code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import matplotlib.pyplot as plt
# # Fundamental Building Blocks
#
# Based on the ideas presented by Cleveland, Ware, and the Gestalt Principles of Visual perception, we're going start talking about the types of charts we want to build and the types of customizations we would entertain, in order to take advantage of these principles (or not violate them).
#
# For <NAME>, this limits our basic elements to just four (4!):
#
# 1. Points
# 2. Lines
# 3. Bars
# 4. Boxes
#
# That's it at least in terms of *quantitative* encoding. For now, we will ignore boxes so that really leaves us points, lines and bars and some combinations.
#
# ## Points
#
# From a `matplotlib` point of view, these do not map exactly onto the functions available. For example, using points for *categorical* data is different than using points for *numerical* data. Additionally, you have to do things differently if you want your chart to be horizontal or vertical.
#
# Here's a categorical chart that uses points, *Dot Chart*. A Dot Chart is preferrable to a Bar Chart when you do not wish to include 0 in the y axis. The dotted lines are visual guidelines, especially in horizontal versions, they will go all the way through like gridlines. Both versions are provided here.
#
# Once you understand how `matplotlib` works, you will see that I have essentially used the regular `plot` function and made up fake x-axis data and gave it categorical labels.
# +
figure = plt.figure(figsize=(5, 3))
xs = [1, 2, 3, 4, 5]
data = [100, 87, 23, 47, 57]
axes = figure.add_subplot(1, 1, 1)
axes.plot(xs, data, "o", color="dimgray")
axes.vlines(xs, [0], data, linestyles='dotted', lw=2)
axes.set_xticks(xs)
axes.set_xlim((0, 6))
axes.set_ylim((0, 110))
axes.set_xticklabels(["A", "B", "C", "D", "E"])
axes.xaxis.grid(False)
# +
figure = plt.figure(figsize=(5, 3))
xs = [1, 2, 3, 4, 5]
data = [100, 87, 23, 47, 57]
axes = figure.add_subplot(1, 1, 1)
axes.plot(xs, data, "o", color="dimgray")
axes.vlines(xs, [0], [110], linestyles='dotted', lw=2)
axes.set_xticks(xs)
axes.set_xlim((0, 6))
axes.set_ylim((0, 110))
axes.set_xticklabels(["A", "B", "C", "D", "E"])
axes.xaxis.grid(False)
# -
# To use points with numerical data only, we use the same command `plot` and create a *Scatter Plot* or *XY-Plot*.
# +
figure = plt.figure(figsize=(5, 3))
xs = [50, 42, 39, 19, 27]
ys = [100, 87, 23, 47, 57]
axes = figure.add_subplot(1, 1, 1)
axes.plot(xs, ys, "o", color="dimgray")
axes.set_xlim((10, 55))
axes.set_ylim((10, 110))
# -
# Notice that there's no specific reason to start at 0 on the x or y scale.
#
# ## Lines
#
# Unless these points follow each other, we should not connect them with lines. If, however, these were readings of two variables that occurred in time, we might connect the points with lines. Note that this uses the Gestalt Principle of Connection...if there is no connection, don't use lines!
# +
figure = plt.figure(figsize=(5, 3))
xs = [50, 42, 39, 19, 27]
ys = [100, 87, 23, 47, 57]
axes = figure.add_subplot(1, 1, 1)
axes.plot(xs, ys, "o-", color="dimgray")
axes.set_xlim((10,60))
axes.set_ylim((10, 110))
axes.set_xticklabels(["A", "B", "C", "D", "E"])
# -
# A single value that is recorded over time is often shown with either just line, points or both. There can be various reasons for each. Just a line is often used where the implication that the numerical measurement is *continuous*. Just points are often used when there are breaks in the measurement (for example, no values for weekends). Points and lines can be used together when we want to emphasize both the continuity and the specific values, especially if we are plotting multiple lines.
#
#
# Note that for `matplotlib`'s purposes, time is categorical. You may need to create an index or use the actual millis on the x-axis and create tick labels for them.
# +
figure = plt.figure(figsize=(5, 3))
xs = [1, 2, 3, 4, 5]
ys = [100, 87, 23, 47, 57]
axes = figure.add_subplot(1, 1, 1)
axes.plot(xs, ys, "o-", color="dimgray")
axes.set_xlim((0, 6))
axes.set_ylim((10, 110))
axes.set_xticks(xs)
axes.set_xticklabels(["Jan", "Feb", "Mar", "Apr", "May"])
# -
# Notice how the Gestalt Principle of Closure describes how your perception projects the line out to June.
#
# ### Bars
#
# We've already done Bars. We will reproduce the chart from above.
# +
sales = [29, 23, 18, 17, 13]
width = 1/1.5
x = range( len( sales))
figure, axes = plt.subplots()
axes.bar(x, sales, width, color="dimgray", align="center")
axes.set_xticks([0, 1, 2, 3, 4])
axes.set_xticklabels(["A", "D", "B", "C", "E"])
axes.yaxis.grid( b=True, which="major")
axes.set_ylim((0, 30))
plt.show()
# -
# So those are the basic chart types. Which one you use, and what secondary changes you make (labels, color, "small multiples") depends on what you're trying to accomplish. Additionally, there will be times when you need to use specialized charts (for example, we'll see a "box-and-whiskers plot" soon) but you should always start with these.
# ## Multiple Data Series
#
# The Playfair Trade Balance chart is an example of a chart with multiple data series (they don't always have to be time related). The two series are exports and imports (and we could derive a 3rd which would be the trade balance). As another example, we can have sales and expenditures for our ten business units.
#
# We can also have data series for different things at different points in time. For example, monthly unemployment rates by state over the last 60 months.
#
# So far our charts have shown a single data series. How do we deal with multiple data series? This largely depends on the base chart (line, bar, dots) and exactly how many series we have...two or 50?
#
# Here's a typical line plot with three data series:
# +
figure = plt.figure(figsize=(5, 3))
xs = [1, 2, 3, 4, 5]
ys1 = [100, 87, 23, 47, 57]
ys2 = [55, 98, 91, 72, 89]
ys3 = [78, 93, 52, 80, 69]
axes = figure.add_subplot(1, 1, 1)
axes.plot(xs, ys1, "o-", color="dimgray", label="ys1")
axes.plot(xs, ys2, "o-", color="dimgray", label="ys2")
axes.plot(xs, ys3, "o-", color="dimgray", label="ys3")
axes.set_xlim((0, 6))
axes.set_ylim((10, 110))
axes.set_xticks(xs)
axes.set_xticklabels(["Jan", "Feb", "Mar", "Apr", "May"])
axes.legend()
plt.show()
plt.close()
# -
# Of course, we can't tell which one is which. We haven't talked about *color* yet but we can use different markers for the lines:
# +
figure = plt.figure(figsize=(5, 3))
xs = [1, 2, 3, 4, 5]
ys1 = [100, 87, 23, 47, 57]
ys2 = [55, 98, 91, 72, 89]
ys3 = [78, 93, 52, 80, 69]
axes = figure.add_subplot(1, 1, 1)
axes.plot(xs, ys1, "o-", color="dimgray", label="ys1")
axes.plot(xs, ys2, "x-", color="dimgray", label="ys2")
axes.plot(xs, ys3, "d-", color="dimgray", label="ys3")
axes.set_xlim((0, 6))
axes.set_ylim((10, 110))
axes.set_xticks(xs)
axes.set_xticklabels(["Jan", "Feb", "Mar", "Apr", "May"])
axes.legend()
plt.show()
plt.close()
# -
# This is still pretty difficult to decode. We have to keep looking at the legend and the points and then back. One of the easier solutions is small multiples...at least once you learn how to get multiple charts in a single plot area.
# +
figure = plt.figure(figsize=(20, 6))
xs = [1, 2, 3, 4, 5]
ys1 = [100, 87, 23, 47, 57]
ys2 = [55, 98, 91, 72, 89]
ys3 = [78, 93, 52, 80, 69]
axes = figure.add_subplot(1, 3, 1)
axes.plot(xs, ys1, "o-", color="dimgray", label="ys1")
axes.set_xlim((0, 6))
axes.set_ylim((10, 110))
axes.set_xticks(xs)
axes.set_xticklabels(["Jan", "Feb", "Mar", "Apr", "May"])
axes.set_xlabel("ys1")
axes = figure.add_subplot(1, 3, 2)
axes.plot(xs, ys2, "o-", color="dimgray", label="ys2")
axes.set_xlim((0, 6))
axes.set_ylim((10, 110))
axes.set_xticks(xs)
axes.set_xticklabels(["Jan", "Feb", "Mar", "Apr", "May"])
axes.set_xlabel("ys2")
axes = figure.add_subplot(1, 3, 3)
axes.plot(xs, ys3, "o-", color="dimgray", label="ys3")
axes.set_xlim((0, 6))
axes.set_ylim((10, 110))
axes.set_xticks(xs)
axes.set_xticklabels(["Jan", "Feb", "Mar", "Apr", "May"])
axes.set_xlabel("ys3")
plt.show()
plt.close()
# -
# Amazingly, it's fairly effortless to detect differences in the series. We can also arrange them vertically if we think that'll make for a better comparison:
# +
figure = plt.figure(figsize=(15, 8))
xs = [1, 2, 3, 4, 5]
ys1 = [100, 87, 23, 47, 57]
ys2 = [55, 98, 91, 72, 89]
ys3 = [78, 93, 52, 80, 69]
axes = figure.add_subplot(3, 1, 1)
axes.plot(xs, ys1, "o-", color="dimgray", label="ys1")
axes.set_xlim((0, 6))
axes.set_ylim((10, 110))
axes.set_xticks(xs)
axes.set_xticklabels(["Jan", "Feb", "Mar", "Apr", "May"])
axes.set_ylabel("ys1")
axes = figure.add_subplot(3, 1, 2)
axes.plot(xs, ys2, "o-", color="dimgray", label="ys2")
axes.set_xlim((0, 6))
axes.set_ylim((10, 110))
axes.set_xticks(xs)
axes.set_xticklabels(["Jan", "Feb", "Mar", "Apr", "May"])
axes.set_ylabel("ys2")
axes = figure.add_subplot(3, 1, 3)
axes.plot(xs, ys3, "o-", color="dimgray", label="ys3")
axes.set_xlim((0, 6))
axes.set_ylim((10, 110))
axes.set_xticks(xs)
axes.set_xticklabels(["Jan", "Feb", "Mar", "Apr", "May"])
axes.set_ylabel("ys3")
plt.show()
plt.close()
# -
# This works for bar charts as well although if you only have two series for a bar chart, there is an alternative.
| fundamentals_2018.9/visualization/blocks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # kNN Clustering - DOHMH New York City Restaurant Inspection Results
#
# Find groups of different business names that might be alternative representations of the same venue. This is an example for the **kNN clustering** supported by **openclean**.
# +
# Open the downloaded dataset to extract the relevant columns and records.
import os
from openclean.pipeline import stream
df = stream(os.path.join('data', '43nn-pn8j.tsv.gz'))
# -
# ## Extract Relevant Records
#
# Get set of distinct business names from *DBA* column.
# +
# Get distinct set of street names. By computing the distinct set of
# street names first we avoid computing keys for each distinct street
# name multiple times.
dba = df.select('DBA').distinct()
print('{} distinct bisiness names (for {} total values)'.format(len(dba), sum(dba.values())))
# +
# Cluster business names using kNN clusterer (with the default n-gram setting)
# using the Levenshtein distance as the similarity measure.
# Remove clusters that contain less than ten distinct values (for display
# purposes).
from openclean.cluster.knn import knn_clusters
from openclean.function.similarity.base import SimilarityConstraint
from openclean.function.similarity.text import LevenshteinDistance
from openclean.function.value.threshold import GreaterThan
# Minimum cluster size. Use ten as default (to limit
# the number of clusters that are printed in the next cell).
minsize = 5
clusters = knn_clusters(
values=dba,
sim=SimilarityConstraint(func=LevenshteinDistance(), pred=GreaterThan(0.9)),
minsize=minsize
)
print('{} clusters of size {} or greater'.format(len(clusters), minsize))
# +
# For each cluster print cluster values, their frequency counts,
# and the suggested common value for the cluster.
def print_cluster(cnumber, cluster):
print('Cluster {} (of size {})\n'.format(cnumber, len(cluster)))
for val, count in cluster.items():
print('{} ({})'.format(val, count))
print('\nSuggested value: {}\n\n'.format(cluster.suggestion()))
# Sort clusters by decreasing number of distinct values.
clusters.sort(key=lambda c: len(c), reverse=True)
for i, cluster in enumerate(clusters):
print_cluster(i + 1, cluster)
# +
# Perform normalization of business names first to get an
# initial set of clusters using key collision clustering.
# Then run kNN clustering on the collision keys.
from collections import Counter
from openclean.cluster.knn import knn_collision_clusters
clusters = knn_collision_clusters(
values=dba,
sim=SimilarityConstraint(func=LevenshteinDistance(), pred=GreaterThan(0.9)),
minsize=minsize
)
print('{} clusters of size {} or greater'.format(len(clusters), minsize))
# +
# Print resulting clusters.
clusters.sort(key=lambda c: len(c), reverse=True)
for i, cluster in enumerate(clusters):
print_cluster(i + 1, cluster)
| examples/notebooks/nyc-restaurant-inspections/NYC Restaurant Inspections - kNN Clusters for Business Name.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# In this part of the exercise, you will implement regularized logistic regression to predict whether microchips from a fabrication plant passes quality assurance (QA). During QA, each microchip goes through various tests to ensure it is functioning correctly.
# Suppose you are the product manager of the factory and you have the test results for some microchips on two different tests. From these two tests, you would like to determine whether the microchips should be accepted or rejected. To help you make the decision, you have a dataset of test results on past microchips, from which you can build a logistic regression model.
#import the libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
#Check out the data file
# !head ../data/ex2data2.txt
data = pd.read_csv('../data/ex2data2.txt', sep=',', header=None, names=['Test1', 'Test2', 'Pass'])
data.head()
data.describe()
# Scale of Test values are similar. I will not apply Standardization. But if means and standart deviations were really far apart, it may have made sense to standardize the data for the optimization purposes. We can see it on a box plot.
plt.figure()
plt.boxplot([data['Test1'], data['Test2']]);
# Let's change the visual style:
plt.style.use('ggplot')
# Finally, let's visualize the data at hand:
# +
fig, ax = plt.subplots(figsize=(12, 8))
ax.scatter(data[data["Pass"] == 0]["Test1"], data[data["Pass"] == 0]["Test2"],
marker='x', label='Failed')
ax.scatter(data[data["Pass"] == 1]["Test1"], data[data["Pass"] == 1]["Test2"],
label='Passed')
ax.legend(frameon = True, fontsize="large", facecolor = "White", framealpha = 0.7)
ax.set_xlabel('Test1')
ax.set_ylabel('Test2');
# -
# A linear seperator will not cut it here, and as a result we are going to implement 5th degree polynomial regression here. It is an overkill but we will use the regularization to get rid of unnecessary elements.
#
# There is a MapFeature module in the homework written in Octave code. Using this code and the code from @jdwittenauer repo, I will transform the Dataset at hand to have the following transformation on our dataframe:
#
# $mapFeature(x) = [x_1, x_2, x_1^2, x_1 x_2,x_2^2,..., x_1x_2^5,x_2^6]$
# +
#let's first keep original data set as I will modify the dataframe
data_or = data.copy()
#There is the MapFeature.m
#I think there was a mistake in his implementation, I corrected it
degree = 7
for i in range(1, degree):
for j in range(0, i+1):
data["T" + str(i-j) + str(j)] = np.power(data["Test1"], i-j) * np.power(data["Test2"], j)
# -
data.drop(['Test1','Test2'], axis=1, inplace=True)
data.insert(1, "Ones", 1)
data.head()
# "As a result of this mapping, our vector of two features (the scores on two QA tests) has been transformed into a 28-dimensional vector. "
#
# We have 28 columns + 1 column for results as well. Finally define the numpy input and output arrays, initial theta value and check the dimensions for them
# +
X = data.iloc[:,1:]
y = data.iloc[:,0]
X = np.asarray(X.values)
y = np.asarray(y.values)
theta = np.zeros(X.shape[1])
#check the dimensions
X.shape, y.shape, theta.shape
# -
# **Regularization**
#
# Regularization means we will add the following item to our old cost function:
#
# $J(\theta) = J_{\text{old}}(\theta) + \frac{\lambda}{2m} \sum_{j=1}^{k}\theta_j^2$
# +
def sigmoid(z):
return 1.0 / (1.0 + np.exp(-z))
def h(theta, X):
"""
Hypothesis Function where
X is an n x k dimensional array of explanatory variables
theta is a array with k elements of multipliers for linear function
Result will be one dimensional vector of n variables
"""
return sigmoid(np.dot(X, theta))
# -
#Now the cost function
def cost_function(theta, Lambda, X, y):
"""
This is a cost function that returns the cost of theta given X and y
X is an n x k dimensional array of explanatory variables
y is a one dimensional array with n elements of explained variables
theta is a vector with k elements of multipliers for linear function
"""
item1 = - y.flatten() * np.log(h(theta, X))
item2 = -(1 - y.flatten()) * np.log(1 - h(theta, X))
item3 = Lambda/(2*X.shape[0]) * np.sum(np.power(theta, 2)[1:])
return np.sum(item1 + item2) / (X.shape[0]) + item3
# $\frac{\partial J(\theta)}{\partial \theta_{j}} = \frac{1}{n} \sum\limits_{i=1}^n (h_{\theta}(x^{i}) - y^i)x_j^i$
def gradient(theta, Lambda, X, y):
"""
This function will take in theta value and calculate the gradient
descent values.
X is an n x k matrix of explanatory variables
y is a n sized array of explained variables
theta is a vector with k elements of multipliers for linear function
"""
errors = h(theta, X) - y
#errors will be calculated more than once, so let's do it once and store it
correction2 = (Lambda/X.shape[0]) * theta
correction2[0] = 0.0
correction = np.sum(errors.reshape((X.shape[0], 1)) * X, axis=0) * (1.0 / X.shape[0])
return correction + correction2
# The cost for Lambda=1 and theta=0 is:
theta=np.zeros(X.shape[1])
cost_function(theta, 1, X, y)
# **Finding the Parameter**
# Finally let's apply our optimization method and find the optimal theta values
# +
import scipy.optimize as opt
Lambda = 1.0
theta = np.zeros(X.shape[1])
result = opt.minimize(fun=cost_function, method='TNC',
jac= gradient, x0=theta, args=(Lambda, X,y), options={'maxiter':400})
result
# -
# Interesting tidbit: if you don't use the gradient function, it takes 750 loops. But if you use it it takes 32. Cost functions are very close.
# Let's check the accuracy rate of our prediction function.
# +
theta_opt = result['x']
def prediction_function(theta, X):
return h(theta, X) >= 0.5
total_corrects = np.sum((y.flatten() == prediction_function(theta_opt, X)))
total_dpoints = X.shape[0]
accuracy_rate = total_corrects/total_dpoints
accuracy_rate
# -
# **Plotting the decision boundary**
#
# We will plot the decision boundary. In order to that I will create a 3d grid with values of test1 and test2 data, and the corresponding $h_{\theta}(X)$ (Acutally we will only find $\theta X$ since $h$ is one to one function). Using this data we can use the plot-contour level functions to find where $\theta X = 0$ since this is where the decision boundary is.
#
# In order to this we need to write the MapFeature function explicitly. It should take in arrays or values and give us polynomial elements of them.
def MapFeature(x1, x2):
"""
This takes in two n elements vector arrays, then builds a
n x 28 dimensional array of features
"""
#flatten the vectors in case:
x1 = x1.flatten()
x2 = x2.flatten()
num_ele = len(x1)
degrees = 6
res_ar = np.ones( (len(x1), 1) )
for i in range(1, degrees+1):
for j in range(0, i+1):
res1 = np.power(x1, i-j)
res2 = np.power(x2, j)
res3 = np.multiply(res1, res2).reshape( (num_ele, 1) )
res_ar = np.hstack(( res_ar, res3 ))
return res_ar
# Let's check if our function work properly:
#The following code checks if there are any non-equal elements.
np.count_nonzero(MapFeature(X[:,1],X[:,2]) != X)
# Now let's define a function to draw counters. I have utilized some of the code from the homework and then also @kaleko's website. In order to repeat the image from the homework, I had to transpose zvals like Prof Ng did in his codes. To be honest, I don't know why we had to do this.
#
# Our function will also show the accuracy rate before it shows the graph it drew. Quite handy.
def Draw_Contour(X, y, Lambda):
#First we need to find optimal Theta
theta_initial = np.zeros(X.shape[1])
result = opt.minimize(fun=cost_function, method='TNC', jac= gradient,
x0=theta_initial, args=(Lambda, X,y), options={'maxiter':400, 'disp':False})
theta = result['x']
#Next define the grids
xvals = np.linspace(-1,1.5,50)
yvals = np.linspace(-1,1.5,50)
zvals = np.zeros((len(xvals),len(yvals)))
for i, xv in enumerate(xvals):
for j, yv in enumerate(yvals):
features = MapFeature(np.array(xv), np.array(yv))
zvals[i, j] = np.dot(features, theta)
zvals = zvals.transpose()
#To be honest I don't know why we transpose but this is the way it is in
#the code provided by Professor Ng.
#Now draw the graph. I reused some code from before.
fig, ax = plt.subplots(figsize=(12, 8))
ax.scatter(X[(y == 0), 1], X[(y == 0), 2], marker='x', label='Failed')
ax.scatter(X[(y == 1), 1], X[(y == 1), 2], label='Passed')
mycontour = ax.contour( xvals, yvals, zvals, [0], label="boundary")
myfmt = { 0:'Lambda = %d'%Lambda}
ax.clabel(mycontour, inline=1, fontsize=15, fmt=myfmt)
ax.legend(frameon = True, fontsize="large", facecolor = "White", framealpha = 0.7)
ax.set_xlabel('Test1')
ax.set_ylabel('Test2')
#Our function will also show the accuracy rate as it draws the graph
total_corrects = np.sum((y.flatten() == prediction_function(theta, X)))
total_dpoints = X.shape[0]
accuracy_rate = total_corrects/total_dpoints
return accuracy_rate
# Let's try our function for different Lambda values, for Lambda = 0, 1, 10
Draw_Contour(X, y, 0.00001)
Draw_Contour(X, y, 1.0)
Draw_Contour(X, y, 10)
| Ex 2 - Logistic Regression/Logistic Regression with Regularization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Graphes : arbres de recherche binaire et quadtrees
# Les graphes sont une notion très importante en informatique. Ils nous seront très utiles pour représenter des réseaux (de transport, de distribution d'énergie etc.) mais aussi pour construire des "espaces de recherche".
# ## Qu'est-ce qu'un graphe ?
# L'histoire des graphes commence en 1735 dans la ville de Königsberg (qui est aujourd'hui l'enclave russe de Kaliningrad).
#
# 
#
# Le mathématicien <NAME> se pose la question de savoir s'il existe ou non une promenade dans les rues de Königsberg permettant, à partir d'un point de départ au choix, de passer une et une seule fois par chaque pont (il y en a 7), et de revenir à son point de départ.
# La réponse est non mais la modélisation mathématique consiste à représenter le problème des sept ponts de Königsberg par un graphe :
#
# 
# Un graphe est constitué de *noeuds* (ici A, B, C, D et E) et *d'arêtes* (I, II, III, IV, V, VI et VII). Il est par exemple possible de passer du noeud A au noeud B puis de revenir à A. C'est ce qu'on appele un *cycle*.
# Un *arbre* est un graphe sans cycle.
#
# 
# Comment coder un graphe en python ? On peut utiliser un dictionnaire :
# dictionnaire konigsberg = {...}
# On peut bien évidemment utiliser des classes d'objets.
#
# ## Noeud
class Node:
"""Définition d'un noeud"""
# Constructeur de noeud
def __init__(self, label = "NaN", content = 0):
self.label = label
self.content = content
# Méthode utilisée par print()
def __str__(self):
t = "Node(" + self.label + ", " + str(self.content) + ")"
return t
a = Node(label = "A", content = 6)
type(a)
print(a)
# ## Arbre de recherche binaire
#
# Commençons par modéliser un arbre de recherche binaire (sera aussi vu en cours d'algorithmique) :
#
# 
class ArbreBin:
"""Arbre binaire"""
# Constructeur d'arbre binaire
def __init__(self, arbre_g, racine, arbre_d):
self.arbre_g = arbre_g
self.racine = racine
self.arbre_d = arbre_d
# A faire !
def __str__(self):
pass
af = ArbreBin(None, Node('F', 8), None)
ag = ArbreBin(None, Node('G', 11), None)
ad = ArbreBin(af, Node('D', 9), ag)
ae = ArbreBin(None, Node('E', 24), None)
ac = ArbreBin(ad, Node('C', 12), ae)
ab = ArbreBin(None, Node('B', 4), None)
aa = ArbreBin(ab, Node('A', 6), ac)
type(aa)
# +
# print(aa) doit donner :
# ArbreBin(ArbreBin(None, Node(B, 4), None), Node(A, 6), ArbreBin(ArbreBin(ArbreBin(None, Node(F, 8), None), Node(D, 9), ArbreBin(None, Node(G, 11), None)), Node(C, 12), ArbreBin(None, Node(E, 24), None)))
# -
class ArbreBin:
"""Arbre binaire"""
# Constructeur d'arbre binaire
def __init__(self, arbre_g, racine, arbre_d):
self.arbre_g = arbre_g
self.racine = racine
self.arbre_d = arbre_d
def __str__(self):
if (self.arbre_g is None):
t_g = "None"
else:
t_g = self.arbre_g.__str__()
if (self.arbre_d is None):
t_d = "None"
else:
t_d = self.arbre_d.__str__()
return "ArbreBin(" + t_g + ", " + self.racine.__str__() + ", " + t_d + ")"
# Ajoute content à l'arbre
#def add(self, node):
# Si strictement inférieur, ajout à gauche
#if (node.content < self.racine.content):
#if self.arbre_g is None:
#left = ...
#else:
#left = ...
#return ArbreBin(...)
# Sinon (supérieur ou égal), ajout à droite
#else:
#if self.arbre_d is None:
#right = ArbreBin(...)
# else:
#right = ...
#return ArbreBin(...)
aa = ArbreBin(None, Node('A', 6), None)
#new_aa = aa.add(Node('B', 7))
# +
#print(new_aa)
# -
class ArbreBin:
"""Arbre binaire"""
# Constructeur d'arbre binaire
def __init__(self, arbre_g, racine, arbre_d):
self.arbre_g = arbre_g
self.racine = racine
self.arbre_d = arbre_d
def __str__(self):
if (self.arbre_g is None):
t_g = "None"
else:
t_g = self.arbre_g.__str__()
if (self.arbre_d is None):
t_d = "None"
else:
t_d = self.arbre_d.__str__()
return "ArbreBin(" + t_g + ", " + self.racine.__str__() + ", " + t_d + ")"
# Ajoute content à l'arbre
def add(self, node):
# Si strictement inférieur, ajout à gauche
if (node.content < self.racine.content):
if self.arbre_g is None:
left = ArbreBin(None, node, None)
else:
left = self.arbre_g.add(node)
return ArbreBin(left, self.racine, self.arbre_d)
# Sinon (supérieur ou égal), ajout à droite
else:
if self.arbre_d is None:
right = ArbreBin(None, node, None)
else:
right = self.arbre_d.add(node)
return ArbreBin(self.arbre_g, self.racine, right)
# Teste si l'arbre contient content
#def contains(self, content):
#if (content == self.racine.content):
#return ...
#elif (content < self.racine.content):
#return ...
#else:
#return ...
aa = ArbreBin(None, Node('A', 6), None)
#new_aa = aa.add(Node('D', 9)).add(Node('E', 24)).add(Node('F', 8)).add(Node('G', 11))
# +
#new_aa.contains(8)
# +
#new_aa.contains(21)
# -
# ## Ok mais à quoi servent les arbres binaires ?
#
# Reprenons l'algorithme de recherche séquentielle vu au TP3.
def rechercher(tableau, valeur):
isFound = False
j = 0
while (j < len(tableau)) and not isFound:
x = tableau[j]
j += 1
if (x == valeur):
isFound = True
return isFound
rechercher([1, 4, 67, 8, 5], 8)
rechercher([1, 4, 67, 8, 5], 0)
# Exercice : ajouter à la classe ArbreBin une méthode permettant de créer un arbre binaire à partir d'un tableau (liste).
class ArbreBin:
"""Arbre binaire"""
# Constructeur d'arbre binaire
def __init__(self, arbre_g, racine, arbre_d):
self.arbre_g = arbre_g
self.racine = racine
self.arbre_d = arbre_d
def __str__(self):
if (self.arbre_g is None):
t_g = "None"
else:
t_g = self.arbre_g.__str__()
if (self.arbre_d is None):
t_d = "None"
else:
t_d = self.arbre_d.__str__()
return "ArbreBin(" + t_g + ", " + self.racine.__str__() + ", " + t_d + ")"
# Ajoute content à l'arbre
def add(self, node):
# Si strictement inférieur, ajout à gauche
if (node.content < self.racine.content):
if self.arbre_g is None:
left = ArbreBin(None, node, None)
else:
left = self.arbre_g.add(node)
return ArbreBin(left, self.racine, self.arbre_d)
# Sinon (supérieur ou égal), ajout à droite
else:
if self.arbre_d is None:
right = ArbreBin(None, node, None)
else:
right = self.arbre_d.add(node)
return ArbreBin(self.arbre_g, self.racine, right)
# Ajoute une liste d'entiers à un arbre
# A faire !
def addL(self, tableau):
pass
# Teste si l'arbre contient content
def contains(self, content):
if (content == self.racine.content):
return True
elif (content < self.racine.content):
return (not (self.arbre_g is None)) and self.arbre_g.contains(content)
else:
return (not (self.arbre_d is None)) and self.arbre_d.contains(content)
aa = ArbreBin(None, Node('Root'), None)
#new_aa = aa.addL([0, 15, 27])
# +
#print(new_aa)
# -
print(aa)
# Maintenant voyons quel est l'algorithme le plus rapide : rechercher ou contains ?
# +
# A faire commme la comparaison des algorithmes de tri
# -
# On voit donc que la recherche dans un arbre bianire est beaucoup plus efficace.
# ## Ok mais quel rapport avec la géomatique ?
#
# Des idées similaires aux arbres binaires (quadtrees, R-trees etc.) sont utilisées dans de nombreux domaines (bases de données, synthèse/compression d'images, simulations, jeux vidéo et géomatique) pour "indexer" les données et augmenter considérablement les performances.
#
# 
#
# Supposons que l'on veuille calculer quels sont les objets proches de B. On va donc devoir faire autant de calcul de distance qu'il y a d'objets même s'il est évident que certains objets ne sont pas "proches". Et si l'on veut connaître tous les objets se trouvant à moins d'une certaine distance les uns des autres (par exemple, où se trouve toutes les antennes relais situées à moins de 100 m d'une école ?), le nombre de calculs augmente encore plus rapidement : 100 pour 10 objets, 10000 pour 100 objets, 1000000 pour 1000 objets etc.
#
# 
#
# Les quadtrees sont des arbres quaternaires, chaque noeud à 4 descendants. Ils permettent d'indexer les objets du plan. Lorsque le nombre d'objets du plan considéré dépasse un certain seuil arbitraire (1 dans notre exemple), le plan est découpé en 4 sous-zones. Et ainsi de suite récursivement. Pour connaître les points "proches" de B, on ne considérera que les points placés dans le même noeud que B, ou ceux appartenant à des noeuds "proches" (ce qui se calcule facilement car on connaît les coordonnées des points aux extrémités des zones).
# (le code ci-dessous n'est pas de moi, je l'adapte de [là](https://jrtechs.net/data-science/implementing-a-quadtree-in-python))
class Point():
def __init__(self, x, y):
self.x = x
self.y = y
class Node():
def __init__(self, x0, y0, w, h, points):
self.x0 = x0
self.y0 = y0
self.width = w
self.height = h
self.points = points
self.children = []
def get_width(self):
return self.width
def get_height(self):
return self.height
def get_points(self):
return self.points
# +
def recursive_subdivide(node, k):
if len(node.points) > k:
w_ = float(node.width/2)
h_ = float(node.height/2)
p = contains(node.x0, node.y0, w_, h_, node.points)
x1 = Node(node.x0, node.y0, w_, h_, p)
recursive_subdivide(x1, k)
p = contains(node.x0, node.y0+h_, w_, h_, node.points)
x2 = Node(node.x0, node.y0+h_, w_, h_, p)
recursive_subdivide(x2, k)
p = contains(node.x0+w_, node.y0, w_, h_, node.points)
x3 = Node(node.x0 + w_, node.y0, w_, h_, p)
recursive_subdivide(x3, k)
p = contains(node.x0+w_, node.y0+h_, w_, h_, node.points)
x4 = Node(node.x0+w_, node.y0+h_, w_, h_, p)
recursive_subdivide(x4, k)
node.children = [x1, x2, x3, x4]
def contains(x, y, w, h, points):
pts = []
for point in points:
if (point.x >= x) and (point.x <= x + w) and (point.y >= y) and (point.y <= y + h):
pts.append(point)
return pts
def find_children(node):
if not node.children:
return [node]
else:
children = []
for child in node.children:
children += (find_children(child))
return children
# +
import random
import matplotlib.pyplot as plt # plotting libraries
import matplotlib.patches as patches
class QTree():
def __init__(self, k, n):
self.threshold = k
self.points = [Point(random.uniform(0, 10), random.uniform(0, 10)) for x in range(n)]
self.root = Node(0, 0, 10, 10, self.points)
def add_point(self, x, y):
self.points.append(Point(x, y))
def get_points(self):
return self.points
def subdivide(self):
recursive_subdivide(self.root, self.threshold)
def graph(self):
fig = plt.figure(figsize=(12, 8))
plt.title("Quadtree")
c = find_children(self.root)
print("Number of segments: %d" %len(c))
areas = set()
for el in c:
areas.add(el.width*el.height)
print("Minimum segment area: %.3f units" %min(areas))
for n in c:
plt.gcf().gca().add_patch(patches.Rectangle((n.x0, n.y0), n.width, n.height, fill=False))
x = [point.x for point in self.points]
y = [point.y for point in self.points]
plt.plot(x, y, 'ro') # plots the points as red dots
plt.show()
# -
test = QTree(3, 100)
test.subdivide()
test.graph()
# + active=""
#
| programmation_python/TP6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.6.2
# language: julia
# name: julia-0.6
# ---
# # Types
# The **type** of a variable tells us the "shape" of the variable, i.e. how to treat / interpret the data stored in the block of memory associated with that variable.
#
# Although it is possible to do much in Julia without thinking or worrying about types, they are always just under the surface, and the true power of the language becomes available through their use.
x = 3
typeof(x)
# For basic ("primitive") types, we can see the raw bits that are associated to a variable:
bits(x) # bitstring in Julia 0.7
y = 3.0
bits(y)
# The internal representations are different, corresponding to the different types.
# We can treat the storage as being of a different type:
z = reinterpret(Int, y)
h = hex(z)
z1 = parse(Int, h, 16) # base 16
bits(z1)
# # Defining our own types
# We have previously seen some examples of how to define a new type in Julia.
#
# A type definition can be thought of as template for a kind (type) of box, that contains certain **fields** containing data.
#
# One of the simplest examples is a "volume" type, representing the volume of some physical or mathematical object:
struct Vol
value
end
Vol
# This defines a new type, called `Vol`, containing one field, called `value`.
#
# It does not yet create an object that has that type. That is done by calling a **constructor** -- a special function with the same name as the type:
V = Vol(3)
V
# `V` is a Julia variable that is of type `Vol`:
typeof(V)
# Its "shape" is thus that of a box containing itself a single variable, which we can access as
V.value
# Since we defined the type `Vol` as `struct`, we cannot modify the contents of the object once it has been created:
V.value = 10
# We could instead make a mutable object:
mutable struct Vol1
value
end
V = Vol1(3)
V.value
V.value = 10
V
# We can change how our objects look by defining a new method of the `show` function:
# +
import Base: show
show(io::IO, V::Vol1) = print(io, "Volume with value ", V.value)
# -
V = Vol1(3)
# We can define e.g. the sum of two volumes:
# +
import Base: +
+(V1::Vol1, V2::Vol1) = Vol1(V1.value + V2.value)
# -
V + V
# But the following does not work, since we haven't defined `*` yet on our type:
2V
# **Exercise**: Define `*` of two `Vol`s and of a `Vol` and a number.
# ## Type annotations
# There is a problem with our definition:
Vol1("hello")
# It doesn't make sense to have a string as a volume. So we should **restrict** which kinds of `value` are allowed, by specifying ("annotating") the type of `value` in the type definition:
struct Vol2
value::Float64
end
Vol2(3.1)
Vol2("hello")
# # Parameterizing a type
# In different contexts, we may want integer volumes, or rational volumes, rather than `Vol`s which contain a floating-point number, e.g. for a 3D printer that makes everything out of cubes of the same size.
#
# We could define the following sequence of different types.
# +
type Vol_Int
value::Int
end
type Vol_Float
value::Float64
end
type Vol_Rational
value::Rational{Int64}
end
# -
Vol_Int(3)
Vol_Int(3.1)
Vol_Float(3.1)
# But clearly this is the wrong way to do it, since we're repeating ourselves, leading to inefficiency and buggy code. (https://en.wikipedia.org/wiki/Don't_repeat_yourself).
#
# Can't Julia itself automatically generate all of these different types?
# ## Specifying type parameters
# What we would like to do is tell Julia that the **type** itself (here, `Int`, `Float64` or `Rational{Int64}`)
# is a special kind of **parameter** that we will specify.
#
# To do so, we use curly braces (`{`, `}`) to specify such **type parameter** `T`:
struct Vol3{T}
value
end
# We can now create objects of type `Vol3` with any type `T`:
V = Vol3{Float64}(3.1)
typeof(V)
V2 = Vol3{Int64}(4)
typeof(V2)
# We see that the types of `V1` and `V2` are *different* (but related), and we have achieved what
# we wanted.
# The type `Vol3` is called a **parametric type**, with **type parameter** `T`. Parameteric types may have several type parameters, as we have already seen with `Array`s:
a = [3, 4, 5]
typeof(a)
# The type parameters here are `Int64`, which is itself a type, and the number `1`.
# ## Improving the solution
# The problem with this solution is the following, which echos what happened at the start of the notebook:
V = Vol3{Int64}(3.1)
typeof(V.value)
# The type `Float64` of the field `V.value` is distinct from the type parameter `Int64` we specified.
# So we have not yet actually captured the pattern of `Vol2`,
# which restricted the `value` field to be of the desired type.
#
# We solve this be specifying the field to **also be of type `T`**, with the **same `T`**:
struct Vol4{T}
value::T
end
# For example,
V = Vol4{Int64}(3)
# If necessary and possible, the argument to the constructor will be converted to the parametric type `T` that we specify:
V = Vol4{Int64}(3.0)
typeof(V.value)
# Now when we try to do
Vol4{Int64}(3.1)
# Julia throws an error, namely an `InexactError()`.
# This means that we are trying to "force" the number 3.1 into a "smaller" type `Int64`, i.e. one in which it can't be represented.
# However, now we seem to be repeating ourselves again: We know that `3.1` is of type `Float64`, and in fact Julia knows this too; so it seems redundant to have to specify it. Can't Julia just work it out? Indeed, it can!:
Vol4(3.1)
# Here, Julia has **inferred** the type `T` from the "inside out". That is, it performed pattern matching to realise that `value::T` is **matched** if `T` is chosen to be `Float64`, and then propagated this same value of `T` **upwards** to the type parameter.
# ## More fields
# **Exercise**: Define a `Point` type that represents a point in 2D, with two fields. What are the options for this type, mirroring the types `Vol1` through `Vol4`?
# ## Summary
# With parametric types, we have the following possibilities:
#
# 1. Julia converts (if possible) to the header type
#
# 2. Julia infers the header type from the inside (through the argument)
#
# # Constructors
# When we define a type, Julia also defines the **constructor functions** that we have been using above. These are functions with exactly the same name as the type.
#
# They can be discovered using `methods`:
struct Vol1
value
end
methods(Vol1)
# We see that Julia provides two default constructors. [Note that the output has changed in Julia 0.7.]
# For parametric types, it is a bit more complicated:
methods(Vol4)
methods(Vol4{Float64})
# ## Outer constructors
# Julia allows us to provide our own constructor functions.
# E.g.
struct Vol1
value
end
struct Vol2
value::Float64
end
Vol2(3)
Vol2("3.1")
# Here, we have tried to provide a numeric string, which is not allowed, since the string is not a number. We can add a constructor to allow this:
Vol2(s::String) = Vol2(parse(Float64, s))
Vol2("3.1")
Vol2("hello")
# We have added a new constructor outside the type definition, so it is called an **outer constructor**.
# ## Constructors that impose a restriction: **inner constructors**
# Now consider the following:
Vol4(3)
V = Vol4(3.5)
V.value
Vol4(-1)
# Oops! A volume cannot be negative, but the constructors so far have no restrictions, and so allow us to make a negative volume. To prevent this, Julia allows us to provide our own constructor, in which any restrictions are enforced.
#
# These constructors are written **within the type definition itself**, and hence are called **inner constructors**.
#
# [In Julia, these are the **only methods** that may be defined inside the type definition. Unlike in object-oriented languages, methods **do not belong to types** in Julia; rather, they exist outside any particular type, and (multiple) dispatch is used instead.]
#
# For example:
struct Vol5
value::Float64
function Vol5(V)
if V < 0
throw(ArgumentError("Volumes cannot be negative"))
end
new(V)
end
end
Vol5(3)
Vol5(-34)
# If we define an inner constructor, then Julia no longer defines the standard constructors; this is why defining an inner constructor gives us exclusive control over how our objects are created.
# Note that we use a special function `new` to actually create the object by filling in the values of the fields.
# If we use an immutable object (defined using `struct`), there is no way of changing the value of the field stored inside the object, so the invariant that `value` must be positive can never be violated.
# # Parametric functions
# Since we now have the ability to make parametric types, we may wish to define parametric functions on those types. E.g.
struct Length{T}
length::T
end
l = Length(10)
function square_area(l)
return l^2
end
# Suppose that we wish to round the area of squares with floating-point side length:
function square_area(l::T) where {T <: AbstractFloat} # method for types T that are subtypes of AbstractFloat
return ceil(Int, l^2)
end
square_area(11.1)
# # Inner constructors for parametric types
struct Vol6{T<:Real}
value::T
function Vol6{T}(V) where {T<:Real} # where specifies that T is a parameter of the parametric function Vol6
if V < 0
throw(ArgumentError("Negative"))
end
return new{T}(V)
end
end
# Here, we have used the syntax for parametric functions to specify a parametric inner constructor
Vol6(3)
Vol6{Float64}(3.1)
# We see that so far, we must explicitly specify the parametric type.
methods(Vol6)
# We can again make Julia infer the type for us:
Vol6(x::T) where {T<:Real} = Vol6{T}(x)
Vol6(3.1)
Vol6(3)
methods(Vol6)
x = 3//4 # rational number
Vol6(x)
# # Fixed-size objects are efficient
struct Vec{T}
x::T
y::T
end
# How efficient is this?
# +
import Base: +
+(f::Vec{T}, g::Vec{T}) where {T} = Vec(f.x + g.x, f.y + g.y)
# -
using BenchmarkTools
@btime +(Vec(1.0, 2.0), Vec(1.0, 2.0))
@btime [1.0, 2.0] + [1.0, 2.0]
# Using fixed-size objects is much more efficient (50 times more efficient)! They are defined in the `StaticArrays.jl` package.
| 08. User-defined types and parametric types.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 모델의 성능 개선하기
# * IBM sample datasets
# https://www.kaggle.com/blastchar/telco-customer-churn
#
# * Demographic info:
# * Gender, SeniorCitizen, Partner, Dependents
# * Services subscribed:
# * PhoneService, MultipleLine, InternetService, OnlineSecurity, OnlineBackup, DeviceProtection, TechSupport, StreamingTV, StreamingMovies
# * Customer account info:
# * CustomerID, Contract, PaperlessBilling, PaymentMethod, MonthlyCharges, TotalCharges, Tenure
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
# +
from IPython.display import set_matplotlib_formats
set_matplotlib_formats('retina')
# -
# ## 데이터 로드하기
# * 전처리된 모델을 로드하기
df = pd.read_csv("data/telco_feature.csv")
df.shape
# customerID 를 인덱스로 설정하기
df = df.set_index("customerID")
df.head()
# ## 전처리
# 결측치를 채워주는 방법도 있지만 일단 제거하도록 합니다.
df = df.dropna()
df["Churn"].value_counts()
# ## 학습, 예측 데이터셋 나누기
# ### 학습, 예측에 사용할 컬럼
# 피처로 사용할 컬럼 지정하기
feature_names = df.columns.tolist()
feature_names.remove("Churn")
feature_names
# ### 정답값이자 예측해야 될 값
# +
# label_name 이라는 변수에 예측할 컬럼의 이름을 담습니다.
label_name = "Churn"
label_name
# -
# ### 문제(feature)와 답안(label)을 나누기
#
# * X, y를 만들어 줍니다.
# * X는 feature, 독립변수, 예) 시험의 문제
# * y는 label, 종속변수, 예) 시험의 정답
# X, y를 만들어 줍니다.
X = df.drop(label_name, axis=1)
y = df[label_name]
# ### 학습, 예측 데이터셋 만들기
# * X_train : 학습 세트 만들기, 행렬, 판다스의 데이터프레임, 2차원 리스트(배열) 구조, 예) 시험의 기출문제
# * y_train : 정답 값을 만들기, 벡터, 판다스의 시리즈, 1차원 리스트(배열) 구조, 예) 기출문제의 정답
# * X_test : 예측에 사용할 데이터세트를 만듭니다. 예) 실전 시험 문제
# * y_test : 예측의 정답값 예) 실전 시험 문제의 정답
# +
# train_test_split 으로 데이터셋 나누기
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42)
# -
X_train.shape, X_test.shape, y_train.shape, y_test.shape
X_train.head(3)
X_test.head(3)
y_train.head(2)
# ## 머신러닝 모델로 예측하기
# +
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier, GradientBoostingClassifier
estimators = [DecisionTreeClassifier(random_state=42),
RandomForestClassifier(random_state=42),
GradientBoostingClassifier(random_state=42)
]
estimators
# -
results = []
for estimator in estimators:
result = []
result.append(estimator.__class__.__name__)
results.append(result)
results
# +
from sklearn.model_selection import RandomizedSearchCV
results = []
for estimator in estimators:
result = []
max_depth = np.random.randint(2, 20, 10)
max_features = np.random.uniform(0.3, 1.0, 10)
param_distributions = {"max_depth": max_depth,
"max_features": max_features}
if estimator.__class__.__name__ != 'DecisionTreeClassifier':
param_distributions["n_estimators"] = np.random.randint(100, 200, 10)
clf = RandomizedSearchCV(estimator,
param_distributions,
n_iter=5,
scoring="accuracy",
n_jobs=-1,
cv=5,
verbose=2
)
clf.fit(X_train, y_train)
result.append(estimator.__class__.__name__)
result.append(clf.best_params_)
result.append(clf.best_score_)
result.append(clf.score(X_test, y_test))
result.append(clf.cv_results_)
results.append(result)
# -
df_result = pd.DataFrame(results,
columns=["estimator", "best_params", "train_score", "test_score", "cv_result"])
df_result
# ## 학습하기
model = DecisionTreeClassifier(max_depth=6, max_features=0.9, random_state=42)
model
# 학습하기
model.fit(X_train, y_train)
# 예측하기
y_predict = model.predict(X_test)
y_predict
# ## 모델 평가하기
# +
# 피처의 중요도를 추출하기
importances = pd.DataFrame({"importances" : model.feature_importances_,
"feature_names" :feature_names})
importances = importances.sort_values("importances", ascending=False)
# -
# 피처의 중요도 시각화 하기
plt.figure(figsize=(10, 20))
sns.barplot(data=importances, x="importances", y="feature_names")
# ### 점수 측정하기
# #### Accuracy
# +
from sklearn.metrics import accuracy_score
accuracy_score(y_test, y_predict)
# -
(y_test == y_predict).mean()
# #### F1 score
# * precision 과 recall의 조화평균
# * [정밀도와 재현율 - 위키백과, 우리 모두의 백과사전](https://ko.wikipedia.org/wiki/%EC%A0%95%EB%B0%80%EB%8F%84%EC%99%80_%EC%9E%AC%ED%98%84%EC%9C%A8)
# +
# plot_confusion_matrix 를 그립니다.
from sklearn.metrics import plot_confusion_matrix
plot_confusion_matrix(model, X_train, y_train)
# +
from sklearn.metrics import classification_report
report = classification_report(y_test, y_predict)
print(report)
# -
| telco-customer-churn/telco_prediction_04_random_search.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .ts
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Typescript 3.3
// language: typescript
// name: typescript
// ---
// # Asynchronous output
//
// Typescript and Node.JS make heavy use of asynchronous execution. ITypescript lets you exercise these asynchronous capabilities, both:
//
// - by updating `stdout` and `stderr` asynchronously, or
// - by updating the ITypescript output asynchronously.
//
// **Note**: This function came from IJavascript, and we port these as static functions.
// ## Updating `stdout` and `stderr` asynchronously
//
// Both streams `stdout` and `stderr` can be written asynchronously. Any text written to these streams will be forwarded back to the latest request from the frontend:
// +
class Counter {
private _n: number = 1;
private _intervalObject: any;
start(n: number, millisec: number){
this._n = n;
this._intervalObject = setInterval(() => {
console.log(this._n--);
if(this._n < 0){
clearInterval(this._intervalObject);
console.warn("Done!");
}
}, millisec);
}
}
let c = new Counter();
c.start(5, 1000);
// -
// ## Updating the ITypescript output asynchronously
//
// ITypescript offers two global definitions to help produce an output asynchronously:
// * IJavascript style: `$$.async()` and `$$.sendResult(result: any)` or `$$.done()`.
// * ITypescript style: `%async on` and `$TS.retrieve(result?: any)`.
//
// When you call `$$.async()`, the ITypescript kernel is instructed not to produce an output. Instead, an output can be produced by calling `$$.sendResult()` or `$TS.retrieve()`.
//
// **Note**: `%async on` should be present at the top of the cell.
// +
// IJavascript style
$$.async();
console.log("Hello!");
setTimeout(
() => {
$$.sendResult("And good bye!");
},
1000
);
// -
// It is also possible to produce a graphical output asynchronously, the same way it is done synchronously, with the difference that `$$done$$()` has to be called to instruct the ITypescript kernel that the output is ready:
// +
// %async on
// ITypescript style
console.log("Hello!");
setTimeout(
() => {
$TS.svg("<svg><circle cx='30px' cy='30px' r='20px'/></svg>");
$TS.done();
},
1000
);
// -
| doc/async.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from jupyter_plotly_dash import JupyterDash
import dash_core_components as dcc
import dash_html_components as html
import dash
from dash.dependencies import Input, Output
from pymongo import MongoClient
import urllib.parse
from bson.json_util import dumps
#TODO: import for their CRUD module
from animalsCRUD import AnimalShelter
# this is a juypter dash application
app = JupyterDash('ModuleFive')
# the application interfaces are declared here
# this application has two input boxes, a submit button, a horizontal line and div for output
app.layout = html.Div(
[
dcc.Input(
id="input_user".format("text"),
type="text",
placeholder="input type {}".format("text")),
dcc.Input(
id="input_passwd".format("password"),
type="password",
placeholder="input type {}".format("password")),
html.Button('Execute', id='submit-val', n_clicks=0),
html.Hr(),
html.Div(id="query-out"),
#TODO: insert unique identifier code here
html.Header("Header"),
html.Tfoot("CS 340.")
]
)
# this is area to define application responses or callback routines
# this one callback will take the entered text and if the submit button is clicked then call the
# mongo database with the find_one query and return the result to the output div
@app.callback(
Output("query-out", "children"),
[Input("input_user".format("text"), "value"),
Input("input_passwd".format("password"),"value"),
Input('submit-val', 'n_clicks')],
[dash.dependencies.State('input_passwd', 'value')]
)
def cb_render(userValue,passValue,n_clicks,buttonValue):
if n_clicks > 0:
###########################
# Data Manipulation / Model
# use CRUD module to access MongoDB
##########################
username = "aacuser"
password = "password"
#TODO: Instantiate CRUD object with above authentication username and password values
testMod5 = AnimalShelter('aacuser', 'password')
# note that MongoDB returns BSON, the pyMongo JSON utility function dumps is used to convert to text
#TODO: Return example query results
tester = testMod5.locate({"animal_type":"Dog","name":"Lucy"})
for y in tester:
return y
app
# -
| Undergrad/CS-340/Project One/python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:new]
# language: python
# name: conda-env-new-py
# ---
# As <NAME> failed to deliver all the legacy *Spitzer* MIPS maps, I have had to collect certian maps. For the Bootes field, these have come from Mattia Vaccari's [Datafusion folders](http://mattiavaccari.net/df/m24/). The files available are:
# ls
from astropy.io import fits
# Lets check format of one of the previous maps
elais_n1=fits.open('../../dmu26/data/ELAIS_N1/MIPS/wp4_elais-n1_mips24_map_v1.0.fits.gz')
elais_n1
elais_n1[0].header
elais_n1[1].header
elais_n1[2].header
elais_n1[3].header
# Ok, so there is a primary header, an image map, an uncertianty map and a coverage map. Lets create the FITS file for Bootes.
hdr = fits.Header()
hdr['PRODUCT'] = 'WP4_Bootes_OFFICIAL_MIPS_24'
hdr['VERSION'] = 'V1.0 '
hdr['FIELD'] = 'Bootes'
hdr['SFILTERS']= 'MIPS_24 '
hdr['DESC'] = 'image, uncertainty and & coverage'
hdr['RIGHTS'] = 'Public '
primary_hdu = fits.PrimaryHDU(header=hdr)
hd1=fits.open('./data/bootes_24um_v2_mosaic.fits.gz')
hd2=fits.open('./data/bootes_24um_v2_mosaic_unc.fits.gz')
hd3=fits.open('./data/bootes_24um_v2_mosaic_cov.fits.gz')
hdul=fits.HDUList([primary_hdu,fits.ImageHDU(hd1[0].data,
header=hd1[0].header),fits.ImageHDU(hd2[0].data, header=hd2[0].header),
fits.ImageHDU(hd3[0].data, header=hd3[0].header)])
hdul.writeto('wp4_bootes_mips24_map_v1.0.fits.gz')
| dmu17/dmu17_Bootes/create_HELP_format_MIPS_map.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Box plots: An investigation into box plots and their use, with example plots.
# ### Notebook structure
# The notebook is split up into the following sections which are based on the project problem statement requirements.
#
# * Section One - Getting started with all necessary python libraries/packages
# * Section Two - Summarise the history of the box plot and situations in which it is used, explaining relevant terminology. (I have combined two of the project statement requirements in section two as I felt they fitted well together).
# * Section Three - Demonstrate the use of the box plot, using data of your choosing.
# * Section Four - Compare the box plot to alternatives.
# * Section Five - References and conclusion
# ### Section one - getting started: importing packages
# The following Python packages are imported for use in this notebook:
# * Mathplotlib.pyplot.py [Mathplotlib.org](https://matplotlib.org). Matplotlib is a Python plotting library and Pyplot is a matplotlib module which provides a MATLAB type interface.
# * NumPy [NumPy](http://www.numpy.org/). NumPy is a Python package for mathematical computing.
# * Seaborn [Seaborn](https://seaborn.pydata.org). Seaborn is a Python package used for plotting data.
# * Pandas [Pandas](https://pandas.pydata.org). Pandas is a Python package for use with data frames.
#
#command below ensures plots display correctly in the notebook
# %matplotlib inline
#below imports all necessary python packages for this notebook
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
import pandas as pd
# ### Section Two - Summarise the history of the box plot and uses. Explain the structure of a boxplot and related terminology.
# Boxplots were first used in 1969 by mathematician [<NAME>](https://en.wikipedia.org/wiki/John_Tukey), below.
# <NAME> was an American mathematician and the founding chairman of the Princeton statistics department.
# Amongst his many contributions to mathematics and statistics one which most people might recognise was the coining of the term "bit" for Binary digits (this, whilst researching at Bell laboratories).
#
# Whilst first using them himself in 1970, Tukey first introduced boxplots in his 1977 book, "Exploratory Data Analysis."
# They are a simple way of visually displaying data distribution based on the five number summary: minimum, first quartile, median, third quartile, and maximum.
#
# In the intervening years they have become one of the most used graphs for statistical display and there are also some variations of the original plot that allow for better display of the distributions within the boxplot than the original. Some of the more common variations are the violin plot and variable width boxplot, both which give a feel for data density. The variations stay true to the Tukey's original boxplot five number summary. It is probably fair to say that it is the advancement of computing and the capabilities of display of data using computers that has led to the many more sophisticated variations of the original box plot which could be created by hand. A paper from 2011,*40 Years of Boxplots by <NAME> and <NAME>*$^{1}$ outlines many of these variations.
#
# Tukey pioneered exploratory data analysis (EDA), believing too great an emphasis was put on confirmatory data analysis (CDA). During my research on Tukey I came across an interesting paper by *<NAME>* on *Exploratory Data Analysis*$^{2}$in which he refers to Tukey's likening EDA to detective work and also highlights the complementary relationship between EDA and CDA.
#
# 
#
# The boxplot is a primary tool in EDA, displaying in a graphical manner the variability in the data. They are easily constructed and during research I came across an online lesson from *University of Texas-Houston*$^{3}$covering boxplots which I would highly recommend a look at.
#
# Boxplots are used to show the distribution of data and show the following five points (which are summary statistics of a dataset.
#
# * data minimum - this is the minimum data point excluding outliers.
# * first quartile this is the 25$^{th}$ Percentile, 25% of data points are below this point.
# * median - this is the 50th percentile, 50% of data points will be above and 50% of points below this point.
# * third quartile - this is the 75$^{th}$ percentile, 75% of data points are below this point.
# * data maximum - this is the maximum data point excluding outliers.
#
# Below I have represented the main features of a box plot.
#
# 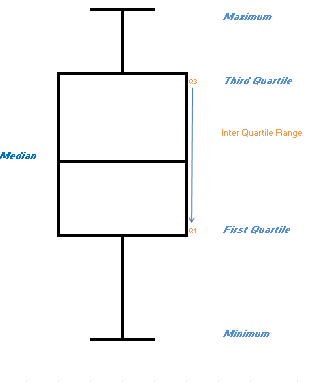
#
#
# It consists of a rectangular box with whiskers extending from each end to the minimum and maximum data points. The rectangle itself goes from the first quartile to the third quartile (called the interquartile range) and a line within the box shows the median value of the data. It allows a simple but highly visually effective method of displaying and comparing data distributions.
#
# For example, if the median is not in the middle of the box the distribution is skewed. Where the median is closer to the bottom, the distribution is skewed right. Where the median is closer to the top, the distribution is skewed left. Therefore, the shape of the boxplot (whisker length and median position) can be used to predict the likely data distribution.
#
# To create a boxplot data is split into four equal sized groups.
# The lowest 25% goes from the minimum to the first quartile **Q1** - **the end of the whisker to the bottom of the box**.
#
# The **box itself shows the next 50% of data** with the **median** (middle value) marked as a line within the box.
#
# The top 25% of data goes from the third quartile **Q3** to the maximum - **the top of the box to the end of the top whisker.**
#
# The range of data from the Third Quartile to First Quartile (Q3-Q1) is called the **interquartile range** (IQR).
#
# Any data points that are more than 1.5 times the box length (IQR) above Q3 or below Q1 may be outliers. Data points more than 3 times the IQR above Q3 or below Q1 are extreme values. Outliers and extreme values are plotted as separate data points rather than as part of the whiskers.
#
# Some possible uses of boxplots would be
# * to compare data from duplicate production lines (to ensure all machines/tools working correctly)
# * to look at change over time where external factors might influence data (as I will do in my examples below).
# * A simple example might be to compare standardised test results of different schools.
# * For large datasets it provides a good first look and would assist in the identification of data that needs cleaning (as I will demonstrate in my second scenario below).
# ### Section Three - Demonstrate the use of the box plot using data of your choosing.
#
# For this I decided to use a few simple examples based on CAO points.
#
# Each year Irish Students undergo a state examination called the Leaving Certificate (LC), see a Wikipedia page about the examination here, [Leaving Cert](https://en.wikipedia.org/wiki/Leaving_Certificate_(Ireland)). For these LC students entry to third level courses in Ireland is via a central application system [CAO](http://www.cao.ie/).
#
# For the purpose of allocating college places Leaving cert results are converted to **applicant points** and Third Level institutions review applicant points & available course places and determine the minimum entry points for the course. Course are then offered to eligible applicants based on their points, in descending order until the course places are all allocated. If an applicant does not take the offered place subsequent rounds of offers are made until the course is filled, with course point reductions being made if necessary. The **course points** will therefore be the total points obtained by the last applicant allocated a course place.
#
# This method means that course popularity and number of available places can result in points changing from year to year, therefore it is to a large extent an example of supply and demand. It also means that the spread of student points can vary from year to year.
#
# In the first simulated example below, example one, a course had a points requirement in 2016 of 365 with a max points of 530, in 2017 a points requirement of 397 with max 495. Intake is unchanged with 39 places allocated in each year.
#
# Below I have plotted the points and also used NumPy to calculate the five boxplot points and the interquartile ranges.
#
# I then used Seaborn to generate the same plot and generated some summary statistics; this is to demonstrate how easily a smart looking plot and summary statistics can be generated by even the most novice user. Something that I feel is extremely relevant in current workplace environments where working with IT systems and data is no longer the sole preserve of the IT department!
#
# In the final part of this section I will bring in a second dataset which represents the variation of points for a different programme, program B across six years. This is intended to show the effect of outliers and unclean data and how a quick visual review with a box plot can highlight data cleansing requirements.
#
# #### Box Plot Example One
## Sample Data One - below I am reading in a file of mocked up CAO points for a course A
df = pd.read_csv("https://raw.githubusercontent.com/Hudsonsue/BOXPLOTS/master/mock%20up%20points.csv", header =0)
#df
# +
## using mathplotlib I am going to create a boxplot of the dataset
boxplot = df.boxplot(column=['201600','201700'], return_type='axes')
plt.title ('Program A, Points Comparison 2016 & 2017')
plt.ylabel('points')
plt.xlabel('term code')
## using numpy I am going to calculate the five points of the box plot and the interquartile range
## minimum
min_pts = np.min(df, axis=0)
print("Min 201600", min_pts[0], " --- Min 201700",min_pts[1])
## Q1
Q1 =np.percentile(df, 25, axis =0)
print ("Q1 201600", Q1[0], "--- Q1 201700", Q1[1])
## median
med =np.median(df,axis =0)
print ("Median 201600", med[0], "--- Median 201700", med[1])
## Q3
Q3 =np.percentile(df, 75, axis=0)
print ("Q3 201600", Q3[0], "--- Q3 201700", Q3[1])
## Maximum
max_pts = np.max(df, axis=0)
print("Max 201600", max_pts[0], " --- Max 201700",max_pts[1])
##Inter Quartile Range
IR = Q3-Q1
print()
print("Interquartile range 201600", (IR[0]))
print("Interquartile range 201700", (IR[1]))
# +
## below I am replicating the above using seaborn.
## I just want to highlight how easily a nice looking plot can be generated.
df1 = pd.DataFrame(data = df, columns = ['201600','201700'])
sns.set(style="darkgrid")
sns.boxplot(x="variable",y = "value", hue = "variable",data=pd.melt(df), palette="Set3").set_title("CAO Points Course A, Comparison of 2016 & 2017")
plt.legend()
plt.ylabel('points')
plt.xlabel('term code')
# below are descriptive statistics for the data. The min, 25%, 50%, 75% and max are the five points of the boxplot
df.describe()
# -
# The example above is demonstrating in 2016 the two common scenarios in CAO points, at the higher end you have a student who has an excess of points but has stuck with the course they want; at the lower end the scenario where the demand was not as high as expected and to fill and make the course viable it is decided to drop points and pull in the last few students in a later round of offers.
#
# In both years the data is skewed right with the part of the box below the median smaller than that above, this tells me that the points of students in the lower part of the box are less varied (e.g. closer together)
#
# When plotted as boxplots it is apparent that whilst the overall spread of points and the max were greater in 2016; the 2017 median was higher as were entry points (bottom point of lower whisker).
#
# The box plots provide a nice visualisation of the course intake across the two years.
# #### Box Plot Example Two
# +
## I am going to read in a second csv to represent the change in points for a programme across a longer timeframe
## of six years.
df3= pd.read_csv("https://raw.githubusercontent.com/Hudsonsue/BOXPLOTS/master/boxplotdata.csv", header = 0)
df3.head(2)
# -
## seaborn box plot of dataset for Course B across six years
## outliers /incorrect data causing plot to be impossible to review
df4 = pd.DataFrame(data = df3, columns = [ '2012','2013', '2014', '2015','2016','2017'] )
sns.boxplot(x="variable", y="value", data= pd.melt(df4), palette="bright").set_title("CAO Points Course B 2012-2017")
plt.ylabel('points')
plt.xlabel('term code')
plt.show()
# Having pulled in the data there are some possible outliers showing:
#
# In 2012 I can see one in 2012 sitting far below the rest of the data, upon investigation it appears to be a typo as it is 41 when all other points in 2012 are three digits and in the range 400-565.
#
# In 2014 I can see some outliers but upon investigation I decide they are valid data points as they are within the range of possible LC points. So I would not remove them.
#
# In 2017 I can see some outliers, upon looking at the plot I can see they are sitting around 900 and 1000. As I am familiar with CAO coding I immediately recognise these as data that should not be in the dataset as they are codes used by CAO for mature and previous defer students and not part of the dataset I wish to use, that of student LC points.
# I hope the above demonstrates the use of boxplots to perform some quick data analysis and to avoid any in-depth analysis of inaccurate datasets. In the next part of the notebook I will demonstrate how the data would look once cleaned and re plotted.
# +
## I am going to read in the cleaned up version of the dataset for programme B, points for a programme
##across a longer timeframe of six years.
df4= pd.read_csv("https://raw.githubusercontent.com/Hudsonsue/BOXPLOTS/master/boxplotdatacleaned.csv", header = 0)
df4.head(2)
# +
## seaborn box plot of dataset for Course B across six years following removal of incorrect data points.
## I have also plotted a strip plot to show how it could be used in conjunction with the box plot to show
##the distribution within the box
#distplot of the cleaned dataset
df5 = pd.DataFrame(data = df4, columns = [ '201200','201300', '201400', '201500','201600','201700'] )
sns.boxplot(x="variable", y="value", palette = "bright", data= pd.melt(df4)).set_title("CAO Points Course B 2012-2017")
plt.ylabel('points')
plt.xlabel('term code')
plt.show()
# strip plot of the cleaned dataset
df6 = pd.DataFrame(data = df5, columns = [ '201200','201300', '201400', '201500','201600','201700']) #making a dataframe for the plot
sns.stripplot(x="variable",y = "value",data=pd.melt(df6), palette="bright").set_title("CAO Points Course B 2012-2017")
plt.ylabel('points') # title of the y axis
plt.xlabel('term code') # title of the x axis
plt.show()
# -
# Once the dataset is cleaned the plots are much easier to review. I added in the strip plot as it complements the boxplot. The rising points for the course from 2012 through to 2016 can be seen from the lowest whisker extremity.
# For example, in 2012 the data has an outlier and the data distribution has a large spread of data. The data is skewed right with the points of the students in the lower section of the box closer to each other than in the upper section.
#
# In 2014 the data is much closer (with two outliers identified) but skewed left with points of the students in the top section of the box closer together.
#
# The most symmetrical of all year plots is 2016 with the points reasonably evenly spread throughout the range.
# In 2017 the data is again skewed left with the points of students in the top section of the box closer together.
# ### Section Four - Compare the box plot to alternatives.
#
# As boxplots are distribution plots the first alternative to spring to mind is the histogram, below I have plotted example one above as seaborn [distribution plots](https://seaborn.pydata.org/generated/seaborn.distplot.html), both superimposed and side by side.
# Whilst it is possible to interpret the plots it is not as visually easy as the box plots where I can compare all six years on the same graph, side by side.
#
# However one limitation of the boxplot versus histograms is that while a dataset might look symmetric (when the median is in the centre of the box), it can’t tell you the shape of the symmetry the way a histogram can. this is outlined in an online article by <NAME> $^{4}$ So you could for example have two more dense regions of data within the box either side of the median but that won't be apparent in a box plot, whereas in a histogram it would be represented by longer bars.
#
# I also investigated the use of a [strip plot](https://seaborn.pydata.org/generated/seaborn.stripplot.html), it shows the distribution of data points and it would be useful in conjunction with the box plots to avoid the above mentioned limitation.
#
# As an aside whilst doing these comparisons it highlighted one major advantage of the boxplot which is that whilst it is lovely in colour the colour is not a necessity which is always an advantage when producing draft documents in the workplace and trying to interpret grayscale!
# +
# plot histograms and strip plots of the data in Example One above to demonstrate alternatives to a boxplot
# plots are using Seaborn with added titles & labels and using colours red and blue for 201600 and 201700 respectively.
sns.set(style="darkgrid")#setting the style for all plots
## the first plot is two histograms superimposed
sns.distplot(df["201600"], label="201600", color="red").set_title("CAO Points Course A")
plt.legend()
sns.distplot(df["201700"], label="201700",color="blue").set_title("CAO Points Course A")
plt.legend()
plt.show()
# the second plot is two side by side histograms
f, axes = plt.subplots(1,2, figsize=(8, 4), sharex=True)
sns.distplot( df["201600"] , color="red", ax=axes[0]).set_title("CAO Points Course A")
sns.distplot( df["201700"] , color="blue", ax=axes[1]).set_title("CAO Points Course A")
plt.show()
# the third plot is a strip plot of the dataset
df1 = pd.DataFrame(data = df, columns = ['201600','201700']) #making a dataframe for the plot
sns.stripplot(x="variable",y = "value", hue = "variable",data=pd.melt(df), palette="Set1").set_title("CAO Points Course A")
plt.legend()#legend
plt.ylabel('points') # title of the y axis
plt.xlabel('term code') # title of the x axis
plt.show()
# -
# ### Section Five- references and conclusion
# **Specific references**
#
# $^{1}$ [40 Years of Boxplots](http://vita.had.co.nz/papers/boxplots.pdf)
#
# $^{2}$[Principles and Procedures of Exploratory Data Analysis -<NAME>](https://pdfs.semanticscholar.org/4016/18eda85f341b0600f49811229cfeb50c2843.pdf)
#
# $^{3}$[Lesson 1.5 Exploratory Data Analysis](https://www.uth.tmc.edu/uth_orgs/educ_dev/oser/L1_5.HTM)
#
# $^{4}$ [What a Boxplot Can Tell You about a Statistical Data Set](https://www.dummies.com/education/math/statistics/what-a-boxplot-can-tell-you-about-a-statistical-data-set/)
#
#
# **General references**
#
# Below are some general references which I used to build up my understanding of box plots and also to assist with
# seaborn plots.
#
# https://en.wikipedia.org/wiki/Box_plot - box plot wiki page
#
# https://www.tutorialspoint.com/python/python_box_plots.htm - box plot tutorial
#
# https://www.mathsisfun.com/data/quartiles.html --reminder of how to work out quartiles!
#
# https://stackoverflow.com/questions/49554139/boxplot-of-multiple-columns-of-a-pandas-dataframe-on-the-same-figure-seaborn --assistance with SNS box plot
#
# https://www.wellbeingatschool.org.nz/information-sheet/understanding-and-interpreting-box-plots - interpreting box lots
#
# https://stats.stackexchange.com/questions/202629/what-information-does-a-box-plot-provide-that-a-histogram-does-not/202663 --discussion around merits of boxplots vs histograms
#
# http://datavizcatalogue.com/blog/box-plot-variations/ -- discussion of some of the variations of box plots
#
# **Conclusion**
#
# When I started this assignment I had never used boxplots nor had I seen them used by others. I am now left wondering why this is the case as they are both easy to construct (even by hand for smaller datasets) and relatively easy to understand.
#
# I think that for general review of stats they are easier to understand than histograms and I was particularly taken by the possibility of showing data trends over years and the fact that black and white plots made as much sense as colour, this is a sharp contrast to working with line graphs and others where colour is needed to distinguish data.
#
# I was also quite taken by strip plots and the combination of box and strip plots is probably one I will use going forward.
#
# ### END
| BOX PLOTS.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Solution is available in the "solution.ipynb"
import tensorflow as tf
# TODO: Convert the following to TensorFlow:
x = tf.constant(10)
y = tf.constant(2)
z = tf.subtract(tf.divide(x,y),tf.cast(tf.constant(1), tf.float64))
# TODO: Print z from a session as the variable output
with tf.Session() as sess:
output = sess.run(z)
# +
### DON'T MODIFY ANYTHING BELOW ###
### Be sure to run all cells above before running this cell ###
import grader
try:
grader.run_grader(output)
except Exception as err:
print(str(err))
# -
| Code/Quiz2_TF_Math/Solution_TF_Math.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# %sh
# mkdir /dbfs/init/
# cat > /dbfs/init/init_selenium.sh <<EOF
wget -N "https://chromedriver.storage.googleapis.com/96.0.4664.45/chromedriver_linux64.zip" -O /tmp/chromedriver.zip
unzip /tmp/chromedriver.zip -d /tmp/chromedriver/
sudo curl -sS -o - https://dl-ssl.google.com/linux/linux_signing_key.pub | apt-key add
sudo echo "deb https://dl.google.com/linux/chrome/deb/ stable main" >> /etc/apt/sources.list.d/google-chrome.list
sudo apt-get -y update
sudo apt-get -y install google-chrome-stable
wget https://dl.google.com/linux/direct/google-chrome-stable_current_amd64.deb
sudo dpkg -i google-chrome-stable_current_amd64.deb
pip install selenium
EOF
| init/Selenium_init.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import tensorflow as tf
x=tf.random.normal(shape=(2,2))
y=tf.random.normal(shape=(2,2))
# $ f(x,y) = \sqrt{(x^2 +y^2)}$
#
# $ \nabla f(x,y) = \frac{\partial f}{\partial x} \hat{\imath} + \frac{\partial f}{\partial y} \hat{\imath}$
with tf.GradientTape() as tape:
tape.watch(x)##I want to calcuate gradient wrt x
f=tf.sqrt(tf.square(x) + tf.square(y))
df_dx=tape.gradient(f,x)
print(df_dx)
with tf.GradientTape() as tape:
tape.watch(y)### I want to calculate gradient of y wrt y
f=tf.sqrt(tf.square(x)+tf.square(y))
df_dy=tape.gradient(f,y)
print(df_dy)
with tf.GradientTape() as tape:
#tape.watch(x)
#tape.watch(y)
f=tf.sqrt(tf.square(x)+tf.square(y))
df_dx,df_dy=tape.gradient(f,[x,y])
print(df_dx)
print(df_dy)
with tf.GradientTape() as tape:
tape.watch(x)
tape.watch(y)
f=tf.sqrt(tf.square(x)+tf.square(y))
df_dx,df_dy=tape.gradient(f,[x,y])
print(df_dx)
print(df_dy)
# # Without using tape we can calculate gradient by using tf.Variable
x1=tf.Variable(tf.random.normal(shape=(2,2)))
y1=tf.Variable(tf.random.normal(shape=(2,2)))
with tf.GradientTape() as tape:
f=tf.sqrt(tf.square(x1)+tf.square(y1))
df_dx,df_dy=tape.gradient(f,[x1,y1])
print(df_dx)
print(df_dy)
x=tf.Variable(3.)
y=tf.Variable(2.)
with tf.GradientTape() as tape:
f=tf.sqrt(tf.square(x)+tf.square(y))
df_dx,df_dy=tape.gradient(f,[x,y])
print(df_dx)
print(df_dy)
# ## Simple Linear Regreassion Example
# $ f(x) =W.x +b $
TRUE_W=3.0
TRUE_B=2.0
NUM_EXAMPLES=1000
x=tf.random.normal(shape=[NUM_EXAMPLES])
noise=tf.random.normal(shape=[NUM_EXAMPLES])
y=x*TRUE_W + TRUE_B + noise
import matplotlib.pyplot as plt
plt.scatter(x,y, c="b")
plt.show()
y=x*TRUE_W + TRUE_B
import matplotlib.pyplot as plt
plt.scatter(x,y, c="b")
plt.show()
# # Lest define a model
class MyModel(tf.Module):
def __init__(self, **kwargs):
super().__init__(**kwargs)
#initialize weights
self.w=tf.Variable(5.0)
self.b=tf.Variable(0.0)
def __call__(self,x):
return self.w*x+self.b
model=MyModel()
model(3)
model.w
model.b
model.variables
# # Lest define the loss function
def MSE_loss(target_y, predicted_y):
error=target_y-predicted_y
squared_error=tf.square(error)
mse=tf.reduce_mean(squared_error)
return mse
plt.scatter(x,y, c="b")
pred_y=model(x)## without training
plt.scatter(x,pred_y, c="r")
plt.show()
current_loss=MSE_loss(y, model(x))
current_loss.numpy()
# # Lets define the traning
def train(model,x,y,learning_rate):
with tf.GradientTape() as tape:
current_loss=MSE_loss(y, model(x))
dc_dw,dc_db=tape.gradient(current_loss,[model.w,model.b])
model.w.assign_sub(learning_rate * dc_dw)
model.b.assign_sub(learning_rate * dc_db)
model=MyModel()
ws,bs=[],[]
epochs=20
learning_rate=0.1
w=model.w.numpy()
b=model.b.numpy()
init_loss=MSE_loss(y, model(x))
print(f"Initial W: {w}, initial bias: {b}, initial_loss:{init_loss}")
for epoch in range(epochs):
train(model,x,y,learning_rate)
ws.append(model.w.numpy())
bs.append(model.b.numpy())
current_loss=MSE_loss(y, model(x))
print(f"For epoch: {epoch}, W: {ws[:-1]}, b: {bs[-1]}, Current_loss: {current_loss}")
plt.plot(range(epochs), ws,'r',range(epochs),bs,'b')
plt.plot([TRUE_W] * epochs,"r--",[TRUE_B]*epochs,"b--")
plt.legend(["w","b","True W","True_B"])
plt.show()
plt.scatter(x,y, c="b")
pred_y=model(x)## after training
plt.scatter(x,pred_y, c="r")
plt.show()
| Calculation of Gradient and build a simple linear model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # LASSO Regression
# +
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
from sklearn.linear_model import Lasso
import matplotlib.pyplot as plt
import seaborn as sns
from dmba import regressionSummary
# %matplotlib inline
# -
# ## Overview
#
# The Lasso is a shrinkage and selection method for linear regression. It minimizes the usual sum of squared errors, with a bound on the sum of the absolute values of the coefficients.
#
# It includes a "penalty" that is based on aggregation of coefficient values. Standardizing the variable is important in this method, as this will prepare your data for wrangling and will allow you to draw clearer conclusions.
#
# In this chapter we will provide an example of how to implement LASSO. This is a feature selection method that shrinks our coefficient estimates ($\hat{\beta}_i$s) towards zero, even setting some to be 0. This helps us create an easier-to-interpret model and reduces the risk of overfitting. (Recall, overfitting my model means I've made a model that fits too closely to the training data but isn't generalizable to other data - therefore, it won't make good predictions.)
#
# We'll implement LASSO using our beloved Toyota Corolla dataset. In the MLR notebook, we learned how to use MLR to make predictions about the price a used Toyota Corolla would sell for based on car features. In this notebook, we'll learn how to use LASSO to make those predictions.
# ### Toyota Corollas Data
# This data represents the sale of used Toyota Corollas in the Netherlands, and our mission is to **predict the price of a car based on its options and features**.
#
# Below, we show descriptions of some of the variables in our dataset.
#
# | Variable | Description |
# |------------|---------------------------------------|
# | Price | Offer price in Euros |
# | Age | Age in months as of August 2004 |
# | Kilometers | Accumulated Kilometers on odometer |
# | Fuel type | Fuel type (Petrol, Diesel, CNG) |
# | HP | Horsepower |
# | Metallic | Metallic color (Yes = 1, No= 0) |
# | Automatic | Automatic (Yes = 1, No = 0) |
# | CC | Cylinder volume in cubic centimeters |
# | Doors | Number of doors |
# | QuartTax | Quarterly road tax in Euros |
# | Weight | Weight in Kilograms |
# ## Data Processing
# Read the data using the ```read_csv``` function from pandas package and save it as the dataframe `car_df`. The data is called `'ToyotaCorolla.csv'`. Use the `.head()` method to make sure the dataframe has been created correctly.
car_df = pd.read_csv('../jb/data/ToyotaCorolla.csv')
car_df.head()
# We use the map function to iterate through a list of values (in this case, the column names, and apply a function to each item in the list. In this case, we want make each item lowercase.)
car_df.columns = map(str.lower, car_df.columns)
car_df.columns
# We can see that `id` and `model` aren't useful to include, as `id` provides a unique but meaningless number for each row, and model is not important, as these are all Toyota Corollas. Let's drop those.
car_df=car_df.drop(columns =['model','id'])
car_df.columns
# Recall that we have to convert categorical variables to numbers. We only have nominal (unordered) categorical variables, so they should be converted to a set of dummy variables.
#
# We've already dropped `model`, so our two remaining categorical variables are `color` and `fuel_type`.
# We use `pd.get_dummies` to dummy-code all the variables. Each categorical variable will generate two or more dummy variables. Dropping the first dummy variable (using the argument `drop_first`) ensures we won't have problems with perfect multicollinearity.
car_df = pd.get_dummies(car_df, drop_first=True)
car_df.columns
# Let's use bracket notation with the new column names to again select the first three rows. What do we see? The first car (row index equal to 0) had a blue car that used diesel. We see below that fuel_type_Diesel = 1, color_Blue = 1, and all the other dummy variables are equal to 0.
#
# For a car with fuel type CNG, what would fuel_type_Diesel and fuel_type_Petrol equal?
#
# How would I know from these new dummy variables that a car is Beige?
car_df[['fuel_type_Diesel', 'fuel_type_Petrol', 'color_Black', 'color_Blue', 'color_Green',
'color_Grey', 'color_Red', 'color_Silver', 'color_Violet',
'color_White', 'color_Yellow']].head(3)
# ---
# ## Model Building
# Now, we can finally move on to what we're really interested in - using a regularization technique called LASSO to help us decide which predictors we should include in our model.
# Let's define our predictors and outcome variable. **Lasso will help me determine which, if any, predictor variables I should drop.**
#
# We'll then create the training and test sets, using our `X` and `y` objects.
# +
outcome = 'price'
predictors = ['age_08_04', 'km', 'fuel_type_Petrol', 'fuel_type_Diesel', 'hp', 'met_color', 'automatic', 'cc', 'doors', 'quarterly_tax', 'weight']
X = car_df[predictors]
y = car_df[outcome]
train_X, valid_X, train_y, valid_y = train_test_split(X, y, test_size=0.4, random_state=1)
train_X
# -
# Remember that in a LASSO model, we penalize large coefficient estimates. The magnitude of a coefficient estimate can vary depending on the units of the variable. We address this by standardizing our data.
# +
#Standardization
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
train_X=scaler.fit_transform(train_X)
valid_X=scaler.transform(valid_X)
# -
# Fitting a LASSO model is very similar to fitting an MLR model, except we'll need to specify the penalty parameter, $\lambda$. Somewhat confusingly, we specify $\lambda$ in the `alpha` argument in the LASSO function. (LASSO is an algorithm in the scikit learn package.)
#
# We need to take the following steps.
# 1. Choose a value for $\lambda$ (the penalty parameter). Below, we set it to equal 1.
# 2. Initialize the LASSO algorithm, setting the $\lambda$ value (alpha argument)
# 3. Fit the model using the training data (this finds the best coefficients AKA the $\hat{\beta}_i$s)
# 4. Use the results (that is, use the coefficient estimates) from the fitted model to predict the outcome (price) for the validation data
# +
# Step 1: Set lambda
lambda_val = 1
# Step 2: load the lasso regression algorithm into an object called "car_lasso"
car_lasso = Lasso(alpha = lambda_val)
# Step 3: fit the lasso model using the training data
car_lasso.fit(train_X, train_y)
# Step 4 use the fitted model to predict the outcome for the validation data
car_lasso_predict = car_lasso.predict(valid_X)
# -
regressionSummary(valid_y, car_lasso_predict)
# Estimated coefficients
car_lasso.coef_
# In the previous step, we used a $\lambda$ value of 1 when fitting the LASSO regression model. However, how do we know whether 1 is a good choice for $\lambda$? We don't!
#
# The beauty of Python is that we can quickly try lots of different $\lambda$ values and find the one that does best, where doing best means minimizing the error. To do this, we'll put the above steps into a for loop, where we fit a LASSO regression model for many different values of $\lambda$. We can then choose the $\lambda$ value that results in the best prediction (lowest error).
#
# ---
# ## Plotting 𝜆 values
# Now, let's use the create a for loop to fit the LASSO regression model for many different $\lambda$ values. Let's break this down into several steps.
#
# First, decide what you want to use as your $\lambda$ values. We put this $\lambda$ values into a list. I've created a few lists of $\lambda$ values below. I've commented out all but one - you can uncomment to try to different lists or even make your own.
#
# For whatever list I create, I save the length of the list (that is, the number of $\lambda$ in my list) as `l_num`.
# +
# Define lambda to be used in LASS0
# lambdas = [0.1,1,5,10]
# lambdas = (0.1,1,5,10,100,500,1000)
lambdas = list(range(1, 40, 2))
# Length of lambda
l_num = len(lambdas)
# -
# Next, create an empty array (sort of a fancy list) to hold the hold the prediction accuracy measure for each fitted model. Remember, we'll have the same number of models as we have $\lambda$s in our list above, so we create an array that is the same length. We put in zeros as placeholders.
# initilize a vector to hold values of RMSE
valid_RMSE = np.zeros(l_num)
# <p>Now we're ready for our for loop! For every single $\lambda$ in our list above, we want to fit a LASSO regression model, using that $\lambda$ value. So, we need to iterate through our list of $\lambda$s. As I iterate through the list, I get two pieces of information about the $\lambda$: the index (it's order in the list) and the actual $\lambda$ value.</p>
# Okay, finally ready to put the for loop code together with the LASSO code! Let's do it.
# enumerate through lambdas with index and val
for ind, val in enumerate(lambdas):
# Initialize the lasso regression by setting the penalty parameter equal to lambda[ind]
car_lasso = Lasso(alpha = val)
# Fit the lasso model using training data
car_lasso.fit(train_X, train_y)
# Calculate the error using the RMSE function and using the validation set data
# Save this RMSE into the appropriate position in the RMSE array
valid_RMSE[ind] = rmse(valid_y,car_lasso.predict(valid_X))
# We've now fit {l_num} models and for each one we've saved the RMSE. Let's look at how they do using a plot!
#
# When you run the code, you'll see that for a smaller lambda value, the RMSE is higher (overfitting). As lambda increases, my RMSE starts to drop; the sweet spot is at around lambda = 5. As my $\lambda$ continutes to increase, my RMSE shoots up, my lambda starts to increase again. For these larger lambdas, I'm penalizing the magnitude of the coefficients to the point where I'm shrinking lots of the coefficients to zero, which means I'm (implicitly) dropping a lot of predictors out of my model. In this case, I'm underfitting.
plt.figure(figsize=(18, 8))
plt.plot(lambdas,valid_RMSE, 'bo-', label=r'$RMSE$ Test set', color="darkred", alpha=0.6, linewidth=3)
plt.xlabel('Lambda value'); plt.ylabel(r'$RMSE$')
#plt.xlim(0, 6)
plt.title(r'Evaluate lasso regression with different lambdas')
plt.legend(loc='best')
plt.grid()
# +
car_lasso = Lasso(alpha = 13)
car_lasso.fit(train_X, train_y)
car_lasso.coef_
for ind, val in enumerate(car_lasso.coef_):
predvar = predictors[ind]
print("The estimated coefficient for",predvar,"is",round(val,2))
# -
# Let's look at the "paths" of the coefficients as $\lambda$ increases. To do this, we take the following steps.
# 1. Set the list of $\lambda$s. Use a wider range to see when the coefficients are pushed down to zero.
# 2. Run the LASSO model for each of the $\lambda$ in the list. Save the coefficients generated from each fitted model
# 3. Create a plot of each of the coefficient estimates as a function of $\lambda$.
#
# Notice how as $\lambda$ increases, the magnitude of more and more of the coefficients are pushed towards zero? By the time $\lambda$ is 100, only `age_08_04`, `hp`, `quarterly_tax`, `weight`, and `km` have non-zero coefficients.
# +
# Set lambdas
lambdas = [0.1,15,50,100,500,1000, 3000]
# Fit lasso regression for each lambda, save coefficients
coefs = []
for val in lambdas:
car_lasso = Lasso(alpha = val)
car_lasso.fit(train_X, train_y)
coefs.append(car_lasso.coef_)
# Generate a plot
# Use log scale so that we can see it a little more easily
fig, ax = plt.subplots(figsize=(30, 10))
ax.plot(lambdas, coefs, marker="o")
ax.set_xscale('log')
plt.xlabel('lambda')
plt.ylabel('Coefficients (beta hats)')
plt.title('Lasso coefficients as a function of choice of lambda')
plt.axis('tight')
plt.legend(predictors,prop={'size': 15})
plt.show()
| jb/LASSO.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] nbgrader={"grade": false, "grade_id": "jupyter", "locked": true, "schema_version": 3, "solution": false}
# For this problem set, we'll be using the Jupyter notebook:
#
# 
# -
# ---
# ## Part A (2 points)
#
# Write a function that returns a list of numbers, such that $x_i=i^2$, for $1\leq i \leq n$. Make sure it handles the case where $n<1$ by raising a `ValueError`.
# + nbgrader={"grade": false, "grade_id": "squares", "locked": false, "schema_version": 3, "solution": true}
def squares(n):
"""Compute the squares of numbers from 1 to n, such that the
ith element of the returned list equals i^2.
"""
### BEGIN SOLUTION
if n < 1:
raise ValueError("n must be greater than or equal to 1")
return [i ** 2 for i in range(1, n + 1)]
### END SOLUTION
# -
# Your function should print `[1, 4, 9, 16, 25, 36, 49, 64, 81, 100]` for $n=10$. Check that it does:
squares(10)
# + nbgrader={"grade": true, "grade_id": "correct_squares", "locked": false, "points": 1, "schema_version": 3, "solution": false}
"""Check that squares returns the correct output for several inputs"""
assert squares(1) == [1]
assert squares(2) == [1, 4]
assert squares(10) == [1, 4, 9, 16, 25, 36, 49, 64, 81, 100]
assert squares(11) == [1, 4, 9, 16, 25, 36, 49, 64, 81, 100, 121]
# + nbgrader={"grade": true, "grade_id": "pedal_squares", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
from pedal import *
from pedal.environments.nbgrader import setup_environment
if setup_environment('squares'):
prevent_ast("If")
prevent_ast("If")
prevent_embedded_answer(1)
prevent_import('numpy')
# -
# + nbgrader={"grade": true, "grade_id": "squares_invalid_input", "locked": false, "points": 1, "schema_version": 3, "solution": false}
"""Check that squares raises an error for invalid inputs"""
try:
squares(0)
except ValueError:
pass
else:
raise AssertionError("did not raise")
try:
squares(-4)
except ValueError:
pass
else:
raise AssertionError("did not raise")
| tests/datafiles/jupyter_notebook_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
# + [markdown] slideshow={"slide_type": "-"}
# # Fast Fourier Transform snippets
#
# ## Documentation
#
# - Numpy implementation: http://docs.scipy.org/doc/numpy/reference/routines.fft.html
# - Scipy implementation: http://docs.scipy.org/doc/scipy/reference/fftpack.html
# -
# ## Import directives
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import cm
# ## Make data
pattern = np.zeros((4, 4))
pattern[1:3,1:3] = 1
pattern
signal = np.tile(pattern, (2, 2))
fig = plt.figure(figsize=(16.0, 10.0))
ax = fig.add_subplot(111)
ax.imshow(signal, interpolation='nearest', cmap=cm.gray)
# ## Fourier transform with Numpy
#
# ### Do the fourier transform
transformed_signal = np.fft.fft2(signal)
#transformed_signal
fig = plt.figure(figsize=(16.0, 10.0))
ax = fig.add_subplot(111)
ax.imshow(abs(transformed_signal), interpolation='nearest', cmap=cm.gray)
# ### Filter
max_value = np.max(abs(transformed_signal))
filtered_transformed_signal = transformed_signal * (abs(transformed_signal) > max_value*0.5)
# +
#filtered_transformed_signal[6, 6] = 0
#filtered_transformed_signal[2, 2] = 0
#filtered_transformed_signal[2, 6] = 0
#filtered_transformed_signal[6, 2] = 0
#filtered_transformed_signal[1, 6] = 0
#filtered_transformed_signal[6, 1] = 0
#filtered_transformed_signal[1, 2] = 0
#filtered_transformed_signal[2, 1] = 0
#filtered_transformed_signal
fig = plt.figure(figsize=(16.0, 10.0))
ax = fig.add_subplot(111)
ax.imshow(abs(filtered_transformed_signal), interpolation='nearest', cmap=cm.gray)
# -
# ### Do the reverse transform
filtered_signal = np.fft.ifft2(filtered_transformed_signal)
#filtered_signal
fig = plt.figure(figsize=(16.0, 10.0))
ax = fig.add_subplot(111)
ax.imshow(abs(filtered_signal), interpolation='nearest', cmap=cm.gray)
# +
#shifted_filtered_signal = np.fft.ifftshift(transformed_signal)
#shifted_filtered_signal
# +
#shifted_transformed_signal = np.fft.fftshift(transformed_signal)
#shifted_transformed_signal
| nb_dev_python/python_numpy_fourier_transform_en.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Import libraries
import requests
import pandas as pd
import re
import seaborn as sns
import matplotlib.pyplot as plt
from bs4 import BeautifulSoup
import numpy as np
import re
import unicodedata
from sklearn.decomposition import PCA
# %matplotlib inline
# -
# In part 1 (top universities) we will create functions to compute and plot the different required ratios that we will reuse in part 2.
#
# To scrap the datas (for both website) we used postman and saw there were a json string we could scrap and parse to get the ranking. For www.topuniversities.com we had to scrap the university specific page to get some more complete informations such as the number of students.
# # www.topuniversities.com
# The ranking is broken and contains only the top 199 universities even though the last university has rank 200, if we want to have 200 universities we should include the university ranked 201, as we want the top 200, we will take only the 199 first universities.
# +
r = requests.get("https://www.topuniversities.com/sites/default/files/qs-rankings-data/357051.txt")
json_dict = r.json()
json_dict = json_dict['data']
json_dict = json_dict[:199]
for e in json_dict:
#Delete useless informations
del e['nid']
del e['logo']
del e['core_id']
del e['score']
del e['cc']
del e['stars']
del e['guide']
#Get the university specific url
r = requests.get("https://www.topuniversities.com/" + e['url'])
soup = BeautifulSoup(r.text, 'html.parser')
#Get faculty members numbers
scrap_list = soup.find_all('div', class_='faculty-main wrapper col-md-4')
if(len(scrap_list) == 0): #If no informations about the faculty members is given
e['# Faculty members'] = 0
e['# Int. faculty members'] = 0
else:
n_list = scrap_list[0].find_all(class_="number")
if(len(n_list) == 2):
e['# Faculty members'] = int(re.sub(',', '', n_list[0].text.strip()))
e['# Int. faculty members'] = int(re.sub(',', '', n_list[1].text.strip()))
else: #If they don't have international faculty member
e['# Faculty members'] = int(re.sub(',', '', n_list[0].text.strip()))
e['# Int. faculty members'] = 0
#Get students numbers
n_list = soup.find_all('div', class_='students-main wrapper col-md-4')
if(len(n_list) != 0):
e['# Students'] = int(re.sub(',', '', n_list[0].find_all(class_="number")[0].text.strip()))
else:
e['# Students'] = 0
n_list = soup.find_all('div', class_='int-students-main wrapper col-md-4')
if(len(n_list) != 0):
e['# Int. students'] = int(re.sub(',', '', n_list[0].find_all(class_="number")[0].text.strip()))
else:
e['# Int. students'] = 0
#We don't need the url anymore, we can delete it
del e['url']
df1 = pd.DataFrame(json_dict)
df1=df1.rename(columns = {'rank_display':'Rank', 'title' : 'University', 'country' : 'Country', 'region' : 'Region'})
df1["Rank"] = df1["Rank"].apply(lambda x: int(re.sub("[^0-9]", "", x)))
df1.index = df1['University']
del df1['University']
# -
# ## Per university ratio
# ### Ratio between faculty members and students
# Best university according to ratio between faculty members and students. We assume that a bigger ratio is better. Even though it might not be desirable, we assume it is the case for this exercise.
# We plot the bar charts only for the first 30 top universites (according to the ratio) to get an idea of the trend.
def uni_ratio_fac_stu(df):
#Compute ratio
df['Ratio Faculty members'] = df['# Faculty members']/df['# Students']
#Sort the rows according to ratio and locate the top 30 ones
df_sorted_ratios = df.sort_values(by=['Ratio Faculty members'], ascending=False).head(30)
#Plot the bar chart and print the best one
fig, ax = plt.subplots(figsize=(20,10))
plot = sns.barplot(y=df_sorted_ratios['Ratio Faculty members'], x=df_sorted_ratios.index, ax=ax)
plot.set_xticklabels(plot.get_xticklabels(), rotation=90)
print("Best university according to ratio between faculty members and students is : " + df_sorted_ratios.iloc[0].name)
#Delete the ratio column of the dataframe
del df_sorted_ratios
uni_ratio_fac_stu(df1)
# As we can see except for the first few universites which clearly have a higher ratio than the others, the ratio tends to decrease in a linear fashion
# Let's compute the best university according to ratio between local and international students. We assume that a ratio of 1 is the best, that is the same number of local and international students.
# Again we only plot the first 30 top ones.
def uni_ratio_int_stu(df):
#Compute ratio
df['Ratio Int. students'] = df['# Int. students']/df['# Students']
#Sort the rows according to ratio and locate the top 30 ones
df_sorted_ratios = df.sort_values(by=['Ratio Int. students'], ascending=False).head(30)
#Plot the bar chart and print the best one
fig, ax = plt.subplots(figsize=(20,10))
plot = sns.barplot(y=df_sorted_ratios['Ratio Int. students'], x=df_sorted_ratios.index, ax=ax)
plot.set_xticklabels(plot.get_xticklabels(), rotation=90)
print("Best university according to ratio between local and international students is : " + df_sorted_ratios.iloc[0].name)
#Delete the ratio column of the dataframe
del df_sorted_ratios
uni_ratio_int_stu(df1)
# Same observations as for the previous bar charts. It is good to note that the universities and their order are not the same as the previous ones.
# ## Per country ratio
# ### Ratio between faculty members and students
# Let's find the best country according to the ratio between faculty members and students. Previous assumptions about what is the best ratio still hold.
def country_ratio_fac_stu(df):
#Group by country and compute the ratios
df['Ratio'] = df['# Faculty members']/df['# Students']
#Sort the rows according to ratios
sorted_df = df.sort_values(by=['Ratio'], ascending=False)
#Plot the bar chart and print the best one
fig, ax = plt.subplots(figsize=(20,10))
plot = sns.barplot(y=sorted_df.Ratio, x=sorted_df.index, ax=ax)
plot.set_xticklabels(plot.get_xticklabels(), rotation=50)
print("Best country according to ratio between faculty members and students is : " + sorted_df.iloc[0].name)
#Delete datas we don't need anymore
del df['Ratio']
del sorted_df
df1_country = df1.groupby(['Country']).agg(np.mean)
country_ratio_fac_stu(df1_country)
# Russia is the exception with a high number in comparison of the others. Again the rest decreases in a linear fashion.
# ### Ratio between local and international students
# Let's find the best country according to the ratio between local and international students. Previous assumptions about what is the best ratio still hold.
def country_ratio_int_stu(df):
#Compute the ratios
df['Ratio'] = df['# Int. students']/df['# Students']
#Sort the rows according to ratios
sorted_df = df.sort_values(by=['Ratio'], ascending=False)
#Plot the bar chart and print the best one
fig, ax = plt.subplots(figsize=(20,10))
plot = sns.barplot(y=sorted_df.Ratio, x=sorted_df.index, ax=ax)
plot.set_xticklabels(plot.get_xticklabels(), rotation=50)
print("Best country according to ratio between local and international students is : " + sorted_df.iloc[0].name)
#Delete datas we don't need anymore
del df
del sorted_df
country_ratio_int_stu(df1_country)
# This bar chart is a bit different than the previous ones. Indeed the first ratios decrease way faster than the last ones but we could still use a combintion of two linear functions to approximate the decrease.
# ## Per region ratio
# ### Ratio between faculty members and students
# Let's find the best region according to the ratio between faculty members and students. Previous assumptions about what is the best ratio still hold.
def region_ratio_fac_stu(df):
#Group by region and compute the ratios
df['Ratio'] = df['# Faculty members']/df['# Students']
#Sort the rows according to ratios
sorted_df = df.sort_values(by=['Ratio'], ascending=False)
#Plot the bar chart and print the best one
fig, ax = plt.subplots(figsize=(20,10))
df = df.groupby(['Region']).agg(sum)
plot = sns.barplot(y=sorted_df.Ratio, x=sorted_df.index, ax=ax)
plot.set_xticklabels(plot.get_xticklabels(), rotation=50)
print("Best region according to ratio between faculty members and students is : " + sorted_df.iloc[0].name)
#Delete datas we don't need anymore
del df['Ratio']
df1_region = df1.groupby(['Region']).agg(np.mean)
region_ratio_fac_stu(df1_region)
# ### Ratio between local and international students
# Let's find the best region according to the ratio between local and international students. Previous assumptions about what is the best ratio still hold.
def region_ratio_int_stu(df):
#Compute the ratios
df['Ratio'] = df['# Int. students']/df['# Students']
#Sort the rows according to ratios
sorted_df = df.sort_values(by=['Ratio'], ascending=False)
#Plot the bar chart and print the best one
fig, ax = plt.subplots(figsize=(20,10))
plot = sns.barplot(y=sorted_df.Ratio, x=sorted_df.index, ax=ax)
plot.set_xticklabels(plot.get_xticklabels(), rotation=50)
print("Best region according to ratio between local and international students is : " + sorted_df.iloc[0].name)
#Delete datas we don't need anymore
del df
del sorted_df
region_ratio_int_stu(df1_region)
# Oceania is clearly the leader here.
# # www.timeshighereducation.com
# ## Scraping
#
# Using Postman when browsing the page with the ranking, I saw that a request for a json file was made. It actually contains all the information we need in a single file.
# Helper to get the region way more easily
countries = dict(df1[['Country', 'Region']].drop_duplicates().values.tolist())
countries['Luxembourg'] = 'Europe'
# URL containing the requested data in json
# +
timeshighereducation_url = "https://www.timeshighereducation.com/sites/default/files/the_data_rankings/world_university_rankings_2018_limit0_369a9045a203e176392b9fb8f8c1cb2a.json"
ranking_brute = requests.get(timeshighereducation_url).json()['data']
# -
infos_to_keep = ["rank", "name", "location", "stats_number_students", "stats_pc_intl_students", "stats_student_staff_ratio"]
column_names = ["Rank", "University", "Country", "# Students", "% Int. students", "% Faculty members"]
# +
# creating dataframe
df2 = pd.DataFrame(ranking_brute[:200], index=range(1, 201), columns=infos_to_keep)
# more expressive column names
df2.columns = column_names
df2["Rank"] = df2["Rank"].apply(lambda x: int(re.sub("[^0-9]", "", x)))
# string to numerical values
df2["# Students"] = pd.to_numeric(df2["# Students"].map(lambda x: x.replace(",", "")))
df2["% Int. students"] = pd.to_numeric(df2["% Int. students"].map(lambda x: x.replace("%", ".")))
df2["% Faculty members"] = pd.to_numeric(df2["% Faculty members"])
# adding number of international students
df2["# Int. students"] = (df2["# Students"] * df2["% Int. students"] / 100).map(round)
# adding number of faculty members
df2["# Faculty members"] = (df2["# Students"] / df2["% Faculty members"]).map(round)
# Changing Russian federation to Russia
df2['Country'].replace('Russian Federation', 'Russia', inplace=True)
# adding regions
df2["Region"] = df2["Country"].map(lambda c: countries[c])
# Usign meaningful index
df2.index = df2['University']
del df2['University']
# Deleting % Int. students because it is not needed anymore
del df2['% Int. students']
del df2["% Faculty members"]
# -
# ## Per university ratio
# ### Ratio between faculty members and students
# Best university according to ratio between faculty members and students. We assume that a bigger ratio is better. Even though it might not be desirable, we assume it is the case for this exercise.
# Again we plots only the first 30 ones.
uni_ratio_fac_stu(df2)
# One university is clearly above the others, then it decreases in a linear fashion.
# ### Ratio between local and international students
# Best university according to ratio between local and international students. We assume that a ratio of 1 is the best, that is the same number of local and international students.
uni_ratio_int_stu(df2)
# We can see that LSE school has 70% of international students which is really impressive.
# ## Per country ratio
# ### Ratio between faculty members and students
# Let's find the best country according to 1) ratio between faculty members and students and 2) ratio between local and international students. Previous assumptions about what is the best ratio still hold.
df2_country = df2.groupby(['Country']).agg(np.mean)
country_ratio_fac_stu(df2_country)
# In Denmark there are around 6 students for a staff member. It's clearly above the average.
# ### Ratio between local and international students
country_ratio_int_stu(df2_country)
# English talking countries are more international (in term of international students ratio) in general and also small european countries like Luxembourg (really impressive how high), Austria and Switzerland.
# Let's find the best region according to 1) ratio between faculty members and students and 2) ratio between local and international students. Previous assumptions about what is the best ratio still hold.
# ## Per region ratio
# ### Ratio between faculty members and students
df2_region = df2.groupby(['Region']).agg(np.mean)
region_ratio_fac_stu(df2_region)
# It's quite suprising to see Africa first. We wouldn't have expect that at first. Although Europe is quite at the back.
# ### Ratio between local and international students
region_ratio_int_stu(df2_region)
# # Quick observations on both ranking
# - Oceania followed by Europe have the biggest ratio between local and internaionl students in both ranking
# - Latin america universities are not present in the top 200 of times higher education
# - Russia tends to have a lot of faculty members, it is at the top (not necessarily first place) in both bar charts.
# - Countries ratios are different between the two rankings which is expected as the universities are not the same, but we can see some countries around the same position in the bar charts of both ranking. This might show that in some cases the universities of a specific country tend to have more or less the same ratio of faculty members or international students. It could come from many factors, such as law enforcment, the culture or the international reputation of a country.
# - More observations are made below.
# # Combining the two websites
# To combine the two rankings, we will be using the name of the universities. However, the names are rarely exactly the same in both rankings, which means that we need to use regex to keep the important part of the name only. <br>
# Sometimes regex will not be enough, and we will need to change the name in one of the dataframe manually.<br>
# We start by working on the cases where regex is not enough.
df2.rename({'ETH Zurich – Swiss Federal Institute of Technology Zurich' : 'ETH Zurich – Swiss Federal Institute of Technology',
'Wageningen University & Research' : 'Wageningen University'}, inplace=True)
df1.rename({'UCL (University College London)': 'University College London',
'KAIST - Korea Advanced Institute of Science & Technology' : 'Korea Advanced Institute of Science and Technology (KAIST)',
'Ludwig-Maximilians-Universität München' : 'LMU Munich',
'Ruprecht-Karls-Universität Heidelberg' : 'Heidelberg University',
'University of North Carolina, Chapel Hill' : 'University of North Carolina at Chapel Hill',
'Trinity College Dublin, The University of Dublin' : 'Trinity College Dublin',
'KIT, Karlsruhe Institute of Technology' : 'Karlsruhe Institute of Technology',
'Humboldt-Universität zu Berlin' : 'Humboldt University of Berlin',
'Freie Universitaet Berlin': 'Free University of Berlin',
'Université de Montréal' : 'University of Montreal',
'Université Pierre et Marie Curie (UPMC)' : 'Pierre and Marie Curie University',
'Technische Universität Berlin (TU Berlin)' : 'Technical University of Berlin',
'Universitat Autònoma de Barcelona' : 'Autonomous University of Barcelona',
'Eberhard Karls Universität Tübingen' : 'University of Tübingen',
'Albert-Ludwigs-Universitaet Freiburg' : 'University of Freiburg',
'Scuola Superiore Sant\'Anna Pisa di Studi Universitari e di Perfezionamento': 'Scuola Superiore Sant’Anna'}, inplace=True)
# And then we use regex on every university name.
# +
regex1 = r'\([^()]*\)'
regex2 = r"(the)|(university)|(of)|(de)|(-)|( )|(’)|(')|\."
p = re.compile('('+regex1+'|'+regex2+')')
for df in [df1, df2]:
df['University_regex'] = df.index.values
# We first take care of the case
df['University_regex'] = df['University_regex'].apply(lambda x: x.casefold())
# Then we remove useless words, spaces, parentheses, their content and so on
df['University_regex'] = df['University_regex'].apply(lambda x: str(re.sub(p, '', x)))
# We then normalize the text to remove accents
df['University_regex'] = df['University_regex'].apply(lambda x: unicodedata.normalize("NFKD", x))
# And swith the encoding to utf-8
df['University_regex'] = df['University_regex'].apply(lambda x: x.rstrip().encode('ascii', errors='ignore')
.decode('utf-8'))
# -
# Once we know that the data is ready to be merged, we apply an inner merge. We chose to use an inner merge instead of an outer merge because it makes more sense to only keep universities present in both rankings.
df_merged = df2.reset_index().merge(df1, how='inner', on='University_regex').set_index('University')
# Once the merge is done, we select and clean the data we are interested in. Moreover, since the number of student is different depending on the ranking, we decided to keep the average.
# +
#Rename columns and put them in a new dataframe
df_all = df_merged.copy()
df_all['# Students (topuni)'] = df_merged['# Students_x']
df_all['# Students (times)'] = df_merged['# Students_y']
df_all['# Int. students (topuni)'] = df_merged['# Int. students_x']
df_all['# Int. students (times)'] = df_merged['# Int. students_y']
df_all['# Faculty members (topuni)'] = df_merged['# Faculty members_x']
df_all['# Faculty members (times)'] = df_merged['# Faculty members_y']
df_all['Rank topuni'] = df_merged['Rank_x']
df_all['Rank times'] = df_merged['Rank_y']
df_all['Country'] = df_merged['Country_x']
df_all['Region'] = df_merged['Region_x']
df_all['Ratio Int. students (topuni)'] = df_merged['Ratio Int. students_x']
df_all['Ratio Int. students (times)'] = df_merged['Ratio Int. students_y']
df_all['Ratio Faculty members (topuni)'] = df_merged['Ratio Faculty members_x']
df_all['Ratio Faculty members (times)'] = df_merged['Ratio Faculty members_y']
#Columns to keep
df_all = df_all[['# Students (topuni)', '# Students (times)',
'# Int. students (topuni)', '# Int. students (times)', '# Faculty members (topuni)',
'# Faculty members (times)', 'Rank topuni', 'Rank times', 'Country', 'Region',
'Ratio Int. students (topuni)', 'Ratio Int. students (times)', 'Ratio Faculty members (topuni)', 'Ratio Faculty members (times)']]
del df_merged
# -
# Plot of the correlation
fig, ax = plt.subplots(figsize=(10,10))
sns.heatmap(df_all.corr(), ax=ax, square=True)
# Above, a few things are interesting to notice:
# - When comparing the same corresponding data for each website, we can clearly distinguish a strong correlation which is something we can expect.
# - There is a negative correlation between the rankings and every other feature (except the # Students) which is logical since the rankings are the only caracteristics where lower is better.
# - There is a higher (absolute) correlation between the ranking of TopUniversity and the number of students than for Times Higher Education. We could guess from that that TopUniversity's criterias for the ranking give more importance to the size of the university than Times Higher Education's criterias.
# - There is an (absolute) correlation between the ratios and the rankings (higher ratios would have better ranks on average).
# - The number of faculty members is correlated with the number of students. Which is logic because it is not desirable to have a university with a lot of faculty members but very few students and inversly.
# - It is good to note that both rankings seem not to always agree on the number of students, faculty members and international students as the correlation is not of 1.
#
#
# *Note that by 'absolute' correlation we mean the absolute value of the correlation*
# ## Best university
# ### Basic method
# To find the best university according to both ranking, we could use the following method: We take the mean of both ranking and the university with the smallest mean is the best university. With this method we try to find the university closest to the top in both rankings.
# +
df_all['Rank mean'] = (df_all['Rank times'] + df_all['Rank topuni'])/2.0
best = df_all.sort_values(by=['Rank mean'], ascending=True).iloc[0]
print("The best university according to both ranking is " + best.name + " with a mean rank score of {}".format(best["Rank mean"]))
del df_all['Rank mean']
# -
# ### Alternative method
# What we could do, is extract the principal components of the data (A PCA, basically) and project the points on the main component.
pca = PCA(n_components=2)
# Creates pairs of ranks for each university
ranks = np.array([df_all[['Rank times']].values.flatten(), df_all[['Rank topuni']].values.flatten()]).T
# Tells the PCA what the data is and then get the data in its new basis
ranks_new_basis = pca.fit(ranks).transform(ranks)
# Get the ratio of importance for each component
pca_score = pca.explained_variance_ratio_
# Get the components
V = pca.components_
# Multiplies the components by their ratio of importance compared to the least important component
ranks_pca_axis_x, ranks_pca_axis_y = V.T * pca_score / pca_score.min()
# We then plotted the different Universities according to their respective ranking and showed the first two main component of the data (green and red).
# +
starting_point = (100, 100)
fig = plt.figure(figsize=(10, 10))
ax = fig.add_subplot(111)
plt.title('Universities ranking')
plt.plot(df_all[['Rank times']].values.flatten(), df_all[['Rank topuni']].values.flatten(), '.')
plt.plot([starting_point[0], starting_point[0] + ranks_pca_axis_x[0]*20], [starting_point[1], starting_point[1] + ranks_pca_axis_y[0]*20], label='Main component')
plt.plot([starting_point[0], starting_point[0] + ranks_pca_axis_x[1]*20], [starting_point[1], starting_point[1] + ranks_pca_axis_y[1]*20], label='Second component')
plt.xlabel('Ranking according to Times')
plt.ylabel('Ranking according to Topuni')
plt.legend()
plt.show()
# -
# Once we have the principle component, we can use it to transform our data and change its basis from (rank times, rank topuni) to (main component, second component)
# +
fig = plt.figure(figsize=(10, 10))
ax = fig.add_subplot(111)
plt.scatter(ranks_new_basis.T[0], ranks_new_basis.T[1])
# Creates main component arrow
plt.plot([-125, 150], [0, 0], c='g', label='Main component')
plt.plot([140, 150], [4, 0], c='g')
plt.plot([140, 150], [-4, 0], c='g')
plt.xlabel('Main component')
plt.ylabel('Second component')
plt.title('Universities ranking in new basis')
plt.legend()
plt.show()
# -
# To get a final ranking system, we simply project the points on the main component axis and rate them depending on how far left they are. The more on the left, the better the rank.
# Keep only main component value of data
ranks_pca1 = ranks_new_basis.T[0]
# Get indices of sorted data
temp = ranks_pca1.argsort()
# Creates new array and then aranges values depending on indices of sorted data
new_ranks = np.empty(len(ranks_pca1), int)
new_ranks[temp] = np.arange(len(ranks_pca1))
# We now add this new rank in the DataFrame and show the 20 first universities in our new ranking system.
df_all['Rank total'] = new_ranks + 1
# Best university according to both ranking :
print("Best university according to PCA is : " + df_all.sort_values(by='Rank total').iloc[0].name)
# As we can see with both the basic and alternative method Stanford University is the best according to both ranking. Which is not surpising considering its excellent reputation.
| 02 - Data from the Web/homework02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#default_exp model.modelmanager
# -
#hide
#from collections.abc import Iterable,Iterator,Generator,Sequence
#from collections import OrderedDict,defaultdict,Counter,namedtuple
# %load_ext autoreload
# %autoreload 2
# # Dataloader
#
# > create dataset and dataloader
# +
# export
from lib.data.lists import *
from lib.model.model import *
from lib.learner.learner import *
from functools import partial
import torch
from torch import Tensor
from torch import nn
class ModelManager():
def __init__(self,model):self.model=model
#@classmethod
#def create_from_model(model:nn.Module):
def find_modules(self,condition):
return find_submodules(self.model, condition)
def summary(self, xb:Tensor, only_leaves=True, print_mod=False):
#device = next(model.parameters()).device
#xb = xb.to(device)
f = lambda hook,mod,inp,out: print(f"\n{mod}\n{out.shape}") if print_mod else print(f"{type(mod)} {out.shape}")
mods = self.find_modules(lambda m: not isinstance(m, nn.Sequential) and not isinstance(m, ResBlock) ) if only_leaves else \
self.model.children()
with Hooks(mods, f) as hooks: self.model(xb)
def grads_summary(self):
modules = self.find_modules( condition=lambda m: not isinstance(m, nn.Sequential) )
for module in modules:
if len(list(module.children()))==0:
requires_grad = [p.requires_grad for p in module.parameters(recurse=False)]
str_requires_grad = "None "
if len(requires_grad) > 0:
str_requires_grad = "False" if sum(requires_grad) == 0 else "True " if sum(requires_grad)==len(requires_grad) else "None"
print(f"requires_grad: {str_requires_grad} : {type(module).__name__}")
def save(self, path, subdir="models"):
mdl_path = Path(path)/subdir
mdl_path.mkdir(exist_ok=True)
st = self.model.state_dict()
torch.save(st, mdl_path/'iw5')
def load(self, path, subdir="models"):
mdl_path = Path(path)/subdir
st = torch.load(mdl_path/'iw5')
self.model.load_state_dict(st)
@staticmethod
def set_grad(module, requires_grad, train_bn=False):
if isinstance(module, (nn.BatchNorm2d)): return
for p in module.parameters(recurse=False):
p.requires_grad_(requires_grad)
def change_requires_grad_(self, modules, requires_grad, train_bn):
condition = lambda m: not isinstance(m, nn.Sequential)
selection = []
for m in modules: selection.extend( ModelManager.find_submodules(m, condition) )
for m in selection: ModelManager.set_grad(m, requires_grad, train_bn)
def freeze( self, train_bn=False ):
self.change_requires_grad_([self.model[0]], requires_grad=False, train_bn=train_bn)
self.change_requires_grad_(self.model[1:], requires_grad=True, train_bn=train_bn)
def unfreeze( self, train_bn=False ):
self.change_requires_grad_(self.model, requires_grad=True, train_bn=train_bn)
def getFirstbatch(self, databunch:DataBunch, normalization:Callback ):
cbfs = [partial(BatchTransformXCallback, tfm = normalization), GetOneBatchCallback]
learn = Learner( self.model, databunch, loss_func=None)
learn.fit(1, opt=None, cb_funcs=cbfs)
cb = learn.find_subcription_by_cls(GetOneBatchCallback)
if cb is None: print("cb is None")
return cb.xb, cb.yb
def adapt_model(self, databunch, normalization):
#get rid of norm
cut = next( i for i,o in enumerate(self.model.children()) if isinstance(o,nn.AdaptiveAvgPool2d) )
m_cut = self.model[:cut]
xb,_ = self.getFirstbatch( databunch, normalization )
pred = m_cut(xb)
ni = pred.shape[1]
self.model = nn.Sequential(
m_cut,
#AdaptiveConcatPool2d(),
nn.AdaptiveAvgPool2d(1),
Flatten(),
nn.Linear(ni, databunch.c_out)
#nn.Linear(ni*2, data.c_out)
)
def predict(self, input_data, tfm_input):
with torch.no_grad():
return self.model( tfm_input(torch.tensor(input_data) ) )
class CnnModelManager(ModelManager):
def initialize(self, uniform:bool=False, a=0.0, nonlinearity="relu"):
#if isinstance(self.model,XResNet):
self.model.initialize(uniform=uniform, a=a, nonlinearity=nonlinearity)
# + active=""
# """
#
#
# def init_cnn_resnet(m):
# if getattr(m, 'bias', None) is not None: nn.init.constant_(m.bias, 0)
# if isinstance(m, (nn.Conv2d,nn.Linear)): nn.init.kaiming_normal_(m.weight)
# for l in m.children(): init_cnn_resnet(l)
#
# def bn_splitter(m):
# def _bn_splitter(l, g1, g2):
# if isinstance(l, nn.BatchNorm2d): g2 += l.parameters()
# elif hasattr(l, 'weight'): g1 += l.parameters()
# for ll in l.children(): _bn_splitter(ll, g1, g2)
#
# g1,g2 = [],[]
# _bn_splitter(m[0], g1, g2)
#
# g2 += m[1:].parameters()
# return g1,g2
# a,b = bn_splitter(learn.model)
# """
#
# -
# # Tests
# + active=""
# # test that we can load mnist or imagenette files
# from ai_pytorch.data.external import *
# import numpy as np
#
# path = untar_data(URLs.MNIST)
# #path = untar_data(URLs.IMAGENETTE_160)
# files = ImageList.from_files( path )
# print( f"path:{path}\nnb-files: {len(files)} Image size: {files[0].size}" )
# + active=""
# ibx_train = files.label_by_func( lambda path: path.parent.parent.name=="training" )
# labels = files.label_by_func( lambda path: int(path.parent.name) )
# uniques_labels = labels.unique()
# train,test = files.split2ways(ibx_train)
# train_labels, test_labels = labels.split2ways(ibx_train)
# ds_train = Dataset(train, train_labels )
# ds_test = Dataset(test, test_labels )
# dataBunch = DataBunch(ds_train, ds_test, c_in=1, c_out=len(uniques_labels))
# print(uniques_labels)
# + active=""
# type(files), type(train), type(test), type(train_labels), type(test_labels), len(files), len(train), len(test), len(train_labels), len(test_labels),
# + active=""
# # test that transformation of and PIL.Image og ndarray image to tensorIUmage and vice versa
# x1 = TensorImage.from_image( files[0] )
# x2 = TensorImage.from_image( files[1] )
#
# gph = Graphics()
# gph.show_image( TensorImage.as_image(x1)/255. )
# gph.show()
# gph.show_image( TensorImage.as_image(x2)/255. )
# + active=""
# # test that the tranformations works
# image = TensorImage.from_image( files[1] )
# affines = AffineTransforms([Rotation(.3,30), ShiftScale(.3,shift=0.5,scale=0.25)])
# image_tfm = affines(image)
#
# gph.show_tensorimage(image)
# gph.show()
# gph.show_tensorimage(image_tfm)
# -
# # Export scripts
from nbdev.export import notebook2script
notebook2script()
| 06_modelmanger.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## Ungraded Lab: Activation in Custom Layers
#
# In this lab, we extend our knowledge of building custom layers by adding an activation parameter. The implementation is pretty straightforward as you'll see below.
#
# ## Imports
import math
import numpy as np
import torch
import torch.nn as nn
from torch.nn.parameter import Parameter
from torchvision.datasets import MNIST
from torch.utils.data import Dataset, DataLoader
import torchvision.transforms as transforms
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
print("device: ", device)
# ## Adding an activation layer
#
# To use the built-in activations in PyTorch, we can specify an activation parameter in the __init__() method of our custom layer class. From there, we can initialize it by using the forward() method. Then, you can pass in the forward computation to this activation.
class SimpleDense(nn.Module):
def __init__(self, in_features, out_features, bias=True, activation=None, device=None, dtype=None):
super(SimpleDense, self).__init__()
factory_kwargs = {'device': device, 'dtype': dtype}
# Input
self.in_features = in_features
# Bias
self.bias = bias
self.apply_activation = False
# Activation
if activation is not None:
self.apply_activation = True
self.activation = getattr(nn.functional, activation)
# Weight
self.weight = Parameter(torch.empty((out_features, in_features), **factory_kwargs))
# Bias
if bias:
self.bias = Parameter(torch.empty(out_features, **factory_kwargs))
else:
self.register_parameter('bias', None)
# Weight and Biase initialization
self._reset_parameters()
def forward(self, input):
x, y = input.shape
if y != self.in_features:
print(f'Wrong Input Features. Please use tensor with {self.in_features} Input Features')
return 0
output = input.matmul(self.weight.t())
if self.bias is not None:
output += self.bias
ret = output
if self.apply_activation:
return self.activation(ret)
return ret
def _reset_parameters(self):
torch.nn.init.kaiming_uniform_(self.weight, a=math.sqrt(5))
if self.bias is not None:
fan_in, _ = torch.nn.init._calculate_fan_in_and_fan_out(self.weight)
bound = 1 / math.sqrt(fan_in)
torch.nn.init.uniform_(self.bias, -bound, bound)
# ## Prepare the Dataset
# Image Transform
transform = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize(mean=(0.5,), std=(0.5,))
])
# Load Dataset
train_data = MNIST(root='./', train=True, download=True, transform=transform)
test_data = MNIST(root='./', train=False, download=True, transform=transform)
# DataLoader
train_loader = DataLoader(dataset=train_data,
batch_size=32,
shuffle=True,
num_workers=2,
pin_memory=True)
val_loader = DataLoader(dataset=test_data,
batch_size=32,
shuffle=True,
num_workers=2,
pin_memory=True)
# Build the Model
model = nn.Sequential(
SimpleDense(in_features=784, out_features=128, activation="relu"),
nn.Dropout(0.2),
SimpleDense(in_features=128, out_features=10),
nn.LogSoftmax(dim=1),
)
model.to(device)
# Optimizer
optimizer = torch.optim.Adam(model.parameters(), lr=0.001)
criterion = torch.nn.CrossEntropyLoss()
# +
# Train the Model
EPOCHS = 5
model.train()
for epoch in range(EPOCHS):
running_loss = 0
correct = 0
for data in train_loader:
images, labels = data
images = images.view(images.shape[0], -1)
images, labels = images.to(device), labels.to(device)
optimizer.zero_grad()
output = model(images)
loss = criterion(output, labels)
loss.backward()
optimizer.step()
pred = output.data.max(1, keepdim=True)[1]
correct += pred.eq(labels.data.view_as(pred)).cpu().sum()
running_loss += loss.item()
print(f"Epoch: {epoch}, loss: {running_loss/len(train_loader)}, accuracy: {correct/len(train_loader.dataset)}")
# Evaluate Trained Model
running_loss = 0
correct = 0
model.eval()
for data in val_loader:
images, labels = data
images = images.view(images.shape[0], -1)
images, labels = images.to(device), labels.to(device)
output = model(images)
loss = criterion(output, labels)
pred = output.data.max(1, keepdim=True)[1]
correct += pred.eq(labels.data.view_as(pred)).cpu().sum()
running_loss += loss.item()
print(f"\nValidation - loss: {running_loss/len(val_loader)}, accuracy: {correct/len(val_loader.dataset)}")
| Course-1 Custom Models, Layers and Loss Functions/Week-3/C1_W3_Lab_3_custom-layer-activation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6 (seaborn-dev)
# language: python
# name: seaborn-dev
# ---
# + active=""
# .. _basic_tutorial:
#
# .. currentmodule:: seaborn
# -
# # Basic plots to show numeric relationships
# + active=""
# <TODO introductory copy>
# -
# %matplotlib inline
import numpy as np
import pandas as pd
import matplotlib as mpl
import matplotlib.pyplot as plt
import seaborn as sns
sns.set(style="darkgrid", color_codes=True)
np.random.seed(sum(map(ord, "basic")))
# + active=""
# Emphasizing continuity with line plots
# --------------------------------------
#
# With some datasets, you may want to understand changes in one variable as a function of another variable that has some sense of continuity. This is most common when one of the variables represents time, either abstractly or with a datetime object. In this situation, a good choice is to draw a line plot. In seaborn, this can be accomplished with the :func:`lineplot` function.
#
# The simplest case is when you have a vector of timepoints and a vector of values. To draw the function relating these variables, pass each to the ``x`` and ``y`` parameters of the :func:`lineplot` function, respectively:
# -
time = pd.date_range("2017-01-01", periods=24 * 31, freq="h")
value = np.random.randn(len(time)).cumsum()
sns.lineplot(x=time, y=value);
# + active=""
# Because :func:`lineplot` assumes that you are most often trying to draw ``y`` as a function of ``x``, the default behavior is to sort the data by the ``x`` values before plotting. However, this can be disabled:
# -
x, y = np.random.randn(2, 1000).cumsum(axis=1)
sns.lineplot(x=x, y=y, sort=False);
# + active=""
# Aggregation and representing uncertainty
# ----------------------------------------
#
# More complex datasets will have multiple measurements for the same value of the ``x`` variable. The default behavior in seaborn is to aggregate the multiple measurements at each ``x`` value by plotting the mean and the 95% confidence interval around the mean:
# -
fmri = sns.load_dataset("fmri")
sns.lineplot(x="timepoint", y="signal", data=fmri);
# + active=""
# The confidence intervals are computed using bootstrapping, which can be time-intensive for larger datasets. It's therefore possible to disable them:
# -
sns.lineplot(x="timepoint", y="signal", data=fmri, ci=None);
# + active=""
# Another good option, especially with larger data, is to represent the spread of the distribution at each timepoint by plotting the standard deviation instead of a confidence interval:
# -
sns.lineplot(x="timepoint", y="signal", data=fmri, ci="sd");
# + active=""
# To turn off aggregation altogether, set the ``estimator`` parameter to ``None``.
# -
sns.lineplot(x="timepoint", y="signal", data=fmri, estimator=None);
# + active=""
# The ``estimator`` parameter can also be used to control what method is used to aggregate the data.
# + active=""
# Plotting subsets of data with semantic mappings
# -----------------------------------------------
#
# Often there will be multiple measurements at each value of ``x`` because we want to know how the relationship between ``x`` and ``y`` changes as a function of other variables. The :func:`lineplot` function allows you to define up to three additional variables that will be used to subset the data. These variables are then semantically mapped by the color (``hue``), width (``size``) and dashes/markers (``style``) used to draw the lines. For example, we can draw two line plots with different colors simply by defining a variable to be used for ``hue`` subsets:
# -
sns.lineplot(x="timepoint", y="signal", hue="event", data=fmri);
# + active=""
# By adding a separate ``style`` variable, we can explore more complex relationships:
# -
sns.lineplot(x="timepoint", y="signal", hue="region", style="event", data=fmri);
# + active=""
# Be cautious about making plots with multiple subset variables. While sometimes informative, they can also be very difficult to parse and interpret. However, even when you are only examining changes across one subset variable, it can be useful to alter both the color and style of the lines, which can make the plot more accessible when printed to black-and-white or viewed by someone with colorblindness:
# -
sns.lineplot(x="timepoint", y="signal", hue="event", style="event", data=fmri);
# + active=""
# Some effort has been put into choosing good defaults so that you do not need to spend time specifying plot attributes for quick exploration, but the way the ``hue`` and ``style`` variables are mapped can be controlled through various parameters to the :func:`lineplot` function:
# -
sns.lineplot(x="timepoint", y="signal", hue="region", style="event",
palette="Set2", hue_order=["frontal", "parietal"],
dashes=["", (1, 1)],
data=fmri);
# + active=""
# By default, the ``style`` variable is represented by drawing lines with different dash patterns, but you can also draw markers with different shapes at the exact position of each observation:
# -
sns.lineplot(x="timepoint", y="signal", hue="region", style="event",
palette="Set2", hue_order=["frontal", "parietal"],
dashes=False, markers=True,
data=fmri);
# + active=""
# In the above examples, the ``hue`` variable takes different categorical values, and the colors of the lines are chosen with an appropriate qualitative colormap. When the ``hue`` variable is instead numeric (specifically, if it can be cast to float), the default behavior is to use a sequential colormap and to make a legend with "ticks" instead of an entry for each line (allowing it to scale to showing many lines):
# -
dots = sns.load_dataset("dots").query("align == 'dots'")
sns.lineplot(x="time", y="firing_rate",
hue="coherence", style="choice",
data=dots);
# + active=""
# A non-default colormap can be selected in this case by passing a colormap name or object, and you can ask for a ``"full"`` legend:
# -
cmap = sns.cubehelix_palette(light=.7, as_cmap=True)
sns.lineplot(x="time", y="firing_rate",
hue="coherence", style="choice",
palette=cmap,
legend="full", data=dots);
# + active=""
# It may happen that, even though the ``hue`` variable is numeric, it is poorly represented by a linear color scale. That's the case here, where the levels of the ``hue`` variable are logarithmically scaled. You can provide specific color values for each line by passing a list or dictionary:
# -
palette = sns.cubehelix_palette(light=.7, n_colors=6)
sns.lineplot(x="time", y="firing_rate",
hue="coherence", style="choice",
palette=palette,
data=dots);
# + active=""
# Another option for semantically mapping a subset is to change the width of its lines, which is accomplished by defining the ``size`` parameter:
# -
sns.lineplot(x="time", y="firing_rate",
size="coherence", style="choice",
data=dots);
# + active=""
# It's possible to control the range of line widths that are spanned by the data using the ``sizes`` parameter. Here we pass a ``(min, max)`` tuple, it it's also possible to pass a list of dictionary to precisely specify the width of each line:
# -
sns.lineplot(x="time", y="firing_rate",
size="coherence", style="choice",
sizes=(1, 2),
data=dots);
# + active=""
# While the ``size`` variable will typically be numeric, it's also possible to map a categorical variable with the width of the lines. Be cautious when doing so, because it will be very difficult to distinguish much more than "thick" vs "thin" lines. However, dashes can be hard to perceive when lines have considerable high-frequency variability, so using different widths may be helpful:
# -
palette = sns.cubehelix_palette(light=.7, n_colors=6)
sns.lineplot(x="time", y="firing_rate",
hue="coherence", size="choice",
palette=palette,
data=dots);
# + active=""
# Plotting with "long" versus "wide" data
# ---------------------------------------
#
# Like many other functions, :func:`lineplot` is most flexible when it is provided with "long-
# form" (or "tidy") data, typically in the form of a DataFrame where each column represents a variable and each row represents an observation. However, data are not always naturally generated or stored in long-form format, and it can be helpful to be able to take a quick look without reformatting. To support this, :func:`lineplot` can visualize a number of different "wide-form" representations if they are passed to ``data``. For instance, you can pass a wide DataFrame, which will draw a line for each column using the index for the ``x`` values:
# -
date = pd.date_range("2017-01-01", periods=365)
vals = np.random.randn(365, 4).cumsum(axis=0)
wide_df = pd.DataFrame(vals, date, list("ABCD"))
sns.lineplot(data=wide_df);
# + active=""
# Numpy arrays are handled similarly, except you lose the label information that you get from pandas:
# -
wide_array = np.asarray(wide_df)
sns.lineplot(data=wide_array);
# + active=""
# You can even pass in a list of objects with heterogeneous indices:
# -
wide_list = [wide_df.loc[:"2017-09-01", "A"], wide_df.loc["2017-05-1":, "B"]]
sns.lineplot(data=wide_list);
| doc/tutorial/basic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# <img src="images/dask_horizontal.svg" align="right" width="30%">
# # Data Storage
# <img src="images/hdd.jpg" width="20%" align="right">
# Efficient storage can dramatically improve performance, particularly when operating repeatedly from disk.
#
# Decompressing text and parsing CSV files is expensive. One of the most effective strategies with medium data is to use a binary storage format like HDF5. Often the performance gains from doing this is sufficient so that you can switch back to using Pandas again instead of using `dask.dataframe`.
#
# In this section we'll learn how to efficiently arrange and store your datasets in on-disk binary formats. We'll use the following:
#
# 1. [Pandas `HDFStore`](http://pandas.pydata.org/pandas-docs/stable/io.html#io-hdf5) format on top of `HDF5`
# 2. Categoricals for storing text data numerically
#
# **Main Take-aways**
#
# 1. Storage formats affect performance by an order of magnitude
# 2. Text data will keep even a fast format like HDF5 slow
# 3. A combination of binary formats, column storage, and partitioned data turns one second wait times into 80ms wait times.
# ## Setup
# Create data if we don't have any
from prep import accounts_csvs
accounts_csvs(3, 1000000, 500)
# ## Read CSV
# First we read our csv data as before.
#
# CSV and other text-based file formats are the most common storage for data from many sources, because they require minimal pre-processing, can be written line-by-line and are human-readable. Since Pandas' `read_csv` is well-optimized, CSVs are a reasonable input, but far from optimized, since reading required extensive text parsing.
import os
filename = os.path.join('data', 'accounts.*.csv')
filename
import dask.dataframe as dd
df_csv = dd.read_csv(filename)
df_csv.head()
# ### Write to HDF5
# HDF5 and netCDF are binary array formats very commonly used in the scientific realm.
#
# Pandas contains a specialized HDF5 format, `HDFStore`. The ``dd.DataFrame.to_hdf`` method works exactly like the ``pd.DataFrame.to_hdf`` method.
target = os.path.join('data', 'accounts.h5')
target
# convert to binary format, takes some time up-front
# %time df_csv.to_hdf(target, '/data')
# same data as before
df_hdf = dd.read_hdf(target, '/data')
df_hdf.head()
# ### Compare CSV to HDF5 speeds
# We do a simple computation that requires reading a column of our dataset and compare performance between CSV files and our newly created HDF5 file. Which do you expect to be faster?
# %time df_csv.amount.sum().compute()
# %time df_hdf.amount.sum().compute()
# Sadly they are about the same, or perhaps even slower.
#
# The culprit here is `names` column, which is of `object` dtype and thus hard to store efficiently. There are two problems here:
#
# 1. How do we store text data like `names` efficiently on disk?
# 2. Why did we have to read the `names` column when all we wanted was `amount`
# ### 1. Store text efficiently with categoricals
# We can use Pandas categoricals to replace our object dtypes with a numerical representation. This takes a bit more time up front, but results in better performance.
#
# More on categoricals at the [pandas docs](http://pandas.pydata.org/pandas-docs/stable/categorical.html) and [this blogpost](http://matthewrocklin.com/blog/work/2015/06/18/Categoricals).
# Categorize data, then store in HDFStore
# %time df_hdf.categorize(columns=['names']).to_hdf(target, '/data2')
# It looks the same
df_hdf = dd.read_hdf(target, '/data2')
df_hdf.head()
# But loads more quickly
# %time df_hdf.amount.sum().compute()
# This is now definitely faster than before. This tells us that it's not only the file type that we use but also how we represent our variables that influences storage performance.
#
# How does the performance of reading depend on the scheduler we use? You can try this with threaded, processes and distributed.
#
# However this can still be better. We had to read all of the columns (`names` and `amount`) in order to compute the sum of one (`amount`). We'll improve further on this with `parquet`, an on-disk column-store. First though we learn about how to set an index in a dask.dataframe.
# ### Exercise
# `fastparquet` is a library for interacting with parquet-format files, which are a very common format in the Big Data ecosystem, and used by tools such as Hadoop, Spark and Impala.
target = os.path.join('data', 'accounts.parquet')
df_csv.categorize(columns=['names']).to_parquet(target)
# Investigate the file structure in the resultant new directory - what do you suppose those files are for?
#
# `to_parquet` comes with many options, such as compression, whether to explicitly write NULLs information (not necessary in this case), and how to encode strings. You can experiment with these, to see what effect they have on the file size and the processing times, below.
# ls -l data/accounts.parquet/
df_p = dd.read_parquet(target)
# note that column names shows the type of the values - we could
# choose to load as a categorical column or not.
df_p.dtypes
# Rerun the sum computation above for this version of the data, and time how long it takes. You may want to try this more than once - it is common for many libraries to do various setup work when called for the first time.
# %time df_p.amount.sum().compute()
# When archiving data, it is common to sort and partition by a column with unique identifiers, to facilitate fast look-ups later. For this data, that column is `id`. Time how long it takes to retrieve the rows corresponding to `id==100` from the raw CSV, from HDF5 and parquet versions, and finally from a new parquet version written after applying `set_index('id')`.
# %%timeit 1000
df_csv.loc[df_csv['id'] == 100]
# %%timeit 1000
df_hdf.loc[df_hdf['id'] == 100]
# %%timeit 10000
df_p.loc[df_p['id'] == 100]
new_target = os.path.join('data', 'accounts_new.parquet')
df_p.set_index('id').to_parquet(new_target)
df_p_new = dd.read_parquet(new_target)
# %%timeit 10000
df_p_new.loc[df_p_new.index == 100]
import dask
dask.__version__
import pandas
pandas.__version__
# ## Remote files
# Dask can access various cloud- and cluster-oriented data storage services such as Amazon S3 or HDFS
#
# Advantages:
# * scalable, secure storage
#
# Disadvantages:
# * network speed becomes bottleneck
#
# The way to set up dataframes (and other collections) remains very similar to before. Note that the data here is available anonymously, but in general an extra parameter `storage_options=` can be passed with further details about how to interact with the remote storage.
#
# ```python
# taxi = dd.read_csv('s3://nyc-tlc/trip data/yellow_tripdata_2015-*.csv',
# storage_options={'anon': True})
# ```
# **Warning**: operations over the Internet can take a long time to run. Such operations work really well in a cloud clustered set-up, e.g., amazon EC2 machines reading from S3 or Google compute machines reading from GCS.
| 07_dataframe_storage.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: anlp37
# language: python
# name: anlp37
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/c-w-m/anlp-tf2/blob/master/chapter5-nlg-with-transformer-gpt/charRNN-text-generation.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# -
# # Imports
# +
import tensorflow as tf
import numpy as np
import pandas as pd
tf.__version__
# +
######## GPU CONFIGS FOR RTX 2070 ###############
## Please ignore if not training on GPU ##
## this is important for running CuDNN on GPU ##
tf.keras.backend.clear_session() #- for easy reset of notebook state
# chck if GPU can be seen by TF
tf.config.list_physical_devices('GPU')
#tf.debugging.set_log_device_placement(True)
gpus = tf.config.experimental.list_physical_devices('GPU')
if gpus:
# Restrict TensorFlow to only use the first GPU
try:
tf.config.experimental.set_memory_growth(gpus[0], True)
tf.config.experimental.set_visible_devices(gpus[0], 'GPU')
logical_gpus = tf.config.experimental.list_logical_devices('GPU')
print(len(gpus), "Physical GPUs,", len(logical_gpus), "Logical GPU")
except RuntimeError as e:
# Visible devices must be set before GPUs have been initialized
print(e)
###############################################
# -
# # Setup Tokenization
# +
chars = sorted(set("abcdefghijklmnopqrstuvwxyz0123456789 -,;.!?:’’’/\|_@#$%ˆ&*˜‘+-=()[]{}' ABCDEFGHIJKLMNOPQRSTUVWXYZ"))
chars = list(chars)
EOS = '<EOS>'
UNK = "<UNK>"
PAD = "<PAD>" # need to move mask to '0'index for TF
chars.append(UNK)
chars.append(EOS) #end of sentence
## need to handle padding characters as well
chars.insert(0, PAD) # now padding should get index of 0
# -
# Creating a mapping from unique characters to indices
char2idx = {u:i for i, u in enumerate(chars)}
idx2char = np.array(chars)
def char_idx(c):
# takes a character and returns an index
# if character is not in list, returns the unknown token
if c in chars:
return char2idx[c]
return char2idx[UNK]
# # Load the Model
# +
# Length of the vocabulary in chars
vocab_size = len(chars)
# The embedding dimension
embedding_dim = 256
# Number of RNN units
rnn_units = 1024
# Batch size
BATCH_SIZE=1
# +
# Define the model
# this one is without padding masking or dropout layer
def build_gen_model(vocab_size, embedding_dim, rnn_units, batch_size):
model = tf.keras.Sequential([
tf.keras.layers.Embedding(vocab_size, embedding_dim,
batch_input_shape=[batch_size, None]),
tf.keras.layers.GRU(rnn_units,
return_sequences=True,
stateful=True,
recurrent_initializer='glorot_uniform'),
tf.keras.layers.Dense(vocab_size)
])
return model
gen_model = build_gen_model(vocab_size, embedding_dim, rnn_units, BATCH_SIZE)
# +
# Now setup the location of the checkpoint
# and load the latest checkpoint
# Directory where the checkpoints will be saved
checkpoint_dir = './training_checkpoints/2020-Oct-01-14-29-55'
gen_model.load_weights(tf.train.latest_checkpoint(checkpoint_dir))
gen_model.build(tf.TensorShape([1, None]))
# -
def generate_text(model, start_string, temperature=0.7, num_generate=75):
# Low temperatures results in more predictable text.
# Higher temperatures results in more surprising text.
# Experiment to find the best setting.
# Converting our start string to numbers (vectorizing)
input_eval = [char2idx[s] for s in start_string]
input_eval = tf.expand_dims(input_eval, 0)
# Empty string to store our results
text_generated = []
# Here batch size == 1
for i in range(num_generate):
predictions = model(input_eval)
# remove the batch dimension
predictions = tf.squeeze(predictions, 0)
# using a categorical distribution to predict the word returned by the model
predictions = predictions / temperature
predicted_id = tf.random.categorical(predictions, num_samples=1)[-1,0].numpy()
# We pass the predicted word as the next input to the model
# along with the previous hidden state
input_eval = tf.expand_dims([predicted_id], 0)
text_generated.append(idx2char[predicted_id])
# lets break is <EOS> token is generated
#if idx2char[predicted_id] == EOS:
# break #end of a sentence reached, lets stop
return (start_string + ''.join(text_generated))
print(generate_text(gen_model, start_string=u"Obama"))
print(generate_text(gen_model, start_string=u"Apple"))
# ## 0.6993 model
print(generate_text(gen_model, start_string=u"Google"))
print(generate_text(gen_model, start_string=u"S&P"))
print(generate_text(gen_model, start_string=u"Market"))
print(generate_text(gen_model, start_string=u"Beyonce"))
# ## 0.7031 model
print(generate_text(gen_model, start_string=u"Apple"))
print(generate_text(gen_model, start_string=u"Google"))
print(generate_text(gen_model, start_string=u"S&P"))
print(generate_text(gen_model, start_string=u"Market"))
print(generate_text(gen_model, start_string=u"Beyonce"))
# ## Different temperature settings with the same model
print(generate_text(gen_model, start_string=u"S&P", temperature=0.1))
print(generate_text(gen_model, start_string=u"S&P", temperature=0.3))
print(generate_text(gen_model, start_string=u"S&P", temperature=0.5))
print(generate_text(gen_model, start_string=u"S&P", temperature=0.7))
print(generate_text(gen_model, start_string=u"S&P", temperature=0.9))
print(generate_text(gen_model, start_string=u"Kim", temperature=0.9))
print(generate_text(gen_model, start_string=u"Kim", temperature=0.7))
print(generate_text(gen_model, start_string=u"Kim", temperature=0.5))
print(generate_text(gen_model, start_string=u"Kim", temperature=0.3))
print(generate_text(gen_model, start_string=u"Kim", temperature=0.1))
# ## in progress model
# +
gen_model2 = build_gen_model(vocab_size, embedding_dim, rnn_units, BATCH_SIZE)
checkpoint_dir = './training_checkpoints/'+ '2020-Jun-02-22-38-17' # -> with 0.6993 loss
#''2020-Jun-02-01-02-14' # -> .7031 loss
gen_model2.load_weights(tf.train.latest_checkpoint(checkpoint_dir))
gen_model2.build(tf.TensorShape([1, None]))
# -
print(generate_text(gen_model2, start_string=u"S&P",
temperature=1, num_generate=75))
print(generate_text(gen_model2, start_string=u"S", temperature=0.7))
print(generate_text(gen_model2, start_string=u"NBA", temperature=0.4))
print(generate_text(gen_model2, start_string=u"Sta", temperature=0.7))
# # Greedy Search with Bigrams
# !wget http://norvig.com/tsv/ngrams-all.tsv.zip
# gzip file masquerading as a ZIP file
# !mv ngrams-all.tsv.zip ngrams-all.tsv.z
# !gunzip ngrams-all.tsv.z
# First 27 rows are characters
# next 669 are bigrams
# !head -n 697 ngrams-all.tsv | tail -n 669 > bigrams.tsv
# file format col1: bigram, col2: overall freq, ignore other cols
from collections import Counter
import csv
counts = Counter()
bitree = {}
totals = 2819662855499
with open("bigrams.tsv", 'r') as fl:
big = csv.reader(fl, delimiter='\t')
for bigram in big:
key = bigram[0]
count = int(bigram[1]) / totals
counts[key] = count
if(key[0] in bitree):
bitree[key[0]][key[1]] = count
else:
#need new subdict
bitree[key[0]] = { key[1] : count }
counts.most_common(10)
print(bitree['T'])
# !pip install anytree
# +
from anytree import Node, RenderTree
# construct 5 letter word o given a bigram
start = 'WI'
compl = Node(start[0], prob=1) # to store comlpetions and probabilities
cnt = 0
def recurse(letter, prob, level, parent):
if level > 2:
return Node(letter, parent=parent, prob=prob*parent.prob)
items = Counter(bitree[letter]).most_common(3)
nd = Node(letter, parent=parent, prob=parent.prob*prob)
for item in items:
rslt = recurse(item[0], item[1], level+1, nd)
return nd
recurse(start[1], 1, 0, compl)
# -
for pre, fill, node in RenderTree(compl):
print("%s%s (%2.8f)" % (pre, node.name, node.prob))
| chapter5-nlg-with-transformer-gpt/charRNN-text-generation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
from itertools import combinations
from sklearn.metrics import f1_score
from sklearn.metrics import accuracy_score
# ### Test npys
path1 = "C:/Users/shant/Desktop/PICT x MIT/WASSA 2022/Ensamble/npys_of_ensemble_models/"
test_BERTBase_OUS_400 = np.load(path1 + 'test_BERTBase-OUS_400.npy')
test_ElectraBase_Augmented_Data_4528_maxlen = np.load(path1 + 'test_ElectraBase-Augmented_Data_4528_maxlen.npy')
test_ElectraBase_Augmented_Data_random_balanced_647 = np.load(path1 + 'test_ElectraBase-Augmented_Data_random_balanced_647.npy')
test_ElectraBase_OUS_400 = np.load(path1 + 'test_ElectraBase-OUS_400.npy')
test_ElectraBase_OUS_500 = np.load(path1 + 'test_ElectraBase-OUS-500.npy')
# ### Valid npys
path2 = "C:/Users/shant/Desktop/PICT x MIT/WASSA 2022/Ensamble/npys_of_ensemble_models/"
val_BERTBase_OUS_400 = np.load(path2 + 'val_BERTBase-OUS_400.npy')
val_ElectraBase_Augmented_Data_4528_maxlen = np.load(path2 + 'val_ElectraBase-Augmented_Data_4528_maxlen.npy')
val_ElectraBase_Augmented_Data_random_balanced_647 = np.load(path2 + 'val_ElectraBase-Augmented_Data_random_balanced_647.npy')
val_ElectraBase_OUS_400 = np.load(path2 + 'val_ElectraBase-OUS_400.npy')
val_ElectraBase_OUS_500 = np.load(path2 + 'val_ElectraBase-OUS-500.npy')
# ### Functions to convert emotions to indices and indices to emotions
# +
e2i = {
"anger": 0,
"disgust": 1,
"fear": 2,
"joy": 3,
"neutral": 4,
"sadness": 5,
"surprise": 6
}
i2e = {v: k for k, v in e2i.items()}
def index_to_emotion(arr):
i2e = {0: 'anger', 1: 'disgust', 2: 'fear', 3: 'joy', 4: 'neutral', 5: 'sadness', 6: 'surprise'}
new_arr = []
for i in range(len(arr)):
new_arr.append(i2e[arr[i]])
return new_arr
def emotion_to_index(arr):
e2i = {'anger': 0, 'disgust': 1, 'fear': 2, 'joy': 3, 'neutral': 4, 'sadness': 5, 'surprise': 6}
new_arr = []
for i in range(len(arr)):
new_arr.append(e2i[arr[i]])
return new_arr
# -
# ### y_true
y_true = pd.read_csv('messages_dev_features_ready_for_WS_2022.tsv', sep = '\t')['emotion'].tolist()
# ### Ensembling Function
# 
def get_outputs(models, model_scores, model_weights, len_predict):
y_pred = []
best_f1 = []
for m in models:
best_f1.append((model_scores[m],m))
best_model = max(best_f1)[-1] # Choosing model with highest f1 score (in case of clashes vote of this mode li prefered)
for i in range(len_predict):
a = [(models[model][i],model) for model in models]
d = dict() # contains weighted votes {label: vote_counts}
for j in a:
if j[0] not in d:
d[j[0]] = model_weights[j[1]]
else:
d[j[0]] += model_weights[j[1]]
votes = [(d[k],k) for k in d]
votes.sort()
if votes[-1][0] == 1: # choosing the label with max votes
y_pred.append(models[best_model][i])
else: # if max votes = 1, choose the prediction of the best_model
y_pred.append(votes[-1][-1])
return y_pred
# ### Models {name: npys}
# +
final_ensemble_models_val = {'val_BERTBase_OUS_400' : val_BERTBase_OUS_400,
'val_ElectraBase_Augmented_Data_4528_maxlen' : val_ElectraBase_Augmented_Data_4528_maxlen,
'val_ElectraBase_Augmented_Data_random_balanced_647' : val_ElectraBase_Augmented_Data_random_balanced_647,
'val_ElectraBase_OUS_400' : val_ElectraBase_OUS_400}
final_ensemble_models_test = {'test_BERTBase_OUS_400' : test_BERTBase_OUS_400,
'test_ElectraBase_Augmented_Data_4528_maxlen' : test_ElectraBase_Augmented_Data_4528_maxlen,
'test_ElectraBase_Augmented_Data_random_balanced_647' : test_ElectraBase_Augmented_Data_random_balanced_647,
'test_ElectraBase_OUS_400' : test_ElectraBase_OUS_400}
# -
# ### Model f1 scores {name: f1 score}
# +
final_ensemble_scores_val = {
'val_BERTBase_OUS_400': 0.591857543223047,
'val_ElectraBase_Augmented_Data_4528_maxlen': 0.5894212727171738,
'val_ElectraBase_Augmented_Data_random_balanced_647': 0.5906288969076003,
'val_ElectraBase_OUS_400': 0.5967307817451174,
}
final_ensemble_scores_test = {
'test_BERTBase_OUS_400': 0.591857543223047,
'test_ElectraBase_Augmented_Data_4528_maxlen': 0.5894212727171738,
'test_ElectraBase_Augmented_Data_random_balanced_647': 0.5906288969076003,
'test_ElectraBase_OUS_400': 0.5967307817451174,
}
# -
# ### Model weights {name: npys}
# +
model_weights_val = {
'val_BERTBase_OUS_400': 1,
'val_ElectraBase_Augmented_Data_4528_maxlen': 1,
'val_ElectraBase_Augmented_Data_random_balanced_647': 1,
'val_ElectraBase_OUS_400': 1,
}
model_weights_test = {
'test_BERTBase_OUS_400': 1,
'test_ElectraBase_Augmented_Data_4528_maxlen': 1,
'test_ElectraBase_Augmented_Data_random_balanced_647': 1,
'test_ElectraBase_OUS_400': 1,
}
# -
# ### Prediction on Validation dataset, calculation of f1 score
y_pred_val = get_outputs(final_ensemble_models_val, final_ensemble_scores_val, model_weights_val, len(y_true))
print(f1_score(emotion_to_index(y_true), y_pred_val, average='macro'))
model_outputs_dev = pd.read_csv(r'C:\Users\shant\Desktop\PICT x MIT\WASSA 2022\Paper Publication functions\model_outputs_dev.csv') #.to_dict()
model_outputs_dev['Ensemble_w1_pred'] = index_to_emotion(y_pred_val)
model_outputs_dev['Ensemble_w1_true'] = y_true
model_outputs_dev = model_outputs_dev[['essay', 'BERTBase_OUS_400_pred', 'BERTBase_OUS_400_true',
'ElectraBase_Augmented_Data_4528_maxlen_pred',
'ElectraBase_Augmented_Data_4528_maxlen_true',
'ElectraBase_Augmented_Data_random_balanced_647_pred',
'ElectraBase_Augmented_Data_random_balanced_647_true',
'ElectraBase_OUS_400_pred', 'ElectraBase_OUS_400_true',
'Ensemble_w1_true', 'Ensemble_w1_pred']]
model_outputs_dev.to_csv(r'C:\Users\shant\Desktop\PICT x MIT\WASSA 2022\Paper Publication functions\model_outputs_dev.csv', index = False)
# ### y_pred test
y_pred_test = get_outputs(final_ensemble_models_test, final_ensemble_scores_test, model_weights_test, len(test_BERTBase_OUS_400))
y_pred_emotions = index_to_emotion(y_pred_test)
pd.DataFrame(y_pred_emotions).to_csv('predictions_EMO.tsv', index = False, sep = '\t', header = None)
| ensemble.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: mmlab
# language: python
# name: mmlab
# ---
from mmdet.apis import init_detector, inference_detector, show_result_pyplot
import mmcv
config_file = 'configs/vfnet/vf_101.py'
# download the checkpoint from model zoo and put it in `checkpoints/`
# url: https://download.openmmlab.com/mmdetection/v2.0/faster_rcnn/faster_rcnn_r50_fpn_1x_coco/faster_rcnn_r50_fpn_1x_coco_20200130-047c8118.pth
checkpoint_file = 'work_dirs/1_vf_101/latest.pth'
# build the model from a config file and a checkpoint file
model = init_detector(config_file, checkpoint_file, device='cuda:0')
# test a single image
img = 'data/coco/test/test10004.jpg'
result = inference_detector(model, img)
# show the results
show_result_pyplot(model, img, result)
# !python label_visualization.py --datasets COCO --img_path data/coco/train/ --label_path data/coco/annotations/coco1.json --cls_list_file ./xf.names
# !conda list
# !python tools/analysis_tools/analyze_logs.py plot_curve work_dirs/cascade_rcnn_r50_fpn_1x_coco/20210707_202839.log.json --keys s0.loss_cls s0.loss_bbox --out losses.pdf
# !python tools/analysis_tools/analyze_logs.py plot_curve work_dirs/1_vf/20210710_031414.log.json --keys <KEY> --out map11vfn.jpg
python tools/train.py configs/cascade_rcnn/cascade_rcnn_r50_fpn_1x_coco.py --work-dir ./work_dirs/2
python infer.py
python DataTransformer.py
python tools/train.py configs/dcn/cascade_rcnn_r50_fpn_dconv_c3-c5_1x_coco.py --work-dir ./work_dirs/4_dcn
anchor:0.38, 0.52, 0.75, 0.92, 1.28, 1.69
python tools/train.py configs/cascade_rcnn/cascade_rcnn_x101_32x4d_fpn_1x_coco.py --work-dir ./work_dirs/3
./tools/dist_train.sh configs/dcn/cascade_rcnn_r50_fpn_dconv_c3-c5_1x_coco.py 2 --work-dir ./work_dirs/6_dcn_OHEM
# +
export CUDA_VISIBLE_DEVICES=0,1
python tools/train.py configs/vfnet/cossume_vfnet.py --work-dir ./work_dirs/2_vf
python tools/train.py work_dirs/cbnet.py --work-dir ./work_dirs/1_cbnet
python tools/train.py configs/vfnet/my_vfnet.py --work-dir ./work_dirs/vf_1603 --seed 2021 --deterministic
python tools/train.py work_dirs/cascade_mix/cascade_rcnn_ohem.py
python tools/train.py work_dirs/vf_R2101-41-swa.py
# +
bash ./tools/dist_train.sh \
configs/vfnet/DRS_vfnet.py \
2 \
--seed 2021 \
--deterministic
cascade_rcnn_r50_rfp_1x_coco.py
bash ./tools/dist_train.sh \
configs/detectors/detectors_cascade.py \
2 \
--seed 2021 \
--deterministic
faster_rcnn_r50_fpn_ohem_1x_coco
bash ./tools/dist_train.sh \
configs/cascade_rcnn/cascade_rcnn_ohem.py \
2 \
--seed 2021 \
--deterministic
# +
python tools/analysis_tools/analyze_logs.py plot_curve work_dirs/1_vf/20210710_031414.log.json \
--keys bbox_mAP \
--out map11vfn.jpg
python tools/test.py \
work_dirs/detectors_cascade_rcnn_r50_1x_coco/detectors_cascade.py \
work_dirs/detectors_cascade_rcnn_r50_1x_coco/epoch_18.pth \
--format-only \
--options "jsonfile_prefix=./results"
python tools/analysis_tools/coco_error_analysis.py \
results.bbox.json \
vision_results \
--ann=data/coco/annotations/coco_val.json
python tools/test.py \
work_dirs/vf_R2101.py \
work_dirs/2_vf_R2101/latest.pth \
--format-only \
--options "jsonfile_prefix=./vfnet_test_R2101"
python tools/misc/browse_dataset.py configs/vfnet/vf_101.py \
--output-dir vis_test
bash ./tools/dist_train.sh \
configs/vfnet/vf_101.py \
2
bash ./tools/dist_train.sh \
work_dirs/vf_R2101.py \
2
# -
python swa.py --model_dir work_dirs/3_vf_R2101_swa --starting_model_id 1 --ending_model_id 12 --save_dir swa_model
python swa.py --model_dir work_dirs/vfX_swa --save_dir work_dirs/swa_model
# !nvidia-smi
coscmd upload mmdetection mmlab/
# rm -rf mmdetection/data
# +
python tools/test.py \
work_dirs/vf_R2101-41.py \
work_dirs/swa_model/swa_1-12.pth \
--format-only \
--options "jsonfile_prefix=./vfnet_swa_R2"
configs/hrnet/cascade_rcnn_hrnetv2p_w18_20e_coco.py
python tools/train.py work_dirs/vf_R2101-41-swa.py
python tools/train.py work_dirs/cascade_rcnn_hrnetv2p_w18_20e_coco.py
python tools/train.py configs/yolox/yolox_s.py
python tools/train.py work_dirs/vf_R2101-41-mosaic.py
python tools/train.py work_dirs/vf_mosaic.py
# -
# echo $http_proxy
# echo $https_proxy
git config --global http.proxy http://10.174.148.31:3928
git config --global https.proxy https://10.174.148.31:3928
| .ipynb_checkpoints/run-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Mukilan-Krishnakumar/NLP_With_Disaster_Tweets/blob/main/NLP_with_Disaster_Tweets_Part_1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="8A5LGczx6X-W"
# Welcome to NLP with Disaster Tweets. We are going to create a baseline NLP model with TensorFlow, this model is similar to the model which I learnt from the [Coursera Course on Natural Language Processing](https://www.coursera.org/learn/natural-language-processing-tensorflow?specialization=tensorflow-in-practice&utm_source=gg&utm_medium=sem&utm_campaign=33-DeepLearningAI-TensorFlow-IN&utm_content=B2C&campaignid=12462557662&adgroupid=120411989496&device=c&keyword=&matchtype=&network=g&devicemodel=&adpostion=&creativeid=510017701427&hide_mobile_promo&gclid=Cj0KCQiA2ZCOBhDiARIsAMRfv9J7y-xaQNirNg9EReDTgRS6rxEsTUV3U7qwa8TiGi2rZk_grBWsjgwaAujhEALw_wcB) taught by [<NAME>](https://www.linkedin.com/in/laurence-moroney/).
#
# I am going to implement the model and improve upon it. This is part 1 of the NLP with Disaster Tweets series and we will gradually improve the model.
#
# 📌 **Note** : As we are trying to get to the juice of model building this part doesn't cover EDA. EDA will be done in the subsequent parts.
#
# Let's get started.
# + [markdown] id="0-dX5W_170Bm"
# ## Downloading Dataset From Kaggle
# + [markdown] id="CbLhTG-W76Zt"
# To download dataset directly from kaggle we need to install kaggle in this machine. We also need to download a file called **kaggle.json**. This can be downloaded from Your Account -> Account -> API -> Generate Token.
#
# We need to upload this file to our colab runtime. Keep in mind that if you are using normal Colab, an uploaded file would be recycled.
#
# We create a folder called kaggle and copy our json file into that folder.
#
# We run `chmod 600` which means only the owner of the file has full read and write acces to it.
#
# We can download the kaggle dataset using `kaggle competitions download nlp-getting-started`.
#
# 📌 **Note** : If you didn't click **Join Competition** in kaggle, you won't be able to download the dataset.
#
# 😂 I did make that mistake so please be careful.
#
# + colab={"base_uri": "https://localhost:8080/"} id="3VPKO7d9itTr" outputId="e1443ed9-905f-47ae-e573-42b7cf1f3d30"
# ! pip install kaggle
# + id="99qONCYUjIKU"
# ! mkdir ~/.kaggle
# + id="9R5m-RHnjIDj"
# ! cp kaggle.json ~/.kaggle/
# + id="ucVtrT5jjH6t"
# ! chmod 600 ~/.kaggle/kaggle.json
# + colab={"base_uri": "https://localhost:8080/"} id="C2Id-Rj_jPFt" outputId="bec70903-6cb6-4009-ae5d-570f6448359f"
# ! kaggle competitions download nlp-getting-started
# + colab={"base_uri": "https://localhost:8080/"} id="RVITaNFMj7Zm" outputId="05501874-92f0-4ce4-b73b-02bf4e4bf9fb"
# ! unzip nlp-getting-started.zip
# + [markdown] id="NpUsuupykryg"
# ## Importing Necessary Modules
#
# We are going to import few necessary python modules to create our model.
#
# We will be importing the following modules:
# * Pandas - For data manupalation and analysis
# * Numpy - For array manipulation
# * Matplotlib.pyplot - For plotting graphs and visualizing data
# * Seaborn - For high level visualization
# * Re - For using Regular Expresions (RegEx)
# * TensorFlow - For building our Neural Network
#
# We will also import `Tokenizer` and `pad_sequences`. As the official documentation states:
# 1. `Tokenizer` : This class allows to vectorize a text corpus, by turning each text into either a sequence of integers (each integer being the index of a token in a dictionary) or into a vector where the coefficient for each token could be binary, based on word count, based on tf-idf.
# 2. `pad_sequences` : This function transforms a list (of length num_samples) of sequences (lists of integers) into a 2D Numpy array of shape (num_samples, num_timesteps). **It essentially adds padding to sentences to make them of equal length.**
#
# + id="YKw2Jn_mkJCe"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import re
import tensorflow as tf
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
# + [markdown] id="aVUKUIJhAH4s"
# We need to convert our csv file into a **Pandas DataFrame**.
#
# After converting we see the first 5 rows.
#
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="iCb7wMoAlvdw" outputId="8e9a4bb7-ee8d-4fdd-86db-28661a576320"
df = pd.read_csv('/content/train.csv')
df_test = pd.read_csv('/content/train.csv')
df.head()
# + colab={"base_uri": "https://localhost:8080/"} id="NUPKmFDoAYOc" outputId="2c75babd-89dc-4fdd-8e06-cf0060f18b08"
df['text']
# + [markdown] id="LdjUsxawAU12"
# ## Cleaning Data
#
# If we scroll a bit to the right in the `df['text']`, we can see many **URLS and Uppercased Words**. URLs are meaningless to our model and using Uppercase words bring redundancy in our word index.
#
# We remove these both using a custom function called cleaningText which removes URLs and lowercases all sentences.
# + id="Kholq_TTs_pX"
def cleaningText(df):
'''
This function gets a dataframe object as an input and removes the URLs from text column and makes every sentence lowercase.
'''
df['text'] = [re.sub(r'http\S+', '', x, flags=re.MULTILINE) for x in df['text']]
df['text'] = df['text'].str.lower()
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="mhIC9kxaoTn9" outputId="b882b610-3145-468f-fd52-17d5633ebd34"
cleaningText(df)
df.head()
# + [markdown] id="le_BWy7eBPqP"
# After running our custom function, we can store the text and label into individual lists.
# + colab={"base_uri": "https://localhost:8080/"} id="p2Kz9Q9Bl8Ga" outputId="ea05f89b-a026-4518-d837-8bded068ccfb"
sentences = [x for x in df['text']]
labels = [x for x in df['target']]
print(sentences)
# + [markdown] id="7Lqkwuo_BYRB"
# We make sure our labels are numerical values and are stored in Numpy arrays by using `np.array`.
#
# We split the data into training and testing data based on 80/20 rule.
#
# We have about 8000 records, we take the first 6090 to be training and the rest to be testing.
# + id="BOAbTEnEw91J"
labels = np.array(labels)
training_sentences = sentences[:6090]
training_labels = labels[:6090]
testing_sentences = sentences[6090:]
testing_labels = labels[6090:]
# + [markdown] id="aYMYnxWSvqTz"
# ## Model Parameters
#
# We need to specify a few things before we build our very own NLP model.
#
# We need to set up `vocab_size`, this is the maximum number of words we can store in our very own dictionary of sorts. We set it to `10000`.
#
# We need to set up `embedding_dim`, embedding is a relatively low-dimensional space into which you can translate high-dimensional vectors. Embeddings make it easier to do machine learning on large inputs like sparse vectors representing words. We set it to `16`.
#
# A tweet can be `280` characheters long, so we will set `max_length` to be `280`.
#
# We do padding on the end, in computer lingo this is called `post-padding`. We will set up `trunc-type` to `post`.
#
# If our model is faced with a new word it has not seen before, it will categorize it to `Out-Of-Vocabulary`, so we will set up `oov_tok` to be `<OOV>`.
#
# What we are going to do is convert all the words in our sentences into a dictionary of sorts (word_index) which allots individual tokens to each words.
#
# Our ML model can never work on text data, so we use this tokenizing mechanism to convert our sentences into sequences, they are numerical representation of our sentences. We pad them to make all the sequences be of same length.
#
# We do the same for testing sequences and labels.
#
# + id="p8xhP7O0on_0"
vocab_size = 10000
embedding_dim = 16
max_length = 280
trunc_type='post'
oov_tok = "<OOV>"
tokenizer = Tokenizer(num_words = vocab_size, oov_token=oov_tok)
tokenizer.fit_on_texts(training_sentences)
word_index = tokenizer.word_index
sequences = tokenizer.texts_to_sequences(training_sentences)
padded = pad_sequences(sequences,maxlen=max_length, truncating=trunc_type)
testing_sequences = tokenizer.texts_to_sequences(testing_sentences)
testing_padded = pad_sequences(testing_sequences,maxlen=max_length)
# + [markdown] id="tpZF6jWjFlnF"
# If you are curious about our word_index we can print them.
# + colab={"base_uri": "https://localhost:8080/"} id="LYWbbUjSyAN6" outputId="4503983e-2547-43d4-b574-29201593b550"
word_index
# + [markdown] id="HwYYWnR9F2wQ"
# ## Model Building
#
# Finally, we getting to juice of this tutorial. We are building our very own ML model.
#
# We will be building a Sequential model.
#
# We use the following layers:
# * Embedding layer - Turns positive integers (indexes) into dense vectors of fixed size. This basically converts our sequences into vectors.
# * GloabalAveragePooling1D - Global average pooling operation for temporal data. It basically computes the maximum of imput channels, finds the most relevant information.
# * Dense layers - One used with activation `relu` for achieving lower loss and another with `sigmoid` for classifying our tweet into either 1 (Disaster) or 0 (Not a Disaster).
#
# We will compile our model with `binary_crossentropy` as our loss because we only have binary classes (1 and 0).
#
# We will use `Adam` optimizer along with `accuracy` as metrics.
#
# We can visualize the layers of our model with `model.summary()`.
# + id="yEsUkx3EyvFm" colab={"base_uri": "https://localhost:8080/"} outputId="b51f1c65-b592-4beb-ee76-37ceb671b83b"
model = tf.keras.Sequential([
tf.keras.layers.Embedding(vocab_size,embedding_dim,input_length = max_length),
tf.keras.layers.GlobalAveragePooling1D(),
tf.keras.layers.Dense(6, activation = "relu"),
tf.keras.layers.Dense(1, activation = "sigmoid")
])
model.compile(loss='binary_crossentropy',optimizer='adam',metrics=['accuracy'])
model.summary()
# + [markdown] id="eREH1U-dHsxs"
# We will make our model run 10 times (`epochs = 10`).
#
# We will fit our model on training data and labels, we will evaluate on testing data.
# + id="ZTvQowbUy8s6" colab={"base_uri": "https://localhost:8080/"} outputId="2adb8103-3acb-48b8-dfda-4bef43557547"
np.random.seed(42)
num_epochs = 10
model.fit( padded, training_labels,epochs = num_epochs, validation_data = (testing_padded, testing_labels))
# + [markdown] id="rACYuaLn06fc"
# 😂 Wow, my model is only able to get 84 % accuracy. This is much better than our model guessing, we can improve this score by doing EDA and building a better model. For now, this is good enough.
| NLP_with_Disaster_Tweets_Part_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="YVt7Z_hzRtIh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 510} outputId="282a9627-2c83-4490-b560-6dbf000fdec9"
from sklearn.svm import LinearSVC
from sklearn.pipeline import Pipeline
from sklearn import metrics
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfTransformer
from sklearn.datasets import fetch_20newsgroups
newsgroups_train = fetch_20newsgroups(subset='train')
newsgroups_test = fetch_20newsgroups(subset='test')
X_train = newsgroups_train.data
X_test = newsgroups_test.data
y_train = newsgroups_train.target
y_test = newsgroups_test.target
text_clf = Pipeline([('vect', CountVectorizer()),
('tfidf', TfidfTransformer()),
('clf', LinearSVC()),
])
text_clf.fit(X_train, y_train)
predicted = text_clf.predict(X_test)
print(metrics.classification_report(y_test, predicted))
| Text Classfication/SVM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Chaine d'information d'un système
# -
# ## Analyse fonctionnelle et structurelle
#
# <img src="https://ericecmorlaix.github.io/img/AnalyseSystemique-GlobaleInformation.svg" alt="Focus sur la chaine d'information">
#
# Dans un système pluritechnique, la chaine d'information permet :
# - d'**acquérir** des informations en provenance de lui même, d'autres systèmes connectés, et de son utilisateur par l'intermédiaire d'une interface homme/machine (IHM) ;
# - de les **traiter** pour contrôler l'action à réaliser sur la matière d'oeuvre ;
# - et de **communiquer** l'état du système à l'utilisateur ou à d'autres systèmes connectés.
#
# ### Exemple du Stepper :
#
# <img src="https://ericecmorlaix.github.io/img/Stepper-Situation.svg" alt="Mise en situation du mini-stepper">
#
#
# La chaine d'information du mini-stepper permet :
# - d'**acquérir** l'information du mouvement d'un step au passage d'un aimant de la pédale devant le capteur [ILS](https://fr.wikipedia.org/wiki/Interrupteur_reed) (Interrupteur à Lame Souple) ;
# - de **traiter** cette information en comptant la durée de l'exercice et le nombre de steps et en calculant le nombre de calories dépensées ;
# - de **communiquer** ces informations à l'utilisateur en les affichant sur l'écran LCD.
#
# <img src="https://ericecmorlaix.github.io/img/Stepper-Information-Complet.svg" alt="Mise en situation du mini-stepper">
#
# ## La fonction "Acquérir" :
#
# - L'acquisition de grandeurs physiques se fait par l'intermédiaires de **capteurs**. Un capteur est un composant qui convertit une grandeur physique en un signal exploitable par l'unité de traitement*.
# > Exemples de capteurs de grandeur physique :
# - sur le stepper : le capteur ILS ;
# - autres :
# - Capteur Infrarouge pour mesurer une température sans contact
# - Barrière optique pour détecter un passage ;
# - Capteur audio pour mesurer un niveu sonore...
#
# - L'acquisition des consignes de l'utilisateur se fait par l'intermédiaire d'une **interface homme/machine** ([IHM](https://www.lebigdata.fr/interface-homme-machine-tout-savoir-sur-les-ihm)) qui intègre des composants capables de convertir des informations humaines en un signal exploitable par l'unité de traitement*.
# > Exemples de composants d'IHM :
# - sur le stepper : Bouton poussoir ;
# - autres :
# - écran tactile...
#
# * **Un signal exploitable par l'unité de traitement** : très souvent ce signal sera de nature électrique, rendu compatible avec le microcontroleur utilisé pour le traitement de l'information...
# ## Typologie des signaux logique, analogique et numérique :
#
# <img src="https://www.lossendiere.com/wp-content/uploads/2016/08/types-informations.jpg" alt="Typologie des signaux logique, analogique et numérique">
#
# Source : [www.lossendiere.com](https://www.lossendiere.com/2016/08/08/acquisition-dune-information/)
#
# <!--
# <img src="https://www.lossendiere.com/wp-content/uploads/2016/08/types-informations.jpg" alt="Typologie des signaux logique, analogique et numérique">
#
#
# -->
#
# ### Ressource vidéo :
# %%HTML
<center>
<iframe width="560" height="315" src="https://www.youtube-nocookie.com/embed/2PckTQZTdBw?start=16" frameborder="0" allow="accelerometer; autoplay; encrypted-media; gyroscope; picture-in-picture" allowfullscreen></iframe>
</center>
# ## Numération et codage de l’information, changement de base
#
# ### Problématique :
#
# Dans notre exemple du stepper, l'information fournie à l'entrée de la chaine par le capteur [ILS](https://fr.wikipedia.org/wiki/Interrupteur_reed) est de type logique (Tout ou Rien [TOR](https://fr.wikipedia.org/wiki/Tout_ou_rien), $0$ ou $1$) or le système de numération adapté pour traiter ce genre d'information est le binaire (à base $2$).
#
# Par conntre, en sortie, l'information à afficher est destinée à l'utilisateur qui lui compte avec un système de numération décimal (à base $10$).
#
# Plus généralement, on le reverra en détail plus tard, lorsque la grandeur physique à acquérir sera de type analogique, il nous faudra la numériser pour la traiter avec un microcontrôleur.
#
# On peut donc d'ores et déjà affirmer que les données qui circuleront dans la chaine d'information seront, à un endroit ou à un autre, numériques (ou pour le moins logique) même si elles représentent autre chose.
#
# Aussi, pour bien comprendre le codage de l'information qui circulent nous devons être capable de convertir une donnée numérique d'un système de numération à l'autre...
# ### Les bases :
#
# #### Système décimal :
#
# C’est le système de numération que nous utilisons tous les jours. C’est un système de base $10$ car il utilise dix symboles différents :
# $$ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9$$
#
# C’est un système positionnel car l’endroit où se trouve le symbole dans le nombre définit sa valeur. Le $2$ du nombre décimal $2356$ n’a pas la même valeur celui du nombre décimal $5623$ :
#
# $2356 = 2 \times 10^3 + 3 \times 10^2 + 5 \times 10^1 + 6 \times 10^0$ ici le $2$ vaut $2000$
#
# $5623 = 5 \times 10^3 + 6 \times 10^2 + 2 \times 10^1 + 3 \times 10^0$ ici le $2$ vaut $20$
# #### Système binaire :
#
# C’est le système de numération utilisé par les « machines numériques ». C’est un système de base $2$ car il utilise deux symboles différents :
# $$ 0, 1$$
#
# Pour distinguer le nombre binaire 10110 du nombre décimal 10110 on indique le code ``0b`` (ou le symbole ``%``) avant le nombre ou l’indice $_2$ (ou $_b$) après le nombre.
#
# $10110_2$ = $1 \times 2^4 + 0 \times 2^3 + 1 \times 2^2 + 1 \times 2^1 + 0 \times 2^0$
#
# $10110_2$ = $1 \times 16 + 0 \times 8 + 1 \times 4 + 1 \times 2 + 0 \times 1$
#
# $10110_2$ = $16 + 4 + 2 + 1$ =$22$
#
# On appel bit (contraction de Binary digIT = BIT) chacun des chiffres d’un nombre binaire.
#
# Avec un bit on peut distinguer deux états d’une information avec les deux nombres soit $1$, soit $0$.
#
# Avec $2$ bits $4= 2^2$
#
# Avec $3$ bits $8 = 2^3$
#
# Avec $8$ bits $256 = 2^8$
#
# Avec n bits on peut former $2^n$ combinaisons (nombres) différentes.
#
# Une suite de quatre bits est un "quartet" , une suite de huit bits est un "octet"
#
# Une suite de 16, 32, 64 bits est un "mot binaire"
# ### Système hexadécimal.
#
# C’est un système de base $16$ qui utilise donc seize symboles différents :
#
# $$ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, A, B, C, D, E, F$$
#
# Pour distinguer un nombre hexadécimal on indique le code ``0x`` (ou le symbole ``$``) avant le nombre ou l’indice $_{16}$ (ou $_h$) après le nombre.
#
# Les lettres A à F correspondent respectivement au nombre décimaux ? $10, 11, 12, 13, 14, 15$
#
# $AC53_{16}$ = $10 \times 16^3 + 12 \times 16^2 + 5 \times 16^1 + 3 \times 16^0$
#
# On utilise le système hexadécimal pour présenter de façon plus condensée un message binaire constitué d'une suite d'octect.
#
# ### Correspondance entre nombres de différentes bases :
# <!--
# <img src="https://ericecmorlaix.github.io/img/Stepper-Correspondance-Partiel.svg" alt="Tableau de correspondance entre bases">
#
#
# | **Décimal** | **Binaire** | **Hexadécimal** | **Décimal** | **Binaire** | **Hexadécimal** |
# |:---------------:|:---------------:|:---------------:|:---------------:|:---------------:|:---------------:|
# |$0$ |$______$ |$____$ | $8$ |$______$ |$____$ |
# |$1$ |$______$ |$____$ | $9$ |$______$ |$____$ |
# |$2$ |$______$ |$____$ | $10$ |$______$ |$____$ |
# |$3$ |$______$ |$____$ | $11$ |$______$ |$____$ |
# |$4$ |$______$ |$____$ | $12$ |$______$ |$____$ |
# |$5$ |$______$ |$____$ | $13$ |$______$ |$____$ |
# |$6$ |$______$ |$____$ | $14$ |$______$ |$____$ |
# |$7$ |$______$ |$____$ | $15$ |$______$ |$____$ |
# -->
#
# <img src="https://ericecmorlaix.github.io/img/Stepper-Correspondance-Complet.svg" alt="Tableau de correspondance entre bases">
#
# ### Changement de base :
#
# #### Conversion d’un nombre décimal en un nombre d’une autre base
#
# - Une méthode de conversion consiste à décomposer le nombre décimal en une somme de puissances de deux.
#
# > Par exemple, pour la conversion : $91$ = $01011011_2$
# >
# > On peut écrire :
# >$$91 = 0 \times 2^7 + 1 \times 2^6 + 0 \times 2^5 + 1 \times 2^4 + 1 \times 2^3 + 0 \times 2^2 + 1 \times 2^1 + 1 \times 2^0$$
# >
# >
# >$$91 = 64 + 16 + 8 + 2 + 1$$
# >
# >En rangeant les puissances de deux dans un tableau, on obtient :
# >
# ><img src="https://ericecmorlaix.github.io/img/Stepper-ConversionBinaire1.svg" alt="Mise en situation du mini-stepper">
#
#
# - Une autre méthode de conversion consiste à diviser le nombre décimal à convertir par la base b et conserver le reste de la division. Le quotient obtenu est divisé par b et conserver le reste. Il faut répéter l’opération sur chaque quotient obtenu.
# > Par exemple, pour la conversion : $91$ = $01011011_2$
# >
# ><img src="https://ericecmorlaix.github.io/img/Stepper-ConversionBinaire2.svg" alt="Mise en situation du mini-stepper">
# >
# >Les restes successifs sont écrits, en commençant par le dernier, de la gauche vers la droite. Cette méthode est dite « Méthode de la division successives ».
#
# #### Conversion d’un nombre hexadécimal en binaire.
#
# Chaque symbole du nombre écrit dans le système hexadécimal est remplacé par son équivalent écrit dans le système binaire.
#
# Exemple : Convertir $ECA_{16}$ = ${1110_2\over E_{16}}{1100_2\over C_{16}}{1010_2\over A_{16}}$ = $1110 1100 1010_2$
#
#
# #### Conversion d’un nombre binaire en hexadécimal.
#
# C’est l’inverse de la précédente. Il faut donc regrouper les 1 et les 0 du nombre par 4 en commençant par la droite, puis chaque groupe est remplacé par le symbole hexadécimal correspondant.
#
# Exemple : Convertir $1100001101111_2$ = ${1_{16}\over 0001_2}{8_{16}\over 1000_2}{6_{16}\over 0110_2}{F_{16}\over 1111_2}$ = $186F_{16}$
#
# ## Exercices :
#
# Convertir $9F2_{16}$ en binaire.
#
# Convertir $001111110101_2$ en hexadécimal.
#
# Convertir en décimal les nombres binaires suivants : $10110_2$ ; $10001101_2$ ; $1111010111_2$
#
# Convertir en binaire les nombres décimaux suivants : $37$ ; $189$ ; $205$ ; $2313$.
#
# Convertir en décimal les nombres hexadécimaux suivants : $92_{16}$ ; $2C0_{16}$ ; $37FD_{16}$.
#
# Convertir en hexadécimal les nombres décimaux suivants : 75 ; 314 ; 25619.
#
# Quelle est l’étendue des nombres définis en hexadécimal sur 6 chiffres ?
#
# Exécuter les opérations $10111101_2 + 101111_2$, $1BF_{16} + A23_{16}$
#
#
#
# ## Vérification avec Python :
#
# La fonction `bin()` permet de convertir un nombre en binaire :
bin(91)
bin(0x5b)
# La fonction `hex()` permet de convertir un nombre en hexadécimal :
hex(91)
hex(0b1011011)
# La fonction `int()` permet de convertir un nombre en décimal :
int(0b1111010111)
int(0x5b)
# ## Humour :
# <img src="data:image/svg+xml;base64,PHN2ZyB4bWxucz0iaHR0cDovL3d3dy53My5vcmcvMjAwMC9zdmciIHhtbG5zOnhsaW5rPSJodHRwOi8vd3d3LnczLm9yZy8xOTk5L3hsaW5rIiB2ZXJzaW9uPSIxLjEiIHdpZHRoPSIxMDc2cHgiIGhlaWdodD0iNDE3cHgiIHZpZXdCb3g9Ii0wLjUgLTAuNSAxMDc2IDQxNyI+PGRlZnMvPjxnPjxnIHRyYW5zZm9ybT0idHJhbnNsYXRlKDIwMC41LDEzMi41KSI+PHN3aXRjaD48Zm9yZWlnbk9iamVjdCBzdHlsZT0ib3ZlcmZsb3c6dmlzaWJsZTsiIHBvaW50ZXItZXZlbnRzPSJhbGwiIHdpZHRoPSI3MDUiIGhlaWdodD0iMTc4IiByZXF1aXJlZEZlYXR1cmVzPSJodHRwOi8vd3d3LnczLm9yZy9UUi9TVkcxMS9mZWF0dXJlI0V4dGVuc2liaWxpdHkiPjxkaXYgeG1sbnM9Imh0dHA6Ly93d3cudzMub3JnLzE5OTkveGh0bWwiIHN0eWxlPSJkaXNwbGF5OiBpbmxpbmUtYmxvY2s7IGZvbnQtc2l6ZTogMTJweDsgZm9udC1mYW1pbHk6IEhlbHZldGljYTsgY29sb3I6IHJnYigwLCAwLCAwKTsgbGluZS1oZWlnaHQ6IDEuMjsgdmVydGljYWwtYWxpZ246IHRvcDsgd2lkdGg6IDcwN3B4OyB3aGl0ZS1zcGFjZTogbm93cmFwOyBvdmVyZmxvdy13cmFwOiBub3JtYWw7IHRleHQtYWxpZ246IGNlbnRlcjsiPjxkaXYgeG1sbnM9Imh0dHA6Ly93d3cudzMub3JnLzE5OTkveGh0bWwiIHN0eWxlPSJkaXNwbGF5OmlubGluZS1ibG9jazt0ZXh0LWFsaWduOmluaGVyaXQ7dGV4dC1kZWNvcmF0aW9uOmluaGVyaXQ7d2hpdGUtc3BhY2U6bm9ybWFsOyI+PHNwYW4gc3R5bGU9ImZvbnQtc2l6ZTogNDhweCI+PGZvbnQgZmFjZT0iQ29taWMgU2FucyBNUyIgc3R5bGU9ImZvbnQtc2l6ZTogNDhweCIgY29sb3I9IiMwMDAwOTkiPklsIHkgYSAxMCBjYXTDqWdvcmllcyBkJ2luZGl2aWR1cyw8YnIgLz5jZXV4IHF1aSBjb21wcmVubmVudCBsZSBiaW5haXJlPGJyIC8+ZXQgbGVzIGF1dHJlcy4uLjwvZm9udD48YnIgLz48L3NwYW4+PC9kaXY+PC9kaXY+PC9mb3JlaWduT2JqZWN0Pjx0ZXh0IHg9IjM1MyIgeT0iOTUiIGZpbGw9IiMwMDAwMDAiIHRleHQtYW5jaG9yPSJtaWRkbGUiIGZvbnQtc2l6ZT0iMTJweCIgZm9udC1mYW1pbHk9IkhlbHZldGljYSI+Jmx0O3NwYW4gc3R5bGU9ImZvbnQtc2l6ZTogNDhweCImZ3Q7Jmx0O2ZvbnQgZmFjZT0iQ29taWMgU2FucyBNUyIgc3R5bGU9ImZvbnQtc2l6ZTogNDhweCIgY29sb3I9IiMwMDAwOTkiJmd0O0lsIHkgYSAxMCBjYXTDqWdvcmllcyBkJ2luZGl2aWR1cywmbHQ7YnImZ3Q7Y2V1eCBxdWkgY29tcHJlbm5lbnQgbGUgYmluYWlyZSZsdDticiZndDtldCBsZXMgYXV0cmVzLi4uJmx0Oy9mb250Jmd0OyZsdDticiZndDsmbHQ7L3NwYW4mZ3Q7PC90ZXh0Pjwvc3dpdGNoPjwvZz48cGF0aCBkPSJNIDI2OS41IDEwNC41IEMgNTUuNSAxMDQuNSAyIDIwNyAxNzMuMiAyMjcuNSBDIDIgMjcyLjYgMTk0LjYgMzcxIDMzMy43IDMzMCBDIDQzMCA0MTIgNzUxIDQxMiA4NTggMzMwIEMgMTA3MiAzMzAgMTA3MiAyNDggOTM4LjI1IDIwNyBDIDEwNzIgMTI1IDg1OCA0MyA2NzAuNzUgODQgQyA1MzcgMjIuNSAzMjMgMjIuNSAyNjkuNSAxMDQuNSBaIiBmaWxsPSJub25lIiBzdHJva2U9IiMwMDAwMDAiIHN0cm9rZS13aWR0aD0iNCIgc3Ryb2tlLW1pdGVybGltaXQ9IjEwIiB0cmFuc2Zvcm09InRyYW5zbGF0ZSgyLDMpIiBvcGFjaXR5PSIwLjI1Ii8+PHBhdGggZD0iTSAyNjkuNSAxMDQuNSBDIDU1LjUgMTA0LjUgMiAyMDcgMTczLjIgMjI3LjUgQyAyIDI3Mi42IDE5NC42IDM3MSAzMzMuNyAzMzAgQyA0MzAgNDEyIDc1MSA0MTIgODU4IDMzMCBDIDEwNzIgMzMwIDEwNzIgMjQ4IDkzOC4yNSAyMDcgQyAxMDcyIDEyNSA4NTggNDMgNjcwLjc1IDg0IEMgNTM3IDIyLjUgMzIzIDIyLjUgMjY5LjUgMTA0LjUgWiIgZmlsbD0ibm9uZSIgc3Ryb2tlPSIjN2YwMGZmIiBzdHJva2Utd2lkdGg9IjQiIHN0cm9rZS1taXRlcmxpbWl0PSIxMCIgcG9pbnRlci1ldmVudHM9Im5vbmUiLz48L2c+PC9zdmc+" width=60% />
#
# <center><a href="https://www.lemonde.fr/blog/binaire/2014/02/10/les-blagues-sur-linformatique-episode-1/">Cliquez ici pour trouver une explication, si besoin..."</a></center>
#
#
# ## Références au programme :
#
# <style type="text/css">
# .tg {border-collapse:collapse;border-spacing:0;}
# .tg td{font-family:Arial, sans-serif;font-size:14px;padding:10px 5px;border-style:solid;border-width:1px;overflow:hidden;word-break:normal;border-color:black;}
# .tg th{font-family:Arial, sans-serif;font-size:14px;font-weight:normal;padding:10px 5px;border-style:solid;border-width:1px;overflow:hidden;word-break:normal;border-color:black;}
# .tg .tg-cv16{font-weight:bold;background-color:#dae8fc;border-color:inherit;text-align:center}
# .tg .tg-xldj{border-color:inherit;text-align:left}
# </style>
# <table class="tg">
# <tr>
# <th class="tg-cv16">Compétences développées</th>
# <th class="tg-cv16">Connaissances associées</th>
# </tr>
# <tr>
# <td class="tg-xldj">Caractériser les échanges d’informations</td>
# <td class="tg-xldj">Natures et caractéristiques des signaux, des
# données, des supports de communication...</td>
# </tr>
# </table>
# + [markdown] slideshow={"slide_type": "-"}
# <a rel="license" href="http://creativecommons.org/licenses/by-sa/4.0/"><img alt="Licence Creative Commons" style="border-width:0" src="https://i.creativecommons.org/l/by-sa/4.0/88x31.png" /></a><br />Ce document est mis à disposition selon les termes de la <a rel="license" href="http://creativecommons.org/licenses/by-sa/4.0/">Licence Creative Commons Attribution - Partage dans les Mêmes Conditions 4.0 International</a>.
#
# Pour toute question, suggestion ou commentaire : <a href="mailto:<EMAIL>"><EMAIL></a>
| SI/ChaineInformation-Numeration_Complet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Amanpatni211/Deep-Learning-from-scratch/blob/main/103120080_ML_Task0_spider.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="JBUcBvtOXr1w"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# + colab={"base_uri": "https://localhost:8080/"} id="qxVdu089XzBN" outputId="20ccd6a7-e20d-4cde-ddc3-b365a638234b"
from google.colab import drive
drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/", "height": 490} id="G9YdrykjX4Gd" outputId="c2c65267-6122-4f59-8e31-46d8ac4689b3"
Dataset =pd.read_csv(r'/content/drive/MyDrive/student-course1.csv',sep=';')
Dataset.head(12)
# + colab={"base_uri": "https://localhost:8080/"} id="SQaIOfr-YPkt" outputId="052b10d7-bb95-425a-eeae-5e2eaaba494c"
obj_Dataset = Dataset.select_dtypes(include=['object']).copy() #columns having non-numerical datatype
print(obj_Dataset.head())
int_Dataset = Dataset.select_dtypes(include=['int64']).copy() #columns having numerical datatype
print(int_Dataset.head())
# + colab={"base_uri": "https://localhost:8080/"} id="rEWH7IUGZDLQ" outputId="ccaecc91-8292-4713-cf4c-3648472802f9"
#Converting object Data into suitable numerical values
hp = list(obj_Dataset.columns)
for i in range(len(hp)):
print(obj_Dataset[hp[i]].value_counts())
for t in range(len(hp)):
obj_Dataset[hp[t]] = obj_Dataset[hp[t]].astype('category')
obj_Dataset[hp[t]] = obj_Dataset[hp[t]].cat.codes
print(obj_Dataset.head())
for i in range(len(hp)):
print(obj_Dataset[hp[i]].value_counts())
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="HsfbjJg-jUyQ" outputId="25cece63-0c46-4faf-ab1a-228492e5035c"
obj_Dataset.head()
# + id="MAIthazJZmql" colab={"base_uri": "https://localhost:8080/", "height": 270} outputId="d7c6ee98-d2f3-4ba5-90a3-a29d974e17cc"
#Making them one again, here all data is numerical
frames = [obj_Dataset, int_Dataset]
final_data = pd.concat(frames,axis=1)
final_data.head()
# + id="bbzt68HYiOCu"
#spliting data in Train and Test
train=final_data.sample(frac=0.8,random_state=200) #random state is a seed value
test=final_data.drop(train.index)
# + id="ZI7qtv36ZyzE" colab={"base_uri": "https://localhost:8080/", "height": 300} outputId="cf5ebc8a-91b7-4589-949a-fdac594d33e6"
Y_trainA = (train.iloc[:, -3:])
Y_trainA.describe() #after running the cell,we realize that marks can range from 0 to 20 (only integers)
# + id="68PzwQw3l-Gy"
x_train = (train.drop(columns=['G1', 'G2', 'G3'], axis=0)).values
y_train1 = (train.iloc[:, -1:]).values
y_train2 = y_train1.T
x_val = (test.drop(columns=['G1', 'G2', 'G3'], axis=0)).values
y_val = (test.iloc[:, -1:]).values
# + id="1LV0LvvzR3LT"
y_train = y_train2.T
# + colab={"base_uri": "https://localhost:8080/", "height": 53} id="LBG3O1w2RzLF" outputId="8627fd9d-1403-4da9-b2c1-c3984f7b56d6"
#here we encode are labels to make the model understand what it means, so we have 21 zeros except at one place there is one ,
#which has index of its VALUE+1
'''a = np.array(y_train2)
y_train = np.zeros((a.size, a.max()+1))
y_train[np.arange(a.size),a] = 1
#example, '6' which is the marks in G1 of 4th instance will be representated as below
print(y_train[4])
print(y_train2[0][4])
'''
# + id="fia1Ew7bBOen"
x_train =x_train.reshape(316,1,30)
# + id="_Na2Qb_mY_JQ"
y_train = list(y_train)
# + colab={"base_uri": "https://localhost:8080/"} id="jQ1IMZ44TttH" outputId="ee962c16-79ab-45f9-fcf5-1d3aa9262f25"
y_train[77][0]
# + id="6GKOedktfQPS"
class Layer:
def __init__(self):
self.input = None
self.output = None
# computes the output Y of a layer for a given input X
def forward_propagation(self, input):
raise NotImplementedError
# computes dE/dX for a given dE/dY (and update parameters if any)
def backward_propagation(self, output_error, learning_rate):
raise NotImplementedError
# inherit from base class Layer
class FCLayer(Layer):
# input_size = number of input neurons
# output_size = number of output neurons
def __init__(self, input_size, output_size):
self.weights = np.random.rand(input_size, output_size) - 0.5
self.bias = np.random.rand(1, output_size) - 0.5
# returns output for a given input
def forward_propagation(self, input_data):
self.input = input_data
self.output = np.dot(self.input,self.weights) + self.bias
return self.output
# computes dE/dW, dE/dB for a given output_error=dE/dY. Returns input_error=dE/dX.
def backward_propagation(self, output_error, learning_rate):
input_error = np.dot(output_error, self.weights.T)
weights_error = np.dot(self.input.T,output_error)
# dBias = output_error
# update parameters
self.weights -= learning_rate * weights_error
self.bias = self.bias - learning_rate *output_error
return input_error
#loss function and its derivative
def mse(y_true, y_pred):
return(np.mean(np.power(y_true-y_pred, 2)));
def mse_prime(y_true, y_pred):
return(2*(y_pred-y_true)/y_true.size);
#activation function and derivative
def tanh(x):
return(np.tanh(x));
def tanh_prime(x):
return(1-np.tanh(x)**2);
'''
# Rectified Linear Unit (ReLU)
def ReLU(x):
return np.maximum(0,x)
# Derivative for ReLU
def ReLU_prime(x):
return 1 if x >0 else 0
'''
def ReLU(x):
y = np.copy(x)
y[y<0] = 0
return y
return relu
def ReLU_prime(x):
y = np.copy(x)
y[y>=0] = 1
y[y<0] = 0
return y
return relu_diff
def sigmoid_prime(x):
return (np.exp(-x))/((np.exp(-x)+1)**2)
def softmax(x):
exps = np.exp(x - x.max())
return exps / np.sum(exps, axis=0)
def softmax_prime(x):
exps = np.exp(x - x.max())
return exps / np.sum(exps, axis=0) * (1 - exps / np.sum(exps, axis=0))
#inherit from base class
class ActivationLayer(Layer):
def __init__(self, activation, activation_prime):
self.activation = activation
self.activation_prime = activation_prime
#return the activation input
def forward_propagation(self, input_data):
self.input = input_data
self.output = self.activation(self.input)
return(self.output)
#return input_error = dE/dX for a given output_error=dE/dY
def backward_propagation(self, output_error, learning_rate):
return(self.activation_prime(self.input) * output_error)
class Network:
def __init__(self):
self.layers = []
self.loss = None
self.loss_prime = None
# add layer to network
def add(self, layer):
self.layers.append(layer)
# set loss to use
def use(self, loss, loss_prime):
self.loss = loss
self.loss_prime = loss_prime
# predict output for given input
def predict(self, input_data):
# sample dimension first
samples = len(input_data)
result = []
# run network over all samples
for i in range(samples):
# forward propagation
output = input_data[i]
for layer in self.layers:
output = layer.forward_propagation(output)
result.append(output)
print(result)
return result
# train the network
def fit(self, x_train, y_train, epochs, learning_rate):
# sample dimension first
samples = len(x_train)
#saving epoch and error in list
epoch_list = []
error_list = []
# training loop
for i in range(epochs):
err = 0
for j in range(samples):
# forward propagation
output = x_train[j]
for layer in self.layers:
output = layer.forward_propagation(output)
# compute loss (for display purpose only)
err += self.loss(y_train[j][0], output)
# backward propagation
error = self.loss_prime(y_train[j][0], output)
'''
print("%%%%%%%%%%%%%") #this is the error it shows , but why is it not leanrning then?
print(error)
print("@@@$")'''
for layer in reversed(self.layers):
error = layer.backward_propagation(error, learning_rate)
# calculate average error on all samples
err /= samples
print('epoch %d/%d error=%f' % (i+1, epochs, err))
epoch_list.append(i+1)
error_list.append(err)
#creating dataframe of epoch and error
df = pd.DataFrame()
df['epoch'] = epoch_list
df['loss'] = error_list
return df
#Network
net = Network()
net.add(FCLayer(30,10))
net.add(ActivationLayer(ReLU, ReLU_prime))
net.add(FCLayer(10, 9))
net.add(ActivationLayer(ReLU, ReLU_prime))
net.use(mse, mse_prime)
net.add(FCLayer(9, 9))
net.add(ActivationLayer(ReLU, ReLU_prime))
net.use(mse, mse_prime)
net.add(FCLayer(9, 5))
net.add(ActivationLayer(ReLU, ReLU_prime))
net.use(mse, mse_prime)
net.add(FCLayer(5, 1))
net.add(ActivationLayer(ReLU, ReLU_prime))
net.use(mse, mse_prime)
df = net.fit(x_train, y_train,epochs=1,learning_rate=0.1)
import plotly.express as px
fig = px.line(df, x='epoch', y='loss',title='Change in loss with respect to Epochs')
fig.show()
# + colab={"base_uri": "https://localhost:8080/"} id="SR1ANfrCr_G_" outputId="24c56e9b-6b6d-4826-c430-c755497c55cf"
out = net.predict(x_val[:1])
print(len(x_val))
# + colab={"base_uri": "https://localhost:8080/", "height": 554} id="GehPmXNYxBzq" outputId="995e8d0f-66ab-43ae-ffaf-56a4ed198ac0"
user_data = []
print("Try your own example")
aman = list(Dataset.columns)
aman=aman[:-3]
for i in range(len(aman)):
b= int(input(aman[i]))
user_data.append(b)
print(user_data)
prediction_on_test_data= net.predict(user_data)
# + id="Mi1IYuqzLr0U"
# DONE-----------------X----------------
| 103120080_ML_Task0_spider.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import sys
def readIn(file, actual):
try:
replay = open(file)
except:
print("file not found")
sys.exit()
lines = replay.readlines()
times = []
for i, line in enumerate(lines):
if i >= 40:
break;
timeRaw = line.split("real")[0].strip()
times.append(float(timeRaw)/actual)
return times
times1 = readIn("exp2/replayTimeSet1.txt", 12.512803)
print(times1)
print()
times2 = readIn("exp3/replayTimeSet1.txt", 17.894)
print(times2)
print()
times3 = readIn("exp3/replayTimeSet2.txt", 9.022)
print(times3)
print()
plt.axis([0, 40, 1, 3])
plt.plot(times1, 'ro', times2, 'bs')
| data_set/plotDiffApp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from PIL import Image
import ia870 as MT
a = MT.iatext('0123456789')
g = MT.iabshow(a,a)
g.shape, g.dtype, g.min(), g.max()
Image.fromarray(g.astype('uint8')*255)
# ## Caso de matriz espaçamento fixo 5x7
# +
txt = '0123'
FontDft = np.array([[0,0,1,1,1,0,0], # 0
[0,1,0,0,1,1,0],
[0,1,0,0,1,1,0],
[0,1,0,1,0,1,0],
[0,1,1,0,0,1,0],
[0,1,1,0,0,1,0],
[0,0,1,1,1,0,0],
[0,0,0,1,0,0,0], # 1
[0,0,1,1,0,0,0],
[0,0,0,1,0,0,0],
[0,0,0,1,0,0,0],
[0,0,0,1,0,0,0],
[0,0,0,1,0,0,0],
[0,0,1,1,1,0,0],
[0,0,1,1,1,0,0], # 2
[0,1,0,0,0,1,0],
[0,0,0,0,0,1,0],
[0,0,1,1,1,0,0],
[0,1,0,0,0,0,0],
[0,1,0,0,0,0,0],
[0,1,1,1,1,1,0],
[0,0,1,1,1,0,0], # 3
[0,1,0,0,0,1,0],
[0,0,0,0,1,0,0],
[0,0,0,1,1,0,0],
[0,0,0,0,0,1,0],
[0,1,0,0,0,1,0],
[0,0,1,1,1,0,0],
],'bool')
FIRST_CHAR =ord('0')
LAST_CHAR = ord('3')
N_CHARS = 1 + LAST_CHAR - FIRST_CHAR
WIDTH_DFT = 7
HEIGHT_DFT = 7
FontDft = np.reshape(FontDft,(HEIGHT_DFT * N_CHARS, WIDTH_DFT))
y = ()
for c in txt:
i = ord(c) - FIRST_CHAR
assert i < N_CHARS,'iatext, code not allowed (%s)' % c
if len(y) == 0:
y = FontDft[i*HEIGHT_DFT:(i+1)*HEIGHT_DFT,:]
else:
y = MT.iaconcat('w',y,FontDft[i*HEIGHT_DFT:(i+1)*HEIGHT_DFT,:])
y.shape
# -
Image.fromarray(MT.iabshow(y,y).astype('uint8')*255)
| dtext.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Reminder
#
#
# <a href="#/slide-1-0" class="navigate-right" style="background-color:blue;color:white;padding:10px;margin:2px;font-weight:bold;">Continue with the lesson</a>
#
# <font size="+1">
#
# By continuing with this lesson you are granting your permission to take part in this research study for the Hour of Cyberinfrastructure: Developing Cyber Literacy for GIScience project. In this study, you will be learning about cyberinfrastructure and related concepts using a web-based platform that will take approximately one hour per lesson. Participation in this study is voluntary.
#
# Participants in this research must be 18 years or older. If you are under the age of 18 then please exit this webpage or navigate to another website such as the Hour of Code at https://hourofcode.com, which is designed for K-12 students.
#
# If you are not interested in participating please exit the browser or navigate to this website: http://www.umn.edu. Your participation is voluntary and you are free to stop the lesson at any time.
#
# For the full description please navigate to this website: <a href="../../gateway-lesson/gateway/gateway-1.ipynb">Gateway Lesson Research Study Permission</a>.
#
# </font>
# + hide_input=true init_cell=true slideshow={"slide_type": "skip"} tags=["Hide"]
# This code cell starts the necessary setup for Hour of CI lesson notebooks.
# First, it enables users to hide and unhide code by producing a 'Toggle raw code' button below.
# Second, it imports the hourofci package, which is necessary for lessons and interactive Jupyter Widgets.
# Third, it helps hide/control other aspects of Jupyter Notebooks to improve the user experience
# This is an initialization cell
# It is not displayed because the Slide Type is 'Skip'
from IPython.display import HTML, IFrame, Javascript, display
from ipywidgets import interactive
import ipywidgets as widgets
from ipywidgets import Layout
import getpass # This library allows us to get the username (User agent string)
# import package for hourofci project
import sys
sys.path.append('../../supplementary') # relative path (may change depending on the location of the lesson notebook)
import hourofci
# Retreive the user agent string, it will be passed to the hourofci submit button
agent_js = """
IPython.notebook.kernel.execute("user_agent = " + "'" + navigator.userAgent + "'");
"""
Javascript(agent_js)
# load javascript to initialize/hide cells, get user agent string, and hide output indicator
# hide code by introducing a toggle button "Toggle raw code"
HTML('''
<script type="text/javascript" src=\"../../supplementary/js/custom.js\"></script>
<input id="toggle_code" type="button" value="Toggle raw code">
''')
# + [markdown] slideshow={"slide_type": "slide"}
# # Types of Computational Systems in Cyberinfrastructure
#
# In this section we will look "under the hood" and cover different types of computational systems that are commonly used in cyberinfrastructure.
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## GPUs
#
# <img src="supplementary/gpu.png" width="400"/>
#
# <small>CC BY 4.0 https://commons.wikimedia.org/wiki/File:NvidiaTesla.jpg</small>
#
# - GPUs – Graphical Processing Units – are a very powerful type of processor (currently being used to display this text on your computer screen)
# - GPUs make up a very large part of the computational power of many computing systems
# - GPUs were originally developed for rendering graphical images, but it turns out that they are very fast for some (but not all) kinds of mathematically-oriented calculations.
#
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Quantum Computers
#
# - Quantum computers are the newest new thing
# - Normal computers work on a very simple principle: you do the same calculation over and over, you get the same results
# - Quantum computers are different
#
# - Rather than operating with a string of things called “bits” each of which is a 0 or a 1 like a current digital computer, Quantum computers operate on things called Qbits that are either 0 or 1 with a certain probability.
# - So when you run a program with a quantum computer, you don’t get an answer. You get a probability distribution of answers
# - Quantum computers are very important for some kinds of challenges but it will be a long time before they matter much to people using GIS applications!
# + [markdown] slideshow={"slide_type": "slide"}
# ### High Throughput Computing (HTC) Systems
#
# - HTC systems have been around for a long time
# - Sometimes certain data analysis problems involved doing lots of analysis (or lots of computations) that can happen pretty much independently
# - So a lot of work is done, and then the results are collected
#
#
# - This is different than the kind of jobs you usually run in parallel on a supercomputer
# - A bit of a good way to think about high performance parallel computing as opposed to high throughput computing is this:
# - If you care about how long one job takes, you’re probably doing high performance parallel computing
# - If you care about how many thousand jobs you run per month, you’re probably doing high throughput computing
# + [markdown] slideshow={"slide_type": "slide"}
# ## Let's take a closer look at High Throughput Computing Systems in GIS (at Clemson University)
#
# <img src="supplementary/htcs.png" width="400"/>
#
# - HTC systems used to analyze GIS data
# - An example problem: calculate the AADT (Annual Average Daily Traffic) through Greenville South Carolina
# - **Problem:** calculate all possible intersects between analyzed traffic routes (1.9 million observations) and all the traffic data collection sites that are spread throughout the city of Greenville
# - The Clemson University HTC system used a very famous and important piece of HTC software called Condor
#
#
# - Read more about the HTC system at Clemson at - https://www.clemsongis.org/high-throughput-computing-for-gis
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### How High Throughput Computing was used
#
# <img src="supplementary/htc.png" width="400"/>
#
# And you can see how this is cyberinfrastructure: lots of data, lots of data storage, broken up and sent across a network to lots of different computers organized into something called a “Condor Pool.” A “Condor Pool” is what a group of computational systems is called within the Condor High Throughput Computing software system.
#
# These images also from https://www.clemsongis.org/high-throughput-computing-for-gis
# + [markdown] slideshow={"slide_type": "slide"}
# ### Another example: The Large Hadron Collider (LHC)
#
# <img src="supplementary/hadron.png" width="400"/>
#
# - The LHC is the single biggest physics experiment in the world
# - It produces lots of small-ish blocks of data
# - The data tell what happened when subatomic particles are smashed together
# - Most of the time nothing new happens
# - So the data analysis task is to look at a whole bunch of data and determine if anything novel has happened
# - PERFECT for HTC – which is a very important kind of cyberinfrastructure
#
# From: https://home.cern/science/accelerators/large-hadron-collider
# + [markdown] slideshow={"slide_type": "slide"}
# <img src="supplementary/congratulations.png" width="400"/>
#
# ## You just learned a lot more detail about what computational systems can be part of cyberinfrastructure systems
#
# Really, anything that can connect to a digital network and can either produce data or do calculations can be considered cyberinfrastructure if it is put to work as part of “infrastructure for knowledge”
#
# <a href="cyberinfrastructure-5.ipynb">Click here to move on to the next segment where you will learn more about the importance of CI in scientific discovery</a>
| beginner-lessons/cyberinfrastructure/cyberinfrastructure-4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (py2)
# language: python
# name: py2
# ---
# +
import igraph
import copy
import networkx as nx
import numpy as np
import pandas as pd
from matplotlib import pylab as plt
import os
import glob
from helpers import *
# %matplotlib inline
#why need download?
#3m resolution
#1. copublish use Ox' license
#2. slider
#recurring problem - needs to look at construction sites
#
# Green info - little engine for
# can we find another source?
#
# +
""" get all files """
files = glob.glob('./matrix_csvs/*')
g_MW_files = sorted([f for f in files if ((f.split('/')[-1].split('_')[1]=='green')and (f.split('/')[-1].split('_')[2]=='MW.csv') ) ])
b_MW_files =sorted([f for f in files if ((f.split('/')[-1].split('_')[1]=='blue') and (f.split('/')[-1].split('_')[2]=='MW.csv') )])
all_MW_files = sorted([f for f in files if ((f.split('/')[-1].split('_')[1]=='all') and (f.split('/')[-1].split('_')[2]=='MW.csv'))])
print all_MW_files
# +
""" get a dict going with files and data """
data = {}
years = range(2007,2018)
years_m = [2007,2008,2009,2010,2012,2013,2014,2015,2016,2017]
for y in years_m:
fname_all = [f for f in all_MW_files if ((str(y)+'.75') in f)][0]
fname_g = [f for f in g_MW_files if ((str(y)+'.75') in f)][0]
fname_b = [f for f in b_MW_files if ((str(y)+'.75') in f)][0]
data[y]={'year':y,
'fname_all':fname_all,
'data_all':0.0,
'fname_g':fname_g,
'data_g':0.0,
'fname_b':fname_b,
'data_b':0.0}
data[2011]={'data_b': 0.0,
'data_g': 0.0,
'year': 2011,
'fname_g': './matrix_csvs/2011.5_green_MW.csv',
'fname_b': './matrix_csvs/2011.5_blue_MW.csv',
'data_all': 0.0,
'fname_all': './matrix_csvs/2011.5_all_MW.csv'}
print data[2016]['data_all']
print data
# -
""" read in all the data """
for y in years:
print y
data[y]['data_all']=pd.read_csv(data[y]['fname_all'], encoding='utf-8').set_index('COMPANY')
data[y]['data_g']=pd.read_csv(data[y]['fname_g'], encoding='utf-8').set_index('COMPANY')
data[y]['data_b']=pd.read_csv(data[y]['fname_b'], encoding='utf-8').set_index('COMPANY')
data[y]['data_ff'] = data[y]['data_all'].subtract(data[y]['data_g'], fill_value=0.0).subtract(data[y]['data_b'], fill_value=0.0)
""" read in the countries """
country_df = pd.read_csv('country_iso_regions.csv', encoding='utf-8').set_index('country')
def softmax(x):
return np.exp(x) / np.sum(np.exp(x), axis=0)
def degree(x):
#print x
#print 'softmax: ', softmax(x)
#raw_input('-->')
return np.sum(softmax(x)/np.max(x))/x.shape[0]
# +
#print data[2016]['data_ff']
regions = country_df.SIPS_REGION.unique()
print list(country_df)
print country_df.SIPS_REGION.unique()
sample_ff = data[2016]['data_ff'].T.merge(country_df[['SIPS_REGION','iso2']], left_index=True, right_on='iso2').groupby('SIPS_REGION').sum().T
sample_g = data[2016]['data_g'].T.merge(country_df[['SIPS_REGION','iso2']], left_index=True, right_on='iso2').groupby('SIPS_REGION').sum().T
sample_b = data[2016]['data_b'].T.merge(country_df[['SIPS_REGION','iso2']], left_index=True, right_on='iso2').groupby('SIPS_REGION').sum().T
sample_all = data[2016]['data_all'].T.merge(country_df[['SIPS_REGION','iso2']], left_index=True, right_on='iso2').groupby('SIPS_REGION').sum().T
print sample_ff.shape
print sample_g[sample_all['AFRICA']>0][['AFRICA']]/sample_all[sample_all['AFRICA']>0][['AFRICA']]
#print sample_b.shape
#print sample_all.shape
#print sample_all[sample_all['AFRICA']>0].shape
#print sample_g[sample_all['AFRICA']>0].shape
#print sample_g.T.sum()/sample_all.T.sum()
print clamp(16)
# -
def str_col(g,b):
return "#{0:02x}{1:02x}{2:02x}".format(clamp(0), clamp(int(g)), clamp(int(b)))
# +
""" let's plot em all """
import matplotlib.patches as mpatches
import matplotlib.lines as mlines
f, axarr = plt.subplots(1, figsize=(16,9))
axarr.set_xlim(2006.3,2018)
axarr.set_xticks(years)
for y in years:
print y
sample_ff = data[y]['data_ff'].T.sum().T
#print sample_ff
sample_g = data[y]['data_g'].T.sum().T
sample_b = data[y]['data_b'].T.sum().T
sample_all = data[y]['data_all'].T.sum().T
#"#{0:02x}{1:02x}{2:02x}".format(clamp(0), clamp(g), clamp(b))
pos = pd.DataFrame(sample_all, columns=['all'])
#print list(pos)
#g = sample_g[sample_all[r]>0][[r]]/sample_all[sample_all[r]>0][[r]]*255.0
#b = sample_b[sample_all[r]>0][[r]]/sample_all[sample_all[r]>0][[r]]*255.0
pos = pos.sort_values('all',ascending=True)/1000.0
pos['cumsum'] = pos.cumsum()
pos['bin']=(pos['cumsum']/pos['cumsum'].max()*100)#.astype(int)
print pos
pos = pos.dropna()
pos['bin'] = pos['bin'].astype(int)
#pos = pos.sort_values('cumsum', ascending=False)
pos['g'] = sample_g[sample_all>0]/sample_all[sample_all>0]*255.0
pos['b'] = sample_b[sample_all>0]/sample_all[sample_all>0]*255.0
pos['color'] = pos.apply(lambda row: str_col(row['g'], row['b']), axis=1)
#print pos
down_sample = pos.groupby('bin').sum()
down_sample['count'] = pos.groupby('bin').count()['all']
down_sample.g = down_sample.g/down_sample['count']
down_sample.b = down_sample.b/down_sample['count']
down_sample['cumsum']=down_sample['all'].cumsum()
down_sample['color']=down_sample.apply(lambda row: str_col(row['g'], row['b']), axis=1)
down_sample['log10count'] = np.log10(down_sample['count'])+np.log10(2.0)
print down_sample
axarr.axvline(x=y, color='k', linestyle='--')
"""
axarr.barh(pos['cumsum'].values-pos[r].values, #position
(1.0-pos['cumsum'].values/(pos['cumsum'].max())*0.9), #width
pos[r].values, #height
y,
align='edge',
color=pos.color.values)
"""
#print down_sample['log10count'].max()
axarr.barh(down_sample['cumsum'].values-down_sample['all'].values, #position
down_sample['log10count']/4.5, #down_sample['log10count'].max(),#(1.0-down_sample['cumsum'].values/(down_sample['cumsum'].max())*0.9), #width
down_sample['all'].values, #height
y,
align='edge',
color=down_sample.color.values)
#<NAME>
#raw_input('-->')
#print pos
axarr.set_ylabel('Generating Capacity [GW]')
axarr.set_ylim(bottom=0.0)
bottom,top = axarr.get_ylim()
axarr.arrow(2007, top, .9, 0, head_width=(top-bottom)*0.03, head_length=0.1, fc='k', ec='k', width=(top-bottom)*0.01)
axarr.text(2006.95, top+(top-bottom)*0.02,'$log_{10}(2*N_{Companies}$)')
green_patch = mpatches.Patch(color='green', label='green data')
blue_patch = mpatches.Patch(color='blue', label='blue data')
black_patch = mpatches.Patch(color='black', label='black data')
f.legend((black_patch,blue_patch,green_patch), ('Fossil Fuels','Nuclear & Large-Scale Hydro','Renewables'), loc='center', bbox_to_anchor=(0.5,0.92), ncol=3)
f.suptitle('Cumulative Generating Capacity: Global', fontsize=20)
f.savefig('./figures/1_'+'global'+'.png')
plt.show()
# +
""" let's plot em all """
import matplotlib.patches as mpatches
import matplotlib.lines as mlines
for r in regions:
print r
f, axarr = plt.subplots(1, figsize=(16,9))
axarr.set_xlim(2006.3,2018)
axarr.set_xticks(years)
for y in years:
print y
sample_ff = data[y]['data_ff'].T.merge(country_df[['SIPS_REGION','iso2']], left_index=True, right_on='iso2').groupby('SIPS_REGION').sum().T
sample_g = data[y]['data_g'].T.merge(country_df[['SIPS_REGION','iso2']], left_index=True, right_on='iso2').groupby('SIPS_REGION').sum().T
sample_b = data[y]['data_b'].T.merge(country_df[['SIPS_REGION','iso2']], left_index=True, right_on='iso2').groupby('SIPS_REGION').sum().T
sample_all = data[y]['data_all'].T.merge(country_df[['SIPS_REGION','iso2']], left_index=True, right_on='iso2').groupby('SIPS_REGION').sum().T
#"#{0:02x}{1:02x}{2:02x}".format(clamp(0), clamp(g), clamp(b))
pos = sample_all[sample_all[r]>0][[r]]
#g = sample_g[sample_all[r]>0][[r]]/sample_all[sample_all[r]>0][[r]]*255.0
#b = sample_b[sample_all[r]>0][[r]]/sample_all[sample_all[r]>0][[r]]*255.0
pos = pos.sort_values(r, ascending=True)/1000.0
pos['cumsum'] = pos[r].cumsum()
pos['bin']=(pos['cumsum']/pos['cumsum'].max()*100).astype(int)
#pos = pos.sort_values('cumsum', ascending=False)
pos['g'] = sample_g[sample_all[r]>0][[r]]/sample_all[sample_all[r]>0][[r]]*255.0
pos['b'] = sample_b[sample_all[r]>0][[r]]/sample_all[sample_all[r]>0][[r]]*255.0
pos['color'] = pos.apply(lambda row: str_col(row['g'], row['b']), axis=1)
#print pos
down_sample = pos.groupby('bin').sum()
down_sample['count'] = pos[['bin',r]].groupby('bin').count()
down_sample.g = down_sample.g/down_sample['count']
down_sample.b = down_sample.b/down_sample['count']
down_sample['cumsum']=down_sample[r].cumsum()
down_sample['color']=down_sample.apply(lambda row: str_col(row['g'], row['b']), axis=1)
down_sample['log10count'] = np.log10(down_sample['count'])+np.log10(2.0)
#print down_sample
axarr.axvline(x=y, color='k', linestyle='--')
"""
axarr.barh(pos['cumsum'].values-pos[r].values, #position
(1.0-pos['cumsum'].values/(pos['cumsum'].max())*0.9), #width
pos[r].values, #height
y,
align='edge',
color=pos.color.values)
"""
print down_sample['log10count'].max()
axarr.barh(down_sample['cumsum'].values-down_sample[r].values, #position
down_sample['log10count']/4.5, #down_sample['log10count'].max(),#(1.0-down_sample['cumsum'].values/(down_sample['cumsum'].max())*0.9), #width
down_sample[r].values, #height
y,
align='edge',
color=down_sample.color.values)
#<NAME>
#raw_input('-->')
#print pos
axarr.set_ylabel('Generating Capacity [GW]')
axarr.set_ylim(bottom=0.0)
bottom,top = axarr.get_ylim()
axarr.arrow(2007, top, .9, 0, head_width=(top-bottom)*0.03, head_length=0.1, fc='k', ec='k', width=(top-bottom)*0.01)
axarr.text(2006.95, top+(top-bottom)*0.02,'$log_{10}(2*N_{Companies}$)')
green_patch = mpatches.Patch(color='green', label='green data')
blue_patch = mpatches.Patch(color='blue', label='blue data')
black_patch = mpatches.Patch(color='black', label='black data')
f.legend((black_patch,blue_patch,green_patch), ('Fossil Fuels','Nuclear & Large-Scale Hydro','Renewables'), loc='center', bbox_to_anchor=(0.5,0.92), ncol=3)
f.suptitle('Cumulative Generating Capacity: '+str(r), fontsize=20)
f.savefig('./figures/1_'+str(r)+'.png')
plt.show()
# +
""" get dregree for all data"""
for y in years:
for data_str in ['data_all','data_g','data_b', 'data_ff']:
df = data[y][data_str]
#degree just with connections
df['degree'] = df[df>0].count(axis=1)
#degree with algo
#df['degree'] = df.apply(lambda row: degree(row/np.sum(row)), axis=1)
cols = [c for c in list(df) if len(c)<3]
df['sum'] = df[cols].sum(axis=1)
df['cumsum'] = df['sum'].cumsum()
data[y][data_str] = df.sort_values('sum', ascending=False)
# +
""" lets plot them degrees """
import matplotlib.patches as mpatches
import matplotlib.lines as mlines
f, axarr = plt.subplots(3,sharex=True, figsize=(16,9))
axarr[0].set_xlim(2006.3,2018)
axarr[0].set_xticks(years)
#axarr[0].set_xtickabels(years)
axarr[0].set_ylim(0,100)
axarr[1].set_ylim(0,100)
axarr[2].set_ylim(0,100)
axarr[0].arrow(2008, 100.0, .9, 0, head_width=5, head_length=0.1, fc='k', ec='k', width=2)
axarr[1].arrow(2008, 100.0, .9, 0, head_width=5, head_length=0.1, fc='k', ec='k', width=2)
axarr[2].arrow(2008, 100.0, .9, 0, head_width=5, head_length=0.1, fc='k', ec='k', width=2)
axarr[0].arrow(2006.8, 50.0, 0, -40, head_width=.1, head_length=10, fc='k', ec='k', width=.01)
axarr[1].arrow(2006.8, 50.0, 0, -40, head_width=.1, head_length=10, fc='k', ec='k', width=.01)
axarr[2].arrow(2006.8, 50.0, 0, -40, head_width=.1, head_length=10, fc='k', ec='k', width=.01)
axarr[0].arrow(2006.8, 50.0, 0, 40, head_width=.1, head_length=10, fc='k', ec='k', width=.01)
axarr[1].arrow(2006.8, 50.0, 0, 40, head_width=.1, head_length=10, fc='k', ec='k', width=.01)
axarr[2].arrow(2006.8, 50.0, 0, 40, head_width=.1, head_length=10, fc='k', ec='k', width=.01)
axarr[0].text(2006.7, 80,'small\ncompanies', ha='right', rotation=90)
axarr[1].text(2006.7, 80,'small\ncompanies', ha='right', rotation=90)
axarr[2].text(2006.7, 80,'small\ncompanies', ha='right', rotation=90)
axarr[0].text(2006.4, 30,'large\ncompanies', ha='left', rotation=90)
axarr[1].text(2006.4, 30,'large\ncompanies', ha='left', rotation=90)
axarr[2].text(2006.4, 30,'large\ncompanies', ha='left', rotation=90)
axarr[0].text(2007.95, 105,'d=1')
axarr[1].text(2007.95, 105,'d=1')
axarr[2].text(2007.95, 105,'d=1')
axarr[0].text(2008.95, 105,'d=5')
axarr[1].text(2008.95, 105,'d=5')
axarr[2].text(2008.95, 105,'d=5')
cols_dict={0:'black',1:'blue',2:'green'}
axarr[0].set_ylabel('%$N_{Companies, Fossil Fuel}$')
axarr[1].set_ylabel('%$N_{Companies, Nuclear & Hydro}$')
axarr[2].set_ylabel('%$N_{Companies, Renewables}$')
degree_means = []
for y in years:
print y
ii=0
for data_str in ['data_ff','data_b','data_g']:
df = data[y][data_str]
degree_means.append(df.degree.mean())
df = df.drop(df[df.degree<1.0].index)
#print df['sum'].sum(axis=0)
verts = np.arange(len(df))/float(len(df))*100
degrees = ((df.degree.rolling(50, min_periods=1).mean())-1.0)/4.0#-df.degree.min())*500
#print 'max degrees', degrees.max()*5
#print degrees
degrees_std = degrees+(df.degree.rolling(50, min_periods=1).std())/4.0#*500
#print 'max std', degrees_std.max()*5
degrees = degrees+y
degrees_std = degrees_std+y
#print degrees.values
#print verts
#print degrees
#print degrees_std
axarr[ii].plot(degrees.values,verts, color=cols_dict[ii])
axarr[ii].fill_betweenx(verts,y,degrees_std.values, alpha=0.3, color=cols_dict[ii])
#axarr[0].plot(degrees_std.values,verts)
#vert = np.arange(len(df))/df['sum'].sum(axis)
axarr[ii].axvline(x=y, color='k', linestyle='--')
#print data[y]['data']
#axarr[2].scatter(df_test['degree'],df_test['cumsum'])
#axarr[2].violinplot(df_test['degree'],positions = [1.0], widths=50, points=10000)
#axarr[2].set_ylabel('ALL_Cumulative_Sum')
#f.savefig(os.path.join('output',path['ID'],'_ndwi_stats.png'))
ii+=1
f.suptitle('Degree Distributions for all Companies', fontsize=20)
gray_patch = mpatches.Patch(color='gray', label='The red data')
black_line = mlines.Line2D([0],[0],color='black')
f.legend((black_line,gray_patch), ('$Rolling\_Mean_{window=50}$','$Rolling\_StdDev_{window=50}$'), loc='center', bbox_to_anchor=(0.5,0.92), ncol=2)
f.savefig('out.png')
plt.show()
print np.mean(degree_means)
# +
print data[2017]['data_all'].degree.rolling(50, min_periods=1).std().argmax()
print data[2017]['data_all'].degree.rolling(50, min_periods=1).std().to_string()
# -
print data[2017]['data_all']# - data[2017]['data_g'] - data[2017]['data_b']
# +
"""Let's do this a fan graph"""
df_all = pd.read_csv(all_MW_files[0], encoding='utf-8').set_index('COMPANY')
df_g = pd.read_csv(g_MW_files[0], encoding='utf-8').set_index('COMPANY')
df_test = pd.DataFrame(df_all.sum(axis=1).sort_values(ascending=False), columns=['sum'])
df_test['cumsum'] = df_test['sum'].cumsum()
#df_test['degree'] = df_all[df_all>0.0001].count(axis=1)
df_test['degree'] = df_all.apply(lambda row: degree(row/np.sum(row)), axis=1)
print df_test.sort_values('sum',ascending=False)
#df_test.plot('cumsum','degree')
#https://stackoverflow.com/questions/28807169/making-a-python-fan-chart-fan-plot
#https://matplotlib.org/examples/pylab_examples/boxplot_demo.html
f, axarr = plt.subplots(3, sharex=True, figsize = (16,9))
axarr[2].scatter(df_test['degree'],df_test['cumsum'])
#axarr[2].violinplot(df_test['degree'],positions = [1.0], widths=50, points=10000)
axarr[2].set_ylabel('ALL_Cumulative_Sum')
#f.savefig(os.path.join('output',path['ID'],'_ndwi_stats.png'))
plt.show()
plt.close()
plt.clf()
#print df_all.T.sum().cumsum()
#print df_g.T.sum()
#
print df_test.shape[1]
test = np.zeros((215))
test[5]=1
test[15]=1
test[24]=1
test[30]=1
print test
print 'test degree: ',degree(test.T/np.sum(test.T))
"algo degree"
#print df_all.apply(lambda row: degree(row/np.sum(row)), axis=1).plot()
#print df_g[df_g>0.0001].count(axis=1).plot()
#print df_all[df_all>0.0001].count(axis=1).plot()
#print df_all.apply() / df_all.T.max()
# -
print df_test
print df_test.degree.rolling(1000, min_periods=1).mean().plot()
print df_test.degree*100
print df_test.degree.rolling(1000, min_periods=1).std().plot()
#print len(df_test.degree.rolling(100, min_periods=1).mean().resample('1'))
print (full_d_range.days)
x_ticks = np.arange(0,full_d_range.days,365)
print (x_ticks.shape)
x_tick_labels = [str(i) for i in range(2008,2019)]
print (len(x_tick_labels))
f, axarr = plt.subplots(3, sharex=True, figsize = (16,9))
axarr[0].scatter(x, y_area, marker='+')
axarr[0].set_ylabel('Area [m^2]')
axarr[0].set_xlim(0,full_d_range.days)
axarr[0].set_xticks(x_ticks)
axarr[0].set_xticklabels(x_tick_labels)
axarr[0].set_title('Extents indicators for '+path['ID']+': '+path['name'])
axarr[1].scatter(x, y_exts, marker='+')
axarr[1].set_ylabel('Fraction')
axarr[1].set_ylim(bottom=0.0)
#print(y_vals)
for y in y_vals:
print (y.shape)
axarr[2].violinplot(y_vals,positions = x, widths=50, points=1000)
axarr[2].set_ylabel('values')
f.savefig(os.path.join('output',path['ID'],'_ndwi_stats.png'))
#plt.show()
#plt.close()
plt.clf()
path_to_matrices = "matrix_csvs/"
years = ["2004"] + ["2007"] + [str(i) for i in range(2014,2018)] #list of all years
quarters = [".0", ".25", ".5", ".75"] #list of all quarters
# +
#Stating period for analisis
y_index = -2
q_index = 3
current_year = years[y_index] + quarters[q_index] #The year and quarter we will do the analisis for
#Importing the files needed
df_all = pd.read_csv(path_to_matrices + current_year + "_all_projection.csv")
names = list(df_all.columns[1:]) #Getting names of nodes
n_countries = len(names)
ADJ_all = np.loadtxt(open(path_to_matrices + current_year + "_all_projection.csv", "rb"), delimiter=",", skiprows=1,
usecols=range(1,n_countries+1)) #getting the adjacency matrix
#files are empty
#pd.read_csv(path_to_matrices + current_year + "_blue_projection.csv")
#Adjmat_blue = np.loadtxt(open(path_to_matrices + current_year + "_blue_projection.csv", "rb"), delimiter=",", skiprows=1,
#usecols=range(1,n_countries+1)) #getting the adjacency matrix
#Adjmat_green = np.loadtxt(open(path_to_matrices + current_year + "_green_projection.csv", "rb"), delimiter=",", skiprows=1,
#usecols=range(1,n_countries+1)) #getting the adjacency matrix
# -
# Note, in year 2004 there is hardly little influence between countries. In year 2017 there is more, although still not a lot (think?)
print(np.diagonal(ADJ_all)) #Is this correct? A "1" in the diagonal would indicate the country is not influenced
#by any other country?
plt.imshow(ADJ_all) #Few influence between countries?
plt.colorbar()
#This is an isolated node.
ADJ_all[0]
#Only small countries can be influenced?
print("Small diagonal value")
for i in range(n_countries):
if np.diagonal(ADJ_all)[i] < 0.5:
print i, " ", names[i]
#PageRank centrality with igraph
G_all = igraph.Graph.Weighted_Adjacency(ADJ_all.tolist() ,mode="directed")
pr_all = G_all.personalized_pagerank(weights=G_all.es["weight"], directed=True)
names_copy = copy.copy(names)
inds = np.array(pr_all).argsort()[::-1][:]
sort_names_pr = np.array(names_copy)[inds]
sort_centrality_pr = np.array(pr_all)[inds]
sort_names_pr, sort_centrality_pr
plt.plot(pr_all, "o")
# ### Loading matrices and making projections
# +
#Extracting bipartite adj mat
ADJ_bip_green = np.loadtxt(open(path_to_matrices + current_year + "_green_MW.csv", "rb"), delimiter=",", skiprows=1,
usecols=range(1,n_countries+1)) #getting the adjacency matrix
ADJ_bip_blue = np.loadtxt(open(path_to_matrices + current_year + "_blue_MW.csv", "rb"), delimiter=",", skiprows=1,
usecols=range(1,n_countries+1)) #getting the adjacency matrix
#Does the all category include the green and blue or is it just the dirty ones? In principle we would like to have
#only the dirty ones, right?
ADJ_bip_all = np.loadtxt(open(path_to_matrices + current_year + "_all_MW.csv", "rb"), delimiter=",", skiprows=1,
usecols=range(1,n_countries+1))
#To get the high emission plants we use the all matrix and remove the "clean" part
ADJ_bip_dirty = ADJ_bip_all - (ADJ_bip_green + ADJ_bip_blue)
#Making the company projection
ADJ_comp_green = (ADJ_bip_green/ADJ_bip_green.sum()).dot(ADJ_bip_green.T/(ADJ_bip_green.T.sum()))
ADJ_comp_blue = (ADJ_bip_blue/ADJ_bip_blue.sum()).dot(ADJ_bip_blue.T/(ADJ_bip_blue.T.sum()))
ADJ_comp_all = (ADJ_bip_all/ADJ_bip_all.sum()).dot(ADJ_bip_all.T/(ADJ_bip_all.T.sum()))
ADJ_comp_dirty = (ADJ_bip_dirty /ADJ_bip_dirty .sum()).dot(ADJ_bip_dirty.T/(ADJ_bip_dirty .T.sum()))
#Making the country projection
ADJ_country_green = (ADJ_bip_green.T/ADJ_bip_green.T.sum()).dot(ADJ_bip_green/(ADJ_bip_green.sum()))
ADJ_country_blue = (ADJ_bip_blue.T/ADJ_bip_blue.T.sum()).dot(ADJ_bip_blue/(ADJ_bip_blue.sum()))
ADJ_country_all = (ADJ_bip_all.T/ADJ_bip_all.T.sum()).dot(ADJ_bip_all/(ADJ_bip_all.sum()))
ADJ_country_dirty = (ADJ_bip_dirty.T /ADJ_bip_dirty.T .sum()).dot(ADJ_bip_dirty/(ADJ_bip_dirty.sum()))
#Getting the percentage of emissions of each type for each country
green_part = [sum(ADJ_bip_green[:, i])/sum(ADJ_bip_all[:, i]) for i in range(n_countries)]
blue_part = [sum(ADJ_bip_blue[:, i])/sum(ADJ_bip_all[:, i]) for i in range(n_countries)]
dirty_part = [sum(ADJ_bip_dirty[:, i])/sum(ADJ_bip_all[:, i]) for i in range(n_countries)]
# -
# # Multilayer analysis
# +
def make_supra_adj(ADJ_list, COUP_list):
"""Function that takes list of adj matrix and coupling and generates the supra adjacency matrix of the
multiplex network
Args:
ADJ_list: list of numpy 2D array's. Each of them is an adjacency matrix
COUP_list: list of numpy 1D array's. Each of them is the vector to be used for coupling the matrices
Return:
Supra Adjacency matrix. Numpy 2D array.
"""
n_layers = len(ADJ_list)
n_nodes = ADJ_list[0].shape[0]
#stacking up adjacency matrix and coupling accordingly
sup_list = []
for i in range(n_layers):
row = []
for j in range(n_layers):
if i == j: #if in diagonal add adjacency matrix
row.append(ADJ_list[i])
#print(ADJ_list[i].shape)
else: #otherwise add the coupling (diagonal since multiplex network)
row.append(np.diag(COUP_list[i])) #if i or j here defined direction of coupling
#print(np.diag(COUP_list[i]).shape)
sup_list.append(row)
return np.bmat(sup_list)
n_nodes = n_countries
n_layers = 3
#The following function is taylored for igraph centrality
def flatten_centrality(centrality_vector, n=n_nodes, l=n_layers, names=names, hubauth=False):
'''Centrality networks in a Multiplex are commonly expressed as a vector of
nlx1 dimension. To have a centrality for each node it is important to flatten
the vector into nx1 dimension. Also, it is important return a list of ranked
nodes and another of their score.
Args
centrality_vector(numpy array): the centrality measure for each node-layer
n(int): number of nodes
l(int): number of layers
names(list of strings): name of nodes (countries)
hubauth(boolean): indicating if the centrality is hub or authority
Return:
sort_names_multi(list of strings): names of countries ordered by centrality
sort_centrality_multi(list of flots): sorted score of nodes
'''
multi_centrality = []
for i in range(n):
cent = 0
for k in range(l):
cent += centrality_vector[i + n*k]
if hubauth:
multi_centrality.append(cent[0])
else:
multi_centrality.append(cent)
node_names = np.array(copy.deepcopy(names))
inds = np.array(multi_centrality).argsort()[::-1][:]
sort_names_multi = node_names[inds]
sort_centrality_multi = np.array(multi_centrality)[inds]
return sort_names_multi, sort_centrality_multi
# -
#Making the supra adjacency matrix
SUPADJ_country = make_supra_adj([ADJ_country_green, ADJ_country_blue, ADJ_country_dirty], [green_part, blue_part, dirty_part])
#making the graph
G_supra = igraph.Graph.Weighted_Adjacency( SUPADJ_country.tolist() ,mode="directed")
#computing pagerank
pr_muliplex = G_supra.personalized_pagerank(weights=G_supra.es["weight"])
#sorting it appropriately
pr_multiplex_sorted = flatten_centrality(pr_muliplex)
pr_multiplex_sorted
# Comparing it with the aggregate pagerank centrality
G_all = igraph.Graph.Weighted_Adjacency( ADJ_all.tolist() ,mode="directed")
pr_all = G_all.personalized_pagerank(weights=G_all.es["weight"])
names_copy = copy.copy(names)
inds = np.array(pr_all).argsort()[:]
#If small countries are first, try including the [::-1 as below]
#inds = np.array(pr_all).argsort()[::-1][:]
sort_names_pr = np.array(names_copy)[inds]
sort_centrality_pr = np.array(pr_all)[inds]
sort_names_pr, sort_centrality_pr
# # Inconsistency between networkx and igraph?
# I tend to trust igraph more, however the discrepancy might mean that values in general are too close. Perhaps networkx power method is not converging?
# +
#PageRank centrality with igraph
G_all = igraph.Graph.Weighted_Adjacency(ADJ_all.tolist() ,mode="directed")
pr_all = G_all.personalized_pagerank(weights=G_all.es["weight"], directed=True)
names_copy = copy.copy(names)
#inds = np.array(pr_all).argsort()[::-1][:]
inds = np.array(pr_all).argsort()[:]
sort_names_pr = np.array(names_copy)[inds]
sort_centrality_pr = np.array(pr_all)[inds]
sort_names_pr, sort_centrality_pr
#PageRank centrality with networkx
G_all_nx = nx.from_numpy_matrix(ADJ_all)
pr_all_nx = nx.pagerank(G_all_nx, weight="weight")
plt.plot(pr_all_nx.values(), "o")
plt.plot(pr_all, ".")
# -
plt.plot(pr_all, pr_all_nx.values(), "o")
for i in range(n_countries):
print i, "nx = ", names[sorted(pr_all_nx.items(), key=lambda x:x[1])[::-1][i][0]], "ig = ", sort_names_pr[i]
#If we want to do Hubs and Auth the following code will be usefull
"""def G_list_hub(G_adj_list):
'''
Args(list): list of graphs adjacency matrices
Return(list): list of adjacency matrices to be used in the block diagonal
of the supra adjacency matrix for hub score
'''
G_hub = []
for g in G_adj_list:
new = np.dot(g, g.transpose())
G_hub.append(new)
return G_hub
def G_list_auth(G_adj_list):
'''
Args(list): list of graphs adjacency matrices
Return(list): list of adjacency matrices to be used in the block diagonal
of the supra adjacency matrix for auth score
'''
G_auth = []
for g in G_adj_list:
new = np.dot( g.transpose(), g)
G_auth.append(new)
return G_auth
def hub_auth_diag(Coup_list):
coup = []
for c in Coup_list:
coup.append(c*c)
return coup"""
pd.__version__
| Basic_Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/librairy/EQAKG/blob/main/test/MuHeQA_Evaluation.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="f0WscfmDXVls"
import pandas as pd
pd.set_option('max_colwidth', 400)
# -
# # VQuAnDa
# + colab={"base_uri": "https://localhost:8080/"} id="Cc8Q2argXdaX" outputId="deb8ecc8-2c74-414d-c345-451c078a6945"
# read csv file
results = pd.read_csv("https://raw.githubusercontent.com/librairy/EQAKG/main/test/VQuAnDa/results/VQuanda.csv", header = 0, encoding = "utf-8", sep=";")
# display DataFrame
print(results.info())
# + colab={"base_uri": "https://localhost:8080/", "height": 300} id="5ZiQG4kKc2bk" outputId="553cb044-8b0d-4d31-a69d-d8e0a575f99d"
results.describe()
# + colab={"base_uri": "https://localhost:8080/", "height": 632} id="j323YYC3emjT" outputId="4889b00d-65d2-4349-a8a8-cd7318c61a3f"
results[results['BLEU Score (SacreBleu)'] == 0.0]
# +
#preguntas cuya respuesta sea numerica
numbers = results[results['Answer'].str.isnumeric()]
numbers
# -
rightNumbers = numbers[numbers['BLEU Score (SacreBleu)'] != 0]
rightNumbers
# +
import requests
queryURL = "http://localhost:5000/eqakg/dbpedia/en?text=true"
# -
for counter,i in enumerate(rightNumbers['Question']):
if counter > 19:
break
files = {
'question': (None, i),
}
response = requests.get(queryURL, files = files)
print("Question:", i)
print("Answer before conversion: ", (response.json())['answer-2'])
print("Generated text: ", (response.json())['text'])
wrongNumbers = numbers[numbers['BLEU Score (SacreBleu)'] == 0]
wrongNumbers
for counter,i in enumerate(wrongNumbers['Question']):
if counter > 19:
break
files = {
'question': (None, i),
}
response = requests.get(queryURL, files = files)
print("Question:", i)
print("Answer before conversion: ", (response.json())['answer-2'])
print("Generated text: ", (response.json())['text'])
#preguntas cuya respuesta sea booleana
booleans = results[(results['Response'] == "true") | (results['Response'] == "false")]
booleans
booleans[(booleans['Answer'] == "yes")]
correctBooleans = booleans[(booleans['Answer'] == "yes") & (booleans['Response'] == "true")]
correctBooleans
for counter,i in enumerate(correctBooleans['Question']):
if counter > 19:
break
files = {
'question': (None, i),
}
response = requests.get(queryURL, files = files)
print("Question:", i)
print("Answer before conversion: ", (response.json())['answer-2'])
print("Generated text: ", (response.json())['text'])
correctBooleans = booleans[(booleans['Answer'] == "no") & (booleans['Response'] == "false")]
correctBooleans
booleans[(booleans['Answer'] == "no")]
# # VANiLLA
# +
# read csv file
results = pd.read_csv("https://raw.githubusercontent.com/librairy/EQAKG/main/test/VANiLLA/VANiLLA.csv", header = 0, encoding = "utf-8", sep=";")
# display DataFrame
print(results.info())
# -
# mean, average, std...
results.describe()
#preguntas cuyo BLEU Score, de SacreBleu, sea 0
results[results['BLEU Score (SacreBleu)'] == 0.0]
#preguntas cuyo BLEU Score, de ntlk, sea 0
results[results['BLEU Score (ntlk)'] == 0.0]
# +
#preguntas cuya respuesta sea numerica
numbers = results[results['Answer'].str.isnumeric()]
numbers
# -
numbers[numbers['BLEU Score (SacreBleu)'] != 0]
#preguntas cuya respuesta sea booleana
booleans = results[(results['Response'] == "true") | (results['Response'] == "false")]
booleans
for counter,i in enumerate(booleans['Question']):
if counter > 19:
break
files = {
'question': (None, i),
}
response = requests.get(queryURL, files = files)
print("Question:", i)
print("Answer before conversion: ", (response.json())['answer-2'])
print("Generated text: ", (response.json())['text'])
# # LC-Quad 2.0
# +
# read csv file
results = pd.read_csv("https://raw.githubusercontent.com/librairy/EQAKG/main/test/LC-QuAD_2.0/results/LC-Quad.csv", header = 0, encoding = "utf-8", sep=";")
# display DataFrame
print(results.info())
# -
# mean, average, std...
results.describe()
#preguntas cuyo BLEU Score, de SacreBleu, sea 0
results[results['BLEU Score (SacreBleu)'] == 0.0]
#preguntas cuyo BLEU Score, de ntlk, sea 0
results[results['BLEU Score (ntlk)'] == 0.0]
#preguntas cuya respuesta sea numerica
numbers = results[results['Answer'].str.isnumeric()]
numbers
correctNumbers = numbers[numbers['BLEU Score (SacreBleu)'] != 0]
correctNumbers
for counter,i in enumerate(correctNumbers['Question']):
if counter > 19:
break
files = {
'question': (None, i),
}
response = requests.get(queryURL, files = files)
print("Question:", i)
print("Answer before conversion: ", (response.json())['answer-2'])
print("Generated text: ", (response.json())['text'])
wrongNumbers = numbers[numbers['BLEU Score (SacreBleu)'] == 0]
wrongNumbers
for counter,i in enumerate(wrongNumbers['Question']):
if counter > 19:
break
files = {
'question': (None, i),
}
response = requests.get(queryURL, files = files)
print("Question:", i)
print("Answer before conversion: ", (response.json())['answer-2'])
print("Generated text: ", (response.json())['text'])
#preguntas cuya respuesta sea booleana
booleans = results[(results['Response'] == "true") | (results['Response'] == "false")]
booleans
correctBooleans = booleans[booleans['BLEU Score (SacreBleu)'] != 0]
correctBooleans
for counter,i in enumerate(correctBooleans['Question']):
if counter > 19:
break
files = {
'question': (None, i),
}
response = requests.get(queryURL, files = files)
print("Question:", i)
print("Answer before conversion: ", (response.json())['answer-2'])
print("Generated text: ", (response.json())['text'])
wrongBooleans = booleans[booleans['BLEU Score (SacreBleu)'] != 0]
wrongBooleans
for counter,i in enumerate(wrongBooleans['Question']):
if counter > 19:
break
files = {
'question': (None, i),
}
response = requests.get(queryURL, files = files)
print("Question:", i)
print("Answer before conversion: ", (response.json())['answer-2'])
print("Generated text: ", (response.json())['text'])
# + [markdown] id="vATtJdRkv8Dh"
# # Test Code
# + colab={"base_uri": "https://localhost:8080/"} id="oRrOVip3v7To" outputId="36e8be7a-eebe-43f1-c73d-01eec90c2697"
"""
!pip install spacy
!python -m spacy download en_core_web_sm
!pip install spacy-entity-linker
"""
# + id="DzIZ8AImx9gU"
"""
!python -m spacy_entity_linker "download_knowledge_base"
"""
# + colab={"base_uri": "https://localhost:8080/"} id="rBNRqZbuwJqs" outputId="16d6d316-8bc8-461a-e6f1-0d55387934c0"
"""
import spacy # version 3.0.6'
# initialize language model
nlp = spacy.load("en_core_web_sm")
# add pipeline (declared through entry_points in setup.py)
nlp.add_pipe("entityLinker", last=True)
doc = nlp("I watched the Pirates of the Caribbean last silvester")
# returns all entities in the whole document
all_linked_entities = doc._.linkedEntities
# iterates over sentences and prints linked entities
for sent in doc.sents:
sent._.linkedEntities.pretty_print()
"""
| test/setup/MuHeQA_Evaluation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: myenv
# language: python
# name: myenv
# ---
# # Getting Financial Data - Google Finance
# ### Introduction:
#
# This time you will get data from a website.
#
#
# ### Step 1. Import the necessary libraries
import pandas as pd
import datetime as dt
from pandas_datareader import data, wb
# ### Step 2. Create your time range (start and end variables). The start date should be 01/01/2015 and the end should today (whatever your today is)
start = dt.datetime(2015, 1, 1)
end = dt.datetime.today()
# ### Step 3. Select the Apple, Tesla, Twitter, IBM, LinkedIn stocks symbols and assign them to a variable called stocks
stocks = ['AAPL', 'TSLA', 'TWTR', 'IBM', 'LNKD']
# ### Step 4. Read the data from yahoo, assign to df and print it
df = data.DataReader(stocks, 'yahoo', start, end)
df
# ### Step 5. What is the type of structure of df ?
df.index.names
# ### Step 6. Print all the Items axis values
# #### To learn more about the Panel structure go to [documentation](http://pandas.pydata.org/pandas-docs/stable/dsintro.html#panel)
df.items
# ### Step 7. Good, now we know the data avaiable. Create a dataFrame called vol, with the Volume values.
vol = df['Volume']
vol.head()
# ### Step 8. Aggregate the data of Volume to weekly
# #### Hint: Be careful to not sum data from the same week of 2015 and other years.
vol_weekly = vol.resample('W').sum()
vol_weekly
# +
vol['week'] = vol.index.week
vol['year'] = vol.index.year
vol_weekly_2 = vol.groupby(['year', 'week']).sum()
# -
vol_weekly_2.head()
# ### Step 9. Find all the volume traded in the year of 2015
vol[vol['year'] == 2015].sum()
| 09_Time_Series/Getting_Financial_Data/Exercises.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
# # Fuzzy Matching of data
#
# ## A common scenario for data scientists is the marketing, operations or business groups give you two sets of similar data with different variables & asks the analytics team to normalize both data sets to have a common record for modelling.
#
#
#
# ### Here is an example of two similar data sets:
#
#
# 
#
#
#
# ### How would you as a data scientist match these two different but similar data sets to have a master record for modelling?
#
# ### Consider that the Sales Data has 10 million records and the # of Customer data has 50,000 records
#
# ### Possible combinations are 500 Billions
#
# ## So an algorithm that tries to match all combinations of the data sets is probably NOT a good way to go
#
# ## Let's see some simple example of how to do something similar in a better way
#
# ### Make sure you installed the necessary Python libraries
#
# ### In *command window*
# * **type pip install fuzzywuzzy**
# * **type pip install python-Levenshtein**
# +
from fuzzywuzzy import fuzz
from fuzzywuzzy import process
print('Running the method ratio()')
fuzz.ratio("ACME Factory", "ACME Factory Inc.")
# -
# ### We can see how the ratio() function is confused by the suffix “Inc.” used in company names, but really the two strings refer to the same entity.
#
# ### This is captured by the partial ratio.
print('Running the method partial_ratio()')
fuzz.partial_ratio("ACME Factory", "ACME Factory Inc.")
# ### More examples
print('Running token_sort_ratio method')
fuzz.token_sort_ratio('<NAME>', '<NAME>')
print('Running token_set_ratio method')
fuzz.token_set_ratio('<NAME>', '<NAME>')
# ### In case we have a list of options and we want to find the closest match(es), we can use the process module:
# +
query = '<NAME>'
choices = ['<NAME>', '<NAME>', 'B. Obama']
# Get a list of matches ordered by score, default limit to 5
process.extract(query, choices)
# [('<NAME>', 95), ('<NAME>', 95), ('B. Obama', 85)]
# If we want only the top one
process.extractOne(query, choices)
# ('<NAME>', 95)
# -
# ## So what are these different method doing and why are they producing somewhat different results
# ### Simple ratio
# **The ratio method compares the whole string and follows the standard Levenshtein distance similarity ratio between two strings**
#
# ### Partial ratio
# **The partial ratio method works on “optimal partial” logic. If the short string k and long string m are considered, the algorithm will score by matching the length of the k string**
#
# ### Token sort ratio
# **The token sort ratio method sorts the tokens alphabetically. Then, the simple ratio method is applied to output the matched percentage**
#
# ### Token set ratio
# **The token set ratio ignores the duplicate words. It is similar to the sort ratio method but more flexible. It basically extracts the common tokens and then applies fuzz.ratio() for comparisons**
# ## Other more sophisticated methods of finding text distance
#
# ## Levenshtein distance
# Levenshtein distance measures the minimum number of insertions, deletions, and substitutions required to change one string into another. This can be a useful measure to use if you think that the differences between two strings are equally likely to occur at any point in the strings. It’s also more useful if you do not suspect full words in the strings are rearranged from each other (see Jaccard similarity further down)
#
# ### pip install textdistance[extras]
import textdistance
# ## Levenshtein distance
# Levenshtein distance measures the minimum number of insertions, deletions, and substitutions required to change one string into another. This can be a useful measure to use if you think that the differences between two strings are equally likely to occur at any point in the strings. It’s also more useful if you do not suspect full words in the strings are rearranged from each other (see Jaccard similarity further down).
textdistance.levenshtein("this is my test", "that test of mine")
textdistance.levenshtein("test this of mine", "this test of mine")
# ## An implementation of Leveenshtein Distance and a test example
import numpy as np
def levenshtein_ratio_and_distance(s, t, ratio_calc = False):
""" levenshtein_ratio_and_distance:
Calculates levenshtein distance between two strings.
If ratio_calc = True, the function computes the
levenshtein distance ratio of similarity between two strings
For all i and j, distance[i,j] will contain the Levenshtein
distance between the first i characters of s and the
first j characters of t
"""
# Initialize matrix of zeros
rows = len(s)+1
cols = len(t)+1
distance = np.zeros((rows,cols),dtype = int)
# Populate matrix of zeros with the indeces of each character of both strings
for i in range(1, rows):
for k in range(1,cols):
distance[i][0] = i
distance[0][k] = k
# Iterate over the matrix to compute the cost of deletions,insertions and/or substitutions
for col in range(1, cols):
for row in range(1, rows):
if s[row-1] == t[col-1]:
cost = 0 # If the characters are the same in the two strings in a given position [i,j] then the cost is 0
else:
# In order to align the results with those of the Python Levenshtein package, if we choose to calculate the ratio
# the cost of a substitution is 2. If we calculate just distance, then the cost of a substitution is 1.
if ratio_calc == True:
cost = 2
else:
cost = 1
distance[row][col] = min(distance[row-1][col] + 1, # Cost of deletions
distance[row][col-1] + 1, # Cost of insertions
distance[row-1][col-1] + cost) # Cost of substitutions
if ratio_calc == True:
# Computation of the Levenshtein Distance Ratio
Ratio = ((len(s)+len(t)) - distance[row][col]) / (len(s)+len(t))
return Ratio
else:
# print(distance) # Uncomment if you want to see the matrix showing how the algorithm computes the cost of deletions,
# insertions and/or substitutions
# This is the minimum number of edits needed to convert string a to string b
return "The strings are {} edits away".format(distance[row][col])
# +
Str1 = "This is my test"
Str2 = "That test of mine"
Distance = levenshtein_ratio_and_distance(Str1,Str2)
print(Distance)
Ratio = levenshtein_ratio_and_distance(Str1,Str2,ratio_calc = True)
print(Ratio)
Str1 = "This is my test"
Str2 = "That test of mine"
# -
# ## Jaro-Winkler
# Jaro-Winkler is another similarity measure between two strings. This algorithm penalizes differences in strings more earlier in the string. A motivational idea behind using this algorithm is that typos are generally more likely to occur later in the string, rather than at the beginning. When comparing “this test” vs. “test this”, even though the strings contain the exact same words (just in different order), the similarity score is just 2/3.
#
# If it matters more that the beginning of two strings in your case are the same, then this could be a useful algorithm to try.
#
# Saying the same in a slightly different way still finds the similarity
textdistance.jaro_winkler("this is my test", "test this test of mine")
textdistance.jaro_winkler("this is my test", "this test of mine")
# ## Jaccard Similarity
# Jaccard similarity measures the shared characters between two strings, regardless of order. In the first example below, we see the first string, “this test”, has nine characters (including the space). The second string, “that test”, has an additional two characters that the first string does not (the “at” in “that”). This measure takes the number of shared characters (seven) divided by this total number of characters (9 + 2 = 11). Thus, 7 / 11 = .636363636363…
# +
textdistance.jaccard("this is my test", "that is my test")
textdistance.jaccard("this is my test", "mine this test of mine")
textdistance.jaccard("this is my test", "my test is this")
# -
# ## Some real life applications of fuzzy matching
# https://towardsdatascience.com/natural-language-processing-for-fuzzy-string-matching-with-python-6632b7824c49
#
# https://www.youtube.com/watch?v=kTS2b6pGElE&t=940s
#
# https://www.youtube.com/watch?v=s0YSKiFdj8Q&t=15s
#
#
# +
#Class example
fuzz.token_set_ratio('The quick brown fox', 'jumped over the lazy dogs')
fuzz.token_sort_ratio('The quick brown fox', 'jumped over the lazy dogs')
fuzz.token_set_ratio('This is my test', 'This test is mine')
fuzz.token_sort_ratio('This is my test', 'This test is mine')
# -
| ExampleNotebooks/Fuzzy Matching of data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Correção de Tarefas
# Verifica as atividades feitas e gera os CSV para serem incluídos nas planilhas de controle de notas da disciplina de DeepLearning da UnB
# +
import requests
from IPython.display import JSON
token = '<PASSWORD>'
url = 'https://api.github.com/repos/deeplearningunb/building-ann/branches?per_page=500'
headers = {
"Authorization": "Token {}".format(token)
}
res = requests.get(url,headers=headers)
JSON(res.json())
# +
import requests
url = 'https://api.github.com/repos/deeplearningunb/building-ann/branches?per_page=500'
headers = {
"Authorization": "Token {}".format(token)
}
res = requests.get(url,headers=headers)
tasks = []
for branch in res.json():
# print(branch['name'],"\t",branch['commit']['url'])
commit = requests.get(branch['commit']['url'],headers=headers)
if commit.json()['author'] != None:
author = commit.json()['author']['login']
# print('\t',author)
files = ""
for file in commit.json()['files']:
filename = file['filename']
additions = file['additions']
deletions = file['deletions']
changes = "-{} +{}".format(deletions,additions)
# print('\t\t',filename,changes)
files = files + filename + " " + changes + "\n"
tasks.append({'name':branch['name'],'url': branch['commit']['url'],'author':author,'files':files.strip()})
import pandas as pd
df1 = pd.DataFrame(tasks)
df1.to_csv('tarefa1.csv',index=False)
df1.head()
# +
# Task 3
import requests
url = 'https://api.github.com/repos/deeplearningunb/building-cnn/branches?per_page=500'
headers = {
"Authorization": "Token {}".format(token)
}
res = requests.get(url,headers=headers)
tasks = []
for branch in res.json():
# print(branch['name'],"\t",branch['commit']['url'])
commit = requests.get(branch['commit']['url'],headers=headers)
if commit.json()['author'] != None:
author = commit.json()['author']['login']
# print('\t',author)
files = ""
for file in commit.json()['files']:
filename = file['filename']
additions = file['additions']
deletions = file['deletions']
changes = "-{} +{}".format(deletions,additions)
# print('\t\t',filename,changes)
files = files + filename + " " + changes + "\n"
tasks.append({'name':branch['name'],'url': branch['commit']['url'],'author':author,'files':files.strip()})
import pandas as pd
df3 = pd.DataFrame(tasks)
df3.to_csv('tarefa3.csv',index=False)
df3.head(30)
# +
# Task 4
import requests
url = 'https://api.github.com/repos/deeplearningunb/building-rnn/branches?per_page=500'
headers = {
"Authorization": "Token {}".format(token)
}
res = requests.get(url,headers=headers)
tasks = []
for branch in res.json():
# print(branch['name'],"\t",branch['commit']['url'])
commit = requests.get(branch['commit']['url'],headers=headers)
if commit.json()['author'] != None:
author = commit.json()['author']['login']
# print('\t',author)
files = ""
for file in commit.json()['files']:
filename = file['filename']
additions = file['additions']
deletions = file['deletions']
changes = "-{} +{}".format(deletions,additions)
# print('\t\t',filename,changes)
files = files + filename + " " + changes + "\n"
tasks.append({'name':branch['name'],'url': branch['commit']['url'],'author':author,'files':files.strip()})
import pandas as pd
df4 = pd.DataFrame(tasks)
df4.to_csv('tarefa4.csv',index=False)
df4.head(30)
# +
# Task 5
import requests
url = 'https://api.github.com/repos/deeplearningunb/building-som/branches?per_page=500'
headers = {
"Authorization": "Token {}".format(token)
}
res = requests.get(url,headers=headers)
tasks = []
for branch in res.json():
# print(branch['name'],"\t",branch['commit']['url'])
commit = requests.get(branch['commit']['url'],headers=headers)
if commit.json()['author'] != None:
author = commit.json()['author']['login']
# print('\t',author)
files = ""
for file in commit.json()['files']:
filename = file['filename']
additions = file['additions']
deletions = file['deletions']
changes = "-{} +{}".format(deletions,additions)
# print('\t\t',filename,changes)
files = files + filename + " " + changes + "\n"
tasks.append({'name':branch['name'],'url': branch['commit']['url'],'author':author,'files':files.strip()})
import pandas as pd
df5 = pd.DataFrame(tasks)
df5.to_csv('tarefa5.csv',index=False)
df5.head(30)
# -
# ## FIM =)
| get-tarefas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # t-Stochastic Neighbor Embedding (t-SNE)
# Reference paper: TODO https://jmlr.org/papers/volume9/vandermaaten08a/vandermaaten08a.pdf
# ## Introduction
# t-SNE is a non-linear dimension reduction technique. This is usefull for visualizing high dimensional data.
#
# TODO add example
# ## Other approches
# - Chernoff faces
# - PCA
# Lets define $p_{j|i}$ as the probability that we pick $x_j$ if we choose the points with probability proportional to a gaussian centered at $x_i$.
#
# More formally if we choose a gaussian of variance $\sigma_i$:
# $$\forall i, \forall j \neq i, p_{j|i}=\frac{\exp\left(-\frac{||x_i-x_j||}{2\sigma_i^2}\right)}{\sum_{k\neq i}\exp\left(-\frac{||x_i-x_k||}{2\sigma_i^2}\right)}$$
# $$\forall i, p_{i|i}=0$$
# Note that the denominator is just here for the normalization constraint:
# $$\sum_{j}p_{j|i}=1$$
#
# Just as we did for the $x_i$, we can define $q_{j|i}$ for the $y_i$.
#
# To measure how faithfully $q_{j|i}$ models $p_{j|i}$, we define the Kullback-Leibler divergence:
# $$C=\sum_{i,j}p_{j|i}\log{\frac{p_{j|i}}{q_{j|i}}}$$
# By the properties of morphims of the logarithm, this just is the cross entropy up to an additive constant.
# $$C=-\sum_{i,j}p_{j|i}\log{q_{j|i}} + \text{ cst}$$
#
# The gradient is:
# $$\frac{\delta C}{\delta y_i}=2\sum_j(p_{j|i}-q_{j|i}+p_{i|j}-q_{i|j})(y_i-y_j)$$
#
# Symetric:
# $$p_{ij}=\frac{p_{j|i}+p_{i|j}}{2n}$$
#
# Gradient for symetric SNE:
# $$\frac{\delta C}{\delta y_i}=4\sum_j(p_{ij}-q_{ij})(y_i-y_j)$$
#
# To prevent crowding:
# $$q_{ij}=\frac{(1+||y_i-y_j||^2)^{-1}}{\sum_{k\neq l}(1+||y_k-y_l||^2)^{-1}}$$
#
# The gradient now becomes:
# $$\frac{\delta C}{\delta y_i}=4\sum_j(p_{ij}-q_{ij})(y_i-y_j)(1+||y_i-y_j||^2)^{-1}$$
# ## Algorithm outline
#
# ### Input
#
# - data set $\mathcal{X}=\{x_1, \dots, x_n\}$
# - optimization parameters: number of iterations $T$, learning rate $\eta$, momentum $\alpha(t)$
# - cost function parameters: perplexity $Per p$
#
# ### Output
#
# low dimentional representation of the data: $\mathcal{Y}^{(T)}=\{y_1, \dots, y_n\}$
#
# ### Initialization
#
# - compute $p_{j|i}$ and $p_{ij}$
# - choose an initial solution $\mathcal{Y}^{(0)}=\{y_1, \dots, y_n\}$
#
# ### For $t=1$ to $T$
#
# - compute $q_{ij}$
# - compute the gradient
# - set $\mathcal{Y}^{(t)}:=\mathcal{Y}^{(t-1)}+\eta\frac{\delta C}{\delta \mathcal{Y}}+\alpha(t)\left(\mathcal{Y}^{(t-1)}-\mathcal{Y}^{(t-2)}\right)$
# ## Imports
# +
# %matplotlib notebook
import numpy as np
import scipy.io as sio
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from matplotlib import cm
from math import ceil, sqrt, sin, exp
# -
# ## Binary search
def binary_search(f, y, a, b, epsilon, max_nb_iter=50):
""" Find a <= x <= b such that |f(x) - y| <= epsilon. Assumes that f is monotone. """
m = (a + b) / 2
fm = f(m)
nb_iter = 0
while abs(fm - y) > epsilon and nb_iter <= max_nb_iter:
if fm > y:
b = m
else:
a = m
m = (a + b) / 2
fm = f(m)
nb_iter += 1
return m
# Test.
binary_search(lambda x: x ** 2, 9, -1, 10, 10 ** (-5))
# ## Toy data generation
def gen_toy_data(n=50, sigma=0.05, show=False):
# Make sure that n is a square.
sqrt_n = ceil(sqrt(n))
n = sqrt_n ** 2
# Sample the manifold.
xs = np.linspace(-1, 1, sqrt_n)
ys = np.linspace(-1, 1, sqrt_n)
xs, ys = np.meshgrid(xs, ys)
zs = xs + np.sin(4 * (xs ** 2 + ys ** 2))
# Add noise.
zs += sigma * np.random.randn(*zs.shape)
X = np.array([xs.flatten(), ys.flatten(), zs.flatten()]).T
# Show the generated data.
if show:
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
ax.plot_surface(xs, ys, zs, cmap=cm.coolwarm, linewidth=0, antialiased=False)
plt.show()
return X
toy_X = gen_toy_data(show=True)
# ## Load the MNIST dataset
# +
max_category_size = 50
digits = [3, 4] # 3 and 4 are cherry picked.
# digits = list(range(10))
# -
# Load the MNIST dataset.
data = sio.loadmat('mnist_digits.mat')
# +
X = data['x']
Y = data['y']
n, dim = X.shape
print(f'The dataset has {n} examples of dimension {dim}.')
# Select only the digits in `digits`.
categories = [[]] * 10
for digit in digits:
category = X[np.argwhere(Y == digit)[:, 0]]
categories[digit] = category[:min(max_category_size, len(category))]
X = np.concatenate([categories[digit] for digit in digits], axis=0)
Y = np.concatenate([i * np.ones(len(category)) for i, category in enumerate(categories)], axis=0)
indices = [[]] * 10
for digit in digits:
indices[digit] = np.argwhere(Y == digit)[:, 0]
n, dim = X.shape
print(f'The trimmed down dataset has {n} examples of dimension {dim}.')
# Add noise to the data.
sigma = .05
noisy_X = X + sigma * np.random.randn(*X.shape)
# -
plt.imshow(np.reshape(X[0],(28,28)), cmap='binary')
plt.show()
# Check the level of noise.
plt.imshow(np.reshape(noisy_X[0],(28,28)), cmap='binary')
plt.show()
# ## PCA
def pca(X, d=1, plot=False, label=''):
# Assert that the data is centered.
mean = np.mean(X, axis=0)
X_centered = np.array([x - mean for x in X])
# Compute the eigenvectors and eigenvalues.
eig_val, eig_vect = np.linalg.eig(X_centered.T @ X_centered)
# Remove the imaginary part.
eig_val = np.real(eig_val)
eig_vect = np.real(eig_vect)
# Sort by eigen value.
sort_idx = np.argsort(eig_val)
sorted_eig_val = eig_val[sort_idx]
sorted_eig_vect = eig_vect[sort_idx]
# Plot the eigenvalues.
if plot:
plt.plot(np.maximum(sorted_eig_val, 10 ** (-11)), label=label)
plt.yscale('log')
# Compute the projection on the subspace.
sub_space = sorted_eig_vect[-d:]
Z = np.array([np.array([np.dot(v, x) for v in sub_space]) for x in X_centered + mean])
return Z
def show_2d_pca(X, nb_points=100):
Z = pca(X, d=2)
for digit in digits:
Z_digit = Z[indices[digit]][:nb_points]
plt.scatter(Z_digit[:, 0], Z_digit[:, 1], label=f'{digit}')
plt.title('MNIST PCA 2D')
plt.legend()
plt.show()
show_2d_pca(X)
show_2d_pca(noisy_X)
# ## t-SNE
def tsne(X, T, eta, alpha, per, toy=False, init_with_pca=True):
""" 2D t-SNE """
n = len(X)
colors = ['green', 'blue', 'orange', 'purple', 'black', 'yelow', 'grey', 'brown', 'cyan', 'red']
# Compute sigma for a given perplexity.
# TODO
sigma = np.array([
binary_search(lambda x: 2 ** (-sum()), per, a, b, 1e-5) for i in rang(n)])
sigma = np.ones(n) * 10 ** (-3) # TODO
# Compute p_{j|i}.
N = np.array([[np.linalg.norm(X[i] - X[j]) ** 2 for i in range(n)] for j in range(n)])
p = np.exp(- N ** 2 / (2 * sigma ** 2))
# Normalize.
for i in range(n):
p[:, i] /= np.sum(p[:, i])
# Compute p_{ji}.
P = (p + p.T) / (2 * n)
# Initial solution.
if init_with_pca:
Y = pca(X, d=2)
else:
Y = np.random.normal(0, 10 ** (-4), size=(n, 2))
Y_old = np.copy(Y)
# Training loop.
for t in range(T):
if t % (T // 10) == 0:
# Plot the actual result.
if toy:
plt.scatter(Y[:, 0], Y[:, 1], label=f'{t}; iter {t}')
else:
for digit in digits:
plt.scatter(Y[indices[digit], 0], Y[indices[digit], 1], label=f'{digit}; iter {t}', c=colors[digit])
plt.legend()
plt.show()
# Compute q_{ij}.
Q = np.zeros((n, n))
for i in range(n):
for j in range(n):
if i != j:
Q[i][j] = 1 / (1 + np.linalg.norm(Y[i] - Y[j]) ** 2)
# Normalize.
Q[i] /= np.sum(Q[i])
# Compute de gradient.
gradient = np.array(
[4 * sum(
(P[i, j] - Q[i, j]) * (Y[i] - Y[j]) / (1 + np.linalg.norm(Y[i] - Y[j]) ** 2)
for j in range(n))
for i in range(n)])
if t % (T // 10) == 0:
# Print the cost and the gradient.
cost = np.sum(P * np.maximum(np.log(np.maximum(P / (np.maximum(Q, 10 ** (-10))), 10 ** (-10))), -10 ** 30))
print(f'Iteration {t}/{T}:\tcost {cost}\t\tgradient {np.linalg.norm(gradient)}')
# Update Y and Y_old.
Y, Y_old = Y + eta * gradient + alpha[t] * (Y - Y_old), Y
return Y
# +
# Parameters.
T = 100
eta = .1
alpha = 0 * np.ones(T)
per = 1
Y = tsne(toy_X, T, eta, alpha, per, toy=True, init_with_pca=False)
plt.legend()
plt.show()
# +
# Parameters.
T = 100
eta = .05
alpha = 0 * np.ones(T)
per = 1
Y = tsne(X, T, eta, alpha, per, toy=False)
# noisy_Y = tsne(noisy_X, T, eta, alpha, per)
# -
# # TODO
# - perplexity/sigma
# - at first use PCA to reduce the dimention to 30
# - remove for loops
# - plot the cost
# - normalize
# - log and exp in base 2
# - springs
# - add gaussian noise to the map points in the early stages of optimization
# - early compression
# - early exaggeration
# - random walk
| t-SNE.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <!--NOTEBOOK_HEADER-->
# *This notebook contains course material from [CBE30338](https://jckantor.github.io/CBE30338)
# by <NAME> (jeff at nd.edu); the content is available [on Github](https://github.com/jckantor/CBE30338.git).
# The text is released under the [CC-BY-NC-ND-4.0 license](https://creativecommons.org/licenses/by-nc-nd/4.0/legalcode),
# and code is released under the [MIT license](https://opensource.org/licenses/MIT).*
# <!--NAVIGATION-->
# < [Path Constraints](http://nbviewer.jupyter.org/github/jckantor/CBE30338/blob/master/notebooks/07.08-Path-Constraints.ipynb) | [Contents](toc.ipynb) | [Zero-Order Hold and Interpolation](http://nbviewer.jupyter.org/github/jckantor/CBE30338/blob/master/notebooks/08.01-Zero-Order-Hold-and-Interpolation.ipynb) ><p><a href="https://colab.research.google.com/github/jckantor/CBE30338/blob/master/notebooks/08.00-Predictive-Control.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open in Google Colaboratory"></a><p><a href="https://raw.githubusercontent.com/jckantor/CBE30338/master/notebooks/08.00-Predictive-Control.ipynb"><img align="left" src="https://img.shields.io/badge/Github-Download-blue.svg" alt="Download" title="Download Notebook"></a>
# # Predictive Control
# <!--NAVIGATION-->
# < [Path Constraints](http://nbviewer.jupyter.org/github/jckantor/CBE30338/blob/master/notebooks/07.08-Path-Constraints.ipynb) | [Contents](toc.ipynb) | [Zero-Order Hold and Interpolation](http://nbviewer.jupyter.org/github/jckantor/CBE30338/blob/master/notebooks/08.01-Zero-Order-Hold-and-Interpolation.ipynb) ><p><a href="https://colab.research.google.com/github/jckantor/CBE30338/blob/master/notebooks/08.00-Predictive-Control.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open in Google Colaboratory"></a><p><a href="https://raw.githubusercontent.com/jckantor/CBE30338/master/notebooks/08.00-Predictive-Control.ipynb"><img align="left" src="https://img.shields.io/badge/Github-Download-blue.svg" alt="Download" title="Download Notebook"></a>
| Mathematics/Mathematical Modeling/08.00-Predictive-Control.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h3>Logistic Regression</h3>
# <h4>Packages Used</h4>
# <ul>
# <li>numpy</li>
# <li>matplotlib</li>
# <li>scipy</li>
# </ul>
# <h3>Import necessary packages</h3>
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import scipy.optimize as op
plt.rcParams['figure.figsize'] = [10, 8]
# <h3>ReadData(data,separator): Helper function to read data</h3>
# <h4> Assumes data is of the form X[0], X[1], ..., X[n], Y</h4>
# <h5>Where X[i] is a feature and Y is the label</h5>
def ReadData(data, separator):
XY = np.genfromtxt(data, delimiter=separator)
m = XY.shape[0]
Y = XY[:, -1].reshape(m, 1)
X = XY[:, 0:-1]
bias = np.zeros((1, 1)) - 24
theta = np.zeros((X.shape[1], 1)) + 0.2
return X, Y, m, bias, theta
# <h3>Normalize(data): Helper function to Normalize data</h3>
def Normalize(data):
Mu = np.mean(X, axis=0)
Sigma = np.std(X, axis=0)
data = ((data-Mu)/Sigma)
return data, Mu, Sigma
# <h3>GradientDescent(theta, X, Y, costweight): Function to calculate the Gradient</h3>
def GradientDescent(theta, X, Y, costweight):
bias = theta.item(0)
theta = np.delete(theta, 0).reshape(len(theta) - 1,1)
H = Sigmoid(np.dot(X, theta) + bias)
diff = H - Y
theta = costweight * np.dot(diff.T,X).T
bias = costweight * sum(diff)
return np.insert(theta,0,bias)
# <h3>CostCalc(X,theta,bias,Y,costweight): Function to calculate cost</h3>
def CostCalc(theta, X, Y, costweight):
bias = theta.item(0)
theta = np.delete(theta, 0).reshape(len(theta) - 1,1)
H = Sigmoid(np.dot(X, theta) + bias)
J = -1 * costweight * sum(Y * np.log(H) + (1-Y) * np.log(1-H))
return J
# <h3>PlotData(theta,X,Y,fignumber=1): Helper function to Plot data, contour plot</h3>
def PlotData(theta,X,Y,fignumber=1):
plt.style.use('ggplot')
plt.figure(fignumber)
plt.subplot(111)
X1 = np.array([[X[i,0],X[i,1]] for i in range(len(X)) if Y[i,0] == 1])
X0 = np.array([[X[i,0],X[i,1]] for i in range(len(X)) if Y[i,0] == 0])
plt.plot(X0[:,0],X0[:,1],'ro',label='class 0')
plt.plot(X1[:,0],X1[:,1],'bo',label='class 1')
plt.ylabel('Feature 2')
plt.xlabel('Feature 1')
plt.legend(bbox_to_anchor=(0.80, 1.15), loc=2,mode="expand", borderaxespad=0.)
ContourPlot(theta, X)
return
# +
def ContourPlot(theta, X):
Y = np.linspace(min(X[:,1]), max(X[:,1]), 500)
X1 = np.linspace(min(X[:,0]), max(X[:,0]), 500)
Z = ContourPlotCalc(theta,X1,Y)
X , Y = np.meshgrid(X1, Y)
CS = plt.contour(X, Y, Z,colors='k', alpha=.5)
CS1 = plt.contourf(X, Y, Z,cmap='RdBu', alpha=.5)
return
################################################################################
################################################################################
def ContourPlotCalc(theta, X,Y):
Z=np.zeros((len(X),len(Y)))
bias = theta.item(0)
theta = np.delete(theta, 0).reshape(len(theta) - 1,1)
for i in range(len(X)):
for j in range(len(Y)):
##Z[j][i] because contour plot needs costs[i][j] Transpose
Z[j][i] = np.dot(np.array([X[i],Y[j]]).reshape(1,2),theta) + bias
Z = np.where(Z > 0.5, 1, 0)
return Z
# -
def Sigmoid(Z):
Z = 1/(1 + np.exp(-Z))
return Z
def Predict(theta, X):
bias = theta.item(0)
theta = np.delete(theta, 0).reshape(len(theta) - 1,1)
P = np.dot(X,theta) + bias
P = np.where(P > 0, 1, 0)
return P
# <h2>Main Code below</h2>
X,Y,m,bias,theta = ReadData('LogRegDS.txt',',')
costweight = 1/m
initial_theta = np.insert(theta,0,bias)
cost = CostCalc(initial_theta,X,Y,costweight)
#print(cost)
initial_theta = np.insert(theta,0,bias)
Result = op.minimize(fun = CostCalc, x0 = initial_theta, args = (X,Y,costweight), method = 'BFGS', jac = GradientDescent)
#print(Result)
cost = Result.fun
trained_theta = Result.x
P = Predict(trained_theta, X)
error = costweight * sum(np.abs(P-Y)) * 100
Accuracy = 100 - error[0]
print(f'Accuracy = {Accuracy} %\ncost = {cost}')
PlotData(trained_theta, X,Y)
| Basic ML algorithms/Logistic Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # "Shell Scripts: env-shebang with Arguments"
# > "What you need to pass arguments to an interpreter found by 'env'."
# - author: jhermann
# - toc: false
# - branch: master
# - badges: true
# - comments: true
# - published: true
# - categories: [linux, know-how]
# - image: images/copied_from_nb/img/linux/linux-shell.png
# 
# ## The Problem
#
# There is an old annoyance that, if you use `env` in a bang path to search the script interpreter in the shell's path, you cannot pass any arguments to it. Instead, all the text after the call to `env` is passed as one single argument, and `env` tries to find this as the executable to invoke, which fails of course when arguments are present.
#
# `env` is not the culprit here, but the very definition of how a bang path works (quoted from the `bash` manpage):
#
# > If the program is a file beginning with ``#!``, the remainder of the first line specifies an interpreter for the program.
# > The shell executes the specified interpreter on operating systems that do not handle this executable format themselves.
# > The **arguments to the interpreter consist of a *single* optional argument** following the interpreter name on the first line… *(emphasis mine)*
#
# So what env gets to see in its ``argv`` array when you write something like ``#! /usr/bin/env python3 -I -S`` is ``['/usr/bin/env', 'python3 -I -S']``. And there is no ``python3 -I -S`` anywhere to be found that could interpret your script. 😞
# ## The Solution
#
# The `env` command in coreutils 8.30 solves this (i.e. Debian Buster only so far, Ubuntu Bionic still has 8.28). The relevant change is introducing a split option (``-S``), designed to handle that special case of getting all arguments mushed together into one.
#
# In the example below, we want to pass the ``-I -S`` options to Python on startup. They increase security of a script, by reducing the possible ways an attacker can insert their malicious code into your runtime environment, as you can see from the help text:
#
# ```
# -I : isolate Python from the user's environment (implies -E and -s)
# -E : ignore PYTHON* environment variables (such as PYTHONPATH)
# -s : don't add user site directory to sys.path; also PYTHONNOUSERSITE
# -S : don't imply 'import site' on initialization
# ```
#
# You can try the following yourself using `docker run --rm -it --entrypoint /bin/bash python:3-slim-buster`:
#
# ```console
# $ cat >isolated <<'.'
# # #!/usr/bin/env -S python3 -I -S
# import sys
# print('\n'.join(sys.path))
# .
# $ chmod +x isolated
# $ ./isolated
# /usr/local/lib/python38.zip
# /usr/local/lib/python3.8
# /usr/local/lib/python3.8/lib-dynload
# ```
#
# Normally, the Python path would include both the current working directory (`/` in this case) as well as site packages (`/usr/local/lib/python3.8/site-packages`).
#
# However, we prevented their inclusion as a source of unanticipated code – and you can be a happy cat again. 😻
| _notebooks/2020-02-28-env_with_arguments.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Regression | | <NAME> | Wednesday March 28th 2018
# +
# Import the necessary libraries
import numpy as np
import pandas as pd
import datetime as date
import matplotlib.pyplot as plt
# %matplotlib inline
import numpy.polynomial.polynomial as poly
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error
import matplotlib
import seaborn as sns
matplotlib.rcParams['figure.figsize'] = [12.0, 8.0]
# -
df = pd.read_csv('./ecommerce-customers.csv', error_bad_lines=False).drop_duplicates()
pd.DataFrame(df.head(100))
df.info()
df.describe()
# # Use seaborn jointplot function to see which fields correlate well with the "Yearly Amount Spent" column. Write your findings.
#
# ### From our analysis it seems Length of Membership correalates very well with a larger average per year spend.
sns.jointplot(df['Avg. Session Length'], df['Yearly Amount Spent'])
sns.jointplot(df['Time on App'], df['Yearly Amount Spent'])
sns.jointplot(df['Time on Website'], df['Yearly Amount Spent'])
sns.jointplot(df['Length of Membership'], df['Yearly Amount Spent'])
sns.pairplot(df)
# +
# sns.lmplot(x='Length of Membership', y='Yearly Amount Spent')
# -
# ## Create two dataframes: one for the target variable ("Yearly Amount Spent"), the other - containing all the rest of numerical features
YAS_df = df['Yearly Amount Spent']
all_but_YAS_df = df[['Avg. Session Length', 'Time on App', 'Time on Website', 'Length of Membership']]
# ## Split the data into a training and test sets. Make a test set size 0.3 and random seed 123
# +
# Generate the full data and split it into a training set and test set.
X_full = all_but_YAS_df['Length of Membership']
Y_full = YAS_df
X_train, X_test, Y_train, Y_test = train_test_split(X_full, Y_full, test_size=0.3, random_state=123)
# -
plt.plot(X_train, Y_train, ".", markersize=20)
plt.title("Test Set")
# ## Fit a regression model on the training set
# +
coefs_lin = poly.polyfit(X_train, Y_train, 1)
X_line = np.linspace(0, 10, 10000)
ffit_lin = poly.polyval(X_line, coefs_lin)
# -
plt.ylim(min(Y_train - 100), max(Y_train + 100))
plt.plot(X_train, Y_train, "r.", markersize=5)
plt.plot(X_line, ffit_lin, "g")
plt.show()
# ## Print out the coefficients of the model
print(coefs_lin)
# ## Make a prediction of the target variable from features dataframe
# +
# Length of Membership would make a good target variable
# -
# ## Calculate the Mean Squared Error (using sklearn.metrics module)
mse_lin = mean_squared_error(Y_train, poly.polyval(X_train, coefs_lin))
print("MSE for a linear regressor: ", mse_lin)
# ## Using Seaborn distplot show the histogram of the residuals - differences between the target variable and predicted target variable
sns.distplot(df['Length of Membership'])
# ## How should we allocate the engineering budget between website development and app development?
# ### App development because the coefficient between time on app and the avg yearly spend is very high. Thus development to get users to spend a small amount more time on the app would result in a relatively large increase in avg yearly spend.
| regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# ## 1) Let's prepare the scripts to be deployed as a Glue Job
# + language="bash"
# mkdir /home/ec2-user/SageMaker/Analytics_Labs/glue_python_shell_sample
# + language="bash"
# cd /home/ec2-user/SageMaker/Analytics_Labs/glue_python_shell_sample
# wget https://raw.githubusercontent.com/angelocarvalho/glue-python-shell-sample/master/glue-python-shell-sample-whl/setup.py
# + language="bash"
# cd /home/ec2-user/SageMaker/Analytics_Labs/glue_python_shell_sample
# python3 setup.py bdist_wheel
# + language="bash"
# cd /home/ec2-user/SageMaker/Analytics_Labs/glue_python_shell_sample/dist
# wget https://raw.githubusercontent.com/angelocarvalho/glue-python-shell-sample/master/etl_with_pandas.py
# +
import simplejson
with open('/opt/ml/metadata/resource-metadata.json') as fh:
metadata = simplejson.loads(fh.read())
accountid = metadata['ResourceArn'].split(':')[4]
# %set_env accountid={accountid}
# %set_env bucket_name=lab-{accountid}
# -
# ## 2) Open the file etl_with_pandas.py and add your bucket name!
# + language="bash"
# aws s3 cp /home/ec2-user/SageMaker/Analytics_Labs/glue_python_shell_sample/dist/glue_python_shell_sample_module-0.1-py3-none-any.whl s3://$bucket_name/lib/
# aws s3 cp /home/ec2-user/SageMaker/Analytics_Labs/glue_python_shell_sample/dist/etl_with_pandas.py s3://$bucket_name/scripts/
# -
# ## 3) Replace the iam role and and the bucket name before running the command bellow
# + language="bash"
# aws glue create-job --name etl_with_pandas \
# --role <<nome_do_seu_iam_role>> \
# --command '{"Name" : "pythonshell", "PythonVersion" : "3", "ScriptLocation" : "s3://<<nome_do_seu_bucket>>/scripts/etl_with_pandas.py"}' \
# --default-arguments '{"--extra-py-files" : "s3://<<nome_do_seu_bucket>>/lib/glue_python_shell_sample_module-0.1-py3-none-any.whl"}'
| LAB11_Glue_Job.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# project: p4
# submitter: zchen697
# partner: none
# -
import project
import math
def get_stat_total(pkmn):
return project.get_hp(pkmn)+project.get_attack(pkmn)+project.get_defense(pkmn)+project.get_sp_atk(pkmn)+project.get_sp_def(pkmn)+project.get_speed(pkmn)
def simple_battle(pkmn1,pkmn2):
a = get_stat_total(pkmn1)
b = get_stat_total(pkmn2)
if a > b and (a - b) <= 300:
return pkmn1
elif b > a and (b - a) <= 300:
return pkmn2
elif a - b > 300:
return pkmn2 + " ran away"
elif b - a > 300:
return pkmn1 + " ran away"
else:
return "Draw"
#q1
simple_battle('Snorunt', 'Starly')
#q2
simple_battle('Snorunt', 'Staravia')
#q3
simple_battle('Chikorita', 'Turtwig')
#q4
simple_battle('Caterpie', 'Melmetal')
#q5
simple_battle('Dragonite', 'Snorunt')
def most_damage(attacker, defender):
physical_damage = 10 * project.get_attack(attacker)/project.get_defense(defender)
special_damage = 10 * project.get_sp_atk(attacker)/project.get_sp_def(defender)
if physical_damage > special_damage:
return physical_damage
else:
return special_damage
#q6
most_damage('Dragonite', 'Rockruff')
#q7
most_damage('Quilava', 'Grovyle')
#q8
most_damage('Goomy', 'Beedrill')
#q9
most_damage('Tepig', 'Charizard')
def num_hits(attacker, defender):
return math.ceil(project.get_hp(defender)/most_damage(attacker,defender))
#q10
num_hits('Gible', 'Goomy')
#q11
num_hits('Aipom', 'Donphan')
#q12
num_hits('Donphan', 'Aipom')
def battle(pkmn1, pkmn2):
#TODO: Return the name of the pkmn that can take more hits from the other
# pkmn. If both pkmn faint within the same number of moves, return the
# string 'Draw'
a = num_hits(pkmn1, pkmn2)
b = num_hits(pkmn2, pkmn1)
if abs(a - b) > 10:
if a > b:
return pkmn1 + " ran away"
else:
return pkmn2 + " ran away"
if a > b and abs(a - b) <10:
return pkmn2
elif a < b and abs(a - b) < 10:
return pkmn1
elif a == b and project.get_speed(pkmn1) == project.get_speed(pkmn2):
return "Draw"
elif a == b and project.get_speed(pkmn1) > project.get_speed(pkmn2):
return pkmn1
else:
return pkmn2
#q13
battle('Scraggy', 'Krabby')
#q14
battle('Charizard', 'Krabby')
#q15
battle('Treecko', 'Litten')
#q16
battle('Treecko', 'Buizel')
#q17
battle('Metapod', 'Talonflame')
#q18
battle('Leavanny', 'Noibat')
def final_battle(pkmn1, pkmn2):
a = project.get_region(pkmn1)
b = project.get_region(pkmn2)
pkmn1_t1 = project.get_type1(pkmn1)
pkmn1_t2 = project.get_type2(pkmn1)
pkmn2_t1 = project.get_type1(pkmn2)
pkmn2_t2 = project.get_type2(pkmn2)
if a != b and pkmn1_t1 != "Flying" and pkmn1_t2 != "Flying" and pkmn2_t1 != "Flying" and pkmn2_t2 != "Flying":
return "Cannot battle"
else:
return battle(pkmn1, pkmn2)
#q19
final_battle('Grotle', 'Roggenrola')
#q20
final_battle('Starly', 'Goodra')
| p4/cs-220-test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.0 64-bit
# language: python
# name: python3
# ---
class Solution(object):
def mySqrt(self, x):
"""
:type x: int
:rtype: int
"""
return int((x**0.5)//1)
class Solution(object):
def mySqrt(self, x):
"""
:type x: int
:rtype: int
"""
i=0
while (i*i<x):
i+=1
if i*i>x:
i-=1
return i
| Problems/69-SqrtX.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/kalz2q/mycolabnotebooks/blob/master/diary.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="UdrKZGG2ClVW"
2021/05/01 3:33 土曜日
アイデア
コロナ時代の学習教材としての Colab
コロナ 学習 対面 オンライン授業 学習教材 ビデオ ストリーミング
コロナ禍の学習教材としての Colab
はじめに
コロナ時代の学習教材としての Colab に注目!!!!
対象は高校生以上なら大丈夫だと思う。 教材作りの方が大変だけど使い回しができるのでみんなで作れば怖くない。
流れ
1. 生徒は https://colab.research.google.com/ にアクセスしてアカウントを作る
2. 教材を Colab で開く URL をメールで配布する。 例 https://colab.research.google.com/github/kalz2q/mycolabnotebooks/blob/master/doingmath01.ipynb
3. 生徒はメニューの"ドライブにコピー"で自分の Colab にコピーする
4. 生徒はセルを実行したり、壊したりしながら実験学習する
メリット・ディメリット
1. だれでも使える。環境設定が不要 (スマホでも使える)
2. 生徒は失敗を恐れずインタラクティブに学習できる
3. 失敗したらもとのファイルからやりなおせばよい
あとがき
# + id="kCIPfBAI1_dE"
2021/05/01 2:33 土曜日
なぜ、github に colab で日記を書くか。
1. バージョン管理ができる (うっかり消しても大丈夫)
2. どこからでも使える。 パソコンが壊れても大丈夫
3. colab (jupyter) のエディターが複数カーソル(multicursor) に対応している (CodeMirror ベース?) とかそこそこ便利
4. google chrome の keep との使い分けができる
5. コードセル/テキストセルの切り替えができる
6. コードセルでは ubuntu (linux) の機能が使える
7. テキストセルでは latex が使えて表現が豊富になる
8. github にセーブできる (もっとも安全なサーバー)
9. git clone して grep とか ubuntu (linux) の機能がつかえて超便利
10. colab には自動保存される。 github にコピーを保存が便利 <= あ、これじゃあわからんね!!!!
思いつき。アイデア。
メニューのキーボードショートカットにはエディター内で使える機能が載っていないのでリストを作る。
ショートカット | 機能 | 備考
--- | --- | ---
Ctrl+C | コピー |
Ctrl+Click | 複数カーソル | muliticursor
てかだれか作っているだろう。
colab editor shortcuts
で検索 check_later
メモ
日本版wikipediaに翻訳記事作成で参加しています
# + [markdown] id="wb8ofcd06ir4"
# kljl; lj;j
#
# + id="yOcz-bF1kfEr"
2021/05/01 2:25 土曜日
ここはパブリックなので、あたりさわりのないことを書く。
お茶の成分はタンニンでコーヒーの成分はカフェイン、というのはなんか違う気がする。
お茶にもカフェインが入っている。 デカフェのコーヒーもコーヒーの味がする。
茶 タンニン コーヒー カフェイン
で検索 => 複雑そう
# + id="JzFAcp-fkhPm"
2021/04/30 20:38
github に書く diary。 メモ代わりには使えるだろう。
| diary.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="https://colab.research.google.com/github/warwickdatascience/beginners-python/blob/master/session_six/session_six_filled_template.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# <center>Spotted a mistake? Report it <a href="https://github.com/warwickdatascience/beginners-python/issues/new">here</a></center>
# # Beginner's Python—Session Six Template
# + [markdown] colab_type="text" id="-Ceh1HnZIsVO"
# ## Functions
# + [markdown] colab_type="text" id="6AZ6H-6jKoSt"
# ### Introduction
#
# -
# Confirm that the type of `print` is `builtin_function_or_method`
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="D4l_2DsZIlgB" outputId="e63ad5b1-2062-4d8c-9ee7-b03a25d72001"
type(print)
# + [markdown] colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="9lLW387eKgNN" outputId="860dcf3b-f3e5-402d-f58f-4d28fb69a6b1"
# Create a function that will print text when ran and run it twice
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="bIr0l5kJI_VV" outputId="c7891804-4acf-4856-e51a-271914a8dc60"
def say_hello():
print("Hello, World!")
say_hello()
print("Once more?")
say_hello()
# -
# Create a function which returns a number and run it, storing the returned value in a variable
# +
def five():
return 5
num = five()
print(num)
# -
# ### Standard Puzzles
# Create a function to print the string 'Hello!' and run this
def greet():
print("Hello")
# Create a function which calls the above function five times
# +
def greet_group():
for i in range(5):
greet()
greet_group()
# -
# Create a function to calculate and return the value of $2 + 2$
# +
def two_plus_two():
return 2 + 2
four = two_plus_two()
print(four)
# -
# ## Function Arguments
# ### Introduction
# Create a function to square an inputted number
# +
def square(num):
print(num ** 2)
square(3)
# -
# Create a function to add two numbers, with the second parameter taking a default value of $1$
def add(a, b=1):
print(a + b)
# ### Standard Puzzles
# Create a function to greet a named person, with fallback default behaviour
def greet_person(name='unknown'):
if name == 'unknown':
print("Hello. What's your name?")
else:
print("Hello,", name)
# Create a function to calculate the area of a rectangle
def rectangle_area(width, height):
return width * height
# ### Bonus Puzzles
# Improve the greeter function above by using `None` as the default value for the name parameter
def greet_person(name=None):
if name is None:
print("Hello. What's your name?")
else:
print("Hello,", name)
# ## Recursion
# ### Introduction
# Create a function to calculate the nth Fibonacci number using recursion
# +
def fibonacci(n):
if n == 0 or n == 1:
return 1
return fibonacci(n-1) + fibonacci(n-2)
fibonacci(10)
# -
# ### Standard Puzzles
# Create a function to calculate $n!$ using recursion
# +
def factorial(n):
if n == 1:
return 1
return n * factorial(n-1)
factorial(4)
# -
# ### Bonus Puzzles
# Refactor the above function using an iterative paradigm
# +
def factorial(n):
prod = 1
for i in range(2, n + 1):
prod *= i
return prod
factorial(4)
# -
# ## Variable Scope
# ### Introduction
# _The examples given in the presentation are not the most suitable to use as exercises, so instead just make sure you've gave them a good look over_
# ### Standard Puzzles
# Predict what values will be printed when running the presentation code. Verify this
# The results will be 10 and 2 respectively. THe first prints the local variable `a` whereas the second prints the global variable
# +
a = 2
def double(b):
a = b * 2
print(a)
double(5)
print(a)
# -
# Rewrite the supplied code to avoid referencing global variables
# +
n = 10
def square(n):
return n ** 2
print(square(n))
| session-six/session_six_filled_template.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # DREAMER Arousal EMI-LSTM 48_16
# Adapted from Microsoft's notebooks, available at https://github.com/microsoft/EdgeML authored by Dennis et al.
# ## Imports
import pandas as pd
import numpy as np
from tabulate import tabulate
import os
import datetime as datetime
import pickle as pkl
# ## DataFrames from CSVs
df = pd.read_csv('/home/sf/data/DREAMER/DREAMER_combined.csv',index_col=0)
# ## Preprocessing
df.columns
# ## Split Ground Truth
filtered_train = df.drop(['Movie', 'Person', 'Arousal','Dominance', 'Valence'], axis=1)
filtered_target = df['Arousal']
filtered_target = filtered_target.replace({1:0,2:1,3:2,4:3,5:4})
print(filtered_target.shape)
print(filtered_train.shape)
y = filtered_target.values.reshape(85744, 128) # 128 is the size of 1 bag, 85744 = (size of the entire set) / 128
# ## Convert to 3D - (Bags, Timesteps, Features)
len(filtered_train.columns)
x = filtered_train.values
print(x.shape)
x = x.reshape(int(len(x) / 128), 128, 16)
print(x.shape)
# ## Filter Overlapping Bags
# filtering bags that overlap with another class
bags_to_remove = []
for i in range(len(y)):
if len(set(y[i])) > 1:
bags_to_remove.append(i)
print(bags_to_remove)
x = np.delete(x, bags_to_remove, axis=0)
y = np.delete(y, bags_to_remove, axis=0)
x.shape
y.shape
# ## Categorical Representation
one_hot_list = []
for i in range(len(y)):
one_hot_list.append(set(y[i]).pop())
categorical_y_ver = one_hot_list
categorical_y_ver = np.array(categorical_y_ver)
categorical_y_ver.shape
x.shape[1]
def one_hot(y, numOutput):
y = np.reshape(y, [-1])
ret = np.zeros([y.shape[0], numOutput])
for i, label in enumerate(y):
ret[i, label] = 1
return ret
# ## Extract 3D Normalized Data with Validation Set
from sklearn.model_selection import train_test_split
import pathlib
x_train_val_combined, x_test, y_train_val_combined, y_test = train_test_split(x, categorical_y_ver, test_size=0.20, random_state=42)
y_test
# +
extractedDir = '/home/sf/data/DREAMER/Arousal/'
timesteps = x_train_val_combined.shape[-2]
feats = x_train_val_combined.shape[-1]
trainSize = int(x_train_val_combined.shape[0]*0.9)
x_train, x_val = x_train_val_combined[:trainSize], x_train_val_combined[trainSize:]
y_train, y_val = y_train_val_combined[:trainSize], y_train_val_combined[trainSize:]
# normalization
x_train = np.reshape(x_train, [-1, feats])
mean = np.mean(x_train, axis=0)
std = np.std(x_train, axis=0)
# normalize train
x_train = x_train - mean
x_train = x_train / std
x_train = np.reshape(x_train, [-1, timesteps, feats])
# normalize val
x_val = np.reshape(x_val, [-1, feats])
x_val = x_val - mean
x_val = x_val / std
x_val = np.reshape(x_val, [-1, timesteps, feats])
# normalize test
x_test = np.reshape(x_test, [-1, feats])
x_test = x_test - mean
x_test = x_test / std
x_test = np.reshape(x_test, [-1, timesteps, feats])
# shuffle test, as this was remaining
idx = np.arange(len(x_test))
np.random.shuffle(idx)
x_test = x_test[idx]
y_test = y_test[idx]
# +
# one-hot encoding of labels
numOutput = 5
y_train = one_hot(y_train, numOutput)
y_val = one_hot(y_val, numOutput)
y_test = one_hot(y_test, numOutput)
extractedDir += '/'
pathlib.Path(extractedDir + 'RAW').mkdir(parents=True, exist_ok = True)
np.save(extractedDir + "RAW/x_train", x_train)
np.save(extractedDir + "RAW/y_train", y_train)
np.save(extractedDir + "RAW/x_test", x_test)
np.save(extractedDir + "RAW/y_test", y_test)
np.save(extractedDir + "RAW/x_val", x_val)
np.save(extractedDir + "RAW/y_val", y_val)
print(extractedDir)
# -
# ls /home/sf/data/DREAMER/Arousal/RAW
np.load('/home/sf/data/DREAMER/Arousal/RAW/x_train.npy').shape
# ## Make 4D EMI Data (Bags, Subinstances, Subinstance Length, Features)
def loadData(dirname):
x_train = np.load(dirname + '/' + 'x_train.npy')
y_train = np.load(dirname + '/' + 'y_train.npy')
x_test = np.load(dirname + '/' + 'x_test.npy')
y_test = np.load(dirname + '/' + 'y_test.npy')
x_val = np.load(dirname + '/' + 'x_val.npy')
y_val = np.load(dirname + '/' + 'y_val.npy')
return x_train, y_train, x_test, y_test, x_val, y_val
def bagData(X, Y, subinstanceLen, subinstanceStride):
numClass = 5
numSteps = 128
numFeats = 16
assert X.ndim == 3
assert X.shape[1] == numSteps
assert X.shape[2] == numFeats
assert subinstanceLen <= numSteps
assert subinstanceLen > 0
assert subinstanceStride <= numSteps
assert subinstanceStride >= 0
assert len(X) == len(Y)
assert Y.ndim == 2
assert Y.shape[1] == numClass
x_bagged = []
y_bagged = []
for i, point in enumerate(X[:, :, :]):
instanceList = []
start = 0
end = subinstanceLen
while True:
x = point[start:end, :]
if len(x) < subinstanceLen:
x_ = np.zeros([subinstanceLen, x.shape[1]])
x_[:len(x), :] = x[:, :]
x = x_
instanceList.append(x)
if end >= numSteps:
break
start += subinstanceStride
end += subinstanceStride
bag = np.array(instanceList)
numSubinstance = bag.shape[0]
label = Y[i]
label = np.argmax(label)
labelBag = np.zeros([numSubinstance, numClass])
labelBag[:, label] = 1
x_bagged.append(bag)
label = np.array(labelBag)
y_bagged.append(label)
return np.array(x_bagged), np.array(y_bagged)
def makeEMIData(subinstanceLen, subinstanceStride, sourceDir, outDir):
x_train, y_train, x_test, y_test, x_val, y_val = loadData(sourceDir)
x, y = bagData(x_train, y_train, subinstanceLen, subinstanceStride)
np.save(outDir + '/x_train.npy', x)
np.save(outDir + '/y_train.npy', y)
print('Num train %d' % len(x))
x, y = bagData(x_test, y_test, subinstanceLen, subinstanceStride)
np.save(outDir + '/x_test.npy', x)
np.save(outDir + '/y_test.npy', y)
print('Num test %d' % len(x))
x, y = bagData(x_val, y_val, subinstanceLen, subinstanceStride)
np.save(outDir + '/x_val.npy', x)
np.save(outDir + '/y_val.npy', y)
print('Num val %d' % len(x))
subinstanceLen = 48
subinstanceStride = 16
extractedDir = '/home/sf/data/DREAMER/Arousal'
from os import mkdir
mkdir('/home/sf/data/DREAMER/Arousal' + '/%d_%d/' % (subinstanceLen, subinstanceStride))
rawDir = extractedDir + '/RAW'
sourceDir = rawDir
outDir = extractedDir + '/%d_%d/' % (subinstanceLen, subinstanceStride)
makeEMIData(subinstanceLen, subinstanceStride, sourceDir, outDir)
np.load('/home/sf/data/DREAMER/Arousal/48_16/y_train.npy').shape
print(x_train.shape)
print(y_train.shape)
from edgeml.graph.rnn import EMI_DataPipeline
from edgeml.graph.rnn import EMI_BasicLSTM, EMI_FastGRNN, EMI_FastRNN, EMI_GRU
from edgeml.trainer.emirnnTrainer import EMI_Trainer, EMI_Driver
import edgeml.utils
def lstm_experiment_generator(params, path = './DSAAR/64_16/'):
"""
Function that will generate the experiments to be run.
Inputs :
(1) Dictionary params, to set the network parameters.
(2) Name of the Model to be run from [EMI-LSTM, EMI-FastGRNN, EMI-GRU]
(3) Path to the dataset, where the csv files are present.
"""
#Copy the contents of the params dictionary.
lstm_dict = {**params}
#---------------------------PARAM SETTING----------------------#
# Network parameters for our LSTM + FC Layer
NUM_HIDDEN = params["NUM_HIDDEN"]
NUM_TIMESTEPS = params["NUM_TIMESTEPS"]
ORIGINAL_NUM_TIMESTEPS = params["ORIGINAL_NUM_TIMESTEPS"]
NUM_FEATS = params["NUM_FEATS"]
FORGET_BIAS = params["FORGET_BIAS"]
NUM_OUTPUT = params["NUM_OUTPUT"]
USE_DROPOUT = True if (params["USE_DROPOUT"] == 1) else False
KEEP_PROB = params["KEEP_PROB"]
# For dataset API
PREFETCH_NUM = params["PREFETCH_NUM"]
BATCH_SIZE = params["BATCH_SIZE"]
# Number of epochs in *one iteration*
NUM_EPOCHS = params["NUM_EPOCHS"]
# Number of iterations in *one round*. After each iteration,
# the model is dumped to disk. At the end of the current
# round, the best model among all the dumped models in the
# current round is picked up..
NUM_ITER = params["NUM_ITER"]
# A round consists of multiple training iterations and a belief
# update step using the best model from all of these iterations
NUM_ROUNDS = params["NUM_ROUNDS"]
LEARNING_RATE = params["LEARNING_RATE"]
# A staging direcory to store models
MODEL_PREFIX = params["MODEL_PREFIX"]
#----------------------END OF PARAM SETTING----------------------#
#----------------------DATA LOADING------------------------------#
x_train, y_train = np.load(path + 'x_train.npy'), np.load(path + 'y_train.npy')
x_test, y_test = np.load(path + 'x_test.npy'), np.load(path + 'y_test.npy')
x_val, y_val = np.load(path + 'x_val.npy'), np.load(path + 'y_val.npy')
# BAG_TEST, BAG_TRAIN, BAG_VAL represent bag_level labels. These are used for the label update
# step of EMI/MI RNN
BAG_TEST = np.argmax(y_test[:, 0, :], axis=1)
BAG_TRAIN = np.argmax(y_train[:, 0, :], axis=1)
BAG_VAL = np.argmax(y_val[:, 0, :], axis=1)
NUM_SUBINSTANCE = x_train.shape[1]
print("x_train shape is:", x_train.shape)
print("y_train shape is:", y_train.shape)
print("x_test shape is:", x_val.shape)
print("y_test shape is:", y_val.shape)
#----------------------END OF DATA LOADING------------------------------#
#----------------------COMPUTATION GRAPH--------------------------------#
# Define the linear secondary classifier
def createExtendedGraph(self, baseOutput, *args, **kwargs):
W1 = tf.Variable(np.random.normal(size=[NUM_HIDDEN, NUM_OUTPUT]).astype('float32'), name='W1')
B1 = tf.Variable(np.random.normal(size=[NUM_OUTPUT]).astype('float32'), name='B1')
y_cap = tf.add(tf.tensordot(baseOutput, W1, axes=1), B1, name='y_cap_tata')
self.output = y_cap
self.graphCreated = True
def restoreExtendedGraph(self, graph, *args, **kwargs):
y_cap = graph.get_tensor_by_name('y_cap_tata:0')
self.output = y_cap
self.graphCreated = True
def feedDictFunc(self, keep_prob=None, inference=False, **kwargs):
if inference is False:
feedDict = {self._emiGraph.keep_prob: keep_prob}
else:
feedDict = {self._emiGraph.keep_prob: 1.0}
return feedDict
EMI_BasicLSTM._createExtendedGraph = createExtendedGraph
EMI_BasicLSTM._restoreExtendedGraph = restoreExtendedGraph
if USE_DROPOUT is True:
EMI_Driver.feedDictFunc = feedDictFunc
inputPipeline = EMI_DataPipeline(NUM_SUBINSTANCE, NUM_TIMESTEPS, NUM_FEATS, NUM_OUTPUT)
emiLSTM = EMI_BasicLSTM(NUM_SUBINSTANCE, NUM_HIDDEN, NUM_TIMESTEPS, NUM_FEATS,
forgetBias=FORGET_BIAS, useDropout=USE_DROPOUT)
emiTrainer = EMI_Trainer(NUM_TIMESTEPS, NUM_OUTPUT, lossType='xentropy',
stepSize=LEARNING_RATE)
tf.reset_default_graph()
g1 = tf.Graph()
with g1.as_default():
# Obtain the iterators to each batch of the data
x_batch, y_batch = inputPipeline()
# Create the forward computation graph based on the iterators
y_cap = emiLSTM(x_batch)
# Create loss graphs and training routines
emiTrainer(y_cap, y_batch)
#------------------------------END OF COMPUTATION GRAPH------------------------------#
#-------------------------------------EMI DRIVER-------------------------------------#
with g1.as_default():
emiDriver = EMI_Driver(inputPipeline, emiLSTM, emiTrainer)
emiDriver.initializeSession(g1)
y_updated, modelStats = emiDriver.run(numClasses=NUM_OUTPUT, x_train=x_train,
y_train=y_train, bag_train=BAG_TRAIN,
x_val=x_val, y_val=y_val, bag_val=BAG_VAL,
numIter=NUM_ITER, keep_prob=KEEP_PROB,
numRounds=NUM_ROUNDS, batchSize=BATCH_SIZE,
numEpochs=NUM_EPOCHS, modelPrefix=MODEL_PREFIX,
fracEMI=0.5, updatePolicy='top-k', k=1)
#-------------------------------END OF EMI DRIVER-------------------------------------#
#-----------------------------------EARLY SAVINGS-------------------------------------#
"""
Early Prediction Policy: We make an early prediction based on the predicted classes
probability. If the predicted class probability > minProb at some step, we make
a prediction at that step.
"""
def earlyPolicy_minProb(instanceOut, minProb, **kwargs):
assert instanceOut.ndim == 2
classes = np.argmax(instanceOut, axis=1)
prob = np.max(instanceOut, axis=1)
index = np.where(prob >= minProb)[0]
if len(index) == 0:
assert (len(instanceOut) - 1) == (len(classes) - 1)
return classes[-1], len(instanceOut) - 1
index = index[0]
return classes[index], index
def getEarlySaving(predictionStep, numTimeSteps, returnTotal=False):
predictionStep = predictionStep + 1
predictionStep = np.reshape(predictionStep, -1)
totalSteps = np.sum(predictionStep)
maxSteps = len(predictionStep) * numTimeSteps
savings = 1.0 - (totalSteps / maxSteps)
if returnTotal:
return savings, totalSteps
return savings
#--------------------------------END OF EARLY SAVINGS---------------------------------#
#----------------------------------------BEST MODEL-----------------------------------#
k = 2
predictions, predictionStep = emiDriver.getInstancePredictions(x_test, y_test, earlyPolicy_minProb,
minProb=0.99, keep_prob=1.0)
bagPredictions = emiDriver.getBagPredictions(predictions, minSubsequenceLen=k, numClass=NUM_OUTPUT)
print('Accuracy at k = %d: %f' % (k, np.mean((bagPredictions == BAG_TEST).astype(int))))
mi_savings = (1 - NUM_TIMESTEPS / ORIGINAL_NUM_TIMESTEPS)
emi_savings = getEarlySaving(predictionStep, NUM_TIMESTEPS)
total_savings = mi_savings + (1 - mi_savings) * emi_savings
print('Savings due to MI-RNN : %f' % mi_savings)
print('Savings due to Early prediction: %f' % emi_savings)
print('Total Savings: %f' % (total_savings))
#Store in the dictionary.
lstm_dict["k"] = k
lstm_dict["accuracy"] = np.mean((bagPredictions == BAG_TEST).astype(int))
lstm_dict["total_savings"] = total_savings
lstm_dict["y_test"] = BAG_TEST
lstm_dict["y_pred"] = bagPredictions
# A slightly more detailed analysis method is provided.
df = emiDriver.analyseModel(predictions, BAG_TEST, NUM_SUBINSTANCE, NUM_OUTPUT)
print (tabulate(df, headers=list(df.columns), tablefmt='grid'))
lstm_dict["detailed analysis"] = df
#----------------------------------END OF BEST MODEL-----------------------------------#
#----------------------------------PICKING THE BEST MODEL------------------------------#
devnull = open(os.devnull, 'r')
for val in modelStats:
round_, acc, modelPrefix, globalStep = val
emiDriver.loadSavedGraphToNewSession(modelPrefix, globalStep, redirFile=devnull)
predictions, predictionStep = emiDriver.getInstancePredictions(x_test, y_test, earlyPolicy_minProb,
minProb=0.99, keep_prob=1.0)
bagPredictions = emiDriver.getBagPredictions(predictions, minSubsequenceLen=k, numClass=NUM_OUTPUT)
print("Round: %2d, Validation accuracy: %.4f" % (round_, acc), end='')
print(', Test Accuracy (k = %d): %f, ' % (k, np.mean((bagPredictions == BAG_TEST).astype(int))), end='')
print('Additional savings: %f' % getEarlySaving(predictionStep, NUM_TIMESTEPS))
#-------------------------------END OF PICKING THE BEST MODEL--------------------------#
return lstm_dict
def experiment_generator(params, path, model = 'lstm'):
if (model == 'lstm'): return lstm_experiment_generator(params, path)
elif (model == 'fastgrnn'): return fastgrnn_experiment_generator(params, path)
elif (model == 'gru'): return gru_experiment_generator(params, path)
elif (model == 'baseline'): return baseline_experiment_generator(params, path)
return
import tensorflow as tf
# cd /home/sf/data/DREAMER/Arousal
# +
# Baseline EMI-LSTM
dataset = 'DREAMER_Arousal_128'
path = '/home/sf/data/DREAMER/Arousal/'+ '/%d_%d/' % (subinstanceLen, subinstanceStride)
#Choose model from among [lstm, fastgrnn, gru]
model = 'lstm'
# Dictionary to set the parameters.
params = {
"NUM_HIDDEN" : 128,
"NUM_TIMESTEPS" : 48, # subinstance length
"ORIGINAL_NUM_TIMESTEPS" : 128,
"NUM_FEATS" : 16,
"FORGET_BIAS" : 1.0,
"NUM_OUTPUT" : 5,
"USE_DROPOUT" : 1, # '1' -> True, '0' -> False
"KEEP_PROB" : 0.75,
"PREFETCH_NUM" : 5,
"BATCH_SIZE" : 32,
"NUM_EPOCHS" : 2,
"NUM_ITER" : 4,
"NUM_ROUNDS" : 10,
"LEARNING_RATE" : 0.001,
"FRAC_EMI" : 0.5,
"MODEL_PREFIX" : dataset + '/model-' + str(model)
}
#Preprocess data, and load the train,test and validation splits.
lstm_dict = lstm_experiment_generator(params, path)
#Create the directory to store the results of this run.
# dirname = ""
# dirname = "./Results" + ''.join(dirname) + "/"+dataset+"/"+model
# pathlib.Path(dirname).mkdir(parents=True, exist_ok=True)
# print ("Results for this run have been saved at" , dirname, ".")
dirname = "" + model
pathlib.Path(dirname).mkdir(parents=True, exist_ok=True)
print ("Results for this run have been saved at" , dirname, ".")
now = datetime.datetime.now()
filename = list((str(now.year),"-",str(now.month),"-",str(now.day),"|",str(now.hour),"-",str(now.minute)))
filename = ''.join(filename)
#Save the dictionary containing the params and the results.
pkl.dump(lstm_dict,open(dirname + "/lstm_dict_" + filename + ".pkl",mode='wb'))
| DREAMER/DREAMER_Arousal_LSTM_48_16.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
#hide
from emdesigner.core import *
# # EMDesigner
#
# > This library is an assistant to electric machine designers. It assists following machine designs through analytical equations.
# * C - core inductor
# * surface mounted permanent magnet synchronous motor
# ## Install
# `pip install emdesigner`
# ## How to use
# To be included:
1+1
| index.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import numpy as np
from pylab import *
import nibabel as nib
def find_contour(img, x, y, used_coords, loop_coords, val):
for ix in range(x-1, x+2):
for iy in range(y-1, y+2):
if ix < 0 or iy < 0 or iy >= len(img) or ix >= len(img[0]):
continue
if img[iy][ix] != 0.0:
if iy not in used_coords or ix not in used_coords[iy]:
loop_coords.append([ix,iy])
val = img[iy][ix]
#print "NESTED (%s, %s), LOOP: %s" % (ix, iy, loop_coords)
# add coord to used_coords
if iy not in used_coords:
used_coords[iy] = {}
if ix not in used_coords[iy]:
used_coords[iy][ix] = True
used_coords, loop_coords, val = find_contour(img, ix, iy, used_coords, loop_coords, val)
return used_coords, loop_coords, val
# ## Test find_contour logic on example grid
# +
test = [[0,0,0,0,0],[0,0,1,0,0],[0,1,0,1,0],[0,0,1,0,0], [0,0,0,0,0]]
test_loop_coords = []
test_used_coords = {}
for y, rows in enumerate(test):
for x, v in enumerate(rows):
#print "START (%s, %s)" % (x,y)
test_used_coords, test_loop_coords, test_loop_category = find_contour(test, x, y, test_used_coords, test_loop_coords, None)
print "FINAL", test_used_coords, test_loop_coords
# -
# ## Find contours on nii image
# Load nii data from disk
nii_obj=nib.load('./segmentation-010.nii')
nii_data = nii_obj.get_fdata()
# Convert nii matrix into per-dicom grid structure
img_data = [[[[None] for x in range(322)] for y in range(1086)] for z in range(20)]
for x, rows in enumerate(nii_data):
for y, depths in enumerate(rows):
for z, v in enumerate(depths):
if v != 0.0:
print z, y, x, v
img_data[z][y][x] = v
# +
all_loops = []
for img in img_data:
loop_coords = []
used_coords = {}
for y, rows in enumerate(img):
for x, v in enumerate(rows):
if v != 0.0:
if y not in used_coords or x not in used_coords[y]:
# loop around for pixels
used_coords, single_loop_coords, loop_category = find_loop(img, x, y, used_coords, [], None)
loop_coords.append({'category': loop_category, 'coords': single_loop_coords})
all_loops.append(loop_coords)
# -
# ## Dump contour data into json file on disk
d = {'dicoms':[]}
for i, img_loops in enumerate(all_loops):
d['dicoms'].append({'contours':img_loops})
with open('contour-data.json', 'w') as outfile:
json.dump(d, outfile)
# ## Generating image display with augmented contours
# +
import matplotlib.pyplot as plt
import matplotlib.patches as patches
import pydicom
from pydicom.data import get_testdata_files
import json, math
import numpy as np
import matplotlib
from shapely.geometry.polygon import LinearRing, Polygon
from matplotlib.collections import PatchCollection
# -
with open('./contours-with-categories.json') as json_file:
contour_data = json.load(json_file)
# ## Display augmented dicoms in a grid
# +
images = [
'./images/001-02.0.dcm', './images/002-06.0.dcm', './images/003-07.0.dcm',
'./images/004-08.0.dcm', './images/005-09.0.dcm', './images/006-10.0.dcm',
'./images/007-11.0.dcm', './images/008-12.0.dcm', './images/009-13.0.dcm',
'./images/010-14.0.dcm', './images/011-15.0.dcm', './images/012-16.0.dcm',
'./images/013-17.0.dcm', './images/014-18.0.dcm', './images/015-19.0.dcm',
'./images/016-20.0.dcm', './images/017-21.0.dcm', './images/018-22.0.dcm',
'./images/019-23.0.dcm', './images/020-24.0.dcm']
n_rows = math.ceil(len(images)*1.0/3)
plt.figure(figsize=(16, 16*n_rows))
for i, img in enumerate(images):
ax = plt.subplot(n_rows, 3, i+1)
ds = pydicom.dcmread(img)
ax.imshow(ds.pixel_array, cmap=plt.cm.bone, vmin=0, vmax=255)
contours = contour_data['dicoms'][i]['contours']
for contour in contours:
if len(contour['coords']) > 2:
poly = Polygon(contour['coords'])
x,y = poly.exterior.xy
ax.plot(x, y, color='#ff0000', alpha=0.4,
linewidth=1, solid_capstyle='round', zorder=2)
plt.suptitle('Tumor Segmentations')
plt.show()
# -
# ## Export augmented dicoms to png files on disk
# +
images = [
'./images/001-02.0.dcm', './images/002-06.0.dcm', './images/003-07.0.dcm',
'./images/004-08.0.dcm', './images/005-09.0.dcm', './images/006-10.0.dcm',
'./images/007-11.0.dcm', './images/008-12.0.dcm', './images/009-13.0.dcm',
'./images/010-14.0.dcm', './images/011-15.0.dcm', './images/012-16.0.dcm',
'./images/013-17.0.dcm', './images/014-18.0.dcm', './images/015-19.0.dcm',
'./images/016-20.0.dcm', './images/017-21.0.dcm', './images/018-22.0.dcm',
'./images/019-23.0.dcm', './images/020-24.0.dcm']
plt.figure(figsize=(2, 16))
for i, img in enumerate(images):
fig,ax = plt.subplots(1)
fig.set_size_inches(18.5, 10.5)
ds = pydicom.dcmread(img)
ax.imshow(ds.pixel_array, cmap=plt.cm.bone, vmin=0, vmax=255)
ax.axes.get_xaxis().set_visible(False)
ax.axes.get_yaxis().set_visible(False)
ax.set_frame_on(False)
contours = contour_data['dicoms'][i]['contours']
for contour in contours:
if len(contour['coords']) > 2:
poly = Polygon(contour['coords'])
x,y = poly.exterior.xy
ax.plot(x, y, color='#ff0000', alpha=0.4,
linewidth=1, solid_capstyle='round', zorder=2)
plt.axis('off')
plt.savefig('./frames/%02d.png' % (i), transparent=False, bbox_inches='tight', pad_inches=0)
plt.show()
# -
# ## Convert png files to mp4
#
# In terminal, run `ffmpeg -framerate 2/1 -i ./frames/%02d.png -c:v libx264 -r 30 demo-video.mp4`
| Generate Visualizations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from __future__ import print_function
import os
import pandas as pd
# %matplotlib inline
from matplotlib import pyplot as plt
#Load the dataset into a pandas.DataFrame
ibm_df = pd.read_csv('datasets/ibm-common-stock-closing-prices.csv')
ibm_df.index = ibm_df['Date']
#Let's find out the shape of the DataFrame
print('Shape of the dataframe:', ibm_df.shape)
#Let's see the top 10 rows
ibm_df.head(10)
#Rename the second column
ibm_df.rename(columns={'IBM common stock closing prices': 'Close_Price'},
inplace=True)
ibm_df.head()
#remove missing values
missing = (pd.isnull(ibm_df['Date'])) & (pd.isnull(ibm_df['Close_Price']))
print('No. of rows with missing values:', missing.sum())
ibm_df = ibm_df.loc[~missing, :]
#To illustrate the idea of moving average we compute a weekly moving average taking
#a window of 5 days instead of 7 days because trading happens only during the weekdays.
ibm_df['5-Day Moving Avg'] = ibm_df['Close_Price'].rolling(5).mean()
fig = plt.figure(figsize=(5.5, 5.5))
ax = fig.add_subplot(2,1,1)
ibm_df['Close_Price'].plot(ax=ax, color='b')
ax.set_title('IBM Common Stock Close Prices during 1962-1965')
ax = fig.add_subplot(2,1,2)
ibm_df['5-Day Moving Avg'].plot(ax=ax, color='r')
ax.set_title('5-day Moving Average')
plt.tight_layout(pad=0.4, w_pad=0.5, h_pad=2.0)
plt.savefig('plots/ch2/B07887_02_14.png', format='png', dpi=300)
# +
#Calculate the moving averages using 'rolling' and 'mean' functions
MA2 = ibm_df['Close_Price'].rolling(window=2).mean()
TwoXMA2 = MA2.rolling(window=2).mean()
MA4 = ibm_df['Close_Price'].rolling(window=4).mean()
TwoXMA4 = MA4.rolling(window=2).mean()
MA3 = ibm_df['Close_Price'].rolling(window=3).mean()
ThreeXMA3 = MA3.rolling(window=3).mean()
# +
#Let's remove NaN from the above variables
MA2 = MA2.ix[~pd.isnull(MA2)]
TwoXMA2 = TwoXMA2.ix[~pd.isnull(TwoXMA2)]
MA4 = MA4.ix[~pd.isnull(MA4)]
TwoXMA4 = TwoXMA4.ix[~pd.isnull(TwoXMA4)]
MA3 = MA3.ix[~pd.isnull(MA3)]
ThreeXMA3 = TwoXMA4.ix[~pd.isnull(ThreeXMA3)]
# +
f, axarr = plt.subplots(3, sharex=True)
f.set_size_inches(5.5, 5.5)
ibm_df['Close_Price'].iloc[:45].plot(color='b', linestyle = '-', ax=axarr[0])
MA2.iloc[:45].plot(color='r', linestyle = '-', ax=axarr[0])
TwoXMA2.iloc[:45].plot(color='r', linestyle = '--', ax=axarr[0])
axarr[0].set_title('2 day MA & 2X2 day MA')
ibm_df['Close_Price'].iloc[:45].plot(color='b', linestyle = '-', ax=axarr[1])
MA4.iloc[:45].plot(color='g', linestyle = '-', ax=axarr[1])
TwoXMA4.iloc[:45].plot(color='g', linestyle = '--', ax=axarr[1])
axarr[1].set_title('4 day MA & 2X4 day MA')
ibm_df['Close_Price'].iloc[:45].plot(color='b', linestyle = '-', ax=axarr[2])
MA3.iloc[:45].plot(color='k', linestyle = '-', ax=axarr[2])
ThreeXMA3.iloc[:45].plot(color='k', linestyle = '--', ax=axarr[2])
axarr[2].set_title('3 day MA & 3X 3day MA')
plt.savefig('plots/ch2/B07887_02_15.png', format='png', dpi=300)
| time series regression/autocorelation, mov avg etc/Moving_Averages.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: py_37_env
# language: python
# name: py_37_env
# ---
import ipyvolume as ipv
import numpy as np
import ipywidgets
ipywidgets.FloatSlider()
x, y, z = np.random.random((3, 10000))
ipv.quickscatter(x, y, z, size=1, marker="sphere")
| inprogress/ML_Med_ipyvol_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="TqvHo5stxc12" colab_type="code" colab={}
import numpy as np
# + [markdown] id="y1fW2YtgyflU" colab_type="text"
# #Sigmoid(x) = 1/(1+e^(-x))
# + id="6Ofksj1xye87" colab_type="code" colab={}
def sigmoid(x):
return 1/(1+np.exp(-x))
# + [markdown] id="stpXAwZcyju0" colab_type="text"
# #d/dx(Sigmoid(x)) = x(1-x)
# + id="SwiFVt_eyh9U" colab_type="code" colab={}
def sigmoid_derivative(x):
return x*(1-x)
# + [markdown] id="pqJVNEo1zum6" colab_type="text"
# ### **Making a simple dataset**
# + [markdown] id="OUOJ08tOzs6r" colab_type="text"
# #inputs
# + id="9_AR6Admym1l" colab_type="code" colab={}
training_inputs = np.array([[0,0,1],
[1,1,1],
[1,0,1],
[0,1,1]])
# + [markdown] id="k-Qo0BuByuVM" colab_type="text"
# #outputs required
# + id="gu0YCFIhyrXE" colab_type="code" colab={}
training_outputs = np.array([[0,1,1,0]]).T
# + [markdown] id="sfHLxdzEzlQb" colab_type="text"
# #Defining synaptics weights
# + id="DsZJpWuryvTz" colab_type="code" colab={}
np.random.seed(1)
synaptic_weights = 2 * np.random.random((3,1)) - 1
# + [markdown] id="G3d5szVAzefa" colab_type="text"
# #synaptic weights before training
# + id="PK5AJwrsziO8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="ef44886d-d19a-4035-b467-96479f03eebe"
print("Random starting synaptic weights: ")
print(synaptic_weights)
# + [markdown] id="qefaIq_Zy8u8" colab_type="text"
# #For training the machine on weightage and the output will be closer as the no. of iterations increases
# + id="SFlBHMaYy5l6" colab_type="code" colab={}
for iteration in range(100000):
input_layer = training_inputs
output = sigmoid(np.dot(input_layer, synaptic_weights))
error = training_outputs - output
adjusment = error * sigmoid_derivative(output)
synaptic_weights += np.dot(input_layer.T, adjusment)
# + [markdown] id="5kg-TNnCzL4z" colab_type="text"
# #Synaptic weights after training
# + id="t2ScTCrNzJDL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="b1110cfd-85ed-455f-8321-0744755332b6"
print('Synaptic Weights after training')
print(synaptic_weights)
# + [markdown] id="ndlVJoQlzRlq" colab_type="text"
# #predicted outputs
# + id="Kmg__FlKzSSO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 102} outputId="10cf44b7-0aac-44ce-ff7c-c8878fd04b5e"
print('Output after training: ')
print(output)
| basic_perceptron.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: RAPIDS Stable
# language: python
# name: rapids-stable
# ---
# ## Tutorial 3: Fuzzy matching and bucketing across events using UMAP and hypergraphs
#
# Part of the [Python security RAPIDS GPU & graph one-liners](Tutorial_0_Intro.ipynb)
#
# All GPU Python data science tutorials: [RAPIDS Academy github](https://github.com/RAPIDSAcademy/rapidsacademy)
#
#
# We often need to map out events, such as for bucketing events into incidents, or finding coordinated bot activity. This can be tough when there are many events, and the correlations are on fuzzy values such as time stamps.
#
# This tutorial is inspired by by <NAME> (Microsoft)'s GraphThePlanet talk on [Intrusion detection with graphs](https://www.slideshare.net/MattSwann1). The original recorded demo used the technique for mapping COVID misinformation bot events on Twitter. We redo it with Project Mordor elastic search APT simulation data.
#
# The analysis walks through:
#
# * Fuzzy match: UMAP nearest-neighbor by generically comparing combinations of many event attributes, e.g., rough time, bytes, and specific username (`cuml`'s `umap`)
# * Exact match: Link on a few individual attributes, such as username (`cudf`, `graphistry`)
# * Visualize (`graphistry`)
#
#
#
# ### Setup
#
# To setup and test your RAPIDS environment (cudf, blazingsql, graphistry, ...), use the [setup guide](setup.ipynb).
# ### Get data
# ! wget https://raw.githubusercontent.com/hunters-forge/mordor/master/datasets/small/windows/privilege_escalation/empire_invoke_runas.tar.gz \
# && ls -alh empire_invoke_runas.tar.gz \
# && tar -xvf empire_invoke_runas.tar.gz \
# && ls -alh empire_invoke_runas_2019-05-18204300.json
# +
# #! rm -f empire_invoke_runas.tar.gz && rm -f empire_invoke_runas_2019-05-18204300.json
# -
# ! head -n 3 empire_invoke_runas_2019-05-18204300.json
# ## Load data
#
# Note that the RAPIDS json logs reader is *not* yet GPU-accelerated, so we use pandas. We recommend using accelerated formats like parquet or csv when possible.
# +
import cudf, cuml, graphistry, json, pandas as pd
import logging
logger = logging.getLogger()
from pandas.api.types import is_datetime64_any_dtype
# +
### Get free Graphistry Hub account & creds at https://www.graphistry.com/get-started
### First run: set to True and fill in creds
### Future runs: set to False and erase your creds
### When done: delete graphistry.json
if False:
#creds = {'token': '...'}
creds = {'username': '***', 'password': '***'}
with open('graphistry.json', 'w') as outfile:
json.dump(creds, outfile)
with open('graphistry.json') as f:
creds = json.load(f)
graphistry.register(
api=3, key='', protocol='https', server='hub.graphistry.com',
**creds)
# -
pd.set_option('display.max_columns', 100)
logger.setLevel(logging.INFO) #DEBUG}
# +
# %%time
raw_df = pd.read_json('./empire_invoke_runas_2019-05-18204300.json',
lines=True)
print('shape', raw_df.shape)
print('columns', raw_df.columns)
raw_df.sample(3)
# -
# ## Flatten & type
#
# Need each column to be a primitive type like a time, str, or int.
#
# * Flatten every json col into multiple primitive cols
# * Time str -> time
#
# +
import ast
def str_to_json(s):
if pd.isna(s):
return {}
try:
return ast.literal_eval(str(s))
except:
#logger.error('oops', exc_info=True)
#logger.error('str: %s', s)
return {}
def flatten_json_str_cols(df, cols):
out_df = df.copy(deep=False)
for col in cols:
flattened_df = pd.io.json.json_normalize(df[col].apply(str_to_json))
#out_df.drop(labels=[col], inplace=True)
logger.debug('%s -> %s', col, flattened_df.columns)
for c in flattened_df.columns:
#print('adding', col, c)
out_df[f'{col}_{c}'] = flattened_df[c]
return out_df
# +
df = raw_df.copy(deep=False)
df['@timestamp'] = pd.to_datetime(df['@timestamp'])
df = flatten_json_str_cols(df, ['event_data', 'host', 'user'])
df['event_data_UtcTime'] = pd.to_datetime(df['event_data_UtcTime'])
print(raw_df.shape, '->', df.shape)
df.sample(3)
# -
{x: len(df[x].astype(str).unique()) for x in df.columns}
# ## Encode columns as values between 0 and 1
#
# * Turns smaller string columns into multiple 0/1 indicator columns ("is_x", "is_y", ...) otherwise drops
# * Good to play with
# +
#pdf -> gdf['id', col_0:(0,1), ...]
def quick_encode(df, cols=None, onehot_threshold=None, id_col=None, normalize=True):
if onehot_threshold is None:
onehot_threshold = min(round(0.1 * len(df)), 100)
logger.debug('threshold on %s -> %s', len(df), onehot_threshold)
if cols is None:
cols = df.columns
df_primitive = df.copy(deep=False)
for col in df.columns:
if df[col].dtype.name == 'object':
df[col] = df[col].astype(str)
gdf = cudf.from_pandas(df)
out_gdf = gdf.copy(deep=False)
if id_col is None:
out_gdf = out_gdf[[]].reset_index().rename(columns={'index': 'id'}, copy=False)
else:
out_gdf = out_gdf[[]]
out_gdf['id'] = gdf[id_col]
#logger.debug('starting cols: %s', out_gdf.columns)
for col in cols:
print('col', col)
if col != id_col and col != 'id':
unique = gdf[col].unique()
if len(unique) < 2:
logger.debug('%s boring', col)
continue
elif gdf[col].dtype.name == 'object':
if len(unique) < onehot_threshold:
logger.debug('%s onehot', col)
out_gdf[col] = gdf[col]
out_gdf = out_gdf.one_hot_encoding(column=col, prefix='onehot_%s' % col, cats=unique)
out_gdf.drop(labels=[col], inplace=True)
else:
logger.debug('%s str -> many uniques; dropping %s', col, len(unique))
continue
elif is_datetime64_any_dtype(df[col]):
logger.debug('%s date->int64', col)
out_gdf[col] = gdf[col].astype('int64')
else:
logger.debug('%s numeric, preserve %s', col, gdf[col].dtype.name)
out_gdf[col] = gdf[col]
continue
else:
logger.debug('%s skip id col', col)
continue
if normalize:
for col in out_gdf.columns:
if col != 'id_col' and col != 'id':
ok = False
out_gdf[col] = out_gdf[col].fillna(0.0)
try:
out_gdf[col] = out_gdf[col].scale()
ok=True
except:
if len(out_gdf[col].unique()) > 0:
mx = out_gdf[col].max()
mn = out_gdf[col].min()
w = mx - mn
if w > 0.0:
out_gdf[col] = out_gdf[col] - out_gdf[col].min()
out_gdf[col] = out_gdf[col] / w
ok = True
if not ok:
out_gdf.drop(labels=[col], inplace=True)
return out_gdf
df_encoded = quick_encode(df).to_pandas()
print('encoded', df_encoded.columns.tolist())
print('new shape', df.shape, '->', df_encoded.shape)
df_encoded.sample(3)
# -
# ### UMAP + K-nn helper
# +
def knn_adjacency_to_edgelist(a_gdf, w_gdf, index_to_id_gdf, drop_far_neighbors=True, drop_far_neighbors_std=1):
#get rid of spurious children seen in small clusters:
#assuming n is "good", threshold as nth-furthest's (avgd dist + 1 stddev)
dist_thresh = None
scalar = 1.0 if drop_far_neighbors else 0.0 #hack to ensure not dropping
if drop_far_neighbors:
dist_thresh = w_gdf[ len(w_gdf.columns) - 1 ].mean() + w_gdf[ len(w_gdf.columns) - 1 ].std() * drop_far_neighbors_std
print('dropping neighbors > %s away (%s neighbor dist mean + std' % (dist_thresh, len(a_gdf.columns)) )
else:
dist_thresh = w_gdf.max().max() #nop, TODO just skip the filter
print('not dropping neighbors')
edges_gdf = cudf.concat([
a_gdf[ [n] ].assign(knn_d=w_gdf[n], knn_n=n) #[[ n, 'knn_d' ]] \
[ w_gdf[n] * scalar <= dist_thresh ]\
.reset_index().rename(columns={'index': 'from_idx', n: 'to_idx'}) #[[ 'from_index', 'to_index', 'kn__d', 'knn_n']] \
.merge(index_to_id_gdf.rename(columns={'id': 'from'}),
left_on='from_idx',
right_on='index').drop(columns=['index', 'from_idx'])\
.merge(index_to_id_gdf.rename(columns={'id': 'to'}),
left_on='to_idx',
right_on='index').drop(columns=['index', 'to_idx']) #[[ 'from', 'to', 'knn_d', 'knn_n']]
for n in a_gdf.columns
], ignore_index=True)
print('%s edges (dropped %s%% edges)' % (
len(edges_gdf),
round(100.0 * (1.0 - (len(edges_gdf) / (len(a_gdf) * len(a_gdf.columns)))))))
edges_no_self_gdf = edges_gdf[ edges_gdf['from'] != edges_gdf['to'] ]
print('%s edges (dropped %s%% edges)' % (
len(edges_no_self_gdf),
round(100.0 * (1.0 - (len(edges_no_self_gdf) / (len(a_gdf) * len(a_gdf.columns - 1)))))))
return edges_no_self_gdf
def nn(projection_gdf, n_neighbors=20, projection_cols=['x','y']):
nn_model = cuml.NearestNeighbors(n_neighbors=n_neighbors)
nn_model.fit(projection_gdf[projection_cols])
(weights_gdf, adjacency_gdf) = nn_model.kneighbors(projection_gdf[projection_cols])
return (weights_gdf, adjacency_gdf)
def nn_edges(projection_gdf, index_to_id, n_neighbors=20, drop_far_neighbors=True, drop_far_neighbors_std=1, projection_cols=['x','y']):
print('computing nn (%s)' % n_neighbors)
(weights_gdf, adjacency_gdf) = nn(projection_gdf, n_neighbors, projection_cols)
edges_gdf = knn_adjacency_to_edgelist(adjacency_gdf, weights_gdf, index_to_id, drop_far_neighbors, drop_far_neighbors_std)
out = edges_gdf.assign(type='knn_%s' % n_neighbors)
out['weight'] = edges_gdf['knn_d']
return out
##FIXME buggy some reason
def with_nn_edges(g, n_neighbors=20, drop_far_neighbors=True, drop_far_neighbors_std=1):
print('with_nn_edges')
xy_gdf = cudf.from_pandas(g._nodes[['x', 'y']])
index_to_id_gdf = cudf.from_pandas(g._nodes[[g._node]]).reset_index()\
.rename(columns={g._node: 'id'})
edges_gdf = nn_edges(xy_gdf, index_to_id_gdf, n_neighbors, drop_far_neighbors, drop_far_neighbors_std, ['x', 'y'])
print('got %s edges' % len(edges_gdf))
edges_pdf = edges_gdf.to_pandas().rename(columns={
'from': g._source,
'to': g._destination
})
return g.edges(pd.concat([ g._edges, edges_pdf], ignore_index=True, sort=False))
def umap_plot(pdf, id_col=None, fit_opts={}, umap_opts={}, nn_opts=None):
if id_col is None:
pdf = pdf.reset_index()
id_col = 'index'
logger.debug('to gdf')
gdf = cudf.from_pandas(pdf)
reducer = cuml.UMAP(**umap_opts)
logger.debug('fit')
embedding = reducer.fit_transform(gdf.drop(columns=[id_col]),
**fit_opts)
nodes_gdf = gdf.assign(x=embedding[0], y=embedding[1])
logger.debug('graphistry')
g = graphistry.nodes(nodes_gdf.to_pandas())\
.bind(node=id_col)\
.bind(source='s', destination='d').edges(pd.DataFrame({
's': [ gdf[id_col].head(1).tolist()[0] ],
'd': [ gdf[id_col].head(1).tolist()[0] ]
})).settings(url_params={'play': 0, 'strongGravity': True})
logger.debug('nn')
if not (nn_opts is None):
g = with_nn_edges(**{'g': g, **nn_opts})
return g
# -
# ### UMAP
#
# * Place similar events next to one another using x/y coordinates
# * Explicitly connect them with edges
# +
# %%time
g = umap_plot(
df_encoded,
id_col='id',
umap_opts={'n_neighbors': 25, 'repulsion_strength': 5, 'n_epochs': 5000},
nn_opts={'drop_far_neighbors_std': 0.5} #set to None for no edges
)
g = g.nodes(
g._nodes[['id', 'x', 'y']]\
.merge(df.reset_index().rename(columns={'index': 'id'}, copy=False),
on='id', how='left'))
print(len(g._nodes), len(g._edges))
# -
g._nodes.sample(3)
g._edges.sample(3)
# ### Visualize
#
# Optionally override some settings
g.plot()
# ## Create graph with physical edges
#
# Instead of UMAP's fuzzy edges, we can directly map explicit event -> entity edges, such as `(event1) --> (ip) <-- (event2)`
#
# * Defaults to `direct=False` to include events as nodes
# * Set `direct=True` to only show entity nodes without events, e.g., `(ip)--[event1]-->(user_name)`
# +
# %%time
# not GPU accelerated.. yet
hg = graphistry.hypergraph(
g._nodes,
[
'computer_name', 'user_name', 'user_identifier',
'event_data_TargetImage', 'event_data_TargetProcessGUID', 'event_data_TargetObject',
'event_id',
'event_data_param1', 'event_data_param2', 'event_data_param3',
'event_data_DestinationIp', 'event_data_SourceIp'
],
direct=False)
edges_df = hg['graph']._edges
print(edges_df.shape)
print(edges_df.sample(3))
hg['graph'].plot( hg['graph']._edges.sample(1000) )
# +
def drop_supernode_edges(g, max_col_links=None):
if max_col_links is None:
max_col_links = len(g._nodes) / 10
logger.info('max_col_links: %s nodes -> %s max', len(g._nodes), max_col_links)
#df[['hit_s': True, g._source: 'a]]
s_counts = g._edges[ g._source ].value_counts()
s_counts = pd.DataFrame(s_counts[ s_counts > max_col_links ])\
.assign(hit_s=True)\
.reset_index()[['hit_s', 'index']]\
.rename(columns={'index': g._source})[['hit_s', g._source]]
#df[['hit_d': True, g._destination: 'a]]
d_counts = g._edges[ g._destination ].value_counts()
d_counts = pd.DataFrame(d_counts[ d_counts > max_col_links ])\
.assign(hit_d=True)\
.reset_index()[['hit_d', 'index']]\
.rename(columns={'index': g._destination})[['hit_d', g._destination]]
edges2 = g._edges.copy(deep=False)
edges2 = edges2.merge(s_counts, on=g._source, how='left')
edges2 = edges2[ ~ edges2['hit_s'].fillna(False) ]
edges2 = edges2.merge(d_counts, on=g._destination, how='left')
edges2 = edges2[ ~ edges2['hit_d'].fillna(False) ]
return edges2
def drop_nan_hyper_edges(g):
edges = g._edges[ ~ g._edges[g._source].str.contains('::nan') ]
edges = edges[ ~ edges[g._destination].str.contains('::nan') ]
return edges
g2 = hg['graph'].edges(drop_supernode_edges(hg['graph']))
print('dropping supernode edges', hg['graph']._edges.shape, '->', g2._edges.shape)
g2 = g2.edges(drop_nan_hyper_edges(g2))
print('dropping nan edges->', g2._edges.shape)
g2.plot()
# -
# ## Combine graph of linked events with linked entities
#
# Final task: try to merge the two above graphs by creating a new graph based on dataframes `g._nodes`, `g._edges`, `g2._nodes`, and `g2._edges`
#
#
# ## Next steps
#
# Part of the [Python security RAPIDS GPU & graph one-liners](Tutorial_0_Intro.ipynb)
#
# All GPU Python data science tutorials: [RAPIDS Academy github](https://github.com/RAPIDSAcademy/rapidsacademy)
| tutorials/security/tour/Tutorial_3_incident_umap_knn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %reload_ext autoreload
# %autoreload 2
# +
import warnings
warnings.filterwarnings('ignore')
from IPython.display import IFrame, clear_output
# +
import os
import time
import logging
from functools import wraps
import pickle
import re
import pandas as pd
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
import plotly.graph_objs as go
import plotly.express as px
from numpy import hstack
from IPython.display import IFrame
from plotly.offline import init_notebook_mode
init_notebook_mode(connected=True)
from plot_utils import *
# +
from sklearn.ensemble import RandomForestClassifier, ExtraTreesClassifier, AdaBoostClassifier, GradientBoostingClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.linear_model import LogisticRegression
from xgboost import XGBClassifier
from lightgbm import LGBMClassifier
from catboost import CatBoostClassifier
from sklearn.preprocessing import MinMaxScaler, StandardScaler, RobustScaler
from sklearn.feature_selection import SelectKBest, f_classif, chi2, mutual_info_classif
from sklearn.utils import shuffle
from sklearn.model_selection import KFold, StratifiedKFold, RepeatedStratifiedKFold, cross_val_score, GridSearchCV, train_test_split
from skopt import BayesSearchCV
from skopt.space import Integer, Real, Categorical
#from sklearn.pipeline import Pipeline
from sklearn.metrics import accuracy_score
# -
folder_path = 'C:/Users/yaass/OneDrive/Bureau/Parser/ransom_datasets'
from helpers import *
from learners import *
def batch_feature_selection(categories,
k_range = [100, 1001],
step=100,
scoring='accuracy',
dataset_prefix='encoded_nested_fileops_',
dataset='file_operations',
label='sublabel',
with_smote=True):
#initialization
k_range_ = range(k_range[0], k_range[1], step)
summaries = dict()
figure_path = 'figures/feature-selection/ransomware/' + dataset + '_' + scoring + '_step' + str(step) + '.html'
#loop through datasets
for category in categories:
#load data
print('-------------------------------------------------')
print(category.capitalize(), ':\n')
if dataset_prefix is not None:
file_name = dataset_prefix + category + '.pkl.gz'
else:
file_name = 'fileops_' + category + '_nested_files.pkl.gz'
df = get_data(file_path = os.path.join(folder_path, file_name), compression='gzip')
#print class distribution
class_distribution(df, label=label)
#build a model with smote
steps = []
if with_smote:
smote_params = {
'category' : 'adaptive',
'over_strategy' : 0.5,
'under_strategy' : 0.8,
'k_neighbors' : 5,
}
X, y = get_X_y(df, label=label)
steps.extend(smote(X, y, **smote_params, fit=False))
steps.append(('classifier', RandomForestClassifier()))
pipe = Pipeline(steps=steps)
#perform grid-search
scores, summary = search_feature_importances(df = df,
k_values = k_range_,
model=pipe,
scoring=scoring,
verbose=False,
label=label)
#append summary results
summaries[category] = summary
#plot scores
fig = plot_evaluation_boxplots(list(scores.values()),
list(scores.keys()),
title='Performance per selected ' + category.capitalize() + ' ' + dataset + ' features [ ' + scoring.capitalize() + '=f(k) ]',
y_axis=scoring.capitalize(),
x_axis='k',
showlegend=False)
#save plot to html
save_figures_to_html(figure_path, [fig])
print()
#compile summary results and save them
overall_summary = summaries_to_df(summaries,
k_range=k_range_,
path='selected-features/k-search-summary/ransomware',
file=dataset + '_' + scoring + '_step' + str(step) + '.csv')
# ## API Calls
categories = ['counts']
batch_feature_selection(categories,
k_range = [100, 301],
step=25,
scoring='accuracy',
dataset_prefix='apistats_',
dataset='apistats',
label='sublabel')
pd.read_csv('selected-features/k-search-summary/ransomware/apistats_accuracy_step25.csv', index_col=0)
# +
selected_k = [250]
for category, k in zip(categories, selected_k):
#load data
file_name = 'apistats_' + category + '.pkl.gz'
df = get_data(file_path = os.path.join(folder_path, file_name), compression='gzip')
#build model
steps = []
smote_params = {
'category' : 'adaptive',
'over_strategy' : 0.5,
'under_strategy' : 0.8,
'k_neighbors' : 5,
}
X, y = get_X_y(df, label='sublabel')
steps.extend(smote(X, y, **smote_params, fit=False))
steps.append(('classifier', RandomForestClassifier()))
pipe = Pipeline(steps=steps)
#save k selected column names
save_selected_features(df = df,
k = k,
model=pipe,
prefix = 'apistats_' + category,
path='selected-features/ransomware',
file= 'apistats_' + category + '.pkl',
label = 'sublabel')
# -
# ## Registry Key Operations
categories = ['opened', 'read', 'written', 'deleted']
batch_feature_selection(categories,
k_range = [100, 1001],
step=50,
scoring='accuracy',
dataset_prefix=None,
dataset='regkeys_operations',
label='sublabel')
pd.read_csv('selected-features/k-search-summary/ransomware/regkeys_operations_accuracy_step50.csv', index_col=0)
# Although some k values gives the highest accuracy, if we take into account **minimizing model complexity and the variance on the accuracy results**, we'll opt for the following k values: <br/><br/>
# **Opened regkeys :** k=150 <br/>
# **Read regkeys :** k=850 <br/>
# **Written regkeys :** k=100 <br/>
# **Deleted regkeys :** k=50 <br/>
#
# **Total regkeys operations selected columns =** 1150 features
# +
selected_k = [150, 850, 100, 50]
for category, k in zip(categories, selected_k):
#load data
file_name = 'regkeys_' + category + '_nested_keys.pkl.gz'
df = get_data(file_path = os.path.join(folder_path, file_name), compression='gzip')
#build model
steps = []
smote_params = {
'category' : 'adaptive',
'over_strategy' : 0.5,
'under_strategy' : 0.8,
'k_neighbors' : 5,
}
X, y = get_X_y(df, label='sublabel')
steps.extend(smote(X, y, **smote_params, fit=False))
steps.append(('classifier', RandomForestClassifier()))
pipe = Pipeline(steps=steps)
#save k selected column names
save_selected_features(df = df,
k = k,
model=pipe,
prefix = 'regkeys_' + category,
path='selected-features/ransomware',
file= 'regkeys_' + category + '_nested_keys.pkl',
label='sublabel')
# -
# ## File Operations
categories = ['opened', 'exists', 'read', 'written', 'created', 'deleted', 'failed', 'recreated']
batch_feature_selection(categories,
k_range = [100, 1001],
step=50,
scoring='accuracy',
dataset_prefix=None,
dataset='file_operations',
label='sublabel')
pd.read_csv('selected-features/k-search-summary/ransomware/file_operations_accuracy_step50.csv', index_col=0)
# Although some k values gives the highest accuracy, if we take into account **minimizing model complexity and the variance on the accuracy results**, we'll opt for the following k values: <br/><br/>
# **Opened files :** k=500 <br/>
# **Exists files :** k=100 <br/>
# **Read files :** k=100 <br/>
# **Written files :** k=300 <br/>
# **Created files :** k=300 <br/>
# **Deleted files :** k=200 <br/>
# **Failed files :** k=100 <br/>
# **Recreated files :** k=50 <br/>
#
# **Total file operations selected columns =** 1650 features
# +
selected_k = [500, 100, 100, 300, 300, 200, 100, 50]
for category, k in zip(categories, selected_k):
#load data
file_name = 'fileops_' + category + '_nested_files.pkl.gz'
df = get_data(file_path = os.path.join(folder_path, file_name), compression='gzip')
#build model
steps = []
smote_params = {
'category' : 'adaptive',
'over_strategy' : 0.5,
'under_strategy' : 0.8,
'k_neighbors' : 5,
}
X, y = get_X_y(df, label='sublabel')
steps.extend(smote(X, y, **smote_params, fit=False))
steps.append(('classifier', RandomForestClassifier()))
pipe = Pipeline(steps=steps)
#save k selected column names
save_selected_features(df = df,
k = k,
model=pipe,
prefix = 'fileops_' + category,
path='selected-features/ransomware',
file= 'fileops_' + category + '_nested_files.pkl',
label='sublabel')
# -
# ## Loaded DLL
batch_feature_selection(categories = ['onehot'],
k_range = [100, 1001],
step=50,
scoring='accuracy',
dataset_prefix='loaded_dll_',
dataset='loaded_dll',
label='sublabel')
pd.read_csv('selected-features/k-search-summary/ransomware/loaded_dll_accuracy_step50.csv', index_col=0)
# +
#load data
file_name = 'loaded_dll_onehot.pkl.gz'
df = get_data(file_path = os.path.join(folder_path, file_name), compression='gzip')
#build model
steps = []
smote_params = {
'category' : 'adaptive',
'over_strategy' : 0.5,
'under_strategy' : 0.8,
'k_neighbors' : 5,
}
X, y = get_X_y(df, label='sublabel')
steps.extend(smote(X, y, **smote_params, fit=False))
steps.append(('classifier', RandomForestClassifier()))
pipe = Pipeline(steps=steps)
#save k selected column names
save_selected_features(df = df,
k = 400,
model=pipe,
prefix = 'loaded_dll',
path='selected-features/ransomware',
file= 'loaded_dll_onehot.pkl',
label='sublabel')
# -
# ## PE Entropy
batch_feature_selection(categories = ['analysis'],
k_range = [100, 1001],
step=50,
scoring='accuracy',
dataset_prefix='pe_entropy_',
dataset='pe_entropy',
label='sublabel')
pd.read_csv('selected-features/k-search-summary/ransomware/pe_entropy_accuracy_step50.csv', index_col=0)
# +
#load data
file_name = 'pe_entropy_analysis.pkl.gz'
df = get_data(file_path = os.path.join(folder_path, file_name), compression='gzip')
#build model
steps = []
smote_params = {
'category' : 'adaptive',
'over_strategy' : 0.5,
'under_strategy' : 0.8,
'k_neighbors' : 5,
}
X, y = get_X_y(df, label='sublabel')
steps.extend(smote(X, y, **smote_params, fit=False))
steps.append(('classifier', RandomForestClassifier()))
pipe = Pipeline(steps=steps)
#save k selected column names
save_selected_features(df = df,
k = 100,
model=pipe,
prefix = 'pe_entropy',
path='selected-features/ransomware',
file= 'pe_entropy_analysis.pkl',
label='sublabel')
# -
# ## PE Imports
# **Libraries :**
batch_feature_selection(categories = ['libraries'],
k_range = [50, 551],
step=25,
scoring='accuracy',
dataset_prefix='pe_imports_',
dataset='pe_imports',
label='sublabel')
pd.read_csv('selected-features/k-search-summary/ransomware/pe_imports_accuracy_step25.csv', index_col=0)
# +
#load data
file_name = 'pe_imports_libraries.pkl.gz'
df = get_data(file_path = os.path.join(folder_path, file_name), compression='gzip')
#build model
steps = []
smote_params = {
'category' : 'adaptive',
'over_strategy' : 0.5,
'under_strategy' : 0.8,
'k_neighbors' : 5,
}
X, y = get_X_y(df, label='sublabel')
steps.extend(smote(X, y, **smote_params, fit=False))
steps.append(('classifier', RandomForestClassifier()))
pipe = Pipeline(steps=steps)
#save k selected column names
save_selected_features(df = df,
k = 75,
model=pipe,
prefix = 'pe_imports_libraries',
path='selected-features/ransomware',
file= 'pe_imports_libraries.pkl',
label='sublabel')
# -
# **Imports per library (most frequent libraries):**
# +
categories = ['kernel32', 'user32', 'advapi32', 'shell32', 'ole32']
batch_feature_selection(categories = categories,
k_range = [100, 1001],
step=50,
scoring='accuracy',
dataset_prefix='pe_imports_',
dataset='pe_imports_top_libraries',
label='sublabel')
# -
pd.read_csv('selected-features/k-search-summary/ransomware/pe_imports_top_libraries_accuracy_step50.csv', index_col=0)
# Although some k values gives the highest accuracy, if we take into account **minimizing model complexity and the variance on the accuracy results**, we'll opt for the following k values: <br/><br/>
# **kernel32 :** k=150 <br/>
# **user32 :** k=200 <br/>
# **advapi32 :** k=400 <br/>
# **shell32 :** k=150 <br/>
# **ole32 :** k=300 <br/>
#
# **Total PE imports selected columns =** 1200 features
# +
selected_k = [150, 200, 400, 150, 300]
for category, k in zip(categories, selected_k):
#load data
file_name = 'pe_imports_' + category + '.pkl.gz'
df = get_data(file_path = os.path.join(folder_path, file_name), compression='gzip')
#build model
steps = []
smote_params = {
'category' : 'adaptive',
'over_strategy' : 0.5,
'under_strategy' : 0.8,
'k_neighbors' : 5,
}
X, y = get_X_y(df, label='sublabel')
steps.extend(smote(X, y, **smote_params, fit=False))
steps.append(('classifier', RandomForestClassifier()))
pipe = Pipeline(steps=steps)
#save k selected column names
save_selected_features(df = df,
k = k,
model=pipe,
prefix = 'pe_imports_' + category,
path='selected-features/ransomware',
file= 'pe_imports_' + category + '.pkl',
label='sublabel')
| model-building/feature-selection-ransomware.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Introduction to Python
#
# What are the benefits of using Python?
#
# * General-purpose.
# * Interpreted.
# * Focuses on readability.
# * Comprehesive standard library.
# * Extended with a large number of third-party packages.
# * Widely used in scientific programming.
#
# This presentation will give a brief into to some key features of Python to help those not familar
# with the language with the remainder of the class. This is in no way a comprehensive introduction
# to either topic, for the sake of time. Excellent tutorials on Python can be found online. If nothing else, Stack Overflow will probably have the answer to any questions you may have regarding Python.
#
# In the next notebook, we will discuss some numerical and scientific Python packages.
# As seen in the first notebook, we can print text by using the print function.
print("I'm Mr. Meeseeks look at me!!!!")
# ## Variables and Types
# In Python you can define a variable and assign it a value. For example:
foo = 1
# foo is the integer 1, so we can add, subtract etc to foo.
foo + 1
# We need to reassign or reuse the variable to keep the add integer
foo2 = foo + 1
print(foo2)
foo = foo + 1
print(foo)
# There are a quite a few different data types in Python, but to name a couple for the sake of time:
# * Strings
# * Floats
# * Integers
# * Lists
# * and more..
# To check a data type we can simply place type in front of that variable:
bar = 3.5
type(bar)
# Python even allows us to combine string variables:
obiwan = "That's no moon... "
kenobi = "It's a space station!"
obiwan_kenobi = obiwan + kenobi
print(obiwan_kenobi)
# ## Containers
# In Python there are also containers. Tuples, lists and dictionaries.
# A list is exactly how it sounds, and list allow us to search via indexing.
# To create at list:
my_list = ['hello', 42, 100000.1, 'I Love Python!!']
# and yes lists can contain different types of variables, floats, strings you name it!
# To index:
print(my_list[3])
# A tuple is a sequence of immutable Python objects. Being sequences, tuples are like lists,
# but the difference between the two is that the tuples cannot be changed unlike lists. These are useful for providing coordinates.
tup = (1, 2, 3)
print(tup)
tup[1] = 6
# With dictionaries, we essential can store information and have keys that we can access. You will see this a lot in
# Py-ART. The fields are stored this way.
arctic = {'animals': 'polar bear, walrus and more',
'size': 13985000,
'size_units': 'km2',
'climate': 'Really cold... but the northern lights are awesome!'}
arctic['animals']
# ## Flow Control Statements
# There are flow control statements in Python. With these we can check different conditions and which of them are met.
a = 10
if a > 10:
print("a is larger than 10")
elif a < 10:
print("a is less than 10")
else:
print("a is equal to 10")
# We can also create loops to do an action to multiple items. Let's take a list for example.
releases = ['Dali', '<NAME>', 'Michelangelo']
for release in releases:
print(release + ' is going to be a Py-ART release!')
# ## Functions
# Functions are a block of reusable code that can perform an action and can even return variables.
# You can also input variables into a function.
# +
def func():
print("Hello world")
func()
# +
def name_func(name):
print("Hello", name)
name_func("<NAME>")
# +
def addition_func(x):
return x + 42
new_num = addition_func(2)
print(new_num)
# -
# ## Classes
# Classes allow for the bundling of data and functionality. Creating a new class
# creates a new type of object.
# +
class Car(object):
engine = 'V4' # class attribute
def start(self): # class method
print("Starting the car with a", self.engine, "engine")
mycar = Car()
# -
type(mycar)
mycar.engine
mycar.start()
mycar.engine = 'V6'
mycar.engine
mycar.start()
# ## Importing
# In Python we can import packages to be used. We will go into more detail on two popular Python packages
# in the next notebook. To import a package:
import turtle
# We can also import a package and assign it a different name.
import numpy as np
# We can also import from within a package.
from numpy import array
# ## Exercise # 1
# Create a function that uses 'if else statement flow control' to determine if the input value is an even number or an odd number.
#
# Hint: In Python, you can get the remainder of division by using the '%' command. For example,
# ```
# 7 % 3
# ```
# The above command will return the remainder of 7 divided by 3, which is one.
# %load section2_answer.py
# ## For and while loops
# When you do data analysis, you will run into many instances where you'll want to repeat a certain operation numerous times. For example, you'll want to go through items in a list and analyze the properties of each item so that you can provide summary statistics. Using the same line over and over again will take too much room and be a hassle. This is where for and while loops come in.
# For loops are useful for iterating over sequential lists. For example, if you want to do the same job, but just increment a variable by 1 before doing the same code, you can do this with a for loop.
print("I'm the Count! I love to count!")
for i in range(10):
print(i)
print("Oh, I'm so excited!")
# You can also iterate over items in a list as well as do incremental counting.
kids = ["Joey", "Maria", "Lamar", "Bobby", "Eduardo", "Liz"]
print("You kids need to be quiet! You're too loud! Get off my lawn!")
for brat in kids:
print(brat + ", get off my lawn!")
# Finally, sometimes you want to wait until a certain condition is met. This is where while loops come in. For example, if you want to flip a coin until you get 100 heads, you don't necessarily know how many flips you'll need in advance. But, with a while loop, we can keep going until we get 100 heads.
# +
import random
def coin_flip():
return random.random() < 0.5
num_flips = 0
num_heads = 0
while num_heads < 100:
num_heads += coin_flip()
num_flips += 1
print("We had to flip the coin " + str(num_flips) + " times.")
| 0.2_Introduction_to_python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="3rqsGAHiVx-n"
# # Gallery web app in `ReactJS` & `Firebase`
#
# **References**
# - Traversy Media, Net Ninja (2020) Build a Photo Gallery With React & Firebase https://www.youtube.com/watch?v=vUe91uOx7R0
# - Source Code by Net Ninja from https://github.com/iamshaunjp/firegram/tree/starter-files
# + [markdown] id="46x7fu2Xw23A"
# ## Firebase setup
# + [markdown] id="JQsliqROVou8"
# ### 1 Create back-end on `Firebase`
# 1. Create a web app on [firebase.google.com](https://firebase.google.com/).
# + [markdown] id="t4ZmoswwVioL"
# ### 2 Import `config.js` in `React App`
#
# 2. Create a folder (`firebase`) in `src` folder, then the conifg file (`config.js`) in it as indicated below. In `config.js`, paste the following snippet from the newly created `firebase project` in your `firebase` account.
#
# 📂 `my-project`
# ├── 📂node_modules
# ├── 📂public
# ├── 📂src
# │ㅤㅤ├── 📂**`firebase`**
# │ㅤㅤ│ㅤㅤ└── 📃**`config.js`**
# ├── 📃package-lock.json
# ├── 📃package.json
# └── 📃README.md
# + id="E1bcgtpjTSV6"
// Import the functions you need from the SDKs you need
import { initializeApp } from "my-project/app";
// TODO: Add SDKs for Firebase products that you want to use
// https://firebase.google.com/docs/web/setup#available-libraries
// Your web app's Firebase configuration
const firebaseConfig = {
apiKey: "",
authDomain: "",
projectId: "my-project",
storageBucket: "my-project-3ra21.appspot.com",
messagingSenderId: "random nuber",
appId: "1:random number:web:random string"
};
// Initialize Firebase
const app = initializeApp(firebaseConfig);
# + [markdown] id="L0omALloV2Y9"
# ### 3 Install `firebase` in `React App`
# + id="WMb7G6ivWBaw"
npm install firebase
# + [markdown] id="DzRp7IEwVbPT"
# ### 4 Import `firebase` and dependencies into `React App`
# + id="skbGM5X9VzYo"
# import * as firebase from 'firebase/app';
import { initializeApp } from "firebase/app";
import 'firebase/storage'; # for static assets like images
import 'firebase/firesotre';
# Initialize Firebase
const app = initializeApp(firebaseConfig); # this like connects the React app with the firebase back-end.
const firestore = getFirestore(app); # initialising 2 service components
const storage = getStorage(app);
export {firestore, storage}; # Make the services available to other documents within React App.
# + [markdown] id="3moaxymjX8fX"
# ### 5 Initialise `firestore` and `storage` on `Firebase` back-end
#
# - select db zone
# - select production mode / `test mode`
# - `test mode` is easier.
# + [markdown] id="_OoLVeLWZasc"
# ### 6 Start a storage service.
# - click start
# - and create an empty storage.
# - click on the `rule` tap.
# - edit the rule as below snippet.
# + id="pf_yrihZYhN-"
rules_version = '2';
service firebase.storage {
match /b/{bucket}/o {
match /{allPaths=**} {
allow read, write; ##: if request.auth != null;
}
}
} # then click publish
| 1 Advanced/Mini Project 4 Gallery - 1 Setup.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Wrangling - Deduplicate Data
#
# Load, deduplicate and pickle the Chicago tree trims data. The wrangling of the tree trims dataset is split into two parts to keep the notebook size manageable. This is part two. Part one is located in the `1-data-wrangling-optimize-verify.ipynb` notebook.
#
# **Project Notebooks Execution Order**
# * [1-data-wrangling-optimize-verify.ipynb](1-data-wrangling-optimize-verify.ipynb)
# * 2-data-wrangling-deduplicate.ipynb
# * [3-exploratory-data-analysis.ipynb](3-exploratory-data-analysis.ipynb)
# * [4-forecasting-opened-requests.ipynb](4-forecasting-opened-requests.ipynb)
# * [5-forecasting-closed-requests.ipynb](5-forecasting-closed-requests.ipynb)
# * [6-summary.ipynb](6-summary.ipynb)
#
# For a full explanation of this project, see the `README.md` file at the project root or the `6-summary.ipynb` notebook.
#
# # Import Packages and Modules
# +
import os
from cycler import cycler
from IPython.core.interactiveshell import InteractiveShell
import matplotlib as mpl
import matplotlib.dates as mdates
import matplotlib.pyplot as plt
from pandas.plotting import register_matplotlib_converters
# -
# # Settings
#
# Configure settings for notebook, plots, files and environmental variables. Set `ast_node_interactivity` to display the output of all expressions in a cell so that more than one data frame can be printed at a time in a single cell. Semicolons at the end of `plt.show();` are used in this notebook to suppress the printing of matplotlib objects in cell outputs. Load functions called in notebook.
# +
# Notebook settings
InteractiveShell.ast_node_interactivity = "all"
# %matplotlib inline
register_matplotlib_converters()
# Plot settings
alpha_mpl = 0.75
color_cycle = ['dodgerblue', '#d5019a', 'midnightblue', 'gray']
plt.style.use('seaborn-darkgrid')
mpl.rcParams['figure.titlesize'] = 18
mpl.rcParams['axes.prop_cycle'] = cycler(color=color_cycle)
mpl.rcParams['axes.titlesize'] = 18
mpl.rcParams['axes.labelsize'] = 14
mpl.rcParams['xtick.labelsize'] = 11
mpl.rcParams['ytick.labelsize'] = 11
mpl.rcParams['legend.fontsize'] = 11
# File settings
data_raw = os.path.abspath('./data/raw/')
data_interim = os.path.abspath('./data/interim/')
data_processed = os.path.abspath('./data/processed/')
# Import preprocessing functions
# %run preprocess_data.py
# -
# # Load Data
#
# ## Create Data Frame
df_tt = read_data('Chicago Tree Trims', data_interim, 'chicago-tree-trims.pkl')
# # Duplicates
#
# Review the dataset for duplicated data that can be dropped from the dataset. How many entire rows are duplicated?
print(
'Number of entire rows that are duplicated: {}'
.format(df_tt.duplicated().sum())
)
# Duplicates in the dataset are identified in the `status` column with either the "Completed - Dup" value or the "Open - Dup" value. (See https://data.cityofchicago.org/Service-Requests/311-Service-Requests-Tree-Trims/uxic-zsuj.) How many records in the dataset are marked as "Completed - Dup" or the "Open - Dup"?
(df_tt['status'].value_counts().to_frame('count'))
# Next, verify whether or not all duplicates in the dataset are correctly classified as duplicates in the `status` column. Start by checking for possible duplicate Service Request Numbers.
#
# # Duplicated Service Request Numbers (SRNs)
#
# Service Request Numbers (SRNs) identify individual tree trim requests. Its possible that there are duplicate SRNs in the dataset and those duplicates may or may not be correctly classified in the `status` column. Check a few scenarios to confirm whether or not SRN duplicates exist and if they are correctly classified.
#
# ## Identified Duplicates
#
# Are there duplicated SRNs in the dataset?
srn_dupes = df_tt[df_tt['service_request_number'].duplicated()].shape[0]
print('Number of duplicated SRNs: {}'.format(srn_dupes))
# What is the `status` value of the duplicated SRNs? Are all duplicated values correctly classified in the `status` column?
(
df_tt[df_tt['service_request_number'].duplicated()]['status']
.value_counts()
.to_frame('count')
)
# There are duplicated SRNs that are not classified as duplicates. Refer to these duplicates as unidentified duplicates.
#
# ## Unidentified Duplicates
#
# Are the unidentified duplicates clustered or spread across the dataset?
# +
dupe_status = ['Completed - Dup', 'Open - Dup']
mask_dupes_unident = (
~(df_tt['status'].isin(dupe_status)) &
(df_tt['service_request_number'].duplicated(keep='first'))
)
srn_dupes_dates = df_tt[mask_dupes_unident]['creation_date_dt']
fig, ax = plt.subplots(figsize=(6, 4))
ax.hist(mdates.date2num(srn_dupes_dates), bins=8, alpha=alpha_mpl)
ax.xaxis.set_major_locator(mdates.YearLocator())
ax.xaxis.set_major_formatter(mdates.DateFormatter('%Y'))
ax.tick_params(axis='x', rotation=90)
ax.set_xlabel('Year')
ax.set_ylabel('Frequency')
ax.set_title('Unidentified SRN Duplicates')
plt.show();
# -
# Unidentified duplicates are primarily clustered in the years 2013-2015. The volume of unidentified duplicates in those years may impact the models, so investigate the unidentified duplicates further to determine if they can be dropped from the dataset.
#
# As for identified duplicates (i.e. `status` values "Completed - Dup" or "Open - Dup"), drop them from the dataset.
print('df_tt shape before: {}'.format(df_tt.shape))
df_tt = df_tt[~df_tt['status'].isin(['Completed - Dup', 'Open - Dup'])]
print('df_tt shape after: {}'.format(df_tt.shape))
# There isn't any additional information at the Chicago Data Portal (see https://data.cityofchicago.org/Service-Requests/311-Service-Requests-Tree-Trims/uxic-zsuj) about why the SRN duplicates are not identified as duplicates in the dataset. Is there any information in the data that would indicate whether or not the unidentified duplicates can be dropped from the dataset?
#
# ### Considerations for Unidentified SRN Duplicates
# * When the dataset is grouped by year and month, as it will be for the models, what percentage of the records are duplicates? Do duplicates play a large or small role in the dataset at the month level?
# * Are there requests in the dataset that were closed en mass (perhaps suggesting a mass data cleanup effort) and, if so, do the SRN duplicates play a large role in those data spikes?
# * How many duplicates are there per SRN? If the size of duplicated groups is three or more, it could complicate the process of deciding which duplicate to keep.
# * Which columns conflict between the duplicated SRNs? (There are likely column that conflict between duplicates as entire rows are not duplicated in the dataset.)
# * Are NaNs in columns causing conflicts between duplicated SRNs?
# * Are there conflicts between the `creation_date_dt` and `completion_date_dt` columns between duplicated SRNs?
#
# Setup filters to identify a subset of rows that include all duplicated SRN records, including the first instance of a duplicate, and a subset of rows that ignores the first instance and keeps all subsequent instances of the duplicate. These filters will be used below for cleanup work.
srn_dupes_all = df_tt.duplicated('service_request_number', keep=False)
srn_dupes_all_idx = df_tt[srn_dupes_all].index
srn_dupes_first = df_tt.duplicated('service_request_number', keep='first')
srn_dupes_first_idx = df_tt[srn_dupes_first].index
# ### Percentage of Duplicates
#
# What percentage of the dataset records are duplicates? Break down the percentage of duplicates by month to understand the impact of duplicates across the dataset. Return the months with the highest rate of duplicates.
print(
'Percentage of SRN duplicates in the dataset: {:.2%}'
.format(srn_dupes_first.sum() / df_tt.shape[0])
)
# The overall percentage of duplicates in the dataset is less than 1%.
# +
def group_by_date(col, df, idx=None):
'''Group data frame by year and month.
Parameters
----------
col : str
Datetime column used to group data frame.
df : pd.DataFrame
Data frame to group.
idx : pd.index, optional
Index used to filter data frame.
Returns
-------
df : pd.DataFrame
Data frame with DatetimeIndex grouped by year and month and
count of records per year and month.
'''
if idx is not None:
df = df[df[col].notnull()].loc[idx].set_index(col)
col_name = 'counts_dupes'
else:
df = df[df[col].notnull()].set_index(col)
col_name = 'counts_all'
groups = [df.index.year, df.index.month]
df = df.groupby(groups).size().to_frame(col_name)
return df
def top_dupes_by_percent(col, df=df_tt, idx=srn_dupes_all_idx):
'''Return top percentage of duplicates per year and month.
Parameters
----------
col : str
Datetime column used to group data frame.
df : pd.DataFrame
Data frame to group.
idx : pd.index, optional
Index used to filter data frame.
Returns
-------
df : pd.DataFrame
Data frame of top percentage of duplicates per year and month.
'''
df_size = group_by_date(col, df)
df_dupes = group_by_date(col, df, idx.values)
df_size = df_size.merge(df_dupes, how='left', left_index=True,
right_index=True)
df_size = df_size.fillna(0)
df_size['percentage'] = df_size['counts_dupes'] / df_size['counts_all']
df_size = df_size[df_size['percentage'] > .0099]
df_size = df_size.style.format({'percentage': '{:.2%}'})
return df_size
top_dupes_by_percent('creation_date_dt')
top_dupes_by_percent('completion_date_dt')
# -
# For a majority of the dataset, the impact of duplicates for any given month is marginal. However, for May 2013, the impact of duplicates for both `completion_date_dt` and `completion_date_dt` variables is high. Also the percentage of duplicate `completion_date_dt` values in November 2014 is nearly 10%.
#
# ### Request Closures En Mass
#
# Are there usually large spikes in request closures and, if so, do the duplicates play a role in those spikes?
#
# Plot the number of closed requests per day to visualize unusual spikes in the closing dates for requests. To smooth out the noise in the line graph below, use a rolling mean (i.e., for every data point, average the data points on either side). Also plot the number of closed requests for unidentified duplicate SRNs.
# +
df_comp_date_counts = (
df_tt[df_tt['completion_date_dt'].notnull()]
.groupby('completion_date_dt')
.size()
.to_frame('count')
.reset_index()
)
x_1 = df_comp_date_counts['completion_date_dt']
y_1 = df_comp_date_counts['count'].rolling(12).mean()
y_median = df_comp_date_counts['count'].median()
df_srn_dupe = (
df_tt[df_tt['completion_date_dt'].notnull()]
.loc[srn_dupes_first_idx]
.groupby('completion_date_dt')
.size()
.to_frame('count')
.reset_index()
)
x_2 = df_srn_dupe['completion_date_dt']
y_2 = df_srn_dupe['count']
fig, ax = plt.subplots(figsize=(11, 4))
ax.plot(x_1, y_1, alpha=alpha_mpl, linewidth=1, label='Rolling Mean')
ax.plot(
x_2,
y_2,
color='C2',
alpha=alpha_mpl,
linewidth=1,
label='Duplicate SRN',
)
ax.axhline(
y_median,
color='C1',
alpha=0.5,
linewidth=1,
linestyle='--',
label='Median',
)
ax.xaxis.set_major_locator(mdates.YearLocator())
ax.xaxis.set_major_formatter(mdates.DateFormatter('%Y'))
ax.set_xlabel('Years')
ax.set_ylabel('Closed Requests')
ax.set_title('Number of Closed Requests per Day')
ax.legend(loc='upper left')
plt.show();
# -
# Aside for May 2013 and November 2014, as indicated above, duplicated SRNs play only a small role the spikes of closed requests.
#
# ### Number of Duplicates per SRN
#
# Group the SRN duplicates and count the number of duplicates per group. What is the range of group sizes?
(
df_tt[srn_dupes_all]
.groupby('service_request_number')
.size()
.value_counts()
.to_frame()
.reset_index()
.rename({'index': 'group_size', 0: 'count'}, axis='columns')
)
# There are a handful of duplicates SRNs where there are 3 duplicates in a group.
#
# ### NaNs
#
# NaNs in columns could be the cause of differences between duplicate SRNs. Out of the duplicate SRNs, which columns contain NaNs? If possible, fill NaN values with populated values from the corresponding SRN duplicate.
df_tt[srn_dupes_all].isnull().sum().to_frame('nan_count')
# How many of the duplicate SRNs with NaNs are in groups of 3? Filling NaN values in groups of three is not as straight forward as doing the same for groups of two.
# +
s_nan_groups = (
df_tt[srn_dupes_all & (df_tt.isnull().any(axis=1))]
.groupby('service_request_number')
.size()
.value_counts()
)
print(
'Number of groups with three records and NaN values: {}'
.format(s_nan_groups.loc[3])
)
# -
# The NaN values in groups of three are isolated to one group. Which columns in that group are NaN?
s_group_three = (
df_tt[srn_dupes_all]
.groupby('service_request_number')
.size()
.loc[lambda x: x == 3]
.index
)
df_tt[
(df_tt['service_request_number'].isin(s_group_three)) &
(df_tt.isnull().any(axis=1))
].isnull().sum().to_frame('nan_count')
# All of the NaN values are isolated to the `location_of_trees` column, so groups of three don't need any modification. Leave the groups of three out of the filling operation below.
#
# Fill the NaN values where populated values are available. Some groups may contain NaN values in all rows for a particular column.
df_tt.loc[srn_dupes_all_idx] = (
df_tt[srn_dupes_all]
.groupby('service_request_number')
.apply(lambda g: g.bfill().ffill())
)
# Verify the fill results.
df_tt[srn_dupes_all].isnull().sum().to_frame('nan_count')
# ### Diff Rows
#
# Now that the NaN values are filled in, which columns are responsible for the conflict between duplicates and how severe are the conflicts? A combination of the severity level of the conflicts and the importance of the column in the forecasting models will determine whether or not to further wrangle the duplicates.
#
# Create a boolean data frame that indicates which columns conflict.
# +
def compare_rows(g):
'''Diff columns between rows.
Within each group, compare the values in the next row against the
values in the previous row. If the values differ, return True, if
not, return False.
Parameters
----------
g : group in pandas GroupBy object
A single group in Pandas GroupBy object.
Returns
-------
df_diff : pd.DataFrame
Data frame of booleans.
'''
df_diff = g.ne(g.shift())
return df_diff
df_diff = (
df_tt[srn_dupes_all]
.sort_values(['service_request_number', 'completion_date_dt'])
.groupby('service_request_number')
.apply(lambda g: compare_rows(g))
)
# -
# Review the results of the last row of each group in df_diff. The last row indicates which columns hold the differences between the rows within groups.
# +
sort = ['service_request_number', 'completion_date_dt']
srn_dupes_last_idx = (
df_tt[srn_dupes_all]
.reset_index()
.sort_values(sort)
.groupby('service_request_number')
.last()['index']
)
df_diff.loc[srn_dupes_last_idx].sum().to_frame('Diff')
# -
# The `completion_date_dt` and `creation_date_dt` columns are important for the forecasting models. The majority of the differences between rows of duplicate SRNs is in the `completion_date_dt` field. There are a minor number of differences in the geolocation related fields. Those geolocation fields are not critical the models and their numbers are low, so leave them as is.
# **Created Date and Completed Date Conflicts**
# What is the range of differences in days between the completion dates for duplicated SRNs? View the distribution in log scale to visualize the long tail more clearly.
# +
diff_completion = (
df_tt[srn_dupes_all]
.sort_values(sort)
.groupby('service_request_number')['completion_date_dt']
.diff().dt.days
.loc[lambda x: x.notnull()]
.astype(int)
.value_counts()
)
fig, ax = plt.subplots(figsize=(8, 4))
ax.hist(diff_completion, bins=25, alpha=alpha_mpl, log=True)
ax.set_xticks(range(0, diff_completion.max(), 50))
ax.set_xlabel('Number of Days')
ax.set_ylabel('Frequency')
ax.set_title('Differences in Days Between Completion Date')
plt.show();
# -
# The majority of the differences in days between conflicting `completion_date_dt` dates fall within a 50 day window with a few outliers.
#
# ### Duplicates Interim Solution
#
# After examining the SRN duplicates from several angles and without a firm conclusion as to why these duplicates are not marked as such in the dataset, the best course of action is to keep the duplicates in the dataset for now. Mark them as duplicates in a new column, fit each model with and without the duplicates and then compare the results between models.
# +
df_tt['is_duplicate'] = False
drop_dupe_idx = (df_tt[srn_dupes_first]
.sort_values(['creation_date_dt', 'completion_date_dt'])
.index)
df_tt.loc[drop_dupe_idx, 'is_duplicate'] = True
print('Number of SRN duplicates: {}'.format(df_tt['is_duplicate'].sum()))
# -
# # Export Data
#
# Export data to `/data/processed/` directory. Pickle the data frame to preserve its attributes for reading into subsequent notebooks.
#
# **Chicago Tree Trims**
export_pickle('chicago-tree-trims', df_tt, data_processed)
| notebooks/chicago-tree-trims-time-series-forecasting/2-data-wrangling-deduplicate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + [markdown] origin_pos=0
# # 暂退法(Dropout)
# :label:`sec_dropout`
#
# 在 :numref:`sec_weight_decay` 中,
# 我们介绍了通过惩罚权重的$L_2$范数来正则化统计模型的经典方法。
# 在概率角度看,我们可以通过以下论证来证明这一技术的合理性:
# 我们已经假设了一个先验,即权重的值取自均值为0的高斯分布。
# 更直观的是,我们希望模型深度挖掘特征,即将其权重分散到许多特征中,
# 而不是过于依赖少数潜在的虚假关联。
#
# ## 重新审视过拟合
#
# 当面对更多的特征而样本不足时,线性模型往往会过拟合。
# 相反,当给出更多样本而不是特征,通常线性模型不会过拟合。
# 不幸的是,线性模型泛化的可靠性是有代价的。
# 简单地说,线性模型没有考虑到特征之间的交互作用。
# 对于每个特征,线性模型必须指定正的或负的权重,而忽略其他特征。
#
# 泛化性和灵活性之间的这种基本权衡被描述为*偏差-方差权衡*(bias-variance tradeoff)。
# 线性模型有很高的偏差:它们只能表示一小类函数。
# 然而,这些模型的方差很低:它们在不同的随机数据样本上可以得出了相似的结果。
#
# 深度神经网络位于偏差-方差谱的另一端。
# 与线性模型不同,神经网络并不局限于单独查看每个特征,而是学习特征之间的交互。
# 例如,神经网络可能推断“尼日利亚”和“西联汇款”一起出现在电子邮件中表示垃圾邮件,
# 但单独出现则不表示垃圾邮件。
#
# 即使我们有比特征多得多的样本,深度神经网络也有可能过拟合。
# 2017年,一组研究人员通过在随机标记的图像上训练深度网络。
# 这展示了神经网络的极大灵活性,因为人类很难将输入和随机标记的输出联系起来,
# 但通过随机梯度下降优化的神经网络可以完美地标记训练集中的每一幅图像。
# 想一想这意味着什么?
# 假设标签是随机均匀分配的,并且有10个类别,那么分类器在测试数据上很难取得高于10%的精度,
# 那么这里的泛化差距就高达90%,如此严重的过拟合。
#
# 深度网络的泛化性质令人费解,而这种泛化性质的数学基础仍然是悬而未决的研究问题。
# 我们鼓励喜好研究理论的读者更深入地研究这个主题。
# 本节,我们将着重对实际工具的探究,这些工具倾向于改进深层网络的泛化性。
#
# ## 扰动的稳健性
#
# 在探究泛化性之前,我们先来定义一下什么是一个“好”的预测模型?
# 我们期待“好”的预测模型能在未知的数据上有很好的表现:
# 经典泛化理论认为,为了缩小训练和测试性能之间的差距,应该以简单的模型为目标。
# 简单性以较小维度的形式展现,
# 我们在 :numref:`sec_model_selection` 讨论线性模型的单项式函数时探讨了这一点。
# 此外,正如我们在 :numref:`sec_weight_decay` 中讨论权重衰减($L_2$正则化)时看到的那样,
# 参数的范数也代表了一种有用的简单性度量。
#
# 简单性的另一个角度是平滑性,即函数不应该对其输入的微小变化敏感。
# 例如,当我们对图像进行分类时,我们预计向像素添加一些随机噪声应该是基本无影响的。
# 1995年,克里斯托弗·毕晓普证明了
# 具有输入噪声的训练等价于Tikhonov正则化 :cite:`Bishop.1995`。
# 这项工作用数学证实了“要求函数光滑”和“要求函数对输入的随机噪声具有适应性”之间的联系。
#
# 然后在2014年,斯里瓦斯塔瓦等人 :cite:`Srivastava.Hinton.Krizhevsky.ea.2014`
# 就如何将毕晓普的想法应用于网络的内部层提出了一个想法:
# 在训练过程中,他们建议在计算后续层之前向网络的每一层注入噪声。
# 因为当训练一个有多层的深层网络时,注入噪声只会在输入-输出映射上增强平滑性。
#
# 这个想法被称为*暂退法*(dropout)。
# 暂退法在前向传播过程中,计算每一内部层的同时注入噪声,这已经成为训练神经网络的常用技术。
# 这种方法之所以被称为*暂退法*,因为我们从表面上看是在训练过程中丢弃(drop out)一些神经元。
# 在整个训练过程的每一次迭代中,标准暂退法包括在计算下一层之前将当前层中的一些节点置零。
#
# 需要说明的是,暂退法的原始论文提到了一个关于有性繁殖的类比:
# 神经网络过拟合与每一层都依赖于前一层激活值相关,称这种情况为“共适应性”。
# 作者认为,暂退法会破坏共适应性,就像有性生殖会破坏共适应的基因一样。
#
# 那么关键的挑战就是如何注入这种噪声。
# 一种想法是以一种*无偏向*(unbiased)的方式注入噪声。
# 这样在固定住其他层时,每一层的期望值等于没有噪音时的值。
#
# 在毕晓普的工作中,他将高斯噪声添加到线性模型的输入中。
# 在每次训练迭代中,他将从均值为零的分布$\epsilon \sim \mathcal{N}(0,\sigma^2)$
# 采样噪声添加到输入$\mathbf{x}$,
# 从而产生扰动点$\mathbf{x}' = \mathbf{x} + \epsilon$,
# 预期是$E[\mathbf{x}'] = \mathbf{x}$。
#
# 在标准暂退法正则化中,通过按保留(未丢弃)的节点的分数进行规范化来消除每一层的偏差。
# 换言之,每个中间活性值$h$以*暂退概率*$p$由随机变量$h'$替换,如下所示:
#
# $$
# \begin{aligned}
# h' =
# \begin{cases}
# 0 & \text{ 概率为 } p \\
# \frac{h}{1-p} & \text{ 其他情况}
# \end{cases}
# \end{aligned}
# $$
#
# 根据此模型的设计,其期望值保持不变,即$E[h'] = h$。
#
# ## 实践中的暂退法
#
# 回想一下 :numref:`fig_mlp`中带有1个隐藏层和5个隐藏单元的多层感知机。
# 当我们将暂退法应用到隐藏层,以$p$的概率将隐藏单元置为零时,
# 结果可以看作是一个只包含原始神经元子集的网络。
# 比如在 :numref:`fig_dropout2`中,删除了$h_2$和$h_5$,
# 因此输出的计算不再依赖于$h_2$或$h_5$,并且它们各自的梯度在执行反向传播时也会消失。
# 这样,输出层的计算不能过度依赖于$h_1, \ldots, h_5$的任何一个元素。
#
# 
# :label:`fig_dropout2`
#
# 通常,我们在测试时不用暂退法。
# 给定一个训练好的模型和一个新的样本,我们不会丢弃任何节点,因此不需要标准化。
# 然而也有一些例外:一些研究人员在测试时使用暂退法,
# 用于估计神经网络预测的“不确定性”:
# 如果通过许多不同的暂退法遮盖后得到的预测结果都是一致的,那么我们可以说网络发挥更稳定。
#
# ## 从零开始实现
#
# 要实现单层的暂退法函数,
# 我们从均匀分布$U[0, 1]$中抽取样本,样本数与这层神经网络的维度一致。
# 然后我们保留那些对应样本大于$p$的节点,把剩下的丢弃。
#
# 在下面的代码中,(**我们实现 `dropout_layer` 函数,
# 该函数以`dropout`的概率丢弃张量输入`X`中的元素**),
# 如上所述重新缩放剩余部分:将剩余部分除以`1.0-dropout`。
#
# + origin_pos=2 tab=["pytorch"]
import torch
from torch import nn
from d2l import torch as d2l
def dropout_layer(X, dropout):
assert 0 <= dropout <= 1
# 在本情况中,所有元素都被丢弃
if dropout == 1:
return torch.zeros_like(X)
# 在本情况中,所有元素都被保留
if dropout == 0:
return X
mask = (torch.rand(X.shape) > dropout).float()
return mask * X / (1.0 - dropout)
# + [markdown] origin_pos=4
# 我们可以通过下面几个例子来[**测试`dropout_layer`函数**]。
# 我们将输入`X`通过暂退法操作,暂退概率分别为0、0.5和1。
#
# + origin_pos=6 tab=["pytorch"]
X= torch.arange(16, dtype = torch.float32).reshape((2, 8))
print(X)
print(dropout_layer(X, 0.))
print(dropout_layer(X, 0.5))
print(dropout_layer(X, 1.))
# + [markdown] origin_pos=8
# ### 定义模型参数
#
# 同样,我们使用 :numref:`sec_fashion_mnist`中引入的Fashion-MNIST数据集。
# 我们[**定义具有两个隐藏层的多层感知机,每个隐藏层包含256个单元**]。
#
# + origin_pos=10 tab=["pytorch"]
num_inputs, num_outputs, num_hiddens1, num_hiddens2 = 784, 10, 256, 256
# + [markdown] origin_pos=12
# ### 定义模型
#
# 我们可以将暂退法应用于每个隐藏层的输出(在激活函数之后),
# 并且可以为每一层分别设置暂退概率:
# 常见的技巧是在靠近输入层的地方设置较低的暂退概率。
# 下面的模型将第一个和第二个隐藏层的暂退概率分别设置为0.2和0.5,
# 并且暂退法只在训练期间有效。
#
# + origin_pos=14 tab=["pytorch"]
dropout1, dropout2 = 0.2, 0.5
class Net(nn.Module):
def __init__(self, num_inputs, num_outputs, num_hiddens1, num_hiddens2,
is_training = True):
super(Net, self).__init__()
self.num_inputs = num_inputs
self.training = is_training
self.lin1 = nn.Linear(num_inputs, num_hiddens1)
self.lin2 = nn.Linear(num_hiddens1, num_hiddens2)
self.lin3 = nn.Linear(num_hiddens2, num_outputs)
self.relu = nn.ReLU()
def forward(self, X):
H1 = self.relu(self.lin1(X.reshape((-1, self.num_inputs))))
# 只有在训练模型时才使用dropout
if self.training == True:
# 在第一个全连接层之后添加一个dropout层
H1 = dropout_layer(H1, dropout1)
H2 = self.relu(self.lin2(H1))
if self.training == True:
# 在第二个全连接层之后添加一个dropout层
H2 = dropout_layer(H2, dropout2)
out = self.lin3(H2)
return out
net = Net(num_inputs, num_outputs, num_hiddens1, num_hiddens2)
# + [markdown] origin_pos=16
# ### [**训练和测试**]
#
# 这类似于前面描述的多层感知机训练和测试。
#
# + origin_pos=18 tab=["pytorch"]
num_epochs, lr, batch_size = 10, 0.5, 256
loss = nn.CrossEntropyLoss()
train_iter, test_iter = d2l.load_data_fashion_mnist(batch_size)
trainer = torch.optim.SGD(net.parameters(), lr=lr)
d2l.train_ch3(net, train_iter, test_iter, loss, num_epochs, trainer)
# + [markdown] origin_pos=20
# ## [**简洁实现**]
#
# 对于深度学习框架的高级API,我们只需在每个全连接层之后添加一个`Dropout`层,
# 将暂退概率作为唯一的参数传递给它的构造函数。
# 在训练时,`Dropout`层将根据指定的暂退概率随机丢弃上一层的输出(相当于下一层的输入)。
# 在测试时,`Dropout`层仅传递数据。
#
# + origin_pos=22 tab=["pytorch"]
net = nn.Sequential(nn.Flatten(),
nn.Linear(784, 256),
nn.ReLU(),
# 在第一个全连接层之后添加一个dropout层
nn.Dropout(dropout1),
nn.Linear(256, 256),
nn.ReLU(),
# 在第二个全连接层之后添加一个dropout层
nn.Dropout(dropout2),
nn.Linear(256, 10))
def init_weights(m):
if type(m) == nn.Linear:
nn.init.normal_(m.weight, std=0.01)
net.apply(init_weights);
# + [markdown] origin_pos=24
# 接下来,我们[**对模型进行训练和测试**]。
#
# + origin_pos=26 tab=["pytorch"]
trainer = torch.optim.SGD(net.parameters(), lr=lr)
d2l.train_ch3(net, train_iter, test_iter, loss, num_epochs, trainer)
# + [markdown] origin_pos=28
# ## 小结
#
# * 暂退法在前向传播过程中,计算每一内部层的同时丢弃一些神经元。
# * 暂退法可以避免过拟合,它通常与控制权重向量的维数和大小结合使用的。
# * 暂退法将活性值$h$替换为具有期望值$h$的随机变量。
# * 暂退法仅在训练期间使用。
#
# ## 练习
#
# 1. 如果更改第一层和第二层的暂退法概率,会发生什么情况?具体地说,如果交换这两个层,会发生什么情况?设计一个实验来回答这些问题,定量描述你的结果,并总结定性的结论。
# 1. 增加训练轮数,并将使用暂退法和不使用暂退法时获得的结果进行比较。
# 1. 当应用或不应用暂退法时,每个隐藏层中激活值的方差是多少?绘制一个曲线图,以显示这两个模型的每个隐藏层中激活值的方差是如何随时间变化的。
# 1. 为什么在测试时通常不使用暂退法?
# 1. 以本节中的模型为例,比较使用暂退法和权重衰减的效果。如果同时使用暂退法和权重衰减,会发生什么情况?结果是累加的吗?收益是否减少(或者说更糟)?它们互相抵消了吗?
# 1. 如果我们将暂退法应用到权重矩阵的各个权重,而不是激活值,会发生什么?
# 1. 发明另一种用于在每一层注入随机噪声的技术,该技术不同于标准的暂退法技术。尝试开发一种在Fashion-MNIST数据集(对于固定架构)上性能优于暂退法的方法。
#
# + [markdown] origin_pos=30 tab=["pytorch"]
# [Discussions](https://discuss.d2l.ai/t/1813)
#
| d2l/pytorch/chapter_multilayer-perceptrons/dropout.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python Data Structures: Linked Lists
#
# _This is a list of contents I have compiling to use in classes._
# _Fell free to share with others and improve._
#
# Everything below should begginers-friendly, ok?
#
# ## About Data Structures
#
# A data strucuture is a particular way of organization data in a program so that can be used effectively.
#
# One aproach is store a list of a itens having the same data-type using the array data structure.
#
# Linked Lists and Arrays are Linear forms of Data Strucuture. The other forms are Tree, Hash and Graphs.
# ## _What are Linked Lists_
#
# The simple and short way to say it: Linked lists are an ordered collection of objects.
# Like a list of days of the week or months in a year or even Pokemons in a team.
#
# Lists where something is followed by other thing and maybe preceded for another thing... Any list ordered list where one thing leads us to another thing.
#
# And that is it.
#
# But lets take a step back and try to understand
# Before going more in depth on what linked lists are and how you can use them, you should first learn how they are structured. Each element of a linked list is called a node, and every node has two different fields:
#
# __Practical Applications__
#
# Linked lists serve a variety of purposes in the real world. They can be used to implement (spoiler alert!) queues or stacks as well as graphs. They’re also useful for much more complex tasks, such as lifecycle management for an operating system application.
#
# Try to think about a sequence of operations that need to be executed in line.
#
# Someone needs to do something delicate like a dinner.
#
# 1. Buy the ingredientes, Next step.<br>
# 2. Check if the ove is ok, Next step.<br>
# 3. Put the main plate in the ove, Next step.<br>
# 4. Boil the water. Next step.<br>
# 5. Prepare the rice. Next step. <br>
# 6. Open the wine<br>
# 7. Serve the dinner.<br>
#
# This is just a example, but I think I made my point.
#
# Ok.
#
# Now lets put our hands in code.
#
# The first thing here, is that we will create a Class to manage our linked lists.
#
# Node class
class Node:
# Function to initialise the node object
def __init__(self, data):
self.data = data # Assign data
self.next = None # Initialize next as null
#
# +
# Linked List class contains a Node object
class LinkedList:
# Function to initialize head
def __init__(self):
self.head = None
def push(self, new_data):
# 1 & 2: Allocate the Node & Put in the data
new_node = Node(new_data)
# 3. Make next of new Node as head
new_node.next = self.head
# 4. Move the head to point to new Node
self.head = new_node
def printNthFromLast(self, n):
temp = self.head # used temp variable
length = 0
while temp is not None:
temp = temp.next
length += 1
# print count
if n > length: # if entered location is greater than length of linked list
print('Location is greater than the' + ' length of LinkedList')
return
temp = self.head
for i in range(0, length - n):
temp = temp.next
#print(temp.data)
# Returns data at given index in linked list
def getNth(self, index):
current = self.head # Initialise temp
count = 0 # Index of current node
# Loop while end of linked list is not reached
while (current):
if (count == index):
return current.data
count += 1
current = current.next
# if we get to this line, the caller was asking for a non-existent element so we assert fail
assert(False)
return 0;
# -
#
# +
llist = LinkedList()
# Use push() to construct below list
# llist.push('Dimanche')
# llist.push('Lundi')
# llist.push('Mardi')
# llist.push('Mercredi')
# llist.push('Jeudi')
# llist.push('Vendredi')
# llist.push('Samedi')
llist.push(1)
llist.push(2)
llist.push(3)
llist.push(4)
llist.push(5)
llist.push(6)
llist.push(7)
#Choose a day
def pick_a_day():
day = int(input("choose a number between 0 and 7: "))
print('List of days: ', llist.head)
print ("The day you choose is at position :", llist.getNth(day))
print ("printNthFromLast :", llist.printNthFromLast(day))
pick_a_day()
# -
#
#
#
# Pretty cool, huh?
#
# And that is it for now.
#
# I try compile the best solutions. Some stuff i did think was not that great, and there are better solution for _for your case_ out there.
#
# I will certainly improve this so keep checking back.
#
# Try to keep your code simple, try to drink more water and keep studing.
#
# __Thanks for reading.__
| Data_Structures_LinkedLists.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 预测波士顿房价
#
# ## 在 SageMaker 中使用 XGBoost(部署)
#
# _机器学习工程师纳米学位课程 | 部署_
#
# ---
#
# 为了介绍 SageMaker 的高阶 Python API,我们将查看一个相对简单的问题。我们将使用[波士顿房价数据集](https://www.cs.toronto.edu/~delve/data/boston/bostonDetail.html)预测波士顿地区的房价中位数。
#
# 高阶 API 的文档位于 [ReadTheDocs 页面](http://sagemaker.readthedocs.io/en/latest/)
#
# ## 一般步骤
#
# 通常,在 notebook 实例中使用 SageMaker 时,你需要完成以下步骤。当然,并非每个项目都要完成每一步。此外,有很多步骤有很大的变化余地,你将在这些课程中发现这一点。
#
# 1. 下载或检索数据。
# 2. 处理/准备数据。
# 3. 将处理的数据上传到 S3。
# 4. 训练所选的模型。
# 5. 测试训练的模型(通常使用批转换作业)。
# 6. 部署训练的模型。
# 7. 使用部署的模型。
#
# 在此 notebook 中,我们将跳过第 5 步 - 测试模型。我们依然会测试模型,但是首先会部署模型,然后将测试数据发送给部署的模型。
# ## 第 0 步:设置 notebook
#
# 先进行必要的设置以运行 notebook。首先,加载所需的所有 Python 模块。
# +
# %matplotlib inline
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.datasets import load_boston
import sklearn.model_selection
# -
# 除了上面的模块之外,我们还需要导入将使用的各种 SageMaker 模块。
# +
import sagemaker
from sagemaker import get_execution_role
from sagemaker.amazon.amazon_estimator import get_image_uri
from sagemaker.predictor import csv_serializer
# This is an object that represents the SageMaker session that we are currently operating in. This
# object contains some useful information that we will need to access later such as our region.
session = sagemaker.Session()
# This is an object that represents the IAM role that we are currently assigned. When we construct
# and launch the training job later we will need to tell it what IAM role it should have. Since our
# use case is relatively simple we will simply assign the training job the role we currently have.
role = get_execution_role()
# -
# ## 第 1 步:下载数据
#
# 幸运的是,我们可以使用 sklearn 检索数据集,所以这一步相对比较简单。
boston = load_boston()
# ## 第 2 步:准备和拆分数据
#
# 因为使用的是整洁的表格数据,所以不需要进行任何处理。但是,我们需要将数据集中的各行拆分成训练集、测试集和验证集。
# +
# First we package up the input data and the target variable (the median value) as pandas dataframes. This
# will make saving the data to a file a little easier later on.
X_bos_pd = pd.DataFrame(boston.data, columns=boston.feature_names)
Y_bos_pd = pd.DataFrame(boston.target)
# We split the dataset into 2/3 training and 1/3 testing sets.
X_train, X_test, Y_train, Y_test = sklearn.model_selection.train_test_split(X_bos_pd, Y_bos_pd, test_size=0.33)
# Then we split the training set further into 2/3 training and 1/3 validation sets.
X_train, X_val, Y_train, Y_val = sklearn.model_selection.train_test_split(X_train, Y_train, test_size=0.33)
# -
# ## 第 3 步:将训练和验证文件上传到 S3
#
# 使用 SageMaker 创建训练作业后,进行训练操作的容器会执行。此容器可以访问存储在 S3 上的数据。所以我们需要将用来训练的数据上传到 S3。我们可以使用 SageMaker API 完成这一步,它会在后台自动处理一些步骤。
#
# ### 将数据保存到本地
#
# 首先,我们需要创建训练和验证 csv 文件,并将这些文件上传到 S3。
# This is our local data directory. We need to make sure that it exists.
data_dir = '../data/boston'
if not os.path.exists(data_dir):
os.makedirs(data_dir)
# +
# We use pandas to save our train and validation data to csv files. Note that we make sure not to include header
# information or an index as this is required by the built in algorithms provided by Amazon. Also, it is assumed
# that the first entry in each row is the target variable.
pd.concat([Y_val, X_val], axis=1).to_csv(os.path.join(data_dir, 'validation.csv'), header=False, index=False)
pd.concat([Y_train, X_train], axis=1).to_csv(os.path.join(data_dir, 'train.csv'), header=False, index=False)
# -
# ### 上传到 S3
#
# 因为目前正在 SageMaker 会话中运行,所以可以使用代表此会话的对象将数据上传到默认的 S3 存储桶中。注意,建议提供自定义 prefix(即 S3 文件夹),以防意外地破坏了其他 notebook 或项目上传的数据。
# +
prefix = 'boston-xgboost-deploy-hl'
val_location = session.upload_data(os.path.join(data_dir, 'validation.csv'), key_prefix=prefix)
train_location = session.upload_data(os.path.join(data_dir, 'train.csv'), key_prefix=prefix)
# -
# ## 第 4 步:训练 XGBoost 模型
#
# 将训练和验证数据上传到 S3 后,我们可以构建 XGBoost 模型并训练它了。我们将使用高阶 SageMaker API 完成这一步,这样的话代码更容易读懂,但是灵活性较差。
#
# 为了构建一个评估器(即我们要训练的对象),我们需要提供训练代码所在的容器的位置。因为我们使用的是内置算法,所以这个容器由 Amazon 提供。但是,容器的完整名称比较长,取决于我们运行所在的区域。幸运的是,SageMaker 提供了一个实用方法,叫做 `get_image_uri`,它可以为我们构建镜像名称。
#
# 为了使用 `get_image_uri` 方法,我们需要向其提供当前所在区域(可以从 session 对象中获得),以及要使用的算法的名称。在此 notebook 中,我们将使用 XGBoost,但是你也可以尝试其他算法。[常见参数](https://docs.aws.amazon.com/sagemaker/latest/dg/sagemaker-algo-docker-registry-paths.html)中列出了 Amazon 的内置算法。
# +
# As stated above, we use this utility method to construct the image name for the training container.
container = get_image_uri(session.boto_region_name, 'xgboost')
# Now that we know which container to use, we can construct the estimator object.
xgb = sagemaker.estimator.Estimator(container, # The name of the training container
role, # The IAM role to use (our current role in this case)
train_instance_count=1, # The number of instances to use for training
train_instance_type='ml.m4.xlarge', # The type of instance ot use for training
output_path='s3://{}/{}/output'.format(session.default_bucket(), prefix),
# Where to save the output (the model artifacts)
sagemaker_session=session) # The current SageMaker session
# -
# 在要求 SageMaker 开始训练作业之前,我们需要设置模型超参数。如果使用 XGBoost 算法,可以设置的超参数有很多,以下只是其中几个。如果你想修改下面的超参数或修改其他超参数,请参阅 [XGBoost 超参数页面](https://docs.aws.amazon.com/sagemaker/latest/dg/xgboost_hyperparameters.html)
xgb.set_hyperparameters(max_depth=5,
eta=0.2,
gamma=4,
min_child_weight=6,
subsample=0.8,
objective='reg:linear',
early_stopping_rounds=10,
num_round=200)
# 完全设置好 estimator 对象后,可以训练它了。我们需要告诉 SageMaker 输入数据是 csv 格式,然后调用 `fit` 方法。
# +
# This is a wrapper around the location of our train and validation data, to make sure that SageMaker
# knows our data is in csv format.
s3_input_train = sagemaker.s3_input(s3_data=train_location, content_type='csv')
s3_input_validation = sagemaker.s3_input(s3_data=val_location, content_type='csv')
xgb.fit({'train': s3_input_train, 'validation': s3_input_validation})
# -
# ## 第 5 步:测试模型
#
# 暂时我们将跳过这一步。我们依然会测试训练过的模型,但是我们将使用部署的模型,而不是设置批转换作业。
#
#
# ## 第 6 步:部署训练过的模型
#
# 将模型拟合训练数据并使用验证数据避免过拟合后,我们可以部署和测试模型了。使用高阶 API 部署模型很简单,我们只需对训练的评估器调用 `deploy` 方法。
#
# **注意:**部署模型是指要求 SageMaker 启动一个计算实例,它将等待接收数据。因此,此计算实例将一直运行,直到你关闭它。这一点很重要,因为部署的端点按照运行时长计费。
#
# 也就是说,**如果你不再使用部署的端点,请关闭!**
xgb_predictor = xgb.deploy(initial_instance_count=1, instance_type='ml.m4.xlarge')
# ## 第 7 步:使用模型
#
# 现在模型已经训练和部署,我们可以向其发送数据并评估结果。因为我们的测试数据集很小,所以调用一次端点就可以发送了。如果测试数据集很大,我们需要拆分数据并分批发送,然后合并结果。
# +
# We need to tell the endpoint what format the data we are sending is in
xgb_predictor.content_type = 'text/csv'
xgb_predictor.serializer = csv_serializer
Y_pred = xgb_predictor.predict(X_test.values).decode('utf-8')
# predictions is currently a comma delimited string and so we would like to break it up
# as a numpy array.
Y_pred = np.fromstring(Y_pred, sep=',')
# -
# 为了查看模型的运行效果,我们可以绘制一个简单的预测值与真实值散点图。如果模型的预测完全准确的话,那么散点图将是一条直线 $x=y$。可以看出,我们的模型表现不错,但是还有改进的余地。
plt.scatter(Y_test, Y_pred)
plt.xlabel("Median Price")
plt.ylabel("Predicted Price")
plt.title("Median Price vs Predicted Price")
# ## 删除端点
#
# 因为我们不再使用部署的端点,所以需要关闭端点。注意,部署的端点按照运行时长计费,所以运行越久,费用越多。
xgb_predictor.delete_endpoint()
# ## 可选步骤:清理数据
#
# SageMaker 上的默认 notebook 实例没有太多的可用磁盘空间。当你继续完成和执行 notebook 时,最终会耗尽磁盘空间,导致难以诊断的错误。完全使用完 notebook 后,建议删除创建的文件。你可以从终端或 notebook hub 删除文件。以下单元格中包含了从 notebook 内清理文件的命令。
# +
# First we will remove all of the files contained in the data_dir directory
# !rm $data_dir/*
# And then we delete the directory itself
# !rmdir $data_dir
# -
| Tutorials/Boston Housing - XGBoost (Deploy) - High Level.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Visualisation of BubbleColumn testing data and Covid19 data
#
# Now we will look a bit deeper into different visualisation options.
#
# Again we will start by reading in the files with read_excel and read_csv.
#
# ## Our data for plotting:
# 1. Bubble column test data (combined data from 3 testruns)
# 2. RKI Covid19 data
import pandas as pd
# ### Read bubblecolumn excel files and combine into one pandas dataframe
df_bub1 = pd.read_excel("../Data/BubbleColumn/Test_01.xlsx",header=[0,1])
df_bub2 = pd.read_excel("../Data/BubbleColumn/Test_02.xlsx",header=[0,1])
df_bub3 = pd.read_excel("../Data/BubbleColumn/Test_03.xlsx",header=[0,1])
df_bub=pd.concat([df_bub1,df_bub2,df_bub2],keys=["Test1","Test2","Test3"],axis=0,names=["Param","Row_Index"],ignore_index=False)
df_bub
df_bub.index.get_level_values(level=0)
# We convert the Param multiindex into an additional column (easier for filtering)
df_bub.reset_index(level=0,inplace=True)
df_bub
# Now we do some data exploration to check for non-numerical or missing values and to check whether the data is as we expect it.
df_bub.describe()
# Let's check the multiindex column names
df_bub.columns.values
# ### We also read in RKI Covid19 data as an example for timeseries and categorical data
df_rki=pd.read_csv("https://www.arcgis.com/sharing/rest/content/items/f10774f1c63e40168479a1feb6c7ca74/data")
df_rki
# Some preprocessing of RKI data to get official results:
# Is the data up to date?
print(df_rki["Datenstand"].unique())
df_rki_temp = df_rki[((df_rki["NeuerFall"]==0) | (df_rki["NeuerFall"]==1))]
# ## Some general tips:
# Always very useful:
# - Data Dictionary: Metadata for your column names. Explanations, units, etc
# - Data Catalogue: Catalogue for with metadata and storage paths for your testing data.
# ## Overview of python visualisation packages
#
# - matplotlib
#
# Widely used visualisation library. Easy to use and has a good online community presence.
#
#
# - pandas built-in plotting library
#
# Single line command to plot the dataframe. Easier to plot scatterplotmatrix using this library compared to matplotlib and bokeh.
#
#
# - bokeh
#
# Visualisations are more appealing and has built in plot configuration tools (zoom in, pan, etc). But takes time to load the visualisation and it is more suited for creating dashboard. Moreover, the documentation is not clear
#
# - seaborn
#
# Plotting based on matplotlib, but with lots of nice preformatting. Optimized for statistical, dataframe plotting
#
# - plotly
#
# Can do contourplots and 3D plots
#
# - altair / Vega-Lite
#
# Statistical visualization library, preformatted. Minimum amount of code required for nice plotting results
#
# ### Keep in mind:
#
# - Check the documentation of the module by using help() function or the ? in front of the function call!
# - Questions, Problems? --> Google! --> One of the best sources is stackoverflow
# - Module features are dependent on the module version! Check your version:
# ```
# import matplotlib
# matplotlib.__version__
# ```
#
# +
# import neessary libraries:
import matplotlib.pyplot as plt
import bokeh.plotting as bp # another plotting option
from bokeh.plotting import figure,output_notebook,show # for plotting
import seaborn as sb # yeat another plotting option
import plotly.express as px
import altair as alt
# interactive widgets
import ipywidgets as widgets # interactive notebooks - make selection etc
from IPython.display import display # to display the widgets in notebook
# some more useful stuff:
import os
import datetime
# -
# ## Plot histograms
# In order to understand the typical distributions of values, you can always start with a histogram
#
# We start with our BubbleColumn testing data
#
# We wil compare the histogram plots from matplotlib library and pandas built-in
# +
# 1. df_test - matplotlib
plt.figure(figsize=(15,5))
plt.hist(df_bub['cam0', 'Max Feret Diameter'].dropna(),bins=25, color='green',alpha=0.7) # Remember to dropna!
plt.xlabel('Max Feret Diameter')
plt.title('Max Feret Diameter')
plt.show()
# 2. df_test - matplotlib: Also applying some filtering to zoom into a smaller range
plt.figure(figsize=(15,5))
plt.hist(df_bub[((df_bub['cam0', 'Max Feret Diameter']>1.2) & (df_bub['cam0', 'Max Feret Diameter']<6.0))]['cam0', 'Max Feret Diameter'].dropna(),bins=25, color='grey',alpha=0.7) # Remember to dropna!
plt.xlabel('Max Feret Diameter')
plt.title('Max Feret Diameter')
plt.show()
# -
# 2. df_test - pandas built-in
df_bub.plot(y=("cam0",'Max Feret Diameter'),kind="hist",bins=25,color="green",alpha=0.7,figsize=(15,5),title='Max Feret Diameter')
# ## How can histograms be extremely valuable?
# With the help of histograms you can already get an idea about outliers:
#
# If you have the data from multiple tests and you want to know how one specific test compares to the overall amount of tests.
help(plt.hist)
# Plot 2 overlaying histograms for comparison.
# To be able to do so, we also need to add the density keyword! Otherwise the bins of the one test will be much much smaller.
plt.figure(figsize=(15,5))
plt.hist(df_bub['cam0', 'Max Feret Diameter'].dropna(),bins=20,density=True, color='blue',label="All tests")
plt.hist(df_bub[df_bub["Param"]=="Test1"]['cam0', 'Max Feret Diameter'].dropna(),bins=20, density=True,color='orange',alpha= 0.35, label="Test 1")
plt.xlabel('Max Feret Diameter')
plt.title('Comparison of one test with the overall amount of test')
plt.legend()
plt.show()
# ## Some simple scatter / line plots
# ### Created in a loop with filtering of a large dataframe
#
# 1. Example: Bubble testing data
# 2. Example: RKI Covid10 cases for different Landkreise
#
# Advantages of a scatter plot over a line plot:
# Whenever you look at a distribution or a change over time, you are not able to see the intensity / density of the datapoints, if you just do a line plot. To get a feeling for the data, it is always better to start with 'point' as marker instead of 'line'
df_bub[df_bub["Param"]=="Test3"]["erg","z_bild "]
# +
plt.figure(figsize=(15,5))
for i in df_bub["Param"].unique():
print(i)
df_temp=df_bub[df_bub["Param"]==i]
x=df_temp["erg","Zeit [ms]"]
y=(df_temp["erg","z_bild "].shift(1)-df_temp["erg","z_bild "])/(df_temp["erg","t_Bilder LabV"].shift(1)-df_temp["erg","t_Bilder LabV"])
plt.scatter(x,y,label=i)
plt.legend()
plt.ylim(0,0.5)
# -
# ## Interactive selection widgets:
# Another option to get the plots for different tests interactively:
#
# In this minimal example you have to run the plot command every time yoiu have changed the Dropdwon values. But of course you can also add a so-called callback to renew the plot automatically, when a dropdown value changes.
# check it out: widget.observe
# At first we create the selection widget for the Testrun
Test_selection=widgets.Dropdown(options=df_bub["Param"].unique(), value="Test2", description="Select one test")
display(Test_selection)
# Then we create the selection widget for the 0th level of the multiindex columns:
Parameter1_selection=widgets.Dropdown(options=df_bub.columns.get_level_values(level=0).unique(), value="cam0", description="Select one parameter")
display(Parameter1_selection)
# Then we create the selection widget for the 1st level of the multiindex columns:
Parameter2_selection=widgets.Dropdown(options=df_bub.loc[:,pd.IndexSlice[["cam0"], :]].columns.get_level_values(1).unique(),
description="Select one parameter")
display(Parameter2_selection)
# +
plt.figure(figsize=(15,5))
df_temp=df_bub[df_bub["Param"]==Test_selection.value]
x=df_temp["erg","Zeit [ms]"]
y=df_temp[Parameter1_selection.value,Parameter2_selection.value]
plt.scatter(x,y,label=str(Parameter1_selection.value)+", "+str(Parameter2_selection.value))
plt.legend()
# -
# ## Now lets have a look at the same plot with different packages
# ### Bokeh --> Interactive plots
# Try the different menu options you can see at the right side of the plot
# This commands let you visualise bokeh below the execution cell
output_notebook()
# +
df_temp=df_bub[df_bub["Param"]==Test_selection.value]
x=df_temp["erg","Zeit [ms]"]
y=df_temp[Parameter1_selection.value,Parameter2_selection.value]
# 1. Bokeh
p = figure(title="Parameter Selection {}, {} for {}".
format(Parameter1_selection.value,Parameter2_selection.value,Test_selection.value),x_axis_type='datetime',
width=800,height=250)
p.circle(x=x,
y=y)
show(p)
# -
# ### Seaborn
# Not interactive, but preformatted for a nice appearance
# +
df_temp=df_bub[df_bub["Param"]==Test_selection.value]
x=df_temp["erg","Zeit [ms]"]
y=df_temp[Parameter1_selection.value,Parameter2_selection.value]
plt.figure(figsize=(15,5))
sb.scatterplot(x,y)
# -
# ### Plotly
# Interactive plots. Here you can see the single values when hovering over the points.
#
# With plotly you can also do 3D plots!
# +
df_temp=df_bub[df_bub["Param"]==Test_selection.value]
x=df_temp["erg","Zeit [ms]"]
y=df_temp[Parameter1_selection.value,Parameter2_selection.value]
fig = px.scatter(x=x, y=y)
fig.show()
# -
# ### And now as well Altair
# +
# Here we need some additional extensions, so maybe you need to install some additional packages to be able to display the plot.
df_temp=df_bub[df_bub["Param"]==Test_selection.value]
x=df_temp["erg","Zeit [ms]"]
y=df_temp[Parameter1_selection.value,Parameter2_selection.value]
chart=alt.Chart(x=x, y=y).interactive()
chart.show()
# -
# ## Doing some line plots in a loop
# Lets look at the Covid19 case numbers for each Landkreis
# +
# Plotting the casenumbers
fig=plt.figure(figsize=(12,10))
ax1=fig.add_subplot(111)
df_rki_lk=df_rki_temp.groupby(["Landkreis","Meldedatum"],as_index=False)[["AnzahlFall"]].sum()
for i in df_rki_lk["Landkreis"].unique():
df=df_rki_lk[df_rki_lk["Landkreis"]==i]
df.set_index("Meldedatum", inplace=True, drop=True)
df.index=pd.to_datetime(df.index,format="%Y-%m-%d")
df.sort_index(inplace=True)
ax1.plot(df["AnzahlFall"],color="grey",alpha=0.3)
if "Berlin" in i:
df_b=df
ax1.plot(df_b["AnzahlFall"],color="red",label="Berlin")
plt.yscale("log")
plt.title("Casenumbers - Reporting Date - for each Landkreis in Germany")
# -
df_rki_lk["Landkreis"].unique()
# ### Create mulitple subplots
#
# Plot different Landkreise in subplots
# +
plt.figure(figsize=(15,7))
plt.subplot(2,2,1)
plt.plot(df_b["AnzahlFall"],'.')
plt.plot(df_b["AnzahlFall"],'-', color="grey", alpha=0.5)
plt.title("Subplot1 - Berlin")
plt.subplot(2,2,2)
plt.plot(df_rki_lk[df_rki_lk["Landkreis"]=="LK Darmstadt-Dieburg"]["AnzahlFall"],'.')
plt.plot(df_rki_lk[df_rki_lk["Landkreis"]=="LK Darmstadt-Dieburg"]["AnzahlFall"],'-', color="grey", alpha=0.5)
plt.title("Subplot 2 - LK Darmstadt-Dieburg")
plt.subplot(2,2,3)
plt.plot(df_rki_lk[df_rki_lk["Landkreis"]=="LK Friesland"]["AnzahlFall"],'.')
plt.plot(df_rki_lk[df_rki_lk["Landkreis"]=="LK Friesland"]["AnzahlFall"],'-', color="grey", alpha=0.5)
plt.title("Subplot 3 - LK Friesland")
plt.subplot(2,2,4)
plt.plot(df_rki_lk[df_rki_lk["Landkreis"]=="LK Heinsberg"]["AnzahlFall"],'.')
plt.plot(df_rki_lk[df_rki_lk["Landkreis"]=="LK Heinsberg"]["AnzahlFall"],'-', color="grey", alpha=0.5)
plt.title("Subplot 4 - LK Heinsberg")
plt.show()
# -
# ## Create a Correlation / Scatterplot matrix
#
# "A scatter plot matrix is a grid (or matrix) of scatter plots used to visualize bivariate relationships between combinations of variables. Each scatter plot in the matrix visualizes the relationship between a pair of variables, allowing many relationships to be explored in one chart."
# (https://pro.arcgis.com/en/pro-app/latest/help/analysis/geoprocessing/charts/scatter-plot-matrix.htm)
# for simplicity we just look at a smaller df with just one level of column indices
df_temp=df_bub[df_bub["Param"]=="Test1"]
df_temp=df_temp.loc[:,pd.IndexSlice[["cam0"], :]]
df_temp.columns.values
df_temp=df_temp[[('cam0', 'Waddel Disk Diameter'),('cam0', 'equi Ellipse Minor'),('cam0', 'Max Feret Diameter'),
('cam0', 'equi Ellipse Minor Axis (Feret)'),('cam0', 'equi Rect Short Side (Feret)')]]
pd.plotting.scatter_matrix(df_temp, figsize=(15, 15), marker='o',
hist_kwds={'bins': 20}, s=1, alpha=.25)
plt.show()
# ### Links to visualisation examples and more:
#
# * Finding the right diagram
# https://www.visual-literacy.org/periodic_table/periodic_table.html
# * Finding the right colormap for our visualisation
# http://colorbrewer2.org/#type=sequential&scheme=BuGn&n=3
# * More visualisation examples:
# https://d3js.org/
# https://docs.bokeh.org/en/latest/docs/gallery.html
# * Broad overview of vsrious tools available in python
# https://github.com/EthicalML/awesome-production-machine-learning
# * Need multiple y-axis?
# https://matplotlib.org/3.1.1/gallery/ticks_and_spines/multiple_yaxis_with_spines.html
| DataVisualization/2_Visualization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Catching feels
# ## Mood prediction pipeline
#
# An elaborate explanation of this experiment is available at http://medium.com/maslo/catching-feels
# +
import re
import os
import random
import sys
import math
import joblib
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import sklearn.ensemble
from sklearn.metrics import *
from sklearn.model_selection import train_test_split
from sklearn.model_selection import RandomizedSearchCV
# -
sns.set(style="white", palette='muted', context='paper')
def get_precision(y_pred, y_true, decimal_places=False):
'''Returns AUC, precision, recall, and accuracy
Input:
y_pred: predicted values of y
y_true: true values of y
Output:
AUC, precision, recall, and accuracy'''
fpr, tpr, _ = roc_curve(y_true, y_pred)
acc_all = accuracy_score(y_true, y_pred)
precision_all = precision_score(y_true, y_pred)
tpr_all = recall_score(y_true, y_pred)
aucs_all = auc(fpr, tpr)
if decimal_places:
acc_all = np.round(acc_all, decimal_places)
aucs_all = np.round(aucs_all, decimal_places)
precision_all = np.round(precision_all, decimal_places)
tpr_all = np.round(tpr_all, decimal_places)
return aucs_all, precision_all, tpr_all, acc_all
# # Loading data
training_summary_data = pd.read_csv(
'/Users/afrah/Desktop/catchingfeels/data/training_summary_data0622.csv')
# keeping track of performance across models
performances = []
# The summary statistics (Median, Variance, Minimum, Maximum) for the following features were used as training features for all prediction models
# * Polynomial coefficients
# * Mel-frequency cepstral coefficients
# * Spectral contrast
# * Spectral centroid
# * Spectral flatness
# * Spectral rolloff
# * Root mean square
# * Zero crossing rate
# +
feature_names = ['mfcc' + str(i) for i in np.arange(20)] + [
'poly' + str(i) for i in np.arange(11)
] + ['spectral_bw', 'spectral_centroid'] + [
'spectral_cnst' + str(i) for i in np.arange(7)
] + ['spectral_flat', 'spectral_rolloff', 'rms', 'zcr']
all_feature_names = []
for j in ['median', 'min', 'max', 'var']:
all_feature_names += [str.join('_', [j, i]) for i in feature_names]
# -
X = np.array(training_summary_data[all_feature_names])
y = training_summary_data.Sex
codes, uniques = pd.factorize(training_summary_data.Actor)
groups_kfold = sklearn.model_selection.GroupShuffleSplit(test_size=.1, train_size=.9,
n_splits=2,
random_state=10)
groups_kfold.get_n_splits(X, y, codes)
train_inds = None
test_inds = None
for train_index, test_index in groups_kfold.split(X, y, codes):
train_inds = train_index
test_inds = test_index
train_status = np.zeros(X.shape[0])
train_status[train_inds] = 1
training_summary_data['train_status'] = pd.Series(train_status)
# +
# training_summary_data[training_summary_data.train_status==False].shape
# -
print(len(test_inds))
training_summary_data[training_summary_data['train_status']==0].groupby(['Emotion','Sex']).count()
# # Sex prediction model
# The purpose of this prediction model is to predict Gender/Sex of the individuals in the dataset based on the training features used in the models. As a gender-specific effect is observed in mood prediction, this model will allow predicting the gender/sex of future Maslo recordings and be used to make appropriate gender-based predictions
X = np.array(training_summary_data[all_feature_names])
y = training_summary_data.Sex
# The summary statistics (Median, Variance, Minimum, Maximum) for the following features were used as training features for all prediction models
# * Polynomial coefficients
# * Mel-frequency cepstral coefficients
# * Spectral contrast
# * Spectral centroid
# * Spectral flatness
# * Spectral rolloff
# * Root mean square
# * Zero crossing rate
# ## Parameter tuning
#
# Parameter tuning is performed on 50% of the training data set to prevent over-fitting to the current dataset.
# +
# parameter_grid = {
# 'n_estimators': [100, 300, 500, 700, 1000, 1500, 2000],
# 'max_features': ['auto', 'sqrt'],
# 'max_depth': [50, 100, 300, 500, 1000, 1500, 2000, None]
# }
# +
# split_i = sklearn.model_selection.GroupShuffleSplit(test_size=.5,
# train_size=0.5,
# n_splits=2)
# split_i.get_n_splits(X, y, codes2)
# clf = RandomizedSearchCV(
# sklearn.ensemble.RandomForestClassifier(random_state=42),
# parameter_grid,
# random_state=42)
# for train_index, test_index in groups_kfold.split(X, y, codes2):
# search = clf.fit(X[train_index,:], y[train_index,:])
# print(search.best_params_)
# break
# -
# ## Cross validation
# Crossvalidation is used to quantify the performance of the model. Repeated Cross-validation (5 repeats, 5 splits = 25 total splits) is used as the dataset is quite small for a problem so complex and thorough cross-validation is needed to quantify the performance of the models.
X_train = X[train_inds, :]
X_test = X[test_inds, :]
y_train = y[train_inds]
y_test_sex = y[test_inds]
sex_rf = sklearn.ensemble.RandomForestClassifier(n_estimators=2000,
max_depth=1000,
max_features='sqrt',
random_state=42).fit(
X_train, y_train)
y_pred_test_sex = sex_rf.predict(X_test)
# ## Feature prediction
plt.figure(dpi=300)
features_importance = pd.Series(sex_rf.feature_importances_,
index=all_feature_names).nlargest(25)
ax = sns.barplot(x=list(features_importance),
y=[str(i) for i in features_importance.index])
ticks = ax.get_yticklabels()
ax.set_yticklabels(ticks, fontsize=5)
plt.title('Gender prediction model')
plt.xlabel('Feature importance')
# # Mood prediction
# ## Gender-nonspecific prediction model
#
# To predict moods, we focus on predicting 'happy', 'angry', 'neutral' and 'sad' labels due to the limited training samples. This model doesnt take the gender/sex of the person into account and makes predictions for all samples
# +
# # Can we predict mood?
training_sub = training_summary_data[
training_summary_data.Emotion.apply( # Excluding other emotions
lambda x: x not in ['surprise', 'disgust', 'fearful', 'calm'])]
X = np.array(training_sub[all_feature_names])
y = np.array(training_sub.Emotion)
codes, uniques = pd.factorize(training_sub['Actor'])
training_sub['pSex'] = sex_rf.predict(X)
# predicted sex
is_female = np.array(training_sub['pSex']=='Female')
# -
# ### Parameter tuning
# +
# X_train, X_test, y_train, y_test = train_test_split(X,
# y,
# test_size=0.5,
# train_size=0.5)
# clf = RandomizedSearchCV(
# sklearn.ensemble.RandomForestClassifier(random_state=42),
# parameter_grid,
# random_state=42)
# search = clf.fit(X_train, y_train)
# print(search.best_params_)
# -
# ### Cross validation
performances = []
for k in range(3):
print(k)
groups_kfold = sklearn.model_selection.GroupShuffleSplit(test_size=.1,
train_size=0.9,
n_splits=2)
# codes used for group splits to avoid the same actor being in train/test splits
groups_kfold.get_n_splits(X, y, codes)
for train_index, test_index in groups_kfold.split(X, y, codes):
female_inds = train_index[is_female[train_index]]
male_inds = train_index[np.where(is_female[train_index]==False)]
female_test_inds = np.where(is_female[test_index]==True)
X_train = X[train_index, :]
X_test = X[test_index, :]
y_train = y[train_index]
y_test = y[test_index]
X_male = X[male_inds,:]
y_male = y[male_inds]
X_female = X[female_inds,:]
y_female = y[female_inds]
print(X_female.shape, X_train.shape, X_male.shape)
mood_mf8_rf = sklearn.ensemble.RandomForestClassifier(
n_estimators=500,
max_depth=1000,
max_features='sqrt',
random_state=42).fit(X_train, y_train)
mood_m8_rf = sklearn.ensemble.RandomForestClassifier(
n_estimators=2000,
max_depth=1000,
max_features='sqrt',
random_state=42).fit(X_male, y_male)
mood_f8_rf = sklearn.ensemble.RandomForestClassifier(
n_estimators=1500,
max_depth=300,
max_features='sqrt',
random_state=42).fit(X_female, y_female)
y_pred_test_mf8 = mood_mf8_rf.predict(X_test)
y_pred_test_m8 = mood_m8_rf.predict(X_test)
y_pred_test_f8 = mood_f8_rf.predict(X_test)
y_pred = y_pred_test_mf8.copy()
for i, val in enumerate(y_pred_test_mf8):
if y_pred_test_f8[i] in =='angry':
y_pred[i] = 'angry'
elif y_pred_test_m8[i] in =='sad':
y_pred[i] = 'sad'
elif i in list(female_test_inds[0]) and y_pred_test_f8[i] in =='happy':
y_pred[i] = 'happy'
# else:
# if y_pred_test_m8[i] in ['angry','sad']:
# y_pred[i] = y_pred_test_m8[i]
print(sum(y_pred!=y_pred_test_mf8))
for mood in np.unique(y_test):
performances += [[mood] + list(
get_precision(np.array(pd.get_dummies(y_pred)[mood]),
np.array(pd.get_dummies(y_test)[mood]), 2)) +
['gender-specific-model']]
performances += [[mood] + list(
get_precision(np.array(pd.get_dummies(y_pred_test_m8)[mood]),
np.array(pd.get_dummies(y_test)[mood]), 2)) +
['males-specific-model']]
performances += [[mood] + list(
get_precision(np.array(pd.get_dummies(y_pred_test_f8)[mood]),
np.array(pd.get_dummies(y_test)[mood]), 2)) +
['females-specific-model']]
performances += [[mood] + list(
get_precision(np.array(pd.get_dummies(y_pred_test_mf8)[mood]),
np.array(pd.get_dummies(y_test)[mood]), 2)) +
['gender-nonspecific-model']]
sum(y_pred_test_mf8[female_test_inds]!=y_pred_test_f8[female_test_inds])
# +
# -
y_pred = y_pred_test_mf8
for i, val in enumerate(y_pred_test_mf8):
if i in list(female_test_inds[0]):
if y_pred_test_f8[i] in ['anger','happy']:
y_pred[i] = y_pred_test_f8[i]
else:
if y_pred_test_m8[i] in ['anger','happy']:
y_pred[i] = y_pred_test_m8[i]
print(y_pred_test_m8[y_pred_test_m8!=y_pred_test_mf8])
print(y_pred_test_mf8[y_pred_test_m8!=y_pred_test_mf8])
# ### Features importance
plt.figure(dpi=300)
features_importance = pd.Series(mood_mf8_rf.feature_importances_,
index=all_feature_names).nlargest(25)
ax = sns.barplot(x=list(features_importance),
y=[str(i) for i in features_importance.index])
ticks = ax.get_yticklabels()
ax.set_yticklabels(ticks, fontsize=5)
plt.title('Gender-nonspecific model')
plt.xlabel('Feature importance')
# ### Features importance
plt.figure(dpi=300)
features_importance = pd.Series(mood_m8_rf.feature_importances_,
index=all_feature_names).nlargest(25)
ax = sns.barplot(x=list(features_importance),
y=[str(i) for i in features_importance.index])
ticks = ax.get_yticklabels()
ax.set_yticklabels(ticks, fontsize=5)
plt.title('Male-specific model')
plt.xlabel('Feature importance')
# ### Features importance
plt.figure(dpi=300)
features_importance = pd.Series(mood_f8_rf.feature_importances_,
index=all_feature_names).nlargest(25)
ax = sns.barplot(x=list(features_importance),
y=[str(i) for i in features_importance.index])
ticks = ax.get_yticklabels()
ax.set_yticklabels(ticks, fontsize=5)
plt.title('Female-specific model')
plt.xlabel('Feature importance')
# # Comparison of models' performance
df_performances = pd.DataFrame(performances)
df_performances.columns = [
'Prediction', 'AUC', 'Precision', 'Recall', 'Accuracy', 'Model'
]
df_performances.to_csv('./df_performances07120-0.9.csv.gz', index=False)
# ## Mood predictions
# +
# df_performances.to_csv('./df_performances20splits_final.csv.gz', index=False)
# -
plt.figure(dpi=200, figsize=(10, 5))
sns.boxplot(x='Prediction',
y='AUC',
hue='Model',
data=df_performances[df_performances.Model != 'sex-prediction'],
palette='Set2')
plt.savefig('../figs/moodmodels_auc.pdf', dpi=300)
plt.figure(dpi=200, figsize=(10, 5))
sns.boxplot(x='Prediction',
y='Precision',
hue='Model',
data=df_performances[df_performances.Model != 'sex-prediction'],
palette='Set2')
plt.savefig('../figs/moodmodels_precision.pdf', dpi=300)
# ## Sex prediction
# +
plt.figure(dpi=300, figsize=(15, 10))
sns.barplot(x='Prediction',
y='AUC',
hue='Prediction',
data=df_performances[df_performances.Model == 'sex-prediction'],
palette='Set2')
plt.savefig('../figs/sex_auc.pdf', dpi=300)
plt.figure(dpi=300, figsize=(15, 10))
sns.barplot(x='Prediction',
y='Precision',
hue='Prediction',
data=df_performances[df_performances.Model == 'sex-prediction'],
palette='Set2')
plt.savefig('../figs/sex_precision.pdf', dpi=300)
# -
# ## Computing median AUC across models-predictions
df_performances.groupby(['Model', 'Prediction']).median()
# 7% angry female, 4% angry male
# 6.5% happy, -0.007% male happy
# 8.3% neutral female; 0% male neutral
# 2% sad; 5% male sad
#
# 8% preciision female, 3%male
# 20%happy; -10%male
# 4% neutral
#
(.84-.81)/.81
# # Comparison of top features across models
#
# Higher MFCCs were more important in female-specific models than male-specific/gender-specific models.
features_arr = np.vstack([
mood_mf8_rf.feature_importances_, mood_m8_rf.feature_importances_,
mood_f8_rf.feature_importances_
])
features_df = pd.DataFrame(features_arr)
features_df.columns = all_feature_names
features_df.index = ['mf', 'm', 'f']
# +
dict_features = {}
for i in all_feature_names:
if 'mfcc' in i:
dict_features[i] = 'darkblue'
elif 'poly' in i:
dict_features[i] = 'orange'
elif 'rms' in i:
dict_features[i] = 'darkgray'
elif 'bw' in i:
dict_features[i] = 'y'
elif 'zcr' in i:
dict_features[i] = 'turquoise'
elif 'rolloff' in i:
dict_features[i] = 'purple'
elif 'flat' in i:
dict_features[i] = 'pink'
elif 'cnst' in i:
dict_features[i] = 'pink'
else:
dict_features[i] = 'cyan'
top_features = np.unique(
list(
pd.Series(mood_mf8_rf.feature_importances_,
index=all_feature_names).nlargest(15).index) + list(
pd.Series(mood_f8_rf.feature_importances_,
index=all_feature_names).nlargest(15).index) +
list(
pd.Series(mood_m8_rf.feature_importances_,
index=all_feature_names).nlargest(15).index))
# -
df = features_df[top_features]
df = df.transpose()
row_colors = df.index.map(dict_features)
sns.clustermap(df, row_colors=row_colors)
plt.savefig('../figs/heatmap_moodmodels.pdf', dpi=300)
import pandas as pd
df = pd.read_csv('./df_performances20splits_final.csv.gz')
df.groupby(['Model', 'Prediction'])
(.82-.74)/.74
| notebooks/.ipynb_checkpoints/Prediction models-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.7 64-bit (''venv'': venv)'
# language: python
# name: python37764bitvenvvenvd33f9026929c40169e2f7d2ac2b15d20
# ---
# # Underscores, Dunders, and More
#
# ### Single leading underscore: \_var
# The underscore prefix is meant as a hint to tell another programmer that a variable or method starting with a single underscore is intended for internal user. This convention is defined in PEP8, the most commonly used Python code style guide
#
# However, this convention isn't enforced by the Python interpreter. Python doesn't have strong distinctions between '*private*' and '*public*' variables
#
# +
class Test:
def __init__(self):
self.foo = 11
self._bar = 23
t = Test()
print(f'Public argument {t.foo}')
print(f'Private argument {t._bar}')
# -
# As you can see, the leading single underscore in _bar did not prevent us from 'reaching into' the class and accessing the value of that variable.
#
# That's because the single underscore prefix in Python is merely an agreed-upon convention-at least when it comes to variable and method names.
#
# Now, if you use a wildcard import to import all the names from the module, Python will not import names with a leading underscore
# +
from resources.mymodule import *
print(external_func())
print(_internal_func())
# -
# Unlike wildcard imports, regular imports are not affected by the leading single underscore naming convetion
# +
import resources.mymodule as mymodule
print(mymodule.external_func())
print(mymodule._internal_func())
# -
# If you stick to the PEP8 recommendation that wildcard imports should be avoided, the all you really need to remember is this:
#
# Single underscores are a Python naming convention that indicates a name is meant for internal use. It is generally not enforced by the Python interpreter and is only meant as a hint to the programmer.
#
# ### Single trailing underscores: var\_
#
# Sometimes the most fitting name for a variables is already taken by keyword in the Python language. Therefore, names like class or def canot be used as variable names in Python. In this case you can append a single underscore to break the naming conflict
# +
def make_object(name, class):
return class
make_object('name', 'class')
# +
def make_object(name, class_):
return class_
make_object('name', 'class')
# -
# ### Double leading underscore: \_\_var
#
# A double underscore prefix causes the Python interpreter to rewrite the attribute name in order to avoid naming conflicts in subclasses. This is also called name mangling the interpreter changes the name of the variable in a way that makes it harder to create collisions when the class is extended later.
#
# +
class Test:
def __init__(self):
self.foo = 11
self._bar = 23
self.__baz = 42
t = Test()
dir(t)
# -
# If you look closely, you'll see there's an attribute called _Test__baz on this object. This is the name mangling that the Python interpreter applies. It does this to protect the variable from getting overriden in subclasses. We are going to create another class that extends the Test class and attempts to override its existing attributes added in the constructor.
# +
class ExtendedTest(Test):
def __init__(self):
super().__init__()
self.foo = 'overridden'
self._bar = 'overridden'
self.__baz = 'overridden'
t2 = ExtendedTest()
print(t2._ExtendedTest__baz)
print(t2._Test__baz)
dir(t2)
# -
# As you can see, \_\_baz got turned into \_ExtendedTest\_\_baz to prevent accidental modification. But the original \_Test\_\_baz is also still around.
#
# Double underscore name mangling is fully transparent to the programmer. Take a took ar the following example that will confirm this. Name mangling affects all names that start with two underscore characters ('dunders') in class context
# +
class ManglingTest:
def __init__(self):
self.__mangled = 'Hello'
def get_mangled(self):
return self.__mangled
def __method(self):
return 42
mangling = ManglingTest()
print(mangling.get_mangled())
print(mangling.__mangled)
# -
mangling.__method()
# Here's another, perhaps surprising, example of name mangling in action:
# +
_ManagledGlobal__mangled = 23
class ManagledGlobal:
def test(self):
return __mangled
print(ManagledGlobal().test())
# -
# The Python interpreter automatically expanded the name __mangled to _ManagledGlobal__managled because it begins with two underscore characters. This demonstrates that name mangling isn't tied to class attributes specifically. It applies to any name starting with two underscore characters that is used in a class context.
#
# Double underscores are often referred to as 'dunders' in the Python community. The reason is that double underscores appear quite often in Python code, and to avoid fatiguing their jaw muscles, Pythonistas ofthe shorten 'double underscore' to 'dunder'.
#
# But that's just yet another quirk in the naming convention. It's like a secret handshake for Python developers.
#
# ### Double leading and trailing underscore: \_\_var\_\_
#
# Perhaps surprisingly, name mangling is not applied if a name starts and ends with double underscores. Variables surrounded by a double underscore prefix and postfix are left unscathed by the Python interpreter.
#
# Hovewer, names that have both leading and trailing double underscores are reserved for special use in the language. This rule covers things like \_\_init\_\_ for object constructors or \_\_call\_\_ to make objects callable.
#
# These dunder methods are often referred to as *magic methods*. They're a core feature in Python and should be used as needed.
#
# Hovewer, as far as naming conventions go, it's best to stay away from using names that start and end with dunders in your own programs to avoid collisions with future changes to the Python language
#
# ### Single underscore: '\_'
#
# Per convention, a single stand-alone underscore is sometimes used as a name to indicate that a variable is temporary or insignificant.
for _ in range(5):
print('Hello, world.')
# Besides its use as a temporary variable, *\_* is a special variable in most Python REPLs that represents the results of the last expression evaluated by the interpreter.
#
# ```
# >>> list()
# []
# >>> _.append(1)
# >>> _.append(2)
# >>> _.append(3)
# >>> _
# [1, 2, 3]
# ```
| Patterns for Cleaner Python/Underscores.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %matplotlib inline
# +
import pickle
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
from sklearn import svm
from sklearn import cross_validation
from sklearn import grid_search
results = pickle.load(open("results-25perc-noloc.pickle", "rb"))
model_results = results[0]
df = results[1]
predictions = results[2]
best_score = results[3]
best_params = results[4]
best_features = results[5]
best_model = results[6]
print "Loaded"
# -
print "Best score:", best_score
#print "Best features:", best_features
print "Best params:", best_params
# Build a histogram with percentages correct for each category
df_test = df[(df["is_test"] == True)]
df_test["prediction"] = predictions
#print df_test.head()
# Compare the percent correct to the results from earlier to make sure things are lined up right
print "Calculated accuracy:", sum(df_test["label"] == df_test["prediction"]) / float(len(df_test))
print "Model accuracy:", best_score
df_correct = df_test[(df_test["label"] == df_test["prediction"])]
df_incorrect = df_test[(df_test["label"] != df_test["prediction"])]
#df_correct.describe()
#df_test.describe()
#plt.hist(correct_labels)
#print df.describe()
print "Correct predictions:", df_correct.groupby(["label"])["prediction"].count()
print "Incorrect predictions:", df_incorrect.groupby(["label"])["prediction"].count()
# Stats of text length for correct and incorrect
print df_correct.describe()
print df_incorrect.describe()
# +
#print model_results
d3_data = {}
for m in model_results:
d3_data[m["feat_name"]] = {}
d3_data[m["feat_name"]]["C"] = []
d3_data[m["feat_name"]]["G"] = []
d3_data[m["feat_name"]]["S"] = []
#print m["feat_name"], m["model_params"], m["model_score"]
for s in m["grid_scores"]:
d3_data[m["feat_name"]]["C"].append(s[0]["C"])
d3_data[m["feat_name"]]["G"].append(s[0]["gamma"])
d3_data[m["feat_name"]]["S"].append(s[1])
#print d3_data
# +
from mpl_toolkits.mplot3d import Axes3D
from matplotlib import cm
from matplotlib.ticker import LinearLocator, FormatStrFormatter
from matplotlib import pylab
pylab.rcParams['figure.figsize'] = (10.0, 8.0)
def d3_plot(X, Y, Z):
fig = plt.figure()
ax = fig.gca(projection='3d')
ax.set_xlabel("C", weight="bold", size="xx-large")
ax.set_xticks([0, 5000, 10000, 15000])
ax.set_xlim(0, max(X))
ax.set_ylabel("gamma", weight="bold", size="xx-large")
ax.set_yticks([0, 1.5, 3, 4.5])
ax.set_ylim(0, max(Y))
ax.set_zlabel("Accuracy", weight="bold", size="xx-large")
#ax.set_zticks([0.5, 0.6, 0.70])
ax.set_zlim(0.5, 0.75)
ax.scatter(X, Y, Z, c='b', marker='o')
ax.zaxis.set_major_locator(LinearLocator(10))
ax.zaxis.set_major_formatter(FormatStrFormatter('%.02f'))
plt.show()
# -
d3_plot(np.array(d3_data["area"]["C"]), np.array(d3_data["area"]["G"]), np.array(d3_data["area"]["S"]))
d3_plot(np.array(d3_data["line"]["C"]), np.array(d3_data["line"]["G"]), np.array(d3_data["line"]["S"]))
d3_plot(np.array(d3_data["word"]["C"]), np.array(d3_data["word"]["G"]), np.array(d3_data["word"]["S"]))
| results/analize_results.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="-K9U3Ab0c7LY"
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras.applications import MobileNetV2
from tensorflow.keras.layers import AveragePooling2D
from tensorflow.keras.layers import Dropout
from tensorflow.keras.layers import Flatten
from tensorflow.keras.layers import Dense
from tensorflow.keras.layers import Input
from tensorflow.keras.models import Model
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.applications.mobilenet_v2 import preprocess_input
from tensorflow.keras.preprocessing.image import img_to_array
from tensorflow.keras.preprocessing.image import load_img
from tensorflow.keras.utils import to_categorical
from sklearn.preprocessing import LabelBinarizer
from sklearn.model_selection import train_test_split
from sklearn.metrics import classification_report
from imutils import paths
import matplotlib.pyplot as plt
import numpy as np
import argparse
import os
# + colab={"base_uri": "https://localhost:8080/"} id="A8ZLmk6Xdnnz" executionInfo={"status": "ok", "timestamp": 1607151799066, "user_tz": -330, "elapsed": 35738, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgAEsAcdZQ86aK_pCyT1HfwDA3G6lZe8WfhE573Jw=s64", "userId": "00439116246834712963"}} outputId="0821d2e6-5c7f-48e6-c667-387d9ba69b05"
from google.colab import drive
drive.mount('/content/drive/')
# + id="Ew6UDwVKe1AU"
#setting root dir for project
rootdir = '/content/drive/My Drive/Mask Detector/Face Mask Detection/'
# + id="GoARlemtgR2Q"
#setting parameters
dataset = rootdir + 'dataset'
plot = 'out.png'
model = rootdir + 'mask_detector.model' ##DNN based face extractor
# + id="ivJ1Iwq-hOWB"
#gathering hyperparameters
INIT_LR = 1e-4
EPOCHS = 20
BS = 32
# + id="13DpF_entoNh"
def _preprocess_numpy_input(x, data_format, mode):
"""Preprocesses a Numpy array encoding a batch of images.
Arguments:
x: Input array, 3D or 4D.
data_format: Data format of the image array.
mode: One of "caffe", "tf" or "torch".
- caffe: will convert the images from RGB to BGR,
then will zero-center each color channel with
respect to the ImageNet dataset,
without scaling.
- tf: will scale pixels between -1 and 1,
sample-wise.
- torch: will scale pixels between 0 and 1 and then
will normalize each channel with respect to the
ImageNet dataset.
Returns:
Preprocessed Numpy array.
"""
if not issubclass(x.dtype.type, np.floating):
x = x.astype(backend.floatx(), copy=False)
print('Ran x.astype')
if mode == 'tf':
x /= 127.5
x -= 1.
return x
elif mode == 'torch':
x /= 255.
mean = [0.485, 0.456, 0.406]
std = [0.229, 0.224, 0.225]
else:
if data_format == 'channels_first':
# 'RGB'->'BGR'
if x.ndim == 3:
x = x[::-1, ...]
else:
x = x[:, ::-1, ...]
else:
# 'RGB'->'BGR'
x = x[..., ::-1]
mean = [103.939, 116.779, 123.68]
std = None
print('here')
# Zero-center by mean pixel
if data_format == 'channels_first':
if x.ndim == 3:
x[0, :, :] -= mean[0]
x[1, :, :] -= mean[1]
x[2, :, :] -= mean[2]
if std is not None:
x[0, :, :] /= std[0]
x[1, :, :] /= std[1]
x[2, :, :] /= std[2]
else:
x[:, 0, :, :] -= mean[0]
x[:, 1, :, :] -= mean[1]
x[:, 2, :, :] -= mean[2]
if std is not None:
x[:, 0, :, :] /= std[0]
x[:, 1, :, :] /= std[1]
x[:, 2, :, :] /= std[2]
else:
x[..., 0] -= mean[0]
x[..., 1] -= mean[1]
x[..., 2] -= mean[2]
if std is not None:
x[..., 0] /= std[0]
x[..., 1] /= std[1]
x[..., 2] /= std[2]
return x
# + colab={"base_uri": "https://localhost:8080/"} id="_qpzxN8ehnxC" executionInfo={"status": "ok", "timestamp": 1607153918213, "user_tz": -330, "elapsed": 564854, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgAEsAcdZQ86aK_pCyT1HfwDA3G6lZe8WfhE573Jw=s64", "userId": "00439116246834712963"}} outputId="66c43fbf-ce85-4c89-be75-065ac0c75f6a"
# grab the list of images in our dataset directory, then initialize
# the list of data (i.e., images) and class images
print("[INFO] loading images...")
imagePaths = list(paths.list_images(dataset))
data = []
labels = []
i = 0;
print(len(imagePaths))
# loop over the image paths
for imagePath in imagePaths:
label = imagePath.split(os.path.sep)[-2]
print('proessing ' + str(i))
i = i + 1
# load the input image (224x224) and preprocess it
image = load_img(imagePath, target_size=(224, 224))
image = img_to_array(image)
image = _preprocess_numpy_input(image,None,'tf')
# update the data and labels lists, respectively
data.append(image)
labels.append(label)
# + id="9A7uM69WtReL"
data = np.array(data, dtype="float32")
labels = np.array(labels)
# + id="cAhl8T4_tWOp"
# perform one-hot encoding on the labels
lb = LabelBinarizer()
labels = lb.fit_transform(labels)
labels = to_categorical(labels)
# + id="p_s9rNaStaYR"
# the data for training and the remaining 25% for testing
(trainX, testX, trainY, testY) = train_test_split(data, labels,
test_size=0.20, stratify=labels, random_state=42)
# + id="8SRnXmfkt-Kr"
# construct the training image generator for data augmentation
aug = ImageDataGenerator(
rotation_range=20,
zoom_range=0.15,
width_shift_range=0.2,
height_shift_range=0.2,
shear_range=0.15,
horizontal_flip=True,
fill_mode="nearest")
# + colab={"base_uri": "https://localhost:8080/"} id="d3zTWDeFuC9P" executionInfo={"status": "ok", "timestamp": 1607153927698, "user_tz": -330, "elapsed": 560342, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgAEsAcdZQ86aK_pCyT1HfwDA3G6lZe8WfhE573Jw=s64", "userId": "00439116246834712963"}} outputId="db68d816-d4fd-49fc-b3b1-f2a2fe160a50"
# load the MobileNetV2 network, ensuring the head FC layer sets are
# left off
baseModel = MobileNetV2(weights="imagenet", include_top=False,
input_tensor=Input(shape=(224, 224, 3)))
# + id="aP81hXSeuHIY"
# construct the head of the model that will be placed on top of the
# the base model
headModel = baseModel.output
headModel = AveragePooling2D(pool_size=(7, 7))(headModel)
headModel = Flatten(name="flatten")(headModel)
headModel = Dense(128, activation="relu")(headModel)
headModel = Dropout(0.5)(headModel)
headModel = Dense(2, activation="softmax")(headModel)
# + id="DBVXCDJ0uMoX"
# place the head FC model on top of the base model (this will become
# the actual model we will train)
model = Model(inputs=baseModel.input, outputs=headModel)
# + id="--htv6tVuP1z"
# loop over all layers in the base model and freeze them so they will
# *not* be updated during the first training process
for layer in baseModel.layers:
layer.trainable = False
# + colab={"base_uri": "https://localhost:8080/"} id="FtKmy3iUuS9B" executionInfo={"status": "ok", "timestamp": 1607153927705, "user_tz": -330, "elapsed": 552731, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgAEsAcdZQ86aK_pCyT1HfwDA3G6lZe8WfhE573Jw=s64", "userId": "00439116246834712963"}} outputId="31c5950e-37d3-40b5-a4ea-4471fa12e00d"
# compile our model
print("[INFO] compiling model...")
opt = Adam(lr=INIT_LR, decay=INIT_LR / EPOCHS)
model.compile(loss="binary_crossentropy", optimizer=opt,
metrics=["accuracy"])
# + colab={"base_uri": "https://localhost:8080/"} id="EA_IndvuuXRH" executionInfo={"status": "ok", "timestamp": 1607156921874, "user_tz": -330, "elapsed": 38358, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgAEsAcdZQ86aK_pCyT1HfwDA3G6lZe8WfhE573Jw=s64", "userId": "00439116246834712963"}} outputId="9e835e2b-0bb1-4b6b-e57f-f76b0d680b7f"
# train the head of the network
print("[INFO] training head...")
H = model.fit(
aug.flow(trainX, trainY, batch_size=BS),
steps_per_epoch=len(trainX) // BS,
validation_data=(testX, testY),
validation_steps=len(testX) // BS,
epochs=EPOCHS)
# + colab={"base_uri": "https://localhost:8080/"} id="md3Hk9916SSs" executionInfo={"status": "ok", "timestamp": 1607156947285, "user_tz": -330, "elapsed": 25424, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgAEsAcdZQ86aK_pCyT1HfwDA3G6lZe8WfhE573Jw=s64", "userId": "00439116246834712963"}} outputId="9d5faa7e-2641-4916-f4a4-0d60178cc811"
print("[INFO] evaluating network...")
predIdxs = model.predict(testX, batch_size=BS)
# + id="yfE_jJ216fds"
# for each image in the testing set we need to find the index of the
# label with corresponding largest predicted probability
predIdxs = np.argmax(predIdxs, axis=1)
# show a nicely formatted classification report
print(classification_report(testY.argmax(axis=1), predIdxs,
target_names=lb.classes_))
# + colab={"base_uri": "https://localhost:8080/"} id="EyTq60FA6l5E" executionInfo={"status": "ok", "timestamp": 1607156949497, "user_tz": -330, "elapsed": 2217, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgAEsAcdZQ86aK_pCyT1HfwDA3G6lZe8WfhE573Jw=s64", "userId": "00439116246834712963"}} outputId="af78109e-9832-43f7-cf2c-00630901eda7"
# serialize the model to disk
print("[INFO] saving mask detector model...")
model.save('/content/drive/My Drive/Mask Detector/Face Mask Detection/mask_detector.model', save_format="h5")
# + colab={"base_uri": "https://localhost:8080/", "height": 863} id="JLyd6yT_7HQu" executionInfo={"status": "ok", "timestamp": 1607156949507, "user_tz": -330, "elapsed": 45, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgAEsAcdZQ86aK_pCyT1HfwDA3G6lZe8WfhE573Jw=s64", "userId": "00439116246834712963"}} outputId="e19e5e4e-b426-4720-da37-80848007aac1"
# plot the training loss and accuracy
N = EPOCHS
plt.style.use("ggplot")
plt.figure()
plt.plot(np.arange(0, N), H.history["loss"], label="train_loss")
plt.plot(np.arange(0, N), H.history["val_loss"], label="val_loss")
plt.plot(np.arange(0, N), H.history["accuracy"], label="train_acc")
plt.plot(np.arange(0, N), H.history["val_accuracy"], label="val_acc")
plt.title("Training Loss and Accuracy")
plt.xlabel("Epoch #")
plt.ylabel("Loss/Accuracy")
plt.legend(loc="lower left")
plt.savefig(plot)
| Mask_Detector_Trainer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.10 64-bit (''torchEnv'': conda)'
# name: python3
# ---
# + [markdown] id="0yAIJhY1M41M"
# # The COVIDNetX challenge
# + [markdown] id="NQyQqzQTM41O"
# <img src="https://www.psycharchives.org/retrieve/096175aa-f7f2-4970-989d-d934c30b5551" alt="drawing" width="400"/>
# -
# The following is a classification challenge using the COVID-X dataset (https://github.com/lindawangg/COVID-Net/blob/master/docs/COVIDx.md).
# The goal is to predict whether a person has COVID-19 or not based on chest X-RAY images.
#
# There are two different categories: `positive` and `negative`.
# `positive` means a person has COVID-19, `negative` means a person
# has not COVID-19.
#
# The metric we use is F1 (https://en.wikipedia.org/wiki/F1_score). The goal
# is to maximize F1.
#
# The data contains images with their associated labels.
# + [markdown] id="QIeCjvyTM418"
# # Exploratory Data Analysis
# + id="V6gETkD5M418"
import pandas as pd
from IPython.display import Image
data_dir = 'data/'
# data_dir = 'data_subset/'
# -
# Now, you can open the submision.csv file (File -> Open) file and download it!
#
# After you download it, you can upload it to the challenge frontend.
# # Baseline simple solution (Logistic Regression on the top of Resnet50 features)
# In this simple baseline solution, we use the learned representation by a Resnet-50 as features
# and use the logistic regression as a model (no fine-tuning here).
# +
import torch
from PIL import Image
from torch.utils.data import Dataset
import torchvision.transforms as transforms
import torchvision
import time
import os
class CustomDataSet(Dataset):
def __init__(self, filenames, transform=None, labels=None):
self.filenames = filenames
self.labels = labels
self.transform = transform
def __len__(self):
return len(self.filenames)
def __getitem__(self, idx):
image = Image.open(self.filenames[idx]).convert("RGB")
label = self.labels[idx] if self.labels is not None else 0
if self.transform:
tensor_image = self.transform(image)
return tensor_image, label
normalize = transforms.Normalize(
mean=[0.485, 0.456, 0.406],
std=[0.229, 0.224, 0.225]
)
transform = transforms.Compose([
transforms.Resize(256),
transforms.CenterCrop(224),
transforms.ToTensor(),
normalize
])
classname_to_index = {"positive": 1, "negative": 0}
index_to_classname = {i:n for n, i in classname_to_index.items()}
def build_dataset(df, split):
filenames = [os.path.join(split, name) for name in df.image]
labels = [classname_to_index[name] for name in df.label]
dataset = CustomDataSet(filenames, transform=transform, labels=labels)
return dataset
df_train = pd.read_csv(data_dir+'train.csv')
df_test = pd.read_csv(data_dir+'submission_valid.csv')
train_dataset = build_dataset(df_train.sample(frac=0.1), data_dir+"train")
test_dataset = build_dataset(df_test.sample(frac=0.1), data_dir+"valid")
len(train_dataset), len(test_dataset)
# +
batch_size = 100
train_loader = torch.utils.data.DataLoader(
train_dataset,
batch_size=batch_size,
shuffle=True
)
test_loader = torch.utils.data.DataLoader(
test_dataset,
batch_size=batch_size,
shuffle=False
)
# +
from tqdm import tqdm
model = torchvision.models.resnet50(pretrained=True)
def extract_features(model, dataloader):
Flist = []
Ylist = []
for X, Y in tqdm(dataloader):
x = model.conv1(X)
x = model.bn1(x)
x = model.relu(x)
x = model.maxpool(x)
x = model.layer1(x)
x = model.layer2(x)
x = model.layer3(x)
x = model.layer4(x)
x = model.avgpool(x)
x = x.view(x.size(0), x.size(1))
Flist.append(x)
Ylist.append(Y)
F = torch.cat(Flist).detach().numpy()
Y = torch.cat(Ylist).detach().numpy()
return F, Y
# +
# resnet50 = torchvision.models.resnet50(pretrained=True)
# resnet50
# vgg16 = torchvision.models.vgg16(pretrained=True)
# vgg16
# -
# %%time
xtrain, ytrain = extract_features(model, train_loader)
# xtest, ytest = extract_features(model, test_loader)
print(xtrain.shape, ytrain.shape)
# print(xtest.shape, ytest.shape)
import numpy as np
idx = np.random.choice(range(len(xtrain)), int(0.8*len(xtrain)), replace=False)
train_idx = np.zeros(len(xtrain), dtype=bool)
train_idx[idx] = True
test_idx = ~train_idx
# %%time
from sklearn.linear_model import LogisticRegression
clf = LogisticRegression(max_iter=1000,class_weight="balanced")
clf.fit(xtrain[train_idx], ytrain[train_idx])
# +
# ypred_test = clf.predict(xtrain[400:])
# ypred_test = [index_to_classname[y] for y in ypred_test]
# print(ypred_test[0:10])
# df = pd.read_csv(data_dir+'submission_valid.csv')
# df["label"] = ypred_test
# df.head()
# +
import seaborn as sns
from sklearn.metrics import confusion_matrix, classification_report
print("############################## TRAIN ############################## ")
y_pred = clf.predict(xtrain[train_idx])
print(classification_report(ytrain[train_idx], y_pred))
print("############################## TEST ############################## ")
y_pred = clf.predict(xtrain[test_idx])
print(classification_report(ytrain[test_idx], y_pred))
# cm = confusion_matrix(ytrain[test_idx], y_pred)
# sns.heatmap(cm, cmap='plasma', annot=True);
# +
# df.to_csv("submission.csv", index=False)
# -
# Now, you can open the submision.csv file (File -> Open) file and download it!
#
# After you download it, you can upload it to the challenge frontend.
# + [markdown] id="Zr6DPp06p39h"
# # Baseline Solution with Resnet-50 Fine-tuning
# + id="PsC0uqOGtQPb"
#Code adapted from https://github.com/pytorch/examples/tree/master/imagenet
import torch
from PIL import Image
from torch.utils.data import Dataset
import torchvision.transforms as transforms
import time
import os
import shutil
# device = "cpu"
device = "cuda"
def train(train_loader, model, criterion, optimizer, epoch):
batch_time = AverageMeter('Time', ':6.3f')
data_time = AverageMeter('Data', ':6.3f')
losses = AverageMeter('Loss', ':.4e')
top1 = AverageMeter('Acc@1', ':6.2f')
top5 = AverageMeter('Acc@5', ':6.2f')
progress = ProgressMeter(
len(train_loader),
[batch_time, data_time, losses, top1, top5],
prefix="Epoch: [{}]".format(epoch))
# switch to train mode
model.train()
end = time.time()
for i, (images, target) in enumerate(train_loader):
# measure data loading time
data_time.update(time.time() - end)
images = images.to(device)
target = target.to(device)
# compute output
output = model(images)
loss = criterion(output, target)
# measure accuracy and record loss
(acc1,) = accuracy(output, target, topk=(1,))
losses.update(loss.item(), images.size(0))
top1.update(acc1[0], images.size(0))
# compute gradient and do SGD step
optimizer.zero_grad()
loss.backward()
optimizer.step()
# measure elapsed time
batch_time.update(time.time() - end)
end = time.time()
if i % 100 == 0:
progress.display(i)
def validate(val_loader, model, criterion):
batch_time = AverageMeter('Time', ':6.3f')
losses = AverageMeter('Loss', ':.4e')
top1 = AverageMeter('Acc@1', ':6.2f')
top5 = AverageMeter('Acc@5', ':6.2f')
progress = ProgressMeter(
len(val_loader),
[batch_time, losses, top1, top5],
prefix='Test: ')
# switch to evaluate mode
model.eval()
with torch.no_grad():
end = time.time()
for i, (images, target) in enumerate(val_loader):
images = images.to(device)
target = target.to(device)
# compute output
output = model(images)
loss = criterion(output, target)
# measure accuracy and record loss
acc1, = accuracy(output, target, topk=(1,))
losses.update(loss.item(), images.size(0))
top1.update(acc1[0], images.size(0))
# measure elapsed time
batch_time.update(time.time() - end)
end = time.time()
# TODO: this should also be done with the ProgressMeter
print(' * Acc@1 {top1.avg:.3f} Acc@5 {top5.avg:.3f}'
.format(top1=top1, top5=top5))
return top1.avg
def save_checkpoint(state, is_best, filename='models/checkpoint.pth.tar'):
torch.save(state, filename)
if is_best:
shutil.copyfile(filename, 'models/model_best.pth.tar')
class AverageMeter(object):
"""Computes and stores the average and current value"""
def __init__(self, name, fmt=':f'):
self.name = name
self.fmt = fmt
self.reset()
def reset(self):
self.val = 0
self.avg = 0
self.sum = 0
self.count = 0
def update(self, val, n=1):
self.val = val
self.sum += val * n
self.count += n
self.avg = self.sum / self.count
def __str__(self):
fmtstr = '{name} {val' + self.fmt + '} ({avg' + self.fmt + '})'
return fmtstr.format(**self.__dict__)
class ProgressMeter(object):
def __init__(self, num_batches, meters, prefix=""):
self.batch_fmtstr = self._get_batch_fmtstr(num_batches)
self.meters = meters
self.prefix = prefix
def display(self, batch):
entries = [self.prefix + self.batch_fmtstr.format(batch)]
entries += [str(meter) for meter in self.meters]
print('\t'.join(entries))
def _get_batch_fmtstr(self, num_batches):
num_digits = len(str(num_batches // 1))
fmt = '{:' + str(num_digits) + 'd}'
return '[' + fmt + '/' + fmt.format(num_batches) + ']'
def adjust_learning_rate(optimizer, epoch, base_lr):
"""Sets the learning rate to the initial LR decayed by 10 every 30 epochs"""
lr = base_lr * (0.1 ** (epoch // 30))
for param_group in optimizer.param_groups:
param_group['lr'] = lr
def accuracy(output, target, topk=(1,)):
"""Computes the accuracy over the k top predictions for the specified values of k"""
with torch.no_grad():
maxk = max(topk)
batch_size = target.size(0)
_, pred = output.topk(maxk, 1, True, True)
pred = pred.t()
correct = pred.eq(target.view(1, -1).expand_as(pred))
res = []
for k in topk:
correct_k = correct[:k].reshape(-1).float().sum(0, keepdim=True)
res.append(correct_k.mul_(100.0 / batch_size))
return res
# + id="5Bu1tNT-p3UZ"
class CustomDataSet(Dataset):
def __init__(self, filenames, transform=None, labels=None):
self.filenames = filenames
self.labels = labels
self.transform = transform
def __len__(self):
return len(self.filenames)
def __getitem__(self, idx):
image = Image.open(self.filenames[idx]).convert("RGB")
label = self.labels[idx] if self.labels is not None else 0
if self.transform:
tensor_image = self.transform(image)
return tensor_image, label
normalize = transforms.Normalize(
mean=[0.485, 0.456, 0.406],
std=[0.229, 0.224, 0.225]
)
transform = transforms.Compose([
transforms.Resize(256),
transforms.CenterCrop(224),
transforms.ToTensor(),
normalize
])
classname_to_index = {"positive": 1, "negative": 0}
nb_classes = len(classname_to_index)
index_to_classname = {idx:name for name, idx in classname_to_index.items()}
def build_dataset(df, split):
filenames = [os.path.join(split, name) for name in df.image]
labels = [classname_to_index[name] for name in df.label]
dataset = CustomDataSet(filenames, transform=transform, labels=labels)
return dataset
df_train_full = pd.read_csv(data_dir+'train.csv')
df_test = pd.read_csv(data_dir+'submission_valid.csv')
nb =int(len(df_train_full)*0.9)
df_train = df_train_full.iloc[0:nb]
df_valid = df_train_full.iloc[nb:]
train_dataset = build_dataset(df_train, data_dir+"train")
valid_dataset = build_dataset(df_valid, data_dir+"train")
test_dataset = build_dataset(df_test, data_dir+"valid")
# + id="dPjDI8tjsfgT"
batch_size = 16
train_loader = torch.utils.data.DataLoader(
train_dataset,
batch_size=batch_size,
shuffle=True
)
val_loader = torch.utils.data.DataLoader(
valid_dataset,
batch_size=batch_size,
shuffle=False
)
test_loader = torch.utils.data.DataLoader(
test_dataset,
batch_size=batch_size,
shuffle=False
)
# + id="_taT2bywsu6k"
import torchvision
import torch.nn as nn
model = torchvision.models.resnet50(pretrained=True).to(device)
model.fc = nn.Linear(2048, nb_classes).to(device)
criterion = nn.CrossEntropyLoss()
lr = 0.0001
optimizer = torch.optim.SGD(model.parameters(), lr=lr, momentum=0.9)
# -
model
# + colab={"base_uri": "https://localhost:8080/", "height": 400} id="hWYh23WvsmSn" outputId="8768e243-47fc-4976-b2cb-d159309fe3a0"
# %%time
best_acc1 = 0.0
for epoch in range(20):
adjust_learning_rate(optimizer, epoch, base_lr=lr)
train(train_loader, model, criterion, optimizer, epoch)
# evaluate on validation set
acc1 = validate(val_loader, model, criterion)
# remember best acc@1 and save checkpoint
is_best = acc1 > best_acc1
best_acc1 = max(acc1, best_acc1)
save_checkpoint({
'epoch': epoch + 1,
'state_dict': model.state_dict(),
'best_acc1': best_acc1,
'optimizer' : optimizer.state_dict(),
}, is_best)
# + [markdown] id="oN_lwHcSM42f"
# # Submission
# + colab={"base_uri": "https://localhost:8080/", "height": 191} id="chzZo_AbvjEC" outputId="964bec8f-28e0-4ec6-f39f-6c6c87cdd130"
# %%time
ypred_test = []
for X, _ in test_loader:
with torch.no_grad():
X = X.to(device)
y = model(X)
y = y.cpu()
_, pred = y.max(dim=1)
pred = pred.tolist()
ypred_test.extend([index_to_classname[p] for p in pred])
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="-hWJzsYTM42j" outputId="9a041915-75c1-406f-fdd7-0e59be803a2d"
print(ypred_test[0:10])
df = pd.read_csv('submission_valid.csv')
df["label"] = ypred_test
df.head()
# + id="RGJkD57eM42v"
df.to_csv("submission.csv", index=False)
# + [markdown] id="zKcxwPiyM42x"
# Now, you can open the submision.csv file (File -> Open) file and download it!
#
# After you download it, you can upload it to the challenge frontend.
| 04_Julich_Challenges/01_Covid19_Challlenge/reference_soln.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Assignment 13 Solutions
# #### 1. Create a function that takes a list and string. The function should remove the letters in the string from the list, and return the list.
# **Examples:**
# `remove_letters(["s", "t", "r", "i", "n", "g", "w"], "string") ➞ ["w"]`
# `remove_letters(["b", "b", "l", "l", "g", "n", "o", "a", "w"], "balloon") ➞ ["b", "g", "w"]`
# `remove_letters(["d", "b", "t", "e", "a", "i"], "edabit") ➞ []`
# +
def remove_letters(in_list,in_string):
in_list_copy = in_list.copy()
for ele in in_string:
if ele in in_list:
in_list.remove(ele)
print(f'remove_letters{in_list_copy,in_string} ➞ {in_list}')
remove_letters(["s", "t", "r", "i", "n", "g", "w"], "string")
remove_letters(["b", "b", "l", "l", "g", "n", "o", "a", "w"], "balloon")
remove_letters(["d", "b", "t", "e", "a", "i"], "edabit")
# -
# #### 2. A block sequence in three dimensions. We can write a formula for this one:
#
# 
#
# Create a function that takes a number (step) as an argument and returns the amount of blocks in that step.
#
# **Examples:**
# `blocks(1) ➞ 5`
# `blocks(5) ➞ 39`
# `blocks(2) ➞ 12`
# +
def blocks(in_num):
depth = in_num*3+((in_num)-1)*1
height = [x for x in range(2,in_num+2)]
print(f'blocks({in_num}) ➞ {depth+sum(height)}')
blocks(1)
blocks(2)
blocks(3)
blocks(4)
blocks(5)
# -
# #### 3. Create a function that subtracts one positive integer from another, without using any arithmetic operators such as -, %, /, +, etc.
# **Examples:**
# `my_sub(5, 9) ➞ 4`
# `my_sub(10, 30) ➞ 20`
# `my_sub(0, 0) ➞ 0`
# +
from operator import sub
def my_sub(in_one,in_two):
output = sub(in_one,in_two) if in_one >= in_two else sub(in_two,in_one)
print(f'my_sub{in_one,in_two} ➞ {output}')
my_sub(5, 9)
my_sub(10, 30)
my_sub(0, 0)
# -
# #### 4. Create a function that takes a string containing money in dollars and pounds sterling (seperated by comma) and returns the sum of dollar bills only, as an integer.
# For the input string:
# 1. `Each amount is prefixed by the currency symbol: $ for dollars and £ for pounds.`
# 2. `Thousands are represented by the suffix k. i.e. $4k = $4,000 and £40k = £40,000`
#
# **Examples:**
# `add_bill("d20,p40,p60,d50") ➞ 20 + 50 = 70`
# `add_bill("p30,d20,p60,d150,p360") ➞ 20 + 150 = 170`
# `add_bill("p30,d2k,p60,d200,p360") ➞ 2 * 1000 + 200 = 2200`
# +
def add_bill(in_string):
out_num = 0
for ele in in_string.split(","):
if 'd' in ele:
if 'k' in ele:
out_num += int(ele.replace('d','').replace('k',''))*1000
else:
out_num += int(ele.replace("d",''))
print(f'add_bill({in_string}) ➞ {out_num}')
add_bill("d20,p40,p60,d50")
add_bill("p30,d20,p60,d150,p360")
add_bill("p30,d2k,p60,d200,p360")
# -
# #### 5. Create a function that flips a horizontal list into a vertical list, and a vertical list into a horizontal list.
# In other words, take an 1 x n list (1 row + n columns) and flip it into a n x 1 list (n rows and 1 column), and vice versa.
#
# **Examples:**
# `flip_list([1, 2, 3, 4]) ➞ [[1], [2], [3], [4]] # Take a horizontal list and flip it vertical.`
# `flip_list([[5], [6], [9]]) ➞ [5, 6, 9] # Take a vertical list and flip it horizontal.`
# `flip_list([]) ➞ []`
# +
def flip_list(in_list):
if len(in_list) > 0:
output = [ele[0] for ele in in_list] if isinstance(in_list[0],list) else [[ele] for ele in in_list]
else:
output = []
print(f'flip_list({in_list}) ➞ {output}')
flip_list([1,2,3,4])
flip_list([[5],[6],[9]])
flip_list([])
| Python Advance Programming/Advance Programming Assignment 13.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/brenoslivio/SME0828_DataScience/blob/master/Projects/2%20-%20An%C3%A1lise%20explorat%C3%B3ria%20de%20dados/Projeto2_Analise_Exploratoria_dos_dados_M.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="LJ7wNY4Uefh6"
# # SME0828 - Introdução à Ciência de Dados
# # Projeto 2: Análise exploratória dos dados
# + [markdown] colab_type="text" id="fscDWzEee1hv"
# ### Alunos
# + [markdown] colab_type="text" id="kIEYP_R3H6YU"
# <NAME>, Nº USP: 9437275
#
# <NAME>, Nº USP: 10276675
#
# <NAME>, Nº USP: 10276661
# + [markdown] colab_type="text" id="lB29Wn4fH0RJ"
# ***Universidade de São Paulo, São Carlos, Brasil.***
# + [markdown] colab_type="text" id="0Wm1OZv73iBq"
# ## Exercícios sugeridos para a atividade avaliativa 2
# + [markdown] colab_type="text" id="S8HH4554-1Rf"
# (PROVISÓRIO)
# Os seguintes exercícios foram sugeridos para entrega em 21 de setembro de 2020 para a disciplina SME0828 oferecida pelo Instituto de Ciências Matemáticas e de Computação (ICMC) da Universidade de São Paulo (USP), a qual foi ministrada pelo professor <NAME>. Tal atividade visa aplicar diferentes técnicas de preparação de dados em Python com o intuito de deixá-los mais suscetíveis a análises estatísticas e computacionais. O conjunto de dados que será mais utilizado é o famoso conjunto [Iris](https://pt.wikipedia.org/wiki/Conjunto_de_dados_flor_Iris#Conjunto_de_dados).
# + [markdown] colab_type="text" id="CHl-yeiA89Ho"
# ## Desenvolvimento
# + [markdown] colab_type="text" id="PzNzEPrHfsoM"
# ### 3 - Obtenha o boxplot de todas as variáveis da flor Iris, para cada espécie.
# -
# Importando-se as bibliotecas necessárias:
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
d_iris = pd.read_csv("/home/gandhi/Downloads/2sem 2020/Introdução à Ciência de Dados/Análise exploratória/iris.csv",
header = (0))
# Utilizando-se da biblioteca seaborn, pode-se criar boxplots para conjuntos de dados de interesse. Assim, abaixo temos a rotina para a obtenção de tais gráficos para cada uma das variáveis do conjunto de dados Iris, separados pela espécie:
#boxplots para comprimento da pétala
plt.figure(figsize=(12, 7))
sns.set_style("darkgrid")
sns.boxplot(x="species", y="petal_length", data=d_iris, palette = "Set2")
plt.xlabel('Espécie', fontsize=18)
plt.ylabel('Comprimento da pétala', fontsize=16)
plt.xticks(fontsize=14)
plt.yticks(fontsize=14)
plt.show(True)
#boxplots para largura da pétala
plt.figure(figsize=(12, 7))
sns.set_style("darkgrid")
sns.boxplot(x="species", y="petal_width", data=d_iris, palette = "Set2")
plt.xlabel('Espécie', fontsize=18)
plt.ylabel('Largura da pétala', fontsize=16)
plt.xticks(fontsize=14)
plt.yticks(fontsize=14)
plt.show(True)
#boxplots para comprimento da sépala
plt.figure(figsize=(12, 7))
sns.set_style("darkgrid")
sns.boxplot(x="species", y="sepal_length", data=d_iris, palette = "Set2")
plt.xlabel('Espécie', fontsize=18)
plt.ylabel('Comprimento da sépala', fontsize=16)
plt.xticks(fontsize=14)
plt.yticks(fontsize=14)
plt.show(True)
#boxplots para largura da sépala
plt.figure(figsize=(12, 7))
sns.set_style("darkgrid")
sns.boxplot(x="species", y="sepal_width", data=d_iris, palette = "Set2")
plt.xlabel('Espécie', fontsize=18)
plt.ylabel('Largura da sépala', fontsize=16)
plt.xticks(fontsize=14)
plt.yticks(fontsize=14)
plt.show(True)
# Outra alternativa ao gráfico boxplot é o gráfico de violino. Tal visualização permite a criação de um gráfico intuitivo no qual pode-se ver onde os dados se acumulam e onde eles são menos frequentes, mostrando a distribuição dos dados ao longo da reta. A visualização do boxplot junto à de violino, permite uma boa análise visual do comportamento dos dados, onde sua mediana se encontra e onde os outliers estão localizados. Segue abaixo gráficos de violino para as mesmas variáveis estudadas anteriormente:
# +
import warnings
warnings.filterwarnings("ignore")
#violinplot para comprimento da pétala
plt.figure(figsize=(8, 8))
sns.set_style("darkgrid")
sns.violinplot(y="species", x="petal_length", data=d_iris, palette = "Set3")
plt.xlabel('Espécie', fontsize=18)
plt.ylabel('Comprimento da pétala', fontsize=16)
plt.xticks(fontsize=14)
plt.yticks(fontsize=14)
plt.show(True)
# -
#violinplots para largura da pétala
plt.figure(figsize=(8, 8))
sns.set_style("darkgrid")
sns.violinplot(y="species", x="petal_width", data=d_iris, palette = "Set2")
plt.xlabel('Espécie', fontsize=18)
plt.ylabel('Largura da pétala', fontsize=16)
plt.xticks(fontsize=14)
plt.yticks(fontsize=14)
plt.show(True)
#violinplots para comprimento da sépala
plt.figure(figsize=(8, 8))
sns.set_style("darkgrid")
sns.violinplot(y="species", x="sepal_length", data=d_iris, palette = "Set2")
plt.xlabel('Espécie', fontsize=18)
plt.ylabel('Comprimento da sépala', fontsize=16)
plt.xticks(fontsize=14)
plt.yticks(fontsize=14)
plt.show(True)
#violinplots para largura da sépala
plt.figure(figsize=(8, 8))
sns.set_style("darkgrid")
sns.violinplot(y="species", x="sepal_width", data=d_iris, palette = "Set2")
plt.xlabel('Espécie', fontsize=18)
plt.ylabel('Largura da sépala', fontsize=16)
plt.xticks(fontsize=14)
plt.yticks(fontsize=14)
plt.show(True)
# Observação: perceba como o gráfico de violino é interessante para a visualização da mediana no caso da largura da pétala. No boxplot não conseguiamos ver a localização da mediana, a qual ficou mais evidente no segundo tipo de visualização. Em contrapartida, o boxplot é mais interessante na localização dos outliers dos conjuntos de dados.
# + [markdown] colab_type="text" id="GUAyONHbftDM"
# ### 6- Considere os dados abaixo, chamado quarteto de Anscombe. Calcule a média, variância, correlação de Pearson e Spearman entre as variáveis x e y. O que você pode dizer sobre esses dados?
# -
# Abaixo, tem-se os vatores de ddos utilizados para a construção do conjunto de dados quarteto de Anscombe:
# + colab={} colab_type="code" id="0BBK-p-rf4m2"
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from scipy.stats import pearsonr, spearmanr
x = [10, 8, 13, 9, 11, 14, 6, 4, 12, 7, 5]
y1 = [8.04, 6.95, 7.58, 8.81, 8.33, 9.96, 7.24, 4.26, 10.84, 4.82, 5.68]
y2 = [9.14, 8.14, 8.74, 8.77, 9.26, 8.10, 6.13, 3.10, 9.13, 7.26, 4.74]
y3 = [7.46, 6.77, 12.74, 7.11, 7.81, 8.84, 6.08, 5.39, 8.15, 6.42, 5.73]
x4 = [8, 8, 8, 8, 8, 8, 8, 19, 8, 8, 8]
y4 = [6.58, 5.76, 7.71, 8.84, 8.47, 7.04, 5.25, 12.50, 5.56, 7.91, 6.89]
# -
# Os quatro conjuntos são formados pelos vetores (x,y1); (x,y2); (x,y3); (x4,y4), e são sobre tais conjuntos que a análise será feita. Primeiramente, calculemos as médias e as variâncias dos vetores presentes nos 4 conjuntos de dados do quarteto de Anscombe:
print("Conjunto x:","média:",np.mean(x),";variância:",np.var(x))
print("Conjunto x4:","média:",np.mean(x4),";variância:",np.var(x4))
print("Conjunto y1:","média:",np.mean(y1),";variância:",np.var(y1))
print("Conjunto y2:","média:",np.mean(y2),";variância:",np.var(y2))
print("Conjunto y3:","média:",np.mean(y3),";variância:",np.var(y3))
print("Conjunto y4:","média:",np.mean(y4),";variância:",np.var(y4))
# Primeiramente, é notório que os conjuntos de dados x e x4 têm médias iguais e variâncias iguais. O mesmo fenômeno ocorre com os conjuntos y1, y2, y3 e y4, que possuem médias com valores muito próximos, assim como suas variâncias. Tal comportamento chama a atenção devido à diferença entre os dados nesses conjuntos.
# Para melhor vizualizar as relação entre as variâveis no quarteto de Anscombe, utilizemos da biblioteca seaborn para fazer gráficos:
sns.scatterplot(x,y1)
plt.xlabel('x', fontsize=18)
plt.ylabel('y1', fontsize=18)
plt.show(True)
sns.scatterplot(x,y2)
plt.xlabel('x', fontsize=18)
plt.ylabel('y2', fontsize=18)
plt.show(True)
sns.scatterplot(x,y3)
plt.xlabel('x', fontsize=18)
plt.ylabel('y3', fontsize=18)
plt.show(True)
sns.scatterplot(x4,y4)
plt.xlabel('x4', fontsize=18)
plt.ylabel('y4', fontsize=18)
plt.show(True)
# É notório que as dsitribuições de pontos nos gráficos de disperção acima possuem comportamentos exremanete diferentes. Cada conjunto de dados parece estabelecer uma relação diferente entre as variáveis que os compoem. Agora, analisemos os coeficientes de spearman e de pearson para cada conjunto:
#Correlação de Pearson:
print("Correlação de Pearson para os conjuntos de dados: \n",
"(x,y1):",pearsonr(x,y1)[0],"\n",
"(x,y2):",pearsonr(x,y2)[0],"\n",
"(x,y3):",pearsonr(x,y3)[0],"\n",
"(x4,y4):",pearsonr(x4,y4)[0],)
#Correlação de Spearman:
print("Correlação de Pearson para os conjuntos de dados: \n",
"(x,y1):",spearmanr(x,y1)[0],"\n",
"(x,y2):",spearmanr(x,y2)[0],"\n",
"(x,y3):",spearmanr(x,y3)[0],"\n",
"(x4,y4):",spearmanr(x4,y4)[0],)
xr = [sorted(x).index(i)+1 for i in x]
yr = [sorted(y2).index(i)+1 for i in y2]
spearmanr(xr,yr)[0]
# Obs.: o teste acima nos permite concluir que a função do coeficiente de spearman ordena os vetores e faz o processo inteiro para a obtenção do coeficiente. Mais sobre a função pode ser encontrado neste [link](https://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.spearmanr.html), no qual podemos ver que ela realmente pode receber vetores não ordenados.
# É notório que o coeficiente de pearson resultou em valores extremanete parecidos para os quatro conjuntos de dados, mesmo estes tendo comportamentos gráficos completamente diferentes. Isso mostra como o coeficiente pode mostrar-se inadequado quando as relações entre os conjuntos de dados não são lineares, e que tal medida deve ser acompanhada de pelo menos uma análise gráfica antes que conclusões sejam tomadas. Por outro lado, o coeficiente de Spearman, o qual analisa se existe uma relação monótona entre as variáveis mostrou-se mais sensível ao contexto e corroborou para a análise. Pode-se notar por exemplo que o gráfico de (x,y2) mostra claramente uma relação quadrática, a qual não foi detectada pelo coeficiente de pearson (o que era esperado pela própria natureza e utilização matemática desta medida) e a qual refletiu no valor do coeficiente de spearman, o qual mostrou-se menor do que para o conjnunto (x,y1) por exemplo, o que era esperado considerando-se os gráficos. Conclui-se assim que por mais que o coeficiente de Pearson seja amplamente utilizado em análises estatísticas e seja muito útil, deve-se tomar cuidado para não fazer uma análise rasa apenas com o seu valor, já que conjuntos de dados com comportamentos muito diferentes podem apresentar coeficientes parecidos e muitas vezes enganosos sobre as verdadeiras propriedades das relações entre as variáveis de estudo. O coeficiente de spearman mostrou-se de grande ajuda nesse caso para mostrar como os dados possuem comportamento diferentes, isso acompanhado de uma simples análise gráfica pode impedir que erros sejam cometidos na análise exploratória.
| Projects/2 - Análise exploratória de dados/Projeto2_Analise_Exploratoria_dos_dados_M.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # OpenCV による画像の読込・書込
import cv2
from matplotlib import pyplot as plt
# ## 画像の読込
img = cv2.imread('../lena.jpg')
# ## matplotlib で表示するため、BGRからRGBに変換する
img = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
# ## 画像の表示
plt.imshow(img)
plt.show()
# ## 画像の反転
img = cv2.flip(img, 1)
plt.imshow(img)
plt.show()
# ## 保存前にRGBからGBRに戻す
img = cv2.cvtColor(img, cv2.COLOR_RGB2BGR)
# ## 画像の保存
cv2.imwrite('../lena_flip.jpg', img)
| samples/opencv/load_and_save.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 알테어 시각화 결과를 장고로 배포하기
#
# - [신교수 파이썬애니웨어 사이트](http://logistex2021.pythonanywhere.com)
# - {'id'; '손님', 'pw'; '0000'}으로 로그인
# - `차트` 메뉴 중 처음 두 하위 메뉴가 알테어로 작성한 차트
# - `알테어 산점도`
# - `알테어 상호작용`
# + [markdown] slideshow={"slide_type": "skip"}
# - [신교수 파이썬애니웨어 사이트를 위한 깃허브 저장소](https://github.com/logistex/pa21)
# -
# - 알테어 관련 패키지 설치
# ```shell
# $ conda install -c conda-forge altair vega_datasets vega
# ```
# - 알테어 차트를 장고로 배포하는 방법
# - 뷰에서 알테어 차트를 json 형식으로 저장하여 템플릿으로 전달
# - 템플릿에서는 전달받은 json 형식 차트 명세를 `spec` 변수에 지정
# - `알테어 산점도` 작성을 위한 코드
# - 뷰 코드
# ```python
# # views.py
# def alt_django(request):
# import altair as alt
# from vega_datasets import data
#
# cars = data.cars()
# chart_json = alt.Chart(cars).mark_circle().encode(
# alt.X('Miles_per_Gallon'),
# alt.Y('Horsepower'),
# alt.Color('Origin'),
# ).to_json() # 차트를 json 형식으로 저장
# return render(request, 'chart/alt_chart.html', {'chart_json': chart_json}) # 저장한 json 형식 차트를 템플릿으로 전달
# ```
#
# - 템플릿 코드
# ```html
# <!DOCTYPE html>
# <html>
# <head>
# <script src="https://cdn.jsdelivr.net/npm/vega@5"></script>
# <script src="https://cdn.jsdelivr.net/npm/vega-lite@3"></script>
# <script src="https://cdn.jsdelivr.net/npm/vega-embed@4"></script>
# </head>
# <body>
# <div id="vis"></div>
# <script type="text/javascript">
# var spec = {{ chart_json|safe }}; /* json 형식 차트를 지정 */
# var opt = {"renderer": "canvas", "actions": false};
# vegaEmbed("#vis", spec, opt);
# </script>
# </body>
# </html>
# ```
# - `알테어 상호작용` 작성을 위한 코드
# - 뷰 코드
# ```python
# # views.py
# def alt_interactive(request):
# import altair as alt
# from vega_datasets import data
#
# domain = ['Europe', 'Japan', 'USA', ]
# range_ = ['red', 'green', 'blue', ]
#
# cars = data.cars()
# # x-축 인코딩에 대한 선택 구간 생성하여 브러쉬로 정의
# brush = alt.selection_interval(encodings=['x'], )
#
# # 브러쉬에 해당하면 진하게, 브러시에서 벗어나면 연하게
# opacity = alt.condition(brush, alt.value(0.9), alt.value(0.1), )
#
# # 연도별 자동차 도수를 개괄하는 도수분포도
# # 연도별 자동차 도수를 선택하는 상호작용적 구간 브러쉬 추가
# overview = alt.Chart(cars).mark_bar().encode(
# alt.X('Year:O', timeUnit='year', # 연도를 추출하고 서수형으로 지정
# axis=alt.Axis(title=None, labelAngle=0), # 축 제목 생략, 축 눈금 레이블 각도 생략
# ),
# alt.Y('count()', title=None), # 도수, 축 제목 생략
# opacity=opacity,
# ).add_selection(
# brush, # 차트에 대한 구간 브러쉬 선택 추가
# ).properties(
# width=800, # 차트 폭 800 픽셀로 설정
# height=150, # 차트 높이 150 픽셀로 설정
# title = {
# 'text': ['', '알테어 상호작용성', ''],
# 'subtitle': ['자동차 히스토그램'],
# },
# )
#
# # 개괄 도수분포도에 대응하는 상세 마력-연비 산점도
# # 브러쉬 선택에 대응하는 산점도 내부 점의 투명도 조절
# detail = alt.Chart(cars).mark_circle().encode(
# alt.X('Miles_per_Gallon', axis=alt.Axis(title='연비 [단위: 갤론 당 마일]'), ),
# alt.Y('Horsepower', axis=alt.Axis(title='마력'), ),
# alt.Color('Origin',
# legend=alt.Legend(
# title='원산지',
# orient='none',
# legendX=820,
# legendY=230,
# ),
# scale=alt.Scale(domain=domain, range=range_, ),
# ),
# opacity=opacity, # 브러쉬 선택에 대응하여 투명도 조절
# ).properties(
# width=800, # 차트 폭을 상단 차트와 동일하게 설정
# height=500,
# title={
# 'text': [''],
# 'subtitle': ['연비-마력 산점도'],
# },
# )
#
# # '&' 연산자로 차트 수직 병합
# interlinked = overview & detail
# interlinked_json = interlinked.to_json()
# return render(request, 'chart/alt_interactive.html', {'interlinked_json': interlinked_json})
# ```
#
# - 템플릿 코드
# ```html
# <!DOCTYPE html>
# <html>
# <head>
# <script src="https://cdn.jsdelivr.net/npm/vega@5"></script>
# <script src="https://cdn.jsdelivr.net/npm/vega-lite@3"></script>
# <script src="https://cdn.jsdelivr.net/npm/vega-embed@4"></script>
# </head>
# <body>
# <div id="vis"></div>
# <script type="text/javascript">
# var spec = {{ interlinked_json|safe }}; /* json 형식 차트를 지정 */
# var opt = {"renderer": "canvas", "actions": false};
# vegaEmbed("#vis", spec, opt);
# </script>
# </body>
# </html>
# ```
| .ipynb_checkpoints/01_altair_django-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: sm-project
# language: python
# name: sm-project
# ---
# # Libraries
# +
## libraries
import sys
import os
import pandas as pd
import seaborn as sns
import string
import matplotlib.pyplot as plt
from wordcloud import WordCloud, STOPWORDS
import re
import numpy as np
from math import cos, asin, sqrt
import time
# %matplotlib inline
# to import Database class from data_collection folder
module_path = os.path.abspath(os.path.join('..')+'/data_collection')
if module_path not in sys.path:
sys.path.append(module_path)
# now that the folder is in the path, ../data_collection/database.py can be imported
from storage_managers.database import Database
# -
# # Data Collection Summary:
# +
db = Database()
# get yelp businesses
yelp_sql = '''SELECT * FROM businesses WHERE url LIKE '%yelp%' '''
yelp_df = db.select_df(yelp_sql)
print('\033[1m- Yelp API search yieleded {} halal-related restaurants in NYC'.format(yelp_df.shape[0]))
# get halal-reviews (reviews that include the word 'halal')
reviews_sql = '''SELECT * FROM reviews'''
reviews_df = db.select_df(reviews_sql)
print('- {} reviews containinf the word Halal were found in those restaurants'.format(reviews_df.shape[0]))
# subset of yelp businesses with halal-reviews
halal_yelp_df = yelp_df[ yelp_df.platform_id.isin(reviews_df.restaurant_id.unique())]
print('- In particular, {} of those restaurants had reveiws that contained the word Halal\033[0m'.format(halal_yelp_df.shape[0]))
# add column with num of scraped reviews per business
counts = reviews_df.restaurant_id.value_counts()
yelp_df = yelp_df.assign(halal_review_count=yelp_df.platform_id.map(dict(zip(counts.index, counts.values))))
yelp_df.halal_review_count = yelp_df.halal_review_count.fillna(0)
# plot histogram of counts per restaurant
plt.figure(figsize=(10,10))
g = sns.distplot(yelp_df.halal_review_count, kde = False)
g.set_title('Distribution of halal-reviews per Yelp business', size=14)
g.set_xlabel('Halal review count', size=14)
g.set_ylabel('Count of incidences', size=14)
plt.plot([3, 3],[0, 3500], linewidth=2, linestyle='dashed', color='r') # cutoff between 2 & 5. To remove noise
plt.show()
# +
# plot histogram of Yelp businesses with more than 10 halal-reviews
fig, ax = plt.subplots(figsize=(14,10))
g = sns.distplot(yelp_df.halal_review_count[yelp_df.halal_review_count>3], kde = False, ax=ax)
g.set_title('Zoom in on Yelp business with more than 3 reviews per business', size=14)
g.set_xlabel('Halal review count', size=14)
g.set_ylabel('Count of incidences', size=14)
# Add labels to the plot
style = dict(size=10)
ax.text(92, 3, 'Burgers By\n Honest Chops', **style)
ax.text(57, 5, 'The Halal Guys\n +2', **style)
ax.text(46, 4, "Sam's Steak\n & Grill+1", **style)
plt.show()
# -
# distribution of reviews' dates
reviews_df.review_date[0]
# df["date"] = df["date"].astype("datetime64")
# df.groupby(df["date"].dt.month).count().plot(kind="bar")
ax, fig = plt.subplots(figsize=(18,8))
reviews_df.review_date = reviews_df['review_date'].astype('datetime64')
reviews_df.groupby([reviews_df.review_date.dt.year, reviews_df.review_date.dt.quarter])['id'].count().plot(kind='bar')
plt.title('Distribution of halal reviews dates', size=14)
plt.xlabel('Review date (quarter, year)', size=14)
plt.show()
# # Target Feature
# ## Halal restaurants from Zabiha.com & Zomato.com
# +
target_df = pd.read_csv('/Users/wesamazaizeh/Desktop/Projects/halal_o_meter/src/data/data_collection/target_feature/zabiha_list_DEPRECATED.csv',\
index_col=0)
print('\033[1m- {} Halal-confirmed restaurants were found on Zabiha.com'.format(sum(target_df['source'] == 'Zabiha'))) # 745 restaurants from Zabiha.com
print('- {} Halal-tagged restaurants were found on Zomato.com\033[0m'.format(sum(target_df['source'] == 'Zomato'))) # 340 restaurants from Zomato.com
target_df['validated_address'] = target_df['validated_address'].str.lower()
target_match = target_df['validated_address'][target_df.source == 'Zabiha'].isin(target_df['validated_address'][target_df.source == 'Zomato'])
target_match_count = [sum(target_match), len(target_match)-sum(target_match)]
plt.figure(figsize=(14,8))
g = sns.barplot(x=['Zabiha & Zomato','Zabiha Only'], y=target_match_count)
g.set_title('Overlap between Zabiha and Zomato (by validated address)', size=14)
g_labels = g.set_xticklabels(g.get_xticklabels(), rotation=45, horizontalalignment='right')
for p in g.patches:
g.annotate('{:.0f}'.format(p.get_height()), (p.get_x()+0.3, p.get_height()),
ha='center', va='bottom', color= 'black')
# -
# ## Is this because of the address validation? What if we comapre by name?
plt.figure(figsize=(14,8))
target_df['name'] = target_df['name'].str.lower()
target_match = target_df['name'][target_df.source == 'Zabiha'].isin(target_df['name'][target_df.source == 'Zomato'])
target_match_count = [sum(target_match), len(target_match)-sum(target_match)]
g = sns.barplot(x=['Zabiha & Zomato','Zabiha Only'], y=target_match_count)
g.set_title('Overlap between Zabiha and Zomato (by restaurant name)', size=14)
g_labels = g.set_xticklabels(g.get_xticklabels(), rotation=45, horizontalalignment='right')
for p in g.patches:
g.annotate('{:.0f}'.format(p.get_height()), (p.get_x()+0.3, p.get_height()),
ha='center', va='bottom', color= 'black')
# ## How many Yelp restaurants (838) with halal reviews match halal-confirmed restaurants (745) from Zabiha.com?
# +
# cleanup yelp address to match validated addresses
yelp_df.address = yelp_df.address.str.lower()
yelp_df.address = yelp_df.address.map(lambda address: re.sub(r'[^A-Za-z0-9, ]+', '', address).split(','))
yelp_df.address = yelp_df.address.map(lambda address: ', '.join([str.strip() for str in address])+', usa')
# only yelp businesses with halal reviews
halal_mask = yelp_df['platform_id'].isin(reviews_df['restaurant_id'].unique())
halal_df = yelp_df[halal_mask]
# only Zabiha results
zabiha_df = target_df[ target_df.source == 'Zabiha']
# overlap with Zabiha.com by address
address_match = halal_df['address'].isin(zabiha_df['validated_address'])
address_match_count = [sum(address_match), len(address_match)-sum(address_match)]
plt.figure(figsize=(14,8))
g = sns.barplot(x=['Match \nby address','No match'], y=address_match_count)
g.set_title('How many of Yelp restaurants with halal-reviews match\naddresses of Zabiha restaurants?')
g_labels = g.set_xticklabels(g.get_xticklabels(), rotation=45, horizontalalignment='right')
for p in g.patches:
g.annotate('{:.0f}'.format(p.get_height()), (p.get_x()+0.3, p.get_height()),
ha='center', va='bottom', color= 'black')
# -
# ## How about Yelp restaurants (838) with halal reviews that match halal-confirmed restaurants (340) from Zomato.com?
# +
# only Zabiha results
zomato_df = target_df[ target_df.source == 'Zomato']
# overlap with Zabiha.com by address
address_match = halal_df['address'].isin(zomato_df['validated_address'])
address_match_count = [sum(address_match), len(address_match)-sum(address_match)]
plt.figure(figsize=(14,8))
g = sns.barplot(x=['Match \nby address','No match'], y=address_match_count)
g.set_title('How many of Yelp restaurants with halal-reviews match\naddresses of Zomato restaurants?')
g_labels = g.set_xticklabels(g.get_xticklabels(), rotation=45, horizontalalignment='right')
for p in g.patches:
g.annotate('{:.0f}'.format(p.get_height()), (p.get_x()+0.3, p.get_height()),
ha='center', va='bottom', color= 'black')
# -
# ## Do the Zabiha list match any restaurants that don't have halal-reviews?
# +
# only yelp businesses withOUT halal reviews
not_halal_mask = yelp_df['platform_id'].isin(reviews_df['restaurant_id'].unique())
not_halal_df = yelp_df[~not_halal_mask]
# overlap with Zabiha.com by address
address_match = not_halal_df['address'].isin(zabiha_df['validated_address'])
address_match_count = [sum(address_match), len(address_match)-sum(address_match)]
plt.figure(figsize=(14,8))
g = sns.barplot(x=['Match \nby address','No match'], y=address_match_count)
g.set_title('How many of Yelp restaurants withOUT halal-reviews match\naddresses of Zabiha restaurants?')
g_labels = g.set_xticklabels(g.get_xticklabels(), rotation=45, horizontalalignment='right')
for p in g.patches:
g.annotate('{:.0f}'.format(p.get_height()), (p.get_x()+0.3, p.get_height()),
ha='center', va='bottom', color= 'black')
# -
# ### How many Zabiha restaurants are within NY state?
ny_or_not = zabiha_df['address'].apply(lambda x: 'NY' in x)
ny_or_not_count = [sum(ny_or_not), len(ny_or_not)-sum(ny_or_not)]
plt.figure(figsize=(14, 8))
g = sns.barplot(x=['NY','Other'], y=ny_or_not_count)
g_labels = g.set_xticklabels(g.get_xticklabels(), rotation=45, horizontalalignment='right')
# ## Are my restaurants of interest even in NYC?
ny_or_not = halal_df['address'].apply(lambda x: ' ny ' in x)
ny_or_not_count = [sum(ny_or_not), len(ny_or_not)-sum(ny_or_not)]
plt.figure(figsize=(14, 8))
g = sns.barplot(x=['NY','Other'], y=ny_or_not_count)
g_labels = g.set_xticklabels(g.get_xticklabels(), rotation=45, horizontalalignment='right')
# # New Approach:
# ## Match restaurants by coordinate distance & name similarity
# ### 1. Add coordinates and image_url data to restaurants of interest (halal_df)
# +
from importlib import reload
import Yelp_business_search
import json
c=0
# halal_df['image_url'] = None
for i, yelp_id in halal_df['platform_id'].iteritems():
if pd.isna(halal_df.loc[i, 'image_url']):
try:
row = Yelp_business_search.get_yelp_business_details(yelp_id)
halal_df.loc[i, 'image_url'] = row['image_url']
halal_df.loc[i, 'lat'] = row['lat']
halal_df.loc[i, 'lng'] = row['lng']
time.sleep(0.2)
except:
row = Yelp_business_search.get_yelp_business_details(yelp_id)
halal_df.loc[i, 'image_url'] = row['image_url']
halal_df.loc[i, 'lat'] = row['lat']
halal_df.loc[i, 'lng'] = row['lng']
time.sleep(0.2)
c+=1
print('[{0}/{1}] id:{2}'.format(c, halal_df.shape[0], yelp_id), end='\r', flush=True )
# -
# ## Find appropriate thresholds
# ### 2.1. Find closest restaurant to the restaurant of interest from Zabiha's list with coordinates
zabiha_df = pd.read_csv('/Users/wesamazaizeh/Desktop/Projects/halal_o_meter/src/data/data_collection/target_feature/zabiha_list.csv')
zabiha_df['lat'] = zabiha_df['coordinates'].apply(lambda x: float(x.split(',')[0]))
zabiha_df['lng'] = zabiha_df['coordinates'].apply(lambda x: float(x.split(',')[1]))
zabiha_df.head()
# +
def distance(lat1, lon1, lat2, lon2):
p = 0.017453292519943295
a = 0.5 - cos((lat2-lat1)*p)/2 + cos(lat1*p)*cos(lat2*p) * (1-cos((lon2-lon1)*p)) / 2
dist = 12742 * asin(sqrt(a)) # [km]
return dist
def closest(v, data):
return min(data, key=lambda p: distance(v['lat'],v['lng'],data[p]['lat'],data[p]['lng']))
halal_dict = {row['name']: {'lat' : row.lat, 'lng' : row.lng} for _,row in halal_df.iterrows()}
zabiha_dict = {row['name']: {'lat' : row.lat, 'lng' : row.lng} for _,row in zabiha_df.iterrows()}
c=0
for i, res_name in halal_df['name'].iteritems():
res = closest(halal_dict[res_name], zabiha_dict)
halal_df.loc[i, 'zabiha_name'] = res
halal_df.loc[i, 'zabiha_lat'] = zabiha_dict[res]['lat']
halal_df.loc[i, 'zabiha_lng'] = zabiha_dict[res]['lng']
halal_df.loc[i, 'zabiha_distance'] = distance(halal_df.loc[i, 'lat'], halal_df.loc[i, 'lng'],
zabiha_dict[res]['lat'], zabiha_dict[res]['lng'])
c+=1
print('Progress: {0}/{1}'.format(c, halal_df.shape[0]), end='\r', flush=True)
# -
# ### 2.2. Add name similarity score
# +
from difflib import SequenceMatcher
import matplotlib.pyplot as plt
from matplotlib.patches import Ellipse
import seaborn as sns
for i,_ in halal_df.iterrows():
s = SequenceMatcher(None, halal_df.loc[i, 'name'], halal_df.loc[i, 'zabiha_name']).ratio()
halal_df.loc[i, 'zabiha_similarity'] = s
a = sns.jointplot(y='zabiha_similarity', x='zabiha_distance', data=halal_df, height=10)
# highlight possible threshhold area
elps = Ellipse((0, 0.8), 350, 0.5,edgecolor='r',facecolor='none')
ax = a.ax_joint
ax.add_artist(elps)
plt.show()
# -
# plot distribution of distance for all zabiha with similarity higher than 0.5 very close to the restaurant of interest
sns.distplot(halal_df[halal_df['zabiha_similarity'] > 0.5]['zabiha_distance'], kde=False, bins=np.arange(0.0, 1.0, 0.01))
plt.xlim(0, 0.5)
plt.plot([.03, .03],[0, 70], linewidth=2, linestyle='dashed', color='r') # cutoff between 2 & 5. To remove noise
plt.show()
halal_df[(halal_df['zabiha_similarity'] > 0.5) & (halal_df['zabiha_distance'] < 0.3)][['name', 'zabiha_name']]
# Criteria:
# - distance less than 30m
# - name similarity score greater than 0.5
mfoodies_df = pd.read_csv('/Users/wesamazaizeh/Desktop/Projects/halal_o_meter/src/data/data_collection/target_feature/muslim_foodies_list.csv')
mfoodies_df['lat'] = zabiha_df['coordinates'].apply(lambda x: float(x.split(',')[0]))
mfoodies_df['lng'] = zabiha_df['coordinates'].apply(lambda x: float(x.split(',')[1]))
mfoodies_df.head()
# +
# mfoodies_dict = {row['name']: {'lat' : row.lat, 'lng' : row.lng} for _,row in mfoodies_df.iterrows()}
c=0
for i, res_name in halal_df['name'].iteritems():
res = closest(halal_dict[res_name], mfoodies_dict)
halal_df.loc[i, 'mfoodies_name'] = res
halal_df.loc[i, 'mfoodies_lat'] = mfoodies_dict[res]['lat']
halal_df.loc[i, 'mfoodies_lng'] = mfoodies_dict[res]['lng']
halal_df.loc[i, 'mfoodies_distance'] = distance(halal_df.loc[i, 'lat'], halal_df.loc[i, 'lng'],
mfoodies_dict[res]['lat'], mfoodies_dict[res]['lng'])
s = SequenceMatcher(None, halal_df.loc[i, 'name'], halal_df.loc[i, 'mfoodies_name']).ratio()
halal_df.loc[i, 'mfoodies_similarity'] = s
c+=1
print('Progress: {0}/{1}'.format(c, halal_df.shape[0]), end='\r', flush=True)
a = sns.jointplot(y='mfoodies_similarity', x='mfoodies_distance', data=halal_df, height=10)
plt.show()
# -
a = sns.jointplot(y='mfoodies_similarity', x='mfoodies_distance', data=halal_df, height=10, xlim=(0,100))
plt.show()
| src/data/EDA/Yelp_EDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Malware Downloader Triage Notes
# > Researching malware downloaders, detection and triage
#
# - toc: true
# - badges: true
# - categories: [downloader,research,detection_engineering,yara,triage]
# ## Overview
# We are going to take a look at two different downloaders, which are so simple they don't have great static detection or information extraction in UnpacMe yet! Our goal is to generate some Yara rules, or maybe some code to identify and extract the relevant info (download URL) in a generic static way.
#
# ### Samples
# - [1a10e2940151982f2ab4f1e62be6e4f53074a2ffb90c7977e16d6a183db98695](https://malshare.com/sample.php?action=detail&hash=1a10e2940151982f2ab4f1e62be6e4f53074a2ffb90c7977e16d6a183db98695)
# - [9211ebf25c3cd3641451c95c50c1d3b7b2a4c53c36fa36564f3c1a177a0cda3d](https://malshare.com/sample.php?action=detail&hash=9211ebf25c3cd3641451c95c50c1d3b7b2a4c53c36fa36564f3c1a177a0cda3d)
#
# ## Triage of `9211...`
#
# The samples has a plaintext URL and the download functionality is not obfuscated/packed and can be identifed using [CAPA](https://github.com/mandiant/capa) as seen in [UnpacMe](https://www.unpac.me/results/9b085f2f-9a1e-43ea-b7ae-814e85e90ddc/#/).
#
# The hardcoded URL is: `http://apuservis.pe/ocultar/fw%d.exe`.
#
# There are 2 loops used to generate ULRs from 1 - 7 calling the URLs:
# - `http://apuservis.pe/ocultar/fw1.exe`
# - `http://apuservis.pe/ocultar/fw2.exe`
# - `http://apuservis.pe/ocultar/fw3.exe`
# - `http://apuservis.pe/ocultar/fw4.exe`
# - `http://apuservis.pe/ocultar/fw5.exe`
# - `http://apuservis.pe/ocultar/fw6.exe`
#
# The file is writtent to `%APPDATA%` with a random file name using a random number `"%08x.exe`.
#
# To execute the file the API `ShellExecuteW` is resolved dynamically using a CRC32 hash then it is called with the `open` command.
#
# +
import requests
HASHDB_HUNT_URL = 'https://hashdb.openanalysis.net/hunt'
HASHDB_HASH_URL = 'https://hashdb.openanalysis.net/hash'
api_hash = 0x1FA8A1D4 + 5
hunt_request = {"hashes": [api_hash]}
r = requests.post(HASHDB_HUNT_URL, json=hunt_request)
print(r.json())
# -
r = requests.get(HASHDB_HASH_URL + '/crc32/' + str(api_hash))
print(r.json())
# ### Possible IOCs
#
# - There is self-delete functionatlity using the batch script:
# `/c ping 127.0.0.1 && del \"%s\" >> NUL`
#
# - They use `GetEnvironmentVariableW(L"ComSpec", Filename, 0x104u)` to get the `cmd.exe` path.
#
# - They have a hardcoded URL `http://apuservis.pe/ocultar/fw%d.exe`
#
# - They have a hardcoded HTTP header `GET %S HTTP/1.1`
#
# - They have a CRC32 hash algo used for the dynamic API resolving
#
# ### Yara rule
#
# ```yaml
# rule download_hunt {
#
# meta:
# description = "Hunt for simple downloaders"
#
# strings:
# $s1 = "/c ping 127.0.0.1 && del \"%s\" >> NUL" wide ascii nocase
# $s2 = "http://" wide ascii
# $s3 = "GET %S HTTP/1.1" wide ascii nocase
# $x1 = { 35 20 83 B8 ED }
# $x2 = { 81 F? 20 83 B8 ED }
#
# condition:
# all of ($s*) and 1 of ($x*)
#
# }
# ```
#
# ** let's also check file size
#
# ** these are small binaries with very few functions, one ID trick might be to try and identify how many functions and only trigger an bins with a few functions... for this we could maybe used CFG count... from @psifertex
#
# > You can see the function table from a CFG binary with dumpbin /loadconfig test.exe
#
#
# #### Yara Rule Revisions
#
# We ran a scan with the above rule over the [MalwareBazaar](https://riskmitigation.ch/yara-scan/index.html) corpus and the results only gave us [one match](https://riskmitigation.ch/yara-scan/results/536840a0d04a2bbc3b63dd1e8cf36c008b1a242d23428c101d7e75157508958b/) (the sample we were originally looking at). This is a **bad** rule.
#
# We are going to loosen the rule to see if we can catch more samples.
#
#
# ```yaml
# rule download_hunt_2 {
#
# meta:
# description = "Hunt for simple downloaders"
#
# strings:
# $s1 = "/c ping 127.0.0.1 && del" wide ascii xor
# $s2 = "http://" wide ascii xor
#
#
# condition:
# all of ($s*)
#
# }
# ```
#
#
#
#
#
# 
# ## Triage of `1a10e...`
#
#
# > twitter: https://twitter.com/malwrhunterteam/status/1535745376766115840
#
#
# There is a PDB path in the binary `C:\Users\H0e4a0r1t\Documents\Visual Studio 2015\Projects\worddy\x64\Release\worddy.pdb`. Based on the username `H0e4a0r1t` we found a possible GitHub https://github.com/h0e4a0r1t and this looks like maybe a "redteam" tools developer??
#
# ### Possible IOCs
#
# - There is an embeded blob that is encrypted with a singl-byte XOR `0x99`
#
# - Once decrypted the blob is a standard Cobalt Strike loader with the following "header" strings.
#
# ```
# Accept: text/html,application/xhtml+xml,application/xml;q=0.9,*/*;q=0.8
# Accept-Language: en-US,en;q=0.5
# Referer: http://code.jquery.com/
# Accept-Encoding: gzip, deflate
# User-Agent: Mozilla/5.0 (Windows NT 6.3; Trident/7.0; rv:11.0) like Gecko
# ```
#
# - The C2 IP is `172.16.17.32`
#
# - To load and launch the decrypted shellcode they use the following APIs
# - VirtualAlloc
# - WriteProcessMemory
# - QueueUserAPC
#
# ### Yara Rule
#
# ```yaml
# rule cs_downloader {
#
# meta:
# description = "Hunt for Cobalt Strike downloader"
#
# strings:
# $s1 = "Accept-Language:" xor(0x01-0xff)
# $s2 = "Referer: http://code.jquery.com/" xor(0x01-0xff)
# $s3 = "Accept-Encoding: gzip, deflate" xor(0x01-0xff)
# $s4 = "User-Agent:" xor(0x01-0xff)
#
#
# condition:
# all of ($s*) and uint32(@s1) == uint32(@s3)
#
# }
# ```
#
#
#
#
# ### Yara Results
#
# We only got 2 hits, this rule also sucks, lol!
#
# ```
# [
# {
# "rule": "cs_downloader",
# "malware": "CobaltStrike",
# "sha256": "e54514b1164508c049733c7dafc97f24ae66d42b8146b0e1a1271f9af7c94d48",
# "mime_type": "application/x-msdownload",
# "virustotal_link": "https://www.virustotal.com/gui/file/e54514b1164508c049733c7dafc97f24ae66d42b8146b0e1a1271f9af7c94d48/detection",
# "malwarebazaar_link": "https://bazaar.abuse.ch/sample/e54514b1164508c049733c7dafc97f24ae66d42b8146b0e1a1271f9af7c94d48/",
# "tags": []
# },
# {
# "rule": "cs_downloader",
# "malware": "CobaltStrike",
# "sha256": "6220127ada00d84b58d718152748cd2c62007b1de92201701dc2968d2b00e31f",
# "mime_type": "application/x-msdownload",
# "virustotal_link": "https://www.virustotal.com/gui/file/6220127ada00d84b58d718152748cd2c62007b1de92201701dc2968d2b00e31f/detection",
# "malwarebazaar_link": "https://bazaar.abuse.ch/sample/6220127ada00d84b58d718152748cd2c62007b1de92201701dc2968d2b00e31f/",
# "tags": []
# }
# ]
# ```
| _notebooks/2022-06-12-downloader-triage.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import seaborn as sns
import warnings as w
import datetime
# ## Cleaning
# ### Cleaning China African Trade Data 1992 - 2018
# +
Exp_China_Africa = 'China-africa trade Data 1992-2018.xlsx'
col_string = "A:BC"
Exp_zh_af_df = pd.read_excel(Exp_China_Africa, skiprows=1, skipfooter = 6, sheet_name = 0, usecols=col_string, parse_dates=['Year'])
Exp_zh_af_df.head(5)
# -
Exp_Zh_final = Exp_zh_af_df.melt(id_vars='Year', var_name='Country', value_name='Export')
print(Exp_Zh_final)
Exp_Zh_final.info()
print('\n')
Exp_Zh_final.head()
Exp_Zh_final.to_csv('Exp_Zh_final.csv')
# ### Cleaning Chinese Imports from Africa 2002 - 2018
# +
Imp_China_Africa = 'China-africa trade Data 1992-2018.xlsx'
col_string = "A:BC"
Imp_zh_af_df = pd.read_excel(Imp_China_Africa, skiprows=1, skipfooter = 6, sheet_name=1,usecols = col_string,
parse_dates=['Year'])
Imp_zh_af_df.head(58)
# -
Imp_zh_final = Imp_zh_af_df.melt(id_vars='Year', var_name='Country', value_name='Import')
Imp_zh_final
Imp_zh_final.info()
print('\n')
Imp_zh_final.head()
Imp_zh_final.to_csv('Imp_zh_final.csv')
# ### Cleaning US Exports from Africa 2002 - 2018
# +
Exp_China_Africa = 'China-africa trade Data 1992-2018.xlsx'
col_string = "A:BB"
Exp_US_af_df = pd.read_excel(Exp_China_Africa, skiprows=1, skipfooter = 6, sheet_name =2, usecols=col_string, parse_dates=['Year'])
Exp_US_af_df['Year'] = pd.DatetimeIndex(Exp_US_af_df['Year']).year
Exp_US_af_df.set_index('Year')
.head(56)
# -
Exp_US_af_df['Annual Total'] = Exp_US_af_df.sum(axis = 1)
Exp_US_Final = Exp_US_af_df.melt(id_vars='Year', var_name='Country', value_name='Export')
Exp_US_Final
Exp_US_Final.to_csv('Exp_US_Final.csv')
# ### Cleaning US Imports from Africa 2002 - 2018
# +
Exp_China_Africa = 'China-africa trade Data 1992-2018.xlsx'
col_string = "A:BB"
Imp_US_af_df = pd.read_excel(Exp_China_Africa, skiprows=1, skipfooter = 5, sheet_name = 3, usecols=col_string, parse_dates=['Year'])
Imp_US_af_df['Year'] = pd.DatetimeIndex(Imp_US_af_df['Year']).year
Imp_US_af_df.set_index('Year')
Imp_US_af_df.head(55)
# -
Imp_US_final = Imp_US_af_df.melt(id_vars='Year', var_name='Country', value_name='Import')
Imp_US_final
Imp_US_final.describe()
Imp_US_final.to_csv('Imp_US_final.csv')
| .ipynb_checkpoints/China Project Cleaning-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (tf2)
# language: python
# name: tf2
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Hyperparameter-Tuning-Demo-using-Keras-Tuner" data-toc-modified-id="Hyperparameter-Tuning-Demo-using-Keras-Tuner-1"><span class="toc-item-num">1 </span>Hyperparameter Tuning Demo using Keras Tuner</a></span><ul class="toc-item"><li><ul class="toc-item"><li><span><a href="#Load-the-data" data-toc-modified-id="Load-the-data-1.0.1"><span class="toc-item-num">1.0.1 </span>Load the data</a></span></li><li><span><a href="#Check-tensorflow-and-kerastuner-versions" data-toc-modified-id="Check-tensorflow-and-kerastuner-versions-1.0.2"><span class="toc-item-num">1.0.2 </span>Check tensorflow and kerastuner versions</a></span></li><li><span><a href="#Import-libraries" data-toc-modified-id="Import-libraries-1.0.3"><span class="toc-item-num">1.0.3 </span>Import libraries</a></span></li><li><span><a href="#Set-random-seeed" data-toc-modified-id="Set-random-seeed-1.0.4"><span class="toc-item-num">1.0.4 </span>Set random seeed</a></span></li><li><span><a href="#Model-without-Hyperparameter-Tuning" data-toc-modified-id="Model-without-Hyperparameter-Tuning-1.0.5"><span class="toc-item-num">1.0.5 </span>Model without Hyperparameter Tuning</a></span></li><li><span><a href="#Building-the-Hypermodel" data-toc-modified-id="Building-the-Hypermodel-1.0.6"><span class="toc-item-num">1.0.6 </span>Building the Hypermodel</a></span></li><li><span><a href="#Initialize-hypermodel" data-toc-modified-id="Initialize-hypermodel-1.0.7"><span class="toc-item-num">1.0.7 </span>Initialize hypermodel</a></span></li><li><span><a href="#Build-Random-Search-Tuner" data-toc-modified-id="Build-Random-Search-Tuner-1.0.8"><span class="toc-item-num">1.0.8 </span>Build Random Search Tuner</a></span></li><li><span><a href="#Run-Random-Search" data-toc-modified-id="Run-Random-Search-1.0.9"><span class="toc-item-num">1.0.9 </span>Run Random Search</a></span></li><li><span><a href="#Evaluate-Random-Search" data-toc-modified-id="Evaluate-Random-Search-1.0.10"><span class="toc-item-num">1.0.10 </span>Evaluate Random Search</a></span></li><li><span><a href="#Build,-Run-and-Evaluate-Hyperband-Tuner" data-toc-modified-id="Build,-Run-and-Evaluate-Hyperband-Tuner-1.0.11"><span class="toc-item-num">1.0.11 </span>Build, Run and Evaluate Hyperband Tuner</a></span></li><li><span><a href="#Build,-Run-and-Evaluate-Bayesian-Optimization-Tuner" data-toc-modified-id="Build,-Run-and-Evaluate-Bayesian-Optimization-Tuner-1.0.12"><span class="toc-item-num">1.0.12 </span>Build, Run and Evaluate Bayesian Optimization Tuner</a></span></li></ul></li></ul></li></ul></div>
# -
# # Hyperparameter Tuning Demo using Keras Tuner
# ### Load the data
# +
from tensorflow.keras.datasets import boston_housing
(x_train, y_train), (x_test, y_test) = boston_housing.load_data()
# -
# ### Check tensorflow and kerastuner versions
# +
import tensorflow as tf
import keras_tuner as kt
from tensorflow.keras import backend as K
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Activation
from tensorflow.keras.layers import Dense, Dropout
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.metrics import categorical_crossentropy
print(tf.__version__)
# -
# ### Import libraries
from sklearn.preprocessing import StandardScaler
from tensorflow.keras import models, layers
from keras_tuner import HyperModel, RandomSearch, Hyperband, BayesianOptimization
# ### Set random seeed
from numpy.random import seed
seed(42)
import tensorflow
tensorflow.random.set_seed(42)
# ### Model without Hyperparameter Tuning
# +
scaler = StandardScaler()
scaler.fit(x_train)
x_train_scaled = scaler.transform(x_train)
x_test_scaled = scaler.transform(x_test)
model = models.Sequential()
model.add(layers.Dense(8, activation='relu', input_shape=(x_train.shape[1],),
kernel_initializer='zeros', bias_initializer='zeros'))
model.add(layers.Dense(16, activation='relu', kernel_initializer='zeros',
bias_initializer='zeros'))
model.add(layers.Dropout(0.1))
model.add(layers.Dense(1))
model.compile(optimizer='rmsprop',loss='mse',metrics=['mse'])
# -
history = model.fit(x_train_scaled, y_train, validation_split=0.2, epochs=10)
mse = model.evaluate(x_test_scaled, y_test)[1]
print('MSE without tuning: {}'.format(mse))
# ### Building the Hypermodel
class RegressionHyperModel(HyperModel):
def __init__(self, input_shape):
self.input_shape = input_shape
def build(self, hp):
model = Sequential()
model.add(
layers.Dense(
units=hp.Int('units', 8, 64, 4, default=8),
activation=hp.Choice(
'dense_activation',
values=['relu', 'tanh', 'sigmoid'],
default='relu'),
input_shape=input_shape,
kernel_initializer='zeros', bias_initializer='zeros'
)
)
model.add(
layers.Dense(
units=hp.Int('units', 16, 64, 4, default=16),
activation=hp.Choice(
'dense_activation',
values=['relu', 'tanh', 'sigmoid'],
default='relu'),
kernel_initializer='zeros', bias_initializer='zeros'
)
)
model.add(
layers.Dropout(
hp.Float(
'dropout',
min_value=0.0,
max_value=0.1,
default=0.005,
step=0.01)
)
)
model.add(layers.Dense(1, kernel_initializer='zeros', bias_initializer='zeros'))
model.compile(
optimizer='rmsprop',loss='mse',metrics=['mse']
)
return model
# ### Initialize hypermodel
input_shape = (x_train.shape[1],)
hypermodel = RegressionHyperModel(input_shape)
# ### Build Random Search Tuner
tuner_rs = RandomSearch(
hypermodel,
objective='mse',
seed=42,
max_trials=10,
executions_per_trial=2, overwrite=True
)
# ### Run Random Search
tuner_rs.search(x_train_scaled, y_train, epochs=10, validation_split=0.2, verbose=0)
# ### Evaluate Random Search
best_model = tuner_rs.get_best_models(num_models=1)[0]
mse_rs = best_model.evaluate(x_test_scaled, y_test)[1]
print('Random search MSE: ', mse_rs)
# ### Build, Run and Evaluate Hyperband Tuner
# +
tuner_hb = Hyperband(
hypermodel,
max_epochs=5,
objective='mse',
seed=42,
executions_per_trial=2,
directory='hb'
)
tuner_hb.search(x_train_scaled, y_train, epochs=10, validation_split=0.2, verbose=0)
best_model = tuner_hb.get_best_models(num_models=1)[0]
mse_hb = best_model.evaluate(x_test_scaled, y_test)[1]
# -
print('Hyperband MSE: ', mse_hb)
# ### Build, Run and Evaluate Bayesian Optimization Tuner
# +
tuner_bo = BayesianOptimization(
hypermodel,
objective='mse',
max_trials=10,
seed=42,
executions_per_trial=2,
directory='bo'
)
tuner_bo.search(x_train_scaled, y_train, epochs=10, validation_split=0.2, verbose=0)
best_model = tuner_bo.get_best_models(num_models=1)[0]
mse_bo = best_model.evaluate(x_test_scaled, y_test)[1]
# -
print('Bayesian Optimization MSE: ', mse_bo)
| Modules/keras_tuner/example_01/Hyperparameter Tuning Demo with Keras Tuner.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Multi-source data import
#
# Sources:
# - MIMIC-III. Covers the years 2001-2012. Has free-text notes.
# - MIMIC-IV. Covers the years 2008 - 2019. Has physician order entry data, reference ranges for lab values, and some other changes. Doesn't have free-text notes as of this writing.
# - UMLS. Provides a common set of concepts that form a central connection point for many other sources such as RxNorm and MeSH.
# - RxNorm. Has drug-drug and drug-disease interactions, indications, contraindications, etc.
# - MeSH. Has broader-narrower relationships among hierarchically-related terms.
# - Pubmed. Has the majority of the world's medical literature in free text, with abstracts freely available an accessible through an API.
# ## Information about each source
# ### MIMIC-III
# Schema of MIMIC-III: https://mit-lcp.github.io/mimic-schema-spy/index.html
# ### MIMIC-IV
# Documentation for MIMIC-IV (no schema on schema spy as of this writing):
# ### RxNorm
# Connect various forms/dosages/routes of a clinical drug to the underlying pharmacologic substance
# 
# Note the "TTY" field from the graph above corresponds to the heading of each box below.
# 
#
# Relate each pharmacologic substance to other drugs with interaction info
# 
#
# Connect clinically relevant properties of drugs
# 
#
# RxNorm main landing page: https://www.nlm.nih.gov/research/umls/rxnorm/index.html
# AMIA article describing RxNorm: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3128404/
# Data downloads: https://www.nlm.nih.gov/research/umls/rxnorm/docs/rxnormfiles.html
# Web-based browser: https://mor.nlm.nih.gov/RxNav/search?searchBy=String&searchTerm=coumadin
# Technical docs: https://www.nlm.nih.gov/research/umls/rxnorm/docs/index.html
#
# The full download of RxNorm files contains a directory called "rrf" with the following contents:
#
# RXNCONSO.RRF 121,180,353 bytes
# RXNDOC.RRF 218,467 bytes
# RXNREL.RRF 503,188,245 bytes
# RXNSAB.RRF 10,698 bytes
# RXNSAT.RRF 502,793,103 bytes
# RXNSTY.RRF 17,996,450 bytes
#
# Archival files for tracking RxNorm historical content:
# RXNATOMARCHIVE.RRF 74,069,962 bytes
# RXNCUICHANGES.RRF 39,589 bytes
# RXNCUI.RRF 1,716,694 bytes
import pandas as pd
# Load RXNREL.RRF into a dataframe
rxnrel = pd.read_csv('/home/tim/Documents/GrApH_AI/Data/RxNorm_full_06072021/rrf/RXNREL.RRF', sep='|', header=None, encoding='utf-8')
rxnrel[:5]
rxnrel.iloc[:,7].value_counts()
# Load RXNSAT.RRF (Simple Concept and Atom Attributes) into a dataframe
columns = ['RXCUI', 'LUI', 'SUI', 'RXAUI', 'STYPE', 'CODE', 'ATUI', 'SATUI', 'ATN', 'SAB', 'ATV', 'SUPPRESS', 'CVF'] # Column headers and descriptions at https://www.nlm.nih.gov/research/umls/rxnorm/docs/techdoc.html#sat
rxnsat = pd.read_csv('/home/tim/Documents/GrApH_AI/Data/RxNorm_full_06072021/rrf/RXNSAT.RRF', sep='|', header=None, encoding='utf-8')
rxnsat = rxnsat.iloc[:,:13] # Drop empty column at index 14
rxnsat.columns = columns
rxnsat[:5]
# RXSAT.RFF table info
#
# |Column|Description|
# |---|---|
# |RXCUI|Unique identifier for concept (concept id)|
# |LUI|Unique identifier for term (no value provided)|
# |SUI|Unique identifier for string (no value provided)|
# |RXAUI|RxNorm atom identifier (RXAUI) or RxNorm relationship identifier (RUI).|
# |STYPE|The name of the column in RXNCONSO.RRF or RXNREL.RRF that contains the identifier to which the attribute is attached, e.g., CUI, AUI.|
# |CODE|"Most useful" source asserted identifier (if the source vocabulary has more than one identifier), or a RxNorm-generated source entry identifier (if the source vocabulary has none.)|
# |ATUI|Unique identifier for attribute|
# |SATUI|Source asserted attribute identifier (optional - present if it exists)|
# |ATN|Attribute name (e.g. NDC). Possible values appear in RXNDOC.RRF and are described on the UMLS Attribute Names page|
# |SAB|Abbreviation of the source of the attribute. Possible values appear in RXNSAB.RRF and are listed on the UMLS Source Vocabularies page|
# |ATV|Attribute value described under specific attribute name on the UMLS Attribute Names page (e.g. 000023082503 where ATN = 'NDC'). A few attribute values exceed 1,000 characters. Many of the abbreviations used in attribute values are explained in RXNDOC.RRF and included UMLS Abbreviations Used in Data Elements page|
# |SUPPRESS|Suppressible flag. Values = O, Y, or N. Reflects the suppressible status of the attribute. N - Attribute is not suppressed. O - Attribute is suppressed at source level. Y - Attribute is suppressed by RxNorm editors.|
# |CVF|Content view flag. RxNorm includes one value, '4096', to denote inclusion in the Current Prescribable Content subset. All rows with CVF='4096' can be found in the subset.|
pd.set_option("display.max_rows", 120)
rxnsat['ATN'].value_counts() #Table listing attribute names and descriptions: https://www.nlm.nih.gov/research/umls/knowledge_sources/metathesaurus/release/attribute_names.html
# ### MED-RT
# Connect medications with other concept types such as diseases, phenotypes, etc.
#
# How MED-RT connects multiple source vocabularies:
# 
# Figure source: https://evs.nci.nih.gov/ftp1/MED-RT/MED-RT%20Documentation.pdf
#
# Sample of some relationships specified in MED-RT:
# 
# Screenshot source: https://www.nlm.nih.gov/research/umls/sourcereleasedocs/current/MED-RT/metarepresentation.html#relationships
#
# MEDRT_MoA_NUIs file is an index of mechanisms of action.
# Sample line from the file:
# Acetylcholine Release Inhibitors [MoA] N0000175770 MED-RT
# Possible ways to store the data:
# - Each line becomes a node with the label "Mechanism_of_Action"
# - Each line becomes a property of a drug node
#
# MEDRT_PE_NUIs file is an index of physiologic effects.
# Sample line from the file:
# Acetylcholine Activity Alteration [PE] N0000008290 MED-RT
# Possible ways to store the data:
# - Each line becomes a node with the label "Physiologic_Effect"
# - Each line becomes a property of an existing UMLS concept node
# ### Excerpt from MED-RT_Schema_v1.xsd
#
# AssociationDef - definition of Association
# <xs:complexType name="AssociationDef">
# <xs:annotation>
# <xs:documentation> This element includes all types of Associations: Synonyms, Term Associations and Concept Associations.
# </xs:documentation>
# </xs:annotation>
# <xs:sequence>
# <xs:element name="namespace" type="xs:token"/>
# <xs:element name="name" type="xs:token"/>
# <!-- name of AssociationType -->
# <xs:group ref="FromElement"/>
# <xs:group ref="ToElement"/>
# <xs:element name="qualifier" type="QualifierDef" minOccurs="0" maxOccurs="unbounded"/>
# </xs:sequence>
# </xs:complexType>
# <xs:group name="ToElement">
# <xs:annotation>
# <xs:documentation> A reference from the local Concept/Term to another Concept/Term (in any Namespace).
# </xs:documentation>
# </xs:annotation>
# <xs:sequence>
# <xs:element name="to_namespace" type="xs:token"/>
# <xs:element name="to_name" type="xs:token">
# <xs:annotation>
# <xs:documentation>MED-RT: Concept Name
# MeSH: Preferred Term
# RxNorm: Preferred Term
# SNOMED CT: FSN Synonym</xs:documentation>
# </xs:annotation>
# </xs:element>
# <!-- name of target Concept/Term -->
# <xs:element name="to_code" type="xs:token" minOccurs="0">
# <xs:annotation>
# <xs:documentation>MED-RT: NUI
# MeSH: Code in Source
# RxNorm: Code in Source
# SNOMED CT: Code in Source</xs:documentation>
# </xs:annotation>
# </xs:element>
# <!-- code of target Term -->
# </xs:sequence>
# </xs:group>
# <xs:group name="FromElement">
# <xs:annotation>
# <xs:documentation> A reference to the local Concept/Term from another Concept/Term (in a different Namespace).
# </xs:documentation>
# </xs:annotation>
# <xs:sequence>
# <xs:element name="from_namespace" type="xs:token"/>
# <xs:element name="from_name" type="xs:token">
# <xs:annotation>
# <xs:documentation>MED-RT: Concept Name
# MeSH: Preferred Term
# RxNorm: Preferred Term
# SNOMED CT: FSN Synonym</xs:documentation>
# </xs:annotation>
# </xs:element>
# <!-- name of source Concept/Term -->
# <xs:element name="from_code" type="xs:token">
# <xs:annotation>
# <xs:documentation>MED-RT: NUI
# MeSH: Code in Source
# RxNorm: Code in Source
# SNOMED CT: Code in Source</xs:documentation>
# </xs:annotation>
# </xs:element>
# <!-- code of source Term -->
# </xs:sequence>
# </xs:group>
# ### FDA's Structured Product Labels
# "The Structured Product Labeling (SPL) is a document markup standard approved by Health Level Seven (HL7) and adopted by FDA as a mechanism for exchanging product and facility information." - U.S. FDA
# SPL Resources: https://www.fda.gov/industry/fda-resources-data-standards/structured-product-labeling-resources
# Download data: https://dailymed.nlm.nih.gov/dailymed/spl-resources-all-drug-labels.cfm
# ### MeSH
# Connect heirarchically-related terms with broader-narrower relationships
# 
# MeSH contributes broader-narrower connections as displayed in the UMLS browser:
# 
#
# RDF format for MeSH: https://id.nlm.nih.gov/mesh/, https://hhs.github.io/meshrdf/
# Concept structure of MeSH: https://www.nlm.nih.gov/mesh/concept_structure.html
#
# ### SemMedDB
#
# SemMedDB can be downloaded as MySQL files or CSV files [here](https://ii.nlm.nih.gov/SemRep_SemMedDB_SKR/SemMedDB/SemMedDB_download.shtml). These are the CSV files:
#
# |TABLE NAME|Size compressed|Size uncompressed|# Rows|
# |---|---|---|---|
# |CITATIONS|152M|1.7G|32,470,549|
# |ENTITY|39G|160.5G|1,555,897,812|
# |GENERIC_CONCEPT|3.9K|9.3K|259|
# |PREDICATION|2.7G|15G|111,846,030|
# |PREDICATION_AUX|3.6G|16.4G|111,846,028|
# |SENTENCE|14G|43.8G|219,049,752|
#
#
# Schema of SemMedDB version 4.2 and later
# 
#
# Let's have a look at each table. Column names are obtained from the [schema documentation](https://ii.nlm.nih.gov/SemRep_SemMedDB_SKR/dbinfo.shtml).
import pandas as pd
columns = ['PMID', 'ISSN', 'DP', 'EDAT', 'PYEAR']
citations = pd.read_csv('/home/tim/Documents/GrApH_AI/Data/SemMedDB/semmedVER43_2021_R_CITATIONS.23871.csv', header=None, names = columns, nrows=100)
citations.head()
# Note that the order of columns in the actual data differs slightly from the documentation
columns = ['ENTITY_ID', 'SENTENCE_ID', 'CUI', 'NAME', 'TYPE', 'GENE_ID', 'GENE_NAME', 'TEXT', 'SCORE', 'START_INDEX', 'END_INDEX']
entity = pd.read_csv('/home/tim/Documents/GrApH_AI/Data/SemMedDB/semmedVER43_2021_R_ENTITY.23871.csv', header=None, names = columns, nrows=100)
entity.head()
columns = ['CONCEPT_ID', 'CUI', 'PREFERRED_NAME']
generic_concept = pd.read_csv('/home/tim/Documents/GrApH_AI/Data/SemMedDB/semmedVER43_2021_R_GENERIC_CONCEPT.csv', header=None, names = columns, nrows=100)
generic_concept.head()
# Note that three spurious columns exist after the last column named below, each containing '/n'
columns = ['PREDICATION_ID', 'SENTENCE_ID', 'PMID', 'PREDICATE', 'SUBJECT_CUI', 'SUBJECT_NAME', 'SUBJECT_SEMTYPE', 'SUBJECT_NOVELTY', 'OBJECT_CUI', 'OBJECT_NAME', 'OBJECT_SEMTYPE', 'OBJECT_NOVELTY']
predication = pd.read_csv('/home/tim/Documents/GrApH_AI/Data/SemMedDB/semmedVER43_2021_R_PREDICATION.23871.csv', usecols = [0,1,2,3,4,5,6,7,8,9,10,11], header=None, names = columns, nrows=100)
predication.head()
# Check what types of predicates exist
pd.set_option("display.max_rows", 120)
predicates = pd.read_csv('/home/tim/Documents/GrApH_AI/Data/SemMedDB/semmedVER43_2021_R_PREDICATION.23871.csv', usecols = [3], header=None)
predicates.value_counts()
columns = ['PREDICATION_AUX_ID', 'PREDICATION_ID', 'SUBJECT_TEXT', 'SUBJECT_DIST', 'SUBJECT_MAXDIST', 'SUBJECT_START_INDEX', 'SUBJECT_END_INDEX', 'SUBJECT_SCORE', 'INDICATOR_TYPE', 'PREDICATE_START_INDEX', 'PREDICATE_END_INDEX', 'OBJECT', 'CURR_TIMESTAMP']
predication_aux = pd.read_csv('/home/tim/Documents/GrApH_AI/Data/SemMedDB/semmedVER43_2021_R_PREDICATION_AUX.23871.csv', usecols = [0,1,2,3,4,5,6,7,8,9,10,11,12], header=None, names = columns, nrows=100)
predication_aux.head()
# Note that the order of columns in the actual data differs slightly from the documentation
columns = ['SENTENCE_ID', 'PMID', 'TYPE', 'NUMBER', 'SENT_START_INDEX', 'SENTENCE', 'SENT_END_INDEX', 'SECTION_HEADER', 'NORMALIZED_SECTION_HEADER']
sentence = pd.read_csv('/home/tim/Documents/GrApH_AI/Data/SemMedDB/semmedVER43_2021_R_SENTENCE.23871.csv', header=None, names = columns, nrows=100)
sentence.head()
for string_of_interest in sentence.SENTENCE[0:5]:
print(string_of_interest, '\n')
# ## Data model to connect the various data sources
# MIMIC-IV d_labitems loinc_code connects to UMLS by LOINC code
| Data_import/Multi-source_import.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
results_path = "qualification_results/"
# ## Load Data
persistence_ssa_results = pd.read_csv(results_path + "rolling_cv_wind_ssa_persistence.csv")
sarima_ssa_results = pd.read_csv(results_path + "rolling_cv_wind_ssa_sarima.csv")
var_ssa_results = pd.read_csv(results_path + "rolling_cv_wind_ssa_var.csv")
hofts_ssa_results = pd.read_csv(results_path + "rolling_cv_wind_ssa_hofts.csv")
cvfts_ssa_results = pd.read_csv(results_path + "rolling_cv_wind_ssa_cvfts.csv")
cmvfts_ssa_results = pd.read_csv(results_path + "rolling_cv_wind_ssa_cmvfts.csv")
lstm_multi_ssa_results = pd.read_csv(results_path + "rolling_cv_wind_ssa_lstm_multi.csv")
lstm_uni_ssa_results = pd.read_csv(results_path + "rolling_cv_wind_ssa_lstm_uni.csv")
mlp_multi_ssa_results = pd.read_csv(results_path + "rolling_cv_wind_ssa_mlp_multi.csv")
mlp_uni_ssa_results = pd.read_csv(results_path + "rolling_cv_wind_ssa_mlp_uni.csv")
RMSE_real = []
for i in cvfts_ssa_results.RMSE:
comp = complex(i)
RMSE_real.append(comp.real)
cvfts_ssa_results['RMSE'] = RMSE_real
U_real = []
for i in cvfts_ssa_results.U:
comp = complex(i)
U_real.append(comp.real)
cvfts_ssa_results['U'] = U_real
def createBoxplot(filename, data, xticklabels, ylabel):
# Create a figure instance
fig = plt.figure(1, figsize=(9, 6))
# Create an axes instance
ax = fig.add_subplot(111)
# Create the boxplot
bp = ax.boxplot(data, patch_artist=True)
## change outline color, fill color and linewidth of the boxes
for box in bp['boxes']:
# change outline color
box.set( color='#7570b3', linewidth=2)
# change fill color
box.set( facecolor = '#1b9e77' )
## change color and linewidth of the whiskers
for whisker in bp['whiskers']:
whisker.set(color='#7570b3', linewidth=2)
## change color and linewidth of the caps
for cap in bp['caps']:
cap.set(color='#7570b3', linewidth=2)
## change color and linewidth of the medians
for median in bp['medians']:
median.set(color='#b2df8a', linewidth=2)
## change the style of fliers and their fill
for flier in bp['fliers']:
flier.set(marker='o', color='#e7298a', alpha=0.5)
## Custom x-axis labels
ax.set_xticklabels(xticklabels)
ax.set_ylabel(ylabel)
plt.show()
fig.savefig(filename, bbox_inches='tight')
# ## Boxplot SSA Multivariate
# +
metric = 'RMSE'
multi_data = [persistence_ssa_results[metric], var_ssa_results[metric], cmvfts_ssa_results[metric], lstm_multi_ssa_results[metric], mlp_multi_ssa_results[metric]]
xticks = ['Persistence','VAR','CMVFTS','LSTM_MULTI','MLP_MULTI']
ylab = 'RMSE'
createBoxplot("boxplot_rmse_wind_ssa_multi", multi_data, xticks, ylab)
# +
metric = 'SMAPE'
multi_data = [persistence_ssa_results[metric], var_ssa_results[metric], cmvfts_ssa_results[metric], lstm_multi_ssa_results[metric], mlp_multi_ssa_results[metric]]
xticks = ['Persistence','VAR','CMVFTS','LSTM_MULTI','MLP_MULTI']
ylab = 'SMAPE'
createBoxplot("boxplot_smape_wind_ssa_multi", multi_data, xticks, ylab)
# +
metric = 'U'
multi_data = [persistence_ssa_results[metric], var_ssa_results[metric], cmvfts_ssa_results[metric], lstm_multi_ssa_results[metric], mlp_multi_ssa_results[metric]]
xticks = ['Persistence','VAR','CMVFTS','LSTM_MULTI','MLP_MULTI']
ylab = 'U Statistic'
createBoxplot("boxplot_u_wind_ssa_multi", multi_data, xticks, ylab)
# -
# ## Improvement table Multivariate
def improvement(metric_model, metric_persistence):
return (1 - (np.mean(metric_model) / np.mean(metric_persistence)))
index = ['Persistence','VAR','CMVFTS','LSTM_MULTI','MLP_MULTI']
columns = ['imp(RMSE)', 'imp(SMAPE)', 'imp(U)']
imp_df = pd.DataFrame(columns=columns, index=index)
# +
metric = 'RMSE'
imp_prst = improvement(persistence_ssa_results[metric], persistence_ssa_results[metric])
imp_var = improvement(var_ssa_results[metric], persistence_ssa_results[metric])
imp_cmvfts = improvement(cmvfts_ssa_results[metric], persistence_ssa_results[metric])
imp_lstm_multi = improvement(lstm_multi_ssa_results[metric], persistence_ssa_results[metric])
imp_mlp_multi = improvement(mlp_multi_ssa_results[metric], persistence_ssa_results[metric])
imp_df['imp('+metric+')'] = [imp_prst, imp_var, imp_cmvfts, imp_lstm_multi, imp_mlp_multi]
# +
metric = 'SMAPE'
imp_prst = improvement(persistence_ssa_results[metric], persistence_ssa_results[metric])
imp_var = improvement(var_ssa_results[metric], persistence_ssa_results[metric])
imp_cmvfts = improvement(cmvfts_ssa_results[metric], persistence_ssa_results[metric])
imp_lstm_multi = improvement(lstm_multi_ssa_results[metric], persistence_ssa_results[metric])
imp_mlp_multi = improvement(mlp_multi_ssa_results[metric], persistence_ssa_results[metric])
imp_df['imp('+metric+')'] = [imp_prst, imp_var, imp_cmvfts, imp_lstm_multi, imp_mlp_multi]
# +
metric = 'U'
imp_prst = improvement(persistence_ssa_results[metric], persistence_ssa_results[metric])
imp_var = improvement(var_ssa_results[metric], persistence_ssa_results[metric])
imp_cmvfts = improvement(cmvfts_ssa_results[metric], persistence_ssa_results[metric])
imp_lstm_multi = improvement(lstm_multi_ssa_results[metric], persistence_ssa_results[metric])
imp_mlp_multi = improvement(mlp_multi_ssa_results[metric], persistence_ssa_results[metric])
imp_df['imp('+metric+')'] = [imp_prst, imp_var, imp_cmvfts, imp_lstm_multi, imp_mlp_multi]
# -
print(imp_df.to_latex())
# ## Boxplot OAHU SSA Univariate
#
# ### SARIMA está FORA ate segunda ordem!!
# +
metric = 'RMSE'
#uni_data = [persistence_ssa_results[metric], sarima_ssa_results[metric], hofts_ssa_results[metric], cvfts_ssa_results[metric], lstm_uni_ssa_results[metric], mlp_uni_ssa_results[metric]]
#xticks = ['Persistence', 'SARIMA', 'HOFTS','CVFTS','LSTM_UNI','MLP_UNI']
uni_data = [persistence_ssa_results[metric], hofts_ssa_results[metric], cvfts_ssa_results[metric], lstm_uni_ssa_results[metric], mlp_uni_ssa_results[metric]]
xticks = ['Persistence', 'HOFTS','NSFTS','LSTM_UNI','MLP_UNI']
ylab = 'RMSE'
createBoxplot("boxplot_rmse_wind_ssa_uni", uni_data, xticks, ylab)
# +
metric = 'SMAPE'
uni_data = [persistence_ssa_results[metric], hofts_ssa_results[metric], cvfts_ssa_results[metric], lstm_uni_ssa_results[metric], mlp_uni_ssa_results[metric]]
xticks = ['Persistence', 'HOFTS','NSFTS','LSTM_UNI','MLP_UNI']
ylab = 'SMAPE'
createBoxplot("boxplot_smape_wind_ssa_uni", uni_data, xticks, ylab)
# +
metric = 'U'
uni_data = [persistence_ssa_results[metric], hofts_ssa_results[metric], cvfts_ssa_results[metric], lstm_uni_ssa_results[metric], mlp_uni_ssa_results[metric]]
xticks = ['Persistence', 'HOFTS','NSFTS','LSTM_UNI','MLP_UNI']
ylab = 'U Statistic'
createBoxplot("boxplot_u_wind_ssa_uni", uni_data, xticks, ylab)
# -
# ## Improvement Table Univariate
index = ['Persistence', 'HOFTS','CVFTS','LSTM_UNI','MLP_UNI']
columns = ['imp(RMSE)', 'imp(SMAPE)', 'imp(U)']
metrics = ['RMSE', 'SMAPE', 'U']
imp_df = pd.DataFrame(columns=columns, index=index)
for metric in metrics:
imp_prst = improvement(persistence_ssa_results[metric], persistence_ssa_results[metric])
imp_hofts = improvement(hofts_ssa_results[metric], persistence_ssa_results[metric])
imp_cvfts = improvement(cvfts_ssa_results[metric], persistence_ssa_results[metric])
imp_lstm_uni = improvement(lstm_uni_ssa_results[metric], persistence_ssa_results[metric])
imp_mlp_uni = improvement(mlp_uni_ssa_results[metric], persistence_ssa_results[metric])
imp_df['imp('+metric+')'] = [imp_prst, imp_hofts, imp_cvfts, imp_lstm_uni, imp_mlp_uni]
print(imp_df.to_latex())
# ## Hybrid Comparison
hybrid_ssa_results = pd.read_csv(results_path + "rolling_cv_wind_ssa_hybrid.csv")
# +
metric = 'RMSE'
multi_data = [persistence_ssa_results[metric], var_ssa_results[metric],lstm_multi_ssa_results[metric], hybrid_ssa_results[metric]]
xticks = ['Persistence','VAR','LSTM_MULTI','Hybrid']
ylab = 'RMSE'
createBoxplot("boxplot_rmse_wind_ssa_hybrid", multi_data, xticks, ylab)
# +
metric = 'SMAPE'
multi_data = [persistence_ssa_results[metric], var_ssa_results[metric],lstm_multi_ssa_results[metric], hybrid_ssa_results[metric]]
xticks = ['Persistence','VAR','LSTM_MULTI','Hybrid']
ylab = 'RMSE'
createBoxplot("boxplot_smape_wind_ssa_hybrid", multi_data, xticks, ylab)
# -
# ## Boxplot Oahu Raw Multivariate
persistence_raw_results = pd.read_csv(results_path + "rolling_cv_wind_raw_persistence.csv")
var_raw_results = pd.read_csv(results_path + "rolling_cv_wind_raw_var.csv")
hofts_raw_results = pd.read_csv(results_path + "rolling_cv_wind_raw_hofts.csv")
cvfts_raw_results = pd.read_csv(results_path + "rolling_cv_wind_raw_cvfts.csv")
cmvfts_raw_results = pd.read_csv(results_path + "rolling_cv_wind_raw_cmvfts.csv")
lstm_multi_raw_results = pd.read_csv(results_path + "rolling_cv_wind_raw_lstm_multi.csv")
lstm_uni_raw_results = pd.read_csv(results_path + "rolling_cv_wind_raw_lstm_uni.csv")
mlp_multi_raw_results = pd.read_csv(results_path + "rolling_cv_wind_raw_mlp_multi.csv")
mlp_uni_raw_results = pd.read_csv(results_path + "rolling_cv_wind_raw_mlp_uni.csv")
# +
RMSE_real = []
for i in cvfts_raw_results.RMSE:
comp = complex(i)
RMSE_real.append(comp.real)
cvfts_raw_results['RMSE'] = RMSE_real
U_real = []
for i in cvfts_raw_results.U:
comp = complex(i)
U_real.append(comp.real)
cvfts_raw_results['U'] = U_real
# +
metric = 'RMSE'
multi_data = [persistence_raw_results[metric], var_raw_results[metric], cmvfts_raw_results[metric], lstm_multi_raw_results[metric], mlp_multi_raw_results[metric]]
xticks = ['Persistence','VAR','CMVFTS','LSTM_MULTI','MLP_MULTI']
ylab = 'RMSE'
createBoxplot("boxplot_rmse_wind_raw_multi", multi_data, xticks, ylab)
# +
metric = 'SMAPE'
multi_data = [persistence_raw_results[metric], var_raw_results[metric], cmvfts_raw_results[metric], lstm_multi_raw_results[metric], mlp_multi_raw_results[metric]]
xticks = ['Persistence','VAR','CMVFTS','LSTM_MULTI','MLP_MULTI']
ylab = 'SMAPE'
createBoxplot("boxplot_smape_wind_raw_multi", multi_data, xticks, ylab)
# +
metric = 'U'
multi_data = [persistence_raw_results[metric], var_raw_results[metric], cmvfts_raw_results[metric], lstm_multi_raw_results[metric], mlp_multi_raw_results[metric]]
xticks = ['Persistence','VAR','CMVFTS','LSTM_MULTI','MLP_MULTI']
ylab = 'U Statistic'
createBoxplot("boxplot_u_wind_raw_multi", multi_data, xticks, ylab)
# -
# ## Improvement Table Raw Multivariate
index = ['Persistence','VAR','CMVFTS','LSTM_MULTI','MLP_MULTI']
columns = ['imp(RMSE)', 'imp(SMAPE)', 'imp(U)']
metrics = ['RMSE', 'SMAPE', 'U']
imp_df = pd.DataFrame(columns=columns, index=index)
# +
for metric in metrics:
imp_prst = improvement(persistence_raw_results[metric], persistence_raw_results[metric])
imp_var = improvement(var_raw_results[metric], persistence_raw_results[metric])
imp_cmvfts = improvement(cmvfts_raw_results[metric], persistence_raw_results[metric])
imp_lstm_multi = improvement(lstm_multi_raw_results[metric], persistence_raw_results[metric])
imp_mlp_multi = improvement(mlp_multi_raw_results[metric], persistence_raw_results[metric])
imp_df['imp('+metric+')'] = [imp_prst, imp_var, imp_cmvfts, imp_lstm_multi, imp_mlp_multi]
print(imp_df.to_latex())
# -
# ## Boxplot Oahu Raw Univariate
# +
metric = 'RMSE'
uni_data = [persistence_raw_results[metric], hofts_raw_results[metric], cvfts_raw_results[metric], lstm_uni_raw_results[metric], mlp_uni_raw_results[metric]]
xticks = ['Persistence', 'HOFTS','NSFTS','LSTM_UNI','MLP_UNI']
ylab = 'RMSE'
createBoxplot("boxplot_rmse_wind_raw_uni", uni_data, xticks, ylab)
# +
metric = 'SMAPE'
uni_data = [persistence_raw_results[metric], hofts_raw_results[metric], cvfts_raw_results[metric], lstm_uni_raw_results[metric], mlp_uni_raw_results[metric]]
xticks = ['Persistence', 'HOFTS','NSFTS','LSTM_UNI','MLP_UNI']
ylab = 'SMAPE'
createBoxplot("boxplot_smape_wind_raw_uni", uni_data, xticks, ylab)
# +
metric = 'U'
uni_data = [persistence_raw_results[metric], hofts_raw_results[metric], cvfts_raw_results[metric], lstm_uni_raw_results[metric], mlp_uni_raw_results[metric]]
xticks = ['Persistence', 'HOFTS','NSFTS','LSTM_UNI','MLP_UNI']
ylab = 'U Statistic'
createBoxplot("boxplot_u_wind_raw_uni", uni_data, xticks, ylab)
# -
# ## Improvement Table Raw Univariate
index = ['Persistence', 'HOFTS','CVFTS','LSTM_UNI','MLP_UNI']
columns = ['imp(RMSE)', 'imp(SMAPE)', 'imp(U)']
metrics = ['RMSE', 'SMAPE', 'U']
imp_df = pd.DataFrame(columns=columns, index=index)
for metric in metrics:
imp_prst = improvement(persistence_raw_results[metric], persistence_raw_results[metric])
imp_hofts = improvement(hofts_raw_results[metric], persistence_raw_results[metric])
imp_cvfts = improvement(cvfts_raw_results[metric], persistence_raw_results[metric])
imp_lstm_uni = improvement(lstm_uni_raw_results[metric], persistence_raw_results[metric])
imp_mlp_uni = improvement(mlp_uni_raw_results[metric], persistence_raw_results[metric])
imp_df['imp('+metric+')'] = [imp_prst, imp_hofts, imp_cvfts, imp_lstm_uni, imp_mlp_uni]
print(imp_df.to_latex())
| notebooks/180802 - Wind Results Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import requests
import urllib
import json
import pandas as pd
import numpy as np
import geopandas as gpd
import shapely
from shapely.geometry import Point, MultiPoint
from shapely import wkt
import shapely.speedups
from shapely.ops import transform, nearest_points
import plotly.express as px
from pyproj import crs
import plotly.graph_objects as go
import os
import gdal
import rasterio
from rasterio.mask import mask
from rasterio.warp import calculate_default_transform, reproject, Resampling
import glob
from functools import partial
import pyproj
import osmnx as ox
from IPython.display import Image
import scipy.ndimage as ndimage
from functools import reduce
### below files are local python programs. Make sure to paste them wherever you're running this notebook
import polygonize as pz
##
import geocoder
from pandana.loaders import osm
import pandana
import pylab as pl
ox.config(log_console=True, use_cache=True)
pl.rcParams["figure.figsize"] = (10,10)
# %pylab inline
def get_city_proj_crs(to_crs, val=0):
"""
Function to indentify local projection for cities dynamically
Input:
to_crs : name of city / country; epsg if known
Returns:
Local epsg (in string)
"""
if isinstance(to_crs, int):
to_crs = to_crs
elif isinstance(to_crs, str):
city, country = to_crs.split(',')
url = "http://epsg.io/?q={}&format=json&trans=1&callback=jsonpFunction".format(city)
r = requests.get(url)
if r.status_code == 200:
js = json.loads(r.text[14:-1])
if js['number_result'] != 0:
lis = []
for i in js['results']:
res = i
if (res['unit'] == 'metre') and (res['accuracy'] == 1.0):
lis.append(res['code'])
if len(lis) == 0:
for i in js['results']:
res = i
if res['unit'] == 'metre':
lis.append(res['code'])
return lis[val]
else:
return lis[val]
else:
if country.strip() == 'United Kingdom of Great Britain and Northern Ireland':
country = 'United Kingdom'
elif country.strip() == 'Venezuela (Bolivarian Republic of)':
country = 'Venezuela'
elif country.strip() == 'Viet Nam':
country = 'Vietnam'
url = "http://epsg.io/?q={}&format=json&trans=1&callback=jsonpFunction".format(country)
r = requests.get(url)
if r.status_code == 200:
js = json.loads(r.text[14:-1])
if js['number_result'] != 0:
lis = []
for i in js['results']:
res = i
if (res['unit'] == 'metre') and (res['accuracy'] == 1.0):
lis.append(res['code'])
if len(lis) == 0:
for i in js['results']:
res = i
if res['unit'] == 'metre':
lis.append(res['code'])
return lis[val]
else:
return lis[val]
def convert_geom_to_shp(shapely_polygon, city, out_crs=None):
string = city.split(',')[0]
df = pd.DataFrame(
{'City': [string],
'geometry': [wkt.dumps(shapely_polygon)]})
df['geometry'] = df['geometry'].apply(wkt.loads)
gdf = gpd.GeoDataFrame(df, geometry='geometry')
if out_crs:
gdf.crs = {'init' : 'epsg:{}'.format(out_crs)}
gdf.to_crs(epsg=4326, inplace=True)
else:
gdf.crs = {'init' : 'epsg:{}'.format(4326)}
return gdf
def getFeatures(gdf):
"""Function to parse features from GeoDataFrame in such a manner that rasterio accepts them"""
import json
return [json.loads(gdf.to_json())['features'][0]['geometry']]
def get_iso(city):
"""
Function to get ISO-3 codes for countries
Input:
city: city name (Ideally in (city, country) format)
Returns:
ISO-3 code for the country
"""
try:
country = city.split(',')[1].strip().lower()
if country == 'south korea': ### incorrect output for South Korea's ISO code with API
return 'kor'
elif country == 'india':
return 'ind'
elif country == 'iran':
return 'irn'
else:
url = "https://restcountries.eu/rest/v2/name/{}".format(country)
r = requests.get(url)
if len(r.json())>1 :
for i in range(len(r.json())):
if country in r.json()[i]['name'].lower():
return r.json()[i]['alpha3Code'].lower()
else:
return r.json()[0]['alpha3Code'].lower()
except IndexError:
url = "https://restcountries.eu/rest/v2/capital/{}".format(city)
r = requests.get(url)
return r.json()[0]['alpha3Code'].lower()
def factors(n):
return set(reduce(list.__add__,
([i, n//i] for i in range(1, int(n**0.5) + 1) if n % i == 0)))
# +
### Performs sliding window cumulative pop_tfa estimation for each pixel
def test_func(values):
#print (values)
return values.sum()
x = np.array([[1,2,3],[4,5,6],[7,8,9]])
footprint = np.array([[1,1,1],
[1,1,1],
[1,1,1]])
# -
def polygonize_raster(ras_path, shp_path, string):
"""
Function to polygonize a raster based on the pixel size of base raster.
Inputs:
ras_path: path to base raster location that is to be polygonized
shp_path: path to where the shapefile will be saved
string: name of the city
Returns:
Geodataframe with polygons equivalent to raster pixels.
"""
print("Polygonizing Raster!!")
import polygonize as pz
outSHPfn = shp_path
lat, lon = pz.main(ras_path,outSHPfn)
sh = gpd.read_file(shp_path)
rio = rasterio.open(ras_path)
sh.crs = rio.meta['crs']
shp_arr = np.array(sh.geometry).reshape(rio.shape[0], rio.shape[1])
pols = []
for row in range(shp_arr.shape[0]-1):
for col in range(shp_arr.shape[1]-1):
pols.append(shapely.geometry.box(shp_arr[row+1][col].x, shp_arr[row+1][col].y, shp_arr[row][col+1].x, shp_arr[row][col+1].y ))
gdf = gpd.GeoDataFrame()
gdf['ID'] = [i for i in range(len(pols))]
gdf['geometry'] = pols
gdf.set_geometry('geometry', inplace=True)
#gdf.crs = {'init':'epsg:4326'}
gdf.crs = rio.meta['crs']
print("Populating avearge height!!")
av_h = []
for i in gdf.geometry:
coords = getFeatures(convert_geom_to_shp(i, string))
out_img, out_transform = mask(dataset=rio, shapes=coords, crop=True)
av_h.append(out_img.sum()/out_img.shape[2])
gdf['avg_height'] = av_h
gdf.to_crs(epsg=4326, inplace=True)
gdf['Lon'] = [i.centroid.x for i in gdf.geometry]
gdf['Lat'] = [i.centroid.y for i in gdf.geometry]
return gdf
def get_population(city, gdf, wp = True, fb = None):
## Assiging Facebook population to pixels
iso = get_iso(city)
wp_pop = []
if fb:
pop = rasterio.open(r"C:\Users\wb542830\OneDrive - WBG\Facebook\population_{}_2018-10-01.tif".format(iso))
else:
pop = rasterio.open(r"M:\Gaurav\GPSUR\Data\WorldPop_2019\{}_ppp_2019.tif".format(iso))
for i in gdf.index:
_gdf = gdf[gdf.index == i]
#_gdf.to_crs(pop.meta['crs'], inplace=True)
_coords = getFeatures(_gdf)
try:
_out_img, _out_transform = mask(dataset=pop, shapes=_coords, crop=True)
outimg = np.nan_to_num(_out_img)
outimg = outimg.reshape(outimg.shape[1], outimg.shape[2])
wp_pop.append(outimg.sum())
except ValueError:
wp_pop.append(0)
return wp_pop
def get_hotspots(city):
string = city.split(',')[0]
### DLR raster file
dest_path = r"M:\Gaurav\GPSUR\Data\DLR Data\{}_WSF3D_AW3D30.tif".format(string)
ras = rasterio.open(dest_path)
## path to shapefile that will be editted
shp_path = r'C:\Users\wb542830\OneDrive - WBG\GPSUR\COVID\shapefiles\{}_ghsl_clip.shp'.format(string)
## Polygonize raster converts raster into polygon
gdf = polygonize_raster(dest_path, shp_path, string)
out_proj = get_city_proj_crs(city)
gdf_copy = gdf.to_crs(epsg= int(out_proj) )
gdf['pixel_area'] = [i.area for i in gdf_copy.geometry]
gdf['tfa'] = [(gdf.avg_height[i] * gdf.pixel_area[i]) / 3 for i in gdf.index]
pop = get_population(city, gdf, wp = True)
if len(pop) < 2:
pop = get_population(city, gdf, wp = False, fb = True)
gdf['pop_2019'] = pop
gdf['pop_2019'] = [i if i>0 else 0 for i in gdf.pop_2019]
gdf['pop_tfa'] = [gdf.pop_2019[i] / gdf.tfa[i] for i in gdf.index]
gdf['pop_tfa'] = [0 if pd.isna(i) else i for i in gdf.pop_tfa]
gdf['pop_tfa'] = [0 if i == np.inf else i for i in gdf.pop_tfa]
gdf['pop_tfa'] = [0 if i == -np.inf else i for i in gdf.pop_tfa]
fac = list(factors(len(gdf)))
a = fac[int(len(fac)/2)]
b = int(len(gdf) / a)
results = ndimage.generic_filter(np.array(gdf.pop_tfa).reshape(a,b), test_func, footprint=footprint)
gdf['poptfa_all'] = results.flatten()
return gdf
# ## %%time
gdf = get_hotspots("Manila, Philippines")
# ### Access to Services
#
# Steps below have to repeated based on different types of services.
x = ox.gdf_from_place("Manila, Philippines", which_result=2).geometry.iloc[0].envelope.bounds
def osm_pois_data(bounds, service):
if service == 'toilets':
amenities = ['toilets', 'washroom', 'restroom']
osm_tags = '"amenity"~"{}"'.format('|'.join(amenities))
elif service == 'water_points':
amenities = ['water_points', 'drinking_water', 'pumps', 'water_pumps', 'well']
osm_tags = '"amenity"~"{}"'.format('|'.join(amenities))
elif service == 'shops':
amenities = ['supermarket', 'convenience', 'general', 'department_stores', 'wholesale', 'grocery', 'general']
osm_tags = '"shop"~"{}"'.format('|'.join(amenities))
else:
osm_tags = None
pois = osm.node_query(bounds[1], bounds[0], bounds[3], bounds[2],tags=osm_tags)
pois['geometry'] = (list(zip(pois.lon,pois.lat)))
pois['geometry'] = pois.geometry.apply(lambda x: Point(x))
pois = gpd.GeoDataFrame(pois, geometry='geometry')
pois.crs = {'init':'epsg:4326'}
return pois
pois = (x, 'water_points')
out_crs = int(get_city_proj_crs("Manila, Philippines"))
pois_crs(epsg=out_crs, inplace=True)
dest = MultiPoint([i for i in pois.geometry])
gdf_copy = gdf.to_crs(epsg=out_crs) ## Converting gdf to local projection
dist = []
for i in gdf_copy.index:
if i % 10000 == 0:
print("{0} of {1} rows processed" .format(i, len(gdf_copy)))
temp_cent = gdf_copy.geometry.iloc[i].centroid
nearest_geoms = nearest_points(temp_cent)
dist.append(nearest_geoms[0].distance(nearest_geoms[1]))
gdf['dis_water'] = dist
gdf['weight'] = [1 / math.sqrt(i) if i > 70 else 1 for i in gdf.dis_water ]
## Adjusting for 'transit pixels'
pop_weight = (gdf.pop_tfa * gdf.weight) / 8
weight_slide = ndimage.generic_filter(np.array(pop_weight).reshape(276,252), test_func, footprint=footprint)
gdf['service_tfa'] = weight_slide.flatten()
## Each pixel's density + service pixel density
gdf['pixel_density'] = gdf.poptfa_all + gdf.service_tfa
gdf.to_file("Manila_Hotspots.shp")
| vulnerability_mapping/GAURAV_Compile_Hotspots.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import networkx as nx
import re
import matplotlib.pyplot as plt
import seaborn as sns
import os.path
import requests
from fa2 import ForceAtlas2
import community
sns.set()
df = pd.read_csv('characters/chars_data.csv')
df.Name = df.Name.replace(' ', '_', regex=True)
df
for char in df.Name:#df.name:
address = 'https://zelda.fandom.com/api.php?'
action="action=query"
titles=f"titles={char}"
content="prop=revisions&rvslots=*&rvprop=content"
dataformat="format=json"
query = '%s%s&%s&%s&%s' % (address, action, titles, content, dataformat)
response = requests.get(query)
content_char = response.text
with open(f'characters_full/{char}.txt', 'w') as f:
f.write(content_char)
pattern1 = r'(?<=\[\[).*?(?=\]\])'
pattern2 = r'(?:Term|Plural)[\w\| \(\)\'\\\s\=]*(?=\})'
G = nx.DiGraph()
for index, row in df.iterrows():
char = row['Name']
role = row.Role
if os.path.isfile(f'characters_full/{char}.txt') :
with open(f'characters_full/{char}.txt') as f:
contents = f.read()
G.add_node(char, role=row.Role, race=row.Race, gender=row.Gender, loc = len(re.findall(r'\w+', contents)))
links = []
if re.findall(pattern2, contents):
links += list(set([re.split(r'\|', name)[2] for name in re.findall(pattern2, contents) if len(re.split(r'\|', name)) > 2]))
links += list(set([re.split(r'\#|\|', name)[0] for name in re.findall(pattern1, contents)]))
links = [x.replace(" ", "_") for x in links]
char_links = list(set(links) & set(df.Name))
for link in char_links:
G.add_edge(char, link)
len(G.nodes)
len(G.edges)
largest_cc = max(nx.weakly_connected_components(G), key=len)
DG = G.subgraph(largest_cc)
UD = DG.to_undirected()
d = dict(UD.degree)
mapping = {"Enemy": 'red', "Boss": 'black', "Ally": 'blue'}
colors = [mapping[UD.nodes[n]['role']] for n in UD.nodes()]
edge_colors = [
"blue" if UD.nodes[edge[0]]['role'] == "Ally" and UD.nodes[edge[1]]['role'] == "Ally"
else "red" if UD.nodes[edge[0]]['role'] in ["Enemy", "Boss"] and UD.nodes[edge[1]]['role'] in ["Enemy", "Boss"]
else "black"
for edge in UD.edges()
]
# +
forceatlas2 = ForceAtlas2(
# Behavior alternatives
outboundAttractionDistribution=False, # Dissuade hubs
linLogMode=False, # NOT IMPLEMENTED
adjustSizes=False, # Prevent overlap (NOT IMPLEMENTED)
edgeWeightInfluence=1.5,
# Performance
jitterTolerance=1.0, # Tolerance
barnesHutOptimize=True,
barnesHutTheta=1.2,
multiThreaded=False, # NOT IMPLEMENTED
# Tuning
scalingRatio=1.0,
strongGravityMode=False,
gravity=1.5,
# Log
verbose=True)
plt.figure(figsize=(12,12))
positions = forceatlas2.forceatlas2_networkx_layout(UD, pos=None, iterations=2000)
# nx.draw_networkx_nodes(UD, positions, node_size=[v*5 for v in list(d.values())], node_color=colors, alpha=0.8)
nx.draw_networkx_nodes(UD, positions, node_size=[v*5 for v in list(d.values())], cmap=cmap, node_color=list(partition.values()), alpha=0.8)
nx.draw_networkx_edges(UD, positions, edge_color=edge_colors, alpha=0.5)
plt.axis('off')
plt.show()
# -
import community.community_louvain as community_louvain
import matplotlib.cm as cm
partition = community_louvain.best_partition(UD)
len(set(partition.values()))
partition
nx.set_node_attributes(UD, partition, "partition")
[node for node in UD.nodes if DG.nodes[node]['partition'] == 1]
partition.values()
plt.hist(partition.values())
plt.show()
cmap = cm.get_cmap('viridis', max(partition.values()) + 1)
nx.draw_networkx_nodes(UD, pos, partition.keys(), node_size=40,
cmap=cmap, node_color=list(partition.values()))
nx.draw_networkx_edges(UD, pos, alpha=0.5)
plt.show()
partition = community.best_partition
| lectures/Assignment 2 Parts 0 and 1.ipynb |
# # Gradient-boosting decision tree (GBDT)
#
# In this notebook, we will present the gradient boosting decision tree
# algorithm and contrast it with AdaBoost.
#
# Gradient-boosting differs from AdaBoost due to the following reason: instead
# of assigning weights to specific samples, GBDT will fit a decision tree on
# the residuals error (hence the name "gradient") of the previous tree.
# Therefore, each new tree in the ensemble predicts the error made by the
# previous learner instead of predicting the target directly.
#
# In this section, we will provide some intuition about the way learners are
# combined to give the final prediction. In this regard, let's go back to our
# regression problem which is more intuitive for demonstrating the underlying
# machinery.
# +
import pandas as pd
import numpy as np
# Create a random number generator that will be used to set the randomness
rng = np.random.RandomState(0)
def generate_data(n_samples=50):
"""Generate synthetic dataset. Returns `data_train`, `data_test`,
`target_train`."""
x_max, x_min = 1.4, -1.4
len_x = x_max - x_min
x = rng.rand(n_samples) * len_x - len_x / 2
noise = rng.randn(n_samples) * 0.3
y = x ** 3 - 0.5 * x ** 2 + noise
data_train = pd.DataFrame(x, columns=["Feature"])
data_test = pd.DataFrame(np.linspace(x_max, x_min, num=300),
columns=["Feature"])
target_train = pd.Series(y, name="Target")
return data_train, data_test, target_train
data_train, data_test, target_train = generate_data()
# +
import matplotlib.pyplot as plt
import seaborn as sns
sns.scatterplot(x=data_train["Feature"], y=target_train, color="black",
alpha=0.5)
_ = plt.title("Synthetic regression dataset")
# -
# As we previously discussed, boosting will be based on assembling a sequence
# of learners. We will start by creating a decision tree regressor. We will set
# the depth of the tree so that the resulting learner will underfit the data.
# +
from sklearn.tree import DecisionTreeRegressor
tree = DecisionTreeRegressor(max_depth=3, random_state=0)
tree.fit(data_train, target_train)
target_train_predicted = tree.predict(data_train)
target_test_predicted = tree.predict(data_test)
# -
# Using the term "test" here refers to data that was not used for training.
# It should not be confused with data coming from a train-test split, as it
# was generated in equally-spaced intervals for the visual evaluation of the
# predictions.
# +
# plot the data
sns.scatterplot(x=data_train["Feature"], y=target_train, color="black",
alpha=0.5)
# plot the predictions
line_predictions = plt.plot(data_test["Feature"], target_test_predicted, "--")
# plot the residuals
for value, true, predicted in zip(data_train["Feature"],
target_train,
target_train_predicted):
lines_residuals = plt.plot([value, value], [true, predicted], color="red")
plt.legend([line_predictions[0], lines_residuals[0]],
["Fitted tree", "Residuals"])
_ = plt.title("Prediction function together \nwith errors on the training set")
# -
# <div class="admonition tip alert alert-warning">
# <p class="first admonition-title" style="font-weight: bold;">Tip</p>
# <p class="last">In the cell above, we manually edited the legend to get only a single label
# for all the residual lines.</p>
# </div>
# Since the tree underfits the data, its accuracy is far from perfect on the
# training data. We can observe this in the figure by looking at the difference
# between the predictions and the ground-truth data. We represent these errors,
# called "Residuals", by unbroken red lines.
#
# Indeed, our initial tree was not expressive enough to handle the complexity
# of the data, as shown by the residuals. In a gradient-boosting algorithm, the
# idea is to create a second tree which, given the same data `data`, will try
# to predict the residuals instead of the vector `target`. We would therefore
# have a tree that is able to predict the errors made by the initial tree.
#
# Let's train such a tree.
# +
residuals = target_train - target_train_predicted
tree_residuals = DecisionTreeRegressor(max_depth=5, random_state=0)
tree_residuals.fit(data_train, residuals)
target_train_predicted_residuals = tree_residuals.predict(data_train)
target_test_predicted_residuals = tree_residuals.predict(data_test)
# +
sns.scatterplot(x=data_train["Feature"], y=residuals, color="black", alpha=0.5)
line_predictions = plt.plot(
data_test["Feature"], target_test_predicted_residuals, "--")
# plot the residuals of the predicted residuals
for value, true, predicted in zip(data_train["Feature"],
residuals,
target_train_predicted_residuals):
lines_residuals = plt.plot([value, value], [true, predicted], color="red")
plt.legend([line_predictions[0], lines_residuals[0]],
["Fitted tree", "Residuals"], bbox_to_anchor=(1.05, 0.8),
loc="upper left")
_ = plt.title("Prediction of the previous residuals")
# -
# We see that this new tree only manages to fit some of the residuals. We will
# focus on a specific sample from the training set (i.e. we know that the
# sample will be well predicted using two successive trees). We will use this
# sample to explain how the predictions of both trees are combined. Let's first
# select this sample in `data_train`.
sample = data_train.iloc[[-2]]
x_sample = sample['Feature'].iloc[0]
target_true = target_train.iloc[-2]
target_true_residual = residuals.iloc[-2]
# Let's plot the previous information and highlight our sample of interest.
# Let's start by plotting the original data and the prediction of the first
# decision tree.
# +
# Plot the previous information:
# * the dataset
# * the predictions
# * the residuals
sns.scatterplot(x=data_train["Feature"], y=target_train, color="black",
alpha=0.5)
plt.plot(data_test["Feature"], target_test_predicted, "--")
for value, true, predicted in zip(data_train["Feature"],
target_train,
target_train_predicted):
lines_residuals = plt.plot([value, value], [true, predicted], color="red")
# Highlight the sample of interest
plt.scatter(sample, target_true, label="Sample of interest",
color="tab:orange", s=200)
plt.xlim([-1, 0])
plt.legend(bbox_to_anchor=(1.05, 0.8), loc="upper left")
_ = plt.title("Tree predictions")
# -
# Now, let's plot the residuals information. We will plot the residuals
# computed from the first decision tree and show the residual predictions.
# +
# Plot the previous information:
# * the residuals committed by the first tree
# * the residual predictions
# * the residuals of the residual predictions
sns.scatterplot(x=data_train["Feature"], y=residuals,
color="black", alpha=0.5)
plt.plot(data_test["Feature"], target_test_predicted_residuals, "--")
for value, true, predicted in zip(data_train["Feature"],
residuals,
target_train_predicted_residuals):
lines_residuals = plt.plot([value, value], [true, predicted], color="red")
# Highlight the sample of interest
plt.scatter(sample, target_true_residual, label="Sample of interest",
color="tab:orange", s=200)
plt.xlim([-1, 0])
plt.legend()
_ = plt.title("Prediction of the residuals")
# -
# For our sample of interest, our initial tree is making an error (small
# residual). When fitting the second tree, the residual in this case is
# perfectly fitted and predicted. We will quantitatively check this prediction
# using the fitted tree. First, let's check the prediction of the initial tree
# and compare it with the true value.
# +
print(f"True value to predict for "
f"f(x={x_sample:.3f}) = {target_true:.3f}")
y_pred_first_tree = tree.predict(sample)[0]
print(f"Prediction of the first decision tree for x={x_sample:.3f}: "
f"y={y_pred_first_tree:.3f}")
print(f"Error of the tree: {target_true - y_pred_first_tree:.3f}")
# -
# As we visually observed, we have a small error. Now, we can use the second
# tree to try to predict this residual.
print(f"Prediction of the residual for x={x_sample:.3f}: "
f"{tree_residuals.predict(sample)[0]:.3f}")
# We see that our second tree is capable of predicting the exact residual
# (error) of our first tree. Therefore, we can predict the value of `x` by
# summing the prediction of all the trees in the ensemble.
y_pred_first_and_second_tree = (
y_pred_first_tree + tree_residuals.predict(sample)[0]
)
print(f"Prediction of the first and second decision trees combined for "
f"x={x_sample:.3f}: y={y_pred_first_and_second_tree:.3f}")
print(f"Error of the tree: {target_true - y_pred_first_and_second_tree:.3f}")
# We chose a sample for which only two trees were enough to make the perfect
# prediction. However, we saw in the previous plot that two trees were not
# enough to correct the residuals of all samples. Therefore, one needs to
# add several trees to the ensemble to successfully correct the error
# (i.e. the second tree corrects the first tree's error, while the third tree
# corrects the second tree's error and so on).
#
# We will compare the generalization performance of random-forest and gradient
# boosting on the California housing dataset.
# +
from sklearn.datasets import fetch_california_housing
from sklearn.model_selection import cross_validate
data, target = fetch_california_housing(return_X_y=True, as_frame=True)
target *= 100 # rescale the target in k$
# +
from sklearn.ensemble import GradientBoostingRegressor
gradient_boosting = GradientBoostingRegressor(n_estimators=200)
cv_results_gbdt = cross_validate(
gradient_boosting, data, target, scoring="neg_mean_absolute_error",
n_jobs=2,
)
# -
print("Gradient Boosting Decision Tree")
print(f"Mean absolute error via cross-validation: "
f"{-cv_results_gbdt['test_score'].mean():.3f} +/- "
f"{cv_results_gbdt['test_score'].std():.3f} k$")
print(f"Average fit time: "
f"{cv_results_gbdt['fit_time'].mean():.3f} seconds")
print(f"Average score time: "
f"{cv_results_gbdt['score_time'].mean():.3f} seconds")
# +
from sklearn.ensemble import RandomForestRegressor
random_forest = RandomForestRegressor(n_estimators=200, n_jobs=2)
cv_results_rf = cross_validate(
random_forest, data, target, scoring="neg_mean_absolute_error",
n_jobs=2,
)
# -
print("Random Forest")
print(f"Mean absolute error via cross-validation: "
f"{-cv_results_rf['test_score'].mean():.3f} +/- "
f"{cv_results_rf['test_score'].std():.3f} k$")
print(f"Average fit time: "
f"{cv_results_rf['fit_time'].mean():.3f} seconds")
print(f"Average score time: "
f"{cv_results_rf['score_time'].mean():.3f} seconds")
# In term of computation performance, the forest can be parallelized and will
# benefit from using multiple cores of the CPU. In terms of scoring
# performance, both algorithms lead to very close results.
#
# However, we see that the gradient boosting is a very fast algorithm to
# predict compared to random forest. This is due to the fact that gradient
# boosting uses shallow trees. We will go into details in the next notebook
# about the hyperparameters to consider when optimizing ensemble methods.
| notebooks/ensemble_gradient_boosting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: py37_pytorch
# language: python
# name: conda-env-py37_pytorch-py
# ---
# # Pre-trained models and transfer learning
#
# Training CNNs can take a lot of time, and a lot of data is required for that task. However, much of the time is spent to learn the best low-level filters that a network is using to extract patterns from images. A natural question arises - can we use a neural network trained on one dataset and adapt it to classifying different images without full training process?
#
# This approach is called **transfer learning**, because we transfer some knowledge from one neural network model to another. In transfer learning, we typically start with a pre-trained model, which has been trained on some large image dataset, such as **ImageNet**. Those models can already do a good job extracting different features from generic images, and in many cases just building a classifier on top of those extracted features can yield a good result.
# +
import torch
import torch.nn as nn
import torchvision
import torchvision.transforms as transforms
import matplotlib.pyplot as plt
from torchinfo import summary
import numpy as np
import os
from pytorchcv import train, plot_results, display_dataset, train_long, check_image_dir
# -
# ## Cats vs. Dogs Dataset
#
# In this unit, we will solve a real-life problem of classifying images of cats and dogs. For this reason, we will use [Kaggle Cats vs. Dogs Dataset](https://www.kaggle.com/c/dogs-vs-cats), which can also be downloaded [from Microsoft](https://www.microsoft.com/en-us/download/details.aspx?id=54765).
#
# Let's download this dataset and extract it into `data` directory (this process may take some time!):
# + tags=[]
if not os.path.exists('data/kagglecatsanddogs_3367a.zip'):
# !wget -P data -q https://download.microsoft.com/download/3/E/1/3E1C3F21-ECDB-4869-8368-6DEBA77B919F/kagglecatsanddogs_3367a.zip
# -
import zipfile
if not os.path.exists('data/PetImages'):
with zipfile.ZipFile('data/kagglecatsanddogs_3367a.zip', 'r') as zip_ref:
zip_ref.extractall('data')
# Unfortunately, there are some corrupt image files in the dataset. We need to do quick cleaning to check for corrupted files. In order not to clobber this tutorial, we moved the code to verify dataset into a module.
check_image_dir('data/PetImages/Cat/*.jpg')
check_image_dir('data/PetImages/Dog/*.jpg')
# Next, let's load the images into PyTorch dataset, converting them to tensors and doing some normalization. We will apply `std_normalize` transform to bring images to the range expected by pre-trained VGG network:
# +
std_normalize = transforms.Normalize(mean=[0.485, 0.456, 0.406],
std=[0.229, 0.224, 0.225])
trans = transforms.Compose([
transforms.Resize(256),
transforms.CenterCrop(224),
transforms.ToTensor(),
std_normalize])
dataset = torchvision.datasets.ImageFolder('data/PetImages',transform=trans)
trainset, testset = torch.utils.data.random_split(dataset,[20000,len(dataset)-20000])
display_dataset(dataset)
# -
# ## Pre-trained models
#
# There are many different pre-trained models available inside `torchvision` module, and even more models can be found on the Internet. Let's see how simplest VGG-16 model can be loaded and used:
vgg = torchvision.models.vgg16(pretrained=True)
sample_image = dataset[0][0].unsqueeze(0)
res = vgg(sample_image)
print(res[0].argmax())
# The result that we have received is a number of an `ImageNet` class, which can be looked up [here](https://gist.github.com/yrevar/942d3a0ac09ec9e5eb3a). We can use the following code to automatically load this class table and return the result:
# +
import json, requests
class_map = json.loads(requests.get("https://s3.amazonaws.com/deep-learning-models/image-models/imagenet_class_index.json").text)
class_map = { int(k) : v for k,v in class_map.items() }
class_map[res[0].argmax().item()]
# -
# Let's also see the architecture of the VGG-16 network:
summary(vgg,input_size=(1,3,224,224))
# In addition to the layer we already know, there is also another layer type called **Dropout**. These layers act as **regularization** technique. Regularization makes slight modifications to the learning algorithm so the model generalizes better. During training, dropout layers discard some proportion (around 30%) of the neurons in the previous layer, and training happens without them. This helps to get the optimization process out of local minima, and to distribute decisive power between different neural paths, which improves overall stability of the network.
#
# ## GPU computations
#
# Deep neural networks, such as VGG-16 and other more modern architectures require quite a lot of computational power to run. It makes sense to use GPU acceleration, if it is available. In order to do so, we need to explicitly move all tensors involved in the computation to GPU.
#
# The way it is normally done is to check the availability of GPU in the code, and define `device` variable that points to the computational device - either GPU or CPU.
#
# +
device = 'cuda' if torch.cuda.is_available() else 'cpu'
print('Doing computations on device = {}'.format(device))
vgg.to(device)
sample_image = sample_image.to(device)
vgg(sample_image).argmax()
# -
#
# ## Extracting VGG features
#
# If we want to use VGG-16 to extract features from our images, we need the model without final classification layers. In fact, this "feature extractor" can be obtained using `vgg.features` method:
res = vgg.features(sample_image).cpu()
plt.figure(figsize=(15,3))
plt.imshow(res.detach().view(-1,512))
print(res.size())
# The dimension of feature tensor is 512x7x7, but in order to visualize it we had to reshape it to 2D form.
#
# Now let's try to see if those features can be used to classify images. Let's manually take some portion of images (800 in our case), and pre-compute their feature vectors. We will store the result in one big tensor called `feature_tensor`, and also labels into `label_tensor`:
bs = 8
dl = torch.utils.data.DataLoader(dataset,batch_size=bs,shuffle=True)
num = bs*100
feature_tensor = torch.zeros(num,512*7*7).to(device)
label_tensor = torch.zeros(num).to(device)
i = 0
for x,l in dl:
with torch.no_grad():
f = vgg.features(x.to(device))
feature_tensor[i:i+bs] = f.view(bs,-1)
label_tensor[i:i+bs] = l
i+=bs
print('.',end='')
if i>=num:
break
# Now we can define `vgg_dataset` that takes data from this tensor, split it into training and test sets using `random_split` function, and train a small one-layer dense classifier network on top of extracted features:
# +
vgg_dataset = torch.utils.data.TensorDataset(feature_tensor,label_tensor.to(torch.long))
train_ds, test_ds = torch.utils.data.random_split(vgg_dataset,[700,100])
train_loader = torch.utils.data.DataLoader(train_ds,batch_size=32)
test_loader = torch.utils.data.DataLoader(test_ds,batch_size=32)
net = torch.nn.Sequential(torch.nn.Linear(512*7*7,2),torch.nn.LogSoftmax()).to(device)
history = train(net,train_loader,test_loader)
# -
# The result is great, we can distinguish between a cat and a dog with almost 98% probability! However, we have only tested this approach on a small subset of all images, because manual feature extraction seems to take a lot of time.
#
# ## Transfer learning using one VGG network
#
# We can also avoid manually pre-computing the features by using the original VGG-16 network as a whole during training. Let's look at the VGG-16 object structure:
print(vgg)
# You can see that the network contains:
# * feature extractor (`features`), comprised of a number of convolutional and pooling layers
# * average pooling layer (`avgpool`)
# * final `classifier`, consisting of several dense layers, which turns 25088 input features into 1000 classes (which is the number of classes in ImageNet)
#
# To train the end-to-end model that will classify our dataset, we need to:
# * **replace the final classifier** with the one that will produce required number of classes. In our case, we can use one `Linear` layer with 25088 inputs and 2 output neurons.
# * **freeze weights of convolutional feature extractor**, so that they are not trained. It is recommended to initially do this freezing, because otherwise untrained classifier layer can destroy the original pre-trained weights of convolutional extractor. Freezing weights can be accomplished by setting `requires_grad` property of all parameters to `False`
# +
vgg.classifier = torch.nn.Linear(25088,2).to(device)
for x in vgg.features.parameters():
x.requires_grad = False
summary(vgg,(1, 3,244,244))
# -
# As you can see from the summary, this model contain around 15 million total parameters, but only 50k of them are trainable - those are the weights of classification layer. That is good, because we are able to fine-tune smaller number of parameters with smaller number of examples.
#
# Now let's train the model using our original dataset. This process will take a long time, so we will use `train_long` function that will print some intermediate results without waiting for the end of epoch. It is highly recommended to run this training on GPU-enabled compute!
# +
trainset, testset = torch.utils.data.random_split(dataset,[20000,len(dataset)-20000])
train_loader = torch.utils.data.DataLoader(trainset,batch_size=16)
test_loader = torch.utils.data.DataLoader(testset,batch_size=16)
train_long(vgg,train_loader,test_loader,loss_fn=torch.nn.CrossEntropyLoss(),epochs=1,print_freq=90)
# -
# It looks like we have obtained reasonably accurate cats vs. dogs classifier! Let's save it for future use!
torch.save(vgg,'data/cats_dogs.pth')
# We can then load the model from file at any time. You may find it useful in case the next experiment destroys the model - you would not have to re-start from scratch.
vgg = torch.load('data/cats_dogs.pth')
# ## Fine-tuning transfer learning
#
# In the previous section, we have trained the final classifier layer to classify images in our own dataset. However, we did not re-train the feature extractor, and our model relied on the features that the model has learned on ImageNet data. If your objects visually differ from ordinary ImageNet images, this combination of features might not work best. Thus it makes sense to start training convolutional layers as well.
#
# To do that, we can unfreeze the convolutional filter parameters that we have previously frozen.
#
# > **Note:** It is important that you freeze parameters first and perform several epochs of training in order to stabilize weights in the classification layer. If you immediately start training end-to-end network with unfrozen parameters, large errors are likely to destroy the pre-trained weights in the convolutional layers.
for x in vgg.features.parameters():
x.requires_grad = True
# After unfreezing, we can do a few more epochs of training. You can also select lower learning rate, in order to minimize the impact on the pre-trained weights. However, even with low learning rate, you can expect the accuracy to drop in the beginning of the training, until finally reaching slightly higher level than in the case of fixed weights.
#
# > **Note:** This training happens much slower, because we need to propagate gradients back through many layers of the network! You may want to watch the first few minibatches to see the tendency, and then stop the computation.
train_long(vgg,train_loader,test_loader,loss_fn=torch.nn.CrossEntropyLoss(),epochs=1,print_freq=90,lr=0.0001)
# ## Other computer vision models
#
# VGG-16 is one of the simplest computer vision architectures. `torchvision` package provides many more pre-trained networks. The most frequently used ones among those are **ResNet** architectures, developed by Microsoft, and **Inception** by Google. For example, let's explore the architecture of the simplest ResNet-18 model (ResNet is a family of models with different depth, you can try experimenting with ResNet-151 if you want to see what a really deep model looks like):
resnet = torchvision.models.resnet18()
print(resnet)
# As you can see, the model contains the same building blocks: feature extractor and final classifier (`fc`). This allows us to use this model in exactly the same manner as we have been using VGG-16 for transfer learning. You can try experimenting with the code above, using different ResNet models as the base model, and see how accuracy changes.
#
# ## Batch Normalization
#
# This network contains yet another type of layer: **Batch Normalization**. The idea of batch normalization is to bring values that flow through the neural network to right interval. Usually neural networks work best when all values are in the range of [-1,1] or [0,1], and that is the reason that we scale/normalize our input data accordingly. However, during training of a deep network, it can happen that values get significantly out of this range, which makes training problematic. Batch normalization layer computes average and standard deviation for all values of the current minibatch, and uses them to normalize the signal before passing it through a neural network layer. This significantly improves the stability of deep networks.
#
# ## Takeaway
#
# Using transfer learning, we were able to quickly put together a classifier for our custom object classification task, and achieve high accuracy. However, this example was not completely fair, because original VGG-16 network was pre-trained to recognize cats and dogs, and thus we were just reusing most of the patterns that were already present in the network. You can expect lower accuracy on more exotic domain-specific objects, such as details on production line in a plant, or different tree leaves.
#
# You can see that more complex tasks that we are solving now require higher computational power, and cannot be easily solved on the CPU. In the next unit, we will try to use more lightweight implementation to train the same model using lower compute resources, which results in just slightly lower accuracy.
| computer-vision-pytorch/6-transfer-learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 10001st prime
#
# # Problem 7
# By listing the first six prime numbers: 2, 3, 5, 7, 11, and 13, we can see that the 6th prime is 13.
#
# What is the 10 001st prime number?
# # Solution 7
def test_primality(n):
if n<4: return True
limit = int(n**0.5)+1
for i in range(2,limit+1):
if n%i == 0: return False
return True
def nth_prime_num(nth):
count = 0
num = 2
while (1):
is_prime = test_primality(num)
if is_prime:
count += 1
else:
num += 1
continue
if count == nth:
return num
num += 1
nth_prime_num(10001)
| solutions/S0007.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 3D Stellar Plot
#
# # WARNING! - This currently plots the supernovae in the wrong location...
# +
# # Todo
# [] Figure out astropy cartesian coordinate directions. Explicit XYZ to UVW mapping.
# [] Figure out SIMBAD reference frame for RA/DEC and then correctly convert SkyCoord object using the same reference frame. See: https://astroquery.readthedocs.io/en/latest/simbad/simbad.html
# >>> customSimbad.add_votable_fields('ra(2;A;ICRS;J2017.5;2000)', 'dec(2;D;ICRS;2017.5;2000)')
#
# In[1]:
#get_ipython().run_line_magic('matplotlib', 'notebook')
# #%matplotlib inline
# In[2]:
import time
from pygments import highlight
from pygments.lexers import PythonLexer
from pygments.formatters import Terminal256Formatter
from pprint import pformat
def pprint_color(obj):
print(highlight(pformat(obj), PythonLexer(), Terminal256Formatter()))
from astropy.table import Table, vstack
from astroquery.simbad import conf
# Available SIMBAD mirrors.
#conf.server = "simbad.u-strasbg.fr"
conf.server = "simbad.harvard.edu" # Harvard's mirror is generally much faster than the main Strasbourg server for US users.
from astroquery.simbad import Simbad
# Set initial config
Simbad.SIMBAD_URL = "http://{}/simbad/sim-script".format(conf.server) # Shouldn't have to do this, probably a bug
Simbad.ROW_LIMIT = 100 # Limit the number of results returned, a zero value "0" will return all rows
# In[3]:
#print(Simbad.SIMBAD_URL)
#print(conf.server)
# In[4]:
#Simbad.list_votable_fields()
# In[5]:
#Simbad.get_field_description('v*')
# In[6]:
Simbad.get_votable_fields()
# In[7]:
Simbad.reset_votable_fields()
# In[8]:
#Simbad.add_votable_fields('coordinates','parallax','otype','ids','ubv','sptype','diameter')
Simbad.add_votable_fields('typed_id','id(NAME)','coo(d;;ICRS;;)','parallax', 'otype','ubv','sptype','fluxdata(B)','fluxdata(V)')
# In[9]:
# http://simbad.u-strasbg.fr/simbad/sim-display?data=otypes
# http://simbad.u-strasbg.fr/simbad/sim-fsam
# http://simbad.u-strasbg.fr/Pages/guide/chF.htx
# plx >= 300 & (otype='Star' & maintype != '**')
# This query selects stars with a Parallax greater than or equal to 300mas and
# where the main object type is a star, filtering out binary star system groupings
# Only include individual members of multiple systems
#result = Simbad.query_criteria('plx >= 300', otype='Star')
# TODO: grab
#result = Simbad.query_criteria('plx >= 300 & (otype="Star" & maintype != "**")')
#result = Simbad.query_criteria('plx >= 200', otype='Star')
#result = Simbad.query_criteria('(plx >= 200) | (plx > 50 & plx < 200 & Vmag < 6 )', otype='Star')
#result = Simbad.query_criteria('(plx > 3 & plx < 100 & Vmag < 3 )', otype='Star')
# Directional query pointed at main Centaurus association (ASSOC II SCO)
# ICRS: 16 15 00.0 -24 12 00
# Galactic: 351.3857 +18.9749
# TBD
# Use association identifier
#result = Simbad.query_criteria("region(circle,'ASSOC II SCO', 10d) & (plx > 3 & plx < 100 & Vmag < 4)", otype="Star")
# Directional query pointed at our two target associations (UCL & LCC)
# Use association identifiers
# Limited by 600sq degree region size constraint
#result = Simbad.query_criteria("(region(circle,'NAME Upper Centaurus Lupus', 5d) | region(circle,'NAME Lower Centaurus Crux', 5d)) & (plx > 3 & plx < 5 & Vmag < 4)", otype="Star")
# Coords
#NAME Upper Centaurus Lupus 15 24 -41.9 331.0211 +12.5023 15.4000000 -41.9000000
# NAME Lower Centaurus Crux 13 19 -57.1 298.5171 +05.4934 12.3166667 -57.1000000
# Draw a galactic box
# Query an identifier
#result = Simbad.query_object("ASSOC II SCO")
# Query multiple identifiers
#result = Simbad.query_objects(['NAME Upper Centaurus Lupus', 'NAME Lower Centaurus Crux'])
#UCL Box from Preibisch & Mamajek / Zeeuw fig 2
#
# Lower left Upper Left Upper Right Lower Right
# Gal 350 0 350 25 312 25 312 0
# Try polygon region
# Too big (948.763 sq. deg)
#result = Simbad.query_criteria("region(polygon gal,350 +0,350 +25,312 +25,312 +0) & (plx > 5 & plx < 8 & Vmag < 8)", otype="Star")
# Lets try halving it
# works!
#result = Simbad.query_criteria("region(polygon gal,350 +0,350 +25,336 +25,336 +0) & (plx > 5 & plx < 8 & Vmag < 8)", otype="Star")
# How about two regions?
# Nope...
#result = Simbad.query_criteria("(region(polygon gal,350 +0,350 +25,336 +25,336 +0) | region(polygon gal,336 +0,336 +25,312 +25,312 +0) ) & (plx > 5 & plx < 8 & Vmag < 8)", otype="Star")
# Separate queries then
result_ucl1 = Simbad.query_criteria("region(polygon gal,350 +0,350 +25,336 +25,336 +0) & (plx > 5 & plx < 10 & Vmag < 6)", otype="Star")
#time.sleep(2)
result_ucl2 = Simbad.query_criteria("region(polygon gal,336 +0,336 +25,312 +25,312 +0) & (plx > 5 & plx < 10 & Vmag < 6)", otype="Star")
#time.sleep(2)
# Vertically stack these two result tables
result_ucl = vstack([result_ucl1, result_ucl2])
# Add subgroup column
result_ucl['subgroup'] = 'UCL'
# Separate queries then
result_lcc1 = Simbad.query_criteria("region(polygon gal,312 -10,312 +21,298 +21,298 -10) & (plx > 5 & plx < 10 & Vmag < 6)", otype="Star")
#time.sleep(2)
result_lcc2 = Simbad.query_criteria("region(polygon gal,298 -10,298 +21,285 +21,285 -10) & (plx > 5 & plx < 10 & Vmag < 6)", otype="Star")
# Vertically stack these two result tables
result_lcc = vstack([result_lcc1, result_lcc2])
# Add subgroup column
result_lcc['subgroup'] = 'LCC'
result = vstack([result_ucl, result_lcc])
print(result)
# In[10]:
result.show_in_notebook()
# In[11]:
import re
# Lifted from: https://stackoverflow.com/questions/21977786/star-b-v-color-index-to-apparent-rgb-color
def bv2rgb(bv):
if bv < -0.40: bv = -0.40
if bv > 2.00: bv = 2.00
r = 0.0
g = 0.0
b = 0.0
if -0.40 <= bv<0.00:
t=(bv+0.40)/(0.00+0.40)
r=0.61+(0.11*t)+(0.1*t*t)
elif 0.00 <= bv<0.40:
t=(bv-0.00)/(0.40-0.00)
r=0.83+(0.17*t)
elif 0.40 <= bv<2.10:
t=(bv-0.40)/(2.10-0.40)
r=1.00
if -0.40 <= bv<0.00:
t=(bv+0.40)/(0.00+0.40)
g=0.70+(0.07*t)+(0.1*t*t)
elif 0.00 <= bv<0.40:
t=(bv-0.00)/(0.40-0.00)
g=0.87+(0.11*t)
elif 0.40 <= bv<1.60:
t=(bv-0.40)/(1.60-0.40)
g=0.98-(0.16*t)
elif 1.60 <= bv<2.00:
t=(bv-1.60)/(2.00-1.60)
g=0.82-(0.5*t*t)
if -0.40 <= bv<0.40:
t=(bv+0.40)/(0.40+0.40)
b=1.00
elif 0.40 <= bv<1.50:
t=(bv-0.40)/(1.50-0.40)
b=1.00-(0.47*t)+(0.1*t*t)
elif 1.50 <= bv<1.94:
t=(bv-1.50)/(1.94-1.50)
b=0.63-(0.6*t*t)
return (r, g, b)
def parse_color_map():
color_map = {}
# Compile regex
colormap_re = re.compile(r'''
([A-Z]{1,2}) # Begins with one or two letters
(\d+\.?\d*)? # Integer or float subclass
\(?
(I|II|III|IV|V)? # Luminosity class, optional
\)?
\s+[0-9.]+\s+[0-9.]+\s+
(\d{3})\s+ # Red
(\d{3})\s+ # Green
(\d{3}) # Blue
''', re.VERBOSE)
# Read in Mitchell Charity's D58 colormap
with open('starcolorsD58.txt') as fp:
for line in fp:
# Skip comments and blank lines
if line.startswith("#") or not line.strip():
#print(line)
continue
#print(line)
spectral_type, subclass, luminosity, r, g, b = colormap_re.findall(line.strip())[0]
#print(colormap_re.findall(line.strip())[0])
# we know the structure of this file, so laziness ensues
#if not subclass == "":
color_map[spectral_type+str(subclass)+luminosity] = (int(r), int(g), int(b))
#else:
# color_map[spectral_type] = (r,g,b)
return color_map
def spectral_class_to_rgb(row, spectral_class, color_map, flux_b, flux_v):
# This is really just an ugly hack for stars which are missing
# B & V magnitude data. Again, just for display purposes.
# Decoding SIMBAD spectral types
# http://simbad.u-strasbg.fr/Pages/guide/chD.htx
# Really rough parsing of spectral classification
# we ignore everything but spectral type, subclass and luminosity class
#print(spectral_class)
# If spectral class is missing
if spectral_class == "":
# Make a rough estimate of color using B-V index
if flux_b and flux_v:
rgb = bv2rgb(flux_b - flux_v)
size = 0
return rgb, size
# Use a random placeholder
spectral_class = 'X0V'
spectral_re = re.compile(r'''
([A-Z]{1,2}) # Begins with one or two letters
(\d+\.?\d*)? # Integer or float subclass
(?:[\+])? # Don't know what to do with this yet :)
(unknown|Ia|Ia\-O|Ia\-O\/Ia|Iab\-b|Ia\/ab|Iab|II|Ib|Ib\-II|III\/IV|II\/III|III|V|IV|IV\/V|V\/VI|VI)? # Luminosity class, optional
''', re.VERBOSE)
try:
spectral_type, subclass, luminosity = spectral_re.findall(spectral_class)[0]
except Exception as e:
print("Regex parsing failed on [{}] with error [{}]".format(spectral_class, e))
spectral_type = 'X'
subclass = '0'
luminosity = 'V'
# Simplify subclass to int
if subclass == "":
pass
#subclass = 0
else:
subclass = int(float(subclass))
# Simplify luminosity class to I,II,III,IV,V
if luminosity == "" or luminosity == "unknown":
if spectral_type.startswith("D"):
# Luminosity is represented by "D", leave blank
# Also, simplify stellar class
spectral_type = "D"
luminosity = ""
size = 0
if spectral_type.startswith("L") or spectral_type.startswith("T") or spectral_type.startswith("Y"):
# Brown dwarf
# Not going to even try on this one, these classifications
# are still being developed
# Set color to brown
size = 0
rgb = (150/255.0,75/255.0,0/255.0)
return rgb, size
else:
luminosity = ""
size = 0
elif luminosity.startswith("III"):
luminosity = "III"
size = 4
elif luminosity.startswith("II"):
luminosity = "II"
size = 6
elif luminosity.startswith("Ia") or luminosity.startswith("Ib"):
luminosity = "I"
size = 8
elif luminosity.startswith("IV"):
luminosity = "IV"
size = 2
elif luminosity.startswith("V"):
luminosity = "V"
size = 0
simplified_sptype = spectral_type + str(subclass) + luminosity
rgb = color_map.get(simplified_sptype, False)
if rgb:
# Color retrieved successfully
# convert from [0-255] to [0-1] range for plotting
rgb = [x / 255.0 for x in rgb]
return rgb, size
else:
# Make a rough estimate of color using B-V index
if flux_b and flux_v:
rgb = bv2rgb(flux_b - flux_v)
size = 0
return rgb, size
elif spectral_type:
# Simplify spectral type and try to fallback to color_map
spectral_type = spectral_type[0]
size = 0
# Try lookup again
rgb = color_map.get(spectral_type, False)
if rgb:
# Color retrieved successfully
# convert from [0-255] to [0-1] range for plotting
rgb = [x / 255.0 for x in rgb]
return rgb, size
else:
# purple :)
pprint_color(result['MAIN_ID', 'FLUX_B', 'FLUX_V', 'SP_TYPE'][row])
rgb = (.5,0,.5)
return rgb, size
else:
# Sigh.. Purple star it is :)
#rgb = (128,0,128)
#pprint_color(result[row])
pprint_color(result['MAIN_ID', 'FLUX_B', 'FLUX_V', 'SP_TYPE'][row])
#print("Failed spectral class [{}]".format(spectral_class))
rgb = (.5,0,.5)
return rgb, size
# In[12]:
from astropy import units as u
from astropy.coordinates import SkyCoord, Distance, search_around_3d
import numpy.ma as ma
import networkx as nx
import numpy as np
from numpy import genfromtxt
# Get dict containing stellar classification to blackbody color approximation in RGB
color_map = parse_color_map()
# Find stars which are associated (binarys, etc..) and only keep the brightest one
# I can't figure out a way to do this using a SIMBAD query yet.
# TODO: Some stars are right at the 0.1 parsec threshold likely due to measurement errors
# Example(SCR J1845-6357A and SCR J1845-6357B) a red dwarf & brown dwarf pairing
# It would be neat to take into account the measurement error data included in SIMBAD
# when estimating the distance threshold. For now, I'm just going to bump this up to 0.15pc
# distance in parsecs for identifying associations
# for reference, proxima <-> alpha centauri ~= .063pc
distance_threshold = .15*u.pc
# Example coordinate conversion using proper motion and radial velocity
# https://docs.astropy.org/en/stable/generated/examples/coordinates/plot_galactocentric-frame.html
#c1 = coord.ICRS(ra=89.014303*u.degree, dec=13.924912*u.degree,
# distance=(37.59*u.mas).to(u.pc, u.parallax()),
# pm_ra_cosdec=372.72*u.mas/u.yr,
# pm_dec=-483.69*u.mas/u.yr,
# radial_velocity=0.37*u.km/u.s)
# convert coordinates in a vectorized form (much faster than just looping over them)
result['sky_coord'] = SkyCoord(result['RA_d__ICRS__'],
result['DEC_d__ICRS__'],
distance=result['PLX_VALUE'].to(u.parsec, equivalencies=u.parallax()),
frame='icrs')
idx1, idx2, sep2d, dist3d = search_around_3d(result['sky_coord'], result['sky_coord'], distance_threshold)
# Do some numpy masking magic, we want to eliminate/mask the indices
# where a star's coordinates were compared to itself. These will also show a zero
# distance. We mask those array indexes which have the same value in both arrays.
# We apply this mask to the first array, which should contain all the indices we
# want to filter out from the main result.
#https://docs.scipy.org/doc/numpy/reference/generated/numpy.ma.masked_where.html
# Compress the results to get rid of "masked" fields for further processing
idx1_pairs = ma.masked_where(idx1 == idx2, idx1).compressed()
idx2_pairs = ma.masked_where(idx1 == idx2, idx2).compressed()
# Perform our first order grouping, values from the same index
# in each array are combined in a new array with sub arrays of
# pairs
first_grouping = [list(x) for x in zip(idx1_pairs,idx2_pairs)]
#pprint_color(first_grouping)
# Create a graph for pairs which are also associated with other pairs
# https://stackoverflow.com/questions/53886120/combine-lists-with-common-elements/53886179#53886179
graph = nx.Graph()
graph.add_edges_from(first_grouping)
# Produce a list of sets containing all association member stars (indexes)
groups = list(nx.connected_components(graph))
# Now process our stellar association groups to find the brightest stars
# skip_list contains indices of all of the dimmer members of each
# association
skip_list = []
for gset in groups:
group = list(gset)
# Try comparing Johnson V mag first
# then fall back to Blue if no V magnitudes available in group
# TODO: Is there a more comprehensive measurement available in
# SIMBAD which could be used for comparison?
min_v_mag_idx = np.argmin(result['FLUX_V'][group])
if np.isnan(np.argmin(result['FLUX_V'][group[min_v_mag_idx]])):
# Try comparing B mags
min_b_mag_idx = np.argmin(result['FLUX_B'][group])
# If B mag min is also NaN, just take it regardless...
del group[min_b_mag_idx]
skip_list.extend(group)
else:
# Remove brightest star from group and
# add all others to skip list
del group[min_v_mag_idx]
skip_list.extend(group)
#print("Fast skiplist: ",skip_list)
### Remove dim association members from results
result.remove_rows(skip_list)
plot_stars = []
for row in range(len(result)):
# Get approximate star color in RGB for plotting
rgb, size = spectral_class_to_rgb(row, result['SP_TYPE'][row].decode(),
color_map,
result['FLUX_B'][row],
result['FLUX_V'][row])
c = result['sky_coord'][row]
subgroup = result['subgroup'][row]
# Clean up label and add leading space (ugly plotting hack)
label_re = re.compile('^(\w*\*|NAME)\s+')
label = ' ' + label_re.sub('', result['MAIN_ID'][row].decode())
plot_star = [label,
float(c.cartesian.x/u.pc),
float(c.cartesian.y/u.pc),
float(c.cartesian.z/u.pc),
rgb,
size,
subgroup]
plot_stars.append(plot_star)
# In[ ]:
#pprint_color(plot_stars)
#for plot_star in plot_stars:
# if plot_star[4] == (0.5, 0, 0.5):
# pprint_color(plot_star)
# In[13]:
#https://matplotlib.org/examples/mplot3d/text3d_demo.html
labels = [' Sun']
xs = [0]
ys = [0]
zs = [0]
rgbs = [[244/255.0,248/255.0,255/255.0]]
sizes = [0]
# LCC
lcc_labels = []
lccxs = []
lccys = []
lcczs = []
lcc_rgbs = []
lcc_sizes = []
# UCL
ucl_labels = []
uclxs = []
uclys = []
uclzs = []
ucl_rgbs = []
ucl_sizes = []
#subgroup = ['']
for row in plot_stars:
if row[6] == 'UCL':
ucl_labels.append(row[0])
uclxs.append(row[1])
uclys.append(row[2])
uclzs.append(row[3])
ucl_rgbs.append(row[4])
ucl_sizes.append(row[5])
elif row[6] == 'LCC':
lcc_labels.append(row[0])
lccxs.append(row[1])
lccys.append(row[2])
lcczs.append(row[3])
lcc_rgbs.append(row[4])
lcc_sizes.append(row[5])
else:
labels.append(row[0])
xs.append(row[1])
ys.append(row[2])
zs.append(row[3])
# Convert int [0-255] to RGB float [0-1]
#rgbs.append([x / 255.0 for x in row[4]])
rgbs.append(row[4])
sizes.append(row[5])
# In[15]:
# For plotting locally (not using a jupyter notebook)
from matplotlib.backends.qt_compat import QtCore, QtWidgets, is_pyqt5
if is_pyqt5():
from matplotlib.backends.backend_qt5agg import (
FigureCanvas, NavigationToolbar2QT as NavigationToolbar)
else:
from matplotlib.backends.backend_qt4agg import (
FigureCanvas, NavigationToolbar2QT as NavigationToolbar)
from mpl_toolkits.mplot3d import Axes3D
import matplotlib.pyplot as plt
import matplotlib.animation as animation
from astropy.visualization import quantity_support
quantity_support()
# In[16]:
# Read in data on historical supernova stars
types = ['int_','float','S10','float','float','int_','int_','int_','int_','float','float','S10','S10','S10']
sn_stars = genfromtxt('supernova_table.csv', dtype=types, delimiter=',', names=True, encoding='utf-8')
# Create astropy table
sn_table = Table(sn_stars)
# Set appropriate units
#sn_table['time_sn'].unit
sn_table['x'].unit = u.pc
sn_table['y'].unit = u.pc
sn_table['z'].unit = u.pc
sn_table['distance'].unit = u.pc
pprint_color(sn_stars)
# Generate coordinates
# convert coordinates in a vectorized form (much faster than just looping over them)
sn_table['sky_coord'] = SkyCoord(x=sn_table['x'],
y=sn_table['y'],
z=sn_table['z'],
frame='icrs',
representation_type='cartesian')
# Group by stellar subgroup
sn_subgroup = sn_table.group_by('subgroup')
lcc_sn_subgroup, ucl_sn_subgroup = sn_subgroup.groups
#lcc_sn_subgroup
#pprint_color(sn_table)
#pprint_color(lcc_sn_subgroup)
#print(rgbs)
# Use dark background for better star visibility
plt.style.use('dark_background')
# Faint grid color for visibility
plt.rcParams['grid.color'] = (0.2,0.2,0.2)
#fig=plt.figure(figsize=(4,35))
#fig = plt.figure(figsize=(10,10))
fig = plt.figure(figsize=(19,19))
#fig = plt.figure(dpi=100)
ax = fig.gca(projection='3d')
#fig, ax = plt.subplots()
#plt3d = plt.figure().gca(projection='3d')
#plt3d = ax.figure()
# Apply stellar name labels
# https://matplotlib.org/api/_as_gen/matplotlib.axes.Axes.text.html
for label, x, y, z in zip(labels, xs, ys, zs):
ax.text(x,y,z,label,None)
# ax.text(x,y,z,label,fontsize=6)
# Plot general stars
ax.scatter(xs, ys, zs, marker='o', linewidth=4, c=rgbs)
# Current LCC region
ax.scatter(lccxs, lccys, lcczs, marker='o', linewidth=lcc_sizes, c='blue', alpha=1, label="Present Day LCC Subgroup")
#ax.scatter(lccxs, lccys, lcczs, marker='+', linewidth=lcc_sizes, edgecolors=lcc_rgbs, c=lcc_rgbs, alpha=1, label="Present Day LCC Subgroup")
# Current UCL region
ax.scatter(uclxs, uclys, uclzs, marker='o', linewidth=ucl_sizes, c='orange', alpha=1, label="Present Day UCL Subgroup")
#ax.scatter(uclxs, uclys, uclzs, marker='s', linewidth=ucl_sizes, edgecolors=ucl_rgbs, c=ucl_rgbs, alpha=1, label="Present Day UCL Subgroup")
#ax.scatter(xs, ys, zs, 'o', linewidth=sizes, edgecolors='white', c=rgbs, alpha=1)
# Chart LCC supernovae subgroup
ax.scatter(lcc_sn_subgroup['z'],
lcc_sn_subgroup['x'],
lcc_sn_subgroup['y'], marker='D', linewidth=1, c='green', alpha=1, label='LCC Supernovae')
# Plot supernovae time in past as label
for label, x, y, z in zip(lcc_sn_subgroup['time_sn'], lcc_sn_subgroup['x'], lcc_sn_subgroup['y'], lcc_sn_subgroup['z']):
ax.text(x,y,z," "+str(label),None,fontsize=8)
#ax.text(x,y,z," "+str(label),(1,1,0),fontsize=8)
# Chart UCL supernovae subgroup
ax.scatter(ucl_sn_subgroup['z'],
ucl_sn_subgroup['x'],
ucl_sn_subgroup['y'], marker='^', linewidth=1, c='purple', alpha=1, label='UCL Supernovae')
# Plot supernovae time in past as label
for label, x, y, z in zip(ucl_sn_subgroup['time_sn'], ucl_sn_subgroup['x'], ucl_sn_subgroup['y'], ucl_sn_subgroup['z']):
#ax.text(x,y,z," "+str(label),(1,-1,0),fontsize=8)
ax.text(x,y,z," "+str(label),None,fontsize=8)
# Tweaking display region and labels
# 3D plot radius from Sun (origin)
#lim = 20*u.pc
lim = 300*u.pc
ax.set_xlim(-lim/2, lim)
ax.set_ylim(-lim, lim)
ax.set_zlim(-lim, lim)
ax.set_xlabel('X (parsec)')
ax.set_ylabel('Y (parsec)')
ax.set_zlabel('Z (parsec)')
# Show galactic plane
# adapted from: https://stackoverflow.com/questions/3461869/plot-a-plane-based-on-a-normal-vector-and-a-point-in-matlab-or-matplotlib
# We'll create two points in the galactic coordinate system, one at the origin (Sun) and one normal
# to the origin at latitude +90deg
#gal_origin = SkyCoord(l=0*u.deg, b=0*u.deg, distance=0*u.pc, frame='galactic')
#gal_normal = SkyCoord(l=0*u.deg, b=90*u.deg, distance=1*u.pc, frame='galactic')
# Or using galactic coordinates (UVW) in a cartesian form
gal_origin = SkyCoord(u=0, v=0, w=0, frame='galactic', representation_type='cartesian')
gal_normal = SkyCoord(u=0, v=0, w=1, frame='galactic', representation_type='cartesian')
# Convert these coordinates to their equivalents into our ICRS reference frame
icrs_origin_sky = gal_origin.transform_to('icrs')
icrs_normal_sky = gal_normal.transform_to('icrs')
# Switch to cartesian representation of these coordinates
icrs_origin_sky.representation_type = 'cartesian'
icrs_normal_sky.representation_type = 'cartesian'
# Numpy array
# Try without intermediate step
icrs_origin = np.array([icrs_origin_sky.x, icrs_origin_sky.y, icrs_origin_sky.z])
icrs_normal = np.array([icrs_normal_sky.x, icrs_normal_sky.y, icrs_normal_sky.z])
pprint_color(icrs_origin)
pprint_color(icrs_normal)
# Compute normal
d = -icrs_origin.dot(icrs_normal)
# Create xy values
xx, yy = np.meshgrid(range(-300,300), range(-300,300))
#xx, yy = np.meshgrid(range(lim/u.pc), range(lim/u.pc))
# Calculate corresponding z
z = (-icrs_normal[0] * xx - icrs_normal[1] * yy - d) * 1. / icrs_normal[2]
# Plot the galactic plane
surf = ax.plot_surface(xx, yy, z, shade=False, color='grey', alpha=0.2, linewidth=0, label='Galactic Plane')
# Fix plotting bug
# https://stackoverflow.com/questions/54994600/pyplot-legend-poly3dcollection-object-has-no-attribute-edgecolors2d
surf._facecolors2d=surf._facecolors3d
surf._edgecolors2d=surf._edgecolors3d
#ax.text(0, -300, 0, "Galactic Plane", None, fontsize=6)
#ax.plot_surface(xx, yy, z)
#plt3d.plot_surface(xx, yy, z)
#ax.w_xaxis.grid(color='r', linestyle='-', linewidth=2)
#ax.grid(alpha=.1)
# Darken axis panes so colored stars show more clearly
ax.xaxis.pane.set_color((0, 0, 0, 1.0))
ax.yaxis.pane.set_color((0, 0, 0, 1.0))
ax.zaxis.pane.set_color((0, 0, 0, 1.0))
#ax.xaxis.pane.set_color((.8, .8, .8, 1.0))
#ax.yaxis.pane.set_color((.8, .8, .8, 1.0))
#ax.zaxis.pane.set_color((.8, .8, .8, 1.0))
#ax.w_xaxis.set_pane_color((0, 0, 0, 1.0))
#ax.w_yaxis.set_pane_color((0, 0, 0, 1.0))
#ax.w_zaxis.set_pane_color((0, 0, 0, 1.0))
ax.legend()
# apply a simple rotation animation
#for angle in range(0, 360):
# ax.view_init(30, angle)
# plt.draw()
# plt.pause(.001)
fig.tight_layout()
#plt.savefig('stellar_neighborhood.png')
plt.show()
# In[ ]:
| research_project/figs/stellar_plot_3d/Supernova Plot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 7.6 List vs. `array` Performance: Introducing `%timeit`
# ### Timing the Creation of a List Containing Results of 6,000,000 Die Rolls
import random
# %timeit rolls_list = \
# [random.randrange(1, 7) for i in range(0, 6_000_000)]
# ### Timing the Creation of an `array` Containing Results of 6,000,000 Die Rolls
import numpy as np
# %timeit rolls_array = np.random.randint(1, 7, 6_000_000)
# ### 60,000,000 and 600,000,000 Die Rolls
# %timeit rolls_array = np.random.randint(1, 7, 60_000_000)
# %timeit rolls_array = np.random.randint(1, 7, 600_000_000)
# ### Customizing the %timeit Iterations
# %timeit -n3 -r2 rolls_array = np.random.randint(1, 7, 6_000_000)
# ### Other IPython Magics
##########################################################################
# (C) Copyright 2019 by Deitel & Associates, Inc. and #
# Pearson Education, Inc. All Rights Reserved. #
# #
# DISCLAIMER: The authors and publisher of this book have used their #
# best efforts in preparing the book. These efforts include the #
# development, research, and testing of the theories and programs #
# to determine their effectiveness. The authors and publisher make #
# no warranty of any kind, expressed or implied, with regard to these #
# programs or to the documentation contained in these books. The authors #
# and publisher shall not be liable in any event for incidental or #
# consequential damages in connection with, or arising out of, the #
# furnishing, performance, or use of these programs. #
##########################################################################
| examples/ch07/snippets_ipynb/07_06.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Train the model
# + pycharm={"name": "#%%\n"}
import os
import datetime
import tensorflow as tf
import tensorflow_datasets as tfds
# -
# ## Prepare training data
# + pycharm={"name": "#%%\n"}
tfds.disable_progress_bar()
# -
# ### Load images from the "cats vs dogs" dataset
# + pycharm={"name": "#%%\n"}
(raw_train, raw_validation, raw_test), metadata = tfds.load(
'cats_vs_dogs',
split=['train[:80%]', 'train[80%:90%]', 'train[90%:]'],
with_info=True,
as_supervised=True,
)
# -
# ### Fit images to network inputs
# + pycharm={"name": "#%%\n"}
IMG_SIZE = 160 # All images will be resized to 160x160
def format_example(image, label):
image = tf.cast(image, tf.float32)
image = (image/127.5) - 1
image = tf.image.resize(image, (IMG_SIZE, IMG_SIZE))
return image, label
train = raw_train.map(format_example)
validation = raw_validation.map(format_example)
test = raw_test.map(format_example)
# -
# ### Shuffle training data
# + pycharm={"name": "#%%\n"}
BATCH_SIZE = 32
SHUFFLE_BUFFER_SIZE = 1000
train_batches = train.shuffle(SHUFFLE_BUFFER_SIZE).batch(BATCH_SIZE)
validation_batches = validation.batch(BATCH_SIZE)
test_batches = test.batch(BATCH_SIZE)
# -
# ## Load the untrained model
# + pycharm={"name": "#%%\n"}
model = tf.keras.models.load_model('saved_model/mobilenetv2-untrained')
# + pycharm={"name": "#%% Configure only the top of the model as trainable\n"}
is_trainable = False
for layer in model.layers:
if layer.name == 'top_start':
is_trainable = True
layer.trainable = is_trainable
# -
# ## Compile the model
# + pycharm={"name": "#%%\n"}
base_learning_rate = 0.0001
model.compile(optimizer=tf.keras.optimizers.RMSprop(lr=base_learning_rate),
loss=tf.keras.losses.BinaryCrossentropy(from_logits=True),
metrics=['accuracy'])
# -
# ## Train the top of the model
# Progress can be visualized using the command:
# ```
# tensorboard --logdir logs/scalars
# ```
# + pycharm={"name": "#%%\n"}
initial_epochs = 10
logdir = "logs/scalars/" + datetime.datetime.now().strftime("%Y%m%d-%H%M%S")
tensorboard_callback = tf.keras.callbacks.TensorBoard(log_dir=logdir)
history = model.fit(train_batches,
epochs=initial_epochs,
validation_data=validation_batches,
callbacks=[tensorboard_callback])
# -
# ## Fine tune the last layers of the network
# + pycharm={"name": "#%%\n"}
n_fine_tune_layers = 30
for layer in model.layers[:-n_fine_tune_layers]:
layer.trainable = False
for layer in model.layers[-n_fine_tune_layers:]:
layer.trainable = True
model.compile(optimizer = tf.keras.optimizers.RMSprop(lr=base_learning_rate/10),
loss=tf.keras.losses.BinaryCrossentropy(from_logits=True),
metrics=['accuracy'])
fine_tune_epochs = 10
total_epochs = initial_epochs + fine_tune_epochs
history_fine = model.fit(train_batches,
epochs=total_epochs,
initial_epoch=history.epoch[-1],
validation_data=validation_batches,
callbacks=[tensorboard_callback])
# -
# ## Save the model
# + pycharm={"name": "#%%\n"}
os.makedirs('saved_model', exist_ok=True)
model.save('saved_model/mobilenetv2')
| 02_train.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
os.environ['CUDA_VISIBLE_DEVICES'] = '3'
os.environ['GOOGLE_APPLICATION_CREDENTIALS'] = '/home/husein/t5/prepare/mesolitica-tpu.json'
import malaya_speech.train.model.conformer as conformer
import malaya_speech.train.model.transducer as transducer
import malaya_speech
import tensorflow as tf
import numpy as np
import json
from glob import glob
import pandas as pd
subwords = malaya_speech.subword.load('transducer-singlish.subword')
featurizer = malaya_speech.tf_featurization.STTFeaturizer(
normalize_per_feature = True
)
# +
n_mels = 80
sr = 16000
maxlen = 18
minlen_text = 1
def mp3_to_wav(file, sr = sr):
audio = AudioSegment.from_file(file)
audio = audio.set_frame_rate(sr).set_channels(1)
sample = np.array(audio.get_array_of_samples())
return malaya_speech.astype.int_to_float(sample), sr
def generate(file):
print(file)
with open(file) as fopen:
audios = json.load(fopen)
for i in range(len(audios)):
try:
audio = audios[i][0]
wav_data, _ = malaya_speech.load(audio, sr = sr)
if (len(wav_data) / sr) > maxlen:
# print(f'skipped audio too long {audios[i]}')
continue
if len(audios[i][1]) < minlen_text:
# print(f'skipped text too short {audios[i]}')
continue
t = malaya_speech.subword.encode(
subwords, audios[i][1], add_blank = False
)
back = np.zeros(shape=(2000,))
front = np.zeros(shape=(200,))
wav_data = np.concatenate([front, wav_data, back], axis=-1)
yield {
'waveforms': wav_data,
'targets': t,
'targets_length': [len(t)],
}
except Exception as e:
print(e)
def preprocess_inputs(example):
s = featurizer.vectorize(example['waveforms'])
mel_fbanks = tf.reshape(s, (-1, n_mels))
length = tf.cast(tf.shape(mel_fbanks)[0], tf.int32)
length = tf.expand_dims(length, 0)
example['inputs'] = mel_fbanks
example['inputs_length'] = length
example.pop('waveforms', None)
example['targets'] = tf.cast(example['targets'], tf.int32)
example['targets_length'] = tf.cast(example['targets_length'], tf.int32)
return example
def get_dataset(
file,
batch_size = 3,
shuffle_size = 20,
thread_count = 24,
maxlen_feature = 1800,
):
def get():
dataset = tf.data.Dataset.from_generator(
generate,
{
'waveforms': tf.float32,
'targets': tf.int32,
'targets_length': tf.int32,
},
output_shapes = {
'waveforms': tf.TensorShape([None]),
'targets': tf.TensorShape([None]),
'targets_length': tf.TensorShape([None]),
},
args = (file,),
)
dataset = dataset.prefetch(tf.contrib.data.AUTOTUNE)
dataset = dataset.map(
preprocess_inputs, num_parallel_calls = thread_count
)
dataset = dataset.padded_batch(
batch_size,
padded_shapes = {
'inputs': tf.TensorShape([None, n_mels]),
'inputs_length': tf.TensorShape([None]),
'targets': tf.TensorShape([None]),
'targets_length': tf.TensorShape([None]),
},
padding_values = {
'inputs': tf.constant(0, dtype = tf.float32),
'inputs_length': tf.constant(0, dtype = tf.int32),
'targets': tf.constant(0, dtype = tf.int32),
'targets_length': tf.constant(0, dtype = tf.int32),
},
)
return dataset
return get
# -
dev_dataset = get_dataset('test-set-imda.json', batch_size = 3)()
features = dev_dataset.make_one_shot_iterator().get_next()
features
training = True
# +
config = malaya_speech.config.conformer_large_encoder_config
config['dropout'] = 0.0
conformer_model = conformer.Model(
kernel_regularizer = None, bias_regularizer = None, **config
)
decoder_config = malaya_speech.config.conformer_large_decoder_config
decoder_config['embed_dropout'] = 0.0
transducer_model = transducer.rnn.Model(
conformer_model, vocabulary_size = subwords.vocab_size, **decoder_config
)
targets_length = features['targets_length'][:, 0]
v = tf.expand_dims(features['inputs'], -1)
z = tf.zeros((tf.shape(features['targets'])[0], 1), dtype = tf.int32)
c = tf.concat([z, features['targets']], axis = 1)
logits = transducer_model([v, c, targets_length + 1], training = training)
# -
decoded = transducer_model.greedy_decoder(v, features['inputs_length'][:, 0], training = training)
decoded
sess = tf.Session()
sess.run(tf.global_variables_initializer())
var_list = tf.get_collection(tf.GraphKeys.GLOBAL_VARIABLES)
saver = tf.train.Saver(var_list = var_list)
saver.restore(sess, 'asr-large-conformer-transducer-singlish/model.ckpt-925000')
wer, cer = [], []
index = 0
while True:
try:
r = sess.run([decoded, features['targets']])
for no, row in enumerate(r[0]):
d = malaya_speech.subword.decode(subwords, row[row > 0])
t = malaya_speech.subword.decode(subwords, r[1][no])
wer.append(malaya_speech.metrics.calculate_wer(t, d))
cer.append(malaya_speech.metrics.calculate_cer(t, d))
index += 1
except Exception as e:
break
np.mean(wer), np.mean(cer)
for no, row in enumerate(r[0]):
d = malaya_speech.subword.decode(subwords, row[row > 0])
t = malaya_speech.subword.decode(subwords, r[1][no])
print(no, d)
print(t)
print()
| pretrained-model/stt/conformer/evaluate/large-singlish.ipynb |
// -*- coding: utf-8 -*-
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .cpp
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: C++17
// language: C++17
// name: xcpp17
// ---
// +
#include <algoviz/SVG.hpp>
AlgoViz::clear();
SVG zeichnung = SVG(400,400,"Die Tardis im Zeitstrudel");
Image tardis = Image("/user-redirect/algoviz/img/tardis.png",10,10,30,30,&zeichnung);
int x = 200;
int y = 100;
int vx = 5;
int vy = 0;
int ax = 0;
int ay = 1;
int winkel = 0;
std::string key;
do {
// Ermittle die zuletzt gedrückte Taste
key = zeichnung.lastKey();
if ( key == "ArrowLeft") {
vx = vx - 1;
} else if ( key == "ArrowRight") {
vx = vx + 1;
} else if ( key == "ArrowUp") {
vy = vy - 1;
} else if ( key == "ArrowDown") {
vy = vy + 1;
} else if ( key == "0") {
vy = 0;
vx = 0;
}
// Beschleunige die Tardis
vy = vy + (200-y)/50;
vx = vx + (200-x)/50;
if ( key == "0" ) {
vx = 0;
vy = 0;
}
// Verändere die Position der Tardis
x = x + vx;
y = y + vy;
// Lasse sie am Rand reflektieren
// Der linke Rand
if ( x < 10 ) {
x = 20-x;
vx = -vx;
}
// Der rechte Rand
if ( x > 389 ) {
x = 778-x;
vx = -vx;
}
// Der obere Rand
if ( y < 10 ) {
y = 20 - y;
vy = -vy;
}
// Der untere Rand
if ( y > 389 ) {
y = 778-y;
vy = -vy;
}
winkel = (winkel+10) % 360;
tardis.rotateTo(winkel);
tardis.moveTo(x,y);
AlgoViz::sleep(10);
} while ( key != "x" );
// -
AlgoViz::clear();
| lessons/02_Grundlagen/.XX_Tardis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Environment (conda_pytorch_p27)
# language: python
# name: conda_pytorch_p27
# ---
% matplotlib notebook
from my_utils import *
import numpy as np
import pandas
import torch, torchvision
import os
# +
full_model = torch.load('./full_model_gpu.pt')
# define the device
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
full_model.to(device);
# -
filenames = os.listdir('./data/test/')
# +
MEAN = [0.485, 0.456, 0.406] # expected by pretrained resnet18
STD = [0.229, 0.224, 0.225] # expected by pretrained resnet18
# define transformations without data augmentation
transforms = torchvision.transforms.Compose([
torchvision.transforms.Resize((224,224)),
torchvision.transforms.ToTensor(), # Expected by pretrained neural network
torchvision.transforms.Normalize(MEAN, STD) # Expected by pretrained neural network
])
# -
test_df = pandas.DataFrame({'Id':filenames})
test_data = TestDataset(test_df,'./data/test/',transform=transforms)
# load the trainind data to get a list of the categories
train_df = pandas.read_csv('./data/train.csv')
train_data = WhaleDataset(train_df,'./data/',transform=None)
categories = train_data.categories
test_dataloader = torch.utils.data.DataLoader(test_data,\
batch_size=4,\
num_workers=4,\
shuffle=False,\
sampler=None)
result = predict(full_model,test_dataloader,device,categories)
result.to_csv('./my_submission_gpu.csv',index=False)
| predict.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + active=""
# Firstly, Check existing packages with the following:
# -
# !python -m pip list
# + active=""
# Secondly, install nbextension to Jupyter Notebook according the following link:
# https://github.com/ipython-contrib/jupyter_contrib_nbextensions
# -
# !python -m pip install jupyter_contrib_nbextensions
# + active=""
# Thirdly, install necessary packages accordingly:
# -
# update pip if necessary
# !python -m pip install --upgrade pip
# install paramiko to access WSL and HPC
# !python -m pip install paramiko==2.7.1
# install pysftp to access HPC
# !python -m pip install pysftp==0.2.8
# install smt for LHS sampling
# !python -m pip install smt
| jupyter/Python_ROM_GUI/InstallPackages.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Simulation to test cost functions for probabilistic path finding
#
# This notebook is a repsonse to https://github.com/ElementsProject/lightning/pull/4771 where <NAME>, <NAME> and I discuss which cost function should be used in Sijkstra computations to test single paths if one wants to incorporate the results from proababilistic path finding research (c.f: https://arxiv.org/abs/2103.08576 )
#
# ## Funding
# This work is funded by NTNU, Bitmex, and independent people tipping me via https://donate.ln.rene-pickhardt.de and via https://patreon.com/renepickhardt
import json
from math import log2
import networkx as nx
import numpy as np
import matplotlib.pyplot as plt
import random
# ## A few helper functions
# +
def import_channels(filename):
f= open(filename,"r")
channels = json.load(f)["channels"]
return channels
def next_hop(path):
for i in range(1,len(path)):
src = path[i-1]
dest = path[i]
yield (src,dest)
def arithmetic_mean(features):
return np.mean(features)
def harmonic_mean(features):
return len(features)/sum(1.0/(f+1) for f in features)
# -
# ## Define cost functions for single features that are potentially used in c-lighting
# +
def fee_function(amt,channel):
return channel["base_fee_millisatoshi"]+amt*channel["fee_per_millionth"]/1000000
def risk_factor_cltv(riskfactor,channel):
cltv_delay = channel["delay"]
BLOCKS_PER_YEAR = 144*365
return riskfactor / 100.0 / BLOCKS_PER_YEAR * cltv_delay
def neg_log_probabilities(amt, channel):
cap = channel["satoshis"]*1000
return -log2((cap+1-amt)/(cap+1))
def linearized_success_probabilities(amt,channel):
cap = channel["satoshis"]*1000
return amt/(cap+1)
# -
# ## Actual cost functions with combination of features
# +
def build_features(amt, channel, risk_factor=0):
features = {
"fees":fee_function(amt,channel),
"neg_log_prob":neg_log_probabilities(amt, channel)}
if risk_factor>0:
features["risk_factor"]= risk_factor_cltv(risk_factor,channel)
return features
def harmonic_probabilistic_cost(amt,channel,risk_factor=0):
features = build_features(amt,channel,risk_factor)
return harmonic_mean(features.values())
def arithmetic_probabilistic_cost(amt,channel,risk_factor=0):
features = build_features(amt,channel,risk_factor)
return arithmetic_mean(list(features.values()))
def rusty_proposed_cost(amt,channel,risk_factor=0):
f = fee_function(amt,channel)
p = linearized_success_probabilities(amt,channel)
r = risk_factor_cltv(risk_factor,channel)
return f*(1+p)+r
# -
# ## Import data from lightning-cli listchannels > channels20210918.json
channels = import_channels("channels20210918.json")
channels[0]
# ## Plot Diagrams
# +
values = list(range(1,20))
plt.title("harmonic mean with cltv on 20k sats channel")
plt.plot(values,[harmonic_probabilistic_cost(amt*1000000,channels[0],1) for amt in values])
plt.grid()
plt.xlabel("amount sent in kilo Sats")
plt.ylabel("cost")
plt.show()
plt.title("harmonic mean without cltv on 20k sats channel")
plt.plot(values,[harmonic_probabilistic_cost(amt*1000000,channels[0]) for amt in values])
plt.grid()
plt.xlabel("amount sent in kilo Sats")
plt.ylabel("cost")
plt.show()
plt.title("arithmetic mean with cltv on 20k sats channel")
plt.plot(values,[arithmetic_probabilistic_cost(amt*1000000,channels[0],1) for amt in values])
plt.grid()
plt.xlabel("amount sent in kilo Sats")
plt.ylabel("cost")
plt.show()
plt.title("arithmetic mean without cltv on 20k sats channel")
plt.plot(values,[arithmetic_probabilistic_cost(amt*1000000,channels[0]) for amt in values])
plt.grid()
plt.xlabel("amount sent in kilo Sats")
plt.ylabel("cost")
plt.show()
plt.title("Rusty Proposed cost on 20k sats channel")
plt.plot(values,[rusty_proposed_cost(amt*1000000,channels[0]) for amt in values])
plt.grid()
plt.xlabel("amount sent in kilo Sats")
plt.ylabel("cost")
plt.show()
# -
# ## Create a graph
def make_known_balance_graph(channels):
D = nx.DiGraph()
for chan in channels:
D.add_edge(chan["source"],chan["destination"],channel=chan)
for s,d in D.edges():
cap = D[s][d]["channel"]["satoshis"]
if s in D[d]:
if s<d:
D[s][d]["balance"]=random.randint(0,cap)
D[d][s]["balance"]=cap - D[s][d]["balance"]
else:
D[s][d]["balance"]=random.randint(0,cap)
return D
D = make_known_balance_graph(channels)
#check visualy that it is uniformly distributed
plt.hist([D[s][d]["balance"]/D[s][d]["channel"]["satoshis"] for s,d in D.edges()])
plt.show()
def prepare_experiment(D,cost_function,amt,risk_factor = 0):
res = nx.DiGraph()
for s,d in D.edges():
if D[s][d]["channel"]["satoshis"]*1000 < amt:
continue
res.add_edge(s,d,weight=cost_function(amt, D[s][d]["channel"],risk_factor))
return res
bos_nodes = [x["public_key"] for x in json.load(open("bos.json","r"))["scores"]]
LN_RENE_PICKHARDT="03efccf2c383d7bf340da9a3f02e2c23104a0e4fe8ac1a880c8e2dc92fbdacd9df"
amt_sats = 100000
harmonic_with_cltv = prepare_experiment(D,harmonic_probabilistic_cost,1000*amt_sats,1)
harmonic_no_cltv = prepare_experiment(D,harmonic_probabilistic_cost,1000*amt_sats)
rusty = res = prepare_experiment(D,rusty_proposed_cost,1000*amt_sats,1)
# +
def can_deliver(path,D,amt):
for s,d in next_hop(path):
if D[s][d]["balance"]<amt:
return False
return True
def evaluate(graph):
lengths = {"s":[],"f":[]}
for dest in bos_nodes:
path = nx.dijkstra_path(graph,LN_RENE_PICKHARDT,dest)
if can_deliver(path,D,amt_sats):
lengths["s"].append(len(path))
else:
lengths["f"].append(len(path))
return lengths
l1 = evaluate(harmonic_with_cltv)
print("probabilistic with CLTV successrate: {:4.2f}%".format(100*len(l1["s"])/len(bos_nodes)))
l2 = evaluate(harmonic_no_cltv)
print("probabilistic no CLTV successrate: {:4.2f}%".format(100*len(l2["s"])/len(bos_nodes)))
l3 = evaluate(rusty)
print("rustyies method successrate: {:4.2f}%".format(100*len(l3["s"])/len(bos_nodes)))
# -
print(np.mean(l1["s"]), np.mean(l1["f"]))
print(np.mean(l2["s"]), np.mean(l2["f"]))
print(np.mean(l3["s"]), np.mean(l3["f"]))
| Probabilistic Pathfinding Simulation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Inaugural Project
# > **Note the following:**
# > 1. This is an example of how to structure your **inaugural project**.
# > 1. Remember the general advice on structuring and commenting your code from [lecture 5](https://numeconcopenhagen.netlify.com/lectures/Workflow_and_debugging).
# > 1. Remember this [guide](https://www.markdownguide.org/basic-syntax/) on markdown and (a bit of) latex.
# > 1. Turn on automatic numbering by clicking on the small icon on top of the table of contents in the left sidebar.
# > 1. The `inauguralproject.py` file includes a function which can be used multiple times in this notebook.
# Imports and set magics:
# +
import numpy as np
# autoreload modules when code is run
# %load_ext autoreload
# %autoreload 2
# local modules
import inauguralproject
# + [markdown] toc-hr-collapsed=true
# # Question 1
# -
# BRIEFLY EXPLAIN HOW YOU SOLVE THE MODEL.
# +
# code for solving the model (remember documentation and comments)
a = np.array([1,2,3])
b = inauguralproject.square(a)
print(b)
# -
# # Question 2
# ADD ANSWER.
# +
# code
# -
# # Question 3
# ADD ANSWER.
# +
# code
# -
# # Question 4
# ADD ANSWER.
# +
# code
# -
# # Question 5
# ADD ANSWER.
# +
# code
# -
# # Conclusion
# ADD CONCISE CONLUSION.
| inauguralproject/inauguralproject.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import requests, json
def match(conditions):
return requests.post("http://localhost:5004/match_api", json=conditions).json()
def get_annotated_data():
url = 'https://storage.gra.cloud.ovh.net/v1/AUTH_32c5d10cb0fe4519b957064a111717e3/models/match_pubmed_affiliations_with_countries_v3.json'
data = requests.get(url).json()
json.dump(data, open('match_pubmed_affiliations_with_countries_v3.json', 'w'), indent=2, ensure_ascii=False)
return data
def compute_precision_recall(match_type, index_prefix=''):
data = get_annotated_data()
nb_TP, nb_FP, nb_FN = 0, 0, 0
false_positive, false_negative = [], []
for ix, d in enumerate(data):
if ix%100==0:
print(ix, end=',')
if d.get(match_type):
res = match({'query': d['label'], 'year':'2020', 'type': match_type, 'index_prefix': index_prefix})
for x in res['results']:
if x in d[match_type]:
nb_TP += 1
else:
nb_FP += 1
false_positive.append(d)
for x in d[match_type]:
if x not in res['results']:
nb_FN += 1
false_negative.append(d)
precision = nb_TP / (nb_TP + nb_FP)
recall = nb_TP / (nb_TP + nb_FN)
res = {'precision' : precision, 'recall' : recall}
return {'res': res, 'false_positive': false_positive, 'false_negative': false_negative}
# -
metrics_country = compute_precision_recall(match_type = 'country', index_prefix='matcher')
metrics_country['res']
metrics_country['false_positive']
metrics_grid = compute_precision_recall(match_type = 'grid', index_prefix='matcher')
metrics_grid['res']
metrics_grid['false_positive']
metrics_rnsr = compute_precision_recall(match_type = 'rnsr', index_prefix='matcher')
metrics_rnsr['res']
| notebooks/evaluation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sympy as sp
from sympy import *
import numpy as np
import matplotlib.pyplot as plt
from scipy.interpolate import interp1d
from scipy.integrate import quad
from scipy.optimize import fmin
import scipy.integrate as integrate
import scipy.special as special
import scipy.stats as st
import sys
font1 = {'size' : 20, 'family':'STIXGeneral'}
from platform import python_version
print(python_version())
# +
#LCDM fractions
mptkm = 3.086*10**(19)
H0 = 67.32/mptkm
Oc = 0.265
Ob = 0.0494
Om = Oc + Ob
Orad = 0.000093
ai = 0.000001
arad=0.0002264 #radiation -DM equality
acmb = 0.0009
Gnewton = 6.67*10**(-11)
def Hub(Om, Orad, a):
return H0*np.sqrt(Om/a**3 + Orad/a**4 + (1-Om-Orad))
def rhoc(a):
return 3*Hub(Om, Orad, a)**2/(8*np.pi/Gnewton)
def Omegac(a):
return Oc/a**3*(H0/Hub(Om,Orad,a))**2
def Omegarad(a):
return Orad/a**4*(H0/Hub(Om,Orad,a))**2
def Omegab(a):
return Ob/a**3*(H0/Hub(Om,Orad,a))**2
# +
fig0 = plt.figure()
plt.figure(figsize=(10,10))
#Load Omega_pbh data at early and late times
dat1 = np.loadtxt("data/8gev+9orad/peakm_5e-7.dat")
dat2 = np.loadtxt("data/8gev+9orad/peakm_2e11.dat")
dat3 = np.loadtxt("data/8gev+9orad/peakm_5e11.dat")
dat4 = np.loadtxt("data/8gev+9orad/peakm_5e13.dat")
dat5 = np.loadtxt("data/8gev+9orad/peakm_5e33.dat")
avals = np.logspace(-6, 0, num=1000)
ax = plt.subplot(2, 1, 1)
plt.xscale('log')
plt.plot(avals, Omegac(avals),linestyle='dashed', color='b', label= '$\Omega_{\\rm cdm}$',alpha=1.)
plt.plot(dat1[:,0], dat1[:,1], label='$M_{pk} = 5\\times 10^{-7}$kg', alpha=0.6)
plt.plot(dat2[:,0], dat2[:,1], label='$M_{pk} = 2\\times 10^{11}$kg', alpha=0.6)
plt.plot(dat3[:,0], dat3[:,1], label='$M_{pk} = 5\\times 10^{11}$kg',alpha=0.6)
plt.plot(dat4[:,0], dat4[:,1], label='$M_{pk} = 5\\times 10^{13}$kg',alpha=0.6)
plt.plot(dat5[:,0], dat5[:,1], label='$M_{pk} = 5\\times 10^{33}$kg',alpha=0.6)
plt.axhline(y=1., xmin=0., xmax=10,color='k',linestyle='dashed')
plt.axvline(acmb,0.,10, color='k', linestyle='dotted')
#plt.text(2e-6, 0.3 , '$T_{\\rm RH} = 10^{8}{\\rm GeV}$', **font1)
ax.tick_params(axis='both', which='major', labelsize=15)
plt.ylim(-0.,1.45)
plt.xlim(ai,1)
plt.ylabel('Density fraction of PBH ($\Omega_{\\rm PBH}$) ',**font1)
plt.xlabel('scale factor (a)', **font1)
plt.legend(loc='best',prop={'size': 16})
plt.tight_layout(pad=3.0)
ax = plt.subplot(2, 1, 2)
plt.xscale('log')
plt.plot(avals, Omegab(avals),linestyle='dashed', color='r', label= '$\Omega_b$',alpha=1.)
plt.plot(dat1[:,0], dat1[:,2], label='$\lambda = 3.4\\times 10^{96}$', alpha=0.6)
plt.plot(dat2[:,0], dat2[:,2], label='$\lambda = 1.9\\times 10^{98}$', alpha=0.6)
plt.plot(dat3[:,0], dat3[:,2], label='$\lambda = 4.8\\times 10^{98}$', alpha=0.6)
plt.plot(dat4[:,0], dat4[:,2], label='$\lambda = 4.7\\times 10^{100}$', alpha=0.6)
plt.plot(dat5[:,0], dat5[:,2], label='$\lambda = 3.5\\times 10^{120}$', alpha=0.6)
plt.axvline(acmb,0.,10, color='k', linestyle='dotted')
ax.tick_params(axis='both', which='major', labelsize=15)
plt.ylim(0,0.2)
plt.xlim(ai,1)
plt.ylabel('Density fraction of baryons ($\Omega_{\\rm b}$) ',**font1)
plt.xlabel('scale factor (a)', **font1)
plt.legend(loc='best',prop={'size': 16})
#plt.setp(plt.subplot(2,1,1).get_xticklabels(), visible=False)
plt.subplots_adjust(hspace=0.2)
plt.subplots_adjust(wspace=0.)
plt.savefig('plots/omega_all.png', format="png", bbox_inches = 'tight')
# +
# Plotting LCDM fractions
fig0 = plt.figure()
plt.figure(figsize=(10,5))
avals = np.logspace(-6, 0, num=1000)
ax = plt.subplot(1, 1, 1)
plt.xscale('log')
plt.plot(avals, Omegac(avals),linestyle='dashed', color='b', label= '$\Omega_c$',alpha=1.)
plt.plot(avals, Omegab(avals),linestyle='dashed', color='g', label= '$\Omega_b$',alpha=1.)
plt.plot(avals, Omegarad(avals),linestyle='dashed', color='r', label= '$\Omega_\gamma$',alpha=1.)
ax.axvspan(ai, 0.000215, alpha=0.5, color='orange')
plt.axhline(y=0.6856, xmin=0., xmax=10,color='k')
ax.tick_params(axis='both', which='major', labelsize=15)
plt.ylim(-0.1,1.5)
plt.xlim(ai,1)
plt.xlabel('scale factor (a)', **font1)
plt.ylabel('Density fraction ($\Omega$) ',**font1)
plt.legend(loc='best',prop={'size': 14})
plt.tight_layout(pad=3.0)
plt.savefig('plots/lcdm_epochs.png', format="png", bbox_inches = 'tight')
# +
fig0 = plt.figure()
plt.figure(figsize=(10,10))
#Load Omega_pbh data at early and late times
dat1 = np.loadtxt("data/8gev+9orad+rem/peakm_5e-7_rem.dat")
dat2 = np.loadtxt("data/8gev+9orad+rem/peakm_2e11_rem.dat")
dat4 = np.loadtxt("data/8gev+9orad+rem/peakm_5e13_rem.dat")
dat1a = np.loadtxt("data/8gev+9orad/peakm_5e-7.dat")
dat2b = np.loadtxt("data/8gev+9orad/peakm_2e11.dat")
dat4c = np.loadtxt("data/8gev+9orad/peakm_5e13.dat")
avals = np.logspace(-6, 0, num=1000)
ax = plt.subplot(2, 1, 1)
plt.xscale('log')
#plt.plot(avals, Omegac(avals),linestyle='dashed', color='b', label= '$\Omega_c$',alpha=1.)
plt.plot(dat1[:,0], dat1[:,1]/dat1a[:,1] , label='$M_{pk} = 5\\times 10^{-7}$kg', alpha=0.6)
plt.plot(dat2[:,0], dat2[:,1]/dat2b[:,1], label='$M_{pk} = 2\\times 10^{11}$kg', alpha=0.6)
#plt.plot(dat3[:,0], dat3[:,1], label='$M_{pk} = 5\\times 10^{11}$kg',alpha=0.6)
#plt.plot(dat4[:,0], dat4[:,1]/dat4c [:,1], label='$M_{pk} = 5\\times 10^{13}$kg',alpha=0.6)
#plt.plot(dat5[:,0], dat5[:,1], label='$M_{pk} = 5\\times 10^{33}$kg',alpha=0.6)
plt.axhline(y=1., xmin=0., xmax=10,color='k',linestyle='dashed')
plt.axvline(acmb,0.,10, color='k', linestyle='dotted')
plt.text(2e-6, 0.3 , '$T_{\\rm RH} = 10^{8}{\\rm GeV}$', **font1)
ax.tick_params(axis='both', which='major', labelsize=15)
plt.ylim(0.,2)
plt.xlim(ai,1)
plt.ylabel('Density fraction of PBH ($\Omega_{\\rm PBH}$) ',**font1)
plt.xlabel('scale factor (a)', **font1)
plt.legend(loc='best',prop={'size': 16})
plt.tight_layout(pad=3.0)
ax = plt.subplot(2, 1, 2)
plt.xscale('log')
#plt.plot(avals, Omegab(avals),linestyle='dashed', color='r', label= '$\Omega_b$',alpha=1.)
plt.plot(dat1[:,0], dat1[:,2]/dat1a[:,2], label='$\lambda = 3.4\\times 10^{96}$', alpha=0.6)
plt.plot(dat2[:,0], dat2[:,2]/dat2b[:,2], label='$\lambda = 1.9\\times 10^{98}$', alpha=0.6)
#plt.plot(dat3[:,0], dat3[:,2], label='$\lambda = 4.8\\times 10^{98}$', alpha=0.6)
#plt.plot(dat4[:,0], dat4[:,2]/dat4c[:,2], label='$\lambda = 4.7\\times 10^{100}$', alpha=0.6)
#plt.plot(dat5[:,0], dat5[:,2], label='$\lambda = 3.5\\times 10^{120}$', alpha=0.6)
plt.axvline(acmb,0.,10, color='k', linestyle='dotted')
ax.tick_params(axis='both', which='major', labelsize=15)
plt.ylim(0.6,1.4)
plt.xlim(ai,1)
plt.ylabel('Density fraction of baryons ($\Omega_{\\rm b}$) ',**font1)
plt.xlabel('scale factor (a)', **font1)
plt.legend(loc='best',prop={'size': 16})
#plt.setp(plt.subplot(2,1,1).get_xticklabels(), visible=False)
plt.subplots_adjust(hspace=0.2)
plt.subplots_adjust(wspace=0.)
plt.savefig('plots/remnants.png', format="png", bbox_inches = 'tight')
# -
| plots.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Copyright 2019 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# -
# <a target="_blank" href="https://colab.research.google.com/github/GoogleCloudPlatform/keras-idiomatic-programmer/blob/master/workshops/Training/Idiomatic%20Programmer%20-%20handbook%203%20-%20Codelab%201.ipynb">
# <img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# # Idiomatic Programmer Code Labs
#
# ## Code Labs #1 - Get Familiar with Hyperparameters
#
# ## Prerequistes:
#
# 1. Familiar with Python
# 2. Completed Handbook 3/Part 10: Training Preparation and Hyperparameters
#
# ## Objectives:
#
# 1. Hand setting Epochs and Mini-Batches
# 2. Use ImageDataGenerator for batch generation.
# 3. Finding good learning rate.
# ## Epochs and Mini-Batches
#
# In this section, we will hand-roll our own code (vs. builtin feeders) to feed the training data for training. We will need to handle the following:
#
# 1. Set a mini-batch size (128) and calculate how many batches will be in the training data.
# 1. Set the number of epochs (number of times we pass the full training data for training)
# 2. Randomly shuffle the training data on each epoch.
# 3. Iterate through the training data on batch at a time.
#
# You fill in the blanks (replace the ??), make sure it passes the Python interpreter.
# +
from keras.datasets import cifar10
import random
# Let's use the CIFAR-10 dataset
(x_train, y_train), (x_test, y_test) = cifar10.load_data()
# We will use a mini-batch size of 128
batch_size = 128
# Calculate the total number of mini-batches in an epoch
# HINT: It has something to do with the (mini) batch size
batches = len(x_train) // ??
# Let's use a seed so we can randomly shuffle both the pixel data and labels the same shuffle.
seed = 101
# Let's do 5 passes (epochs) over the dataset
epochs = 5
for epoch in range(epochs):
# Shuffle the dataset at the beginning of each epoch
# HINT: We have to shuffle the image data and labels from the training data
random.seed(seed)
random.shuffle(??)
random.seed(seed)
random.shuffle(??)
# Set a new seed for the next shuffle
seed += random.randint(0, 100)
# Iterate (sequential) through the shuffled training data, one batch at a time.
for batch in range(batches):
# Get the next batch of data
# HINT: if the begin of the batch is at location X, then the end is X + batc
x_batch = x_train[batch * batch_size:(batch+??) * batch_size]
y_batch = y_train[batch * batch_size:(batch+??) * batch_size]
print("Epoch", epoch+1, "Batch", batch+1)
print("Done - the last line above this should be: Epoch 5, Batch 390")
# -
# ## ImageDataGenerator and Batch Generation
#
# In this section, we will use the **Keras** ImageDataGenerator to automatically generate out mini-batches (vs. hand generating them), and shuffling the training data on each epoch.
# +
from keras.preprocessing.image import ImageDataGenerator
# Let's use the CIFAR-10 dataset
(x_train, y_train), (x_test, y_test) = cifar10.load_data()
# We will use a mini-batch size of 128
batch_size = 128
# Calculate the total number of mini-batches in an epoch
batches = len(x_train) // batch_size
# instantiate an Image Data generator object
# HINT: Image Data generator is a big giveaway
datagen = ??()
# Let's do 5 passes (epochs) over the dataset
epochs = 5
for epoch in range(epochs):
# Use generator to create batches
# HINT: The method is about flowing data from in-memory (vs. on-disk)
batch = 0
for x_batch, y_batch in datagen.??(x_train, y_train, batch_size=batch_size, shuffle=True):
batch += 1 # Keep track of the number of batches so far.
print("Epoch", epoch+1, "Batch", batch)
# At the end of the training data, let's loop around for the next epoch.
if batch == batches:
break
print("Done - the last line above this should be: Epoch 5, Batch 390")
# -
# ## Learning Rate
#
# Let's show how to do short epochs to get a feel on what might be the right learning rate for your training.
# +
from keras import Sequential, optimizers
from keras.layers import Conv2D, MaxPooling2D, Flatten, Dense
from keras.utils import to_categorical
import numpy as np
# Let's use the CIFAR-10 dataset
(x_train, y_train), (x_test, y_test) = cifar10.load_data()
# Normalize the pixel data
x_train = (x_train / 255.0).astype(np.float32)
x_test = (x_test / 255.0).astype(np.float32)
# One-hot encode the labels
y_train = to_categorical(y_train)
y_test = to_categorical(y_test)
def convNet(input_shape, nclasses):
model = Sequential()
model.add(Conv2D(32, kernel_size=(3, 3),
activation='relu',
input_shape=input_shape))
model.add(Conv2D(64, (3, 3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(Dense(nclasses, activation='softmax'))
return model
# Create a simple CNN and set learning rate very high (0.1))
# HINT: how would you abbreviate learning rate?
model = convNet((32, 32, 3), 10)
model.compile(loss='categorical_crossentropy',
optimizer=optimizers.Adam(??=0.1),
metrics=['accuracy'])
# Let's take a fraction of the training data to test the learning rate (2%)
x_tmp = x_train[0:1000]
y_tmp = y_train[0:1000]
# Let's run 3 epochs at learning rate = 0.1
model.fit(x_tmp, y_tmp, epochs=3, batch_size=32, verbose=1)
# -
# Argh, it's horrible. The loss on the first epoch is high (14.0+) and then never goes down - like it's stuck.
#
# Hum, okay now you experiment with different learning rates to find one where the loss goes down rapidly and a steady increase in accuracy.
# +
model = convNet((32, 32, 3), 10)
# Pick your own learning rate until the results are good.
# HINT: It's going to be a lot smaller than 0.1
model.compile(loss='categorical_crossentropy',
optimizer=optimizers.Adam(lr=??),
metrics=['accuracy'])
# Let's run 3 epochs at your learning rate
model.fit(x_tmp, y_tmp, epochs=3, batch_size=32, verbose=1)
# -
# ## End of Code Lab
| workshops/Training/Idiomatic Programmer - handbook 3 - Codelab 1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (base)
# language: python
# name: base
# ---
import numpy as np
import math
import matplotlib.pyplot as plt
# # %matplotlib inline
import mpld3
# mpld3.enable_notebook()
file=open("kalmann.txt",'r')
pos=np.loadtxt("kalmann.txt",delimiter=',',usecols=[0,1])
vel=np.loadtxt("kalmann.txt",skiprows=1,delimiter=',',usecols=[2,3])
vel=np.insert(vel,[0],0,axis=0)
# defining varibles
del_t=0.075
sigma_ax=0.03
sigma_ay=0.02
sigma_x=0.1
sigma_y=0.1
sigma_vx=5
sigma_vy=5
pos_x0,pos_y0=pos[0]
p=1000
pred_state=[]
est_state=[]
pred_p=[]
est_p=[]
F=np.zeros((6,6))
F[0,:3]=(1,del_t,0.5*del_t**2)
F[1,:3]=(0,1,del_t)
F[2,2]=1
F[3,3:6]=(1,del_t,0.5*del_t**2)
F[4,3:6]=(0,1,del_t)
F[5,5]=1
FT=np.transpose(F)
# F
# +
# Q=np.zeros((6,6))
# Q[0,:3]=(0.05*del_t**5*sigma_ax**2,0.125*del_t**4*sigma_ax**2,(1/6)*del_t**3*sigma_ax**2)
# Q[1,:3]=(0,(1/3)*del_t**3*sigma_ax**2,0.5*del_t**2*sigma_ax**2)
# Q[2,:3]=(0,0,1*sigma_ax**2*del_t)
# Q[3,3:6]=(0.05*del_t**5*sigma_ay**2,0.125*del_t**4*sigma_ay**2,(1/6)*del_t**3*sigma_ay**2)
# Q[4,3:6]=(0,(1/3)*del_t**3*sigma_ay**2,0.5*del_t**2*sigma_ay**2)
# Q[5,3:6]=(0,0,1*sigma_ay**2*del_t)
# # Q=Q*sigma_ay**2
# # Q-Continuos noise
# -
Q=np.zeros((6,6))
Q[0,:3]=(0.25*del_t**4*sigma_ax**2,0.5*del_t**3*sigma_ax**2,0.5*del_t**2*sigma_ax**2)
Q[1,:3]=(0,del_t**2*sigma_ax**2,del_t*sigma_ax**2)
Q[2,:3]=(0,0,1*sigma_ax**2)
Q[3,3:6]=(0.25*del_t**4*sigma_ay**2,0.5*del_t**3*sigma_ay**2,0.5*del_t**2*sigma_ay**2)
Q[4,3:6]=(0,del_t**2*sigma_ay**2,del_t*sigma_ay**2)
Q[5,3:6]=(0,0,1*sigma_ay**2)
# Q=Q*sigma_ay**2
# Q-Discrete noise
R=np.zeros((4,4))
R[0,0],R[1,1],R[2,2],R[3,3]=(sigma_x**2,sigma_vx**2,sigma_y**2,sigma_vy**2)
# R
H=np.zeros((4,6))
H[0,0],H[1,1],H[2,3],H[3,4] =(1,1,1,1)
HT=np.transpose(H)
# H
temp=np.zeros((6,1))
temp[:,0]=[pos_x0,0,0,pos_y0,0,0]
est_state.append(temp)
# est_state[0]
temp=np.diag((p,p,p,p,p,p))
est_p.append(temp)
# est_p[0]
def state_extrapolate(xn):
xnext=np.dot(F,xn)
return xnext
def cov_extrapolate(pn):
pnext=np.dot(np.dot(F,pn),FT) +Q
return pnext
def measure(n):
z=np.zeros((4,1))
z[:,0]=(pos[n,0],vel[n,0],pos[n,1],vel[n,1])
return z
def K_gain(n):
kn=np.dot(np.dot(pred_p[n-1],HT),(np.linalg.inv(np.dot(np.dot(H,pred_p[n-1]),HT)+R)))
return kn
def state_update(xn_1,kn,zn):
xn=xn_1 + np.dot(kn,(zn-np.dot(H,xn_1)))
return xn
def cov_update(pn_1,kn):
idm=(np.identity(6)-np.dot(kn,H))
idmT=np.transpose(idm)
pn=np.dot(np.dot(idm,pn_1),idmT)+ np.dot(np.dot(kn,R),np.transpose(kn))
return pn
# +
pred_state.append(state_extrapolate(est_state[0]))
pred_p.append(cov_extrapolate(est_p[0]))
for i in range(1,len(vel)):
print("\n---------------------------------\n\nIteration :-",i)
zi=measure(i)
ki=K_gain(i)
est_state.append(state_update(pred_state[i-1],ki,zi))
print("Estimated position(x,y) :- (%g,%g)" % (est_state[i][0],est_state[i][3]))
est_p.append(cov_update(pred_p[i-1],ki))
pred_state.append(state_extrapolate(est_state[i]))
print("Uncertainity(x,y) :- (%g,%g)" % (est_p[i][0,0],est_p[i][3,3]))
pred_p.append(cov_extrapolate(est_p[i]))
# -
i=np.arange(1,len(vel))
ar=[]
br=[]
m=[]
for p in range(1,len(vel)):
ar.append(pred_state[p][0])
br.append(est_state[p][0])
m.append(pos[p][0])
# pred_state[2][0]
# plt.plot(i,ar,color='orange')
plt.plot(i,br,color='blue')
plt.plot(i,m,color='red')
plt.title("comparision of estimated x(blue) and measured x(red)")
plt.show()
# comparision of estimated x(blue) and measured x(red)
i=np.arange(1,len(vel))
ar=[]
br=[]
m=[]
for p in range(1,len(vel)):
ar.append(pred_state[p][3])
br.append(est_state[p][3])
m.append(pos[p][1])
# pred_state[2][0]
# plt.plot(i,ar,color='orange')
plt.plot(i,br,color='blue')
plt.plot(i,m,color='red')
plt.title("comparision of estimated y(blue) and measured y(red)")
plt.show()
# comparision of estimated y(blue) and measured y(red)
i=np.arange(1,len(vel))
ar=[]
br=[]
m=[]
for p in range(1,len(vel)):
ar.append(est_state[p][0])
br.append(est_state[p][3])
m.append(pos[p][0])
# pred_state[2][0]
plt.plot(ar,br,color='blue')
plt.title("estimated trajectory")
plt.show()
# plt.plot(i,br)
# plt.plot(i,m)
# estimated trajectory
i=np.arange(1,len(vel))
ar=[]
br=[]
m=[]
for p in range(1,len(vel)):
ar.append(pos[p][0])
br.append(pos[p][1])
# m.append(pos[p][0])
# pred_state[2][0]
plt.plot(ar,br,color='red')
plt.title("measured trajectory")
plt.show()
# plt.plot(i,br)
# plt.plot(i,m)
# measured trajectory
i=np.arange(1,len(vel))
ar=[]
br=[]
ar1=[]
br1=[]
ar2=[]
br2=[]
m=[]
for p in range(1,len(vel)):
ar.append(est_state[p][0])
br.append(est_state[p][3])
ar1.append(pred_state[p][0])
br1.append(pred_state[p][3])
ar2.append(pos[p][0])
br2.append(pos[p][1])
# m.append(pos[p][0])
# pred_state[2][0]
plt.plot(ar,br,color='blue')
# plt.plot(ar1,br1,color='orange')
plt.plot(ar2,br2,color='red')
plt.title("measured(red) and estimated(blue) trjectory")
# plt.plot(i,br)
# plt.plot(i,m)
# measured(red) and estimated(blue) trjectory
plt.show()
| Task 3/code/Final-Corrected.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
df = pd.read_csv('https://archive.ics.uci.edu/ml/\
machine-learning-databases/breast-cancer-wisconsin/wdbc.data',header = None)
#把字符串类标转换成整数
from sklearn.preprocessing import LabelEncoder
X = df.loc[:,2:].values
y = df.loc[:,1].values
le = LabelEncoder()
y = le.fit_transform(y)
le.transform(['M','B'])
from sklearn.cross_validation import train_test_split
X_train,X_test,y_train,y_test = train_test_split(X,y,test_size = 0.2,random_state = 1)
#在流水线中集成操作
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
from sklearn.linear_model import LogisticRegression
from sklearn.pipeline import Pipeline
pipe_lr = Pipeline([('scl',StandardScaler()),('pca',PCA(n_components = 2)),
('clf',LogisticRegression(random_state = 1))])
pipe_lr.fit(X_train,y_train)
print('Test Accuracy: %.3f' % pipe_lr.score(X_test,y_test))
#k折交叉验证
import numpy as np
from sklearn.cross_validation import StratifiedKFold
kfold = StratifiedKFold(y = y_train,n_folds = 10,random_state = 1)
scores = []
for k,(train,test) in enumerate(kfold):
pipe_lr.fit(X_train[train],y_train[train])
score = pipe_lr.score(X_train[test],y_train[test])
scores.append(score)
print('Fold: %s,Class dist.: %s,Acc: %.3f' % (k+1,np.bincount(y_train[train]),score))
print('CV accuracy: %.3f +/- %.3f' % (np.mean(scores),np.std(scores)))
#绘制学习曲线
import matplotlib.pyplot as plt
from sklearn.learning_curve import learning_curve
pipe_lr = Pipeline([('scl',StandardScaler()),
('clf',LogisticRegression(penalty = 'l2',random_state = 0))])
train_sizes,train_scores,test_scores = \
learning_curve(estimator = pipe_lr,X = X_train,y = y_train,
train_sizes = np.linspace(0.1,1.0,10),cv = 10,n_jobs = 1)
train_mean = np.mean(train_scores,axis = 1)
train_std = np.std(train_scores,axis = 1)
test_mean = np.mean(test_scores,axis = 1)
test_std = np.std(test_scores,axis = 1)
plt.plot(train_sizes,train_mean,color = 'blue',marker = 'o',
markersize = 5,label = 'training accuracy')
plt.fill_between(train_sizes,train_mean + train_std,train_mean-train_std,
alpha = 0.15,color = 'blue')
plt.plot(train_sizes,test_mean,color = 'green',linestyle = '--',
marker = 's', markersize = 5,label = 'validation accuracy')
plt.fill_between(train_sizes,test_mean + test_std,test_mean-test_std,
alpha = 0.15,color = 'green')
plt.grid()
plt.xlabel('Number of train samples')
plt.ylabel('Accuracy')
plt.legend(loc = 'lower right')
plt.ylim([0.8,1.0])
plt.show()
#绘制验证曲线
from sklearn.learning_curve import validation_curve
param_range = [0.001,0.01,0.1,1.0,10.0,100.0]
train_scores,test_scores = validation_curve(estimator = pipe_lr,
X = X_train,y = y_train,param_name = 'clf__C',
param_range = param_range,cv = 10)
train_mean = np.mean(train_scores,axis = 1)
train_std = np.std(train_scores,axis = 1)
test_mean = np.mean(test_scores,axis = 1)
test_std = np.std(test_scores,axis = 1)
plt.plot(param_range,train_mean,color = 'blue',marker = 'o',
markersize = 5,label = 'training accuracy')
plt.fill_between(param_range,train_mean + train_std,train_mean-train_std,
alpha = 0.15,color = 'blue')
plt.plot(param_range,test_mean,color = 'green',linestyle = '--',
marker = 's', markersize = 5,label = 'validation accuracy')
plt.fill_between(param_range,test_mean + test_std,test_mean-test_std,
alpha = 0.15,color = 'green')
plt.grid()
plt.xscale('log')
plt.legend(loc = 'lower right')
plt.xlabel('Parameter C')
plt.ylabel('Accuracy')
plt.ylim([0.8,1.0])
plt.show()
#使用网络搜索调优超参
from sklearn.grid_search import GridSearchCV
from sklearn.svm import SVC
pipe_svc = Pipeline([('scl',StandardScaler()),
('clf',SVC(random_state = 1))])
param_range = [0.0001,0.001,0.01,0.1,1.0,10.0,100.0,1000.0]
param_grid = [{'clf__C':param_range,'clf__kernel':['linear']},
{'clf__C':param_range,'clf__gamma':param_range,'clf__kernel':['rbf']}]
gs = GridSearchCV(estimator = pipe_svc,param_grid = param_grid,
scoring = 'accuracy',cv = 10,n_jobs = -1)
gs = gs.fit(X_train,y_train)
print(gs.best_score_)
print(gs.best_params_)
clf =gs.best_estimator_
clf.fit(X_train,y_train)
print('Test accuracy: %.3f' % clf.score(X_test,y_test))
#通过嵌套交叉验证选择所需算法
from sklearn.model_selection import cross_val_score
gs = GridSearchCV(estimator = pipe_svc,param_grid = param_grid,
scoring = 'accuracy',cv = 10,n_jobs = -1)
scores = cross_val_score(gs,X,y,scoring = 'accuracy',cv = 5)
print('CV accuracy: %.3f +/- %.3f' % (np.mean(scores),np.std(scores)))
from sklearn.tree import DecisionTreeClassifier
gs = GridSearchCV(estimator = DecisionTreeClassifier(random_state = 0),
param_grid = [{'max depth':[1,2,3,4,5,6,7,None]}],
scoring = 'accuracy',cv = 5)
scores = cross_val_score(gs,X_train,y_train,scoring = 'accuracy',cv = 5)
print('CV accuracy: %.3f +/- %.3f' % (np.mean(scores),np.std(scores)))
#绘制混淆矩阵
from sklearn.metrics import confusion_matrix
pipe_svc.fit(X_train,y_train)
y_pred = pipe_svc.predict(X_test)
confmat = confusion_matrix(y_true = y_test,y_pred = y_pred)
print(confmat)
# +
#绘制ROC曲线
from sklearn.metrics import roc_curve,auc
from scipy import interp
X_train2 = X_train[:,[4,14]]
cv = StratifiedKFold(y_train,n_folds = 3,random_state = 1)
fig = plt.figure(figsize = (7,5))
mean_tpr = 0.0
mean_fpr = np.linspace(0,1,100)
all_tpr = []
for i,(train,test) in enumerate(cv):
probas = pipe_lr.fit(X_train2[train],y_train[train]).predict_proba(X_train2[test])
fpr,tpr,thresholds = roc_curve(y_train[test],probas[:,1],pos_label = 1)
mean_tpr += interp(mean_fpr,fpr,tpr)
mean_tpr[0] = 0.0
roc_auc = auc(fpr,tpr)
plt.plot(fpr,tpr,lw = 1,label = 'ROC fold %d (area = %0.2f)' %(i+1,roc_auc))
plt.plot([0,1],[0,1],linestyle = '--',color = (0.6,0.6,0.6),label = 'random guessing')
mean_tpr /= len(cv)
mean_tpr[-1] /= 1.0
mean_auc = auc(mean_fpr,mean_tpr)
plt.plot(mean_fpr,mean_tpr,'k--',label = 'mean ROC (area = %0.2f)' % mean_auc,lw = 2)
plt.plot([0,0,1],[0,1,1],lw = 2,linestyle = ':',color = 'black',label = 'perfect performance')
plt.xlim([-0.05,1.05])
plt.ylim([-0.05,1.05])
plt.xlabel('false positive rate')
plt.ylabel('true positive rate')
plt.title('Receiver Operator Characteristic')
plt.legend(loc = "lower right")
plt.show()
# -
| ch06model_optimize.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from neuron.mnist import mnist_loader
import neuron.mnist.network_sandbox as network
import matplotlib.pyplot as plt
import matplotlib.cm as cm
import numpy as np
import pandas as pd
import seaborn as sns
from neuron.activation_functions import sigmoid
from neuron.mnist import stepik_original as stepik
np.set_printoptions(precision=5)
% load_ext autoreload
% autoreload 2
# -
# ### The networks are fully reconciled. With full data set you may even use different mnist parser. The final weights should be reconciled
# # PARSE DATA
# +
import os
os.chdir("C:\\Users\\mkapchenko\\Dropbox\\perso\\GitHub\\Neuron\\notebooks")
training_data, validation_data, test_data = mnist_loader.load_data_wrapper()
n = 50000
nepochs = 30
nb_batches = 10
batch_size = 784//nb_batches
mini_res = list(training_data)[0:n]
mini_test = list(test_data)[0:n]
# to have the same data
minX = np.array(mini_res[0][0])
miny = np.array(mini_res[0][1])
# -
(X, y), (validation_X, validation_y), (Xtest, ytest) = mnist_loader.perf_load_data_wrapper()
n = 50000
(X, y) = (X[0:n], y[0:n])
# # LEARNING
# ### Neuron
# +
# %%time
# 28 s
# minX, miny = X[0:n], y[0:n]
netw = network.Network([784, 30, 10])
netw.SGD(X, y, epochs=nepochs, batch_size=batch_size, learning_rate = 3., test_data = (Xtest, ytest));
# -
# ### Stepik
# %%time
# 9min 9s
# Stepik network
netstepik = stepik.Network([784, 30, 10])
netstepik.SGD(mini_res, epochs=nepochs, mini_batch_size=batch_size, eta=3.0, test_data=mini_test)
# # RECON
# ### Final weights
for layer in range(2):
print(f'Layer {layer} recon: ' , (netstepik.weights[layer] - netw.weights[layer]).mean())
| notebooks/gold standard/recon_sandbox.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.5 64-bit
# metadata:
# interpreter:
# hash: 1ee38ef4a5a9feb55287fd749643f13d043cb0a7addaab2a9c224cbe137c0062
# name: python3
# ---
# # Lab Two
# ---
#
# For this lab we're going to get into logic
#
# Our Goals are:
# - Using Conditionals
# - Using Loops
# - Creating a Function
# - Using a Class
# + tags=[]
# Create an if statement
if True:
print("Done")
# + tags=[]
# Create an if else statement
if False:
print("If Statement Printing")
else:
print("Else Statement Printing")
# + tags=[]
# Create an if elif else statement
if False:
print("If Statement Printing")
elif True:
print("Elif Statement Printing")
else:
print("Else Statement Printing")
# + tags=[]
# Create a for loop using range(). Go from 0 to 9. Print out each number.
for number in range(10):
print(number)
# + tags=[]
# Create a for loop iterating through this list and printing out the value.
arr = ['Blue', 'Yellow', 'Red', 'Green', 'Purple', 'Magenta', 'Lilac']
for color in arr:
print(color)
# Get the length of the list above and print it.
print(len(arr))
# + tags=[]
# Create a while loop that ends after 6 times through. Print something for each pass.
index = 0
while index < 6:
print(index)
index += 1
# + tags=[]
# Create a function to add 2 numbers together. Print out the number. I messed up. Not required.
def addition(input1, input2):
total = input1 + input2
print(total)
addition(10, 94)
# + tags=[]
# Create a function that tells you if a number is odd or even and print the result. I messed up. Not required.
def odd_or_even(number):
if number % 2 == 0:
print("Number is even")
else:
print("Number is odd")
odd_or_even(0)
# + tags=[]
# Initialize an instance of the following class. Use a variable to store the object and then call the info function to print out the attributes.
class Dog(object):
def __init__(self, name, height, weight, breed):
self.name = name
self.height = height
self.weight = weight
self.breed = breed
def info(self):
print("Name:", self.name)
print("Weight:", str(self.weight) + " Pounds")
print("Height:", str(self.height) + " Inches")
print("Breed:", self.breed)
Butters = Dog("Butters", "20", "14", "Corgi")
Butters.info()
# -
| JupyterNotebooks/Labs/Lab 2 Solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: fengine
# language: python
# name: fengine
# ---
# ## Gaussian Transformation with Feature-Engine
#
# Scikit-learn has recently released transformers to do Gaussian mappings as they call the variable transformations. The PowerTransformer allows to do Box-Cox and Yeo-Johnson transformation. With the FunctionTransformer, we can specify any function we want.
#
# The transformers per se, do not allow to select columns, but we can do so using a third transformer, the ColumnTransformer
#
# Another thing to keep in mind is that Scikit-learn transformers return NumPy arrays, and not dataframes, so we need to be mindful of the order of the columns not to mess up with our features.
#
# ## Important
#
# Box-Cox and Yeo-Johnson transformations need to learn their parameters from the data. Therefore, as always, before attempting any transformation it is important to divide the dataset into train and test set.
#
# In this demo, I will not do so for simplicity, but when using this transformation in your pipelines, please make sure you do so.
#
#
# ## In this demo
#
# We will see how to implement variable transformations using Scikit-learn and the House Prices dataset.
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import scipy.stats as stats
import feature_engine.transformation as vt
# -
data = pd.read_csv('../houseprice.csv')
data.head()
# ## Plots to assess normality
#
# To visualise the distribution of the variables, we plot a histogram and a Q-Q plot. In the Q-Q pLots, if the variable is normally distributed, the values of the variable should fall in a 45 degree line when plotted against the theoretical quantiles. We discussed this extensively in Section 3 of this course.
# +
# plot the histograms to have a quick look at the variable distribution
# histogram and Q-Q plots
def diagnostic_plots(df, variable):
# function to plot a histogram and a Q-Q plot
# side by side, for a certain variable
plt.figure(figsize=(15,6))
plt.subplot(1, 2, 1)
df[variable].hist()
plt.subplot(1, 2, 2)
stats.probplot(df[variable], dist="norm", plot=plt)
plt.show()
# -
diagnostic_plots(data, 'LotArea')
diagnostic_plots(data, 'GrLivArea')
# ## LogTransformer
lt = vt.LogTransformer(variables = ['LotArea', 'GrLivArea'])
lt.fit(data)
# variables that will be transformed
lt.variables
data_tf = lt.transform(data)
diagnostic_plots(data_tf, 'LotArea')
# transformed variable
diagnostic_plots(data_tf, 'GrLivArea')
# ## ReciprocalTransformer
rt = vt.ReciprocalTransformer(variables = ['LotArea', 'GrLivArea'])
rt.fit(data)
data_tf = rt.transform(data)
diagnostic_plots(data_tf, 'LotArea')
# transformed variable
diagnostic_plots(data_tf, 'GrLivArea')
# ## ExponentialTransformer
et = vt.PowerTransformer(variables = ['LotArea', 'GrLivArea'])
et.fit(data)
data_tf = et.transform(data)
diagnostic_plots(data_tf, 'LotArea')
# transformed variable
diagnostic_plots(data_tf, 'GrLivArea')
# ## BoxCoxTransformer
bct = vt.BoxCoxTransformer(variables = ['LotArea', 'GrLivArea'])
bct.fit(data)
# these are the exponents for the BoxCox transformation
bct.lambda_dict_
data_tf = bct.transform(data)
diagnostic_plots(data_tf, 'LotArea')
# transformed variable
diagnostic_plots(data_tf, 'GrLivArea')
# ## Yeo-Johnson Transformer
#
# Yeo-Johnson Transformer will be available in the next release of Feauture-Engine!!!
yjt = vt.YeoJohnsonTransformer(variables = ['LotArea', 'GrLivArea'])
yjt.fit(data)
# these are the exponents for the Yeo-Johnson transformation
yjt.lambda_dict_
data_tf = yjt.transform(data)
diagnostic_plots(data_tf, 'LotArea')
# transformed variable
diagnostic_plots(data_tf, 'GrLivArea')
| Section-07-Variable-Transformation/07.03-Gaussian-transformation-feature-engine.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from sklearn.datasets import make_regression, make_moons
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
from sklearn.neural_network import MLPRegressor
from sklearn.ensemble import RandomForestRegressor
from sklearn.metrics import r2_score, explained_variance_score
from scipy.stats import pearsonr, spearmanr
# +
def interval_transform(x, a, b):
m = x.min()
ma = x.max()
alpha_inv = (1 - m/ma)*ma/(a - b)
alpha = 1/alpha_inv
beta = b - alpha*m
f = lambda x: alpha*x + beta
return f(x)
def make_noise_feature(x):
n_features = x.shape[1]
n_samples = x.shape[0]
weights = np.random.uniform(1e-4, 1e-2, n_features)
noise = np.random.normal(1, 5, n_samples)
signal = np.sum(weights*x, -1)
return signal + noise
def calculate_pvalues(df,
method = spearmanr
):
"""
Assumes df with only numeric entries clean of null entries.
"""
dfcols = pd.DataFrame(columns=df.columns)
pvalues = dfcols.transpose().join(dfcols, how='outer')
for r in df.columns:
for c in df.columns:
pvalues[r][c] = round(method(df[r], df[c])[1], 4)
return pvalues
def correlation_matrix(df,
method = "pearson",
annot_bool = False,
annot_size = 20
):
# Compute the correlation matrix
corr = df.corr(method = method)
if annot_bool:
annot = corr.copy()
if method == "pearson":
sig_meth = pearsonr
else:
sig_meth = spearmanr
pval = calculate_pvalues(df, sig_meth)
# create three masks
r0 = corr.applymap(lambda x: '{:.2f}'.format(x))
r1 = corr.applymap(lambda x: '{:.2f}*'.format(x))
r2 = corr.applymap(lambda x: '{:.2f}**'.format(x))
r3 = corr.applymap(lambda x: '{:.2f}***'.format(x))
# apply them where appropriate --this could be a single liner
annot = annot.where(pval>0.1,r0)
annot = annot.where(pval<=0.1,r1)
annot = annot.where(pval<=0.05,r2)
annot = annot.mask(pval<=0.01,r3)
# Generate a mask for the upper triangle
mask = np.zeros_like(corr, dtype=np.bool)
mask[np.triu_indices_from(mask)] = True
# Set up the matplotlib figure
f, ax = plt.subplots(figsize=(11, 11))
# Generate a custom diverging colormap
cmap = sns.diverging_palette(220, 10, as_cmap=True)
# Draw the heatmap with the mask and correct aspect ratio
sns.heatmap(corr, mask=mask, cmap=cmap, center=0,
square=True, linewidths=.5, cbar_kws={"shrink": .5},
annot = annot,
fmt = "",
annot_kws={"size": annot_size},
vmin = -1,
vmax = 1,
)
# +
n_info = 3
n_redu = 0
n_samples=2000
#making nonlinear decision boundaries requires multiple blob like features
X1, y1 = make_regression(
n_samples=n_samples,
n_features=3,
n_informative=n_info,
#n_redundant=n_redu,
shuffle=False,
random_state=42,
#difficulty
effective_rank=2,
noise=0.6,
tail_strength=0.2,
bias=12,
)
X2, y2 = make_regression(
n_samples=n_samples,
n_features=3,
n_informative=n_info,
#n_redundant=n_redu,
shuffle=False,
random_state=6,
#difficulty
effective_rank=1,
noise=1.1,
tail_strength=0.3,
bias=10,
)
#X3, y3 = make_moons(n_samples=2*n_samples, noise=1, random_state=42)
X = np.concatenate([X1, X2], axis=0)
y = np.concatenate([y1, y2], axis=0)
data = np.concatenate([X, np.expand_dims(y, -1)], -1)
data = pd.DataFrame(data)
# -
print(X.shape)
data.head()
correlation_matrix(data, annot_bool=True, annot_size=15)
plt.hist(y)
# To educationify the data we scale and transform the dataset:
#
# For the informative features we'll use:
# * GPA [0, 4] unit:grade
# * Attendance [0, 100] unit:percent
# * Passed percent of classes [0, 100] unit:percent
#
# For the redundant we'll use:
# * Sex [0, 1] unit:integer class
# * Ethnicity [0, 1, 2] unit:integer class
# * HSGPA [0, 4] unit:grade
#
fig, axs = plt.subplots(nrows=n_info, figsize=(5, 10 ))
for i in range(n_info):
ax = axs[i]
data[i].plot(kind="hist", ax=ax)
attendance_column = interval_transform(data[2], 0, 100)
plt.hist(attendance_column)
gpa_column = interval_transform(data[1], 1, 4)
plt.hist(gpa_column)
passed_column = interval_transform(data[0], 0, 100)
plt.hist(passed_column)
sex_column = make_noise_feature(X)
sex_column = (sex_column > sex_column.mean()).astype(int)
plt.hist(sex_column)
hsgpa_column = interval_transform(make_noise_feature(X), 0, 4)
plt.hist(hsgpa_column)
ethn_column = make_noise_feature(X)
ethn_column = pd.qcut(ethn_column, q=[0, .25, .5, 1], labels=[0, 1, 2])
plt.hist(ethn_column)
fci_post = interval_transform(y, 0, 30)
plt.hist(fci_post)
gpa_column.shape
# +
full_data = np.concatenate(
[
np.expand_dims(gpa_column, axis=-1),
np.expand_dims(attendance_column, axis=-1),
np.expand_dims(passed_column, axis=-1),
np.expand_dims(sex_column, axis=-1),
np.expand_dims(hsgpa_column, axis=-1),
np.expand_dims(ethn_column, axis=-1),
np.expand_dims(fci_post, axis=-1)
],
axis=1
)
columns = [
"cGPA",
"attendance",
"passed_percent",
"sex",
"hsGPA",
"ethnicity",
"fci_post"]
df_full = pd.DataFrame(full_data,
columns=columns)
# -
df_full.head()
# +
comb = [(1, 0), (2, 0), (2, 1)]
fig, axs = plt.subplots(nrows=len(comb), figsize=(10, 7))
for i in range(len(comb)):
sns.scatterplot(full_data[:,comb[i][0]], full_data[:,comb[i][1]], hue=y, ax=axs[i], alpha=0.3)
# -
correlation_matrix(df_full, annot_bool=True, annot_size=15)
# +
t_X = X.copy()
fd = full_data[:, :-1].copy()
for i in range(t_X.shape[1]):
t_X[:,i] = (t_X[:,i] - t_X[:,i].mean())/t_X[:,i].std()
for i in [0, 1, 2, -1]:
fd[:,i] = (fd[:,i] - fd[:,i].mean())/fd[:,i].std()
data_c = [c for c in columns if c != "fci_post"]
clf_data = df_full[data_c].values
clf_targets = df_full["fci_post"].values
#x_train, x_test, y_train, y_test = train_test_split(t_X, y, shuffle=True)
x_train, x_test, y_train, y_test = train_test_split(fd, full_data[:,-1], shuffle=True)
# +
model = LinearRegression()
model.fit(x_train, y_train)
mlp_model = MLPRegressor(
activation="relu",
hidden_layer_sizes=[100, ]*20,
max_iter=1000,
early_stopping=True,
validation_fraction=0.2,
alpha=0.1,
beta_1=0.8,
learning_rate_init=0.0001
)
mlp_model.fit(x_train, y_train)
rf_model = RandomForestRegressor(max_features=3)
rf_model.fit(x_train, y_train)
# -
print("LR model", r2_score(y_test, model.predict(x_test)))
print("MLP model", r2_score(y_test, mlp_model.predict(x_test)))
print("RF model", r2_score(y_test, rf_model.predict(x_test)))
pd.to_pickle(df_full, "regression_data.pkl")
| data-creation/make_regression_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 2. Setup and Joining
# We assume that all data was reported and recorded correctly, and that no players share the same name and pick.
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
pd.set_option('display.max_columns', None)
player_avgs = pd.read_csv('player_avgs.csv')
nba_combine = pd.read_csv('nba_draft_combine_all_years.csv')
player_avgs.head()
nba_combine.head()
# We'll rename the `Draft pick` column in `nba_combine` to `Pk`, in order to facilitate joining the two datasets later:
nba_combine.rename(columns = {'Draft pick': 'Pk'}, inplace = True)
# Then, we need to clean the players' names in the `player_avgs` dataframe, so that the part after the slash is not included.
player_avgs.Player = [name.split('\\')[0] for name in player_avgs.Player]
player_avgs.Player[0:5]
# Now, we can merge the two datasets using `pd.merge`. We'll join them based on both player name and which pick they were, since we have a case where two players had the exact same name (<NAME>) but were drafted in different positions. There are no other duplicated names.
combine_careers_raw = pd.merge(nba_combine, player_avgs, on = ['Player', 'Pk'], how = 'inner')
# combine_careers_raw.to_csv('combine_careers.csv')
combine_careers_raw.head()
# the duplicated player name is <NAME>
combine_careers_raw.Player.value_counts().head()
# they were drafted at different picks
# joining on both player name and pick position allows us to avoid cross-product joins
combine_careers_raw[combine_careers_raw.Player == '<NAME>'][['Player', 'Pk']]
# ## 3. Basic Cleaning
# Let's look at the structure of our data. We see below that we have 368 rows and 44 columns, but we have quite a few missing values in several columns. There's also seems to be several redundant columns that could be removed (e.g., `Year_x` and `Year_y`).
combine_careers_raw.info()
# #### A. Removing Unnecessary Columns
# We immediately notice that the first column is unnecessary, so we can remove it.
combine_careers = combine_careers_raw.copy()
combine_careers.drop('Unnamed: 0', 1, inplace = True)
# We also notice that we have two year columns. Are there any discrepancies between the two? The answer is yes:
year_diff = combine_careers[['Year_x', 'Year_y']][combine_careers.Year_x != combine_careers.Year_y]
combine_careers.iloc[year_diff.index][['Player', 'Year_x', 'Year_y']]
# A quick look at these players' NBA combine results shows that `Year_y` is the correct column (for both [2010](https://www.nbadraft.net/2010-nba-draft-combine-official-measurements/) and [2017](https://www.nbadraft.net/2017-nba-draft-combine-measurements/) draftees). A brief check against the [official NBA Combine data](https://stats.nba.com/draft/combine-anthro/?SeasonYear=2016-17) shows that player statistics are still accurate, with some minor rounding differences. We'll go ahead and delete the `Year_x` column, and rename `Year_y` to `Year`.
combine_careers.drop('Year_x', 1, inplace = True)
combine_careers.rename(columns = {'Year_y': 'Year'}, inplace = True)
# Other columns we can delete: `Lg` and `Rd` (which stand for league and draft round, respectively). These are redundant because all players are in the NBA league, and because [draft round is redundant information if we know the draft pick](https://www.nba.com/nba-draft-lottery-explainer) (picks 1-30 are in round 1, and picks 31-60 are in round 2).
# all players are in the NBA
combine_careers.Lg.value_counts()
# all picks 1-30 are in Round 1, and all picks 31-60 are in Round 2
combine_careers.groupby(['Rd', 'Pk']).Player.count().unstack()
combine_careers.drop(['Lg', 'Rd'], 1, inplace = True)
# Let's take a final look at our (relatively clean) dataset. We'll deal with all the missing values next.
combine_careers.shape
combine_careers.head()
# #### B. Removing Players Who Never Played
# We can easily see that there are a number of players who participated in the draft combine, were drafted, but never played a single game.
combine_careers[combine_careers.G.isnull()].Player
# Let's go ahead and delete these rows, since they won't help us learn about how athleticism affects career.
combine_careers = combine_careers[combine_careers.G.notnull()]
# no more players who didn't play any games
len(combine_careers[combine_careers.G.isnull()].Player)
combine_careers.shape
# #### C. Simplifying Positions
# Players who have in-between positions may be denoted in two different ways (e.g., "G-F" and "F-G"). However, for our purposes, we don't need to worry about this distinction, so we'll combine "G-F" and "F-G", and then "F-C" and "C-F" (there's really no such thing as a "G-C" in the NBA, nor in our data).
combine_careers.loc[combine_careers['Pos'] == 'F-C', 'Pos'] = 'C-F'
combine_careers.loc[combine_careers['Pos'] == 'F-G', 'Pos'] = 'G-F'
combine_careers.Pos.value_counts()
# Let's write this dataset to CSV.
combine_careers.set_index('Player').to_csv('cc.csv')
| produce_base_dataset.ipynb |