code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.3
# language: julia
# name: julia-1.6
# ---
using Plots; pyplot()
using LaTeXStrings
using BasicBVP1D
using LinearAlgebra
default(xtickfont=font(14), ytickfont=font(14), guidefont=font(14),
legendfont=font(12), lw=2,ms=8)
# +
f = x-> (π)^2 * sin(π * x);
u_exact = x-> sin(π * x);
a = 0.;
b = 1.;
n = 7;
problem = SpectralBVPProblem(a, b, n, f, DirichletBC(0), DirichletBC(0));
assemble_system!(problem);
u = solve_bvp(problem);
plot(problem.x, u_exact.(problem.x), label="Exact")
scatter!(problem.x, real.(u),label="FD Soln")
# -
err = @. u - u_exact(problem.x);
@show norm(err, Inf)
| BasicBVP1D/notebooks/Spectral Dirichlet Test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # La data visualisation avec Python
# + slideshow={"slide_type": "fragment"}
import numpy as np
import pandas as pd
# + [markdown] slideshow={"slide_type": "fragment"}
# De nombreux packages sont disponibles en Python pour représenter des données
# de manière attractive.
#
# Le plus connu est Matplotlib et sert de base à de nombreux autres packages.
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Construction de graphiques avec matplotlib
#
# ### Afficher des graphiques
#
# Matplotlib est un package complet pour la data visualisation. Il est basé sur la
# construction de graphiques en utilisant des commandes simples.
#
# On commence par importer le module pyplot du package :
# + slideshow={"slide_type": "fragment"}
import matplotlib.pyplot as plt
# + [markdown] slideshow={"slide_type": "fragment"}
# Si vous travaillez dans un notebook Jupyter, il vous faudra ajouter une ligne au
# début de votre notebook afin de faire en sorte que le graphique apparaisse dans le
# notebook à la suite de votre code sous la forme d’une image statique :
# + slideshow={"slide_type": "fragment"}
# %matplotlib inline
# + [markdown] slideshow={"slide_type": "subslide"}
# Le principe du développement de graphiques avec Matplotlib est toujours le
# même : on peuple notre graphique petit à petit et, une fois le graphique terminé, on soumet le code et on affiche le graphique (avec plt.show() dans votre IDE classique ou en soumettant la cellule dans votre notebook).
#
# Par ailleurs, il existe deux manières de construire des graphiques avec Matplotlib.
#
# On peut enchaîner les traitements et finalement afficher un graphique :
# + slideshow={"slide_type": "subslide"}
# on crée une figure
plt.figure(figsize=(7,4))
# on commence par un sous-graphique
plt.subplot(2,2,1)
plt.plot(np.random.randn(100))
# on enchaine avec un second sous-graphique
plt.subplot(2,2,2)
plt.plot(np.random.random(size=100),"+",label="data1")
plt.plot(np.random.random(size=100),"+",label="data2")
plt.legend()
# on ajoute un troisième sous-graphique
plt.subplot(2,2,3)
plt.plot(np.random.randn(100),"r.")
# on finit avec le dernier sous-graphique
plt.subplot(2,2,4)
plt.plot(np.random.randn(100), "g--")
#plt.show() #si dans un IDE classique
# + [markdown] slideshow={"slide_type": "subslide"}
# On peut utiliser des objets et travailler dessus :
# + slideshow={"slide_type": "subslide"}
# on crée une figure avec 4 sous-graphiques
fig, ax=plt.subplots(2,2)
# on construit les 4 graphiques
ax[0,0].plot(np.random.randn(100))
ax[0,1].plot(np.random.random(size=100),"+",label="data1")
ax[0,1].plot(np.random.random(size=100),"+",label="data2")
ax[0,1].legend()
ax[1,0].plot(np.random.randn(100),"r.")
ax[1,1].plot(np.random.randn(100), "g--")
# on sauvegarde l'image dans un fichier
plt.savefig("./data/mes_graphiques2.jpg")
#plt.show() #si dans un IDE classique
# + [markdown] slideshow={"slide_type": "subslide"}
# Nous aurons tendance à privilégier la première approche qui est bien adaptée aux
# notebooks mais nous serons amenés à utiliser parfois la seconde notamment dans
# le cadre de graphiques plus interactifs.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Définir son premier graphique
#
# Si on désire dessiner un graphique simple, on pourra utiliser la fonction `plot()` :
#
# - On a différentes fonctions pour modifier le graphique :
# - `plt.title()`, `plt.xlabel()`, `plt.ylabel()`, `plt.axis`
#
#
# - En enchainant les `plt.plot()`, on peut avoir plusieurs courbes par graphique
# + slideshow={"slide_type": "subslide"}
plt.figure(figsize=(7,3))
plt.plot(np.random.randn(100))
# + [markdown] slideshow={"slide_type": "subslide"}
# On peut simplement ajouter des graphiques et des caractéristique à notre figure :
# + slideshow={"slide_type": "subslide"}
plt.figure(figsize=(7,3))
# on génère un premier graphique (rouge avec une ligne continue entre les points)
plt.plot(np.random.randn(100),'r-' )
# on génère un second graphique (bleu avec une ligne pointillée entre les points)
plt.plot(np.random.randn(100),'b--' )
# on ajoute des infos sur le titre, les noms des axes
plt.title("Evolution du prix des actions sur 100 jours")
plt.xlabel("Temps")
plt.ylabel("Valeur")
# + [markdown] slideshow={"slide_type": "slide"}
# ### Nuage de points avec plt.scatter
#
# On peut utiliser la fonction `plt.scatter()` pour dessiner un nuage de points.
# + slideshow={"slide_type": "fragment"}
# on génère des données
x=np.random.random(size=100)
y=np.random.random(size=100)
taille = np.random.random(size=100)*100
couleurs=np.random.random(size=100)*100
# + slideshow={"slide_type": "subslide"}
# s représente la taille des points
# c représente les couleurs (on peut avoir une seule couleur)
# cmap permet de fournir à Matplotlib une palette de couleurs
plt.scatter(x,y,s=taille, c=couleurs, cmap=plt.get_cmap("jet"))
plt.xlabel("X")
plt.ylabel("Y")
plt.title("Représentation de points aléatoires")
plt.colorbar()
# + [markdown] slideshow={"slide_type": "subslide"}
# **Exercice :**
#
# Utilisez les données AirBnB pour représenter les coordonnées géographiques sous forme de nuage de points
# -
# on commence par récupérer les données
listing=pd.read_csv("./data/airbnb.csv",index_col=0)
'review_scores_location'
plt.figure(figsize=(8,5))
plt.scatter("longitude","latitude",s=1, c="review_scores_location", data=listing,
cmap=plt.get_cmap("rainbow"))
plt.colorbar()
plt.title("Notes des appartements à Paris")
plt.axis('off')
plt.savefig("notes_airbnb_paris.jpg")
# + [markdown] slideshow={"slide_type": "slide"}
# ### La construction d’histogrammes
#
# Les histogrammes sont des outils de description des données extrêmement importants afin d’aider à déterminer la distribution sous-jacente à chaque variable quantitative.
#
# Si nous désirons créer deux histogrammes associés à deux variables ayant les mêmes échelles sur le même graphique, nous utilisons le code suivant :
# + slideshow={"slide_type": "fragment"}
# on génère les données
data1=np.random.randn(100000)+1
data2=np.random.randn(100000)
# + slideshow={"slide_type": "subslide"}
# 1er histogramme
plt.hist(data1, bins=50, color="blue", label="données 1", alpha=0.5)
# 2nd histogramme
plt.hist(data2, bins=50, color="red", label="données 2", alpha = 0.5)
plt.title("Histogrammes")
plt.ylabel("Fréquences")
plt.legend()
# + [markdown] slideshow={"slide_type": "subslide"}
# Dans ce code, on utilise *label=* pour donner un nom aux données.
#
# Le *alpha=* est utilisé afin d’afficher les graphiques en transparence et gère le degré d’opacité des graphiques.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Seaborn pour des représentations plus élaborées
# ### Utilisation de Seaborn
#
# Seaborn est un autre package intéressant pour la création de graphiques. Il est
# basé sur Matplotlib et en utilise les principes. Son principal intérêt réside dans la création de graphiques plus spécifiques en quelques lignes de code.
# + slideshow={"slide_type": "fragment"}
import seaborn as sns
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Le box-plot ou la boîte à moustaches
# + [markdown] slideshow={"slide_type": "fragment"}
# Un box plot (appelé aussi une boîte à moustache) est un graphique utilisé fréquemment pour l’exploration des données. Il permet de visualiser pour une variable ou pour un groupe d’individus le comportement global des individus.
# + [markdown] slideshow={"slide_type": "fragment"}
# Nous utiliserons ici des données de la région Île-de-France sur les communes de
# la (données open data Île-de-France, voir le début du chapitre 4 pour une description des données).
#
# On représente le box-plot du nombre de naissances par commune en fonction du département d’Île-de-France :
# + slideshow={"slide_type": "subslide"}
# on récupère les données
data_idf=pd.read_csv("./data/base-dpt.csv",sep=";")
sns.boxplot(data_idf["DEP"], data_idf["NAIS0914"]/ data_idf["P09_POP"])
plt.title("Box plot du nombre de naissances par habitant par département")
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Les pairplot() de Seaborn ou la matrice de graphiques
#
# Le pairplot de Seaborn est un outil très efficace pour représenter les variables
# quantitatives d’un DataFrame (à condition de ne pas trop en avoir).
#
# Un pairplot va construire une figure avec à l’intérieur une matrice de graphiques.
#
# Les éléments sur la diagonale sont des distplot et les éléments hors de la diagonale sont des nuages de points qui croisent les variables deux à deux.
#
# Si nous essayons de représenter ce graphique par département sur les données
# de la région Ile-de-France :
# + slideshow={"slide_type": "skip"}
sns.pairplot(data=data_idf, hue='DEP',
vars=['P14_POP', 'SUPERF','NAIS0914', 'DECE0914'],height=2,
plot_kws={"s": 25},markers=['x','o','v','^','<','+','>',"*"])
# + [markdown] slideshow={"slide_type": "slide"}
# Il existe bien d'autres fonctionnalités de Matplotlib et Seaborn, n'hésitez pas à consulter la documentation des ces packages :
# - https://matplotlib.org/
# - https://seaborn.pydata.org/
| 07_Data_visualisation_matplotlib_seaborn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
from sklearn.model_selection import cross_val_score
from collections import Counter
from sklearn.cluster import KMeans, AgglomerativeClustering
# +
DATA='ugrin2020-vehiculo-usado-multiclase/'
TRAIN=DATA+'train.csv'
TEST=DATA+'test.csv'
PREPROCESSED_DATA='preprocessed_data/'
RESULTS='results/'
test = pd.read_csv(TEST)
test_ids = test['id']
# Carga de datos ya procesados
X=np.load(PREPROCESSED_DATA+'binScale-unbalanced.npz')
train = X['arr_0']
class_labels = X['arr_1']
test = X['arr_2']
X.close()
# -
r=50
train.shape
test.shape
data=np.vstack((train,test))
data.shape
data=np.hstack(((data),np.zeros((data.shape[0],5))))
for i in range(train.shape[0]):
j=-label[i]
data[i][int(j)]=r
K=5
results = KMeans(n_clusters=K, random_state=25).fit(data)
cluster_labels=results.labels_
centroids=results.cluster_centers_
df=pd.DataFrame(data)
df['cluster']=cluster_labels
Counter(class_labels)
Counter(cluster_labels) # K-means
Counter(cluster_labels) # Ward
K=5
results = AgglomerativeClustering(distance_threshold=None, n_clusters=K).fit(data)
cluster_labels=results.labels_
| practica3/clustering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Programming Assignment
#
# ## Saving and loading models, with application to the EuroSat dataset
#
# ### Instructions
#
# In this notebook, you will create a neural network that classifies land uses and land covers from satellite imagery. You will save your model using Tensorflow's callbacks and reload it later. You will also load in a pre-trained neural network classifier and compare performance with it.
#
# Some code cells are provided for you in the notebook. You should avoid editing provided code, and make sure to execute the cells in order to avoid unexpected errors. Some cells begin with the line:
#
# `#### GRADED CELL ####`
#
# Don't move or edit this first line - this is what the automatic grader looks for to recognise graded cells. These cells require you to write your own code to complete them, and are automatically graded when you submit the notebook. Don't edit the function name or signature provided in these cells, otherwise the automatic grader might not function properly. Inside these graded cells, you can use any functions or classes that are imported below, but make sure you don't use any variables that are outside the scope of the function.
#
# ### How to submit
#
# Complete all the tasks you are asked for in the worksheet. When you have finished and are happy with your code, press the **Submit Assignment** button at the top of this notebook.
#
# ### Let's get started!
#
# We'll start running some imports, and loading the dataset. Do not edit the existing imports in the following cell. If you would like to make further Tensorflow imports, you should add them here.
# +
#### PACKAGE IMPORTS ####
# Run this cell first to import all required packages. Do not make any imports elsewhere in the notebook
import tensorflow as tf
from tensorflow.keras.preprocessing.image import load_img, img_to_array
from tensorflow.keras.models import Sequential, load_model
from tensorflow.keras.layers import Dense, Flatten, Conv2D, MaxPooling2D
from tensorflow.keras.callbacks import ModelCheckpoint, EarlyStopping
import os
import numpy as np
import pandas as pd
# If you would like to make further imports from tensorflow, add them here
# -
# 
#
# #### The EuroSAT dataset
#
# In this assignment, you will use the [EuroSAT dataset](https://github.com/phelber/EuroSAT). It consists of 27000 labelled Sentinel-2 satellite images of different land uses: residential, industrial, highway, river, forest, pasture, herbaceous vegetation, annual crop, permanent crop and sea/lake. For a reference, see the following papers:
# - Eurosat: A novel dataset and deep learning benchmark for land use and land cover classification. <NAME>, <NAME>, <NAME>, <NAME>. IEEE Journal of Selected Topics in Applied Earth Observations and Remote Sensing, 2019.
# - Introducing EuroSAT: A Novel Dataset and Deep Learning Benchmark for Land Use and Land Cover Classification. <NAME>, <NAME>, <NAME>. 2018 IEEE International Geoscience and Remote Sensing Symposium, 2018.
#
# Your goal is to construct a neural network that classifies a satellite image into one of these 10 classes, as well as applying some of the saving and loading techniques you have learned in the previous sessions.
# #### Import the data
#
# The dataset you will train your model on is a subset of the total data, with 4000 training images and 1000 testing images, with roughly equal numbers of each class. The code to import the data is provided below.
# +
# Run this cell to import the Eurosat data
def load_eurosat_data():
data_dir = 'data/'
x_train = np.load(os.path.join(data_dir, 'x_train.npy'))
y_train = np.load(os.path.join(data_dir, 'y_train.npy'))
x_test = np.load(os.path.join(data_dir, 'x_test.npy'))
y_test = np.load(os.path.join(data_dir, 'y_test.npy'))
return (x_train, y_train), (x_test, y_test)
(x_train, y_train), (x_test, y_test) = load_eurosat_data()
x_train = x_train / 255.0
x_test = x_test / 255.0
# -
# #### Build the neural network model
# You can now construct a model to fit to the data. Using the Sequential API, build your model according to the following specifications:
#
# * The model should use the input_shape in the function argument to set the input size in the first layer.
# * The first layer should be a Conv2D layer with 16 filters, a 3x3 kernel size, a ReLU activation function and 'SAME' padding. Name this layer 'conv_1'.
# * The second layer should also be a Conv2D layer with 8 filters, a 3x3 kernel size, a ReLU activation function and 'SAME' padding. Name this layer 'conv_2'.
# * The third layer should be a MaxPooling2D layer with a pooling window size of 8x8. Name this layer 'pool_1'.
# * The fourth layer should be a Flatten layer, named 'flatten'.
# * The fifth layer should be a Dense layer with 32 units, a ReLU activation. Name this layer 'dense_1'.
# * The sixth and final layer should be a Dense layer with 10 units and softmax activation. Name this layer 'dense_2'.
#
# In total, the network should have 6 layers.
# +
#### GRADED CELL ####
# Complete the following function.
# Make sure to not change the function name or arguments.
def get_new_model(input_shape):
"""
This function should build a Sequential model according to the above specification. Ensure the
weights are initialised by providing the input_shape argument in the first layer, given by the
function argument.
Your function should also compile the model with the Adam optimiser, sparse categorical cross
entropy loss function, and a single accuracy metric.
"""
model = Sequential([
Conv2D(16, (3, 3), padding='same', activation='relu', input_shape=input_shape, name='conv_1'),
Conv2D(8, (3, 3), padding='same', activation='relu', name='conv_2'),
MaxPooling2D((8, 8), name='pool_1'),
Flatten(name='flatten'),
Dense(32, activation='relu', name='dense_1'),
Dense(10, activation='softmax', name='dense_2')
])
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
return model
# -
# #### Compile and evaluate the model
# +
# Run your function to create the model
model = get_new_model(x_train[0].shape)
# +
# Run this cell to define a function to evaluate a model's test accuracy
def get_test_accuracy(model, x_test, y_test):
"""Test model classification accuracy"""
test_loss, test_acc = model.evaluate(x=x_test, y=y_test, verbose=0)
print('accuracy: {acc:0.3f}'.format(acc=test_acc))
# +
# Print the model summary and calculate its initialised test accuracy
model.summary()
get_test_accuracy(model, x_test, y_test)
# -
# #### Create checkpoints to save model during training, with a criterion
#
# You will now create three callbacks:
# - `checkpoint_every_epoch`: checkpoint that saves the model weights every epoch during training
# - `checkpoint_best_only`: checkpoint that saves only the weights with the highest validation accuracy. Use the testing data as the validation data.
# - `early_stopping`: early stopping object that ends training if the validation accuracy has not improved in 3 epochs.
# +
#### GRADED CELL ####
# Complete the following functions.
# Make sure to not change the function names or arguments.
def get_checkpoint_every_epoch():
"""
This function should return a ModelCheckpoint object that:
- saves the weights only at the end of every epoch
- saves into a directory called 'checkpoints_every_epoch' inside the current working directory
- generates filenames in that directory like 'checkpoint_XXX' where
XXX is the epoch number formatted to have three digits, e.g. 001, 002, 003, etc.
"""
checkpoint_path = 'checkpoints_every_epoch/checkpoint_' + str(epoch)
checkpoints = ModelCheckpoint(filepath=checkpoint_path,
save_freq='epoch',
save_weights_only=True,
verbose=1)
return checkpoints
def get_checkpoint_best_only():
"""
This function should return a ModelCheckpoint object that:
- saves only the weights that generate the highest validation (testing) accuracy
- saves into a directory called 'checkpoints_best_only' inside the current working directory
- generates a file called 'checkpoints_best_only/checkpoint'
"""
checkpoint_path = 'checkpoints_best_only/checkpoint'
checkpoints = ModelCheckpoint(filepath=checkpoint_path,
frequency='epoch',
monitor='val_accuracy',
save_weights_only=True,
save_best_only=True,
verbose=1)
return checkpoints
# +
#### GRADED CELL ####
# Complete the following function.
# Make sure to not change the function name or arguments.
def get_early_stopping():
"""
This function should return an EarlyStopping callback that stops training when
the validation (testing) accuracy has not improved in the last 3 epochs.
HINT: use the EarlyStopping callback with the correct 'monitor' and 'patience'
"""
callback = EarlyStopping(monitor='val_accuracy', patience=3)
return callback
# +
# Run this cell to create the callbacks
checkpoint_every_epoch = get_checkpoint_every_epoch()
checkpoint_best_only = get_checkpoint_best_only()
early_stopping = get_early_stopping()
# -
# #### Train model using the callbacks
#
# Now, you will train the model using the three callbacks you created. If you created the callbacks correctly, three things should happen:
# - At the end of every epoch, the model weights are saved into a directory called `checkpoints_every_epoch`
# - At the end of every epoch, the model weights are saved into a directory called `checkpoints_best_only` **only** if those weights lead to the highest test accuracy
# - Training stops when the testing accuracy has not improved in three epochs.
#
# You should then have two directories:
# - A directory called `checkpoints_every_epoch` containing filenames that include `checkpoint_001`, `checkpoint_002`, etc with the `001`, `002` corresponding to the epoch
# - A directory called `checkpoints_best_only` containing filenames that include `checkpoint`, which contain only the weights leading to the highest testing accuracy
# +
# Train model using the callbacks you just created
callbacks = [checkpoint_every_epoch, checkpoint_best_only, early_stopping]
model.fit(x_train, y_train, epochs=50, validation_data=(x_test, y_test), callbacks=callbacks)
# -
# #### Create new instance of model and load on both sets of weights
#
# Now you will use the weights you just saved in a fresh model. You should create two functions, both of which take a freshly instantiated model instance:
# - `model_last_epoch` should contain the weights from the latest saved epoch
# - `model_best_epoch` should contain the weights from the saved epoch with the highest testing accuracy
#
# _Hint: use the_ `tf.train.latest_checkpoint` _function to get the filename of the latest saved checkpoint file. Check the docs_ [_here_](https://www.tensorflow.org/api_docs/python/tf/train/latest_checkpoint).
# +
#### GRADED CELL ####
# Complete the following functions.
# Make sure to not change the function name or arguments.
def get_model_last_epoch(model):
"""
This function should create a new instance of the CNN you created earlier,
load on the weights from the last training epoch, and return this model.
"""
filename = tf.train.latest_checkpoint('checkpoints_every_epoch')
model.load_weights(filename)
return model
def get_model_best_epoch(model):
"""
This function should create a new instance of the CNN you created earlier, load
on the weights leading to the highest validation accuracy, and return this model.
"""
filename = tf.train.latest_checkpoint('checkpoints_best_only')
model.load_weights(filename)
return model
# +
# Run this cell to create two models: one with the weights from the last training
# epoch, and one with the weights leading to the highest validation (testing) accuracy.
# Verify that the second has a higher validation (testing) accuarcy.
model_last_epoch = get_model_last_epoch(get_new_model(x_train[0].shape))
model_best_epoch = get_model_best_epoch(get_new_model(x_train[0].shape))
print('Model with last epoch weights:')
get_test_accuracy(model_last_epoch, x_test, y_test)
print('')
print('Model with best epoch weights:')
get_test_accuracy(model_best_epoch, x_test, y_test)
# -
# #### Load, from scratch, a model trained on the EuroSat dataset.
#
# In your workspace, you will find another model trained on the `EuroSAT` dataset in `.h5` format. This model is trained on a larger subset of the EuroSAT dataset and has a more complex architecture. The path to the model is `models/EuroSatNet.h5`. See how its testing accuracy compares to your model!
# +
#### GRADED CELL ####
# Complete the following functions.
# Make sure to not change the function name or arguments.
def get_model_eurosatnet():
"""
This function should return the pretrained EuroSatNet.h5 model.
"""
model = load_model('models/EuroSatNet.h5')
return model
# +
# Run this cell to print a summary of the EuroSatNet model, along with its validation accuracy.
model_eurosatnet = get_model_eurosatnet()
model_eurosatnet.summary()
get_test_accuracy(model_eurosatnet, x_test, y_test)
# -
# Congratulations for completing this programming assignment! You're now ready to move on to the capstone project for this course.
| Getting Started with TensorFlow 2/Week 4/Week 4 Programming Assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Preprocessing Data
# - dealing with categorical features
# - handling missing data
# - encoding
# - pipelines
# - centering & scaling data
# ## Dealing with Categorical Features
# - sklearn: OneHotEncoder()
# - pandas: get_dummies()
# +
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split, GridSearchCV
from sklearn.linear_model import Ridge, Lasso
sns.set()
# -
cars = pd.read_csv('datasets/cars.csv')
cars.head()
cars_dummies = pd.get_dummies(cars, drop_first=True)
# +
X, y = cars_dummies.drop('mpg', axis=1), cars_dummies['mpg']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=.25)
ridge = Ridge(normalize=True)
params = {'alpha': np.linspace(0.1, 10, 15), 'solver': ['auto', 'svd', 'lsqr', 'cholesky', 'saga']}
ridge_cv = GridSearchCV(ridge, params, cv=5).fit(X_train, y_train)
print(f'Best parameters: {ridge_cv.best_params_}')
print(f'Best score: {ridge_cv.best_score_}')
# -
# best features
# +
gapminder = pd.read_csv('datasets/gapminder.csv')
# Create a boxplot of life expectancy per region
gapminder.boxplot('life', 'Region', rot=60)
plt.show()
# -
gapminder_dummies = pd.get_dummies(gapminder, drop_first=True)
gapminder_dummies.columns
# +
from sklearn.model_selection import cross_val_score
X, y = gapminder_dummies.drop('life', axis=1), gapminder_dummies['life']
# Instantiate a ridge regressor: ridge
ridge = Ridge(alpha=.5, normalize=True)
# Perform 5-fold cross-validation: ridge_cv
ridge_cv = cross_val_score(ridge, X, y, cv=5)
# Print the cross-validated scores
print(ridge_cv)
print(np.mean(ridge_cv))
# -
# # Handling Missing Data
# Missing data can be of different types:
# - 0
# - ?
# - -1
diabetes = pd.read_csv('datasets/diabetes.csv')
diabetes.info()
# +
diabetes.insulin.replace(0, np.nan, inplace=True)
diabetes.triceps.replace(0, np.nan, inplace=True)
diabetes.bmi.replace(0, np.nan, inplace=True)
diabetes.info()
# +
# imputer
from sklearn.impute import SimpleImputer
X = diabetes.drop('diabetes', axis=1)
y = diabetes['diabetes']
imp = SimpleImputer(strategy='mean')
imp.fit(X)
X = imp.transform(X)
# -
# ## Pipeline
# +
from sklearn.pipeline import make_pipeline
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import classification_report, confusion_matrix
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=.35, stratify=y)
imputer = SimpleImputer(strategy='mean')
logreg = LogisticRegression()
pipeline = make_pipeline(imputer, logreg)
pipeline.fit(X_train, y_train)
print(f'Train Score: {pipeline.score(X_train, y_train)}')
print(f'Test Score: {pipeline.score(X_test, y_test)}')
y_pred = pipeline.predict(X_test)
print(confusion_matrix(y_test, y_pred))
print(classification_report(y_test, y_pred))
# -
# # Centering & Scaling
# - standardization
#
red_wine = pd.read_csv('datasets/red_wine.csv', sep=';')
print(red_wine.info())
red_wine.head()
# +
from sklearn.preprocessing import scale
X, y = red_wine.drop('quality', axis=1), red_wine['quality']
X_scaled = scale(X)
print(np.mean(X_scaled), np.std(X_scaled))
# +
# scaling with pipeline
from sklearn.neighbors import KNeighborsClassifier
from sklearn.preprocessing import StandardScaler
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=.3, stratify=y)
pipeline = make_pipeline(StandardScaler(), KNeighborsClassifier())
pipeline.fit(X_train, y_train)
print('With Standardization')
print(classification_report(y_test, pipeline.predict(X_test)))
print('Without Standardization')
knn = KNeighborsClassifier().fit(X_train, y_train)
print(classification_report(y_test, knn.predict(X_test)))
# +
# gridsearchcv with pipeline
from sklearn.model_selection import GridSearchCV
from sklearn.pipeline import Pipeline
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=.3, stratify=y)
pipeline = Pipeline([('scaler', StandardScaler()), ('knn', KNeighborsClassifier())])
params = dict(knn__n_neighbors=np.arange(1, 50))
gridsearch = GridSearchCV(pipeline, params)
gridsearch.fit(X_train, y_train)
print(f'Best Parameters: {gridsearch.best_params_}')
print(f'Best Score: {gridsearch.best_score_}')
# +
from sklearn.svm import SVC
# Setup the pipeline
steps = [('scaler', StandardScaler()),
('SVM', SVC())]
pipeline = Pipeline(steps)
# Specify the hyperparameter space
parameters = {'SVM__C':[1, 10, 100],
'SVM__gamma':[0.1, 0.01]}
# Create train and test sets
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=.2, random_state=21)
# Instantiate the GridSearchCV object: cv
cv = GridSearchCV(pipeline, parameters, cv=3)
# Fit to the training set
cv.fit(X_train, y_train)
# Predict the labels of the test set: y_pred
y_pred = cv.predict(X_test)
# Compute and print metrics
print("Accuracy: {}".format(cv.score(X_test, y_test)))
print(classification_report(y_test, y_pred))
print("Tuned Model Parameters: {}".format(cv.best_params_))
# +
# gapminder with pipeline
from sklearn.linear_model import ElasticNet
X, y = gapminder.drop(['life', 'Region'], axis=1), gapminder['life']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=.35)
steps = [('imputation', SimpleImputer()), ('scaler', StandardScaler()), ('elasticnet', ElasticNet())]
pipeline = Pipeline(steps)
parameters = {'elasticnet__l1_ratio': np.linspace(0, 1, 30)}
gridsearch = GridSearchCV(pipeline, parameters, cv=3)
gridsearch.fit(X_train, y_train)
# Compute and print the metrics
r2 = gridsearch.score(X_test, y_test)
print("Tuned ElasticNet Alpha: {}".format(gridsearch.best_params_))
print("Tuned ElasticNet R squared: {}".format(r2))
# -
| datacamp_ml/ml_classification_regression/preprocessing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Application Prototype overview
#
# ## The application prototype for this project is built using HTML, Bootstrap, JavaScript and JSON.
#
# ### It provides following features:
# * Search General Practices by their id, post code or name
# * View prescription data trends for Jan-17 to Dec-17
# * Add GPs of interest to focus group for further analysis
# * In detailed view i.e. Trends and Focus Group, following information is shown:
# * General GP information:
# * Name, id, CCG name, Total Patient Size, Average Age, Index of Multiple Deprivition etc.
# * Patient Age Distibution as pie chart
# * GP Health Indicators information as radar chart
#
#
# ## The application code base following components.
#
# ### The codebase primarility contains HTML, JavaScript and data files (JSON and CSV)
#
# ### HTML Files:
# #### ClusterAnalysis.html - Cluster analysis page
# - single page with 3D cluster visualization
# - parameter selection
# - general information
# - costs information
#
# #### Home.html - Application main page
# - search GPs
# - options to manage and view focus group
#
# #### ViewFocusGroup.html - view focus groups for selected GPs
# #### ViewTrends.html - view trends for selected GP
# ### JavaScript files
# #### clusters.js
# * handles the code related to load of cluster data and visualization
# #### costs.js
# * handles the generation of costs plot. Reused in Trends and FocusGroup pages
#
# #### home.js
# * handles update HOME.html related code
# #### search.js
# * handles search opertions, communication with openprescribing.net API and update results
#
# #### trends.js
# * handles ViewTrends.html updates
#
# #### focusGroup.js
# * handles ViewFocusGroups.html updates
#
# #### drugGroups.js
# * handles drugGroups related code - retrieval of costs
#
# #### gpProfiles.js
# * handles client logic related to GPProfiles
#
# #### utils.js
# * Utility funcitons
| src/DataExploration/.ipynb_checkpoints/PrototypeCodeBase-Explanation-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Pedestrian Detection
import cv2
# +
cap = cv2.VideoCapture('pedestrians.avi')
p_cascade = cv2.CascadeClassifier('pedestrian.xml')
# +
while True:
ret, img = cap.read()
if (type(img) == type(None)):
break
gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
p = p_cascade.detectMultiScale(gray,1.3,2)
for(a,b,c,d) in p:
cv2.rectangle(img,(a,b),(a+c,b+d),(0,255,210),4)
cv2.imshow('video', img)
if cv2.waitKey(33) == 27:
break
cv2.destroyAllWindows()
# -
| Vehicles and Pedestrian Detection/Pedestrian Detection/pedestrian.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exploring Datasets with Python
# In this short demo we will analyse a given dataset from 1978, which contains information about politicians having affairs.
#
# To analyse it, we will use a [Jupyter Notebook](http://jupyter.org/), which is basically a *REPL++* for Python. Entering a command with shift executes the line and prints the result.
# To work with common files like CSV, JSON, Excel files etc., we will use [Pandas](http://pandas.pydata.org/), _an open source, BSD-licensed library providing high-performance, easy-to-use data structures and data analysis tools for the Python programming language_™. Let's import it!
# Our dataset is given as a CSV file. Pandas provides an easy way to read our file with `read_csv`. The path of the file to read is relative to our notebook file. The path can also be an URL, supporting HTTP, FTP and also S3 if your data is stored inside an AWS S3 Bucket!
# The first thing we will check is the size of our dataset. We can use `info()` to get the number of entries of each column.
# Now we know how many data is inside our file. Pandas is smart enough to parse the column titles by itself and estimate the data types of each column.
#
# You may be curious how the data looks like. Let's see by using `head()`, which will print the first 5 rows.
# We can access a column of our dataset by using bracket notation and the name of the column.
# For categorical features like `sex`, you can also get the distributions of each value by using `value_counts()`.
# But what about numerical values? It definitly makes no sense to count each distinct value. Therefore, we can use `describe()`.
# You can also access values like `mean` or `max` directly with the corrsponding methods. Let's see who is the oldest cheater!
# This works for the whole dataframe as well. Pandas knows which values are numerical based on the datatype and hides the categorical features for you.
# There is also an easy way to filter your dataset. Let's say we want to have a subset of our data containing only woman. This is also possible with the bracket notation!
# The above statement returns a new dataframe (not a copy, modifying this data will modify the original as well), which can be accessed like before. Let's see how the numerical distribution is for our females.
# We can also create new rows. Specify the new column name in brackets and provide a function to set the data. We will create a new column containing True or False, wheather or not the person is below 30.
# We can use this to normalize our columns with better values. Take for example `religious`. The number have the following meaning: 1 = not, 2 = mildly, 3 = fairly, 4 = strongly. We can easily replace them inline with the following code.
rel_meanings = ['not', 'mildly', 'fairly', 'strongly']
# This should be enought about Pandas. Let's get some visualisations!
# ## Visualize Data
# To visualize our data, we will use [Seaborn](https://seaborn.pydata.org), a Python visualization library based on matplotlib. It provides a high-level interface for drawing attractive statistical graphics. Let's import it.
# To see our charts directly in our notebook, we have to execute the following:
# %matplotlib inline
sns.set()
sns.set_context('talk')
# Seaborn together with Pandas makes it pretty easy to create charts to analyze our data. We can pass our Dataframes and Series directly into Seaborn methods. We will see how in the following sections.
# ### Univariate Plotting
# Let's start by visualizing the distribution of the age our our people. We can achieve this with a simple method called `distplot` by passing our series of ages as argument.
# The chart above calculates a kernel density as well. To get a real histogram, we have to disable the `kde` feature. We can also increase to number of buckets for our histogram by setting `bins` to 50.
# Interesting! The ages of the people in this dataset seem to end with two or seven.
#
# We can do the same for every numerical column, e.g. the years of marriage.
# The average age of our people is around 32, but the most people are married for more than 14 years!
# ### Bivariate Plotting
# Numbers get even more interesting when we can compare them to other numbers! Lets start comparing the number of years married vs the number of affairs. Seaborn provides us with a method called `jointplot` for this use case.
# To get a better feeling of how the number of affairs is affected by the number of years married, we can use a regression model by specifying `kind` as `reg`.
# We can also use a kernel to kompare the density of two columns against each other, e.g. `age` and `ym`.
# We can get an even better comparison by plotting everything vs everything! Seaborn provides this with the `pairplot` method.
# You won't see any special in this data. We need to separate them by some kind of criteria. We can use our categorical values to do this! Seaborn uses a parameter called `hue` to do this. Let's separate our data by `sex` first. To make things even more interesting, let's create a regression for every plot, too!
# To get even better separation, we can use `lmplot` to compare just the fields we need.
#
# Let's say we're interested in the number of affairs vs years married. We also whant to separate them by `sex`, `child` and `religious`. We will use `sns.lmplot(x="ym", y="nbaffairs", hue="sex", col="child", row="religious", data=affairs)` to achieve this.
# Here are some categorical plots to explore the dataset even further.
# We can also get the correlations between the values by using Pandas builtin method `corr()`.
# Feed these stats into Seaborns `heatmap` method will provide us with the visual representation.
| Exploring Datasets Yourself.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.12 64-bit
# language: python
# name: python3
# ---
# ## ME4: Decision Tree Classifiers
# #### Collaborators
#
# Notes: This homework can be done individually or by a troup of two. If this is done by a group, please write the names of students who work together. Make sure that each individual makes own submission(s).
#
# - <NAME>
#
# - <NAME>
#
#
# ### Setup
# First, let's import a few common modules, ensure MatplotLib plots figures inline and prepare a function to save the figures. We also check that Python 3.5 or later is installed (although Python 2.x may work, it is deprecated so we strongly recommend you use Python 3 instead), as well as Scikit-Learn ≥0.20.
# +
# Python ≥3.5 is required
import sys
assert sys.version_info >= (3, 5)
# Scikit-Learn ≥0.20 is required
import sklearn
assert sklearn.__version__ >= "0.20"
# Common imports
import numpy as np
import os
from sklearn.metrics import confusion_matrix
from sklearn.metrics import classification_report
# To plot pretty figures
# %matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rc('axes', labelsize=14)
mpl.rc('xtick', labelsize=12)
mpl.rc('ytick', labelsize=12)
# to make this notebook's output stable across runs
np.random.seed(42)
# Where to save the figures
PROJECT_ROOT_DIR = "."
CHAPTER_ID = "decision_trees"
IMAGES_PATH = os.path.join(PROJECT_ROOT_DIR, "images", CHAPTER_ID)
os.makedirs(IMAGES_PATH, exist_ok=True)
def save_fig(fig_id, tight_layout=True, fig_extension="png", resolution=300):
path = os.path.join(IMAGES_PATH, fig_id + "." + fig_extension)
print("Saving figure", fig_id)
if tight_layout:
plt.tight_layout()
plt.savefig(path, format=fig_extension, dpi=resolution)
# -
# ## Part 0
#
# - Read each cell of the examples below, run and check the outputs.
# ### Confusion matrix simple example 1 - binary classification
# +
y_true1 = [1, 0, 0, 1, 1, 0, 1, 1, 0]
y_pred1 = [1, 1, 0, 1, 1, 0, 1, 1, 1]
confusion_mat1 = confusion_matrix(y_true1, y_pred1)
print(confusion_mat1)
# +
# Print classification report
target_names2 = ['Class-0', 'Class-1']
result_metrics = classification_report(y_true1, y_pred1, target_names=target_names2)
print(result_metrics)
# -
# ### Confusion matrix simple example 2 - multiclass classification
# +
y_true2 = [1, 0, 0, 2, 1, 0, 3, 3, 3]
y_pred2 = [1, 1, 0, 2, 1, 0, 1, 3, 3]
confusion_mat2 = confusion_matrix(y_true2, y_pred2)
print(confusion_mat2)
# +
# Print classification report
target_names2 = ['Class-0', 'Class-1', 'Class-2', 'Class-3']
result_metrics2 = classification_report(y_true2, y_pred2, target_names=target_names2)
print(result_metrics2)
# -
# ### Decision Tree Classifier
#
# - dataset: iris dataset
# +
from IPython.display import Image
Image("images/iris.png")
# -
# #### Data visualization of the iris dataset before we start training and testing a model
#
# - iris.csv is stored in a local folder 'data'.
# +
import matplotlib.pyplot as plt
import pandas as pd
# read data from CSV file to dataframe
iris = pd.read_csv('./data/iris.csv')
# make sure you understand the type of the object
print(type(iris))
# check the top five and the botoom five data tuples
print(iris.head())
print(iris.tail())
# scatter matrix plot
pd.plotting.scatter_matrix(iris);
plt.figure()
# -
# ## Decision Trees
#
# - Read the details of decision tree classifier
#
# https://scikit-learn.org/stable/modules/generated/sklearn.tree.DecisionTreeClassifier.html
#
# - Check out the difference between model parameters and hyper parameters:
#
# https://towardsdatascience.com/model-parameters-and-hyperparameters-in-machine-learning-what-is-the-difference-702d30970f6
#
# ## A simple example of DT modeling
#
# - We first start the modeling without k-cross validation here but show step-by-step code segments how to train a model and test it.
# ### Load data
#
# - For the following code, we use sklearn.datasets package for loading a dataset instead of reading a data file stored on a local machine.
# +
from sklearn.datasets import load_iris
iris = load_iris()
# make sure that you understand the type of the object
print(type(iris))
print(iris)
# -
# ### Split the data to training and testing
# +
from sklearn.model_selection import train_test_split
X = iris.data # sepal length and width, petal length and width
y = iris.target
#print(X)
# split the data 70% for training, 30% for test data
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=3, test_size=0.20)
# -
# ## Training
# ### Learing using training data
#
# - use Gini index measure
#
# *** Notes: you can also use gain information (entropy) measure by setting criterion="entropy" in the model
# +
from sklearn.tree import DecisionTreeClassifier
tree_clf = DecisionTreeClassifier(max_depth=2, criterion='gini')
tree_clf.fit(X_train, y_train)
# -
# ## Testing
# ### Evaluating the model using testing data
# +
y_pred = tree_clf.predict(X_test)
y_pred
# -
# ## Model performance
# ### Confusion matrix
# +
# plot a confusion matrix
confusion_mat = confusion_matrix(y_test, y_pred)
print(confusion_mat)
# -
# ### Model performance summary
# +
# Print classification report
target_names = iris.target_names
result_metrics = classification_report(y_test, y_pred, target_names=target_names)
print(result_metrics)
# +
# you can access each class's metrics from result_metrics
# you can access each class's metrics from result_metrics
# output_dict should be set to True
result_metrics_dict = classification_report(y_test, y_pred, target_names=target_names, output_dict=True)
print(result_metrics_dict)
# an example that shows how to access the value of precision metric of class 'setosa'
print(result_metrics_dict['setosa']['precision'])
# -
# ### Draw a decision tree
#
# - You may need to install graphviz package!
# +
from graphviz import Source
from sklearn.tree import export_graphviz
export_graphviz(
tree_clf,
out_file=os.path.join(IMAGES_PATH, "iris_tree.dot"),
feature_names=iris.feature_names,
class_names=iris.target_names,
rounded=True,
filled=True
)
Source.from_file(os.path.join(IMAGES_PATH, "iris_tree.dot"))
# -
# ### Important features from a decision tree using gini index
#
# - Decision tree classifier
#
# https://scikit-learn.org/stable/modules/tree.html#classification
#
# - Metrics
#
# https://scikit-learn.org/stable/modules/model_evaluation.html
# plot important features
def plot_feature_importances(clf, feature_names):
c_features = len(feature_names)
plt.barh(range(c_features), clf.feature_importances_)
plt.xlabel("Feature importance")
plt.ylabel("Feature name")
plt.yticks(np.arange(c_features), feature_names)
# +
clf1 = DecisionTreeClassifier(criterion='gini').fit(X_train, y_train)
print('Accuracy of DT classifier on training set: {:.2f}'
.format(clf1.score(X_train, y_train)))
print('Accuracy of DT classifier on test set: {:.2f}'
.format(clf1.score(X_test, y_test)))
plt.figure(figsize=(8,4), dpi=60)
# call the function above
plot_feature_importances(clf1, iris.feature_names)
plt.show()
print('Feature importances: {}'.format(tree_clf.feature_importances_))
# +
clf2 = DecisionTreeClassifier(criterion='entropy').fit(X_train, y_train)
print('Accuracy of DT classifier on training set: {:.2f}'
.format(clf2.score(X_train, y_train)))
print('Accuracy of DT classifier on test set: {:.2f}'
.format(clf2.score(X_test, y_test)))
plt.figure(figsize=(8,4), dpi=60)
# call the function above
plot_feature_importances(clf2, iris.feature_names)
plt.show()
print('Feature importances: {}'.format(tree_clf.feature_importances_))
# -
# ## k-Cross Validation
#
# - using KFold function with freedom
# +
from sklearn.model_selection import KFold # import k-fold validation
kf = KFold(n_splits=3, random_state=None, shuffle=True) # Define the split - into 2 folds
kf.get_n_splits(X) # returns the number of splitting iterations in the cross-validator
print(kf)
# -
# ### Applying k-Cross Validation
# +
tree_clf = DecisionTreeClassifier(max_depth=2, random_state=42)
for train_index, test_index in kf.split(X):
print("TRAIN:", train_index, "TEST:", test_index)
X_train, X_test = X[train_index], X[test_index]
y_train, y_test = y[train_index], y[test_index]
tree_clf.fit(X_train, y_train)
y_pred = tree_clf.predict(X_test)
# Print classification report
target_names = iris.target_names
print(classification_report(y_test, y_pred, target_names=target_names))
# -
# # Predicting classes and class probabilities
# +
# Class0 (setona)
prob = tree_clf.predict_proba([[5.0, 3.4, 1.3, 0.2]])
# check predictions for different samples
# Class1 (versicolor)
#prob = tree_clf.predict_proba([[7.1, 3.1, 4.8, 1.4]])
# Class2 (virginica)
#prob = tree_clf.predict_proba([[6.4, 2.7, 4.9, 1.8]])
print(prob)
# +
# predict class1 (versicolor)
predicted = tree_clf.predict([[5.0, 3.4, 1.3, 0.2]])
print(predicted)
# -
# ## ME4
#
# ### Part 1
# ## Construct decision trees
#
# #### 1. Construct a decision tree using the following parameters
#
# - Use information gain (entropy) measure
# - Apply k=10 cross validation and print a summary of statistics (performance evaluation) for each fold
#
#
# #### 2. Compare the performance results with those of the decision tree using Gini index measure in the above example
#
# #### 3. For both trees, change the following parameters and observe the changes:
#
# - The depth of tree: currently max_depth=2 in the model training step. Change the depth 3, 4, 5 and check if this affects the overall results.
#
# - The k value for cross validation is currently set to 3. Change k value, k = 5, 7, 10 and check if this affects the overall results.
# ## Part 2
#
# 1. See DT examples at:
#
# https://www.kaggle.com/dmilla/introduction-to-decision-trees-titanic-dataset
#
# 2. Discuss about different ways to handle the following types of data for decision tree classification.
#
# - text data (strings): in the case a dataset includes non-numerical data.
#
# - continuous data like age, weight, income, etc.
#
kf = KFold(n_splits=10, random_state=None, shuffle=True)
kf.get_n_splits(X)
clf_entropy = DecisionTreeClassifier(criterion='entropy', max_depth=5)
for train_index, test_index in kf.split(X):
X_train, X_test = X[train_index], X[test_index]
y_train, y_test = y[train_index], y[test_index]
clf_entropy.fit(X_train, y_train)
y_pred = clf_entropy.predict(X_test)
# Print classification report
target_names = iris.target_names
result_metrics = classification_report(y_test, y_pred, target_names=target_names)
print(result_metrics)
kf = KFold(n_splits=10, random_state=None, shuffle=True)
kf.get_n_splits(X)
clf_gini = DecisionTreeClassifier(criterion='gini', max_depth=5)
for train_index, test_index in kf.split(X):
X_train, X_test = X[train_index], X[test_index]
y_train, y_test = y[train_index], y[test_index]
clf_gini.fit(X_train, y_train)
y_pred = clf_gini.predict(X_test)
# Print classification report
target_names = iris.target_names
result_metrics = classification_report(y_test, y_pred, target_names=target_names)
print(result_metrics)
# Changing the max_depth in the decision tree does change the overall results. On the entropy tree, we can see that the overall accuracy of the model increases as there are more depths. On the gini tree, we see that the middle splits are more accurate than the beginning and ending splits.
#
# When changing the number of k-fold splits by increasing from 3 to 5 to 7 to 10, it changes the overall results of the trees. On the entropy tree, we can see that the value in the precision column that from 3 to 5 splits there is not a lot of variation, but 7 and 10 have more dramatic changes. On the gini tree, we can see that as the number of k increases, the more accurate it becomes at the later folds.
#
# For both trees, when we averaged out the data they yielded the same results.
#
# In order to combat different types of data that may not be numeric, we could group them into categories represented by a numeric value. An example would be if the input data was countries, we could convert each country name to a corresponding number based on region and use the split point way of categorizing to choose which path to travel down the tree. To handle numeric data like age, weight, and income, we can use the same strategy of the split points (greater than or less than) to categorize the tree.
# ### Submission(s): Each individual student should make own submission.
#
# - Upload the notebook on your Git repo and provide an URL link in your summar.
#
# - Submit your summar to Canvas
#
# From this modeling exercise, I learned how decision trees work and how to combine them with the k-fold cross validation. I also learned how increasing the amount of folds and depths of the tree change the accuracy of the model and that when we average out the data between the gini and entropy trees, that they yield the same results. Something which I struggled with was figuring out what data to read for the tree in the exercise and I was really confused.
| ME4_DecisionTrees/decision_trees.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SageMath 7.3
# language: ''
# name: sagemath
# ---
# <h3>Diccionarios</h3>
# <p>Disponemos, en $"PALABRAS2.txt"$ de una lista de unas 120000 palabras en inglés, y queremos organizar la información en un diccionario de SAGE. Como el uso de este diccionario será para averiguar si una palabra pertenece o no al inglés nos conviene usar como claves los posibles grupos de tres letras iniciales y valores listas de palabras que comienzan por esas tres letras de la clave.</p>
def diccionario():
dicc = {}
infile = open("PALABRAS2.txt","r")
for palabra in infile.readlines():
if dicc.has_key(palabra[:3]):
dicc[palabra[:3]].append(palabra[:-1])
else:
dicc[palabra[:3]]=[palabra[:-1]]
infile.close()
return dicc
dicc = diccionario()
print dicc["GEO"]
print dicc["MAT"]
def comprueba(C):
if C in dicc[C[:3]]:
return True
else:
return False
time comprueba('GEOMETRY')
time comprueba('ABCDEFG')
# <h4>Usando listas</h4>
def listado():
L = []
infile = open("PALABRAS2.txt","r")
for palabra in infile.readlines():
L.append(palabra[:-1])
return L
L = listado()
len(L)
L[100]
def comprueba_L(C):
if C in L:
return True
else:
return False
time comprueba_L('GEOMETRY')
time comprueba_L('ABCDEFG')
# <p>Los tiempos que obtenemos son menores para el diccionario, pero conviene aplicar ambos métodos a un texto corto:</p>
def comprueba_texto(C):
cont = 0
L = C.split()
for palabra in L:
if comprueba(palabra) == True:
cont += 1
return cont
texto = "THROUGH THE USE ABSTRACTION AND LOGICAL REASONING MATHEMATICS DEVELOPED FROM COUNTING CALCULATION MEASUREMENT AND THE SYSTEMATIC STUDY THE SHAPES AND MOTIONS PHYSICAL OBJECTS PRACTICAL MATHEMATICS HAS BEEN HUMAN ACTIVITY FOR FAR BACK WRITTEN RECORDS EXIST RIGOROUS ARGUMENTS FIRST APPEARED GREEK MATHEMATICS MOST NOTABLY EUCLIDS ELEMENTS MATHEMATICS DEVELOPED RELATIVELY SLOW PACE UNTIL THE RENAISSANCE WHEN MATHEMATICAL INNOVATIONS INTERACTING WITH NEW SCIENTIFIC DISCOVERIES LED RAPID INCREASE THE RATE MATHEMATICAL DISCOVERY THAT CONTINUES THE PRESENT DAY"
time comprueba_texto(texto)
def comprueba_texto2(C):
cont = 0
L = C.split()
for palabra in L:
if comprueba_L(palabra) == True:
cont += 1
return cont
time comprueba_texto2(texto)
# <p>Usando un diccionario seguimos obteniendo resultados mejores. Por supuesto, en una situación real, con textos mucho más largos, la diferencia en tiempos debe ser grande.</p>
# <h4>Versión abstracta</h4>
# <p>Generamos una lista de $10^6$ enteros aleatorios en el intervalo $[100,10^7]$ que convertimos en cadenas de caracteres (palabras). Enteros aleatorios significa que todos los enteros del intervalo tienen,a priori, la misma probabilidad de aparecer en la lista $L$.</p>
L = [str(randint(100,10^7)) for muda in srange(10^6)]
# <h5>Con listas</h5>
def comprobador(L):
cont = 0
for muda in srange(10^3):
if muda%100 == 0:
print "Van otros 100"
if str(randint(1,10^7)) in L:
cont += 1
return cont
time comprobador(L)
# <h5>Con diccionarios</h5>
def diccionario2():
dicc = {}
for palabra in L:
if dicc.has_key(palabra[:3]):
dicc[palabra[:3]].append(palabra[:-1])
else:
dicc[palabra[:3]]=[palabra[:-1]]
return dicc
dicc2 = diccionario2()
def comprueba2(C):
if C in dicc2[C[:3]]:
return True
else:
return False
def comprobador2():
cont = 0
for muda in srange(10^3):
if muda%100 == 0:
print "Van otros 100"
if comprueba2(str(randint(100,10^7))) == True:
cont += 1
return cont
time comprobador2()
# <p>Vemos la ventaja enorme, cuando la cantidad de información a manejar es muy grande, de estructurar la información en un diccionario frente a la versión mucho más amorfa de una lista.</p>
# <p>Ésto no debería sorprendernos: el diccionario tiene unas $1000$ claves y cada una de ellas tendrá como valor una lista de alrededor de $1000$ enteros. Una búsqueda en el diccionario equivale, más o menos, a dos búsquedas en listas de longitud $1000$, mientras que una búsqueda en una lista de longitud $10^6$ es mucho más costosa porque en el peor caso hay que recorrer casi toda la lista buscando nuestro entero.</p>
max([len(dicc2[key]) for key in dicc2])
min([len(dicc2[key]) for key in dicc2])
| 2_Curso/Laboratorio/SAGE-noteb/IPYNB/CRIPT/49-PROGR-diccionario.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.7 64-bit (''Testing'': conda)'
# name: python3
# ---
# # [All you need to know about skip connections](https://www.analyticsvidhya.com/blog/2021/08/all-you-need-to-know-about-skip-connections/)
# Skip connections allow us to overcome the degradation problem/vanishing gradients problem where deep networks begin to perform worse than shallow networks. As the network gets deeper it become hard (read, impossible) for the weights to be propagated back to the earlier layers. Skip connections allow us to pass these weights forward from earlier layers to later layers allowing more fine grained detail to remain in the network at the end alongside more abstracted features.
# ## ResNet - A residual block
# First we implement a residual block using skip connections in PyTorch.
# +
# import required libraries
import torch
from torch import nn
import torch.nn.functional as F
import torchvision
# basic resdidual block of ResNet
# This is generic in the sense, it could be used for downsampling of features.
class ResidualBlock(nn.Module):
def __init__(self, in_channels, out_channels, stride=[1, 1], downsample=None):
"""
A basic residual block of ResNet
Parameters
----------
in_channels: Number of channels that the input have
out_channels: Number of channels that the output have
stride: strides in convolutional layers
downsample: A callable to be applied before addition of residual mapping
"""
super(ResidualBlock, self).__init__()
self.conv1 = nn.Conv2d(
in_channels, out_channels, kernel_size=3, stride=stride[0],
padding=1, bias=False
)
self.conv2 = nn.Conv2d(
out_channels, out_channels, kernel_size=3, stride=stride[1],
padding=1, bias=False
)
self.bn = nn.BatchNorm2d(out_channels)
self.downsample = downsample
def forward(self, x):
residual = x
# applying a downsample function before adding it to the output
if(self.downsample is not None):
residual = downsample(residual)
out = F.relu(self.bn(self.conv1(x)))
out = self.bn(self.conv2(out))
# note that adding residual before activation
out = out + residual
out = F.relu(out)
return out
# +
# downsample using 1*1 convolution
downsample = nn.Sequential(
nn.Conv2d(64, 128, kernel_size=1, stride=2, bias=False),
nn.BatchNorm2d(128)
)
# First 5 layers of ResNet34
resnet_blocks = nn.Sequential(
nn.Conv2d(3, 64, kernel_size=7, stride=2, padding=3, bias=False),
nn.MaxPool2d(kernel_size=2, stride=2),
ResidualBlock(64,64),
ResidualBlock(64,64),
ResidualBlock(64,128, stride=[2,1], downsample=downsample)
)
# checking the shape
inputs = torch.rand(1,3,100,100) # single 100*100 colour image
outputs = resnet_blocks(inputs)
print(outputs.shape) # shape would be (1,128,13,13)
# -
# one could also use pretrained weight of ResNet trained on ImageNet
resnet34 = torchvision.models.resnet34(pretrained=True)
| Skip Connections.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import requests
import json
from bs4 import BeautifulSoup as bs
import numpy as np
import sys
# +
page = 'https://www.springfieldspringfield.co.uk/episode_scripts.php?tv-show=game-of-thrones'
prefix = 'https://www.springfieldspringfield.co.uk/'
def get_episodes():
result = []
info = requests.get(page).text
soup = bs(info, 'html.parser')
season_list = soup.find_all('div', attrs={'class':'season-episodes'})
for s in season_list:
episode_list = s.find_all('a', attrs={'class':'season-episode-title'})
for e in episode_list:
href = e.get('href')
seid = href[-6:]
url = prefix + href
esoup = bs(requests.get(url).content, 'html.parser')
print(seid)
try:
etext = esoup.find('div', {'class':'scrolling-script-container'})
for br in etext.find_all("br"):
br.replace_with("\n")
etext = etext.text.strip().replace('\r', '')
ename = esoup.find('h3').text
for text in etext.split('\n'):
document = seid + '\t' + ename + '\t' + text
result.append(document)
# print(document)
except:
print('miss')
return result
# -
results = get_episodes()
with open('got.dat', 'w+') as f:
for r in results:
f.write(r)
f.write('\n')
| Src/Archived/got/nb.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Atlantic Hurricanes from 2005-2015
# 
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
data = pd.read_csv("https://people.sc.fsu.edu/~jburkardt/data/csv/hurricanes.csv")
data
df = data.iloc[:,np.r_[2:len(data.columns)]]
df.columns = ['2005', '2006', '2007', '2008', '2009', '2010', '2011', '2012', '2013', '2014', '2015']
yearly_totals = df.sum(axis=0)
yearly_totals
plt.title('# Hurricanes and Tropical Storms in the Atlantic')
yearly_totals.plot.bar()
| notebooks/jupyter-notebook-demo/analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import sys
import numpy as np
from numpy import linalg as LA
from pyDOE import *
import george
from george.kernels import ExpSquaredKernel
import matplotlib.pyplot as plt
#Create design in area and depth
design = lhs(6, samples=36)
idepth_lims=[24.8,26.8] # this is corresponds to the definiotion of idepth=ilim+1; idepth corresponds to median i-band depth, see https://github.com/LSSTDESC/ObsStrat/tree/static/static
area_lims=[7500,20000]
shear_m_lims=[0.002,0.02]
# Rachel: 0.01 is too optimistic 0.02 instead, 0.05 is too opti, choose 0.08
sigma_z_lims=[0.05,0.01] #Gaussian photo-z: sig_z is the 68% credible region around the Gaussian tomographic mean.
sig_delta_z_lims=[0.001,0.005] # Gaussian prior on delta_z
sig_sigma_z_lims=[0.003,0.006] # Gaussian prior on sigma_z
design[:,0]=area_lims[0]+(area_lims[1]-area_lims[0])*design[:,0]
design[:,1]=idepth_lims[0]+(idepth_lims[1]-idepth_lims[0])*design[:,1]
design[:,2]=shear_m_lims[0]+(shear_m_lims[1]-shear_m_lims[0])*design[:,2]
design[:,3]=sigma_z_lims[0]+(sigma_z_lims[1]-sigma_z_lims[0])*design[:,3]
design[:,4]=sig_delta_z_lims[0]+(sig_delta_z_lims[1]-sig_delta_z_lims[0])*design[:,4]
design[:,5]=sig_sigma_z_lims[0]+(sig_sigma_z_lims[1]-sig_sigma_z_lims[0])*design[:,5]
plt.scatter(design[:,0],design[:,1])
plt.title("Emulator Points")
plt.xlabel("Area in sqr deg")
plt.ylabel("median i-band depth")
plt.show()
# +
# Compute redshift distribution parameters based on fitting functions derived here
# https://github.com/LSSTDESC/ObsStrat/tree/static/static
# idepth=design[:,1]
# idepth[0]=24
#
# areavalues=design[:,0]
# LSSneff=37.8*10**(0.359*(ilim - 25))
# LSSz0 = 0.00627*(ilim-25)**2+0.0188*(ilim-25)+0.272
# LSSalpha = 0.0125*(ilim-25)**2-0.025*(ilim-25)+0.909
# WLneff = 4.33*(idepth-25)**2+7.03*(idepth-25)+10.49
# WLz0 = -0.0125*(idepth-25)+0.193
# WLalpha = -0.069*(idepth-25)+0.876
# new fitting formula from Xiao
area=design[:,0]
idepth=design[:,1]
shear_m=design[:,2]
design[:,3]=(design[:,3]-sigma_z_lims[0])/(sigma_z_lims[1]-sigma_z_lims[0])
sigma_z=design[:,3]
sig_delta_z=design[:,4]
sig_sigma_z= design[:,5]
#check with Xiao on how he exactly derived this
WLneff = 10.47*10**(0.3167*(idepth - 25.))
WLz0 = -0.0125*(idepth-25)+0.193
WLalpha = -0.069*(idepth-25)+0.876
ilim=idepth-1
LSSneff=37.8*10**(0.359*(ilim - 25))
LSSz0 = 0.00627*(ilim-25)**2+0.0188*(ilim-25)+0.272
LSSalpha = 0.0125*(ilim-25)**2-0.025*(ilim-25)+0.909
# -
# Tabulate design and derived parameters
from tabulate import tabulate
sigma_z=0.01+(0.1-0.01)*design[:,3]
table=zip(area,idepth,WLneff,WLz0,WLalpha,LSSneff,LSSz0,LSSalpha,shear_m,sigma_z,sig_delta_z,sig_sigma_z)
header=["area","idepth","WLneff","WLz0","WLalpha","LSSneff","LSSz0","LSSalpha","shear_m","sigma_z","sig_delta_z","sig_sigma_z"]
print tabulate(table, header)
# +
# Create redshift distribution files
zbin=300
zrange=[i for i in np.linspace(0.000001,3.5,zbin+1)]
zmin=zrange[0:300]
zmax=zrange[1:301]
zmid=[0.0]*zbin
for i in range(zbin):
zmid[i]=(zmin[i]+zmax[i])/2
nzWL=[0.0]*zbin
nzLSS=[0.0]*zbin
plt.figure(0)
plt.figure(1)
for i in range(len(design[:,1])):
f = open('zdistris/wl_redshift_model%d_WLz0%e_WLalpha%e.txt'%(i,WLz0[i],WLalpha[i]),'w')
g = open('zdistris/LSS_redshift_model%d_LSSz0%e_LSSalpha%e.txt'%(i,LSSz0[i],LSSalpha[i]),'w')
for j in range(zbin):
nzWL[j]=zmid[j]**2.0*np.exp(-((zmid[j]/WLz0[i])**WLalpha[i]))
nzLSS[j]=zmid[j]**2.0*np.exp(-((zmid[j]/LSSz0[i])**LSSalpha[i]))
f.write('%e %e %e %e\n'%(zmin[j],zmid[j],zmax[j],nzWL[j]))
g.write('%e %e %e %e\n'%(zmin[j],zmid[j],zmax[j],nzLSS[j]))
plt.figure(0)
plt.plot(zmid,nzWL)
plt.figure(1)
plt.plot(zmid,nzLSS)
f.close()
g.close()
plt.show()
# -
| emulator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python for Psychologists - Session 5
# ## Functions
# ### Function definition and return values
#
# We have already met functions like `len()` or `print()`. A function usually *does* something. The elements we pass to it are called *parameters* or *arguments*. Luckily, we are not dependent on Python providing exactly those functions we need. We can built our own functions! The syntax is very simple:
#
# ```python
# def function_name(argument1, argument2):
# # stuff I want
# # the function
# # to do
# return some_output # optional; see below...
# ```
#
# The indentation (with tab) is, again, very important!
#
#
# Define a function that adds its two arguments. Call the function "adder".
def adder(summand1, summand2):
result = summand1 + summand2
return result
adder(1,3)
# Let's see what happens if we comment out the last line (removing the `return()` command).
def adder(summand1, summand2):
result = summand1+summand2
# return result
adder(1,3)
# Apparently nothing! Although, internally, the variable "result" has been created, we have no access to it, because we didn't ask the function to return it to us.
#
# If a function returns something, hence, depends on how it has been defined. We already know two functions that do the same thing with one major difference being that one of them returns something and the other one does not:
# *my_list.pop()* and *my_list.remove()*
#
# Of course, we also know that the `pop()` function uses the index of the element, while `remove()` expects the content if what is to be removed. Let's ignore this difference for a second and concentrate on whether the functions return something or not.
#
# Let's take a look at this again.
# In one cell, create a list with numbers from 0 to 4 and use the `pop()` function to delete the second element from the list.
# In a second cell, look at "my_list" again.
#
# Then, in yet another cell, create the same list again and use the `remove()` function to remove the element with the value 1. Again, in a second cell, take a look at "my_list" again.
my_list=[0,1,2,3,4,5]
my_list.pop(1)
my_list
my_list=[0,1,2,3,4,5]
my_list.remove(1)
my_list
# What have we observed? Although only the `pop()` function returned a value (which in this case was the the value stored at the second position), *both functions did the exact same thing* independent of whether they had a return value or not.
# Another way of accessing a value from within a function is to use the `print()` function. However, when using `print()` we won't be able to assign the result to a new variable as the print-function simply prints the result to the output line. Copy the function "adder" from above and replace the "return..." line by a line that prints the result to the console/out cell.
def adder(summand1, summand2):
result = summand1+summand2
print(result)
adder(1,3)
# ### Keyword arguments and default values
#
# By setting default values, we can call functions without further specification of (some of) its parameters. We can add a default value to a function parameter like this:
#
# ```python
# def some_function(argument1 = "default_value"):
# # something the
# # function should
# # be doing
# return something # optional
# ```
#
# Like this, the function above needs no parameters passed to it and will work anyway, simply because we have given to it some value to work with in the definition.
#
# Try to define a function that takes a number and subtracts 5 from it. If asked to, the function can also subtract some other user-defined number. Give the function the name "subtractor". Then try the function without giving any further specification of the number being subtracted. Afterwards also try to modify the number subtracted from the input number when calling the function.
def subtractor(minuend, subtrahend = 5):
return minuend - subtrahend
subtractor(10)
subtractor(10,2)
# In the "subtractor" function above we had to enter the minuend and the subtrahend values in a certain order that corresponded to the order of the arguments given to the function during definition. Instead of passing arguments solely by their **position** they can also be passed by **keywords**, or by a mixture of both:
#
# ```python
# def some_function(arg1, arg2, arg3):
# return arg1+arg2+arg3
#
# some_function(value_arg1, arg3 = value_arg3, arg2 = value_arg2)
#
# ```
#
#
# In the code line above, the function `some_function()` was called with a positional argument, and then two keyword arguments.
#
# Take the subtractor function and pass it two keyword arguments in a different order than defined (i.e., first pass the keyword argument for the subtrahend, then pass the keyword argument for the minuend).
subtractor(subtrahend = 3, minuend = 10)
# ### Local and global variables
#
# Variable that have been defined inside a function are so called "local variables". "Local" refers to the fact, that variables that have been defined inside a function cannot be accessed from "outside", i.e., from the main part of the script outside the function definition. Let's illustrate this. Use your `adder()` function with two random integer inputs. Look at the function definition again. As you can see we defined a variable "return" inside the function. After execution of the function, however, we won't be able to access the variable "result". Try this!
result
# As you can see, there is no variable called `return` on the global level that we could access from outside the function. The only thing that `return` does for us is to return the *value* of the variable `result`, however, it doesn't return the object/variable per se.
#
# In principle, it is possible to define and change global variable inside a function definition for them to be accessible outside the function again. The keyword for this is `global`:
#
# ```python
# def fun():
# global some_global_variable_name
# # do something
# ```
#
# We won´t cover that here, but read [here](https://www.w3schools.com/python/gloss_python_global_variables.asp) for further information.
# ### Unlimited number of arguments
#
# If we want a function to be able to accept any number of arguments we can use `*args`. The name "args", by the way, is again variable and could be anything. Important is the preceding "`*`".
# +
def new_adder(*args):
x = 0
for a in args:
x = x + a
return x
new_adder(1,2,3,4,5)
# -
# We can also allow for an unlimited number of keyword arguments:
#
# +
def some_function(**kwargs):
for a, b in kwargs.items(): # kwargs is basically a dictionary
print(str(a) + " = " + str(b))
some_function(otter="cutest", dogs="extremely cute")
| session5/Session5_Functions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: jup3.7.3
# language: python
# name: jup3.7.3
# ---
# From https://github.com/MARDAScience/SediNet/blob/master/notebooks/SediNet_Continuous_SievedSand_sieveplus4Prcs.ipynb
#
# This dataset has only 400 images, but they are large ~(3000x2000). You can to predict the PX values and plot it similar to the notebook above
# %pylab inline
import pandas as pd
from tqdm.auto import tqdm
from pathlib import Path
import lasio
from pprint import pprint
import PIL
import shutil
from IPython.display import display
data_in = Path('../../data/raw/SediNet')
csv_file = pd.read_csv(data_in/'grain_size_sieved_sands'/'data_pescadero_sieve.csv')
csv_file
for i, row in csv_file.sample(5).iterrows():
break
im = PIL.Image.open(data_in/row['files'])
print(im.size, im, row)
im.resize((256, 256))
# TODO:
# - copy only ones we use to processed
# - break into smaller images
| notebooks/00 Data prep/09-mc-grainsize-sedinet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Analyzing public cloud and hybrid networks
#
# Public cloud and hybrid networks can be hard to debug and secure. Many of the standard tools (e.g., traceroute) do not work in the cloud setting, even though the types of paths that can emerge are highly complicated, depending on whether the endpoints are in the same region, different regions, across physical and virtual infrastructure, or whether public or private IPs of cloud instances are being used.
#
# At same time, the fast pace of evolution of these networks, where new subnets and instances can be spun up rapidly by different groups of people, creates a significant security risk. Network engineers need tools than can provide comprehensive guaratnees that services and applications are available and secure as intended at all possible times.
#
# In this notebook, we show how Batfish can predict and help debug network paths for cloud and hybrid networks and how it can guarantee that the network's availability and security posture is exactly as desired.
#
# 
# +
# Import packages
# %run startup.py
bf = Session(host="localhost")
def show_first_trace(trace_answer_frame):
"""
Prints the first trace in the answer frame.
In the presence of multipath routing, Batfish outputs all traces
from the source to destination. This function picks the first one.
"""
if len(trace_answer_frame) == 0:
print("No flows found")
else:
show("Flow: {}".format(trace_answer_frame.iloc[0]['Flow']))
show(trace_answer_frame.iloc[0]['Traces'][0])
def is_reachable(start_location, end_location, headers=None):
"""
Checks if the start_location can reach the end_location using specified packet headers.
All possible headers are considered if headers is None.
"""
ans = bf.q.reachability(pathConstraints=PathConstraints(startLocation=start_location,
endLocation=end_location),
headers=headers).answer()
return len(ans.frame()) > 0
# -
# ## Initializing the Network and Snapshot
#
# SNAPSHOT_PATH below can be updated to point to a custom snapshot directory. See [instructions](https://github.com/batfish/batfish/wiki/Packaging-snapshots-for-analysis) for how to package data for analysis.
# +
# Initialize a network and snapshot
NETWORK_NAME = "hybrid-cloud"
SNAPSHOT_NAME = "snapshot"
SNAPSHOT_PATH = "networks/hybrid-cloud"
bf.set_network(NETWORK_NAME)
bf.init_snapshot(SNAPSHOT_PATH, name=SNAPSHOT_NAME, overwrite=True)
# -
# The network snapshot that we just initialized is illustrated below. It has a datacenter network with the standard leaf-spine design on the left. Though not strictly necessary, we have included a host `srv-101` in this network to enable end-to-end analysis. The exit gateway of the datacenter connects to an Internet service provider (ASN 65200) that we call `isp_dc`.
#
# The AWS network is shown on the right. It is spread across two regions, `us-east-2` and `us-west-2`. Each region has two VPCs, one of which is meant to host Internet-facing services and the other is meant to host only private services. Subnets in the public-facing VPCs use an Internet gateway to send and receive traffic outside of AWS. The two VPCs in a region peer via a transit gateway. Each VPC has two subnets, and we have some instances running as well.
#
# The physical network connects to the AWS network using IPSec tunnels, shown in pink, between `exitgw` and the two transit gateways. BGP sessions run atop these tunnels to make endpoints aware of prefixes on the other side.
#
# You can view configuration files that we used [here](https://github.com/batfish/pybatfish/tree/master/jupyter_notebooks/networks/hybrid-cloud). The AWS portion of the configuration is in the aws_configs subfolder. It has JSON files obtained via AWS APIs. An example script that packages AWS data into a Batfish snapshot is [here](https://github.com/ratulm/bf-aws-snapshot).
#
# 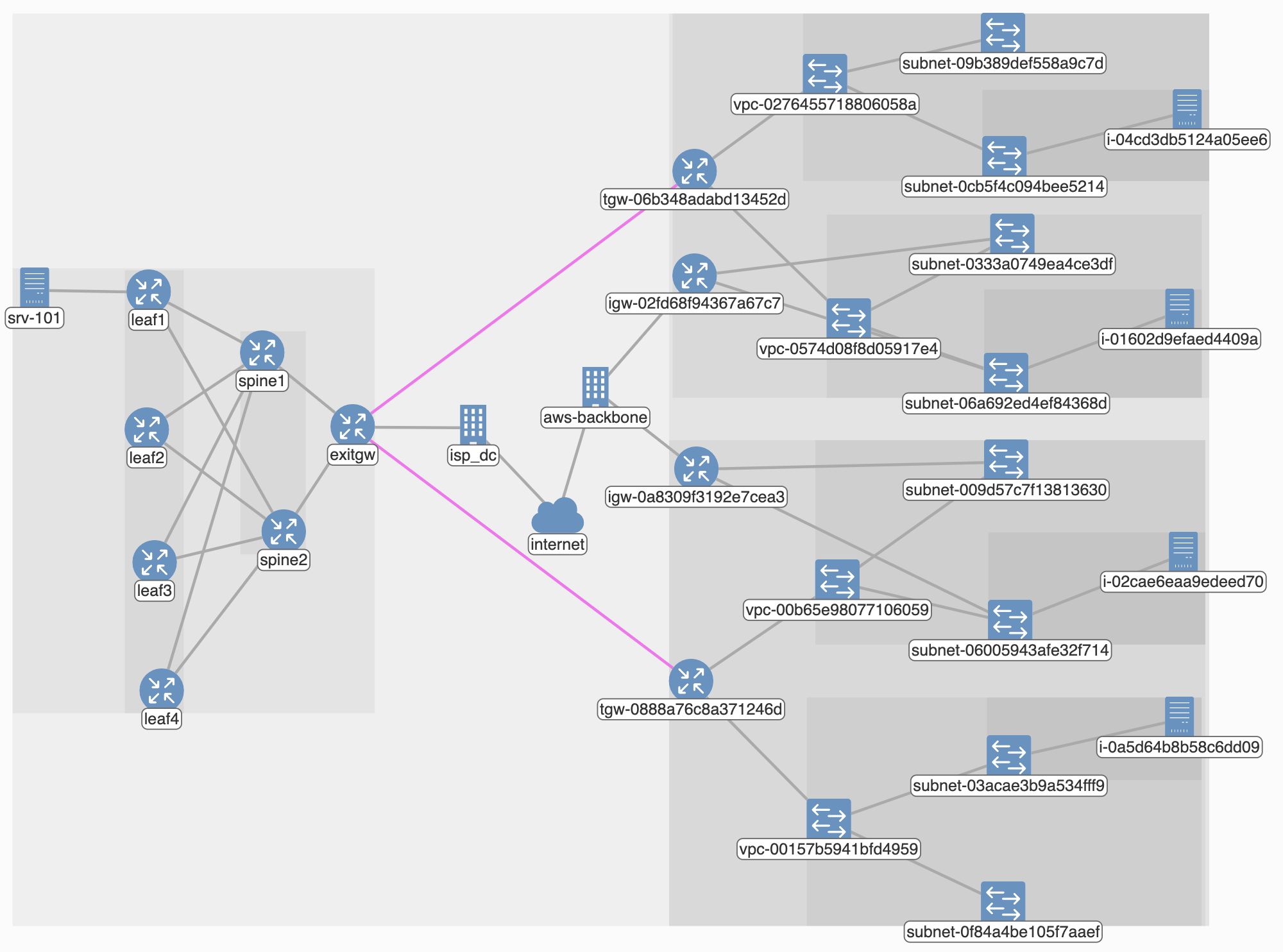
# ## Analyzing network paths
#
# Batfish can help analyze cloud and hybrid networks by showing how exactly traffic flows (or not) in the network, which can help debug and fix configuration errors. Batfish can also help ensure that the network is configured exactly as desired, with respect to reachability and security policies. We illustrate these types of analysis below.
#
# First, lets define a couple of maps to help with the analysis.
# +
#Instances in AWS in each region and VPC type (public, private)
hosts = {}
hosts["east2_private"] = "i-04cd3db5124a05ee6"
hosts["east2_public"] = "i-01602d9efaed4409a"
hosts["west2_private"] = "i-0a5d64b8b58c6dd09"
hosts["west2_public"] = "i-02cae6eaa9edeed70"
#Public IPs of instances in AWS
public_ips = {}
public_ips["east2_public"] = "192.168.127.12" # of i-01602d9efaed4409a
public_ips["west2_public"] = "172.16.17.32" # of i-02cae6eaa9edeed70
# -
# ### Paths across VPCs within an AWS region
#
# To see how traffic flows between two instances in the same region but across different VPCs, say, from `hosts["east2_private"]` to `hosts["east2_public"]`, we can run a traceroute query across them as follows.
#
# In the query below, we use the name of the instance as the destination for the traceroute. This makes Batfish pick the instance's private (i.e., non-Elastic) IP (`10.20.1.207`). It does not pick the public IP because that those IPs do not reside on instances but are used by the Internet gateway to NAT instance's traffic in and out of AWS (see [documentation](https://docs.aws.amazon.com/vpc/latest/userguide/VPC_Internet_Gateway.html)). If an instance has multiple private IPs, Batfish will pick one at random. To make Batfish use a specific IP, supply that IP as the argument to the `dstIps` parameter.
# traceroute between instances in the same region, using SSH
ans = bf.q.traceroute(startLocation=hosts["east2_private"],
headers=HeaderConstraints(dstIps=hosts["east2_public"],
applications="ssh")).answer()
show_first_trace(ans.frame())
# The trace above shows how traffic goes from `host["east2_private"]` to `host["east2_public"]` -- via the source subnet and VPC, then to the transit gateway, and finally to the destination VPC and subnet. Along the way, it also shows where the flow encounters security groups (at both instances) and network ACLs (at subnets). In this instance, all security groups and network ACLs permit this particular flow.
#
# This type of insight into traffic paths, which helps understand and debug network configuration, is difficult to obtain otherwise. Traceroutes on the live AWS network do not yield any information if the flow does not make it through, and do not show why or where a packet is dropped.
# ### Paths across AWS regions
#
# The traceroute query below shows paths across instances in two different regions.
# traceroute between instances across region using the destination's private IP
ans = bf.q.traceroute(startLocation=hosts["east2_public"],
headers=HeaderConstraints(dstIps=hosts["west2_public"],
applications="ssh")).answer()
show_first_trace(ans.frame())
# We see that such traffic does not reach the destination but instead is dropped by the AWS backbone (ASN 16509). This happens because, in our network, there is no (transit gateway or VPC) peering between VPCs in different regions. So, the source subnet is unaware of the address space of the destination subnet, which makes it use the default route that points to the Internet gateway (`igw-02fd68f94367a67c7`). The Internet gateway forwards the packet to `aws-backbone`, after NAT'ing its source IP. The packet is eventually dropped as it is using a private address as destination. Recall that using the instance name as destination amounts to using its private IP.
#
# The behavior is different if we use the public IP instead, as shown below.
# traceroute betwee instances across region using the destination's public IP
ans = bf.q.traceroute(startLocation=hosts["east2_public"],
headers=HeaderConstraints(dstIps=public_ips["west2_public"],
applications="ssh")).answer()
show_first_trace(ans.frame())
# This traceroute starts out like the previous one, up until the AWS backbone (`isp_16509`) -- from source subnet to the Internet gateway which forwards it to the backbone, after source NAT'ing the packet. The backbone carries it to the internet gateway in the destination region (`igw-0a8309f3192e7cea3`), and this gateway NATs the packet's destination from the public IP to the instance's private IP.
# ## Connectivity between DC and AWS
#
# A common mode to connect to AWS is using VPNs and BGP, that is, establish IPSec tunnels between exit gateways on the physical side and AWS gateways and run BGP on top of these tunnels to exchange prefixes. Incompatibility in either IPSec or BGP settings on the two sides means that connectivity between the DC and AWS will not work.
#
# Batfish can determine if the two sides are compatibly configured with respect to IPSec and BGP settings and if those sessions will come up.
#
# The query below lists the status of all IPSec sessions between the `exitgw` and AWS transit gateways (specified using the regular expression `^tgw-` that matches those node names). This filtering lets us ignore any other IPSec sessions that may exist in our network and focus on DC-AWS connectivity.
# show the status of all IPSec tunnels between exitgw and AWS transit gateways
ans = bf.q.ipsecSessionStatus(nodes="exitgw", remoteNodes="/^tgw-/").answer()
show(ans.frame())
# In the output above, we see all expected tunnels. Each transit gateways has two established sessions to `exitgw`. The default AWS behavior is to have two IPSec tunnels between gateways and physical nodes.
#
# Now that we know IPSec tunnels are working, we can check BGP sessions. The query below lists the status of all BGP sessions where one end is an AWS transit gateway.
# show the status of all BGP sessions between exitgw and AWS transit gateways
ans = bf.q.bgpSessionStatus(nodes="exitgw", remoteNodes="/^tgw-/").answer()
show(ans.frame())
# The output above shows that all BGP sessions are established as expected.
# ### Paths from the DC to AWS
#
# Finally, lets look at paths from the datacenter to AWS. The query below does that using the private IP of the public instance in us-east-2 region.
# traceroute from DC host to an instances using private IP
ans = bf.q.traceroute(startLocation="srv-101",
headers=HeaderConstraints(dstIps=hosts["east2_public"],
applications="ssh")).answer()
show_first_trace(ans.frame())
# We see that this traffic travels on the IPSec links between the datacenter's `exitgw` and the transit gateway in the destination region (`tgw-06b348adabd13452d`), and then makes it to the destination instance after making it successfully past the network ACL on the subnet node and the security group on the instance.
#
# A different path emerges if we use the public IP of the same instance, as shown below.
# traceroute from DC host to an instances using public IP
ans = bf.q.traceroute(startLocation="srv-101",
headers=HeaderConstraints(dstIps=public_ips["east2_public"],
applications="ssh")).answer()
show_first_trace(ans.frame())
# We now see that the traffic traverses the Internet via `isp_65200` and the Internet gateway (`igw-02fd68f94367a67c7`), which NATs the destination address of the packet from the public to the private IP.
# ## Evaluating the network's availability and security
#
# In addition to helping you understand and debug network paths, Batfish can also help ensure that the network is correctly configured with respect to its availability and security policies.
#
# As examples, the queries below evaluate which instances are or are not reachable from the Internet.
# +
# compute which instances are open to the Internet
reachable_from_internet = [key for (key, value) in hosts.items() if is_reachable("internet", value)]
print("\nInstances reachable from the Internet: {}".format(sorted(reachable_from_internet)))
# compute which instances are NOT open to the Internet
unreachable_from_internet = [key for (key, value) in hosts.items() if not is_reachable("internet", value)]
print("\nInstances NOT reachable from the Internet: {}".format(sorted(unreachable_from_internet)))
# -
# We see that Batfish correctly computes that the two instances in the public subnets are accessible from the Internet, and the other two are not.
#
# We can compare the answers produced by Batfish to what is expected based on network policy. This comparison can ensure that all instances that are expected to host public-facing services are indeed reachable from the Internet, and all instances that are expecpted to host private services are indeed not accessible from the Internet.
#
# We can similarly compute which instances are reachable from hosts in the datacenter, using the query like the following.
# compute which instances are reachable from data center
reachable_from_dc = [key for (key,value) in hosts.items() if is_reachable("srv-101", value)]
print("\nInstances reachable from the DC: {}".format(sorted(reachable_from_dc)))
# We see that all four instances are accessible from the datacenter host.
#
# Batfish allows a finer-grained evaluation of security policy as well. In our network, our intent is that the public instances should only allow SSH traffic. Let us see if this invariant actually holds.
tcp_non_ssh = HeaderConstraints(ipProtocols="tcp", dstPorts="!22")
reachable_from_internet_non_ssh = [key for (key, value) in hosts.items()
if is_reachable("internet", value, tcp_non_ssh)]
print("\nInstances reachable from the Internet with non-SSH traffic: {}".format(
sorted(reachable_from_internet_non_ssh)))
# We see that, against our policy, the public-facing instance allows non-SSH traffic. To see examples of such traffic, we can run the following query.
ans = bf.q.reachability(pathConstraints=PathConstraints(startLocation="internet",
endLocation=hosts["east2_public"]),
headers=tcp_non_ssh).answer()
show_first_trace(ans.frame())
# We thus see that our misconfigured public instance allows TCP traffic to port 3306 (MySQL).
#
# In this and earlier reachability queries, we are not specifying anything about the flow to Batfish. It automatically figures out that the flow from the Internet that can reach `hosts["east2_public"]` must have `192.168.127.12` as its destination address, which after NAT'ing becomes the private IP of the instance. Such exhaustive analysis over all possible header spaces is unique to Batfish, which makes it an ideal tool for comprehensive availability and security analysis.
#
# Batfish can also diagnose *why* certain traffic makes it past security groups and network ACLs. For example, we can run the testFilters question as below to reveal why the flow above made it past the security group on `hosts["east2_public"]`.
flow=ans.frame().iloc[0]['Flow'] # the rogue flow uncovered by Batfish above
ans = bf.q.testFilters(nodes=hosts["east2_public"],
filters="~INGRESS_ACL~eni-01997085076a9b98a",
headers=HeaderConstraints(srcIps=flow.srcIp,
dstIps="10.20.1.207", # destination IP after the NAT at Step 3 above
srcPorts=flow.srcPort,
dstPorts=flow.dstPort,
ipProtocols=flow.ipProtocol)).answer()
show(ans.frame())
# The "Trace" column shows that the flow was permitted because the security group "launch-wizard-1" has a matching rule called "Connectivity test." (Perhaps someone added this rule to test connectivity but forgot to remove it.)
#
# Such introspection capability is indispensable for complex security groups and network ACLs. See [this notebook](https://github.com/batfish/pybatfish/tree/master/jupyter_notebooks/Analyzing%20ACLs%20and%20Firewall%20Rules.ipynb) for a more detailed illustration of these capabilities of Batfish.
# ## Summary
#
# Batfish allows you to analyze, debug, and secure your cloud and hybrid networks. It can shed light on different types of traffic paths between different types of endpoints (e.g., intra-region, cross-region, across hybrid links), and it can reveal the detailed availability and security posture of the network.
#
# Want to learn more? Come find us on [Slack](https://join.slack.com/t/batfish-org/shared_invite/enQtMzA0Nzg2OTAzNzQ1LTcyYzY3M2Q0NWUyYTRhYjdlM2IzYzRhZGU1NWFlNGU2MzlhNDY3OTJmMDIyMjQzYmRlNjhkMTRjNWIwNTUwNTQ) or [GitHub](https://github.com/batfish/batfish)
| jupyter_notebooks/Analyzing public and hybrid cloud networks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Bag of Words Meets Bags of Popcorn
#
# 
#
# * https://www.kaggle.com/c/word2vec-nlp-tutorial
#
# ### [자연 언어 처리 - 위키백과, 우리 모두의 백과사전](https://ko.wikipedia.org/wiki/%EC%9E%90%EC%97%B0_%EC%96%B8%EC%96%B4_%EC%B2%98%EB%A6%AC)
#
# * 자연 언어 처리(自然言語處理) 또는 자연어 처리(自然語處理)는 인간이 발화하는 언어 현상을 기계적으로 분석해서 컴퓨터가 이해할 수 있는 형태로 만드는 자연 언어 이해 혹은 그러한 형태를 다시 인간이 이해할 수 있는 언어로 표현하는 제반 기술을 의미한다. (출처 : 위키피디아)
#
# ### 자연어처리(NLP)와 관련 된 캐글 경진대회
# * [Sentiment Analysis on Movie Reviews | Kaggle](https://www.kaggle.com/c/sentiment-analysis-on-movie-reviews)
# * [Toxic Comment Classification Challenge | Kaggle](https://www.kaggle.com/c/jigsaw-toxic-comment-classification-challenge)
# * [Spooky Author Identification | Kaggle](https://www.kaggle.com/c/spooky-author-identification)
#
#
# ## 튜토리얼 개요
# ### 파트1
# * 초보자를 대상으로 기본 자연어 처리를 다룬다.
#
# ### 파트2, 3
# * Word2Vec을 사용하여 모델을 학습시키는 방법과 감정분석에 단어 벡터를 사용하는 방법을 본다.
#
# * 파트3는 레시피를 제공하지 않고 Word2Vec을 사용하는 몇 가지 방법을 실험해 본다.
#
# * 파트3에서는 K-means 알고리즘을 사용해 군집화를 해본다.
#
# * 긍정과 부정 리뷰가 섞여있는 100,000만개의 IMDB 감정분석 데이터 세트를 통해 목표를 달성해 본다.
#
#
# ### 평가 - ROC 커브(Receiver-Operating Characteristic curve)
# * TPR(True Positive Rate)과 FPR(False Positive Rate)을 각각 x, y 축으로 놓은 그래프
# * 민감도 TPR
# - 1인 케이스에 대해 1로 예측한 비율
# - 암환자를 진찰해서 암이라고 진단함
# * 특이도 FPR
# - 0인 케이스에 대해 1로 잘못 예측한 비율
# - 암환자가 아닌데 암이라고 진단함
#
# * X, Y가 둘 다 [0, 1] 범위이고 (0, 0)에서 (1, 1)을 잇는 곡선이다.
# * ROC 커브의 및 면적이 1에 가까울 수록(왼쪽 꼭지점에 다가갈수록) 좋은 성능
#
# * 참고 :
# * [New Sight :: Roc curve, AUR(AUCOC), 민감도, 특이도](http://newsight.tistory.com/53)
# * [ROC의 AUC 구하기 :: 진화하자 - 어디에도 소속되지 않기](http://adnoctum.tistory.com/121)
# * [Receiver operating characteristic - Wikipedia](https://en.wikipedia.org/wiki/Receiver_operating_characteristic)
#
# ### Use Google's Word2Vec for movie reviews
# * 자연어 텍스트를 분석해서 특정단어를 얼마나 사용했는지, 얼마나 자주 사용했는지, 어떤 종류의 텍스트인지 분류하거나 긍정인지 부정인지에 대한 감정분석, 그리고 어떤 내용인지 요약하는 정보를 얻을 수 있다.
# * 감정분석은 머신러닝(기계학습)에서 어려운 주제로 풍자, 애매모호한 말, 반어법, 언어 유희로 표현을 하는데 이는 사람과 컴퓨터에게 모두 오해의 소지가 있다. 여기에서는 Word2Vec을 통한 감정분석을 해보는 튜토리얼을 해본다.
# * Google의 Word2Vec은 단어의 의미와 관계를 이해하는 데 도움
# * 상당수의 NLP기능은 nltk모듈에 구현되어 있는데 이 모듈은 코퍼스, 함수와 알고리즘으로 구성되어 있다.
# * 단어 임베딩 모형 테스트 : [Korean Word2Vec](http://w.elnn.kr/search/)
#
# ### BOW(bag of words)
# * 가장 간단하지만 효과적이라 널리쓰이는 방법
# * 장, 문단, 문장, 서식과 같은 입력 텍스트의 구조를 제외하고 각 단어가 이 말뭉치에 얼마나 많이 나타나는지만 헤아린다.
# * 구조와 상관없이 단어의 출현횟수만 세기 때문에 텍스트를 담는 가방(bag)으로 생각할 수 있다.
# * BOW는 단어의 순서가 완전히 무시 된다는 단점이 있다. 예를 들어 의미가 완전히 반대인 두 문장이 있다고 하다.
# - `it's bad, not good at all.`
# - `it's good, not bad at all.`
# * 위 두 문장은 의미가 전혀 반대지만 완전히 동일하게 반환된다.
# * 이를 보완하기 위해 n-gram을 사용하는 데 BOW는 하나의 토큰을 사용하지만 n-gram은 n개의 토큰을 사용할 수 있도록 한다.
#
# * [Bag-of-words model - Wikipedia](https://en.wikipedia.org/wiki/Bag-of-words_model)
# # 파트 1
#
# NLP는?
# NLP(자연어처리)는 텍스트 문제에 접근하기 위한 기술집합이다.
# 이 튜토리얼에서는 IMDB 영화 리뷰를 로딩하고 정제하고 간단한 BOW(Bag of Words) 모델을 적용하여 리뷰가 추천인지 아닌지에 대한 정확도를 예측한다.
#
#
# ### 시작하기 전에
# 이 튜토리얼은 파이썬으로 되어 있으며, NLP에 익숙하다면 파트2 로 건너뛰어도 된다.
# +
import pandas as pd
"""
header = 0 은 파일의 첫 번째 줄에 열 이름이 있음을 나타내며
delimiter = \t 는 필드가 탭으로 구분되는 것을 의미한다.
quoting = 3은 쌍따옴표를 무시하도록 한다.
"""
# QUOTE_MINIMAL (0), QUOTE_ALL (1),
# QUOTE_NONNUMERIC (2) or QUOTE_NONE (3).
# 레이블인 sentiment 가 있는 학습 데이터
train = pd.read_csv('data/labeledTrainData.tsv',
header=0, delimiter='\t', quoting=3)
# 레이블이 없는 테스트 데이터
test = pd.read_csv('data/testData.tsv',
header=0, delimiter='\t', quoting=3)
train.shape
# -
train.tail(3)
test.shape
test.tail()
train.columns.values
# 레이블인 'sentiment'가 없다. 이 데이터를 기계학습을 통해 예측한다.
test.columns.values
train.info()
train.describe()
train['sentiment'].value_counts()
# html 태그가 섞여있기 때문에 이를 정제해줄 필요가 있음
train['review'][0][:700]
# ### 데이터 정제 Data Cleaning and Text Preprocessing
# 기계가 텍스트를 이해할 수 있도록 텍스트를 정제해 준다.
#
# 신호와 소음을 구분한다. 아웃라이어데이터로 인한 오버피팅을 방지한다.
#
# 1. BeautifulSoup(뷰티풀숩)을 통해 HTML 태그를 제거
# 2. 정규표현식으로 알파벳 이외의 문자를 공백으로 치환
# 3. NLTK 데이터를 사용해 불용어(Stopword)를 제거
# 4. 어간추출(스테밍 Stemming)과 음소표기법(Lemmatizing)의 개념을 이해하고 SnowballStemmer를 통해 어간을 추출
#
#
# ### 텍스트 데이터 전처리 이해하기
#
# (출처 : [트위터 한국어 형태소 분석기](https://github.com/twitter/twitter-korean-text))
#
# **정규화 normalization (입니닼ㅋㅋ -> 입니다 ㅋㅋ, 샤릉해 -> 사랑해)**
#
# * 한국어를 처리하는 예시입니닼ㅋㅋㅋㅋㅋ -> 한국어를 처리하는 예시입니다 ㅋㅋ
#
# **토큰화 tokenization**
#
# * 한국어를 처리하는 예시입니다 ㅋㅋ -> 한국어Noun, 를Josa, 처리Noun, 하는Verb, 예시Noun, 입Adjective, 니다Eomi ㅋㅋKoreanParticle
#
# **어근화 stemming (입니다 -> 이다)**
#
# * 한국어를 처리하는 예시입니다 ㅋㅋ -> 한국어Noun, 를Josa, 처리Noun, 하다Verb, 예시Noun, 이다Adjective, ㅋㅋKoreanParticle
#
#
# **어구 추출 phrase extraction**
#
# * 한국어를 처리하는 예시입니다 ㅋㅋ -> 한국어, 처리, 예시, 처리하는 예시
#
# Introductory Presentation: [Google Slides](https://docs.google.com/presentation/d/10CZj8ry03oCk_Jqw879HFELzOLjJZ0EOi4KJbtRSIeU/)
# * 뷰티풀숩이 설치되지 않았다면 우선 설치해 준다.
# ```
# # !pip install BeautifulSoup4
# ```
# 설치 및 버전확인
# !pip show BeautifulSoup4
# +
from bs4 import BeautifulSoup
example1 = BeautifulSoup(train['review'][0], "html5lib")
print(train['review'][0][:700])
example1.get_text()[:700]
# -
# 정규표현식을 사용해서 특수문자를 제거
import re
# 소문자와 대문자가 아닌 것은 공백으로 대체한다.
letters_only = re.sub('[^a-zA-Z]', ' ', example1.get_text())
letters_only[:700]
# 모두 소문자로 변환한다.
lower_case = letters_only.lower()
# 문자를 나눈다. => 토큰화
words = lower_case.split()
print(len(words))
words[:10]
# ### 불용어 제거(Stopword Removal)
#
# 일반적으로 코퍼스에서 자주 나타나는 단어는 학습 모델로서 학습이나 예측 프로세스에 실제로 기여하지 않아 다른 텍스트와 구별하지 못한다. 예를들어 조사, 접미사, i, me, my, it, this, that, is, are 등 과 같은 단어는 빈번하게 등장하지만 실제 의미를 찾는데 큰 기여를 하지 않는다. Stopwords는 "to"또는 "the"와 같은 용어를 포함하므로 사전 처리 단계에서 제거하는 것이 좋다. NLTK에는 153 개의 영어 불용어가 미리 정의되어 있다. 17개의 언어에 대해 정의되어 있으며 한국어는 없다.
#
#
# ### NLTK data 설치
# * http://corazzon.github.io/nltk_data_install
import nltk
from nltk.corpus import stopwords
stopwords.words('english')[:10]
# stopwords 를 제거한 토큰들
words = [w for w in words if not w in stopwords.words('english')]
print(len(words))
words[:10]
# ### 스테밍(어간추출, 형태소 분석)
# 출처 : [어간 추출 - 위키백과, 우리 모두의 백과사전](https://ko.wikipedia.org/wiki/%EC%96%B4%EA%B0%84_%EC%B6%94%EC%B6%9C)
#
#
# * 어간 추출(語幹 抽出, 영어: stemming)은 어형이 변형된 단어로부터 접사 등을 제거하고 그 단어의 어간을 분리해 내는 것
# * "message", "messages", "messaging" 과 같이 복수형, 진행형 등의 문자를 같은 의미의 단어로 다룰 수 있도록 도와준다.
# * stemming(형태소 분석): 여기에서는 NLTK에서 제공하는 형태소 분석기를 사용한다. 포터 형태소 분석기는 보수적이고 랭커스터 형태소 분석기는 좀 더 적극적이다. 형태소 분석 규칙의 적극성 때문에 랭커스터 형태소 분석기는 더 많은 동음이의어 형태소를 생산한다. [참고 : 모두의 데이터 과학 with 파이썬(길벗)](http://www.gilbut.co.kr/book/bookView.aspx?bookcode=BN001787)
# 포터 스태머의 사용 예
stemmer = nltk.stem.PorterStemmer()
print(stemmer.stem('maximum'))
print("The stemmed form of running is: {}".format(stemmer.stem("running")))
print("The stemmed form of runs is: {}".format(stemmer.stem("runs")))
print("The stemmed form of run is: {}".format(stemmer.stem("run")))
# 랭커스터 스태머의 사용 예
from nltk.stem.lancaster import LancasterStemmer
lancaster_stemmer = LancasterStemmer()
print(lancaster_stemmer.stem('maximum'))
print("The stemmed form of running is: {}".format(lancaster_stemmer.stem("running")))
print("The stemmed form of runs is: {}".format(lancaster_stemmer.stem("runs")))
print("The stemmed form of run is: {}".format(lancaster_stemmer.stem("run")))
# 처리 전 단어
words[:10]
# +
from nltk.stem.snowball import SnowballStemmer
stemmer = SnowballStemmer('english')
words = [stemmer.stem(w) for w in words]
# 처리 후 단어
words[:10]
# -
# ### Lemmatization 음소표기법
#
# 언어학에서 음소 표기법 (또는 lemmatization)은 단어의 보조 정리 또는 사전 형식에 의해 식별되는 단일 항목으로 분석 될 수 있도록 굴절 된 형태의 단어를 그룹화하는 과정이다.
# 예를 들어 동음이의어가 문맥에 따라 다른 의미를 갖는데
# <pre>
# 1) *배*가 맛있다.
# 2) *배*를 타는 것이 재미있다.
# 3) 평소보다 두 *배*로 많이 먹어서 *배*가 아프다.
# </pre>
# 위에 있는 3개의 문장에 있는 "배"는 모두 다른 의미를 갖는다. <br/>
# 레마타이제이션은 이때 앞뒤 문맥을 보고 단어의 의미를 식별하는 것이다.
# 영어에서 meet는 meeting으로 쓰였을 때 회의를 뜻하지만 meet 일 때는 만나다는 뜻을 갖는데 그 단어가 명사로 쓰였는지 동사로 쓰였는지에 따라 적합한 의미를 갖도록 추출하는 것이다.
#
# * 참고 :
# - [Stemming and lemmatization](https://nlp.stanford.edu/IR-book/html/htmledition/stemming-and-lemmatization-1.html)
# - [Lemmatisation - Wikipedia](https://en.wikipedia.org/wiki/Lemmatisation)
# +
from nltk.stem import WordNetLemmatizer
wordnet_lemmatizer = WordNetLemmatizer()
print(wordnet_lemmatizer.lemmatize('fly'))
print(wordnet_lemmatizer.lemmatize('flies'))
words = [wordnet_lemmatizer.lemmatize(w) for w in words]
# 처리 후 단어
words[:10]
# -
# * 하지만 이 튜토리얼에서는 Stemming과 Lemmatizing을 소개만 해서 stemming 코드를 별도로 추가하였다.
#
#
# # 문자열 처리
# * 위에서 간략하게 살펴본 내용을 바탕으로 문자열을 처리해 본다.
def review_to_words( raw_review ):
# 1. HTML 제거
review_text = BeautifulSoup(raw_review, 'html.parser').get_text()
# 2. 영문자가 아닌 문자는 공백으로 변환
letters_only = re.sub('[^a-zA-Z]', ' ', review_text)
# 3. 소문자 변환
words = letters_only.lower().split()
# 4. 파이썬에서는 리스트보다 세트로 찾는게 훨씬 빠르다.
# stopwords 를 세트로 변환한다.
stops = set(stopwords.words('english'))
# 5. Stopwords 불용어 제거
meaningful_words = [w for w in words if not w in stops]
# 6. 어간추출
stemming_words = [stemmer.stem(w) for w in meaningful_words]
# 7. 공백으로 구분된 문자열로 결합하여 결과를 반환
return( ' '.join(stemming_words) )
clean_review = review_to_words(train['review'][0])
clean_review
# 첫 번째 리뷰를 대상으로 전처리 해줬던 내용을 전체 텍스트 데이터를 대상으로 처리한다.
# 전체 리뷰 데이터 수 가져오기
num_reviews = train['review'].size
num_reviews
# +
"""
clean_train_reviews = []
캐글 튜토리얼에는 range가 xrange로 되어있지만
여기에서는 python3를 사용하기 때문에 range를 사용했다.
"""
# for i in range(0, num_reviews):
# clean_train_reviews.append( review_to_words(train['review'][i]))
"""
하지만 위 코드는 어느 정도 실행이 되고 있는지 알 수가 없어서
5000개 단위로 상태를 찍도록 개선했다.
"""
# clean_train_reviews = []
# for i in range(0, num_reviews):
# if (i + 1)%5000 == 0:
# print('Review {} of {} '.format(i+1, num_reviews))
# clean_train_reviews.append(review_to_words(train['review'][i]))
"""
그리고 코드를 좀 더 간결하게 하기 위해 for loop를 사용하는
대신 apply를 사용하도록 개선
"""
# # %time train['review_clean'] = train['review'].apply(review_to_words)
"""
코드는 한 줄로 간결해 졌지만 여전히 오래 걸림
"""
# CPU times: user 1min 15s, sys: 2.3 s, total: 1min 18s
# Wall time: 1min 20s
# +
# 참고 : https://gist.github.com/yong27/7869662
# http://www.racketracer.com/2016/07/06/pandas-in-parallel/
from multiprocessing import Pool
import numpy as np
def _apply_df(args):
df, func, kwargs = args
return df.apply(func, **kwargs)
def apply_by_multiprocessing(df, func, **kwargs):
# 키워드 항목 중 workers 파라메터를 꺼냄
workers = kwargs.pop('workers')
# 위에서 가져온 workers 수로 프로세스 풀을 정의
pool = Pool(processes=workers)
# 실행할 함수와 데이터프레임을 워커의 수 만큼 나눠 작업
result = pool.map(_apply_df, [(d, func, kwargs)
for d in np.array_split(df, workers)])
pool.close()
# 작업 결과를 합쳐서 반환
return pd.concat(list(result))
# -
# %time clean_train_reviews = apply_by_multiprocessing(\
# train['review'], review_to_words, workers=4)
# %time clean_test_reviews = apply_by_multiprocessing(\
# test['review'], review_to_words, workers=4)
# ### 워드 클라우드
# - 단어의 빈도 수 데이터를 가지고 있을 때 이용할 수 있는 시각화 방법
# - 단순히 빈도 수를 표현하기 보다는 상관관계나 유사도 등으로 배치하는 게 더 의미 있기 때문에 큰 정보를 얻기는 어렵다.
# +
from wordcloud import WordCloud, STOPWORDS
import matplotlib.pyplot as plt
# # %matplotlib inline 설정을 해주어야지만 노트북 안에 그래프가 디스플레이 된다.
# %matplotlib inline
def displayWordCloud(data = None, backgroundcolor = 'white', width=800, height=600 ):
wordcloud = WordCloud(stopwords = STOPWORDS,
background_color = backgroundcolor,
width = width, height = height).generate(data)
plt.figure(figsize = (15 , 10))
plt.imshow(wordcloud)
plt.axis("off")
plt.show()
# -
# 학습 데이터의 모든 단어에 대한 워드 클라우드를 그려본다.
# %time displayWordCloud(' '.join(clean_train_reviews))
# 테스트 데이터의 모든 단어에 대한 워드 클라우드를 그려본다.
# %time displayWordCloud(' '.join(clean_test_reviews))
# 단어 수
train['num_words'] = clean_train_reviews.apply(lambda x: len(str(x).split()))
# 중복을 제거한 단어 수
train['num_uniq_words'] = clean_train_reviews.apply(lambda x: len(set(str(x).split())))
# 첫 번째 리뷰에
x = clean_train_reviews[0]
x = str(x).split()
print(len(x))
x[:10]
# +
import seaborn as sns
fig, axes = plt.subplots(ncols=2)
fig.set_size_inches(18, 6)
print('리뷰별 단어 평균 값 :', train['num_words'].mean())
print('리뷰별 단어 중간 값', train['num_words'].median())
sns.distplot(train['num_words'], bins=100, ax=axes[0])
axes[0].axvline(train['num_words'].median(), linestyle='dashed')
axes[0].set_title('리뷰별 단어 수 분포')
print('리뷰별 고유 단어 평균 값 :', train['num_uniq_words'].mean())
print('리뷰별 고유 단어 중간 값', train['num_uniq_words'].median())
sns.distplot(train['num_uniq_words'], bins=100, color='g', ax=axes[1])
axes[1].axvline(train['num_uniq_words'].median(), linestyle='dashed')
axes[1].set_title('리뷰별 고유한 단어 수 분포')
# -
# ### [Bag-of-words model - Wikipedia](https://en.wikipedia.org/wiki/Bag-of-words_model)
#
# 다음의 두 문장이 있다고 하자,
# ```
# (1) John likes to watch movies. Mary likes movies too.
# (2) John also likes to watch football games.
# ```
# 위 두 문장을 토큰화 하여 가방에 담아주면 다음과 같다.
#
# ```
# [
# "John",
# "likes",
# "to",
# "watch",
# "movies",
# "Mary",
# "too",
# "also",
# "football",
# "games"
# ]
# ```
#
# 그리고 배열의 순서대로 가방에서 각 토큰이 몇 번 등장하는지 횟수를 세어준다.
# ```
# (1) [1, 2, 1, 1, 2, 1, 1, 0, 0, 0]
# (2) [1, 1, 1, 1, 0, 0, 0, 1, 1, 1]
# ```
# => 머신러닝 알고리즘이 이해할 수 있는 형태로 바꿔주는 작업이다.
#
# 단어 가방을 n-gram을 사용해 bigram 으로 담아주면 다음과 같다.
# ```
# [
# "<NAME>",
# "likes to",
# "to watch",
# "watch movies",
# "Mary likes",
# "likes movies",
# "movies too",
# ]
# ```
# => 여기에서는 CountVectorizer를 통해 위 작업을 한다.
#
#
# ### 사이킷런의 CountVectorizer를 통해 피처 생성
# * 정규표현식을 사용해 토큰을 추출한다.
# * 모두 소문자로 변환시키기 때문에 good, Good, gOod이 모두 같은 특성이 된다.
# * 의미없는 특성을 많이 생성하기 때문에 적어도 두 개의 문서에 나타난 토큰만을 사용한다.
# * min_df로 토큰이 나타날 최소 문서 개수를 지정할 수 있다.
# +
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.pipeline import Pipeline
# 튜토리얼과 다르게 파라메터 값을 수정
# 파라메터 값만 수정해도 캐글 스코어 차이가 많이 남
vectorizer = CountVectorizer(analyzer = 'word',
tokenizer = None,
preprocessor = None,
stop_words = None,
min_df = 2, # 토큰이 나타날 최소 문서 개수
ngram_range=(1, 3),
max_features = 20000
)
vectorizer
# -
# 속도 개선을 위해 파이프라인을 사용하도록 개선
# 참고 : https://stackoverflow.com/questions/28160335/plot-a-document-tfidf-2d-graph
pipeline = Pipeline([
('vect', vectorizer),
])
# %time train_data_features = pipeline.fit_transform(clean_train_reviews)
train_data_features
train_data_features.shape
vocab = vectorizer.get_feature_names()
print(len(vocab))
vocab[:10]
# +
# 벡터화 된 피처를 확인해 봄
import numpy as np
dist = np.sum(train_data_features, axis=0)
for tag, count in zip(vocab, dist):
print(count, tag)
pd.DataFrame(dist, columns=vocab)
# -
pd.DataFrame(train_data_features[:10].toarray(), columns=vocab).head()
# ### [랜덤 포레스트 - 위키백과, 우리 모두의 백과사전](https://ko.wikipedia.org/wiki/%EB%9E%9C%EB%8D%A4_%ED%8F%AC%EB%A0%88%EC%8A%A4%ED%8A%B8)
# 랜덤 포레스트의 가장 핵심적인 특징은 임의성(randomness)에 의해 서로 조금씩 다른 특성을 갖는 트리들로 구성된다는 점이다. 이 특징은 각 트리들의 예측(prediction)들이 비상관화(decorrelation) 되게하며, 결과적으로 일반화(generalization) 성능을 향상시킨다. 또한, 임의화(randomization)는 포레스트가 노이즈가 포함된 데이터에 대해서도 강인하게 만들어 준다.
# +
from sklearn.ensemble import RandomForestClassifier
# 랜덤포레스트 분류기를 사용
forest = RandomForestClassifier(
n_estimators = 100, n_jobs = -1, random_state=2018)
forest
# -
# %time forest = forest.fit(train_data_features, train['sentiment'])
from sklearn.model_selection import cross_val_score
# %time score = np.mean(cross_val_score(\
# forest, train_data_features, \
# train['sentiment'], cv=10, scoring='roc_auc'))
# 위에서 정제해준 리뷰의 첫 번째 데이터를 확인
clean_test_reviews[0]
# 테스트 데이터를 벡터화 함
# %time test_data_features = pipeline.transform(clean_test_reviews)
test_data_features = test_data_features.toarray()
test_data_features
# 벡터화 된 단어로 숫자가 문서에서 등장하는 횟수를 나타낸다
test_data_features[5][:100]
# 벡터화 하며 만든 사전에서 해당 단어가 무엇인지 찾아볼 수 있다.
# vocab = vectorizer.get_feature_names()
vocab[8], vocab[2558], vocab[2559], vocab[2560]
# 테스트 데이터를 넣고 예측한다.
result = forest.predict(test_data_features)
result[:10]
# 예측 결과를 저장하기 위해 데이터프레임에 담아 준다.
output = pd.DataFrame(data={'id':test['id'], 'sentiment':result})
output.head()
output.to_csv('data/tutorial_1_BOW_{0:.5f}.csv'.format(score), index=False, quoting=3)
output_sentiment = output['sentiment'].value_counts()
print(output_sentiment[0] - output_sentiment[1])
output_sentiment
fig, axes = plt.subplots(ncols=2)
fig.set_size_inches(12,5)
sns.countplot(train['sentiment'], ax=axes[0])
sns.countplot(output['sentiment'], ax=axes[1])
# 첫 번째 제출을 할 준비가 되었다.
# 리뷰를 다르게 정리하거나 'Bag of Words' 표현을 위해 다른 수의 어휘 단어를 선택하거나 포터 스테밍 등을 시도해 볼 수 있다.
# 다른 데이터세트로 NLP를 시도해 보려면 로튼 토마토(Rotten Tomatoes)를 해보는 것도 좋다.
#
# * 로튼 토마토 데이터 셋을 사용하는 경진대회 : [Sentiment Analysis on Movie Reviews | Kaggle](https://www.kaggle.com/c/sentiment-analysis-on-movie-reviews)
# +
# 파마레터를 조정해 가며 점수를 조금씩 올려본다.
# uni-gram 사용 시 캐글 점수 0.84476
print(436/578)
# tri-gram 사용 시 캐글 점수 0.84608
print(388/578)
# 어간추출 후 캐글 점수 0.84780
print(339/578)
# 랜덤포레스트의 max_depth = 5 로 지정하고
# CountVectorizer의 tokenizer=nltk.word_tokenize 를 지정 후 캐글 점수 0.81460
print(546/578)
# 랜덤포레스트의 max_depth = 5 는 다시 None으로 변경
# CountVectorizer max_features = 10000개로 변경 후 캐글 점수 0.85272
print(321/578)
# CountVectorizer의 tokenizer=nltk.word_tokenize 를 지정 후 캐글 점수 0.85044
print(326/578)
# CountVectorizer max_features = 10000개로 변경 후 캐글 점수 0.85612
print(305/578)
# 0.85884
print(296/578)
print(310/578)
# -
| word2vec-nlp-tutorial/tutorial-part-1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 18659, "status": "ok", "timestamp": 1610382574744, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggpw7xw-lyk6u6l92QjpI7MlI7qjJuuciCpwrUd=s64", "userId": "03770692095188133952"}, "user_tz": -420} id="e4abR9zSaWNk" outputId="0c32d57f-a411-4648-8144-7db8ae862454"
# Mount Google Drive
from google.colab import drive # import drive from google colab
ROOT = "/content/drive" # default location for the drive
print(ROOT) # print content of ROOT (Optional)
drive.mount(ROOT) # we mount the google drive at /content/drive
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 1573, "status": "ok", "timestamp": 1610382829592, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggpw7xw-lyk6u6l92QjpI7MlI7qjJuuciCpwrUd=s64", "userId": "03770692095188133952"}, "user_tz": -420} id="rYQ5KUgrSClH" outputId="a4f91034-cf58-4cbc-bd3b-9a68557f9591"
# %cd "/content/drive/My Drive/Projects/quantum_image_classifier/PennyLane/Data Reuploading Classifier"
# + executionInfo={"elapsed": 5512, "status": "ok", "timestamp": 1610357980284, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggpw7xw-lyk6u6l92QjpI7MlI7qjJuuciCpwrUd=s64", "userId": "03770692095188133952"}, "user_tz": -420} id="gk5AKGKcYGOo"
# !pip install pennylane
from IPython.display import clear_output
clear_output()
# + id="GigSJusGbx1b"
import os
def restart_runtime():
os.kill(os.getpid(), 9)
restart_runtime()
# + executionInfo={"elapsed": 855, "status": "ok", "timestamp": 1610357982999, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggpw7xw-lyk6u6l92QjpI7MlI7qjJuuciCpwrUd=s64", "userId": "03770692095188133952"}, "user_tz": -420} id="HoLmJLkIX810"
# # %matplotlib inline
import matplotlib.pyplot as plt
from mpl_toolkits.axes_grid1 import make_axes_locatable
import numpy as np
# + [markdown] id="vZFNOwFXoY8N"
# # Loading Raw Data
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 3065, "status": "ok", "timestamp": 1610357986720, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggpw7xw-lyk6u6l92QjpI7MlI7qjJuuciCpwrUd=s64", "userId": "03770692095188133952"}, "user_tz": -420} id="IvdFsGCVof9g" outputId="57b0c866-93c0-45e7-a833-90119b85cf6c"
import tensorflow as tf
(x_train, y_train), (x_test, y_test) = tf.keras.datasets.mnist.load_data()
x_train = x_train[:, 0:27, 0:27]
x_test = x_test[:, 0:27, 0:27]
# + executionInfo={"elapsed": 1254, "status": "ok", "timestamp": 1610357989363, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggpw7xw-lyk6u6l92QjpI7MlI7qjJuuciCpwrUd=s64", "userId": "03770692095188133952"}, "user_tz": -420} id="c6zvGFvIoxAN"
x_train_flatten = x_train.reshape(x_train.shape[0], x_train.shape[1]*x_train.shape[2])/255.0
x_test_flatten = x_test.reshape(x_test.shape[0], x_test.shape[1]*x_test.shape[2])/255.0
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 996, "status": "ok", "timestamp": 1610357989364, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggpw7xw-lyk6u6l92QjpI7MlI7qjJuuciCpwrUd=s64", "userId": "03770692095188133952"}, "user_tz": -420} id="Rmj1dzaso00h" outputId="e6c46dd3-4962-4412-8c06-7ccc3a10679a"
print(x_train_flatten.shape, y_train.shape)
print(x_test_flatten.shape, y_test.shape)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 932, "status": "ok", "timestamp": 1610357989714, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggpw7xw-lyk6u6l92QjpI7MlI7qjJuuciCpwrUd=s64", "userId": "03770692095188133952"}, "user_tz": -420} id="d10VoIC6o5_I" outputId="0732b267-d0a4-47cd-c1b4-28acd63e5bf8"
x_train_0 = x_train_flatten[y_train == 0]
x_train_1 = x_train_flatten[y_train == 1]
x_train_2 = x_train_flatten[y_train == 2]
x_train_3 = x_train_flatten[y_train == 3]
x_train_4 = x_train_flatten[y_train == 4]
x_train_5 = x_train_flatten[y_train == 5]
x_train_6 = x_train_flatten[y_train == 6]
x_train_7 = x_train_flatten[y_train == 7]
x_train_8 = x_train_flatten[y_train == 8]
x_train_9 = x_train_flatten[y_train == 9]
x_train_list = [x_train_0, x_train_1, x_train_2, x_train_3, x_train_4, x_train_5, x_train_6, x_train_7, x_train_8, x_train_9]
print(x_train_0.shape)
print(x_train_1.shape)
print(x_train_2.shape)
print(x_train_3.shape)
print(x_train_4.shape)
print(x_train_5.shape)
print(x_train_6.shape)
print(x_train_7.shape)
print(x_train_8.shape)
print(x_train_9.shape)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 932, "status": "ok", "timestamp": 1610357990859, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggpw7xw-lyk6u6l92QjpI7MlI7qjJuuciCpwrUd=s64", "userId": "03770692095188133952"}, "user_tz": -420} id="snFw4LqepFOl" outputId="6fa81d41-9174-4053-b2db-c4933edfcab8"
x_test_0 = x_test_flatten[y_test == 0]
x_test_1 = x_test_flatten[y_test == 1]
x_test_2 = x_test_flatten[y_test == 2]
x_test_3 = x_test_flatten[y_test == 3]
x_test_4 = x_test_flatten[y_test == 4]
x_test_5 = x_test_flatten[y_test == 5]
x_test_6 = x_test_flatten[y_test == 6]
x_test_7 = x_test_flatten[y_test == 7]
x_test_8 = x_test_flatten[y_test == 8]
x_test_9 = x_test_flatten[y_test == 9]
x_test_list = [x_test_0, x_test_1, x_test_2, x_test_3, x_test_4, x_test_5, x_test_6, x_test_7, x_test_8, x_test_9]
print(x_test_0.shape)
print(x_test_1.shape)
print(x_test_2.shape)
print(x_test_3.shape)
print(x_test_4.shape)
print(x_test_5.shape)
print(x_test_6.shape)
print(x_test_7.shape)
print(x_test_8.shape)
print(x_test_9.shape)
# + [markdown] id="SAxUS6Lhp95g"
# # Selecting the dataset
#
# Output: X_train, Y_train, X_test, Y_test
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 832, "status": "ok", "timestamp": 1610357995835, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggpw7xw-lyk6u6l92QjpI7MlI7qjJuuciCpwrUd=s64", "userId": "03770692095188133952"}, "user_tz": -420} id="f--pX5Oto_XB" outputId="3a0814dc-fa71-4a5b-cc07-701a022af134"
X_train = np.concatenate((x_train_list[0][:200, :], x_train_list[1][:200, :]), axis=0)
Y_train = np.zeros((X_train.shape[0],), dtype=int)
Y_train[200:] += 1
X_train.shape, Y_train.shape
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 573, "status": "ok", "timestamp": 1610357996969, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggpw7xw-lyk6u6l92QjpI7MlI7qjJuuciCpwrUd=s64", "userId": "03770692095188133952"}, "user_tz": -420} id="W_SHH9e3rqwG" outputId="4f626106-a04f-4f44-d1f9-591c0fd1c96c"
X_test = np.concatenate((x_test_list[0][:500, :], x_test_list[1][:500, :]), axis=0)
Y_test = np.zeros((X_test.shape[0],), dtype=int)
Y_test[500:] += 1
X_test.shape, Y_test.shape
# + [markdown] id="LrebzTO1z-or"
# # Dataset Preprocessing
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 788, "status": "ok", "timestamp": 1610358000137, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggpw7xw-lyk6u6l92QjpI7MlI7qjJuuciCpwrUd=s64", "userId": "03770692095188133952"}, "user_tz": -420} id="tpIJBmVAz-os" outputId="066f3c86-cdcf-48f7-cfe7-b4e203f6a139"
X_train = X_train.reshape(X_train.shape[0], 27, 27)
X_test = X_test.reshape(X_test.shape[0], 27, 27)
X_train.shape, X_test.shape
# + [markdown] id="ockEle2Ez-os"
# # Quantum
# + executionInfo={"elapsed": 4679, "status": "ok", "timestamp": 1610358007092, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggpw7xw-lyk6u6l92QjpI7MlI7qjJuuciCpwrUd=s64", "userId": "03770692095188133952"}, "user_tz": -420} id="jP9aoKRGz-os"
import pennylane as qml
from pennylane import numpy as np
from pennylane.optimize import AdamOptimizer, GradientDescentOptimizer
qml.enable_tape()
from tensorflow.keras.utils import to_categorical
# Set a random seed
np.random.seed(2020)
# + executionInfo={"elapsed": 826, "status": "ok", "timestamp": 1610358054620, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggpw7xw-lyk6u6l92QjpI7MlI7qjJuuciCpwrUd=s64", "userId": "03770692095188133952"}, "user_tz": -420} id="BFo9kVhAz-ot"
# Define output labels as quantum state vectors
def density_matrix(state):
"""Calculates the density matrix representation of a state.
Args:
state (array[complex]): array representing a quantum state vector
Returns:
dm: (array[complex]): array representing the density matrix
"""
return state * np.conj(state).T
label_0 = [[1], [0]]
label_1 = [[0], [1]]
state_labels = [label_0, label_1]
# -
dm_labels = [density_matrix(state_labels[i]) for i in range(2)]
dm_labels
dm_labels[0]
# + executionInfo={"elapsed": 975, "status": "ok", "timestamp": 1610358057231, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggpw7xw-lyk6u6l92QjpI7MlI7qjJuuciCpwrUd=s64", "userId": "03770692095188133952"}, "user_tz": -420} id="fYmu1Jchz-ot"
n_qubits = 2 # number of class
dev_fc = qml.device("default.qubit", wires=n_qubits)
@qml.qnode(dev_fc)
def q_fc(params, inputs):
"""A variational quantum circuit representing the DRC.
Args:
params (array[float]): array of parameters
inputs = [x, y]
x (array[float]): 1-d input vector
y (array[float]): single output state density matrix
Returns:
float: fidelity between output state and input
"""
# layer iteration
for l in range(len(params[0])):
# qubit iteration
for q in range(n_qubits):
# gate iteration
for g in range(int(len(inputs)/3)):
qml.Rot(*(params[0][l][3*g:3*(g+1)] * inputs[3*g:3*(g+1)] + params[1][l][3*g:3*(g+1)]), wires=q)
return [qml.expval(qml.Hermitian(dm_labels[i], wires=[i])) for i in range(n_qubits)]
# + executionInfo={"elapsed": 859, "status": "ok", "timestamp": 1610358059994, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggpw7xw-lyk6u6l92QjpI7MlI7qjJuuciCpwrUd=s64", "userId": "03770692095188133952"}, "user_tz": -420} id="UNYRb5MCz-ot"
dev_conv = qml.device("default.qubit", wires=9)
@qml.qnode(dev_conv)
def q_conv(conv_params, inputs):
"""A variational quantum circuit representing the Universal classifier + Conv.
Args:
params (array[float]): array of parameters
x (array[float]): 2-d input vector
y (array[float]): single output state density matrix
Returns:
float: fidelity between output state and input
"""
# layer iteration
for l in range(len(conv_params[0])):
# RY layer
# height iteration
for i in range(3):
# width iteration
for j in range(3):
qml.RY((conv_params[0][l][3*i+j] * inputs[i, j] + conv_params[1][l][3*i+j]), wires=(3*i+j))
# entangling layer
for i in range(9):
if i != (9-1):
qml.CNOT(wires=[i, i+1])
return qml.expval(qml.PauliZ(0) @ qml.PauliZ(1) @ qml.PauliZ(2) @ qml.PauliZ(3) @ qml.PauliZ(4) @ qml.PauliZ(5) @ qml.PauliZ(6) @ qml.PauliZ(7) @ qml.PauliZ(8))
# -
a = np.zeros((2, 1, 9))
q_conv(a, X_train[0, 0:3, 0:3])
a = np.zeros((2, 1, 9))
q_fc(a, X_train[0, 0, 0:9])
# + executionInfo={"elapsed": 811, "status": "ok", "timestamp": 1610358062091, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggpw7xw-lyk6u6l92QjpI7MlI7qjJuuciCpwrUd=s64", "userId": "03770692095188133952"}, "user_tz": -420} id="zzVunJV1z-ou"
from keras import backend as K
# Alpha Custom Layer
class class_weights(tf.keras.layers.Layer):
def __init__(self):
super(class_weights, self).__init__()
w_init = tf.random_normal_initializer()
self.w = tf.Variable(
initial_value=w_init(shape=(1, 2), dtype="float32"),
trainable=True,
)
def call(self, inputs):
return (inputs * self.w)
# + executionInfo={"elapsed": 2804, "status": "ok", "timestamp": 1610358067284, "user": {"displayName": "<NAME>", "photoUrl": "<KEY>", "userId": "03770692095188133952"}, "user_tz": -420} id="6IozHMJhz-ou"
# Input image, size = 27 x 27
X = tf.keras.Input(shape=(27,27), name='Input_Layer')
# Specs for Conv
c_filter = 3
c_strides = 2
# First Quantum Conv Layer, trainable params = 18*L, output size = 13 x 13
num_conv_layer_1 = 1
q_conv_layer_1 = qml.qnn.KerasLayer(q_conv, {"conv_params": (2, num_conv_layer_1, 9)}, output_dim=(1), name='Quantum_Conv_Layer_1')
size_1 = int(1+(X.shape[1]-c_filter)/c_strides)
q_conv_layer_1_list = []
# height iteration
for i in range(size_1):
# width iteration
for j in range(size_1):
temp = q_conv_layer_1(X[:, 2*i:2*(i+1)+1, 2*j:2*(j+1)+1])
temp = tf.keras.layers.Reshape((1,))(temp)
q_conv_layer_1_list += [temp]
concat_layer_1 = tf.keras.layers.Concatenate(axis=1)(q_conv_layer_1_list)
reshape_layer_1 = tf.keras.layers.Reshape((size_1, size_1))(concat_layer_1)
# Second Quantum Conv Layer, trainable params = 18*L, output size = 6 x 6
num_conv_layer_2 = 1
q_conv_layer_2 = qml.qnn.KerasLayer(q_conv, {"conv_params": (2, num_conv_layer_2, 9)}, output_dim=(1), name='Quantum_Conv_Layer_2')
size_2 = int(1+(reshape_layer_1.shape[1]-c_filter)/c_strides)
q_conv_layer_2_list = []
# height iteration
for i in range(size_2):
# width iteration
for j in range(size_2):
temp = q_conv_layer_2(reshape_layer_1[:, 2*i:2*(i+1)+1, 2*j:2*(j+1)+1])
temp = tf.keras.layers.Reshape((1,))(temp)
q_conv_layer_2_list += [temp]
concat_layer_2 = tf.keras.layers.Concatenate(axis=1)(q_conv_layer_2_list)
reshape_layer_2 = tf.keras.layers.Reshape((size_2, size_2, 1))(concat_layer_2)
# Max Pooling Layer, output size = 9
max_pool_layer = tf.keras.layers.MaxPooling2D(pool_size=(2, 2), strides=None, name='Max_Pool_Layer')(reshape_layer_2)
reshape_layer_3 = tf.keras.layers.Reshape((9,))(max_pool_layer)
# Quantum FC Layer, trainable params = 18*L*n_class + 2, output size = 2
num_fc_layer = 1
q_fc_layer_0 = qml.qnn.KerasLayer(q_fc, {"params": (2, num_fc_layer, 9)}, output_dim=2)(reshape_layer_3)
# Alpha Layer
alpha_layer_0 = class_weights()(q_fc_layer_0)
model = tf.keras.Model(inputs=X, outputs=alpha_layer_0)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 39184, "status": "ok", "timestamp": 1610358109156, "user": {"displayName": "<NAME>", "photoUrl": "https://<KEY>", "userId": "03770692095188133952"}, "user_tz": -420} id="-kazExRcz-ov" outputId="11a34545-b4dd-425d-d246-8d0d260c9da8"
model(X_train[0:3, :, :])
# +
import keras.backend as K
# def custom_loss(y_true, y_pred):
# return K.sum(((y_true.shape[1]-2)*y_true+1)*K.square(y_true-y_pred))/len(y_true)
def custom_loss(y_true, y_pred):
loss = K.square(y_true-y_pred)
#class_weights = y_true*(weight_for_1-weight_for_0) + weight_for_0
#loss = loss * class_weights
return K.sum(loss)/len(y_true)
# + executionInfo={"elapsed": 1168, "status": "ok", "timestamp": 1610358110335, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggpw7xw-lyk6u6l92QjpI7MlI7qjJuuciCpwrUd=s64", "userId": "03770692095188133952"}, "user_tz": -420} id="wZ2e6t93z-ow"
opt = tf.keras.optimizers.Adam(learning_rate=0.1)
model.compile(opt, loss='mse', metrics=["accuracy"])
# +
cp_val_acc = tf.keras.callbacks.ModelCheckpoint(filepath="./Model/2_QConv2ent_QFC_valacc.hdf5",
monitor='val_accuracy', verbose=1, save_weights_only=True, save_best_only=True, mode='max')
cp_val_loss = tf.keras.callbacks.ModelCheckpoint(filepath="./Model/2_QConv2ent_QFC_valloss.hdf5",
monitor='val_loss', verbose=1, save_weights_only=True, save_best_only=True, mode='min')
# -
filepath = "./Model_2/2_QConv2ent_QFC_saved-model-{epoch:02d}.hdf5"
checkpoint = tf.keras.callbacks.ModelCheckpoint(filepath, monitor='val_acc', verbose=1,
save_weights_only=True, save_best_only=False, mode='auto')
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 23954180, "status": "ok", "timestamp": 1610382086131, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggpw7xw-lyk6u6l92QjpI7MlI7qjJuuciCpwrUd=s64", "userId": "03770692095188133952"}, "user_tz": -420} id="GKW8hv-9z-ow" outputId="a004b2f4-c3e3-4aa1-cef5-0d28d21a6d57"
H = model.fit(X_train, to_categorical(Y_train), epochs=10, batch_size=32, initial_epoch=0,
validation_data=(X_test, to_categorical(Y_test)), verbose=1,
callbacks=[checkpoint, cp_val_acc, cp_val_loss])
# -
# history for 10 different weights set
H.history
model.summary()
# fix ent
H.history
# fix ent
model.weights
# # Exploring the results
# +
X_train = np.concatenate((x_train_list[0][:20, :], x_train_list[1][:20, :]), axis=0)
Y_train = np.zeros((X_train.shape[0],), dtype=int)
Y_train[20:] += 1
X_train.shape, Y_train.shape
# +
X_test = np.concatenate((x_test_list[0][:20, :], x_test_list[1][:20, :]), axis=0)
Y_test = np.zeros((X_test.shape[0],), dtype=int)
Y_test[20:] += 1
X_test.shape, Y_test.shape
# +
X_train = X_train.reshape(X_train.shape[0], 27, 27)
X_test = X_test.reshape(X_test.shape[0], 27, 27)
X_train.shape, X_test.shape
# -
# ## First Layer
# +
qconv_1_weights = np.array([[[ 1.6704171 , -0.37210035, -1.4763509 , -0.09573141,
-1.4124248 , -0.5384489 , -1.5873678 , -0.4178921 ,
0.86764395]],
[[ 0.07841697, 0.06181896, -0.11658821, 0.08088297,
-0.22606611, -0.47507533, 0.20372277, 0.43287253,
-0.4662246 ]]])
qconv_1_weights.shape
# +
# Input image, size = 27 x 27
X = tf.keras.Input(shape=(27,27), name='Input_Layer')
# Specs for Conv
c_filter = 3
c_strides = 2
# First Quantum Conv Layer, trainable params = 18*L, output size = 13 x 13
num_conv_layer_1 = 1
q_conv_layer_1 = qml.qnn.KerasLayer(q_conv, {"conv_params": (2, num_conv_layer_1, 9)}, output_dim=(1), name='Quantum_Conv_Layer_1')
size_1 = int(1+(X.shape[1]-c_filter)/c_strides)
q_conv_layer_1_list = []
# height iteration
for i in range(size_1):
# width iteration
for j in range(size_1):
temp = q_conv_layer_1(X[:, 2*i:2*(i+1)+1, 2*j:2*(j+1)+1])
temp = tf.keras.layers.Reshape((1,))(temp)
q_conv_layer_1_list += [temp]
concat_layer_1 = tf.keras.layers.Concatenate(axis=1)(q_conv_layer_1_list)
reshape_layer_1 = tf.keras.layers.Reshape((size_1, size_1))(concat_layer_1)
qconv1_model = tf.keras.Model(inputs=X, outputs=reshape_layer_1)
# -
qconv1_model(X_train[0:1])
qconv1_model.get_layer('Quantum_Conv_Layer_1').set_weights([qconv_1_weights])
qconv1_model.weights
# +
preprocessed_img_train = qconv1_model(X_train)
preprocessed_img_test = qconv1_model(X_test)
data_train = preprocessed_img_train.numpy().reshape(-1, 13*13)
np.savetxt('./2_QConv2ent_QFC-Filter1_Image_Train.txt', data_train)
data_test = preprocessed_img_test.numpy().reshape(-1, 13*13)
np.savetxt('./2_QConv2ent_QFC-Filter1_Image_Test.txt', data_test)
print(data_train.shape, data_test.shape)
# -
# ## Second Layer
# +
qconv_2_weights = np.array([[[ 2.531285 , -0.53907233, 1.2279214 , -0.04873934,
-1.9625113 , -0.31179625, 1.4537607 , -0.3272767 ,
1.5368826 ]],
[[ 0.8165296 , 0.3640772 , 1.7864715 , 0.5245018 ,
-1.6975542 , -0.21682522, 2.097541 , 0.49968052,
2.2059586 ]]])
qconv_2_weights.shape
# +
# Input image, size = 27 x 27
X = tf.keras.Input(shape=(27,27), name='Input_Layer')
# Specs for Conv
c_filter = 3
c_strides = 2
# First Quantum Conv Layer, trainable params = 18*L, output size = 13 x 13
num_conv_layer_1 = 1
q_conv_layer_1 = qml.qnn.KerasLayer(q_conv, {"conv_params": (2, num_conv_layer_1, 9)}, output_dim=(1), name='Quantum_Conv_Layer_1')
size_1 = int(1+(X.shape[1]-c_filter)/c_strides)
q_conv_layer_1_list = []
# height iteration
for i in range(size_1):
# width iteration
for j in range(size_1):
temp = q_conv_layer_1(X[:, 2*i:2*(i+1)+1, 2*j:2*(j+1)+1])
temp = tf.keras.layers.Reshape((1,))(temp)
q_conv_layer_1_list += [temp]
concat_layer_1 = tf.keras.layers.Concatenate(axis=1)(q_conv_layer_1_list)
reshape_layer_1 = tf.keras.layers.Reshape((size_1, size_1))(concat_layer_1)
# Second Quantum Conv Layer, trainable params = 18*L, output size = 6 x 6
num_conv_layer_2 = 1
q_conv_layer_2 = qml.qnn.KerasLayer(q_conv, {"conv_params": (2, num_conv_layer_2, 9)}, output_dim=(1), name='Quantum_Conv_Layer_2')
size_2 = int(1+(reshape_layer_1.shape[1]-c_filter)/c_strides)
q_conv_layer_2_list = []
# height iteration
for i in range(size_2):
# width iteration
for j in range(size_2):
temp = q_conv_layer_2(reshape_layer_1[:, 2*i:2*(i+1)+1, 2*j:2*(j+1)+1])
temp = tf.keras.layers.Reshape((1,))(temp)
q_conv_layer_2_list += [temp]
concat_layer_2 = tf.keras.layers.Concatenate(axis=1)(q_conv_layer_2_list)
reshape_layer_2 = tf.keras.layers.Reshape((size_2, size_2, 1))(concat_layer_2)
qconv2_model = tf.keras.Model(inputs=X, outputs=reshape_layer_2)
# -
qconv2_model(X_train[0:1])
qconv2_model.get_layer('Quantum_Conv_Layer_1').set_weights([qconv_1_weights])
qconv2_model.get_layer('Quantum_Conv_Layer_2').set_weights([qconv_2_weights])
qconv2_model.weights
# +
preprocessed_img_train = qconv2_model(X_train)
preprocessed_img_test = qconv2_model(X_test)
data_train = preprocessed_img_train.numpy().reshape(-1, 6*6)
np.savetxt('./2_QConv2ent_QFC-Filter2_Image_Train.txt', data_train)
data_test = preprocessed_img_test.numpy().reshape(-1, 6*6)
np.savetxt('./2_QConv2ent_QFC-Filter2_Image_Test.txt', data_test)
print(data_train.shape, data_test.shape)
# -
# ## Quantum States
X_train.shape, X_test.shape
# +
q_fc_weights = np.array([[[ 0.3726357 , -1.0686979 , 0.24734715, 2.5706375 ,
3.4816115 , -0.3167716 , 1.2021828 , 0.19082105,
0.0209959 ]],
[[-0.213224 , 1.235254 , 2.298314 , 2.2815244 ,
-1.1340573 , 0.8286312 , 0.6926544 , -1.0297745 ,
0.14662777]]])
q_fc_weights.shape
# -
pred_train = model.predict(X_train)
pred_test = model.predict(X_test)
np.argmax(pred_train, axis=1)
np.argmax(pred_test, axis=1)
# +
# Input image, size = 27 x 27
X = tf.keras.Input(shape=(27,27), name='Input_Layer')
# Specs for Conv
c_filter = 3
c_strides = 2
# First Quantum Conv Layer, trainable params = 18*L, output size = 13 x 13
num_conv_layer_1 = 1
q_conv_layer_1 = qml.qnn.KerasLayer(q_conv, {"conv_params": (2, num_conv_layer_1, 9)}, output_dim=(1), name='Quantum_Conv_Layer_1')
size_1 = int(1+(X.shape[1]-c_filter)/c_strides)
q_conv_layer_1_list = []
# height iteration
for i in range(size_1):
# width iteration
for j in range(size_1):
temp = q_conv_layer_1(X[:, 2*i:2*(i+1)+1, 2*j:2*(j+1)+1])
temp = tf.keras.layers.Reshape((1,))(temp)
q_conv_layer_1_list += [temp]
concat_layer_1 = tf.keras.layers.Concatenate(axis=1)(q_conv_layer_1_list)
reshape_layer_1 = tf.keras.layers.Reshape((size_1, size_1))(concat_layer_1)
# Second Quantum Conv Layer, trainable params = 18*L, output size = 6 x 6
num_conv_layer_2 = 1
q_conv_layer_2 = qml.qnn.KerasLayer(q_conv, {"conv_params": (2, num_conv_layer_2, 9)}, output_dim=(1), name='Quantum_Conv_Layer_2')
size_2 = int(1+(reshape_layer_1.shape[1]-c_filter)/c_strides)
q_conv_layer_2_list = []
# height iteration
for i in range(size_2):
# width iteration
for j in range(size_2):
temp = q_conv_layer_2(reshape_layer_1[:, 2*i:2*(i+1)+1, 2*j:2*(j+1)+1])
temp = tf.keras.layers.Reshape((1,))(temp)
q_conv_layer_2_list += [temp]
concat_layer_2 = tf.keras.layers.Concatenate(axis=1)(q_conv_layer_2_list)
reshape_layer_2 = tf.keras.layers.Reshape((size_2, size_2, 1))(concat_layer_2)
# Max Pooling Layer, output size = 9
max_pool_layer = tf.keras.layers.MaxPooling2D(pool_size=(2, 2), strides=None, name='Max_Pool_Layer')(reshape_layer_2)
reshape_layer_3 = tf.keras.layers.Reshape((9,))(max_pool_layer)
maxpool_model = tf.keras.Model(inputs=X, outputs=reshape_layer_3)
# -
maxpool_model(X_train[0:1])
maxpool_model.get_layer('Quantum_Conv_Layer_1').set_weights([qconv_1_weights])
maxpool_model.get_layer('Quantum_Conv_Layer_2').set_weights([qconv_2_weights])
# +
maxpool_train = maxpool_model(X_train)
maxpool_test = maxpool_model(X_test)
maxpool_train.shape, maxpool_test.shape
# +
n_qubits = 1 # number of class
dev_state = qml.device("default.qubit", wires=n_qubits)
@qml.qnode(dev_state)
def q_fc_state(params, inputs):
# layer iteration
for l in range(len(params[0])):
# qubit iteration
for q in range(n_qubits):
# gate iteration
for g in range(int(len(inputs)/3)):
qml.Rot(*(params[0][l][3*g:3*(g+1)] * inputs[3*g:3*(g+1)] + params[1][l][3*g:3*(g+1)]), wires=q)
#return [qml.expval(qml.Hermitian(density_matrix(state_labels[i]), wires=[i])) for i in range(n_qubits)]
return qml.expval(qml.Hermitian(density_matrix(state_labels[0]), wires=[0]))
# -
q_fc_state(np.zeros((2,1,9)), maxpool_train[0])
q_fc_state(q_fc_weights, maxpool_train[0])
# +
train_state = np.zeros((len(X_train), 2), dtype=np.complex_)
test_state = np.zeros((len(X_test), 2), dtype=np.complex_)
for i in range(len(train_state)):
q_fc_state(q_fc_weights, maxpool_train[i])
temp = np.flip(dev_state._state)
train_state[i, :] = temp
for i in range(len(test_state)):
q_fc_state(q_fc_weights, maxpool_test[i])
temp = np.flip(dev_state._state)
test_state[i, :] = temp
# -
train_state.shape, test_state.shape
# +
# sanity check
print(((np.conj(train_state) @ density_matrix(state_labels[0])) * train_state)[:, 0] > 0.5)
print(((np.conj(test_state) @ density_matrix(state_labels[0])) * test_state)[:, 0] > 0.5)
# -
np.savetxt('./2_QConv2ent_QFC-State_Train_all_samples.txt', train_state)
np.savetxt('./2_QConv2ent_QFC-State_Test_all_samples.txt', test_state)
# ## Random Starting State
# +
# Input image, size = 27 x 27
X = tf.keras.Input(shape=(27,27), name='Input_Layer')
# Specs for Conv
c_filter = 3
c_strides = 2
# First Quantum Conv Layer, trainable params = 18*L, output size = 13 x 13
num_conv_layer_1 = 1
q_conv_layer_1 = qml.qnn.KerasLayer(q_conv, {"conv_params": (2, num_conv_layer_1, 9)}, output_dim=(1), name='Quantum_Conv_Layer_1')
size_1 = int(1+(X.shape[1]-c_filter)/c_strides)
q_conv_layer_1_list = []
# height iteration
for i in range(size_1):
# width iteration
for j in range(size_1):
temp = q_conv_layer_1(X[:, 2*i:2*(i+1)+1, 2*j:2*(j+1)+1])
temp = tf.keras.layers.Reshape((1,))(temp)
q_conv_layer_1_list += [temp]
concat_layer_1 = tf.keras.layers.Concatenate(axis=1)(q_conv_layer_1_list)
reshape_layer_1 = tf.keras.layers.Reshape((size_1, size_1))(concat_layer_1)
# Second Quantum Conv Layer, trainable params = 18*L, output size = 6 x 6
num_conv_layer_2 = 1
q_conv_layer_2 = qml.qnn.KerasLayer(q_conv, {"conv_params": (2, num_conv_layer_2, 9)}, output_dim=(1), name='Quantum_Conv_Layer_2')
size_2 = int(1+(reshape_layer_1.shape[1]-c_filter)/c_strides)
q_conv_layer_2_list = []
# height iteration
for i in range(size_2):
# width iteration
for j in range(size_2):
temp = q_conv_layer_2(reshape_layer_1[:, 2*i:2*(i+1)+1, 2*j:2*(j+1)+1])
temp = tf.keras.layers.Reshape((1,))(temp)
q_conv_layer_2_list += [temp]
concat_layer_2 = tf.keras.layers.Concatenate(axis=1)(q_conv_layer_2_list)
reshape_layer_2 = tf.keras.layers.Reshape((size_2, size_2, 1))(concat_layer_2)
# Max Pooling Layer, output size = 9
max_pool_layer = tf.keras.layers.MaxPooling2D(pool_size=(2, 2), strides=None, name='Max_Pool_Layer')(reshape_layer_2)
reshape_layer_3 = tf.keras.layers.Reshape((9,))(max_pool_layer)
# Quantum FC Layer, trainable params = 18*L*n_class + 2, output size = 2
num_fc_layer = 1
q_fc_layer_0 = qml.qnn.KerasLayer(q_fc, {"params": (2, num_fc_layer, 9)}, output_dim=2)(reshape_layer_3)
# Alpha Layer
alpha_layer_0 = class_weights()(q_fc_layer_0)
model_random = tf.keras.Model(inputs=X, outputs=alpha_layer_0)
model_maxpool_random = tf.keras.Model(inputs=X, outputs=reshape_layer_3)
# -
model_random(X_train[0:1])
model_random.weights
random_weights = np.array([[[-0.52412415, 0.31337726, -0.0668911 , -0.43626598,
-0.10543117, -0.11661649, -0.22867104, 0.26161313,
0.03067034]],
[[-0.3362496 , 0.52573454, -0.12595582, 0.4650234 ,
0.10775095, 0.2522235 , 0.06523472, 0.32300556,
0.42630672]]])
# +
maxpool_train = model_maxpool_random(X_train)
maxpool_test = model_maxpool_random(X_test)
maxpool_train.shape, maxpool_test.shape
# +
n_qubits = 1 # number of class
dev_state = qml.device("default.qubit", wires=n_qubits)
@qml.qnode(dev_state)
def q_fc_state(params, inputs):
# layer iteration
for l in range(len(params[0])):
# qubit iteration
for q in range(n_qubits):
# gate iteration
for g in range(int(len(inputs)/3)):
qml.Rot(*(params[0][l][3*g:3*(g+1)] * inputs[3*g:3*(g+1)] + params[1][l][3*g:3*(g+1)]), wires=q)
#return [qml.expval(qml.Hermitian(density_matrix(state_labels[i]), wires=[i])) for i in range(n_qubits)]
return qml.expval(qml.Hermitian(density_matrix(state_labels[0]), wires=[0]))
# -
q_fc_state(np.zeros((2,1,9)), maxpool_train[0])
q_fc_state(random_weights, maxpool_train[21])
# +
train_state = np.zeros((len(X_train), 2), dtype=np.complex_)
test_state = np.zeros((len(X_test), 2), dtype=np.complex_)
for i in range(len(train_state)):
q_fc_state(random_weights, maxpool_train[i])
temp = np.flip(dev_state._state)
train_state[i, :] = temp
for i in range(len(test_state)):
q_fc_state(random_weights, maxpool_test[i])
temp = np.flip(dev_state._state)
test_state[i, :] = temp
# -
train_state.shape, test_state.shape
# +
# sanity check
print(((np.conj(train_state) @ density_matrix(state_labels[0])) * train_state)[:, 0] > 0.5)
print(((np.conj(test_state) @ density_matrix(state_labels[0])) * test_state)[:, 0] > 0.5)
# -
np.savetxt('./2_QConv2ent_QFC-RandomState_Train_all_samples.txt', train_state)
np.savetxt('./2_QConv2ent_QFC-RandomState_Test_all_samples.txt', test_state)
# ## Constructing quantum state with 10 set of weights
random_weights = np.array(model.get_weights()[0], dtype=float)
# +
QFC_weights_list = [random_weights]
for i in range(10):
if i == 9:
model.load_weights('./Model_2/2_PCA_QFC_saved-model-' + str(i+1) + '.hdf5')
else:
model.load_weights('./Model_2/2_PCA_QFC_saved-model-0' + str(i+1) + '.hdf5')
QFC_weights_list += [np.array(model.get_weights()[0], dtype=float)]
len(QFC_weights_list)
# +
n_qubits = 1
dev_state = qml.device("default.qubit", wires=n_qubits)
@qml.qnode(dev_state)
def q_fc_state(params, inputs):
"""A variational quantum circuit representing the DRC.
Args:
params (array[float]): array of parameters
inputs = [x, y]
x (array[float]): 1-d input vector
y (array[float]): single output state density matrix
Returns:
float: fidelity between output state and input
"""
# layer iteration
for l in range(len(params[0])):
# qubit iteration
for q in range(n_qubits):
# gate iteration
for g in range(int(len(inputs)/3)):
qml.Rot(*(params[0][l][3*g:3*(g+1)] * inputs[3*g:3*(g+1)] + params[1][l][3*g:3*(g+1)]), wires=q)
return qml.expval(qml.Hermitian(dm_labels[0], wires=[0]))
# -
q_fc_state(QFC_weights_list[0], X_test[0])
# +
for k in range(10+1):
train_state = np.zeros((len(X_train), 2), dtype=np.complex_)
test_state = np.zeros((len(X_test), 2), dtype=np.complex_)
for i in range(len(train_state)):
q_fc_state(QFC_weights_list[k], X_train[i])
temp = np.flip(dev_state._state)
train_state[i, :] = temp
for i in range(len(test_state)):
q_fc_state(QFC_weights_list[k], X_test[i])
temp = np.flip(dev_state._state)
test_state[i, :] = temp
print(train_state.shape, test_state.shape)
np.savetxt('./Model_2/2_PCA_QFC-State_Train_Epoch=' + str(k) + '.txt', train_state)
np.savetxt('./Model_2/2_PCA_QFC-State_Test_Epoch=' + str(k) + '.txt', test_state)
# -
# # Finish
first_10_epoch = H.history
first_10_epoch
# initial 10 epoch
model.get_weights()
# # Making animations
random_weights = model.get_weights()
random_weights
# +
weights_list = [random_weights]
for i in range(10):
if i == 9:
model.load_weights('./Model_2/2_QConv2ent_QFC_saved-model-' + str(i+1) + '.hdf5')
weights_list += [model.get_weights()]
else:
model.load_weights('./Model_2/2_QConv2ent_QFC_saved-model-0' + str(i+1) + '.hdf5')
weights_list += [model.get_weights()]
len(weights_list)
# +
# Input image, size = 27 x 27
X = tf.keras.Input(shape=(27,27), name='Input_Layer')
# Specs for Conv
c_filter = 3
c_strides = 2
# First Quantum Conv Layer, trainable params = 18*L, output size = 13 x 13
num_conv_layer_1 = 1
q_conv_layer_1 = qml.qnn.KerasLayer(q_conv, {"conv_params": (2, num_conv_layer_1, 9)}, output_dim=(1), name='Quantum_Conv_Layer_1')
size_1 = int(1+(X.shape[1]-c_filter)/c_strides)
q_conv_layer_1_list = []
# height iteration
for i in range(size_1):
# width iteration
for j in range(size_1):
temp = q_conv_layer_1(X[:, 2*i:2*(i+1)+1, 2*j:2*(j+1)+1])
temp = tf.keras.layers.Reshape((1,))(temp)
q_conv_layer_1_list += [temp]
concat_layer_1 = tf.keras.layers.Concatenate(axis=1)(q_conv_layer_1_list)
reshape_layer_1 = tf.keras.layers.Reshape((size_1, size_1))(concat_layer_1)
qconv1_model = tf.keras.Model(inputs=X, outputs=reshape_layer_1)
# +
X_2 = tf.keras.Input(shape=(size_1,size_1), name='Input_Layer_2')
# Second Quantum Conv Layer, trainable params = 18*L, output size = 6 x 6
num_conv_layer_2 = 1
q_conv_layer_2 = qml.qnn.KerasLayer(q_conv, {"conv_params": (2, num_conv_layer_2, 9)}, output_dim=(1), name='Quantum_Conv_Layer_2')
size_2 = int(1+(reshape_layer_1.shape[1]-c_filter)/c_strides)
q_conv_layer_2_list = []
# height iteration
for i in range(size_2):
# width iteration
for j in range(size_2):
temp = q_conv_layer_2(X_2[:, 2*i:2*(i+1)+1, 2*j:2*(j+1)+1])
temp = tf.keras.layers.Reshape((1,))(temp)
q_conv_layer_2_list += [temp]
concat_layer_2 = tf.keras.layers.Concatenate(axis=1)(q_conv_layer_2_list)
reshape_layer_2 = tf.keras.layers.Reshape((size_2, size_2, 1))(concat_layer_2)
qconv2_model = tf.keras.Model(inputs=X_2, outputs=reshape_layer_2)
# +
X_3 = tf.keras.Input(shape=(size_2,size_2,1), name='Input_Layer_3')
# Max Pooling Layer, output size = 9
max_pool_layer = tf.keras.layers.MaxPooling2D(pool_size=(2, 2), strides=None, name='Max_Pool_Layer')(X_3)
reshape_layer_3 = tf.keras.layers.Reshape((9,))(max_pool_layer)
maxpool_model = tf.keras.Model(inputs=X_3, outputs=reshape_layer_3)
# +
n_qubits = 1 # number of class
dev_state = qml.device("default.qubit", wires=n_qubits)
@qml.qnode(dev_state)
def q_fc_state(params, inputs):
# layer iteration
for l in range(len(params[0])):
# qubit iteration
for q in range(n_qubits):
# gate iteration
for g in range(int(len(inputs)/3)):
qml.Rot(*(params[0][l][3*g:3*(g+1)] * inputs[3*g:3*(g+1)] + params[1][l][3*g:3*(g+1)]), wires=q)
#return [qml.expval(qml.Hermitian(density_matrix(state_labels[i]), wires=[i])) for i in range(n_qubits)]
return qml.expval(qml.Hermitian(density_matrix(state_labels[0]), wires=[0]))
# -
a = qconv1_model(X_train[0:1])
b = qconv2_model(a)
c = maxpool_model(b)
d = q_fc_state(weights_list[0][2], c)
print(d)
print('ok')
X_train.shape, X_test.shape
for k in range(11):
qconv1_model.get_layer('Quantum_Conv_Layer_1').set_weights([weights_list[k][0]])
qconv2_model.get_layer('Quantum_Conv_Layer_2').set_weights([weights_list[k][1]])
print('Model ' + str(k) + ' got loaded.')
preprocessed_img_train_1 = qconv1_model(X_train)
preprocessed_img_test_1 = qconv1_model(X_test)
data_train = preprocessed_img_train_1.numpy().reshape(-1, 13*13)
np.savetxt('./Model_2/2_QConv2ent_QFC/2_QConv2ent_QFC-Filter1_Image_Train_Epoch=' + str(k) + '.txt', data_train)
data_test = preprocessed_img_test_1.numpy().reshape(-1, 13*13)
np.savetxt('./Model_2/2_QConv2ent_QFC/2_QConv2ent_QFC-Filter1_Image_Test_Epoch=' + str(k) + '.txt', data_test)
print('Filter 1 finished.')
preprocessed_img_train_2 = qconv2_model(preprocessed_img_train_1)
preprocessed_img_test_2 = qconv2_model(preprocessed_img_test_1)
data_train = preprocessed_img_train_2.numpy().reshape(-1, 6*6)
np.savetxt('./Model_2/2_QConv2ent_QFC/2_QConv2ent_QFC-Filter2_Image_Train_Epoch=' + str(k) + '.txt', data_train)
data_test = preprocessed_img_test_2.numpy().reshape(-1, 6*6)
np.savetxt('./Model_2/2_QConv2ent_QFC/2_QConv2ent_QFC-Filter2_Image_Test_Epoch=' + str(k) + '.txt', data_test)
print('Filter 2 finished.')
maxpool_train = maxpool_model(preprocessed_img_train_2)
maxpool_test = maxpool_model(preprocessed_img_test_2)
train_state = np.zeros((len(X_train), 2), dtype=np.complex_)
test_state = np.zeros((len(X_test), 2), dtype=np.complex_)
for i in range(len(train_state)):
q_fc_state(weights_list[k][2], maxpool_train[i])
temp = np.flip(dev_state._state)
train_state[i, :] = temp
for i in range(len(test_state)):
q_fc_state(weights_list[k][2], maxpool_test[i])
temp = np.flip(dev_state._state)
test_state[i, :] = temp
np.savetxt('./Model_2/2_QConv2ent_QFC/2_QConv2ent_QFC-State_Train_Epoch=' + str(k) + '.txt', train_state)
np.savetxt('./Model_2/2_QConv2ent_QFC/2_QConv2ent_QFC-State_Test_Epoch=' + str(k) + '.txt', test_state)
print('Quantum State finished.')
H.history
| PennyLane/Data Reuploading Classifier/Buat Paper/2_QConv2ent_QFC (best).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
# +
# %config IPython.matplotlib.backend = "retina"
from matplotlib import rcParams
rcParams["figure.dpi"] = 150
rcParams["savefig.dpi"] = 150
# import maelstrom as ms
import numpy as np
import corner
import pandas as pd
import matplotlib.pyplot as plt
# -
import exoplanet as xo
xo.__version__
# +
kicid = 9651065
metadata = np.loadtxt(f"data/kic{kicid}_metadata.csv", delimiter=",", skiprows=1)
nu_arr = metadata[::6]
times, mags = np.loadtxt(f"data/kic{kicid}_lc.txt",usecols=(0,1)).T
# Let's get rid of the crappy early data
start = 3000
times = times[start:]
mags = mags[start:]
# Subtract midpoint
time_mid = (times[0] + times[-1]) / 2.
times -= time_mid
nu_arr = np.loadtxt(f"data/kic{kicid}_metadata.csv", delimiter=",", skiprows=1)[::6]
nu_arr = nu_arr[:3]
mags *= 1000.
plt.plot(times, mags, ".k")
# +
import theano.tensor as tt
import pymc3 as pm
from exoplanet.orbits import get_true_anomaly
nu_arr = np.array(nu_arr)
t = times
y = mags
with pm.Model() as model:
# Parameters: Use normal priors when possible - Uniforms can be hard to sample
logperiod = pm.Normal("logperiod", mu=np.log(272.), sd=100)
period = pm.Deterministic("period", tt.exp(logperiod))
t0 = pm.Normal("t0", mu=0.0, sd=100.0)
varpi = xo.distributions.Angle("varpi")
# varpi = pm.Normal("varpi", mu=0, sd=2*np.pi)
# varpi_vec = pm.Normal("varpi_vec", shape=2, testval=np.array([1.0, 0.0]))
# varpi = pm.Deterministic("varpi", tt.arctan2(varpi_vec[0], varpi_vec[1]))
eccen = pm.Uniform("eccen", lower=1e-5, upper=1.0 - 1e-5, testval=0.1)
logs = pm.Normal('logs', mu=np.log(np.std(y)), sd=100)
lighttime = pm.Normal('lighttime', mu=0.0, sd=100.0, shape=len(nu_arr))
# Better parameterization for the reference time
# see: https://github.com/dfm/exoplanet/blob/master/exoplanet/orbits/keplerian.py
sinw = tt.sin(varpi)
cosw = tt.cos(varpi)
opsw = 1 + sinw
E0 = 2 * tt.arctan2(tt.sqrt(1-eccen)*cosw, tt.sqrt(1+eccen)*opsw)
M0 = E0 - eccen * tt.sin(E0)
tref = pm.Deterministic("tref", t0 - M0 * period / (2*np.pi))
# Mean anom
M = 2.0 * np.pi * (t - tref) / period
# True anom
f = get_true_anomaly(M, eccen + tt.zeros_like(M))
psi = - (1 - tt.square(eccen)) * tt.sin(f+varpi) / (1 + eccen*tt.cos(f))
# tau in d
tau = (lighttime / 86400.)[None,:] * psi[:,None]
# Just sample in the weights parameters too. This seems to be faster
factor = 2. * np.pi * nu_arr[None, :]
arg = factor * t[:, None] - factor * tau
mean_flux = pm.Normal("mean_flux", mu=0.0, sd=100.0)
W_hat_cos = pm.Normal("W_hat_cos", mu=0.0, sd=100.0, shape=len(nu_arr))
W_hat_sin = pm.Normal("W_hat_sin", mu=0.0, sd=100.0, shape=len(nu_arr))
model_tensor = tt.dot(tt.cos(arg), W_hat_cos[:, None])
model_tensor += tt.dot(tt.sin(arg), W_hat_sin[:, None])
model_tensor = tt.squeeze(model_tensor) + mean_flux
# Condition on the observations
pm.Normal("obs", mu=model_tensor, sd=tt.exp(logs), observed=y)
# -
# Let's be a little more clever about the optimization:
with model:
map_soln = xo.optimize(start=model.test_point, vars=[mean_flux, W_hat_cos, W_hat_sin])
map_soln = xo.optimize(start=map_soln, vars=[logs, mean_flux, W_hat_cos, W_hat_sin])
map_soln = xo.optimize(start=map_soln, vars=[lighttime, t0])
map_soln = xo.optimize(start=map_soln, vars=[logs, mean_flux, W_hat_cos, W_hat_sin])
map_soln = xo.optimize(start=map_soln, vars=[logperiod, t0])
map_soln = xo.optimize(start=map_soln, vars=[eccen, varpi])
map_soln = xo.optimize(start=map_soln, vars=[logperiod, t0])
map_soln = xo.optimize(start=map_soln, vars=[lighttime])
map_soln = xo.optimize(start=map_soln, vars=[eccen, varpi])
map_soln = xo.optimize(start=map_soln, vars=[logs, mean_flux, W_hat_cos, W_hat_sin])
map_soln = xo.optimize(start=map_soln)
map_soln
with model:
tau_val = xo.utils.eval_in_model(tau, map_soln)
plt.plot(times, tau_val*86400-np.mean(tau_val), lw=1)
plt.xlabel('Time (d)')
plt.ylabel('Time delay (s)')
# +
with model:
lc_val = xo.utils.eval_in_model(model_tensor, map_soln)
plt.plot(times, mags, ".k")
plt.plot(times, lc_val, color="C0", lw=1)
plt.tight_layout()
# plt.xlim(0,5)
#plt.ylim(-1, 5)
plt.xlabel("Time (d)")
plt.ylabel("L(t)");
plt.title('yup')
# -
np.random.seed(42)
sampler = xo.PyMC3Sampler(start=50, window=50, finish=300)
with model:
burnin = sampler.tune(tune=3000, start=map_soln,
step_kwargs=dict(target_accept=0.9))
trace = sampler.sample(draws=3000)
pm.summary(trace, varnames=["period", "lighttime", "t0", "varpi", "eccen", "logs", "mean_flux", "W_hat_cos", "W_hat_sin"])
import corner
samples = pm.trace_to_dataframe(trace, varnames=["period", "t0", "varpi", "eccen", "logs"])
for k in samples.columns:
if "_" in k:
samples[k.replace("_", "")] = samples[k]
del samples[k]
corner.corner(samples);
| using_exoplanet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.6 64-bit (''ds-Flask'': conda)'
# name: python37664bitdsflaskconda5d6acf79da9542b3aeeb7103e7ad852b
# ---
from utils import openaq
api=openaq.OpenAQ()
status,body=api.measurements(city='Los Angeles', parameter='pm25')
t=body['results']
out=[]
for i in t:
out.append((i['date']['utc'],i['value']))
# !pwd
from sqlite3 import connect
conn = connect("db.sqlite3")
curs=conn.cursor()
result=curs.execute("SELECT * FROM record r WHERE r.value >= 10").fetchall()
result[1]
| unit-1-build/app/testing.ipynb |
# +++
# title = "Working with Scikit-Learn"
# +++
#
# This guide goes through how to use this package with the Scikit-Learn package.
# ## Load the train and test datasets
#
# We'll first get the train and test splits for the `musk` dataset (completely unrelated to <NAME>).
# +
from tabben.datasets import OpenTabularDataset
train = OpenTabularDataset('./temp', 'musk') # train split by default
test = OpenTabularDataset('./temp', 'musk', split='test') # should only be used ONCE!
print(f'The {train.name} dataset is a {train.task} task with {train.num_classes} classes.')
# -
X_fulltrain, y_fulltrain = train.numpy()
# In order to tune some hyperparameters, we'll need our own validation split (not the test set). We'll do an 80-20 split and stratify on the class.
# +
from sklearn.model_selection import train_test_split
X_train, X_valid, y_train, y_valid = train_test_split(
X_fulltrain, y_fulltrain,
train_size=0.8,
stratify=y_fulltrain
)
# -
# ## Create and train a model
#
# Next, we'll create a $k$-Nearest Neighbors model and train it on our train split.
# +
from sklearn.neighbors import KNeighborsClassifier
model = KNeighborsClassifier()
# -
model.fit(X_train, y_train)
# And we'll evaluate it on our *validation* set, using a simple accuracy metric.
model.score(X_valid, y_valid)
# ## In a larger data processing pipeline
#
# However, it might be the case that we want to use a sklearn pipeline to do some data preprocessing like feature normalization, one-hot encoding, etc. or explore the effect of, say, turning continuous attributes into binary ones.
# +
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import Binarizer, StandardScaler
pipeline = make_pipeline(
StandardScaler(with_std=False),
Binarizer(),
KNeighborsClassifier(),
)
# -
pipeline.fit(X_train, y_train)
pipeline.score(X_valid, y_valid)
# ---
#
# This code was last run using the following package versions (if you're looking at the webpage which doesn't have the output, see the notebook for versions):
# +
from importlib.metadata import version
packages = ['scikit-learn', 'tabben']
for pkg in packages:
print(f'{pkg}: {version(pkg)}')
| docs/notebooks/integrations/scikit-learn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Ensemble-Trees" data-toc-modified-id="Ensemble-Trees-1"><span class="toc-item-num">1 </span>Ensemble Trees</a></span><ul class="toc-item"><li><span><a href="#Bagging" data-toc-modified-id="Bagging-1.1"><span class="toc-item-num">1.1 </span>Bagging</a></span></li><li><span><a href="#Random-Forest" data-toc-modified-id="Random-Forest-1.2"><span class="toc-item-num">1.2 </span>Random Forest</a></span></li><li><span><a href="#Implementation" data-toc-modified-id="Implementation-1.3"><span class="toc-item-num">1.3 </span>Implementation</a></span></li><li><span><a href="#Feature-Importance" data-toc-modified-id="Feature-Importance-1.4"><span class="toc-item-num">1.4 </span>Feature Importance</a></span></li><li><span><a href="#Extra-Trees" data-toc-modified-id="Extra-Trees-1.5"><span class="toc-item-num">1.5 </span>Extra Trees</a></span></li></ul></li><li><span><a href="#Reference" data-toc-modified-id="Reference-2"><span class="toc-item-num">2 </span>Reference</a></span></li></ul></div>
# +
# code for loading the format for the notebook
import os
# path : store the current path to convert back to it later
path = os.getcwd()
os.chdir(os.path.join('..', 'notebook_format'))
from formats import load_style
load_style(css_style = 'custom2.css', plot_style = False)
# +
os.chdir(path)
# 1. magic for inline plot
# 2. magic to print version
# 3. magic so that the notebook will reload external python modules
# 4. magic to enable retina (high resolution) plots
# https://gist.github.com/minrk/3301035
# %matplotlib inline
# %load_ext watermark
# %load_ext autoreload
# %autoreload 2
# %config InlineBackend.figure_format = 'retina'
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.datasets import load_iris
from sklearn.metrics import mean_squared_error
from sklearn.model_selection import train_test_split, GridSearchCV
from sklearn.tree import DecisionTreeRegressor, DecisionTreeClassifier
from sklearn.ensemble import RandomForestRegressor, ExtraTreesRegressor
# %watermark -a 'Ethen' -d -t -v -p numpy,pandas,matplotlib,sklearn
# -
# # Ensemble Trees
# Some of the materials builds on top of the [previous documentation/implementation on decision trees](http://nbviewer.jupyter.org/github/ethen8181/machine-learning/blob/master/trees/decision_tree.ipynb), thus it might be best to walk through that one first.
#
# Ensembling is a very popular method for improving the predictive performance of machine learning models. Let's pretend that instead of building a single model to solve a binary classification problem, you created **five independent models**, and each model was correct about 70% of the time. If you combined these models into an "ensemble" and used their majority vote as a prediction, how often would the ensemble be correct?
# +
# generate 1000 random numbers (between 0 and 1) for each model,
# representing 1000 observations
np.random.seed(1234)
mod1 = np.random.rand(1000)
mod2 = np.random.rand(1000)
mod3 = np.random.rand(1000)
mod4 = np.random.rand(1000)
mod5 = np.random.rand(1000)
# each model independently predicts 1 (the "correct response")
# if random number was at least 0.3
preds1 = np.where(mod1 > 0.3, 1, 0)
preds2 = np.where(mod2 > 0.3, 1, 0)
preds3 = np.where(mod3 > 0.3, 1, 0)
preds4 = np.where(mod4 > 0.3, 1, 0)
preds5 = np.where(mod5 > 0.3, 1, 0)
# how accurate was each individual model?
print(preds1.mean())
print(preds2.mean())
print(preds3.mean())
print(preds4.mean())
print(preds5.mean())
# +
# average the predictions, and then round to 0 or 1
# you can also do a weighted average, as long as the weight adds up to 1
ensemble_preds = np.round((preds1 + preds2 + preds3 + preds4 + preds5) / 5).astype(int)
# how accurate was the ensemble?
print(ensemble_preds.mean())
# -
# ## Bagging
# The primary weakness of decision trees is that they don't tend to have the best predictive accuracy and the result can be very unstable. This is partially due to the fact that we were using greedy algorithm to choose the rule/feature to split the tree. Hence a small variations in the data might result in a completely different tree being generated. Fortunately, this problem can be mitigated by training an ensemble of decision trees and use these trees to form a "forest".
#
# This first idea we'll introduce is **Bagging**. **Bagging**, short for **bootstrap aggregation** is a general procedure for reducing the variance of a machine learning algorithm, although it can used with any type of method, it is most commonly applied to tree-based models. The way it works is:
#
# Given a training set $X = x_1, ..., x_n$ with responses $Y = y_1, ..., y_n$, bagging repeatedly ($B$ times) selects a random sample with replacement (a.k.a bootstrap sample) of the training set and fits trees to these newly generated samples:
#
# For $b = 1, ..., B$:
#
# 1. Sample, with replacement, $n$ training examples from $X$, $Y$; call these $X_b$, $Y_b$. Note that the bootstrap sample should be the same size as the original training set
# 2. Train a tree, $f_b$, on $X_b$, $Y_b$. For these individual tree, we should allow them to grow deeper (increase the max_depth parameter) so that they have low bias/high variance
#
# After training, predictions for unseen samples $x'$ can be made by averaging the predictions from all the individual regression trees on $x'$:
#
#
# $$
# \begin{align}
# f' = \frac {1}{B}\sum _{b=1}^{B}{f}_{b}(x')
# \end{align}
# $$
#
# Or by taking the majority vote in the case of classification trees. If you are wondering why bootstrapping is a good idea, the rationale is:
#
# We wish to ask a question of a population but we can't. Instead, we take a sample and ask the question to it instead. Now, how confident we should be that the sample answer is close to the population answer obviously depends on the structure of population. One way we might learn about this is to take samples from the population again and again, ask them the question, and see how variable the sample answers tended to be. But often times this isn't possible (we wouldn't relaunch the Titanic and crash it into another iceberg), thus we can use the information in the sample we actually have to learn about it.
#
# This is a reasonable thing to do because not only is the sample you have the best and the only information you have about what the population actually looks like, but also because most samples will, if they're randomly chosen, look quite like the population they came from. In the end, sampling with replacement is just a convenient way to treat the sample like it's a population and to sample from it in a way that reflects its shape.
# ## Random Forest
#
# Random Forest is very similar to bagged trees. Exactly like bagging, we create an ensemble of decision trees using bootstrapped samples of the training set. When building each tree, however, each time a split is considered, a random sample of $m$ features is chosen as split candidates from the full set of $p$ features. The split is only allowed to use one of those $m$ features to generate the best rule/feature to split on.
#
# - For **classification**, $m$ is typically chosen to be, $\sqrt{p}$, the square root of $p$.
# - For **regression**, $m$ is typically chosen to be somewhere between $p/3$ and $p$.
#
# The whole point of choosing a new random sample of features for every single tree at every single split is to correct for decision trees' habit of overfitting to their training set. Suppose there is one very strong feature in the data set, when using bagged trees, most of the trees will use that feature as the top split, resulting in an ensemble of similar trees that are highly correlated. By randomly leaving out candidate features from each split, Random Forest "decorrelates" the trees, such that the averaging process can further reduce the variance of the resulting model.
# ## Implementation
# Here, we will use the [Wine Quality Data Set](https://archive.ics.uci.edu/ml/datasets/Wine+Quality) to test our implementation. This [link](https://archive.ics.uci.edu/ml/machine-learning-databases/wine-quality/winequality-white.csv) should download the .csv file. The task is to predict the quality of the wine (a scale of 1 ~ 10) given some of its features. We'll build three types of regression model, decision tree, bagged decision tree and random forest on the training set and compare the result on the test set.
# +
wine = pd.read_csv('winequality-white.csv', sep = ';')
# train/test split the features and response column
y = wine['quality'].values
X = wine.drop('quality', axis = 1).values
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size = 0.2, random_state = 1234)
print('dimension of the dataset: ', wine.shape)
wine.head()
# -
# this cell simply demonstrates how to create boostrap samples
# we create an array of numbers from 1 to 20
# create the boostrap sample on top of that
np.random.seed(1)
nums = np.arange(1, 21)
print('original:', nums)
print('bootstrap: ', np.random.choice(nums, size = 20, replace = True))
class RandomForest:
"""
Regression random forest using scikit learn's
decision tree as the base tree
Parameters
----------
n_estimators: int
the number of trees that you're going built
on the bagged sample (you can even shutoff
the bagging procedure for some packages)
max_features: int
the number of features that you allow
when deciding which feature to split on
all the other parameters for a decision tree like
max_depth or min_sample_split also applies to Random Forest,
it is just not used here as that is more
related to a single decision tree
"""
def __init__(self, n_estimators, max_features):
self.n_estimators = n_estimators
self.max_features = max_features
def fit(self, X, y):
# for each base-tree models:
# 1. draw bootstrap samples from the original data
# 2. train the tree model on that bootstrap sample, and
# during training, randomly select a number of features to
# split on each node
self.estimators = []
for i in range(self.n_estimators):
boot = np.random.choice(y.shape[0], size = y.shape[0], replace = True)
X_boot, y_boot = X[boot], y[boot]
tree = DecisionTreeRegressor(max_features = self.max_features)
tree.fit(X_boot, y_boot)
self.estimators.append(tree)
return self
def predict(self, X):
# for the prediction, we average the
# predictions made by each of the bagged tree
pred = np.empty((X.shape[0], self.n_estimators))
for i, tree in enumerate(self.estimators):
pred[:, i] = tree.predict(X)
pred = np.mean(pred, axis = 1)
return pred
# +
# compare the results between a single decision tree,
# bagging and random forest, the lower the mean square
# error, the better
tree = DecisionTreeRegressor()
tree.fit(X_train, y_train)
tree_y_pred = tree.predict(X_test)
print('tree: ', mean_squared_error(y_test, tree_y_pred))
# bagged decision tree
# max_feature = None simply uses all features
bag = RandomForest(n_estimators = 50, max_features = None)
bag.fit(X_train, y_train)
bag_y_pred = bag.predict(X_test)
print('bagged tree: ', mean_squared_error(y_test, bag_y_pred))
# random forest using a random one third of the features at every split
rf = RandomForest(n_estimators = 50, max_features = 1 / 3)
rf.fit(X_train, y_train)
rf_y_pred = rf.predict(X_test)
print('random forest: ', mean_squared_error(y_test, rf_y_pred))
# use library to confirm results are comparable
rf_reg = RandomForestRegressor(n_estimators = 50, max_features = 1 / 3)
rf_reg.fit(X_train, y_train)
rf_reg_y_pred = rf_reg.predict(X_test)
print('random forest library: ', mean_squared_error(y_test, rf_reg_y_pred))
# -
# ## Feature Importance
# When using Bagging with decision tree or using Random Forest, we can increase the predictive accuracy of individual tree. These methods, however, do decrease model interpretability, because it is no longer possible to visualize all the trees that are built to form the "forest". Fortunately, we can still obtain an overall summary of feature importance from these models. The way feature importance works is as follows (there are many ways to do it, this is the implementation that scikit-learn uses):
#
# We first compute the feature importance values of a single tree:
#
# - We can initialize an array `feature_importances` of all zeros with size `n_features`
# - We start building the tree and for each internal node that splits on feature $i$ we compute the information gain (error reduction) of that node multiplied by the proportion of samples that were routed to the node and add this quantity to `feature_importances[i]`
#
# The information gain (error reduction) depends on the impurity criterion that you use (e.g. Gini, Entropy for classification, MSE for regression). Its the impurity of the set of examples that gets routed to the internal node minus the sum of the impurities of the two partitions created by the split.
#
# Now, recall that these Ensemble Tree models simply consists of a bunch of individual trees, hence after computing the `feature_importance` values across all individual trees, we sum them up and take the average across all of them (normalize the values to sum up to 1 if necessary).
#
# Building on top of the [previous documentation/implementation on decision trees](http://nbviewer.jupyter.org/github/ethen8181/machine-learning/blob/master/trees/decision_tree.ipynb), we add the code to compute the feature importance. The code is not shown here, but can be obtained [here](https://github.com/ethen8181/machine-learning/blob/master/trees/tree.py) for those that are interested in the implementation.
# +
from tree import Tree
# load a sample dataset
iris = load_iris()
iris_X = iris.data
iris_y = iris.target
# train model and print the feature importance
tree = Tree()
tree.fit(iris_X, iris_y)
print(tree.feature_importance)
# use library to confirm result
# note that the result might not always be the same
# because of decision tree's high variability
clf = DecisionTreeClassifier(criterion = 'entropy', min_samples_split = 10, max_depth = 3)
clf.fit(iris_X, iris_y)
print(clf.feature_importances_)
# -
# For ensemble tree, we simply sum all the feauture importance up and take the average (normalize it to sum up to 1 if necessary). Thus, we will not go through the process of building that from scratch, we'll simply visualize the feature importance of the regression Random Forest that we've previously trained on the wine dataset.
def vis_importance(estimator, feature_names, threshold = 0.05):
"""
Visualize the relative importance of predictors.
Parameters
----------
estimator : sklearn-like ensemble tree model
A tree estimator that contains the attribute
``feature_importances_``.
feature_names : str 1d array or list[str]
Feature names that corresponds to the
feature importance.
threshold : float, default 0.05
Features that have importance scores lower than this
threshold will not be presented in the plot, this assumes
the feature importance sum up to 1.
"""
if not hasattr(estimator, 'feature_importances_'):
msg = '{} does not have the feature_importances_ attribute'
raise ValueError(msg.format(estimator.__class__.__name__))
imp = estimator.feature_importances_
feature_names = np.asarray(feature_names)
mask = imp > threshold
importances = imp[mask]
idx = np.argsort(importances)
scores = importances[idx]
names = feature_names[mask]
names = names[idx]
y_pos = np.arange(1, len(scores) + 1)
if hasattr(estimator, 'estimators_'):
# apart from the mean feature importance, for scikit-learn we can access
# each individual tree's feature importance and compute the standard deviation
tree_importances = np.asarray([tree.feature_importances_
for tree in estimator.estimators_])
importances_std = np.std(tree_importances[:, mask], axis = 0)
scores_std = importances_std[idx]
plt.barh(y_pos, scores, align = 'center', xerr = scores_std)
else:
plt.barh(y_pos, scores, align = 'center')
plt.yticks(y_pos, names)
plt.xlabel('Importance')
plt.title('Feature Importance Plot')
# +
# change default figure and font size
plt.rcParams['figure.figsize'] = 8, 6
plt.rcParams['font.size'] = 12
# visualize the feature importance of every variable
vis_importance(rf_reg, wine.columns[:-1])
# -
# **Caveat:**
#
# One thing to keep in mind when using the impurity based feature importance ranking is that when the dataset has two (or more) correlated features, then from the model's point of view, any of these correlated features can be used as the predictor, with no preference of one over the others. But once one of them is used, the importance of others is significantly reduced since the impurity they can effectively remove has already been removed by the first feature. As a consequence, they will have a lower reported importance. This is not an issue when we want to use feature selection to reduce overfitting, since it makes sense to remove features that are mostly duplicated by other features. But when we're interpreting the data, it can lead to incorrect conclusions that one of the variables is a strong predictor while the others in the same group are unimportant, while actually they are very close in terms of their relationship with the response variable.
#
# The effect of this phenomenon for Random Forest is somewhat reduced thanks to random selection of features at each node creation, but in general the effect is not removed completely. In the following example, we have three correlated variables $X_0$, $X_1$, $X_2$, and no noise in the data, with the output variable being the sum of the three features:
# +
size = 10000
np.random.seed(10)
X_seed = np.random.normal(0, 1, size)
X0 = X_seed + np.random.normal(0, 0.1, size)
X1 = X_seed + np.random.normal(0, 0.1, size)
X2 = X_seed + np.random.normal(0, 0.1, size)
X_012 = np.array([ X0, X1, X2 ]).T
Y = X0 + X1 + X2
rf = RandomForestRegressor(n_estimators = 20, max_features = 2)
rf.fit(X_012, Y)
print('Scores for X0, X1, X2:', np.round(rf.feature_importances_, 3))
# -
# When we compute the feature importances, we see that some of the features have higher importance than the others, while their “true” importance should be very similar. One thing to point out though is that the difficulty of interpreting the importance/ranking of correlated variables is not Random Forest specific, but applies to most model based feature selection methods. This is why it often best practice to remove correlated features prior to training the model.
#
# **Advantages of Random Forests:**
#
# - Require very little feature engineering (e.g. standardization)
# - Easy to use, as it rarely requires parameter tuning to achieve compelling and robust performance
# - Provides a more reliable estimate of feature importance compare to other black-box methods (e.g. deep learning, support vector machine)
# - Performance and computation wise, it is very competitive. Although you can typically find a model that beats Random Forest for any given dataset (typically a deep learning or gradient boosting algorithm), it’s never by much, and it usually takes much longer to train and tune those model
# ## Extra Trees
# What distinguishes Extra Trees from Random Forest is:
#
# - We use the entire training set instead of a bootstrap sample of the training set (but can also be trained on a bootstrapped sample as well if we wish)
# - Just like Random Forest, when choosing rules/features at a split, a random subset of candidate features is used, but now, instead of looking at all the thresholds to find the best the best split, thresholds (for the split) are chosen completely at random for each candidate feature and the best of these randomly generated thresholds is picked as the splitting rule. We all know that tree-based methods employ a greedy algorithm when choosing the feature to split on. Thus, we can think of this as taking an extra step in trying to migitate this drawback
#
# > Based on [Stackoverflow: RandomForestClassifier vs ExtraTreesClassifier in scikit learn](http://stackoverflow.com/questions/22409855/randomforestclassifier-vs-extratreesclassifier-in-scikit-learn?rq=1)
# > In practice, RFs are often more compact than ETs. ETs are generally cheaper to train from a computational point of view but can grow much bigger. ETs can sometime generalize better than RFs but it's hard to guess when it's the case without trying both first (and tuning n_estimators, max_features and min_samples_split by cross-validated grid search).
# +
# grid search on a range of max features and compare
# the performance between Extra Trees and Random Forest
param_name = 'max_features'
max_features_options = np.arange(4, 10)
fit_param = {param_name: max_features_options}
rf_reg = RandomForestRegressor(n_estimators = 30)
et_reg = ExtraTreesRegressor(n_estimators = 30)
gs_rf = GridSearchCV(rf_reg, fit_param, n_jobs = -1)
gs_et = GridSearchCV(et_reg, fit_param, n_jobs = -1)
gs_rf.fit(X_train, y_train)
gs_et.fit(X_train, y_train)
# visualize the performance on the cross validation test score
gs_rf_mean_score = gs_rf.cv_results_['mean_test_score']
gs_et_mean_score = gs_et.cv_results_['mean_test_score']
mean_scores = [gs_rf_mean_score, gs_et_mean_score]
labels = ['RF', 'ET']
for score, label in zip(mean_scores, labels):
plt.plot(max_features_options, score, label = label)
plt.legend()
plt.ylabel('MSE')
plt.xlabel(param_name)
plt.xlim( np.min(max_features_options), np.max(max_features_options) )
plt.show()
# -
# It is not always the case that Random Forest will outperform Extra Trees making it a method that's worth trying. As for interpretation, Extra Trees is simply another kind of Ensemble Tree method, hence we can still access the `feature_importance_` attribute to see which predictors were contributing a lot to explaining the response.
# # Reference
# - [Notebook: Ensembling](http://nbviewer.jupyter.org/github/justmarkham/DAT8/blob/master/notebooks/18_ensembling.ipynb)
# - [Notebook: useR machine learning tutorial Random Forest](http://nbviewer.jupyter.org/github/ledell/useR-machine-learning-tutorial/blob/master/random-forest.ipynb)
# - [Blog: Selecting good features – Part III: random forests](http://blog.datadive.net/selecting-good-features-part-iii-random-forests/)
# - [Blog: The Unreasonable Effectiveness of Random Forests](https://medium.com/rants-on-machine-learning/the-unreasonable-effectiveness-of-random-forests-f33c3ce28883#.pv7i5ien9)
# - [StackExchange: Explaining to laypeople why bootstrapping works](http://stats.stackexchange.com/questions/26088/explaining-to-laypeople-why-bootstrapping-works/)
# - [Stackoverflow: RandomForestClassifier vs ExtraTreesClassifier in scikit learn](http://stackoverflow.com/questions/22409855/randomforestclassifier-vs-extratreesclassifier-in-scikit-learn?rq=1)
# - [Stackoverflow: How are feature_importances in RandomForestClassifier determined?](http://stackoverflow.com/questions/15810339/how-are-feature-importances-in-randomforestclassifier-determined)
| trees/random_forest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Scikit-learn
#
# La página oficial de Scitit-learn es [aquí](https://scikit-learn.org/stable/index.html).
#
#
# ## ¿Cómo instalar biblioteca en Python?
#
# En la consola _Qt Console_, se escribe el comando:
#
# pip install _nombre_de_libreria_
#
#
# ### Ejemplo: Regresión Lineal Simple
#
# Ahora veamos un ejemplo, siguiendo los pasos para hacer un ajuste.
# Se cargan las librerías que se van a utilizar
import numpy as np
import math
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
import sklearn
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
from sklearn.metrics import r2_score
from sklearn.metrics import mean_squared_error
# +
## 1) EXTRAER DATOS
# Los datos pueden encontrarse en diferentes formatos, en nuestro caso están en formato csv.
# Se carga la base de datos y se definen las varibles X y Y
datos = pd.read_csv('Salary_Data.csv') #Se encuentra en la misma carpeta que el jupyter notebook
X = datos.iloc[:, :-1].values #Años de experiencia
Y = datos.iloc[:, 1].values #Salario
print("type(datos): ",type(datos))
print("type(X)",type(X))
print("type(Y)",type(Y))
# +
## 2) ANÁLISIS EXPLORATORIO
# Se realiza una descripción analítica de los datos.
# Se muestran los primeros 5 datos del data frame
datos.head()
#Los datos corresponden al salario de 30 personas y el número de años de experiencia que tienen.
##Información del data frame
#datos.info
#print(datos)
#print(X)
#print(Y)
# -
# Se cuenta el número de NaN's por columna
datos.isnull().sum()
# Cuenta los valores repetidos de la columna 1 (años de experiencia)
datos["YearsExperience"].value_counts().head()
# Cuenta los valores repetidos de la columna 2 (salario)
datos["Salary"].value_counts().head()
# Dimensiones del data frame
datos.shape #Se tienen 30 renglones (personas) y 2 columnas (Años de experiencia y Salario)
# Se cambian los NA's por el promedio de la columna
#Nota: En este caso no hay NA's
datos["YearsExperience"] = datos["YearsExperience"].fillna(datos["YearsExperience"].mean())
datos["YearsExperience"].head()
# +
## 3) VISUALIZACIÓN DE LOS DATOS
# Para entender mejor los datos es necesario graficarlos.
sns.distplot(Y)#Salario
# -
plt.hist(X, bins=[0,2,4,6,8,10,12]) #Se hace la división por cada 2 años
plt.title('Histograma de los años de experiencia')
plt.xlabel('Años')
plt.ylabel('Número de personas')
plt.show()
# +
print(min(Y)) #37731.0
print(max(Y)) #122391.0
plt.hist(Y, bins=[30000,40000,50000,60000,70000,
80000,90000,100000,110000,120000,
130000]) #Cada 10 mil a partir de 30 mil.
plt.title('Histograma del salario')
plt.xlabel('Años')
plt.ylabel('Número de personas')
plt.show()
# -
sns.boxplot(x="YearsExperience", data=datos)
round(np.mean(X),2) #Promedio de años de experiencia
sns.boxplot(x="Salary", data=datos)
round(np.mean(Y),2) #Promedio de salario
#Se muestra la correlación entre las variables
sns.pairplot(datos)
# +
## 4) DIVIDIR LOS DATOS
# Se separan los datos en 2 grupos (usualmente 80% y 20%):
# i) Para entrenar al modelo (80%)
# ii) Para probar el modelo (20%)
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, #Se indican los vectores que se van a dividir
test_size = 0.2, #Se indica el porcentajede los datos para probar el modelo
random_state = 0) #Se fija la semilla
# Nota: Tomar la muestra aleatoria es muy importante porque en caso de que los datos estén
#ordenados el algoritmo no aprende adecuadamente. Por ejemplo si tenemos 80 sanos y 20 enfermos,
#sólo se tomarían los 80 sanos (por ser los primeros 80).
print("El vector X_train tiene ",len(X_train), " elementos.")
print("El vector Y_train tiene ",len(Y_train), " elementos.")
print("El vector X_test tiene ",len(X_test), " elementos.")
print("El vector Y_test tiene ",len(Y_test), " elementos.")
# -
## 5) CONSTRUIR UN MODELO
# En este ejemplo vamos a elegir un modelo de regresión lineal simple para "X_train"
regresor = LinearRegression()
regresor.fit(X_train, Y_train)
# Se grafican los vectores de entrenamiento.
plt.scatter(X_train, Y_train, color = 'red')
plt.plot(X_train, regresor.predict(X_train), color = 'blue')
plt.title('Datos de entrenamiento')
plt.xlabel('Años de experiencia')
plt.ylabel('Salario')
plt.show()
# Se grafican los vectores de prueba.
plt.scatter(X_test, Y_test, color = 'red') #Puntos utilizados para la prueba del modelo
plt.plot(X_train, regresor.predict(X_train), color = 'blue') #Recta correspondiente a los datos de entrenamiento
plt.title('Datos de prueba')
plt.xlabel('Años de experiencia')
plt.ylabel('Salario')
plt.show()
## 6) PREDICCIONES
# Se hacen las predicciones con "X_test"
Y_pred = regresor.predict(X_test)
#print(Y_pred)
print("El vector Y_pred tiene ",len(Y_pred), " elementos.")
# Se grafican los resultados de la predicción.
plt.scatter(X_test, Y_pred, color = 'red')
plt.plot(X_train, regresor.predict(X_train), color = 'blue') #Recta correspondiente a los datos de entrenamiento
plt.title('Predicciones')
plt.xlabel('Años de experiencia')
plt.ylabel('Salario')
plt.show()
# #### 7) EVALUACIÓN DEL MODELO
# Veamos cómo se comporta el modelo:
#
# 7.1 Calcular $R^{2}$
#
# 7.2 Calcular los errores absolutos $(real - estimado)$ y graficarlos
#
# 7.3 Calcular los errores relativos $\left( \dfrac{\text{real - estimado}}{\text{real}} \right)$ y graficarlos
#
# 7.4 Graficar valores estimados vs valores reales
#
# 7.5 Calcular el error cuadrático: $(real − estimado)^{2}$
#
# 7.6 Calcular el error cuadrático medio: $\dfrac{\displaystyle \sum_{i = 1}^{n} (real_{i} − estimado_{i})^{2}}{n}$
#
#7.1 Calcular R^2
r_cuadrada = r2_score(Y_test,Y_pred)
round(r_cuadrada,3) #Porcentaje de los datos explicados por el modelo
# +
#7.2 Calcular los errores absolutos (real - estimado) y graficarlos
err_abs = Y_test-Y_pred
print(np.around(err_abs,2))
X = range(1,len(err_abs)+1)
plt.scatter(X, err_abs, color = 'blue')
plt.plot(X, np.zeros(len(err_abs)), color = 'red') #Recta en Y = 0
plt.title('Errores absolutos (real - estimado)')
plt.ylabel('Errores absolutos')
plt.show()
# +
#7.3 Calcular los errores relativos [(real - estimado)/real] y graficarlos
err_rel = err_abs/Y_test
print(np.around(err_rel,3))
X = range(1,len(err_rel)+1)
plt.scatter(X, err_rel, color = 'blue')
plt.plot(X, np.zeros(len(err_abs)), color = 'red') #Recta en Y = 0
plt.title('Errores relativos [(real - estimado)/real]')
plt.ylabel('Errores relativos')
plt.show()
# -
#7.4 Graficar valores estimados vs valores reales
X = range(1,len(Y_test)+1)
plt.plot(X, Y_test, color = 'blue') #Recta de valores reales
plt.plot(X, Y_pred, color = 'green') #Recta de valores estimados
plt.title('Valores estimados vs valores reales')
plt.ylabel('Salario')
plt.show()
#7.5 Calcular el error cuadrático = (real − estimado)^2
#print(np.around(err_abs,2))
err_cuad = pow(err_abs,2)
print(err_cuad)
#7.6 Calcular el error cuadrático medio = (1/n) * \sum (real − estimado)^2
'''
Indica qué tan cerca está la línea de la regresión lineal de los valores estimados.
i) Se elevan al cuadrado los errores absolutos.
ii) Se suman.
iii) Se divide el resultado entre el número de datos estimados.
'''
err_cuad_medio = mean_squared_error(Y_test, Y_pred)
print(round(err_cuad_medio,2))
print(round(math.sqrt(err_cuad_medio),2))#Raíz cuadrada del error cuadrático medio
#Graficamos los errores cuadráticos
X = range(1,len(err_cuad)+1)
Y= np.repeat(err_cuad_medio, len(err_cuad))
plt.scatter(X, err_cuad, color = 'blue')
plt.plot(X,Y , color = 'red') #Recta en Y = err_cuad_medio
plt.title('Errores cuadráticos: (real − estimado)^2')
plt.ylabel('Errores cuadráticos')
plt.show()
# ### Ejemplo: Regresión Lineal Múltiple
# +
## 1) EXTRAER DATOS
# Los datos pueden encontrarse en diferentes formatos, en nuestro caso están en formato csv.
# Se carga la base de datos y se definen las varibles X y Y
datos = pd.read_csv('50_Startups.csv') #Se encuentra en la misma carpeta que el jupyter notebook
X = pd.DataFrame(datos.iloc[:, :-1].values) #Se convierte en data frame
X.columns = datos.columns[:-1] #Ponemos los nombres de las columnas
Y = datos.iloc[:, 4].values #Ganancia
print("type(datos): ",type(datos))
print("type(X)",type(X))
print("type(Y)",type(Y))
# +
## 2) ANÁLISIS EXPLORATORIO
# Se realiza una descripción analítica de los datos.
# Se muestran los primeros 5 datos del data frame
datos.head()
# Los datos corresponden a la información de 50 empresas nuevas. La información que se tiene de cada una de ellas es:
#R&D = Research and development: Gastos de investigación y desarrollo
#Administration: Gastos administrativos
#Marketing Spend: Gastos en las técnicas de comercialización de algún producto
#State: Estado en el que se encuentra la empresa
#Profit: Ganancia
# -
# Dimensiones del data frame
datos.shape
# Se cuenta el número de NaN's por columna
datos.isnull().sum()
# Cuenta los valores repetidos de la columna 1 (R&D Spend)
datos["R&D Spend"].value_counts().head()
# Cuenta los valores repetidos de la columna 4 (R&D Spend)
datos["State"].value_counts()
#Se muestran las variables dummies
pd.get_dummies(X['State']).head()
# +
'''Se convierte la columna "State" en una columna categórica.
Se elimina la columna de "California" porque se puede obtener su valor cuando las
columnas "Florida" y "New York" son ambas 0 en el i-ésimo renglón.'''
estados = pd.get_dummies(X['State'],drop_first=True)
estados.head()
# -
#Se cambia la columna "State" por las variables dummies creadas
print(X.head())
X=X.drop('State',axis=1)
X=pd.concat([X,estados],axis=1)
X.head()
# +
## 3) VISUALIZACIÓN DE LOS DATOS
# Para entender mejor los datos es necesario graficarlos.
sns.distplot(Y)#Ganancia
# +
#R&D = Research and development: Gastos de investigación y desarrollo
print(min(X['R&D Spend']))
print(max(X['R&D Spend']))
#Histograma de la columna "R&D Spend"
plt.hist(X['R&D Spend'], bins=[0,25000,50000,75000,100000,
125000,150000,175000]) #División cada 25 mil
plt.title('Histograma de la columna "R&D Spend"')
plt.xlabel('Unidades monetarias')
plt.ylabel('Número de empresas')
plt.show()
# +
#Administration: Gastos administrativos
print(min(X['Administration']))
print(max(X['Administration']))
#Histograma de la columna "Administration"
plt.hist(X['Administration'], bins=[50000,75000,100000,125000,
150000,175000,200000]) #División cada 25 mil
plt.title('Histograma de la columna "Administration"')
plt.xlabel('Unidades monetarias')
plt.ylabel('Número de empresas')
plt.show()
# +
#Marketing Spend: Gastos en las técnicas de comercialización de algún producto
print(min(X['Marketing Spend']))
print(max(X['Marketing Spend']))
#Histograma de la columna "Marketing Spend"
plt.hist(X['Marketing Spend'], bins=[0,50000,100000,150000,200000,250000,
300000,350000,400000,450000,500000]) #División cada 50 mil
plt.title('Histograma de la columna "Marketing Spend"')
plt.xlabel('Unidades monetarias')
plt.ylabel('Número de empresas')
plt.show()
# -
#State: Estado en el que se encuentra la empresa
#Histograma de la columna "State"
plt.hist(datos['State'])
plt.title('Histograma de la columna "State"')
plt.xlabel('Estado')
plt.ylabel('Número de empresas') #17 17 16
plt.show()
sns.boxplot(x="R&D Spend", data=datos)
round(np.mean(X['R&D Spend']),2) #Promedio de gastos por investigación y desarrollo
sns.boxplot(x="Administration", data=datos)
round(np.mean(X['Administration']),2) #Promedio de gastos por administración
sns.boxplot(x="Marketing Spend", data=datos)
round(np.mean(X['Marketing Spend']),2) #Promedio de gastos por "marketing"
sns.boxplot(x=Y, data=datos)
round(np.mean(Y),2) #Promedio de ganancias
min(Y) #Outlier
#Se muestra la correlación entre las variables
sns.pairplot(datos)
# +
## 4) DIVIDIR LOS DATOS
# Se separan los datos en 2 grupos (usualmente 80% y 20%):
# i) Para entrenar al modelo (80%)
# ii) Para probar el modelo (20%)
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, #Se indican los vectores que se van a dividir
test_size = 0.2, #Se indica el porcentajede los datos para probar el modelo
random_state = 0) #Se fija la semilla
# Nota: Tomar la muestra aleatoria es muy importante porque en caso de que los datos estén
#ordenados el algoritmo no aprende adecuadamente. Por ejemplo si tenemos 80 sanos y 20 enfermos,
#sólo se tomarían los 80 sanos (por ser los primeros 80).
# -
## 5) CONSTRUIR UN MODELO
# En este ejemplo vamos a elegir un modelo de regresión lineal simple para "X_train"
regresor = LinearRegression()
regresor.fit(X_train, Y_train)
## 6) PREDICCIONES
# Se hacen las predicciones con "X_test"
Y_pred = regresor.predict(X_test)
#print(X_test) #data frame
#print(Y_pred)
# +
# Se grafican los resultados de la predicción.
X = range(1,len(Y_pred)+1)
plt.scatter(X, Y_pred, color = 'blue')
plt.title('Predicciones')
plt.ylabel('Ganancia')
plt.show()
#Nota: No estamos graficando contra ninguna variable explicativa (R&D, Administration, Marketing Spend, State).
# -
# #### 7) EVALUACIÓN DEL MODELO
# Veamos cómo se comporta el modelo:
#
# 7.1 Calcular $R^{2}$ ajustada $ = 1 - \dfrac{(1 - R^{2}) (n-1)}{n - p - 1}$, donde
#
# $R^{2}:$ R cuadrada de los datos
#
# $n:$ Número de datos para entrenar al modelo
#
# $p:$ Número de variables independientes
#
# 7.2 Calcular los errores absolutos $(real - estimado)$ y graficarlos
#
# 7.3 Calcular los errores relativos $\left( \dfrac{\text{real - estimado}}{\text{real}} \right)$ y graficarlos
#
# 7.4 Graficar valores estimados vs valores reales
#
# 7.5 Calcular el error cuadrático: $(real − estimado)^{2}$
#
# 7.6 Calcular el error cuadrático medio: $\dfrac{\displaystyle \sum_{i = 1}^{n} (real_{i} − estimado_{i})^{2}}{n}$
#
# +
#7.1 Calcular R^2 ajustada
r_cuadrada = r2_score(Y_test,Y_pred)
print('R^2 = ',round(r_cuadrada,3)) #Porcentaje de los datos explicados por el modelo
#R^2 ajustada
n = len(Y_train)
p = X_train.shape[1]
r_cuad_aj = 1 - (((1-r_cuadrada)*(n-1))/(n-p-1))
print('n = ',n)
print('p = ',p)
print('r_cuad_aj = ',round(r_cuad_aj,3))
# +
#7.2 Calcular los errores absolutos (real - estimado) y graficarlos
err_abs = Y_test-Y_pred
print(np.around(err_abs,2))
X = range(1,len(err_abs)+1)
plt.scatter(X, err_abs, color = 'blue')
plt.plot(X, np.zeros(len(err_abs)), color = 'red') #Recta en Y = 0
plt.title('Errores absolutos (real - estimado)')
plt.ylabel('Errores absolutos')
plt.show()
# +
#7.3 Calcular los errores relativos [(real - estimado)/real] y graficarlos
err_rel = err_abs/Y_test
print(np.around(err_rel,3))
X = range(1,len(err_rel)+1)
plt.scatter(X, err_rel, color = 'blue')
plt.plot(X, np.zeros(len(err_abs)), color = 'red') #Recta en Y = 0
plt.title('Errores relativos [(real - estimado)/real]')
plt.ylabel('Errores relativos')
plt.show()
# -
#7.4 Graficar valores estimados vs valores reales
X = range(1,len(Y_test)+1)
plt.plot(X, Y_test, color = 'blue') #Recta de valores reales
plt.plot(X, Y_pred, color = 'green') #Recta de valores estimados
plt.title('Valores estimados vs valores reales')
plt.ylabel('Ganancia')
plt.show()
#7.5 Calcular el error cuadrático = (real − estimado)^2
#print(np.around(err_abs,2))
err_cuad = pow(err_abs,2)
print(err_cuad)
#Graficamos los errores cuadráticos
X = range(1,len(err_cuad)+1)
Y= np.repeat(err_cuad_medio, len(err_cuad))
plt.scatter(X, err_cuad, color = 'blue')
plt.plot(X,Y , color = 'red') #Recta en Y = err_cuad_medio
plt.title('Errores cuadráticos: (real − estimado)^2')
plt.ylabel('Errores cuadráticos')
plt.show()
#7.6 Calcular el error cuadrático medio = (1/n) * \sum (real − estimado)^2
'''
Indica qué tan cerca está la línea de la regresión lineal de los valores estimados.
i) Se elevan al cuadrado los errores absolutos.
ii) Se suman.
iii) Se divide el resultado entre el número de datos estimados.
'''
err_cuad_medio = mean_squared_error(Y_test, Y_pred)
print(round(err_cuad_medio,2))
print(round(math.sqrt(err_cuad_medio),2))#Raíz cuadrada del error cuadrático medio
| M1/.ipynb_checkpoints/1_4 Scikit-learn-Linear Regression-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from matplotlib import pyplot as plt
import numpy as np
import seaborn as sns
import glob
all_files = sorted(glob.glob('data/[0-9]*.csv'))
filename = all_files[2]
dates = ['Monday, 2019-02-08', 'Tuesday, 2019-02-09',
'Wednesday, 2019-02-10', 'Thursday, 2019-02-08',
'Friday, 2019-02-08']
current_date = dates[2]
filename
# +
df = pd.read_csv(filename, index_col=None, sep= ';', header=0)
# create dummy column
df['dummy']= 1
# create datetime index
df.timestamp = pd.to_datetime(df['timestamp'])
df = df.set_index('timestamp')
# create 4 different dataframes for the location and then stack on top of each other:
df_dairy = df[df.location == "dairy"]
df_drinks = df[df.location == "drinks"]
df_fruit = df[df.location == "fruit"]
df_spices = df[df.location == "spices"]
# groupby location
df_dairy_plot = df_dairy.groupby('timestamp')[['dummy']].count().resample(rule='30MIN').sum().reset_index()
df_drinks_plot = df_drinks.groupby('timestamp')[['dummy']].count().resample(rule='30MIN').sum().reset_index()
df_fruit_plot = df_fruit.groupby('timestamp')[['dummy']].count().resample(rule='30MIN').sum().reset_index()
df_spices_plot = df_spices.groupby('timestamp')[['dummy']].count().resample(rule='30MIN').sum().reset_index()
sns.set_style("whitegrid")
plt.rcParams['figure.figsize'] = (13, 7)
# Values of each group
bars1 = list(df_dairy_plot.dummy)
bars2 = list(df_drinks_plot.dummy)
bars3 = list(df_fruit_plot.dummy)
bars4 = list(df_spices_plot.dummy)
# Bottoms
height_bars_1 = bars1
height_bars_1_2 = np.add(bars1, bars2).tolist()
height_bars_1_2_3 = np.add(height_bars_1_2, bars3).tolist()
# The position of the bars on the x-axis
xtick_pos = list(range(1,31))
# Names of group and bar width
barWidth = 1
labels = ['7:00', '7:30', '8:00', '8:30', '9:00', '9:30', '10:00', '10:30',
'11:00', '11:30', '12:00', '12:30', '13:00', '13:30', '14:00', '14:30',
'15:00', '15:30', '16:00', '16:30', '17:00', '17:30', '18:00', '18:30',
'19:00', '19:30', '20:00', '20:30', '21:00', '21:30']
# Create bottom bars
plot_dairy = plt.bar(xtick_pos, bars1,
color='darkgrey', width=barWidth, label='dairy')
# Create middle bars (1) , on top of the first ones
plot_drinks = plt.bar(xtick_pos, bars2, bottom=height_bars_1,
color='blue', width=barWidth, label='drinks')
# Create middle bars (2) , on top of the second ones
plot_fruit = plt.bar(xtick_pos, bars3, bottom=height_bars_1_2,
color='darkorange',width=barWidth, label='fruit')
# Create top bars , on top of the third ones
plot_spices = plt.bar(xtick_pos, bars4, bottom=height_bars_1_2_3,
color='green', width=barWidth, label='spices')
# Custom X axis
plt.xticks(xtick_pos, labels, fontweight='normal',rotation=90)
plt.xlabel("Time of day", fontsize=13)
plt.ylabel('Customer count', fontsize=13)
plt.xlim(xmin=0.5, xmax=30.5)
plt.legend(handles=[plot_dairy, plot_drinks, plot_fruit, plot_spices],
bbox_to_anchor=(1.0, 1.015), loc='upper left', fontsize=13)
# create titles
print("Saving: ", current_date)
plt.title(f'Distribution of customer count in the supermark sections over time ({current_date})', fontsize=15)
#plt.show()
# save figure
plt.savefig(f'plots/cus_dist_in_sections_overtime_{current_date}.png', dpi=300)
# -
| 04_stacked_barplots_customer_in_location_over_time.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load in
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import random
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
from plotly.offline import download_plotlyjs, init_notebook_mode, plot, iplot
import plotly.offline as py
py.init_notebook_mode(connected=True)
import plotly.graph_objs as go
# Input data files are available in the "../input/" directory.
# For example, running this (by clicking run or pressing Shift+Enter) will list the files in the input directory
from subprocess import check_output
# +
print(check_output(["ls", "./"]).decode("utf8"))
# Any results you write to the current directory are saved as output.
# -
## Loading events dataset
df_events = pd.read_csv("./events.csv")
## Loading ginf dataset, which has some important data to merge with event dataset
df_ginf = pd.read_csv("./ginf.csv")
## Selecting only the features I need. `id_odsp` serves as a unique identifier that will be used to
## union the 2 datasets
df_ginf = df_ginf[['id_odsp', 'date', 'league', 'season', 'country']]
## Join the 2 datasets to add 'date', 'league', 'season', and 'country' information to the main dataset
df_events = df_events.merge(df_ginf, how='left')
## Naming the leagues with their popular names, which will make thinks much clear for us
leagues = {'E0': 'Premier League', 'SP1': 'La Liga',
'I1': 'Serie A', 'F1': 'League One', 'D1': 'Bundesliga'}
## Apply the mapping
df_events['league'] = df_events['league'].map(leagues)
# +
## Events type 1
event_type_1 = pd.Series([
'Announcement',
'Attempt',
'Corner',
'Foul',
'Yellow card',
'Second yellow card',
'Red card',
'Substitution',
'Free kick won',
'Offside',
'Hand ball',
'Penalty conceded'], index=[[item for item in range(0, 12)]])
## Events type 2
event_type2 = pd.Series(['Key Pass', 'Failed through ball', 'Sending off', 'Own goal'],
index=[[item for item in range(12, 16)]])
## Match side
side = pd.Series(['Home', 'Away'], index=[[item for item in range(1, 3)]])
## Shot place
shot_place = pd.Series([
'Bit too high',
'Blocked',
'Bottom left corner',
'Bottom right corner',
'Centre of the goal',
'High and wide',
'Hits the bar',
'Misses to the left',
'Misses to the right',
'Too high',
'Top centre of the goal',
'Top left corner',
'Top right corner'
], index=[[item for item in range(1, 14)]])
## Outcome of shot
shot_outcome = pd.Series(['On target', 'Off target', 'Blocked', 'Hit the bar'],
index=[[item for item in range(1, 5)]])
## Location of shot
location = pd.Series([
'Attacking half',
'Defensive half',
'Centre of the box',
'Left wing',
'Right wing',
'Difficult angle and long range',
'Difficult angle on the left',
'Difficult angle on the right',
'Left side of the box',
'Left side of the six yard box',
'Right side of the box',
'Right side of the six yard box',
'Very close range',
'Penalty spot',
'Outside the box',
'Long range',
'More than 35 yards',
'More than 40 yards',
'Not recorded'
],
index=[[item for item in range(1, 20)]])
## Players' body part
bodypart = pd.Series(['right foot', 'left foot', 'head'], index=[[item for item in range(1, 4)]])
## Assist method
assist_method = pd.Series(['None', 'Pass', 'Cross', 'Headed pass', 'Through ball'],
index=[item for item in range(0, 5)])
## Situation
situation = pd.Series(['Open play', 'Set piece', 'Corner', 'Free kick'],
index=[item for item in range(1, 5)])
# -
## Utility function to plot bar plots with similar configuration
def plot_barplot(data, x_ticks, x_labels, y_labels, title, color='muted', num=10, figsize=8):
## Set standard style as whitegrid (this also could be customized via param)
sns.set_style("whitegrid") # Proposed themes: darkgrid, whitegrid, dark, white, and ticks
## Set a figure with custom figsize
plt.figure(figsize=(num, figsize))
## Plottin data
ax = sns.barplot(x = [j for j in range(0, len(data))], y=data.values, palette=color)
## Setting ticks extracted from data indexes
ax.set_xticks([j for j in range(0, len(data))])
## Set labels of the chart
ax.set_xticklabels(x_ticks, rotation=45)
ax.set(xlabel = x_labels, ylabel = y_labels, title = title)
ax.plot();
plt.tight_layout()
# +
## Count of events occurecies
events_series = df_events['event_type'].value_counts()
## Plotting chart
plot_barplot(events_series, event_type_1.values,
"Event type", "Number of events", "Event types", 'Accent_r', 10, 5)
# +
## Filtering out dataframe to extract attemtps which resulted in goals
df_shot_places = df_events[(df_events['event_type'] == 1) &
(df_events['is_goal'] == 1)]['shot_place'].value_counts()
## Plotting the chart
plot_barplot(df_shot_places, shot_place[[3, 4, 5, 13, 12]], 'Shot places', 'Number of events',
'Shot places resulting in goals', 'RdYlBu_r', 8, 5)
# +
## Filtering out dataframe to extract attemtps which resulted in goals
df_shot_places = df_events[(df_events['event_type'] == 1) &
(df_events['is_goal'] == 0)]['shot_place'].value_counts()
## Plotting the chart
plot_barplot(df_shot_places, shot_place, 'Shot places', 'Number of events',
'Shot places no resulting in goals', 'BuGn_r', 8, 5)
# +
## Copying original dataframe
df_shot_places_ed = df_events.copy()
## Grouping data by shot places
df_shot_places_ed = df_events.groupby('shot_place', as_index=False).count().sort_values('id_event',
ascending=False).dropna()
## Mapping dataframe index to shot places labels available in the dictionary file
df_shot_places_ed['shot_place'] = df_shot_places_ed['shot_place'].map(shot_place)
## Plotting the chart
plot_barplot(df_shot_places_ed['id_event'], df_shot_places_ed['shot_place'],
'Shot places',
'Number of events',
'Shot places',
'BuGn_r', 8, 5)
# +
## Grouping attempts by team
grouping_by_offensive = df_events[df_events['is_goal']==1].groupby('event_team')
## Sorting the values
grouping_by_offensive = grouping_by_offensive.count().sort_values(by='id_event', ascending=False)[:10]
teams = grouping_by_offensive.index
scores = grouping_by_offensive['id_event']
## Plotting the teams
plot_barplot(scores, teams, 'Teams', '# of goals', 'Most offensive teams', 'GnBu_d', 8, 6)
# +
## Grouping attempts by team
grouping_by_offensive = df_events[df_events['is_goal']==1].groupby('event_team')
## Sorting the values
grouping_by_offensive = grouping_by_offensive.count().sort_values(by='id_event', ascending=True)[:10]
teams = grouping_by_offensive.index
scores = grouping_by_offensive['id_event']
## Plotting the teams
plot_barplot(scores, teams, 'Teams', '# of goals', 'Less offensive teams', 'GnBu_d', 8, 6)
# +
## grouping by player when is goal
grouping_by_offensive_player = df_events[df_events['is_goal']==1].groupby('player')
## Couting and sorting the number of goals by player, then pick the top 10
grouping_by_offensive_player = grouping_by_offensive_player.count().sort_values(by='id_event',
ascending=False)[:10]
## Extracting player names
players = grouping_by_offensive_player.index
## Extracting values (# of goals)
scores = grouping_by_offensive_player['id_event']
## Plotting the chart
plot_barplot(scores, players, 'Players', '# of score', 'Most offensive players', 'GnBu_d', 8, 6)
# +
## Loading events dataset
df_events1 = pd.read_csv("./events.csv")
## Loading ginf dataset, which has some important data to merge with event dataset
df_ginf1 = pd.read_csv("./ginf.csv")
## Selecting only the features I need. `id_odsp` serves as a unique identifier that will be used to
## union the 2 datasets
## Join the 2 datasets to add 'date', 'league', 'season', and 'country' information to the main dataset
df_events1 = df_events1.merge(df_ginf1, how='left')
df_events1 = df_events1[['id_odsp', 'id_event', 'league', 'season', 'ht', 'at', 'event_team', 'is_goal']]
## Naming the leagues with their popular names, which will make thinks much clear for us
leagues = {'E0': 'Premier League', 'SP1': 'La Liga',
'I1': 'Serie A', 'F1': 'League One', 'D1': 'Bundesliga'}
## Apply the mapping
df_events1['league'] = df_events1['league'].map(leagues)
# cats = ['id_odsp', 'id_event', 'league', 'season', 'ht', 'at', 'event_team']
# d = dict.fromkeys(cats,'category')
# df_events1 = df_events1.astype(d)
# -
df_events1.info()
import warnings
warnings.filterwarnings('ignore')
Leagues = df_events1['league'].unique()
Seasons = list(df_events1['season'].unique())
print(Leagues)
print(Seasons)
df_events1[df_events1['league']== 'Bundesliga'][df_events1['season'] == 2012]['ht'].unique()
for league in Leagues[::-1]:
if league == 'Premier League' :
print('No details on {}'.format(league))
else:
for season in Seasons[::-1]:
print('****Informations about {}'.format(league) + ' ' +'{} ****'.format(season))
Stats = []
Teams = df_events1[df_events1['league']== league][df_events1['season'] == season]['ht'].unique()
Games = df_events1[df_events1['league']== league][df_events1['season'] == season]['id_odsp'].unique()
for game in Games:
Events = df_events1[df_events1['league']== league][df_events1['season'] == season][df_events1['id_odsp']==game]
ht= Events.iloc[1,4]
at= Events.iloc[1,5]
Butat=0
Butht=0
for j in range(1,Events.shape[0]):
if Events.iloc[j,7] == 1 :
if Events.iloc[j,6] == ht:
Butat +=1
else:
Butht +=1
item = [ht, at, Butht, Butat]
Stats.append(item)
Stats = np.array(Stats)
df_Stats = pd.DataFrame({'Teamht': Stats[:,0], 'Butht':Stats[:,2], 'Teamat': Stats[:,1], 'But at':Stats[:,3] })
results = []
for team in Teams:
data_team = df_Stats.loc[(df_Stats['Teamht'] == team)|(df_Stats['Teamat'] == team)]
nbrgoals = 0
for j in range(1, data_team.shape[0]):
if data_team.iloc[j,2]== team:
nbrgoals += int(data_team.iloc[j,0])
else:
nbrgoals += int(data_team.iloc[j,1])
elem = [team, nbrgoals]
results.append(elem)
results = np.array(results)
ids = np.argsort(results[:,1])[::-1]
results[:,1] = results[:,1][ids]
results[:,0] = results[:,0][ids]
df_results = pd.DataFrame({'ATeam': results[:,0], 'ButsE': results[:,1]})
print(df_results)
Leagues
# +
## grouping by player when is goal
goal = df_events[df_events['is_goal']==1]
## Plotting the hist
# fig=plt.figure(figsize=(13,10))
plt.hist(goal.time, 100)
plt.xlabel("TIME",fontsize=10)
plt.ylabel("# of goals",fontsize=10)
plt.title("goal counts vs time",fontsize=15)
x=goal.groupby(by='time')['time'].count().sort_values(ascending=False).index[0]
y=goal.groupby(by='time')['time'].count().sort_values(ascending=False).iloc[0]
x1=goal.groupby(by='time')['time'].count().sort_values(ascending=False).index[1]
y1=goal.groupby(by='time')['time'].count().sort_values(ascending=False).iloc[1]
plt.text(x=x-10,y=y+10,s='time:'+str(x)+',max:'+str(y),fontsize=12,fontdict={'color':'red'})
plt.text(x=x1-10,y=y1+10,s='time:'+str(x1)+',the 2nd max:'+str(y1),fontsize=12,fontdict={'color':'black'})
plt.show()
# -
redCards = df_events[df_events['event_type'] == 6]['event_team']
# +
## Count of events occurecies
redCards_series = redCards.value_counts()[:10]
## Plotting chart
plot_barplot(redCards_series, redCards_series.index,
"Event_team", "Number of Red Cards", "Red Cards per team", 'gist_earth', 10, 5)
# -
yellowCards = df_events[df_events['event_type'] == (4 or 5)]['event_team']
# +
## Count of events occurecies
yellowCards_series = yellowCards.value_counts()[:10]
## Plotting chart
plot_barplot(yellowCards_series, yellowCards_series.index,
"Event_team", "Number of yellow Cards", "Yellow Cards per team", 'gist_earth', 10, 5)
# -
# When do the red cards occur?
reds = plt.hist(redCards.event_team, 100, color="red")
plt.xlabel("Time")
plt.ylabel("Red Cards")
plt.title("When Red Cards Occur")
plt.show()
# Penalties
penalties=df_events[df_events["location"]==14]
# Check the shot place
for i in range(14):
if sum(penalties["shot_place"]==i)==0:
print(i)
# +
top_left=sum(penalties["shot_place"]==12)
bot_left=sum(penalties["shot_place"]==3)
top_right=sum(penalties["shot_place"]==13)
bot_right=sum(penalties["shot_place"]==4)
centre=sum(penalties["shot_place"]==5)+sum(penalties["shot_place"]==11)
missed=sum(penalties["shot_place"]==1)+sum(penalties["shot_place"]==6)+sum(penalties["shot_place"]==7)+sum(penalties["shot_place"]==8)+sum(penalties["shot_place"]==9)+sum(penalties["shot_place"]==10)
labels_pen=["top left","bottom left","centre","top right","bottom right","missed"]
num_pen=[top_left,bot_left,centre,top_right,bot_right,missed]
colors_pen=["red", "aqua","royalblue","yellow","violet","m"]
plt.pie(num_pen,labels=labels_pen,colors=colors_pen,autopct='%1.1f%%',startangle=60,explode=(0,0,0,0,0,0.2))
plt.axis('equal')
plt.title("Percentage of each placement of penalties",fontsize=18,fontweight="bold")
fig=plt.gcf()
fig.set_size_inches(8,6)
plt.show()
# +
# success rate of penalties
scored_pen=penalties[penalties["is_goal"]==1]
pen_rightfoot=scored_pen[scored_pen["bodypart"]==1].shape[0]
pen_leftfoot=scored_pen[scored_pen["bodypart"]==2].shape[0]
penalty_combi=pd.DataFrame({"right foot":pen_rightfoot,"left foot":pen_leftfoot},index=["Scored"])
penalty_combi.plot(kind="bar")
plt.title("Penalties scored (Right/Left foot)",fontsize=14,fontweight="bold")
penalty_combi
# -
def pen_stats(player):
player_pen=penalties[penalties["player"]==player]
right_attempt=player_pen[player_pen["bodypart"]==1]
right_attempt_scored=right_attempt[right_attempt["is_goal"]==1].shape[0]
right_attempt_missed=right_attempt[right_attempt["is_goal"]==0].shape[0]
left_attempt=player_pen[player_pen["bodypart"]==2]
left_attempt_scored=left_attempt[left_attempt["is_goal"]==1].shape[0]
left_attempt_missed=left_attempt[left_attempt["is_goal"]==0].shape[0]
scored=pd.DataFrame({"right foot":right_attempt_scored,"left foot":left_attempt_scored},index=["Scored"])
missed=pd.DataFrame({"right foot":right_attempt_missed,"left foot":left_attempt_missed},index=["Missed"])
combi=scored.append(missed)
return combi
pen_stats("<NAME>")
pen_stats("<NAME>")
pen_stats("<NAME>")
pen_stats('<NAME>')
# +
# for player in list(players):
# print(player+ '\n')
# print(pen_stats(player))
# +
# penalties["player"].unique()
# -
def pen_full_stats(player):
player_pen=penalties[penalties["player"]==player]
scored_pen=player_pen[player_pen["is_goal"]==1]
missed_pen=player_pen[player_pen["is_goal"]==0]
top_left_rightfoot=scored_pen[scored_pen["shot_place"]==12][scored_pen["bodypart"]==1].shape[0]
top_left_leftfoot=scored_pen[scored_pen["shot_place"]==12][scored_pen["bodypart"]==2].shape[0]
bot_left_rightfoot=scored_pen[scored_pen["shot_place"]==3][scored_pen["bodypart"]==1].shape[0]
bot_left_leftfoot=scored_pen[scored_pen["shot_place"]==3][scored_pen["bodypart"]==2].shape[0]
top_right_rightfoot=scored_pen[scored_pen["shot_place"]==13][scored_pen["bodypart"]==1].shape[0]
top_right_leftfoot=scored_pen[scored_pen["shot_place"]==13][scored_pen["bodypart"]==2].shape[0]
bot_right_rightfoot=scored_pen[scored_pen["shot_place"]==4][scored_pen["bodypart"]==1].shape[0]
bot_right_leftfoot=scored_pen[scored_pen["shot_place"]==4][scored_pen["bodypart"]==2].shape[0]
centre_rightfoot=scored_pen[scored_pen["shot_place"]==5][scored_pen["bodypart"]==1].shape[0]+scored_pen[scored_pen["shot_place"]==11][scored_pen["bodypart"]==1].shape[0]
centre_leftfoot=scored_pen[scored_pen["shot_place"]==5][scored_pen["bodypart"]==2].shape[0]+scored_pen[scored_pen["shot_place"]==11][scored_pen["bodypart"]==2].shape[0]
scored_without_recorded_loc_rightfoot=scored_pen[scored_pen["shot_place"].isnull()][scored_pen["bodypart"]==1].shape[0]
scored_without_recorded_loc_leftfoot=scored_pen[scored_pen["shot_place"].isnull()][scored_pen["bodypart"]==2].shape[0]
missed_rightfoot=missed_pen[missed_pen["bodypart"]==1].shape[0]
missed_leftfoot=missed_pen[missed_pen["bodypart"]==2].shape[0]
right_foot=pd.DataFrame({"Top Left Corner":top_left_rightfoot,"Bottom Left Corner":bot_left_rightfoot,"Top Right Corner":top_right_rightfoot,"Bottom Right Corner":bot_right_rightfoot,"Centre":centre_rightfoot,"Unrecorded placement":scored_without_recorded_loc_rightfoot,"Missed":missed_rightfoot},index=["Right Foot attempt"])
left_foot=pd.DataFrame({"Top Left Corner":top_left_leftfoot,"Bottom Left Corner":bot_left_leftfoot,"Top Right Corner":top_right_leftfoot,"Bottom Right Corner":bot_right_leftfoot,"Centre":centre_leftfoot,"Unrecorded placement":scored_without_recorded_loc_leftfoot,"Missed":missed_leftfoot},index=["Left Foot attempt"])
fullstats=right_foot.append(left_foot)
fullstats=fullstats[["Top Right Corner","Bottom Right Corner","Top Left Corner","Bottom Left Corner","Centre","Unrecorded placement","Missed"]]
return fullstats
import warnings
warnings.filterwarnings('ignore')
pen_full_stats("lionel messi")
pen_full_stats("<NAME>")
def stats(player):
player_pen=df_events[df_events["player"]==player]
right_attempt=player_pen[player_pen["bodypart"]==1]
right_attempt_scored=right_attempt[right_attempt["is_goal"]==1].shape[0]
right_attempt_missed=right_attempt[right_attempt["is_goal"]==0].shape[0]
left_attempt=player_pen[player_pen["bodypart"]==2]
left_attempt_scored=left_attempt[left_attempt["is_goal"]==1].shape[0]
left_attempt_missed=left_attempt[left_attempt["is_goal"]==0].shape[0]
head_attempt=player_pen[player_pen["bodypart"]==3]
head_attempt_scored=head_attempt[head_attempt["is_goal"]==1].shape[0]
head_attempt_missed=head_attempt[head_attempt["is_goal"]==0].shape[0]
scored=pd.DataFrame({"right foot":right_attempt_scored,"left foot":left_attempt_scored, "head": head_attempt_scored},index=["Scored"])
missed=pd.DataFrame({"right foot":right_attempt_missed,"left foot":left_attempt_missed, "head": head_attempt_missed},index=["Missed"])
combi=scored.append(missed)
return combi
stats('lionel messi')
stats("cristiano ronaldo")
def full_stats(player):
player_pen=df_events[df_events["player"]==player]
scored_pen=player_pen[player_pen["is_goal"]==1]
missed_pen=player_pen[player_pen["is_goal"]==0]
top_left_rightfoot=scored_pen[scored_pen["shot_place"]==12][scored_pen["bodypart"]==1].shape[0]
top_left_leftfoot=scored_pen[scored_pen["shot_place"]==12][scored_pen["bodypart"]==2].shape[0]
top_left_head = scored_pen[scored_pen["shot_place"]==12][scored_pen["bodypart"]==3].shape[0]
bot_left_rightfoot=scored_pen[scored_pen["shot_place"]==3][scored_pen["bodypart"]==1].shape[0]
bot_left_leftfoot=scored_pen[scored_pen["shot_place"]==3][scored_pen["bodypart"]==2].shape[0]
bot_left_head = scored_pen[scored_pen["shot_place"]==3][scored_pen["bodypart"]==3].shape[0]
top_right_rightfoot=scored_pen[scored_pen["shot_place"]==13][scored_pen["bodypart"]==1].shape[0]
top_right_leftfoot=scored_pen[scored_pen["shot_place"]==13][scored_pen["bodypart"]==2].shape[0]
top_right_head = scored_pen[scored_pen["shot_place"]==13][scored_pen["bodypart"]==3].shape[0]
bot_right_rightfoot=scored_pen[scored_pen["shot_place"]==4][scored_pen["bodypart"]==1].shape[0]
bot_right_leftfoot=scored_pen[scored_pen["shot_place"]==4][scored_pen["bodypart"]==2].shape[0]
bot_right_head = scored_pen[scored_pen["shot_place"]==4][scored_pen["bodypart"]==3].shape[0]
centre_rightfoot=scored_pen[scored_pen["shot_place"]==5][scored_pen["bodypart"]==1].shape[0]+scored_pen[scored_pen["shot_place"]==11][scored_pen["bodypart"]==1].shape[0]
centre_leftfoot=scored_pen[scored_pen["shot_place"]==5][scored_pen["bodypart"]==2].shape[0]+scored_pen[scored_pen["shot_place"]==11][scored_pen["bodypart"]==2].shape[0]
centre_head = scored_pen[scored_pen["shot_place"]==5][scored_pen["bodypart"]==3].shape[0]
scored_without_recorded_loc_rightfoot=scored_pen[scored_pen["shot_place"].isnull()][scored_pen["bodypart"]==1].shape[0]
scored_without_recorded_loc_leftfoot=scored_pen[scored_pen["shot_place"].isnull()][scored_pen["bodypart"]==2].shape[0]
scored_without_recorded_loc_head=scored_pen[scored_pen["shot_place"].isnull()][scored_pen["bodypart"]==3].shape[0]
missed_rightfoot=missed_pen[missed_pen["bodypart"]==1].shape[0]
missed_leftfoot=missed_pen[missed_pen["bodypart"]==2].shape[0]
missed_head=missed_pen[missed_pen["bodypart"]==3].shape[0]
right_foot=pd.DataFrame({"Top Left Corner":top_left_rightfoot,"Bottom Left Corner":bot_left_rightfoot,"Top Right Corner":top_right_rightfoot,"Bottom Right Corner":bot_right_rightfoot,"Centre":centre_rightfoot,"Unrecorded placement":scored_without_recorded_loc_rightfoot,"Missed":missed_rightfoot},index=["Right Foot attempt"])
left_foot=pd.DataFrame({"Top Left Corner":top_left_leftfoot,"Bottom Left Corner":bot_left_leftfoot,"Top Right Corner":top_right_leftfoot,"Bottom Right Corner":bot_right_leftfoot,"Centre":centre_leftfoot,"Unrecorded placement":scored_without_recorded_loc_leftfoot,"Missed":missed_leftfoot},index=["Left Foot attempt"])
head=pd.DataFrame({"Top Left Corner":top_left_head,"Bottom Left Corner":bot_left_head,"Top Right Corner":top_right_head,"Bottom Right Corner":bot_right_head,"Centre":centre_head,"Unrecorded placement":scored_without_recorded_loc_head,"Missed":missed_head},index=["Head attempt"])
fullstats=right_foot.append(left_foot.append(head))
fullstats=fullstats[["Top Right Corner","Bottom Right Corner","Top Left Corner","Bottom Left Corner","Centre","Unrecorded placement","Missed"]]
return fullstats
full_stats('<NAME>')
full_stats("<NAME>")
# +
# Most effective player
## Grouping by player when attempt == 1
grouped_by_player = df_events[df_events['event_type'] == 1].groupby('player').count()
## Grouping by player when is goal and attempt == 1
grouped_by_player_goals = df_events[(df_events['event_type'] == 1) &
(df_events['is_goal'] == 1)].groupby('player').count()
## Grouping by player when is not goal and attempt == 1
grouped_by_player_not_goals = df_events[(df_events['event_type'] == 1) &
(df_events['is_goal'] == 0)].groupby('player').count()
## Setting a threshold to filter out players with a small number of attempts, which can lead to a lack
## of consistency in the final result
threshold = grouped_by_player['id_event'].std()
grouped_by_player_is_goal = df_events[df_events['is_goal'] == 1].groupby('player').count()
## filtering players that have at least more than average number of chances to skip noise
## For example, a player that has 2 attempts and 1 goal has a very high effectiveness, even though,
## this particular player has not created many chances for his team
grouped_by_player_is_goal_filtered = grouped_by_player_goals[grouped_by_player_goals['id_event'] > threshold]
grouped_by_players_not_goal_filtered = grouped_by_player_not_goals[grouped_by_player_not_goals['id_event'] > threshold]
## Total number of attemtps
total = grouped_by_players_not_goal_filtered['id_event'] + grouped_by_player_is_goal_filtered['id_event']
## Dividing the total of attempts by the attemtps which ended up in goals
result = total/grouped_by_player_is_goal_filtered['id_event']
## Dropping NaN values
result.dropna(inplace=True)
## Sorting results
sorted_results = result.sort_values(ascending=True)
## Extracting players names
players = sorted_results[:10].index
## Extracting value of effectiveness
effectiveness = sorted_results[:10]
## Plotting results
plot_barplot(effectiveness, players, 'Players',
'# of shots needed to score a goal',
'Most effective players',
'RdBu_r', 8, 6)
# +
# Most effective player
## Grouping by player when attempt == 1
grouped_by_player = df_events[df_events['event_type'] == 1].groupby('event_team').count()
## Grouping by player when is goal and attempt == 1
grouped_by_player_goals = df_events[(df_events['event_type'] == 1) &
(df_events['is_goal'] == 1)].groupby('event_team').count()
## Grouping by player when is not goal and attempt == 1
grouped_by_player_not_goals = df_events[(df_events['event_type'] == 1) &
(df_events['is_goal'] == 0)].groupby('event_team').count()
## Setting a threshold to filter out players with a small number of attempts, which can lead to a lack
## of consistency in the final result
threshold = grouped_by_player['id_event'].std()
grouped_by_player_is_goal = df_events[df_events['is_goal'] == 1].groupby('event_team').count()
## filtering players that have at least more than average number of chances to skip noise
## For example, a player that has 2 attempts and 1 goal has a very high effectiveness, even though,
## this particular player has not created many chances for his team
grouped_by_player_is_goal_filtered = grouped_by_player_goals[grouped_by_player_goals['id_event'] > threshold]
grouped_by_players_not_goal_filtered = grouped_by_player_not_goals[grouped_by_player_not_goals['id_event'] > threshold]
## Total number of attemtps
total = grouped_by_players_not_goal_filtered['id_event'] + grouped_by_player_is_goal_filtered['id_event']
## Dividing the total of attempts by the attemtps which ended up in goals
result = total/grouped_by_player_is_goal_filtered['id_event']
## Dropping NaN values
result.dropna(inplace=True)
## Sorting results
sorted_results = result.sort_values(ascending=True)
## Extracting players names
players = sorted_results[:10].index
## Extracting value of effectiveness
effectiveness = sorted_results[:10]
## Plotting results
plot_barplot(effectiveness, players, 'Players',
'# of shots needed to score a goal',
'Most effective players')
# +
grouped_by_player = df_events[df_events['event_type'] == 1].groupby('player').count()
rouped_by_player_goals = df_events[(df_events['event_type'] == 1) &
(df_events['is_goal'] == 1)].groupby('player').count()
grouped_by_player_not_goals = df_events[(df_events['event_type'] == 1) &
(df_events['is_goal'] == 0)].groupby('player').count()
threshold = grouped_by_player['is_goal'].std()
grouped_by_player_is_goal = df_events[df_events['is_goal'] == 1].groupby('player').count()
grouped_by_player_is_goal_filtered = grouped_by_player_is_goal[grouped_by_player_is_goal['is_goal'] >= threshold ]
grouped_by_players_not_goal_filtered = grouped_by_player_not_goals[grouped_by_player_not_goals['is_goal'] >= threshold ]
## Total number of attemtps
total = grouped_by_players_not_goal_filtered['id_event'] + grouped_by_player_is_goal_filtered['id_event']
## Dividing the total of attempts by the attemtps which ended up in goals
result = total/grouped_by_player_is_goal_filtered['id_event']
## Dropping NaN values
result.dropna(inplace=True)
## Sorting results
sorted_results = result.sort_values(ascending=True)
## Extracting players names
players = sorted_results[:10].index
## Extracting value of effectiveness
effectiveness = sorted_results[:10]
## Plotting results
plot_barplot(effectiveness, players, 'Players',
'# of shots needed to score a goal',
'Most effective players',
'RdBu_r', 8, 6)
# -
threshold
# +
grouped_by_player = df_events[df_events['event_type'] == 1].groupby('event_team').count()
rouped_by_player_goals = df_events[(df_events['event_type'] == 1) &
(df_events['is_goal'] == 1)].groupby('event_team').count()
grouped_by_player_not_goals = df_events[(df_events['event_type'] == 1) &
(df_events['is_goal'] == 0)].groupby('event_team').count()
threshold = grouped_by_player['is_goal'].std()
grouped_by_player_is_goal = df_events[df_events['is_goal'] == 1].groupby('event_team').count()
grouped_by_player_is_goal_filtered = grouped_by_player_is_goal
grouped_by_players_not_goal_filtered = grouped_by_player_not_goals
## Total number of attemtps
total = grouped_by_players_not_goal_filtered['id_event'] + grouped_by_player_is_goal_filtered['id_event']
## Dividing the total of attempts by the attemtps which ended up in goals
result = total/grouped_by_player_is_goal_filtered['id_event']
## Dropping NaN values
result.dropna(inplace=True)
## Sorting results
sorted_results = result.sort_values(ascending=True)
## Extracting players names
players = sorted_results[:10].index
## Extracting value of effectiveness
effectiveness = sorted_results[:10]
## Plotting results
plot_barplot(effectiveness, players, 'Teams',
'# of shots needed to score a goal',
'Most effective teams',
'Blues_r', 8, 6)
# +
## Creating a dataframe with total of attempts and total goals
result_df = pd.DataFrame({'total': total.dropna(), 'is_goal': grouped_by_player_is_goal_filtered['id_event']})
## Sorting values by total
result_df.sort_values('total', ascending=False, inplace=True)
## Setting style to dark
sns.set(style="darkgrid")
## Creating figure
f, ax = plt.subplots(figsize=(10, 6))
## Plotting chart
sns.set_color_codes("pastel")
sns.barplot(x="total",
y=result_df.index,
data=result_df,
label="# of attempts", color="b")
sns.set_color_codes("muted")
sns.barplot(x='is_goal',
y=result_df.index,
data=result_df,
label="# of goals", color="b")
ax.legend(ncol=2, loc="lower right", frameon=True)
ax.set(ylabel="Players",
xlabel="Number of goals x attempts", title='Player effectiveness')
each = result_df['is_goal'].values
the_total = result_df['total'].values
x_position = 50
for i in range(len(ax.patches[:15])):
ax.text(ax.patches[i].get_width() - x_position, ax.patches[i].get_y() +.50,
str(round((each[i]/the_total[i])*100, 2))+'%')
sns.despine(left=True, bottom=True)
f.tight_layout()
# -
grouped_by_player_is_goal_filtered['id_event']
players = sorted_results[:10].index
players
threshold
grouped_by_player_is_goal_filtered
grouped_by_player = df_events[df_events['event_type'] == 1].groupby('event_team').count()
rouped_by_player_goals = df_events[(df_events['event_type'] == 1) &
(df_events['is_goal'] == 1)].groupby('event_team').count()
grouped_by_player_not_goals = df_events[(df_events['event_type'] == 1) &
(df_events['is_goal'] == 0)].groupby('event_team').count()
threshold = grouped_by_player['id_event'].std()
grouped_by_player_is_goal = df_events[df_events['is_goal'] == 1].groupby('event_team').count()
grouped_by_player_is_goal_filtered = grouped_by_player_goals[grouped_by_player_goals['id_event'] > threshold]
threshold
grouped_by_player_is_goal
| Football Dataset Analysis/.ipynb_checkpoints/Data_analysis_football-checkpoint.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6 with Spark
# language: python3
# name: python36
# ---
# Welcome to exercise one of week two of “Apache Spark for Scalable Machine Learning on BigData”. In this exercise you’ll read a DataFrame in order to perform a simple statistical analysis. Then you’ll rebalance the dataset. No worries, we’ll explain everything to you, let’s get started.
#
# Let’s create a data frame from a remote file by downloading it:
#
#
#
# This notebook is designed to run in a IBM Watson Studio Apache Spark runtime. In case you are running it in an IBM Watson Studio standard runtime or outside Watson Studio, we install Apache Spark in local mode for test purposes only. Please don't use it in production.
!pip install --upgrade pip
if not ('sc' in locals() or 'sc' in globals()):
print('It seems you are note running in a IBM Watson Studio Apache Spark Notebook. You might be running in a IBM Watson Studio Default Runtime or outside IBM Waston Studio. Therefore installing local Apache Spark environment for you. Please do not use in Production')
from pip import main
main(['install', 'pyspark==2.4.5'])
from pyspark import SparkContext, SparkConf
from pyspark.sql import SparkSession
sc = SparkContext.getOrCreate(SparkConf().setMaster("local[*]"))
spark = SparkSession \
.builder \
.getOrCreate()
# +
# delete files from previous runs
!rm -f hmp.parquet*
# download the file containing the data in PARQUET format
!wget https://github.com/IBM/coursera/raw/master/hmp.parquet
# create a dataframe out of it
df = spark.read.parquet('hmp.parquet')
# register a corresponding query table
df.createOrReplaceTempView('df')
# -
# Let’s have a look at the data set first. This dataset contains sensor recordings from different movement activities as we will see in the next week’s lectures. X, Y and Z contain accelerometer sensor values whereas the class field contains information about which movement has been recorded. The source field is optional and can be used for data lineage since it contains the file name of the original file where the particular row was imported from.
#
# More details on the data set can be found here:
# https://github.com/wchill/HMP_Dataset
#
df.show()
df.printSchema()
# This is a classical classification data set. One thing we always do during data analysis is checking if the classes are balanced. In other words, if there are more or less the same number of example in each class. Let’s find out by a simple aggregation using SQL.
spark.sql('select class,count(*) from df group by class').show()
# As you can see there is quite an imbalance between classes. Before we dig into this, let’s re-write the same query using the DataFrame API – just in case you are not familiar with SQL. As we’ve learned before, it doesn’t matter if you express your queries with SQL or the DataFrame API – it all gets boiled down into the same execution plan optimized by Tungsten and accelerated by Catalyst. You can even mix and match SQL and DataFrame API code if you like.
#
# Again, more details on the API can be found here:
# https://spark.apache.org/docs/latest/api/python/pyspark.sql.html#pyspark.sql.DataFrame
#
df.groupBy('class').count().show()
# Let’s create a bar plot from this data. We’re using the pixidust library, which is Open Source, because of its simplicity. But any other library like matplotlib is fine as well.
# + pixiedust={"displayParams": {"handlerId": "barChart", "keyFields": "class", "legend": "true", "mpld3": "false", "orientation": "horizontal", "rendererId": "matplotlib", "sortby": "Values ASC", "valueFields": "count"}}
import pixiedust
from pyspark.sql.functions import col
counts = df.groupBy('class').count().orderBy('count')
display(counts)
# -
# This looks nice, but it would be nice if we can aggregate further to obtain some quantitative metrics on the imbalance like, min, max, mean and standard deviation. If we divide max by min we get a measure called minmax ration which tells us something about the relationship between the smallest and largest class. Again, let’s first use SQL for those of you familiar with SQL. Don’t be scared, we’re used nested sub-selects, basically selecting from a result of a SQL query like it was a table. All within on SQL statement.
spark.sql('''
select
*,
max/min as minmaxratio -- compute minmaxratio based on previously computed values
from (
select
min(ct) as min, -- compute minimum value of all classes
max(ct) as max, -- compute maximum value of all classes
mean(ct) as mean, -- compute mean between all classes
stddev(ct) as stddev -- compute standard deviation between all classes
from (
select
count(*) as ct -- count the number of rows per class and rename it to ct
from df -- access the temporary query table called df backed by DataFrame df
group by class -- aggrecate over class
)
)
''').show()
# The same query can be expressed using the DataFrame API. Again, don’t be scared. It’s just a sequential expression of transformation steps. You now an choose which syntax you like better.
# +
from pyspark.sql.functions import col, min, max, mean, stddev
df \
.groupBy('class') \
.count() \
.select([
min(col("count")).alias('min'),
max(col("count")).alias('max'),
mean(col("count")).alias('mean'),
stddev(col("count")).alias('stddev')
]) \
.select([
col('*'),
(col("max") / col("min")).alias('minmaxratio')
]) \
.show()
# -
# Now it’s time for you to work on the data set. First, please create a table of all classes with the respective counts, but this time, please order the table by the count number, ascending.
$$$ your code goes here
# Pixiedust is a very sophisticated library. It takes care of sorting as well. Please modify the bar chart so that it gets sorted by the number of elements per class, ascending. Hint: It’s an option available in the UI once rendered using the display() function.
$$$ your code goes here
# Imbalanced classes can cause pain in machine learning. Therefore let’s rebalance. In the flowing we limit the number of elements per class to the amount of the least represented class. This is called undersampling. Other ways of rebalancing can be found here:
#
# https://machinelearningmastery.com/tactics-to-combat-imbalanced-classes-in-your-machine-learning-dataset/
# +
from pyspark.sql.functions import min
# create a lot of distinct classes from the dataset
classes = [row[0] for row in df.select('class').distinct().collect()]
# compute the number of elements of the smallest class in order to limit the number of samples per calss
min = df.groupBy('class').count().select(min('count')).first()[0]
# define the result dataframe variable
df_balanced = None
# iterate over distinct classes
for cls in classes:
# only select examples for the specific class within this iteration
# shuffle the order of the elements (by setting fraction to 1.0 sample works like shuffle)
# return only the first n samples
df_temp = df \
.filter("class = '"+cls+"'") \
.sample(False, 1.0) \
.limit(min)
# on first iteration, assing df_temp to empty df_balanced
if df_balanced == None:
df_balanced = df_temp
# afterwards, append vertically
else:
df_balanced=df_balanced.union(df_temp)
# -
# Please verify, by using the code cell below, if df_balanced has the same number of elements per class. You should get 6683 elements per class.
$$$
| IBM_skillsnetwork/2_coursera_bd/week2/a6_w2_ex1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.4 64-bit (''venv_scratch2'': venv)'
# language: python
# name: python3
# ---
import sys
sys.path.append('..')
import numpy as np
from common.my_util import preprocess
text = 'You say goodbye and I say hello.'
corpus, word_to_id, id_to_word = preprocess(text)
# # 共起表現
# - 単語の周辺(ウィンドウサイズ分)に出てくる単語をカウント
# 共起行列
C = np.array([
[0, 1, 0, 0, 0, 0, 0],
[1, 0, 1, 0, 1, 1, 0],
[0, 1, 0, 1, 0, 0, 0],
[0, 0, 1, 0, 1, 0, 0],
[0, 1, 0, 1, 0, 0, 0],
[0, 1, 0, 0, 0, 0, 1],
[0, 0, 0, 0, 0, 1, 0],
], dtype=np.int32)
print(C[0]) # 単語IDが0のベクトル
print(C[word_to_id['goodbye']]) # goodbyeのベクトル
for idx, word_id in enumerate(corpus):
print(idx, word_id)
# +
import sys
sys.path.append('..')
from common.util import preprocess , create_co_matrix, cos_similarity
text = 'You say goodby and I say hello.'
corpus, word_to_id, id_to_word = preprocess(text) # 小文字に変換される
vocab_size = len(word_to_id)
C = create_co_matrix(corpus, vocab_size) # 共起行列の作成
# youとiのコサイン類似度
c0 = C[word_to_id['you']] # youの単語ベクトル
c1 = C[word_to_id['i']] # iの単語ベクトル
print(cos_similarity(c0, c1)) # 0.7と比較的高い値が出る
# -
arr = np.array([10,1000,1])
arr.argsort()
| ch02/ch02-3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### visualize the correlation coefficients between features (wheat dataset)
import pandas as pd
import matplotlib.pyplot as plt
# load the data
flc = '/Users/pinqingkan/Desktop/Codes/Course_edX_PythonDataScience/03_ExploratoryAnalysis/Datasets/'
fname = flc + 'wheat.data'
X = pd.read_csv(fname, index_col = 0)
Z = X.columns
Z
# calculate correlation
Y = X.corr()
Y
plt.imshow(Y, cmap = 'bwr', vmin = -.8, vmax = .8)
#ticks = [i for i in range(len(Y.columns))]
#plt.xticks(ticks, Y.columns)
plt.colorbar()
plt.show()
| Lab3_ExploratoryAnalysis/assignment6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Hull Moving Average
# https://www.incrediblecharts.com/indicators/hull-moving-average.php
#
# https://www.fidelity.com/learning-center/trading-investing/technical-analysis/technical-indicator-guide/hull-moving-average
# + outputHidden=false inputHidden=false
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import math
import warnings
warnings.filterwarnings("ignore")
# yfinance is used to fetch data
import yfinance as yf
yf.pdr_override()
# + outputHidden=false inputHidden=false
# input
symbol = 'AAPL'
start = '2018-01-01'
end = '2019-01-01'
# Read data
df = yf.download(symbol,start,end)
# View Columns
df.head()
# + outputHidden=false inputHidden=false
import talib as ta
# + outputHidden=false inputHidden=false
n = 30
df['WMA_1'] = ta.WMA(df['Adj Close'], timeperiod=n/2) * 2
df['WMA_2'] = df['WMA_1'] - ta.WMA(df['Adj Close'], timeperiod=n)
df['HMA'] = ta.WMA(df['WMA_2'], timeperiod=math.sqrt(n))
df = df.drop(['WMA_1', 'WMA_2'], axis=1)
# + outputHidden=false inputHidden=false
df.tail()
# + outputHidden=false inputHidden=false
fig = plt.figure(figsize=(14,10))
ax1 = plt.subplot(2, 1, 1)
ax1.plot(df['Adj Close'])
ax1.set_title('Stock '+ symbol +' Closing Price')
ax1.set_ylabel('Price')
ax2 = plt.subplot(2, 1, 2)
ax2.plot(df['HMA'], label='Hull Moving Average', color='red')
#ax2.axhline(y=0, color='blue', linestyle='--')
ax2.grid()
ax2.set_ylabel('Hull Moving Average')
ax2.set_xlabel('Date')
ax2.legend(loc='best')
# -
# ## Candlestick with Hull Moving Average
# + outputHidden=false inputHidden=false
from matplotlib import dates as mdates
import datetime as dt
dfc = df.copy()
dfc['VolumePositive'] = dfc['Open'] < dfc['Adj Close']
#dfc = dfc.dropna()
dfc = dfc.reset_index()
dfc['Date'] = pd.to_datetime(dfc['Date'])
dfc['Date'] = dfc['Date'].apply(mdates.date2num)
dfc.head()
# + outputHidden=false inputHidden=false
from mpl_finance import candlestick_ohlc
fig = plt.figure(figsize=(14,10))
ax1 = plt.subplot(2, 1, 1)
candlestick_ohlc(ax1,dfc.values, width=0.5, colorup='g', colordown='r', alpha=1.0)
ax1.xaxis_date()
ax1.xaxis.set_major_formatter(mdates.DateFormatter('%d-%m-%Y'))
ax1.grid(True, which='both')
ax1.minorticks_on()
ax1v = ax1.twinx()
colors = dfc.VolumePositive.map({True: 'g', False: 'r'})
ax1v.bar(dfc.Date, dfc['Volume'], color=colors, alpha=0.4)
ax1v.axes.yaxis.set_ticklabels([])
ax1v.set_ylim(0, 3*df.Volume.max())
ax1.set_title('Stock '+ symbol +' Closing Price')
ax1.set_ylabel('Price')
ax2 = plt.subplot(2, 1, 2)
ax2.plot(df['HMA'], label='Hull Moving Average', color='red')
#ax2.axhline(y=0, color='blue', linestyle='--')
ax2.grid()
ax2.set_ylabel('Hull Moving Average')
ax2.set_xlabel('Date')
ax2.legend(loc='best')
| Python_Stock/Technical_Indicators/Hull_Moving_Average.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PythonData
# language: python
# name: pythondata
# ---
# # Clean Google Mobility Data
#
#
# Dependencies and Setup
import json
import os
import pandas as pd
import urllib.request
import requests
# from config import db_pwd, db_user
from sqlalchemy import create_engine
# ## Store Google CSV into DataFrame
csv_file = "Resources/google_mob_US.csv"
google_data_df = pd.read_csv(csv_file)
google_data_df.head()
#
# ### Rename the dataframe with select columns
google_data_df = google_data_df.rename(columns = {"State":'states', "date":'dates', "retail_and_recreation":'retail_recreation',
"grocery_and_pharmacy":'grocery_pharmacy', "parks":'parks', "transit_stations":"transit", "workplaces":"workplaces", "residential":"residential"})
google_data_df.head()
google_df = google_data_df[["states", "dates", "retail_recreation",
"grocery_pharmacy", "parks", "transit", "workplaces", "residential"]]
google_df.head()
# +
# Reseting the index and saving the cleaned file to csv
# clean_google_mob_US_df = clean_google_mob_US_df.reset_index(drop = True)
# clean_google_mob_US_df.to_csv("../Data/clean_google_mob_US.csv")
# -
# ## Adding the 30 Day Moving Average
google_df["SMA_retail_recreation"] = google_df.iloc[:,2].rolling(window=30).mean()
google_df["SMA_grocery_pharmacy"] = google_df.iloc[:,3].rolling(window=30).mean()
google_df["SMA_parks"] = google_df.iloc[:,4].rolling(window=30).mean()
google_df["SMA_transit"] = google_df.iloc[:,5].rolling(window=30).mean()
google_df["SMA_workplaces"] = google_df.iloc[:,6].rolling(window=30).mean()
google_df["SMA_residential"] = google_df.iloc[:,7].rolling(window=30).mean()
google_df.head()
# Reseting the index and saving the cleaned file to csv
google_us_df = google_df.reset_index(drop = True)
google_us_df.to_csv("data/google_us.csv")
# ## Test Data - Virginia
# Selecting only the data for the US. This dropped the data to 456634 rows × 14 columns
google_mob_VA = google_df.loc[google_df["states"] == "Virginia"]
google_mob_VA = google_mob_VA.reset_index(drop = True)
google_mob_VA.head()
# Grouping by date, so we can get all the data for all states into one date
# skipnabool, default is True, and all NA/null values are excluded, when computing the result.
data_by_date_VA_df = pd.DataFrame(google_mob_VA.groupby("dates").mean())
data_by_date_VA_df.reset_index(inplace = True)
data_by_date_VA_df.head()
# Reseting the index and saving the cleaned file to csv
data_by_date_VA_df.to_csv("data/google_mob_VAA.csv")
# Grouping by date, so we can get all the data for all states into one date
# skipnabool, default is True, and all NA/null values are excluded, when computing the result.
data_by_date_US_df = pd.DataFrame(google_df.groupby("dates").mean())
data_by_date_US_df.reset_index(inplace = True)
data_by_date_US_df.head()
# Reseting the index and saving the cleaned file to csv
data_by_date_US_df.to_csv("data/google_mob_US.csv")
# ## US States 30 Day Moving Average DF
# Review previous df for the entire U.S - Noting number of rows
google_df.head()
google_df.columns
# Groupby state and date to return the moving average (30 days)
data_us_df = pd.DataFrame(google_df.groupby(['states','dates']).mean())
data_us_df.reset_index(inplace = True)
data_us_df
# Export the US data as a csv file
data_us_df.to_csv("data/data_us.csv")
# +
# start_date = "2020-02-15"
# end_date = "2020-03-01"
# mask = (google_df["dates"] > start_date) & (google_df["dates"] <= end_date)
# cut_date_df = google_df.loc[mask]
# cut_date_df
# -
# ## Store NYT COVID cases and deaths CSV into DataFrame
csv_file = "Resources/COVID-states.csv"
covid_data_df = pd.read_csv(csv_file)
covid_data_df.head()
covid_us = covid_data_df.rename(columns = {"state":'states', "fips": 'fips', "date":'dates', "cases":'cases',
"deaths":'deaths'})
covid_us.head()
# ### Connect to local database
rds_connection_string = f"{db_user}:{{db_pwd}}@localhost:5432/mobility_db"
engine = create_engine(f'postgresql://{rds_connection_string}')
# ### Check for tables
engine.table_names()
# ### Use pandas to load csv converted DataFrame into database
google_us.to_sql(name='google_data', con=engine, if_exists='append', index=False)
covid_us.to_sql(name='covid_data', con=engine, if_exists='append', index=False)
data_us_df.to_sql(name='us_data', con=engine, if_exists='append', index=False)
# ### Confirm data has been added by querying the tables
pd.read_sql_query('select * from google_data', con=engine).head(10)
pd.read_sql_query('select * from covid_data', con=engine).head(10)
| COMPLETE/Google Data/Clean Google Data - Project 3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# !pip install mysql-connector-python
# +
from sqlalchemy import create_engine
source = create_engine('mysql+mysqlconnector://test:test123@172.20.0.2/test')
# -
# !pip install pandas
# +
import pandas as pd
pd.read_sql('select now()', con=source)
| docker/Jupyter_MYSQL_Docker.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Read data from a CARTO SQL Query
#
# This example illustrates how to read data from a CARTO table using a SQL Query.
#
# > Use this when you need to see or modify the data locally. If you only need to visualize the data, just use the query directly in the Layer: `Layer('SELECT * FROM dataset_name')`
#
# _Note: You'll need [CARTO Account](https://carto.com/signup) credentials to reproduce this example._
# +
from cartoframes.auth import set_default_credentials
set_default_credentials('cartoframes')
# +
from cartoframes import read_carto
gdf = read_carto("SELECT * FROM starbucks_brooklyn WHERE revenue > 1200000")
gdf.head()
# +
from cartoframes.viz import Layer
Layer(gdf)
| docs/examples/data_management/read_carto_query.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="o3cL5pykvDUW"
# Medical Cost Data
# + id="uimBB_ZjHOAN"
import tensorflow as tf
import pandas as pd
import matplotlib.pyplot as plt
# + [markdown] id="n8yL5mGouzqu"
# Veri setini link uzeerinden direkt okuyabiliriz bu sekilde
# + colab={"base_uri": "https://localhost:8080/", "height": 417} id="Mo5cCMDruFLg" outputId="db4ed704-d337-44c5-9008-02e277df982b"
insurance = pd.read_csv("https://raw.githubusercontent.com/stedy/Machine-Learning-with-R-datasets/master/insurance.csv")
insurance
# + colab={"base_uri": "https://localhost:8080/"} id="lX_UIizJuyB2" outputId="2261ecb3-30ee-4740-8ee8-b54b2b803fb4"
insurance["smoker"] , insurance['age']
# + [markdown] id="7210Ky-2vhbt"
# Goruldugu gibi bazi veriler numerik iken bazilari degil. Tum verilerimizi numerik degerlere yapmaliyiz.(numeric encoding)
# + [markdown] id="6dzdGuG9v0c2"
# **One-Hot Encoding**
#
# [Nasil biseye benzeniyor](https://i.imgur.com/mtimFxh.png)
# kullanalim be gorelim
# + colab={"base_uri": "https://localhost:8080/", "height": 437} id="1nlzRtl9vfPf" outputId="bdfd3687-18d1-4df6-adfe-7b738fce5265"
insurance_one_hot = pd.get_dummies(insurance)
insurance_one_hot
# + [markdown] id="7AusrojryLXm"
# Features and Labels
# + id="NbdNJm6xw_oo"
X = insurance_one_hot.drop("charges",axis=1) #charges coloumsu haric hepsi
y = insurance_one_hot["charges"] #charges coloumsu
# + colab={"base_uri": "https://localhost:8080/", "height": 223} id="Ka4EAbsJyw6p" outputId="8aa51305-00e6-428c-96a1-4b9574c59ca5"
X.head()
# + colab={"base_uri": "https://localhost:8080/"} id="NRstd5tXy_S_" outputId="08c664ec-3a67-4f6c-a904-43274be24011"
y.head()
# + [markdown] id="RqMXxelpzNmi"
# Train ve Test kumelerini hazirlayalim
# + colab={"base_uri": "https://localhost:8080/"} id="G_aXLKG3zAcX" outputId="7243a69b-782e-441b-a8bc-7815a42c2699"
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size= 0.2, random_state = 42)
len(X), len(X_train), len(X_test)
# + colab={"base_uri": "https://localhost:8080/", "height": 437} id="knWPgFubzvIw" outputId="c5f2f8f3-ca88-4a94-b341-61302a40a7a0"
X_train
# + [markdown] id="TcqrBqqk0gno"
# indexlerden de gordugunuz gibi data siralamalari rastgele karistirilimis sekilde
# + [markdown] id="nPVpS7oRsyRV"
# ### Modeli olusturalim
# + id="i1y8yoOh0fwo"
#create a model
insurance_model = tf.keras.Sequential([
tf.keras.layers.Dense(10),
tf.keras.layers.Dense(1)
])
#compile the model
insurance_model.compile(loss= tf.keras.losses.mae,
optimizer= tf.keras.optimizers.SGD(),
metrics= ["mae"]
)
#fit the model
insurance_model.fit(X_train, y_train, epochs= 100)
# + colab={"base_uri": "https://localhost:8080/"} id="Uv76IY1ctopd" outputId="5dd68f6a-825c-4c63-b14f-fbc039510b71"
#sonuclari test kumesinde test edelim
insurance_model.evaluate(X_test, y_test)
# + [markdown] id="dt3EgToevMjj"
# Modelimizin performansi iyi cikmadi bu yuzden gelsitirmeyi deneyelim
# + [markdown] id="SribSgj8va_R"
# 2 farkli yol deneyelim
#
# 1. Ekstra gizli katman ekleyelim ve optimizeri Adam olarak degistirelim
# 2. Daha uzun surede egitmeye calisalim
# + colab={"base_uri": "https://localhost:8080/"} id="AxZoMIFAusEy" outputId="3ade01fe-d5cb-4f25-e9de-c2df57d8d5d7"
#create a model
insurance_model_2 = tf.keras.Sequential([
tf.keras.layers.Dense(100),
tf.keras.layers.Dense(10),
tf.keras.layers.Dense(1)
])
#compile the model
insurance_model_2.compile(loss= tf.keras.losses.mae,
optimizer= tf.keras.optimizers.Adam(),#optimizeri da degistirmis olduk
metrics= ["mae"]
)
#fit the model
insurance_model_2.fit(X_train, y_train, epochs= 100,verbose= 0)
# + colab={"base_uri": "https://localhost:8080/"} id="7JrWi9XwwAiz" outputId="5dd16d04-d39d-43b8-90ac-c14df9356722"
#sonuclari test kumesinde test edelim
insurance_model_2.evaluate(X_test, y_test)
# + [markdown] id="E-tcrHSvwxmZ"
# Goruldugu gibi modelimizin performansi gelisti
# + [markdown] id="R5ieW4bQxI_w"
# Daha uzun surede egitmeyi deneyelim epochs = 200
# + id="kLYn83cWwJPq"
#create a model
insurance_model_3 = tf.keras.Sequential([
tf.keras.layers.Dense(100),
tf.keras.layers.Dense(10),
tf.keras.layers.Dense(1)
])
#compile the model
insurance_model_3.compile(loss= tf.keras.losses.mae,
optimizer= tf.keras.optimizers.Adam(),#optimizeri da degistirmis olduk
metrics= ["mae"]
)
#fit the model
history = insurance_model_3.fit(X_train, y_train, epochs= 200,verbose= 0)
# + colab={"base_uri": "https://localhost:8080/"} id="8C--rocUxZx6" outputId="af785aa9-0a57-404a-a06e-aa19f5b34bdd"
#sonuclari test kumesinde test edelim
insurance_model_3.evaluate(X_test, y_test)
# + [markdown] id="K3oyrrxCx6Dg"
# Modelim epochs sayisini 200 yaptigimizda daha da iyi bir performansa ulasmis oldu.
# + [markdown] id="PzK1sUJVzB8X"
# Modelimizin epochs(donem) boyunca gelisimine bakalim
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="yFog5vvjxeX5" outputId="da78a562-1e7e-4f95-d139-f46f9443d418"
pd.DataFrame(history.history).plot()
plt.ylabel("loss")
plt.xlabel("epochs")
# + [markdown] id="IuWoQNsb2JfG"
# Peki modeli ne kadar sure egitmeliyiz en ideal epoch sayisini nedir ?
#
# Bu uzerinde calistiginiz probleme gore degisiklik gostermektedir. Bu yuzden Tensorflow un bu konudaki bir cozumu olarak EarlyStopping yani erken durdurmayi kullanabilirsiniz.
#
# EarlyStopping modelimizin gelsiimi belirli bir seviyeye geldiginde veya artik gelismeyi durdurdugunda egitmimizi epoch sayimiza ulasmadan erken durdurmamaizi saglar.
# + [markdown] id="7uYcKo1K3LtO"
# ##Preporccessing Data (normalization and standardization)
#
# Normalization
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="PkdYwT9ayd3i" outputId="134a37a9-c321-4309-da62-33e84859256a"
X["age"].plot(kind= "hist")
# + [markdown] id="JKq5reK14L-F"
# Yas verilerini inceledik peki bunlarin hepsini 0-1 arasinda degerlere karsilik olarak sikistirsaydik. Iste buna normalization denir.
# + colab={"base_uri": "https://localhost:8080/", "height": 417} id="tViH_YgK4FWJ" outputId="6b7e3172-a4eb-41f0-9e83-e1501461e3af"
import pandas as pd
import tensorflow as tf
import matplotlib.pyplot as plt
#tekrar datamizi alalim
insurance = pd.read_csv("https://raw.githubusercontent.com/stedy/Machine-Learning-with-R-datasets/master/insurance.csv")
insurance
# + id="nSjRvNxV5nWW"
from sklearn.compose import make_column_transformer
from sklearn.preprocessing import MinMaxScaler, OneHotEncoder
from sklearn.model_selection import train_test_split
#create a column transformer(sutunlar icin transformer olusturalim )
ct = make_column_transformer(
(MinMaxScaler(),["age", "bmi", "children"]), #tum degerler 0-1 arasina sikistirdik
(OneHotEncoder(handle_unknown="ignore"),["sex","smoker", "region"])
)
#X and y
X = insurance.drop("charges",axis=1)
y = insurance["charges"]
#train test split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size= 0.2, random_state = 42)
len(X), len(X_train), len(X_test)
#fit column transformer to our training data
ct.fit(X_train)
X_train_normal =ct.transform(X_train)
X_test_normal =ct.transform(X_test)
# + [markdown] id="jrS55YiV8s2M"
# Nasil gorundugune bir bakalim
# + colab={"base_uri": "https://localhost:8080/"} id="HXqtI6zm8ESe" outputId="f3cce76b-087a-417c-fa60-50a94a2a677b"
X_train.loc[1]
# + colab={"base_uri": "https://localhost:8080/"} id="Mig8UKco8-68" outputId="ac52a545-2060-4156-a05e-db1c2f214b71"
X_train_normal[1]
# + [markdown] id="4ps2okDc9d27"
# Tum bu islemden sonra tekrar modelimiz fit edelim
# + id="TlHFpest9YD8"
#create a model
insurance_model_4 = tf.keras.Sequential([
tf.keras.layers.Dense(100),
tf.keras.layers.Dense(10),
tf.keras.layers.Dense(1)
])
#compile the model
insurance_model_4.compile(loss= tf.keras.losses.mae,
optimizer= tf.keras.optimizers.Adam(),#optimizeri da degistirmis olduk
metrics= ["mae"]
)
#fit the model
history = insurance_model_4.fit(X_train_normal, y_train, epochs= 100,verbose= 0)
# + colab={"base_uri": "https://localhost:8080/"} id="VAIMdq8N-Bl2" outputId="91e5f935-9bca-489b-b95a-a104d7c85603"
#sonuclari test kumesinde test edelim
insurance_model_4.evaluate(X_test_normal, y_test)
# + [markdown] id="L0peXyMC-bgz"
# Datamızı normalıze etmeden onceki ayni modelimizdeki sonucla arasindaki farki gorebiliyoruz
# + id="_PwoHzOM-SFN"
#9/9 [==============================] - 0s 2ms/step - loss: 5104.4990 - mae: 5104.4990
[5104.4990234375, 5104.4990234375]
| Regression2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
def read_csfile(csfile):
content = np.array(np.load(csfile).tolist())
dtype = np.load(csfile).dtype
header = []
for key in dtype.fields.keys():
header.append(key)
header = np.array(header)
return header, content
particle_file = 'P15_J35_passthrough_particles.cs'
header, content = read_csfile(particle_file)
header
content.shape
content[:10,1]
# %matplotlib inline
from matplotlib import pyplot as plt
mic1 = np.where(content[:,1]==content[0,1])
plt.scatter(content[mic1,5],content[mic1,6])
plt.show()
| notebooks/Analyse-picks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Giải phẫu cấu trúc Matplotlib
#
# ### BS. Lê Ngọc Khả Nhi
#
# # Giới thiệu
# Matplotlib là thư viện đồ họa thống kê chính trong Python, được thiết kế bởi <NAME> cách đây 18 năm (2002) như một giao thức mô phỏng theo ngôn ngữ Matlab. Sau khi Hunter qua đời năm 2012, thư viện tiếp tục được phát triển bởi <NAME> và <NAME>.
#
# Matplotlib là nền tảng về đồ họa trong ngôn ngữ Python, vì hầu hết những API đồ họa thống kê khác trong Python như seaborn, pandas, plotnine, bokeh, holoview, ... đều dựa trên matplotlib.
#
# Tuy không phô bày một cú pháp trong sáng như ggplot2 của R, nhưng matplotlib dễ học hơn nhiều so với base R graphics, đầy đủ tính năng cho nghiên cứu khoa học và một số loại biểu đồ vẽ bằng matplotlib có phẩm chất mỹ thuật không hề thua kém R.
#
# Matplotlib thực ra rất dễ học, khó khăn chủ yếu khi thực hành là do người dùng bị nhầm lẫn giữa 2 phong cách viết code: (1) sử dụng trực tiếp function và method và (2) tiếp cận theo hướng OOP - gồm subplots, Gridspec và Axes.
#
# Trong bài thực hành này, Nhi sẽ hệ thống lại những yếu tố quan trọng nhất trong cú pháp matplotlib, cho phép thỏa mãn tất cả nhu cầu trong công việc hằng ngày của nghiên cứu sinh.
#
# +
## Ignore warnings
import warnings
warnings.filterwarnings('ignore')
# Data preparation
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from itertools import product, combinations
import matplotlib.gridspec as plt_gs
# -
# Trong bài sử dụng dữ liệu minh họa là iris
# +
path = 'https://vincentarelbundock.github.io/Rdatasets/csv/datasets/iris.csv'
df = pd.read_csv(path, index_col=0)
df
# -
# # Vòng đời của một figure
#
# Khi bạn vẽ một tấm hình bằng matplotlib, nó được gọi là figure.
#
# Figure được sinh ra bằng method plt.figure(), trong đó bạn có thể tùy chỉnh nhiều thuộc tính, quan trọng nhất là kích thước figsize như 1 tuple, và độ phân giải dpi.
#
# Vòng đời của figure chỉ chấm dứt khi bạn trình diễn nó bằng plt.show(), hoặc sao lưu bằng plt.savefig(), kèm hoặc không kèm tắt tính năng display bằng method plt.close().
#
# Một khi figure chưa display hoặc sao lưu, ta có thể tiếp tục vẽ rất nhiều thành phần bên trong nó, sử dụng các hàm đồ họa của matplotlib, hoặc seaborn (method của seaborn tương thích hoàn toàn với pyplot).
#
# Bên trong 1 tấm hình (figure) có thể chứa rất nhiều graphs hay biểu đồ vấn đề là làm sao tổ chức trình bày những graph này theo mục tiêu của người dùng.
# +
plt.figure() # Figure sinh ra
... # tha hồ vẽ
plt.show() # đóng và trình diễn
# +
fig_1 = plt.figure() # Tạo object figure mới
...
plt.savefig() # sao lưu figure hiện hành
plt.close(fig_1) # Đóng figure
# -
# # Sử dụng trực tiếp hàm đồ họa từ pyplot
#
# Sau khi import module pyplot từ matplotlib với tên viết tắt là plt, người dùng có 2 cách làm việc, hoặc dùng functions/methods trực tiếp từ plt, hoặc khởi tạo những objects (phong cách code OOP).
#
# Cách làm thứ nhất tiện lợi khi bạn chỉ cần tạo 1 graph duy nhất, cấu tạo đơn giản bên trong figure, hoặc chồng lắp những layer cùng bản chất trên một graph duy nhất.
#
# Thí dụ, nếu bạn chỉ cần vẽ 1 scatter_plot đơn, có thể gọi trực tiếp plt.scatter()
# +
plt.figure(figsize = (6,5))
plt.scatter(df['Sepal.Length'],df['Petal.Length'])
plt.xlabel('Sepal.Length')
plt.ylabel('Petal.Length')
plt.show()
# -
# Mỗi lần dùng 1 hàm đồ họa trực tiếp từ plt, bất kể hàm nào, nó chồng 1 layer lên figure hiện hành, nhưng vẫn nằm trên cùng 1 graph, ta lợi dụng hành vi này để vẽ những biểu đồ có yếu tố phân nhóm nhưng cùng bản chất, thí dụ time series, scatter plot, density plot... bằng vòng lặp for
# +
plt.figure(figsize = (6,5))
for i in df['Species'].unique():
filt_df = df[df['Species'] == i]
plt.scatter(filt_df['Sepal.Length'],filt_df['Petal.Length'], label = i)
plt.xlabel('Sepal.Length')
plt.ylabel('Petal.Length')
plt.legend()
plt.show()
# -
# vì seaborn cũng chính là matplotlib, nó cũng tuân theo quy tắc này:
# +
plt.figure(figsize = (6,5))
for i in df['Species'].unique():
filt_df = df[df['Species'] == i]
sns.kdeplot(filt_df['Sepal.Length'], label = i)
plt.xlabel('Sepal.Length')
plt.ylabel('Petal.Length')
plt.show()
# -
# Tuy nhiên, một khi bạn quên chưa đóng figure và muốn vẽ 2 graph khác nhau, kết quả có thể rất kì dị, không theo ý muốn:
# +
plt.figure(figsize = (6,5))
plt.scatter(df['Sepal.Length'],df['Petal.Length'])
plt.plot(df['Sepal.Length'], 'r')
plt.xlabel('Sepal.Length')
plt.ylabel('Petal.Length')
plt.show()
# +
plt.figure(figsize = (6,5))
plt.scatter(df['Sepal.Length'],df['Petal.Length'])
for i in df['Species'].unique():
filt_df = df[df['Species'] == i]
sns.kdeplot(filt_df['Sepal.Length'], label = i)
plt.xlabel('Sepal.Length')
plt.ylabel('Petal.Length')
plt.show()
# -
# Nếu ta muốn vẽ hàng loạt figure cùng bản chất, có thể đóng figure sau mỗi iteration trong một vòng lặp, bằng cách này bạn vẫn đang sử dụng function và method, chưa cần dùng đến tính năng OOP, và hiệu quả vẫn có thể chấp nhận được:
# +
plt.figure()
for i in combinations(df.drop(['Species'], axis = 1).columns, 2):
for j in df['Species'].unique():
filt_df = df[df['Species'] == j]
plt.scatter(filt_df[i[0]], filt_df[i[1]], label = j)
plt.xlabel(i[0])
plt.ylabel(i[1])
plt.legend()
plt.show()
# -
# ## Sử dụng subplot
#
# Lưu ý, trường hợp trên ta có 6 biểu đồ scatter plot nối tiếp nhau, tuy nhiên mỗi biểu đồ chính là 1 figure riêng biệt, và mỗi figure này chỉ chứa 1 graph bên trong, sau khi figure trước được đóng, thì figure mới tiếp theo sinh ra, cho đến figure cuối cùng.
#
# Đó là cách làm thủ công, nhược điểm của nó, đó là bạn sẽ có 6 figure rời rạc, bạn không thể lưu lại hình ảnh trong notebook thành 1 figure duy nhất, mà chỉ có thể lưu 6 files ảnh rời rạc.
#
# Cách làm đúng, đó là trình bày 6 graphs trong 1 figure; sử dụng method plt.subplot()
#
# Thí dụ sau, Nhi tổ chức 1 figure có 4 hàng, 1 cột, trong cùng 1 figure, bằng plt.subplot(411), sau đó + vị trí i của biến cần vẽ kde_plot, như vậy 412 là graph ở vị trí thứ 2 trong figure 4 hàng x 1 cột.
# +
plt.figure(figsize = (6,8))
for i,k in zip(range(4), df.drop(['Species'], axis = 1).columns):
plt.subplot(411 + i)
for j in df['Species'].unique():
filt_df = df[df['Species'] == j]
sns.kdeplot(filt_df[k], shade = True, alpha = 0.3, label = j)
plt.xlabel(k)
plt.tight_layout()
plt.show()
# -
# Tương tự, 141, 142, 143... là cấu trúc figure có 1 hàng x 4 cột:
# +
plt.figure(figsize = (8,8))
for i,k in zip(range(4), df.drop(['Species'], axis = 1).columns):
plt.subplot(141 + i)
for j in df['Species'].unique():
filt_df = df[df['Species'] == j]
sns.kdeplot(filt_df[k], shade = True, alpha = 0.3, label = j, vertical = True)
plt.xlabel(k)
plt.tight_layout()
plt.show()
# -
# # Phong cách OOP
#
# Bây giờ, ta sẽ dùng phong cách OOP để tạo ra object figrue và axes là các phân vùng cho graphs
#
# ## Sử dụng Subplots
#
# Cách làm thứ nhất để tạo ra figure có cấu trúc phức tạp, đó là dùng plt.subplots, với arguments về số hàng, số cột.
#
# Thí dụ: fig, axes = plt.subplots(3,4, figsize = (6,6)) tạo ra 1 figure có 3 hàng, 4 cột và kích thước figure là 6x6 inches
fig, axes = plt.subplots(3,4, figsize = (6,6))
# trong đó, axes có bản chất là numpy array, trong thí dụ này là array có shape = 3,4
axes
axes.shape
axes[1]
# Bạn có thể hình dung, axes[0] là hàng thứ nhất, axes[2] là hàng thứ 3...
axes[0][1] # hàng thứ nhất, cột thứ 2
# Trong thí dụ sau, ta xếp 6 scatter plot vào một ma trận 3 hàng, 2 cột, sử dụng thủ thuật axes.flatten() để biến numpy array 2 chiều thành 1 chiều.
#
# Cách làm này chỉ dùng khi cả 6 biểu đồ có cùng bản chất (vẽ ra bởi cùng 1 hàm).
#
# Thí dụ bạn muốn vẽ kde plot cho hàng loạt biến, đây là cách phù hợp nhất
# +
fig, axes = plt.subplots(3,2, figsize = (6,8))
for i,ax in zip(combinations(df.drop(['Species'], axis = 1).columns, 2),
axes.flatten()):
for j in df['Species'].unique():
filt_df = df[df['Species'] == j]
ax.scatter(filt_df[i[0]], filt_df[i[1]], label = j)
ax.set_xlabel(i[0])
ax.set_ylabel(i[1])
ax.legend()
plt.tight_layout()
plt.show()
# -
# Trong trường hợp bạn muốn vẽ nhiều loại biểu đồ bằng nhiều hàm khác nhau, và xếp mỗi loại vào cùng 1 cột, cũng có thể dùng axes và vòng lặp:
# +
fig, axes = plt.subplots(4,2, figsize = (6,8), sharex=True, sharey = True)
for i,ax,target in zip(range(4), axes, df.drop(['Species'], axis = 1).columns):
sns.boxplot(data = df, x = 'Species', y = target, ax = ax[0])
sns.violinplot(data = df, x = 'Species', y = target, ax = ax[1])
plt.tight_layout()
plt.show()
# +
fig, axes = plt.subplots(4,3, figsize = (8,8), sharex=False, sharey = False)
for i,ax,target in zip(range(4), axes, df.drop(['Species'], axis = 1).columns):
sns.boxplot(data = df, x = 'Species', y = target, ax = ax[0])
sns.violinplot(data = df, x = 'Species', y = target, ax = ax[1])
for j in df['Species'].unique():
filt_df = df[df['Species'] == j]
sns.kdeplot(filt_df[target], ax = ax[2], shade = True, alpha = 0.3)
ax[2].set_xlabel(target)
plt.tight_layout()
plt.show()
# -
# ## Gridspec
#
# Subplots chí thích hợp cho những quy trình đơn giản (các graph theo 1 pattern nào đó, tỉ lệ diện tích như nhau...), nhưng không dùng được khi bạn cần vẽ 1 figure phức tạp, thí dụ ghép 2 density plot vào trục hoành/tung của 1 scatter plot.
#
# Lúc này, ta dùng Gridspec, phương pháp này chia figure ra thành một lưới nhiều ô có tọa độ hàng và cột như 1 numpy array.
#
# Tiếp theo, ta dùng subplot để tạo ra các axes có tọa độ và diện tích khác nhau, tùy ý, miễn là vẫn nằm trong phạm vi lưới tọa độ của gridspec.
#
# Kết quả như sau:
# +
fig = plt.figure(figsize = (8,8))
gs = plt_gs.GridSpec(3,3)
ax1 = plt.subplot(gs[0,:-1])
ax2 = plt.subplot(gs[1:,:-1])
ax3 = plt.subplot(gs[1:,-1])
for j in df['Species'].unique():
filt_df = df[df['Species'] == j]
sns.kdeplot(filt_df['Sepal.Length'], ax = ax1, shade = True, alpha = 0.3)
sns.kdeplot(filt_df['Petal.Length'], ax = ax3, vertical = True, shade = True, alpha = 0.3)
ax2.scatter(filt_df['Sepal.Length'], filt_df['Petal.Length'], label = j)
ax2.set_xlabel('Sepal.Length')
ax2.set_ylabel('Petal.Length')
plt.tight_layout()
plt.show()
# -
# Gridspec có thể tạo figure có cấu trúc tương đối phức tạp, tuy nhiên vẫn còn nhược điểm là không cho phép tinh chỉnh kích thước, tọa độ những axes bên trong theo ý thích, mà chỉ có thể đặt cứng nhắc vào 1 vùng nào đó, giống như xếp đồ vào từng ngăn trong chiếc hộp.
#
# Do đó gridspec thích hợp cho những hình vẽ có cấu trúc nhiều graphs riêng biệt, không chồng lắp lên nhau, canh hàng hoặc cột thẳng, đều.
#
#
# ## Sử dụng Axes
#
# Một tính năng cao cấp hơn, đó là Axes, nó cho phép tự do tạo ra axes với kích thước tùy thích, đặt ở bất cứ vị trí nào bạn muốn, miễn là nó vẫn nằm trong giới hạn của figure.
#
# Mỗi axis được tạo ra như 1 hình chữ nhật với 4 thuộc tính (left, bottom, width, height), left là khoảng cách tính từ lề bên trái của figure, bottom là khoảng cách tính từ đáy figure, width và height là chiều rộng và chiều cao; tất cả tính bằng tỉ lệ từ 0-1 so với kích thước của figure.
#
# Ta có thể dùng axes để vẽ ra những figure ô cùng phức tạp, thí dụ như sau:
# +
plt.figure(1, figsize=(8, 8))
left, bottom = 0.01, 0.1
w_kde = h_kde = 0.2
left_h = left + w_kde + 0.02
width = height = 0.65
bottom_h = bottom + height + 0.02
rect_y = [left, bottom, w_kde-0.04, height]
rect_main = [left_h, bottom, width, height]
rect_x = [left_h, bottom_h + 0.01, width, h_kde]
ax_main = plt.axes(rect_main)
ax_x = plt.axes(rect_x)
ax_y = plt.axes(rect_y)
ax_reg = plt.axes([left_h + 0.35, bottom + 0.08, 0.25, 0.25])
for j in df['Species'].unique():
filt_df = df[df['Species'] == j]
sns.kdeplot(filt_df['Sepal.Length'], ax = ax_x, shade = True, alpha = 0.3)
sns.kdeplot(filt_df['Petal.Length'], ax = ax_y, vertical = True, shade = True, alpha = 0.3)
ax_main.scatter(filt_df['Sepal.Length'], filt_df['Petal.Length'], label = j)
sns.regplot(data = df,
x = 'Sepal.Length',
y = 'Petal.Length',
ax = ax_reg,
scatter_kws={'alpha':0.2,'color':'red','s': 5},
line_kws={'color': 'black', 'lw':1})
ax_main.set_xlabel('Sepal.Length')
ax_main.set_ylabel('Petal.Length')
ax_y.invert_xaxis()
ax_y.legend().remove()
ax_x.legend().remove()
ax_main.legend()
plt.tight_layout()
# -
# # Tổng kết
#
# 1) Figure là hình vẽ tổng thể, bên trong figure chứa 1 hay nhiều graphs
#
# 2) Method seaborn cũng chính là matplotlib, khi không xác định ax, hàm seaborn tương đương sử dụng hàm ptt trực tiếp, và chồng lên graph hiện hành, khi xác định ax, seaborn method cho phép gán graph mới vào axis tùy chọn.
#
# 3) Không bắt buộc dùng OOP, chỉ cần dùng hàm pyplot trực tiếp, hoặc subplot rời rạc là đủ cho hơn 90% nhu cầu công việc.
#
# 4) Có 3 cấp độ tùy chỉnh cấu trúc figure bằng OOP: đơn giản nhất là subplots, cao hơn là gridspec, và tinh tế nhất là Axes.
#
# 5) Sử dụng subplots khi cần vẽ hàng loạt hình cùng bản chất, có kích thước như nhau
#
# 6) Sử dụng gridspec để vẽ những graphs có kích thước khác nhau nhưng tách biệt, canh thẳng hàng/cột, không chồng lắp và số lượng giới hạn.
#
# 7) Sử dụng Axes để vẽ những graphs tự do về tọa độ và kích thước, có thể chồng lên nhau.
| external_research_tutorials/bsNhi_lectures/Giai phau matplotlib.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="-3fR5n1TDCpw" colab_type="code" colab={}
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# + id="-XlwWfHmDiD3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="6dc3489e-c897-4c56-c09f-7fcd0bac8053" executionInfo={"status": "ok", "timestamp": 1583267120373, "user_tz": -60, "elapsed": 495, "user": {"displayName": "<NAME>\u015b", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Giri_N5UJuhCf-cKwdy3CXW6oVHsYTHqXRi5XeJiQ=s64", "userId": "16417033357933413602"}}
# cd '/content/drive/My Drive/Colab Notebooks/matrix/matrix_two/dw_matrix_car'
# + id="yZxAVplkDuNg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="0417630f-9be9-4516-962e-5f4b6b95d8cb" executionInfo={"status": "ok", "timestamp": 1583266960672, "user_tz": -60, "elapsed": 2147, "user": {"displayName": "<NAME>\u015b", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Giri_N5UJuhCf-cKwdy3CXW6oVHsYTHqXRi5XeJiQ=s64", "userId": "16417033357933413602"}}
# ls
# + id="C93bAYugEIqu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="4f853d91-8b43-4f60-a89d-4c4133736b3f" executionInfo={"status": "ok", "timestamp": 1583267123828, "user_tz": -60, "elapsed": 2442, "user": {"displayName": "<NAME>\u015b", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Giri_N5UJuhCf-cKwdy3CXW6oVHsYTHqXRi5XeJiQ=s64", "userId": "16417033357933413602"}}
df = pd.read_hdf('data/car.h5')
df.shape
# + id="EZ5OwqXGETe2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="f8999ca8-699a-459a-a4f0-763219cd9a7f" executionInfo={"status": "ok", "timestamp": 1583267152556, "user_tz": -60, "elapsed": 510, "user": {"displayName": "<NAME>\u015b", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Giri_N5UJuhCf-cKwdy3CXW6oVHsYTHqXRi5XeJiQ=s64", "userId": "16417033357933413602"}}
df.columns.values
# + id="ipr2_VRFEo5P" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 170} outputId="af7b0a73-4167-44da-8823-fee0d1fccd9d" executionInfo={"status": "ok", "timestamp": 1583267255575, "user_tz": -60, "elapsed": 525, "user": {"displayName": "<NAME>\u015b", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Giri_N5UJuhCf-cKwdy3CXW6oVHsYTHqXRi5XeJiQ=s64", "userId": "16417033357933413602"}}
df['price_value'].describe()
# + id="3Kr5zoRpFRKy" colab_type="code" colab={}
def group_and_barplot(feat_groupby, feat_agg='price_value', agg_funcs=[np.mean, np.median, np.size], feat_sort='mean', top=50, subplots=True):
return (
df
.groupby(feat_groupby)[feat_agg]
.agg(agg_funcs)
.sort_values(by=feat_sort, ascending=False)
.head(top)
).plot(kind='bar', figsize=(15,5), subplots=subplots)
# + id="ElUiktM6FiPy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 437} outputId="6f9693d0-51ef-47e7-d4c0-b5783b917cff" executionInfo={"status": "ok", "timestamp": 1583268608255, "user_tz": -60, "elapsed": 2088, "user": {"displayName": "<NAME>\u015b", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Giri_N5UJuhCf-cKwdy3CXW6oVHsYTHqXRi5XeJiQ=s64", "userId": "16417033357933413602"}}
group_and_barplot('param_marka-pojazdu');
# + id="1xrlhUpHKN7G" colab_type="code" colab={}
| day2_visualisation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Unit Testing ML Code: Hands-on Exercise (Input Values)
#
# ## In this notebook we will explore unit tests to validate input data using a basic schema
#
# #### We will use a classic toy dataset: the Iris plants dataset, which comes included with scikit-learn
# Dataset details: https://scikit-learn.org/stable/datasets/index.html#iris-plants-dataset
#
# As we progress through the course, the complexity of examples will increase, but we will start with something basic. This notebook is designed so that it can be run in isolation, once the setup steps described below are complete.
#
# ### Setup
#
# Let's begin by importing the dataset and the libraries we are going to use. Make sure you have run `pip install -r requirements.txt` on requirements file located in the same directory as this notebook. We recommend doing this in a separate virtual environment (see dedicated setup lecture).
#
# If you need a refresher on jupyter, pandas or numpy, there are some links to resources in the section notes.
# + pycharm={"is_executing": false, "name": "#%%\n"}
from sklearn import datasets
import pandas as pd
import numpy as np
# Access the iris dataset from sklearn
iris = datasets.load_iris()
# Load the iris data into a pandas dataframe. The `data` and `feature_names`
# attributes of the dataset are added by default by sklearn. We use them to
# specify the columns of our dataframes.
iris_frame = pd.DataFrame(iris.data, columns=iris.feature_names)
# Create a "target" column in our dataframe, and set the values to the correct
# classifications from the dataset.
iris_frame['target'] = iris.target
# + pycharm={"is_executing": false, "name": "#%%\n"}
# View the first 5 rows of our dataframe.
iris_frame.head(5)
# -
# View summary statistics for our dataframe.
iris_frame.describe()
# ### Now that we have our data loaded, we will create a simplified pipeline.
#
# This pipeline is a class for encapsulating all the related functionality for our model. As the course unfolds, we will work with more complex pipelines, including those provided by third party libraries.
#
# We train a logistic regression model to classify the flowers from the Iris dataset.
# +
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
class SimplePipeline:
def __init__(self):
self.frame = None
# Shorthand to specify that each value should start out as
# None when the class is instantiated.
self.X_train, self.X_test, self.y_train, self.Y_test = None, None, None, None
self.model = None
self.load_dataset()
def load_dataset(self):
"""Load the dataset and perform train test split."""
# fetch from sklearn
dataset = datasets.load_iris()
# remove units ' (cm)' from variable names
self.feature_names = [fn[:-5] for fn in dataset.feature_names]
self.frame = pd.DataFrame(dataset.data, columns=self.feature_names)
self.frame['target'] = dataset.target
# we divide the data set using the train_test_split function from sklearn,
# which takes as parameters, the dataframe with the predictor variables,
# then the target, then the percentage of data to assign to the test set,
# and finally the random_state to ensure reproducibility.
self.X_train, self.X_test, self.y_train, self.y_test = train_test_split(
self.frame[self.feature_names], self.frame.target, test_size=0.65, random_state=42)
def train(self, algorithm=LogisticRegression):
# we set up a LogisticRegression classifier with default parameters
self.model = algorithm(solver='lbfgs', multi_class='auto')
self.model.fit(self.X_train, self.y_train)
def predict(self, input_data):
return self.model.predict(input_data)
def get_accuracy(self):
# use our X_test and y_test values generated when we used
# `train_test_split` to test accuracy.
# score is a method on the Logisitic Regression that
# returns the accuracy by default, but can be changed to other metrics, see:
# https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LogisticRegression.html#sklearn.linear_model.LogisticRegression.score
return self.model.score(X=self.X_test, y=self.y_test)
def run_pipeline(self):
"""Helper method to run multiple pipeline methods with one call."""
self.load_dataset()
self.train()
# +
pipeline = SimplePipeline()
pipeline.run_pipeline()
accuracy_score = pipeline.get_accuracy()
# note that f' string interpolation syntax requires python 3.6
# https://www.python.org/dev/peps/pep-0498/
print(f'current model accuracy is: {accuracy_score}')
# -
# ### Test Inputs
#
# Now that we have our basic pipeline, we are in a position to test the input data.
#
# Best practice is to use a schema. A schema is a collection of rules which specify the expected values for a set of fields. Below we show a simple schema (just using a nested dictionary) for the Iris dataset. Later in the course we will look at more complex schemas, using some of the common Python libraries for data validation.
#
# The schema specifies the maximum and minimum values that can be taken by each variable. We can learn these values from the data, as we have done for this demo, or these values may come from specific domain knowledge of the subject.
iris_schema = {
'sepal length': {
'range': {
'min': 4.0, # determined by looking at the dataframe .describe() method
'max': 8.0
},
'dtype': float,
},
'sepal width': {
'range': {
'min': 1.0,
'max': 5.0
},
'dtype': float,
},
'petal length': {
'range': {
'min': 1.0,
'max': 7.0
},
'dtype': float,
},
'petal width': {
'range': {
'min': 0.1,
'max': 3.0
},
'dtype': float,
}
}
# +
import unittest
import sys
class TestIrisInputData(unittest.TestCase):
def setUp(self):
# `setUp` will be run before each test, ensuring that you
# have a new pipeline to access in your tests. See the
# unittest docs if you are unfamiliar with unittest.
# https://docs.python.org/3/library/unittest.html#unittest.TestCase.setUp
self.pipeline = SimplePipeline()
self.pipeline.run_pipeline()
def test_input_data_ranges(self):
# get df max and min values for each column
max_values = self.pipeline.frame.max()
min_values = self.pipeline.frame.min()
# loop over each feature (i.e. all 4 column names)
for feature in self.pipeline.feature_names:
# use unittest assertions to ensure the max/min values found in the dataset
# are less than/greater than those expected by the schema max/min.
self.assertTrue(max_values[feature] <= iris_schema[feature]['range']['max'])
self.assertTrue(min_values[feature] >= iris_schema[feature]['range']['min'])
def test_input_data_types(self):
data_types = self.pipeline.frame.dtypes # pandas dtypes method
for feature in self.pipeline.feature_names:
self.assertEqual(data_types[feature], iris_schema[feature]['dtype'])
# -
# setup code to allow unittest to run the above tests inside the jupyter notebook.
suite = unittest.TestLoader().loadTestsFromTestCase(TestIrisInputData)
unittest.TextTestRunner(verbosity=1, stream=sys.stderr).run(suite)
# ### Data Input Test: Hands-on Exercise
# Change either the schema or the input data (not the model config) so that the test fails. Do you understand why the test is failing?
| exercise_notebooks/unit_testing_exercise/unit_testing_input_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
from sklearn import datasets
iris = datasets.load_iris()
X = iris.data # Input variables
y0 = iris.target # Output labels as integers
from keras.utils import to_categorical
y = to_categorical(y0) # One-hot encoding of output labels
import numpy as np
np.random.seed(0)
from sklearn.utils import shuffle
X, y0, y = shuffle(X, y0, y)
from keras import models
from keras import layers
model = models.Sequential()
model.add(layers.Dense(10, activation="relu", input_dim=X.shape[1]))
model.add(layers.Dense(3, activation="softmax"))
model.compile(optimizer="adam",
loss="categorical_crossentropy",
metrics=["accuracy"])
model.fit(X, y, epochs=500, validation_split=0.2)
# Let's predict probabilities for the first two examples.
model.predict(X[0:2,:])
# Now let's fit the same model but this time encoding the data as integers
model2 = models.Sequential()
model2.add(layers.Dense(10, activation="relu", input_dim=X.shape[1]))
model2.add(layers.Dense(3, activation="softmax"))
model2.compile(optimizer="adam",
loss="sparse_categorical_crossentropy",
metrics=["accuracy"])
model2.fit(X, y0, epochs=500, validation_split=0.2)
| notes/4.2 Iris.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Package loading and basic configurations
# +
# %reload_ext autoreload
# %autoreload 2
# load dependencies'
import pandas as pd
import geopandas as gpd
from envirocar import TrackAPI, DownloadClient, BboxSelector, ECConfig
# create an initial but optional config and an api client
config = ECConfig()
track_api = TrackAPI(api_client=DownloadClient(config=config))
# -
# # Querying enviroCar Tracks
# The following cell queries tracks from the enviroCar API. It defines a bbox for the area of Münster (Germany) and requests 50 tracks. The result is a GeoDataFrame, which is a geo-extended Pandas dataframe from the GeoPandas library. It contains all information of the track in a flat dataframe format including a specific geometry column.
# +
bbox = BboxSelector([
6.8, # min_x
51.0, # min_y
8.9, # max_x
52.2 # max_y
])
# issue a query
track_df = track_api.get_tracks(bbox=bbox, num_results=50) # requesting 50 tracks inside the bbox
track_df
# -
# # Visualization using MatPlotlib package:
# Using the matplotlib package I have created a histogram for the Intake Pressure value. And the result shows that for approximately more the 2000 points the pressure ranges from approximately 20 to 35 kPa and so on.
import matplotlib.pyplot as plt
track_df['Intake Pressure.value'].hist()
plt.show()
# # Statistics:
# Using pandas summarized the count, mean, standard deviation and IQR values of all the columns using describe().
print(df.describe())
track_df.plot(figsize=(8, 10))
# # Inspecting a single Track
some_track_id = track_df['track.id'].unique()[9]
some_track = track_df[track_df['track.id'] == some_track_id]
some_track.plot()
ax = some_track['Consumption.value'].plot()
ax.set_title("Consumption")
ax.set_ylabel(some_track['Consumption.unit'][0])
ax
# ## Interactive Map
# The following map-based visualization makes use of folium. It allows to visualizate geospatial data based on an interactive leaflet map. Since the data in the GeoDataframe is modelled as a set of Point instead of a LineString, we have to manually create a polyline
# +
import folium
lats = list(some_track['geometry'].apply(lambda coord: coord.y))
lngs = list(some_track['geometry'].apply(lambda coord: coord.x))
avg_lat = sum(lats) / len(lats)
avg_lngs = sum(lngs) / len(lngs)
m = folium.Map(location=[avg_lat, avg_lngs], zoom_start=13)
folium.PolyLine([coords for coords in zip(lats, lngs)], color='blue').add_to(m)
m
| examples/G1-04-11--Sivakumar-enviroCar_PY_Notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import networkx as nx
from networkx.algorithms.operators.binary import difference, symmetric_difference
import os
import json
import matplotlib.pyplot as plt
from zipfile import ZipFile
from tqdm.notebook import tqdm
def plot_graph(graph):
nx.draw(graph, with_labels=True,
node_color='skyblue', node_size=2200,
width=3, edge_cmap=plt.cm.OrRd,
arrowstyle='->',arrowsize=20,
font_size=10, font_weight="bold",
pos=nx.random_layout(init, seed=13))
plt.show()
ctr = 0
predicted_transformations = []
actual_transformations = []
factorials = {}
num_lines = sum(1 for line in open('C:/Users/Admin/Projects/Transformation Driven Visual Learning/data/data.jsonl', 'r'))
with open('C:/Users/Admin/Projects/Transformation Driven Visual Learning/data/data.jsonl', 'r') as f:
for line in tqdm(f, total=num_lines):
# print(line)
entry = json.loads(line)
data = {
'init_state': entry['states'][0]['objects'],
'init_image_file': entry['states'][0]['images']['Camera_Center'],
'final_state': entry['states'][1]['objects'],
'final_image_file': entry['states'][1]['images']['Camera_Center'],
'transformation': entry['transformations']
}
init_state = data['init_state']
final_state = data['final_state']
n = len(init_state)
objects = ['obj'+str(i) for i in range(n)]
init_edges = []
final_edges = []
init = nx.DiGraph()
final = nx.DiGraph()
init = nx.DiGraph()
final = nx.DiGraph()
attributes = {'colors' : [],
'materials' : [],
'shapes' : [],
'sizes' : [],
'positions': [],
}
# next cell
nodes = []
for i in range(n):
nodes.append(init_state[i]['color'])
nodes.append(init_state[i]['material'])
nodes.append(tuple(init_state[i]['position']))
nodes.append(init_state[i]['shape'])
nodes.append(init_state[i]['size'])
nodes.append(final_state[i]['color'])
nodes.append(final_state[i]['material'])
nodes.append(tuple(final_state[i]['position']))
nodes.append(final_state[i]['shape'])
nodes.append(final_state[i]['size'])
nodes = nodes + objects
init.add_nodes_from(nodes)
final.add_nodes_from(nodes)
# init.add_nodes_from(objects)
# final.add_nodes_from(objects)
# next cell
import itertools
if n in factorials.keys():
objectSet = factorials[n]
else:
objectSet = [list(i) for i in itertools.permutations(objects)]
ni = len(objectSet)
factorials[n] = objectSet
# next cell
def edge_creator(state, id=0):
objects = objectSet[id]
edges = []
for i in range(n):
edges.append((objects[i], state[i]['shape']))
edges.append((objects[i], state[i]['color']))
edges.append((objects[i], state[i]['size']))
edges.append((objects[i], tuple(state[i]['position'])))
edges.append((objects[i], state[i]['material']))
return edges
# next cell
init_edges = edge_creator(init_state)
i = 0
final_edges = edge_creator(final_state)
init.add_edges_from(init_edges)
final.add_edges_from(final_edges)
diff = symmetric_difference(init, final)
minChanges = len(diff.edges())
final.remove_edges_from(final_edges)
bestDiffGraph = diff
for i in range(1, ni):
final_edges = edge_creator(final_state, i)
diff = symmetric_difference(init, final)
final.remove_edges_from(final_edges)
changes = len(diff.edges())
if minChanges > changes:
minChanges = changes
bestDiffGraph = diff
if minChanges <= 8:
break
# next cell
# nx.draw(bestDiffGraph, with_labels=True,
# node_color='skyblue', node_size=2200,
# width=3, edge_cmap=plt.cm.OrRd,
# arrowstyle='->',arrowsize=20,
# font_size=10, font_weight="bold",
# pos=nx.random_layout(init, seed=13))
# next cell
subGraphs = []
for object in objects:
subGraph = nx.DiGraph()
subGraph.add_node(object)
nodes = bestDiffGraph.neighbors(object)
for node in nodes:
subGraph.add_node(node)
subGraph.add_edge(object, node)
subGraphs.append(subGraph)
changes = []
for i in range(len(subGraphs)):
nodes = list(subGraphs[i].nodes())
print(nodes)
if len(nodes) == 3:
changes.append([(nodes[0], nodes[1]), (nodes[0], nodes[2])])
predicted_transformations.append(changes)
# init.remove_edges_from(init_edges)
# init.remove_nodes_from(nodes)
# final.remove_nodes_from(nodes)
# print(predicted_transformations)
break
print(changes)
print(init.has_edge(*changes[0]))
init.edges()
nodes
with open('C:/Users/Admin/Projects/Transformation Driven Visual Learning/predicted_transformations.json', 'w') as fp:
json.dump(predicted_transformations, fp)
| Graph Based Predictor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/droidadroit/age-and-gender-classification/blob/master/AgeGender.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="7QQe6I3yhCY_" colab_type="text"
# # Introduction
# This is the implementation of [Age and Gender Classification by <NAME> and <NAME>](https://talhassner.github.io/home/projects/cnn_agegender/CVPR2015_CNN_AgeGenderEstimation.pdf) using PyTorch as the deep learning framework and Google Colab as the training ground. The network proposed in the paper has five convolutional layers and three fully connected layers, and is simple enough to understand and get familiar with PyTorch and Colab. Try it!
#
# Instead, if you just want to test the model on an image, go to the **Testing on an image** section.
# + [markdown] id="XCz5QH4dlh_H" colab_type="text"
# # Setup
# + [markdown] id="arTKN7rkgZsR" colab_type="text"
# Ensure that you are using Python 3.6 and GPU as the hardware accelerator. To check this, go to **Runtime -> Change runtime type** in the menu bar of Colab.
# + [markdown] id="O5NLRAhllX29" colab_type="text"
# ## Mounting the drive
# + id="uRVNRKKSVNLK" colab_type="code" colab={}
from google.colab import drive
drive.mount('/content/gdrive')
# + [markdown] id="Kpr2iINzADlV" colab_type="text"
# ## Directory structure
# Our project is in the directory **AgeGenderClassification**. Create this directory and the sub-directories inside it along with the downloaded **AgeGender.ipynb**.
#
# content
# -------- gdrive
# ---------------- My Drive
# ------------------------ **AgeGenderClassification**
# -------------------------------- **data**
# -------------------------------- **models**
# -------------------------------- **results**
# -------------------------------- **AgeGender.ipynb**
#
# + [markdown] id="wQvenRKomUkx" colab_type="text"
# ## Installing PyTorch
# This script installs PyTorch and torchvision.
# + id="MRHSqYujmgn-" colab_type="code" colab={}
from os.path import exists
from wheel.pep425tags import get_abbr_impl, get_impl_ver, get_abi_tag
platform = '{}{}-{}'.format(get_abbr_impl(), get_impl_ver(), get_abi_tag())
# cuda_output = !ldconfig -p|grep cudart.so|sed -e 's/.*\.\([0-9]*\)\.\([0-9]*\)$/cu\1\2/'
accelerator = cuda_output[0] if exists('/dev/nvidia0') else 'cpu'
# !pip install -q http://download.pytorch.org/whl/{accelerator}/torch-0.4.1-{platform}-linux_x86_64.whl torchvision
# + [markdown] id="UkNgiJoSm87s" colab_type="text"
# ## Imports
# + id="BRaqXIPonB8j" colab_type="code" colab={}
import torch
import torch.autograd.variable as Variable
import torchvision
import torchvision.transforms as transforms
import torch.optim as optim
import torch.nn as nn
import torch.nn.functional as F
import torchvision.utils as utils
from torch.utils.data import Dataset, DataLoader
# + id="1wIv9zyxtHxt" colab_type="code" colab={}
import numpy as np
import os
import matplotlib.pyplot as plt
from PIL import Image
from shutil import copyfile
# + [markdown] id="0_z6Q972xi32" colab_type="text"
# # Preparing dataloaders
# + [markdown] id="s_KtxziRPrVI" colab_type="text"
# ## **Raw data**
#
#
# + [markdown] id="8csd2D2v4-0O" colab_type="text"
# ### Downloading data
#
#
# We use the Adience dataset consisting unfiltered faces ([Link](http://www.cslab.openu.ac.il/download/adiencedb/AdienceBenchmarkOfUnfilteredFacesForGenderAndAgeClassification/aligned.tar.gz)). Then, we unzip it.
# The first cell below downloads the data for you and places it in the **data** directory. The second cell unzips the data.
# + id="4Tqb-HEKTLLo" colab_type="code" colab={}
# !wget --user adiencedb --password adience http://www.cslab.openu.ac.il/download/adiencedb/AdienceBenchmarkOfUnfilteredFacesForGenderAndAgeClassification/aligned.tar.gz -P "/content/gdrive/My Drive/AgeGenderClassification/data"
# + id="NPeBSiNqimxw" colab_type="code" colab={}
# !tar xvzf "/content/gdrive/My Drive/AgeGenderClassification/data/aligned.tar.gz" -C "/content/gdrive/My Drive/AgeGenderClassification/data/"
# + [markdown] id="88F5gWj5o4bh" colab_type="text"
# ### Downloading folds
#
# All five folds used in this paper are present [here](https://github.com/GilLevi/AgeGenderDeepLearning/tree/master/Folds/train_val_txt_files_per_fold). Download the **train_val_txt_files_per_fold** folder and place it in **My Drive/AgeGenderClassification/data**.
#
# + [markdown] id="2VtoAJghnpLn" colab_type="text"
# ## Data loading
# + id="squb12xguXqd" colab_type="code" colab={}
PATH_TO_FOLDS = "/content/gdrive/My Drive/AgeGenderClassification/data/train_val_txt_files_per_fold"
PATH_TO_DATA = "/content/gdrive/My Drive/AgeGenderClassification/data"
PATH_TO_IMAGE_FOLDERS = PATH_TO_DATA + "/aligned"
# + [markdown] id="O2Czi4va4QKo" colab_type="text"
# ### Creating a Dataset class
#
# We create a class **`AdienceDataset`** that extends **`Dataset`**. This class helps us in feeding the input data to the network in minibatches.
#
# [This](https://pytorch.org/tutorials/beginner/data_loading_tutorial.html) is a useful tutorial on how to load and augment data in PyTorch.
# + id="BiUufbSinr1K" colab_type="code" colab={}
class AdienceDataset(Dataset):
def __init__(self, txt_file, root_dir, transform):
self.txt_file = txt_file
self.root_dir = root_dir
self.transform = transform
self.data = self.read_from_txt_file()
def __len__(self):
return len(self.data)
def read_from_txt_file(self):
data = []
f = open(self.txt_file)
for line in f.readlines():
image_file, label = line.split()
label = int(label)
if 'gender' in self.txt_file:
label += 8
data.append((image_file, label))
return data
def __getitem__(self, idx):
img_name, label = self.data[idx]
image = Image.open(self.root_dir + '/' + img_name)
if self.transform:
image = self.transform(image)
return {
'image': image,
'label': label
}
# + [markdown] id="WAx-8hZNsXFM" colab_type="text"
# ### Transforms
# Every image is first resized to a `256x256` image and then cropped to a `227x227` image before being fed to the network.
#
# **`transforms_list`** is the list of transforms we would like to apply to the input data. Apart from training the neural network without any transformations, we can also train the network using the following transforms (also called as data augmentation techniques):
# * random horizontal flip
# * random crop and random horizontal flip
#
# We don't perform any transformation on the images during validation and testing.
#
# + id="_wMCsfG1sY4s" colab_type="code" colab={}
transforms_list = [
transforms.Resize(256),
transforms.CenterCrop(227),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
transforms.RandomCrop(227)
]
transforms_dict = {
'train': {
0: list(transforms_list[i] for i in [0, 1, 3]), # no transformation
1: list(transforms_list[i] for i in [0, 1, 2, 3]), # random horizontal flip
2: list(transforms_list[i] for i in [0, 4, 2, 3]) # random crop and random horizontal flip
},
'val': {
0: list(transforms_list[i] for i in [0, 1, 3])
},
'test': {
0: list(transforms_list[i] for i in [0, 1, 3])
}
}
# + [markdown] id="iURni95m4sGW" colab_type="text"
# ### Dataloader
# The **`DataLoader`** class in PyTorch helps us iterate through the dataset. This is where we input **`minibatch_size`** to our algorithm.
# + id="ox_PryLS4u2x" colab_type="code" colab={}
def get_dataloader(s, c, fold, transform_index, minibatch_size):
"""
Args:
s: A string. Equals either "train", "val", or "test".
c: A string. Equals either "age" or "gender".
fold: An integer. Lies in the range [0, 4] as there are five folds present.
transform_index: An integer. The transforms in the list correesponding
to this index in the dictionary will be applied on the images.
minibatch_size: An integer.
Returns:
An instance of the DataLoader class.
"""
txt_file = f'{PATH_TO_FOLDS}/test_fold_is_{fold}/{c}_{s}.txt'
root_dir = PATH_TO_IMAGE_FOLDERS
transformed_dataset = AdienceDataset(txt_file, root_dir,
transforms.Compose(transforms_dict[s][transform_index]))
dataloader = DataLoader(transformed_dataset, batch_size=minibatch_size, shuffle=True, num_workers=4)
return dataloader
# + [markdown] id="9lrstEuSm1qH" colab_type="text"
# # Network
# + id="o9feSnLcFq2q" colab_type="code" colab={}
device = torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')
# + id="EJ2QrXRPmCGs" colab_type="code" colab={}
PATH_TO_MODELS = "/content/gdrive/My Drive/AgeGenderClassification/models"
# + [markdown] id="SZh1psZ6mY6d" colab_type="text"
# ## Defining the network
# This is the network as described in the [paper](https://talhassner.github.io/home/projects/cnn_agegender/CVPR2015_CNN_AgeGenderEstimation.pdf).
# + id="pd91hyHvmhYP" colab_type="code" colab={}
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.conv1 = nn.Conv2d(3, 96, 7, stride = 4, padding = 1)
self.pool1 = nn.MaxPool2d(3, stride = 2, padding = 1)
self.norm1 = nn.LocalResponseNorm(size = 5, alpha = 0.0001, beta = 0.75)
self.conv2 = nn.Conv2d(96, 256, 5, stride = 1, padding = 2)
self.pool2 = nn.MaxPool2d(3, stride = 2, padding = 1)
self.norm2 = nn.LocalResponseNorm(size = 5, alpha = 0.0001, beta = 0.75)
self.conv3 = nn.Conv2d(256, 384, 3, stride = 1, padding = 1)
self.pool3 = nn.MaxPool2d(3, stride = 2, padding = 1)
self.norm3 = nn.LocalResponseNorm(size = 5, alpha = 0.0001, beta = 0.75)
self.fc1 = nn.Linear(18816, 512)
self.dropout1 = nn.Dropout(0.5)
self.fc2 = nn.Linear(512, 512)
self.dropout2 = nn.Dropout(0.5)
self.fc3 = nn.Linear(512, 10)
self.apply(weights_init)
def forward(self, x):
x = F.leaky_relu(self.conv1(x))
x = self.pool1(x)
x = self.norm1(x)
x = F.leaky_relu(self.conv2(x))
x = self.pool2(x)
x = self.norm2(x)
x = F.leaky_relu(self.conv3(x))
x = self.pool3(x)
x = self.norm3(x)
x = x.view(-1, 18816)
x = self.fc1(x)
x = F.leaky_relu(x)
x = self.dropout1(x)
x = self.fc2(x)
x = F.leaky_relu(x)
x = self.dropout2(x)
x = F.log_softmax(self.fc3(x), dim=1)
return x
# + id="uaI0Bs-Rxio-" colab_type="code" colab={}
def weights_init(m):
if isinstance(m, nn.Conv2d) or isinstance(m, nn.Linear):
nn.init.normal_(m.weight, mean=0, std=1e-2)
# + id="FrLA7cfxj4JE" colab_type="code" colab={}
criterion = nn.NLLLoss()
# + [markdown] id="qCTDc4XEyqOq" colab_type="text"
# ## Hyperparameters
# Try playing with these! While the **`minibatch_size`** and **`lr`** are pulled from the paper, **`num_epochs`** is set empirically.
# + id="w5Vm2Tb2yzYT" colab_type="code" colab={}
minibatch_size = 50
num_epochs = 60
lr = 0.0001 # initial learning rate
# + [markdown] id="G5NPiENkmpYl" colab_type="text"
# ## Training the network
# We save the network to the drive and compute the loss on validation set after every **`checkpoint_frequency`** number of iterations. We decrease the learning by a tenth after 10,000 iterations using the **`MultiStepLR`** class of PyTorch.
# + id="NnkM7TgymrED" colab_type="code" colab={}
def train(net, train_dataloader, epochs, filename, checkpoint_frequency, val_dataloader=None):
"""
Args:
net: An instance of PyTorch's Net class.
train_dataloader: An instance of PyTorch's Dataloader class.
epochs: An integer.
filename: A string. Name of the model saved to drive.
checkpoint_frequency: An integer. Represents how frequent (in terms
of number of iterations) the model should be saved to drive.
val_dataloader: An instance of PyTorch's Dataloader class.
Returns:
net: An instance of PyTorch's Net class. The trained network.
training_loss: A list of numbers that represents the training loss at each checkpoint.
validation_loss: A list of numbers that represents the validation loss at each checkpoint.
"""
net.train()
optimizer = optim.Adam(net.parameters(), lr)
scheduler = optim.lr_scheduler.MultiStepLR(optimizer, milestones=[10000])
training_loss, validation_loss = [], []
checkpoint = 0
iteration = 0
running_loss = 0
for epoch in range(epochs):
for i, batch in enumerate(train_dataloader):
scheduler.step()
optimizer.zero_grad()
images, labels = batch['image'].to(device), batch['label'].to(device)
outputs = net(images)
loss = criterion(outputs, labels)
running_loss += float(loss.item())
loss.backward()
optimizer.step()
if (iteration+1) % checkpoint_frequency == 0 and val_dataloader is not None:
training_loss.append(running_loss/checkpoint_frequency)
validation_loss.append(validate(net, val_dataloader))
print(f'minibatch:{i}, epoch:{epoch}, iteration:{iteration}, training_error:{training_loss[-1]}, validation_error:{validation_loss[-1]}')
save_network(net, f'{filename}_checkpoint{checkpoint}')
checkpoint += 1
running_loss = 0
iteration += 1
return net, training_loss, validation_loss
# + [markdown] id="x7WndciFmv0C" colab_type="text"
# ## Validation
# We evaluate the performance (in terms of loss) of the trained network on validation set.
# + id="8Es1uGKQnK9D" colab_type="code" colab={}
def validate(net, dataloader):
net.train()
total_loss = 0
with torch.no_grad():
for i, batch in enumerate(dataloader):
images, labels = batch['image'].to(device), batch['label'].to(device)
outputs = net(images)
loss = criterion(outputs, labels)
total_loss += float(loss.item())
return total_loss/(i+1)
# + id="sUyMnXqwauw9" colab_type="code" colab={}
def get_validation_error(c, fold, train_transform_index):
filename = get_model_filename(c, fold, train_transform_index)
net = Net().to(device)
net.load_state_dict(torch.load(f'{PATH_TO_MODELS}/{filename}'))
return validate(net, get_dataloader('val', c, fold, 0, minibatch_size))
# + [markdown] id="gAto0iO4zPLw" colab_type="text"
# ## Testing
# We evaluate the performance (in terms of accuracy) of the trained network on the test set.
# + id="fCUCNrL0zR3n" colab_type="code" colab={}
def test(net, dataloader, c):
result = {
'exact_match': 0,
'total': 0
}
if c == 'age':
result['one_off_match'] = 0
with torch.no_grad():
net.eval()
for i, batch in enumerate(dataloader):
images, labels = batch['image'].to(device), batch['label'].to(device)
outputs = net(images)
outputs = torch.tensor(list(map(lambda x: torch.max(x, 0)[1], outputs))).to(device)
result['total'] += len(outputs)
result['exact_match'] += sum(outputs == labels).item()
if c == 'age':
result['one_off_match'] += (sum(outputs==labels) +
sum(outputs==labels-1) +
sum(outputs==labels+1)).item()
return result
# + [markdown] id="mSq61rZ-yOPZ" colab_type="text"
# ## Saving the network
# + id="YzgCR7VqyQU6" colab_type="code" colab={}
def save_network(net, filename):
torch.save(net.state_dict(), f'{PATH_TO_MODELS}/{filename}.pt')
# + [markdown] id="hl1owOOw0ZeU" colab_type="text"
# # Execution
# + [markdown] id="fpye5rRQv2e6" colab_type="text"
# ### Picking the best model for a fold
# **`train_save()`** trains the network using the **`train()`** function and then, using the validation losses returned by this function at all checkpoints, chooses the model with least validation error. This function also plots a graph of training and validation errors over the iterations.
#
# **Usage:**
#
# For e.g., if you want to train the network for **`age`** using **`fold=2`** and **`train_transform_index=2`**,
# ```
# train_save('age', 2, 2)
# ```
#
#
# + id="ravxG5ce0bz0" colab_type="code" colab={}
def train_save(c, fold, train_transform_index, checkpoint_frequency=50):
"""
Args:
c: A string. Equals either "age" or "gender".
fold: An integer. Lies in the range [0, 4] as there are five folds present.
train_transform_index: An integer. The transforms in the list correesponding
to this index in the dictionary will be applied on the images.
checkpoint_frequency: An integer. Represents how frequent (in terms
of number of iterations) the model should be saved to drive.
Returns:
validation_loss: A list of numbers that represents the validation loss at each checkpoint.
"""
trained_net, training_loss, validation_loss = train(
Net().to(device),
get_dataloader('train', c, fold, train_transform_index, minibatch_size),
num_epochs,
f'{fold}_{c}_train_{train_transform_index}',
checkpoint_frequency,
get_dataloader('val', c, fold, 0, minibatch_size)
)
plt.plot(list(map(lambda x: checkpoint_frequency * x, (list(range(1, len(validation_loss)+1))))), validation_loss, label='validation_loss')
plt.plot(list(map(lambda x: checkpoint_frequency * x, (list(range(1, len(training_loss)+1))))), training_loss, label='training_loss')
plt.legend(bbox_to_anchor=(0., 1.02, 1., .102), loc=3,
ncol=2, mode="expand", borderaxespad=0.)
plt.xlabel('iterations')
plt.ylabel('loss')
plt.show()
choose_model_with_least_val_error(c, fold, train_transform_index, validation_loss)
return validation_loss
# + id="1bYLxmb_JupM" colab_type="code" colab={}
def choose_model_with_least_val_error(c, fold, train_transform_index, validation_loss):
index = validation_loss.index(min(validation_loss))
filename = f'{fold}_{c}_train_{train_transform_index}'
for file in os.listdir(PATH_TO_MODELS):
if file.startswith(filename):
if file.startswith(f'{filename}_checkpoint{index}'):
pass
else:
os.remove(f'{PATH_TO_MODELS}/{file}')
# + [markdown] id="F5R7ySD9xJdW" colab_type="text"
# ### Picking the best model among all the folds
#
# Using **`pick_best_model()`**, we can pick the model among various folds that gives us the best validation accuracy. The best model's name is appended with **_best **in the **models** directory.
#
# **Usage:**
#
# To pick the best model for **`age`**,
# ```
# pick_best_model('age')
# ```
#
#
# To pick the best model for **`gender`**,
#
# ```
# pick_best_model('gender')
# ```
# + id="sKRrMiSI6KsJ" colab_type="code" colab={}
def pick_best_model(c):
"""
Args:
s: A string. Equals either "train", "val", or "test".
c: A string. Equals either "age" or "gender".
"""
def fn_filter(file):
file_split = file.split('_')
return True if (len(file_split) == 5 and file_split[1] == c) else False
def fn_map(file):
file_split = file.split('_')
return get_validation_error(c, file_split[0], file_split[3])
files = list(filter(fn_filter, os.listdir(PATH_TO_MODELS)))
val_errors = list(map(fn_map, files))
min_val_error, file = min(zip(val_errors, files))
best_model = f'{PATH_TO_MODELS}/{file.split(".")[0]}_best.pt'
copyfile(f'{PATH_TO_MODELS}/{file}', best_model)
print(f'Picking {best_model} as the best model for {c}...')
# + [markdown] id="VhEJNE-PyNqE" colab_type="text"
# ### Calculating performance/accuracy
#
# We can check the performance of any model using the **`get_performance()`** function.
#
# **Usage:**
#
# To know the performance on **`age`** classification,
# ```
# get_performance('age')
# ```
#
#
# To know the performance on **`gender`** classification,
#
# ```
# get_performance('gender')
# ```
#
#
# + id="OsrQjMrtESM8" colab_type="code" colab={}
def get_performance(c):
"""
Args:
c: A string. Equals either "age" or "gender".
Returns:
A dictionary containing accuracy (and one-off accuracy for age) of the model.
"""
file = get_best_model_filename(c).split('_')
return get_performance_of_a_model('test', file[1], file[0], file[3])
# + id="ghOw4Hnpa2wg" colab_type="code" colab={}
def get_best_model_filename(c):
def fn_filter(file):
file_split = file.split('_')
return True if (len(file_split) == 6 and file_split[1] == c) else False
return list(filter(fn_filter, os.listdir(PATH_TO_MODELS)))[0]
# + id="Qo_p5XeGUVmd" colab_type="code" colab={}
def get_performance_of_a_model(s, c, fold, train_transform_index):
"""
Args:
s: A string. Equals either "train", "val", or "test".
c: A string. Equals either "age" or "gender".
fold: An integer. Lies in the range [0, 4] as there are five folds present.
transform_index: An integer. The transforms in the list correesponding
to this index in the dictionary will be applied on the images.
Returns:
A dictionary containing accuracy (and one-off accuracy for age) of the model.
"""
filename = get_model_filename(c, fold, train_transform_index)
net = Net().to(device)
net.load_state_dict(torch.load(f'{PATH_TO_MODELS}/{filename}'))
performance = test(
net,
get_dataloader(s, c, fold, 0, minibatch_size),
c
)
if c == 'age':
return {
'accuracy': performance['exact_match']/performance['total'],
'one-off accuracy': performance['one_off_match']/performance['total']
}
elif c == 'gender':
return {
'accuracy': performance['exact_match']/performance['total']
}
# + id="KQgMcSVibjYa" colab_type="code" colab={}
def get_model_filename(c, fold, train_transform_index):
start_of_filename = f'{fold}_{c}_train_{train_transform_index}_checkpoint'
for file in os.listdir(PATH_TO_MODELS):
if file.startswith(start_of_filename):
return file
# + [markdown] id="aj0XrK-f75EG" colab_type="text"
# ### How to run?
#
# 1. **Run `train_save()`:**
#
#
# > You can do this for different combinations of **`c`**, **`fold`**, and **`train_transform_index`**, where **`c={'age','gender'}`**, **`fold={0,1,2,3,4}`**, and **`train_transform_index={0,1,2}`**.
#
# > I suggest you to first train the network on all the folds for **either** **`age`** or **`gender`** and then proceed to the setp 2. Then, follow the same steps for the other class. Also, use **`train_transform_index=2`** as it gives smaller validation error due to random flipping and cropping.
#
# > **Note:** It is just the network's architecture that is the same for age and gender. They both are trained independently. Ultimately, we will be having two different networks with the same architecture, one to classify age and the other to classify gender.
#
# 2. **Picking the best model:**
#
# > Call **`pick_best_model()`** on either **`age`** or **`gender`**.
#
# 3. **Know the performance:**
#
# > Call **`get_performance()`** on either **`age`** or **`gender`** to know the final performance of the network on the test set.
#
#
#
#
# + [markdown] id="PqPEPILSzrKW" colab_type="text"
# # Results
# + [markdown] id="Qi1mAy6-CeOK" colab_type="text"
# ## Age
# + [markdown] id="IRZmuqBYvsKO" colab_type="text"
# ### Plots of validation and training losses:
# These are graphs we get on calling **`train_save()`** on age for different folds.
#
# #### **Fold 0**
# 
#
# ---
#
# #### **Fold 1**
# 
#
# ---
#
# #### **Fold 2**
# 
#
# ---
#
# #### **Fold 3**
# 
#
# ---
#
# #### **Fold 4**
# 
#
#
# + [markdown] id="oQ483io4Ctax" colab_type="text"
# ### Accuracy
# These are the accurcies we get on calling **`get_performance()`** on **`age`**.
#
# ```
# {'accuracy': 0.5272136474411048, 'one-off accuracy': 0.8399675060926076}
# ```
#
#
# + [markdown] id="5JNvwScSwz8u" colab_type="text"
# ## Gender
# + [markdown] id="QD6Y3GnOQpLl" colab_type="text"
# ### Plots of validation and training losses:
# These are graphs we get on calling **`train_save()`** on gender for different folds.
#
# #### **Fold 0**
# 
#
# ---
#
# #### **Fold 1**
# 
#
# ---
#
# #### **Fold 2**
# 
#
# ---
#
# #### **Fold 3**
# 
#
# ---
#
# #### **Fold 4**
# 
# + [markdown] id="WzBBnRYpNqsH" colab_type="text"
# ### Accuracy
# This is the accuracy we get on calling **`get_performance()`** on **`gender`**.
#
# ```
# {'accuracy': 0.8437770719029744}
# ```
#
#
# + [markdown] id="fmgl8BZyZToa" colab_type="text"
# # Testing on an image
# + [markdown] id="5nLgK_DMQVpL" colab_type="text"
# You can either download the trained models for age and gender classification from [here](https://github.com/droidadroit/age-and-gender-classification/tree/master/models) into the **models** directory or rename the best models for age and gender upon training with **age.pt** and **gender.pt** respectively.
#
#
# For the prediction of age and gender from an image, simply call **`test()`** providing the path (in your Google Drive) of the image as the argument.
#
# **Usage:**
# `test("/content/gdrive/My Drive/AgeGenderClassification/test.jpeg")`
# + id="aeIlSTv2KAQt" colab_type="code" colab={}
PATH_TO_AGE_MODEL = f'{PATH_TO_MODELS}/age.pt'
PATH_TO_GENDER_MODEL = f'{PATH_TO_MODELS}/gender.pt'
# + id="m8UQqvFby3XK" colab_type="code" colab={}
mapping = {
0: '0-2 years',
1: '4-6 years',
2: '8-13 years',
3: '15-20 years',
4: '25-32 years',
5: '38-43 years',
6: '48-53 years',
7: '60 years and above',
8: 'male',
9: 'female'
}
# + id="VG9zRkATZW1C" colab_type="code" colab={}
def test_on_a_class(c, image_tensor):
with torch.no_grad():
net = Net().to(device)
net.load_state_dict(torch.load(f'{PATH_TO_MODELS}/{c}.pt'))
net.eval()
output = net(image_tensor)
output = torch.max(output, 1)[1].to(device)
result = f'{c} = {mapping[output.item()]}'
return result
# + id="Vlrupi7_EXYY" colab_type="code" colab={}
def test(path):
image = Image.open(path)
plt.imshow(image)
image = transforms.Compose(transforms_dict['test'][0])(image)
image.unsqueeze_(0)
image = image.to(device)
print(test_on_a_class('age', image))
print(test_on_a_class('gender', image))
# + [markdown] id="EpNbkMh_RNKg" colab_type="text"
# ## Examples
# + colab_type="code" id="wAvhu1z6RIRX" outputId="934134d1-5d7d-4024-c7de-7d9658338e1b" colab={"base_uri": "https://localhost:8080/", "height": 360}
test("/content/gdrive/My Drive/AgeGenderClassification/test2.jpg")
# + colab_type="code" outputId="10edc639-a419-4b9d-e31d-5d7433e1f51b" id="-EqG8ushRHfv" colab={"base_uri": "https://localhost:8080/", "height": 384}
test("/content/gdrive/My Drive/AgeGenderClassification/test6.jpeg")
# + [markdown] id="Ki0hQatZBRTA" colab_type="text"
# # Issues while running:
#
#
# *
# **```
# AttributeError: module 'PIL.Image' has no attribute 'register_extensions'
# ```**
#
#
# > In such cases, restart the runtime by pressing **Ctrl + M** or going to **Runtime->Restart runtime** in the menu bar of Colab. Then, run the code again.
#
#
#
#
#
| AgeGender.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (tensorflow)
# language: python
# name: tensorflow
# ---
import numpy as np
import cv2
import trimesh
import os
from paz.core import Pose6D
import json
from mrcnn import visualize, utils
from paz.core import ops
from mrcnn import visualize, utils
import tensorflow as tf
import paz.processors as pr
import matplotlib.pyplot as plt
from pose_estimation.solver import PnPSolver
from pose_estimation.pose_evaluation import PoseError
from pose_estimation.pose import compute_poses
import pyrender
ROOT_DIR = os.path.abspath("../../../")
MESH_DIR = '/home/incendio/Documents/Thesis/YCBVideo_detector/color_meshes'
class_names = ['background', '002_master_chef_can', '003_cracker_box',
'004_sugar_box', '005_tomato_soup_can', '006_mustard_bottle',
'007_tuna_fish_can', '008_pudding_box', '009_gelatin_box',
'010_potted_meat_can', '011_banana', '019_pitcher_base',
'021_bleach_cleanser', '024_bowl',
'025_mug', '035_power_drill',
'036_wood_block', '037_scissors',
'040_large_marker', '051_large_clamp',
'052_extra_large_clamp', '061_foam_brick']
def evaluation(gt_mask, pred_mask, true_id, name):
pnp = PnPSolver(gt_mask, true_id, name)
gt_points3d, image2d = pnp.get_points()
camera = pnp.compute_camera_matrix()
_, true_R, true_T, inliers = ops.solve_PNP(gt_points3d, image2d, camera, ops.UPNP)
# model_point = pnp.get_object_points(image2d)
# model_point = np.diag(model_point)[..., np.newaxis].T
# true_R, true_T, _ = pnp.solve_PnP()
est_pnp = PnPSolver(pred_mask, true_id, name)
points3d, image2d = est_pnp.get_points()
_, est_R, est_T, est_inliers = ops.solve_PNP(points3d, image2d, camera, ops.UPNP)
# est_R, est_T, _ = est_pnp.solve_PnP()
error = PoseError(gt_points3d, est_R, est_T, true_R, true_T)
add = error.add()
adi = error.adi()
rot_error = error.rotational_error()
trans_error = error.translational_error()
return add, adi, rot_error, trans_error
# +
data_dir = os.path.join(ROOT_DIR, 'for_pose_data/train')
mask_dir = data_dir + '/mask/'
with open(data_dir+'/pose_info.json', 'r') as f:
pose_data = json.load(f)
camera_to_world = np.array([[ 7.0710e-01, -4.0824e-01, 5.7735e-01, 9.9751e-01],
[ 6.0047e-17, 8.1649e-01, 5.7735e-01, 9.8064e-01],
[-7.0710e-01, -4.0824e-01, 5.7735e-01, 1.0218],
[ 0., 0., 0., 1.]])
world_to_camera = np.array([[ 0.70710678, 0., -0.70710678, 0.01674194],
[-0.40824829, 0.81649658, -0.40824829, -0.01203142],
[ 0.57735027, 0.57735027, 0.57735027, -1.73205081],
[ 0., 0., 0., 1.]])
for name in os.listdir(mask_dir):
pose_id, _ = name.split('.')
_, class_id = pose_id.split('_')
# pose = np.array(pose_data[pose_id])
gt_pose = np.dot(world_to_camera, np.array(pose_data[pose_id]))
mask = cv2.imread(mask_dir + '/' + name)
mask = cv2.cvtColor(mask, cv2.COLOR_BGR2RGB)
class_name = class_names[int(class_id)]
# +
#Ground-truth saved poses
pnp = PnPSolver(mask, int(class_id), class_name, color=(255, 0, 0), dimension=[.08, .08])
camera = pnp.compute_camera_matrix()
rot_test = gt_pose[:3, :3]
pose6D = Pose6D.from_rotation_vector(cv2.Rodrigues(rot_test)[0], gt_pose[:, 3][:3], class_name)
dimensions = {class_name: [.08, .08]}
pose = {'pose6D': pose6D, 'image': mask, 'color': (0, 255, 0)}
draw = pr.DrawBoxes3D(camera, dimensions)
args, projected_points = draw(pose)
fig = plt.figure(figsize=(10, 10))
plt.imshow(args['image'])
# -
#Poses from PNP_RANSAC algorithm
pnp = PnPSolver(mask, int(class_id), class_name, color=(255, 0, 0), dimension=[.08, .08])
points3d, image2d = pnp.get_points()
_, est_R, est_T, est_inliers = ops.solve_PNP(points3d, image2d, pnp.camera, ops.UPNP)
# camera = pnp.compute_camera_matrix()
# points3d, image2D = pnp.get_points()
# (_, rotation, translation, _) = ops.solve_PNP(points3d, image2D, camera, ops.UPNP)
gt_poses = pnp.solve_PnP()
args, pr_points = pnp.visualize_3D_boxes(mask, gt_poses)
fig = plt.figure(figsize=(10, 10))
plt.imshow(args['image'])
true_R = cv2.Rodrigues(rot_test)[0]
true_T = gt_pose[:, 3][:3]
error = PoseError(points3d, true_R, true_T, est_R, est_T)
add = error.add()
adi = error.adi()
rot_error = error.rotational_error()
trans_error = error.translational_error()
print(add, adi, rot_error, trans_error)
np.dot(world_to_camera, pose)
gt_poses
# print(np.round(pose[:3, :3]))
gt_trans = pose[:, 3][:3]
pred_trans = translation
print('saved pose..', gt_trans)
print('from pnp algorithm', pred_trans)
print(np.linalg.norm(gt_trans - pred_trans))
# r_est, _ = cv2.Rodrigues(rotation)
# print(np.rad2deg(np.arccos((np.trace(cv2.Rodrigues(rotation)[0].dot(pose[:3, :3])) - 1) / 2)))
# print(cv2.Rodrigues(rotation)[0])
# print(translation)
# +
world_to_camera = np.array([[ 0.70710678, 0., -0.70710678, 0.01674194],
[-0.40824829, 0.81649658, -0.40824829, -0.01203142],
[ 0.57735027, 0.57735027, 0.57735027, -1.73205081],
[ 0., 0., 0., 1.,]])
camera_to_world = np.array([[ 7.0710e-01, -4.0824e-01, 5.7735e-01, 9.9751e-01],
[ 6.0047e-17, 8.1649e-01, 5.7735e-01, 9.8064e-01],
[-7.0710e-01, -4.0824e-01, 5.7735e-01, 1.0218],
[ 0., 0., 0., 1.]])
for name in os.listdir(mask_dir):
pose_id, _ = name.split('.')
pose = np.dot(world_to_camera, np.array(pose_data[pose_id]))
# print(pose)
mesh_path = os.path.join(MESH_DIR, '03_sugar_box.obj')
mesh = trimesh.load(mesh_path)
# before = mesh.visual.vertex_colors[:, :3]
mesh.visual.vertex_colors[:, :3] = [10, 20, 30]
vertex_colors = mesh.visual.vertex_colors[:, :3]
mesh.apply_transform(pose)
# after = mesh.visual.vertex_colors[:, :3]
mesh = pyrender.Mesh.from_trimesh(mesh)
camera = pyrender.PerspectiveCamera(yfov=np.pi / 3.0, aspectRatio=1.0)
s = np.sqrt(2)/2
camera_pose = np.array([[0.0, -s, s, 0.3],
[1.0, 0.0, 0.0, 0.0],
[0.0, s, s, 0.35],
[0.0, 0.0, 0.0, 1.0]])
renderer = pyrender.OffscreenRenderer(viewport_width=320, viewport_height=320)
scene = pyrender.Scene(bg_color=np.zeros(4), ambient_light=np.ones(3))
scene.add(mesh)
scene.add(camera, pose=camera_pose)
img, _ = renderer.render(scene, flags=pyrender.constants.RenderFlags.FLAT)
# scene = pyrender.Scene()
# scene.add(mesh)
# pyrender.Viewer(scene, use_raymond_lighting=True)
# -
plt.imshow(img)
print(vertex_colors)
# +
data_dir = os.path.join(ROOT_DIR, 'dummy_data/train')
mask_dir = data_dir + '/mask/'
with open(data_dir+'/pose_info.json', 'r') as f:
pose_data = json.load(f)
masks = []
world_to_camera = np.array([[ 0.70710678, 0., -0.70710678, 0.01674194],
[-0.40824829, 0.81649658, -0.40824829, -0.01203142],
[ 0.57735027, 0.57735027, 0.57735027, -1.73205081],
[ 0., 0., 0., 1.,]])
def get_transformed_mesh(true_id, pose):
for name in os.listdir(MESH_DIR):
class_id = name.split('_')[0]
if int(class_id) == true_id:
mesh_path = os.path.join(MESH_DIR, name)
mesh = trimesh.load(mesh_path)
mesh = mesh.apply_transform(pose)
vertex_colors = mesh.visual.vertex_colors[:, :3]
return mesh, vertex_colors
for name in os.listdir(mask_dir):
pose_id, _ = name.split('.')
pose = np.dot(world_to_camera, np.array(pose_data[pose_id]))
class_id = int(name.split('_')[1].split('.')[0])
mesh, vertex_colors = get_transformed_mesh(class_id, pose)
mask = cv2.imread(mask_dir + name)
mask = cv2.cvtColor(mask, cv2.COLOR_BGR2RGB)
rows, cols, _ = np.where(mask > 0)
for index in range(len(rows)):
x, y = rows[index], cols[index]
x_index = np.where(mask[x, y, :].all() == vertex_colors.all())
print(x_index)
# r_index = np.where(vertex_colors[:, 0] == R)[0]
# g_index = np.where(vertex_colors[:, 1] == G)[0]
# b_index = np.where(vertex_colors[:, 2] == B)[0]
# d = [r_index, g_index, b_index]
# print(set(d[0]).intersection(*d))
# rg = np.intersect1d(r_index, g_index)
# rb = np.intersect1d(r_index, b_index)
# bg = np.intersect1d(b_index, g_index)
# if (rg.all() and rb.all() and bg.all()):
# +
path = './data_verification/data_inspection.json'
with open(path, 'r') as f:
data = json.load(f)
class_names = ['background', '002_master_chef_can', '003_cracker_box',
'004_sugar_box', '005_tomato_soup_can', '006_mustard_bottle',
'007_tuna_fish_can', '008_pudding_box', '009_gelatin_box',
'010_potted_meat_can', '011_banana', '019_pitcher_base',
'021_bleach_cleanser', '024_bowl',
'025_mug', '035_power_drill',
'036_wood_block', '037_scissors',
'040_large_marker', '051_large_clamp',
'052_extra_large_clamp', '061_foam_brick']
img = cv2.imread('./data_verification/original.png')
image = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
class_id = np.array(data['class_id'])
boxes = np.array(data['boxes'])
masks = np.array(data['masks'])
print(boxes)
visualize.display_rgb_instances(image, boxes, masks, class_id, class_names, figsize=(10, 10))
# -
| samples/ycbv/verify_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Array (Dizi) Yapısı
# ## İçerik
# * [Arrays](#1)
# * [Dynamic Array](#2)
# * [Dynamic Array with Python](#3)
# * [Array İş Mülakatları Soru-Cevap](#4)
# * [Array Python Challenge/Problem](#5)
# * [Neler Öğrendik](#6)
# <a id="1"></a>
# ## Arrays
# * 1 mesafe sensörü düşünün. Saniye de 1 veri yollayan bir sensör.
# * Bu verileri depolamak için variable tanımlayabilir miyiz?
# * Peki variable yerine ne kullanabiliriz? ARRAYS
# * Value depolamamıza yarayan yapılara array diyoruz.
# * Örn: [1,2,3,4,5] = higher level abstraction yani real world'de böyle kullanılır.
# * Bu array 5 elemanlı bir array.
# * Arrayin içinde ki her bir elemanın yerini indexler belirler.
# * 2 sayısı yani değeri index 1 dedir.
# * Biz tüm bu kursumuzda python kullandığımız gibi data structure konusunda da python kullanacağız ve python da index sıfırdan başlar.
import numpy as np
array = np.array([[1,2,3,4,5]]) # vector 1D
print(array)
print("Boyut: ",array.shape)
# * Peki array tek boyulu olmak zorunda mı? Hayır 2 boyutlu hatta 3-4 boyutlu bile olabilir. (Optional: tensor is lower or higher dimensinal array)
# * 2D array satır ve sütunlardan oluşur. Row - Column
array2D = np.array([[1,2,3,4,5],
[6,7,8,9,10]])
print(array2D)
print("boyut: ",array2D.shape)
print("2. satır 4. sütun: ",array2D[1,3])
# <a id="2"></a>
# ## Dynamic Array
# * Dynamic array boyutunu önceden belirtmek zorunda olmadığımız daha sonra eleman ekleyip çıkartabileceğimiz yapılara denir.
# * Dynamic array growable and resizable olarak tanımlanır.
# * Dynamic array neredeyse tüm programlama dillerinde çok sık kullanılan bir yapıdır.
# * 
# <a id="3"></a>
# ## Dynamic Array with Python
# * Anlamak için kendi dynamic array class'ımızı yaratacağız.
# * Eğer class ile ilgili sorunlarız var ise bu linkte ki benim dersimin object oriented kısmından öğrenebilirsiniz:
# * https://www.udemy.com/python-sfrdan-uzmanlga-programlama-1/
# +
import ctypes # yeni array yaratmak icin kullanacagiz
class DynamicArray(object):
# initialize (constructor)
def __init__(self):
self.n = 0 # eleman sayisi
self.capacity = 1 # kapasite
self.A = self.make_array(self.capacity)
def __len__(self):
"""
return array icerisinde eleman sayisi
"""
return self.n
def __getitem__(self,k):
"""
return index k'da ki eleman(value)
"""
if not 0 <= k < self.n:
return IndexError("k is out of bounds !")
return self.A[k]
def append(self,eleman):
"""
array'e eleman ekler
"""
# eger kapasite dolu ise kapasiteyi iki katina cikar
if self.n == self.capacity:
self._resize(2*self.capacity)
self.A[self.n] = eleman # eleman ekleme
self.n += 1 # eleman sayisi bir arttir
def _resize(self,new_cap):
"""
array kapasitesini arttir
"""
B = self.make_array(new_cap) # yeni array yap
# eski array (A) icerisindeki degerleri yeni arraye(B) icine tasi
for k in range(self.n):
B[k] = self.A[k]
self.A = B # arrayi guncelle
self.capacity = new_cap # kapasite guncelle
def make_array(self,new_cap):
"""
return yeni array
"""
return (new_cap*ctypes.py_object)()
# -
# obje tanimla
arr = DynamicArray()
# append new element 1
arr.append(1)
print(arr[0])
# append new element 1 , 3
arr.append(3)
print(arr[0],arr[1])
# append new element 1 , 3 ,5
arr.append(5)
print(arr[0],arr[1],arr[2])
# <a id="4"></a>
# ## Array İş Mülakatları Soru-Cevap
# 1. Dynamic Array neden önemli?
# * Automatic resizing
# 2. Array ile Dynamic Array arasındaki fark nedir?
# * Array is fixed size. Başlangıçta yani array yaratırken size belirlemek lazım
# * Dynamic array de başlangıçta size belirlemeye gerek yok
# 3. Dynamic Array Advantages and Disadvantages ?
# * Advantages:
# * Fast lookups: istenilen arrayde ki degeri elde etmesi
# * Variable size: resizeable
# * Disadvantages:
# * Slow worst-case appends: eger yer yoksa kapasiteyi arttırmak lazım ama yavas.
# * Costly inserts and deletes
# * 
# 4. Size Nasıl Artıyor?
# * 2 katına çıkıyor
# 5. Which of the following operations is not O(1) for an array of sorted data. You may assume that array elements are distinct.
# * (A) Find the ith largest element
# * (B) Delete an element
# * (C) Find the ith smallest element
# * (D) All of the above
# * (B)
# <a id="5"></a>
# ## Array Python Challenge/Problem
# * Brute force çözmeye çalışın yani kağıt kalem düz mantık for döngüleri filan kullanara, metot kullanmadan.
# * Eğer soruyu çözemezseniz bırakın 1 gün sonra tekrar bakın
# * Array Python Challenges/Problems
# 1. Word Split
# ### 1) Word Split
# * input = ["deeplearning", "d,dll,a,deep,dee,base,lear,learning"]
# * output = ["deep,learning"]
def wordSplit(liste):
word = list((liste[0])) # "deeplearning" => ["d","e"...]
d = liste[1].split(",") # ["d","dll","a","deep","dee","base","lear","learning"]
for i in range(1,len(word)):
c = word[:]
c.insert(i," ")
x , y = "".join(c).split() # "d", "eeplearning" "de","eplearning"
if x in d and y in d:
return x + " , "+ y
return "bulamadim (no way)"
print(wordSplit(["deeplearning", "d,dll,a,deep,dee,base,lear,learning"]))
print(wordSplit(["deeplear2ning", "d,dll,a,deep,dee,base,lear,learning"]))
print(wordSplit(["deeplear2ning", "d,dll,a,deep,dee,base,lear,lear2ning"]))
# <a id="6"></a>
# ## Neler Öğrendik
# * Arrays
# * Dynamic Array
# * Dynamic Array with Python
# * Array İş Mülakatları Soru-Cevap
# * Array Python Challenge/Problem
#
# * TAVSİYE: Array ile ilgili soru çözün. Mesela https://coderbyte.com/
| Data Structures/Arrays/Array (Dizi) Yapısı.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="giR0ibyi9GZm"
# # **Set Up**
# + id="DZEFKIjKyMxw" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1617673674143, "user_tz": 240, "elapsed": 382, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}} outputId="261fd6db-7ab1-4a96-f80e-3c2db32537b3"
# import libraries
import pandas as pd
import numpy as np
# %matplotlib inline
import math
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import pickle
import os
import datetime
from wordcloud import WordCloud
from tensorflow import keras
import tensorflow as tf
from keras.callbacks import Callback, EarlyStopping, ModelCheckpoint
from keras.models import Model
from keras.layers import Input, Reshape, Dot
from keras.layers.embeddings import Embedding
print("Keras version " + keras.__version__)
print("Tensorflow version " + tf.__version__)
# + id="8yw6-4Tq2kBX" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1617673521442, "user_tz": 240, "elapsed": 20599, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}} outputId="1ebff91c-e5c1-47be-afaf-86b5ff671e77"
from google.colab import drive
drive.mount("/content/drive")
# + [markdown] id="QPzFFR3i9iV-"
# # **Data Loading and Preprocessing**
# + id="nzDU60oQ2pZG" executionInfo={"status": "ok", "timestamp": 1617673523197, "user_tz": 240, "elapsed": 354, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}}
base_path = '/content/drive/My Drive/DATA2040_Final_Project_YARD'
# + id="ksMBFb5UV7NG" executionInfo={"status": "ok", "timestamp": 1617673526275, "user_tz": 240, "elapsed": 2609, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}}
ratings = pd.read_csv(base_path + '/data/ratings.csv', sep='\t', encoding='latin-1',
usecols=['user_id', 'movie_id', 'user_emb_id', 'movie_emb_id', 'rating'])
# + colab={"base_uri": "https://localhost:8080/", "height": 534} id="Ye80pMPCWFZm" executionInfo={"status": "ok", "timestamp": 1617673527382, "user_tz": 240, "elapsed": 305, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}} outputId="aea6b170-d76e-43a6-ba36-b501c7085c4c"
ratings
# + colab={"base_uri": "https://localhost:8080/"} id="BURhMhv9WMUw" executionInfo={"status": "ok", "timestamp": 1617673529177, "user_tz": 240, "elapsed": 283, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}} outputId="ed9e60cb-6ef5-425e-8021-bf26c13df07a"
max_userid = ratings['user_id'].drop_duplicates().max()
max_movieid = ratings['movie_id'].drop_duplicates().max()
print(max_userid)
print(max_movieid)
# + id="5e52Tozo2JGL" executionInfo={"status": "ok", "timestamp": 1617673530862, "user_tz": 240, "elapsed": 628, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}}
users = pd.read_csv(base_path + '/data/users.csv', sep='\t', encoding='latin-1',
usecols=['user_id', 'gender', 'zipcode', 'age_desc', 'occ_desc'])
# + colab={"base_uri": "https://localhost:8080/", "height": 534} id="SCvEQn4m3H4_" executionInfo={"status": "ok", "timestamp": 1617673531877, "user_tz": 240, "elapsed": 243, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}} outputId="29059ccc-1195-4c64-fdce-3980ae613fb5"
users
# + id="kH9Vvx5L6KTi" executionInfo={"status": "ok", "timestamp": 1617673534013, "user_tz": 240, "elapsed": 408, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}}
movies = pd.read_csv(base_path + '/data/movies.csv', sep='\t', encoding='latin-1',
usecols=['movie_id', 'title', 'genres'])
# + colab={"base_uri": "https://localhost:8080/", "height": 534} id="utCVhoGyWcrc" executionInfo={"status": "ok", "timestamp": 1617673534929, "user_tz": 240, "elapsed": 399, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}} outputId="b205dd00-948f-43ef-c3fa-2301348aed31"
movies
# + colab={"base_uri": "https://localhost:8080/"} id="UUJYGu9yWuNw" executionInfo={"status": "ok", "timestamp": 1617673536813, "user_tz": 240, "elapsed": 403, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}} outputId="9d47b459-9915-4d93-9ce8-d54c1c667621"
# Create training set
RNG_SEED = 32
shuffled_ratings = ratings.sample(frac=1., random_state=RNG_SEED)
# Shuffling users
Users = shuffled_ratings['user_emb_id'].values
print('Users:', Users, ', shape =', Users.shape)
# Shuffling movies
Movies = shuffled_ratings['movie_emb_id'].values
print('Movies:', Movies, ', shape =', Movies.shape)
# Shuffling ratings
Ratings = shuffled_ratings['rating'].values
print('Ratings:', Ratings, ', shape =', Ratings.shape)
# + [markdown] id="KIJ2nLRr9rZk"
# # **Exploratory Data Analysis**
# + [markdown] id="IwCV-9wiGYXh"
# **MOVIES**
# + colab={"base_uri": "https://localhost:8080/"} id="1oeAdHUhm_RK" executionInfo={"status": "ok", "timestamp": 1617673539515, "user_tz": 240, "elapsed": 265, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}} outputId="17c51f2e-ce72-4298-c712-7d4b32fdf55f"
# get movie shape
movies.shape
# + [markdown] id="I8tA_IT7pV5Z"
# There are 3883 movies in the dataset. The first column contains the movie ID (integers), the second column contains the movie titles (along with the year it was released), and the third column contains the movie genres.
# + colab={"base_uri": "https://localhost:8080/", "height": 260} id="QfQALp1dorMQ" executionInfo={"status": "ok", "timestamp": 1617673541385, "user_tz": 240, "elapsed": 245, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}} outputId="10096b2d-da74-4a0c-98b7-eb6c4ffdb537"
movies.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 380} id="4Gih5TaSn0Ji" executionInfo={"status": "ok", "timestamp": 1617673542816, "user_tz": 240, "elapsed": 281, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}} outputId="29aac919-299f-4deb-be24-502fad230f1b"
movies.describe()
# + colab={"base_uri": "https://localhost:8080/"} id="AWyO92OEqjP5" executionInfo={"status": "ok", "timestamp": 1617673544163, "user_tz": 240, "elapsed": 235, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}} outputId="3e238acc-d787-4517-95b1-b312be97424f"
# check for missing values
movies.isnull().sum(axis=0)
# + colab={"base_uri": "https://localhost:8080/"} id="rCd4sxkOqrl3" executionInfo={"status": "ok", "timestamp": 1617673545791, "user_tz": 240, "elapsed": 231, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}} outputId="a6b2a4ea-54b5-4483-ab52-f23a2f288137"
# check count of different genres
movies['genres'].value_counts()
# + colab={"base_uri": "https://localhost:8080/"} id="dugRcolCq20A" executionInfo={"status": "ok", "timestamp": 1617673547197, "user_tz": 240, "elapsed": 311, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}} outputId="5d83165c-c2ca-4b9a-9c4d-0ec51ae4099c"
# check number of unique genres
movies['genres'].value_counts().nunique()
# + id="0U5FfV9Xq7yY" executionInfo={"status": "ok", "timestamp": 1617673548831, "user_tz": 240, "elapsed": 428, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}}
# merge movies and ratings
movies_ratings = pd.merge(movies, ratings)
# + colab={"base_uri": "https://localhost:8080/", "height": 260} id="Abnm3pllriiZ" executionInfo={"status": "ok", "timestamp": 1617673550458, "user_tz": 240, "elapsed": 312, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}} outputId="612c925d-4274-4e58-9509-63493bb46a4d"
movies_ratings.head()
# + colab={"base_uri": "https://localhost:8080/"} id="ExzwYdYEryFu" executionInfo={"status": "ok", "timestamp": 1617673551825, "user_tz": 240, "elapsed": 255, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}} outputId="730092b2-282a-4e3c-f348-3e1e7e700bce"
top10 = movies_ratings.groupby('title').size().sort_values(ascending=False)[:10]
top10
# + colab={"base_uri": "https://localhost:8080/", "height": 590} id="DJHlHpy7VhS_" executionInfo={"status": "ok", "timestamp": 1617673554649, "user_tz": 240, "elapsed": 1555, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}} outputId="775c5ae7-f006-48c9-e50f-0ce2f4768c03"
#Plot Top 10 Movies Viewed
movies_ratings.groupby('title')['user_id'].count().sort_values(ascending = False)[:10].plot(kind ='barh', figsize = (10,9), color= "pink")
plt.xlabel('Count')
plt.ylabel('Movie Titles')
plt.title('Top 10 Viewed Movies')
plt.savefig(base_path + '/figures/plt/' + 'Bar chart Top 10 Movies Viewed.jpg',bbox_inches='tight')
plt.show
# + [markdown] id="uwNfcj0gWD_r"
# The graph shows the top 10 movies viewed (each user can only be counted once) by count. American Beauty is the highest followed by the 3 Star Wars Movies.
# + colab={"base_uri": "https://localhost:8080/"} id="8pCMRFCiTQVL" executionInfo={"status": "ok", "timestamp": 1617673556363, "user_tz": 240, "elapsed": 292, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}} outputId="391d2f5d-6b74-489b-8671-3818f32d3e21"
#counts unique genres
n = len(pd.unique(movies_ratings ['genres']))
print("No.of.unique values :", n)
# + id="orOaxQfrTmlN" executionInfo={"status": "ok", "timestamp": 1617673557759, "user_tz": 240, "elapsed": 252, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}}
#define a function that counts the number of times each genre appear:
def count_word(df, ref_col, liste):
keyword_count = dict()
for s in liste: keyword_count[s] = 0
for liste_keywords in movies_ratings[ref_col].str.split('|'):
if type(liste_keywords) == float and pd.isnull(liste_keywords): continue
for s in liste_keywords:
if pd.notnull(s): keyword_count[s] += 1
# convert the dictionary in a list to sort the keywords by frequency
keyword_occurences = []
for k,v in keyword_count.items():
keyword_occurences.append([k,v])
keyword_occurences.sort(key = lambda x:x[1], reverse = True)
return keyword_occurences, keyword_count
# + id="ny8bBZaiTzkh" executionInfo={"status": "ok", "timestamp": 1617673561985, "user_tz": 240, "elapsed": 2975, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}}
genre_labels = set()
for s in movies_ratings['genres'].str.split('|').values:
genre_labels = genre_labels.union(set(s))
# + colab={"base_uri": "https://localhost:8080/"} id="zXFrgj5wTm2p" executionInfo={"status": "ok", "timestamp": 1617673566448, "user_tz": 240, "elapsed": 3785, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}} outputId="26aee761-d691-4eb9-d6be-2641404c4e01"
keyword_occurences, dum = count_word(movies_ratings, 'genres', genre_labels)
#keyword_occurences
trunc_occurences = keyword_occurences[0:50]
trunc_occurences
# + colab={"base_uri": "https://localhost:8080/", "height": 492} id="slA4jDJtUOgO" executionInfo={"status": "ok", "timestamp": 1617673568614, "user_tz": 240, "elapsed": 1116, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}} outputId="4f86abbc-f784-4d2f-83fd-239982c5b364"
# Bar graph of popularity of genres
fig = plt.figure(1, figsize=(18,13))
ax2 = fig.add_subplot(2,1,2)
y_axis = [i[1] for i in trunc_occurences]
x_axis = [k for k,i in enumerate(trunc_occurences)]
x_label = [i[0] for i in trunc_occurences]
plt.xticks(rotation=85, fontsize = 15)
plt.yticks(fontsize = 15)
plt.xticks(x_axis, x_label)
plt.ylabel("Number of Occurences", fontsize = 24, labelpad = 0)
ax2.bar(x_axis, y_axis, align = 'center', color = 'purple')
plt.title("Popularity of Genres",fontsize = 30)
plt.savefig(base_path + '/figures/plt/' + 'Bar graph popularity of genres.jpg',bbox_inches='tight')
plt.show()
# + [markdown] id="FzJMeCRfVLfj"
# The bar graph shows the number of occurences for each genre(genre was edited because there were too many genres). Some movies had multiple genres (ex. romantic comedy woud count as romance and comedy) We can see that comedy and drama are the most present genres.
# + colab={"base_uri": "https://localhost:8080/", "height": 358} id="mZXagWG-V8Zt" executionInfo={"status": "ok", "timestamp": 1617673679907, "user_tz": 240, "elapsed": 1085, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}} outputId="4c34b77d-d566-4d6a-84c7-088b8ed71c31"
#Wordcloud visualization of popularity of genres
# Function that control the color of the words
def random_color_func(word=None, font_size=None, position=None,
orientation=None, font_path=None, random_state=None):
h = int(360.0 * tone / 255.0)
s = int(100.0 * 255.0 / 255.0)
l = int(100.0 * float(random_state.randint(70, 120)) / 255.0)
return "hsl({}, {}%, {}%)".format(h, s, l)
#Finally, the result is shown as a wordcloud:
words = dict()
trunc_occurences = keyword_occurences[0:50]
for s in trunc_occurences:
words[s[0]] = s[1]
tone = 100 # define the color of the words
f, ax = plt.subplots(figsize=(14, 6))
wordcloud = WordCloud(width=550,height=300, background_color='black',
max_words=1628,relative_scaling=0.7,
color_func = random_color_func,
normalize_plurals=False)
wordcloud.generate_from_frequencies(words)
plt.imshow(wordcloud, interpolation="bilinear")
plt.axis('off')
plt.savefig(base_path + '/figures/plt/' + 'Wordcloud visualization of popularity of genres.jpg',bbox_inches='tight')
plt.show()
# + [markdown] id="f33zIy0VWlmg"
# This Wordcloud visualization of popularity of genres shows the same things as the bar grah above which is the number of occurences for each genre(genre was edited because there were too many genres). Some movies had multiple genres (ex. romantic comedy woud count as romance and comedy) Again, we can see that comedy and drama are the most present genres.
# + [markdown] id="jh9dMS4pC6WD"
# **RATINGS**
# + colab={"base_uri": "https://localhost:8080/", "height": 380} id="4kJSeb4TtIpR" executionInfo={"status": "ok", "timestamp": 1617673581795, "user_tz": 240, "elapsed": 425, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}} outputId="594a59f5-2a05-4b0f-c306-a81548410407"
ratings.describe()
# + colab={"base_uri": "https://localhost:8080/", "height": 306} id="iUpwjyuAtXcB" executionInfo={"status": "ok", "timestamp": 1617673584194, "user_tz": 240, "elapsed": 806, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}} outputId="9cca842a-f94c-47a0-f28c-d0eb20011ac5"
# plot a bar chart of ratings count
ratings.value_counts(ratings['rating']).sort_index().plot.bar(color=["orange"])
plt.title('Distribution of Movie Ratings', fontsize=20)
plt.ylabel('count', fontsize=16)
plt.xlabel(ratings['rating'].name, fontsize=16)
plt.xticks(rotation=0, ha='right')
plt.savefig(base_path + '/figures/plt/' + 'Bar chart ratings count.jpg',bbox_inches='tight')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 306} id="HdMD_7y1Tm9n" executionInfo={"status": "ok", "timestamp": 1617673585755, "user_tz": 240, "elapsed": 561, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}} outputId="af9ecfa8-936f-4dc7-d52a-1b519226f8d4"
# plot a bar chart of ratings count
ratings.value_counts(ratings['rating'], normalize=True).sort_index().plot.bar(color=["yellow"])
plt.title('Distribution of Movie Ratings', fontsize=20)
plt.ylabel('proportion', fontsize=16)
plt.xlabel(ratings['rating'].name, fontsize=16)
plt.xticks(rotation=0, ha='right')
plt.savefig(base_path + '/figures/plt/' + 'Bar chart ratings proportion.jpg',bbox_inches='tight')
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="jZdMzOtdUK7F" executionInfo={"status": "ok", "timestamp": 1617673587359, "user_tz": 240, "elapsed": 200, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}} outputId="d4d22f81-d0b8-47c2-9329-9e2cf24e17fe"
# get exact class proportions
ratings.value_counts(ratings['rating'], normalize=True)
# + colab={"base_uri": "https://localhost:8080/"} id="glUzMlCoInZj" executionInfo={"status": "ok", "timestamp": 1617673588829, "user_tz": 240, "elapsed": 229, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}} outputId="4ec8f208-31c1-4f1d-dd0c-438733fde512"
# mean ratings
np.mean(ratings['rating'])
# + [markdown] id="DCFa8mGoGxSB"
# The bar plot shows the distribution of movie ratings. The mode (i.e. most common) rate is 4, and the distribution is left-skewed. The mean rating is 3.58.
#
# In terms of class balance, 34.9% of the ratings are 4s, 26.1% are 3s, 22.6% are 5s, 10.7% are 2s, and 5.6% are 1s.
# + colab={"base_uri": "https://localhost:8080/", "height": 480} id="8wlNtO5xNmPP" executionInfo={"status": "ok", "timestamp": 1617673590758, "user_tz": 240, "elapsed": 861, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}} outputId="8624adb8-b3a8-459e-b62a-888fc3ed8cba"
#Visualize the User Rating of the Movie “Toy Story”
movies_ratings[movies_ratings.title == 'Toy Story (1995)'].groupby('rating')['user_id'].count().plot(kind = 'bar', color = 'green',figsize = (8,7))
plt.xlabel('Rating')
plt.ylabel('Number of Users')
plt.title('Bar Chart of User Rating of Toy Story')
plt.xticks(rotation=0, ha='right')
plt.savefig(base_path + '/figures/plt/' + 'Bar chart User Rating of the Movie “Toy Story”.jpg',bbox_inches='tight')
plt.show
# + [markdown] id="rT_MkJ7IONVM"
# The bar plot shows the distribution of movie ratings for Toy Story by user. Clearly, we can see that Toy Story has received very good reviews give that it has mostly 4 and 5 stars
# + [markdown] id="DBruhJpHDDPr"
# **USERS**
# + [markdown] id="8LhFWX_NDuuy"
# **Gender**
# + colab={"base_uri": "https://localhost:8080/"} id="x55az-WzvZwk" executionInfo={"status": "ok", "timestamp": 1617673592754, "user_tz": 240, "elapsed": 230, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}} outputId="e3403a01-9f84-48d2-d0d3-61ee3655f362"
# get gender count
users['gender'].value_counts()
# + colab={"base_uri": "https://localhost:8080/", "height": 248} id="dBL2GW_xybUw" executionInfo={"status": "ok", "timestamp": 1617673594039, "user_tz": 240, "elapsed": 603, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}} outputId="fa66f138-c6a6-4e5d-b76f-c86eddfdc3cc"
# plot a pie chart of gender distribution
gender_data = users.value_counts(users['gender'])
plt.pie(gender_data, labels = ["male", "female"], autopct='%.2f', colors=["lightblue", "pink"], textprops={'fontsize': 14})
plt.savefig(base_path + '/figures/plt/' + 'Pie chart of gender distribution.jpg',bbox_inches='tight')
plt.show()
# + [markdown] id="cD-sagc_GItG"
# From the pie chart, 71.7% of the users are male, and 28.3% are female. Thus, the data is imbalanced towards male users.
# + [markdown] id="fASSkNASD2Ax"
# **Age range**
# + colab={"base_uri": "https://localhost:8080/", "height": 284} id="Ho7qHWph0ern" executionInfo={"status": "ok", "timestamp": 1617673596105, "user_tz": 240, "elapsed": 975, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}} outputId="d3826580-48f1-48e0-b57c-5ceeb6c2e5f8"
# replace below 18 values with 0-18, for plot order
users['age_desc'] = users['age_desc'].replace('Under 18', '0-18')
users.head()
# plot a bar chart of user age ranges
users.value_counts(users['age_desc']).sort_index().plot.bar(color=["orange"])
plt.ylabel('count', fontsize=16)
plt.xlabel('user age range', fontsize=16)
plt.xticks(rotation=0, ha='right')
plt.savefig(base_path + '/figures/plt/' + 'Bar chart of user age ranges.jpg',bbox_inches='tight')
plt.show()
# + [markdown] id="0aVo8H9-E902"
# From the bar plot of user age ranges, the age distribution has its mode at 25-34, and appears to have a right skew. Overall, most users appear to be young adults between 18-44.
# + colab={"base_uri": "https://localhost:8080/"} id="p-nqnidQ8PCo" executionInfo={"status": "ok", "timestamp": 1617673597360, "user_tz": 240, "elapsed": 237, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}} outputId="82b68d0d-c288-4182-8d2f-1bc6353f6583"
users.value_counts(users['age_desc'])
# + id="FGvdO3Uv0lVM" executionInfo={"status": "ok", "timestamp": 1617673599296, "user_tz": 240, "elapsed": 253, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYl_tqOFZ5W2cE-YU8KG1J1TwyYbIywmPedKYmNQ=s64", "userId": "14394843052745211035"}}
age_ranges = users['age_desc'].unique()
# + [markdown] id="ZJdHIwFaD_oJ"
# **Occupation**
# + colab={"base_uri": "https://localhost:8080/", "height": 284} id="PD7bwacf1hs9" executionInfo={"status": "ok", "timestamp": 1617673601139, "user_tz": 240, "elapsed": 981, "user": {"displayName": "<NAME>", "photoUrl": "<KEY>", "userId": "14394843052745211035"}} outputId="5f86dd5c-f0f8-42df-c850-41a39a17061a"
# get list of occupations
users.value_counts(users['occ_desc']).nunique()
# plot a sorted horizontal bar chart of user occupations
users.value_counts(users['occ_desc']).sort_values(ascending=True).plot.barh(color=["orange"])
plt.ylabel('count', fontsize=16)
plt.xlabel('user occupations', fontsize=16)
plt.xticks(rotation=0, ha='right')
plt.savefig(base_path + '/figures/plt/' + 'Sorted horizontal bar chart of user occupations.jpg',bbox_inches='tight')
plt.show()
# + [markdown] id="AQpOLXqu96DD"
# There are 21 different occupations in the dataset. The most common occupation in the Movie Lens user data is college/grad student, followed by 'other, and then 'executive managerial'. The top 5 most common occuaptions also include academic/educator, and technician engineering. Thus, users appear to be relatively highly educated.
| src/clean/blog post 1/EDA_Blog_Post_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from bs4 import BeautifulSoup
import numpy as np
import requests
import re
import itertools
import random
import pandas as pd
np.random.seed(42)
url="https://www.bindingdb.org/validation_sets/index-1.jsp"
r = requests.get(url)
soup = BeautifulSoup(r.content,'html.parser')
properstruct = re.compile('http://www.rcsb.org/pdb/explore/explore.do')
rows = soup.find_all('tr')
imp_rows = [i for i in rows if len(i.find_all('a')) > 0]
rec_to_pdb = dict()
list_of_pdbs=[]
system = ""
list_rec = []
for idx, i in enumerate(imp_rows):
name = i.span
if name is not None:
if len(list_rec):
rec_to_pdb[system]=list_rec
list_rec = []
system = name.text
continue
#non-system names should make it here
if 'bold' in i.attrs['class']:
pdbid = [j.text for j in i.find_all('a',href=properstruct)]
list_rec+= pdbid
list_of_pdbs+= pdbid
if len(list_rec):
rec_to_pdb[system]=list_rec
groups = [*rec_to_pdb]
train = round(0.8*len(groups))
val=round(1.0/3*len(groups))
np.random.shuffle(groups)
train_data = pd.read_csv('training_input.txt', delimiter=' ', header =None)
train_data.columns = ['label','reglabel','og', 'lig1','lig2']
train_data['recs'] = train_data['og'].astype(str).str[:4]
train_data.head()
#train_data['rec'] = pd.Series(train_data.apply(lambda x: re.match(x,rec_of_row)[0],axis=1), index=train_data.index)
pdb_to_ind = dict()
for pdb in list_of_pdbs:
pdb_to_ind[pdb]=train_data.index[train_data['recs'] == pdb].tolist()
del train_data['recs']
# +
perm1_train=groups[:train]
remainder=groups[train:]
perm1_val = remainder[:len(remainder)//2]
perm1_test= remainder[len(remainder)//2:]
print(len(perm1_train),len(perm1_val),len(perm1_test))
def make_csvs(train_data, groups, name, rec_to_pdb, pdb_to_ind):
csv_out=pd.DataFrame()
for val in groups:
rec =rec_to_pdb[val]
for r in rec:
all_rec = train_data.loc[pdb_to_ind[r]]
csv_out = csv_out.append(all_rec)
csv_out.to_csv(name+'.txt',sep=' ',header=False,index=False)
return csv_out.shape[0]
double_check = 0
for grouping,name in zip([perm1_train,perm1_val,perm1_test],['train','val','test']):
val = make_csvs(train_data, grouping,name,rec_to_pdb, pdb_to_ind)
print(float(val)/train_data.shape[0])
double_check += val
# -
print(double_check)
print(train_data.shape[0])
97+42
'text'.split()
| Train_Test_CV_Splitter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# ## Datos generales
# +
from sklearn.preprocessing import MinMaxScaler
from sklearn.model_selection import RepeatedStratifiedKFold
from sklearn.naive_bayes import GaussianNB
from project.feature_selection.feature_selector import FSPipeline
from project.feature_selection.feature_selector import FSPipelineEvaluator
from project.feature_selection.feature_selector import FeatureSelection
metric = "gmean"
norm = ("minmax", MinMaxScaler()) # probar con standard scaler
estimator = ("naiveBayes", GaussianNB())
cv = RepeatedStratifiedKFold(5, n_repeats=30, random_state=None)
max_n_features = 100
corpus_id = "ma_100"
# -
from project.utils import read_datasets
datasets = read_datasets("ma")
# ## Pipelines de métodos de selección
# +
# Fisher
fisher_pipeline = FSPipeline([
norm,
("fs", FeatureSelection("fisher", max_n_features)),
estimator,
])
ev_fs = FSPipelineEvaluator([fisher_pipeline,
],
cv,
metric,
max_n_features)
# -
# %time res = ev_fs._run(corpus_id, datasets, n_jobs=None, reset=True)
res.results['dataset_prostate_singh'].plot()
# .## res.save()
# +
# ReliefF
reliefF_pipeline = FSPipeline([
norm,
("fs", FeatureSelection("reliefF", max_n_features)),
estimator,
])
# Chi2
chi2_pipeline = FSPipeline([
norm,
("fs", FeatureSelection("chi_square", max_n_features)),
estimator,
])
# Random Forest
random_forest_pipeline = FSPipeline([
norm,
("fs", FeatureSelection("random_forest", max_n_features)),
estimator,
])
# -
from project.utils import read_datasets
datasets = read_datasets("ma")
| Feature-Selection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:tflearn]
# language: python
# name: conda-env-tflearn-py
# ---
# +
#https://github.com/kyamaz/openql-notes/blob/master/docs/20180219/sympy_programing_1_handout.ipynb
#0. [準備]必要なライブラリをインポートする
from sympy import *
from sympy.physics.quantum import *
from sympy.physics.quantum.qubit import Qubit,QubitBra
from sympy.physics.quantum.gate import X,Y,Z,H,S,T,CNOT,SWAP, CPHASE
from sympy.physics.quantum.gate import IdentityGate as _I
# +
#↓SymPy で良い感じに表示するためのおまじない
from sympy.printing.dot import dotprint
init_printing()
# +
#1. 計算に必要な量子ビット(量子レジスタ)を準備して、その値を初期化する
#計算に必要な量子ビットは、sympy.physics.quantum.qubit.Qubitクラス(実態はケット・ベクトル)を必要なビット数を
#初期化して作成します。
#例1:変数 $a$ に、2量子ビット $ \vert 00 \rangle $ を準備します。
a = Qubit('00')
print(a)
a
# +
#例3:8量子ビットをすべて $0$ として準備します。
q_8 = Qubit('0'*8)
pprint(q_8)
q_8
# -
#例4:(後述で説明する計算方法を使いますが)計算を行って、変数 $c$ の状態ベクトルを準備します。
#2量子ビットの重ね合わせ
c = qapply(H(1)*H(0)*a)
print(c)
c
# +
#Qubitクラスには次のような関数やプロパティが定義されています。
q = Qubit('000')
# サイズ(量子ビット数)を返す
print("nqubits=%d"% q.nqubits)
print("len=%d" % len(q))
print("dimension=%d" % q.dimension)
print(q.qubit_values)
q.flip(1) # 引数で指定された位置(*)の右から2番目の量子ビットを反転します.
# -
#2. 量子計算をユニタリ行列(ゲート演算子)で記述する
#作用させるユニタリ行列を標準で準備されているユニタリ行列のテンソル積で表します。
#
#まず最初に、標準で定義されているユニタリ行列を見て見ましょう。
#
#1量子ビット操作
#引数に渡す数字は、作用させる量子ビットの位置を表します。
#
#パウリ演算子
X(0).get_target_matrix() # σ_x, get_target_matrix() で行列表現を取り出します
Y(0).get_target_matrix() # σ_y
Z(0).get_target_matrix() # σ_z
H(0).get_target_matrix()
S(0).get_target_matrix()
represent(S(0), nqubits=1) # represent は、行列を nqubits で表現します。
Dagger(S(0))
Dagger(S(0).get_target_matrix()) # get_target_matrix() はDaggerカッコの中
represent(Dagger(S(0)), nqubits=1)
def Sdg(n): return S(n)**(-1) # S^{\dagger} としては、この演算子を使います。
represent(Sdg(0),nqubits=1)
T(0)
represent(T(0),nqubits=1)
Dagger(T(0))
represent(Dagger(T(0)),nqubits=1)
Dagger(T().get_target_matrix()) # get_target_matrix() はDaggerカッコの中
def Tdg(n): return T(n)**(-1) # T^{\dagger} としては、この演算子を使います。
represent(Tdg(0),nqubits=1)
"""
2量子ビット操作
引数に渡す数字は2つ、作用させる量子ビットの位置を表します。省略すると 0 のみとなるため、そのあとの計算がうまくできません。
必ず、引数を指定するようにしましょう。
CNOT操作
次の $ 4 \times 4 $ の行列です。 get_target_matrix() では表示できません。represent() を使います。
$ \left( \begin{array}{cccc} 1 & 0 & 0 & 0 \\ 0 & 1 & 0 & 0 \\ 0 & 0 & 0 & 1 \\ 0 & 0 & 1 & 0 \end{array} \right) $
"""
CX=CNOT(1,0)
print("controls=%s,targets=%s,gate=%s" % (CX.controls, CX.targets, CX.gate)) # CNOTのプロパティ
# CX.get_target_matrix() は、XGate になってしまいます。
# pprint(CX.get_target_matrix())
represent(CX,nqubits=2) # 行列形式で直接表現する方法
"""
SWAP操作
指定された量子ビットを入れ替える操作です。
"""
pprint(SWAP(0,1).get_target_matrix())
represent(SWAP(0,1),nqubits=2) # 行列形式で直接表現する方法
# +
"""
CPHASE操作
制御Zゲート、Control-Z、CZ と呼ばれ、次の行列です。
$ \left( \begin{array}{cccc} 1 & 0 & 0 & 0 \\ 0 & 1 & 0 & 0 \\ 0 & 0 & 1 & 0 \\ 0 & 0 & 0 & -1 \end{array} \right) $
"""
CPHASE(0,1)
# -
represent(CPHASE(0,1),nqubits=2)
Op1 = X(1)*_I(0)
pprint(TensorProduct(X().get_target_matrix(),_I().get_target_matrix())) # Op1 を行列形式で直接プリントできる方法があるかも?なかったら作ります
Op1
# 標準の1量子ビット操作のユニタリ演算は、その掛け算の順番よりも、指定した量子ビット(引数)がポイントとなります。
# 順番が大切になるのは、同じ位置の量子ビットに作用する場合で、積の交換はできません。
Op1_ = _I(0)*X(1)
Op1_
Op2 = _I(1)*X(0)
pprint(TensorProduct(_I().get_target_matrix(),X().get_target_matrix())) # Op2 を行列形式で直接プリントできる方法があるかも?なかったら作ります
Op2
# # !pip install graphviz
import graphviz
from sympy.printing.dot import dotprint
from graphviz import Source
Source(dotprint(Op1)) # Jupyter NotebookやQtConsoleでは、ツリー構造が表示できます。
Source(dotprint(Op2)) # 内部的な構造のため、通常の量子計算では不要です。
# +
def hadamard(s,n):
h = H(s)
for i in range(s+1,n+s):
h = H(i)*h
return h
h_8 = hadamard(0,8)
print(h_8)
print("\n")
pprint(h_8) # = pretty_print(h_8)
# h8.get_target_matrix() は実装がなく、エラーになります
h_8
# -
h_4 = hadamard(0,4)
represent(h_4,nqubits=4)
#量子回路図
#ユニタリ行列のテンソル積は、量子回路として、量子回路図で表現できます。
#SymPy には、量子回路図を描画するための仕組みが備わっています。量子回路図を描いてみましょう。
# そのために、まずは、必要なライブラリを読み込んで準備します。
# %matplotlib inline
import matplotlib.pyplot as plt
from sympy.physics.quantum.circuitplot import CircuitPlot,labeller, Mz,CreateOneQubitGate
#CNOTゲートを描いてみましょう。
CircuitPlot(CNOT(1,0), 2, labels=labeller(2,'\phi')[::-1]) # labellerでつくられる添え字の上下が逆?
#(ダガー)付きのゲート演算子など、ゲートの自作もできます。
Sdag = CreateOneQubitGate('Sdg','S^{\dagger}')
Tdag = CreateOneQubitGate('Tdg','T^{\dagger}')
CircuitPlot(Sdag(0)*Tdag(0),nqubits=1)
CircuitPlot(CNOT(1,0)*S(1), 2, labels=labeller(2)[::-1]) # 操作の順番に注意してください。
CircuitPlot(S(1)*H(0)*CNOT(1,0)*S(1)*H(1), 2, labels=labeller(2)[::-1])
#(参考)Z基底での測定を描くときは、Mz を使って表現します。
CircuitPlot(Mz(0)*Mz(1)*S(1)*H(0)*CNOT(1,0)*S(1)*H(1), 2, labels=labeller(2)[::-1])
# +
#3. ユニタリ行列を量子ビットに作用する
#ここまでのユニタリ行列の準備では、実際の行列計算は行われておりません。
#SymPy の特徴である「変数を変数のまま」扱ってきました。変数は単なる記号でしかありませんでした。
#実際に変数を計算するのは、 qapply() という関数で行います。計算したい記述を qapply 関数の引数に指定して計算します。
#直前で回路図を描いた計算をしてみましょう。
qapply(S(1)*H(0)*CNOT(1,0)*S(1)*H(1)*Qubit('00'))
# -
hadamard_8=qapply(h_8*q_8)
hadamard_8
"""
4. 測定する
ここまでの量子計算(qapply)では、量子状態のまま計算されます。
シミュレーション上の理論的な計算では、qapplyまで行えば、目的が達成することが多いです。
SymPy には、観測のためのメソッドも備わっています。
measure_all()では、引数で渡された量子ビットのすべての状態の測定確率を計算で求めて出力します。
そのほかにも幾つか観測のためのメソッドが提供されます。例えば、measure_partial()は、第一引数で指定された量子ビットに対して、第二引数で指定された部分的な振幅の測定を実行します。
"""
from sympy.physics.quantum.qubit import measure_all, measure_partial, measure_all_oneshot, measure_partial_oneshot
measure_all(hadamard_8)
for i in range(8):
pprint(measure_all_oneshot(hadamard_8))
#(例題1)Toffoli(CCX), CCCX, CCCCX ...
from sympy.physics.quantum.gate import CGateS
def CCX(c1,c2,t): return CGateS((c1,c2),X(t))
def Toffoli(c1,c2,t): return CGateS((c1,c2),X(t))
CircuitPlot(CCX(1,2,0),3,labels=labeller(3)[::-1])
represent(CCX(1,2,0),nqubits=3)
def CCCX(c1,c2,c3,t): return CGateS((c1,c2,c3),X(t))
CircuitPlot(CCCX(1,2,3,0),4,labels=labeller(4)[::-1])
def CCCCX(c1,c2,c3,c4,t): return CGateS((c1,c2,c3,c4),X(t))
CircuitPlot(CCCCX(1,2,3,4,0),5,labels=labeller(5)[::-1])
# +
"""
(例題2)重ね合わせ状態の中の、ある状態のみマーキングする
重ね合わせ状態の中の、ある状態 $ s_t $ のみに、−1掛ける操作(ここでは、マーキングを呼びます)を考えます。
たとえば、2量子ビットの重ね合わせ状態 $\displaystyle \frac{1}{2} \left(\ |\ 00\ \rangle\ +\ |\ 01\ \rangle\ +\ |\ 10\ \rangle\ +\ |\ 11\ \rangle\ \right) $ を考えるとき、
状態 $\;|\ 10\ \rangle\;$ をマーキングするとします。
マーキング後の状態は、$\displaystyle \frac{1}{2} \left(\ |\ 00\ \rangle\ +\ |\ 01\ \rangle\ -\ |\ 10\ \rangle\ +\ |\ 11\ \rangle\ \right) $ となることを目指します。
3量子ビットのマーキング操作を試してみましょう
"""
h_3 = hadamard(0,3)
target_state_3 = qapply(h_3*Qubit('000')) #3量子ビットの重ね合わせ状態を準備します。
def CCZ(c1,c2,t): return (H(t)*CCX(c1,c2,t)*H(t)) # CCZ演算子を定義します。
# -
mark_7 = CCZ(1,2,0)
qapply(mark_7*target_state_3)
mark_6 = X(0)*CCZ(1,2,0)*X(0)
qapply(mark_6*target_state_3)
mark_5 = X(1)*CCZ(1,2,0)*X(1)
qapply(mark_5*target_state_3)
mark_4 = X(1)*X(0)*CCZ(1,2,0)*X(1)*X(0)
qapply(mark_4*target_state_3)
mark_3 = X(2)*CCZ(1,2,0)*X(2)
qapply(mark_3*target_state_3)
mark_2 = X(2)*X(0)*CCZ(1,2,0)*X(2)*X(0)
qapply(mark_2*target_state_3)
mark_1 = X(2)*X(1)*CCZ(1,2,0)*X(2)*X(1)
qapply(mark_1*target_state_3)
mark_0 = X(2)*X(1)*X(0)*CCZ(1,2,0)*X(2)*X(1)*X(0)
qapply(mark_0*h_3*Qubit('000'))
# +
"""
(例題3)重ね合わせ状態の中に、マーキングした状態があるかを見る
著名な操作「Grover のアルゴリズム」を試してみましょう。
(ヒント1)平均値周りの反転操作:$\displaystyle D_{n} = H_{n} \cdot \Big( 2\ |\ 0\ \rangle\langle\ 0\ |_{n}\ -\ I_{n} \Big) \cdot H_{n} $ を使います。
(ヒント2)試行回数は、$\displaystyle \mathcal{O}(\sqrt{n}) $
"""
# d_3 = h_3 * X(0)*X(1)*X(2) * H(0)*CCX(1,2,0)*H(0) * X(0)*X(1)*X(2) * h_3 # グローバル位相(絶対位相)の差に注意
# d_3 = h_3 * X(0)*X(1)*X(2) * CGateS((1,2), Z(0)) * X(0)*X(1)*X(2) * h_3
def DOp(n): return (Qubit('0'*n)*QubitBra('0'*n)*2-_I(0)) # ゲート操作で計算するには、上記コメントのような演算になります。
d_3 = h_3 * DOp(3) * h_3 # 平均値周りの反転操作
represent(d_3,nqubits=3)
# +
#3量子ビットで状態|7>を探す
ret1=qapply(d_3*mark_7*target_state_3)
measure_all(ret1)
# -
ret2=qapply(d_3*mark_7*ret1) # シミュレーター計算ですから、上で一度観測した ret1 を使っても観測の影響なく次の計算に利用可能。
measure_all(ret2)
# +
over_1=qapply(d_3*mark_7*ret2)# 試行回数が多いとどうなるでしょう。# 試行回数が多
measure_all(over_1)
# -
over_2=qapply(d_3*mark_7*over_1) # 試行回数が多いとどうなるでしょう。# 試行回数が多
measure_all(over_2)
#遠しで計算すると、回路が長くなってきています。少し時間かかります。
from sympy.physics.quantum.gate import gate_simp
search_7_in_3qubit = gate_simp(d_3*mark_7*d_3*mark_7)
ret3=qapply(search_7_in_3qubit*target_state_3)
ret3
print(measure_all(ret3))
for i in range(10):
pprint(measure_all_oneshot(ret3))
d_3_gate = h_3 * X(0)*X(1)*X(2) * H(0)*CCX(1,2,0)*H(0) * X(0)*X(1)*X(2) * h_3
CircuitPlot(gate_simp(d_3_gate*mark_7*d_3_gate*mark_7*h_3),nqubits=3,labels=labeller(3)[::-1])
"""
OpenQL 発のPythonライブラリ QuantPy を紹介します
QuantPy は、フロントエンドを SymPy を使っています。
「量子ビットの準備」「量子操作(ゲート演算子)の記述」は SymPy で行います。
「量子ビットにゲート演算子を作用」する qapply() を置き換えて動作します。
"""
from quantpy.sympy.qapply import qapply
# sympy.physics.quantum.qapply を上書きします。sympy.physics.quantum のインポートの後に読み込んでください。
from quantpy.sympy.executor.classical_simulation_executor import ClassicalSimulationExecutor
classical_simulator = ClassicalSimulationExecutor()
for i in range(8):
print(qapply(H(2)*H(1)*H(0)*Qubit('000'), executor=classical_simulator))
from quantpy.sympy.executor.ibmq_executor import IBMQExecutor
# APItoken = '<PASSWORD>'
# ibmq_simulator = IBMQExecutor(api_key=APItoken, backend='ibmqx2')
ibmq_simulator = IBMQExecutor(backend='local_qasm_simulator')
qpy_ret=qapply(H(2)*H(1)*H(0)*Qubit('000'), executor=ibmq_simulator)
print(qpy_ret)
from quantpy.sympy.expr_extension import *
sympy_expr_add_operators()
circuit = X(0)>>X(1)>>X(2)
circuit
#(Homework)Toffoli 再び
#課題1) Toffoli ゲートを、基本的な量子ゲートだけで表してください。
def ToffoliGate_plot(q0,q1,q2):
return T(q0)*S(q1)*CNOT(q0,q1)*Tdag(q1)*CNOT(q0,q1)\
*H(q2)*Tdag(q1)*T(q2)*CNOT(q0,q2)*Tdag(q2)*CNOT(q1,q2)\
*T(q2)*CNOT(q0,q2)*Tdag(q2)*CNOT(q1,q2)*H(q2)
CircuitPlot(ToffoliGate_plot(2,1,0), nqubits=3)
def Tdg(q): return (T(q)**(-1))
def ToffoliGate(q0,q1,q2):
return T(q0)*S(q1)*CNOT(q0,q1)*Tdg(q1)*CNOT(q0,q1)\
*H(q2)*Tdg(q1)*T(q2)*CNOT(q0,q2)*Tdg(q2)*CNOT(q1,q2)\
*T(q2)*CNOT(q0,q2)*Tdg(q2)*CNOT(q1,q2)*H(q2)
ToffoliGate(2,1,0)
# qapply(ToffoliGate(2,1,0)*Qubit('110'), dagger=True) # 残念ながら、計算ができませんでした。
def CCCX_with_ancilla(q0,q1,q2,t,a0,a1,a2):
return CNOT(q0,a0)*Toffoli(q1,a0,a1)*Toffoli(q2,a1,a2)*CNOT(a2,t)*Toffoli(q2,a1,a2)*Toffoli(q1,a0,a1)*CNOT(q0,a0)
CircuitPlot(CCCX_with_ancilla(6,5,4,3,2,1,0),nqubits=7,labels=(labeller(3,'a')+['t']+labeller(3,)))
qapply(CCCX_with_ancilla(6,5,4,3,2,1,0)*Qubit('1111000'))
qapply(CCCX_with_ancilla(6,5,4,3,2,1,0)*Qubit('1110000'))
# +
"""
付録A:ブラ・ベクトル
複素共役を表すための dagger( $ \dagger $ ) をQubitオプジェクトに作用させると、ブラ・ベクトルになります。
また、内積も計算できます。
シンボルのままで内積にするには、単にブラ・ベクトル QubitBra とケット・ベクトル Qubit を掛ける(*)だけです。
実際の値を計算するには、この内積オブジェクトのdoit()メソッドで計算します。
"""
from sympy.physics.quantum.dagger import Dagger
Dagger(q)
# -
type(Dagger(q))
ip = Dagger(q)*q
ip
ip.doit()
# +
"""
付録B:量子フーリエ変換
"""
from sympy.physics.quantum.qft import QFT
fourier_3 = QFT(0,3).decompose()
fourier_3
# -
figure_qft3 = CircuitPlot(fourier_3, nqubits=3)
qapply(fourier_3 * Qubit('000'))
# +
"""
付録C:QASM
SymPy独自のQASMでゲートを記述できます。
ただし、OpenQASM とは異なるフォーマットです。注意してください。
"""
from sympy.physics.quantum.qasm import Qasm
qprog1 = Qasm(
'qubit q0',
'qubit q1',
'h q0',
'cnot q0,q1')
CircuitPlot(qprog1.get_circuit(),2,labels=labeller(2))
# -
qprog2 = Qasm("def c-S,1,'S'",
"def c-T,1,'T'",
"qubit j_0",
"qubit j_1",
"qubit j_2",
"h j_0",
"c-S j_1,j_0",
"c-T j_2,j_0",
"nop j_1",
"h j_1",
"c-S j_2,j_1",
"h j_2",
"swap j_0,j_2")
qprog2.plot()
qapply(qprog2.get_circuit()*Qubit('000'))
# +
qasm_lines = """\
def CU,1,'U'
def CU2,1,'U^2'
def CU4,1,'U^4'
def c-S,1,'S'
def c-T,1,'T'
qubit j_0,0 # QFT qubits
qubit j_1,0
qubit j_2,0
qubit s_0 # U qubits
h j_0 # equal superposition
h j_1
h j_2
CU4 j_0,s_0 # controlled-U
CU2 j_1,s_0
CU j_2,s_0
h j_0 # QFT
c-S j_0,j_1
h j_1
nop j_0
c-T j_0,j_2
c-S j_1,j_2
h j_2
nop j_0
nop j_0
nop j_1
measure j_0 # final measurement
measure j_1
measure j_2"""
qprog3 = Qasm(*qasm_lines.splitlines())
qprog3.plot()
# -
| docs/20180219/sympy_programing_1_handout-mac-20180820.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Library design for CTP-05, Intronic RNA
#
#
# by <NAME>
#
# This library design is based on target regions designed by Stephen
# ## 0. Imports
# +
# %run "E:\Users\puzheng\Documents\Startup_py3.py"
sys.path.append(r"E:\Users\puzheng\Documents")
import ImageAnalysis3
from ImageAnalysis3 import get_img_info, visual_tools, corrections, library_tools
from ImageAnalysis3.library_tools import LibraryDesigner as ld
from ImageAnalysis3.library_tools import LibraryTools as lt
# %matplotlib notebook
print(os.getpid())
# -
# ## 1. Load probes and convert to dic
# +
# folder for genomic info
genome_folder = r'E:\Genomes\hg38'
# Library directories
pool_folder = r'X:\Libraries\CTP-07'
# folder for sub-pool
library_folder = os.path.join(pool_folder, 'chr21_intronic')
# folder for fasta sequences
sequence_folder = os.path.join(library_folder, 'sequences')
# folder to save result probes
save_folder = os.path.join(library_folder, 'reports')
# +
import csv
source_folder = r'E:\Users\puzheng\Documents\Libraries\CTP-05\Candidate_Probes'
probe_file = 'Chr21genes_96Intron.csv'
probes = []
with open(os.path.join(source_folder, probe_file),'r') as csvfile:
reader = csv.reader(csvfile, delimiter=',', quotechar='|')
header = next(reader,None)
for row in reader:
_source_probe = {h.lower():info for h,info in zip(header, row)}
_probe = {}
for _key,_value in _source_probe.items():
if 'position' in _key and 'position' not in _probe:
_probe['position'] = int(_value)
if 'gene' in _key and 'gene' not in _probe:
if '_' in _value:
_probe['gene'] = _value.split('_')[0]
else:
_probe['gene'] = _value
if 'sequence' in _key and 'sequence' not in _probe:
_probe['sequence'] = _value
probes.append(_probe)
# merge into dictionary
probe_dic = {}
for _probe in probes:
if _probe['gene'] not in probe_dic:
probe_dic[_probe['gene']] = [_probe]
else:
probe_dic[_probe['gene']].append(_probe)
position_file = "chr21_intron_positions.csv"
position_dic = {}
with open(os.path.join(source_folder, position_file),'r') as csvfile:
reader = csv.reader(csvfile, delimiter=',', quotechar='|')
for row in reader:
position_dic[row[0]] = int(row[1])
# sort dictionary and append probe id, position
for _gene, _pb_list in probe_dic.items():
_sorted_pb_list = sorted(_pb_list, key=lambda v:int(v['position']))
# extract position
_gene_start = position_dic[_gene]
_gene_end = _sorted_pb_list[-1]['position'] + len(_sorted_pb_list[-1]['sequence'])+_gene_start
for _i, _probe in enumerate(_sorted_pb_list):
_sorted_pb_list[_i]['id'] = _i
_sorted_pb_list[_i]['region'] = f'chr21:{_gene_start}-{_gene_end}'
print(len(probe_dic))
# +
#>chr21:26842909-26847909_gene_ADAMTS1_pb_0_pos_0_readouts_[NDB_1147_u,NDB_1147_u,NDB_1147_u]
probe_dic['APP']
# -
# ## 3. Screening genes and probes
# keep the most 5' probes and exclude genes without enough probes
min_num_probe = 12 # this number is from MERFISH experience as well as Long Cai 2018 paper
max_num_probe = 80 # this is just an arbitrary number larger than 2x min_num_probe
# initialize
kept_probe_dic = {}
for _gene, _pb_list in sorted(probe_dic.items(), key=lambda v:position_dic[v[0]]):
if len(_pb_list) < min_num_probe:
print(f"Gene: {_gene} has probes less than {min_num_probe}, skip")
else:
kept_probe_dic[_gene] = sorted(_pb_list, key=lambda p:int(p["position"]))[:min(len(_pb_list), max_num_probe)]
print("Number of genes kept:", len(kept_probe_dic))
# save
pickle.dump(kept_probe_dic, open(os.path.join(library_folder, 'filtered_candidate_dict.pkl'), 'wb'))
# ### save to fasta file
# +
# biopython for SeqRecord
from Bio import SeqIO
from Bio.Seq import Seq
from Bio.SeqRecord import SeqRecord
from Bio.Alphabet import IUPAC
# blast
from Bio.Blast.Applications import NcbiblastnCommandline
from Bio.Blast import NCBIXML
# -
# ## 4. Append barcode info
# design gene_readout_dict
gene_readout_dict = {_gene:['u'+str(i)]*3 for i,_gene in enumerate(kept_probe_dic)}
# load primers and readouts
reload(library_tools.probes)
primers = library_tools.probes.load_primers([4,3])
unique_readouts = library_tools.probes.load_readouts(len(gene_readout_dict), 'NDB_new', _num_colors=3, _start_id=29)
readout_dict = {'u':unique_readouts}
kept_probe_dic['APP']
# Assemble probes
reload(library_tools.probes)
cand_probes, readout_summary = library_tools.probes.Assemble_probes(library_folder, kept_probe_dic, gene_readout_dict,
readout_dict, primers, rc_targets=True, save=True)
# ## 5. Check quality
# +
# biopython for SeqRecord
from Bio import SeqIO
from Bio.Seq import Seq
from Bio.SeqRecord import SeqRecord
from Bio.Alphabet import IUPAC
# blast
from Bio.Blast.Applications import NcbiblastnCommandline
from Bio.Blast import NCBIXML
# +
# folder for this library
# candidate full-length probe filename
candidate_full_name = 'candidate_probes.fasta'
# load full probes
full_records = []
with open(os.path.join(library_folder, candidate_full_name), 'r') as handle:
for record in SeqIO.parse(handle, "fasta"):
full_records.append(record)
print(f"Total probe loaded: {len(full_records)}")
# +
import ImageAnalysis3.library_tools.quality_check as qc
reload(qc)
primer_check = qc._check_primer_usage(full_records, primers[0], primers[1])
print(f"primer_check:{primer_check}")
reg_size_dic, len_check = qc._check_region_size(full_records, min_size=15)
print(f"len_check:{len_check}")
reg_readout_dic, reg2readout_check = qc._check_region_to_readouts(full_records, readout_dict)
print(f"reg2readout_check:{reg2readout_check}")
readout_reg_dic, readout2reg_check = qc._check_readout_to_region(reg_readout_dic, full_records, readout_dict, target_len=30)
print(f"readout2reg_check:{readout2reg_check}")
int_map = qc._construct_internal_map(full_records, library_folder)
readout_count_dic, readout_count_check = qc._check_readout_in_probes(readout_reg_dic, reg_size_dic, int_map, readout_dict)
print(f"readout_count_check:{readout_count_check}")
kept_records, removed_count = qc._check_between_probes(full_records, int_map)
# save kept records
with open(os.path.join(library_folder, 'filtered_full_probes.fasta'), 'w') as output_handle:
SeqIO.write(kept_records, output_handle, "fasta")
# -
# ## blast
qc.Blast_probes(kept_records, library_folder)
kept_pbs, blast_keep_dic, hard_count_list, soft_count_list = qc.Screening_Probes_by_Blast(library_folder, 60,
hard_thres=29)
kept_pbs
# +
final_probe_folder = os.path.join(library_folder, 'final_probes')
if not os.path.exists(final_probe_folder):
os.makedirs(final_probe_folder)
reload(qc)
primer_check = qc._check_primer_usage(kept_pbs, primers[0], primers[1])
print(f"primer_check:{primer_check}")
reg_size_dic, len_check = qc._check_region_size(kept_pbs, min_size=15)
print(f"len_check:{len_check}")
reg_readout_dic, reg2readout_check = qc._check_region_to_readouts(kept_pbs, readout_dict)
print(f"reg2readout_check:{reg2readout_check}")
readout_reg_dic, readout2reg_check = qc._check_readout_to_region(reg_readout_dic, kept_pbs, readout_dict, target_len=30)
print(f"readout2reg_check:{readout2reg_check}")
int_map = qc._construct_internal_map(kept_pbs, library_folder)
readout_count_dic, readout_count_check = qc._check_readout_in_probes(readout_reg_dic, reg_size_dic, int_map, readout_dict)
print(f"readout_count_check:{readout_count_check}")
kept_records, removed_count = qc._check_between_probes(kept_pbs, int_map)
# save kept records
with open(os.path.join(final_probe_folder, 'extra_filtered_full_probes.fasta'), 'w') as output_handle:
SeqIO.write(kept_records, output_handle, "fasta")
# -
len(kept_records)
str(kept_records[0].seq)[40:70]
| Library_design/CTP-07/CTP-07_Chr21_intronic_RNA_newNDB.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import cython
import timeit
import math
# %load_ext cython
# # Native code compilation
#
# We will see how to convert Python code to native compiled code. We will use the example of calculating the pairwise distance between a set of vectors, a $O(n^2)$ operation.
#
# For native code compilation, it is usually preferable to use explicit for loops and minimize the use of `numpy` vectorization and broadcasting because
#
# - It makes it easier for the `numba` JIT to optimize
# - It is easier to "cythonize"
# - It is easier to port to C++
#
# However, use of vectors and matrices is fine especially if you will be porting to use a C++ library such as Eigen.
# ## Timing code
# ### Manual
# +
import time
def f(n=1):
start = time.time()
time.sleep(n)
elapsed = time.time() - start
return elapsed
# -
f(1)
# ### Clock time
# +
# %%time
time.sleep(1)
# -
# ### Using `timeit`
#
# The `-r` argument says how many runs to average over, and `-n` says how many times to run the function in a loop per run.
# %timeit time.sleep(0.01)
# %timeit -r3 time.sleep(0.01)
# %timeit -n10 time.sleep(0.01)
# %timeit -r3 -n10 time.sleep(0.01)
# ### Time unit conversions
#
# ```
# 1 s = 1,000 ms
# 1 ms = 1,000 µs
# 1 µs = 1,000 ns
# ```
# ## Profiling
#
# If you want to identify bottlenecks in a Python script, do the following:
#
# - First make sure that the script is modular - i.e. it consists mainly of function calls
# - Each function should be fairly small and only do one thing
# - Then run a profiler to identify the bottleneck function(s) and optimize them
#
# See the Python docs on [profiling Python code](https://docs.python.org/3/library/profile.html)
# Profiling can be done in a notebook with %prun, with the following readouts as column headers:
#
# - ncalls
# - for the number of calls,
# - tottime
# - for the total time spent in the given function (and excluding time made in calls to sub-functions),
# - percall
# - is the quotient of tottime divided by ncalls
# - cumtime
# - is the total time spent in this and all subfunctions (from invocation till exit). This figure is accurate even for recursive functions.
# - percall
# - is the quotient of cumtime divided by primitive calls
# - filename:lineno(function)
# - provides the respective data of each function
# +
def foo1(n):
return np.sum(np.square(np.arange(n)))
def foo2(n):
return sum(i*i for i in range(n))
def foo3(n):
[foo1(n) for i in range(10)]
foo2(n)
def foo4(n):
return [foo2(n) for i in range(100)]
def work(n):
foo1(n)
foo2(n)
foo3(n)
foo4(n)
# +
# %%time
work(int(1e5))
# -
# %prun -q -D work.prof work(int(1e5))
import pstats
p = pstats.Stats('work.prof')
p.print_stats()
pass
p.sort_stats('time', 'cumulative').print_stats('foo')
pass
p.sort_stats('ncalls').print_stats(5)
pass
# ## Optimizing a function
#
# Our example will be to optimize a function that calculates the pairwise distance between a set of vectors.
#
# We first use a built-in function from`scipy` to check that our answers are right and also to benchmark how our code compares in speed to an optimized compiled routine.
from scipy.spatial.distance import squareform, pdist
n = 100
p = 100
xs = np.random.random((n, p))
sol = squareform(pdist(xs))
# %timeit -r3 -n10 squareform(pdist(xs))
# ## Python
# ### Simple version
def pdist_py(xs):
"""Unvectorized Python."""
n, p = xs.shape
A = np.zeros((n, n))
for i in range(n):
for j in range(n):
for k in range(p):
A[i,j] += (xs[i, k] - xs[j, k])**2
A[i,j] = np.sqrt(A[i,j])
return A
# Note that we
#
# - first check that the output is **right**
# - then check how fast the code is
func = pdist_py
print(np.allclose(func(xs), sol))
# %timeit -r3 -n10 func(xs)
# ### Exploiting symmetry
def pdist_sym(xs):
"""Unvectorized Python."""
n, p = xs.shape
A = np.zeros((n, n))
for i in range(n):
for j in range(i+1, n):
for k in range(p):
A[i,j] += (xs[i, k] - xs[j, k])**2
A[i,j] = np.sqrt(A[i,j])
A += A.T
return A
func = pdist_sym
print(np.allclose(func(xs), sol))
# %timeit -r3 -n10 func(xs)
# ### Vectorizing inner loop
def pdist_vec(xs):
"""Vectorize inner loop."""
n, p = xs.shape
A = np.zeros((n, n))
for i in range(n):
for j in range(i+1, n):
A[i,j] = np.sqrt(np.sum((xs[i] - xs[j])**2))
A += A.T
return A
func = pdist_vec
print(np.allclose(func(xs), sol))
# %timeit -r3 -n10 func(xs)
# ### Broadcasting and vectorizing
#
# Note that the broadcast version does twice as much work as it does not exploit symmetry.
def pdist_numpy(xs):
"""Fully vectroized version."""
return np.sqrt(np.square(xs[:, None] - xs[None, :]).sum(axis=-1))
func = pdist_numpy
print(np.allclose(func(xs), sol))
# %timeit -r3 -n10 squareform(func(xs))
# ## JIT with `numba`
# We use the `numba.jit` decorator which will trigger generation and execution of compiled code when the function is first called.
from numba import jit
# ### Using `jit` as a function
pdist_numba_py = jit(pdist_py, nopython=True, cache=True)
func = pdist_numba_py
print(np.allclose(func(xs), sol))
# %timeit -r3 -n10 func(xs)
# ### Using `jit` as a decorator
@jit(nopython=True, cache=True)
def pdist_numba_py_1(xs):
"""Unvectorized Python."""
n, p = xs.shape
A = np.zeros((n, n))
for i in range(n):
for j in range(n):
for k in range(p):
A[i,j] += (xs[i, k] - xs[j, k])**2
A[i,j] = np.sqrt(A[i,j])
return A
func = pdist_numba_py_1
print(np.allclose(func(xs), sol))
# %timeit -r3 -n10 func(xs)
# ### Can we make the code faster?
#
# Note that in the inner loop, we are updating a matrix when we only need to update a scalar. Let's fix this.
@jit(nopython=True, cache=True)
def pdist_numba_py_2(xs):
"""Unvectorized Python."""
n, p = xs.shape
A = np.zeros((n, n))
for i in range(n):
for j in range(n):
d = 0.0
for k in range(p):
d += (xs[i, k] - xs[j, k])**2
A[i,j] = np.sqrt(d)
return A
func = pdist_numba_py_2
print(np.allclose(func(xs), sol))
# %timeit -r3 -n10 func(xs)
# ### Can we make the code even faster?
#
# We can also try to exploit symmetry.
@jit(nopython=True, cache=True)
def pdist_numba_py_sym(xs):
"""Unvectorized Python."""
n, p = xs.shape
A = np.zeros((n, n))
for i in range(n):
for j in range(i+1, n):
d = 0.0
for k in range(p):
d += (xs[i, k] - xs[j, k])**2
A[i,j] = np.sqrt(d)
A += A.T
return A
func = pdist_numba_py_sym
print(np.allclose(func(xs), sol))
# %timeit -r3 -n10 func(xs)
# ### Does `jit` work with vectorized code?
pdist_numba_vec = jit(pdist_vec, nopython=True, cache=True)
# %timeit -r3 -n10 pdist_vec(xs)
func = pdist_numba_vec
print(np.allclose(func(xs), sol))
# %timeit -r3 -n10 func(xs)
# ### Does `jit` work with broadcasting?
pdist_numba_numpy = jit(pdist_numpy, nopython=True, cache=True)
# %timeit -r3 -n10 pdist_numpy(xs)
func = pdist_numba_numpy
try:
print(np.allclose(func(xs), sol))
# %timeit -r3 -n10 func(xs)
except Exception as e:
print(e)
# #### We need to use `reshape` to broadcast
def pdist_numpy_(xs):
"""Fully vectroized version."""
return np.sqrt(np.square(xs.reshape(n,1,p) - xs.reshape(1,n,p)).sum(axis=-1))
pdist_numba_numpy_ = jit(pdist_numpy_, nopython=True, cache=True)
# %timeit -r3 -n10 pdist_numpy_(xs)
func = pdist_numba_numpy_
print(np.allclose(func(xs), sol))
# %timeit -r3 -n10 func(xs)
# ### Summary
#
# - `numba` appears to work best with converting fairly explicit Python code
# - This might change in the future as the `numba` JIT compiler becomes more sophisticated
# - Always check optimized code for correctness
# - We can use `timeit` magic as a simple way to benchmark functions
# ## Cython
#
# Cython is an Ahead Of Time (AOT) compiler. It compiles the code and replaces the function invoked with the compiled version.
#
# In the notebook, calling `%cython -a` magic shows code colored by how many Python C API calls are being made. You want to reduce the yellow as much as possible.
# + magic_args="-a " language="cython"
#
# import numpy as np
#
# def pdist_cython_1(xs):
# n, p = xs.shape
# A = np.zeros((n, n))
# for i in range(n):
# for j in range(i+1, n):
# d = 0.0
# for k in range(p):
# d += (xs[i,k] - xs[j,k])**2
# A[i,j] = np.sqrt(d)
# A += A.T
# return A
# -
def pdist_base(xs):
n, p = xs.shape
A = np.zeros((n, n))
for i in range(n):
for j in range(i+1, n):
d = 0.0
for k in range(p):
d += (xs[i,k] - xs[j,k])**2
A[i,j] = np.sqrt(d)
A += A.T
return A
# %timeit -r3 -n1 pdist_base(xs)
func = pdist_cython_1
print(np.allclose(func(xs), sol))
# %timeit -r3 -n1 func(xs)
# ## Cython with static types
#
# - We provide types for all variables so that Cython can optimize their compilation to C code.
# - Note `numpy` functions are optimized for working with `ndarrays` and have unnecessary overhead for scalars. We therefor replace them with math functions from the C `math` library.
# + magic_args="-a " language="cython"
#
# import cython
# import numpy as np
# cimport numpy as np
# from libc.math cimport sqrt, pow
#
# @cython.boundscheck(False)
# @cython.wraparound(False)
# def pdist_cython_2(double[:, :] xs):
# cdef int n, p
# cdef int i, j, k
# cdef double[:, :] A
# cdef double d
#
# n = xs.shape[0]
# p = xs.shape[1]
# A = np.zeros((n, n))
# for i in range(n):
# for j in range(i+1, n):
# d = 0.0
# for k in range(p):
# d += pow(xs[i,k] - xs[j,k],2)
# A[i,j] = sqrt(d)
# for i in range(1, n):
# for j in range(i):
# A[i, j] = A[j, i]
# return A
# -
func = pdist_cython_2
print(np.allclose(func(xs), sol))
# %timeit -r3 -n1 func(xs)
# ## Wrapping C++ cdoe
# ### Function to port
# ```python
# def pdist_base(xs):
# n, p = xs.shape
# A = np.zeros((n, n))
# for i in range(n):
# for j in range(i+1, n):
# d = 0.0
# for k in range(p):
# d += (xs[i,k] - xs[j,k])**2
# A[i,j] = np.sqrt(d)
# A += A.T
# return A
# ```
# ### First check that the function works as expected
# +
# %%file main.cpp
#include <iostream>
#include <Eigen/Dense>
#include <cmath>
using std::cout;
// takes numpy array as input and returns another numpy array
Eigen::MatrixXd pdist(Eigen::MatrixXd xs) {
int n = xs.rows() ;
int p = xs.cols();
Eigen::MatrixXd A = Eigen::MatrixXd::Zero(n, n);
for (int i=0; i<n; i++) {
for (int j=i+1; j<n; j++) {
double d = 0;
for (int k=0; k<p; k++) {
d += std::pow(xs(i,k) - xs(j,k), 2);
}
A(i, j) = std::sqrt(d);
}
}
A += A.transpose().eval();
return A;
}
int main() {
using namespace Eigen;
MatrixXd A(3,2);
A << 0, 0,
3, 4,
5, 12;
std::cout << pdist(A) << "\n";
}
# + language="bash"
#
# g++ -o main.exe main.cpp -I./eigen3
# + language="bash"
#
# ./main.exe
# -
A = np.array([
[0, 0],
[3, 4],
[5, 12]
])
squareform(pdist(A))
# ### Now use the boiler plate for wrapping
# +
# %%file wrap.cpp
<%
cfg['compiler_args'] = ['-std=c++11']
cfg['include_dirs'] = ['./eigen3']
setup_pybind11(cfg)
%>
#include <pybind11/pybind11.h>
#include <pybind11/eigen.h>
// takes numpy array as input and returns another numpy array
Eigen::MatrixXd pdist(Eigen::MatrixXd xs) {
int n = xs.rows() ;
int p = xs.cols();
Eigen::MatrixXd A = Eigen::MatrixXd::Zero(n, n);
for (int i=0; i<n; i++) {
for (int j=i+1; j<n; j++) {
double d = 0;
for (int k=0; k<p; k++) {
d += std::pow(xs(i,k) - xs(j,k), 2);
}
A(i, j) = std::sqrt(d);
}
}
A += A.transpose().eval();
return A;
}
PYBIND11_PLUGIN(wrap) {
pybind11::module m("wrap", "auto-compiled c++ extension");
m.def("pdist", &pdist);
return m.ptr();
}
# +
import cppimport
import numpy as np
code = cppimport.imp("wrap")
print(code.pdist(A))
# -
func = code.pdist
print(np.allclose(func(xs), sol))
# %timeit -r3 -n1 func(xs)
# # Pythran and Transonic
#
# [Pythran](https://github.com/serge-sans-paille/pythran) is an ahead-of-time (AOT) compiler for a subset of the Python language, with a focus on scientific computing. It takes a Python module annotated with a few interface description and turns it into a native Python module with the same interface, but (hopefully) faster.
#
# [Transonic](https://transonic.readthedocs.io) is a pure Python package (requiring Python >= 3.6) to easily accelerate modern Python-Numpy code with different accelerators (like Cython, Pythran, Numba, etc…) opportunistically (i.e. if/when they are available). However, Transonic is a very young project so as of today, only Pythran is supported. Here, we use Transonic to use Pythran as a just-in-time compiler.
#
# +
from transonic import jit
# transonic import numpy as np
functions = {"py": pdist_py, "sym": pdist_sym, "vec": pdist_vec}
functions_trans = {key: jit(func) for key, func in functions.items()}
for func in functions_trans.values():
assert np.allclose(func(xs), sol)
# +
from transonic import wait_for_all_extensions
wait_for_all_extensions()
# -
for key, func in functions.items():
print(key, "not compiled:")
assert np.allclose(func(xs), sol)
# %timeit -r3 -n10 func(xs)
func = functions_trans[key]
print(key, "compiled:")
assert np.allclose(func(xs), sol)
# %timeit -r10 -n100 func(xs)
| notebook/S13D_Native_Code_Compilation_Recap.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] _cell_guid="0bd69447-6990-4972-b967-11acf97246d1" _execution_state="idle" _uuid="07183b7e9b11dfa4a0bafe6eab0379ffeb25d3d8"
# # Titanic Prediction using Python
# ### A huge thank you to <NAME> and his Udemy course for teaching me https://www.udemy.com/python-for-data-science-and-machine-learning-bootcamp/learn/v4
# + [markdown] _cell_guid="6b340d12-953b-41b0-bd50-62776d644e02" _execution_state="idle" _uuid="409e41d03ae8eb482d8b422f40ecf89cbc97788d"
# ## Imports and reading in files
# + _cell_guid="f3869157-12f9-41e9-94be-4bdc73c090b5" _execution_state="idle" _uuid="4fdd229f5baf94d1a132155873b5e6159ea1d0f4"
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
from subprocess import check_output
#print(check_output(["ls", "../input"]).decode("utf8"))
import warnings
#warnings.filterwarnings('ignore')
# + _cell_guid="5f7408e7-a053-4861-a7db-b3d1e00fb77a" _execution_state="idle" _uuid="693cd43662518d8af1d562b71396cc599f1bc1e3"
df = pd.read_csv("../input/train.csv")
# + _cell_guid="0129b56b-c792-4fc6-95b5-1afd24d5c4ba" _execution_state="idle" _uuid="773c008ff7a7d8c395078a838ecf5eb6c4fcb6a5"
#Check that the file was read in properly and explore the columns
df.head()
# + [markdown] _cell_guid="52938632-55cd-4bd5-ad9a-0dc905de9f70" _execution_state="idle" _uuid="0dd8f07fd00550568d3a1b88595068f7f0bd31ef"
# ## Data exploration
# -
counter = df.isnull().sum()
counter.map(lambda x : x/df.count().max()*100)
# + _cell_guid="59ccc104-493b-437c-994c-5a7a19d07955" _execution_state="idle" _uuid="dd0daa67fe04087fb33fb472f183a28986273bbd"
plt.figure(figsize=(12,8))
sns.heatmap(df.isnull(),cbar=True, yticklabels=False, cmap='viridis')
# + [markdown] _cell_guid="90cfc754-6257-4a06-9454-fc585c9da834" _execution_state="idle" _uuid="3fb06bc89e244f2c3b9de608208544059729753b"
# From the heat map, we can see that a lot of the 'Cabin' row information is missing. However, while the age column is also missing some data, we can use imputation to fill in some of the data later. Additionally, the 'Embarked' column has so few rows missing, that we can just delete those.
# + _cell_guid="d3742711-40fa-47b6-bcb0-a95ec4560144" _execution_state="idle" _uuid="62febb18cdedc10a5ba1681b33fac82f675a239b"
sns.set_style('darkgrid')
# + _cell_guid="cdf4187b-43e6-41af-abd9-5e9f4611ac4b" _execution_state="idle" _uuid="3124ce9a5a3286c41362194dbbc51b3641161c09"
sns.countplot(x='Survived', data=df, hue='Pclass')
# + [markdown] _cell_guid="5e579544-b525-4eca-a095-5913f08457e7" _execution_state="idle" _uuid="515cacd2ff70f77a21effc3ff6dffa634f2294ae"
# We can see here that those who did not survive were predominantly from the 3rd Passenger Class (Pclass).
# + _cell_guid="debc9053-784d-4934-b2c8-6fa8d4b3f749" _execution_state="idle" _uuid="ab72e9a84a399ef71160eb27bf4361cda91600e4"
sns.countplot(x='SibSp', data=df, hue='Survived')
# + _cell_guid="428c6f9e-e35f-4152-9365-b844545281b8" _execution_state="idle" _uuid="00459b863e0127cb0d2f9bd7ecffd9aa9fe115f9"
df['Fare'].hist(bins=40)
# + [markdown] _cell_guid="5f05fe07-7732-46f5-90e1-3d9d33d1bcd6" _execution_state="idle" _uuid="00cbc7c157074efb35bdc52cd58880b4e5921d17"
# Here, we impute the age of those we do not have information on. We use a boxplot to estimate the median age of each class, and impute that into the age for the rows with missing age.
# + _cell_guid="248f2650-2310-4291-9a0b-ec3160e592ff" _execution_state="idle" _uuid="24f0efcbfe90c76105d846286854fde301a74efb"
plt.figure(figsize=(12,6))
sns.boxplot(x='Pclass', y='Age', data=df)
# + _cell_guid="2f094d5d-3ac5-4b03-907a-b6e36a2489e8" _execution_state="idle" _uuid="2b4dfd63718b9172b1ade9ac75cda872b9fe07fe"
def inpute_age(cols):
Age = cols[0]
Pclass = cols[1]
if pd.isnull(Age):
if Pclass == 1:
return 37
elif Pclass == 2:
return 29
else: return 24
else: return Age
# + _cell_guid="87547ef8-2f60-4104-b8b7-978ba770cd17" _execution_state="idle" _uuid="0720c1ff1a23be5509864450825fc1f2d82d74f0"
df['Age']=df[['Age','Pclass']].apply(inpute_age, axis=1)
# + _cell_guid="c4170754-00c0-4f2f-800c-ed19d2462cd5" _execution_state="idle" _uuid="de28c0c2adf5af525b83df19afbc7fe83ad48fe0"
plt.figure(figsize=(12,8))
sns.heatmap(df.isnull(),cbar=False, yticklabels=False, cmap='viridis')
# + [markdown] _cell_guid="53f6a54e-1bbb-4df1-9419-8f3f5c34f6fa" _execution_state="idle" _uuid="5f1fe7e3b6954d257800c9dc46cc8861fa21bffb"
# You can see now the data is cleaner, but we still need to clean the 'Cabin' and 'Embarked' columns. For now, we will simply drop the 'Cabin' column and drop the rows where 'Embarked' is missing.
# + _cell_guid="036f5642-eadb-4344-b690-4b4c6ae61d44" _execution_state="idle" _uuid="92745eafe0f4c319f0b706c02dfe99187105fd58"
df.drop('Cabin', axis=1, inplace=True)
# + _cell_guid="054b705f-d37e-4178-a80c-8314bcf21f98" _execution_state="idle" _uuid="5a44851769b5c5612af406d7d4bfd040d3cb4b1c"
plt.figure(figsize=(12,6))
sns.heatmap(df.isnull(),cbar=False, yticklabels=False, cmap='viridis')
# + _cell_guid="6fcd7796-aefd-4193-936d-d669b20039e0" _execution_state="idle" _uuid="6c411ada1594fb55afb29c67f808462df60ddb76"
df.dropna(inplace=True)
# + _cell_guid="4b45a567-c5d1-469c-a559-b7ddbe386c40" _execution_state="idle" _uuid="5fff94d248bcc6e8226932a56ed1084b21b877d1"
plt.figure(figsize=(12,6))
sns.heatmap(df.isnull(),cbar=False, yticklabels=False, cmap='viridis')
# + [markdown] _cell_guid="2a8d2ca0-a05c-4dfd-972e-5f6647797022" _execution_state="idle" _uuid="fb2c0385a4f2fe30b22da60d85c583ec9dbb2bd4"
# The data is now clean of null values, but we still need to take care of objects that a machine learning algorithm can't handle, namely strings.
# + _cell_guid="9d0f603b-d655-4b57-ae3a-a4d030420116" _execution_state="idle" _uuid="380361912c14a39eff883d29096567344725c510"
df.info()
# + [markdown] _cell_guid="829126a5-d93d-4bad-ac9a-f53711e3f247" _execution_state="idle" _uuid="f49c76a58ec6a2898d4147df524e03c605299787"
# We can see that 'Name', 'Sex', 'Ticket', and 'Embarked' are all objects. In this case, they indeed are all strings. We will use Pandas built in getDummies() funciton to convert those to numbers.
# + _cell_guid="2c7c8330-6a64-435b-ad3a-a3fc0894f3af" _execution_state="idle" _uuid="a9cc4cd395c0eef5328154c1460d31fcbcee27da"
#We make a new 'Male columns because getDummies will drop one the the dummy variables
#to ensure linear independence.
df['Male'] = pd.get_dummies(df['Sex'], drop_first=True)
# + _cell_guid="ed08af19-a1f6-4064-8141-a213c3e6dfb9" _execution_state="idle" _uuid="7de16a191a73564bc406aaf4b833e332edc17c0e"
#The embarked column indicates where the passenger boarded the Titanic.
#It has three values ['S','C','Q']
embarked = pd.get_dummies(df['Embarked'], drop_first=True)
df = pd.concat([df, embarked], axis=1)
# + _cell_guid="48274961-7341-4b58-83d3-f02199b6db40" _execution_state="idle" _uuid="f0f12671cbbc5d018366a852b00368a5ff03475f"
#These columns do not provide us any information for the following reasons:
#PassengerID: we consider 'PassengerID' a randomly assigned ID thus not correlated with surviability
#Name: we are not performing any feature extraction from the name, so we must drop tihs non-numerical column
#Sex: the 'Male' column already captures all information about the sex of the passenger
#Ticket: we are not performing any feature extraction, so we must drop this non-numerical column
#Embarked: we have extracted the dummy values, so those two numerical dummy values encapsulate all the embarked info
df.drop(['PassengerId', 'Name', 'Sex', 'Ticket', 'Embarked'], axis=1, inplace=True)
# + _cell_guid="1088380d-feca-48f9-8932-6b49218e849a" _execution_state="idle" _uuid="1f8a997db6bee8d669aa6818d1f48d0dc3a6b5f5"
#Take a look at our new dataframe
df.head()
# + _cell_guid="1920c67b-641b-40b5-b015-77d17ebbcf44" _execution_state="idle" _uuid="06ef694ed1bc6f6bb574b57ab41dd97859cfb2e5"
df.info()
# + [markdown] _cell_guid="b8506c54-2dd5-46ab-ab6c-38a1f82d1110" _execution_state="idle" _uuid="539c55ea8337dd0b1792d7b4f2c4226f984ad9fc"
# ##Build and train the model
# + _cell_guid="d4fa8385-85ab-416a-aafd-13415a9512b3" _execution_state="idle" _uuid="ded9b4360db02587f1977fc3d47b946d4797fb7b"
#Seperate the feature columns from the target column
X = df.drop('Survived', axis=1)
y = df['Survived']
# + _cell_guid="c7c72a22-2346-406b-a470-392c4083b392" _execution_state="idle" _uuid="92a1213ede40ded8e80153fa8fe841bd5bad10ef"
#Split the data into two. I don't think this is necessary since there are two files.
#I will keep this here for now
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X,y, test_size=0.3)
# + _cell_guid="8da84909-b2f7-4aa0-be5d-5c46cab71d1b" _execution_state="idle" _uuid="accd29cc3a075b81bd13129df840054224e8b396"
from sklearn.linear_model import LogisticRegression
logmodel = LogisticRegression()
logmodel.fit(X, y)
# -
# Let's get our model accuracy based on our test data
predicted = logmodel.predict(X_test)
print('accuracy is: {:.0%}'.format((predicted == y_test).sum()/y_test.count()))
# + _cell_guid="ef5f966b-b8c6-4b9d-b3ee-455dbd53e35e" _execution_state="idle" _uuid="45fab0ccf873872eca09527ae9fa47fceee0e3b2"
#Read in the test data
test_df = pd.read_csv('../input/test.csv')
# + _cell_guid="946f68e4-d359-48a0-aeb7-969bc4d3f4f4" _execution_state="idle" _uuid="0b86f9e7faaff6fe763dbe3cce6b0071a6ae4858"
#Clean the test data the same way we did the training data
test_df['Age']=test_df[['Age','Pclass']].apply(inpute_age, axis=1)
test_df.drop('Cabin', axis=1, inplace=True)
test_df.dropna(inplace=True)
test_df['Male'] = pd.get_dummies(test_df['Sex'], drop_first=True)
embarked = pd.get_dummies(test_df['Embarked'], drop_first=True)
test_df = pd.concat([test_df, embarked], axis=1)
pass_ids = test_df['PassengerId']
test_df.drop(['PassengerId', 'Name', 'Sex', 'Ticket', 'Embarked'], axis=1, inplace=True)
# + _cell_guid="9ec7a484-3259-47f6-a7ae-067979bc39b8" _execution_state="idle" _uuid="f8b930fdfa969b55c8ef15a27c8d91454983d5bb"
test_df.tail()
# + _cell_guid="70d73cb3-1c06-48cc-8fc4-538b42cdaddf" _execution_state="idle" _uuid="12b9c3816e860c4682dc89ed5939e7c5d19975ed"
predictions = logmodel.predict(test_df)
# + _cell_guid="323d6dcd-de82-41bd-b1d0-5772c1c6f412" _execution_state="idle" _uuid="065e3c6fa7df9db4388a8ab7415f091bc7074826"
submission = pd.DataFrame({
"PassengerId": pass_ids,
"Survived": predictions
})
submission.to_csv('titanic.csv', index=False)
| #4/notebooks/Titanic_logistic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] nbgrader={}
# # Numpy Exercise 2
# + [markdown] nbgrader={}
# ## Imports
# + nbgrader={}
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
# + [markdown] nbgrader={}
# ## Factorial
# + [markdown] nbgrader={}
# Write a function that computes the factorial of small numbers using `np.arange` and `np.cumprod`.
# + nbgrader={"checksum": "4e72a0f70d32914a77d8d31ae0c2fb5a", "solution": true}
def np_fact(n):
if n == 0:
return 1
else:
#This puts the numbers 1 to n in steps of one in an array
numbers = np.arange(1.0, n+1, 1.0)
#The cumprod command makes a list of the factorials up to n!
#The [-1] call takes the last element in the array (n!)
product = numbers.cumprod(0)[-1]
return product
np_fact(6)
# + deletable=false nbgrader={"checksum": "fcb54de452abd89e3b1818a34678d4b4", "grade": true, "grade_id": "numpyex02a", "points": 4}
assert np_fact(0)==1
assert np_fact(1)==1
assert np_fact(10)==3628800
assert [np_fact(i) for i in range(0,11)]==[1,1,2,6,24,120,720,5040,40320,362880,3628800]
# + [markdown] nbgrader={}
# Write a function that computes the factorial of small numbers using a Python loop.
# + nbgrader={"checksum": "9341aa5eb68a3ce57e5cce1cd9192021", "solution": true}
def loop_fact(n):
answer = 1
for i in range(n):
#The answer is multipled by 1 to n (n!)
answer = answer * (i+1)
return answer
loop_fact(6)
# + deletable=false nbgrader={"checksum": "a9946f0fdca1abf66ddfb0454ab10113", "grade": true, "grade_id": "numpyex02b", "points": 4}
assert loop_fact(0)==1
assert loop_fact(1)==1
assert loop_fact(10)==3628800
assert [loop_fact(i) for i in range(0,11)]==[1,1,2,6,24,120,720,5040,40320,362880,3628800]
# + [markdown] nbgrader={}
# Use the `%timeit` magic to time both versions of this function for an argument of `50`. The syntax for `%timeit` is:
#
# ```python
# # # %timeit -n1 -r1 function_to_time()
# ```
# + deletable=false nbgrader={"checksum": "6cff4e8e53b15273846c3aecaea84a3d", "solution": true}
# %timeit -n1 -r1 np_fact(5)
# %timeit -n1 -r1 loop_fact(5)
# + [markdown] nbgrader={}
# In the cell below, summarize your timing tests. Which version is faster? Why do you think that version is faster?
# + [markdown] deletable=false nbgrader={"checksum": "84e52188175a63ea7106817b9b063eef", "grade": true, "grade_id": "numpyex02c", "points": 2, "solution": true}
# In np_fact, the computer must first create an array of elements 1, 2, ... (n-1) to n. Then, the cumprod function creates an array of all the factorials to n! (1!, 2!,... (n-1)!, n!).
#
# In loop_fact, the computer takes each numbers 0 to (n-1), adds one to it and multiplies it to a variable, n times.
#
# np_fact is slower because it has a greater amount of operations to complete. loop_fact is more simple, as it requires taking an integer in a range, adding one, multiplying by a variable and reassigning the variabe. Whereas in np_fact, a range of numbers is divided by step size, then cumprod creates an array containing each number's factorial and I only call the last element in that array.
| assignments/assignment03/NumpyEx02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] deletable=true editable=true
# # Initializing Supervised Learning
# + [markdown] deletable=true editable=true
# ## 1. Purpose
#
# Apply supervised learning methods to the data
# + [markdown] deletable=true editable=true
# ## 2. Preparation
#
# ### 2.0 Import Required Packages
# +
# #%reset
# + deletable=true editable=true
## Data Mining
import pandas as pd
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
# %matplotlib inline
plt.style.use('ggplot')
## Classification Method
from sklearn import preprocessing, neighbors ## KNN
import sklearn.linear_model as skl_lm ## Linear Methods (Logit)
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis ## Linear Discricimant Analysis
from sklearn.svm import SVC ## Support Vector Machine
from sklearn.naive_bayes import BernoulliNB ## Naive Bayes
from sklearn.ensemble import RandomForestClassifier ## Random Forest
## Model Selection and Metrics
from sklearn.model_selection import train_test_split, LeaveOneOut
from sklearn.metrics import confusion_matrix, classification_report, precision_recall_curve, roc_curve, auc
# + [markdown] deletable=true editable=true
# ### 2.1 Load Data
# + deletable=true editable=true
## All Data
#alldata = pd.read_csv("../../data/alldata_traincode_170510.csv", encoding='CP932')
#allWrdMat10 = pd.read_csv("../../data/allWrdMat10.csv.gz", encoding='CP932')
## Test on Cabinet Data
cabinetdata = pd.read_csv('../data_public/cabinetdata_traincode.csv', encoding='CP932', na_values='NA')
cabinetWrdMat10 = pd.read_csv('../data_public/cabinetWrdMat10.csv', encoding='CP932', na_values='NA')
## Test on LDP Data
#ldpdata = pd.read_csv('../data_public/ldpdata_traincode.csv', encoding='CP932', na_values='NA')
#ldpWrdMat10 = pd.read_csv('../data_public/ldpWrdMat10.csv', encoding='CP932', na_values='NA')
## Test on Politics Data (FOR THIS EXAMPLE)
#politicsdata = pd.read_csv('../data_public/politicsdata_traincode.csv', encoding='CP932', na_values='NA')
#politicsWrdMat10 = pd.read_csv('../data_public/politicsWrdMat10.csv', encoding='CP932', na_values='NA')
# + deletable=true editable=true
cabinetdata.head(3)
# + deletable=true editable=true
cabinetWrdMat10.head(3)
# + [markdown] deletable=true editable=true
# ### 2.2 Prepare Training Data and Testing Data
# + deletable=true editable=true
targetx = cabinetWrdMat10
targety = cabinetdata
X = np.array(targetx.loc[targety['train'] == 1,:])
y = np.array(targety.loc[targety['train'] == 1,'codeN'])
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=.3, random_state=0)
# + [markdown] deletable=true editable=true
# ## 3. Conduct Classification
# + [markdown] deletable=true editable=true
# ### 3.1 K Nearest Neighbors
# + deletable=true editable=true
## With k = 10
k = 10
knn1 = neighbors.KNeighborsClassifier(n_neighbors=k) ##knn estimation
knn1.fit(X_train, y_train)
## On Test Set
proby = knn1.predict_proba(X_test)[:,1] # predicted Y
knn1fpr, knn1tpr, _ = roc_curve(y_test, proby) #ROC Curve
knn1auc = auc(knn1fpr, knn1tpr) # Area Under Curve (AUC)
knn1prec, knn1rec, _ = precision_recall_curve(y_test, proby) #PR Curve
## With k = 50
k = 50
knn2 = neighbors.KNeighborsClassifier(n_neighbors=k) ##knn estimation
knn2.fit(X_train, y_train)
## On Test Set
proby = knn2.predict_proba(X_test)[:,1] # predicted Y
knn2fpr, knn2tpr, _ = roc_curve(y_test, proby) #ROC Curve
knn2auc = auc(knn2fpr, knn2tpr) # Area Under Curve (AUC)
knn2prec, knn2rec, _ = precision_recall_curve(y_test, proby) #PR Curve
# + [markdown] deletable=true editable=true
# ### 3.2 Logistic Regression (With Newton Conjugate Solver)
# + deletable=true editable=true
## Logistic Regression
logit = skl_lm.LogisticRegression(solver='newton-cg')
logit.fit(X_train,y_train)
## On Test Set
proby = logit.predict_proba(X_test)[:,1] # predicted Y
logitfpr, logittpr, _ = roc_curve(y_test, proby) #ROC Curve
logitauc = auc(logitfpr, logittpr) # Area Under Curve (AUC)
logitprec, logitrec, _ = precision_recall_curve(y_test, proby) #PR Curve
# + [markdown] deletable=true editable=true
# ### 3.3 Linear Discriminant Analysis
# + deletable=true editable=true
## Linear Discriminant Analysis
lda = LinearDiscriminantAnalysis()
lda.fit(X_train,y_train)
## On Test Set
proby = lda.predict_proba(X_test)[:,1] # predicted Y
ldafpr, ldatpr, _ = roc_curve(y_test, proby) #ROC Curve
ldaauc = auc(ldafpr, ldatpr) # Area Under Curve (AUC)
ldaprec, ldarec, _ = precision_recall_curve(y_test, proby) #PR Curve
# + [markdown] deletable=true editable=true
# ### 3.4 Support Vector Machine
# + deletable=true editable=true
## Support Vector Machines
svmcl = SVC(probability=True)
svmcl.fit(X_train,y_train)
## On Test Set
proby = svmcl.predict_proba(X_test)[:,1] # predicted Y
svmfpr, svmtpr, _ = roc_curve(y_test, proby) #ROC Curve
svmauc = auc(svmfpr, svmtpr) # Area Under Curve (AUC)
svmprec, svmrec, _ = precision_recall_curve(y_test, proby) #PR Curve
# -
# ### 3.5 Bernoulli Naive Bayes
# +
## Bernoulli Naive Bayes
nb = BernoulliNB()
nb.fit(X_train, y_train)
## On Test Set
proby = nb.predict_proba(X_test)[:,1] # predicted Y
nbfpr, nbtpr, _ = roc_curve(y_test, proby) #ROC Curve
nbauc = auc(nbfpr, nbtpr) # Area Under Curve (AUC)
nbprec, nbrec, _ = precision_recall_curve(y_test, proby) #PR Curve
# -
# ### 3.5 Random Forest
# +
## Random Forest
rf = RandomForestClassifier(n_estimators=200)
rf.fit(X_train, y_train)
## On Test Set
proby = rf.predict_proba(X_test)[:,1] # predicted Y
rffpr, rftpr, _ = roc_curve(y_test, proby) #ROC Curve
rfauc = auc(rffpr, rftpr) # Area Under Curve (AUC)
rfprec, rfrec, _ = precision_recall_curve(y_test, proby) #PR Curve
# + [markdown] deletable=true editable=true
# ## 4. Compare The Result
# + [markdown] deletable=true editable=true
# ### 4.1 ROC Curve
# + deletable=true editable=true
plt.rcParams["figure.figsize"] = (17,4)
f, (ax02, ax1, ax2, ax3, ax4, ax5), = plt.subplots(1, 6, sharey=True)
f.text(0.5, 0.02, 'False Positive Rate', ha='center', va='center', fontsize = 15)
f.text(0.06, 0.5, 'True Positive Rate', ha='center', va='center', rotation='vertical', fontsize = 15)
ax01.plot(knn1fpr, knn1tpr, 'k-', label = 'ROC Curve \n(Area = %0.3f)' % knn1auc)
ax01.plot([0, 1], [0, 1], color='gray', lw=2, linestyle='--')
ax01.set_title('KNN (K=10)');
ax01.set_xlim(-0.05,1.05)
ax01.set_ylim(-0.05,1.05)
ax01.legend(loc="lower right")
ax02.plot(knn2fpr, knn2tpr, 'k-', label = 'ROC Curve \n(Area = %0.3f)' % knn2auc)
ax02.plot([0, 1], [0, 1], color='gray', lw=2, linestyle='--')
ax02.set_title('KNN (K=50)');
ax02.set_xlim(-0.05,1.05)
ax02.set_ylim(-0.05,1.05)
ax02.legend(loc="lower right")
ax1.plot(logitfpr, logittpr, 'r-', label = 'ROC Curve \n(Area = %0.3f)' % logitauc)
ax1.plot([0, 1], [0, 1], color='gray', lw=2, linestyle='--')
ax1.set_title('Logit');
ax1.set_xlim(-0.05,1.05)
ax1.set_ylim(-0.05,1.05)
ax1.legend(loc="lower right")
ax2.plot(ldafpr, ldatpr, 'g-', label = 'ROC Curve \n(Area = %0.3f)' % ldaauc)
ax2.plot([0, 1], [0, 1], color='gray', lw=2, linestyle='--')
ax2.set_title('LDA');
ax2.set_xlim(-0.05,1.05)
ax2.set_ylim(-0.05,1.05)
ax2.legend(loc="lower right")
ax3.plot(svmfpr, svmtpr, 'b-', label = 'ROC Curve \n(Area = %0.3f)' % svmauc)
ax3.plot([0, 1], [0, 1], color='gray', lw=2, linestyle='--')
ax3.set_title('SVM');
ax3.set_xlim(-0.05,1.05)
ax3.set_ylim(-0.05,1.05)
ax3.legend(loc="lower right")
ax4.plot(nbfpr, nbtpr, 'm-', label = 'ROC Curve \n(Area = %0.3f)' % nbauc)
ax4.plot([0, 1], [0, 1], color='gray', lw=2, linestyle='--')
ax4.set_title('Naive Bayes');
ax4.set_xlim(-0.05,1.05)
ax4.set_ylim(-0.05,1.05)
ax4.legend(loc="lower right")
ax5.plot(rffpr, rftpr, 'c-', label = 'ROC Curve \n(Area = %0.3f)' % rfauc)
ax5.plot([0, 1], [0, 1], color='gray', lw=2, linestyle='--')
ax5.set_title('Random Forest');
ax5.set_xlim(-0.05,1.05)
ax5.set_ylim(-0.05,1.05)
ax5.legend(loc="lower right")
plt.show()
# + [markdown] deletable=true editable=true
# ### 4.2 PR Curve
# + deletable=true editable=true
plt.rcParams["figure.figsize"] = (17,4)
f, (ax02, ax1, ax2, ax3, ax4, ax5) = plt.subplots(1, 6, sharey=True)
f.text(0.5, 0.02, 'Recall', ha='center', va='center', fontsize = 15)
f.text(0.06, 0.5, 'Precision', ha='center', va='center', rotation='vertical', fontsize = 15)
ax01.plot(knn1rec, knn1prec, 'k-', label = 'PR Curve')
ax01.set_title('KNN (K=10)');
ax01.set_xlim(-0.05,1.05)
ax01.set_ylim(-0.05,1.05)
ax01.legend(loc="lower right")
ax02.plot(knn2rec, knn2prec, 'k-', label = 'PR Curve')
ax02.set_title('KNN (K=50)');
ax02.set_xlim(-0.05,1.05)
ax02.set_ylim(-0.05,1.05)
ax02.legend(loc="lower right")
ax1.plot(logitrec, logitprec, 'r-', label = 'PR Curve')
ax1.set_title('Logit');
ax1.set_xlim(-0.05,1.05)
ax1.set_ylim(-0.05,1.05)
ax1.legend(loc="lower right")
ax2.plot(ldarec, ldaprec, 'g-', label = 'PR Curve')
ax2.set_title('LDA');
ax2.set_xlim(-0.05,1.05)
ax2.set_ylim(-0.05,1.05)
ax2.legend(loc="lower right")
ax3.plot(svmrec, svmprec, 'b-', label = 'PR Curve')
ax3.set_title('SVM');
ax3.set_xlim(-0.05,1.05)
ax3.set_ylim(-0.05,1.05)
ax3.legend(loc="lower right")
ax4.plot(nbrec, nbprec, 'm-', label = 'PR Curve')
ax4.set_title('Naive Bayes');
ax4.set_xlim(-0.05,1.05)
ax4.set_ylim(-0.05,1.05)
ax4.legend(loc="lower right")
ax5.plot(rfrec, rfprec, 'c-', label = 'PR Curve')
ax5.set_title('Random Forest');
ax5.set_xlim(-0.05,1.05)
ax5.set_ylim(-0.05,1.05)
ax5.legend(loc="lower right")
plt.show()
# -
| notebooks/.ipynb_checkpoints/Supervised Learning-Copy1-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # I am missing you
#
# Using https://data.medicare.gov/Hospital-Compare/Hospital-Readmissions-Reduction-Program/9n3s-kdb3
#
# In October 2012, CMS began reducing Medicare payments for Inpatient Prospective Payment System hospitals with excess readmissions. Excess readmissions are measured by a ratio, by dividing a hospital’s number of “predicted” 30-day readmissions for heart attack, heart failure, pneumonia, chronic obstructive pulmonary disease, hip/knee replacement, and coronary artery bypass graft surgery by the number that would be “expected,” based on an average hospital with similar patients. A ratio greater than 1.0000 indicates excess readmissions.
# %matplotlib inline
import numpy as np
import pandas as pd
import missingno as mno
from stemgraphic.alpha import stem_graphic as sga
# !head data/Hospital_Readmissions_Reduction_Program.csv
df = pd.read_csv('data/Hospital_Readmissions_Reduction_Program.csv')
df.describe(include='all')
mno.matrix(df.sample(250));
# What you don't know: VISU.AI scores this at around 30% invalid data. stemgraphic.alpha can help us, as it can cope with mixed data, and let us visualize it as stem-and-leaf:
sga(df['Excess Readmission Ratio'].tolist(), display=250);
# Just for that one column, we found many 0.x and 1.x values, but also, almost as many strings starting with No....
df['Excess Readmission Ratio'].unique()
for col in df.columns:
try:
df[col][df[col] == 'Not Available'] = np.nan
except TypeError:
pass
mno.matrix(df.sample(1000))
# Let's move on to the other columns now.
#
# First column, Hospital Name, is text. Most common way to visualize words? yep... word cloud. But...
from wordcloud import WordCloud
wordcloud = WordCloud().generate(' '.join(df['Hospital Name'].values))
import matplotlib.pyplot as plt
plt.rcParams["figure.figsize"] = (15,10)
plt.imshow(wordcloud, interpolation='bilinear')
plt.axis("off");
# # Just for North Carolina
#
# We can make this prettier, but...
from os import path
from PIL import Image
nc_mask = np.array(Image.open(path.join('.', "nc_mask.jpg")))
wordcloud = WordCloud(background_color="white", mask=nc_mask, width=800, height=400)
wordcloud.generate(' '.join(df['Hospital Name'][df.State=='NC'].values));
plt.imshow(wordcloud, interpolation='bilinear')
plt.axis("off");
# What the word cloud is supposed to answer: ranking
#
# There are better options.
from stemgraphic.alpha import word_freq_plot, word_sunburst
# I'm lazy so I'll just ingest the whole file and let the visualization do its thing.
word_sunburst('data/Hospital_Readmissions_Reduction_Program.csv', top=10, sort_by='count');
word_freq_plot('data/Hospital_Readmissions_Reduction_Program.csv', top=40, sort_by='count');
# Again, "Not Available" is showing up. A lot. And what about, as we go up the chart, "to", "report", "few", "too", all have the same frequency. "Too few to report", perhaps? Visualizations can really help to understand what kind of data we are dealing with.
| 01-missing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Lecture 14: Interacting with the filesystem
#
# CSCI 1360: Foundations for Informatics and Analytics
# + [markdown] slideshow={"slide_type": "slide"}
# ## Overview and Objectives
#
# So far, all the data we've worked with have either been manually instantiated as NumPy arrays or lists of strings, or randomly generated. Here we'll finally get to go over reading to and writing from the filesystem. By the end of this lecture, you should be able to:
# + [markdown] slideshow={"slide_type": "fragment"}
# - Implement a basic file reader / writer using built-in Python tools
# - Use exception handlers to make your interactions with the filesystem robust to failure
# - Use Python tools to move around the filesystem
# + [markdown] slideshow={"slide_type": "slide"}
# ## Part 1: Interacting with text files
# + [markdown] slideshow={"slide_type": "fragment"}
# Text files are probably the most common and pervasive format of data. They can contain almost anything: weather data, stock market activity, literary works, and raw web data.
# + [markdown] slideshow={"slide_type": "fragment"}
# Text files are also convenient for your own work: once some kind of analysis has finished, it's nice to dump the results into a file you can inspect later.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Reading an entire file
# + [markdown] slideshow={"slide_type": "fragment"}
# So let's jump into it! Let's start with something simple; say...the text version of <NAME>'s *Alice in Wonderland*?
# + slideshow={"slide_type": "fragment"}
file_object = open("alice.txt", "r")
contents = file_object.read()
print(contents[:71])
file_object.close()
# + [markdown] slideshow={"slide_type": "fragment"}
# Yep, I went there.
# + [markdown] slideshow={"slide_type": "slide"}
# Let's walk through the code, line by line. First, we have a call to a function `open()` that accepts two arguments:
# + slideshow={"slide_type": "fragment"}
file_object = open("alice.txt", "r")
# + [markdown] slideshow={"slide_type": "fragment"}
# - The first argument is the *file path*. It's like a URL, except to a file on your computer. It should be noted that, unless you specify a leading forward slash `"/"`, Python will interpret this path to be *relative* to wherever the Python script is that you're running with this command.
# + [markdown] slideshow={"slide_type": "fragment"}
# - The second argument is the *mode*. This tells Python whether you're reading from a file, writing to a file, or appending to a file. We'll come to each of these.
# + [markdown] slideshow={"slide_type": "fragment"}
# These two arguments are part of the function `open()`, which then returns a *file descriptor*. You can think of this kind of like the reference / pointer discussion we had in our prior functions lecture: `file_object` is a reference to the file.
# + [markdown] slideshow={"slide_type": "slide"}
# The next line is where the magic happens:
# + slideshow={"slide_type": "fragment"}
contents = file_object.read()
# + [markdown] slideshow={"slide_type": "fragment"}
# In this line, we're calling the method `read()` on the file reference we got in the previous step. This method goes into the file, pulls out *everything* in it, and sticks it all in the variable `contents`. One big string!
# + slideshow={"slide_type": "fragment"}
print(contents[:71])
# + [markdown] slideshow={"slide_type": "fragment"}
# ...of which I then print the first 71 characters, which contains the name of the book and the author. Feel free to print the entire string `contents`; it'll take a few seconds, as you're printing the whole book!
# + [markdown] slideshow={"slide_type": "slide"}
# Finally, the last and possibly most important line:
# + slideshow={"slide_type": "fragment"}
file_object.close()
# + [markdown] slideshow={"slide_type": "fragment"}
# This statement explicitly closes the file reference, effectively shutting the valve to the file.
# + [markdown] slideshow={"slide_type": "fragment"}
# **Do not** underestimate the value of this statement. There are weird errors that can crop up when you forget to close file descriptors. It can be difficult to remember to do this, though; in other languages where you have to manually allocate and release any memory you use, it's a bit easier to remember. Since Python handles all that stuff for us, it's not a force of habit to explicitly shut off things we've turned on.
# + [markdown] slideshow={"slide_type": "slide"}
# Fortunately, there's an alternative we can use!
# + slideshow={"slide_type": "fragment"}
with open("alice.txt", "r") as file_object:
contents = file_object.read()
print(contents[:71])
# + [markdown] slideshow={"slide_type": "fragment"}
# This code works identically to the code before it. The difference is, by using a `with` block, Python intrinsically closes the file descriptor at the end of the block. Therefore, no need to remember to do it yourself! Hooray!
# + [markdown] slideshow={"slide_type": "slide"}
# Let's say, instead of *Alice in Wonderland*, we had some behemoth of a piece of literature: something along the lines of *War and Peace* or even an entire encyclopedia. Essentially, not something we want to read into Python *all at once.* Fortunately, we have an alternative:
# + slideshow={"slide_type": "fragment"}
with open("alice.txt", "r") as file_object:
num_lines = 0
for line_of_text in file_object:
print(line_of_text)
num_lines += 1
if num_lines == 5: break
# + [markdown] slideshow={"slide_type": "fragment"}
# We can use a `for` loop just as we're used to doing with lists. In this case, at each iteration, Python will hand you **exactly 1** line of text from the file to handle it however you'd like.
# + [markdown] slideshow={"slide_type": "slide"}
# Of course, if you still want to read in the entire file at once, but really like the idea of splitting up the file line by line, there's a function for that, too:
# + slideshow={"slide_type": "fragment"}
with open("alice.txt", "r") as file_object:
lines_of_text = file_object.readlines()
print(lines_of_text[0])
# + [markdown] slideshow={"slide_type": "fragment"}
# By using `readlines()` instead of plain old `read()`, we'll get back a list of strings, where each element of the list is a single line in the text file. In the code snippet above, I've printed the first line of text from the file.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Writing to a file
# + [markdown] slideshow={"slide_type": "fragment"}
# We've so far seen how to *read* data from a file. What if we've done some computations and want to save our results to a file?
# + slideshow={"slide_type": "fragment"}
data_to_save = "This is important data. Definitely worth saving."
with open("outfile.txt", "w") as file_object:
file_object.write(data_to_save)
# + [markdown] slideshow={"slide_type": "fragment"}
# You'll notice two important changes from before:
# + [markdown] slideshow={"slide_type": "fragment"}
# 1. Switch the `"r"` argument in the `open()` function to `"w"`. You guessed it: we've gone from **R**eading to **W**riting.
# 2. Call `write()` on your file descriptor, and pass in the data you want to write to the file (in this case, `data_to_save`).
# + [markdown] slideshow={"slide_type": "fragment"}
# If you try this using a new notebook on JupyterHub (or on your local machine), you should see a new text file named "`outfile.txt`" appear in the same directory as your script. Give it a shot!
# + [markdown] slideshow={"slide_type": "slide"}
# Some notes about writing to a file:
# + [markdown] slideshow={"slide_type": "fragment"}
# - If the file you're writing to does NOT currently exist, Python will try to create it for you. In most cases this should be fine (but we'll get to outstanding cases in Part 3 of this lecture).
# + [markdown] slideshow={"slide_type": "fragment"}
# - If the file you're writing to DOES already exist, Python will overwrite everything in the file with the new content. As in, **everything that was in the file before will be erased**.
# + [markdown] slideshow={"slide_type": "fragment"}
# That second point seems a bit harsh, doesn't it? Luckily, there is recourse.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Appending to an existing file
# + [markdown] slideshow={"slide_type": "fragment"}
# If you find yourself in the situation of writing to a file multiple times, and wanting to keep what you wrote to the file previously, then you're in the market for *appending* to a file.
# + [markdown] slideshow={"slide_type": "fragment"}
# This works *exactly* the same as writing to a file, with one small wrinkle:
# + slideshow={"slide_type": "fragment"}
data_to_save = "This is ALSO important data. BOTH DATA ARE IMPORTANT."
with open("outfile.txt", "a") as file_object:
file_object.write(data_to_save)
# + [markdown] slideshow={"slide_type": "fragment"}
# The only change that was made was switching the `"w"` in the `open()` method to `"a"` for, you guessed it, **A**ppend. If you look in `outfile.txt`, you should see both lines of text we've written.
# + [markdown] slideshow={"slide_type": "slide"}
# Some notes on appending to files:
# + [markdown] slideshow={"slide_type": "fragment"}
# - If the file does NOT already exist, then using "a" in `open()` is functionally identical to using "w".
# + [markdown] slideshow={"slide_type": "fragment"}
# - You only need to use append mode if you *closed* the file descriptor to that file previously. If you have an open file descriptor, you can call `write()` multiple times; each call will append the text to the previous text. It's only when you *close* a descriptor, but then want to open up another one to the *same file*, that you'd need to switch to append mode.
# + [markdown] slideshow={"slide_type": "slide"}
# Let's put together what we've seen by writing to a file, appending more to it, and then reading what we wrote.
# + slideshow={"slide_type": "fragment"}
data_to_save = "This is important data. Definitely worth saving.\n"
with open("outfile.txt", "w") as file_object:
file_object.write(data_to_save)
# + slideshow={"slide_type": "fragment"}
data_to_save = "This is ALSO important data. BOTH DATA ARE IMPORTANT."
with open("outfile.txt", "a") as file_object:
file_object.write(data_to_save)
# + slideshow={"slide_type": "fragment"}
with open("outfile.txt", "r") as file_object:
contents = file_object.readlines()
print("LINE 1: {}".format(contents[0]))
print("LINE 2: {}".format(contents[1]))
# + [markdown] slideshow={"slide_type": "slide"}
# ## Part 2: Preventing errors
# + [markdown] slideshow={"slide_type": "fragment"}
# This aspect of programming hasn't been very heavily emphasized--that of error handling--because for the most part, data science is about building models and performing computations so you can make inferences from your data.
# + [markdown] slideshow={"slide_type": "fragment"}
# ...except, of course, from nearly every survey that says your average data scientist spends the *vast* majority of their time cleaning and organizing their data.
# + [markdown] slideshow={"slide_type": "fragment"}
# 
# + [markdown] slideshow={"slide_type": "slide"}
# Data is messy and computers are fickle. Just because that file was there yesterday doesn't mean it'll still be there tomorrow. When you're reading from and writing to files, you'll need to put in checks to make sure things are behaving the way you expect, and if they're not, that you're handling things gracefully.
# + [markdown] slideshow={"slide_type": "fragment"}
# We're going to become good friends with `try` and `except` whenever we're dealing with files. For example, let's say I want to read again from that *Alice in Wonderland* file I had:
# + slideshow={"slide_type": "fragment"}
with open("alicee.txt", "r") as file_object:
contents = file_object.readlines()
print(contents[0])
# + [markdown] slideshow={"slide_type": "fragment"}
# Whoops. In this example, I simply misnamed the file. In practice, maybe the file was moved; maybe it was renamed; maybe you're getting the file from the user and they incorrectly specified the name. Maybe the hard drive failed, or any number of other "acts of God." Whatever the reason, **your program should be able to handle missing files.**
# + [markdown] slideshow={"slide_type": "slide"}
# You could probably code this up yourself:
# + slideshow={"slide_type": "fragment"}
filename = "alicee.txt"
try:
with open(filename, "r") as file_object:
contents = file_object.readlines()
print(contents[0])
except FileNotFoundError:
print("Sorry, the file '{}' does not seem to exist.".format(filename))
# + [markdown] slideshow={"slide_type": "fragment"}
# Pay attention to this: this will most likely show up on future assignments / exams, and you'll be expected to properly handle missing files or incorrect filenames.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Part 3: Moving around the filesystem
# + [markdown] slideshow={"slide_type": "fragment"}
# Turns out, you can automate a significant chunk of the double-clicking-around that you do on a Windows machine looking for files. Python has an `os` module that is very powerful.
# + [markdown] slideshow={"slide_type": "fragment"}
# There are a ton of utilities in this module--I encourage you to [check out everything it can do](https://docs.python.org/3.5/library/os.html)--but I'll highlight a few of my favorites here.
# + [markdown] slideshow={"slide_type": "slide"}
# ### `getcwd`
# + [markdown] slideshow={"slide_type": "fragment"}
# This is one of your mainstays: it tells you the full path to where your Python program is currently executing.
# + [markdown] slideshow={"slide_type": "fragment"}
# "cwd" is shorthand for "current working directory."
# + slideshow={"slide_type": "fragment"}
import os
print(os.getcwd())
# + [markdown] slideshow={"slide_type": "slide"}
# ### `chdir`
# + [markdown] slideshow={"slide_type": "fragment"}
# You know where you are using `getcwd`, but you actually want to be somewhere else. `chdir` is the Python equivalent of typing `cd` on the command line, or quite literally double-clicking a folder.
# + slideshow={"slide_type": "fragment"}
print(os.getcwd())
# Go up one directory.
os.chdir("..")
print(os.getcwd())
# + [markdown] slideshow={"slide_type": "slide"}
# ### `listdir`
# + [markdown] slideshow={"slide_type": "fragment"}
# Now you've made it into your directory of choice, but you need to know what files exist. You can use `listdir` to, literally, list the directory contents.
# + slideshow={"slide_type": "fragment"}
for item in os.listdir("."): # A dot "." means the current directory
print(item)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Modifying the filesystem
# + [markdown] slideshow={"slide_type": "fragment"}
# There are a ton of functions at your disposal to actually make changes to the filesystem.
# + [markdown] slideshow={"slide_type": "fragment"}
# - `os.mkdir` and `os.rmdir`: create and delete directories, respectively
# + [markdown] slideshow={"slide_type": "fragment"}
# - `os.remove` and `os.unlink`: delete files (both are equivalent)
# + [markdown] slideshow={"slide_type": "fragment"}
# - `os.rename`: renames a file or directory to something else (equivalent to "move", or "`mv`")
# + [markdown] slideshow={"slide_type": "slide"}
# ### `os.path`
# + [markdown] slideshow={"slide_type": "fragment"}
# The base `os` module has a lot of high-level, basic tools for interacting with the filesystem. If you find that your needs exceed what this module can provide, it has a submodule for more specific filesystem interactions.
# + [markdown] slideshow={"slide_type": "fragment"}
# For instance: testing if a file or directory even exists at all?
# + slideshow={"slide_type": "fragment"}
import os.path
if os.path.exists("/Users/squinn"):
print("Path exists!")
else:
print("No such directory.")
# + slideshow={"slide_type": "fragment"}
if os.path.exists("/something/arbitrary"):
print("Path exists!")
else:
print("No such directory.")
# + [markdown] slideshow={"slide_type": "slide"}
# Once you know a file or directory exists, you can get even more specific: is it a *file*, or a *directory*?
# + [markdown] slideshow={"slide_type": "fragment"}
# Use `os.path.isdir` and `os.path.isfile` to find out.
# + slideshow={"slide_type": "fragment"}
if os.path.exists("/Users/squinn") and os.path.isdir("/Users/squinn"):
print("It exists, and it's a directory.")
else:
print("Something was false.")
# + [markdown] slideshow={"slide_type": "slide"}
# ### `join`
# + [markdown] slideshow={"slide_type": "fragment"}
# This is a relatively unassuming function that is quite possibly the single most useful one; I certainly find myself using it all the time.
# + [markdown] slideshow={"slide_type": "fragment"}
# To illustrate: you're running an image hosting site. You store your images on a hard disk, perhaps at `C:\\images\\`. Within that directory, you stratify by user: each user has their own directory, which has the same name as their username on the site, and all the images that user uploads are stored in their folder.
# + [markdown] slideshow={"slide_type": "fragment"}
# For example, if I was a user and my username was `squinn`, my uploaded images would be stored at `C:\\images\\squinn\\`. A different user, `hunter2`, would have their images stored at `C:\\images\\hunter2\\`. And so on.
# + [markdown] slideshow={"slide_type": "slide"}
# Let's say I've uploaded a new image, `my_cat.png`. I need to stitch a full path together to *move* the image to that path.
# + [markdown] slideshow={"slide_type": "fragment"}
# One way to do it would be hard-coded (hard-core?):
# + slideshow={"slide_type": "fragment"}
img_name = "my_cat.png"
username = "squinn"
base_path = "C:\\images"
full_path = base_path + "\\" + username + "\\" + img_name
print(full_path)
# + [markdown] slideshow={"slide_type": "fragment"}
# That...works. I mean, it works, but it ain't pretty. Also, this will fail **miserably** if you take this code verbatim and run it on a \*nix machine!
# + [markdown] slideshow={"slide_type": "slide"}
# Enter `join`. This not only takes the hard-coded-ness out of the process, but is also **operating system aware**: that is, it will add the needed directory separator for your specific OS, without any input on your part.
# + slideshow={"slide_type": "fragment"}
import os.path
img_name = "my_cat.png"
username = "squinn"
base_path = "C:\\images"
full_path = os.path.join(base_path, username, img_name)
print(full_path)
# + [markdown] slideshow={"slide_type": "fragment"}
# (of course, the slashes are flipped the "wrong" way, because *I'm on a Unix system*, and Python detects that and inserts the correct-facing slashes--but you see how it works in practice!)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Review Questions
#
# Some questions to discuss and consider:
#
# 1: In Part 1 of the lecture when we read in and printed the first few lines of *Alice in Wonderland*, you'll notice each line of text has a blank line in between. Explain why, and how to fix it (so that printing each line shows text of the book the same way you'd see it in the file).
#
# 2: Describe the circumstances under which append "a" mode and write "w" mode are identical.
#
# 3: `os.path.join` can accept any number of file paths. How would it look if you wrote out the function header for `join`?
#
# 4: I'm in a folder, `"/some/folder/"`. I want to create a list of all the `.png` images in the folder. Write a function that will do this.
#
# 5: For whatever reason, I've decided I don't like `os.path.exists` and don't want to use it. How could I structure my code in such a way that I interact with the filesystem at certain paths without checking first if they even exist, while also preventing catastrophic crashes?
# + [markdown] slideshow={"slide_type": "slide"}
# ## Course Administrivia
# + [markdown] slideshow={"slide_type": "fragment"}
# - How is A6 going?
# + [markdown] slideshow={"slide_type": "fragment"}
# - No new assignment this week!
# + [markdown] slideshow={"slide_type": "fragment"}
# - More on the midterm coming soon...
# + [markdown] slideshow={"slide_type": "slide"}
# ## Additional Resources
#
# 1. <NAME>. *Python Crash Course*, Chapter 10. 2016. ISBN-13: 978-1593276034
# 2. <NAME>. *Python for Data Analysis*, Chapter 6. 2013. ISBN-13: 978-1449319793
| lectures/L14.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SageMath 8.1
# language: ''
# name: sagemath
# ---
# + [markdown] deletable=true editable=true
# # 05. Random Variables, Expectations, Data, Statistics, Arrays and Tuples, Iterators and Generators
#
# ## [Inference Theory 1](https://lamastex.github.io/scalable-data-science/infty/2018/01/)
#
# ©2018 <NAME>. [Attribution 4.0 International (CC BY 4.0)](https://creativecommons.org/licenses/by/4.0/)
# + [markdown] deletable=true editable=true
# ### Topics
#
# 1. Continuous Random Variables
# - Expectations
# - Data and Statistics
# - Sample Mean
# - Sample Variance
# - Order Statistics
# - Frequencies
# - Empirical Mass Function
# - Empirical Distribution Function
# - Live Data Ingestion with Try-Catch
# - Arrays
# - Tuples
# - Iterators
# - Generators
#
#
# # Random Variables
#
# A random variable is a mapping from the sample space $\Omega$ to the set of real numbers $\mathbb{R}$. In other words, it is a numerical value determined by the outcome of the experiment.
#
# We already saw *discrete random variables* that take values in a discrete set, of two types:
#
# - those with with finitely many values, eg. the two values in $\{0,1\}$ for the Bernoulli$(\theta)$ RV
# - those with *countably infinitely many* values, eg. values in the set of all non-negative integers: $\{0,1,2,\ldots\}$, for the 'infinite coin tossing experiment' that records the number of consecutive 'Tails' you see before the first 'Heads' occurs.
#
# Now, we will see the other main type of real-valued random variable.
#
# ## Continuous random variable
#
# When a random variable takes on values in the continuum we call it a continuous RV.
#
# ### Examples
#
# - Volume of water that fell on the Southern Alps yesterday (See video link below)
# - Vertical position above sea level, in micrometers, since the original release of a pollen grain at the head waters of a river
# - Resting position in degrees of a roulettet wheel after a brisk spin
#
# ## Probability Density Function
#
# A RV $X$ with DF $F$ is called continuous if there exists a piece-wise continuous function $f$, called the probability density function (PDF) $f$ of $X$, such that, for any $a$, $b \in \mathbb{R}$ with $a < b$,
#
# $$
# P(a < X \le b) = F(b)-F(a) = \int_a^b f(x) \ dx \ .
# $$
#
#
# The following hold for a continuous RV $X$ with PDF $f$:
#
# For any $x \in \mathbb{R}$, $P(X=x)=0$.
# Consequentially, for any $a,b \in \mathbb{R}$ with $a \le b$
# $$P(a < X < b ) = P(a < X \le b) = P(a \leq X \le b) = P(a \le X < b)$$
# By the fundamental theorem of calculus, except possibly at finitely many points (where the continuous pieces come together in the piecewise-continuous $f$):
# $$f(x) = \frac{d}{dx} F(x)$$
# And of course $f$ must satisfy:
# $$\int_{-\infty}^{\infty} f(x) \ dx = P(-\infty < X < \infty) = 1$$
#
#
# ### You try at home
# Watch the Khan Academy [video about probability density functions](https://youtu.be/Fvi9A_tEmXQ) to warm-up to the meaning behind the maths above. Consider the continuous random variable $Y$ that measures the exact amount of rain tomorrow in inches. Think of the probability space $(\Omega,\mathcal{F},P)$ underpinning this random variable $Y:\Omega \to \mathbb{Y}$. Here the sample space, range or support of the random variable $Y$ denoted by $\mathbb{Y} = [0,\infty) =\{y : 0 \leq y < \infty\}$.
#
# + deletable=true editable=true
def showURL(url, ht=500):
"""Return an IFrame of the url to show in notebook with height ht"""
from IPython.display import IFrame
return IFrame(url, width='95%', height=ht)
showURL('https://en.wikipedia.org/wiki/Integral',600)
# + [markdown] deletable=true editable=true
# ## The Uniform$(0,1)$ RV
#
# The Uniform$(0,1)$ RV is a continuous RV with a probability density function (PDF) that takes the value 1 if $x \in [0,1]$ and $0$ otherwise. Formally, this is written
#
#
# $$
# \begin{equation}
# f(x) = \mathbf{1}_{[0,1]}(x) =
# \begin{cases}
# 1 & \text{if } 0 \le x \le 1 ,\\
# 0 & \text{otherwise}
# \end{cases}
# \end{equation}
# $$
#
#
# and its distribution function (DF) or cumulative distribution function (CDF) is:
#
#
# $$
# \begin{equation}
# F(x) := \int_{- \infty}^x f(y) \ dy =
# \begin{cases}
# 0 & \text{if } x < 0 , \\
# x & \text{if } 0 \le x \leq 1 ,\\
# 1 & \text{if } x > 1
# \end{cases}
# \end{equation}
# $$
#
#
# Note that the DF is the identity map in $[0,1]$.
#
# The PDF, CDF and inverse CDF for a Uniform$(0,1)$ RV are shown below
#
# <img src="images/Uniform01ThreeCharts.png" alt="Uniform01ThreeCharts" width=500>
#
# The Uniform$(0,1)$ is sometimes called the Fundamental Model.
#
# The Uniform$(0,1)$ distribution comes from the Uniform$(a,b)$ family.
#
# $$
# \begin{equation}
# f(x) = \mathbf{1}_{[a,b]}(x) =
# \begin{cases}
# \frac{1}{(b-a)} & \text{if } a \le x \le b,\\
# 0 & \text{otherwise}
# \end{cases}
# \end{equation}
# $$
#
# This is saying that if $X$ is a Uniform$(a,b)$ RV, then all values of $x$ between $a$ and $b$, i.e., $a \le x \le b$, are equally probable. The Uniform$(0,1)$ RV is the member of the family where $a=0$, $b=1$.
#
# The PDF and CDF for a Uniform$(a,b)$ RV are shown from wikipedia below
#
# <table style="width:95%">
# <tr>
# <th><img src="https://upload.wikimedia.org/wikipedia/commons/thumb/9/96/Uniform_Distribution_PDF_SVG.svg/500px-Uniform_Distribution_PDF_SVG.svg.png" alt="500px-Uniform_Distribution_PDF_SVG.svg.png" width=250></th>
# <th><img src="https://upload.wikimedia.org/wikipedia/commons/thumb/6/63/Uniform_cdf.svg/500px-Uniform_cdf.svg.png" alt="wikipedia image 500px-Uniform_cdf.svg.png" width=250></th>
# </tr>
# </table>
#
# You can dive deeper into this family of random vaiables <a href="https://en.wikipedia.org/wiki/Uniform_distribution_(continuous)">here</a>.
#
# SageMath has a function for simulating samples from a Uniform$(a,b)$ distribution. We will learn more about this later in the course. Let's go ahead and use it to simulate samples from it below.
# + deletable=true editable=true
uniform(-1,1) # reevaluate the cell to see how the samples change upon each re-evaluation
# + deletable=true editable=true
# + [markdown] deletable=true editable=true
# # Expectations
#
# The *expectation* of $X$ is also known as the *population mean*, *first moment*, or *expected value* of $X$.
#
# $$
# \begin{equation}
# E\left(X\right) := \int x \, dF(x) =
# \begin{cases}
# \sum_x x \, f(x) & \qquad \text{if }X \text{ is discrete} \\
# \int x \, f(x)\,dx & \qquad \text{if } X \text{ is continuous}
# \end{cases}
# \end{equation}
# $$
#
# Sometimes, we denote $E(X)$ by $E X$ for brevity. Thus, the expectation is a single-number summary of the RV $X$ and may be thought of as the average.
#
# In general though, we can talk about the Expectation of a function $g$ of a RV $X$.
#
# The Expectation of a function $g$ of a RV $X$ with DF $F$ is:
#
# $$
# \begin{equation}
# E\left(g(X)\right) := \int g(x)\,dF(x) =
# \begin{cases}
# \sum_x g(x) f(x) & \qquad \text{if }X \text{ is discrete} \\
# \int g(x) f(x)\,dx & \qquad \text{if } X \text{ is continuous}
# \end{cases}
# \end{equation}
# $$
#
#
# provided the sum or integral is well-defined. We say the expectation exists if
#
#
# $$
# \begin{equation}
# \int \left|g(x)\right|\,dF(x) < \infty \ .
# \end{equation}
# $$
#
# When we are looking at the Expectation of $X$ itself, we have $g(x) = x$
#
# Thinking about the Expectations like this, can you see that the familiar Variance of X is in fact the Expection of $g(x) = (x - E(X))^2$?
#
# The variance of $X$ (a.k.a. second moment)
#
# Let $X$ be a RV with mean or expectation $E(X)$. The variance of $X$ denoted by $V(X)$ or $VX$ is
#
# $$
# V(X) := E\left((X-E(X))^2\right) = \int (x-E(X))^2 \,d F(x)
# $$
#
# provided this expectation exists. The standard deviation denoted by $\sigma(X) := \sqrt{V(X)}$.
#
# Thus variance is a measure of ``spread'' of a distribution.
#
# The $k$-th moment of a RV comes from the Expectation of $g(x) = x^k$.
#
# We call
#
# $$
# E(X^k) = \int x^k\,dF(x)
# $$
#
#
# the $k$-th moment of the RV $X$ and say that the $k$-th moment exists when $E(|X|^k) < \infty$.
#
#
# ## Properties of Expectations
#
#
#
# 1. If the $k$-th moment exists and if $j<k$ then the $j$-th moment exists.
# - If $X_1,X_2,\ldots,X_n$ are RVs and $a_1,a_2,\ldots,a_n$ are constants, then $E \left( \sum_{i=1}^n a_i X_i \right) = \sum_{i=1}^n a_i E(X_i)$
# - Let $X_1,X_2,\ldots,X_n$ be independent RVs, then
# - $E \left( \prod_{i=1}^n X_i \right) = \prod_{i=1}^{n} E(X_i)$
# - $V(X) = E(X^2) - (E(X))^2$
# - If $a$ and $b$ are constants, then: $V \left(aX + b\right) = a^2V(X) $
# - If $X_1,X_2,\ldots,X_n$ are independent and $a_1,a_2,\ldots,a_n$ are constants, then: $V \left( \sum_{i=1}^n a_i X_i \right) = \sum_{i=1}^n a_i^2 V(X_i)$
#
# ### You try at home
#
# Watch the Khan Academy videos about [probability density functions](https://youtu.be/Fvi9A_tEmXQ) and [expected value](https://youtu.be/j__Kredt7vY) if you want to get another angle on the material more slowly step-by-step:
# + [markdown] deletable=true editable=true
# ## The population mean and variance of the Bernoulli$(\theta)$ RV
#
# We have already met the discrete Bernoulli$(\theta)$ RV. Remember, that if we have an event $A$ with $P(A) = \theta$, then a Bernoulli$(\theta)$ RV $X$ takes the value $1$ if "$A$ occurs" with probability $\theta$ and $0$ if "$A$ does not occur" with probability $1-\theta$.
#
# In other words, the indicator function $\mathbf{1}_A$ of "$A$ occurs" with probability $\theta$ is the Bernoulli$(\theta)$ RV.
#
# For example, flip a fair coin.
# Consider the event that it turns up heads. Since the coin is fair, the probability of this event $\theta$ is $\frac{1}{2}$. If we define an RV $X$ that takes the value 1 if the coin turns up heads ("event coin turns up heads occurs") and 0 otherwise, then we have a Bernoulli$(\theta = \frac{1}{2})$ RV.
#
# We all saw that given a parameter $\theta \in [0,1]$, the probability mass function (PMF) for the Bernoulli$(\theta)$ RV $X$ is:
#
# $$
# f(x;\theta) = \theta^x (1-\theta)^{1-x} \mathbf{1}_{\{0,1\}}(x) =
# \begin{cases}
# \theta & \text{if } x=1 \ ,\\
# 1-\theta & \text{if } x=0 \ ,\\
# 0 & \text{otherwise}
# \end{cases}
# $$
#
#
# and its DF is:
#
# $$
# \begin{equation}
# F(x;\theta) =
# \begin{cases}
# 1 & \text{if } 1 \le x \ ,\\
# 1-\theta & \text{if } 0 \le x < 1 \ ,\\
# 0 & \text{otherwise}
# \end{cases}
# \end{equation}
# $$
#
# Now let's look at some expectations: the population mean and variance of an RV $X \thicksim$ Bernoulli$(\theta)$.
#
# Because $X$ is a discrete RV, our expectations use sums rather than integrals.
#
# The first moment or expectation is:
#
# $$
# \begin{array}{lcl}
# E(X) & = & \displaystyle\sum_{x=0}^{1}xf(x;\theta) \\ &=& (0 \times (1-\theta)) + (1 \times \theta)\\ &=& 0 + \theta\\ &=& \theta\\
# \end{array}
# $$
#
# The second moment is:
# $$\begin{array}{lcl}
# E(X^2) &=& \displaystyle\sum_{x=0}^{1}x^2f(x;\theta) \\
# &=& (0^2 \times (1-\theta)) + (1^2 \times \theta)\\ &=& 0 + \theta\\ &=& \theta
# \end{array}
# $$
#
# The variance is:
# $$
# \begin{array}{lcl} V(X) &=& E(X^2) - \left(E(X)\right)^2\\ &=& \theta - \theta^2\\ &=& \theta(1-\theta)
# \end{array}
# $$
#
# We can see that $E(X)$ and $V(X)$ will vary with the parameter $\theta$. This is why we subscript $E$ and $V$ with $\theta$, to emphasise that the values depend on the parameter.
#
# $$E_{\theta}(X) = \theta$$
#
# $$V_{\theta}(X) = \theta(1-\theta)$$
#
# We can use Sage to do a simple plot to see how $E_{\theta}(X)$ and $V_{\theta}(X)$ vary with $\theta$.
# + deletable=true editable=true
def bernoulliPopMean(th):
'''A function to find the population mean for an RV distributed Bernoulli(theta).
parameter th is the distribution parameter theta.'''
return th
def bernoulliPopVariance(th):
'''A function to find the population variance for an RV distributed Bernoulli(theta).
parameter th is the distribution parameter theta.'''
return th*(1-th)
# + deletable=true editable=true
# ?points
# + deletable=true editable=true
# using a list comprehension into Geometric primitive of 'points' we can plot as follows
p = points([(x*0.01,bernoulliPopMean(x*0.01)) for x in range(0,100)], rgbcolor="blue") # assign the plot to p
p += points([(x*0.01,bernoulliPopVariance(x*0.01)) for x in range(0,100)], rgbcolor="red") # add variance plot to p
show(p, figsize=[6,3])
# + [markdown] deletable=true editable=true
# Note how the variance is maximized at $\theta=\frac{1}{2}$.
# + [markdown] deletable=true editable=true
# ## The population mean and variance of the Uniform$(0,1)$ RV
#
# Now let's look at the the population mean and variance of a continuous RV $X \thicksim$ Uniform$(0,1)$.
#
# Because $X$ is a continuous RV, our expectations use integrals.
#
# $$
# \begin{array}{lcl} E(X) &=&\int_{x=0}^1 x f(x)\, dx\\ &=& \int_{x=0}^1 x \ 1 \, dx\\ &=& \frac{1}{2} \left( x^2 \right]_{x=0}^{x=1}\\ &=& \frac{1}{2} \left( 1-0 \right)\\ &=& \frac{1}{2} \end{array}
# $$
#
#
#
# $$
# \begin{array}{lcl} E(X^2) &=& \int_{x=0}^1 x^2 f(x)\, dx \\ &=& \int_{x=0}^1 x^2 \ 1 \, dx\\ &=& \frac{1}{3} \left( x^3 \right]_{x=0}^{x=1}\\ &=& \frac{1}{3} \left( 1-0 \right)\\ &=& \frac{1}{3}\\ \end{array}
# $$
#
# $$
# \begin{array}{lcl} V(X) &=& E(X^2) - \left(E(X)\right)^2\\ &=&\frac{1}{3} - \left( \frac{1}{2} \right)^2\\ &=& \frac{1}{3} - \frac{1}{4}\\ &=& \frac{1}{12} \end{array}
# $$
#
#
#
# ### Winnings on Average
#
# Think about playing a game where we draw $x \thicksim X$ and I pay you $r(x)$ ($r$ is some reward function, a function of $x$ that says what your reward is when $x$ is drawn). Then, your average winnings from the game is the sum (or integral), over all the possible values of $x$, of $r(x) \times$ the chance that $X=x$.
#
# Put formally, if $Y= r(X)$, then
#
# $$E(Y) = E(r(X)) = \int r(x) \,dF(x)$$
# + [markdown] deletable=true editable=true
# ## Probability is an Expectation
#
# Remember when we first talked about the probability of some event $A$, we talked about the idea of the probability of $A$ as the long term relative frequency of $A$?
#
# Now consider some event $A$ and a reward function $r(x) = \mathbf{1}_A(x)$.
#
# Recall that $\mathbf{1}_A(x) = 1$ if $ x \in A$ and $\mathbf{1}_A(x) = 0$ if $ x \notin A$: the reward is 1 if $x \in A$ and 0 otherwise.
#
# $$
# \begin{array}{lcl} \text{If } X \text{ is continuous } E(\mathbf{1}_A(X)) &=& \int \mathbf{1}_A(x)\, dF(x)\\ &=& \int_A f(x)\, dx\\ &=& P(X \in A) = P(A)\\ \text{If } X \text{ is discrete } E(\mathbf{1}_A(X)) &=& \mathbf{1}_A(x)\, dF(x)\\ &=& \sum_{x \in A} f(x)\\ &=& P(X \in A) = P(A) \\ \end{array}
# $$
#
# This says that probability is a special case of expectation: the probability of $A$ is the expectation that $A$ will occur.
#
# Take a Uniform$(0,1)$ RV $X$. What would you say the probability that an observation of this random variable is $\le 0.5$ is, ie what is $P(X \le 0.5)$?
#
# Let's use SageMath to simulate some observations for us and look at the relative frequency of observations $\le 0.5$:
# + deletable=true editable=true
uniform(0,1) # remember calling this each time changes the outcome - reevaluate this cell and see
# + deletable=true editable=true
countObOfInterest = 0 # variable to count observations of interest
numberOfObs = 1000 # variable to control how many observations we simulate
obOfInterest = 0.5 # variable for observation of interest
for i in range(numberOfObs): # loop to simulate observations
if uniform(0,1) <= obOfInterest: # conditional statement to check observation
countObOfInterest += 1 # accumulate count of observation of interest
print "The relative frequency of x <=", obOfInterest.n(digits=2), \
" was", RR(countObOfInterest/numberOfObs).n(digits=3) # just formatting out print output
# + [markdown] deletable=true editable=true
# Or, we could look at a similar simulation for a discrete RV, say a Bernoulli$(\frac{1}{2})$ RV.
#
# Another way of thinking about the Bernoulli$(\frac{1}{2})$ RV is that it has a discrete uniform distribution over $\{0,1\}$. It can take on a finite number of values (0 and 1 only) and the probabilities of observing either of these two values are are equal.
#
# This could be modelling the event that we get a head when we throw a fair coin. For this we'll use the `randint(0,1)` function to simulate the observed value of our RV $X$.
# + deletable=true editable=true
randint(0,1) # try again and again
# + deletable=true editable=true
countObOfInterest = 0 # variable to count observations of interest
numberOfObs = 1000 # variable to control how many observations we simulate
obOfInterest = 1 # variable for observation of interest
for i in range(numberOfObs): # loop to simulate observations
if randint(0,1) == obOfInterest: # conditional statement to check observation
countObOfInterest += 1 # accumulate count of observation of interest
print "The relative frequency of x ==", obOfInterest, \
" was", RR(countObOfInterest/numberOfObs).n(digits=3) # just formatting out print output
# + [markdown] deletable=true editable=true
# ## The de Moivre$(\frac{1}{k}, \frac{1}{k}, \ldots, \frac{1}{k})$ RV
#
# We have seen that a Bernoulli$(\theta)$ RV has two outcomes (0, and 1). What if we are interested in modelling situations where there are more than two outcomes of interest? For example, we could use a $Bernoulli(\frac{1}{2})$ RV to model whether the outcome of the flip of a fair coin is a head, but we can't use it for modelling a RV which is the number we get when we toss a six-sided die.
#
# So, now, we will consider a natural generalization of the Bernoulli$(\theta)$ RV with more than two outcomes. This is called the de Moivre$(\frac{1}{k}, \frac{1}{k}, \ldots, \frac{1}{k})$ RV (after Abraham de Moivre, 1667-1754), one of the first great analytical probabalists).
#
# A de Moivre$(\frac{1}{k}, \frac{1}{k}, \ldots, \frac{1}{k})$ RV $X$ has a discrete uniform distribution over $\{1, 2, ..., k\}$: there are $k$ possible equally probable ('equiprobable') values that the RV can take.
#
# If we are rolling a die and $X$ is the number on die, then $k=6$.
#
# Or think of the New Zealand Lotto game. There are 40 balls in the machine, numbered $1, 2, \ldots, 40$. The number on the first ball out of the machine can be modelled as a de Moivre$(\frac{1}{40}, \frac{1}{40}, \ldots, \frac{1}{40})$ RV.
#
# We say that an RV $X$ is de Moivre$(\frac{1}{k}, \frac{1}{k}, \ldots, \frac{1}{k})$ distributed if its probability mass function PMF is:
#
# $$
# f \left(x; \left( \frac{1}{k}, \frac{1}{k}, \ldots, \frac{1}{k} \right) \right) = \begin{cases} 0 & \quad \text{if } x \notin \{1,2,\ldots,k\}\\ \frac{1}{k} & \quad \text{if } x \in \{1,2,\ldots,k\} \end{cases}
# $$
#
# We can find the expectation:
# $$
# \begin{array}{lcl} E(X) & = & \sum_{x=1}^k xP(X=x)\\ &=& (1 \times \frac{1}{k}) + (2 \times \frac{1}{k}) + \ldots + (k \times \frac{1}{k})\\ &=& (1 + 2 + \dots + k)\frac{1}{k}\\ &=& \frac{k(k+1)}{2}\frac{1}{k}\\ &=& \frac{k+1}{2} \, , \end{array}
# $$
#
# the second moment:
# $$
# \begin{array}{lcl} E(X^2) & =& \sum_{x=1}^k x^2P(X=x)\\ & =& (1^2 \times \frac{1}{k}) + (2^2 \times \frac{1}{k}) + \ldots + (k^2 \times \frac{1}{k})\\ &=& (1^2 + 2^2 + \dots + k^2)\frac{1}{k}\\ &=& \frac{k(k+1)(2k+1)}{6}\frac{1}{k}\\ &=& \frac{2k^2+3k+1}{6}\, , \end{array}
# $$
#
# and finally the variance:
# $$
# \begin{array}{lcl} V(X) &=& E(X^2) - \left(E(X)\right)^2\\ &=& \frac{2k^2+3k+1}{6} - \left( \frac{k+1}{2} \right)^2\\ &=&\frac{2k^2+3k+1}{6} - \frac{k^2 + 2k +1}{4}\\ &=& \frac{4(2k^2 + 3k + 1) - 6(k^2 + 2k + 1) }{24}\\ &=& \frac{8k^2 + 12k + 4 - 6k^2 - 12k - 6 }{24}\\ &=& \frac{2k^2-2}{24} \\ &=& \frac{k^2-1}{12} \, . \end{array}
# $$
#
# We coud use the Sage `randint` function to simulate the number on the first ball in a Lotto draw:
# + deletable=true editable=true
myList=[]
nn=10000
for i in range(nn):
myList.append(randint(1,40))
len([x for x in myList if x==20])/RR(nn)
# + deletable=true editable=true
1/40.
# + [markdown] deletable=true editable=true
# # Statistics
#
# The official NZ Government site for statistics about New Zealand is http://www.stats.govt.nz/ and that for Sweden is http://www.scb.se/en/About-us/official-statistics-of-sweden/
#
# Take a tour through it!
#
# ## Data and statistics
#
# In general, given some probability triple $(\Omega, \mathcal{F}, P)$, let the function $X$ measure the outcome $\omega$ from the sample space $\Omega$.
#
# $$X(\omega): \Omega \rightarrow \mathbb{X}$$
#
# The RV $X$ is called **data**, if $X$ is the finest available measurement of $\Omega$.
#
# $\mathbb{X}$ is called the data space (sample space of the data $X$).
#
# $X(\omega)=x$ is the outcome $\omega$ measured by $X$ and is called the observed data or the realisation of $X$.
#
# Often the measurements made by $X$ are real numbers or structures of real numbers or structures that are a mixture of combinatorial objects and real numbers (they typically arise from more general data objects/structures, like addresses, images, videos, trajectories, networks, etc.).
#
# Recall the definition of a real-valued or $\mathbb{R}$-valued random variable we have already seen (an $\mathbb{R}$-valued random variable $X$ is a specific function or map from the sample space to the real line $\mathbb{R}$). We have also seen that $X$ can in fact be a real-vector-valued or $\mathbb{R}^n$-valued random variable $X=(X_1,X_2,\ldots,X_n)$, i.e. a vector of $\mathbb{R}$-valued random variables or simply a random vector. A random variable can be much more general as we will see in the sequel - it can be image-valued, movie-valued, mp3-song-valued, graph-valued, etc.
#
# When we talked about an experiment involving two IID Bernoulli$(\frac{1}{2})$ RVs above (tossing two coins), we listed the different results we might get as (0,0), (1,0), (0,1), (1,1).
#
# Say we observe the outcome $\omega$ = (H, H) (two heads). Our $Bernoulli$ random vector $X$ measures this as (1,1). Thus, (1,1) is the observed data or the realisation of $X$.
#
# ## So what is a statistic?
#
# A *statistic* is any *[measureable function](https://en.wikipedia.org/wiki/Measurable_function)* of the data: $T(x): \mathbb{X} \rightarrow \mathbb{T}$ or simply that $\mathbb{X} \overset{T}{\rightarrow} \mathbb{T}$.
#
# Thus, a statistic $T$ is also an RV that takes values in the space $\mathbb{T}$.
#
# When $x \in \mathbb{X}$ is the observed data, $T(x)=t$ is the observed statistic of the observed data $x$, i.e., $x \mapsto t$.
#
# ### Question: Can Statistic be the Data?
#
# <p style="color:#F8F9F9";>Consider the identity map... with $T(x)=x$ and $\mathbb{X} = \mathbb{T}$.</p>
#
# #### Remember $\Omega$ is always under the hood!
#
# $$
# \Omega \overset{X}{\rightarrow} \mathbb{X} \overset{T}{\rightarrow} \mathbb{T}, \quad \text{with} \quad \omega \mapsto x \mapsto t.
# $$
#
# Generally, one cannot explicity work with $\Omega$ (recall $\Omega$ for the indicator random variable of *rain on Southern Alps tomorrow!*) but can work with the following axiomatic cascade towards more concretely measurable events:
#
# - observable events in a sigma-algebra $\mathcal{F}(\Omega)$ or $\mathcal{F}_{\Omega}$ on $\Omega$
# - that are captured by the RV $X$ as measurable sets in $\mathcal{F}(\mathbb{X})$, the sigma-algebra over the data space $\mathbb{X}$
# - and typically projected further by the statistic $T$, another RV that measurable maps the sigm-algebra $\mathcal{F}(\mathbb{X})$ to the sigma-algebra $\mathcal{F}(\mathbb{T})$.
#
#
# + [markdown] deletable=true editable=true
# ## Example 2: New Zealand Lotto Data and a Probability Model of it as IID de Moivre RVs
#
# ### New Zealand lotto data
# Let's look at our pre-made New Zealand lotto data. This is the winning first ball drawn from a NZ Lotto machine for several years.
# + deletable=true editable=true
# These lotto draws of the first ball from NZ Lotto was already downloaded and processed for you
listBallOne = [4, 3, 11, 35, 23, 12, 14, 13, 15, 19, 36, 18, 37, 39, 37, 35, 39, 1, 24, 29, 38, 18, 40, 35, \
12, 7, 14, 23, 21, 35, 14, 32, 19, 2, 1, 34, 39, 29, 7, 20, 2, 40, 28, 4, 30, 34, 20, 37, 9, 24,\
36, 4, 22, 1, 31, 12, 16, 29, 36, 5, 21, 23, 30, 39, 38, 22, 13, 6, 14, 30, 40, 21, 5, 12, 28, 27,\
13, 18, 19, 23, 2, 10, 37, 31, 40, 4, 25, 4, 17, 6, 34, 26, 38, 35, 3, 38, 14, 40, 3, 30, 21, 4,\
24, 34, 27, 14, 25, 18, 21, 1, 25, 39, 18, 40, 18, 11, 5, 37, 33, 26, 29, 26, 36, 33, 18, 32, 3, 1,\
5, 22, 39, 25, 12, 21, 23, 12, 31, 1, 35, 8, 32, 24, 34, 14, 26, 4, 3, 31, 17, 22, 24, 10, 29, 40,\
4, 8, 26, 11, 8, 18, 25, 22, 8, 30, 10, 14, 32, 14, 5, 35, 3, 32, 40, 17, 39, 7, 21, 4, 35, 9, 16,\
30, 30, 11, 28, 22, 38, 5, 16, 27, 16, 23, 22, 1, 27, 32, 30, 24, 32, 29, 11, 3, 26, 19, 22, 25, 3,\
34, 31, 17, 16, 31, 20, 29, 10, 2, 17, 36, 6, 34, 11, 7, 22, 28, 13, 15, 20, 39, 16, 10, 25, 1, 37,\
14, 28, 35, 20, 39, 3, 39, 20, 40, 6, 20, 17, 26, 27, 4, 24, 40, 16, 24, 7, 8, 25, 16, 15, 8, 29, 13,\
16, 39, 2, 24, 24, 23, 24, 37, 39, 40, 5, 11, 13, 6, 24, 1, 5, 7, 15, 38, 3, 35, 10, 22, 19, 3, 21,\
39, 38, 4, 30, 17, 15, 9, 32, 28, 7, 12, 6, 37, 25, 4, 8, 30, 7, 31, 12, 21, 31, 13, 2, 20, 14, 40,\
32, 23, 10, 1, 35, 35, 32, 16, 25, 13, 20, 33, 27, 2, 26, 12, 5, 34, 20, 7, 34, 38, 20, 8, 5, 11, 17,\
10, 36, 34, 1, 36, 6, 7, 37, 22, 33, 7, 32, 18, 8, 1, 37, 25, 35, 29, 23, 11, 19, 7, 21, 30, 23, 12,\
10, 26, 21, 9, 9, 25, 2, 14, 16, 14, 25, 40, 8, 28, 19, 8, 35, 22, 23, 27, 31, 36, 22, 33, 22, 15, 3,\
37, 8, 2, 22, 39, 3, 6, 13, 33, 18, 37, 28, 3, 17, 8, 2, 36, 1, 14, 38, 5, 31, 34, 16, 37, 2, 40, 14,\
16, 21, 40, 5, 21, 24, 24, 38, 26, 38, 33, 20, 25, 7, 33, 12, 22, 34, 34, 20, 38, 12, 20, 7, 28, 26,\
30, 13, 40, 36, 29, 11, 31, 15, 9, 13, 17, 32, 18, 9, 24, 6, 40, 1, 1, 9, 13, 28, 19, 5, 7, 27, 12,\
3, 34, 26, 20, 28, 28, 25, 21, 23, 6, 15, 19, 30, 10, 13, 8, 11, 38, 7, 33, 12, 16, 11, 40, 25, 32,\
34, 1, 32, 31, 33, 15, 39, 9, 25, 39, 30, 35, 20, 34, 3, 30, 17, 24, 20, 15, 10, 25, 6, 39, 19, 20,\
23, 16, 17, 31, 25, 8, 17, 15, 31, 20, 19, 33, 11, 37, 31, 4, 12, 37, 7, 40, 8, 22, 3, 25, 35, 8, 9,\
14, 13, 33, 4, 2, 1, 31, 24, 8, 13, 19, 34, 10, 32, 35, 28, 11, 10, 31, 25, 8, 6, 13, 33, 19, 35, 19,\
8, 21, 10, 40, 36, 16, 27, 31, 1, 18, 36, 40, 18, 37, 18, 24, 33, 34, 31, 6, 10, 24, 8, 7, 24, 27, 12,\
19, 23, 5, 33, 20, 2, 32, 33, 6, 13, 5, 25, 7, 31, 40, 1, 30, 37, 19, 27, 40, 28, 3, 24, 36, 7, 22,\
20, 21, 36, 38, 15, 11, 37, 21, 4, 13, 9, 12, 13, 34, 30, 8, 23, 40, 4, 13, 6, 4, 22, 35, 2, 35, 20,\
9, 28, 9, 13, 33, 19, 5, 38, 24, 18, 37, 10, 25, 25, 31, 3, 13, 25, 35, 1, 36, 21, 3, 22, 23, 7, 6,\
26, 11, 6, 1, 24, 2, 25, 38, 3, 16, 16, 20, 22, 12, 8, 27, 38, 10, 39, 9, 37, 30, 33, 12, 4, 32, 2,\
29, 6, 34, 2, 3, 12, 9, 1, 22, 40, 38, 9, 18, 40, 17, 5, 17, 26, 17, 26, 6, 7, 18, 10, 27, 24, 39, 1,\
3, 26, 38, 2, 12, 5, 7, 38, 2, 8, 30, 35, 18, 19, 29, 37, 5, 27, 35, 40, 14, 25, 15, 20, 32, 22, 9, 1,\
8, 14, 38, 27, 23, 24, 15, 29, 7, 4, 19, 6, 21, 27, 23, 21, 35, 32, 13, 27, 34, 1, 11, 36, 24, 23, 13,\
2, 33, 25, 18, 1, 10, 5, 27, 1, 36, 36, 11, 3, 31, 30, 31, 39, 7, 21, 25, 28, 38, 2, 3, 40, 10, 40,\
12, 22, 20, 16, 14, 30, 16, 19, 33, 32, 30, 19, 36, 16, 27, 7, 18, 38, 14, 14, 33, 29, 24, 21, 22, 15,\
25, 27, 25, 37, 35, 34, 11, 19, 35, 10, 30, 8, 11, 20, 7, 27, 19, 16, 21, 13, 6, 29, 35, 13, 31, 23,\
26, 10, 18, 39, 38, 5, 16, 33, 21, 31, 21, 23, 32, 35, 2, 24, 11, 25, 30, 7, 18, 32, 38, 22, 27, 2, 6,\
31, 24, 34, 33, 15, 39, 21, 9, 1, 8, 38, 37, 40, 14, 2, 25, 30, 16, 6, 36, 27, 28, 8, 17, 37, 15, 29,\
27, 30, 30, 19, 15, 13, 34, 5, 24, 18, 40, 37, 1, 28, 17, 32, 8, 34, 5, 6, 31, 8, 9, 28, 26, 40, 40,\
9, 23, 36, 28, 24, 33, 18, 36, 6, 22, 29, 6, 6, 25, 15, 29, 18, 38, 20, 26, 30, 17, 30, 32, 33, 19,\
10, 29, 25, 24, 19, 28, 38, 3, 24, 12, 28, 29, 29, 20, 12, 11, 12, 21, 11, 24, 36, 3, 3, 5, 28, 2,\
8, 30, 23, 4, 40, 28, 6, 31, 37, 25, 9, 23, 20, 20, 16, 38, 21, 35, 18, 3, 15, 40, 19, 33, 34, 20,\
3, 11, 34, 35, 10, 32, 23, 10, 29, 13, 12, 6, 30, 7, 5, 4, 29, 22, 22, 2, 26, 24, 7, 13, 26, 27, 27,\
15, 12, 18, 38, 33, 4, 11, 20, 33, 21, 5, 26, 10, 22, 36, 3, 4, 35, 35, 16, 32, 5, 19, 23, 24, 40,\
25, 30, 10, 9, 23, 12, 40, 21, 29, 18, 17, 15, 32, 2, 35, 7, 30, 4, 2, 16, 6, 8, 35]
# + deletable=true editable=true
len(listBallOne) # there are 1114 draws from
# + [markdown] deletable=true editable=true
# ### Data Space
# The data space is every possible sequence of ball numbers that we could have got in these 1114 draws. $\mathbb{X} = \{1, 2, \ldots, 40\}^{1114}$. There are $40^{1114}$ possible sequences and
#
# ### Probability model
# We can think of this list `listBallOne` of sample size $n=1114$ as $x = (x_1,x_2,\ldots,x_{1114})$, the realisation of a random vector $X = (X_1, X_2,\ldots, X_{1114})$ where $X_1, X_2,\ldots, X_{1114} \overset{IID}{\thicksim} de Moivre(\frac{1}{40}, \frac{1}{40}, \ldots, \frac{1}{40})$.
#
# $P \left( (X_1, X_2, \ldots, X_{1114}) = (x_1, x_2, \ldots, x_{1114}) \right) = \frac{1}{40} \times \frac{1}{40} \times \ldots \times \frac{1}{40} = \left(\frac{1}{40}\right)^{1114}$ if $(x_1, x_2, \ldots, x_{1114}) \in \mathbb{X} = \{1, 2, \ldots, 40\}^{1114}$
#
# Our data is just one of the $40^{1114}$ possible points in this data space.
#
# ## Some statistics
#
# ### Sample mean
#
# From a given sequence of RVs $X_1, X_2, \ldots, X_n$, or a random vector $X = (X_1, X_2, \ldots, X_n)$, we can obtain another RV called the sample mean (technically, the $n$-sample mean):
#
# $$T_n((X_1, X_2, \ldots, X_n)) = \bar{X_n}((X_1, X_2, \ldots, X_n)) :=\frac{1}{n}\displaystyle\sum_{i=1}^nX_i$$
#
# We write $\bar{X_n}((X_1, X_2, \ldots, X_n))$ as $\bar{X}_n$,
#
# and its realisation $\bar{X_n}((x_1, x_2, \ldots, x_n))$ as $\bar{x_n}$.
#
# By the properties of expectations that we have seen before,
#
# $$E(\bar{X_n}) = E \left(\frac{1}{n}\sum_{i=1}^nX_i \right) = \frac{1}{n}E\left(\sum_{i=1}^nX_i\right) = \frac{1}{n}\sum_{i=1}^nE(X_i)$$
#
# And because every $X_i$ is identically distributed with the same expectation, say $E(X_1)$, then
#
# $$E(\bar{X_n}) = \frac{1}{n}\sum_{i=1}^nE(X_i)= \frac{1}{n} \times n \times E(X_1) = E(X_1)$$
#
# Similarly, we can show that,
#
# $$V(\bar{X_n}) = V\left(\frac{1}{n}\sum_{i=1}^nX_i\right) = \frac{1}{n^2}V\left(\sum_{i=1}^nX_i\right)$$
#
# And because every $X_i$ is independently and identically distributed with the same variance, say $V(X_1)$, then
#
# $$V(\bar{X_n}) = \frac{1}{n^2}V\left(\sum_{i=1}^nX_i \right) = \frac{1}{n^2} \times n \times V(X_1) = \frac{1}{n} V(X_1)$$
#
# ### Sample variance
#
# Sample variance is given by $$ \frac{1}{n}\sum_{i=1}^n \left( X_i - \bar{X}_n \right)^2$$ Sometimes, we divide by $n-1$ instead of $n$. It is a measure of spread from the sample.
#
# Similarly, we can look at a sample standard deviation = $\sqrt{\text{sample variance}}$
#
# ### numpy for numerical Python
# Python has a nice module called `numpy` for numerical computing akin to MATLAB. We will import `numpy` to use some of its basic statistical capabilities for our Lotto data. To be able to use these capabilities, it is easiest to convert our Lotto Data into a type called a `numpy.array` or `np.array` if we `import numpy as np`. (You will be looking more closely at `numpy` `array`s in the You Try sections below).
# + deletable=true editable=true
import numpy as np
#import array, mean, var, std # make pylab stuff we need accessible in Sage
arrayBallOne = np.array(listBallOne) # make the array out of the list
arrayBallOne # disclose the list
# + deletable=true editable=true
arrayBallOne. # hit the Tab after the . and see the available methods on numpy arrays
# + [markdown] deletable=true editable=true
# Now we can get the some sample statistics for the lotto ball one data.
#
# ### Sample mean of NZ Lotto data
# + deletable=true editable=true
arrayBallOne.mean() # this calls the mean method and gives our first statistic, the sample mean
# + deletable=true editable=true
arrayBallOne.mean # if you just call the method without '()' then you are not evaluating it
# + [markdown] deletable=true editable=true
# ### Sample vairance and sample standard deviation of NZ Lotto data
# + deletable=true editable=true
arrayBallOne.var()
# + deletable=true editable=true
arrayBallOne.std()
# + deletable=true editable=true
sqrt(arrayBallOne.var()) # just checking std is indeed sqrt(var)
# + [markdown] deletable=true editable=true
# ## Order statistics
#
# The $k$th order statistic of a sample is the $k$th smallest value in the sample. Order statistics that may be of particular interest include the smallest (minimum) and largest (maximum) values in the sample.
# + deletable=true editable=true
arrayBallOne.min() # sample minimum statistic
# + deletable=true editable=true
arrayBallOne.max() # sample maximum statistic
# + deletable=true editable=true
arrayBallOneSorted = np.array(listBallOne) # make the array out of the list for sorting next
arrayBallOneSorted.sort() # sort the array to get the order statistic - this sorts arrayBallOneSorted
# + deletable=true editable=true
arrayBallOneSorted # the sorted data array - numpy automatically displays with intermediates as ...
# + deletable=true editable=true
arrayBallOne # the original data array
# + [markdown] deletable=true editable=true
# ## Frequencies
#
# The next most interesting statistics we will visit is the frequencies of the ball numbers.
#
# With lots of discrete data like this, we may be interested in the frequency of each value, or the number of times we get a particular value. This can be formalized as the following process of adding indicator functions of the data points $(X_1,X_2,\ldots,X_n)$: $$ \sum_{i=1}^n \mathbf{1}_{\{X_i\}}(x) $$
#
# We can use the SageMath/Python dictionary to give us a mapping from ball numbers in the list to the count of the number of times each ball number comes up.
#
# ### A Function to Count Elements in a Sequence
#
# Although we are doing this with the Lotto data, mapping a list like this seems like a generally useful thing to be able to do, and so we write it as a function `makeFreqDict(myDataSeq)` and then use the function on our Lotto data. This is a good example of reusing useful code through a function. We will rely on this function in the sequel for other input data.
#
# We can use `long` integers if we need to count a lot of items or the usual `int` type in Python to count the elements in `myDataSeq`. We can use the default value for the second parameter named `one` to be of type `int` by the following second argument with **default parameter value**: `one=int(1)` to `makeFreqDict`. This will enable us to specify `one=long(1)` if we want the counting to be done in `long` integers if necessary. See [https://docs.python.org/2.0/ref/function.html](https://docs.python.org/2.0/ref/function.html) for default parameters in functions.
#
# Also Python's integer types are supposed to get as big as one needs according to this discussion:
#
# - [https://stackoverflow.com/questions/9860588/maximum-value-for-long-integer](https://stackoverflow.com/questions/9860588/maximum-value-for-long-integer)
#
# We will still pass in the additional argument for pedagogical reasons and to avoid the more heavy-duty SageMath's type of `'sage.rings.integer.Integer'`.
# + deletable=true editable=true
# Note that long is a Python long integer
myLongInteger = long(1)
print(myLongInteger)
type(myLongInteger)
# + deletable=true editable=true
type(1)
# + deletable=true editable=true
def makeFreqDict(myDataSeq, one = int(1)):
'''Make a frequency mapping out of a sequence of data - list, array, str.
Param myDataList, a list of data.
Return a dictionary mapping each unique data value to its frequency count.'''
freqDict = {} # start with an empty dictionary
for res in myDataSeq:
if res in freqDict: # the data value already exists as a key
freqDict[res] = freqDict[res] + one #int(1) # add 1 to the count
else: # the data value does not exist as a key value
freqDict[res] = one # add a new key-value pair for this new data value, frequency 1
return freqDict # return the dictionary created
# + deletable=true editable=true
myExampleDataList = [1,2,2,3,3,3] # a list with one 1, two 2's and three 3's
myExampleFreqDict = makeFreqDict(myExampleDataList) # the frequencies are returned as a dictionary
myExampleFreqDict
# + deletable=true editable=true
myExampleFreqDict.items() # this returns the (k,v) or key,value pairs in the dictionary as a list
# + deletable=true editable=true
myExampleFreqDict.keys() # this returns the keys
# + deletable=true editable=true
myExampleFreqDict.values() # this returns the values which happens to be same in our case!
# + deletable=true editable=true
myExampleFreqLongsDict = makeFreqDict(myExampleDataList,long(1)) # the long frequencies are returned as a dictionary
myExampleFreqLongsDict
# + deletable=true editable=true
myExampleFreqIntDict = makeFreqDict(myExampleDataList, int(1)) # the int frequencies are returned as a dictionary
myExampleFreqIntDict
# + deletable=true editable=true
myExampleFreqDict = makeFreqDict(myExampleDataList) # the frequencies are returned as a dictionary with default one
myExampleFreqDict
# + deletable=true editable=true
myExampleFreqIntDict == myExampleFreqDict # th last two freq dictionaries are the same!
# + [markdown] deletable=true editable=true
# Note that the function `makeFreqDict` also works on other sequence types like `numpy.arrays`, and `str` or sequence of bytes.
# + deletable=true editable=true
myExampleDataArray = np.array(myExampleDataList)
makeFreqDict(myExampleDataArray)
# + deletable=true editable=true
myExampleDataStr = '122333'
makeFreqDict(myExampleDataStr)
# + [markdown] deletable=true editable=true
# Let us use the `makeFreqDict` to find the frequencies of the balls that popped out first in the 1114 NZ Lotto draws.
# + deletable=true editable=true
ballOneFreqs = makeFreqDict(listBallOne) # call the function
print(ballOneFreqs) # disclose it
# + [markdown] deletable=true editable=true
# Thus, balls labelled by the number 1 come up 29 times in the first ball drawn (Ball One) out of 1,114 Lotto trials. Similarly, the number 2 comes up 28 times, ... 40 comes up 37 times. Of course, these numbers would be different if you have downloaded a more recent data file with additional trials!
#
# So what? Well, we'd hope that the lotto is fair, i.e. that the probability of each ball coming up with any of the available numbers is the same for each number: the probability that Ball One is a 1 is the same as the probability that it is 2, is the same as the probability that it is 3,..... , is the same as the probability that it is 40. If the lotto is fair, the number that comes up on each ball should be a discrete uniform random variable. More formally, we would expect it to be the de Moivre$(\frac{1}{40}, \frac{1}{40}, \ldots, \frac{1}{40})$ RV as lectured. Over the long term, we'd expect the number of times each number comes up on a given trial to be about the same as the number of times any other number comes up on that trial.
#
# We have data from 1987 to 2008, and a first step to assessing the fairness of the lotto (for Ball One, anyway) could be to just visualise the data. We can use the list of points we created above and the SageMath plotting function points to plot a simple graphic like this.
#
# Here we are plotting the frequency with which each number comes up against the numbers themselves, but it is a bit hard to see what number on the ball each red point relates to. To deal with this we add dotted lines going up from the number on the horizontal axis to the corresponding (number, frequency) tuple plotted as a red point.
#
# For these plots let us use SageMath Graphics primitives: `point`, `points` and `line`, and their `+` operations for superimpositions.
# + deletable=true editable=true
# ?point # Returns either a 2-dimensional or 3-dimensional point or "sum" of points as Graphics object
# + deletable=true editable=true
# ?line # Returns either a 2-dimensional or 3-dimensional line depending on value of points as Graphics object
# + deletable=true editable=true
print(ballOneFreqs.items())
# + deletable=true editable=true
# DATA we plot each item (k,f) in ballOneFreqs.items() as a red points at (k,f), k is ball number, f is frequency
lottoPlotCounts = point(ballOneFreqs.items(), rgbcolor="red")
# MODEL next loop through each key k and add a blue dotted line that goes from (k, 0) to (k, ballOneFreqs[k])
for k in ballOneFreqs.keys():
lottoPlotCounts += line([(k, 0),(k, ballOneFreqs[k])], rgbcolor="blue", linestyle=":")
show(lottoPlotCounts, figsize=[8,3])
# + [markdown] deletable=true editable=true
# ## Empirical mass function - just another statistic
#
# What about plotting the height of a point as the relative frequency rather than the frequency? The relative frequency for a number is the count (frequency) for that number divided by the sample size, i.e., the total number of trials. This can be formalized as the following process of adding normalized indicator functions of the data points $(X_1,X_2,\ldots,X_n)$: $$\frac{1}{n} \sum_{i=1}^n \mathbf{1}_{\{X_i\}}(x)$$
#
# In the next section on *List comprehensions and tuples* below we will show how the guts of the `makeEMF` function works. For now just evaluate the next cell and use the `makeEMF` so that you can just concentrate on the data for now. All `makeEMF` does is turn the frequencies in the above plot to relative frequences that sum to $1$ instead of the sample size $n=1114$.
#
# Now, we plot the points based on relative frequencies. What we have made is another statistic called the empirical mass function.
# + deletable=true editable=true
def makeEMF(myDataList):
'''Make an empirical mass function from a data list.
Param myDataList, list of data to make emf from.
Return list of tuples comprising (data value, relative frequency) ordered by data value.'''
freqs = makeFreqDict(myDataList) # make the frequency counts mapping
totalCounts = RR(sum(freqs.values())) # make mpfr_real to make relFreqs are fractions
relFreqs = [fr/totalCounts for fr in freqs.values()] # use a list comprehension
numRelFreqPairs = zip(freqs.keys(), relFreqs) # zip the keys and relative frequencies together
return numRelFreqPairs
# + deletable=true editable=true
numRelFreqPairs = makeEMF(listBallOne) # make a list of unique data values and their relative frequencies
lottoPlotEMF = point(numRelFreqPairs, rgbcolor = "purple")
for k in numRelFreqPairs: # for each tuple in the list
kkey, kheight = k # unpack tuple
lottoPlotEMF += line([(kkey, 0),(kkey, kheight)], rgbcolor="blue", linestyle=":")
show(lottoPlotEMF, figsize=[8,3])
# + [markdown] deletable=true editable=true
# Let us plot the probability mass function (PMF) of the hypothesized probability model for a fair NZ Lotto, i.e., the PMF of the RV $X$ ~ de Moivre$(1/40,1/40, ... , 1/40)$ and overlay it on the empirical mass function of the observed data. We will meet list comprehension properly in a future worksheet.
# + deletable=true editable=true
# list comprehension by the constant 1/40 for f(x)=1/40, x=1,2,...,40
ballOneEqualProbs = [1/40 for x in range(1,41,1)]
# + deletable=true editable=true
print(ballOneEqualProbs)
# + deletable=true editable=true
# make a list of (x,f(x)) tuples for the PDF using zip
numEqualProbsPairs = zip(ballOneFreqs.keys(), ballOneEqualProbs)
print(numEqualProbsPairs) # print the list of tuple
# + deletable=true editable=true
numRelFreqPairs = makeEMF(listBallOne) # make a list of unique data values and their relative frequencies
# make the EMF plot
lottoPlotEMF = point(numRelFreqPairs, rgbcolor = "purple")
for k in numRelFreqPairs: # for each tuple in the list
kkey, kheight = k # unpack tuple
lottoPlotEMF += line([(kkey, 0),(kkey, kheight)], rgbcolor="blue", linestyle=":")
ballOneEqualProbs = [1/40 for x in range(1,41,1)] # make sure we have the equal probabilities
numEqualProbsPairs = zip(ballOneFreqs.keys(), ballOneEqualProbs) # and the pairs
# make a plot of the equal probability pairs
equiProbabledeMoivre40PMF = point(numEqualProbsPairs, rgbcolor = "green")
for e in numEqualProbsPairs: # for each tuple in list
ekey, eheight = e # unpack tuple
equiProbabledeMoivre40PMF += line([(ekey, 0),(ekey, eheight)], rgbcolor="green", linestyle=":")
show(equiProbabledeMoivre40PMF + lottoPlotEMF, figsize=[8,3]) # plot the PMF and the PDF
# + [markdown] deletable=true editable=true
# ### Empirical Distribution Function
#
# Another extremely important statistics of the observed data is called the empirical distribution function (EDF). The EDF is the empirical or data-based distribution function (DF) just like the empirical mass function (EMF) is the empirical or data-based probability mass function (PMF). This can be formalized as the following process of adding indicator functions of the half-lines beginning at the data points
# $[X_1,+\infty),[X_2,+\infty),\ldots,[X_n,+\infty)$:
# $$\widehat{F}_n (x) = \frac{1}{n} \sum_{i=1}^n \mathbf{1}_{[X_i,+\infty)}(x) $$
#
# Next, let's make a function called `makeEDF` to return the empirical distribution function from a list of data points.
# + deletable=true editable=true
def makeEDF(myDataList):
'''Make an empirical distribution function from a data list.
Param myDataList, list of data to make emf from.
Return list of tuples comprising (data value, cumulative relative frequency) ordered by data value.
'''
freqs = makeFreqDict(myDataList) # make the frequency counts mapping
totalCounts = RR(sum(freqs.values())) # make mpfr_real to make relFreqs as fractions
relFreqs = [fr/totalCounts for fr in freqs.values()] # use a list comprehension
relFreqsArray = np.array(relFreqs)
cumFreqs = list(relFreqsArray.cumsum())
numCumFreqPairs = zip(freqs.keys(), cumFreqs) # zip the keys and culm relative frequencies together
return numCumFreqPairs
# + deletable=true editable=true
numCumFreqPairs = makeEDF(listBallOne)
lottoPlotEDF = points(numCumFreqPairs, rgbcolor = "black", faceted = true)
for k in range(len(numCumFreqPairs)):
x, kheight = numCumFreqPairs[k] # unpack tuple
previous_x = 0
previous_height = 0
if k > 0:
previous_x, previous_height = numCumFreqPairs[k-1] # unpack previous tuple
lottoPlotEDF += line([(previous_x, previous_height),(x, previous_height)], rgbcolor="grey")
lottoPlotEDF += points((x, previous_height),rgbcolor = "white", faceted = true)
lottoPlotEDF += line([(x, previous_height),(x, kheight)], rgbcolor="blue", linestyle=":")
show(lottoPlotEDF, figsize=(8,6))
# + [markdown] deletable=true editable=true
# ## Real-world Data Ingestion
#
# Let us download the New York Power Ball winning lottery numbers since 2010 from this US government site:
#
# - [https://catalog.data.gov/dataset/lottery-powerball-winning-numbers-beginning-2010](https://catalog.data.gov/dataset/lottery-powerball-winning-numbers-beginning-2010)
#
# using this url as a comma-separated-variable or csv file:
#
# - [https://data.ny.gov/api/views/d6yy-54nr/rows.csv?accessType=DOWNLOAD](https://data.ny.gov/api/views/d6yy-54nr/rows.csv?accessType=DOWNLOAD)
#
# #### This is a live fetch! So we have to figure it out on the fly as new records are added to the winning New York power ball data.
# + deletable=true editable=true
from urllib import *
NYpowerball = urlopen('https://data.ny.gov/api/views/d6yy-54nr/rows.csv?accessType=DOWNLOAD').read().decode('utf-8')
# + deletable=true editable=true
type(NYpowerball) # this is a unicode sequence
# + deletable=true editable=true
NYpowerballLines = NYpowerball.split('\n') # this splits the unicode by the end of line character '\n' in a list
# + deletable=true editable=true
type(NYpowerballLines) # we have a list of lines now
# + deletable=true editable=true
NYpowerballLines[0:10] # we can see the first 10 lines now and the first 0-index row is the header line
# + deletable=true editable=true
len(NYpowerballLines) # looks like there are 919 items (lines) in the list
# + deletable=true editable=true
NYpowerballLines[1] # this grabs the first row of data - the winning numbers from Feb 03 2010
# + deletable=true editable=true
NYpowerballLines[1].split(',') # this further split the row by comma character ',' into another list
# + deletable=true editable=true
NYpowerballLines[1].split(',')[1] # this grabs the second element of the list with index 1
# + deletable=true editable=true
NYpowerballLines[1].split(',')[1].split(' ') # this splits the 6 integers by white space ' '
# + deletable=true editable=true
NYpowerballLines[1].split(',')[1].split(' ')[0] # this grabs the first element of this list with 0-index
# + deletable=true editable=true
int(NYpowerballLines[1].split(',')[1].split(' ')[0]) # now we can turn this string into a Python int
# + [markdown] deletable=true editable=true
# That was not too hard. But what we really want is the first winning number across all the draws.
#
# We can simply apply the expression in the last cell and loop it through each item say `line` in `NYpowerballLines`:
#
# - `int(line[1].split(',')[1].split(' ')[0])`
#
# So let's do it.
# + deletable=true editable=true
firstWins = [] # initialize an empty list to append to it each winning first number
for line in NYpowerballLines:
win1 = int(line.split(',')[1].split(' ')[0])
firstWins.append(win1)
# + [markdown] deletable=true editable=true
# Opps, we got a `ValueError: invalid literal for int() with base 10: 'Winning'`.
# + deletable=true editable=true
NYpowerballLines[0] # remember our header line at index 0 has 'Winning' a string ad this can't be cast to int
# + [markdown] deletable=true editable=true
# We need to handle such error carefully, as data downloaded from public sources will generally come woth such caveats and more complicated issues in general.
#
# A proper way to handle such errors safely is called 'Exception Handling' using `try` and `catch`. Briefly from the [docs](https://docs.python.org/2/tutorial/errors.html#handling-exceptions):
#
# > The `try` statement works as follows.
# >
# > - First, the try clause (the statement(s) between the try and except keywords) is executed.
# > - If no exception occurs, the except clause is skipped and execution of the try statement is finished.
# > - If an exception occurs during execution of the try clause, the rest of the clause is skipped. Then if its type matches the exception named after the except keyword, the except clause is executed, and then execution continues after the try statement.
# > - If an exception occurs which does not match the exception named in the except clause, it is passed on to outer try statements; if no handler is found, it is an unhandled exception and execution stops with a message as shown above.
# + deletable=true editable=true
# let's be cautious and process only the first 10 lines
firstWins = [] # initialize an empty list to append to it each winning first number
for line in NYpowerballLines[0:10]:
try:
win1 = int(line.split(',')[1].split(' ')[0])
firstWins.append(win1)
except ValueError:
print "Err.. numbers only - input was: ", line
# + deletable=true editable=true
firstWins # looks like we gracefully converted the first winning numbers for lines 2-10 indexed 1-9
# + deletable=true editable=true
# now let's bravely process all the lines
firstWins = [] # initialize an empty list to append to it each winning first number
for line in NYpowerballLines:
try:
win1 = int(line.split(',')[1].split(' ')[0])
firstWins.append(win1)
except ValueError:
print "Err.. numbers only - line was: ", line
# + [markdown] deletable=true editable=true
# Now we have a new error `IndexError: list index out of range`. But let's see how many numbers have been appended in our list `firstWins`.
# + deletable=true editable=true
len(firstWins)
# + deletable=true editable=true
NYpowerballLines[918] # this grabs the second last row of data - the winning numbers from May 12 2018
# + deletable=true editable=true
len(NYpowerballLines) # but there is another line at index 918
# + deletable=true editable=true
NYpowerballLines[918] # contents of the last line looks empty! this is our problem
# + [markdown] deletable=true editable=true
# So, we need to handle this new exception.
#
# From the [docs](https://docs.python.org/2/tutorial/errors.html#handling-exceptions) we note that:
#
# > A `try` statement may have more than one `except` clause, to specify handlers for different exceptions. At most one handler will be executed....
#
# Let's have an `except` clause for `IndexError` next.
# + deletable=true editable=true
# now let's bravely process all the lines
firstWins = [] # initialize an empty list to append to it each winning first number
for line in NYpowerballLines:
try:
win1 = int(line.split(',')[1].split(' ')[0])
firstWins.append(win1)
except ValueError:
print "Err.. numbers only - line was: ", line
except IndexError:
print "list length had IndexError - line was: ", line
# + deletable=true editable=true
len(firstWins) # now we got an exception-handled data processing done!
# + deletable=true editable=true
print firstWins[0:10], "...", firstWins[906:916] # the first and last 10 balls in our list
# + [markdown] deletable=true editable=true
# Congratulations! You have taken your first step towards being a data scientist!
#
# Data scientists are expected to directly work with data and not merely load data from pre-made libraries.
#
# More crucially, *mathematical statistical data scientists* are expected to be comfortable with axiomatic probability and mathematical statistical theory (we will learn more in the sequel).
#
# Finally, *computational mathematical statistical data scientists*, students being trained here, can apply (and even develop as needed in cooperation with domain experts) the right model, methods and theory for a given data science problem and solve it using the most appropriate computational tools.
# + [markdown] deletable=true editable=true
# # Arrays
#
# ## YouTrys!
#
# We have already talked about lists in SAGE. Lists are great but are also general: lists are designed to be able to cope with any sort of data but that also means they don't have some of the specific functionality we might like to be able to analyse numerical data sets. Arrays provide a way of representing and manipulating numerical data known an a matrix in maths lingo. Matrices are particularly useful for data science.
#
# The lists we have been dealing with were one-dimensional. An array and the matrix it represents in a computer can be multidimensional. The most common forms of array that we will meet are one-dimensional and two-dimensional. You can imagine a two-dimensional array as a grid, which has rows and columns. Take for instance a 3-by-6 array (i.e., 3 rows and 6 columns).
#
# <table border="1"><colgroup> <col width="20" /> <col width="30" /> <col width="20" /> <col width="30" /> <col width="20" /> <col width="30" /> </colgroup>
# <tbody>
# <tr>
# <td>0</td>
# <td>1</td>
# <td>2</td>
# <td>3</td>
# <td>4</td>
# <td>5</td>
# </tr>
# <tr>
# <td>6</td>
# <td>7</td>
# <td>8</td>
# <td>9</td>
# <td>10</td>
# <td>11</td>
# </tr>
# <tr>
# <td>12</td>
# <td>13</td>
# <td>14</td>
# <td>15</td>
# <td>16</td>
# <td>17</td>
# </tr>
# </tbody>
# </table>
#
# The array type is not available in the basic SageMath package, so we import a library package called numpy which includes arrays.
# + deletable=true editable=true
import numpy as np
# this is just to print
print 'numpy is imported'
# + [markdown] deletable=true editable=true
# Because the arrays are part of numpy, not the basic SageMath package, we don't just say `array`, but instead qualify `array` with the name of the module it is in. So, we say `np.array` since we did `import numpy as np`.
#
# The cell below makes a two-dimensional 3-by-6 (3 rows by 6 columns) array by specifying the rows, and each element in each row, individually.
# + deletable=true editable=true
array1 = np.array([[0,1,2,3,4,5],[6,7,8,9,10,11],[12,13,14,15,16,17]]) # make an array the hard way
array1
# + [markdown] deletable=true editable=true
# We can use the `array`'s `shape` method to find out its **shape**. The shape method for a two-dimensional array returns an ordered pair in the form `(rows,columns)`. In this case it tells us that we have 3 rows and 6 columns.
# + deletable=true editable=true
array1.shape
# + [markdown] deletable=true editable=true
# Another way to make an `array` is to start with a one-dimensional array and `resize` it to give the `shape` we want, as we do in the two cells below.
#
# We start by making a one-dimensional array using the `range` function we have met earlier.
# + deletable=true editable=true
array2 = np.array([int(i) for i in range(18)]) # make an one dimensional array of Python int's
array2
# + [markdown] deletable=true editable=true
# The array's `resize` allows us to specify a new shape for the same array. We are going to make a two-dimensional array with 3 rows and 6 columns. Note that the shape you specify must be compatible with the number of elements in the array.
#
# In this case, `array2` has 18 elements and we specify a new shape of 3 rows and 6 columns, which still makes 18 = 3*6 elements.
#
# If the new shape you specify won't work with the array, you'll get an error message.
# + deletable=true editable=true
array2.resize(3,6) # use the array's resize method to change the shape of the array 18=3*6
array2
# + [markdown] deletable=true editable=true
# If you check the type for `array2`, you might be surprised to find that it is a `numpy.ndarray`. What happened to `numpy.array`? The array we are using is the NumPy N-dimensional array, or `numpy.ndarray`.
# + deletable=true editable=true
type(array2)
# + [markdown] deletable=true editable=true
# Tensors and general multidimensional arrays are possible via `resize` method.
# + deletable=true editable=true
array2.resize(3,2,3) # use the array's resize method to change the shape of the array 18=3*2*3
array2
# + [markdown] deletable=true editable=true
# The `range` function will only give us ranges of integers. The numpy module includes a more flexible function `arange` (that's `arange` with one `r` -- think of it as something like a-for-array-range -- not 'arrange' with two r's). The `arange` function takes parameters specifying the `start_argument`, `stop_argument` and `step_argument` values (similar to `range`), but is not restricted to integers and returns a one-dimensional array rather than a list.
#
# Here we use `arange` to make an array of numbers from `0.0` (`start_argument`) going up in steps of `0.1` (`step_argument`) to `0.18` - `0.1` and just as with range, the last number we get in the array is `stop_argument - step_argument` = `0.18` - `0.01` = `0.17`.
# + deletable=true editable=true
array3 = np.arange(0.0,0.18,0.01) # the pylab.arange(start, stop, step)
array3
# + deletable=true editable=true
type(array3) # type is actually a numpy.ndarray
# + deletable=true editable=true
array3.shape # the shape is 1 dimensional
# + [markdown] deletable=true editable=true
# ### Resize and Reshape
#
# The `resize` method resizes the array it is applied to.
#
# The `reshape` method will leave the original array unchanged but return a new array, based on the original one, of the required new shape.
# + deletable=true editable=true
array3.resize(9,2) # which we can resize into a 9 by 2 array
array3
# + deletable=true editable=true
array3.shape # try to see the shape of array3 now
# + deletable=true editable=true
array4 = array3.reshape(6,3) # reshape makes a new array of the specified size 6 by 3
array4
# + deletable=true editable=true
array4.shape # try to see the shape of array4 now
# + deletable=true editable=true
array3.shape # try to see the shape of array3 now, i.e. after it was reshaped into array4
# + [markdown] deletable=true editable=true
# So the reshape does leave the original array unchanged.
# + [markdown] deletable=true editable=true
# ## Arrays: indexing, slicing and copying
#
# Remember indexing into lists to find the elements at particular positions? We can do this with arrays, but we need to specify the index we want for each dimension of the array. For our two-dimensional arrays, we need to specify the row and column index, i.e. use a format like [`row_index`,`column_index`]. For exampe, [0,0] gives us the element in the first column of the first row (as with lists, array indices start from 0 for the first element).
# + deletable=true editable=true
array4 = np.arange(0.0, 0.18, 0.01) # make sure we have array4
array4.resize(6,3)
array4
# + [markdown] deletable=true editable=true
# `array4[0,0]` gives the element in the first row and first column while `array4[5,2]` gives us the element in the third column (index 2) of the sixth row (index 5).
# + deletable=true editable=true
array4[0,0]
# + deletable=true editable=true
array4[5,2]
# + [markdown] deletable=true editable=true
# We can use the colon to specify a range of columns or rows. For example, `[0:4,0]` gives us the elements in the first column (index 0) of rows with indices from 0 through 3. Note that the row index range `0:4` gives rows with indices starting from index 0 and ending at index `3=4-1`, i.e., indices `0` through `3`.
# + deletable=true editable=true
array4[0:4,0]
# + [markdown] deletable=true editable=true
# Similarly we could get all the elements in the second column (column index 1) of rows with indices 2 through 4.
# + deletable=true editable=true
array4[2:5,1]
# + [markdown] deletable=true editable=true
# The colon on its own gives everything, so a column index of `:` gives all the columns. Thus we can get, for example, all elements of a particular row -- in this case, the third row.
# + deletable=true editable=true
array4[2,:]
# + [markdown] deletable=true editable=true
# Or all the elements of a specified column, in this case the first column.
# + deletable=true editable=true
array4[:,0]
# + [markdown] deletable=true editable=true
# Or all the elements of a range of rows (think of this as like slicing the array horizontally to obtain row indices 1 through 4=5-1).
# + deletable=true editable=true
array4[1:5,:]
# + [markdown] deletable=true editable=true
# Or all the elements of a range of columns (think of this as slicing the array vertically to obtain column indices 1 through 2).
# + deletable=true editable=true
array4[:,1:3]
# + [markdown] deletable=true editable=true
# Naturally, we can slice both horizontally and vertically to obtain row indices 2 through 4 and column indices 0 through 1, as follows:
# + deletable=true editable=true
array4[2:5,0:2]
# + [markdown] deletable=true editable=true
# Finally, `[:]` gives a copy of the whole array. This copy of the original array can be assigned to a new name for furter manipulation without affecting the original array.
# + deletable=true editable=true
CopyOfArray4 = array4[:] # assign a copy of array4 to the new array named CopyOfArray4
CopyOfArray4 # disclose CopyOfArray4
# + deletable=true editable=true
CopyOfArray4.resize(9,2) # resize CopyOfArray4 from a 6-by-3 array to a 9-by-2 array
CopyOfArray4 # disclose CopyOfArray4 as it is now
# + deletable=true editable=true
array4 # note that our original array4 has remained unchanged with size 6-by-3
# + [markdown] deletable=true editable=true
# ## Useful arrays
#
# NumPy provides quick ways of making some useful kinds of arrays. Some of these are shown in the cells below.
#
# An array of zeros or ones of a particular shape. Here we ask for shape (2,3), i.e., 2 rows and 3 columns.
# + deletable=true editable=true
arrayOfZeros = np.zeros((2,3)) # get a 2-by-3 array of zeros
arrayOfZeros
# + deletable=true editable=true
arrayOfOnes = np.ones((2,3)) # get a 2-by-3 array of ones
arrayOfOnes
# + [markdown] deletable=true editable=true
# An array for the identity matrix, i.e., square (same number of elements on each dimension) matrix with 1's along the diagonal and 0's everywhere else.
# + deletable=true editable=true
iden = np.identity(3) # get a 3-by-3 identity matrix with 1's along the diagonal and 0's elsewhere
iden
# + [markdown] deletable=true editable=true
# ## Useful functions for sequences
#
# Now we are going to demonstrate some useful methods of sequences. A `list` is a sequence, and so is a `tuple`.
#
# There are differences between them (recall that tuples are immutable) but in many cases we can use the same methods on them because they are both sequences.
#
# We will demonstrate with a list and a tuple containing the same data values. The data could be the results of three IID Bernoulli trials..
# + deletable=true editable=true
obsDataList = [0, 1, 1]
obsDataList
# + deletable=true editable=true
obsDataTuple = tuple(obsDataList)
obsDataTuple
# + [markdown] deletable=true editable=true
# A useful operation we can perform on a tuple is to count the number of times a particular element, say 0 or 1 in our ObsDataTuple, occurs. This is a statistic of our data called the sample frequency of the element of interest.
# + deletable=true editable=true
obsDataTuple.count(0) # frequency of 0 using the tuple
# + deletable=true editable=true
obsDataList.count(1) # frequency of 1 using the list
# + [markdown] deletable=true editable=true
# Another useful operation we can perform on a tuple is to sum all elements in our `obsDataTuple` or further scale the sum by the sample size. These are also statistics of our data called the sample sum and the sample mean, respectively. Try the same things with `obsDataList`.
# + deletable=true editable=true
sum(obsDataTuple) # sample sum
# + deletable=true editable=true
sum(obsDataTuple)/3 # sample mean using sage.rings.Rational
# + deletable=true editable=true
obsDataTuple.count(1) / 3 # alternative expression for sample mean in IID Bernoulli trials
# + [markdown] deletable=true editable=true
# We used a lot of the techniques we have seen so far when we wanted to get the relative frequency associated with each ball in the lotto data.
#
# Let's revist the code that made relative frequencies and fully understand each part of it now.
# + deletable=true editable=true
ballOneFreqs = makeFreqDict(listBallOne) # call the function to make the dictionary
totalCounts = sum(ballOneFreqs.values())
relFreqs = []
for k in ballOneFreqs.keys():
relFreqs.append(k/totalCounts)
numRelFreqPairs = zip(ballOneFreqs.keys(), relFreqs) # zip the keys and relative frequencies together
print(numRelFreqPairs)
# + [markdown] deletable=true editable=true
# We could also do it with a list comprehension as follows:
# + deletable=true editable=true
ballOneFreqs = makeFreqDict(listBallOne) # call the function to make the dictionary
totalCounts = sum(ballOneFreqs.values())
relFreqs = [k/totalCounts for k in ballOneFreqs.keys()]
numRelFreqPairs = zip(ballOneFreqs.keys(), relFreqs) # zip the keys and relative frequencies together
print(numRelFreqPairs)
# + [markdown] deletable=true editable=true
# ## More on tuples and sequences in general
#
# First, we can make an empty tuple like this:
# + deletable=true editable=true
empty = ()
len(empty)
# + [markdown] deletable=true editable=true
# Secondly, if we want a tuple with only one element we have to use a trailing comma. If you think about it from the computer's point of view, if it can't see the trailing comma, how is it to tell that you want a tuple not just a single number?
# + deletable=true editable=true
aFineSingletonTuple = (1, )
type(aFineSingletonTuple)
# + deletable=true editable=true
unFineSingletonTuple = (1)
type(unFineSingletonTuple)
# + deletable=true editable=true
unFineSingletonTuple = ((1))
type(unFineSingletonTuple)
# + deletable=true editable=true
aFineSingletonTuple = ('lonelyString', )
type(aFineSingletonTuple)
# + deletable=true editable=true
unFineSingletonTuple = ('lonelyString')
type(unFineSingletonTuple)
# + [markdown] deletable=true editable=true
# You can make tuples out almost any object. It's as easy as putting the things you want in ( ) parentheses and separating them by commas.
# + deletable=true editable=true
myTupleOfStuff = (1, ZZ(1), QQ(1), RR(1), int(1), long(1), float(1))
myTupleOfStuff
# + deletable=true editable=true
1==ZZ(1) # remember? will this be True/False?
# + [markdown] deletable=true editable=true
# Actually, you don't even need the ( ). If you present SageMath (Python) with a sequence of object separated by commas, the default result is a tuple.
# + deletable=true editable=true
myTuple2 = 60, 400 # assign a tuple to the variable named myTuple2
type(myTuple2)
# + [markdown] deletable=true editable=true
# A statement like the one in the cell above is known as tuple packing (more generally, sequence packing).
#
# When we work with tuples we also often use the opposite of packing - Python's very useful sequence unpacking capability. This is, literally, taking a sequence and unpacking or extracting its elements.
#
#
# + deletable=true editable=true
x, y = myTuple2 # tuple unpacking
print x
print y
x * y
# + [markdown] deletable=true editable=true
# The statement below is an example of *multiple assignment*, which you can see now is really just a combination of tuple packing followed by unpacking.
# + deletable=true editable=true
x, y = 600.0, 4000.0
print x
print y
x/y
# + [markdown] deletable=true editable=true
# Let's try to implement a for loop with a tuple -- it will work just like the for loop with a list.
# + deletable=true editable=true
for x in (1,2,3):
print x^2
# + [markdown] deletable=true editable=true
# Next let's try a list comprehension on the tuple. This creates a list as expected.
# + deletable=true editable=true
[x^2 for x in (1,2,3)]
# + [markdown] deletable=true editable=true
# But if we try the same comprehension with tuples, i.e., with `()` insteqad of `[]` we get a `generator object`. This will be covered next by first easing into functional programming in Python.
# + deletable=true editable=true
(x^2 for x in (1,2,3))
# + [markdown] deletable=true editable=true
# # Functional Programming in Python
#
# Let us take at the basics of functional programming and what Python has to offer on this front at:
#
# - [https://docs.python.org/2/howto/functional.html#functional-programming-howto](https://docs.python.org/2/howto/functional.html#functional-programming-howto)
# + deletable=true editable=true
showURL("https://docs.python.org/2/howto/functional.html#functional-programming-howto",600)
# + [markdown] deletable=true editable=true
# ## Iterators
#
# From [https://docs.python.org/2/howto/functional.html#iterators](https://docs.python.org/2/howto/functional.html#iterators):
#
# > An iterator is an object representing a stream of data; this object returns the data one element at a time. A Python iterator must support a method called next() that takes no arguments and always returns the next element of the stream. If there are no more elements in the stream, next() must raise the StopIteration exception. Iterators don’t have to be finite, though; it’s perfectly reasonable to write an iterator that produces an infinite stream of data.
#
# > The built-in [iter()](https://docs.python.org/2/library/functions.html#iter) function takes an arbitrary object and tries to return an iterator that will return the object’s contents or elements, raising [TypeError](https://docs.python.org/2/library/exceptions.html#exceptions.TypeError) if the object doesn’t support iteration. Several of Python’s built-in data types support iteration, the most common being lists and dictionaries. An object is called an iterable object if you can get an iterator for it.
#
# > You can experiment with the iteration interface manually:
# + deletable=true editable=true
L = [1,2,3]
it = iter(L)
it
# + deletable=true editable=true
type(it)
# + deletable=true editable=true
it.next()
# + deletable=true editable=true
it.next()
# + deletable=true editable=true
it.next()
# + deletable=true editable=true
it.next()
# + [markdown] deletable=true editable=true
# > Iterators can be materialized as lists or tuples by using the list() or tuple() constructor functions:
# + deletable=true editable=true
L = [1,2,3]
it = iter(L)
t = tuple(it)
t
# + deletable=true editable=true
L = [1,2,3]
it = iter(L)
l = list(it)
l
# + deletable=true editable=true
it = iter(L)
max(it) # you can call functions on iterators
# + [markdown] deletable=true editable=true
# ## Generators
#
# "Naive Tuple Comprehension" creates a `generator` in SageMath/Python. From [https://docs.python.org/2/glossary.html#term-generator](https://docs.python.org/2/glossary.html#term-generator):
#
# > **generator**
# > A function which returns an iterator. It looks like a normal function except that it contains yield statements for producing a series of values usable in a for-loop or that can be retrieved one at a time with the next() function. Each yield temporarily suspends processing, remembering the location execution state (including local variables and pending try-statements). When the generator resumes, it picks-up where it left-off (in contrast to functions which start fresh on every invocation).
#
# > **generator expression**
# > An expression that returns an iterator. It looks like a normal expression followed by a for expression defining a loop variable, range, and an optional if expression. The combined expression generates values for an enclosing function:
#
# **List comprehensions (listcomps) and Generator Expressions (genexps)** from
# - [https://docs.python.org/2/howto/functional.html#generator-expressions-and-list-comprehensions](https://docs.python.org/2/howto/functional.html#generator-expressions-and-list-comprehensions)
#
# >With a list comprehension, you get back a Python list; stripped_list is a list containing the resulting lines, not an iterator. Generator expressions return an iterator that computes the values as necessary, not needing to materialize all the values at once. This means that list comprehensions aren’t useful if you’re working with iterators that return an infinite stream or a very large amount of data. Generator expressions are preferable in these situations.
# > Generator expressions are surrounded by parentheses (“()”) and list comprehensions are surrounded by square brackets (“[]”). Generator expressions have the form:
#
# ```
# ( expression for expr in sequence1
# if condition1
# for expr2 in sequence2
# if condition2
# for expr3 in sequence3 ...
# if condition3
# for exprN in sequenceN
# if conditionN )
# ```
# + [markdown] deletable=true editable=true
# > Two common operations on an iterator’s output are 1) performing some operation for every element, 2) selecting a subset of elements that meet some condition. For example, given a list of strings, you might want to strip off trailing whitespace from each line or extract all the strings containing a given substring.
#
# > List comprehensions and generator expressions (short form: “listcomps” and “genexps”) are a concise notation for such operations, borrowed from the functional programming language Haskell (https://www.haskell.org/). You can strip all the whitespace from a stream of strings with the following code:
# + deletable=true editable=true
line_list = [' line 1\n', 'line 2 \n', 'line 3 \n']
# List comprehension -- returns list
stripped_list = [line.strip() for line in line_list]
# Generator expression -- returns a generator
stripped_iter = (line.strip() for line in line_list)
# + deletable=true editable=true
stripped_list # returns a list
# + deletable=true editable=true
type(stripped_iter) # returns a generator
# + deletable=true editable=true
stripped_iter # returns only an genexpr generator object
# + deletable=true editable=true
list(stripped_iter) # the generator can be materialized into a list
# + deletable=true editable=true
tuple(stripped_iter) # the generator can be materialized into a tuple - it's already been emptied!
# + deletable=true editable=true
# Generator expression -- returns a generator
stripped_iter = (line.strip() for line in line_list) # let's create a new generator
tuple(stripped_iter) # the generator can be materialized into a tuple now
# + [markdown] deletable=true editable=true
# Once again we have emptied the `stripped_iter` so we only get an empty list when we materilize it into a list.
# + deletable=true editable=true
list(stripped_iter)
# + deletable=true editable=true
showURL("https://docs.python.org/2/howto/functional.html#generator-expressions-and-list-comprehensions",500)
# + deletable=true editable=true
myGenerator = (x^2 for x in (1,2,3)) # this is actually a generator
# + deletable=true editable=true
type(myGenerator)
# + deletable=true editable=true
myGenerator.next()
# + deletable=true editable=true
myGenerator.next()
# + deletable=true editable=true
myGenerator.next()
# + deletable=true editable=true
myGenerator.next()
# + [markdown] deletable=true editable=true
# We can pass a generator to a function like `sum` to evaluate it:
# + deletable=true editable=true
myGenerator = (i*i for i in range(10))
# + deletable=true editable=true
sum(myGenerator)
# + [markdown] deletable=true editable=true
# See [https://docs.python.org/2/howto/functional.html#generators](https://docs.python.org/2/howto/functional.html#generators) for more details:
#
# > Generators are a special class of functions that simplify the task of writing iterators. Regular functions compute a value and return it, but generators return an iterator that returns a stream of values.
#
# > You’re doubtless familiar with how regular function calls work in Python or C. When you call a function, it gets a private namespace where its local variables are created. When the function reaches a return statement, the local variables are destroyed and the value is returned to the caller. A later call to the same function creates a new private namespace and a fresh set of local variables. But, what if the local variables weren’t thrown away on exiting a function? What if you could later resume the function where it left off? This is what generators provide; they can be thought of as resumable functions.
#
# > Here’s the simplest example of a generator function:
# + deletable=true editable=true
def generate_ints(N):
for i in range(N):
yield i
# + [markdown] deletable=true editable=true
# > Any function containing a yield keyword is a generator function; this is detected by Python’s [bytecode](https://docs.python.org/2/glossary.html#term-bytecode) compiler which compiles the function specially as a result.
#
# > When you call a generator function, it doesn’t return a single value; instead it returns a generator object that supports the iterator protocol. On executing the yield expression, the generator outputs the value of i, similar to a return statement. The big difference between yield and a return statement is that on reaching a yield the generator’s state of execution is suspended and local variables are preserved. On the next call to the generator’s .next() method, the function will resume executing.
#
# > Here’s a sample usage of the generate_ints() generator:
# + deletable=true editable=true
gen = generate_ints(3)
gen
# + deletable=true editable=true
gen.next()
# + deletable=true editable=true
gen.next()
# + deletable=true editable=true
gen.next()
# + deletable=true editable=true
gen.next()
# + deletable=true editable=true
gen. # see the methods available on gen by hitting TAB after the 'gen.' try to `close()` the generator
# + [markdown] deletable=true editable=true
# Using list comprehensions and generator expressions one can do functional programming in Python. But this is not the same as doing pure functional programming in a language such as haskell (see [https://www.haskell.org/](https://www.haskell.org/)) that is designed specifically for pure funcitonal programming.
# + [markdown] deletable=true editable=true
# ## Keep learning
#
# We have taken several exercises above from http://docs.python.org/tutorial/datastructures.html.
#
# This is a very useful site to look for user-friendly information about Python in general (it can't help you with anything that Sage overlays on top of Python though, or with many of the specialised modules that you may want to import into your Sage notebook).
# + [markdown] deletable=true editable=true
# # A Chalenging YouTry!
#
# Recall how we downloaded *Pride and Prejudice* and processed it as a String and break it by Chapters. This is at our disposal - all we need to do is copy-paste the right set of cells from earlier here to have the string from that Book by fetching live from the Guthenberg Project again.
#
# Think about what algorithmic constructs and methods one will need to `split` each sentence by `words` it contains and count the number of each distinct word.
#
# Now that you have understood `for` loops, `list` comprehensions and anonymous `function`s, and can learn about the needed methods on strings for splitting (which you can search by adding a `.` after a `srt` and hitting the `Tab` button to look through exixting methods), the `dictionary` data structure, and already seen how to count the number of ball labels, you are ready for the challenge.
#
# Process the English words in the book *Pride and Prejudice* by <NAME> and obtain the empirical mass function of the 50 most frequent words. You may report it as a dictionary. If you have more time them try to plot the empirical mass function (at least for the top 20 words) as a picture/image as we did above.
# + deletable=true editable=true
# + deletable=true editable=true
# + [markdown] deletable=true editable=true
# # A YouTryIfYouWantTo on a Really Interesting Dataset
#
# ## Swedish Election Outcomes 2018
#
# See: [http://www.lamastex.org/datasets/public/elections/2018/sv/README](http://www.lamastex.org/datasets/public/elections/2018/sv/README)!
#
# It should say something like the following:
#
#
# This was obtained by processing using the scripts at:
#
# - https://gitlab.com/tilo.wiklund/swedis-election-data-scraping
#
# Download and extract using `tar`:
# ```
# $ wget http://www.lamastex.org/datasets/public/elections/2018/sv/final.tgz
# $ tar zxvf final.tgz
# ```
#
# Now you have all of the data in the csv file:
#
# ```
# $ head final.csv
# region,municipality,district,party,votes
# Blekinge län,Karlshamn,0 - Centrala Asarum,S,519
# Blekinge län,Karlshamn,0 - Centrala Asarum,SD,311
# Blekinge län,Karlshamn,0 - Centrala Asarum,M,162
# Blekinge län,Karlshamn,0 - Centrala Asarum,V,82
# Blekinge län,Karlshamn,0 - Centrala Asarum,KD,53
# Blekinge län,Karlshamn,0 - Centrala Asarum,C,37
# Blekinge län,Karlshamn,0 - Centrala Asarum,L,37
# Blekinge län,Karlshamn,0 - Centrala Asarum,MP,32
# Blekinge län,Karlshamn,0 - Centrala Asarum,BLANK,13
# ```
#
# ### Flex your Pythonic Muscles and create any interesting statistics of the dataset!
#
# This is just for fun and is a **YouTryIfYouWantTo**.
# + deletable=true editable=true
| _infty/2018/01/jp/05.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
import numpy as np
import time
import matplotlib.pyplot as plt
from scipy import interpolate
sys.path.append(r'C:\Chuji\Code_and_Data\MyCode')
import Circuit_Simulator
import PulseGenerator as PG
from toolfunc import *
from toolfunc.adia_analysis import *
from toolfunc.pulse_filter import *
import scipy.optimize as sci_op
from toolfunc import DE
# +
raw_config = Circuit_Simulator.RawConfig(qubit_num=3,dimension=3,circuit_type=1,initial_state='ground',sampling_rate=1e9)
raw_config.load_default_value(modulation=True,decoherence=False,use_capacitance=False)
flux_pulse = np.linspace(0/7.5,4/7.5,400)
freq_array = 8.5e9-flux_pulse * (8.5e9 - 1e9)
raw_config.setValue('Q1 f01_max',6.0e9)
raw_config.setValue('Q2 f01_max',8.5e9)
raw_config.setValue('Q3 f01_max',5.4e9)
raw_config.setValue('Q1 f01_min',1e9)
raw_config.setValue('Q2 f01_min',1e9)
raw_config.setValue('Q3 f01_min',1e9)
raw_config.setValue('Q1 Ec',0.25e9)
raw_config.setValue('Q2 Ec',0.30e9)
raw_config.setValue('Q3 Ec',0.25e9)
raw_config.setValue('r12',0.018)
raw_config.setValue('r23',0.018)
raw_config.setValue('r13',0.0015)
raw_config.setValue('Q2 Voltage period',-1)
raw_config.setValue('Q2 Voltage operating point',0.00)
raw_config.setValue('Q2 Flux',flux_pulse)
simu_config = Circuit_Simulator.read_config(raw_config.get_dict())
Simulator = Circuit_Simulator.Simulator(simu_config)
Simulator.show_pulse()
# +
Simulator.performsimulation(solver_type=2,resample_factor=1,eigen_cloest_to_bare=False ,sort_by_maximum_overlap=True,gap=12e6)
fig = plt.figure(figsize=[6.4,6])
ax = fig.add_subplot(111)
eigen_trace = Simulator.EigenResult.get_Ener_gap_trace('101-100-001+000')
ax.plot(freq_array[0:400],-eigen_trace[0:400]/1e6)
ax.set_yscale('log')
# -
# %matplotlib inline
def cost_func_distor(pulse_params,*args):
gate_time,SRATE,f_term,factor_r,T_reflec=args
# str_idx = bin(int(factor_idx))[2:6].zfill(4)
factor1=1.0
factorc=1.0
factor2=1.0
lamb1 = pulse_params
total_len = gate_time + 12e-9+4*T_reflec
Seq=PG.Sequence(total_len=total_len,sample_rate=SRATE,complex_trace=False)
Seq.clear_pulse(tips_on=False)
Seq.add_pulse('Adiabatic',t0=gate_time/2+12e-9/2,width=gate_time,plateau=0e-9,frequency=0,F_Terms=f_term,Lcoeff=np.array(lamb1),Q1_freq=6.0e9,
CPLR_idle_freq=(7.87e9-6e9)*factorc+6e9,Q2_freq=6e9+(5.4e9-6e9)*factor2,constant_coupling=False,r1c=0.018*factor1,r2c=0.018*factor2,r12=0.0015*factorc,anhar_CPLR=-300e6*factorc,
anhar_Q1=-250e6*factor1,anhar_Q2=-250e6*factor2,negative_amplitude=False,dfdV=(7.87e9-6e9)*factorc+6e9-1e9,gap_threshold=8e6,freqpoints=301,pulsepoints=601)
Seq.add_filter('Gauss Low Pass',300e6)
Seq.add_filter('Reflection',*(factor_r,T_reflec))
flux_pulse=Seq.get_sequence()
raw_config = Circuit_Simulator.RawConfig(qubit_num=3,dimension=3,circuit_type=1,initial_state='-Z+Z+Z',sampling_rate=SRATE)
raw_config.load_default_value(modulation=True,decoherence=False,use_capacitance=False)
raw_config.setValue('Q1 f01_max',6.0e9)
raw_config.setValue('Q2 f01_max',7.87e9)
raw_config.setValue('Q3 f01_max',5.4e9)
raw_config.setValue('Q1 f01_min',1e9)
raw_config.setValue('Q2 f01_min',1e9)
raw_config.setValue('Q3 f01_min',1e9)
raw_config.setValue('Q1 Ec',0.25e9)
raw_config.setValue('Q2 Ec',0.3e9)
raw_config.setValue('Q3 Ec',0.25e9)
raw_config.setValue('r12',0.018)
raw_config.setValue('r23',0.018)
raw_config.setValue('r13',0.0015)
raw_config.setValue('Q2 Voltage period',-1)
raw_config.setValue('Q2 Voltage operating point',0)
raw_config.setValue('Q2 Flux',flux_pulse)
simu_config = Circuit_Simulator.read_config(raw_config.get_dict())
Simulator = Circuit_Simulator.Simulator(simu_config)
Simulator.performsimulation(solver_type=1)
Simulator.UnitaryResult.get_U(-1)
Simulator.UnitaryResult.get_subspace_operator(['000','001','100','101'])
Simulator.UnitaryResult.remove_single_qubit_gate()
Simulator.UnitaryResult.set_Target_gate('CZ')
Simulator.UnitaryResult.get_Gate_Fidelity()
fidelity = Simulator.UnitaryResult.Gate_Fidelity
return 1 - fidelity
np.linspace(-0.1,0.1,21)
len(np.arange(0.5e-9,10.1e-9,0.5e-9))
# +
SRATE=6e9
gate_time=30e-9
f_terms=1
Tr_arr = np.arange(0.5e-9,30.1e-9,1e-9)
factor_r_arr = np.linspace(-0.1,0.1,21)
gate_fidelity_one = np.zeros([len(Tr_arr),len(factor_r_arr)])
gate_params_one = np.zeros([len(Tr_arr),len(factor_r_arr)])
raw_initial_seeds=np.array([0.8])
ii = 0
for T_r in Tr_arr:
jj = 0
for factor_r in factor_r_arr:
time_start = time.time()
DATA = sci_op.minimize(cost_func_distor,raw_initial_seeds,args=(gate_time,SRATE,f_terms,factor_r,T_r), method='Nelder-Mead', options={'disp': True,'ftol':5e-5,'xtol':5e-5,'maxiter':30})
gate_fidelity_one[ii,jj] = DATA.fun
gate_params_one[ii,jj] = DATA.x
print('fidelity',DATA.fun)
print(time.time()-time_start)
np.savetxt(r'C:\Chuji\Latex_Papers\Mypapers\ZZ_coupling_20210205\fig_zz\Robustness\params_one_distor_0ns_30ns.txt',gate_params_one )
np.savetxt(r'C:\Chuji\Latex_Papers\Mypapers\ZZ_coupling_20210205\fig_zz\Robustness\error_one_distor_0ns_30ns.txt',gate_fidelity_one )
jj+=1
ii+=1
# -
# +
gate_time=60e-9
SRATE=10e9
f_term=2
factor_r=-0.06
T_reflex=4e-9
factor1=1.0
factorc=1.0
factor2=1.0
lamb1 = [0.8,-0.1]
total_len = gate_time + 8e-9+4*T_reflex
Seq=PG.Sequence(total_len=total_len,sample_rate=SRATE,complex_trace=False)
Seq.clear_pulse(tips_on=False)
Seq.add_pulse('Adiabatic',t0=gate_time/2+12e-9/2,width=gate_time,plateau=0e-9,frequency=0,F_Terms=f_term,Lcoeff=np.array(lamb1),Q1_freq=6.0e9,
CPLR_idle_freq=(7.87e9-6e9)*factorc+6e9,Q2_freq=6e9+(5.4e9-6e9)*factor2,constant_coupling=False,r1c=0.018*factor1,r2c=0.018*factor2,r12=0.0015*factorc,anhar_CPLR=-300e6*factorc,
anhar_Q1=-250e6*factor1,anhar_Q2=-250e6*factor2,negative_amplitude=False,dfdV=(7.87e9-6e9)*factorc+6e9-1e9,gap_threshold=8e6,freqpoints=301,pulsepoints=601)
Seq.add_filter('Gauss Low Pass',300e6)
Seq.add_filter('Reflection',*(factor_r,T_reflex))
flux_pulse=Seq.get_sequence()
plt.plot(flux_pulse)
# -
# %matplotlib inline
# +
SRATE=10e9
f_terms=1
gate_time_arr = np.arange(12e-9,60.1e-9,1e-9)
for fac_idx in np.arange(0,7.1,1):
gate_fidelity_one = []
gate_params_one = []
raw_initial_seeds=np.array([3])
for gate_time in gate_time_arr:
time_start = time.time()
DATA = sci_op.minimize(cost_func_inhomogeneity,raw_initial_seeds,args=(gate_time,SRATE,f_terms,fac_idx,0.05), method='Nelder-Mead', options={'disp': True,'ftol':1e-5,'xtol':1e-5,'maxiter':30})
# DATA = sci_op.minimize(cost_func,raw_initial_seeds,args=(gate_time,SRATE,f_terms), method='Nelder-Mead', options={'disp': True,'ftol':1e-5,'xtol':1e-5,'maxiter':30})
gate_fidelity_one.append(DATA.fun)
gate_params_one.append(DATA.x)
raw_initial_seeds =DATA.x*0.92
print('gate time',gate_time)
print('fidelity',DATA.fun)
np.savetxt(r'C:\Chuji\Latex_Papers\Mypapers\ZZ_coupling_20210205\fig_zz\fig3_data\params_one_5_inhomo'+str(fac_idx)+'.txt',gate_params_one )
np.savetxt(r'C:\Chuji\Latex_Papers\Mypapers\ZZ_coupling_20210205\fig_zz\fig3_data\error_one_5_inhomo'+str(fac_idx)+'.txt',gate_fidelity_one )
np.savetxt(r'C:\Chuji\Latex_Papers\Mypapers\ZZ_coupling_20210205\fig_zz\fig3_data\gate_time_one_5_inhomo'+str(fac_idx)+'.txt',gate_time_arr )
# -
| examples/code for 'Coupler-assisted C-phase gate'/Robustness/Robustness to pulse distortion Tg=30ns.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/llpk79/DS-Unit-2-Regression-1/blob/master/Paul_K_doing_even_more_linear_regression.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="uD-iMXCvNabf" colab_type="code" outputId="313eaa14-2e75-4124-acc6-8ce464d90683" colab={"base_uri": "https://localhost:8080/", "height": 197}
import pandas as pd
# %matplotlib inline
from mpl_toolkits.mplot3d import Axes3D
from sklearn.metrics import mean_squared_error as mse, r2_score as r2, mean_absolute_error as mae
import numpy as np
from sklearn.linear_model import LinearRegression as LR
import matplotlib.pyplot as plt
columns = ['Year','Incumbent','Other','Incumbent Votes']
data = [[1952,"Stevenson","Eisenhower",44.6],
[1956,"Eisenhower","Stevenson",57.76],
[1960,"Nixon","Kennedy",49.91],
[1964,"Johnson","Goldwater",61.34],
[1968,"Humphrey","Nixon",49.60],
[1972,"Nixon","McGovern",61.79],
[1976,"Ford","Carter",48.95],
[1980,"Carter","Reagan",44.70],
[1984,"Reagan","Mondale",59.17],
[1988,"Bush, Sr.","Dukakis",53.94],
[1992,"Bush, Sr.","Clinton",46.55],
[1996,"Clinton","Dole",54.74],
[2000,"Gore","Bush, Jr.",50.27],
[2004,"Bush, Jr.","Kerry",51.24],
[2008,"McCain","Obama",46.32],
[2012,"Obama","Romney",52.00],
[2016,"Clinton","Trump",48.2]]
votes = pd.DataFrame(data=data, columns=columns)
columns = ['Year','Income growth']
data = [[1952,2.40],
[1956,2.89],
[1960, .85],
[1964,4.21],
[1968,3.02],
[1972,3.62],
[1976,1.08],
[1980,-.39],
[1984,3.86],
[1988,2.27],
[1992, .38],
[1996,1.04],
[2000,2.36],
[2004,1.72],
[2008, .10],
[2012, .95],
[2016, .10]]
growth = pd.DataFrame(data=data, columns=columns)
columns = ['Year','Fatalities']
data = [[1952,190],
[1956, 0],
[1960, 0],
[1964, 1],
[1968,146],
[1972, 0],
[1976, 2],
[1980, 0],
[1984, 0],
[1988, 0],
[1992, 0],
[1996, 0],
[2000, 0],
[2004, 4],
[2008, 14],
[2012, 5],
[2016, 5]]
deaths = pd.DataFrame(data=data, columns=columns)
df = votes.merge(growth).merge(deaths)
df.head()
# + id="n5vvvN55ONGE" colab_type="code" outputId="e858d19c-32d2-4ec5-99b4-cf9bf51d5e90" colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY> "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 112}
from google.colab import files
files.upload()
# + id="og2iuWn1R0mo" colab_type="code" outputId="b0c34449-0ac9-4258-dfb5-148aeb83ad5e" colab={"base_uri": "https://localhost:8080/", "height": 197}
price_index = pd.read_csv('CPIAUCSL.csv')
price_index.head()
# + id="-FCAj1kkSOON" colab_type="code" outputId="11254772-fa67-4dfd-8fd1-d2363c508067" colab={"base_uri": "https://localhost:8080/", "height": 318}
prices = []
for year in df['Year']:
time = str(year) + '-01-01'
prices.append(price_index[price_index['DATE'] == time]['CPIAUCSL'].values[0])
prices
# + id="KE7CYRhZTnBX" colab_type="code" colab={}
df['price index'] = prices
# + id="VvPWwJaMTrV7" colab_type="code" outputId="ce6446f8-41ff-4cce-aca0-d1b35cad2415" colab={"base_uri": "https://localhost:8080/", "height": 214}
df.head()
# + id="IKq52lewYZPb" colab_type="code" colab={}
features = ['Income growth', 'price index']
label = 'Incumbent Votes'
train = df.query('Year < 2000')
test = df.query('Year >= 2000')
X_train_0 = train[features]
y_train = train[label]
X_test_0 = test[features]
y_test = test[label]
# + id="fng8EVxTavG3" colab_type="code" outputId="9c5fbb43-5cd2-49f2-9021-4b981667beee" colab={"base_uri": "https://localhost:8080/", "height": 87}
model_0 = LR()
model_0.fit(X_train_0, y_train)
y_pred_0 = model_0.predict(X_test_0)
error_0 = mae(y_test, y_pred_0)
print(f'MAE for model_0 is {error_0}')
print(f'MSE for model_0 is {mse(y_test, y_pred_0)}')
print(f'RMSE for model_0 is {(mse(y_test, y_pred_0))**0.5}')
print(f'R2 for model_0 is {r2(y_test, y_pred_0)}')
# + id="qYJhmQI-a2D8" colab_type="code" colab={}
# features = ['Fatalities', 'price index']
# X_train_1 = train[features]
# X_test_1 = test[features]
# model_1 = LR()
# model_1.fit(X_train_1, y_train)
# y_pred_1 = model_1.predict(X_test_1)
# error_1 = mae(y_test, y_pred_1)
# print(f'MAE for model_1 is {error_1}')
# print(f'MSE for model_0 is {mse(y_test, y_pred_1)}')
# print(f'RMSE for model_0 is {(mse(y_test, y_pred_1)**0.5)}')
# print(f'R2 for model_0 is {r2(y_test, y_pred_1)}')
# + id="iPiFpnUdhU7m" colab_type="code" outputId="d75e80f8-96ac-4832-a160-62dfa222882e" colab={"base_uri": "https://localhost:8080/", "height": 34}
y_pred_0
# + id="AKVeDFOPdZK_" colab_type="code" outputId="4b66a6b7-2f40-407a-94b1-a6f2ee6193d0" colab={"base_uri": "https://localhost:8080/", "height": 354}
years = [2000, 2004, 2008, 2012, 2016]
data=df.query('Year >= 2000')
fig = plt.figure(figsize=(5, 5))
ax = fig.gca()
ax.scatter('Year', 'Incumbent Votes', data=data)
ax.scatter(x=years, y=y_pred_0, color='red')
ax.set_ylabel('% Total Vote', fontsize=14)
ax.set_xlabel('Year', fontsize=14)
ax.set_xlim(1999, 2017)
for year, pct, act in zip(years, y_pred_0, data['Incumbent Votes']):
plt.text(year + .15, pct - .1, s=f'{pct:.1f}', fontsize=11)
plt.text(year + .15, act - .1, s=f'{act:.1f}', fontsize=11)
plt.title(r"Predicted (red) incumbent vote totals vs acutal totals.", fontsize=14);
# + id="a-QqZCHkO6Jr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="2a182f76-eb0b-4a47-b2c9-410d9ccc42e3"
inter, coef_1, coef_2 = model_0.intercept_, *model_0.coef_
inter, coef_1, coef_2
# + id="uHt8KnLHOm8b" colab_type="code" colab={}
def plot_2_feature_regression(model, features, label, train, test):
feature_1, feature_2 = features
predicted = model.predict(test[features])
intercept, coef_1, coef_2 = model.intercept_, *model.coef_
# Define grid space for plane.
X1 = np.linspace(train[feature_1].min(), train[feature_1].max(), 5)
X2 = np.linspace(train[feature_2].min(), train[feature_2].max(), 5)
X, Y = np.meshgrid(X1, X2)
# Apply model params to linespace values and save in a third matrix.
Z = np.zeros(X.shape)
for r in zip(range(X.shape[0])):
for c in range(X.shape[1]):
Z[r,c] = coef_1 * X[r,c] + coef_2 * Y[r,c] + intercept
# Do plotty things.
fig = plt.figure(figsize=(7, 7))
ax = fig.add_subplot(111, projection='3d')
# Plot points for train set.
ax.scatter3D(train[feature_1], train[feature_2], train[label], color='b')
# Plot points for test set.
ax.scatter3D(test[feature_1], test[feature_2], test[label], color='r')
# Plot predictions.
ax.scatter3D(test[feature_1], test[feature_2], predicted, color='black')
# Draw plane of best fit
ax.plot_surface(X, Y, Z, color='g', alpha=.25)
# Project predictions onto plane.
for x, y, z, label in zip(test[feature_1], test[feature_2], predicted, test[label]):
# I'm fairly certain that coef_1 and coef_2 belong here, not sure of the third param
# or on deciding what order they go in, but 1/intercept keeps everything in the
# ballpark.
best_fit = np.array([coef_1, coef_2, 1/intercept])
our_vector = np.array([x, y, z])
vector_scaler = np.dot(our_vector, best_fit) / np.dot(best_fit, best_fit)
scaled_best_fit = best_fit * vector_scaler
end_points = our_vector - scaled_best_fit
# print('beep')
# print(end_points)
# Plot lines from predicted to true values.
ax.plot((x, x), (y, y), (z, label), color='gray', linestyle='--')
# Plot lines from predicted values to somewhere.
ax.plot((x, end_points[0]), (y, end_points[1]), (z, end_points[2]), color='black', linestyle=':')
ax.set_xlabel(f'{feature_1}')
ax.set_ylabel(f'{feature_2}')
ax.set_zlabel(f'{label}')
# Set veiwing angle.
ax.view_init(15, 120)
# + id="ERFtprhTP_5V" colab_type="code" outputId="b5801b6f-cf45-4a1c-d6a8-<KEY>" colab={"base_uri": "https://localhost:8080/", "height": 411}
features = ['Income growth', 'price index']
plot_2_feature_regression(model_0, features, label, train, test)
# + id="3N8DdbAmra92" colab_type="code" colab={}
| module3-doing-linear-regression/Paul_K_doing_even_more_linear_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Deploy Model
# ## Init Model
# + language="bash"
#
# pio init-model \
# --model-server-url http://prediction-scikit.community.pipeline.io/ \
# --model-type scikit \
# --model-namespace default \
# --model-name scikit_balancescale \
# --model-version v1 \
# --model-path .
# -
# ## Deploy Model (CLI)
# + language="bash"
#
# pio deploy
# -
# ## TODO: Deploy Model (REST)
| jupyterhub/notebooks/scikit/scikit_balancescale/03_DeployModel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import cv2
import numpy as np
def preprocess(img, imgsize, jitter, random_placing=False):
"""
Image preprocess for yolo input
Pad the shorter side of the image and resize to (imgsize, imgsize)
Args:
img (numpy.ndarray): input image whose shape is :math:`(H, W, C)`.
Values range from 0 to 255.
imgsize (int): target image size after pre-processing
jitter (float): amplitude of jitter for resizing
random_placing (bool): if True, place the image at random position
Returns:
img (numpy.ndarray): input image whose shape is :math:`(C, imgsize, imgsize)`.
Values range from 0 to 1.
info_img : tuple of h, w, nh, nw, dx, dy.
h, w (int): original shape of the image
nh, nw (int): shape of the resized image without padding
dx, dy (int): pad size
"""
h, w, _ = img.shape
img = img[:, :, ::-1]
assert img is not None
#尺寸大小的随机抖动,jitter越大,长宽的的变化越大
if jitter > 0:
# add jitter
dw = jitter * w
dh = jitter * h
new_ar = (w + np.random.uniform(low=-dw, high=dw))\
/ (h + np.random.uniform(low=-dh, high=dh))
else:
new_ar = w / h
if new_ar < 1:
nh = imgsize
nw = nh * new_ar
else:
nw = imgsize
nh = nw / new_ar
nw, nh = int(nw), int(nh)
#图像填充位置的随机性
if random_placing:
dx = int(np.random.uniform(imgsize - nw))
dy = int(np.random.uniform(imgsize - nh))
else:
dx = (imgsize - nw) // 2
dy = (imgsize - nh) // 2
img = cv2.resize(img, (nw, nh))
sized = np.ones((imgsize, imgsize, 3), dtype=np.uint8) * 127
sized[dy:dy+nh, dx:dx+nw, :] = img
info_img = (h, w, nh, nw, dx, dy)
return sized, info_img
jitter=0.1
andom_placing=True
img_size=416
img=cv2.imread('data/1.jpg')
# print(img.shape)
# img = cv2.cvtColor(img, cv2.COLOR_RGB2HSV)
# img[:,:,2] = img[:,:,2]+10
# img = cv2.cvtColor(img, cv2.COLOR_HSV2RGB)
# # cv2.imwrite('img_or.jpg',img)
# cv2.imwrite('img_dis3.jpg',img)
sized, info_img=preprocess(img, img_size, jitter=jitter,random_placing=andom_placing)
# print(sized.shape)
# sized=sized[:,:,::-1]
# cv2.imshow('imgs',img)
# cv2.imshow('img',sized)
cv2.waitKey()
def random_distort(img, hue, saturation, exposure):
"""
perform random distortion in the HSV color space.
Args:
img (numpy.ndarray): input image whose shape is :math:`(H, W, C)`.
Values range from 0 to 255.
hue (float): random distortion parameter.
saturation (float): random distortion parameter.
exposure (float): random distortion parameter.
Returns:
img (numpy.ndarray)
"""
#hue 调整色彩度,sat 调整对比度, exp调整亮度
dhue = np.random.uniform(low=-hue, high=hue)
dsat = rand_scale(saturation)
dexp = rand_scale(exposure)
img = cv2.cvtColor(img, cv2.COLOR_RGB2HSV)
img = np.asarray(img, dtype=np.float32) / 255.
img[:, :, 1] *= dsat
img[:, :, 2] *= dexp
H = img[:, :, 0] + dhue
if dhue > 0:
H[H > 1.0] -= 1.0
else:
H[H < 0.0] += 1.0
img[:, :, 0] = H
img = (img * 255).clip(0, 255).astype(np.uint8)
img = cv2.cvtColor(img, cv2.COLOR_HSV2RGB)
img = np.asarray(img, dtype=np.float32)
return img
def rand_scale(s):
"""
calculate
random scaling factor
Args:
s (float): range of the random scale.
Returns:
random scaling factor (float) whose range is
from 1 / s to s .
"""
scale = np.random.uniform(low=1, high=s)
if np.random.rand() > 0.5:
return scale
return 1 / scale
hue=0.2
saturation=1.5
exposure=1.5
img2 = random_distort(sized, hue, saturation, exposure)
sized=sized[:,:,::-1]
# img2 = cv2.cvtColor(img2, cv2.COLOR_HSV2RGB)
img2=img2[:,:,::-1]
cv2.imwrite('img_or.jpg',img)
cv2.imwrite('img_dis.jpg',img2)
# cv2.waitKey()
| image_arguement.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/probml/probml-notebooks/blob/main/notebooks/sg_mcmc_jax.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="CQdfeQ_pOh9S"
# # SG-MCMC-JAX library
#
# https://github.com/jeremiecoullon/SGMCMCJax
#
#
# + [markdown] id="VQ5U7hR8QKbp"
# #Setup
# + id="mJawbDo3g5Dx"
# If running in TPU mode
import jax.tools.colab_tpu
jax.tools.colab_tpu.setup_tpu()
# + id="0fopeFuiObaN"
# %%capture
# !pip install sgmcmcjax
# + colab={"base_uri": "https://localhost:8080/"} id="Qc4tvflueuqM" outputId="7c7b0c1a-7d41-4bca-b148-c92c59695906"
import jax
import jax.numpy as jnp
from jax import jit, grad, vmap, random
print(jax.__version__)
print(jax.devices())
# + id="v4WVl4nqPD1w"
from sgmcmcjax.samplers import build_sgld_sampler
from sgmcmcjax.kernels import build_sgld_kernel, build_psgld_kernel, build_sgldAdam_kernel, build_sghmc_kernel
from tqdm.auto import tqdm
import matplotlib.pyplot as plt
import numpy as np
key = random.PRNGKey(0)
# + [markdown] id="nm1WIaOCOzod"
# # Gaussian posterior
#
# https://sgmcmcjax.readthedocs.io/en/latest/nbs/gaussian.html
# + id="PY7HiHcAOicY"
# define model in JAX
def loglikelihood(theta, x):
return -0.5*jnp.dot(x-theta, x-theta)
def logprior(theta):
return -0.5*jnp.dot(theta, theta)*0.01
# generate dataset
N, D = 10_000, 100
key = random.PRNGKey(0)
mu_true = random.normal(key, shape=(D,))
X_data = random.normal(key, shape=(N, D)) + mu_true
# build sampler
batch_size = int(0.1*N)
dt = 1e-5
my_sampler = build_sgld_sampler(dt, loglikelihood, logprior, (X_data,), batch_size)
# + colab={"base_uri": "https://localhost:8080/", "height": 83, "referenced_widgets": ["06ea1f92111d42c59f5743f7f19ddecd", "31d5332f37ac479d9af2fb057f62919a", "08104e55ab654e6e879a690db22c330c", "044100d715714b419e185d40d82e466e", "143541dd2f7a4f1190c31124c2e1a4b9", "a600713bb198461392c65c1f4f81b885", "<KEY>", "3551fc911d954ad8b20c25d8957b3a6e", "a87ad53eea084f00ad0cd510f1283859", "8e2c1822d9884a76b54d0b8377baac64", "692f3d95b0664943817257244324af24"]} id="C4saWTbOPCsa" outputId="9a1f211f-3661-4538-92bc-a0e91ef8805f"
# %%time
Nsamples = 10_000
samples = my_sampler(key, Nsamples, jnp.zeros(D))
# + colab={"base_uri": "https://localhost:8080/"} id="uuwhhcWGPHtg" outputId="3c61568a-eb0b-474b-b994-b3ec3a918e54"
print(samples.shape)
mu_est = jnp.mean(samples, axis=0)
print(jnp.allclose(mu_true, mu_est, atol=1e-1))
print(mu_true[:10])
print(mu_est[:10])
# + colab={"base_uri": "https://localhost:8080/", "height": 341} id="D68_hyWjXaZY" outputId="52b8c09c-1e01-4742-fafa-eb78a5634d95"
data = (X_data,)
init_fn, sgld_kernel, get_params = build_sgld_kernel(dt, loglikelihood, logprior, data, batch_size)
key, subkey = random.split(key)
params = random.normal(subkey, shape=(D,))
key, subkey = random.split(key)
state = init_fn(subkey, params)
print(state)
# + [markdown] id="IE268RvOQaRn"
# # Logistic regression
#
# https://sgmcmcjax.readthedocs.io/en/latest/nbs/logistic_regression.html
# + colab={"base_uri": "https://localhost:8080/", "height": 368} id="mz6uwMYfP6h7" outputId="b2575f27-03f3-4a42-a26d-69530aba264c"
from models.logistic_regression import gen_data, loglikelihood, logprior
key = random.PRNGKey(42)
dim = 10
Ndata = 100000
theta_true, X, y_data = gen_data(key, dim, Ndata)
# + [markdown] id="dh7cMKRaQknp"
# # Bayesian neural network
#
# https://sgmcmcjax.readthedocs.io/en/latest/nbs/BNN.html
# + colab={"base_uri": "https://localhost:8080/", "height": 368} id="KkvqqmuPQelA" outputId="dc5c931e-6842-4245-e6f1-08a2513002a8"
from models.bayesian_NN.NN_data import X_train, X_test, y_train, y_test
from models.bayesian_NN.NN_model import init_network, loglikelihood, logprior, accuracy
from sgmcmcjax.kernels import build_sgld_kernel
from tqdm.auto import tqdm
# + [markdown] id="9hchex74Qwf9"
# # FLAX CNN
#
# https://sgmcmcjax.readthedocs.io/en/latest/nbs/Flax_MNIST.html
# + id="mOK2K-J2REyH"
# %%capture
# !pip install --upgrade git+https://github.com/google/flax.git
# + id="iy3WEKDyQmfw"
import tensorflow_datasets as tfds
from flax import linen as nn
# + id="D-g1gVeSRqa1"
class CNN(nn.Module):
"""A simple CNN model."""
@nn.compact
def __call__(self, x):
x = nn.Conv(features=32, kernel_size=(3, 3))(x)
x = nn.relu(x)
x = nn.avg_pool(x, window_shape=(2, 2), strides=(2, 2))
x = nn.Conv(features=64, kernel_size=(3, 3))(x)
x = nn.relu(x)
x = nn.avg_pool(x, window_shape=(2, 2), strides=(2, 2))
x = x.reshape((x.shape[0], -1)) # flatten
x = nn.Dense(features=256)(x)
x = nn.relu(x)
x = nn.Dense(features=10)(x)
x = nn.log_softmax(x)
return x
# + colab={"base_uri": "https://localhost:8080/", "height": 273, "referenced_widgets": ["0fa5bf6d413d49da90cfdf6bc1c93ad7", "836a784d8eff4de490a1466c51145e5c", "72149a1caaa340ecb0bd1541ed3966af", "d3d0416899f349e79e40a35ca96304aa", "e2edef161ef14aac91f1df4aa97b82e4", "e00004a6fdf14570b808fa4cd2e45696", "8ab28b5ef74d4cd9933430b42796e760", "<KEY>", "3c127d0bc41c42ca8bad12c7efe2cf83", "<KEY>", "fc4c9953df514699bd4a07551a8587af"]} id="e2WL0nyNRqse" outputId="aec3d2b1-39b2-4921-b35c-7fc185ee1bb1"
cnn = CNN()
def loglikelihood(params, x, y):
x = x[jnp.newaxis] # add an axis so that it works for a single data point
logits = cnn.apply({'params':(params)}, x)
label = jax.nn.one_hot(y, num_classes=10)
return jnp.sum(logits*label)
def logprior(params):
return 1.
@jit
def accuracy_cnn(params, X, y):
target_class = y
predicted_class = jnp.argmax(cnn.apply({'params':(params)}, X), axis=1)
return jnp.mean(predicted_class == target_class)
def get_datasets():
"""Load MNIST train and test datasets into memory."""
ds_builder = tfds.builder('mnist')
ds_builder.download_and_prepare()
train_ds = tfds.as_numpy(ds_builder.as_dataset(split='train', batch_size=-1))
test_ds = tfds.as_numpy(ds_builder.as_dataset(split='test', batch_size=-1))
train_ds['image'] = jnp.float32(train_ds['image']) / 255.
test_ds['image'] = jnp.float32(test_ds['image']) / 255.
return train_ds, test_ds
train_ds, test_ds = get_datasets()
X_train_s = train_ds['image']
y_train_s = jnp.array(train_ds['label'])
X_test_s = test_ds['image']
y_test_s = jnp.array(test_ds['label'])
data = (X_train_s, y_train_s)
batch_size = int(0.01*data[0].shape[0])
print(batch_size)
# + colab={"base_uri": "https://localhost:8080/"} id="4FyicuEFTUJu" outputId="00c2509c-6457-4142-fb23-d6acb32af628"
params = cnn.init(key, jnp.ones([1,28,28,1]))
print(params.keys())
params = params['params']
print(params['Dense_0']['kernel'].shape)
print(params['Dense_1']['kernel'].shape)
# + id="WjzaI5agRy8P"
def run_sgmcmc(key, Nsamples, init_fn, my_kernel, get_params,
record_accuracy_every=50):
"Run SGMCMC sampler and return the test accuracy list"
accuracy_list = []
params = cnn.init(key, jnp.ones([1,28,28,1]))['params']
key, subkey = random.split(key)
state = init_fn(subkey, params)
for i in tqdm(range(Nsamples)):
key, subkey = random.split(key)
state = my_kernel(i, subkey, state)
if i % record_accuracy_every==0:
test_acc = accuracy_cnn(get_params(state), X_test_s, y_test_s)
accuracy_list.append((i, test_acc))
return accuracy_list
# + id="7zV76Cc3co-M"
kernel_dict = {}
# SGLD
init_fn, kernel, get_params = build_sgld_kernel(5e-6, loglikelihood, logprior, data, batch_size)
kernel = jit(kernel)
kernel_dict['sgld'] = (init_fn, kernel, get_params)
# SG-HMC
init_fn, kernel, get_params = build_sghmc_kernel(
1e-6, 4, loglikelihood, logprior, data, batch_size, compiled_leapfrog=False)
kernel = jit(kernel)
kernel_dict['sghmc'] = (init_fn, kernel, get_params)
# PSGLD
init_fn, kernel, get_params = build_psgld_kernel(1e-3, loglikelihood, logprior, data, batch_size)
kernel = jit(kernel)
kernel_dict['psgld'] = (init_fn, kernel, get_params)
# SGLD-Adam
init_fn, kernel, get_params = build_sgldAdam_kernel(1e-2, loglikelihood, logprior, data, batch_size)
kernel = jit(kernel)
kernel_dict['sgldAdam'] = (init_fn, kernel, get_params)
# + colab={"base_uri": "https://localhost:8080/", "height": 301, "referenced_widgets": ["515eee086faa4af192a4f15b732cdc12", "11d6cf780e094a74820cb1f4a59f82c2", "ccacbc5cf85c4f58ba6b04c1bd10f44a", "27463680b4964bb9b7cbfd6e3810a0a7", "a98650350f0245d4ad43eeb67db4d475", "<KEY>", "d74b51168bfc468ab6077ac1a3e9b510", "956879b98b6543a3b77edb3f13519d2a", "<KEY>", "c9f56d7307fa4cc69ec42eabae759d3f", "0d85e43f2cf8466c8620e59ae2150873", "00ce0f32a72d47b6b9ff3ea81baaba06", "37a1fa07d5114025a67808fe91d63d10", "<KEY>", "533fc883a3514a1ebad42b986435185e", "<KEY>", "<KEY>", "<KEY>", "1e6755b199c548e7a793ba287c563673", "5ac5e621d9b743678efd266c2a183ddd", "<KEY>", "<KEY>", "78771cfa75b54d5fb94e9d3874426398", "<KEY>", "<KEY>", "30d2304c9d4d43fca97feae501a52f1b", "<KEY>", "3dd51604e7e941eb99644b251ae461e6", "60998fdf80ef4cad96dadda247ae038b", "<KEY>", "ea7c18e53f174f4f9fb6f13797f851f9", "<KEY>", "d0173557582c4cd189745d4700ec42ef", "c20aaacdea57496cb7a1be74a524966a", "<KEY>", "<KEY>", "e98f2e686d184706ae55f1a756c3112a", "<KEY>", "<KEY>", "22f8e96a941b4fe591734c48755468be", "7a8c03f483ac44f796d4795c96ea5b46", "0dfb815cdcea47d3adcb5604ef0a1241", "e5450ca369044b3480f42568a05003ef", "798ce80bf7104f55affa9dd149f19c4e"]} id="vcKM1NGCdSmm" outputId="8de4010e-90a0-404b-ad9d-c2ba9b95e937"
Nsamples = 500
acc_dict = {}
for (name, fns) in kernel_dict.items():
print(name)
init_fn, kernel, get_params = fns
accuracy_list = run_sgmcmc(random.PRNGKey(0), Nsamples, init_fn, kernel, get_params)
print(accuracy_list)
acc_dict[name] = accuracy_list
# + colab={"base_uri": "https://localhost:8080/", "height": 321} id="q11jhQ2IjCdw" outputId="941e23ba-d27d-468e-827c-3a8c66435f86"
plt.figure()
for name, acc_list in acc_dict.items():
steps = [s for (s,a) in acc_list]
accs = [a for (s,a) in acc_list]
plt.plot(steps, accs, '-o', label=name)
plt.legend(fontsize=16)
plt.title("Test accuracy of SGMCMC samplers on MNIST with a CNN")
plt.xlabel("Iterations", size=20)
plt.ylabel("Test accuracy", size=20)
| notebooks/sg_mcmc_jax.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Sequence-to-sequence LSTM outlier detector deployment
# Wrap a keras seq2seq-LSTM python model for use as a prediction microservice in seldon-core and deploy on seldon-core running on Minikube or a Kubernetes cluster using GCP.
# ## Dependencies
# - [helm](https://github.com/helm/helm)
# - [minikube](https://github.com/kubernetes/minikube)
# - [s2i](https://github.com/openshift/source-to-image) >= 1.1.13
#
# Python packages:
# - keras: pip install keras
# - tensorflow: https://www.tensorflow.org/install/pip
# ## Task
# The outlier detector needs to spot anomalies in electrocardiograms (ECG's). The dataset "ECG5000" contains 5000 ECG's, originally obtained from [Physionet](https://physionet.org/cgi-bin/atm/ATM) under the name "BIDMC Congestive Heart Failure Database(chfdb)", record "chf07". The data has been pre-processed in 2 steps: first each heartbeat is extracted, and then each beat is made equal length via interpolation. The data is labeled and contains 5 classes. The first class which contains almost 60% of the observations is seen as "normal" while the others are outliers. The seq2seq-LSTM algorithm is trained on some heartbeats from the first class and needs to flag the other classes as anomalies. The plot below shows an example ECG for each of the classes.
# 
# ## Train locally
#
# Train on some inlier ECG's. The data can be downloaded [here](http://www.timeseriesclassification.com/description.php?Dataset=ECG5000) and should be extracted in the [data](./data) folder.
# !python train.py \
# --dataset './data/ECG5000_TEST.arff' \
# --data_range 0 2627 \
# --minmax \
# --timesteps 140 \
# --encoder_dim 20 \
# --decoder_dim 40 \
# --output_activation 'sigmoid' \
# --dropout 0 \
# --learning_rate 0.005 \
# --loss 'mean_squared_error' \
# --epochs 100 \
# --batch_size 32 \
# --validation_split 0.2 \
# --print_progress \
# --save
# The plot below shows a typical prediction (*red line*) of an inlier (class 1) ECG compared to the original (*blue line*) after training the seq2seq-LSTM model.
# 
# On the other hand, the model is not good at fitting ECG's from the other classes, as illustrated in the chart below:
# 
# The predictions in the above charts are made on ECG's the model has not seen before. The differences in scale are due to the sigmoid output layer and do not affect the prediction accuracy.
# ## Test using Kubernetes cluster on GCP or Minikube
# Run the outlier detector as a model or a transformer. If you want to run the anomaly detector as a transformer, change the SERVICE_TYPE variable from MODEL to TRANSFORMER [here](./.s2i/environment), set MODEL = False and change ```OutlierSeq2SeqLSTM.py``` to:
#
# ```python
# from CoreSeq2SeqLSTM import CoreSeq2SeqLSTM
#
# class OutlierSeq2SeqLSTM(CoreSeq2SeqLSTM):
# """ Outlier detection using a sequence-to-sequence (seq2seq) LSTM model.
#
# Parameters
# ----------
# threshold (float) : reconstruction error (mse) threshold used to classify outliers
# reservoir_size (int) : number of observations kept in memory using reservoir sampling
# """
# def __init__(self,threshold=0.003,reservoir_size=50000,model_name='seq2seq',load_path='./models/'):
#
# super().__init__(threshold=threshold,reservoir_size=reservoir_size,
# model_name=model_name,load_path=load_path)
# ```
MODEL = True
# Pick Kubernetes cluster on GCP or Minikube.
MINIKUBE = True
if MINIKUBE:
# !minikube start --memory 4096
else:
# !gcloud container clusters get-credentials standard-cluster-1 --zone europe-west1-b --project seldon-demos
# Create a cluster-wide cluster-admin role assigned to a service account named “default” in the namespace “kube-system”.
# !kubectl create clusterrolebinding kube-system-cluster-admin --clusterrole=cluster-admin \
# --serviceaccount=kube-system:default
# !kubectl create namespace seldon
# Add current context details to the configuration file in the seldon namespace.
# !kubectl config set-context $(kubectl config current-context) --namespace=seldon
# Create tiller service account and give it a cluster-wide cluster-admin role.
# !kubectl -n kube-system create sa tiller
# !kubectl create clusterrolebinding tiller --clusterrole cluster-admin --serviceaccount=kube-system:tiller
# !helm init --service-account tiller
# Check deployment rollout status and deploy seldon/spartakus helm charts.
# !kubectl rollout status deploy/tiller-deploy -n kube-system
# !helm install ../../../helm-charts/seldon-core-operator --name seldon-core --set usage_metrics.enabled=true --namespace seldon-system
# Check deployment rollout status for seldon core.
# !kubectl rollout status deploy/seldon-controller-manager -n seldon-system
# Install Ambassador API gateway
# !helm install stable/ambassador --name ambassador --set crds.keep=false
# !kubectl rollout status deployment.apps/ambassador
# If Minikube used: create docker image for outlier detector inside Minikube using s2i. Besides the transformer image and the demo specific model image, the general model image for the Seq2Seq LSTM outlier detector is also available from Docker Hub as ***seldonio/outlier-s2s-lstm-model:0.1***.
if MINIKUBE & MODEL:
# !eval $(minikube docker-env) && \
# s2i build . seldonio/seldon-core-s2i-python3:0.4 seldonio/outlier-s2s-lstm-model-demo:0.1
elif MINIKUBE:
# !eval $(minikube docker-env) && \
# s2i build . seldonio/seldon-core-s2i-python3:0.4 seldonio/outlier-s2s-lstm-transformer:0.1
# Install outlier detector helm charts and set *threshold* and *reservoir_size* hyperparameter values.
if MODEL:
# !helm install ../../../helm-charts/seldon-od-model \
# --name outlier-detector \
# --namespace=seldon \
# --set model.type=seq2seq \
# --set model.seq2seq.image.name=seldonio/outlier-s2s-lstm-model-demo:0.1 \
# --set model.seq2seq.threshold=0.002 \
# --set model.seq2seq.reservoir_size=50000 \
# --set oauth.key=oauth-key \
# --set oauth.secret=oauth-secret \
# --set replicas=1
else:
# !helm install ../../../helm-charts/seldon-od-transformer \
# --name outlier-detector \
# --namespace=seldon \
# --set outlierDetection.enabled=true \
# --set outlierDetection.name=outlier-s2s-lstm \
# --set outlierDetection.type=seq2seq \
# --set outlierDetection.seq2seq.image.name=seldonio/outlier-s2s-lstm-transformer:0.1 \
# --set outlierDetection.seq2seq.threshold=0.002 \
# --set outlierDetection.seq2seq.reservoir_size=50000 \
# --set oauth.key=oauth-key \
# --set oauth.secret=oauth-secret \
# --set model.image.name=seldonio/outlier-s2s-lstm-model:0.1
# ## Port forward Ambassador
#
# Run command in terminal:
# ```
# kubectl port-forward $(kubectl get pods -n seldon -l app.kubernetes.io/name=ambassador -o jsonpath='{.items[0].metadata.name}') -n seldon 8003:8080
# ```
# ## Import rest requests, load data and test requests
# +
import numpy as np
from utils import get_payload, rest_request_ambassador, send_feedback_rest, ecg_data
ecg_data, ecg_labels = ecg_data(dataset='TRAIN')
X = ecg_data[0,:].reshape(1,ecg_data.shape[1],1)
label = ecg_labels[0].reshape(1)
print(X.shape)
print(label.shape)
# -
# Test the rest requests with the generated data. It is important that the order of requests is respected. First we make predictions, then we get the "true" labels back using the feedback request. If we do not respect the order and eg keep making predictions without getting the feedback for each prediction, there will be a mismatch between the predicted and "true" labels. This will result in errors in the produced metrics.
request = get_payload(X)
response = rest_request_ambassador("outlier-detector","seldon",request,endpoint="localhost:8003")
# If the outlier detector is used as a transformer, the output of the anomaly detection is added as part of the metadata. If it is used as a model, we send model feedback to retrieve custom performance metrics.
if MODEL:
send_feedback_rest("outlier-detector","seldon",request,response,0,label,endpoint="localhost:8003")
# ## Analytics
# Install the helm charts for prometheus and the grafana dashboard
# !helm install ../../../helm-charts/seldon-core-analytics --name seldon-core-analytics \
# --set grafana_prom_admin_password=password \
# --set persistence.enabled=false \
# --namespace seldon
# ## Port forward Grafana dashboard
# Run command in terminal:
# ```
# kubectl port-forward $(kubectl get pods -n seldon -l app=grafana-prom-server -o jsonpath='{.items[0].metadata.name}') -n seldon 3000:3000
# ```
# You can then view an analytics dashboard inside the cluster at http://localhost:3000/dashboard/db/prediction-analytics?refresh=5s&orgId=1. Your IP address may be different. get it via minikube ip. Login with:
#
# Username : admin
#
# password : password (as set when starting seldon-core-analytics above)
#
# Import the outlier-detector-s2s-lstm dashboard from ../../../helm-charts/seldon-core-analytics/files/grafana/configs.
# ## Run simulation
#
# - Sample random ECG from dataset.
# - Get payload for the observation.
# - Make a prediction.
# - Send the "true" label with the feedback if the detector is run as a model.
#
# It is important that the prediction-feedback order is maintained. Otherwise there will be a mismatch between the predicted and "true" labels.
#
# View the progress on the grafana "Outlier Detection" dashboard. Most metrics need the outlier detector to be run as a model since they need model feedback.
import time
n_requests = 100
n_samples, sample_length = ecg_data.shape
for i in range(n_requests):
idx = np.random.choice(n_samples)
X = ecg_data[idx,:].reshape(1,sample_length,1)
label = ecg_labels[idx].reshape(1)
request = get_payload(X)
response = rest_request_ambassador("outlier-detector","seldon",request,endpoint="localhost:8003")
if MODEL:
send_feedback_rest("outlier-detector","seldon",request,response,0,label,endpoint="localhost:8003")
time.sleep(1)
if MINIKUBE:
# !minikube delete
| components/outlier-detection/seq2seq-lstm/seq2seq_lstm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import torch
from torch.autograd import Variable
from torchvision.datasets import MNIST
from torchvision.transforms import ToTensor
from capsule_net import CapsNetWithReconstruction, CapsNet, ReconstructionNet
# initialize network classes
capsnet = CapsNet(3, 10)
reconstructionnet = ReconstructionNet(16, 10)
model = CapsNetWithReconstruction(capsnet, reconstructionnet)
# Load trained model
MODEL_PATH = 'checkpoints/250_model_dict_3routing_reconstructionTrue.pth'
model.load_state_dict(torch.load(MODEL_PATH))
dataset = MNIST('../data/MNIST', train=False, transform=ToTensor())
# +
# (1x28x28 tensor input)
def get_digit_caps(model, image):
input_ = Variable(image.unsqueeze(0), volatile=True)
digit_caps, probs = model.capsnet(input_)
return digit_caps
# takes digit_caps output and target label
def get_reconstruction(model, digit_caps, label):
target = Variable(torch.LongTensor([label]), volatile=True)
reconstruction = model.reconstruction_net(digit_caps, target)
return reconstruction.data.cpu().numpy()[0].reshape(28, 28)
# create reconstructions with perturbed digit capsule
def dimension_perturbation_reconstructions(model, digit_caps, label, dimension, dim_values):
reconstructions = []
for dim_value in dim_values:
digit_caps_perturbed = digit_caps.clone()
digit_caps_perturbed[0, label, dimension] = dim_value
reconstruction = get_reconstruction(model, digit_caps_perturbed, label)
reconstructions.append(reconstruction)
return reconstructions
# -
# %matplotlib inline
import matplotlib.pyplot as plt
# Get reconstructions
images = []
reconstructions = []
for i in range(8):
image_tensor, label = dataset[i]
digit_caps = get_digit_caps(model, image_tensor)
reconstruction = get_reconstruction(model, digit_caps, label)
images.append(image_tensor.numpy()[0])
reconstructions.append(reconstruction)
# Plot reconstructions
fig, axs = plt.subplots(2, 8, figsize=(16, 4))
axs[0, 0].set_ylabel('Org image', size='large')
axs[1, 0].set_ylabel('Reconstruction', size='large')
for i in range(8):
axs[0, i].imshow(images[i], cmap='gray')
axs[1, i].imshow(reconstructions[i], cmap='gray')
axs[0, i].set_yticks([])
axs[0, i].set_xticks([])
axs[1, i].set_yticks([])
axs[1, i].set_xticks([])
digit, label = dataset[0]
perturbed_reconstructions = []
perturbation_values = [0.05*i for i in range(-5, 6)]
digit_caps = get_digit_caps(model, digit)
for dimension in range(16):
perturbed_reconstructions.append(
dimension_perturbation_reconstructions(model, digit_caps, label,
dimension, perturbation_values)
)
fig, axs = plt.subplots(16, 11, figsize=(11*1.5, 16*1.5))
for i in range(16):
axs[i, 0].set_ylabel('dim {}'.format(i), size='large')
for j in range(11):
axs[i, j].imshow(perturbed_reconstructions[i][j], cmap='gray')
axs[i, j].set_yticks([])
axs[i, j].set_xticks([])
| capsule_viz.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tensorflow
# language: python
# name: tensorflow
# ---
# +
import tensorflow as tf
import tensorflow.examples.tutorials.mnist.input_data as input_data
# Config params
learning_rate = 0.1
logs_path = "/tmp/mnist_act/"
training_epochs = 50
batch_size = 1024
n_input = 784
n_classes = 10
# Load mnist data set (train, validation, test)
mnist = input_data.read_data_sets("MNIST_data", one_hot=True)
# Input
with tf.name_scope("Input"):
X = tf.placeholder(tf.float32, shape=[None, n_input], name="input_X")
Y = tf.placeholder(tf.float32, shape=[None, n_classes], name="labels_Y")
# Attach a lot of summaries to a Tensor
def variable_summaries(var):
with tf.name_scope('summaries'):
mean = tf.reduce_mean(var)
tf.summary.scalar('mean', mean)
with tf.name_scope('stddev'):
stddev = tf.sqrt(tf.reduce_mean(tf.square(var - mean)))
tf.summary.scalar('stddev', stddev)
tf.summary.scalar('max', tf.reduce_max(var))
tf.summary.scalar('min', tf.reduce_min(var))
tf.summary.histogram('histogram', var)
# Create a layer in neural network (Weights, bias, activations
def nn_layer(input_tensor, input_dim, output_dim, layer_name, act=tf.nn.sigmoid):
with tf.name_scope(layer_name):
with tf.name_scope('weights'):
weights = tf.Variable(tf.random_normal([input_dim, output_dim]), name="Weights")
#weights = tf.get_variable(shape = [input_dim, output_dim], name=layer_name+"_Weights", initializer=tf.contrib.layers.xavier_initializer(uniform=True, seed=None, dtype=tf.float32))
variable_summaries(weights)
with tf.name_scope('biases'):
biases = tf.Variable(tf.random_normal([1, output_dim]), name="biases")
variable_summaries(biases)
with tf.name_scope('Wx_plus_b'):
preactivate = tf.matmul(input_tensor, weights) + biases
tf.summary.histogram('pre_activations', preactivate)
activations = act(preactivate, name='activation')
tf.summary.histogram('activations', activations)
return activations
# Define Layers
hidden_1 = nn_layer(X, n_input, 16, 'layer_1')
hidden_2 = nn_layer(hidden_1, 16, 16, 'layer_2')
output_layer = nn_layer(hidden_2, 16, 10, 'output', act=tf.identity)
# Few metrics
with tf.name_scope('metrics'):
yHat = tf.nn.softmax(output_layer)
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits=output_layer, labels=Y))
correct_prediction = tf.equal(tf.argmax(Y, 1), tf.argmax(yHat, 1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
tf.summary.scalar("cost", cost)
tf.summary.scalar("accuracy", accuracy)
# Optimizer
with tf.name_scope('train'):
optimizer = tf.train.GradientDescentOptimizer(learning_rate)
train_step = optimizer.minimize(cost)
# Create session
sess = tf.Session()
sess.run(tf.global_variables_initializer())
writer = tf.summary.FileWriter(logs_path, graph=tf.get_default_graph())
# Merge all the summaries and write them out to /tmp/mnist_logs (by default)
metrics = tf.summary.merge_all()
# Train model
for epoch in range(training_epochs):
total_batches = int(mnist.train.num_examples/batch_size)
for batch in range(total_batches):
batch_xs, batch_ys = mnist.train.next_batch(batch_size)
cv, _, summary = sess.run([cost, train_step, metrics], feed_dict={X:batch_xs, Y:batch_ys})
a = sess.run([accuracy], feed_dict={X:mnist.test.images, Y:mnist.test.labels})
# Generate summary event for TensorBoard visualization
step = epoch * total_batches + batch
writer.add_summary(summary, step)
print (epoch, batch, cv, a)
# -
| ml/.ipynb_checkpoints/multi_layer_mnist_tb-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 自定义层
from __future__ import absolute_import, division, print_function
import tensorflow as tf
# ## 网络层layer的常见操作
# 通常机器学习模型可以表示为简单网络层的堆叠与组合,而tensorflow就提供了常见的网络层,为我们编写神经网络程序提供了便利。 TensorFlow2推荐使用tf.keras来构建网络层,tf.keras来自原生keras,用其来构建网络具有更好的可读性和易用性。
# +
layer = tf.keras.layers.Dense(100)
# 也可以添加输入维度限制
layer = tf.keras.layers.Dense(100, input_shape=(None, 20))
# -
# 可以在文档中查看预先存在的图层的完整列表。 它包括Dense,Conv2D,LSTM,BatchNormalization,Dropout等等。
#
# 每个层都可以当作一个函数,然后以输入的数据作为函数的输入
# +
layer(tf.ones([6, 6]))
print(layer.variables) # 包含了权重和偏置
# -
print(layer.kernel, layer.bias) # 也可以分别取出权重和偏置
# ## 实现自定义网络层
# 实现自己的层的最佳方法是扩展tf.keras.Layer类并实现:
#
# - ```__init__()```函数,你可以在其中执行所有与输入无关的初始化
# - ```build()```函数,可以获得输入张量的形状,并可以进行其余的初始化
# - ```call()```函数,构建网络结构,进行前向传播
#
# 实际上,你不必等到调用```build()```来创建网络结构,您也可以在 ```__init__()``` 中创建它们。 但是,在```build()```中创建它们的优点是它可以根据图层将要操作的输入的形状启用后期的网络构建。 另一方面,在 ```__init__``` 中创建变量意味着需要明确指定创建变量所需的形状。
# +
class MyDense(tf.keras.layers.Layer):
def __init__(self, n_outputs):
super(MyDense, self).__init__()
self.n_outputs = n_outputs
def build(self, input_shape):
self.kernel = self.add_variable('kernel',
shape=[int(input_shape[-1]),
self.n_outputs])
def call(self, input):
return tf.matmul(input, self.kernel)
layer = MyDense(10)
print(layer(tf.ones([6, 5])))
print(layer.trainable_variables)
# -
# ## 网络层组合
# 机器学习模型中有很多是通过叠加不同的结构层组合而成的,如resnet的每个残差块就是“卷积+批标准化+残差连接”的组合。
#
# 在tensorflow2中要创建一个包含多个网络层的的结构,一般继承与tf.keras.Model类。
# +
# 残差块
class ResnetBlock(tf.keras.Model):
def __init__(self, kernel_size, filters):
super(ResnetBlock, self).__init__(name='resnet_block')
# 每个子层卷积核数
filter1, filter2, filter3 = filters
# 三个子层,每层1个卷积加一个批正则化
# 第一个子层, 1*1的卷积
self.conv1 = tf.keras.layers.Conv2D(filter1, (1,1))
self.bn1 = tf.keras.layers.BatchNormalization()
# 第二个子层, 使用特点的kernel_size
self.conv2 = tf.keras.layers.Conv2D(filter2, kernel_size, padding='same')
self.bn2 = tf.keras.layers.BatchNormalization()
# 第三个子层,1*1卷积
self.conv3 = tf.keras.layers.Conv2D(filter3, (1,1))
self.bn3 = tf.keras.layers.BatchNormalization()
def call(self, inputs, training=False):
# 堆叠每个子层
x = self.conv1(inputs)
x = self.bn1(x, training=training)
x = self.conv2(x)
x = self.bn2(x, training=training)
x = self.conv3(x)
x = self.bn3(x, training=training)
# 残差连接
x += inputs
outputs = tf.nn.relu(x)
return outputs
resnetBlock = ResnetBlock(2, [6,4,9])
# 数据测试
print(resnetBlock(tf.ones([1,3,9,9])))
# 查看网络中的变量名
print([x.name for x in resnetBlock.trainable_variables])
# -
| 202_user-defined_layer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1 - Risk and return models
#
#
# In this section, we compare how well the different risk models predict an out-of-sample covariance matrix, and how well the different returns models predict out-of-sample returns.
#
# ## Risk models
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import pypfopt
from pypfopt import risk_models, expected_returns, plotting
pypfopt.__version__
# +
df = pd.read_csv("data/stock_prices.csv", parse_dates=True, index_col="date")
past_df, future_df = df.iloc[:-250], df.iloc[-250:]
future_cov = risk_models.sample_cov(future_df)
sample_cov = risk_models.sample_cov(past_df)
plotting.plot_covariance(sample_cov)
plotting.plot_covariance(future_cov)
plt.show()
# -
# We can see that visually, the sample covariance does not capture some of the new features of the covariance matrix, for example the highly correlated group of FAANG stocks. We may be able to improve this by using an exponentially-weighted covariance matrix, which gives more weight to recent data. We can also look at how each model predicts future variance.
# +
future_variance = np.diag(future_cov)
mean_abs_errors = []
risk_methods = [
"sample_cov",
"semicovariance",
"exp_cov",
"ledoit_wolf",
"ledoit_wolf_constant_variance",
"ledoit_wolf_single_factor",
"ledoit_wolf_constant_correlation",
"oracle_approximating",
]
for method in risk_methods:
S = risk_models.risk_matrix(df, method=method)
variance = np.diag(S)
mean_abs_errors.append(np.sum(np.abs(variance - future_variance)) / len(variance))
xrange = range(len(mean_abs_errors))
plt.barh(xrange, mean_abs_errors)
plt.yticks(xrange, risk_methods)
plt.show()
# -
# We can see that the exponential covariance matrix is a much better estimator of future variance compared to the other models. Its mean absolute error is 2%, which is actually pretty good. Let's visually compare the exponential cov matrix to the realised future cov matrix:
exp_cov = risk_models.exp_cov(past_df)
plotting.plot_covariance(exp_cov)
plotting.plot_covariance(future_cov)
plt.show()
# ## Returns
#
# What about returns? Will the exponentially-weighted returns similarly be the best performer?
# +
future_rets = expected_returns.mean_historical_return(future_df)
mean_abs_errors = []
return_methods = [
"mean_historical_return",
"ema_historical_return",
"james_stein_shrinkage",
"capm_return",
]
for method in return_methods:
mu = expected_returns.return_model(past_df, method=method)
mean_abs_errors.append(np.sum(np.abs(mu - future_rets)) / len(mu))
xrange = range(len(mean_abs_errors))
plt.barh(xrange, mean_abs_errors)
plt.yticks(xrange, return_methods)
plt.show()
# -
# The exponential moving average is marginally better than the others, but the improvement is almost unnoticeable. We also note that the mean absolute deviations are above 25%, meaning that if your expected annual returns are 10%, on average the realised annual return could be anywhere from a 15% loss to a 35% gain. This is a massive range, and gives some context to the advice in the docs suggesting that you optimise without providing an estimate of returns.
# +
fig, axs = plt.subplots( 1, len(return_methods),sharey=True, figsize=(15,10))
for i, method in enumerate(return_methods):
mu = expected_returns.return_model(past_df, method=method)
axs[i].set_title(method)
mu.plot.barh(ax=axs[i])
# -
# The good news is that we see a good degree of agreement (apart from the `ema` method). Between the three methods. We also observe that James-Stein srhinkage is quite conservative in shrinking asset prices to the grand mean, resulting in less variation than simply taking the mean.
| cookbook/1-RiskReturnModels.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # EGM722 - Week 5 Practical: Vector and raster operations using python
#
# ## Overview
#
# Up to now, we have worked with either vector data or raster data, but we haven't really used them together. In this week's practical, we'll learn how we can combine these two data types, and see some examples of different analyses, such as zonal statistics or sampling raster data, that we can automate using python.
#
# ## Objectives
# - learn how to use `rasterstats` to perform zonal statistics
# - learn how to handle exceptions using try...except
# - rasterize polygon data using `rasterio`
# - learn how to mask and select (index) rasters using vector data
# - see additional plotting examples using matplotlib
#
# ## Data provided
#
# In the data\_files folder, you should have the following:
# - LCM2015_Aggregate_100m.tif
# - NI_DEM.tif
#
#
# ## 1. Getting started
#
# In this practical, we'll look at a number of different GIS tasks related to working with both raster and vector data in python, as well as a few different python and programming concepts. To get started, run the cell below.
# +
# %matplotlib notebook
import numpy as np
import rasterio as rio
import geopandas as gpd
import matplotlib.pyplot as plt
from rasterstats import zonal_stats
plt.rcParams.update({'font.size': 22}) # update the font size for our plots to be size 22
# -
# ## 2. Zonal statistics
# In GIS, [_zonal statistics_](https://pro.arcgis.com/en/pro-app/latest/tool-reference/spatial-analyst/how-zonal-statistics-works.htm) is a process whereby you calculate statistics for the pixels of a raster in different groups, or zones, defined by properties in another dataset. In this example, we're going to use the Northern Ireland County border dataset from Week 2, along with a re-classified version of the Northern Ireland [Land Cover Map](https://catalogue.ceh.ac.uk/documents/47f053a0-e34f-4534-a843-76f0a0998a2f) 2015[<sup id="fn1-back">1</sup>](#fn1 "footnote 1").
#
# The Land Cover Map tells, for each pixel, what type of land cover is associated with a location - that is, whether it's woodland (and what kind of woodland), grassland, urban or built-up areas, and so on. For our re-classified version of the dataset, we're working with the aggregate class data, re-sampled to 100m resolution from the original 25m resolution.
#
# The raster data type is _unsigned integer_ with a _bitdepth_ of 8 bits - that is, it has a range of possible values from 0 to 255. Even though it has this range of possible values, we only use 10 (11) of them:
#
# | Raster value | Aggregate class name |
# | :------------|:---------------------------|
# | 0 | No Data |
# | 1 | Broadleaf woodland |
# | 2 | Coniferous woodland |
# | 3 | Arable |
# | 4 | Improved grassland |
# | 5 | Semi-natural grassland |
# | 6 | Mountain, heath, bog |
# | 7 | Saltwater |
# | 8 | Freshwater |
# | 9 | Coastal |
# | 10 | Built-up areas and gardens |
#
# In this part of the practical, we'll try to work out the percentage of the entire country that is covered by each of these different landcovers, as well as each of the different counties. To start, we'll load the `LCM2015_Aggregate_100m.tif` raster, as well as the counties shapefile from Week 2:
# +
# open the land cover raster and read the data
with rio.open('data_files/LCM2015_Aggregate_100m.tif') as dataset:
xmin, ymin, xmax, ymax = dataset.bounds
crs = dataset.crs
landcover = dataset.read(1)
affine_tfm = dataset.transform
# now, load the county dataset from the week 2 folder
counties = gpd.read_file('../Week2/data_files/Counties.shp').to_crs(crs)
# -
# Next, we'll define a function that takes an array, and returns a __dict__ object containing the count (number of pixels) for each of the unique values in the array:
#
# ```python
# def count_unique(array, nodata=0):
# '''
# Count the unique elements of an array.
#
# :param array: Input array
# :param nodata: nodata value to ignore in the counting
#
# :returns count_dict: a dictionary of unique values and counts
# '''
# count_dict = {}
# for val in np.unique(array):
# if val == nodata:
# continue
# count_dict[str(val)] = np.count_nonzero(array == val)
# return count_dict
# ```
#
# Here, we have two input parameters: the first, `array`, is our array (or raster data). The next, `nodata`, is the value of the array that we should ignore. We then define an empty __dict__ (`count_dict = {}`).
#
# With [`numpy.unique()`](https://numpy.org/doc/stable/reference/generated/numpy.unique.html), we get an array containing the unique values of the input array. Note that this works for data like this raster, where we have a limited number of pre-defined values. For something like a digital elevation model, which represents continuous floating-point values, we wouldn't want to use this approach to bin the data.
#
# Next, for each of the different unique values `val`, we find all of the locations in `array` that have that value (`array == val`). Note that this is actually a boolean array, with values of either `True` where `array == val`, and `False` where `array != val`. [`numpy.count_nonzero()`](https://numpy.org/doc/stable/reference/generated/numpy.count_nonzero.html) the counts the number of non-zero (in this case, `True`) values in the array - that is, this:
#
# ```python
# np.count_nonzero(array == val)
# ```
#
# tells us the number of pixels in `array` that are equal to `val`. We then assign this to our dictionary with a key that is a __str__ representation of the value, before returning our `count_dict` variable at the end of the function.
#
# Run the cell below to define the function and run it on our `landcover` raster.
# +
def count_unique(array, nodata=0):
'''
Count the unique elements of an array.
:param array: Input array
:param nodata: nodata value to ignore in the counting
:returns count_dict: a dictionary of unique values and counts
'''
count_dict = {}
for val in np.unique(array):
if val == nodata:
continue
count_dict[str(val)] = np.count_nonzero(array == val)
return count_dict
unique_landcover = count_unique(landcover)
print(unique_landcover)
# -
# So this provides us with a __dict__ object with keys corresponding to each of the unique values (1-10).
#
# <span style="color:#009fdf;font-size:1.1em;font-weight:bold">Can you work out the percentage area of Northern Ireland that is covered by each of the 10 landcover classes?</span>
# In the following cell, we use [`rasterstats.zonal_stats()`](https://pythonhosted.org/rasterstats/manual.html#zonal-statistics) with our `counties` and `landcover` datasets to do the same exercise as above (counting unique pixel values). Rather than counting the pixels in the entire raster, however, we want to count the number of pixels with each land cover value that fall within a specific area defined by the features in the `counties` dataset:
# +
county_stats = zonal_stats(counties, landcover, affine=affine_tfm, categorical=True, nodata=0)
print(county_stats[0])
# -
# ## 3. The zip built-in
#
# This isn't a very readable result, though. If we want to interpret the results for each county, we have to know what land cover name corresponds to each of the values in the raster. One way that we could do this is by writing a function that re-names each of the keys in the __dict__. This example shows one way we could do this: the function takes the original __dict__ object (_dict_in_), as well as a list of the 'old' keys (_old_names_), and the corresponding 'new' keys (_new_names_).
def rename_dict(dict_in, old_names, new_names):
'''
Rename the keys of a dictionary, given a list of old and new keynames
:param dict_in: the dictionary to rename
:param old_names: a list of old keys
:param new_names: a list of new key names
:returns dict_out: a dictionary with the keys re-named
'''
dict_out = {}
for new, old in zip(new_names, old_names):
dict_out[new] = dict_in[old]
return dict_out
# For this function, we're also making use of the built-in function `zip()` ([documentation](https://docs.python.org/3.8/library/functions.html#zip)). In Python 3, `zip()` returns an __iterator__ object that combines elements from each of the iterable objects passed as arguments. As an example:
# +
x = [1, 2, 3, 4]
y = ['a', 'b', 'c', 'd']
list(zip(x, y))
# -
# So, with `zip(x, y)`, each of the elements of `x` is paired with the corresponding element from `y`. If `x` and `y` are different lengths, `zip(x, y)` will only use up to the shorter of the two:
# +
x = [1, 2, 3]
list(zip(x, y))
# -
# Let's see what happens when we run our function `rename_dict()` using the stats for our first county (County Tyrone - remember that the output from zonal_stats will have correspond to the rows of our input vector data):
# +
old_names = [float(i) for i in range(1, 11)]
new_names = ['Broadleaf woodland', 'Coniferous woodland', 'Arable', 'Improved grassland',
'Semi-natural grassland', 'Mountain, heath, bog', 'Saltwater', 'Freshwater',
'Coastal', 'Built-up areas and gardens']
rename_dict(county_stats[0], old_names, new_names)
# -
# Have a look at the _keys_ for `county_stats` - you should notice that there are no pixels corresponding to landcover class 7 (Saltwater), which makes sense given that County Tyrone is an inland county:
print(county_stats[0].keys())
print(county_stats[0])
# To run this for each of our counties, we could run some checks to make sure that we only try to access keys that exist in `dict_in`. For example, we could add an `if` statement to the function:
#
# ```python
# def rename_dict(dict_in, old_names, new_names):
# dict_out = {}
# for new, old in zip(new_names, old_names)
# if old in dict_in.keys():
# dict_out[new] = dict_in[old]
# else:
# continue
# return dict_out
# ```
#
# But, this is also an example of an exception that isn't necessarily something that requires us to stop executing our program. We don't expect each landcover type to be present in each county, so we don't want our program to stop as soon as it finds out that one of the counties doesn't have a particular landcover type.
#
# ## 4. Handling Exceptions with try ... except
# Python provides a way to handle these kind of exceptions: the [try...except](https://realpython.com/python-exceptions/#the-try-and-except-block-handling-exceptions) block:
# ```python
#
# try:
# # run some code
# except:
# # run this if the try block causes an exception
# ```
#
# In general, it's [not recommended](https://www.python.org/dev/peps/pep-0008/#programming-recommendations) to just have a bare `except:` clause, as this will make it harder to interrupt a program. In our specific case, we only want the interpreter to ignore `KeyError` exceptions - if there are other problems, we still need to know about those:
def rename_dict(dict_in, old_names, new_names):
'''
Rename the keys of a dictionary, given a list of old and new keynames
:param dict_in: the dictionary to rename
:param old_names: a list of old keys
:param new_names: a list of new key names
:returns dict_out: a dictionary with the keys re-named
'''
dict_out = {}
for new, old in zip(new_names, old_names):
try:
dict_out[new] = dict_in[old]
except KeyError:
continue
return dict_out
# Notice how for each pair of names, we try to assign the value corresponding to `old` in `dict_in`. If `old` is not a valid key for `dict_in`, we just move onto the next one. Now, let's run this new function on `county_stats[0]` again:
rename_dict(county_stats[0], old_names, new_names)
# We'll do one last thing before moving on here. Just like with the __dict__ outputs of `zonal_stats()`, the __list__ of __dict__ objects isn't very readable. Let's create a new __dict__ object that takes the county names as keys, and returns the re-named __dict__ objects for each:
# +
renamed_list = [rename_dict(d, old_names, new_names) for d in county_stats] # create a list of renamed dict objects
nice_names = [n.title() for n in counties.CountyName]
stats_dict = dict(zip(nice_names, renamed_list))
print(stats_dict['Tyrone'])
print(stats_dict['Antrim'])
# -
# Depending on how we're using the data, it might be easier to keep the output of `zonal_stats()` as-is, rather than using these long, complicated keys. For visualization and readability purposes, though, it helps to be able to easily and quickly understand what the outputs actually represent.
#
# <span style="color:#009fdf;font-size:1.1em;font-weight:bold">What is the total area (in km<sup>2</sup>) covered by "Mountain, heath, bog" in County Down?</span>
# ## 5. Rasterizing vector data using rasterio
# `rasterstats` provides a nice tool for quickly and easily extracting zonal statistics from a raster using vector data. Sometimes, though, we might want to _rasterize_ our vector data - for example, in order to mask our raster data, or to be able to select pixels. To do this, we can use the [`rasterio.features`](https://rasterio.readthedocs.io/en/latest/api/rasterio.features.html) module:
import rasterio.features # we have imported rasterio as rio, so this will be rio.features (and rasterio.features)
# `rasterio.features`has a number of different methods, but the one we are interested in here is `rasterize()`:
#
# ```
# rio.features.rasterize(
# shapes,
# out_shape=None,
# fill=0,
# out=None,
# transform=Affine(1.0, 0.0, 0.0,
# 0.0, 1.0, 0.0),
# all_touched=False,
# merge_alg=<MergeAlg.replace: 'REPLACE'>,
# default_value=1,
# dtype=None,
# )
# Docstring:
# Return an image array with input geometries burned in.
#
# Warnings will be raised for any invalid or empty geometries, and
# an exception will be raised if there are no valid shapes
# to rasterize.
#
# Parameters
# ----------
# shapes : iterable of (`geometry`, `value`) pairs or iterable over
# geometries. The `geometry` can either be an object that
# implements the geo interface or GeoJSON-like object. If no
# `value` is provided the `default_value` will be used. If `value`
# is `None` the `fill` value will be used.
# out_shape : tuple or list with 2 integers
# Shape of output numpy ndarray.
# fill : int or float, optional
# Used as fill value for all areas not covered by input
# geometries.
# ...
# ```
#
# Here, we pass an __iterable__ (__list__, __tuple__, __array__, etc.) that contains (__geometry__, __value__) pairs. __value__ determines the pixel values in the output raster that the __geometry__ overlaps. If we don't provide a __value__, it takes the `default_value` or the `fill` value.
#
# So, to create a rasterized version of our county outlines, we could do the following:
#
# ```python
# shapes = list(zip(counties['geometry'], counties['COUNTY_ID']))
#
# county_mask = rio.features.rasterize(shapes=shapes, fill=0,
# out_shape=landcover.shape, transform=affine_tfm)
# ```
#
# The first line uses `zip()` and `list()` to create a list of (__geometry__, __value__) pairs, and the second line actually creates the rasterized array, `county_mask`. Note that in the call to `rasterio.features.rasterize()`, we have to set the output shape (`out_shape`) of the raster, as well as the `transform` - that is, how we go from pixel coordinates in the array to real-world coordinates. Since we want to use this rasterized output with our `landcover`, we use the `shape` of the `landcover` raster, as well as its `transform` (`affine_tfm`) - that way, the outputs will line up as we expect. Run the cell below to see what the output looks like:
# +
shapes = list(zip(counties['geometry'], counties['COUNTY_ID']))
county_mask = rio.features.rasterize(shapes=shapes, fill=0,
out_shape=landcover.shape, transform=affine_tfm)
plt.figure()
plt.imshow(county_mask) # visualize the rasterized output
# -
# As you can see, this provides us with an __array__ whose values correspond to the `COUNTY_ID` of the county feature at that location (check the `counties` __GeoDataFrame__ again to see which county corresponds to which ID). In the next section, we'll see how we can use arrays like this to investigate our data further.
#
# ## 6. Masking and indexing rasters
# So far, we've seen how we can index an array (or a list, a tuple, ...) using simple indexing (e.g., `myList[0]`) or _slicing_ (e.g., `myList[2:4]`). `numpy` arrays, however, can [actually be indexed](https://numpy.org/doc/stable/reference/arrays.indexing.html) using other arrays of type `bool` (the elements of the array are boolean (`True`/`False`) values). In this section, we'll see how we can use this, along with our rasterized vectors, to select and investigate values from a raster using boolean indexing.
#
# To start, we'll open our dem raster - note that this raster has the same georeferencing information as our landcover raster, so we don't have to load all of that information, just the raster band:
with rio.open('data_files/NI_DEM.tif') as dataset:
dem = dataset.read(1)
# From the previous section, we have an array with values corresponding each of the counties of Northern Ireland. Using `numpy`, we can use this array to select elements of other rasters by creating a _mask_, or a boolean array - that is, an array with values of `True` and `False`. For example, we can create a mask corresponding to County Antrim (`COUNTY_ID=1`) like this:
#
# ```python
# county_antrim = county_mask == 1
# ```
# Let's see what this mask looks like:
# +
county_antrim = county_mask == 1
plt.figure()
plt.imshow(county_antrim)
# -
# We can also combine expressions using functions like [`np.logical_and()`](https://numpy.org/doc/stable/reference/generated/numpy.logical_and.html) or [`np.logical_or()`](https://numpy.org/doc/stable/reference/generated/numpy.logical_or.html). If we wanted to create a mask corresponding to both County Antrim and County Down, we could do the following:
# +
antrim_and_down = np.logical_or(county_mask == 3, county_mask == 1)
plt.figure()
plt.imshow(antrim_and_down)
# -
# We could then find the mean elevation of these two counties by indexing, or selecting, pixels from `dem` using our mask:
ad_elevation = dem[antrim_and_down]
print('Mean elevation: {:.2f} m'.format(ad_elevation.mean()))
# Now let's say we wanted to investigate the two types of woodland we have, broadleaf and conifer. One thing we might want to look at is the area-elevation distribution of each type. To do this, we first have to select the pixels from the DEM that correspond to the broadleaf woodlands, and all of the pixels corresponding to conifer woodlands:
broad_els = dem[landcover == 1] # get all dem values where landcover = 1
conif_els = dem[landcover == 2] # get all dem values where landcover = 2
# Now, we have two different arrays, `broad_els` and `conif_els`, each corresponding to the DEM pixel values of each landcover type. We can plot a histogram of these arrays using [`plt.hist()`](https://matplotlib.org/stable/api/_as_gen/matplotlib.pyplot.hist.html), but this will only tell us the number of pixels - for the area-elevation distribution, we have to convert the pixel counts into areas by multiplying with the pixel area (100 m x 100 m).
#
# First, though, we can use `numpy.histogram()`, along with an array representing our elevation bins, to produce a count of the number of pixels with an elevation that falls within each bin. Let's try elevations ranging from 0 to 600 meters, with a spacing of 5 meters:
# +
el_bins = np.arange(0, 600, 5) # create an array of values ranging from 0 to 600, spaced by 5.
broad_count, _ = np.histogram(broad_els, el_bins) # bin the broadleaf elevations using the elevation bins
conif_count, _ = np.histogram(conif_els, el_bins) # bin the conifer elevations using the elevation bins
broad_area = broad_count * 100 * 100 # convert the pixel counts to an area by multipling by the pixel size in x, y
conif_area = conif_count * 100 * 100
# -
# Finally, we can plot the area-elevation distribution for each land cover type using [`matplotlib.pyplot.bar()`](https://matplotlib.org/stable/api/_as_gen/matplotlib.pyplot.bar.html):
# +
fig, ax = plt.subplots(1, 1, figsize=(8, 8)) # create a new figure and axes object
# plot the area-elevation distributions using matplotlib.pyplot.bar(), converting from sq m to sq km:
_ = ax.bar(el_bins[:-1], broad_area / 1e6, align='edge', width=5, alpha=0.8, label='Broadleaf Woodland')
_ = ax.bar(el_bins[:-1], conif_area / 1e6, align='edge', width=5, alpha=0.8, label='Conifer Woodland')
ax.set_xlim(0, 550) # set the x limits of the plot
ax.set_ylim(0, 30) # set the y limits of the plot
ax.set_xlabel('Elevation (m)') # add an x label
ax.set_ylabel('Area (km$^2$)') # add a y label
ax.legend() # add a legend
# -
# From this, we can clearly see that Conifer woodlands tend to be found at much higher elevations than Broadleaf woodlands, and at a much larger range of elevations (0-500 m, compared to 0-250 m or so). With these samples (`broad_els`, `conif_els`), we can also calculate statistics for each of these samples using `numpy` functions such as `np.mean()`, `np.median()`, `np.std()`, and so on.
#
# <span style="color:#009fdf;font-size:1.1em;font-weight:bold">Of the 10 different landcover types shown here, which one has the highest mean elevation? What about the largest spread in elevation values?</span>
# ## Next steps
#
# That's all for this practical. In lieu of an an additional exercise this week, spend some time working on your project - are there concepts or examples from this practical that you can incorporate into your project?
#
# ### Footnotes
# [<sup id="fn1">1</sup>](#fn1-back)<NAME>.; <NAME>.; <NAME>.; <NAME>.; <NAME>.; <NAME>. (2017). Land Cover Map 2015 (25m raster, N. Ireland). NERC Environmental Information Data Centre. [doi:10.5285/47f053a0-e34f-4534-a843-76f0a0998a2f](https://doi.org/10.5285/47f053a0-e34f-4534-a843-76f0a0998a2f)</span>
| Week5/Practical5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Dive 11: Movie Reviews
#
# Today, we'll build a naive bayes classifier to judge whether a movie review is positive or negative. In today's exercise, we'll be using a sample of movie reviews [compiled by Stanford University's CS Department](http://ai.stanford.edu/~amaas/data/sentiment/).
#
# ***
#
# <img src="https://grantmlong.com/teaching/spring2019/lectures/img/l11_r7.png" width=600/>
#
# Courtesy of [@AmznMovieRevws](https://twitter.com/AmznMovieRevws)
# +
import os
# %matplotlib inline
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.naive_bayes import BernoulliNB
from sklearn.model_selection import KFold, train_test_split
from sklearn.metrics import accuracy_score, recall_score, precision_score
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import GradientBoostingClassifier
random_state = 20181118
# -
corpus_df = pd.read_csv('https://grantmlong.com/data/sentiment_corpus.csv')
corpus_df.shape
# #### Inspect df contents.
for i in np.random.choice(corpus_df.index.values, 4):
print(corpus_df.label.values[i])
print(corpus_df.text.values[i], '\n\n')
#
# # Process Input Data
# #### Split Train and Holdout Data Sets
# +
x_train, x_holdout, y_train, y_holdout = train_test_split(
corpus_df['text'],
corpus_df['label'], test_size=0.1,
random_state=random_state
)
train_df = pd.DataFrame(x_train, columns=['text'])
train_df['label'] = y_train
holdout_df = pd.DataFrame(x_holdout, columns=['text'])
holdout_df['label'] = y_holdout
train_df.reset_index(inplace=True)
holdout_df.reset_index(inplace=True)
print(train_df.shape[0], np.mean((train_df.label=='positive')*1))
print(holdout_df.shape[0], np.mean((holdout_df.label=='positive')*1))
# -
# ### Create Function to Vectorize Text
# The functionality `scikit-learn`'s [`CountVectorizer`](https://scikit-learn.org/stable/modules/generated/sklearn.feature_extraction.text.CountVectorizer.html) makes it really easy to transform words to text.
def featurize_text(df, n_features=1000, max_df=0.50, min_df=50):
vectorizer = CountVectorizer(max_df=max_df,
min_df=min_df,
max_features=n_features,
stop_words='english')
X = (vectorizer.fit_transform(df.text)>0)*1
Y = (df.label=='positive')*1
return X, Y, vectorizer
# ### Inspect Vectorized Data
X, Y, vectorizer = featurize_text(train_df, n_features=1000)
# +
word_df = pd.DataFrame(
data = X.todense(),
columns=vectorizer.get_feature_names()
)
count_df = pd.DataFrame(
data = [word_df.sum(),
word_df.loc[train_df.label=='positive'].sum(),
word_df.loc[train_df.label=='negative'].sum()],
index = ['total_count', 'pos_count', 'neg_count']
).transpose()
count_df['pos_share'] = count_df.pos_count/count_df.total_count*100
count_df['neg_share'] = count_df.neg_count/count_df.total_count*100
# -
(count_df[['total_count', 'pos_share', 'neg_share']]
.sort_values(by='total_count', ascending=True)[:30])
# # Cross Validate Vectorization Parameters
# Here we use `scikit-learn` to train a naive bayes classifier to rate the sentiment of each movie review. For more details of `scikit-learn` see [here](https://scikit-learn.org/stable/modules/naive_bayes.html).
# * What are the hyperparameters for this model?
# * Does it makes sense to train more than one?
# * What should we expect to see from our learning curves? Will more words overfit?
k_fold = KFold(n_splits=5, random_state=random_state)
def get_cv_results(df, n_features=1000, max_df=0.80, min_df=50):
nbayes = BernoulliNB()
X, Y, _ = featurize_text(df, n_features, max_df, min_df)
results = []
for train, test in k_fold.split(X):
nbayes.fit(X[train, :], Y[train])
y_predicted = nbayes.predict(X[test, :])
accuracy = accuracy_score(Y[test], y_predicted)
results.append(accuracy)
return np.mean(results), np.std(results)
get_cv_results(train_df)
# +
hp_values = [XXX]
all_mu = []
all_sigma = []
for m in hp_values:
mu, sigma = get_cv_results(train_df, n_features=m)
all_mu.append(mu)
all_sigma.append(sigma)
print(m, mu, sigma)
# -
plt.figure(figsize=(14, 5))
plt.plot(hp_values, all_mu)
plt.ylabel('Cross Validation Accuracy')
plt.xlabel('Number of Features')
# # Train Optimal Model
X, Y, vectorizer = featurize_text(train_df, n_features=XXX)
nbayes = BernoulliNB()
nbayes.fit(X, Y)
# #### Create Test Feature and Target Variables using the vectorizer we produced above.
# * Why do we use the vectorizer produced above instead of creating a new one?
# +
X_test = vectorizer.transform(holdout_df.text)
Y_test = (holdout_df.label=='positive')*1
# Generate predicted labels (positive=1)
y_predicted = nbayes.predict(X_test)
# Generate predicted probabilities
y_probpos = nbayes.predict_proba(X_test)[:, 1]
print(X_test.shape)
# -
# #### Inspect in-sample performance.
np.random.seed(random_state + 0)
for i in np.random.choice(holdout_df.index.values, 4):
print('Index: %i' % i)
print('Prob. Positive: %0.3f' % nbayes.predict_proba(X_test[i])[0][1])
print('Label: %s\n' % holdout_df.label.values[i])
print(holdout_df.text.values[i], '\n\n')
# #### Report Overall Out of Sample Performance
# * Based on these results, do you think our classifier is overfitting?
# +
def report_performance(classifier):
y_predicted = classifier.predict(X_test)
print('No. of test samples: %i' % len(y_predicted))
print('Accuracy: %0.1f%%' % (accuracy_score(Y_test, y_predicted)*100))
print('Precision: %0.1f%%' % (precision_score(Y_test, y_predicted)*100))
print('Recall: %0.1f%%' % (recall_score(Y_test, y_predicted)*100))
report_performance(nbayes)
# -
# ### Examine our false positives
np.random.seed(random_state + 0)
for i in np.random.choice(holdout_df.loc[(Y_test==0) & (y_predicted==1)].index.values, 2):
print('Index: %i' % i)
print('Prob. Positive: %0.3f' % y_probpos[i])
print('True Label: %s\n' % holdout_df.label.values[i])
print(holdout_df.text.values[i], '\n\n')
# ### Examine our false negatives
np.random.seed(random_state + 0)
for i in np.random.choice(holdout_df.loc[(Y_test==1) & (y_predicted==0)].index.values, 2):
print('Index: %i' % i)
print('Prob. Positive: %0.3f' % nbayes.predict_proba(X_test[i])[0][1])
print('True Label: %s\n' % holdout_df.label.values[i])
print(holdout_df.text.values[i], '\n\n')
# ### Examine borderline cases
# +
np.random.seed(random_state + 0)
border_threshold = 0.05
borderline = ((y_probpos>(0.5-border_threshold)) &
(y_probpos<(0.5+border_threshold)))
print('No. on border: %i' % sum(borderline), '\n')
for i in np.random.choice(holdout_df.loc[borderline].index.values, 2):
print('Index: %i' % i)
print('Prob. Positive: %0.3f' % nbayes.predict_proba(X_test[i])[0][1])
print('True Label: %s\n' % holdout_df.label.values[i])
print(holdout_df.text.values[i], '\n\n')
# -
# ### Benchmark Against Other Models
# * How will our performance with Naive Bayes live up to a logistic regression or gradient boosting machine?
# +
logreg = LogisticRegression(
random_state=random_state,
solver='lbfgs'
)
logreg.fit(X, Y)
report_performance(logreg)
# +
gbm = GradientBoostingClassifier(
random_state=random_state,
max_depth=20,
n_estimators=100
)
gbm.fit(X, Y)
report_performance(gbm)
# -
# # Take the classifier for a spin! Review a movie.
# * Does our naive bayes classifier perform like we'd expect it to?
# * How does it handle things like sarcasm? Negation?
#
# ***
#
# To evaluate our classifier, we'll also rely on some additional reviews helpfully compiled by [@AmznMovieRevws](https://twitter.com/AmznMovieRevws)
#
def score_review(example, model):
test = vectorizer.transform([example])
print('Words included: %i' % test.toarray().sum())
print('Prob. Positive: %0.3f' % nbayes.predict_proba(test)[0][1])
print(example)
# +
example = '''
This was a really great movie. One of the best I've seen in a while!
'''
score_review(example, nbayes)
# +
example = '''
I hated this move. The plot was so slow and the acting was terrible.
'''
score_review(example, nbayes)
# -
# 
#
# +
example = '''
False Advertising. Don't be fooled. Although this movie is called "The Rock" it has NOTHING to do with WWE sports
entertainer Dwayne "The Rock" Johnson.
'''
score_review(example, nbayes)
# -
# 
#
# +
example = '''
Misleading. There are no wolves in this movie.
'''
score_review(example, nbayes)
# -
# 
#
# +
example = '''
If you were going to see this movie because, like me, you love chocolate and and wanted to learn more about
the process prepare to go elsewhere. This movie has no chocolate to offer - no milk chocolate, no dark
chocolate, nothing about the abuses in the chocolate industry, and not even any hot chocolate on a cold evening.
I wish I were one of the dead characters in the movie, as I would have felt more than I did watching this.
'''
score_review(example, nbayes)
# -
# 
#
# +
example = '''
Expected a superhero named the tick to bite people and infect them with a series of horrible illnesses.
Not impressed.
'''
score_review(example, nbayes)
# +
example = '''
There is nothing funny about a tick being a super hero. Ticks cause lyme disease and ruin lives.
'''
score_review(example, nbayes)
# -
| lecture-11/DataDive-Lecture11-Reviews.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <small><small><i>
# All the IPython Notebooks in this **Python Examples** series by Dr. <NAME> are available @ **[GitHub](https://github.com/milaan9/90_Python_Examples)**
# </i></small></small>
# # Python Program to Iterate Over Dictionaries Using for Loop
#
# In this example, you will learn to iterate over dictionaries using **`for`** loop.
#
# To understand this example, you should have the knowledge of the following **[Python programming](https://github.com/milaan9/01_Python_Introduction/blob/main/000_Intro_to_Python.ipynb)** topics:
#
# * **[Python for Loop](https://github.com/milaan9/03_Python_Flow_Control/blob/main/005_Python_for_Loop.ipynb)**
# * **[Python Dictionary](https://github.com/milaan9/02_Python_Datatypes/blob/main/005_Python_Dictionary.ipynb)**
# +
# Example 1: Access both key and value using items()
dt = {'a': 'time', 'b': 'money', 'c': 'health'}
for key, value in dt.items():
print(key, value)
'''
>>Output/Runtime Test Cases:
a time
b money
c health
'''
# -
# **Explanation:**
#
# * Using a **`for`** loop, pass two loop variables **`key`** and **`value`** for iterable **`dt.items()`**. **`items()`** returns the **`key:value`** pairs.
# * Print **`key`** and **`value`**.
# +
# Example 2: Access both key and value without using items()
dt = {'a': 'time', 'b': 'money', 'c': 'health'}
for key in dt:
print(key, dt[key])
'''
>>Output/Runtime Test Cases:
a time
b money
c health
'''
# -
# **Explanation:**
#
# * Iterate through the dictionary using a **`for`** loop.
# * Print the loop variable **`key`** and **`value`** at key (i.e. **`dt[key]`**).
#
# However, the more pythonic way is example 1.
# +
# Example 3: Return keys or values explicitly
dt = {'a': 'time', 'b': 'money', 'c': 'health'}
for key in dt.keys():
print(key)
for value in dt.values():
print(value)
'''
>>Output/Runtime Test Cases:
a
b
c
time
money
health
'''
# -
# **Explanation:**
#
# You can use **`keys()`** and **`values()`** to explicitly return keys and values of the dictionary respectively.
| 03_Python_Flow_Control_examples/014_iterate_over_dictionaries_using_for_loop.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
# +
def fun(x):
return [x[0] + 0.5 * (x[0] - x[1])**3 - 1.0,
0.5 * (x[1] - x[0])**3 + x[1]]
def jac(x):
return np.array([[1 + 1.5 * (x[0] - x[1])**2, -1.5 * (x[0] - x[1])**2], [-1.5 * (x[1] - x[0])**2,
1 + 1.5 * (x[1] - x[0])**2]])
# -
from scipy import optimize
sol = optimize.root(fun, [0, 0], jac=jac, method='hybr')
sol.x
sol
sol.x
| HW3/test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lab 01: Tabular Data
#
# This lab is presented with some revisions from [<NAME> at Cal Poly](https://web.calpoly.edu/~dsun09/index.html) and his [Data301 Course](http://users.csc.calpoly.edu/~dsun09/data301/lectures.html)
#
# ### When you have filled out all the questions, submit via [Tulane Canvas](https://tulane.instructure.com/)
# What does data look like? For most people, the first image that comes to mind is a spreadsheet, with numbers neatly arranged in a table of rows and columns. One goal of this book is to get you to think beyond tables of numbers---to recognize that the words in a book and the markers on a map are also data to be collected, processed, and analyzed. But a lot of data is still organized into tables, so it is important to know how to work with **tabular data**.
# Let's look at a tabular data set. Shown below are the first 5 rows of a data set about the passengers on the Titanic. This data set contains information about each passenger (e.g., name, sex, age), their journey (e.g., the fare they paid, their destination), and their ultimate fate (e.g., whether they survived or not, the lifeboat they were on).
#
# <img src="../images/titanic_data.png" width="800">
# In a tabular data set, each row represents a distinct observation (or entity) and each column a distinct variable. Each **observation** is an entity being measured, and **variables** are the attributes we measure. In the Titanic data set above, each row represents a passenger on the Titanic. For each passenger, 14 variables have been recorded, including `pclass` (their ticket class: 1, 2, or 3) and `boat` (which lifeboat they were on, if they survived).
# ## Storing Data on Disk and in Memory
# How do we represent tabular data on disk so that it can be saved for later or shared with someone else? The Titanic data set above is saved in a file called `titanic.csv`. Let's peek inside this file using the shell command `head`.
#
# _Jupyter Tip_: To run a shell command inside a Jupyter notebook, simply prefix the shell command by the `!` character.
#
# _Jupyter Tip_: To run a cell, click on it and press the "play" button in the toolbar above. (Alternatively, you can press `Shift+Enter` on the keyboard.)
# !head ../data/titanic.csv
# The first line of this file contains the names of the variables, separated by commas. Each subsequent line contains the values of those variables for a passenger. The values appear in the same order as the variable names in the first line and are also separated by commas. Because the values in this file are separated (or _delimited_) by commas, this file is called a **comma-separated values** file, or **CSV** for short. CSV files typically have a `.csv` file extension, but not always.
#
# Although commas are by far the most common delimiter, you may encounter tabular data files that use tabs, semicolons (;), or pipes (|) as delimiters.
# How do we represent this information in memory so that it can be manipulated efficiently? In Python, the `pandas` library provides a convenient data structure for storing tabular data, called the `DataFrame`.
import pandas as pd
pd.DataFrame
# To read a file from disk into a `pandas` `DataFrame`, we can use the `read_csv` function in `pandas`. The first line of code below reads the Titanic dataset into a `DataFrame` called `df`. The second line calls the `.head()` method of `DataFrame`, which returns a new `DataFrame` consisting of just the first few rows (or "head") of the original.
df = pd.read_csv("../data/titanic.csv")
df.head()
# _Jupyter Tip_: When you execute a cell in a Jupyter notebook, the result of the last line is automatically printed. To suppress this output, you can do one of two things:
#
# - Assign the result to a variable, e.g., `df_head = df.head()`.
# - Add a semicolon to the end of the line, e.g., `df.head();`.
#
# I encourage you to try these out by modifying the code above and re-running the cell!
# Now that the tabular data is in memory as a `DataFrame`, we can manipulate it by writing Python code.
# ## Observations
#
# Recall that **observations** are the rows in a tabular data set. It is important to think about what each row represents, or the **unit of observation**, before starting a data analysis. In the Titanic `DataFrame`, the unit of observation is a passenger. This makes it easy to answer questions about passengers (e.g., "What percentage of passengers survived?") but harder to answer questions about families (e.g., "What percentage of families had at least one surviving member?")
# What if we instead had one row per _family_, instead of one row per _passenger_? We could still store information about _how many_ members of each family survived, but this representation would make it difficult to store information about _which_ members survived.
#
# There is no single "best" representation of the data. The right representation depends on the question you are trying to answer: if you are studying families on the Titanic, then you might want the unit of observation to be a family, but if you need to know which passengers survived, then you might prefer that it be a passenger. No matter which representation you choose, it is important to be conscious of the unit of observation.
# ### The Row Index
#
# In a `DataFrame`, each observation is identified by an index. You can determine the index of a `DataFrame` by looking for the **bolded** values at the beginning of each row when you print the `DataFrame`. For example, notice how the numbers **0**, **1**, **2**, **3**, **4**, ... above are bolded, which means that this `DataFrame` is indexed by integers starting from 0. This is the default index when you read in a data set from disk into `pandas`, unless you explicitly specify otherwise.
# Since each row represents one passenger, it might be useful to re-index the rows by the name of the passenger. To do this, we call the `.set_index()` method of `DataFrame`, passing in the name of the column we want to use as the index. Notice how `name` now appears at the very left, and the passengers' names are all bolded. This is how you know that `name` is the index of this `DataFrame`.
df.set_index("name").head()
# _Warning_: The `.set_index()` method does _not_ modify the original `DataFrame`. It returns a _new_ `DataFrame` with the specified index. To verify this, let's look at `df` again after running the above code.
df.head()
# Nothing has changed! If you want to save the `DataFrame` with the new index, you have to explicitly assign it to a variable.
df_by_name = df.set_index("name")
df_by_name.head()
# If you do not want the modified `DataFrame` to be stored in a new variable, you can either assign the result back to itself:
#
# `df = df.set_index("name")`
#
# or use the `inplace=True` argument, which will modify the `DataFrame` in place:
#
# `df.set_index("name", inplace=True)`.
#
# These two commands should only be run once. If you try to run them a second time, you will get an error. Don't just take my word for it---create a cell below and try it! The reason for the error is: after the command is executed the first time, `name` is no longer a column in `df`, since it is now in the index. When the command is run again, `pandas` will try (and fail) to find a column called `name`.
#
# Thus, the interactivity of Jupyter notebooks is both a blessing and a curse. It allows us to see the results of our code immediately, but it makes it easy to lose track of the state, especially if you run a cell twice or out of order. Remember that Jupyter notebooks are designed to be run from beginning to end. Keep this in mind as you run other people's notebooks and as you organize your own notebooks.
# ### Selecting Rows
#
# Now that we have set the (row) index of the `DataFrame` to be the passengers' names, we can use the index to select specific passengers. To do this, we use the `.loc` selector. The `.loc` selector takes in a label and returns the row(s) corresponding to that index label.
#
# For example, if we wanted to find the data for the father of the Allison family, we would pass in the label "Allison, Master. <NAME>" to `.loc`. Notice the square brackets.
df_by_name.loc["Allison, Master. <NAME>"]
# Notice that the data for a single row is printed differently. This is no accident. If we inspect the type of this data structure:
type(df_by_name.loc["Allison, Master. <NAME>"])
# we see that it is not a `DataFrame`, but a different data structure called a `Series`.
# `.loc` also accepts a _list_ of labels, in which case it returns multiple rows, one row for each label in the list. So, for example, if we wanted to select all 4 members of the Allison family from `df_by_name`, we would pass in a list with each of their names.
df_by_name.loc[[
"Allison, Master. <NAME>",
"Allison, Miss. <NAME>",
"Allison, Mr. <NAME>",
"Allison, Mrs. <NAME> (<NAME>)"
]]
# Notice that when there are multiple rows, the resulting data is stored in a `DataFrame`.
#
# The members of the Allison family happen to be consecutive rows of the `DataFrame`. If you want to select a consecutive set of rows, you do not need to type the index of every row that you want. Instead, you can use **slice notation**. The slice notation `a:b` allows you to select all rows from `a` to `b`. So another way we could have selected all four members of the Allison family is to write:
df_by_name.loc["Allison, Master. <NAME>":"Allison, Mrs. <NAME> (<NAME>)"]
# This behavior of the slice may be surprising to you if you are a Python veteran. We will say more about this in a second.
# What if you wanted to inspect the 100th row of the `DataFrame`, but didn't know the index label for that row? You can use `.iloc` to **select by position** (in contrast to `.loc`, which **selects by label**).
#
# Remember that `pandas` (and Python in general) uses zero-based indexing, so the position index of the 100th row is 99.
df_by_name.iloc[99]
# You can also select multiple rows by position, either by passing in a list:
df_by_name.iloc[[99, 100]]
# or by using slice notation:
df_by_name.iloc[99:101]
# Notice the difference between how slice notation works for `.loc` and `.iloc`.
#
# - `.loc[a:b]` returns the rows from `a` up to `b`, _including_ `b`.
# - `.iloc[a:b]` returns the rows from `a` up to `b`, _not including_ `b`.
#
# So to select the rows in positions 99 and 100, we do `.iloc[99:101]` because we want the rows from position 99 up to 101, _not including 101_. This is consistent with the behavior of slices elsewhere in Python. For example, the slice `1:2` applied to a list returns one element, not two.
test = ["a", "b", "c", "d"]
test[1:2]
# ### What Makes a Good Index?
#
# Something odd happens if we look for "Mr. <NAME>" in this `DataFrame`. Although we only ask for one label, we get two rows back.
df_by_name.loc["Kelly, Mr. James"]
# This happened because there were two passengers on the Titanic named "<NAME>". In general, a good row index should uniquely identify observations in the data set. Names are often, but not always, unique. The best row indexes are usually IDs that are guaranteed to be unique.
#
# Another common row index is time. If each row represents a measurement in time, then it makes sense to have the date or the timestamp be the index.
# ## Variables
# Recall that **variables** (or attributes) are the columns in a tabular data set. They are the measurements that we make on each observation.
# ### Selecting Variables
#
# Suppose we want to select the `age` column from the `DataFrame` above. There are three ways to do this.
# 1\. Use `.loc`, specifying both the rows and columns. (_Note:_ The colon `:` is Python shorthand for "all".)
df.loc[:, "age"]
# 2\. Access the column as you would a key in a `dict`.
df["age"]
# 3\. Access the column as an attribute of the `DataFrame`.
df.age
# Method 3 (attribute access) is the most concise. However, it does not work if the variable name contains spaces or special characters, begins with a number, or matches an existing attribute of `DataFrame`. For example, if `df` had a column called `head`, `df.head` would not return the column because `df.head` already means something else, as we have seen.
# Notice that the data structure used to store a single column is again a `Series`, not a `DataFrame`. So single rows and columns are stored in `Series`.
# To select multiple columns, you would pass in a _list_ of variable names, instead of a single variable name. For example, to select both the `age` and `sex` variables, we could do one of the following:
# +
# METHOD 1
df.loc[:, ["age", "sex"]].head()
# METHOD 2
df[["age", "sex"]].head()
# -
# Note that there is no way to generalize attribute access (Method 3 above) to select multiple columns.
# ### The Different Types of Variables
# There is a fundamental difference between variables like `age` and `fare`, which can be measured on a numeric scale, and variables like `sex` and `home.dest`, which cannot.
#
# Variables that can be measured on a numeric scale are called **quantitative variables**. Just because a variable happens to contain numbers does not necessarily make it "quantitative". For example, consider the variable `survived` in the Titanic data set. Each passenger either survived or didn't. This data set happens to use 1 for "survived" and 0 for "died", but these numbers do not reflect an underlying numeric scale.
#
# Variables that are not quantitative but take on a limited set of values are called **nominal or categorical variables**. For example, the variable `sex` takes on one of two possible values ("female" or "male"), so it is a categorical variable. So is the variable `home.dest`, which takes on a larger, but still limited, set of values. We call each possible value of a categorical variable a "category". Although categories are usually non-numeric (as in the case of `sex` and `home.dest`), they are sometimes numeric. For example, the variable `survived` in the Titanic data set is a categorical variable with two categories (1 if the passenger survived, 0 if they didn't), even though those values are numbers. With a categorical variable, one common analysis question is, "How many observations are there in each category?".
#
# Some variables do not fit neatly into either category. For example, the variable `name` in the Titanic data set is obviously not quantitative, but it is not categorical either because it does not take on a limited set of values. Generally speaking, every passenger will have a different name (the two <NAME> notwithstanding), so it does not make sense to analyze the frequencies of different names, as one might do with a categorical variable. We will group variables like `name`, that are neither quantitative nor categorical, into an "other" category.
#
# Every variable can be classified into one of these three **types**: quantitative, categorical, or other. The type of the variable often dictates the kind of analysis we do and the kind of visualizations we make, as we will see later in this chapter.
#
# `pandas` tries to infer the type of each variable automatically. If every value in a column (except for missing values) can be cast to a number, then `pandas` will treat that variable as quantitative. Otherwise, the variable is treated as categorical. To see the type that Pandas inferred for a variable, simply select that variable using the methods above and look for its `dtype`. A `dtype` of `float64` or `int64` indicates that the variable is quantitative. For example, the `age` variable above had a `dtype` of `float64`, so it is quantitative. On the other hand, if we look at the `sex` variable,
df.sex
# its `dtype` is `object`, so `pandas` will treat it as a categorical variable. Sometimes, this check can yield surprises. For example, if you only looked the first few rows of `df`, you might expect `ticket` to be a quantitative variable. But if we actually look at its `dtype`:
df.ticket
# it appears to be an `object`. That is because there are some values in this column that contain non-numeric characters. For example:
df.ticket[9]
# As long as there is one value in the column that cannot be cast to a numeric type, the entire column will be treated as categorical, and the individual values will be strings (notice the quotes around even a number like 24160, indicating that `pandas` is treating it as a string).
df.ticket[0]
# If you wanted `pandas` to treat this variable as quantitative, you can use the `to_numeric()` function. However, you have to specify what to do for values like `'PC 17609'` that cannot be converted to a number. The `errors="coerce"` option tells `pandas` to treat these values as missing (`NaN`).
pd.to_numeric(df.ticket, errors="coerce")
# If we wanted to keep this change, we would assign this column back to the original `DataFrame`, as follows:
#
# `df.ticket = pd.to_numeric(df.ticket, errors="coerce")`.
#
# But since `ticket` does not appear to be a quantitative variable, this is not actually a change we want to make.
# There are also categorical variables that `pandas` infers as quantitative because the values happen to be numbers. As we discussed earlier, the `survived` variable is categorical, but the values happen to be coded as 1 or 0. To force `pandas` to treat this as a categorical variable, you can cast the values to strings. Notice how the `dtype` changes:
df.survived.astype(str)
# In this case, this is a change that we actually want to keep, so we assign the modified column back to the `DataFrame`.
df.survived = df.survived.astype(str)
# ## Summary
#
# - Tabular data is stored in a data structure called a `DataFrame`.
# - Rows represent observations; columns represent variables.
# - Single rows and columns are stored in a data structure called a `Series`.
# - The row index should be a set of labels that uniquely identify observations.
# - To select rows by label, we use `.loc[]`. To select rows by (0-based) position, we use `.iloc[]`.
# - To select columns, we can use `.loc` notation (specifying both the rows and columns we want, separated by a comma), key access, or attribute access.
# - Variables can be quantitative, categorical, or other.
# - Pandas will try to infer the type, and you can check the type that Pandas inferred by looking at the `dtype`.
# # Exercises
# **Exercise 1.** Consider the variable `pclass` in the Titanic data set, which is 1, 2, or 3, depending on whether the passenger was in 1st, 2nd, or 3rd class.
#
# - What type of variable is this: quantitative, categorical, or other? (_Hint:_ One useful test is to ask yourself, "Does it make sense to add up values of this variable?" If the variable can be measured on a numeric scale, then it should make sense to add up values of that variable.)
# - Did `pandas` correctly infer the type of this variable? If not, convert this variable to the appropriate type.
# pclass would be categorical, because although it is numeric we cannot add these numbers. It is a limited set of values that we can use to categorize a person by class.
# since the value was numerical, pandas did not correctly infer the type.
df.pclass = df.pclass.astype(str)
# Exercises 2-7 deal with the Tips data set (`../data/tips.csv`). You can learn more about this data set on the first page of [this reference](http://www.ggobi.org/book/chap-data.pdf).
# **Exercise 2.** Read in the Tips data set into a `pandas` `DataFrame` called `tips`.
#
# - What is the unit of observation in this data set?
# - For each variable in the data set, identify it as quantitative, categorical, or other, based on your understanding of each variable. Did `pandas` correctly infer the type of each variable?
# - *Hint:* Check out the [Pandas dtype command](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.dtypes.html)
tips = pd.read_csv("../data/tips.csv")
tips.head()
# the unit of observation is a customer
# total_bill: quantitative; tip: quantitative; sex: categorical; smoker: categorical; day: categorical; time: categorical; size: quantitative
tips.dtypes
# pandas correctly inferred each variable type
# **Exercise 3.** Make the day of the week the index of the `DataFrame`.
#
# - What do you think will happen when you call `tips.loc["Thur"]`? Try it. What happens?
# - Is this a good variable to use as the index? Explain why or why not.
tips.set_index("day").head()
# at this point since I didn't actually modify the DataFrame's index, I think it will return an error
tips.loc["Thur"]
# it did return an error!
# this isn't a great variable to use as an index because there are lots of instances of each day; a proper index would have a unique value for each item
# **Exercise 4.** Make sure the index of the `DataFrame` is the default (i.e., 0, 1, 2, ...). If you changed it away from the default in the previous exercise, you can use `.reset_index()` to reset it.
#
# - How do you think `tips.loc[50]` and `tips.iloc[50]` will compare? Now try it. Was your prediction correct?
# - How do you think `tips.loc[50:55]` and `tips.iloc[50:55]` will compare? Now try it. Was your prediction correct?
# .loc selects by label, so it will try to return an index labeled 50. .iloc[50] will return the 49th row
tips.loc[50]
# tips.loc[50] was a little different than expected. I expected nothing but it returned the value at row 50
tips.iloc[50]
# this returned the exact same thing, I assume because the index of the original DataFrame corresponds to the actual
#index of each row. I did not think that far ahead so this is not what I predicted, but it makes sense.
# **Exercise 5.** How do you think `tips.loc[50]` and `tips.loc[[50]]` will compare? Now try it. Was your prediction correct? Are these the same or different? Why?
# tips.loc[50] will be the same as above and return the object labeled 50.
# tips.loc[[50]] looks like the set up for a list of labels, only this list just contains one value.
# I think tips.loc[[50]] will return the same thing as tips.loc[50]
tips.loc[[50]]
# I was sort of right; it returned the same information except for the dtype.
# This is because the first method displays as a series while the second method displays as a DataFrame.
# **Exercise 6.** What data structure is used to represent a single column, such as `tips["total_bill"]`? How could you modify this code to obtain a `DataFrame` consisting of just one column, `total_bill`?
# The data structure to represent a single column is a Series.
# I could modify it to get a DataFrame like this:
tips[["total_bill"]]
# **Exercise 7.** Create a new `DataFrame` from the Tips data that consists of just information about the table (i.e., whether or not there was a smoker, the day and time they visited the restaurant, and the size of the party), without information about the check or who paid.
#
# (There are many ways to do this. How many ways can you find?)
tips[["smoker", "day", "time", "size"]]
tips.loc[:, ["smoker", "day", "time", "size"]]
# ### When you have filled out all the questions, submit via [Tulane Canvas](https://tulane.instructure.com/)
| _labs/Lab01/Lab01-Tabular Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Refine the Data
import pandas as pd
df = pd.read_csv('tweets.csv', sep="\t")
df.head()
# To get the date of the title - we will need the following algorithm
# - If the string contains **hours** we can consider it **1 day**
# - And if the string has **day**, we pick the number preceding the **day**
#
# To apply this algorithm, we need to be able to pick these words and digits from a string. For that we will use Regular Expression.
# ## Introduction to Regular Expression (Regex)
#
# Regular expression is a way of selecting text using symbols in a string.
#
# Refer to the following links for an interactive playground
# - [http://regexr.com](http://regexr.com/)
# - [http://regex101.com/](http://regex101.com/)
import re
test_string = "Hello world, welcome to 2016."
# We can pass the whole string and re.search will give the first occurence of the value
# re.search - This function searches for first occurrence of RE pattern within string.
a = re.search('Hello world, welcome to 2016',test_string)
a
a.group()
# Match the first letters in the string
a = re.search('.',test_string)
a.group()
# Match all the letters in the string
a = re.search('.*',test_string)
a.group()
a = re.search('Hello',test_string)
print(a)
# ** Some basic symbols**
#
# **`?`**
#
# The question mark indicates zero or one occurrences of the preceding element. For example, colou?r matches both "color" and "colour".
#
# **`\*`**
#
# The asterisk indicates zero or more occurrences of the preceding element. For example, ab*c matches "ac", "abc", "abbc", "abbbc", and so on.
#
# **`\+`**
# The plus sign indicates one or more occurrences of the preceding element. For example, ab+c matches "abc", "abbc", "abbbc", and so on, but not "ac".
#
a = re.search('\w.',test_string)
print(a)
a = re.search('\w*',test_string)
print(a)
# ### Exercises
# Write a regex to remove URL link from the tweets
| notebook/twitter/Refine.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### <NAME> - BMI 203 - 3/24/17
import pickle as pkl
import matplotlib.pyplot as plt
# #### (1) Neural Network Implementation
#
# The auto-encoder problem is implemented in the function train_autoencoder with arguments (8,3,8) in the file neural_net.py. An example call is shown in the __main__.py file.
#
# This is generalized to DNA sequences by encoding each sequence with the following scheme. Each nucleotide in
# a sequence string is converted to a 4-bit number with three 0's and one 1 according to the following mapping:
#
# A: 1000
#
# G: 0100
#
# T: 0010
#
# C: 0001
#
# This takes a DNA sequence of length 17 and turns it into a vector of length 17*4 = 68. This 68-bit vector can then be
# used on a neural network with input layer of size 68. The DNA neural network is implemented using the same code as the autoencoder with a 68x20x1 architecture.
#
# #### (2/3) Learning Procedure and Training on Negative Data
#
# I created a 68x20x1 neural network to accept DNA input in the form described above. I then train this network on all the positive sequences in the file rap1-lieb-positives.txt and a subset of the negative sequences in the file yeast-upstream-1k-negative.fa. For every 1000bp negative sequence, I look at every 17-bp substring. If this 17-bp substring is one of the positive sequences, I do not include any 17-bp substring from this negative sequence in the negative training set. If the 1000bp negative sequence does not contain any 17-bp substring that is a positive sequence I include it in a master set of negative sequences. Once I have the master set of negative sequences, I take a random subset of the master set such that the final ratio of positive to negative training points is 10:1. This is to avoid over training on negative data since there are so many more negative data points.
# #### (4) Cross Validation and Changing Parameters
#
# Shown below are the results of three different testing methods I used for changing parameters of the neural network. I ran cross validation by aggregating all the training data in the manner described above then partitioning the master set of training data into 5 smaller subsets. I trained a neural network for each of these 5 subsets and then tested the trained network on the combined other 4 sets. In the case of cross validation, I was hoping to see very little fluctuation in the network response between the different training sets. As a rudimentary measure, I used the ratio of the number of guessed positives from the network to the number of actual positives in the validation set and the same ratio for negative binding sites. While I was happy to see little variance between networks, I realized that this was because the network converges to uniform output values for all inputs as described below in (5).
#
# I wrote functions to vary the hidden layer size and number of batch gradient descent iterations to test the corresponding network behavior. The same metrics are shown as for cross validation though the results once again reflect the network's convergence to uniform output values.
#
#
# Print results for cross-validation
results_0 = pkl.load(open('results_training_0.pkl', 'rb'))
results_1 = pkl.load(open('results_training_1.pkl', 'rb'))
results_2 = pkl.load(open('results_training_2.pkl', 'rb'))
results_3 = pkl.load(open('results_training_3.pkl', 'rb'))
results_4 = pkl.load(open('results_training_4.pkl', 'rb'))
results = [results_0, results_1, results_2, results_3, results_4]
for i in range(5):
num_positives, num_negatives, guess_positives, guess_negatives = results[i]
positive_ratio = guess_positives / num_positives
negative_ratio = guess_negatives / num_negatives
print('--------Training Set ' + str(i) + '----------------------------')
print(positive_ratio, negative_ratio)
print('--------------------------------------------------')
# Print results for varying hidden layer size
results_layer_size_1 = pkl.load(open('results_layer_size_1.pkl', 'rb'))
results_layer_size_5 = pkl.load(open('results_layer_size_5.pkl', 'rb'))
results_layer_size_10 = pkl.load(open('results_layer_size_10.pkl', 'rb'))
results_layer_size_20 = pkl.load(open('results_layer_size_20.pkl', 'rb'))
results_layer_size_50 = pkl.load(open('results_layer_size_50.pkl', 'rb'))
results_layer_size_100 = pkl.load(open('results_layer_size_100.pkl', 'rb'))
results = [results_layer_size_1, results_layer_size_5, results_layer_size_10, results_layer_size_20, results_layer_size_50, results_layer_size_100]
sizes = [1,5,10,20,50,100]
for i in range(6):
num_positives, num_negatives, guess_positives, guess_negatives = results[i]
# positive_ratio = guess_positives / num_positives
# negative_ratio = guess_negatives / num_negatives
print('--------Hidden Layer Size ' + str(sizes[i]) + '----------------------------')
print(positive_ratio, negative_ratio)
print('----------------------------------------------------')
# Print results for varying number of iterations
results_iterations_1 = pkl.load(open('results_iterations_1.pkl', 'rb'))
results_iterations_10 = pkl.load(open('results_iterations_10.pkl', 'rb'))
results_iterations_50 = pkl.load(open('results_iterations_50.pkl', 'rb'))
results_iterations_100 = pkl.load(open('results_iterations_100.pkl', 'rb'))
results_iterations_1000 = pkl.load(open('results_iterations_1000.pkl', 'rb'))
results = [results_iterations_1, results_iterations_10, results_iterations_50, results_iterations_100, results_iterations_1000]
iterations = [1,10,50,100,1000]
for i in range(5):
num_positives, num_negatives, guess_positives, guess_negatives = results[i]
# positive_ratio = guess_positives / num_positives
# negative_ratio = guess_negatives / num_negatives
print('--------Number Iterations ' + str(iterations[i]) + '----------------------------')
print(positive_ratio, negative_ratio)
print('-------------------------------------------------------')
# #### (5) Sequence Likelihood Predictions
#
# Sequence likelihood predictions can be found in the file likelihood_scores.tsv where each sequence is printed with its corresponding likelihood score seperated by a tab. Looking at the neural network output, it looks like all the scores have converged to the same value close to 0. This did not happen with my autoencoder on 8-bit numbers and I believe this is a product of the training scheme I used. By taking every substring of length 17 from the negative sequences, I may have included too much repetitive information from each negative sequence in the training data and in doing so may have washed out important features of the training set.
#
# The final trained neural network (weights, biases, and activations) can be found in the pickle file DNA_net.pkl.
| GabeReder_BMI203_FinalProject.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a name="top"></a>
# <div style="width:1000 px">
#
# <div style="float:right; width:98 px; height:98px;">
# <img src="https://raw.githubusercontent.com/Unidata/MetPy/master/src/metpy/plots/_static/unidata_150x150.png" alt="Unidata Logo" style="height: 98px;">
# </div>
#
# <h1>Pythonic Data Analysis</h1>
# <h3>Unidata Python Workshop</h3>
#
# <div style="clear:both"></div>
# </div>
#
# <hr style="height:2px;">
#
# <div style="float:right; width:250 px"><img src="http://matplotlib.org/_images/date_demo.png" alt="METAR" style="height: 300px;"></div>
#
# ### Questions
# 1. How can we employ Python language features to make complicated analysis require less code?
# 1. How can we make multi panel plots?
# 1. What can be done to eliminate repeated code that operates on sequences of objects?
# 1. How can functions be used to encapsulate calculations and behaviors?
#
# ### Objectives
# 1. <a href="#basicfunctionality">From the Time Series Plotting Episode</a>
# 1. <a href="#multipanel">Multi-panel plots</a>
# 1. <a href="#iteration">Iteration and Enumeration</a>
# 1. <a href="#functions">Functions</a>
# 1. <a href="#argskwargs">Args and Kwargs</a>
# 1. <a href="#plottingiteration">Plotting with Iteration</a>
# 1. <a href="#multifile">Plotting Multiple Files</a>
# <a name="basicfunctionality"></a>
# ## 1. From Time Series Plotting Episode
# Here's the basic set of imports and data reading functionality that we established in the [Basic Time Series Plotting](../Time_Series/Basic%20Time%20Series%20Plotting.ipynb) notebook.
# +
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib.dates import DateFormatter, DayLocator
from siphon.simplewebservice.ndbc import NDBC
# %matplotlib inline
# +
# Read in some data
df = NDBC.realtime_observations('42039')
# Trim to the last 7 days
df = df[df['time'] > (pd.Timestamp.utcnow() - pd.Timedelta(days=7))]
# -
# <a href="#top">Top</a>
# <hr style="height:2px;">
# <a name="multipanel"></a>
# ## 2. Multi-panel Plots
#
# Often we wish to create figures with multiple panels of data. It's common to separate variables of different types into these panels. We also don't want to create each panel as an individual figure and combine them in a tool like Illustrator - imagine having to do that for hundreds of plots!
#
# Previously we specified subplots individually with `plt.subplot()`. We can instead use the `subplots` method to specify a number of rows and columns of plots in our figure, which returns the figure and all of the axes (subplots) we ask for in a single call:
# +
# ShareX means that the axes will share range, ticking, etc. for the x axis
fig, (ax1, ax2) = plt.subplots(1, 2, sharex=True, figsize=(18, 6))
# Panel 1
ax1.plot(df.time.values, df.wind_speed, color='tab:orange', label='Windspeed')
ax1.set_xlabel('Time')
ax1.set_ylabel('Speed')
ax1.set_title('Measured Winds')
ax1.legend(loc='upper left')
ax1.grid(True)
# Not repeated only by sharing x
ax1.xaxis.set_major_formatter(DateFormatter('%m/%d'))
ax1.xaxis.set_major_locator(DayLocator())
# Panel 2
ax2.plot(df.time.values, df.pressure, color='black', label='Pressure')
ax2.set_xlabel('Time')
ax2.set_ylabel('hPa')
ax2.set_title('Atmospheric Pressure')
ax2.legend(loc='upper left')
ax2.grid(True)
plt.suptitle('Buoy 42039 Data', fontsize=24)
# -
# So even with the sharing of axis information, there's still a lot of repeated code. This current version with just two parameters might still be ok, but:
#
# - What if we had more data being plotted on each axes?
# - What if we had many subplots?
# - What if we wanted to change one of the parameters?
# - What if we wanted to plot data from different files on the same plot?
# <a href="#top">Top</a>
# <hr style="height:2px;">
# <a name="iteration"></a>
# ## 3. Iteration and Enumeration
# Iterating over lists is a very useful tool to reduce the amount of repeated code you write. We're going to start out by iterating over a single list with a `for` loop. Unlike C or other common scientific languages, Python 'knows' how to iterate over certain objects without you needing to specify an index variable and do the book keeping on that.
# +
my_list = ['2001 A Space Obyssey',
'The Princess Bride',
'Monty Python and the Holy Grail']
for item in my_list:
print(item)
# -
# Using the `zip` function we can even iterate over multiple lists at the same time with ease:
# +
my_other_list = ['I\'m sorry, Dave. I\'m afraid I can\'t do that.',
'My name is <NAME>ontoya.',
'It\'s only a flesh wound.']
for item in zip(my_list, my_other_list):
print(item)
# -
# That's really handy, but needing to access each part of each item with an index like `item[0]` isn't very flexible, requires us to remember the layout of the item, and isn't best practice. Instead we can use Python's unpacking syntax to make things nice and intuitive.
for reference, quote in zip(my_list, my_other_list):
print(reference, '-', quote)
# <div class="alert alert-success">
# <b>EXERCISE</b>:
# <ul>
# <li>Make two new lists named <code>plot_variables</code> and <code>plot_names</code>. Populate them
# with the variable name and plot label string for windspeed and pressure.</li>
# <li>Using the unpacking syntax, write a for loop that prints a sentence describing the action
# that would be taken (i.e. Plotting variable wind_speed as Windspeed</li>
# </ul>
# </div>
# +
# YOUR CODE GOES HERE
# -
# <div class="alert alert-info">
# <b>SOLUTION</b>
# </div>
# +
# # %load solutions/zip.py
# -
# `zip` can also be used to "unzip" items.
zipped_list = [(1, 2),
(3, 4),
(5, 6)]
unzipped = zip(*zipped_list)
print(list(unzipped))
# Let's break down what happened there. Zip pairs elements from all of the input arguements and hands those back to us. So effectively out `zip(*zipped_list)` is `zip((1, 2), (3, 4), (5, 6))`, so the first element from each input is paired (1, 3, 5), etc. You can think of it like unzipping or transposing.
#
# ---
# We can use the `enumerate` function to 'count through' an iterable object as well. This can be useful when placing figures in certain rows/columns or when a counter is needed.
for i, quote in enumerate(my_other_list):
print(i, ' - ', quote)
# <div class="alert alert-success">
# <b>EXERCISE</b>:
# <ul>
# <li>Combine what you've learned about enumeration and iteration to produce the following output:</li>
# </ul>
# <code>0 - 2001 A Space Obyssey - I'm sorry, Dave. I'm afraid I can't do that.
# 1 - The Princess Bride - My name is <NAME>.
# 2 - Monty Python and the Holy Grail - It's only a flesh wound.</code>
# </div>
# +
# YOUR CODE GOES HERE
# -
# <div class="alert alert-info">
# <b>SOLUTION</b>
# </div>
# +
# # %load solutions/enumerate.py
# -
# <a href="#top">Top</a>
# <hr style="height:2px;">
# <a name="functions"></a>
# ## 4. Functions
# You're probably already familiar with Python functions, but here's a quick refresher. Functions are used to house blocks of code that we can run repeatedly. Paramters are given as inputs, and values are returned from the function to where it was called. In the world of programming you can think of functions like paragraphs, they encapsulate a complete idea/process.
#
# Let's define a simple function that returns a value:
def silly_add(a, b):
return a + b
# We've re-implemented add which isn't incredibly exiciting, but that could be hundreds of lines of a numerical method, making a plot, or some other task. Using the function is simple:
result = silly_add(3, 4)
print(result)
# <div class="alert alert-success">
# <b>EXERCISE</b>:
# <ul>
# <li>Write a function that returns powers of 2. (i.e. calling <code>myfunc(4)</code> returns 2^4)</li>
# <li><b>Bonus</b>: Using for loop iteration, print all powers of 2 from 0 to 24.</li>
# </ul>
# </div>
# +
# Your code goes here
def myfunc(power):
return 2**power
for i in range(25):
print(myfunc(i))
# -
# <div class="alert alert-info">
# <b>SOLUTION</b>
# </div>
# +
# # %load solutions/functions.py
# -
# ### Reading buoy data with a function
# Let's create a function to read in buoy data and trim it down to the last 7 days by only providing the buoy number to the function.
def read_buoy_data(buoy, days=7):
# Read in some data
df = NDBC.realtime_observations(buoy)
# Trim to the last 7 days
df = df[df['time'] > (pd.Timestamp.utcnow() - pd.Timedelta(days=days))]
return df
df = read_buoy_data('42039')
df
# <a href="#top">Top</a>
# <hr style="height:2px;">
# <a name="argskwargs"></a>
# ## 5. Args and Kwargs
# Within a function call, we can also set optional arguments and keyword arguments (abbreviated args and kwargs in Python). Args are used to pass a *variable* length list of *non-keyword* arguments. This means that args don't have a specific keyword they are attached to, and are used in the order provided. Kwargs are arguments that are attached to specific keywords, and therefore have a specific use within a function.
# ### Args Example
# +
def arg_func(*argv):
for arg in argv:
print (arg)
arg_func('Welcome', 'to', 'the', 'Python', 'Workshop')
# -
# ### Kwargs Example
# Create a function to conduct all basic math operations, using a kwarg
def silly_function(a, b, operation=None):
if operation == 'add':
return a + b
elif operation == 'subtract':
return a - b
elif operation == 'multiply':
return a * b
elif operation == 'division':
return a / b
else:
raise ValueError('Incorrect value for "operation" provided.')
print(silly_function(3, 4, operation='add'))
print(silly_function(3, 4, operation='multiply'))
# Kwargs are commonly used in MetPy, matplotlib, pandas, and many other Python libraries (in fact we've used them in almost every notebook so far!).
# <a href="#top">Top</a>
# <hr style="height:2px;">
# <a name="plottingiteration"></a>
# ## 6. Plotting with Iteration
# Now let's bring what we've learned about iteration to bear on the problem of plotting. We'll start with a basic example and roll into a more involved system at the end.
#
# To begin, let's make an arbitrary number of plots in a single row:
# +
# A list of names of variables we want to plot
plot_variables = ['wind_speed', 'pressure']
# Make our figure, now choosing number of subplots based on length of variable name list
fig, axes = plt.subplots(1, len(plot_variables), sharex=True, figsize=(18, 6))
# Loop over the list of subplots and names together
for ax, var_name in zip(axes, plot_variables):
ax.plot(df.time.values, df[var_name])
# Set label/title based on variable name--no longer hard-coded
ax.set_ylabel(var_name)
ax.set_title(f'Buoy {var_name}')
# Set up our formatting--note lack of repetition
ax.grid(True)
ax.set_xlabel('Time')
ax.xaxis.set_major_formatter(DateFormatter('%m/%d'))
ax.xaxis.set_major_locator(DayLocator())
# -
# It's a step forward, but we've lost a lot of formatting information. The lines are both blue, the labels as less ideal, and the title just uses the variable name. We can use some of Python's features like dictionaries, functions, and string manipulation to help improve the versatility of the plotter.
#
# To start out, let's get the line color functionality back by using a Python dictionary to hold that information. Dictionaries can hold any data type and allow you to access that value with a key (hence the name key-value pair). We'll use the variable name for the key and the value will be the color of line to plot.
colors = {'wind_speed': 'tab:orange', 'wind_gust': 'tab:olive', 'pressure': 'black'}
# To access the value, just access that element of the dictionary with the key.
colors['pressure']
# Now let's apply that to our plot. We'll use the same code from the previous example, but now look up the line color in the dictionary.
# +
fig, axes = plt.subplots(1, len(plot_variables), sharex=True, figsize=(18, 6))
for ax, var_name in zip(axes, plot_variables):
# Grab the color from our dictionary and pass it to plot()
color = colors[var_name]
ax.plot(df.time.values, df[var_name], color)
ax.set_ylabel(var_name)
ax.set_title(f'Buoy {var_name}')
ax.grid(True)
ax.set_xlabel('Time')
ax.xaxis.set_major_formatter(DateFormatter('%m/%d'))
ax.xaxis.set_major_locator(DayLocator())
# -
# That's already much better. We need to be able to plot multiple variables on the wind speed/gust plot though. In this case, we'll allow a list of variables for each plot to be given and iterate over them. We'll store this in a list of lists. Each plot has its own list of variables!
# +
plot_variables = [['wind_speed', 'wind_gust'], ['pressure']]
fig, axes = plt.subplots(1, len(plot_variables), sharex=True, figsize=(18, 6))
for ax, var_names in zip(axes, plot_variables):
for var_name in var_names:
# Grab the color from our dictionary and pass it to plot()
color = colors[var_name]
ax.plot(df.time.values, df[var_name], color)
ax.set_ylabel(var_name)
ax.set_title(f'Buoy {var_name}')
ax.grid(True)
ax.set_xlabel('Time')
ax.xaxis.set_major_formatter(DateFormatter('%m/%d'))
ax.xaxis.set_major_locator(DayLocator())
# -
# <div class="alert alert-success">
# <b>EXERCISE</b>:
# <ul>
# <li>Create a dictionary of linestyles in which the variable name is the key and the linestyle is the value.</li>
# <li>Use that dictionary to modify the code below to plot the lines with the styles you specified.</li>
# </ul>
# </div>
# +
# Create your linestyles dictionary and modify the code below
fig, axes = plt.subplots(1, len(plot_variables), sharex=True, figsize=(18, 6))
for ax, var_names in zip(axes, plot_variables):
for var_name in var_names:
# Grab the color from our dictionary and pass it to plot()
color = colors[var_name]
ax.plot(df.time.values, df[var_name], color)
ax.set_ylabel(var_name)
ax.set_title(f'Buoy {var_name}')
ax.grid(True)
ax.set_xlabel('Time')
ax.xaxis.set_major_formatter(DateFormatter('%m/%d'))
ax.xaxis.set_major_locator(DayLocator())
# -
# <div class="alert alert-info">
# <b>SOLUTION</b>
# </div>
# +
# # %load solutions/looping1.py
# -
# We're almost back to where to started, but in a much more versatile form! We just need to make the labels and titles look nice. To do that, let's write a function that uses some string manipulation to clean up the variable names and give us an axis/plot title and legend label.
def format_varname(varname):
parts = varname.split('_')
title = parts[0].title()
label = varname.replace('_', ' ').title()
return title, label
# +
fig, axes = plt.subplots(1, len(plot_variables), sharex=True, figsize=(18, 6))
linestyles = {'wind_speed': '-', 'wind_gust': '--', 'pressure': '-'}
for ax, var_names in zip(axes, plot_variables):
for var_name in var_names:
title, label = format_varname(var_name)
color = colors[var_name]
linestyle = linestyles[var_name]
ax.plot(df.time.values, df[var_name], color, linestyle=linestyle, label=label)
ax.set_ylabel(title)
ax.set_title(f'Buoy {title}')
ax.grid(True)
ax.set_xlabel('Time')
ax.xaxis.set_major_formatter(DateFormatter('%m/%d'))
ax.xaxis.set_major_locator(DayLocator())
ax.legend(loc='upper left')
# -
# <a href="#top">Top</a>
# <hr style="height:2px;">
# <a name="multifile"></a>
# ## 7. Plotting Multiple Files
#
# Finally, let's plot data for two buoys on the same figure by iterating over a list of file names. We can use enumerate to plot each file on a new row of the figure. We will also create a function to read in the buoy data and avoid all of that repeated code.
# +
buoys = ['42039', '42022']
fig, axes = plt.subplots(len(buoys), len(plot_variables), sharex=True, figsize=(14, 10))
for row, buoy in enumerate(buoys):
df = read_buoy_data(buoy)
for col, var_names in enumerate(plot_variables):
ax = axes[row,col]
for var_name in var_names:
title, label = format_varname(var_name)
color = colors[var_name]
linestyle = linestyles[var_name]
ax.plot(df.time.values, df[var_name], color, linestyle=linestyle, label=label)
ax.set_ylabel(title)
ax.set_title(f'Buoy {buoy} {title}')
ax.grid(True)
ax.set_xlabel('Time')
ax.xaxis.set_major_formatter(DateFormatter('%m/%d'))
ax.xaxis.set_major_locator(DayLocator())
# -
# <a href="#top">Top</a>
# <hr style="height:2px;">
| pages/workshop/Pythonic_Data_Analysis/Pythonic Data Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
# %matplotlib inline
import matplotlib
matplotlib.rcParams["figure.figsize"]=(20,10)
imported_data= pd.read_csv('/Users/kushalsedhai/Downloads/Bengaluru_House_Data.csv')
imported_data
imported_data.groupby('area_type')['area_type'].agg('count')
imported_data_2= imported_data.drop(['availability','balcony','society','area_type'],axis=1)
imported_data_2
imported_data_2.isnull().sum()
imported_data_3=imported_data_2.dropna()
imported_data_3.isnull().sum()
imported_data_3['size'].unique()
imported_data_3['bhk']= imported_data_3['size'].apply(lambda x: int(x.split(' ')[0]))
imported_data_3.head()
imported_data_3['bhk'].unique()
imported_data_3[imported_data_3.bhk>20]
imported_data_3.total_sqft.unique()
def is_float(x):
try:
float(x)
except:
return False
return True
imported_data_3[~imported_data_3['total_sqft'].apply(is_float)]
def convert_sqft_to_num(x):
tokens = x.split('-')
if len(tokens)==2:
return(float(tokens[0])+float(tokens[1]))/2
try:
return float(x)
except:
return None
imported_data_4 = imported_data_3.copy()
imported_data_4['total_sqft']= imported_data_4['total_sqft'].apply(convert_sqft_to_num)
imported_data_4.head()
# Create a new price per sqft column
imported_data_5 = imported_data_4.copy()
imported_data_5['price_per_sqft']= imported_data_5['price']*100000/imported_data_5['total_sqft']
imported_data_5.head()
len(imported_data_5.location.unique())
imported_data_5.location = imported_data_5.location.apply(lambda x:x.strip())
location_stats = imported_data_5.groupby('location')['location'].agg('count').sort_values(ascending=False)
location_stats
len(location_stats[location_stats<=10])
location_stats_less_than_10 = location_stats [location_stats<=10]
location_stats_less_than_10
len(imported_data_5.location.unique())
imported_data_5.location = imported_data_5.location.apply(lambda x: 'other' if x in location_stats_less_than_10 else x)
imported_data_5.location
len(imported_data_5.location.unique())
imported_data_5[imported_data_5.total_sqft/imported_data_5.bhk<300].head()
imported_data_5.shape
imported_data_6= imported_data_5[~(imported_data_5.total_sqft/imported_data_5.bhk<300)]
imported_data_6.shape
imported_data_6.price_per_sqft.describe()
# +
def remove_pps_outliers(df):
df_out= pd.DataFrame()
for key, subdf in df.groupby('location'):
m = np.mean(subdf.price_per_sqft)
st = np.std(subdf. price_per_sqft)
reduced_df= subdf[(subdf.price_per_sqft>(m-st)) & (subdf.price_per_sqft<=(m+st))]
df_out= pd.concat([df_out , reduced_df], ignore_index=True)
return df_out
imported_data_7=remove_pps_outliers(imported_data_6)
imported_data_7.shape
# +
def plot_scatter_chart(df , location):
bhk2=df[(df.location==location) & (df.bhk==2)]
bhk3=df[(df.location==location) & (df.bhk==3)]
matplotlib.rcParams['figure.figsize'] = (15,10)
plt.scatter(bhk2.total_sqft , bhk2.price , color= "blue", label="2 BHK" , s=50)
plt.scatter(bhk3.total_sqft , bhk3.price , marker ='+', color= "green", label="3 BHK" , s=50)
plt.xlabel("Total Square Feet Area")
plt.ylabel("Price Per Square Feet")
plt.title(location)
plt.legend()
plot_scatter_chart(imported_data_7 , "<NAME>")
# +
def plot_scatter_chart(df , location):
bhk2=df[(df.location==location) & (df.bhk==2)]
bhk3=df[(df.location==location) & (df.bhk==3)]
matplotlib.rcParams['figure.figsize'] = (15,10)
plt.scatter(bhk2.total_sqft , bhk2.price , color= "blue", label="2 BHK" , s=50)
plt.scatter(bhk3.total_sqft , bhk3.price , marker ='+', color= "green", label="3 BHK" , s=50)
plt.xlabel("Total Square Feet Area")
plt.ylabel("Price Per Square Feet")
plt.title(location)
plt.legend()
plot_scatter_chart(imported_data_7 , "Hebbal")
# -
def remove_bhk_outliers(df):
exclude_indices =np.array([])
for location, location_df in df.groupby('location'):
bhk_stats = {}
for bhk, bhk_df in location_df.groupby('bhk'):
bhk_stats[bhk] = {
'mean': np.mean(bhk_df.price_per_sqft),
'std': np.std(bhk_df.price_per_sqft),
'count': bhk_df.shape[0]
}
for bhk, bhk_df in location_df.groupby('bhk'):
stats= bhk_stats.get(bhk-1)
if stats and stats['count']>5:
exclude_indices = np.append(exclude_indices, bhk_df[bhk_df.price_per_sqft <(stats['mean'])].index.values)
return df.drop(exclude_indices , axis= 'index')
imported_data_8= remove_bhk_outliers(imported_data_7)
imported_data_8.shape
plot_scatter_chart(imported_data_8 , "<NAME>")
plot_scatter_chart(imported_data_8 , "Hebbal")
import matplotlib
matplotlib.rcParams["figure.figsize"] = (20,10)
plt.hist(imported_data_8.price_per_sqft,rwidth=0.8)
plt.xlabel("Price Per Square Feet")
plt.ylabel("Count")
imported_data_8.bath.unique()
imported_data_8[imported_data_8.bath>10]
plt.hist(imported_data_8.bath , rwidth=0.8)
plt.xlabel("Number of Bathrooms")
plt.ylabel("Count")
imported_data_8[imported_data_8.bath>imported_data_8.bhk+2]
imported_data_9=imported_data_8[imported_data_8.bath<imported_data_8.bhk+2]
imported_data_9.shape
imported_data_10=imported_data_9.drop(['price_per_sqft', 'size' ] ,axis=1)
imported_data_10.head(3)
dummies = pd.get_dummies(imported_data_10.location)
dummies.head(3)
imported_data_11 = pd.concat([imported_data_10 , dummies.drop('other', axis=1)], axis=1)
imported_data_11.head(3)
imported_data_12=imported_data_11.drop('location', axis=1)
imported_data_12.head(3)
X = imported_data_12.drop('price', axis=1)
X.head()
y = imported_data_12.price
y.head()
from sklearn.model_selection import train_test_split
X_train , X_test , y_train , y_test = train_test_split (X,y,test_size=0.2,random_state=10)
from sklearn.linear_model import LinearRegression
lr_clf = LinearRegression()
lr_clf.fit(X_train, y_train)
lr_clf.score(X_test, y_test)
# +
from sklearn.model_selection import ShuffleSplit
from sklearn.model_selection import cross_val_score
cv = ShuffleSplit(n_splits=5 , test_size=0.2, random_state=0)
cross_val_score(LinearRegression(), X, y, cv=cv)
# +
from sklearn.model_selection import GridSearchCV
from sklearn.linear_model import Lasso
from sklearn.tree import DecisionTreeRegressor
def find_best_model_using_gridsearchcv(X,y):
algos = {
'linear_regression' : {
'model': LinearRegression(),
'params': {
'normalize': [True, False]
}
},
'lasso': {
'model': Lasso(),
'params': {
'alpha': [1,2],
'selection': ['random', 'cyclic']
}
},
'decision_tree': {
'model': DecisionTreeRegressor(),
'params': {
'criterion' : ['mse','friedman_mse'],
'splitter': ['best','random']
}
}
}
scores = []
cv = ShuffleSplit(n_splits=5, test_size=0.2, random_state=0)
for algo_name, config in algos.items():
gs = GridSearchCV(config['model'], config['params'], cv=cv, return_train_score=False)
gs.fit(X,y)
scores.append({
'model': algo_name,
'best_score': gs.best_score_,
'best_params': gs.best_params_
})
return pd.DataFrame(scores,columns=['model','best_score','best_params'])
find_best_model_using_gridsearchcv(X,y)
# -
def predict_price(location,sqft,bath,bhk):
loc_index = np.where(X.columns==location)[0][0]
x = np.zeros(len(X.columns))
x[0] = sqft
x[1] = bath
x[2] = bhk
if loc_index >= 0:
x[loc_index] = 1
return lr_clf.predict([x])[0]
predict_price('Indira Nagar', 1000 ,2 ,2)
predict_price('Rajaji Nagar', 1000 ,2 ,2)
#Export the model to a pickle file
import pickle
with open('bengalaru_home_price_modle_pickle','wb') as f:
pickle.dump(lr_clf,f)
X.columns
import json
columns = {
'data_columns' : [col.lower() for col in X.columns]
}
with open("columns.json","w") as f:
f.write(json.dumps(columns))
| apartment_price_model/Bengalaru apartment price prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import slate3k as slate
import pandas as pd
import re
import numpy as np
from tika import parser
# ##### ```tika``` uses a java pipeline and may require access to firewall, but it's the best parser I've found.
raw = parser.from_file('Films_Illustrating_Character_Strengths_and_Virtues.pdf')
# with open('Films_Illustrating_Character_Strengths_and_Virtues.pdf', 'rb') as f:
# extracted_text = slate.PDF(f)
# +
header1 = "Positive Psychology at the Movies"
header2 = "Appendices"
header3 = "From: <NAME>., & Wedding, D. : Using Films to Build Virtues and Character Strengths. © 2008 Hogrefe & Huber Publishers (http://www.hogrefe.com/)"
char_strengths = ['WISDOM: Creativity (originality, ingenuity)',
' WISDOM: Curiosity (interest, novelty-seeking, openness to experience)',
' WISDOM: Open-mindedness (judgment, critical thinking)',
' WISDOM: Love of Learning',
' WISDOM: Perspective (wisdom)',
' COURAGE: Bravery (valor)',
' COURAGE: Integrity (authenticity, honesty)',
' COURAGE: Vitality (zest, enthusiasm, vigor, energy)',
' COURAGE: Persistence (perseverance, industriousness)',
' HUMANITY: Love',
'HUMANITY: Kindness (generosity, nurturance, care, compassion, altruistic love, niceness)',
' HUMANITY: Social Intelligence (emotional, personal intelligence)',
' JUSTICE: Citizenship (social responsibility, loyalty, teamwork)',
' JUSTICE: Fairness',
'JUSTICE: Leadership ',
' TEMPERANCE: Forgiveness and Mercy',
' TEMPERANCE: Humility and Modesty',
' TEMPERANCE: Prudence',
' TEMPERANCE: Self-Regulation (self-control)',
' TRANSCENDENCE: Appreciation of Beauty and Excellence (awe, wonder, elevation)',
' TRANSCENDENCE: Grati-tude',
' TRANSCENDENCE: Hope (optimism, future mindedness, future orientation)',
' TRANSCENDENCE: Humor (playfulness)',
' TRANSCENDENCE: Spirituality (religiousness, faith, purpose, meaning)'
]
# -
def find(string, substring):
return [m.start() for m in re.finditer(substring, string)]
# +
content = raw['content']
content = content[content.find("WISDOM"):]
content = content[:content.find("View publication")]
content = content.replace("\n", "")
content = content.replace(header1, "")
content = content.replace(header2, "")
content = content.replace(header3, "")
content = content.strip()
# +
lines = []
all_indices = find(content, 'Ψ ')
for i, index in enumerate(all_indices):
if i == 0:
lines.append(content[0:index+1])
else:
lines.append(content[all_indices[i-1]+2:index+1])
# +
csv_dict = {}
count_score = []
char_found = -1
for line in lines:
previous_char = char_found
for strength in char_strengths:
# if any char strength is found in the line
if line.find(strength.strip()) != -1:
current_strength = strength.strip()
char_found += 1
if char_found != previous_char:
clear_line = line.replace(current_strength, "").replace('Ψ', "").strip()
# if there is a movie left
if clear_line != "":
csv_dict[clear_line] = current_strength.strip()
count_score.append([float(line.count('Ψ'))])
else:
# if the movie already has a char strength:
if line.replace('Ψ', "").strip() in list(csv_dict.keys()):
count_score[list(csv_dict.keys()).index(line.replace('Ψ', "").strip())].append(float(line.count('Ψ')))
csv_dict[line.replace('Ψ', "").strip()] += ';' + current_strength.strip()
else:
count_score.append([float(line.count('Ψ'))])
csv_dict[line.replace('Ψ', "").strip()] = current_strength.strip()
# +
dataframe_dict = {}
df_columns = [i.strip() for i in char_strengths]
for i, key in enumerate(csv_dict.keys()):
binary_list = []
binary_list.append(np.array(count_score[i]).mean())
for strength in df_columns:
if strength in csv_dict[key].split(';'):
binary_list.append('1')
else:
binary_list.append('0')
dataframe_dict[key.replace('Ψ', "").strip()] = binary_list
# -
movie_df = pd.DataFrame.from_dict(dataframe_dict, orient='index', columns=df_columns.insert(0, 'Meaningfulness'))
# +
renames = ['Meaningfulness',
'Creativity (Wisdom)',
'Curiosity (Wisdom)',
'Open-mindedness (Wisdom)',
'Love of learning (Wisdom)',
'Perspective (Wisdom)',
'Bravery (Courage)',
'Integrity (Courage)',
'Vitality (Courage)',
'Persistence (Courage)',
'Love (Humanity)',
'Kindness (Humanity)',
'Social Intelligence (Humanity)',
'Citizenship (Justice)',
'Fairness (Justice)',
'Leadership (Justice)',
'Forgiveness and mercy (Temperance)',
'Humility and modesty (Temperance)',
'Prudence (Temperance)',
'Self-regulation (Temperance)',
'Appreciation of Beauty and Excellence (Transcendence)',
'Gratitude (Transcendence)',
'Hope (Transcendence)',
'Humor (Transcendence)',
'Spirituality (Transcendence)'
]
renaming_dict = {}
for i, col in enumerate(list(movie_df.columns)):
renaming_dict[col] = renames[i]
movie_df.rename(columns = renaming_dict, inplace = True)
movie_df['movie_id'] = range(0, len(movie_df))
movie_df.to_csv('paper_dataset.csv')
| database/csv_from_pdf.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] cell_id="00000-1880b0fe-0687-4cdf-a63b-26c7845a0227" tags=[]
# # Introduction
#
#
# Taking derivatives is an ubiquitous tool that covers a wide range of applications - from mathematics, to physics and computer science, etc.
#
# In machine learning, differentiation is extensively applied to the optimization problem by performing the gradient descent to minimize the cost function.
# Using a fast and versatile differentiation method, people can also get the second order derivatives and hessian matrices.
#
# Differentiation is also applied to solve everyday problems. For example, knowing the initial decomposition level of a contaminated food,
# we can define a differential equation to predict how food will decay in the future and set an appropriate expiration date.
#
# However, traditional methods of differentiation such as symbolic differentiation and numerical differentiation might be computationally expensive and less robust. Also, these methods do not scale well to vector functions with multiple variable inputs, which are widely used to solve real world problems.
#
# In this package, SmartDiff is a software package which implements a forward mode Automatic Differentiation (AD) package with chain rule to help users take derivatives easily. It automatically breaks the complex functions into small independent elementary operations based parentheses and operators. Then it calculates the derivatives in the small chunks step by step, moving outward to the entire function, to give the result. Therefore, SmartDiff is widely **generalizable** to different functions and **consumes very little memory**. It also computes the derivative of different components of the function in parallel, further speeding up the computation.
#
# + [markdown] cell_id="00001-e4cec731-17d4-4c6a-8917-08832f84a123" tags=[]
# # Background
#
# ## Chain Rule
# This automatic differentiation (AD) is performed mainly based on the chain rule.
# The basic idea is to split the function into hierarchies of chunks containing the input variables and the built-in elementary operations in SmartDiff.
# Therefore, from the innermost chunk(s), the calculation can be expressed in a series of steps iteratively, in which SmartDiff only considers one elementary operation at a time.
# After the evaluation of both the function and its derivatives, the iteration can be traced in the computation graph.
#
# Based on the key definition of chain rule, and known that $F(x) = f(g(x))$, we can represent it in multiple steps as follows:
# $\frac{\partial}{\partial x}(F(x)) =\frac{\partial}{\partial x}f(g(x))=f'(g(x))g'(x)$
#
# From the equation above, as long as we can compute the derivative of $f(x)$ and $g(x)$, we can get the derivative of $F(x)$.
#
# Also, with such forward method, SmartDiff is able to evaluate each step of the calculation and keep track of function values and derivatives at each step, which makes the calculation faster and more robust.
#
# ## Graph Structure of Calculation
# SmartDiff is implemented based on a graph structure. For each node on the graph, it contains a variable or the result following one elementary operation.
# For example, given a multivariable function $f(x,y) = (x^2+1)+\exp{(y)}$. To evaluate the function at $(x,y) = (1,1)$, we can write the evaluation trace as:
#
# | `Trace` | ` Operation` | `Value` | ` Derivative ` | `Der to x `| `Der to y` |
# |: -----------------:|:------------------:|:----------------------:|:-----------------------:|:-----------:|:--------------------------------:|
# | $x_1$ | $x$ | 1 | $ \dot{x}$ | 1 | 0 |
# | $x_2$ | $y$ | 1 | $ \dot{y}$ | 0 | 1 |
# | $v_1$ | $x_1^2$ | 1 | $2x_1\dot{x_1}$ | 2 | 0 |
# | $v_2$ | $v_1+1$ | 2 | $\dot{v_1}$ | 2 | 0 |
# | $v_3$ | $\exp{(x_2)}$ | $e$ | $\exp{(x_2)}\dot{x_2}$ | 0 | $e$ |
# | $f$ | $v_2+v_3$ | $e+2$ | $\dot{v_2}+\dot{v_3}$ | 2 | $e$ |
#
# The traces above split the function into chunks of elementary operations and can be easily put into a graph structure as following, which is helpful for the differentiation calculation.
#
# 
#
# ## Elementary Functions
# SmartDiff is mainly a combination of elementary operations in calculus, which contains:
#
# 1. Arithmetic symbols (+, -, ×, /), parentheses, constant functions ($2$, $\pi$, $e$, etc.)
# 2. Logarithmic functions, exponential functions, trigonometric functions, square root function, hyperbolic functions, etc.
# + [markdown] cell_id="00002-2ac024b1-909e-4391-aab6-b46bf4a48214" tags=[]
# # How to use SmartDiff
#
# 1. How do you envision that a user will interact with your package?
#
# The user first installs SmartDiff and calls the main script in the commandline and interacts with the Graphic User Interface (GUI). There is no need to import or instantiate python objects.
#
# ```python
# pip install SmartDiff
# python /dir/to/SmartDiff/main.py
# ```
#
# + [markdown] cell_id="00003-11b22b12-f2b2-4145-8659-afb3a171594e" tags=[]
# # SmartDiff Graphic User Interface (GUI)
#
# ### Procedure:
#
# Once SmartDiff is installed, it outputs the derivative(s) in the following steps.
#
# 1. The user chooses the dimension of the vector function from a drop down list (1,2,3) and the number of one-dimensional real-valued variables from a drop down list (1,2,3)
#
# 2. The user inputs a specific data point (required)
#
# 3. The user inputs a vector function by each component of the vector function
#
# 4. The user checks the confirmation message, showing the function and the variable values.
#
# 5. The user can choose to see the function value and the trace table in addition to the differentiation result.
#
# 6. The user clicks Ok to differentiate.
#
# 7. The program shows the results based on user specifications
#
# ### Advantages:
#
# Using the buttons of functions, operators, and digits, the GUI can directly translate the user input into a defined string for latex rendering and a python expression for function evaluation and differentiation. This also alleviates the need to check for typos and undefined functions within the user input.
#
# + [markdown] cell_id="00004-53ac6421-7441-4699-b81e-3cc972a98951" tags=[]
# # Software Organization
#
# ### 1. Directory structure:
# The following is the structure of the project after users download it from our github repository.
# The package we are developing is SmartDiff and the few files outside of it would be used in its packaging and distribution.
# ```python
# cs107-FinalProject/
# setup.py
# requirement.txt
# README.md
# LICENSE
# docs/
# milestone1.ipynb
# graph.png
#
# SmartDiff/
# __init__.py
# main.py # start up script
# preprocess/ # This is a (sub)package
# __init__.py
# parse_gui_input.py # A GUI input parser module
# solvers/ # calculate derivatives and function values at the given point
# __init__.py
# element_op.py # define the derivatives of elementary operations
# integrator.py # go through the preprocessed python expression and call the corresponding elementary operations
# find_value.py # calculate the function value based on preprocessed output
# trace_table/ # generate the trace table during forward AD
# __init__.py
# gen_table.py # generate the table based on integrator.py output
# visualize_table.py # generate a .png file of the trace table
# GUI/ # create a PyQt5-based GUI that takes in user input and displays computation output
# __init__.py
# UI_main.py # coordinate between different UI submodules
#
# layout/ # configure the window layout for each step
# __init__.py
# step1.py
# step2.py
# step3.py
# step4.py
# step5.py
# step6.py
# step7.py
# all.py # the window configuration shared by all steps
# shared.py # the window configuration shared by step 2 and 3
#
# config_layout/ # Changes the window based on user input in the previous step
# __init__.py
# confit.py
#
# msg/ # generate instructions and error message (sometimes based on other modules) for the user (all steps)
# __init__.py
# error_msg.py
# instruction.py
#
# tests/ # extensive tests of each module and boundaries between different modules
# __init__.py
# preprocess_test.py
# solvers_test.py
# UI_test.py
# UI_preprocess_test.py
# preprocess_solvers_test.py
# solvers_UI.py
# ```
#
# ### 2. Testing Procedure
# For each function, we will have some basic test cases in the docstring. For extensive testing between the boundaries of different modules, we will implement several test functions for each module
# on separate files under /tests. These tests will include the functions *element_op_tests*, *integrator_tests*, *find_value_tests*, and *trace_table_tests*, which will check the correct definition of
# derivatives for elementary operations, the successful implementation of those derivatives in a given equation, and the correct generation of a trace table, respectively. All of the tests functions will
# be called under a main test calling function, *all_tests*, to ensure that all the tests are running. The automatic testing of our code will be conducted using TravisCI and the proportion of code
# tested will be monitored by CodeCov.
#
# ### 3. Packaging and Package Distribution
# We will be using Python Package Index (PyPI) to pack up our project and distribute it to the Python user community.
# The PyPI of our package is "smartdiff", the same name to be used in pip install.
# setup.py file is a build script of setuptools and will be used to generate distribution archive files. Then we generate our project API token and upload the files to TestPyPI.
#
# ### 4. Other Considerations
# Since users may have different local environments from the one developments are based on, we are researching into methods that can help users install the dependencies of smartdiff.
# We found two ways to handle dependency requirements:
#
# 1. install_require() function from the setuptools package.
#
# 2. specify our own requirement file.
#
# Basic idea of SmartDiff:
#
# From the user input in GUI, we can simultaneously
#
# 1. render a latex expression of the function
#
# 2. get a python expression
#
# For each component of the vector function, we divide it into chunks by parentheses (priority) and operators, then start from the innermost chunk(s) and evaluate the derivatives outward.
# + [markdown] cell_id="00005-f740c177-ed1f-4ef7-bfe5-0abbe1cf97e0" tags=[]
# # Implementation
#
# ### 1. Core Data Structures
# One of the core data structures is stack.
# When users type in the equations from left to right, they may create a few nested expressions. So the input parser needs to keep track of the terms at each level. Whenever we encounter an open parenthesis, we need to start a new level on top of the stack and add expression at that nested level as operand tuples.
# Whenever a level terminates (i.e. the closing parenthesis is encountered), we need to pop the expression off from the top of stack and evaluate its derivative with respect to the target variable.
# If a terminated level of expression does not contain the target variable, we can simply leave it as a generic operand as a whole and continue the parsing.
#
# ### 2. Classes, Methods and Name Attributes
# For now, we have identified 5 basic classes as follows. However, we may add or delete some of them depending on how the fundamental algorithm works.
# - Operand() class
# - type: "numeral" or "var" since the type of each operand can either be a variable or a numerical value. If it is a mixed type, as long as there is a variable, its type will be "var".
# - value: string representation of the variable or numerical value
# - has_target_var: True if this operand contains the target variable (since this operand will be treated as a variable in the outer expression levels); False otherwise.
#
# - Operator() class
# This is an Enum class that holds the string ID of each mathematical operators the differentiator can handle. Its attributes are the uppercase name of each operator.
#
# - ElementaryGrad() class
# This class has a method for each type of operator above. Each method returns an new Operand object which is the derivative with respect to one input operand.
# The base case of evaluating the derivative is 1. taking derivative of a single operand 2. taking derivative of two operands connected with an operator with respect to one operand.
#
# - InputScanner() class
# This class provides two methods described below:
# - tokenize(input_str, dx_str): scans the input string and outputs a nested list of operand-operator tuples and an operand for "dx_str".
# - to_latex_format(input_str, dx_str): converts the input string into a latex string to be displayed in the GUI.
#
# - Differentiator() class
# This class takes evaluates the derivative of a tokenized expression.
# - grad(expr, dx): It processes a nested list of tuples and iteratively computes the derivative with respect to the operand "dx".
#
# - GUI() class
# This class will be based on the python GUI package, PyQt5 and uses the following predefined python classes to
# inform the user with instructions and error messages, direct the user to input the information, and display the output:
# - QComboBox, QLineEdit, QPushButton, QLabel, QCheckBox, QMessageBox
#
#
# ### 3. External Dependencies
#
# - PyQt5, for the UI
# - NumPy, for the solvers
# - Pandas, for generating the trace table
# - Pydoc, for viewing the function docstring
# - doctest, for unit testing of all the cases a function deals with (in a function docstring)
# - pytest, for testing files in test (extensive testing of boundaries between different modules)
# - setuptools & wheel, for packaging and distributing the code
# - twine, for uploading the distribution archive files
#
# ### 4. Handle Elementary Functions like sin, sqrt, log, and exp (and all the others)
#
# We will be using operator overloading to define the derivative of each elementary functions in SmartDiff/solvers. For example, $\sin(x)$ will be mapped to $\cos(x)$ and $x+y$ will still be $x+y$.
# + [markdown] cell_id="00006-7f2b52a1-95f7-4513-864b-96787ffc3942" tags=[]
# # Feedback
#
# ## Milestone1
#
# #### Feedback from <NAME>:
# On background part: having computational graph example will be nice. - Rest is pretty good
#
#
# #### Our updates:
# We created the computational graph structure to better describute such calculation in Background/Gaph Structure of Calculation.
# We also updated the directory structure to insert the image we use.
#
| docs/milestone1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sh
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Bash
# language: bash
# name: bash
# ---
# # Identifying contamination
# It is always a good idea to check that your data is from the species you expect it to be. A very useful tool for this is [Kraken](https://www.ebi.ac.uk/research/enright/software/kraken). In this tutorial we will go through how you can use Kraken to check your samples for contamination.
#
# __Note if using the Sanger cluster:__ Kraken is run as part of the automatic qc pipeline and you can retreive the results using the `pf qc` script. For more information, run `pf man qc`.
#
# Before we start, change into the `QC/qc-training-notebooks/` directory:
cd ~/QC/qc-training-notebooks
# ## Setting up a database
# To run Kraken you need to either build a database or download an existing one. The standard database is very large (33 GB), but thankfully there are some smaller, pre-built databased available. To download the smallest of them, the 4 GB MiniKraken, run this command (this may take a few minutes to complete):
wget http://ccb.jhu.edu/software/kraken/dl/minikraken_20171019_4GB.tgz
# Then all you need to do is un-tar it:
tar -zxvf minikraken_20171019_4GB.tgz
# This version of the database is constructed from complete bacterial, archaeal, and viral genomes in RefSeq, however it contains only around 3 percent of the kmers from the original kraken database (more information [here](https://ccb.jhu.edu/software/kraken/)). If the pre-packaged databases are not quite what you are looking for, you can create your own customized database instead. Details about this can be found [here](http://ccb.jhu.edu/software/kraken/MANUAL.html#custom-databases).
#
# __Note if using the Sanger cluster:__ There are several pre-built databases available centrally on the Sanger cluster. For more information, please contact the Pathogen Informatics team.
# ## Running Kraken
# To run Kraken, you need to provide the path to the database you just created. By default, the input files are assumed to be in FASTA format, so in this case we also need to tell Kraken that our input files are in FASTQ format, gzipped, and that they are paired end reads (this may take a while to complete):
kraken --db ./minikraken_20171013_4GB --output kraken_results \
--fastq-input --gzip-compressed --paired \
'data/13681_1#18_1.fastq.gz' 'data/13681_1#18_2.fastq.gz'
# The five columns in the file that's generated are:
#
# 1. "C"/"U": one letter code indicating that the sequence was either classified or unclassified.
# 2. The sequence ID, obtained from the FASTA/FASTQ header.
# 3. The taxonomy ID Kraken used to label the sequence; this is 0 if the sequence is unclassified.
# 4. The length of the sequence in bp.
# 5. A space-delimited list indicating the LCA mapping of each k-mer in the sequence.
#
# To get a better overview you can create a kraken report:
kraken-report --db ./minikraken_20171013_4GB \
kraken_results > kraken-report
# ## Looking at the results
# Let's have a closer look at the kraken_report for the sample. If for some reason your kraken-run failed there is a prebaked kraken-report at data/kraken-report
head -n 20 kraken-report
# The six columns in this file are:
#
# 1. Percentage of reads covered by the clade rooted at this taxon
# 2. Number of reads covered by the clade rooted at this taxon
# 3. Number of reads assigned directly to this taxon
# 4. A rank code, indicating (U)nclassified, (D)omain, (K)ingdom, (P)hylum, (C)lass, (O)rder, (F)amily, (G)enus, or (S)pecies. All other ranks are simply '-'.
# 5. NCBI taxonomy ID
# 6. Scientific name
#
# ## Exercises
# __Q1: What is the most prevalent species in this sample?__
# __Q2: Are there clear signs of contamination?__
# __Q3: What percentage of reads could not be classified?__
# ## Heterozygous SNPs
# For bacteria, another thing that you can look at to detect contamination is if there are heterozygous SNPs in your samples. Simply put, if you align your reads to a reference, you would expect any snps to be homozygous, i.e. if one read differs from the reference genome, then the rest of the reads that map to that same location will also do so:
#
# __Homozygous SNP__
# Ref: CTTGAGACGAAATCACTAAAAAACGTGACGACTTG
# Read1: CTTGAGtCG
# Read2: CTTGAGtCGAAA
# Read3: GAGtCGAAATCACTAAAA
# Read4: GtCGAAATCA
#
# But if there is contamination, this may not be the case. In the example below, half of the mapped reads have the T allele and half have the A.
#
# __Heterozygous SNP__
# Ref: CTTGAGACGAAATCACTAAAAAACGTGACGACTTG
# Read1: CTTGAGtCG
# Read2: CTTGAGaCGAAA
# Read3: GAGaCGAAATCACTAAAA
# Read4: GtCGAAATCA
#
# __Note if using the Sanger cluster:__ Heterozygous SNPs are calculated as part of the automated QC pipeline. The result for each lane is available in the file heterozygous_snps_report.txt.
#
# Congratulations! You have reached the end of this tutorial. You can find the answers to all the questions of the tutorial [here](contamination-answers.ipynb).
#
# To revisit the previous section [click here](assessment.ipynb). Alternatively you can head back to the [index page](index.ipynb)
| qc-training-notebooks/contamination.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# [](http://rpi.analyticsdojo.com)
# <center><h1>Intro to Tensorflow - MINST</h1></center>
# <center><h3><a href = 'http://rpi.analyticsdojo.com'>rpi.analyticsdojo.com</a></h3></center>
#
#
# [](https://colab.research.google.com/github/rpi-techfundamentals/fall2018-materials/blob/master/10-deep-learning/06-tensorflow-minst.ipynb)
#
# + [markdown] slideshow={"slide_type": "subslide"}
#
# Adopted from [Hands-On Machine Learning with Scikit-Learn and TensorFlow by <NAME>](https://github.com/ageron/handson-ml).
#
#
# Apache License
# Version 2.0, January 2004
# http://www.apache.org/licenses/
# TERMS AND CONDITIONS FOR USE, REPRODUCTION, AND DISTRIBUTION
#
# [For full license see repository.](https://github.com/ageron/handson-ml/blob/master/LICENSE)
#
#
# **Chapter 10 – Introduction to Artificial Neural Networks**
#
#
# -
# _This notebook contains all the sample code and solutions to the exercices in chapter 10._
# # Setup
# First, let's make sure this notebook works well in both python 2 and 3, import a few common modules, ensure MatplotLib plots figures inline and prepare a function to save the figures:
# +
# Common imports
import numpy as np
import os
# to make this notebook's output stable across runs
def reset_graph(seed=42):
tf.reset_default_graph()
tf.set_random_seed(seed)
np.random.seed(seed)
# To plot pretty figures
# %matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
plt.rcParams['axes.labelsize'] = 14
plt.rcParams['xtick.labelsize'] = 12
plt.rcParams['ytick.labelsize'] = 12
# Where to save the figures
PROJECT_ROOT_DIR = "/home/jovyan/techfundamentals-fall2017-materials/classes/13-deep-learning"
def save_fig(fig_id, tight_layout=True):
path = os.path.join(PROJECT_ROOT_DIR, 'images', fig_id + ".png")
print("Saving figure", fig_id)
if tight_layout:
plt.tight_layout()
plt.savefig(path, format='png', dpi=300)
# -
# ### MNIST
# - Very common machine learning library with goal to classify digits.
# - This example is using MNIST handwritten digits, which contains 55,000 examples for training and 10,000 examples for testing. The digits have been size-normalized and centered in a fixed-size image (28x28 pixels) with values from 0 to 1. For simplicity, each image has been flattened and converted to a 1-D numpy array of 784 features (28*28).
# 
#
# More info: http://yann.lecun.com/exdb/mnist/
#
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets("/tmp/data/")
X_train = mnist.train.images
X_test = mnist.test.images
y_train = mnist.train.labels.astype("int")
y_test = mnist.test.labels.astype("int")
print ("Training set: ", X_train.shape,"\nTest set: ", X_test.shape)
# List a few images and print the data to get a feel for it.
images = 2
for i in range(images):
#Reshape
x=np.reshape(X_train[i], [28, 28])
print(x)
plt.imshow(x, cmap=plt.get_cmap('gray_r'))
plt.show()
# print("Model prediction:", preds[i])
# ## TFLearn: Deep learning library featuring a higher-level API for TensorFlow
#
# - TFlearn is a modular and transparent deep learning library built on top of Tensorflow.
# - It was designed to provide a higher-level API to TensorFlow in order to facilitate and speed-up experimentations
# - Fully transparent and compatible with Tensorflow
# - [DNN classifier](https://www.tensorflow.org/api_docs/python/tf/contrib/learn/DNNClassifier)
# - `hidden_units` list of hidden units per layer. All layers are fully connected. Ex. [64, 32] means first layer has 64 nodes and second one has 32.
# - [Scikit learn wrapper for TensorFlow Learn Estimator](https://www.tensorflow.org/api_docs/python/tf/contrib/learn/SKCompat)
# - See [tflearn documentation](http://tflearn.org/).
#
import tensorflow as tf
config = tf.contrib.learn.RunConfig(tf_random_seed=42) # not shown in the config
feature_cols = tf.contrib.learn.infer_real_valued_columns_from_input(X_train)
# List of hidden units per layer. All layers are fully connected. Ex. [64, 32] means first layer has 64 nodes and second one has 32.
dnn_clf = tf.contrib.learn.DNNClassifier(hidden_units=[300,100], n_classes=10,
feature_columns=feature_cols, config=config)
dnn_clf = tf.contrib.learn.SKCompat(dnn_clf) # if TensorFlow >= 1.1
dnn_clf.fit(X_train, y_train, batch_size=50, steps=4000)
# +
#We can use the sklearn version of metrics
from sklearn import metrics
y_pred = dnn_clf.predict(X_test)
#This calculates the accuracy.
print("Accuracy score: ", metrics.accuracy_score(y_test, y_pred['classes']) )
#Log Loss is a way of score classes probabilsitically
print("Logloss: ",metrics.log_loss(y_test, y_pred['probabilities']))
# -
# ### Tensorflow
# - Direct access to Python API for Tensorflow will give more flexibility
# - Like earlier, we will define the structure and then run the session.
# - set placeholders
# +
import tensorflow as tf
n_inputs = 28*28 # MNIST
n_hidden1 = 300 # hidden units in first layer.
n_hidden2 = 100
n_outputs = 10 # Classes of output variable.
# -
#Placehoder
reset_graph()
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int64, shape=(None), name="y")
def neuron_layer(X, n_neurons, name, activation=None):
with tf.name_scope(name):
n_inputs = int(X.get_shape()[1])
stddev = 2 / np.sqrt(n_inputs)
init = tf.truncated_normal((n_inputs, n_neurons), stddev=stddev)
W = tf.Variable(init, name="kernel")
b = tf.Variable(tf.zeros([n_neurons]), name="bias")
Z = tf.matmul(X, W) + b
if activation is not None:
return activation(Z)
else:
return Z
with tf.name_scope("dnn"):
hidden1 = neuron_layer(X, n_hidden1, name="hidden1", activation=tf.nn.relu)
hidden2 = neuron_layer(hidden1, n_hidden2, name="hidden2", activation=tf.nn.relu)
logits = neuron_layer(hidden2, n_outputs, name="outputs")
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name="loss")
# +
learning_rate = 0.01
with tf.name_scope("train"):
optimizer = tf.train.GradientDescentOptimizer(learning_rate)
training_op = optimizer.minimize(loss)
# -
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32))
init = tf.global_variables_initializer()
saver = tf.train.Saver()
# ### Running the Analysis over 40 Epocs
# - 40 passes through entire dataset.
# -
n_epochs = 40
batch_size = 50
with tf.Session() as sess:
init.run()
for epoch in range(n_epochs):
for iteration in range(mnist.train.num_examples // batch_size):
X_batch, y_batch = mnist.train.next_batch(batch_size)
sess.run(training_op, feed_dict={X: X_batch, y: y_batch})
acc_train = accuracy.eval(feed_dict={X: X_batch, y: y_batch})
acc_test = accuracy.eval(feed_dict={X: mnist.test.images,
y: mnist.test.labels})
print("Epoc:", epoch, "Train accuracy:", acc_train, "Test accuracy:", acc_test)
save_path = saver.save(sess, "./my_model_final.ckpt")
with tf.Session() as sess:
saver.restore(sess, "./my_model_final.ckpt") # or better, use save_path
X_new_scaled = mnist.test.images[:20]
Z = logits.eval(feed_dict={X: X_new_scaled})
y_pred = np.argmax(Z, axis=1)
print("Predicted classes:", y_pred)
print("Actual classes: ", mnist.test.labels[:20])
# +
from IPython.display import clear_output, Image, display, HTML
def strip_consts(graph_def, max_const_size=32):
"""Strip large constant values from graph_def."""
strip_def = tf.GraphDef()
for n0 in graph_def.node:
n = strip_def.node.add()
n.MergeFrom(n0)
if n.op == 'Const':
tensor = n.attr['value'].tensor
size = len(tensor.tensor_content)
if size > max_const_size:
tensor.tensor_content = "<stripped %d bytes>"%size
return strip_def
def show_graph(graph_def, max_const_size=32):
"""Visualize TensorFlow graph."""
if hasattr(graph_def, 'as_graph_def'):
graph_def = graph_def.as_graph_def()
strip_def = strip_consts(graph_def, max_const_size=max_const_size)
code = """
<script>
function load() {{
document.getElementById("{id}").pbtxt = {data};
}}
</script>
<link rel="import" href="https://tensorboard.appspot.com/tf-graph-basic.build.html" onload=load()>
<div style="height:600px">
<tf-graph-basic id="{id}"></tf-graph-basic>
</div>
""".format(data=repr(str(strip_def)), id='graph'+str(np.random.rand()))
iframe = """
<iframe seamless style="width:1200px;height:620px;border:0" srcdoc="{}"></iframe>
""".format(code.replace('"', '"'))
display(HTML(iframe))
# -
show_graph(tf.get_default_graph())
# ## Using `dense()` instead of `neuron_layer()`
# Note: the book uses `tensorflow.contrib.layers.fully_connected()` rather than `tf.layers.dense()` (which did not exist when this chapter was written). It is now preferable to use `tf.layers.dense()`, because anything in the contrib module may change or be deleted without notice. The `dense()` function is almost identical to the `fully_connected()` function, except for a few minor differences:
# * several parameters are renamed: `scope` becomes `name`, `activation_fn` becomes `activation` (and similarly the `_fn` suffix is removed from other parameters such as `normalizer_fn`), `weights_initializer` becomes `kernel_initializer`, etc.
# * the default `activation` is now `None` rather than `tf.nn.relu`.
# * a few more differences are presented in chapter 11.
n_inputs = 28*28 # MNIST
n_hidden1 = 300
n_hidden2 = 100
n_outputs = 10
# +
reset_graph()
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int64, shape=(None), name="y")
# -
with tf.name_scope("dnn"):
hidden1 = tf.layers.dense(X, n_hidden1, name="hidden1",
activation=tf.nn.relu)
hidden2 = tf.layers.dense(hidden1, n_hidden2, name="hidden2",
activation=tf.nn.relu)
logits = tf.layers.dense(hidden2, n_outputs, name="outputs")
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name="loss")
# +
learning_rate = 0.01
with tf.name_scope("train"):
optimizer = tf.train.GradientDescentOptimizer(learning_rate)
training_op = optimizer.minimize(loss)
# -
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32))
init = tf.global_variables_initializer()
saver = tf.train.Saver()
# +
n_epochs = 20
n_batches = 50
with tf.Session() as sess:
init.run()
for epoch in range(n_epochs):
for iteration in range(mnist.train.num_examples // batch_size):
X_batch, y_batch = mnist.train.next_batch(batch_size)
sess.run(training_op, feed_dict={X: X_batch, y: y_batch})
acc_train = accuracy.eval(feed_dict={X: X_batch, y: y_batch})
acc_test = accuracy.eval(feed_dict={X: mnist.test.images, y: mnist.test.labels})
print(epoch, "Train accuracy:", acc_train, "Test accuracy:", acc_test)
save_path = saver.save(sess, "./my_model_final.ckpt")
# -
show_graph(tf.get_default_graph())
| site/_build/html/_sources/notebooks/09-deep-learning1/06_tensorflow_minst.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.7 64-bit (''aipy'': conda)'
# name: python3
# ---
# # Image classifier project
# ## Configure notebook
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
# ## Import modules
# +
import time
import matplotlib.pyplot as plt
import numpy as np
import torch
from torch import nn
from torch import optim
import torch.nn.functional as F
from torchvision import datasets, transforms, models, utils
# -
# ## Configure torch device
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# ## Configure the locations of the source data
data_dir = 'flowers'
training_data_dir = data_dir + '/train'
validation_data_dir = data_dir + '/valid'
testing_data_dir = data_dir + '/test'
# ## Define transforms, datasets and dataloaders
# +
# Define transforms for the training, validation, and testing sets
normalised_transform = transforms.Normalize([0.485, 0.456, 0.406],
[0.229, 0.224, 0.225])
training_transforms = transforms.Compose([transforms.RandomRotation(30),
transforms.RandomResizedCrop(224),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
normalised_transform])
validation_transforms = transforms.Compose([transforms.Resize(255),
transforms.CenterCrop(224),
transforms.ToTensor(),
normalised_transform])
testing_transforms = transforms.Compose([transforms.Resize(255),
transforms.CenterCrop(224),
transforms.ToTensor(),
normalised_transform])
# Load the datasets with ImageFolder
training_dataset = datasets.ImageFolder(training_data_dir,
transform=training_transforms)
validation_dataset = datasets.ImageFolder(validation_data_dir,
transform=validation_transforms)
testing_dataset = datasets.ImageFolder(testing_data_dir,
transform=testing_transforms)
# Define the dataloaders using the image datasets and trainforms
training_dataloader = torch.utils.data.DataLoader(training_dataset,
batch_size=64,
shuffle=True,
pin_memory=True)
validation_dataloader = torch.utils.data.DataLoader(validation_dataset,
batch_size=64,
shuffle=True,
pin_memory=True)
testing_dataloader = torch.utils.data.DataLoader(testing_dataset,
batch_size=64,
shuffle=True,
pin_memory=True)
# +
import json
with open('cat_to_name.json', 'r') as file:
cat_to_name = json.load(file)
len(cat_to_name)
# -
# ## Define the model
def create_model_densenet121(hidden_size=256, lr=0.003):
model = models.densenet121(pretrained=True)
# Freeze parameters so we don't backprop through them
for param in model.parameters():
param.requires_grad = False
model.classifier = nn.Sequential(nn.Linear(1024, hidden_size),
nn.ReLU(),
nn.Dropout(0.2),
nn.Linear(hidden_size, 102),
nn.LogSoftmax(dim=1))
criterion = nn.NLLLoss()
# Only train the classifier parameters, feature parameters are frozen
optimizer = optim.Adam(model.classifier.parameters(), lr)
return model, optimizer, criterion
def create_model_densenet121_1():
model = models.densenet121(pretrained=True)
# Freeze parameters so we don't backprop through them
for param in model.parameters():
param.requires_grad = False
model.classifier = nn.Sequential(nn.Linear(1024, 512),
nn.ReLU(),
nn.Dropout(0.2),
nn.Linear(512, 256),
nn.ReLU(),
nn.Dropout(0.2),
nn.Linear(256, 102),
nn.LogSoftmax(dim=1))
criterion = nn.NLLLoss()
# Only train the classifier parameters, feature parameters are frozen
optimizer = optim.Adam(model.classifier.parameters(), lr=0.0005)
return model, optimizer, criterion
def create_model_resnet101():
model = models.resnet101(pretrained=True)
# Freeze parameters so we don't backprop through them
for param in model.parameters():
param.requires_grad = False
model.fc = nn.Sequential(nn.Linear(2048, 256),
nn.ReLU(),
nn.Dropout(0.2),
nn.Linear(256, 102),
nn.LogSoftmax(dim=1))
criterion = nn.NLLLoss()
# Only train the classifier parameters, feature parameters are frozen
optimizer = optim.Adam(model.fc.parameters(), lr=0.003)
return model, optimizer, criterion
def create_model_vgg11():
model = models.vgg11(pretrained=True)
# Freeze parameters so we don't backprop through them
for param in model.parameters():
param.requires_grad = False
model.classifier[-1] = nn.Sequential(nn.Linear(4096, 256),
nn.ReLU(),
nn.Dropout(0.2),
nn.Linear(256, 102),
nn.LogSoftmax(dim=1))
criterion = nn.NLLLoss()
# Only train the classifier parameters, feature parameters are frozen
optimizer = optim.Adam(model.classifier.parameters(), lr=0.003)
return model, optimizer, criterion
# ## Train the model
# +
epochs = 15
steps = 0
hidden_size = 256
lr = 0.0005
model, optimizer, criterion = create_model_densenet121(hidden_size, lr)
model.to(device)
training_losses, validation_losses = [], []
for e in range(epochs):
start_time = time.time()
total_training_loss = 0
model.train()
for images, expected_results in training_dataloader:
images, expected_results = images.to(device), expected_results.to(device)
optimizer.zero_grad()
log_ps = model(images)
loss = criterion(log_ps, expected_results)
total_training_loss += loss.item()
loss.backward()
optimizer.step()
total_validation_loss = 0
total_correctly_predicted = 0
model.eval()
with torch.no_grad():
for images, expected_results in validation_dataloader:
images, expected_results = images.to(device), expected_results.to(device)
log_ps = model(images)
loss = criterion(log_ps, expected_results)
total_validation_loss += loss.item()
ps = torch.exp(log_ps)
top_p, top_class = ps.topk(1, dim=1)
correct_predictions = top_class == expected_results.view(*top_class.shape)
total_correctly_predicted += correct_predictions.sum().item()
mean_training_loss = total_training_loss / len(training_dataloader.dataset)
mean_validation_loss = total_validation_loss / len(validation_dataloader.dataset)
mean_correctly_predicted = total_correctly_predicted / len(validation_dataloader.dataset)
training_losses.append(mean_training_loss)
validation_losses.append(mean_validation_loss)
duration = time.time() - start_time
print("Epoch: {}/{}.. ".format(e+1, epochs),
"Training Loss: {:.5f}.. ".format(mean_training_loss),
"Validation Loss: {:.5f}.. ".format(mean_validation_loss),
"Accuracy: {:.3f}.. ".format(mean_correctly_predicted),
"Duration: {:.1f}s".format(duration))
# -
plt.plot(training_losses, label='Training loss')
plt.plot(validation_losses, label='Validation loss')
plt.legend(frameon=False);
# ## Save and restore the model
checkpoint_file_path = "flwr_class_model.pth"
# +
checkpoint = {'feature_model': 'densenet121',
'hidden_size': hidden_size,
'model_state': model.state_dict()}
torch.save(checkpoint, checkpoint_file_path)
# -
checkpoint = None
checkpoint = torch.load(checkpoint_file_path)
model, optimizer, criterion = create_model_densenet121(hidden_size=checkpoint['hidden_size'])
model.load_state_dict(checkpoint['model_state'])
# ## Test the model
# +
from PIL import Image
def process_image(image):
''' Scales, crops, and normalizes a PIL image for a PyTorch model,
returns an Numpy array
'''
shortest_side_size = 256
crop_size = 224
scale_factor = min(image.size) / shortest_side_size
new_size = tuple([int(x / scale_factor) for x in image.size])
image = image.resize(new_size)
crop_width = (image.width - crop_size) // 2
crop_height = (image.height - crop_size) // 2
image = image.crop((crop_width,
crop_height,
image.width - crop_width,
image.height - crop_height))
np_image = np.array(image)
mean = np.array([0.485, 0.456, 0.406])
std = np.array([0.229, 0.224, 0.225])
np_image = (np_image / 255 - mean) / std
np_image = np.transpose(np_image, (2, 0, 1))
return np_image
# -
def imshow(image, ax=None, title=None):
"""Imshow for Tensor."""
if ax is None:
fig, ax = plt.subplots()
# PyTorch tensors assume the color channel is the first dimension
# but matplotlib assumes is the third dimension
image = image.numpy().transpose((1, 2, 0))
# Undo preprocessing
mean = np.array([0.485, 0.456, 0.406])
std = np.array([0.229, 0.224, 0.225])
image = std * image + mean
# Image needs to be clipped between 0 and 1 or it looks like noise when displayed
image = np.clip(image, 0, 1)
ax.imshow(image)
return ax
# +
image_path = "flowers/test/5/image_05159.jpg"
with Image.open(image_path) as im:
imshow(torch.from_numpy(process_image(im)))
# -
# ## Another test tool
def view_image_prediction(img, ps):
''' Function for viewing an image and it's predicted classes.
'''
ps = ps.data.numpy().squeeze()
fig, (ax1, ax2) = plt.subplots(figsize=(6,9), ncols=2)
ax1.imshow(img.resize_(1, 224, 224).numpy().squeeze())
ax1.axis('off')
ax2.barh(np.arange(10), ps)
ax2.set_aspect(0.1)
ax2.set_yticks(np.arange(102))
ax2.set_yticklabels(np.arange(102))
ax2.set_title('Class Probability')
ax2.set_xlim(0, 1.1)
plt.tight_layout()
def imshow2(inp, title=None):
"""Imshow for Tensor."""
inp = inp.numpy().transpose((1, 2, 0))
mean = np.array([0.485, 0.456, 0.406])
std = np.array([0.229, 0.224, 0.225])
inp = std * inp + mean
inp = np.clip(inp, 0, 1)
plt.imshow(inp)
if title is not None:
plt.title(title)
plt.pause(0.001) # pause a bit so that plots are updated
def visualize_model(model, dataloader, num_images=6):
was_training = model.training
model.eval()
images_so_far = 0
fig = plt.figure()
with torch.no_grad():
for i, (inputs, labels) in enumerate(dataloader):
inputs = inputs.to(device)
labels = labels.to(device)
outputs = model(inputs)
_, preds = torch.max(outputs, 1)
for j in range(inputs.size()[0]):
images_so_far += 1
ax = plt.subplot(num_images//2, 2, images_so_far)
ax.axis('off')
#print(cat_to_name[str(preds[j].item() + 1)])
ax.set_title('predicted: {}, actual: {}'.format(cat_to_name[str(preds[j].item() + 1)],
cat_to_name[str(labels[j].item() + 1)]))
imshow2(inputs.cpu().data[j])
if images_so_far == num_images:
model.train(mode=was_training)
return
model.train(mode=was_training)
model.to(device)
visualize_model(model, testing_dataloader, num_images=10)
| Sandpit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Internal Mutations Report
# This notebook contains developmental code for analyzing mutations in SARS-CoV-2 samples released by Andersen Lab.
import pandas as pd
from path import Path
from mutations import *
from onion_trees import *
from bjorn_support import *
from shutil import copy
df = pd.read_csv('/home/al/analysis/alab_mutations_13-01-2021/alab_substitutions_wide_13-01-2021.csv')
df.head()
df['samples'].iloc[0]
msa_fp = Path('/home/al/analysis/mutations/S501Y/msa_reference.fa')
patient_zero = 'NC_045512.2'
fa_fp = Path('/home/al/analysis/mutations/S501Y/sequences_2020-12-20_12-04.fasta')
ref_fp = '/home/al/data/test_inputs/NC045512.fasta'
align_fasta_reference(fa_fp, num_cpus=25, ref_fp=ref_fp)
cns = AlignIO.read(msa_fp, 'fasta')
print(f"Initial cleaning...")
seqs, ref_seq = process_cns_seqs(cns, patient_zero,
start_pos=0, end_pos=30000)
seqsdf = identify_replacements_per_sample(seqs, None, ref_seq, GENE2POS)
seqsdf.columns
subs = (seqsdf.groupby(['gene', 'ref_codon', 'alt_codon', 'pos', 'ref_aa',
'codon_num', 'alt_aa'])
.agg(
num_samples=('idx', 'nunique'),
# first_detected=('date', 'min'),
# last_detected=('date', 'max'),
# locations=('location', uniq_locs),
# location_counts=('location',
# lambda x: np.unique(x, return_counts=True)),
samples=('idx', 'unique')
)
.reset_index())
subs.to_csv('b117_usa_substitutions.csv', index=False)
# dels
start_pos = 265
end_pos =29674
gene2pos = GENE2POS
seqs, ref_seq = process_cns_seqs(cns, patient_zero, start_pos, end_pos)
dels = identify_deletions_per_sample(seqs, None, 1)
del_seqs = (dels.groupby(['relative_coords', 'del_len'])
.agg(samples=('idx', 'unique'),
num_samples=('idx', 'nunique'))
# first_detected=('date', 'min'),
# last_detected=('date', 'max'),
# locations=('location', uniq_locs),
# location_counts=('location',
# lambda x: np.unique(x, return_counts=True)))
.reset_index()
.sort_values('num_samples'))
# del_seqs['locations'] = del_seqs['location_counts'].apply(lambda x: list(x[0]))
# del_seqs['location_counts'] = del_seqs['location_counts'].apply(lambda x: list(x[1]))
del_seqs['type'] = 'deletion'
# adjust coordinates to account for the nts trimmed from beginning e.g. 265nts
del_seqs['absolute_coords'] = del_seqs['relative_coords'].apply(adjust_coords, args=(start_pos+1,))
del_seqs['pos'] = del_seqs['absolute_coords'].apply(lambda x: int(x.split(':')[0]))
# approximate the gene where each deletion was identified
del_seqs['gene'] = del_seqs['pos'].apply(map_gene_to_pos)
del_seqs = del_seqs.loc[~del_seqs['gene'].isna()]
# filter our substitutions in non-gene positions
del_seqs = del_seqs.loc[del_seqs['gene']!='nan']
# compute codon number of each substitution
del_seqs['codon_num'] = del_seqs.apply(compute_codon_num, args=(gene2pos,), axis=1)
# fetch the reference codon for each substitution
del_seqs['ref_codon'] = del_seqs.apply(get_ref_codon, args=(ref_seq, gene2pos), axis=1)
# fetch the reference and alternative amino acids
del_seqs['ref_aa'] = del_seqs['ref_codon'].apply(get_aa)
# record the 5 nts before each deletion (based on reference seq)
del_seqs['prev_5nts'] = del_seqs['absolute_coords'].apply(lambda x: ref_seq[int(x.split(':')[0])-5:int(x.split(':')[0])])
# record the 5 nts after each deletion (based on reference seq)
del_seqs['next_5nts'] = del_seqs['absolute_coords'].apply(lambda x: ref_seq[int(x.split(':')[1])+1:int(x.split(':')[1])+6])
del_seqs[['type', 'gene', 'absolute_coords', 'del_len', 'pos',
'ref_aa', 'codon_num', 'num_samples','samples',
'ref_codon', 'prev_5nts', 'next_5nts'
]].to_csv('b117_usa_deletions.csv', index=False)
in_dir = Path('/home/al/code/HCoV-19-Genomics/consensus_sequences/')
out_dir = Path('/home/al/analysis/mutations/alab_git/cns_seqs')
ref_fp = '/home/al/data/test_inputs/NC045512.fasta'
# fa_fp = concat_fasta(in_dir, out_dir)
# +
# align_fasta_reference(fa_fp, num_cpus=25, ref_fp=ref_fp)
# -
in_dir.listdir()[0]
for filename in in_dir.listdir():
if ((filename.endswith('fa') or filename.endswith('fasta')) and ('JOR' not in filename)):
copy(filename, '/home/al/analysis/mutations/alab_git/b117/fa')
meta = pd.read_csv('/home/al/analysis/mutations/alab_git/nextstrain_groups_neherlab_ncov_S.N501_metadata.tsv', sep='\t')
strains = meta['Strain'].unique().tolist()
len(strains)
gisaid_seqs = Path('/home/al/analysis/mutations/gisaid/data/sequences_2020-12-29_08-08.fasta')
seqs_data = SeqIO.to_dict(SeqIO.parse(gisaid_seqs, "fasta"))
len(seqs_data.keys())
my_seqs = {}
for name in seqs_data.keys():
if name in strains:
my_seqs[name] = seqs_data[name]
len(my_seqs)
out_dir = Path('/home/al/analysis/mutations/alab_git/b117')
with open(out_dir/'b117_seqs.fa', 'w') as handle:
SeqIO.write(my_seqs.values(), handle, 'fasta')
concat_fasta('/home/al/analysis/mutations/alab_git/b117/fa',
'/home/al/analysis/mutations/alab_git/b117/alab_usa')
fasta_filepath = out_dir/'b117_seqs.fa'
align_fasta_reference(fasta_filepath, ref_fp=)
consensus_data = SeqIO.to_dict(SeqIO.parse(aligned_seqs, "fasta"))
fa_fp = Path('/home/al/analysis/mutations/alab_git/b117/b117_usa.fasta')
msa_fp = align_fasta_reference(fa_fp, ref_fp='/home/al/data/test_inputs/NC045512.fasta')
msa_fp =
patient_zero = 'NC_045512.2'
cns = AlignIO.read(msa_fp, 'fasta')
print(f"Initial cleaning...")
seqs, ref_seq = process_cns_seqs(cns, patient_zero,
start_pos=0, end_pos=30000)
seqsdf = identify_replacements_per_sample(seqs, None, ref_seq, GENE2POS)
seqsdf.columns
subs = (seqsdf.groupby(['gene', 'ref_codon', 'alt_codon', 'pos', 'ref_aa',
'codon_num', 'alt_aa'])
.agg(
num_samples=('idx', 'nunique'),
# first_detected=('date', 'min'),
# last_detected=('date', 'max'),
# locations=('location', uniq_locs),
# location_counts=('location',
# lambda x: np.unique(x, return_counts=True)),
samples=('idx', 'unique')
)
.reset_index())
subs.to_csv('b117_usa_substitutions.csv', index=False)
# dels
start_pos = 265
end_pos =29674
gene2pos = GENE2POS
seqs, ref_seq = process_cns_seqs(cns, patient_zero, start_pos, end_pos)
dels = identify_deletions_per_sample(seqs, None, 1)
del_seqs = (dels.groupby(['relative_coords', 'del_len'])
.agg(samples=('idx', 'unique'),
num_samples=('idx', 'nunique'))
# first_detected=('date', 'min'),
# last_detected=('date', 'max'),
# locations=('location', uniq_locs),
# location_counts=('location',
# lambda x: np.unique(x, return_counts=True)))
.reset_index()
.sort_values('num_samples'))
# del_seqs['locations'] = del_seqs['location_counts'].apply(lambda x: list(x[0]))
# del_seqs['location_counts'] = del_seqs['location_counts'].apply(lambda x: list(x[1]))
del_seqs['type'] = 'deletion'
# adjust coordinates to account for the nts trimmed from beginning e.g. 265nts
del_seqs['absolute_coords'] = del_seqs['relative_coords'].apply(adjust_coords, args=(start_pos+1,))
del_seqs['pos'] = del_seqs['absolute_coords'].apply(lambda x: int(x.split(':')[0]))
# approximate the gene where each deletion was identified
del_seqs['gene'] = del_seqs['pos'].apply(map_gene_to_pos)
del_seqs = del_seqs.loc[~del_seqs['gene'].isna()]
# filter our substitutions in non-gene positions
del_seqs = del_seqs.loc[del_seqs['gene']!='nan']
# compute codon number of each substitution
del_seqs['codon_num'] = del_seqs.apply(compute_codon_num, args=(gene2pos,), axis=1)
# fetch the reference codon for each substitution
del_seqs['ref_codon'] = del_seqs.apply(get_ref_codon, args=(ref_seq, gene2pos), axis=1)
# fetch the reference and alternative amino acids
del_seqs['ref_aa'] = del_seqs['ref_codon'].apply(get_aa)
# record the 5 nts before each deletion (based on reference seq)
del_seqs['prev_5nts'] = del_seqs['absolute_coords'].apply(lambda x: ref_seq[int(x.split(':')[0])-5:int(x.split(':')[0])])
# record the 5 nts after each deletion (based on reference seq)
del_seqs['next_5nts'] = del_seqs['absolute_coords'].apply(lambda x: ref_seq[int(x.split(':')[1])+1:int(x.split(':')[1])+6])
del_seqs[['type', 'gene', 'absolute_coords', 'del_len', 'pos',
'ref_aa', 'codon_num', 'num_samples','samples',
'ref_codon', 'prev_5nts', 'next_5nts'
]].to_csv('b117_usa_deletions.csv', index=False)
del_seqs[['type', 'gene', 'absolute_coords', 'del_len', 'pos',
'ref_aa', 'codon_num', 'num_samples','samples',
'ref_codon', 'prev_5nts', 'next_5nts'
]].to_csv('b117_usa_deletions.csv', index=False)
del
dels = identify_deletions_per_sample(seqs, None, 1)
dels
seqsdf['pos'] = seqsdf['pos'] + 1
positions = [3267, 5388, 6954, 11288, 23063, 23271, 23604, 23709, 24506, 24914,
27972, 28048, 28111, 28280, 28977]
seqsdf[seqsdf['pos'].isin(positions)]
in_dir = Path('/home/al/code/HCoV-19-Genomics/consensus_sequences/')
out_dir = Path('/home/al/analysis/mutations/alab_git')
meta_fp = Path('/home/al/code/HCoV-19-Genomics/metadata.csv')
msa_fp = '/home/al/analysis/mutations/alab_git/cns_seqs_aligned.fa'
patient_zero = 'NC_045512.2'
cns = AlignIO.read(msa_fp, 'fasta')
seqs, ref_seq = process_cns_seqs(cns, patient_zero,
start_pos=0, end_pos=30000)
seqs_df = identify_replacements_per_sample(seqs,
meta_fp,
ref_seq,
GENE2POS)
# +
q957l = seqs_df.loc[(seqs_df['gene']=='S') & (seqs_df['codon_num']==957) & (seqs_df['alt_aa']=='L')]
q957l.date.min(), q957l.date.max()
# -
seqs_df['location'].unique()
jor_seqs = seqs_df[seqs_df['location'].str.contains('Jordan')]
print(jor_seqs['idx'].unique().shape)
mutation_filter = (jor_seqs['gene']=='S') & (jor_seqs['codon_num']==957) & (jor_seqs['alt_aa']=='L')
jor_seqs['q957l'] = False
jor_seqs.loc[mutation_filter, 'q957l'] = True
jor_seqs['q957l'].value_counts()
# **Out of the 8046 total substitution-based mutations, 388 were Q957L among our Jordanian samples that have been released so far.**
jor_seqs['date'].min(), jor_seqs['date'].max()
jor_seqs[jor_seqs['q957l']]['date'].min(), jor_seqs[jor_seqs['q957l']]['date'].max()
jor_seqs['idx']
jor_seqs.columns
all_samples = jor_seqs['idx'].unique()
len(all_samples)
mutant_samples = jor_seqs[jor_seqs['q957l']]['idx'].unique()
len(mutant_samples)
jor_seqs.loc[:, 'is_nonsyn_mutation'] = False
jor_seqs.loc[jor_seqs['alt_aa']!=jor_seqs['ref_aa'], 'is_nonsyn_mutation'] = True
jor_seqs.loc[:, 'is_S_nonsyn_mutation'] = False
jor_seqs.loc[(jor_seqs['alt_aa']!=jor_seqs['ref_aa']) & (jor_seqs['gene']=='S'), 'is_S_nonsyn_mutation'] = True
mnthly_cnts = jor_seqs.groupby('month').agg(total_samples=('idx', 'nunique'),
mutated_samples=('q957l', 'sum'))
mnthly_cnts['mutation_freq'] = mnthly_cnts['mutated_samples'] / mnthly_cnts['total_samples']
mnthly_cnts
jor_seqs[['is_nonsyn_mutation', 'is_S_nonsyn_mutation']].sum() / len(all_samples)
jor_seqs[jor_seqs['idx'].isin(mutant_samples)][['is_nonsyn_mutation', 'is_S_nonsyn_mutation']].sum() / len(mutant_samples)
jor_seqs[~jor_seqs['idx'].isin(mutant_samples)][['is_nonsyn_mutation', 'is_S_nonsyn_mutation']].sum() / (len(all_samples) - len(mutant_samples))
# +
print("All Samples\n")
print(f"{len(all_samples)} total samples from Jordan")
print(f"An average of 11.59 non-synonymous mutations per sample")
print(f"An average of 1.98 non-synonymous mutations in S gene per sample\n")
print("Mutated Samples\n")
print(f"{len(mutant_samples)} total samples from Jordan with Q957L mutation")
print(f"An average of 12.47 non-synonymous mutations per sample")
print(f"An average of 2.20 non-synonymous mutations in S gene per sample\n")
print("Non-mutated Samples\n")
print(f"{len(all_samples)-len(mutant_samples)} total samples from Jordan without Q957L mutation")
print(f"An average of 8.25 non-synonymous mutations per sample")
print(f"An average of 1.18 non-synonymous mutations in S gene per sample")
# -
mutants = jor_seqs[(jor_seqs['idx'].isin(mutant_samples))].groupby('idx').agg(nonsyn_mutations=('is_nonsyn_mutation', 'sum'), s_nonsyn_mutations=('is_S_nonsyn_mutation', 'sum'))
print(mutants.shape)
mutants.head()
nonmutants = jor_seqs[(~jor_seqs['idx'].isin(mutant_samples))].groupby('idx').agg(nonsyn_mutations=('is_nonsyn_mutation', 'sum'), s_nonsyn_mutations=('is_S_nonsyn_mutation', 'sum'))
print(nonmutants.shape)
nonmutants.head()
ttest_ind(mutants['nonsyn_mutations'].values, nonmutants['nonsyn_mutations'].values)
ttest_ind(mutants['s_nonsyn_mutations'].values, nonmutants['s_nonsyn_mutations'].values)
df = pd.read_csv("/home/al/analysis/2020-12-24_release/insertions.csv")
df
codons_of_interest = [69, 70, 144, 145, 681, 716, 982, 1118, 501, 570]
df[(df['gene']=='S') & (df['codon_num'].isin(codons_of_interest))].sort_values('num_samples', ascending=False).iloc[0].values
dels = pd.read_csv("/home/al/analysis/2020-12-24_release/deletions.csv")
dels[(dels['gene']=='S') & (dels['codon_num'].isin(codons_of_interest))].sort_values('num_samples', ascending=False)
subs = pd.read_csv("/home/al/code/HCoV-19-Genomics/variants/substitutions_22-12-2020.csv")
subs[(subs['gene']=='S') & (subs['codon_num'].isin(codons_of_interest))].sort_values('num_samples', ascending=False)
codons_of_interest = [501, 957, 67, 68, 69, 417, 484]
(df[(df['gene']=='S') & (df['codon_num'].isin(codons_of_interest))]
.sort_values('num_samples', ascending=False)
.iloc[:99])
ddf = pd.read_csv("/home/al/analysis/2020-12-24_release/deletions.csv")
(ddf[(ddf['gene']=='S') & (ddf['codon_num'].isin(codons_of_interest))]
.sort_values('num_samples', ascending=False)
.iloc[:99])
df = pd.read_csv("/home/al/data/release_summary_csv/release_summary_24_12_2020.csv")
def check_date(x):
if type(x)==str: return True
return False
df = df[df['Collection date'].apply(check_date)]
Path.isfile()
# clean and process sample collection dates
df = df.loc[(df['Collection date']!='Unknown')
& (df['Collection date']!='1900-01-00')]
df.loc[df['Collection date'].str.contains('/'), 'Collection date'] = df['Collection date'].apply(lambda x: x.split('/')[0])
df['date'] = pd.to_datetime(df['Collection date'])
df.columns
df['idx'] = df['SEARCH SampleID'].apply(lambda x: x.split('-')[1]).astype(int)
df.loc[df['idx']>5325, 'New sequences ready for release'] = 'Yes'
df.to_csv("/home/al/data/release_summary_csv/release_summary_24_12_2020.csv", index=False)
import pandas as pd
from bjorn import *
from bjorn_support import *
from onion_trees import *
import gffutils
import math
from mutations import *
import ast
subs = pd.read_csv("/home/al/analysis/mutations/gisaid/gisaid_replacements_aggregated_19-12-2020.csv")
dels = pd.read_csv("/home/al/analysis/mutations/gisaid/gisaid_deletions_aggregated_19-12-2020.csv")
strain_dels = dels[(dels['gene']=='S') & (dels['codon_num'].isin([67, 68, 69]))].drop_duplicates(subset=['absolute_coords']).reset_index(drop=True)
strain_subs = subs[(subs['gene']=='S') & (subs['codon_num'].isin([501, 417, 484]))].drop_duplicates(subset=['gene', 'codon_num', 'alt_aa']).reset_index(drop=True)
strain_subs.shape, strain_dels.shape
strain_subs.columns
def is_in(x, loc):
for i in eval(x):
if loc in i.lower():
return True
return False
strain_subs['isin_SD'] = strain_subs['locations'].apply(is_in, args=('san diego',))
strain_subs['isin_CA'] = strain_subs['divisions'].apply(is_in, args=('california',))
strain_subs['isin_US'] = strain_subs['countries'].apply(is_in, args=('usa',))
strain_dels.columns
strain_dels['isin_SD'] = strain_dels['locations'].apply(is_in, args=('san diego',))
strain_dels['isin_CA'] = strain_dels['divisions'].apply(is_in, args=('california',))
strain_dels['isin_US'] = strain_dels['countries'].apply(is_in, args=('usa',))
strain_dels.to_csv('/home/al/analysis/mutations/S501Y/gisaid_strain_deletions.csv', index=False)
strain_subs.to_csv('/home/al/analysis/mutations/S501Y/gisaid_strain_substitutions.csv', index=False)
strain_subs
in_dir = Path('/home/al/code/HCoV-19-Genomics/consensus_sequences/')
out_dir = Path('/home/al/analysis/mutations/alab_git')
meta_fp = Path('/home/al/code/HCoV-19-Genomics/metadata.csv')
fasta_fp = concat_fasta(in_dir, out_dir/'cns_seqs')
msa_fp = align_fasta_reference(fasta_fp, num_cpus=20, ref_fp='/home/gk/code/hCoV19/db/NC045512.fasta')
msa_fp = '/home/al/analysis/mutations/alab_git/cns_seqs_aligned.fa'
subs = identify_replacements(msa_fp, meta_fp)
subs.explode('samples')['samples'].unique().shape
subs.sort_values('num_samples', ascending=False).to_csv(out_dir/"substitutions_22-12-2020_orig.csv", index=False)
dels = identify_deletions(msa_fp, meta_fp, min_del_len=1)
dels.sort_values('num_samples', ascending=False).to_csv(out_dir/"deletions_22-12-2020_orig.csv", index=False)
align_fasta_reference(seqs_fp, num_cpus=25, ref_fp=ref_fp)
# ## CNS Mutations Report
analysis_folder = Path('/home/al/code/HCoV-19-Genomics/consensus_sequences/')
meta_fp = Path('/home/al/code/HCoV-19-Genomics/metadata.csv')
ref_path = Path('/home/gk/code/hCoV19/db/NC045512.fasta')
patient_zero = 'NC_045512.2'
in_fp = '/home/al/analysis/mutations/S501Y/msa_aligned.fa'
subs = identify_replacements(in_fp, meta_fp)
subs.head()
dels = identify_deletions(in_fp, meta_fp, patient_zero)
dels
dels[dels['gene']=='S'].sort_values('num_samples', ascending=False)#.to_csv('S_deletions_consensus.csv', index=False)
identify_insertions(in_fp, patient_zero).to_csv('test.csv', index=False)
# ## dev
GENE2POS = {
'5UTR': {'start': 0, 'end': 265},
'ORF1ab': {'start': 265, 'end': 21555},
'S': {'start': 21562, 'end': 25384},
'ORF3a': {'start': 25392, 'end': 26220},
'E': {'start': 26244, 'end': 26472},
'M': {'start': 26522, 'end': 27191},
'ORF6': {'start': 27201, 'end': 27387},
'ORF7a': {'start': 27393, 'end': 27759},
'ORF7b': {'start': 27755, 'end': 27887},
'ORF8': {'start': 27893, 'end': 28259},
'N': {'start': 28273, 'end': 29533},
'ORF10': {'start': 29557, 'end': 29674},
'3UTR': {'start': 29674, 'end': 29902}
}
in_dir = '/home/al/analysis/mutations/fa/'
out_dir = '/home/al/analysis/mutations/msa/'
# !rm -r /home/al/analysis/mutations
# !mkdir /home/al/analysis/mutations
# !mkdir /home/al/analysis/mutations/fa
for filename in analysis_folder.listdir():
if (filename.endswith('fa') or filename.endswith('fasta')):
copy(filename, '/home/al/analysis/mutations/fa/')
# print(filename)
copy(ref_path, in_dir)
in_dir = '/home/al/analysis/mutations/fa/'
out_dir = '/home/al/analysis/mutations/msa'
concat_fasta(in_dir, out_dir)
align_fasta_reference('/home/al/analysis/mutations/msa.fa', num_cpus=12, ref_fp=ref_path)
cns = AlignIO.read('/home/al/analysis/mutations/msa_aligned.fa', 'fasta')
ref_seq = get_seq(cns, patient_zero)
len(ref_seq)
seqs = get_seqs(cns, 0, 30000)
seqsdf = (pd.DataFrame(index=seqs.keys(), data=seqs.values(), columns=['sequence'])
.reset_index().rename(columns={'index': 'idx'}))
# +
# seqsdf
# -
def find_replacements(x, ref):
return [f'{i}:{n}' for i, n in enumerate(x)
if n!=ref[i] and n!='-' and n!='n']
seqsdf['replacements'] = seqsdf['sequence'].apply(find_replacements, args=(ref_seq,))
seqsdf = seqsdf.explode('replacements')
seqsdf['pos'] = -1
seqsdf.loc[~seqsdf['replacements'].isna(), 'pos'] = seqsdf.loc[~seqsdf['replacements'].isna(), 'replacements'].apply(lambda x: int(x.split(':')[0]))
seqsdf = seqsdf.loc[seqsdf['pos']!=-1]
def compute_codon_num(x, gene2pos: dict):
pos = x['pos']
ref_pos = gene2pos[x['gene']]['start']
return math.ceil((pos - ref_pos + 1) / 3)
seqsdf['gene'] = seqsdf['pos'].apply(map_gene_to_pos)
seqsdf = seqsdf.loc[~seqsdf['gene'].isna()]
seqsdf = seqsdf.loc[seqsdf['gene']!='nan']
seqsdf['codon_num'] = seqsdf.apply(compute_codon_num, args=(GENE2POS,), axis=1)
def get_ref_codon(x, ref_seq, gene2pos: dict):
ref_pos = gene2pos[x['gene']]['start']
codon_start = ref_pos + ((x['codon_num'] - 1) * 3)
return ref_seq[codon_start: codon_start+3].upper()
seqsdf['ref_codon'] = seqsdf.apply(get_ref_codon, args=(ref_seq, GENE2POS), axis=1)
def get_alt_codon(x, gene2pos: dict):
ref_pos = gene2pos[x['gene']]['start']
codon_start = ref_pos + ((x['codon_num'] - 1) * 3)
return x['sequence'][codon_start: codon_start+3].upper()
seqsdf['alt_codon'] = seqsdf.apply(get_alt_codon, args=(GENE2POS,), axis=1)
def get_aa(codon: str):
CODON2AA = {
'ATA':'I', 'ATC':'I', 'ATT':'I', 'ATG':'M',
'ACA':'T', 'ACC':'T', 'ACG':'T', 'ACT':'T',
'AAC':'N', 'AAT':'N', 'AAA':'K', 'AAG':'K',
'AGC':'S', 'AGT':'S', 'AGA':'R', 'AGG':'R',
'CTA':'L', 'CTC':'L', 'CTG':'L', 'CTT':'L',
'CCA':'P', 'CCC':'P', 'CCG':'P', 'CCT':'P',
'CAC':'H', 'CAT':'H', 'CAA':'Q', 'CAG':'Q',
'CGA':'R', 'CGC':'R', 'CGG':'R', 'CGT':'R',
'GTA':'V', 'GTC':'V', 'GTG':'V', 'GTT':'V',
'GCA':'A', 'GCC':'A', 'GCG':'A', 'GCT':'A',
'GAC':'D', 'GAT':'D', 'GAA':'E', 'GAG':'E',
'GGA':'G', 'GGC':'G', 'GGG':'G', 'GGT':'G',
'TCA':'S', 'TCC':'S', 'TCG':'S', 'TCT':'S',
'TTC':'F', 'TTT':'F', 'TTA':'L', 'TTG':'L',
'TAC':'Y', 'TAT':'Y', 'TAA':'_', 'TAG':'_',
'TGC':'C', 'TGT':'C', 'TGA':'_', 'TGG':'W',
}
return CODON2AA.get(codon, 'nan')
seqsdf['ref_aa'] = seqsdf['ref_codon'].apply(get_aa)
seqsdf['alt_aa'] = seqsdf['alt_codon'].apply(get_aa)
seqsdf = seqsdf.loc[seqsdf['alt_aa']!='nan']
seqsdf.columns
meta = pd.read_csv(meta_fp)
print(seqsdf['idx'].unique().shape)
seqsdf = pd.merge(seqsdf, meta, left_on='idx', right_on='fasta_hdr')
print(seqsdf['idx'].unique().shape)
seqsdf = seqsdf.loc[(seqsdf['collection_date']!='Unknown')
& (seqsdf['collection_date']!='1900-01-00')]
seqsdf.loc[seqsdf['collection_date'].str.contains('/'), 'collection_date'] = seqsdf['collection_date'].apply(lambda x: x.split('/')[0])
seqsdf['date'] = pd.to_datetime(seqsdf['collection_date'])
seqsdf['date'].min()
# +
# (seqsdf.groupby(['gene', 'ref_aa', 'codon_num', 'alt_aa'])
# .agg(
# num_samples=('ID', 'nunique')))
# -
def uniq_locs(x):
return np.unique(x)
def loc_counts(x):
_, counts = np.unique(x, return_counts=True)
return counts
subs = (seqsdf.groupby(['gene', 'pos', 'ref_aa', 'codon_num', 'alt_aa'])
.agg(
num_samples=('ID', 'nunique'),
first_detected=('date', 'min'),
last_detected=('date', 'max'),
locations=('location', uniq_locs),
location_counts=('location', loc_counts),
samples=('ID', 'unique')
)
.reset_index())
subs['pos'] = subs['pos'] + 1
(subs[subs['gene']=='S'].sort_values('num_samples', ascending=False)
.to_csv('S_mutations_consensus.csv', index=False))
# ## Consolidate metadata ID and fasta headers
def fix_header(x):
if 'Consensus' in x:
return x.split('_')[1]
else:
return x.split('/')[2]
seqsdf['n_ID'] = seqsdf['idx'].apply(fix_header)
seqsdf['n_ID'] = seqsdf['n_ID'].str.replace('ALSR', 'SEARCH')
meta = pd.read_csv(meta_fp)
meta['n_ID'] = meta['ID'].apply(lambda x: '-'.join(x.split('-')[:2]))
seqsdf['n_ID'] = seqsdf['n_ID'].apply(lambda x: '-'.join(x.split('-')[:2]))
tmp = pd.merge(seqsdf, meta, on='n_ID')
# +
# tmp[tmp['ID'].str.contains('2112')]
# +
# seqsdf
# -
set(meta['n_ID'].unique()) - set(tmp['n_ID'].unique())
seqsdf['idx'].unique().shape
meta['ID'].unique().shape
s = seqsdf[['n_ID', 'idx']].drop_duplicates()
new_meta = pd.merge(meta, s, on='n_ID', how='left')
(new_meta.drop(columns=['n_ID'])
.rename(columns={'idx': 'fasta_hdr'})
.to_csv('metadata.csv', index=False))
new_meta.shape
new_meta
len(ref_seq)
| notebooks/internal_mutations-Copy1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 2D Nuclear Segmentation with PanOpticFPN
# +
import os
import errno
import numpy as np
import tensorflow as tf
import deepcell
# -
# ## Load the data
#
# ### Download the data from `deepcell.datasets`
#
# `deepcell.datasets` provides access to a set of annotated live-cell imaging datasets which can be used for training cell segmentation and tracking models.
# All dataset objects share the `load_data()` method, which allows the user to specify the name of the file (`path`), the fraction of data reserved for testing (`test_size`) and a `seed` which is used to generate the random train-test split.
# Metadata associated with the dataset can be accessed through the `metadata` attribute.
# +
# Download the data (saves to ~/.keras/datasets)
filename = 'HeLa_S3.npz'
test_size = 0.2 # % of data saved as test
seed = 0 # seed for random train-test split
(X_train, y_train), (X_test, y_test) = deepcell.datasets.hela_s3.load_data(filename, test_size=test_size, seed=seed)
print('X.shape: {}\ny.shape: {}'.format(X_train.shape, y_train.shape))
# -
# ### Set up filepath constants
# +
# the path to the data file is currently required for `train_model_()` functions
# NOTE: Change DATA_DIR if you are not using `deepcell.datasets`
DATA_DIR = os.path.expanduser(os.path.join('~', '.keras', 'datasets'))
DATA_FILE = os.path.join(DATA_DIR, filename)
# confirm the data file is available
assert os.path.isfile(DATA_FILE)
# +
# Set up other required filepaths
# If the data file is in a subdirectory, mirror it in MODEL_DIR and LOG_DIR
PREFIX = os.path.relpath(os.path.dirname(DATA_FILE), DATA_DIR)
ROOT_DIR = '/data' # TODO: Change this! Usually a mounted volume
MODEL_DIR = os.path.abspath(os.path.join(ROOT_DIR, 'models', PREFIX))
LOG_DIR = os.path.abspath(os.path.join(ROOT_DIR, 'logs', PREFIX))
# create directories if they do not exist
for d in (MODEL_DIR, LOG_DIR):
try:
os.makedirs(d)
except OSError as exc: # Guard against race condition
if exc.errno != errno.EEXIST:
raise
# -
# ## Create the RetinaMask Model
#
# Here we instantiate a `RetinaMask` model from `deepcell.model_zoo` for object detection and masking.
# +
from deepcell.utils.retinanet_anchor_utils import get_anchor_parameters
# Generate backbone information from the data
backbone_levels, pyramid_levels, anchor_params = get_anchor_parameters(y_train.astype('int'))
num_classes = 1 # "object" is the only class
# +
from deepcell import model_zoo
backbone = 'resnet50'
# training model is `retinanet` while prediction model is `retinanet_bbox`
model = model_zoo.RetinaMask(
backbone=backbone,
input_shape=X_train.shape[1:],
panoptic=True,
num_semantic_heads=2,
num_semantic_classes=[1, 1],
class_specific_filter=False,
num_classes=num_classes,
backbone_levels=backbone_levels,
pyramid_levels=pyramid_levels,
anchor_params=anchor_params)
# -
# ## Prepare for training
#
# ### Set up training parameters.
#
# There are a number of tunable hyper parameters necessary for training deep learning models:
#
# **model_name**: Incorporated into any files generated during the training process.
#
# **n_epoch**: The number of complete passes through the training dataset.
#
# **lr**: The learning rate determines the speed at which the model learns. Specifically it controls the relative size of the updates to model values after each batch.
#
# **optimizer**: The TensorFlow module [tf.keras.optimizers](https://www.tensorflow.org/api_docs/python/tf/keras/optimizers) offers optimizers with a variety of algorithm implementations. DeepCell typically uses the Adam or the SGD optimizers.
#
# **lr_sched**: A learning rate scheduler allows the learning rate to adapt over the course of model training. Typically a larger learning rate is preferred during the start of the training process, while a small learning rate allows for fine-tuning during the end of training.
#
# **batch_size**: The batch size determines the number of samples that are processed before the model is updated. The value must be greater than one and less than or equal to the number of samples in the training dataset.
#
# **min_objects**: The minimum number of objects in every training image. Images with fewer than `min_objects` objects will be discarded.
# +
from tensorflow.keras.optimizers import SGD, Adam
from deepcell.utils.train_utils import rate_scheduler
model_name = '{}_retinanet'.format(backbone)
n_epoch = 5 # Number of training epochs
lr = 1e-5
optimizer = Adam(lr=lr, clipnorm=0.001)
lr_sched = rate_scheduler(lr=lr, decay=0.99)
batch_size = 1
min_objects = 3
transforms = ['inner-distance', 'outer-distance']
transforms_kwargs = {}
# -
# ### Create the DataGenerators
#
# The `RetinaNetDataGenerator` outputs a raw image (`X`) with bounding boxes and classifications for every object in the labeled data (`y`).
# +
from deepcell.image_generators import RetinaNetGenerator
# this will do preprocessing and realtime data augmentation
datagen = RetinaNetGenerator(
rotation_range=180,
zoom_range=(.8, 1.2),
horizontal_flip=True,
vertical_flip=True)
datagen_val = RetinaNetGenerator()
# +
from deepcell.utils.retinanet_anchor_utils import guess_shapes
train_data = datagen.flow(
{'X': X_train, 'y': y_train},
seed=seed,
include_bbox=True,
include_masks=True,
panoptic=True,
transforms=transforms,
transforms_kwargs=transforms_kwargs,
pyramid_levels=pyramid_levels,
min_objects=min_objects,
anchor_params=anchor_params,
compute_shapes=guess_shapes,
batch_size=batch_size)
val_data = datagen_val.flow(
{'X': X_test, 'y': y_test},
seed=seed,
include_bbox=True,
include_masks=True,
panoptic=True,
transforms=transforms,
transforms_kwargs=transforms_kwargs,
pyramid_levels=pyramid_levels,
min_objects=min_objects,
anchor_params=anchor_params,
compute_shapes=guess_shapes,
batch_size=batch_size)
# -
# ### Compile the model with a loss function
#
# Each output is trained with it's own loss function, regression loss is used for the bounding box task while a different loss is used for the categorical task. Both of these are encapsulated by the `RetinaNetLosses` class.
# +
from deepcell.losses import RetinaNetLosses
retinanet_losses = RetinaNetLosses(
sigma=3.0, alpha=0.25, gamma=2.0,
mask_size=(28, 28), iou_threshold=0.5)
panoptic_weight = 0.1
def semantic_loss(n_classes):
def _semantic_loss(y_pred, y_true):
if n_classes > 1:
return panoptic_weight * losses.weighted_categorical_crossentropy(
y_true, y_pred, n_classes=n_classes)
return panoptic_weight * tf.keras.losses.MSE(y_true, y_pred)
return _semantic_loss
loss = {
'regression': retinanet_losses.regress_loss,
'classification': retinanet_losses.classification_loss,
'masks': retinanet_losses.mask_loss
}
# Give losses for all of the semantic heads
for layer in model.layers:
if layer.name.startswith('semantic'):
n_classes = layer.output_shape[-1]
loss[layer.name] = semantic_loss(num_classes)
# -
model.compile(loss=loss, optimizer=optimizer)
# ## Train the RetinaMask model
#
# Create a prediction model, convert the data generators to `Datasets`, and call `fit()` on the compiled model, along with a default set of callbacks.
# +
from tensorflow.python.data import Dataset
# Some versions of TensorFlow 2.x do not handle multiple
# outputs in X or y, but is compatible with a Dataset instead.
input_type_dict = {'input': tf.float32, 'boxes_input': tf.float32}
input_shape_dict = {
'input': tuple([None] + list(train_data.x.shape[1:])),
'boxes_input': (None, None, 4)
}
output_type_dict = {
'regression': tf.float32,
'classification': tf.float32,
'masks': tf.float32,
}
output_shape_dict = {
'regression': (None, None, None),
'classification': (None, None, None),
'masks': (None, None, None),
}
train_dataset = Dataset.from_generator(
lambda: train_data,
(input_type_dict, output_type_dict),
output_shapes=(input_shape_dict, output_shape_dict))
val_dataset = Dataset.from_generator(
lambda: val_data,
(input_type_dict, output_type_dict),
output_shapes=(input_shape_dict, output_shape_dict))
for i, n in enumerate(range(len(transforms))):
output_type_dict['semantic_{}'.format(i)] = tf.float32
output_shape_dict['semantic_{}'.format(i)] = (None, None, None, n)
# +
from deepcell.model_zoo.retinamask import retinamask_bbox
prediction_model = retinamask_bbox(
model,
nms=True,
num_semantic_heads=len(transforms),
panoptic=True,
anchor_params=anchor_params,
class_specific_filter=False)
# +
from deepcell.utils.train_utils import get_callbacks
from deepcell.utils.train_utils import count_gpus
from deepcell.callbacks import RedirectModel, Evaluate
model_path = os.path.join(MODEL_DIR, '{}.h5'.format(model_name))
num_gpus = count_gpus()
print('Training on', num_gpus, 'GPUs.')
train_callbacks = get_callbacks(
model_path,
lr_sched=lr_sched,
save_weights_only=num_gpus >= 2,
monitor='val_loss',
verbose=1)
eval_callback = RedirectModel(
Evaluate(val_data,
iou_threshold=0.5,
score_threshold=.01,
max_detections=100,
weighted_average=True),
prediction_model)
loss_history = model.fit(
train_data,
steps_per_epoch=train_data.y.shape[0] // batch_size,
epochs=n_epoch,
validation_data=val_data,
validation_steps=val_data.y.shape[0] // batch_size,
callbacks=train_callbacks)
# -
# ## Predict on test data
#
# Use the trained model to predict on new data and post-process the results into a label mask.
# +
# Set up the prediction model
from deepcell import model_zoo
# Set up the prediction model
prediction_model = model_zoo.retinamask_bbox(
model,
nms=True,
anchor_params=anchor_params,
num_semantic_heads=2,
panoptic=True,
class_specific_filter=False)
# +
import matplotlib.pyplot as plt
import os
import time
import numpy as np
from skimage.transform import resize
from skimage.measure import label
from skimage.exposure import equalize_hist, rescale_intensity
from deepcell_toolbox import retinamask_semantic_postprocess
from deepcell.utils.plot_utils import draw_detections, draw_masks
index = np.random.randint(low=0, high=X_test.shape[0])
print('Image Number:', index)
image, mask = X_test[index:index + 1], y_test[index:index + 1]
results = prediction_model.predict(image)
image = 0.01 * np.tile(np.expand_dims(image[0, ..., 0], axis=-1), (1, 1, 3))
mask = np.squeeze(mask)
boxes = results[-6]
scores = results[-5]
labels = results[-4]
masks = results[-3]
semantic = results[-2:]
label_image = retinamask_semantic_postprocess(results[:-1])
# # copy to draw on
draw = image.copy()
# draw the masks
draw_masks(draw, boxes[0], scores[0], masks[0], score_threshold=0.5)
display_image = image.copy()
display_image = rescale_intensity(display_image, out_range=(-1, 1))
fig, axes = plt.subplots(ncols=3, nrows=2, figsize=(20, 20), sharex=True, sharey=True)
ax = axes.ravel()
ax[0].imshow(display_image[..., -1], cmap='jet')
ax[0].set_title('Source Image')
ax[1].imshow(mask, cmap='jet')
ax[1].set_title('Ground Truth Labels')
ax[2].imshow(draw, cmap='jet')
ax[2].set_title('Detections')
ax[3].imshow(semantic[0][0, ..., 0], cmap='jet')
ax[3].set_title('Inner Distance')
ax[4].imshow(semantic[1][0, ..., 0], cmap='jet')
ax[4].set_title('Outer Distance')
ax[5].imshow(label_image[0], cmap='jet')
ax[5].set_title('Label Image')
plt.tight_layout()
plt.show()
# -
#
| notebooks/training/retinanets/PanOpticFPN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/apmoore1/target_aspect_unique/blob/master/texts_contain_the_same_aspect_more_than_once.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="HtPBz5ZkwQiQ" colab_type="code" colab={}
# %%capture
# !pip install git+git://github.com/apmoore1/target-extraction.git@v0.0.2#egg=target_extraction
# + [markdown] id="PaxhVGPry8we" colab_type="text"
# # SemEval 2015 and 2016 datasets do the texts contain the same aspect more than once?
#
# The title is the question we are going to answer here. We are only going to look at sentences that contain (target, aspect, sentiment) where aspect can also be an (entity, aspect) pair. The reason why an aspect or (entity, aspect) pair can occur more than once in a text is because it is attached to a target that is within the text. Thus there could be the following case:
#
# ```
# The CPU memory is great, but the RAM is not
# ```
#
# Given this text we could have the following annotations (memory, MEMORY, positive) and (RAM, MEMORY, negative) where the following represent (target, aspect, sentiment). Thus in this case the MEMORY aspect has occured twice in the same text. The reason we want to see if this occurs or not in the SemEval datasets is to see if we could treat the target sentiment problem as just a text/sentence level aspect based sentiment analysis task. Further whether when trying to identify aspects whether this has to be done at the target level or if it could be done at the text/sentence level. As shown by this example if it was treated a text level aspect based sentiment problem it would have to get one of the two samples wrong as the aspect occurs twice with two different sentiments.
#
# To analysis this we first need to load the datasets. The datasets that have the following annotations (target, aspect, sentiment) are:
# 1. [Restaurant SemEval 2015 Train.](http://alt.qcri.org/semeval2015/task12/index.php?id=data-and-tools)
# 2. [Restaurant SemEval 2015 Test.](http://alt.qcri.org/semeval2015/task12/index.php?id=data-and-tools)
# 3. [Restaurant SemEval 2016 Train.](http://alt.qcri.org/semeval2016/task5/index.php?id=data-and-tools)
# 4. [Restaurant SemEval 2016 Test.](http://alt.qcri.org/semeval2016/task5/index.php?id=data-and-tools)
#
# There are more SemEval datasets from the 2016 edition that contian this annotation format. However for now we are only going to look at these datasets.
# + id="RIwiYZEExsIw" colab_type="code" colab={}
from collections import Counter
from pathlib import Path
from google.colab import files
from target_extraction.dataset_parsers import semeval_2016
from target_extraction.data_types import TargetTextCollection
# SemEval 2014 Laptop and Restaurant
semeval_dataset_fp = {'Restaurant Train 2015': Path('ABSA-15_Restaurants_Train_Final.xml'),
'Restaurant Test 2015': Path('ABSA15_Restaurants_Test.xml'),
'Restaurant Train 2016': Path('ABSA16_Restaurants_Train_SB1_v2.xml'),
'Restaurant Test 2016': Path('EN_REST_SB1_TEST_2016.xml')}
semeval_dataset = {}
for dataset_name, fp in semeval_dataset_fp.items():
if not fp.exists():
print(f'Upload {dataset_name}')
files.upload()
semeval_dataset[dataset_name] = semeval_2016(fp, conflict=True)
# + id="QSjLIO9A2eXq" colab_type="code" colab={}
# SemEval Restaurant
restaurant_train = semeval_dataset['Restaurant Train 2015']
restaurant_test = semeval_dataset['Restaurant Test 2015']
restaurant_combined_2015 = TargetTextCollection.combine(restaurant_train,
restaurant_test)
restaurant_combined_2015.name = 'Restaurant 2015'
restaurant_train = semeval_dataset['Restaurant Train 2016']
restaurant_test = semeval_dataset['Restaurant Test 2016']
restaurant_combined_2016 = TargetTextCollection.combine(restaurant_train,
restaurant_test)
restaurant_combined_2016.name = 'Restaurant 2016'
# + [markdown] id="5AUUa25Mv31y" colab_type="text"
# To confirm that we have the data correct we shall check that the total number of aspects/categories for the datasets matches those that are in the papers:
# 1. For Restaurant 2015 we should have 2499 according to [Pontiki et al. 2015](https://www.aclweb.org/anthology/S15-2082.pdf)
# 2. For Restaurant 2016 we should have 3366 according to [Pontiki et al. 2016](https://www.aclweb.org/anthology/S16-1002.pdf)
# + id="X4b8mS9EvX_d" colab_type="code" outputId="d612ff6c-c0a0-44ee-8715-a97ed9ee136c" colab={"base_uri": "https://localhost:8080/", "height": 51}
print(f'Number of aspects in Restaurant 2015 '
f'{restaurant_combined_2015.number_categories()}')
print(f'Number of aspects in Restaurant 2016 '
f'{restaurant_combined_2016.number_categories()}')
# + [markdown] id="qX15s2ijwRL_" colab_type="text"
# It would appear from above that we have parsed that dataset correctly as we match the number of aspects in the original paper.
#
# We now move on to see how many sentences and the number of aspects that are affected if we treat the task as a text level aspect task instead of taking into the target:
# + id="MhHX2T6Nwd9u" colab_type="code" outputId="74731c79-b8f8-4554-ad64-21431c679159" colab={"base_uri": "https://localhost:8080/", "height": 119}
def aspects_affected(dataset: TargetTextCollection) -> None:
number_texts_wrong = 0
number_wrong = 0
number_text_wrong_diff = 0
number_diff_wrong = 0
for key, value in dataset.items():
aspects_in_text = value['categories']
all_aspect_sentiments = value['target_sentiments']
if aspects_in_text is None:
assert value['targets'] is None
continue
number_aspects = len(aspects_in_text)
aspect_count = Counter(aspects_in_text)
aspect_count_diff = number_aspects - len(aspect_count)
if aspect_count_diff != 0:
number_texts_wrong += 1
number_wrong += aspect_count_diff
# This is a more fine grained analysis that looks at if the aspects that
# do occur more than once in a text, occur more than once with a
# different sentiment in the text. As if they all have the same sentiment
# to some degree you can just get anyway with not duplicating the aspect
# even though the aspects should be attached to a target.
number_text_wrong_diff_bool = False
aspects_occur_more_than_once = [aspect for aspect, count in aspect_count.items()
if count > 1]
for aspect in aspects_occur_more_than_once:
aspect_sentiments = set()
for index, sentiment in enumerate(all_aspect_sentiments):
if aspects_in_text[index] == aspect:
aspect_sentiments.add(sentiment)
if len(aspect_sentiments) > 1:
number_text_wrong_diff_bool = True
number_diff_wrong += aspect_count[aspect]
if number_text_wrong_diff_bool:
number_text_wrong_diff += 1
#print(aspects_occur_more_than_once)
#print(aspect_count)
number_samples = dataset.number_categories()
percent_wrong = round((number_wrong / float(number_samples)) * 100, 2)
percent_text_wrong = round((number_texts_wrong / float(number_samples)) * 100, 2)
percent_diff_wrong = round((number_diff_wrong / float(number_samples)) * 100, 2)
percent_diff_text_wrong = round((number_text_wrong_diff / float(number_samples)) * 100, 2)
print(f'For the dataset {dataset.name} which contains {number_samples} '
f'samples and {len(dataset)} texts\n{number_texts_wrong}'
f'({percent_text_wrong}%) texts and {number_wrong}'
f'({percent_wrong}%) samples are affected.\n{number_text_wrong_diff}'
f'({percent_diff_text_wrong}%) texts and {number_diff_wrong}'
f'({percent_diff_wrong}%) samples are affected with respect to having a '
'different sentiment.')
aspects_affected(restaurant_combined_2015)
aspects_affected(restaurant_combined_2016)
# + [markdown] id="Q4epKq_57AR6" colab_type="text"
# As we can see from the two datasets it shows that up to 11% of samples are affected and up to 8% of texts are affected, whereby the texts can contain more than one of the same aspect. However only up to 4% of samples and up to 2% of texts contain more than one of the same aspect, but where those same aspects have different sentiments like the example given at the start of the notebook on the aspect MEMORY. This shows that in the majority of cases when an aspect does occur more than once for the same text it does so with the same sentiment, thus allowing you to some degree to get anyway with using aspect based text level methods and ignoring the target. However by doing so you will not be able to accurately classify up to 4% of samples that do contain texts with multiple of the same aspects with different sentiments.
#
# # How many implicit targets are there?
#
# In these datasets some of the targets are implicit. This can be found when the target value is `None` while the aspect exisits. An example of this can be seen below:
# + id="gpiak8WC8DGX" colab_type="code" outputId="4919d789-6b6d-41bf-faa1-69ec0de496e0" colab={"base_uri": "https://localhost:8080/", "height": 136}
for key, value in restaurant_combined_2015['P#3:3'].items():
print(f'{key}: {value}')
# + [markdown] id="1XsFOYFn8kZL" colab_type="text"
# This particular example shows how difficult it would be to identify these implicit examples as the aspect already appears explictly in the text. However we are here to answer the question of how many implicit targets there are:
# + id="CuRt8S-H85-W" colab_type="code" outputId="01d91c00-777f-4f7a-8d3a-6e2d12eae311" colab={"base_uri": "https://localhost:8080/", "height": 85}
def implicit_targets(dataset: TargetTextCollection) -> None:
number_implicit_texts = 0
number_implicit_targets = 0
for key, value in dataset.items():
targets = value['targets']
aspects = value['categories']
if targets is None:
continue
text_is_implicit = False
for target, aspect in zip(targets, aspects):
if target is None:
assert not aspect is None
number_implicit_targets += 1
text_is_implicit = True
if text_is_implicit:
number_implicit_texts += 1
number_samples = dataset.number_categories()
percent_texts = round((number_implicit_texts / float(number_samples)) * 100, 2)
percent_targets = round((number_implicit_targets / float(number_samples)) * 100, 2)
print(f'For the dataset {dataset.name} which contains {number_samples} '
f'samples and {len(dataset)} texts\n{number_implicit_texts}'
f'({percent_texts}%) texts and {number_implicit_targets}'
f'({percent_targets}%) samples are implicit.')
implicit_targets(restaurant_combined_2015)
implicit_targets(restaurant_combined_2016)
# + [markdown] id="UVihSbCj_2DT" colab_type="text"
# We can see that at least 25% of the samples contain implicit sentiment!
| texts_contain_the_same_aspect_more_than_once.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="JHijuspuuMkO"
from google.colab import drive
drive.mount('/content/drive')
# + id="0u6oSdNWHhpm"
# !pip install emoji
# + id="8N_7mKSDF2X0"
# !pip install ekphrasis
# + id="aZRAL3V1w2w_"
pip install plotly==4.5.4
# + id="k1BgltZfpfIb"
# !pip install transformers==4.2.1
# + id="uR494XZ0FZ5h"
import numpy as np
import pandas as pd
import string
from nltk.corpus import stopwords
import re
import os
from ekphrasis.classes.preprocessor import TextPreProcessor
from ekphrasis.classes.tokenizer import SocialTokenizer
from ekphrasis.dicts.emoticons import emoticons
# + id="ZjOCTC2LFZ5p"
text_processor = TextPreProcessor(
# terms that will be normalized
normalize=['url', 'email', 'percent', 'money', 'phone', 'user',
'time', 'url', 'date', 'number'],
# terms that will be annotated
annotate={"hashtag", "allcaps", "elongated", "repeated",
'emphasis', 'censored'},
fix_html=True, # fix HTML tokens
# corpus from which the word statistics are going to be used
# for word segmentation
segmenter="twitter",
# corpus from which the word statistics are going to be used
# for spell correction
corrector="twitter",
unpack_hashtags=True, # perform word segmentation on hashtags
unpack_contractions=True, # Unpack contractions (can't -> can not)
spell_correct_elong=True, # spell correction for elongated words
# select a tokenizer. You can use SocialTokenizer, or pass your own
# the tokenizer, should take as input a string and return a list of tokens
tokenizer=SocialTokenizer(lowercase=True).tokenize,
# list of dictionaries, for replacing tokens extracted from the text,
# with other expressions. You can pass more than one dictionaries.
dicts=[emoticons]
)
# + id="BBrkiHZsFZ5v"
def print_text(texts,i,j):
for u in range(i,j):
print(texts[u])
print()
# + id="mC2SiOlMFZ51"
df = pd.read_csv('/content/drive/My Drive/offenseval/olid-training-v1.0.tsv',delimiter='\t',encoding='utf-8')
print(list(df.columns.values)) #file header
print(df.head(5)) #last N rows
# + id="n5uO1nfcyhw7"
df.replace(np.NaN, 'NA', inplace=True)
# + id="QDSHL-Uz13Aj"
df.head(5)
# + id="UEVYS5LfFZ57"
text_array = df["tweet"]
labels = df["subtask_a"]
labels_target = df["subtask_b"]
print(len(text_array))
print_text(text_array,0,10)
# + id="IiJRyIwRiTEi"
original = text_array
# + id="GbgC7u_4jIO7"
from collections import Counter
# + id="wvsUCosR5ck9"
df_test_labels_b = pd.read_csv('/content/drive/My Drive/offenseval/labels-levelb.csv', header=None)
print(len(df_test_labels_b))
lol = df_test_labels_b[1]
print(Counter(lol))
df_test_labels_b.head(5)
# + id="Gr6FcNLp8QzI"
labels_target_test = []
# + id="TNkBXvuXFZ6B"
df_test_text = pd.read_csv('/content/drive/My Drive/offenseval/testset-levela.tsv',delimiter='\t',encoding='utf-8')
print(list(df_test_text.columns.values)) #file header
print(df_test_text.head(5)) #first N rows
df_test_labels = pd.read_csv('/content/drive/My Drive/offenseval/labels-levela.csv', header=None)
print(list(df_test_labels.columns.values))
print(df_test_labels.head(5))
count = 0
j = 0
for i in range(0,len(df_test_text["id"])):
if df_test_labels[1][i] == "OFF":
if df_test_labels[0][i] == df_test_labels_b[0][j]:
labels_target_test.append(df_test_labels_b[1][j])
j = j + 1
else:
labels_target_test.append("NA")
print(len(df_test_text["id"]))
print(count)
text_array_test = df_test_text["tweet"]
labels_test = df_test_labels[1]
print("Checking length of validation set")
print(len(text_array_test),len(labels_test))
# + id="a6jRCqVmQX4J"
original_test = text_array_test
# + id="-bSNIGHn7ZNl"
Counter(labels_target_test)
# + id="wkOC5ONKFZ6I"
#removing website names
def remove_website(text):
return " ".join([word if re.search("r'https?://\S+|www\.\S+'|((?i).com$|.co|.net)",word)==None else "" for word in text.split(" ") ])
# Training set
text_array = text_array.apply(lambda text: remove_website(text))
print_text(text_array,0,10)
print("**************************************************************************")
# Validation set
text_array_test = text_array_test.apply(lambda text: remove_website(text))
print_text(text_array_test,0,10)
# + id="rKBHJaEkFZ6N"
# Functions for chat word conversion
f = open("/content/drive/My Drive/offenseval/slang.txt", "r")
chat_words_str = f.read()
chat_words_map_dict = {}
chat_words_list = []
for line in chat_words_str.split("\n"):
if line != "":
cw = line.split("=")[0]
cw_expanded = line.split("=")[1]
chat_words_list.append(cw)
chat_words_map_dict[cw] = cw_expanded
chat_words_list = set(chat_words_list)
def chat_words_conversion(text):
new_text = []
for w in text.split():
if w.upper() in chat_words_list:
new_text.append(chat_words_map_dict[w.upper()])
else:
new_text.append(w)
return " ".join(new_text)
# + id="t8bYKRrNFZ6R"
# Chat word conversion
# Training set
text_array = text_array.apply(lambda text: chat_words_conversion(text))
print_text(text_array,0,10)
print_text(original,0,10)
print("********************************************************************************")
# Validation set
text_array_test = text_array_test.apply(lambda text: chat_words_conversion(text))
print_text(text_array_test,0,10)
# + id="NK9YMXntG4p_"
os.chdir("/content/drive/My Drive/offenseval")
print(os.getcwd())
# + id="tAVp4vfVFZ6W"
#Function for emoticon conversion
from emoticons import EMOTICONS
def convert_emoticons(text):
for emot in EMOTICONS:
text = re.sub(u'('+emot+')', " ".join(EMOTICONS[emot].replace(",","").split()), text)
return text
#testing the emoticon function
text = "Hello :-) :-)"
text = convert_emoticons(text)
print(text + "\n")
# + id="hdkDKvx9FZ6c"
# Emoticon conversion
# Training set
text_array = text_array.apply(lambda text: convert_emoticons(text))
print_text(text_array,0,10)
print("**********************************************************************************")
# Validation set
text_array_test = text_array_test.apply(lambda text: convert_emoticons(text))
print_text(text_array_test,0,10)
# + id="19UZfUjlFZ6h"
# FUnction for removal of emoji
import emoji
def convert_emojis(text):
text = emoji.demojize(text, delimiters=(" ", " "))
text = re.sub("_|-"," ",text)
return text
# Training set
text_array = text_array.apply(lambda text: convert_emojis(text))
print_text(text_array,0,10)
print("**************************************************************************")
# Validation set
text_array_test = text_array_test.apply(lambda text: convert_emojis(text))
print_text(text_array_test,0,10)
# + id="jn8WX-8FHbjC"
os.chdir("/content")
print(os.getcwd())
# + id="HZITistjFZ6m"
# Ekphrasis pipe for text pre-processing
def ekphrasis_pipe(sentence):
cleaned_sentence = " ".join(text_processor.pre_process_doc(sentence))
return cleaned_sentence
# Training set
text_array = text_array.apply(lambda text: ekphrasis_pipe(text))
print("Training set completed.......")
#Validation set
text_array_test = text_array_test.apply(lambda text: ekphrasis_pipe(text))
print("Test set completed.......")
# + id="w4LXWqIKFZ6q"
print_text(text_array,0,10)
print("************************************************************************")
print_text(text_array_test,0,10)
# + id="f9WO_7JIFZ6v"
# Removing unnecessary punctuations
PUNCT_TO_REMOVE = "\"$%&'()+,-./;=[\]^_`{|}~"
def remove_punctuation(text):
return text.translate(str.maketrans('', '', PUNCT_TO_REMOVE))
# Training set
text_array = text_array.apply(lambda text: remove_punctuation(text))
print_text(text_array,0,10)
print("********************************************************************")
# Validation set
text_array_test = text_array_test.apply(lambda text: remove_punctuation(text))
print_text(text_array_test,0,10)
# + id="uQmxE8_s1Bbk"
# print_text(text_array,3550,3555)
print_text(original,9540,9555)
# + id="dsNw3V7MFZ62"
# Finding length of longest array
maxLen = len(max(text_array,key = lambda text: len(text.split(" "))).split(" "))
print(maxLen)
# + id="PXlK2jraFZ68"
u = lambda text: len(text.split(" "))
sentence_lengths = []
for x in text_array:
sentence_lengths.append(u(x))
print(sorted(sentence_lengths)[-100:])
print(len(sentence_lengths))
# + [markdown] id="7q94LkYyFZ7A"
# <h2>Text pre-processing complete</h2>
# + id="L6iCqNnyFZ7X"
# Count of each label in dataset
from collections import Counter
# Printing training set counts for analysis
print("Elements: ",set(labels))
print("Length: ",len(labels))
print(Counter(labels))
print("**************************************************************************")
# Printing validation set counts for analysis
print("Elements: ",set(labels_test))
print("Length: ",len(labels_test))
print(Counter(labels_test))
# + id="005HnK3xFZ7b"
Y = []
Y_test = []
# Training set
for i in range(0,len(labels)):
if(labels[i] == "OFF"):
Y.append(0)
if(labels[i] == "NOT"):
Y.append(1)
# Validation set
for i in range(0,len(labels_test)):
if(labels_test[i] == "OFF"):
Y_test.append(0)
if(labels_test[i] == "NOT"):
Y_test.append(1)
# + id="BGOgKNrEBqXs"
Y_target = []
Y_target_test = []
# Training set
for i in range(0,len(labels_target)):
if(labels_target[i] == "NA"):
Y_target.append(0)
if(labels_target[i] == "TIN"):
Y_target.append(1)
if(labels_target[i] == "UNT"):
Y_target.append(2)
# Validation set
for i in range(0,len(labels_target_test)):
if(labels_target_test[i] == "NA"):
Y_target_test.append(0)
if(labels_target_test[i] == "TIN"):
Y_target_test.append(1)
if(labels_target_test[i] == "UNT"):
Y_target_test.append(2)
# + id="nXy6Wxh2FZ7f"
# Testing the conversion into integers
for i in range(200,210):
print(text_array_test[i])
print(labels_test[i],Y_test[i])
print(labels_target_test[i],Y_target_test[i])
print()
# + id="k-95uJgsFZ7k"
# Verifying train set
X = np.asarray(list(text_array))
Y = np.asarray(list(Y))
Y_target = np.asarray(list(Y_target))
print(type(X))
print(type(Y))
print(type(Y_target))
print(np.shape(X),np.shape(Y),np.shape(Y_target))
# Verifying validation set
X_test = np.asarray(list(text_array_test))
Y_test = np.asarray(list(Y_test))
Y_target_test = np.asarray(list(Y_target_test))
print(type(X_test))
print(type(Y_test))
print(type(Y_target_test))
print(np.shape(X_test),np.shape(Y_test),np.shape(Y_target_test))
# + id="Se9u5rOU4DTf"
print(Counter(Y))
print(Counter(Y_test))
# + id="hRqxpkqDnRrY"
print(X_test[0])
print(Y_test[0])
print(labels_test[0])
print(Y_target_test[0])
print(labels_target_test[0])
# + [markdown] id="TZvFBMRRFZ7p"
# <h2>Shuffling training and validation data</h2>
# + id="t9P3ZmhZFZ7r"
from sklearn.utils import shuffle
from sklearn.model_selection import train_test_split
# + id="y6-U6ElWFZ7y"
print(Counter(labels))
print(Counter(labels_test))
print(Counter(labels_target))
print(Counter(labels_target_test))
# + id="4EQXXOb-FZ73"
# Converting to one hot vectors
def convert_to_one_hot(Y, C):
Y = np.eye(C)[Y.reshape(-1)] #u[Y] helps to index each element of Y index at u. U here is a class array
return Y
# + id="WydeObXQFZ77"
Y_oh_train = convert_to_one_hot(np.array(Y), C = 2)
Y_oh_test = convert_to_one_hot(np.array(Y_test), C = 2)
Y_oh_target_train = convert_to_one_hot(np.array(Y_target), C = 3)
Y_oh_target_test = convert_to_one_hot(np.array(Y_target_test), C = 3)
print(np.shape(Y_oh_train))
print(np.shape(Y_oh_target_test))
index = 0
print(labels[index], Y[index], "is converted into one hot", Y_oh_train[index])
print(labels_target[index], Y_target[index], "is converted into one hot", Y_oh_target_train[index])
# + [markdown] id="Vk9YydcsFZ8C"
# <h2>Model using ALBERT</h2>
# + id="J-Lxn5eGvDAE"
import tensorflow as tf
import os
import numpy as np
import pandas as pd
import string
from nltk.corpus import stopwords
import re
import os
from collections import Counter
# + id="w35qjBz-vEo3"
from transformers import RobertaTokenizerFast, TFRobertaModel, TFBertModel, BertTokenizerFast, ElectraTokenizerFast, TFElectraModel, AlbertTokenizerFast, TFAlbertModel, XLNetTokenizerFast, TFXLNetModel, MPNetTokenizerFast, TFMPNetModel
from tensorflow import keras
from tensorflow.keras import layers
from tensorflow.keras import backend as K
from tensorflow.keras.callbacks import ModelCheckpoint
from tensorflow.keras.layers import RepeatVector, Concatenate, Dense, Activation, Dot, BatchNormalization, Dropout
from sklearn.metrics import classification_report
from sklearn.metrics import f1_score
# + id="d97gRRE9y383"
print(tf.__version__)
# + id="JSTFXg6DYo8O"
resolver = tf.distribute.cluster_resolver.TPUClusterResolver(tpu='grpc://' + os.environ['COLAB_TPU_ADDR'])
tf.config.experimental_connect_to_cluster(resolver)
tf.tpu.experimental.initialize_tpu_system(resolver)
print("All devices: ", tf.config.list_logical_devices('TPU'))
# + id="4Za2kQSGvVWI"
tokenizer = AlbertTokenizerFast.from_pretrained('albert-large-v2')
# + id="DL_GaDaavdNk"
X = list(X)
X_test = list(X_test)
# + id="l2vEqLJDzNnE"
model_train_x, model_val_x, Y_train, Y_val = train_test_split(X, Y, test_size=0.05, random_state=44)
# + id="ce2tYBEpvdJo"
train_encodings = tokenizer(model_train_x, max_length=100, truncation=True, padding="max_length", return_tensors='tf')
val_encodings = tokenizer(model_val_x, max_length=100, truncation=True, padding="max_length", return_tensors='tf')
test_encodings = tokenizer(X_test, max_length=100, truncation=True, padding="max_length", return_tensors='tf')
# + id="gE2O9UySazhE"
cluster_encodings = tokenizer(X, max_length=100, truncation=True, padding="max_length", return_tensors='tf')
# + id="L0VMHbInvcVi"
print(np.shape(train_encodings["input_ids"]))
print(np.shape(val_encodings["input_ids"]))
print(np.shape(test_encodings["input_ids"]))
print(np.shape(cluster_encodings["input_ids"]))
# + id="nYs7gYzsvcOa"
print(train_encodings["input_ids"][0])
print("***************************************************************************")
print(val_encodings["input_ids"][0])
print("***************************************************************************")
print(test_encodings["input_ids"][0])
# + [markdown] id="wdKykEUBmu5M"
# <h3> Subtask A</h3>
# + id="CRJHpbxaFZ86"
def Offense_classifier(input_shape):
"""
Function creating the model's graph.
Arguments:
input_shape -- shape of the input,(max_len,)
word_to_vec_map -- dictionary mapping every word in a vocabulary into its 100-dimensional vector representation
word_to_index -- dictionary mapping from words to their indices in the vocabulary (13 million words)
Returns:
model -- a model instance in Keras
"""
model = TFAlbertModel.from_pretrained('albert-large-v2')
layer = model.layers[0]
# Define sentence_indices as the input of the graph, it should be of shape input_shape and dtype 'int32' (as it contains indices).
inputs = keras.Input(shape=input_shape, dtype='int32')
input_masks = keras.Input(shape=input_shape, dtype='int32')
embeddings = layer([inputs, input_masks])[1]
X = BatchNormalization(momentum=0.99, epsilon=0.001, center=True, scale=True)(embeddings)
# Add dropout with a probability of 0.1
X = Dropout(0.1)(X)
X = Dense(128,activation='elu',kernel_regularizer=keras.regularizers.l2(0.001))(X)
X = Dense(32,activation='elu',kernel_regularizer=keras.regularizers.l2(0.001))(X)
X = Dense(3,activation='elu',kernel_regularizer=keras.regularizers.l2(0.01))(X)
X = Dense(32,activation='elu',kernel_regularizer=keras.regularizers.l2(0.001))(X)
X = BatchNormalization(momentum=0.99, epsilon=0.001, center=True, scale=True)(X)
X = Dense(128,activation='elu',kernel_regularizer=keras.regularizers.l2(0.001))(X)
X = Dense(1,activation='linear',kernel_regularizer=keras.regularizers.l2(0.01))(X)
# Add a sigmoid activation
X = Activation('sigmoid')(X)
# Create Model instance which converts sentence_indices into X.
model = keras.Model(inputs=[inputs,input_masks], outputs=[X])
return model
# + id="MoCY8kjG0DC5"
strategy = tf.distribute.TPUStrategy(resolver)
# + id="BLnuscUj0IjO"
class EvaluationMetric(keras.callbacks.Callback):
def __init__(self, trial_encodings, trial_masks, Y_test):
super(EvaluationMetric, self).__init__()
self.trial_encodings = trial_encodings
self.trial_masks = trial_masks
self.Y_test = Y_test
def on_epoch_begin(self, epoch, logs={}):
print("\nTraining...")
def on_epoch_end(self, epoch, logs={}):
print("\nEvaluating...")
trial_prediction = self.model.predict([self.trial_encodings,self.trial_masks])
pred = []
for i in range(0,len(self.Y_test)):
num = trial_prediction[i]
if(num > 0.5):
num = 1
else:
num = 0
pred.append(num)
from sklearn.metrics import classification_report
print(classification_report(Y_test, pred, digits=3))
evaluation_metric = EvaluationMetric(test_encodings["input_ids"], test_encodings["attention_mask"], Y_test)
# + id="nCOUVfDy0IYr"
with strategy.scope():
model = Offense_classifier((100,))
optimizer = keras.optimizers.Adam(learning_rate=1e-5)
loss_fun = [
tf.keras.losses.BinaryCrossentropy()
]
metric = ['acc']
model.compile(optimizer=optimizer, loss=loss_fun, metrics=metric)
# + id="kQdfWk222Act"
model.summary()
# + id="rM223PhsK5Ir"
neg, pos = np.bincount(Y)
total = neg + pos
print('Examples:\n Total: {}\n Positive: {} ({:.2f}% of total)\n'.format(
total, pos, 100 * pos / total))
# + id="mCtur56mLoep"
class_weight = {}
maxi = max(neg, pos)
weight_for_0 = (maxi / (maxi + neg))
weight_for_1 = (maxi / (maxi + pos))
class_weight = {0: weight_for_0, 1: weight_for_1}
print('Weight for class 0: {:.2f}'.format(weight_for_0))
print('Weight for class 1: {:.2f}'.format(weight_for_1))
# + id="6mEBrm9NFZ9W"
from keras.callbacks import ModelCheckpoint
checkpoint = ModelCheckpoint(filepath='/content/olid_albert_val.{epoch:03d}.h5',
monitor='val_acc',
verbose=1,
save_weights_only=True,
period=1)
# + id="m5OEmpHyzDQ-"
print(Counter(Y))
print(Counter(Y_test))
# + id="6_Ejap9-cMoj"
print(Counter(Y_train))
print(Counter(Y_val))
# + id="1Otd9w3JbVbq"
print(len(train_encodings["input_ids"]),len(val_encodings["input_ids"]))
# + id="7IfquNEaFZ9i"
# val 0.05
history = model.fit(
x = [train_encodings["input_ids"], train_encodings["attention_mask"]],
y = Y_train,
validation_data = ([val_encodings["input_ids"],val_encodings["attention_mask"]],Y_val),
callbacks = [evaluation_metric, checkpoint],
batch_size = 64,
shuffle=True,
epochs=6,
class_weight = class_weight
)
# + [markdown] id="-C9gJOp-3pK6"
# <h4>Training Curves</h4>
# + id="4EzL7l_M1HXA"
history = history
import matplotlib.pyplot as plt
plt.plot(history.history['acc'])
plt.plot(history.history['val_acc'])
plt.title('Model accuracy')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(['Train', 'Validation'], loc='lower right')
plt.show()
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('Model loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train', 'Validation'], loc='lower right')
plt.show()
# + id="Dhp7jUyPWVf5"
# model.load_weights("/content/drive/MyDrive/OLID Transformer weights/olid_albert(0.05).005.h5")
# + id="qQ06iVLk6Y4V"
# model.save_weights("/content/drive/MyDrive/OLID Transformer weights/olid_albert(0.05).005.h5")
# + [markdown] id="IlcmHgY2QL8u"
# <h4>Test Set Statistics</h4>
# + id="dlkXuAvTFZ9z"
answer = model.predict([test_encodings["input_ids"], test_encodings["attention_mask"]])
# + id="yAh-EgxOS4I1"
pred = []
sample = df_test_text["tweet"]
count = 0
for i in range(0,len(X_test)):
num = answer[i]
if(num >= 0.5):
num = 1
else:
num = 0
pred.append(num)
print(count)
# + id="AqVOLWrFkhnv"
con_mat = tf.math.confusion_matrix(labels=Y_test, predictions=pred)
print(con_mat)
# + id="zeT_iT6HvKF2"
import seaborn as sns
import matplotlib.pyplot as plt
# + id="5YRvE7emtL9M"
figure = plt.figure(figsize=(8, 8))
sns.set(font_scale=1.75)
sns.heatmap(con_mat, annot=True,cmap=plt.cm.viridis,fmt='d', xticklabels=["Offensive","Not Offensive"], yticklabels=["Offensive","Not Offensive"],annot_kws={"size": 15})
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
plt.show()
# + id="2jj5ISVyu6Mc"
from sklearn.metrics import f1_score, classification_report
# + id="hznYcG3LtVhx"
f1_score(Y_test, pred, average='macro')
# + id="Hi9Q-xbecQRM"
print(classification_report(Y_test, pred, target_names=["offensive", "not offensive"], digits=3))
# + [markdown] id="wVY5iNvkS5aB"
# <h3>Train set analysis</h3>
# + id="Nudw40bYS5Co"
answer_train = model.predict([cluster_encodings["input_ids"], cluster_encodings["attention_mask"]])
# + id="LJ8p8hcOTIoE"
pred = []
sample = original
count = 0
for i in range(0,len(Y)):
num = answer_train[i]
lol = num
if(num > 0.5):
num = 1
else:
num = 0
pred.append(num)
if(num != Y[i] and Y[i] == 0 and lol >=0.8):
print(answer_train[i])
print("Original label: ",labels[i])
print("Without pre-processing: ",sample[i])
print("With pre-processing: ",X[i])
lol = ""
count += 1
if(num == 0):
lol = "Offensive"
if(num == 1):
lol = "Not Offensive"
print("Predicted: " + lol)
print()
print(count)
# + [markdown] id="wDHYC0jVktN0"
# <h3>Training examination</h3>
# + id="D_JXxlJUIE4I"
import plotly
import plotly.graph_objs as go
import plotly.express as px
# + id="Jx8jYvYCng6n"
# 3 neuron output
model.layers[-6].name
# + id="LJS7gXgJFCCa"
cluster_dense_3 = keras.Model(inputs=model.input, outputs=model.layers[-6].output)
with strategy.scope():
cluster_3 = cluster_dense_3.predict([cluster_encodings["input_ids"], cluster_encodings["attention_mask"]])
# + id="zn_QbPYsk9es"
pred_train = []
temp = 0
for i in range(0,len(Y)):
num = answer_train[i]
if(num >= 0.5):
num = 1
else:
num = 0
pred_train.append(num)
print(temp)
# + id="eeHnrAsbpq0b"
flag = []
count = 0
x_ = []
y_ = []
z_ = []
for i in range(0,len(Y)):
count = count + 1
x_.append(cluster_3[i][1])
y_.append(cluster_3[i][0])
z_.append(cluster_3[i][2])
if( answer_train[i] > 0.3 and answer_train[i] < 0.7 ):
flag.append(2)
if( answer_train[i] > 0 and answer_train[i] <= 0.3 ):
flag.append(0)
if( answer_train[i] >= 0.7 and answer_train[i] < 1 ):
flag.append(1)
# + id="aodbtdXNvHhz"
Counter(flag)
# + id="G5W6RP9YlCnx"
con_mat = tf.math.confusion_matrix(labels=Y, predictions=pred_train)
print(con_mat)
# + id="2Xr1InDUIS7_"
pred_colour = []
for i in range(0,len(flag)):
if flag[i] == 2:
pred_colour.append("Neutral")
if flag[i] == 1:
pred_colour.append("Not Offensive")
if flag[i] == 0:
pred_colour.append("Offensive")
test_df = pd.DataFrame({'x':x_, 'y':y_, 'z':z_, 'Labels':pred_colour})
fig = px.scatter_3d(test_df, x='x', y='y', z='z', color='Labels')
fig.update_traces(
marker={
'size': 1,
'opacity': 0.7,
'colorscale' : 'Oryel',
}
)
fig.update_layout(legend= {'itemsizing': 'constant', 'font_size':18}, font_size=15, scene_aspectmode='cube')
fig.update_layout(width = 850, height = 750)
fig.update_layout(margin=dict(l=0, r=0, b=0, t=0))
# + id="7MBmam1bzqK_"
pred_colour = []
for i in range(0,len(flag)):
if pred_train[i] == 1:
pred_colour.append("Not Offensive")
if pred_train[i] == 0:
pred_colour.append("Offensive")
test_df = pd.DataFrame({'X':x_, 'Y':y_, 'Z':z_, 'Labels:':pred_colour})
fig = px.scatter_3d(test_df, x='X', y='Y', z='Z', color='Labels:')
fig.update_traces(
marker={
'size': 1,
'opacity': 1,
'colorscale' : 'rainbow',
}
)
fig.update_layout(legend= {'itemsizing': 'constant'}, font_size=14, scene_aspectmode='cube')
fig.update_layout(width = 850, height = 750)
fig.update_layout(margin=dict(l=0, r=0, b=0, t=0))
# + [markdown] id="NvADyeWtkyCa"
# <h3>Traning examination end</h3>
# + [markdown] id="gPUPHuPieZk7"
# <h1>CLUSTERING</h1>
# + [markdown] id="hCoFfK-90oxq"
# <h3>PLM layer</h3>
# + id="JZTddCJb0oJD"
model.layers[-8].name
# + id="Pk32a2tk0oFC"
cluster_bert = keras.Model(inputs=model.input, outputs=model.layers[-8].output)
with strategy.scope():
cl_bert = cluster_bert.predict([cluster_encodings["input_ids"], cluster_encodings["attention_mask"]])
# + id="5r7-V0NqLGqd"
len(cl_bert)
# + id="HVtIKDOf0oBM"
flag_bert = []
count = 0
x_bert = []
y_bert = []
z_bert = []
for i in range(0,len(Y)):
count = count + 1
x_bert.append(cl_bert[i][0])
y_bert.append(cl_bert[i][1])
z_bert.append(cl_bert[i][2])
if( answer_train[i] > 0.28 and answer_train[i] < 0.8 ):
flag_bert.append(2)
if( answer_train[i] > 0 and answer_train[i] <= 0.28 ):
flag_bert.append(0)
if( answer_train[i] >= 0.8 and answer_train[i] < 1 ):
flag_bert.append(1)
print(count)
# + [markdown] id="QcuTHL6oADGT"
# <p>k-means PLM layer</p>
# + id="xgg7vxiCfX9C"
from sklearn.cluster import KMeans
# + id="rDh5fAcH0n9H"
kmeans_bert = KMeans(n_clusters=3, random_state=44).fit(cl_bert)
y_kmeans_bert = kmeans_bert.predict(cl_bert)
# + id="S-RuV1hD0n4w"
Counter(y_kmeans_bert)
# + id="KB90wC8s0n0Y"
Counter(flag_bert)
# + id="7n1QV3RZ0nvi"
# 1 index values are offensive
# 0 index values are not offensive
# 2 index values are neutral
count = 0
for i in range(0,len(flag_bert)):
if flag_bert[i] == 1 and y_kmeans_bert[i] == 1:
count = count + 1
print(count)
# + id="CkPj1SQh2xwF"
for i in range(0,len(flag_bert)):
if(y_kmeans_bert[i] == 0):
y_kmeans_bert[i] = 2
elif(y_kmeans_bert[i] == 1):
y_kmeans_bert[i] = 1
else:
y_kmeans_bert[i] = 0
# + id="veXnzAlUMw7R"
flag_bert = []
count = 0
x_bert = []
y_bert = []
z_bert = []
for i in range(0,len(Y)):
count = count + 1
x_bert.append(cl_bert[i][0])
y_bert.append(cl_bert[i][1])
z_bert.append(cl_bert[i][2])
if( answer_train[i] > 0.3 and answer_train[i] < 0.7 ):
flag_bert.append(2)
if( answer_train[i] > 0 and answer_train[i] <= 0.3 ):
flag_bert.append(0)
if( answer_train[i] >= 0.7 and answer_train[i] < 1 ):
flag_bert.append(1)
print(count)
# + id="yshzveBsM4Fm"
Counter(flag_bert)
# + id="yUCI6rBn2xka"
con_mat = tf.math.confusion_matrix(labels=flag_bert, predictions=y_kmeans_bert)
print(con_mat)
# + id="ctpCXXhOo8Cf"
import sklearn
print(sklearn.metrics.classification_report(flag_bert, y_kmeans_bert, output_dict=False, digits=3))
# + id="cCwmwVNe2xVm"
from sklearn.metrics.pairwise import cosine_similarity
from scipy.spatial.distance import cosine
# + id="-88zP4sK2xHi"
centers_bert = kmeans_bert.cluster_centers_
# + id="cq0TowJm0nex"
svns_off = []
for i in range(0,len(Y_test)):
off = cosine(cl_bert[i], centers_bert[2])/2
svns_off.append(1-off)
print(len(svns_off))
# + id="oRWKQ8RI4Ral"
svns_noff = []
for i in range(0,len(Y_test)):
noff = cosine(cl_bert[i], centers_bert[1])/2
svns_noff.append(1-noff)
print(len(svns_noff))
# + id="eKo_zWN74RWB"
svns_neu = []
for i in range(0,len(Y_test)):
neu = cosine(cl_bert[i], centers_bert[0])/2
svns_neu.append(1-neu)
print(len(svns_neu))
# + [markdown] id="xakK3QUgNM8G"
# <h5>k-means PLM PLot</h5>
# + id="U6gKpMTzZYLu"
import plotly
import plotly.graph_objs as go
import plotly.express as px
# + id="cgdXokQMXnFF"
pred_colour = []
for i in range(0,len(y_kmeans_bert)):
if y_kmeans_bert[i] == 2:
pred_colour.append("Neutral")
if y_kmeans_bert[i] == 1:
pred_colour.append("Not Offensive")
if y_kmeans_bert[i] == 0:
pred_colour.append("Offensive")
test_df = pd.DataFrame({'SVNS Offensive':svns_off, 'SVNS Not Offensive':svns_noff, 'SVNS Neutral':svns_neu, 'Labels:':pred_colour})
fig = px.scatter_3d(test_df, x='SVNS Offensive', y='SVNS Not Offensive', z='SVNS Neutral', color='Labels:')
fig.update_traces(
marker={
'size': 1,
'opacity': 1,
'colorscale' : 'viridis',
},
)
fig.update_layout(legend= {'itemsizing': 'constant'}, font_size=14, scene_aspectmode='cube')
fig.update_layout(width = 850, height = 750)
fig.update_layout(margin=dict(l=0, r=0, b=0, t=0))
# + id="RT9ZDinc0FyH"
pred_kalbert = []
for i in range(0,len(Y_test)):
if(svns_off[i] > svns_noff[i]):
pred_kalbert.append(0)
else:
pred_kalbert.append(1)
print(classification_report(Y_test, pred_kalbert, output_dict=False, digits=3))
# + id="uCymB9LL0FqL"
con_mat = tf.math.confusion_matrix(labels=Y_test, predictions=pred_kalbert)
print(con_mat)
# + [markdown] id="lkQoSZi6exAc"
# <p> GMM model </p>
# + id="3Ty06IQHewe7"
from sklearn.mixture import GaussianMixture
# + id="L4PkC0bPfAB9"
gmm_bert = GaussianMixture(n_components=3, random_state = 44).fit(cl_bert)
# + id="r87Jsg3oe_UH"
mean_bert = gmm_bert.means_
cov_bert = gmm_bert.covariances_
print(np.shape(mean_bert))
print(np.shape(cov_bert))
# + id="KakHZ0KYe_QM"
labels_bert = gmm_bert.predict(cl_bert)
# + id="NWqCpdnbKLPt"
flag_bert = []
count = 0
x_bert = []
y_bert = []
z_bert = []
for i in range(0,len(X)):
count = count + 1
x_bert.append(cl_bert[i][0])
y_bert.append(cl_bert[i][1])
z_bert.append(cl_bert[i][2])
if( answer_train[i] > 0.28 and answer_train[i] < 0.8 ):
flag_bert.append(2)
if( answer_train[i] > 0 and answer_train[i] <= 0.28 ):
flag_bert.append(0)
if( answer_train[i] >= 0.8 and answer_train[i] < 1 ):
flag_bert.append(1)
print(count)
# + id="q3sXMOuJKefR"
Counter(flag_bert)
# + id="tqRelHjIhQhq"
# 1 index values are offensive
# 0 index values are not offensive
# 2 index values are neutral
count = 0
for i in range(0,len(flag_bert)):
if flag_bert[i] == 1 and labels_bert[i] == 1:
count = count + 1
print(count)
# + id="IvseXiiahQD0"
for i in range(0,len(flag_bert)):
if(labels_bert[i] == 0):
labels_bert[i] = 2
elif(labels_bert[i] == 1):
labels_bert[i] = 1
else:
labels_bert[i] = 0
# + id="RuN6XZY9Kuys"
flag_bert = []
count = 0
x_bert = []
y_bert = []
z_bert = []
for i in range(0,len(X)):
count = count + 1
x_bert.append(cl_bert[i][0])
y_bert.append(cl_bert[i][1])
z_bert.append(cl_bert[i][2])
if( answer_train[i] > 0.3 and answer_train[i] < 0.7 ):
flag_bert.append(2)
if( answer_train[i] > 0 and answer_train[i] <= 0.3 ):
flag_bert.append(0)
if( answer_train[i] >= 0.7 and answer_train[i] < 1 ):
flag_bert.append(1)
print(count)
# + id="ZWmgNWKahjLk"
con_mat = tf.math.confusion_matrix(labels=flag_bert, predictions=labels_bert)
print(con_mat)
# + id="3od2lBfD3Z41"
import sklearn
print(sklearn.metrics.classification_report(flag_bert, labels_bert, output_dict=False, digits=3))
# + id="05ZEaAn-Ftq8"
prob_bert = gmm_bert.predict_proba(cl_bert)
prob_bert = prob_bert.T
# + [markdown] id="sq43uk25N8-Y"
# <h5>GMM PLM Plot</h5>
# + id="fITkqpNDaNtR"
pred_colour = []
for i in range(0,len(y_kmeans_bert)):
if labels_bert[i] == 2:
pred_colour.append("Neutral")
if labels_bert[i] == 1:
pred_colour.append("Not Offensive")
if labels_bert[i] == 0:
pred_colour.append("Offensive")
test_df = pd.DataFrame({'SVNS Offensive':prob_bert[2], 'SVNS Non Offensive':prob_bert[1], 'SVNS Neutral':prob_bert[0], 'Labels:':pred_colour})
fig = px.scatter_3d(test_df, x='SVNS Offensive', y='SVNS Non Offensive', z='SVNS Neutral', color='Labels:')
fig.update_traces(
marker={
'size': 1.8,
'opacity': 1,
'colorscale' : 'viridis',
}
)
fig.update_layout(legend= {'itemsizing': 'constant'}, font_size=14, scene_aspectmode='cube')
fig.update_layout(width = 950, height = 750)
fig.update_layout(margin=dict(l=0, r=0, b=0, t=0))
# + [markdown] id="HpYTb160z1N4"
# <h3>Dense 3 layer</h3>
# + id="gsnmLYS8IKwY"
from sklearn.preprocessing import normalize
# + id="-gdI2YnUypsL"
cl_norm = normalize(cluster_3, norm='l2', axis=1)
# + id="2leCxqUczL9r"
flag_3 = []
count = 0
x_ = []
y_ = []
z_ = []
for i in range(0,len(X)):
count = count + 1
x_.append(cl_norm[i][0])
y_.append(cl_norm[i][1])
z_.append(cl_norm[i][2])
if( answer_train[i] > 0.28 and answer_train[i] < 0.8 ):
flag_3.append(2)
if( answer_train[i] > 0 and answer_train[i] <= 0.28 ):
flag_3.append(0)
if( answer_train[i] >= 0.8 and answer_train[i] < 1 ):
flag_3.append(1)
print(count)
# + [markdown] id="MntcgtMyAYZN"
# <p>k-means Dense 3</p>
# + id="jF5bqOYoM9nb"
kmeans_3 = KMeans(n_clusters=3, random_state=44).fit(cl_norm)
y_kmeans_3 = kmeans_3.predict(cl_norm)
# + id="42ugjvzrPOhp"
Counter(y_kmeans_3)
# + id="EGNkI4XGSJeq"
Counter(flag_3)
# + id="G3QrGYwJPXkh"
# 1 index values are offensive
# 0 index values are not offensive
# 2 index values are neutral
count = 0
for i in range(0,len(flag_3)):
if flag_3[i] == 1 and y_kmeans_3[i] == 1:
count = count + 1
print(count)
# + id="Uq2TyIP3PYcM"
for i in range(0,len(flag_3)):
if(y_kmeans_3[i] == 0):
y_kmeans_3[i] = 2
elif(y_kmeans_3[i] == 1):
y_kmeans_3[i] = 1
else:
y_kmeans_3[i] = 0
# + id="XKSpqPoiPeSG"
flag_3 = []
count = 0
x_ = []
y_ = []
z_ = []
for i in range(0,len(X)):
count = count + 1
x_.append(cl_norm[i][2])
y_.append(cl_norm[i][1])
z_.append(cl_norm[i][0])
if( answer_train[i] > 0.3 and answer_train[i] < 0.7 ):
flag_3.append(2)
if( answer_train[i] > 0 and answer_train[i] <= 0.3 ):
flag_3.append(0)
if( answer_train[i] >= 0.7 and answer_train[i] < 1 ):
flag_3.append(1)
print(count)
# + id="74XZjQHoow4J"
Counter(flag_3)
# + id="7YMxovgxcgLU"
con_mat = tf.math.confusion_matrix(labels=flag_3, predictions=y_kmeans_3)
print(con_mat)
# + id="MBGH7CdfRdFl"
import sklearn
print(sklearn.metrics.classification_report(flag_3, y_kmeans_3, output_dict=False, digits=3))
# + [markdown] id="J3angcCcHEw5"
# <p>Transition phase</p>
# + id="66dJKDnVdvPV"
pred_colour = []
for i in range(0,len(y_kmeans_bert)):
if y_kmeans_3[i] == 2:
pred_colour.append("Neutral")
if y_kmeans_3[i] == 1:
pred_colour.append("Not Offensive")
if y_kmeans_3[i] == 0:
pred_colour.append("Offensive")
test_df = pd.DataFrame({'X':x_, 'Y':y_, 'Z':z_, 'Labels:':pred_colour})
fig = px.scatter_3d(test_df, x='X', y='Y', z='Z', color='Labels:')
fig.update_traces(
marker={
'size': 1,
'opacity': 1,
'colorscale' : 'viridis',
}
)
fig.update_layout(legend= {'itemsizing': 'constant'}, font_size=14, scene_aspectmode='cube')
fig.update_layout(width = 850, height = 750)
fig.update_layout(margin=dict(l=0, r=0, b=0, t=0))
# + [markdown] id="QVkZLf0wHLol"
# <p>Original predictions</p>
# + id="qiNW3HXMfO-i"
pred_colour = []
for i in range(0,len(y_kmeans_bert)):
if pred_train[i] == 1:
pred_colour.append("Not Offensive")
if pred_train[i] == 0:
pred_colour.append("Offensive")
test_df = pd.DataFrame({'X':x_, 'Y':y_, 'Z':z_, 'Labels:':pred_colour})
fig = px.scatter_3d(test_df, x='X', y='Y', z='Z', color='Labels:')
fig.update_traces(
marker={
'size': 1,
'opacity': 1,
'colorscale' : 'rainbow',
}
)
fig.update_layout(legend= {'itemsizing': 'constant'}, font_size=14, scene_aspectmode='cube')
fig.update_layout(width = 850, height = 750)
fig.update_layout(margin=dict(l=0, r=0, b=0, t=0))
# + [markdown] id="R7d1l_ubROD6"
# <h4>End of transition capture</h4>
# + id="cQHolZXSuzK2"
from sklearn.metrics.pairwise import cosine_similarity
from scipy.spatial.distance import cosine
# + id="JSkLQp_7uy6B"
centers_3 = kmeans_3.cluster_centers_
print(centers_3)
# + id="USgTEi4ruyos"
svns_off = []
for i in range(0,len(Y_test)):
off = cosine(cl_norm[i], centers_3[2])/2
svns_off.append(1-off)
print(len(svns_off))
# + id="o7Hxo-QLvt1U"
svns_noff = []
for i in range(0,len(Y_test)):
noff = cosine(cl_norm[i], centers_3[1])/2
svns_noff.append(1-noff)
print(len(svns_noff))
# + id="rF5b8CvBvvGo"
svns_neu = []
for i in range(0,len(Y_test)):
neu = cosine(cl_norm[i], centers_3[0])/2
svns_neu.append(1-neu)
print(len(svns_neu))
# + [markdown] id="XZad5lE_Om-6"
# <h5>kmeans Dense 3 Plot</h5>
# + id="zDi2Kr1EdDoU"
pred_colour = []
for i in range(0,len(y_kmeans_bert)):
if y_kmeans_3[i] == 2:
pred_colour.append("Neutral")
if y_kmeans_3[i] == 1:
pred_colour.append("Not Offensive")
if y_kmeans_3[i] == 0:
pred_colour.append("Offensive")
test_df = pd.DataFrame({'SVNS Offensive':svns_off, 'SVNS Not Offensive':svns_noff, 'SVNS Neutral':svns_neu, 'Labels:':pred_colour})
fig = px.scatter_3d(test_df, x='SVNS Offensive', y='SVNS Not Offensive', z='SVNS Neutral', color='Labels:')
fig.update_traces(
marker={
'size': 1,
'opacity': 1,
'colorscale' : 'viridis',
}
)
fig.update_layout(legend= {'itemsizing': 'constant'}, font_size=14, scene_aspectmode='cube')
fig.update_layout(width = 850, height = 750)
fig.update_layout(margin=dict(l=0, r=0, b=0, t=0))
# + id="7GvWVjw_1YXS"
pred_dalbert = []
for i in range(0,len(Y_test)):
if(svns_off[i] > svns_noff[i]):
pred_dalbert.append(0)
else:
pred_dalbert.append(1)
print(classification_report(Y_test, pred_dalbert, output_dict=False, digits=3))
# + id="Y6hbh_nJ1YNV"
con_mat = tf.math.confusion_matrix(labels=Y_test, predictions=lolol)
print(con_mat)
# + [markdown] id="0UxQLMFmyX_G"
# <p> GMM model </p>
# + id="KJ3TUE0zyXEr"
gmm_3 = GaussianMixture(n_components=3, random_state = 44).fit(cl_norm)
# + id="ebI2i-bryW7A"
mean_norm = gmm_3.means_
cov_norm = gmm_3.covariances_
print(np.shape(mean_norm))
print(np.shape(cov_norm))
# + id="7G4NEJI2yWyI"
labels_norm = gmm_3.predict(cl_norm)
# + id="--NR_8RmWNdY"
flag_3 = []
count = 0
x_ = []
y_ = []
z_ = []
for i in range(0,len(X)):
count = count + 1
x_.append(cl_norm[i][2])
y_.append(cl_norm[i][1])
z_.append(cl_norm[i][0])
if( answer_train[i] > 0.28 and answer_train[i] < 0.8 ):
flag_3.append(2)
if( answer_train[i] > 0 and answer_train[i] <= 0.28 ):
flag_3.append(0)
if( answer_train[i] >= 0.8 and answer_train[i] < 1 ):
flag_3.append(1)
print(count)
# + id="YP-hLHbbSa8R"
Counter(labels_norm)
# + id="aFJc8jcvWRuc"
Counter(flag_3)
# + id="lVx299gsyWqC"
# 1 index values are offensive
# 0 index values are not offensive
# 2 index values are neutral
count = 0
for i in range(0,len(flag_3)):
if flag_3[i] == 2 and labels_norm[i] == 0:
count = count + 1
print(count)
# + id="Bwj7FBuSzEGw"
for i in range(0,len(flag_3)):
if(labels_norm[i] == 0):
labels_norm[i] = 2
elif(labels_norm[i] == 1):
labels_norm[i] = 1
else:
labels_norm[i] = 0
# + id="GPH-CLYbWtmD"
flag_3 = []
count = 0
x_ = []
y_ = []
z_ = []
for i in range(0,len(X)):
count = count + 1
x_.append(cl_norm[i][2])
y_.append(cl_norm[i][1])
z_.append(cl_norm[i][0])
if( answer_train[i] > 0.3 and answer_train[i] < 0.7 ):
flag_3.append(2)
if( answer_train[i] > 0 and answer_train[i] <= 0.3 ):
flag_3.append(0)
if( answer_train[i] >= 0.7 and answer_train[i] < 1 ):
flag_3.append(1)
print(count)
# + id="STLy6RirzECt"
con_mat = tf.math.confusion_matrix(labels=flag_3, predictions=labels_norm)
print(con_mat)
# + id="K04A25tK40Y8"
import sklearn
print(sklearn.metrics.classification_report(flag_3, labels_norm, output_dict=False, digits=3))
# + id="nzerzBvA5yh0"
prob_norm = gmm_3.predict_proba(cl_norm)
prob_norm = prob_norm.T
# + [markdown] id="4PFUDsrmPLb8"
# <h5>GMM Dense 3 Plot</h5>
# + id="Ata8zfmjg6lj"
pred_colour = []
for i in range(0,len(y_kmeans_bert)):
if labels_norm[i] == 2:
pred_colour.append("Neutral")
if labels_norm[i] == 1:
pred_colour.append("Not Offensive")
if labels_norm[i] == 0:
pred_colour.append("Offensive")
test_df = pd.DataFrame({'SVNS Offensive':prob_norm[2], 'SVNS Not Offensive':prob_norm[1], 'SVNS Neutral':prob_norm[0], 'Labels:':pred_colour})
fig = px.scatter_3d(test_df, x='SVNS Offensive', y='SVNS Not Offensive', z='SVNS Neutral', color='Labels:')
fig.update_traces(
marker={
'size': 1.5,
'opacity': 1,
'colorscale' : 'viridis',
}
)
fig.update_layout(legend= {'itemsizing': 'constant'}, font_size=14, scene_aspectmode='cube')
fig.update_layout(width = 850, height = 750)
fig.update_layout(margin=dict(l=0, r=0, b=0, t=0))
# + [markdown] id="LoSh2HkVM-FR"
# <h3>Dense 3 layer end</h3>
# + [markdown] id="I9KReciSt2OQ"
# <h3>Batch Norm layer</h3>
# + id="jy39KUfk4YCu"
model.layers[-4].name
# + id="3T2ar9of0Duo"
cluster_32 = keras.Model(inputs=model.input, outputs=model.layers[-4].output)
with strategy.scope():
cl_32 = cluster_32.predict([test_encodings["input_ids"], test_encodings["attention_mask"]])
# + id="ge2aGeyg4jqM"
flag_32 = []
count = 0
x_32 = []
y_32 = []
z_32 = []
for i in range(0,len(X)):
count = count + 1
x_32.append(cl_32[i][0])
y_32.append(cl_32[i][1])
z_32.append(cl_32[i][2])
if( answer_train[i] > 0.28 and answer_train[i] < 0.8 ):
flag_32.append(2)
if( answer_train[i] > 0 and answer_train[i] <= 0.28 ):
flag_32.append(0)
if( answer_train[i] >= 0.8 and answer_train[i] < 1 ):
flag_32.append(1)
print(count)
# + [markdown] id="QoM1uO00AxUV"
# <p>k-means BatchNorm layer</p>
# + id="6nv_fX296vdI"
kmeans_32 = KMeans(n_clusters=3, random_state=44).fit(cl_32)
y_kmeans_32 = kmeans_32.predict(cl_32)
# + id="UlkhMcNKJejg"
Counter(y_kmeans_32)
# + id="EakhhsbKfjfX"
Counter(flag_32)
# + id="LnXxczDPWVlZ"
# 2 index values are offensive
# 0 index values are not offensive
# 0 index values are neutral
count = 0
for i in range(0,len(y_kmeans_32)):
if flag_32[i] == 1 and y_kmeans_32[i] == 1:
count = count + 1
print(count)
# + id="8yPxtiMV_trW"
for i in range(0,len(y_kmeans_32)):
if(y_kmeans_32[i] == 0):
y_kmeans_32[i] = 0
elif(y_kmeans_32[i] == 1):
y_kmeans_32[i] = 1
else:
y_kmeans_32[i] = 2
# + id="Toe6CE1ilYGj"
flag_32 = []
count = 0
x_32 = []
y_32 = []
z_32 = []
for i in range(0,len(X)):
count = count + 1
x_32.append(cl_32[i][0])
y_32.append(cl_32[i][1])
z_32.append(cl_32[i][2])
if( answer_train[i] > 0.3 and answer_train[i] < 0.7 ):
flag_32.append(2)
if( answer_train[i] > 0 and answer_train[i] <= 0.3 ):
flag_32.append(0)
if( answer_train[i] >= 0.7 and answer_train[i] < 1 ):
flag_32.append(1)
print(count)
# + id="nC4X5_bGfAwo"
con_mat = tf.math.confusion_matrix(labels=flag_32, predictions=y_kmeans_32)
print(con_mat)
# + id="MpGwKo4jqQPO"
import sklearn
print(sklearn.metrics.classification_report(flag_32, y_kmeans_32, output_dict=False, digits=3))
# + id="ZIBWvBFEyyoF"
centers_32 = kmeans_32.cluster_centers_
# + id="8sisL0IvdrTP"
svns_off = []
for i in range(0,len(Y_test)):
off = cosine(cl_32[i], centers_32[0])/2
svns_off.append(1-off)
print(len(svns_off))
# + id="LqDGo4o8k8Ph"
svns_noff = []
for i in range(0,len(Y_test)):
noff = cosine(cl_32[i], centers_32[1])/2
svns_noff.append(1-noff)
print(len(svns_noff))
# + id="o7abIoiRk8CH"
svns_neu = []
for i in range(0,len(Y_test)):
neu = cosine(cl_32[i], centers_32[2])/2
svns_neu.append(1-neu)
print(len(svns_neu))
# + [markdown] id="Hv6x7n4oA39u"
# <p>k-means BatchNorm Plot</p>
# + id="xbsHcP9Jkjnf"
pred_colour = []
for i in range(0,len(y_kmeans_bert)):
if y_kmeans_32[i] == 2:
pred_colour.append("Neutral")
if y_kmeans_32[i] == 1:
pred_colour.append("Not Offensive")
if y_kmeans_32[i] == 0:
pred_colour.append("Offensive")
test_df = pd.DataFrame({'SVNS Offensive':svns_off, 'SVNS Not Offensive':svns_noff, 'SVNS Neutral':svns_neu, 'Labels:':pred_colour})
fig = px.scatter_3d(test_df, x='SVNS Offensive', y='SVNS Not Offensive', z='SVNS Neutral', color='Labels:')
fig.update_traces(
marker={
'size': 1,
'opacity': 1,
'colorscale' : 'viridis',
}
)
fig.update_layout(legend= {'itemsizing': 'constant'}, font_size=14, scene_aspectmode='cube')
fig.update_layout(width = 850, height = 750)
fig.update_layout(margin=dict(l=0, r=0, b=0, t=0))
# + id="ECwsT3mx2aMk"
pred_BNalbert = []
for i in range(0,len(Y_test)):
if(svns_off[i] > svns_noff[i]):
pred_BNalbert.append(0)
else:
pred_BNalbert.append(1)
print(classification_report(Y_test, pred_BNalbert, output_dict=False, digits=3))
# + id="cRBgCmSa2aEJ"
con_mat = tf.math.confusion_matrix(labels=Y_test, predictions=pred_BNalbert)
print(con_mat)
# + [markdown] id="s2xr2iN58eQe"
# <p> GMM Model BatchNorm</p>
# + id="dnfQ9GrSm2Hh"
gmm_32 = GaussianMixture(n_components=3, random_state = 44).fit(cl_32)
# + id="HN1DnstL8hfC"
mean_32 = gmm_32.means_
cov_32 = gmm_32.covariances_
print(np.shape(mean_32))
print(np.shape(cov_32))
# + id="qCCf6p2W8hbh"
labels_32 = gmm_32.predict(cl_32)
# + id="Gq5znocnV2VF"
flag_32 = []
count = 0
x_32 = []
y_32 = []
z_32 = []
for i in range(0,len(X)):
count = count + 1
x_32.append(cl_32[i][0])
y_32.append(cl_32[i][1])
z_32.append(cl_32[i][2])
if( answer_train[i] > 0.28 and answer_train[i] < 0.8 ):
flag_32.append(2)
if( answer_train[i] > 0 and answer_train[i] <= 0.28 ):
flag_32.append(0)
if( answer_train[i] >= 0.8 and answer_train[i] < 1 ):
flag_32.append(1)
print(count)
# + id="V5AgvLFl-4On"
Counter(flag_32)
# + id="6A7HVs_P8hX2"
# 1 index values are offensive
# 0 index values are not offensive
# 2 index values are neutral
count = 0
for i in range(0,len(flag_32)):
if flag_32[i] == 2 and labels_32[i] == 0:
count = count + 1
print(count)
# + id="MjsJFtnd8hT5"
for i in range(0,len(flag_32)):
if(labels_32[i] == 0):
labels_32[i] = 2
elif(labels_32[i] == 1):
labels_32[i] = 1
else:
labels_32[i] = 0
# + id="SFMHwFz5YY8Q"
flag_32 = []
count = 0
x_32 = []
y_32 = []
z_32 = []
for i in range(0,len(X)):
count = count + 1
x_32.append(cl_32[i][0])
y_32.append(cl_32[i][1])
z_32.append(cl_32[i][2])
if( answer_train[i] > 0.3 and answer_train[i] < 0.7 ):
flag_32.append(2)
if( answer_train[i] > 0 and answer_train[i] <= 0.3 ):
flag_32.append(0)
if( answer_train[i] >= 0.7 and answer_train[i] < 1 ):
flag_32.append(1)
print(count)
# + id="_CV6mSbW8hP2"
con_mat = tf.math.confusion_matrix(labels=flag_32, predictions=labels_32)
print(con_mat)
# + id="ZU2xjTjG5i6o"
import sklearn
print(sklearn.metrics.classification_report(flag_32, labels_32, output_dict=False, digits=3))
# + id="7w8LxWjbDpkG"
prob_32 = gmm_32.predict_proba(cl_32)
prob_32 = prob_32.T
# + [markdown] id="SDMEd04XBI73"
# <p>GMM BatchNorm Plot</p>
# + id="H6Nfvvi1lTsO"
pred_colour = []
for i in range(0,len(y_kmeans_bert)):
if labels_32[i] == 2:
pred_colour.append("Neutral")
if labels_32[i] == 1:
pred_colour.append("Not Offensive")
if labels_32[i] == 0:
pred_colour.append("Offensive")
test_df = pd.DataFrame({'SVNS Offensive':prob_32[2], 'SVNS Not Offensive':prob_32[1], 'SVNS Neutral':prob_32[0], 'Labels:':pred_colour})
fig = px.scatter_3d(test_df, x='SVNS Offensive', y='SVNS Not Offensive', z='SVNS Neutral', color='Labels:')
fig.update_traces(
marker={
'size': 1.5,
'opacity': 1,
'colorscale' : 'viridis',
}
)
fig.update_layout(legend= {'itemsizing': 'constant'}, font_size=14, scene_aspectmode='cube')
fig.update_layout(width = 850, height = 750)
fig.update_layout(margin=dict(l=0, r=0, b=0, t=0))
| Models/Albert.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# #%matplotlib tk
# %load_ext autoreload
# %autoreload 2
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:100% !important; }</style>"))
import matplotlib
import matplotlib.pyplot as plt
from matplotlib.path import Path
import os
import numpy as np
import scipy.io as sio
import matplotlib.pyplot as plt
import time
# pytorch for GPU
import torch
import LocaNMF
# +
# allen map folder
mapfolder='/home/cat/code/self_initiated_alex_locaNMF/locaNMF/data/AllenMap/'
# data location
Uc_fname = '/media/cat/1TB/data/yuki/IA2am_Mar9_30Hz_aligned_Uc_500SVD_GPU_registered.npy'
Vc_fname = '/media/cat/1TB/data/yuki/IA2am_Mar9_30Hz_aligned_Vc_500SVD_GPU.npy'
root_dir = os.path.split(Vc_fname)[0]+'/'
# brain mask
fname_mask = '/media/cat/1TB/data/yuki/IA2am_Mar9_30Hz_aligned_Uc_aligned_brainmask.npy'
plt.imshow(np.load(fname_mask))
# user params
minrank = 1; maxrank = 1; # rank = how many components per brain region. Set maxrank to around 10 for regular dataset.
rank_range = (minrank, maxrank, 1)
min_pixels = 100 # minimum number of pixels in Allen map for it to be considered a brain region
loc_thresh = 50 # Localization threshold, i.e. percentage of area restricted to be inside the 'Allen boundary'
r2_thresh = 0.99 # Fraction of variance in the data to capture with LocaNMF
## [OPTIONAL] if cuda support, uncomment following lines
if True:
os.environ["CUDA_DEVICE_ORDER"]="PCI_BUS_ID"
os.environ["CUDA_VISIBLE_DEVICES"]="0"
device='cuda'
else:
# else, if on cpu
device='cpu'
# +
# data = np.load('/media/cat/1TB/data/yuki/IA2am_Mar9_30Hz_aligned_Uc_500SVD_GPU_registered.npy')
# print (data.shape)
# data = np.load('/media/cat/1TB/data/yuki/IA2am_Mar9_30Hz_aligned_Uc_aligned_brainmask.npy')
# print (data.shape)
# data = np.load('/media/cat/1TB/data/yuki/IA2am_Mar9_30Hz_aligned_Vc_500SVD_GPU.npy')
# print (data.shape)
# -
# Load Allen Brain Map
# Get data from this link--> https://www.dropbox.com/s/q4m1v151o06zsdv/Vc_Uc.mat?dl=0
# Get map here if you do not have it--> https://www.dropbox.com/s/d35xt7e6l2ywiol/preprocessed_allenDorsalMap.mat?dl=0
dorsalMapScaled = sio.loadmat(mapfolder+'preprocessed_allenDorsalMap.mat')['dorsalMapScaled'].astype(float)
print (dorsalMapScaled.shape)
plt.imshow(dorsalMapScaled)
plt.show()
# +
# Load Vc and Uc data
# Brainmask limits x and y
new_x, new_y = 580, 540
#V=arrays['Vc']
V=np.load(Vc_fname)
#U=arrays['Uc'][:new_y, :new_x, :]
U=np.load(Uc_fname)[:new_y, :new_x, :]
print ("V : ", V.shape)
print ("U : ", U.shape)
# +
# load Uc spatial components and crop if required
class crop(object):
def __init__(self,data, fname):
self.data = data
self.fname = fname
self.define_ROI()
def define_ROI(self):
#if os.path.exists(self.fname[:-4]+".npy")==False:
roi_coords = self.select_ROI()
return (roi_coords)
def select_ROI(self):
''' Function to crop field-of-view of video
'''
self.fig, self.ax = plt.subplots()
self.coords=[]
self.sample_image = data[1]
print (self.sample_image.shape)
self.ax.imshow(self.sample_image)#, vmin=0.0, vmax=0.02)
#self.ax.set_title(ROI_name)
#figManager = plt.get_current_fig_manager()
#figManager.window.showMaximized()
self.cid = self.fig.canvas.mpl_connect('button_press_event', self.on_click)
plt.show(block=False)
def on_click(self, event):
''' Mouse click function that catches clicks and plots them on top of existing image
'''
if event.inaxes is not None:
print (event.ydata, event.xdata)
self.coords.append((event.ydata, event.xdata))
#for j in range(len(self.coords)):
for k in range(2):
for l in range(2):
self.sample_image[int(event.ydata)-1+k,int(event.xdata)-1+l]=np.max(self.sample_image)
self.ax.imshow(self.sample_image)
self.fig.canvas.draw()
else:
print ('Exiting')
plt.close()
#self.fig.canvas.mpl_disconnect(self.cid)
np.save(self.fname[:-4]+"_coords.npy", self.coords)
return
f_out = 'crop file name'
if False:
if os.path.exists(f_out)==False:
data = np.load(fname).transpose(2,0,1)
#data = np.load('/media/cat/10TB/in_vivo/tim/yuki/IA2/tif_files/IA2am_Mar9_30Hz/IA2am_Mar9_30Hz_aligned_Uc_500SVD_GPU.npy')
print (data.shape)
crop(data, fname)
# +
# load coords and visualized cropped data
#f_out = fname[:-4]+"_brainmask.npy"
if False:
if os.path.exists(f_out)==False:
data = np.load(fname).transpose(2,0,1)
coords = np.load(fname[:-4]+"_coords.npy")
data = data[0]
#Search points outside and black them out:
all_points = []
for i in range(data.shape[0]):
for j in range(data.shape[1]):
all_points.append([i,j])
all_points = np.array(all_points)
vertixes = np.array(coords)
vertixes_path = Path(vertixes)
mask = vertixes_path.contains_points(all_points)
print (mask.shape)
counter=0
coords_save=[]
images_processed = data
print (images_processed.shape)
for i in range(images_processed.shape[0]):
for j in range(images_processed.shape[1]):
if mask[counter] == False:
images_processed[i][j]=np.nan
coords_save.append([i,j])
counter+=1
final_mask = images_processed
np.save(fname[:-4]+"_brainmask.npy", final_mask)
fig, ax = plt.subplots()
ax.imshow(images_processed)
plt.show()
# +
#brainmask=~np.isnan(arrays['brainmask'][:new_y, :new_x])
#brainmask_full=~np.isnan(arrays['brainmask'])
#trueareas=arrays['trueareas'].flatten()
mask = np.load(fname_mask)
brainmask=~np.isnan(mask[:new_y, :new_x])
brainmask_full=~np.isnan(mask)
print ("branimask: ", brainmask.shape)
print ("branimask full: ", brainmask_full.shape)
plt.imshow(brainmask)
plt.show()
# -
# ax=plt.subplot(121)
# plt.imshow(brainmask_full)
# ax=plt.subplot(122)
# plt.imshow(brainmask)
# plt.show()
# Divide up region based Allen map into left and right sides
dorsalMapScaled[:,:int(dorsalMapScaled.shape[1]/2)] = dorsalMapScaled[:,:int(dorsalMapScaled.shape[1]/2)] * -1
dorsalMapScaled = -dorsalMapScaled[:new_y, :new_x]
# +
# Check that data has the correct shapes. V [K_d x T], U [X x Y x K_d], brainmask [X x Y]
if V.shape[0]!=U.shape[-1]: print('Wrong dimensions of U and V!')
print("Rank of video : %d" % V.shape[0]); print("Number of timepoints : %d" % V.shape[1]);
# Plot the maximum U for each pixel
plotmap=np.zeros((dorsalMapScaled.shape)); plotmap.fill(np.nan); plotmap[brainmask]=dorsalMapScaled[brainmask]
plt.imshow(plotmap,cmap='Spectral'); plt.axis('off'); plt.title('Allen region map'); plt.show();
plt.imshow(np.max(U,axis=2)); plt.axis('off'); plt.title('True A'); plt.show()
# -
# Perform the LQ decomposition. Time everything.
t0_global = time.time()
t0 = time.time()
q, r = np.linalg.qr(V.T)
time_ests={'qr_decomp':time.time() - t0}
# # Initialize LocaNMF
# +
# Put in data structure for LocaNMF
video_mats = (np.copy(U[brainmask]), r.T)
del U
# -
# GET LABELS FOR EACH CELL FROM SHREYA
# region_mats[0] = [unique regions x pixels] the mask of each region
# region_mats[1] = [unique regions x pixels] the distance penalty of each region
# region_mats[2] = [unique regions] area code
region_mats = LocaNMF.extract_region_metadata(brainmask,
dorsalMapScaled,
min_size=min_pixels)
# +
# GET LABELS FOR EACH CELL FROM SHREYA
region_metadata = LocaNMF.RegionMetadata(region_mats[0].shape[0],
region_mats[0].shape[1:],
device=device)
region_metadata.set(torch.from_numpy(region_mats[0].astype(np.uint8)),
torch.from_numpy(region_mats[1]),
torch.from_numpy(region_mats[2].astype(np.int64)))
# -
# GET LABELS FOR EACH CELL FROM SHREYA
if device=='cuda': torch.cuda.synchronize()
print('v SVD Initialization')
t0 = time.time()
region_videos = LocaNMF.factor_region_videos(video_mats,
region_mats[0],
rank_range[1],
device=device)
if device=='cuda': torch.cuda.synchronize()
print("\'-total : %f" % (time.time() - t0))
time_ests['svd_init'] = time.time() - t0
low_rank_video = LocaNMF.LowRankVideo(
(int(np.sum(brainmask)),) + video_mats[1].shape, device=device
)
low_rank_video.set(torch.from_numpy(video_mats[0].T),
torch.from_numpy(video_mats[1]))
# # LocaNMF
if device=='cuda': torch.cuda.synchronize()
print('v Rank Line Search')
t0 = time.time()
locanmf_comps = LocaNMF.rank_linesearch(low_rank_video,
region_metadata,
region_videos,
maxiter_rank=maxrank-minrank+1,
maxiter_lambda=500,
maxiter_hals=20,
lambda_step=1.35,
lambda_init=1e-8,
loc_thresh=loc_thresh,
r2_thresh=r2_thresh,
rank_range=rank_range,
verbose=[True, False, False],
sample_prop=(1,1),
device=device
)
if device=='cuda': torch.cuda.synchronize()
print("\'-total : %f" % (time.time() - t0))
time_ests['rank_linesearch'] = time.time() - t0
print("Number of components : %d" % len(locanmf_comps))
# +
# Evaluate R^2
_,r2_fit=LocaNMF.evaluate_fit_to_region(low_rank_video,
locanmf_comps,
region_metadata.support.data.sum(0),
sample_prop=(1, 1))
print("R^2 fit on all data : %f" % r2_fit)
time_ests['global_time'] = time.time()-t0_global
# -
# # Reformat spatial and temporal matrices, plot, and save
# Assigning regions to components
region_ranks = []; region_idx = []
for rdx in torch.unique(locanmf_comps.regions.data, sorted=True):
region_ranks.append(torch.sum(rdx == locanmf_comps.regions.data).item())
region_idx.append(rdx.item())
areas=np.repeat(region_mats[2],region_ranks,axis=0)
# +
# Get LocaNMF spatial and temporal components
A=locanmf_comps.spatial.data.cpu().numpy().T
A_reshape=np.zeros((brainmask_full.shape[0],brainmask_full.shape[1],A.shape[1])); A_reshape.fill(np.nan)
A_reshape[brainmask_full,:]=A
C=np.matmul(q,locanmf_comps.temporal.data.cpu().numpy().T).T
# +
# Plotting all the regions' components
A_validmask=np.zeros((brainmask_full.shape[0],brainmask_full.shape[1])); A_validmask.fill(np.nan)
for rdx, i in zip(region_idx, np.cumsum(region_ranks)-1):
fig, axs = plt.subplots(1 + int((1+region_ranks[rdx]) / 4), 4,
figsize=(16,(1 + int((1+region_ranks[rdx]) / 4)) * 4))
axs = axs.reshape((int(np.prod(axs.shape)),))
A_validmask[brainmask_full] = locanmf_comps.distance.data[i].cpu()==0
axs[0].imshow(A_validmask)
axs[0].set_title("Region: {}".format(rdx+1)); axs[0].axis('off')
# only for simulated data
#A_validmask[brainmask_full]=video_mats[0][:,np.where(areas[i]==trueareas)[0][0]]
#axs[1].imshow(A_validmask)
#axs[1].set_title("True A: {}".format(rdx+1)); axs[1].axis('off')
axs[2].imshow(A_reshape[:,:,i])
axs[2].set_title("LocaNMF A: {}".format(i+1)); axs[2].axis('off')
# axs[3].plot(V[np.where(areas[i]==trueareas)[0][0],:3000].T,'k'); axs[3].plot(C[i,:3000].T,'r');
# axs[3].set_title("True & LocaNMF C: {}".format(i+1));axs[3].axis('off');
# if i==0: axs[3].legend(('True','LocaNMF'))
plt.show()
# -
# Plot the maximum U for each pixel
plotmap=np.zeros((dorsalMapScaled.shape)); plotmap.fill(np.nan); plotmap[brainmask]=dorsalMapScaled[brainmask]
plt.imshow(plotmap,cmap='Spectral'); plt.axis('off'); plt.title('Allen region map'); plt.show();
plt.imshow(np.max(A_reshape,axis=2)); plt.axis('off'); plt.title('True A'); plt.show()
# Plot the distribution of lambdas.
# If lots of values close to the minimum, decrease lambda_init.
# If lots of values close to the maximum, increase maxiter_lambda or lambda_step.
plt.hist(np.log(locanmf_comps.lambdas.data.cpu()), bins=torch.unique(locanmf_comps.lambdas.data).shape[0])
plt.show()
datafolder = os.path.split(fname)[0]+"/"
sio.savemat(datafolder+'locanmf_decomp_loc'+str(loc_thresh)+'.mat',
{'C':C,
'A':A_reshape,
'lambdas':locanmf_comps.lambdas.data.cpu().numpy(),
'areas':areas,
'r2_fit':r2_fit,
'time_ests':time_ests
})
torch.cuda.empty_cache()
# +
# make random data times;
import numpy as np
import matplotlib.pyplot as plt
# load time traces from
import csv
fname = '/media/cat/1TB/data/yuki/dlc_models/yuki-cat-2020-05-11/videos/IA2pm_Apr22_Week2_30HzDLC_resnet50_yukiMay11shuffle1_1030000.csv'
with open(fname, newline='') as csvfile:
data = list(csv.reader(csvfile))
labels = data[1]
print (labels)
traces = np.array(data[3:])
print (traces.shape)
start_idx = 10
lever = np.float32(traces[:,start_idx:start_idx+3])
fig=plt.figure()
plt.plot(lever[:,0], c='blue')
plt.plot(lever[:,1], c='red')
plt.show()
# -
| older/self_initiated_paper/older/NMF_yuki_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# # Bay Area Bike Share Analysis
#
# ## Introduction
#
# > **Tip**: Quoted sections like this will provide helpful instructions on how to navigate and use an iPython notebook.
#
# [Bay Area Bike Share](http://www.bayareabikeshare.com/) is a company that provides on-demand bike rentals for customers in San Francisco, Redwood City, Palo Alto, Mountain View, and San Jose. Users can unlock bikes from a variety of stations throughout each city, and return them to any station within the same city. Users pay for the service either through a yearly subscription or by purchasing 3-day or 24-hour passes. Users can make an unlimited number of trips, with trips under thirty minutes in length having no additional charge; longer trips will incur overtime fees.
#
# In this project, you will put yourself in the shoes of a data analyst performing an exploratory analysis on the data. You will take a look at two of the major parts of the data analysis process: data wrangling and exploratory data analysis. But before you even start looking at data, think about some questions you might want to understand about the bike share data. Consider, for example, if you were working for Bay Area Bike Share: what kinds of information would you want to know about in order to make smarter business decisions? Or you might think about if you were a user of the bike share service. What factors might influence how you would want to use the service?
#
# **Question 1**: Write at least two questions you think could be answered by data.
#
# **Answer**:
# What's the availability of the bikes to the user?
# What's the average duration of usage by the users?
# What's the most popular routes?
# When is the peak usage time of bikes?
#
# > **Tip**: If you double click on this cell, you will see the text change so that all of the formatting is removed. This allows you to edit this block of text. This block of text is written using [Markdown](http://daringfireball.net/projects/markdown/syntax), which is a way to format text using headers, links, italics, and many other options. You will learn more about Markdown later in the Nanodegree Program. Hit **Shift** + **Enter** or **Shift** + **Return**.
# ## Using Visualizations to Communicate Findings in Data
#
# As a data analyst, the ability to effectively communicate findings is a key part of the job. After all, your best analysis is only as good as your ability to communicate it.
#
# In 2014, Bay Area Bike Share held an [Open Data Challenge](http://www.bayareabikeshare.com/datachallenge-2014) to encourage data analysts to create visualizations based on their open data set. You’ll create your own visualizations in this project, but first, take a look at the [submission winner for Best Analysis](http://thfield.github.io/babs/index.html) from <NAME>. Read through the entire report to answer the following question:
#
# **Question 2**: What visualizations do you think provide the most interesting insights? Are you able to answer either of the questions you identified above based on Tyler’s analysis? Why or why not?
#
# **Answer**:
# Where do people bike share, he was able to gather a lot of additional events such as temp, popular sports game, strike into analysing the behavior of the riders.
# Most of it yes, except for how available is the bikes to the riders wasn't answered.
# It would have been useful to see if riders had any issues trying to get a bike during which hour of the day and at which particular station.
# ## Data Wrangling
#
# Now it's time to explore the data for yourself. Year 1 and Year 2 data from the Bay Area Bike Share's [Open Data](http://www.bayareabikeshare.com/open-data) page have already been provided with the project materials; you don't need to download anything extra. The data comes in three parts: the first half of Year 1 (files starting `201402`), the second half of Year 1 (files starting `201408`), and all of Year 2 (files starting `201508`). There are three main datafiles associated with each part: trip data showing information about each trip taken in the system (`*_trip_data.csv`), information about the stations in the system (`*_station_data.csv`), and daily weather data for each city in the system (`*_weather_data.csv`).
#
# When dealing with a lot of data, it can be useful to start by working with only a sample of the data. This way, it will be much easier to check that our data wrangling steps are working since our code will take less time to complete. Once we are satisfied with the way things are working, we can then set things up to work on the dataset as a whole.
#
# Since the bulk of the data is contained in the trip information, we should target looking at a subset of the trip data to help us get our bearings. You'll start by looking at only the first month of the bike trip data, from 2013-08-29 to 2013-09-30. The code below will take the data from the first half of the first year, then write the first month's worth of data to an output file. This code exploits the fact that the data is sorted by date (though it should be noted that the first two days are sorted by trip time, rather than being completely chronological).
#
# First, load all of the packages and functions that you'll be using in your analysis by running the first code cell below. Then, run the second code cell to read a subset of the first trip data file, and write a new file containing just the subset we are initially interested in.
#
# > **Tip**: You can run a code cell like you formatted Markdown cells by clicking on the cell and using the keyboard shortcut **Shift** + **Enter** or **Shift** + **Return**. Alternatively, a code cell can be executed using the **Play** button in the toolbar after selecting it. While the cell is running, you will see an asterisk in the message to the left of the cell, i.e. `In [*]:`. The asterisk will change into a number to show that execution has completed, e.g. `In [1]`. If there is output, it will show up as `Out [1]:`, with an appropriate number to match the "In" number.
# import all necessary packages and functions.
import csv
from datetime import datetime
import numpy as np
import pandas as pd
from babs_datacheck import question_3
from babs_visualizations import usage_stats, usage_plot
from IPython.display import display
# %matplotlib inline
# +
# file locations
file_in = '201402_trip_data.csv'
file_out = '201309_trip_data.csv'
with open(file_out, 'w') as f_out, open(file_in, 'r') as f_in:
# set up csv reader and writer objects
in_reader = csv.reader(f_in)
out_writer = csv.writer(f_out)
# write rows from in-file to out-file until specified date reached
while True:
datarow = next(in_reader)
# trip start dates in 3rd column, m/d/yyyy HH:MM formats
if datarow[2][:9] == '10/1/2013':
break
out_writer.writerow(datarow)
# -
# ### Condensing the Trip Data
#
# The first step is to look at the structure of the dataset to see if there's any data wrangling we should perform. The below cell will read in the sampled data file that you created in the previous cell, and print out the first few rows of the table.
# +
sample_data = pd.read_csv('201309_trip_data.csv')
display(sample_data.head())
# -
# In this exploration, we're going to concentrate on factors in the trip data that affect the number of trips that are taken. Let's focus down on a few selected columns: the trip duration, start time, start terminal, end terminal, and subscription type. Start time will be divided into year, month, and hour components. We will also add a column for the day of the week and abstract the start and end terminal to be the start and end _city_.
#
# Let's tackle the lattermost part of the wrangling process first. Run the below code cell to see how the station information is structured, then observe how the code will create the station-city mapping. Note that the station mapping is set up as a function, `create_station_mapping()`. Since it is possible that more stations are added or dropped over time, this function will allow us to combine the station information across all three parts of our data when we are ready to explore everything.
# +
# Display the first few rows of the station data file.
station_info = pd.read_csv('201402_station_data.csv')
display(station_info.head())
# This function will be called by another function later on to create the mapping.
def create_station_mapping(station_data):
"""
Create a mapping from station IDs to cities, returning the
result as a dictionary.
"""
station_map = {}
for data_file in station_data:
with open(data_file, 'r') as f_in:
# set up csv reader object - note that we are using DictReader, which
# takes the first row of the file as a header row for each row's
# dictionary keys
weather_reader = csv.DictReader(f_in)
for row in weather_reader:
station_map[row['station_id']] = row['landmark']
return station_map
# -
# You can now use the mapping to condense the trip data to the selected columns noted above. This will be performed in the `summarise_data()` function below. As part of this function, the `datetime` module is used to **p**arse the timestamp strings from the original data file as datetime objects (`strptime`), which can then be output in a different string **f**ormat (`strftime`). The parsed objects also have a variety of attributes and methods to quickly obtain
#
# There are two tasks that you will need to complete to finish the `summarise_data()` function. First, you should perform an operation to convert the trip durations from being in terms of seconds to being in terms of minutes. (There are 60 seconds in a minute.) Secondly, you will need to create the columns for the year, month, hour, and day of the week. Take a look at the [documentation for datetime objects in the datetime module](https://docs.python.org/2/library/datetime.html#datetime-objects). **Find the appropriate attributes and method to complete the below code.**
def summarise_data(trip_in, station_data, trip_out):
"""
This function takes trip and station information and outputs a new
data file with a condensed summary of major trip information. The
trip_in and station_data arguments will be lists of data files for
the trip and station information, respectively, while trip_out
specifies the location to which the summarized data will be written.
"""
# generate dictionary of station - city mapping
station_map = create_station_mapping(station_data)
with open(trip_out, 'w') as f_out:
# set up csv writer object
out_colnames = ['duration', 'start_date', 'start_year',
'start_month', 'start_hour', 'weekday',
'start_city', 'end_city', 'subscription_type']
trip_writer = csv.DictWriter(f_out, fieldnames = out_colnames)
trip_writer.writeheader()
for data_file in trip_in:
with open(data_file, 'r') as f_in:
# set up csv reader object
trip_reader = csv.DictReader(f_in)
# collect data from and process each row
for row in trip_reader:
new_point = {}
# convert duration units from seconds to minutes
### Question 3a: Add a mathematical operation below ###
### to convert durations from seconds to minutes. ###
new_point['duration'] = float(row['Duration']) ________
# reformat datestrings into multiple columns
### Question 3b: Fill in the blanks below to generate ###
### the expected time values. ###
trip_date = datetime.strptime(row['Start Date'], '%m/%d/%Y %H:%M')
new_point['start_date'] = trip_date.strftime('%Y-%m-%d')
new_point['start_year'] = trip_date.________
new_point['start_month'] = trip_date.________
new_point['start_hour'] = trip_date.________
new_point['weekday'] = trip_date.________
# remap start and end terminal with start and end city
new_point['start_city'] = station_map[row['Start Terminal']]
new_point['end_city'] = station_map[row['End Terminal']]
# two different column names for subscribers depending on file
if 'Subscription Type' in row:
new_point['subscription_type'] = row['Subscription Type']
else:
new_point['subscription_type'] = row['Subscriber Type']
# write the processed information to the output file.
trip_writer.writerow(new_point)
# **Question 3**: Run the below code block to call the `summarise_data()` function you finished in the above cell. It will take the data contained in the files listed in the `trip_in` and `station_data` variables, and write a new file at the location specified in the `trip_out` variable. If you've performed the data wrangling correctly, the below code block will print out the first few lines of the dataframe and a message verifying that the data point counts are correct.
# +
# Process the data by running the function we wrote above.
station_data = ['201402_station_data.csv']
trip_in = ['201309_trip_data.csv']
trip_out = '201309_trip_summary.csv'
summarise_data(trip_in, station_data, trip_out)
# Load in the data file and print out the first few rows
sample_data = pd.read_csv(trip_out)
display(sample_data.head())
# Verify the dataframe by counting data points matching each of the time features.
question_3(sample_data)
# -
# > **Tip**: If you save a jupyter Notebook, the output from running code blocks will also be saved. However, the state of your workspace will be reset once a new session is started. Make sure that you run all of the necessary code blocks from your previous session to reestablish variables and functions before picking up where you last left off.
#
# ## Exploratory Data Analysis
#
# Now that you have some data saved to a file, let's look at some initial trends in the data. Some code has already been written for you in the `babs_visualizations.py` script to help summarize and visualize the data; this has been imported as the functions `usage_stats()` and `usage_plot()`. In this section we'll walk through some of the things you can do with the functions, and you'll use the functions for yourself in the last part of the project. First, run the following cell to load the data, then use the `usage_stats()` function to see the total number of trips made in the first month of operations, along with some statistics regarding how long trips took.
# +
trip_data = pd.read_csv('201309_trip_summary.csv')
usage_stats(trip_data)
# -
# You should see that there are over 27,000 trips in the first month, and that the average trip duration is larger than the median trip duration (the point where 50% of trips are shorter, and 50% are longer). In fact, the mean is larger than the 75% shortest durations. This will be interesting to look at later on.
#
# Let's start looking at how those trips are divided by subscription type. One easy way to build an intuition about the data is to plot it. We'll use the `usage_plot()` function for this. The second argument of the function allows us to count up the trips across a selected variable, displaying the information in a plot. The expression below will show how many customer and how many subscriber trips were made. Try it out!
usage_plot(trip_data, 'subscription_type')
# Seems like there's about 50% more trips made by subscribers in the first month than customers. Let's try a different variable now. What does the distribution of trip durations look like?
usage_plot(trip_data, 'duration')
# Looks pretty strange, doesn't it? Take a look at the duration values on the x-axis. Most rides are expected to be 30 minutes or less, since there are overage charges for taking extra time in a single trip. The first bar spans durations up to about 1000 minutes, or over 16 hours. Based on the statistics we got out of `usage_stats()`, we should have expected some trips with very long durations that bring the average to be so much higher than the median: the plot shows this in a dramatic, but unhelpful way.
#
# When exploring the data, you will often need to work with visualization function parameters in order to make the data easier to understand. Here's where the third argument of the `usage_plot()` function comes in. Filters can be set for data points as a list of conditions. Let's start by limiting things to trips of less than 60 minutes.
usage_plot(trip_data, 'duration', ['duration < 60'])
# This is looking better! You can see that most trips are indeed less than 30 minutes in length, but there's more that you can do to improve the presentation. Since the minimum duration is not 0, the left hand bar is slighly above 0. We want to be able to tell where there is a clear boundary at 30 minutes, so it will look nicer if we have bin sizes and bin boundaries that correspond to some number of minutes. Fortunately, you can use the optional "boundary" and "bin_width" parameters to adjust the plot. By setting "boundary" to 0, one of the bin edges (in this case the left-most bin) will start at 0 rather than the minimum trip duration. And by setting "bin_width" to 5, each bar will count up data points in five-minute intervals.
usage_plot(trip_data, 'duration', ['duration < 60'], boundary = 0, bin_width = 5)
# **Question 4**: Which five-minute trip duration shows the most number of trips? Approximately how many trips were made in this range?
#
# **Answer**: Replace this text with your response!
# Visual adjustments like this might be small, but they can go a long way in helping you understand the data and convey your findings to others.
#
# ## Performing Your Own Analysis
#
# Now that you've done some exploration on a small sample of the dataset, it's time to go ahead and put together all of the data in a single file and see what trends you can find. The code below will use the same `summarise_data()` function as before to process data. After running the cell below, you'll have processed all the data into a single data file. Note that the function will not display any output while it runs, and this can take a while to complete since you have much more data than the sample you worked with above.
# +
station_data = ['201402_station_data.csv',
'201408_station_data.csv',
'201508_station_data.csv' ]
trip_in = ['201402_trip_data.csv',
'201408_trip_data.csv',
'201508_trip_data.csv' ]
trip_out = 'babs_y1_y2_summary.csv'
# This function will take in the station data and trip data and
# write out a new data file to the name listed above in trip_out.
summarise_data(trip_in, station_data, trip_out)
# -
# Since the `summarise_data()` function has created a standalone file, the above cell will not need to be run a second time, even if you close the notebook and start a new session. You can just load in the dataset and then explore things from there.
trip_data = pd.read_csv('babs_y1_y2_summary.csv')
display(trip_data.head())
# #### Now it's your turn to explore the new dataset with `usage_stats()` and `usage_plot()` and report your findings! Here's a refresher on how to use the `usage_plot()` function:
# - first argument (required): loaded dataframe from which data will be analyzed.
# - second argument (required): variable on which trip counts will be divided.
# - third argument (optional): data filters limiting the data points that will be counted. Filters should be given as a list of conditions, each element should be a string in the following format: `'<field> <op> <value>'` using one of the following operations: >, <, >=, <=, ==, !=. Data points must satisfy all conditions to be counted or visualized. For example, `["duration < 15", "start_city == 'San Francisco'"]` retains only trips that originated in San Francisco and are less than 15 minutes long.
#
# If data is being split on a numeric variable (thus creating a histogram), some additional parameters may be set by keyword.
# - "n_bins" specifies the number of bars in the resultant plot (default is 10).
# - "bin_width" specifies the width of each bar (default divides the range of the data by number of bins). "n_bins" and "bin_width" cannot be used simultaneously.
# - "boundary" specifies where one of the bar edges will be placed; other bar edges will be placed around that value (this may result in an additional bar being plotted). This argument may be used alongside the "n_bins" and "bin_width" arguments.
#
# You can also add some customization to the `usage_stats()` function as well. The second argument of the function can be used to set up filter conditions, just like how they are set up in `usage_plot()`.
usage_stats(trip_data)
usage_plot(trip_data)
# Explore some different variables using the functions above and take note of some trends you find. Feel free to create additional cells if you want to explore the dataset in other ways or multiple ways.
#
# > **Tip**: In order to add additional cells to a notebook, you can use the "Insert Cell Above" and "Insert Cell Below" options from the menu bar above. There is also an icon in the toolbar for adding new cells, with additional icons for moving the cells up and down the document. By default, new cells are of the code type; you can also specify the cell type (e.g. Code or Markdown) of selected cells from the Cell menu or the dropdown in the toolbar.
#
# One you're done with your explorations, copy the two visualizations you found most interesting into the cells below, then answer the following questions with a few sentences describing what you found and why you selected the figures. Make sure that you adjust the number of bins or the bin limits so that they effectively convey data findings. Feel free to supplement this with any additional numbers generated from `usage_stats()` or place multiple visualizations to support your observations.
# Final Plot 1
usage_plot(trip_data)
# **Question 5a**: What is interesting about the above visualization? Why did you select it?
#
# **Answer**: Replace this text with your response!
# Final Plot 2
usage_plot(trip_data)
# **Question 5b**: What is interesting about the above visualization? Why did you select it?
#
# **Answer**: Replace this text with your response!
# ## Conclusions
#
# Congratulations on completing the project! This is only a sampling of the data analysis process: from generating questions, wrangling the data, and to exploring the data. Normally, at this point in the data analysis process, you might want to draw conclusions about our data by performing a statistical test or fitting the data to a model for making predictions. There are also a lot of potential analyses that could be performed on the data which are not possible with only the code given. Instead of just looking at number of trips on the outcome axis, you could see what features affect things like trip duration. We also haven't looked at how the weather data ties into bike usage.
#
# **Question 6**: Think of a topic or field of interest where you would like to be able to apply the techniques of data science. What would you like to be able to learn from your chosen subject?
#
# **Answer**: Replace this text with your response!
#
# > **Tip**: If we want to share the results of our analysis with others, we aren't limited to giving them a copy of the jupyter Notebook (.ipynb) file. We can also export the Notebook output in a form that can be opened even for those without Python installed. From the **File** menu in the upper left, go to the **Download as** submenu. You can then choose a different format that can be viewed more generally, such as HTML (.html) or
# PDF (.pdf). You may need additional packages or software to perform these exports.
| Bay_Area_Bike_Share_Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Self Governing Neural Networks (SGNN): the Projection Layer
#
# > A SGNN's word projections preprocessing pipeline in scikit-learn
#
# In this notebook, we'll use T=80 random hashing projection functions, each of dimensionnality d=14, for a total of 1120 features per projected word in the projection function P.
#
# Next, we'll need feedforward neural network (dense) layers on top of that (as in the paper) to re-encode the projection into something better. This is not done in the current notebook and is left to you to implement in your own neural network to train the dense layers jointly with a learning objective. The SGNN projection created hereby is therefore only a preprocessing on the text to project words into the hashing space, which becomes spase 1120-dimensional word features created dynamically hereby. Only the CountVectorizer needs to be fitted, as it is a char n-gram term frequency prior to the hasher. This one could be computed dynamically too without any fit, as it would be possible to use the [power set](https://en.wikipedia.org/wiki/Power_set) of the possible n-grams as sparse indices computed on the fly as (indices, count_value) tuples, too.
# +
import sklearn
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.pipeline import Pipeline, FeatureUnion
from sklearn.random_projection import SparseRandomProjection
from sklearn.base import BaseEstimator, TransformerMixin
from sklearn.metrics.pairwise import cosine_similarity
from collections import Counter
from pprint import pprint
# -
# ## Preparing dummy data for demonstration:
# +
class SentenceTokenizer(BaseEstimator, TransformerMixin):
# char lengths:
MINIMUM_SENTENCE_LENGTH = 10
MAXIMUM_SENTENCE_LENGTH = 200
def fit(self, X, y=None):
return self
def transform(self, X):
return self._split(X)
def _split(self, string_):
splitted_string = []
sep = chr(29) # special separator character to split sentences or phrases.
string_ = string_.strip().replace(".", "." + sep).replace("?", "?" + sep).replace("!", "!" + sep).replace(";", ";" + sep).replace("\n", "\n" + sep)
for phrase in string_.split(sep):
phrase = phrase.strip()
while len(phrase) > SentenceTokenizer.MAXIMUM_SENTENCE_LENGTH:
# clip too long sentences.
sub_phrase = phrase[:SentenceTokenizer.MAXIMUM_SENTENCE_LENGTH].lstrip()
splitted_string.append(sub_phrase)
phrase = phrase[SentenceTokenizer.MAXIMUM_SENTENCE_LENGTH:].rstrip()
if len(phrase) >= SentenceTokenizer.MINIMUM_SENTENCE_LENGTH:
splitted_string.append(phrase)
return splitted_string
with open("./data/How-to-Grow-Neat-Software-Architecture-out-of-Jupyter-Notebooks.md") as f:
raw_data = f.read()
test_str_tokenized = SentenceTokenizer().fit_transform(raw_data)
# Print text example:
print(len(test_str_tokenized))
pprint(test_str_tokenized[3:9])
# -
# ## Creating a SGNN preprocessing pipeline's classes
class WordTokenizer(BaseEstimator, TransformerMixin):
def fit(self, X, y=None):
return self
def transform(self, X):
begin_of_word = "<"
end_of_word = ">"
out = [
[
begin_of_word + word + end_of_word
for word in sentence.replace("//", " /").replace("/", " /").replace("-", " -").replace(" ", " ").split(" ")
if not len(word) == 0
]
for sentence in X
]
return out
# +
char_ngram_range = (1, 4)
char_term_frequency_params = {
'char_term_frequency__analyzer': 'char',
'char_term_frequency__lowercase': False,
'char_term_frequency__ngram_range': char_ngram_range,
'char_term_frequency__strip_accents': None,
'char_term_frequency__min_df': 2,
'char_term_frequency__max_df': 0.99,
'char_term_frequency__max_features': int(1e7),
}
class CountVectorizer3D(CountVectorizer):
def fit(self, X, y=None):
X_flattened_2D = sum(X.copy(), [])
super(CountVectorizer3D, self).fit_transform(X_flattened_2D, y) # can't simply call "fit"
return self
def transform(self, X):
return [
super(CountVectorizer3D, self).transform(x_2D)
for x_2D in X
]
def fit_transform(self, X, y=None):
return self.fit(X, y).transform(X)
# +
import scipy.sparse as sp
T = 80
d = 14
hashing_feature_union_params = {
# T=80 projections for each of dimension d=14: 80 * 14 = 1120-dimensionnal word projections.
**{'union__sparse_random_projection_hasher_{}__n_components'.format(t): d
for t in range(T)
},
**{'union__sparse_random_projection_hasher_{}__dense_output'.format(t): False # only AFTER hashing.
for t in range(T)
}
}
class FeatureUnion3D(FeatureUnion):
def fit(self, X, y=None):
X_flattened_2D = sp.vstack(X, format='csr')
super(FeatureUnion3D, self).fit(X_flattened_2D, y)
return self
def transform(self, X):
return [
super(FeatureUnion3D, self).transform(x_2D)
for x_2D in X
]
def fit_transform(self, X, y=None):
return self.fit(X, y).transform(X)
# -
# ## Fitting the pipeline
#
# Note: at fit time, the only thing done is to discard some unused char n-grams and to instanciate the random hash, the whole thing could be independent of the data, but here because of discarding the n-grams, we need to "fit" the data. Therefore, fitting could be avoided all along, but we fit here for simplicity of implementation using scikit-learn.
# +
params = dict()
params.update(char_term_frequency_params)
params.update(hashing_feature_union_params)
pipeline = Pipeline([
("word_tokenizer", WordTokenizer()),
("char_term_frequency", CountVectorizer3D()),
('union', FeatureUnion3D([
('sparse_random_projection_hasher_{}'.format(t), SparseRandomProjection())
for t in range(T)
]))
])
pipeline.set_params(**params)
result = pipeline.fit_transform(test_str_tokenized)
print(len(result), len(test_str_tokenized))
print(result[0].shape)
# -
# ## Let's see the output and its form.
# +
print(result[0].toarray().shape)
print(result[0].toarray()[0].tolist())
print("")
# The whole thing is quite discrete:
print(set(result[0].toarray()[0].tolist()))
# We see that we could optimize by using integers here instead of floats by counting the occurence of every entry.
print(Counter(result[0].toarray()[0].tolist()))
# -
# ## Checking that the cosine similarity before and after word projection is kept
#
# Note that this is a yet low-quality test, as the neural network layers above the projection are absent, so the similary is not yet semantic, it only looks at characters.
# +
word_pairs_to_check_against_each_other = [
# Similar:
["start", "started"],
["prioritize", "priority"],
["twitter", "tweet"],
["Great", "great"],
# Dissimilar:
["boat", "cow"],
["orange", "chewbacca"],
["twitter", "coffee"],
["ab", "ae"],
]
before = pipeline.named_steps["char_term_frequency"].transform(word_pairs_to_check_against_each_other)
after = pipeline.named_steps["union"].transform(before)
for i, word_pair in enumerate(word_pairs_to_check_against_each_other):
cos_sim_before = cosine_similarity(before[i][0], before[i][1])[0,0]
cos_sim_after = cosine_similarity( after[i][0], after[i][1])[0,0]
print("Word pair tested:", word_pair)
print("\t - similarity before:", cos_sim_before,
"\t Are words similar?", "yes" if cos_sim_before > 0.5 else "no")
print("\t - similarity after :", cos_sim_after ,
"\t Are words similar?", "yes" if cos_sim_after > 0.5 else "no")
print("")
# -
# ## Next up
#
# So we have created the sentence preprocessing pipeline and the sparse projection (random hashing) function. We now need a few feedforward layers on top of that.
#
# Also, a few things could be optimized, such as using the power set of the possible n-gram values with a predefined character set instead of fitting it, and the Hashing's fit function could be avoided as well by passing the random seed earlier, because the Hasher doesn't even look at the data and it only needs to be created at some point. This would yield a truly embedding-free approach. Free to you to implement this. I wanted to have something that worked first, leaving optimization for later.
#
#
# ## License
#
#
# BSD 3-Clause License
#
#
# Copyright (c) 2018, <NAME>
#
# All rights reserved.
#
#
# ## Extra links
#
# ### Connect with me
#
# - [LinkedIn](https://ca.linkedin.com/in/chevalierg)
# - [Twitter](https://twitter.com/guillaume_che)
# - [GitHub](https://github.com/guillaume-chevalier/)
# - [Quora](https://www.quora.com/profile/Guillaume-Chevalier-2)
# - [YouTube](https://www.youtube.com/c/GuillaumeChevalier)
# - [Dev/Consulting](http://www.neuraxio.com/en/)
#
# ### Liked this piece of code? Did it help you? Leave a [star](https://github.com/guillaume-chevalier/SGNN-Self-Governing-Neural-Networks-Projection-Layer/stargazers), [fork](https://github.com/guillaume-chevalier/SGNN-Self-Governing-Neural-Networks-Projection-Layer/network/members) and share the love!
#
| SGNN-demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
sys.path.append('../../') # or just install the module
# +
# %load_ext autoreload
# %autoreload 2
import numpy as np
from fuzzytools.datascience.xerror import XError
dummy_xe = XError(None)
print(dummy_xe)
xe1 = XError(np.random.uniform(10,15, size=11))
xe2 = XError(np.random.uniform(20,29, size=12))
xe3 = XError(np.random.uniform(51,52, size=13))
xe4 = XError(np.random.uniform(49,53, size=14))
xe5 = sum([xe1, xe2])
xe6 = xe1+2
print(xe1.get_size(), xe2.get_size(), xe5.get_size(), xe6.get_size())
xe7 = xe6/10
print(xe1)
print('xe2',xe2, xe2.get_size(), xe2.is_1d())
print('xe3',xe3)
print('xe4',xe4)
print('xe5',xe5)
print('xe6',xe6)
print('xe7',xe7)
# -
print(xe7==xe7)
print(xe7==xe6)
xe7.get_rawrepr('f')
# +
# %load_ext autoreload
# %autoreload 2
xe = XError(np.random.uniform(100,101, size=10))
print(xe)
xe = XError(np.random.uniform(1e-7,1e-6, size=10))
print(xe)
xe = XError(np.random.uniform(-1e-7,-1e-6, size=10))
print(xe)
# -
xe1.get_item(0)
| tests/datascience/xerror.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import json
import re
import ast
import json
from graphviz import Digraph
import pandas as pd
# color the graph
import graph_tool.all as gt
import copy
import matplotlib.colors as mcolors
import sys
import matplotlib.colors as mcolors
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
sns.set_style("whitegrid")
import pandas as pd
import os
import json
import itertools
import matplotlib
import matplotlib.ticker as ticker
worker_color = {'10.255.23.108': '#e41a1c',
'10.255.23.109': '#984ea3',
'10.255.23.110': '#ff7f00',
'10.255.23.115': '#4daf4a'}
results_dir = '/local0/serverless/serverless/ipythons/plots'
stats_dir='/opt/dask-distributed/benchmark/stats'
# +
# get all benchmarks
def get_benchmarks():
benchmarks = {}
for _file in os.listdir(stats_dir):
try:
app = _file.split('.', 1)[0]
assert os.path.isfile(os.path.join(stats_dir, f'{app}.g')) \
and os.path.isfile(os.path.join(stats_dir, f'{app}.json')) \
and os.path.isfile(os.path.join(stats_dir, f'{app}.colors'))
bnch, scheduler, _ = app.split('_', 2)
#scheduler = 'vanilla'
benchmarks[app] = {'app': app, 'scheduler': scheduler, 'benchmark': bnch}
except AssertionError:
pass
return benchmarks
# -
# +
worker_color = {'10.255.23.108': '#e41a1c',
'10.255.23.109': '#984ea3',
'10.255.23.110': '#ff7f00',
'10.255.23.115': '#4daf4a'}
def color_assignment(benchmark, task_style):
cfile = f'/opt/dask-distributed/benchmark/stats/{benchmark}.colors'
with open(cfile, 'r') as cfd:
raw = cfd.read().split('\n')
for ln in raw:
if not ln:
continue
task_name, actual = ln.split(',')
if task_name not in task_style:
task_style[task_name] = {}
task_style[task_name]['actual'] = actual
def build_graph(benchmark, task_style):
css_colors = list(mcolors.CSS4_COLORS.keys())
gfile = f'/opt/dask-distributed/benchmark/stats/{benchmark}.g'
with open(gfile, 'r') as fd:
raw = fd.read().split('\n')
g = gt.Graph(directed=True)
vid_to_vx = {}
name_to_vid = {}
g.vertex_properties['name'] = g.new_vertex_property("string")
g.vertex_properties['color'] = g.new_vertex_property("string")
g.vertex_properties['worker'] = g.new_vertex_property("string")
g.vertex_properties['icolor'] = g.new_vertex_property("int")
g.vertex_properties['simcolor'] = g.new_vertex_property("string")
g.vertex_properties['isimcolor'] = g.new_vertex_property("string")
for ln in raw:
if ln.startswith('v'):
_, vid, name = ln.split(',', 2)
v = g.add_vertex()
vid_to_vx[vid] = v
name_to_vid[name] = vid
g.vp.name[v] = name
try:
g.vp.icolor[v] = int(task_style[name]['actual'])
#if g.vp.icolor[v] >= len(css_colors):
#g.vp.color[v] = mcolors.CSS4_COLORS[css_colors[0]]
#else:
g.vp.color[v] = mcolors.CSS4_COLORS[css_colors[int(task_style[name]['actual'])]]
except KeyError:
print(f'Keyerror for {name}')
raise NameError('Error')
#g.vp.color[v] = 'yellow'
#g.vp.icolor[v] = 2
for ln in raw:
if ln.startswith('e'):
_, vsrc, vdst, _ = ln.split(',', 3)
g.add_edge(vid_to_vx[vsrc], vid_to_vx[vdst])
return g
keys = list(mcolors.CSS4_COLORS.keys())
def update_runtime_state(benchmark, g, task_style):
print('process', benchmark)
tasks = []
jfile = f'/opt/dask-distributed/benchmark/stats/{benchmark}.json'
with open(jfile, 'r') as fd:
stats = ast.literal_eval(fd.read())
#print(json.dumps(stats, indent=4))
print('stat size is', len(stats))
min_ts = sys.maxsize
for s in stats:
task_style[s]['output_size'] = stats[s]['msg']['nbytes']
task_style[s]['input_size'] = 0
task_style[s]['remote_read'] = 0
task_style[s]['local_read'] = 0
task_style[s]['worker'] = stats[s]['worker'].split(':')[1].replace('/', '')
startsstops = stats[s]['msg']['startstops']
for ss in startsstops:
if ss['action'] == 'inputsize': continue
if ss['action'] == 'compute':
task_style[s]['compute_end'] = ss['stop']
task_style[s]['compute_start'] = ss['start']
task_style[s]['runtime'] = ss['stop'] - ss['start']
if ss['start'] < min_ts: min_ts = ss['start']
if ss['stop'] < min_ts: min_ts = ss['stop']
print(min_ts)
for s in stats:
startsstops = stats[s]['msg']['startstops']
min_start = sys.maxsize
max_end = 0
transfer_stop = 0
transfer_start = sys.maxsize
for ss in startsstops:
if ss['action'] == 'inputsize': continue
if ss['start'] < min_start: min_start = ss['start']
if ss['stop'] > max_end: max_end = ss['stop']
if ss['action'] == 'compute':
compute_stop = ss['stop']
compute_start = ss['start']
run_time = ss['stop'] - ss['start']
if ss['action'] == 'transfer':
#print(ss['start'] - min_ts, ss['stop'] - min_ts)
if ss['start'] < transfer_start:
transfer_start = ss['start']
if ss['stop'] > transfer_stop:
transfer_stop = ss['stop']
#print('transfer start', ss['start'] - min_ts,
# 'transfer stop', ss['stop'] - min_ts)
if transfer_stop == 0:
transfer_stop = compute_start
transfer_start = compute_start
#print('***transfer start', transfer_start - min_ts,
# '****transfer stop', transfer_stop - min_ts)
tasks.append({'name': s, 'start_ts': min_start - min_ts, 'end_ts': max_end - min_ts,
'compute_start': compute_start - min_ts, 'compute_stop': compute_stop - min_ts,
'transfer_start': transfer_start - min_ts, 'transfer_stop': transfer_stop - min_ts,
'worker': stats[s]['worker'].split(':')[1].replace('/', '')})
#print('\n')
#total amount of data accessed, data accessed remotely, data accessed locally
for v in g.vertices():
for vi in v.in_neighbors():
task_style[g.vp.name[v]]['input_size'] += task_style[g.vp.name[vi]]['output_size']
if task_style[g.vp.name[v]]['worker'] == task_style[g.vp.name[vi]]['worker']:
task_style[g.vp.name[v]]['local_read'] += task_style[g.vp.name[vi]]['output_size']
else:
task_style[g.vp.name[v]]['remote_read'] += task_style[g.vp.name[vi]]['output_size']
for v in g.vertices():
g.vp.worker[v] = task_style[g.vp.name[v]]['worker']
#g.vp.color[v] = colors[task_style[g.vp.name[v]]['worker']]
#Check the slack for the prefetching
bw = 10*(1<<27) # 10 Gbps (1<<30)/(1<<3)
not_from_remote = 0
for v in g.vertices():
parents_end = []
for vi in v.in_neighbors():
parents_end.append(task_style[g.vp.name[vi]]['compute_end'])
if len(parents_end):
max_end = max(parents_end)
for vi in v.in_neighbors():
if max_end == task_style[g.vp.name[vi]]['compute_end'] and task_style[g.vp.name[vi]]['worker'] != task_style[g.vp.name[v]]['worker']:
#print(f'Slack come from local chain')
not_from_remote += 1
#print(f'slack for {g.vp.name[v]}: {round(1000*(max(parents_end) - min(parents_end)), 2)}msec',
# '\t runtime:', round(1000*task_style[g.vp.name[vi]]['runtime'], 4), 'msec',
# '\t remote read', task_style[g.vp.name[v]]['remote_read']/bw)
#print(not_from_remote)
return tasks
def plot_graph(g):
policy = b.split('_')[1]
print('policy is', policy)
dg = Digraph('G', filename=f'{b}.gv', format='png')
for v in g.vertices():
#print(g.vp.name[v])
vname = g.vp.name[v].split('-', 1)
vname = vname[1] if len(vname) > 1 else vname[0]
#dg.attr('node', shape='ellipse', style='filled', color=g.vp.color[v])
dg.attr('node', shape='ellipse', style="filled,solid",
penwidth="3",
fillcolor=g.vp.color[v] if "chaincolor" in policy else "#f0f0f0",
color=worker_color[g.vp.worker[v]])
color = '-'
if 'chaincolor' in policy:
color = g.vp.icolor[v]
dg.node(f'{vname}, color({color})')
#print(g.vp.name[v], g.vp.icolor[v], g.vp.worker[v])
for e in g.edges():
vname = g.vp.name[e.source()].split('-', 1)
sname = vname[1] if len(vname) > 1 else vname[0]
vname = g.vp.name[e.target()].split('-', 1)
tname = vname[1] if len(vname) > 1 else vname[0]
dg.edge(f'{sname}, color({g.vp.icolor[e.source()] if "chaincolor" in policy else "-"})',
f'{tname}, color({g.vp.icolor[e.target()] if "chaincolor" in policy else "-"})')
dg.view(f'/local0/serverless/serverless/ipythons/plots/{b}', quiet=False)
def plot_graph2(g):
policy = b.split('_')[1]
dg = Digraph('G', filename=f'{b.split("_")[0]}.gv', format='png')
for v in g.vertices():
vname = g.vp.name[v].split('-', 1)
vname = vname[1] if len(vname) > 1 else vname[0]
dg.attr('node', shape='ellipse', style='filled', color='#252525')
dg.attr('node', shape='ellipse', style="filled,solid",
penwidth="3",
fillcolor= "#f0f0f0",
color='#252525')
dg.node(f'{vname}')
for e in g.edges():
vname = g.vp.name[e.source()].split('-', 1)
sname = vname[1] if len(vname) > 1 else vname[0]
vname = g.vp.name[e.target()].split('-', 1)
tname = vname[1] if len(vname) > 1 else vname[0]
dg.edge(f'{sname}',
f'{tname}')
dg.view(f'/local0/serverless/serverless/ipythons/plots/{b.split("_")[0]}', quiet=False)
def format_xticks(x, pos=None):
return x
#return str(int(x*1000))
def plot_gannt_chart(tasks, benchmark):
#_max = df['runtime'].max()
#print(benchmark, df['runtime'].max())
sns.set_style("ticks")
sns.set_context("paper", font_scale=1)
sns.set_context(rc = {'patch.linewidth': 1.5, 'patch.color': 'black'})
plt.rc('font', family='serif')
fig, ax = plt.subplots(figsize=(10,8))
ax.set_xlabel('Time (second)', fontsize=16)
sns.despine()
ax.yaxis.grid(color='#99999910', linestyle=(0, (5, 10)), linewidth=0.4)
ax.set_axisbelow(True)
ax.tick_params(axis='both', which='major', labelsize=14)
ax.yaxis.set_ticks_position('both')
ax.xaxis.set_major_formatter(ticker.FuncFormatter(format_xticks))
ax.set_yticklabels([])
#ax.set_xlim([0, _max])
# Setting graph attribute
ax.grid(True)
base = 0
size = 8
margin = 3
workers_load={}
for ts in tasks:
#print(json.dumps(ts, indent=4))
#print(ts['name'], ts['start_ts'], ts['end_ts'], ts['worker'])
if ts['worker'] not in workers_load:
workers_load[ts['worker']] = []
#workers_load[ts['worker']].append((ts['start_ts'], ts['end_ts'] - ts['start_ts']))
ax.broken_barh([(ts['transfer_start'], ts['transfer_stop'] - ts['transfer_start'])], (base, size),
edgecolors =worker_color[ts['worker']], facecolors = worker_color[ts['worker']])
ax.broken_barh([(ts['compute_start'], ts['compute_stop'] - ts['compute_start'])], (base, size),
edgecolors = worker_color[ts['worker']], facecolors='#f0f0f0')
vname = ts['name'].split('-', 1)
sname = vname[1] if len(vname) > 1 else vname[0]
ax.text(x=ts['compute_start'] + (ts['compute_stop'] - ts['compute_start'])/2,
y=base + size/2,
s=sname,
ha='center',
va='center',
color='black')
base += (size + margin)
jobname, policy, _ = benchmark.split('_')
#ax.axes.yaxis.set_visible(False)
ax.set_ylabel(f'{benchmark}', fontsize=14)
ax.set_title(f'{jobname}, {policy}', fontsize=16)
leg1 = ax.legend(['10.255.23.108', '10.255.23.109', '10.255.23.109', '10.255.23.115'],
title='Communication cost', ncol = 1, loc='lower center')
leg1.legendHandles[0].set_color(worker_color['10.255.23.108'])
leg1.legendHandles[1].set_color(worker_color['10.255.23.109'])
leg1.legendHandles[2].set_color(worker_color['10.255.23.110'])
leg1.legendHandles[3].set_color(worker_color['10.255.23.115'])
leg2 = ax.legend(['10.255.23.108', '10.255.23.109', '10.255.23.109', '10.255.23.115'],
title='Computation cost',loc='lower right')
leg2.legendHandles[0].set_edgecolor(worker_color['10.255.23.108'])
leg2.legendHandles[1].set_edgecolor(worker_color['10.255.23.109'])
leg2.legendHandles[2].set_edgecolor(worker_color['10.255.23.110'])
leg2.legendHandles[3].set_edgecolor(worker_color['10.255.23.115'])
leg2.legendHandles[0].set_facecolor('#f0f0f0')
leg2.legendHandles[1].set_facecolor('#f0f0f0')
leg2.legendHandles[2].set_facecolor('#f0f0f0')
leg2.legendHandles[3].set_facecolor('#f0f0f0')
#end_2_end = df[df['exp'] == benchmark]['runtime']
#ax.vlines(end_2_end, ymin=0, ymax=base, label = end_2_end, linestyles='--', color='#bdbdbd')
ax.add_artist(leg1)
fig.savefig(f'{os.path.join("/local0/serverless/serverless/ipythons/plots",benchmark)}.gannt.png',
format='png', dpi=200)
plt.show()
def plot_runtime(name, df):
sns.set_style("ticks")
sns.set_context("paper", font_scale=1)
sns.set_context(rc = {'patch.linewidth': 1.5, 'patch.color': 'black'})
plt.rc('font', family='serif')
fig, ax = plt.subplots(figsize=(8,5))
sns.barplot(x='scheduler', y='runtime', #, hue = 'scheduler',
order = ['optimal', 'vanilla', 'optplacement', 'chaincolorrr', 'chaincolorch', 'random'],
palette = ['#fbb4ae','#fdb462', '#8dd3c7', '#80b1d3', '#bebada' , '#fb8072'],
data=df, ax=ax)
sns.despine()
ax.yaxis.grid(color='#99999910', linestyle=(0, (5, 10)), linewidth=0.4)
ax.set_axisbelow(True)
ax.tick_params(axis='x', which='major', labelsize=14, rotation=15)
ax.tick_params(axis='y', which='major', labelsize=14)
ax.yaxis.set_ticks_position('both')
ax.set_ylabel('Runtime (sec)', fontsize=16)
ax.set_xlabel(f'Benchmark: {df["benchmark"].unique()[0]}', fontsize=16)
#ax.legend(fontsize=14, ncol=3)
plt.tight_layout()
fig.savefig(f'/local0/serverless/serverless-sim/results/runtime_{name}.png',
format='png', dpi=200)
plt.show()
#dt = pd.read_csv('/local0/serverless/task-bench/dask/stats.csv')
#dt.head(5)
#gdf = dt.groupby('name')
benchmarks = ['tree.w10.s8.1G1B_vanilla_3d70b39c'] #get_benchmarks()
#for name, df in gdf:
#plot_runtime(name, df)
#continue
#if 'spread' not in name: continue
for b in benchmarks:
#if name not in b: continue
task_style = {}
color_assignment(b, task_style)
g = build_graph(b, task_style)
tasks = update_runtime_state(b, g, task_style)
plot_graph(g)
plot_graph2(g)
plot_gannt_chart(tasks, b)
#break
#break
# -
| ipythons/executions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import copy
import numpy as np
import skimage
from skimage.transform import warp
from matplotlib import pyplot as plt
import time
class SkimageAugmentor:
def __init__(self, cfg, **kwargs):
self.cfg = cfg
def aug_image(self, image, **kwargs):
# exposure
image = skimage.exposure.adjust_gamma(image, gamma=0.9, gain=1)
image = skimage.exposure.adjust_sigmoid(image, cutoff=0.5, gain=10, inv=False)
image = skimage.exposure.adjust_log(image, gain=1, inv=False)
# transform
tform = skimage.transform.SimilarityTransform(scale=(0.99, 1.01), rotation=0.1, translation=(-10, 10))
image = warp(image, tform)
tform = skimage.transform.EuclideanTransform(matrix=None, rotation=0.1, translation=(-10, 10))
image = warp(image, tform)
tform = skimage.transform.AffineTransform(matrix=None, scale=(0.9, 1.1), rotation=0.1, shear=0.1, translation=(-10, 10))
image = warp(image, tform)
tform = skimage.transform.ProjectiveTransform(matrix=np.array([[0.99,0.01,0],[0.01,0.99,0],[0,0,1]]))
image = warp(image, tform)
# utils
image = skimage.util.random_noise(image, mode='gaussian', seed=None, clip=True)
# filters
image = skimage.filters.gaussian(image, sigma=1, output=None, mode='nearest', cval=0, multichannel=None, preserve_range=False, truncate=4.0)
image = skimage.filters.median(image, selem=None, out=None, mask=None, shift_x=False, shift_y=False, mode='nearest', cval=0.0, behavior='ndimage')
image = skimage.filters.laplace(image, ksize=3, mask=None)
image = skimage.filters.meijering(image, sigmas=range(1, 5, 2), alpha=None, black_ridges=True)
image = skimage.filters.sato(image, sigmas=range(1, 5, 2), black_ridges=True)
image = skimage.filters.frangi(image, sigmas=range(1, 5, 2), scale_range=None, scale_step=None, beta1=None, beta2=None, alpha=0.5, beta=0.5, gamma=15, black_ridges=True)
image = skimage.filters.hessian(image, sigmas=range(1, 5, 2), scale_range=None, scale_step=None, beta1=None, beta2=None, alpha=0.5, beta=0.5, gamma=15, black_ridges=True)
return image
skimageAugmentor = SkimageAugmentor(None)
org_image = skimage.io.imread("../data/01.jpg", as_gray=False)
plt.imshow(org_image)
# -
# image
aug_img = skimageAugmentor.aug_image(org_image)
plt.imshow(aug_img)
print(np.max(aug_img.astype(np.float) - org_image.astype(np.float)))
print("image augmentation is done!")
| skimage/skimage_augmentor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Wide & Deep Recommender Demo
# Wide and Deep Learning Model, proposed by Google in 2016, is a DNN-Linear mixed model. Wide and deep learning has been used for Google App Store for their app recommendation.
#
# In this tutorial, we use Recommender API of Analytics Zoo to build a wide linear model and a deep neural network, which is called Wide&Deep model, and use optimizer of BigDL to train the neural network. Wide&Deep model combines the strength of memorization and generalization. It's useful for generic large-scale regression and classification problems with sparse input features (e.g., categorical features with a large number of possible feature values).
# ## Intialization
# * import necessary libraries
# +
from zoo.models.recommendation import *
from zoo.models.recommendation.utils import *
from zoo.common.nncontext import init_nncontext
import os
import sys
import datetime as dt
from bigdl.dataset.transformer import *
from bigdl.dataset.base import *
from bigdl.nn.criterion import *
from bigdl.optim.optimizer import *
from bigdl.util.common import *
import matplotlib
matplotlib.use('agg')
import matplotlib.pyplot as plt
# %pylab inline
# -
# * Initilaize NN context, it will get a SparkContext with optimized configuration for BigDL performance.
sc = init_nncontext("WideAndDeep Example")
# ## Data Preparation
# * Download and read movielens 1M rating data, understand the dimension.
# +
from bigdl.dataset import movielens
movielens_data = movielens.get_id_ratings("/tmp/movielens/")
min_user_id = np.min(movielens_data[:,0])
max_user_id = np.max(movielens_data[:,0])
min_movie_id = np.min(movielens_data[:,1])
max_movie_id = np.max(movielens_data[:,1])
rating_labels= np.unique(movielens_data[:,2])
print(movielens_data.shape)
print(min_user_id, max_user_id, min_movie_id, max_movie_id, rating_labels)
# -
# * Transform ratings into dataframe, read user and item data into dataframes.
# +
sqlContext = SQLContext(sc)
from pyspark.sql.types import *
from pyspark.sql import Row
Rating = Row("userId", "itemId", "label")
User = Row("userId", "gender", "age" ,"occupation")
Item = Row("itemId", "title" ,"genres")
ratings = sc.parallelize(movielens_data) \
.map(lambda line: map(int, line)) \
.map(lambda r: Rating(*r))
ratingDF = sqlContext.createDataFrame(ratings)
users= sc.textFile("/tmp/movielens/ml-1m/users.dat")\
.map(lambda line: line.split("::")[0:4])\
.map(lambda line: (int(line[0]), line[1], int(line[2]), int(line[3])))\
.map(lambda r: User(*r))
userDF = sqlContext.createDataFrame(users)
items = sc.textFile("/tmp/movielens/ml-1m/movies.dat") \
.map(lambda line: line.split("::")[0:3]) \
.map(lambda line: (int(line[0]), line[1], line[2].split('|')[0])) \
.map(lambda r: Item(*r))
itemDF = sqlContext.createDataFrame(items)
# -
# * Join data together, and transform data. For example, gender is going be used as categorical feature, occupation and gender will be used as crossed features.
# +
from pyspark.sql.functions import col, udf
gender_udf = udf(lambda gender: categorical_from_vocab_list(gender, ["F", "M"], start=1))
bucket_cross_udf = udf(lambda feature1, feature2: hash_bucket(str(feature1) + "_" + str(feature2), bucket_size=100))
genres_list = ["Crime", "Romance", "Thriller", "Adventure", "Drama", "Children's",
"War", "Documentary", "Fantasy", "Mystery", "Musical", "Animation", "Film-Noir", "Horror",
"Western", "Comedy", "Action", "Sci-Fi"]
genres_udf = udf(lambda genres: categorical_from_vocab_list(genres, genres_list, start=1))
allDF = ratingDF.join(userDF, ["userId"]).join(itemDF, ["itemId"]) \
.withColumn("gender", gender_udf(col("gender")).cast("int")) \
.withColumn("age-gender", bucket_cross_udf(col("age"), col("gender")).cast("int")) \
.withColumn("genres", genres_udf(col("genres")).cast("int"))
allDF.show(5)
# -
# * Speficy data feature information shared by the WideAndDeep model and its feature generation. Here, we use occupation gender for wide base part, age and gender crossed as wide cross part, genres and gender as indicators, userid and itemid for embedding.
bucket_size = 100
column_info = ColumnFeatureInfo(
wide_base_cols=["occupation", "gender"],
wide_base_dims=[21, 3],
wide_cross_cols=["age-gender"],
wide_cross_dims=[bucket_size],
indicator_cols=["genres", "gender"],
indicator_dims=[19, 3],
embed_cols=["userId", "itemId"],
embed_in_dims=[max_user_id, max_movie_id],
embed_out_dims=[64, 64],
continuous_cols=["age"])
# * Transform data to RDD of Sample. We use optimizer of BigDL directly to train the model, it requires data to be provided in format of RDD(Sample). A Sample is a BigDL data structure which can be constructed using 2 numpy arrays, feature and label respectively. The API interface is Sample.from_ndarray(feature, label). Wide&Deep model need two input tensors, one is SparseTensor for the Wide model, another is a DenseTensor for the Deep model.
rdds = allDF.rdd.map(lambda row: to_user_item_feature(row, column_info))
trainPairFeatureRdds, valPairFeatureRdds = rdds.randomSplit([0.8, 0.2], seed= 1)
valPairFeatureRdds.persist()
train_data= trainPairFeatureRdds.map(lambda pair_feature: pair_feature.sample)
test_data= valPairFeatureRdds.map(lambda pair_feature: pair_feature.sample)
# ## Create the Wide&Deep model.
# * In Analytics Zoo, it is simple to build Wide&Deep model by calling WideAndDeep API. You need specify model type, and class number, as well as column information of features according to your data. You can also change other default parameters in the network, like hidden layers. The model could be fed into an Optimizer of BigDL or NNClassifier of analytics-zoo. Please refer to the document for more details. In this example, we demostrate how to use optimizer of BigDL.
wide_n_deep = WideAndDeep(5, column_info, "wide_n_deep")
# ## Create optimizer and train the model
# +
# Create an Optimizer
batch_size = 8000
optimizer = Optimizer(
model=wide_n_deep,
training_rdd=train_data,
criterion=ClassNLLCriterion(),
optim_method=Adam(learningrate = 0.001, learningrate_decay=0.00005),
end_trigger=MaxEpoch(10),
batch_size=batch_size)
# Set the validation logic
optimizer.set_validation(
batch_size=batch_size,
val_rdd=test_data,
trigger=EveryEpoch(),
val_method=[Top1Accuracy(), Loss(ClassNLLCriterion())]
)
log_dir='/tmp/bigdl_summaries/'
app_name='wide_n_deep-'+dt.datetime.now().strftime("%Y%m%d-%H%M%S")
train_summary = TrainSummary(log_dir=log_dir,
app_name=app_name)
val_summary = ValidationSummary(log_dir=log_dir,
app_name=app_name)
optimizer.set_train_summary(train_summary)
optimizer.set_val_summary(val_summary)
print("saving logs to %s" % (log_dir + app_name))
# -
# Train the network. Wait some time till it finished.. Voila! You've got a trained model
# %%time
# Boot training process
optimizer.optimize()
print("Optimization Done.")
# ## Prediction and recommendation
# * Zoo models make inferences based on the given data using model.predict(val_rdd) API. A result of RDD is returned. predict_class returns the predicted label.
# +
results = wide_n_deep.predict(test_data)
results.take(5)
results_class = wide_n_deep.predict_class(test_data)
results_class.take(5)
# -
# * In the Analytics Zoo, Recommender has provied 3 unique APIs to predict user-item pairs and make recommendations for users or items given candidates.
# * Predict for user item pairs
userItemPairPrediction = wide_n_deep.predict_user_item_pair(valPairFeatureRdds)
for result in userItemPairPrediction.take(5): print(result)
# * Recommend 3 items for each user given candidates in the feature RDDs
userRecs = wide_n_deep.recommend_for_user(valPairFeatureRdds, 3)
for result in userRecs.take(5): print(result)
# * Recommend 3 users for each item given candidates in the feature RDDs
itemRecs = wide_n_deep.recommend_for_item(valPairFeatureRdds, 3)
for result in itemRecs.take(5): print(result)
# ## Evaluate the trained model
# %%time
evaluate_result=wide_n_deep.evaluate(test_data, 2800, [Top1Accuracy()])
print("Top1 accuracy: %s" % evaluate_result[0].result)
# ## Draw the convergence curve
# +
loss = np.array(train_summary.read_scalar("Loss"))
top1 = np.array(val_summary.read_scalar("Top1Accuracy"))
plt.figure(figsize = (12,12))
plt.subplot(2,1,1)
plt.plot(loss[:,0],loss[:,1],label='loss')
plt.xlim(0,loss.shape[0]+10)
plt.grid(True)
plt.title("loss")
plt.subplot(2,1,2)
plt.plot(top1[:,0],top1[:,1],label='top1')
plt.xlim(0,loss.shape[0]+10)
plt.title("top1 accuracy")
plt.grid(True)
# -
valPairFeatureRdds.unpersist()
| apps/recommendation-wide-n-deep/wide_n_deep.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import sys
# Modify the path
sys.path.append("..")
import pandas as pd
import yellowbrick as yb
import matplotlib.pyplot as plt
from sklearn.datasets import load_diabetes
# +
import matplotlib.pyplot as plt
import numpy as np
from yellowbrick.base import Visualizer
from yellowbrick.exceptions import YellowbrickValueError
##########################################################################
## TargetVisualizer Base Class
##########################################################################
class TargetVisualizer(Visualizer):
"""
The base class for target visualizers, generic enough to support any
computation on a single vector, y. This Visualizer is based on the
LabelEncoder in sklearn.preprocessing, which only accepts a target y.
"""
def fit(self, y):
"""
Fit the visualizer to the target y. Note that this visualizer breaks
the standard estimator interface, and therefore cannot be used inside
of pipelines, but must be used separately; similar to how the
LabelEncoder is used.
"""
raise NotImplementedError(
"target visualizers must implement a fit method"
)
##########################################################################
## Balanced Binning Reference
##########################################################################
class BalancedBinningReference(TargetVisualizer):
"""
BalancedBinningReference generates a histogram with vertical lines
showing the recommended value point to bin your data so they can be evenly
distributed in each bin.
Parameters
----------
ax : matplotlib Axes, default: None
This is inherited from FeatureVisualizer and is defined within
``BalancedBinningReference``.
target : string, default: "Frequency"
The name of the ``y`` variable
bins : number of bins to generate the histogram, default: 4
kwargs : dict
Keyword arguments that are passed to the base class and may influence
the visualization as defined in other Visualizers.
Attributes
----------
bin_edges : binning reference values
Examples
--------
>>> visualizer = BalancedBinningReference()
>>> visualizer.fit(y)
>>> visualizer.poof()
Notes
-----
These parameters can be influenced later on in the visualization
process, but can and should be set as early as possible.
"""
def __init__(self, ax=None, target=None, bins=4, **kwargs):
super(BalancedBinningReference, self).__init__(ax, **kwargs)
self.target = target
self.bins = bins
def draw(self, y, **kwargs):
"""
Draws a histogram with the reference value for binning as vertical
lines.
Parameters
----------
y : an array of one dimension or a pandas Series
"""
# draw the histogram
hist, bin_edges = np.histogram(y, bins=self.bins)
self.bin_edges_ = bin_edges
self.ax.hist(y, bins=self.bins, color=kwargs.pop("color", "#6897bb"), **kwargs)
# add vetical line with binning reference values
plt.vlines(bin_edges,0,max(hist),colors=kwargs.pop("colors", "r"))
def fit(self, y, **kwargs):
"""
Sets up y for the histogram and checks to
ensure that ``y`` is of the correct data type.
Fit calls draw.
Parameters
----------
y : an array of one dimension or a pandas Series
kwargs : dict
keyword arguments passed to scikit-learn API.
"""
#throw an error if y has more than 1 column
if y.ndim > 1:
raise YellowbrickValueError("y needs to be an array or Series with one dimension")
# Handle the target name if it is None.
if self.target is None:
self.target = 'Frequency'
self.draw(y)
return self
def poof(self, **kwargs):
"""
Creates the labels for the feature and target variables.
"""
self.ax.set_xlabel(self.target)
self.finalize(**kwargs)
def finalize(self, **kwargs):
"""
Finalize executes any subclass-specific axes finalization steps.
The user calls poof and poof calls finalize.
Parameters
----------
kwargs: generic keyword arguments.
"""
for tk in self.ax.get_xticklabels():
tk.set_visible(True)
for tk in self.ax.get_yticklabels():
tk.set_visible(True)
##########################################################################
## Quick Method
##########################################################################
def balanced_binning_reference(y, ax=None, target='Frequency', bins=4, **kwargs):
"""
BalancedBinningReference generates a histogram with vertical lines
showing the recommended value point to bin your data so they can be evenly
distributed in each bin.
Parameters
----------
y : an array of one dimension or a pandas Series
ax : matplotlib Axes, default: None
This is inherited from FeatureVisualizer and is defined within
``BalancedBinningReference``.
target : string, default: "Frequency"
The name of the ``y`` variable
bins : number of bins to generate the histogram, default: 4
kwargs : dict
Keyword arguments that are passed to the base class and may influence
the visualization as defined in other Visualizers.
"""
# Initialize the visualizer
visualizer = BalancedBinningReference(ax=ax, bins=bins, target=target, **kwargs)
# Fit and poof the visualizer
visualizer.fit(y)
visualizer.poof()
# +
def balanced_binning_reference():
# Load a regression data set
data = load_diabetes()
# Extract the target variable
y = data['target']
# Instantiate and fit the visualizer
visualizer = BalancedBinningReference()
visualizer.fit(y)
return visualizer.poof()
# -
balanced_binning_reference()
| examples/rebeccabilbro/balanced_binning_docs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: VPython
# language: python
# name: vpython
# ---
# +
# Fun some setup for the project
# Silence annoying pytorch deprecated warnings
import warnings
warnings.filterwarnings("ignore")
import matplotlib.pyplot as plt
from eval_nll import *
import numpy as np
# %matplotlib inline
# for auto-reloading extenrnal modules
# see http://stackoverflow.com/questions/1907993/autoreload-of-modules-in-ipython
# %load_ext autoreload
# %autoreload 2
# -
# # Letter_low Dataset Notebook. Treat Letter-low as the normal dataset and Letter-med / Letter-high as the anomalous. Namely, we treat noise as the anomaly.
nll_dir = 'nll/Letter-low'
# +
# Step 1
# Load the training and test nll predictions to test generalization
train_nlls = load_avg_nlls(nll_dir, 'train')
test_nlls = load_avg_nlls(nll_dir, 'test')
# Plot distribtutions
fig, ax_compare = compare_dist([train_nlls, test_nlls], ['Letter-low Train', 'Letter-low Test'], 'Train vs. Test Distributions')
# +
# Step 2
# Compare the test graphs from the normal class nlls with the anomalous graph nlls
anom_nlls_med = load_avg_nlls(nll_dir, 'Letter-med')
anom_nlls_high = load_avg_nlls(nll_dir, 'Letter-high')
nlls = [train_nlls, test_nlls, anom_nlls_med, anom_nlls_high]
labels = ['Low Noise (norm)', 'Low Noise Test (norm)', 'Med Noise (anom)', 'High Noise (anom)']
fig, ax_compare = compare_dist(nlls, labels, 'N Vs. All other letter distributions')
# +
# Step 2
# Compare the test graphs from the normal class nlls with the anomalous graph nlls
anom_nlls_high = load_avg_nlls(nll_dir, 'Letter-high')
nlls = [train_nlls, test_nlls, anom_nlls_high]
labels = ['Low Noise (norm)', 'Low Noise Test (norm)', 'High Noise (anom)']
fig, ax_compare = compare_dist(nlls, labels, 'N Vs. All other letter distributions')
# -
| Letters-Noise-GraphRNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: py37astro
# language: python
# name: py37astro
# ---
# +
# %matplotlib notebook
import sys
from pathlib import Path
SRC_ROOT_DIR_0 = '/g/wsl_projs/practical-astronomy'
SRC_ROOT_DIR_1 = '/g/wsl_projs/practical-astronomy/myastro/'
sys.path.insert(0, SRC_ROOT_DIR_0)
sys.path.insert(1, SRC_ROOT_DIR_1)
# %load_ext autoreload
# %autoreload 2
# +
import myastro.timeconv as tc
import myastro.coord as co
import myastro.orbit as ob
from myastro.orbit import OrbObject
from timeconv import sin_dgms, cos_dgms, tan_dgms
import numpy as np
from toolz import pipe, compose
import toolz as tz
from functools import partial
from itertools import starmap
import myastro.util as ut
from operator import itemgetter
import matplotlib.pyplot as plt
from matplotlib import pyplot
import matplotlib.animation as animation
from matplotlib.animation import FuncAnimation
from matplotlib.patches import Ellipse
# +
#commets
# Ellipse (e=0.5) Parabola (e=1.0) Hyperbola (e=1.5)
# +
#Position in the orbit
#Although the equation for the conic section gives every possible poisition at which a comet may be found during ti
# its orbit, it does not establish at what time this will occur.
# To solve this proble, we will use the law of equal areas.
# The total area S of the ellipse is
# S = pi*a^2*sqrt(1-e^2)
# dx_dt = -np.sqrt(GM/a)*np.sin(e_anomaly/(1-e*cos(e_anomaly)))
# dy_dt = ...
# x =
# +
obliq = np.deg2rad(23.5)
r_eclip = np.array([1,2,3])
# -
r_equat = ob.Rx_3d(-obliq).dot(r_eclip)
r_equat
# +
def rect2polar(x,y,z):
r = np.sqrt(x*x+y*y+z*z)
phi = np.arctan2(y,x)
theta = np.arctan2(z,np.sqrt(x*x+y*y))
return r, phi, theta
def rec2polar_v(vect):
return np.array(rec2polar(*vect))
def polar2rect(r,phi,theta) :
r_cos_theta = r*np.cos(theta)
return r_cos_theta*np.cos(phi), r_cos_theta*np.sin(phi), r*np.sin(theta)
def polar2rect_v(vect):
return np.array(polar2rect(*vect))
# -
def rect2polar2(r_v):
return np.array(rect2polar(*r_v))
compose(rect2polar2,np.array)((1,2,3))
compose(rect2polar,*tuple)(rv)
# +
def function1(function):
def wrapper(*args, **kwargs):
tuple = args[0]
result = function(*args, **kwargs)
return np.array(result)
print("welcome to edureka")
return wrapper
@function1
def function2(name):
print(f"{name}")
function2("pythonista")
# -
np.array((1,2))
| notebooks/comets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Visualizing Chipotle's Data
# This time we are going to pull data directly from the internet.
# Special thanks to: https://github.com/justmarkham for sharing the dataset and materials.
#
# ### Step 1. Import the necessary libraries
# +
import pandas as pd
import matplotlib.pyplot as plt
from collections import Counter
# set this so the graphs open internally
# %matplotlib inline
# -
# ### Step 2. Import the dataset from this [address](https://raw.githubusercontent.com/justmarkham/DAT8/master/data/chipotle.tsv).
# ### Step 3. Assign it to a variable called chipo.
chipo = pd.read_csv('https://raw.githubusercontent.com/justmarkham/DAT8/master/data/chipotle.tsv', sep="\t")
chipo.info()
# ### Step 4. See the first 10 entries
chipo.head(10)
# ### Step 5. Create a histogram of the top 5 items bought
chipo['item_price'] = chipo['item_price'].apply(lambda x: float(str(x).replace('$', '')) )
top = chipo.groupby(['item_name'])['quantity'].value_counts()\
.rename('item_qtd')\
.reset_index()\
.sort_values('item_qtd', ascending=False)
chipo_5 = top.loc[top.index[0:5]]
chipo_5
chipo_5.hist(bins=10);
chipo_5.plot.bar(x='item_name', y='item_qtd', rot=60);
# ### Step 6. Create a scatterplot with the number of items orderered per order price
# #### Hint: Price should be in the X-axis and Items ordered in the Y-axis
items = chipo.sort_values('item_price', ascending=False)
items
# +
items.plot.scatter(x='item_price', y='quantity', s=200, marker='^', color='green', figsize=(16, 8));
plt.xlabel('Item Price $');
plt.ylabel('Quantity');
plt.show();
# -
# ### Step 7. BONUS: Create a question and a graph to answer your own question.
# +
# Qual o pedido com mais quantidade compradas
# -
pedidos = chipo.groupby('order_id')['quantity'].sum().rename('qtd.').reset_index().sort_values('qtd.', ascending=False)
top_10 = pedidos.loc[pedidos.index[0:10]]
top_10
top_10.plot.bar(x='order_id', y='qtd.', rot=60, figsize=(16,8));
plt.xlabel('Pedidos');
plt.ylabel('Quantidades de itens');
| Analysis/Practices/Pandas/07_Visualization/Chipotle/Exercises.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
# ## DataTypes and Attributes
# ndarray => n dimensional array
a1 = np.array([1, 2, 3])
a1
type(a1)
a2 = np.array([[1, 2.2, 3],
[4, 5, 6.5]])
a3 = np.array([[[123, 2.2, 3],
[42, 5, 6.5],
[7, 8, 9]],
[[10, 11, 12],
[13, 14, 15],
[16, 17, 18]]])
a2
a3
a1.shape
a2.shape
a3.shape
a4 = np.array([[[123, 2.2, 3],
[42, 5, 6.5]],
[[10, 11, 12],
[13, 14, 15]]])
a4
a4.shape
a1.ndim, a2.ndim, a3.ndim, a4.ndim
a1.size, a2.size, a3.size, a4.size
type(a1)
# +
# create dataframe from a NumPy array
import pandas as pd
df = pd.DataFrame(a2)
df
# -
# ## 2. Creating arrays
ones = np.ones(2)
ones
zeros = np.zeros((2, 4))
zeros
range_array = np.arange(0,10,2)
range_array
random_array = np.random.randint(0, 10, size=(2,4))
random_array
np.random.random((3, 4))
np.random.rand(3, 4)
# +
# pseudo-random numbers
np.random.seed(seed=42)
random_array_2 = np.random.randint(10, size=(5,3))
random_array_2
# -
np.random.seed(7)
np.random.random((5, 3))
# ## 3. Viewing arrays and matrices
np.unique(random_array_2)
a2[0]
a2[0][0]
a3[0][0][0]
a2[:2]
a2[:5]
a3[:2]
a3[0, 0, 0]
a3
a3[:2, :1, :2]
a5 = np.random.randint(10, size=(2, 3, 4, 5))
a5
# get only first 4 vectors of inner most array
a5[:, :, :, :4]
# ## 4. Manipulating and comparing Arrays
# ### Arithmetic
a1
ones = np.ones(3)
ones
a1 + ones
a1 - ones
a1 * ones
a2
a1 * a2
a3
a1 * a3
a1 / ones
a2 / a1
a2 // a1 # floor division
a2 ** 2
np.square(a2)
a2 % 2 # remainder
np.log(a1)
# ### Aggregation
my_list = [1, 2, 3]
sum(my_list)
np.sum(a1)
massive_array = np.random.random(100000)
massive_array
# %timeit sum(massive_array) # python's sum()
# %timeit np.sum(massive_array) # numpy sum()
np.mean(a4)
np.std(a4)
high_var_array = np.array([1, 100, 200, 300, 4000, 5000])
low_var_array = np.array([2, 4, 6, 8, 10])
np.var(high_var_array), np.var(low_var_array)
# average distance of array elements from the mean
np.std(high_var_array), np.std(low_var_array)
np.mean(high_var_array), np.mean(low_var_array)
# +
# %matplotlib inline
import matplotlib.pyplot as plt
plt.hist(high_var_array)
plt.show()
# -
plt.hist(low_var_array)
plt.show()
# ## Reshaping and Transposing
a2
a3
a2.shape
a3.shape
a2.reshape(2, 3, 1) * a3
# transpose
a2.T
a2.shape, a2.T.shape
# ## Dot product
np.random.seed(42)
mat1 = np.random.randint(10, size=(5, 3))
mat2 = np.random.randint(10, size=(5, 3))
mat1
mat2
# Element wise multiplication - Hadamard product
mat1 * mat2
np.dot(mat1, mat2.T)
# ## Comparision operators
a1 > a2
a1 >= a2
# ### Sorting
np.sort(a2)
| intro-to-numpy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Surface
# +
# %matplotlib notebook
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.axes_grid1 import make_axes_locatable
from mpl_toolkits.mplot3d import Axes3D
from matplotlib import cm
# +
fig, ax = plt.subplots(subplot_kw={"projection": "3d"})
x = y = np.linspace(-3,3,100)
X, Y = np.meshgrid(x, y)
Z = X**4+Y**4-16*X*Y
surf = ax.plot_surface(X,Y,Z, cmap=cm.coolwarm,
linewidth=0, antialiased=False)
ax.set_zlim(0, 200)
fig.colorbar(surf, shrink=0.5, aspect=5)
ax.set_xlabel('X Axis')
ax.set_ylabel('Y Axis')
plt.show()
# -
# ## Gradient Descent
# +
def E(u,v):
return u**4+v**4-16*u*v
eta=0.01
x=1.2; y=1.2
print (0,'\t','x=', x,'\t','y=',y,'\t', 'E=',E(x,y))
for i in range(0,30):
g=4*x**3-16*y
h=4*y**3-16*x
x=x-eta*g
y=y-eta*h
print (i+1,'\t','x=',round(x,3),'\t','y=',round(y,3),'\t','E=',round(E(x,y),3))
# -
# ## Linear Regression Revisited
# We will redo the example of multivariate-data in linear regression using gradient descent.
data = np.genfromtxt('../../data/multivar_simulated/data.csv',skip_header=1,delimiter=',')
Y = data[:,1]
X1 = data[:,2:]
O = np.ones(shape=(X.shape[0],1))
X = np.concatenate([X1,O],axis=1)
X.shape
# The error function is given by
# $$ E = \sum_{j=1}^{N} (y_j-\sum_{s=1}^{k+1} x_{js}m_{s})^2 .$$
# Write a function for $E$.
# +
#def Er(M):
# formula here
# return the result
# -
def Er(M):
F=Y-X@M
return sum(F*F)
# The gradient of $E$ is given by
# $$ \nabla E = -2 X^{\intercal}Y + 2
# X^{\intercal}XM. $$
# Write a function for $\nabla E$.
# +
#def GE(M):
# return formula here
# -
def GE(M):
return -2*X.transpose()@Y+2*X.transpose()@X@M
# Choose initial values.
# +
#eta=
#iter_num=
#M=np.array([?,?,?])
# -
eta=0.0001
iter_num=10000
M=np.array([0,0,0])
# Calculate the initial error.
Er(M)
# Run a loop for gradient descent and print the values of M and Er(M).
# +
#Write a loop here
#
#print M and Er(M)
# -
for i in range(iter_num):
M=M-eta*GE(M)
print(M, Er(M))
# Compare the result with the previous result from Linear Regression which was
#
# [ 1.78777492, -3.47899986, 6.0608333 ]
#
#
# ## Newton's Method
# +
def E(u,v):
return u**4+v**4-16*u*v
eta=1
x=1.2; y=1.2
print (0,'\t','x=', x,'\t','y=',y, '\t','E=',E(x,y))
for i in range(0,10):
d=9*x**2*y**2-16
g=(3*x**3*y**2 -8*y**3 -16*x)/d
h=(3*x**2*y**3 -8*x**3 -16*y)/d
x=x-eta*g
y=y-eta*h
print (i+1,'\t','x=', round(x,3),'\t','y=',round(y,3), '\t','E=',round(E(x,y),3))
# -
# ### Haberman's Survival Data Set
#
# https://archive.ics.uci.edu/ml/datasets/Haberman%27s+Survival
import numpy as np
import pandas as pd
from sklearn import datasets, linear_model
from sklearn.model_selection import train_test_split
from matplotlib import pyplot as plt
df=pd.read_csv("../data/haberman.data",header=None)
X=df.iloc[:,:-1].values
y=df.iloc[:,-1].values
X
# ### Scaling Features
#
# As you can see the data from various columns have various value ranges. Having the features in same range makes the algorithm to converge faster than using the features which are not scaled to be in same range.
from sklearn.preprocessing import MinMaxScaler
numcols=[0,1,2]
mms=MinMaxScaler()
X=mms.fit_transform(df[numcols])
X
# ### Splitting the data
#
# We split the data set into two parts: one for train and the other for test.
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3)
print(X_train.shape, y_train.shape)
print(X_test.shape, y_test.shape)
# We add one column consisting of ones.
n_train=len(y_train)
new_col=np.ones((n_train,1))
X_train_modified=np.append(X_train,new_col,1)
X_train_modified
n_test=len(y_test)
new_col=np.ones((n_test,1))
X_test_modified=np.append(X_test,new_col,1)
# Define the function $\sigma(x) = \dfrac {e^x}{e^x+1}= \dfrac 1 {1+e^{-x}}$.
# +
#def sigmoid(x):
# return the function
# -
def sigmoid(x):
return 1/ (1 + np.exp(-x))
# Define the error function
# $$ E (\mathbf{w}) = - \frac 1 N \sum_{n=1}^N \{ t_n \ln y_n + (1-t_n) \ln (1-y_n)\}, $$
# where $y_n=\sigma(w_1 x_{n1}+ w_2 x_{n2} + \cdots + w_k x_{nk}+w_{k+1} )$. This function will be obtained in Logistic Regression.
# +
#def Er(w):
# yn=
# return ???
# -
def Er(w):
yn=sigmoid(X_train_modified@w)
return -(1/n_train)*sum((2-y_train).reshape(n_train,1)*np.log(yn)\
+(y_train-1).reshape(n_train,1)*np.log(1-yn))
w=np.array([0,0,0,0])
Er(w)
w=[[0],[0],[0],[0]]
Er(w)
# Define the gradient of $E$.
# +
#def gradE(w):
# return the function
# -
def gradE(ww):
yw=sigmoid(X_train_modified@ww)
return (y_train+(yw-2).reshape(1,n_train))[0]@X_train_modified/n_train
gradE(w)
# Set the initial values.
# +
#w=[[?],[?],[?],[?]]
#eta=
#iter_num=
# -
w=[[0],[0],[0],[0]]
eta=0.0005
iter_num=10000
# Run a loop for gradient descent.
# +
#for i in range(iter_num):
#
# -
for i in range(iter_num):
w=w-eta*gradE(w).reshape(4,1)
if i%1000==0:
print(w,Er(w))
print(w)
# We compute the accuracy of the trained model.
y_pred=(sigmoid(X_test_modified@w).round()).reshape([1,n_test])
y_test=y_test%2
print("TraiWe compute the accuracy of the trained model.n Accuracy:", sum(y_test==y_pred[0])*100/n_test,"%")
| GradientDescent/lab/gradient-descent-with-solutions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import re
import pandas as pd
from Bio import SeqIO
from Bio.SeqRecord import SeqRecord
from Bio.SeqFeature import SeqFeature, FeatureLocation
from Bio import AlignIO
from collections import Counter
from ete3 import Tree
# +
#1. Download all coronaviruses from Vipr with human host. Select all possible metadata fields and download as a fasta
# +
#2. Name this file 'human_cov_genome.fasta' and put it in the seasonal-cov/data/ directory
# -
#3. Run add_subtype_to_virus_fastas function
add_subtype_to_virus_fastas("../data/human_cov_genome.fasta")
#4. Run fix_fasta_dateformat function
fix_fasta_dateformat("../data/human_cov_genome_annotated.fasta")
# +
#5. In terminal, run: `snakemake` from seasonal-cov/
# -
#6. Run find_unlabeled_subtypes function
find_unlabled_subtypes('../data/human_cov_full.fasta')
#7. Run separate_virus_fastas function for each virus
viruses = ['oc43', '229e', 'hku1', 'nl63']
for virus in viruses:
separate_virus_fastas(virus)
# +
#8. Toggle each virus's Snakefile to GENE = ["full"].
# In terminal, run: `snakemake` from within each virus directory
# -
#9. Run extract_genes function for each virus
# viruses = ['oc43', '229e', 'hku1', 'nl63']
viruses = ['hku1']
for virus in viruses:
extract_genes(virus)
# +
#10. Toggle each virus's Snakefile to GENE = ["replicase1ab", ...].
# In terminal, run: `snakemake` from within each virus directory
# -
#Append virus subtype to fasta fields, to ultimately create column in fauna
def add_subtype_to_virus_fastas(input_fasta):
output_fasta = str(input_fasta.replace('.fasta',''))+"_annotated.fasta"
#subtypes to look for
cov_subtypes = ['OC43', 'HKU1', 'NL63', '229E']
cov_types = {'OC43':'beta', 'HKU1':'beta', 'NL63':'alpha', '229E':'alpha'}
sequences = []
with open(input_fasta, "r") as handle:
for record in SeqIO.parse(handle, "fasta"):
new_record_list = record.description.split('|')
#Annotate subtypes
new_record_list = new_record_list+['None', 'None', 'full']
for subtype in cov_subtypes:
if subtype in record.description:
new_record_list[-3] = subtype.lower()
new_record_list[-2] = cov_types[subtype]
#New fasta fields format: 'gb-id|strain|date|host|country|virus species|subtype|type|sequence_locus'
new_record_description = '|'.join(new_record_list)
sequences.append(SeqRecord(record.seq, id=new_record_description, description=new_record_description))
SeqIO.write(sequences, output_fasta, "fasta")
def fix_fasta_dateformat(input_fasta):
with open(input_fasta, "r") as handle:
new_cov_records = []
for record in SeqIO.parse(handle, "fasta"):
date = record.description.split('|')[3].replace('_','-')
if len(date)==4:
date = date+'-XX-XX'
if len(date)==7:
date = date+'-XX'
fix_date_description = record.description.split('|')
fix_date_description[3] = date
fix_date_description.pop(2)
fix_date_description.pop(5)
fix_date_description = '|'.join(fix_date_description)
new_record = SeqRecord(record.seq, id= fix_date_description,
name= fix_date_description, description= fix_date_description)
new_cov_records.append(new_record)
SeqIO.write(new_cov_records, "../data/human_cov_full.fasta", 'fasta')
# +
#Known viruses for each subtype
known = {'oc43':['KF963229', 'KF530087', 'KF530059', 'LC506876', 'KU131570', 'LC506782', 'LC506896'],
'hku1':['KR055515', 'KF430197', 'KF686339', 'KR055516', 'MF996629', 'MH940245'],
'nl63': ['KM055607', 'KT359837', 'KM055597', 'KM055602', 'JX104161', 'KY862037', 'MF996663'],
'229e': ['JX503060', 'LC005741', 'KM055524', 'KM055568', 'KJ866103', 'GU068548', 'KY369908', 'KT359754'],
'mers': ['KJ156950', 'KT357808', 'MK129253', 'KX034094', 'KJ156890'],
'sars1':['AY345986', 'GU553363', 'JN247396', 'DQ182595'],
'sars2':['MT326086', 'MT159717', 'MT304486']
}
def find_unlabled_subtypes(input_fasta):
cov_tree = Tree('../results/tree_cov_full.nwk', format=1)
cov_metadata = pd.read_csv('../results/metadata_cov_full.tsv', delimiter = '\t').set_index('strain')
subtyped_viruses = {key: set() for key in known.keys()}
for node in cov_tree.iter_descendants("postorder"):
leafs = node.get_leaves()
descendents = [leaf.name for leaf in leafs]
for subtype in known.keys():
this_subtype = known[subtype]
remove_subtype = {key:val for key, val in known.items() if key != subtype}
other_subtypes = [item for sublist in list(remove_subtype.values()) for item in sublist]
#check if node has all known viruses of a subtype
if all(elem in descendents for elem in this_subtype):
#check that node doesn't have any viruses of other known subtypes
if not any(elem in descendents for elem in other_subtypes):
subtyped_viruses[subtype].update(descendents)
output_fasta = str(input_fasta.replace('.fasta',''))+"_subtyped.fasta"
add_newly_subtyped_to_virus_fastas(input_fasta, subtyped_viruses, output_fasta)
manually_add_subtype(output_fasta)
make_strainname(output_fasta)
def make_strainname(output_fasta):
new_strainname_records = []
with open(output_fasta, "r") as handle:
for record in SeqIO.parse(handle, "fasta"):
new_record_list = record.description.split('|')
#Make strain name from subtype, accession number, strain, and date
year = str(new_record_list[2])[0:4]
strain_name = str(new_record_list[-3])+'/'+str(new_record_list[0])+'/'+str(new_record_list[1])+'/'+year
new_record_list = [strain_name] + new_record_list
#New fasta fields format: 'strain_name|gb-id|strain|date|host|country|virus species|subtype|type|sequence_locus'
new_record_description = '|'.join(new_record_list)
new_record = SeqRecord(record.seq, id= new_record_description,
name= new_record_description, description= new_record_description)
new_strainname_records.append(new_record)
SeqIO.write(new_strainname_records, output_fasta, "fasta")
def add_newly_subtyped_to_virus_fastas(input_fasta, subtyped_viruses, output_fasta):
cov_types = {'oc43':'beta', 'hku1':'beta', 'nl63':'alpha', '229e':'alpha', 'mers':'beta', 'sars1':'beta', 'sars2':'beta'}
new_subtype_records = []
with open(input_fasta, "r") as handle:
for record in SeqIO.parse(handle, "fasta"):
new_record_list = record.description.split('|')
#Annotate subtypes
for subtype in known.keys():
strains = subtyped_viruses[subtype]
if new_record_list[0] in strains:
new_record_list[-3] = subtype
new_record_list[-2] = cov_types[subtype]
new_record_description = '|'.join(new_record_list)
new_record = SeqRecord(record.seq, id= new_record_description,
name= new_record_description, description= new_record_description)
new_subtype_records.append(new_record)
SeqIO.write(new_subtype_records, output_fasta, "fasta")
#manually label all OC43 from Ren et al paper
def manually_add_subtype(output_fasta):
ren_oc43_strains = ['JN129834','JN129835','KF572816','KF572807',
'KF572817','KF572818','KF572819', 'KF572864',
'KF572820', 'KF572842', 'KF572843', 'KF572844',
'KF572845', 'KF572846', 'KF572847', 'KF572848',
'KF572849', 'KF572850', 'KF572851', 'KF572852',
'KF572853', 'KF572854', 'KF572855', 'KF572856',
'KF572857', 'KF572858', 'KF572859', 'KF572860',
'KF572861', 'KF572862', 'KF572863', 'KF572824',
'KF572826', 'KF572827', 'KF572828', 'KF572830',
'KF572868', 'KF572872', 'KF572865', 'KF572866',
'KF572867', 'KF572870', 'KF572871', 'KF572834',
'KF572835', 'KF572836', 'KF572837', 'KF572821',
'KF572822', 'KF572823', 'KF572825', 'KF572804',
'KF572805', 'KF572806', 'KF572808', 'KF572809',
'KF572838', 'KF572810', 'KF572839', 'KF572811',
'KF572812', 'KF572813', 'KF572814', 'KF572841',
'KF572831', 'KF572832', 'KF572833']
manually_named_records = []
with open(output_fasta, "r") as handle:
for record in SeqIO.parse(handle, "fasta"):
new_record_list = record.description.split('|')
if new_record_list[0] in ren_oc43_strains:
new_record_list[-3] = 'oc43'
new_record_list[-2] = 'beta'
new_record_description = '|'.join(new_record_list)
new_record = SeqRecord(record.seq, id= new_record_description,
name= new_record_description, description= new_record_description)
manually_named_records.append(new_record)
SeqIO.write(manually_named_records, output_fasta, "fasta")
# -
def separate_virus_fastas(virus):
output_fasta = "../"+str(virus)+"/data/"+str(virus)+"_full.fasta"
sequences = []
with open("../data/human_cov_full_subtyped.fasta", "r") as handle:
for record in SeqIO.parse(handle, "fasta"):
if virus in record.description:
#Fix date formatting
date = record.description.split('|')[3].replace('_','-')
if len(date)==4:
date = date+'-XX-XX'
if len(date)==7:
date = date+'-XX'
new_record_list = record.description.split('|')
new_record_list[3] = date
new_record_description = '|'.join(new_record_list)
#Fasta fields format: 'gb-id|strain|segment|date|host|country|subtype|virus species'
sequences.append(SeqRecord(record.seq, id=new_record_description, description=new_record_description))
SeqIO.write(sequences, output_fasta, "fasta")
#Gene positions for each virus
def get_virus_genes(virus):
if virus == 'nl63':
genes_dict = {'replicase1ab':"replicase polyprotein 1ab", 'spike':"spike protein", 'protein3':"protein 3",
'envelope':"envelope protein", 'membrane':"membrane protein", 'nucleocapsid':"nucleocapsid protein",
's1':'spike_subdomain1', 's2':'spike_subdomain2', 'rdrp':'rna_depedent_rna_polymerase'}
elif virus == '229e':
genes_dict = {'replicase1ab':"replicase polyprotein 1ab", 'replicase1a': "replicase polyprotein 1a", 'spike':"surface glycoprotein",
'protein4a':"4a protein", 'protein4b':"4b protein",
'envelope':"envelope protein", 'membrane':"membrane protein", 'nucleocapsid':"nucleocapsid protein",
's1':'spike_subdomain1', 's2':'spike_subdomain2', 'rdrp':'rna_depedent_rna_polymerase'}
elif virus == 'hku1':
genes_dict = {'replicase1ab':"orf1ab polyprotein", 'he':"hemagglutinin-esterase glycoprotein",
'spike':"spike glycoprotein", 'nonstructural4':"non-structural protein",
'envelope':"small membrane protein", 'membrane':"membrane glycoprotein",
'nucleocapsid':"nucleocapsid phosphoprotein", 'nucleocapsid2':"nucleocapsid phosphoprotein 2",
's1':'spike_subdomain1', 's2':'spike_subdomain2', 'rdrp':'rna_depedent_rna_polymerase'}
elif virus == 'oc43':
genes_dict = {'replicase1ab':"replicase polyprotein", 'nonstructural2a':"NS2a protein",
'he':"HE protein", 'spike':"S protein", 'nonstructural2':"NS2 protein",
'envelope':"NS3 protein", 'membrane':"M protein",
'nucleocapsid':"N protein", 'n2protein':"N2 protein",
's1':'spike_subdomain1', 's2':'spike_subdomain2', 'rdrp':'rna_depedent_rna_polymerase'}
return genes_dict
# +
#first 6 aas of each domain
#from uniprot: NL63 (Q6Q1S2), 229e(P15423), oc43 (P36334), hku1 (Q0ZME7)
s1_domains = {'nl63': 'FFTCNS', '229e': 'CQTTNG', 'oc43': 'AVIGDL', 'hku1': 'AVIGDF'}
s2_domains = {'nl63': 'SSDNGI', '229e': 'IIAVQP', 'oc43': 'AITTGY', 'hku1': 'SISASY'}
rdrp_domains_start = {'oc43': 'SKDTNF', '229e': 'SFDNSY', 'nl63': 'SVDISY', 'hku1': 'SKDLNF'}
rdrp_domains_end = {'oc43': 'RSAVMQ', '229e': 'KSTVLQ', 'nl63': 'NSTILQ', 'hku1': 'KSAVMQ'}
# -
def get_s1_coords(virus):
spike_reference = '../'+str(virus)+'/config/'+str(virus)+'_spike_reference.gb'
with open(spike_reference, "r") as handle:
for record in SeqIO.parse(handle, "genbank"):
nt_seq = record.seq
aa_seq = record.seq.translate()
s1_regex = re.compile(f'{s1_domains[virus]}.*(?={s2_domains[virus]})')
s1_aa = s1_regex.search(str(aa_seq)).group()
s1_aa_coords = [(aa.start(0), aa.end(0)) for aa in re.finditer(s1_regex, str(aa_seq))][0]
s1_nt_coords = [s1_aa_coords[0]*3, s1_aa_coords[1]*3]
return s1_nt_coords
def get_s2_coords(virus):
spike_reference = '../'+str(virus)+'/config/'+str(virus)+'_spike_reference.gb'
with open(spike_reference, "r") as handle:
for record in SeqIO.parse(handle, "genbank"):
nt_seq = record.seq
aa_seq = record.seq.translate()
s2_regex = re.compile(f'{s2_domains[virus]}.*')
s2_aa = s2_regex.search(str(aa_seq)).group()
s2_aa_coords = [(aa.start(0), aa.end(0)) for aa in re.finditer(s2_regex, str(aa_seq))][0]
s2_nt_coords = [s2_aa_coords[0]*3, s2_aa_coords[1]*3]
return s2_nt_coords
def get_rdrp_coords(virus):
replicase_reference = '../'+str(virus)+'/config/'+str(virus)+'_replicase1ab_reference.gb'
with open(replicase_reference, "r") as handle:
for record in SeqIO.parse(handle, "genbank"):
nt_seq = record.seq
aa_seq = record.seq.translate()
rdrp_regex = re.compile(f'{rdrp_domains_start[virus]}.*{rdrp_domains_end[virus]}')
rdrp_aa = rdrp_regex.search(str(aa_seq)).group()
rdrp_aa_coords = [(aa.start(0), aa.end(0)) for aa in re.finditer(rdrp_regex, str(aa_seq))][0]
rdrp_nt_coords = [rdrp_aa_coords[0]*3, rdrp_aa_coords[1]*3]
return rdrp_nt_coords
#Gene positions for each virus
def get_gene_position_test(virus, gene, sequence):
genes_dict = get_virus_genes(virus)
for seq_record in SeqIO.parse("../"+str(virus)+"/config/"+str(virus)+"_full_reference.gb", "genbank"):
for feature in seq_record.features:
if feature.type == 'CDS':
if gene != 's1' and gene != 's2' and gene != 'rdrp':
if feature.qualifiers['product'] == [genes_dict[gene]]:
gene_nt = feature.location.extract(sequence)
if gene == 's1':
if feature.qualifiers['product'] == [genes_dict['spike']]:
s1_nt_coords = get_s1_coords(virus)
gene_nt = feature.location.extract(sequence)[s1_nt_coords[0]:s1_nt_coords[1]]
if gene == 's2':
if feature.qualifiers['product'] == [genes_dict['spike']]:
s2_nt_coords = get_s2_coords(virus)
gene_nt = feature.location.extract(sequence)[s2_nt_coords[0]:s2_nt_coords[1]]
if gene == 'rdrp':
if feature.qualifiers['product'] == [genes_dict['replicase1ab']]:
rdrp_nt_coords = get_rdrp_coords(virus)
gene_nt = feature.location.extract(sequence)[rdrp_nt_coords[0]:rdrp_nt_coords[1]]
return gene_nt
#From aligned .fasta, extract just the portion of the genome encoding each gene
def extract_genes(virus):
aligned_fasta = "../"+str(virus)+"/results/aligned_"+str(virus)+"_full.fasta"
original_fasta = "../"+str(virus)+"/data/"+str(virus)+"_full.fasta"
genes_dict = get_virus_genes(virus)
genes = [k for k,v in genes_dict.items()]
for gene in genes:
output_fasta = "../"+str(virus)+"/data/"+str(virus)+"_"+str(gene)+".fasta"
gene_sequences = {}
with open(aligned_fasta, "r") as handle:
alignment = SeqIO.parse(handle, "fasta")
for aligned_record in alignment:
gene_nt = get_gene_position_test(virus, gene, aligned_record.seq)
gene_nt_str = str(gene_nt)
#Throw out sequences that don't cover gene
num_unaligned_gene = Counter(gene_nt_str)['N']
if num_unaligned_gene < (len(gene_nt)/2):
gene_sequences[aligned_record.id] = gene_nt
gene_entries = []
with open(original_fasta, "r") as handle_2:
metadata = SeqIO.parse(handle_2, "fasta")
for meta_record in metadata:
strain_accession = meta_record.id.split('|')[0]
if str(strain_accession) in gene_sequences.keys():
gene_record = SeqRecord(gene_sequences[strain_accession], id=meta_record.id, description=meta_record.id)
gene_entries.append(gene_record)
SeqIO.write(gene_entries, output_fasta, "fasta")
| data-wrangling/postdownload_formatting_for_rerun.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import csv
import gzip
import random
import sqlite3
import sys
import time
import itertools
import json
import numpy as np
import colorspacious
from tensorflow.keras.models import Model
from tensorflow.keras.layers import Input, Dense, SeparableConv1D, Activation
import tensorflow as tf
import sklearn.utils
import matplotlib.pyplot as plt
import numba
# Force CPU operation for deterministic and reproducible results
# Disable parallelism per https://github.com/NVIDIA/framework-determinism/blob/28091e4fb1483685fc78b7ab844a5ae6ddf55a14/README.md#cpu
tf.config.set_visible_devices([], "GPU")
tf.config.threading.set_intra_op_parallelism_threads(1)
tf.config.threading.set_inter_op_parallelism_threads(1)
# Needed for deterministic results
SEED = 567687
random.seed(SEED)
np.random.seed(SEED)
tf.random.set_seed(SEED)
with open("top-sets.json") as infile:
TOP_SETS_RGB = json.load(infile)
TOP_SETS_RGB = {int(i): TOP_SETS_RGB[i] for i in TOP_SETS_RGB}
ALL_NUM_COLORS = TOP_SETS_RGB.keys()
DB_FILE = "../survey-results/results.db"
npz = np.load("../color-name-model/colornamemodel.npz")
COLOR_NAMES = list(npz["names"])
BCT_IDX = npz["bct_idxs"]
COLOR_NAME_IDX = npz["name_idxs"]
ENSEMBLE_COUNT = 100
# ## Color conversion and sorting functions
#
# Data are all stored as 8-bit RGB values and need to be converted to CAM02-UCS.
#
# We also need to be able to sort along the three CAM02-UCS axes.
# +
def to_jab(color):
"""
Convert hex color code (without `#`) to CAM02-UCS.
"""
rgb = [(int(i[:2], 16), int(i[2:4], 16), int(i[4:], 16)) for i in color]
jab = [colorspacious.cspace_convert(i, "sRGB255", "CAM02-UCS") for i in rgb]
return np.array(jab)
def sort_colors_by_j(colors):
"""
Sorts colors by CAM02-UCS J' axis.
"""
return colors[np.lexsort(colors[:, ::-1].T, 0)]
def sort_colors_by_a(colors):
"""
Sorts colors by CAM02-UCS a' axis.
"""
return colors[np.argsort(colors[:, ::-1].T[1])]
def sort_colors_by_b(colors):
"""
Sorts colors by CAM02-UCS b' axis.
"""
return colors[np.argsort(colors[:, ::-1].T[2])]
# -
# ## Data loading
#
# Survey data are loaded from a SQLite database, and the complete set of color sets is loaded from a text file. The 8-bit RGB values are converted to CAM02-UCS and sorted along the three CAM02-UCS axes.
# +
# Load survey data
cycle_data = {}
cycle_targets = {}
min_count = 1e10
conn = sqlite3.connect(DB_FILE)
c = conn.cursor()
for num_colors in ALL_NUM_COLORS:
count = 0
cycle_data[num_colors] = []
cycle_targets[num_colors] = []
for row in c.execute(
f"SELECT c1, c2, o, cp, sp FROM picks WHERE length(c1) = {num_colors * 7 - 1}"
):
count += 1
orders = [[int(c) for c in o] for o in row[2].split(",")]
# Convert to Jab [CAM02-UCS based]
jab1 = to_jab(row[0].split(","))
jab2 = to_jab(row[1].split(","))
# Add cycle data
for i in range(4):
if i != row[3] - 1:
cycle_data[num_colors].append(
np.array((jab1[orders[row[3] - 1]], jab2[orders[i]])).flatten()
)
cycle_targets[num_colors].append(0)
cycle_data[num_colors] = np.array(cycle_data[num_colors])
cycle_targets[num_colors] = np.array(cycle_targets[num_colors])
min_count = min(min_count, count)
print(num_colors, count)
conn.close()
# -
# Convert color set data
TOP_SETS = {}
for num_colors in ALL_NUM_COLORS:
TOP_SETS[num_colors] = to_jab(TOP_SETS_RGB[num_colors]).flatten()
# Best color set
orderings = {}
orderings_rgb = {}
for num_colors in ALL_NUM_COLORS:
orderings[num_colors] = np.array(
list(itertools.permutations(TOP_SETS[num_colors].reshape(-1, 3)))
).reshape(-1, 3 * num_colors)
orderings_rgb[num_colors] = np.array(
list(itertools.permutations(TOP_SETS_RGB[num_colors]))
)
def construct_network(num_colors):
#
# Construct network
#
conv_size = 5
layer1 = Dense(units=5, activation="elu", name="l1")
layer2 = Dense(units=5, activation="elu", name="l2")
layer3 = SeparableConv1D(5, conv_size, padding="same", activation="elu", name="l3")
layer4 = SeparableConv1D(3, conv_size, padding="same", activation="elu", name="l4")
layer5 = SeparableConv1D(1, conv_size, padding="same", activation="elu", name="l5")
# Create network
input_a = Input(shape=(3 * num_colors,))
inputs_a = [input_a[:, i * 3 : (i + 1) * 3] for i in range(num_colors)]
# Share layers between colors
x_a = [layer1(i / 100) for i in inputs_a]
x_a = [layer2(i) for i in x_a]
# Combine colors into sets
x_a = tf.keras.layers.concatenate(
[tf.keras.backend.expand_dims(i, 1) for i in x_a], axis=1
)
# Share layers between color sets
x_a = layer3(x_a)
x_a = layer4(x_a)
x_a = layer5(x_a)
# Average outputs
x_a = tf.math.reduce_mean(x_a, axis=1)
# Final non-linear activation
layer_nm1 = Activation("sigmoid", name=f"score{num_colors}")
x_a = layer_nm1(x_a)
# For evaluation
return Model(inputs=input_a, outputs=x_a)
scoring_model = {nc: construct_network(nc) for nc in ALL_NUM_COLORS}
# ## Evaluate model accuracy
#
# Scores are compared to calculate model accuracy. All data are used (no train / test split).
#
# As the ten-color data were not used during design, hyperparameter tuning, or training, they serve as a validation set. These data were only evaluated after the model was finalized.
survey_scores_a = {nc: [] for nc in ALL_NUM_COLORS}
survey_scores_b = {nc: [] for nc in ALL_NUM_COLORS}
for nc in ALL_NUM_COLORS:
for i in range(ENSEMBLE_COUNT):
with gzip.open(f"weights/cycle_model_weights_{i:03d}.h5.gz", "rb") as infile:
infile.endswith = lambda x: x == ".h5" # Monkey-patch for format detection
scoring_model[nc].load_weights(infile, by_name=True)
survey_scores_a[nc].append(
scoring_model[nc].predict(cycle_data[nc][:, : nc * 3]).flatten()
)
survey_scores_b[nc].append(
scoring_model[nc].predict(cycle_data[nc][:, nc * 3 :]).flatten()
)
survey_scores_a[nc] = np.array(survey_scores_a[nc])
survey_scores_b[nc] = np.array(survey_scores_b[nc])
for nc in ALL_NUM_COLORS:
acc = np.mean((survey_scores_a[nc] > survey_scores_b[nc]) ^ cycle_targets[nc])
print(f"accuracy {nc:2d}: {acc:.5f}")
# ## Predict scores for full set of color sets
#
# A score is predicted for each color set in the full set of color sets. Furthermore, the variability within the ensemble is also evaluated.
# Score color cycles
scores = {nc: [] for nc in ALL_NUM_COLORS}
for nc in ALL_NUM_COLORS:
for i in range(ENSEMBLE_COUNT):
with gzip.open(f"weights/cycle_model_weights_{i:03d}.h5.gz", "rb") as infile:
infile.endswith = lambda x: x == ".h5" # Monkey-patch for format detection
scoring_model[nc].load_weights(infile, by_name=True)
scores[nc].append(scoring_model[nc].predict(orderings[nc]).flatten())
scores[nc] = np.array(scores[nc])
cycle_scores = {}
for nc in ALL_NUM_COLORS:
bootstrap = np.array(
[np.mean(sklearn.utils.resample(scores[nc]), axis=0) for _ in range(1000)]
)
cycle_scores[f"mean{nc:02d}"] = scores[nc].mean(axis=0)
cycle_scores[f"error{nc:02d}"] = bootstrap.std(axis=0)
# ## Calculate final cycle scores
#
# The final cycle scores combine the machine learning derived scores with a perceptual distance score and a luminance distance score. These scores are calculated to favor cycles that include easier to differentiate colors at the beginning of the cycle, both for color and grayscale. Additionally, only cycles that do not contain two colors in the first half of the cycle that would be identified by the same basic color term are considered.
top_cycles = {}
def to_jab_cvd(color):
rgb255 = [(int(i[:2], 16), int(i[2:4], 16), int(i[4:], 16)) for i in color]
rgb = [
(int(i[:2], 16) / 255, int(i[2:4], 16) / 255, int(i[4:], 16) / 255)
for i in color
]
jab = [colorspacious.cspace_convert(i, "sRGB1", "CAM02-UCS") for i in rgb]
jab_deut = [
colorspacious.cspace_convert(
i,
{"name": "sRGB1+CVD", "cvd_type": "deuteranomaly", "severity": 100},
"CAM02-UCS",
)
for i in rgb
]
jab_prot = [
colorspacious.cspace_convert(
i,
{"name": "sRGB1+CVD", "cvd_type": "protanomaly", "severity": 100},
"CAM02-UCS",
)
for i in rgb
]
jab_trit = [
colorspacious.cspace_convert(
i,
{"name": "sRGB1+CVD", "cvd_type": "tritanomaly", "severity": 100},
"CAM02-UCS",
)
for i in rgb
]
return (
np.array(rgb255),
np.array(jab),
np.array(jab_deut),
np.array(jab_prot),
np.array(jab_trit),
)
@numba.njit
def cam02de(c1, c2):
diff = np.abs(c1 - c2)
return np.sqrt(np.sum(diff * diff))
BCT_IDX_WHITE = COLOR_NAMES.index("white")
@numba.njit
def score_permuations(
permutations, set_rgb, set_jab, set_jab_deut, set_jab_prot, set_jab_trit
):
unique_name_count = len(
set([BCT_IDX[i[0] + i[1] * 256 + i[2] * 256 ** 2] for i in set_rgb[1:]])
)
permutation_scores = []
for order in permutations:
# Score cycle by calculating minimum perceptual distance as each color is added and averaging distances
dists = []
color_names = [BCT_IDX_WHITE]
dist = 1000
J_dist = 1000
name_score = True
for i in range(1, len(set_jab)):
j = order[i - 1] + 1
for k in [0] + list(order[: i - 1] + 1):
dist = min(dist, cam02de(set_jab[j], set_jab[k]))
dist = min(dist, cam02de(set_jab_deut[j], set_jab_deut[k]))
dist = min(dist, cam02de(set_jab_prot[j], set_jab_prot[k]))
dist = min(dist, cam02de(set_jab_trit[j], set_jab_trit[k]))
J_dist = min(J_dist, abs(set_jab[j][0] - set_jab[k][0]))
color_names.append(
BCT_IDX[set_rgb[j, 0] + set_rgb[j, 1] * 256 + set_rgb[j, 2] * 256 ** 2]
)
if (
i < len(set_jab) - 1
and i <= unique_name_count
and len(set(color_names)) < len(color_names)
):
name_score = False
break
dists.append(dist * J_dist)
if name_score:
permutation_scores.append(
np.mean(np.array(dists, dtype=np.float64))
) # Order score (higher is better)
else:
permutation_scores.append(
0.0
) # Duplicate color name in first half of cycle
return np.array(permutation_scores)
# ### Six colors
set_rgb, set_jab, set_jab_deut, set_jab_prot, set_jab_trit = to_jab_cvd(
["ffffff"] + TOP_SETS_RGB[6]
)
permutations = np.array(list(itertools.permutations(np.arange(6))))
scores_mean = np.mean(scores[6], axis=0)
with_same_first_color = permutations[:, 0] == permutations[np.argmax(scores_mean)][0]
permutations = permutations[with_same_first_color]
scores_mean = scores_mean[with_same_first_color]
permutation_scores = score_permuations(
permutations, set_rgb, set_jab, set_jab_deut, set_jab_prot, set_jab_trit
)
print(np.sum(permutation_scores > 0))
cycle_scores["first06"] = with_same_first_color
cycle_scores["access06"] = permutation_scores.astype(np.float32)
plt.scatter(scores_mean, permutation_scores)
tallies6 = permutation_scores * scores_mean
np.where(tallies6 == np.max(tallies6))
# +
idx = np.where(tallies6 == np.sort(np.unique(tallies6))[-1])[0][0]
n = len(orderings_rgb[6][0])
fig, ax = plt.subplots(figsize=(3, n * 0.5))
# Get height and width
w, Y = fig.get_dpi() * fig.get_size_inches()
h = Y / (n + 1)
for i, name in enumerate(orderings_rgb[6][with_same_first_color][idx]):
row = i
y = Y - (row * h) - h
xi_line = w * (0.05)
xf_line = w * (0.25)
xi_text = w * (0.3)
ax.text(
xi_text,
y,
"#" + name,
fontsize=(h * 0.6),
horizontalalignment="left",
verticalalignment="center",
)
ax.hlines(y + h * 0.1, xi_line, xf_line, color="#" + name, linewidth=(h * 0.6))
ax.set_xlim(0, w)
ax.set_ylim(0, Y)
ax.set_axis_off()
fig.subplots_adjust(left=0, right=1, top=1, bottom=0, hspace=0, wspace=0)
plt.show()
# -
tmp = orderings_rgb[6][with_same_first_color][idx]
top_cycles[6] = list(tmp)
print("Names:")
for i in range(6):
i = COLOR_NAME_IDX[
int(tmp[i][0:2], 16)
+ int(tmp[i][2:4], 16) * 256
+ int(tmp[i][4:], 16) * 256 ** 2
]
print(i, COLOR_NAMES[i])
# ### Eight colors
set_rgb, set_jab, set_jab_deut, set_jab_prot, set_jab_trit = to_jab_cvd(
["ffffff"] + TOP_SETS_RGB[8]
)
permutations = np.array(list(itertools.permutations(np.arange(8))))
scores_mean = np.mean(scores[8], axis=0)
with_same_first_color = permutations[:, 0] == permutations[np.argmax(scores_mean)][0]
permutations = permutations[with_same_first_color]
scores_mean = scores_mean[with_same_first_color]
permutation_scores = score_permuations(
permutations, set_rgb, set_jab, set_jab_deut, set_jab_prot, set_jab_trit
)
print(np.sum(permutation_scores > 0))
cycle_scores["first08"] = with_same_first_color
cycle_scores["access08"] = permutation_scores.astype(np.float32)
plt.scatter(scores_mean, permutation_scores)
tallies8 = permutation_scores * scores_mean
np.where(tallies8 == np.max(tallies8))
# +
idx = np.where(tallies8 == np.sort(np.unique(tallies8))[-1])[0][0]
n = len(orderings_rgb[8][0])
fig, ax = plt.subplots(figsize=(3, n * 0.5))
# Get height and width
w, Y = fig.get_dpi() * fig.get_size_inches()
h = Y / (n + 1)
for i, name in enumerate(orderings_rgb[8][with_same_first_color][idx]):
row = i
y = Y - (row * h) - h
xi_line = w * (0.05)
xf_line = w * (0.25)
xi_text = w * (0.3)
ax.text(
xi_text,
y,
"#" + name,
fontsize=(h * 0.6),
horizontalalignment="left",
verticalalignment="center",
)
ax.hlines(y + h * 0.1, xi_line, xf_line, color="#" + name, linewidth=(h * 0.6))
ax.set_xlim(0, w)
ax.set_ylim(0, Y)
ax.set_axis_off()
fig.subplots_adjust(left=0, right=1, top=1, bottom=0, hspace=0, wspace=0)
plt.show()
# -
tmp = orderings_rgb[8][with_same_first_color][idx]
top_cycles[8] = list(tmp)
print("Names:")
for i in range(8):
i = COLOR_NAME_IDX[
int(tmp[i][0:2], 16)
+ int(tmp[i][2:4], 16) * 256
+ int(tmp[i][4:], 16) * 256 ** 2
]
print(i, COLOR_NAMES[i])
# ### Ten colors
set_rgb, set_jab, set_jab_deut, set_jab_prot, set_jab_trit = to_jab_cvd(
["ffffff"] + TOP_SETS_RGB[10]
)
permutations = np.array(list(itertools.permutations(np.arange(10))))
scores_mean = np.mean(scores[10], axis=0)
with_same_first_color = permutations[:, 0] == permutations[np.argmax(scores_mean)][0]
permutations = permutations[with_same_first_color]
scores_mean = scores_mean[with_same_first_color]
permutation_scores = score_permuations(
permutations, set_rgb, set_jab, set_jab_deut, set_jab_prot, set_jab_trit
)
print(np.sum(permutation_scores > 0))
cycle_scores["first10"] = with_same_first_color
cycle_scores["access10"] = permutation_scores.astype(np.float32)
plt.scatter(scores_mean, permutation_scores)
tallies10 = permutation_scores * scores_mean
np.where(tallies10 == np.max(tallies10))
# +
idx = np.where(tallies10 == np.sort(np.unique(tallies10))[-1])[0][0]
n = len(orderings_rgb[10][0])
fig, ax = plt.subplots(figsize=(3, n * 0.5))
# Get height and width
w, Y = fig.get_dpi() * fig.get_size_inches()
h = Y / (n + 1)
for i, name in enumerate(orderings_rgb[10][with_same_first_color][idx]):
row = i
y = Y - (row * h) - h
xi_line = w * (0.05)
xf_line = w * (0.25)
xi_text = w * (0.3)
ax.text(
xi_text,
y,
"#" + name,
fontsize=(h * 0.6),
horizontalalignment="left",
verticalalignment="center",
)
ax.hlines(y + h * 0.1, xi_line, xf_line, color="#" + name, linewidth=(h * 0.6))
ax.set_xlim(0, w)
ax.set_ylim(0, Y)
ax.set_axis_off()
fig.subplots_adjust(left=0, right=1, top=1, bottom=0, hspace=0, wspace=0)
plt.show()
# -
tmp = orderings_rgb[10][with_same_first_color][idx]
top_cycles[10] = list(tmp)
print("Names:")
for i in range(10):
i = COLOR_NAME_IDX[
int(tmp[i][0:2], 16)
+ int(tmp[i][2:4], 16) * 256
+ int(tmp[i][4:], 16) * 256 ** 2
]
print(i, COLOR_NAMES[i])
# ### Save results
np.savez_compressed('cycle-scores.npz', **cycle_scores)
with open('top-cycles.json', 'w') as outfile:
json.dump(top_cycles, outfile)
| aesthetic-models/cycle-evaluation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Introduction to Data Science
#
# ### Introduction to Pandas
# +
#import pylab
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import os
import pathlib
# %matplotlib inline
# #%matplotlib notebook
# -
# ### Pandas Data Structures: Series
obj = pd.Series([4, 7, -5, 3, 5])
obj
obj.values
obj.index
obj.index = ['Bob', 'Steve', 'Jeff', 'Ryan', 'Fernie']
obj
obj['Bob']
obj2 = pd.Series([4, 7, -5, 3], index=['d', 'b', 'a', 'c'])
obj2
obj2['c']
obj2[['c', 'a', 'd']]
obj2[obj2 < 0]
obj2 * 2
np.exp(obj2)
sdata = {'Ohio': 35000, 'Texas': 71000, 'Oregon': 16000, 'Utah': 5000}
obj3 = pd.Series(sdata)
obj3
states = ['California', 'Ohio', 'Oregon', 'Texas']
obj4 = pd.Series(sdata, index=states)
obj4
pd.isnull(obj4)
pd.notnull(obj4)
obj3 + obj4
obj4.name = 'population'
obj4.index.name = 'state'
obj4
# ### Pandas Data Structures: Dataframe
data = {'state': ['Ohio', 'Ohio', 'Ohio', 'Nevada', 'Nevada'],'year': [2000, 2001, 2002, 2001, 2002],'pop': [1.5, 1.7, 3.6, 2.4, 2.9]}
frame = pd.DataFrame(data)
frame
frame['pop']
pd.DataFrame(data, columns=['year', 'state', 'pop'])
frame2 = pd.DataFrame(data, columns=['year', 'state', 'pop', 'debt'],index=['one', 'two', 'three', 'four', 'five'])
frame2
frame2['nova'] = 13
frame2
frame2.nova = 23
frame2
frame2.columns
frame2['state']
frame2.state
#frame2.loc['three']
frame2.loc['three','state']
frame2['debt'] = 16.5
frame2
frame2['debt'] = np.arange(5.)
frame2
val = pd.Series([-1.2, -1.5, -1.7], index=['two', 'four', 'five'])
frame2['debt'] = val
frame2
frame2['eastern'] = frame2.state == 'Ohio'
frame2
del frame2['eastern']
frame2.columns
transpose = frame2.pivot(index= 'year', columns='state', values='pop')
transpose
pop = {'Nevada': {2001: 2.4, 2002: 2.9},'Ohio': {2000: 1.5, 2001: 1.7, 2002: 3.6}}
frame3 = pd.DataFrame(pop)
frame3
frame3.T
pd.DataFrame(pop, index=[2001, 2002, 2003])
pdata = {'Ohio': frame3['Ohio'][:-1],'Nevada': frame3['Nevada'][:2]}
pd.DataFrame(pdata)
frame3.index.name = 'year'; frame3.columns.name = 'state'
frame3
pop = {'Nevada': {2001: 2.4, 2002: 2.9},'Ohio': {2000: 1.5, 2001: 1.7, 2002: 3.6}}
frame4 = pd.DataFrame(pop)
frame4
frame4.loc[2000,'Nevada'] = 2
frame4
frame5 = pd.concat([frame4, frame4])
frame5
frame5.drop_duplicates(['Nevada'])
dates = pd.date_range("20160101", periods=10)
data = np.random.random((10,3))
column_names = ['Column1', 'Column2', 'Column3']
df = pd.DataFrame(data, index=dates, columns=column_names)
df.head(10)
df[1:3]
df['20160104':'20160107']
df.loc['20160101':'20160102',['Column1','Column3']]
df.iloc[3:5, 0:2]
df.describe()
df.sort_index(axis=0, ascending=False,) # inplace=True)
df.sort_values(by='Column2')
# +
dates1 = pd.date_range("20160101", periods=6)
data1 = np.random.random((6,2))
column_names1 = ['ColumnA', 'ColumnB']
dates2 = pd.date_range("20160101", periods=7)
data2 = np.random.random((7,2))
column_names2 = ['ColumnC', 'ColumnD']
df1 = pd.DataFrame(data1, index=dates1, columns=column_names1)
df2 = pd.DataFrame(data2, index=dates2, columns=column_names2)
# -
df1.head()
df2.head()
df1.join(df2)
# +
df3 = df1.join(df2)
# add a column to df to group on
df3['ProfitLoss'] = pd.Series(['Profit',
'Loss',
'Profit',
'Profit',
'Profit',
'Loss',
'Profit',
'Profit',
'Profit',
'Loss'], index=dates)
# -
df3.head()
df3.groupby('ProfitLoss').mean()
# idmin & idmax
df3['ColumnA'].idxmax()
df3['ColumnA'].idxmin()
# ne()
df = pd.DataFrame()
df['x'] = [0,0,0,0,0,0,1,2,3,4,5,6,7]
df['x'].ne(0)
df['x'].ne(0).idxmax()
df['x'].nsmallest(3)
df.nlargest(3, 'x')
| notebooks/Intro_Pandas.ipynb |