh_per_country = ln_per_country(l_a_b_fossils,data2018["tes-gdp"],data2015["tes-gdp"])
vg_per_country = ln_per_country(l_a_b_fossils,data2018["gdp-pop"],data2015["gdp-pop"])
vp_per_country = ln_per_country(l_a_b_fossils,data2018["pop"],data2015["pop"])
vw_2015_2018 = mt.exp(vw_per_country.sum())
vu_2015_2018 = mt.exp(vu_per_country.sum())
vh_2015_2018 = mt.exp(vh_per_country.sum())
vg_2015_2018 = mt.exp(vg_per_country.sum())
vp_2015_2018 = mt.exp(vp_per_country.sum())
Vtot_2015_2018 = vw_2015_2018 * vu_2015_2018 * vh_2015_2018 * vg_2015_2018 * vp_2015_2018
V_2015_2018 = pd.DataFrame([(vw_2015_2018, vu_2015_2018, vh_2015_2018, vg_2015_2018, vp_2015_2018, Vtot_2015_2018)],
columns =('Vw_2015_2018', 'Vu_2015_2018', 'Vh_2015_2018', 'Vg_2015_2018', 'Vp_2015_2018', 'Vtot_2015_2018'))
V_2015_2018
# -
| IDA-LMDI.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 2 qubits gate
# Let's check the 2 qubits gate.
#
# ## What we'll learn this time
# 1. 2qubits gate
# 2. Implementation example
#
# ## Install Blueqat
# Install Blueqat from pip.
# + colab={"base_uri": "https://localhost:8080/", "height": 136} colab_type="code" id="Z2z6oBnToiqa" outputId="413f3f56-1b22-40f7-fcff-4621af4596d4"
# !pip install blueqat
# -
# ## Two qubit gate
# Two qubits gate is mainly one qubit gate with a control bit added to it.
#
# ### CX, CY, CZ
# CX, CY, CZ gates can be implemented by the following.
#
# | CX | CY | CZ |
# |:-:|:-:|:-:|
# |`cx`or`cnot`|`cy`|`cz`|
# #### Implementation
# Let's implement these gates.
#
# ###### CX gate
# +
from blueqat import Circuit
import math
Circuit().cx[0,1].m[:].run(shots=100)
#or
Circuit().cnot[0,1].m[:].run(shots=100)
# -
# ###### CY gate
Circuit().cy[0,1].m[:].run(shots=100)
# ###### CZ gate
Circuit().cz[0,1].m[:].run(shots=100)
# ## CX, CY, CZ gates
# The control gate consists of control bit and target bit.
#
# The control gate uses two qubits. These convert the second bit when the first bit is $|1\rangle$.
# Here is a commonly used CX gate as an example.
#
# <img src="../tutorial-ja/img/007/007_0.png" width="60%">
#
# As you can see, when the first bit is $|1\rangle$, the second bit is mapped to the X gate.
# Other control gates are the same.
# ### CRx, CRy, CRz,
# CRx, CRy, CRz gates can be implemented by the following.
#
# | CRx | CRy | CRz |
# |:-:|:-:|:-:|
# |`crx(θ)`|`cry(θ)`|`crz(θ)`|
#
# $\theta$ is the same as RX, RY and RZ gate.
# ###### CRx gate
Circuit().crx(math.pi/2)[0,1].m[:].run(shots=100)
# ###### CRy gate
Circuit().cry(math.pi/2)[0,1].m[:].run(shots=1000)
# ###### CRz gate
Circuit().crz(math.pi/2)[0,1].m[:].run(shots=1000)
# ### CU
# CU gates can be implemented by the following.
#
# | CU |
# |:-:|
# |`cu(θ,φ,λ,γ)`|
#
# $\theta, \phi, \lambda, \gamma$ is the same as U gate.
Circuit().cu(math.pi/2, 0, math.pi, -math.pi/2)[0,1].m[:].run(shots=100)
# ## Swap Gate
# Swap gate is the gate to exchange value of 2qubts. By using CX gate we can realize swap gate.
# ## Circuit
# By using 3CX gate we can use swap gate.
#the fist X gate is data input to check if the swap gate works
Circuit().x[0].cx[0,1].cx[1,0].cx[0,1].m[:].run(shots=1)
# By applying 3 CX gate continuously you can use swap gate. The first qubit of 1 is exchanged and we finally get 0 as 0th qubit and 1 as 1st qubit
# ## Ising Gate
# An ising gate is a gate that rotates two qubits at the same time.
# ### Rxx, Ryy, Rzz
# Rxx,Ryy,Rzz gates can be implemented by the following.
#
# | Rxx | Ryy | Rzz |
# |:-:|:-:|:-:|
# |`rxx(θ)`|`ryy(θ)`|`rzz(θ)`|
# ###### RXX gate
Circuit(2).rxx(math.pi/2)[0,1].m[:].run(shots=100)
# ###### RYY gate
Circuit(2).ryy(math.pi/2)[0,1].m[:].run(shots=100)
# ###### RZZ gate
Circuit().rzz(math.pi/2)[0,1].m[:].run(shots=100)
# ## Rxx, Ryy, Rzz gates
# The rotation of one qubit was represented by two states, 0 and 1.
# The rotation of two qubits is represented by four states, 00, 01, 10, and 11.
# #### Rxx gate
#
# The matrix looks like the following.
#
# $$
# Rxx(\theta) = \left(\begin{array}{cccc}
# \cos\theta &0&0&-i\sin\theta\\
# 0& \cos\theta &-i\sin\theta&0\\
# 0&-i\sin\theta& \cos\theta &0\\
# -i\sin\theta&0&0& \cos\theta
# \end{array}\right)\\
# $$
#
# And the action on each bit is:
#
# <img src="../tutorial-ja/img/ising_0.png" width="55%">
# #### Ryy gate
#
# The matrix looks like the following.
#
# $$
# Ryy(\theta) = \left(\begin{array}{cccc}
# \cos\theta &0&0&i\sin\theta\\
# 0& \cos\theta &-i\sin\theta&0\\
# 0&-i\sin\theta& \cos\theta &0\\
# i\sin\theta&0&0& \cos\theta
# \end{array}\right)
# $$
#
# And the action on each bit is:
#
# <img src="../tutorial-ja/img/ising_1.png" width="55%">
# #### Rzz gate
#
# The matrix looks like the following.
#
# $$
# Rzz(\theta) = \left(\begin{array}{cccc}
# e^{\frac{\theta}{2}i} &0&0&0\\
# 0& e^{-\frac{\theta}{2}i} &0&0\\
# 0&0& e^{-\frac{\theta}{2}i} &0\\
# 0&0&0& e^{\frac{\theta}{2}i}
# \end{array}\right)
# $$
#
# And the action on each bit is:
#
# <img src="../tutorial-ja/img/ising_2.png" width="35%">
| tutorial/006_2gate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: heom
# language: python
# name: heom
# ---
# # A *quantum_HEOM* tutorial
# ## Part 1: Quick Run
# Run the following cell (``shift`` + ``enter``) to quickly plot the time evolution of a QuantumSystem. You can play around with the settings and re-run the cell to see changes in the dynamics. Alternatively, see the tutorial in _Part 2_ for a breakdown of the settings options.
# +
# %matplotlib inline
from quantum_system import QuantumSystem
# Define the sites and settings for the quantum system.
sites = 5
settings = {'atomic_units': True,
'interaction_model': 'nearest neighbour cyclic',
'dynamics_model': 'dephasing lindblad',
'decay_rate': 0.1,
'time_interval': 0.1,
'timesteps': 300}
# Initialise QuantumSystem with the defined sites and settings
q = QuantumSystem(sites=sites, **settings)
# 3D complex space-time plot of dynamics.
q.plot_time_evolution(view_3d=True, elements='all')
# -
# ## Part 2: Tutorial
# ### Imports
# %matplotlib inline
from quantum_system import QuantumSystem
# ### Set up the QuantumSystem
# Define the number of sites that are present in the quantum system:
sites = 3
# Now define the some settings that dictate how the sites interact, and how the dynamics of the density matrix are defined. After each colon ``:`` fill in your choices. The options are as follows:
#
# * ``'atomic_units':`` choose from ``True`` or ``False``
# * ``'interaction_model':`` choose from ``'nearest neighbour cyclic'`` or ``'nearest neighbour linear'``
# * ``'dynamics_model':`` choose from ``'simple'`` or ``'dephasing lindblad'``
# * ``'decay_rate':`` choose a non-negative float value, i.e. ``0.15``
# * ``'time_interval':`` choose a positive float value, i.e. ``0.1``
# * ``'timesteps':`` choose a positive integer value, i.e ``3000``
settings = {'atomic_units': True,
'interaction_model': 'nearest neighbour cyclic',
'dynamics_model': 'dephasing lindblad',
'decay_rate': 0.15,
'time_interval': 0.1,
'timesteps': 300}
# Now run the next cell to initialise a QuantumSystem object with your chosen number of sites and settings:
q = QuantumSystem(sites=sites, **settings)
# ### Plot the time evolution
# The time evolution of the system under the interactions and dynamics you have chosen can now be plotted. Run the following cell to do so. The options for the 2 arguments can be changed as follows:
#
# * ``view_3d`` choose from ``True`` (makes a complex space-time plot) or ``False`` (plots the evolution of just the real amplitudes of the diagonal elements of the density matrix).
# * ``elements`` can be specified as a list of the elements of the density matrix you wish to plot, i.e. ``elements=['11', '12', '21', '22']`` or as a string of one of the following values; ``'all'``, ``'diagonals'``, or ``'off-diagonals'``.
# * **Note**: choosing ``view_3d=False`` makes the string options ``'all'`` and ``'diagonals'`` have the same effect (as only diagonals are plotted), and ``'off-diagonals'`` as plotting none of the elements. You can still specify an array to choose the diagonals you wish to plot.
q.plot_time_evolution(view_3d=True, elements='all')
# It is possible to redefine some settings and replot the time evolution. This can be done by reassigning the attribute for the setting you want to change, and recalling the ``plot_time_evolution()`` method. For example, one could change the ``interaction_model`` to a ``'nearest neighbour linear'`` one and then re-plot in the following way:
q.interaction_model = 'nearest neighbour linear'
q.plot_time_evolution(view_3d=True, elements='all')
| quantum_heom/tutorial_define_system_plot_evolution.ipynb |
# ---
# title: "Iterables with enumerate() and zip()"
# date: 2020-04-12T14:41:32+02:00
# author: "<NAME>"
# type: technical_note
# draft: false
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Using `enumerate()`
# `enumerate()` returns an enumerate object that produces a sequence of tuples, and each of the tuples is an index-value pair.
# +
mutants = ['<NAME>',
'<NAME>',
'<NAME>',
'<NAME>',
'<NAME>']
# Create a list of tuples
mutant_list = list(enumerate(mutants))
print(mutant_list)
# Unpack and print the tuple pairs
for index1,value1 in enumerate(mutants):
print(index1, value1)
# Change the start index
for index2,value2 in enumerate(mutants, start=1):
print(index2, value2)
# -
# ### Using `zip()`
# takes any number of iterables and returns a zip object that is an iterator of tuples. If you wanted to print the values of a zip object, you can convert it into a list and then print it. Printing just a zip object will not return the values unless you unpack it first.
mutants=['<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>']
aliases=['prof x', 'iceman', 'nightcrawler', 'magneto', 'shadowcat']
powers=['telepathy', 'thermokinesis', 'teleportation', 'magnetokinesis', 'intangibility']
# +
# Create a list of tuples
mutant_data = list(zip(mutants,aliases,powers))
print(mutant_data)
# Create a zip object using the three lists
mutant_zip = zip(mutants,aliases,powers)
print(mutant_zip)
# Unpack the zip object and print the tuple values
for value1,value2,value3 in mutant_zip:
print(value1, value2, value3)
# -
# ### Using * and zip to 'unzip'
# `*` unpacks an iterable such as a list or a tuple into positional arguments in a function call.
# +
# Create a zip object from mutants and powers
z1 = zip(mutants,powers)
# Print the tuples in z1 by unpacking with *
print(*z1)
# Re-create a zip object from mutants and powers, as the print(*) exhausted all elements of z1
z1 = zip(mutants,powers)
# 'Unzip' the tuples in z1 by unpacking with * and zip(): result1, result2
result1, result2 = zip(*z1)
# Check if unpacked tuples are equivalent to original tuples
print(result1 == mutants)
print(result2 == powers)
| courses/datacamp/notes/python/basics/iterators_enum_zip.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
parts = ['C:\\Users\\kaush\\Downloads\\patterns-part1.csv', 'C:\\Users\\kaush\\Downloads\\patterns-part2.csv', 'C:\\Users\\kaush\\Downloads\\patterns-part3.csv', 'C:\\Users\\kaush\\Downloads\\patterns-part4.csv']
new_path = 'C:\\Users\\kaush\\Downloads\\october_safegraph.csv'
monthly_patterns_raw = pd.concat(map(pd.read_csv, parts)).reset_index(drop=True)
monthly_patterns_raw.head()
monthly_patterns_raw = monthly_patterns_raw[monthly_patterns_raw['region'] == 'CA']
monthly_patterns_raw['county_tract'] = (monthly_patterns_raw['poi_cbg'] / 1000000).astype(int)
monthly_patterns_raw = monthly_patterns_raw[monthly_patterns_raw['county_tract'] == 60371]
monthly_patterns_raw.to_csv(new_path, encoding='utf-8', index=False)
| code/SafeGraphPreprocessing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:bpm3]
# language: python
# name: conda-env-bpm3-py
# ---
import pandas as pd
from sklearn.metrics import roc_auc_score, brier_score_loss
import os
import glob
import csv
import numpy as np
import re
def calculate_stability(group, col="predicted"):
group["diff"] = abs(group[col].shift(-1) - group[col])
return(group["diff"].mean(skipna=True))
def extract_event_nr(s):
m = re.match(r'.*_(\d{1,2})$', s)
if m:
return int(m.group(1))
else:
return 1
def extract_case_id(s):
m = re.match(r'(.*)_\d{1,2}$', s)
if m:
return m.group(1)
else:
return s
# # Original
results_dir = "results_stability"
for filename in glob.glob("results_detailed/*test*"):
print(filename)
dt_results = pd.read_csv(filename, sep=";")
dt_results.case_id = dt_results.case_id.astype(str)
if "lstm" not in filename:
if "single" in filename:
dt_results["nr_events"] = dt_results.case_id.apply(extract_event_nr)
dt_results["case_id"] = dt_results.case_id.apply(extract_case_id)
dataset_name = dt_results.dataset.iloc[0]
if "params" in dt_results.columns:
method_name = dt_results.params.iloc[0]
else:
method_name = dt_results.method.iloc[0]
cls_method = dt_results.cls.iloc[0]
with open(os.path.join(results_dir, "results_auc_stability_%s_%s_%s.csv" % (dataset_name, method_name, cls_method)), 'w') as csvfile:
spamwriter = csv.writer(csvfile, delimiter=';', quoting=csv.QUOTE_NONE)
spamwriter.writerow(["dataset", "method", "cls", "nr_events", "metric", "score"])
for nr_events, group in dt_results.groupby("nr_events"):
auc = np.nan if len(set(group.actual)) < 2 else roc_auc_score(group.actual, group.predicted)
spamwriter.writerow([dataset_name, method_name, cls_method, nr_events, "auc", auc])
stab_by_case = dt_results.groupby("case_id").apply(calculate_stability)
spamwriter.writerow([dataset_name, method_name, cls_method, -1, "stability", 1 - stab_by_case.mean()])
# # Smoothing
# +
import pickle
with open("n_test_cases.pickle", "rb") as fin:
test_cases_dict = pickle.load(fin)
df_test_cases = pd.DataFrame(test_cases_dict)
df_test_cases = df_test_cases.stack().reset_index()
df_test_cases.columns = ["nr_events", "dataset", "n_cases"]
# -
results_dir = "results_stability_smoothed"
for filename in glob.glob("results_detailed/*test*"):
print(filename)
dt_results = pd.read_csv(filename, sep=";")
dt_results.case_id = dt_results.case_id.astype(str)
if "lstm" not in filename:
if "single" in filename:
dt_results["nr_events"] = dt_results.case_id.apply(extract_event_nr)
dt_results["case_id"] = dt_results.case_id.apply(extract_case_id)
dataset_name = dt_results.dataset.iloc[0]
if "params" in dt_results.columns:
method_name = dt_results.params.iloc[0]
dt_results = dt_results.drop(["params"], axis=1)
else:
method_name = dt_results.method.iloc[0]
dt_results = dt_results.drop(["method"], axis=1)
cls_method = dt_results.cls .iloc[0]
dt_results = dt_results.drop(["dataset", "cls"], axis=1)
dt_results.nr_events = dt_results.nr_events.astype(int)
betas = [0, 0.1, 0.25, 0.5, 0.75, 0.9]
smoothed_preds = dt_results[dt_results.nr_events==1]
for beta in betas:
smoothed_preds["smoothed_pred_%s" % beta] = smoothed_preds["predicted"]
for nr_events in range(2, dt_results.nr_events.max()+1):
tmp_merged = pd.merge(dt_results[dt_results.nr_events==nr_events], smoothed_preds[smoothed_preds.nr_events==(nr_events-1)].drop(["predicted", "nr_events"], axis=1), on=["case_id", "actual"])
for beta in betas:
tmp_merged["smoothed_pred_%s" % beta] = beta * tmp_merged["smoothed_pred_%s" % beta] + (1-beta) * tmp_merged.predicted
smoothed_preds = pd.concat([smoothed_preds, tmp_merged], axis=0)
with open(os.path.join(results_dir, "results_auc_stability_%s_%s_%s.csv" % (dataset_name, method_name, cls_method)), 'w') as csvfile:
spamwriter = csv.writer(csvfile, delimiter=';', quoting=csv.QUOTE_NONE)
spamwriter.writerow(["dataset", "method", "cls", "beta", "metric", "score"])
for beta in betas:
aucs = []
for nr_events, group in smoothed_preds.groupby("nr_events"):
auc = np.nan if len(set(group.actual)) < 2 else roc_auc_score(group.actual, group["smoothed_pred_%s" % beta])
aucs.append((auc, dataset_name, nr_events))
dt_aucs = pd.DataFrame(aucs)
dt_aucs.columns = ["score", "dataset", "nr_events"]
dt_aucs = pd.merge(dt_aucs, df_test_cases, on=["dataset", "nr_events"])
dt_aucs["score"].fillna(0, inplace=True)
auc = np.average(dt_aucs["score"], weights=dt_aucs["n_cases"])
spamwriter.writerow([dataset_name, method_name, cls_method, beta, "auc", auc])
stab_by_case = smoothed_preds.groupby("case_id").apply(calculate_stability, col="smoothed_pred_%s" % beta)
spamwriter.writerow([dataset_name, method_name, cls_method, beta, "stability", 1 - stab_by_case.mean()])
# # Brier scores
results_dir = "results_stability_brier"
for filename in glob.glob("results_detailed/*test*"):
print(filename)
dt_results = pd.read_csv(filename, sep=";")
dt_results.case_id = dt_results.case_id.astype(str)
if "lstm" not in filename:
if "sepsis" in filename:
dt_results.case_id = dt_results.case_id.str.replace("missing_caseid", "missing")
max_underscores_in_caseid = dt_results.case_id.apply(lambda x: len(x.split("_"))).max()
if "single" in filename:
dt_results["nr_events"] = dt_results.case_id.apply(lambda x: 1 if len(x.split("_")) < max_underscores_in_caseid else x.split("_")[-1])
dt_results["case_id"] = dt_results.case_id.apply(lambda x: x if len(x.split("_")) < max_underscores_in_caseid else x.split("_")[0])
dataset_name = dt_results.dataset.iloc[0]
if "params" in dt_results.columns:
method_name = dt_results.params.iloc[0]
else:
method_name = dt_results.method.iloc[0]
cls_method = dt_results.cls.iloc[0]
with open(os.path.join(results_dir, "results_auc_stability_%s_%s_%s.csv" % (dataset_name, method_name, cls_method)), 'w') as csvfile:
spamwriter = csv.writer(csvfile, delimiter=';', quoting=csv.QUOTE_NONE)
spamwriter.writerow(["dataset", "method", "cls", "nr_events", "metric", "score"])
for nr_events, group in dt_results.groupby("nr_events"):
brier = brier_score_loss(group.actual, group.predicted)
spamwriter.writerow([dataset_name, method_name, cls_method, nr_events, "brier", brier])
brier = brier_score_loss(dt_results.actual, dt_results.predicted)
spamwriter.writerow([dataset_name, method_name, cls_method, -1, "brier", brier])
| evaluate_accuracy_stability.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
Operation Sorted List Hash Table Binary Search Tree AVL Tree
put/insert O(n) O(1) O(log2n) O(log2n)
get/access O(log2n) O(1) O(log2n) O(log2n)
in/search O(log2n) O(1) O(log2n) O(log2n)
del/remove O(n) O(1) O(log2n) O(log2n)
# +
""" Binary Tree
The total number of nodes in a perfectly balanced binary tree is (2^(h+1)) −1,
where h represents the height of the tree.
Ref: GeeksforGeeks + Tushar
"""
from collections import deque
class Node:
""" A node structure """
def __init__(self, val):
""" A utility function to create a new node """
self.val = val
self.left = None
self.right = None
def insertLeft(node,newNodeValue): # Time (O(n))
""" function to insert element in a given node's left child in binary tree """
newNode = Node(newNodeValue)
if node.left == None:
node.left = newNode
else:
newNode.left = node.left
node.left = newNode
def insertRight(node,newNodeValue):
""" function to insert element in a given node's right child in binary tree """
newNode = Node(newNodeValue)
if node.right == None:
node.right = newNode
else:
newNode.right = node.right
node.right = newNode
def getRightChild(node):
return node.right
def getLeftChild(node):
return node.left
def setRootVal(node,value):
node.val = value
def getRootVal(node):
return node.val
def insert(root, key): # Time: O(n), Space: O(n)
"""
Given a binary tree and a key, insert the key into the binary tree
at first position available in level order (BFS way). The idea is to do
iterative level order traversal of the given tree using queue.
If we find a node whose left child is empty, we make new key as
left child of the node. Else if we find a node whose right child
is empty, we make new key as right child. We keep traversing the
tree until we find a node whose either left or right is empty."""
queue = deque()
queue.append(root)
# Do level order (BFS) traversal until we find an empty place.
while (len(queue) > 0):
root = queue.popleft()
if (root.left == None):
root.left = Node(key)
return
else:
queue.append(root.left)
if (root.right == None):
root.right = Node(key)
return
else:
queue.append(root.right)
def delete(root, key): # Time: O(n), Space: O(n)
"""
Function to delete element in binary tree
Given a binary tree, delete a node from it by making sure that tree shrinks
from the bottom (i.e. the deleted node is replaced by bottom most and
rightmost node). This different from BST deletion. Here we do not have
any order among elements, so we replace with last element.
Algorithm:
1. Starting at root, find the deepest and rightmost node in
binary tree and node which we want to delete.
2. Replace the deepest rightmost node’s data with node to be deleted.
3. Then delete the deepest rightmost node.
"""
if root is None:
return
queue = deque()
queue.append(root)
current = None
nodeToBeDeleted = None
while len(queue)>0: # Do level order traversal to find deepest node (current)
current = queue.popleft() # and node to be deleted (nodeToBeDeleted)
if (current.val == key): # node with the given key found
nodeToBeDeleted = current
if current.left:
queue.append(current.left)
if current.right:
queue.append(current.right)
if nodeToBeDeleted == None: # key not found
return
nodeToBeDeleted.val = current.val # current would be the deepest (usually rightmost) leaf node
deleteDeepestNode(root, current)
def deleteDeepestNode(root, node): # Time: O(n), Space: O(n)
"""
Helper function for delete(): deletes the given deepest
node (node) in binary tree. """
queue = deque()
queue.append(root)
while (len(queue)>0): # Do level order traversal until last node
temp = queue.popleft()
if temp.right:
if temp.right == node:
temp.right = None
return
else:
queue.append(temp.right)
if temp.left:
if temp.left == node:
temp.left = None
return
else:
queue.append(temp.left)
def printLevelOrder(root): # Time: O(n), Space: O(n)
"""
BFS of Binary Tree : Iterative Method to print the height of binary tree
Ref: https://www.youtube.com/watch?v=9PHkM0Jri_4&list=PLrmLmBdmIlpv_jNDXtJGYTPNQ2L1gdHxu&index=9
"""
if root is None: # Base Case
return
queue = deque() # Create an empty queue for level order traversal
queue.append(root) # Enqueue Root and initialize height
while(len(queue) > 0):
node = queue.popleft()
print(node.val, end=' '), # Print front of queue and remove it from queue
if node.left:
queue.append(node.left) # Enqueue left child
if node.right:
queue.append(node.right)# Enqueue right child
print()
def printRevereseLevelOrder(root): # Time: O(n), Space: O(n)
"""BFS of Binary Tree : Iterative Method to print nodes on each level from bottom up (in reverse order)
of a binary tree.
Ref: https://www.youtube.com/watch?v=D2bIbWGgvzI&list=PLrmLmBdmIlpv_jNDXtJGYTPNQ2L1gdHxu&index=14
"""
if root is None: # Base Case
return
stack = deque()
queue = deque()
queue.append(root)
while(len(queue) > 0):
node = queue.popleft()
if node.right:
queue.append(node.right) # Enqueue right child
if node.left:
queue.append(node.left) # Enqueue left child
stack.append(node) # Push on to stack
while len(stack) > 0:
current = stack.pop() # Pop from stack
print(current.val, end=' '),
print()
def printSpiralOrder(root): # Time: O(n), Space: O(n)
"""BFS of Binary Tree : Iterative Method to print nodes on each level of a binary tree top to bottom
in a SPIRAL order.
Ref: https://www.youtube.com/watch?v=vjt5Y6-1KsQ&list=PLrmLmBdmIlpv_jNDXtJGYTPNQ2L1gdHxu&index=15
"""
if root is None: # Base Case
return
stack1 = deque()
stack2 = deque()
stack1.append(root)
while (len(stack1) > 0) or (len(stack2) > 0):
while len(stack1) > 0:
root = stack1.pop()
if root.left:
stack2.append(root.left)
if root.right:
stack2.append(root.right)
print(root.val, end=" ")
while len(stack2) > 0:
root = stack2.pop()
if root.right:
stack1.append(root.right)
if root.left:
stack1.append(root.left)
print(root.val, end=" ")
print()
def printPreorder_recursive(root): # Time: O(n), Space: O(1)
"""
DFS: A function to do preorder tree traversal
Time: O(n), Space: O(1)"""
if root:
print(root.val, end=' '), # First print the data of node
printPreorder_recursive(root.left) # Then recur on left child
printPreorder_recursive(root.right) # Finally recur on right child
def printInorder_recursive(root): # Time: O(n), Space: O(1)
"""
DFS: A function to do inorder tree traversal
Time: O(n), Space: O(1)"""
if root:
printInorder_recursive(root.left) # First recur on left child
print(root.val, end=' '), # then print the data of node
printInorder_recursive(root.right) # now recur on right child
def printPostorder_recursive(root): # Time: O(n), Space: O(1)
"""
DFS: A function to do postorder tree traversal
Time: O(n), Space: O(1)"""
if root:
printPostorder_recursive(root.left) # First recur on left child
printPostorder_recursive(root.right) # the recur on right child
print(root.val, end=' '), # now print the data of node
def printPreorder_iterative(root): # Time: O(n), Space: O(n)
"""
DFS: A function to do preorder tree traversal in iterative fashion using stack. Time: O(n), Space: O(n)
Ref: https://www.youtube.com/watch?v=elQcrJrfObg&list=PLrmLmBdmIlpv_jNDXtJGYTPNQ2L1gdHxu&index=11
"""
if root == None:
return
stack = deque()
stack.append(root)
while len(stack) > 0:
root = stack.pop()
print(root.val, end= " ")
if root.right:
stack.append(root.right)
if root.left:
stack.append(root.left)
def printInorder_iterative(root): # # Time: O(n), Space: O(n)
"""
DFS: A function to do inorder tree traversal in iterative fashion using stack.
Time: O(n), Space: O(n)
Ref: https://www.youtube.com/watch?v=nzmtCFNae9k&list=PLrmLmBdmIlpv_jNDXtJGYTPNQ2L1gdHxu&index=12
"""
if root == None:
return
stack = deque()
continueTraversal = True
while(continueTraversal):
if root != None:
stack.append(root)
root = root.left
else:
if len(stack) == 0:
continueTraversal = False
root = stack.pop()
print(root.val, end=" ")
root = root.right
def printPostorder_iterative(root): # Time: O(n), Space: O(n)
"""
DFS: A function to do postorder tree traversal in iterative fashion using two stacks.
Ref: https://www.youtube.com/watch?v=qT65HltK2uE&list=PLrmLmBdmIlpv_jNDXtJGYTPNQ2L1gdHxu&index=10
"""
if root == None:
return
stack1 = deque()
stack2 = deque()
stack1.append(root)
while len(stack1) > 0:
root = stack1.pop()
s2.append(root)
if root.left != None:
stack1.append(root.left)
if root.right != None:
stack1.append(root.right)
while len(stack2) > 0:
root = stack2.pop()
print(root.val, end=" ")
def search(root, find_val): # Time: O(n), Space: O(1)
"""Return True if the value
is in the tree, return
False otherwise."""
return preorder_search(root, find_val)
def preorder_search(root, find_val): # Time: O(n), Space: O(1)
"""Helper method - use this to create a
recursive search solution."""
if root:
if root.val == find_val:
return True
else:
return preorder_search(root.left, find_val) or preorder_search(root.right, find_val)
return False
def size(node): # Time: O(n), Space: O(1)
"""
#i.e., how many total nodes in a tree
The function Compute the "size" of a tree. Size is the
number of nodes in a tree.
Ref: https://www.youtube.com/watch?v=NA8B84DZYSA&list=PLrmLmBdmIlpv_jNDXtJGYTPNQ2L1gdHxu&index=5
"""
if node is None: # Base Case : Tree is empty
return 0
leftSize = size(node.left)
rightSize = size(node.right)
return 1 + leftSize + rightSize
def height(node): # Time: O(n), Space: O(1)
"""
The function Compute the "height" of a tree. Height is the
number of nodes along the longest path from the root node
down to the farthest leaf node.
Ref: https://www.youtube.com/watch?v=_SiwrPXG9-g&list=PLrmLmBdmIlpv_jNDXtJGYTPNQ2L1gdHxu&index=6
"""
if node is None: # Base Case : Tree is empty
return 0
# If tree is not empty then height = 1 + max of left
# height and right heights
return 1 + max(height(node.left) , height(node.right))
def diameter(root): # Time: O(n), Space: O(1)
"""
Function to get the diamtere of a binary tree.
The diameter of a Tree is the largest of the following quantities:
-- the diameter of Tree’s left subtree
-- the diameter of Tree’s right subtree
-- the longest path between leaves that goes through the root of Tree
(This can be computed from the heights of the subtrees of Tree).
"""
if root is None: # Base case
return 0
lheight = height(root.left) # Height of left subtree
rheight = height(root.right) # Height of right subtree
longest_path = 1 + lheight + rheight # The longest path between leaves that
# goes thru the root of the tree
ldiameter = diameter(root.left) # Diameter of left subtree
rdiameter = diameter(root.right) # Diameter of right subtree
return max(longest_path, max(ldiameter, rdiameter))
def detectSameTree(root1, root2): # Time: O(n), Space: O(1)
""" Return True/False whether the given trees are same.
Ref: https://www.youtube.com/watch?v=ySDDslG8wws&list=PLrmLmBdmIlpv_jNDXtJGYTPNQ2L1gdHxu&index=4
"""
if root1 == None:
return root2 == None
if root2 == None:
return root1 == None
if root1.val == root2.val:
return detectSameTree(root1.left, root2.left) and detectSameTree(root1.right, root2.right)
else:
return False
def rootToLeafSumBinaryTree(root, sum_value, path): # Time: O(n), Space: O(n)
"""
Ref: https://www.youtube.com/watch?v=Jg4E4KZstFE&list=PLrmLmBdmIlpv_jNDXtJGYTPNQ2L1gdHxu&index=7
Time: O(n), Space: O(n) (for the path variable)
"""
if root == None:
return False
if root.left is None and root.right is None: # leaf node
if root.val == sum_value:
path.append(root.val)
return True
else:
return False
if rootToLeafSumBinaryTree(root.left, sum_value - root.val, path):
path.append(root.left.val)
return True
if rootToLeafSumBinaryTree(root.right, sum_value - root.val, path):
path.append(root.right.val)
return True
return False
def findAncestors(root,k): # Time: O(n), Space: O(n) (if skewed tree)
""" Return ancestors (list) of a given node in a tree. """
def findAncestors_rec(root, k, ancestor_list):
if root == None:
return False
elif root.val == k:
return True
elif (findAncestors_rec(root.left, k, ancestor_list) or findAncestors_rec(root.right, k, ancestor_list)):
ancestor_list.append(root.val)
return True
return False
ancestor_list = []
findAncestors_rec(root, k, ancestor_list)
return ancestor_list
def lca(root, node1, node2): # Time: O(n), Space O(1):
"""Lowest common ancestor of a given pair of nodes in a binary tree
Ref: https://www.youtube.com/watch?v=13m9ZCB8gjw&list=PLrmLmBdmIlpv_jNDXtJGYTPNQ2L1gdHxu&index=17
"""
if root == None:
return None
if root == node1 or root == node2:
return root
left = lca(root.left, node1, node2)
right = lca(root.right, node1, node2)
if left != None and right != None:
return root
if left == None and right == None:
return None
if left != None:
return left
else:
return right
def isBST_iterative(root): # Time: O(n) , Space: 0(n)
"""
Ref code: https://github.com/mission-peace/interview/blob/master/src/com/interview/tree/IsBST.java
"""
if root == None:
return True
stack = []
node = root
prev = float("-inf")
current = None
while (True):
if (node != None):
stack.append(node)
node = node.left
else:
if len(stack) == 0:
return False
node = stack.pop()
current = node.val
if current < prev:
return False
prev = current
node = node.right
return True
def isBST_recursive(root): # Time: O(n) , Space: O(1)
"""
Ref: https://www.youtube.com/watch?v=MILxfAbIhrE&list=PLrmLmBdmIlpv_jNDXtJGYTPNQ2L1gdHxu&index=8
"""
def isBST_rec_Helper(root, minValue, maxValue):
if root == None:
return True
if root.val <= minValue or root.val >= maxValue:
return False
return isBST_rec_Helper(root.left, minValue, root.val) and isBST_rec_Helper(root.right, root.val, maxValue)
minValue = float("-inf")
maxValue = float("inf")
return isBST_rec_Helper(root, minValue, maxValue)
def largestBST(root): # Time: O(n^2), Space: O(1)
"""
Consider a skewed tree for worst time scenario.
# Returns size of the largest BST subtree in a Binary Tree (inefficient version).
Ref: https://www.geeksforgeeks.org/find-the-largest-subtree-in-a-tree-that-is-also-a-bst/"""
if isBST_recursive(root):
return size(root)
else:
return max(largestBST(root.left), largestBST(root.right))
def largestBST_Optimized(root): # Time: O(n), Space: O(n)
"""
Returns size of the largest BST subtree in a Binary Tree (efficient version).
Ref vid: https://www.youtube.com/watch?v=4fiDs7CCxkc&list=PLrmLmBdmIlpv_jNDXtJGYTPNQ2L1gdHxu&index=19
Ref Code: https://github.com/mission-peace/interview/blob/master/python/tree/largest_bst_in_binary_tree.py
"""
class TreeStatus:
def __init__(self):
self.isBST = True
self.size = 0
self.min = float("inf")
self.max = float("-inf")
def largest(root):
if root is None:
return TreeStatus()
leftTreeStatus = largest(root.left)
rightTreeStatus = largest(root.right)
tree_stat = TreeStatus()
if (leftTreeStatus.isBST == False or rightTreeStatus.isBST == False
or leftTreeStatus.max > root.val or rightTreeStatus.min <= root.val):
tree_stat.isBST = False
tree_stat.size = max(leftTreeStatus.size, rightTreeStatus.size)
return tree_stat
tree_stat.isBST = True
tree_stat.size = 1 + leftTreeStatus.size + rightTreeStatus.size
tree_stat.min = leftTreeStatus.min if root.left is not None else root.val
tree_stat.max = rightTreeStatus.max if root.right is not None else root.val
return tree_stat
tree_status = largest(root)
return tree_status.size
def find_nodes_k_distance_from_root(root, k): # Time: O(n), Space: O(n)
""" Find Nodes at 'k' distance from the Root. """
def find_k_recursive(root, k, res):
if root is None:
return
elif k == 0:
res.append(root.val)
else:
find_k_recursive(root.left, k-1, res)
find_k_recursive(root.right, k-1, res)
res = []
find_k_recursive(root, k, res)
return res
def binaryTreeToCircularLinkedList(root): # Time: (n) time, Space: O(1)
"""
Python Program that converts (In-Place) a tree to a Circular Doubly LinkedList
and then returns the head of the resulting LinkedList.
Red vid: https://www.youtube.com/watch?v=Dte6EF1nHNo
Ref: https://www.geeksforgeeks.org/convert-a-binary-tree-to-a-circular-doubly-link-list/
"""
def concatenate(leftList, rightList):
"""
Helper function to binaryTreeToCircularLinkedList() function.
A function that appends rightList at the end of leftList.
Please note that both rightList and leftList are
circular doubly linked list themselves.
"""
# If either of the list is empty then return the other list
if (leftList == None): return rightList
if (rightList == None): return leftList
leftEnd = leftList.left # Store the End Node of leftList
rightEnd = rightList.left # Store the End Node of rightList
# Connect the End node of LeftList
# with the first Node of the rightList
leftEnd.right = rightList
rightList.left = leftEnd
# Connect Left of first node points to
# the End node in the list
leftList.left = rightEnd
# Connect Right of End node refers to
# the first node of the List
rightEnd.right = leftList
return leftList
if (root == None):
return None
# Recursively convert left and right subtrees
leftList = binaryTreeToCircularLinkedList(root.left)
rightList = binaryTreeToCircularLinkedList(root.right)
# Make a circular linked list of single node (or root). To do so,
# make the right and left pointers of this node point to itself
root.left = root
root.right = root
# Step 1 (concatenate the left list with the list with single node, i.e., current node)
# Step 2 (concatenate the returned list with the right List)
root = concatenate(leftList, root)
root = concatenate(root, rightList)
return root
def displayCircularLinkedList(head):
"""Helper function to binaryTreeToCircularLinkedList() function.
Functon that displays Circular Link List
"""
print("Circular Linked List is :")
if head is None:
return
current = head
first = 1
while (head != current or first == 1):
print(current.val, end = " ")
current = current.right
first = 0
print("")
def test_connect_same_level_siblings():
""" Given a binary tree, connect its siblings (i.e., root.next) at each level,
last sibling in each lebvel points to Null.
https://www.educative.io/collection/page/5642554087309312/5679846214598656/250002
"""
def connect_same_level_siblings(root): # O(n), O(1)
if root == None:
return
root.next = None
while root != None:
root = connect_next_level(root)
def connect_next_level(root): # O(n), O(1)
current = root
next_level_head = None
prev = None
while current != None:
if current.left != None and current.right != None:
if next_level_head == None:
next_level_head = current.left
current.left.next = current.right
if prev != None:
prev.next = current.left
prev = current.right
elif current.left != None:
if next_level_head == None:
next_level_head = current.left
if prev != None:
prev.next = current.left
prev = current.left
elif current.right != None:
if next_level_head == None:
next_level_head = current.right
if prev != None:
prev.next = current.right
prev = current.right
current = current.next
if prev != None:
prev.next = None
return next_level_head # end of method
def get_level_order_using_next(root): # O(n), O(n) - BFS
output = []
while root != None:
head = root
next_head = None
while head != None:
output.append(head.val)
if next_head == None:
if head.left != None:
next_head = head.left
else:
next_head = head.right
head = head.next
root = next_head
return output
root = Node(100)
for node in [25, 350, 15, 75, 300, 400, 10, 50, 200, 325, 375]:
insert(root, node)
print("print given tree: ")
print_tree_2D(root)
connect_same_level_siblings(root)
v2 = get_level_order_using_next(root)
print("print level order by using next pointer: ", v2)
print("root.next: ", root.next) #None
print("root.left.next.val: ", root.left.next.val) #350
print("root.left.right.next.val: ", root.left.right.next.val) #300
print("root.right.left.next.val: ", root.right.left.next.val) #400
print("root.right.right.next: " , root.right.right.next) #None
"""=================================== Parse Tree for Numerical Equations ==================================="""
def buildParsparseTree(expression_string):
if len(expression_string) == 0: # input empty
return None
expression_list = expression_string.split()
queue = deque()
parseTree = Node('')
queue.append(parseTree) # enqueue
currentTree = parseTree
for i in expression_list:
if i == '(':
# insert left of CurrentTree
newNode = Node('')
if currentTree.left is None:
currentTree.left = newNode
else:
newNode.left = currentTree.left
currentTree.left = newNode
queue.append(currentTree) # enqueue
currentTree = currentTree.left
elif i not in ['+', '-', '*', '/', ')']:
currentTree.val = int(i)
parent = queue.popleft() # dequeue
currentTree = parent
elif i in ['+', '-', '*', '/']:
currentTree.val = i
# insert right of CurrentTree
newNode = Node('')
if currentTree.right is None:
currentTree.right = newNode
else:
newNode.right = currentTree.right
currentTree.right = newNode
queue.append(currentTree) # enqueue
currentTree = currentTree.right
elif i == ')':
currentTree = queue.popleft() # dequeue
else:
raise ValueError
return parseTree
def evaluate(parseTree):
""" Evaluate the numerical expression in the parse tree (recursive)"""
if parseTree is None: # input None/empty
return None
import operator
operators = { '+': operator.add,
'-': operator.sub,
'*': operator.mul,
'/': operator.truediv
}
if parseTree.left and parseTree.right:
fn = operators[parseTree.val]
return fn(evaluate(parseTree.left), evaluate(parseTree.right) )
else:
return parseTree.val
def printParenthesizedTree(parseTree):
value = ""
if parseTree:
value = '(' + printParenthesizedTree(parseTree.left)
value = value + str(parseTree.val)
value = value + printParenthesizedTree(parseTree.right) + ')'
return value
def print_tree_2D(root):
""" Wrapper over print2DUtil()
Ref: https://www.geeksforgeeks.org/print-binary-tree-2-dimensions/
"""
def print2DUtil(root, space, space_between_levels):
"""# Function to print binary tree in 2D. It does reverse inorder traversal
Ref: https://www.geeksforgeeks.org/print-binary-tree-2-dimensions/
"""
# Base case
if (root == None):
return
# Increase distance between levels
space += space_between_levels
# Process right first
print2DUtil(root.right, space, space_between_levels)
# Print current node after space count
print()
for _ in range(space_between_levels, space):
print(end = " ")
print(root.val)
# Process left
print2DUtil(root.left, space, space_between_levels)
space = 0 # Pass initial space count as 0
space_between_levels = 10 # constant variable, i.e., 10 spaces between levels
print2DUtil(root, space, space_between_levels)
"""================================== TEST ================================="""
# Driver Program to test above functions
print("===================BINARY TREE===================")
""" 3
/ \
2 7
/ \ / \
1 5 6 9
/
8
"""
root = Node(3)
for _ in [2,7,1,5,6,9,8]:
insert(root, _)
print_tree_2D(root)
root2 = Node(3)
for _ in [2,7,1,5,6,9,8]:
insert(root2, _)
print("same tree? ", detectSameTree(root, root2))
print("Diameter of the given binary tree is ", diameter(root))
print("(BFS) Level Order Traversal of binary tree is -")
printLevelOrder(root)
print("\n(BFS) Reverese Level Order Traversal of binary tree is -")
printRevereseLevelOrder(root)
print("\n(BFS) Spiral Order Traversal of binary tree is -")
printSpiralOrder(root)
print("\n(DFS) Preorder traversal of binary tree is -")
printPreorder_recursive(root)
print("\n(DFS) Inorder traversal of binary tree is -")
printInorder_recursive(root)
print("\n(DFS) Postorder traversal of binary tree is -")
printPostorder_recursive(root)
# Test search
print("\nSearcing for 9, found? ", search(root, 9)) # Should be True
print("Searching for 51, found? ", search(root, 51)) # Should be False
# Test if tree is BST
print("\nIs this tree a BST? Answer: ", isBST_recursive(root)) # False
print("Is this tree a BST? Answer: ", isBST_recursive(root.right)) # True
print("Is this tree a BST (iterative)? Answer: ", isBST_iterative(root)) # False
print("Is this tree a BST (iterative)? Answer: ", isBST_iterative(root.right)) # True
# Lowest Common Ancestor
lca1 = lca(root, root.left.left, root.right.right.left)
print("\nLCA of (1,8)? Answer: ", lca1.val) # 3 (i.e., root)
lca2 = lca(root, root.right.left, root.right.right.left)
print("LCA of (6,8)? Answer: ", lca2.val) # 7 (i.e., root.right)
lca3 = lca(root, root.left, root.left.left)
print("LCA of (2,1)? Answer: ", lca3.val) # 2 (i.e., root.left)
# Test largest BST within the tree
print("\nSize of the largest BST within this tree is: ", largestBST(root))
print("(Optimized) Size of the largest BST within this tree is: ", largestBST_Optimized(root))
# Test insertion
key = 12
print("\nInorder traversal before insertion of key:")
printInorder_recursive(root)
insert(root, key);
print("\nBINARY TREE AFTER INSERTION of 12: ")
"""Constructed binary tree after insertion:
3
/ \
2 7
/ \ / \
1 5 6 9
/ \
8 12
"""
print_tree_2D(root)
print("Diameter of the given binary tree is ", diameter(root))
print("\nInorder traversal after insertion of key:")
printInorder_recursive(root)
print("\nBinary tree after deletion of key=2: ")
delete(root, 2);
print_tree_2D(root)
print("\nInorder traversal after deletion of key:")
printInorder_recursive(root)
# Test largest BST within the tree after insertion
print("\nSize of the largest BST within this tree is: ", largestBST(root))
print("(Optimized) Size of the largest BST within this tree is: ", largestBST_Optimized(root))
print("\n================== CONVERT BINARY TREE TO CIRCULAR DOUBLY LINKED LIST ================== ")
print("Given binary tree:")
print_tree_2D(root)
head = binaryTreeToCircularLinkedList(root)
print("\nAfter converting ", end=" "),
displayCircularLinkedList(head)
print("\n================== CONNECT SAME LEVEL SIBLINGS ================== ")
test_connect_same_level_siblings()
print("\n===================== TEST PROGRAM FOR PARSE TREES (USE CASE: NUMERICAL EQ.) ===================")
parse_tree = buildParsparseTree("( ( 10 + 5 ) * 3 )")
print("Print the parse tree with parenthesis: ", printParenthesizedTree(parse_tree))
print_tree_2D(parse_tree)
print("Inorder traversal: ", end ="")
printInorder_recursive(parse_tree)
print("\nPreorder traversal: ", end ="")
printPreorder_recursive(parse_tree)
print("\nPostorder traversal: ", end ="")
printPostorder_recursive(parse_tree)
print("\nEvaluting the above expression: ", evaluate(parse_tree))
# +
"""BST Implementation Educative"""
from collections import deque
class Node:
def __init__(self, val):
self.val = val
self.left = None
self.right = None
def insert(self, val):
"""Time: Θ(h)"""
if self is None:
self = Node(val)
return
current = self
parent = None
while current != None:
parent = current
if val < current.val:
current = current.left
else:
current = current.right
if val < parent.val:
parent.left = Node(val)
else:
parent.right = Node(val)
def insert_recursive(self, val):
"""Time: Θ(h)"""
if self is None:
self = Node(val)
return
if val < self.val:
if self.left:
self.left.insert_recursive(val)
else:
self.left = Node(val)
return
else:
if self.right:
self.right.insert_recursive(val)
else:
self.right = Node(val)
return
def search(self, val):
"""Time: Θ(log(n))"""
if self is None:
return self
current = self
while current != None and val != current.val:
if val < current.val:
current = current.left
else:
current = current.right
return current
def search_recursive(self, val):
"""Time: Θ(log(n))"""
if self is None or self.val == val:
return self
elif val < self.val:
if self.left:
return self.left.search_recursive(val)
else:
return None
else:
if self.right:
return self.right.search_recursive(val)
else:
return None
def delete(self, val):
"""Time: Θ(h)"""
if self is None:
return False
current = self
while current and current.val != val:
parent = current
if current.val < val:
current = current.right
else:
current = current.left
if current is None or current.val != val: # not found
return False
elif current.right is None and current.left is None: # leaf node
if current.val < parent.val:
parent.left = None
else:
parent.right = None
return True
elif current.right and current.left is None: # has right subtree
if current.val < parent.val:
parent.left = current.right
else:
parent.right = current.right
return True
elif current.right is None and current.left: # has left subtree
if current.val < parent.val:
parent.left = current.left
else:
parent.right = current.left
return True
elif current.right and current.left: # has both right & left subtrees
replaceParent = current
replaceNode = current.right
while replaceNode.left:
replaceParent = replaceNode
replaceNode = replaceNode.left
current.val = replaceNode.val
if replaceNode.right:
if replaceNode.val < replaceParent.val:
replaceParent.left = replaceNode.right
else:
replaceParent.right = replaceNode.right
return True
else:
if replaceNode.val < replaceParent.val:
replaceParent.left = None
else:
replaceParent.right = None
return True
class BST:
def __init__(self, val):
self.root = Node(val)
def insert(self, val):
self.root.insert(val)
def search(self, val):
return self.root.search(val)
def delete(self, val):
return self.root.delete(val)
def minValueNode(root): # Time: Θ(log(n))
"""
Given a non-empty binary search tree, return the node
with minimum key value found in that tree. Note that the
entire tree does not need to be searched.
"""
current = root
while(current.left != None): # loop down to find the leftmost leaf
current = current.left
return current
def print_levelOrder(root):
"""Time: Θ(log(n)), Space: Θ(n)"""
if root is None:
return
q = deque()
q.append(root) # enqueue
while len(q) > 0:
node = q.popleft() # dequeue
print(node.val, end=" ")
if node.left:
q.append(node.left)
if node.right:
q.append(node.right)
def print_preorder(root):
"""Time: Θ(log(n)), Space: Θ(1)"""
if root != None:
print(root.val, end=" "),
print_preorder(root.left)
print_preorder(root.right)
def print_inorder(root):
"""Time: Θ(log(n)), Space: Θ(1)"""
if root != None:
print_inorder(root.left)
print(root.val, end=" "),
print_inorder(root.right)
def print_postorder(root):
"""Time: Θ(log(n)), Space: Θ(1)"""
if root != None:
print_postorder(root.left)
print_postorder(root.right)
print(root.val, end=" "),
def isBST(root): # Time: O(n) , Space: O(1)
""" Ref: https://www.youtube.com/watch?v=MILxfAbIhrE&list=PLrmLmBdmIlpv_jNDXtJGYTPNQ2L1gdHxu&index=8 """
def isBST_rec(root, minValue, maxValue):
if root == None:
return True
if root.val <= minValue or root.val >= maxValue:
return False
return isBST_rec(root.left, minValue, root.val) and isBST_rec(root.right, root.val, maxValue)
minValue = float("-inf")
maxValue = float("inf")
return isBST_rec(root, minValue, maxValue)
def lca(root, node1, node2): # Time: O(h) , Space: O(1)
""" Lowest common ancestor of two nodes in a BST.
Time: O(h), where h is the height of the tree. """
if root.val > max(node1.val, node2.val):
return lca(root.left, node1, node2)
elif root.val < min(node1.val, node2.val):
return lca(root.right, node1, node2)
else:
return root
def findKthMax(root,k): # Time: O(n), Space: O(n)
""" Find the Kth maximum (value) node in the BST. """
def inOrderTraverse(node, treeList):
""" Helper recursive function to traverse the tree inorder """
if node != None:
inOrderTraverse(node.left, treeList)
treeList.append(node.val)
inOrderTraverse(node.right, treeList)
treeList = []
inOrderTraverse(root, treeList)
if (len(treeList) - k) >= 0: # kth value exists in resulting list
return treeList[-k]
return None
def findAncestors(root, k): # Time: O(log(n)), Space: O(n)
""" """
ancestors = []
current = root
while current:
if current.val > k:
ancestors.append(current.val)
current = current.left
elif current.val < k:
ancestors.append(current.val)
current = current.right
else: # when k == current.val
# add this line if you want to include the sought node itself in the result
# ancestors.append(current.val)
return ancestors[::-1] # [::-1] reverses the array
return []
def print_tree_2D(root):
"""
Function to print binary tree in 2D. It does reverse inorder traversal
Ref: https://www.geeksforgeeks.org/print-binary-tree-2-dimensions/
"""
def print2DUtil(root, space, count):
""" Recursive function to print a tree. """
# Base case
if (root == None):
return
# Increase distance between levels
space += count
# Process right first
print2DUtil(root.right, space, count)
# Print current node after space
# count
print()
for i in range(count, space):
print(end = " ")
print(root.val)
# Process left
print2DUtil(root.left, space, count)
space= 0 # Pass initial space count as 0
count = 10
print2DUtil(root, space, count)
"""================================== TEST ================================="""
#Driver Program to test above functions
"""
Constructed binary search tree is
10
/ \
-10 30
\ / \
8 25 60
/ \ \ \
6 9 28 78
"""
tree = BST(10)
for _ in [-10, 30, 8, 6, 9, 25, 60, 28, 78]:
tree.insert(_)
print("===================BINARY SEARCH TREE===================")
COUNT = [10]
print_tree_2D(tree.root)
print("\n(DFS) Preorder traversal of BST is -")
print_preorder(tree.root)
print("\n(DFS) Inorder traversal of BST is -")
print_inorder(tree.root)
print("\n(DFS) Postorder traversal of BST is -")
print_postorder(tree.root)
print("\n(BFS) Level Order traversal of BST is -")
print_levelOrder(tree.root)
print("\nFind the 3rd Max value of the node in the tree: ", findKthMax(tree.root, 3))
print("Find the ancestors of 28: ", findAncestors(tree.root, 28))
# Test if tree is BST
print("Is this tree a BST? Answer: ", isBST(tree.root))
# -
sentence = ' hello '
''.join(sentence.split())
# +
# Program to convert binary tree to BST
""" Program to convert a given binary tree to BST.
Binary Tree to Binary Search Tree Conversion: Example 1
Input:
10
/ \
2 7
/ \
8 4
Output:
8
/ \
4 10
/ \
2 7
Binary Tree to Binary Search Tree Conversion: Example 2
Input:
10
/ \
30 15
/ \
20 5
Output:
15
/ \
10 20
/ \
5 30
Algorithm: Following is a 3 step solution for converting Binary
tree to Binary Search Tree:
1) Create a temp array arr[] that stores inorder traversal of the tree.
This step takes O(n) time and O(n) space.
2) Sort the temp array arr[]. Time complexity of this step depends upon
the sorting algorithm. In the following implementation, Quick Sort is
used which takes (n^2) time. This can be done in O(nLogn) time using Heap Sort
or Merge Sort.
3) Again do inorder traversal of tree and copy array elements to tree nodes
one by one. This step takes O(n) time.
"""
def print_tree_2D(root):
"""
Function to print binary tree in 2D. It does reverse inorder traversal
Ref: https://www.geeksforgeeks.org/print-binary-tree-2-dimensions/
"""
def print2DUtil(root, space, count):
""" Recursive function to print a tree. """
# Base case
if (root == None):
return
# Increase distance between levels
space += count
# Process right first
print2DUtil(root.right, space, count)
# Print current node after space
# count
print()
for i in range(count, space):
print(end = " ")
print(root.val)
# Process left
print2DUtil(root.left, space, count)
space= 0 # Pass initial space count as 0
count = 10
print2DUtil(root, space, count)
class Node:
""" A binary tree node """
def __init__(self, data):
self.val = data
self.left = None
self.right = None
"""=================================== Binary Tree to BST ==================================="""
def binaryTreeToBST(root): # Time: O(nlog(n)), Space: O(n)
if root is None: # Base Case: Tree is empty
return
arr = [] # Create the temp array
storeInorder(root, arr) # and in it store the inorderTraveral of tree
arr.sort() # Sort the array
arrayToBST(arr, root) # Copy array elements back to binary tree
def storeInorder(root, array):
""" Helper function to store the inroder traversal of a tree. """
if root:
storeInorder(root.left, array) # First store the left subtree
array.append(root.val) # Copy the root's data
storeInorder(root.right, array) # Finally store the right subtree
def arrayToBST(arr, node):
""" Helper function that copies contents of sorted array to Binary tree """
if node is None: # Base Case
return
arrayToBST(arr, node.left) # First update the left subtree
node.val = arr[0] # update root's data
arr.pop(0) # delete the value from array
arrayToBST(arr, node.right) # Finally update the right subtree
# Driver program to test above function
root = Node(10)
root.left = Node(30)
root.right = Node(15)
root.left.left = Node(20)
root.right.right= Node(5)
print("\n================== CONVERT BINARY TREE TO BST ================== ")
print("Given Binary Tree: ")
print_tree_2D(root)
binaryTreeToBST(root) # Convert binary tree to BST
print("Converted BST: ")
print_tree_2D(root)
print("Following is the inorder traversal of the converted BST")
printInorder_recursive(root)
# +
""" AVL Tree: Time: Θ(log(n))
Source: https://www.geeksforgeeks.org/avl-tree-set-2-deletion/
https://www.geeksforgeeks.org/avl-tree-set-2-deletion/
AVL tree is a special kind of binary search tree that automatically makes sure
that the tree remains balanced at all times. This tree is called an AVL tree
and is named for its inventors:
<NAME> and <NAME>.
To implement our AVL tree we need to keep track of a balance factor for each
node in the tree. We do this by looking at the heights of the left and right
subtrees for each node. More formally, we define the balance factor for a node
as the difference between the height of the left subtree
and the height of the right subtree.
balanceFactor = height(leftSubTree) − height(rightSubTree)
If the balance factor is zero then the tree is perfectly in balance.
For purposes of implementing an AVL tree, and gaining the benefit of having
a balanced tree we will define a tree to be in balance if the balance factor
is -1, 0, or 1. Once the balance factor of a node in a tree is outside this
range we will need to have a procedure to bring the tree back into balance.
"""
class Node:
""" Generic tree node class """
def __init__(self, val):
self.val = val
self.left = None
self.right = None
self.height = 1
class AVL_Tree:
""" AVL tree class which supports the insert operation. """
def insert(self, root, key):
""" Recursive function to insert key in subtree rooted with node
and returns new root of subtree. """
# Step 1 - Perform normal BST
if not root:
return Node(key)
elif key < root.val:
root.left = self.insert(root.left, key)
else:
root.right = self.insert(root.right, key)
# Step 2 - Update the height of the ancestor node
root.height = 1 + max(self.getHeight(root.left), self.getHeight(root.right))
# Step 3 - Get the balance factor
balance = self.getBalance(root)
# Step 4 - If the node is unbalanced, then try out the 4 cases
if balance > 1 and key < root.left.val: # Case 1 - Left Left
return self.rightRotate(root)
if balance > 1 and key > root.left.val: # Case 2 - Left Right
root.left = self.leftRotate(root.left)
return self.rightRotate(root)
if balance < -1 and key > root.right.val: # Case 3 - Right Right
return self.leftRotate(root)
if balance < -1 and key < root.right.val: # Case 4 - Right Left
root.right = self.rightRotate(root.right)
return self.leftRotate(root)
return root
def delete(self, root, key):
"""
Recursive function to delete a node with
given key from subtree with given root.
It returns root of the modified subtree.
"""
# Step 1 - Perform standard BST delete
if not root:
return root
elif key < root.val:
root.left = self.delete(root.left, key)
elif key > root.val:
root.right = self.delete(root.right, key)
else: # root.val == key
if root.left is None:
temp = root.right
root = None
return temp
elif root.right is None:
temp = root.left
root = None
return temp
# elif root.left and root.left:
temp = self.getMinValueNode(root.right)
root.val = temp.val
root.right = self.delete(root.right,
temp.val)
# If the tree has only one node,
# simply return it
if root is None:
return root
# Step 2 - Update the height of the
# ancestor node
root.height = 1 + max(self.getHeight(root.left),
self.getHeight(root.right))
# Step 3 - Get the balance factor
balance = self.getBalance(root)
# Step 4 - If the node is unbalanced,
# then try out the 4 cases
# Case 1 - Left Left
if balance > 1 and self.getBalance(root.left) >= 0:
return self.rightRotate(root)
# Case 2 - Right Right
if balance < -1 and self.getBalance(root.right) <= 0:
return self.leftRotate(root)
# Case 3 - Left Right
if balance > 1 and self.getBalance(root.left) < 0:
root.left = self.leftRotate(root.left)
return self.rightRotate(root)
# Case 4 - Right Left
if balance < -1 and self.getBalance(root.right) > 0:
root.right = self.rightRotate(root.right)
return self.leftRotate(root)
return root
def leftRotate(self, root):
node = root.right
temp = node.left
# Perform rotation
node.left = root
root.right = temp
# Update heights
root.height = 1 + max(self.getHeight(root.left),
self.getHeight(root.right))
node.height = 1 + max(self.getHeight(node.left),
self.getHeight(node.right))
# Return the new root
return node
def rightRotate(self, root):
node = root.left
temp = node.right
# Perform rotation
node.right = root
root.left = temp
# Update heights
root.height = 1 + max(self.getHeight(root.left),
self.getHeight(root.right))
node.height = 1 + max(self.getHeight(node.left),
self.getHeight(node.right))
# Return the new root
return node
def getHeight(self, root):
if not root:
return 0
return root.height
def getBalance(self, root):
if not root:
return 0
return self.getHeight(root.left) - self.getHeight(root.right)
def getMinValueNode(self, root):
current = root
while current.left != None:
current = current.left
return current
def preOrder(self, root):
if not root:
return
print("{0} ".format(root.val), end="")
self.preOrder(root.left)
self.preOrder(root.right)
def print_tree_2D(root):
"""
Function to print binary tree in 2D. It does reverse inorder traversal
Ref: https://www.geeksforgeeks.org/print-binary-tree-2-dimensions/
"""
def print2DUtil(root, space, count):
""" Recursive function to print a tree. """
# Base case
if (root == None):
return
# Increase distance between levels
space += count
# Process right first
print2DUtil(root.right, space, count)
# Print current node after space
# count
print()
for i in range(count, space):
print(end = " ")
print(root.val)
# Process left
print2DUtil(root.left, space, count)
space= 0 # Pass initial space count as 0
count = 10
print2DUtil(root, space, count)
# Driver program to test above function
"""
The constructed AVL Tree would be
30
/ \
20 40
/ \ \
10 25 50
"""
myTree = AVL_Tree()
root = None
root = myTree.insert(root, 10)
root = myTree.insert(root, 20)
root = myTree.insert(root, 30)
root = myTree.insert(root, 40)
root = myTree.insert(root, 50)
root = myTree.insert(root, 25)
root = myTree.insert(root, 33)
root = myTree.insert(root, 51)
print_tree_2D(root)
# Preorder Traversal
print("Preorder traversal of the", "constructed AVL tree is")
myTree.preOrder(root)
print()
# Delete
root = myTree.delete(root, 10)
root = myTree.delete(root, 20)
print_tree_2D(root)
# Preorder Traversal
print("Preorder Traversal after deletion of nodes -")
myTree.preOrder(root)
print()
# +
""" Red Black Tree: Time: Θ(log(n))
Red-Black Tree is a self-balancing Binary Search Tree (BST)
where every node follows following rules:
1) Every node has a color either red or black.
2) Root of tree is always black
3) There are no two adjacent red nodes
(A red node cannot have a red parent or red child).
4) Every path from a node (including root) to any of its
descendant NULL node has the same number of black nodes.
Source: https://www.geeksforgeeks.org/red-black-tree-set-1-introduction-2/
"""
"""An implementation of red-black trees, based on the description in
Introduction to Algorithms (Cormen, Leiserson, Rivest), Chapter 14.
# https://www.hashcollision.org/hkn/python/red_black/red_black.py
"""
"""Each node can be colored RED or BLACK."""
RED = "RED"
BLACK = "BLACK"
class NilNode(object):
def __init__(self):
self.color = BLACK
"""We define NIL to be the leaf sentinel of our tree."""
NIL = NilNode()
class Node(object):
def __init__(self, key, color=RED, left=NIL, right=NIL, p=NIL):
"""Constructs a single node of the red-black tree.
Key is the key that has an ordering.
color is RED or BLACK.
left and right are the left and right subtrees.
p is the parent Node.
"""
assert color in (RED, BLACK)
self.color = color
self.key = key
self.left = left
self.right = right
self.parent = p
class Tree(object):
def __init__(self, root=NIL):
self.root = root
def left_rotate(tree, x):
"""Left-rotates node x on tree tree.
x
/ \
a y
/ \
b g
mutates into:
y
/ \
x g
/ \
a b
Used for maintaining tree balance.
"""
assert (x.right != NIL)
y = x.right
x.right = y.left
if y.left != NIL:
y.left.parent = x
y.parent = x.parent
if x.parent == NIL:
tree.root = y
elif x == x.parent.left:
x.parent.left = y
else:
x.parent.right = y
y.left = x
x.parent = y
def right_rotate(tree, x):
"""Right-rotates node x on tree tree.
x
/ \
y g
/ \
a b
mutates into:
y
/ \
a x
/ \
b g
Used for maintaining tree balance.
"""
assert (x.left != NIL)
y = x.left
x.left = y.right
if y.right != NIL:
y.right.parent = x
y.parent = x.parent
if x.parent == NIL:
tree.root = y
elif x == x.parent.right:
x.parent.right = y
else:
x.parent.left = y
y.right = x
x.parent = y
def tree_insert(tree, z):
"""Inserts node 'z' into binary tree 'tree'."""
y = NIL
x = tree.root
while x != NIL:
y = x
if z.key < x.key:
x = x.left
else:
x = x.right
z.p = y
if y == NIL:
tree.root = z
elif z.key < y.key:
y.left = z
else:
y.right = z
def rb_insert(tree, x):
"""Does an insertion of 'x' into the red-black tree 'tree'. The
algorithm here is a little subtle, but is explained in CLR."""
tree_insert(tree, x)
x.color = RED
while x != tree.root and x.p.color == RED:
if x.p == x.p.p.left:
y = x.p.p.right
if y.color == RED:
x.p.color = BLACK
y.color = BLACK
x.p.p.color = RED
x = x.p.p
else:
if x == x.p.right:
x = x.p
left_rotate(tree, x)
x.p.color = BLACK
x.p.p.color = RED
right_rotate(tree, x.p.p)
else:
y = x.p.p.left
if y.color == RED:
x.p.color = BLACK
y.color = BLACK
x.p.p.color = RED
x = x.p.p
else:
if x == x.p.left:
x = x.p
right_rotate(tree, x)
x.p.color = BLACK
x.p.p.color = RED
left_rotate(tree, x.p.p)
tree.root.color = BLACK
def tree_minimum(x):
"""Returns the minimal element of the subtree rooted at 'x'."""
while x.left != NIL:
x = x.left
return x
def tree_maximum(x):
"""Returns the maximal element of the subtree rooted at 'x'."""
while x.right != NIL:
x = x.right
return x
def tree_successor(x):
"""Returns the inorder successor of node 'x'."""
if x.right != NIL:
return tree_minimum(x.right)
y = x.p
while y != NIL and x == y.right:
x = y
y = y.p
return y
def tree_predecessor(x):
"""Returns the inorder predecessor of node 'x'."""
if x.left != NIL:
return tree_maximum(x.left)
y = x.p
while y != NIL and x == y.left:
x =y
y = y.p
return y
def tree_height(node):
"""Returns the height of a subtree rooted by node 'node'."""
if node == NIL: return 0
return max(1 + tree_height(node.left), 1 + tree_height(node.right))
def tree_count_internal(node):
"""Returns the number of internal nodes in the subtree rooted at 'node'."""
if node == NIL: return 0
return 1 + tree_count_internal(node.left) + tree_count_internal(node.right)
######################################################################
## Unit tests
import unittest
class RedBlackTest(unittest.TestCase):
def setUp(self):
pass
def tearDown(self):
pass
def testRotationOneElementTree(self):
"""Checks to see that left and right rotation on one-element
trees throws AssertionErrors."""
tree = Tree()
tree.root = Node(5)
self.assertRaises(AssertionError, left_rotate, tree, tree.root)
self.assertRaises(AssertionError, right_rotate, tree, tree.root)
def testBinaryInsertion(self):
"""Checks that we get
5
/ \
3 7
/ /
1 6
"""
one, three, five, six, seven = map(lambda x: Node(x),
[1, 3, 5, 6, 7])
tree = Tree()
tree_insert(tree, five)
tree_insert(tree, three)
tree_insert(tree, seven)
tree_insert(tree, one)
tree_insert(tree, six)
self.assertEquals(tree.root, five)
self.assertEquals(tree.root.left, three)
self.assertEquals(tree.root.left.left, one)
self.assertEquals(tree.root.right, seven)
self.assertEquals(tree.root.right.left, six)
def testTreeInsertHundredElements(self):
MAX = 100
nodes = map(Node, range(MAX))
tree = Tree()
for n in nodes: tree_insert(tree, n)
self.assertEquals(nodes[0], tree_minimum(tree.root))
for i in range(MAX-1):
self.assertEquals(nodes[i], tree_predecessor(nodes[i+1]))
## Worse case input to a binary tree should produce one long chain!
self.assertEquals(MAX, tree_height(tree.root))
def testRotationTwoElementTree(self):
"""Check to see that
x
/
y
transforms to
y
\
x
and back again.
"""
tree = Tree()
x = Node('x', 'BLACK')
y = Node('y', 'RED', p=x)
x.left = y
tree.root = x
right_rotate(tree, x)
self.assertEquals(tree.root, y)
self.assertEquals(tree.root.right, x)
self.assertEquals(tree.root.left, NIL)
self.assertEquals((tree.root.right.left, tree.root.right.right),
(NIL, NIL))
left_rotate(tree, y)
self.assertEquals(tree.root, x)
self.assertEquals(tree.root.left, y)
self.assertEquals(tree.root.right, NIL)
self.assertEquals((tree.root.left.left, tree.root.left.right),
(NIL, NIL))
def testRbInsertOneElement(self):
tree = Tree()
one = Node('1')
rb_insert(tree, one)
self.assertEquals(tree.root, one)
self.assertEquals(tree.root.color, BLACK)
def testRbInsertTwoElements(self):
tree = Tree()
nodes = map(Node, range(2))
for n in nodes:
rb_insert(tree, n)
self.assertEquals(tree.root, nodes[0])
self.assertEquals(tree.root.right, nodes[1])
def testRbInsertThreeElements(self):
"""
We expect to get:
1b
/ \
0r 2r
"""
tree = Tree()
nodes = map(Node, range(3))
for n in nodes:
rb_insert(tree, n)
self.assertEquals(tree.root, nodes[1])
self.assertEquals(tree.root.left, nodes[0])
self.assertEquals(tree.root.right, nodes[2])
self.assertEquals([RED, BLACK, RED], map(lambda n: n.color,
nodes))
self.assertEquals(nodes[0], tree_minimum(tree.root))
self.assertEquals(nodes[1], tree_successor(tree_minimum(tree.root)))
self.assertEquals(nodes[2], tree_successor
(tree_successor(tree_minimum(tree.root))))
self.assertEquals(NIL, tree_successor
(tree_successor(tree_successor
(tree_minimum(tree.root)))))
self.assertEquals(nodes[1], tree_predecessor(nodes[2]))
self.assertEquals(nodes[0], tree_predecessor(nodes[1]))
def testRbFourElements(self):
"""
1b
/ \
0b 2b
\
3r
"""
MAX = 4
nodes = map(Node, range(MAX))
tree = Tree()
for n in nodes: rb_insert(tree, n)
self.assertEquals(nodes[0], tree_minimum(tree.root))
for i in range(MAX-1):
self.assertEquals(nodes[i], tree_predecessor(nodes[i+1]))
self.assertEquals(nodes[1], tree.root)
self.assertEquals(nodes[0], tree.root.left)
self.assertEquals(nodes[2], tree.root.right)
self.assertEquals(nodes[3], tree.root.right.right)
self.assertEquals([BLACK, BLACK, BLACK, RED],
map(lambda n: n.color, nodes))
def isRbTreeHeightCorrect(self, tree):
"""By definition, the height of the resulting red-black tree
is less than or equal to 2* lg(number_of_nodes + 1)."""
def lg(x):
import math
return math.log(x) / math.log(2)
return (tree_height(tree.root)
<= 2 * lg(tree_count_internal(tree.root) + 1))
def testRbHundredElements(self, MAX=100):
nodes = map(Node, range(MAX))
tree = Tree()
for n in nodes: rb_insert(tree, n)
self.assertEquals(nodes[0], tree_minimum(tree.root))
for i in range(MAX-1):
self.assertEquals(nodes[i], tree_predecessor(nodes[i+1]))
def lg(x):
import math
return math.log(x) / math.log(2)
## By definition, the height of the resulting red-black tree
## is less than or equal to 2* lg(number_of_nodes + 1).
self.assert_(self.isRbTreeHeightCorrect(tree))
self.assertEquals(MAX, tree_count_internal(tree.root))
def testRbThousandElements(self):
self.testRbHundredElements(MAX=1000)
# if __name__ == '__main__':
# unittest.main()
# +
""" =================== OPTIONAL ========================
Binary Tree -- Alternate Array Implementation"""
def BinaryTree(r):
return [r, [], []]
def insertLeft(root,newBranch):
t = root.pop(1)
if len(t) > 1:
root.insert(1,[newBranch,t,[]])
else:
root.insert(1,[newBranch, [], []])
return root
def insertRight(root,newBranch):
t = root.pop(2)
if len(t) > 1:
root.insert(2,[newBranch,[],t])
else:
root.insert(2,[newBranch,[],[]])
return root
def getRootVal(root):
return root[0]
def setRootVal(root,newVal):
root[0] = newVal
def getLeftChild(root):
return root[1]
def getRightChild(root):
return root[2]
r = BinaryTree(3)
insertLeft(r,4)
insertLeft(r,5)
insertRight(r,6)
insertRight(r,7)
l = getLeftChild(r)
print(l)
setRootVal(l,9)
print(r)
insertLeft(l,11)
print(r)
print(getRightChild(getRightChild(r)))
# +
import heapq
minheap = [1,2,3,4,5,6,7,8,9,10,11,12,13,14,15]
heapq.heapify(minheap) # for a min heap
print(minheap)
# pop from minheap
while minheap:
print(heapq.heappop(minheap), end=" ") # pop from maxheap
print()
maxheap = [1,2,3,4,5,6,7,8,9,10,11,12,13,14,15]
heapq._heapify_max(maxheap)
print(maxheap)
while maxheap:
print(heapq._heappop_max(maxheap), end=" ") # pop from maxheap
print()
# +
from heapq import *
def find_Kth_smallest(lists, k):
minHeap = []
# put the 1st element of each list in the min heap
for list in enumerate(lists):
arrIdentifier, arr = list[0], list[1]
index = 0
heappush(minHeap, (arr[0], index, arrIdentifier))
# take the smallest(top) element form the min heap, if the running count is equal to k return the number
numberCount, number = 0, 0
while minHeap:
number, index, arrIdentifier = heappop(minHeap)
arr = lists[arrIdentifier]
numberCount += 1
if numberCount == k:
break
# if the array of the top element has more elements, add the next element to the heap
if len(arr) > index+1:
heappush(minHeap, (arr[index+1], index+1, arrIdentifier))
return number
def main():
print("Kth smallest number is: " +
str(find_Kth_smallest([[2, 6, 8], [3, 6, 7], [1, 3, 4]], 5)))
main()
# -
| DS_Algo/Trees.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/eremeich/hlll_course/blob/master/Python_Deep_Learning_And_Fine_Tuning.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="CZApnopCetN3" colab_type="text"
# # Using convnets with small datasets
#
#
# + id="YyX0droDqTxB" colab_type="code" outputId="8a41a091-ff8e-4557-d286-ef2c3d16a9c9" colab={"base_uri": "https://localhost:8080/", "height": 97}
import keras
keras.__version__
# + id="tpnUPI5VqirF" colab_type="code" colab={}
import os, shutil
# + [markdown] id="ZjlyVp5WDCvn" colab_type="text"
# ## Downloading the data
#
# The cats vs. dogs dataset that we will use isn't packaged with Keras. It was made available by Kaggle.com as part of a computer vision competition in late 2013, back when convnets weren't quite mainstream.
#
# https://www.kaggle.com/c/dogs-vs-cats/data
#
#
# + id="dxYrt1l7w8EP" colab_type="code" outputId="f34ca809-586f-4e49-ca33-c4cdeaa54533" colab={"base_uri": "https://localhost:8080/", "height": 122}
from google.colab import drive
drive.mount('/content/gdrive')
# + id="OW4jt_hJxcyD" colab_type="code" outputId="99e13de8-595a-468b-8128-4d50cf803e22" colab={"base_uri": "https://localhost:8080/", "height": 34}
# !ls /content/gdrive/My\ Drive/dogs-vs-cats/
# !apt-get -qq install -y unzip
# + id="N2Pxl38mtVky" colab_type="code" outputId="21db0623-b3c4-4e62-e657-4b1afa0cdfc0" colab={"base_uri": "https://localhost:8080/", "height": 578}
# !apt-get update
# !apt-get -qq install -y libarchive-dev && pip install -q -U libarchive
# !apt-get -qq install -y unzip
import libarchive
# + id="qDaJwpLC3j_8" colab_type="code" colab={}
# !unzip /content/gdrive/My\ Drive/dogs-vs-cats/train.zip
# # !ls train | wc -l
# # !rm -rf small
# + id="1kGMrQfYG7Po" colab_type="code" outputId="07b831cc-ffef-4f38-b4c6-1c8c98090e50" colab={"base_uri": "https://localhost:8080/", "height": 34}
# !ls
# + [markdown] id="EHgsGVgcEFG5" colab_type="text"
# After downloading and uncompressing it, we will create a new dataset containing three subsets: a training set with 1000 samples of each class, a validation set with 500 samples of each class, and finally a test set with 500 samples of each class.
# + id="FPCA839Oqn-r" colab_type="code" colab={}
# !rm -rf small
# The path to the directory where the original
# dataset was uncompressed
original_dataset_dir = 'train'
# The directory where we will
# store our smaller dataset
base_dir = 'small'
os.mkdir(base_dir)
# Directories for our training,
# validation and test splits
train_dir = os.path.join(base_dir, 'train')
os.mkdir(train_dir)
validation_dir = os.path.join(base_dir, 'validation')
os.mkdir(validation_dir)
test_dir = os.path.join(base_dir, 'test')
os.mkdir(test_dir)
# Directory with our training cat pictures
train_cats_dir = os.path.join(train_dir, 'cats')
os.mkdir(train_cats_dir)
# Directory with our training dog pictures
train_dogs_dir = os.path.join(train_dir, 'dogs')
os.mkdir(train_dogs_dir)
# Directory with our validation cat pictures
validation_cats_dir = os.path.join(validation_dir, 'cats')
os.mkdir(validation_cats_dir)
# Directory with our validation dog pictures
validation_dogs_dir = os.path.join(validation_dir, 'dogs')
os.mkdir(validation_dogs_dir)
# Directory with our validation cat pictures
test_cats_dir = os.path.join(test_dir, 'cats')
os.mkdir(test_cats_dir)
# Directory with our validation dog pictures
test_dogs_dir = os.path.join(test_dir, 'dogs')
os.mkdir(test_dogs_dir)
# Copy first 1000 cat images to train_cats_dir
fnames = ['cat.{}.jpg'.format(i) for i in range(1000)]
for fname in fnames:
src = os.path.join(original_dataset_dir, fname)
dst = os.path.join(train_cats_dir, fname)
shutil.copyfile(src, dst)
# Copy next 500 cat images to validation_cats_dir
fnames = ['cat.{}.jpg'.format(i) for i in range(1000, 1500)]
for fname in fnames:
src = os.path.join(original_dataset_dir, fname)
dst = os.path.join(validation_cats_dir, fname)
shutil.copyfile(src, dst)
# Copy next 500 cat images to test_cats_dir
fnames = ['cat.{}.jpg'.format(i) for i in range(1500, 2000)]
for fname in fnames:
src = os.path.join(original_dataset_dir, fname)
dst = os.path.join(test_cats_dir, fname)
shutil.copyfile(src, dst)
# Copy first 1000 dog images to train_dogs_dir
fnames = ['dog.{}.jpg'.format(i) for i in range(1000)]
for fname in fnames:
src = os.path.join(original_dataset_dir, fname)
dst = os.path.join(train_dogs_dir, fname)
shutil.copyfile(src, dst)
# Copy next 500 dog images to validation_dogs_dir
fnames = ['dog.{}.jpg'.format(i) for i in range(1000, 1500)]
for fname in fnames:
src = os.path.join(original_dataset_dir, fname)
dst = os.path.join(validation_dogs_dir, fname)
shutil.copyfile(src, dst)
# Copy next 500 dog images to test_dogs_dir
fnames = ['dog.{}.jpg'.format(i) for i in range(1500, 2000)]
for fname in fnames:
src = os.path.join(original_dataset_dir, fname)
dst = os.path.join(test_dogs_dir, fname)
shutil.copyfile(src, dst)
# + id="C3HMKjb63EcO" colab_type="code" outputId="5f3eb0a5-0d7e-496b-cbc3-fd9c4bcdfad9" colab={"base_uri": "https://localhost:8080/", "height": 323}
# !apt-get -qq install -y tree
# !tree small -d
# + id="uKbXLRPQ4k84" colab_type="code" outputId="2a6a954c-4753-4926-dd1b-657e2dd78def" colab={"base_uri": "https://localhost:8080/", "height": 34}
print('total training cat images:', len(os.listdir(train_cats_dir)))
# + id="JhGQA2cG5cLy" colab_type="code" outputId="bde371d3-0a43-4b49-e37e-accbc06a2dee" colab={"base_uri": "https://localhost:8080/", "height": 34}
print('total training dog images:', len(os.listdir(train_dogs_dir)))
# + id="GiuYsFC65g5b" colab_type="code" outputId="91b5305e-42fd-4ba3-8ee2-f1bcc65ff6bb" colab={"base_uri": "https://localhost:8080/", "height": 34}
print('total validation cat images:', len(os.listdir(validation_cats_dir)))
# + id="Q3pIIRPl5k9o" colab_type="code" outputId="1f2ded18-a04d-4433-a342-166f51f989e8" colab={"base_uri": "https://localhost:8080/", "height": 34}
print('total validation dog images:', len(os.listdir(validation_dogs_dir)))
# + id="Wb1YUd9L5owt" colab_type="code" outputId="428f45d2-36ee-467c-efd5-349075c58cce" colab={"base_uri": "https://localhost:8080/", "height": 34}
print('total test cat images:', len(os.listdir(test_cats_dir)))
# + id="I00qAJTD5wJK" colab_type="code" outputId="0330489f-9d5d-43ba-b1dd-14c73b417daf" colab={"base_uri": "https://localhost:8080/", "height": 34}
print('total test dog images:', len(os.listdir(test_dogs_dir)))
# + [markdown] id="Ga-xz1PlFA3W" colab_type="text"
# ## Building our network
#
# Our convnet will be a stack of alternated Conv2D (with relu activation) and MaxPooling2D layers.
# + id="u5hpiQ0B50Y5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 173} outputId="f0ccd48f-5c03-4327-d643-a80b33f2e776"
from keras import layers
from keras import models
model = models.Sequential()
model.add(layers.Conv2D(32, (3, 3), activation='relu',
input_shape=(150, 150, 3)))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Conv2D(64, (3, 3), activation='relu'))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Conv2D(128, (3, 3), activation='relu'))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Conv2D(128, (3, 3), activation='relu'))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Flatten())
model.add(layers.Dense(512, activation='relu'))
model.add(layers.Dense(1, activation='sigmoid'))
# + [markdown] id="T7jpv19EFk0_" colab_type="text"
# Let's take a look at how the dimensions of the feature maps change with every successive layer:
# + id="nLcp-b9lCYxR" colab_type="code" outputId="bfc68155-a89b-4b09-b023-6cfaeee12e36" colab={"base_uri": "https://localhost:8080/", "height": 527}
model.summary()
# + [markdown] id="DKJfmUG_Fuwl" colab_type="text"
# For our compilation step, we'll go with the RMSprop optimizer as usual. Since we ended our network with a single sigmoid unit, we will use binary crossentropy as our loss (as a reminder, check out the table in Chapter 4, section 5 for a cheatsheet on what loss function to use in various situations).
# + id="RQoe1jq1CipN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 156} outputId="4e180940-0e1d-4462-fb0b-3003dfd3a766"
from keras import optimizers
model.compile(loss='binary_crossentropy',
optimizer=optimizers.RMSprop(lr=1e-4),
metrics=['acc'])
# + [markdown] id="0QLUjrpMF-sH" colab_type="text"
# ## Data preprocessing
#
# Data should be formatted into appropriately pre-processed floating point tensors before being fed into our network. Currently, our data sits on a drive as JPEG files, so the steps for getting it into our network are roughly:
#
# * Read the picture files.
# * Decode the JPEG content to RBG grids of pixels.
# * Convert these into floating point tensors.
# * Rescale the pixel values (between 0 and 255) to the [0, 1] interval (as you know, neural networks prefer to deal with small input values).
#
# It may seem a bit daunting, but thankfully Keras has utilities to take care of these steps automatically. Keras has a module with image processing helper tools, located at keras.preprocessing.image. In particular, it contains the class ImageDataGenerator which allows to quickly set up Python generators that can automatically turn image files on disk into batches of pre-processed tensors. This is what we will use here.
# + id="_ZLUYxOyComq" colab_type="code" outputId="c0c865bd-24c2-491f-8773-0959320bd59b" colab={"base_uri": "https://localhost:8080/", "height": 51}
from keras.preprocessing.image import ImageDataGenerator
# All images will be rescaled by 1./255
train_datagen = ImageDataGenerator(rescale=1./255)
test_datagen = ImageDataGenerator(rescale=1./255)
train_generator = train_datagen.flow_from_directory(
# This is the target directory
train_dir,
# All images will be resized to 150x150
target_size=(150, 150),
batch_size=20,
# Since we use binary_crossentropy loss, we need binary labels
class_mode='binary')
validation_generator = test_datagen.flow_from_directory(
validation_dir,
target_size=(150, 150),
batch_size=20,
class_mode='binary')
# + [markdown] id="k7By6TyEGmch" colab_type="text"
# Let's take a look at the output of one of these generators: it yields batches of 150x150 RGB images (shape (20, 150, 150, 3)) and binary labels (shape (20,)). 20 is the number of samples in each batch (the batch size). Note that the generator yields these batches indefinitely: it just loops endlessly over the images present in the target folder. For this reason, we need to break the iteration loop at some point.
# + id="PbyD7GnuDLZ4" colab_type="code" outputId="f7ee5e77-582a-4dc5-86c0-81684d3b0b94" colab={"base_uri": "https://localhost:8080/", "height": 51}
for data_batch, labels_batch in train_generator:
print('data batch shape:', data_batch.shape)
print('labels batch shape:', labels_batch.shape)
break
# + [markdown] id="YXVbodH0G1Pq" colab_type="text"
# Let's fit our model to the data using the generator. We do it using the fit_generator method, the equivalent of fit for data generators like ours. It expects as first argument a Python generator that will yield batches of inputs and targets indefinitely, like ours does. Because the data is being generated endlessly, the generator needs to know example how many samples to draw from the generator before declaring an epoch over. This is the role of the steps_per_epoch argument: after having drawn steps_per_epoch batches from the generator, i.e. after having run for steps_per_epoch gradient descent steps, the fitting process will go to the next epoch. In our case, batches are 20-sample large, so it will take 100 batches until we see our target of 2000 samples.
#
# When using fit_generator, one may pass a validation_data argument, much like with the fit method. Importantly, this argument is allowed to be a data generator itself, but it could be a tuple of Numpy arrays as well. If you pass a generator as validation_data, then this generator is expected to yield batches of validation data endlessly, and thus you should also specify the validation_steps argument, which tells the process how many batches to draw from the validation generator for evaluation.
# + id="yWxJIby3E_z_" colab_type="code" outputId="090fb59a-3b3a-4eec-e54f-90eb2651dee1" colab={"base_uri": "https://localhost:8080/", "height": 1000}
import pickle
history = model.fit_generator(
train_generator,
steps_per_epoch=100,
epochs=100,
validation_data=validation_generator,
validation_steps=50)
# + [markdown] id="vAhhvutUHH0j" colab_type="text"
# It is good practice to always save your models after training:
# + id="tHB644PwNqxn" colab_type="code" outputId="03ad44c6-bb6c-4148-f559-7f8cfdbd2a8b" colab={"base_uri": "https://localhost:8080/", "height": 34}
#model.save('/content/gdrive/My\ Drive/dogs-vs-cats/cats_and_dogs_small_1.h5')
#model.load_weights('/content/gdrive/My\ Drive/dogs-vs-cats/cats_and_dogs_small_1.h5')
model.save('cats_and_dogs_small_1.h5')
model.load_weights('cats_and_dogs_small_1.h5')
scores = model.evaluate_generator(validation_generator, 100)
print("%s: %.2f%%" % (model.metrics_names[1], scores[1]*100))
# + [markdown] id="6xfHjdPlHUUk" colab_type="text"
# Let's plot the loss and accuracy of the model over the training and validation data during training:
# + id="uqqJBlZ9OA0s" colab_type="code" outputId="11342228-b85b-4755-a9a0-6ce89c18564d" colab={"base_uri": "https://localhost:8080/", "height": 545}
import matplotlib.pyplot as plt
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(acc))
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
# + [markdown] id="kRYfK_uFHcft" colab_type="text"
# These plots are characteristic of overfitting. Our training accuracy increases linearly over time, until it reaches nearly 100%, while our validation accuracy stalls at 70-72%. Our validation loss reaches its minimum after only five epochs then stalls, while the training loss keeps decreasing linearly until it reaches nearly 0.
#
# Because we only have relatively few training samples (2000), overfitting is going to be our number one concern. You already know about a number of techniques that can help mitigate overfitting, such as dropout and weight decay (L2 regularization). We are now going to introduce a new one, specific to computer vision, and used almost universally when processing images with deep learning models: data augmentation.
# + id="G3Fi2zP1OGEY" colab_type="code" colab={}
# for item in globals():
# if 'istory' in item:
# print(item)
# + id="C2bMCjPtOsSy" colab_type="code" colab={}
# globals()['history']
# + [markdown] id="agjuULZDUEjH" colab_type="text"
# ### Using data augmentation
#
# Overfitting is caused by having too few samples to learn from, rendering us unable to train a model able to generalize to new data. Given infinite data, our model would be exposed to every possible aspect of the data distribution at hand: we would never overfit. Data augmentation takes the approach of generating more training data from existing training samples, by "augmenting" the samples via a number of random transformations that yield believable-looking images. The goal is that at training time, our model would never see the exact same picture twice. This helps the model get exposed to more aspects of the data and generalize better.
#
# In Keras, this can be done by configuring a number of random transformations to be performed on the images read by our ImageDataGenerator instance.
# + id="CH_sgQTBPDiT" colab_type="code" colab={}
datagen = ImageDataGenerator(
rotation_range=40,
width_shift_range=0.2,
height_shift_range=0.2,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True,
fill_mode='nearest')
# + [markdown] id="4y1r9nHGr9u_" colab_type="text"
# These are just a few of the options available (for more, see the Keras documentation). Let's quickly go over what we just wrote:
#
# * rotation_range is a value in degrees (0-180), a range within which to randomly rotate pictures.
# * width_shift and height_shift are ranges (as a fraction of total width or height) within which to randomly translate pictures vertically or horizontally.
# * shear_range is for randomly applying shearing transformations.
# * zoom_range is for randomly zooming inside pictures.
# * horizontal_flip is for randomly flipping half of the images horizontally -- relevant when there are no assumptions of horizontal asymmetry (e.g. real-world pictures).
# * fill_mode is the strategy used for filling in newly created pixels, which can appear after a rotation or a width/height shift.
#
# Let's take a look at our augmented images:
# + id="VW7KagxpUKFm" colab_type="code" outputId="bca43c89-cd44-4b83-f66b-9e0a2f1a4152" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# This is module with image preprocessing utilities
from keras.preprocessing import image
fnames = [os.path.join(train_cats_dir, fname) for fname in os.listdir(train_cats_dir)]
# We pick one image to "augment"
img_path = fnames[3]
# Read the image and resize it
img = image.load_img(img_path, target_size=(150, 150))
# Convert it to a Numpy array with shape (150, 150, 3)
x = image.img_to_array(img)
# Reshape it to (1, 150, 150, 3)
x = x.reshape((1,) + x.shape)
# The .flow() command below generates batches of randomly transformed images.
# It will loop indefinitely, so we need to `break` the loop at some point!
i = 0
for batch in datagen.flow(x, batch_size=1):
plt.figure(i)
imgplot = plt.imshow(image.array_to_img(batch[0]))
i += 1
if i % 4 == 0:
break
plt.show()
# + [markdown] id="Ku--F_J5smMh" colab_type="text"
# If we train a new network using this data augmentation configuration, our network will never see twice the same input. However, the inputs that it sees are still heavily intercorrelated, since they come from a small number of original images -- we cannot produce new information, we can only remix existing information. As such, this might not be quite enough to completely get rid of overfitting. To further fight overfitting, we will also add a Dropout layer to our model, right before the densely-connected classifier:
# + id="ZUMrd6QAVxV6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 122} outputId="eaffa285-6abd-4557-cf56-63a9a17f1d69"
model = models.Sequential()
model.add(layers.Conv2D(32, (3, 3), activation='relu',
input_shape=(150, 150, 3)))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Conv2D(64, (3, 3), activation='relu'))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Conv2D(128, (3, 3), activation='relu'))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Conv2D(128, (3, 3), activation='relu'))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Flatten())
model.add(layers.Dropout(0.5))
model.add(layers.Dense(512, activation='relu'))
model.add(layers.Dense(1, activation='sigmoid'))
model.compile(loss='binary_crossentropy',
optimizer=optimizers.RMSprop(lr=1e-4),
metrics=['acc'])
# + [markdown] id="yu4jrttGsuMS" colab_type="text"
# Let's train our network using data augmentation and dropout:
# + id="TF6sGR8mWQCd" colab_type="code" outputId="099d72d6-8556-41e6-a3e6-7bf968fa2ee1" colab={"base_uri": "https://localhost:8080/", "height": 1000}
train_datagen = ImageDataGenerator(
rescale=1./255,
rotation_range=40,
width_shift_range=0.2,
height_shift_range=0.2,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True,)
# Note that the validation data should not be augmented!
test_datagen = ImageDataGenerator(rescale=1./255)
train_generator = train_datagen.flow_from_directory(
# This is the target directory
train_dir,
# All images will be resized to 150x150
target_size=(150, 150),
batch_size=32,
# Since we use binary_crossentropy loss, we need binary labels
class_mode='binary')
validation_generator = test_datagen.flow_from_directory(
validation_dir,
target_size=(150, 150),
batch_size=32,
class_mode='binary')
history = model.fit_generator(
train_generator,
steps_per_epoch=100,
epochs=100,
validation_data=validation_generator,
validation_steps=50)
# + id="EipWCa82WTrN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="0a617056-5fcf-46d8-fc1e-4a09e02c722a"
model.save('cats_and_dogs_small_2.h5')
model.load_weights('cats_and_dogs_small_2.h5')
scores = model.evaluate_generator(validation_generator, 100)
# workers=12)
print("%s: %.2f%%" % (model.metrics_names[1], scores[1]*100))
# + id="ivdG9Y7gisqW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 545} outputId="09039842-1825-45c8-8a00-fd91c20d49a7"
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(acc))
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
# + id="P93aFFM5yXz3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="1bb61e7b-34c9-4823-bb0e-ccd1be8d3ad0"
# # !mkdir gdrive/My\ Drive/DeepLearningAndNN/models
# model.save('gdrive/My Drive/DeepLearningAndNN/models/dropout.h5')
# model.load_weights('gdrive/My Drive/DeepLearningAndNN/models/dropout.h5')
scores = model.evaluate_generator(validation_generator, 100)
# workers=12)
print("%s: %.2f%%" % (model.metrics_names[1], scores[1]*100))
# + [markdown] id="P9_QeBmTtD7G" colab_type="text"
# Thanks to data augmentation and dropout, we are no longer overfitting: the training curves are rather closely tracking the validation curves. We are now able to reach an accuracy of 82%, a 15% relative improvement over the non-regularized model.
#
# By leveraging regularization techniques even further and by tuning the network's parameters (such as the number of filters per convolution layer, or the number of layers in the network), we may be able to get an even better accuracy, likely up to 86-87%. However, it would prove very difficult to go any higher just by training our own convnet from scratch, simply because we have so little data to work with. As a next step to improve our accuracy on this problem, we will have to leverage a pre-trained model, which will be the focus of the next two sections.
# + [markdown] id="WwnbCLVjjuTv" colab_type="text"
# # Using a pre-trained convnet
#
# Let's instantiate the VGG16 model:
#
#
# + id="WcDU_VeTi-Sk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="fddd5131-56f2-4e97-d75c-8c8b4aed9f48"
from keras.applications import VGG16
conv_base = VGG16(weights='imagenet',
include_top=False,
input_shape=(150, 150, 3))
# + [markdown] id="74J8ia4S12SK" colab_type="text"
# We passed three arguments to the constructor:
#
# * weights, to specify which weight checkpoint to initialize the model from
# * include_top, which refers to including or not the densely-connected classifier on top of the network. By default, this densely-connected classifier would correspond to the 1000 classes from ImageNet. Since we intend to use our own densely-connected classifier (with only two classes, cat and dog), we don't need to include it.
# * input_shape, the shape of the image tensors that we will feed to the network. This argument is purely optional: if we don't pass it, then the network will be able to process inputs of any size.
#
#
# Here's the detail of the architecture of the VGG16 convolutional base: it's very similar to the simple convnets that you are already familiar with.
# + id="rIsrVPjb1Taa" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 799} outputId="8f4d1425-8f18-4ba9-a1d1-f2c04a01761b"
conv_base.summary()
# + [markdown] id="4o6EUria2sXC" colab_type="text"
# The final feature map has shape (4, 4, 512). That's the feature on top of which we will stick a densely-connected classifier.
#
# At this point, there are two ways we could proceed:
#
# * Running the convolutional base over our dataset, recording its output to a Numpy array on disk, then using this data as input to a standalone densely-connected classifier similar to those you have seen in the first chapters of this book. This solution is very fast and cheap to run, because it only requires running the convolutional base once for every input image, and the convolutional base is by far the most expensive part of the pipeline. However, for the exact same reason, this technique would not allow us to leverage data augmentation at all.
# * Extending the model we have (conv_base) by adding Dense layers on top, and running the whole thing end-to-end on the input data. This allows us to use data augmentation, because every input image is going through the convolutional base every time it is seen by the model. However, for this same reason, this technique is far more expensive than the first one.
# + id="h-NsY-Oe2QJO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="346687ef-348c-4369-db8a-140e7b2b07c4"
import os
import numpy as np
from keras.preprocessing.image import ImageDataGenerator
# base_dir = '/Users/fchollet/Downloads/cats_and_dogs_small'
# base_dir = './'
base_dir = 'small'
train_dir = os.path.join(base_dir, 'train')
validation_dir = os.path.join(base_dir, 'validation')
test_dir = os.path.join(base_dir, 'test')
# print(train_dir)
# print(validation_dir)
# print(test_dir)
datagen = ImageDataGenerator(rescale=1./255)
batch_size = 20
def extract_features(directory, sample_count):
features = np.zeros(shape=(sample_count, 4, 4, 512))
labels = np.zeros(shape=(sample_count))
generator = datagen.flow_from_directory(
directory,
target_size=(150, 150),
batch_size=batch_size,
class_mode='binary')
i = 0
for inputs_batch, labels_batch in generator:
features_batch = conv_base.predict(inputs_batch)
features[i * batch_size : (i + 1) * batch_size] = features_batch
labels[i * batch_size : (i + 1) * batch_size] = labels_batch
i += 1
if i * batch_size >= sample_count:
# Note that since generators yield data indefinitely in a loop,
# we must `break` after every image has been seen once.
break
return features, labels
train_features, train_labels = extract_features(train_dir, 2000)
validation_features, validation_labels = extract_features(validation_dir, 1000)
test_features, test_labels = extract_features(test_dir, 1000)
# + id="hC9X9x3PGO56" colab_type="code" colab={}
train_features = np.reshape(train_features, (2000, 4 * 4 * 512))
validation_features = np.reshape(validation_features, (1000, 4 * 4 * 512))
test_features = np.reshape(test_features, (1000, 4 * 4 * 512))
# + id="ygaTVmwUOrj4" colab_type="code" outputId="1057c3f0-2ce3-4a85-e62a-53af18fdd5f1" colab={"base_uri": "https://localhost:8080/", "height": 1000}
from keras import models
from keras import layers
from keras import optimizers
model = models.Sequential()
model.add(layers.Dense(256, activation='relu', input_dim=4 * 4 * 512))
model.add(layers.Dropout(0.5))
model.add(layers.Dense(1, activation='sigmoid'))
model.compile(optimizer=optimizers.RMSprop(lr=2e-5),
loss='binary_crossentropy',
metrics=['acc'])
history = model.fit(train_features, train_labels,
epochs=30,
batch_size=20,
validation_data=(validation_features, validation_labels))
# + id="_BzK19bCOvHF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 545} outputId="6bf72306-8c02-4b2b-8d4f-a11f738f18cc"
import matplotlib.pyplot as plt
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(acc))
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
# + [markdown] id="0Iz_8BUsQevD" colab_type="text"
# ### the second technique we mentioned for doing feature extraction,
# which is much slower and more expensive, but which allows us to leverage data augmentation during training: extending the conv_base model and running it end-to-end on the inputs.
#
# Note that this technique is in fact so expensive that you should only attempt it if you have access to a GPU: it is absolutely intractable on CPU. If you cannot run your code on GPU, then the previous technique is the way to go.
# + id="5F7owL1yPEYS" colab_type="code" colab={}
from keras import models
from keras import layers
model = models.Sequential()
model.add(conv_base)
model.add(layers.Flatten())
model.add(layers.Dense(256, activation='relu'))
model.add(layers.Dense(1, activation='sigmoid'))
# + id="IXCTIW9HQz-Z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 289} outputId="048f69c9-0012-4255-a068-e33da9b7d228"
model.summary()
# + [markdown] id="jQZSaBVyRl-J" colab_type="text"
# Before we compile and train our model, a very important thing to do is to freeze the convolutional base. "Freezing" a layer or set of layers means preventing their weights from getting updated during training.
#
# In Keras, freezing a network is done by setting its trainable attribute to False:
# + id="Cw6WbDUPQ23g" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="2f7b69f7-bac3-4aaa-c704-a65dffcd0d0e"
print('This is the number of trainable weights '
'before freezing the conv base:', len(model.trainable_weights))
# + id="l_uPjb31R1sG" colab_type="code" colab={}
conv_base.trainable = False
# + id="xIMvhNGCR31c" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="98c3a734-f395-4b62-c7ee-36ce087d2620"
print('This is the number of trainable weights '
'after freezing the conv base:', len(model.trainable_weights))
# + id="UKgaclo7R541" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 545} outputId="c29220fe-7c18-47f4-b6f6-03dc735e9c73"
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(acc))
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
# + [markdown] id="DKYFjzBDY6oY" colab_type="text"
# Now we can start training our model, with the same data augmentation configuration that we used in our previous example:
# + id="hxD3ZrXDSjGb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="2215811a-71f9-4dc2-ae80-3fbf204255e2"
from keras.preprocessing.image import ImageDataGenerator
train_datagen = ImageDataGenerator(
rescale=1./255,
rotation_range=40,
width_shift_range=0.2,
height_shift_range=0.2,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True,
fill_mode='nearest')
# Note that the validation data should not be augmented!
test_datagen = ImageDataGenerator(rescale=1./255)
train_generator = train_datagen.flow_from_directory(
# This is the target directory
train_dir,
# All images will be resized to 150x150
target_size=(150, 150),
batch_size=20,
# Since we use binary_crossentropy loss, we need binary labels
class_mode='binary')
validation_generator = test_datagen.flow_from_directory(
validation_dir,
target_size=(150, 150),
batch_size=20,
class_mode='binary')
model.compile(loss='binary_crossentropy',
optimizer=optimizers.RMSprop(lr=2e-5),
metrics=['acc'])
# history = model.fit_generator(
# train_generator,
# steps_per_epoch=100,
# epochs=30,
# validation_data=validation_generator,
# validation_steps=50,
# verbose=2)
history = model.fit_generator(
train_generator,
steps_per_epoch=100,
epochs=30,
validation_data=validation_generator,
validation_steps=50)
# + id="UEuYIWRBY-Xj" colab_type="code" colab={}
model.save('cats_and_dogs_small_3.h5')
# + id="at8No1ll3eLx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 545} outputId="510d409d-fd30-4a08-f4c7-3f13b35d2e9c"
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(acc))
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
# + id="iGHRyhit3iMb" colab_type="code" colab={}
conv_base.trainable = True
set_trainable = False
for layer in conv_base.layers:
if layer.name == 'block5_conv1':
set_trainable = True
if set_trainable:
layer.trainable = True
else:
layer.trainable = False
# + id="dVo4FnCb3qdX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="4d02c4c8-273c-4952-a525-beb7291a0545"
model.compile(loss='binary_crossentropy',
optimizer=optimizers.RMSprop(lr=1e-5),
metrics=['acc'])
history = model.fit_generator(
train_generator,
steps_per_epoch=100,
epochs=100,
validation_data=validation_generator,
validation_steps=50)
# + id="kc2jgJYu3zJ1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c419d084-80be-474f-d1cd-433a34786aea"
model.save('cats_and_dogs_small_4.h5')
model.load_weights('cats_and_dogs_small_4.h5')
scores = model.evaluate_generator(validation_generator, 100)
print("%s: %.2f%%" % (model.metrics_names[1], scores[1]*100))
# + id="MIbJrZwAvurG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 545} outputId="a3b4426e-0879-4a32-939c-0ea4416a141a"
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(acc))
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
# + id="zcVNADvavwz9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 545} outputId="967c2d43-8f8a-4bd7-c6ab-f4d86fd6c679"
def smooth_curve(points, factor=0.8):
smoothed_points = []
for point in points:
if smoothed_points:
previous = smoothed_points[-1]
smoothed_points.append(previous * factor + point * (1 - factor))
else:
smoothed_points.append(point)
return smoothed_points
plt.plot(epochs,
smooth_curve(acc), 'bo', label='Smoothed training acc')
plt.plot(epochs,
smooth_curve(val_acc), 'b', label='Smoothed validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
plt.plot(epochs,
smooth_curve(loss), 'bo', label='Smoothed training loss')
plt.plot(epochs,
smooth_curve(val_loss), 'b', label='Smoothed validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
# + id="e4L3gkX1v3DM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="18ffab0a-9ece-4bb3-a713-506902036ac3"
test_generator = test_datagen.flow_from_directory(
test_dir,
target_size=(150, 150),
batch_size=20,
class_mode='binary')
test_loss, test_acc = model.evaluate_generator(test_generator, steps=50)
print('test acc:', test_acc)
# + id="cNxa2R71v9cd" colab_type="code" colab={}
| Python_Deep_Learning_And_Fine_Tuning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.2 64-bit
# language: python
# name: python3
# ---
# --- Welcome to Project Better Help! ---
#
# Background
#
# BetterHelp is a mobile app that I have always wondered if their way of (virtual therapy sessions and ability to reach out to your therapist instantly via messaging in crisis situations) delivering psychological therapy really works -- even after post-covid times?
#
# So, I took the liberty to analyse what their customers say.
#
#
# I start by scraping Better Help's most recent (starting in the beginning of 2022) Appstore and Google Play reviews and write them into a file.
#download and import necessary modules
import sys
# !{sys.executable} -m pip install google-play-scraper
# !{sys.executable} -m pip install app_store_scraper
from app_store_scraper import AppStore
import pandas as pd
import numpy as np
import json
import datetime
#Appstore scraping
betterhelp_appstore = AppStore(country='us', app_name='betterhelp-therapy', app_id = '995252384')
betterhelp_appstore.review(after= datetime.datetime(2021, 12, 31)) #retrieve most recent reviews from beginning of 2022
display(betterhelp_appstore[:5])
betterhelp_appstore = pd.DataFrame(betterhelp_appstore.reviews)
betterhelp_appstore.columns
keep_col = ['rating','review','date']
betterhelp_appstore = betterhelp_appstore[keep_col]
betterhelp_appstore = betterhelp_appstore.sort_values(by = 'date')
display(betterhelp_appstore[:5])
#Google Play scraping
from google_play_scraper import Sort, reviews_all
betterhelp_googleplay = reviews_all('com.betterhelp', Sort.NEWEST)
betterhelp_googleplay = pd.DataFrame.from_dict(betterhelp_googleplay, orient='columns')
betterhelp_googleplay = betterhelp_googleplay[betterhelp_googleplay['at'] > '2021-12-31'] #retrieve comments starting from 2022
keep_col = ['score','content','at']
betterhelp_googleplay = betterhelp_googleplay[keep_col]
betterhelp_googleplay = betterhelp_googleplay.sort_values(by = 'at')
display(betterhelp_googleplay[:5])
| Project-BetterHelp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
# -
# ## Exercise 1
# - load the dataset: `../data/international-airline-passengers.csv`
# - inspect it using the `.info()` and `.head()` commands
# - use the function `pd.to_datetime()` to change the column type of 'Month' to a datatime type
# - set the index of df to be a datetime index using the column 'Month' and the `df.set_index()` method
# - choose the appropriate plot and display the data
# - choose appropriate scale
# - label the axes
# - load the dataset: ../data/international-airline-passengers.csv
df = pd.read_csv('../data/international-airline-passengers.csv')
# - inspect it using the .info() and .head() commands
df.info()
df.head()
# +
# - use the function to_datetime() to change the column type of 'Month' to a datatime type
# - set the index of df to be a datetime index using the column 'Month' and tthe set_index() method
df['Month'] = pd.to_datetime(df['Month'])
df = df.set_index('Month')
# -
df.head()
# +
# - choose the appropriate plot and display the data
# - choose appropriate scale
# - label the axes
df.plot()
# -
# ## Exercise 2
# - load the dataset: `../data/weight-height.csv`
# - inspect it
# - plot it using a scatter plot with Weight as a function of Height
# - plot the male and female populations with 2 different colors on a new scatter plot
# - remember to label the axes
# - load the dataset: ../data/weight-height.csv
# - inspect it
df = pd.read_csv('../data/weight-height.csv')
df.head()
df.info()
df.describe()
df['Gender'].value_counts()
# - plot it using a scatter plot with Weight as a function of Height
_ = df.plot(kind='scatter', x='Height', y='Weight')
# +
# - plot the male and female populations with 2 different colors on a new scatter plot
# - remember to label the axes
# this can be done in several ways, showing 2 here:
males = df[df['Gender'] == 'Male']
females = df.query('Gender == "Female"')
fig, ax = plt.subplots()
males.plot(kind='scatter', x='Height', y='Weight',
ax=ax, color='blue', alpha=0.3,
title='Male & Female Populations')
females.plot(kind='scatter', x='Height', y='Weight',
ax=ax, color='red', alpha=0.3)
# -
df['Gendercolor'] = df['Gender'].map({'Male': 'blue', 'Female': 'red'})
df.head()
df.plot(kind='scatter',
x='Height',
y='Weight',
c=df['Gendercolor'],
alpha=0.3,
title='Male & Female Populations')
fig, ax = plt.subplots()
ax.plot(males['Height'], males['Weight'], 'ob',
females['Height'], females['Weight'], 'or', alpha=0.3)
plt.xlabel('Height')
plt.ylabel('Weight')
plt.title('Male & Female Populations')
# ## Exercise 3
# - plot the histogram of the heights for males and for females on the same plot
# - use alpha to control transparency in the plot comand
# - plot a vertical line at the mean of each population using `plt.axvline()`
# +
males['Height'].plot(kind='hist',
bins=50,
range=(50, 80),
alpha=0.3,
color='blue')
females['Height'].plot(kind='hist',
bins=50,
range=(50, 80),
alpha=0.3,
color='red')
plt.title('Height distribution')
plt.legend(["Males", "Females"])
plt.xlabel("Heigth (in)")
plt.axvline(males['Height'].mean(), color='blue', linewidth=2)
plt.axvline(females['Height'].mean(), color='red', linewidth=2)
# +
males['Height'].plot(kind='hist',
bins=200,
range=(50, 80),
alpha=0.3,
color='blue',
cumulative=True,
normed=True)
females['Height'].plot(kind='hist',
bins=200,
range=(50, 80),
alpha=0.3,
color='red',
cumulative=True,
normed=True)
plt.title('Height distribution')
plt.legend(["Males", "Females"])
plt.xlabel("Heigth (in)")
plt.axhline(0.8)
plt.axhline(0.5)
plt.axhline(0.2)
# -
# ## Exercise 4
# - plot the weights of the males and females using a box plot
# - which one is easier to read?
# - (remember to put in titles, axes and legends)
dfpvt = df.pivot(columns = 'Gender', values = 'Weight')
dfpvt.head()
dfpvt.info()
dfpvt.plot(kind='box')
plt.title('Weight Box Plot')
plt.ylabel("Weight (lbs)")
# ## Exercise 5
# - load the dataset: `../data/titanic-train.csv`
# - learn about scattermatrix here: http://pandas.pydata.org/pandas-docs/stable/visualization.html
# - display the data using a scattermatrix
df = pd.read_csv('../data/titanic-train.csv')
df.head()
from pandas.tools.plotting import scatter_matrix
_ = scatter_matrix(df.drop('PassengerId', axis=1), figsize=(10, 10))
| solutions/2 Data exploration Exercises Solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: ROS Python 2
# language: python
# name: ros_python
# ---
# # Example of MoveIt usage
#
# This is an example of MoveIt
# +
import sys
import copy
import rospy
import rospy as rp
import moveit_commander
import moveit_msgs.msg
import geometry_msgs.msg
from math import pi
from std_msgs.msg import String
from moveit_commander.conversions import pose_to_list
# -
moveit_commander.roscpp_initialize(sys.argv)
rospy.init_node('move_group_python_interface_tutorial',
anonymous=True)
robot = moveit_commander.RobotCommander()
scene = moveit_commander.PlanningSceneInterface()
group = moveit_commander.MoveGroupCommander("arm")
group
group.get_name()
# +
# group.set_end_effector_link('gripper_eef')
# -
group.has_end_effector_link()
print robot.get_current_state()
group_variable_values = group.get_current_joint_values()
group_variable_values
group.set_joint_value_target(group.get_random_joint_values())
plan2 = group.plan()
group.execute(plan2)
group.get_current_pose()
pose_target = geometry_msgs.msg.Pose()
pose_target.orientation.w = 1
pose_target.position.x = 0.5
pose_target.position.y = 0.5
pose_target.position.z = 0.5
group.set_start_state_to_current_state()
group.set_pose_target(pose_target)
planx = group.plan()
# +
from matplotlib import pyplot as plt
# %matplotlib inline
import numpy as np
plt_list = []
for el in planx.joint_trajectory.points:
plt_list.append(el.positions)
for i in range(len(plt_list[0])):
plt.plot(np.asarray(plt_list)[:, i])
plt.show()
# -
# ?group.execute
group.execute(planx)
group.get_random_joint_values()
group.compute_cartesian_path([pose_target], 0.1, 0.1)
| demos/Demo_Notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # This works but I have abandoned these strategies because of catalyst issues. t2 are the most current strategies.
#
# ## Enigma Catalyst Notes
#
# ### Types of runs
#
# - Default: live=false, simulate_orders=false
# - Backtest: live=false, simulate_orders=false, start and end
# - Paper Trading: live=true, simulate_orders=true
# - Live Trading: live=true, simulate_orders=false
#
# ### Integration for future V2 algos
#
# - More parameters instead of hard-coded values in algorithm
# - Including signals and indicators from library for reuse
#
# - https://github.com/enigmampc/catalyst/blob/a062e1826f3e420b24c22776e04650b9e2a0719e/docs/live-trading-blueprint.md
#
# - Maybe go back to my first tactic of using and modifying titan algorithms. To much problems with data strutures
# +
from os import getcwd
import pandas as pd
from kryptobot.portfolio.manager import Manager
config = getcwd() + '/config.json'
manager = Manager(config=config)
# Works with catalyst supported backtest exchanges
params = {
'strategy': 'simple_moving_average',
'algo_namespace': 'simple_moving_average',
'type': 'core',
'default': {
'capital_base': 1000,
'data_frequency': 'daily',
'exchange_name': 'bittrex',
'pair': 'doge_btc',
'start': '2017-06-15',
'end': '2018-06-15',
'profit_target_percentage': 1.1,
'fixed_stoploss_percentage': .95,
'trailing_stoploss_percentage': .90,
'order_quantity': 100,
'position_limit': 1000
},
'custom': {
'long_window': 1,
'short_window': 4
},
'portfolio': {
'name': 'default'
}
}
# Tries to rerun ingest although should have the data
# Maybe related to having asset integer ids instead of string in cryptopia?
# params = {
# 'strategy': 'simple_moving_average',
# 'algo_namespace': 'simple_moving_average',
# 'type': 'core',
# 'default': {
# 'capital_base': 1000,
# 'data_frequency': 'minute',
# 'exchange_name': 'cryptopia',
# 'pair': 'etn_btc',
# 'start': '2017-06-15',
# 'end': '2018-06-15',
# 'profit_target_percentage': 1.1,
# 'fixed_stoploss_percentage': .95,
# 'trailing_stoploss_percentage': .90,
# 'order_quantity': 100,
# 'position_limit': 1000
# },
# 'custom': {
# 'long_window': 30,
# 'short_window': 120
# },
# 'portfolio': {
# 'name': 'default'
# }
# }
# Took a while but ran
# params = {
# 'strategy': 'simple_moving_average',
# 'algo_namespace': 'simple_moving_average',
# 'type': 'core',
# 'default': {
# 'capital_base': 1000,
# 'data_frequency': 'minute',
# 'exchange_name': 'hitbtc',
# 'pair': 'smart_btc',
# 'start': '2017-10-31',
# 'end': '2017-11-09',
# 'profit_target_percentage': 1.1,
# 'fixed_stoploss_percentage': .95,
# 'trailing_stoploss_percentage': .90,
# 'order_quantity': 100,
# 'position_limit': 1000
# },
# 'custom': {
# 'long_window': 30,
# 'short_window': 120
# },
# 'portfolio': {
# 'name': 'default'
# }
# }
# Confusing bcolz errors
# params = {
# 'strategy': 'simple_moving_average',
# 'algo_namespace': 'simple_moving_average',
# 'type': 'core',
# 'default': {
# 'capital_base': 1000,
# 'data_frequency': 'daily',
# 'exchange_name': 'binance',
# 'pair': 'eth_usdt',
# 'start': '2017-08-17',
# 'end': '2018-06-24',
# 'profit_target_percentage': 1.1,
# 'fixed_stoploss_percentage': .95,
# 'trailing_stoploss_percentage': .90,
# 'order_quantity': 100,
# 'position_limit': 1000
# },
# 'custom': {
# 'long_window': 30,
# 'short_window': 120
# },
# 'portfolio': {
# 'name': 'default'
# }
# }
manager.run_strategy(params)
# +
import pandas as pd
data = pd.read_csv('/root/.catalyst/data/csv/binance-daily.csv')
data.head(200)
# +
import pandas as pd
import pickle
from kryptobot.strategies.core.simple_moving_average import SimpleMovingAverage
strategy = SimpleMovingAverage()
analyze = strategy.get_analyze()
# %matplotlib inline
data = pd.read_pickle('/root/.catalyst_pickles/57.pickle')
data.tail(300)
# data.head(300)
# analyze(None, data)
# -
| notebooks/core_strategies.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="ZQM3ihaUPdOc" colab_type="text"
# # **Kaggle Plant Pathology Classification with Xception Model**
# + id="48chSoqxL7qv" colab_type="code" outputId="1a6bab61-1544-4e50-a5b5-f72e0db078a1" executionInfo={"status": "ok", "timestamp": 1589486941069, "user_tz": -120, "elapsed": 699, "user": {"displayName": "<NAME> <NAME>\u00fcller", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GimIm9584b_oU_X3u9Vf3_Qlx6l2UcnDKs7fkmrFQ=s64", "userId": "04601735386306688307"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
from google.colab import drive
drive.mount('/content/drive')
# + id="vz5cHb6ANag3" colab_type="code" colab={}
from __future__ import absolute_import, division, print_function, unicode_literals
import os
import shutil
import cv2
import numpy as np
try:
# Use the %tensorflow_version magic if in colab.
# %tensorflow_version 1.x
except Exception:
pass
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras.layers import Dense, Activation, Input, Dropout, MaxPooling2D, Flatten, BatchNormalization, GaussianNoise, GlobalAveragePooling2D
from tensorflow.keras.models import Model
import matplotlib.pyplot as plt
from tensorflow.keras.applications.xception import Xception
from tensorflow.keras.layers import GlobalAveragePooling2D
from keras.preprocessing.image import ImageDataGenerator
import pandas as pd,numpy as np,pylab as pl
# %matplotlib inline
# + id="0npSV0ymMwRt" colab_type="code" colab={}
#### Trainings DATAFRAME #########
from IPython.display import Image
data_dir = '/content/drive/My Drive/Colab Notebooks/Plant Pathology/'
training_images = '/content/drive/My Drive/Colab Notebooks/Plant Pathology/images/'
traindf=pd.read_csv('/content/drive/My Drive/Colab Notebooks/Plant Pathology/train.csv', dtype=str)
traindf['healthy'] = traindf['healthy'].astype(str)
traindf['multiple_diseases'] = traindf['multiple_diseases'].astype(str)
traindf['rust'] = traindf['rust'].astype(str)
traindf['scab'] = traindf['scab'].astype(str)
def append_ext(fn):
return fn+".jpg"
traindf["image_id"]=traindf["image_id"].apply(append_ext)
# + [markdown] id="9-yXjFlZWeZT" colab_type="text"
# ## **Data preprocessing**
# + id="LJACAXJAeR21" colab_type="code" outputId="926e37aa-d976-4f37-9fcd-d4d63b3934ef" executionInfo={"status": "ok", "timestamp": 1589486942984, "user_tz": -120, "elapsed": 2483, "user": {"displayName": "Julia und <NAME>\u00fcller", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GimIm9584b_oU_X3u9Vf3_Qlx6l2UcnDKs7fkmrFQ=s64", "userId": "04601735386306688307"}} colab={"base_uri": "https://localhost:8080/", "height": 51}
#### DATA PREPROZESSING ####
image_size = 224 # All images will be resized to 224x224
batch_size = 32
train_datagen_I = keras.preprocessing.image.ImageDataGenerator(
rescale=1./255,
rotation_range=0.3,
width_shift_range=0.15,
height_shift_range=0.15,
zoom_range=0.15,
horizontal_flip=True,
vertical_flip=True,
validation_split=0.25
)
validation_datagen_I=ImageDataGenerator(rescale=1./255.,validation_split=0.25)
train_generator_I = train_datagen_I.flow_from_dataframe(dataframe=traindf, directory=training_images,
x_col='image_id',
y_col=['healthy','multiple_diseases','rust','scab'],
target_size=(image_size, image_size),
batch_size=batch_size,
class_mode='raw',
subset='training',
shuffle=True,
seed=7)
validation_generator_I = validation_datagen_I.flow_from_dataframe(dataframe=traindf, directory=training_images,
x_col='image_id',
y_col=['healthy','multiple_diseases','rust','scab'],
target_size=(image_size, image_size),
class_mode='raw',
batch_size=batch_size,
shuffle=True,
subset='validation',
seed=7)
# + [markdown] id="gx2eTOrgWLYt" colab_type="text"
# ### **Show images (optional)**
# + colab_type="code" outputId="077478d6-e45f-4915-fe00-f02918a96ece" executionInfo={"status": "ok", "timestamp": 1589486945833, "user_tz": -120, "elapsed": 5314, "user": {"displayName": "<NAME> <NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GimIm9584b_oU_X3u9Vf3_Qlx6l2UcnDKs7fkmrFQ=s64", "userId": "04601735386306688307"}} id="zVIkAi_XzEsf" colab={"base_uri": "https://localhost:8080/", "height": 729}
#### SHOW AUGMENTED IMAGES ####
import typing
def plot_images_with_labels(
images: typing.List[int], labels: np.ndarray, class_names: typing.List[str]) -> None:
if len(images) != 9:
images = images[:9]
labels = labels[:9]
labels = labels.astype(int)
plt.figure(figsize=(12, 12))
index = 0
for image, label in zip(images, labels):
index += 1
plt.subplot(3, 3, index)
plt.imshow(image)
plt.title("\n Class: " + format(class_names[label.argmax()]) + "\n" + format(label))
plt.axis("off")
# plot some augmented images
X_augmented, y_augmented = next(train_generator_I)
plot_images_with_labels((X_augmented + 1) / 2, y_augmented, ['healthy','multiple_diseases','rust','scab'])
# + [markdown] id="r_GKEpPfaUBT" colab_type="text"
# ## **Transfer learning model with Xception pre-train model and a new Fully-Connected-Classifier**
# * ### with fine-tuning
# * ### with data augmentation
# * ### Optimizer RMSprop(lr=0.0001)
# + colab_type="code" id="qJlQ6jbn99nz" colab={}
#### BILD THE MODEL WITH XCEPTION AND RMSprop OPTIMIZER ####
n_classes = 4
# build a transfer learning model with Xception and a new Fully-Connected-Classifier
base_model = Xception(
weights='imagenet',
include_top=False
)
for layer in base_model.layers:
layer.trainable = True
model = GlobalAveragePooling2D()(base_model.output)
model = Dropout(0.5)(model)
# include new Fully-Connected-Classifier
output_layer = Dense(n_classes, activation='softmax')(model)
# create Model
model = Model(base_model.input, output_layer)
# + [markdown] id="FVJkXLuh_QTY" colab_type="text"
# ### Fit the model
# + id="QqmwprDorjrW" colab_type="code" outputId="2048873a-47b0-41c7-b635-b9efc9c7548e" executionInfo={"status": "ok", "timestamp": 1589489349440, "user_tz": -120, "elapsed": 2408889, "user": {"displayName": "<NAME> <NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GimIm9584b_oU_X3u9Vf3_Qlx6l2UcnDKs7fkmrFQ=s64", "userId": "04601735386306688307"}} colab={"base_uri": "https://localhost:8080/", "height": 1000}
# Saving the model
# Directory where the checkpoints will be saved
checkpoint_dir = data_dir + 'training_checkpoints'
#os.makedirs(checkpoint_dir, exist_ok = True)
# Name of the checkpoint files
checkpoint_prefix = os.path.join(checkpoint_dir, "ckpt_training_xception_accuracy")
checkpoint_callback=tf.keras.callbacks.ModelCheckpoint(
filepath=checkpoint_prefix,
save_weights_only=True)
model.compile(
optimizer=tf.keras.optimizers.RMSprop(lr=0.0001),
loss='categorical_crossentropy',
metrics=['accuracy']
)
steps_per_epoch = train_generator_I.n // batch_size
validation_steps = validation_generator_I.n // batch_size
print(steps_per_epoch)
historyI = model.fit_generator(train_generator_I,
steps_per_epoch = steps_per_epoch,
epochs=25,
validation_data=validation_generator_I,
callbacks=[checkpoint_callback])
# + id="c2t2YIarwmDl" colab_type="code" outputId="87e54e3c-5275-4ff0-f6d0-c24ec39371e3" executionInfo={"status": "ok", "timestamp": 1589489387833, "user_tz": -120, "elapsed": 2447266, "user": {"displayName": "Julia und <NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GimIm9584b_oU_X3u9Vf3_Qlx6l2UcnDKs7fkmrFQ=s64", "userId": "04601735386306688307"}} colab={"base_uri": "https://localhost:8080/", "height": 1000}
#### DATA PREDICTION ####
data_dir = '/content/drive/My Drive/Colab Notebooks/Plant Pathology/'
test_images = '/content/drive/My Drive/Colab Notebooks/Plant Pathology/images/'
#display(Image(training_images + '14.png'))
#display(Image(training_images + 'Test_0.jpg'))
testdf=pd.read_csv('/content/drive/My Drive/Colab Notebooks/Plant Pathology/test.csv', dtype=str)
def append_ext(fn):
return fn+".jpg"
testdf["image_id"]=testdf["image_id"].apply(append_ext)
test_datagen = keras.preprocessing.image.ImageDataGenerator(
rescale=1./255
)
image_size = 224 # All images will be resized to 224x224
batch_size = 1
test_generator = test_datagen.flow_from_dataframe(dataframe=testdf, directory=test_images,
x_col='image_id',
target_size=(image_size, image_size),
batch_size=batch_size,
class_mode=None,
shuffle=False)
filenames = test_generator.filenames
nb_samples = len(filenames)
steps = test_generator.n // batch_size
predict = model.predict_generator(test_generator, 100)
#np.savetxt(data_dir + "predict_test.csv", predict, fmt="%10.3f")
np.set_printoptions(formatter={'float': lambda x: "{0:0.3f}".format(x)})
print("Accuracy = ", predict)
# + id="NrEeEA2TzSem" colab_type="code" colab={}
def plot_learning_curve(
title: str, x: int, y: int, y_test: int, ylim: float = 0.6
) -> None:
plt.figure()
plt.title(title)
axes = plt.gca()
axes.set_ylim([ylim, 1])
plt.xlabel("Epoch")
plt.ylabel("Accuracy")
train_sizes = x
train_scores = y
test_scores = y_test
plt.grid()
plt.plot(
train_sizes,
train_scores,
"o-",
color=(177 / 255, 6 / 255, 58 / 255),
label="Training accuracy",
)
plt.plot(
train_sizes,
test_scores,
"o-",
color=(246 / 255, 168 / 255, 0),
label="Validation accuracy",
)
plt.legend(loc="best")
def plot_history(title: str, history: "History", ylim: float = 0.6) -> None:
y = history.history["acc"]
y_test = history.history["val_acc"]
plot_learning_curve(title, np.arange(1, 1 + len(y)), y, y_test, ylim)
# + id="3XLtWagu3s7b" colab_type="code" outputId="913746b4-8ee5-46b2-e952-226bcb49d853" executionInfo={"status": "ok", "timestamp": 1589489387836, "user_tz": -120, "elapsed": 2447245, "user": {"displayName": "Julia und <NAME>\u00fcller", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GimIm9584b_oU_X3u9Vf3_Qlx6l2UcnDKs7fkmrFQ=s64", "userId": "04601735386306688307"}} colab={"base_uri": "https://localhost:8080/", "height": 295}
plot_history('Performance with fine-tuning and data augmentation', historyI, 0)
#plot_history('Pretrain model performance with fine-tuning with data augmentation', historyII, 0)
#plot_history('Pretrain model performance with fine-tuning without data augmentation', historyIII, 0)
# + [markdown] id="U7ebvPW88mS4" colab_type="text"
# ## Summary
# ### I chose Xception as my pre-trained model. Without fine-tuning, the results were not at all satisfactory - accuracy about 0.5. It didn't matter which optimizer I used or how the learning rate was set.
# ### The breakthrough came through fine-tuning. The RMSprop Optimizer was able to achieve better results than Adam or SGD - Accuracy 0.92-0.93
| ADL_JupyterNotebooks/Practical Project/Alpha Version/Kaggle_Plant_Pathology_Classification_Accurycy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Email Network creation to excel
import pandas as pd
import numpy as np
import networkx as nx
from networkx.algorithms import bipartite
#Load Edge List
file = '../data/initial/email-edges.txt'
edge_list = pd.read_table(file, delim_whitespace=True, names=('Senders', 'Recievers'))
# This is a randomly picked data set taken from: (<NAME>, Danon, L, Diaz-Guilera, A, Giralt, <NAME>.
# Phys. Rev. E 68 , art. no. 065103 (2003).) in order to ilustrate a simple Network Analysis retrieving general measures from it
edge_list.head(5)
len(edge_list['Recievers'].unique())
len(edge_list['Recievers'])
# # Creation of the Graph
G = nx.read_edgelist(file)
len(G.nodes())
len(G.edges())
# The library that we are using avoid repetition of edges, therefore if Alice send a message to Bob, the edge is already created an it will not count the entry 'Bob send a message to Alice'. That explains the difference between the lenght of the edgelist (edge_list) and the Graph (G)
# We could create the graph using the edge_list to confirm it
#Creating the Graph2
G2 = nx.Graph()
G2.add_nodes_from(edge_list['Senders'].unique())
G2.add_edges_from(
[(row['Recievers'], row['Senders']) for idx, row in edge_list.iterrows()])
len(G2.nodes())
len(G2.edges())
# # General Network Stats
# +
Connected_Network = nx.is_connected(G)
Density = len(G.edges())*2/(len(G.nodes())*(len(G.nodes())-1))
Number_of_Edges = len(G.edges())
Number_of_Nodes = len(G.nodes())
Separated_Components = nx.number_connected_components(G)
Assortativity = nx.degree_assortativity_coefficient(G)
Clustering = nx.average_clustering(G)
Diameter = nx.diameter(G)
print(
"EMAIL NETWORK" + '\n'
"Connected Network: " + str(Connected_Network) +'\n'
"Number of Nodes: "+ str(Number_of_Nodes) +'\n'
"Number of Edges: "+ str(Number_of_Edges) +'\n'
"Density: "+ str(Density) +'\n'
"Separated Components: "+ str(Separated_Components) +'\n'
"Assortativity: "+ str(Assortativity) + '\n'
"Average Clustering Coefficient: "+ str(Clustering) + '\n'
"Diameter : " + str(Diameter) + '\n'
)
# -
# # Stats per node
# Stats 1: Degree List -- For Excel: df_degrees
degrees = [(node,val) for (node, val) in G.degree()]
df_degrees = pd.DataFrame(degrees)
df_degrees.columns = ["Senders", "Degree"]
df_degrees = df_degrees.set_index("Senders")
# Stats 2: Neighbour Degree -- For Excel: df_nei_degrees
df_nei_degrees = pd.DataFrame(nx.average_neighbor_degree(G).items())
df_nei_degrees.columns = ["Senders", "Neighbour Degree"]
df_nei_degrees = df_nei_degrees.set_index("Senders")
# Stats 3: Clustering -- For Excel: df_cluster_node
cluster_node = nx.clustering(G)
df_cluster_node = pd.DataFrame(cluster_node.items())
df_cluster_node.columns = ["Senders", "Clustering"]
df_cluster_node = df_cluster_node.set_index("Senders")
# Stats 4: Eccentricity -- For Excel: dist
Eccentricity_G = nx.eccentricity(G)
dist=pd.DataFrame(Eccentricity_G.items())
dist.columns=['Senders', 'Eccentricity']
dist=dist.set_index('Senders')
# Stats 5.1: Degree Centrality -- For Excel: df_degree_centrality
degree_centrality=nx.degree_centrality(G)
df_degree_centrality=pd.DataFrame(degree_centrality.items())
df_degree_centrality.columns=["Senders", "Degree Centrality"]
df_degree_centrality=df_degree_centrality.set_index("Senders")
# Stats 5.2: Closeness Centrality -- For Excel: df_closeness_centrality
closeness_centrality=nx.closeness_centrality(G)
df_closeness_centrality=pd.DataFrame(closeness_centrality.items())
df_closeness_centrality.columns=["Senders", "Closeness"]
df_closeness_centrality=df_closeness_centrality.set_index("Senders")
# Stats 5.3: Betweeness Centrality -- For Excel: df_betweeness_centrality
betweeness_centrality=nx.betweenness_centrality(G)
df_betweeness_centrality=pd.DataFrame(betweeness_centrality.items())
df_betweeness_centrality.columns=["Senders", "Betweeness"]
df_betweeness_centrality=df_betweeness_centrality.set_index("Senders")
# Stats 5.4: Random Walk Centrality -- For Excel: df_rw_centrality
rw_centrality=nx.current_flow_betweenness_centrality(G)
df_rw_centrality=pd.DataFrame(rw_centrality.items())
df_rw_centrality.columns=["Senders", "RandomWalk"]
df_rw_centrality=df_rw_centrality.set_index("Senders")
# Final Excel Creation
df_full_excel = pd.concat([
df_degrees,
df_nei_degrees,
df_cluster_node,
dist,
df_degree_centrality,
df_closeness_centrality,
df_betweeness_centrality,
df_rw_centrality
],
axis=1)
file = '../data/output/Email_Network_Stats.xlsx'
writer = pd.ExcelWriter(file)
df_full_excel.to_excel(writer)
writer.save()
| jupyter-notebook/Email_Network_and_Stats.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# #Getting started with Theano
# This lab was created by <NAME>
# The following timer counts down to a five minute warning before the lab instance shuts down. You should get a pop up at the five minute warning reminding you to save your work! If you are about to run out of time, please see the [Post-Lab](#Post-Lab-Summary) section for saving this lab to view offline later.
# <iframe id="timer" src="timer/timer.html" width="100%" height="120px"></iframe>
# ---
# Before we begin, let's verify [WebSockets](http://en.wikipedia.org/wiki/WebSocket) are working on your system. To do this, execute the cell block below by giving it focus (clicking on it with your mouse), and hitting Ctrl-Enter, or pressing the play button in the toolbar above. If all goes well, you should see some output returned below the grey cell. If not, please consult the [Self-paced Lab Troubleshooting FAQ](https://developer.nvidia.com/self-paced-labs-faq#Troubleshooting) to debug the issue.
print "The answer should be three: " + str(1+2)
# Let's execute the cell below to display information about the GPUs running on the server.
# !nvidia-smi
# ## Introduction
#
# Theano is a Python library that allows you to write symbolic mathematical expressions and compile them onto different computational architectures (in particular, CPU and GPU). It was developed by machine learning researchers at the University of Montreal. Its use is not limited to machine learning applications, but it was designed with machine learning in mind. It's especially good for machine learning techniques which are CPU-intensive and benefit from parallelization (e.g. large neural networks).
#
# In this class we will demonstrate Theano's explicit representation of symbolic maths and it's ability to compile that representation into efficient code to be applied to numerical data. We are going to work through one of the most fundamental applications of articial neural networks - the multi-layer perceptron. This example will allow us to see a complete neural network being defined and trained from scratch, but the process we will follow in Theano is essentially the same for some of the much more complicated networks, such as Convolutional Neural Networks (CNNs), that we have seen in our previous classes. We will finish the class by getting you started in extending the multi-layer perceptron example into a CNN.
# ## Key Theano concepts
# We will begin by introducing some important general concepts that form the basis for all computation in Theano.
#
# First we must import Theano into Python. Notice that Theano is using the GPU - there is no more GPU configuration required when using Theano, the GPU will be leveraged for all applicable computations. This was achieved by creating a file `.theanorc` file in our home directory that specified the flag `device=gpu`. This configuration file is referenced by Theano when it is imported into Python.
#
# Execute the cell below to import Theano into Python.
#
# You will know the lab is processing when you see a solid circle in the top-right of the window that looks like this: 
# Otherwise, when it is idle, you will see the following: 
# If you ever feel like a cell has run for to long, you can stop it with the stop button in the toolbar.
# For troubleshooting, please see [Self-paced Lab Troubleshooting FAQ](https://developer.nvidia.com/self-paced-labs-faq#Troubleshooting) to debug the issue.
#
import numpy as np
import theano
# By convention, the tensor submodule is loaded as T
import theano.tensor as T
# ### Symbolic variables
#
# In Theano algorithms are defined symbolically. This means that variables don't have to have explicit values. The `theano.tensor` sub-module has various symbolic variable types. Below is an example of a simple symbolic calculation using scalar variables.
# +
# Define a scalar variable
a = T.scalar('a')
# Define another variable as a squared
b = a**2
# b will also be a theano variable
print type(b)
# We can inspect how b is defined using theano's pretty print function
print theano.pp(b)
# -
# ### Functions
#
# To compute things in Theano using actual numerical values you define symbolic functions.
# We can't compute anything with a, b and c yet.
# We need to define a theano function first.
# The first argument of theano.function defines the inputs
f = theano.function([a],b)
print f(2)
# In some cases you can use a symbolic variable's eval method.
# This can be more concise and convenient that defining a function.
# If it has inputs, the eval method takes a Python dictionary where
# the keys are theano variables and the values are values for those
# variables.
print b.eval({a: 2})
# ### theano.tensor
#
# Theano has variable types for vectors, matrices and tensors. The `theano.tensor` sub-module has various functions for performing operations on these variables.
# +
# Define symbolic matrix and vector variables
A = T.matrix('A')
x = T.vector('x')
# Define a symbolic function using these variables
y = T.dot(A, x)
# Define a theano function to compute y
linear_func = theano.function([A, x], y)
# Supply numeric values for A and x
print linear_func(np.array([[1,2,3],[4,5,6]], dtype=theano.config.floatX),
np.array([1,2,3], dtype=theano.config.floatX))
# -
# `theano.config.floatX` is a configuration variable that Theano also obtains from the `.theanorc` file. Here it is used to specify the numerical type of the variables A and x. If you are using GPU acceleration then `floatX` should be `float32`.
# ### Shared variables
#
# Shared variables are Theano variables that do have an explcit value that can be get/set and is shared across all functions which use the variable.
shared_var = theano.shared(np.array([[1,2],[3,4]], dtype=theano.config.floatX))
print shared_var.type()
print shared_var.get_value()
shared_var.set_value(np.array([[3, 4], [2, 1]], dtype=theano.config.floatX))
print shared_var.get_value()
shared_squared = shared_var**2
# The first argument of theano.function (inputs) tells Theano what the arguments to the compiled function should be.
# Note that because shared_var is shared, it already has a value, so it doesn't need to be an input to the function.
# Therefore, Theano implicitly considers shared_var an input to a function using shared_squared and so we don't need
# to include it in the inputs argument of theano.function.
function_1 = theano.function([], shared_squared)
print function_1()
# ### updates
#
# The value of a shared variable can be updated in a function by using the `updates` argument of `theano.function`.
# We can also update the state of a shared var in a function
subtract = T.matrix('subtract')
# updates takes a dict where keys are shared variables and values are the new value the shared variable should take
# Here, updates will set shared_var = shared_var - subtract
function_2 = theano.function([subtract], shared_var, updates={shared_var: shared_var - subtract})
print "shared_var before subtracting [[1, 1], [1, 1]] using function_2:"
print shared_var.get_value()
# Subtract [[1, 1], [1, 1]] from shared_var
function_2(np.array([[1, 1], [1, 1]], dtype=theano.config.floatX))
print "shared_var after calling function_2:"
print shared_var.get_value()
# Note that this also changes the output of function_1, because shared_var is shared!
print "New output of function_1() (shared_var**2):"
print function_1()
# ## Example 1: Logistic regression
# Now that we have seen the basic building blocks used in Theano we going to see a more practical application. We're going to generate some toy data, simply two classes of points embedded in 2D space and seperated by the line $y=x$.
# +
# Import some necessary Python visualization modules
# %matplotlib inline
import matplotlib.pyplot as plt
from IPython import display
# Generate 50 random 2-D points
x = np.random.random((50,2)).astype('float32')
# If point is above y=x then class is 1,
# otherwise class is 0
y = np.zeros((50,)).astype('int32')
y[x[:,0]>x[:,1]] = 1
# Plot the points
plt.scatter(x[:,0], x[:,1], c=y, s=100)
plt.show()
# -
# Logistic regression is a probabilistic linear classifier. That means it uses a linear model to predict the probability of the class that a data sample belongs to, where a data sample is a real-valued input vector. Logistic regression achieves this by projecting these input vectors onto a set of hyperplanes, each one representing a class, and the distance of the data sample from the hyperplane boundary represents the probability that it is a member of that class.
#
# Logistic regression is parameterized by a weight matrix $W$ and a bias vector $b$. The probability that an input vector $x$ belongs to class $i$ can be written as:
#
# $$\Pr(Y=i \mid x,W,b)=softmax_i(Wx+b)$$
#
# The linear function $Wx+b$ is equivalent to an artificial neural network layer where all input neurons are fully-connected to all output neurons. The number of input neurons is the same as the number of dimensions in the input vector. The dimensions of $W$ determine how many neurons are in the outut layer and this is equal to the number of classes. The $softmax$ function simply "squashes" an arbitrary real-valued vector into a vector of the same dimensions but with values in the range (0,1) - in this example, softmax is applied to the output of $Wx+b$ so that we get probabilities of class membership.
#
# Using Theano to express the mathematics, we define a logistic regression model as a Python class with a method that initializes the parameters as random real-valued numbers in [0,1] and feeds-forward an input vector. We also define a method that will allow us to use the model in it's current state to classify new data samples.
class LogisticRegression(object):
def __init__(self, input, n_in, n_out):
# Initialize pseudo-random number generator
rng = np.random.RandomState(1234)
# Randomly generate weight matrix W
W_values = np.asarray(
rng.uniform(
low=-np.sqrt(6. / (n_in + n_out)),
high=np.sqrt(6. / (n_in + n_out)),
size=(n_in, n_out)
),
dtype=theano.config.floatX
)
self.W = theano.shared(value=W_values, name='W', borrow=True)
# Randomly initialize bias vector b
b_values = np.zeros((n_out,), dtype=theano.config.floatX)
self.b = theano.shared(value=b_values, name='b', borrow=True)
# Define feed-forward function
self.p_y_given_x = T.nnet.softmax(T.dot(input, self.W) + self.b)
# Specify parameters of layer
self.params = [self.W, self.b]
def predict(self, input):
# Function to apply layer feedforward to new input data
return T.nnet.softmax(T.dot(input, self.W) + self.b)
# Notice that we define the two parameters of the logistic regression as Theano shared variables. This declares them both as being symbolic Theano variables that mantain a persistent state throughout training the model. Shared variables must have an explicit value set - they are not purely symbolic.
#
# We can test our new class by passing in a random input vector.
# +
# Define theano shared variables for the input data and labels
shared_x = theano.shared(np.asarray(x, dtype=theano.config.floatX), borrow=True)
shared_y = theano.shared(np.asarray(y, dtype=theano.config.floatX), borrow=True)
shared_y = T.cast(shared_y, 'int32')
# Initialize a logistic regressor with 2-dimensional inputs
# and 2 classes for output with the first row of x as input
lr = LogisticRegression(shared_x[0,:], 2, 2)
# Feed the first data sample from x through the regressor and print output
print lr.p_y_given_x.eval()
# -
# Because the feed-forward step of the logistic regression is just multiplication by a 2-dimensional matrix, we can actually pass into our class a whole matrix of input vectors where each row represents an individual data sample. This is a good example of why GPU acceleration can be of great benefit in neural network training - these types of matrix multiplication commonly occur and can be parallelized.
# Initialize a logistic regressor with 2-dimensional inputs
# and 2 classes for output but now with all of x as input
lr = LogisticRegression(shared_x, 2, 2)
# Print the first 10 data samples from x through the regressor and print output
print lr.p_y_given_x.eval()[:10]
# So far our model is not very useful as all of the parameters have random values. In order to train the parameters of the network we must define a loss function. A loss function measures how close the models predicted classification of a training data sample is to the actual known classification. The training process will attempt to find values of $W$ and $b$ that will minimize this loss value across all of our training data samples. We will use the common negative log-likelihood as the loss. This is equivalent to maximizing the likelihood of the training set $D$ under the model parameterized by $W$ and $b$. The loss is defined as:
#
# $$L=-\sum_{i=0}^{|D|}log(P(Y=y^{(i)}|x^{(i)},W,B))$$
#
# In Theano, we define the negative log loss function in the following way by creating another method operating on our logistic regression class.
# +
def negative_log_likelihood(self, y):
# y corresponds to a vector that gives for each data sample
# the correct label
return -T.mean(T.log(self.p_y_given_x)[T.arange(y.shape[0]), y])
LogisticRegression.negative_log_likelihood = negative_log_likelihood
# -
# To train the model we will use an algorithm called stochastic gradient descent (SGD) with mini-batches (sometimes also called minibatch gradient descent (MGD)). We will not expose the complete details of SGD in this introductory class, but the essential algorithm proceeds as follows. First, we feed-forward a mini-batch of data samples through the model to get probabilistic class predictions. Second, we compute the loss by comparing the predictions to the true class labels we have as part of our dataset. Third, we update the model parameters in such a way that next time we feed-forward the same mini-batch through the model the loss will be lower.
#
# In order to update the parameters we must compute whether to make each one bigger or smaller to reduce the loss. We work this out by computing what are called the *gradients* of the loss function with respect to the parameters. In mathematical terms these are the partial derivatives of the loss function with respect to each of the parameters. Intuitively, these tell you which direction and with what magnitude you must modify each of the parameters to reduce the loss.
#
# Theano makes computing gradients very simple and this is a huge benefit. Due to the symbolic definition of the loss function in terms of the parameters, we can simply call the Theano *grad* function to work out the gradients - no calculus required!
#
# When you execute the cell below you will define a Theano function called `train`. Defining this function causes Theano to compile our symbolic definition of the SGD algorithm into efficient GPU-enabled code that will operate on our numerical input data. Notice that at no point in defining this function does an explicit numerical value need to be assigned to any of our variables - it is more like writing math than code!
# +
# Initialize logistic regression model
lr = LogisticRegression(shared_x, 2, 2)
learning_rate = 0.1
# Allocate symbolic variables for the input data
X = T.matrix('X')
Y = T.ivector('Y')
# Begin training function definition based on symbolic mathematical operations
# Specify loss function
cost = lr.negative_log_likelihood(Y)
# Specify parameters to be updated by SGD algorithm
params = lr.params
# Compute gradients of parameters with respect to loss
grads = T.grad(cost, params)
# Compute parameter updates using SGD with momentum
# We use the values 0.1 for learning rate and 0.9 for momentum
updates = []
for p, g in zip(params, grads):
mparam_i = theano.shared(np.zeros(p.get_value().shape, dtype=theano.config.floatX))
v = 0.9 * mparam_i - learning_rate * g
updates.append((mparam_i, v))
updates.append((p, p + v))
# Define theano function to be compiled to perform training update based on numerical data
train = theano.function(
inputs=[],
outputs=cost,
updates=updates,
givens={
X: shared_x[:],
Y: shared_y[:]
}
)
# -
# When you execute the cell below you will see our logistic regression model train on the randomly generated data. After every 10 training iterations you will see the decision boundary between the two classes updated.
# +
# Initialize decision surface for display
x_min, x_max = x[:, 0].min() - 0.1, x[:, 0].max() + 0.1
y_min, y_max = x[:, 1].min() - 0.1, x[:, 1].max() + 0.1
xx, yy = np.meshgrid(np.arange(x_min, x_max, .01),
np.arange(y_min, y_max, .01))
# Perform 300 training iterations
for i in range(0,300):
# Perform SGD update
cost = train()
# Compute decision surface
Z = np.argmax(lr.predict(np.c_[xx.ravel(), yy.ravel()].astype('float32')).eval(),1)
Z = Z.reshape(xx.shape)
# Every 10 iterations, visualize decision surface
if i%10==0:
display.clear_output(wait=True)
print 'Iteration ' + str(i) + ' of 300'
print 'Cost: ' + str(cost)
plt.contourf(xx, yy, Z, levels=[-1,0,1], colors=['b','r'])
plt.scatter(x[:,0], x[:,1], c=y, s=100)
display.display(plt.gcf())
# -
# We see that we are able to quickly learn a decision boundary between the two classes of points which allows the logistic regression function to accurately classify them. Now let's see what happens if we have a more complicated distribution of points in our two classes.
#
# ####Q #1:
#
# What is the effect of reducing the learning rate value in the train function? Try setting the *learning_rate* variable to 0.001 and then executing the last two cells again.
#
# **A**: See [Answer #1 below](#Answer-#1)
#
# Next we will generate a new set of points where one class lies within the circle defined by $x^2+y^2=0.5$ and the other class lies outside it.
# +
# Generate 100 random points
# Class one is points inside x^2 + y^2 = 0.5
# Class two is points outside x^2 + y^2 = 0.5
x = 2 * np.random.random((100,2)).astype('float32') - 1
y = np.zeros((100,)).astype('int32')
y[np.sum(np.square(x),1) > 0.5] = 1
# Plot the points
plt.scatter(x[:,0], x[:,1], c=y, s=100)
plt.show()
# -
# When you execute the cell below you will define a Theano function called `train`. This function causes Theano to compile our symbolic definition of the SGD algorithm into efficient GPU-enabled code that will operate on our numerical input data.
# +
# Define theano shared variables for the second set of input data and labels
shared_x = theano.shared(np.asarray(x, dtype=theano.config.floatX), borrow=True)
shared_y = theano.shared(np.asarray(y, dtype=theano.config.floatX), borrow=True)
shared_y = T.cast(shared_y, 'int32')
# Initialize logistic regression model
lr = LogisticRegression(shared_x, 2, 2)
# Allocate symbolic variables for the input data
X = T.matrix('X')
Y = T.ivector('Y')
# Specify parameters to be updated by SGD algorithm
params = lr.params
# Begin training function definition based on symbolic mathematical operations
# Specify loss function
cost = lr.negative_log_likelihood(Y)
# Compute gradients of parameters with respect to loss
grads = T.grad(cost, params)
# Compute parameter updates using SGD with momentum
# We use the values 0.1 for learning rate and 0.9 for momentum
updates = []
for p, g in zip(params, grads):
mparam_i = theano.shared(np.zeros(p.get_value().shape, dtype=theano.config.floatX))
v = 0.9 * mparam_i - 0.1 * g
updates.append((mparam_i, v))
updates.append((p, p + v))
# Define theano function to be compiled to perform training update based on numerical data
train = theano.function(
inputs=[],
outputs=cost,
updates=updates,
givens={
X: shared_x[:],
Y: shared_y[:]
}
)
# -
# When you execute the cell below you will see our logistic regression model train on the second, more complicated, set of randomly generated data. After every 10 training iterations you will see the decision boundary between the two classes updated.
# +
# Initialize decision surface for display
x_min, x_max = x[:, 0].min() - 0.1, x[:, 0].max() + 0.1
y_min, y_max = x[:, 1].min() - 0.1, x[:, 1].max() + 0.1
xx, yy = np.meshgrid(np.arange(x_min, x_max, .01),
np.arange(y_min, y_max, .01))
# Perform 300 training iterations
for i in range(0,300):
# Perform SGD update
cost = train()
# Compute decision surface
Z = np.argmax(lr.predict(np.c_[xx.ravel(), yy.ravel()].astype('float32')).eval(),1)
Z = Z.reshape(xx.shape)
# Every 10 iterations, visualize decision surface
if i%10==0:
display.clear_output(wait=True)
print 'Iteration ' + str(i) + ' of 300'
print 'Cost: ' + str(cost)
plt.contourf(xx, yy, Z, levels=[-1,0,1], colors=['b','r'])
plt.scatter(x[:,0], x[:,1], c=y, s=100)
display.display(plt.gcf())
# -
# This time we see that our logistic regression model is unable to find a decision boundary that can accurately seperate the two classes. Due to the random parameter initialization and random data generation the exact outcome will be different each time, but often you will find that the model converges on a decision boundary where all data points get classified in the same class.
#
# The reason this logistic regression model cannot accurately seperate the two classes is because it is limited to only finding a linear function, i.e. a straight line, that can seperate them. In order to better classify this data we must increase the complexity of function that our model can learn - Theano enables us to easily do this as a logical extension to our current model.
# ## Example 2: Multilayer perceptron
# As mentioned earlier, the logistic regression model we defined is equivalent to an artificial neural network with a single layer of input neurons fully connected to a single layer of output neurons. Another name for a network of this type is a *(single-layer) perceptron*. We can extend this simple model to give it the ability to learn more complex non-linear decision boundaries by adding another layer to the neural network between the inputs and the outputs. We call this additional layer a *hidden* layer. When we do so we create what is called a *multilayer perceptron*. The mathematical form of this additional layer is exactly the same as before. The only difference is that we now use a hyperbolic tangent (tanh) activation function in the first hidden layer and keep the softmax activation in the output layer.
class hidden(object):
def __init__(self, input, n_in, n_out):
# Initialize pseudo-random number generator
rng = np.random.RandomState(1234)
# Randomly generate weight matrix W
W_values = np.asarray(
rng.uniform(
low=-np.sqrt(6. / (n_in + n_out)),
high=np.sqrt(6. / (n_in + n_out)),
size=(n_in, n_out)
),
dtype=theano.config.floatX
)
self.W = theano.shared(value=W_values, name='W', borrow=True)
# Randomly initialize bias vector b
b_values = np.zeros((n_out,), dtype=theano.config.floatX)
self.b = theano.shared(value=b_values, name='b', borrow=True)
# Define feed-forward function
self.hidden = T.tanh(T.dot(input, self.W) + self.b)
# Specify parameters of layer
self.params = [self.W, self.b]
def feedforward(self, input):
# Function to apply layer feedforward to new input data
return T.tanh(T.dot(input, self.W) + self.b)
# When you execute the cell below you will define a Theano function called `train`. This function causes Theano to compile our symbolic definition of the SGD algorithm into efficient GPU-enabled code that will operate on our numerical input data.
# +
# Initialize multi-layer perceptron
# i.e., initialize a hidden layer followed by a logistic regression layer
layer0 = hidden(shared_x, 2, 10)
layer1 = LogisticRegression(layer0.hidden, 10, 2)
# Allocate symbolic variables for the input data
X = T.matrix('X')
Y = T.ivector('Y')
# Begin training function definition based on symbolic mathematical operations
# Specify loss function
cost = layer1.negative_log_likelihood(Y)
# Specify parameters to be updated by SGD algorithm
params = layer0.params + layer1.params
# Compute gradients of parameters with respect to loss
grads = T.grad(cost, params)
# Compute parameter updates using SGD with momentum
# We use the values 0.1 for learning rate and 0.9 for momentum
updates = []
for p, g in zip(params, grads):
mparam_i = theano.shared(np.zeros(p.get_value().shape, dtype=theano.config.floatX))
v = 0.9 * mparam_i - 0.1 * g
updates.append((mparam_i, v))
updates.append((p, p + v))
# Define theano function to be compiled to perform training update based on numerical data
train = theano.function(
inputs=[],
outputs=cost,
updates=updates,
givens={
X: shared_x[:],
Y: shared_y[:]
}
)
# -
# When you execute the cell below you will see our multi-layer perceptron model train on the second, more complicated, set of randomly generated data. After every 10 training iterations you will see the decision boundary between the two classes updated.
# +
# Initialize decision surface for display
x_min, x_max = x[:, 0].min() - 0.1, x[:, 0].max() + 0.1
y_min, y_max = x[:, 1].min() - 0.1, x[:, 1].max() + 0.1
xx, yy = np.meshgrid(np.arange(x_min, x_max, .01),
np.arange(y_min, y_max, .01))
# Perform 300 training iterations
for i in range(0,300):
# Perform SGD update
cost = train()
# Compute decision surface
Z = np.argmax(layer1.predict(layer0.feedforward((np.c_[xx.ravel(), yy.ravel()].astype('float32')))).eval(),1)
Z = Z.reshape(xx.shape)
# Every 10 iterations, visualize decision surface
if i%10==0:
display.clear_output(wait=True)
print 'Iteration ' + str(i) + ' of 300'
print 'Cost: ' + str(cost)
plt.contourf(xx, yy, Z, levels=[-1,0,1], colors=['b','r'])
plt.scatter(x[:,0], x[:,1], c=y, s=100)
display.display(plt.gcf())
# -
# We now see that the addition of a second layer in our neural network allows us to learn the more complicated decision boundary required to seperate the two classes.
# ## Recap
#
# We have seen so far that in Theano we can define an artificial neural network layer-by-layer as a sequence of Python classes. We have also seen that we can define the loss function as a method on the final layer's class. We can then define a Theano function to train this network and we can make use of the Theano grad function so that we don't have to analytically work out the gradients of our loss function.
#
# Furthermore, we have seen that Theano's close syntactical resemblance to NumPy makes it very easy to integrate Theano functions into a Python data processing pipeline.
#
# This basic recipe for defining and training artificial neural networks in Theano can be extended to develop the most complicated, cutting edge networks being used in Deep Learning research today. To see some examples of more complicated network types being defined and trained in Theano I recommend you look at the [Deep Learning Tutorials](http://deeplearning.net/tutorial/) made available by the Theano developers.
# ## Example 3: Convolutional layers
#
# The `theano.tensor` sub-module offers many functions for operating on symbolic variables. One such function is called `theano.tensor.signal.conv2d` and it is used for implementing the convolutional layers that are critical in computer vision applications.
#
# `theano.tensor.signal.conv2d` takes two symbolic inputs:
#
# - a 4D tensor corresponding to a mini-batch of input images. The shape of the tensor is: [mini-batch size, number of input feature maps, image height, image width].
# - a 4D tensor corresponding to the weight matrix $W$. The shape of this tensor is: [number of feature maps at layer m, number of feature maps at layer m-1, filter height, filter width].
#
# Below is Theano code for implementing a convolutional layer. The layer has 3 input feature maps, i.e. an RGB image, of size 512x512. We use two convolutional filters with 9x9 receptive fields.
# +
from theano.tensor.nnet import conv
# Initialize 4D tensor for input
input = T.tensor4(name='input')
# Specify height and width of convolutional filters
filter_shape = (9, 9)
# Initialize shared variable for weights
w_shp = (2,3,filter_shape[0],filter_shape[1])
w_bound = np.sqrt(3 * np.prod(filter_shape))
W = theano.shared(np.asarray(
np.random.uniform(
low=-1.0 / w_bound,
high=1.0 / w_bound,
size=w_shp),
dtype=input.dtype), name='W')
# build symbolic expression that computes the convolution
# of the input with the filters in W
conv_out = conv.conv2d(input, W)
# build symbolic expression to apply sigmoid activation function
# NOTE: for simplicity we are ommiting the bias vector that
# would also normally be added
output = T.nnet.sigmoid(conv_out)
# create theano function to compute filtered images
f = theano.function([input], output)
# -
# Let's see what happens when we apply our layer to Lena...
# +
# import the lena image from skimage
import skimage.data
lena = skimage.data.lena()
# put the image in 4D tensor of shape (1,3, height, width)
img = np.asarray(lena, dtype='float32') / 256
img_ = img.transpose(2, 0, 1).reshape(1, 3, 512, 512)
# apply convolutional layer
filtered_img = f(img_)
plt.figure(figsize=(10,5))
# plot the original and first and second components of output
plt.subplot(1, 3, 1); plt.axis('off'); plt.imshow(img)
plt.gray();
# recall that the convOp output (filtered image) is actually a "minibatch",
# of size 1 here, so we take index 0 in the first dimension:
plt.subplot(1, 3, 2); plt.axis('off'); plt.imshow(filtered_img[0, 0, :, :])
plt.subplot(1, 3, 3); plt.axis('off'); plt.imshow(filtered_img[0, 1, :, :])
plt.show()
# -
# Notice that randomly initialized filters act very much like edge detectors.
#
# ####Q #2:
#
# What is the effect of changing the convolutional filter size? Try the values (50, 50) and (2, 2) by modifying the ``filter_shape`` variable and then re-execute the last two cells.
#
# **A**: See [Answer #2 below](#Answer-#2)
# ## Theano based Deep Learning libraries
#
# A number of lightweight Deep Learning focused libraries have been built using Theano. Two of the most popular ones are [Lasagne](https://github.com/Lasagne/Lasagne) and [Keras](https://github.com/fchollet/keras).
#
# The motivation for developing these libraries is to create lightweight interfaces for Theano that are more concise for neural network specific development. In other words, these libraries are less expressive than Theano in general but can enable more rapid development for Deep Learning specific use cases.
#
# Below is a demonstration of how to implement our multi-layer perceptron again, but this time using Keras. Our choice to show Keras is arbitrary, we do not necessarily recommend it over Lasagne or any other similar library - we recommend you learn about each of them to find the best fit for your needs.
# +
# Import required keras sub-modules
from keras.models import Sequential
from keras.layers.core import Dense, Activation
from keras.optimizers import SGD
from keras.utils import np_utils
# Convert training labels to "one-hot" representation
Y = np_utils.to_categorical(y, 2)
# specify mlp model
model = Sequential()
model.add(Dense(2, 10, init='lecun_uniform'))
model.add(Activation('tanh'))
model.add(Dense(10, 2, init='lecun_uniform'))
model.add(Activation('softmax'))
# specify training parameters
sgd = SGD(lr=0.1, momentum=0.9)
model.compile(loss='binary_crossentropy', optimizer=sgd)
# train the model
model.fit(x, Y, batch_size=100, nb_epoch=300, verbose=1)
# -
# Notice that we can define and train the same network in far fewer lines of code.
#
# Below we use the trained Keras model to generate our decision surface. The actual code to use the model to generate new predictions is just one line.
# +
# Initialize decision surface for display
x_min, x_max = x[:, 0].min() - 0.1, x[:, 0].max() + 0.1
y_min, y_max = x[:, 1].min() - 0.1, x[:, 1].max() + 0.1
xx, yy = np.meshgrid(np.arange(x_min, x_max, .01),
np.arange(y_min, y_max, .01))
Z = np.c_[xx.ravel(), yy.ravel()]
# use keras model to make predictions across decision surface
Z = model.predict(Z, batch_size=100,verbose=1)
# Plot decision surface
Z = np.argmax(Z, 1)
Z = Z.reshape(xx.shape)
plt.jet()
plt.contourf(xx, yy, Z)
plt.scatter(x[:,0], x[:,1], c=y, s=100)
plt.show()
# -
# ####Q #3:
#
# What is the effect of changing the batch size to 1 for training? Do this by modifying the ``batch_size`` parameter in ``model.fit(...)`` above.
#
# **A**: See [Answer #3 below](#Answer-#3)
# ## Post-Lab Summary
#
# If you would like to download this lab for later viewing, it is recommended you go to your browsers File menu (not the Jupyter notebook file menu) and save the complete web page. This will ensure the images are copied down as well.
#
# ### More information
#
# For more information on using Theano, visit [http://deeplearning.net/software/theano/](http://deeplearning.net/software/theano/). A description of the framework, how to use it, and plenty of examples similar to this lesson are posted.
#
# To learn more about these other topics, please visit:
# * GPU accelerated machine learning: [http://www.nvidia.com/object/machine-learning.html](http://www.nvidia.com/object/machine-learning.html)
# * Theano: [http://deeplearning.net/software/theano/](http://deeplearning.net/software/theano/)
# * Torch: [http://torch.ch/](http://torch.ch/)
# * DIGITS: [https://developer.nvidia.com/digits](https://developer.nvidia.com/digits)
# * cuDNN: [https://developer.nvidia.com/cudnn](https://developer.nvidia.com/cudnn)
#
# ### Deep Learning Lab Series
#
# Make sure to check out the rest of the classes in this Deep Learning lab series. You can find them [here](https://developer.nvidia.com/deep-learning-courses).
#
# ### Acknowledgements
#
# The logistic regression example used was inspired by <NAME>'s [ConvnetJS](http://cs.stanford.edu/people/karpathy/convnetjs/demo/classify2d.html) demonstration. The "Key Theano concepts" section was largely adapted from <NAME>'s [Theano tutorial](http://nbviewer.ipython.org/github/craffel/theano-tutorial/blob/master/Theano%20Tutorial.ipynb). The "Convolutional layers" section was adapted from the Theano Deep Learning tutorial [here](http://deeplearning.net/tutorial/lenet.html#lenet).
# ## Lab Answers
#
# ### Answer #1
# Reducing the learning rate causes the decision boundary to change much more slowly.
#
# [Return to question](#Q-#1:)
#
# ### Answer #2
# You will see that larger filters caused a more blurred feature map and smaller filters caused a sharper feature map.
#
# [Return to question](#Q-#2:)
#
# ### Answer #3
# Training becomes much slower. This is because we are no longer able to benefit from the speed-up gained by parallelizing the processing of multiple input data samples at the same time on the GPU.
#
# [Return to question](#Q-#3:)
| Getting started with Theano.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # *Catbars* step by step
#
# *Simple bars, four features*
from catbars import Bars
import pandas as pd
df = pd.DataFrame({'numbers' : [1, 10, 2, 3],
'name' : ['one', 'ten', 'two', 'three'],
'parity' : ['odd', 'even', 'even', 'odd'],
'french_name' : ['un', 'dix', 'deux', 'trois']})
df
# # Raw bars
#
# The first feature is the only positional (required) argument. It is a numerical feature.
#
# First bars in the list appear on the top of the figure.
Bars(df['numbers'])
Bars([1e9, 1e3, 1e6, 1])
# ## With ```auto_scale```
# *Catbars* automatically adapts the x scale to the data.
#
# Therefore, if the bar orders of magnitude are too different, the logarithmic scale is used.
Bars([1e9, 1e3, 1e6, 1],
auto_scale = True)
# You can chose the color you want.
Bars(df['numbers'],
default_color = 'purple')
# # Up to three features can be visualized in addition to the first numerical feature
#
# ### Right labels
Bars(df['numbers'],
right_labels = df['name'])
# ### Right and classic labels
Bars(df['numbers'],
right_labels = df['name'],
left_labels = df['french_name'])
# ### Color management (the fourth feature)
#
# If there are more categories than available colors, residual categories are mapped to a default color.
#
# Default available colors (SEE BELOW ```tints``` in the configuration section) are assigned first to the most common categories in the selected slice.
#
# The whole list is considered to do the mapping if ```global_view``` is enabled.
#
# You can override the way **Catbars** manage colors by specifying:
# - the ```color_dic``` argument (a dictionary mapping color categories to actual colors) or
# - the ```tints``` argument (the list of colors used to perform automatic mapping).
#
# Those two ways are mutually exclusive.
Bars(df['numbers'],
right_labels = df['name'],
left_labels = df['french_name'],
colors = df['parity'])
# **The figure automatically adapt to long right labels.**
Bars(df['numbers'],
right_labels = ['one', 'a very long right label', 'two', 'threee'],
left_labels = df['french_name'],
colors = df['parity'])
# Another example for the ```colors```argument.
#
# Moreover, you can add a vertical line.
Bars(df['numbers'],
right_labels = df['name'],
left_labels = df['french_name'],
colors = df['name'],
line_dic = {'number': 5,
'color': 'black',
'label': 'a limit'})
# Colors to use can be specified.
Bars(df['numbers'],
right_labels = df['name'],
left_labels = df['french_name'],
colors = df['name'],
color_dic = {'three': 'red'})
# The default label for residual categories can be overriden.
Bars(df['numbers'],
right_labels = df['name'],
left_labels = df['french_name'],
colors = df['name'],
color_dic = {'three': 'red'},
default_label = 'other numbers',
default_color = 'gray')
# ***
# # Sorting and slicing
#
# The sort is performed on a copy (no side effects) in descending order.
#
# It is *slice-wide* by default (*list-wide* if ```global_view```is enabled).
#
# The ```slice``` tuple is one-based and applied to all features.
#
# #### About the ```'rank'``` option
# - You can use the ```'rank'``` option with ```labels``` or ```left_labels``` to create an index.
# - This index restarts from 1 for the selected slice by default. The index is *list-wide* if ```global_view``` is enabled.
#
Bars(df['numbers'],
right_labels = df['name'],
left_labels = 'rank',
sort = True)
Bars(df['numbers'],
right_labels = df['name'],
left_labels = 'rank',
sort = True,
slice = (3,4))
# **When ```global_view``` is enabled, among other consequences, the data bounds are the whole list bounds.**
#
# This function can be useful to split one long chart into several smaller charts in a consistent way.
Bars(df['numbers'],
right_labels = df['name'],
left_labels = 'rank',
sort = True,
slice = (3,4),
global_view = True)
Bars(df['numbers'],
right_labels = df['name'],
sort = True,
slice = (1,2),
global_view = True)
# **There is another option named ```'proportion'```.**
#
# It computes the proportion of each bar relative to the slice total (or the list total if ```global_view```is enabled).
#
# **This option allows bar charts to replace pie charts.**
Bars(df['numbers'],
left_labels = 'rank',
colors = df['numbers'],
right_labels = 'proportion',
sort = True)
# ***
# # Titles for publication: title, xlabel, ylabel, legend_title
#
# With ```Catbars```, what you see in your notebook is what you get.
#
# Given the selected font sizes and the figure size (```figsize```and ```dpi```), the data space adapt to your labels and title texts.
#
# However, if your texts are definitely too long for the available pixels, you can try to split your texts in lines with ```\n```, you can extend the figure or you can reduce font sizes (SEE the configuraton topic).
#
# **The following figure is the same size than the previous one !**
#
Bars(df['numbers'],
right_labels = df['name'],
left_labels = df['french_name'],
colors = df['parity'],
sort = True,
xlabel = 'Feature 1\nThis feature is a very interesting feature\nthat requires many explanations.',
ylabel = 'Feature 2 and 3\nBars can be tagged with two labels',
title = 'With $\mathtt{Catbars}$,\nyou concatenate bars\n and manage labels',
legend_title = 'Feature 4\nThe legend text can be very long')
# ***
# # Advanced configuration
#
# The ```conf``` class attribute lists all the advanced parameters you can redefine by using simple key word arguments.
#
# If it is not enough, you can rewrite the **conf.py** module which is a decorator for the Matplotlib settings. This file is normally located in the library top level directory (```sys.path``` and the command ```pip show catbars ``` can be useful to locate that directory).
Bars.conf
Bars(df['numbers'],
right_labels = df['name'],
left_labels = df['french_name'],
colors = df['parity'],
sort = True,
xlabel = 'Feature 1\nThis feature is a very interesting feature\nthat requires many explanations',
ylabel = 'Feature 2 and 3\nBars can be tagged with two labels',
title = 'With $\mathtt{Catbars}$,\nyou concatenate bars',
legend_title = 'Feature 4\nThe legend text can be very long',
legend_visible = True,
figsize = (8,9),
pad = 0.15,
title_pad = 0.15,
margin = 0,
tints = ['blue', 'red'],
data_font_size = 15)
# ***
# ## For more information
help(Bars)
| documentation/notebooks/catbars_step_by_step.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <img src="header_anwender.png" align="left"/>
# # Anwendungsbeispiel Import of image data with augmentation and classification
#
# Das Ziel dieses Beispieles ist es die Organisation, den Import und die Vorbereitung von Bilddaten für eine Klassifikation zu erklären. Dabei werden folgende Schritte durchgeführt:
#
# - Dynamisches Laden und entpacken der Bilddaten von einer externen Quelle
# - Review der Organisation auf dem Filesystem
# - Laden der Daten
# - Transformationen
# - Augmentierung
# - Training
# - Analyse
# - Verbesserung
#
# Der verwendete Datensatz heisst caltech101[3] mit 101 Klassen und jeweils 40 bis 800 Bildern pro Klasse. Die Bilder haben 200 - 300 Pixel Auflösung in Farbe.
#
# Quellen für die Beispiele und Daten:
#
# - [1] [https://machinelearningmastery.com/how-to-develop-a-cnn-from-scratch-for-cifar-10-photo-classification/](https://machinelearningmastery.com/how-to-develop-a-cnn-from-scratch-for-cifar-10-photo-classification/)
# - [2] [https://github.com/bhavul/Caltech-101-Object-Classification](https://github.com/bhavul/Caltech-101-Object-Classification)
# - [3] [http://www.vision.caltech.edu/Image_Datasets/Caltech101/](http://www.vision.caltech.edu/Image_Datasets/Caltech101/)
#
#
# +
#
# Abdrehen von Fehlermeldungen
#
from warnings import simplefilter
# ignore all future warnings
simplefilter(action='ignore', category=FutureWarning)
simplefilter(action='ignore', category=Warning)
simplefilter(action='ignore', category=RuntimeWarning)
#
# Import der Module
#
import os
import logging
import tarfile
import operator
import random
from urllib.request import urlretrieve
from PIL import Image
import numpy as np
import matplotlib.pyplot as plt
import sklearn
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import LabelEncoder
#
# Tensorflow und Keras
#
import tensorflow as tf
from tensorflow.keras.utils import to_categorical
from tensorflow.keras.models import Sequential, Model
from tensorflow.keras.layers import Conv2D, Input, Dropout, Activation, Dense, MaxPooling2D, Flatten, GlobalAveragePooling2D
from tensorflow.keras.optimizers import Adadelta
from tensorflow.keras.callbacks import ModelCheckpoint
from tensorflow.keras.callbacks import EarlyStopping
from tensorflow.keras.preprocessing.image import ImageDataGenerator
#
# Für GPU Support
#
tflogger = tf.get_logger()
tflogger.setLevel(logging.ERROR)
tf.compat.v1.logging.set_verbosity(tf.compat.v1.logging.ERROR )
physical_devices = tf.config.list_physical_devices('GPU')
print(physical_devices)
if len(physical_devices) > 0:
tf.config.experimental.set_memory_growth(physical_devices[0], True)
#
# Einstellen der Grösse von Diagrammen
#
plt.rcParams['figure.figsize'] = [16, 9]
#
# Ausgabe der Versionen
#
print('working on keras version {} on tensorflow {} using sklearn {}'.format ( tf.keras.__version__, tf.version.VERSION, sklearn.__version__ ) )
# -
# # Hilfsfunktionen
urlDataSource = 'http://www.vision.caltech.edu/Image_Datasets/Caltech101/101_ObjectCategories.tar.gz'
localExtractionFolder = 'data/caltech101'
localDataArchive = 'data/caltech101/caltech101.tar.gz'
# +
#
# Laden der Daten von einer URL
#
def download_dataset(url,dataset_file_path):
if os.path.exists(localDataArchive):
print("archive already downloaded.")
else:
print("started loading archive from url {}".format(url))
filename, headers = urlretrieve(url, dataset_file_path)
print("finished loading archive from url {}".format(url))
def extract_dataset(dataset_file_path, extraction_directory):
if (not os.path.exists(extraction_directory)):
os.makedirs(extraction_directory)
if (dataset_file_path.endswith("tar.gz") or dataset_file_path.endswith(".tgz")):
tar = tarfile.open(dataset_file_path, "r:gz")
tar.extractall(path=extraction_directory)
tar.close()
elif (dataset_file_path.endswith("tar")):
tar = tarfile.open(dataset_file_path, "r:")
tar.extractall(path=extraction_directory)
tar.close()
print("extraction of dataset from {} to {} done.".format(dataset_file_path,extraction_directory) )
# -
# # Laden der Daten
#
# Laden der Daten ausführen
#
download_dataset(urlDataSource,localDataArchive)
#
# Extrahieren der Daten
#
extract_dataset(localDataArchive,localExtractionFolder)
# # Organisation von Bilddaten auf dem Filesystem
#
# Eine gute Einführung in das Thema ist zu finden unter
#
# - [Brownlee](https://machinelearningmastery.com/how-to-load-large-datasets-from-directories-for-deep-learning-with-keras/)
# - [Sarkar](https://towardsdatascience.com/a-single-function-to-streamline-image-classification-with-keras-bd04f5cfe6df)
#
# <img src="info.png" align="left"/>
# # Erzeugen der Trainingsdaten
# +
#
# Hilfsfunktionen
#
def get_images(object_category, data_directory):
if (not os.path.exists(data_directory)):
print("data directory not found.")
return
obj_category_dir = os.path.join(os.path.join(data_directory,"101_ObjectCategories"),object_category)
images = [os.path.join(obj_category_dir,img) for img in os.listdir(obj_category_dir)]
return images
def return_images_per_category(data_directory):
folder = os.path.join(data_directory,"101_ObjectCategories")
#print(folder)
categories=[d for d in os.listdir(folder) if os.path.isdir(os.path.join(folder,d))]
#print(categories)
return categories
#
# Lesen der Bilddaten aus einer Datei. Anpassen der Größe auf 300x200 (Breite x Höhe) Pixel.
#
def read_image(image_path):
#img = cv2.imread(image_path, cv2.IMREAD_COLOR)
#img = cv2.resize(img, (300,200), interpolation=cv2.INTER_CUBIC)
im = Image.open(image_path).convert("RGB").resize((300,200))
np_img = np.array(im)
return np_img
#
# Sammelfunktion die alle Kategorien durchgeht und die Files sammelt
#
def create_training_data(data_directory,fraction):
i = 0
X = []
Y = []
print("started to read dataset from {}.".format(data_directory) )
for category in return_images_per_category(data_directory):
if category == 'BACKGROUND_Google':
continue
print(".",end='')
for image in get_images(category, data_directory):
if not image.endswith('.jpg'):
continue
if random.uniform(0, 1) > fraction:
continue
X.insert(i, read_image(image) )
Y.insert(i, category )
i += 1
print("finished reading dataset.")
X = np.array(X)
return X,Y
# +
#
# Erzeugen der Trainingsdaten. Der Faktor fraction bestimmt, wieviele Daten wirklich in den Speicher geladen werden.
# Achtung: diese Funktion kümmert sich nicht um die Gleichverteilung der Klassen.
#
X, Y = create_training_data(localExtractionFolder,fraction=0.4)
print('data X={}, y={}'.format(X.shape, len(Y)) )
# -
print(Y)
#
# Transformation der Labels in one-hot encoding
#
label_encoder = LabelEncoder()
Y_integer_encoded = label_encoder.fit_transform(Y)
Y_one_hot = to_categorical(Y_integer_encoded)
Y_one_hot.shape
# +
#
# Normalisieren der Bilddaten
#
X_normalized = ( X.astype(np.float64) / 255 ) + 0.001
#
# Löschen von X um Speicher gezielt freizumachen
#
del X
# +
#
# Split der Daten in Train und Test(validation) Datensätze
#
X_train, X_validation, Y_train, Y_validation = train_test_split(X_normalized, Y_one_hot, test_size=0.25, random_state=42)
del X_normalized
#
# gültige Werte in X_train, X_validation, Y_train, Y_validation, label_encoder, data_directory
#
# -
# # Prüfen der Daten
# +
#
# Form der Daten
#
print('train: X=%s, y=%s' % (X_train.shape, Y_train.shape))
print('test: X=%s, y=%s' % (X_validation.shape, Y_validation.shape))
#
# Plot von Bildern
#
for i in range(9):
plt.subplot(330 + 1 + i)
plt.imshow(X_train[i])
plt.show()
# -
# # Bauen eines Modelles
#
# Erzeugen eines einfache Modelles
#
def createModel():
model = Sequential()
model.add(Conv2D(16, (3,3), activation='relu', input_shape=(200,300,3)))
model.add(Conv2D(32, (3,3), activation='relu'))
model.add(MaxPooling2D(pool_size=2, strides=2))
model.add(Dropout(0.2))
model.add(Conv2D(64, (3,3), activation='relu'))
model.add(Conv2D(128, (3,3), activation='relu'))
model.add(MaxPooling2D(pool_size=2, strides=2))
model.add(Dropout(0.4))
model.add(Flatten())
model.add(Dense(101, activation='softmax'))
return model
#
# Compile und Training des Modelles
#
model_cnn = createModel()
model_cnn.compile(loss='categorical_crossentropy',optimizer='adam', metrics=['accuracy'])
#
# Callbacks steuern das Speichern von Checkpoints und eine Überwachung gegen Overfitting.
#
callbacks = [ModelCheckpoint('model_cnn_weights.h5', monitor='val_acc', save_best_only=True),
EarlyStopping(monitor='val_loss', patience=4, verbose=1, mode='auto')]
history = model_cnn.fit(X_train, Y_train, batch_size=16, epochs=6, verbose=1, validation_data=(X_validation,Y_validation), callbacks=callbacks)
#
# Evaluierung des Modelles
#
_, acc = model_cnn.evaluate(X_validation, Y_validation, verbose=0)
print('accuracy {:.3f} '.format(acc) )
#
# Ausgabe des Trainingsverlaufes
#
def summarize_diagnostics(history,modelname):
plt.subplot(211)
plt.title('Cross Entropy Loss')
plt.plot(history.history['loss'], color='blue', label='train')
plt.plot(history.history['val_loss'], color='lightblue', label='test')
plt.subplot(212)
plt.title('Classification Accuracy')
plt.plot(history.history['accuracy'], color='green', label='train')
plt.plot(history.history['val_accuracy'], color='lightgreen', label='test')
plt.subplots_adjust(hspace=0.5)
plt.savefig( 'results/' + modelname + '_plot.png')
plt.show()
plt.close()
summarize_diagnostics(history,'05_model_cnn')
# # Optimiertes Laden der Bilder
#
# Die bisherige Ladefunktion hat alle Bilder in den Speicher geladen. Das führt schnell dazu, dass der Hauptspeicher ausgeht. Daher benötigen wir eine Funktion, die Bilder der Reihe nach in den Speicher lädt und für das Training zur Verfügung stellt.
#
# Eine solche Funktion kann mit einem python **Generator** implementiert werden. Die Erklärung von Generatoren ist hier zu finden [2]. Das Tutorial zum Laden mit Generatoren ist hier [1] zu finden.
#
# Quellen:
# - [1] [https://towardsdatascience.com/a-single-function-to-streamline-image-classification-with-keras-bd04f5cfe6df](https://towardsdatascience.com/a-single-function-to-streamline-image-classification-with-keras-bd04f5cfe6df)
# - [2] [https://www.python-kurs.eu/generatoren.php](https://www.python-kurs.eu/generatoren.php)
#
#
# <img src="info.png" align="left"/>
#
#
# Anlegen eines Generators für Bilder
#
datagen = ImageDataGenerator()
it_train = datagen.flow(X_train, Y_train, batch_size=16)
#
# Training
#
model_cnn = createModel()
model_cnn.compile(loss='categorical_crossentropy',optimizer='adam', metrics=['accuracy'])
#
# Neue Funktion fit_generator
#
steps = int(X_train.shape[0] / 16)
history = model_cnn.fit_generator(it_train, steps_per_epoch=steps, epochs=6, validation_data=(X_validation,Y_validation), verbose=1, callbacks=callbacks)
#
# Evaluierung
#
_, acc = model_cnn.evaluate(X_validation, Y_validation, verbose=0)
print('accuracy {:.3f} '.format(acc) )
summarize_diagnostics(history,'model_cnn_gen')
# # Optimierung durch Augmentierung
#
# Augmentierung erweitert den Trainingsdatensatz um künstlich erzeugte Bilder. Damit wird erreicht, dass ein Modell robuster wird und sich nicht auf einzelne Pixel bezieht. Methoden der Augmentierung für Bilder sind:
#
# - Breite und Höhe des Bildinhaltes ändern (width_shift_range, height_shift_range)
# - Spiegelung (flip)
# - Rotation (rotation_range)
# - Zoomen (zoom_range)
# - Helligkeit (brightness_range)
# - Verzerrung (shear_range)
#
# Das Zufügen von Rauschen kann in Keras nicht direkt über den [ImageDataGenerator](https://keras.io/preprocessing/image/) eingestellt werden. Dies wird aber durch die Verwendung von Dropout annähernd simuliert.
#
# <img src="info.png" align="left"/>
#
#
# Anlegen eines Generators für Bilder
#
datagen = ImageDataGenerator(width_shift_range=0.1, height_shift_range=0.1, horizontal_flip=True, rotation_range=5, zoom_range=0.1)
# prepare iterator
it_train = datagen.flow(X_train, Y_train, batch_size=16)
#
# Training
#
steps = int(X_train.shape[0] / 16)
model_cnn = createModel()
model_cnn.compile(loss='categorical_crossentropy',optimizer='adam', metrics=['accuracy'])
history = model_cnn.fit_generator(it_train, steps_per_epoch=steps, epochs=24, validation_data=(X_validation,Y_validation), verbose=1, callbacks=callbacks)
#
# Evaluierung
#
_, acc = model_cnn.evaluate(X_validation, Y_validation, verbose=0)
print('accuracy {:.3f} '.format(acc) )
summarize_diagnostics(history,'05_model_cnn_aug')
| 05 Anwendungsbeispiel Importing of image data with augmentation and classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/objectc/CNN-with-TensorFlow2.0-and-Keras/blob/master/Resnet_residual_block.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="CnpkAcPxZGzD" colab_type="code" colab={}
#import needed classes
from tensorflow import keras
from tensorflow.keras.datasets import cifar10
from tensorflow.keras.layers import Dense,Conv2D,MaxPooling2D,Flatten,AveragePooling2D,Dropout,BatchNormalization,Activation, Input
from tensorflow.keras.models import Model
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.callbacks import LearningRateScheduler
from tensorflow.keras.callbacks import ModelCheckpoint
from math import ceil
import os
from tensorflow.keras.preprocessing.image import ImageDataGenerator
# + id="IwGns0UdZKhU" colab_type="code" colab={}
def Unit(x,filters,pool=False):
res = x
if pool:
x = MaxPooling2D(pool_size=(2, 2))(x)
res = Conv2D(filters=filters,kernel_size=[1,1],strides=(2,2),padding="same")(res)
out = BatchNormalization()(x)
out = Activation("relu")(out)
out = Conv2D(filters=filters, kernel_size=[3, 3], strides=[1, 1], padding="same")(out)
out = BatchNormalization()(out)
out = Activation("relu")(out)
out = Conv2D(filters=filters, kernel_size=[3, 3], strides=[1, 1], padding="same")(out)
out = keras.layers.add([res,out])
return out
# + id="jtG2k0coZN6W" colab_type="code" colab={}
#Define the model
def MiniModel(input_shape):
images = Input(input_shape)
net = Conv2D(filters=32, kernel_size=[3, 3], strides=[1, 1], padding="same")(images)
net = Unit(net,32)
net = Unit(net,32)
net = Unit(net,32)
net = Unit(net,64,pool=True)
net = Unit(net,64)
net = Unit(net,64)
net = Unit(net,128,pool=True)
net = Unit(net,128)
net = Unit(net,128)
net = Unit(net, 256,pool=True)
net = Unit(net, 256)
net = Unit(net, 256)
net = BatchNormalization()(net)
net = Activation("relu")(net)
net = Dropout(0.25)(net)
net = AveragePooling2D(pool_size=(4,4))(net)
net = Flatten()(net)
net = Dense(units=10,activation="softmax")(net)
model = Model(inputs=images,outputs=net)
return model
# + id="br7gwp0HZXH6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="b4452a12-7d8a-4c44-927a-0f33c2c4248f"
#load the cifar10 dataset
(train_x, train_y) , (test_x, test_y) = cifar10.load_data()
#normalize the data
train_x = train_x/255
test_x = test_x/255
#Subtract the mean image from both train and test set
# train_x = train_x - train_x.mean()
# test_x = test_x - test_x.mean()
#Divide by the standard deviation
# train_x = train_x / train_x.std(axis=0)
# test_x = test_x / test_x.std(axis=0)
datagen = ImageDataGenerator(rotation_range=10,
width_shift_range=5. / 32,
height_shift_range=5. / 32,
rescale=1./255,
horizontal_flip=True)
# Compute quantities required for featurewise normalization
# (std, mean, and principal components if ZCA whitening is applied).
datagen.fit(train_x)
#Encode the labels to vectors
train_y = keras.utils.to_categorical(train_y,10)
test_y = keras.utils.to_categorical(test_y,10)
#define a common unit
input_shape = (32,32,3)
model = MiniModel(input_shape)
#Print a Summary of the model
model.summary()
#Specify the training components
model.compile(optimizer=Adam(0.001),loss="categorical_crossentropy",metrics=["accuracy"])
# + id="ErjiTblBc7mz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="d431f211-4873-44da-b067-884f72901bd6"
epochs = 50
steps_per_epoch = ceil(50000/128)
# Fit the model on the batches generated by datagen.flow().
model.fit(datagen.flow(train_x, train_y, batch_size=128),
validation_data=(test_x,test_y),
epochs=epochs,
steps_per_epoch=steps_per_epoch,
verbose=1
)
#Evaluate the accuracy of the test dataset
accuracy = model.evaluate(x=test_x,y=test_y,batch_size=128)
model.save("cifar10model.h5")
| Resnet_residual_block.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: clouds113_kernel
# language: python
# name: clouds113_kernel
# ---
# ### Cumulative contributions to clc_32
#
# **For Figure 10**
# +
import os
import shap
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
from scipy.interpolate import make_interp_spline, BSpline
import importlib
importlib.reload(matplotlib)
importlib.reload(plt)
# -
# *Load SHAP files*
# +
r2b4_shap_files = []
for file in os.listdir('../shap_values'):
if file.startswith('r2b4_shap'):
r2b4_shap_files.append('./shap_values/'+file)
r2b4_file_count = len(r2b4_shap_files)
# +
r2b5_shap_files = []
for file in os.listdir('../shap_values'):
if file.startswith('r2b5_shap'):
r2b5_shap_files.append('./shap_values/'+file)
r2b5_file_count = len(r2b5_shap_files)
# +
# Append the averages of all shap value files we have for r2b4 and r2b5 each
r2b4_shap_values = [np.load(r2b4_shap_files[i]) for i in range(r2b4_file_count)]
r2b4_shap_means = [np.mean(r2b4_shap_values[i], axis=0) for i in range(r2b4_file_count)]
r2b5_shap_values = [np.load(r2b5_shap_files[i]) for i in range(r2b5_file_count)]
r2b5_shap_means = [np.mean(r2b5_shap_values[i], axis=0) for i in range(r2b5_file_count)]
# -
# *Feature names*
# +
# R2B4
r2b4_feature_names = []
feat_names = ['qv', 'qc', 'qi', 'temp', 'pres', 'rho', 'zg']
for s in feat_names:
for i in range(21, 48):
r2b4_feature_names.append('%s_%d'%(s, i))
r2b4_feature_names.append('fr_lake')
r2b4_feature_names = np.array(r2b4_feature_names)
remove_fields = [27, 162, 163, 164]
r2b4_feature_names = np.delete(r2b4_feature_names, remove_fields)
# +
# R2B5
r2b5_feature_names = []
feat_names = ['qv', 'qc', 'qi', 'temp', 'pres', 'zg']
for s in feat_names:
for i in range(21, 48):
r2b5_feature_names.append('%s_%d'%(s, i))
r2b5_feature_names.append('fr_land')
r2b5_feature_names = np.array(r2b5_feature_names)
remove_fields = [27, 28, 29, 30, 31, 32, 135, 136, 137]
r2b5_feature_names = np.delete(r2b5_feature_names, remove_fields)
# +
# Intersecting and unique features
features_intersect = np.intersect1d(r2b4_feature_names, r2b5_feature_names)
only_in_r2b5 = set(r2b5_feature_names).difference(set(features_intersect))
only_in_r2b4 = set(r2b4_feature_names).difference(set(features_intersect))
# +
# For every feature in features_intersect, we extract the means
r2b4_shap_means_intersect = []
r2b5_shap_means_intersect = []
for s in features_intersect:
feature_ind_r2b4 = np.where(r2b4_feature_names==s)[0][0]
feature_ind_r2b5 = np.where(r2b5_feature_names==s)[0][0]
r2b4_shap_means_intersect.append([r2b4_shap_means[i][feature_ind_r2b4] for i in range(r2b4_file_count)])
r2b5_shap_means_intersect.append([r2b5_shap_means[i][feature_ind_r2b5] for i in range(r2b5_file_count)])
# List with as many entries as there are intersecting features. Each entry has as many entries as there are shap value files.
assert len(r2b4_shap_means_intersect) == len(r2b5_shap_means_intersect) == len(features_intersect)
assert len(r2b4_shap_means_intersect[0]) == r2b4_file_count
assert len(r2b5_shap_means_intersect[0]) == r2b5_file_count
r2b4_shap_means_intersect = np.array(r2b4_shap_means_intersect)
r2b5_shap_means_intersect = np.array(r2b5_shap_means_intersect)
assert r2b4_shap_means_intersect.shape == (len(features_intersect), r2b4_file_count)
assert r2b5_shap_means_intersect.shape == (len(features_intersect), r2b5_file_count)
# -
np.sum(r2b5_shap_means_intersect[:2], axis=0)
# *Cumulative plot*
# +
# Sum over feature contributions from the entire column
r2b4_var_type = []
r2b5_var_type = []
for var in ['qv', 'qi', 'qc', 'zg', 'pres', 'temp']:
inds = [ind for ind in np.arange(len(features_intersect)) if features_intersect[ind].startswith(var)]
r2b4_var_type.append(np.sum(r2b4_shap_means_intersect[inds], axis=0))
r2b5_var_type.append(np.sum(r2b5_shap_means_intersect[inds], axis=0))
r2b4_var_type = np.array(r2b4_var_type)
r2b5_var_type = np.array(r2b5_var_type)
assert r2b4_var_type.shape == (6, r2b4_file_count)
assert r2b5_var_type.shape == (6, r2b5_file_count)
# +
# Errorbars: Maximum possible deviation
# R2B4
r2b4_err_lower = r2b4_var_type[:, 0] - np.min(r2b4_var_type, axis=1)
r2b4_err_upper = np.max(r2b4_var_type, axis=1) - r2b4_var_type[:, 0]
# R2B5
r2b5_err_lower = r2b5_var_type[:, 0] - np.min(r2b5_var_type, axis=1)
r2b5_err_upper = np.max(r2b5_var_type, axis=1) - r2b5_var_type[:, 0]
# -
# *SHAP vertical profiles for qi and qv*
# +
inds = [ind for ind in np.arange(len(features_intersect)) if features_intersect[ind].startswith('qv')]
i = 21
qv_r2b4 = []
qv_r2b5 = []
for ind in inds:
qv_r2b4.append(r2b4_shap_means_intersect[ind])
qv_r2b5.append(r2b5_shap_means_intersect[ind])
i += 1
# +
inds = [ind for ind in np.arange(len(features_intersect)) if features_intersect[ind].startswith('qi')]
i = 21
qi_r2b4 = []
qi_r2b5 = []
for ind in inds:
qi_r2b4.append(r2b4_shap_means_intersect[ind])
qi_r2b5.append(r2b5_shap_means_intersect[ind])
i += 1
# -
# *SHAP Dependence Plots*
#
# Conditional expectation line is computed over all seeds. <br>
# Point cloud is shown only for one seed.
# +
# To provide the Dependence Plot the corresponding 10000 NARVAL samples
r2b4_narval_r2b5_samples = np.load('./shap_values/r2b4_narval_r2b5_samples_layer_32_seed_100_train_samples_10000_narval_samples_10000.npy')
r2b5_narval_r2b5_samples = np.load('./shap_values/r2b5_narval_r2b5_samples_layer_32_seed_100_train_samples_7931_narval_samples_10000-constructed_base_value.npy')
# The 10000 NARVAL samples should be seed-independent
r2b4_narval_samples = ['./shap_values/'+file for file in os.listdir('./shap_values') if file.startswith('r2b4_narval_r2b5_samples')]
r2b5_narval_samples = ['./shap_values/'+file for file in os.listdir('./shap_values') if file.startswith('r2b5_narval_r2b5_samples')]
for i in range(len(r2b4_narval_samples)):
assert np.all(np.load(r2b4_narval_samples[0]) == np.load(r2b4_narval_samples[i]))
for i in range(len(r2b5_narval_samples)):
assert np.all(np.load(r2b5_narval_samples[0]) == np.load(r2b5_narval_samples[i]))
# +
# Needed to scale back to native units
# R2B4 Column-based model
r2b4_feature_means = np.array([2.62572183e-06,2.72625252e-06,2.74939600e-06,3.30840599e-06,6.62808605e-06,1.75788934e-05,4.56026919e-05,1.05190041e-04,2.05702805e-04,3.57870694e-04,5.71860616e-04,8.86342854e-04,1.40607454e-03,2.11394275e-03,2.96908898e-03,3.83956666e-03,4.85640761e-03,6.05059066e-03,7.37936039e-03,8.88779732e-03,1.05374548e-02,1.20163575e-02,1.32316365e-02,1.40249843e-02,1.44862015e-02,1.47169496e-02,1.49353026e-02,4.10339294e-14,1.09916165e-10,5.08967307e-11,9.79269311e-14,7.81782591e-13,1.59702138e-12,1.06302286e-08,1.03287141e-07,2.32342195e-07,4.52571159e-07,9.59800950e-07,2.75292262e-06,5.47922031e-06,6.96345062e-06,7.10544829e-06,8.49121303e-06,1.14876828e-05,1.62598283e-05,2.54900781e-05,3.60999973e-05,3.30096121e-05,1.50384025e-05,3.37482390e-06,9.94423396e-07,3.95924469e-07,2.27436437e-07,1.47661800e-14,4.78581565e-11,6.02759292e-09,7.85422277e-08,3.42838766e-07,1.03181587e-06,2.10645844e-06,2.66487045e-06,2.04870326e-06,1.01504965e-06,4.92335725e-07,2.89430485e-07,1.73665966e-07,6.58006285e-08,1.47246476e-08,2.46884148e-09,2.97776000e-10,2.23559883e-11,1.53999974e-12,9.41478240e-13,7.94546431e-13,5.45907918e-13,2.42190024e-13,1.03934147e-13,3.65539123e-14,1.55304439e-14,1.05358904e-14,2.08525301e+02,2.02078330e+02,1.96095922e+02,1.96231880e+02,2.02855933e+02,2.11649673e+02,2.21128411e+02,2.30533497e+02,2.39352824e+02,2.47584803e+02,2.54661191e+02,2.60677478e+02,2.65862196e+02,2.70101506e+02,2.74180293e+02,2.77942434e+02,2.81490486e+02,2.84592019e+02,2.87187378e+02,2.89183826e+02,2.90680284e+02,2.92060146e+02,2.93733091e+02,2.95405966e+02,2.96851675e+02,2.97902688e+02,2.98445713e+02,4.89214262e+03,6.41523961e+03,8.33266520e+03,1.08240088e+04,1.37001631e+04,1.70600136e+04,2.07632553e+04,2.48067011e+04,2.90956820e+04,3.37499929e+04,3.85120640e+04,4.34081851e+04,4.86049928e+04,5.36237056e+04,5.88910085e+04,6.39346849e+04,6.89441862e+04,7.37692055e+04,7.83390226e+04,8.26346116e+04,8.66219041e+04,9.01095619e+04,9.33174822e+04,9.59079770e+04,9.79586311e+04,9.94225005e+04,1.00212068e+05,8.18133756e-02,1.10740562e-01,1.48196795e-01,1.92120442e-01,2.35155442e-01,2.80640720e-01,3.26932112e-01,3.74686426e-01,4.23310942e-01,4.74706421e-01,5.26574353e-01,5.79757246e-01,6.36319453e-01,6.90718806e-01,7.46907098e-01,7.99472206e-01,8.50742657e-01,8.99725050e-01,9.46097691e-01,9.90226000e-01,1.03168333e+00,1.06722030e+00,1.09810057e+00,1.12163306e+00,1.13970514e+00,1.15248459e+00,1.15929123e+00,1.61339519e+04,1.47406270e+04,1.34213890e+04,1.21742147e+04,1.09970932e+04,9.88816591e+03,8.84571032e+03,7.86812123e+03,6.95389299e+03,6.10160326e+03,5.30990022e+03,4.57749545e+03,3.90316351e+03,3.28574924e+03,2.72418251e+03,2.21749985e+03,1.76487258e+03,1.36564177e+03,1.01936294e+03,7.25868116e+02,4.85363829e+02,2.98613214e+02,1.67518124e+02,9.62005988e+01,2.51278402e-03])
r2b4_feature_stds = np.array([1.47224807e-07,2.51491754e-07,2.82187441e-07,5.93724945e-07,1.51594298e-06,6.30296895e-06,2.22274936e-05,6.22211930e-05,1.43226310e-04,2.86338200e-04,4.85240662e-04,7.58182385e-04,1.13414912e-03,1.54548518e-03,1.89771471e-03,2.12619203e-03,2.33918584e-03,2.55623874e-03,2.79223044e-03,2.96787488e-03,2.98239542e-03,2.88134285e-03,2.89833854e-03,3.00368460e-03,3.09352534e-03,3.13736360e-03,3.16548101e-03,9.42811508e-12,3.44228546e-09,1.59084922e-09,5.93648639e-12,3.42805209e-11,8.49979059e-11,1.42841729e-07,8.71018616e-07,1.61722240e-06,2.73813894e-06,4.98860589e-06,1.09156013e-05,1.92056078e-05,2.58635856e-05,2.67312436e-05,2.88542767e-05,3.27361856e-05,3.80907041e-05,4.95573286e-05,6.64222088e-05,6.35284725e-05,4.02976359e-05,1.60662423e-05,8.31400034e-06,4.98668318e-06,4.18798497e-06,4.72870218e-12,9.65155904e-09,1.53906014e-07,1.30802262e-06,3.95656270e-06,7.01268576e-06,9.21713269e-06,9.56682609e-06,7.50242259e-06,4.54526978e-06,2.74400532e-06,1.68254503e-06,9.94899145e-07,4.90686500e-07,1.74423527e-07,6.26854136e-08,1.35744721e-08,2.72159858e-09,2.39259137e-10,6.37191578e-11,5.72720348e-11,4.81701846e-11,2.14967902e-11,1.01020924e-11,4.65900766e-12,3.62080895e-12,3.08853822e-12,2.14124189e+00,2.04636219e+00,2.28842093e+00,2.18477413e+00,1.60619664e+00,1.23447911e+00,1.36168420e+00,1.59257820e+00,1.77424088e+00,1.85501643e+00,1.85519231e+00,1.74569793e+00,1.50736838e+00,1.37280694e+00,1.30119982e+00,1.30107249e+00,1.42479076e+00,1.62621509e+00,1.88375876e+00,2.13026250e+00,2.32127917e+00,2.48017927e+00,2.58637194e+00,2.59273129e+00,2.57883413e+00,2.58848551e+00,2.63916745e+00,5.25025104e+01,6.13379273e+01,6.86279405e+01,8.39005677e+01,1.08798431e+02,1.33773907e+02,1.50246438e+02,1.57891625e+02,1.59368430e+02,1.59393957e+02,1.63436704e+02,1.78941090e+02,2.18262201e+02,2.74998258e+02,3.62779581e+02,4.62465472e+02,5.85167856e+02,7.29868327e+02,8.80813572e+02,1.05716175e+03,1.23096824e+03,1.39147497e+03,1.57507360e+03,1.69639911e+03,1.79506877e+03,1.86809404e+03,1.89542576e+03,9.42444728e-04,1.10053780e-03,1.74702325e-03,2.61821961e-03,2.72889747e-03,2.27853670e-03,1.76545251e-03,1.59704425e-03,1.94238096e-03,2.47019694e-03,3.01007451e-03,3.43331975e-03,3.85832590e-03,4.47209741e-03,5.18871128e-03,5.98561751e-03,7.30225606e-03,9.21774294e-03,1.16181468e-02,1.47943226e-02,1.80870903e-02,2.07868612e-02,2.28726988e-02,2.36245337e-02,2.39326331e-02,2.39750201e-02,2.37309135e-02,4.37962105e-01,1.18129232e+00,1.95876778e+00,3.13352980e+00,4.84460444e+00,7.25070491e+00,1.05220551e+01,1.48284694e+01,2.03242942e+01,2.71311569e+01,3.53196384e+01,4.48911206e+01,5.57614000e+01,6.77483165e+01,8.05664007e+01,9.38318753e+01,1.07080624e+02,1.19799754e+02,1.31470485e+02,1.41617295e+02,1.49856634e+02,1.55939200e+02,1.60153490e+02,1.61897063e+02,1.10524044e-02])
# R2B5 Column-based (fold 2) model
r2b5_feature_means = np.array([2.57681365e-06,2.60161901e-06,2.86229890e-06,3.49524686e-06,6.32444387e-06,1.62852938e-05,4.26197236e-05,1.00492283e-04,2.10850387e-04,3.96992495e-04,6.62768743e-04,1.00639902e-03,1.42273038e-03,1.89269379e-03,2.42406883e-03,2.97704256e-03,3.52303812e-03,4.15430913e-03,4.89285256e-03,5.71192194e-03,6.58451740e-03,7.47955824e-03,8.42949837e-03,9.18162558e-03,9.58900058e-03,9.80246788e-03,9.98071441e-03,2.57897497e-16,1.24502901e-08,5.43912468e-07,1.97554777e-06,2.10205332e-06,3.45718981e-06,4.17987790e-06,4.89876027e-06,6.03250921e-06,6.71487544e-06,7.71281746e-06,9.96528417e-06,1.40351017e-05,1.87534642e-05,2.15523809e-05,1.77725032e-05,1.10700238e-05,6.98113679e-06,5.98240074e-06,8.03857856e-06,1.55278994e-05,1.98903187e-13,1.45240003e-10,2.39426913e-08,5.63226688e-07,3.10209365e-06,6.64324795e-06,8.83422658e-06,9.89681102e-06,9.97096463e-06,7.74324652e-06,4.95774608e-06,2.61087000e-06,1.29680563e-06,7.46596833e-07,4.94444102e-07,3.51674311e-07,2.61199355e-07,2.03219747e-07,1.66907845e-07,1.42871199e-07,1.25114261e-07,1.11956533e-07,1.02782118e-07,9.86031894e-08,9.95790399e-08,1.06733810e-07,1.26921172e-07,2.10924633e+02,2.07944695e+02,2.05115507e+02,2.03204784e+02,2.06103772e+02,2.12329817e+02,2.19299382e+02,2.26348890e+02,2.33352039e+02,2.40105681e+02,2.46401637e+02,2.52153555e+02,2.57207037e+02,2.61575645e+02,2.65446543e+02,2.68951996e+02,2.72093136e+02,2.74765728e+02,2.76963041e+02,2.78775116e+02,2.80398659e+02,2.81959850e+02,2.83501227e+02,2.84935364e+02,2.86119192e+02,2.86867707e+02,2.87046277e+02,4.78805278e+03,6.25615004e+03,8.06726288e+03,1.03500805e+04,1.30603494e+04,1.61944127e+04,1.97232230e+04,2.36181577e+04,2.78401230e+04,3.23377105e+04,3.70511232e+04,4.19785078e+04,4.70365400e+04,5.21124420e+04,5.72512536e+04,6.23517142e+04,6.72989145e+04,7.20972394e+04,7.66740332e+04,8.09510300e+04,8.49437983e+04,8.85136468e+04,9.16490946e+04,9.42529147e+04,9.63348759e+04,9.77633315e+04,9.86144363e+04,1.61343240e+04,1.47416307e+04,1.34230525e+04,1.21768751e+04,1.10012039e+04,9.89431495e+03,8.85470770e+03,7.88104473e+03,6.97198713e+03,6.12617252e+03,5.34218664e+03,4.61854836e+03,3.95376191e+03,3.34629894e+03,2.79465640e+03,2.29750295e+03,1.85381761e+03,1.46282067e+03,1.12390793e+03,8.36771545e+02,6.01482480e+02,4.18667943e+02,2.90324051e+02,2.20122534e+02,2.57179068e-01])
r2b5_feature_stds = np.array([1.66577356e-07,2.69438906e-07,6.32166532e-07,1.46870734e-06,2.84939866e-06,8.72797379e-06,2.96195352e-05,8.32385500e-05,1.93655438e-04,3.82345501e-04,6.27888913e-04,9.30858552e-04,1.27418047e-03,1.61904466e-03,1.95753088e-03,2.23604988e-03,2.49372225e-03,2.83062031e-03,3.22013981e-03,3.62381600e-03,4.05060687e-03,4.53912094e-03,5.14120557e-03,5.61150119e-03,5.82135854e-03,5.92232391e-03,6.02114792e-03,1.93770206e-12,1.94386132e-07,2.96883744e-06,8.75974976e-06,1.02724976e-05,1.44929996e-05,1.65663508e-05,1.81326398e-05,2.09805520e-05,2.41254125e-05,2.82129201e-05,3.57128254e-05,4.76374494e-05,5.95853155e-05,6.60615445e-05,5.80449728e-05,4.41472861e-05,3.63224833e-05,3.84500230e-05,5.48299167e-05,1.01230094e-04,2.84426774e-10,8.67755936e-08,2.57513880e-06,8.53816046e-06,1.97356234e-05,2.80242488e-05,3.05461589e-05,3.17141059e-05,3.22684724e-05,2.65101493e-05,1.83177779e-05,1.05168506e-05,6.02113023e-06,4.48008643e-06,3.22093921e-06,2.22409748e-06,1.62946826e-06,1.31793070e-06,1.14119306e-06,1.02620335e-06,9.44881472e-07,8.91631794e-07,8.56775098e-07,8.35271824e-07,8.19217124e-07,8.09879379e-07,8.37114763e-07,4.61938080e+00,5.32560366e+00,6.59828260e+00,8.11723979e+00,6.41348334e+00,3.53782199e+00,3.22568870e+00,5.51353694e+00,7.69704358e+00,9.19195458e+00,1.00629480e+01,1.04422426e+01,1.05152774e+01,1.05114012e+01,1.05471048e+01,1.07545816e+01,1.10865116e+01,1.13965606e+01,1.17078707e+01,1.20696061e+01,1.24923716e+01,1.28886157e+01,1.32697644e+01,1.37077879e+01,1.42377515e+01,1.47618886e+01,1.53836576e+01,1.65668010e+02,2.26139751e+02,3.23181791e+02,4.81891512e+02,6.96129876e+02,9.23059045e+02,1.13288883e+03,1.30878137e+03,1.44327690e+03,1.53977206e+03,1.61213693e+03,1.68432353e+03,1.78325130e+03,1.93151619e+03,2.15397041e+03,2.45262514e+03,2.80472214e+03,3.20634251e+03,3.63386944e+03,4.07078962e+03,4.52036282e+03,4.94232873e+03,5.33456701e+03,5.66290433e+03,5.92823270e+03,6.11058960e+03,6.23266007e+03,1.66988637e+00,4.50375687e+00,7.46700178e+00,1.19425370e+01,1.84570049e+01,2.76286540e+01,4.08392722e+01,6.01544270e+01,8.66284809e+01,1.20094159e+02,1.59157271e+02,2.01903044e+02,2.46894999e+02,2.92578673e+02,3.37234060e+02,3.79561515e+02,4.19219129e+02,4.56022726e+02,4.89322766e+02,5.18456198e+02,5.42710799e+02,5.61319067e+02,5.74407679e+02,5.79710837e+02,4.23033734e-01])
# -
def conditional_line(shap_values, narval_samples, feature_names, feature, eps = 1e-4):
feature_ind = np.where(feature_names=='%s_32'%feature)[0][0]
xvals = narval_samples[:, feature_ind]
yvals = np.mean(np.array(shap_values), axis=0)[:, feature_ind] # Average of shap values over all seeds
k_max = int(np.floor(max(xvals)/eps))
b = []
for k in range(k_max): # Stop after we reached the maximum value for x
b.append(np.mean([yvals[i] for i in range(len(yvals)) if k*eps <= xvals[i] < (k+1)*eps])) # Basically using bins here
# Corresponding x-values
# a = eps*np.arange(k_max) + eps/2
a = eps*np.arange(k_max)
# We have nans if there are no points in k*eps <= xvals[i] < (k+1)*eps. We simply remove these
a_new = [a[i] for i in range(len(b)) if ~np.isnan(b[i])]
b_new = [b[i] for i in range(len(b)) if ~np.isnan(b[i])]
# We use a spline of degree 3 to draw a smooth line
xnew = np.linspace(min(a_new), max(a_new), 200)
spl = make_interp_spline(a_new, b_new, k=3)
y_smooth = spl(xnew)
return xnew, y_smooth
# +
fig = plt.figure(figsize=(18,5))
# plt.subplots_adjust(bottom=0.1)
label_size=20
## First plot
ax = fig.add_subplot(131)
# Scale back to native units
feature_ind_r2b4 = np.where(r2b4_feature_names=='qi_32')[0][0]
r2b4_narval_r2b5_samples_qi = r2b4_narval_r2b5_samples*r2b4_feature_stds[feature_ind_r2b4] + r2b4_feature_means[feature_ind_r2b4]
feature_ind_r2b5 = np.where(r2b5_feature_names=='qi_32')[0][0]
r2b5_narval_r2b5_samples_qi = r2b5_narval_r2b5_samples*r2b5_feature_stds[feature_ind_r2b5] + r2b5_feature_means[feature_ind_r2b5]
# The narval samples should be the same in their original unnormalized space
assert np.all(np.abs(r2b5_narval_r2b5_samples_qi[:, feature_ind_r2b5] - r2b4_narval_r2b5_samples_qi[:, feature_ind_r2b4]) < 1e-10)
# Average SHAP values
r2b4_mean = np.mean(np.array(r2b4_shap_values)[:, :, feature_ind_r2b4], dtype=np.float64)
r2b5_mean = np.mean(np.array(r2b5_shap_values)[:, :, feature_ind_r2b5], dtype=np.float64)
# Put the one with the larger range second
sdp = shap.dependence_plot(feature_ind_r2b4, r2b4_shap_values[0], features=r2b4_narval_r2b5_samples_qi, ax=ax,
feature_names=r2b4_feature_names, interaction_index=None, show=False, color='blue', dot_size=5, alpha=0.6)
sdp = shap.dependence_plot(feature_ind_r2b5, r2b5_shap_values[0], features=r2b5_narval_r2b5_samples_qi, ax=ax,
feature_names=r2b5_feature_names, interaction_index=None, show=False, color='orange', dot_size=5, alpha=0.6)
# It's the same as:
# sdp = shap.dependence_plot(0, r2b5_shap_values[0][:, feature_ind_r2b5:(feature_ind_r2b5+1)], features=r2b5_narval_r2b5_samples_qi[:, feature_ind_r2b5:(feature_ind_r2b5+1)],
# ax=ax, feature_names=r2b5_feature_names, interaction_index=None, show=False, color='orange', dot_size=5)
# Plot showing averages
qi_min = np.min(r2b4_narval_r2b5_samples_qi[:, feature_ind_r2b4])
qi_max = np.max(r2b4_narval_r2b5_samples_qi[:, feature_ind_r2b4])
plt.plot([qi_min, qi_max], [r2b4_mean, r2b4_mean], 'b--', linewidth=1.5)
plt.plot([qi_min, qi_max], [r2b5_mean, r2b5_mean], color='orange', linestyle='--', linewidth=1.5)
# Legend
ax.annotate('NARVAL R2B4 model', xy=(0.5,0.84),xycoords='axes fraction', color='blue', fontsize=14)
ax.annotate('QUBICC R2B5 model', xy=(0.5,0.9),xycoords='axes fraction', color='orange', fontsize=14)
plt.xlabel('$q_i$_32 [kg/kg]', fontsize=label_size)
plt.ylabel('SHAP values for clc_32', fontsize=label_size)
# Conditional averages. The choice of eps has a large influence on the plot
m = 50
xnew, y_smooth = conditional_line(r2b4_shap_values, r2b4_narval_r2b5_samples_qi, r2b4_feature_names, 'qi', eps = 5*1e-6) # eps = 2*1e-5
ax.plot(xnew[:m], y_smooth[:m], linewidth=4)
xnew, y_smooth = conditional_line(r2b5_shap_values, r2b5_narval_r2b5_samples_qi, r2b5_feature_names, 'qi', eps = 5*1e-6)
ax.plot(xnew[:m], y_smooth[:m], linewidth=4)
plt.ylim((-6.279482202575369, 87.43461904261041)) # Taken from the qv plot
## Second plot
ax_2 = fig.add_subplot(132)
# Scale back to native units
feature_ind_r2b4 = np.where(r2b4_feature_names=='qv_32')[0][0]
r2b4_narval_r2b5_samples_qv = r2b4_narval_r2b5_samples*r2b4_feature_stds[feature_ind_r2b4] + r2b4_feature_means[feature_ind_r2b4]
feature_ind_r2b5 = np.where(r2b5_feature_names=='qv_32')[0][0]
r2b5_narval_r2b5_samples_qv = r2b5_narval_r2b5_samples*r2b5_feature_stds[feature_ind_r2b5] + r2b5_feature_means[feature_ind_r2b5]
# The narval samples should be the same in their original unnormalized space
assert np.all(np.abs(r2b5_narval_r2b5_samples_qv[:, feature_ind_r2b5] - r2b4_narval_r2b5_samples_qv[:, feature_ind_r2b4]) < 1e-10)
# Average SHAP values
r2b4_mean = np.mean(np.array(r2b4_shap_values)[:, :, feature_ind_r2b4], dtype=np.float64)
r2b5_mean = np.mean(np.array(r2b5_shap_values)[:, :, feature_ind_r2b5], dtype=np.float64)
# Put the one with the larger range second
sdp_2 = shap.dependence_plot(feature_ind_r2b4, r2b4_shap_values[0], features=r2b4_narval_r2b5_samples_qv, ax=ax_2,
feature_names=r2b4_feature_names, interaction_index=None, show=False, color='blue', dot_size=5, alpha=0.6)
sdp_2 = shap.dependence_plot(feature_ind_r2b5, r2b5_shap_values[0], features=r2b5_narval_r2b5_samples_qv, ax=ax_2,
feature_names=r2b5_feature_names, interaction_index=None, show=False, color='orange', dot_size=5, alpha=0.6)
# Plot showing averages
qv_min = np.min(r2b4_narval_r2b5_samples_qv[:, feature_ind_r2b4])
qv_max = np.max(r2b4_narval_r2b5_samples_qv[:, feature_ind_r2b4])
plt.plot([qv_min, qv_max], [r2b4_mean, r2b4_mean], 'b--', linewidth=1.5)
plt.plot([qv_min, qv_max], [r2b5_mean, r2b5_mean], color='orange', linestyle='--', linewidth=1.5)
# Conditional averages. The choice of eps has a large influence on the plot
xnew, y_smooth = conditional_line(r2b4_shap_values, r2b4_narval_r2b5_samples_qv, r2b4_feature_names, 'qv', eps = 4*1e-4)
ax_2.plot(xnew, y_smooth, linewidth=4)
xnew, y_smooth = conditional_line(r2b5_shap_values, r2b5_narval_r2b5_samples_qv, r2b5_feature_names, 'qv', eps = 4*1e-4)
ax_2.plot(xnew, y_smooth, linewidth=4)
plt.gca().ticklabel_format(axis='x', style='sci', scilimits=(-2,2))
# ax_2.xaxis.set_major_formatter(FormatStrFormatter('%E'))
plt.xlabel('$q_v$_32 [kg/kg]', fontsize=label_size)
plt.ylabel(' ')
# plt.savefig('figures/shap_dependence_plots.pdf')
# -
# plt.plot(qv_diffs, np.arange(27, 27+len(qi_diffs)), 'bo')
line_r2b5 = plt.plot(qv_r2b5, np.arange(21, 21+len(qv_r2b5)), '.', color='orange')
line_r2b4 = plt.plot(qv_r2b4, np.arange(21, 21+len(qv_r2b5)), '.', color='blue')
plt.ylabel('Vertical layer')
plt.xlabel('SHAP value for qv')
plt.title('qv')
# plt.xlabel('SHAP value difference')
plt.legend([line_r2b5[0], line_r2b4[0]], ['R2B5 QUBICC model', 'R2B4 NARVAL model'])
plt.grid(b=True)
# plt.legend(['NARVAL - QUBICC qi'])
plt.gca().invert_yaxis()
# plt.plot(qi_diffs, np.arange(27, 27+len(qi_diffs)), 'bo')
line_r2b5 = plt.plot(qi_r2b5, np.arange(21, 21+len(qi_r2b5)), '.', color='orange')
line_r2b4 = plt.plot(qi_r2b4, np.arange(21, 21+len(qi_r2b5)), '.', color='blue')
plt.ylabel('Vertical layer')
plt.xlabel('SHAP value for qi')
plt.title('qi')
# plt.xlabel('SHAP value difference')
plt.legend([line_r2b5[0], line_r2b4[0]], ['R2B5 QUBICC model', 'R2B4 NARVAL model'])
plt.grid(b=True)
# plt.legend(['NARVAL - QUBICC qi'])
plt.gca().invert_yaxis()
# +
# Bars show minimum and maximum value
x_labels = ['qv','qi', 'qc', 'zg', 'pres','temp']
x = np.arange(len(x_labels)) # Label locations!
width = 0.4
fig = plt.figure()
ax = fig.add_subplot(111, ylabel='Sum of SHAP values', title='Contributions to clc_32 from the entire column')
ax.axhline(np.sum(r2b5_var_type[:, 0]), xmin=0, xmax=1, color='orange', linewidth=1)
ax.bar(np.arange(len(x_labels))-width/2, r2b5_var_type[:, 0], width=width, color='orange', yerr=np.array([r2b5_err_lower, r2b5_err_upper]),\
align='center', alpha=0.5, ecolor='black', capsize=5)
ax.bar(np.arange(len(x_labels))+width/2, r2b4_var_type[:, 0], width=width, color='blue', yerr=np.array([r2b4_err_lower, r2b4_err_upper]),\
align='center', alpha=0.5, ecolor='black', capsize=5)
ax.set_xticks(x)
ax.set_xticklabels(x_labels)
ax.legend(['R2B5 QUBICC model bias', 'R2B5 QUBICC model', 'R2B4 NARVAL model'])
ax.axhline(0, xmin=0, xmax=1, color='gray', linewidth=.5, ls='--')
# plt.savefig('figures/shap_clc_32_cumulative.pdf', bbox_inches='tight')
# -
# **All in one plot**
# +
# matplotlib.rcParams # To see all parameters of matplotlib
# +
import matplotlib
# Increase the general font size in plots
size_plots_label = 22
matplotlib.rcParams['legend.fontsize'] = size_plots_label
matplotlib.rcParams['axes.labelsize'] = size_plots_label # For an axes xlabel and ylabel
matplotlib.rcParams['axes.titlesize'] = size_plots_label+2 # For an axes xlabel and ylabel
matplotlib.rcParams['xtick.labelsize'] = size_plots_label
matplotlib.rcParams['ytick.labelsize'] = size_plots_label
# Averaged over the NARVAL region
zg_mean_narval = [20785,19153,17604,16134,14741,13422,12175,10998,9890,8848,
7871,6958,6107,5317,4587,3915,3300,2741,2237,1787,1390,1046,
754,515,329,199,128] # in meters
zg_mean_narval = np.round(np.array(zg_mean_narval)/1000, decimals=1) # in kilometers
# Averaged globally
zg_mean_qubicc = [20785,19153,17604,16134,14742,13424,12178,11002,9896,8857,
7885,6977,6133,5351,4630,3968,3363,2814,2320,1878,1490,1153,
867,634,452,324,254] # in meters
zg_mean_qubicc = np.round(np.array(zg_mean_qubicc)/1000, decimals=1) # in kilometers
green='#004D40'
red='#D81B60'
blue='#1E88E5'
# +
fig = plt.figure(figsize=(30,11))
# plt.subplots_adjust(bottom=0.1)
# # Increase the general font size
# matplotlib.rcParams['legend.fontsize'] = 'x-large'
# matplotlib.rcParams['axes.labelsize'] = 'xx-large' # For an axes xlabel and ylabel
# matplotlib.rcParams['xtick.labelsize'] = 'xx-large'
# matplotlib.rcParams['ytick.labelsize'] = 'xx-large'
# label_size=20 # For the dependence plots
## First plot
ax1 = fig.add_subplot(121, ylabel='$\Sigma$(SHAP values) / |Samples|')
# Bars show minimum and maximum value
x_labels = ['$q_v$','$q_i$', '$q_c$', '$z_g$', '$p$','$T$']
x = np.arange(len(x_labels)) # Label locations!
width = 0.4
ax1.axhline(np.sum(r2b5_var_type[:, 0]), xmin=0, xmax=1, color='orange', linewidth=1)
ax1.bar(np.arange(len(x_labels))-width/2, r2b5_var_type[:, 0], width=width, color='orange', yerr=np.array([r2b5_err_lower, r2b5_err_upper]),\
align='center', alpha=0.5, ecolor='black', capsize=5)
ax1.bar(np.arange(len(x_labels))+width/2, r2b4_var_type[:, 0], width=width, color='blue', yerr=np.array([r2b4_err_lower, r2b4_err_upper]),\
align='center', alpha=0.5, ecolor='black', capsize=5)
ax1.set_xticks(x)
ax1.set_xticklabels(x_labels)
ax1.set_title(r'$\bf{(a)}$ Summed SHAP values from the entire grid column ', fontsize=size_plots_label, pad=12)
ax1.legend(['R2B5 QUBICC model bias', 'R2B5 QUBICC model', 'R2B4 NARVAL model'])
ax1.axhline(0, xmin=0, xmax=1, color='gray', linewidth=.5, ls='--')
## Second plot
ax2 = fig.add_subplot(243, ylabel='Vertical layer')
# plt.plot(qv_diffs, np.arange(27, 27+len(qi_diffs)), 'bo')
line_r2b5 = ax2.plot(qv_r2b5, np.arange(21, 21+len(qv_r2b5)), '.', color='orange')
line_r2b4 = ax2.plot(qv_r2b4, np.arange(21, 21+len(qv_r2b5)), '.', color='blue')
# plt.xlabel('SHAP value difference')
ax2.legend([line_r2b5[0], line_r2b4[0]], ['R2B5 QUBICC model', 'R2B4 NARVAL model'], markerscale=3) # markerscale makes dots larger/readable in the legend!
ax2.grid(b=True)
ax2.set_title(r'$\bf{(b)}$ Mean SHAP values of $q_v$ per layer', fontsize=size_plots_label, pad=12)
# plt.legend(['NARVAL - QUBICC qi'])
plt.gca().invert_yaxis()
## Third plot
ax3 = fig.add_subplot(244)
# plt.plot(qi_diffs, np.arange(27, 27+len(qi_diffs)), 'bo')
line_r2b5 = ax3.plot(qi_r2b5, np.arange(21, 21+len(qi_r2b5)), '.', color='orange')
line_r2b4 = ax3.plot(qi_r2b4, np.arange(21, 21+len(qi_r2b5)), '.', color='blue')
# plt.xlabel('SHAP value difference')
# ax3.legend([line_r2b5[0], line_r2b4[0]], ['R2B5 QUBICC model', 'R2B4 NARVAL model'])
ax3.grid(b=True)
ax3.set_title(r'$\bf{(c)}$ Mean SHAP values of $q_i$ per layer', fontsize=size_plots_label, pad=12)
# plt.legend(['NARVAL - QUBICC qi'])
plt.gca().invert_yaxis()
## Forth plot
ax4 = fig.add_subplot(247)
# Scale back to native units
feature_ind_r2b4 = np.where(r2b4_feature_names=='qv_32')[0][0]
r2b4_narval_r2b5_samples_qv = r2b4_narval_r2b5_samples*r2b4_feature_stds[feature_ind_r2b4] + r2b4_feature_means[feature_ind_r2b4]
feature_ind_r2b5 = np.where(r2b5_feature_names=='qv_32')[0][0]
r2b5_narval_r2b5_samples_qv = r2b5_narval_r2b5_samples*r2b5_feature_stds[feature_ind_r2b5] + r2b5_feature_means[feature_ind_r2b5]
# Average SHAP values
r2b4_mean = np.mean(np.array(r2b4_shap_values)[:, :, feature_ind_r2b4], dtype=np.float64)
r2b5_mean = np.mean(np.array(r2b5_shap_values)[:, :, feature_ind_r2b5], dtype=np.float64)
# Put the one with the larger range second
sdp_2 = shap.dependence_plot(feature_ind_r2b4, r2b4_shap_values[0], features=r2b4_narval_r2b5_samples_qv, ax=ax4,
feature_names=r2b4_feature_names, interaction_index=None, show=False, color='blue', dot_size=5, alpha=0.7)
sdp_2 = shap.dependence_plot(feature_ind_r2b5, r2b5_shap_values[0], features=r2b5_narval_r2b5_samples_qv, ax=ax4,
feature_names=r2b5_feature_names, interaction_index=None, show=False, color='orange', dot_size=5, alpha=0.7)
# Plot showing averages
qv_min = np.min(r2b4_narval_r2b5_samples_qv[:, feature_ind_r2b4])
qv_max = np.max(r2b4_narval_r2b5_samples_qv[:, feature_ind_r2b4])
plt.plot([qv_min, qv_max], [r2b4_mean, r2b4_mean], 'b--', linewidth=1.5)
plt.plot([qv_min, qv_max], [r2b5_mean, r2b5_mean], color='orange', linestyle='--', linewidth=1.5)
# Conditional averages. The choice of eps has a large influence on the plot
xnew, y_smooth = conditional_line(r2b4_shap_values, r2b4_narval_r2b5_samples_qv, r2b4_feature_names, 'qv', eps = 4*1e-4)
ax4.plot(xnew, y_smooth, linewidth=4)
xnew, y_smooth = conditional_line(r2b5_shap_values, r2b5_narval_r2b5_samples_qv, r2b5_feature_names, 'qv', eps = 4*1e-4)
ax4.plot(xnew, y_smooth, linewidth=4)
plt.gca().ticklabel_format(axis='x', style='sci', scilimits=(-2,2))
# ax_2.xaxis.set_major_formatter(FormatStrFormatter('%E'))
# Legend
ax4.annotate(r'$\bf{(d)}$', xy=(0.1,0.84),xycoords='axes fraction', fontsize=size_plots_label)
# ax4.annotate('NARVAL R2B4 model', xy=(0.1,0.84),xycoords='axes fraction', color='blue', fontsize=14)
# ax4.annotate('QUBICC R2B5 model', xy=(0.1,0.9),xycoords='axes fraction', color='orange', fontsize=14)
ax4.set_xlabel('$q_v$_32 [kg/kg]', fontsize=size_plots_label)
ax4.set_ylabel('SHAP value', fontsize=size_plots_label)
ax4.tick_params(labelsize=size_plots_label)
qv_ylim = plt.ylim()
## Fifth plot
ax5 = fig.add_subplot(248)
# Scale back to native units
feature_ind_r2b4 = np.where(r2b4_feature_names=='qi_32')[0][0]
r2b4_narval_r2b5_samples_qi = r2b4_narval_r2b5_samples*r2b4_feature_stds[feature_ind_r2b4] + r2b4_feature_means[feature_ind_r2b4]
feature_ind_r2b5 = np.where(r2b5_feature_names=='qi_32')[0][0]
r2b5_narval_r2b5_samples_qi = r2b5_narval_r2b5_samples*r2b5_feature_stds[feature_ind_r2b5] + r2b5_feature_means[feature_ind_r2b5]
# Average SHAP values
r2b4_mean = np.mean(np.array(r2b4_shap_values)[:, :, feature_ind_r2b4], dtype=np.float64)
r2b5_mean = np.mean(np.array(r2b5_shap_values)[:, :, feature_ind_r2b5], dtype=np.float64)
# Put the one with the larger range second
sdp = shap.dependence_plot(feature_ind_r2b4, r2b4_shap_values[0], features=r2b4_narval_r2b5_samples_qi, ax=ax5,
feature_names=r2b4_feature_names, interaction_index=None, show=False, color='blue', dot_size=5, alpha=0.7, xmax="percentile(99.85)") # Better to cut off at a high percentile
sdp = shap.dependence_plot(feature_ind_r2b5, r2b5_shap_values[0], features=r2b5_narval_r2b5_samples_qi, ax=ax5,
feature_names=r2b5_feature_names, interaction_index=None, show=False, color='orange', dot_size=5, alpha=0.7, xmax="percentile(99.85)")
# Plot showing averages
qi_min = np.min(r2b4_narval_r2b5_samples_qi[:, feature_ind_r2b4])
qi_max = np.max(r2b4_narval_r2b5_samples_qi[:, feature_ind_r2b4])
ax5.plot([qi_min, qi_max], [r2b4_mean, r2b4_mean], 'b--', linewidth=1.5)
ax5.plot([qi_min, qi_max], [r2b5_mean, r2b5_mean], color='orange', linestyle='--', linewidth=1.5)
# Conditional averages. The choice of eps has a large influence on the plot
m = 50
xnew, y_smooth = conditional_line(r2b4_shap_values, r2b4_narval_r2b5_samples_qi, r2b4_feature_names, 'qi', eps = 8*1e-6) # eps = 2*1e-5
ax5.plot(xnew[:m], y_smooth[:m], linewidth=4)
xnew, y_smooth = conditional_line(r2b5_shap_values, r2b5_narval_r2b5_samples_qi, r2b5_feature_names, 'qi', eps = 8*1e-6)
ax5.plot(xnew[:m], y_smooth[:m], linewidth=4)
# Legend
ax5.annotate(r'$\bf{(e)}$', xy=(0.7,0.84),xycoords='axes fraction', fontsize=size_plots_label)
ax5.set_xlabel('$q_i$_32 [kg/kg]', fontsize=size_plots_label)
ax5.set_ylabel('', fontsize=size_plots_label)
ax5.tick_params(labelsize=size_plots_label)
plt.ylim(qv_ylim)
plt.savefig('figures/shap_clc_32_all_plots.pdf')
# plt.show()
# -
# All subplots pertain only SHAP values for clc_32!
#
# For each input feature, the SHAP values were first averaged over all 10000 NARVAL R02B05 samples.
# If we now focus on a specific feature, such as q_v, then we can draw a plot like the one shown in **b)**.
# We do this for every seed, so we can see some error bars in **(b)**.
# If we sum up all SHAP values shown in **(b)**, we receive the first bar in plot **(a)**, including the error bars.
#
# Now why is the Model bias the sum of all SHAP values divided by the number of samples? <br>
# -> Well it's because we want to explain the average bias of the model on layer 32, not the one produced if we multiply the bias by the number of parameters!
#
# **d) and e):** <br>
# Each dot shows one sample: A pair of input feature and corresponding SHAP value
# Thick lines show the average SHAP values conditioned on small bins (respectively cover around 1/10 of the size of the range of input values).
# We do not plot it for qi_32 > 2 kg/kg due to the low density of points there. The NARVAL samples we evaluate SHAP on are exactly the same for the QUBICC and the NARVAL model.
| additional_content/plots_offline_paper/SHAP_cumulative_contribution_plot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="9vyzYqcR2lCe" outputId="6b377ddd-e8a3-4f98-f304-2bec3d3a505d"
import numpy
x=numpy.array([342,546,234,665,452])
x.min()
# + colab={"base_uri": "https://localhost:8080/"} id="LJ45SWcG2vN_" outputId="b95f9102-2ec3-4e98-f510-60c9c9895903"
x.max()
# + colab={"base_uri": "https://localhost:8080/", "height": 283} id="yTq-M7Tj2zyk" outputId="02fab8fa-2832-4cc3-937d-c44d065f6a06"
x=[1,2,3,4,5,6]
y=[3,34,45,67,76,93]
import matplotlib.pyplot as ABC
ABC.plot(x,y)
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="9NpAK9p370AA" outputId="fad47e1b-99b7-4ff2-ecd1-e5ffbb87ebbc"
x={"kerala":"trivandrum","tamil nadu":"chennai"}
x["kerala"]
# + colab={"base_uri": "https://localhost:8080/"} id="tyYuKgaR9X3W" outputId="cf64a8c7-c6ee-4e12-d019-8c0960572f47"
x.keys()
# + colab={"base_uri": "https://localhost:8080/"} id="Ifc6EZQC9ntR" outputId="0637cfee-1728-41ab-e1b7-ee37e307aece"
x.values()
# + id="GyDeKF2b9qXa"
x={"kerala":"trivandrum","tamil nadu":"madras"}
x["tamil nadu"]="chennai"
# + colab={"base_uri": "https://localhost:8080/"} id="upvZodSf-SLz" outputId="45ac8f48-0cd3-451a-f0c5-63d2bc2d99ef"
x
# + colab={"base_uri": "https://localhost:8080/"} id="VQ3eZOFn-Xh6" outputId="3fed113a-dd57-4293-faf3-c21c06273b6f"
x["karnataka"]="bengaluru"
x
# + id="YrDd9gZ4_Epe"
del x["karnataka"]
# + colab={"base_uri": "https://localhost:8080/"} id="QS3pXa57_Iej" outputId="f163a9d1-3e06-412d-eff3-b47a41fa668f"
x
# + id="_x0pRqaG_JvN"
import numpy as AB
# + id="0fCwIL97BuCZ"
x=numpy.array([[1,2,3,4,5],[4,6,3,7,4]])
# + colab={"base_uri": "https://localhost:8080/"} id="6TxJr1CEByEM" outputId="29c01abb-2f30-4d75-9dd7-36ebfa229211"
x
# + colab={"base_uri": "https://localhost:8080/"} id="uf9v8AlGBzYV" outputId="666d5b9c-a6dc-40c5-e7b4-d3f3094f2be7"
x.shape
# + colab={"base_uri": "https://localhost:8080/"} id="0h777-jfB17o" outputId="5e7f7824-f4a9-456e-e460-570d1e8c6298"
x[0:]
# + id="OeIZBNi6B4so"
y=[1,2,4,5]
# + colab={"base_uri": "https://localhost:8080/"} id="4j9LEr_YB80Q" outputId="9b319f86-3338-4eea-ac4a-38fbcf7cf0d4"
y[0]
# + id="WdBtl_ThB-ry"
| numpy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.7 64-bit
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
df = pd.read_json('real_experiences_m_ch.jsonl', lines=True)
df.head()
# # Time point
#
# Time point is current point in time since the start of the shift in seconds.
# **My interpretation** order timestamp - shift starts (in seconds)
df[['shift_start', 'shift_end','order_timestamp']] = df[['shift_start','shift_end', 'order_timestamp']].apply(pd.to_datetime, infer_datetime_format=True)
df['current_point_in_time'] = (df['order_timestamp'] - df['shift_start']).dt.total_seconds()
df['time_limit_of_the_shift'] = (df['shift_end'] - df['order_timestamp']).dt.total_seconds()
df.head()
# # Time budget
# Extract estimated arrival to depot
#
# Conditions:
# 1. len(unloaded_drops) != 0
# 2. last visit is to depot
time = pd.to_datetime(df.route_update.values[0]['unloaded_drops'][-1]['estimated_arrival'])
time_series = pd.Series(time)
def extract_arrival_time_to_depot(route_update, shift_end):
if len(route_update['unloaded_drops']) == 0:
raise ValueError(f"unloaded drops cannot be empty")
last_drop = route_update['unloaded_drops'][-1]['drop_type']
if last_drop != "DEPOT_VISIT":
raise ValueError(f"last unloaded drop should be depot visit")
estimated_arrival = pd.to_datetime(route_update['unloaded_drops'][-1]['estimated_arrival'], infer_datetime_format=True)
return (shift_end - estimated_arrival).total_seconds()
df['time_budget'] = df.apply(lambda x: extract_arrival_time_to_depot(x['route_update'], x['shift_end']), axis=1)
df.head()
# # Flexibility
def extract_flexibility(route_update, shift_end, order_timestamp):
customer_drops = []
for drop in range(len(route_update['unloaded_drops'])):
if route_update['unloaded_drops'][drop]['drop_type'] == 'CUSTOMER_DROP':
customer_drops.append(route_update['unloaded_drops'][drop])
if len(customer_drops) == 0:
return (shift_end - order_timestamp).total_seconds() # Not sure
summed_time = 0
for customer in customer_drops:
arrival = pd.to_datetime(customer['estimated_arrival'], infer_datetime_format=True)
deadline = pd.to_datetime(customer['delivery_window']['end'], infer_datetime_format=True)
diff = (deadline - arrival).total_seconds()
summed_time += diff
return summed_time / len(customer_drops)
df['flexibility'] = df.apply(lambda x: extract_flexibility(x['route_update'], x['shift_end'], x['order_timestamp']), axis=1)
df.head()
# # Average distance per customer
def average_distance_per_customer(route_update):
customer_drops = []
summed_distance = 0
for drop in range(len(route_update['unloaded_drops'])):
if route_update['unloaded_drops'][drop]['drop_type'] == 'CUSTOMER_DROP':
customer_drops.append(route_update['unloaded_drops'][drop])
summed_distance += route_update['unloaded_drops'][drop]['leg_distance']
n_customers = len(customer_drops)
if not n_customers:
return 0
return summed_distance / n_customers
df['avg_distance_per_customer'] = df.apply(lambda x: average_distance_per_customer(x['route_update']), axis=1)
df.head()
# # Time budget in worst case
def time_budget_in_worst_case(route_update, shift_end):
time = []
for drop in range(len(route_update['unloaded_drops'])):
arrival = pd.to_datetime(route_update['unloaded_drops'][drop]['estimated_arrival'], infer_datetime_format=True)
time.append(arrival)
mean = np.mean(pd.Series(time))
std = np.std(pd.Series(time))
return (shift_end - (mean + 2 * std)).total_seconds()
df['time_budget_in_worst_case'] = df.apply(lambda x: time_budget_in_worst_case(x['route_update'], x['shift_end']), axis=1)
df.head()
# # Baseline
from sklearn.linear_model import SGDRegressor, LinearRegression
from sklearn.metrics import mean_squared_error
from sklearn.preprocessing import StandardScaler
train = df.loc[df.episode_index < 19]
train = train[['revenue', 'time_budget', 'flexibility', 'avg_distance_per_customer', 'time_budget_in_worst_case']]
test = df.loc[df.episode_index >= 19]
test = test[['revenue', 'time_budget', 'flexibility', 'avg_distance_per_customer', 'time_budget_in_worst_case']]
X = train.copy()
y = X.pop('revenue')
# Preview of the training data:
X.head()
y_real = test.pop('revenue')
reg = LinearRegression().fit(X, y)
y_pred = reg.predict(test)
mean_squared_error(y_real, y_pred)
reg.coef_
reg.intercept_
import matplotlib.pyplot as plt
# +
plt.figure(figsize=(10,10))
plt.scatter(y_real, y_pred, c='crimson')
p1 = max(max(y_pred), max(y_real))
p2 = min(min(y_pred), min(y_real))
plt.plot([p1, p2], [p1, p2], 'b-')
plt.xlabel('True Values', fontsize=15)
plt.ylabel('Predictions', fontsize=15)
plt.axis('equal')
plt.show()
| model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Informes de mortalidad
#
# Actualizado diariamente, este documento se [visualiza mejor aquí](https://nbviewer.jupyter.org/github/jaimevalero/COVID-19/blob/master/jupyter/Momo.ipynb).
#
# Datos del Sistema de Monitorización de la Mortalidad diaria, que incluye las defunciones por todas las causas procedentes de 3.929 registros civiles informatizados, que representan el 92% de la población española.
#
#
# Cargamos datos
import Loading_data
from matplotlib import pyplot as plt
import warnings
warnings.filterwarnings('ignore')
import pandas as pd
from IPython.display import display, HTML
df = pd.read_csv('https://momo.isciii.es/public/momo/data')
df.to_csv('/tmp/momo.csv')
df.head()
# +
import janitor
import datetime
def pipeline_basic_with_query(df,query):
''' Basic filtering, using janitor
Carga de datos, enriquecimiento de fechas y filtro por query configurable
'''
LISTA_COLUMNAS_A_BORRAR = ['Unnamed: 0',
'defunciones_observadas_lim_inf',
'defunciones_observadas_lim_sup',
'defunciones_esperadas',
'defunciones_esperadas_q01',
'defunciones_esperadas_q99']
return (
df
# Quitar: columnas
.remove_columns(LISTA_COLUMNAS_A_BORRAR)
.clean_names()
# Enriquecer: fechas con columnas de años, mes y año-mes
.rename_column( "fecha_defuncion", "date")
.to_datetime('date')
.join_apply(lambda x: x['date'].strftime('%Y') , new_column_name="date_year" )
.join_apply(lambda x: x['date'].strftime('%m') , new_column_name="date_month" )
.join_apply(lambda x: x['date'].strftime('%m-%d') , new_column_name="date_month_day" )
.join_apply(lambda x: x['date'].strftime('%U') , new_column_name="date_week" )
.join_apply(lambda x: x['date'].strftime('%Y-%m') , new_column_name="date_year_month" )
.join_apply(lambda x: x['date'].strftime('%Y-%U') , new_column_name="date_year_week" )
# Filtrar:por query
.filter_on( query )
.set_index('date')
)
def pipeline_basic(df):
query = 'ambito == "nacional" & nombre_gedad == "todos" & nombre_sexo == "todos" '
return pipeline_basic_with_query(df,query)
def extraer_defunciones_anuales_por_periodo(periodo_de_tiempo,query):
'''Extrae el cuadro de comparativa por week, or year '''
def pipeline_agregado_anual(periodo_de_tiempo,df,year):
''' Saca un dataframe de los datos agrupados por año'''
return (
df
.filter_on('date_year == "'+year+'"' )
.groupby_agg( by='date_'+periodo_de_tiempo, agg='sum', agg_column_name="defunciones_observadas", new_column_name="agregados")
.rename_column( "agregados", year)
.join_apply(lambda x: x['date_'+periodo_de_tiempo] , new_column_name=periodo_de_tiempo )
.set_index('date_'+periodo_de_tiempo)
[[periodo_de_tiempo,year]]
.drop_duplicates()
)
def pipeline_comparativa_anual(periodo_de_tiempo,df_2018,df_2019,df_2020):
''' Mergea tres dataframes de año, por periodo de tiempo'''
return (
df_2018
.merge( df_2019, on=periodo_de_tiempo, how='right')
.merge( df_2020, on=periodo_de_tiempo, how='left')
.sort_naturally(periodo_de_tiempo)
.set_index(periodo_de_tiempo)
.join_apply(lambda x: x['2020'] - x['2019'] , new_column_name="resta 2020 y 2019" )
)
# Sacamos los datos y limpiamos
df = pd.read_csv('')
df_basic = pipeline_basic_with_query(df,query)
# Sacamos los datos agrupados por años
muertes_2018 = pipeline_agregado_anual(periodo_de_tiempo,df=df_basic,year='2018')
muertes_2019 = pipeline_agregado_anual(periodo_de_tiempo,df=df_basic,year='2019')
muertes_2020 = pipeline_agregado_anual(periodo_de_tiempo,df=df_basic,year='2020')
# Generamos un solo cuadro, con columna por año
df_comparativa_años = pipeline_comparativa_anual(periodo_de_tiempo,muertes_2018,muertes_2019,muertes_2020)
return df_comparativa_años
def debug_extraer_defunciones_anuales_por_periodo():
""" Solo para depurar"""
query = 'ambito == "nacional" & nombre_gedad == "todos" & nombre_sexo == "todos" '
df_muertes_anuales_por_semana = extraer_defunciones_anuales_por_periodo("week",query)
df_muertes_anuales_por_mes = extraer_defunciones_anuales_por_periodo("month",query)
return df_muertes_anuales_por_semana , df_muertes_anuales_por_mes
#df1, df2 = debug_extraer_defunciones_anuales_por_periodo()
#df1
# -
# ## Sacamos el grafico comparativo de fallecimiento, para los años 2019 y 2020, por semana
# +
from matplotlib import pyplot as plt
from IPython.display import display, HTML
import pandas as pd
import numpy as np
periodo_de_tiempo="week"
query = 'ambito == "nacional" & nombre_gedad == "todos" & nombre_sexo == "todos" '
df = extraer_defunciones_anuales_por_periodo(periodo_de_tiempo,query)
fig = plt.figure(figsize=(8, 6), dpi=80)
plt.xticks(rotation=90)
for ca in ['2018','2019','2020']:
plt.plot(df[ca])
plt.legend(df.columns)
plt.xlabel(periodo_de_tiempo)
plt.ylabel("Deaths by " + periodo_de_tiempo)
fig.suptitle('Comparativa de fallecimientos por año, según MOMO', fontsize=20)
plt.show()
periodo_de_tiempo="week"
query = 'ambito == "nacional" & nombre_gedad == "todos" & nombre_sexo == "todos" '
df = extraer_defunciones_anuales_por_periodo(periodo_de_tiempo,query)
df.style.format({"2020": "{:20,.0f}",
"2018": "{:20,.0f}",
"2019": "{:20,.0f}",
"resta 2020 y 2019": "{:20,.0f}", }).background_gradient(cmap='Wistia',subset=['resta 2020 y 2019'])
# +
def get_current_year_comparison(query):
"""Saca muertos del año en curso en el ambito como argumento"""
df = pd.read_csv('/tmp/momo.csv')
df = pipeline_basic_with_query(df,query)
semana_actual = df.tail(1).date_week.values[0]
year_actual = df.tail(1).date_year.values[0]
date_month_day_actual= df.tail(1).date_month_day.values[0]
year_last = str(int(year_actual)-1)
death_this_year_today = df.query( f"date_year == '{year_actual}' ").defunciones_observadas.sum()
deaht_last_year_today = df.query( f"date_year == '{year_last}' and date_month_day <= '{date_month_day_actual}' ").defunciones_observadas.sum()
deaths_this_year_excess = death_this_year_today - deaht_last_year_today
return deaths_this_year_excess
query = f""" ambito == "nacional" & nombre_gedad == "todos" & nombre_sexo == "todos" """
deaths_this_year_excess = get_current_year_comparison(query)
display(HTML(f"<h4 id='excedente'>Excdente de muertes de este año, respecto al año anterior:</h4><h2>{deaths_this_year_excess:,.0f} </h2>"))
# -
query = f""" nombre_ambito == "Madrid, Comunidad de" & nombre_gedad == "todos" & nombre_sexo == "todos" """
deaths_this_year_excess = get_current_year_comparison(query)
display(HTML(f"<h4 id='excedentemadrid'>Excedente de muertes de este año en Madrid, respecto al año anterior:</h4><h2>{deaths_this_year_excess:,.0f} </h2>"))
# +
# Sacamos las muertes en madrid de hombres y de mujeres
import numpy as np
import seaborn as sns
def pipeline_comparativa_semestral_diaria(df):
return (
df
.filter_on(" defunciones_observadas > 0")
.remove_columns(['nombre_gedad','ambito','cod_ambito','cod_ine_ambito','nombre_ambito','cod_sexo','cod_gedad','date_year','date_week','date_month','date_year_week'])
.rename_column( "nombre_sexo" , "sexo")
.rename_column( "date_year_month", "mes")
)
# Sacamos los datos de 2019
df = pd.read_csv('/tmp/momo.csv')
query = ' date_year == "2019" & nombre_ambito == "Madrid, Comunidad de" & nombre_gedad == "todos" & nombre_sexo != "todos" & date_month < "13" '
df_madrid_2019 = pipeline_basic_with_query(df,query)
df_madrid_2019 = pipeline_comparativa_semestral_diaria(df_madrid_2019)
# Sacamos los datos de 2020
df = pd.read_csv('/tmp/momo.csv')
query = ' date_year == "2020" & nombre_ambito == "Madrid, Comunidad de" & nombre_gedad == "todos" & nombre_sexo != "todos" & date_month < "13" '
df_madrid_2020 = pipeline_basic_with_query(df,query)
df_madrid_2020 = pipeline_comparativa_semestral_diaria(df_madrid_2020)
df_madrid_2019
# +
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
display(HTML("<h2>Distribucion muertes en Madrid </h2>"))
display(HTML("<h3>Comparativa de defunciones, entre el primer semestre de 2019 y el del 2020</h3>"))
f, axes = plt.subplots(1 , 2 ,figsize=(16, 7), sharex=True)
sns.despine(left=True)
# Mismo limites, para poder comparar entre años
axes[0].set_ylim([0,500])
axes[1].set_ylim([0,500])
sns.violinplot(x="mes", y="defunciones_observadas", hue="sexo",
data=df_madrid_2019, split=True, scale="count", ax=axes[0] )
sns.violinplot(x="mes", y="defunciones_observadas", hue="sexo",
data=df_madrid_2020, split=True, scale="count", ax=axes[1])
# -
# Aux functions
def print_categorical_variables(df):
""" Get a dict with categorical variables"""
my_dict = {}
cols = df.columns
num_cols = df._get_numeric_data().columns
# Show categorical values
categorical = list(set(cols) - set(num_cols))
for i in categorical :
if 'echa' not in i.lower() : my_dict[i] = df[i].unique()
return my_dict
df = pd.read_csv('/tmp/momo.csv')
my_dict = print_categorical_variables(df)
my_dict
momo2020 = pd.read_csv("/root/scripts/COVID-19/data/momo2019.csv", sep='\t',)
# +
periodo_de_tiempo="week"
query = 'ambito == "nacional" & nombre_gedad == "todos" & nombre_sexo == "todos" '
df = extraer_defunciones_anuales_por_periodo(periodo_de_tiempo,query)
# -
df = pd.read_csv('/tmp/momo.csv')
periodo_de_tiempo="week"
year=2021
def pipeline_agregado_anual(periodo_de_tiempo,df,year):
''' Saca un dataframe de los datos agrupados por año'''
return (
df
.filter_on('date_year == "'+year+'"' )
.groupby_agg( by='date_'+periodo_de_tiempo, agg='sum', agg_column_name="defunciones_observadas", new_column_name="agregados")
.rename_column( "agregados", year)
.join_apply(lambda x: x['date_'+periodo_de_tiempo] , new_column_name=periodo_de_tiempo )
.set_index('date_'+periodo_de_tiempo)
[[periodo_de_tiempo,year]]
.drop_duplicates()
)
pipeline_agregado_anual)
| jupyter/.ipynb_checkpoints/Momo-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Basic usage of SFRmaker in a scripting context
# This example illustrates basic usage of SFRmaker in a scripting context. For examples of using SFRmaker with a configuration file, see the MERAS and Tyler Forks examples.
import numpy as np
import matplotlib.pyplot as plt
import flopy
import sfrmaker
# ### Input requirements
# The most basic input requirements of SFRmaker are hydrography and a model grid. Optionally, a model and a DEM can be input, as demonstrated below. See the [documentation](https://aleaf.github.io/sfrmaker/inputs.html) for a more detailed description of inputs.
#
# #### Hydrography
# In this example, we will use data that has been downloaded from [NHDPlus](https://nhdplus.com/NHDPlus/NHDPlusV2_data.php). The original file structure in the download has been maintained, allowing us to simply supply SFRmaker with a path to the NHDPlus files:
NHDPlus_paths = '../tylerforks/NHDPlus/'
# If we were dealing with more than one drainage basin, the NHDPlus file paths could be included in a list:
NHDPlus_paths_list = ['/NHDPlusGL/NHDPlus04/',
'/NHDPlusMS/NHDPlus07/']
# ### Creating a ``Lines`` instance from NHDPlus
# The ``sfrmaker.Lines`` class includes functionality for reading and processing hydrography flowlines. This example shows how to create a ``Lines`` instance from NHDPlus data.
#
# For large hydrography datasets, it is advantageous to filter the data when it is read in. The ``filter`` argument to ``sfrmaker.Lines`` accepts a shapefile path or tuple of bounding box coordinates. In either case, a bounding box tuple is created and passed to the ``filter`` method in the [``fiona`` package](https://fiona.readthedocs.io/en/latest/manual.html), which is fast.
lns = sfrmaker.Lines.from_nhdplus_v2(NHDPlus_paths='../tylerforks/NHDPlus/',
filter='../tylerforks/grid.shp')
# Alternatively, ``sfrmaker.Lines`` can be instantiated with separate arguments for each NHDPlus file used:
lines = sfrmaker.Lines.from_nhdplus_v2(NHDFlowlines='../tylerforks/NHDPlus/NHDSnapshot/Hydrography/NHDFlowline.shp',
PlusFlowlineVAA='../tylerforks/NHDPlus/NHDPlusAttributes/PlusFlowlineVAA.dbf',
PlusFlow='../tylerforks/NHDPlus/NHDPlusAttributes/PlusFlow.dbf',
elevslope='../tylerforks/NHDPlus/NHDPlusAttributes/elevslope.dbf',
filter=(-90.625, 46.3788, -90.4634, 46.4586))
# ### Creating a ``Lines`` instance from custom hydrography
# Alternatively, a ``Lines`` instance can be created from any hydrography that includes the pertinent attribute fields, which must be specified:
custom_lines = sfrmaker.Lines.from_shapefile(shapefile='../meras/flowlines.shp',
id_column='COMID', # arguments to sfrmaker.Lines.from_shapefile
routing_column='tocomid',
width1_column='width1',
width2_column='width2',
up_elevation_column='elevupsmo',
dn_elevation_column='elevdnsmo',
name_column='GNIS_NAME',
attr_length_units='feet', # units of source data
attr_height_units='feet', # units of source data
)
# ### Specifying a model grid from a flopy `StructuredGrid` instance
# The next step is to specify a model grid. One option is to specify a flopy `StructuredGrid` instance. SFRmaker will then use this internally to create an instance of its own `StructuredGrid` class. In this case, the row and column spacing must be provided in the units of the projected coordinate reference system (CRS) that the model is in, which is typically meters. In this case, our model grid spacing is 250 feet, so we have to convert.
#
# Specifying a CRS (via the ``proj4`` argument) is also important, as it allows SFRmaker to automatically reproject any input data to the same CRS as the model grid. [The best way to do this is with an EPSG code](https://proj.org/faq.html#what-is-the-best-format-for-describing-coordinate-reference-systems), as shown below.
#
# See the flopy documentation for more details about `StructuredGrid`.
# +
delr = np.array([250 * 0.3048] * 160) # cell spacing along a row
delc = np.array([250 * 0.3048] * 111) # cell spacing along a column
flopy_grid = flopy.discretization.StructuredGrid(delr=delr, delc=delc,
xoff=682688, yoff=5139052, # lower left corner of model grid
angrot=0, # grid is unrotated
# projected coordinate system of model (UTM NAD27 zone 15 North)
proj4='epsg:26715'
)
# -
# ### Specifying a model grid from a shapefile
# Another option is to create an SFRmaker `StructuredGrid` directly using a shapefile. While the basic underpinings are in place for SFRmaker to support unstructured grids, this option hasn't been fully implemented yet.
#
# Attribute fields with row and column information must be specified. An polygon defining the area where the SFR network will be created can optionally be specified here, or later in the creation of ``SFRData`` (see below).
grid = sfrmaker.StructuredGrid.from_shapefile(shapefile='../tylerforks/grid.shp',
icol='i', # attribute field with row information
jcol='j', # attribute field with column information
active_area='../tylerforks/active_area.shp'
)
# With specification of ``active_area``, the grid created above has an **``isfr``** array attribute designating which cells can have SFR:
plt.imshow(grid.isfr, interpolation='nearest')
# ### Using the ``modelgrid`` attached to a flopy model
# If no grid is supplied as input, SFRmaker will try to use ``modelgrid`` attribute attached to a supplied flopy model instance. This only works if ``modelgrid`` is valid. Loading a flopy model with a valid model grid requires the grid information to be specified in the namefile header, and that the model be in the same units as the projected CRS. See the flopy documentation for more details
# ### Specifying a model
# While a model is not required to run SFRmaker, specifying a model is advantageous in that it allows SFRmaker to assign valid model layers for reaches. Models are specified as flopy model instances, which can be loaded or created from scratch (see the flopy documentation). In this case, we are loading a model:
m = flopy.modflow.Modflow.load('tf.nam', model_ws='../tylerforks/tylerforks')
m
# ### Creating an SFRData instance
# The ``sfrmaker.SFRData`` class is the primary object for creating or modifying an SFR dataset. A ``SFRData`` instance can be created from the ``Lines`` class using the ``to_sfr()`` method.
#
# Either a flopy ``StructuredGrid`` or sfrmaker ``StructuredGrid`` can be supplied. While MODFLOW and flopy support specification of length units, these aren't always specified in the model input files, so it is good practice to specifiy the units explicitly to SFRmaker.
sfrdata = lines.to_sfr(grid=flopy_grid, model=m, model_length_units='feet')
# ### The ``SFRData`` class
# Internally, the SFRData class mostly uses the data model for the MODFLOW-2005 style SFR2 package (which includes MODFLOW-NWT), which organizes the input into segments and reaches. Segment and reach data are stored in the ``reach_data`` and ``segment_data`` attribute DataFrames. In addition to the MODFLOW-2005 data, reach numbers and their routing connections (as needed for MODFLOW-6) are also stored. On writing of SFR package input, MODFLOW-6 style input can be created via the ``sfrmaker.mf5to6`` module.
sfrdata.reach_data.head()
sfrdata.segment_data.head()
# ### Sampling streambed top elevations from a DEM
# While the above dataset contains streambed top elevations read from NHDPlus, a DEM can be sampled to obtain more accurate elevations. If the DEM elevation units are specified, SFRmaker will convert the elevations to model units if needed. See the Streambed_elevation_demo notebook for more information on how this works.
sfrdata.set_streambed_top_elevations_from_dem('../tylerforks/dem_26715.tif', dem_z_units='meters')
# ### Assigning layers to the reaches
# Once we have a valid set of streambed elevations, the reaches can be assigned to model layers.
sfrdata.assign_layers()
sfrdata.reach_data.head()
# ### Running diagnostics
# `SFRData` includes a `run_diagnostics()` method that executes the Flopy checker on the SFR package input. While the Flopy checks are only implemented for MODFLOW-2005 style SFR packages, `run_diagnostics()` works on MODFLOW-6 packages as well, by working of an attached MODFLOW-2005 representation of the MODFLOW-6 SFR input. The following checks are executed:
# * **NaNs** (not a number values, which will cause MODFLOW to crash)
# * **consecutive segment and reach numbering**
# * **segment numbering that only increases** in the downstream direction
# * **circular routing sequences** (reaches or segments routing back to themselves)
# * **routing connection proximity:** The length of routing connections, as determined by the distance between their cell centers, is compared to 1.25 times the diagonal distance across the cell containing the upstream reach. This benchmark length is equivalent to a diagonal connection between two square cells, where one cell had a spacing of 1.5 times the other cell (the maximum recommended increase in spacing between two adjacent cells; see Anderson and others, 2015). While this routing proximity check can be helpful for identifying egregious routing issues, in practice, it may not be uncommon for valid routing connections to violate this rule. For example, if the ``one_reach_per_cell`` option is used in constructing the SFR package with a large grid size, there may be numerous routing connections that extend across several cells. When non-adjacent routing connections are identified, the user can check the shapefile of routing connections to verify the results.
# * **overlapping conductances:** This check identifies instances of collocated reaches, where more than one reach has a connection to the groundwater flow solution in that cell. Collocated reaches may or may not present an issue for a particular model solution, but can potentially promote instability or spurious circulation of flow between the collocated reaches.
# * **spurious streambed top elevations:** Values < -10 or > 15,000 are flagged
# * **downstream rises in streambed elevation**
# * **inconsistencies between streambed elevation and the model grid:** This check looks for streambed bottoms below their respective cell bottoms, which will cause MODFLOW to halt execution, and streambed tops that are above the model top. The latter issue will not prevent MODFLOW from running, but may indicate problems with the input if the differences are large. For coarse grids in areas of high stream density (where there are often multiple streams in a cell), it may be impossible to ensure that all streambed top elevations are at or below the model top. Oftentimes, these reaches are dry anyways (for example, and ephemeral gully that runs off a bluff into a perennial stream, where the cell elevation is based on the elevation of the perennial stream). In any case, it is good practice to check the largest violations by comparing the SFRmaker shapefile output to a DEM and other data in a GIS environment.
# * **suprious slopes:** slopes of less than 0.0001 (which can lead to excessive stages with the SFR package's Manning approximation) and greater than 1.0 are flagged.
sfrdata.run_diagnostics(verbose=False)
# ### Writing an SFR package
# Now we can write the package. By default, a MODFLOW-2005 style package is written to the current working directory or the model workspace of the attached model instance.
sfrdata.write_package()
# ### Writing a MODFLOW-6 SFR package
# Alternatively, a MODFLOW-6 style SFR package can be written by specifying ``version='mf6'``
sfrdata.write_package(version='mf6')
# ### Writing tables
# The ``reach_data`` and ``segment_data`` tables can be written out to csv files for further processing:
sfrdata.write_tables()
# and then read back in to create an ``SFRData`` instance
sfrdata2 = sfrmaker.SFRData.from_tables('tables/tf_sfr_reach_data.csv', 'tables/tf_sfr_segment_data.csv',
grid=grid)
# ### Writing shapefiles
# Shapefiles can also be written to visualize the SFR package. These include:
# * hydrography linestrings associated with each reach
# * model cell polygons associated with each reach
# * outlet locations where water is leaving the model
# * routing connections (as lines drawn between cell centers)
# * the locations of specified inflows or any other period data
# * observation locations (if observations were supplied as input)
sfrdata.write_shapefiles()
| examples/Notebooks/SFRmaker_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [Task Scheduler](https://leetcode.com/problems/task-scheduler/)。给一大写字母的数组,代表$26$种任务的集合。CPU任一时间片可以挑选任一任务去执行,但是同种任务的两次执行之间需要有$n$个时间片的间隔。问执行完所有任务最少需要多少时间片。如```tasks = ["A","A","A","B","B","B"]```,```n = 2```,执行时间片为:```A -> B -> idle -> A -> B -> idle -> A -> B```。
#
# 思路:首先对同种任务进行计数,同种任务之间的执行至少要间隔$n$,那么易得最短时间取决于数量最多的那种任务。易得完成$k$个同种任务至少需要$(k-1)*(n+1)+1$个时间片,若数量最多的任务有$s$种,那么至少需要$(k-1)*(n+1)+s$个时间片。
def leastInterval(tasks, n: int) -> int:
res = len(tasks)
cnts = [0]*26
for ch in tasks:
cnts[ord(ch)-65] += 1
cnts.sort(reverse=True)
neet_t = (cnts[0]-1)*(n+1)
for cnt in cnts:
if cnt == cnts[0]:
neet_t += 1
return max(res, neet_t)
# [Fraction to Recurring Decimal](https://leetcode.com/problems/fraction-to-recurring-decimal/)。将分数转换成小数形式,当出现循环小数时,将循环部分加上括号。
#
# 思路:首先明确分数中分子为除数,分母为被除数。手动做一下除法就不难发现,当某一次的余数在之前出现过时,肯定就会出现循环小数,所以该题解法是先正常的做除法,并记录每一次余数出现的位置,当余数重复时即出现循环小数。
def fractionToDecimal(numerator: int, denominator: int) -> str:
res = '-' if numerator*denominator < 0 else '' # 符号
numerator, denominator = abs(numerator), abs(denominator)
integer, remainder = divmod(numerator, denominator) # 首个商与余数
res += str(integer)
if remainder:
res += '.'
else:
return res
remainder_idx = dict() # 记录余数位置的字典
while remainder:
if remainder not in remainder_idx:
remainder_idx[remainder] = len(res)
integer, remainder = divmod(remainder*10, denominator)
res += str(integer)
else:
idx = remainder_idx[remainder]
res = '{}({})'.format(res[:idx], res[idx:])
break
return res
# 求倒数第$Q$个排列,**2019网易笔试题**。给$n$个自然数,然后指定其中一个排列,假设选中的是第$Q$个排列,求倒数第$Q$个排列。如$n=3$,选中排列$123$,它是第$1$个排列,那么要求倒数第$i$个排列$321$。
#
# 思路:找规律,第$Q$个排列和倒数第$Q$个排列,对应位的数字相加恰好等于$n+1$。
def func(n, num):
num = list(str(num))
res = list()
for bit in num:
res.append(n+1-int(bit))
return int(''.join(map(str, res)))
# [Beautiful Arrangement II](https://leetcode.com/problems/beautiful-arrangement-ii/)。给$n$个正整数,重排这些正整数,使得任意两相邻值的绝对差值只有$k$种取值。如$[1,2,3]$只有一种绝对差值$1$。
#
# 思路:找规律。易得```func(4,1)=[1,2,3,4]```,```func(4,2)=[4,1,2,3]```,```func(4,3)=[1,4,2,3]```。发现交替放置$k$个大小值如```[1,4,3,2]```可以构成$k-1$种差值,而令最后一种差值为$1$,顺序放置剩余的元素即可。在$[1,n]$区间设置首位双指针,若$k$为奇数,则放置$i$;若$k$为偶数则放置$j$。
def constructArray(n: int, k: int):
res = list()
i, j = 1, n
while i <= j:
if k > 1:
if k & 1: # 奇数
res.append(i)
i += 1
else: # 偶数
res.append(j)
j -= 1
k -= 1
else:
res.append(i)
i += 1
return res
# [Number of Digit One](https://leetcode.com/problems/number-of-digit-one/)。求$[1,n]$范围内$1$出现的次数。
#
# 思路:直接取出最大值$n$,由低往高依次将每一位都置成$1$,然后判断高位与地位各有多少种取值。设由掩码$mask$提取出的数字是$idx$,有以下三种情况;
# - $idx=0$,该位若要置$1$的话那么高位数字只能先减$1$,高位数字在$[0,high-1]$任取,低位数字任取,即$high\times{mask}$;
# - $idx=1$,当高位数字在$[0,high-1]$时,低位数字任取,当高位数字取最大值时,低位只能在$[0,low]$里取值,即$high\times{mask}+low+1$;
# - $idx>1$,高位数字可在$[0,high]$中任取,低位数字任取,即$(high+1)\times{mask}$。
# +
def countDigitOne(n: int) -> int:
res = 0
mask = 1
while mask <= n:
idx = n//mask % 10
high = n//mask//10
low = n % mask
if idx == 0:
res += high*mask
elif idx == 1:
res += high*mask+low+1
else:
res += (high+1)*mask
mask *= 10
return res
countDigitOne(11)
# -
# [Nth Digit](https://leetcode.com/problems/nth-digit/)。给一自然数的顺序排序,求该字符串中的第$n$位数字。
#
# 思路:$1$位数有$9$个数字,$2$位数有$90$个数字,$3$位数有$900$个数字。按该规律$n$位数有$9\times{10}^{n-1}$个数字。
def findNthDigit(n: int) -> int:
if n < 10:
return n
bits = 2
n -= 10
while n > 9*(10**(bits-1))*bits:
n -= 9*(10**(bits-1))*bits
bits += 1
start = 10**(bits-1) # bits位数字的起点数字,1,10,100,...
number = start+n//bits # 第n位在哪一个数字中
return str(number)[n % bits]
# [Count Primes](https://leetcode.com/problems/count-primes/)。计数$n$以内的质数个数。
#
# 思路:设置一个长度为$n$的状态数组,以倍数法来标记非质数,然后对该数组求和即可。有一个数学定理,$n$以内的最大质数不会超过$\sqrt{n}$。
def countPrimes(self, n: int) -> int:
if n < 2:
return 0
import math
thresh = int(math.sqrt(n))
isPrime = [True]*n
isPrime[0] = isPrime[1] = False # 0,1不算质数
for i in range(2, thresh+1):
if isPrime[i]:
isPrime[i*i:n:i] = [False]*len(isPrime[i*i:n:i])
return sum(isPrime)
# [Roman to Integer](https://leetcode.com/problems/roman-to-integer/)。**2019老虎证券笔试题**。罗马数字转阿拉伯数字。
#
# 思路:找规律题。如$IV=4$而$VI=6$,若两罗马字符$ab$中$a\ge{b}$,说明$a$是正的,反之表明$a$是负的。
def romanToInt(s: str) -> int:
lookup = {'I': 1, 'V': 5, 'X': 10, 'L': 50,
'C': 100, 'D': 500, 'M': 1000}
res = 0
for i in range(len(s)-1):
if lookup[s[i]] < lookup[s[i+1]]:
res -= lookup[s[i]]
else:
res += lookup[s[i]]
return res+lookup[s[-1]]
# [Gray Code](https://leetcode.com/problems/gray-code/)。指定二进制的位数$n$,输出在该位数下的十进制格雷码序列。
#
# 思路:纯规律题。
# $$
# \begin{align}
# n=0 \ & [0] \\
# n=1 \ & [0,1] \\
# n=2 \ & [00,01,11,10] \\
# n=3 \ & [000,001,011,010,110,111,101,100] \\
# \end{align}
# $$
# 观察到每下一级的格雷码可以对半分成两份,前半部分与上一级数字相同,只是在高位加$0$;而后半部分则是上一级序列的逆序,然后再高位加$1$,相当于加$2^{(i-1)}$。
def grayCode(n: int):
res=[0]
if n==0:
return res
for i in range(1,n+1):
res.extend([item+2**(i-1) for item in res[::-1]])
return res
# [1-bit and 2-bit Characters](https://leetcode.com/problems/1-bit-and-2-bit-characters/)。现有两种特殊字符,第一种字符被编码成$0$,第二种字符编码成$10$和$11$。现给一编码串,判断该编码串的是否是以第一种字符结尾。
#
# 思路:两种编码有明显的区别,第一种字符编码以$0$开头,一位;第二种字符编码以$1$开头,两位。使用工作指针线性扫描字串,遇到$0$前进一位,遇到$1$前进两位,判断指针能否停在最后一位即可。
def isOneBitCharacter(bits) -> bool:
n = len(bits)
idx = 0
while idx < n-1:
if bits[idx] == 0:
idx += 1
else:
idx += 2
return idx == n-1
# [Self Dividing Numbers](https://leetcode.com/problems/self-dividing-numbers/)。给定一个范围,求出范围内所有的SDN。SDN定义为能被自身中所有数字整除的数,并且SDN不允许包含0。
#
# 思路:逻辑实现题。
def selfDividingNumbers(left: int, right: int):
res=list()
if left>right:
return res
for num in range(left,right+1):
tmp=num
while tmp:
bit=tmp%10
if bit==0 or num%bit!=0:
break
else:
tmp//=10
if tmp==0:
res.append(num)
return res
# 判断点是否在多边形内。**2019去哪儿笔试题**。首先给出用户坐标,后跟的是多边形的点的坐标,保证顺序。判断该点是否在多边形内部。
#
# 思路:经典计算几何题,思路来源于[网上](https://www.cnblogs.com/anningwang/p/7581545.html)。由于笔试不允许使用Python,所以只测试了两个最简单的用例,不保证正确性。
# +
import sys
def func(x_u, y_u, xs, ys):
res = False
n = len(xs)
i, j = 0, n - 1
while i < n:
if ((ys[i] > y_u) != (ys[j] > y_u)
and (x_u < (xs[j] - xs[i]) * (y_u - ys[i]) / (ys[j] - ys[i]) + xs[i])):
res = not res
j = i
i += 1
return res
coord = sys.stdin.readline().strip().split()
x_u, y_u = list(map(float, coord[0].split(',')))
xs = list(map(float, [item.split(',')[0] for item in coord[1:]]))
ys = list(map(float, [item.split(',')[1] for item in coord[1:]]))
print(xs, ys, x_u, y_u)
print(func(x_u, y_u, xs, ys))
| Algorithm/OnlineExam/Python/Math.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# !pip install tensorflow
# +
import numpy as np
def batch(inputs, max_sequence_length=None):
"""
Args:
inputs:
list of sentences (integer lists)
max_sequence_length:
integer specifying how large should `max_time` dimension be.
If None, maximum sequence length would be used
Outputs:
inputs_time_major:
input sentences transformed into time-major matrix
(shape [max_time, batch_size]) padded with 0s
sequence_lengths:
batch-sized list of integers specifying amount of active
time steps in each input sequence
"""
sequence_lengths = [len(seq) for seq in inputs]
batch_size = len(inputs)
if max_sequence_length is None:
max_sequence_length = max(sequence_lengths)
inputs_batch_major = np.zeros(shape=[batch_size, max_sequence_length], dtype=np.int32) # == PAD
for i, seq in enumerate(inputs):
for j, element in enumerate(seq):
inputs_batch_major[i, j] = element
# [batch_size, max_time] -> [max_time, batch_size]
inputs_time_major = inputs_batch_major.swapaxes(0, 1)
return inputs_time_major, sequence_lengths
def random_sequences(length_from, length_to,
vocab_lower, vocab_upper,
batch_size):
""" Generates batches of random integer sequences,
sequence length in [length_from, length_to],
vocabulary in [vocab_lower, vocab_upper]
"""
if length_from > length_to:
raise ValueError('length_from > length_to')
def random_length():
if length_from == length_to:
return length_from
return np.random.randint(length_from, length_to + 1)
while True:
yield [
np.random.randint(low=vocab_lower,
high=vocab_upper,
size=random_length()).tolist()
for _ in range(batch_size)
]
# +
import numpy as np
import tensorflow as tf
tf.reset_default_graph()
sess = tf.InteractiveSession()
# -
| src/nlpia/book/examples/Untitled3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
from cloudmesh.common.Shell import Shell
from pprint import pprint
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib
sns.set_theme(style="whitegrid")
import pandas as pd
# +
experiments = 10
points = [10,
100,
1000,
10000]
points = points + list(range(100000, 1000001, 50000))
print (points)
df = pd.DataFrame(
{"Size": points}
)
df = df.set_index('Size')
# -
df
for experiment in range(0,experiments):
label="gregor-mac"
log = f"result-{label}-{experiment}.log"
os.system(f"rm {log}")
for n in points:
print (n)
os.system (f"mpiexec -n 4 python count-click.py --n {n} --max_number 10 --find 8 --label gregor-mac"
f"| tee -a {log}")
lines = Shell.grep("csv,Result:", log)
print(lines)
values = []
times = []
for line in lines.splitlines():
msg = line.split(",")[7]
t = line.split(",")[4]
total, overall, trials, find, label = msg.split(" ")
values.append(int(overall))
times.append(float(t))
print (t, overall)
#data = pd.DataFrame(values, times, columns=["Values", "Time"])
#print (data.describe())
#sns.lineplot(data=data, palette="tab10", linewidth=2.5)
# df["Size"] = values
df[f"Time_{experiment}"] = times
print(df)
# +
df = df.rename_axis(columns="Time")
df
# -
sns.lineplot(data=df, markers=True)
plt.show()
plt.savefig(f'benchmark-{label}.png')
plt.savefig(f'benchmark-{label}.pdf')
plt.show()
| examples/count/sweep-fancy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
#-*- coding: utf-8 -*-
import object_detector.file_io as file_io
import cv2
import numpy as np
import object_detector.utils as utils
import matplotlib.pyplot as plt
import progressbar
import pandas as pd
class Evaluator(object):
def __init__(self):
self._recall_precision = None
self._dataset = None
def eval_average_precision(self, test_image_files,
annotation_path,
detector,
window_dim, window_step, pyramid_scale):
"""Public function to calculate average precision of the detector.
Parameters
----------
test_image_files : list of str
list of test image filenames to evaluate detector's performance
annotation_path : str
annotation directory path for test_image_files
detector : Detector
instance of Detector class
window_dim : list
(height, width) order of sliding window size
window_step : list
(height_step, width_step) order of sliding window step
pyramid_scale : float
scaling ratio of building image pyramid
Returns
----------
average_precision : float
evaluated score for the detector and test images on average precision.
Examples
--------
"""
patches = []
probs = []
gts = []
# setup the progress bar
widgets = ["Running for each Test image as gathering patches and its probabilities: ",
progressbar.Percentage(), " ", progressbar.Bar(), " ", progressbar.ETA()]
pbar = progressbar.ProgressBar(maxval=len(test_image_files), widgets=widgets).start()
for i, image_file in enumerate(test_image_files):
test_image = cv2.imread(image_file)
test_image = cv2.cvtColor(test_image, cv2.COLOR_BGR2GRAY)
boxes, probs_ = detector.run(test_image,
window_dim, window_step, pyramid_scale,
threshold_prob=0.0,
show_result=False,
show_operation=False)
truth_bb = self._get_truth_bb(image_file, annotation_path)
ious = self._calc_iou(boxes, truth_bb)
is_positive = ious > 0.5
patches += boxes.tolist()
probs += probs_.tolist()
gts += is_positive.tolist()
pbar.update(i)
pbar.finish()
probs = np.array(probs)
gts = np.array(gts)
self._calc_precision_recall(probs, gts)
average_precision = self._calc_average_precision()
return average_precision
def plot_recall_precision(self):
"""Function to plot recall-precision graph.
It should be performed eval_average_precision() before this function is called.
"""
range_offset = 0.1
if self._recall_precision is None:
raise ValueError('Property _recall_precision is not calculated. To calculate this, run eval_average_precision() first.')
recall_precision = self._recall_precision
plt.plot(recall_precision[:, 0], recall_precision[:, 1], "r-")
plt.plot(recall_precision[:, 0], recall_precision[:, 1], "ro")
plt.axis([0 - range_offset, 1 + range_offset, 0 - range_offset, 1 + range_offset])
plt.xlabel("recall")
plt.ylabel("precision")
plt.show()
@property
def dataset(self):
if self._dataset is None:
raise ValueError('Property _dataset is not calculated. To calculate this, run eval_average_precision() first.')
d = {"probability": self._dataset[:,0], 'ground truth': self._dataset[:,1].astype(np.bool_)}
df = pd.DataFrame(data=d, columns = ["probability", 'ground truth'])
return df
def _calc_average_precision(self):
inter_precisions = []
for i in range(11):
recall = float(i) / 10
inter_precisions.append(self._calc_interpolated_precision(recall))
return np.array(inter_precisions).mean()
def _calc_precision_recall(self, probs, ground_truths):
probs = np.array(probs)
ground_truths = np.array(ground_truths)
dataset = np.concatenate([probs.reshape(-1,1), ground_truths.reshape(-1,1)], axis=1)
dataset = dataset[dataset[:, 0].argsort()[::-1]]
n_gts = len(dataset[dataset[:, 1] == 1])
n_relevant = 0.0
n_searched = 0.0
recall_precision = []
for data in dataset:
n_searched += 1
if data[1] == 1:
n_relevant += 1
recall = n_relevant / n_gts
precision = n_relevant / n_searched
recall_precision.append((recall, precision))
if recall == 1.0:
break
self._dataset = dataset
self._recall_precision = np.array(recall_precision)
def _calc_interpolated_precision(self, desired_recall):
recall_precision = self._recall_precision
inter_precision = recall_precision[recall_precision[:,0] >= desired_recall]
inter_precision = inter_precision[:, 1]
inter_precision = max(inter_precision)
return inter_precision
def _calc_iou(self, boxes, truth_box):
y1 = boxes[:, 0]
y2 = boxes[:, 1]
x1 = boxes[:, 2]
x2 = boxes[:, 3]
y1_gt = truth_box[0]
y2_gt = truth_box[1]
x1_gt = truth_box[2]
x2_gt = truth_box[3]
xx1 = np.maximum(x1, x1_gt)
yy1 = np.maximum(y1, y1_gt)
xx2 = np.minimum(x2, x2_gt)
yy2 = np.minimum(y2, y2_gt)
w = np.maximum(0, xx2 - xx1 + 1)
h = np.maximum(0, yy2 - yy1 + 1)
intersections = w*h
As = (x2 - x1 + 1) * (y2 - y1 + 1)
B = (x2_gt - x1_gt + 1) * (y2_gt - y1_gt + 1)
ious = intersections.astype(float) / (As + B -intersections)
return ious
# Todo : extractor module과 중복되는 내용 제거
def _get_truth_bb(self, image_file, annotation_path):
image_id = utils.get_file_id(image_file)
annotation_file = "{}/annotation_{}.mat".format(annotation_path, image_id)
bb = file_io.FileMat().read(annotation_file)["box_coord"][0]
return bb
# -
| _writing/object detector.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Make predictions
#
# This script uses the classifiers with the highest accuracy to get the LIs and predictions for all cases
# ### import modules
# + tags=["hide-cell"]
# %matplotlib inline
# +
import os
import fnmatch
import numpy as np
import pandas as pd
from scipy import stats
import matplotlib.pylab as plt
import seaborn as sns
from sklearn.svm import SVC
from sklearn import preprocessing, model_selection, metrics
from nilearn import plotting
import pickle
# -
# ### get absolute directory of project
# +
# after converstion to .py, we can use __file__ to get the module folder
try:
thisDir = os.path.realpath(__file__)
# in notebook form, we take the current working directory (we need to be in 'notebooks/' for this!)
except:
thisDir = '.'
# convert relative path into absolute path, so this will work with notebooks and py modules
supDir = os.path.abspath(os.path.join(os.path.dirname(thisDir), '..'))
supDir
# -
sns.set_style('white')
sns.set_context('poster')
# ### load labeler
myLabeler = pickle.load(open('%s/models/myLabeler.p' % supDir, 'rb'))
# + tags=["hide-cell"]
myLabeler.classes_
# -
# ### collect the parameters that allow for above-chance prediction
aboveDf = pd.read_csv('%s/models/aboveDf_clf_1d_drop.csv' % supDir,index_col=[0,1])
# + tags=["hide-cell"]
aboveDf.T
# -
clfDict = pickle.load(open('%s/models/clf_1d_drop.p' % supDir))
# + tags=["hide-cell"]
clfDict[95][3-0]
# -
# ### predictions for one value and one parameter set
def makePred(x, roiPc, tThresh, clfDict, myLabeler=myLabeler):
thisClf = clfDict[roiPc][tThresh]['clf']
thisScaler = clfDict[roiPc][tThresh]['scaler']
xArr = np.array(x)
xScaled = thisScaler.transform(xArr.reshape(1, -1 * xArr.shape[-1]))
y_pred = thisClf.predict_proba(xScaled)
df = pd.DataFrame(y_pred).T
idx = [myLabeler.inverse_transform([x])[-1] for x in thisClf.classes_]
df.index = idx
return df
# Example:
# + tags=["hide-cell"]
thesePreds = makePred([0.0], 0, 3, clfDict)
# + tags=["hide-cell"]
thesePreds
# -
# ### predictions for one patient, for all above-chance parameters
def changeDf(df):
idx1 = df.columns.get_level_values(0).astype(float)
idx2 = df.columns.get_level_values(1)
mIdx = pd.MultiIndex.from_arrays([idx2, idx1])
df.columns = mIdx
df.sort_index(axis=1,inplace=True)
return df
# Example Patient:
# + tags=["hide-cell"]
pCsv = '%s/data/interim/csv/roiLaterality_pat0399_b.csv' % supDir
# + tags=["hide-cell"]
pName = 'pat_%s' % (pCsv.split('_pat')[-1].split('.')[0])
pName
# + tags=["hide-cell"]
pDf = pd.read_csv(pCsv, index_col=[0], header=[0, 1])
pDf = changeDf(pDf)
# + tags=["hide-cell"]
pDf.tail()
# -
def getP(pDf, pName, roiSize, thresh, dims, myLabeler=myLabeler):
if dims == 1:
liValue = pDf.loc[roiSize, 'LI'].loc[thresh]
thisDf = pd.DataFrame([liValue], index=[pName], columns=['LI'])
elif dims == 2:
diffValue = pDf.loc[roiSize, 'L-R'].loc[thresh]
diffDf = pd.DataFrame([diffValue], index=[pName], columns=['L-R'])
addValue = pDf.loc[roiSize, 'L+R'].loc[thresh]
addDf = pd.DataFrame([addValue], index=[pName], columns=['L+R'])
thisDf = pd.concat([diffDf, addDf], axis=1)
return thisDf
# + tags=["hide-cell"]
getP(pDf, pName, roiSize=50, thresh=5.8, dims=1)
# -
def makeBestPreds(pCsv, aboveDf, clfDict, dims):
pName = 'pat_%s' % (pCsv.split('_pat')[-1].split('.')[0])
pDf = pd.read_csv(pCsv, index_col=[0], header=[0, 1])
pDf = changeDf(pDf)
valueDict = {}
predDf = pd.DataFrame()
# here we loop through the aboveDf, which has in its index
# all parameters that we want
# get the table with the roi size
for pc in aboveDf.index.levels[0]:
# get the data for the threshold
for t in aboveDf.loc[pc].index:
thisParam = getP(pDf, pName, pc, t, dims)
# store the value
thisVals = list(thisParam.loc[pName])
valueDict[str(pc) + '_' + str(t)] = thisVals
# make predictions, these are like df's
try:
thisPred = makePred(thisVals, pc, t, clfDict)
except:
thisPred = pd.DataFrame({
'bilateral': 0,
'left': 0,
'right': 0,
'inconclusive': 1
},
index=[0]).T
#store predictions
thisPred = thisPred.T
thisPred.index = [str(pc) + '_' + str(t)]
predDf = pd.concat([predDf, thisPred])
if dims == 1:
valueDf = pd.DataFrame(valueDict, index=['LI']).T
elif dims == 2:
valueDf = pd.DataFrame(valueDict, index=['L-R', 'L+R']).T
# average
meanValueDf = pd.DataFrame(valueDf.mean())
meanPredDf = pd.DataFrame(predDf.mean())
meanDf = pd.concat([meanValueDf,meanPredDf]).T
meanDf.index = [pName]
return valueDf, predDf, meanDf
# Example:
# + tags=["hide-cell"]
valueDf, predDf, meanDf = makeBestPreds(pCsv, aboveDf, clfDict, dims=1)
# + tags=["hide-cell"]
meanDf
# + tags=["hide-cell"]
fuDf = predDf.copy()
fuDf.index = pd.MultiIndex.from_tuples(list([x.split('_') for x in fuDf.index]))
# -
def changeIdx(df):
idx1 = df.index.get_level_values(0).astype(int)
idx2 = df.index.get_level_values(1).astype(float)
mIdx = pd.MultiIndex.from_arrays([idx2, idx1])
df.index = mIdx
df.sort_index(axis=0,inplace=True)
return df
# + tags=["hide-cell"]
fuDf = changeIdx(fuDf)
# + tags=["hide-cell"]
fig = plt.figure(figsize=(16,6))
for i,c in enumerate(fuDf.columns):
ax = plt.subplot(1,fuDf.columns.shape[-1],i+1)
thisDf = fuDf.loc[:,[c]].unstack()[c].T
sns.heatmap(thisDf,cmap='rainbow',vmin=0,vmax=1,axes=ax)
ax.set_title(c)
plt.tight_layout()
plt.show()
# -
def makeAllComputations(pCsv, dims, drop, sigLevel=0.001):
dropStr = ['full', 'drop'][drop]
dimStr = ['1d', '2d'][dims - 1]
# load the classifier
clfDict = pickle.load(
open('%s/models/clf_%s_%s.p' % (supDir, dimStr, dropStr), 'rb'))
accDict = pickle.load(
open('%s/models/acc_%s_%s.p' % (supDir, dimStr, dropStr), 'rb'))
aboveDf = pd.read_csv(
'%s/models/aboveDf_clf_%s_%s.csv' % (supDir, dimStr, dropStr),
index_col=[0, 1])
# compute
valueDf, predDf, meanDf = makeBestPreds(pCsv, aboveDf, clfDict, dims=dims)
# if we compute the 1-dimensional LI and do not want to model inconclusive cases,
# we still need to handle cases where division by zero occurs
# therefore, we compute the proportion of cases where neither of the 3 main classes was predicted
if dims == 1 and drop == True:
meanDf.loc[:,'inconclusive'] = 1 - meanDf.loc[:,['left','bilateral','right']].sum(axis=1)
return valueDf, predDf, meanDf
# + tags=["hide-cell"]
valueDf, predDf, meanDf = makeAllComputations(pCsv, dims=2, drop=True)
# + tags=["hide-cell"]
valueDf.tail()
# + tags=["hide-cell"]
predDf.tail()
# + tags=["hide-cell"]
meanDf
# -
# ### do all variations
from datetime import datetime
# + tags=["hide-cell"]
def makeP(pCsv):
pName = 'pat%s' % (pCsv.split('_pat')[-1].split('.')[0])
bigDf = pd.DataFrame()
for myDim in [1, 2]:
for myDrop in [True, False]:
dimStr = ['1d', '2d'][myDim - 1]
dropStr = ['full', 'drop'][myDrop]
#print myDim, myDrop, datetime.now()
valueDf, predDf, meanDf = makeAllComputations(
pCsv, dims=myDim, drop=myDrop)
valueDf.to_csv('%s/data/processed/csv/values_%s_%s_%s.csv' %
(supDir, pName, dimStr, dropStr))
predDf.to_csv('%s/data/processed/csv/predictions_%s_%s_%s.csv' %
(supDir, pName, dimStr, dropStr))
meanDf.index = pd.MultiIndex.from_arrays([[dimStr], [dropStr]])
bigDf = pd.concat([bigDf, meanDf])
bigDf.to_csv('%s/data/processed/csv/meanTable_%s.csv' % (supDir, pName))
return bigDf
# + tags=["show-cell"]
#def makeP(pFolder, pName):
#
# pCsv = '%s/roiLaterality_%s.csv' % (pFolder, pName)
#
# bigDf = pd.DataFrame()
#
# for myDim in [2]:
# for myDrop in [False]:
#
# dimStr = ['1d', '2d'][myDim - 1]
# dropStr = ['full', 'drop'][myDrop]
#
# valueDf, predDf, meanDf = makeAllComputations(
# pCsv, dims=myDim, drop=myDrop)
#
# valueDf.to_csv(
# '%s/values_%s_%s_%s.csv' % (pFolder, pName, dimStr, dropStr))
# predDf.to_csv('%s/predictions_%s_%s_%s.csv' % (pFolder, pName,
# dimStr, dropStr))
#
# meanDf.index = pd.MultiIndex.from_arrays([[dimStr], [dropStr]])
# bigDf = pd.concat([bigDf, meanDf])
#
# bigDf.to_csv('%s/meanTable_%s.csv' % (pFolder, pName))
#
# return bigDf
# + tags=["hide-cell"]
meanDf = makeP(pCsv)
# + tags=["hide-cell"]
meanDf
# -
# ## do this for all patients
# ### collect all patients
# + tags=["hide-cell"]
my_train = pickle.load(open('../models/my_nest.p', 'rb'))
my_test = pickle.load(open('../models/my_test.p', 'rb'))
my_all = my_train + my_test
len(my_all)
# + tags=["hide-cell"]
csvList = [
'../data/interim/csv/%s' % x for x in os.listdir('../data/interim/csv/')
if x.startswith('roiLaterality_pat')
]
csvList.sort()
# + tags=["hide-cell"]
def makeDf(csvList,trainOrTest):
df = pd.DataFrame()
for pat in csvList:
for lab in trainOrTest:
if lab[-1] in pat:
thisDf = pd.DataFrame([pat], index=[[lab[0]], [lab[1]]])
df = pd.concat([df, thisDf])
df.columns = ['csv']
df.sort_index(inplace=True)
return df
# + tags=["hide-cell"]
dfAll = makeDf(csvList,trainOrTest=my_all)
# + tags=["hide-cell"]
dfAll.shape
# + tags=["hide-cell"]
dfAll.tail()
# -
# ### run for all patients
# + tags=["hide-cell"]
doneList = [
x.split('meanTable_')[-1].split('.')[0]
for x in os.listdir('%s/data/processed/csv/' % supDir)
if x.startswith('meanTable_')
]
len(doneList)
# + tags=["hide-cell"]
for p in dfAll.index:
if p[1] not in doneList:
pCsv = dfAll.loc[p,'csv']
print datetime.now(),pCsv
meanDf = makeP(pCsv)
| notebooks/09-mw-apply-classifier-to-data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# # Support Vector Machine with Polynomial Kernel
# +
from sklearn import datasets
import numpy as np
import pandas as pd
from sklearn.linear_model import LinearRegression
from sklearn import svm, preprocessing
raw_data=pd.read_csv("sampled3.csv")
print (raw_data.head(3))
npMatrix = np.matrix(raw_data)
X, y = npMatrix[1:,1:34], npMatrix[1:,34:]
# +
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
sc.fit(X)
X = sc.transform(X)
# +
from sklearn.cross_validation import train_test_split
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.3)
# -
clf=svm.SVR(kernel = 'poly')
clf.fit(X_train, y_train[:,0])
clf.score(X_test, y_test[:,0])
| SVM_poly_kernel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="zrEyXRrG5iNp"
# # UST Into the Wormhole
# > A decentralised stablecoin into a decentralised bridge
#
# - toc:true
# - branch: master
# - badges: true
# - comments: false
# - author: <NAME>
# - categories: [Terra, Wormhole]
# - hide: false
#
# + colab={"base_uri": "https://localhost:8080/", "height": 283} id="BpIzaSvW5hgs" outputId="a52bd53f-2a22-4243-f0ba-14cc29bbbdab"
#hide
#Imports & settings
# !pip install plotly --upgrade
import pandas as pd
import plotly.express as px
import plotly.graph_objects as go
from plotly.subplots import make_subplots
# %matplotlib inline
# %load_ext google.colab.data_table
# %load_ext rpy2.ipython
# %R options(tidyverse.quiet = TRUE)
# %R options(lubridate.quiet = TRUE)
# %R options(jsonlite.quiet = TRUE)
# %R suppressMessages(library(tidyverse))
# %R suppressMessages(library(lubridate))
# %R suppressMessages(library(jsonlite))
# %R suppressMessages(options(dplyr.summarise.inform = FALSE))
# + id="JCW7BPHs52rP"
#hide
# %%R
#Grab wormhole bridge query from Flipside
df_wh = fromJSON('https://api.flipsidecrypto.com/api/v2/queries/d0c057eb-cbe7-49e4-8f5a-77b3672105e6/data/latest', simplifyDataFrame = TRUE)
#fix the column names
names(df_wh)<-tolower(names(df_wh))
#Change the date to date format
df_wh$block_timestamp <- parse_datetime(df_wh$block_timestamp)
df_wh <- df_wh %>% rename(bridge_chain_id = wormhole_chain_id)
#join chain ids
chains <- tibble(bridge_chain_id = c(0,1,2,3,4,5,6),
bridge_chain = c('Unknown','Solana','Ethereum','Terra','BSC','Polygon','Harmony'))
df <- df_wh %>% left_join(chains, by = "bridge_chain_id")
#create a date field
df$date <- floor_date(df$block_timestamp, unit = 'day')
# clip by date to remove the last part date
#df <- df %>% filter(date < '2021-11-17')
#Grab the token labels
labels <- read_csv("https://github.com/scottincrypto/analytics/raw/master/data/wormhole_bridge_assets.csv", show_col_types = FALSE)
#join the token lables
df <- df %>%
left_join(labels, by = "denom")
tx_by_day <- df %>%
group_by(date) %>%
summarise(qty = sum(amount),
tx_count = n(),
av_tx_size = qty / tx_count)
tx_by_day_excl_network_acc <- df %>%
filter(user != 'terra1dtzfwgzt8xa70zkm8gqzwz0n4zrqtngdpqejx5') %>%
group_by(date) %>%
summarise(qty = sum(amount),
tx_count = n(),
av_tx_size = qty / tx_count)
#unique wallets
unique_wallets <- df %>%
filter(user != 'terra1dtzfwgzt8xa70zkm8gqzwz0n4zrqtngdpqejx5') %>%
group_by(user) %>%
summarise(first_date = min(date)) %>%
ungroup() %>%
group_by(first_date) %>%
summarise(unique_wallets = n()) %>%
mutate(cum_unique = cumsum(unique_wallets),
growth_rate = unique_wallets / cum_unique * 100)
#split by chain
tx_by_dest_chain <- df %>%
filter(user != 'terra1dtzfwgzt8xa70zkm8gqzwz0n4zrqtngdpqejx5') %>%
group_by(bridge_chain, date) %>%
summarise(qty = sum(amount),
tx_count = n(),
av_tx_size = qty / tx_count)
tx_by_wallet <- df %>%
group_by(user) %>%
summarise(qty = sum(amount),
tx_count = n(),
av_tx_size = qty / tx_count)
sol_by_day_of_week <- tx_by_dest_chain %>%
filter(bridge_chain == 'Solana') %>%
filter(date != ymd('2021-11-04')) %>%
mutate(weekday = weekdays(date, abbreviate = F),
sort = wday(date)) %>%
group_by(weekday, sort) %>%
summarise(av_qty = mean(qty)) %>%
arrange(sort)
# + [markdown] id="UnEwm_pE9NkW"
# # Sending UST into the Wormhole
#
# The Wormhole Bridge went live on the Terra network at the end of September 2021. Wormhole is a cross chain network which allows traffic between blockchains with different consensus mechanisms. This is done in a decentralised manner, avoiding the need for the centralised liquidity pools which power many existing bridges. When the Wormhole bridge connected to Terra, it allowed connectivity between Terra, Solana, Ethereum, Polygon and Binance Smart Chain. This was a huge step forward for a multi-chain world, as Terra and Solana are very different technologies to the other three EVM compatible chains.
#
# Initial use of the Wormhole bridge to & from Terra was light but is steadily increasing. As was outlined in [this post,](https://scottincrypto.github.io/analytics/terra/wormhole/2021/11/17/A-Flight-Through-a-Terra-Wormhole.html) most of the traffic passing over the Wormhole Bridge to & from Terra is UST - the decentralised stablecoin native to Terra. This post will look more deeply at UST being sent from Terra over the Wormhole Bridge.
# + [markdown] id="h54CspAJA6La"
# # How much UST is being sent?
#
# The graph below shows the amount of UST sent from the Terra network since the Wormhole Bridge was attached. The graph is a little curious - there is a background rate of a million or so UST per day, puncuated by some very large volume days. This is worth some exploration.
# + colab={"base_uri": "https://localhost:8080/", "height": 511} id="ufxQ-_R1VLNI" outputId="5e8cf8b7-2023-4fe6-b569-ca9bfc5cf4cd"
#hide_input
#UST Sent via Wormhole
# df_p = %R tx_by_day
fig = px.bar(df_p
, x = "date"
, y = "qty"
, labels=dict(date="Date", qty="UST Sent")
, title= "UST Sent via Wormhole"
, template="simple_white", width=800, height=800/1.618
)
fig.update_layout(legend=dict(
yanchor="top",
y=0.99,
xanchor="right",
x=0.99,
title_text=None
))
fig.update_yaxes(title_text='UST')
fig.update_xaxes(title_text=None)
fig.show()
# + [markdown] id="Ir7idhK3CRRy"
# ## Investigating the big sends
#
# The table below shows the top 10 senders of UST over the Wormhole Bridge in the time period of the chart above. Notice the top user has sent over $124m UST in only 12 transactions. The next largest user has sent 84m UST, but over many more transactions.
#
# In investigating user our top wallet, terra..ejx5, we identified a single transaction which was 42m UST in size. View this on [finder.terra.money here](https://finder.terra.money/mainnet/tx/C1213BE51F64E9F320C0D39B0AAF36C289CF586BE2496E80463B95BED87CB22D).
#
# We know this transaction went to Ethereum (recipient chain 2) but the recipient address is obfuscated in the transaction so we can't easily identify it. By looking for transactions on the Ethereum Wormhole Bridge contract at around the time of this transaction, we found this one: [this one](https://etherscan.io/tx/0xb827b16a9125dcbeffa839bd5be65bd35690b176e0f868abe4fa5afd31d18f2c). It shows our 42m UST, which is now a Wormhole wrapped version of UST, being claimed at the other end. Further transactions on this wallet identify that the 42m UST was [deposited as liquidity](https://etherscan.io/tx/0xf3d25987e15237905c7efa87332a50ce83cf8dc704b00804551bdcea763c8bcb) into the Shuttle/Wormhole Pool. This Pool on the Ethereum network serves to exchange wrapped UST tokens from the legacy Shuttle bridge for Wormhole wrapped UST ready for use on the new bridge. This pool is described in the [Wormhole documentation](https://medium.com/terra-money/wormhole-v2-for-terra-the-ui-walkthrough-595ca6649ae8) and is not a for-profit dex pool. Instead, this is part of the infrastructure used to eventually convert from the Shuttle bridge to the Wormhole bridge permanently.
#
# Our conclusion is that our big sending address, terra..ejx5, belongs to the one of the Terra/Wormhole teams and is being used for network operations to stand up the bridge. Since we are looking at usage, it's worthwhile to exclude this address from any further analysis as it doesn't represent organic usage of the bridge.
#
# + colab={"base_uri": "https://localhost:8080/", "height": 305} id="HSW2qwIrCRw9" outputId="7cc69987-2604-44d5-c181-af5c4bf7469b"
#hide_input
# %R tx_by_wallet %>% arrange(desc(qty)) %>% head(10)
# + [markdown] id="NUX9SL_dJIks"
# # How much UST is being sent - revisited
#
# The graph below shows the amount of UST being sent per day from the Terra network via the Wormhole bridge, this time excluding the network operations account we identified earlier. Once the bridge ramped up for the first week, it has settled into a range of 400k -> 7m per day, with a typical day being around 1.5m UST sent to other networks.
# + colab={"base_uri": "https://localhost:8080/", "height": 511} id="P75QhmG7JQma" outputId="a92fde26-915f-47e0-d456-4fda42ce5da9"
#hide_input
#UST Sent via Wormhole expluding the network liquidity account
# df_p = %R tx_by_day_excl_network_acc
fig = px.bar(df_p
, x = "date"
, y = "qty"
, labels=dict(date="Date", qty="UST Sent")
, title= "UST Sent via Wormhole, network ops count excluded"
, template="simple_white", width=800, height=800/1.618
)
fig.update_layout(legend=dict(
yanchor="top",
y=0.99,
xanchor="right",
x=0.99,
title_text=None
))
fig.update_yaxes(title_text='UST')
fig.update_xaxes(title_text=None)
fig.show()
# + [markdown] id="3A_4C3i7K4du"
# # Where is the UST going?
#
# As mentioned in the intro, the Wormhole bridge connects Terra to 4 other chains - Ethereum, Solana, Polygon and Binance Smart Chain (BSC). The table below shows how much UST has gone to each chain since the bridge began (excluding the network ops account). We can see that most of the UST sent has gone to Solana - 57m UST, around 83% of all the UST traffic. Ethereum is next largest with 8m UST or 12%, and Polygon & BSC having comparatively small amounts.
# + colab={"base_uri": "https://localhost:8080/", "height": 176} id="xjGcwz5nKbLt" outputId="f2b189d8-f42d-4ef5-ecf0-893491b09bed"
#hide_input
# %R tx_by_dest_chain %>% group_by (bridge_chain) %>% summarise(qty_sent = sum(qty)) %>% mutate(percentage = qty_sent / sum(qty_sent)*100 )%>% arrange(desc(qty_sent))
# + [markdown] id="STABzaaBMeq9"
# The graph below shows the UST sent by destination chain plotted daily, excluding the network ops account. We see a few large transactions dominating BSC, Ethereum and Polygon, but the network traffic over Solana is both larger and more consistent than the EVM chains. It appears that sending UST to Solana is a use case that the market was looking for, and has been met with the Terra->Solana Wormhole bridge.
# + colab={"base_uri": "https://localhost:8080/", "height": 511} id="l-0n4nZiG_k5" outputId="acb542d5-be23-425a-ef61-bcf06b34ed38"
#hide_input
# User Counts
# df_p = %R tx_by_dest_chain %>% select(-tx_count) %>% select (-av_tx_size) %>% pivot_wider(names_from = bridge_chain, values_from = qty, values_fill = 0) %>% arrange(date)
fig = make_subplots(rows=2, cols=2, subplot_titles=("BSC", "Ethereum", "Polygon", "Solana"))
fig.append_trace(go.Bar(x=df_p["date"], y=df_p["BSC"], name="UST"), row=1, col=1)
fig.append_trace(go.Bar(x=df_p["date"], y=df_p["Ethereum"], name="UST"), row=2, col=1)
fig.append_trace(go.Bar(x=df_p["date"], y=df_p["Polygon"], name="UST"), row=1, col=2)
fig.append_trace(go.Bar(x=df_p["date"], y=df_p["Solana"], name="UST"), row=2, col=2)
fig.update_layout(width=800, height=800/1.618, title_text="UST Sent by Destionation Chain")
fig.update_layout(template="simple_white", showlegend=False)
fig.update_yaxes(title_text='Value Bridged UST', row=1, col=1)
fig.update_yaxes(title_text='Value Bridged UST', row=2, col=1)
fig.show()
# + [markdown] id="_X3fkAeTNi5H"
# # Estimating User Count
# User count is challenging in blockchain environments as people may operate multiple wallets. The fallback for this problem is to use unique wallet addresses, which gives us an upper bound on the number of users. For wallets sending UST via the Wormhole bridge, we get the following profile of unique accounts over time. At the time of publication, just under 2000 unique wallets were recorded sending UST over Wormhole
# + colab={"base_uri": "https://localhost:8080/", "height": 511} id="rTK3UZH_VvsS" outputId="23120664-e586-4023-f95a-1adc7d966847"
#hide_input
#cumulative wallets
# df_p = %R unique_wallets
fig = px.bar(df_p
, x = "first_date"
, y = "cum_unique"
, labels=dict(date="Date", av_tx_size="Av Tx Size")
, title= "Unique Wallets Sending UST over Wormhole Bridge"
, template="simple_white", width=800, height=800/1.618
)
fig.update_layout(legend=dict(
yanchor="top",
y=0.99,
xanchor="right",
x=0.99,
title_text=None
))
fig.update_yaxes(title_text='Wallet Count')
fig.update_xaxes(title_text=None)
fig.show()
# + [markdown] id="70G3_QPUYJaT"
# The unique wallet growth rate is consistent but steady, ranging from 1.5% -> 3% daily after some rapid initial growth
# + colab={"base_uri": "https://localhost:8080/", "height": 511} id="7rd1aXkHWdaX" outputId="c8628f47-d1c2-466e-e52e-d6ff5d48ec5a"
#hide_input
#cumulative wallets growth rate
# df_p = %R unique_wallets
fig = px.bar(df_p
, x = "first_date"
, y = "growth_rate"
, labels=dict(date="Date", av_tx_size="Av Tx Size")
, title= "Growth rate of unique wallets sending UST over Wormhole Bridge"
, template="simple_white", width=800, height=800/1.618
)
fig.update_layout(legend=dict(
yanchor="top",
y=0.99,
xanchor="right",
x=0.99,
title_text=None
))
fig.update_yaxes(title_text='Growth Rate')
fig.update_xaxes(title_text=None)
fig.show()
# + [markdown] id="Kfzf21aeaFPo"
# # Transaction Size
#
# The charts below show the average transaction sizes of UST sent to each of the target chains, excluding the network ops address. Each chain has a small number of large transactions which skew the data on those days. Outside of these, we see relatively small values bridged to BSC. This makes sense in the context that BSC has very low transaction fees, and bridging a few hundred UST is still relatively efficient. This contrasts with Ethereum - the lower bound of the average transaction is closer to 2-3000 UST. This shows that users are sensitive to the Ethereum gas fees associated with claiming the tokens on the other side of the bridge which will typically be on the order of 100-200 USD.
#
# Polygon is an outlier when compared with BSC and Ethereum. With low transaction fees, it might be expected that there are small transaction sizes sent to Polygon in line with BSC. Instead we see that the typical day has transaction sizes in the 10-20k UST range. This can be explained by a relatively small number of users doing large transactions, indicating that retail users haven't discovered this pathway to Polygon just yet.
#
# Solana transaction sizes stand out compared to the other chains. They are more consistent, ranging from 5k UST on a slow day to 30-60k on a bigger day. There are no days with zero or very low transaction size, supporting the higher usage shown above.
# + colab={"base_uri": "https://localhost:8080/", "height": 511} id="IQxxJQo5aVik" outputId="2f408efa-c026-4def-c1bf-0153856d92d4"
#hide_input
# Tx size
# df_p = %R tx_by_dest_chain %>% select(-tx_count) %>% select (-qty) %>% pivot_wider(names_from = bridge_chain, values_from = av_tx_size, values_fill = 0) %>% arrange(date)
fig = make_subplots(rows=2, cols=2, subplot_titles=("BSC", "Ethereum", "Polygon", "Solana"))
fig.append_trace(go.Bar(x=df_p["date"], y=df_p["BSC"], name="UST"), row=1, col=1)
fig.append_trace(go.Bar(x=df_p["date"], y=df_p["Ethereum"], name="UST"), row=2, col=1)
fig.append_trace(go.Bar(x=df_p["date"], y=df_p["Polygon"], name="UST"), row=1, col=2)
fig.append_trace(go.Bar(x=df_p["date"], y=df_p["Solana"], name="UST"), row=2, col=2)
fig.update_layout(width=800, height=800/1.618, title_text="Transaction Size by Destionation Chain")
fig.update_layout(template="simple_white", showlegend=False)
fig.update_yaxes(title_text='Tx Size UST', row=1, col=1)
fig.update_yaxes(title_text='Tx Size UST', row=2, col=1)
fig.show()
# + [markdown] id="5YX2NvbRdv8F"
# # What's going on with Solana?
#
# We have seen above that the volume and patterns of UST sent to Solana is different to the three EVM chains (BSC, Ethereum, Polygon). The Solana data is shown in the chart below (excluding network ops wallet). There appears to be a cyclical component to the usage, possibly based on a weekday/weekend cycle.
# + colab={"base_uri": "https://localhost:8080/", "height": 511} id="024BkwINeEcW" outputId="93aee616-a974-4654-fc4a-14b0f196cb26"
#hide_input
#UST Sent via Wormhole expluding the network liquidity account
# df_p = %R tx_by_dest_chain %>% filter(bridge_chain == 'Solana')
fig = px.bar(df_p
, x = "date"
, y = "qty"
, labels=dict(date="Date", qty="UST Sent")
, title= "UST Sent to Solana via Wormhole"
, template="simple_white", width=800, height=800/1.618
)
fig.update_layout(legend=dict(
yanchor="top",
y=0.99,
xanchor="right",
x=0.99,
title_text=None
))
fig.update_yaxes(title_text='UST')
fig.update_xaxes(title_text=None)
fig.show()
# + [markdown] id="3Yc-1n7vgKiU"
# The chart below shows the average volume of UST sent to Solana from Terra by day of the week. The very large day on the 4th of November was excluded as an outlier, as were the network operations transactions. There seems to be an influence based on day of the week - Monday sees transaction volumes of nearly 2m UST, and Saturdays at 0.6m UST as the low point. At face value this doesn't quite align with a normal work week. The timestamps, however are in UTC. APAC markets are ahead of UTC - Korea, a country known to use the Terra network heavily via the Chai payments network, is UTC+9 hours. The chart below aligns with a timezone such as this, with activity on Sunday night UTC being Monday morning in Asia. We can draw two conclusions from this:
#
# - Bridging to Solana is being done for work (or at work) as more transactions occur on weekdays as opposed to weekends
# - It is likely that this activity is occurring in APAC markets due to the observed timezone offest
# + colab={"base_uri": "https://localhost:8080/", "height": 511} id="xoP7BINEgJ6o" outputId="4111360a-1f00-499c-d9d1-0b1073cf0729"
#hide_input
#UST Sent via Wormhole expluding the network liquidity account
# df_p = %R sol_by_day_of_week
fig = px.bar(df_p
, x = "weekday"
, y = "av_qty"
, labels=dict(date="Date", qty="UST Sent")
, title= "UST Sent to Solana via Wormhole"
, template="simple_white", width=800, height=800/1.618
)
fig.update_layout(legend=dict(
yanchor="top",
y=0.99,
xanchor="right",
x=0.99,
title_text=None
))
fig.update_yaxes(title_text='UST')
fig.update_xaxes(title_text=None)
fig.show()
# + [markdown] id="foFTl9M8Jgbt"
# # Conclusions
#
# The Wormhole Network is a new cross-chain bridge network, linking blockchains with different consensus mechanisms in a decentralised manner. The connection of Wormhole into the Terra network has unleashed another avenue for UST to take over the world. UST is the most commonly sent token sent via the Wormhole bridge from Terra, with typical volumes of 1.5m UST per day being sent. Most of the volume is going to Solana, with only sporadic transaction volume going to the three EVM chains - Ethereum, BSC and Polygon. Unique wallet counts have been growing steadily, at daily rates of 1.5-3%. There are around 2000 unique wallets so far which have sent UST via this bridge. Transaction sizes vary by destination, with some evidence that transaction fees may impact this size. Finally, we saw that UST sent to Solana appeared to show commercial use, potentially in an APAC timezone due to the timing of the transaction volume.
#
# All data was sourced from the curated on-chain data tables at [Flipside Crypto](https://flipsidecrypto.com/)
| _notebooks/2021-12-01-UST-into-the-Wormhole.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tf_gpu
# language: python
# name: tf_gpu
# ---
# # Introduction to neural networks
# ## Questions
# ### 2
# A xor B = A ^ ~B v (~A ^ B)
#
# Rysunek w zeszycie
# ### 3
# Perceptron jest zbieżny jedynie w przypadku gdy dane są liniowo separowalne.
# ### 4
# Użycie logistycznej funkcji aktywacji było podyktowane tym, że gradient tej funkcji jest niezerowy w całej swojej dziedzinie. Z tego powodu algorytm gradientu prostego nie zatrzymywał się.
# ### 5
# - RELU max(0, x)
# - Leaky RELU max(/alpha x, x), /alpha /in (0, 1)
# - sigmoid e^(-x) / (1 - e^(-x))
# +
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
n = 100
zeros = np.zeros(n)
x = np.linspace(-10, 10, n)
relu = np.maximum(zeros, x)
plt.plot(x, relu, 'b-')
alpha = 0.1
l_relu = np.maximum(alpha*x, x)
plt.plot(x, l_relu, 'r-')
sigm = 1 / (1 + np.exp(-x))
plt.plot(x, sigm, 'g-')
plt.xlim((-10, 10))
plt.ylim((-2, 2))
plt.show()
# -
# ### 6
# Y = XW + b
#
# X.dim = (-1, 10)
# W_u.dim = (10, 50) b_u.dim = (50)
# W_wy.dim = (50, 3) b_wy.dim = (3)
#
# Y.dim = (-1, 3)
#
# Y = X_wy * (X * W_u + b_u) + b_wy
# ### 7
# ham / spam - jeden neuron, aktywacja sigmoid(logistyczna), wartość neuronu określa prawdopodobieństwo należenia próbki do spamu
#
# MNIST - dziesięć neuronów, aktywacja softmax
#
# Housing - jeden neuron, aktywacja brak / liniowa
# ### 8
# Propagacja wsteczna - algorytm poprawiania wag sieci neuronowej wykorzystujący odwrotne różnioczkowanie automatyczne
#
# Odwrotne różniczkowanie automatyczne - algorytm automatycznie obliczający gradienty w sieci neuronowej
# ### 9
# - liczba warstw
# - liczba neuronów w warstwie
# - występowanie biasu
# - funkcja aktywacji
# - współczynnik uczenia
# - liczba próbek w batchu
#
# W przypadku przetrenowania należy spróbować zmniejszyć liczbę neuronów w warstwach lub liczbę warstw.
# ### 10
# MNIST
# +
import os
import numpy as np
import tensorflow as tf
from tensorflow.keras.datasets import mnist
from tensorflow.keras.layers import Dense, Flatten
from tensorflow.keras.models import Sequential
from tensorflow.keras.losses import *
from tensorflow.keras.optimizers import *
from tensorflow.keras.activations import *
from matplotlib import pyplot as plt
# %matplotlib inline
# -
(X_train_full, y_train_full), (X_test, y_test) = mnist.load_data()
plt.imshow(X_train_full[0], cmap='binary')
y_train_full.shape
X_train, y_train = X_train_full[:50_000] / 255., y_train_full[:50_000]
X_valid, y_valid = X_train_full[50_000:] / 255., y_train_full[50_000:]
tensorboard_path = os.path.join(os.curdir, 'my_logs')
def get_run_logdir():
import time
run_id = time.strftime('run_%Y_%m_%d-%H_%M_%S')
return os.path.join(tensorboard_path, run_id)
def build_model(hidden_layer_size):
model = Sequential()
model.add(Flatten(input_shape=(28,28)))
model.add(Dense(hidden_layer_size, activation='relu'))
model.add(Dense(hidden_layer_size, activation='relu'))
model.add(Dense(10, activation='softmax'))
return model
# #### Finding best learingn rate
# +
K = tf.keras.backend
class ExponensionalLR(tf.keras.callbacks.Callback):
def __init__(self, factor):
self.factor = factor
self.lr = []
self.loss = []
def on_batch_end(self, batch, logs):
self.lr.append(K.get_value(self.model.optimizer.lr))
self.loss.append(logs['loss'])
K.set_value(self.model.optimizer.lr, self.factor * self.model.optimizer.lr )
# -
model = build_model(256)
model.compile(loss='sparse_categorical_crossentropy',
optimizer=SGD(lr=1e-3),
metrics=['accuracy'])
model.summary()
lr_clb = ExponensionalDecayLR(1.005)
history = model.fit(X_train, y_train,
validation_data=(X_valid, y_valid),
callbacks=[lr_clb], epochs=1)
# +
import numpy
def smooth(x,window_len=11,window='hanning'):
if x.ndim != 1:
raise ValueError("smooth only accepts 1 dimension arrays.")
if x.size < window_len:
raise ValueError("Input vector needs to be bigger than window size.")
if window_len<3:
return x
if not window in ['flat', 'hanning', 'hamming', 'bartlett', 'blackman']:
raise ValueError("Window is on of 'flat', 'hanning', 'hamming', 'bartlett', 'blackman'")
s=numpy.r_[2*x[0]-x[window_len-1::-1],x,2*x[-1]-x[-1:-window_len:-1]]
if window == 'flat': #moving average
w=numpy.ones(window_len,'d')
else:
w=eval('numpy.'+window+'(window_len)')
y=numpy.convolve(w/w.sum(),s,mode='same')
return y[window_len:-window_len+1]
plt.plot(lr_clb.lr, smooth(np.array(lr_clb.loss), window_len=30))
plt.gca().set_xscale('log')
plt.ylim((0, 2.))
# +
tf.keras.backend.clear_session()
model = build_model(256)
model.compile(loss='sparse_categorical_crossentropy',
optimizer=SGD(lr=2e-1),
metrics=['accuracy'])
model.summary()
# -
tensorboard_cb = tf.keras.callbacks.TensorBoard(get_run_logdir())
earlystop_cb = tf.keras.callbacks.EarlyStopping(patience=10, restore_best_weights=True)
model.fit(X_train, y_train, validation_data=(X_valid, y_valid), epochs=50,
callbacks=[tensorboard_cb, earlystop_cb])
# %load_ext tensorboard
# %tensorboard --logdir=./my_logs --port=6006
| my_solutions/Chapter_10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.6 64-bit (''aligned-umap'': conda)'
# name: python3
# ---
import umap
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# Importing the data
p2 = pd.read_csv('Data/TTI_Pillar2/DistanceMatrixJaccardOriginPillar2.csv')
sgss = pd.read_csv('Data/TTI_SGSS/DistanceMatrixJaccardOriginSGSS.csv')
css = pd.read_csv('Data/CovidSymptomStudy/JaccardSymptomWide.csv')
cis = pd.read_csv('Data/CommunityInfectionSurvey/DistanceMatrixJaccard.csv')
# Name the first column as the symptom columnm.
p2 = p2.rename({p2.columns[0]: 'symptom'}, axis='columns')
sgss = sgss.rename({sgss.columns[0]: 'symptom'}, axis='columns')
css = css.rename({css.columns[0]: 'symptom'}, axis='columns')
cis = cis.rename({cis.columns[0]: 'symptom'}, axis='columns')
# Extract the jaccard distance matrices
p2_jaccard = p2.loc[:, p2.columns != 'symptom'].values
sgss_jaccard = sgss.loc[:, sgss.columns != 'symptom'].values
css_jaccard = css.loc[:, css.columns != 'symptom'].values
cis_jaccard = cis.loc[:, cis.columns != 'symptom'].values
# Extract the symptom names
p2_symptoms = p2.loc[:, p2.columns == 'symptom']
sgss_symptoms = sgss.loc[:, sgss.columns == 'symptom']
css_symptoms = css.loc[:, css.columns == 'symptom']
cis_symptoms = cis.loc[:, cis.columns == 'symptom']
# Each dataset stores the variables with slightly different names. We can join the the symptom_name_category lookup table to get a standardised naming convention for symptoms.
symptom_name_category_lookup = pd.read_csv('Data/Lookups/SymptomNameCategoryLookup.csv')
# We need to subset the lookup to get appropiate lookups for each dataset.
ctas_lookup = symptom_name_category_lookup[symptom_name_category_lookup.dataset == 'CTAS']
css_lookup = symptom_name_category_lookup[symptom_name_category_lookup.dataset == 'Zoe']
cis_lookup = symptom_name_category_lookup[symptom_name_category_lookup.dataset == 'ONS']
# Join the symptoms to create lookup, and retain only the useful data
p2_symptoms = pd.merge(left = p2_symptoms, right = ctas_lookup, left_on = 'symptom', right_on='symptom_name_raw')[['symptom', 'symptom_id', 'symptom_name_formatted', 'category']]
sgss_symptoms = pd.merge(left = sgss_symptoms, right = ctas_lookup, left_on = 'symptom', right_on='symptom_name_raw')[['symptom', 'symptom_id', 'symptom_name_formatted', 'category']]
css_symptoms = pd.merge(left = css_symptoms, right = css_lookup, left_on = 'symptom', right_on='symptom_name_raw')[['symptom', 'symptom_id', 'symptom_name_formatted', 'category']]
cis_symptoms = pd.merge(left = cis_symptoms, right = cis_lookup, left_on = 'symptom', right_on='symptom_name_raw')[['symptom', 'symptom_id', 'symptom_name_formatted', 'category']]
# Create a list containing a list of all the symptom_id's in each datset, and then work out which symptom id's are common across all datasets.
# +
symptom_ids = [
p2_symptoms.symptom_name_formatted.values,
sgss_symptoms.symptom_name_formatted.values,
css_symptoms.symptom_name_formatted.values,
cis_symptoms.symptom_name_formatted.values
]
shared_ids = symptom_ids[0]
for id_set in symptom_ids:
shared_ids = np.intersect1d(shared_ids, id_set)
shared_ids
# -
# Create a utlity function that produces the mapping between symptoms in different datasets.
def map_symptoms(vector_from, vector_to):
mapping = {}
for num_from, symptom_id_from in enumerate(vector_from.symptom_name_formatted.values):
if symptom_id_from in shared_ids:
for num_to, symptom_id_to in enumerate(vector_to.symptom_name_formatted.values):
if symptom_id_to == symptom_id_from:
mapping[num_from] = num_to
return mapping
# Specify a sequence of datasets, and then create a list of mappings between those datasets.
distance_matrix_list = [p2_jaccard, sgss_jaccard, css_jaccard, cis_jaccard]
relation_dict_list = [map_symptoms(p2_symptoms, sgss_symptoms), map_symptoms(sgss_symptoms, css_symptoms), map_symptoms(css_symptoms, cis_symptoms)]
# Perform the alignment
# +
aligned_mapper = umap.AlignedUMAP(
n_neighbors=[4]*4,
min_dist = 0.01,
n_components = 2,
alignment_window_size=4,
alignment_regularisation=1e-2,
learning_rate = 0.1,
random_state=42,
metric='precomputed')
# perform the alignment
aligned_mapper.fit(distance_matrix_list, relations = relation_dict_list)
# -
# utility function to find create some useable axis
def axis_bounds(embedding):
left, right = embedding.T[0].min(), embedding.T[0].max()
bottom, top = embedding.T[1].min(), embedding.T[1].max()
adj_h, adj_v = (right - left) * 0.1, (top - bottom) * 0.1
return [left - adj_h, right + adj_h, bottom - adj_v, top + adj_v]
# Quickly plot the output. We have more advanced plotting code in the R visualisation notebooks that plot points and labels using size and colour, etc.
fig, axs = plt.subplots(2, 2, figsize=(10, 10))
ax_bound = axis_bounds(np.vstack(aligned_mapper.embeddings_))
for i, ax in enumerate(axs.flatten()):
ax.scatter(*aligned_mapper.embeddings_[i].T)
ax.axis(ax_bound)
plt.tight_layout()
plt.show()
# saving outputs
p2_embedding = pd.DataFrame(aligned_mapper.embeddings_[0])
sgss_embedding = pd.DataFrame(aligned_mapper.embeddings_[1])
css_embedding = pd.DataFrame(aligned_mapper.embeddings_[2])
cis_embedding = pd.DataFrame(aligned_mapper.embeddings_[3])
p2_embedding[['symptom_name_raw', 'symptom_name_formatted', 'category']] = p2_symptoms[['symptom', 'symptom_name_formatted', 'category']]
sgss_embedding[['symptom_name_raw', 'symptom_name_formatted', 'category']] = sgss_symptoms[['symptom', 'symptom_name_formatted', 'category']]
css_embedding[['symptom_name_raw', 'symptom_name_formatted', 'category']] = css_symptoms[['symptom', 'symptom_name_formatted', 'category']]
cis_embedding[['symptom_name_raw', 'symptom_name_formatted', 'category']] = cis_symptoms[['symptom', 'symptom_name_formatted', 'category']]
p2_embedding.to_csv('Data/Alignments/AlignedUMAP/CoreSymptomAlignment/P2.csv')
sgss_embedding.to_csv('Data/Alignments/AlignedUMAP/CoreSymptomAlignment/SGSS.csv')
css_embedding.to_csv('Data/Alignments/AlignedUMAP/CoreSymptomAlignment/CSS.csv')
cis_embedding.to_csv('Data/Alignments/AlignedUMAP/CoreSymptomAlignment/CIS.csv')
| AlignedUMAP/FullDatasetAlignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !pip install scikit-learn
# +
import numpy as np
import pandas as pd
from pathlib import Path
from collections import Counter
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler, LabelEncoder, MinMaxScaler
from sklearn.metrics import accuracy_score
from sklearn.utils import resample
from sklearn.linear_model import LogisticRegression
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import confusion_matrix, classification_report, roc_curve, auc
import matplotlib.pyplot as plt
from sklearn.metrics import mean_absolute_error, make_scorer
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.impute import SimpleImputer
from sklearn.model_selection import KFold
from sklearn.model_selection import train_test_split
from sklearn.model_selection import cross_val_score
from sklearn.base import BaseEstimator, TransformerMixin
from sklearn.linear_model import BayesianRidge
from sklearn.preprocessing import OneHotEncoder
# -
train_df = pd.read_csv(Path('Resources/2019loans.csv'))
test_df = pd.read_csv(Path('Resources/2020Q1loans.csv'))
# Display for check
display(train_df)
display(test_df)
# Convert categorical data to numeric and separate target feature for training data
X_train = pd.get_dummies(train_df.drop(columns=['loan_status']))
y_train = train_df['loan_status']
# Convert categorical data to numeric and separate target feature for testing data
X_test = pd.get_dummies(test_df.drop(columns=['loan_status']))
y_test = test_df['loan_status']
# add missing dummy variables to testing set
for col in X_train.columns:
if col not in X_test.columns:
X_test[col] = 0
X_train
X_test
# Hypothesis - In doing research online, the random forest classifier model usually performs better than the logistic regression model, so I assume the random forest classifier model will out perform the logistic regression model in our example as well.
# +
# Train the Logistic Regression model on the unscaled data and print the model score
from sklearn.linear_model import LogisticRegression
classifier = LogisticRegression()
classifier
classifier.fit(X_train, y_train)
print(f"Training Data Score: {classifier.score(X_train, y_train)}")
print(f"Testing Data Score: {classifier.score(X_test, y_test)}")
# -
# Train a Random Forest Classifier model and print the model score
from sklearn.ensemble import RandomForestClassifier
clf = RandomForestClassifier(random_state=1, n_estimators=500).fit(X_train, y_train)
print(f"Training Data Score: {clf.score(X_train, y_train)}")
print(f"Testing Data Score: {clf.score(X_test, y_test)}")
# 2nd Hypothesis - Since scaling changes the range of the datasets, I assume scaling the data will cause it to be even more accurate. Still assuming the random forest classifier model will out perform the logisitic regression model.
# Scale the data
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler().fit(X_train)
X_train_scaled = scaler.transform(X_train)
X_test_scaled = scaler.transform(X_test)
# +
# Train the Logistic Regression model on the scaled data and print the model score
classifier.fit(X_train_scaled, y_train)
print(f"Training Data Score: {classifier.score(X_train_scaled, y_train)}")
print(f"Testing Data Score: {classifier.score(X_test_scaled, y_test)}")
# -
# Train a Random Forest Classifier model on the scaled data and print the model score
srfc = RandomForestClassifier(random_state=1, n_estimators=500).fit(X_train_scaled, y_train)
print(f"Training Data Score: {srfc.score(X_train_scaled, y_train)}")
print(f"Testing Data Score: {srfc.score(X_test_scaled, y_test)}")
# Conclusion - The random forest classifier model did outperform the logistic regression model on accuracy when the data was and was not scaled. The Logisitic regression model was more accurate when the data was scaled. But the random forest classifier model performed similarly regardless if the data was scaled or not.
| Credit Risk Evaluator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.5 64-bit
# name: python3
# ---
# + [markdown] id="K-N4ha5EG6ky" colab_type="text"
# # **Curso Python Para Análise de dados**
# + [markdown] id="P5yjJd4Se7a8" colab_type="text"
# # **Estrutura de dados**
# + [markdown] id="0VKL1NCNhUfG" colab_type="text"
# ## **Listas**
# + id="XHAFPHWmfFoB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="5dbb8e99-09ba-40db-f848-f2395ded9c73"
#Criando uma lista chamada animais
animais = [1,2,3]
animais
# + id="5v-w_ftdga0J" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="8ded31ab-a939-4f4d-caac-ef0a281bcfb6"
animais = ["cachorro", "gato", 12345, 6.5]
animais
# + id="bJkOFrGfgrKR" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="273b6c48-3245-47d8-b8d4-e37865e52b21"
#Imprimindo o primeiro elemento da lista
animais[0]
# + id="M-2mXu3mgvuS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="81236f47-7507-4682-ee71-96d05ad1eb41"
#Imprimindo o 4 elemento da lista
animais[3]
# + id="RDCW4hs_gxyT" colab_type="code" colab={}
#Substituindo o primeiro elemento da lista
animais[0] = "papagaio"
# + id="QJrP-830gzvh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e69ecaaa-d6a3-48ee-e08a-33b3c2417ad8"
animais
# + id="3QFbAI6Og0wP" colab_type="code" colab={}
#Removendo gato da lista
animais.remove("gato")
# + id="b8g1bs42g31p" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c4fa21bc-0375-4c4f-d645-1fa535ac3559"
animais
# + id="xfu4p_g3g7OS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="1a511e7d-4616-4fe1-eaab-154d1b8a84d1"
len(animais)
# + id="0HedcjoFg-5m" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="04ebec34-a75f-404a-b719-0ca710a24059"
"gato" in animais
# + id="VDZ6nZU2hDdf" colab_type="code" colab={}
lista = [500, 30, 300, 80, 10]
# + id="VjrYRfHJhGhL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="ad010fc6-b64f-4b11-ec1c-67ef22b93da0"
max(lista)
# + id="jkeXKWQPhJRr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="b8d73be6-aa7f-4129-bd57-9a8c294d5c77"
min(lista)
# + id="20T_hCwAhMsU" colab_type="code" colab={}
animais.append(["leão", "Cachorro"])
# + id="WgWWMNJXhQdB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="b2a0495e-b1ee-469d-bae5-e771c20e1efb"
animais
# + id="Cl-M69f5hSsD" colab_type="code" colab={}
animais.extend(["cobra", 6])
# + id="_scJEZbWMFBW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9ae58baf-1d1c-408d-9026-4a74e5a7f4a4"
animais
# + id="kNn0DiyxheHC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="b4a75f67-c2f5-4972-9ee1-17c6481c394a"
animais.count("leão")
# + id="VgoLBevzhinC" colab_type="code" colab={}
lista.sort()
# + id="mPJTKqXMhrDB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="aaa017a3-80a9-47ae-895a-63d6de385d18"
lista
# + [markdown] id="VHmSI4EcoX-h" colab_type="text"
# # **Tuplas**
# + id="Qrv07mzWocDT" colab_type="code" colab={}
#As tuplas usam parênteses como sintaxe
tp = ("Banana", "Maçã", 10, 50)
# + id="r7ZMMLn8pt7R" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d2ad4094-b8d6-489f-a605-36c4b569bd8b"
#Retornando o primeiro elemento
tp[0]
# + id="jKRnh-DbpzBj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 180} outputId="04b2babe-4507-4279-8cfc-64e62f3c20cf"
#Diferente das listas as tuplas são imutáveis, o que quer dizer que não podemos alterar os seus elementos
tp[0] = "Laranja"
# + id="a-KaNhePE0nb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="ddf3ebbd-355a-467e-d238-0c9601e07c61"
tp.count("Maçã")
# + id="ntXTi7MaFAX7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="595f3409-7948-4919-cff4-787d350423b7"
tp[0:2]
# + [markdown] id="Q0Ltz4kcFQps" colab_type="text"
# # **Dicionários**
# + id="43OFknPUFO68" colab_type="code" colab={}
#Para criar um dicionário utilizamos as {}
dc = {"Maçã":20, "Banana":10, "Laranja":15, "Uva":5} #Dicionários trabalham com o condeito chave e valor
# + id="dCH9PefxGBdZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="4017b350-db9b-4404-8f2b-1d41d9e983bf"
dc
# + id="iyGxfz72FxNa" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="4e85b169-8639-4402-ddcf-5a954e6acd9f"
#Acessando o valor de um dicionário através da chave
dc["Maçã"]
# + id="6WwkohdpF5Cq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="053f88f7-b499-47d5-c741-25f0a394e2c8"
#Atualizando o valor da Maçã
dc["Maçã"] = 25
dc
# + id="_50MV-XaGFae" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="53879989-4805-4017-f5a9-9294ba60d86a"
#Retornando todas as chaves do dicionário
dc.keys()
# + id="AHLXfikMGRS4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="1beaa637-f1dd-4122-80a5-5d317c993943"
#Retornando os valores do dicionário
dc.values()
# + id="co2yQ4IrGX8n" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f8f5dc72-2f3d-433b-c54b-cea91173e306"
#Verificando se já existe uma chave no dicionário e caso não exista inserir
dc.setdefault("Limão", 22)
# + id="M9_v4jAZGyz6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e8754510-9f3c-4560-c4c1-133cbc06cf9b"
dc
# + id="l2-4mE1LG0CC" colab_type="code" colab={}
| analise-de-dados-com-pandas/Notebooks/Aula2_Estrutura_de_dados.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from IPython.display import IFrame, YouTubeVideo, SVG, HTML
import pandas as pd
import numpy as np
print(pd.__version__)
print(np.__version__)
url = 'https://github.com/codeforamerica/ohana-api/blob/master/data/sample-csv/mail_addresses.csv'
df = pd.read_csv('https://github.com/prasertcbs/basic-dataset/raw/master/iris.csv')
df.species.value_counts()
df = pd.read_csv('https://github.com/codeforamerica/ohana-api/blob/master/data/sample-csv/contacts.csv')
df=pd.read_csv('https://spotifycharts.com/regional/global/daily/latest/download')
df.head()
df = pd.read_excel('https://github.com/prasertcbs/tutorial/raw/master/BornInGeneration.xlsx')
df.head()
| data-import.ipynb |