code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # 3. I/O handling
import numpy as np
import torch
# set logger and enforce reproducibility
from GPErks.log.logger import get_logger
from GPErks.utils.random import set_seed
log = get_logger()
seed = 8
set_seed(seed) # reproducible sampling
#
# <br/>
#
# **2D function example**: Currin et al. (1988)
#
# $f(x) = \left[1 - \exp{\left(1 - \dfrac{1}{2 x_2}\right)}\right]\,\left(\dfrac{2300 x_{1}^3 + 1900 x_{1}^2 + 2092 x_{1} + 60}{100 x_{1}^3 + 500 x_{1}^2 + 4 x_{1} + 20}\right)$
#
# <br/>
#
# function to learn (normally a high-dimensional, expensive deterministic model)
from GPErks.utils.test_functions import currin_exp
f = lambda X: np.array([currin_exp(x) for x in X])
D = 2
# build dataset
from GPErks.gp.data.dataset import Dataset
dataset = Dataset.build_from_function(
f,
D,
n_train_samples=20,
n_test_samples=25,
design="lhs",
seed=seed,
)
# choose likelihood
from gpytorch.likelihoods import GaussianLikelihood
likelihood = GaussianLikelihood()
# choose mean function
from gpytorch.means import LinearMean
mean_function = LinearMean(input_size=dataset.input_size)
# choose kernel
from gpytorch.kernels import RBFKernel, ScaleKernel
kernel = ScaleKernel(RBFKernel(ard_num_dims=dataset.input_size))
# choose metrics
from torchmetrics import MeanSquaredError, R2Score
metrics = [MeanSquaredError(), R2Score()]
# define experiment
from GPErks.gp.experiment import GPExperiment
experiment = GPExperiment(
dataset,
likelihood,
mean_function,
kernel,
n_restarts=3,
metrics=metrics,
seed=seed # reproducible training
)
# dump experiment in config file
config_file = "./example_3.ini"
experiment.save_to_config_file(config_file)
# +
# load experiment from config file
del experiment
from GPErks.gp.experiment import load_experiment_from_config_file
experiment = load_experiment_from_config_file(
config_file,
dataset # notice that we still need to provide the dataset used!
)
# -
# choose training options: device + optimizer
device = "cpu"
optimizer = torch.optim.Adam(experiment.model.parameters(), lr=0.1)
# train model
from GPErks.train.emulator import GPEmulator
emulator = GPEmulator(experiment, device)
# +
# snapshotting
from GPErks.serialization.path import posix_path
from GPErks.train.snapshot import (
EveryEpochSnapshottingCriterion,
EveryNEpochsSnapshottingCriterion,
NeverSaveSnapshottingCriterion
)
import os
snapshot_dir = posix_path(os.getcwd(), "snapshot", "example_3")
train_restart_template = "restart_{restart}"
train_epoch_template = "epoch_{epoch}.pth"
snapshot_file = train_epoch_template
snpc = EveryEpochSnapshottingCriterion(
posix_path(snapshot_dir, train_restart_template),
snapshot_file
)
# training
emulator.train(optimizer, snapshotting_criterion=snpc)
# -
# inference on stored test set
from GPErks.perks.inference import Inference
inference = Inference(emulator)
inference.summary()
# +
# loading emulator
best_model_file = posix_path(
snapshot_dir,
"best_model.pth"
)
best_model_state = torch.load(best_model_file, map_location=torch.device(device))
emulator1 = GPEmulator(experiment, device)
emulator1.model.load_state_dict(best_model_state)
# -
inference = Inference(emulator1)
inference.summary()
|
notebooks/example_3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import ase.io as io
from ase.build import cut
from ase.spacegroup import crystal
a = 9.04
skutterudite = crystal(('Co', 'Sb'),
basis=[(0.25, 0.25, 0.25), (0.0, 0.335, 0.158)],
spacegroup=204,
cellpar=[a, a, a, 90, 90, 90])
# Create a new atoms instance with Co at origo including all atoms on the
# surface of the unit cell
cosb3 = cut(skutterudite, origo=(0.25, 0.25, 0.25), extend=1.01)
# Define the atomic bonds to show
bondatoms = []
symbols = cosb3.get_chemical_symbols()
for i in range(len(cosb3)):
for j in range(i):
if (symbols[i] == symbols[j] == 'Co' and
cosb3.get_distance(i, j) < 4.53):
bondatoms.append((i, j))
elif (symbols[i] == symbols[j] == 'Sb' and
cosb3.get_distance(i, j) < 2.99):
bondatoms.append((i, j))
# Create nice-looking image using povray
io.write('spacegroup-cosb3.pov', cosb3,
transparent=True,
display=False,
run_povray=True,
camera_type='perspective',
background=(0, 0, 0, 1.),
canvas_width=1200,
radii=0.4,
rotation='90y',
bondlinewidth=0.07,
bondatoms=bondatoms)
# -
# # POVRay Settings
#
# To get transparency modify pov ray .pov file to include
#
# ```
# background {srgbt <0.00, 0.00, 0.00, 1.00>}
# ```
#
# and the .ini file to contain
#
# ```
# Output_Alpha=True
# ```
#
# You can also adjust the .pov file to change the distance to the spotlight source, and adjust the diffuse lighting conditions on `ase3`.
2400 / 1.5664649389
# +
from arpes.utilities.bz import hex_cell
a = 5.761
c = 12.178
alloy = crystal(
symbols=['S', 'Nb', 'Fe', 'Nb'],
basis=[(1./3, 0, 3./8,), (1./3, 2./3, 0), (1./3, 2./3, 1./4), (0, 0, 0)],
cell=hex_cell(a=a, c=c),
spacegroup=182,
)
fenbs2 = cut(alloy, origo=(0, 0, 0.28), extend=(3.01, 3.01, 6.91), clength=2.1, tolerance=1)
# Define the atomic bonds to show
bondatoms = []
symbols = fenbs2.get_chemical_symbols()
for i in range(len(fenbs2)):
for j in range(i):
if (symbols[i] in {'Nb', 'S'} and symbols[j] in {'S', 'Nb'} and symbols[i] != symbols[j] and
fenbs2.get_distance(i, j) < 4):
bondatoms.append((i, j))
# Create nice-looking image using povray
io.write('spacegroup-fenbs2.pov', fenbs2,
transparent=False,
display=False,
run_povray=True,
camera_type='perspective',
canvas_width=1200,
radii=0.4,
rotation='5y,-80x',
bondlinewidth=0.07,
bondatoms=bondatoms)
#view(alloy, viewer='x3d')
# -
from ase.build import graphene_nanoribbon
graphene =
# +
a = 5.761
c = 12.178
graphene = graphene_nanoribbon(3, 3, sheet=True)
overlayer = graphene.copy()
# Define the atomic bonds to show
bondatoms = []
symbols = graphene.get_chemical_symbols()
for i in range(len(graphene)):
for j in range(i):
if graphene.get_distance(i, j) < 1.5:
bondatoms.append((i, j))
# Create nice-looking image using povray
io.write('spacegroup-graphene.pov', graphene,
transparent=False,
display=False,
run_povray=True,
camera_type='perspective',
canvas_width=1200,
radii=0.4,
rotation='0y,-90x',
bondlinewidth=0.07,
bondatoms=bondatoms)
#view(alloy, viewer='x3d')
# +
from ase.build import mx2, stack
ws2 = mx2('WS2', size=(30, 30, 1))
# Define the atomic bonds to show
bondatoms = []
symbols = ws2.get_chemical_symbols()
ls = []
for i in ws2:
ls.append(i.position)
ps = np.stack(ls)
vs = squareform(pdist(ps))
for i in range(vs.shape[0]):
for j in range(i):
if vs[i,j] < 3:
bondatoms.append((i, j))
# Create nice-looking image using povray
colors.jmol_colors[74] = (44. / 255, 132. / 255, 30. / 255) # W
colors.jmol_colors[16] = (109. / 255, 193. / 255, 96. / 255)
colors.cpk_colors[74] = (44. / 255, 132. / 255, 30. / 255) # W
colors.cpk_colors[16] = (109. / 255, 193. / 255, 96. / 255)
#colors.jmol_colors[74] = (229. / 255, 139. / 255, 22. / 255) # W
#colors.jmol_colors[34] = (224. / 255, 169. / 255, 98. / 255) # W
#colors.cpk_colors[74] = (229. / 255, 139. / 255, 22. / 255) # W
#colors.cpk_colors[34] = (224. / 255, 169. / 255, 98. / 255) # W
io.write('spacegroup-ws2.pov', ws2,
transparent=True,
display=False,
run_povray=False,
camera_type='orthographic',
canvas_width=1200,
background=(0, 0, 0, 1.,), # weird alpha convention
radii=0.4,
rotation='0y,0x',
bondlinewidth=0.07,
bondatoms=bondatoms)
#view(alloy, viewer='x3d')
# W 44, 132, 30
# S 109, 193, 96
# W 229, 139, 22
# Se 224, 169, 98
# -
from ase.data import colors
colors.jmol_colors = colors_copy
# +
# io.write?
# -
from scipy.spatial.distance import pdist, squareform
squareform(pdist(ps))
ps.shape
np.matmul
# +
fig, ax = plt.subplots(1, 3, figsize=(15, 5,))
arr = xr.DataArray(np.linspace(0, 1, 100), coords={'x': np.linspace(0, 0.1, 100)}, dims=['x'])
arr.plot(ax=ax[0])
copied = arr.copy(deep=True)
copied.coords['x'] = copied.coords['x'] + 0.05
copied.plot(ax=ax[1])
arr.plot(ax=ax[2])
for axi in ax:
axi.set_xlim([0, 0.15])
axi.set_ylim([0, 1])
# +
fig, ax = plt.subplots(1, 3, figsize=(15, 5,))
arr = xr.DataArray(np.linspace(0, 1, 100), coords={'x': np.linspace(0, 0.1, 100)}, dims=['x'])
arr.plot(ax=ax[0])
copied = arr.copy(deep=True)
copied.coords['x'] += 0.05
copied.plot(ax=ax[1])
arr.plot(ax=ax[2])
|
front/public/notebooks/crystal_structure_example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from PIL import Image
import numpy as np
import matplotlib.pyplot as plt
from scipy import stats
import math
# %matplotlib inline
# # Patient 8 THY
# ## 3M Littmann Data
image = Image.open('3M_zhy_Post.bmp')
image
x = image.size[0]
y = image.size[1]
print(x)
print(y)
# +
matrix = []
points = []
integrated_density = 0
for i in range(x):
matrix.append([])
for j in range(y):
matrix[i].append(image.getpixel((i,j)))
#integrated_density += image.getpixel((i,j))[1]
#points.append(image.getpixel((i,j))[1])
# -
# ### Extract Red Line Position
# +
redMax = 0
xStore = 0
yStore = 0
for xAxis in range(x):
for yAxis in range(y):
currentPoint = matrix[xAxis][yAxis]
if currentPoint[0] == 255 and currentPoint[1] < 10 and currentPoint[2] < 10:
redMax = currentPoint[0]
xStore = xAxis
yStore = yAxis
print(xStore, yStore)
# -
# ### Extract Blue Points
# +
redline_pos = 74
gain = 120
absMax = 0
littmannArr = []
points_vertical = []
theOne = 0
for xAxis in range(x):
for yAxis in range(y):
currentPoint = matrix[xAxis][yAxis]
# Pickup Blue points
if currentPoint[2] == 255 and currentPoint[0] < 100 and currentPoint[1] < 100:
points_vertical.append(yAxis)
#print(points_vertical)
# Choose the largest amplitude
for item in points_vertical:
if abs(item-redline_pos) > absMax:
absMax = abs(item-redline_pos)
theOne = item
littmannArr.append((theOne-redline_pos)*gain)
absMax = 0
theOne = 0
points_vertical = []
# -
fig = plt.figure()
s = fig.add_subplot(111)
s.plot(littmannArr, linewidth=0.6, color='blue')
# # Ascul Pi Data
pathBase = 'C://Users//triti//OneDrive//Dowrun//Text//Manuscripts//Data//ZhaoHongye//AusculPi_Post//'
filename = 'Numpy_Array_File_2020-07-06_17_45_18.npy'
line = pathBase + filename
arr = np.load(line)
arr
arr.shape
fig = plt.figure()
s = fig.add_subplot(111)
s.plot(arr[0], linewidth=1.0, color='black')
fig = plt.figure()
s = fig.add_subplot(111)
s.plot(arr[:,200], linewidth=1.0, color='black')
# +
start = 366
end = 1168
start_adj = int(start * 2583 / 3000)
end_adj = int(end * 2583 / 3000)
# -
fig = plt.figure()
s = fig.add_subplot(111)
s.plot(arr[start_adj:end_adj,500], linewidth=0.6, color='black')
start_adj-end_adj
fig = plt.figure()
s = fig.add_subplot(111)
s.plot(littmannArr, linewidth=0.6, color='blue')
asculArr = arr[start_adj:end_adj,500]
fig = plt.figure()
s = fig.add_subplot(111)
s.plot(asculArr, linewidth=0.6, color='black')
# ## Preprocess the two array
# +
asculArr_processed = []
littmannArr_processed = []
for ascul in asculArr:
asculArr_processed.append(math.fabs(ascul))
for item in littmannArr:
littmannArr_processed.append(math.fabs(item))
# -
fig = plt.figure()
s = fig.add_subplot(111)
s.plot(asculArr_processed, linewidth=1.0, color='black')
fig = plt.figure()
s = fig.add_subplot(111)
s.plot(littmannArr_processed, linewidth=1.0, color='blue')
len(littmannArr)
len(asculArr)
fig = plt.figure()
s = fig.add_subplot(111)
s.plot(asculArr_processed[:100], linewidth=1.0, color='black')
fig = plt.figure()
s = fig.add_subplot(111)
s.plot(littmannArr_processed[:100], linewidth=1.0, color='blue')
# ### Coeffient
stats.pearsonr(asculArr_processed, littmannArr_processed)
stats.pearsonr(asculArr_processed[:100], littmannArr_processed[:100])
# ### Fitness
stats.chisquare(asculArr_processed[:80], littmannArr_processed[2:82])
def cosCalculate(a, b):
l = len(a)
sumXY = 0
sumRootXSquare = 0
sumRootYSquare = 0
for i in range(l):
sumXY = sumXY + a[i]*b[i]
sumRootXSquare = sumRootXSquare + math.sqrt(a[i]**2)
sumRootYSquare = sumRootYSquare + math.sqrt(b[i]**2)
cosValue = sumXY / (sumRootXSquare * sumRootYSquare)
return cosValue
cosCalculate(asculArr_processed, littmannArr_processed)
|
WaveformExtraction_zhy_Post.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/yohanesnuwara/ccs-gundih/blob/master/main/05_visualize_synthetic_seismic.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="_ERrfy7_HbeR" colab_type="text"
# # Visualize Synthetic Seismic Data of Gundih $CO_2$ Injection
# + id="fpmN_MTNyQuh" colab_type="code" colab={}
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# + id="jNmITRpiyBIH" colab_type="code" outputId="34dceef0-80ee-4cab-fd01-0c2340cad5d6" colab={"base_uri": "https://localhost:8080/", "height": 117}
# !git clone https://www.github.com/yohanesnuwara/ccs-gundih
# + id="3IN7ggMsyWM2" colab_type="code" colab={}
# !pip install obspy
# + [markdown] id="h8vZ5pTtHaQc" colab_type="text"
# ## Synthetic Seismic at Baseline (before injection)
# + id="Ghnvp1kxyhdx" colab_type="code" colab={}
from obspy.io.segy.segy import _read_segy
baseline = _read_segy('/content/ccs-gundih/data/seismics/Nov2019_M0.sgy', headonly=True)
# + id="ryvSN3fbyz_I" colab_type="code" outputId="5c7ccc71-1ee5-47f1-e35e-b20050c0b628" colab={"base_uri": "https://localhost:8080/", "height": 33}
baseline
# + id="JvUKGBFly1oC" colab_type="code" outputId="de64a87a-7f36-4e21-8a92-b114c72ab2ab" colab={"base_uri": "https://localhost:8080/", "height": 159}
one_trace_m0 = baseline.traces[190]
plt.figure(figsize=(16,2))
plt.plot(one_trace_m0.data)
plt.show()
# + id="zeIkldVMzCKN" colab_type="code" outputId="a16d426d-1869-4657-c8af-c8f346f6a7d4" colab={"base_uri": "https://localhost:8080/", "height": 87}
data_m0 = np.stack(t.data for t in baseline.traces)
data_m0.shape # traces, time samples
# + id="Xy7OVsCwzcCl" colab_type="code" outputId="9c523d7b-e664-433d-e1a3-f422962d3d88" colab={"base_uri": "https://localhost:8080/", "height": 33}
vm = np.percentile(data_m0, 99)
print("The 99th percentile is {:.0f}; the max amplitude is {:.0f}".format(vm, data_m0.max()))
# + id="vY_x-iekzLMa" colab_type="code" outputId="bad6ed6b-b0b5-416a-eb7c-478c6af2019e" colab={"base_uri": "https://localhost:8080/", "height": 421}
plt.figure(figsize=(15,6))
plt.imshow(data_m0.T, cmap="Greys", vmin=-0.05, vmax=0.05, aspect='auto', interpolation='lanczos')
plt.colorbar()
plt.title('Synthetic Post-Stack Seismic Section of Baseline', size=20, pad=15)
plt.xlabel('Trace', size=13); plt.ylabel('Time (ms)', size=13)
plt.xlim(0, 225); plt.ylim(775, 620)
# plt.grid(True, axis='y', color='black')
plt.show()
# + [markdown] id="ZCOjxYanIYQd" colab_type="text"
# ## Synthetic Seismic after 10 years injection (Monitor, M2)
# + id="tHVq_8twIgRv" colab_type="code" outputId="97d3a151-0d37-4705-ac12-76c455ac29bd" colab={"base_uri": "https://localhost:8080/", "height": 159}
monitor2 = _read_segy('/content/ccs-gundih/data/seismics/Nov2019_M2.sgy', headonly=True)
one_trace_m2 = monitor2.traces[190]
plt.figure(figsize=(16,2))
plt.plot(one_trace_m2.data)
plt.show()
# + id="fIoCHIO_I_BT" colab_type="code" outputId="d1b1d0bd-63a8-4508-9381-ed01da837111" colab={"base_uri": "https://localhost:8080/", "height": 474}
data_m2 = np.stack(t.data for t in monitor2.traces)
plt.figure(figsize=(15,6))
plt.imshow(data_m2.T, cmap="Greys", vmin=-0.05, vmax=0.05, aspect='auto', interpolation='lanczos')
plt.colorbar()
plt.title('Synthetic Post-Stack Seismic Section of Post-Injection 10 Years', size=20, pad=15)
plt.xlabel('Trace', size=13); plt.ylabel('Time (ms)', size=13)
plt.xlim(0, 225); plt.ylim(775, 620)
# plt.grid(True, axis='y', color='black')
plt.show()
# + [markdown] id="oixDnAMsKHtW" colab_type="text"
# ## Original Seismic Data
# + id="0ru1BkJVKMgI" colab_type="code" outputId="bd88ec57-f8ff-41bb-d453-788cbe457018" colab={"base_uri": "https://localhost:8080/", "height": 33}
ori = _read_segy('/content/ccs-gundih/data/seismics/2D_Xline_12680.sgy', headonly=True)
ori
# + id="quEJwEghKkK-" colab_type="code" outputId="093d863a-d5dd-4c85-940a-6b96934858a6" colab={"base_uri": "https://localhost:8080/", "height": 157}
one_trace_ori = ori.traces[400]
plt.figure(figsize=(16,2))
plt.plot(one_trace_ori.data)
plt.show()
# + id="kgAKv9VpK0kL" colab_type="code" outputId="8405658b-89e3-4f96-ebfd-c49e7af188ef" colab={"base_uri": "https://localhost:8080/", "height": 87}
data_ori = np.stack(i.data for i in ori.traces)
vm = np.percentile(data_ori, 99)
print("The 99th percentile is {:.0f}; the max amplitude is {:.0f}".format(vm, data_ori.max()))
# + id="wXUaqWe2LNbq" colab_type="code" outputId="fb52936e-0426-4f73-b00d-3fe55a2f3c74" colab={"base_uri": "https://localhost:8080/", "height": 421}
plt.figure(figsize=(15,6))
plt.imshow(data_ori.T, cmap="Greys", vmin=-4300, vmax=4300, aspect='auto', interpolation='lanczos')
plt.colorbar()
plt.title('Original Post-Stack Seismic Section', size=20, pad=15)
plt.xlabel('Trace', size=13); plt.ylabel('Time (ms)', size=13)
plt.xlim(450, 560); plt.ylim(680, 560)
# plt.grid(True, axis='y', color='grey')
plt.show()
# + [markdown] id="9ZYnCWjEV_o_" colab_type="text"
# ## Velocity Model
# + id="0MXDaZmeWCwW" colab_type="code" outputId="109749d5-088b-4507-c2ed-8a559be50664" colab={"base_uri": "https://localhost:8080/", "height": 33}
vel = _read_segy('/content/ccs-gundih/data/seismics/2D_Vel_Xline_12680.sgy', headonly=True)
vel
# + id="1ENDdgBsWLVZ" colab_type="code" outputId="199bb8a4-cdd2-4f91-fea8-31bd795c4400" colab={"base_uri": "https://localhost:8080/", "height": 87}
data_vel = np.stack(i.data for i in vel.traces)
vm = np.percentile(data_vel, 99)
print("The 99th percentile is {:.0f}; the max amplitude is {:.0f}".format(vm, data_vel.max()))
# + id="P1wgm3QCWWlR" colab_type="code" outputId="cfb2e441-74d7-4860-9570-36c6a7d58908" colab={"base_uri": "https://localhost:8080/", "height": 421}
plt.figure(figsize=(15,6))
plt.imshow(data_vel.T, cmap="jet", vmin=3000, vmax=5300, aspect='auto', interpolation='lanczos')
plt.colorbar()
plt.title('Seismic P-Wave Velocity Section', size=20, pad=15)
plt.xlabel('Trace', size=13); plt.ylabel('Time (ms)', size=13)
plt.xlim(450, 560); plt.ylim(1375, 1125)
# plt.grid(True, axis='y', color='grey')
plt.show()
|
main/05_visualize_synthetic_seismic.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# [](https://www.pythonista.io)
# # Seguridad y autenticación.
# ```
# src/api_segura/
# ├── main.py
# ├── config.py
# ├── manage.py
# ├── requirements.txt
# ├── api
# │ ├── api.py
# │ ├── auth
# │ ├── crud.py
# │ ├── __init__.py
# │ └── schemas.py
# └── data
# └── __init__.py
# ```
# %cd src/api_segura
# !uvicorn main:app --host 0.0.0.0 --reload
# <p style="text-align: center"><a rel="license" href="http://creativecommons.org/licenses/by/4.0/"><img alt="Licencia Creative Commons" style="border-width:0" src="https://i.creativecommons.org/l/by/4.0/80x15.png" /></a><br />Esta obra está bajo una <a rel="license" href="http://creativecommons.org/licenses/by/4.0/">Licencia Creative Commons Atribución 4.0 Internacional</a>.</p>
# <p style="text-align: center">© <NAME>. 2022.</p>
|
22_api_segura.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <table width="100%"> <tr>
# <td style="background-color:#ffffff;">
# <a href="http://qworld.lu.lv" target="_blank"><img src="..\images\qworld.jpg" width="35%" align="left"> </a></td>
# <td style="background-color:#ffffff;vertical-align:bottom;text-align:right;">
# prepared by <NAME> (<a href="http://qworld.lu.lv/index.php/qlatvia/" target="_blank">QLatvia</a>)
# <br>
# updated by <NAME> | August 28, 2019
# </td>
# </tr></table>
# <table width="100%"><tr><td style="color:#bbbbbb;background-color:#ffffff;font-size:11px;font-style:italic;text-align:right;">This cell contains some macros. If there is a problem with displaying mathematical formulas, please run this cell to load these macros. </td></tr></table>
# $ \newcommand{\bra}[1]{\langle #1|} $
# $ \newcommand{\ket}[1]{|#1\rangle} $
# $ \newcommand{\braket}[2]{\langle #1|#2\rangle} $
# $ \newcommand{\dot}[2]{ #1 \cdot #2} $
# $ \newcommand{\biginner}[2]{\left\langle #1,#2\right\rangle} $
# $ \newcommand{\mymatrix}[2]{\left( \begin{array}{#1} #2\end{array} \right)} $
# $ \newcommand{\myvector}[1]{\mymatrix{c}{#1}} $
# $ \newcommand{\myrvector}[1]{\mymatrix{r}{#1}} $
# $ \newcommand{\mypar}[1]{\left( #1 \right)} $
# $ \newcommand{\mybigpar}[1]{ \Big( #1 \Big)} $
# $ \newcommand{\sqrttwo}{\frac{1}{\sqrt{2}}} $
# $ \newcommand{\dsqrttwo}{\dfrac{1}{\sqrt{2}}} $
# $ \newcommand{\onehalf}{\frac{1}{2}} $
# $ \newcommand{\donehalf}{\dfrac{1}{2}} $
# $ \newcommand{\hadamard}{ \mymatrix{rr}{ \sqrttwo & \sqrttwo \\ \sqrttwo & -\sqrttwo }} $
# $ \newcommand{\vzero}{\myvector{1\\0}} $
# $ \newcommand{\vone}{\myvector{0\\1}} $
# $ \newcommand{\vhadamardzero}{\myvector{ \sqrttwo \\ \sqrttwo } } $
# $ \newcommand{\vhadamardone}{ \myrvector{ \sqrttwo \\ -\sqrttwo } } $
# $ \newcommand{\myarray}[2]{ \begin{array}{#1}#2\end{array}} $
# $ \newcommand{\X}{ \mymatrix{cc}{0 & 1 \\ 1 & 0} } $
# $ \newcommand{\Z}{ \mymatrix{rr}{1 & 0 \\ 0 & -1} } $
# $ \newcommand{\Htwo}{ \mymatrix{rrrr}{ \frac{1}{2} & \frac{1}{2} & \frac{1}{2} & \frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & \frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} & \frac{1}{2} } } $
# $ \newcommand{\CNOT}{ \mymatrix{cccc}{1 & 0 & 0 & 0 \\ 0 & 1 & 0 & 0 \\ 0 & 0 & 0 & 1 \\ 0 & 0 & 1 & 0} } $
# $ \newcommand{\norm}[1]{ \left\lVert #1 \right\rVert } $
# <h2> <font color="blue"> Solutions </font> for Multiple Qubits </h2>
# <a id="task1"></a>
# <h3> Task 1 </h3>
#
# Find the state of the composite system.
# <h3> Solution </h3>
#
# $$ \Bigg(\frac{1}{\sqrt{3}} \ket{0} + \frac{\sqrt{2}}{\sqrt{3}} \ket{1} \Bigg ) \otimes \Bigg( -\frac{1}{\sqrt{2}} \ket{0} + \frac{1}{\sqrt{2}} \ket{1} \Bigg )=$$
#
#
# $$ -\frac{1}{\sqrt{6}}\big( \ket{0} \otimes \ket{0} \big) + \frac{1}{\sqrt{6}} \big( \ket{0} \otimes \ket{1} \big) -\frac{\sqrt{2}}{\sqrt{6}} \big( \ket{1} \otimes \ket{0} \big) + \frac{\sqrt{2}}{\sqrt{6}} \big( \ket{1} \otimes \ket{1} \big) = $$
#
# $$ -\frac{1}{\sqrt{6}} \ket{00} + \frac{1}{\sqrt{6}} \ket{01} -\frac{\sqrt{2}}{\sqrt{6}} \ket{10} + \frac{\sqrt{2}}{\sqrt{6}} \ket{11}. $$
# <a id = "task2"></a>
# <h3> Task 2 </h3>
#
# Pick one of the following cases (2, 3, or 4), and verify the correctness of all three different ways for this selected case.
# <hr>
# <h3> Solution </h3>
# <h4> Case 2: Let's find $ H^{\otimes 2} \ket{01} $ (in three different ways) </h4>
# <ul>
# <li> Direct matrix-vector multiplication:
# $$
# H^{\otimes 2} \ket{01}
# = \Htwo \myvector{0 \\ 1 \\ 0 \\ 0}
# = \myrvector{ \frac{1}{2} \\ - \frac{1}{2} \\ \frac{1}{2} \\ - \frac{1}{2} } .
# $$ </li>
# <li> We calculate the quantum state of each state, and then find the quantum state of the composite system.
# $$
# H\ket{0} \otimes H \ket{1}
# = \vhadamardzero \otimes \vhadamardone
# = \myrvector{ \frac{1}{2} \\ - \frac{1}{2} \\ \frac{1}{2} \\ - \frac{1}{2} }.
# $$ </li>
# <li> We make calculations with $ \ket{0} $ and $ \ket{1} $.
# $$
# H \ket{0} \otimes H \ket{1}
# = \mypar{ \frac{1}{\sqrt{2}} \ket{0} + \frac{1}{\sqrt{2}} \ket{1} }
# \otimes \mypar{ \frac{1}{\sqrt{2}} \ket{0} - \frac{1}{\sqrt{2}} \ket{1} }
# = \frac{1}{2} \ket{00} - \frac{1}{2} \ket{01} + \frac{1}{2} \ket{10} - \frac{1}{2} \ket{11}
# = \myrvector{ \frac{1}{2} \\ - \frac{1}{2} \\ \frac{1}{2} \\ - \frac{1}{2} }.
# $$
# </ul>
#
# <h4> Case 3: Let's find $ H^{\otimes 2} \ket{10} $ (in three different ways) </h4>
# <ul>
# <li> Direct matrix-vector multiplication:
# $$
# H^{\otimes 2} \ket{10}
# = \Htwo \myvector{0 \\ 0 \\ 1 \\ 0}
# = \myrvector{ \frac{1}{2} \\ \frac{1}{2} \\ - \frac{1}{2} \\ - \frac{1}{2} } .
# $$ </li>
# <li> We calculate the quantum state of each state, and then find the quantum state of the composite system.
# $$
# H\ket{1} \otimes H \ket{0}
# = \vhadamardone \otimes \vhadamardzero
# = \myrvector{ \frac{1}{2} \\ \frac{1}{2} \\ - \frac{1}{2} \\ - \frac{1}{2} }.
# $$ </li>
# <li> We make calculations with $ \ket{0} $ and $ \ket{1} $.
# $$
# H \ket{1} \otimes H \ket{0}
# = \mypar{ \frac{1}{\sqrt{2}} \ket{0} - \frac{1}{\sqrt{2}} \ket{1} }
# \otimes \mypar{ \frac{1}{\sqrt{2}} \ket{0} + \frac{1}{\sqrt{2}} \ket{1} }
# = \frac{1}{2} \ket{00} + \frac{1}{2} \ket{01} - \frac{1}{2} \ket{10} - \frac{1}{2} \ket{11}
# = \myrvector{ \frac{1}{2} \\ \frac{1}{2} \\ - \frac{1}{2} \\ -\frac{1}{2} }.
# $$
# </ul>
#
# <h4> Case 4: Let's find $ H^{\otimes 2} \ket{11} $ (in three different ways) </h4>
# <ul>
# <li> Direct matrix-vector multiplication:
# $$
# H^{\otimes 2} \ket{11}
# = \Htwo \myvector{0 \\ 0 \\ 0 \\ 1}
# = \myrvector{ \frac{1}{2} \\ - \frac{1}{2} \\ - \frac{1}{2} \\ \frac{1}{2} } .
# $$ </li>
# <li> We calculate the quantum state of each state, and then find the quantum state of the composite system.
# $$
# H\ket{1} \otimes H \ket{1}
# = \vhadamardone \otimes \vhadamardone
# = \myrvector{ \frac{1}{2} \\ - \frac{1}{2} \\ - \frac{1}{2} \\ \frac{1}{2} }.
# $$ </li>
# <li> We make calculations with $ \ket{0} $ and $ \ket{1} $.
# $$
# H \ket{1} \otimes H \ket{1}
# = \mypar{ \frac{1}{\sqrt{2}} \ket{0} - \frac{1}{\sqrt{2}} \ket{1} }
# \otimes \mypar{ \frac{1}{\sqrt{2}} \ket{0} - \frac{1}{\sqrt{2}} \ket{1} }
# = \frac{1}{2} \ket{00} - \frac{1}{2} \ket{01} - \frac{1}{2} \ket{10} + \frac{1}{2} \ket{11}
# = \myrvector{ \frac{1}{2} \\ - \frac{1}{2} \\ - \frac{1}{2} \\ \frac{1}{2} }.
# $$
# </ul>
# <a id="task3"></a>
# <h3> Task 3 </h3>
#
#
# If the following vectors are valid quantum states defined with real numbers, then what can be the values of $a$ and $b$?
#
# $$
# \ket{v} = \myrvector{-0.1 \\ a \\ -0.3 \\ 0.4 \\ 0.5 \\ 0.06 \\ 0.07 \\ -0.08}
# ~~~~~ \mbox{and} ~~~~~
# \ket{u} = \myrvector{ \frac{1}{\sqrt{2}} \\ \frac{1}{\sqrt{b}} \\ -\frac{1}{\sqrt{5}} \\ \frac{1}{\sqrt{10}} }.
# $$
# <h3>Solution</h3>
#
# +
# vector |v>
print("vector |v>")
values = [-0.1, -0.3, 0.4, 0.5,0.06,0.07,-0.08]
total = 0 # summation of squares
for i in range(len(values)):
total += values[i]**2; # add the square of each value
print("total is ",total)
print("the missing part is",1-total)
print("so, the value of 'a' can be",(1-total)**0.5,"or",-(1-total)**0.5) # square root of the missing part
print()
print("vector |u>")
values = [1/(2**0.5), -1/(5**0.5),1/(10**0.5)]
total = 0 # summation of squares
for i in range(len(values)):
total += values[i]**2; # add the square of each value
print("total is ",total)
print("the missing part is",1-total)
# the missing part is 1/b, square of 1/sqrt(b)
# thus, b is 1/missing_part
print("so, the value of 'b' should be",1/(1-total))
# -
#
# <a id="task4"></a>
# <h3> Task 4</h3>
#
# Create a quantum circuit with two qubits: qreg[0] and qreg[1].
#
# <i> Qiskit combines these two qubits as qreg[1]$\otimes$qreg[0]. </i>
#
# Apply h-gate to qreg[0].
#
# Apply first x-gate and then a z-gate to qreg[1].
#
# Calculate the quantum states of both qubits and then the quantum state of composite system (qreg[1]$\otimes$qreg[0]) on paper.
#
# Read quantum state of the quantum circuit.
#
# Compare your results.
# <h3> Solution </h3>
# The quantum state of qreg[1] is $ \Z \X \vzero = \Z \vone= \myrvector{ 0 \\ -1 } $.
#
# The quantum state of qreg[0] is $ \hadamard \myvector{ 1 \\ 0} = \vhadamardzero$.
#
# The quantum state of qreg[1]$\otimes$qreg[0] is $ \myvector{ 0 \\ -1 } \otimes \vhadamardzero = \myvector{ 0 \vhadamardzero \\ -1 \vhadamardzero } = \myvector{0 \\ 0 \\ -\frac{1}{\sqrt{2} } \\ -\frac{1}{\sqrt{2} } } $.
# +
# we can print the values by using python
print(0,0,round(-1/(2**0.5),4),round(-1/(2**0.5),4) )
# +
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
from math import pi
qreg = QuantumRegister(2)
creg = ClassicalRegister(2)
mycircuit1 = QuantumCircuit(qreg,creg)
mycircuit1.x(qreg[1])
mycircuit1.z(qreg[1])
mycircuit1.h(qreg[0])
job = execute(mycircuit1,Aer.get_backend('statevector_simulator'),optimization_level=0)
current_quantum_state=job.result().get_statevector(mycircuit1)
for amplitude in current_quantum_state:
print( round(amplitude.real,4) )
# -
# <a id="task5"></a>
# <h3> Task 5</h3>
#
# Create a quantum ciruit with 5 qubits.
#
# Apply h-gate (Hadamard operator) to each qubit.
#
# Apply z-gate ($Z$ operator) to randomly picked qubits. (i.e., $ mycircuit.z(qreg[i]) $)
#
# Apply h-gate to each qubit.
#
# Measure each qubit.
#
# Execute your program 1000 times.
#
# Compare the outcomes of the qubits affected by z-gates, and the outcomes of the qubits not affected by z-gates.
#
# Does z-gate change the outcome?
#
# Why?
# <h3> Solution </h3>
# +
# import all necessary objects and methods for quantum circuits
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
# import randrange for random choices
from random import randrange
number_of_qubit = 5
# define a quantum register with 5 qubits
qreg = QuantumRegister(number_of_qubit)
# define a classical register with 5 bits
creg = ClassicalRegister(number_of_qubit)
# define our quantum circuit
mycircuit2 = QuantumCircuit(qreg,creg)
# apply h-gate to all qubits
for i in range(number_of_qubit):
mycircuit2.h(qreg[i])
# apply z-gate to randomly picked qubits
for i in range(number_of_qubit):
if randrange(2) == 0: # the qubit with index i is picked to apply z-gate
mycircuit2.z(qreg[i])
# apply h-gate to all qubits
for i in range(number_of_qubit):
mycircuit2.h(qreg[i])
# measure all qubits
mycircuit2.measure(qreg,creg)
print("Everything looks fine, let's continue ...")
# +
# draw the circuit
mycircuit2.draw()
# +
# execute the circuit 1000 times in the local simulator
job = execute(mycircuit2,Aer.get_backend('qasm_simulator'),shots=1000)
counts = job.result().get_counts(mycircuit2)
for outcome in counts: # for each key-value in dictionary
reverse_outcome = ''
for i in outcome: # each string can be considered as a list of characters
reverse_outcome = i + reverse_outcome # each new symbol comes before the old symbol(s)
print(reverse_outcome,"is observed",counts[outcome],"times")
# -
# We start in state $ \ket{0} $ in each qubit.
#
# If a qubit is affected by z-gate, then its final value is changed to $ \ket{1} $.
#
# If a qubit is not affected by z-gate, then its final value is $ \ket{0} $.
# <a id="task6"></a>
# <h3> Task 6 [Extra] </h3>
#
# Design a quantum circuit with 10 quantum bits and 10 classical bits.
#
# For each quantum bit, flip a coin, and apply x-gate if the outcome is head.
#
# Measure your quantum bits.
#
# Execeute your circuit 128 times.
#
# Repeat this task as mush as you want, and enjoy your random choices.
# <h3>Solution</h3>
# +
# we import all necessary methods and objects
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
from random import randrange
# we will use 10 quantum bits and 10 classical bits
qreg3 = QuantumRegister(10)
creg3 = ClassicalRegister(10)
mycircuit3 = QuantumCircuit(qreg3,creg3)
# we will store the index of each qubit to which x-gate is applied
picked_qubits=[]
for i in range(10):
if randrange(2) == 0: # Assume that 0 is Head and 1 is Tail
mycircuit3.x(qreg3[i]) # apply x-gate
print("x-gate is applied to the qubit with index",i)
picked_qubits.append(i) # i is picked
# measurement
mycircuit3.measure(qreg3,creg3)
print("Everything looks fine, let's continue ...")
# +
# draw the circuit
mycircuit3.draw(reverse_bits=True)
#mycircuit3.draw(output='mpl',reverse_bits=True)
# reexecute me if you DO NOT see the circuit diagram
# +
# execute the circuit and read the results
job = execute(mycircuit3,Aer.get_backend('qasm_simulator'),shots=128)
counts = job.result().get_counts(mycircuit3)
print(counts)
|
bronze/B38_Multiple_Qubits_Solutions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Create an L-layer neural network (classification)
#
# Since the goal is to learn the details of neural networks, this model is implemented using numpy matrix operations. We also have added l2 and dropout regularization options.
#
# ## Model parameters
#
# * X = $n_x$ by m array where training vectors are in columns
# * Y = 1 by m array of true class values
# * $n_x$ = number of predictors
# * m = number of training examples
# * L = number of layers
# * n = list containing number of neurons per layer, n[0] = $n_x$
# * Wi = n[i] by n[i-1] array of model parameters for layer i
# * bi = n[i] by i array of bias parameters for layer i
# * dWi = n[i] by n[i-1] array of partial derivatives of the cost function with respect to the corresponding parameters of Wi
# * dbi = n[i] by i array of partial derivatives of the cost function with respect to the corresponding bias parameters
# * regularization = None or 'l2'
# * lambd = regularization parameter for l2
import numpy as np
# activation function for output vector
def sigmoid(z):
return 1/(1+np.exp(-z))
# activation function for hidden layers
def relu(z):
return np.maximum(0, z)
# derivative of the relu function. We keep the shape of the input array for vectorization purposes. Strictly
# speaking, the total derivative should be a (much larger) diagonal matrix
def der_relu(z):
return (z>0).astype(int)
def predict(X, parameters):
'''Predict binary class value given input X and model parameters. The activation function
for each hidden layer is a ReLU and the output activation function is a sigmoind. Since this
function is only used for model predictions, no intermediate values of the model are cached
or returned to the user.
Input:
X = n_x by m array of feature vectors
parameters = dictionary of parameters Wi and bi for the neural network
Output:
A = 1 by m array of predicted class values.'''
# Calculate the number of layers in the network
L = len(parameters)//2
# hidden layer computations
A = X
for i in range(1, L):
A = relu(np.dot(parameters['W'+str(i)], A) + parameters['b'+str(i)])
# output layer
A = sigmoid(np.dot(parameters['W'+str(L)], A) + parameters['b'+str(L)])
# predict class
A = (A >= 0.5)
return A.astype(int)
def initialize(n):
'''Initialize model parameters using the input specification. We use He initialization since we have ReLU
activation functions.
Input:
n = length L+1 list where n[i] is the number of neurons in layer i
Output:
parameters = dictionary containing initialized model parameters Wi and bi'''
parameters = dict()
for i in range(1, len(n)):
parameters['W'+str(i)] = np.random.randn(n[i], n[i-1])*(2/np.sqrt(n[i-1]))
parameters['b'+str(i)] = np.zeros((n[i], 1))
return parameters
def propogate(X, Y, parameters, regularization=None):
'''Perform forward and backward propogation.
Input:
X = n_x by m array containing training examples
Y = 1 by m array containing true binary class value for training examples
parameters = dictionary of model parameters Wi and bi
Output:
gradient = dictionary of gradient arrays dWi and dbi corresponding to model parameters Wi and bi'''
m = X.shape[1] # infer the number of training examples from X
L = len(parameters)//2 # infer the number of layers from the number of parameters
sum_vec = np.full((m, 1), 1) # vector used in gradient calculations
cache = dict() # cache for results of intermediate steps
# forward propogation
# hidden layer computations
A = X # A stores the activation of the previous layer, initializes to training vectors
cache['A0'] = A
for i in range(1, L):
Z = np.dot(parameters['W'+str(i)], A) + parameters['b'+str(i)] # linear step
A = relu(Z) # activation
cache['Z'+str(i)] = Z # store intermediate values for use in gradient calcululation
cache['A'+str(i)] = A
# output layer
Z = np.dot(parameters['W'+str(L)], A) + parameters['b'+str(L)] # linear step
A = sigmoid(Z) # activation
cache['Z'+str(L)] = Z # store intermediate values for use in gradient calculation
cache['A'+str(L)] = A
#backward propogation
gradient = dict()
# output layer
dA = (1/m)*(A-Y) # tracking matrix for chain rule
gradient['dW'+str(L)] = np.dot(dA, cache['A'+str(L-1)].T) # dWL
gradient['db'+str(L)] = np.dot(dA, sum_vec) # dbL
# hidden layers
for i in reversed(range(1, L)):
dA = np.dot(dA.T, parameters['W'+str(i+1)])*der_relu(cache['Z'+str(i)].T) # update dA. See page 3 of notes for derivation/proof
dA = dA.T
gradient['dW'+str(i)] = np.dot(dA, cache['A'+str(i-1)].T) # dWi
gradient['db'+str(i)] = np.dot(dA, sum_vec) # dbi
return gradient
def cost(Yhat, Y, eps=0.000000001, regularization=None, lambd=0, parameters=None):
'''Calculate cost of prediction Yhat. The small value eps is used to prevent log(0) error.
Input:
Yhat = 1 by m array of class probabilities
Y = 1 by m array of true binary class
eps = small value to prevent log(0) error
regularization = None or l2
lambd = l2 regularization constant
parameters = dictionary of model parameters
Output:
cost = cost of the given prediction Yhat'''
m = Yhat.shape[1] # infer the number of training examples
if regularization == 'l2':
L = len(parameters)//2 # infer the number of layers
l2_sum = 0 # sum squares of parameters
for i in range(1, L+1):
l2_sum += np.sum(parameters['W'+str(i)]**2)
return -(1/m)*np.sum(Y*np.log(Yhat + eps)+(1-Y)*np.log(1-Yhat + eps)) + (lambd/(2*m))*l2_sum
else:
return -(1/m)*np.sum(Y*np.log(Yhat + eps)+(1-Y)*np.log(1-Yhat + eps))
def fit(X, Y, parameters, learning_rate=0.01, iterations=2000, regularization=None, lambd=0, print_cost=False):
L = len(parameters)//2
m = X.shape[0]
for i in range(iterations):
gradient = propogate(X, Y, parameters, regularization=regularization)
# print cost every 1000 iterations
if (i%1000 == 0) and (print_cost == True):
current_cost = cost(predict(X, parameters), Y, regularization=regularization, lambd=lambd, parameters=parameters)
print('Cost after {} iterations: {:.12f}'.format(i, current_cost))
# update parameters
if regularization == 'l2':
for i in range(1, L+1):
parameters['W'+str(i)] = (1-lambd/m)*parameters['W'+str(i)] - learning_rate*gradient['dW'+str(i)]
parameters['b'+str(i)] = parameters['b'+str(i)] - learning_rate*gradient['db'+str(i)]
else:
for i in range(1, L+1):
parameters['W'+str(i)] = parameters['W'+str(i)] - learning_rate*gradient['dW'+str(i)]
parameters['b'+str(i)] = parameters['b'+str(i)] - learning_rate*gradient['db'+str(i)]
return parameters
def L_layer_model(X_train, Y_train, X_test, Y_test, n, learning_rate=0.01, iterations=2000, regularization=None, lambd=0, print_cost=False):
# initialize model parameters
parameters = initialize(n)
# fit model
parameters = fit(X_train, Y_train, parameters, learning_rate=learning_rate, iterations=iterations, regularization=regularization, lambd=lambd, print_cost=print_cost)
train_predictions = predict(X_train, parameters)
train_accuracy = 100-np.average(np.abs(Y_train-train_predictions))*100
test_predictions = predict(X_test, parameters)
test_accuracy = 100-np.average(np.abs(Y_test-test_predictions))*100
print('Training set accuracy: {:.4f}%'.format(train_accuracy))
print('Test set accuracy: {:.4f}%'.format(test_accuracy))
return parameters
# ## Test the functions with a random input
# Test
X = np.random.randn(4,5)
Y = np.array([[1, 1, 0, 0, 1]])
n = [4, 6, 5, 4, 3, 7, 2, 1]
parameters = initialize(n)
relu(X)
sigmoid(X)
predict(X, parameters)
# +
# predict?
# -
gradient = propogate(X, Y, parameters)
parameters['b4'] - 0.001*gradient['db4']
parameters = fit(X, Y, parameters, regularization='l2', lambd=0.01)
Yhat = predict(X, parameters)
cost(Yhat, Y, regularization='l2', lambd=0.01, parameters=parameters)
parameters = L_layer_model(X, Y, X, Y, n, learning_rate=0.5, iterations=20000, regularization='l2', lambd=0.1)
parameters['W1']
# ## Build an 'or' network
#
# Test the L layer model by building a simple network to represent an or gate.
X = np.array([[1, 1, 0, 0], [1, 0, 1, 0]])
Y = np.array([[1, 1, 1, 0]])
n = [2, 4, 1]
# model with single hidden layer of four neurons
parameters = L_layer_model(X, Y, X, Y, n, learning_rate=0.9, iterations=2000, regularization='l2', lambd=0.1, print_cost=True)
# add a second hidden layer
n = [2, 4, 2, 1]
parameters = L_layer_model(X, Y, X, Y, n, learning_rate=0.1, iterations=20000, print_cost=False)
# Simple networks are best in this situation
n = [2, 2, 1]
parameters = L_layer_model(X, Y, X, Y, n, learning_rate=0.1, iterations=20000, regularization='l2', lambd=0.01)
predict(np.array([[1], [0]]), parameters)
# After running the above tests successfully, it appears that the L layer model is functional.
# ## Test the L layer model on interview data set
#
# We import an interview data set to investigate the performance of an L layer neural network on a larger data set with more predictors.
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
import pandas as pd
# +
train_location = '' # insert file path here
train_df = pd.read_csv(train_location)
# +
# Clean data. Details can be found in other notebook
train_df = train_df.dropna()
# Correct days so that all are spelled out
train_df['x35'] = train_df['x35'].map(lambda x: 'wednesday' if x=='wed' else 'thursday' if (x=='thur' or x=='thurday') else 'friday' if x=='fri' else x)
# Correct sept. to Sept and Dev to Dec in column x68
train_df['x68'] = train_df['x68'].map(lambda x: 'Jan' if x=='January' else 'Sept' if x=='sept.' else 'Dec' if x=='Dev' else x)
# Transform columns x34, x35, x68, and x93 to dummy variables
train_df = pd.get_dummies(train_df, columns=['x34', 'x35', 'x68', 'x93'])
# Transform columns x41 and x45 to floats
train_df['x41'] = train_df['x41'].map(lambda x: x.lstrip('$'))
train_df['x41'] = pd.to_numeric(train_df['x41'])
train_df['x45'] = train_df['x45'].map(lambda x: x.rstrip('%'))
train_df['x45'] = pd.to_numeric(train_df['x45'])
# Take the first 1000 samples for some quick tests
train_df = train_df.iloc[:1000,:]
train_df.shape
# -
# Note that there are 127 data columns in the data set. The column labeled 'y' contains the true binary class of each training example.
# +
# split data into train/test sets
X_train, X_test, Y_train, Y_test = train_test_split(train_df.drop('y', axis=1), (train_df['y']), test_size=0.2, random_state=42)
# scale data using the standard scaler in sklearn
scaler = StandardScaler()
X_train = scaler.fit_transform(X_train)
X_test = scaler.transform(X_test)
# Get numpy columns for Y
Y_train = Y_train.values.reshape(len(Y_train),1)
Y_test = Y_test.values.reshape(len(Y_test),1)
# -
X_train.shape
# Run model with one hidden layer of three neurons.
# note that samples are in rows of train_df, so the transposes are fed into the models
params = L_layer_model(X_train.T, Y_train.T, X_test.T, Y_test.T, n=[126, 3, 1], learning_rate=0.1, iterations=30000, regularization='l2', lambd=0.01, print_cost=True)
# Results
# * Training accuracy = 98.5%
# * Test accuracy = 84.5%
# We appear to be overfitting the model to the data. We can try increasing the regularization to prevent this problem.
params = L_layer_model(X_train.T, Y_train.T, X_test.T, Y_test.T, n=[126, 3, 1], learning_rate=0.1, iterations=30000, regularization='l2', lambd=0.5, print_cost=True)
# Results indicate that increasing the value of lambd reduces the amount of variance.
params = L_layer_model(X_train.T, Y_train.T, X_test.T, Y_test.T, n=[126, 3, 1], learning_rate=0.1, iterations=30000, regularization='l2', lambd=0.6, print_cost=True)
# Results indicate that increasing the value of lambd further does not necessarily improve performance any more.
#
# We can try models with more hidden layers/neurons.
params = L_layer_model(X_train.T, Y_train.T, X_test.T, Y_test.T, n=[126, 6, 3, 1], learning_rate=0.1, iterations=30000)
params = L_layer_model(X_train.T, Y_train.T, X_test.T, Y_test.T, n=[126, 63, 3, 1], learning_rate=0.1, iterations=30000)
params = L_layer_model(X_train.T, Y_train.T, X_test.T, Y_test.T, n=[126, 63, 3, 1], learning_rate=0.1, iterations=30000, regularization='l2', lambd=0.05)
# Let's compare the neural networks to simple logistic regression.
params = L_layer_model(X_train.T, Y_train.T, X_test.T, Y_test.T, n=[126, 1], learning_rate=0.01, iterations=30000)
params = L_layer_model(X_train.T, Y_train.T, X_test.T, Y_test.T, n=[126, 1], learning_rate=0.1, iterations=30000)
params = L_layer_model(X_train.T, Y_train.T, X_test.T, Y_test.T, n=[126, 1], learning_rate=1, iterations=30000)
params = L_layer_model(X_train.T, Y_train.T, X_test.T, Y_test.T, n=[126, 1], learning_rate=0.005, iterations=30000)
# These results indicate that neural networks don't have performance gains over simple logistic regression when the amount of available data is small.
# Increase the amount of data to 10000 samples to evaluate performance.
# +
train_location = '/Users/connorodell/Documents/Data_Science/learning/exercise_03_train.csv'
train_df = pd.read_csv(train_location)
# +
# Clean data. Details can be found in other notebook
train_df = train_df.dropna()
# Correct days so that all are spelled out
train_df['x35'] = train_df['x35'].map(lambda x: 'wednesday' if x=='wed' else 'thursday' if (x=='thur' or x=='thurday') else 'friday' if x=='fri' else x)
# Correct sept. to Sept and Dev to Dec in column x68
train_df['x68'] = train_df['x68'].map(lambda x: 'Jan' if x=='January' else 'Sept' if x=='sept.' else 'Dec' if x=='Dev' else x)
# Transform columns x34, x35, x68, and x93 to dummy variables
train_df = pd.get_dummies(train_df, columns=['x34', 'x35', 'x68', 'x93'])
# Transform columns x41 and x45 to floats
train_df['x41'] = train_df['x41'].map(lambda x: x.lstrip('$'))
train_df['x41'] = pd.to_numeric(train_df['x41'])
train_df['x45'] = train_df['x45'].map(lambda x: x.rstrip('%'))
train_df['x45'] = pd.to_numeric(train_df['x45'])
train_df = train_df.iloc[:10000,:]
train_df.shape
# +
# split data into train/test sets
X_train, X_test, Y_train, Y_test = train_test_split(train_df.drop('y', axis=1), (train_df['y']), test_size=0.2, random_state=42)
# scale data using the standard scaler in sklearn
scaler = StandardScaler()
X_train = scaler.fit_transform(X_train)
X_test = scaler.transform(X_test)
# Get numpy columns for Y
Y_train = Y_train.values.reshape(len(Y_train),1)
Y_test = Y_test.values.reshape(len(Y_test),1)
# -
X_train.shape
params = L_layer_model(X_train.T, Y_train.T, X_test.T, Y_test.T, n=[126, 63, 3, 1], learning_rate=0.1, iterations=30000, regularization='l2', lambd=0.01)
params = L_layer_model(X_train.T, Y_train.T, X_test.T, Y_test.T, n=[126, 63, 3, 1], learning_rate=0.1, iterations=200000)
# With more data, test accuracy has improved on the order of 10%.
#
# We can see if the logistic regression model has similary performance gains with more data.
params = L_layer_model(X_train.T, Y_train.T, X_test.T, Y_test.T, n=[126, 1], learning_rate=0.1, iterations=30000)
# Results indicate that more data does not improve the logistic regression model. We can interpret this to mean that the model has too much bias.
# Now we test the models with the full data set.
# +
train_location = '/Users/connorodell/Documents/Data_Science/learning/exercise_03_train.csv'
train_df = pd.read_csv(train_location)
# +
# Clean data. Details can be found in other notebook
train_df = train_df.dropna()
# Correct days so that all are spelled out
train_df['x35'] = train_df['x35'].map(lambda x: 'wednesday' if x=='wed' else 'thursday' if (x=='thur' or x=='thurday') else 'friday' if x=='fri' else x)
# Correct sept. to Sept and Dev to Dec in column x68
train_df['x68'] = train_df['x68'].map(lambda x: 'Jan' if x=='January' else 'Sept' if x=='sept.' else 'Dec' if x=='Dev' else x)
# Transform columns x34, x35, x68, and x93 to dummy variables
train_df = pd.get_dummies(train_df, columns=['x34', 'x35', 'x68', 'x93'])
# Transform columns x41 and x45 to floats
train_df['x41'] = train_df['x41'].map(lambda x: x.lstrip('$'))
train_df['x41'] = pd.to_numeric(train_df['x41'])
train_df['x45'] = train_df['x45'].map(lambda x: x.rstrip('%'))
train_df['x45'] = pd.to_numeric(train_df['x45'])
#train_df = train_df.iloc[:10000,:]
#train_df.shape
# +
# split data into train/test sets
X_train, X_test, Y_train, Y_test = train_test_split(train_df.drop('y', axis=1), (train_df['y']), test_size=0.2, random_state=42)
# scale data using the standard scaler in sklearn
scaler = StandardScaler()
X_train = scaler.fit_transform(X_train)
X_test = scaler.transform(X_test)
# Get numpy columns for Y
Y_train = Y_train.values.reshape(len(Y_train),1)
Y_test = Y_test.values.reshape(len(Y_test),1)
# -
# Test various hyperparameters to evaluate relative performance.
params = L_layer_model(X_train.T, Y_train.T, X_test.T, Y_test.T, n=[126, 63, 3, 1], learning_rate=0.1, iterations=2000, regularization='l2', lambd=0.01, print_cost=True)
params = L_layer_model(X_train.T, Y_train.T, X_test.T, Y_test.T, n=[126, 63, 6, 1], learning_rate=0.1, iterations=2000, print_cost=True)
params = L_layer_model(X_train.T, Y_train.T, X_test.T, Y_test.T, n=[126, 63, 6, 1], learning_rate=0.1, iterations=30000, regularization='l2', lambd=0.01)
params = L_layer_model(X_train.T, Y_train.T, X_test.T, Y_test.T, n=[126, 63, 10, 10, 1], learning_rate=0.1, iterations=30000, regularization='l2', lambd=0.01)
params = L_layer_model(X_train.T, Y_train.T, X_test.T, Y_test.T, n=[126, 63, 6, 1], learning_rate=0.1, iterations=30000, regularization='l2', lambd=0.05)
params = L_layer_model(X_train.T, Y_train.T, X_test.T, Y_test.T, n=[126, 63, 6, 1], learning_rate=0.1, iterations=30000, regularization='l2', lambd=0.05)
params = L_layer_model(X_train.T, Y_train.T, X_test.T, Y_test.T, n=[126, 63, 6, 1], learning_rate=0.1, iterations=30000, regularization='l2', lambd=0.05)
params = L_layer_model(X_train.T, Y_train.T, X_test.T, Y_test.T, n=[126, 63, 6, 1], learning_rate=0.1, iterations=30000, regularization='l2', lambd=0.06)
params = L_layer_model(X_train.T, Y_train.T, X_test.T, Y_test.T, n=[126, 63, 6, 1], learning_rate=0.1, iterations=30000, regularization='l2', lambd=0.07)
params = L_layer_model(X_train.T, Y_train.T, X_test.T, Y_test.T, n=[126, 63, 6, 1], learning_rate=0.1, iterations=30000, regularization='l2', lambd=0.08)
params = L_layer_model(X_train.T, Y_train.T, X_test.T, Y_test.T, n=[126, 63, 6, 1], learning_rate=0.1, iterations=30000, regularization='l2', lambd=0.09)
params = L_layer_model(X_train.T, Y_train.T, X_test.T, Y_test.T, n=[126, 63, 6, 1], learning_rate=0.1, iterations=30000, regularization='l2', lambd=0.1)
# Compare to simple linear regression on the full data set.
params = L_layer_model(X_train.T, Y_train.T, X_test.T, Y_test.T, n=[126, 1], learning_rate=0.1, iterations=30000)
# Again, simple linear regression sees no performance gains from additional data.
|
DeepLearning/L_layer_network.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# # Leaderboard statistics exploration
# %pylab inline
import pandas as pd
import seaborn as sns
from __future__ import print_function
import datetime
df = pd.read_csv('./data/predict-west-nile-virus_public_leaderboard.csv');
df['SubmissionDate'] = pd.to_datetime(df['SubmissionDate'])
df['Date'] = df['SubmissionDate'].dt.date
total = len(df['TeamName'].unique())
print("total number of teams: {}".format(total))
len(df.index)
winner = df.sort_values(by=['Score'],ascending=False).iloc[0]
print("Leaderboard winner : {}, score: {}".format(winner['TeamName'],winner['Score']));
plt.plot(df['SubmissionDate'],df['Score'],'bo',alpha=0.01)
plt.figure(figsize=(8,6))
ndf = df.groupby("Date").mean().reset_index()
ndf['Date'] = pd.to_datetime(ndf['Date'])
plt.plot(ndf["Date"], ndf["Score"],'bo');
plt.xlim(datetime.datetime(2015,4,22),datetime.datetime(2015,6,17));
plt.xlabel('Date');
plt.ylabel('Daily average score');
plt.xticks(rotation=45,ha='right');
df.columns
max_scores = df.groupby('TeamName').max()['Score'].get_values()
no_submissions = df.groupby('TeamName').count()['Score'].get_values()
lbs = pd.DataFrame()
lbs['MaxScore'] = max_scores
lbs['NoSubmissions'] = no_submissions
plt.figure(figsize=(8,6))
sns.regplot(x="NoSubmissions", y="MaxScore", data=lbs,x_estimator=np.mean,fit_reg=False)
plt.ylim(0.5,0.9);
plt.xlim(0,70);
# ## No. of submissions per team
plt.figure(figsize=(8,6))
plt.hist(no_submissions,bins=50);
plt.ylabel('Count');
plt.xlabel('Submissions');
# ## Track the top ten competition teams
winners = df.groupby('TeamName').max().reset_index().\
sort_values(by=['Score'],ascending=False).iloc[:10]['TeamName']
# +
#plot background
sns.set_palette(sns.color_palette("Set2", 10));
plt.figure(figsize=(8,6))
ndf = df.groupby("Date").mean().reset_index()
ndf['Date'] = pd.to_datetime(ndf['Date'])
plt.plot(ndf["Date"], ndf["Score"],'bo',label='Average');
plt.xlabel('Date');
plt.ylabel('Daily average score');
#plot team progressions
ndf = df.groupby(["Date","TeamName"]).max().reset_index()
for winner in winners:
series = ndf[ndf["TeamName"]==winner]
plt.plot(series['Date'],series['Score'],label=winner);
plt.legend(loc=0);
plt.xlim(datetime.datetime(2015,4,22),datetime.datetime(2015,6,17));
plt.xticks(rotation=45,ha='right');
# -
|
Leaderboard_Statistics_Exploration.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
sys.path.append("..") # Adds higher directory to python modules path.
from pathlib import Path
import glob
import numpy as np
import tensorflow as tf
import pickle
import matplotlib.pyplot as plt
import random
import pickle
import os
import config
import data
import random
from natsort import natsorted
import lfp
import gym
arm = 'UR5'
TEST_DATASET = "UR5_slow_gripper_test"
print('Using local setup')
WORKING_PATH = Path().absolute().parent
print(f'Working path: {WORKING_PATH}')
os.chdir(WORKING_PATH)
STORAGE_PATH = WORKING_PATH
print(f'Storage path: {STORAGE_PATH}')
TRAIN_DATA_PATHS = [STORAGE_PATH/'data'/x for x in ["pybullet/UR5" , "pybullet/UR5_high_transition" ,"pybullet/UR5_slow_gripper"]]
TEST_DATA_PATH = STORAGE_PATH/'data'/TEST_DATASET
# -
import roboticsPlayroomPybullet
env = gym.make('UR5PlayAbsRPY1Obj-v0')
env.render('human')
_ = env.reset()
env.render('playback')
env.instance.calc_state()['observation'][0:7]
env.step(np.array([ -1.91859640e-02, 1.93180365e-01, 0.2, 0.0,
0.0, 0.0, -7.02553025e-06]))
plt.figure(figsize = (20,20))
plt.imshow(env.instance.calc_state()['img'][:,:,:])
# # Replays the teleop data
#
# - This little loop of code replays the teleop data, and optionally saves the images to create an image dataset
# - Every 30 steps it resets state, because minor errors in the physics compound
TRAIN_DATA_PATHS
# +
for DIR in TRAIN_DATA_PATHS:
DIR = str(DIR)
# DIR = str(TRAIN_DATA_PATHS[0]) # glob/natsorted prefer strings
obs_act_path = DIR+'/obs_act_etc/'
o, a, ag = [], [], []
for demo in natsorted(os.listdir(obs_act_path)):
traj = np.load(obs_act_path+demo+'/data.npz')
print(demo, len(traj['obs']))
o.append(traj['obs']), a.append(traj['acts']), ag.append(traj['achieved_goals'])
print('________________________', len(np.vstack(o)))
o, a, ag = np.vstack(o), np.vstack(a), np.vstack(ag)
# +
import time
jp = traj['joint_poses']
ag = traj['achieved_goals']
for i in range(0, len(jp)):
time.sleep(0.02)
env.instance.reset_arm_joints(env.instance.arm, jp[i,:])
env.instance.reset_object_pos(ag[])
# -
o.shape
env.reset(o[0,:])
d = a
for i in range(0, d.shape[1]):
plt.hist(d[:,i], bins=1000)
#plt.xlim(-0.2,0.2)
plt.show()
d = a - o[:, :7]
for i in range(0, d.shape[1]):
plt.hist(d[:,i], bins=1000)
plt.xlim(-0.2,0.2)
plt.show()
d = d[1:] - d[:-1]
d = o[150000:150020]
f = a[150000:150020]
for i in range(0, d.shape[1]):
plt.plot(np.linspace(0,len(d),len(d)), d[:,i])
plt.plot(np.linspace(0,len(d),len(d)), f[:,i])
plt.show()
# +
import scipy.misc
from IPython.display import display, clear_output
keys = ['obs', 'acts', 'achieved_goals', 'joint_poses', 'target_poses', 'acts_quat', 'acts_rpy_rel', 'velocities', 'obs_quat']
#
for DIR in TRAIN_DATA_PATHS:
DIR = str(DIR) # glob/natsorted prefer strings
obs_act_path = DIR+'/obs_act_etc/'
for demo in natsorted(os.listdir(obs_act_path)):
print(demo)
start_points = natsorted(glob.glob(DIR+'/states_and_ims/'+str(demo)+'/env_states/*.bullet'))
traj = np.load(obs_act_path+demo+'/data.npz')
d = {k:traj[k] for k in keys}
acts = d['acts']
set_len = len(acts)
start = 0
end= min(start+30, set_len)
print(DIR+'/states_and_ims/'+str(demo)+'/ims')
try:
os.makedirs(DIR+'/states_and_ims/'+str(demo)+'/ims')
except:
pass
for start_point in start_points:
env.p.restoreState(fileName=start_point)
env.instance.updateToggles() # need to do it when restoring, colors not carried over
for i in range(start, end):
o,r,_,_ = env.step(acts[i])
start += 30
end = min(start+30, set_len)
# -
# +
import scipy.misc
from IPython.display import display, clear_output
keys = ['obs', 'acts', 'achieved_goals', 'joint_poses', 'target_poses', 'acts_quat', 'acts_rpy_rel', 'velocities', 'obs_quat', 'gripper_proprioception']
#
for DIR in TRAIN_DATA_PATHS:
obs_act_path = DIR/'obs_act_etc/'
obs_act_path2 = DIR + 'obs_act_etc2/'
for demo in natsorted(os.listdir(obs_act_path)):
print(demo)
start_points = natsorted(glob.glob(DIR+'/states_and_ims/'+str(demo)+'/env_states/*.bullet'))
traj = np.load(obs_act_path+demo+'/data.npz')
d = {k:traj[k] for k in keys}
acts = d['acts']
set_len = len(acts)
start = 0
end= min(start+30, set_len)
print(DIR+'/states_and_ims/'+str(demo)+'/ims')
try:
os.makedirs(DIR+'/states_and_ims/'+str(demo)+'/ims')
except:
pass
for start_point in start_points:
env.p.restoreState(fileName=start_point)
env.panda.updateToggles() # need to do it when restoring, colors not carried over
for i in range(start, end):
#scipy.misc.imsave(DIR+'/states_and_ims/'+str(demo)+'/ims/'+str(i)+'.jpg', o['img'])
o,r,_,_ = env.step(acts[i])
# clear_output(wait=True)
# fig = plt.imshow(scipy.misc.imread(DIR+'/states_and_ims/'+str(demo)+'/ims/'+str(i)+'.jpg'))
# plt.show()
#time.sleep(0.05)
start += 30
end = min(start+30, set_len)
# try:
# os.makedirs(obs_act_path2+demo)
# except:
# pass
# np.savez(obs_act_path2+demo+'/data', obs=d['obs'], acts=d['acts'], achieved_goals=d['achieved_goals'],
# joint_poses=d['joint_poses'],target_poses=d['target_poses'], acts_quat=d['acts_quat'],
# acts_rpy_rel=d['acts_rpy_rel'], velocities = d['velocities'],
# obs_quat=d['obs_quat'], gripper_proprioception=d['gripper_proprioception'])
# +
env.p.restoreState(fileName=path)
vid_path = 'output/videos/trajectory.mp4'
with imageio.get_writer(vid_path, mode='I') as writer:
for i in range(start, start+WINDOW_SIZE):
o ,r, d, _ = env.step(actions[i,:])
writer.append_data(o['img'])
clear_output(wait=True)
fig = plt.imshow(o['img'])
plt.show()
# +
keys = ['obs', 'acts', 'achieved_goals', 'joint_poses', 'target_poses', 'acts_quat', 'acts_rpy_rel', 'velocities', 'obs_quat', 'gripper_proprioception']
for DIR in [TRAIN_DATA_PATHS[1]]:
obs_act_path = os.path.join(DIR, 'obs_act_etc/')
for demo in natsorted(os.listdir(obs_act_path)):
if int(demo)>18:
print(demo)
start_points = natsorted(glob.glob(str(DIR/'states_and_ims'/str(demo)/'env_states/*.bullet')))
traj = np.load(obs_act_path+demo+'/data.npz')
d = {k:traj[k] for k in keys}
acts = d['acts']
set_len = len(acts)
start = 0
end= min(start+30, set_len)
gripper_proprioception = []
for start_point in start_points:
env.p.restoreState(fileName=start_point)
for i in range(start, end):
o,r,_,_ = env.step(acts[i])
#print(d['gripper_proprioception'][i])
time.sleep(0.015)
start += 30
end = min(start+30, set_len)
# +
#dataset, cnt = data.create_single_dataset(dataset_path)
def load_data(path, keys):
dataset = {k:[] for k in keys+['sequence_index','sequence_id']}
obs_act_path = os.path.join(path, 'obs_act_etc/')
for demo in natsorted(os.listdir(obs_act_path)):
print(demo)
traj = np.load(obs_act_path+demo+'/data.npz')
for k in keys:
d = traj[k]
if len(d.shape) < 2:
d = np.expand_dims(d, axis = 1) # was N, should be N,1
dataset[k].append(d.astype(np.float32))
timesteps = len(traj['obs'])
dataset['sequence_index'].append(np.arange(timesteps, dtype=np.int32).reshape(-1, 1))
dataset['sequence_id'].append(np.full(timesteps, fill_value=int(demo), dtype=np.int32).reshape(-1, 1))
# convert to numpy
for k in keys+['sequence_index','sequence_id']:
dataset[k] = np.vstack(dataset[k])
return dataset
keys = ['obs', 'acts', 'achieved_goals', 'joint_poses', 'target_poses', 'acts_rpy', 'acts_rpy_rel', 'velocities', 'obs_rpy', 'obs_rpy_inc_obj', 'gripper_proprioception']
dataset = load_data(UR5, keys)
#transition_dataset = load_data(UR5_25, keys)
# -
import tensorflow_probability as tfp
tfd = tfp.distributions
tfb = tfp.bijectors
tfpl = tfp.layers
scaling = np.array([256.0/4, 256.0/2]).astype(np.float32)
# +
def logistic_mixture(inputs, quantized = True):
weightings, mu, scale = inputs
print(mu.shape, scaling.shape, scale.shape, weightings.shape)
mu = mu*np.expand_dims(scaling,1)
print(mu)
dist = tfd.Logistic(loc=mu, scale=scale)
if quantized:
dist = tfd.QuantizedDistribution(
distribution=tfd.TransformedDistribution(
distribution=dist,
bijector=tfb.Shift(shift=-0.5)),
low=-128.,
high=128.
)
mixture_dist = tfd.MixtureSameFamily(
mixture_distribution=tfd.Categorical(logits=weightings),
components_distribution=dist,
validate_args=True
)
print(mixture_dist)
if quantized:
quantized_scale = 1/scaling
mixture_dist = tfd.TransformedDistribution(
distribution=mixture_dist,
bijector=tfb.Scale(scale=quantized_scale)
)
return mixture_dist
mu = np.array([[[-1.5, 0.4, 0.4],[-0.2, 0.3, 0.3]]]).astype(np.float32)
std = np.array([[[1.0,1.0,1],[1.0,1.0,1]]]).astype(np.float32)
weights = np.array([[[1,1,1],[1,1,1]]]).astype(np.float32)
m = logistic_mixture((weights,mu,std))
#m = logistic_mixture(([1], [0.06], [1]))
# -
m.sample()
samples = np.array([m.sample().numpy() for i in range(0,100)])
samples.shape
samples[:,0]
plt.hist(np.array(samples[:,:,0]), bins=100)
plt.plot(np.linspace(-0.5, 0.5, 100),m.log_prob(np.linspace(-0.5, 0.5, 100)))
# +
# Coverage analysis
np.set_printoptions(suppress=True)
ag = dataset['achieved_goals']
t_ag = transition_dataset['achieved_goals']
# +
def see_diff(ag):
diff_ag = abs(np.sum(ag[1:]-ag[:-1],axis = -1))
print(sum(diff_ag == 0))
plt.plot(diff_ag)
see_diff(ag[:150000])
see_diff(t_ag[:150000])
# +
mins = np.min(dataset['achieved_goals'], axis = 0)
maxes = np.max(dataset['achieved_goals'], axis = 0)
bins = np.linspace(mins,maxes+0.01, 11)
idx = 0
qs = []
for idx in range(0,ag.shape[1]):
quantiles = np.digitize(dataset['achieved_goals'][:,idx], bins[:,idx])
qs.append(quantiles)
qs = np.array(qs).T
qs.shape
np.unique(qs, axis=0).shape[0]
# -
from tqdm import tqdm
step2 = []
count2 = []
for i in tqdm(np.linspace(1, len(qs), 10)):
i = int(i)
step2.append(i)
count2.append(np.unique(qs[:i], axis=0).shape[0])
import matplotlib.pyplot as plt
#plt.plot(step, count)
plt.plot(step2, count2)
import matplotlib.pyplot as plt
plt.plot(step, count)
d['']
# +
print(obs_act_path2+demo)
try:
os.makedirs(obs_act_path2+demo)
except:
pass
np.savez(obs_act_path2+demo+'/data', obs=d['obs'], acts=d['acts'], achieved_goals=d['achieved_goals'],
joint_poses=d['joint_poses'],target_poses=d['target_poses'], acts_rpy=d['acts_rpy'],
acts_rpy_rel=d['acts_rpy_rel'], velocities = d['velocities'],
obs_rpy=d['obs_rpy'], gripper_proprioception=d['gripper_proprioception'])
# -
d['obs']
np.load(obs_act_path2+demo+'/data.npz', allow_pickle=True)['obs']
os.make_dirs(obs_act_path2)
env.step(acts[i])
print(start_points)
rpy_obs = 'obs_rpy' #'rpy_obs'
def load_data(path, keys):
dataset = {k:[] for k in keys+['sequence_index','sequence_id']}
obs_act_path = os.path.join(path, 'obs_act_etc/')
for demo in natsorted(os.listdir(obs_act_path)):
print(demo)
traj = np.load(obs_act_path+demo+'/data.npz')
for k in keys:
dataset[k].append(traj[k].astype(np.float32))
timesteps = len(traj['obs'])
dataset['sequence_index'].append(np.arange(timesteps, dtype=np.int32).reshape(-1, 1))
dataset['sequence_id'].append(np.full(timesteps, fill_value=int(demo), dtype=np.int32).reshape(-1, 1))
# convert to numpy
for k in keys+['sequence_index','sequence_id']:
dataset[k] = np.vstack(dataset[k])
return dataset
keys = ['obs', 'acts', 'achieved_goals', 'joint_poses', 'target_poses', 'acts_rpy', 'acts_rpy_rel', 'velocities', 'obs_rpy']
dataset = load_data(PYBULLET_DATA_DIR, keys)
# +
obs_act_path = os.path.join(path, 'obs_act_etc/')
starts = []
idxs = []
fs = []
for f in natsorted(os.listdir(obs_act_path)):
potential_start_points = glob.glob(TEST_DIR+'/states_and_ims/'+str(f)+'/env_states/*.bullet')
potential_start_idxs = [int(x.replace('.bullet','').replace(f"{TEST_DIR}/states_and_ims/{str(f)}/env_states/", "")) for x in potential_start_points]
folder = [f]*len(potential_start_idxs)
[starts.append(x) for x in potential_start_points], [idxs.append(x) for x in potential_start_idxs], [fs.append(x) for x in folder]
# +
descriptions = {
1: 'lift up',
2: 'take down',
3: 'door left',
4: 'door right',
5: 'drawer in',
6: 'drawer out',
7: 'pick place',
8: 'press button',
9: 'dial on',
10: 'dial off',
11: 'rotate block left',
12: 'rotate block right',
13: 'stand up block',
14: 'knock down block',
15: 'block in cupboard right',
16: 'block in cupboard left',
17: 'block in drawer',
18: 'block out of drawer',
19: 'block out of cupboard right',
20: 'block out of cupboard left',
}
# +
trajectory_labels = {}
done = []
# +
import time
for i in range(0,len(starts)):
if starts[i] not in done:
data = np.load(TEST_DIR+'obs_act_etc/'+str(fs[i])+'/data.npz')
traj_len = 40#random.randint(40,50)
end = min(len(data['acts'])-1,idxs[i]+traj_len )
acts = data['acts_rpy'][idxs[i]:end]
value = "r"
while value == "r":
env.p.restoreState(fileName=starts[i])
for a in range(0, len(acts)):
env.step(acts[a])
time.sleep(0.01)
value = input("Label:")
if value == 's':
break
elif value == 'r':
pass
else:
trajectory_labels[starts[i]] = descriptions[int(value)]
done.append(starts[i])
np.savez("trajectory_labels", trajectory_labels=trajectory_labels, done=done)
# -
len(starts)
for k,v in trajectory_labels.items():
if v == 'knock':
trajectory_labels[k] = 'knock down block'
starts[i]
left = np.load(TEST_DIR+'left_right.npz')['left']
right = np.load(TEST_DIR+'left_right.npz')['right']
# +
left_complete = []
right_complete = []
for pth in left:
f = pth.split('/')[7]
i = pth.split('/')[9].replace('.bullet', '')
data = np.load(TEST_DIR+'obs_act_etc/'+f+'/data.npz')
o = data['obs'][int(i):int(i)+40]
a = data['acts_rpy'][int(i):int(i)+40]
pth = pth.replace('/content/drive/My Drive/Robotic Learning/UR5_25Hz_test_suite/', TEST_DIR)
left_complete.append((pth, o, a))
for pth in right:
f = pth.split('/')[7]
i = pth.split('/')[9].replace('.bullet', '')
data = np.load(TEST_DIR+'obs_act_etc/'+f+'/data.npz')
o = data['obs'][int(i):int(i)+40]
a = data['acts_rpy'][int(i):int(i)+40]
pth = pth.replace('/content/drive/My Drive/Robotic Learning/UR5_25Hz_test_suite/', TEST_DIR)
right_complete.append((pth, o, a))
# -
for i in range(0,50):
pth, obs, acts = left_complete[np.random.choice(len(left_complete))]
env.p.restoreState(fileName=pth)
for a in range(0, len(acts)):
env.step(acts[a])
time.sleep(0.001)
for i in range(0,50):
pth, obs, acts = right_complete[np.random.choice(len(right_complete))]
env.p.restoreState(fileName=pth)
for a in range(0, len(acts)):
env.step(acts[a])
time.sleep(0.001)
obs_left = np.array([x[1] for x in left_complete])
obs_right = np.array([x[1] for x in right_complete])
# +
import seaborn as sns
fig, axs = plt.subplots(ncols=4, nrows=5,figsize=(20, 20),)
for x in range(0, obs_left.shape[2]):
shape = obs_left.shape
sns.distplot(np.reshape(obs_left[:], [shape[0] * shape[1], shape[2]])[:,x], hist=True, kde=True,
bins=int(180/5), color = 'darkblue',
hist_kws={'edgecolor':'black'},
kde_kws={'linewidth': 4}, ax=axs[mapping[x][0], mapping[x][1]])
shape = obs_right.shape
sns.distplot(np.reshape(obs_right[:], [shape[0] * shape[1], shape[2]])[:,x], hist=True, kde=True,
bins=int(180/5), color = 'orange',
hist_kws={'edgecolor':'orange'},
kde_kws={'linewidth': 4}, ax=axs[mapping[x][0], mapping[x][1]])
plt.show()
# +
acts_left = np.array([x[2] for x in left_complete])
acts_right = np.array([x[2] for x in right_complete])
import seaborn as sns
fig, axs = plt.subplots(ncols=4, nrows=2,figsize=(20, 20),)
for x in range(0, acts_left.shape[2]):
shape = acts_left.shape
sns.distplot(np.reshape(acts_left[:], [shape[0] * shape[1], shape[2]])[:,x], hist=True, kde=True,
bins=int(180/5), color = 'darkblue',
hist_kws={'edgecolor':'black'},
kde_kws={'linewidth': 4}, ax=axs[mapping[x][0], mapping[x][1]])
shape = acts_right.shape
sns.distplot(np.reshape(acts_right[:], [shape[0] * shape[1], shape[2]])[:,x], hist=True, kde=True,
bins=int(180/5), color = 'orange',
hist_kws={'edgecolor':'orange'},
kde_kws={'linewidth': 4}, ax=axs[mapping[x][0], mapping[x][1]])
plt.show()
# +
mapping = []
for i in range(0,5):
for j in range(0,4):
mapping.append([i,j])
mapping
# -
obs_left.shape[2]-1
# +
arm_pos = [0.29, -0.01, 0.51]
b= [0.25, 0.11, 0.02]
realsense_y= translation[2] - bb[0]
realsense_x = translation[1] - bb[1]
realsense_z = translation[0] - bb[2]
# -
# Testing camera transforms
camera_coord = (20,20)
plt.scatter(camera_coord[0], 480-camera_coord[1], s=40)
plt.xlim(0,480)
plt.ylim(0,480)
import math
def gripper_frame_to_robot_frame(x,y, angle):
y=-y
X = x*math.cos(angle) - y*math.sin(angle)
Y = x*math.sin(angle) + y*math.cos(angle)
return X, Y
current_angle = 0.22
gripper_frame_to_robot_frame(0.02,-0.02, math.pi/2)
path = os.getcwd()+ '/sapien_simulator/config/ur5e.srdf' # '/ocrtoc_task/urdf/ur5e.urdf'
p.loadURDF(path)
#
height =
# +
os.path.exists(path)
# +
# Testing that diversity does increase with more training data
t_it = iter(train_dataset)
mins = np.min(dataset['obs_rpy'], axis = 0)
maxes = np.max(dataset['obs_rpy'], axis = 0)
shape = dataset['obs_rpy'].shape[1]
bins = np.linspace(mins,maxes+0.01, 11)
def get_quantisation(ags, bins):
qs = []
for idx in range(0 , shape):
quantiles = np.digitize(ags[:, idx], bins[:,idx])
qs.append(quantiles)
return np.array(qs).T
batch = t_it.next()
o = tf.reshape(batch['obs'][:,:,:], (-1, OBS_DIM))
coverage = get_quantisation(o, bins)
shapes = []
for i in range(0,10):
batch = t_it.next()
o = tf.reshape(batch['obs'][:,:,:], (-1, OBS_DIM))
c = get_quantisation(o, bins)
coverage = np.unique(np.concatenate([coverage, c], 0), axis = 0)
shapes.append(coverage.shape[0])
np.unique(get_quantisation(dataset['obs_rpy'], bins), axis = 0).shape
plt.plot([120215]*11)
plt.plot(old)
plt.plot(shapes)
plt.plot(one)
plt.title("Unique states observed in batches with shuffle size N")
plt.legend(['Unique values', 40, 10, 1])
|
experimental/Review and Analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# When you submit this code in Coding Ninjas it raises TLE for last two test cases. Do you know why?
# Exactly swapping takes most of the time
# // Refer the optimised code which is written below of this code//
class Node:
def __init__(self, data):
self.data = data
self.next = None
def length(head):
cnt = 0
while head:
cnt += 1
head = head.next
return cnt
def ele(head, index):
cnt = 0
curr = head
while curr:
if cnt == index:
return curr.data
cnt += 1
curr = curr.next
return curr.data
def swap_nodes(head, i, j):
if ((i == 0 or j == 0) and (abs(i-j) == 1)):
curr1 = head
head = head.next
adv1 = head.next
head.next = curr1
curr1.next = adv1
elif i == 0 or j == 0:
if i == 0:
curr1 = head
ptr2 = head
cnt = 1
while cnt < j:
cnt += 1
ptr2 = ptr2.next
curr2 = ptr2.next
adv1 = curr1.next
adv2 = curr2.next
head = curr2
curr2.next = adv1
ptr2.next = curr1
curr1.next = adv2
elif j == 0:
curr2 = head
ptr1 = head
cnt = 1
while cnt < i:
cnt += 1
ptr1 = ptr1.next
curr1 = ptr1.next
adv2 = curr2.next
adv1 = curr1.next
head = curr1
curr1.next = adv2
ptr1.next = curr2
curr2.next = adv1
elif abs(i-j) == 1:
if i < j:
cnt1 = 1
cnt2 = 1
ptr1 = head
ptr2 = head
while cnt1 < i:
cnt1 += 1
ptr1 = ptr1.next
curr1 = ptr1.next
adv1 = curr1.next
while cnt2 < j:
cnt2 += 1
ptr2 = ptr2.next
curr2 = ptr2.next
adv2 = curr2.next
ptr1.next = curr2
curr2.next = curr1
curr1.next = adv2
elif j < i:
cnt1 = 1
cnt2 = 1
ptr1 = head
ptr2 = head
while cnt2 < j:
cnt2 += 1
ptr2 = ptr2.next
curr2 = ptr2.next
adv2 = curr2.next
while cnt1 < i:
cnt1 += 1
ptr1 = ptr1.next
curr1 = ptr1.next
adv1 = curr1.next
ptr2.next = curr1
curr1.next = curr2
curr2.next = adv1
else:
cnt1, cnt2 = 1,1
ptr1, ptr2 = head, head
while cnt2 < j:
cnt2 += 1
ptr2 = ptr2.next
curr2 = ptr2.next
adv2 = curr2.next
while cnt1 < i:
cnt1 += 1
ptr1 = ptr1.next
curr1 = ptr1.next
adv1 = curr1.next
ptr1.next = curr2
curr2.next = adv1
ptr2.next = curr1
curr1.next = adv2
return head
def bubbleSortLL(head):
leng = length(head)
for i in range(0, leng - 1):
for j in range(0, leng - i - 1):
if ele(head,j) > ele(head,j+1):
head = swap_nodes(head, j, j+1)
return head
def ll(arr):
if len(arr) == 0:
return None
head = Node(arr[0])
tail = head
for data in arr[1:]:
tail.next = Node(data)
tail = tail.next
return head
def printll(head):
while head:
print(head.data, end = " ")
head = head.next
print()
arr = list(int(i) for i in input().strip().split(' '))
l = ll(arr[:-1])
l = bubbleSortLL(l)
printll(l)
# +
# // Optimised code //
class Node:
def __init__(self, data):
self.data = data
self.next = None
def bubbleSortLL(head) :
end = None
while end != head:
p = head
while p.next != end:
q = p.next
if p.data > q.data:
p.data, q.data = q.data, p.data
p = p.next
end = p
return end
def ll(arr):
if len(arr) == 0:
return None
head = Node(arr[0])
tail = head
for data in arr[1:]:
tail.next = Node(data)
tail = tail.next
return head
def printll(head):
while head:
print(head.data, end = " ")
head = head.next
print()
arr = list(int(i) for i in input().strip().split(' '))
l = ll(arr[:-1])
l = bubbleSortLL(l)
printll(l)
# -
|
10 Linked List-2/10.14 Bubble Sort Using Linked List.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Tidy Data In Python
#
# A Python exercise motivated by this nice article from <NAME>: [Tidy Data](http://vita.had.co.nz/papers/tidy-data.pdf). reading this paper is a nice prerequisite for this notebook.
#
# The code in this notebook was originally developed and commented by [<NAME>](https://www.ibm.com/developerworks/community/blogs/jfp?lang=en) in a blog post on "[Tidy Data In Python](https://www.ibm.com/developerworks/community/blogs/jfp/entry/Tidy_Data_In_Python?lang=en)".
#
# Let's start.
#
# We only need two Python packages here.
import pandas as pd
import numpy as np
# Let's show readers which versions we are using.
pd.__version__
# ## Introduction
#
# A messy data set. *Messy* is used as defined in Hadley Wickham's paper: any data set that is not tidy. Messy data sets are often convenient for showing them to human as they are compact. This form is often used in publications.
messy = pd.DataFrame({'First' : ['John', 'Jane', 'Mary'],
'Last' : ['Smith', 'Doe', 'Johnson'],
'Treatment A' : [np.nan, 16, 3],
'Treatment B' : [2, 11, 1]})
messy
# People may prefer the transpose view of that data set
messy.T
# Messy data sets aren't that easy to process by statistical or machine learning packages. These often assume that examples are provided as rows in a matrix whose columns are example featureThis is precisely what a tidy data set is. Applying the `melt()` function to it creates a tidy version of it. We sort the result by name to make it easier to read.
tidy = pd.melt(messy, id_vars=['First','Last'])
tidy
# The values are fine but column names aren't really meaningful. Fortinately, the `melt()` function has arguments for renaming them.
tidy = pd.melt(messy, id_vars=['First','Last'], var_name='treatment', value_name='result')
tidy
# ## A simple melt example
messy = pd.DataFrame({'row' : ['A', 'B', 'C'],
'a' : [1, 2, 3],
'b' : [4, 5, 6],
'c' : [7, 8, 9]})
messy
pd.melt(messy, id_vars='row')
tidy = pd.melt(messy, id_vars='row', var_name='dimension', value_name='length')
tidy
# Pivot is almost the inverse of melt
messy1 = tidy.pivot(index='row',columns='dimension',values='length')
messy1
# This is almost the same as the orginal dataframe, except that row is used as index. We can move it back to a row easily.
messy1.reset_index(inplace=True)
messy1
# Last step is to remove the name for the set of columns.
messy1.columns.name = ''
messy1
# This is the same as the original dataframe, up to column reordering.
# ## Column headers are values, not variable names
#
# This is the first issue with messy data in Hadley's paper. Let's first create the dataframe used as an example. For practical reasons, it was simpler to first construct the transpose of it, then process it to get the data set used in the article.
# +
messy = pd.DataFrame({'Agnostic' : [27, 34, 60, 81, 76, 137],
'Atheist' : [12, 27, 37, 52, 35, 70],
'Buddhist' : [27, 21, 30, 34, 33, 58],
'Catholic' : [418, 617, 732, 670, 638, 1116],
"Don't know/refused" : [15, 14, 15, 11, 10, 35],
'Evangelical Prot' : [575, 869, 1064, 982, 881, 1486],
'Hindu' : [1, 9, 7, 9, 11, 34],
'Historically Black Prot' : [228, 244, 236, 238, 197, 223],
"Jehovah's Witness" : [20, 27, 24, 24, 21, 30],
'Jewish' : [19, 19, 25, 25, 30, 95],
})
def transpose(df, columns):
df = df.T.copy()
df.reset_index(inplace=True)
df.columns = columns
return df
messy = transpose(messy, ['religion', '<$10k', '$10-20k', '$20-30k', '$30-40k', '$40-50k', '$50-75k'])
messy
# -
# Again, the `melt()` function is our friend. We sort the result by religion to make it easier to read.
tidy = pd.melt(messy, id_vars = ['religion'], var_name='income', value_name='freq')
tidy.sort_values(by=['religion'], inplace=True)
tidy.head()
# ## Variables are stored in both rows and columns
#
# This example is a little trickier. We first read the input data as a data frame. This data is available at https://github.com/hadley/tidy-data/blob/master/data/tb.csv
#
# I've cloned it therefore it is in my local `data` directory.
#
# Reading it is easy. We remove the `new_sp_` prefix appearing in most columns, and we rename a couple of columns as well.
url = "https://raw.githubusercontent.com/hadley/tidy-data/master/data/tb.csv"
tb = pd.read_csv(url)
tb.columns = tb.columns.str.replace('new_sp_','')
tb.rename(columns = {'new_sp' : 'total', 'iso2' : 'country'},
inplace=True)
tb.head()
# Let's use year 2000, and drop few columns, to stay in sync with Wickham's article.
messy = tb[tb['year'] == 2000].copy()
messy.drop(['total','m04','m514','f04','f514'], axis=1, inplace=True)
messy.head(10)
messy.iloc[:,:11].head(10)
# The `melt()` function is useful, but is not enough. Let's use it still.
molten = pd.melt(messy, id_vars=['country', 'year'], value_name='cases')
molten.sort_values(by=['year', 'country'], inplace=True)
molten.head(10)
# What isn't really nice is the encoding of sex and age ranges as a string in the `variable` column. Let's process the dataset to create two additional columns, one for the sex, and one for the age range. We then remove the `variable` column. The tidy form also makes it easy to remove the values where the age is `u`.
# +
def parse_age(s):
s = s[1:]
if s == '65':
return '65+'
else:
return s[:-2]+'-'+s[-2:]
tidy = molten[molten['variable'] != 'mu'].copy()
tidy['sex'] = tidy['variable'].apply(lambda s: s[:1])
tidy['age'] = tidy['variable'].apply(parse_age)
tidy = tidy[['country', 'year', 'sex', 'age', 'cases']]
tidy.head(10)
# -
# ## Variables are stored in both rows and columns
#
# This example is really tricky. Let's first create the dataframe. This time, I create it using an array instead of a dictionary, just for the fun of doing something a bit different.
columns = ['id', 'year', 'month', 'element', 'd1', 'd2', 'd3', 'd4', 'd5', 'd6', 'd7', 'd8']
data = [['MX17004', 2010, 1, 'tmax', np.nan, np.nan, np.nan, np.nan, np.nan, np.nan, np.nan, np.nan, ],
['MX17004', 2010, 1, 'tmin', np.nan, np.nan, np.nan, np.nan, np.nan, np.nan, np.nan, np.nan, ],
['MX17004', 2010, 2, 'tmax', np.nan, 27.3, 24.1, np.nan, np.nan, np.nan, np.nan, np.nan, ],
['MX17004', 2010, 2, 'tmin', np.nan, 14.4, 14.4, np.nan, np.nan, np.nan, np.nan, np.nan, ],
['MX17004', 2010, 3, 'tmax', np.nan, np.nan, np.nan, np.nan, 32.1, np.nan, np.nan, np.nan, ],
['MX17004', 2010, 3, 'tmin', np.nan, np.nan, np.nan, np.nan, 14.2, np.nan, np.nan, np.nan, ],
['MX17004', 2010, 4, 'tmax', np.nan, np.nan, np.nan, np.nan, np.nan, np.nan, np.nan, np.nan, ],
['MX17004', 2010, 4, 'tmin', np.nan, np.nan, np.nan, np.nan, np.nan, np.nan, np.nan, np.nan, ],
['MX17004', 2010, 5, 'tmax', np.nan, np.nan, np.nan, np.nan, np.nan, np.nan, np.nan, np.nan, ],
['MX17004', 2010, 5, 'tmin', np.nan, np.nan, np.nan, np.nan, np.nan, np.nan, np.nan, np.nan,]
]
messy = pd.DataFrame(data=data, columns=columns); messy
# Most of the values are not relevant. However, filtering the NaN values is imposible here. We need to melt the dataframe first.
molten = pd.melt(messy,
id_vars=['id', 'year','month','element',],
var_name='day');
molten.dropna(inplace=True)
molten = molten.reset_index(drop=True)
molten
# This dataframe is not in tidy form yet. First, the column `element` contains variable names. Second, the columns `year, month, day` represent one variable: the date. Let's fix the latter problem first.
# +
def f(row):
return "%d-%02d-%02d" % (row['year'], row['month'], int(row['day'][1:]))
molten['date'] = molten.apply(f,axis=1)
molten = molten[['id', 'element','value','date']]
molten
# -
# Now we need to pivot the element column.
tidy = molten.pivot(index='date',columns='element',values='value')
tidy
# Wait a minute.
#
# Where is the id?
#
# One way to keep it, is to move the id to an index with the `groupby()` function, and apply `pivot()` inside each group.
tidy = molten.groupby('id').apply(pd.DataFrame.pivot,
index='date',
columns='element',
values='value')
tidy
# We are almost there. We simply have to move id back as a column with the `reset_index()`.
tidy.reset_index(inplace=True)
tidy
# We get rid of the `element` name.
tidy.columns.name = ''
tidy
# Et Voilà!
# ## Multiple types in one table
#
# This example is used to illustrate two of the above problems.
#
# Let's create it. It is an excerpt from the Billboard top hits for 2000.
# +
columns = ['year','artist','track','time','date entered','wk1','wk2','wk3',]
data = [[2000,"2,Pac","Baby Don't Cry","4:22","2000-02-26",87,82,72,],
[2000,"2Ge+her","The Hardest Part Of ...","3:15","2000-09-02",91,87,92,],
[2000,"3 Doors Down","Kryptonite","3:53","2000-04-08",81,70,68,],
[2000,"98^0","Give Me Just One Nig...","3:24","2000-08-19",51,39,34,],
[2000,"A*Teens","Dancing Queen","3:44","2000-07-08",97,97,96,],
[2000,"Aaliyah","I Don't Wanna","4:15","2000-01-29",84,62,51,],
[2000,"Aaliyah","Try Again","4:03","2000-03-18",59,53,38,],
[2000,"Adams,Yolanda","Open My Heart","5:30","2000-08-26",76,76,74]
]
messy = pd.DataFrame(data=data, columns=columns)
messy
# -
# This dataset is messy because there are several observations per row, in the columns wk1, wk2, wk3. We can get one observation per row by metling the dataset.
molten = pd.melt(messy,
id_vars=['year','artist','track','time','date entered'],
var_name = 'week',
value_name = 'rank',
)
molten.sort_values(by=['date entered','week'], inplace=True)
molten.head()
# We can clean the dataset further, first by turning week into number
molten['week'] = molten['week'].apply(lambda s: int(s[2:]))
molten.head()
# Second, we need the starting date of the week for each observation, instead of the date the track entered.
# +
from datetime import datetime, timedelta
def increment_date(row):
date = datetime.strptime(row['date entered'], "%Y-%m-%d")
return date + timedelta(7) * (row['week'] - 1)
molten['date'] = molten.apply(increment_date, axis=1)
molten.drop('date entered', axis=1, inplace=True)
molten.head()
# -
# Last, this dataset is denormalized. This is fine for most statistical and machine learning packages, but we might want to normalize it. It means that we should group information that is repeated every week for a track in a separate table. This information appears in columns `year ,artist, track, time`.
tidy_track = molten[['year','artist','track','time']]\
.groupby(['year','artist','track'])\
.first()
tidy_track.reset_index(inplace=True)
tidy_track.reset_index(inplace=True)
tidy_track.rename(columns = {'index':'id'}, inplace=True)
tidy_track
tidy_rank = pd.merge(molten, tidy_track, on='track')
tidy_rank = tidy_rank[['id', 'date', 'rank']]
tidy_rank.head()
|
Tidy-Data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Tuple
# Tupuls are similar to list but the value cannot be changed after creating the tupul. All te operattions in tupuls are similar to the list operations
# #### Indexing
# 
# ### Operations in Tuple
#define a tuple
numbers = (1,2,3,4,5,6,7,8,9,10,11,12,13,14,15)
#Get length of the tuple
print(len(numbers))
#Get the index of a element
print(numbers.index(12))
#Count how many times a element is repeted
print(numbers.count(12))
#prints complete list
print(numbers)
#prints first element of the list
print(numbers[0])
#prints elements starting from 3rd to 5th
print(numbers[2:5])
#prints elements starting from 3rd to last
print(numbers[2:])
# +
#prints the elements of string from 3 to 7 skipping one element.
#This is extended slice syntax. The general form is [start:stop:step].
print(numbers[3:7:1])
# -
#reverse a tuple
print(numbers[::-1])
#Repete tuple
print(numbers * 2)
|
Day1/g__tuple_example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 04 - Preprocessing & Model Training 🤖
#
# ## Problem
#
# How can we generate short descriptions including a list of popular beer styles from brewery reviews with natural language processing (NLP)?
#
# NOTE: This is similar to how Google Maps provides short blurbs for businesses such as “From scratch, Northern Italian dining.”
#
# For example, "Spacious warehouse brewery with daily food trucks. Allows dogs. Features IPAs, Hazy IPAs, and high-gravity stouts."
#
# ## Notebook Objectives
#
# * Understand the importance of creating a model training development data set.
# * Correctly identify when to create dummy features or one-hot encoded features.
# * Understand the importance of magnitude standardization.
# * Apply the train and test split to the development dataset effectively.
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import warnings
import json
import time
import math
from pprint import pprint
import random
random.seed(42)
# Progress bar
# from tqdm import tqdm, trange
from tqdm.notebook import tqdm, trange
# NLP Toolkit
import spacy
from spacy import displacy
from spacy.util import minibatch, compounding, decaying
# %matplotlib inline
# -
# Set default plot size
plt.rcParams['figure.figsize'] = (10, 5)
BASE_MODEL_PATH = "../models/base_model"
# Load small English language model for sentence separation.
nlp = spacy.load("en_core_web_sm")
# Load data
df = pd.read_csv('../data/raw/brewery_reviews.csv', index_col=0, parse_dates=['date'])
df.head()
# We don't need reviews without text for this work
df.dropna(inplace=True)
# Count sentences
df['sents'].sum()
# Separate reviews into sentences for annotation.
# WARNING: THIS TAKES A LONG TIME (~8 minutes)
sentence_list = []
for review in tqdm(df['review_text']):
doc = nlp(review)
for sent in doc.sents:
sentence_list.append(sent.text)
# Get sentence length (i.e. number of tokens per sentence)
sentence_length = [len(sent) for sent in sentence_list]
# Visualize
sns.histplot(data=sentence_length)
plt.ylabel('Count')
plt.xlabel('Sentence Length (characters)')
plt.show()
# Create DataFrame of sentence
sent_df = pd.DataFrame(sentence_length)
# Inspect
sent_df.describe()
def find_outliers(array):
"""
Returns a tuple of (lower, upper) cut-offs so you can do something like:
`sent_list_cleaned = [sent for sent in sent_list if len(sent) >= lower and len(sent) <= upper]`
"""
data = np.array(array)
# calculate interquartile range
q25, q75 = np.percentile(data, 25), np.percentile(data, 75)
iqr = q75 - q25
print(f'Percentiles: 25th={q25}, 75th={q75}, IQR={iqr}')
# calculate the outlier cutoff
cut_off = iqr * 1.5
lower, upper = q25, q75 + cut_off
print(f'Cut-offs: lower={lower}, upper={upper}')
# identify outliers
outliers = [x for x in data if x < lower or x > upper]
print(f'Identified outliers: {len(outliers)}')
# remove outliers
outliers_removed = [x for x in data if x >= lower and x <= upper]
print(f'Non-outlier observations: {len(outliers_removed)}')
return lower, upper
lower, upper = find_outliers(sentence_length)
# Create list of sentences of a non-outlier length
sents_cleaned = [sent for sent in sentence_list if len(sent) >= lower and len(sent) <= upper]
# +
# Write random sentences to text file for import into Doccano
sents_np = np.array(sents_cleaned)
sizes = [100, 500, 1000, 5000]
for size in sizes:
random_sents = np.random.choice(sents_np, size=size, replace=False)
with open(f'../data/processed/brewery-review-sentences-{size}.txt','w') as write:
write.write("\n".join(random_sents))
# -
# ## Full Review Text vs. Sentences
#
# After loading 500 sentences into Doccano and starting to annotate, I realized the context may be more helpful from a user experience perspective and useful for a model training persepctive.
sns.histplot(data=df, x='tokens')
plt.xlabel('# Tokens (Words) in Review')
plt.ylabel('Count')
plt.show()
lower, upper = find_outliers(df['tokens'])
df_cleaned = df[(df['tokens'] >= lower) & (df['tokens'] <= upper)]
len(df_cleaned)
# +
# Write random sentences to text file for import into Doccano
reviews_np = np.array(df_cleaned['review_text'])
sizes = [100, 500, 1000, 5000]
for size in sizes:
random_reviews = np.random.choice(reviews_np, size=size, replace=False)
with open(f'../data/processed/brewery-reviews-{size}.txt','w') as write:
write.write("\n".join(random_reviews))
# -
# ## Model Training v1
# +
# Load the TRAIN_DATA from the Doccano export
filename = ("../data/annotations/final-training-set/reviews-annotated-ner-200-p2.json")
TRAIN_DATA = []
with open(filename) as file:
lines = file.readlines()
for train_data in lines:
data = json.loads(train_data)
if len(data['labels']) == 0 and data['annotation_approver'] == None: continue
ents = [tuple(entity[:3]) for entity in data['labels']]
TRAIN_DATA.append((data['text'],{'entities':ents}))
with open('{}'.format(filename.replace('json','txt')),'w') as write:
write.write(str(TRAIN_DATA))
print("[INFO] Stored the spacy training data and filename is {}".format(filename.replace('json','txt')))
# +
# Load the LABELS from the Doccano export
labels_filename = ("../data/annotations/final-training-set/labels.json")
LABELS = []
with open(labels_filename) as file:
labels = json.load(file)
LABELS = [label['text'] for label in labels]
print(LABELS)
# +
# Transfer Learning
# With an existing model, add the labels and save as the `base_model`
model = "en_core_web_md"
nlp = spacy.load(model)
print(f"Loading model '{model}'")
# Get the NER component so we can add labels
ner = nlp.get_pipe("ner")
# Add labels
for label in LABELS:
print(f"Adding LABEL '{label}'")
ner.add_label(label)
# Save the model to disk
nlp.to_disk(BASE_MODEL_PATH)
print(f"Saved base model to {BASE_MODEL_PATH}")
# +
# Train the model
# Load base_model
nlp = spacy.load(BASE_MODEL_PATH)
# Number of iterations
n_iter=100
# Drop to prevent over-fitting
drop = decaying(0.6, 0.2, 1e-4)
# Resume training since we're using a base model
optimizer = nlp.resume_training()
# Minibatch sizes
sizes = compounding(1.0, 4.0, 1.001)
loss = []
pipe_exceptions = ["ner", "trf_wordpiecer", "trf_tok2vec"]
other_pipes = [pipe for pipe in nlp.pipe_names if pipe not in pipe_exceptions]
# only train NER
with nlp.disable_pipes(*other_pipes), warnings.catch_warnings():
# show warnings for misaligned entity spans once
warnings.filterwarnings("once", category=UserWarning, module='spacy')
# batch up the examples using spaCy's minibatch
# NOTE: These are defaults from https://spacy.io/usage/training#ner
with trange(n_iter) as pbar:
for _ in pbar:
# Randomize the training data
random.shuffle(TRAIN_DATA)
# Create batches
batches = minibatch(TRAIN_DATA, size=sizes)
# Train and update
losses = {}
for batch in batches:
texts, annotations = zip(*batch)
nlp.update(texts, annotations, sgd=optimizer, drop=next(drop), losses=losses)
# Logging
loss.append(losses['ner'])
pbar.set_postfix(loss=losses['ner'])
# -
plt.plot(range(n_iter), loss)
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.show()
# Save Model
nlp.to_disk("../models/v5/model-final/")
# ## Model Training v2
#
# 1. Load in annotations from Doccano
# 2. Randomize and split into train and test sets
# 3. Use spacy CLI to train and save model
#
# ### Reference
#
# * https://medium.com/@justindavies/training-spacy-ner-models-with-doccano-8d8203e29bfa
# +
# Load the annotations from the Doccano export
filename = ("../data/annotations/final-training-set/reviews-annotated-ner-200-p2.json")
# Array of tuples [(TEXT, {'entities': ENTITIES[]})]
DATA = []
with open(filename) as file:
lines = file.readlines()
for annot_data in lines:
data = json.loads(annot_data)
# Ignore annotations without labels and no approval
if len(data['labels']) == 0 and data['annotation_approver'] == None: continue
# Create entities and append
ents = [tuple(entity[:3]) for entity in data['labels']]
DATA.append((data['text'], {'entities': ents}))
print(f"Number of examples: {len(DATA)}")
# -
# Convert Doccano JSONL to spaCy-compatible JSONL
with open(filename) as file:
lines = file.readlines()
for line in lines:
line = json.loads(line)
# Ignore annotations without labels and no approval
if len(line['labels']) == 0 and line['annotation_approver'] == None: continue
# Rename "labels" to "entities"
if "labels" in line:
line["entities"] = line.pop("labels")
else:
line["entities"] = []
# Reorganize the entities
tmp_ents = []
for entity in line["entities"]:
if entity[2] in LABELS:
tmp_ents.append({"start": entity[0], "end": entity[1], "label": entity[2]})
line["entities"] = tmp_ents
with open(filename.replace('.json','-spacy.jsonl'), 'a+') as write:
write.write(json.dumps({"entities": line["entities"], "text": line["text"]}))
write.write("\n")
# +
# Randomize and split into train and test
spacy_filename = filename.replace('.json','-spacy.jsonl')
train_set_ratio = 0.9
with open(spacy_filename) as file:
DATA = file.readlines()
random.shuffle(DATA)
n = math.floor(train_set_ratio * len(DATA))
train_data = DATA[:n]
test_data = DATA[n:]
print(f"train: {len(train_data)}")
print(f"test: {len(test_data)}")
# +
train_filename = spacy_filename.replace('.jsonl',f'-train-{len(train_data)}.jsonl')
test_filename = spacy_filename.replace('.jsonl',f'-dev-{len(test_data)}.jsonl')
with open(train_filename, 'w') as write:
write.write("".join(train_data))
print("[INFO] Stored the spacy training data at {}".format(train_filename))
with open(test_filename, 'w') as write:
write.write("".join(test_data))
print("[INFO] Stored the spacy dev data at {}".format(test_filename))
# +
# Convert JSONL to spaCy JSON
# python -m spacy convert ./data/annotations/final-training-set/reviews-annotated-ner-200-p2-spacy-train.jsonl ./data/annotations/final-training-set/ --lang en
# python -m spacy convert ./data/annotations/final-training-set/reviews-annotated-ner-200-p2-spacy-test.jsonl ./data/annotations/final-training-set/ --lang en
# Test / Debug Data
# python -m spacy debug-data en ./data/annotations/final-training-set/reviews-annotated-ner-200-p2-spacy-train.json ./data/annotations/final-training-set/reviews-annotated-ner-200-p2-spacy-test.json --base-model ./models/base_model --pipeline ner
# Train model
# python -m spacy train en ./models/v4/ ./data/annotations/final-training-set/reviews-annotated-ner-200-p2-spacy-train.json ./data/annotations/final-training-set/reviews-annotated-ner-200-p2-spacy-test.json -b ./models/base_model -p ner
# Evaluate model
# python -m spacy evaluate ./models/v4/model-final ./data/annotations/final-training-set/reviews-annotated-ner-200-p2-spacy-test.json --displacy-path reports/evaluations/v4
# -
# ## Evaluate Model (manually)
# Load saved model
nlp = nlp.from_disk("../models/v5/model-final")
# Set up colors and options for displaCy
colors = {"FEATURE": "#80CBC4", "BEER_STYLE": "#FDD835", "LOCATION": "#C5CAE9", "BREWERY": "#AED581"}
options = {"colors": colors}
# +
# Test trained model with real data
with open('../data/raw/reviews_san-diego_half-door-brewing-co.txt') as file:
reviews = file.readlines()
for review in reviews:
doc = nlp(review)
displacy.render(doc, style="ent", options=options)
# +
# Test trained model
with open('../data/raw/reviews_san-diego_alesmith.txt') as file:
reviews = file.readlines()
for review in reviews:
doc = nlp(review)
displacy.render(doc, style="ent", options=options)
# -
|
notebooks/03 - Model Training.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from bokeh.plotting import figure
from bokeh.embed import notebook_div
from bokeh.io import show, push_notebook, output_notebook
output_notebook()
plot = figure(plot_width=500, plot_height=500)
plot.circle([1,2], [3,4])
show(plot)
# cd ..
import blackscholes as bs
import numpy as np
from ipywidgets import interact
k = 1
s_space = np.linspace(0.0001,2,100)
delta_surface = np.vstack((bs.blackScholes(t, s_space, k, 0.2)['delta'] for t in np.linspace(1,0.0001,100)))
# +
# N = 500
# x = np.linspace(0, 10, N)
# y = np.linspace(0, 10, N)
# xx, yy = np.meshgrid(x, y)
# d = np.sin(xx)*np.cos(yy)
# p = figure(x_range=(0, 10), y_range=(0, 10), plot_width=500, plot_height=500)
p = figure(plot_width=500, plot_height=500)
# must give a vector of image data for image parameter
img = p.image(image=[delta_surface], x=0, y=0, dw=10, dh=10, palette="Viridis256")
show(p, notebook_handle=True)
# -
img.data_source
def update(vola=0.1, k=1):
# k = 1
s_space = np.linspace(0.0001, 2, 100)
delta_surface = np.vstack((bs.blackScholes(t, s_space, k, vola)['delta'] for t in np.linspace(1, 0.0001, 100)))
img.data_source.data['image'] = [delta_surface]
push_notebook()
interact(update, vola=(0.0001, 1., 0.01), k=(0.01, 4.0, 0.01));
# ### Policy
delta_surface.tostring()
k = 1
s_space = np.linspace(0.0001,2,100)
np.array([2, 3, 1]).dot(np.array([t, s, 1]))
delta_surface = np.vstack((np.arrat, s_space, k, 0.2)['delta'] for t in np.linspace(1,0.0001,100)))
# ## lskl
# +
from pythreejs import *
import numpy as np
from IPython.display import display
vertices = np.asarray([
[0, 0, 0],
[0, 0, 1],
[0, 1, 0],
[0, 1, 1],
[1, 0, 0],
[1, 0, 1],
[1, 1, 0],
[1, 1, 1]
], dtype='float32')
faces = np.asarray([
[0, 1, 3],
[0, 2, 3],
[0, 2, 4],
[2, 4, 6],
[0, 1, 4],
[1, 4, 5],
[2, 3, 6],
[3, 6, 7],
[1, 3, 5],
[3, 5, 7],
[4, 5, 6],
[5, 6, 7]
])
vertexcolors = np.asarray([(0,0,0), (0,0,1), (0,1,0), (1,0,0), (0,1,1), (1,0,1), (1,1,0), (1,1,1)])
cubeGeometry = PlainBufferGeometry(vertices=vertices, faces=faces, colors = vertexcolors)
myobjectCube = Mesh(geometry=cubeGeometry, material = LambertMaterial(vertexColors = 'VertexColors'))
cCube = PerspectiveCamera(position=[3, 3, 3], fov=20,
children=[DirectionalLight(color='#ffffff', position=[-3, 5, 1], intensity=0.5)])
sceneCube = Scene(children=[myobjectCube, AmbientLight(color='#dddddd')])
rendererCube = Renderer(camera=cCube, background='black', background_opacity=1,
scene = sceneCube, controls=[OrbitControls(controlling=cCube)])
display(rendererCube)
# -
import pythreejs
from traitlets import link, dlink
from ipywidgets import HTML, Text
# +
nx, ny = (20, 20)
xmax=1
x = np.linspace(-xmax, xmax, nx)
y = np.linspace(-xmax, xmax, ny)
xx, yy = np.meshgrid(x, y)
z = xx ** 2 - yy ** 2
#z[6,1] = float('nan')
surf_g = SurfaceGeometry(z=list(z[::-1].flat),
width=2 * xmax,
height=2 * xmax,
width_segments=nx - 1,
height_segments=ny - 1)
surf = Mesh(geometry=surf_g, material=LambertMaterial(map=height_texture(z[::-1], 'YlGnBu_r')))
surfgrid = SurfaceGrid(geometry=surf_g, material=LineBasicMaterial(color='black'))
hover_point = Mesh(geometry=SphereGeometry(radius=0.05), material=LambertMaterial(color='hotpink'))
scene = Scene(children=[surf, surfgrid, hover_point, AmbientLight(color='#777777')])
c = PerspectiveCamera(position=[0, 3, 3], up=[0, 0, 1],
children=[DirectionalLight(color='white', position=[3, 5, 1], intensity=0.6)])
click_picker = Picker(root=surf, event='dblclick')
hover_picker = Picker(root=surf, event='mousemove')
renderer = Renderer(camera=c, scene = scene, controls=[OrbitControls(controlling=c), click_picker, hover_picker])
def f(change):
value = change['new']
print('Clicked on %s' % value)
point = Mesh(geometry=SphereGeometry(radius=0.05),
material=LambertMaterial(color='red'),
position=value)
scene.children = list(scene.children) + [point]
click_picker.observe(f, names=['point'])
link((hover_point, 'position'), (hover_picker, 'point'))
h = HTML()
def g(change):
h.value = 'Pink point at (%.3f, %.3f, %.3f)' % tuple(change['new'])
g({'new': hover_point.position})
hover_picker.observe(g, names=['point'])
display(h)
display(renderer)
|
notebooks/Visualize_Policy.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Feature Extraction with Data Augmentation
# ### Imports
# +
import tensorflow as tf
from tensorflow import keras
from keras import optimizers
from keras.models import Sequential
from keras.layers import Conv2D, MaxPooling2D, Dense, Flatten, Dropout
from keras.utils import np_utils
from keras.callbacks import Callback
from keras.preprocessing.image import ImageDataGenerator
from keras.applications import VGG16
from tensorflow.python.keras.callbacks import TensorBoard
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
from matplotlib import rcParams
from time import time
import os
import wandb
from wandb.keras import WandbCallback
import numpy as np
# -
# ### Logging code
# +
# TENSORBOARD_LOGS_DIR = f"dandc-{int(time())}"
# tensorboard = TensorBoard(log_dir=f"logs/{TENSORBOARD_LOGS_DIR}", write_images=True, histogram_freq=1, write_grads=True)
# run = wandb.init()
# config = run.config
# -
# ### Network Configuration
config = {
'img_width': 150,
'img_height': 150,
'first_layer_conv_width': 3,
'first_layer_conv_height': 3,
'dense_layer_size': 256,
'epochs': 30,
'optimizer': "adam",
'hidden_nodes': 100
}
# ### Images Dataset
base_dir = 'E:\kaggle\dogs-and-cats\sm_dataset'
train_dir = os.path.join(base_dir, 'train')
validation_dir = os.path.join(base_dir, 'validation')
test_dir = os.path.join(base_dir, 'test')
# ### Data Generation
# +
train_datagen = ImageDataGenerator(
rescale=1./255,
rotation_range=40,
width_shift_range=0.2,
height_shift_range=0.2,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True,
fill_mode='nearest'
)
test_datagen = ImageDataGenerator(rescale=1./255)
train_generator = train_datagen.flow_from_directory(train_dir, target_size=(150, 150), batch_size=20, class_mode='binary')
validation_generator = test_datagen.flow_from_directory(validation_dir, target_size=(150, 150), batch_size=20, class_mode='binary')
# -
# ### Inspect Generators
for data_batch, labels_batch in train_generator:
print('data batch shape:', data_batch.shape)
print('labels batch shape:', labels_batch.shape)
break
# +
# %matplotlib inline
rcParams['figure.figsize'] = 11, 8
fig, ax = plt.subplots(1,4)
for data_batch, labels_batch in train_generator:
ax[0].imshow(data_batch[0])
ax[1].imshow(data_batch[1])
ax[2].imshow(data_batch[2])
ax[3].imshow(data_batch[3])
break
# -
# ### Base Convolutional Mode
conv_base = VGG16(
weights = 'imagenet',
include_top = False,
input_shape = (150, 150, 3)
)
conv_base.summary()
# ### Create Model
model = Sequential()
model.add(conv_base)
model.add(Flatten())
model.add(Dropout(0.5))
model.add(Dense(config['dense_layer_size'], activation='relu'))
model.add(Dropout(0.4))
model.add(Dense(1, activation="sigmoid"))
model.summary()
# ### Freeze Convolutional Layer
print('This is the number of trainable weights before freezing the conv base:',
len(model.trainable_weights))
conv_base.trainable = True
for layer in conv_base.layers:
if layer.name == 'block5_conv1':
layer.trainable = True
else:
layer.trainable = False
print('This is the number of trainable weights after freezing the conv base:', len(model.trainable_weights))
# ### Compile Model
model.compile(
loss='binary_crossentropy',
optimizer=optimizers.RMSprop(lr=1e-5),
metrics=['accuracy'])
# ### Training
class TensorBoardWrapper(TensorBoard):
'''Sets the self.validation_data property for use with TensorBoard callback.'''
def __init__(self, batch_gen, nb_steps, b_size, **kwargs):
super(TensorBoardWrapper, self).__init__(**kwargs)
self.batch_gen = batch_gen # The generator.
self.nb_steps = nb_steps # Number of times to call next() on the generator.
#self.batch_size = b_size
def on_epoch_end(self, epoch, logs):
# Fill in the `validation_data` property. Obviously this is specific to how your generator works.
# Below is an example that yields images and classification tags.
# After it's filled in, the regular on_epoch_end method has access to the validation_data.
imgs, tags = None, None
for s in range(self.nb_steps):
ib, tb = next(self.batch_gen)
if imgs is None and tags is None:
imgs = np.zeros(((self.nb_steps * self.batch_size,) + ib.shape[1:]), dtype=np.float32)
tags = np.zeros(((self.nb_steps * self.batch_size,) + tb.shape[1:]), dtype=np.uint8)
imgs[s * ib.shape[0]:(s + 1) * ib.shape[0]] = ib
tags[s * tb.shape[0]:(s + 1) * tb.shape[0]] = tb
self.validation_data = [imgs, tags, np.ones(imgs.shape[0])]
return super(TensorBoardWrapper, self).on_epoch_end(epoch, logs)
tbw = TensorBoardWrapper(validation_generator, nb_steps=50 // 20, b_size=20, log_dir='./log',
histogram_freq=1,
write_graph=False, write_grads=True)
history = model.fit_generator(train_generator, epochs=config['epochs'], steps_per_epoch=100, validation_data=validation_generator, validation_steps=50, callbacks=[])
# ### Save model
model.save('candd06.h5')
# ### Loss & Accuracy
# +
def smooth_curve(points, factor=0.8):
smoothed_points = []
for point in points:
if smoothed_points:
previous = smoothed_points[-1]
smoothed_points.append(previous * factor + point * (1 - factor))
else:
smoothed_points.append(point)
return smoothed_points
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(1, len(acc) + 1)
plt.plot(epochs, smooth_curve(acc), 'bo', label='Training Acc')
plt.plot(epochs, smooth_curve(val_acc), 'b', label='Validation Acc')
plt.title('Training and Validation Accuracy')
plt.legend()
plt.show()
# -
# ### Training and Validation loss
plt.plot(epochs, smooth_curve(loss), 'bo', label='Training Loss')
plt.plot(epochs, smooth_curve(val_loss), 'b', label='Validation Loss')
plt.title('Training and Validation Loss')
plt.legend()
plt.show()
|
cnn/cats-and-dogs/cats-and-dogs-ffe-fine-tunning.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="aWR6ocRVB6bu" colab_type="text"
# ## Unsupervised Learning
# #### <NAME>
# #### 12-03-2019
#
# The dataset used for this analysis can be found in the sklearn Library in Python.
#
# ### About t-SNE
# t-distributed stochastic neighbor embedding (t-SNE), is one of the unsupervised learning method for visualisation. It maps high dimensional space into a 2 or 3 dimensional space which can be visualised. Specifically, it models each high-dimensional object by a two- or three-dimensional point in such a way that similar objects are modeled by nearby points and dissimilar objects are modeled by distant points with high probability.
#
# ### Purpose of this analysis
# For our Unsupervised Algorithm to model each high-dimensional object by a two- or three-dimensional point in such a way that similar objects are modeled by nearby points and dissimilar objects are modeled by distant points with high probability.
# + id="H_q3rBf1Bz_t" colab_type="code" outputId="157c67ed-d4aa-42af-8307-15c002c3a5cd" colab={"base_uri": "https://localhost:8080/", "height": 347}
# importing libraries
from sklearn import datasets
from sklearn.manifold import TSNE
import matplotlib.pyplot as plt
# load dataset
iris_df = datasets.load_iris()
# defining the model
model = TSNE(learning_rate=100)
# fitting the model
transformed = model.fit_transform(iris_df.data)
# plotting 2d t-sne
x_axis = transformed[:, 0]
y_axis = transformed[:, 1]
plt.scatter(x_axis, y_axis, c=iris_df.target)
plt.show()
# + [markdown] id="p0uy9H4vhnl9" colab_type="text"
# Here as the Iris dataset has four features(4d) it is transformed and represented in two dimensional figure. Similarly t-SNE model can be applied to a dataset which has n-features.
|
Data Science Lit/t_SNE_clustering.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import glob
import pandas as pd
import os
import pyspark
import warnings
warnings.filterwarnings(action='ignore')
# from pyspark import SparkContext
# sc = SparkContext()
# from pyspark.sql import SQLContext
# sqlContext = SQLContext(sc)
from pyspark.sql import SparkSession
spark = SparkSession \
.builder \
.appName("Python Spark SQL basic example") \
.getOrCreate()
# -
os.getcwd()
# +
#Read Clicks Data
# files_path =
files_path = glob.glob( '**/*.csv',recursive=True)
files_path = files_path
print(files_path)
# df_clicks = pd.DataFrame()
# for file in files_path:
# print(file)
# df = pd.read_parquet(file)
# df_clicks = df_clicks.append(df)
# print(df_clicks.shape)
# +
from pyspark.sql.types import StructType, StructField, IntegerType,StringType,DoubleType
schema = StructType([
StructField("STATION", StringType(), True),
StructField("YEARMONTH", StringType(), True),
StructField("KPI", StringType(), True),
StructField("VALUE", DoubleType(), True),
StructField("FLAG1", StringType(), True),
StructField("FLAG2", StringType(), True),
StructField("FLAG3", StringType(), True),
StructField("FLAG4", StringType(), True)])
schema_less = StructType([
StructField("STATION", StringType(), True),
StructField("YEARMONTH", StringType(), True),
StructField("KPI", StringType(), True),
StructField("VALUE", DoubleType(), True),
StructField("COUNTRY_CODE", DoubleType(), True)])
# -
main_df = spark.createDataFrame([],schema = schema_less)
main_df.show()
col_names = ['STATION','YEARMONTH','KPI','VALUE','COUNTRY_CODE']
files_path
files_path =files_path[0:7]
files_path
for i in files_path:
print(i)
df = spark.read.csv(i,inferSchema=False,schema = schema)
df = df.withColumn('COUNTRY_CODE', df.STATION.substr(0,2))
df = df.filter(df.COUNTRY_CODE == 'US').select(col_names).filter(df.KPI.isin(['PRCP','SNOW','SNWD','TMAX','TAVG','TMIN']))
main_df = main_df.unionByName(df)
pivotDF = main_df.groupBy("STATION","YEARMONTH").pivot("KPI").sum("VALUE")
# pivotDF.printSchema()
stn_df = spark.read.parquet('output/stn.parquet')
# print(pivotDF.count())
join_df = pivotDF.join(stn_df,how='left',on='STATION')
# print(join_df.count())
join_df = join_df.withColumn("YEAR", join_df.YEARMONTH.substr(0,4)).withColumn("MONTH", join_df.YEARMONTH.substr(5,2)).withColumn("DAY", join_df.YEARMONTH.substr(7,2))
join_df.write.parquet('output/full_data.parquet')
# +
# join_df = join_df.fillna(subset=['SNOW','PRCP'],value=0.0)
# +
# join_df.show()
# -
import pyspark.sql.functions as F
agg_df = join_df.groupBy("YEAR","MONTH","STATE_CITY_NAME").agg(F.max("SNOW").alias("MAX_SNOW"), \
F.min("SNOW").alias("MIN_SNOW"), \
F.max("PRCP").alias("MAX_PRCP"), \
F.min("PRCP").alias("MIN_PRCP"), \
F.mean("SNOW").alias("MEAN_SNOW"), \
F.mean("PRCP").alias("MEAN_PRCP"), \
F.percentile_approx("SNOW", 0.5).alias("MEDIAN_SNOW"), \
F.percentile_approx("PRCP", 0.5).alias("MEDIAN_PRCP"), \
F.mean("LONG").alias("MEAN_LONG"), \
F.mean("LAT").alias("MEAN_LAT") )
agg_year_only_df = join_df.groupBy("YEAR","STATE_CITY_NAME").agg(F.max("SNOW").alias("MAX_SNOW"), \
F.min("SNOW").alias("MIN_SNOW"), \
F.max("PRCP").alias("MAX_PRCP"), \
F.min("PRCP").alias("MIN_PRCP"), \
F.mean("SNOW").alias("MEAN_SNOW"), \
F.mean("PRCP").alias("MEAN_PRCP"), \
F.percentile_approx("SNOW", 0.5).alias("MEDIAN_SNOW"), \
F.percentile_approx("PRCP", 0.5).alias("MEDIAN_PRCP"), \
F.mean("LONG").alias("MEAN_LONG"), \
F.mean("LAT").alias("MEAN_LAT") )
agg_year_only_df.write.parquet('output/agg_data_year_level.parquet')
agg_df.write.parquet('output/agg_df_y_m.parquet')
|
Weather Data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Engineering
# +
import numpy as np
import pandas as pd
calendar = pd.read_csv("../data/calendar.csv")
sales = pd.read_csv("../data/sales_train_validation.csv")
# sample_submission = pd.read_csv("../data/sample_submission.csv")
sell_prices = pd.read_csv("../data/sell_prices.csv")
# -
calendar.set_index('d')
sell_prices.set_index(["store_id", "item_id"])
# +
items = sales["item_id"].unique()
a = []
for s in items:
a.append(s)
if s=="FOODS_3_827":
break
print(len(a), set(items)-set(a))
# +
# sample = sales.sample(frac=0.1, replace=False, random_state=1)
# len(sample)
# +
# # This is disaster, Too slow
# from tqdm.notebook import tqdm
# with tqdm(total=len(sample)) as pbar:
# dfarray = []
# for index, row in sample.iterrows():
# s = sell_prices
# s = s[s["store_id"]==row["store_id"]]
# s = s[s["item_id"]==row["item_id"]]
# weekly_idx = 0
# acc_weekly_sale = 0
# prev_weekly_sale = 0
# cc = 0
# for i in range(1, 1914):
# d = calendar[calendar["d"]=="d_{}".format(i)]
# wm_yr_wk = int(d["wm_yr_wk"])
# ss = s[s["wm_yr_wk"]==wm_yr_wk]
# sell_price = None
# if len(ss)!=0:
# sell_price = ss["sell_price"]
# rolling_mean = None
# if i > 3:
# rolling_mean = 0
# rolling_mean += row["d_{}".format(i)]
# rolling_mean += row["d_{}".format(i-1)]
# rolling_mean += row["d_{}".format(i-2)]
# rolling_mean /= 3
# if wm_yr_wk == weekly_idx:
# acc_weekly_sale += row["d_{}".format(i)]
# cc += 1
# else:
# weekly_idx = wm_yr_wk
# prev_weekly_sale = acc_weekly_sale
# acc_weekly_sale = row["d_{}".format(i)]
# cc = 1
# dfarray.append({
# "item_id": row["item_id"],
# "dept_id": row["dept_id"],
# "cat_id": row["cat_id"],
# "store_id": row["store_id"],
# "state_id": row["state_id"],
# "day": i,
# "wm_yr_wk": d["wm_yr_wk"],
# "wday": d["wday"],
# "month": d["month"],
# "year": d["year"],
# "event_name_1": d["event_name_1"],
# "event_type_1": d["event_type_1"],
# "event_name_2": d["event_name_2"],
# "event_type_2": d["event_type_2"],
# "snap_CA": d["snap_CA"],
# "snap_TX": d["snap_TX"],
# "snap_WI": d["snap_WI"],
# "sell_price": sell_price,
# "sale": row["d_{}".format(i)],
# "shifted_sale1": row["d_{}".format(i-1)] if i > 1 else None,
# "shifted_sale2": row["d_{}".format(i-2)] if i > 2 else None,
# "shifted_sale3": row["d_{}".format(i-3)] if i > 3 else None,
# "shifted_sale4": row["d_{}".format(i-4)] if i > 4 else None,
# "shifted_sale5": row["d_{}".format(i-5)] if i > 5 else None,
# "rolling_mean": rolling_mean,
# "prev_weekly_sale": prev_weekly_sale,
# "acc_weekly_sale": acc_weekly_sale,
# "avg_sale_this_week": acc_weekly_sale*1.0/cc,
# "next_sale": row["d_{}".format(i+1)] if i < 1913 else None,
# })
# if index%200 == 0:
# df = pd.DataFrame(dfarray)
# df.to_csv("expand_sale_{}.csv".format(index))
# pbar.update(1)
# # break
# +
# df = pd.DataFrame(dfarray)
# df.to_csv("expand_sale.csv")
# -
# ## Privot data v2
#
# I still cannot run this code on my machine (it ate all my RAM) so I decided to run on colab.
# https://colab.research.google.com/drive/1CzOnHyEAUIX6OLJtXV09TOXXosXaieZa#scrollTo=GN6oYkAXEzix
#
# +
# df = sales.melt(id_vars=["id", "item_id", "dept_id", "cat_id", "store_id", "state_id"], var_name="d", value_name="sale")
# itemIds = sales["item_id"].unique()
# from tqdm.notebook import tqdm
# with tqdm(total=len(itemIds)) as pbar:
# for item in itemIds:
# subDf = df[df["id"]==item]
# p = pd.merge(subDf, calendar, on='d')
# p2 = pd.merge(p, sell_prices, on=["store_id", "item_id", "wm_yr_wk"], how="left")
# p2.to_csv("dataframes/expand_df_{}.csv".format(item))
# print(item, "DONE")
# pbar.update(1)
# -
# # Loading new datasets
#
itemIds = sales["item_id"].unique()
itemIds = ["HOBBIES_1_{:03d}".format(i) for i in range(1, 51)]
filenames = ["expand_df_"+s for s in itemIds]
df = pd.read_csv("dataframes/{}.csv".format(filenames[0]))
df.sort_values(by=['store_id', "date"], inplace=True)
df.reset_index(inplace=True)
df.head()
dfv2 = df.drop(["Unnamed: 0", "id", "item_id", "dept_id", "cat_id"], axis=1)
dfv2.head()
# # Feature Engineering
# +
# "shifted_sale1": row["d_{}".format(i-1)] if i > 1 else None,
# "shifted_sale2": row["d_{}".format(i-2)] if i > 2 else None,
# "shifted_sale3": row["d_{}".format(i-3)] if i > 3 else None,
# "shifted_sale4": row["d_{}".format(i-4)] if i > 4 else None,
# "shifted_sale5": row["d_{}".format(i-5)] if i > 5 else None,
# "rolling_mean": rolling_mean,
# "prev_weekly_sale": prev_weekly_sale,
# "acc_weekly_sale": acc_weekly_sale,
# "avg_sale_this_week": acc_weekly_sale*1.0/cc,
# "next_sale": row["d_{}".format(i+1)] if i < 1913 else None,
import math
def get_features(df):
for i in range(1, 6):
df[f"shift_t{i}"] = df.groupby('store_id')["sale"].shift(i)
df["rolling_mean"] = df["sale"].shift(1).rolling(3, min_periods=1).mean()
df["rolling_decay_mean"] = df["shift_t1"].copy()
for i in range(2, 4):
df["rolling_decay_mean"] += math.pow(0.9, i-1) * df[f"shift_t{i}"]
df["rolling_decay_mean"] = df["rolling_decay_mean"]/3.0
weekly_sale = df.groupby(['store_id', "wm_yr_wk"])["sale"].sum().reset_index()
weekly_sale["prev_weekly_sale"] = weekly_sale.groupby('store_id')["sale"].shift(1)
weekly_sale.drop(["sale"], axis=1, inplace=True)
df = pd.merge(df, weekly_sale, on=["store_id", "wm_yr_wk"], how="left")
df["acc_sale_by_week"] = df.groupby(['store_id', "wm_yr_wk"])["sale"].cumsum()
#skew, kurt??
#is_weekend
return df
def get_label(df):
return df.groupby('store_id')["sale"].shift(-1)
dfv3 = get_features(dfv2)
next_sale = get_label(dfv3)
# -
pd.set_option('display.expand_frame_repr', False)
pd.set_option('display.max_columns', 500)
dfv3.columns
dfv3.head(1)
# +
X = dfv3.drop(["index", "d", "date", "wm_yr_wk", "weekday"], axis=1)
X["wday"] = X["wday"].astype('category')
X["month"] = X["month"].astype('category')
X["year"] = X["year"].astype('category')
# eventNames = X["event_name_1"].unique().tolist()+X["event_name_2"].unique().tolist()
# This prove that event1 unique values covers event2
a = set(X["event_name_1"].unique().tolist())
b = set(X["event_name_2"].unique().tolist())
print(b - a)
#TODO: deal with events
# X1 = pd.get_dummies(X, prefix=['event1'], columns = ['event_name_1'], dummy_na=True, drop_first=True)
# X2 = pd.get_dummies(X, prefix=['event2'], columns = ['event_name_2'], dummy_na=True, drop_first=True)
X["holiday"] = pd.notna(X["event_name_1"])
X = X.drop(["event_name_1", "event_name_2", "event_type_1", "event_type_2"], axis=1)
# Ignore where sell_price is nan which indicates that it did not sell in that day
X["pred"] = next_sale
X = X.dropna(subset=['sell_price', "pred"])
Y = X["pred"]
X = X.drop(["pred"], axis=1)
X.head()
# +
from sklearn import preprocessing, metrics
cat = ['store_id', 'state_id']
for feature in cat:
encoder = preprocessing.LabelEncoder()
X[feature] = encoder.fit_transform(X[feature])
# -
X["snap_CA"] = X["snap_CA"].astype('int')
X["snap_TX"] = X["snap_TX"].astype('int')
X["snap_WI"] = X["snap_WI"].astype('int')
X.head()
# # Train LBGM
# +
from sklearn.model_selection import train_test_split
import lightgbm as lgb
x_train, x_test, y_train, y_test = train_test_split(X, Y, test_size=0.2)
params = {
# 'boosting_type': 'gbdt',
'metric': 'rmse',
'objective': 'poisson',
'n_jobs': -1,
'seed': 20,
'learning_rate': 0.1,
'alpha': 0.1,
'lambda': 0.1,
'bagging_fraction': 0.66,
'bagging_freq': 2,
'colsample_bytree': 0.77}
train_set = lgb.Dataset(x_train, y_train)
test_set = lgb.Dataset(x_test, y_test)
model = lgb.train(params, train_set, num_boost_round = 2000, early_stopping_rounds = 200, valid_sets = [train_set, test_set], verbose_eval = 100)
# joblib.dump(model, 'lgbm_0.sav')
val_pred = model.predict(x_test, num_iteration=model.best_iteration)
val_score = np.sqrt(metrics.mean_squared_error(val_pred, y_test))
print(f'RMSE: {val_score}')
# # def predict(test, submission):
# # predictions = test[['id', 'date', 'demand']]
# # predictions = pd.pivot(predictions, index = 'id', columns = 'date', values = 'demand').reset_index()
# # predictions.columns = ['id'] + ['F' + str(i + 1) for i in range(28)]
# # evaluation_rows = [row for row in submission['id'] if 'evaluation' in row]
# # evaluation = submission[submission['id'].isin(evaluation_rows)]
# # validation = submission[['id']].merge(predictions, on = 'id')
# # final = pd.concat([validation, evaluation])
# # final.to_csv('submission.csv', index = False)
# -
|
tk/Simple models.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="O9I9rz0pTamX"
# # Basic RoBERTa Fine-tuning Example
# + [markdown] colab_type="text" id="EiowR0WNTd1C"
# In this notebook, we will:
#
# * Train a RoBERTa base model on MRPC and evaluate its performance
# + [markdown] colab_type="text" id="rXbD_U1_VDnw"
# ## Setup
# + [markdown] colab_type="text" id="tC9teoazUnW8"
# #### Install dependencies
#
# First, we will install libraries we need for this code.
# + colab={} colab_type="code" id="8aU3Z9szuMU9"
# %%capture
# !git clone https://github.com/jiant-dev/jiant.git
# !cd jiant
# !git checkout d26c213c742d36f8909f3a910694c8a90da416f1
# !cd ..
# + colab={} colab_type="code" id="hMUKEH2YvFPv"
# %%capture
# This Colab notebook already has its CUDA-runtime compatible versions of torch and torchvision installed
# !pip install -r jiant/requirements-no-torch.txt
# Install pyarrow for nlp
# !pip install pyarrow==0.16.0
# + [markdown] colab_type="text" id="KGJcCmRzU1Qb"
# #### Download data
#
# Next, we will download task data.
# + colab={} colab_type="code" id="jKCz8VksvFlN"
# %%capture
# Download MRPC data
# !PYTHONPATH=/content/jiant python jiant/jiant/scripts/download_data/runscript.py \
# download \
# --tasks mrpc \
# --output_path=/content/tasks/
# + [markdown] colab_type="text" id="rQKSAhYzVIlv"
# ## `jiant` Pipeline
# + colab={} colab_type="code" id="v88oXqmBvFuK"
import sys
sys.path.insert(0, "/content/jiant")
# + colab={} colab_type="code" id="ibmMT7CXv1_P"
import jiant.proj.main.tokenize_and_cache as tokenize_and_cache
import jiant.proj.main.export_model as export_model
import jiant.proj.main.scripts.configurator as configurator
import jiant.proj.main.runscript as main_runscript
import jiant.shared.caching as caching
import jiant.utils.python.io as py_io
import jiant.utils.display as display
import os
# + [markdown] colab_type="text" id="HPZHyLOlVp07"
# #### Download model
#
# Next, we will download a `roberta-base` model. This also includes the tokenizer.
# + colab={} colab_type="code" id="K06qUGjkKWa7"
export_model.lookup_and_export_model(
model_type="roberta-base",
output_base_path="./models/roberta-base",
)
# + [markdown] colab_type="text" id="dV-T-8r1V0wf"
# #### Tokenize and cache
#
# With the model and data ready, we can now tokenize and cache the inputs features for our task. This converts the input examples to tokenized features ready to be consumed by the model, and saved them to disk in chunks.
# + colab={} colab_type="code" id="22bNWQajO4zm"
# Tokenize and cache each task
task_name = "mrpc"
tokenize_and_cache.main(tokenize_and_cache.RunConfiguration(
task_config_path=f"./tasks/configs/{task_name}_config.json",
model_type="roberta-base",
model_tokenizer_path="./models/roberta-base/tokenizer",
output_dir=f"./cache/{task_name}",
phases=["train", "val"],
))
# + [markdown] colab_type="text" id="JJ-mWSQQWJsw"
# We can inspect the first examples of the first chunk of each task.
# + colab={} colab_type="code" id="iLk_X0KypUyr"
row = caching.ChunkedFilesDataCache("./cache/mrpc/train").load_chunk(0)[0]["data_row"]
print(row.input_ids)
print(row.tokens)
# + [markdown] colab_type="text" id="3MBuH19IWOr0"
# #### Writing a run config
#
# Here we are going to write what we call a `jiant_task_container_config`. This configuration file basically defines a lot of the subtleties of our training pipeline, such as what tasks we will train on, do evaluation on, batch size for each task. The new version of `jiant` leans heavily toward explicitly specifying everything, for the purpose of inspectability and leaving minimal surprises for the user, even as the cost of being more verbose.
#
# We use a helper "Configurator" to write out a `jiant_task_container_config`, since most of our setup is pretty standard.
#
# **Depending on what GPU your Colab session is assigned to, you may need to lower the train batch size.**
# + colab={} colab_type="code" id="pQYtl7xTKsiP"
jiant_run_config = configurator.SimpleAPIMultiTaskConfigurator(
task_config_base_path="./tasks/configs",
task_cache_base_path="./cache",
train_task_name_list=["mrpc"],
val_task_name_list=["mrpc"],
train_batch_size=8,
eval_batch_size=16,
epochs=3,
num_gpus=1,
).create_config()
os.makedirs("./run_configs/", exist_ok=True)
py_io.write_json(jiant_run_config, "./run_configs/mrpc_run_config.json")
display.show_json(jiant_run_config)
# + [markdown] colab_type="text" id="-UF501yoXHBi"
# To briefly go over the major components of the `jiant_task_container_config`:
#
# * `task_config_path_dict`: The paths to the task config files we wrote above.
# * `task_cache_config_dict`: The paths to the task features caches we generated above.
# * `sampler_config`: Determines how to sample from different tasks during training.
# * `global_train_config`: The number of total steps and warmup steps during training.
# * `task_specific_configs_dict`: Task-specific arguments for each task, such as training batch size and gradient accumulation steps.
# * `taskmodels_config`: Task-model specific arguments for each task-model, including what tasks use which model.
# * `metric_aggregator_config`: Determines how to weight/aggregate the metrics across multiple tasks.
# + [markdown] colab_type="text" id="BBKkvXzdYPqZ"
# #### Start training
#
# Finally, we can start our training run.
#
# Before starting training, the script also prints out the list of parameters in our model.
# + colab={} colab_type="code" id="JdwWPgjQWx6I"
run_args = main_runscript.RunConfiguration(
jiant_task_container_config_path="./run_configs/mrpc_run_config.json",
output_dir="./runs/mrpc",
model_type="roberta-base",
model_path="./models/roberta-base/model/roberta-base.p",
model_config_path="./models/roberta-base/model/roberta-base.json",
model_tokenizer_path="./models/roberta-base/tokenizer",
learning_rate=1e-5,
eval_every_steps=500,
do_train=True,
do_val=True,
do_save=True,
force_overwrite=True,
)
main_runscript.run_loop(run_args)
# + [markdown] colab_type="text" id="4SXcuHFIYp6Y"
# At the end, you should see the evaluation scores for MRPC.
|
examples/notebooks/jiant_Basic_Example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 322} colab_type="code" id="c2ut_4TUa9YI" outputId="0db74731-aa27-490b-d40c-36f32e4f8a82"
# %tensorflow_version 1.x
# %cd /content
# !pip install nose
# !rm -rf boltzmann-machines
# !git clone --single-branch --depth=1 --branch with-models https://github.com/hannesdm/boltzmann-machines.git
# %cd boltzmann-machines
import numpy as np
import matplotlib.pyplot as plt
plt.rcParams['image.cmap'] = 'gray'
plt.rcParams['axes.grid'] = False
from examples import callable_dbm_mnist
from keras.datasets import mnist
from sklearn.neural_network import BernoulliRBM
from sklearn import preprocessing
from sklearn.metrics import accuracy_score
from boltzmann_machines import DBM
from boltzmann_machines.rbm import BernoulliRBM
from boltzmann_machines.utils import im_plot, Stopwatch
# + [markdown] colab_type="text" id="h76_RkCZbw1g"
# ## Load the mnist data
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" id="umyVA70DYEmM" outputId="d21a7f08-d0b4-494a-e906-ae01fb49e321"
(X_train, Y_train), (X_test, Y_test) = mnist.load_data()
X_train = X_train.reshape((X_train.shape[0], -1))
X_test = X_test.reshape((X_test.shape[0],-1))
X_train = X_train.astype('float32') / 255
X_test = X_test.astype('float32') / 255
lb = preprocessing.LabelBinarizer()
Y_train_one_hot = lb.fit_transform(Y_train)
Y_test_one_hot = lb.transform(Y_test)
# + [markdown] colab_type="text" id="HegMeKq0b8s0"
# ## Load a pretrained model and visualise the components
# To prevent losing too much time on training a deep model, a working pretrained RBM will be used. <br/>
# **Exercise** Observe the components (weights of each neuron) of the DBM and compare with the components of the previously trained RBM in exercise 1. What is the difference?<br/>
# Can you explain the difference between the components of the first and second layer of the DBM?
#
# + colab={"base_uri": "https://localhost:8080/", "height": 373} colab_type="code" id="s0X2KqR_fUU5" outputId="d4a2b54c-92cc-4004-a4e3-2935e9eee46d"
rbm1 = BernoulliRBM.load_model('/content/boltzmann-machines/models/dbm_mnist_rbm1/')
rbm2 = BernoulliRBM.load_model('/content/boltzmann-machines/models/dbm_mnist_rbm2/')
dbm = DBM.load_model('/content/boltzmann-machines/models/dbm_mnist/')
dbm.load_rbms([rbm1, rbm2])
# + colab={"base_uri": "https://localhost:8080/", "height": 599} colab_type="code" id="6VM7wmSKfxHl" outputId="cd5edae6-0357-44b3-cd71-5915458e2a3c"
W1_joint = dbm.get_tf_params(scope='weights')['W']
fig = plt.figure(figsize=(10, 10))
im_plot(W1_joint.T, shape=(28, 28), imshow_params={'cmap': plt.cm.gray})
plt.savefig("figure2.png")
# + colab={"base_uri": "https://localhost:8080/", "height": 599} colab_type="code" id="7EXVU-3_f4dA" outputId="e2cc7e45-0103-400c-81d6-129b09a42d02"
W2_joint = dbm.get_tf_params(scope='weights')['W_1']
U_joint = W1_joint.dot(W2_joint)
fig = plt.figure(figsize=(10, 10))
im_plot(U_joint.T, shape=(28, 28), imshow_params={'cmap': plt.cm.gray})
plt.savefig("figure3.png")
# + [markdown] colab_type="text" id="HYaG0cv2gtfG"
# ## Sample the DBM
# **Exercise** Comment on the quality of the samples and compare them with the samples from the RBM.<br/>
# Do you see a difference in quality? Explain why.
# + colab={"base_uri": "https://localhost:8080/", "height": 616} colab_type="code" id="pqILfLzKgAv-" outputId="2dac812f-e00c-42c0-b25f-a4197f392505"
gibbs_steps = 100
with Stopwatch(verbose=True) as s:
V = dbm.sample_v(n_gibbs_steps=gibbs_steps)
fig = plt.figure(figsize=(10, 10))
im_plot(V, shape=(28, 28),
imshow_params={'cmap': plt.cm.gray})
plt.savefig("figure4.png")
|
assignment 4/DBM.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] _uuid="570fb89bb4f7838b1d0fdff70b2935790e5dbdbe" lang="en"
# <center>
# <img src="https://habrastorage.org/files/fd4/502/43d/fd450243dd604b81b9713213a247aa20.jpg">
#
# ## [mlcourse.ai](mlcourse.ai) – Open Machine Learning Course
#
# <center>Auteur: [<NAME>](http://yorko.github.io) <br>
# Traduit et édité par [<NAME>](https://www.linkedin.com/in/isvforall/), [<NAME>](https://www.linkedin.com/in/datamove/), [<NAME>](https://www.linkedin.com/in/anastasiamanokhina/), [<NAME>](https://www.linkedin.com/in/yuanyuanpao/) et [<NAME>](https://github.com/oussou-dev)<br>Contenu soumis à la licence [Creative Commons CC BY-NC-SA 4.0](https://creativecommons.org/licenses/by-nc-sa/4.0/).
# + [markdown] _uuid="998fc5f76227ab1df9460a7b05c508304b14b3a6"
# # <center> Devoir #1 (démo)
# ## <center> Analyse exploratoire des données avec Pandas
#
# + [markdown] lang="fr"
# **Dans cette tâche, vous devez utiliser la librairie Pandas pour répondre à quelques questions liées au dataset (ensemble de données) [Adulte](https://archive.ics.uci.edu/ml/datasets/Adult). (Vous n'avez pas à télécharger les données - vous les trouverez ci-joints). Choisissez vos réponses via ce [formulaire](https://docs.google.com/forms/d/1uY7MpI2trKx6FLWZte0uVh3ULV4Cm_tDud0VDFGCOKg). C'est une version de démonstration. Ainsi, en soumettant le formulaire, vous verrez un lien vers le fichier .ipynb de la solution.**
# + [markdown] _uuid="c322301ac858c748f8f80e63cf6f734fc70dde30" lang="en"
# Valeurs uniques de toutes les caractéristiques (pour plus d'informations, veuillez consulter les liens ci-dessus):
# - `age`: continuous.
# - `workclass`: Private, Self-emp-not-inc, Self-emp-inc, Federal-gov, Local-gov, State-gov, Without-pay, Never-worked.
# - `fnlwgt`: continuous.
# - `education`: Bachelors, Some-college, 11th, HS-grad, Prof-school, Assoc-acdm, Assoc-voc, 9th, 7th-8th, 12th, Masters, 1st-4th, 10th, Doctorate, 5th-6th, Preschool.
# - `education-num`: continuous.
# - `marital-status`: Married-civ-spouse, Divorced, Never-married, Separated, Widowed, Married-spouse-absent, Married-AF-spouse.
# - `occupation`: Tech-support, Craft-repair, Other-service, Sales, Exec-managerial, Prof-specialty, Handlers-cleaners, Machine-op-inspct, Adm-clerical, Farming-fishing, Transport-moving, Priv-house-serv, Protective-serv, Armed-Forces.
# - `relationship`: Wife, Own-child, Husband, Not-in-family, Other-relative, Unmarried.
# - `race`: White, Asian-Pac-Islander, Amer-Indian-Eskimo, Other, Black.
# - `sex`: Female, Male.
# - `capital-gain`: continuous.
# - `capital-loss`: continuous.
# - `hours-per-week`: continuous.
# - `native-country`: United-States, Cambodia, England, Puerto-Rico, Canada, Germany, Outlying-US(Guam-USVI-etc), India, Japan, Greece, South, China, Cuba, Iran, Honduras, Philippines, Italy, Poland, Jamaica, Vietnam, Mexico, Portugal, Ireland, France, Dominican-Republic, Laos, Ecuador, Taiwan, Haiti, Columbia, Hungary, Guatemala, Nicaragua, Scotland, Thailand, Yugoslavia, El-Salvador, Trinadad&Tobago, Peru, Hong, Holand-Netherlands.
# - `salary`: >50K,<=50K
# + [markdown] lang="fr"
# Valeurs uniques de toutes les caractéristiques (pour plus d'informations, veuillez consulter les liens ci-dessus):
#
# - `age`: continuous.
# - `workclass`: Private, Self-emp-not-inc, Self-emp-inc, Federal-gov, Local-gov, State-gov, Without-pay, Never-worked.
# - `fnlwgt`: continuous.
# - `education`: Bachelors, Some-college, 11th, HS-grad, Prof-school, Assoc-acdm, Assoc-voc, 9th, 7th-8th, 12th, Masters, 1st-4th, 10th, Doctorate, 5th-6th, Preschool.
# - `education-num`: continuous.
# - `marital-status`: Married-civ-spouse, Divorced, Never-married, Separated, Widowed, Married-spouse-absent, Married-AF-spouse.
# - `occupation`: Tech-support, Craft-repair, Other-service, Sales, Exec-managerial, Prof-specialty, Handlers-cleaners, Machine-op-inspct, Adm-clerical, Farming-fishing, Transport-moving, Priv-house-serv, Protective-serv, Armed-Forces.
# - `relationship`: Wife, Own-child, Husband, Not-in-family, Other-relative, Unmarried.
# - `race`: White, Asian-Pac-Islander, Amer-Indian-Eskimo, Other, Black.
# - `sex`: Female, Male.
# - `capital-gain`: continuous.
# - `capital-loss`: continuous.
# - `hours-per-week`: continuous.
# - `native-country`: United-States, Cambodia, England, Puerto-Rico, Canada, Germany, Outlying-US(Guam-USVI-etc), India, Japan, Greece, South, China, Cuba, Iran, Honduras, Philippines, Italy, Poland, Jamaica, Vietnam, Mexico, Portugal, Ireland, France, Dominican-Republic, Laos, Ecuador, Taiwan, Haiti, Columbia, Hungary, Guatemala, Nicaragua, Scotland, Thailand, Yugoslavia, El-Salvador, Trinadad&Tobago, Peru, Hong, Holand-Netherlands.
# - `salary`: >50K,<=50K
# + _uuid="d3eb2411e23b7db11e4cbbb498a42752442a6f4c"
import numpy as np
import pandas as pd
# + _uuid="c5119d1b8151029b4ff57735c4279b795ae492a2"
# lire le csv
data = pd.read_csv(
"adult.data.csv"
) # destination du fichier à adapter (../dossier/adult.data.csv)
# afficher les 5 premières lignes du dataset
data.head()
# + [markdown] lang="fr"
# **1. Combien d'hommes et de femmes (* caractéristique sex * ) sont représentés dans ce dataset ?**
# + _uuid="d32cafcbe966dbdfa40c3017d16d6f8c96bfb3e2"
# ton code
# + [markdown] lang="fr"
# **2. Quel est l’âge moyen des femmes?**
# + _uuid="64f60d928855107d8980875656dcdf3587d91646"
# ton code
# + [markdown] lang="fr"
# **3. Quel est le pourcentage de citoyens allemands (caractéristique *native-country*)?**
# + _uuid="ca9807509692638bd945671ed7d8b4a5ad1f3626"
# ton code
# + [markdown] lang="fr"
# **4-5. Quels sont la moyenne et la déviation standard (écart-type) de l’âge pour ceux qui gagnent plus de 50k par an (caractéristique * salary *) et ceux qui gagnent moins de 50k par an?**
# + _uuid="97948f0115f949913d211627151b5605bcd204fd"
# ton code
# + [markdown] lang="fr"
# **6. Est-il vrai que les personnes qui gagnent plus de 50k ont au moins terminé leurs études secondaires? (* education - Bachelors, Prof-school, Assoc-acdm, Assoc-voc, Masters * ou * Doctorat *)**
# + _uuid="58f38ebfa2b0e1418a6a1a680d0743697f9876a2"
# ton code
# + [markdown] lang="fr"
# **7. Affichez les statistiques d'âge pour chaque race (caractéristique * race *) et chaque sexe (caractéristique * sex *). Utilisez * groupby () * et * describe () *. Trouvez l'âge maximum des hommes de race * Amer-Indian-Eskimo *.**
# + _uuid="a4567b72200b583f7f2ee1583405d636c12169db"
# ton code
# + [markdown] lang="fr"
# **8. Parmi ceux qui gagnent plus de 50k, quel est l'état matrimonial (caractéristique marital-status) : marié ou célibataire le plus représenté ?
# On considère comme mariés ceux qui ont un marital-status commençant par Married (Married-civ-spouse, Married-spouse-absent or Married-AF-spouse), les autres sont considérés comme célibataires (bachelors).**
# + _uuid="5bb3a8524fcbd794707075ccf6aaf8dd0f30a3dd"
# ton code
# + [markdown] lang="fr"
# **9. Quel est le nombre d'heures maximum de travail par semaine effectué par une personne (caractéritique * hours-per-week *)?
# Combien de personnes travaillent autant, et quel est le pourcentage de celles qui gagnent beaucoup (>50k) parmi elles?**
# + _uuid="3571468172fc4bcf34e10b3d6ab358d4e41f7ef4"
# ton code
# + [markdown] lang="fr"
# **10. Calculez le temps de travail moyen (*hours-per-week*) pour ceux qui sont peu et beaucoup payés (*salary*) pour chaque pays (*native-country*). Qu'obtient-on pour le Japon?**
# + _uuid="75a93411f1e3b519afcc27c026aaa0c67eb526e9"
# ton code
|
jupyter_french/assignments_demo/a1-demo-pandas-and-uci-adult-dataset-fr_def.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# +
import importlib
import theano.tensor as T
import sys, os
sys.path.append("/home/bl3/PycharmProjects/GeMpy/")
sys.path.append("/home/bl3/PycharmProjects/pygeomod/pygeomod")
import GeoMig
#import geogrid
#importlib.reload(GeoMig)
importlib.reload(GeoMig)
import numpy as np
os.environ['CUDA_LAUNCH_BLOCKING'] = '1'
np.set_printoptions(precision = 15, linewidth= 300, suppress = True)
from mpl_toolkits.mplot3d import axes3d
import matplotlib.pyplot as plt
from matplotlib import cm
# -
# +
test = GeoMig.GeoMigSim_pro2(c_o = np.float32(-0.1),range = 17)
test.create_regular_grid_3D(0,10,0,10,0,10,20,20,20)
test.theano_set_3D_nugget_degree0()
# +
layer_1 = np.array([[1,5,7], [9,5,7]], dtype = "float32")
layer_2 = np.array([[2,5,1],[7,5,1]], dtype = "float32")
dip_pos_1 = np.array([2,5,6], dtype = "float32")
dip_pos_2 = np.array([6.,4,6], dtype = "float32")
dip_pos_3 = np.array([8,4,5], dtype = "float32")
dip_angle_1 = float(0)
dip_angle_2 = float(45)
layers = np.asarray([layer_1,layer_2])
dips = np.asarray([dip_pos_1])#, dip_pos_3])
dips_angles = np.asarray([dip_angle_1], dtype="float32")
azimuths = np.asarray([0], dtype="float32")
polarity = np.asarray([1], dtype="float32")
#print (dips_angles)
rest = np.vstack((i[1:] for i in layers))
ref = np.vstack((np.tile(i[0],(np.shape(i)[0]-1,1)) for i in layers))
dips_angles.dtype
rest = rest.astype("float32")
ref = ref.astype("float32")
dips = dips.astype("float32")
dips_angles = dips_angles.astype("float32")
type(dips_angles)
# -
rest, ref
rest, ref
# +
G_x = np.sin(np.deg2rad(dips_angles)) * np.sin(np.deg2rad(azimuths)) * polarity
G_y = np.sin(np.deg2rad(dips_angles)) * np.cos(np.deg2rad(azimuths)) * polarity
G_z = np.cos(np.deg2rad(dips_angles)) * polarity
G_x, G_y, G_z
# -
test.geoMigueller(dips,dips_angles,azimuths,polarity, rest, ref)[0]
_,h1 = np.argmin((abs(test.grid - ref[0])).sum(1)), test.geoMigueller(dips,dips_angles,azimuths,polarity, rest, ref)[0][np.argmin((abs(test.grid - ref[0])).sum(1))]
_, h2 =np.argmin((abs(test.grid - ref[1])).sum(1)), test.geoMigueller(dips,dips_angles,azimuths,polarity, rest, ref)[0][np.argmin((abs(test.grid - ref[1])).sum(1))]
print(test.geoMigueller(dips,dips_angles,azimuths,polarity, rest, ref)[0][np.argmin((abs(test.grid - ref[0])).sum(1))])
for i in range(rest.shape[0]):
print(test.geoMigueller(dips,dips_angles,azimuths,polarity, rest, ref)[0][np.argmin((abs(test.grid - rest[i])).sum(1))])
test.geoMigueller(dips,dips_angles,azimuths,polarity, rest, ref)[0][np.argmin((abs(test.grid - rest[0])).sum(1))]
rest
sol = test.geoMigueller(dips,dips_angles,azimuths,polarity, rest, ref)[0].reshape(200,200,200, order = "C")[:,:,::-1].transpose()
#sol = np.swapaxes(sol,0,1)
G_x, G_y, G_z = test.geoMigueller(dips,dips_angles,azimuths,polarity, rest, ref)[-3:]
G_x, G_y, G_z
# +
import numpy as np
from mpl_toolkits.mplot3d import Axes3D
import matplotlib.pyplot as plt
import matplotlib
import matplotlib.cm as cmx
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
h = np.array([h1,h2])
cm = plt.get_cmap("jet")
cNorm = matplotlib.colors.Normalize(vmin=h.min(), vmax=h.max())
scalarMap = cmx.ScalarMappable(norm=cNorm, cmap=cm)
sol = test.geoMigueller(dips,dips_angles,azimuths,polarity, rest, ref)[0].reshape(200,200,200,
order = "C")[:,:,:]
#sol = np.swapaxes(sol,0,1)
from skimage import measure
isolines = np.linspace(h1,h2,2)
#vertices = measure.marching_cubes(sol, isolines[0], spacing = (0.2,0.2,0.2),
# gradient_direction = "descent")[0]
for i in isolines[0:10]:
vertices = measure.marching_cubes(sol, i, spacing = (0.05,0.05,0.05),
gradient_direction = "ascent")[0]
ax.scatter(vertices[::40,0],vertices[::40,1],vertices[::40,2],color=scalarMap.to_rgba(i),
alpha = 0.2) #color=scalarMap.to_rgba(vertices[::10,2])
ax.scatter(layers[0][:,0],layers[0][:,1],layers[0][:,2], s = 50, c = "r" )
ax.scatter(layers[1][:,0],layers[1][:,1],layers[1][:,2], s = 50, c = "g" )
ax.quiver3D(dips[:,0],dips[:,1],dips[:,2], G_x,G_y,G_z, pivot = "tail", linewidths = 2)
ax.set_xlabel("x")
ax.set_ylabel("y")
ax.set_zlabel("z")
ax.set_xlim(0,10)
ax.set_ylim(0,10)
ax.set_zlim(0,10)
#ax.scatter(simplices[:,0],simplices[:,1],simplices[:,2])
# -
test.c_o.set_value(-0.56)
test.c_o.get_value()
test.geoMigueller(dips,dips_angles,azimuths,polarity, rest, ref,1,1,1,1,1,1)[1]
c_sol = np.array(([-7.2386541205560206435620784759521484375E-14],
[-1.5265566588595902430824935436248779296875E-14],
[-1.154631945610162802040576934814453125E-14],
[6.21724893790087662637233734130859375E-15],
[-5.9952043329758453182876110076904296875E-15],
[7.99360577730112709105014801025390625E-15],
[2.220446049250313080847263336181640625E-15],
[-3.641531520770513452589511871337890625E-14],
[8.0380146982861333526670932769775390625E-14],
[0.8816416857576581111999303175252862274646759033203125],
[9.355249580684368737593104015104472637176513671875],
[-0.1793850547262900996248191631821100600063800811767578125],
[0.047149729032205163481439313954979297704994678497314453125],
[-8.994519501910499315044944523833692073822021484375],
[ 0.4451793036427798000431721447966992855072021484375],
[-1.7549816402777651536126768405665643513202667236328125],
[0.0920938443689063301889063950511626899242401123046875],
[0.36837537747562587586713789278292097151279449462890625])).squeeze()
c_sol.squeeze()
c_sol=np.array([ -0.07519608514102089913411219868066837079823017120361328125,
0,
3.33264951481644633446421721600927412509918212890625,
1.3778510792932487927231477442546747624874114990234375,
-2.295940519242440469582788864499889314174652099609375,
])
# +
import pymc as pm
a = pm.Uniform('a', lower=-1.1, upper=1.1, )
b = pm.Uniform('b', lower=-1.1, upper=1.1, )
c = pm.Uniform('c', lower=-1.1, upper=1.1, )
d = pm.Uniform('d', lower=-1.1, upper=1.1, )
e = pm.Uniform('e', lower=-1.1, upper=1.1, )
f = pm.Uniform('f', lower=-1.1, upper=1.1, )
@pm.deterministic
def this(value = 0, a = a ,b = b,c = c,d = d,e= e,f =f, c_sol = c_sol):
sol = test.geoMigueller(dips,dips_angles,azimuths,polarity, rest, ref,
a,b,-3*b,d,e,f)[1]
#error = abs(sol-c_sol)
#print (error)
return sol
like= pm.Normal("likelihood", this, 1./np.square(0.0000000000000000000000000000000000000000000000001),
value = c_sol, observed = True, size = len(c_sol)
)
model = pm.Model([a,b,c,d,e,f, like])
# -
M = pm.MAP(model)
M.fit()
a.value, b.value, c.value,d.value, e.value, f.value, this.value, c_sol, test.geoMigueller(dips,dips_angles,azimuths,polarity, rest, ref,1,1,1,1,1,1)[1]
a.value, b.value, c.value,d.value, e.value, f.value, this.value, c_sol, test.geoMigueller(dips,dips_angles,azimuths,polarity, rest, ref,1,1,1,1,1,1)[1]
a.value, b.value, c.value,d.value, e.value, f.value, this.value, c_sol, test.geoMigueller(dips,dips_angles,azimuths,polarity, rest, ref,1,1,1,1,1,1)[1]
a.value, b.value, c.value,d.value, e.value, f.value, this.value, c_sol, test.geoMigueller(dips,dips_angles,azimuths,polarity, rest, ref,a,b,-3*b,d,1,1)[1]
-3*b.value
a.value, b.value, c.value,d.value, e.value, f.value, this.value, c_sol, test.geoMigueller(dips,dips_angles,azimuths,polarity, rest, ref,1,1,1,1,1,1)[1]
a.value, b.value, c.value,d.value, e.value, f.value, this.value, c_sol, test.geoMigueller(dips,dips_angles,azimuths,polarity, rest, ref,1,1,1,1,1,1)[1]
a.value, b.value, c.value,d.value, e.value, f.value, this.value, c_sol, test.geoMigueller(dips,dips_angles,azimuths,polarity, rest, ref,1,1,1,1,1,1)[1]
a.value, b.value, c.value,d.value, e.value, f.value, this.value, c_sol, test.geoMigueller(dips,dips_angles,azimuths,polarity, rest, ref,1,1,1,1,1,1)[1]
test.geoMigueller(dips,dips_angles,azimuths,polarity, rest, ref,0,0,-0.33,0,1,1)[1]
# # Test with all variables
a.value, b.value, c.value,d.value,e.value,f.value, this.value, c_sol, test.geoMigueller(dips,dips_angles,azimuths,polarity, rest, ref,a,b,1,1,1,1)[1]
a.value, b.value, c.value,d.value,e.value,f.value, this.value, c_sol, test.geoMigueller(dips,dips_angles,azimuths,polarity, rest, ref,1,1,1,1,1,1)[1]
importlib.reload(GeoMig)
test = GeoMig.GeoMigSim_pro2(c_o = np.float32(-0.1),range = 17)
test.create_regular_grid_3D(0,10,0,10,0,10,20,20,20)
test.theano_set_3D_nugget_degree0()
# +
import matplotlib.pyplot as plt
# %matplotlib inline
G_x, G_y, G_z = test.geoMigueller(dips,dips_angles,azimuths,polarity, rest, ref,1,1,1,1,1,1)[-3:]
sol = test.geoMigueller(dips,dips_angles,azimuths,polarity, rest, ref,a,b,-3*b,d,1,-1)[0].reshape(20,20,20)
def plot_this_crap(direction):
fig = plt.figure()
ax = fig.add_subplot(111)
if direction == "x":
plt.arrow(dip_pos_1[1],dip_pos_1[2], dip_pos_1_v[1]-dip_pos_1[1],
dip_pos_1_v[2]-dip_pos_1[2], head_width = 0.2)
plt.arrow(dip_pos_2[1],dip_pos_2[2],dip_pos_2_v[1]-dip_pos_2[1],
dip_pos_2_v[2]-dip_pos_2[2], head_width = 0.2)
plt.plot(layer_1[:,1],layer_1[:,2], "o")
plt.plot(layer_2[:,1],layer_2[:,2], "o")
plt.plot(layer_1[:,1],layer_1[:,2], )
plt.plot(layer_2[:,1],layer_2[:,2], )
plt.contour( sol[25,:,:] ,30,extent = (0,10,0,10) )
if direction == "y":
plt.quiver(dips[:,0],dips[:,2], G_x,G_z, pivot = "tail")
plt.plot(layer_1[:,0],layer_1[:,2], "o")
plt.plot(layer_2[:,0],layer_2[:,2], "o")
plt.plot(layer_1[:,0],layer_1[:,2], )
plt.plot(layer_2[:,0],layer_2[:,2], )
plt.contour( sol[:,10,:].T ,30,extent = (0,10,0,10) )
if direction == "z":
plt.arrow(dip_pos_1[0],dip_pos_1[1], dip_pos_1_v[0]-dip_pos_1[0],
dip_pos_1_v[1]-dip_pos_1[1], head_width = 0.2)
plt.arrow(dip_pos_2[0],dip_pos_2[1],dip_pos_2_v[0]-dip_pos_2[0],
dip_pos_2_v[1]-dip_pos_2[1], head_width = 0.2)
plt.plot(layer_1[:,0],layer_1[:,1], "o")
plt.plot(layer_2[:,0],layer_2[:,1], "o")
plt.plot(layer_1[:,0],layer_1[:,1], )
plt.plot(layer_2[:,0],layer_2[:,1], )
plt.contour( sol[:,:,25] ,30,extent = (0,10,0,10) )
#plt.colorbar()
#plt.xlim(0,10)
#plt.ylim(0,10)
plt.colorbar()
plt.title("GeoBulleter v 0.1")
# -
plot_this_crap("y")
a.value, b.value
test.geoMigueller(dips,dips_angles,azimuths,polarity, rest, ref,1,1,1,1,1,1)[1]
c_sol
# +
h,j,k =sol[5,10,35], sol[25,5,5], sol[30,15,-25]
layer_1 = np.array([[1,5,7],[5,5,7],[6,5,7], [9,5,7]], dtype = "float32")
layer_2 = np.array([[1,5,1],[5,5,1],[9,5,1]], dtype = "float32")
print(sol[5,25,35], sol[25,25,35], sol[30,25,35], sol[45,25,35])
print(sol[5,25,5], sol[25,25,5], sol[45,25,5])
# -
list(layer_1[0]*5)
# +
interfaces_aux = test.geoMigueller(dips,dips_angles,azimuths,polarity,
rest, ref)[0]
h = sol[10,20,30]# interfaces_aux[np.argmin(abs((test.grid - ref[0]).sum(1)))]
k = sol[30,15,25]# interfaces_aux[np.argmin(abs((test.grid - dips[0]).sum(1)))]
j = sol[45,25,5]#interfaces_aux[np.argmin(abs((test.grid - dips[-1]).sum(1)))]
h,k,j
# -
dips[-1], ref[0]
sol[30,15,25], sol[30,15,25]
sol = test.geoMigueller(dips,dips_angles,azimuths,polarity, rest, ref)[0].reshape(50,50,50, order = "C")
sol = np.swapaxes(sol,0,1)
plt.contour(sol[:,25,:].transpose())
# +
"""Export model to VTK
Export the geology blocks to VTK for visualisation of the entire 3-D model in an
external VTK viewer, e.g. Paraview.
..Note:: Requires pyevtk, available for free on: https://github.com/firedrakeproject/firedrake/tree/master/python/evtk
**Optional keywords**:
- *vtk_filename* = string : filename of VTK file (default: output_name)
- *data* = np.array : data array to export to VKT (default: entire block model)
"""
vtk_filename = "noddyFunct2"
extent_x = 10
extent_y = 10
extent_z = 10
delx = 0.2
dely = 0.2
delz = 0.2
from pyevtk.hl import gridToVTK
# Coordinates
x = np.arange(0, extent_x + 0.1*delx, delx, dtype='float64')
y = np.arange(0, extent_y + 0.1*dely, dely, dtype='float64')
z = np.arange(0, extent_z + 0.1*delz, delz, dtype='float64')
# self.block = np.swapaxes(self.block, 0, 2)
gridToVTK(vtk_filename, x, y, z, cellData = {"geology" : sol})
# -
len(x)
surf_eq.min()
np.min(z)
layers[0][:,0]
G_x = np.sin(np.deg2rad(dips_angles)) * np.sin(np.deg2rad(azimuths)) * polarity
G_y = np.sin(np.deg2rad(dips_angles)) * np.cos(np.deg2rad(azimuths)) * polarity
G_z = np.cos(np.deg2rad(dips_angles)) * polarity
a
data = [trace1, trace2]
layout = go.Layout(
xaxis=dict(
range=[2, 5]
),
yaxis=dict(
range=[2, 5]
)
)
fig = go.Figure(data=data, layout=layout)
# +
import lxml
# lxml??
# +
# Random Box
#layers = [np.random.uniform(0,10,(10,2)) for i in range(100)]
#dips = np.random.uniform(0,10, (60,2))
#dips_angles = np.random.normal(90,10, 60)
#rest = (np.vstack((i[1:] for i in layers)))
#ref = np.vstack((np.tile(i[0],(np.shape(i)[0]-1,1)) for i in layers))
#rest;
# -
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
X, Y, Z = axes3d.get_test_data(0.05)
cset = ax.contour(X, Y, Z, cmap=cm.coolwarm)
ax.clabel(cset, fontsize=9, inline=1)
print(X)
plt.show()
import matplotlib.pyplot as plt
% matplotlib inline
plt.contour( sol.reshape(100,100) ,30,extent = (0,10,0,10) )
# +
import matplotlib.pyplot as plt
% matplotlib inline
dip_pos_1_v = np.array([np.cos(np.deg2rad(dip_angle_1))*1,
np.sin(np.deg2rad(dip_angle_1))]) + dip_pos_1
dip_pos_2_v = np.array([np.cos(np.deg2rad(dip_angle_2))*1,
np.sin(np.deg2rad(dip_angle_2))]) + dip_pos_2
plt.arrow(dip_pos_1[0],dip_pos_1[1], dip_pos_1_v[0]-dip_pos_1[0],
dip_pos_1_v[1]-dip_pos_1[1], head_width = 0.2)
plt.arrow(dip_pos_2[0],dip_pos_2[1],dip_pos_2_v[0]-dip_pos_2[0],
dip_pos_2_v[1]-dip_pos_2[1], head_width = 0.2)
plt.plot(layer_1[:,0],layer_1[:,1], "o")
plt.plot(layer_2[:,0],layer_2[:,1], "o")
plt.plot(layer_1[:,0],layer_1[:,1], )
plt.plot(layer_2[:,0],layer_2[:,1], )
plt.contour( sol.reshape(100,100) ,30,extent = (0,10,0,10) )
#plt.colorbar()
#plt.xlim(0,10)
#plt.ylim(0,10)
plt.title("GeoBulleter v 0.1")
print (dip_pos_1_v, dip_pos_2_v, layer_1)
# -
# # CPU
# %%timeit
sol = test.geoMigueller(dips,dips_angles,rest, ref)[0]
test.geoMigueller.profile.summary()
# +
sys.path.append("/home/bl3/anaconda3/lib/python3.5/site-packages/PyEVTK-1.0.0-py3.5.egg_FILES/pyevtk")
nx = 50
ny = 50
nz = 50
xmin = 1
ymin = 1
zmin = 1
grid = sol
var_name = "Geology"
#from evtk.hl import gridToVTK
import pyevtk
from pyevtk.hl import gridToVTK
# define coordinates
x = np.zeros(nx + 1)
y = np.zeros(ny + 1)
z = np.zeros(nz + 1)
x[1:] = np.cumsum(delx)
y[1:] = np.cumsum(dely)
z[1:] = np.cumsum(delz)
# plot in coordinates
x += xmin
y += ymin
z += zmin
print (len(x), x)
gridToVTK("GeoMigueller", x, y, z,
cellData = {var_name: grid})
# -
# ## GPU
# %%timeit
sol = test.geoMigueller(dips,dips_angles,azimuths,polarity, rest, ref);
test.geoMigueller.profile.summary()
importlib.reload(GeoMig)
test = GeoMig.GeoMigSim_pro2()
# +
from theano import function, config, shared, sandbox
import theano.tensor as T
import numpy
import time
vlen = 10 * 30 * 768 # 10 x #cores x # threads per core
iters = 1000
rng = numpy.random.RandomState(22)
x = shared(numpy.asarray(rng.rand(vlen), config.floatX))
f = function([], T.exp(x))
print(f.maker.fgraph.toposort())
t0 = time.time()
for i in range(iters):
r = f()
t1 = time.time()
print("Looping %d times took %f seconds" % (iters, t1 - t0))
print("Result is %s" % (r,))
if numpy.any([isinstance(x.op, T.Elemwise) for x in f.maker.fgraph.toposort()]):
print('Used the cpu')
else:
print('Used the gpu')
# +
from theano import function, config, shared, sandbox
import theano.tensor as T
import numpy
import time
vlen = 10 * 30 * 768 # 10 x #cores x # threads per core
iters = 1000
rng = numpy.random.RandomState(22)
x = shared(numpy.asarray(rng.rand(vlen), config.floatX))
f = function([], T.exp(x))
print(f.maker.fgraph.toposort())
t0 = time.time()
for i in range(iters):
r = f()
t1 = time.time()
print("Looping %d times took %f seconds" % (iters, t1 - t0))
print("Result is %s" % (r,))
if numpy.any([isinstance(x.op, T.Elemwise) for x in f.maker.fgraph.toposort()]):
print('Used the cpu')
else:
print('Used the gpu')
# +
from theano import function, config, shared, sandbox
import theano.tensor as T
import numpy
import time
vlen = 10 * 30 * 768 # 10 x #cores x # threads per core
iters = 1000
rng = numpy.random.RandomState(22)
x = shared(numpy.asarray(rng.rand(vlen), config.floatX))
f = function([], T.exp(x))
print(f.maker.fgraph.toposort())
t0 = time.time()
for i in range(iters):
r = f()
t1 = time.time()
print("Looping %d times took %f seconds" % (iters, t1 - t0))
print("Result is %s" % (r,))
if numpy.any([isinstance(x.op, T.Elemwise) for x in f.maker.fgraph.toposort()]):
print('Used the cpu')
else:
print('Used the gpu')
# -
np.set_printoptions(precision=2)
test.geoMigueller(dips,dips_angles,rest, ref)[1]
# +
# T.fill_diagonal?
# +
import matplotlib.pyplot as plt
% matplotlib inline
dip_pos_1_v = np.array([np.cos(np.deg2rad(dip_angle_1))*1,
np.sin(np.deg2rad(dip_angle_1))]) + dip_pos_1
dip_pos_2_v = np.array([np.cos(np.deg2rad(dip_angle_2))*1,
np.sin(np.deg2rad(dip_angle_2))]) + dip_pos_2
plt.arrow(dip_pos_1[0],dip_pos_1[1], dip_pos_1_v[0]-dip_pos_1[0],
dip_pos_1_v[1]-dip_pos_1[1], head_width = 0.2)
plt.arrow(dip_pos_2[0],dip_pos_2[1],dip_pos_2_v[0]-dip_pos_2[0],
dip_pos_2_v[1]-dip_pos_2[1], head_width = 0.2)
plt.plot(layer_1[:,0],layer_1[:,1], "o")
plt.plot(layer_2[:,0],layer_2[:,1], "o")
plt.plot(layer_1[:,0],layer_1[:,1], )
plt.plot(layer_2[:,0],layer_2[:,1], )
plt.contour( sol.reshape(50,50) ,30,extent = (0,10,0,10) )
#plt.colorbar()
#plt.xlim(0,10)
#plt.ylim(0,10)
plt.title("GeoBulleter v 0.1")
print (dip_pos_1_v, dip_pos_2_v, layer_1)
# +
n = 10
#a = T.horizontal_stack(T.vertical_stack(T.ones(n),T.zeros(n)), T.vertical_stack(T.zeros(n), T.ones(n)))
a = T.zeros(n)
print (a.eval())
#U_G = T.horizontal_stack(([T.ones(n),T.zeros(n)],[T.zeros(n),T.ones(n)]))
# -
T.stack?ö+aeg
# +
x_min = 0
x_max = 10
y_min = 0
y_max = 10
z_min = 0
z_max = 10
nx = 2
ny = 2
nz = 2
g = np.meshgrid(
np.linspace(x_min, x_max, nx, dtype="float32"),
np.linspace(y_min, y_max, ny, dtype="float32"),
np.linspace(z_min, z_max, nz, dtype="float32"), indexing="ij"
)
np.vstack(map(np.ravel, g)).T.astype("float32")
# -
map(np.ravel, g)
np.ravel(g, order = "F")
g
# +
# np.transpose?
# -
from scipy.optimize import basinhopping
c_sol, test.geoMigueller(dips,dips_angles,azimuths,polarity, rest, ref,1,1,1,1,1,1)[1]
def func2d(x):
return abs((test.geoMigueller(dips,dips_angles,azimuths,polarity, rest, ref,x[0],x[1],x[2],x[3],1,1)[1] - c_sol)).sum()
minimizer_kwargs = {"method": "BFGS"}
x0 = [0.1, 0.1,0.1,0.1]
ret = basinhopping(func2d, x0, minimizer_kwargs=minimizer_kwargs,
niter=200)
ret
ret
ret
|
.ipynb_checkpoints/Geothealler big function 3D-Testing-degree0-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # International Suicide Rates 1985-2016: Exploratory Data Analysis
#
# ### Author: <NAME>
# ## Description of dataset:
#
# The data have been collected from a Kaggle dataset (https://www.kaggle.com/russellyates88/suicide-rates-overview-1985-to-2016#master.csv). The data include socio-economic data for individuals who completed suicide in various countries between 1985 to 2016. As I was not involved in collection or curation, I cannot attest to the validity of the underlying data, but will proceed to use the dataset for exploratory data analysis purposes.
#
# In the U.S., rates of suicide have been increasing in nearly every state, according to the most recent Vital Signs report by the Centers for Disease Control and Prevention (CDC). It states that "in 2016, nearly 45,000 Americans age 10 or older died by suicide...which is the 10th leading cause of death and is one of just three leading causes that are on the rise" (https://www.cdc.gov/nchs/pressroom/sosmap/suicide-mortality/suicide.htm).
#
# While the phenomenon of suicide has a causal connection to psychiatric disorders that precipitate suicidal ideation, it is also worth exploring relationships between suicide rates across different socioeconomic and geographic regions (including similarities and contrasts between the U.S. and other countries). Statistically signficant relationships that emerge could guide clinical practice, community awareness and preventive efforts, and ultimately help shape health policy at the federal, state, and local levels of government.
#
# The dataset includes the following variables: country, year, sex, age, suicides_no, population, suicides/100k pop, country-year, HDI for year, gdp_for_year, gdp_per_capita, and generation.
#
# ## Exploratory Data Analysis
# +
#Load required Python packages into local environment
import numpy as np
import pandas as pd
import seaborn as sns
import ast, json
from datetime import datetime
import matplotlib.pyplot as plt
% matplotlib inline
# +
#Read in the file to construct a Pandas dataframe:
suicide_df = pd.read_csv('/Users/dannywitt/Desktop/IDS_705_ML/Homework/ids705/assignments/suicide_data.csv')
suicide_df.head()
# -
#Describe the summary statistics of the dataset:
suicide_df.describe()
#Describe the different data types of the variables in the dataset:
print(suicide_df.dtypes)
# +
#Check to see if any missing values (NaN):
print('Are there any missing values?: {}'.format(suicide_df.isnull().values.any()))
print('')
#How many values are missing values? And, where are these missing values located
#(i.e., how are they distributed across variables)?
print(suicide_df.isnull().sum())
# +
#Since there are many missing in one variable, I'm going to remove variable named "HDI for year" due to
#large number of missing values:
suicide_df_v2 = suicide_df.drop(columns=['HDI for year'])
#Analyze how many unique values exist in the categorical variables:
category_var = ['country','year','sex','age','generation']
suicide_df_v2[category_var].nunique()
# +
# Check the distribution of target variable 'suicides_no'
# Plot 1 - Distribution of the target variable
sns.distplot(suicide_df_v2['suicides_no'], color='#7f181b', kde=True, hist=False) ;
# +
# Analysis of top 10 country with highest median suicide numbers
# I decided to do a median value because the distribution (as seen above) is skewed to the right
df_v3 = suicide_df_v2.groupby(by=['country']).median()[['suicides_no','population','gdp_per_capita ($)']].sort_values('suicides_no',ascending=False).reset_index()
cat = list(df_v3.head(10)['country'])
print('Top 10 country with highest suicide median ')
print(df_v3.head(10))
# +
import seaborn as sns; sns.set()
import matplotlib
import matplotlib.pyplot as plt
# I'm interested in understanding how the top 10 countries' (based on number of suicides) suicide rates
#have changed year by year from 1985-2016.
df_v4 = suicide_df_v2.groupby(by=['country','year']).median()['suicides_no'].reset_index()
matplotlib.rcParams['figure.figsize'] = [10,5]
gs = matplotlib.gridspec.GridSpec(2,1)
ax1 = plt.subplot(gs[0,0])
ax2 = plt.subplot(gs[1,0])
# Plot 2: Number of suicides committed in top 3 highest rate countries:
for i in cat[:3] :
sns.lineplot(data=df_v4[df_v4['country']==i], x='year', y='suicides_no', ax=ax1) ;
ax1.legend(cat[:3], loc=7, bbox_to_anchor=(1.3, 0.5)) ;
ax1.set_title('Total Annual Suicide Counts of Top 3 Countries', size=15, fontweight='bold') ;
# Plot 3: Line plot depicting change in total suicides/yr by other countries
for i in cat[3:] :
sns.lineplot(data=df_v4[df_v4['country']==i], x='year', y='suicides_no', ax=ax2) ;
ax2.legend(cat[3:], loc=7, bbox_to_anchor=(1.3, 0.5)) ;
ax2.set_xlabel('Total Annual Suicide Counts of Remaining Countries', size=15, fontweight='bold') ;
# +
# Visualize the distribution of suicide numbers across (1) gender/sex and (2) age:
matplotlib.rcParams['figure.figsize'] = [15,5]
gs = matplotlib.gridspec.GridSpec(1,2)
ax1 = plt.subplot(gs[0,0])
ax2 = plt.subplot(gs[0,1])
# Plot 4: Distribution of suicide count by sex
sns.barplot(data=suicide_df_v2, x='sex', y='suicides_no', hue='age', ax=ax1
,hue_order=['5-14 years','15-24 years','25-34 years','35-54 years','55-74 years','75+ years']) ;
ax1.set_title('Distribution of Annual Suicide Count by Sex', size=15, fontweight='bold') ;
# Plot 5:
sns.barplot(data=suicide_df_v2, x='sex', y='suicides/100k pop', hue='age', ax=ax2
,hue_order=['5-14 years','15-24 years','25-34 years','35-54 years','55-74 years','75+ years']) ;
ax2.set_title('Distribution of Annual Suicide Count by Sex (per 100,000 people)', size=15, fontweight='bold') ;
# +
# Make new variable 'continent_new' that identifies the continent for each country where a suicide occurred:
country = suicide_df_v2['country'].unique()
new_val = ['Europe','Central America','South America','Asia','Central America'
,'Australia','Europe','Asia','Central America','Asia'
,'Central America','Europe','Europe','Central America'
,'Europe','South America','Europe','Africa'
,'North America','South America','South America','Central America','Europe','Central America'
,'Asia','Europe','Europe','Central America','South America'
,'Central America','Europe','Oceania','Europe','Europe','Asia'
,'Europe','Europe','Central America','Central America','South America','Europe'
,'Europe','Europe','Asia','Europe','Central America','Asia'
,'Asia','Oceania','Asia','Asia','Europe'
,'Europe','Europe','Asia','Asia','Europe'
,'Africa','North America','Asia','Europe','Europe'
,'Oceania','Central America','Europe','Asia','Central America','South America'
,'Asia','Europe','Europe','Central America','Asia'
,'Asia','Europe','Europe'
,'Central America','Central America'
,'Central America','Europe','Europe'
,'Africa','Asia','Europe','Europe','Africa'
,'Europe','Asia','South America','Europe','Europe'
,'Asia','Central America','Asia','Asia'
,'Europe','Asia','Europe'
,'North America','South America','Asia']
new_var = []
for i in range(len(country)) :
n = len(suicide_df_v2[suicide_df_v2['country']==country[i]])
for j in range(n) :
new_var.append(new_val[i])
suicide_df_v2['continent_new'] = new_var
# +
# Count how many country in each continent recorded in the dataset:
continent = list(suicide_df_v2['continent_new'].unique())
new_val = pd.Series(new_val)
count = []
print('Count of country in each continent recorded in the dataset:')
for i in continent :
n = len(new_val[new_val==i])
count.append(n)
print(i,':',n)
# +
# Check the effect of variable 'generation'
matplotlib.rcParams['figure.figsize'] = [16,5]
gs = matplotlib.gridspec.GridSpec(1,3, width_ratios=[60,8,6])
ax1 = plt.subplot(gs[0,0])
# Plot 6 - Total suicide counts per country:
sns.barplot(data=suicide_df_v2[suicide_df_v2['continent_new'].isin(['North America',
'Europe',
'South America',
'Asia',
'Australia',
'Central America',
'Oceania',
'Africa'])], x='continent_new', y='suicides_no', ax=ax1
,hue_order=['G.I. Generation','Silent','Boomers','Generation X','Millennials','Generation Z'], hue='generation',) ;
ax1.set_title('Total Suicide Counts per Country', size=15, fontweight='bold') ;
# -
# ## Summary and Future Directions:
#
# On initial investigation, there was a range in annual suicides per country between 0 to approximately 27,000 deaths. It is unclear if data are missing and it is unlikely that no suicides occurred in any given country, given other international epidemiology studies citing the mental health phenomenon. Thus, I would be cautious about drawing conclusions with regard to every country's suicide rates. However, there was a large sample of data reported for the U.S. so I proceeded to do exploratory data analysis.
#
# This dataset included 101 unique countries, 32 different years, 2 sex identifiers, 6 age groups, and 6 generational cohort labels. The highest annual suicide counts occurred in the U.S., Russia, and Japan. However, since 1985, the annual death totals in Russia have decreased and stayed relatively similar for Japan; in the U.S., the annual suicide deaths has increased continuously from 1985-2016. Other countries saw a decrease in suicide deaths between 1985-2016, except for South Korea, which had an increase in suicide deaths particularly increasing after the year 2000.
#
# Death from suicide was higher in males than females, which conforms with epidemiological studies which suggest that while women might attempt suicide at higher rates, males complete suicide at higher rates. The highest rates of suicide (adjusted for share of population) were the 75+ age group in both males and females, while the highest total annual deaths occurred in the 35-54 year old cohorts in both males and females.
#
# Finally, I compared annual suicide death counts across different continents (by grouping the countries respectively) and observed higher annual suicide deaths in North America compared to other continents. However, it is unclear how accurate and valid data collection were across all listed countries and continents. The "Baby Boomer" generation showed the highest death counts from suicide across all continents, and this seems to relate to previous findings that the 35-54 y.o and 75+ age groups had the highest annual suicide counts.
#
# These are the initial interpretations of the data, however future data analysis is warranted to better understand other socio-economic factors that may relate to suicide rates in various countries.
#
|
Assignment_1_exploratory_data_analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # 2018.11.01: Network inference from time series of cartegorical variables
# ## eps = 1 if A, -1 otherwise
import sys,os
import numpy as np
from scipy import linalg
from sklearn.preprocessing import OneHotEncoder
import matplotlib.pyplot as plt
# %matplotlib inline
# +
# setting parameter:
np.random.seed(1)
n = 10 # number of positions
m = 5 # number of values at each position
l = 4*((n*m)**2) # number of samples
g = 1.
# +
def itab(n,m):
i1 = np.zeros(n)
i2 = np.zeros(n)
for i in range(n):
i1[i] = i*m
i2[i] = (i+1)*m
return i1.astype(int),i2.astype(int)
i1tab,i2tab = itab(n,m)
# -
# generate coupling matrix w0:
def generate_coupling(n,m,g):
nm = n*m
w = np.random.normal(0.0,g/np.sqrt(nm),size=(nm,nm))
for i in range(n):
i1,i2 = i1tab[i],i2tab[i]
w[i1:i2,:] -= w[i1:i2,:].mean(axis=0)
for i in range(n):
i1,i2 = i1tab[i],i2tab[i]
w[:,i1:i2] -= w[:,i1:i2].mean(axis=1)[:,np.newaxis]
return w
w0 = generate_coupling(n,m,g)
# 2018.10.27: generate time series by MCMC
def generate_sequences_MCMC(w,n,m,l):
#print(i1tab,i2tab)
# initial s (categorical variables)
s_ini = np.random.randint(0,m,size=(l,n)) # integer values
#print(s_ini)
# onehot encoder
enc = OneHotEncoder(n_values=m)
s = enc.fit_transform(s_ini).toarray()
#print(s)
ntrial = 100
for t in range(l-1):
h = np.sum(s[t,:]*w[:,:],axis=1)
for i in range(n):
i1,i2 = i1tab[i],i2tab[i]
k = np.random.randint(0,m)
for itrial in range(ntrial):
k2 = np.random.randint(0,m)
while k2 == k:
k2 = np.random.randint(0,m)
if np.exp(h[i1+k2]- h[i1+k]) > np.random.rand():
k = k2
s[t+1,i1:i2] = 0.
s[t+1,i1+k] = 1.
return s
s = generate_sequences_MCMC(w0,n,m,l)
# +
#print(s[:5])
# -
# recover s0 from s
s0 = np.argmax(s.reshape(-1,m),axis=1).reshape(-1,n)
def eps_ab_func(s0,m):
l,n = s0.shape
eps = np.zeros((n,l-1,m,m))
eps[:,:,:] = -1.
for i in range(n):
for t in range(l-1):
#eps[i,t,:,int(s0[t+1,i])] = -1.
eps[i,t,int(s0[t+1,i]),:] = 1.
return eps
eps_ab_all = eps_ab_func(s0,m)
# +
l = s.shape[0]
s_av = np.mean(s[:-1],axis=0)
ds = s[:-1] - s_av
c = np.cov(ds,rowvar=False,bias=True)
#print(c)
c_inv = linalg.pinv(c,rcond=1e-15)
#print(c_inv)
nm = n*m
nloop = 5
wini = np.random.normal(0.0,g/np.sqrt(nm),size=(nm,nm))
w_infer = np.zeros((nm,nm))
for i in range(n):
i1,i2 = i1tab[i],i2tab[i]
w_true = w0[i1:i2,:]
w = wini[i1:i2,:].copy()
#h = s[1:,i1:i2].copy()
for iloop in range(nloop):
h = np.dot(s[:-1],w.T)
for ia in range(m):
for t in range(l-1):
if s[t+1,i1+ia] == 1.:
ha = 0.
for ib in range(m):
if ib != ia:
hab = (h[t,ia] - h[t,ib])
if hab != 0: ha += hab/np.tanh(hab/2.)
h[t,ia] = ha/float(m)
h_av = h.mean(axis=0)
dh = h - h_av
dhds = dh[:,:,np.newaxis]*ds[:,np.newaxis,:]
dhds_av = dhds.mean(axis=0)
w = np.dot(dhds_av,c_inv)
mse = ((w_true - w)**2).mean()
slope = (w_true*w).sum()/(w_true**2).sum()
print(i,iloop,mse,slope)
w_infer[i1:i2,:] = w
# -
plt.scatter(w0,w_infer)
plt.plot([-0.3,0.3],[-0.3,0.3],'r--')
|
old_versions/ref_initial/1main_time_series-v8.1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction
#
# <font color = green> <strong>MUST READ:</strong></font> This Notebook tutorial will allow you to setup with Azure account for Azure connection using TNGTLS. By using this Notebook, its possible to update account credentials to Azure cli, Register the Signer and device.
#
# Before running this Notebook,
# 1. <font><strong>docs/Azure_iot_hub_details.csv</strong></font> file should have updated with azure iot hub name.
# 2. Make sure CryptoAuth Trust Platform is having factory reset program
# ### 1.1.Tutorial Prerequisites:
#
#
# The following code is runs on python 3.x environment. This step of the tutorial will attempt to load all the necessary modules and their dependencies on your machine. If the above modules are already installed you can safely step over the next Tutorial step.
#
# > <font color = green>pip install -U module_name</font>
# <font color = orange> <strong>Note</strong></font>
# 1. Installation time for prerequisites depends upon system and network speed.
# 2. Installing prerequisites for the first time takes more time and watch the kernel status for progress. Following image helps to locate the Kernel status,
# <center><img src="../../../../assets/notebook/img/kerner_status.png" alt="**Check Kernel Status**" /></center>
#
#
# 3. Installing prerequisites gives the following error and it can be safely ignored. Functionality remains unaffected.
# - <font color = orange> azure-cli 2.0.76 has requirement colorama~=0.4.1, but you'll have colorama 0.3.9 which is incompatible.</font>
# - <font color = orange> azure-cli 2.0.76 has requirement pytz==2019.1, but you'll have pytz 2019.3 which is incompatible. </font>
# - <font color = orange> Extension 'azure-cli-iot-ext' is already installed. </font>
# +
import sys, os
home_path = os.path.dirname(os.path.dirname(os.path.dirname(os.path.dirname(os.path.realpath(os.getcwd())))))
module_path = os.path.join(home_path, 'assets', 'python')
if not module_path in sys.path:
sys.path.append(module_path)
from requirements_helper import requirements_installer
obj = requirements_installer(os.path.join(home_path, 'assets', 'requirements.txt'))
# -
# !az extension add --name azure-cli-iot-ext
# ### Load the necessary modules and helper functions
# +
from helper_azure import *
from ipywidgets import *
from IPython.display import display
from tkinter import Tk, filedialog
from cryptoauthlib import *
from cryptoauthlib.iface import *
import binascii
from trustplatform import *
print("Importing modules - Successful")
#TFLXTLS device I2C address 0x6C; #TNGTLS device I2C address 0x6A;
common_helper.connect_to_secure_element('ATECC608', ATCAKitType.ATCA_KIT_I2C_IFACE, 0x6A)
print("Device initialization - Successful")
# Request the Serial Number
serial_number = bytearray(9)
assert atcab_read_serial_number(serial_number) == Status.ATCA_SUCCESS
device_reg_id = binascii.hexlify(serial_number).decode("utf-8").upper()
print ("Device ID: {}".format(device_reg_id))
# Verify Certificate support for Azure connection
unsupported_otp = "Rsuy5YJh"
otpcode = bytearray(32)
ATCA_ZONE_OTP = 1
assert atcab_read_zone(ATCA_ZONE_OTP, 0, 0, 0, otpcode, 32) == Status.ATCA_SUCCESS
assert unsupported_otp != otpcode[:8].decode('utf-8'), "TNGTLS connected doesnt support Azure connection! Try with another TNGTLS device"
# -
# The following steps assist to perform
# 1. Azure account login
# 2. Registering the Device
#
# #### Azure account login (1. Account Login)
# This helps to update user Azure credentials to Azure-cli. Azure-cli will be used for any interactions with Azure cloud from the PC.This can be skipped if the azure login update to azure cli is already done.
#
# #### Register the Device (2. Register device)
# This will register the device ID of the TNGTLS to Azure iot hub account. For registering TNGTLS device, its required to provide Manifest file and associated verification certificate. If Manifest file and verification certificates are not available, run the Trust&GO resource generation to generate. Load Manifest and Validation certs before clicking the 'Register device' button. This should register the devices in Manifest file to Azure cloud.
# ### 1. Azure account login
# This step helps to initialise the Azure cli tools with azure account credentials.
# +
def execute_azure_login(b):
Azure_login.button_style = azure_account_login()
Azure_login = widgets.Button(description = "Azure login")
Azure_login.on_click(execute_azure_login)
# -
# ### 3. Register Device ID from Manifest
# This step of the tutorial registers the device ID to the Azure IOT Hub.
# +
def execute_device_register(b):
reg_device.button_style = ''
if not manifest_file.data or not validation_cert.data:
print('Step2a & Step2b should be executed first before proceeding to this step\r\n')
return None
register_device(json.loads(manifest_file.data[0]), validation_cert.data[0])
manifest_file = FileUpload(description='Step2a. Load Manifest JSON File', accept='.json', layout=widgets.Layout(width='auto'), multiple=False)
validation_cert = FileUpload(description='Step2b. Load Validation CERT File', accept='.crt', layout=widgets.Layout(width='auto'), multiple=False)
reg_device = widgets.Button(description = "Step2. Register device", layout=widgets.Layout(width='auto'))
reg_device.on_click(execute_device_register)
display(widgets.HBox((Azure_login, widgets.VBox((manifest_file, validation_cert, reg_device)))))
|
TrustnGO/05_cloud_connect/notebook/azure/TNGTLS_azure_connect.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# # Machine Learning artifacts management
# This notebook contains steps and code to demonstrate how to manage and clean up Watson Machine Learning instance. This notebook contains steps and code to work with [ibm-watson-machine-learning](https://pypi.python.org/pypi/ibm-watson-machine-learning) library available in PyPI repository. This notebook introduces commands for listing artifacts, getting artifacts details and deleting them.
#
# Some familiarity with Python is helpful. This notebook uses Python 3.
# ## Learning goals
#
# The learning goals of this notebook are:
#
# - List Watson Machine Learning artifacts.
# - Get artifacts details.
# - Delete artifacts.
#
#
# ## Contents
#
# This notebook contains the following parts:
#
# 1. [Setup](#setup)
# 2. [Manage pipelines](#pipelines)
# 3. [Manage model definitions](#model_definitions)
# 4. [Manage models](#models)
# 5. [Manage functions](#functions)
# 6. [Manage experiments](#experiments)
# 7. [Manage trainings](#trainings)
# 8. [Manage deployments](#deployments)
# 9. [Summary and next steps](#summary)
# <a id="setup"></a>
# ## 1. Set up the environment
#
# Before you use the sample code in this notebook, you must perform the following setup tasks:
#
# - Create a <a href="https://console.ng.bluemix.net/catalog/services/ibm-watson-machine-learning/" target="_blank" rel="noopener no referrer">Watson Machine Learning (WML) Service</a> instance (a free plan is offered and information about how to create the instance can be found <a href="https://dataplatform.ibm.com/docs/content/analyze-data/wml-setup.html" target="_blank" rel="noopener no referrer">here</a>).
# ### Connection to WML
#
# Authenticate the Watson Machine Learning service on IBM Cloud. You need to provide platform `api_key` and instance `location`.
#
# You can use [IBM Cloud CLI](https://cloud.ibm.com/docs/cli/index.html) to retrieve platform API Key and instance location.
#
# API Key can be generated in the following way:
# ```
# ibmcloud login
# ibmcloud iam api-key-create API_KEY_NAME
# ```
#
# In result, get the value of `api_key` from the output.
#
#
# Location of your WML instance can be retrieved in the following way:
# ```
# ibmcloud login --apikey API_KEY -a https://cloud.ibm.com
# ibmcloud resource service-instance WML_INSTANCE_NAME
# ```
#
# In result, get the value of `location` from the output.
# **Tip**: Your `Cloud API key` can be generated by going to the [**Users** section of the Cloud console](https://cloud.ibm.com/iam#/users). From that page, click your name, scroll down to the **API Keys** section, and click **Create an IBM Cloud API key**. Give your key a name and click **Create**, then copy the created key and paste it below.
#
# You can also get service specific apikey by going to the [**Service IDs** section of the Cloud Console](https://cloud.ibm.com/iam/serviceids). From that page, click **Create**, then copy the created key and paste it below.
#
# **Action**: Enter your `api_key` and `location` in the following cell.
api_key = 'PASTE YOUR PLATFORM API KEY HERE'
location = 'PASTE YOUR INSTANCE LOCATION HERE'
wml_credentials = {
"apikey": api_key,
"url": 'https://' + location + '.ml.cloud.ibm.com'
}
# ### Install and import the `ibm-watson-machine-learning` package
# **Note:** `ibm-watson-machine-learning` documentation can be found <a href="http://ibm-wml-api-pyclient.mybluemix.net/" target="_blank" rel="noopener no referrer">here</a>.
# !pip install -U ibm-watson-machine-learning
# +
from ibm_watson_machine_learning import APIClient
client = APIClient(wml_credentials)
# -
# ### Working with spaces
#
# First of all, you need to create a space that will be used for your work with Watson Machine Learning. If you do not have space already created, you can use [Deployment Spaces Dashboard](https://dataplatform.cloud.ibm.com/ml-runtime/spaces?context=cpdaas) to create one.
#
# - Click New Deployment Space
# - Create an empty space
# - Select Cloud Object Storage
# - Select Watson Machine Learning instance and press Create
# - Copy `space_id` and paste it below
#
# **Tip**: You can also use SDK to prepare the space for your work. More information [here](). ##TODO
#
# **Action**: Assign space ID below
space_id = 'PASTE YOUR SPACE ID HERE'
# You can use `list` method to print all existing spaces.
client.spaces.list(limit=10)
# To be able to interact with all resources available in Watson Machine Learning, you need to set **space** which you will be using.
client.set.default_space(space_id)
# <a id="pipelines"></a>
# ## 2. Manage pipelines
# List existing pipelines. If you want to list only part of pipelines use `client.pipelines.list(limit=n_pipelines)`.
client.pipelines.list(limit=10)
# Get pipelines details. If you want to get only part of pipelines details use `client.pipelines.get_details(limit=n_pipelines)`.
#
# You can get each pipeline details by calling `client.pipelines.get_details()` and providing pipeline id from listed pipelines.
pipelines_details = client.pipelines.get_details(limit=10)
print(pipelines_details)
# Delete all pipelines. You can delete one pipeline by calling `client.pipelines.delete()` and providing pipeline id from listed pipelines.
for pipeline in pipelines_details['resources']:
client.pipelines.delete(pipeline['metadata']['id'])
# <a id="model_definitions"></a>
# ## 3. Manage model definitions
# List existing model definitions. If you want to list only part of model definitions use `client.model_definitions.list(limit=n_model_definitions)`.
client.model_definitions.list(limit=10)
# Get model definiton details by copying model definition uid from above cell and running `client.model_definitions.get_details(model_definition_guid)`.
model_definition_guid = "PUT_YOUR_MODEL_DEFINITION_GUID"
model_definitions_details = client.model_definitions.get_details(model_definition_guid)
print(model_definitions_details)
# Delete model definitions by calling `client.model_definitions.delete(model_definition_guid)`.
client.model_definitions.delete(model_definition_guid)
# <a id="models"></a>
# ## 4. Manage models
# List existing models. If you want to list only part of models use `client.repository.list_models(limit=n_models)`.
client.repository.list_models(limit=10)
# Get model details by copying model uid from above cell and running `client.repository.get_details(model_guid)`.
model_guid = "PUT_YOUR_MODEL_GUID"
model_details = client.repository.get_details(model_guid)
print(model_details)
# Delete model from repository by calling `client.repository.delete(model_guid)`.
client.repository.delete(model_guid)
# <a id="functions"></a>
# ## 5. Manage functions
# List existing functions. If you want to list only part of functions use `client.repository.list_functions(limit=n_functions)`.
client.repository.list_functions(limit=10)
# Get function details by copying function uid from above cell and running `client.repository.get_details(function_guid)`.
function_guid = "PUT_YOUR_FUNCTION_GUID"
function_details = client.repository.get_details(function_guid)
print(function_details)
# Delete function from repository by calling `client.repository.delete(function_guid)`.
client.repository.delete(function_guid)
# <a id="experiments"></a>
# ## 6. Manage experiments
# List existing experiments. If you want to list only part of experiments use `client.pipelines.list(limit=n_experiments)`.
client.experiments.list(limit=10)
# Get experiments details. If you want to get only part of experiments details use `client.experiments.get_details(limit=n_experiments)`.
#
#
# You can get each experiment details by calling `client.experiments.get_details()` and providing experiment id from listed experiments.
experiments_details = client.experiments.get_details()
print(experiments_details)
# Delete all experiments. You can delete one experiment by calling `client.experiments.delete()` and providing experiment id from listed experiments.
for experiment in experiments_details['resources']:
client.experiments.delete(experiment['metadata']['id'])
# <a id="trainings"></a>
# ## 7. Manage trainings
# List existing trainings. If you want to list only part of trainings use `client.training.list(limit=n_trainings)`.
client.training.list(limit=10)
# Get trainings details. If you want to get only part of trainings details use `client.training.get_details(limit=n_trainings)`.
#
# You can get each training details by calling `client.training.get_details()` and providing training id from listed trainings.
trainings_details = client.training.get_details(limit=10)
print(trainings_details)
# Delete all trainings. You can delete one training by calling `client.training.cancel()` and providing training id from listed trainings.
#
# **Note** The `client.training.cancel()` method has `hard_delete` parameter. Please change it to:
#
# - True - to delete the completed or canceled training runs.
# - False - to cancel the currently running training run.
#
# Default value is `False`.
for training in trainings_details['resources']:
client.training.cancel(training['metadata']['id'])
# <a id="deployments"></a>
# ## 8. Manage deployments
# List existing deployments. If you want to list only part of deployments use `client.deployments.list(limit=n_deployments)`.
# + pycharm={"name": "#%%\n"}
client.deployments.list(limit=10)
# -
# Get deployments details. If you want to get only part of deployments details use `client.deployments.get_details(limit=n_deployments)`.
#
# You can get each deployment details by calling `client.deployments.get_details()` and providing deployment id from listed deployments.
# + pycharm={"name": "#%%\n"}
deployments_details = client.deployments.get_details()
print(deployments_details)
# -
# Delete all deployments. You can delete one deployment by calling `client.deployments.delete()` and providing deployment id from listed deployments.
# + pycharm={"name": "#%%\n"}
for deployment in deployments_details['resources']:
client.deployments.delete(deployment['metadata']['id'])
# -
# <a id="summary"></a>
# ## 9. Summary and next steps
# You successfully completed this notebook! You learned how to use ibm-watson-machine-learning client for Watson Machine Learning instance management and clean up. Check out our _[Online Documentation](https://console.ng.bluemix.net/docs/services/PredictiveModeling/index.html?pos=2)_ for more samples, tutorials, documentation, how-tos, and blog posts.
# ### Authors
#
# **<NAME>**, Software Engineer at IBM.
# Copyright © 2020 IBM. This notebook and its source code are released under the terms of the MIT License.
|
notebooks/python_sdk/instance-management/Machine Learning artifacts management.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Pyspark (local)
# language: python
# name: pyspark_local
# ---
# ## Beregning av statistiske størrelser med pyspark
# ### I denne noten viser vi hvordan vi kan beregne statistiske størrelser i pyspark. Utgangspunkt for beregninger er parquet datasett som leses inn fra dapla.
# #### Innhenter verktøy fra bibliotek
# Import-stegene henter inn bibliotek med kode og funksjoner utviklet ekstern.
from datetime import datetime
from pyspark.sql import SparkSession
from pyspark.sql.types import *
from pyspark.sql import SQLContext
import pyspark.sql.functions as F
import numpy as np
from pyspark.sql import Row
# #### Vis tilgjengelige datasett
# Kjører metoden read.path for å få oversikt over hvilke parquet datasett som er tilrettelagt i tilknytning til veilderens lagringsområde i sky. <br> Oversikten blir lest inn i egen dataframe - df_datsets.<br>
# Aktuelt lagringsområde blir lagt inn som parameter (string objekt som vi definerer selv) INNDATA_PATH.
INNDATA_PATH = '/felles/veiledning/datasett/*'
df_datasets = spark.read.path(INNDATA_PATH)
# df_datsets skrives ut i output vindu.
df_datasets.show(100, False)
# Parquet datasettene df_sammensatt_pyspark og df_sammensatt_python som vises når man kjører paragrafen over, skal i utgangspunktet være identiske.<br>
# De er produsert i henholdsvis pyspark og python. I vårt eksempel tar vi utgangspunkt i df_sammensatt_pyspark.
# #### Leser inn parquet datasett
# Leser inn parquet datasett df_sammensatt_pyspark, selekterer de variable vi skal bruke og etablerer dataframen df_areal_bnp_og_innbyggerantall
df_areal_bnp_og_innbyggerantall = spark.read.path('/felles/veiledning/datasett/df_sammensatt_pyspark').select('Land', 'Areal', 'BNP', 'Innbyggerantall')
df_areal_bnp_og_innbyggerantall.show()
# #### Beregninger
# Vi ønsker som utgangspunkt å beregne størrelsene count, mean, std, min, max og median for alle variable i df_areal_bnp_og_innbyggerantall. Ved å kjøre pyspark metoden describe på datasettet får vi ut størrelsene count, mean, std, min og max. Dvs at pyspark describe har ikke støtte for å beregne medianen.
# ### Beregninger - eksempel 1
# Vi ønsker som utgangspunkt å beregne størrelsene count, mean, std, min, max og median for alle variable i df_areal_bnp_og_innbyggerantall. Ved å kjøre metoden summary på datasettet får vi ut størrelsene count, mean, std, min, max, 25% percentile, 50% percentile og 75% percentile. Vi betrakter 50% percentile som medianen. Dvs at vi får ut alle de størrelser vi ønsker å beregne i dette eksempelet kun ved å describe metoden i pandas.
# ##### 1. Kjører ut statistiske størrelser
# Kjører ut statistiske størrelser fra dataframe df_areal_bnp_og_innbyggerantall med metoden summary. Resultat fra metode blir lagt i dataframen df_stat.
df_stat = df_areal_bnp_og_innbyggerantall.summary()
df_stat.show()
# ##### 2. Transformerer df_stat
# Transformerer df_stat (med bl.a. pivot funksjonen) slik at rader i df_stat blir kolonner i dataframe df_stat_piv. Formål er å gjøre output mer oversiktlig ved at alle størrelser på en gitt variabel blir liggende i samme rad. Dette kan være hensiktsmessig i de tilfeller antall "statistikkvaiable" er høyt (dvs langt høyere enn de fire som inngår i eksempel).
var_lst = list(df_stat.columns)
var_lst.remove('summary')
i = 1
for col in var_lst:
df_tmp = df_stat.groupBy().pivot('summary').agg(F.sum(col)).withColumn('variabel',F.lit(col))
if i == 1:
df_stat_piv = df_tmp
else:
df_stat_piv = df_stat_piv.unionByName(df_tmp)
i = i + 1
df_stat_piv.show(100, False)
# ### Beregninger eksempel 2
# I eksempel 1 fikk vi beregnet de størrelsene vi var ute etter kun ved å bruke metoden summary. Hvis vi ønsker å beregne størrelser som ikke er støttet av summary metoden kan vi bruke andre funksjoner og/eller bibliotek.
#
# I eksempel 2 ønsker vi å beregne skewness gjennom å bruke en funksjon fra pyspark.sql modulen. Videre ønsker vi i samme steg å inkludere skewness "sammen" med størrelsene vi får ved å kjøre summary metoden. Resultat skal transformeres på tilsvarende måte som i eksempel 1.
#
# Vi ønsker å etablere datasett hvor skeness er satt sammen med størrelsene vi får fra summary. For å få til dette må vi gjennomføre flere steg.
# ##### 1. Kjører ut statistiske størrelser med describe
# Kjører ut statistiske størrelser fra dataframe df_areal_bnp_og_innbyggerantall med metoden describe. Resultat fra metode blir lagt i dataframen df_stat.
df_stat_adv = df_areal_bnp_og_innbyggerantall.summary()
df_stat_adv.show()
# Printer schemaet til df_stat_adv
df_stat_adv.printSchema()
# ##### 2. Beregner skewness
# Beregner størrelsen skewness for hver kolonne, med unntak av summary kolonne, i df_areal_bnp_og_innbyggerantall med funksjonen skewness. Beregningene gjøres i en for loop. Resultat fra funksjon blir returnert inn i egne dataframes (ett pr variabel fra dataframen df_stat_adv). Datasettene kobles så sammen slik at vi får ett datasett der skewness for alle variable er samlet i en rad.
var_lst = list(df_stat_adv.columns)
var_lst.remove('summary')
i = 1
for col in var_lst:
df_tmp = df_areal_bnp_og_innbyggerantall.agg(F.skewness(col).alias(col))
if i == 1:
df_stat_ske = df_tmp
else:
df_stat_ske = df_stat_ske.join(df_tmp)
i = i + 1
df_stat_ske = df_stat_ske.withColumn("summary", F.lit('skewness'))
df_stat_ske.show()
# ##### 3. Slår sammen df_stat og df_stat_ske
# Slår sammen df_stat og df_stat_ske. Viktig at man her bruker unionByName fordi rekkefølgen på variablene ikke er identisk.
df_stat_sammensatt = df_stat_adv.unionByName(df_stat_ske)
df_stat_sammensatt.show()
# ##### 4. Transformerer df_stat_sammensatt
# Transformerer df_stat (med bl.a. pivot funksjonen) slik at rader i df_stat blir kolonner i dataframe df_stat_piv. Formål er å gjøre output mer oversiktlig ved at alle størrelser på en gitt variabel blir liggende i samme rad. Dette kan være hensiktsmessig i de tilfeller antall "statistikkvariable" er høyt (dvs langt høyere enn de fire som inngår i eksempel).
# +
var_lst = list(df_stat_sammensatt.columns)
var_lst.remove('summary')
i = 1
for col in var_lst:
df_tmp = df_stat_sammensatt.groupBy().pivot('summary').agg(F.sum(col)).withColumn('variabel',F.lit(col))
if i == 1:
df_stat_sammensatt_piv = df_tmp
else:
df_stat_sammensatt_piv = df_stat_sammensatt_piv.unionByName(df_tmp)
i = i + 1
# -
df_stat_sammensatt_piv.show(100, False)
|
utforske/deskriptiv_statistikk_pyspark.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Kulturnav - platser
# +
import urllib3, json
import pandas as pd
http = urllib3.PoolManager()
pd.set_option("display.max.columns", None)
urlbase = "https://kulturnav.org/api/search/entityType:Place?start="
dftot = pd.DataFrame()
for i in range(1,10000,20):
url = urlbase + str(i)
if ((i-1) % 1000) == 0:
print('*', end='', flush=True)
r = http.request('GET', url)
try:
data = json.loads(r.data)
dftot = dftot.append(pd.DataFrame(data),sort=False)
except:
print("\tError in i=",i)
# -
dftot.info()
dftot
|
Kulturnav - platser.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# Inspiration from https://github.com/cran/RadOnc/blob/master/R/read.DVH.R
#
# And http://stackoverflow.com/questions/3914454/python-how-to-loop-through-blocks-of-lines
# %load_ext watermark
# %watermark -a '<NAME>' -u -d -v -p numpy,pandas,matplotlib
import numpy as np
import itertools
import json
import re
# +
file = 'case1_AAA.txt'
file_header = open(file, 'r').readlines()[:15]
file_body = open(file, 'r').readlines()[16:] # first 16 lines are header, for some reason was not opening header
plan = {} # to contain all file plan info
# parse the header here
for line in file_header:
if line.replace(u'\ufeff', '').startswith('Patient Name'):
plan['Patient Name'] = line.replace(u'\ufeff', '').split(':')[1].rstrip()
elif line.startswith('Patient ID'):
plan['Patient ID'] = line.split(':')[1].rstrip()
elif line.startswith('Plan:'):
plan['Plan'] = line.split(':')[1].rstrip()
elif line.startswith('Prescribed dose [Gy]'):
plan['Prescribed dose [Gy]'] = line.split(':')[1].rstrip()
else:
pass
#############################
## get all structures
Structures = [] # create list
print(len(file_body))
for i, line in enumerate(file_body):
matchObj = re.match( r'^Structure: (.*?$)', line, re.M|re.X)
if matchObj:
Structures.append(matchObj.group(1))
plan['Structures'] = Structures
#############################
plan
# -
i = 0
for key, Structure_group in itertools.groupby(file_body, lambda line: line.startswith('Structure:')): # group on Structure
print(i)
for key, sub_group in itertools.groupby(Structure_group, lambda line: line=='\n'): # Group on empty lines
if not key:
for i, line in enumerate(sub_group):
if line.startswith('Relative dose'):
data = [] # create a list so can use the append . np arrays must be init with length
for line in sub_group:
if not line.startswith('Relative dose') and len(line.split()) == 3 : # 3 cols of data
aa = [float(i) for i in line.split()]
#data = np.asarray(data)
data.append(aa)
else:
info = {};
for line in sub_group:
if len(line.split()) == 2 :
field,value=line.split(':')
if isinstance(value, str): # if a string
value=value.strip()
else:
value = float(value)
info[field]=value
i = i + 1
# +
from decimal import *
line = ' 0 0 100'
print(len(line.split()))
bb = [float(i) for i in line.split()]
bb
#results =[float(i) for i in bb]
#results
# -
|
pyparsing/Import eclipse dvh text files 3-10-2016.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.0 64-bit
# name: python390jvsc74a57bd063fd5069d213b44bf678585dea6b12cceca9941eaf7f819626cde1f2670de90d
# ---
# +
import pandas as pd
from IPython.display import display
red_wine = pd.read_csv('data/red-wine.csv')
# Create training and validation splits - Training data 70% and Validation data 30%
df_train = red_wine.sample(frac=0.7, random_state=0)
df_valid = red_wine.drop(df_train.index)
display(df_train.head(4))
# Scale to [0, 1] - This scales each feature to lie in the interval [0,1] neural networks tend to work best with this.
max_ = df_train.max(axis=0)
min_ = df_train.min(axis=0)
df_train = (df_train - min_) / (max_ - min_)
df_valid = (df_valid - min_) / (max_ - min_)
# Split features and target - The target feature is the "quality"
X_train = df_train.drop('quality', axis=1)
X_valid = df_valid.drop('quality', axis=1)
y_train = df_train['quality']
y_valid = df_valid['quality']
# -
print(X_train.shape) # Print the shape of the training dataset (i.e. no. of rows and columns)
# +
# Import keras
from tensorflow import keras
from tensorflow.keras import layers
# Define the Sequential model: 3 layers with ReLU activation and 512 neurons each.
model = keras.Sequential([
layers.Dense(512, activation='relu', input_shape=[11]),
layers.Dense(512, activation='relu'),
layers.Dense(512, activation='relu'),
layers.Dense(1),
])
# +
# Add the loss function and the optimiser to the model (compile it in)
model.compile(
optimizer='adam',
loss='mae',
)
# +
# Fit the model: specify validation data, set batch_size as 256 rows (So this feeds the optimiser 256 rows of training data at a time), set the number of epochs as 10 (Number of times the model goes all the way through the dataset)
# Defined as a history object in order to keep a record of the loss produced
history = model.fit(
X_train, y_train,
validation_data=(X_valid, y_valid),
batch_size=256,
epochs=10,
)
# Keras updates you on the loss as the model trains...
# +
# This converts the training history into a Pandas dataframe:
history_df = pd.DataFrame(history.history)
# Uses Pandas native plot method to plot the changing loss
history_df['loss'].plot();
# As the epochs go by the loss levels off (This happens when the model has learned all that it can therefore no more epochs required)
# -
|
Kaggle_ML_Tutorials/IntoDeepLearning/SGDtutorial.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import fitz
import os
import pickle
import math
from bs4 import BeautifulSoup
import numpy as np
# +
index_pdf_df = pd.read_csv("G:\\ESA_downloads\\Demo_Alignment_Sheets\\Index_2_PDF.csv",
encoding= 'unicode_escape')
index_pdf_df = index_pdf_df.rename(columns={"Data ID": "DataID"})
index_pdf_df.head()
pdfs = []
for index, row in index_pdf_df.iterrows():
pdfs.append(str(row["DataID"]))
len(pdfs)
# -
#index_file = pd.read_csv(path + "Data_Files_v2\\book1.csv", index_col = 0, encoding= 'unicode_escape')
index_file = pd.read_csv("G:\ESA_downloads\ESA_bundled_with_VCs\ESA_website_ENG.csv",
encoding= 'unicode_escape')
print(len(index_file))
count = 0
not_needed_pdfs = []
for index, row in index_file.iterrows():
pdf = str(row["Data ID"])
if pdf in pdfs:
continue
else:
count = count +1
not_needed_pdfs.append(str(row["Data ID"]))
print(count)
len(set(not_needed_pdfs)
len(set(not_needed_pdfs))
set(not_needed_pdfs)
import glob
#### Reading the Index 1 file - Dataframe
path = "G:\\ESA_downloads\\Demo_Alignment_Sheets\\"
path_pdfs = path + "Data_Files_v2\\PDFs\\*.pdf"
count = 0
pdf_paths = []
for path_pdf in glob.glob(path_pdfs, recursive=True):
count = count + 1
if '444427' in path_pdf:
print(path_pdf)
444427 in pdfs
|
berdi/Section_06_Alignment_Sheets_index_merge/Sanity Checks.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/jonkrohn/ML-foundations/blob/master/notebooks/2-linear-algebra-ii.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="aTOLgsbN69-P"
# # Linear Algebra II: Matrix Operations
# + [markdown] colab_type="text" id="yqUB9FTRAxd-"
# This topic, *Linear Algebra II: Matrix Operations*, builds on the basics of linear algebra. It is essential because these intermediate-level manipulations of tensors lie at the heart of most machine learning approaches and are especially predominant in deep learning.
#
# Through the measured exposition of theory paired with interactive examples, you’ll develop an understanding of how linear algebra is used to solve for unknown values in high-dimensional spaces as well as to reduce the dimensionality of complex spaces. The content covered in this topic is itself foundational for several other topics in the *Machine Learning Foundations* series, especially *Probability & Information Theory* and *Optimization*.
# + [markdown] colab_type="text" id="d4tBvI88BheF"
# Over the course of studying this topic, you'll:
#
# * Develop a geometric intuition of what’s going on beneath the hood of machine learning algorithms, including those used for deep learning.
# * Be able to more intimately grasp the details of machine learning papers as well as all of the other subjects that underlie ML, including calculus, statistics, and optimization algorithms.
# * Reduce the dimensionalty of complex spaces down to their most informative elements with techniques such as eigendecomposition, singular value decomposition, and principal components analysis.
# + [markdown] colab_type="text" id="Z68nQ0ekCYhF"
# **Note that this Jupyter notebook is not intended to stand alone. It is the companion code to a lecture or to videos from <NAME>'s [Machine Learning Foundations](https://github.com/jonkrohn/ML-foundations) series, which offer detail on the following:**
#
# *Review of Matrix Properties*
#
# * Modern Linear Algebra Applications
# * Tensors, Vectors, and Norms
# * Matrix Multiplication
# * Matrix Inversion
# * Identity, Diagonal and Orthogonal Matrices
#
# *Segment 2: Eigendecomposition*
#
# * Eigenvectors
# * Eigenvalues
# * Matrix Determinants
# * Matrix Decomposition
# * Applications of Eigendecomposition
#
# *Segment 3: Matrix Operations for Machine Learning*
#
# * Singular Value Decomposition (SVD)
# * The Moore-Penrose Pseudoinverse
# * The Trace Operator
# * Principal Component Analysis (PCA): A Simple Machine Learning Algorithm
# * Resources for Further Study of Linear Algebra
# + [markdown] colab_type="text" id="ty8zFJj9j3Ui"
# ## Segment 1: Review of Matrix Properties
# + colab={} colab_type="code" id="ixvqsCeFj3Uj"
import numpy as np
import torch
# + [markdown] colab_type="text" id="E0LgmflFj3Um"
# ### Vector Transposition
# -
x = np.array([25, 2, 5])
x
x.shape
# + colab={} colab_type="code" id="hvLQEylJj3Un" outputId="201ba11e-2904-4084-e9d9-b865072e6e93"
x = np.array([[25, 2, 5]])
x
# -
x.shape
# + colab={} colab_type="code" id="D6XtfKZ8j3Ut" outputId="3ce3d51d-a5ca-4513-9296-d69e8d45e8cc"
x.T
# + colab={} colab_type="code" id="IN1_aSVQj3Uw" outputId="e37e7b5c-ed16-4955-faa0-e76ce1d7f112"
x.T.shape
# + colab={} colab_type="code" id="K-_nrgesj3U4" outputId="4917c288-7b5d-47b2-a878-e3da375f58b0"
x_p = torch.tensor([25, 2, 5])
x_p
# + colab={} colab_type="code" id="X0YuG9Pmj3U6" outputId="734a426f-6503-4298-845a-e936513e8066"
x_p.T
# + colab={} colab_type="code" id="S_bRLc5wj3U8" outputId="2530e985-5684-433d-c576-603916d2daec"
x_p.view(3, 1) # "view" because we're changing output but not the way x is stored in memory
# + [markdown] colab_type="text" id="llPpXVPHj3U_"
# **Return to slides here.**
# + [markdown] colab_type="text" id="43tlq2huj3U_"
# ## $L^2$ Norm
# + colab={} colab_type="code" id="wF1B2qL1j3VA" outputId="a24057fd-93a3-4513-b95c-9eaa8220045d"
x
# + colab={} colab_type="code" id="SW2iYHE8j3VC" outputId="26ebcfff-3220-4351-fa90-e2c21adeb6b3"
(25**2 + 2**2 + 5**2)**(1/2)
# + colab={} colab_type="code" id="BGiMFU0pj3VE" outputId="165d1e73-3514-4c3b-c955-24012c2b2b2c"
np.linalg.norm(x)
# + [markdown] colab_type="text" id="3YBk8ta2j3VI"
# So, if units in this 3-dimensional vector space are meters, then the vector $x$ has a length of 25.6m
# + colab={} colab_type="code" id="jjS4_w_Rj3VI"
# the following line of code will fail because torch.norm() requires input to be float not integer
# torch.norm(p)
# + colab={} colab_type="code" id="fgadi8SFj3VK" outputId="a8819539-9dbe-4bb5-dbde-943ef6c1d789"
torch.norm(torch.tensor([25, 2, 5.]))
# + [markdown] colab_type="text" id="qHti3Xslj3VM"
# **Return to slides here.**
# + [markdown] colab_type="text" id="4xDLTPutj3VN"
# ### Matrices
# + colab={} colab_type="code" id="Fep93KC7j3VN" outputId="daa76d80-2294-4994-c268-0a6274d183a1"
X = np.array([[25, 2], [5, 26], [3, 7]])
X
# + colab={} colab_type="code" id="S_yMqWdSj3VP" outputId="8b2f690e-a383-41b3-fe4b-c8fa8b6d4735"
X.shape
# + colab={} colab_type="code" id="RDkn2n92j3VX" outputId="0491c341-f2ab-42d9-ae41-306c55b292cb"
X_p = torch.tensor([[25, 2], [5, 26], [3, 7]])
X_p
# + colab={} colab_type="code" id="SzpKBCqKj3VY" outputId="3ea34673-a489-418e-a900-f8b33ce4f758"
X_p.shape
# + [markdown] colab_type="text" id="5jL7I-iWj3Vg"
# **Return to slides here.**
# + [markdown] colab_type="text" id="pC1rWTlVj3Vg"
# ### Matrix Transposition
# + colab={} colab_type="code" id="srh333wTj3Vg" outputId="faa97c11-28ce-4a47-f2fc-d4e80e30c8c0"
X
# + colab={} colab_type="code" id="e7siMBsRj3Vi" outputId="8d2b7052-32d3-4e18-a47d-71f7721099cf"
X.T
# + colab={} colab_type="code" id="YF3iqTkUj3Vk" outputId="0289f154-12bd-4cd7-e448-311260d662cf"
X_p.T
# + [markdown] colab_type="text" id="mXo5iD5Xj3Vm"
# **Return to slides here.**
# + [markdown] colab_type="text" id="loFYZ-pXj3Vm"
# ### Matrix Multiplication
# + [markdown] colab_type="text" id="BrHCDYrzj3Vm"
# Scalars are applied to each element of matrix:
# + colab={} colab_type="code" id="Yf3WIZ6Jj3Vn" outputId="d1771acb-2ace-4539-8ca6-3f1c474b7cb3"
X*3
# + colab={} colab_type="code" id="Pk-lY78Nj3Vp" outputId="3a677d4d-9921-42bf-c631-6e38fac78e33"
X*3+3
# + colab={} colab_type="code" id="g_sJ0NI8j3Vq" outputId="10874a31-e67d-47f7-cd9a-25e686206b37"
X_p*3
# + colab={} colab_type="code" id="jebPN-iPj3Vs" outputId="81de14ff-554f-444c-ec7f-eb2e4bfbdea1"
X_p*3+3
# + [markdown] colab_type="text" id="-a4o6abYj3Vu"
# Using the multiplication operator on two tensors of the same size in PyTorch (or Numpy or TensorFlow) applies element-wise operations. This is the **Hadamard product** (denoted by the $\odot$ operator, e.g., $A \odot B$) *not* **matrix multiplication**:
# + colab={} colab_type="code" id="JtRT2V0cj3Vu" outputId="dc363187-2e45-43b7-e308-098f90a68649"
A = np.array([[3, 4], [5, 6], [7, 8]])
A
# + colab={} colab_type="code" id="LLJYG8MIj3Vw" outputId="b3c07ca8-bec3-4400-ba59-e930d578abb3"
X
# + colab={} colab_type="code" id="MtXoLfKbj3Vx" outputId="98ccafbb-a46f-4b7f-e9d2-b6f51cf96dba"
X * A
# + colab={} colab_type="code" id="8T09ZO4ij3Vz" outputId="573b4ae2-9ca9-49c9-a2b8-086a83ef4af6"
A_p = torch.tensor([[3, 4], [5, 6], [7, 8]])
A_p
# + colab={} colab_type="code" id="VBadBzQJj3V1" outputId="4deeea6d-e38b-43f0-8352-561cd540b366"
X_p * A_p
# + [markdown] colab_type="text" id="s_4kMhF7j3V4"
# Matrix multiplication with a vector:
# + colab={} colab_type="code" id="LAJEstCLj3V5" outputId="1294b0db-7eed-40dd-a98a-8834e248b2d1"
b = np.array([1, 2])
b
# + colab={} colab_type="code" id="ZOxK6XEXj3V8" outputId="d219f59e-304f-4f02-e80b-b79cdf93cb9b"
np.dot(A, b) # even though technically dot products is between 2 vectors
# + colab={} colab_type="code" id="0Y3Ohltbj3V-" outputId="10c44bd8-6249-4f87-e505-37f97f53d546"
b_p = torch.tensor([1, 2])
b_p
# + colab={} colab_type="code" id="LioexYZ_j3WB" outputId="9a565e69-359a-419b-faf9-1af930e1fa65"
torch.matmul(A_p, b_p)
# + [markdown] colab_type="text" id="6Wa5CB28j3WC"
# Matrix multiplication with two matrices:
# + colab={} colab_type="code" id="a8ZXM-T4j3WD" outputId="e43c31b0-b959-448d-e33f-7a1e5950e2b8"
B = np.array([[1, 9], [2, 0]])
B
# + colab={} colab_type="code" id="xE0DA5Xrj3WF" outputId="d56f767e-6057-4f5f-8799-9821e9a39210"
np.dot(A, B) # note first column is same as Xb
# + colab={} colab_type="code" id="bBvYO9jFj3WH" outputId="07ad5cfa-6827-45a7-f416-8c9c80cd7b7a"
B_p = torch.tensor([[1, 9], [2, 0]])
B_p
# + colab={} colab_type="code" id="o2nEnn3Mj3WI" outputId="c22cb10a-d2b3-4e1c-cc09-c5e01ff7a36f"
torch.matmul(A_p, B_p)
# + [markdown] colab_type="text" id="eCFqu8LWj3WJ"
# ### Matrix Inversion
# + colab={} colab_type="code" id="Hhr16-XVj3WK" outputId="9e0aa4f3-a438-4d73-a8e8-b63535d94ba1"
X = np.array([[4, 2], [-5, -3]])
X
# + colab={} colab_type="code" id="FIGmFK9oj3WM" outputId="6f685f9b-f1cb-49ee-8f27-52a644d24a76"
Xinv = np.linalg.inv(X)
Xinv
# + colab={} colab_type="code" id="WWEFauNsj3WO" outputId="517fc8a2-de81-4341-d761-68c866a9be62"
y = np.array([4, -7])
y
# + colab={} colab_type="code" id="egKyXcZhj3WR" outputId="1f6f59e4-82d3-4a65-8d2e-251f5dc5da98"
w = np.dot(Xinv, y)
w
# -
# Show that $y = Xw$:
np.dot(X, w)
# + colab={} colab_type="code" id="jro-lItLj3WT" outputId="417d4901-675b-4641-a791-f48a088ac0b1"
X_p = torch.tensor([[4, 2], [-5, -3.]]) # note that torch.inverse() requires floats
X_p
# + colab={} colab_type="code" id="o4HfI57Nj3WW" outputId="6863c818-6832-4ff3-e78d-126fe19df83f"
Xinv_p = torch.inverse(X_p)
Xinv_p
# + colab={} colab_type="code" id="dy88LRcyj3WY" outputId="95a6c7e0-c066-4ee1-cec3-5e7dfbf4a813"
y_p = torch.tensor([4, -7.])
y_p
# + colab={} colab_type="code" id="1wB3r-ggj3Wa" outputId="3cb28b7a-75cc-4a7b-d707-399cd171361b"
w_p = torch.matmul(Xinv_p, y_p)
w_p
# -
torch.matmul(X_p, w_p)
# + [markdown] colab_type="text" id="oWXikiwFj3Wb"
# **Return to slides here.**
# + [markdown] colab_type="text" id="R4XpcVX2j3Wc"
# ## Segment 2: Eigendecomposition
# + [markdown] colab_type="text" id="et4bgYFDj3Wr"
# ### Eigenvectors and Eigenvalues
# + [markdown] colab_type="text" id="gLjGas2ij3Ws"
# Let's say we have a vector $v$:
# + colab={} colab_type="code" id="zZvzKGRkj3Ws" outputId="180686db-9448-4d12-ca99-08e0f8fa5dcf"
v = np.array([3, 1])
v
# + [markdown] colab_type="text" id="4UlRvHvZj3Wt"
# Let's plot $v$ using Hadrien Jean's handy `plotVectors` function (from [this notebook](https://github.com/hadrienj/deepLearningBook-Notes/blob/master/2.7%20Eigendecomposition/2.7%20Eigendecomposition.ipynb) under [MIT license](https://github.com/hadrienj/deepLearningBook-Notes/blob/master/LICENSE)).
# + colab={} colab_type="code" id="Hl54713Rj3Wt"
import matplotlib.pyplot as plt
# + colab={} colab_type="code" id="fmyWW7Xvj3Wu"
def plotVectors(vecs, cols, alpha=1):
"""
Plot set of vectors.
Parameters
----------
vecs : array-like
Coordinates of the vectors to plot. Each vectors is in an array. For
instance: [[1, 3], [2, 2]] can be used to plot 2 vectors.
cols : array-like
Colors of the vectors. For instance: ['red', 'blue'] will display the
first vector in red and the second in blue.
alpha : float
Opacity of vectors
Returns:
fig : instance of matplotlib.figure.Figure
The figure of the vectors
"""
plt.figure()
plt.axvline(x=0, color='#A9A9A9', zorder=0)
plt.axhline(y=0, color='#A9A9A9', zorder=0)
for i in range(len(vecs)):
x = np.concatenate([[0,0],vecs[i]])
plt.quiver([x[0]],
[x[1]],
[x[2]],
[x[3]],
angles='xy', scale_units='xy', scale=1, color=cols[i],
alpha=alpha)
# + colab={} colab_type="code" id="8uP1xgnLj3Wx" outputId="75194e3a-37ce-4185-ec47-3638ebe86448"
plotVectors([v], cols=['lightblue'])
_ = plt.xlim(-1, 5)
_ = plt.ylim(-1, 5)
# + [markdown] colab_type="text" id="ydGgh5lRj3Wy"
# "Applying" a matrix to a vector (i.e., performing matrix-vector multiplication) can linearly transform the vector, e.g, rotate it or rescale it.
# + [markdown] colab_type="text" id="ReW9fF-sj3Wy"
# The identity matrix, introduced earlier, is the exception that proves the rule: Applying an identity matrix does not transform the vector:
# + colab={} colab_type="code" id="0riv3SS8j3Wz" outputId="3ae4298a-4d45-4a30-bd1f-3bba9d7a8d50"
I = np.array([[1, 0], [0, 1]])
I
# + colab={} colab_type="code" id="nc7nn6VDj3W0" outputId="d0ea9120-1072-441c-e0ca-3599862879d1"
Iv = np.dot(I, v)
Iv
# + colab={} colab_type="code" id="AXQBaq2fj3W1" outputId="80a7263f-b229-47d0-99cd-f712436e26c6"
v == Iv
# + colab={} colab_type="code" id="SP81dIksj3W3" outputId="9375a9ab-bf9d-446d-f977-aa1337a13deb"
plotVectors([Iv], cols=['blue'])
_ = plt.xlim(-1, 5)
_ = plt.ylim(-1, 5)
# + [markdown] colab_type="text" id="JcdkCchSj3W4"
# In contrast, let's see what happens when we apply (some non-identity matrix) $A$ to the vector $v$:
# + colab={} colab_type="code" id="R-EsIE7cj3W4" outputId="8bbeea02-94cf-4347-eb2a-230d585beaf3"
A = np.array([[-1, 4], [2, -2]])
A
# + colab={} colab_type="code" id="DZ0rxeKTj3W5" outputId="3190599a-7754-47d0-9916-0a6153ac148e"
Av = np.dot(A, v)
Av
# + colab={} colab_type="code" id="RCAfNJ8hj3W7" outputId="eccd3efb-e938-4eb9-c313-ec88612f3089"
plotVectors([v, Av], ['lightblue', 'blue'])
_ = plt.xlim(-1, 5)
_ = plt.ylim(-1, 5)
# + colab={} colab_type="code" id="7cMA5yMij3W8" outputId="e2ce5ca4-4749-40c7-b947-3e41ac7bb601"
# a second example:
v2 = np.array([2, 1])
plotVectors([v2, np.dot(A, v2)], ['lightgreen', 'green'])
_ = plt.xlim(-1, 5)
_ = plt.ylim(-1, 5)
# + [markdown] colab_type="text" id="gRqpCA6qj3W-"
# We can concatenate several vectors together into a matrix (say, $V$), where each column is a separate vector. Then, whatever linear transformations we apply to $V$ will be independently applied to each column (vector):
# + colab={} colab_type="code" id="dsRo3xPFj3W-" outputId="7ed6be60-29ec-4dcd-aadb-9c1c5809dd15"
v
# + colab={} colab_type="code" id="nQDdsKXNj3XA" outputId="17255afd-aa09-4f00-cc5e-45ce87cb92f0"
# recall that we need to convert array to 2D to transpose into column, e.g.:
np.matrix(v).T
# + colab={} colab_type="code" id="Zb9cSgjnj3XB"
v3 = np.array([-3, -1]) # mirror image of x over both axes
v4 = np.array([-1, 1])
# + colab={} colab_type="code" id="EsdcEFXYj3XC" outputId="82d690a4-4609-4066-8cba-35d2ec7a3dc9"
V = np.concatenate((np.matrix(v).T,
np.matrix(v2).T,
np.matrix(v3).T,
np.matrix(v4).T),
axis=1)
V
# + colab={} colab_type="code" id="t3CSo0Hrj3XD" outputId="b568ad45-390d-40dd-ad88-e74e35f36172"
IV = np.dot(I, V)
IV
# + colab={} colab_type="code" id="a6M61JEWj3XE" outputId="2bacb4b7-accc-48a1-f966-9717fcc04343"
AV = np.dot(A, V)
AV
# + colab={} colab_type="code" id="wGmvjr85j3XG"
# function to convert column of matrix to 1D vector:
def vectorfy(mtrx, clmn):
return np.array(mtrx[:,clmn]).reshape(-1)
# + colab={} colab_type="code" id="L2eQ4w83j3XH" outputId="6a972e6d-6011-4842-ebab-c4368e1ee456"
vectorfy(V, 0)
# + colab={} colab_type="code" id="_o4HOgEwj3XI" outputId="9f4c5c00-997d-40a8-cca1-caa2e8377def"
vectorfy(V, 0) == v
# + colab={} colab_type="code" id="btsivNB_j3XK" outputId="63d1db88-f0c4-414a-ed8f-67b37244fc02"
plotVectors([vectorfy(V, 0), vectorfy(V, 1), vectorfy(V, 2), vectorfy(V, 3),
vectorfy(AV, 0), vectorfy(AV, 1), vectorfy(AV, 2), vectorfy(AV, 3)],
['lightblue', 'lightgreen', 'lightgray', 'orange',
'blue', 'green', 'gray', 'red'])
_ = plt.xlim(-4, 6)
_ = plt.ylim(-5, 5)
# + [markdown] colab_type="text" id="4n31lIDpj3XL"
# Now that we can appreciate linear transformation of vectors by matrices, let's move on to working with eigenvectors and eigenvalues.
#
# An **eigenvector** (*eigen* is German for "typical"; we could translate *eigenvector* to "characteristic vector") is a special vector $v$ such that when it is transformed by some matrix (let's say $A$), the product $Av$ has the exact same direction as $v$.
#
# An **eigenvalue** is a scalar (traditionally represented as $\lambda$) that simply scales the eigenvector $v$ such that the following equation is satisfied:
#
# $Av = \lambda v$
# + [markdown] colab_type="text" id="G7zVjpW-j3XL"
# Easiest way to understand this is to work through an example:
# + colab={} colab_type="code" id="GbXJtk1Ej3XL" outputId="4fe3b949-175c-4a22-873a-7690d3e4c443"
A
# + [markdown] colab_type="text" id="0t2JsM6Wj3XN"
# Eigenvectors and eigenvalues can be derived algebraically (e.g., with the [QR algorithm](https://en.wikipedia.org/wiki/QR_algorithm), which was independent developed in the 1950s by both [Vera Kublanovskaya](https://en.wikipedia.org/wiki/Vera_Kublanovskaya) and John Francis), however this is outside scope of today's class. We'll cheat with NumPy `eig()` method, which returns a tuple of:
#
# * a vector of eigenvalues
# * a matrix of eigenvectors
# + colab={} colab_type="code" id="CYWmgM7jj3XN"
lambdas, V = np.linalg.eig(A)
# + [markdown] colab_type="text" id="OVUp_p6-j3XO"
# The matrix contains as many eigenvectors as there are columns of A:
# + colab={} colab_type="code" id="7iHlAMa7j3XO" outputId="9a4a2b4c-401c-4587-cd11-6f60751d4ce7"
V # each column is a separate eigenvector v
# + [markdown] colab_type="text" id="EBSmc2JLj3XP"
# With a corresponding eigenvalue for each eigenvector:
# + colab={} colab_type="code" id="QYJbCydNj3XP" outputId="527f7f51-b2b5-4f0c-f840-bd5b144c2eba"
lambdas
# + [markdown] colab_type="text" id="OoZPiaBMj3XR"
# Let's confirm that $Av = \lambda v$ for the first eigenvector:
# + colab={} colab_type="code" id="n1QZ57TJj3XR" outputId="32386f00-070a-4c6a-eb30-3d2d09c45a4b"
v = V[:,0]
v
# + colab={} colab_type="code" id="6vwkWMiIj3XV" outputId="1b274e96-01d7-4104-fbe7-f91ae5826fe9"
lambduh = lambdas[0] # note that "lambda" is reserved term in Python
lambduh
# + colab={} colab_type="code" id="Leh9n8QBj3XW" outputId="3165479f-bb06-422d-bbcf-5e8b4b35e66e"
Av = np.dot(A, v)
Av
# + colab={} colab_type="code" id="PROIJU30j3XX" outputId="833d6e53-6d1b-4759-f5cf-3e28af96ff30"
lambduh * v
# + colab={} colab_type="code" id="0bT3vjoQj3XY" outputId="dc53ddfc-5703-407e-ec3a-e146d779841c"
plotVectors([Av, v], ['blue', 'lightblue'])
_ = plt.xlim(-1, 2)
_ = plt.ylim(-1, 2)
# + [markdown] colab_type="text" id="tKQI4691j3XZ"
# And again for the second eigenvector of A:
# + colab={} colab_type="code" id="riOJuqz3j3XZ" outputId="992b8b02-ef09-4de5-db15-bffcedc722a2"
v2 = V[:,1]
v2
# + colab={} colab_type="code" id="QveeHYhDj3Xa" outputId="9985e974-655f-4acb-dc38-e5fac398ae7b"
lambda2 = lambdas[1]
lambda2
# + colab={} colab_type="code" id="TDk1VoVIj3Xb" outputId="ccc3fa42-455a-4491-f75e-86d6f9c464f6"
Av2 = np.dot(A, v2)
Av2
# + colab={} colab_type="code" id="smlYxxgpj3Xc" outputId="e36bef99-5e6a-42a9-8669-910e636676df"
lambda2 * v2
# + colab={} colab_type="code" id="3IigKHp0j3Xd" outputId="c2dc34fb-5384-47de-a325-4e053417563c"
plotVectors([Av, v, Av2, v2],
['blue', 'lightblue', 'green', 'lightgreen'])
_ = plt.xlim(-1, 4)
_ = plt.ylim(-3, 2)
# + [markdown] colab_type="text" id="VF9uLWjOj3Xe"
# Using the PyTorch `eig()` method, we can do exactly the same:
# + colab={} colab_type="code" id="EcJa6w0mj3Xe" outputId="8980d896-85e5-4081-d373-fc61e3cba9d8"
A
# + colab={} colab_type="code" id="9WvCqoRij3Xf" outputId="309a5053-1af0-4859-9dd6-8101dc78aaff"
A_p = torch.tensor([[-1, 4], [2, -2.]]) # must be float for PyTorch eig()
A_p
# + colab={} colab_type="code" id="27FbfPXGj3Xg" outputId="df1ccbc4-31a0-4440-e006-b08a5c4f5daa"
eigens = torch.eig(A_p, eigenvectors=True)
eigens
# + colab={} colab_type="code" id="gz2_l7ebj3Xh" outputId="3f82b260-845e-4208-fe69-405a73f4fa09"
v_p = eigens.eigenvectors[:,0]
v_p
# + colab={} colab_type="code" id="VrYaxNCRj3Xj" outputId="565234c5-973f-4992-8281-e2125d751a70"
lambda_p = eigens.eigenvalues[0][0]
lambda_p
# + colab={} colab_type="code" id="SUq1UGH7j3Xl" outputId="0a1ab88b-c83c-4938-df90-d25c13f3336f"
Av_p = torch.matmul(A_p, v_p)
Av_p
# + colab={} colab_type="code" id="co1VNLIej3Xn" outputId="7a6ab193-43d9-405d-f55f-6b70a05da0ae"
lambda_p * v_p
# + colab={} colab_type="code" id="1b47vG92j3Xo" outputId="8c7f2adb-b54e-4adf-f928-f9ca2646874e"
v2_p = eigens.eigenvectors[:,1]
v2_p
# + colab={} colab_type="code" id="B-evpW17j3Xp" outputId="7e7c3cc5-ebfa-4c00-cf82-3c2623758be3"
lambda2_p = eigens.eigenvalues[1][0]
lambda2_p
# + colab={} colab_type="code" id="2d5gjh2sj3Xq" outputId="56ed7ce8-aaf4-4b9e-d9fa-ec5f8f6d31c5"
Av2_p = torch.matmul(A_p, v2_p)
Av2_p
# + colab={} colab_type="code" id="8-ki3i8dj3Xr" outputId="1d1b5f93-370f-405e-fac5-eba55e9267d9"
lambda2_p * v2_p
# + colab={} colab_type="code" id="s2p8yb_Zj3Xs" outputId="fca8be0a-eb8c-4f1b-bf3e-12883a5cdd09"
plotVectors([Av_p.numpy(), v_p.numpy(), Av2_p.numpy(), v2_p.numpy()],
['blue', 'lightblue', 'green', 'lightgreen'])
_ = plt.xlim(-1, 4)
_ = plt.ylim(-3, 2)
# + [markdown] colab_type="text" id="6-HqwyESj3Xt"
# ### Eigenvectors in >2 Dimensions
# + [markdown] colab_type="text" id="M01JPwToj3Xt"
# While plotting gets trickier in higher-dimensional spaces, we can nevertheless find and use eigenvectors with more than two dimensions. Here's a 3D example (there are three dimensions handled over three rows):
# + colab={} colab_type="code" id="HWsBDEMgj3Xt" outputId="da9e86f2-da5f-4586-cb43-5a4a997492db"
X
# + colab={} colab_type="code" id="Y-uUMyRFj3Xv"
lambdas_X, V_X = np.linalg.eig(X)
# + colab={} colab_type="code" id="virh7GVFj3Xw" outputId="9de3d28c-610b-4b8a-e913-8cd84f2383ec"
V_X # one eigenvector per column of X
# + colab={} colab_type="code" id="9yHkmEd0j3Xw" outputId="f2c36081-ca90-451c-d6dd-b1bf4416abc0"
lambdas_X # a corresponding eigenvalue for each eigenvector
# + [markdown] colab_type="text" id="Qp3qPeUxj3Xy"
# Confirm $Xv = \lambda v$ for an example vector:
# + colab={} colab_type="code" id="dUEfbThhj3Xy" outputId="d28db68f-5855-4ed1-bf62-b1972ed93461"
v_X = V_X[:,0]
v_X
# + colab={} colab_type="code" id="xhnF5asDj3X0" outputId="86a568fe-27d1-456b-bcc7-aa5fff4e8c84"
lambda_X = lambdas_X[0]
lambda_X
# + colab={} colab_type="code" id="K3bA6vRzj3X1" outputId="be53cc8d-00a2-4d6c-dd34-9dcc9b2906fd"
np.dot(X, v_X) # matrix multiplication
# + colab={} colab_type="code" id="UfN3hk0Gj3X2" outputId="249e8bca-d69e-4202-ab61-1d3232629e52"
lambda_X * v_X
# + [markdown] colab_type="text" id="VcjTZv24j3X3"
# **Exercises**:
#
# 1. Use PyTorch to confirm $Xv = \lambda v$ for the first eigenvector of $X$.
# 2. Confirm $Xv = \lambda v$ for the remaining eigenvectors of $X$ (you can use NumPy or PyTorch, whichever you prefer).
# + [markdown] colab_type="text" id="5z5AdeHKj3X4"
# **Return to slides here.**
# + [markdown] colab_type="text" id="F44cMjS8j3Wc"
# ### 2x2 Matrix Determinants
# + colab={} colab_type="code" id="GjsXPZEsj3Wc" outputId="4c12d175-8993-4c94-b590-8602378c8663"
X
# + colab={} colab_type="code" id="4tg8BDzOj3We" outputId="276a9faf-2755-4277-c4c2-16fc8afb48bf"
np.linalg.det(X)
# + [markdown] colab_type="text" id="87wpY5hUj3Wg"
# **Return to slides here.**
# + colab={} colab_type="code" id="o5jp6vkNj3Wg" outputId="365f8b28-32a5-479a-ab8c-a2b0b2c67219"
N = np.array([[-4, 1], [-8, 2]])
N
# + colab={} colab_type="code" id="ejdkdN7Lj3Wi" outputId="8ecb038d-ad39-4109-8c92-42af5d01351e"
np.linalg.det(N)
# + colab={} colab_type="code" id="nbIrtcaCj3Wj"
# Uncommenting the following line results in a "singular matrix" error
# Ninv = np.linalg.inv(N)
# + colab={} colab_type="code" id="kBT5VC5Nj3Wl"
N = torch.tensor([[-4, 1], [-8, 2.]]) # must use float not int
# + colab={} colab_type="code" id="MQA5g0gGj3Wm" outputId="251da4e8-92b8-45ae-e63a-aa4a66f3ba22"
torch.det(N)
# + [markdown] colab_type="text" id="XBIJB7tfj3Wn"
# **Return to slides here.**
# + [markdown] colab_type="text" id="rwwoY1k6j3Wn"
# ### Generalizing Determinants
# + colab={} colab_type="code" id="bqjRKaCaj3Wn" outputId="4a38488f-5ab7-4959-f813-6fb3443e85ee"
X = np.array([[1, 2, 4], [2, -1, 3], [0, 5, 1]])
X
# + colab={} colab_type="code" id="uimvv39Nj3Wp" outputId="2c2a0732-0270-4bf5-e06e-956af66afb53"
np.linalg.det(X)
# -
# ### Determinants & Eigenvalues
# + colab={} colab_type="code" id="g0uEEY2qj3X6" outputId="46b324e0-bd95-4c8b-f91b-2ea188b0e0a4"
lambdas, V = np.linalg.eig(X)
lambdas
# + colab={} colab_type="code" id="41P8dP9Gj3X8" outputId="a54bd6e4-055c-4999-a78b-325ede395597"
np.product(lambdas)
# -
# **Return to slides here.**
# + colab={} colab_type="code" id="a7Bleu07j3X-" outputId="1a190bab-cc1c-4961-9dee-b56a3479ea44"
np.abs(np.product(lambdas))
# + colab={} colab_type="code" id="rMPe8LOXj3X_" outputId="d3f2f157-85f7-49e0-e084-c8329901c297"
B = np.array([[1, 0], [0, 1]])
B
# + colab={} colab_type="code" id="zlhnOiNzj3YA" outputId="aeef875f-c003-48de-e140-d51bec5bc5f8"
plotVectors([vectorfy(B, 0), vectorfy(B, 1)],
['lightblue', 'lightgreen'])
_ = plt.xlim(-1, 3)
_ = plt.ylim(-1, 3)
# + colab={} colab_type="code" id="Fjpem_6Ij3YB" outputId="5357a768-5ce5-4506-e7ef-2fcb8eff821b"
N
# + colab={} colab_type="code" id="2BhgWTvaj3YC" outputId="b5977945-7ed4-4301-a884-ebc5d179198a"
np.linalg.det(N)
# + colab={} colab_type="code" id="O3XSySPaj3YE" outputId="99134bfc-b3d2-4f17-80eb-6256fc8d0c08"
NB = np.dot(N, B)
NB
# + colab={} colab_type="code" id="GLiyz0nxj3YF" outputId="6e0f19af-b700-4a5d-d9ef-f83e1d15bc93"
plotVectors([vectorfy(B, 0), vectorfy(B, 1), vectorfy(NB, 0), vectorfy(NB, 1)],
['lightblue', 'lightgreen', 'blue', 'green'])
_ = plt.xlim(-6, 6)
_ = plt.ylim(-9, 3)
# + colab={} colab_type="code" id="mqEkY-8lj3YH" outputId="75b1aa94-044e-4950-a7e2-89cf6fa15414"
I
# + colab={} colab_type="code" id="zwmKA0wLj3YI" outputId="b7ec8bd6-8bfa-48f6-c2b5-c6109c7546d1"
np.linalg.det(I)
# + colab={} colab_type="code" id="xaIq_dUKj3YJ" outputId="0cf90877-a9e7-4863-dc6a-07d96b314efb"
IB = np.dot(I, B)
IB
# + colab={} colab_type="code" id="8y6svzaLj3YK" outputId="a4f1d555-790a-445e-eff8-7027f658143b"
plotVectors([vectorfy(B, 0), vectorfy(B, 1), vectorfy(IB, 0), vectorfy(IB, 1)],
['lightblue', 'lightgreen', 'blue', 'green'])
_ = plt.xlim(-1, 3)
_ = plt.ylim(-1, 3)
# + colab={} colab_type="code" id="Xj4bnqVcj3YL" outputId="41795c0a-8a56-4bef-9097-1788cd5d906f"
J = np.array([[-0.5, 0], [0, 2]])
J
# + colab={} colab_type="code" id="D2ODTOpAj3YM" outputId="5b6498e0-9bef-45eb-fb30-7869eace08ee"
np.linalg.det(J)
# + colab={} colab_type="code" id="3kHJ7Q2Ij3YN" outputId="12d2b176-944d-4a1e-c140-b489ea91b5d2"
np.abs(np.linalg.det(J))
# + colab={} colab_type="code" id="ABciINQwj3YO" outputId="e3c94eca-4cf9-4e78-cf0c-8adabc234e97"
JB = np.dot(J, B)
JB
# + colab={} colab_type="code" id="5nWa22ZNj3YO" outputId="14671995-3dd7-4ba6-815d-88cba03e8302"
plotVectors([vectorfy(B, 0), vectorfy(B, 1), vectorfy(JB, 0), vectorfy(JB, 1)],
['lightblue', 'lightgreen', 'blue', 'green'])
_ = plt.xlim(-1, 3)
_ = plt.ylim(-1, 3)
# + colab={} colab_type="code" id="u-OI7xBWj3YQ"
doubleI = I*2
# + colab={} colab_type="code" id="iKoTu9sbj3YR" outputId="b4de617b-60c9-43ff-9084-d105fb6bc62e"
np.linalg.det(doubleI)
# + colab={} colab_type="code" id="kTb0o2Ydj3YS" outputId="847204c0-ed6c-430b-f8fc-20d6a3c6e74f"
doubleIB = np.dot(doubleI, B)
doubleIB
# + colab={} colab_type="code" id="3J2ou_zSj3YT" outputId="d8646e60-d161-4963-e2f9-ef26726853db"
plotVectors([vectorfy(B, 0), vectorfy(B, 1), vectorfy(doubleIB, 0), vectorfy(doubleIB, 1)],
['lightblue', 'lightgreen', 'blue', 'green'])
_ = plt.xlim(-1, 3)
_ = plt.ylim(-1, 3)
# + [markdown] colab_type="text" id="Av0R8fddj3Yb"
# **Return to slides here.**
# + [markdown] colab_type="text" id="MXNajp3Ej3Yb"
# ### Eigendecomposition
# + [markdown] colab_type="text" id="wQt403xbj3Yb"
# The **eigendecomposition** of some matrix $A$ is
#
# $A = V \Lambda V^{-1}$
#
# Where:
#
# * As in examples above, $V$ is the concatenation of all the eigenvectors of $A$
# * $\Lambda$ (upper-case $\lambda$) is the diagonal matrix diag($\lambda$). Note that the convention is to arrange the lambda values in descending order; as a result, the first eigenvector (and its associated eigenvector) may be a primary characteristic of the matrix $A$.
# + colab={} colab_type="code" id="W7LmR3YGj3Yb" outputId="4b5c460a-b49c-4241-c544-0f62f0ea33d2"
# This was used earlier as a matrix X; it has nice clean integer eigenvalues...
A = np.array([[4, 2], [-5, -3]])
A
# + colab={} colab_type="code" id="37zeBrqhj3Yc"
lambdas, V = np.linalg.eig(A)
# + colab={} colab_type="code" id="b7LtIIMJj3Yd" outputId="7d9608df-6e8f-42e0-8004-462eb48267f2"
V
# + colab={} colab_type="code" id="q1uuRwcdj3Ye" outputId="8383e3f8-ef3b-421b-8347-e26a1f0db56b"
Vinv = np.linalg.inv(V)
Vinv
# + colab={} colab_type="code" id="_InHRuS1j3Yf" outputId="0191817d-bf9b-455c-deec-1c4d068065b8"
Lambda = np.diag(lambdas)
Lambda
# + [markdown] colab_type="text" id="KSBnzTBZj3Yg"
# Confirm that $A = V \Lambda V^{-1}$:
# + colab={} colab_type="code" id="pG1E3yLYj3Yg" outputId="59e6239c-e5ee-453f-8491-6ca89ffcc9ff"
np.dot(V, np.dot(Lambda, Vinv))
# + [markdown] colab_type="text" id="JTmKZk8fj3Yh"
# Eigendecomposition is not possible with all matrices. And in some cases where it is possible, the eigendecomposition involves complex numbers instead of straightforward real numbers.
#
# In machine learning, however, we are typically working with real symmetric matrices, which can be conveniently and efficiently decomposed into real-only eigenvectors and real-only eigenvalues. If $A$ is a real symmetric matrix then...
#
# $A = Q \Lambda Q^T$
#
# ...where $Q$ is analogous to $V$ from the previous equation except that it's special because it's an orthogonal matrix.
# + colab={} colab_type="code" id="GpZLd9Ozj3Yh" outputId="ad679f93-087a-4021-caf5-9bf80eac47c4"
A = np.array([[2, 1], [1, 2]])
A
# + colab={} colab_type="code" id="EJOgExEZj3Yj"
lambdas, Q = np.linalg.eig(A)
# + colab={} colab_type="code" id="9qouDzN5j3Yk" outputId="19d23366-93e8-4e84-a74d-040bb382748d"
lambdas
# + colab={} colab_type="code" id="JZFyXQzkj3Yl" outputId="782e731a-d0f8-404e-9226-f01bf2c9d7ab"
Lambda = np.diag(lambdas)
Lambda
# + colab={} colab_type="code" id="BLXaGoVBj3Yl" outputId="86a3fdd6-05aa-41b7-f832-1b7c65806ee3"
Q
# + [markdown] colab_type="text" id="_eq_1nssj3Ym"
# Recalling that $Q^TQ = QQ^T = I$, can demonstrate that $Q$ is an orthogonal matrix:
# + colab={} colab_type="code" id="TcavBhdEj3Ym" outputId="590b2ba2-418e-40aa-eb7d-5c0550d74988"
np.dot(Q.T, Q)
# + colab={} colab_type="code" id="xup113b8j3Yo" outputId="df2493e1-e0a1-4499-8234-3feacc1ff9e3"
np.dot(Q, Q.T)
# + [markdown] colab_type="text" id="UnOfuIf7j3Yo"
# Let's confirm $A = Q \Lambda Q^T$:
# + colab={} colab_type="code" id="k4DukMWJj3Yo" outputId="eafffd5d-0ffb-4636-c225-5fb9c15d09f8"
np.dot(Q, np.dot(Lambda, Q.T))
# + [markdown] colab_type="text" id="fdJaTKnsj3Yp"
# **Exercises**:
#
# 1. Use PyTorch to decompose the matrix $P$ (below) into its components $V$, $\Lambda$, and $V^{-1}$. Confirm that $P = V \Lambda V^{-1}$.
# 2. Use PyTorch to decompose the symmetric matrix $S$ (below) into its components $Q$, $\Lambda$, and $Q^T$. Confirm that $S = Q \Lambda Q^T$.
# + colab={} colab_type="code" id="_RVUCVlvj3Yp" outputId="7abc66ce-1fa3-4c1b-cea9-1da3d0f8ade6"
P = torch.tensor([[25, 2, -5], [3, -2, 1], [5, 7, 4.]])
P
# + colab={} colab_type="code" id="GjKZ_AWLj3Yq" outputId="b9ee6619-60b4-4fb5-c88b-0fe313039635"
S = torch.tensor([[25, 2, -5], [2, -2, 1], [-5, 1, 4.]])
S
# + [markdown] colab_type="text" id="1OFq3uGaj3Yq"
# **Return to slides here.**
# + [markdown] colab_type="text" id="gKam0tJOj3Yr"
# ## Segment 3: Matrix Operations for ML
# + [markdown] colab_type="text" id="j-wbn7omj3Yr"
# ### Singular Value Decomposition (SVD)
# + [markdown] colab_type="text" id="x2SHytttj3Yr"
# As on slides, SVD of matrix $A$ is:
#
# $A = UDV^T$
#
# Where:
#
# * $U$ is an orthogonal $m \times m$ matrix; its columns are the **left-singular vectors** of $A$.
# * $V$ is an orthogonal $n \times n$ matrix; its columns are the **right-singular vectors** of $A$.
# * $D$ is a diagonal $m \times n$ matrix; elements along its diagonal are the **singular values** of $A$.
# + colab={} colab_type="code" id="V7hR4Htdj3Yr" outputId="d0399dd8-fde2-4c20-8015-e825117c76a9"
A = np.array([[-1, 2], [3, -2], [5, 7]])
A
# + colab={} colab_type="code" id="ihj2XfMQj3Ys"
U, d, VT = np.linalg.svd(A) # V is already transposed
# + colab={} colab_type="code" id="DUfP2aaTj3Yv" outputId="37f47eca-2391-46b3-993f-923dc20eb034"
U
# + colab={} colab_type="code" id="s_Fkoarvj3Yw" outputId="ed097007-023b-43cb-9e9a-f4d7bc925466"
VT
# + colab={} colab_type="code" id="wNSRDcfsj3Yx" outputId="5e0565b6-c57a-4d08-ec2e-da33d7cc956f"
d
# + colab={} colab_type="code" id="Lbxh2rYoj3Yy" outputId="a62efd6d-c1d3-463b-8c7b-4621bf170349"
np.diag(d)
# + colab={} colab_type="code" id="V47I3B87j3Y0" outputId="6e0dbbd4-f080-4f50-f56a-f1fbc95248a1"
D = np.concatenate((np.diag(d), [[0, 0]]), axis=0)
D
# + colab={} colab_type="code" id="9euCs5vvj3Y2" outputId="75c652fe-01bb-45a1-d9a6-89e91bf83e54"
np.dot(U, np.dot(D, VT))
# + [markdown] colab_type="text" id="u-WCBOzKj3Y3"
# SVD and eigendecomposition are closely related to each other:
#
# * Left-singular vectors of $A$ = eigenvectors of $AA^T$.
# * Right-singular vectors of $A$ = eigenvectors of $A^TA$.
# * Non-zero singular values of $A$ = square roots of eigenvectors of $AA^T$ = square roots of eigenvectors of $A^TA$
#
# **Exercise**: Using the matrix `P` from the preceding PyTorch exercises, demonstrate that these three SVD-eigendecomposition equations are true.
# + [markdown] colab_type="text" id="CWEOMqUUj3Y3"
# ### Image Compression via SVD
# + [markdown] colab_type="text" id="XmRvLo_Tj3Y3"
# The section features code adapted from [<NAME>'s](https://gist.github.com/frankcleary/4d2bd178708503b556b0).
# + colab={} colab_type="code" id="luD8Y98Vj3Y3"
import time
from PIL import Image
# + [markdown] colab_type="text" id="thPmYUx4j3Y4"
# Fetch photo of Oboe, a terrier, with the book *Deep Learning Illustrated*:
# + colab={} colab_type="code" id="bPUItNUVj3Y4" outputId="cb38ee6c-a746-4318-c8a6-f6fed7d9b163"
# ! wget https://raw.githubusercontent.com/jonkrohn/DLTFpT/master/notebooks/oboe-with-book.jpg
# + colab={} colab_type="code" id="6lx_Frl6j3Y6" outputId="a9b87d11-2d9a-4d28-db5c-512985470336"
img = Image.open('oboe-with-book.jpg')
plt.imshow(img)
# + [markdown] colab_type="text" id="XYmg1Fa8j3Y6"
# Convert image to grayscale so that we don't have to deal with the complexity of multiple color channels:
# + colab={} colab_type="code" id="uki3S6w0j3Y7" outputId="97a2a67e-bc40-4de2-f162-9777a4e5c8b3"
imggray = img.convert('LA')
plt.imshow(imggray)
# + [markdown] colab_type="text" id="eVwgrA0Jj3Y9"
# Convert data into numpy matrix, which doesn't impact image data:
# + colab={} colab_type="code" id="wHijyFgUj3Y9" outputId="6a6edcbf-aed9-461c-95ab-f9353287c330"
imgmat = np.array(list(imggray.getdata(band=0)), float)
imgmat.shape = (imggray.size[1], imggray.size[0])
imgmat = np.matrix(imgmat)
plt.imshow(imgmat, cmap='gray')
# + [markdown] colab_type="text" id="x8VCD3lyj3Y-"
# Calculate SVD of the image:
# + colab={} colab_type="code" id="kbBLn2Csj3Y-"
U, sigma, V = np.linalg.svd(imgmat)
# + [markdown] colab_type="text" id="ApybkCdLj3Y-"
# As eigenvalues are arranged in descending order in diag($\lambda$) so to are singular values, by convention, arranged in descending order in $D$ (or, in this code, diag($\sigma$)). Thus, the first left-singular vector of $U$ and first right-singular vector of $V$ may represent the most prominent feature of the image:
# + colab={} colab_type="code" id="rZTwlhGxj3Y_" outputId="1560da9a-70a4-4529-ff7d-f6df772e608e"
reconstimg = np.matrix(U[:, :1]) * np.diag(sigma[:1]) * np.matrix(V[:1, :])
plt.imshow(reconstimg, cmap='gray')
# + [markdown] colab_type="text" id="4p2cEqIoj3Y_"
# Additional singular vectors improve the image quality:
# + colab={} colab_type="code" id="f5-6LEbij3ZA" outputId="41a72677-6f00-46b7-d440-7c18546bfee3"
for i in [2, 4, 8, 16, 32, 64]:
reconstimg = np.matrix(U[:, :i]) * np.diag(sigma[:i]) * np.matrix(V[:i, :])
plt.imshow(reconstimg, cmap='gray')
title = "n = %s" % i
plt.title(title)
plt.show()
# + [markdown] colab_type="text" id="IjfUJ4wNj3ZA"
# With 64 singular vectors, the image is reconstructed quite well, however the data footprint is much smaller than the original image:
# + colab={} colab_type="code" id="hXQy4TzCj3ZB" outputId="28b856f8-c3e0-4da1-ee3d-7dbf12fc92de"
imgmat.shape
# -
full_representation = 4032*3024
full_representation
svd64_rep = 64*4032 + 64 + 64*3024
svd64_rep
# + colab={} colab_type="code" id="YwdA6taLj3ZD" outputId="fd993012-1e41-4817-9037-ee5867b4ed15"
svd64_rep/full_representation
# + [markdown] colab_type="text" id="_HdtL8p7j3ZD"
# Specifically, the image represented as 64 singular vectors is 3.7% of the size of the original!
# + [markdown] colab_type="text" id="lBQnc4uGj3ZE"
# **Return to slides here.**
# + [markdown] colab_type="text" id="FEnTYLbBj3ZE"
# ### The Moore-Penrose Pseudoinverse
# + [markdown] colab_type="text" id="q4_MX4D5j3ZE"
# Let's calculate the pseudoinverse $A^+$ of some matrix $A$ using the formula from the slides:
#
# $A^+ = VD^+U^T$
# + colab={} colab_type="code" id="gbbiRcmzj3ZE" outputId="ed4921a7-8360-46b3-e928-5d554b2353b9"
A
# + [markdown] colab_type="text" id="TFNYVdZ4j3ZF"
# As shown earlier, the NumPy SVD method returns $U$, $d$, and $V^T$:
# + colab={} colab_type="code" id="I6NjgNbjj3ZF"
U, d, VT = np.linalg.svd(A)
# + colab={} colab_type="code" id="_sROQTAzj3ZG" outputId="eb770b9f-7790-4749-ec31-f6fbff959c11"
U
# + colab={} colab_type="code" id="drDSW-vkj3ZG" outputId="392d0405-73d8-4b22-90bd-48596c41ccfd"
VT
# + colab={} colab_type="code" id="Sr3jIv4Jj3ZH" outputId="b4affc7f-02c0-42e9-ce4f-08b3b9029b67"
d
# + [markdown] colab_type="text" id="7yalmt19j3ZI"
# To create $D^+$, we first invert the non-zero values of $d$:
# + colab={} colab_type="code" id="HEvRLwQfj3ZI" outputId="12e4dd08-d35d-4949-93c9-c766d91c23fd"
D = np.diag(d)
D
# + colab={} colab_type="code" id="6O2LADtLj3ZJ" outputId="d90103ed-14cc-4d0a-de03-c9f433904535"
1/8.669
# + colab={} colab_type="code" id="y1ZbYtmIj3ZJ" outputId="5da3ca1f-3392-4112-aa3b-718d634987b0"
1/4.104
# + [markdown] colab_type="text" id="yO1tNES9j3ZK"
# ...and then we would take the tranpose of the resulting matrix.
#
# Because $D$ is a diagonal matrix, this can, however, be done in a single step by inverting $D$:
# + colab={} colab_type="code" id="Y0sjcP3Ej3ZK" outputId="237289cf-5784-4d92-da00-e8489a7be541"
Dinv = np.linalg.inv(D)
Dinv
# + [markdown] colab_type="text" id="PXJnvna-j3ZM"
# The final $D^+$ matrix needs to have a shape that can undergo matrix multiplication in the $A^+ = VD^+U^T$ equation. These dimensions can be obtained from $A$:
# + colab={} colab_type="code" id="yK1PadPFj3ZM" outputId="01fc6273-3895-4e20-81be-bbcec80b783e"
A.shape[0]
# + colab={} colab_type="code" id="E5bfwPDSj3ZM" outputId="fb6e523d-8dc3-4b4f-b355-882e710861da"
A.shape[1]
# + colab={} colab_type="code" id="Ghxh-1P4j3ZN" outputId="d6023ea0-0525-475d-e7e7-39018582b66e"
Dplus = np.zeros((3, 2)).T
Dplus
# + colab={} colab_type="code" id="UTfh42jEj3ZO" outputId="6f847066-ba6b-4f2a-f106-9f114066fd46"
Dplus[:2, :2] = Dinv
Dplus
# + [markdown] colab_type="text" id="6Xt4NYHuj3ZO"
# Now we have everything we need to calculate $A^+$ with $VD^+U^T$:
# + colab={} colab_type="code" id="ZtWN_wnij3ZO" outputId="46b2627b-ebd8-4f12-d554-e2a0d49ce031"
np.dot(VT.T, np.dot(Dplus, U.T))
# + [markdown] colab_type="text" id="3syT7-hCj3ZP"
# Working out this derivation is helpful for understanding how Moore-Penrose pseudoinverses work, but unsurprisingly NumPy is loaded with an existing method `pinv()`:
# + colab={} colab_type="code" id="fh0nDMeLj3ZP" outputId="ea3f09c0-2046-42ea-ae93-121ab9e7e585"
np.linalg.pinv(A)
# + [markdown] colab_type="text" id="xNrIfpAij3ZS"
# **Exercise**
#
# Use the `torch.svd()` method to calculate the pseudoinverse of `A_p`, confirming that your result matches the output of `torch.pinverse(A_p)`:
# + colab={} colab_type="code" id="W2635vlEj3ZS" outputId="702b613f-1748-4af8-b515-6c9e887ce8b2"
A_p = torch.tensor([[-1, 2], [3, -2], [5, 7.]])
A_p
# + colab={} colab_type="code" id="ZW4SsUOlj3ZT" outputId="86d514f8-1fac-4ed1-b6ec-00c167066667"
torch.pinverse(A_p)
# + [markdown] colab_type="text" id="KlXBgI3Nj3ZT"
# **Return to slides here.**
# + [markdown] colab_type="text" id="xMnIqjpfj3ZT"
# For regression problems, we typically have many more cases ($n$, or rows of $X$) than features to predict ($m$, or columns of $X$). Let's solve a miniature example of such an overdetermined situation.
#
# We have eight data points ($n$ = 8):
# + colab={} colab_type="code" id="2Ft4PXaTj3ZU"
x1 = [0, 1, 2, 3, 4, 5, 6, 7.] # E.g.: Dosage of drug for treating Alzheimer's disease
y = [1.86, 1.31, .62, .33, .09, -.67, -1.23, -1.37] # E.g.: Patient's "forgetfulness score"
# -
title = 'Clinical Trial'
xlabel = 'Drug dosage (mL)'
ylabel = 'Forgetfulness'
# + colab={} colab_type="code" id="OiMAISFBj3ZW" outputId="f4739a01-fe06-432a-ede9-0ee8eadba0ca"
fig, ax = plt.subplots()
plt.title(title)
plt.xlabel(xlabel)
plt.ylabel(ylabel)
_ = ax.scatter(x1, y)
# + [markdown] colab_type="text" id="GWZUFeqzj3ZX"
# Although it appears there is only one predictor ($x_1$), we need a second one (let's call it $x_0$) in order to allow for a $y$-intercept (therefore, $m$ = 2). Without this second variable, the line we fit to the plot would need to pass through the origin (0, 0). The $y$-intercept is constant across all the points so we can set it equal to `1` across the board:
# + colab={} colab_type="code" id="RpAIoxydj3ZX" outputId="23fba07b-daf5-4e46-da2f-e2018dbf11c8"
x0 = np.ones(8)
x0
# + [markdown] colab_type="text" id="_bkwC8Wnj3ZY"
# Concatenate $x_0$ and $x_1$ into a matrix $X$:
# + colab={} colab_type="code" id="x56TMNFMj3ZY" outputId="f5d261ff-1fd8-449f-f780-81de2ac91abe"
X = np.concatenate((np.matrix(x0).T, np.matrix(x1).T), axis=1)
X
# + [markdown] colab_type="text" id="v7TomkyCj3ZY"
# From the slides, we know that we can compute the weights $w$ using the pseudoinverse of $w = X^+y$:
# + colab={} colab_type="code" id="iRYhw-N0j3ZZ" outputId="b6fd116b-99d7-475d-f951-9408f4b41b2f"
w = np.dot(np.linalg.pinv(X), y)
w
# + [markdown] colab_type="text" id="N2SoGsRNj3ZZ"
# The first weight corresponds to the $y$-intercept of the line, which is typically denoted as $b$:
# + colab={} colab_type="code" id="nLvuVmBGj3ZZ" outputId="12b69842-6e34-4e93-b0ac-decb8271efe6"
b = np.asarray(w).reshape(-1)[0]
b
# + [markdown] colab_type="text" id="96XCC8Z-j3Za"
# While the second weight corresponds to the slope of the line, which is typically denoted as $m$:
# + colab={} colab_type="code" id="HHTNUSJCj3Za" outputId="94d24206-e622-4d78-f037-063e9122831d"
m = np.asarray(w).reshape(-1)[1]
m
# + [markdown] colab_type="text" id="lvGCTZRqj3Zc"
# With the weights we can plot the line to confirm it fits the points:
# + colab={} colab_type="code" id="q9SAiUyej3Zc" outputId="e648a5c1-25d1-410d-d801-c61a2c15e944"
fig, ax = plt.subplots()
plt.title(title)
plt.xlabel(xlabel)
plt.ylabel(ylabel)
ax.scatter(x1, y)
x_min, x_max = ax.get_xlim()
y_min, y_max = m*x_min + b, m*x_max + b
ax.set_xlim([x_min, x_max])
_ = ax.plot([x_min, x_max], [y_min, y_max])
# + [markdown] colab_type="text" id="rJnSV2afj3Zd"
# ### The Trace Operator
# + [markdown] colab_type="text" id="dq9uqorvj3Zd"
# Denoted as Tr($A$). Simply the sum of the diagonal elements of a matrix: $$\sum_i A_{i,i}$$
# + colab={} colab_type="code" id="vOdkry9ij3Zd" outputId="ac2b9194-6212-4053-ecef-54d342b6bfa9"
A = np.array([[25, 2], [5, 4]])
A
# + colab={} colab_type="code" id="zwh8-KTNj3Ze" outputId="b146e756-11e4-47b4-e859-b8cdc78b9da8"
25 + 4
# + colab={} colab_type="code" id="7LxzUu37j3Zf" outputId="0dcb4d75-a641-44af-aba2-15f4c94a5a2c"
np.trace(A)
# + [markdown] colab_type="text" id="dUeQKrYMj3Zg"
# The trace operator has a number of useful properties that come in handy while rearranging linear algebra equations, e.g.:
#
# * Tr($A$) = Tr($A^T$)
# * Assuming the matrix shapes line up: Tr(ABC) = Tr(CAB) = Tr(BCA)
# + [markdown] colab_type="text" id="kuKYTjskj3Zg"
# In particular, the trace operator can provide a convenient way to calculate a matrix's Frobenius norm: $$||A||_F = \sqrt{\mathrm{Tr}(AA^\mathrm{T})}$$
# + [markdown] colab_type="text" id="JcqTZnimj3Zg"
# **Exercise**
#
# Using the matrix `A_p`:
#
# 1. Identify the PyTorch trace method and the trace of the matrix.
# 2. Further, use the PyTorch Frobenius norm method (for the left-hand side of the equation) and the trace method (for the right-hand side of the equation) to demonstrate that $||A||_F = \sqrt{\mathrm{Tr}(AA^\mathrm{T})}$
# + colab={} colab_type="code" id="rQhYWOFvj3Zg" outputId="bc8c70ae-d1f9-4e7c-ee18-a6112c8b8096"
A_p
# + [markdown] colab_type="text" id="QXuZgAUgj3Zh"
# **Return to slides here.**
# + [markdown] colab_type="text" id="1rQOXPyaj3Zh"
# ### Principal Component Analysis
# + [markdown] colab_type="text" id="X3_Etgo4j3Zh"
# This PCA example code is adapted from [here](https://jupyter.brynmawr.edu/services/public/dblank/CS371%20Cognitive%20Science/2016-Fall/PCA.ipynb).
# + colab={} colab_type="code" id="ubl3WdRWj3Zh"
from sklearn import datasets
iris = datasets.load_iris()
# + colab={} colab_type="code" id="ZE2yvfEbj3Zi" outputId="7b6d1699-db20-4877-b648-1c2c10d2d0ce"
iris.data.shape
# + colab={} colab_type="code" id="fa9fcMl2j3Zi" outputId="365fbdc9-b667-4def-85c6-d361b6038fdc"
iris.get("feature_names")
# + colab={} colab_type="code" id="8O9xwrOLj3Zj" outputId="893e01f9-2886-4e18-f312-b5d393b49d1a"
iris.data[0:6,:]
# + colab={} colab_type="code" id="YoodmvRsj3Zj"
from sklearn.decomposition import PCA
# + colab={} colab_type="code" id="PcJwICbtj3Zk"
pca = PCA(n_components=2)
# + colab={} colab_type="code" id="bNb6txoIj3Zk"
X = pca.fit_transform(iris.data)
# + colab={} colab_type="code" id="plS7skQGj3Zl" outputId="bb83578c-2310-4c32-edaa-6728d4dd063d"
X.shape
# + colab={} colab_type="code" id="wC_j-7Xyj3Zl" outputId="c4c4f893-7c7c-4405-ab00-6d1698ed7863"
X[0:6,:]
# + colab={} colab_type="code" id="O_aNxFn5j3Zm" outputId="a26eaac4-92af-4cc4-a419-dc49e255a01b"
plt.scatter(X[:, 0], X[:, 1])
# + colab={} colab_type="code" id="bTck5c93j3Zm" outputId="c6f22246-3117-4be9-dd5c-ac0f53c1851a"
iris.target.shape
# + colab={} colab_type="code" id="IzGhB6NTj3Zn" outputId="7f1b1fec-b7f6-4779-adf6-331765856e1f"
iris.target[0:6]
# + colab={} colab_type="code" id="DQ8oRWsWj3Zn" outputId="26174c04-7ae5-4763-f958-fa111b20ad50"
unique_elements, counts_elements = np.unique(iris.target, return_counts=True)
np.asarray((unique_elements, counts_elements))
# + colab={} colab_type="code" id="VAIoVTYWj3Zo" outputId="438276eb-4a39-41dc-8733-8bad0a51f9d1"
list(iris.target_names)
# + colab={} colab_type="code" id="JlZX_2vQj3Zo" outputId="c80aad27-9433-4662-e836-89f2f4b28e39"
plt.scatter(X[:, 0], X[:, 1], c=iris.target)
# + [markdown] colab_type="text" id="w1Y8YA2oj3Zp"
# **Return to slides here.**
# + colab={} colab_type="code" id="8i9d2Li1j3Zp"
|
notebooks/2-linear-algebra-ii.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using dominant colors to determine genre
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# +
from matplotlib import image
from scipy.cluster.vq import whiten
from scipy.cluster.vq import kmeans
from timeit import default_timer as timer
def get_dominant_color(id):
start = timer()
img = image.imread(r'../Data/Images/' + str(id) + '.jpg')
np_img = np.reshape(img, (img.shape[0]*img.shape[1], img.shape[2]))
np_flat_img = whiten(np_img)
cluster_centers, distortion = kmeans(np_flat_img, 2)
std_red = np_img[:, 0].std()
std_green = np_img[:, 1].std()
std_blue = np_img[:, 0].std()
results = {}
for index, cluster_center in enumerate(cluster_centers):
int_color = int('{:02x}{:02x}{:02x}'.format(
int(cluster_center[0] * red_std),
int(cluster_center[1] * green_std),
int(cluster_center[2] * blue_std)), 16)
key = 'color_{}'.format(index)
results[key] = int_color
end = timer()
print('{0}: {1:.2f}s'.format(id, end - start))
return results
# -
movies = pd.read_csv(r'../Data/movies_df.csv', index_col=0)
temp = movies[:50].apply(lambda x: get_dominant_color(x['id']), axis=1, result_type='expand')
df = pd.merge(movies, temp, left_index=True, right_index=True, )
df.head()
sns.pairplot(df, vars=['color_0', 'color_1'], hue='Drama')
from sklearn.model_selection import train_test_split
df.columns
df.describe()
y = df[['Action']]
X = df[['color_0', 'color_1']]
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, stratify=y)
from sklearn.multiclass import OneVsRestClassifier
from sklearn.svm import SVC
model = OneVsRestClassifier(SVC(gamma='auto'))
model.fit(X_train,y_train)
predictions = model.predict(X_test)
from sklearn.metrics import confusion_matrix, classification_report
print(classification_report(y_test,predictions))
|
Scripts/DominantColor.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# <img src="img/full-colour-logo-UoB.png" alt="Drawing" style="width: 200px;"/>
#
# # Introduction to Programming for Engineers
#
# ## Python 3
#
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# # 11 Simulation : Grids and Multi-Agent Systems
# ## CLASS MATERIAL
#
# <br> <a href='#Grid-BasedSimulations'>1. Grid-Based Simulations</a>
# <br> <a href='#SpatialVectorization'>2. Spatial Vectorization</a>
# <br><a href='#ReviewExercises'>3. Review Exercises</a>
# + [markdown] slideshow={"slide_type": "slide"}
# ### Lesson Goal
#
# Learn how to construct grid and particle/agent based simulations and visualise the results through animation.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Fundamental programming concepts
#
# - Using numpy arrays to contruct efficient grid and particle based simulations.
# - Create animations of the results (.gif and .mp4 videos)
# + [markdown] slideshow={"slide_type": "slide"}
# Computer simulation can be a useful tool for solving and visualising the behaviour of large collections of agents across multiple timesteps.
#
# __Agents__: individuals that interact with one another and their environment.
#
# Example agents:
# - dicretised areas of an environment having one of a set of states
# - particles in a chemical reaction
# - swarm robots or biological organisms (flocking birds, humans)
# + [markdown] slideshow={"slide_type": "slide"}
# Using `for` loops is one way to cycle through each agent in a collection.
#
# For example, to cycle through every cell of an orthoganol grid, we could use:
#
# ```for x in X:
# for y in Y:
# ...
# ```
#
# However, this is computationally costly and programatically inelegant.
# + [markdown] slideshow={"slide_type": "slide"}
# Python provides useful tools for:
# - quickly and concisely solving the rules governing a simulation for large groups of agents.
# - visualising the results as animations
# + [markdown] slideshow={"slide_type": "slide"}
# <a id='Grid-BasedSimulations'></a>
# # 1. Grid-Based Simulations
#
# <br> <a href='#ExampleWildfire'>1.1 Example : Wildfire</a>
# <br> <a href='#ExampleGameLife'>1.2 Example : Game of Life</a>
#
#
# Orthoganol grids are a popular choice for spatial representation of environments as they allow for fast and easy solutions using numpy arrays.
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# <a id='ExampleWildfire'></a>
# ## 1.1 Example : Wildfire
# We can use a grid to model the spread of a fire.
#
# __Problem setting:__
# - A bounded area represented as an orthoganol grid is covered by either
# - vegetation (`Fuel`)
# - `Clear` ground
# - The centre-most cell is on fire (`Burning`)
#
# The fire spreads with the following rules:
# >If a cell is on fire in timestep $t$:
# >- the cell is cleared in timestep $t+1$
# >- all orthoganol neighbours congaining fuel are `Burning` in timestep $t+1$
#
# <br><img src="img/neighbours.png" alt="Drawing" style="width: 200px;"/>
# + [markdown] slideshow={"slide_type": "slide"}
# Install packages
# imageio
# -
import numpy as np
import imageio
from numpy import ma as ma
# + slideshow={"slide_type": "slide"}
prob = .6 # probability of a cell being fuel
timesteps = 300 # simulation time
height = 100 # grid height
width = 100 # grid width
# + slideshow={"slide_type": "slide"}
# Cell States
C = 0 # Clear
F = 1 # Fuel
B = 2 # Burning
# + slideshow={"slide_type": "slide"}
# array : grid at each timestep
states = np.zeros((timesteps, height, width))
np.random.seed(0) # the same random selection each time
# Create an initial random distribution of clear (0) and fuel (1) cells
states[0] = np.random.choice([C,F], # possible cell value
size=[height, width], # size of array
p=[1-prob,prob]) # probability of each value
# set centre cell on fire
states[0,height//2, width//2] = B
print(states[0])
# + [markdown] slideshow={"slide_type": "slide"}
# The grid has a one cell border that is not updated at each timestep.
#
# ```
# for x in range(1,width-1):
# for y in range(1,height-1):
# ```
#
# <br><img src="img/neighbours.png" alt="Drawing" style="width: 200px;"/>
# -
# Let's forst consider the solution using `for` loops:
# + slideshow={"slide_type": "slide"}
def iterate():
"Cycles through each cell and checks if it should be clear, fuel or burning"
for t in range(1,timesteps): # At every timestep ...
states[t] = states[t-1].copy() # ... make a copy of the previous timestep.
for x in range(1,width-1): # Loop through each cell (x,y value) on the grid
for y in range(1,height-1):
if np.array_equal(states[t-1,x,y], B): # If cell is on fire...
states[t,x,y] = C # ... clear cell
if np.array_equal(states[t-1,x+1,y], F): # ...and set any orthoganol neighbours on fire
states[t,x+1,y] = B
if np.array_equal(states[t-1,x-1,y], F):
states[t,x-1,y] = B
if np.array_equal(states[t-1,x,y+1], F):
states[t,x,y+1] = B
if np.array_equal(states[t-1,x,y-1], F):
states[t,x,y-1] = B
# + slideshow={"slide_type": "slide"}
iterate()
# + slideshow={"slide_type": "slide"}
def make_gif(name):
"Crops the simulation to a 200 frames (limit for gif). Saves animation as gif"
# a grid to hold a colour representation of each cell
coloured = np.zeros((timesteps, height, width, 3),
dtype=np.uint8)
# Color
for t in range(timesteps): # each timestep
coloured[t] = np.where(states[t].reshape(height, width, 1) == 0,
np.array([139,69,19]).reshape(1,3) ,
coloured[t])
coloured[t] = np.where(states[t].reshape(height, width, 1) == 1,
np.array([0,255,0]).reshape(1,3) ,
coloured[t])
coloured[t] = np.where(states[t].reshape(height, width, 1) == 2,
np.array([255,0,0]).reshape(1,3) ,
coloured[t])
# Crop
cropped = coloured[
:200,
1:height-1,
1:width-1
]
#imageio.mimsave('./img/wildfire.gif', cropped)
imageio.mimsave(name, cropped)
# + slideshow={"slide_type": "slide"}
make_gif('./img/wildfire.gif')
# + [markdown] slideshow={"slide_type": "-"}
# <img src="img/wildfire.gif" alt="Drawing" style="width: 300px;"/>
# + [markdown] slideshow={"slide_type": "slide"}
# Let's revisit the function `iterate`.
#
# Instead of using for loops, we can create a set of rules that allow us to treat the environment as a single array.
#
#
# + slideshow={"slide_type": "slide"}
def iterate():
"Cycles through each cell and checks if it should be clear, fuel or burning"
for t in range(1,timesteps): # At every timestep ...
states[t] = states[t-1].copy() # ... make a copy of the previous timestep.
burn = ma.make_mask( states[t-1]==B )*1 # Make a map of burning cells
print(burn)
if burn.size==1: # If no cells are burning, stop the simulation
break
states[t] = states[t] - burn * 2 # Extinguish any burning cells from previous timestep
# Ignite any orthoganol neightbours of burning cells from previous timestep
states[t,1:-1,1:-1] = states[t,1:-1,1:-1] + (states[t,1:-1,1:-1] *
np.sign(
burn[ :-2, 1:-1] +
burn[1:-1, :-2] +
burn[1:-1, 2:] +
burn[2: , 1:-1]
)
)
# + [markdown] slideshow={"slide_type": "slide"}
# ##### What does this code do?
# Let's look at what each part of the new function does...
# + [markdown] slideshow={"slide_type": "slide"}
# We cycle through each timestep, copying the previous timestep as before:
# ```python
# def iterate():
# "Cycles through each cell : clear, fuel or burning?"
# for t in range(1,timesteps): # At every timestep ...
# states[t] = states[t-1].copy() # ...copy previous timestep
# ```
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ```python
# burn = ma.make_mask(states[t-1]==B)*1 # map of burning cells
# ```
#
# The function `make_mask` makes a mask of an array containing boolean values where a certain condition is true or false.
#
#
# `a = np.array([[2, 3],[1,4]])`
#
# `ma.make_mask(a==2)`
#
# __`>> [[True, False],[False, False]]`__
#
# `ma.make_mask(a > 1) * 1`
#
# __`>> [[1, 1],[0, 1]]`__
#
#
#
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ```
# if burn.size==1: # If no cells are burning, stop the simulation
# break
# ```
#
# If the condition is not True for any cell in the grid, `ma.make_map` will return a single value, 0.
#
# There are several ways to deal with this, one way is to `break` out of the loop.
# + [markdown] slideshow={"slide_type": "slide"}
# Elementwise subtraction of the map of burning cells from the grid.
#
# `# Extinguish any burning cells from previous timestep
# states[t] = states[t] - burn * 2 `
# + [markdown] slideshow={"slide_type": "slide"}
# The sum of the four orthogonal neighbours of each cell can be found by finding the sum of the arrays one cell to the left, right, top and bottom of the region of interest `states[t,1:-1,1:-1` .
#
# `burn[ :-2, 1:-1] + burn[1:-1, :-2] + burn[1:-1, 2:] + burn[2:,1:-1]`
# + [markdown] slideshow={"slide_type": "-"}
# <img src="img/vectorize_array.png" alt="Drawing" style="width: 500px;"/>
# + [markdown] slideshow={"slide_type": "slide"}
# ```
# # Ignite any orthoganol neightbours of burning cells from previous timestep
# states[t,1:-1,1:-1] = (states[t,1:-1,1:-1 +
# (states[t,1:-1,1:-1] *
# np.sign(
# burn[ :-2, 1:-1] +
# burn[1:-1, :-2] +
# burn[1:-1, 2:] +
# burn[2: , 1:-1]
# )
# )
# ```
# + [markdown] slideshow={"slide_type": "slide"}
# Let's run the code using the new function:
# + slideshow={"slide_type": "slide"}
import numpy as np
import imageio
prob = .6 # probability of a cell being fuel
timesteps = 300 # simulation time
height = 100 # grid height
width = 100 # grid width
# Cell States
C = 0 # Clear
F = 1 # Fuel
B = 2 # Burning
# + slideshow={"slide_type": "slide"}
# # array : grid at each timestep
states = np.zeros((timesteps, height, width))
# Create an initial random distribution of clear (0) and fuel (1) cells
states[0] = np.random.choice([C,F], # possible cell value
size=[height, width], # size of array
p=[1-prob,prob]) # probability of each value
# set centre cell on fire
states[0,height//2, width//2] = B
iterate()
make_gif('./img/wildfire.gif')
# + [markdown] slideshow={"slide_type": "slide"}
# <img src="img/wildfire.gif" alt="Drawing" style="width: 300px;"/>
# + [markdown] slideshow={"slide_type": "slide"}
# This type of simulation is known as a __cellular automata__.
#
# __Cellular Automata__
# <br>A discrete model studied in computer science, mathematics, physics, complexity science, theoretical biology and microstructure modeling.
#
# <br><img src="img/bacteria_compressed.gif
# " alt="Drawing" style="width: 400px;"/>
#
# A regular grid of cells, each in one of a finite number of states, such as on and off.
# + [markdown] slideshow={"slide_type": "slide"}
# <a id='ExampleGameLife'></a>
# ## 1.2 Example : Game of Life
#
#
#
# The best-known example of a cellular automaton.
#
# Devised by the British mathematician <NAME> in 1970.
#
# Zero-player game : evolution is determined by initial state (no input from human players).
# + [markdown] slideshow={"slide_type": "slide"}
# Model space : An infinite two-dimensional orthogonal grid of square cells.
#
# Each cell has two possible states: live or dead.
#
# Every cell interacts with its 8 neighbours : cells that are directly horizontally, vertically, or diagonally adjacent.
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# At each timestep:
#
# - Any live cell with fewer than two live neighbours dies, as if by underpopulation.
# - Any live cell with more than three live neighbours dies, as if by overcrowding.
# - Any live cell with two or three live neighbours lives, unchanged, to the next generation.
# - Any dead cell with exactly three live neighbours becomes a live cell.
# + slideshow={"slide_type": "slide"}
prob = 0.2 # probability of a cell being fuel
timesteps = 300 # simulation time
height = 100 # grid height
width = 100 # grid width
# Cell States
ON = 1
OFF = 0
# + [markdown] slideshow={"slide_type": "slide"}
# Populate the grid, as before
# +
# populate grid with random on/off - more off than on
# # array : grid at each timestep
states = np.zeros((timesteps, height, width))
np.random.seed(0)
# Create an initial random distribution of OFF (0) and ON (1) cells
states[0] = np.random.choice([OFF,ON], # possible cell value
size=[height, width], # size of array
p=[1-prob,prob]) # probability of each value
print(states[0])
# + [markdown] slideshow={"slide_type": "slide"}
# We need to update the `iterate` function to be run at each timestep.
#
# There are two key differences from the last simulation.
#
# 1. The game is played on an infinite surface : a toroid
# <br><img src="img/torus.gif" alt="Drawing" style="width: 300px;"/>
#
# 2. Different rules to determine the next state
# + [markdown] slideshow={"slide_type": "slide"}
# In contrast to the previous example (which had a bounding border of one cell):
#
# - The upper neighbour of a cell in the top row is in the bottom row
# - The lower neighbour of a cell in the bottom row is in the top row
# - The left neighbour of a cell in the furtherst left column is in the furthest right column
# - The right neighbour of a cell in the furtherst right column is in the furthest left column
# + [markdown] slideshow={"slide_type": "slide"}
# We can use the `roll` function which rolls array elements along a given axis.
#
# `np.roll( array,
# number of places to shift,
# axis to shift
# )`
# + slideshow={"slide_type": "slide"}
def iterate():
for t in range(1,timesteps): # At every timestep ...
states[t] = states[t-1].copy() # ... make a copy of the previous timestep.
# compute 8-neghbor sum
# using toroidal boundary conditions - x and y wrap around
total = (np.roll(states[t], 1, 0) +
np.roll(states[t], -1, 0) +
np.roll(states[t], 1, 1) +
np.roll(states[t], -1, 1) +
np.roll(np.roll(states[t], 1, 0), 1, 1) +
np.roll(np.roll(states[t], -1, 0), -1, 1) +
np.roll(np.roll(states[t], 1, 0), -1, 1) +
np.roll(np.roll(states[t], -1, 0), 1, 1))
# apply game of life rules
states[t] = np.where((states[t-1] == ON) & ((total < 2) | (total > 3)),
OFF,
states[t])
states[t] = np.where((states[t-1] == OFF) & (total == 3),
ON,
states[t])
# + slideshow={"slide_type": "slide"}
iterate()
# + slideshow={"slide_type": "slide"}
make_gif('./img/game_of_life.gif')
# -
# <img src="img/game_of_life.gif" alt="Drawing" style="width: 300px;"/>
# + [markdown] slideshow={"slide_type": "slide"}
# The game of life has several patterns that oscillate : repeating infinitely until the game is stopped.
#
# The glider is a pattern that travels across the board in Conway's Game of Life.
#
# <img src="img/Animated_glider_emblem.gif" alt="Drawing" style="width: 100px;"/>
#
# The mutation and movement of a "glider".
# + slideshow={"slide_type": "slide"}
# populate grid
states = np.zeros((timesteps, height, width))
glider = [[1, 0, 0],
[0, 1, 1],
[1, 1, 0]]
states[0, :3, :3] = glider
iterate()
make_gif('./img/game_of_life_glider.gif')
# + [markdown] slideshow={"slide_type": "slide"}
# <img src="img/game_of_life_glider.gif" alt="Drawing" style="width: 300px;"/>
# + [markdown] slideshow={"slide_type": "slide"}
# An early question posed about the Game of Life was whether any configurations exist which result in asymptotically unbounded growth.
#
# The following example is one of the most compact configurations which display unbounded growth.
#
# Note : growth is unbounded on an infinite grid. On a torroidal grid growth is unbounded until the edge is reached, after that different results are produced.
# + slideshow={"slide_type": "slide"}
# populate grid
height = 30
width = 40
states = np.zeros((timesteps, height, width))
unbounded = [[1, 1, 1, 0, 1],
[1, 0, 0, 0, 0],
[0, 0, 0, 1, 1],
[0, 1, 1, 0, 1],
[1, 0, 1, 0, 1]]
states[0, 15:20, 18:23] = unbounded
iterate()
make_gif('./img/game_of_life_unbounded.gif')
# + [markdown] slideshow={"slide_type": "slide"}
# <img src="img/game_of_life_unbounded.gif" alt="Drawing" style="width: 300px;"/>
# + [markdown] slideshow={"slide_type": "slide"}
# The earliest known instance of unbounded growth is the "Glider Gun".
#
# It is an oscillating pattern that creates an infinite series of gliders.
# + slideshow={"slide_type": "slide"}
# populate grid
height = 100
width = 100
states = np.zeros((timesteps, height, width))
glider_gun =\
[[0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0],
[0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,1,0,0,0,0,0,0,0,0,0,0,0],
[0,0,0,0,0,0,0,0,0,0,0,0,1,1,0,0,0,0,0,0,1,1,0,0,0,0,0,0,0,0,0,0,0,0,1,1],
[0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,1,0,0,0,0,1,1,0,0,0,0,0,0,0,0,0,0,0,0,1,1],
[1,1,0,0,0,0,0,0,0,0,1,0,0,0,0,0,1,0,0,0,1,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0],
[1,1,0,0,0,0,0,0,0,0,1,0,0,0,1,0,1,1,0,0,0,0,1,0,1,0,0,0,0,0,0,0,0,0,0,0],
[0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,1,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0],
[0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0],
[0,0,0,0,0,0,0,0,0,0,0,0,1,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0]]
states[0, 1:10, 1:37] = glider_gun
iterate()
make_gif('./img/game_of_life_glider_gun.gif')
# + [markdown] slideshow={"slide_type": "slide"}
# <img src="img/game_of_life_glider_gun.gif" alt="Drawing" style="width: 300px;"/>
# + [markdown] slideshow={"slide_type": "slide"}
# By applying different:
# - sets of rules
# - initial conditions
# to the grid, systems such as chemical reactions, population dynamics and biological systems such as neurons and patterning on shells/skin.
# + [markdown] slideshow={"slide_type": "slide"}
# We are now going to consider a different method of producing animation that can be applied not only to grid-based simulation but to any figure constructed using `matplotlib`.
# + [markdown] slideshow={"slide_type": "slide"}
# This will be convenient for the simulations of rigid body dynamics that we will study next week.
#
#
# + slideshow={"slide_type": "slide"}
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.animation as animation
from matplotlib.animation import FFMpegWriter
from matplotlib import animation, rc
from IPython.display import HTML
# %matplotlib inline
# + [markdown] slideshow={"slide_type": "slide"}
# These are the steps for creating an animation using the `matplotlib.animation` package:
#
# 1. Variables
# 1. Figure window
# 1. Axes within the figure window.
# 1. Object(s) to animate
# 1. Function (e.g. `animate`) to update simulation as a function of time
# 1. Use the function `animation.FuncAnimation` to create the animation.
# + slideshow={"slide_type": "slide"}
# 1. Variables
prob = 0.2 # probability of a cell being fuel
timesteps = 300 # simulation time
height = 100 # grid height
width = 100 # grid width
# Cell States
ON = 1
OFF = 0
# + slideshow={"slide_type": "slide"}
# populate a single 2D grid
#states = np.zeros((height, width))
np.random.seed(0)
# Create an initial random distribution of clear (0) and fuel (1) cells
states = np.random.choice([OFF,ON], # possible cell value
size=[height, width], # size of array
p=[1-prob,prob]) # probability of each value
print(states[0])
# + slideshow={"slide_type": "slide"}
fig, ax = plt.subplots() # 2,3 Figure + axes
mat = ax.matshow(states) # 4. Object to animate : input array as a colour matrix
# + [markdown] slideshow={"slide_type": "slide"}
# We need to write a function that updates a figure of the grid each time it is called.
#
# The function should have __one input, the frame number__, even if it is not used in the function.
# + slideshow={"slide_type": "slide"}
# 5. Function to update simulation
def iterate(data):
"A function to update the game of life simulation using the neighbour rules"
# Make states a glocal variable
global states
# Copy grid
newState = states.copy()
# Compute 8-neghbor sum
total = (np.roll(newState, 1, 0) +
np.roll(newState, -1, 0) +
np.roll(newState, 1, 1) +
np.roll(newState, -1, 1) +
np.roll(np.roll(newState, 1, 0), 1, 1) +
np.roll(np.roll(newState, -1, 0), -1, 1) +
np.roll(np.roll(newState, 1, 0), -1, 1) +
np.roll(np.roll(newState, -1, 0), 1, 1))
# Apply rules
newState = np.where((states == ON) & ((total < 2) | (total > 3)), OFF, newState)
newState = np.where((states == OFF) & (total == 3), ON, newState)
# Update figure using the set_data function
mat.set_data(newState)
states = newState
# For animation the current figure must be returned
return [mat]
# + [markdown] slideshow={"slide_type": "slide"}
# The function to create the animation `FuncAnimation` takes several arguments, including the `iterate` function.
# +
# 6. Create animation object
ani = animation.FuncAnimation(fig, # The figure object to animate
iterate, # Function to call at each frame
interval=50, # Delay between frames ms (default = 2)
frames=30 # Number of frames
)
# Create the animation
ani
#writer = FFMpegWriter(fps=15, metadata=dict(artist='Me'), bitrate=1800)
ani.save("img/game_of_life_anim.mp4", writer=writer)
# + slideshow={"slide_type": "slide"}
from IPython.display import HTML
HTML(ani.to_jshtml())
# + [markdown] slideshow={"slide_type": "slide"}
# <a id='SpatialVectorization'></a>
# # 2. Spatial Vectorization
#
#
# Where elements share the same computation but interact with only a subgroup of other elements.
#
# This was already the case for the cellular automata examples.
#
# In more realistic situations there is an added difficulty because the subgroup is often dynamic and needs to be updated at each iteration:
# - particle systems where particles interact mostly with local neighbours.
# - "boids" that simulate flocking behaviors.
#
# <img src="img/flocking.gif" alt="Drawing" style="width: 300px;"/>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Example : Boids
#
# An artificial life program, developed by <NAME> in 1986, which simulates the flocking behaviour of birds.
#
# The name "boid" corresponds to a shortened version of "bird-oid object", which refers to a bird-like object.
# + [markdown] slideshow={"slide_type": "slide"}
# Boids is an example of emergent behavior; the complexity of Boids arises from the interaction of individual agents (the boids).
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# Interaction is governed by a set of simple rules that determine the boids acceleration and velocity:
#
# - __separation__: steer to avoid crowding local flock-mates
# - __alignment__: steer towards the average heading of local flock-mates
# - __cohesion__: steer to move toward the average position (center of mass) of local flock-mates
#
#
# <img src="img/boids.png" alt="Drawing" style="width: 400px;"/>
# + slideshow={"slide_type": "slide"}
import numpy as np
# + slideshow={"slide_type": "slide"}
boid_count = 10 # number of boids
# + slideshow={"slide_type": "slide"}
height = 2000
width = 2000
limits = np.array([height, width]) # define edges of simulation environment
# + slideshow={"slide_type": "slide"}
np.random.seed(0)
# create 2 x boid_count array of random values in range [0,1]
positions = np.random.rand(2, boid_count)
# Mulitply by size of environment to get random poitions
positions *= limits[:, np.newaxis]
# initial position of each boid
positions
# + [markdown] slideshow={"slide_type": "slide"}
# #### What does this code do?
#
# The `newaxis` expression is used to increase the dimension of the existing array by one more dimension
#
# `limits[:, np.newaxis] `
#
#
# This reshapes the 1-dimensional array `limits` to allow the following multiplication to take place.
#
# Multiply a 2×10 array by a 2×1 array – and get a 2×10 array.
# <br>`positions = np.random.rand(2, boid_count) * limits[:, np.newaxis] `
#
# (In this case, it is has the same result as `limits[:].reshape(2,1)`)
# + [markdown] slideshow={"slide_type": "slide"}
# \begin{align*}
# p &= p_{min} + D(p_{max} - p_{min})\\
# \begin{bmatrix} p_x \\ p_y \end{bmatrix}
# &= \begin{bmatrix} p_{x,min} \\ p_{y,min} \end{bmatrix}
# + D
# \left(
# \begin{bmatrix} p_{x,max} \\ p_{y,max} \end{bmatrix} -
# \begin{bmatrix} p_{x,min} \\ p_{y,min} \end{bmatrix}
# \right)
# \end{align*}
#
# where $D$ is a random number in range 0 to 1
# -
def new_flock(count, minimum, maximum):
return (minimum[:, np.newaxis] +
np.random.rand(2, count) *
(maximum - minimum)[:, np.newaxis])
# + [markdown] slideshow={"slide_type": "slide"}
# Let’s assume that we want our initial positions to vary between 100 and 200 in the x axis, and 900 and 1100 in the y axis.
#
# We can generate random positions within these constraints with:
# -
positions = new_flock(boid_count,
np.array([100, 900]),
np.array([200, 1100]))
# + [markdown] slideshow={"slide_type": "slide"}
# Each bird will also need a starting velocity.
#
# Let’s make these random.
#
# Let's let initial x velocities range over [0,10] and the y velocities over [−20,20].
# -
velocities = new_flock(boid_count,
np.array([0, -20]),
np.array([10, 20]))
velocities
# + [markdown] slideshow={"slide_type": "slide"}
# To find the position in the next timestep according to:
#
# $p_t = p_{t-1} + v_{t-1}$
#
# where:
# <br>$p=$ position of each bird
# <br>$v=$ velocity of each bird
# -
positions += velocities
# + [markdown] slideshow={"slide_type": "slide"}
# ### Animate
# Now we can animate our Boids, again by using the matplotlib animation tools.
# + slideshow={"slide_type": "slide"}
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.animation as animation
from matplotlib.animation import FFMpegWriter
from matplotlib import animation, rc
from IPython.display import HTML
# %matplotlib inline
# + [markdown] slideshow={"slide_type": "slide"}
# These are the steps for creating an animation using the `matplotlib.animation` package:
#
# 1. Variables
# 1. Figure window
# 1. Axes within the figure window.
# 1. Object(s) to animate
# 1. Function (e.g. `animate`) to update simulation as a function of time
# 1. Use the function `animation.FuncAnimation` to create the animation.
# + slideshow={"slide_type": "slide"}
# 1. Variables : define positions and velocities
positions = new_flock(100, np.array([100, 900]), np.array([200, 1100]))
velocities = new_flock(100, np.array([0, -20]), np.array([10, 20]))
# 2. Figure window
figure = plt.figure()
# 3. Axes within the figure window
axes = plt.axes(xlim=(0, limits[0]), # choose the figure size
ylim=(0, limits[1]))
# 4. Object to animate : scatter plot
scatter = axes.scatter(positions[0, :], # x values
positions[1, :], # y values
marker='o',
edgecolor='k',
lw=0.5)
# + [markdown] slideshow={"slide_type": "slide"}
# The function to run to update each frame of the animation:
# +
def update_boids(positions, velocities):
"Updates the position of each boid"
positions += velocities
def animate(frame):
update_boids(positions, velocities)
# change the positions in the figure
scatter.set_offsets(positions.transpose())
# + [markdown] slideshow={"slide_type": "slide"}
# Create the animation using `FuncAnimation`:
# -
anim = animation.FuncAnimation(figure, # The figure object to animate
animate, # Function to call at each frame
frames=50, # Delay between frames ms (default = 2)
interval=50) # Number of frames
# + slideshow={"slide_type": "slide"}
anim.save('img/boids_basic.mp4')
# + slideshow={"slide_type": "slide"}
from IPython.display import HTML
HTML(anim.to_jshtml())
# + [markdown] slideshow={"slide_type": "slide"}
# ### Cohesion
#
# Boids try to fly towards the centre of mass of the group
# +
positions = new_flock(4, np.array([100, 900]), np.array([200, 1100]))
velocities = new_flock(4, np.array([0, -20]), np.array([10, 20]))
COM = np.mean(positions, 1) # mean x and y position (along row axis)
positions.shape
# + [markdown] slideshow={"slide_type": "slide"}
# Find the x and y components of the vector from each position to the middle.
# -
cohesion = positions - COM[:, np.newaxis]
# + [markdown] slideshow={"slide_type": "slide"}
# Substract the velocity pulling the boid towarsd the COM from the total velcoity of each individual.
# +
cohesion_strength = 0.01
velocities = velocities - (cohesion * cohesion_strength)
# + [markdown] slideshow={"slide_type": "slide"}
# Let’s update our function, and animate that:
# +
def update_boids(positions, velocities):
# Cohesion
cohesion_strength = 0.01 # cohesion parameter
COM = np.mean(positions, 1) # centre of mass of flock
cohesion = positions - COM[:, np.newaxis] # distance from each boid to COM
velocities -= cohesion * cohesion_strength # update velocities
positions += velocities # update positions
def animate(frame):
update_boids(positions, velocities)
scatter.set_offsets(positions.transpose())
# + slideshow={"slide_type": "slide"}
positions = new_flock(4, np.array([100, 900]), np.array([200, 1100]))
velocities = new_flock(4, np.array([0, -20]), np.array([10, 20]))
anim = animation.FuncAnimation(figure,
animate,
frames=200,
interval=50)
# + slideshow={"slide_type": "slide"}
anim.save('img/boids_cohesion.mp4')
# + slideshow={"slide_type": "slide"}
HTML(anim.to_jshtml())
# + [markdown] slideshow={"slide_type": "slide"}
# ### Seperation
# Avoid collisions with nearby flockmates
# + [markdown] slideshow={"slide_type": "slide"}
# We want to find the distance from every boid to every other boid (including itself).
#
# We can use broadcasting to subtract every value in an array from every value an array.
#
# Example:
# + slideshow={"slide_type": "slide"}
a = np.array([1, 2, 3, 4])
a
# + slideshow={"slide_type": "slide"}
a[np.newaxis, :]
# -
a[:, np.newaxis]
a[np.newaxis, :] - a[:, np.newaxis]
# + slideshow={"slide_type": "-"}
(a[np.newaxis, :] - a[:, np.newaxis]).shape
# + [markdown] slideshow={"slide_type": "slide"}
# We can subtract all x,y positions...
# -
positions[:, np.newaxis, :].shape
# + [markdown] slideshow={"slide_type": "slide"}
# ...from all x,y positions
# -
positions[:, :, np.newaxis].shape
# + slideshow={"slide_type": "slide"}
separations = positions[:, np.newaxis, :] - positions[:, :, np.newaxis]
# -
separations.shape
# + slideshow={"slide_type": "slide"}
positions = new_flock(4, np.array([100, 900]), np.array([200, 1100]))
velocities = new_flock(4, np.array([0, -20]), np.array([10, 20]))
# + [markdown] slideshow={"slide_type": "slide"}
# Find the elementwise sum of squared x and y value for each distance
# -
sum_square_dis = np.sum(separations**2, 0)
sum_square_dis.shape
# + slideshow={"slide_type": "slide"}
alert_distance = 2000
close_birds = sum_square_dis < alert_distance
close_birds
# + [markdown] slideshow={"slide_type": "slide"}
# Find the direction distances only to those birds which are too close:
# +
separations_if_close = np.copy(separations) # copy of all distances
far_map = np.logical_not(close_birds) # map of far away birds
far_map
# + [markdown] slideshow={"slide_type": "slide"}
# Set x and y values in separations_if_close to zero if they are far away:
# -
separations_if_close[0, :, :][far_map] = 0 # x values
separations_if_close[1, :, :][far_map] = 0 # y values
separations_if_close
separations_if_close.shape
# + [markdown] slideshow={"slide_type": "slide"}
# Sum of x distance and y distance to closest boids for each boid
# -
np.sum(separations_if_close, 2)
# + [markdown] slideshow={"slide_type": "slide"}
# Add the velocity seperating the boids to the total velcoity of each individual.
# -
velocities = velocities + np.sum(separations_if_close, 2)
# + [markdown] slideshow={"slide_type": "slide"}
# Let's update the `update_boids` function to include the seperation rule:
# +
def update_boids(positions, velocities):
# Cohesion
cohesion_strength = 0.01 # cohesion parameter
COM = np.mean(positions, 1) # centre of mass of flock
cohesion = positions - COM[:, np.newaxis] # distance from each boid to COM
velocities -= cohesion * cohesion_strength # update velocities
# Seperation
separations = positions[:, np.newaxis, :] - positions[:, :, np.newaxis] # all distances between boids
sum_squared_dis = np.sum(separations**2, 0) # sum of squared distances
alert_distance = 100 # threshold for seperation
far_map = sum_squared_dis > alert_distance # boolean map of far boids
separations_if_close = np.copy(separations) # copy of all distances between boids
separations_if_close[0, :, :][far_map] = 0 # remove far boids
separations_if_close[1, :, :][far_map] = 0
velocities += np.sum(separations_if_close, 1) # update velocity of boids
positions += velocities # update positions
def animate(frame):
update_boids(positions, velocities)
scatter.set_offsets(positions.transpose())
# + slideshow={"slide_type": "slide"}
positions = new_flock(100, np.array([100, 900]), np.array([200, 1100]))
velocities = new_flock(100, np.array([0, -20]), np.array([10, 20]))
anim = animation.FuncAnimation(figure,
animate,
frames=200,
interval=50)
# + slideshow={"slide_type": "slide"}
anim.save('img/boids_seperation.mp4')
HTML(anim.to_jshtml())
# + [markdown] slideshow={"slide_type": "slide"}
# ### Alignment
#
# Attempt to match velocity with nearby flockmates
# + slideshow={"slide_type": "slide"}
def update_boids(positions, velocities):
# Cohesion
cohesion_strength = 0.01 # cohesion parameter
COM = np.mean(positions, 1) # centre of mass of flock
cohesion = positions - COM[:, np.newaxis] # distance from each boid to COM
velocities -= cohesion * cohesion_strength # update velocities
# Seperation
separations = positions[:, np.newaxis, :] - positions[:, :, np.newaxis] # all distances between boids
sum_squared_dis = np.sum(separations**2, 0) # sum of squared distances
alert_distance = 100 # threshold for seperation
far_map = sum_squared_dis > alert_distance # boolean map of far boids
separations_if_close = np.copy(separations) # copy of all distances between boids
separations_if_close[0, :, :][far_map] = 0 # remove far boids
separations_if_close[1, :, :][far_map] = 0
velocities += np.sum(separations_if_close, 1) # update velocity of boids
# Alignment
velocity_dif = velocities[:, np.newaxis, :] - velocities[:, :, np.newaxis] # all differences in velocity between boids
form_flying_distance = 10000 # threshold for formation flying
form_flying_strength = 0.125 # formation flying parameter
very_far_map = sum_squared_dis > form_flying_distance # boolean map of very far boids
velocity_dif_if_close = np.copy(velocity_dif) # copy of all differences in velocity between boids
velocity_dif_if_close[0, :, :][very_far_map] = 0 # remove very far boids
velocity_dif_if_close[1, :, :][very_far_map] = 0
velocities -= np.mean(velocity_dif_if_close, 1) * form_flying_strength # update velocity of boids
positions += velocities # update positions
# + slideshow={"slide_type": "slide"}
def animate(frame):
update_boids(positions, velocities)
scatter.set_offsets(positions.transpose())
# + slideshow={"slide_type": "slide"}
positions = new_flock(100, np.array([100, 900]), np.array([200, 1100]))
velocities = new_flock(100, np.array([0, -20]), np.array([10, 20]))
anim = animation.FuncAnimation(figure,
animate,
frames=400,
interval=50)
# + slideshow={"slide_type": "slide"}
anim.save('img/boids_alignment.mp4')
HTML(anim.to_jshtml())
# + [markdown] slideshow={"slide_type": "slide"}
# Our boids eventually travel outside of the plot window.
#
# Like the previous simulations we can setconditions that effect the boundary of the simulation.
#
# In the boids simulation, the boids avoid the walls by *steering*, heading away from the wall on a heading normal to it's surface.
#
# The wall does not provide a physical boundary in this case.
#
# The boid can, for example, fly through the boundary.
# + [markdown] slideshow={"slide_type": "slide"}
# We define parameters for the distance from the wall at which we want a repulsive force to act.
# -
wall_alert_distance = 1
wall_strength = 1
# + slideshow={"slide_type": "slide"}
positions = new_flock(10, np.array([100, 900]), np.array([200, 1100]))
velocities = new_flock(10, np.array([0, -20]), np.array([10, 20]))
# + slideshow={"slide_type": "slide"}
right = np.copy(positions[0, :]) - width # x distance from roght wall
far_map = right < wall_alert_distance # map of x vals far from wall
right[far_map] = 0 # remove boids far from wall
velocities[0, :] += right * -wall_strength # update velocity of boids close to wall
# + slideshow={"slide_type": "slide"}
def update_boids(positions, velocities):
# Cohesion
cohesion_strength = 0.01 # cohesion parameter
COM = np.mean(positions, 1) # centre of mass of flock
cohesion = positions - COM[:, np.newaxis] # distance from each boid to COM
velocities -= cohesion * cohesion_strength # update velocities
# Seperation
separations = positions[:, np.newaxis, :] - positions[:, :, np.newaxis] # all distances between boids
sum_squared_dis = np.sum(separations**2, 0) # sum of squared distances
alert_distance = 100 # threshold for seperation
far_map = sum_squared_dis > alert_distance # boolean map of far boids
separations_if_close = np.copy(separations) # copy of all distances between boids
separations_if_close[0, :, :][far_map] = 0 # remove far boids
separations_if_close[1, :, :][far_map] = 0
velocities += np.sum(separations_if_close, 1) # update velocity of boids
# Alignment
velocity_dif = velocities[:, np.newaxis, :] - velocities[:, :, np.newaxis] # all differences in velocity between boids
form_flying_distance = 10000 # threshold for formation flying
form_flying_strength = 0.125 # formation flying parameter
very_far_map = sum_squared_dis > form_flying_distance # boolean map of very far boids
velocity_dif_if_close = np.copy(velocity_dif) # copy of all differences in velocity between boids
velocity_dif_if_close[0, :, :][very_far_map] = 0 # remove very far boids
velocity_dif_if_close[1, :, :][very_far_map] = 0
velocities -= np.mean(velocity_dif_if_close, 1) * form_flying_strength # update velocity of boids
# Simulate wall
wall_alert_distance = 1
wall_strength = 3
left = np.copy(positions[0, :]) # x distance from left wall
far_map = left > wall_alert_distance # map of x vals far from left wall
left[far_map] = 0 # remove boids far from wall
velocities[0, :] += left * -wall_strength # update velocity of boids close to wall
right = np.copy(positions[0, :]) - width # x distance from right wall
far_map = right < wall_alert_distance # map of x vals far from wall
right[far_map] = 0 # remove boids far from wall
velocities[0, :] += right * -wall_strength # update velocity of boids close to wall
top = np.copy(positions[1, :])
far_map = top > wall_alert_distance
top[far_map] = 0
velocities[1, :] += top * -wall_strength
bottom = np.copy(positions[1, :]) - height
far_map = bottom < wall_alert_distance
bottom[far_map] = 0
velocities[1, :] += bottom * -wall_strength
positions += velocities # update positions
def animate(frame):
update_boids(positions, velocities)
scatter.set_offsets(positions.transpose())
# + slideshow={"slide_type": "slide"}
# bounce off right wall
positions = new_flock(100, np.array([500, 900]), np.array([700, 1100]))
velocities = new_flock(100, np.array([0, -20]), np.array([10, 20]))
anim.save('img/boids_rightbounce.mp4')
HTML(anim.to_jshtml())
# + slideshow={"slide_type": "slide"}
# bounce off left wall
positions = new_flock(100, np.array([500, 900]), np.array([700, 1100]))
velocities = new_flock(100, np.array([0, -20]), np.array([10, 20])) # right bounce
anim.save('img/boids_leftbounce.mp4')
HTML(anim.to_jshtml())
# + slideshow={"slide_type": "slide"}
# bounce off top wall
positions = new_flock(100, np.array([500, 900]), np.array([700, 1100]))
velocities = new_flock(100, np.array([-20, -10]), np.array([20, 0])) # bottom bounce
anim.save('img/boids_topbounce.mp4')
HTML(anim.to_jshtml())
# + slideshow={"slide_type": "slide"}
# bounce off top wall
positions = new_flock(100, np.array([500, 900]), np.array([700, 1100]))
velocities = new_flock(100, np.array([-20, 0]), np.array([20, 10])) # top bounce
anim.save('img/boids_bottombounce.mp4')
HTML(anim.to_jshtml())
# + slideshow={"slide_type": "slide"}
# random direction
positions = new_flock(100, np.array([500, 900]), np.array([700, 1100]))
velocities = new_flock(100, np.array([-40, -60]), np.array([40, 50])) # top bounce
anim = animation.FuncAnimation(figure,
animate,
frames=200,
interval=20)
anim.save('img/boids_alignment.mp4')
HTML(anim.to_jshtml())
# + [markdown] slideshow={"slide_type": "slide"}
# To add more complex environemnts and behaviours, we need a new appraoch that allows us to handle large numbers of individuals.
#
# <img src="img/boids01.gif" alt="Drawing" style="width: 300px;"/>
#
# This will be the topic of next weeek's class.
# + [markdown] slideshow={"slide_type": "slide"}
# By creating layers of numpy arrays you can produce simulate complex and realistic behaviour quickly without requiring much memory.
#
# For example, you could introduce another array `accelerations`:
# - gravity acting on every particle
# - acceleration due to a collision
# + [markdown] slideshow={"slide_type": "slide"}
# # Summary
# - Grid-based simulations are a fast and computationally light way to model complex and rule-based systems and environments.
# - Numpy arrays are a useful way to represent a grid-based simulation due to built-in elementwise operations and broadcasting.
# - Python provides a number of ways for creating animated visualisations of simulation data :
# - capturing an RGB colour array as a .gif file
# - using the `matplotlib.animation` package to animate a figure
#
# - Python arrays are can also be useful way to represent large collections of interacting (simple!) agents.
#
# <br><img src="img/Reaction-Diffusion_3D.gif
# " alt="Drawing" style="width: 200px;"/>
# + [markdown] slideshow={"slide_type": "slide"}
# <a id='ReviewExercises'></a>
# # 3. Review Exercises
# + [markdown] slideshow={"slide_type": "slide"}
# # Review Exercise 1 : Game of Life
#
# Create recreating the game of life example.
#
# Try populating the grid with different starting conditions (you can find some famous initial conditions by searching the internet).
#
# Try changing the rules of the game.
#
# Save the results of your simulation as an animation.
# + [markdown] slideshow={"slide_type": "slide"}
# # Review Exercise 2 : Particles in a box
#
# In the boids simulation, the boids avoid the walls by *steering*, heading away from the wall on a heading normal to it's surface.
#
# The wall does not provide a physical boundary in this case.
#
# The boid can, for example, fly through the boundary.
# + [markdown] slideshow={"slide_type": "slide"}
# Let's change the problem setting slightly.
#
# - 10 particles
# - random starting position
# - random starting velocity
# - particles can occupy the same space (ignore collisions with other particles)
# - if a particle collides with the simulation boundary, it bounces off...
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# The basic principle when bouncing is that the incoming angle shoudl be equal to the outgong angle.
#
# <img src="img/ball.png" alt="Drawing" style="width: 300px;"/>
# + [markdown] slideshow={"slide_type": "slide"}
# We can buid a simple kinematic model of the particle collision by neglecting forces (e.g. friction):
# - if a particle hits a surface parallel to x axis, $v_y = v_y \times - 1$#
# - if a particle hits a surface parallel to y axis, $v_x = v_x \times - 1$
# + [markdown] slideshow={"slide_type": "slide"}
# Build a simulation of the problem setting described.
#
# Save an animation of your simulation.
# + slideshow={"slide_type": "slide"}
import io
import base64
from IPython.display import HTML
video = io.open('img/bouncing_particles.mp4', 'r+b').read()
encoded = base64.b64encode(video)
HTML(data='''<video width = "500" alt="test" controls>
<source src="data:video/mp4;base64,{0}" type="video/mp4" />
</video>'''.format(encoded.decode('ascii')))
# + slideshow={"slide_type": "slide"}
|
11_Simulation1__ClassMaterial.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="LcO5BFI4I7HB"
# # Deoxyfluorination with Sulfonyl Fluorides: Navigating Reaction Space with Machine Learning
#
# DOI: 10.1021/jacs.8b01523
#
# <NAME>. Soc. 2018, 140, 5004−5008
#
# ## Defining protos for reaction data in Figure 1
# + [markdown] colab_type="text" id="FkGs9wsG6hbC"
# Colab set-up: install schema
# + [markdown] colab_type="text" id="QpV2Y-uBQJog"
# Import schema and helper functions
# + colab={} colab_type="code" id="WNYlhmmX64wp"
from datetime import datetime
from ord_schema.proto import reaction_pb2
from ord_schema.proto import dataset_pb2
from ord_schema.units import UnitResolver
from ord_schema import validations
from ord_schema import message_helpers
unit_resolver = UnitResolver()
# + [markdown] colab_type="text" id="rbVTyluBQNZI"
# # Define exemplary reaction
#
# This will be a single prototypical reaction from Table 1. We will copy it and use it as a template for filling out the full results table.
# + colab={"base_uri": "https://localhost:8080/", "height": 85} colab_type="code" id="kX_Pp3fA7UWB" outputId="487cf4bc-b168-4328-c75b-8f9581a46164"
reaction = reaction_pb2.Reaction()
reaction.identifiers.add(value=r'deoxyfluorination', type='NAME')
reaction
# + [markdown] colab_type="text" id="ameuJjZIQUl0"
# Define reaction inputs
# - The first input is a pre-mixed stock solution of an alcohol in THF (1a for the prototype reaction)
# - The second input is a pre-mixed stock solution of a sulfonyl fluoride (4-chlorobenzenesulfonyl fluoride for the prototype reaction)
# - The third addition is a pure base (DBU for the prototype reaction)
# + colab={} colab_type="code" id="-tHbGCMiyhE3"
# Input 1a: 0.1 mmol alcohol in 125 uL THF
input_1a = reaction.inputs['alcohol in THF']
input_1a.addition_order = 1
solute = input_1a.components.add()
solvent = input_1a.components.add()
solute.reaction_role = reaction_pb2.ReactionRole.REACTANT
solute.identifiers.add(value=r'c1ccccc1CCC(O)C', type='SMILES')
solute.amount.moles.CopyFrom(unit_resolver.resolve('0.1 mmol'))
solute.preparations.add().type = reaction_pb2.CompoundPreparation.PreparationType.NONE
solute.is_limiting = True
solvent.reaction_role = reaction_pb2.ReactionRole.SOLVENT
solvent.identifiers.add(value=r'THF', type='NAME')
solvent.identifiers.add(value=r'C1CCCO1', type='SMILES')
solvent.amount.volume.CopyFrom(unit_resolver.resolve('125 uL'))
solvent.preparations.add().type = reaction_pb2.CompoundPreparation.PreparationType.DRIED
## Note: more lengthy way to specify volume w/o unit resolver
# solvent.volume.value = 125
# solvent.volume.units = reaction_pb2.Volume.MICROLITER
# + colab={} colab_type="code" id="YS5VpUzn3AK-"
# Input sf: 0.11 mmol sulfonyl fluoride in 125 uL THF
input_sf = reaction.inputs['sulfonyl fluoride']
input_sf.addition_order = 2
solute = input_sf.components.add()
solvent = input_sf.components.add()
solute.reaction_role = reaction_pb2.ReactionRole.REACTANT
solute.identifiers.add(value=r'4-chlorobenzenesulfonyl fluoride', type='NAME')
solute.identifiers.add(value=r'Clc1ccc(S(=O)(=O)F)cc1', type='SMILES')
solute.amount.moles.CopyFrom(unit_resolver.resolve('0.11 mmol'))
solute.preparations.add().type = reaction_pb2.CompoundPreparation.PreparationType.SYNTHESIZED
solvent.reaction_role = reaction_pb2.ReactionRole.SOLVENT
solvent.identifiers.add(value=r'THF', type='NAME')
solvent.identifiers.add(value=r'C1CCCO1', type='SMILES')
solvent.amount.volume.CopyFrom(unit_resolver.resolve('125 uL'))
solvent.preparations.add().type = reaction_pb2.CompoundPreparation.PreparationType.DRIED
# + colab={} colab_type="code" id="lR1ff-uq71TL"
# Input base: 0.15 mmol pure
input_base = reaction.inputs['base']
input_base.addition_order = 3
base = input_base.components.add()
base.reaction_role = reaction_pb2.ReactionRole.REAGENT
base.identifiers.add(value=r'N\2=C1\N(CCCCC1)CCC/2', type='SMILES')
base.amount.moles.CopyFrom(unit_resolver.resolve('0.15 mmol'))
base.preparations.add().type = reaction_pb2.CompoundPreparation.PreparationType.NONE
base.source.vendor = 'Sigma-Millipore'
# + [markdown] colab_type="text" id="Fuoxwji2RaZA"
# Define reaction setup & conditions
# + colab={} colab_type="code" id="m2Ud6UqK8-LL"
# Reactions performed in 1 mL sealed glass vial
reaction.setup.vessel.CopyFrom(
reaction_pb2.Vessel(
type='VIAL',
material=dict(type='GLASS'),
volume=unit_resolver.resolve('1 mL')
)
)
reaction.setup.vessel.attachments.add(type='CAP')
reaction.setup.is_automated = False
# + colab={} colab_type="code" id="W_mov8JIYQUC"
# No temperature control = ambient conitions
t_conds = reaction.conditions.temperature
t_conds.control.type = t_conds.TemperatureControl.AMBIENT
# + colab={} colab_type="code" id="zygxzC8ziJTr"
# Vials were sealed under ambient conditions
p_conds = reaction.conditions.pressure
p_conds.control.type = p_conds.PressureControl.SEALED
p_conds.atmosphere.type = p_conds.Atmosphere.AIR
# + colab={} colab_type="code" id="TjyZ_Urkix6_"
# Vials contained stir bars at 600 rpm
s_conds = reaction.conditions.stirring
s_conds.type = s_conds.STIR_BAR
s_conds.rate.type = s_conds.StirringRate.HIGH # qualitative
s_conds.rate.rpm = 600
# + [markdown] colab_type="text" id="lNGei36QRlzg"
# No additional safety notes or observations
# + colab={} colab_type="code" id="OZm3Src4i0ro"
# No safety notes
# reaction.notes.safety_notes = ''
# No reaction observations
# observation = reaction.observations.add()
# observation.time =
# observation.comment =
# + colab={} colab_type="code" id="gFDmbbfF3Xjj"
# Only workup is addition of NMR standard
workup = reaction.workups.add(type='ADDITION')
solute = workup.input.components.add()
solvent = workup.input.components.add()
solute.reaction_role = reaction_pb2.ReactionRole.INTERNAL_STANDARD
solute.identifiers.add(value=r'C1=CC=C2C(=C1)C=CC=C2F', type='SMILES')
solute.identifiers.add(value=r'1-fluoronaphthalene', type='NAME')
solute.amount.moles.CopyFrom(unit_resolver.resolve('0.1 mmol'))
solvent.reaction_role = reaction_pb2.ReactionRole.WORKUP
solvent.identifiers.add(value=r'chloroform', type='NAME')
solvent.identifiers.add(value=r'ClC(Cl)Cl', type='SMILES')
solvent.amount.volume.CopyFrom(unit_resolver.resolve('250 uL'))
# + [markdown] colab_type="text" id="_hFXowZCRrW7"
# After 48 hours, the crude reaction mixture is characterized by 19F NMR to estimate the yield
#
# This represents one "outcome" with one characterized "product"
# + colab={} colab_type="code" id="nMZKW4ZSkyeY"
outcome = reaction.outcomes.add()
outcome.reaction_time.CopyFrom(unit_resolver.resolve('48 hrs'))
# Analyses: 19F NMR
outcome.analyses['19f nmr of crude'].type = reaction_pb2.Analysis.NMR_OTHER
outcome.analyses['19f nmr of crude'].details = ('19F NMR using 1 equiv 1-fluoro'
'naphthalene in 250 uL chloroform as internal standard')
outcome.analyses['19f nmr of crude'].instrument_manufacturer = 'Bruker'
# Define product identity
prod_2a = outcome.products.add(is_desired_product=True)
prod_2a.identifiers.add(type='SMILES', value='c1ccccc1CCC(F)C')
prod_2a.reaction_role = reaction_pb2.ReactionRole.PRODUCT
# Define product yield from Figure 1 in the main test
# Note: described in SI, 4.8% is the variance of the yield including variance
# in the experiment itself. NMR yields itself has precision of <= 2%
# accoring to SI Section II.
prod_2a.measurements.add(type='YIELD', analysis_key='19f nmr of crude',
percentage=dict(value=40, precision=4.8), uses_internal_standard=True)
# The "19f nmr of crude" analysis was used to confirm both identity and yield
prod_2a.measurements.add(type='IDENTITY', analysis_key='19f nmr of crude')
# Reaction provenance
reaction.provenance.city = r'Princeton, NJ'
reaction.provenance.doi = r'10.1021/jacs.8b01523'
reaction.provenance.publication_url = r'https://pubs.acs.org/doi/10.1021/jacs.8b01523'
reaction.provenance.record_created.time.value = datetime.now().strftime("%m/%d/%Y, %H:%M:%S")
reaction.provenance.record_created.person.CopyFrom(reaction_pb2.Person(
name='<NAME>', organization='MIT', orcid='0000-0002-8271-8723', email='<EMAIL>'
))
# + [markdown] colab_type="text" id="GRBoi75yS1Gn"
# Validate and examine this final prototypical reaction entry -- note that this is just a single entry from the results table in Figure 1
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="yM9bpRndSpul" outputId="aacf1117-1b5d-41a1-f4e4-33be27f5189e"
validations.validate_message(reaction)
print(reaction)
# + [markdown] colab_type="text" id="riR6W8geS9JI"
# # Define full set of compounds and conditions
#
# We will make extensive use of the ```message_helpers.build_compound``` helper function and define our own custom helper function to use the prototypical reaction example and replace only select entries
# + colab={} colab_type="code" id="YJv51yW-qWBT"
# Defining all major reactants, reagents, and products used in screen
compounds = {
# Reactants
'1a': message_helpers.build_compound(r'c1ccccc1CCC(O)C'),
'2a': message_helpers.build_compound(r'c1ccccc1CCC(F)C'),
'1b': message_helpers.build_compound(r'c1ccccc1-c2ccc(CO)cc2'),
'2b': message_helpers.build_compound(r'c1ccccc1-c2ccc(CF)cc2'),
'1c': message_helpers.build_compound(r'O[C@@H]1C[C@H](OCC2=CC=CC=C2)C1'),
'2c': message_helpers.build_compound(r'F[C@@H]1C[C@H](OCC2=CC=CC=C2)C1'),
'1d': message_helpers.build_compound(r'O[C@H]1C[C@@H](C(OC)=O)N(C(OC(C)(C)C)=O)C1'),
'2d': message_helpers.build_compound(r'F[C@H]1C[C@@H](C(OC)=O)N(C(OC(C)(C)C)=O)C1'),
# Sulfonyl fluorides
'3-Cl': message_helpers.build_compound(r'Clc1ccc(S(=O)(=O)F)cc1'),
'PyFluor': message_helpers.build_compound(r'O=S(C1=CC=CC=N1)(F)=O', name='PyFluor'),
'3-CF3': message_helpers.build_compound(r'O=S(C1=CC=C(C(F)(F)F)C=C1)(F)=O'),
'3-NO2': message_helpers.build_compound(r'O=S(C1=CC=C([N+]([O-])=O)C=C1)(F)=O'),
'PBSF': message_helpers.build_compound(r'C(C(C(F)(F)S(=O)(=O)F)(F)F)(C(F)(F)F)(F)F',
vendor='Acros'),
# Bases
'DBU': message_helpers.build_compound(r'N\2=C1\N(CCCCC1)CCC/2', name='DBU',
vendor='Millipore-Sigma'),
'MTBD': message_helpers.build_compound(r'CN1CCCN2C1=NCCC2', name='MTBD',
vendor='Sigma-Millipore'),
'BTMG': message_helpers.build_compound(r'CC(C)(C)N=C(N(C)C)N(C)C', name='BTMG',
vendor='Millipore-Sigma'),
'BTPP': message_helpers.build_compound(r'CC(C)(C)N=P(N1CCCC1)(N2CCCC2)N3CCCC3',
name='BTPP', vendor='Millipore-Sigma'),
}
# + colab={} colab_type="code" id="_w2k1q_Us-oZ"
# Define yield tables
data = [ # reactant, product, sulfonyl fluoride, base, yield percent
# 2a
('1a', '2a', '3-Cl', 'DBU', 40),
('1a', '2a', '3-Cl', 'MTBD', 54),
('1a', '2a', '3-Cl', 'BTMG', 41),
('1a', '2a', '3-Cl', 'BTPP', 42),
('1a', '2a', 'PyFluor', 'DBU', 57),
('1a', '2a', 'PyFluor', 'MTBD', 59),
('1a', '2a', 'PyFluor', 'BTMG', 49),
('1a', '2a', 'PyFluor', 'BTPP', 53),
('1a', '2a', '3-CF3', 'DBU', 52),
('1a', '2a', '3-CF3', 'MTBD', 69),
('1a', '2a', '3-CF3', 'BTMG', 57),
('1a', '2a', '3-CF3', 'BTPP', 60),
('1a', '2a', '3-NO2', 'DBU', 54),
('1a', '2a', '3-NO2', 'MTBD', 63),
('1a', '2a', '3-NO2', 'BTMG', 55),
('1a', '2a', '3-NO2', 'BTPP', 51),
('1a', '2a', 'PBSF', 'DBU', 39),
('1a', '2a', 'PBSF', 'MTBD', 60),
('1a', '2a', 'PBSF', 'BTMG', 61),
('1a', '2a', 'PBSF', 'BTPP', 65),
# 2b
('1b', '2b', '3-Cl', 'DBU', 11),
('1b', '2b', '3-Cl', 'MTBD', 36),
('1b', '2b', '3-Cl', 'BTMG', 83),
('1b', '2b', '3-Cl', 'BTPP', 92),
('1b', '2b', 'PyFluor', 'DBU', 12),
('1b', '2b', 'PyFluor', 'MTBD', 26),
('1b', '2b', 'PyFluor', 'BTMG', 57),
('1b', '2b', 'PyFluor', 'BTPP', 77),
('1b', '2b', '3-CF3', 'DBU', 17),
('1b', '2b', '3-CF3', 'MTBD', 36),
('1b', '2b', '3-CF3', 'BTMG', 83),
('1b', '2b', '3-CF3', 'BTPP', 99),
('1b', '2b', '3-NO2', 'DBU', 21),
('1b', '2b', '3-NO2', 'MTBD', 41),
('1b', '2b', '3-NO2', 'BTMG', 83),
('1b', '2b', '3-NO2', 'BTPP', 91),
('1b', '2b', 'PBSF', 'DBU', 23),
('1b', '2b', 'PBSF', 'MTBD', 37),
('1b', '2b', 'PBSF', 'BTMG', 48),
('1b', '2b', 'PBSF', 'BTPP', 68),
# 2c
('1c', '2c', '3-Cl', 'DBU', 1),
('1c', '2c', '3-Cl', 'MTBD', 1),
('1c', '2c', '3-Cl', 'BTMG', 1),
('1c', '2c', '3-Cl', 'BTPP', 3),
('1c', '2c', 'PyFluor', 'DBU', 1),
('1c', '2c', 'PyFluor', 'MTBD', 2),
('1c', '2c', 'PyFluor', 'BTMG', 1),
('1c', '2c', 'PyFluor', 'BTPP', 1),
('1c', '2c', '3-CF3', 'DBU', 3),
('1c', '2c', '3-CF3', 'MTBD', 4),
('1c', '2c', '3-CF3', 'BTMG', 5),
('1c', '2c', '3-CF3', 'BTPP', 12),
('1c', '2c', '3-NO2', 'DBU', 11),
('1c', '2c', '3-NO2', 'MTBD', 12),
('1c', '2c', '3-NO2', 'BTMG', 12),
('1c', '2c', '3-NO2', 'BTPP', 22),
('1c', '2c', 'PBSF', 'DBU', 83),
('1c', '2c', 'PBSF', 'MTBD', 79),
('1c', '2c', 'PBSF', 'BTMG', 89),
('1c', '2c', 'PBSF', 'BTPP', 82),
# 2d
('1d', '2d', '3-Cl', 'DBU', 25),
('1d', '2d', '3-Cl', 'MTBD', 30),
('1d', '2d', '3-Cl', 'BTMG', 29),
('1d', '2d', '3-Cl', 'BTPP', 30),
('1d', '2d', 'PyFluor', 'DBU', 45),
('1d', '2d', 'PyFluor', 'MTBD', 40),
('1d', '2d', 'PyFluor', 'BTMG', 39),
('1d', '2d', 'PyFluor', 'BTPP', 33),
('1d', '2d', '3-CF3', 'DBU', 41),
('1d', '2d', '3-CF3', 'MTBD', 47),
('1d', '2d', '3-CF3', 'BTMG', 39),
('1d', '2d', '3-CF3', 'BTPP', 33),
('1d', '2d', '3-NO2', 'DBU', 48),
('1d', '2d', '3-NO2', 'MTBD', 48),
('1d', '2d', '3-NO2', 'BTMG', 40),
('1d', '2d', '3-NO2', 'BTPP', 32),
('1d', '2d', 'PBSF', 'DBU', 59),
('1d', '2d', 'PBSF', 'MTBD', 74),
('1d', '2d', 'PBSF', 'BTMG', 64),
('1d', '2d', 'PBSF', 'BTPP', 58),
]
# + [markdown] colab_type="text" id="51jtOEFNTccF"
# # Define all reactions in Table 1
#
# The only aspects of reaction data that vary are (a) the identity of the solute of the ```'alcohol in THF'``` input, the identity of the solute in the ```'sulfonyl fluoride'``` input, and the identity of the sole component in the ```'base'``` input. The vessel, reaction setup, solvent, timing, temperature, etc. are all constant. The yield, of course, also changes, but the instrument and type of analysis (19F NMR) is constant.
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="3ZLtrKI254si" outputId="5f902c93-b8f1-49b4-c413-829540aaccfa"
reactions = []
for alc, prod, sf, base, y in data:
# Start with template
this_condition = reaction_pb2.Reaction()
this_condition.CopyFrom(reaction)
# Modify species:
# - identifiers, vendor, prep ONLY
def modify(cpd_from, cpd_to):
del cpd_to.identifiers[:]
del cpd_to.preparations[:]
for identifier in cpd_from.identifiers:
cpd_to.identifiers.add().CopyFrom(identifier)
if cpd_from.source.vendor:
cpd_to.source.vendor = cpd_from.source.vendor
for preparation in cpd_from.preparations:
cpd_to.preparations.add().CopyFrom(preparation)
modify(compounds[alc], this_condition.inputs['alcohol in THF'].components[0])
modify(compounds[sf], this_condition.inputs['sulfonyl fluoride'].components[0])
modify(compounds[base], this_condition.inputs['base'].components[0])
del this_condition.outcomes[0].products[0].identifiers[:]
this_condition.outcomes[0].products[0].identifiers.extend(compounds[prod].identifiers)
# Modify yield
this_condition.outcomes[0].products[0].measurements[0].percentage.value = y
# Validate
validations.validate_message(this_condition)
# Save
reactions.append(this_condition)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="ith5vNbfC05O" outputId="8d55b989-5af0-46e3-ca72-1ff5877b4a63"
print(f'Generated {len(reactions)} reactions')
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="3sR-EPjuVbYk" outputId="732f72ce-675d-4d73-b09f-818c719fbf37"
# Inspect random reaction from this set
reactions[42]
# -
# ### Package reactions into a dataset
dataset = dataset_pb2.Dataset(
name='Deoxyfluorination screen',
description='Reactions from Figure 1 of DOI: 10.1021/jacs.8b01523',
reactions=reactions,
)
# message_helpers.write_message(dataset, 'ord-nielsen-example.pbtxt')
# + [markdown] colab_type="text" id="B0j2SWS73Xj4"
# ## Prepare for machine learning
# Given the list of reactions (defined here or retrieved through other means), we can prepare a DataFrame that summarizes the conditions and outcomes using only the fields that vary.
# + colab={} colab_type="code" id="lOTNUsdI3Xj4"
import pandas as pd
# + colab={"base_uri": "https://localhost:8080/", "height": 521} colab_type="code" id="UAk33ry13Xj6" outputId="caf445b2-5ec4-4a9d-cadb-9ae67d77afed"
data_dict = {
'reaction_id': [],
'alcohol': [],
'sulfonyl fluoride': [],
'base': [],
'product': [],
'yield': [],
}
for reaction in reactions:
data_dict['reaction_id'].append(reaction.reaction_id)
data_dict['alcohol'].append(
message_helpers.smiles_from_compound(reaction.inputs['alcohol in THF'].components[0])
)
data_dict['sulfonyl fluoride'].append(
message_helpers.smiles_from_compound(reaction.inputs['sulfonyl fluoride'].components[0])
)
data_dict['base'].append(
message_helpers.smiles_from_compound(reaction.inputs['base'].components[0])
)
data_dict['product'].append(
message_helpers.smiles_from_compound(reaction.outcomes[0].products[0])
)
data_dict['yield'].append(reaction.outcomes[0].products[0].measurements[0].percentage.value)
pd.DataFrame.from_dict(data_dict)
# + [markdown] colab_type="text" id="OFDZxKKl3Xj-"
# # Text description (work in progress)
# + colab={} colab_type="code" id="ClVlt8Bb3Xj_"
from ord_schema.visualization import generate_text
# + colab={"base_uri": "https://localhost:8080/", "height": 54} colab_type="code" id="kQFcEZqb3XkC" outputId="e960f683-92ed-4b28-9bc6-25f57b09e856"
generate_text.generate_text(reactions[1])
# + colab={} colab_type="code" id="a3joIRKYRXS4"
|
examples/submissions/2_Nielsen_Deoxyfluorination_Screen/example_nielsen.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="DFY-Kv5FaGxT" colab_type="text"
# # Semantic Segmentation with DeepLabV3
#
# ## CityScapes Dataset
#
# [Repository](https://github.com/srihari-humbarwadi/DeepLabV3_Plus-Tensorflow2.0) | [Pre-trained Model](https://drive.google.com/file/d/1wRXyIGUVRws3BJHX-UrNDSZGDzUzgVMx/view) | [Dataset](https://www.cityscapes-dataset.com/)
# + [markdown] id="udWA105ca0Z9" colab_type="text"
# ## Setting Up Environment
# + id="na-egK7PZmm5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 217} executionInfo={"status": "ok", "timestamp": 1593761676494, "user_tz": -120, "elapsed": 50168, "user": {"displayName": "artificialint Workshop", "photoUrl": "", "userId": "12022852047375758782"}} outputId="8f187e83-ee5b-4751-dcbd-c0409ba2ae9c"
# Clone Repo
# !git clone --recursive https://github.com/srihari-humbarwadi/DeepLabV3_Plus-Tensorflow2.0; \
# rm -dr sample_data; \
# mv DeepLabV3_Plus-Tensorflow2.0/* .; \
# rm -drf DeepLabV3_Plus-Tensorflow2.0; \
# mkdir inferences
# Update Source Code
# !sed -i 's/from tensorflow.python.keras.applications import keras_modules_injection/# from tensorflow.python.keras.applications import keras_modules_injection/g' resnet50.py
# !sed -i 's/@keras_modules_injection/# @keras_modules_injection/g' resnet50.py
# !awk '/from .imagenet_utils import _obtain_input_shape/ { print; print "from tensorflow import keras"; next }1' resnet/resnet50.py > tmp && mv tmp resnet/resnet50.py
# !awk '/# Determine proper input shape/ { print; print " keras_utils = keras.utils"; next }1' resnet/resnet50.py > tmp && mv tmp resnet/resnet50.py
# !awk '/# Determine proper input shape/ { print; print " models = keras.models"; next }1' resnet/resnet50.py > tmp && mv tmp resnet/resnet50.py
# !awk '/# Determine proper input shape/ { print; print " layers = keras.layers"; next }1' resnet/resnet50.py > tmp && mv tmp resnet/resnet50.py
# !awk '/# Determine proper input shape/ { print; print " backend = keras.backend"; next }1' resnet/resnet50.py > tmp && mv tmp resnet/resnet50.py
# Download Pretrained Model
# !fileid="1wRXyIGUVRws3BJHX-UrNDSZGDzUzgVMx"; \
# filename="top_weights.h5"; \
# curl -c ./cookie -s -L "https://drive.google.com/uc?export=download&id=${fileid}" > /dev/null; \
# curl -Lb ./cookie "https://drive.google.com/uc?export=download&confirm=`awk '/download/ {print $NF}' ./cookie`&id=${fileid}" -o ${filename}
# + [markdown] id="o-aMUpQvbLiS" colab_type="text"
# ## Reconstructing Model
# + id="YUFocNj-a8cq" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1593761678576, "user_tz": -120, "elapsed": 52230, "user": {"displayName": "artificialint Workshop", "photoUrl": "", "userId": "12022852047375758782"}}
import os
import pickle
import numpy as np
import cv2
import matplotlib.pyplot as plt
import tensorflow as tf
from tensorflow.python.keras.preprocessing.image import load_img, img_to_array
from tensorflow.keras.applications.resnet50 import preprocess_input
from deeplab import DeepLabV3Plus
# + id="b5uIZftza91o" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 146} executionInfo={"status": "ok", "timestamp": 1593761693910, "user_tz": -120, "elapsed": 67555, "user": {"displayName": "artificialint Workshop", "photoUrl": "", "userId": "12022852047375758782"}} outputId="f3c49d15-4f71-45fe-a510-738e8323e7d9"
# Instantiate Model
h, w = 800, 1600
with open('cityscapes_dict.pkl', 'rb') as f:
id_to_color = pickle.load(f)['color_map']
model = DeepLabV3Plus(h, w, 34)
model.load_weights('top_weights.h5')
# + [markdown] id="ovgmOFF9bSew" colab_type="text"
# ## Making an Inference
# + [markdown] id="Z1RNrsogbWqM" colab_type="text"
# Either upload an image or download it from a URL:
# + id="Bsd78sWIbWQv" colab_type="code" colab={}
# EITHER: Upload Image Input
from google.colab import files
files.upload()
# Don't forget to put the image into the inferences folder
# + id="mY-qprXmbUYh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 72} executionInfo={"status": "ok", "timestamp": 1593761696697, "user_tz": -120, "elapsed": 70336, "user": {"displayName": "artificialint Workshop", "photoUrl": "", "userId": "12022852047375758782"}} outputId="67994c7f-190d-41ac-f3c9-136bf0b79b3a"
# OR: Download Image Input
# !imageurl="https://www.researchgate.net/profile/Varun_Jampani/publication/319056828/figure/fig3/AS:667765274845187@1536219056188/Qualitative-results-from-the-Cityscapes-dataset-Observe-how-NetWarp-PSPNet-is-able-to.jpg"; \
# filetype='jpg'; \
# curl ${imageurl} > inferences/input.${filetype}
# + id="R-Boy4pSbpKH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 184} executionInfo={"status": "ok", "timestamp": 1593761709416, "user_tz": -120, "elapsed": 83049, "user": {"displayName": "artificialint Workshop", "photoUrl": "", "userId": "12022852047375758782"}} outputId="8bb55ec5-465d-454c-eb99-84aec65c58da"
# Load Image
image_dir = 'inferences'
image_list = os.listdir(image_dir)
image_list.sort()
image = load_img(f'{image_dir}/{image_list[0]}')
image = img_to_array(image)
# Preprocess and Make Inference
image = cv2.resize(image, (w, h))
x = image.copy()
y = np.argmax(np.squeeze(
model.predict(preprocess_input(np.expand_dims(x, axis=0)))), axis=2)
# Turn Inference to Image
img_color = image.copy()
for i in np.unique(y):
if i in id_to_color:
img_color[y == i] = id_to_color[i]
disp = img_color.copy()
# Visualise Inference
alpha = 0.5
cv2.addWeighted(image, alpha, img_color, 1 - alpha, 0, img_color)
out = np.concatenate([image/255, img_color/255, disp/255], axis=1)
plt.figure(figsize=(24, 12))
plt.xticks([])
plt.yticks([])
plt.imshow(img_color/255.0)
plt.imshow(out)
# + [markdown] id="oiZNiQtQmIfl" colab_type="text"
# ## Interpreting the Inference
#
# [cityscapesScripts](https://github.com/mcordts/cityscapesScripts/blob/master/cityscapesscripts/helpers/labels.py)
# + id="fAnYoPCkmQCX" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1593762044180, "user_tz": -120, "elapsed": 976, "user": {"displayName": "artificialint Workshop", "photoUrl": "", "userId": "12022852047375758782"}}
# Metadata from cityscapesScripts
from collections import namedtuple
Label = namedtuple( 'Label' , [
'name' , # The identifier of this label, e.g. 'car', 'person', ... .
# We use them to uniquely name a class
'id' , # An integer ID that is associated with this label.
# The IDs are used to represent the label in ground truth images
# An ID of -1 means that this label does not have an ID and thus
# is ignored when creating ground truth images (e.g. license plate).
# Do not modify these IDs, since exactly these IDs are expected by the
# evaluation server.
'trainId' , # Feel free to modify these IDs as suitable for your method. Then create
# ground truth images with train IDs, using the tools provided in the
# 'preparation' folder. However, make sure to validate or submit results
# to our evaluation server using the regular IDs above!
# For trainIds, multiple labels might have the same ID. Then, these labels
# are mapped to the same class in the ground truth images. For the inverse
# mapping, we use the label that is defined first in the list below.
# For example, mapping all void-type classes to the same ID in training,
# might make sense for some approaches.
# Max value is 255!
'category' , # The name of the category that this label belongs to
'categoryId' , # The ID of this category. Used to create ground truth images
# on category level.
'hasInstances', # Whether this label distinguishes between single instances or not
'ignoreInEval', # Whether pixels having this class as ground truth label are ignored
# during evaluations or not
'color' , # The color of this label
] )
labels = [
# name id trainId category catId hasInstances ignoreInEval color
Label( 'unlabeled' , 0 , 255 , 'void' , 0 , False , True , ( 0, 0, 0) ),
Label( 'ego vehicle' , 1 , 255 , 'void' , 0 , False , True , ( 0, 0, 0) ),
Label( 'rectification border' , 2 , 255 , 'void' , 0 , False , True , ( 0, 0, 0) ),
Label( 'out of roi' , 3 , 255 , 'void' , 0 , False , True , ( 0, 0, 0) ),
Label( 'static' , 4 , 255 , 'void' , 0 , False , True , ( 0, 0, 0) ),
Label( 'dynamic' , 5 , 255 , 'void' , 0 , False , True , (111, 74, 0) ),
Label( 'ground' , 6 , 255 , 'void' , 0 , False , True , ( 81, 0, 81) ),
Label( 'road' , 7 , 0 , 'flat' , 1 , False , False , (128, 64,128) ),
Label( 'sidewalk' , 8 , 1 , 'flat' , 1 , False , False , (244, 35,232) ),
Label( 'parking' , 9 , 255 , 'flat' , 1 , False , True , (250,170,160) ),
Label( 'rail track' , 10 , 255 , 'flat' , 1 , False , True , (230,150,140) ),
Label( 'building' , 11 , 2 , 'construction' , 2 , False , False , ( 70, 70, 70) ),
Label( 'wall' , 12 , 3 , 'construction' , 2 , False , False , (102,102,156) ),
Label( 'fence' , 13 , 4 , 'construction' , 2 , False , False , (190,153,153) ),
Label( 'guard rail' , 14 , 255 , 'construction' , 2 , False , True , (180,165,180) ),
Label( 'bridge' , 15 , 255 , 'construction' , 2 , False , True , (150,100,100) ),
Label( 'tunnel' , 16 , 255 , 'construction' , 2 , False , True , (150,120, 90) ),
Label( 'pole' , 17 , 5 , 'object' , 3 , False , False , (153,153,153) ),
Label( 'polegroup' , 18 , 255 , 'object' , 3 , False , True , (153,153,153) ),
Label( 'traffic light' , 19 , 6 , 'object' , 3 , False , False , (250,170, 30) ),
Label( 'traffic sign' , 20 , 7 , 'object' , 3 , False , False , (220,220, 0) ),
Label( 'vegetation' , 21 , 8 , 'nature' , 4 , False , False , (107,142, 35) ),
Label( 'terrain' , 22 , 9 , 'nature' , 4 , False , False , (152,251,152) ),
Label( 'sky' , 23 , 10 , 'sky' , 5 , False , False , ( 70,130,180) ),
Label( 'person' , 24 , 11 , 'human' , 6 , True , False , (220, 20, 60) ),
Label( 'rider' , 25 , 12 , 'human' , 6 , True , False , (255, 0, 0) ),
Label( 'car' , 26 , 13 , 'vehicle' , 7 , True , False , ( 0, 0,142) ),
Label( 'truck' , 27 , 14 , 'vehicle' , 7 , True , False , ( 0, 0, 70) ),
Label( 'bus' , 28 , 15 , 'vehicle' , 7 , True , False , ( 0, 60,100) ),
Label( 'caravan' , 29 , 255 , 'vehicle' , 7 , True , True , ( 0, 0, 90) ),
Label( 'trailer' , 30 , 255 , 'vehicle' , 7 , True , True , ( 0, 0,110) ),
Label( 'train' , 31 , 16 , 'vehicle' , 7 , True , False , ( 0, 80,100) ),
Label( 'motorcycle' , 32 , 17 , 'vehicle' , 7 , True , False , ( 0, 0,230) ),
Label( 'bicycle' , 33 , 18 , 'vehicle' , 7 , True , False , (119, 11, 32) ),
Label( 'license plate' , -1 , -1 , 'vehicle' , 7 , False , True , ( 0, 0,142) ),
]
# + id="A2OH_2A0ntq0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 417} executionInfo={"status": "ok", "timestamp": 1593763030734, "user_tz": -120, "elapsed": 1034, "user": {"displayName": "artificialint Workshop", "photoUrl": "", "userId": "12022852047375758782"}} outputId="f24e870b-45a3-46db-f9b9-71aa59b5af12"
counts = np.unique(y, return_counts=True)
total = y.size
print([idx for idx, val in enumerate(counts[0]) if val == 1])
for label in labels:
label_idx = label.id
counter_idx = None
for idx, val in enumerate(counts[0]):
if val == label_idx:
counter_idx = idx
if counter_idx:
percentage = 100 * counts[1][counter_idx] / total
print(f'{percentage:>10.5f}% {label.name}')
# + [markdown] id="gkLqkHOnyysP" colab_type="text"
# Developed by the City Intelligence Lab, Austrian Institute of Technology GmbH
|
colab-notebooks/semantic-segmentation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import cv2
import matplotlib.pyplot as plt
import numpy as np
# # Q31 アフィン変換(スキュー)
# (1)アフィン変換を用いて、出力(1)のようなX-sharing(dx = 30)画像を作成せよ。
#
# (2)アフィン変換を用いて、出力2のようなY-sharing(dy = 30)画像を作成せよ。
#
# (3)アフィン変換を用いて、出力3のような幾何変換した(dx = 30, dy = 30)画像を作成せよ。
#
# このような画像はスキュー画像と呼ばれ、画像を斜め方向に伸ばした画像である。
# +
def skew_img(img,sx,sy):
h,w,l = img.shape
M = np.array([[1,sx/h,0],[sy/w,1,0]],dtype=np.float32)
out = cv2.warpAffine(img,M,(w+sx,h+sy))
return out
img = cv2.imread("imori.jpg")
sx = 30
sy = 30
result1 = skew_img(img,sx,0)
result2 = skew_img(img,0,sy)
result3 = skew_img(img,sx,sy)
plt.figure(facecolor="white",figsize=(10,3))
plt.subplot(1,4,1)
plt.imshow(cv2.cvtColor(img, cv2.COLOR_BGR2RGB))
plt.subplot(1,4,2)
plt.imshow(cv2.cvtColor(result1, cv2.COLOR_BGR2RGB))
plt.subplot(1,4,3)
plt.imshow(cv2.cvtColor(result2, cv2.COLOR_BGR2RGB))
plt.subplot(1,4,4)
plt.imshow(cv2.cvtColor(result3, cv2.COLOR_BGR2RGB))
plt.show()
# -
# # Q32. フーリエ変換
#
# 二次元離散フーリエ変換(DFT)を実装し、imori.jpgをグレースケール化したものの周波数のパワースペクトルを表示せよ。 また、逆二次元離散フーリエ変換(IDFT)で画像を復元せよ。
# +
img = cv2.imread("imori_gray.jpg")
f = np.fft.fft2(img)
fshift = np.fft.fftshift(f)
magnitude_spectrum = 20*np.log(np.abs(fshift))
plt.subplot(121),plt.imshow(img, cmap = 'gray')
plt.title('Input Image'), plt.xticks([]), plt.yticks([])
plt.subplot(122),plt.imshow(magnitude_spectrum, cmap = 'gray')
plt.title('Magnitude Spectrum'), plt.xticks([]), plt.yticks([])
plt.show()
# -
magnitude_spectrum
|
Question_31_40/.ipynb_checkpoints/MyAnswer-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="uqNjQjCJ_aSt" executionInfo={"status": "error", "timestamp": 1617022227422, "user_tz": 180, "elapsed": 4231, "user": {"displayName": "<NAME>\u00f5es", "photoUrl": "<KEY>", "userId": "05351605997823750375"}} outputId="7a26df52-7992-4e05-83f8-e8f8336d3bd9" colab={"base_uri": "https://localhost:8080/", "height": 367}
import os
import glob
import random
import shutil
from tempfile import gettempdir
from IPython.display import clear_output, Image
from torchvision.datasets.folder import pil_loader
from model import Net
from utils import pil_to_model_tensor_transform
import consts
# UTKFace constants
MALE = 0
FEMALE = 1
WHITE = 0
BLACK = 1
ASIAN = 2
INDIAN = 3
OTHER = 4
# User constants
dset_path = os.path.join('.', 'data', 'UTKFace', 'unlabeled')
tempdir = gettempdir()
# + id="8dzEdzAa_aSy" outputId="15abd643-44ef-40dc-9ced-1c24b5609761"
consts.NUM_Z_CHANNELS = 50 # we have two trained models, with 50 and 100
net = Net()
load_path = {50: r".\trained_models\2018_09_08\01_44\epoch76", 100: r"C:\Users\Mattan\Downloads\epoch_200_no_tf"}[consts.NUM_Z_CHANNELS]
net.load(load_path, slim=True) # slim tells the net to load only the encoder and generator
# + id="TKb9utNq_aS0" outputId="4756ca42-f330-4ac7-a039-6a5320aad40e"
# Game 1: Age Progression/Regression
# Set the attributes of a random person you want to test
age = 38
gender = FEMALE
race = OTHER
image_path = random.choice(glob.glob(os.path.join(dset_path, '{a}_{g}_{r}*'.format(a=age, g=gender, r=race))))
Image(filename=image_path) # Will select and show a person with the attributes you selected
# + id="J8_CpoXY_aS0" outputId="e0e1c433-17a3-4f5a-b7e0-5b6a857f456b"
# Game 1: Age Progression/Regression
image_tensor = pil_to_model_tensor_transform(pil_loader(image_path))
Image(filename=net.test_single(image_tensor=image_tensor, age=age, gender=gender, target=tempdir, watermark=False))
|
aging_game.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Pixel to Pixel Generative Adversarial Networks
#
# [Pixel to Pixel Generative Adversarial Networks](https://arxiv.org/abs/1611.07004) applies [Conditional Generative Adversarial Networks](https://arxiv.org/abs/1411.1784) as a general-purpose solution to image-to-image translation problems. These networks not only learn the mapping from input image to output image, but also learn a loss function to train this mapping.
#
# With pixel2pixel GAN, it is possible to train different type of image translation tasks with small datasets. In this tutorial, we will train on three image translation tasks: facades with 400 images from [CMP Facades dataset](http://cmp.felk.cvut.cz/%7Etylecr1/facade/), cityscapes with 2975 images from [Cityscapes training set](https://www.cityscapes-dataset.com/) and maps with 1096 training images scraped from Google Maps.
#
# For harder problems such as [edges2shoes](http://vision.cs.utexas.edu/projects/finegrained/utzap50k/) and [edges2handbags](https://github.com/junyanz/iGAN), it may be important to train on far larger datasets, which takes significantly more time. You can try them with [Multiple GPUs](http://gluon.mxnet.io/chapter07_distributed-learning/multiple-gpus-gluon.html).
# +
from __future__ import print_function
import os
import matplotlib as mpl
import tarfile
import matplotlib.image as mpimg
from matplotlib import pyplot as plt
import mxnet as mx
from mxnet import gluon
from mxnet import ndarray as nd
from mxnet.gluon import nn, utils
from mxnet.gluon.nn import Dense, Activation, Conv2D, Conv2DTranspose, \
BatchNorm, LeakyReLU, Flatten, HybridSequential, HybridBlock, Dropout
from mxnet import autograd
import numpy as np
# -
# ## Set Training parameters
# +
epochs = 100
batch_size = 10
use_gpu = True
ctx = mx.gpu() if use_gpu else mx.cpu()
lr = 0.0002
beta1 = 0.5
lambda1 = 100
pool_size = 50
# -
# ## Download and Preprocess Dataset
#
# We first train on facades dataset. We need to crop images to input images and output images. Notice that pixel2pixel GAN is capable to train these tasks bidirectional. You can set ```is-reversed=True``` to switch input and output image patterns.
dataset = 'facades'
# We first resize images to size 512 * 256. Then normalize image pixel values to be between -1 and 1.
# +
img_wd = 256
img_ht = 256
train_img_path = '%s/train' % (dataset)
val_img_path = '%s/val' % (dataset)
def download_data(dataset):
if not os.path.exists(dataset):
url = 'https://people.eecs.berkeley.edu/~tinghuiz/projects/pix2pix/datasets/%s.tar.gz' % (dataset)
os.mkdir(dataset)
data_file = utils.download(url)
with tarfile.open(data_file) as tar:
tar.extractall(path='.')
os.remove(data_file)
def load_data(path, batch_size, is_reversed=False):
img_in_list = []
img_out_list = []
for path, _, fnames in os.walk(path):
for fname in fnames:
if not fname.endswith('.jpg'):
continue
img = os.path.join(path, fname)
img_arr = mx.image.imread(img).astype(np.float32)/127.5 - 1
img_arr = mx.image.imresize(img_arr, img_wd * 2, img_ht)
# Crop input and output images
img_arr_in, img_arr_out = [mx.image.fixed_crop(img_arr, 0, 0, img_wd, img_ht),
mx.image.fixed_crop(img_arr, img_wd, 0, img_wd, img_ht)]
img_arr_in, img_arr_out = [nd.transpose(img_arr_in, (2,0,1)),
nd.transpose(img_arr_out, (2,0,1))]
img_arr_in, img_arr_out = [img_arr_in.reshape((1,) + img_arr_in.shape),
img_arr_out.reshape((1,) + img_arr_out.shape)]
img_in_list.append(img_arr_out if is_reversed else img_arr_in)
img_out_list.append(img_arr_in if is_reversed else img_arr_out)
return mx.io.NDArrayIter(data=[nd.concat(*img_in_list, dim=0), nd.concat(*img_out_list, dim=0)],
batch_size=batch_size)
download_data(dataset)
train_data = load_data(train_img_path, batch_size, is_reversed=True)
val_data = load_data(val_img_path, batch_size, is_reversed=True)
# -
# Visualize 4 images:
# +
def visualize(img_arr):
plt.imshow(((img_arr.asnumpy().transpose(1, 2, 0) + 1.0) * 127.5).astype(np.uint8))
plt.axis('off')
def preview_train_data():
img_in_list, img_out_list = train_data.next().data
for i in range(4):
plt.subplot(2,4,i+1)
visualize(img_in_list[i])
plt.subplot(2,4,i+5)
visualize(img_out_list[i])
plt.show()
preview_train_data()
# -
# ## Defining the networks
#
# Both generator and discriminator use modules of the form convolution-BatchNorm-ReLu.
#
# The key for generator is U-net architecture adding skip connections which shuttle low-level infomation shared between input and output images across net.
# 
#
# PatchGAN – that only penalizes structure at the scale of patches is applied as disciminator architecture. This discriminator tries to classify if each N × N patch in an image is real or fake. We run this discriminator convolutionally across the image, averaging all responses to provide the ultimate output of netD.
# +
# Define Unet generator skip block
class UnetSkipUnit(HybridBlock):
def __init__(self, inner_channels, outer_channels, inner_block=None, innermost=False, outermost=False,
use_dropout=False, use_bias=False):
super(UnetSkipUnit, self).__init__()
with self.name_scope():
self.outermost = outermost
en_conv = Conv2D(channels=inner_channels, kernel_size=4, strides=2, padding=1,
in_channels=outer_channels, use_bias=use_bias)
en_relu = LeakyReLU(alpha=0.2)
en_norm = BatchNorm(momentum=0.1, in_channels=inner_channels)
de_relu = Activation(activation='relu')
de_norm = BatchNorm(momentum=0.1, in_channels=outer_channels)
if innermost:
de_conv = Conv2DTranspose(channels=outer_channels, kernel_size=4, strides=2, padding=1,
in_channels=inner_channels, use_bias=use_bias)
encoder = [en_relu, en_conv]
decoder = [de_relu, de_conv, de_norm]
model = encoder + decoder
elif outermost:
de_conv = Conv2DTranspose(channels=outer_channels, kernel_size=4, strides=2, padding=1,
in_channels=inner_channels * 2)
encoder = [en_conv]
decoder = [de_relu, de_conv, Activation(activation='tanh')]
model = encoder + [inner_block] + decoder
else:
de_conv = Conv2DTranspose(channels=outer_channels, kernel_size=4, strides=2, padding=1,
in_channels=inner_channels * 2, use_bias=use_bias)
encoder = [en_relu, en_conv, en_norm]
decoder = [de_relu, de_conv, de_norm]
model = encoder + [inner_block] + decoder
if use_dropout:
model += [Dropout(rate=0.5)]
self.model = HybridSequential()
with self.model.name_scope():
for block in model:
self.model.add(block)
def hybrid_forward(self, F, x):
if self.outermost:
return self.model(x)
else:
return F.concat(self.model(x), x, dim=1)
# Define Unet generator
class UnetGenerator(HybridBlock):
def __init__(self, in_channels, num_downs, ngf=64, use_dropout=True):
super(UnetGenerator, self).__init__()
#Build unet generator structure
unet = UnetSkipUnit(ngf * 8, ngf * 8, innermost=True)
for _ in range(num_downs - 5):
unet = UnetSkipUnit(ngf * 8, ngf * 8, unet, use_dropout=use_dropout)
unet = UnetSkipUnit(ngf * 8, ngf * 4, unet)
unet = UnetSkipUnit(ngf * 4, ngf * 2, unet)
unet = UnetSkipUnit(ngf * 2, ngf * 1, unet)
unet = UnetSkipUnit(ngf, in_channels, unet, outermost=True)
with self.name_scope():
self.model = unet
def hybrid_forward(self, F, x):
return self.model(x)
# Define the PatchGAN discriminator
class Discriminator(HybridBlock):
def __init__(self, in_channels, ndf=64, n_layers=3, use_sigmoid=False, use_bias=False):
super(Discriminator, self).__init__()
with self.name_scope():
self.model = HybridSequential()
kernel_size = 4
padding = int(np.ceil((kernel_size - 1)/2))
self.model.add(Conv2D(channels=ndf, kernel_size=kernel_size, strides=2,
padding=padding, in_channels=in_channels))
self.model.add(LeakyReLU(alpha=0.2))
nf_mult = 1
for n in range(1, n_layers):
nf_mult_prev = nf_mult
nf_mult = min(2 ** n, 8)
self.model.add(Conv2D(channels=ndf * nf_mult, kernel_size=kernel_size, strides=2,
padding=padding, in_channels=ndf * nf_mult_prev,
use_bias=use_bias))
self.model.add(BatchNorm(momentum=0.1, in_channels=ndf * nf_mult))
self.model.add(LeakyReLU(alpha=0.2))
nf_mult_prev = nf_mult
nf_mult = min(2 ** n_layers, 8)
self.model.add(Conv2D(channels=ndf * nf_mult, kernel_size=kernel_size, strides=1,
padding=padding, in_channels=ndf * nf_mult_prev,
use_bias=use_bias))
self.model.add(BatchNorm(momentum=0.1, in_channels=ndf * nf_mult))
self.model.add(LeakyReLU(alpha=0.2))
self.model.add(Conv2D(channels=1, kernel_size=kernel_size, strides=1,
padding=padding, in_channels=ndf * nf_mult))
if use_sigmoid:
self.model.add(Activation(activation='sigmoid'))
def hybrid_forward(self, F, x):
out = self.model(x)
#print(out)
return out
# -
# ## Construct networks, Initialize parameters, Setup Loss Function and Optimizer
# We use binary cross entropy and L1 loss as loss functions. L1 loss can be used to capture low frequencies in images.
# +
def param_init(param):
if param.name.find('conv') != -1:
if param.name.find('weight') != -1:
param.initialize(init=mx.init.Normal(0.02), ctx=ctx)
else:
param.initialize(init=mx.init.Zero(), ctx=ctx)
elif param.name.find('batchnorm') != -1:
param.initialize(init=mx.init.Zero(), ctx=ctx)
# Initialize gamma from normal distribution with mean 1 and std 0.02
if param.name.find('gamma') != -1:
param.set_data(nd.random_normal(1, 0.02, param.data().shape))
def network_init(net):
for param in net.collect_params().values():
param_init(param)
def set_network():
# Pixel2pixel networks
netG = UnetGenerator(in_channels=3, num_downs=8)
netD = Discriminator(in_channels=6)
# Initialize parameters
network_init(netG)
network_init(netD)
# trainer for the generator and the discriminator
trainerG = gluon.Trainer(netG.collect_params(), 'adam', {'learning_rate': lr, 'beta1': beta1})
trainerD = gluon.Trainer(netD.collect_params(), 'adam', {'learning_rate': lr, 'beta1': beta1})
return netG, netD, trainerG, trainerD
# Loss
GAN_loss = gluon.loss.SigmoidBinaryCrossEntropyLoss()
L1_loss = gluon.loss.L1Loss()
netG, netD, trainerG, trainerD = set_network()
# -
# ## Image pool for discriminator
# We use history image pool to help discriminator memorize history errors instead of just comparing current real input and fake output.
class ImagePool():
def __init__(self, pool_size):
self.pool_size = pool_size
if self.pool_size > 0:
self.num_imgs = 0
self.images = []
def query(self, images):
if self.pool_size == 0:
return images
ret_imgs = []
for i in range(images.shape[0]):
image = nd.expand_dims(images[i], axis=0)
if self.num_imgs < self.pool_size:
self.num_imgs = self.num_imgs + 1
self.images.append(image)
ret_imgs.append(image)
else:
p = nd.random_uniform(0, 1, shape=(1,)).asscalar()
if p > 0.5:
random_id = nd.random_uniform(0, self.pool_size - 1, shape=(1,)).astype(np.uint8).asscalar()
tmp = self.images[random_id].copy()
self.images[random_id] = image
ret_imgs.append(tmp)
else:
ret_imgs.append(image)
ret_imgs = nd.concat(*ret_imgs, dim=0)
return ret_imgs
# ## Training Loop
# We recommend to use gpu to boost training. After a few epochs, we can see images silimar to building structure are generated.
# +
from datetime import datetime
import time
import logging
def facc(label, pred):
pred = pred.ravel()
label = label.ravel()
return ((pred > 0.5) == label).mean()
def train():
image_pool = ImagePool(pool_size)
metric = mx.metric.CustomMetric(facc)
stamp = datetime.now().strftime('%Y_%m_%d-%H_%M')
logging.basicConfig(level=logging.DEBUG)
for epoch in range(epochs):
tic = time.time()
btic = time.time()
train_data.reset()
iter = 0
for batch in train_data:
############################
# (1) Update D network: maximize log(D(x, y)) + log(1 - D(x, G(x, z)))
###########################
real_in = batch.data[0].as_in_context(ctx)
real_out = batch.data[1].as_in_context(ctx)
fake_out = netG(real_in)
fake_concat = image_pool.query(nd.concat(real_in, fake_out, dim=1))
with autograd.record():
# Train with fake image
# Use image pooling to utilize history images
output = netD(fake_concat)
fake_label = nd.zeros(output.shape, ctx=ctx)
errD_fake = GAN_loss(output, fake_label)
metric.update([fake_label,], [output,])
# Train with real image
real_concat = nd.concat(real_in, real_out, dim=1)
output = netD(real_concat)
real_label = nd.ones(output.shape, ctx=ctx)
errD_real = GAN_loss(output, real_label)
errD = (errD_real + errD_fake) * 0.5
errD.backward()
metric.update([real_label,], [output,])
trainerD.step(batch.data[0].shape[0])
############################
# (2) Update G network: maximize log(D(x, G(x, z))) - lambda1 * L1(y, G(x, z))
###########################
with autograd.record():
fake_out = netG(real_in)
fake_concat = nd.concat(real_in, fake_out, dim=1)
output = netD(fake_concat)
real_label = nd.ones(output.shape, ctx=ctx)
errG = GAN_loss(output, real_label) + L1_loss(real_out, fake_out) * lambda1
errG.backward()
trainerG.step(batch.data[0].shape[0])
# Print log infomation every ten batches
if iter % 10 == 0:
name, acc = metric.get()
logging.info('speed: {} samples/s'.format(batch_size / (time.time() - btic)))
logging.info('discriminator loss = %f, generator loss = %f, binary training acc = %f at iter %d epoch %d'
%(nd.mean(errD).asscalar(),
nd.mean(errG).asscalar(), acc, iter, epoch))
iter = iter + 1
btic = time.time()
name, acc = metric.get()
metric.reset()
logging.info('\nbinary training acc at epoch %d: %s=%f' % (epoch, name, acc))
logging.info('time: %f' % (time.time() - tic))
# Visualize one generated image for each epoch
fake_img = fake_out[0]
visualize(fake_img)
plt.show()
train()
# -
# ## Results
# Generate images with generator.
#
# +
def print_result():
num_image = 4
img_in_list, img_out_list = val_data.next().data
for i in range(num_image):
img_in = nd.expand_dims(img_in_list[i], axis=0)
plt.subplot(2,4,i+1)
visualize(img_in[0])
img_out = netG(img_in.as_in_context(ctx))
plt.subplot(2,4,i+5)
visualize(img_out[0])
plt.show()
print_result()
# -
# ## Other dataset experiments
# Run experiments on cityscapes and maps datasets
# +
datasets = ['cityscapes', 'maps']
is_reversed = False
batch_size = 64
for dataset in datasets:
train_img_path = '%s/train' % (dataset)
val_img_path = '%s/val' % (dataset)
download_data(dataset)
train_data = load_data(train_img_path, batch_size, is_reversed=is_reversed)
val_data = load_data(val_img_path, batch_size, is_reversed=is_reversed)
print("Preview %s training data:" % (dataset))
preview_train_data()
netG, netD, trainerG, trainerD = set_network()
train()
print("Training result for %s" % (dataset))
print_result()
# -
# ## Citation
#
# CMP Facades dataset:
# @INPROCEEDINGS{
# Tylecek13,
# author = {<NAME>, Radim {\v S}{\' a}ra},
# title = {Spatial Pattern Templates for Recognition of Objects with Regular Structure},
# booktitle = {Proc. GCPR},
# year = {2013},
# address = {Saarbrucken, Germany},
# }
#
# Cityscapes training set:
# @inproceedings{Cordts2016Cityscapes,
# title={The Cityscapes Dataset for Semantic Urban Scene Understanding},
# author={<NAME> and <NAME> Ramos, <NAME>, Timo and Enzweiler, Markus and <NAME> <NAME> <NAME>},
# booktitle={Proc. of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR)},
# year={2016}
# }
|
chapter14_generative-adversarial-networks/pixel2pixel.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Converting between the 4-metric $g_{\mu\nu}$ and ADM variables $\left\{\gamma_{ij}, \alpha, \beta^i\right\}$ or BSSN variables $\left\{h_{ij}, {\rm cf}, \alpha, {\rm vet}^i\right\}$
# ## Author: <NAME>
#
# [comment]: <> (Abstract: TODO)
#
# ### We will often find it useful to convert between the 4-metric $g_{\mu\nu}$ and the ADM or BSSN variables. This notebook documents the NRPy+ Python module [`BSSN.ADMBSSN_tofrom_4metric`](../edit/BSSN/ADMBSSN_tofrom_4metric.py), which provides that functionality.
#
# **Module Status:** <font color='orange'><b> Self-validated, some additional tests performed </b></font>
#
# **Validation Notes:** This tutorial module has been confirmed to be self-consistent with its corresponding NRPy+ module, as documented [below](#code_validation). In addition, the construction of $g_{\mu\nu}$ and $g^{\mu\nu}$ from BSSN variables has passed the test $g^{\mu\nu}g_{\mu\nu}=4$ [below](#validationcontraction). **Additional validation tests may have been performed, but are as yet, undocumented. (TODO)**
#
# ### NRPy+ Source Code for this module: [BSSN/ADMBSSN_tofrom_4metric.py](../edit/BSSN/ADMBSSN_tofrom_4metric.py)
#
# ## Introduction:
#
# <a id='toc'></a>
#
# # Table of Contents
# $$\label{toc}$$
#
# This module is organized as follows
#
# 1. [Step 1](#setup_ADM_quantities): `setup_ADM_quantities(inputvars)`: If `inputvars="ADM"` declare ADM quantities $\left\{\gamma_{ij},\beta^i,\alpha\right\}$; if `inputvars="ADM"` define ADM quantities in terms of BSSN quantities
# 1. [Step 2](#admbssn_to_fourmetric): Write 4-metric $g_{\mu\nu}$ and its inverse $g^{\mu\nu}$ in terms of ADM or BSSN quantities
# 1. [Step 2.a](#admbssn_to_fourmetric_lower): 4-metric $g_{\mu\nu}$ in terms of ADM or BSSN quantities
# 1. [Step 2.b](#admbssn_to_fourmetric_inv): 4-metric inverse $g^{\mu\nu}$ in terms of ADM or BSSN quantities
# 1. [Step 2.c](#validationcontraction): Validation check: Confirm $g_{\mu\nu}g^{\mu\nu}=4$
# 1. [Step 3](#fourmetric_to_admbssn): Write ADM/BSSN metric quantities in terms of 4-metric $g_{\mu\nu}$ (Excludes extrinsic curvature $K_{ij}$ or the BSSN $\bar{A}_{ij}$, $K$)
# 1. [Step 3.a](#adm_ito_fourmetric_validate): ADM in terms of 4-metric validation: Confirm $\gamma_{ij}\gamma^{ij}=3$
# 1. [Step 3.b](#bssn_ito_fourmetric_validate): BSSN in terms of 4-metric validation: Confirm $\bar{\gamma}_{ij}\bar{\gamma}^{ij}=3$
# 1. [Step 4](#code_validation): Code Validation against `BSSN.ADMBSSN_tofrom_4metric` NRPy+ module
# 1. [Step 5](#latex_pdf_output): Output this module to $\LaTeX$-formatted PDF
# <a id='setup_ADM_quantities'></a>
#
# # Step 1: `setup_ADM_quantities(inputvars)`: If `inputvars="ADM"` declare ADM quantities $\left\{\gamma_{ij},\beta^i,\alpha\right\}$; if `inputvars="ADM"` define ADM quantities in terms of BSSN quantities \[Back to [top](#toc)\]
# $$\label{setup_ADM_quantities}$$
# +
import sympy as sp
import NRPy_param_funcs as par
import indexedexp as ixp
import sys
def setup_ADM_quantities(inputvars):
if inputvars == "ADM":
gammaDD = ixp.declarerank2("gammaDD", "sym01")
betaU = ixp.declarerank1("betaU")
alpha = sp.symbols("alpha", real=True)
elif inputvars == "BSSN":
import BSSN.ADM_in_terms_of_BSSN as AitoB
# Construct gamma_{ij} in terms of cf & gammabar_{ij}
AitoB.ADM_in_terms_of_BSSN()
gammaDD = AitoB.gammaDD
# Next construct beta^i in terms of vet^i and reference metric quantities
import BSSN.BSSN_quantities as Bq
Bq.BSSN_basic_tensors()
betaU = Bq.betaU
alpha = sp.symbols("alpha", real=True)
else:
print("inputvars = " + str(inputvars) + " not supported. Please choose ADM or BSSN.")
sys.exit(1)
return gammaDD,betaU,alpha
# -
# <a id='admbssn_to_fourmetric'></a>
#
# # Step 2: Write 4-metric $g_{\mu\nu}$ and its inverse $g^{\mu\nu}$ in terms of ADM or BSSN variables \[Back to [top](#toc)\]
# $$\label{admbssn_to_fourmetric}$$
#
# <a id='admbssn_to_fourmetric_lower'></a>
#
# ## Step 2.a: 4-metric $g_{\mu\nu}$ in terms of ADM or BSSN variables \[Back to [top](#toc)\]
# $$\label{admbssn_to_fourmetric_lower}$$
#
# Given ADM variables $\left\{\gamma_{ij},\beta^i,\alpha \right\}$, which themselves may be written in terms of the rescaled BSSN curvilinear variables $\left\{h_{ij},{\rm cf},\mathcal{V}^i,\alpha \right\}$ for our chosen reference metric via simple function calls to `ADM_in_terms_of_BSSN()` and `BSSN_quantities.BSSN_basic_tensors()`, we are to construct the 4-metric $g_{\mu\nu}$.
#
# We accomplish this via Eq. 2.122 (which can be trivially derived from the ADM 3+1 line element) of Baumgarte & Shapiro's *Numerical Relativity* (henceforth B&S):
# $$
# g_{\mu\nu} = \begin{pmatrix}
# -\alpha^2 + \beta^k \beta_k & \beta_i \\
# \beta_j & \gamma_{ij}
# \end{pmatrix},
# $$
# where the shift vector $\beta^i$ is lowered via (Eq. 2.121):
#
# $$\beta_k = \gamma_{ik} \beta^i.$$
def g4DD_ito_BSSN_or_ADM(inputvars):
# Step 0: Declare g4DD as globals, to make interfacing with other modules/functions easier
global g4DD
# Step 1: Check that inputvars is set to a supported value
gammaDD,betaU,alpha = setup_ADM_quantities(inputvars)
# Step 2: Compute g4DD = g_{mu nu}:
# To get \gamma_{\mu \nu} = gamma4DD[mu][nu], we'll need to construct the 4-metric, using Eq. 2.122 in B&S:
g4DD = ixp.zerorank2(DIM=4)
# Step 2.a: Compute beta_i via Eq. 2.121 in B&S
betaD = ixp.zerorank1()
for i in range(3):
for j in range(3):
betaD[i] += gammaDD[i][j] * betaU[j]
# Step 2.b: Compute beta_i beta^i, the beta contraction.
beta2 = sp.sympify(0)
for i in range(3):
beta2 += betaU[i] * betaD[i]
# Step 2.c: Construct g4DD via Eq. 2.122 in B&S
g4DD[0][0] = -alpha ** 2 + beta2
for mu in range(1, 4):
g4DD[mu][0] = g4DD[0][mu] = betaD[mu - 1]
for mu in range(1, 4):
for nu in range(1, 4):
g4DD[mu][nu] = gammaDD[mu - 1][nu - 1]
# <a id='admbssn_to_fourmetric_inv'></a>
#
# ## Step 2.b: Inverse 4-metric $g^{\mu\nu}$ in terms of ADM or BSSN variables \[Back to [top](#toc)\]
# $$\label{admbssn_to_fourmetric_inv}$$
#
# B&S also provide a convenient form for the inverse 4-metric (Eq. 2.119; also Eq. 4.49 in [Gourgoulhon](https://arxiv.org/pdf/gr-qc/0703035.pdf)):
# $$
# g^{\mu\nu} = \gamma^{\mu\nu} - n^\mu n^\nu =
# \begin{pmatrix}
# -\frac{1}{\alpha^2} & \frac{\beta^i}{\alpha^2} \\
# \frac{\beta^i}{\alpha^2} & \gamma^{ij} - \frac{\beta^i\beta^j}{\alpha^2}
# \end{pmatrix},
# $$
# where the unit normal vector to the hypersurface is given by $n^{\mu} = \left(\alpha^{-1},-\beta^i/\alpha\right)$.
def g4UU_ito_BSSN_or_ADM(inputvars):
# Step 0: Declare g4UU as globals, to make interfacing with other modules/functions easier
global g4UU
# Step 1: Check that inputvars is set to a supported value
gammaDD,betaU,alpha = setup_ADM_quantities(inputvars)
# Step 2: Compute g4UU = g_{mu nu}:
# To get \gamma^{\mu \nu} = gamma4UU[mu][nu], we'll need to use Eq. 2.119 in B&S.
g4UU = ixp.zerorank2(DIM=4)
# Step 3: Construct g4UU = g^{mu nu}
# Step 3.a: Compute gammaUU based on provided gammaDD:
gammaUU, gammaDET = ixp.symm_matrix_inverter3x3(gammaDD)
# Then evaluate g4UU:
g4UU = ixp.zerorank2(DIM=4)
g4UU[0][0] = -1 / alpha**2
for mu in range(1,4):
g4UU[0][mu] = g4UU[mu][0] = betaU[mu-1]/alpha**2
for mu in range(1,4):
for nu in range(1,4):
g4UU[mu][nu] = gammaUU[mu-1][nu-1] - betaU[mu-1]*betaU[nu-1]/alpha**2
# <a id='validationcontraction'></a>
#
# ## Step 2.c: Validation check: Confirm $g_{\mu\nu}g^{\mu\nu}=4$ \[Back to [top](#toc)\]
# $$\label{validationcontraction}$$
#
# Next we compute $g^{\mu\nu} g_{\mu\nu}$ as a validation check. It should equal 4:
g4DD_ito_BSSN_or_ADM("BSSN")
g4UU_ito_BSSN_or_ADM("BSSN")
sum = 0
for mu in range(4):
for nu in range(4):
sum += g4DD[mu][nu]*g4UU[mu][nu]
if sp.simplify(sum) == sp.sympify(4):
print("TEST PASSED!")
else:
print("TEST FAILED: "+str(sum)+" does not apparently equal 4.")
sys.exit(1)
# <a id='fourmetric_to_admbssn'></a>
#
# # Step 3: Write ADM/BSSN metric quantities in terms of 4-metric $g_{\mu\nu}$ (Excludes extrinsic curvature $K_{ij}$, the BSSN $a_{ij}$, $K$, and $\lambda^i$) \[Back to [top](#toc)\]
# $$\label{fourmetric_to_admbssn}$$
#
# Given $g_{\mu\nu}$, we now compute ADM/BSSN metric quantities, excluding extrinsic curvature.
#
# Let's start by computing the ADM quantities in terms of the 4-metric $g_{\mu\nu}$
#
# Recall that
# $$
# g_{\mu\nu} = \begin{pmatrix}
# -\alpha^2 + \beta^k \beta_k & \beta_i \\
# \beta_j & \gamma_{ij}
# \end{pmatrix}.
# $$
#
# From this equation we immediately obtain $\gamma_{ij}$. However we need $\beta^i$ and $\alpha$. After computing the inverse of $\gamma_{ij}$, $\gamma^{ij}$, we raise $\beta_j$ via $\beta^i=\gamma^{ij} \beta_j$ and then compute $\alpha$ via $\alpha = \sqrt{\beta^k \beta_k - g_{00}}$. To convert to BSSN variables $\left\{h_{ij},{\rm cf},\mathcal{V}^i,\alpha \right\}$, we need only convert from ADM via function calls to [`BSSN.BSSN_in_terms_of_ADM`](../edit/BSSN/BSSN_in_terms_of_ADM.py) ([**tutorial**](Tutorial-BSSN_in_terms_of_ADM.ipynb)).
def BSSN_or_ADM_ito_g4DD(inputvars):
# Step 0: Declare output variables as globals, to make interfacing with other modules/functions easier
if inputvars == "ADM":
global gammaDD,betaU,alpha
elif inputvars == "BSSN":
global hDD,cf,vetU,alpha
else:
print("inputvars = " + str(inputvars) + " not supported. Please choose ADM or BSSN.")
sys.exit(1)
# Step 1: declare g4DD as symmetric rank-4 tensor:
g4DD = ixp.declarerank2("g4DD","sym01",DIM=4)
# Step 2: Compute gammaDD & betaD
betaD = ixp.zerorank1()
gammaDD = ixp.zerorank2()
for i in range(3):
betaD[i] = g4DD[0][i]
for j in range(3):
gammaDD[i][j] = g4DD[i+1][j+1]
# Step 3: Compute betaU
# Step 3.a: Compute gammaUU based on provided gammaDD
gammaUU, gammaDET = ixp.symm_matrix_inverter3x3(gammaDD)
# Step 3.b: Use gammaUU to raise betaU
betaU = ixp.zerorank1()
for i in range(3):
for j in range(3):
betaU[i] += gammaUU[i][j]*betaD[j]
# Step 4: Compute alpha = sqrt(beta^2 - g_{00}):
# Step 4.a: Compute beta^2 = beta^k beta_k:
beta_squared = sp.sympify(0)
for k in range(3):
beta_squared += betaU[k]*betaD[k]
# Step 4.b: alpha = sqrt(beta^2 - g_{00}):
alpha = sp.sqrt(sp.simplify(beta_squared) - g4DD[0][0])
# Step 5: If inputvars == "ADM", we are finished. Return.
if inputvars == "ADM":
return
# Step 6: If inputvars == "BSSN", convert ADM to BSSN & return hDD, cf,
import BSSN.BSSN_in_terms_of_ADM as BitoA
dummyBU = ixp.zerorank1()
BitoA.gammabarDD_hDD( gammaDD)
BitoA.cf_from_gammaDD(gammaDD)
BitoA.betU_vetU( betaU,dummyBU)
hDD = BitoA.hDD
cf = BitoA.cf
vetU = BitoA.vetU
# <a id='adm_ito_fourmetric_validate'></a>
#
# ## Step 3.a: ADM in terms of 4-metric validation: Confirm $\gamma_{ij}\gamma^{ij}=3$ \[Back to [top](#toc)\]
# $$\label{adm_ito_fourmetric_validate}$$
#
# Next we compute $\gamma^{ij} \gamma_{ij}$ as a validation check. It should equal 3:
# +
BSSN_or_ADM_ito_g4DD("ADM")
gammaUU, gammaDET = ixp.symm_matrix_inverter3x3(gammaDD)
sum = sp.sympify(0)
for i in range(3):
for j in range(3):
sum += gammaDD[i][j]*gammaUU[i][j]
if sp.simplify(sum) == sp.sympify(3):
print("TEST PASSED!")
else:
print("TEST FAILED: "+str(sum)+" does not apparently equal 3.")
sys.exit(1)
# -
# <a id='bssn_ito_fourmetric_validate'></a>
#
# ## Step 3.b: BSSN in terms of 4-metric validation: Confirm $\bar{\gamma}_{ij}\bar{\gamma}^{ij}=3$ \[Back to [top](#toc)\]
# $$\label{bssn_ito_fourmetric_validate}$$
#
# Next we compute $\bar{\gamma}_{ij}\bar{\gamma}^{ij}$ as a validation check. It should equal 3:
# +
import reference_metric as rfm
par.set_parval_from_str("reference_metric::CoordSystem","SinhCylindrical")
rfm.reference_metric()
BSSN_or_ADM_ito_g4DD("BSSN")
gammabarDD = ixp.zerorank2()
for i in range(3):
for j in range(3):
# gammabar_{ij} = h_{ij}*ReDD[i][j] + gammahat_{ij}
gammabarDD[i][j] = hDD[i][j] * rfm.ReDD[i][j] + rfm.ghatDD[i][j]
gammabarUU, gammabarDET = ixp.symm_matrix_inverter3x3(gammabarDD)
sum = sp.sympify(0)
for i in range(3):
for j in range(3):
sum += gammabarDD[i][j]*gammabarUU[i][j]
if sp.simplify(sum) == sp.sympify(3):
print("TEST PASSED!")
else:
print("TEST FAILED: "+str(sum)+" does not apparently equal 3.")
sys.exit(1)
# -
# <a id='code_validation'></a>
#
# ## Step 4: Code Validation against `BSSN.ADMBSSN_tofrom_4metric` NRPy+ module \[Back to [top](#toc)\]
# $$\label{code_validation}$$
#
# Here, as a code validation check, we verify agreement in the SymPy expressions for BrillLindquist initial data between
# 1. this tutorial and
# 2. the NRPy+ [BSSN.ADMBSSN_tofrom_4metric](../edit/BSSN/ADMBSSN_tofrom_4metric.py) module.
#
# By default, we analyze these expressions in SinhCylindrical coordinates, though other coordinate systems may be chosen.
# +
par.set_parval_from_str("reference_metric::CoordSystem","SinhCylindrical")
rfm.reference_metric()
import BSSN.ADMBSSN_tofrom_4metric as AB4m
for inputvars in ["BSSN","ADM"]:
g4DD_ito_BSSN_or_ADM(inputvars)
AB4m.g4DD_ito_BSSN_or_ADM(inputvars)
for i in range(4):
for j in range(4):
print(inputvars+" input: g4DD["+str(i)+"]["+str(j)+"] - g4DD_mod["+str(i)+"]["
+str(j)+"] = "+str(g4DD[i][j]-AB4m.g4DD[i][j]))
g4UU_ito_BSSN_or_ADM(inputvars)
AB4m.g4UU_ito_BSSN_or_ADM(inputvars)
for i in range(4):
for j in range(4):
print(inputvars+" input: g4UU["+str(i)+"]["+str(j)+"] - g4UU_mod["+str(i)+"]["
+str(j)+"] = "+str(g4UU[i][j]-AB4m.g4UU[i][j]))
BSSN_or_ADM_ito_g4DD("BSSN")
AB4m.BSSN_or_ADM_ito_g4DD("BSSN")
print("BSSN QUANTITIES (ito 4-metric g4DD)")
print("cf - mod_cf = " + str(cf - AB4m.cf))
print("alpha - mod_alpha = " + str(alpha - AB4m.alpha))
for i in range(3):
print("vetU["+str(i)+"] - mod_vetU["+str(i)+"] = " + str(vetU[i] - AB4m.vetU[i]))
for j in range(3):
print("hDD["+str(i)+"]["+str(j)+"] - mod_hDD["+str(i)+"]["+str(j)+"] = "
+ str(hDD[i][j] - AB4m.hDD[i][j]))
BSSN_or_ADM_ito_g4DD("ADM")
AB4m.BSSN_or_ADM_ito_g4DD("ADM")
print("ADM QUANTITIES (ito 4-metric g4DD)")
print("alpha - mod_alpha = " + str(alpha - AB4m.alpha))
for i in range(3):
print("betaU["+str(i)+"] - mod_betaU["+str(i)+"] = " + str(betaU[i] - AB4m.betaU[i]))
for j in range(3):
print("gammaDD["+str(i)+"]["+str(j)+"] - mod_gammaDD["+str(i)+"]["+str(j)+"] = "
+ str(gammaDD[i][j] - AB4m.gammaDD[i][j]))
# -
# <a id='latex_pdf_output'></a>
#
# # Step 4: Output this module to $\LaTeX$-formatted PDF file \[Back to [top](#toc)\]
# $$\label{latex_pdf_output}$$
#
# The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename [Tutorial-ADMBSSN_tofrom_4metric.pdf](Tutorial-ADMBSSN_tofrom_4metric.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means.)
# !jupyter nbconvert --to latex --template latex_nrpy_style.tplx Tutorial-ADMBSSN_tofrom_4metric.ipynb
# !pdflatex -interaction=batchmode Tutorial-ADMBSSN_tofrom_4metric.tex
# !pdflatex -interaction=batchmode Tutorial-ADMBSSN_tofrom_4metric.tex
# !pdflatex -interaction=batchmode Tutorial-ADMBSSN_tofrom_4metric.tex
# !rm -f Tut*.out Tut*.aux Tut*.log
|
Tutorial-ADMBSSN_tofrom_4metric.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import random
import math
import csv
import time
import os
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from matplotlib.ticker import (MultipleLocator, AutoMinorLocator)
from sklearn.neural_network import MLPRegressor
from sklearn.model_selection import GridSearchCV
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import MinMaxScaler,StandardScaler
from sklearn.metrics import accuracy_score,top_k_accuracy_score, mean_squared_error, mean_absolute_error
from sklearn.calibration import CalibratedClassifierCV
from sklearn.multioutput import MultiOutputRegressor
from helper_func import prediction_plot,data_prep
from joblib import dump, load
numexpr=100000
noise=0.01
csv_path=f"./data/train_data{numexpr}_noise{noise}.csv"
df = pd.read_csv(csv_path,index_col=0)
#Data preparation step
df,X,y, X_train, X_test, y_train, y_test,scaled_X_train,scaled_X_test=data_prep(df)
SVM_REG= load('models/SVM_REGR.joblib')
#Calcualte predictions based on test data
start = time.time()
y_pred_fin=SVM_REG.predict(scaled_X_test)
end = time.time()
print('SVM predicting taken ', end-start,' seconds')
#Display accuracy
# Evaluate the regressor
mse_one = mean_squared_error((y_test['mean']-1)/10, (y_pred_fin[:,0]-1)/10)
mse_two = mean_squared_error(y_test['sd']*4, y_pred_fin[:,1]*4)
print(f'MSE for first regressor: {mse_one} - second regressor: {mse_two}')
mae_one = mean_absolute_error((y_test['mean']-1)/10, (y_pred_fin[:,0]-1)/10)
mae_two = mean_absolute_error(y_test['sd']*4, y_pred_fin[:,1]*4)
print(f'MAE for first regressor: {mae_one} - second regressor: {mae_two}')
#Plot diameter prediction vs actual
prediction_plot(real_val=(y_test["mean"].values-1)/10,predic_val=(y_pred_fin[:,0]-1)/10,ax_lim_low=0.1,ax_lim_high=1,majr_tick=0.1,mnr_tick=0.05,ax_label='Mean Diameter')
#Plot estiamted vs real standard deviations
prediction_plot(real_val=y_test["sd"].values*4,predic_val=y_pred_fin[:,1]*4,ax_lim_low=0.01,ax_lim_high=1,majr_tick=0.1,mnr_tick=0.05,ax_label='SD')
|
Deploy_SVM_REG.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
# +
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import gpplot as gpp
import anchors
from poola import core as pool
import core_functions as fns
gpp.set_aesthetics(palette='Set2')
# -
# ## Functions
# +
#QC
def add_guide_seq_col(df, guide_col = 'sgRNA'):
df_match = df.copy()
guide_seq_list = []
for i, row in enumerate(df_match.loc[:,'sgRNA']):
split_row = row.split('_')
guide_seq =split_row[-1]
guide_seq_list.append(guide_seq)
df_match['match_guide'] = pd.Series(guide_seq_list)
return df_match
def get_grouped_controls(df, control_name):
'''
Inputs:
1. df: guide-gene annotation df
2. control_name: list of string identifiers for controls
Outputs:
1. grouped_genes: controls grouped into pseudogenes with
'''
ungrouped_df = pd.DataFrame()
for control in control_name:
control_condition = (df['Gene Symbol'].str.contains(control, na=False))
control_rows = df[control_condition]
#if all controls have same Gene Symbol
if len(set(control_rows['Gene Symbol'])) == 1:
ungrouped_df = df.copy()
# Give controls unique names before grouping
ungrouped_df.loc[control_condition, 'Gene Symbol'] = control + control_rows['Guide']
ungrouped_genes = ungrouped_df.dropna()
grouped_genes = pool.group_pseudogenes(ungrouped_genes[['Guide', 'Gene Symbol']], pseudogene_size=3, gene_col='Gene Symbol', control_regex = control_name)
return grouped_genes
def match_pseudogenes(df, guide_mapping, control_name):
'''
Inputs:
1. df: data frame to which grouped pseudogene annotations will be merged
2. guide_mapping: guide-gene annotation data frame
3. control_name: list containing substrings that identify controls
'''
grouped_genes = get_grouped_controls(guide_mapping, control_name)
grouped_df = pd.merge(grouped_genes, df, on = 'Guide', how = 'outer', suffixes = ['', '_old'])
grouped_df.loc[:,'Gene Symbol'] = grouped_df.loc[:,'Gene Symbol'].fillna(grouped_df['Gene Symbol_old'])
grouped_df = grouped_df.drop('Gene Symbol_old', axis = 1)
return grouped_df
# -
def run_guide_residuals(lfc_df, paired_lfc_cols):
'''
Calls get_guide_residuals function from anchors package to calculate guide-level residual z-scores
Input:
1. lfc_df: data frame with log-fold changes (relative to pDNA)
'''
modified = []
unperturbed = []
#reference_df: column1 = modifier condition, column2 = unperturbed column
ref_df = pd.DataFrame(columns=['modified', 'unperturbed'])
row = 0 #row index for reference df
for pair in paired_lfc_cols:
#number of resistant pops in pair = len(pair)-1
res_idx = 1
#if multiple resistant populations, iterate
while res_idx < len(pair):
ref_df.loc[row, 'modified'] = pair[res_idx]
ref_df.loc[row, 'unperturbed'] = pair[0]
res_idx +=1
row +=1
print(ref_df)
#input lfc_df, reference_df
#guide-level
residuals_lfcs, all_model_info, model_fit_plots = anchors.get_guide_residuals(lfc_df, ref_df)
return residuals_lfcs, all_model_info, model_fit_plots
# ## Data Summary
#
# * Cell line: Calu-3
# * Library: Gattinara
# +
reads_act = pd.read_csv('../../Data/Reads/Hsu/Calu3_Activation.txt', sep = '\t', error_bad_lines=False)
pDNA_reads_CalSetA = pd.read_excel('../../Data/Reads/Goujon/Calu3/Calabrese/CalabreseSetApDNAReads.xlsx', sheet_name= 'SetA pXPR_109 raw reads', header = 3)
pDNA_reads_CalSetA = pDNA_reads_CalSetA.copy()[['sgRNA Sequence', 'pDNA']]
reads_act_match = add_guide_seq_col(reads_act)
reads_all = pd.merge(pDNA_reads_CalSetA, reads_act_match, left_on = 'sgRNA Sequence', right_on = 'match_guide', how = 'outer')
# Columns to include
# Activation:
# Treatment: L-SAM_D5_r1, M-SAM_D5_r2, R4, R5
# Control: J-SAM_D0_r1, K-SAM_D0_r2, R4C, R5C
treatment_cols = ['L-SAM_D5_r1', 'M-SAM_D5_r2', 'R4', 'R5']
control_cols = ['J-SAM_D0_r1', 'K-SAM_D0_r2', 'R4C', 'R5C']
base_cols = ['sgRNA', 'Gene', 'pDNA']
cols_to_include = base_cols + control_cols + treatment_cols
reads = reads_all[cols_to_include]
reads
reads = reads.rename(columns={'Gene':'Gene Symbol'})
reads = reads.rename(columns={'sgRNA':'Guide'})
reads = match_pseudogenes(reads.copy(), reads.copy()[['Guide', 'Gene Symbol']], control_name=['NO-TARGET'])
reads
# +
#Calculate lognorm
cols = reads.columns[2:].to_list() #reads columns = start at 3rd column
lognorms = fns.get_lognorm(reads.dropna(), cols = cols)
lognorms
# -
# ## Quality Control
# ### Population Distributions
# +
#Calculate log-fold change relative to pDNA
target_cols = list(lognorms.columns[3:])
pDNA_lfc = fns.calculate_lfc(lognorms,target_cols)
# Pair cols
paired_cols = []
i = 0
j = 0
for i, t in enumerate(treatment_cols):
for j ,c in enumerate(control_cols):
if i == j:
c_new = c + '_lfc'
t_new = t + '_lfc'
pair = [c_new, t_new]
paired_cols.append(pair)
else:
continue
paired_cols = True, paired_cols
#Plot population distributions of log-fold changes
fns.lfc_dist_plot(pDNA_lfc, paired_cols=paired_cols, filename = 'Calu3_Act_Hsu', figsize=(6, 12))
# -
# ### Control Distributions
fns.control_dist_plot(pDNA_lfc, paired_cols=paired_cols, control_name=['NO-TARGET'], filename='Calu3_Act_Hsu', figsize=(6, 12))
# ## Gene-level analysis
# ### Residual z-scores
# +
#Calculate z-scores of lfc residuals
lfc_df = pDNA_lfc.copy().drop(['Gene Symbol'], axis = 1)
lfc_df = lfc_df.dropna()
lfc_df = lfc_df.drop_duplicates()
paired_lfc_cols = paired_cols[1]
guide_residuals_lfcs, all_model_info, model_fit_plots = run_guide_residuals(lfc_df, paired_lfc_cols)
guide_mapping = pDNA_lfc[['Guide', 'Gene Symbol']]
gene_residuals = anchors.get_gene_residuals(guide_residuals_lfcs, guide_mapping)
# -
gene_residual_sheet = fns.format_gene_residuals(gene_residuals, guide_min=2, guide_max=4, ascending=True)
guide_residual_sheet = pd.merge(guide_mapping, guide_residuals_lfcs, on = 'Guide', how = 'outer')
with pd.ExcelWriter('../../Data/Processed/GEO_submission_v2/Calu3_Act_Hsu_v1.xlsx') as writer:
gene_residual_sheet.to_excel(writer, sheet_name='Calu3_KO_avg_zscore', index =False)
reads.to_excel(writer, sheet_name='Calu3_KO_genomewide_reads', index =False)
guide_mapping.to_excel(writer, sheet_name='Calu3_KO_guide_mapping', index =False)
with pd.ExcelWriter('../../Data/Processed/Individual_screens_v2/Calu3_Act_Hsu_indiv_v1.xlsx') as writer:
gene_residuals.to_excel(writer, sheet_name='condition_genomewide_zscore', index =False)
guide_residual_sheet.to_excel(writer, sheet_name='guide-level_zscore', index =False)
# ## Compare to Goujon data
# +
from adjustText import adjust_text
# Read Goujon Gattinara processed data
Goujon_Calu3_Cal = pd.read_excel('../../Data/Processed/GEO_submission_v2/Calu3_Calabrese_Goujon_v3.xlsx')
HsuvsGoujon_Calu3_act = pd.merge(Goujon_Calu3_Cal, gene_residual_sheet, on = 'Gene Symbol', suffixes = ['_Goujon', '_Hsu'])
fig, ax = plt.subplots(figsize=(2,2))
ax = gpp.point_densityplot(HsuvsGoujon_Calu3_act, 'residual_zscore_avg_Goujon', 'residual_zscore_avg_Hsu', s=6)
ax = gpp.add_correlation(data=HsuvsGoujon_Calu3_act, x='residual_zscore_avg_Goujon', y='residual_zscore_avg_Hsu', loc='upper left', fontsize = 7)
top_ranked_Goujon = HsuvsGoujon_Calu3_act.nlargest(20, 'residual_zscore_avg_Goujon')
top_ranked_Hsu = HsuvsGoujon_Calu3_act.nlargest(20, 'residual_zscore_avg_Hsu')
top_ranked_Vero = pd.concat([top_ranked_Goujon, top_ranked_Hsu]).reset_index(drop = True).drop_duplicates()
bottom_ranked_Goujon = HsuvsGoujon_Calu3_act.nsmallest(20, 'residual_zscore_avg_Goujon')
bottom_ranked_Hsu = HsuvsGoujon_Calu3_act.nsmallest(20, 'residual_zscore_avg_Hsu')
bottom_ranked_Vero = pd.concat([bottom_ranked_Goujon, bottom_ranked_Hsu]).reset_index(drop = True).drop_duplicates()
ranked_Goujon = pd.concat([top_ranked_Goujon, bottom_ranked_Goujon])
ranked_Hsu = pd.concat([top_ranked_Hsu, bottom_ranked_Hsu])
# Label gene hits common to both screens
common_ranked_Calu3_act = pd.merge(ranked_Goujon, ranked_Hsu, on =['Gene Symbol', 'residual_zscore_avg_Goujon', 'residual_zscore_avg_Hsu'], how = 'inner')
# ranked_Vero = pd.concat([top_ranked_Vero, bottom_ranked_Vero])
sns.scatterplot(data = common_ranked_Calu3_act, x='residual_zscore_avg_Goujon', y='residual_zscore_avg_Hsu', color = sns.color_palette('Set2')[0], edgecolor=None, s=6, rasterized=True)
texts= []
for j, row in common_ranked_Calu3_act.iterrows():
texts.append(ax.text(row['residual_zscore_avg_Goujon']+0.25, row['residual_zscore_avg_Hsu'], row['Gene Symbol'], fontsize=7,
color = 'black'))
# ensures text labels are non-overlapping
adjust_text(texts)
plt.title('Calu-3 Activation Hsu vs Goujon', fontsize=7)
plt.xlabel('Mean z-score (Goujon)', fontsize=7)
plt.ylabel('Mean z-score (Hsu)', fontsize=7)
plt.xticks(fontsize=7)
plt.yticks(fontsize=7)
ax.set_box_aspect(1)
sns.despine()
gpp.savefig('../../Figures/Scatterplots/Calu3act_HsuvsGoujon_scatterplot.pdf', dpi=300)
# -
|
PrimaryScreens/Hsu_Calu3_Activation_v1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
# +
doi_df_journal = pd.DataFrame({
"DOI": [
"10.1371/journal.pcbi.1004668",
"10.21105/joss.01035",
"10.12688/f1000research.18866.2",
"10.1038/s41598-019-52881-4",
"10.1186/s12859-019-3171-0"],
"journal": [
"PLoS Comput Biol",
"JOSS",
"F1000Res",
"Sci Rep",
"BMC Bioinformatics]"]})
doi_df_first_author = pd.DataFrame({
"DOI": [
"10.1371/journal.pcbi.1004668",
"10.21105/joss.01035",
"10.21105/joss.01006",
"10.12688/f1000research.18866.2",
"10.1186/s12859-019-3171-0"],
"first_author": [
"Blischak",
"Sparks",
"Granger",
"Thang",
"Chen]"]})
# -
doi_df_journal
doi_df_first_author
doi_df_first_author.merge(doi_df_journal, on="DOI")
doi_df_first_author.merge(doi_df_journal, on="DOI", how="left")
doi_df_first_author.merge(doi_df_journal, on="DOI", how="right")
doi_df_first_author.merge(doi_df_journal, on="DOI", how="inner")
doi_df_first_author.merge(doi_df_journal, on="DOI", how="outer")
openapc = pd.read_csv("https://raw.githubusercontent.com/OpenAPC/openapc-de/master/data/fuberlin/APC_FU_Berlin_2015.csv")
openapc
openapc[openapc.euro >= 2000.0]
openapc["high_price"] = openapc["euro"] * 1000000
openapc
# +
def replace_berlin(cell):
return cell.replace("Berlin", "Buxtehude")
openapc["new_insitution"] = openapc["institution"].apply(replace_berlin)
# -
openapc
|
code/Pandas_merge_und_apply.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import csv
import matplotlib.pyplot as plt
import matplotlib.patches as patches
from PIL import Image
def parsetxt(filename):
with open(filename) as f:
defectTypes = {}
for i in range(3):
f.readline()
line = f.readline()
defectTypes[i] = line.strip().split()
for j in range(len(defectTypes[i])):
defectTypes[i][j] = int(defectTypes[i][j])
f.readline()
return defectTypes
def parsecsv(filename, defectTypes):
labels = {}
with open(filename) as csvfile:
reader = csv.reader(csvfile)
temp = 0
for row in reader:
if (temp == 0):
temp = 1
if (
row[5] != 'X' or
row[6] != 'Y' or
row[11] != 'Width' or
row[12] != 'Height'
):
continue
#raise ValueError()
else:
label = int(row[0])
X = float(row[5])
Y = float(row[6])
width = int(row[11])
height = int(row[12])
defectType = 5
if ( (len(defectTypes[0]) != 0) and (defectTypes[0][0]) <= label <= (defectTypes[0][1])):
defectType = 0
elif ((len(defectTypes[1]) != 0) and (defectTypes[1][0]) <= label <= (defectTypes[1][1])):
defectType = 1
elif ((len(defectTypes[2]) != 0) and (defectTypes[2][0]) <= label <= (defectTypes[2][1])):
defectType = 2
labels[label] = [defectType, X, Y, width, height]
return labels
def labelImage(imagename, labels):
im = Image.open(imagename)
fig, ax = plt.subplots(1)
ax.imshow(im)
for i in range(1,len(labels)+1):
x = labels[i][1] - labels[i][3] / 2 - 10
y = labels[i][2] - labels[i][4] / 2 - 10
width = labels[i][3] + 20
height = labels[i][4] + 20
if(labels[i][0] == 0):
color = 'r'
elif(labels[i][0] == 1):
color = 'g'
elif(labels[i][0] == 2):
color = 'b'
else:
color = 'y'
rect = patches.Rectangle((x, y), width, height, linewidth = 1, edgecolor = color, facecolor = 'none')
ax.add_patch(rect)
ax.text(x, y, i)
plt.show()
def calculateBoundingBoxes(defect, x,y, w,h):
X1 = x -(w/2)
Y1 = y - (h/2)
X2 = x + (w/2)
Y2 = y + (h/2)
return [defect, round(X1,2), round(Y1,2), round(X2,2), round(Y2,2)]
with open('filenames.txt') as f:
filename = f.readline().strip('\n')
print(filename)
while filename:
print(filename)
filetxt = 'logs/' + filename + '_log.txt'
filecsv = 'results/' + filename + '_results.csv'
count0 = 0
count1 = 0
count2 = 0
defectTypes_gt = parsetxt(filetxt)
#print(defectTypes_gt)
labels_gt = parsecsv(filecsv, defectTypes_gt)
outfile = 'bounding_boxes/' + filename + '.txt'
fout = open(outfile, mode = 'w+')
for i in range(1, len(labels_gt) + 1):
if(labels_gt[i][0] == 0 or labels_gt[i][0] == 1 or labels_gt[i][0] == 2 ):
if (labels_gt[i][0] == 0):
count0 += 1
elif (labels_gt[i][0] == 1):
count1 += 1
elif (labels_gt[i][0] == 2):
count2 += 1
mylist = calculateBoundingBoxes(labels_gt[i][0], labels_gt[i][1], labels_gt[i][2], labels_gt[i][3], labels_gt[i][4])
fout.write(str(mylist[0]) +' ' + str(mylist[1]) + ' ' + str(mylist[2]) + ' ' + str(mylist[3]) + ' '+ str(mylist[4]) + '\n')
fout.close()
print(str(count0) + ' ' + str(count1) + ' ' + str(count2))
filename = f.readline().strip('\n')
|
Archive/TF_VERSION_MASKRCNN/datasets/ellipse/defectextraction/Data3TypesYminXminYmaxXmax5/BoundingBoxes.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Loops and Control Structures
#
# We will study these two topics together, because they are so commonly found in association with one another. Let's say you want to go through your list of students, and only print the names of students older than 25. How?
#
# We will how look at "iteration" over both List and Hash data structrues.
#
# ## Iteration - "looping"
#
# "looping" (iteration) means to do the same thing many times. Imagine this code:
#
# mylist = [123, 334, 223, 197, 901, 344]
# print mylist[0]
# print mylist[1]
# print mylist[2]
# print mylist[3]
# print mylist[4]
# print mylist[5]
#
# Why is this bad? (discuss... there are many reasons!)
#
# This is why we need Loops - they "abstract" the problem of doing the same thing, on a different value, some arbitrary number of times.
#
# ### while , for
#
# The two basic kinds of iterations are "while" and "for"
#
# * "while" means "while a certain condition is true"
# * "for" means "for a certain number of times"
#
# They look like this:
#
# while CONDTION:
# do something here
#
#
# for ITERATING:
# do something here
#
# Here is an example of while and for loops:
#
# +
myage = 0
while myage < 18: # note the ":" colon character!
print("I am still young, at ", myage)
myage = myage + 1
#myage+=1
print("Now that I am ", myage, " I am old!")
# +
myage = 0
for myage in range(1,18): # note the ":", and the use of "in" to assign the next value to the variable
print("I am ", myage, " years old")
print("at the end I am ", myage, " years old")
# -
# ## if/else
#
# Sometimes, you want to do something if a condition is true, but do something else if the condition is false. For that, you can use the "else" statement. It looks like this:
#
myage = 0
for myage in range(1,18): # note the ":", and the use of "in" to assign the next value to the variable
if myage % 2 == 0: # % is the modulo operator - gives you the remainder when dividing myage by 2
# note that the comparison operator is "==". IT IS NOT "=" (= is used to assign a value!)
# see end of Lesson 4 to review the comparison and mathematical operators!
print("I am", myage, "my age is an even number")
else:
print("I am", myage, "my age is an odd number")
#
# ### More complex condtionals - elif
#
# if this, else if (elif) that, else the other thing.
#
# you can use as many elif's as you want!
#
# <pre>
#
#
# </pre>
myage = 0
for myage in range(1,18): # note the ":", and the use of "in" to assign the next value to the variable
if (myage % 2 == 0) & (myage % 4 == 0):
print("I am", myage, "my age is divisible by BOTH 2 (logical AND!) 4")
elif myage % 4 == 0: # note the order of my "if" statements... what happens if I switch the %2 and the %4? Try!
print("I am", myage, "my age is divisible by 4")
elif myage % 2 == 0:
print("I am", myage, "my age is divisible by 2")
else:
print("I am", myage, "my age is an odd number")
#con elif no se hacen todos los if, como si en vez de elif fueran if
#NOTE: this is a surprisingly common interview question for a data scientist!!!!
#
# # DANGER!!!!!!!!
#
#
#
# Note that loops can be dangerous!! For example, in a "while" loop, if the condition is never met, then the loop will never end! This is an "infinite loop", and they can be very very very bad, if your statements are, for example, writing to a file, or interacting with a database!
#
# for example (DO NOT TRY THIS!!!!!!!!)
#
# age = 0
# while age != 21: # != "not equal to" condition will never be met - our age is always an even number
# print(age)
# age += 2 # an abbreviation for "age = age + 2"
#
# You should always be careful that the condition will be met; however, if you cannot be careful, there is a way to do a "sanity check" in your code so that you break the loop under a certain condition. The command is "break", and it also is associated with a conditional phrase:
#
# age = 0
# while (age != 21): # condition will never be met, because our age is always an even number
# print(age)
# age += 2 # an abbreviation for "age = age + 2"
# if age > 100: # set a "sanity check" can't be more than 100!
# break
#
# ---es buena idea siempre poner el brek en los loops
age = 0
while (age != 21): # this condition will never be met, because our age is always an even number!!!!
print(age)
age += 2 # an abbreviation for "age = age + 2"
if age > 30: # once you are 30, your life has ended... break the loop
break
#
# Note that the use of "break" can fool you into thinking that everything was OK! The loop ends, and the program continues to run.... so it isn't PERFECT, but it can at least help minimize damage!
#
# A better way do this is to "raise an exception" - this allows you to send a message explaining what went wrong. In this introductory section, we wont discuss the topic of Exceptions beyond the fact that they exist, and are used as follows:
#
age = 0
while (age != 21): # this condition will never be met, because our age is always an even number!!!!
print(age)
age += 2 # an abbreviation for "age = age + 2"
if age > 30: # once you are 30, your life has ended... break the loop
raise Exception("the age became greater than 30, and this should not happen")
break
# # "Safer" ways to iterate/loop
#
# There are some safe ways to iterate:
#
# 1. using "in" on a list is safer
# 2. using "in" on a list index is safer
#
# for example:
# +
students = ["Mark", "Jonas", "Michele", "Alberto"]
for name in students:
print("student's name is ", name)
# you can determine the length of an array using the "len" function:
print()
print( len(students))
print()
for index in range(len(students)):
print("student #", index, "is named", students[index])
# -
#
# You can do similar things with Dictionaries. Often, you will want to separate your data based on the "keys" of the Dictionary. To do this, use the "keys" method on your dictionary. for example:
#
# students = {"Mark": 50, "John": 45}
# print( students.keys() )
#
students = {"Mark": 50, "John": 45}
print(students.keys())
print(students.values()) # this also works if you want to extract all of the values as a list - uncommon...
#
# Note that the output is telling us that the "keys" method returns us an object of type "dict_keys"... I will simply tell you that dict_keys can be used as an iterator (exactly like "range" is!). So you can say:
#
# +
students = {"Mark": 50, "John": 45}
for student in students.keys():
print("student named", student, "is", students[student], "years old")
|
Lesson 5a - Loops, conditionals, and control structures.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
## Job1: simple wordcount on term-document tuples to compute tf
### Map
# Input: (docname, contents):
# Output: ((term,docname), 1)
### Reduce
# Input: ((term,docname), 1)
# Output: ((term,docname), n)
########################################### mapper1.py
import os
import sys
for line in sys.stdin:
line = line.strip()
terms = line.split(" ")
path = os.environ['mapreduce_map_input_file'].split('/')
docname = path[-1]
for term in terms:
term = term.strip('''!()-[]{};:'"\,<>./?@#$%^&*_~''').lower()
print('%s\t%s' % (term + '_' + docname, 1))
########################################### reducer1.py
import sys
current_pair = None
current_count = 0
pair = None
for line in sys.stdin:
line = line.strip()
pair, count = line.split('\t', 1)
# convert count (currently a string) to int
try:
count = int(count)
except ValueError:
# count was not a number, so silently
# ignore/discard this line
continue
if current_pair == pair:
current_count += count
else:
if current_pair:
# write result to STDOUT
print ('%s\t%s' % (current_pair, current_count))
current_count = count
current_pair = pair
if current_pair == pair:
print ('%s\t%s' % (current_pair, current_count))
# +
## Job2: append document frequency d to term_doc pairs
### Map
# Input: ((term,docname), n)
# Output: (term, (docname,n,1))
### Reduce
# Input: (term, (docname,n,1))
# Output: ((term,docname), (n,d))
########################################### mapper2.py
import sys
for line in sys.stdin:
term,rest = line.split('_')
docname,n = rest.split('\t')
try:
n = int(n)
except ValueError:
continue
print('%s\t%s' % (term, docname + '_' + n + '_' + 1))
########################################### reducer2.py
import sys
current_term = None
doc_list = []
n_list=[]
current_count = 0
term = None
doc = None
for line in sys.stdin:
term,rest = line.split('\t',1)
doc,n,count = rest.split('_', 2)
# convert count (currently a string) to int
try:
count = int(count)
except ValueError:
# count was not a number, so silently
# ignore/discard this line
continue
if current_term == term:
doc_list.append(doc)
n_list.append(n)
current_count += count
else:
if current_term:
for i,document in enumerate(doc_list):
print ('%s\t%s' % (current_term + '_' + document, n_list[i] + '_' + str(current_count)))
doc_list = []
n_list = []
current_count = count
current_term = term
doc_list.append(doc)
n_list.append(n)
if current_term == term:
for i,document in enumerate(doc_list):
print ('%s\t%s' % (current_term + '_' + document, n_list[i] + '_' + str(current_count)))
# +
## Job3: compute tf-idfs
### Map
# Input: ((term,docname), (n,d))
# Output: ((term,docname), tfidf)
### Reduce ----pass
# Input:
# Output:
########################################### mapper3.py
import sys
import math
for line in sys.stdin:
pair,rest = line.split('/t',1)
n,d = rest.split('_',1)
try:
n = int(n)
d = int (d)
except ValueError:
continue
## to calculate these with normalizing terms we need to compute:
# 1. the total of words in each documents
# 2. the total number of documents in the corpus
# look into Hadoop global variables/ counters
tf = n
idf = math.log(1/(1+d))
tfidf = tf * idf
print('%s\t%s' % (pair, tfidf))
# -
minutes Jan 31st meeting:
- computation of Nj and D
-
|
tfidf_mapreduce_DR_v1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # "FixRes: 'Fixing the train-test resolution discrepancy'"
#
# > "'Fixing the train-test resolution discrepancy' from Facebook AI Research, NeurIPS 2019"
#
# - toc: false
# - badges: false
# - comments: true
# - author: <NAME>
# - image: images/20202_04_15_horse_train_224.png
# - categories: [papers, training technique, classification]
# ## TL;DR
# The paper outlines two easy-to-implement tips to improve your image classification test results:
# 1. Do your inference on the test set at a **higher resolution** than your train set
# 2. Fine-tune the **last layers** of your CNN classifier (i.e. the linear layer(s) after your pooling layer) at the higher test resolution
#
# ## Overview
#
# This article is a quick summary of ['Fixing the train-test resolution discrepancy'](https://arxiv.org/abs/1906.06423) from <NAME>, <NAME>, <NAME>, <NAME> from Facebook AI Research, presented at NeurIPS 2019, with additional data from the note ['Fixing the train-test resolution discrepancy: FixEfficientNet'](https://arxiv.org/pdf/2003.08237.pdf) from the same authors
#
#
# ## Results
#
# - Facebook AI Research (FAIR) used this technique to **achieve a new SOTA result on Imagenet** (**`88.5%`** top-1 accuracy) using EfficientNet (using extra data)
#
# 
#
# - The authors also claim that it can **enable faster training** by training at a lower resolution while still attaining similr/better results
# > twitter: https://twitter.com/HugoTouvron/status/1242071277415870470
# ## But Why?
# Using typical training transforms such as `RandomResizedCrop` result in objects in training images appearing **larger** than they do in the test set. Have a look at the example from the paper below.
#
# Our original image is resized to `224 x 224` before it is shown to the model. `RandomResizedCrop` is used to resize our training image (and add a little regularisation) while for the test image a simple center crop is taken. As a result of these different resizing methods, the size of the white horse in the top left training image is much larger than what would be shown to the model in the test set. It is this **difference in object (e.g. horse) size** that the authors say that their FixRes technique addresses
#
# 
#
# In other words:
#
# >...resizing the input images in pre-processing changes the distribution of objects sizes. Since different pre-processing protocols are used at training and testing time, the size distribution differs in the two cases.
#
# ## How? - Two Tricks
# 1. Test at a Higher Resolution
#
# Simply testing at a higher resolution should yield a performance improvement. Here, the authors show ImageNet top-1 test set accuracy trained at `224 x 224`, you can see that the optimal test resolution was `288 x 288`:
#
# 
#
# (This behaviour was previously been shown in 2016 in ["Identity Mappings in Deep Residual Networks"](https://arxiv.org/pdf/1603.05027.pdf)). Alternatively if you don't want to/cannot test at higher resolution, then training at a lower resolution is said to deliver the same accuracy, while **enabling you to train faster** (as you will be able to use a larger batch size with your smaller image resolutions)
#
# 2. Fine-tuning of later (classifier) layers of your CNN model
#
# >For the convolutional part of the CNN, comprising linear convolution, subsampling, ReLU, and similar layers, changing the input crop size is approximately transparent because the receptive field is unaffected by the input size. However, for classification the network must be terminated by a pooling operator (usually average pooling) in order to produce a fixed-size vector. **Changing the size of the input crop strongly affects the activation statistics of this layer**.
#
# When fine-tuning, the authors recommend using **test-time augmentation, not the previous training augmentation** as it is simplest and performs well. Using training augmentations gave only slightly better results.
#
#
#
# ## Similarity to Fast.ai's Progressive Resizing
#
# Interestingly this technique is a little similar to `Progressive Resizing`, first espoused in the [fast.ai](www.fast.ai) deep learning course. The idea behind Progressive Resizing is that you first train at a lower resolution before increasing resolution and training again, albeit you're always training the entire network as opposed fine-tuning the classifier layers as described above. Nevertheless, it makes me wonder if both the FixRes and Progressive Resizing training techniques work via correcting for the same Train/Test object size mis-match?
#
# Any thoughts, comments, suggestions I'd love to hear from you [@mcgenergy](www.twitter.com/mcgenergy) on Twitter 😃
|
_notebooks/2020-04-15-fixres.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# name: python2
# ---
# + [markdown] id="x0xsfKkoFi1H" colab_type="text"
# # Linear Least-Squares Algorithms for Temporal Difference Learning
# + [markdown] id="3BTzn41GF0CA" colab_type="text"
# ### What is Linear Least Squares Temporal Difference Learning?
#
# The linear least-squares function approximation aims to linearly estimate a function given pairwise samples of observed inputs and outputs.. Since we know that TD(0) with linear approximation converges asympotically to a fixed point, LSTD provides means to do this. It accomplishes this by building estimates of C matrix and d vector to directly solve for d + CBeta_{lambda} = 0 directly.
#
# + [markdown] id="dj3H9v-PQdR5" colab_type="text"
# #Installing Pycolab and Gym
# + id="GzDh44mnPndc" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 9}], "base_uri": "https://localhost:8080/", "height": 408} outputId="8c78da99-f35f-40ac-e376-6e843da0ebbe" executionInfo={"status": "ok", "timestamp": 1521656017370, "user_tz": 240, "elapsed": 9229, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-N2vm2KOF7iE/AAAAAAAAAAI/AAAAAAAACL8/3K7KcnaQmY0/s50-c-k-no/photo.jpg", "userId": "111336538593611088506"}}
# ! pip install git+https://github.com/deepmind/pycolab.git
# ! pip install git+https://github.com/openai/gym.git
# + id="FXjTty6nPqP3" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import curses
import sys
import gym
import numpy as np
import time
from pycolab import ascii_art
from pycolab import human_ui
from pycolab.prefab_parts import sprites as prefab_sprites
import matplotlib.pyplot as plt
# + [markdown] id="CbhOKm7aQb-i" colab_type="text"
# # Environment
# + [markdown] id="Ei0kotutoW-2" colab_type="text"
# ## Common Environment Methods
# + id="ArYaF8cBkl-o" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
def obsv2state(obs):
""" Convert pycolab's observation to int state.
The state is flattened in our case and is (0,12).
Args:
obs: Pycolab's Observation object.
Returns:
Integer state from (0,12).
"""
state = np.array(obs.layers['P'], dtype=np.float).flatten()
states = np.flatnonzero(state)
assert len(states) == 1, "There should be just one P."
return states[0]
def displayenv(obs, display_visits=False, stored_chain=None):
""" Display the environment from observation.
Args:
obs: Pycolab's Observation object.
display_visits: Whether to highlight the previously visited state.
stored_chain: Previously stored values for the chain. This is used if displaying the old visits.
Returns:
Returns updated stored_chain to be used in next step to highlight previously visitied state.
"""
chain = 125 * np.array(obs.layers['P'], dtype=np.float) #The agent/player sprite P is indicated by
if display_visits:
current_visits = 15.50 * np.array(obs.layers['P'], dtype=np.float) #Indicates new visits to states
if stored_chain is not None:
stored_chain += current_visits
chain += stored_chain #For visualization
else:
stored_chain = current_visits
if 'G' in obs.layers:
chain += 175 * np.array(obs.layers['G'], dtype=np.float) #Indicates Goal distinctly
plt.figure(figsize=(3,4))
plt.axis('off')
plt.imshow(chain, cmap='Reds')
return stored_chain
# + [markdown] id="9qbJs4mLoiYK" colab_type="text"
# ## Boyan's Chain
# + id="F8_eeTOMQFD0" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
# --------------------------13-State Markov Chain From Boyan----------------------------------#
BOYANCHAIN_STATES = ['P G']
# --------------------------Features for Boyan Chain------------------------------------------#
# For value function approximation, each state is represented by four features, as follows:
# the representations for states 12, 8, 4, and 0 are, respectively, [1, 0, 0, 0], [0, 1, 0, 0],
# [0, 0, 1, 0], and [0, 0, 0, 1]; and the representations for the other states are obtained by
# linearly interpolating between these.
# --------------------------------------------------------------------------------------------#
BOYANCHAIN_FEATURES = np.array([[1.00, 0.00, 0.00, 0.00],
[0.75, 0.25, 0.00, 0.00],
[0.50, 0.50, 0.00, 0.00],
[0.25, 0.75, 0.00, 0.00],
[0.00, 1.00, 0.00, 0.00],
[0.00, 0.75, 0.25, 0.00],
[0.00, 0.50, 0.50, 0.00],
[0.00, 0.25, 0.75, 0.00],
[0.00, 0.00, 1.00, 0.00],
[0.00, 0.00, 0.75, 0.25],
[0.00, 0.00, 0.50, 0.50],
[0.00, 0.00, 0.25, 0.75],
[0.00, 0.00, 0.00, 0.00]])
class PlayerSpriteBoyanChain(prefab_sprites.MazeWalker):
"""A `Sprite` for our player."""
def __init__(self, corner, position, character):
"""Inform superclass that we cant walk through #."""
super(PlayerSpriteBoyanChain, self).__init__(
corner, position, character, impassable='#')
def update(self, actions, board, layers, backdrop, things, the_plot):
del backdrop, things # Unused.
#Get current position in the chain
info, position = self.position
# Apply motion commands.
if actions == 0: # Only one action ==> left to right.
if layers["G"][0, position+1]: # If current state is just adjacent to the goal state Reward = -2
self._east(board, the_plot)
the_plot.add_reward(-2)
else: # From any other 11 states, not adjacent to goal or goal Reward = -3
if (np.random.rand() > 0.5):
self._east(board, the_plot)
self._east(board, the_plot)
else:
self._east(board, the_plot)
the_plot.add_reward(-3)
if layers["G"][self.position]: # If current state is Goal State, Terminate Episode
the_plot.terminate_episode()
class BoyansChain(gym.Env):
def __init__(self):
self.game= ascii_art.ascii_art_to_game(
BOYANCHAIN_STATES,
what_lies_beneath=' ',
sprites={'P': PlayerSpriteBoyanChain})
obs, reward, gamma = self.game.its_showtime()
def step(self):
""" Take given action and return the next state observation and reward.
"""
obs, reward, gamma = self.game.play(0)
return obs, reward, self.game.game_over, ""
def reset(self):
""" Resets the game and returns the observation for the start state.
"""
self.game= ascii_art.ascii_art_to_game(
BOYANCHAIN_STATES,
what_lies_beneath=' ',
sprites={'P': PlayerSpriteBoyanChain})
obs, reward, gamma = self.game.its_showtime()
return obs
# + [markdown] id="Y5zSTVxzobQj" colab_type="text"
# **Testing Boyan Chain Environment**
# + id="ssZvYmc9oe2W" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}, {"item_id": 2}, {"item_id": 3}, {"item_id": 4}, {"item_id": 5}, {"item_id": 6}, {"item_id": 7}, {"item_id": 8}], "base_uri": "https://localhost:8080/", "height": 401} outputId="1c2ebccd-b320-49ab-9f21-cd24c7bec428" executionInfo={"status": "ok", "timestamp": 1521670430886, "user_tz": 240, "elapsed": 818, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "105824936464368788271"}}
game = BoyansChain()
obs = game.reset()
stored_chain = displayenv(obs, display_visits=True)
returnGt = 0
game_over = False
#Play until terminates
while not(game_over):
obs, reward, game_over, _ = game.step()
returnGt += reward
stored_chain = displayenv(obs, display_visits=True, stored_chain=stored_chain)
# + [markdown] id="h6VXczxBQmue" colab_type="text"
# ## Bradtke & Barto
# + id="z3dSY8r-Qwv9" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
# --------------------------5-State Markov Chain From Bradtke----------------------------------#
BRADTKE_STATES = ['P ']
# --------------------------Transition probabilities, reward matrix and features for Bradtke Chain -------#
TRANSITION_MATRIX = np.array([[0.42, 0.13, 0.14, 0.03, 0.28],
[0.25, 0.08, 0.16, 0.35, 0.15],
[0.08, 0.20, 0.33, 0.17, 0.22],
[0.36, 0.05, 0.00, 0.51, 0.07],
[0.17, 0.24, 0.19, 0.18, 0.22]])
REWARD_MATRIX = np.array([[104.66, 29.69, 82.36, 37.49, 68.82],
[ 75.86, 29.24, 100.37, 0.31, 35.99],
[ 57.68, 65.66, 56.95, 100.44, 47.63],
[ 96.23, 14.01, 0.88, 89.77, 66.77],
[ 70.35, 23.69, 73.41, 70.70, 85.41]])
BRADTKE_FEATURE_MATRIX = np.array([[74.29, 34.61, 73.48, 53.29, 7.79],
[61.60, 48.07, 34.68, 36.19, 82.02],
[97.00, 4.88, 8.51, 87.89, 5.17],
[41.10, 40.13, 64.63, 92.67, 31.09],
[ 7.76, 79.82, 43.78, 8.56, 61.11]])
class PlayerSpriteBradtkeChain(prefab_sprites.MazeWalker):
"""A `Sprite` for our player."""
def __init__(self, corner, position, character):
"""Inform superclass that we cant walk through #."""
super(PlayerSpriteBradtkeChain, self).__init__(
corner, position, character, impassable='#')
def update(self, actions, board, layers, backdrop, things, the_plot):
del backdrop, things # Unused.
#Get current position in the chain
info, position = self.position
# Apply motion commands.
if actions == 0:
# Choose next state for the current state randomly.
new_position = np.argmax(np.random.multinomial(n=1, pvals=TRANSITION_MATRIX[position, :]))
# Get the next reward from next and current state.
the_plot.add_reward(REWARD_MATRIX[position, new_position])
# Move to next state.
self._teleport((0, new_position))
class BradtkeChain(gym.Env):
def __init__(self):
self.game= ascii_art.ascii_art_to_game(
BRADTKE_STATES,
what_lies_beneath=' ',
sprites={'P': PlayerSpriteBradtkeChain})
obs, reward, gamma = self.game.its_showtime()
def step(self):
""" Take given action and return the next state observation and reward.
"""
obs, reward, gamma = self.game.play(0)
return obs, reward, self.game.game_over, ""
def reset(self):
""" Resets the game and returns the observation for the start state.
"""
self.game= ascii_art.ascii_art_to_game(
BRADTKE_STATES,
what_lies_beneath=' ',
sprites={'P': PlayerSpriteBradtkeChain})
obs, reward, gamma = self.game.its_showtime()
return obs
# + [markdown] id="_yveoKX7t_qF" colab_type="text"
# **Testing Five States Environment**
# + id="7skU28IEpJFG" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}, {"item_id": 2}, {"item_id": 3}, {"item_id": 4}, {"item_id": 5}, {"item_id": 6}, {"item_id": 7}, {"item_id": 8}, {"item_id": 9}, {"item_id": 10}, {"item_id": 11}, {"item_id": 12}], "base_uri": "https://localhost:8080/", "height": 833} outputId="bea5af48-5ce6-4bd3-8131-f5448148bc7c" executionInfo={"status": "ok", "timestamp": 1521671298414, "user_tz": 240, "elapsed": 1170, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "105824936464368788271"}}
game = BradtkeChain()
obs = game.reset()
stored_chain = displayenv(obs, display_visits=True)
returnGt = 0
game_over = False
max_steps = 10
step = 0
#Play until terminates
while not(game_over):
obs, reward, game_over, _ = game.step()
returnGt += reward
storedchain = displayenv(obs, display_visits=True, stored_chain=stored_chain)
step += 1
if step > max_steps:
break
# + [markdown] id="TaOfHrbqhBYM" colab_type="text"
# # LSTD(lamda)
# + [markdown] id="SNFgV011trxw" colab_type="text"
# ***Offline LSTD Implementation***
# + id="18y5xslKhG84" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
#--------------------------------------------------------------------
#
# Given: simulation model, featurizer, and \lambda for TD($\lambda$)
# Output: a coefficient vector Beta for which Vpi(x) = Beta . Phi(x)
#
#
#
#--------------------------------------------------------------------
def offlineLSTD(game, featurizer, lstd_gamma, lstd_lambda, episodes=1000, max_timesteps=1000):
A, b = 0, 0
betas = []
for n in range(episodes):
#Choose a start state xt E X
obs = game.reset()
state = obsv2state(obs)
game_over = False
#Set eligibility trace vector Zt = Featurizer(xt)
phi_t = featurizer[state]
traces = phi_t
timesteps = 0
#While xt != End, Repeat:
while not(game_over) and timesteps < max_timesteps:
#Simulate on step of the chain, producing a reward Rt and next state xt+1
obs, reward, game_over, _ = game.step()
next_state = obsv2state(obs)
phi_tp = featurizer[next_state]
A = A + np.outer(traces, phi_t - lstd_gamma * phi_tp)
b = b + traces * reward # b = b + eligibility * reward
traces = lstd_gamma * lstd_lambda * traces + phi_tp # Set $Z_t+1 = \lambda * Z_t + phi(x_t+1)
timesteps = timesteps + 1
phi_t = phi_tp
#Whenever updated coefficients are desired: Set Beta = Ainv.b
beta = np.dot(np.linalg.inv(A),b)
betas.append(beta)
return A, b, np.array(betas)
# + [markdown] id="rYRYWIc9tXL0" colab_type="text"
# ***Recursive LSTD Implementation***
# + id="hljvp7eltXkd" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
#--------------------------------------------------------------------
#
# Given: simulation model, featurizer, and \lambda for TD($\lambda$)
# Output: a coefficient vector Beta for which Vpi(x) = Beta . Phi(x)
#
#
#
#--------------------------------------------------------------------
def recursiveLSTD(game, featurizer, lstd_gamma, lstd_lambda, episodes=1000, max_timesteps=1000, init_eps=10e-10):
Ainv = np.eye(featurizer.shape[1]) * 1/init_eps
b = np.zeros(featurizer.shape[1])
betas = []
for n in range(episodes):
#Choose a start state xt E X
obs = game.reset()
state = obsv2state(obs)
#storedchain = displayenv(obs, displayvisits=True)
#Set eligibility trace vector Zt = Featurizer(xt)
phi_t = featurizer[state].reshape(-1,1)
traces = phi_t
timesteps = 0
#While xt != End, Repeat:
game_over = False
while not(game_over) and timesteps < max_timesteps:
#Simulate on step of the chain, producing a reward Rt and next state xt+1
obs, reward, game_over, _ = game.step()
next_state = obsv2state(obs)
phi_tp = featurizer[next_state].reshape(-1,1)
u = traces
v = (phi_t - lstd_gamma * phi_tp)
Ainv = Ainv - np.dot(Ainv, np.dot(np.dot(u, v.T), Ainv))/(1 + np.dot(v.T, np.dot(Ainv, u)))
b = b + traces.reshape(-1) * reward #b = b + eligibility * reward
traces = lstd_gamma * lstd_lambda * traces + phi_tp #Set $Z_t+1 = \lambda * Z_t + phi(x_t+1)
timesteps = timesteps + 1
phi_t = phi_tp
#Whenever updated coefficients are desired: Set Beta = Ainv.b
beta = np.dot(Ainv,b)
betas.append(beta)
return Ainv, b, np.array(betas)
# + [markdown] id="iJ-TRhwMVj7Z" colab_type="text"
# # Empirical Analysis
# + [markdown] id="C-68b5TRr8BA" colab_type="text"
# ## Boyan's Chain: Performance of Offline LSTD for Different Lambda Values
# + id="VShj_xVlSY5d" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}], "base_uri": "https://localhost:8080/", "height": 386} outputId="9edfb0d4-6d1f-49f3-d92e-d33c3d667959" executionInfo={"status": "ok", "timestamp": 1521679582364, "user_tz": 240, "elapsed": 91232, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "105824936464368788271"}}
#-------------------------------------------------------------------------------------------#
#---RMSE Comparison of different lambdas for Boyan Chain using Offline LSTD (OLSTD)---------#
#-------------------------------------------------------------------------------------------#
lambdas = [0.00, 0.25, 0.40, 0.50, 0.75, 1.00]
trials = 10
episodes = 1000
lstd_gamma = 1.00
size = (len(lambdas), episodes)
#In fact, this domain’s optimal V π function is exactly linear in these features: the optimal coefficients β∗λ are (−24, −16, −8, 0).
beta_Optimal = np.array([-24, -16, -8, 0])
game = BoyansChain()
featurizer = BOYANCHAIN_FEATURES
V_Optimal = np.dot(beta_Optimal, BOYANCHAIN_FEATURES.T)
errors = np.zeros(size)
legends = []
total_time = 0
for i,lambda_current in enumerate(lambdas):
for _ in range(trials):
_, _, betas = offlineLSTD(game, featurizer, lstd_gamma, lambda_current, episodes)
predicted_Vs = np.dot(betas, BOYANCHAIN_FEATURES.T)
errors[i] += np.linalg.norm(predicted_Vs - V_Optimal, axis=1)
errors[i] /= trials
legend = "Offline-LSTD = {:}".format(lambda_current)
legends.append(legend)
plt.semilogx(range(episodes), errors[i], linewidth=1)
plt.xlabel("Trajectory Number", fontsize=14)
plt.ylabel("RMS error of value function over all states", fontsize=14)
plt.title("Boyan's Chain: Performance of Offline LSTD for Different Lambda Values", fontsize=14)
plt.legend(legends, prop={'size': 16}, loc='upper right', fancybox=True, framealpha=0.5)
plt.show()
# + [markdown] id="cJjmLqDFsCgQ" colab_type="text"
# ## Boyan's Chain: Performance of Recursive LSTD for Different Lambda Values
# + id="zuLCoJAb4Z8w" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}], "base_uri": "https://localhost:8080/", "height": 386} cellView="code" outputId="6b41f117-a385-4b19-865f-de7879a928fd" executionInfo={"status": "ok", "timestamp": 1521679650044, "user_tz": 240, "elapsed": 67116, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "105824936464368788271"}}
#-------------------------------------------------------------------------------------------#
#---RMSE Comparison of different lambdas for Boyan Chain using Recursive LSTD (OLSTD)-------#
#-------------------------------------------------------------------------------------------#
lambdas = [0.00, 0.25, 0.40, 0.50, 0.75, 1.00]
trials = 10
episodes = 1000
lstd_gamma = 1.00
size = (len(lambdas), episodes)
#In fact, this domain’s optimal V π function is exactly linear in these features: the optimal coefficients β∗λ are (−24, −16, −8, 0).
beta_Optimal = [-24, -16, -8, 0]
game = BoyansChain()
featurizer = BOYANCHAIN_FEATURES
V_Optimal = np.dot(beta_Optimal, BOYANCHAIN_FEATURES.T)
errors = np.zeros(size)
legends = []
for i,lambda_current in enumerate(lambdas):
for _ in range(trials):
_, _, betas = recursiveLSTD(game, featurizer, lstd_gamma, lambda_current, episodes)
predicted_Vs = np.dot(betas, BOYANCHAIN_FEATURES.T)
errors[i] += np.linalg.norm(predicted_Vs - V_Optimal, axis=1)
errors[i] /= trials
legend = "Recursive-LSTD = {:}".format(lambda_current)
legends.append(legend)
plt.semilogx(range(episodes), errors[i], linewidth=1)
plt.xlabel("Trajectory Number", fontsize=14)
plt.ylabel("RMS error of value function over all states", fontsize=14)
plt.title("Boyan's Chain: Performance of Recursive LSTD for Different Lambda Values", fontsize=14)
plt.legend(legends, prop={'size': 16}, loc='upper right', fancybox=True, framealpha=0.5)
plt.show()
# + [markdown] id="xcBp5K0CscP4" colab_type="text"
# ## Bradtke Chain: Performance of Offline LSTD for Different Lambda Values
# + id="ImrLPIZDdY2g" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}], "base_uri": "https://localhost:8080/", "height": 386} outputId="d6bc9bfe-2d00-4472-d91b-a186ef90b130" executionInfo={"status": "ok", "timestamp": 1521677142088, "user_tz": 240, "elapsed": 592288, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "105824936464368788271"}}
#-------------------------------------------------------------------------------------------#
#---RMSE Comparison of different lambdas for Bradtke Chain using Offline LSTD (OLSTD)-------#
#-------------------------------------------------------------------------------------------#
lambdas = [0.00, 0.25, 0.40, 0.50, 0.75, 1.00]
trials = 10
episodes = 1000
lstd_gamma = 0.8
size = (len(lambdas),episodes)
max_timesteps = 100
#In fact, this domain’s optimal V π function is exactly linear in these features: the optimal coefficients β∗λ are (−24, −16, −8, 0).
game = BradtkeChain()
featurizer = BRADTKE_FEATURE_MATRIX
V_Optimal = np.dot(beta_optimal_bradtke, BRADTKE_FEATURE_MATRIX.T)
errors = np.zeros(size)
legends = []
for i,lambdacurrent in enumerate(lambdas):
for _ in range(10):
_, _, betas = offlineLSTD(game, featurizer, lstd_gamma, lambdacurrent, episodes, max_timesteps)
predicted_Vs = np.dot(betas, BRADTKE_FEATURE_MATRIX.T)
errors[i] += np.linalg.norm(predicted_Vs, axis=1)
errors[i] /= trials
legend = "Offline-LSTD = {:}".format(lambdacurrent)
legends.append(legend)
plt.semilogx(range(episodes), errors[i], linewidth=1)
plt.xlabel("Trajectory Number", fontsize=14)
plt.ylabel("L2-norm of V_\pi", fontsize=14)
plt.title("Bradtke Chain: Performance of Offline LSTD for Different Lambda Values", fontsize=14)
plt.legend(legends, prop={'size': 16}, loc='upper right', fancybox=True, framealpha=0.5)
plt.show()
# + [markdown] id="CiSd9zPwF-rY" colab_type="text"
# ## Bradtke Chain: Performance of Recursive LSTD for Different Lambda Values
# + id="1sXBEypwWNzw" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}], "base_uri": "https://localhost:8080/", "height": 386} outputId="c469352e-9351-4700-da06-66475a540341" executionInfo={"status": "ok", "timestamp": 1521678642450, "user_tz": 240, "elapsed": 499156, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "105824936464368788271"}}
#-------------------------------------------------------------------------------------------#
#---RMSE Comparison of different lambdas for Bradtke Chain using Recursive LSTD (RLSTD)-------#
#-------------------------------------------------------------------------------------------#
lambdas = [0.00, 0.25, 0.40, 0.50, 0.75, 1.00]
trials = 10
episodes = 1000
lstd_gamma = 0.8
size = (len(lambdas),episodes)
max_timesteps = 100
#In fact, this domain’s optimal V π function is exactly linear in these features: the optimal coefficients β∗λ are (−24, −16, −8, 0).
game = BradtkeChain()
featurizer = BRADTKE_FEATURE_MATRIX
errors = np.zeros(size)
legends = []
for i,lambda_current in enumerate(lambdas):
for _ in range(10):
_, _, betas = recursiveLSTD(game, featurizer, lstd_gamma, lambda_current, episodes, max_timesteps)
predicted_Vs = np.dot(betas, BRADTKE_FEATURE_MATRIX.T)
errors[i] += np.linalg.norm(predicted_Vs, axis=1)
errors[i] /= trials
legend = "Recursive-LSTD = {:}".format(lambda_current)
legends.append(legend)
plt.semilogx(range(episodes), errors[i], linewidth=1)
plt.xlabel("Trajectory Number", fontsize=14)
plt.ylabel("L2-norm of V_\pi", fontsize=14)
plt.title("Bradtke Chain: Performance of Recursive LSTD for Different Lambda Values", fontsize=14)
plt.legend(legends, prop={'size': 16}, loc='upper right', fancybox=True, framealpha=0.5)
plt.show()
# + [markdown] id="sabg6A9M6-_b" colab_type="text"
# ## Discussion
# + [markdown] id="DkQPj4Ok6RIY" colab_type="text"
# In this set of experiments we evaluated our implementation of Offline-LSTD and Recursive-LSTD on two set of environments: Boyan's chain from [1] and Bradtke's chain from [2]. We ran each of these for 1000 episodes. As Bradtke's chain doesn't have a terminal state, we ran Bradtke's chain for 100 steps. $\gamma$ for Boyan's chain was set to $1$ and for Bradtke's chain to $0.8$.
#
# As we knew the optimal value for Boyan's chain, we plotted the RMSE over episodes for these experiments. We see that for all values of $\lambda$ the RMSE quickly coverges to $0$. So, the choice of $\lambda$ doesn't have much impact of either Offline or Recursive LSTD algorithm.
#
# For Bradtke's chain, we plotted the L2-norm of $V_\pi$. We see the same behavior that $V_pi$ coverges to same value for all values of $\lambda$ almost at the same rate i.e. the value of $\lambda$ doesn't have much impact of convergence. We choose $\gamma=0.8$ randomly and didn't run our experiments for various values of $\gamma$
# + [markdown] id="W2qIqnNSem12" colab_type="text"
# ## Equivalence of LSTD(1) and linear regression
# + [markdown] id="S8x56ioE7PRq" colab_type="text"
# **Read the section “Equivalence of LSTD(1) and linear regression” in the appendix of Boyan’s paper. Verify this claim empirically by computing the matrices A and b in the “supervised learning” approach. Compare the matrices with those that you would have obtained by LSTD(lambda=1) in part 1 and 2. Check if the solution (value function) found by the “supervised learning” approach matches the solution found by LSTD.**
# + id="s5qu3N4wXxEg" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
def build_features_returns(game, featurizer, episodes=1000):
features = []
returns = []
for n in range(episodes):
feats = []
rewards = []
game_over = False
#Choose a start state xt E X
obs = game.reset()
state = obsv2state(obs)
#While xt != End, Repeat:
while not(game_over):
#Simulate on step of the chain, producing a reward Rt and next state xt+1
feats.append(featurizer[state])
obs, reward, game_over, _ = game.step()
next_state = obsv2state(obs)
rewards.append(reward)
state = next_state
features.extend(feats)
returns.extend([np.sum(rewards[t:]) for t in range(len(rewards))])
return np.array(features), np.array(returns)
# + id="eWgsfSdaaUq6" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 3}], "base_uri": "https://localhost:8080/", "height": 898} outputId="78fc7503-dad9-40d4-e478-9e25a8c2ad4c" executionInfo={"status": "ok", "timestamp": 1521677737586, "user_tz": 240, "elapsed": 3498, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "105824936464368788271"}}
game = BoyansChain()
lstd_lambda = 1
lstd_gamma = 1
featurizer = BOYANCHAIN_FEATURES
episodes = 1000
errors = np.zeros(episodes)
np.random.seed(2204)
Phi, Y = build_features_returns(game, featurizer, episodes)
# A = X^TX
A_lsq = np.dot(Phi.T, Phi)
# b = X^TY
b_lsq = np.dot(Phi.T, Y)
# W = A^{-1}b
Beta_lsq = np.dot(np.linalg.inv(A_lsq),b_lsq)
print('_' * 100)
print("Ainv, b, beta and V_pi for Linear Least Square Regression")
print('_' * 100)
print(np.linalg.inv(A_lsq))
print('-' * 50)
print(b_lsq)
print('-' * 50)
print(Beta_lsq)
print('-' * 50)
print(np.dot(Beta_lsq, BOYANCHAIN13STATES_FEATURES.T))
np.random.seed(2204)
A_olstd, b_olstd, Beta_olstd = offlineLSTD(game, featurizer, lstd_gamma, lstd_lambda, episodes)
print('\n')
print('_' * 100)
print("Ainv, b, beta and V_pi for Offline LSTD for \lambda=1")
print('_' * 100);
print(np.linalg.inv(A_olstd))
print('-' * 50)
print(b_olstd)
print('-' * 50)
print(Beta_olstd[-1])
print('-' * 50)
print(np.dot(Beta_olstd[-1], BOYANCHAIN13STATES_FEATURES.T))
np.random.seed(2204)
Ainv_rlstd, b_rlstd, Beta_rlstd = recursiveLSTD(game, featurizer, lstd_gamma, lstd_lambda, episodes)
print('\n')
print('_' * 100)
print("Ainv, b, beta and V_pi for Recursive LSTD for \lambda=1")
print('_' * 100);
print(Ainv_rlstd)
print('-' * 50)
print(b_rlstd)
print('-' * 50)
print(Beta_rlstd[-1])
print('-' * 50)
print(np.dot(Beta_rlstd[-1], BOYANCHAIN13STATES_FEATURES.T))
# + [markdown] id="xhLuW7bQ6Nx-" colab_type="text"
# ### Discussion
# + [markdown] id="FSfUyqjc6fc9" colab_type="text"
# **Equivalence of LSTD(1) and linear regression**
# + [markdown] id="UC9o-y696R9S" colab_type="text"
# In this set of experiments, we aim to prove that for $\lambda=1$, the LSTD($\lambda$) algorithm is equivalent to Linear Least Square Regression. We empirically show that for Boyan's chain, the $A$ ($Ainv$), $b$, $\beta$ and $V_\pi$ for Offline-LSTD($\lambda$), Recursive-LSTD($\lambda$) and Least Square Linear Regression converge to the same value.
#
# We achieved this by first building the dataset for linear least square regression as mentioned in Appendix of Boyan's paper. The features $\Phi$ (Phi) and returns $Y$ (cumulated rewards) were built by running to game for $episode$=1000 number of times. Then using these features and returns, we do Linear Least Square update to compute $\beta$. $A$ and $b$ were the byproducts of this computations.
#
# Similarly, for both Recursive as well as Offline LSTD($\lambda$), we ran the experiments $episodes$=1000. We also retured the $A$ ($Ainv$ in case of recursive) and $b$ from our lstd methods.
#
|
dynaq/FunctionApproximation/LSTDlambda.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:dev_pytorch17]
# language: python
# name: conda-env-dev_pytorch17-py
# ---
# + colab={} colab_type="code" id="qdB23YiUA02F"
# %matplotlib inline
# %reload_ext autoreload
# %autoreload 2
# %load_ext line_profiler
# +
import sys, os, warnings, time, logging, re
import torch
import pytorch_lightning as pl
from pytorch_lightning.core.decorators import auto_move_data
from pathlib import Path
import PIL
import numpy as np
import matplotlib.pyplot as plt
import torchvision
from tqdm.autonotebook import tqdm
from lightning_addons.progressbar import ProgressBar
from lightning_addons.progressplotter import ProgressPlotter
from lightning_addons.progressprinter import ProgressPrinter
warnings.filterwarnings("ignore", category=UserWarning)
warnings.filterwarnings("ignore", category=DeprecationWarning)
# this makes lightning reports not look like errors
pl._logger.handlers = [logging.StreamHandler(sys.stdout)]
# -
# # ImageDataset
# +
def get_files(path, regex_filter=None):
if regex_filter is not None:
return [
Path(file)
for file in os.listdir(path)
if re.search(regex_filter, str(file))
]
return os.listdir(path)
class ImageDataset(torch.utils.data.Dataset):
def __init__(self, root, img_transforms=None, seg_transforms=None):
self.root = root
if img_transforms:
self.img_transform = torchvision.transforms.Compose(img_transforms)
else:
self.img_transform = None
if seg_transforms:
self.seg_transform = torchvision.transforms.Compose(seg_transforms)
else:
self.seg_transform = None
seg_file_filter = ".*_seg\.tif"
img_file_filter = ".*\.jpg"
self.img_filenames = get_files(root, img_file_filter)
self.seg_filenames = get_files(root, seg_file_filter)
def __getitem__(self, index):
img = PIL.Image.open(self.root / self.img_filenames[index])
seg = PIL.Image.open(self.root / self.seg_filenames[index])
if self.img_transform:
img = self.img_transform(img)
if self.seg_transform:
seg = self.seg_transform(seg)
return {"image": img, "segmentation": seg}
def __len__(self):
return len(self.img_filenames)
dataset = ImageDataset("/fastdata/ISPRS_BENCHMARK_DATASETS/Potsdam/processed/annotated_images/patches_300x300/training")
len(dataset)
# -
# # PotsdamDataModule
# mean and std for "image"
image_mean, image_std = (torch.tensor([0.3421, 0.3642, 0.3392]), torch.tensor([0.1402, 0.1360, 0.1415]))
segmentation_mean, segmentation_std = (torch.tensor([0.3725, 0.6933, 0.7947]), torch.tensor([0.4819, 0.4601, 0.4021]))
# +
class PotsdamDataModule(pl.LightningDataModule):
def __init__(self, data_dir: Path = './',
batch_size: int = 64, num_workers: int = 8,
x_norm_mean: float = 0.5, x_norm_std: float = 5,
y_norm_mean: float = 0.5, y_norm_std: float = 5,
):
super().__init__()
self.data_dir = Path(data_dir)
self.batch_size = batch_size
self.num_workers = num_workers
# self.dims is returned when you call dm.size()
# Setting default dims here because we know them.
# Could optionally be assigned dynamically in dm.setup()
self.dims = (3, 256, 256)
self.img_transforms = [
torchvision.transforms.Resize(self.dims[1]),
torchvision.transforms.ToTensor(),
torchvision.transforms.Normalize(x_norm_mean, x_norm_std)
]
self.seg_transforms = [
torchvision.transforms.Resize(self.dims[1]),
torchvision.transforms.ToTensor(),
torchvision.transforms.Normalize(y_norm_mean, y_norm_std)
]
def setup(self, stage=None):
if stage == 'fit' or stage is None:
self.train_ds = ImageDataset(self.data_dir / "training", self.img_transforms, self.seg_transforms)
if stage == 'test' or stage is None:
self.test_ds = ImageDataset(self.data_dir / "test", self.img_transforms, self.seg_transforms)
def train_dataloader(self):
return torch.utils.data.DataLoader(self.train_ds, batch_size=self.batch_size, num_workers=self.num_workers, pin_memory=True, shuffle=True)
def test_dataloader(self):
return torch.utils.data.DataLoader(self.test_ds, batch_size=self.batch_size, num_workers=self.num_workers, pin_memory=True)
x_mean = image_mean
x_std = image_std
y_mean = segmentation_mean
y_std = segmentation_std
# y_mean = 0.5
# y_std = 0.5
batch_size = 64
data = PotsdamDataModule(
"/fastdata/ISPRS_BENCHMARK_DATASETS/Potsdam/processed/annotated_images/patches_300x300/",
batch_size,
x_norm_mean=x_mean,
x_norm_std=x_std,
y_norm_mean=y_mean,
y_norm_std=y_std,
)
data.prepare_data()
data.setup()
batch = next(iter(data.test_dataloader()))
# -
batch["image"].shape, batch["segmentation"].shape
for i in range(0, batch_size, 8):
f, axs = plt.subplots(1, 2)
axs[0].imshow(batch["image"][i].permute(1, 2, 0).mul(image_std).add(image_mean))
# axs[1].hist(batch["image"][i].reshape(-1).numpy())
axs[1].imshow(batch["segmentation"][i].permute(1, 2, 0).mul(segmentation_std).add(segmentation_mean))
# ## find dataset mean and std
# +
# # compute mean and std etc
# # input_key = "image"
# # target_key = "segmentation"
# input_key = "segmentation"
# target_key = "image"
# def collect_stats(sample):
# image = sample[input_key]
# return image.sum((1, 2)), (image ** 2).sum((1, 2))#, image.mean((1, 2)), image.min((1, 2)), image.max((1, 2))
# all_stats = [collect_stats(sample) for sample in tqdm(data.train_dataloader().dataset)]
# sums, squared_sums= [torch.stack(stats, 0) for stats in (zip(*all_stats))]
# # , means, mins, maxs
# ds_mean = sums.mean(0) / (256 * 256)
# # ds_mean = means.mean(0)
# ds_std = (squared_sums.mean(0) / (256 * 256) - ds_mean ** 2).sqrt()
# ds_mean, ds_std
# -
import torch.nn as nn
import functools
# +
class Up(nn.Module):
def __init__(self, in_channels, out_channels, bias=True, upsample="conv"):
super().__init__()
self.upsample = upsample
if upsample == "conv":
self.conv = nn.ConvTranspose2d(in_channels, out_channels, kernel_size=4, stride=2, padding=1, bias=bias)
elif upsample == "bilinear":
self.upsampler = nn.Upsample(scale_factor=2.0, mode='bilinear', align_corners=True)
self.conv = nn.ConvTranspose2d(in_channels, out_channels, kernel_size=3, stride=1, padding=1, bias=bias)
def forward(self, x):
if self.upsample == "conv":
return self.conv(x)
elif self.upsample == "bilinear":
x_u = self.upsampler(x)
return self.conv(x_u)
Up(128, 64, upsample="bilinear")(torch.empty(2, 128, 64, 64)).shape, Up(128, 64, upsample="conv")(torch.empty(2, 128, 64, 64)).shape
# +
class UnetGenerator(nn.Module):
"""Create a Unet-based generator"""
def __init__(self, input_nc, output_nc, num_downs, ngf=64, norm_layer=nn.BatchNorm2d, use_dropout=False, upsample="conv"):
"""Construct a Unet generator
Parameters:
input_nc (int) -- the number of channels in input images
output_nc (int) -- the number of channels in output images
num_downs (int) -- the number of downsamplings in UNet. For example, # if |num_downs| == 7,
image of size 128x128 will become of size 1x1 # at the bottleneck
ngf (int) -- the number of filters in the last conv layer
norm_layer -- normalization layer
We construct the U-Net from the innermost layer to the outermost layer.
It is a recursive process.
"""
super(UnetGenerator, self).__init__()
# construct unet structure
unet_block = UnetSkipConnectionBlock(ngf * 8, ngf * 8, input_nc=None, submodule=None, norm_layer=norm_layer, innermost=True, upsample=upsample) # add the innermost layer
for i in range(num_downs - 5): # add intermediate layers with ngf * 8 filters
unet_block = UnetSkipConnectionBlock(ngf * 8, ngf * 8, input_nc=None, submodule=unet_block, norm_layer=norm_layer, use_dropout=use_dropout, upsample=upsample)
# gradually reduce the number of filters from ngf * 8 to ngf
unet_block = UnetSkipConnectionBlock(ngf * 4, ngf * 8, input_nc=None, submodule=unet_block, norm_layer=norm_layer, upsample=upsample)
unet_block = UnetSkipConnectionBlock(ngf * 2, ngf * 4, input_nc=None, submodule=unet_block, norm_layer=norm_layer, upsample=upsample)
unet_block = UnetSkipConnectionBlock(ngf, ngf * 2, input_nc=None, submodule=unet_block, norm_layer=norm_layer, upsample=upsample)
self.model = UnetSkipConnectionBlock(output_nc, ngf, input_nc=input_nc, submodule=unet_block, outermost=True, norm_layer=norm_layer, upsample=upsample) # add the outermost layer
def forward(self, input):
"""Standard forward"""
return self.model(input)
class UnetSkipConnectionBlock(nn.Module):
"""Defines the Unet submodule with skip connection.
X -------------------identity----------------------
|-- downsampling -- |submodule| -- upsampling --|
"""
def __init__(self, outer_nc, inner_nc, input_nc=None,
submodule=None, outermost=False, innermost=False,
norm_layer=nn.BatchNorm2d, use_dropout=False, output_activation=nn.Tanh(), upsample="conv"):
"""Construct a Unet submodule with skip connections.
Parameters:
outer_nc (int) -- the number of filters in the outer conv layer
inner_nc (int) -- the number of filters in the inner conv layer
input_nc (int) -- the number of channels in input images/features
submodule (UnetSkipConnectionBlock) -- previously defined submodules
outermost (bool) -- if this module is the outermost module
innermost (bool) -- if this module is the innermost module
norm_layer -- normalization layer
use_dropout (bool) -- if use dropout layers.
"""
super(UnetSkipConnectionBlock, self).__init__()
self.outermost = outermost
if type(norm_layer) == functools.partial:
use_bias = norm_layer.func == nn.InstanceNorm2d
else:
use_bias = norm_layer == nn.InstanceNorm2d
if input_nc is None:
input_nc = outer_nc
downconv = nn.Conv2d(input_nc, inner_nc, kernel_size=4,
stride=2, padding=1, bias=use_bias)
downrelu = nn.LeakyReLU(0.2, True)
downnorm = norm_layer(inner_nc)
uprelu = nn.ReLU(True)
upnorm = norm_layer(outer_nc)
if outermost:
upconv = Up(inner_nc * 2, outer_nc, upsample=upsample)
down = [downconv]
up = [uprelu, upconv, output_activation]
model = down + [submodule] + up
elif innermost:
upconv = Up(inner_nc, outer_nc, bias=use_bias, upsample=upsample)
down = [downrelu, downconv]
up = [uprelu, upconv, upnorm]
model = down + up
else:
upconv = Up(inner_nc * 2, outer_nc, bias=use_bias, upsample=upsample)
down = [downrelu, downconv, downnorm]
up = [uprelu, upconv, upnorm]
if use_dropout:
model = down + [submodule] + up + [nn.Dropout(0.5)]
else:
model = down + [submodule] + up
self.model = nn.Sequential(*model)
def forward(self, x):
if self.outermost:
return self.model(x)
else: # add skip connections
return torch.cat([x, self.model(x)], 1)
input_channels = 3
output_channels = 3
num_downs = 7
generator = UnetGenerator(input_channels, output_channels, num_downs, upsample="bilinear")
generator(torch.empty(2, 3, 256, 256)).shape
# +
class NLayerDiscriminator(nn.Module):
"""Defines a PatchGAN discriminator"""
def __init__(self, input_nc, ndf=64, n_layers=3, norm_layer=nn.BatchNorm2d):
"""Construct a PatchGAN discriminator
Parameters:
input_nc (int) -- the number of channels in input images
ndf (int) -- the number of filters in the last conv layer
n_layers (int) -- the number of conv layers in the discriminator
norm_layer -- normalization layer
"""
super(NLayerDiscriminator, self).__init__()
if type(norm_layer) == functools.partial: # no need to use bias as BatchNorm2d has affine parameters
use_bias = norm_layer.func == nn.InstanceNorm2d
else:
use_bias = norm_layer == nn.InstanceNorm2d
kw = 4
padw = 1
sequence = [nn.Conv2d(input_nc, ndf, kernel_size=kw, stride=2, padding=padw), nn.LeakyReLU(0.2, True)]
nf_mult = 1
nf_mult_prev = 1
for n in range(1, n_layers): # gradually increase the number of filters
nf_mult_prev = nf_mult
nf_mult = min(2 ** n, 8)
sequence += [
nn.Conv2d(ndf * nf_mult_prev, ndf * nf_mult, kernel_size=kw, stride=2, padding=padw, bias=use_bias),
norm_layer(ndf * nf_mult),
nn.LeakyReLU(0.2, True)
]
nf_mult_prev = nf_mult
nf_mult = min(2 ** n_layers, 8)
sequence += [
nn.Conv2d(ndf * nf_mult_prev, ndf * nf_mult, kernel_size=kw, stride=1, padding=padw, bias=use_bias),
norm_layer(ndf * nf_mult),
nn.LeakyReLU(0.2, True)
]
sequence += [nn.Conv2d(ndf * nf_mult, 1, kernel_size=kw, stride=1, padding=padw)] # output 1 channel prediction map
self.model = nn.Sequential(*sequence)
def forward(self, input):
"""Standard forward."""
return self.model(input)
ndf = 64
discriminator = NLayerDiscriminator(input_channels, ndf, n_layers=3)
# -
t = torch.empty(batch_size, input_channels, 256, 256)
discriminator(t).shape
# +
class Pix2Pix(pl.LightningModule):
def __init__(
self,
input_channels,
height,
width,
output_channels,
lr,
batch_size,
beta1=0.5,
l1_lambda=100,
):
super().__init__()
self.save_hyperparameters()
data_shape = (input_channels, width, height)
num_downs = 8 # 8: 1x1, 7: 2x2
self.generator = UnetGenerator(input_channels, output_channels, num_downs, upsample="bilinear")
ndf = 64
self.discriminator = NLayerDiscriminator(input_channels + output_channels, ndf, n_layers=3)
self.test_x = None
self.test_y = None
self.example_input_array = torch.zeros(batch_size, input_channels, height, width)
@auto_move_data
def forward(self, x):
return self.generator(x)
def configure_optimizers(self):
optimizer_generator = torch.optim.Adam(self.generator.parameters(), lr=self.hparams.lr, betas=(self.hparams.beta1, 0.999))
optimizer_discriminator = torch.optim.Adam(self.discriminator.parameters(), lr=self.hparams.lr, betas=(self.hparams.beta1, 0.999))
optimizers = [optimizer_generator, optimizer_discriminator]
return optimizers
def gan_criterion(self, preds, targets):
return torch.nn.functional.mse_loss(preds, targets)
def l1_criterion(self, preds, targets):
return torch.nn.functional.l1_loss(preds, targets)
def training_step(self, batch, batch_nb, optimizer_idx):
real_x_imgs, real_y_imgs = batch["image"], batch["segmentation"]
fake_y_imgs = self.generator(real_x_imgs)
if optimizer_idx == 0:
fake_y_real_x = torch.cat((fake_y_imgs, real_x_imgs), 1)
fake_pred = self.discriminator(fake_y_real_x)
valid = torch.ones_like(fake_pred)
fake = torch.zeros_like(fake_pred)
gan_loss = self.gan_criterion(fake_pred, valid)
l1_loss = self.l1_criterion(fake_y_imgs, real_y_imgs)
loss = gan_loss + self.hparams.l1_lambda * l1_loss
self.log("g_loss", loss)
self.log("gan_loss", gan_loss)
self.log("l1_loss", l1_loss)
elif optimizer_idx == 1:
real_y_real_x = torch.cat((real_y_imgs, real_x_imgs), 1)
real_pred = self.discriminator(real_y_real_x)
valid = torch.ones_like(real_pred)
fake = torch.zeros_like(real_pred)
fake_y_real_x = torch.cat((fake_y_imgs.detach(), real_x_imgs), 1)
fake_pred = self.discriminator(fake_y_real_x)
real_loss = self.gan_criterion(real_pred, valid)
fake_loss = self.gan_criterion(fake_pred, fake)
loss = (real_loss + fake_loss) / 2
self.log("d_loss", loss)
self.log("real_loss", real_loss)
self.log("fake_loss", fake_loss)
self.log("loss", loss)
return loss
def on_epoch_end(self):
if self.test_x is not None and self.test_y is not None:
test_x = self.test_x.to(self.device).permute(0, 2, 3, 1).mul(x_std.to(self.device)).add(x_mean.to(self.device)).permute(0, 3, 1, 2)
test_y = self.test_y.to(self.device).permute(0, 2, 3, 1).mul(y_std.to(self.device)).add(y_mean.to(self.device)).permute(0, 3, 1, 2)
fake_y_imgs = self.generator(test_x).permute(0, 2, 3, 1).mul(y_std.to(self.device)).add(y_mean.to(self.device)).permute(0, 3, 1, 2)
output = torch.cat((test_x, test_y, fake_y_imgs))
grid = torchvision.utils.make_grid(output)#.add(1.0).div(2.0).clip(0.0, 1.0)
self.logger[0].experiment.add_image(
"generated_images", grid, self.current_epoch
)
def set_test_batch(self, test_batch, n_samples=8):
self.test_x = test_batch["image"][:n_samples]
self.test_y = test_batch["segmentation"][:n_samples]
lr = 1e-4
model = Pix2Pix(*data.size(), output_channels, lr, batch_size)
model.set_test_batch(next(iter(data.test_dataloader())))
# +
# pl.core.memory.ModelSummary(model, mode="full")
# +
tb_logger = pl.loggers.TensorBoardLogger("/home/mtadmin/projects/tensorboard_logs")
logger = [tb_logger]
# logger = []
plotter = ProgressPlotter()
callbacks = [
# ProgressBar(),
ProgressPrinter(),
plotter,
]
trainer = pl.Trainer(
max_epochs=50,
# fast_dev_run=True,
gpus=1,
logger=logger,
callbacks=callbacks,
precision=16,
limit_train_batches=0.1,
)
start_time = time.time()
trainer.fit(model, data)
print(f"time {time.time() - start_time:.2}")
# -
# %debug
|
tutorial-pix2pix.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # DAT210x - Programming with Python for DS
# ## Module5- Lab5
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib
matplotlib.style.use('ggplot') # Look Pretty
# -
# ### A Convenience Function
def plotDecisionBoundary(model, X, y):
fig = plt.figure()
ax = fig.add_subplot(111)
padding = 0.6
resolution = 0.0025
colors = ['royalblue','forestgreen','ghostwhite']
# Calculate the boundaris
x_min, x_max = X[:, 0].min(), X[:, 0].max()
y_min, y_max = X[:, 1].min(), X[:, 1].max()
x_range = x_max - x_min
y_range = y_max - y_min
x_min -= x_range * padding
y_min -= y_range * padding
x_max += x_range * padding
y_max += y_range * padding
# Create a 2D Grid Matrix. The values stored in the matrix
# are the predictions of the class at at said location
xx, yy = np.meshgrid(np.arange(x_min, x_max, resolution),
np.arange(y_min, y_max, resolution))
# What class does the classifier say?
Z = model.predict(np.c_[xx.ravel(), yy.ravel()])
Z = Z.reshape(xx.shape)
# Plot the contour map
cs = plt.contourf(xx, yy, Z, cmap=plt.cm.terrain)
# Plot the test original points as well...
for label in range(len(np.unique(y))):
indices = np.where(y == label)
plt.scatter(X[indices, 0], X[indices, 1], c=colors[label], label=str(label), alpha=0.8)
p = model.get_params()
plt.axis('tight')
plt.title('K = ' + str(p['n_neighbors']))
# ### The Assignment
# Load up the dataset into a variable called `X`. Check `.head` and `dtypes` to make sure you're loading your data properly--don't fail on the 1st step!
# +
# .. your code here ..
# -
# Copy the `wheat_type` series slice out of `X`, and into a series called `y`. Then drop the original `wheat_type` column from the `X`:
# +
# .. your code here ..
# -
# Do a quick, "ordinal" conversion of `y`. In actuality our classification isn't ordinal, but just as an experiment...
# +
# .. your code here ..
# -
# Do some basic nan munging. Fill each row's nans with the mean of the feature:
# +
# .. your code here ..
# -
# Split `X` into training and testing data sets using `train_test_split()`. Use `0.33` test size, and use `random_state=1`. This is important so that your answers are verifiable. In the real world, you wouldn't specify a random_state:
# +
# .. your code here ..
# -
# Create an instance of SKLearn's Normalizer class and then train it using its .fit() method against your _training_ data. The reason you only fit against your training data is because in a real-world situation, you'll only have your training data to train with! In this lab setting, you have both train+test data; but in the wild, you'll only have your training data, and then unlabeled data you want to apply your models to.
# +
# .. your code here ..
# -
# With your trained pre-processor, transform both your training AND testing data. Any testing data has to be transformed with your preprocessor that has ben fit against your training data, so that it exist in the same feature-space as the original data used to train your models.
# +
# .. your code here ..
# -
# Just like your preprocessing transformation, create a PCA transformation as well. Fit it against your training data, and then project your training and testing features into PCA space using the PCA model's `.transform()` method. This has to be done because the only way to visualize the decision boundary in 2D would be if your KNN algo ran in 2D as well:
# +
# .. your code here ..
# -
# Create and train a KNeighborsClassifier. Start with `K=9` neighbors. Be sure train your classifier against the pre-processed, PCA- transformed training data above! You do not, of course, need to transform your labels.
# +
# .. your code here ..
# -
# I hope your KNeighbors classifier model from earlier was named 'knn'
# If not, adjust the following line:
plotDecisionBoundary(knn, X_train, y_train)
# Display the accuracy score of your test data/labels, computed by your KNeighbors model. You do NOT have to run `.predict` before calling `.score`, since `.score` will take care of running your predictions for you automatically.
# +
# .. your code here ..
# -
# ### Bonus
# Instead of the ordinal conversion, try and get this assignment working with a proper Pandas get_dummies for feature encoding. You might have to update some of the `plotDecisionBoundary()` code.
plt.show()
|
Module5/.ipynb_checkpoints/Module5 - Lab5-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"is_executing": false}
# -*- coding: utf-8 -*-
"""
Created on Wed Oct 9 15:33:37 2019
@author: Birdman
"""
from sklearn import tree
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.externals import joblib
from sklearn.metrics import precision_score, recall_score
import csv
# + pycharm={"name": "#%%\n", "is_executing": false}
def get_features(temp,xss):
temp.append(xss.lower().count("script"))
temp.append(xss.lower().count("java"))
temp.append(xss.lower().count("iframe"))
temp.append(xss.lower().count("<"))
temp.append(xss.lower().count(">"))
temp.append(xss.lower().count("\""))
temp.append(xss.lower().count("\'"))
temp.append(xss.lower().count("%"))
temp.append(xss.lower().count("("))
temp.append(xss.lower().count(")"))
return temp
# + pycharm={"name": "#%%\n", "is_executing": false}
if __name__ == '__main__':
xss=[]
label = []
temp = []
data = []
LaBel = []
with open(r'xssed.csv','r',encoding="utf-8") as csvfile:
csv_reader = csv.reader(csvfile)
for row in csv_reader:
xss.append(row[0])
label.append(1)
csvfile.close()
with open(r'dmzo_nomal.csv','r',encoding="utf-8") as csvfile:
csv_reader = csv.reader(csvfile)
for row in csv_reader:
xss.append(row[0])
label.append(0)
csvfile.close()
for i in range(0,len(xss)):
temp = get_features(temp,xss[i])
data.append(temp)
LaBel.append(label[i])
temp = []
data=np.array(data)
LaBel=np.array(LaBel)
#print(data[:10])
#print(LaBel[:10])
X_train,X_test,y_train,y_test=train_test_split(data,LaBel,test_size=0.3,random_state=0)
sc = StandardScaler()
sc.fit(X_train)
#对数据集进行处理
xss_tree = tree.DecisionTreeClassifier(criterion='gini', splitter='best', max_depth=None, min_samples_split=2,
min_samples_leaf=1, min_weight_fraction_leaf=0.0, max_features=10, random_state=None, max_leaf_nodes=None,
min_impurity_decrease=0.0, min_impurity_split=None, presort=False)
print("***************start****************")
xss_tree_save=xss_tree.fit(X_train,y_train)
print("**************saveing***************")
# joblib.dump(xss_tree_save,'url_tree.model')
print("Training score:%f" % (xss_tree.score(X_train, y_train)))
print("Testing score:%f" % (xss_tree.score(X_test, y_test)))
y_pred = xss_tree.predict(X_test)
print("Training precision score:%f" % (precision_score(y_test, y_pred)))
print("Training recall score:%f" % (recall_score(y_test, y_pred)))
#test
xss="<script>alert('1')</script>"
test=[]
test = np.array(get_features(test,xss))
if xss_tree.predict([test]) == 1:
print("test result: BAD")
else:
print("test result: GOOD")
|
Homework/2019/Task3/8/code/trainXSS.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# # Lab 1: Single-qubit and multi-qubit states, quantum teleportation
# In this lab, you will learn how to write `Qiskit` code and investigate single-qubit and multi-qubit states using the `qpshere` visualization that you learned in lecture 1.
#
# If you have not used Jupyter notebooks before, take a look at the following video to quickly get started.
# - https://www.youtube.com/watch?v=jZ952vChhuI
#
# Remember, to run a cell in Jupyter notebooks, you press `Shift` + `Return/Enter` on your keyboard.
# ### Installing necessary packages
# Before we begin, you will need to install some prerequisites into your environment. Run the cell below to complete these installations. At the end, the cell outputs will be cleared.
# +
# !pip install -U -r grading_tools/requirements.txt
from IPython.display import clear_output
clear_output()
# -
# # Single-qubit states
# In lecture, you learned that single qubit states can be written down generally as
#
# $$\sqrt{1-p}\vert0\rangle + e^{i\phi}\sqrt{p}\vert1\rangle$$
#
# Here, $p$ is the probability that a measurement of the state in the computational basis $\{\vert0\rangle, \vert1\rangle\}$ will have the outcome $1$, and $\phi$ is the phase between the two computational basis states.
#
# Single-qubit gates can then be used to manipulate this quantum state by changing either $p$, $\phi$, or both.
#
# Let's begin by creating a single-qubit quantum circuit. We can do this in `Qiskit` using the following:
# +
from qiskit import QuantumCircuit
mycircuit = QuantumCircuit(1)
mycircuit.draw('mpl')
# -
# The above quantum circuit does not contain any gates. Therefore, if you start in any state, say $\vert0\rangle$, applying this circuit to your state doesn't change the state.
#
# To see this clearly, let's create the statevector $\vert0\rangle$. In `Qiskit`, you can do this using the following:
# +
from qiskit.quantum_info import Statevector
sv = Statevector.from_label('0')
# -
# You can see what's contained in the object `sv`:
sv
# The vector itself can be found by writing
sv.data
# As you can see, the above matches what you learned in lecture. Recall that $$\vert0\rangle = \begin{bmatrix}1\\0\end{bmatrix}$$
#
# We can now apply the quantum circuit `mycircuit` to this state by using the following:
new_sv = sv.evolve(mycircuit)
# Once again, you can look at the new statevector by writing
new_sv
# As you can see, the statevector hasn't changed. Recall the concept of state projection that you learned in lecture. You can compute the projection of `new_sv` onto `sv` by writing
# +
from qiskit.quantum_info import state_fidelity
state_fidelity(sv, new_sv)
# -
# As you can see, the projection of `new_sv` onto `sv` is 1, indicating that the two states are identical. You can visualize this state using the `qsphere` by writing
# +
from qiskit.visualization import plot_state_qsphere
plot_state_qsphere(sv.data)
# -
# As you learned in lecture 1, applying an $X$ gate flips the qubit from the state $\vert0\rangle$ to the state $\vert1\rangle$. To see this clearly, we will first create a single-qubit quantum circuit with the $X$ gate.
# +
mycircuit = QuantumCircuit(1)
mycircuit.x(0)
mycircuit.draw('mpl')
# -
# Now, we can apply this circuit onto our state by writing
sv = Statevector.from_label('0')
new_sv = sv.evolve(mycircuit)
new_sv
# As you can see, the statevector now corresponds to that of the state $\vert1\rangle$. Recall that
#
# $$\vert1\rangle = \begin{bmatrix}0\\1\end{bmatrix}$$
# Now, the projection of `new_sv` onto `sv` is
state_fidelity(new_sv, sv)
# This is not surprising. Recall from the lecture that the states $\vert0\rangle$ and $\vert1\rangle$ are orthogonal. Therefore, $\langle0\vert1\rangle = 0$. The state can be shown on the `qsphere` by writing
plot_state_qsphere(new_sv.data)
# Similarly, we can create the state $$\frac{1}{\sqrt{2}}\left(\vert0\rangle + \vert1\rangle\right)$$
# by applying a Hadamard gate as you learned in lecture. Here is how we can create the state and visualize it in `Qiskit`:
sv = Statevector.from_label('0')
mycircuit = QuantumCircuit(1)
mycircuit.h(0)
mycircuit.draw('mpl')
new_sv = sv.evolve(mycircuit)
print(new_sv)
plot_state_qsphere(new_sv.data)
# As you can see above, the state has equal components of $\vert0\rangle$ and $\vert1\rangle$. The size of the circle is proportional to the probability of measuring each basis state in the statevector. As a result, you can see that the size of the circles is half of the size of the circles in our previous visualizations.
# Recall from lecture that we can also create other superpositions with different phase. Let's create $$\frac{1}{\sqrt{2}}\left(\vert0\rangle - \vert1\rangle\right)$$ which can be done by applying the Hadamard gate on the state $\vert1\rangle$.
# +
sv = Statevector.from_label('1')
mycircuit = QuantumCircuit(1)
mycircuit.h(0)
new_sv = sv.evolve(mycircuit)
print(new_sv)
plot_state_qsphere(new_sv.data)
# -
# This time, the bottom circle, corresponding to the basis state $\vert1\rangle$ has a different color corresponding to the phase of $\phi = \pi$. This is because the coefficient of $\vert1\rangle$ in the state $$\frac{1}{\sqrt{2}}\left(\vert0\rangle - \vert1\rangle\right)$$ is $-1$, which is equal to $e^{i\pi}$.
#
# Other phases can also be created by applying different gates. The $T$ and $S$ gates apply phases of $+\pi/4$ and $+\pi/2$, respectively. The widget below helps you see different gates, and their actions on single-qubit quantum states.
from resources.qiskit_textbook.widgets import gate_demo
gate_demo(qsphere=True)
# A summary of the operations of the most common gates on single-qubit states is given by the handy image below, where the phases are shown in degrees.
#
# 
# # Multi-qubit states
# Similar to the discussion above, you can also explore multi-qubit gates in `Qiskit`. In lecture, you learned about Bell states, and how they can be generated using quantum gates. We will demonstrate below how to create the Bell state $$\frac{1}{\sqrt{2}}\left(\vert00\rangle + \vert11\rangle\right)$$ from the state $\vert00\rangle$. We'll start by visualizing the state $\vert00\rangle$ using the same procedure:
sv = Statevector.from_label('00')
plot_state_qsphere(sv.data)
# Next, we use the Hadamard gate described above, along with a controlled-X gate, to create the Bell state.
mycircuit = QuantumCircuit(2)
mycircuit.h(0)
mycircuit.cx(0,1)
mycircuit.draw('mpl')
# The result of this quantum circuit on the state $\vert00\rangle$ is found by writing
new_sv = sv.evolve(mycircuit)
print(new_sv)
plot_state_qsphere(new_sv.data)
# Note how this looks very similar to a single-qubit superposition with zero phase. Following entanglement, it is no longer possible to treat the two qubits individually, and they must be considered to be one system.
#
# To see this clearly, we can see what would happen if we measured the Bell state above 1000 times.
# +
counts = new_sv.sample_counts(shots=1000)
from qiskit.visualization import plot_histogram
plot_histogram(counts)
# -
# As you can see above, all measurements give either the result `00` or `11`. In other words, if the measurement outcome for one of the qubits is known, then the outcome for the other is fully determined.
# ### Ungraded exercise 1
#
# Can you create the state $$\frac{1}{\sqrt{2}}\left(\vert01\rangle + \vert10\rangle\right)$$ using a similar procedure?
# ### Ungraded exercise 2
#
# Can you create the state $$\frac{1}{\sqrt{2}}\left(\vert01\rangle - \vert10\rangle\right)$$ using a similar procedure?
# # Measurements
# In the above example, we simulated the action of a measurement by sampling counts from the statevector. A measurement can explicitly be inserted into a quantum circuit as well. Here is an example that creates the same Bell state and applies a measurement.
mycircuit = QuantumCircuit(2, 2)
mycircuit.h(0)
mycircuit.cx(0,1)
mycircuit.measure([0,1], [0,1])
mycircuit.draw('mpl')
# Two new features appeared in the circuit compared to our previous examples.
#
# - First, note that we used a second argument in the `QuantumCircuit(2,2)` command. The second argument says that we will be creating a quantum circuit that contains two qubits (the first argument), and two classical bits (the second argument).
# - Second, note that the `measure` command takes two arguments. The first argument is the set of qubits that will be measured. The second is the set of classical bits onto which the outcomes from the measurements of the qubits will be stored.
# Since the above quantum circuit contains non-unitaries (the measurement gates), we will use `Qiskit`'s built-in `Aer` simulators to run the circuit. To get the measurement counts, we can use the following code:
from qiskit import Aer, execute
simulator = Aer.get_backend('qasm_simulator')
result = execute(mycircuit, simulator, shots=10000).result()
counts = result.get_counts(mycircuit)
plot_histogram(counts)
# As you can see, the measurement outcomes are similar to when we sampled counts from the statevector itself.
# # Graded exercise 1: Quantum teleportation
# In this graded exercise, you will teleport the quantum state
# $$\sqrt{0.70}\vert0\rangle + \sqrt{0.30}\vert1\rangle$$ from Alice's qubit to Bob's qubit. Recall that the teleportation algorithm consists of four major components:
#
# 1. Initializing the state to be teleported. We will do this on Alice's qubit `q0`.
# 2. Creating entanglement between two qubits. We will use qubits `q1` and `q2` for this. Recall that Alice owns `q1`, and Bob owns `q2`.
# 3. Applying a Bell measurement on Alice's qubits `q0` and `q1`.
# 4. Applying classically controlled operations on Bob's qubit `q2` depending on the outcomes of the Bell measurement on Alice's qubits.
#
# This exercise guides you through each of these steps.
# ### Initializing the state to be teleported
# First, create a quantum circuit that creates the state $$\sqrt{0.70}\vert0\rangle + \sqrt{0.30}\vert1\rangle$$ You can do this by using `Qiskit`'s `initialize` function.
def initialize_qubit(given_circuit, qubit_index):
import numpy as np
#from qiskit import initialize
### WRITE YOUR CODE BETWEEN THESE LINES - START
given_circuit.initialize([np.sqrt(0.7),np.sqrt(0.3)],qubit_index)
### WRITE YOUR CODE BETWEEN THESE LINES - END
return given_circuit
# Next, we need to create entanglement between Alice's and Bob's qubits.
def entangle_qubits(given_circuit, qubit_Alice, qubit_Bob):
### WRITE YOUR CODE BETWEEN THESE LINES - START
given_circuit.h(qubit_Alice)
given_circuit.cx(qubit_Alice,qubit_Bob)
### WRITE YOUR CODE BETWEEN THESE LINES - END
return given_circuit
# Next, we need to do a Bell measurement of Alice's qubits.
def bell_meas_Alice_qubits(given_circuit, qubit1_Alice, qubit2_Alice, clbit1_Alice, clbit2_Alice):
### WRITE YOUR CODE BETWEEN THESE LINES - START
given_circuit.cx(qubit1_Alice, qubit2_Alice)
given_circuit.h(qubit1_Alice)
given_circuit.measure(qubit1_Alice,clbit1_Alice)
given_circuit.measure(qubit2_Alice,clbit2_Alice)
### WRITE YOUR CODE BETWEEN THESE LINES - END
return given_circuit
# Finally, we apply controlled operations on Bob's qubit. Recall that the controlled operations are applied in this order:
#
# - an $X$ gate is applied on Bob's qubit if the measurement coutcome of Alice's second qubit, `clbit2_Alice`, is `1`.
# - a $Z$ gate is applied on Bob's qubit if the measurement coutcome of Alice's first qubit, `clbit1_Alice`, is `1`.
def controlled_ops_Bob_qubit(given_circuit, qubit_Bob, clbit1_Alice, clbit2_Alice):
print(clbit1_Alice,clbit2_Alice)
### WRITE YOUR CODE BETWEEN THESE LINES - START
given_circuit.x(qubit_Bob).c_if(clbit2_Alice, 1) # Apply gates if the registers
given_circuit.z(qubit_Bob).c_if(clbit1_Alice, 1)
### WRITE YOUR CODE BETWEEN THESE LINES - END
return given_circuit
# The next lines of code put everything together. **You do not need to modify anything below, but you will need to run the cell to submit your solution.**
# +
### imports
from qiskit import QuantumRegister, ClassicalRegister
### set up the qubits and classical bits
all_qubits_Alice = QuantumRegister(2)
all_qubits_Bob = QuantumRegister(1)
creg1_Alice = ClassicalRegister(1)
creg2_Alice = ClassicalRegister(1)
### quantum teleportation circuit here
# Initialize
mycircuit = QuantumCircuit(all_qubits_Alice, all_qubits_Bob, creg1_Alice, creg2_Alice)
initialize_qubit(mycircuit, 0)
mycircuit.barrier()
# Entangle
entangle_qubits(mycircuit, 1, 2)
mycircuit.barrier()
# Do a Bell measurement
bell_meas_Alice_qubits(mycircuit, all_qubits_Alice[0], all_qubits_Alice[1], creg1_Alice, creg2_Alice)
mycircuit.barrier()
# Apply classically controlled quantum gates
controlled_ops_Bob_qubit(mycircuit, all_qubits_Bob[0], creg1_Alice, creg2_Alice)
### Look at the complete circuit
print(mycircuit.draw(output='text'))
### store the circuit as the submitted answer
answer = mycircuit
# -
# Then, grade your solution by running the cell below. Provide always the same name and email, as the one you wrote during the course sign up.
# +
name = '<NAME>'
email = '<EMAIL>'
from grading_tools import grade
grade(answer, name, email, 'lab1', 'ex1')
# -
# # Additional reading
#
# - You can watch a video on building the quantum teleportation quantum circuit here: https://www.youtube.com/watch?v=mMwovHK2NrE&list=PLOFEBzvs-Vvp2xg9-POLJhQwtVktlYGbY&index=6&t=0s
#
# - For additional details about the quantum teleportation algorithm, including the principle of deferred measurement, you can refer to the Qiskit Textbook's section on the algorithm here: https://qiskit.org/textbook/ch-algorithms/teleportation.html
#
# - The `1 minute Qiskit` episode entitled `What is the qsphere?` succinctly describes the Qsphere visualization tool that we used in this lab. You can find it here: https://youtu.be/4SoK2h4a7us
|
Labs/lab1/ex1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <center> In-class Assignment 1 </center>
# **Task 1. Word game**
#
# You need to write a program for a word game.
# The first player think about a word and save it using input() function and only lowercase letters.
#
# The second player provides new words in the same format. In total, the user needs to make 15 moves (name 15 new words after the first word). Each new word must start with the same letter as the first word. If the word has already been called earlier (is duplicated) or the first letter of a new word is not the same as the first letter of the first word, the game ends with the phrase "The rules of the game are violated."
# +
# The first player comes up with a word
word = input('Name a word:')
# Second player names other new words
new_word = input('Next word starting from the same letter: ')
# YOUR CODE HERE
# -
# **Task 2. String preprocessing**.
#
# 1. This is a downloaded paragraph from the main page of the United Nations website. Remove all symbols for new lines ("\n"). Transform a list of all sentences into one text (= one string).
#
# 2. Create a dictionary, where the keys are the words and the values are the numbers how many times each word occured in a given paragraph). What is the most frequent word in this paragraph?
#
# For instance:
# ```python
# >>> your_dict['freedom']
# Out: 2
# >>>your_dict['human']
# Out: 4
# ```
paragraph = ["What Are Human Rights? Human rights are rights inherent to all human \n",
"beings, regardless of race, sex, nationality, ethnicity, language, religion, or \n",
"any other status. Human rights include the right to life and liberty, freedom \n",
"from slavery and torture, freedom of opinion and expression, the right to work \n",
"and education, and many more.", " Everyone is entitled to these rights, without ",
"discrimination. \nInternational human rights law lays down the obligations of Governments to act in \n",
"certain ways or to refrain from certain acts, in order to promote and protect human \n",
"rights and fundamental freedoms of individuals or groups. \n",
"One of the great achievements of the United Nations is the creation of a comprehensive \n",
"body of human rights law — a universal and internationally protected code to which all \n",
"nations can subscribe and all people aspire. The United Nations has defined a broad range \n",
"of internationally accepted rights, including civil, cultural, economic, political and \n",
"social rights. It has also established mechanisms to promote and protect these rights \n",
"and to assist states in carrying out their responsibilities."]
# +
# YOUR CODE HERE
# -
# **Task 3. G7 Summit**
#
# You are on the organizing committee of the G7 summit. You need to check in all the participants of the summit in a hotel with double rooms, guided by the rule: ```participants can live in one room if they are delegates from the same country.``` <br>
#
# You have been given a dictionary with information on all participants.
#
# Write a function that returns ```True```, if it is possible to check in $X$ participants ($X$ is the number of participants) in $\frac{X}{2}$ rooms without violating the rule and ```False``` otherwise.
#
# ```Input:```
participants = {
'<NAME>' : {'sex': 'Male', 'country': 'The United Kingdom'},
'<NAME>' : {'sex': 'Male', 'country': 'The United Kingdom'},
'<NAME>' : {'sex': 'Female', 'country': 'Germany'},
'<NAME>' : {'sex': 'Female', 'country': 'Germany'},
'<NAME>' : {'sex': 'Male', 'country': 'Italy'},
'<NAME>' : {'sex': 'Female', 'country': 'Italy'},
'<NAME>' : {'sex': 'Male', 'country': 'Canada'},
'<NAME>' : {'sex': 'Female', 'country': 'Canada'},
'<NAME>' : {'sex': 'Male', 'country': 'Canada'},
'<NAME>' : {'sex': 'Female', 'country': 'Canada'},
'<NAME>' : {'sex': 'Male', 'country': 'France'},
'<NAME>' : {'sex': 'Male', 'country': 'France'},
'<NAME>' : {'sex': 'Male', 'country': 'Japan'},
'Ren Suzuki' : {'sex': 'Male', 'country': 'Japan'},
'<NAME>' : {'sex': 'Male', 'country': 'USA'},
'<NAME>' : {'sex': 'Male', 'country': 'USA'}
}
# ```Output```: ```True``` (because there is even number of participants from each country).
# ```Hint:``` Note that the values in a dictionary are also the dictionaries. If you want to access to the value of a particular delegate:
#
# ```python
# >>> participants['Ren Suzuki']['country']
# Out: Japan
# ```
def check_in(participants):
# YOUR CODE HERE
|
week6/In_class_A1_196.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <small><small><i>
# All the IPython Notebooks in **Python Functions** lecture series by Dr. <NAME> are available @ **[GitHub](https://github.com/milaan9/04_Python_Functions)**
# </i></small></small>
# # Python Functions
#
# In this class, you'll learn about functions, what a function is, the syntax, components, and types of functions. Also, you'll learn to create a function in Python.
# # What is a function in Python?
#
# In Python, a **function is a block of organized, reusable (DRY- Don’t Repeat Yourself) code with a name** that is used to perform a single, specific task. It can take arguments and returns the value.
#
# Functions help break our program into smaller and modular chunks. As our program grows larger and larger, functions make it more organized and manageable.
#
# Furthermore, it improves efficiency and reduces errors because of the reusability of a code.
# ## Types of Functions
#
# Python support two types of functions
#
# 1. **[Built-in](https://github.com/milaan9/04_Python_Functions/tree/main/002_Python_Functions_Built_in)** function
# 2. **[User-defined](https://github.com/milaan9/04_Python_Functions/blob/main/Python_User_defined_Functions.ipynb)** function
#
# 1.**Built-in function**
#
# The functions which are come along with Python itself are called a built-in function or predefined function. Some of them are:
# **`range()`**, **`print()`**, **`input()`**, **`type()`**, **`id()`**, **`eval()`** etc.
#
# **Example:** Python **`range()`** function generates the immutable sequence of numbers starting from the given start integer to the stop integer.
#
# ```python
# >>> for i in range(1, 10):
# >>> print(i, end=' ')
#
# 1 2 3 4 5 6 7 8 9
# ```
#
# 2. **User-defined function**
#
# Functions which are created by programmer explicitly according to the requirement are called a user-defined function.
# **Syntax:**
#
# ```python
# def function_name(parameter1, parameter2):
# """docstring"""
# # function body
# # write some action
# return value
# ```
#
# <div>
# <img src="img/f1.png" width="550"/>
# </div>
# ## Defining a Function
#
# 1. **`def`** is a keyword that marks the start of the function header.
#
# 2. **`function_name`** to uniquely identify the function. Function naming follows the same **[rules of writing identifiers in Python](https://github.com/milaan9/01_Python_Introduction/blob/main/005_Python_Keywords_and_Identifiers.ipynb)**.
#
# 2. **`parameter`** is the value passed to the function. They are optional.
#
# 3. **`:`** (colon) to mark the end of the function header.
#
# 4. **`function body`** is a block of code that performs some task and all the statements in **`function body`** must have the same **indentation** level (usually 4 spaces).
#
# 5. **"""docstring"""** documentation string is used to describe what the function does.
#
# 6. **`return`** is a keyword to return a value from the function.. A return statement with no arguments is the same as return **`None`**.
#
# >**Note:** While defining a function, we use two keywords, **`def`** (mandatory) and **`return`** (optional).
#
# **Example:**
#
# ```python
# >>> def add(num1,num2): # Function name: 'add', Parameters: 'num1', 'num2'
# >>> print("Number 1: ", num1) # Function body
# >>> print("Number 2: ", num2) # Function body
# >>> addition = num1 + num2 # Function body
# >>> return addition # return value
#
#
# >>> res = add(2, 4) # Function call
# >>> print("Result: ", res)
# ```
# ## Defining a function without any parameters
#
# Function can be declared without parameters.
# +
# Example 1:
def greet():
print("Welcome to Python for Data Science")
# call function using its name
greet()
# +
# Example 2:
def add_two_numbers ():
num_one = 3
num_two = 6
total = num_one + num_two
print(total)
add_two_numbers() # calling a function
# +
# Example 3:
def generate_full_name ():
first_name = 'Milaan'
last_name = 'Parmar'
space = ' '
full_name = first_name + space + last_name
print(full_name)
generate_full_name () # calling a function
# -
# ## Defining a function without parameters and `return` value
#
# Function can also return values, if a function does not have a **`return`** statement, the value of the function is None. Let us rewrite the above functions using **`return`**. From now on, we get a value from a function when we call the function and print it.
# +
# Example 1:
def add_two_numbers ():
num_one = 3
num_two = 6
total = num_one + num_two
return total
print(add_two_numbers())
# +
# Example 2:
def generate_full_name ():
first_name = 'Milaan'
last_name = 'Parmar'
space = ' '
full_name = first_name + space + last_name
return full_name
print(generate_full_name())
# -
# ## Defining a function with parameters
#
# In a function we can pass different data types(number, string, boolean, list, tuple, dictionary or set) as a parameter.
# ### Single Parameter:
#
# If our function takes a parameter we should call our function with an argument
# +
# Example 1: Gereeting
def greet(name):
"""
This function greets to the person passed in as a parameter
"""
print("Hello, " + name + ". Good morning!") # No output!
# +
# Example 2:
def sum_of_numbers(n):
total = 0
for i in range(n+1):
total+=i
print(total)
print(sum_of_numbers(10)) # 55
print(sum_of_numbers(100)) # 5050
# -
# ### Two Parameter:
#
# A function may or may not have a parameter or parameters. A function may also have two or more parameters. If our function takes parameters we should call it with arguments.
# +
# Example 1:
def course(name, course_name):
print("Hello", name, "Welcome to Python for Data Science")
print("Your course name is", course_name)
course('Arthur', 'Python') # call function
# -
# ## Defining a function with parameters and `return` value
# +
# Example 1:
def greetings (name): # single parameter
message = name + ', welcome to Python for Data Science'
return message
print(greetings('Milaan'))
# +
# Example 2:
def add_ten(num): # single parameter
ten = 10
return num + ten
print(add_ten(90))
# +
# Example 3:
def square_number(x): # single parameter
return x * x
print(square_number(3))
# +
# Example 4:
def area_of_circle (r): # single parameter
PI = 3.14
area = PI * r ** 2
return area
print(area_of_circle(10))
# +
# Example 5:
def calculator(a, b): # two parameter
add = a + b
return add # return the addition
result = calculator(30, 6) # call function & take return value in variable
print("Addition :", result) # Output Addition : 36
# +
# Example 6:
def generate_full_name (first_name, last_name): # two parameter
space = ' '
full_name = first_name + space + last_name
return full_name
print('Full Name: ', generate_full_name('Milaan','Parmar'))
# +
# Example 7:
def sum_two_numbers (num_one, num_two): # two parameter
sum = num_one + num_two
return sum
print('Sum of two numbers: ', sum_two_numbers(1, 9))
# +
# Example 8:
def calculate_age (current_year, birth_year): # two parameter
age = current_year - birth_year
return age;
print('Age: ', calculate_age(2021, 1819))
# +
# Example 9:
def weight_of_object (mass, gravity): # two parameter
weight = str(mass * gravity)+ ' N' # the value has to be changed to a string first
return weight
print('Weight of an object in Newtons: ', weight_of_object(100, 9.81))
# -
# ## Function `return` Statement
#
# In Python, to return value from the function, a **`return`** statement is used. It returns the value of the expression following the returns keyword.
#
# **Syntax:**
#
# ```python
# def fun():
# statement-1
# statement-2
# statement-3
# .
# .
# return [expression]
# ```
#
# The **`return`** value is nothing but a outcome of function.
#
# * The **`return`** statement ends the function execution.
# * For a function, it is not mandatory to return a value.
# * If a **`return`** statement is used without any expression, then the **`None`** is returned.
# * The **`return`** statement should be inside of the function block.
# ### Return Single Value
print(greet("Cory"))
# Here, **`None`** is the returned value since **`greet()`** directly prints the name and no **`return`** statement is used.
# ## Passing Arguments with Key and Value
#
# If we pass the arguments with key and value, the order of the arguments does not matter.
# +
# Example 1:
def print_fullname(firstname, lastname):
space = ' '
full_name = firstname + space + lastname
print(full_name)
print(print_fullname(firstname = 'Milaan', lastname = 'Parmar'))
# +
# Example 2:
def add_two_numbers (num1, num2):
total = num1 + num2
print(total)
print(add_two_numbers(num2 = 3, num1 = 2)) # Order does not matter
# -
# If we do not **`return`** a value with a function, then our function is returning **`None`** by default. To return a value with a function we use the keyword **`return`** followed by the variable we are returning. We can return any kind of data types from a function.
# +
# Example 1: with return statement
def print_fullname(firstname, lastname):
space = ' '
full_name = firstname + space + lastname
return full_name
print(print_fullname(firstname = 'Milaan', lastname = 'Parmar'))
# +
# Example 2: with return statement
def add_two_numbers (num1, num2):
total = num1 + num2
return total
print(add_two_numbers(num2 = 3, num1 = 2)) # Order does not matter
# +
# Example 3:
def absolute_value(num):
"""This function returns the absolute
value of the entered number"""
if num >= 0:
return num
else:
return -num
print(absolute_value(2))
print(absolute_value(-4))
# +
# Example 4:
def sum(a,b): # Function 1
print("Adding the two values")
print("Printing within Function")
print(a+b)
return a+b
def msg(): # Function 2
print("Hello")
return
total=sum(10,20)
print('total : ',total)
msg()
print("Rest of code")
# +
# Example 5:
def is_even(list1):
even_num = []
for n in list1:
if n % 2 == 0:
even_num.append(n)
# return a list
return even_num
# Pass list to the function
even_num = is_even([2, 3, 46, 63, 72, 83, 90, 19])
print("Even numbers are:", even_num)
# -
# ### Return Multiple Values
#
# You can also return multiple values from a function. Use the return statement by separating each expression by a comma.
# +
# Example 1:
def arithmetic(num1, num2):
add = num1 + num2
sub = num1 - num2
multiply = num1 * num2
division = num1 / num2
# return four values
return add, sub, multiply, division
a, b, c, d = arithmetic(10, 2) # read four return values in four variables
print("Addition: ", a)
print("Subtraction: ", b)
print("Multiplication: ", c)
print("Division: ", d)
# -
# ### Return Boolean Values
# +
# Example 1:
def is_even (n):
if n % 2 == 0:
print('even')
return True # return stops further execution of the function, similar to break
return False
print(is_even(10)) # True
print(is_even(7)) # False
# -
# ### Return a List
# +
# Example 1:
def find_even_numbers(n):
evens = []
for i in range(n + 1):
if i % 2 == 0:
evens.append(i)
return evens
print(find_even_numbers(10))
# -
# ## How to call a function in python?
#
# Once we have defined a function, we can call it from another function, program or even the Python prompt. To call a function we simply type the function name with appropriate parameters.
#
# <div>
# <img src="img/f2.png" width="500"/>
# </div>
greet('Alan')
# >**Note:** Try running the above code in the Python program with the function definition to see the output.
# +
# Example 1:
def wish(name):
"""
This function wishes to the person passed in as a parameter
"""
print("Happy birthday, " + name + ". Hope you have a wonderful day!")
wish('Bill')
# +
# Example 2:
def greetings (name = 'Clark'):
message = name + ', welcome to Python for Data Science'
return message
print(greetings())
print(greetings('Milaan'))
# +
# Example 3:
def generate_full_name (first_name = 'Milaan', last_name = 'Parmar'):
space = ' '
full_name = first_name + space + last_name
return full_name
print(generate_full_name())
print(generate_full_name('Ethan','Hunt'))
# +
# Example 4:
def calculate_age (birth_year,current_year = 2021):
age = current_year - birth_year
return age;
print('Age: ', calculate_age(1821))
# +
# Example 5:
def swap(x, y):
"""
This function swaps the value of two variables
"""
temp = x; # value of x will go inside temp
x = y; # value of y will go inside x
y = temp; # value of temp will go inside y
print("value of x is:", x)
print("value of y is:", y)
return # "return" is optional
x = 6
y = 9
swap(x, y) #call function
# +
# Example 6:
def even_odd(n):
if n % 2 == 0: # check number is even or odd
print(n, 'is a Even number')
else:
print(n, 'is a Odd Number')
even_odd(9) # calling function by its name
# +
# Example 7:
def weight_of_object (mass, gravity = 9.81):
weight = str(mass * gravity)+ ' N' # the value has to be changed to string first
return weight
print('Weight of an object in Newtons: ', weight_of_object(100)) # 9.81 - average gravity on Earth's surface
print('Weight of an object in Newtons: ', weight_of_object(100, 1.62)) # gravity on the surface of the Moon
# -
#
# ## Docstrings
#
# The first string after the function header is called the **docstring** and is short for documentation string. It is a descriptive text (like a comment) written by a programmer to let others know what block of code does.
#
# Although **optional**, documentation is a good programming practice. Unless you can remember what you had for dinner last week, always document your code.
#
# It is being declared using triple single quotes **`''' '''`** or triple-double quote **`""" """`** so that docstring can extend up to multiple lines.
#
# We can access docstring using doc attribute **`__doc__`** for any object like list, tuple, dict, and user-defined function, etc.
#
# In the above example, we have a docstring immediately below the function header.
print(greet.__doc__)
# To learn more about docstrings in Python, visit **[Python Docstrings](https://github.com/milaan9/04_Python_Functions/blob/main/Python_Docstrings.ipynb)**.
# ## Function `pass` Statement
#
# In Python, the **`pass`** is the keyword, which won’t do anything. Sometimes there is a situation where we need to define a syntactically empty block. We can define that block using the **`pass`** keyword.
#
# When the interpreter finds a **`pass`** statement in the program, it returns no operation.
# +
# Example 1:
def addition(num1, num2):
# Implementation of addition function in comming release
# Pass statement
pass
addition(10, 2)
# -
# ## 💻 Exercises ➞ <span class='label label-default'>Functions</span>
#
# ### Exercises ➞ <span class='label label-default'>Level 1</span>
#
# 1. Area of a circle is calculated as follows: **area = π x r x r** and **perimeter = 2 x π x r**. Write a function that calculates **`area_of_circle`** and **`perimeter_of_circle`**.
# 2. Write a function called **`add_all_nums`** which takes arbitrary number of arguments and sums all the arguments. Check if all the list items are number types. If not do give a reasonable feedback.
# 3. Temperature in **°C** can be converted to **°F** using this formula: **°F = (°C x 9/5) + 32**. Write a function which converts **°C to °F**, **`convert_celsius_2_fahrenheit`**.
# 4. Write a function called **`check_season`**, it takes a month parameter and returns the season: Autumn, Winter, Spring or Summer.
# 5. Write a function called **`calculate_slope`** which return the slope of a linear equation
# 6. Quadratic equation is calculated as follows: **ax² + bx + c = 0**. Write a function which calculates solution set of a quadratic equation, **`solve_quadratic_eqn`**.
# 7. Declare a function named **`print_list`**. It takes a list as a parameter and it prints out each element of the list.
# 8. Declare a function named **`reverse_list`**. It takes an array as a parameter and it returns the reverse of the array (use loops).
#
# - ```py
# print(reverse_list([1, 2, 3, 4, 5]))
# #[5, 4, 3, 2, 1]
# print(reverse_list1(["A", "B", "C"]))
# #["C", "B", "A"]
# ```
#
# 9. Declare a function named **`capitalize_list_items`**. It takes a list as a parameter and it returns a capitalized list of items
# 10. Declare a function named **`add_item`**. It takes a list and an item parameters. It returns a list with the item added at the end.
#
# - ```py
# food_staff = ['Potato', 'Tomato', 'Mango', 'Milk']
# print(add_item(food_staff, 'Fungi')) #['Potato', 'Tomato', 'Mango', 'Milk', 'Fungi']
# numbers = [2, 3, 7, 9]
# print(add_item(numbers, 5)) #[2, 3, 7, 9, 5]
# ```
#
# 11. Declare a function named **`remove_item`**. It takes a list and an item parameters. It returns a list with the item removed from it.
#
# - ```py
# food_staff = ['Potato', 'Tomato', 'Mango', 'Milk']
# print(remove_item(food_staff, 'Mango')) # ['Potato', 'Tomato', 'Milk']
# numbers = [2, 3, 7, 9]
# print(remove_item(numbers, 3)) # [2, 7, 9]
# ```
#
# 12. Declare a function named **`sum_of_numbers`**. It takes a number parameter and it adds all the numbers in that range.
#
# - ```py
# print(sum_of_numbers(5)) # 15
# print(sum_all_numbers(10)) # 55
# print(sum_all_numbers(100)) # 5050
# ```
#
# 13. Declare a function named **`sum_of_odds`**. It takes a number parameter and it adds all the odd numbers in that range.
# 14. Declare a function named **`sum_of_even`**. It takes a number parameter and it adds all the even numbers in that - range.
#
# ### Exercises ➞ <span class='label label-default'>Level 2</span>
#
# 1. Declare a function named **`evens_and_odds`**. It takes a positive integer as parameter and it counts number of evens and odds in the number.
#
# - ```py
# print(evens_and_odds(100))
# #The number of odds are 50.
# #The number of evens are 51.
# ```
#
# 2. Call your function **`factorial`**, it takes a whole number as a parameter and it return a factorial of the number
# 3. Call your function **`is_empty`**, it takes a parameter and it checks if it is empty or not
# 4. Write different functions which take lists. They should **`calculate_mean`**, **`calculate_median`**, **`calculate_mode`**, **`calculate_range`**, **`calculate_variance`**, **`calculate_std`** (standard deviation).
#
# ### Exercises ➞ <span class='label label-default'>Level 3</span>
#
# 1. Write a function called **`is_prime`**, which checks if a number is prime.
# 2. Write a functions which checks if all items are unique in the list.
# 3. Write a function which checks if all the items of the list are of the same data type.
# 4. Write a function which check if provided variable is a valid python variable
# 5. Go to the data folder and access the **[countries-data.py](https://github.com/milaan9/03_Python_Flow_Control/blob/main/countries_details_data.py)** file.
#
# - Create a function called the **`most_spoken_languages`** in the world. It should return 10 or 20 most spoken languages in the world in descending order
# - Create a function called the **`most_populated_countries`**. It should return 10 or 20 most populated countries in descending order.
|
001_Python_Functions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="pUEqzkNmnyYH"
# # [HW2] Topic Modeling
# 1. Crawling News
# 2. Preprocessing
# 3. Build Term-Document Matrix
# 4. Topic modeling
# 5. Visualization
#
# ```
# 🔥 이번 시간에는 Topic Modeling를 직접 크롤링한 뉴스 데이터에 대해서 수행해보는 시간을 갖겠습니다.
#
# 먼저 네이버에서 뉴스 기사를 간단하게 크롤링합니다.
# 기본적인 전처리 이후 Term-document Matrix를 만들고 이를 non-negative factorization을 이용해 행렬 분해를 하여 Topic modeling을 수행합니다.
#
# t-distributed stochastic neighbor embedding(T-SNE) 기법을 이용해 Topic별 시각화를 진행합니다.
# ```
# + id="NIARcrg_oNMN" outputId="d406e33b-dba0-4b71-dcc0-01ec933ee7e8" colab={"base_uri": "https://localhost:8080/"}
# !pip install newspaper3k
# !pip install konlpy
# + id="aPu9TCFEoK_m"
# 크롤링에 필요한 패키지 설치
from bs4 import BeautifulSoup
from newspaper import Article
from time import sleep
from time import time
from dateutil.relativedelta import relativedelta
from datetime import datetime
from multiprocessing import Pool
import json
import requests
import re
import sys
# + [markdown] id="oU2ipk6ZvZXf"
# ```
# 💡 Crawling(크롤링)이란?
#
# 크롤링은 웹 페이지에서 필요한 데이터를 추출해내는 작업을 말합니다.
# 이번 시간에는 정적 페이지인 네이버의 뉴스 신문 기사 웹페이지를 크롤링합니다.
#
# HTML은 설명되어 있는 자료가 많기 때문에 생략하도록 하겠습니다.
# HTML 구조 파악 및 태그에 대한 설명은 아래 참고자료를 살펴봐주세요 !
# ```
#
# 참고: [위키피디아: 정적페이지](https://ko.wikipedia.org/wiki/%EC%A0%95%EC%A0%81_%EC%9B%B9_%ED%8E%98%EC%9D%B4%EC%A7%80)
#
# 참고: [생활코딩: HTML](https://opentutorials.org/course/2039)
# + id="c0UYQWmsnHTF"
def crawl_news(query: str=None, crawl_num: int=1000, workers: int=4):
'''뉴스 기사 텍스트가 담긴 list를 반환합니다.
Keyword arguments:
query -- 검색어 (default None)
crawl_num -- 수집할 뉴스 기사의 개수 (defualt 1000)
workers -- multi-processing시 사용할 thread의 개수 (default 4)
'''
url = 'https://search.naver.com/search.naver?where=news&sm=tab_jum&query={}'
articleList = []
crawled_url = set()
keyboard_interrupt = False
t = time()
idx = 0
page = 1
# 서버에 url 요청의 결과를 선언
res = requests.get(url.format(query))
sleep(0.5)
# res를 parsing할 parser를 선언
bs = BeautifulSoup(res.text, 'html.parser')
with Pool(workers) as p:
while idx < crawl_num:
table = bs.find('ul', {'class': 'list_news'})
li_list = table.find_all('li', {'id': re.compile('sp_nws.*')})
area_list = [li.find('div', {'class':'news_area'}) for li in li_list]
a_list = [area.find('a', {'class':'news_tit'}) for area in area_list]
for n in a_list[:min(len(a_list), crawl_num-idx)]:
articleList.append(n.get('title'))
idx += 1
page += 1
pages = bs.find('div', {'class': 'sc_page_inner'})
next_page_url = [p for p in pages.find_all('a') if p.text == str(page)][0].get('href')
req = requests.get('https://search.naver.com/search.naver' + next_page_url)
bs = BeautifulSoup(req.text, 'html.parser')
return articleList
# + [markdown] id="8ln1Wih0XVnt"
# ```
# 🔥 이제 '구글'이라는 이름으로 뉴스 기사 1000개의 제목을 크롤링하겠습니다.
# ```
# + id="G6CAFa6J56yJ"
query = '구글'
articleList = crawl_news(query)
# + id="UDIS5V1yAcRT" outputId="551cc979-b549-48f5-f5c0-8aa794c2f4fe" colab={"base_uri": "https://localhost:8080/"}
articleList[:10]
# + [markdown] id="J9HG4LxOaa1r"
# ```
# 🔥 태거(tagger)를 이용해 한글 명사와 알파벳만을 추출해서 term-document matrix (tdm)을 만들겠습니다.
#
# 태거(tagger)는 tokenization에서 조금 더 자세히 다루도록 하겠습니다.
# ```
#
# 참고: [konlpy: morph analyzer](https://konlpy-ko.readthedocs.io/ko/v0.4.3/morph/)
# + [markdown] id="Mdr-aTytaCLr"
# ## Preprocessing
# + id="HUN9k5DXaCLs" outputId="7c4b75b8-a48c-436a-a5f8-63b61b41ce96" colab={"base_uri": "https://localhost:8080/"}
from konlpy.tag import Okt
from collections import Counter
import json
# Okt 형태소 분석기 선언
t = Okt()
words_list_ = []
vocab = Counter()
tag_set = set(['Noun', 'Alpha'])
stopwords = set(['글자'])
for i, article in enumerate(articleList):
if i % 100 == 0:
print(i)
# tagger를 이용한 품사 태깅
words = t.pos(article, norm=True, stem=True)
############################ ANSWER HERE ################################
# TODO: 다음의 조건을 만족하는 단어의 리스트를 완성하세요.
# 조건 1: 명사와 알파벳 tag를 가진 단어
# 조건 2: 철자 길이가 2이상인 단어
# 조건 3: stopwords에 포함되지 않는 단어
words = [w for w,t in words if t in tag_set and len(w) > 1 and w not in stopwords]
#########################################################################
vocab.update(words)
words_list_.append((words, article))
vocab = sorted([w for w, freq in vocab.most_common(10000)])
word2id = {w: i for i, w in enumerate(vocab)}
words_list = []
for words, article in words_list_:
words = [w for w in words if w in word2id]
if len(words) > 10:
words_list.append((words, article))
del words_list_
# + [markdown] id="5L1BkQeaaCLv"
# ## Build document-term matrix
# + [markdown] id="r5YfokxJf86R"
# ```
# 🔥 이제 document-term matrix를 만들어보겠습니다.
# document-term matrix는 (문서 개수 x 단어 개수)의 Matrix입니다.
# ```
#
# 참고: [Document-Term Matrix](https://wikidocs.net/24559)
# + id="MI5weWREaCLv" outputId="0c3177a2-a43c-4b3a-a2fc-24823553d826" colab={"base_uri": "https://localhost:8080/"}
from sklearn.feature_extraction.text import TfidfTransformer
import numpy as np
dtm = np.zeros((len(words_list), len(vocab)), dtype=np.float32)
print(dtm.shape)
for i, (words, article) in enumerate(words_list):
for word in words:
dtm[i, word2id[word]] += 1
print('dtm : ', dtm)
dtm = TfidfTransformer().fit_transform(dtm)
# + [markdown] id="I0xYqpVtmgoL"
# ```
# 🔥 document-term matrix를 non-negative factorization(NMF)을 이용해 행렬 분해를 해보겠습니다.
#
# 💡 Non-negative Factorization이란?
#
# NMF는 주어진 행렬 non-negative matrix X를 non-negative matrix W와 H로 행렬 분해하는 알고리즘입니다.
# 이어지는 코드를 통해 W와 H의 의미에 대해 파악해봅시다.
# ```
# 참고: [Non-negative Matrix Factorization](https://angeloyeo.github.io/2020/10/15/NMF.html)
# + [markdown] id="zRWNuIguaCLy"
# ## Topic modeling
# + id="erhwS2ntQBSD"
# Non-negative Matrix Factorization
from sklearn.decomposition import NMF
K=5
nmf = NMF(n_components=K, alpha=0.1)
# + [markdown] id="5I24Y78G3VvO"
# ```
# 🔥 sklearn의 NMF를 이용해 W와 H matrix를 구해봅시다.
# W는 document length x K, H는 K x term length의 차원을 갖고 있습니다.
# W의 하나의 row는 각각의 feature에 얼만큼의 가중치를 줄 지에 대한 weight입니다.
# H의 하나의 row는 하나의 feature를 나타냅니다.
#
# 우선 하나의 Topic (H의 n번째 row)에 접근해서 해당 topic에 대해 값이 가장 높은 20개의 단어를 출력해보겠습니다.
# ```
# + id="v2TY6gu4QH1o" outputId="b799f5f0-4be9-41b9-908f-83a0bf995b5a" colab={"base_uri": "https://localhost:8080/"}
W = nmf.fit_transform(dtm)
H = nmf.components_
# + [markdown] id="kbNWlTAv5Zn2"
# ```
# 🔥 우선 하나의 Topic (H의 n번째 row)에 접근해서 해당 topic에 대해 값이 가장 높은 20개의 단어를 출력해보겠습니다.
# ```
# + id="uSzB5PBuaCL2" outputId="bc775213-b4f9-4903-d534-b8ed590cf5c3" colab={"base_uri": "https://localhost:8080/"}
for k in range(K):
print(f"{k}th topic")
for index in H[k].argsort()[::-1][:20]:
print(vocab[index], end=' ')
print()
# + [markdown] id="PTP6lUft5C4j"
# ```
# 🔥 이번에는 W에서 하나의 Topic (W의 n번째 column)에 접근해서 해당 topic에 대해 값이 가장 높은 3개의 뉴스 기사 제목을 출력해보겠습니다.
# ```
# + id="K1CniIn7aCL5" outputId="bb3ca8e3-1731-4500-bd49-72ca6c0d55e8" colab={"base_uri": "https://localhost:8080/"}
for k in range(K):
print(f"==={k}th topic===")
for index in W[:, k].argsort()[::-1][:3]:
print(words_list[index][1])
print('\n')
# + [markdown] id="yLOplJlO5tFi"
# ```
# ❓ 2번째 토픽에 대해 가장 높은 가중치를 갖는 제목 5개를 출력해볼까요?
# ```
# + id="Jfm4fK4jaCL9" outputId="0d9f78d1-b0f7-4a8e-bae4-35050b7cdbd9" colab={"base_uri": "https://localhost:8080/"}
#TODO
index_list = W[:, 2].argsort()[::-1][:5]
for index in index_list:
print(words_list[index][1])
# + [markdown] id="b7AZ3SyS6Y5L"
# ```
# 🔥 이번에는 t-SNE를 이용해 Topic별 시각화를 진행해보겠습니다.
#
# 💡 t-SNE는 무엇인가요?
#
# t-Stochastic Neighbor Embedding(t-SNE)은 고차원의 벡터를
# 저차원(2~3차원) 벡터로 데이터간 구조적 특징을 유지하며 축소를 하는 방법 중 하나입니다.
#
# 주로 고차원 데이터의 시각화를 위해 사용됩니다.
# ```
#
# 참고: [lovit: t-SNE](https://lovit.github.io/nlp/representation/2018/09/28/tsne/#:~:text=t%2DSNE%20%EB%8A%94%20%EA%B3%A0%EC%B0%A8%EC%9B%90%EC%9D%98,%EC%9D%98%20%EC%A7%80%EB%8F%84%EB%A1%9C%20%ED%91%9C%ED%98%84%ED%95%A9%EB%8B%88%EB%8B%A4.)
#
# 참고: [ratsgo: t-SNE](https://ratsgo.github.io/machine%20learning/2017/04/28/tSNE/)
# + [markdown] id="H1HlyoakaCMA"
# ## Visualization
# + id="rqajA3lDaCMB" outputId="091e4379-4c15-49cf-c60d-9fe9ca8f0411" colab={"base_uri": "https://localhost:8080/"}
from sklearn.manifold import TSNE
# n_components = 차원 수
tsne = TSNE(n_components=2, init='pca', verbose=1)
# W matrix에 대해 t-sne를 수행합니다.
W2d = tsne.fit_transform(W)
# 각 뉴스 기사 제목마다 가중치가 가장 높은 topic을 저장합니다.
topicIndex = [v.argmax() for v in W]
# + id="pc76X-jSaCME" outputId="6489acce-0e79-4027-a30a-3588fe42e852" colab={"base_uri": "https://localhost:8080/", "height": 634}
from bokeh.models import HoverTool
from bokeh.palettes import Category20
from bokeh.io import show, output_notebook
from bokeh.plotting import figure, ColumnDataSource
output_notebook()
# 사용할 툴들
tools_to_show = 'hover,box_zoom,pan,save,reset,wheel_zoom'
p = figure(plot_width=720, plot_height=580, tools=tools_to_show)
source = ColumnDataSource(data={
'x': W2d[:, 0],
'y': W2d[:, 1],
'id': [i for i in range(W.shape[0])],
'document': [article for words, article in words_list],
'topic': [str(i) for i in topicIndex], # 토픽 번호
'color': [Category20[K][i] for i in topicIndex]
})
p.circle(
'x', 'y',
source=source,
legend='topic',
color='color'
)
# interaction
p.legend.location = "top_left"
hover = p.select({'type': HoverTool})
hover.tooltips = [("Topic", "@topic"), ('id', '@id'), ("Article", "@document")]
hover.mode = 'mouse'
show(p)
|
06_Natural_Language_Processing/sol/[HW25_Problem]_Topic_Modeling.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="xBF9RPBhw2we"
# ### Dataset Reading
# + id="pN4tMIn1w2wg" executionInfo={"status": "ok", "timestamp": 1603190663089, "user_tz": -330, "elapsed": 3282, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="c5cac573-b711-4a7b-c6fb-03229be37505" colab={"base_uri": "https://localhost:8080/", "height": 289}
import pandas as pd
data = pd.read_excel('drive/My Drive/Constraint_Competition_Dataset/Constraint_Covid-19_English_Train.xlsx')
pd.set_option('display.max_colwidth',150)
data.head()
# + id="O9ABoWjOw2wl" executionInfo={"status": "ok", "timestamp": 1603190663953, "user_tz": -330, "elapsed": 4127, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="f6773b8e-8b88-47e9-8b36-a21fabba6f75" colab={"base_uri": "https://localhost:8080/", "height": 35}
data.shape
# + id="JSKI3CX6w2wp" executionInfo={"status": "ok", "timestamp": 1603190663954, "user_tz": -330, "elapsed": 4114, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="a32bdd68-98a0-4ad8-ddf1-d24abc6ec7e7" colab={"base_uri": "https://localhost:8080/", "height": 87}
print(data.dtypes)
# + [markdown] id="XNsif5VGw2ws"
# ### Making of "label" Variable
# + id="gwE60IAxw2ws" executionInfo={"status": "ok", "timestamp": 1603190663955, "user_tz": -330, "elapsed": 4102, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="10da318d-291c-4283-c958-669794685c2c" colab={"base_uri": "https://localhost:8080/", "height": 121}
label = data['label']
label.head()
# + [markdown] id="ShrD5Y7ew2wv"
# ### Checking Dataset Balancing
# + id="kFui_Mz3w2wv" executionInfo={"status": "ok", "timestamp": 1603190663956, "user_tz": -330, "elapsed": 4092, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="c1bc63bd-c15e-4d34-a9c7-645237a319f5" colab={"base_uri": "https://localhost:8080/", "height": 349}
print(label.value_counts())
import matplotlib.pyplot as plt
label.value_counts().plot(kind='bar', color='blue')
# + [markdown] id="MRSdKLNiw2wx"
# ### Convering label into "0" or "1"
# + id="0ESnvF3Vw2wy" executionInfo={"status": "ok", "timestamp": 1603190663957, "user_tz": -330, "elapsed": 4080, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="ee166daf-4ab4-4463-818b-370c090eb7bc" colab={"base_uri": "https://localhost:8080/", "height": 35}
import numpy as np
classes_list = ["fake","real"]
label_index = data['label'].apply(classes_list.index)
final_label = np.asarray(label_index)
print(final_label[:10])
# + id="NSuVpENKGBWU" executionInfo={"status": "ok", "timestamp": 1603190663957, "user_tz": -330, "elapsed": 4077, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
from keras.utils.np_utils import to_categorical
label_twoDimension = to_categorical(final_label, num_classes=2)
# + id="GtlQzqdpGMBM" executionInfo={"status": "ok", "timestamp": 1603190663958, "user_tz": -330, "elapsed": 4062, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="a2a5d9aa-0e6f-4d9a-ff4e-7cf171d60631" colab={"base_uri": "https://localhost:8080/", "height": 191}
print(label_twoDimension[:10])
# + [markdown] id="s2JSVKo3w2w0"
# ### Making of "text" Variable
# + id="-VK4ScnGw2w1" executionInfo={"status": "ok", "timestamp": 1603190663959, "user_tz": -330, "elapsed": 4048, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="e5635fb7-4127-48e9-e992-012c555eaf87" colab={"base_uri": "https://localhost:8080/", "height": 228}
text = data['tweet']
text.head(10)
# + [markdown] id="tbKm17HIw2w3"
# ### Dataset Pre-processing
# + id="_Sf_xhO6w2w7" executionInfo={"status": "ok", "timestamp": 1603190664809, "user_tz": -330, "elapsed": 4895, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
import re
def text_clean(text):
''' Pre process and convert texts to a list of words '''
text=text.lower()
# Clean the text
text = re.sub(r"[^A-Za-z0-9^,!.\/'+-=]", " ", text)
text = re.sub(r"what's", "what is ", text)
text = re.sub(r"I'm", "I am ", text)
text = re.sub(r"\'s", " ", text)
text = re.sub(r"\'ve", " have ", text)
text = re.sub(r"can't", "cannot ", text)
text = re.sub(r"wouldn't", "would not ", text)
text = re.sub(r"shouldn't", "should not ", text)
text = re.sub(r"shouldn", "should not ", text)
text = re.sub(r"didn", "did not ", text)
text = re.sub(r"n't", " not ", text)
text = re.sub(r"i'm", "i am ", text)
text = re.sub(r"\'re", " are ", text)
text = re.sub(r"\'d", " would ", text)
text = re.sub(r"\'ll", " will ", text)
text = re.sub('https?://\S+|www\.\S+', "", text)
text = re.sub(r",", " ", text)
text = re.sub(r"\.", " ", text)
text = re.sub(r"!", " ! ", text)
text = re.sub(r"\/", " ", text)
text = re.sub(r"\^", " ^ ", text)
text = re.sub(r"\+", " + ", text)
text = re.sub(r"\-", " - ", text)
text = re.sub(r"\=", " = ", text)
text = re.sub(r"'", " ", text)
text = re.sub(r"(\d+)(k)", r"\g<1>000", text)
text = re.sub(r":", " : ", text)
text = re.sub(r" e g ", " eg ", text)
text = re.sub(r" b g ", " bg ", text)
text = re.sub(r" u s ", " american ", text)
text = re.sub(r"\0s", "0", text)
text = re.sub(r" 9 11 ", "911", text)
text = re.sub(r"e - mail", "email", text)
text = re.sub(r"j k", "jk", text)
text = re.sub(r"\s{2,}", " ", text)
text = re.sub(r"[0-9]", "", text)
# text = re.sub(r"rt", " ", text)
return text
# + id="5_JQL5rRw2xA" executionInfo={"status": "ok", "timestamp": 1603190664810, "user_tz": -330, "elapsed": 4891, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
clean_text = text.apply(lambda x:text_clean(x))
# + id="A_uqquBZw2xE" executionInfo={"status": "ok", "timestamp": 1603190664811, "user_tz": -330, "elapsed": 4876, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="7fcbb23e-89f7-4667-b6cb-d4e00bf6f188" colab={"base_uri": "https://localhost:8080/", "height": 228}
clean_text.head(10)
# + [markdown] id="AGYA06eJw2xJ"
# ### Removing stopwords
# + id="JBLDOKifw2xK" executionInfo={"status": "ok", "timestamp": 1603190665502, "user_tz": -330, "elapsed": 5553, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="eedc4390-e661-4fe7-e2f0-ca39ca6141c3" colab={"base_uri": "https://localhost:8080/", "height": 52}
import nltk
from nltk.corpus import stopwords
nltk.download('stopwords')
def stop_words_removal(text1):
text1=[w for w in text1.split(" ") if w not in stopwords.words('english')]
return " ".join(text1)
# + id="dwSLSw3Nw2xN" executionInfo={"status": "ok", "timestamp": 1603190683633, "user_tz": -330, "elapsed": 23681, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
clean_text_ns=clean_text.apply(lambda x: stop_words_removal(x))
# + id="OFjJCsd_w2xQ" executionInfo={"status": "ok", "timestamp": 1603190683637, "user_tz": -330, "elapsed": 23668, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="0574432a-8c32-4b12-93c7-2940e0378c80" colab={"base_uri": "https://localhost:8080/", "height": 228}
print(clean_text_ns.head(10))
# + [markdown] id="Vxq3KDt4w2xS"
# ### Lemmatization
# + id="FlGoDlLmw2xT" executionInfo={"status": "ok", "timestamp": 1603190683638, "user_tz": -330, "elapsed": 23653, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="c40b5301-5f00-4d33-eaf5-6dbcf491754a" colab={"base_uri": "https://localhost:8080/", "height": 70}
"""# Lemmatization
import nltk
nltk.download('wordnet')
from nltk.stem import WordNetLemmatizer
lemmatizer = WordNetLemmatizer()
def word_lemmatizer(text):
lem_text = "".join([lemmatizer.lemmatize(i) for i in text])
return lem_text"""
# + id="desz-r2qw2xW" executionInfo={"status": "ok", "timestamp": 1603190683638, "user_tz": -330, "elapsed": 23636, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="055d659e-c7cf-4aea-9d4d-51a553689b09" colab={"base_uri": "https://localhost:8080/", "height": 35}
"""clean_text_lem = clean_text_ns.apply(lambda x : word_lemmatizer(x))"""
# + id="OuhsiibOw2xY" executionInfo={"status": "ok", "timestamp": 1603190683638, "user_tz": -330, "elapsed": 23624, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="c84e853f-a33e-437b-b517-22950f95fb2d" colab={"base_uri": "https://localhost:8080/", "height": 35}
"""print(clean_text_lem.head(10))"""
# + [markdown] id="96IyUsaow2xa"
# ### Stemming
# + id="2TuWAy4bw2xb" executionInfo={"status": "ok", "timestamp": 1603190683639, "user_tz": -330, "elapsed": 23623, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
# Stemming
from nltk.stem import PorterStemmer
stemmer = PorterStemmer()
def word_stemmer(text):
stem_text = "".join([stemmer.stem(i) for i in text])
return stem_text
# + id="ivl__lJWw2xe" executionInfo={"status": "ok", "timestamp": 1603190683639, "user_tz": -330, "elapsed": 23620, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
clean_text_stem = clean_text_ns.apply(lambda x : word_stemmer(x))
# + id="qoMbymPmw2xf" executionInfo={"status": "ok", "timestamp": 1603190683640, "user_tz": -330, "elapsed": 23609, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="473fc26d-3698-4412-e0a4-6b4f6ee6a8ae" colab={"base_uri": "https://localhost:8080/", "height": 228}
print(clean_text_stem.head(10))
# + id="0UFWzUEcw2xh" executionInfo={"status": "ok", "timestamp": 1603190683640, "user_tz": -330, "elapsed": 23607, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
# final_text = [x for x in clean_text_lem if len(x) > 3]
# + id="15kD9mAWw2xj" executionInfo={"status": "ok", "timestamp": 1603190683641, "user_tz": -330, "elapsed": 23605, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
#print(final_text)
# + [markdown] id="LyORidvKw2xl"
# ### Tokenization using "keras"
# + id="feW2fI8Dw2xl" executionInfo={"status": "ok", "timestamp": 1603190683641, "user_tz": -330, "elapsed": 23602, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
import keras
import tensorflow
from keras.preprocessing.text import Tokenizer
tok_all = Tokenizer(filters='!"#$%&\'()*+,-./:;<=>?@[\\]^_`{|}~', lower=True, char_level = False)
tok_all.fit_on_texts(clean_text_stem)
# + [markdown] id="pVf7lAKJw2xo"
# ### Making Vocab for words
# + id="LtBxjGZKw2xo" executionInfo={"status": "ok", "timestamp": 1603190683641, "user_tz": -330, "elapsed": 23587, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="6f51943b-30ce-4050-bf42-384d9531442c" colab={"base_uri": "https://localhost:8080/", "height": 35}
vocabulary_all = len(tok_all.word_counts)
print(vocabulary_all)
# + id="PKAhcecYw2xr" executionInfo={"status": "ok", "timestamp": 1603190685292, "user_tz": -330, "elapsed": 25213, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="fcdd8386-a230-4a12-85a7-d6c1dd4ec27c" colab={"base_uri": "https://localhost:8080/", "height": 55}
l = tok_all.word_index
print(l)
# + [markdown] id="wLKyeIYHw2xu"
# ### encoding or sequencing
# + id="5tTNFeyrw2xu" executionInfo={"status": "ok", "timestamp": 1603190685293, "user_tz": -330, "elapsed": 25199, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="bee4c902-5425-4d0d-8d53-82a45dbe7df7" colab={"base_uri": "https://localhost:8080/", "height": 52}
encoded_clean_text_stem = tok_all.texts_to_sequences(clean_text_stem)
print(clean_text_stem[1])
print(encoded_clean_text_stem[1])
# + [markdown] id="ao425zSrw2xw"
# ### Pre-padding
# + id="mJB28ImAw2xw" executionInfo={"status": "ok", "timestamp": 1603190685295, "user_tz": -330, "elapsed": 25197, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
from keras.preprocessing import sequence
max_length = 100
padded_clean_text_stem = sequence.pad_sequences(encoded_clean_text_stem, maxlen=max_length, padding='pre')
# + [markdown] id="lEigFn5fWFAs"
# # Test Data Pre-processing
# + [markdown] id="4zQ1QbtFWX_J"
# # Data test Reading
# + id="F0wlDEHwWOlx" executionInfo={"status": "ok", "timestamp": 1603190685295, "user_tz": -330, "elapsed": 25184, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="828a267f-8c87-4ec2-dad4-0b5cdc77c40a" colab={"base_uri": "https://localhost:8080/", "height": 254}
data_t = pd.read_excel('drive/My Drive/Constraint_Competition_Dataset/Constraint_Covid-19_English_Val.xlsx')
pd.set_option('display.max_colwidth',150)
data_t.head()
# + id="W5bwz_-dWyui" executionInfo={"status": "ok", "timestamp": 1603190685296, "user_tz": -330, "elapsed": 25171, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="19dd6bbb-7be2-484e-94e7-8bf1865e2a15" colab={"base_uri": "https://localhost:8080/", "height": 35}
data_t.shape
# + id="ntkVP_FiW4vn" executionInfo={"status": "ok", "timestamp": 1603190685297, "user_tz": -330, "elapsed": 25153, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="ef17fd5b-a16f-4fee-e7e2-4a83ebae1859" colab={"base_uri": "https://localhost:8080/", "height": 87}
print(data_t.dtypes)
# + [markdown] id="Ocyn5IEDXAr7"
# # Making of "label" Variable
# + id="bAglc2pzXDpJ" executionInfo={"status": "ok", "timestamp": 1603190685298, "user_tz": -330, "elapsed": 25138, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="943a5d9f-a500-4476-a5c2-603193a67595" colab={"base_uri": "https://localhost:8080/", "height": 121}
label_t = data_t['label']
label_t.head()
# + [markdown] id="VVxcyv1uYhUV"
# # Checking Dataset Balancing
# + id="2GJE9j_OW5kG" executionInfo={"status": "ok", "timestamp": 1603190685298, "user_tz": -330, "elapsed": 25124, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="cddae900-a637-485e-892c-eadb300bc0ea" colab={"base_uri": "https://localhost:8080/", "height": 347}
print(label_t.value_counts())
import matplotlib.pyplot as plt
label_t.value_counts().plot(kind='bar', color='red')
# + [markdown] id="Kq3obUM1Y3v3"
# # Convering label into "0" or "1"
# + id="0V7LGxK_ZA4S" executionInfo={"status": "ok", "timestamp": 1603190685299, "user_tz": -330, "elapsed": 25109, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="45f6455f-0e40-4b03-9114-7dfcad80e571" colab={"base_uri": "https://localhost:8080/", "height": 35}
import numpy as np
classes_list_t = ["fake","real"]
label_t_index = data_t['label'].apply(classes_list_t.index)
final_label_t = np.asarray(label_t_index)
print(final_label_t[:10])
# + id="4Ve8y_srZA75" executionInfo={"status": "ok", "timestamp": 1603190685300, "user_tz": -330, "elapsed": 25106, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
from keras.utils.np_utils import to_categorical
label_twoDimension_t = to_categorical(final_label_t, num_classes=2)
# + id="3rmVyCfKZSxz" executionInfo={"status": "ok", "timestamp": 1603190685300, "user_tz": -330, "elapsed": 25092, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="e366a1fe-81b0-4cfa-89c2-8ebd2377ec74" colab={"base_uri": "https://localhost:8080/", "height": 191}
print(label_twoDimension_t[:10])
# + [markdown] id="R5NMHXF6ZZJj"
# # Making of "text" Variable
# + id="BFFgaFBHZomG" executionInfo={"status": "ok", "timestamp": 1603190685301, "user_tz": -330, "elapsed": 25077, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="4e15c4e5-7f70-426a-e20f-4cfbc0ee3985" colab={"base_uri": "https://localhost:8080/", "height": 228}
text_t = data_t['tweet']
text_t.head(10)
# + [markdown] id="wdok08rOZwro"
# # **Dataset Pre-processing**
# 1. Remove unwanted words
# 2. Stopwords removal
# 3. Stemming
# 4. Tokenization
# 5. Encoding or Sequencing
# 6. Pre-padding
# + [markdown] id="QrxT9sK5bUs3"
# ### 1. Removing Unwanted Words
# + id="eapxovvvavlO" executionInfo={"status": "ok", "timestamp": 1603190685991, "user_tz": -330, "elapsed": 25763, "user": {"displayName": "RAUSHAN Raj", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
import re
def text_clean(text):
''' Pre process and convert texts to a list of words '''
text=text.lower()
# Clean the text
text = re.sub(r"[^A-Za-z0-9^,!.\/'+-=]", " ", text)
text = re.sub(r"what's", "what is ", text)
text = re.sub(r"I'm", "I am ", text)
text = re.sub(r"\'s", " ", text)
text = re.sub(r"\'ve", " have ", text)
text = re.sub(r"can't", "cannot ", text)
text = re.sub(r"wouldn't", "would not ", text)
text = re.sub(r"shouldn't", "should not ", text)
text = re.sub(r"shouldn", "should not ", text)
text = re.sub(r"didn", "did not ", text)
text = re.sub(r"n't", " not ", text)
text = re.sub(r"i'm", "i am ", text)
text = re.sub(r"\'re", " are ", text)
text = re.sub(r"\'d", " would ", text)
text = re.sub(r"\'ll", " will ", text)
text = re.sub('https?://\S+|www\.\S+', "", text)
text = re.sub(r",", " ", text)
text = re.sub(r"\.", " ", text)
text = re.sub(r"!", " ! ", text)
text = re.sub(r"\/", " ", text)
text = re.sub(r"\^", " ^ ", text)
text = re.sub(r"\+", " + ", text)
text = re.sub(r"\-", " - ", text)
text = re.sub(r"\=", " = ", text)
text = re.sub(r"'", " ", text)
text = re.sub(r"(\d+)(k)", r"\g<1>000", text)
text = re.sub(r":", " : ", text)
text = re.sub(r" e g ", " eg ", text)
text = re.sub(r" b g ", " bg ", text)
text = re.sub(r" u s ", " american ", text)
text = re.sub(r"\0s", "0", text)
text = re.sub(r" 9 11 ", "911", text)
text = re.sub(r"e - mail", "email", text)
text = re.sub(r"j k", "jk", text)
text = re.sub(r"\s{2,}", " ", text)
text = re.sub(r"[0-9]", "", text)
# text = re.sub(r"rt", " ", text)
return text
# + id="ZKXhURU5a0q-" executionInfo={"status": "ok", "timestamp": 1603190685991, "user_tz": -330, "elapsed": 25759, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
clean_text_t = text_t.apply(lambda x:text_clean(x))
# + id="4R6Paqqia0y_" executionInfo={"status": "ok", "timestamp": 1603190685992, "user_tz": -330, "elapsed": 25749, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="1d6e95b1-d13e-4f35-dbcb-c05e0746bcb1" colab={"base_uri": "https://localhost:8080/", "height": 228}
clean_text_t.head(10)
# + [markdown] id="lyxeJ7xtbB5-"
# ### 2. Removing Stopwords
# + id="yfdc4WLNbIYP" executionInfo={"status": "ok", "timestamp": 1603190685992, "user_tz": -330, "elapsed": 25738, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="6b64226a-e1aa-49f6-8f5c-65a5df365192" colab={"base_uri": "https://localhost:8080/", "height": 52}
import nltk
from nltk.corpus import stopwords
nltk.download('stopwords')
def stop_words_removal(text1):
text1=[w for w in text1.split(" ") if w not in stopwords.words('english')]
return " ".join(text1)
# + id="7lH4FtPtbfmc" executionInfo={"status": "ok", "timestamp": 1603190691478, "user_tz": -330, "elapsed": 31221, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
clean_text_t_ns=clean_text_t.apply(lambda x: stop_words_removal(x))
# + id="xSzxQQE0bfpw" executionInfo={"status": "ok", "timestamp": 1603190691481, "user_tz": -330, "elapsed": 31215, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="facf59f1-6d17-4ae8-d00e-fc54ecff65b2" colab={"base_uri": "https://localhost:8080/", "height": 228}
print(clean_text_t_ns.head(10))
# + [markdown] id="9VkXLxaMbpqb"
# ### 3. Stemming
# + id="2gEVoc0fbu1m" executionInfo={"status": "ok", "timestamp": 1603190691482, "user_tz": -330, "elapsed": 31213, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
# Stemming
from nltk.stem import PorterStemmer
stemmer = PorterStemmer()
def word_stemmer(text):
stem_text = "".join([stemmer.stem(i) for i in text])
return stem_text
# + id="RnIAjbL7bvon" executionInfo={"status": "ok", "timestamp": 1603190691482, "user_tz": -330, "elapsed": 31210, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
clean_text_t_stem = clean_text_t_ns.apply(lambda x : word_stemmer(x))
# + id="hywyHMQ8bz9B" executionInfo={"status": "ok", "timestamp": 1603190691483, "user_tz": -330, "elapsed": 31202, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="a07f06c3-1d0e-4c31-8fb3-72ad97a20d63" colab={"base_uri": "https://localhost:8080/", "height": 228}
print(clean_text_t_stem.head(10))
# + [markdown] id="gNW4AywXb4ZL"
# ### 4. Tokenization
# + id="F-79JOmgb_io" executionInfo={"status": "ok", "timestamp": 1603190691483, "user_tz": -330, "elapsed": 31199, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
import keras
import tensorflow
from keras.preprocessing.text import Tokenizer
tok_test = Tokenizer(filters='!"#$%&\'()*+,-./:;<=>?@[\\]^_`{|}~', lower=True, char_level = False)
tok_test.fit_on_texts(clean_text_t_stem)
# + id="4YCYJtTKcKe-" executionInfo={"status": "ok", "timestamp": 1603190691484, "user_tz": -330, "elapsed": 31191, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="500ac6ef-0918-422e-993c-bd56bc5d922c" colab={"base_uri": "https://localhost:8080/", "height": 35}
vocabulary_all_test = len(tok_test.word_counts)
print(vocabulary_all_test)
# + id="9UCJEGCMcOri" executionInfo={"status": "ok", "timestamp": 1603190691484, "user_tz": -330, "elapsed": 31182, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="83f06fcd-19e0-46a4-f583-9913884bf13f" colab={"base_uri": "https://localhost:8080/", "height": 55}
test_list = tok_test.word_index
print(test_list)
# + [markdown] id="qZeXZbM5cPm5"
# ### 5. Encoding or Sequencing
# + id="88IUoE2tcavl" executionInfo={"status": "ok", "timestamp": 1603190691485, "user_tz": -330, "elapsed": 31174, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="7bcf7ba2-433b-443a-899f-3cdda88141e8" colab={"base_uri": "https://localhost:8080/", "height": 52}
encoded_clean_text_t_stem = tok_all.texts_to_sequences(clean_text_t_stem)
print(clean_text_t_stem[0])
print(encoded_clean_text_t_stem[0])
# + [markdown] id="2qg4xgewcjLG"
# ### 6. Pre-padding
# + id="arj7T2r1coOw" executionInfo={"status": "ok", "timestamp": 1603190691485, "user_tz": -330, "elapsed": 31172, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
from keras.preprocessing import sequence
max_length = 100
padded_clean_text_t_stem = sequence.pad_sequences(encoded_clean_text_t_stem, maxlen=max_length, padding='pre')
# + [markdown] id="QfhyZliqgYTb"
# # GloVe Embedding
# + id="G4S7PI9cw2xy" executionInfo={"status": "ok", "timestamp": 1603190725849, "user_tz": -330, "elapsed": 65526, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="44a5cf07-adb1-4c0b-8cbe-eab5ac61d8ef" colab={"base_uri": "https://localhost:8080/", "height": 35}
# GloVe Embedding link - https://nlp.stanford.edu/projects/glove/
import os
import numpy as np
embeddings_index = {}
f = open('drive/My Drive/HASOC Competition Data/Copy of glove.6B.300d.txt')
for line in f:
values = line.split()
word = values[0]
coefs = np.asarray(values[1:], dtype='float32')
embeddings_index[word] = coefs
f.close()
print('Loaded %s word vectors.' % len(embeddings_index))
# + id="7-9fLmPZzlP_" executionInfo={"status": "ok", "timestamp": 1603190725850, "user_tz": -330, "elapsed": 65524, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
embedding_matrix = np.zeros((vocabulary_all+1, 300))
for word, i in tok_all.word_index.items():
embedding_vector = embeddings_index.get(word)
if embedding_vector is not None:
embedding_matrix[i] = embedding_vector
# + [markdown] id="oM5OmlqZgrLy"
# # **CNN Model**
# + id="r2VGeKXv0vOz" executionInfo={"status": "ok", "timestamp": 1603190725850, "user_tz": -330, "elapsed": 65522, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
from keras.preprocessing import sequence
from keras.preprocessing import text
import numpy as np
from keras.models import Sequential
from keras.layers import Dense, Dropout, Activation
from keras.layers import Embedding, LSTM , Bidirectional
from keras.layers import Conv1D, Flatten
from keras.preprocessing import text
from keras.models import Sequential,Model
from keras.layers import Dense ,Activation,MaxPool1D,Conv1D,Flatten,Dropout,Activation,Dropout,Input,Lambda,concatenate
from keras.utils import np_utils
from nltk.corpus import stopwords
from nltk.tokenize import RegexpTokenizer
from nltk.stem.porter import PorterStemmer
import nltk
import csv
import pandas as pd
from keras.preprocessing import text as keras_text, sequence as keras_seq
# + id="qr8uLf-q0lPJ" executionInfo={"status": "ok", "timestamp": 1603190732566, "user_tz": -330, "elapsed": 72236, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
Bi_lstm1_network = Sequential()
# Adding Embedding layer
Bi_lstm1_network.add(Embedding(vocabulary_all+1,300,weights = [embedding_matrix], input_length=max_length, trainable= False))
# Adding 1 Bi-Lstm layers
Bi_lstm1_network.add(Bidirectional(LSTM(128, return_sequences=False)))
Bi_lstm1_network.add(Dropout(0.2))
# Adding Dense layer
Bi_lstm1_network.add(Dense(64,activation="relu"))
Bi_lstm1_network.add(Dropout(0.3))
Bi_lstm1_network.add(Dense(2,activation="sigmoid"))
# + id="iqV6VLZ83HH6" executionInfo={"status": "ok", "timestamp": 1603190732567, "user_tz": -330, "elapsed": 72228, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="a39c0180-05af-4d65-ee99-6827e91ca373" colab={"base_uri": "https://localhost:8080/", "height": 364}
Bi_lstm1_network.summary()
# + id="80QTgAc6BMJ1" executionInfo={"status": "ok", "timestamp": 1603190732568, "user_tz": -330, "elapsed": 72227, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
from keras.optimizers import Adam
Bi_lstm1_network.compile(loss = "binary_crossentropy", optimizer=Adam(lr=0.00003), metrics=["accuracy"])
# + id="h9c9ECvp7P9f" executionInfo={"status": "ok", "timestamp": 1603190732569, "user_tz": -330, "elapsed": 72219, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="59b21ead-b184-4940-8011-59e59c0baa9e" colab={"base_uri": "https://localhost:8080/", "height": 754}
from keras.utils.vis_utils import plot_model
plot_model(Bi_lstm1_network, to_file='Bi_lstm1_network.png', show_shapes=True, show_layer_names=True)
# + id="LR0JsV_kAcRY" executionInfo={"status": "ok", "timestamp": 1603190732569, "user_tz": -330, "elapsed": 72217, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
from keras.callbacks import EarlyStopping, ReduceLROnPlateau,ModelCheckpoint
earlystopper = EarlyStopping(patience=8, verbose=1)
reduce_lr = ReduceLROnPlateau(monitor='val_loss', factor=0.9,
patience=2, min_lr=0.00001, verbose=1)
# + [markdown] id="fMnqgj6rhDVR"
# ### **Model Fitting or Training**
# + id="5nbnfnRZAv1M" executionInfo={"status": "ok", "timestamp": 1603191731289, "user_tz": -330, "elapsed": 179226, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="351c9803-215f-4e5f-eff1-218aa8878cf5" colab={"base_uri": "https://localhost:8080/", "height": 1000}
hist = Bi_lstm1_network.fit(padded_clean_text_stem,label_twoDimension,epochs=100,batch_size=64,callbacks=[earlystopper, reduce_lr])
# + [markdown] id="T5W_uxCThTLl"
# # log loss
# + id="X9DBoQg8Cf1G" executionInfo={"status": "ok", "timestamp": 1603191731290, "user_tz": -330, "elapsed": 179214, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
Bi_lstm1_network_predictions = Bi_lstm1_network.predict(padded_clean_text_t_stem)
# + id="bJQznoSlJ5bT" executionInfo={"status": "ok", "timestamp": 1603191731291, "user_tz": -330, "elapsed": 179208, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="07bd340d-3aea-4ffa-c0b1-f867000972ee" colab={"base_uri": "https://localhost:8080/", "height": 35}
from sklearn.metrics import log_loss
log_loss_test= log_loss(label_twoDimension_t,Bi_lstm1_network_predictions)
log_loss_test
# + [markdown] id="MryQdO5YhdEz"
# # Classification Report
# + id="3UhoxZljKBVs" executionInfo={"status": "ok", "timestamp": 1603191731292, "user_tz": -330, "elapsed": 179200, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
predictions = np.zeros_like(Bi_lstm1_network_predictions)
predictions[np.arange(len(Bi_lstm1_network_predictions)), Bi_lstm1_network_predictions.argmax(1)] = 1
# + id="pNAHulQqKP80" executionInfo={"status": "ok", "timestamp": 1603191731292, "user_tz": -330, "elapsed": 179194, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="9c8b68f1-d871-484d-fea9-212b5a1d964f" colab={"base_uri": "https://localhost:8080/", "height": 35}
predictionInteger=(np.argmax(predictions, axis=1))
predictionInteger
# + id="p4zH_CHRSkji" executionInfo={"status": "ok", "timestamp": 1603191731292, "user_tz": -330, "elapsed": 179183, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="a6be7af1-7ef3-46e0-f8a6-4eb26249ca19" colab={"base_uri": "https://localhost:8080/", "height": 35}
'''pred_label = np.array(predictionInteger)
df = pd.DataFrame(data=pred_label , columns=["task1"])
print(df)'''
# + id="gMcD5cG7XLL9" executionInfo={"status": "ok", "timestamp": 1603191731293, "user_tz": -330, "elapsed": 179174, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
# df.to_csv("submission_EN_A.csv", index=False)
# + id="HE-j9PERKXBE" executionInfo={"status": "ok", "timestamp": 1603191731816, "user_tz": -330, "elapsed": 179689, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="079cde56-c794-4dd2-f8cf-cc3e4af288cf" colab={"base_uri": "https://localhost:8080/", "height": 191}
from sklearn.metrics import classification_report
print(classification_report(label_twoDimension_t,predictions))
# + [markdown] id="WqNX-4ljhwsu"
# # Epoch v/s Loss Plot
# + id="Dk322X4pKjEQ" executionInfo={"status": "ok", "timestamp": 1603191731816, "user_tz": -330, "elapsed": 179679, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="7fcbfe6d-94d9-46b4-fc6b-93396d4c9ccf" colab={"base_uri": "https://localhost:8080/", "height": 295}
from matplotlib import pyplot as plt
plt.plot(hist.history["loss"],color = 'red', label = 'train_loss')
#plt.plot(hist.history["val_loss"],color = 'blue', label = 'val_loss')
plt.title('Loss Visualisation')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.savefig('Bi_lstm1_HASOC_Eng_lossPlot.pdf',dpi=1000)
from google.colab import files
files.download('Bi_lstm1_HASOC_Eng_lossPlot.pdf')
# + [markdown] id="A5eYuEVbh0Qi"
# # Epoch v/s Accuracy Plot
# + id="BSDEzNM1LKmp" executionInfo={"status": "ok", "timestamp": 1603191731817, "user_tz": -330, "elapsed": 179670, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="d0256e39-efbd-4019-b17f-ef201866f6b3" colab={"base_uri": "https://localhost:8080/", "height": 295}
plt.plot(hist.history["accuracy"],color = 'red', label = 'train_accuracy')
#plt.plot(hist.history["val_accuracy"],color = 'blue', label = 'val_accuracy')
plt.title('Accuracy Visualisation')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.legend()
plt.savefig('Bi_lstm1_HASOC_Eng_accuracyPlot.pdf',dpi=1000)
files.download('Bi_lstm1_HASOC_Eng_accuracyPlot.pdf')
# + [markdown] id="5v-PNBwUh6fK"
# # Area under Curve-ROC
# + id="rIga22ZbL5Lg" executionInfo={"status": "ok", "timestamp": 1603191733407, "user_tz": -330, "elapsed": 181250, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
pred_train = Bi_lstm1_network.predict(padded_clean_text_stem)
pred_test = Bi_lstm1_network.predict(padded_clean_text_t_stem)
# + id="rWKVJtN1Mz_d" executionInfo={"status": "ok", "timestamp": 1603191733409, "user_tz": -330, "elapsed": 181233, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
import numpy as np
import matplotlib.pyplot as plt
from itertools import cycle
from sklearn import svm, datasets
from sklearn.metrics import roc_curve, auc
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import label_binarize
from sklearn.multiclass import OneVsRestClassifier
from scipy import interp
def plot_AUC_ROC(y_true, y_pred):
n_classes = 2 #change this value according to class value
# Compute ROC curve and ROC area for each class
fpr = dict()
tpr = dict()
roc_auc = dict()
for i in range(n_classes):
fpr[i], tpr[i], _ = roc_curve(y_true[:, i], y_pred[:, i])
roc_auc[i] = auc(fpr[i], tpr[i])
# Compute micro-average ROC curve and ROC area
fpr["micro"], tpr["micro"], _ = roc_curve(y_true.ravel(), y_pred.ravel())
roc_auc["micro"] = auc(fpr["micro"], tpr["micro"])
############################################################################################
lw = 2
# Compute macro-average ROC curve and ROC area
# First aggregate all false positive rates
all_fpr = np.unique(np.concatenate([fpr[i] for i in range(n_classes)]))
# Then interpolate all ROC curves at this points
mean_tpr = np.zeros_like(all_fpr)
for i in range(n_classes):
mean_tpr += interp(all_fpr, fpr[i], tpr[i])
# Finally average it and compute AUC
mean_tpr /= n_classes
fpr["macro"] = all_fpr
tpr["macro"] = mean_tpr
roc_auc["macro"] = auc(fpr["macro"], tpr["macro"])
# Plot all ROC curves
plt.figure()
plt.plot(fpr["micro"], tpr["micro"],
label='micro-average ROC curve (area = {0:0.2f})'
''.format(roc_auc["micro"]),
color='deeppink', linestyle=':', linewidth=4)
plt.plot(fpr["macro"], tpr["macro"],
label='macro-average ROC curve (area = {0:0.2f})'
''.format(roc_auc["macro"]),
color='navy', linestyle=':', linewidth=4)
colors = cycle(['aqua', 'darkorange'])
#classes_list1 = ["DE","NE","DK"]
classes_list1 = ["Non-duplicate","Duplicate"]
for i, color,c in zip(range(n_classes), colors,classes_list1):
plt.plot(fpr[i], tpr[i], color=color, lw=lw,
label='{0} (AUC = {1:0.2f})'
''.format(c, roc_auc[i]))
plt.plot([0, 1], [0, 1], 'k--', lw=lw)
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic curve')
plt.legend(loc="lower right")
#plt.show()
plt.savefig('Bi_lstm1_HASOC_Eng_Area_RocPlot.pdf',dpi=1000)
files.download('Bi_lstm1_HASOC_Eng_Area_RocPlot.pdf')
# + id="i3tsqxDENNB6" executionInfo={"status": "ok", "timestamp": 1603191733410, "user_tz": -330, "elapsed": 181221, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="338a8925-5e32-4111-e2dc-0bf77c11bfa4" colab={"base_uri": "https://localhost:8080/", "height": 333}
plot_AUC_ROC(label_twoDimension_t,pred_test)
# + id="6boPbARON83n" executionInfo={"status": "ok", "timestamp": 1603191733410, "user_tz": -330, "elapsed": 181209, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
|
BiLSTM Models/With GloVe/BiLSTM1 Covid-19.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data from <NAME>'s GitHub
#
# (See Excel sheet [MatchChart 0.2.0](https://github.com/JeffSackmann/tennis_MatchChartingProject/blob/master/MatchChart%200.2.0.xlsm) for full description)
#
# Each row is one point.
# Volunteers manually input the rally in the '1st' or '2nd' column, depending on whether it's a first or second serve (the other features, like the score, are automatically populated).
#
# Numbers are used to indicate direction and depth, while letters are used to specific shot types (e.g. 'f' stands for 'forehand') and error types ('n' stands for 'net').
#
# A few symbols are used for other purposes, such as types of errors (e.g. '@' means 'unforced error' and '+' indicates an approach shot).
#
# ### Serve
#
# Direction:
#
# + 4 = wide
# + 5 = body
# + 6 = down the T
# + 0 = unknown
#
# Fault types:
#
# + n = net
# + w = wide (in either direction)
# + d = deep
# + x = both wide and deep
# + g = foot faults
# + e = unknown
# + ! = shank
# + V = time violation
# + c = let
#
#
# + \+ = serve-and-volley attempt
#
# EXAMPLES
# + 6 = a serve down the T that lands in the box.
# + 4 = a wide serve that lands in the box
# + 6x = a serve down the T that is both wide and deep.
# + 5d = a body serve that lands deep
# + cc4e = two lets, followed by a wide serve that is out, but you don't know in which direction.
#
# + 4+w = a serve-and-volley attempt on which a wide serve lands deep
#
# ### Serve outcome
#
# + \* = ace
# + \# = unreturnable (the other player touches but cannot return)
# + 'return code' \+ \# = forced error (example: 6f\#)
# + 'return code' \+ @ = unforced error (example: 6f2d@)
#
# ### Rally sequence
#
# Types of shot:
#
#
# + f = forehand groundstroke
# + b = backhand groundstroke
#
# + r = forehand slice
# + s = backhand slice
#
# + v = forehand volley
# + z = backhand volley
#
# + o = standard overhead/smash
# + p = backhand overhead/smash
#
# + u = forehand drop shot
# + y = backhand drop shot
#
# + l = forehand lob
# + m = backhand lob
#
# + h = forehand half-volley
# + i = backhand half-volley
#
# + j = forehand swinging volley
# + k = backhand swinging volley
#
# + t = all trick shots
#
# + q = any unknown shot
#
# Direction (optional):
#
# + 1 = to a right-hander's forehand side / left-hander's backhand side
# + 2 = down the middle of the court
# + 3 = to a right-hander's backhand side / left-hander's forehand side
# + 0 = unknown
#
# ### Rally outcome
#
# + \* = winning shot
# + @ = unforced error
# + \# = forced error
#
# Error types:
# + n = net
# + w = wide
# + d = deep
# + x = wide and deep
# + ! = shank
# + e = unknown
#
# Example:
# 5f2f1f1v2n@ = body serve, forehand down the middle, forehand crosscourt, forehand crosscourt, volley down the middle into the net (unforced error).
#
# ### Serve return depth
#
# + 7 = within the service boxes
# + 8 = behind the service line, but closer to the service line than the baseline
# + 9 = closer to the baseline than the service line.
# + 0 = unknown
#
# ### Court position (optional)
#
# + \+ = approach shots
#
# + \- = at the net
# + = = at the baseline
#
# + ; = hit the net cord
#
# + ^ = stop volleys/drop volley
#
# ## Special situations
#
# + S = point to the server
# + R = point to the returner
# + P = point penalty against the server
# + Q = point penalty against the returner
#
# 'Notes' column : challenges, medical timeouts, rain delays, on-court coaching, time violation warnings...
|
Shot encodings.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
# # 1. Human capital accumulation
# Consider a worker living in **two periods**, $t \in \{1,2\}$.
#
# In each period she decides whether to **work ($l_t = 1$) or not ($l_t = 0$)**.
#
# She can *not* borrow or save and thus **consumes all of her income** in each period.
# If she **works** her **consumption** becomes:
#
# $$c_t = w h_t l_t\,\,\text{if}\,\,l_t=1$$
#
# where $w$ is **the wage rate** and $h_t$ is her **human capital**.
#
# If she does **not work** her consumption becomes:
#
# $$c_t = b\,\,\text{if}\,\,l_t=0$$
#
# where $b$ is the **unemployment benefits**.
# Her **utility of consumption** is:
#
# $$ \frac{c_t^{1-\rho}}{1-\rho} $$
#
# Her **disutility of working** is:
#
# $$ \gamma l_t $$
# From period 1 to period 2, she **accumulates human capital** according to:
#
# $$ h_2 = h_1 + l_1 +
# \begin{cases}
# 0 & \text{with prob. }0.5 \\
# \Delta & \text{with prob. }0.5
# \end{cases} \\
# $$
#
# where $\Delta$ is a **stochastic experience gain**.
# In the **second period** the worker thus solves:
#
# $$
# \begin{eqnarray*}
# v_{2}(h_{2}) & = &\max_{l_{2}} \frac{c_2^{1-\rho}}{1-\rho} - \gamma l_2
# \\ & \text{s.t.} & \\
# c_{2}& = & w h_2 l_2 \\
# l_{2}& \in &\{0,1\}
# \end{eqnarray*}
# $$
# In the **first period** the worker thus solves:
#
# $$
# \begin{eqnarray*}
# v_{1}(h_{1}) &=& \max_{l_{1}} \frac{c_1^{1-\rho}}{1-\rho} - \gamma l_1 + \beta\mathbb{E}_{1}\left[v_2(h_2)\right]
# \\ & \text{s.t.} & \\
# c_1 &=& w h_1 l_1 \\
# h_2 &=& h_1 + l_1 + \begin{cases}
# 0 & \text{with prob. }0.5\\
# \Delta & \text{with prob. }0.5
# \end{cases}\\
# l_{1} &\in& \{0,1\}\\
# \end{eqnarray*}
# $$
#
# where $\beta$ is the **discount factor** and $\mathbb{E}_{1}\left[v_2(h_2)\right]$ is the **expected value of living in period two**.
# The **parameters** of the model are:
rho = 2
beta = 0.96
gamma = 0.1
w = 2
b = 1
Delta = 0.1
# The **relevant levels of human capital** are:
h_vec = np.linspace(0.1,1.5,100)
# **Question 1:** Solve the model in period 2 and illustrate the solution (including labor supply as a function of human capital).
#
# **Question 2:** Solve the model in period 1 and illustrate the solution (including labor supply as a function of human capital).
#
# **Question 3:** Will the worker never work if her potential wage income is lower than the unemployment benefits she can get? Explain and illustrate why or why not.
# # 2. AS-AD model
# Consider the following **AS-AD model**. The **goods market equilibrium** is given by
#
# $$ y_{t} = -\alpha r_{t} + v_{t} $$
#
# where $y_{t}$ is the **output gap**, $r_{t}$ is the **ex ante real interest** and $v_{t}$ is a **demand disturbance**.
# The central bank's **Taylor rule** is
#
# $$ i_{t} = \pi_{t+1}^{e} + h \pi_{t} + b y_{t}$$
#
# where $i_{t}$ is the **nominal interest rate**, $\pi_{t}$ is the **inflation gap**, and $\pi_{t+1}^{e}$ is the **expected inflation gap**.
# The **ex ante real interest rate** is given by
#
# $$ r_{t} = i_{t} - \pi_{t+1}^{e} $$
# Together, the above implies that the **AD-curve** is
#
# $$ \pi_{t} = \frac{1}{h\alpha}\left[v_{t} - (1+b\alpha)y_{t}\right]$$
# Further, assume that the **short-run supply curve (SRAS)** is given by
#
# $$ \pi_{t} = \pi_{t}^{e} + \gamma y_{t} + s_{t}$$
#
# where $s_t$ is a **supply disturbance**.
# **Inflation expectations are adaptive** and given by
#
# $$ \pi_{t}^{e} = \phi\pi_{t-1}^{e} + (1-\phi)\pi_{t-1}$$
# Together, this implies that the **SRAS-curve** can also be written as
#
# $$ \pi_{t} = \pi_{t-1} + \gamma y_{t} - \phi\gamma y_{t-1} + s_{t} - \phi s_{t-1} $$
# The **parameters** of the model are:
# +
par = {}
par['alpha'] = 5.76
par['h'] = 0.5
par['b'] = 0.5
par['phi'] = 0
par['gamma'] = 0.075
# -
# **Question 1:** Use the ``sympy`` module to solve for the equilibrium values of output, $y_t$, and inflation, $\pi_t$, (where AD = SRAS) given the parameters ($\alpha$, $h$, $b$, $\alpha$, $\gamma$) and $y_{t-1}$ , $\pi_{t-1}$, $v_t$, $s_t$, and $s_{t-1}$.
#
# **Question 2:** Find and illustrate the equilibrium when $y_{t-1} = \pi_{t-1} = v_t = s_t = s_{t-1} = 0$. Illustrate how the equilibrium changes when instead $v_t = 0.1$.
# **Persistent disturbances:** Now, additionaly, assume that both the demand and the supply disturbances are AR(1) processes
#
# $$ v_{t} = \delta v_{t-1} + x_{t} $$
# $$ s_{t} = \omega s_{t-1} + c_{t} $$
#
# where $x_{t}$ is a **demand shock**, and $c_t$ is a **supply shock**. The **autoregressive parameters** are:
par['delta'] = 0.80
par['omega'] = 0.15
# **Question 3:** Starting from $y_{-1} = \pi_{-1} = s_{-1} = 0$, how does the economy evolve for $x_0 = 0.1$, $x_t = 0, \forall t > 0$ and $c_t = 0, \forall t \geq 0$?
# **Stochastic shocks:** Now, additionally, assume that $x_t$ and $c_t$ are stochastic and normally distributed
#
# $$ x_{t}\sim\mathcal{N}(0,\sigma_{x}^{2}) $$
# $$ c_{t}\sim\mathcal{N}(0,\sigma_{c}^{2}) $$
#
# The **standard deviations of the shocks** are:
par['sigma_x'] = 3.492
par['sigma_c'] = 0.2
# **Question 4:** Simulate the AS-AD model for 1,000 periods. Calculate the following five statistics:
#
# 1. Variance of $y_t$, $var(y_t)$
# 2. Variance of $\pi_t$, $var(\pi_t)$
# 3. Correlation between $y_t$ and $\pi_t$, $corr(y_t,\pi_t)$
# 4. Auto-correlation between $y_t$ and $y_{t-1}$, $corr(y_t,y_{t-1})$
# 5. Auto-correlation between $\pi_t$ and $\pi_{t-1}$, $corr(\pi_t,\pi_{t-1})$
# **Question 5:** Plot how the correlation between $y_t$ and $\pi_t$ changes with $\phi$. Use a numerical optimizer or root finder to choose $\phi\in(0,1)$ such that the simulated correlation between $y_t$ and $\pi_t$ comes close to 0.31.
# **Quesiton 6:** Use a numerical optimizer to choose $\sigma_x>0$, $\sigma_c>0$ and $\phi\in(0,1)$ to make the simulated statistics as close as possible to US business cycle data where:
#
# 1. $var(y_t) = 1.64$
# 2. $var(\pi_t) = 0.21$
# 3. $corr(y_t,\pi_t) = 0.31$
# 4. $corr(y_t,y_{t-1}) = 0.84$
# 5. $corr(\pi_t,\pi_{t-1}) = 0.48$
# # 3. Exchange economy
# Consider an **exchange economy** with
#
# 1. 3 goods, $(x_1,x_2,x_3)$
# 2. $N$ consumers indexed by \\( j \in \{1,2,\dots,N\} \\)
# 3. Preferences are Cobb-Douglas with log-normally distributed coefficients
#
# $$ \begin{eqnarray*}
# u^{j}(x_{1},x_{2},x_{3}) &=&
# \left(x_{1}^{\beta_{1}^{j}}x_{2}^{\beta_{2}^{j}}x_{3}^{\beta_{3}^{j}}\right)^{\gamma}\\
# & & \,\,\,\beta_{i}^{j}=\frac{\alpha_{i}^{j}}{\alpha_{1}^{j}+\alpha_{2}^{j}+\alpha_{3}^{j}} \\
# & & \,\,\,\boldsymbol{\alpha}^{j}=(\alpha_{1}^{j},\alpha_{2}^{j},\alpha_{3}^{j}) \\
# & & \,\,\,\log(\boldsymbol{\alpha}^j) \sim \mathcal{N}(\mu,\Sigma) \\
# \end{eqnarray*} $$
#
# 4. Endowments are exponentially distributed,
#
# $$
# \begin{eqnarray*}
# \boldsymbol{e}^{j} &=& (e_{1}^{j},e_{2}^{j},e_{3}^{j}) \\
# & & e_i^j \sim f, f(z;\zeta) = 1/\zeta \exp(-z/\zeta)
# \end{eqnarray*}
# $$
# Let $p_3 = 1$ be the **numeraire**. The implied **demand functions** are:
#
# $$
# \begin{eqnarray*}
# x_{i}^{\star j}(p_{1},p_{2},\boldsymbol{e}^{j})&=&\beta^{j}_i\frac{I^j}{p_{i}} \\
# \end{eqnarray*}
# $$
#
# where consumer $j$'s income is
#
# $$I^j = p_1 e_1^j + p_2 e_2^j +p_3 e_3^j$$
# The **parameters** and **random preferences and endowments** are given by:
# +
# a. parameters
N = 50000
mu = np.array([3,2,1])
Sigma = np.array([[0.25, 0, 0], [0, 0.25, 0], [0, 0, 0.25]])
gamma = 0.8
zeta = 1
# b. random draws
seed = 1986
np.random.seed(seed)
# preferences
alphas = np.exp(np.random.multivariate_normal(mu, Sigma, size=N))
betas = alphas/np.reshape(np.sum(alphas,axis=1),(N,1))
# endowments
e1 = np.random.exponential(zeta,size=N)
e2 = np.random.exponential(zeta,size=N)
e3 = np.random.exponential(zeta,size=N)
# -
# **Question 1:** Plot the histograms of the budget shares for each good across agents.
#
# Consider the **excess demand functions:**
#
# $$ z_i(p_1,p_2) = \sum_{j=1}^N x_{i}^{\star j}(p_{1},p_{2},\boldsymbol{e}^{j}) - e_i^j$$
#
# **Question 2:** Plot the excess demand functions.
# **Quesiton 3:** Find the Walras-equilibrium prices, $(p_1,p_2)$, where both excess demands are (approximately) zero, e.g. by using the following tâtonnement process:
#
# 1. Guess on $p_1 > 0$, $p_2 > 0$ and choose tolerance $\epsilon > 0$ and adjustment aggressivity parameter, $\kappa > 0$.
# 2. Calculate $z_1(p_1,p_2)$ and $z_2(p_1,p_2)$.
# 3. If $|z_1| < \epsilon$ and $|z_2| < \epsilon$ then stop.
# 4. Else set $p_1 = p_1 + \kappa \frac{z_1}{N}$ and $p_2 = p_2 + \kappa \frac{z_2}{N}$ and return to step 2.
# **Question 4:** Plot the distribution of utility in the Walras-equilibrium and calculate its mean and variance.
# **Question 5:** Find the Walras-equilibrium prices if instead all endowments were distributed equally. Discuss the implied changes in the distribution of utility. Does the value of $\gamma$ play a role for your conclusions?
|
examproject/exam_2019.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="9Xp5YoON6uU4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="3ff200a3-b2e2-4e36-bab0-cb37e03992ed"
from keras.preprocessing.image import load_img, img_to_array, save_img
target_image_path = '/content/drive/My Drive/Colab Notebooks/4.jpeg'
style_reference_image_path = '/content/drive/My Drive/Colab Notebooks/3.jpeg'
# 생성된 사진의 차원
width, height = load_img(target_image_path).size
img_height = 400
img_width = int(width * img_height / height)
# + id="zL0SY434-Xh9" colab_type="code" colab={}
import numpy as np
from keras.applications import vgg19
def preprocess_image(image_path):
img = load_img(image_path, target_size=(img_height, img_width))
img = img_to_array(img)
img = np.expand_dims(img, axis=0)
img = vgg19.preprocess_input(img)
return img
def deprocess_image(x):
# ImageNet의 평균 픽셀 값을 더합니다
x[:, :, 0] += 103.939
x[:, :, 1] += 116.779
x[:, :, 2] += 123.68
# 'BGR'->'RGB'
x = x[:, :, ::-1]
x = np.clip(x, 0, 255).astype('uint8')
return x
# + id="49AhoHINDHDM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="82596b4d-5d06-4a01-8661-966a1f5e006e"
from keras import backend as K
target_image = K.constant(preprocess_image(target_image_path))
style_reference_image = K.constant(preprocess_image(style_reference_image_path))
# 생성된 이미지를 담을 플레이스홀더
combination_image = K.placeholder((1, img_height, img_width, 3))
# 세 개의 이미지를 하나의 배치로 합칩니다
input_tensor = K.concatenate([target_image,
style_reference_image,
combination_image], axis=0)
# 세 이미지의 배치를 입력으로 받는 VGG 네트워크를 만듭니다.
# 이 모델은 사전 훈련된 ImageNet 가중치를 로드합니다
model = vgg19.VGG19(input_tensor=input_tensor,
weights='imagenet',
include_top=False)
print('모델 로드 완료.')
# + id="wjJ4TuiHC-VV" colab_type="code" colab={}
def content_loss(base, combination):
return K.sum(K.square(combination - base))
# + id="pNohT4lSFipx" colab_type="code" colab={}
def gram_matrix(x):
features = K.batch_flatten(K.permute_dimensions(x, (2, 0, 1)))
gram = K.dot(features, K.transpose(features))
return gram
def style_loss(style, combination):
S = gram_matrix(style)
C = gram_matrix(combination)
channels = 3
size = img_height * img_width
return K.sum(K.square(S - C)) / (4. * (channels ** 2) * (size ** 2))
# + id="DXY3fhNUGAcT" colab_type="code" colab={}
def total_variation_loss(x):
a = K.square(
x[:, :img_height - 1, :img_width - 1, :] - x[:, 1:, :img_width - 1, :])
b = K.square(
x[:, :img_height - 1, :img_width - 1, :] - x[:, :img_height - 1, 1:, :])
return K.sum(K.pow(a + b, 1.25))
# + id="Adi4OOVMGcf5" colab_type="code" colab={}
# 층 이름과 활성화 텐서를 매핑한 딕셔너리
outputs_dict = dict([(layer.name, layer.output) for layer in model.layers])
# 콘텐츠 손실에 사용할 층
content_layer = 'block5_conv2'
# 스타일 손실에 사용할 층
style_layers = ['block1_conv1',
'block2_conv1',
'block3_conv1',
'block4_conv1',
'block5_conv1']
# 손실 항목의 가중치 평균에 사용할 가중치
total_variation_weight = 1e-4
style_weight = 1.
content_weight = 0.025
# 모든 손실 요소를 더해 하나의 스칼라 변수로 손실을 정의합니다
loss = K.variable(0.)
layer_features = outputs_dict[content_layer]
target_image_features = layer_features[0, :, :, :]
combination_features = layer_features[2, :, :, :]
loss = loss+ content_weight * content_loss(target_image_features,
combination_features)
for layer_name in style_layers:
layer_features = outputs_dict[layer_name]
style_reference_features = layer_features[1, :, :, :]
combination_features = layer_features[2, :, :, :]
sl = style_loss(style_reference_features, combination_features)
loss = loss+ (style_weight / len(style_layers)) * sl
loss = loss+ total_variation_weight * total_variation_loss(combination_image)
# + id="R31kj71CI-MO" colab_type="code" colab={}
# 손실에 대한 생성된 이미지의 그래디언트를 구합니다
grads = K.gradients(loss, combination_image)[0]
# 현재 손실과 그래디언트의 값을 추출하는 케라스 Function 객체입니다
fetch_loss_and_grads = K.function([combination_image], [loss, grads])
class Evaluator(object):
def __init__(self):
self.loss_value = None
self.grads_values = None
def loss(self, x):
assert self.loss_value is None
x = x.reshape((1, img_height, img_width, 3))
outs = fetch_loss_and_grads([x])
loss_value = outs[0]
grad_values = outs[1].flatten().astype('float64')
self.loss_value = loss_value
self.grad_values = grad_values
return self.loss_value
def grads(self, x):
assert self.loss_value is not None
grad_values = np.copy(self.grad_values)
self.loss_value = None
self.grad_values = None
return grad_values
evaluator = Evaluator()
# + id="q9oDLG1nNfKE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 102} outputId="f724a892-50e6-4380-dd1e-17741a93fe62"
from scipy.optimize import fmin_l_bfgs_b
import time
result_prefix = 'style_transfer_result'
iterations = 20
x = preprocess_image(target_image_path)
x = x.flatten()
for i in range(iterations):
print("loop ",i)
start_time = time.time()
x, min_val, info = fmin_l_bfgs_b(evaluator.loss, x, fprime=evaluator.grads, maxfun=20)
print('now loss :', min_val)
img = x.copy().reshape((img_height, img_width, 3))
img = deprocess_image(img)
fname = result_prefix + '_at_iteration_%d.png' % i
save_img(fname, img)
print('save image :', fname)
end_time =time.time()
print(i, ' 번째 반복 완료 : ', end_time-start_time)
# + id="uaAUSm7KOfNg" colab_type="code" colab={}
|
8/8-3-1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.4 64-bit
# name: python374jvsc74a57bd07945e9a82d7512fbf96246d9bbc29cd2f106c1a4a9cf54c9563dadf10f2237d4
# ---
# # 04 - Grouping
# ### Step 1. Import the necessary libraries
import pandas as pd
# ### Step 2. Create the DataFrame with the following values:
raw_data = {'regiment': ['Nighthawks', 'Nighthawks', 'Nighthawks', 'Nighthawks', 'Dragoons', 'Dragoons', 'Dragoons', 'Dragoons', 'Scouts', 'Scouts', 'Scouts', 'Scouts'],
'company': ['1st', '1st', '2nd', '2nd', '1st', '1st', '2nd', '2nd','1st', '1st', '2nd', '2nd'],
'name': ['Miller', 'Jacobson', 'Ali', 'Milner', 'Cooze', 'Jacon', 'Ryaner', 'Sone', 'Sloan', 'Piger', 'Riani', 'Ali'],
'preTestScore': [4, 24, 31, 2, 3, 4, 24, 31, 2, 3, 2, 3],
'postTestScore': [25, 94, 57, 62, 70, 25, 94, 57, 62, 70, 62, 70]}
# ### Step 3. Assign it to a variable called regiment.
regimen=pd.DataFrame(raw_data)
regimen
# ### Step 4. What is the mean preTestScore from the regiment Nighthawks?
Nighthawks=regimen.groupby('regiment').mean()
Nighthawks.iloc[1,0]
Nighthawks.loc["Nighthawks"][0]
regimen[regimen["regiment"]=="Nighthawks"].groupby("regiment").mean()
#
# ### Step 5. Present general statistics by company
regimen.groupby('regiment').describe()
# ### Step 6. What is the mean of each company's preTestScore?
# + jupyter={"outputs_hidden": false}
regimen.groupby('regiment').mean()
# -
# ### Step 7. Present the mean preTestScores grouped by regiment and company
# + jupyter={"outputs_hidden": false}
regimen.groupby(['regiment',"company"]).mean()
# -
# ### Step 8. Present the mean preTestScores grouped by regiment and company without heirarchical indexing
pretestscore=regimen.groupby(['regiment',"company"]).mean().unstack()
pretestscore
# ### Step 9. Group the entire dataframe by regiment and company
regimen.groupby(["regiment","company"]).count().unstack()
# ### Step 10. What is the number of observations in each regiment and company
regimen.groupby(["regiment","company"]).count()
# ### Step 11. Iterate over a group and print the name and the whole data from the regiment
for x,y in regimen.groupby("regiment"):
print(x)
print("---")
print(y)
print("++++")
|
week4_EDA_np_pd_json_apis_regex/day5_matplotlib_I_api/exercises/nuevo_panda/Regiment.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/npgeorge/DS-Unit-2-Applied-Modeling/blob/master/Nick_George_Assignment_1_Applied_Modeling.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="nCc3XZEyG3XV"
# Lambda School Data Science
#
# *Unit 2, Sprint 3, Module 1*
#
# ---
#
#
# # Define ML problems
#
# You will use your portfolio project dataset for all assignments this sprint.
#
# ## Assignment
#
# Complete these tasks for your project, and document your decisions.
#
# - [ ] Choose your target. Which column in your tabular dataset will you predict?
# - [ ] Is your problem regression or classification?
# - [ ] How is your target distributed?
# - Classification: How many classes? Are the classes imbalanced?
# - Regression: Is the target right-skewed? If so, you may want to log transform the target.
# - [ ] Choose which observations you will use to train, validate, and test your model.
# - Are some observations outliers? Will you exclude them?
# - Will you do a random split or a time-based split?
# - [ ] Choose your evaluation metric(s).
# - Classification: Is your majority class frequency > 50% and < 70% ? If so, you can just use accuracy if you want. Outside that range, accuracy could be misleading. What evaluation metric will you choose, in addition to or instead of accuracy?
# - [ ] Begin to clean and explore your data.
# - [ ] Begin to choose which features, if any, to exclude. Would some features "leak" future information?
# + id="sB5a_oDsDemj" colab_type="code" colab={}
#use first principles to figure out end data frame
#what are the foundational differences, keep it small, build from there.
#due to the many data frames, my goal is to build one comprehensive data frame
#this one data frame will be used to try to predict who will win each game.
#target
#'outcome' column from the game csv
#regression
#win, loss, or tie will numerically represented
#target distribution
#need to explore
#I will train/test/val on each season separately.
#If this doesn't work, I will try multiple seasons at a time.
#Perhaps a 'returning_players' parameter giving a % of how many players stay YoY could be useful
# + id="yklRxcFUQn-0" colab_type="code" colab={}
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
#upload local file
from google.colab import files
import io
# + id="JGzaklYcBqcv" colab_type="code" colab={}
#choose file button
uploaded = files.upload()
# + id="-C-ejDciBo75" colab_type="code" colab={}
#uploading the game df to start exploration
df = pd.read_csv(io.BytesIO(uploaded['game.csv']))
# + id="43PUS3_LGcJC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 351} outputId="15536973-a29e-40f3-b7a9-2f8f75484567"
df.head()
# + id="T2UF-9RINr-C" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 367} outputId="cffd45b3-d000-4451-d903-f0fec7cb07cc"
#bruins
#if a model works for one team, it *could* work broadly for others
#this may turn out to be a bad assumption
#this filter returns the df for the Bruins
#or statement for home and away games
bruins_id_condition = (df['home_team_id'] == 6) | (df['away_team_id'] == 6)
df_bruins = df[bruins_id_condition]
df_bruins.head()
# + id="VCKfVNZ0VvDa" colab_type="code" colab={}
#seasons
#the NHL seasons runs from early October to Early April, Playoffs through mid June
#there are 6 seasons in this data set
#i'll split the set by seasons
#as some years are very different than others for certain teams
#in case seasons started early one year
#2010 to 2011 season
season_10to11 = (df['date_time'] > '2010-09-01') & (df['date_time'] < '2011-07-01')
#2011 to 2012
season_11to12 = (df['date_time'] > '2011-09-01') & (df['date_time'] < '2012-07-01')
#2012 to 2013
season_12to13 = (df['date_time'] > '2012-09-01') & (df['date_time'] < '2013-07-01')
#2013 to 2014
season_13to14 = (df['date_time'] > '2013-09-01') & (df['date_time'] < '2014-07-01')
#2014 to 2015
season_14to15 = (df['date_time'] > '2014-09-01') & (df['date_time'] < '2015-07-01')
#2015 to 2016
season_15to16 = (df['date_time'] > '2015-09-01') & (df['date_time'] < '2016-07-01')
#2016 to 2017
season_16to17 = (df['date_time'] > '2016-09-01') & (df['date_time'] < '2017-07-01')
#2017 to 2018
season_17to18 = (df['date_time'] > '2017-09-01') & (df['date_time'] < '2018-07-01')
#2018 to 2019
season_18to19 = (df['date_time'] > '2018-09-01') & (df['date_time'] < '2019-07-01')
#need to try to implement a function here
# + id="P8Ue6oyioinz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 626} outputId="c0caaa3c-4ff0-4caa-942c-54292b3f3624"
#pass the season condition, sort season from beginning to end
df_10to11 = df[season_10to11].sort_values(by=['date_time'])
df_10to11
# + id="SvyC6cL2e9_6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 301} outputId="7a2cce2d-42e2-4213-d5b2-a1a68a5591b5"
#bruins 2010 to 2011, won the Stanley Cup this year
bruins_id_condition = (df_10to11['home_team_id'] == 6) | (df_10to11['away_team_id'] == 6)
df_bruins_10to11 = df_10to11[bruins_id_condition]
df_bruins_10to11.tail() #last game won in April - Stanley Cup Win
|
Nick_George_Assignment_1_Applied_Modeling.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] deletable=true editable=true
# <h1> Getting started with TensorFlow </h1>
#
# In this notebook, you play around with the TensorFlow Python API.
# + deletable=true editable=true
import tensorflow as tf
import numpy as np
print(tf.__version__)
# + [markdown] deletable=true editable=true
# <h2> Adding two tensors </h2>
#
# First, let's try doing this using numpy, the Python numeric package. numpy code is immediately evaluated.
# + deletable=true editable=true
a = np.array([5, 3, 8])
b = np.array([3, -1, 2])
c = np.add(a, b)
print(c)
# + [markdown] deletable=true editable=true
# The equivalent code in TensorFlow consists of two steps:
# <p>
# <h3> Step 1: Build the graph </h3>
# + deletable=true editable=true
a = tf.constant([5, 3, 8])
b = tf.constant([3, -1, 2])
c = tf.add(a, b)
print(c)
# + [markdown] deletable=true editable=true
# c is an Op ("Add") that returns a tensor of shape (3,) and holds int32. The shape is inferred from the computation graph.
#
# Try the following in the cell above:
# <ol>
# <li> Change the 5 to 5.0, and similarly the other five numbers. What happens when you run this cell? </li>
# <li> Add an extra number to a, but leave b at the original (3,) shape. What happens when you run this cell? </li>
# <li> Change the code back to a version that works </li>
# </ol>
#
# <p/>
# <h3> Step 2: Run the graph
# + deletable=true editable=true
with tf.Session() as sess:
result = sess.run(c)
print(result)
# + [markdown] deletable=true editable=true
# <h2> Using a feed_dict </h2>
#
# Same graph, but without hardcoding inputs at build stage
# + deletable=true editable=true
a = tf.placeholder(dtype=tf.int32, shape=(None,)) # batchsize x scalar
b = tf.placeholder(dtype=tf.int32, shape=(None,))
c = tf.add(a, b)
with tf.Session() as sess:
result = sess.run(c, feed_dict={
a: [3, 4, 5],
b: [-1, 2, 3]
})
print(result)
# + [markdown] deletable=true editable=true
# <h2> Heron's Formula in TensorFlow </h2>
#
# The area of triangle whose three sides are $(a, b, c)$ is $\sqrt{s(s-a)(s-b)(s-c)}$ where $s=\frac{a+b+c}{2}$
#
# Look up the available operations at https://www.tensorflow.org/api_docs/python/tf
# + deletable=true editable=true
def compute_area(sides):
# slice the input to get the sides
a = sides[:,0] # 5.0, 2.3
b = sides[:,1] # 3.0, 4.1
c = sides[:,2] # 7.1, 4.8
# Heron's formula
s = (a + b + c) * 0.5 # (a + b) is a short-cut to tf.add(a, b)
areasq = s * (s - a) * (s - b) * (s - c) # (a * b) is a short-cut to tf.multiply(a, b), not tf.matmul(a, b)
return tf.sqrt(areasq)
with tf.Session() as sess:
# pass in two triangles
area = compute_area(tf.constant([
[5.0, 3.0, 7.1],
[2.3, 4.1, 4.8]
]))
result = sess.run(area)
print(result)
# + [markdown] deletable=true editable=true
# <h2> Placeholder and feed_dict </h2>
#
# More common is to define the input to a program as a placeholder and then to feed in the inputs. The difference between the code below and the code above is whether the "area" graph is coded up with the input values or whether the "area" graph is coded up with a placeholder through which inputs will be passed in at run-time.
# + deletable=true editable=true
with tf.Session() as sess:
sides = tf.placeholder(tf.float32, shape=(None, 3)) # batchsize number of triangles, 3 sides
area = compute_area(sides)
result = sess.run(area, feed_dict = {
sides: [
[5.0, 3.0, 7.1],
[2.3, 4.1, 4.8]
]
})
print(result)
# + [markdown] deletable=true editable=true
# ## tf.eager
#
# tf.eager allows you to avoid the build-then-run stages. However, most production code will follow the lazy evaluation paradigm because the lazy evaluation paradigm is what allows for multi-device support and distribution.
# <p>
# One thing you could do is to develop using tf.eager and then comment out the eager execution and add in the session management code.
#
# <b> You may need to click on Reset Session to try this out </b>
# + deletable=true editable=true
import tensorflow as tf
tf.enable_eager_execution()
def compute_area(sides):
# slice the input to get the sides
a = sides[:,0] # 5.0, 2.3
b = sides[:,1] # 3.0, 4.1
c = sides[:,2] # 7.1, 4.8
# Heron's formula
s = (a + b + c) * 0.5 # (a + b) is a short-cut to tf.add(a, b)
areasq = s * (s - a) * (s - b) * (s - c) # (a * b) is a short-cut to tf.multiply(a, b), not tf.matmul(a, b)
return tf.sqrt(areasq)
area = compute_area(tf.constant([
[5.0, 3.0, 7.1],
[2.3, 4.1, 4.8]
]))
print(area)
# + [markdown] deletable=true editable=true
# ## Challenge Exercise
#
# Use TensorFlow to find the roots of a fourth-degree polynomial using [Halley's Method](https://en.wikipedia.org/wiki/Halley%27s_method). The five coefficients (i.e. $a_0$ to $a_4$) of
# <p>
# $f(x) = a_0 + a_1 x + a_2 x^2 + a_3 x^3 + a_4 x^4$
# <p>
# will be fed into the program, as will the initial guess $x_0$. Your program will start from that initial guess and then iterate one step using the formula:
# <img src="https://wikimedia.org/api/rest_v1/media/math/render/svg/142614c0378a1d61cb623c1352bf85b6b7bc4397" />
# <p>
# If you got the above easily, try iterating indefinitely until the change between $x_n$ and $x_{n+1}$ is less than some specified tolerance. Hint: Use [tf.while_loop](https://www.tensorflow.org/api_docs/python/tf/while_loop)
# + [markdown] deletable=true editable=true
# Copyright 2017 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License
|
courses/machine_learning/deepdive/03_tensorflow/a_tfstart.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] run_control={"frozen": false, "read_only": false}
# Please go through the "building_strategies" notebook before looking at this notebook.
#
# Lets work on optimizing our strategy parameters for the Bollinger Band© we previously built. We will include a function from the examples folder called build_example_strategy which builds this strategy.
# + run_control={"frozen": false, "read_only": false}
import math
from types import SimpleNamespace
import pandas as pd
import numpy as np
import pyqstrat as pq
from pyqstrat.evaluator import compute_sharpe, compute_sortino, compute_maxdd_pct, compute_amean, compute_rolling_dd
from pyqstrat.evaluator import compute_periods_per_year
from pyqstrat.notebooks.support.build_example_strategy import build_example_strategy
pq.ContractGroup.clear()
# -
# Lets try to optimize the the lookback period for the moving average and the number of standard deviations for the bands.
#
# We will try to optimize the sharpe ratio but also look at the sortino and max drawdown as we optimize the sharpe.
#
# To do this, we have to write a generator function and a cost function. The generator produces all the combinations of parameters you want to optimize. The cost function will run the strategy for each parameter combination provided by the generator and return whatever metric you want to optimize, as well as any other metrics you want to see at the same time.
#
# In this case, our cost metric will be the sharpe ratio of the strategy but we will also look at sortino and drawdowns at the same time. We also look at the number of trades generate and ignore results as unreliable if the number of trades is less than 10.
#
# The optimizer uses multiple processes to run as fast as possible. You can set the number of processes you want to use using the max_processes argument to the Optimizer constructor. If you don't set this, the optimizer will the same number of processes as the CPU cores on your machine. You may want to increase this number if your backtesting is I/O bound, and the CPU cores are often idle waiting for disk or other resources.
#
# In this case, we are optimizing 2 parameters at the same time, but we can optimize 1 parameter or more than 2 as well.
# + run_control={"frozen": false, "read_only": false}
def generator():
for lookback_period in np.arange(5, 50, 10):
for num_std in np.arange(1, 5, 0.5): # number of standard deviations the bands are away from the SMA
costs = (yield {'lookback_period' : lookback_period, 'num_std' : num_std})
yield
def cost_func(suggestion):
strategy_context = SimpleNamespace(lookback_period = suggestion['lookback_period'], num_std = suggestion['num_std'])
strategy = build_example_strategy(strategy_context)
strategy.run()
returns_df = strategy.df_returns().set_index('timestamp')
num_trades = len(strategy.df_trades())
if num_trades < 10: return np.nan, {}
returns = returns_df.ret.values
equity = returns_df.equity.values
dates = returns_df.index.values
periods_per_year = compute_periods_per_year(dates)
amean = compute_amean(returns, periods_per_year)
sharpe = compute_sharpe(returns, amean, periods_per_year)
sortino = compute_sortino(returns, amean, periods_per_year)
rolling_dd = compute_rolling_dd(dates, equity)
maxdd = compute_maxdd_pct(rolling_dd[1])
return sharpe, {'sortino' : sortino, 'maxdd' : maxdd, 'num_trades' : num_trades}
optimizer = pq.Optimizer('example', generator(), cost_func, max_processes = 8)
optimizer.run(raise_on_error = True)
optimizer.plot_3d(x = 'lookback_period', y = 'num_std', hspace = 0.5);
# -
# There seems to be a region around a lookback period of 35 bars and 3 standard deviations that has a positive sharpe and sortino. The actual points you provided to the plot are shown with red X markers.
#
# Let's build a contour plot to see the same data in a different view.
# + run_control={"frozen": false, "read_only": false}
optimizer.plot_3d(x = 'lookback_period', y = 'num_std', plot_type = 'contour', hspace = 0.5, );
# -
# Lets look at the actual values of the sharpes, sortinos and drawdowns
# + run_control={"frozen": false, "read_only": false}
optimizer.df_experiments(ascending = False)
# + run_control={"frozen": false, "read_only": false}
optimizer.df_experiments(sort_column = 'maxdd', ascending = False)
# -
# Let's run the strategy at a lookback period of 35 bars with 3 standard deviations to verify the results
# + run_control={"frozen": false, "read_only": false}
strategy = build_example_strategy(SimpleNamespace(lookback_period = 35, num_std = 3))
strategy.run()
strategy.evaluate_returns();
# -
# This confirms that lookback period somewhere around 35 bars with 3 standard deviations gives us a positive sharpe and sortino and lower drawdowns. This is a toy example, and in real life you would want to run with a lot more data, look at the stability of this region and do other out of sample testing before you can use this strategy with real money.
optimizer.df_experiments()
|
pyqstrat/notebooks/optimizing_strategies.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Linear Regression in eager mode
import time
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
sess_config = tf.ConfigProto(gpu_options=tf.GPUOptions(allow_growth=True))
tf.enable_eager_execution()
def read_birth_life_data(filename):
"""Read in birth_life_2010.txt and return:
data in the form of NumPy array
n_samples: number of samples
"""
text = open(filename, 'r').readlines()[1:]
data = [line[:-1].split('\t') for line in text]
births = [float(line[1]) for line in data]
lifes = [float(line[2]) for line in data]
data = list(zip(births, lifes))
n_samples = len(data)
data = np.asarray(data, dtype=np.float32)
return data, n_samples
# +
DATA_FILE = '../datasets/birth_life_2010.txt'
data, n_samples = read_birth_life_data(DATA_FILE)
print("data.shape: {}\n".format(data.shape))
print(data)
# -
# ### `tf.data` 사용하여 dataset 만들기
dataset = tf.data.Dataset.from_tensor_slices((data[:, 0], data[:, 1]))
print(dataset)
import tensorflow.contrib.eager as tfe
# ## Phase1 : Build a graph
# +
# Create variables.
w = tfe.Variable(0.0)
b = tfe.Variable(0.0)
# Define the linear predictor.
def prediction(x):
return x * w + b
# Define loss functions of the form: L(y, y_predicted)
def squared_loss(y, y_predicted):
return (y - y_predicted) ** 2
def huber_loss(y, y_predicted, m=1.0):
"""Huber loss."""
t = y - y_predicted
# Note that enabling eager execution lets you use Python control flow and
# specificy dynamic TensorFlow computations. Contrast this implementation
# to the graph-construction one found in `utils`, which uses `tf.cond`.
return t ** 2 if tf.abs(t) <= m else m * (2 * tf.abs(t) - m)
# -
# ## Phase 2 : Training a model
def train(loss_fn):
"""Train a regression model evaluated using `loss_fn`."""
print('Training; loss function: ' + loss_fn.__name__)
optimizer = tf.train.GradientDescentOptimizer(learning_rate=0.01)
# Define the function through which to differentiate.
def loss_for_example(x, y):
return loss_fn(y, prediction(x))
# `grad_fn(x_i, y_i)` returns (1) the value of `loss_for_example`
# evaluated at `x_i`, `y_i` and (2) the gradients of any variables used in
# calculating it.
grad_fn = tfe.implicit_value_and_gradients(loss_for_example)
start = time.time()
for epoch in range(100):
total_loss = 0.0
for x_i, y_i in tfe.Iterator(dataset):
loss, gradients = grad_fn(x_i, y_i)
# Take an optimization step and update variables.
optimizer.apply_gradients(gradients)
total_loss += loss
if epoch % 10 == 0:
print('Epoch {0}: {1}'.format(epoch, total_loss / n_samples))
print('Took: %f seconds' % (time.time() - start))
print('Eager execution exhibits significant overhead per operation. '
'As you increase your batch size, the impact of the overhead will '
'become less noticeable. Eager execution is under active development: '
'expect performance to increase substantially in the near future!')
train(huber_loss)
# ### Plot the result
plt.plot(data[:,0], data[:,1], 'bo')
# The `.numpy()` method of a tensor retrieves the NumPy array backing it.
# In future versions of eager, you won't need to call `.numpy()` and will
# instead be able to, in most cases, pass Tensors wherever NumPy arrays are
# expected.
plt.plot(data[:,0], data[:,0] * w.numpy() + b.numpy(), 'r',
label="huber regression")
plt.legend()
plt.show()
|
03.regression/01-2.linear.regression.eager.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Analysis Tools
# # Assignment: Testing a Potential Moderator
#
# Following is the Python program I wrote to fulfill the last assignment of the [Data Analysis Tools online course](https://www.coursera.org/learn/data-analysis-tools/home/welcome).
#
# I used [Jupyter Notebook](http://nbviewer.jupyter.org/github/ipython/ipython/blob/3.x/examples/Notebook/Index.ipynb) as it is a pretty way to write code and present results.
#
# ## Research question for this assignment
#
# Using the [Gapminder database](http://www.gapminder.org/), I found a significant [correlation](PearsonCorrelation.ipynb) between the income per person (the explanatory variable) and the residential electricity consumption (the response variable). For this exercice, I would like to see if the urban rate is a potential moderator.
#
# ## Data management
#
# For the question I'm interested in, the countries for which data are missing will be discarded. As missing data in Gapminder database are replace directly by `NaN` no special data treatment is needed.
# + hide_input=false
# Magic command to insert the graph directly in the notebook
# %matplotlib inline
# Load a useful Python libraries for handling data
import pandas as pd
import numpy as np
import statsmodels.formula.api as smf
import scipy.stats as stats
import seaborn as sns
import matplotlib.pyplot as plt
from IPython.display import Markdown, display
# -
# Read the data
data_filename = r'gapminder.csv'
data = pd.read_csv(data_filename, low_memory=False)
data = data.set_index('country')
# General information on the Gapminder data
# + variables={"len(data)": "<p><strong>NameError</strong>: name 'data' is not defined</p>\n", "len(data.columns)": "<p><strong>NameError</strong>: name 'data' is not defined</p>\n"}
display(Markdown("Number of countries: {}".format(len(data))))
display(Markdown("Number of variables: {}".format(len(data.columns))))
# -
subdata2 = (data[['incomeperperson', 'urbanrate', 'relectricperperson']]
.assign(income=lambda x: pd.to_numeric(x['incomeperperson'], errors='coerce'),
urbanrate=lambda x: pd.to_numeric(x['urbanrate'], errors='coerce'),
electricity=lambda x: pd.to_numeric(x['relectricperperson'], errors='coerce'))
.dropna())
# ## Data analysis
sns.distplot(subdata2.income)
plt.xlabel("Income per person (constant 2000 US$)")
_ = plt.title("Distribution of the income per person")
sns.distplot(subdata2.electricity)
plt.xlabel("Residential electricity consumption (kWh)")
_ = plt.title("Distribution of the residential electricity consumption")
sns.distplot(subdata2.urbanrate)
plt.xlabel("Urban rate (%)")
_ = plt.title("Urban rate distribution")
# ## Correlation test
sns.regplot(x='income', y='electricity', data=subdata2)
plt.xlabel('Income per person (2000 US$)')
plt.ylabel('Residential electricity consumption (kWh)')
_ = plt.title('Scatterplot for the association between the income and the residential electricity consumption')
# +
correlation, pvalue = stats.pearsonr(subdata2['income'], subdata2['electricity'])
display(Markdown("The correlation coefficient is {:.3g} and the associated p-value is {:.3g}.".format(correlation, pvalue)))
display(Markdown("And the coefficient of determination is {:.3g}.".format(correlation**2)))
# -
# The Pearson test proves a significant positive relationship between income per person and residential electricity consumption as the p-value is below 0.05.
#
# Moreover, the square of the correlation coefficient, i.e. the coefficient of determination, is 0.425. This means that we can predict 42.5% of the variability of residential electricity consumption knowing the income per person.
#
# ## Potential moderator
#
# Now comes the analysis of Pearson correlation between different urban rate group to see if the urban rate is a moderator on the relationship between income per person and residential electricity consumption.
# +
def urban_group(row):
if row['urbanrate'] < 25.0:
return '0%<=..<25%'
elif row['urbanrate'] < 50.0:
return '25%<=..<50%'
elif row['urbanrate'] < 75.0:
return '50%<=..<75%'
else:
return '75%=<'
subdata3 = subdata2.copy()
subdata3['urban_group'] = pd.Categorical(subdata3.apply(lambda x: urban_group(x), axis=1))
# +
summary = dict()
for group in subdata3.urban_group.cat.categories:
moderator_group = subdata3[subdata3['urban_group'] == group]
summary[group] = stats.pearsonr(moderator_group['income'], moderator_group['electricity'])
df = (pd.DataFrame(summary)
.rename(index={0:'Pearson r', 1:'p-value'}))
# -
df2 = (df.stack()
.unstack(level=0))
df2.index.name = 'Urban rate'
df2
# For all urban rate categories, the p-value is below the threshold of 0.05. Therefore the urban rate does not moderate the relationship between income per person and residential electricity consumption. In other words the residential electricity consumption has a significant positive relationship in regard to the income per person whatever the urban rate in the country.
#
# By plotting the scatter plots of the four groups, we can see that the correlation is indeed present for all of them. One corollary finding from the graphics below is the tendency of countries with higher income per person to have higher urban rate.
# +
g = sns.FacetGrid(subdata3.reset_index(),
col='urban_group', hue='urban_group', col_wrap=2, size=4)
_ =g.map(sns.regplot, 'income', 'electricity')
# -
# This conclude the final assignment of this second course.
#
# > If you are interested into data sciences, follow me on [Tumblr](http://fcollonval.tumblr.com/).
|
PotentialModerator.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# cd /data/alv/CTCF_degron/analysis/ChIPseq/ChIP4/input
# ls
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
from bioframe.tools import intersect
from bioframe import read_table
from itertools import combinations
# -
import bioframe
chromsizes = bioframe.fetch_chromsizes('hg19')
chromosomes = list(chromsizes.index)
fnames = ["NT-CTCF-summits-sort-merge.bed"]
x = read_table(fnames[0],schema='bed')
x = x[x.chrom.isin(chromosomes)].reset_index(drop=True)
print(len(x))
x
# +
bedpes = []
gsep_min=200000
gsep_max=400000
df = x
for chrom in chromosomes[:-2]:
# create pairwise combinations of indices of CTCF peaks for a given chrom:
pairwise_index = pd.DataFrame(list(combinations(df.index[df['chrom']==chrom],2)))
# left,right of BEDPE ...
left=df.iloc[pairwise_index[0]].reset_index(drop=True)[["chrom","start","end"]]
right=df.iloc[pairwise_index[1]].reset_index(drop=True)[["chrom","start",'end']]
# concat them ...
cols = ["chrom1","start1","end1","chrom2","start2","end2"]
bedpe = pd.concat([left,right],ignore_index=True,axis=1).rename(dict(enumerate(cols)),axis=1)
# assert distances:
genomic_seps = bedpe['start2']-bedpe['start1']
#assert (genomic_seps>=0).all()
# accumulate
bedpes.append(bedpe[(genomic_seps>gsep_min)&(genomic_seps<gsep_max)])
bedpe_fin = pd.concat(bedpes, ignore_index=True)
bedpe_fin.to_csv("/data/alv/CTCF_degron/analysis/for-paper/draft4/figure2bis/dots/mao/G4-pairwise-200-400kb.bedpe")
# +
fff = ["G4-pairwise-200-400kb.bedpe",
]
for f in fff:
df = pd.read_csv("/data/alv/CTCF_degron/analysis/for-paper/draft4/figure2bis/dots/mao/"+f,index_col=0)
df.to_csv("/data/alv/CTCF_degron/analysis/for-paper/draft4/figure2bis/dots/mao/"+f+".tsv",index=False,sep='\t')
# -
|
Notebook_Archive/CTCFpeaks-pairwise-from-ZT.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# %pylab
# %matplotlib inline
# ### Part 1
def parse_paths(fn):
paths = []
with open(fn) as data:
for path in data.readlines():
start,_,stop = path.split()
paths += [[eval(start),eval(stop)]]
return paths
print(paths:=parse_paths('d5p1_test.txt'))
def map_danger_1(paths):
sz = np.array(paths).max()+1
diagram = np.zeros([sz,sz],dtype='uint32')
for path in paths:
if path[0][0]==path[1][0] or path[0][1]==path[1][1]: # horizonta and vertical
xrange = (min(path[0][0],path[1][0]),
max(path[0][0],path[1][0]))
yrange = (min(path[0][1],path[1][1]),
max(path[0][1],path[1][1]))
for i in range(xrange[0],xrange[1]+1):
for j in range(yrange[0],yrange[1]+1):
diagram[j,i]+=1
return diagram
diagram = map_danger_1(paths)
assert len(np.where(diagram>1)[0])==5
paths=parse_paths('d5p1.txt')
diagram = map_danger_1(paths)
print(f'The answer is {len(np.where(diagram>1)[0])}')
# ### Part 2
def map_danger_2(paths):
sz = np.array(paths).max()+1
diagram = np.zeros([sz,sz],dtype='uint32')
for path in paths:
xrange = (min(path[0][0],path[1][0]),
max(path[0][0],path[1][0]))
yrange = (min(path[0][1],path[1][1]),
max(path[0][1],path[1][1]))
if path[0][0]==path[1][0] or path[0][1]==path[1][1]: # horizonta and vertical
xrange = (min(path[0][0],path[1][0]),
max(path[0][0],path[1][0]))
yrange = (min(path[0][1],path[1][1]),
max(path[0][1],path[1][1]))
for i in range(xrange[0],xrange[1]+1):
for j in range(yrange[0],yrange[1]+1):
diagram[j,i]+=1
elif abs(path[0][0]-path[1][0])==abs(path[0][1]-path[1][1]): # must be diagonal by design -- trust but verify
xrange_ = np.arange(xrange[0],xrange[1]+1)
if path[0][0]>path[1][0]:
xrange_ = xrange_[::-1]
yrange_ = np.arange(yrange[0],yrange[1]+1)
if path[0][1]>path[1][1]:
yrange_ = yrange_[::-1]
for pos in zip(xrange_,yrange_):
i,j=pos
diagram[j,i]+=1
else:
print(f'Something is wrong with {path}')
return diagram
paths=parse_paths('d5p1_test.txt')
diagram = map_danger_2(paths)
print(diagram)
assert len(np.where(diagram>1)[0])==12
paths=parse_paths('d5p1.txt')
diagram = map_danger_2(paths)
#print(diagram)
print(f'The answer is {len(np.where(diagram>1)[0])}')
|
Day 5 - Hydrothermal Venture.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <div class="alert alert-info">
# **Note:**
#
# You can download this demo as a Jupyter notebook [here](https://github.com/ejhigson/dyPolyChord/blob/master/docs/demo.ipynb) and run it interactively yourself.
# </div>
#
# # dyPolyChord Demo
#
# The main user-facing function is ``dyPolyChord.run_dypolychord``, which performs dynamic nested sampling.
#
# Likelihoods and priors are specified within a Python callable, which can be used to run PolyChord on the likelihood and prior with an input settings dictionary. Tools for making such a callable are provided in ``pypolychord_utils.py`` (python likelihoods and priors) and ``polychord_utils.py`` (compiled C++ and Fortran likelihoods and priors).
#
# In addition the user can specify PolyChord settings (as a dictionary), and can choose whether to prioritise parameter estimation or evidence calculation via the ``dynamic_goal`` argument - see the [dynamic nested sampling paper (Higson et al., 2019)](https://doi.org/10.1007/s11222-018-9844-0) for an explanation.
# ## Compiled (C++ or Fortran) likelihoods and priors
#
# C++ and Fortran likelihoods used by PolyChord can also be used by ``dyPolyChord`` (they must be able to read settings from .ini files). These must be compiled to executables within the PolyChord directory, via commands such as
#
# $ make gaussain # PolyChord gaussian example
#
# or
#
# $ make polychord_CC_ini # PolyChord template C++ likelihood which reads .ini file
#
# See the PolyChord README for more details. ``dyPolyChord`` simply needs the file path to the executable, which it runs via ``os.system`` - settings are specified by writing temporary .ini files.
# +
import dyPolyChord.polychord_utils
import dyPolyChord
# Definte the distribution to sample (likelihood, prior, number of dimensions)
ex_command = './gaussian' # path to compiled executable likelihood
# The prior must be specified as strings via .ini files. get_prior_block_str provides a
# convenient function for making such PolyChord-formatted strings. See its docstring and
# the PolyChord documentation for more details
ndim = 10
prior_str = dyPolyChord.polychord_utils.get_prior_block_str(
'gaussian', # name of prior - see PolyChord for a list of allowed priors
[0.0, 10.0], # parameters of the prior
ndim)
# Make a callable for running PolyChord
my_callable = dyPolyChord.polychord_utils.RunCompiledPolyChord(
ex_command, prior_str)
# Specify sampler settings (see run_dynamic_ns.py documentation for more details)
dynamic_goal = 1.0 # whether to maximise parameter estimation or evidence accuracy.
ninit = 100 # number of live points to use in initial exploratory run.
nlive_const = 500 # total computational budget is the same as standard nested sampling with nlive_const live points.
settings_dict = {'file_root': 'gaussian',
'base_dir': 'chains',
'seed': 1}
# Run dyPolyChord
dyPolyChord.run_dypolychord(my_callable, dynamic_goal, settings_dict,
ninit=ninit, nlive_const=nlive_const)
# -
# ## Python likelihoods and priors
#
# Python likelihoods and priors must be defined as functions or callable classes, just as for running pypolychord (PolyChord's python wrapper). Otherwise the process is very similar to that with compiled likelihoods.
#
# Note that pypolychord used to be called PyPolyChord before PolyChord v1.15. ``dyPolyChord`` is compatible with both the new and old names; if pypolychord cannot be imported then we try importing PyPolyChord instead.
# +
import dyPolyChord.python_likelihoods as likelihoods # Import some example python likelihoods
import dyPolyChord.python_priors as priors # Import some example python priors
import dyPolyChord.pypolychord_utils
import dyPolyChord
# Definte the distribution to sample (likelihood, prior, number of dimensions)
ndim = 10
likelihood = likelihoods.Gaussian(sigma=1.0)
prior = priors.Gaussian(sigma=10.0)
# Make a callable for running PolyChord
my_callable = dyPolyChord.pypolychord_utils.RunPyPolyChord(
likelihood, prior, ndim)
# Specify sampler settings (see run_dynamic_ns.py documentation for more details)
dynamic_goal = 1.0 # whether to maximise parameter estimation or evidence accuracy.
ninit = 100 # number of live points to use in initial exploratory run.
nlive_const = 500 # total computational budget is the same as standard nested sampling with nlive_const live points.
settings_dict = {'file_root': 'gaussian',
'base_dir': 'chains',
'seed': 1}
# Run dyPolyChord
dyPolyChord.run_dypolychord(my_callable, dynamic_goal, settings_dict,
ninit=ninit, nlive_const=nlive_const)
# -
# ## Parallelisation
#
#
# #### Compiled likelihoods and priors
#
# To run compiled likelihoods in parallel with MPI, specify an mpirun command in the `mpi_str` argument when initializing your `RunPyPolyChord` object. For example to run with 8 processes, use
my_callable = dyPolyChord.polychord_utils.RunCompiledPolyChord(
ex_command, prior_str, mpi_str='mpirun -np 8')
# The callable can then be used with `run_dypolychord` as normal.
#
# #### Python likelihoods and priors
#
# You must import `mpi4py`, create an `MPI.COMM_WORLD` object and pass it to `run_dypolychord` as an argument.
# +
from mpi4py import MPI
comm = MPI.COMM_WORLD
dyPolyChord.run_dypolychord(my_callable, dynamic_goal, settings_dict,
ninit=ninit, nlive_const=nlive_const, comm=comm)
# -
# You can then run your script with mpirun:
#
# $ mpirun -np 8 my_dypolychord_script.py
#
# #### Repeated runs
#
# If you want to perform a number of independent `dyPolyChord` calculations (such as repeating the same calculation many times) then, as this is "embarrassingly parallel", you don't need MPI and can instead perform many `dyPolyChord` runs in parallel using python's `concurrent.futures`. This also allows reliable random seeding for reproducible results, which is not possible with MPI due to the unpredictable order in which slave processes are called by PolyChord. Note that for this to work PolyChord must be installed without MPI.
#
# For an example of this type of usage, see the code used to make the results for the dynamic nested sampling paper (https://github.com/ejhigson/dns).
# ## Checking the output
# Running ``dyPolyChord`` produces PolyChord-format output files in `settings['base_dir']`. These output files can be analysed in the same way as other PolyChord output files.
#
# One convenient package for doing this in python is ``nestcheck`` (http://nestcheck.readthedocs.io/en/latest/). We can use it to load and analyse the results from the 10-dimensional Gaussian produced by running the second cell in this demo (the example python likelihood and prior).
# +
import nestcheck.data_processing
import nestcheck.estimators as e
# load the run
run = nestcheck.data_processing.process_polychord_run(
'gaussian', # = settings['file_root']
'chains') # = settings['base_dir']
print('The log evidence estimate using the first run is {}'
.format(e.logz(run)))
print('The estimateed the mean of the first parameter is {}'
.format(e.param_mean(run, param_ind=0)))
# -
# As an illustration, lets use ``nestcheck`` to check ``dyPolyChord``'s allocation of live points roughly matches the distribution of posterior mass, which it should do when the dynamic goal setting equals 1. For a detailed explanation of this type of plot, see Figure 4 in the dynamic nested sampling paper ([Higson et al., 2019](https://doi.org/10.1007/s11222-018-9844-0)) and its caption.
# +
import numpy as np
import matplotlib.pyplot as plt
import nestcheck.ns_run_utils
# %matplotlib inline
# get the sample's estimated logX co-ordinates and relative posterior mass
logx = nestcheck.ns_run_utils.get_logx(run['nlive_array'])
logw = logx + run['logl']
w_rel = np.exp(logw - logw.max())
# plot nlive and w_rel on same axis
fig = plt.figure()
ax1 = fig.add_subplot(111)
ax2 = ax1.twinx()
l1 = ax1.plot(logx, run['nlive_array'], label='number of live points', color='blue')
l2 = ax2.plot(logx, w_rel, label='relative posterior mass', color='black', linestyle='dashed')
lines = l1 + l2
ax1.legend(lines, [l.get_label() for l in lines], loc=0)
ax1.set_xlabel('estimated $\log X$')
ax1.set_xlim(right=0.0)
ax1.set_ylim(bottom=0.0)
ax1.set_ylabel('number of live points')
ax2.set_ylim(bottom=0.0)
ax2.set_yticks([])
ax2.set_ylabel('relative posterior mass')
plt.show()
# -
# It looks like `dyPolyChord`'s allocation of samples closely matches the regions with high posterior mass, as expected.
#
# Note that this plot is only approximate as the $\log X$ values ($x$ coordinates) are estimated from just the one run, and are correlated with the posterior mass estimates. For a more accurate version, see Figure 4 of [Higson et al., (2019)](https://doi.org/10.1007/s11222-018-9844-0).
|
docs/demo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib
import matplotlib.pyplot as plt
from plots import *
from math import sqrt
# +
import matplotlib.pyplot as plt
from matplotlib import rc
import seaborn as sns
rc('font', **{'family': 'sans-serif',
'sans-serif': ['Computer Modern Roman']})
params = {'axes.labelsize': 12,
'font.size': 12,
'legend.fontsize': 12,
'xtick.labelsize': 10,
'ytick.labelsize': 10,
'text.usetex': True,
'figure.figsize': (8, 6)}
plt.rcParams.update(params)
sns.set_context("poster")
sns.set_style("ticks")
# -
xls = pd.ExcelFile("Final Plotting/Experiment - 2_1.xlsx")
df1 = pd.read_excel(xls, sheet_name="VGG_cifar10", index_col=0)
df2 = pd.read_excel(xls, sheet_name="VGG_cifar100", index_col=0)
df3 = pd.read_excel(xls, sheet_name="VGG_SVHN", index_col=0)
df4 = pd.read_excel(xls, sheet_name="VGG_FashionMNIST", index_col = 0)
# # VGG CIFAR10
df1.head()
df1 = df1.rename(columns = {"Unnamed: 1":"% Pruned", "Accuracy on CIFAR10":"CIFAR10", "Unnamed: 3":"CIFAR100", "Unnamed: 4":"Random"})
df1 = df1.iloc[1:]
df1.dropna(inplace = True)
x = np.float64(df1["% Pruned"].values)
cifar_10 = np.float64(df1["CIFAR10"].values)
cifar_100 = np.float64(df1["CIFAR100"].values)
rd = np.float64(df1["Random"].values)
idx_vals = [0,20,36,48.8,59.04,67.23,73.79,86.58,93.13,96.48,98.2,99.08,99.53,99.81,99.88]
idx = []
for i in range(len(x)):
for j in range(len(idx_vals)):
if x[i] == idx_vals[j]:
idx.append(i)
idx.sort()
plt.figure(figsize=(10,7))
plt.plot([i for i in range(len(idx))], [cifar_10[i]/100 for i in idx], label='CIFAR-10', marker = '.')
plt.plot([i for i in range(len(idx))], [rd[i]/100 for i in idx], label="Random", marker='^')
plt.plot([i for i in range(len(idx))], [cifar_100[i]/100 for i in idx], label='CIFAR-100', marker='*')
plt.title("CIFAR-10")
plt.xlabel("Fraction of weights pruned")
plt.ylabel("CIFAR-10 test accuracy\n at convergence")
tick_lbls = [np.round(x[i]/100, 3) for i in idx]
plt.xticks(range(len(idx)), tick_lbls, rotation = -45)
plt.grid(axis='y')
plt.legend(loc="best",title="Ticket Source")
# plt.ylim(0.5,0.90)
plt.tight_layout()
plt.savefig("finalplots/Exp2_VGG_CIFAR10_N.pdf")
# # VGG CIFAR 100
df2.head()
df2 = df2.rename(columns = {"Unnamed: 1":"% Pruned", "Accuracy on CIFAR100":"CIFAR10", "Unnamed: 3":"CIFAR100", "Unnamed: 4":"Random"})
df2 = df2.loc[:, ~df2.columns.str.contains('^Unnamed')]
df2.dropna(inplace = True)
df2 = df2.iloc[1:]
df2.head()
x = np.float64(df2["% Pruned"].values)
cifar_10 = np.float64(df2["CIFAR10"].values)
cifar_100 = np.float64(df2["CIFAR100"].values)
rd = np.float64(df2["Random"].values)
idx_vals = [0,20,36,48.8,59.04,67.23,73.79,86.58,93.13,96.48,98.2,99.08,99.53,99.81,99.88]
idx = []
for i in range(len(x)):
for j in range(len(idx_vals)):
if x[i] == idx_vals[j]:
idx.append(i)
idx.sort()
plt.figure(figsize=(10,7))
plt.plot([i for i in range(len(idx))], [cifar_10[i]/100 for i in idx], label='CIFAR-10', marker = '.')
plt.plot([i for i in range(len(idx))], [rd[i]/100 for i in idx], label="Random", marker='^')
plt.plot([i for i in range(len(idx))], [cifar_100[i]/100 for i in idx], label='CIFAR-100', marker='*')
plt.title("CIFAR-100")
plt.xlabel("Fraction of weights pruned")
plt.ylabel("CIFAR-100 test accuracy\n at convergence")
tick_lbls = [np.round(x[i]/100, 3) for i in idx]
plt.xticks(range(len(idx)), tick_lbls, rotation = -45)
plt.grid(axis='y')
plt.legend(loc="best",title="Ticket Source")
plt.ylim(0.4,0.75)
plt.tight_layout()
plt.savefig("finalplots/Exp2_VGG_CIFAR100_N1.pdf")
# # SVHN
df3.head()
df3 = df3.rename(columns = {"Unnamed: 1":"% Pruned", "Accuracy on SVHN":"CIFAR10", "Unnamed: 3":"CIFAR100", "Unnamed: 4":"SVHN","Unnamed: 5":"Random"})
df3.dropna(inplace = True)
df3 = df3.iloc[1:]
df3.head()
x = np.float64(df3["% Pruned"].values)
cifar_10 = np.float64(df3["CIFAR10"].values)
cifar_100 = np.float64(df3["CIFAR100"].values)
svhn = np.float64(df3["SVHN"].values)
rd = np.float64(df3["Random"].values)
idx_vals = [0,20,36,48.8,59.04,67.23,73.79,86.58,93.13,96.48,98.2,99.08,99.53,99.81,99.88]
idx = []
for i in range(len(x)):
for j in range(len(idx_vals)):
if x[i] == idx_vals[j]:
idx.append(i)
idx.sort()
# +
plt.figure(figsize=(10,7))
plt.plot([i for i in range(len(idx))], [cifar_10[i]/100 for i in idx], label='CIFAR-10', marker = '.')
plt.plot([i for i in range(len(idx))], [rd[i]/100 for i in idx], label="Random", marker='^')
plt.plot([i for i in range(len(idx))], [cifar_100[i]/100 for i in idx], label='CIFAR-100', marker='*')
plt.plot([i for i in range(len(idx))], [svhn[i]/100 for i in idx], label='SVHN', marker='x')
plt.title("SVHN")
plt.xlabel("Fraction of weights pruned")
plt.ylabel("SVHN test accuracy\n at convergence")
tick_lbls = [np.round(x[i]/100, 3) for i in idx]
plt.xticks(range(len(idx)), tick_lbls, rotation = -45)
plt.grid(axis='y')
plt.legend(loc="best",title="Ticket Source")
plt.ylim(0.75,0.99)
plt.tight_layout()
plt.savefig("finalplots/Exp2_VGG_SVHN_N.pdf")
# -
df4.head()
df4 = df4.rename(columns = {"Unnamed: 1":"% Pruned", "Accuracy on FashionMNIST":"CIFAR10", "Unnamed: 3":"CIFAR100", "Unnamed: 4":"FashionMNIST","Unnamed: 5":"Random"})
df4.dropna(inplace = True)
df4 = df4.iloc[1:]
df4.head()
x = np.float64(df4["% Pruned"].values)
cifar_10 = np.float64(df4["CIFAR10"].values)
cifar_100 = np.float64(df4["CIFAR100"].values)
fmnist = np.float64(df4["FashionMNIST"].values)
rd = np.float64(df4["Random"].values)
idx_vals = [0,20,36,48.8,59.04,67.23,73.79,86.58,93.13,96.48,98.2,99.08,99.53,99.81,99.88]
idx = []
for i in range(len(x)):
for j in range(len(idx_vals)):
if x[i] == idx_vals[j]:
idx.append(i)
idx.sort()
# +
plt.figure(figsize=(15,7))
plt.plot([i for i in range(len(idx))], [cifar_10[i]/100 for i in idx], label='CIFAR-10', marker = '.', alpha=0.9)
plt.plot([i for i in range(len(idx))], [rd[i]/100 for i in idx], label="Random", marker='^', alpha=0.5)
plt.plot([i for i in range(len(idx))], [cifar_100[i]/100 for i in idx], label='CIFAR-100', marker='*', alpha=0.5)
plt.plot([i for i in range(len(idx))], [fmnist[i]/100 for i in idx], label='FashionMNIST', marker='x', alpha=0.5)
plt.title("FashionMNIST")
plt.xlabel("Fraction of weights pruned")
plt.ylabel("FashionMNIST test accuracy\n at convergence")
tick_lbls = [np.round(x[i]/100, 3) for i in idx]
plt.xticks(range(len(idx)), tick_lbls, rotation = -45)
plt.grid(axis='y')
plt.legend(loc="best",title="Ticket Source")
plt.ylim(0.0,0.9)
plt.tight_layout()
plt.savefig("finalplots/Exp2_VGG_FashionMNIST1.pdf")
|
plots/codes/exp2_vgg.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Example of performing linear least squares fittings
#
# First we import numpy and matplotlib as usual
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
# Now, let's generate some random data about a trend line
# +
#set a random number seed
np.random.seed(119)
#set number of data points
npoints = 50
#set x
x = np.linspace(0,10.,npoints)
#set slope, intecept , and scatter rms
m = 2.0
b = 1.0
sigma = 2.0 #error, width of guassian
#generate y points
y = m*x + b + np.random.normal(scale=sigma,size=npoints)
y_err = np.full(npoints,sigma)
# -
# # Let's just plot the data first
f = plt.figure(figsize=(7,7))
plt.errorbar(x,y,sigma,fmt='ro')
plt.xlabel('x')
plt.ylabel('y')
plt.show()
# # Method #1, polyfit()
m_fit, b_fit = np.poly1d(np.polyfit(x ,y , 1, w =1. / y_err)) #weight the uncertainties
print(m_fit, b_fit)
y_fit = m_fit * x + b_fit
# # Plot result
f = plt.figure(figsize=(7,7))
plt.errorbar(x,y,yerr=y_err,fmt = 'ro',label = 'data')
plt.plot(x,y_fit,label = 'fit')
plt.xlabel('x')
plt.ylabel('y')
plt.legend(loc=2,frameon=False)
plt.show()
# # Method #2, scipy + optimize
# +
#import optimize from scipy
from scipy import optimize
#define the finction to the fit
def f_line(x, m ,b):
return m*x + b
#perform the fit
params, params_cov = optimize.curve_fit(f_line,x,y,sigma=y_err)
m_fit = params[0]
b_fit = params[1]
print(m_fit,b_fit)
# -
# # Plot the result
f = plt.figure(figsize=(7,7))
plt.errorbar(x,y,yerr=y_err,fmt='ro',label='data')
plt.plot(x,y_fit,label='fit')
plt.xlabel('x')
plt.ylabel('y')
plt.legend(loc=0,frameon=False)
plt.show()
# # We can perform much more complicated fits...
#
# +
#redefine x and y
npoints = 50
x = np.linspace(0.,2*np.pi,npoints)
#make y a complicated function
a = 3.4
b = 2.1
c = 0.27
d = -1.3
sig = 0.6
y = a * np.sin( b*x +c) + d + np.random.normal(scale=sig,size=npoints)
y_err = np.full(npoints,sig)
f = plt.figure(figsize=(7,7))
plt.errorbar(x,y,yerr=y_err,fmt='o')
plt.xlabel('x')
plt.ylabel('y')
plt.show()
# -
# ## Perform a fit using scipy.optimize.curve_fit()
# +
#import optimize from scipy
from scipy import optimize
#define the function to fit
def f_line(x, a, b, c, d):
return a * np.sin(b*x +c) + d
#perform the fit
params, params_cov = optimize.curve_fit(f_line,x,y,sigma=y_err,p0=[1,2.,0.1,-0.1])
a_fit = params[0]
b_fit = params[1]
c_fit = params[2]
d_fit = params[3]
print(a_fit,b_fit,c_fit,d_fit)
y_fit = a_fit * np.sin(b_fit * x + c_fit) + d_fit
# -
# # Plot the fit
f = plt.figure(figsize=(7,7))
plt.errorbar(x,y,yerr=y_err,fmt='o',label='data')
plt.plot(x,y_fit,label='fit')
plt.xlabel('x')
plt.ylabel('y')
plt.legend(loc=0,frameon=False)
plt.show()
|
astr_119_session_7.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Amazon Fine Food Reviews Analysis
#
#
# Data Source: https://www.kaggle.com/snap/amazon-fine-food-reviews <br>
#
# Amazon fine food review dataset is medium large dataset which consists.<br>
#
# Number of reviews: 568,454<br>
# Number of users: 256,059<br>
# Number of products: 74,258<br>
# Timespan: Oct 1999 - Oct 2012<br>
# Number of Dimensions/Columns in data: 10
#
# Each Attribute Information:
#
# 1. Id - just a sequence of reviews
# 2. ProductId - unique identifier for the product
# 3. UserId - unqiue identifier for the user
# 4. ProfileName - profile name of the user
# 5. HelpfulnessNumerator - number of users who found the review helpful
# 6. HelpfulnessDenominator - number of users who indicated whether they found the review helpful or not(total)
# 7. Score - rating between 1 and 5
# 8. Time - timestamp for the review
# 9. Summary - brief summary of the review
# 10. Text - text of the review
#
#
# ### Problem Statement:
# Given a review, determine whether the review is positive (rating of 4 or 5) or negative (rating of 1 or 2).
# Just avoiding rating 3 because it considered as neutral.
#
# %matplotlib inline
import warnings
warnings.filterwarnings("ignore")
import sqlite3
import pandas as pd
import numpy as np
import nltk
import matplotlib.pyplot as plt
from sklearn.feature_extraction.text import CountVectorizer
import re
# Tutorial about Python regular expressions: https://pymotw.com/2/re/
from nltk.corpus import stopwords
from bs4 import BeautifulSoup
import pickle
from nltk.stem import PorterStemmer
from wordcloud import WordCloud, STOPWORDS
from tqdm import tqdm
# #### Data Loading
# * To load data here we have to perform sqlite queries with python.
# FIrst we have to create connection to the database we have
connection = sqlite3.connect("database.sqlite")
data = pd.read_sql_query(""" SELECT * FROM Reviews """, connection)
# After reading we have to close the connection
connection.close()
data.head(3)
data.shape
data.info()
data.describe()["Score"]
# **Observation**
# * As we can clearly see that mean rating is 4.1 and minimum is 1 and maximum is 5.
# * And memory used by data is 43.4 MegaBytes
# * And There is not any null values avalaible in any column.
# * Number of revies we have is approx 5.6 Millions.
# Lets see distributions of ratings
ratings = dict(data["Score"].value_counts())
x = list(ratings.keys())
y = list(ratings.values())
plt.figure(figsize=(10,6))
plt.bar(x,y)
plt.title("Rating distribution")
plt.ylabel("Nmber of rviews")
plt.xlabel("Ratings")
plt.show()
# As we can see that maximum number of reviews have 5 ratings.
duplicates = data.duplicated(["UserId","ProfileName","Time","Text"])
print(sum(duplicates))
# As we can see we have 174512 duplicates in our dataset.so we have to remove that because they does not have any benefits in our model.
# To drop duplicates
data.drop_duplicates(subset={"UserId","ProfileName","Time","Text"}, keep='first', inplace=True)
data.shape
# Now we have only 393933 row of reviews.
# Now lets drop the row which has 3 ratings
index_of_three = data[ data['Score'] == 3 ].index
data.drop(index_of_three, inplace = True)
data.shape
# Now we have new shape (364164,10)
# Give reviews with Score>3 a 1(positive) rating, and reviews with a score<3 a 0(negative) rating.
def partition(x):
if x < 3:
return '0'
return '1'
data["Score"] = data["Score"].map(partition)
data.head(3)
# Till now we have cleared our data besed on number or rows and the score type and also dropped duplicates.<br>
# Now time to preprocess data<br>
# ### Data preprocessing
# Hence in the Preprocessing phase we do the following in the order below:-
#
# * Begin by removing the html tags
# * Remove any punctuations or limited set of special characters like , or . or # etc.
# * Check if the word is made up of english letters and is not alpha-numeric
# * Check to see if the length of the word is greater than 2 (as it was researched that there is no adjective in 2-letters)
# * Convert the word to lowercase
# * Remove Stopwords
# * Finally Snowball Stemming the word (it was obsereved to be better than Porter Stemming)<br>
#
# After which we collect the words used to describe positive and negative reviews
# * Here we taking only 100000 review to work faster and due to lack of memory
# Make 100000 samples
sample_data = data.sample(100000)
sample_data.shape
# https://stackoverflow.com/a/47091490/4084039
import re
def decontracted(phrase):
# specific
phrase = re.sub(r"won't", "will not", phrase)
phrase = re.sub(r"can\'t", "can not", phrase)
# general
phrase = re.sub(r"n\'t", " not", phrase)
phrase = re.sub(r"\'re", " are", phrase)
phrase = re.sub(r"\'s", " is", phrase)
phrase = re.sub(r"\'d", " would", phrase)
phrase = re.sub(r"\'ll", " will", phrase)
phrase = re.sub(r"\'t", " not", phrase)
phrase = re.sub(r"\'ve", " have", phrase)
phrase = re.sub(r"\'m", " am", phrase)
return phrase
stopwords= set(['br', 'the', 'i', 'me', 'my', 'myself', 'we', 'our', 'ours', 'ourselves', 'you', "you're", "you've",\
"you'll", "you'd", 'your', 'yours', 'yourself', 'yourselves', 'he', 'him', 'his', 'himself', \
'she', "she's", 'her', 'hers', 'herself', 'it', "it's", 'its', 'itself', 'they', 'them', 'their',\
'theirs', 'themselves', 'what', 'which', 'who', 'whom', 'this', 'that', "that'll", 'these', 'those', \
'am', 'is', 'are', 'was', 'were', 'be', 'been', 'being', 'have', 'has', 'had', 'having', 'do', 'does', \
'did', 'doing', 'a', 'an', 'the', 'and', 'but', 'if', 'or', 'because', 'as', 'until', 'while', 'of', \
'at', 'by', 'for', 'with', 'about', 'against', 'between', 'into', 'through', 'during', 'before', 'after',\
'above', 'below', 'to', 'from', 'up', 'down', 'in', 'out', 'on', 'off', 'over', 'under', 'again', 'further',\
'then', 'once', 'here', 'there', 'when', 'where', 'why', 'how', 'all', 'any', 'both', 'each', 'few', 'more',\
'most', 'other', 'some', 'such', 'only', 'own', 'same', 'so', 'than', 'too', 'very', \
's', 't', 'can', 'will', 'just', 'don', "don't", 'should', "should've", 'now', 'd', 'll', 'm', 'o', 're', \
've', 'y', 'ain', 'aren', "aren't", 'couldn', "couldn't", 'didn', "didn't", 'doesn', "doesn't", 'hadn',\
"hadn't", 'hasn', "hasn't", 'haven', "haven't", 'isn', "isn't", 'ma', 'mightn', "mightn't", 'mustn',\
"mustn't", 'needn', "needn't", 'shan', "shan't", 'shouldn', "shouldn't", 'wasn', "wasn't", 'weren', "weren't", \
'won', "won't", 'wouldn', "wouldn't"])
import nltk
nltk.download('wordnet')
# Combining all the above stundents
from tqdm import tqdm
# To make lemetize words
from nltk.stem import WordNetLemmatizer
lemmatizer = WordNetLemmatizer()
prep_reviews = []
ratings = []
for row in tqdm(range(sample_data.shape[0])):
# Here we are giving more importance to the Summary becuse it contains more acurate and more information about text itself.
sentance = 2*(sample_data["Summary"].iloc[row]+" ") + sample_data["Text"].iloc[row]
sentance = re.sub(r"http\S+", "", sentance)
# To remove any html and XML
sentance = BeautifulSoup(sentance, 'lxml').get_text()
# To change can't to can not
sentance = decontracted(sentance)
sentance = re.sub("\S*\d\S*", "", sentance).strip()
# Keep only english letters
sentance = re.sub('[^A-Za-z]+', ' ', sentance)
p_stammer = PorterStemmer()
# Remove stop words and convert it into lower case
prep_sent = ""
for word in sentance.split():
if word.lower() not in stopwords:
word = lemmatizer.lemmatize(word)
prep_sent += word.lower()+" "
prep_reviews.append(prep_sent.strip())
ratings.append(sample_data["Score"].iloc[row])
prep_reviews[1:5]
prep_review_df = pd.DataFrame({"Reviews":prep_reviews,"rating":ratings})
prep_review_df.to_csv("AFFR_preprocessed_100k.csv",index=False)
# +
# Make word count vectorizer
def text_splitter(text):
return text.split()
vectorizer = CountVectorizer(tokenizer = text_splitter)
review_vector = vectorizer.fit_transform(prep_review_df['Reviews'].values.astype(str))
# -
review_vector.shape
# To check the feature names or vocabulary of our data
print(vectorizer.get_feature_names()[20:30])
review_vector[0].toarray().shape
column_sum = review_vector.sum(axis=0).tolist()[0]
# * Now we see that our vocabulary size is 60140
# * And now each word is represented by the 60140 dimension vector
#
review_dict = dict(zip(vectorizer.get_feature_names(),column_sum))
# +
# Ploting word cloud
#Initializing WordCloud using frequencies of tags.
wordcloud = WordCloud(background_color='black', width=1600, height=800,).generate_from_frequencies(review_dict)
fig = plt.figure(figsize=(12,8))
# To show the generated image
plt.imshow(wordcloud)
plt.axis('off')
plt.tight_layout(pad=0)
fig.savefig("vocab.png")
plt.show()
# -
# * The Big ones are more frequent words in this dataset
# ## Now we are done with dasic cleaning and preprocessing Then go to featurization notebook to understand featurization on text data.
|
.ipynb_checkpoints/EDA-Amazon fine food reviews-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Testing the models (90-5-5)
# +
import tensorflow as tf
config = tf.ConfigProto()
config.gpu_options.allow_growth = True
sess = tf.Session(config=config)
import os, shutil
from keras.preprocessing.image import ImageDataGenerator
import numpy as np
base_dir = r'tiles'
test_dir = os.path.join(base_dir, 'test')
test_datagen = ImageDataGenerator(rescale=1./255)
test_generator = test_datagen.flow_from_directory( test_dir, target_size=(1024, 1024),
batch_size=9, class_mode='binary', shuffle=False)
from keras.models import load_model
import warnings
warnings.filterwarnings(action="ignore")
import matplotlib.pyplot as plt
import numpy as np
from keras.models import load_model
import numpy as np
from sklearn.metrics import classification_report, confusion_matrix
import itertools
from collections import Counter
from math import pow
def plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, cm[i, j],
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('Actual class')
plt.xlabel('Predicted class')
# -
# # VGG-3
# ## 1
model = load_model('Models1/VGG3-1.h5')
test_steps_per_epoch = np.math.ceil(test_generator.samples / test_generator.batch_size)
test_loss, test_acc = model.evaluate_generator(test_generator, test_steps_per_epoch)
print('Test score: ', test_loss) #Loss on test
print('Test accuracy: ', test_acc)
# +
from sklearn.metrics import f1_score, precision_score, recall_score, confusion_matrix, classification_report
model.evaluate_generator(generator=test_generator, steps=test_steps_per_epoch)
test_generator.reset()
predictions = model.predict_generator(test_generator, steps=test_steps_per_epoch, verbose=0,use_multiprocessing=False,workers = 0) #pickle_safe = True, workers = 1
y_pred = np.rint(predictions)
Y_pred = predictions
Y_pred_classes = y_pred
Y_true = test_generator.classes
true_classes = test_generator.classes
class_labels = list(test_generator.class_indices.keys())
print(class_labels)
target_names = ['Road', 'No Road']
print(confusion_matrix(test_generator.classes, y_pred))
report = classification_report(test_generator.classes, y_pred,target_names=target_names, digits=4)
print(report)
# -
confusion_mtx= confusion_matrix(Y_pred_classes, Y_true)
plt.figure(figsize = (6.5,4.5))
plot_confusion_matrix(confusion_mtx, classes = ['Road', 'No Road'], cmap='Blues')
from sklearn.metrics import roc_auc_score
roc_auc_score = roc_auc_score(test_generator.classes, predictions)
roc_auc_score
# ## 2
model = load_model('Models1/VGG3-2.h5')
test_steps_per_epoch = np.math.ceil(test_generator.samples / test_generator.batch_size)
test_loss, test_acc = model.evaluate_generator(test_generator, test_steps_per_epoch)
print('Test score: ', test_loss) #Loss on test
print('Test accuracy: ', test_acc)
# +
from sklearn.metrics import f1_score, precision_score, recall_score, confusion_matrix, classification_report
model.evaluate_generator(generator=test_generator, steps=test_steps_per_epoch)
test_generator.reset()
predictions = model.predict_generator(test_generator, steps=test_steps_per_epoch, verbose=0,use_multiprocessing=False,workers = 0) #pickle_safe = True, workers = 1
y_pred = np.rint(predictions)
Y_pred = predictions
Y_pred_classes = y_pred
Y_true = test_generator.classes
true_classes = test_generator.classes
class_labels = list(test_generator.class_indices.keys())
print(class_labels)
target_names = ['Road', 'No Road']
print(confusion_matrix(test_generator.classes, y_pred))
report = classification_report(test_generator.classes, y_pred,target_names=target_names, digits=4)
print(report)
# -
confusion_mtx= confusion_matrix(Y_pred_classes, Y_true)
plt.figure(figsize = (6.5,4.5))
plot_confusion_matrix(confusion_mtx, classes = ['Road', 'No Road'], cmap='Blues')
from sklearn.metrics import roc_auc_score
roc_auc_score = roc_auc_score(test_generator.classes, predictions)
roc_auc_score
# ## 3
model = load_model('Models1/VGG3-3.h5')
test_steps_per_epoch = np.math.ceil(test_generator.samples / test_generator.batch_size)
test_loss, test_acc = model.evaluate_generator(test_generator, test_steps_per_epoch)
print('Test score: ', test_loss) #Loss on test
print('Test accuracy: ', test_acc)
# +
from sklearn.metrics import f1_score, precision_score, recall_score, confusion_matrix, classification_report
model.evaluate_generator(generator=test_generator, steps=test_steps_per_epoch)
test_generator.reset()
predictions = model.predict_generator(test_generator, steps=test_steps_per_epoch, verbose=0,use_multiprocessing=False,workers = 0) #pickle_safe = True, workers = 1
y_pred = np.rint(predictions)
Y_pred = predictions
Y_pred_classes = y_pred
Y_true = test_generator.classes
true_classes = test_generator.classes
class_labels = list(test_generator.class_indices.keys())
print(class_labels)
target_names = ['Road', 'No Road']
print(confusion_matrix(test_generator.classes, y_pred))
report = classification_report(test_generator.classes, y_pred,target_names=target_names, digits=4)
print(report)
# -
confusion_mtx= confusion_matrix(Y_pred_classes, Y_true)
plt.figure(figsize = (6.5,4.5))
plot_confusion_matrix(confusion_mtx, classes = ['Road', 'No Road'], cmap='Blues')
from sklearn.metrics import roc_auc_score
roc_auc_score = roc_auc_score(test_generator.classes, predictions)
roc_auc_score
# # VGG-5
# ## 1
model = load_model('Models1/VGG5-1.h5')
test_steps_per_epoch = np.math.ceil(test_generator.samples / test_generator.batch_size)
test_loss, test_acc = model.evaluate_generator(test_generator, test_steps_per_epoch)
print('Test score: ', test_loss) #Loss on test
print('Test accuracy: ', test_acc)
# +
from sklearn.metrics import f1_score, precision_score, recall_score, confusion_matrix, classification_report
model.evaluate_generator(generator=test_generator, steps=test_steps_per_epoch)
test_generator.reset()
predictions = model.predict_generator(test_generator, steps=test_steps_per_epoch, verbose=0,use_multiprocessing=False,workers = 0) #pickle_safe = True, workers = 1
y_pred = np.rint(predictions)
Y_pred = predictions
Y_pred_classes = y_pred
Y_true = test_generator.classes
true_classes = test_generator.classes
class_labels = list(test_generator.class_indices.keys())
print(class_labels)
target_names = ['Road', 'No Road']
print(confusion_matrix(test_generator.classes, y_pred))
report = classification_report(test_generator.classes, y_pred,target_names=target_names, digits=4)
print(report)
# -
confusion_mtx= confusion_matrix(Y_pred_classes, Y_true)
plt.figure(figsize = (6.5,4.5))
plot_confusion_matrix(confusion_mtx, classes = ['Road', 'No Road'], cmap='Blues')
from sklearn.metrics import roc_auc_score
roc_auc_score = roc_auc_score(test_generator.classes, predictions)
roc_auc_score
# ## 2
model = load_model('Models1/VGG5-2.h5')
test_steps_per_epoch = np.math.ceil(test_generator.samples / test_generator.batch_size)
test_loss, test_acc = model.evaluate_generator(test_generator, test_steps_per_epoch)
print('Test score: ', test_loss) #Loss on test
print('Test accuracy: ', test_acc)
# +
from sklearn.metrics import f1_score, precision_score, recall_score, confusion_matrix, classification_report
model.evaluate_generator(generator=test_generator, steps=test_steps_per_epoch)
test_generator.reset()
predictions = model.predict_generator(test_generator, steps=test_steps_per_epoch, verbose=0,use_multiprocessing=False,workers = 0) #pickle_safe = True, workers = 1
y_pred = np.rint(predictions)
Y_pred = predictions
Y_pred_classes = y_pred
Y_true = test_generator.classes
true_classes = test_generator.classes
class_labels = list(test_generator.class_indices.keys())
print(class_labels)
target_names = ['Road', 'No Road']
print(confusion_matrix(test_generator.classes, y_pred))
report = classification_report(test_generator.classes, y_pred,target_names=target_names, digits=4)
print(report)
# -
confusion_mtx= confusion_matrix(Y_pred_classes, Y_true)
plt.figure(figsize = (6.5,4.5))
plot_confusion_matrix(confusion_mtx, classes = ['Road', 'No Road'], cmap='Blues')
from sklearn.metrics import roc_auc_score
roc_auc_score = roc_auc_score(test_generator.classes, predictions)
roc_auc_score
# ## 3
model = load_model('Models1/VGG5-3.h5')
test_steps_per_epoch = np.math.ceil(test_generator.samples / test_generator.batch_size)
test_loss, test_acc = model.evaluate_generator(test_generator, test_steps_per_epoch)
print('Test score: ', test_loss) #Loss on test
print('Test accuracy: ', test_acc)
# +
from sklearn.metrics import f1_score, precision_score, recall_score, confusion_matrix, classification_report
model.evaluate_generator(generator=test_generator, steps=test_steps_per_epoch)
test_generator.reset()
predictions = model.predict_generator(test_generator, steps=test_steps_per_epoch, verbose=0,use_multiprocessing=False,workers = 0) #pickle_safe = True, workers = 1
y_pred = np.rint(predictions)
Y_pred = predictions
Y_pred_classes = y_pred
Y_true = test_generator.classes
true_classes = test_generator.classes
class_labels = list(test_generator.class_indices.keys())
print(class_labels)
target_names = ['Road', 'No Road']
print(confusion_matrix(test_generator.classes, y_pred))
report = classification_report(test_generator.classes, y_pred,target_names=target_names, digits=4)
print(report)
# -
confusion_mtx= confusion_matrix(Y_pred_classes, Y_true)
plt.figure(figsize = (6.5,4.5))
plot_confusion_matrix(confusion_mtx, classes = ['Road', 'No Road'], cmap='Blues')
from sklearn.metrics import roc_auc_score
roc_auc_score = roc_auc_score(test_generator.classes, predictions)
roc_auc_score
# # VGG-from-scratch
# ## 1
model = load_model('Models1/vgg16_noweights-1.h5')
test_steps_per_epoch = np.math.ceil(test_generator.samples / test_generator.batch_size)
test_loss, test_acc = model.evaluate_generator(test_generator, test_steps_per_epoch)
print('Test score: ', test_loss) #Loss on test
print('Test accuracy: ', test_acc)
# +
from sklearn.metrics import f1_score, precision_score, recall_score, confusion_matrix, classification_report
model.evaluate_generator(generator=test_generator, steps=test_steps_per_epoch)
test_generator.reset()
predictions = model.predict_generator(test_generator, steps=test_steps_per_epoch, verbose=0,use_multiprocessing=False,workers = 0) #pickle_safe = True, workers = 1
y_pred = np.rint(predictions)
Y_pred = predictions
Y_pred_classes = y_pred
Y_true = test_generator.classes
true_classes = test_generator.classes
class_labels = list(test_generator.class_indices.keys())
print(class_labels)
target_names = ['Road', 'No Road']
print(confusion_matrix(test_generator.classes, y_pred))
report = classification_report(test_generator.classes, y_pred,target_names=target_names, digits=4)
print(report)
# -
confusion_mtx= confusion_matrix(Y_pred_classes, Y_true)
plt.figure(figsize = (6.5,4.5))
plot_confusion_matrix(confusion_mtx, classes = ['Road', 'No Road'], cmap='Blues')
from sklearn.metrics import roc_auc_score
roc_auc_score = roc_auc_score(test_generator.classes, predictions)
roc_auc_score
# ## 2
model = load_model('Models1/vgg16_noweights-2.h5')
test_steps_per_epoch = np.math.ceil(test_generator.samples / test_generator.batch_size)
test_loss, test_acc = model.evaluate_generator(test_generator, test_steps_per_epoch)
print('Test score: ', test_loss) #Loss on test
print('Test accuracy: ', test_acc)
# +
from sklearn.metrics import f1_score, precision_score, recall_score, confusion_matrix, classification_report
model.evaluate_generator(generator=test_generator, steps=test_steps_per_epoch)
test_generator.reset()
predictions = model.predict_generator(test_generator, steps=test_steps_per_epoch, verbose=0,use_multiprocessing=False,workers = 0) #pickle_safe = True, workers = 1
y_pred = np.rint(predictions)
Y_pred = predictions
Y_pred_classes = y_pred
Y_true = test_generator.classes
true_classes = test_generator.classes
class_labels = list(test_generator.class_indices.keys())
print(class_labels)
target_names = ['Road', 'No Road']
print(confusion_matrix(test_generator.classes, y_pred))
report = classification_report(test_generator.classes, y_pred,target_names=target_names, digits=4)
print(report)
# -
confusion_mtx= confusion_matrix(Y_pred_classes, Y_true)
plt.figure(figsize = (6.5,4.5))
plot_confusion_matrix(confusion_mtx, classes = ['Road', 'No Road'], cmap='Blues')
from sklearn.metrics import roc_auc_score
roc_auc_score = roc_auc_score(test_generator.classes, predictions)
roc_auc_score
# ## 3
model = load_model('Models1/vgg16_noweights-3.h5')
test_steps_per_epoch = np.math.ceil(test_generator.samples / test_generator.batch_size)
test_loss, test_acc = model.evaluate_generator(test_generator, test_steps_per_epoch)
print('Test score: ', test_loss) #Loss on test
print('Test accuracy: ', test_acc)
# +
from sklearn.metrics import f1_score, precision_score, recall_score, confusion_matrix, classification_report
model.evaluate_generator(generator=test_generator, steps=test_steps_per_epoch)
test_generator.reset()
predictions = model.predict_generator(test_generator, steps=test_steps_per_epoch, verbose=0,use_multiprocessing=False,workers = 0) #pickle_safe = True, workers = 1
y_pred = np.rint(predictions)
Y_pred = predictions
Y_pred_classes = y_pred
Y_true = test_generator.classes
true_classes = test_generator.classes
class_labels = list(test_generator.class_indices.keys())
print(class_labels)
target_names = ['Road', 'No Road']
print(confusion_matrix(test_generator.classes, y_pred))
report = classification_report(test_generator.classes, y_pred,target_names=target_names, digits=4)
print(report)
# -
confusion_mtx= confusion_matrix(Y_pred_classes, Y_true)
plt.figure(figsize = (6.5,4.5))
plot_confusion_matrix(confusion_mtx, classes = ['Road', 'No Road'], cmap='Blues')
from sklearn.metrics import roc_auc_score
roc_auc_score = roc_auc_score(test_generator.classes, predictions)
roc_auc_score
|
RECONOCIMIENTO/Evaluacion/Testing1024.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="6htBPnT8tYS3"
# !pip install transformers[sentencepiece] datasets
# !pip install accelerate cloud-tpu-client==0.10 torch==1.9.0 https://storage.googleapis.com/tpu-pytorch/wheels/torch_xla-1.9-cp37-cp37m-linux_x86_64.whl
# !pip install git+https://github.com/huggingface/accelerate
# + id="1u9XNcsl2QX2" colab={"base_uri": "https://localhost:8080/"} outputId="da77c1e8-36a6-4cfe-a44b-5e31a1c9dbe6"
import torch
import torch.optim as optim
from torch.utils.data import DataLoader
from accelerate import Accelerator, DistributedType
from datasets import load_dataset, load_metric
from transformers import (
AutoTokenizer,
get_scheduler,
set_seed,
)
from tqdm.auto import tqdm
import datasets
import transformers
# + id="fq1tYS5RAY-C" colab={"base_uri": "https://localhost:8080/", "height": 481, "referenced_widgets": ["3a05f806388e4225aa1ff791fbe414a9", "f927540235fd453f8334476739f61dbe", "3f2ce719d61b4543b85a9e0d3ea20f58", "75a3e9cac2b84bce8f1e83ab6bb956ea", "b0bde2611c034bb4a3b186de29febad4", "<KEY>", "fc8d6622c6824ceca8443cb1158e1f1b", "f9b67a70a9ff4e1aad7c831e51be42ab", "<KEY>", "<KEY>", "b20ad301841b403c91341a0646f414d7", "3e30822ee1434f6c87803a0167437506", "d4feae3df403454dae806a3f6790fd7a", "<KEY>", "e5c95a52cf764880af5f84c345a10798", "e41c8b5c2ed94950a743a23d78b78459", "<KEY>", "9fa5582b703b48e2857b07a3f64d9346", "94705141ebed4bc293d006866398178e", "8a406679dd9b4967ae0a66763b2d502f", "<KEY>", "d1b5595811f34d46a00c3194c6a3073c", "<KEY>", "6c2dda088eca434ca97180c11253bbe5", "006c3a63879c4a6d8ab2a89a74836bac", "07b441a9f4674ec1a4cea785956e0634", "c81dcc10c5484e3b880b498d18862edb", "<KEY>", "<KEY>", "ff0af4b2a4654d51bdb118835cadfcb3", "161f2bd821dc4adda156b354ec82132f", "58f8f8f752e042d590b49a2ac7ff3df8", "67da1001dd30413e922e0ce63da368b6", "<KEY>", "4d50c6aadeea42bcaecfa375ba9afe5e", "<KEY>", "<KEY>", "<KEY>", "20629c856e804480b24a44ffbfca84df", "39f662a12acf45adb240561a90106fa9", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "067730585b5044ebadc12af50259b602", "19aec383178144fcaf4a62c05772aff1", "<KEY>", "d2ff5fc73ad54fe4a6bbf455a8c577cd", "f683fcc717ea47f3908f1a0a5f4236c0", "7a901c481c744869a2772efe38e01992", "58c89a47b42546e4939e719987cf32b4", "3869a16601034f3a99b5a55de966b9a6", "165ead0b6155471e90f2cc9516845dee", "f647622ab43f49e58689095d037d22b5", "<KEY>", "<KEY>", "aad3afc9aa144fe0a0e853799404a824", "da8004562ca0489c81e096da1c92c6bd", "95a87ca1e16b456085964c3e39d6236a", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "5776a83376334baf86afe7987778052f", "73016be79dea422789dc7de7f2925d12", "97d66a7d6a7c40ae93b963f81b60e195", "967ffdc7e6dd4d618a6148589eddd385", "1edce7e7ba484a0598d47bd7e36fae21", "<KEY>", "ed5e4c8bbcc24db98deae8ed0372edae", "44f3aee9475c4cd895ea5625c63421a0", "<KEY>", "6d0529a5da39439da504f3f7d2bdde29", "6073407dd5984edb8777a7071ad4bf6f", "<KEY>", "<KEY>", "9772d09816544efb8d024a7fd3e87201", "<KEY>", "a0f98154f94b435091802a60f144ea8e", "cd3fcb274ae74ad8acd9ce497fa1b99e", "1c3751a970db4c3e9af500fb9fdc1622", "<KEY>", "<KEY>", "e9323c2bfbe4454f9109e0ac19e97393", "<KEY>", "<KEY>", "ede220c06a8140e38133ed70efd63c83", "<KEY>", "9f0718047fbe4ba8843f23ff2eefe279", "e8d8b23d330d4cd48a69bab4caba4c51", "<KEY>", "12292d1864214b55a760905240f5448d", "c198392d2091475da1cf38750cc9b09c", "<KEY>", "de61e99014e3423399600240720dab26", "3315685e66334357a765dbc87c3ed76d", "<KEY>", "455945cfdef048548b7e8ea1d5fda954", "aab8ec026fb1417d911d530ee52c64ab"]} outputId="d720320a-c574-4656-f9ca-7f61e333d579"
raw_dataset = load_dataset("squad")
raw_dataset
# + [markdown] id="00hcrwB_eW9a"
# # View samples
# + id="7aM7dLzQtUQy" colab={"base_uri": "https://localhost:8080/"} outputId="c3039d8f-c910-4eb5-f8c7-c1f2c912c6a1"
print("Context: ", raw_dataset["train"][0]["context"])
print("Question: ", raw_dataset["train"][0]["question"])
print("Answer: ", raw_dataset["train"][0]["answers"])
# + id="2iFis6sXtgPx" colab={"base_uri": "https://localhost:8080/"} outputId="3e5e8e9d-636c-4ddd-bd08-88d17a70133f"
print(raw_dataset["validation"][0]["answers"])
print(raw_dataset["validation"][2]["answers"])
# + id="jTGdiwBftsyR" colab={"base_uri": "https://localhost:8080/"} outputId="e863a483-68e4-4a56-d6cb-d8546a60c77a"
print(raw_dataset["validation"][2]["context"])
print(raw_dataset["validation"][2]["question"])
# + id="YKazsUtBtw3f" colab={"base_uri": "https://localhost:8080/", "height": 145, "referenced_widgets": ["0b7ece0cf45c4c6a82e3a4edd0e3d0ac", "35e73f9ad5204a25a57843aa03e862c7", "7e74e101155e428d9b425d9d8f63e0ed", "45c281e3d7a8404e8ad5d90734844420", "8fee2439d0cc443faa1c9beedadf9e9f", "fbc2a8f03a384109b00d2f40b0e39ab9", "c770bd80014f40ce85daa523eeedd344", "5fce52afa28d44aaa3d37a339a9e58cf", "a2a4634f32df4c4a8be99278d0ccddab", "95b5a6898a6244bdb0f0911e98db9ae8", "c067c379b1224a699557d02de18906db", "12a27eaf62154854a6d6dd8e62cbb361", "11b52363e3054577b1bd3ad5056f13c5", "b8a871afcc17463381e1bb3af4e1fd95", "50f317f1501046aa9d31e768285cdd65", "52ad84ac6ceb45f9beb5423c4fcad005", "463680268e38472e9a8ff2ee12e39b35", "<KEY>", "<KEY>", "<KEY>", "cadab57f792c48c8be9637c0c1ed19c7", "651affe842ba48938f9a126cd03189b3", "be915295f2ee452f849854165749b467", "<KEY>", "bc6c9356391c42e39bc5e1353baf4172", "1301016b6b99451597cafe977e947e6f", "<KEY>", "<KEY>", "ec77ff5e61114f6087797815f3e50eee", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "9b0ae22815744a03a1597625041ce1c1", "<KEY>", "<KEY>", "a0819f48928b4374aed7ce11dc6cda77", "<KEY>", "<KEY>", "47f28168ed22450eaefe9cf780a2092d", "9992f830d6ce4ce8a3eb3f75870ff53f", "35b1db2320e14e63bbacef3052625dc0", "<KEY>", "<KEY>"]} outputId="ce570dd8-ce39-4366-e46d-ef8d34a56fde"
from transformers import AutoTokenizer
model_checkpoint = "bert-base-cased"
tokenizer = AutoTokenizer.from_pretrained(model_checkpoint)
# + id="1AmZVcC6uJ9z" colab={"base_uri": "https://localhost:8080/", "height": 140} outputId="32ab6986-60af-4658-b449-5c9071245c65"
context = raw_dataset["train"][0]["context"]
question = raw_dataset["train"][0]["question"]
inputs = tokenizer(question, context)
tokenizer.decode(inputs["input_ids"])
# + [markdown] id="KffUXTKWedQO"
# # Preprocessing
# + id="m6sGCKFAuwyn"
max_length = 384
stride = 128
def preprocess_training_examples(examples):
questions = [q.strip() for q in examples["question"]]
inputs = tokenizer(
questions,
examples["context"],
max_length=max_length,
truncation="only_second",
stride=stride,
return_overflowing_tokens=True,
return_offsets_mapping=True,
padding="max_length",
)
offset_mapping = inputs.pop("offset_mapping")
sample_map = inputs.pop("overflow_to_sample_mapping")
answers = examples["answers"]
start_positions = []
end_positions = []
for i, offset in enumerate(offset_mapping):
sample_idx = sample_map[i]
answer = answers[sample_idx]
start_char = answer["answer_start"][0]
end_char = answer["answer_start"][0] + len(answer["text"][0])
sequence_ids = inputs.sequence_ids(i)
# Find the start and end of the context
idx = 0
while sequence_ids[idx] != 1:
idx += 1
context_start = idx
while sequence_ids[idx] == 1:
idx += 1
context_end = idx - 1
# If the answer is not fully inside the context, label is (0, 0)
if offset[context_start][0] > start_char or offset[context_end][1] < end_char:
start_positions.append(0)
end_positions.append(0)
else:
# Otherwise it's the start and end token positions
idx = context_start
while idx <= context_end and offset[idx][0] <= start_char:
idx += 1
start_positions.append(idx - 1)
idx = context_end
while idx >= context_start and offset[idx][1] >= end_char:
idx -= 1
end_positions.append(idx + 1)
inputs["start_positions"] = start_positions
inputs["end_positions"] = end_positions
return inputs
# + id="M5tEI_BLu8Am" colab={"base_uri": "https://localhost:8080/", "height": 67, "referenced_widgets": ["f1660ec373084cd084422679a59c9142", "<KEY>", "b4363b1ee5d34d2fb364a1b1fbec0bcc", "ee8725cfec1c44a099da9c46f97f787f", "<KEY>", "<KEY>", "<KEY>", "70c45ad127b44e83992fb4710eda12a9", "50b54e5e20f84aab9638308ead8da399", "<KEY>", "7a300c189afa43d89166a8a0e43a64f5"]} outputId="2d173f50-2501-4d6f-c6d0-7747bbd21483"
train_dataset = raw_dataset["train"].map(
preprocess_training_examples,
batched=True,
remove_columns=raw_dataset["train"].column_names,
)
len(raw_dataset["train"]), len(train_dataset)
# + id="3VQmYkCevI6N"
def preprocess_validation_examples(examples):
questions = [q.strip() for q in examples["question"]]
inputs = tokenizer(
questions,
examples["context"],
max_length=max_length,
truncation="only_second",
stride=stride,
return_overflowing_tokens=True,
return_offsets_mapping=True,
padding="max_length",
)
sample_map = inputs.pop("overflow_to_sample_mapping")
example_ids = []
for i in range(len(inputs["input_ids"])):
sample_idx = sample_map[i]
example_ids.append(examples["id"][sample_idx])
sequence_ids = inputs.sequence_ids(i)
offset = inputs["offset_mapping"][i]
inputs["offset_mapping"][i] = [
o if sequence_ids[k] == 1 else None for k, o in enumerate(offset)
]
inputs["example_id"] = example_ids
return inputs
# + id="oQzdAIXqvKFy" colab={"base_uri": "https://localhost:8080/", "height": 67, "referenced_widgets": ["242526d050c142b28c9152d154438621", "5c923e1031224681b604afef92216bba", "f2ca3a1f0c5040a88303d3b6906ab7f9", "<KEY>", "baeede144c434f60a175465e8d586995", "e4121f45dd5f4ab3bf1636d55d09a722", "a6d5b07209914565891452351429be9c", "9fb005456ec04853a0e0e1ff349c087e", "1442ff9d60b94fafa920d4a890184e09", "aee5158149e941ab887adfce119af017", "ed7423bca19648f287c4131223e1c8a6"]} outputId="32adc998-b244-4d46-aa37-67625ab319fb"
validation_dataset = raw_dataset["validation"].map(
preprocess_validation_examples,
batched=True,
remove_columns=raw_dataset["validation"].column_names,
)
len(raw_dataset["validation"]), len(validation_dataset)
# + id="tzfCxJcIes4_" colab={"base_uri": "https://localhost:8080/", "height": 81, "referenced_widgets": ["1086deddc0804dfc977ac4716db9e201", "1e7b032edebb4c72b80e4ba8b25c181b", "fc26777c71b747e3808128cfb7029687", "bbd5b9d122304da3aca1e7cc29fae60a", "91e3380e91a94d6895efd88ce51867a4", "<KEY>", "41e0be9f4fcf4a3fa78024488fd537a0", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "895594aa9cea48c7adb287e03f284401", "9e15ce179a6b4554ba036122fa6ad4a1", "<KEY>", "470cbc8e4adb4434827a659517c5163d", "cff3952a3b7246c69d96a927168e868e"]} outputId="348becd5-ab70-4e3f-9d04-ccd737197581"
metric = load_metric("squad")
# + id="evC6T6MXdrQp"
import numpy as np
import collections
n_best = 20
max_answer_length = 30
def compute_metrics(start_logits, end_logits, features, examples):
example_to_features = collections.defaultdict(list)
for idx, feature in enumerate(features):
example_to_features[feature["example_id"]].append(idx)
predicted_answers = []
for example in tqdm(examples):
example_id = example["id"]
context = example["context"]
answers = []
# Loop through all features associated with that example
for feature_index in example_to_features[example_id]:
start_logit = start_logits[feature_index]
end_logit = end_logits[feature_index]
offsets = features[feature_index]["offset_mapping"]
start_indexes = np.argsort(start_logit)[-1 : -n_best - 1 : -1].tolist()
end_indexes = np.argsort(end_logit)[-1 : -n_best - 1 : -1].tolist()
for start_index in start_indexes:
for end_index in end_indexes:
# Skip answers that are not fully in the context
if offsets[start_index] is None or offsets[end_index] is None:
continue
# Skip answers with a length that is either < 0 or > max_answer_length
if (
end_index < start_index
or end_index - start_index + 1 > max_answer_length
):
continue
answer = {
"text": context[offsets[start_index][0] : offsets[end_index][1]],
"logit_score": start_logit[start_index] + end_logit[end_index],
}
answers.append(answer)
# Select the answer with the best score
if len(answers) > 0:
best_answer = max(answers, key=lambda x: x["logit_score"])
predicted_answers.append(
{"id": example_id, "prediction_text": best_answer["text"]}
)
else:
predicted_answers.append({"id": example_id, "prediction_text": ""})
theoretical_answers = [{"id": ex["id"], "answers": ex["answers"]} for ex in examples]
return metric.compute(predictions=predicted_answers, references=theoretical_answers)
# + [markdown] id="UGmn4VhJeiWB"
# # Testing evaluation scripts
# + colab={"base_uri": "https://localhost:8080/", "height": 177, "referenced_widgets": ["a5b632c7d3594562b0dd55278d864ff9", "198159a3766443e49180f430f67c628a", "fb6b4d93c2d04894871fe1160e30f1b4", "3942a610318f4a2faed742429f0435cd", "abcd8d7d703e40ec9d6e0c1e8092f285", "62b2372371294491b74bad23f995a0b6", "<KEY>", "7267e7aff52544aebed32729d2e2e7e2", "74c55526272746c3871710c7a15bb2f0", "331aa0175c864fbca86d4ddf3598ed48", "f191dc1157414eaca6cd86c9cf5be1b4", "7a86c1ff82a141ffaedfa562a45269c5", "4828564dab6440b49426b8ad57effef5", "715c4a3bca70482fa44e72ae8f0704db", "<KEY>", "1df941f755c143e79cd3ab717cc0ce69", "<KEY>", "a91faeeb266845a5aff2ea9f25317276", "<KEY>", "<KEY>", "5c33a72bcac64b3d9ea275b7e016de4f", "<KEY>", "<KEY>", "0db65adf576347839f5d0524bb6e020b", "<KEY>", "bf07dda3e1ca475b8524944e4ae50f75", "1105a3ef6b6e4c16b7502e22ef715b28", "a78f852bc6e84bea82277847850c79a8", "0d6fbf534d7844d5b320ea327db44cca", "1a0adcf998fc46e493b2a6326635f24e", "015fe554ee004b059c136e46dde46f49", "68622e7a73c24fefad666df301ab89ff", "7c0cc469beb942a082ff19c6714ea787", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "c9e34da5d05d4d8e9cfcffb0f08c11e6", "<KEY>", "f476b2d0ade9487d9a6243d1bb92a5bc", "6ca1989018d346bd99de61a3c624ae6e", "c5263a5d2fb84980a4f863ad8f276c62", "865f9d0b18a64cac879c0c993d343401", "d4441262624e48eea1fda7092fe24236", "<KEY>", "259289674338457ba715432ee373a8f7", "<KEY>", "<KEY>", "f28147a1a1f14d47b10e6f900702e468", "8c485cece5164c0f80f48fefdb089ecf", "<KEY>", "<KEY>", "f4e7d852e3854d7192352eec557c8e03", "<KEY>", "<KEY>"]} id="74FW-ZWmw1jM" outputId="4c9b4c4a-d05a-465c-9659-728381e9b904"
small_eval_set = raw_dataset["validation"].select(range(100))
trained_checkpoint = "distilbert-base-cased-distilled-squad"
tokenizer = AutoTokenizer.from_pretrained(trained_checkpoint)
eval_set = small_eval_set.map(
preprocess_validation_examples,
batched=True,
remove_columns=raw_dataset["validation"].column_names,
)
# + id="2pMqolhAgvZL"
import torch
from transformers import AutoModelForQuestionAnswering
# + colab={"base_uri": "https://localhost:8080/", "height": 49, "referenced_widgets": ["6f648663ec784808a477178894da6a39", "dc5bcba8d51943ada6eaa09e17bb508c", "84edb3ef1edd438d985a4dfe14e3b5c1", "<KEY>", "ce43709a596140789a9f08e9ed1e7dca", "b5ac43abfdb74e07a7f7883765392c72", "<KEY>", "d8235e793bc2400d99e0e27811d8beda", "e7bce21cc2c84e1f8a26a097e7ebed4a", "ced92c0d8fd4462b8bfebcff1fdd6722", "7abb9a76b6bc4aba9821f406110a4190"]} id="ciRWp6nsw_fZ" outputId="e3c96c39-ce9f-43a7-8376-aa863cee6856"
eval_set_for_model = eval_set.remove_columns(["example_id", "offset_mapping"])
eval_set_for_model.set_format("torch")
device = torch.device("cuda") if torch.cuda.is_available() else torch.device("cpu")
batch = {k: eval_set_for_model[k].to(device) for k in eval_set_for_model.column_names}
trained_model = AutoModelForQuestionAnswering.from_pretrained(trained_checkpoint).to(
device
)
with torch.no_grad():
outputs = trained_model(**batch)
# + id="1DjlYKAdxQCv"
start_logits = outputs.start_logits.cpu().numpy()
end_logits = outputs.end_logits.cpu().numpy()
# + id="IVl2Vu76wqz9"
example_to_features = collections.defaultdict(list)
for idx, feature in enumerate(eval_set):
example_to_features[feature["example_id"]].append(idx)
# + id="Y-U7TDN4wKEo"
predicted_answers = []
for example in small_eval_set:
example_id = example["id"]
context = example["context"]
answers = []
for feature_index in example_to_features[example_id]:
start_logit = start_logits[feature_index]
end_logit = end_logits[feature_index]
offsets = eval_set["offset_mapping"][feature_index]
start_indexes = np.argsort(start_logit)[-1 : -n_best - 1 : -1].tolist()
end_indexes = np.argsort(end_logit)[-1 : -n_best - 1 : -1].tolist()
for start_index in start_indexes:
for end_index in end_indexes:
# Skip answers that are not fully in the context
if offsets[start_index] is None or offsets[end_index] is None:
continue
# Skip answers with a length that is either < 0 or > max_answer_length.
if (
end_index < start_index
or end_index - start_index + 1 > max_answer_length
):
continue
answers.append(
{
"text": context[offsets[start_index][0] : offsets[end_index][1]],
"logit_score": start_logit[start_index] + end_logit[end_index],
}
)
best_answer = max(answers, key=lambda x: x["logit_score"])
predicted_answers.append({"id": example_id, "prediction_text": best_answer["text"]})
# + id="47fonZFBxVJU"
theoretical_answers = [
{"id": ex["id"], "answers": ex["answers"]} for ex in small_eval_set
]
# + colab={"base_uri": "https://localhost:8080/"} id="IF-VsnGvxiiV" outputId="2ee8567f-fe98-4b21-fab7-ef75d8dab10c"
print(predicted_answers[0])
print(theoretical_answers[0])
# + colab={"base_uri": "https://localhost:8080/"} id="KIPBgIIKxWQI" outputId="1a1bdbca-5b13-44e6-acf9-43983934e807"
metric.compute(predictions=predicted_answers, references=theoretical_answers)
# + colab={"base_uri": "https://localhost:8080/", "height": 66, "referenced_widgets": ["5263f69b77e0426d95b0f21c4aa2772e", "7375fd7c08ad45b8821127cb706ab970", "3a37c777920a46fe976f022fce9bbecf", "8c036af0d1904281a3e6c7edadf6d08b", "e3e3619692ba4346a2916f306596de0f", "5bb7b060594b45ca89179513ff696d99", "b4ae09a9af7c499387ccb10f5507c182", "92523a42db8d4ae8ad4244632ff34ded", "66bea59fa1d14557b6d182d2ea257cdd", "e657562149354627b6d9e7ff8996d0f7", "7ed64b14df7d488f8d4930e30c7f60c0"]} id="633I1OnaxpMK" outputId="f7c91806-98ba-4450-fe5b-68786bda0e0d"
compute_metrics(start_logits, end_logits, eval_set, small_eval_set)
# + [markdown] id="1SFUAhaCetJE"
# # Training
# + id="-Fn5wXjK2azN"
hyperparameters = {
"learning_rate": 2e-5,
"num_epochs": 3,
"train_batch_size": 8, # Actual batch size will this x 8
"eval_batch_size": 32, # Actual batch size will this x 8
"seed": 42,
}
# + id="lMWD2sJL2jXC"
from torch.optim import AdamW
from torch.utils.data import DataLoader
from transformers import default_data_collator
# + id="5fdg78NC22UD"
def training_function():
# Initialize accelerator
accelerator = Accelerator()
# To have only one message (and not 8) per logs of Transformers or Datasets, we set the logging verbosity
# to INFO for the main process only.
if accelerator.is_main_process:
datasets.utils.logging.set_verbosity_warning()
transformers.utils.logging.set_verbosity_info()
else:
datasets.utils.logging.set_verbosity_error()
transformers.utils.logging.set_verbosity_error()
train_dataset.set_format("torch")
validation_set = validation_dataset.remove_columns(["example_id", "offset_mapping"])
validation_set.set_format("torch")
train_dataloader = DataLoader(
train_dataset,
shuffle=True,
collate_fn=default_data_collator,
batch_size=hyperparameters["train_batch_size"],
)
eval_dataloader = DataLoader(
validation_set,
collate_fn=default_data_collator,
batch_size=hyperparameters["eval_batch_size"],
)
# The seed need to be set before we instantiate the model, as it will determine the random head.
set_seed(hyperparameters["seed"])
# Instantiate the model, let Accelerate handle the device placement.
model = AutoModelForQuestionAnswering.from_pretrained(model_checkpoint)
# Instantiate optimizer
optimizer = optim.AdamW(params=model.parameters(), lr=hyperparameters["learning_rate"])
# Prepare everything
# There is no specific order to remember, we just need to unpack the objects in the same order we gave them to the
# prepare method.
model, optimizer, train_dataloader, eval_dataloader = accelerator.prepare(
model, optimizer, train_dataloader, eval_dataloader
)
num_epochs = hyperparameters["num_epochs"]
# Instantiate learning rate scheduler after preparing the training dataloader as the prepare method
# may change its length.
num_train_epochs = 3
num_update_steps_per_epoch = len(train_dataloader)
num_training_steps = num_train_epochs * num_update_steps_per_epoch
lr_scheduler = get_scheduler(
"linear",
optimizer=optimizer,
num_warmup_steps=0,
num_training_steps=num_training_steps,
)
# Instantiate a progress bar to keep track of training. Note that we only enable it on the main
# process to avoid having 8 progress bars.
progress_bar = tqdm(range(num_training_steps), disable=not accelerator.is_main_process)
# Now we train the model
for epoch in range(num_epochs):
model.train()
for step, batch in enumerate(train_dataloader):
outputs = model(**batch)
loss = outputs.loss
accelerator.backward(loss)
optimizer.step()
lr_scheduler.step()
optimizer.zero_grad()
progress_bar.update(1)
model.eval()
start_logits = []
end_logits = []
accelerator.print("Evaluation!")
for batch in tqdm(eval_dataloader, disable=not accelerator.is_main_process):
with torch.no_grad():
outputs = model(**batch)
start_logits.append(accelerator.gather(outputs.start_logits).cpu().numpy())
end_logits.append(accelerator.gather(outputs.end_logits).cpu().numpy())
start_logits = np.concatenate(start_logits)
end_logits = np.concatenate(end_logits)
start_logits = start_logits[: len(validation_dataset)]
end_logits = end_logits[: len(validation_dataset)]
metrics = compute_metrics(
start_logits, end_logits, validation_dataset, raw_dataset["validation"]
)
accelerator.print(f"epoch {epoch}:", metrics)
# Save and upload
accelerator.wait_for_everyone()
unwrapped_model = accelerator.unwrap_model(model)
unwrapped_model.save_pretrained("qa_bert", save_function=accelerator.save)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000, "referenced_widgets": ["f3437963740146f1adf170f968b55d41", "e3bc35d25fc44ed1939860f6c5aedef2", "a321c81233a04595a2b25679c76b8b65", "035d46433d5f48f1893386b4d37e2ee6", "0f47872aaa664cc6a5413c02832813d0", "7cb45de93c4748ff9e50c8c736dfb84f", "5dad745ff30f4556b3f73d9c3b4d29c8", "a470f8ddb0ad4f20ad9e9897d8409a86", "c0f6ad4356444ea7852d488e8a03db7f", "048ffd210d564f9f9919afd534fa527c", "e2e2dd177b934535850ca184c7b85b71", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "8b9db0ebc1cb4ab095a4270857f1e3d3", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "e611f561777c4938871d8e6d123d9a70", "06de1adddf114fbf96f470ca782a826c", "4cd726b051cc47ed9923b2ee6accc75d", "b4dda7d57aa44dd39c725b5e05981327", "<KEY>", "<KEY>", "<KEY>", "08de96e272994610b4d729eaf5c6f91a", "a0e0394172d44a7bae068d38d45fbf7e", "7a7f1937d1104187a7e24dd80240d144", "<KEY>", "d37be048591c4b6b9ce56b3e42b75057", "612c4577138d43ec85a297af42ba02b2", "<KEY>", "<KEY>", "<KEY>", "937c28a4c93a47888ad619e72b2ffdb0", "934b23f57e7240ed8484bfe54a01f44e", "<KEY>", "ba1a2c06b1eb4e07aaa23ddaff1560e2", "<KEY>", "<KEY>", "<KEY>", "73de50a45a42401b879abac13d9df1b4", "a746b372ef3f47a5a17fbd57d378d730", "<KEY>", "<KEY>", "<KEY>", "8e609ee1d3cc4e67b9bf8834e86ca2e6", "fce16be02ba8440ab061aa1504e2f570", "<KEY>", "<KEY>", "<KEY>", "0e6432d4b4044347ba08d07f85dff546", "<KEY>", "<KEY>", "037db94530b44307aebde0de3fde0195", "<KEY>", "39614277172c4549a30864c865681bef", "<KEY>", "<KEY>", "3d4134bec8234e1abae160210c108575", "<KEY>", "5d0dd457cda8414aa3d75a09f889be9c", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "fd35ed5c6e594076a7f40a6c519a047f", "a85e0d071ece42fbb27a98e328d48e3e", "bec9f3f6299f40b49e491de3dae19062", "<KEY>", "<KEY>", "ee040e8254094721ada2bcd77c41ec1d", "80ff110dd08641dead35a37c16dc179e", "<KEY>", "dd75a51e6a174c7eae1e8b8012a694e6", "141200f8cc334eaab26a240620e9381e", "<KEY>", "<KEY>", "ca8faaddfcba4d63bd178be4be9739ad", "<KEY>", "e6f5e6d427eb47eb978af938ffd67190", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "12d7255622b74e12b74aaa90bd08ef4c", "<KEY>", "9214c01ceaf04f54925915ff34637003", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "45a32464e3a043818dc757274692fa8d", "cfb002bed55242829ff220ec61ca4e10", "<KEY>", "ebd2f19df4ea4923b6ddf6fb1baede12", "9ebe597df77149468fd7f413e8ede703", "45113ced3e6b409cb0edcf5fd9e90dc3", "b03e4a5b0dbc4024a9b7e2b9b56afb16", "<KEY>", "<KEY>", "6ce7da7cbe5a4eeabbb99d3f5aba5bcf", "<KEY>", "8c45af61e7924f5ba9e8d36a70acab87", "<KEY>", "50fdffda4df343a486209dde45823145", "<KEY>", "964f16d44d3e43988f8efd8514a2cc46", "<KEY>", "<KEY>", "<KEY>", "e9f66fa925f04b3992f4f221d385d64e", "<KEY>", "81878475a2534a9e8f1856abe15e4880", "<KEY>", "<KEY>", "<KEY>", "acbd4a6e9ea141f08bad004431f3f47f", "e48892dd3ca8484bb2b0a29489c17449", "<KEY>", "<KEY>", "<KEY>", "076bf98634464d43a1a16021be7a61d3", "<KEY>", "e7654a28d895476db0f838ec16c2dcac", "07d99a8a15c44408a642b5d357c58921", "<KEY>", "da6333961eda4462b96dc4e6aeec72eb", "5b4ed58fd50f4b2086a2747f6ad5a3dc", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "650c5a3de7b647b9be30e59e53865d4b", "2750baad54bf495882e38652c506a2b6", "<KEY>", "<KEY>", "15bf79427d254dd1851e5d2aba90ad76", "5c947145cffa40f389f6c512dc05e65f", "<KEY>", "91153ecf80c44a00bde649adb2aedfa2", "277702c9a5db4da1b4efe3af7c690f97", "b83380f899ce4dd0b628fce4d29b7bbe", "6d2f59d06cd74030b676df0a7c08e440", "d6746d47859b4c57ad2b0217a3f8730a", "<KEY>", "<KEY>", "db3b6658ba084ec883af099fbe393904", "<KEY>", "f01efd6d17d94ed2846e11b55d3d927e", "a683099264fb497694274af7ee43ca57", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "1f1c276a8c4e4c0584e1f7da7c6ae855", "9052779c448a43d994a7ae89f8b58476", "46b133fd40f047c1a26a2c22579ce7db", "<KEY>", "fd3494d5eddf43d6b655fa45af419fa9", "<KEY>", "44542f9a6e1b48d1acae446f1f6b17ad", "e51ed5d6d51448df9d4b5e6c06581d05", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "025e8e9b88d14c2b9a09c96a02ec1e86", "fe14d59004324b9faf44c20e6a8d3b02", "<KEY>", "<KEY>", "<KEY>", "478a3110ff8549a5b1d66e56276ec58d", "<KEY>", "c5fc77b2a8f74ff480eee3b512ac3a59", "85e7315fa2ce4ac39e9d6e974960b7ad", "<KEY>", "<KEY>", "7d86e7d2f0754dd1ab3eeb1d7c1ebdfe", "c7c7d3083322447f9b48c7ed79eb8837", "<KEY>", "<KEY>", "874529809666470da7588d4d24af5c8a", "648a00dfb08e4b1ca2861832dc06e48f", "e1ee3c9e06b94b72afef019c7cef61b4", "<KEY>", "404e5d1b010b4a76bd33e665e5e2d69b", "<KEY>", "1ed3e33dc63d4a7985dd69f46289c0a9", "<KEY>", "9accfe111dce4e80b0d0fba96be7347f", "<KEY>", "<KEY>", "26ba9489366845458cfb1748de856ce1", "d5252855b7d047f78c5e80d421e0ab8f", "eafe2a0ef1d7456e9f8fe4cbfa2bb16b", "<KEY>", "77c40ddace5d41c7b8e2fa7b100fbe36", "<KEY>", "<KEY>", "d0f515fc39d244c2ac314faab54b560e", "<KEY>", "<KEY>", "d46859fed3e64b5ea7ebcdf7dae08cce", "<KEY>", "4b8e5c8c01c342b88a3888096b00de37", "7b6a7b30e7b345a9bc9b4f3574a57bea", "<KEY>", "934acc399c024ffa9eaaea476de7a9f4", "704f51ed02d54660a2cb70948e1c3b57", "2c5ed35572de491bb1b93d944950aefd", "<KEY>", "53d8b082a27742f8af100e26272d51b1", "<KEY>", "<KEY>", "788ead8f27fe402bb7be094b5627aba3", "<KEY>", "<KEY>", "<KEY>", "5c369f22fa424a9b9ab9b6ee27e7ef02", "6b8e79276a1345348631d04d42aeb1ba", "<KEY>", "<KEY>", "a7b4d0c9bb794e1181aff22ee747a7f1", "f1e849a7760e44bdad2e7a180008e9f3", "d2c65ab3626041eaae9c1fefa4e9f83a", "<KEY>", "935d0f6e988a4d408b9831884669c86a", "<KEY>", "<KEY>", "2a064ae36dbc45e9b4a58816f9a494b4", "93dd58f3c9174a2fa489086eca91c771", "e088cb0ff1f94ebebe2e6c373dfe68e7", "<KEY>", "<KEY>", "8f57409f31aa4a3d90574ffb932c7a41", "f976152c3a32478bbe4a51b4e06c2245", "<KEY>", "299b31a027714c1ca9366f12195e21c4", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "eeae199c7be046b7a51d686f8d712bb7", "<KEY>", "3670daa8900640f0b9f9be366c4af18a", "1150159b4fd44bd68e318dd65d873ee7", "2d9c18d6e8034581934924e6970e0ff0", "96d493a52583484f9a2b57e0f27b02ca", "<KEY>", "dac61efd477f4114a719e2b37977a937", "97de0063eca84e469971cec49e79bde3", "16be496e14d74d90a1c7e9ac1e412551", "<KEY>", "808e6e6ba086447caa4a50f1ca1c4d60", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "f984aac097ef4affa8b1d4db46622ccf", "<KEY>", "6364c9d1990642ed826b7de32f93f36e", "<KEY>", "f9daf18dc0d84c3e97527a3c17dce602", "ef4eff42d424465caa77d8a53a256e10", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "db6815701a2c4b1e903dc7a23d7b1f26", "3d40fc6479cb4ad788ddd22fffe847d3", "bd5dfa6ba45d489283c0e4b9820ffdbb", "21db8803910343bab93eb8cea6388b13", "52622c6adc2d4f46a1269166356d8507", "dea194982d5345eb921747075f9a8bee", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "58474e075a924315928587a4f78081ff", "0a566c9aafd942b29c308e09010482e7", "7ddcea6b27b443ee8c60015cfae58d00", "cc7e095988e4473880860f3f5df9e849", "87bb1cea22a64443b3ed2fe783b73921", "8d70988f4c114cabb9fba55dc4a969f0", "c602e99b43ad4c668a5a1905f769d006", "99a25fb06503486186addc08fc9ea594", "<KEY>", "9e76a63f7c68453aafd07f5c9c9fade3", "<KEY>", "009e6306dd024d25836876ad4d56cf35", "95e269d70f0f4339ad0269c79f083393", "<KEY>", "<KEY>", "050cf44b498e4652933602f6b254310a", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "da93733b2a134f3f829d15e5c1db90a0", "<KEY>", "<KEY>", "64b56e762aa94ad19940bd7ad1995684", "<KEY>", "<KEY>", "a0781ccdc7a94f8283f38360a7ca06ba", "<KEY>", "d8b0eeebd1154e5b87b1f93e1a559dc6", "d30c0086c091428ea582e4549549d60a", "7e5db675c2df431db4195d5bee2a498c", "b2f9e6aa3e214932a2d612ee745355e8", "28efbf9aca274c318ef89e76afb4dc9d", "bb3fb48959e14ce3be6acd9f18e6c0fc", "<KEY>", "ab8df52823a74e0d8f334e77877590f3", "<KEY>", "<KEY>", "<KEY>", "ff2b009664c94aae8c46e783203f4714", "<KEY>", "<KEY>", "<KEY>", "3cf43e6cecd449658bd37d6ce8886e1b", "<KEY>", "<KEY>", "<KEY>", "a31adaeff2fe4e69a34ec290e0b29fbb", "f91d5930e99f4180bca1804c5864a4b6", "caf89ab45c4e4c31ab4530e13d2d5e7c", "2a66dc52e4854a06b21d04e43aba8a83", "557149776f3a488f9ff0fe5f829cc71b", "<KEY>", "fe4e35e1391c41199dabdd60b9a4097b", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "eecabc90da4848c0a9f6651cdb978e33", "<KEY>", "<KEY>", "e29f1572544140cfb75042f28e783392", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "cce71361b1f0486c832dd008e1ee977a", "<KEY>", "a6b9cdd3aa364a5f9b8bed218eaf5a18", "79a88e866d4f4a3381de9ea1b12fced2", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "2904d059300447f0b9890a6d9aa96df5", "<KEY>", "<KEY>", "<KEY>", "e94d6ef1cb55493ea231ba2ed3e53aa1", "<KEY>", "5b489c052a8c4efab6642209c2cd0d35", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "d69064ba7c0a45b59979d39ea17c29ea", "710691e6c47a41a38c037a4cdfe469b8", "1894874d258040aaba44f2a588606610", "<KEY>", "<KEY>", "9137e0d30ff14a5f8016ceaf429cd007", "fe253f74e7fb480383da7ec03438e038", "4c49a83869bb4a509c66892e6edd928b", "c1afa8fa3daa4cc6a1f7aa66c8c21920", "2ae9fa3138f7431fa3b78a5ab60d6c84", "ccb6e6cc882e47e4add4271e7970c2ee", "f6075dcebed34d62a2b510c16e74acf1", "0d61d6833e0c47a586756c64ca481ce4", "<KEY>", "<KEY>", "<KEY>", "5fb4ef60758c4a68bff814aedfc16928", "49096c218b0a4605a7dd5b91e08b0e89", "7e259a038aab486986cc813e537cab12", "<KEY>", "b18209c916024e39983e281483e858f3", "<KEY>", "<KEY>", "<KEY>", "<KEY>"]} id="EBXLbJDSue4A" outputId="9199fbec-7d04-4637-ef85-d0c4ecbe77b2"
from accelerate import notebook_launcher
notebook_launcher(training_function)
# + colab={"base_uri": "https://localhost:8080/"} id="MbRE-Mvidu_9" outputId="97ad5d75-661f-41eb-ddca-3dd89298f742"
tokenizer.save_pretrained("qa_bert")
# + id="u_nH9yb0d9JG" colab={"base_uri": "https://localhost:8080/"} outputId="54dfdcf0-5247-46c7-d4a4-76db325beb94"
from transformers import pipeline
# Replace this with your own checkpoint
model_checkpoint = "qa_bert"
question_answerer = pipeline("question-answering", model=model_checkpoint)
context = """
🤗 Transformers is backed by the three most popular deep learning libraries — Jax, PyTorch and TensorFlow — with a seamless integration
between them. It's straightforward to train your models with one before loading them for inference with the other.
"""
question = "Which deep learning libraries back 🤗 Transformers?"
question_answerer(question=question, context=context)
|
huggingface_course/extractive_QA.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Jupyter Notebooks
# Este formato de arquivo facilita a combinação perfeita de texto e código. O texto pode ser simples ou formatado com [Markdown](https://daringfireball.net/projects/markdown/). O código pode ser escrito em mais de 40 linguagens, incluindo Python, R e Scala. Mais importante, o código pode ser alterado quando o notebook é aberto em um servidor local (isto é, no seu computador). Alternativamente, ele pode ser visualizado simplesmente por meio de um repositório do github ou do site `nbviewer`.
#
# ## Google Colab
# O [Google Colaboratory](https://colab.research.google.com/), ou simplesmente Colab, é um serviço de nuvem gratuito hospedado pelo Google para incentivar a pesquisa em Aprendizado de Máquina e Inteligência Artificial.
#
# Sua interface é similar ao jupyter notebooks, sendo composto por uma lista de células que podem conter textos explicativos (Markdown) ou códigos executáveis e suas respectivas saídas. Ao entrar no site, você já é apresentado a um Notebook de exemplo, semelhante a este:
#
# 
#
# Esse notebook explica algumas características do Colab e mostra como dar os primeiros passos (recomendo a leitura). Algumas das principais características do Colab são:
#
# * Como roda em uma máquina do google, não precisamos realizar qualquer configuração.
# * É simples de compartilhar, igual a qualquer arquivo contido no drive.
#
# Para criar seu próprio notebook, clique em >File, na parte superior esquerda, logo em seguida em >New Notebook.
#
# ### Escolha a sintaxe de uma célula
# Notebooks são construídos de células. As células podem ter vários formatos internos, incluindo *código* e *Markdown* para texto. Você pode selecionar o formato desejado em um menu suspenso na parte superior.
#
# Se você quiser digitar palavras, use "Markdown"; Se você quiser escrever código, escolha "código".
#
# ### Mover entre as células
# Para executar uma determinada célula, digite [shift-enter], ativo nessa célula. Você pode executar todas as células com Cell> Run all; outras variações estão disponíveis nesse menu suspenso.
1+1
|
aulas/00_Intro.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Documento para creación de ploteos para documento de conclusiones de ensayos dinámicos
# * Autor: <NAME>
# * Fecha inicio: 12/3/2020
# * Fecha última modificación: 12/3/2020
# +
# Inicialización
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
from scipy.signal import find_peaks
# +
# Definiciones de funciones
def fourier_spectrum( nsamples, data, deltat, logdb, power, rms ):
"""Given nsamples of real voltage data spaced deltat seconds apart,
find the spectrum of the data (its frequency components). If logdb,
return in dBV, otherwise linear volts. If power, return the power
spectrum, otherwise the amplitude spectrum. If rms, use RMS volts,
otherwise use peak-peak volts. Also return the number of frequency
samples, the frequency sample spacing and maximum frequency. Note:
The results from this agree pretty much with my HP 3582A FFT
Spectrum Analyzer,
although that has higher dynamic range than the 8 bit scope."""
data_freq = np.fft.rfft(data * np.hanning( nsamples ))
nfreqs = data_freq.size
data_freq = data_freq / nfreqs
ascale = 4
if( rms ):
ascale = ascale / ( 2 * np.sqrt(2) )
if( power ):
spectrum = ( ascale * absolute(data_freq) )**2
if( logdb ):
spectrum = 10.0 * np.log10( spectrum )
else:
spectrum = ascale * np.absolute(data_freq)
if( logdb ):
spectrum = 20.0 * log10( spectrum )
freq_step = 1.0 / (deltat * 2 * nfreqs);
max_freq = nfreqs * freq_step
return( nfreqs, freq_step, max_freq, spectrum )
# -
# ## Apertura de archivos de ensayos
# Los archivos abiertos corresponden a la configuración de ensayos planificados. Cada uno de ellos corresponden a la siguiente configuración,
# * Js11: Banco de sensor inductivo con carga a 1kOhm
# * Js12: Banco de sensor inductivo configurado a 2kOhm
# * Js13: Banco de sensor inductivo configurado a 4,7kOhm
# * Js14: Banco de sensor inductivo configurado a 10kOhm
#
# Cada conjunto de ensayos corresponden a una masa determinada.
#
# Para el caso de este informe se relevan los siguientes archivos,
#
# | Nro ensayo | R configuración | Sensor | Masas elegida | Registro |
# |------|------|------|------|------|
# | 21 | Js11| izquierdo | 2 | Ensayo s_alim 18 CH...|
# | 22 | Js12| izquierdo | 2 | Ensayo s_alim 19 CH...|
# | 23 | Js13| izquierdo | 2 | Ensayo s_alim 20 CH...|
# | 24 | Js14| izquierdo | 2 | Ensayo s_alim 21 CH...|
#
# Definición de variables de ensayos
fecha = '08/02/2020'
rotor = 'rotor número 2'
sensor = 'izquierdo'
tipo_sensor = 'optico'
# +
# Definición de ruta de archivos de ensayos
ruta_ensayos = 'Registro osciloscopio' + '/'
Js11 = 'Ensayo s_alim 18'
Js12 = 'Ensayo s_alim 20'
Js13 = 'Ensayo s_alim 22'
Js14 = 'Ensayo s_alim 24'
canal_1 = 'CH1'
canal_2 = 'CH2'
extension = '.npz'
direccion_salida = 'Conclusiones' + '/'
extension_salida = 'png'
nombre_salida_a = 'Conclusion 5_a'
nombre_salida_b = 'Conclusion 5_b'
archivo_salida_a = direccion_salida + nombre_salida_a + '.' + extension_salida
archivo_salida_b = direccion_salida + nombre_salida_b + '.' + extension_salida
ensayo_1_1 = ruta_ensayos + Js11 + ' ' + canal_1 + extension
ensayo_1_2 = ruta_ensayos + Js11 + ' ' + canal_2 + extension
ensayo_2_1 = ruta_ensayos + Js12 + ' ' + canal_1 + extension
ensayo_2_2 = ruta_ensayos + Js12 + ' ' + canal_2 + extension
ensayo_3_1 = ruta_ensayos + Js13 + ' ' + canal_1 + extension
ensayo_3_2 = ruta_ensayos + Js13 + ' ' + canal_2 + extension
ensayo_4_1 = ruta_ensayos + Js14 + ' ' + canal_1 + extension
ensayo_4_2 = ruta_ensayos + Js14 + ' ' + canal_2 + extension
with np.load(ensayo_1_1) as archivo:
time_V1_1 = archivo['x']
V1_1 = archivo['y']
with np.load(ensayo_1_2) as archivo:
time_V1_2 = archivo['x']
V1_2 = archivo['y']
with np.load(ensayo_2_1) as archivo:
time_V2_1 = archivo['x']
V2_1 = archivo['y']
with np.load(ensayo_2_2) as archivo:
time_V2_2 = archivo['x']
V2_2 = archivo['y']
with np.load(ensayo_3_1) as archivo:
time_V3_1 = archivo['x']
V3_1 = archivo['y']
with np.load(ensayo_3_2) as archivo:
time_V3_2 = archivo['x']
V3_2 = archivo['y']
with np.load(ensayo_4_1) as archivo:
time_V4_1 = archivo['x']
V4_1 = archivo['y']
with np.load(ensayo_4_2) as archivo:
time_V4_2 = archivo['x']
V4_2 = archivo['y']
# -
# ## Descripción y elección de canales de medición
#
# Cada ensayo consta de dos mediciones realizadas con el osciloscopio rigol DS1052Z. El canal 1 representa las mediciones relevadas sobre el circuito óptico, el canal 2 representa las tensión medida sobre el sensor inductivo de la balanceadora.
#
# En esta sección se podrá decidir entre una medición y otra.
# +
# Elección entre las dos mediciones 'optica' o 'inductivo'
if (tipo_sensor == 'optico'):
tiempo_1 = time_V1_1
tiempo_2 = time_V2_1
tiempo_3 = time_V3_1
tiempo_4 = time_V4_1
V1 = V1_1
V2 = V2_1
V3 = V3_1
V4 = V4_1
elif (tipo_sensor == 'inductivo'):
tiempo_1 = time_V1_2
tiempo_2 = time_V2_2
tiempo_3 = time_V3_2
tiempo_4 = time_V4_2
V1 = V1_2
V2 = V2_2
V3 = V3_2
V4 = V4_2
# -
# ## Recorte de señales medidas
# Esta acción se hace necesaria debido a que la adquisición por parte del osciloscopio tiene en sus últimos valores un tramo de datos que no corresponden a la adquisición.
# +
# Recortador de la imagen
ini_cut = np.empty(1)
ini_cut = 0
fin_cut = np.empty(1)
#Definición de cantidad de puntos a recortar desde el final
fin_cut = V1.size - 20
V1_cort = V1[ ini_cut: fin_cut]
tiempo_1_cort = tiempo_1[ ini_cut: fin_cut ]
V2_cort = V2[ ini_cut: fin_cut]
tiempo_2_cort = tiempo_2[ ini_cut: fin_cut ]
V3_cort = V3[ ini_cut: fin_cut]
tiempo_3_cort = tiempo_3[ ini_cut: fin_cut ]
V4_cort = V4[ ini_cut: fin_cut]
tiempo_4_cort = tiempo_4[ ini_cut: fin_cut ]
# -
# ## Creción de variables para cálculo de transformada de Fourier
# El cálculo de fft de cada una de las señales medida requiere variables como cantidad de muestras y el intervalo temporal entre cada muestra.
## Creación de variables para función fourier_spectrum
# Para V1
nro_muestras_V1 = V1_cort.size
deltat_V1 = tiempo_1_cort[1] - tiempo_1_cort[0]
#Para V2
nro_muestras_V2 = V2_cort.size
deltat_V2 = tiempo_2_cort[1] - tiempo_2_cort[0]
# Para V3
nro_muestras_V3 = V3_cort.size
deltat_V3 = tiempo_3_cort[1] - tiempo_3_cort[0]
# Para V4
nro_muestras_V4 = V4_cort.size
deltat_V4 = tiempo_4_cort[1] - tiempo_4_cort[0]
# ## Cálculo de transformadas
# +
# Cálculo de transformada de fourier para V1
( nfreqs_V1, freq_step_V1, max_freq_V1, spectrum_V1 ) = fourier_spectrum( nro_muestras_V1, V1_cort, deltat_V1, False, False, True )
# Presentación de datos principales en consola del espectro de V1
print ("Freq step", freq_step_V1, "Max freq", max_freq_V1, "Freq bins",nfreqs_V1)
# Cálcula de transformada de fourier para V2
( nfreqs_V2, freq_step_V2, max_freq_V2, spectrum_V2 ) = fourier_spectrum( nro_muestras_V2, V2_cort, deltat_V2, False, False, True )
# Presentación de datos principales en consola del espectro de V2
("Freq step", freq_step_V2, "Max freq", max_freq_V2, "Freq bins", nfreqs_V2)
# Presentación de datos principales en consola del espectro de V1
print ("Freq step", freq_step_V2, "Max freq", max_freq_V2, "Freq bins",nfreqs_V2)
# Cálculo de transformada de fourier para V3
( nfreqs_V3, freq_step_V3, max_freq_V3, spectrum_V3 ) = fourier_spectrum( nro_muestras_V3, V3_cort, deltat_V3, False, False, True )
# Presentación de datos principales en consola del espectro de V1
print ("Freq step", freq_step_V3, "Max freq", max_freq_V3, "Freq bins",nfreqs_V3)
# Cálculo de transformada de fourier para V1
( nfreqs_V4, freq_step_V4, max_freq_V4, spectrum_V4 ) = fourier_spectrum( nro_muestras_V4, V4_cort, deltat_V4, False, False, True )
# Presentación de datos principales en consola del espectro de V1
print ("Freq step", freq_step_V4, "Max freq", max_freq_V4, "Freq bins",nfreqs_V4)
# -
# ## Creación de gráfico temporal de todos los ensayos
fig, axs = plt.subplots(2, 2, figsize=(15,15))
fig.suptitle('Ensayo sobre ' + rotor + ' medido en sensor ' + sensor + ' ' + tipo_sensor + ' fecha ' + fecha )
axs[0,0].plot(tiempo_1_cort, V1_cort)
axs[0,0].set_title('Tension ensayo Js11')
axs[0,0].grid(True)
#axs[0,1].set_xlim( 0, 100 )
axs[0,1].plot( tiempo_2_cort, V2_cort, 'tab:red')
axs[0,1].set_title('Tension ensayo Js12')
axs[0,1].grid(True)
axs[1,0].plot(tiempo_3_cort, V3_cort, 'tab:orange')
axs[1,0].set_title('Tension ensayo Js13')
axs[1,0].grid(True)
#axs[1,1].set_xlim( 0, 100 )
axs[1,1].plot( tiempo_4_cort, V4_cort, 'tab:green')
axs[1,1].set_title('Tension ensayo Js14')
axs[1,1].grid(True)
# ## Creación de gráfico de espectro de mediciones y representación de picos
# +
# Creción de eje de frecuencias para cada gráfico de espectro
freqs_V1 = np.arange( 0, max_freq_V1, freq_step_V1 )
freqs_V2 = np.arange( 0, max_freq_V2, freq_step_V2 )
freqs_V3 = np.arange( 0, max_freq_V3, freq_step_V3 )
freqs_V4 = np.arange( 0, max_freq_V4, freq_step_V4 )
# Acondicionamiento de vector de frecuencias creado para evitar problemas si la cantidad de puntos es par o impar
freqs_V1 = freqs_V1[0:spectrum_V1.size]
freqs_V2 = freqs_V2[0:spectrum_V2.size]
freqs_V3 = freqs_V3[0:spectrum_V3.size]
freqs_V4 = freqs_V4[0:spectrum_V4.size]
# Búsque da picos en espectro con su umbral
umbral = 0.0095
picos_V1, _ = find_peaks(spectrum_V1, height=umbral)
picos_V2, _ = find_peaks(spectrum_V2, height=umbral)
picos_V3, _ = find_peaks(spectrum_V3, height=umbral)
picos_V4, _ = find_peaks(spectrum_V4, height=umbral)
# Representación en subplot de gráficos como vienen e invertidos
fig, axs = plt.subplots(2, 2, figsize=(16,9))
fig.suptitle('Espectros de ' + rotor + ' medido en sensor ' + sensor + ' ' + tipo_sensor + ' fecha ' + fecha + '(Conclusión 6)' )
axs[0,0].set_xlim( 0, 100 )
axs[0,0].plot(freqs_V1, spectrum_V1)
axs[0,0].plot(freqs_V1[picos_V1], spectrum_V1[picos_V1], "x")
axs[0,0].plot(np.ones(spectrum_V1.size)*umbral, "--", color="gray")
axs[0,0].set_title('Espectro sensor ' + tipo_sensor + ' Js11')
axs[0,0].grid(True)
axs[0,1].set_xlim( 0, 100 )
axs[0,1].plot( freqs_V2, spectrum_V2, 'tab:red')
axs[0,1].plot(freqs_V2[picos_V2], spectrum_V2[picos_V2], "x")
axs[0,1].plot(np.ones(spectrum_V2.size)*umbral, "--", color="gray")
axs[0,1].set_title('Espectro sensor ' + tipo_sensor + ' Js12')
axs[0,1].grid(True)
axs[1,0].set_xlim( 0, 100 )
axs[1,0].plot(freqs_V3, spectrum_V3, 'tab:orange')
axs[1,0].plot(freqs_V3[picos_V3], spectrum_V3[picos_V3], "x")
axs[1,0].plot(np.ones(spectrum_V3.size)*umbral, "--", color="gray")
axs[1,0].set_title('Espectro sensor ' + tipo_sensor + ' Js13')
axs[1,0].grid(True)
axs[1,1].set_xlim( 0, 100 )
axs[1,1].plot( freqs_V4, spectrum_V4, 'tab:green')
axs[1,1].plot(freqs_V4[picos_V4], spectrum_V4[picos_V4], "x")
axs[1,1].plot(np.ones(spectrum_V4.size)*umbral, "--", color="gray")
axs[1,1].set_title('Espectro sensor ' + tipo_sensor + ' Js14')
axs[1,1].grid(True)
plt.savefig(archivo_salida_a, dpi=300, facecolor='w', edgecolor='w',
orientation='portrait', papertype='a4', format=extension_salida, quality= 50)
plt.show()
# -
# ## Comparación de picos de fundamental y armónicos
# Comparación de cada uno de los picos en frecuencia y rms
# +
fig, axs = plt.subplots(1, 1, figsize=(16,9))
fig.suptitle('Picos de espectros en ensayos sobre ' + rotor + ' medido en sensor ' + sensor + ' ' + tipo_sensor + ' fecha ' + fecha )
axs.plot(freqs_V1[picos_V1[0]], spectrum_V1[picos_V1[0]], "x", color = "green", label = 'Js11')
axs.plot(freqs_V2[picos_V2[0]], spectrum_V2[picos_V2[0]], "o", color = "blue", label = 'Js12')
axs.plot(freqs_V3[picos_V3[0]], spectrum_V3[picos_V3[0]], "s", color = "red", label = 'Js13')
axs.plot(freqs_V4[picos_V4[0]], spectrum_V4[picos_V4[0]], "^", color = "black", label = 'Js14')
axs.set_title('Primer pico')
axs.grid(True)
axs.legend(loc="upper left")
plt.savefig(archivo_salida_b, dpi=300, facecolor='w', edgecolor='w',
orientation='portrait', papertype='a4', format=extension_salida, quality= 50)
plt.show()
# -
# ## Valores de picos
# +
print('Frecuencia 1er pico Js11', np.around(freqs_V1[picos_V1[0]], decimals = 3), '\nAmplitud 1er pico Js11', np.around(spectrum_V1[picos_V1[0]], decimals = 4 ), '\n')
print('Frecuencia 1er pico Js12', np.around(freqs_V2[picos_V2[0]], decimals = 3), '\nAmplitud 1er pico Js12', np.around(spectrum_V2[picos_V2[0]], decimals = 4 ), '\n')
print('Frecuencia 1er pico Js13', np.around(freqs_V3[picos_V3[0]], decimals = 3), '\nAmplitud 1er pico Js13', np.around(spectrum_V3[picos_V3[0]], decimals = 4 ), '\n')
print('Frecuencia 1er pico Js14', np.around(freqs_V4[picos_V4[0]], decimals = 3), '\nAmplitud 1er pico Js14', np.around(spectrum_V4[picos_V4[0]], decimals = 4 ), '\n')
print('Frecuencia 2do pico Js11', np.around(freqs_V1[picos_V1[1]], decimals = 3), '\nAmplitud 2do pico Js11', np.around(spectrum_V1[picos_V1[1]], decimals = 4 ), '\n')
print('Frecuencia 2do pico Js12', np.around(freqs_V2[picos_V2[1]], decimals = 3), '\nAmplitud 2do pico Js12', np.around(spectrum_V2[picos_V2[1]], decimals = 4 ), '\n')
print('Frecuencia 2do pico Js13', np.around(freqs_V3[picos_V3[1]], decimals = 3), '\nAmplitud 2do pico Js13', np.around(spectrum_V3[picos_V3[1]], decimals = 4 ), '\n')
print('Frecuencia 2do pico Js14', np.around(freqs_V4[picos_V4[1]], decimals = 3), '\nAmplitud 2do pico Js14', np.around(spectrum_V4[picos_V4[1]], decimals = 4 ), '\n')
|
Ensayo 20200208/Lectura datos/Conclusion 5.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import random
import pandas as pd
import numpy as np
from scipy.spatial.distance import hamming
# -
df = pd.read_feather('./data.feather')
df.info()
user1 = random.sample(df['User-ID'].drop_duplicates().to_list(), 1)[0]
user1
user_data = df[df['User-ID'] == user1]
user_data
PERC_OF_DATA_TO_USE = 100
user_rating_counts = df['User-ID'].value_counts()
cutoff_point = int(user_rating_counts.shape[0]*(PERC_OF_DATA_TO_USE/100.0))
users_with_most_ratings = user_rating_counts[:cutoff_point]
book_rating_counts = df['ISBN'].value_counts()
cutoff_point = int(book_rating_counts.shape[0]*(PERC_OF_DATA_TO_USE/100.0))
books_with_most_ratings = book_rating_counts[:cutoff_point]
df = df[df.apply(lambda rating: rating['User-ID'] in users_with_most_ratings, axis=1)]
df = df[df.apply(lambda rating: rating['ISBN'] in books_with_most_ratings, axis=1)]
df = df.append(user_data, ignore_index=True).drop_duplicates()
df.info()
user_item_df = df.drop(columns=['Book-Title', 'Book-Author', 'Book-Year-Of-Publication', 'Book-Publisher']).dropna().reset_index(drop=True)
user_item_df['Book-Rating'] = user_item_df['Book-Rating'].astype(np.int16)
user_item_df.head()
user_item_matrix = pd.pivot_table(user_item_df, values='Book-Rating', index='User-ID', columns='ISBN')
user_item_matrix
df = df[df['User-ID'] != user1]
def hamming_distance(user1, user2, user_item_matrix):
try:
user1_ratings = user_item_matrix.transpose()[user1]
user2_ratings = user_item_matrix.transpose()[user2]
distance = hamming(user1_ratings, user2_ratings)
except:
distance = np.NaN
return distance
user1_ratings = user_item_matrix.transpose()[user1]
user2_ratings = user_item_matrix.transpose()[random.sample(df['User-ID'].drop_duplicates().to_list(), 1)[0]]
distance = hamming(user1_ratings,user2_ratings)
distance
df["Distance"] = df["User-ID"].apply(lambda user2: hamming_distance(user1, user2, user_item_matrix))
df.head()
df.sort_values(["Distance"], ascending=True)
RECOMMENDATION_AMOUNT = 3
print(user1)
neighbours_amount = RECOMMENDATION_AMOUNT*2
k_nearest_users = df[df['User-ID'] != user1].sort_values(["Distance"], ascending=True)["User-ID"].drop_duplicates()[:neighbours_amount]
k_nearest_users
nn_ratings = user_item_matrix[user_item_matrix.index.isin(k_nearest_users)]
nn_ratings
books_read = user_item_matrix.transpose()[user1].dropna().index
books_read
avg_rating = nn_ratings.apply(np.nanmean).dropna()
avg_rating
avg_rating = avg_rating[~avg_rating.index.isin(books_read)]
avg_rating
avg_rating.sort_values(ascending=False)
recommended_books = avg_rating.sort_values(ascending=False).index[:RECOMMENDATION_AMOUNT]
recommended_books
df[df['ISBN'].apply(lambda isbn: isbn in recommended_books)].drop(columns=['User-ID', 'Book-Rating']).sort_values(["Distance"], ascending=True).drop_duplicates()[:RECOMMENDATION_AMOUNT]
|
User-item_recommender.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <b>Calule a derivada da função dada. Simplifique as respostas.</b>
# <b>11. $y = x^2 + 2x + 3$</b>
# <b>Aplicando a regra da soma $\frac{d}{dx}(f+g) = \frac{d}{dx}(f) + \frac{d}{dx}(g) $</b><br><br>
# $\frac{d}{dx}(x^2) + \frac{d}{dx}(2x) + \frac{d}{dx}(3)$<br><br>
#
# <b>Aplicando a regra da potência: </b> $\frac{d}{dx}(x^a) = a\cdot x^{a-1}$<br><br>
#
# $\frac{d}{dx}(x^2) = 2x^{2-1}$<br><br>
# $\frac{d}{dx}(x^2) = 2x$<br><br><br>
#
# $\frac{d}{dx}(2x) = 2x^{1-1}$<br><br>
# $\frac{d}{dx}(2x) = 2x^0$<br><br>
# $\frac{d}{dx}(2x) = 2$<br><br><br>
#
# <b>Derivada de uma constante é zero</b><br><br>
# $\frac{d}{dx}(3) = 0$<br><br>
#
# $\frac{d}{dx}(x^2) + \frac{d}{dx}(2x) + \frac{d}{dx}(3) = 2x + 2$<br>
#
#
#
|
Problemas 2.2/11.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy
import sys
import nltk
nltk.download('stopwords')
from nltk.tokenize import RegexpTokenizer
from nltk.corpus import stopwords
from keras.models import Sequential
from keras.utils import np_utils
from keras.layers import Dense,Dropout,LSTM
from keras.callbacks import ModelCheckpoint
file=open("frankestein.txt").read()
def tokenize_words(input):
input=input.lower()
tokenizer=RegexTokenizer(r'\w+')
tokens=tokenizer.tokenize(input)
filtered=filter(lambda token:token not in stopwords.words('english'),tokens)
return "".join(filtered)
processed_input=tokenize_words(file)
# +
chars=sorted(list(set(processed_input)))
chat_ro_num=dict(c,i) for i,c in enumerate(chars)
# -
input_len=len(processed_input)
vocab_len=len(chars)
print("Total number of characters:",input_len)
print("total vocab:",vocab_len)
seq_length=100
x_data=[]
y_data=[]
for i in range(0,input_len-seq_length):
in_seq=processed_input[i:i +seq_length]
out_seq=processed_input[i + seq_length]
x_data.append([char_to_num[char] for char in in_seq])
y_data.append(char_to_num[out_seq])
n_patterns=len(x_data)
print("Total Pattern",n_pattern)
# +
X=numpy.reshape(x_data,(n_pattern,seq_length,1))
X=X/float(vocab_len)
# -
y=np.utils.to_categorical(y_data)
model=Sequential()
model.add(LSTM(256,input_shape=(X.shape[2]),return_sequences=True))
model.add(Dropout(0.2))
model.add(LSMT(128))
model.add(Dropout(0.2))
model.add(Dense(y.shape[1],activation='softmax'))
model.compile(loss='categorical_crossentropy',optimizer='adam')
filepath="model_weight_saved.hdf5"
checkpoint=Modelcheckpoint(filepath,monitor='loss',verbose=1,save_best_only=True,mode='min')
desired_callbacks=[checkpoint]
filename="model_weights_saved.hdf5"
model.load_weights(filename)
model.compile(loss='categorical_crossentropy',optimizer='adam')
# +
num_to_char=dict((i,c)for i,c in enumerate(chars))
start=numpy.random.randint(0,len(x_data)-1)
pattern("random seed:")
print("\".".join([num_to_char[value] for value in pattern])"\"")
# -
for i in range(1000):
x=numpy.reshape(pattern,(1,len(pattern),1))
x=xfloat(vocab_len)
prediction=model.predict(x,verbose=0)
index=numpy.argmax(prediction)
result=num_to_char[index]
seq_in=[num_to_char[value] for value in pattern]
sys.stdout.write(result)
pattern.append(index)
pattern=pattern[1:len(pattern)]
|
text_generation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Serving MLflow models
#
# Out of the box, MLServer supports the deployment and serving of MLflow models with the following features:
#
# - Loading of MLflow Model artifacts.
# - Support of dataframes, dict-of-tensors and tensor inputs.
#
# In this example, we will showcase some of this features using an example model.
# +
from IPython.core.magic import register_line_cell_magic
@register_line_cell_magic
def writetemplate(line, cell):
with open(line, 'w') as f:
f.write(cell.format(**globals()))
# -
# ## Training
#
# The first step will be to train and serialise a MLflow model.
# For that, we will use the [linear regression examle from the MLflow docs](https://www.mlflow.org/docs/latest/tutorials-and-examples/tutorial.html).
# + tags=[]
# # %load src/train.py
# Original source code and more details can be found in:
# https://www.mlflow.org/docs/latest/tutorials-and-examples/tutorial.html
# The data set used in this example is from
# http://archive.ics.uci.edu/ml/datasets/Wine+Quality
# <NAME>, <NAME>, <NAME>, <NAME> and <NAME>.
# Modeling wine preferences by data mining from physicochemical properties.
# In Decision Support Systems, Elsevier, 47(4):547-553, 2009.
import warnings
import sys
import pandas as pd
import numpy as np
from sklearn.metrics import mean_squared_error, mean_absolute_error, r2_score
from sklearn.model_selection import train_test_split
from sklearn.linear_model import ElasticNet
from urllib.parse import urlparse
import mlflow
import mlflow.sklearn
from mlflow.models.signature import infer_signature
import logging
logging.basicConfig(level=logging.WARN)
logger = logging.getLogger(__name__)
def eval_metrics(actual, pred):
rmse = np.sqrt(mean_squared_error(actual, pred))
mae = mean_absolute_error(actual, pred)
r2 = r2_score(actual, pred)
return rmse, mae, r2
if __name__ == "__main__":
warnings.filterwarnings("ignore")
np.random.seed(40)
# Read the wine-quality csv file from the URL
csv_url = (
"http://archive.ics.uci.edu/ml"
"/machine-learning-databases/wine-quality/winequality-red.csv"
)
try:
data = pd.read_csv(csv_url, sep=";")
except Exception as e:
logger.exception(
"Unable to download training & test CSV, "
"check your internet connection. Error: %s",
e,
)
# Split the data into training and test sets. (0.75, 0.25) split.
train, test = train_test_split(data)
# The predicted column is "quality" which is a scalar from [3, 9]
train_x = train.drop(["quality"], axis=1)
test_x = test.drop(["quality"], axis=1)
train_y = train[["quality"]]
test_y = test[["quality"]]
alpha = float(sys.argv[1]) if len(sys.argv) > 1 else 0.5
l1_ratio = float(sys.argv[2]) if len(sys.argv) > 2 else 0.5
with mlflow.start_run():
lr = ElasticNet(alpha=alpha, l1_ratio=l1_ratio, random_state=42)
lr.fit(train_x, train_y)
predicted_qualities = lr.predict(test_x)
(rmse, mae, r2) = eval_metrics(test_y, predicted_qualities)
print("Elasticnet model (alpha=%f, l1_ratio=%f):" % (alpha, l1_ratio))
print(" RMSE: %s" % rmse)
print(" MAE: %s" % mae)
print(" R2: %s" % r2)
mlflow.log_param("alpha", alpha)
mlflow.log_param("l1_ratio", l1_ratio)
mlflow.log_metric("rmse", rmse)
mlflow.log_metric("r2", r2)
mlflow.log_metric("mae", mae)
tracking_url_type_store = urlparse(mlflow.get_tracking_uri()).scheme
model_signature = infer_signature(train_x, train_y)
# Model registry does not work with file store
if tracking_url_type_store != "file":
# Register the model
# There are other ways to use the Model Registry,
# which depends on the use case,
# please refer to the doc for more information:
# https://mlflow.org/docs/latest/model-registry.html#api-workflow
mlflow.sklearn.log_model(
lr,
"model",
registered_model_name="ElasticnetWineModel",
signature=model_signature,
)
else:
mlflow.sklearn.log_model(lr, "model", signature=model_signature)
# -
# !python src/train.py
# The training script will also serialise our trained model, leveraging the [MLflow Model format](https://www.mlflow.org/docs/latest/models.html).
# By default, we should be able to find the saved artifact under the `mlruns` folder.
[experiment_file_path] = !ls -td ./mlruns/0/* | head -1
model_path = os.path.join(experiment_file_path, "artifacts", "model")
print(model_path)
# !ls {model_path}
# ## Serving
#
# Now that we have trained and serialised our model, we are ready to start serving it.
# For that, the initial step will be to set up a `model-settings.json` that instructs MLServer to load our artifact using the MLflow Inference Runtime.
# %%writetemplate ./model-settings.json
{{
"name": "wine-classifier",
"implementation": "mlserver_mlflow.MLflowRuntime",
"parameters": {{
"uri": "{model_path}"
}}
}}
# Now that we have our config in-place, we can start the server by running `mlserver start .`. This needs to either be ran from the same directory where our config files are or pointing to the folder where they are.
#
# ```shell
# mlserver start .
# ```
#
# Since this command will start the server and block the terminal, waiting for requests, this will need to be ran in the background on a separate terminal.
# ### Send test inference request
#
# We now have our model being served by `mlserver`.
# To make sure that everything is working as expected, let's send a request from our test set.
# For that, we can use the Python types that `mlserver` provides out of box, or we can build our request manually.
#
# Note that, the request specifies the value `pd` as its *content type*, whereas every input specifies the *content type* `np`.
# These parameters will instruct MLServer to:
#
# - Convert every input value to a NumPy array, using the data type and shape information provided.
# - Group all the different inputs into a Pandas DataFrame, using their names as the column names.
#
# To learn more about how MLServer uses content type parameters, you can check this [worked out example](../content-type/README.md).
# +
import requests
inference_request = {
"inputs": [
{
"name": "fixed acidity",
"shape": [1],
"datatype": "FP32",
"data": [7.4],
},
{
"name": "volatile acidity",
"shape": [1],
"datatype": "FP32",
"data": [0.7000],
},
{
"name": "citric acid",
"shape": [1],
"datatype": "FP32",
"data": [0],
},
{
"name": "residual sugar",
"shape": [1],
"datatype": "FP32",
"data": [1.9],
},
{
"name": "chlorides",
"shape": [1],
"datatype": "FP32",
"data": [0.076],
},
{
"name": "free sulfur dioxide",
"shape": [1],
"datatype": "FP32",
"data": [11],
},
{
"name": "total sulfur dioxide",
"shape": [1],
"datatype": "FP32",
"data": [34],
},
{
"name": "density",
"shape": [1],
"datatype": "FP32",
"data": [0.9978],
},
{
"name": "pH",
"shape": [1],
"datatype": "FP32",
"data": [3.51],
},
{
"name": "sulphates",
"shape": [1],
"datatype": "FP32",
"data": [0.56],
},
{
"name": "alcohol",
"shape": [1],
"datatype": "FP32",
"data": [9.4],
},
]
}
endpoint = "http://localhost:8080/v2/models/wine-classifier/infer"
response = requests.post(endpoint, json=inference_request)
response.json()
# -
# As we can see in the output above, the predicted quality score for our input wine was `5.57`.
# ### MLflow Scoring Protocol
#
# MLflow currently ships with an [scoring server with its own protocol](https://www.mlflow.org/docs/latest/models.html#deploy-mlflow-models).
# In order to provide a drop-in replacement, the MLflow runtime in MLServer also exposes a custom endpoint which matches the signature of the MLflow's `/invocations` endpoint.
#
# As an example, we can try to send the same request that sent previously, but using MLflow's protocol.
# Note that, in both cases, the request will be handled by the same MLServer instance.
# +
import requests
inference_request = {
"columns": [
"alcohol",
"chlorides",
"citric acid",
"density",
"fixed acidity",
"free sulfur dioxide",
"pH",
"residual sugar",
"sulphates",
"total sulfur dioxide",
"volatile acidity",
],
"data": [[7.4,0.7,0,1.9,0.076,11,34,0.9978,3.51,0.56,9.4]],
}
endpoint = "http://localhost:8080/invocations"
response = requests.post(endpoint, json=inference_request)
response.json()
# -
# As we can see above, the predicted quality for our input is `5.57`, matching the prediction we obtained above.
# ### MLflow Model Signature
#
# MLflow lets users define a [_model signature_](https://www.mlflow.org/docs/latest/models.html#model-signature-and-input-example), where they can specify what types of inputs does the model accept, and what types of outputs it returns.
# Similarly, the [V2 inference protocol](https://github.com/kubeflow/kfserving/tree/master/docs/predict-api/v2) employed by MLServer defines a [_metadata endpoint_](https://github.com/kubeflow/kfserving/blob/master/docs/predict-api/v2/required_api.md#model-metadata) which can be used to query what inputs and outputs does the model accept.
# However, even though they serve similar functions, the data schemas used by each one of them are not compatible between them.
#
# To solve this, if your model defines a MLflow model signature, MLServer will convert _on-the-fly_ this signature to a metadata schema compatible with the V2 Inference Protocol.
# This will also include specifying any extra [content type](../content-type/README.md) that is required to correctly decode / encode your data.
#
# As an example, we can first have a look at the model signature saved for our MLflow model.
# This can be seen directly on the `MLModel` file saved by our model.
#
#
#
# !cat {model_path}/MLmodel
# We can then query the metadata endpoint, to see the model metadata inferred by MLServer from our test model's signature.
# For this, we will use the `/v2/models/wine-classifier/` endpoint.
# +
import requests
endpoint = "http://localhost:8080/v2/models/wine-classifier"
response = requests.get(endpoint)
response.json()
# -
# As we should be able to see, the model metadata now matches the information contained in our model signature, including any extra content types necessary to decode our data correctly.
|
docs/examples/mlflow/README.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # MAB experiment
#
# In this notebook we will simulate a multi armed bandit experiment, using different approaches to solve it
# +
import os
import sys
sys.path.append('/Users/fci02/Documents/GitHub/decisioning-analysis/test_and_learn/production_modules/')
import numpy as np
import pandas as pd
from scipy.stats import f_oneway
from tqdm import tqdm
from contextual_mab.experiments.framework import MABFramework
from contextual_mab.experiments.framework import run_experiment
from contextual_mab.data_utils.data_generation import generate_experimental_dataset
import matplotlib.pyplot as plt
import seaborn as sns
# -
# ## Experimental data generation
# +
n_1 = 5000
noise_scale_1=5.
cw_1 = [.05,.03,.01]
ct_prm_1 = {'X1':{'loc':0.,'scale':1.},
'X2':{'loc':10.,'scale':5.},
'X3':{'loc':-.3,'scale':.5}}
ord_prm_1 = {'O1':{'start':1,'stop':11,'weights':[.1]*10},
'O2':{'start':1,'stop':4,'weights':[.3,.4,.3]}}
catg_prm_1 = {'C1':{'levels':['Yes','No'],'weights':[.4,.6]},
'C2':{'levels':['Green','Yellow'],'weights':[.2,.8]},
'C3':{'levels':['A','B','C'],'weights':[.4,.1,.5]}}
weights_1 = [[-0.85427315, 1.16572722, 0.8890073, -0.57988417, 0.15193386,
-0.28800726, -0.06606457, 0.36732494, -0.03666541, -0.61067511],
[ 0.46719077, -1.64435559, 0.69791627, -0.14981489, 0.26023682,
0.67528998, 1.52150038, 1.05417964, 0.37329345, 0.79700709],
[ 1.62075116, 0.15865047, -0.85728784, 1.23667642, -0.58707557,
0.05713119, -0.47387454, 0.51293855, -0.55820087, -2.14815787]]
# +
n_2 = 5000
cw_2 = [.02,.07,.03]
ct_prm_2 = {'X1':{'loc':5.,'scale':1.},
'X2':{'loc':1.,'scale':5.},
'X3':{'loc':1.3,'scale':.5}}
ord_prm_2 = {'O1':{'start':1,'stop':11,'weights':[.2,.2,.05,.05,.05,.05,.1,.1,.1,.1]},
'O2':{'start':1,'stop':4,'weights':[.1,.6,.3]}}
catg_prm_2 = {'C1':{'levels':['Yes','No'],'weights':[.1,.9]},
'C2':{'levels':['Green','Yellow'],'weights':[.5,.5]},
'C3':{'levels':['A','B','C'],'weights':[.6,.2,.2]}}
noise_scale_2=5.
weights_2 = [[ 0.2249653, 0.54732847, 0.76620536, 0.7461608, -0.76568111,
-0.13241893, -1.82046231, -0.47742618, -0.96465132, -0.68848216],
[-0.04717597, 1.49105257, -0.6332578, -1.03206255, -1.30374031,
-0.48575409, 0.01466847, 0.54927814, 0.72014772, 0.42807199],
[-0.56907754, 0.04875765, 0.89346343, 0.62619356, -2.19116666,
1.70168624, 0.34768686, 0.26208243, 1.27787397, -2.07476064]]
# +
n_3 = 5000
cw_3 = [.01,.03,.06]
ct_prm_3 = {'X1':{'loc':-2.,'scale':1.},
'X2':{'loc':-20.,'scale':5.},
'X3':{'loc':11.3,'scale':.5}}
ord_prm_3 = {'O1':{'start':1,'stop':11,'weights':[.5,.0,.0,.0,.05,.05,.1,.1,.1,.1]},
'O2':{'start':1,'stop':4,'weights':[.5,.1,.4]}}
catg_prm_3 = {'C1':{'levels':['Yes','No'],'weights':[.5,.5]},
'C2':{'levels':['Green','Yellow'],'weights':[.8,.2]},
'C3':{'levels':['A','B','C'],'weights':[.35,.35,.3]}}
noise_scale_3=5.
weights_3 = [[ 0.2249653, 0.54732847, 0.76620536, 0.7461608, -0.76568111,
-0.13241893, -1.82046231, -0.47742618, -0.96465132, -0.68848216],
[-0.04717597, 1.49105257, -0.6332578, -1.03206255, -1.30374031,
-0.48575409, 0.01466847, 0.54927814, 0.72014772, 0.42807199],
[-0.56907754, 0.04875765, 0.89346343, 0.62619356, -2.19116666,
1.70168624, 0.34768686, 0.26208243, 1.27787397, -2.07476064]]
# -
seed = 0
experiment_data = generate_experimental_dataset([n_1,n_2,n_3],
[cw_1,cw_2,cw_3],
[ct_prm_1,ct_prm_2,ct_prm_3],
[ord_prm_1,ord_prm_2,ord_prm_3],
[catg_prm_1,catg_prm_2,catg_prm_3],
[noise_scale_1,noise_scale_2,noise_scale_3],
[weights_1,weights_2,weights_3],
seed,
output_info=True)
# ## Single run
# ### ABC greedy
# +
batch_size = 2000
run_experiment(experiment_data, batch_size, MABFramework,{'strategy':'static-one-fits-all','n_actions':3,'static_min_steps':2})
# -
# ### Thompson Sampling
run_experiment(experiment_data, batch_size, MABFramework,{'strategy':'dynamic-one-fits-all','n_actions':3,'alphas':[1.,1.,1.],'betas':[1.,1.,1.]})
# ### Contextual Thompson Sampling
# #### Bayesian Logistic
from contextual_mab.predictive_models.bayesian_logistic import BayesianLogisticRegression
action_cols = [column for column in experiment_data.columns if 'action' in column]
predictors = pd.get_dummies(experiment_data.drop(columns=action_cols),drop_first=True).columns.tolist()
predictors
run_experiment(experiment_data, batch_size, MABFramework,{'strategy':'contextual-one-fits-one','n_actions':3,
'modelling_approach':BayesianLogisticRegression,
'modelling_approach_pms':{'n_samples':500,'n_chains':2,
'predictors':predictors.copy(),
'tune':1000,
'check_prog':False}})
# #### Bootstrap Oracles
from contextual_mab.predictive_models.oracles import BootstrapOracle
from sklearn.linear_model import LogisticRegression
logistic_params = {'solver':'lbfgs',
'max_iter':500,
'random_state':0}
run_experiment(experiment_data, batch_size, MABFramework,{'strategy':'contextual-one-fits-one','n_actions':3,
'modelling_approach':BootstrapOracle,
'modelling_approach_pms':{'n_bootstrap':1000,
'learner_class':LogisticRegression,
'learner_class_params':logistic_params,
'check_prog':False}})
# ## Running many experiments
batch_size = 2000
abc_approval_rates = []
for seed in range(10):
experiment_data = generate_experimental_dataset([n_1,n_2,n_3],
[cw_1,cw_2,cw_3],
[ct_prm_1,ct_prm_2,ct_prm_3],
[ord_prm_1,ord_prm_2,ord_prm_3],
[catg_prm_1,catg_prm_2,catg_prm_3],
[noise_scale_1,noise_scale_2,noise_scale_3],
[weights_1,weights_2,weights_3],
seed)
abc_approval_rates.append(run_experiment(experiment_data, batch_size, MABFramework,{'strategy':'static-one-fits-all','n_actions':3,'static_min_steps':2}))
ts_approval_rates = []
for seed in range(10):
experiment_data = generate_experimental_dataset([n_1,n_2,n_3],
[cw_1,cw_2,cw_3],
[ct_prm_1,ct_prm_2,ct_prm_3],
[ord_prm_1,ord_prm_2,ord_prm_3],
[catg_prm_1,catg_prm_2,catg_prm_3],
[noise_scale_1,noise_scale_2,noise_scale_3],
[weights_1,weights_2,weights_3],
seed)
ts_approval_rates.append(run_experiment(experiment_data, batch_size, MABFramework,{'strategy':'dynamic-one-fits-all','n_actions':3,'alphas':[1.,1.,1.],'betas':[1.,1.,1.]}))
from statsmodels.stats.weightstats import ztest
ztest(abc_approval_rates,ts_approval_rates)
|
examples/Contextual_MAB_experiment.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1 align = 'center'> Neural Networks Demystified </h1>
# <h2 align = 'center'> Part 6: Training </h2>
#
#
# <h4 align = 'center' > @stephencwelch </h4>
from IPython.display import YouTubeVideo
YouTubeVideo('9KM9Td6RVgQ')
# So far we’ve built a neural network in python, computed a cost function to let us know how well our network is performing, computed the gradient of our cost function so we can train our network, and last time we numerically validated our gradient computations. After all that work, it’s finally time to train our neural network.
# Back in part 3, we decided to train our network using gradient descent. While gradient descent is conceptually pretty straight forward, its implementation can actually be quite complex- especially as we increase the size and number of layers in our neural network. If we just march downhill with consistent step sizes, we may get stuck in a local minimum or flat spot, we may move too slowly and never reach our minimum, or we may move to quickly and bounce out of our minimum. And remember, all this must happen in high-dimensional space, making things significantly more complex. Gradient descent is a wonderfully clever method, but provides no guarantees that we will converge to a good solution, that we will converge to a solution in a certain amount of time, or that we will converge to a solution at all.
# The good and bad news here is that this problem is not unique to Neural Networks - there’s an entire field dedicated to finding the best combination of inputs to minimize the output of an objective function: the field of Mathematical Optimization. The bad news is that optimization can be a bit overwhelming; there are many different techniques we could apply to our problem.
# Part of what makes the optimization challenging is the broad range of approaches covered - from very rigorous, theoretical methods to hands-on, more heuristics-driven methods. <NAME>’s 1998 publication Efficient BackProp presents an excellent review of various optimization techniques as applied to neural networks.
# Here, we’re going to use a more sophisticated variant on gradient descent, the popular Broyden-Fletcher-Goldfarb-Shanno numerical optimization algorithm. The BFGS algorithm overcomes some of the limitations of plain gradient descent by estimating the second derivative, or curvature, of the cost function surface, and using this information to make more informed movements downhill. BFGS will allow us to find solutions more often and more quickly.
# We’ll use the BFGS implementation built into the scipy optimize package, specifically within the minimize function. To use BFGS, the minimize function requires us to pass in an objective function that accepts a vector of parameters, input data, and output data, and returns both the cost and gradients. Our neural network implementation doesn’t quite follow these semantics, so we’ll use a wrapper function to give it this behavior. We’ll also pass in initial parameters, set the jacobian parameter to true since we’re computing the gradient within our neural network class, set the method to BFGS, pass in our input and output data, and some options. Finally, we’ll implement a callback function that allows us to track the cost function value as we train the network. Once the network is trained, we’ll replace the original, random parameters, with the trained parameters.
# %pylab inline
#Import code from previous videos:
from partFive import *
from scipy import optimize
class trainer(object):
def __init__(self, N):
#Make Local reference to network:
self.N = N
def callbackF(self, params):
self.N.setParams(params)
self.J.append(self.N.costFunction(self.X, self.y))
def costFunctionWrapper(self, params, X, y):
self.N.setParams(params)
cost = self.N.costFunction(X, y)
grad = self.N.computeGradients(X,y)
return cost, grad
def train(self, X, y):
#Make an internal variable for the callback function:
self.X = X
self.y = y
#Make empty list to store costs:
self.J = []
params0 = self.N.getParams()
options = {'maxiter': 200, 'disp' : True}
_res = optimize.minimize(self.costFunctionWrapper, params0, jac=True, method='L-BFGS-B', \
args=(X, y), options=options, callback=self.callbackF)
self.N.setParams(_res.x)
self.optimizationResults = _res
# If we plot the cost against the number of iterations through training, we should see a nice, monotonically decreasing function. Further, we see that the number of function evaluations required to find the solution is less than 100, and far less than the 10^27 function evaluation that would have been required to find a solution by brute force, as shown in part 3. Finally, we can evaluate our gradient at our solution and see very small values – which make sense, as our minimum should be quite flat.
NN = Neural_Network()
T = trainer(NN)
T.train(X,y)
plot(T.J)
grid(1)
xlabel('Iterations')
ylabel('Cost')
NN.costFunctionPrime(X,y)
# The more exciting thing here is that we finally have a trained network that can predict your score on a test based on how many hours you sleep and how many hours you study the night before. If we run our training data through our forward method now, we see that our predictions are excellent. We can go one step further and explore the input space for various combinations of hours sleeping and hours studying, and maybe we can find an optimal combination of the two for your next test.
NN.forward(X)
y
# +
#Test network for various combinations of sleep/study:
hoursSleep = linspace(0, 10, 100)
hoursStudy = linspace(0, 5, 100)
#Normalize data (same way training data way normalized)
hoursSleepNorm = hoursSleep/10.
hoursStudyNorm = hoursStudy/5.
#Create 2-d versions of input for plotting
a, b = meshgrid(hoursSleepNorm, hoursStudyNorm)
#Join into a single input matrix:
allInputs = np.zeros((a.size, 2))
allInputs[:, 0] = a.ravel()
allInputs[:, 1] = b.ravel()
# -
allOutputs = NN.forward(allInputs)
# +
#Contour Plot:
yy = np.dot(hoursStudy.reshape(100,1), np.ones((1,100)))
xx = np.dot(hoursSleep.reshape(100,1), np.ones((1,100))).T
CS = contour(xx,yy,100*allOutputs.reshape(100, 100))
clabel(CS, inline=1, fontsize=10)
xlabel('Hours Sleep')
ylabel('Hours Study')
# +
#3D plot:
##Uncomment to plot out-of-notebook (you'll be able to rotate)
# #%matplotlib qt
from mpl_toolkits.mplot3d import Axes3D
fig = plt.figure()
ax = fig.gca(projection='3d')
surf = ax.plot_surface(xx, yy, 100*allOutputs.reshape(100, 100), \
cmap=cm.jet)
ax.set_xlabel('Hours Sleep')
ax.set_ylabel('Hours Study')
ax.set_zlabel('Test Score')
# -
# Our results look pretty reasonable, and we see that for our model, sleep actually has a bigger impact on your grade than studying – something I wish I had realized when I was in school. So we’re done, right?
# Nope.
#
# We’ve made possibly the most dangerous and tempting error in machine learning – overfitting. Although our network is performing incredibly well (maybe too well) on our training data – that doesn’t mean that our model is a good fit for the real world, and that’s what we’ll work on next time.
|
Lectures/Lecture-06/code/Part 6 Training.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] graffitiCellId="id_wzbr23x"
# ### Problem Statement
#
# The Tower of Hanoi is a puzzle where we have three rods and `n` disks. The three rods are:
# 1. source
# 2. destination
# 3. auxiliary
#
# Initally, all the `n` disks are present on the source rod. The final objective of the puzzle is to move all disks from the source rod to the destination rod using the auxiliary rod. However, there are some rules according to which this has to be done:
# 1. Only one disk can be moved at a time.
# 2. A disk can be moved only if it is on the top of a rod.
# 3. No disk can be placed on the top of a smaller disk.
#
# You will be given the number of disks `num_disks` as the input parameter.
#
# For example, if you have `num_disks = 3`, then the disks should be moved as follows:
#
# 1. move disk from source to auxiliary
# 2. move disk from source to destination
# 3. move disk from auxiliary to destination
#
# You must print these steps as follows:
#
# S A
# S D
# A D
#
# Where S = source, D = destination, A = auxiliary
#
# + graffitiCellId="id_8tcr5o8"
def tower_of_Hanoi(num_disks):
"""
:param: num_disks - number of disks
TODO: print the steps required to move all disks from source to destination
"""
pass
# + [markdown] graffitiCellId="id_rh9jy5w"
# <span class="graffiti-highlight graffiti-id_rh9jy5w-id_aaedpt9"><i></i><button>Show Solution</button></span>
# + [markdown] graffitiCellId="id_6dm5twe"
# #### Compare your results with the following test cases
# * num_disks = 2
#
# solution
# S A
# S D
# A D
#
# * num_disks = 3
#
# solution
# S D
# S A
# D A
# S D
# A S
# A D
# S D
#
# * num_disks = 4
#
# solution
# S A
# S D
# A D
# S A
# D S
# D A
# S A
# S D
# A D
# A S
# D S
# A D
# S A
# S D
# A D
|
recursion/.ipynb_checkpoints/Tower-of-Hanoi-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import yahoo_fin.stock_info as si
evaluationDF = pd.read_csv('../ProjectDatasets/final_recommendations.csv', sep=',', index_col='ticker')
evaluationDF.head(50)
|
ProjectCode/003_examine_results.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import datetime as dt
# +
path = "Resources/cities.csv"
df = pd.read_csv(path)
df
# +
#Converting to HTML
html = df.to_html()
print(html)
# -
code = open("code_export.txt", "w")
code.write(html)
code.close()
|
CSV to HTML.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
from nltk import FreqDist
from nltk.tokenize import word_tokenize, sent_tokenize
from nltk.stem.wordnet import WordNetLemmatizer
lemma = WordNetLemmatizer()
from nltk.corpus import stopwords
stop = stopwords.words('english')
from bs4 import BeautifulSoup
from urllib.request import urlopen
from gensim.models import Phrases
from gensim.models.phrases import Phraser
import os
from collections import Counter
import string
punctuations = list(string.punctuation)
#Add some more punctuation, as the list doesn't cover all cases.
punctuations.extend(['”', '–', '``', "''"])
stop = stop + punctuations
# +
url = urlopen('http://www.iflscience.com/technology/new-device-generate-electricity-anywhere-natural-temperature-changes/')
soup = BeautifulSoup(url.read().decode('utf8'), "lxml")
text = '\n\n'.join(map(lambda p: p.text, soup.find_all('p')))
#text = text[text.find('enumerated'):]
title = soup.find('h1').text.strip()
print(title, '\n', '_' * 60, '\n', text)
# +
def intersection(sent1, sent2):
s1 = [i for i in word_tokenize(sent1) if i not in punctuations and i not in stop]
s2 = [i for i in word_tokenize(sent2) if i not in punctuations and i not in stop]
intersection = [i for i in s1 if i in s2]
return len(intersection) / ((len(s1) + len(s2)) / 2)
def get_summary(text, limit=3):
sentences = sent_tokenize(text)
matrix = [[intersection(sentences[i], sentences[j]) for i in range(0,len(sentences))] for j in range(0,len(sentences))]
scores = {sentences[i]: sum(matrix[i]) for i in range(len(matrix))}
sents = sorted(scores, key=scores.__getitem__, reverse=True)[:limit]
best_sents = [i[0] for i in sorted([(i, text.find(i)) for i in sents], key=lambda x: x[0])]
return best_sents
def summarize(text, limit=3):
summary = get_summary(text, limit)
print(title)
print()
print(' '.join(summary))
# -
summarize(text,2)
# +
def intersection(sent1, sent2):
#As sentences are lists of tokens, there is no need to split them.
intersection = [i for i in sent1 if i in sent2]
return len(intersection) / ((len(sent1) + len(sent2)) / 2)
def split_sentences(sents):
sentence_stream = [[i for i in word_tokenize(sent) if i not in stop] for sent in sents]
bigram = Phrases(sentence_stream, min_count=2, threshold=2, delimiter=b'_')
bigram_phraser = Phraser(bigram)
bigram_tokens = bigram_phraser[sentence_stream]
trigram = Phrases(bigram_tokens,min_count=2, threshold=2, delimiter=b'_')
trigram_phraser = Phraser(trigram)
trigram_tokens = trigram_phraser[bigram_tokens]
return [i for i in trigram_tokens]
def get_summary(text, limit=3):
sents = sent_tokenize(text)
sentences = split_sentences(sents)
matrix = [[intersection(sentences[i], sentences[j]) for i in range(0,len(sentences))] for j in range(0,len(sentences))]
scores = {sents[i]: sum(matrix[i]) for i in range(len(matrix))}
sents = sorted(scores, key=scores.__getitem__, reverse=True)[:limit]
best_sents = [i[0] for i in sorted([(i, text.find(i)) for i in sents], key=lambda x: x[0])]
return best_sents
# -
summarize(text, 5)
# ### Word Frequency
# +
def score_sentences(words, sentences):
#Return scores for sentences.
scores = Counter()
#Words - list of words and their scores, first element is the word, second - its score.
for word in words:
for i in range(0, len(sentences)):
#If word is also in title, then add double score to the sentence.
if word[0] in sentences[i] and word[0] in title:
scores[i] += 2 * word[1]
elif word[0] in sentences[i]:
scores[i] += word[1]
sentence_scores = sorted(scores.items(), key=scores.__getitem__, reverse=True)
return sentence_scores
def split_sentences(sents):
sentence_stream = [[i for i in word_tokenize(sent) if i not in stop] for sent in sents]
bigram = Phrases(sentence_stream, min_count=2, threshold=2, delimiter=b'_')
bigram_phraser = Phraser(bigram)
bigram_tokens = bigram_phraser[sentence_stream]
trigram = Phrases(bigram_tokens,min_count=2, threshold=2, delimiter=b'_')
trigram_phraser = Phraser(trigram)
trigram_tokens = trigram_phraser[bigram_tokens]
all_words = [i for j in trigram_tokens for i in j]
frequent_words = [i for i in Counter(all_words).most_common() if i[1] > 1]
sentences = [i for i in trigram_tokens]
return frequent_words, sentences
def get_summary(text, limit=3):
sents = sent_tokenize(text)
frequent_words, sentences = split_sentences(sents)
sentence_scores = score_sentences(frequent_words, sentences)
limited_sents = [sents[num] for num, count in sentence_scores[:limit]]
best_sents = [i[0] for i in sorted([(i, text.find(i)) for i in limited_sents], key=lambda x: x[0])]
return best_sents
def summarize(text, limit=3):
summary = get_summary(text, limit)
print(title)
print()
print(' '.join(summary))
# -
summarize(text, 2)
|
notebooks/NLP_Text_Summarization.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# <small><i>This notebook was prepared by [<NAME>](https://github.com/donnemartin). Source and license info is on [GitHub](https://github.com/donnemartin/interactive-coding-challenges).</i></small>
# # Solution Notebook
# ## Problem: Implement a graph.
#
# * [Constraints](#Constraints)
# * [Test Cases](#Test-Cases)
# * [Algorithm](#Algorithm)
# * [Code](#Code)
# * [Unit Test](#Unit-Test)
# ## Constraints
#
# * Is the graph directed?
# * Yes
# * Do the edges have weights?
# * Yes
# ## Test Cases
#
# Input:
# * `add_edge(source, destination, weight)`
#
# ```
# graph.add_edge(0, 1, 5)
# graph.add_edge(0, 5, 2)
# graph.add_edge(1, 2, 3)
# graph.add_edge(2, 3, 4)
# graph.add_edge(3, 4, 5)
# graph.add_edge(3, 5, 6)
# graph.add_edge(4, 0, 7)
# graph.add_edge(5, 4, 8)
# graph.add_edge(5, 2, 9)
# ```
#
# Result:
# * `source` and `destination` nodes within `graph` are connected with specified `weight`.
#
# Note:
# * We'll be using an OrderedDict to make the outputs more consistent and simplify our testing.
# * Graph will be used as a building block for more complex graph challenges.
# ## Algorithm
#
# ### Node
#
# Node will keep track of its:
# * id
# * visit state
# * adjacent
# * key: node
# * value: weight
#
# ### Graph
#
# Graph will keep track of its:
# * nodes
# * key: node id
# * value: node
#
# #### add_node
#
# * Create a node with the input id
# * Add the newly created node to the list of nodes
#
# Complexity:
# * Time: O(1)
# * Space: O(1)
#
# #### add_edge
#
# * If the source node is not in the list of nodes, add it
# * If the dest node is not in the list of nodes, add it
# * Add a connection from the source node to the dest node with the given edge weight
#
# Complexity:
# * Time: O(1)
# * Space: O(1)
# ## Code
# +
# %%writefile graph.py
from collections import OrderedDict
from enum import Enum # Python 2 users: Run pip install enum34
class State(Enum):
unvisited = 1
visited = 2
visiting = 3
class Node:
def __init__(self, id):
self.id = id
self.visit_state = State.unvisited
self.adjacent = OrderedDict() # key = node, val = weight
def __str__(self):
return str(self.id)
class Graph:
def __init__(self):
self.nodes = OrderedDict() # key = node id, val = node
def add_node(self, id):
node = Node(id)
self.nodes[id] = node
return node
def add_edge(self, source, dest, weight=0):
if source not in self.nodes:
self.add_node(source)
if dest not in self.nodes:
self.add_node(dest)
self.nodes[source].adjacent[self.nodes[dest]] = weight
# -
# %run graph.py
# ## Unit Test
# +
# %%writefile test_graph.py
from nose.tools import assert_equal
class TestGraph(object):
def test_graph(self):
graph = Graph()
for id in range(0, 6):
graph.add_node(id)
graph.add_edge(0, 1, 5)
graph.add_edge(0, 5, 2)
graph.add_edge(1, 2, 3)
graph.add_edge(2, 3, 4)
graph.add_edge(3, 4, 5)
graph.add_edge(3, 5, 6)
graph.add_edge(4, 0, 7)
graph.add_edge(5, 4, 8)
graph.add_edge(5, 2, 9)
assert_equal(graph.nodes[0].adjacent[graph.nodes[1]], 5)
assert_equal(graph.nodes[0].adjacent[graph.nodes[5]], 2)
assert_equal(graph.nodes[1].adjacent[graph.nodes[2]], 3)
assert_equal(graph.nodes[2].adjacent[graph.nodes[3]], 4)
assert_equal(graph.nodes[3].adjacent[graph.nodes[4]], 5)
assert_equal(graph.nodes[3].adjacent[graph.nodes[5]], 6)
assert_equal(graph.nodes[4].adjacent[graph.nodes[0]], 7)
assert_equal(graph.nodes[5].adjacent[graph.nodes[4]], 8)
assert_equal(graph.nodes[5].adjacent[graph.nodes[2]], 9)
print('Success: test_graph')
def main():
test = TestGraph()
test.test_graph()
if __name__ == '__main__':
main()
# -
# %run -i test_graph.py
|
graphs_trees/graph/graph_solution.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <h1>Fantasy Basketball Part Two </h1>
# <h2> Introduction </h2>
#
# If you're familiar with fantasy basketball and worked through our **first example** you may have noticed that the lineup we solved for doesn't actually fit the criteria for many popular contests. It was meant as a ground-floor example into mathematical optimization, so make sure to dive into that one first for background on the problem. Additionally, we developed a predictive model to forecast each player's fantasy points, and we'll again use that forecast (without going through the modeling process again).
#
# In this follow-up example, we will extend the optimization model to fulfill the requirements of a typical [DraftKings](https://www.draftkings.com/help/rules/nba) lineup. Specifically, we'll need to change the model to:
# - Allow some players to be selected for up to two positions (e.g. LeBron James can fill the PF or SF position)
# - Increase the roster from five to eight players
# - Ensure the three new roster slots consist of one guard (SG or PG), one forward (SF or PF), and one utility player (any position)
#
# A quick recap of what we saw in the prior example: When selecting a 5-player lineup, there were *a lot* of possibilities; staying under a salary cap wasn't all that easy; and implementing intuitive decision rules can lead to suboptimal lineups -- providing a glimpse into why mathematical optimization should be part of everyone's analytics toolkit.
#
# **The repository for this project can be accessed by following this link:**
#
# <h2> Objective and Prerequisites </h2>
#
# As with the first example, the goal here is to build upon our findings of the previous example and select the optimal lineup of players of the National Basketball Association (NBA) that would produce the highest total of fantasy points, which are composed of a player's in-game stats.
#
# As mentioned above, we will make a lineup that meets DraftKings contest rules. There were over 2 million possible lineups when selecting a 5-player team that contained one of each position. Can you figure out the number of possible lineups for this example? Here's a hint: It's a lot. Remember that there are 25 PGs, 23 SFs, 22 SGs, 19 PFs, and 9 Cs. The answer (approximately) will be given a bit later.
#
# Also as mentioned, this example will use the same forecasting results as our fantasy basketball beginner's example, which used historical data from the 2016-2017 and 2017-2018 seasons to predict each player's fantasy points for games on 12/25/2017.
#
# This example assumes you have experience using Python for data manipulation and requires the installation of the following packages:
#
# - **pandas**: for data analysis and manipulation
# - **math**: for mathematical manipulations
# - **gurobipy**: for utilizing Gurobi to formulate and solve the optimization problem
#
# We'll also explore a few different ways to write summations and constraints in gurobipy, so you can find what works best for you. A quick note: Any output you see similar to <gurobi.Constr \*Awaiting Model Update\*> can be ignored and semi-colons have been added to cells to suppress that output.
# <h2> Problem Statement and Solution Approach</h2>
#
# By formuating a model that allows some players to be assigned to more than one position and by also expanding the roster, we are making the model much more complex.
#
# Our final lineup needs to include each of the following:
# - Point Guard (PG)
# - Shooting Guard (SG)
# - Shooting Forward (SF)
# - Power Forward (PF)
# - Center (C)
# - Guard (SG,PG)
# - Forward (SF,PF)
# - Utility (PG, SG, SF, PF, C)
#
# The solution of the problem consists of two components: 1) **fantasy points forecast** and 2) **lineup optimization**.
#
# We'll start by loading a dataset for the eligible players, showing their potential positions, salaries, as well as the forecasts for their upcoming performance. This information will act as an input to our optimization model which will guarantee that we satisfy the rules of the DraftKings contest while maximizing the total fantasy points of our team.
# <h3> Fantasy Points Forecast </h3>
#
# This section will be short as we are using the output of the predictive model from the first part of this example. We'll see that in this version some players have a *Main Position* and *Alternative Position*, which will let players fill different position slots if eligible. For example, <NAME> can fill the PG or SG position, though <NAME> is only available as a PG.
#
# We begin by loading the necessary libraries to solve our problem and importing the data from the predictive model in part one of the problem.
# %pip install gurobipy
import pandas as pd #importing pandas
import math #importing math
from gurobipy import Model, GRB, quicksum #importing Gurobi
player_predictions = pd.read_csv('https://raw.githubusercontent.com/Gurobi/modeling-examples/master/fantasy_basketball_1_2/results_target_advanced.csv') #load processed dataset
player_predictions.sort_values(by='PredictedFantasyPoints',ascending=False).head(20)
# Have you figured out the total number of possible lineups yet? It's around $3.6 \times 10^{11}$, which is a lot.
#
# This is where mathematical optimization and Gurobi are best utilized: To efficiently explore a huge decision space and provide a much needed tool for optimal decision-making.
# <h3> Optimal DraftKings Lineup Selection </h3>
#
# As we set up our optimization model, we first need to make some definitions. Some of this is the same as before, but as this example is a bit more complicated, we'll need to be a bit more thorough in some definitions.
#
# **Sets and Indices**
#
# $i$ is the index for the set of all players
#
# $j$ is the index for the set of basketball positions (PG,SG,SF,PF,C)
#
# **Input Parameters**
#
# $p_{i}$: the predicted fantasy points of player $i$
#
# $s_{i}$: the salary of player $i$
#
# $S$: our total available salary
# +
players = player_predictions["Player"].tolist()
positions = player_predictions["MainPosition"].unique().tolist()
salaries = player_predictions["Salary"].tolist()
fantasypoints = player_predictions["PredictedFantasyPoints"].tolist()
S = 50000
salary_dict = {players[i]: salaries[i] for i in range(len(players))}
points_dict = {players[i]: fantasypoints[i] for i in range(len(players))}
m = Model()
# -
# **Decision Variables**
#
# Since it is possible for certain players to fill one of two positions, we need to map each player to their eligible positions. Additionally, we need to add the position index to our decision variable. Instead of a binary variable (i.e. a variable that only takes the values of 0 or 1) $y_i$ we have $y_{i,j}$.
#
# $y_{i,j}$: This variable is equal to 1 if player $i$ is selected at position $j$; and 0 otherwise.
# +
mainposition = list(zip(player_predictions.Player, player_predictions.MainPosition))
alternativeposition = list(zip(player_predictions.Player, player_predictions.AlternativePosition))
indices = mainposition+alternativeposition
player_pos_map = [t for t in indices if not any(isinstance(n, float) and math.isnan(n) for n in t)]
y = m.addVars(player_pos_map, vtype=GRB.BINARY, name="y")
# -
# **Objective Function**
#
# The objective function of our problem is to maximize the total fantasy points of our lineup, same as the last time but using the differently indexed decision variable.
#
# \begin{align}
# Max \hspace{0.2cm} Z = \sum_{i,j} p_{i} \cdot y_{i,j}
# \end{align}
m.setObjective(quicksum([points_dict[i]*y[i,j] for i,j in player_pos_map]), GRB.MAXIMIZE)
# **Constraints**
#
# Our model still requires each of the primary basketball positions (PG, SG, SF, PF, C) to be filled. Last time we required exactly one of each, so these constraints were equalities (note that this also implied our roster size of five players). In this version of our model, we need *at least one* of each position since it is possible to use, for example, only one power forward (PF).
#
# For each position $j$:
# \begin{align}
# \sum_{i} y_{i,j} \geq 1
# \end{align}
#
# Here is where we have a couple of options in how to add these constraints. The first, intuitively, is to loop through the set of positions. The second uses a slightly different function to add the constraints (*addConstr* -> *addConstrs*). This function incorporates the for loop directly as an argument. **Running both will double up on the constraints in the model, which isn't best practice, so take a look at each approach and comment one out before running the cell.**
# +
# option 1 for writing the above constraints
for j in positions:
m.addConstr(quicksum([y[i,j] for i, pos in player_pos_map if pos==j])>=1, name = "pos" + j)
# option 2, a slightly more compact way of adding the same set of constraints
m.addConstrs((quicksum([y[i,j] for i, pos in player_pos_map if pos==j])>=1 for j in positions), name = "pos");
# -
# Now let's work with the additional slots of our new roster: the guard, forward, and utility slots. Here it's worth mentioning that there are different ways to formulate the same problem in mathematical optimization. Some approaches can be better than others and writing efficient models will become an important skill as you tackle larger and more complex problems.
#
# Let's first consider the additional slot for the guard position, which can be filled by a PG or a SG. Considering the overall roster we want to create, we have already guaranteed one PG and one SG (so two total guards). To make sure we get one more additional guard, a constraint needs to added that says the total number of guards needs to be *at least* three.
#
# \begin{align}
# \sum_{i} y_{i,j} \geq 3, position\space j \space is\space PG\space or \space SG
# \end{align}
#
# The same needs to be done for forwards (SF or PF)
# \begin{align}
# \sum_{i} y_{i,j} \geq 3, position\space j \space is\space SF\space or \space PF
# \end{align}
#
# It is important that we use inequalities here because of the utility position we'll address in a little bit.
m.addConstr(quicksum([y[i,j] for i, j in player_pos_map if (j=='PG' or j=='SG')])>=3)
m.addConstr(quicksum([y[i,j] for i, j in player_pos_map if (j=='SF' or j=='PF')])>=3);
# Now that we have a position index for our decision variable, we need to ensure each player is assigned to *at most* one position (either their primary or alternative, but not both). To do this we sum across each position $j$ for each player and limit that summation to one.
#
# For each player $i$,
# \begin{align}
# \sum_{j} y_{i,j} \leq 1
# \end{align}
#
# Here we'll use another way to sum over one of the variable's indices by appending *.sum* to the end and replacing the index we want to sum over with "\*". This is useful because each player is not eligible for each position and this syntax automatically sums over the second index.
m.addConstrs((y.sum(i, "*") <= 1 for i in players), name="max_one");
# A good exercise would be to rewrite other summations using this syntax and to try and the above constraint using quicksum.
#
# So far we have addressed seven of our eight lineup slots. Since the utility slot can have any position, all we need is to require the total number of players selected to be eight by setting the sum over all players and positions to that value.
#
# \begin{align}
# \sum_{i,j} y_{i,j} = 8
# \end{align}
m.addConstr(quicksum([y[i,j] for i,j in player_pos_map]) == 8, name="full_lineup");
# Finally, we need to stay under the salary cap $S$, which in a typical DraftKings contest is $\$50,000$:
#
# \begin{align}
# \sum_{i,j} s_{i} \cdot y_{i,j} \leq S
# \end{align}
cap = m.addConstr(quicksum(salary_dict[i]*y[i,j] for i,j in player_pos_map) <= S, name="salary")
# In this last constraint, we stored it to be able to easily get information about this constraint after the model runs and the model is updated just as if we didn't store the constraint. Time to find the optimal lineup.
m.optimize() # optimize our model
# Notice the output above says we have 167 binary variables, which is much less than $98 \cdot 5 = 490$ variables we'd have if we mapped all players to all basketball positions. Additionally, we would need more constraints to eliminate ineligible player/position combinations. While this wouldn't make much difference in this small example, formulating efficient models is a valuable skill in mathematical optimization.
#
# Let's display our optimal lineup.
# +
player_selections = []
for v in m.getVars():
if (abs(v.x) > 1e-6):
player_selections.append(tuple(y)[v.index])
df = pd.DataFrame(player_selections, columns = ['Player','Assigned Position'])
df = df.merge(pd.DataFrame(list(salary_dict.items()), columns=['Player', 'Salary']), left_on=['Player'], right_on=['Player'])
lineup = df.merge(pd.DataFrame(list(points_dict.items()), columns=['Player', 'Predicted Points']), left_on=['Player'], right_on=['Player'])
lineup.sort_values(by=['Assigned Position'])
# -
print('Total fantasy score: ', round(m.objVal,2))
print('Remaining salary: ', cap.Slack)
# The last print statement uses the constraint we stored earlier. The *Slack* attribute of a constraint will show any gap between each side of the inequality. For this application, it's the gap between the salary cap and the amount of salary used in the optimal lineup, which shows there is $600 in unused salary.
# <h2> Conclusion </h2>
#
# In this notebook we finished up a problem that started from a raw data set containing NBA player box score data and ended in, if our predictive model holds, an optimal fantasy basketball lineup.
#
# Specifically in this part, we:
# - Expanded the initial model to reflect the true complexity of creating a fantasy basketball lineup
# - Discovered multiple ways to add a set of constraints to a model
# - Utilized two summation commands
# - Used attributes of a model to get more information about the optimal solution
#
# Overall, this two-part example displayed how, even with the best predictive model, making optimal decisions is still a complicated exercise. Along with machine learning techniques, mathematical optimization is an essential part of a well-developed analytical toolbox.
|
fantasy_basketball_1_2/fantasy_basketball_part2_gcl.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import requests as rq
from bs4 import BeautifulSoup
# # Test on one link
url = 'https://en.wikipedia.org//wiki/Operation_Skorpion'
# url = 'https://en.wikipedia.org//wiki/Siege_of_Giarabub'
# url = 'https://en.wikipedia.org/wiki/Siege_of_Giarabub'
# url = 'https://en.wikipedia.org/wiki/Battle_of_Dakar'
# url = 'https://en.wikipedia.org/wiki/Vilnius_Offensive'
url
def get_dom(url):
response = rq.get(url)
response.raise_for_status()
return BeautifulSoup(response.content, 'html.parser')
info = get_dom(url)
# ## Processing one link
#
# - actors
# - date
# - geolocation
# - who wins
# - casualities
# - map?
table = info.find('table','infobox vevent')
def _table_to_dict(table):
result = {}
for row in table.find_all('tr'):
key = next(row.th.stripped_strings)
value = row.td.get_text().strip()
result[key] = value
return result
def _get_main_info(table):
main = [el for el in table.tbody.find_all('tr', recursive=False) if 'Location' in el.get_text()][0]
return {'main': _table_to_dict(main) }
_get_main_info(table)
# # Additional
# +
def _parse_row(row):
'''parse secondory info row
as dict of info points - for allies and axis
'''
cells = row.find_all('td', recursive=False)
return [cell.get_text().strip() for cell in cells]
def _find_row_by_header(table, string):
header = table.tbody.find('tr', text=string)
if header is not None:
return header.next_sibling
def _additional(table):
keywords = (
'Belligerents',
'Commanders and leaders',
'Strength',
'Casualties and losses',
)
result = {}
for keyword in keywords:
try:
data = _find_row_by_header(table, keyword)
if data:
result[keyword] = _parse_row(data)
except Exception as e:
raise Exception(keyword, e)
return result
# -
_additional(table)
# ## Test on a few
urls = {
'Dakar': 'https://en.wikipedia.org/wiki/Battle_of_Dakar',
'Brest': 'https://en.wikipedia.org/wiki/Battle_for_Brest',
'Torpedo Alley': 'https://en.wikipedia.org/wiki/Torpedo_Alley',
'Moravo': 'https://en.wikipedia.org/wiki/Battle_of_Morava%E2%80%93Ivan'
}
# +
def _parse_battle_page(url):
print(url)
dom = _default_collect(url)
dom['url'] = url
table = dom.find('table','infobox vevent')
if table is None:
return {}
data = _get_main_info(table)
additional = _additional(table)
data.update(additional)
return data
# -
_parse_battle_page(url)
result = {k:_parse_battle_page(v) for k, v in urls.items()}
result['Moravo']
result['Brest']
|
Chapter07/7_Scraping_part_2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python3
# ---
# <script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
# <script>
# window.dataLayer = window.dataLayer || [];
# function gtag(){dataLayer.push(arguments);}
# gtag('js', new Date());
#
# gtag('config', 'UA-59152712-8');
# </script>
#
# # Two Formulations of Maxwell's equations in Cartesian Coordinates
#
# ## Authors: <NAME> & <NAME>
# ### Formatting improvements courtesy <NAME>
#
# [comment]: <> (Abstract: TODO)
#
# **Notebook Status:** <font color='orange'><b> Self-Validated </b></font>
#
# **Validation Notes:** This tutorial notebook has been confirmed to be self-consistent with its corresponding NRPy+ module, as documented below, [here](#code_validation_sys1) and [here](#code_validation). **Additional validation tests may have been performed, but are as yet, undocumented. (TODO)**
#
# ### NRPy+ Source Code for this module:
# * [Maxwell/MaxwellCartesian_Evol.py](../edit/Maxwell/MaxwellCartesian_Evol.py)
# * [Maxwell/MaxwellCartesian_ID.py](../edit/Maxwell/MaxwellCartesian_ID.py)
#
# ## Introduction:
#
# This tutorial will draw on previous work done by <NAME> on [Maxwell's equations in Curvilinear Coordinates](Tutorial-MaxwellCurvilinear.ipynb), which itself drew on the two formulations described in [Illustrating Stability Properties of Numerical Relativity in Electrodynamics](https://arxiv.org/abs/gr-qc/0201051). This will be done to aid construction of an Einstein Toolkit thorn, which will itself be built in the [Tutorial-ETK_thorn-Maxwell](Tutorial-ETK_thorn-Maxwell.ipynb) tutorial notebook.
#
# The construction of our equations here will be nearly identical; however, by assuming Cartesian coordinates, we are able to make several simplifications and eliminate the need for [reference_metric.py](../edit/reference_metric.py) as a dependency.
#
# We will begin with their System I. While the Curvilinear version of this code assumed flat spacetime, we will be constructing the equations in a general spacetime. This allows us to simply replace the reference metric "hatted" quantities with their general counterparts.
# <a id='toc'></a>
#
# # Table of Contents:
# $$\label{toc}$$
#
# This notebook is organized as follows
#
# 1. [Step 1](#initializenrpy): Import core NRPy+ modules and set NRPy+ parameters
# 1. [Step 2](#laplacian): Constructing the Covariant Laplacian
# 1. [Step 3](#violation): Measuring the Constraint violation
# 1. [Step 4](#code_validation_sys1): Code Validation against `Maxwell.MaxwellCartesian_Evol` NRPy+ Module (System I)
# 1. [Step 5](#code_validation_sys2): Code Validation against `Maxwell.MaxwellCartesian_Evol` NRPy+ Module (System II)
# 1. [Step 6](#id): Constructing the Initial Data
# 1. [Step 7](#code_validation): Code Validation against `Maxwell.MaxwellCartesian_ID` NRPy+ Module
# 1. [Step 8](#latex_pdf_output): Output this notebook to $\LaTeX$-formatted PDF file
# <a id='initializenrpy'></a>
#
# # Step 1: Import core NRPy+ modules and set NRPy+ parameters \[Back to [top](#toc)\]
# $$\label{initializenrpy}$$
# +
#Step P1: Import needed Python modules
import NRPy_param_funcs as par
import indexedexp as ixp
import grid as gri
import finite_difference as fin
from outputC import *
#Step P2: Name this Python module
thismodule = "MaxwellCartesian"
#Step 0: Set the spatial dimension parameter to 3.
par.set_parval_from_str("grid::DIM", 3)
DIM = par.parval_from_str("grid::DIM")
# Step 1: Set the finite differencing order to 4.
par.set_parval_from_str("finite_difference::FD_CENTDERIVS_ORDER", 4)
# Step 2: Register gridfunctions that are needed as input.
psi = gri.register_gridfunctions("EVOL", ["psi"])
# Step 3a: Declare the rank-1 indexed expressions E_{i}, A_{i},
# and \partial_{i} \psi. Derivative variables like these
# must have an underscore in them, so the finite
# difference module can parse the variable name properly.
ED = ixp.register_gridfunctions_for_single_rank1("EVOL", "ED")
AD = ixp.register_gridfunctions_for_single_rank1("EVOL", "AD")
psi_dD = ixp.declarerank1("psi_dD")
x,y,z = gri.register_gridfunctions("AUX",["x","y","z"])
## Step 3b: Declare the conformal metric tensor and its first
# derivative. These are needed to find the Christoffel
# symbols, which we need for covariant derivatives.
gammaDD = ixp.register_gridfunctions_for_single_rank2("AUX","gammaDD", "sym01") # The AUX or EVOL designation is *not*
# used in diagnostic modules.
gammaDD_dD = ixp.declarerank3("gammaDD_dD","sym01")
gammaDD_dDD = ixp.declarerank4("gammaDD_dDD","sym01_sym23")
gammaUU = ixp.declarerank3("gammaUU","sym01")
detgamma = gri.register_gridfunctions("AUX",["detgamma"])
gammaUU, detgamma = ixp.symm_matrix_inverter3x3(gammaDD)
gammaUU_dD = ixp.declarerank3("gammaDD_dD","sym01")
# Define the Christoffel symbols
GammaUDD = ixp.zerorank3(DIM)
for i in range(DIM):
for k in range(DIM):
for l in range(DIM):
for m in range(DIM):
GammaUDD[i][k][l] += (sp.Rational(1,2))*gammaUU[i][m]*\
(gammaDD_dD[m][k][l] + gammaDD_dD[m][l][k] - gammaDD_dD[k][l][m])
# Step 3b: Declare the rank-2 indexed expression \partial_{j} A_{i},
# which is not symmetric in its indices.
# Derivative variables like these must have an underscore
# in them, so the finite difference module can parse the
# variable name properly.
AD_dD = ixp.declarerank2("AD_dD", "nosym")
# Step 3c: Declare the rank-3 indexed expression \partial_{jk} A_{i},
# which is symmetric in the two {jk} indices.
AD_dDD = ixp.declarerank3("AD_dDD", "sym12")
# Step 4: Calculate first and second covariant derivatives, and the
# necessary contractions.
# First covariant derivative
# D_{j} A_{i} = A_{i,j} - \Gamma^{k}_{ij} A_{k}
AD_dcovD = ixp.zerorank2()
for i in range(DIM):
for j in range(DIM):
AD_dcovD[i][j] = AD_dD[i][j]
for k in range(DIM):
AD_dcovD[i][j] -= GammaUDD[k][i][j] * AD[k]
# -
# <a id='laplacian'></a>
#
# # Step 2: Constructing the Covariant Laplacian \[Back to [top](#toc)\]
# $$\label{laplacian}$$
#
# One particular difficulty we will encounter here is taking the covariant Laplacian of a vector. This will here take the form of $D_j D^j A_i$. We will start with the outer derivative after lowering the index on the second operator. So, we see that
# \begin{align}
# D_j D^j A_i &= D_j (\gamma^{jk} D_k A_i) \\
# &= (D_k A_i) D_j \gamma^{jk} + \gamma^{jk} D_j D_k A_i \\
# &= \gamma^{jk} [\partial_j (D_k A_i) - \Gamma^l_{ij} D_k A_l - \Gamma^l_{jk} D_l A_i],
# \end{align}
# dropping the first term from the second line because $D_j \gamma^{jk} = 0$ Next, we will again apply the covariant derivative to $A_i$. First, however, we should consider that
# \begin{align}
# D_k A_i &= \partial_k A_i - \Gamma^l_{ik} A_l \\
# & = \partial_k A_i - \gamma^{lm} \Gamma_{mik} A_l \\
# & = \partial_k A_i - \Gamma_{mik} A^m, \\
# \end{align}
# where $\Gamma_{ljk} = \frac{1}{2} (\partial_k \gamma_{lj} + \partial_j \gamma_{kl} - \partial_l \gamma_{jk})$ is the Christoffel symbol of the first kind. Note how we were able to use the raising operator to switch the height of two indices; we will use this in upcoming steps. Thus, our expression becomes
# \begin{align}
# D_j D^j A_i &= \gamma^{jk} [\partial_j \partial_k A_i - \partial_j (\Gamma_{lik} A^l) - \Gamma^l_{ij} \partial_k A_l + \Gamma^m_{ij} \Gamma_{lmk} A^l - \Gamma^l_{jk} \partial_l A_i + \Gamma^m_{jk} \Gamma_{lim} A^l] \\
# &= \gamma^{jk} [\partial_j \partial_k A_i - \underbrace{\partial_j (\Gamma_{lik} A^l)}_{\text{sub-term}} - \Gamma^l_{ij} \partial_k A_l + \Gamma^m_{ij} \Gamma^l_{mk} A_l - \Gamma^l_{jk} \partial_l A_i + \Gamma^m_{jk} \Gamma^l_{im} A_l].
# \end{align}
# Let's focus on the underbraced sub-term for a moment. Expanding this using the product rule and the definition of the Christoffel symbol,
# \begin{align}
# \partial_j (\Gamma_{lik} A^l) &= A^l \partial_j \Gamma_{lik} + \Gamma_{lik} \partial_j A^l \\
# &= A^l \partial_j (\partial_k \gamma_{li} + \partial_i \gamma_{kl} - \partial_l \gamma_{ik}) + \Gamma_{lik} \partial_j (\gamma^{lm} A_m) \\
# &= A^l (\gamma_{li,kj} + \gamma_{kl,ij} - \gamma_{ik,lj}) + \Gamma_{lik} (\gamma^{lm} A_{m,j} + A_m \gamma^{lm}{}_{,j}), \\
# \end{align}
# where commas in subscripts denote partial derivatives.
#
# So, the Laplacian becomes
# \begin{align}
# D_j D^j A_i &= \gamma^{jk} [A_{i,jk} -
# \underbrace{A^l (\gamma_{li,kj} + \gamma_{kl,ij} - \gamma_{ik,lj})}_{\text{Term 1}} +
# \underbrace{\Gamma_{lik} (\gamma^{lm} A_{m,j} + A_m \gamma^{lm}{}_{,j})}_{\text{Term 2}} -
# \underbrace{(\Gamma^l_{ij} A_{l,k} + \Gamma^l_{jk} A_{i,l})}_{\text{Term 3}} +
# \underbrace{(\Gamma^m_{ij} \Gamma^l_{mk} A_l + \Gamma ^m_{jk} \Gamma^l_{im} A_l)}_{\text{Term 4}}]; \\
# \end{align}
# we will now begin to contruct these terms individually.
# +
# First, we must construct the lowered Christoffel symbols:
# \Gamma_{ijk} = \gamma_{il} \Gamma^l_{jk}
# And raise the index on A:
# A^j = \gamma^{ij} A_i
GammaDDD = ixp.zerorank3()
AU = ixp.zerorank1()
for i in range(DIM):
for j in range(DIM):
AU[j] += gammaUU[i][j] * AD[i]
for k in range(DIM):
for l in range(DIM):
GammaDDD[i][j][k] += gammaDD[i][l] * GammaUDD[l][j][k]
# Covariant second derivative (the bracketed terms):
# D_j D^j A_i = \gamma^{jk} [A_{i,jk} - A^l (\gamma_{li,kj} + \gamma_{kl,ij} - \gamma_{ik,lj})
# + \Gamma_{lik} (\gamma^{lm} A_{m,j} + A_m \gamma^{lm}{}_{,j})
# - (\Gamma^l_{ij} A_{l,k} + \Gamma^l_{jk} A_{i,l})
# + (\Gamma^m_{ij} \Gamma^l_{mk} A_l + \Gamma ^m_{jk} \Gamma^l_{im} A_l)
AD_dcovDD = ixp.zerorank3()
for i in range(DIM):
for j in range(DIM):
for k in range(DIM):
AD_dcovDD[i][j][k] = AD_dDD[i][j][k]
for l in range(DIM):
# Terms 1 and 3
AD_dcovDD[i][j][k] -= AU[l] * (gammaDD_dDD[l][i][k][j] + gammaDD_dDD[k][l][i][j] - \
gammaDD_dDD[i][k][l][j]) \
+ GammaUDD[l][i][j] * AD_dD[l][k] + GammaUDD[l][j][k] * AD_dD[i][l]
for m in range(DIM):
# Terms 2 and 4
AD_dcovDD[i][j][k] += GammaDDD[l][i][k] * (gammaUU[l][m] * AD_dD[m][j] + AD[m] * gammaUU_dD[l][m][j]) \
+ GammaUDD[m][i][j] * GammaUDD[l][m][k] * AD[l] \
+ GammaUDD[m][j][k] * GammaUDD[l][i][m] * AD[l]
# Covariant divergence
# D_{i} A^{i} = \gamma^{ij} D_{j} A_{i}
DivA = 0
# Gradient of covariant divergence
# DivA_dD_{i} = \gamma^{jk} A_{k;\hat{j}\hat{i}}
DivA_dD = ixp.zerorank1()
# Covariant Laplacian
# LapAD_{i} = \gamma^{jk} A_{i;\hat{j}\hat{k}}
LapAD = ixp.zerorank1()
for i in range(DIM):
for j in range(DIM):
DivA += gammaUU[i][j] * AD_dcovD[i][j]
for k in range(DIM):
DivA_dD[i] += gammaUU[j][k] * AD_dcovDD[k][j][i]
LapAD[i] += gammaUU[j][k] * AD_dcovDD[i][j][k]
# Step 5: Define right-hand sides for the evolution.
ArhsD = ixp.zerorank1()
ErhsD = ixp.zerorank1()
for i in range(DIM):
ArhsD[i] = -ED[i] - psi_dD[i]
ErhsD[i] = -LapAD[i] + DivA_dD[i]
psi_rhs = -DivA
# Step 6: Generate C code for System I Maxwell's evolution equations,
# print output to the screen (standard out, or stdout).
#lhrh_list = []
#for i in range(DIM):
# lhrh_list.append(lhrh(lhs=gri.gfaccess("rhs_gfs", "AD" + str(i)), rhs=ArhsD[i]))
# lhrh_list.append(lhrh(lhs=gri.gfaccess("rhs_gfs", "ED" + str(i)), rhs=ErhsD[i]))
#lhrh_list.append(lhrh(lhs=gri.gfaccess("rhs_gfs", "psi"), rhs=psi_rhs))
#fin.FD_outputC("stdout", lhrh_list)
# -
# <a id='violation'></a>
#
# # Step 3: Measuring the Constraint violation \[Back to [top](#toc)\]
# $$\label{violation}$$
#
# To evaluate our results, we will need to measure how much our simulation differs from what should be physically possible. We will do this with the constraint equation $D_i E^i = 4 \pi \rho_e$; specifically, we will measure the constraint violation
# \begin{align}
# \mathcal{C} &= D_i E^i - 4 \pi \rho_e \\
# &= D_i (\gamma^{ij} E_j) - 4 \pi \rho_e \\
# &= \gamma^{ij} D_i E_j + E_j D_i \gamma^{ij} - 4 \pi \rho_e \\
# &= \gamma^{ij} D_i E_j,
# \end{align}
# since the covariant derivative of the metric tensor is $0$ and $\rho_e=0$ in free space. So, $\mathcal{C} = \gamma^{ij} (E_{j,i} - \Gamma^b_{ij} E_b)$, which will be valid for both systems.
ED_dD = ixp.declarerank2("ED_dD","nosym")
Cviolation = gri.register_gridfunctions("AUX", ["Cviolation"])
Cviolation = sp.sympify(0)
for i in range(DIM):
for j in range(DIM):
Cviolation += gammaUU[i][j] * ED_dD[j][i]
for b in range(DIM):
Cviolation -= gammaUU[i][j] * GammaUDD[b][i][j] * ED[b]
# <a id='code_validation_sys1'></a>
#
# # Step 4: Code Validation against `Maxwell.MaxwellCartesian_Evol` NRPy+ Module (System I) \[Back to [top](#toc)\]
# $$\label{code_validation_sys1}$$
#
# Here, as a code validation check, we verify agreement in the SymPy expressions for the RHSs of Maxwell's equations (in System I) between
#
# 1. this tutorial and
# 2. the NRPy+ [Maxwell.MaxwellCartesian](../edit/Maxwell/MaxwellCartesian_Evol.py) module.
#
# +
# Reset the list of gridfunctions, as registering a gridfunction
# twice will spawn an error.
gri.glb_gridfcs_list = []
# Step 18: Call the MaxwellCartesian_Evol() function from within the
# Maxwell/MaxwellCartesian_Evol.py module,
# which should do exactly the same as in Steps 1-16 above.
import Maxwell.MaxwellCartesian_Evol as mwevol
par.set_parval_from_str("System_to_use","System_I")
mwevol.MaxwellCartesian_Evol()
print("Consistency check between MaxwellCartesian tutorial and NRPy+ module: ALL SHOULD BE ZERO.")
print("psi_rhs - mwevol.psi_rhs = " + str(psi_rhs - mwevol.psi_rhs))
for i in range(DIM):
print("ArhsD["+str(i)+"] - mwevol.ArhsD["+str(i)+"] = " + str(ArhsD[i] - mwevol.ArhsD[i]))
print("ErhsD["+str(i)+"] - mwevol.ErhsD["+str(i)+"] = " + str(ErhsD[i] - mwevol.ErhsD[i]))
print("Cviolation - mwevol.Cviolation = " + str(Cviolation - mwevol.Cviolation))
# -
# We will now build the equations for System II.
# +
# We inherit here all of the definitions from System I, above
# Step 7a: Register the scalar auxiliary variable \Gamma
Gamma = gri.register_gridfunctions("EVOL", ["Gamma"])
# Step 7b: Declare the ordinary gradient \partial_{i} \Gamma
Gamma_dD = ixp.declarerank1("Gamma_dD")
# Step 8a: Construct the second covariant derivative of the scalar \psi
# \psi_{;\hat{i}\hat{j}} = \psi_{,i;\hat{j}}
# = \psi_{,ij} - \Gamma^{k}_{ij} \psi_{,k}
psi_dDD = ixp.declarerank2("psi_dDD", "sym01")
psi_dcovDD = ixp.zerorank2()
for i in range(DIM):
for j in range(DIM):
psi_dcovDD[i][j] = psi_dDD[i][j]
for k in range(DIM):
psi_dcovDD[i][j] += - GammaUDD[k][i][j] * psi_dD[k]
# Step 8b: Construct the covariant Laplacian of \psi
# Lappsi = ghat^{ij} D_{j} D_{i} \psi
Lappsi = 0
for i in range(DIM):
for j in range(DIM):
Lappsi += gammaUU[i][j] * psi_dcovDD[i][j]
# Step 9: Define right-hand sides for the evolution.
ArhsD = ixp.zerorank1()
ErhsD = ixp.zerorank1()
for i in range(DIM):
ArhsD[i] = -ED[i] - psi_dD[i]
ErhsD[i] = -LapAD[i] + Gamma_dD[i]
psi_rhs = -Gamma
Gamma_rhs = -Lappsi
# Step 10: Generate C code for System II Maxwell's evolution equations,
# print output to the screen (standard out, or stdout).
#lhrh_list = []
#for i in range(DIM):
# lhrh_list.append(lhrh(lhs=gri.gfaccess("rhs_gfs", "AD" + str(i)), rhs=ArhsD[i]))
# lhrh_list.append(lhrh(lhs=gri.gfaccess("rhs_gfs", "ED" + str(i)), rhs=ErhsD[i]))
#lhrh_list.append(lhrh(lhs=gri.gfaccess("rhs_gfs", "psi"), rhs=psi_rhs))
#lhrh_list.append(lhrh(lhs=gri.gfaccess("rhs_gfs", "Gamma"), rhs=Gamma_rhs))
#fin.FD_outputC("stdout", lhrh_list)
# -
# <a id='code_validation_sys2'></a>
#
# # Step 5: Code Validation against `Maxwell.MaxwellCartesian_Evol` NRPy+ Module (System II) \[Back to [top](#toc)\]
# $$\label{code_validation_sys2}$$
#
# Here, as a code validation check, we verify agreement in the SymPy expressions for the RHSs of Maxwell's equations (in System II) between
#
# 1. this tutorial and
# 2. the NRPy+ [Maxwell.MaxwellCartesian](../edit/Maxwell/MaxwellCartesian_Evol.py) module.
#
# +
# Reset the list of gridfunctions, as registering a gridfunction
# twice will spawn an error.
gri.glb_gridfcs_list = []
# Step 18: Call the MaxwellCartesian_Evol() function from within the
# Maxwell/MaxwellCartesian_Evol.py module,
# which should do exactly the same as in Steps 1-16 above.
par.set_parval_from_str("System_to_use","System_II")
mwevol.MaxwellCartesian_Evol()
print("Consistency check between MaxwellCartesian tutorial and NRPy+ module: ALL SHOULD BE ZERO.")
print("psi_rhs - mwevol.psi_rhs = " + str(psi_rhs - mwevol.psi_rhs))
print("Gamma_rhs - mwevol.Gamma_rhs = " + str(Gamma_rhs - mwevol.Gamma_rhs))
for i in range(DIM):
print("ArhsD["+str(i)+"] - mwevol.ArhsD["+str(i)+"] = " + str(ArhsD[i] - mwevol.ArhsD[i]))
print("ErhsD["+str(i)+"] - mwevol.ErhsD["+str(i)+"] = " + str(ErhsD[i] - mwevol.ErhsD[i]))
print("Cviolation - mwevol.Cviolation = " + str(Cviolation - mwevol.Cviolation))
# -
# <a id='id'></a>
#
# # Step 6: Constructing the Initial Data \[Back to [top](#toc)\]
# $$\label{id}$$
#
# Now that we have evolution equations in place, we must construct the initial data that will be evolved by the solver of our choice. We will start from the analytic solution to this system of equations, given in [Illustrating Stability Properties of Numerical Relativity in Electrodynamics](https://arxiv.org/abs/gr-qc/0201051) as
# \begin{align}
# A^{\hat{\phi}} &= \mathcal{A} \sin \theta \left( \frac{e^{-\lambda v^2}-e^{-\lambda u^2}}{r^2} - 2 \lambda \frac{ve^{-\lambda v^2}-ue^{-\lambda u^2}}{r} \right), \\
# \end{align}
# for vanishing scalar potential $\psi$, where $\mathcal{A}$ gives the amplitude, $\lambda$ describes the size of the wavepacket, $u = t+r$, and $v = t-r$. Other components of this field are $0$.To get initial data, then, we simply set $t=0$; since $\psi=0$, $E_i = \partial_t A_i$. Thus, our initial data becomes the equations
# \begin{align}
# A^{\hat{\phi}} &= 0 \\
# E^{\hat{\phi}} &= 8 \mathcal{A} r \sin \theta \lambda^2 e^{-\lambda r^2} \\
# \psi &= 0
# \end{align}
# where the non-$\hat{\phi}$ components are set to 0. We still will need to convert $E^i$ in spherical-like coordinates and then lower its index. Using the standard transformations for coordinates and unit vectors,
# \begin{align}
# E^{\hat{x}} &= -\frac{y E^{\hat{\phi}}(x,y,z)}{\sqrt{x^2+y^2}} \\
# E^{\hat{y}} &= \frac{x E^{\hat{\phi}}(x,y,z)}{\sqrt{x^2+y^2}} \\
# E^{\hat{z}} &= 0. \\
# \end{align}
# We can lower the index in the usual way.
#
# For system II, we will also need to set initial data for $\Gamma$. Since $\Gamma = -\partial_t \psi$ and we have chosen $\psi(t=0) = 0$, $\Gamma(t=0) = 0$.
# +
# Step 1: Declare free parameters intrinsic to these initial data
amp,lam = par.Cparameters("REAL",thismodule,
["amp","lam"],
[1.0,1.0]) # __name__ = "MaxwellCartesian_ID", this module's name
# Step 2: Set the initial data
AidD = ixp.zerorank1()
EidD = ixp.zerorank1()
EidU = ixp.zerorank1()
# Set the coordinate transformations:
radial = sp.sqrt(x*x + y*y + z*z)
polar = sp.atan2(sp.sqrt(x*x + y*y),z)
EU_phi = 8*amp*radial*sp.sin(polar)*lam*lam*sp.exp(-lam*radial*radial)
EidU[0] = -(y * EU_phi)/sp.sqrt(x*x + y*y)
EidU[1] = (x * EU_phi)/sp.sqrt(x*x + y*y)
# The z component (2)is zero.
for i in range(DIM):
for j in range(DIM):
EidD[i] += gammaDD[i][j] * EidU[j]
psi_ID = sp.sympify(0)
Gamma_ID = sp.sympify(0)
# -
# <a id='code_validation'></a>
#
# # Step 7: Code Validation against `Maxwell.MaxwellCartesian_ID` NRPy+ Module \[Back to [top](#toc)\]
# $$\label{code_validation}$$
#
# Here, as a code validation check, we verify agreement in the SymPy expressions for the initial data we intend to use between
#
# 1. this tutorial and
# 2. the NRPy+ [Maxwell.MaxwellCartesian_ID](../edit/Maxwell/MaxwellCartesian_ID.py) module.
#
# Since the initial data is identical between the two systems for $E_i$, $A_i$, and $\psi$, so checking system I should be redundant; we will do it anyways, to be sure.
# +
# Reset the list of gridfunctions, as registering a gridfunction
# twice will spawn an error.
gri.glb_gridfcs_list = []
par.set_parval_from_str("System_to_use","System_I")
import Maxwell.MaxwellCartesian_ID as mwid
mwid.MaxwellCartesian_ID()
print("System I consistency check between MaxwellCartesian tutorial and NRPy+ module;\n ALL SHOULD BE ZERO:")
print("psi_ID - mwid.psi_ID = " + str(psi_ID - mwid.psi_ID))
for i in range(DIM):
print("AidD["+str(i)+"] - mwid.AidD["+str(i)+"] = " + str(AidD[i] - mwid.AidD[i]))
print("EidD["+str(i)+"] - mwid.EidD["+str(i)+"] = " + str(EidD[i] - mwid.EidD[i]))
# -
# Finally, we will repeat the check with system II initial data.
# +
# Reset the list of gridfunctions, as registering a gridfunction
# twice will spawn an error.
gri.glb_gridfcs_list = []
par.set_parval_from_str("System_to_use","System_II")
mwid.MaxwellCartesian_ID()
print("System II consistency check between MaxwellCartesian tutorial and NRPy+ module;\n ALL SHOULD BE ZERO:")
print("psi_ID - mwid.psi_ID = " + str(psi_ID - mwid.psi_ID))
print("Gamma_ID - mwid.Gamma_ID = " + str(Gamma_ID - mwid.Gamma_ID))
for i in range(DIM):
print("AidD["+str(i)+"] - mwid.AidD["+str(i)+"] = " + str(AidD[i] - mwid.AidD[i]))
print("EidD["+str(i)+"] - mwid.EidD["+str(i)+"] = " + str(EidD[i] - mwid.EidD[i]))
# -
# <a id='latex_pdf_output'></a>
#
# # Step 8: Output this notebook to $\LaTeX$-formatted PDF file \[Back to [top](#toc)\]
# $$\label{latex_pdf_output}$$
#
# The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename
# [Tutorial-MaxwellCartesian.pdf](Tutorial-MaxwellCartesian.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means.)
# !jupyter nbconvert --to latex --template latex_nrpy_style.tplx --log-level='WARN' Tutorial-MaxwellCartesian.ipynb
# !pdflatex -interaction=batchmode Tutorial-MaxwellCartesian.tex
# !pdflatex -interaction=batchmode Tutorial-MaxwellCartesian.tex
# !pdflatex -interaction=batchmode Tutorial-MaxwellCartesian.tex
# !rm -f Tut*.out Tut*.aux Tut*.log
|
Tutorial-MaxwellCartesian.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Single Source Shortest Path (SSSP)
#
# In this notebook, we will use cuGraph to compute the shortest path from a starting vertex to everyother vertex in our training dataset.
#
# Notebook Credits
# * Original Authors: <NAME> and <NAME>
# * available since rerlease 0.6
# * Last Edit: 10/28/2019
#
#
# RAPIDS Versions: 0.10.0
#
# Test Hardware
#
# * GV100 32G, CUDA 10.0
#
#
#
# ## Introduction
#
# Single source shortest path computes the shortest paths from the given starting vertex to all other reachable vertices.
#
# To compute SSSP for a graph in cuGraph we use:
# **cugraph.sssp(G, source)**
#
# Input
# * __G__: cugraph.Graph object
# * __source__: int, Index of the source vertex
#
# Returns
# * __df__: a cudf.DataFrame object with two columns:
# * df['vertex']: The vertex identifier for the vertex
# * df['distance']: The computed distance from the source vertex to this vertex
#
#
# ## cuGraph Notice
# The current version of cuGraph has some limitations:
#
# * Vertex IDs need to be 32-bit integers.
# * Vertex IDs are expected to be contiguous integers starting from 0.
#
# cuGraph provides the renumber function to mitigate this problem. Input vertex IDs for the renumber function can be either 32-bit or 64-bit integers, can be non-contiguous, and can start from an arbitrary number. The renumber function maps the provided input vertex IDs to 32-bit contiguous integers starting from 0. cuGraph still requires the renumbered vertex IDs to be representable in 32-bit integers. These limitations are being addressed and will be fixed soon.
# ### Test Data
# We will be using the Zachary Karate club dataset
# *<NAME>, An information flow model for conflict and fission in small groups, Journal of
# Anthropological Research 33, 452-473 (1977).*
#
#
# 
#
# This is a small graph which allows for easy visual inspection to validate results.
# __Note__: The Karate dataset starts with vertex ID 1 which the cuGraph analytics assume a zero-based starting ID.
# Import needed libraries
import cugraph
import cudf
from collections import OrderedDict
# ### Read the data and adjust the vertex IDs
# Test file - using the clasic Karate club dataset.
datafile='../data/karate-data.csv'
gdf = cudf.read_csv(datafile, names=["src", "dst"], delimiter='\t', dtype=["int32", "int32"])
# +
# Need to shift the vertex IDs to start with zero rather than one (next version of cuGraph will fix this issue)
gdf["src_0"] = gdf["src"] - 1
gdf["dst_0"] = gdf["dst"] - 1
# The SSSP algorithm requires that there are weights. Just use 1.0 here (equivalent to BFS)
gdf["data"] = 1.0
# -
gdf.head()
# ### Create a Graph and call SSSP
# create a Graph
G = cugraph.Graph()
G.from_cudf_edgelist(gdf, source='src_0', destination='dst_0', edge_attr='data')
# Call cugraph.sssp to get the distances from vertex 0:
df = cugraph.sssp(G, 0)
# Find the farthest vertex from the source using the distances:
# __note__ the vertex ID is shifted back to 1-based so that it can be seen on picture above
bestDist = df['distance'][0]
bestVert = df['vertex'][0]
for i in range(len(df)):
if df['distance'][i] > bestDist:
bestDist = df['distance'][i]
bestVert = df['vertex'][i]
print("Farthest vertex is " + str(bestVert + 1) + " with distance of " + str(bestDist))
# There are a number of vertices with the same distance of 3
# ___
# Copyright (c) 2019, <NAME>.
#
# Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
# ___
|
cugraph/traversal/SSSP.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Линейная регрессия
#
#
# ## Мотивация
#
# Почему мы изучаем линеную регрессию?
# - широко используема
# - легко использовать (не нужно тюнить много параметров)
# - относительно легко интерпретируема
# - является базой для понимания более сложных алгоритмов
#
# ## Библиотеки
#
# Мы познакомимся с двумя библиотеками: [Statsmodels](http://statsmodels.sourceforge.net/) и [scikit-learn](http://scikit-learn.org/stable/). Первая будет полезна вам для эконометрики и интерпретации моделей, вторая это в целом более популярная библиотека для машинного обучения, которая предлагает множество других методов помимо линейной регресии.
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
# ## Мы рассмотрим Advertising Dataset
#
# read data into a DataFrame
data = pd.read_csv('Advertising.csv', index_col=0)
data.head()
# Что такое **признаки / регрессоры / независимые переменные / "фичи" / факторы**?
# - TV: доллары, потраченные на ТВ рекламу для одного продукта на данном рынке (в тысячах долларов)
# - Radio: доллары, потраченные на радио рекламу
# - Newspaper: доллары, потраченные на рекламу в газетах
#
# Что такое **таргет / зависимая переменная**?
# - Sales: продажи одного продукта на данном рынке (в тысячах)
# print the shape of the DataFrame
data.shape
# В нашей выборке мы имеем 200 **наблюдений / "семплов"** (200 рынков в наборе данных)
# visualize the relationship between the features and the response using scatterplots
fig, axs = plt.subplots(1, 3, sharey=True)
data.plot(kind='scatter', x='TV', y='Sales', ax=axs[0], figsize=(16, 8))
data.plot(kind='scatter', x='Radio', y='Sales', ax=axs[1])
data.plot(kind='scatter', x='Newspaper', y='Sales', ax=axs[2])
plt.show()
# ## Вопросы на которые мы обычно хотим знать ответы
#
# Давайте представим, что вы работаете на компанию, которая производит и продает этот товар. Компания может задать вам следующий вопрос: на основании этих данных, как мы должны тратить наши рекламные деньги в будущем?
#
# Этот общий вопрос может привести вас к более конкретным вопросам:
# 1. Существует ли связь между рекламой и продажами?
# 2. Насколько сильны эти отношения?
# 3. Какие типы рекламы способствуют росту продаж?
# 4. Зная расходы на рекламу на конкретном рынке,можем ли мы предсказать продажи?
#
# ## Одномерная линейная регрессия
#
# Простая линейная регрессия - это подход к прогнозированию **количественной переменной** с использованием **одного признака**:
#
# $y = \beta_0 + \beta_1x$
#
# - $y$ это таргет
# - $x$ это признак
# - $\beta_0$ это свободный член
# - $\beta_1$ это коэффициент при признаке x
#
# Вместе $\beta_0$ and $\beta_1$ называются **коэффициентами модели**. Чтобы создать свою модель, вы должны "выучить" значения этих коэффициентов. И как только мы "выучим" эти коэффициенты, мы сможем использовать модель для прогнозирования продаж!
# ## Оценка ("обучение") коэффициентов модели
#
#
# Вообще говоря, коэффициенты оцениваются с использованием **критерия наименьших квадратов**, что означает, что мы находим линию (математически), которая минимизирует **сумму квадратов остатков** (или «сумму квадратов ошибок»):
# <img src="images/08_estimating_coefficients.png">
# Что изображено на схеме?
# - Черные точки - это **наблюдаемые значения** x и y.
# - Синяя линия - это наша **линия полученная при минимизации квадратов ошибок**.
# - Красные линии - это **остатки**, которые представляют собой расстояния между наблюдаемыми значениями и линией наименьших квадратов.
#
# Как коэффициенты модели соотносятся с линией наименьших квадратов?
# - $ \ beta_0 $ - это **точка пересечения** (значение $ y $, когда $ x $ = 0)
# - $ \ beta_1 $ - это **наклон** (изменение $ y $, деленное на изменение $ x $)
#
# Вот графическое изображение этих вычислений:
# <img src="images/08_slope_intercept.png">
# Давайте попробуем использовать **Statsmodels** для оценки коэффициентов модели на наших данных
# +
# this is the standard import if you're using "formula notation" (similar to R)
import statsmodels.formula.api as smf
# create a fitted model in one line
lm = smf.ols(formula='Sales ~ TV', data=data).fit()
# print the coefficients
lm.params
# -
# ## Интерпретация коэффициентов модели
#
# Как мы интерпретируем TV коэффициент ($\beta_1$)?
# - Дополнительные 1000 долларов, потраченные на телевизионную рекламу, увеличивают продажы на 47,537 товаров.
#
# Обратите внимание, что если бы увеличение расходов на телевизионную рекламу было связано с **снижением** продаж, $\beta_1$ был бы **отрицательным**.
# ## Использование модели для прогнозирования
#
# Допустим, появился новый рынок, на котором расходы на телевизионную рекламу составили **$50,000**. Что бы мы спрогнозировали для продаж на этом рынке?
#
# $$y = \beta_0 + \beta_1x$$
# $$y = 7.032594 + 0.047537 \times 50$$
# manually calculate the prediction
7.032594 + 0.047537*50
# Таким образом, мы могли бы предсказать продажи **9409 товаров** на этом рынке.
#
# Конечно, мы также можем использовать Statsmodels для прогнозирования:
# you have to create a DataFrame since the Statsmodels formula interface expects it
X_new = pd.DataFrame({'TV': [50]})
X_new.head()
# use the model to make predictions on a new value
lm.predict(X_new)
# ## Построение линии наименьших квадратов
#
# Давайте сделаем прогнозы для **наименьших и наибольших наблюдаемых значений x**, а затем используем предсказанные значения для построения линии наименьших квадратов:
# create a DataFrame with the minimum and maximum values of TV
X_new = pd.DataFrame({'TV': [data.TV.min(), data.TV.max()]})
X_new.head()
# make predictions for those x values and store them
preds = lm.predict(X_new)
preds
# +
# first, plot the observed data
data.plot(kind='scatter', x='TV', y='Sales')
# then, plot the least squares line
plt.plot(X_new, preds, c='red', linewidth=2)
plt.show()
# -
# ## Насколько хорошо модель соответствует данным?
#
# Одним из способом оценки качества линейной модели является значение **R-квадрат**. R-квадрат-это **это доля объяснённой суммы квадратов в общей**:
#
# 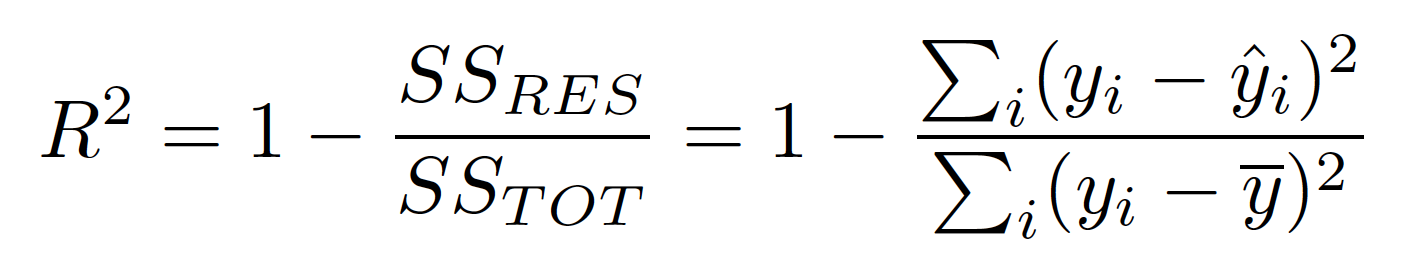
#
# Чем выше R-квадрат, тем лучше, потому что это означает, что большая дисперсия объясняется моделью. Вот пример того, как выглядит R-квадрат":
# <img src="images/08_r_squared.png">
# Вы можете видеть, что **синяя линия** объясняет некоторую дисперсию в данных (R-квадрат=0,54), **зеленая линия** объясняет большую часть дисперсии (R-квадрат=0,64), а **красная линия** еще больше подходит к данным обучения (R-квадрат=0,66). (Красная линия выглядит так, как будто она **слишком сильно подстраивается**?)
#
# Давайте вычислим значение R-квадрата для нашей простой линейной модели:
# print the R-squared value for the model
lm.rsquared
# Является ли это "хорошим" значением R-квадрата? Трудно сказать. Порог для хорошего значения R-квадрата широко зависит от области. Поэтому он наиболее полезен в качестве инструмента для **сравнения различных моделей**.
# ## Множественная линейная регрессия
#
# Простая линейная регрессия может быть легко расширена для включения нескольких объектов. Это называется **множественной линейной регрессией** (вообще, когда говорят о линейной регрессии подразумевают именно ее):
#
# $y = \beta_0 + \beta_1x_1 + ... + \beta_nx_n$
#
# Каждый $x$ представляет собой отдельный признак и имеет свой собственный коэффициент. В этом случае:
#
# $y = \beta_0 + \beta_1 \times TV + \beta_2 \times Radio + \beta_3 \times Newspaper$
#
# Давайте воспользуемся Statsmodels для оценки этих коэффициентов:
# +
# create a fitted model with all three features
lm = smf.ols(formula='Sales ~ TV + Radio + Newspaper', data=data).fit()
# print the coefficients
lm.params
# -
# Как мы интерпретируем эти коэффициенты? Для данной суммы расходов на рекламу на радио и в газетах **увеличение расходов на рекламу на телевидении на 1000 долларов увеличивает продажы на 45,765**.
#
# ## Несколько слов про значимость полученных результатов
#
# Вы должны понимать, что **коэффициенты полученные вами это случайные величины**. В примере выше, например, можно сказать, что **реклама в газетах уменьшает количество проданного товара**, тк коэффициент перед сооствествующим коэффициентом отрицательный. Однако, из-за того, что коэффициент маленький **так могло произойти случайно**. Чтобы оценить это вам понадобяться знания, которые вы получите позже :) Однако, даже сейчас вы можете посмотреть отчет, который предлагает Statsmodels и сделать "примерные" выводы о значимости каждого коэффициента
# print a summary of the fitted model
lm.summary()
# Что можно увидеть по результатам выше?
#
# - Расходы на рекламу на телевидении и радио **положительно связаны** с продажами, в то время как расходы на рекламу в газетах **незначимо отрицательно связаны** с продажами.
# - Эта модель имеет более высокий **R-квадрат** (0,897), чем предыдущая модель, что означает, что эта модель обеспечивает лучшее соответствие данным, чем модель, включающая только TV.
# ## Переобучение (overfitting)
#
# **R-квадрат всегда будет увеличиваться по мере добавления новых признаков в модель**, даже если они не связаны с таргетом. Таким образом, выбор модели с наибольшим R-квадратом не является надежным подходом для выбора наилучшей линейной модели.
#
# 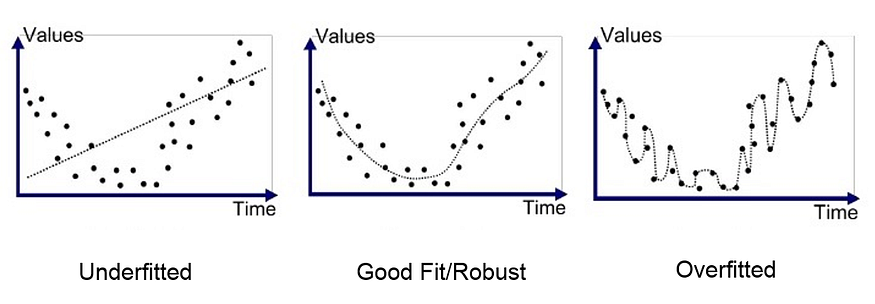
# only include TV and Radio in the model
lm = smf.ols(formula='Sales ~ TV + Radio', data=data).fit()
lm.rsquared
# add Newspaper to the model (which we believe has no association with Sales)
lm = smf.ols(formula='Sales ~ TV + Radio + Newspaper', data=data).fit()
lm.rsquared
# +
data["Random"] = np.random.normal(0, 1, data.shape[0])
lm = smf.ols(formula='Sales ~ TV + Radio + Newspaper + Random', data=data).fit()
lm.rsquared
# -
# Что делать? **Тестировать на независимой выборке**! Этим мы займемся на следующем семинаре, а сейчас scikit-learn!
# ## Линейная регрессия в scikit-learn
#
# +
# create X and y
feature_cols = ['TV', 'Radio', 'Newspaper']
X = data[feature_cols]
y = data.Sales
# follow the usual sklearn pattern: import, instantiate, fit
from sklearn.linear_model import LinearRegression
lm = LinearRegression()
lm.fit(X, y)
# print intercept and coefficients
print(lm.intercept_)
print(lm.coef_)
# -
# pair the feature names with the coefficients
list(zip(feature_cols, lm.coef_))
# predict for a new observation
lm.predict(np.array([[100, 25, 25]]))
# calculate the R-squared
lm.score(X, y)
# Заметьте, что **scikit-learn предоставляет сильно меньше информации** по сравнению с Statsmodels
# ## Обработка категориальных предикторов с двумя категориями
#
# До сих пор все наши предсказатели были числовыми. Что, если один из наших предсказателей был категориальным?
#
# Давайте создадим новый бинарный признак под названием **Size** с двумя возможными значениями: **маленький или большой**:
# +
import numpy as np
# set a seed for reproducibility
np.random.seed(12345)
# create a Series of booleans in which roughly half are True
nums = np.random.rand(len(data))
mask_large = nums > 0.5
# initially set Size to small, then change roughly half to be large
data['Size'] = 'small'
data.loc[mask_large, 'Size'] = 'large'
data.head()
# -
# Для scikit-learn нам нужно представить все данные **в числовом формате**. Если функция имеет только две категории, мы можем просто создать **фиктивную переменную**, которая представляет категории как бинарное значение:
# create a new Series called IsLarge
data['IsLarge'] = data.Size.map({'small':0, 'large':1})
data.head()
# +
# create X and y
feature_cols = ['TV', 'Radio', 'Newspaper', 'IsLarge']
X = data[feature_cols]
y = data.Sales
# instantiate, fit
lm = LinearRegression()
lm.fit(X, y)
# print coefficients
list(zip(feature_cols, lm.coef_))
# -
# Как мы интерпретируем **IsLarge коэффициент**? Для данного объема расходов на рекламу на телевидении/радио/в газетах наличие большого рынка связано со средним **увеличением** продаж на 57,42 (по сравнению с небольшим рынком, который называется **базовым значением**).
#
# Если бы мы изменили кодировку 0/1 и вместо этого создали признак "IsSmall"? Коэффициент будет тем же самым, за исключением того, что он будет отрицательным, а не положительным.
# ## Обработка категориальных предикторов с более чем двумя категориями
#
# Давайте создадим новый признак под названием **Area** и случайным образом назначим наблюдения **rural, suburban, или urban**:
# +
# set a seed for reproducibility
np.random.seed(123456)
# assign roughly one third of observations to each group
nums = np.random.rand(len(data))
mask_suburban = (nums > 0.33) & (nums < 0.66)
mask_urban = nums >= 0.66
data['Area'] = 'rural'
data.loc[mask_suburban, 'Area'] = 'suburban'
data.loc[mask_urban, 'Area'] = 'urban'
data.head()
# -
# Мы должны представлять область численно, но мы не можем просто кодировать ее как 0=Сельская, 1=Пригородная, 2=городская, потому что это означало бы упорядоченное отношение между пригородом и городом (и, таким образом, город каким-то образом "дважды" является пригородной категорией).
#
# Вместо этого мы создаем **еще одну фиктивную переменную**:
# +
# create three dummy variables using get_dummies, then exclude the first dummy column
area_dummies = pd.get_dummies(data.Area, prefix='Area').iloc[:, 1:]
# concatenate the dummy variable columns onto the original DataFrame (axis=0 means rows, axis=1 means columns)
data = pd.concat([data, area_dummies], axis=1)
data.head()
# -
# Вот как мы интерпретируем кодировку:
# - **сельский** кодируется как Area_suburban=0 и Area_urban=0
# - **suburban** кодируется как Area_suburban=1 и Area_urban=0
# - **urban** кодируется как Area_suburban=0 и Area_urban=1
#
# Почему нам нужны только **две фиктивные переменные, а не три?** потому что две дамми переменные захватывают всю информацию об признаке Area и неявно определяют сельскую местность как базовое значение. (В общем случае, если у вас есть категориальный признак с k уровнями, вы создаете фиктивные переменные k-1.)
#
# Если это сбивает с толку, подумайте о том, почему нам нужна только одна фиктивная Переменная для размера (IsLarge), а не две фиктивные переменные (IsSmall и IsLarge).
#
# Давайте включим в модель две новые фиктивные переменные:
# +
# create X and y
feature_cols = ['TV', 'Radio', 'Newspaper', 'IsLarge', 'Area_suburban', 'Area_urban']
X = data[feature_cols]
y = data.Sales
# instantiate, fit
lm = LinearRegression()
lm.fit(X, y)
# print coefficients
list(zip(feature_cols, lm.coef_))
# -
# Как мы интерпретируем коэффициенты?
# - Если все остальные переменные фиксированны, то быть **пригородным** районом, влечет **снижение** продаж на 106,56 (по сравнению с базовым уровнем, который является сельским).
# - Быть **городским** районом связано со средним **увеличением** продаж на 268,13 (по сравнению с сельским районом).
#
# **Заключительное замечание о фиктивном кодировании:** если у вас есть категории, которые можно ранжировать (например, полностью не согласен, не согласен, нейтрален, согласен, полностью согласен), вы можете использовать одну фиктивную переменную и представлять категории численно (например, 1, 2, 3, 4, 5).
# ## Что мы не рассмотрели?
#
# - Допущения линейной регрессии
# - Нелинейные засисимости
# - Обнаружение коллинеарности
# - И многое другое! Этот ноутбук рассказывает только про самые базовые вещи!
#
# ## References
#
# - Этот ноутбук является переводом и адаптацией под наш курс данного ноутбука - https://github.com/justmarkham/DAT4/blob/master/notebooks/08_linear_regression.ipynb
#
#
# - Визуализация от нашего ассистента :)
# https://kawaiiuroboros.github.io/linear-regression/
|
sem10_11_linearRegression/sem10_linearRegression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/dan-a-iancu/airm/blob/master/Project_Portfolio_Management/Project_Portfolio_Management_Template.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="7VtkH-sn3oBT"
# This notebook implements the solution to the **Project Portfolio Management** mini-case. It assumes you are familiar with the case and the model.
# + [markdown] id="vb4A3Z6sy5J_"
# ____
# # Basic Setup
#
# Import useful modules, read the data and store it in data frames, and set up some useful Python lists. You may want to expand this section and make sure you understand how the data is organized, and also read the last part where the Python lists are created, as these may be very useful when you build your model.
# + colab={"base_uri": "https://localhost:8080/"} id="WcFPypqhXbcv" cellView="form" outputId="a99183bf-e182-4790-a772-12d9f57c5624"
#@markdown We first import some useful modules.
# Python ≥3.5 is required
import sys
assert sys.version_info >= (3, 5)
# import numpy
import numpy as np
import urllib.request # for file downloading
# Import pandas for data-frames
import pandas as pd
pd.options.display.max_rows = 15
pd.options.display.float_format = "{:,.2f}".format
from IPython.display import display
# Make sure Matplotlib runs inline, for nice figures
# %matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rc('axes', labelsize=14)
mpl.rc('xtick', labelsize=12)
mpl.rc('ytick', labelsize=12)
import matplotlib.ticker as ticker
# install Gurobi (our linear optimization solver)
# !pip install -i https://pypi.gurobi.com gurobipy
from gurobipy import *
# some modules to create local directories for CBC (to avoid issues with solving multiple models)
import os
def new_local_directory(name):
full_path = os.path.join(".", name)
os.makedirs(full_path, exist_ok=True)
return full_path
# install the latest version of seaborn for nicer graphics
# #!pip install --prefix {sys.prefix} seaborn==0.11.0 &> /dev/null
import seaborn as sns
# Ignore useless some warnings
import warnings
warnings.simplefilter(action="ignore")
print("Completed successfully!")
# + [markdown] id="Aqf-EBXAAgn1"
# ## Load the case data into Pandas data frames
# + [markdown] id="17bUiOG3EYfu"
# We first download an Excel file with all the data from Github.
# + colab={"base_uri": "https://localhost:8080/"} id="oO9JJFpMEX8K" outputId="c7d7db27-ae23-439b-c256-035ea92603ab"
#@markdown Download the entire data as an Excel file from Github
url_Excel = 'https://github.com/dan-a-iancu/airm/blob/master/Project_Portfolio_Management/Project_Data.xlsx?raw=true'
local_file = "Portfolio_Project_Data.xlsx" # name of local file where you want to store the downloaded file
urllib.request.urlretrieve(url_Excel, local_file) # download from website and save it locally
# + [markdown] id="nb7NvPyuyoGh"
# Read in and store the data in suitable dataframes.
# + colab={"base_uri": "https://localhost:8080/", "height": 377} id="EUxSwrdEyJDs" outputId="23ab5bd3-eb3f-4e9f-da28-a74543800fb6"
#@markdown Create a dataframe with the information on projects
# Read in the information about the available projects
projectData = pd.read_excel("Portfolio_Project_Data.xlsx", sheet_name = "Projects", index_col = 0)
# Have a look:
display(projectData)
# + colab={"base_uri": "https://localhost:8080/", "height": 137} id="kXWS_YKrSl05" outputId="33d9ab53-aceb-4401-9665-72c1e901a368"
#@markdown Create a dataframe with the information on available resources (this is useful in **Q5**)
# Read in the information about the available projects
resourceData = pd.read_excel("Portfolio_Project_Data.xlsx", sheet_name = "Resources", index_col = 0)
# Have a look:
display(resourceData)
# + [markdown] id="XRdz0phvd9Z1"
# Also set up any other problem data/parameters, such as the initial capital available.
# + id="Tx8pnOVOd6_u"
initialCapital = 25000
# + [markdown] id="8yhR3xIVyJ13"
# ## Create Python lists based on the data-frames
#
# __NOTE__: Make sure you understand what the __lists__ created here are! These will be very helpful when creating the model.
# + id="oi5sktA8j3PF" colab={"base_uri": "https://localhost:8080/"} outputId="43aeec45-0790-4615-d477-693954f998b7"
#@markdown Some useful lists for building all the models
# the list with project names (A,B, ...)
allProjects = list(projectData.index)
print("This is the list of all the project names:")
print(allProjects)
# the unique locations / continents
allLocations = list(projectData["Location"].unique())
print("\nThis is the list of unique locations:")
print(allLocations)
# + colab={"base_uri": "https://localhost:8080/"} id="FP9z_8hAS25M" outputId="b65aa55c-131f-4bbf-dc50-cc13c672a666"
#@markdown The following lists will be useful in **Q5**
# the list with periods when the projects could be scheduled
allPeriods = list(resourceData.columns)
print("These are periods when the projects could be scheduled:")
print(allPeriods)
# the types of resources needed
allResources = list(resourceData.index)
print("\nThese are the unique resources needed to execute the projects:")
print(allResources)
# + [markdown] id="ZZVNUzIEtYwJ"
# **Q1**
# + id="19TAA2O5tZsi"
|
Project_Portfolio_Management/Project_Portfolio_Management_Template.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
"""
Hand Tracing Module
By: <NAME>
Website: https://github.com/ShubhGurukul
"""
import cv2
import mediapipe as mp
import time
import math
import numpy as np
class handDetector():
def __init__(self, mode=False, maxHands=2, detectionCon=0.5, trackCon=0.5):
self.mode = mode
self.maxHands = maxHands
self.detectionCon = detectionCon
self.trackCon = trackCon
self.mpHands = mp.solutions.hands
self.hands = self.mpHands.Hands(self.mode, self.maxHands,
self.detectionCon, self.trackCon)
self.mpDraw = mp.solutions.drawing_utils
self.tipIds = [4, 8, 12, 16, 20]
def findHands(self, img, draw=True):
imgRGB = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
self.results = self.hands.process(imgRGB)
# print(results.multi_hand_landmarks)
if self.results.multi_hand_landmarks:
for handLms in self.results.multi_hand_landmarks:
if draw:
self.mpDraw.draw_landmarks(img, handLms,
self.mpHands.HAND_CONNECTIONS)
return img
def findPosition(self, img, handNo=0, draw=True):
xList = []
yList = []
bbox = []
self.lmList = []
if self.results.multi_hand_landmarks:
myHand = self.results.multi_hand_landmarks[handNo]
for id, lm in enumerate(myHand.landmark):
# print(id, lm)
h, w, c = img.shape
cx, cy = int(lm.x * w), int(lm.y * h)
xList.append(cx)
yList.append(cy)
# print(id, cx, cy)
self.lmList.append([id, cx, cy])
if draw:
cv2.circle(img, (cx, cy), 5, (255, 0, 255), cv2.FILLED)
xmin, xmax = min(xList), max(xList)
ymin, ymax = min(yList), max(yList)
bbox = xmin, ymin, xmax, ymax
if draw:
cv2.rectangle(img, (xmin - 20, ymin - 20), (xmax + 20, ymax + 20),
(0, 255, 0), 2)
return self.lmList, bbox
def fingersUp(self):
fingers = []
# Thumb
if self.lmList[self.tipIds[0]][1] > self.lmList[self.tipIds[0] - 1][1]:
fingers.append(1)
else:
fingers.append(0)
# Fingers
for id in range(1, 5):
if self.lmList[self.tipIds[id]][2] < self.lmList[self.tipIds[id] - 2][2]:
fingers.append(1)
else:
fingers.append(0)
# totalFingers = fingers.count(1)
return fingers
def findDistance(self, p1, p2, img, draw=True,r=15, t=3):
x1, y1 = self.lmList[p1][1:]
x2, y2 = self.lmList[p2][1:]
cx, cy = (x1 + x2) // 2, (y1 + y2) // 2
if draw:
cv2.line(img, (x1, y1), (x2, y2), (255, 0, 255), t)
cv2.circle(img, (x1, y1), r, (255, 0, 255), cv2.FILLED)
cv2.circle(img, (x2, y2), r, (255, 0, 255), cv2.FILLED)
cv2.circle(img, (cx, cy), r, (0, 0, 255), cv2.FILLED)
length = math.hypot(x2 - x1, y2 - y1)
return length, img, [x1, y1, x2, y2, cx, cy]
def main():
pTime = 0
cTime = 0
cap = cv2.VideoCapture(1)
detector = handDetector()
while True:
success, img = cap.read()
img = detector.findHands(img)
lmList, bbox = detector.findPosition(img)
if len(lmList) != 0:
print(lmList[4])
cTime = time.time()
fps = 1 / (cTime - pTime)
pTime = cTime
cv2.putText(img, str(int(fps)), (10, 70), cv2.FONT_HERSHEY_PLAIN, 3,
(255, 0, 255), 3)
cv2.imshow("Image", img)
cv2.waitKey(1)
if __name__ == "__main__":
main()
# -
# +
import cv2
import mediapipe as mp
import time
cap = cv2.VideoCapture(0)
mpHands = mp.solutions.hands
hands = mpHands.Hands()
mpDraw = mp.solutions.drawing_utils
pTime = 0
cTime = 0
while True:
success, img = cap.read()
imgRGB = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
results = hands.process(imgRGB)
# print(results.multi_hand_landmarks)
if results.multi_hand_landmarks:
for handLms in results.multi_hand_landmarks:
for id, lm in enumerate(handLms.landmark):
# print(id, lm)
h, w, c = img.shape
cx, cy = int(lm.x * w), int(lm.y * h)
print(id, cx, cy)
# if id == 4:
cv2.circle(img, (cx, cy), 15, (255, 0, 255), cv2.FILLED)
mpDraw.draw_landmarks(img, handLms, mpHands.HAND_CONNECTIONS)
cTime = time.time()
fps = 1 / (cTime - pTime)
pTime = cTime
cv2.putText(img, str(int(fps)), (10, 70), cv2.FONT_HERSHEY_PLAIN, 3,
(255, 0, 255), 3)
cv2.imshow("Image", img)
cv2.waitKey(1)
# -
# # %%writefile HandTrackingModule.py
"""
Hand Tracing Module
By: <NAME>
Website: https://github.com/ShubhGurukul
"""
import cv2
import mediapipe as mp
import time
class handDetector():
def __init__(self, mode=False, maxHands=2, detectionCon=0.5, trackCon=0.5):
self.mode = mode
self.maxHands = maxHands
self.detectionCon = detectionCon
self.trackCon = trackCon
self.mpHands = mp.solutions.hands
self.hands = self.mpHands.Hands(self.mode, self.maxHands,
self.detectionCon, self.trackCon)
self.mpDraw = mp.solutions.drawing_utils
def findHands(self, img, draw=True):
imgRGB = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
self.results = self.hands.process(imgRGB)
# print(results.multi_hand_landmarks)
if self.results.multi_hand_landmarks:
for handLms in self.results.multi_hand_landmarks:
if draw:
self.mpDraw.draw_landmarks(img, handLms,
self.mpHands.HAND_CONNECTIONS)
return img
def findPosition(self, img, handNo=0, draw=True):
lmList = []
if self.results.multi_hand_landmarks:
myHand = self.results.multi_hand_landmarks[handNo]
for id, lm in enumerate(myHand.landmark):
# print(id, lm)
h, w, c = img.shape
cx, cy = int(lm.x * w), int(lm.y * h)
# print(id, cx, cy)
lmList.append([id, cx, cy])
if draw:
cv2.circle(img, (cx, cy), 15, (255, 0, 255), cv2.FILLED)
return lmList
def main():
pTime = 0
cTime = 0
cap = cv2.VideoCapture(1)
detector = handDetector()
while True:
success, img = cap.read()
img = detector.findHands(img)
lmList = detector.findPosition(img)
if len(lmList) != 0:
print(lmList[4])
cTime = time.time()
fps = 1 / (cTime - pTime)
pTime = cTime
cv2.putText(img, str(int(fps)), (10, 70), cv2.FONT_HERSHEY_PLAIN, 3,
(255, 0, 255), 3)
cv2.imshow("Image", img)
cv2.waitKey(1)
if __name__ == "__main__":
main()
# !pip install pycaw
import cv2
import time
import numpy as np
import HandTrackingModule as htm
import math
from ctypes import cast, POINTER
from comtypes import CLSCTX_ALL
from pycaw.pycaw import AudioUtilities, IAudioEndpointVolume
################################
wCam, hCam = 640, 480
################################
cap = cv2.VideoCapture(1)
cap.set(3, wCam)
cap.set(4, hCam)
pTime = 0
detector = htm.handDetector(detectionCon=0.7)
devices = AudioUtilities.GetSpeakers()
interface = devices.Activate(
IAudioEndpointVolume._iid_, CLSCTX_ALL, None)
volume = cast(interface, POINTER(IAudioEndpointVolume))
# volume.GetMute()
# volume.GetMasterVolumeLevel()
volRange = volume.GetVolumeRange()
minVol = volRange[0]
maxVol = volRange[1]
vol = 0
volBar = 400
volPer = 0
while True:
success, img = cap.read()
img = detector.findHands(img)
lmList = detector.findPosition(img, draw=False)
if len(lmList) != 0:
# print(lmList[4], lmList[8])
x1, y1 = lmList[4][1], lmList[4][2]
x2, y2 = lmList[8][1], lmList[8][2]
cx, cy = (x1 + x2) // 2, (y1 + y2) // 2
cv2.circle(img, (x1, y1), 15, (255, 0, 255), cv2.FILLED)
cv2.circle(img, (x2, y2), 15, (255, 0, 255), cv2.FILLED)
cv2.line(img, (x1, y1), (x2, y2), (255, 0, 255), 3)
cv2.circle(img, (cx, cy), 15, (255, 0, 255), cv2.FILLED)
length = math.hypot(x2 - x1, y2 - y1)
# print(length)
# Hand range 50 - 300
# Volume Range -65 - 0
vol = np.interp(length, [50, 300], [minVol, maxVol])
volBar = np.interp(length, [50, 300], [400, 150])
volPer = np.interp(length, [50, 300], [0, 100])
print(int(length), vol)
volume.SetMasterVolumeLevel(vol, None)
if length <50:
cv2.circle(img, (cx, cy), 15, (0, 255, 0), cv2.FILLED)
cv2.rectangle(img, (50, 150), (85, 400), (255, 0, 0), 3)
cv2.rectangle(img, (50, int(volBar)), (85, 400), (255, 0, 0), cv2.FILLED)
cv2.putText(img, f'{int(volPer)} %', (40, 450), cv2.FONT_HERSHEY_COMPLEX,
1, (255, 0, 0), 3)
cTime = time.time()
fps = 1 / (cTime - pTime)
pTime = cTime
cv2.putText(img, f'FPS: {int(fps)}', (40, 50), cv2.FONT_HERSHEY_COMPLEX,
1, (255, 0, 0), 3)
cv2.imshow("Img", img)
cv2.imshow("Video",img) # and showing on screen
if cv2.waitKey(1) & 0xff == ord('q'):
break
# # Finger Counter
'''
Finger Counter Module
By: <NAME>
Website: https://github.com/ShubhGurukul
'''
import cv2
import time
import os
import HandTrackingModule as htm
wCam, hCam = 640, 480
cap = cv2.VideoCapture(1)
cap.set(3, wCam)
cap.set(4, hCam)
folderPath = "FingerImages"
myList = os.listdir(folderPath)
print(myList)
overlayList = []
for imPath in myList:
image = cv2.imread(f'{folderPath}/{imPath}')
# print(f'{folderPath}/{imPath}')
overlayList.append(image)
print(len(overlayList))
pTime = 0
detector = htm.handDetector(detectionCon=0.75)
tipIds = [4, 8, 12, 16, 20]
while True:
success, img = cap.read()
img = detector.findHands(img)
lmList = detector.findPosition(img, draw=False)
# print(lmList)
if len(lmList) != 0:
fingers = []
# Thumb
if lmList[tipIds[0]][1]< lmList[tipIds[0] - 1][1]:
fingers.append(1)
else:
fingers.append(0)
# 4 Fingers
for id in range(1, 5):
if lmList[tipIds[id]][2] < lmList[tipIds[id] - 2][2]:
fingers.append(1)
else:
fingers.append(0)
# print(fingers)
totalFingers = fingers.count(1)
print(totalFingers)
h, w, c = overlayList[totalFingers - 1].shape
img[0:h, 0:w] = overlayList[totalFingers - 1]
cv2.rectangle(img, (20, 225), (170, 425), (0, 255, 0), cv2.FILLED)
cv2.putText(img, str(totalFingers), (45, 375), cv2.FONT_HERSHEY_PLAIN,
10, (255, 0, 0), 25)
cTime = time.time()
fps = 1 / (cTime - pTime)
pTime = cTime
cv2.putText(img, f'FPS: {int(fps)}', (400, 70), cv2.FONT_HERSHEY_PLAIN,
3, (255, 0, 0), 3)
cv2.imshow("Image", img)
# cv2.waitKey(1)
if cv2.waitKey(1) & 0xff == ord('q'):
break
# # AI Personal Trainer
# ## PoseModule
# +
'''
Pose Module
By: <NAME>
Website: https://github.com/ShubhGurukul
'''
# # %%writefile PoseModule.py
import cv2
import mediapipe as mp
import time
import math
class poseDetector():
def __init__(self, mode=False, upBody=False, smooth=True,
detectionCon=0.5, trackCon=0.5):
self.mode = mode
self.upBody = upBody
self.smooth = smooth
self.detectionCon = detectionCon
self.trackCon = trackCon
self.mpDraw = mp.solutions.drawing_utils
self.mpPose = mp.solutions.pose
self.pose = self.mpPose.Pose(self.mode, self.upBody, self.smooth,
self.detectionCon, self.trackCon)
def findPose(self, img, draw=True):
imgRGB = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
self.results = self.pose.process(imgRGB)
if self.results.pose_landmarks:
if draw:
self.mpDraw.draw_landmarks(img, self.results.pose_landmarks,
self.mpPose.POSE_CONNECTIONS)
return img
def findPosition(self, img, draw=True):
self.lmList = []
if self.results.pose_landmarks:
for id, lm in enumerate(self.results.pose_landmarks.landmark):
h, w, c = img.shape
# print(id, lm)
cx, cy = int(lm.x * w), int(lm.y * h)
self.lmList.append([id, cx, cy])
if draw:
cv2.circle(img, (cx, cy), 5, (255, 0, 0), cv2.FILLED)
return self.lmList
def findAngle(self, img, p1, p2, p3, draw=True):
# Get the landmarks
x1, y1 = self.lmList[p1][1:]
x2, y2 = self.lmList[p2][1:]
x3, y3 = self.lmList[p3][1:]
# Calculate the Angle
angle = math.degrees(math.atan2(y3 - y2, x3 - x2) -
math.atan2(y1 - y2, x1 - x2))
if angle &lt; 0:
angle += 360
# print(angle)
# Draw
if draw:
cv2.line(img, (x1, y1), (x2, y2), (255, 255, 255), 3)
cv2.line(img, (x3, y3), (x2, y2), (255, 255, 255), 3)
cv2.circle(img, (x1, y1), 10, (0, 0, 255), cv2.FILLED)
cv2.circle(img, (x1, y1), 15, (0, 0, 255), 2)
cv2.circle(img, (x2, y2), 10, (0, 0, 255), cv2.FILLED)
cv2.circle(img, (x2, y2), 15, (0, 0, 255), 2)
cv2.circle(img, (x3, y3), 10, (0, 0, 255), cv2.FILLED)
cv2.circle(img, (x3, y3), 15, (0, 0, 255), 2)
cv2.putText(img, str(int(angle)), (x2 - 50, y2 + 50),
cv2.FONT_HERSHEY_PLAIN, 2, (0, 0, 255), 2)
return angle
def main():
cap = cv2.VideoCapture('PoseVideos/1.mp4')
pTime = 0
detector = poseDetector()
while True:
success, img = cap.read()
img = detector.findPose(img)
lmList = detector.findPosition(img, draw=False)
if len(lmList) != 0:
print(lmList[14])
cv2.circle(img, (lmList[14][1], lmList[14][2]), 15, (0, 0, 255), cv2.FILLED)
cTime = time.time()
fps = 1 / (cTime - pTime)
pTime = cTime
cv2.putText(img, str(int(fps)), (70, 50), cv2.FONT_HERSHEY_PLAIN, 3,
(255, 0, 0), 3)
cv2.imshow("Image", img)
cv2.waitKey(1)
if __name__ == "__main__":
main()
# -
# ## AI Trainer
import cv2
import numpy as np
import time
import PoseModule as pm
# cap = cv2.VideoCapture("AiTrainer/curls.mp4")
detector = pm.poseDetector()
count = 0
dir = 0
pTime = 0
while True:
success, img = cap.read()
img = cv2.resize(img, (1280, 720))
# img = cv2.imread("AiTrainer/test.jpg")
img = detector.findPose(img, False)
lmList = detector.findPosition(img, False)
# print(lmList)
if len(lmList) != 0:
# Right Arm
angle = detector.findAngle(img, 12, 14, 16)
# # Left Arm
#angle = detector.findAngle(img, 11, 13, 15,False)
per = np.interp(angle, (210, 310), (0, 100))
bar = np.interp(angle, (220, 310), (650, 100))
# print(angle, per)
# Check for the dumbbell curls
color = (255, 0, 255)
if per == 100:
color = (0, 255, 0)
if dir == 0:
count += 0.5
dir = 1
if per == 0:
color = (0, 255, 0)
if dir == 1:
count += 0.5
dir = 0
print(count)
# Draw Bar
cv2.rectangle(img, (1100, 100), (1175, 650), color, 3)
cv2.rectangle(img, (1100, int(bar)), (1175, 650), color, cv2.FILLED)
cv2.putText(img, f'{int(per)} %', (1100, 75), cv2.FONT_HERSHEY_PLAIN, 4,
color, 4)
# Draw Curl Count
cv2.rectangle(img, (0, 450), (250, 720), (0, 255, 0), cv2.FILLED)
cv2.putText(img, str(int(count)), (45, 670), cv2.FONT_HERSHEY_PLAIN, 15,
(255, 0, 0), 25)
cTime = time.time()
fps = 1 / (cTime - pTime)
pTime = cTime
cv2.putText(img, str(int(fps)), (50, 100), cv2.FONT_HERSHEY_PLAIN, 5,
(255, 0, 0), 5)
cv2.imshow("Image", img)
cv2.waitKey(1)
# # AI Virtual Mouse Project(Not Completed)
# +
# # !pip install autopy
# -
import cv2
import numpy as np
import HandTrackingModule as htm
import time
import autopy
# +
wCam, hCam = 640, 480
frameR = 100 # Frame Reduction
smoothening = 7
#########################
pTime = 0
plocX, plocY = 0, 0
clocX, clocY = 0, 0
cap = cv2.VideoCapture(1)
cap.set(3, wCam)
cap.set(4, hCam)
detector = htm.handDetector(maxHands=1)
wScr, hScr = autopy.screen.size()
# print(wScr, hScr)
while True:
# 1. Find hand Landmarks
success, img = cap.read()
img = detector.findHands(img)
lmList, bbox = detector.findPosition(img)
# 2. Get the tip of the index and middle fingers
if len(lmList) != 0:
x1, y1 = lmList[8][1:]
x2, y2 = lmList[12][1:]
# print(x1, y1, x2, y2)
# 3. Check which fingers are up
fingers = detector.fingersUp()
# print(fingers)
cv2.rectangle(img, (frameR, frameR), (wCam-frameR, hCam-frameR),
(255, 0, 255), 2)
# 4. Only Index Finger : Moving Mode
if fingers[1] == 1 and fingers[2] == 0:
# 5. Convert Coordinates
x3 = np.interp(x1, (frameR, wCam - frameR), (0, wScr))
y3 = np.interp(y1, (frameR, hCam - frameR), (0, hScr))
# 6. Smoothen Values
clocX = plocX + (x3 - plocX) / smoothening
clocY = plocY + (y3 - plocY) / smoothening
# 7. Move Mouse
autopy.mouse.move(wScr - clocX, clocY)
cv2.circle(img, (x1, y1), 15, (255, 0, 255), cv2.FILLED)
plocX, plocY = clocX, clocY
# 8. Both Index and middle fingers are up : Clicking Mode
if fingers[1] == 1 and fingers[2] == 1:
# 9. Find distance between fingers
length, img, lineInfo = detector.findDistance(8, 12, img)
print(length)
# 10. Click mouse if distance short
if length < 40:
cv2.circle(img, (lineInfo[4], lineInfo[5]),
15, (0, 255, 0), cv2.FILLED)
autopy.mouse.click()
# 11. Frame Rate
cTime = time.time()
fps = 1 / (cTime - pTime)
pTime = cTime
cv2.putText(img, str(int(fps)), (20, 50), cv2.FONT_HERSHEY_PLAIN, 3,
(255, 0, 0), 3)
# 12. Display
cv2.imshow('Image', img)
if cv2.waitKey(1) & 0xff == ord('q'):
break
# -
|
All_Is_Done.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="_f-Sm2rSFHzi" colab_type="code" outputId="b233428f-d800-4020-9a87-6e84fbe2a987" executionInfo={"status": "ok", "timestamp": 1583402251101, "user_tz": -60, "elapsed": 15539, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiyVjJipparYJ7zM1JucsAJQgPgEHY3Bh_x9pwjgQ=s64", "userId": "14237109751796858774"}} colab={"base_uri": "https://localhost:8080/", "height": 493}
# !pip install --upgrade tables
# !pip install eli5
# !pip install xgboost
# + id="Nvc9A2v3H_CV" colab_type="code" outputId="12fc31e2-aa9f-40c9-fc2b-dd7833300695" executionInfo={"status": "ok", "timestamp": 1583402222020, "user_tz": -60, "elapsed": 26412, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiyVjJipparYJ7zM1JucsAJQgPgEHY3Bh_x9pwjgQ=s64", "userId": "14237109751796858774"}} colab={"base_uri": "https://localhost:8080/", "height": 122}
from google.colab import drive
drive.mount('/content/drive')
# + id="UfrsZF7cITSH" colab_type="code" colab={}
import pandas as pd
import numpy as np
# + id="h04TZz5kIgxa" colab_type="code" colab={}
from sklearn.dummy import DummyRegressor
from sklearn.tree import DecisionTreeRegressor
from sklearn.ensemble import RandomForestRegressor
# + id="VH0ioG0WJEJ3" colab_type="code" colab={}
import xgboost as xgb
# + id="93pPWXyrJHq_" colab_type="code" colab={}
from sklearn.metrics import mean_absolute_error as mae
from sklearn.model_selection import cross_val_score, KFold
# + id="cYF70JQNJcQd" colab_type="code" colab={}
import eli5
# + id="mBhy0zj0JgXV" colab_type="code" colab={}
from eli5.sklearn import PermutationImportance
# + id="QlSFb96LJl_l" colab_type="code" outputId="20165b6c-8ca4-43cf-f89b-dc0d4163e819" executionInfo={"status": "ok", "timestamp": 1583402758850, "user_tz": -60, "elapsed": 638, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiyVjJipparYJ7zM1JucsAJQgPgEHY3Bh_x9pwjgQ=s64", "userId": "14237109751796858774"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# %cd /content/drive/My\ Drive/Colab\ Notebooks/matrix/matrix_two/dw_matrix_car
# + id="bSi7buohKK9j" colab_type="code" outputId="36f19434-352a-4371-f768-a84edee11845" executionInfo={"status": "ok", "timestamp": 1583402811771, "user_tz": -60, "elapsed": 4532, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiyVjJipparYJ7zM1JucsAJQgPgEHY3Bh_x9pwjgQ=s64", "userId": "14237109751796858774"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
df = pd.read_hdf("data/car.h5")
df.shape
# + id="XgiPxgepKW8y" colab_type="code" colab={}
# + [markdown] id="TCsSFRAqKkmg" colab_type="text"
# ## Features Engineering
# + id="peJIRpWhKpFx" colab_type="code" outputId="c856f2cc-42d9-42cc-aaf9-8ab357a902ed" executionInfo={"status": "ok", "timestamp": 1583403233498, "user_tz": -60, "elapsed": 2379, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiyVjJipparYJ7zM1JucsAJQgPgEHY3Bh_x9pwjgQ=s64", "userId": "14237109751796858774"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
SUFFIX_CAT = '_cat'
for feat in df.columns:
# skip list columns
if isinstance(df[feat][0], list): continue
factorized_values = df[feat].factorize()[0]
if SUFFIX_CAT in feat: # guard against multiply restarts of this notebook
df[feat] = factorized_values
else:
df[feat + SUFFIX_CAT] = factorized_values
#cat_feats = [x for x in df.columns if SUFFIX_CAT in x and 'price' not in x]
cat_feats = [x for x in df.columns if SUFFIX_CAT in x]
cat_feats = [x for x in cat_feats if 'price' not in x]
len(cat_feats)
# + id="N_Kk2KNPLA5x" colab_type="code" colab={}
def run_model(model, feats):
X = df[ feats].values
y = df.price_value.values
scores = cross_val_score(model, X, y, cv = 3, scoring='neg_mean_absolute_error') # runs fit and cross-validation
return np.mean(scores), np.std(scores)
# + [markdown] id="xwgDKj5BNQFK" colab_type="text"
# ## Decision Tree
# + id="PmzteAEYNQyV" colab_type="code" outputId="58be0542-8650-4f35-a3d8-4e829d9b96bf" executionInfo={"status": "ok", "timestamp": 1583411708448, "user_tz": -60, "elapsed": 5226, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiyVjJipparYJ7zM1JucsAJQgPgEHY3Bh_x9pwjgQ=s64", "userId": "14237109751796858774"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
model = DecisionTreeRegressor(max_depth=5,random_state=0) # only one tree
run_model(model, cat_feats)
# + id="P1gedHBpNVBG" colab_type="code" colab={}
# + [markdown] id="are1Voc3NVi4" colab_type="text"
# # Random Forrest
# + id="Q4WJsBu3NXpu" colab_type="code" outputId="3dfcf8f7-917d-4e47-be8f-8fc2f6b87527" executionInfo={"status": "ok", "timestamp": 1583404023176, "user_tz": -60, "elapsed": 129700, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiyVjJipparYJ7zM1JucsAJQgPgEHY3Bh_x9pwjgQ=s64", "userId": "14237109751796858774"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
m2 = RandomForestRegressor(max_depth=5, n_estimators=50, random_state=0) # 50 trees
run_model(m2, cat_feats)
# + [markdown] id="kZqjKKuTOFcw" colab_type="text"
# ## XGBoost
# + id="fDUYrSiDOG93" colab_type="code" outputId="50f6df2d-bc82-4a50-9692-128332d36ce6" executionInfo={"status": "ok", "timestamp": 1583404173072, "user_tz": -60, "elapsed": 61006, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiyVjJipparYJ7zM1JucsAJQgPgEHY3Bh_x9pwjgQ=s64", "userId": "14237109751796858774"}} colab={"base_uri": "https://localhost:8080/", "height": 85}
xgb_params ={'max_depth':5, 'n_estimators':50, 'learning_rate':0.1, 'seed':0}
m3 = xgb.XGBRegressor(**xgb_params)
run_model(m3, cat_feats)
# + id="USjYC__PPGPh" colab_type="code" outputId="4a34993b-ac18-4862-9d22-554aad9dcdbb" executionInfo={"status": "ok", "timestamp": 1583404374106, "user_tz": -60, "elapsed": 30995, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiyVjJipparYJ7zM1JucsAJQgPgEHY3Bh_x9pwjgQ=s64", "userId": "14237109751796858774"}} colab={"base_uri": "https://localhost:8080/", "height": 153}
m4 = xgb.XGBRegressor(**xgb_params)
m4.fit(X,y)
# + id="u6ftyOchQMII" colab_type="code" outputId="c4b24281-66e4-4ccf-db99-b9d05b0f4f2c" executionInfo={"status": "ok", "timestamp": 1583404734872, "user_tz": -60, "elapsed": 333940, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiyVjJipparYJ7zM1JucsAJQgPgEHY3Bh_x9pwjgQ=s64", "userId": "14237109751796858774"}} colab={"base_uri": "https://localhost:8080/", "height": 391}
imp = PermutationImportance(m4, random_state=0).fit(X,y)
eli5.show_weights(imp, feature_names=cat_feats)
# + [markdown] id="aNir9UMlhsVk" colab_type="text"
# ## Simplify model - use only most influential features
# + id="moGPJtXlQcB3" colab_type="code" colab={}
feats = ['param_stan_cat',
'param_rok-produkcji_cat',
'param_napęd_cat',
'param_faktura-vat_cat',
'param_moc_cat',
'param_skrzynia-biegów_cat',
'param_marka-pojazdu_cat',
'feature_kamera-cofania_cat',
'param_typ_cat',
'param_pojemność-skokowa_cat',
'seller_name_cat',
'param_wersja_cat',
'feature_wspomaganie-kierownicy_cat',
'param_model-pojazdu_cat',
'feature_system-start-stop_cat',
'param_kod-silnika_cat',
'feature_asystent-pasa-ruchu_cat',
'feature_łopatki-zmiany-biegów_cat',
'feature_światła-led_cat',
'feature_czujniki-parkowania-przednie_cat']
# + id="i5B6UxbXgCGo" colab_type="code" outputId="dbbc7249-e852-4d8d-ed8c-c060c1ba5f0e" executionInfo={"status": "ok", "timestamp": 1583408558159, "user_tz": -60, "elapsed": 681, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiyVjJipparYJ7zM1JucsAJQgPgEHY3Bh_x9pwjgQ=s64", "userId": "14237109751796858774"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
len(cat_feats),len(feats)
# + id="ETqkX-6zgS01" colab_type="code" outputId="0dd3068d-4da9-4328-d1c8-b8de2a4e8942" executionInfo={"status": "ok", "timestamp": 1583408673953, "user_tz": -60, "elapsed": 14378, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiyVjJipparYJ7zM1JucsAJQgPgEHY3Bh_x9pwjgQ=s64", "userId": "14237109751796858774"}} colab={"base_uri": "https://localhost:8080/", "height": 85}
run_model(xgb.XGBRegressor(**xgb_params), feats)
# + [markdown] id="LADtmjkWiQnb" colab_type="text"
# ## Recover numerical information lost during categorization
# + id="I_R8NOrbgrvs" colab_type="code" outputId="254287f6-52e4-452d-a474-fe658ad70211" executionInfo={"status": "ok", "timestamp": 1583409532930, "user_tz": -60, "elapsed": 730, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiyVjJipparYJ7zM1JucsAJQgPgEHY3Bh_x9pwjgQ=s64", "userId": "14237109751796858774"}} colab={"base_uri": "https://localhost:8080/", "height": 187}
df['param_rok-produkcji'].unique()
# + id="hFEA_tcNi9n0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 136} outputId="74ad1586-6bea-4503-e60a-a07afdd382eb" executionInfo={"status": "ok", "timestamp": 1583409877972, "user_tz": -60, "elapsed": 643, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiyVjJipparYJ7zM1JucsAJQgPgEHY3Bh_x9pwjgQ=s64", "userId": "14237109751796858774"}}
df['param_rok-produkcji'].map(lambda r: -1 if r is None else int(r)).unique()
# + id="JF0JsV4akX2n" colab_type="code" colab={}
df['param_rok-produkcji_NUM'] = df['param_rok-produkcji'].map(lambda r: -1 if r is None else int(r))
# + id="EJ4GlKn1lpL6" colab_type="code" colab={}
feats2 = ['param_stan_cat',
'param_rok-produkcji_NUM',
'param_napęd_cat',
'param_faktura-vat_cat',
'param_moc_cat',
'param_skrzynia-biegów_cat',
'param_marka-pojazdu_cat',
'feature_kamera-cofania_cat',
'param_typ_cat',
'param_pojemność-skokowa_cat',
'seller_name_cat',
'param_wersja_cat',
'feature_wspomaganie-kierownicy_cat',
'param_model-pojazdu_cat',
'feature_system-start-stop_cat',
'param_kod-silnika_cat',
'feature_asystent-pasa-ruchu_cat',
'feature_łopatki-zmiany-biegów_cat',
'feature_światła-led_cat',
'feature_czujniki-parkowania-przednie_cat']
# + id="ELce0Jw7lwjq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="9ad2facc-7789-4217-cc77-d9426e037733" executionInfo={"status": "ok", "timestamp": 1583410023831, "user_tz": -60, "elapsed": 13883, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiyVjJipparYJ7zM1JucsAJQgPgEHY3Bh_x9pwjgQ=s64", "userId": "14237109751796858774"}}
run_model(xgb.XGBRegressor(**xgb_params), feats2)
# + id="9_dMrQ_Zl1bk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="82e26739-31e8-4b18-c2bb-b0548668b8e2" executionInfo={"status": "ok", "timestamp": 1583410116944, "user_tz": -60, "elapsed": 642, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiyVjJipparYJ7zM1JucsAJQgPgEHY3Bh_x9pwjgQ=s64", "userId": "14237109751796858774"}}
df['param_moc'].value_counts()
# + id="LJ2kvd98mGiE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 680} outputId="4927da42-369f-4d13-d8e6-8b7d4d803b8a" executionInfo={"status": "ok", "timestamp": 1583410396397, "user_tz": -60, "elapsed": 745, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiyVjJipparYJ7zM1JucsAJQgPgEHY3Bh_x9pwjgQ=s64", "userId": "14237109751796858774"}}
df['param_moc'].map(lambda x: -1 if x is None else int(x.split(' ')[0])).unique()
# + id="grMVuiiwnvlG" colab_type="code" colab={}
df['param_moc_NUM'] = df['param_moc'].map(lambda x: -1 if x is None else int(x.split(' ')[0]))
# + id="c8VJ-LdOnD6z" colab_type="code" colab={}
feats3 = ['param_stan_cat',
'param_rok-produkcji_NUM',
'param_napęd_cat',
'param_faktura-vat_cat',
'param_moc_NUM',
'param_skrzynia-biegów_cat',
'param_marka-pojazdu_cat',
'feature_kamera-cofania_cat',
'param_typ_cat',
'param_pojemność-skokowa_cat',
'seller_name_cat',
'param_wersja_cat',
'feature_wspomaganie-kierownicy_cat',
'param_model-pojazdu_cat',
'feature_system-start-stop_cat',
'param_kod-silnika_cat',
'feature_asystent-pasa-ruchu_cat',
'feature_łopatki-zmiany-biegów_cat',
'feature_światła-led_cat',
'feature_czujniki-parkowania-przednie_cat']
# + id="snvrVhvjn5Us" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="cc66ba5f-5570-4948-bee5-34d81f904298" executionInfo={"status": "ok", "timestamp": 1583410578343, "user_tz": -60, "elapsed": 13775, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiyVjJipparYJ7zM1JucsAJQgPgEHY3Bh_x9pwjgQ=s64", "userId": "14237109751796858774"}}
run_model(xgb.XGBRegressor(**xgb_params), feats3)
# + id="_RPDyU1Sn6f9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="303f1796-e1b0-468b-ebd0-c6433ea5ec9b" executionInfo={"status": "ok", "timestamp": 1583410680348, "user_tz": -60, "elapsed": 712, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiyVjJipparYJ7zM1JucsAJQgPgEHY3Bh_x9pwjgQ=s64", "userId": "14237109751796858774"}}
df['param_pojemność-skokowa'].unique()
# + id="0T4YVUe8oUPc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="987d6093-b253-4cbe-82e6-7e83f04d62d4" executionInfo={"status": "ok", "timestamp": 1583410884000, "user_tz": -60, "elapsed": 666, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiyVjJipparYJ7zM1JucsAJQgPgEHY3Bh_x9pwjgQ=s64", "userId": "14237109751796858774"}}
df['param_pojemność-skokowa'].map(lambda x: -1 if x is None else int(x.split('cm')[0].replace(' ',''))).unique()
# + id="3d5JRydnoyCO" colab_type="code" colab={}
df['param_pojemność-skokowa_NUM'] = df['param_pojemność-skokowa'].map(lambda x: -1 if x is None else int(x.split('cm')[0].replace(' ','')))
# + id="iEwKCjH9plIk" colab_type="code" colab={}
feats4 = ['param_stan_cat',
'param_rok-produkcji_NUM',
'param_napęd_cat',
'param_faktura-vat_cat',
'param_moc_NUM',
'param_skrzynia-biegów_cat',
'param_marka-pojazdu_cat',
'feature_kamera-cofania_cat',
'param_typ_cat',
'param_pojemność-skokowa_NUM',
'seller_name_cat',
'param_wersja_cat',
'feature_wspomaganie-kierownicy_cat',
'param_model-pojazdu_cat',
'feature_system-start-stop_cat',
'param_kod-silnika_cat',
'feature_asystent-pasa-ruchu_cat',
'feature_łopatki-zmiany-biegów_cat',
'feature_światła-led_cat',
'feature_czujniki-parkowania-przednie_cat']
# + id="xGFnaNezpq1T" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="0b10cc04-2031-4236-f805-f46714edb79c" executionInfo={"status": "ok", "timestamp": 1583412177663, "user_tz": -60, "elapsed": 14027, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiyVjJipparYJ7zM1JucsAJQgPgEHY3Bh_x9pwjgQ=s64", "userId": "14237109751796858774"}}
run_model(xgb.XGBRegressor(**xgb_params), feats4)
# + id="p8ILzbtopt5c" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d8e503da-5f81-4812-fcfb-86f06f22260a" executionInfo={"status": "ok", "timestamp": 1583412212473, "user_tz": -60, "elapsed": 13910, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiyVjJipparYJ7zM1JucsAJQgPgEHY3Bh_x9pwjgQ=s64", "userId": "14237109751796858774"}}
run_model(xgb.XGBRegressor(**xgb_params, objective="reg:squarederror"), feats4)
# + id="CT23SQKiuLv_" colab_type="code" colab={}
|
day4.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# The `show()` method of the `Particelset` class is capable of plotting the particle locations and velocity fields in scalar and vector form. In this notebook, we demonstrate these capabilities using the GlobCurrent dataset. We begin by importing the relevant modules.
# %matplotlib inline
from parcels import FieldSet, ParticleSet, JITParticle, AdvectionRK4
from datetime import timedelta, datetime
# We then instatiate a `FieldSet` with the velocity field data from GlobCurrent dataset.
filenames = {'U': "GlobCurrent_example_data/20*.nc",
'V': "GlobCurrent_example_data/20*.nc"}
variables = {'U': 'eastward_eulerian_current_velocity',
'V': 'northward_eulerian_current_velocity'}
dimensions = {'lat': 'lat',
'lon': 'lon',
'time': 'time'}
fieldset = FieldSet.from_netcdf(filenames, variables, dimensions)
# Next, we instantiate a `ParticeSet` composed of `JITParticles`:
pset = ParticleSet.from_line(fieldset=fieldset, size=5, pclass=JITParticle,
start=(31, -31), finish=(32, -31), time=datetime(2002, 1, 1))
# Given this `ParticleSet`, we can now explore the different features of the `show()` method. To start, let's simply call `show()` with no arguments.
pset.show()
# Then, let's advect the particles starting on January 1, 2002 for a week.
pset.execute(AdvectionRK4, runtime=timedelta(days=7), dt=timedelta(minutes=5))
# If we call `show()` again, we will see that the particles have been advected:
pset.show()
# To plot without the continents on the same plot, add `land=False`.
pset.show(land=False)
# To set the domain of the plot, we specify the domain argument. The format `domain` expects is `[max lat, min lat, max lon, min lon]`. Note that the plotted domain is found by interpolating the user-specified domain onto the velocity grid. For instance,
pset.show(domain=[-31, -35, 33, 26])
# We can also easily display a scalar contour plot of a single component of the velocity vector field. This is done by setting the `field` argument equal to the desired scalar velocity field.
pset.show(field=fieldset.U)
# To plot the scalar U velocity field at a different date and time, we set the argument `show_time` equal to a `datetime` or `timedelta` object or simply the number of seconds since the time origin. For instance, let's view the U field on January, 10, 2002 at 2 PM.
pset.show(field=fieldset.U, show_time=datetime(2002, 1, 10, 2))
# Note that the particle locations do not change, but remain at the locations corresponding to the end of the last integration. To remove them from the plot, we set the argument `with_particles` equal to `False`.
pset.show(field=fieldset.U, show_time=datetime(2002, 1, 10, 2), with_particles=False)
# By setting the `field` argument equal to `vector`, we can display the velocity in full vector form.
pset.show(field='vector')
# The normalized vector field is colored by speed. To control the maximum speed value on the colorbar, set the `vmax` argument equal to the desired value.
pset.show(field='vector', vmax=3.0, domain=[-31, -39, 33, 18])
# We can change the projection of the plot by providing one of the [projections](https://scitools.org.uk/cartopy/docs/v0.15/crs/projections.html) from `cartopy`. For example, to plot on a Robinson projection , we use `projection=cartopy.crs.Robinson()`. Note that not all projections support gridlines, so these may not be shown.
try: # Within a try/pass for unit testing on machines without cartopy installed
import cartopy
pset.show(field='vector', vmax=3.0, domain=[-31, -39, 33, 18], projection=cartopy.crs.Robinson())
except:
pass
# If we want to save the file rather than show it, we set the argument `savefile` equal to the `'path/to/save/file'`.
pset.show(field='vector', vmax=3.0, domain=[-31, -39, 33, 18], land=True, savefile='particles')
|
parcels/examples/tutorial_plotting.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Concatenar y apendizar data sets
import pandas as pd
red_wine = pd.read_csv("/Users/nuelcodes/Data-Science-Python/datasets/wine/winequality-red.csv", sep=";")
white_wine = pd.read_csv("/Users/nuelcodes/Data-Science-Python/datasets/wine/winequality-white.csv", sep=";")
red_wine.head()
white_wine.head()
red_wine.columns.values
red_wine.shape
white_wine.columns.values
white_wine.shape
# En python tenemos dos tipos de ejes
#
# * axis = 0 denota el eje horizontal
# * axis = 1 denota el eje vertical
wine_data = pd.concat([red_wine, white_wine], axis= 0)
wine_data.shape
wine_data.head()
# Scramble
data1 = wine_data.head(10)
data2 = wine_data[300:310]
data3 = wine_data.tail(10)
wine_scramble = pd.concat([data1, data2, data3], axis=0)
wine_scramble
wine_scramble = pd.concat([data2, data3, data1], axis=0)
wine_scramble
# # Datos distribuidos
data = pd.read_csv("/Users/nuelcodes/Data-Science-Python/datasets/distributed-data/001.csv")
data.head()
# * Importar el primer fichero
# * Hacemos un buble para ir recorriendo todos y cada uno de los ficheros
# * Importante tener una consistencia en el nombre de los ficheros
# * Cada uno de ellos debe apendizarse (añadir al final) del primer fichero que ya habiamos cargado
# * Repetimos el bucle hasta que no queden ficheros
# +
filepath = "/Users/nuelcodes/Data-Science-Python/datasets/distributed-data/"
data = pd.read_csv("/Users/nuelcodes/Data-Science-Python/datasets/distributed-data/001.csv")
for i in range(2,333):
if i < 10:
filename = "00" + str(i)
if 10 <= i < 100:
filename = "0" + str(i)
if i >= 100:
filename = str(i)
file = filepath + filename + ".csv"
temp_data = pd.read_csv(file)
data = pd.concat([data, temp_data], axis = 0)
# -
data.shape
data.tail()
data.head()
# +
import matplotlib.pyplot as plt
plt.hist(data["ID"])
# -
plt.hist(data["sulfate"])
plt.hist(data["nitrate"])
# # Joins de datasets
filepath = "/Users/nuelcodes/Data-Science-Python/datasets/athletes/"
data_main = pd.read_csv(filepath + "Medals.csv", encoding= "ISO-8859-1")
data_main.head()
a = data_main["Athlete"].unique().tolist()
len(a)
data_main.shape
# Lugar de donde vienen los atletas
data_country = pd.read_csv(filepath + "Athelete_Country_Map.csv", encoding= "ISO-8859-1")
data_country.head()
data_country.shape
data_country[data_country["Athlete"] == "Aleksandar Ciric"]
# Deporte al cual se le asocia el atleta
data_sports = pd.read_csv(filepath + "Athelete_Sports_Map.csv", encoding= "ISO-8859-1")
data_sports.head()
len(data_sports)
data_sports[
(data_sports["Athlete"] == "<NAME>") |
(data_sports["Athlete"] == "<NAME>") |
(data_sports["Athlete"] == "<NAME>")
]
data_country_dp = data_country.drop_duplicates(subset = "Athlete")
# comparar la longitud de data_country_dp con a
# donde a es los atletas unicos del data set original
len(data_country_dp) == len(a)
len(data_country_dp)
# Hacer un merge del data principal con el data del pais
data_main_country = pd.merge(left = data_main, right = data_country_dp,
left_on="Athlete", right_on="Athlete")
data_main_country.tail()
data_main_country.shape
data_main_country[data_main_country["Athlete"] == "<NAME>"]
# Eliminando los duplicados del data set de Deportes
data_sports_dp = data_sports.drop_duplicates(subset="Athlete")
# Comparando longitud data_sports_dp con el data set sin duplicados del data set original
len(data_sports_dp) == len(a)
# Merge final
data_final = pd.merge(left = data_main_country, right = data_sports_dp,
left_on = "Athlete", right_on = "Athlete")
data_final.head()
len(data_final)
# # Tipos de Joins
from IPython.display import Image
import numpy as np
# **Inner Join <= A (Left Join), B (Right Join) <= Outer Join**
# +
# Los atletas que resultaron, se utilizaran para eliminar informacion de ellos
out_athletes = np.random.choice(data_main["Athlete"], size = 6, replace = False)
# -
out_athletes
# +
data_country_dlt = data_country_dp[(~data_country_dp["Athlete"].isin(out_athletes)) &
(data_country_dp["Athlete"] != "<NAME>")]
data_sports_dlt = data_sports_dp[(~data_sports_dp["Athlete"].isin(out_athletes)) &
(data_sports_dp["Athlete"] != "<NAME>")]
data_main_dlt = data_main[(~data_main["Athlete"].isin(out_athletes)) &
(data_main["Athlete"] != "<NAME>")]
# -
data_country_dlt.head()
len(data_country_dlt)
len(data_sports_dlt)
len(data_main_dlt)
len(data_country_dp) - len(data_country_dlt)
# ## Inner Join
#
# * Devuelve un data frame con las filas que tienen valor tanto en el primer como en el segundo data frame que estamos uniendo
# * El número de filas será igual al número de filas **comunes** que tengan ambos data sets
# * Data Set A tiene 60 filas
# * Data Set B tiene 30 filas
# * Ambos comparten 30 filas
# * Entonces A Inner Join B tendrá 30 filas
# * En términos de teoría de conjuntos, se trata de la intersección de los dos conjuntos
Image(filename="/Users/nuelcodes/Data-Science-Python/notebooks/resources/inner-join.png")
# +
# Unir un data set que contenga toda la información, con uno que le falte una parte
# data_main contiene toda la informacion
# data_country_dlt le falta la informacion de 7 atletas
merged_inner = pd.merge(left = data_main, right = data_country_dlt, how = "inner", left_on = "Athlete", right_on = "Athlete")
# -
len(merged_inner)
merged_inner.head()
# ## Left Join
#
# * Devuelve un data frame con las filas que tuvieran valor en el dataset de la izquierda, sin importar si tienen correspondencia en la de la derecha o no.
# * Las filas del data frame final que no correspondan a ningula fila del data frame derecho, tendrán NA's en las columnas del data frame derecho.
# * El número de filas será igual al número de filas del data frame izquierdo
# * Data Set A tiene 60 filas
# * Data Set B tiene 50 filas
# * Entonces A Left Join B tendrá 60 filas
# * En términos de teoría de conjuntos, se trata del propio data set de la izquierda quien, además tiene la intersección en su interior
Image(filename="/Users/nuelcodes/Data-Science-Python/notebooks/resources/left-join.png")
merged_left = pd.merge(left = data_main, right = data_country_dlt, how = "left", left_on = "Athlete", right_on = "Athlete")
len(merged_left)
merged_left.head()
# ## Right Join
#
# * Devuelve un data frame con las filas que tuvieran valor en el dataset de la derecha, sin importar si tienen correspondencia en el de la izquierda o no.
# * Las filas del data frame final que no correspondan a ninguna fila del data frame izquierdo, tendrán NA's en las columnas del data frame izquierdo.
# * Data Set A tiene 60 filas
# * Data Set B tiene 50 filas
# * Entonces A Right Join B tendrá 50 filas
# * En términos de teoría de conjuntos, se trata del propio data set de la derecha quien, además tiene la intersección en su interior.
Image("/Users/nuelcodes/Data-Science-Python/notebooks/resources/right-join.png")
merged_right = pd.merge(left = data_main_dlt, right = data_country_dp, how = "right", left_on = "Athlete", right_on = "Athlete")
len(merged_right)
merged_right.head(10)
# ## Outer Join
#
# * Devuleve un data frame con todas las filas de ambos, reemplazando las ausencias de uno o de otro con NA's en la región específica.
# * Las filas del data frame final que no correspondan a ninguna fila del data frame derecho (o izquierdo), tendrán NA's en las columnas del data frame derecho (o izquierdo).
# * El número de filas será igual al máximo número de filas de ambos data frames.
# * Data Set A tinen 60 filas
# * Data Set B tiene 50 filas
# * Ambos comparten 30 filas
# * Entonces A Outer Join B tendrá 60 + 50 -30 = 80 filas
# * En términos de teoría de conjuntos, se trata de la unión de conjuntos.
Image("/Users/nuelcodes/Data-Science-Python/notebooks/resources/outer-join.png")
data_country_mc = data_country_dlt.append(
{
"Athlete" : "<NAME>",
"Country" : "Mexico"
},ignore_index = True
)
data_country_mc
merged_outer = pd.merge(left = data_main, right = data_country_mc, how = "outer", left_on = "Athlete", right_on = "Athlete")
len(merged_outer)
merged_outer.tail()
len(data_main)
len(data_main_dlt)
len(data_country_dp)
len(data_country_dlt)
len(merged_inner)
len(merged_left)
len(merged_right)
len(merged_outer)
|
scratch/T2 - 4 - Data Cleaning - Concatenacion de datos.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
# <script>
# window.dataLayer = window.dataLayer || [];
# function gtag(){dataLayer.push(arguments);}
# gtag('js', new Date());
#
# gtag('config', 'UA-59152712-8');
# </script>
#
# # Start-to-Finish Example: `GiRaFFE_NRPy` 3D tests
#
# ### Author: <NAME>
#
# ### Adapted from [Start-to-Finish Example: Head-On Black Hole Collision](../Tutorial-Start_to_Finish-BSSNCurvilinear-Two_BHs_Collide.ipynb)
#
# ## This module implements a basic GRFFE code to evolve one-dimensional GRFFE waves.
#
# ### NRPy+ Source Code for this module:
# * [GiRaFFEfood_NRPy/GiRaFFEfood_NRPy_Exact_Wald.py](../../edit/in_progress/GiRaFFEfood_NRPy/GiRaFFEfood_NRPy_Exact_Wald.py) [\[**tutorial**\]](Tutorial-GiRaFFEfood_NRPy_Exact_Wald.ipynb) Generates Exact Wald initial data
# * [GiRaFFEfood_NRPy/GiRaFFEfood_NRPy_Aligned_Rotator.py](../../edit/in_progress/GiRaFFEfood_NRPy/GiRaFFEfood_NRPy_Aligned_Rotator.py) [\[**tutorial**\]](Tutorial-GiRaFFEfood_NRPy_Aligned_Rotator.ipynb) Generates Aligned Rotator initial data
# * [GiRaFFEfood_NRPy/GiRaFFEfood_NRPy_1D_tests.py](../../edit/in_progress/GiRaFFEfood_NRPy/GiRaFFEfood_NRPy_1D_tests.py) [\[**tutorial**\]](Tutorial-GiRaFFEfood_NRPy_1D_tests.ipynb) Generates Alfvén Wave initial data.
# * [GiRaFFE_NRPy/Afield_flux.py](../../edit/in_progress/GiRaFFE_NRPy/Afield_flux.py) [\[**tutorial**\]](Tutorial-GiRaFFE_NRPy-Afield_flux.ipynb) Generates the expressions to find the flux term of the induction equation.
# * [GiRaFFE_NRPy/GiRaFFE_NRPy_A2B.py](../../edit/in_progress/GiRaFFE_NRPy/GiRaFFE_NRPy_A2B.py) [\[**tutorial**\]](Tutorial-GiRaFFE_NRPy-Afield_flux.ipynb) Generates the driver to compute the magnetic field from the vector potential/
# * [GiRaFFE_NRPy/GiRaFFE_NRPy_BCs.py](../../edit/in_progress/GiRaFFE_NRPy/GiRaFFE_NRPy_BCs.py) [\[**tutorial**\]](Tutorial-GiRaFFE_NRPy-BCs.ipynb) Generates the code to apply boundary conditions to the vector potential, scalar potential, and three-velocity.
# * [GiRaFFE_NRPy/GiRaFFE_NRPy_C2P_P2C.py](../../edit/in_progress/GiRaFFE_NRPy/GiRaFFE_NRPy_C2P_P2C.py) [\[**tutorial**\]](Tutorial-GiRaFFE_NRPy-C2P_P2C.ipynb) Generates the conservative-to-primitive and primitive-to-conservative solvers.
# * [GiRaFFE_NRPy/GiRaFFE_NRPy_Metric_Face_Values.py](../../edit/in_progress/GiRaFFE_NRPy/GiRaFFE_NRPy_Metric_Face_Values.py) [\[**tutorial**\]](Tutorial-GiRaFFE_NRPy-Metric_Face_Values.ipynb) Generates code to interpolate metric gridfunctions to cell faces.
# * [GiRaFFE_NRPy/GiRaFFE_NRPy_PPM.py](../../edit/in_progress/GiRaFFE_NRPy/GiRaFFE_NRPy_PPM.py) [\[**tutorial**\]](Tutorial-GiRaFFE_NRPy-PPM.ipynb) Genearates code to reconstruct primitive variables on cell faces.
# * [GiRaFFE_NRPy/GiRaFFE_NRPy_Source_Terms.py](../../edit/in_progress/GiRaFFE_NRPy/GiRaFFE_NRPy_Source_Terms.py) [\[**tutorial**\]](Tutorial-GiRaFFE_NRPy-Source_Terms.ipynb) Generates the expressions to find the flux term of the Poynting flux evolution equation.
# * [GiRaFFE_NRPy/Stilde_flux.py](../../edit/in_progress/GiRaFFE_NRPy/Stilde_flux.py) [\[**tutorial**\]](Tutorial-GiRaFFE_NRPy-Stilde_flux.ipynb) Generates the expressions to find the flux term of the Poynting flux evolution equation.
# * [../GRFFE/equations.py](../../edit/GRFFE/equations.py) [\[**tutorial**\]](../Tutorial-GRFFE_Equations-Cartesian.ipynb) Generates code necessary to compute the source terms.
# * [../GRHD/equations.py](../../edit/GRHD/equations.py) [\[**tutorial**\]](../Tutorial-GRHD_Equations-Cartesian.ipynb) Generates code necessary to compute the source terms.
#
# Here we use NRPy+ to generate the C source code necessary to set up initial data for an Alfvén wave (see [the original GiRaFFE paper](https://arxiv.org/pdf/1704.00599.pdf)). Then we use it to generate the RHS expressions for [Method of Lines](https://reference.wolfram.com/language/tutorial/NDSolveMethodOfLines.html) time integration based on the [explicit Runge-Kutta fourth-order scheme](https://en.wikipedia.org/wiki/Runge%E2%80%93Kutta_methods) (RK4).
# <a id='toc'></a>
#
# # Table of Contents
# $$\label{toc}$$
#
# This notebook is organized as follows
#
# 1. [Step 1](#initializenrpy): Set core NRPy+ parameters for numerical grids
# 1. [Step 2](#grffe): Output C code for GRFFE evolution
# 1. [Step 2.a](#mol): Output macros for Method of Lines timestepping
# 1. [Step 3](#gf_id): Import `GiRaFFEfood_NRPy` initial data modules
# 1. [Step 4](#cparams): Output C codes needed for declaring and setting Cparameters; also set `free_parameters.h`
# 1. [Step 5](#mainc): `GiRaFFE_NRPy_standalone.c`: The Main C Code
# <a id='setup'></a>
#
# # Step 1: Set up core functions and parameters for solving GRFFE equations \[Back to [top](#toc)\]
# $$\label{setup}$$
#
# +
import shutil, os, sys # Standard Python modules for multiplatform OS-level functions
# First, we'll add the parent directory to the list of directories Python will check for modules.
nrpy_dir_path = os.path.join("..")
if nrpy_dir_path not in sys.path:
sys.path.append(nrpy_dir_path)
# Step P1: Import needed NRPy+ core modules:
from outputC import outCfunction, lhrh # NRPy+: Core C code output module
import sympy as sp # SymPy: The Python computer algebra package upon which NRPy+ depends
import finite_difference as fin # NRPy+: Finite difference C code generation module
import NRPy_param_funcs as par # NRPy+: Parameter interface
import grid as gri # NRPy+: Functions having to do with numerical grids
import indexedexp as ixp # NRPy+: Symbolic indexed expression (e.g., tensors, vectors, etc.) support
import cmdline_helper as cmd # NRPy+: Multi-platform Python command-line interface
# Step P2: Create C code output directory:
Ccodesdir = os.path.join("GiRaFFE_standalone_Ccodes/")
# First remove C code output directory if it exists
# Courtesy https://stackoverflow.com/questions/303200/how-do-i-remove-delete-a-folder-that-is-not-empty
# # !rm -r ScalarWaveCurvilinear_Playground_Ccodes
shutil.rmtree(Ccodesdir, ignore_errors=True)
# Then create a fresh directory
cmd.mkdir(Ccodesdir)
# Step P3: Create executable output directory:
outdir = os.path.join(Ccodesdir,"output/")
cmd.mkdir(Ccodesdir)
cmd.mkdir(outdir)
# Step P5: Set timestepping algorithm (we adopt the Method of Lines)
REAL = "double" # Best to use double here.
default_CFL_FACTOR= 0.5 # (GETS OVERWRITTEN WHEN EXECUTED.) In pure axisymmetry (symmetry_axes = 2 below) 1.0 works fine. Otherwise 0.5 or lower.
# Step P6: Set the finite differencing order to 2.
par.set_parval_from_str("finite_difference::FD_CENTDERIVS_ORDER",2)
thismodule = "Start_to_Finish-GiRaFFE_NRPy-1D_tests"
TINYDOUBLE = par.Cparameters("REAL", thismodule, "TINYDOUBLE", 1e-100)
import GiRaFFE_NRPy.GiRaFFE_NRPy_Main_Driver as md
# par.set_paramsvals_value("GiRaFFE_NRPy.GiRaFFE_NRPy_C2P_P2C::enforce_speed_limit_StildeD = False")
par.set_paramsvals_value("GiRaFFE_NRPy.GiRaFFE_NRPy_C2P_P2C::enforce_current_sheet_prescription = False")
# -
# <a id='grffe'></a>
#
# # Step 2: Output C code for GRFFE evolution \[Back to [top](#toc)\]
# $$\label{grffe}$$
#
# We will first write the C codes needed for GRFFE evolution. We have already written a module to generate all these codes and call the functions in the appropriate order, so we will import that here. We will take the slightly unusual step of doing this before we generate the initial data functions because the main driver module will register all the gridfunctions we need. It will also generate functions that, in addition to their normal spot in the MoL timestepping, will need to be called during the initial data step to make sure all the variables are appropriately filled in.
#
# All of this is handled with a single call to `GiRaFFE_NRPy_Main_Driver_generate_all()`, which will register gridfunctions, write all the C code kernels, and write the C code functions to call those.
md.GiRaFFE_NRPy_Main_Driver_generate_all(Ccodesdir)
# <a id='mol'></a>
#
# ## Step 2.a: Output macros for Method of Lines timestepping \[Back to [top](#toc)\]
# $$\label{mol}$$
#
# Now, we generate the code to implement the method of lines using the fourth-order Runge-Kutta algorithm.
# +
RK_method = "RK4"
# Step 3: Generate Runge-Kutta-based (RK-based) timestepping code.
# As described above the Table of Contents, this is a 3-step process:
# 3.A: Evaluate RHSs (RHS_string)
# 3.B: Apply boundary conditions (post_RHS_string, pt 1)
import MoLtimestepping.C_Code_Generation as MoL
from MoLtimestepping.RK_Butcher_Table_Dictionary import Butcher_dict
RK_order = Butcher_dict[RK_method][1]
cmd.mkdir(os.path.join(Ccodesdir,"MoLtimestepping/"))
MoL.MoL_C_Code_Generation(RK_method,
RHS_string = """
GiRaFFE_NRPy_RHSs(¶ms,auxevol_gfs,RK_INPUT_GFS,RK_OUTPUT_GFS);""",
post_RHS_string = """
GiRaFFE_NRPy_post_step(¶ms,xx,auxevol_gfs,RK_OUTPUT_GFS,n+1);\n""",
outdir = os.path.join(Ccodesdir,"MoLtimestepping/"))
# -
# <a id='gf_id'></a>
#
# # Step 3: Import `GiRaFFEfood_NRPy` initial data modules \[Back to [top](#toc)\]
# $$\label{gf_id}$$
#
# With the preliminaries out of the way, we will write the C functions to set up initial data. There are two categories of initial data that must be set: the spacetime metric variables, and the GRFFE plasma variables. We will set up the spacetime first.
# +
# There are several initial data routines we need to test. We'll control which one we use with a string option
initial_data = "ExactWald" # Valid options: "ExactWald", "AlignedRotator"
spacetime = "ShiftedKerrSchild" # Valid options: "ShiftedKerrSchild", "flat"
if spacetime == "ShiftedKerrSchild":
# Exact Wald is more complicated. We'll need the Shifted Kerr Schild metric in Cartesian coordinates.
import BSSN.ShiftedKerrSchild as sks
sks.ShiftedKerrSchild(True)
import reference_metric as rfm
par.set_parval_from_str("reference_metric::CoordSystem","Cartesian")
rfm.reference_metric()
# Use the Jacobian matrix to transform the vectors to Cartesian coordinates.
par.set_parval_from_str("reference_metric::CoordSystem","Spherical")
rfm.reference_metric()
Jac_dUCart_dDrfmUD,Jac_dUrfm_dDCartUD = rfm.compute_Jacobian_and_inverseJacobian_tofrom_Cartesian()
# Transform the coordinates of the Jacobian matrix from spherical to Cartesian:
par.set_parval_from_str("reference_metric::CoordSystem","Cartesian")
rfm.reference_metric()
tmpa,tmpb,tmpc = sp.symbols("tmpa,tmpb,tmpc")
for i in range(3):
for j in range(3):
Jac_dUCart_dDrfmUD[i][j] = Jac_dUCart_dDrfmUD[i][j].subs([(rfm.xx[0],tmpa),(rfm.xx[1],tmpb),(rfm.xx[2],tmpc)])
Jac_dUCart_dDrfmUD[i][j] = Jac_dUCart_dDrfmUD[i][j].subs([(tmpa,rfm.xxSph[0]),(tmpb,rfm.xxSph[1]),(tmpc,rfm.xxSph[2])])
Jac_dUrfm_dDCartUD[i][j] = Jac_dUrfm_dDCartUD[i][j].subs([(rfm.xx[0],tmpa),(rfm.xx[1],tmpb),(rfm.xx[2],tmpc)])
Jac_dUrfm_dDCartUD[i][j] = Jac_dUrfm_dDCartUD[i][j].subs([(tmpa,rfm.xxSph[0]),(tmpb,rfm.xxSph[1]),(tmpc,rfm.xxSph[2])])
gammaSphDD = ixp.zerorank2()
for i in range(3):
for j in range(3):
gammaSphDD[i][j] += sks.gammaSphDD[i][j].subs(sks.r,rfm.xxSph[0]).subs(sks.th,rfm.xxSph[1])
betaSphU = ixp.zerorank1()
for i in range(3):
betaSphU[i] += sks.betaSphU[i].subs(sks.r,rfm.xxSph[0]).subs(sks.th,rfm.xxSph[1])
alpha = sks.alphaSph.subs(sks.r,rfm.xxSph[0]).subs(sks.th,rfm.xxSph[1])
gammaDD = rfm.basis_transform_tensorDD_from_rfmbasis_to_Cartesian(Jac_dUrfm_dDCartUD, gammaSphDD)
unused_gammaUU,gammaDET = ixp.symm_matrix_inverter3x3(gammaDD)
sqrtgammaDET = sp.sqrt(gammaDET)
betaU = rfm.basis_transform_vectorD_from_rfmbasis_to_Cartesian(Jac_dUrfm_dDCartUD, betaSphU)
# Description and options for this initial data
desc = "Generate a spinning black hole with Shifted Kerr Schild metric."
loopopts_id ="AllPoints,Read_xxs"
elif spacetime == "flat":
gammaDD = ixp.zerorank2(DIM=3)
for i in range(3):
for j in range(3):
if i==j:
gammaDD[i][j] = sp.sympify(1) # else: leave as zero
betaU = ixp.zerorank1() # All should be 0
alpha = sp.sympify(1)
# Description and options for this initial data
desc = "Generate a flat spacetime metric."
loopopts_id ="AllPoints" # we don't need to read coordinates for flat spacetime.
name = "set_initial_spacetime_metric_data"
values_to_print = [\
lhrh(lhs=gri.gfaccess("auxevol_gfs","gammaDD00"),rhs=gammaDD[0][0]),\
lhrh(lhs=gri.gfaccess("auxevol_gfs","gammaDD01"),rhs=gammaDD[0][1]),\
lhrh(lhs=gri.gfaccess("auxevol_gfs","gammaDD02"),rhs=gammaDD[0][2]),\
lhrh(lhs=gri.gfaccess("auxevol_gfs","gammaDD11"),rhs=gammaDD[1][1]),\
lhrh(lhs=gri.gfaccess("auxevol_gfs","gammaDD12"),rhs=gammaDD[1][2]),\
lhrh(lhs=gri.gfaccess("auxevol_gfs","gammaDD22"),rhs=gammaDD[2][2]),\
lhrh(lhs=gri.gfaccess("auxevol_gfs","betaU0"),rhs=betaU[0]),\
lhrh(lhs=gri.gfaccess("auxevol_gfs","betaU1"),rhs=betaU[1]),\
lhrh(lhs=gri.gfaccess("auxevol_gfs","betaU2"),rhs=betaU[2]),\
lhrh(lhs=gri.gfaccess("auxevol_gfs","alpha"),rhs=alpha),\
]
outCfunction(
outfile = os.path.join(Ccodesdir,name+".h"), desc=desc, name=name,
params ="const paramstruct *params,REAL *xx[3],REAL *auxevol_gfs",
body = fin.FD_outputC("returnstring",values_to_print,params="outCverbose=False"),
loopopts = loopopts_id)
# -
# Now, we will write out the initial data function for the GRFFE variables.
# +
import GiRaFFEfood_NRPy.GiRaFFEfood_NRPy as gid
if initial_data=="ExactWald":
gid.GiRaFFEfood_NRPy_generate_initial_data(ID_type = initial_data, stagger_enable = False,M=sks.M,KerrSchild_radial_shift=sks.r0,gammaDD=gammaDD,sqrtgammaDET=sqrtgammaDET)
desc = "Generate exact Wald initial test data for GiRaFFEfood_NRPy."
elif initial_data=="SplitMonopole":
gid.GiRaFFEfood_NRPy_generate_initial_data(ID_type = initial_data, stagger_enable = False,M=sks.M,a=sks.a,KerrSchild_radial_shift=sks.r0,alpha=alpha,betaU=betaSphU,gammaDD=gammaDD,sqrtgammaDET=sqrtgammaSphDET)
desc = "Generate Split Monopole initial test data for GiRaFFEfood_NRPy."
elif initial_data=="AlignedRotator":
gf.GiRaFFEfood_NRPy_generate_initial_data(ID_type = initial_data, stagger_enable = True)
desc = "Generate aligned rotator initial test data for GiRaFFEfood_NRPy."
else:
print("Unsupported Initial Data string "+initial_data+"! Supported ID: ExactWald, or SplitMonopole")
name = "initial_data"
values_to_print = [\
lhrh(lhs=gri.gfaccess("out_gfs","AD0"),rhs=gid.AD[0]),\
lhrh(lhs=gri.gfaccess("out_gfs","AD1"),rhs=gid.AD[1]),\
lhrh(lhs=gri.gfaccess("out_gfs","AD2"),rhs=gid.AD[2]),\
lhrh(lhs=gri.gfaccess("auxevol_gfs","ValenciavU0"),rhs=gid.ValenciavU[0]),\
lhrh(lhs=gri.gfaccess("auxevol_gfs","ValenciavU1"),rhs=gid.ValenciavU[1]),\
lhrh(lhs=gri.gfaccess("auxevol_gfs","ValenciavU2"),rhs=gid.ValenciavU[2]),\
# lhrh(lhs=gri.gfaccess("auxevol_gfs","BU0"),rhs=gid.BU[0]),\
# lhrh(lhs=gri.gfaccess("auxevol_gfs","BU1"),rhs=gid.BU[1]),\
# lhrh(lhs=gri.gfaccess("auxevol_gfs","BU2"),rhs=gid.BU[2]),\
lhrh(lhs=gri.gfaccess("out_gfs","psi6Phi"),rhs=sp.sympify(0))\
]
outCfunction(
outfile = os.path.join(Ccodesdir,name+".h"), desc=desc, name=name,
params ="const paramstruct *params,REAL *xx[3],REAL *auxevol_gfs,REAL *out_gfs",
body = fin.FD_outputC("returnstring",values_to_print,params="outCverbose=False"),
loopopts ="AllPoints,Read_xxs")
# -
# <a id='cparams'></a>
#
# # Step 4: Output C codes needed for declaring and setting Cparameters; also set `free_parameters.h` \[Back to [top](#toc)\]
# $$\label{cparams}$$
#
# Based on declared NRPy+ Cparameters, first we generate `declare_Cparameters_struct.h`, `set_Cparameters_default.h`, and `set_Cparameters[-SIMD].h`.
#
# Then we output `free_parameters.h`, which sets initial data parameters, as well as grid domain & reference metric parameters, applying `domain_size` and `sinh_width`/`SymTP_bScale` (if applicable) as set above
# +
# Step 3.e: Output C codes needed for declaring and setting Cparameters; also set free_parameters.h
# Step 3.e.i: Generate declare_Cparameters_struct.h, set_Cparameters_default.h, and set_Cparameters[-SIMD].h
par.generate_Cparameters_Ccodes(os.path.join(Ccodesdir))
# Step 3.e.ii: Set free_parameters.h
with open(os.path.join(Ccodesdir,"free_parameters.h"),"w") as file:
file.write("""// Override parameter defaults with values based on command line arguments and NGHOSTS.
params.Nxx0 = atoi(argv[1]);
params.Nxx1 = atoi(argv[2]);
params.Nxx2 = atoi(argv[3]);
params.Nxx_plus_2NGHOSTS0 = params.Nxx0 + 2*NGHOSTS;
params.Nxx_plus_2NGHOSTS1 = params.Nxx1 + 2*NGHOSTS;
params.Nxx_plus_2NGHOSTS2 = params.Nxx2 + 2*NGHOSTS;
// Step 0d: Set up space and time coordinates
// Step 0d.i: Declare \Delta x^i=dxx{0,1,2} and invdxx{0,1,2}, as well as xxmin[3] and xxmax[3]:
const REAL xxmin[3] = {-1.5,-0.1,-0.1};
const REAL xxmax[3] = { 1.5, 0.1, 0.1};
//const REAL xxmin[3] = {-1.5,-1.5,-1.5};
//const REAL xxmax[3] = { 1.5, 1.5, 1.5};
params.dxx0 = (xxmax[0] - xxmin[0]) / ((REAL)params.Nxx0+1);
params.dxx1 = (xxmax[1] - xxmin[1]) / ((REAL)params.Nxx1+1);
params.dxx2 = (xxmax[2] - xxmin[2]) / ((REAL)params.Nxx2+1);
printf("dxx0,dxx1,dxx2 = %.5e,%.5e,%.5e\\n",params.dxx0,params.dxx1,params.dxx2);
params.invdx0 = 1.0 / params.dxx0;
params.invdx1 = 1.0 / params.dxx1;
params.invdx2 = 1.0 / params.dxx2;
const int poison_grids = 0;
// Standard GRFFE parameters:
params.GAMMA_SPEED_LIMIT = 2000.0;
params.diss_strength = 0.1;
""")
if initial_data=="ExactWald":
with open(os.path.join(Ccodesdir,"free_parameters.h"),"a") as file:
file.write("""params.r0 = 0.4;
params.a = 0.0;
""")
# -
# <a id='bc_functs'></a>
#
# # Step 4: Set up boundary condition functions for chosen singular, curvilinear coordinate system \[Back to [top](#toc)\]
# $$\label{bc_functs}$$
#
# Next apply singular, curvilinear coordinate boundary conditions [as documented in the corresponding NRPy+ tutorial notebook](Tutorial-Start_to_Finish-Curvilinear_BCs.ipynb)
#
# ...But, for the moment, we're actually just using this because it writes the file `gridfunction_defines.h`.
import CurviBoundaryConditions.CurviBoundaryConditions as cbcs
cbcs.Set_up_CurviBoundaryConditions(os.path.join(Ccodesdir,"boundary_conditions/"),Cparamspath=os.path.join("../"),enable_copy_of_static_Ccodes=False)
# <a id='mainc'></a>
#
# # Step 5: `GiRaFFE_NRPy_standalone.c`: The Main C Code \[Back to [top](#toc)\]
# $$\label{mainc}$$
# +
# Part P0: Define REAL, set the number of ghost cells NGHOSTS (from NRPy+'s FD_CENTDERIVS_ORDER),
# and set the CFL_FACTOR (which can be overwritten at the command line)
with open(os.path.join(Ccodesdir,"GiRaFFE_NRPy_REAL__NGHOSTS__CFL_FACTOR.h"), "w") as file:
file.write("""
// Part P0.a: Set the number of ghost cells, from NRPy+'s FD_CENTDERIVS_ORDER
#define NGHOSTS """+str(3)+"""
#define NGHOSTS_A2B """+str(2)+"""
// Part P0.b: Set the numerical precision (REAL) to double, ensuring all floating point
// numbers are stored to at least ~16 significant digits
#define REAL """+REAL+"""
// Part P0.c: Set the CFL Factor. Can be overwritten at command line.
REAL CFL_FACTOR = """+str(default_CFL_FACTOR)+";")
# +
# %%writefile $Ccodesdir/GiRaFFE_NRPy_standalone.c
// Step P0: Define REAL and NGHOSTS; and declare CFL_FACTOR. This header is generated in NRPy+.
#include "GiRaFFE_NRPy_REAL__NGHOSTS__CFL_FACTOR.h"
#include "declare_Cparameters_struct.h"
const int NSKIP_1D_OUTPUT = 1;
// Step P1: Import needed header files
#include "stdio.h"
#include "stdlib.h"
#include "math.h"
#include "time.h"
#include "stdint.h" // Needed for Windows GCC 6.x compatibility
#ifndef M_PI
#define M_PI 3.141592653589793238462643383279502884L
#endif
#ifndef M_SQRT1_2
#define M_SQRT1_2 0.707106781186547524400844362104849039L
#endif
// Step P2: Declare the IDX4S(gf,i,j,k) macro, which enables us to store 4-dimensions of
// data in a 1D array. In this case, consecutive values of "i"
// (all other indices held to a fixed value) are consecutive in memory, where
// consecutive values of "j" (fixing all other indices) are separated by
// Nxx_plus_2NGHOSTS0 elements in memory. Similarly, consecutive values of
// "k" are separated by Nxx_plus_2NGHOSTS0*Nxx_plus_2NGHOSTS1 in memory, etc.
#define IDX4S(g,i,j,k) \
( (i) + Nxx_plus_2NGHOSTS0 * ( (j) + Nxx_plus_2NGHOSTS1 * ( (k) + Nxx_plus_2NGHOSTS2 * (g) ) ) )
#define IDX4ptS(g,idx) ( (idx) + (Nxx_plus_2NGHOSTS0*Nxx_plus_2NGHOSTS1*Nxx_plus_2NGHOSTS2) * (g) )
#define IDX3S(i,j,k) ( (i) + Nxx_plus_2NGHOSTS0 * ( (j) + Nxx_plus_2NGHOSTS1 * ( (k) ) ) )
#define LOOP_REGION(i0min,i0max, i1min,i1max, i2min,i2max) \
for(int i2=i2min;i2<i2max;i2++) for(int i1=i1min;i1<i1max;i1++) for(int i0=i0min;i0<i0max;i0++)
#define LOOP_ALL_GFS_GPS(ii) _Pragma("omp parallel for") \
for(int (ii)=0;(ii)<Nxx_plus_2NGHOSTS_tot*NUM_EVOL_GFS;(ii)++)
// Step P3: Set gridfunction macros
#include "boundary_conditions/gridfunction_defines.h"
// Step P4: Include the RHS, BC, and primitive recovery functions
#include "GiRaFFE_NRPy_Main_Driver.h"
// Step P5: Include the initial data functions
#include "set_initial_spacetime_metric_data.h"
#include "initial_data.h"
// main() function:
// Step 0: Read command-line input, set up grid structure, allocate memory for gridfunctions, set up coordinates
// Step 1: Set up scalar wave initial data
// Step 2: Evolve scalar wave initial data forward in time using Method of Lines with RK4 algorithm,
// applying quadratic extrapolation outer boundary conditions.
// Step 3: Output relative error between numerical and exact solution.
// Step 4: Free all allocated memory
int main(int argc, const char *argv[]) {
paramstruct params;
#include "set_Cparameters_default.h"
// Step 0a: Read command-line input, error out if nonconformant
if(argc != 4 || atoi(argv[1]) < NGHOSTS || atoi(argv[2]) < NGHOSTS || atoi(argv[3]) < NGHOSTS) {
printf("Error: Expected three command-line arguments: ./GiRaFFE_NRPy_standalone [Nx] [Ny] [Nz],\n");
printf("where Nx is the number of grid points in the x direction, and so forth.\n");
printf("Nx,Ny,Nz MUST BE larger than NGHOSTS (= %d)\n",NGHOSTS);
exit(1);
}
// Step 0c: Set free parameters, overwriting Cparameters defaults
// by hand or with command-line input, as desired.
#include "free_parameters.h"
#include "set_Cparameters-nopointer.h"
// ... and then set up the numerical grid structure in time:
const REAL t_final = 0.5;
const REAL CFL_FACTOR = 0.5; // Set the CFL Factor
// Step 0c: Allocate memory for gridfunctions
const int Nxx_plus_2NGHOSTS_tot = Nxx_plus_2NGHOSTS0*Nxx_plus_2NGHOSTS1*Nxx_plus_2NGHOSTS2;
// Step 0k: Allocate memory for gridfunctions
#include "MoLtimestepping/RK_Allocate_Memory.h"
REAL *restrict auxevol_gfs = (REAL *)malloc(sizeof(REAL) * NUM_AUXEVOL_GFS * Nxx_plus_2NGHOSTS_tot);
REAL *evol_gfs_exact = (REAL *)malloc(sizeof(REAL) * NUM_EVOL_GFS * Nxx_plus_2NGHOSTS_tot);
REAL *auxevol_gfs_exact = (REAL *)malloc(sizeof(REAL) * NUM_AUXEVOL_GFS * Nxx_plus_2NGHOSTS_tot);
// For debugging, it can be useful to set everything to NaN initially.
if(poison_grids) {
for(int ii=0;ii<NUM_EVOL_GFS * Nxx_plus_2NGHOSTS_tot;ii++) {
y_n_gfs[ii] = 1.0/0.0;
y_nplus1_running_total_gfs[ii] = 1.0/0.0;
//k_odd_gfs[ii] = 1.0/0.0;
//k_even_gfs[ii] = 1.0/0.0;
diagnostic_output_gfs[ii] = 1.0/0.0;
evol_gfs_exact[ii] = 1.0/0.0;
}
for(int ii=0;ii<NUM_AUXEVOL_GFS * Nxx_plus_2NGHOSTS_tot;ii++) {
auxevol_gfs[ii] = 1.0/0.0;
auxevol_gfs_exact[ii] = 1.0/0.0;
}
}
// Step 0d: Set up coordinates: Set dx, and then dt based on dx_min and CFL condition
// This is probably already defined above, but just in case...
#ifndef MIN
#define MIN(A, B) ( ((A) < (B)) ? (A) : (B) )
#endif
REAL dt = CFL_FACTOR * MIN(dxx0,MIN(dxx1,dxx2)); // CFL condition
int Nt = (int)(t_final / dt + 0.5); // The number of points in time.
//Add 0.5 to account for C rounding down integers.
// Step 0e: Set up cell-centered Cartesian coordinate grids
REAL *xx[3];
xx[0] = (REAL *)malloc(sizeof(REAL)*Nxx_plus_2NGHOSTS0);
xx[1] = (REAL *)malloc(sizeof(REAL)*Nxx_plus_2NGHOSTS1);
xx[2] = (REAL *)malloc(sizeof(REAL)*Nxx_plus_2NGHOSTS2);
for(int j=0;j<Nxx_plus_2NGHOSTS0;j++) xx[0][j] = xxmin[0] + (j-NGHOSTS+1)*dxx0;
for(int j=0;j<Nxx_plus_2NGHOSTS1;j++) xx[1][j] = xxmin[1] + (j-NGHOSTS+1)*dxx1;
for(int j=0;j<Nxx_plus_2NGHOSTS2;j++) xx[2][j] = xxmin[2] + (j-NGHOSTS+1)*dxx2;
// Step 1: Set up initial data to be exact solution at time=0:
REAL time = 0.0;
set_initial_spacetime_metric_data(¶ms,xx,auxevol_gfs);
initial_data(¶ms,xx,auxevol_gfs,y_n_gfs);
// Fill in the remaining quantities
apply_bcs_potential(¶ms,y_n_gfs);
driver_A_to_B(¶ms,y_n_gfs,auxevol_gfs);
//override_BU_with_old_GiRaFFE(¶ms,auxevol_gfs,0);
GiRaFFE_NRPy_prims_to_cons(¶ms,auxevol_gfs,y_n_gfs);
apply_bcs_velocity(¶ms,auxevol_gfs);
// Extra stack, useful for debugging:
GiRaFFE_NRPy_cons_to_prims(¶ms,xx,auxevol_gfs,y_n_gfs);
//GiRaFFE_NRPy_prims_to_cons(¶ms,auxevol_gfs,y_n_gfs);
//GiRaFFE_NRPy_cons_to_prims(¶ms,xx,auxevol_gfs,y_n_gfs);
//GiRaFFE_NRPy_prims_to_cons(¶ms,auxevol_gfs,y_n_gfs);
//GiRaFFE_NRPy_cons_to_prims(¶ms,xx,auxevol_gfs,y_n_gfs);
for(int n=0;n<=Nt;n++) { // Main loop to progress forward in time.
//for(int n=0;n<=1;n++) { // Main loop to progress forward in time.
// Step 1a: Set current time to correct value & compute exact solution
time = ((REAL)n)*dt;
/* Step 2: Validation: Output relative error between numerical and exact solution, */
if((n)%NSKIP_1D_OUTPUT ==0) {
// Step 2c: Output relative error between exact & numerical at center of grid.
const int i0mid=Nxx_plus_2NGHOSTS0/2;
const int i1mid=Nxx_plus_2NGHOSTS1/2;
const int i2mid=Nxx_plus_2NGHOSTS2/2;
char filename[100];
sprintf(filename,"out%d-%08d.txt",Nxx0,n);
FILE *out2D = fopen(filename, "w");
for(int i0=0;i0<Nxx_plus_2NGHOSTS0;i0++) {
const int idx = IDX3S(i0,i1mid,i2mid);
fprintf(out2D,"%.16e %.16e %.16e %.16e %.16e %.16e %.16e %.16e %.16e %.16e %.16e %.16e %.16e %.16e\n",
xx[0][i0],
auxevol_gfs[IDX4ptS(BU0GF,idx)],auxevol_gfs[IDX4ptS(BU1GF,idx)],auxevol_gfs[IDX4ptS(BU2GF,idx)],
y_n_gfs[IDX4ptS(AD0GF,idx)],y_n_gfs[IDX4ptS(AD1GF,idx)],y_n_gfs[IDX4ptS(AD2GF,idx)],
y_n_gfs[IDX4ptS(STILDED0GF,idx)],y_n_gfs[IDX4ptS(STILDED1GF,idx)],y_n_gfs[IDX4ptS(STILDED2GF,idx)],
auxevol_gfs[IDX4ptS(VALENCIAVU0GF,idx)],auxevol_gfs[IDX4ptS(VALENCIAVU1GF,idx)],auxevol_gfs[IDX4ptS(VALENCIAVU2GF,idx)],
y_n_gfs[IDX4ptS(PSI6PHIGF,idx)]);
}
fclose(out2D);
//set_initial_spacetime_metric_data(¶ms,xx,auxevol_gfs_exact);
//initial_data(¶ms,xx,auxevol_gfs_exact,evol_gfs_exact);
// Fill in the remaining quantities
//driver_A_to_B(¶ms,evol_gfs_exact,auxevol_gfs_exact);
//GiRaFFE_NRPy_prims_to_cons(¶ms,auxevol_gfs_exact,evol_gfs_exact);
sprintf(filename,"out%d-%08d_exact.txt",Nxx0,n);
FILE *out2D_exact = fopen(filename, "w");
for(int i0=0;i0<Nxx_plus_2NGHOSTS0;i0++) {
const int idx = IDX3S(i0,i1mid,i2mid);
fprintf(out2D_exact,"%.16e %.16e %.16e %.16e %.16e %.16e %.16e %.16e %.16e %.16e %.16e %.16e %.16e %.16e\n",
xx[0][i0],
auxevol_gfs_exact[IDX4ptS(BU0GF,idx)],auxevol_gfs_exact[IDX4ptS(BU1GF,idx)],auxevol_gfs_exact[IDX4ptS(BU2GF,idx)],
evol_gfs_exact[IDX4ptS(AD0GF,idx)],evol_gfs_exact[IDX4ptS(AD1GF,idx)],evol_gfs_exact[IDX4ptS(AD2GF,idx)],
evol_gfs_exact[IDX4ptS(STILDED0GF,idx)],evol_gfs_exact[IDX4ptS(STILDED1GF,idx)],evol_gfs_exact[IDX4ptS(STILDED2GF,idx)],
auxevol_gfs_exact[IDX4ptS(VALENCIAVU0GF,idx)],auxevol_gfs_exact[IDX4ptS(VALENCIAVU1GF,idx)],auxevol_gfs_exact[IDX4ptS(VALENCIAVU2GF,idx)],
evol_gfs_exact[IDX4ptS(PSI6PHIGF,idx)]);
}
fclose(out2D_exact);
}
// Step 3: Evolve scalar wave initial data forward in time using Method of Lines with RK4 algorithm,
// applying quadratic extrapolation outer boundary conditions.
// Step 3.b: Step forward one timestep (t -> t+dt) in time using
// chosen RK-like MoL timestepping algorithm
#include "MoLtimestepping/RK_MoL.h"
} // End main loop to progress forward in time.
// Step 4: Free all allocated memory
#include "MoLtimestepping/RK_Free_Memory.h"
free(auxevol_gfs);
free(auxevol_gfs_exact);
free(evol_gfs_exact);
for(int i=0;i<3;i++) free(xx[i]);
return 0;
}
# +
cmd.C_compile(os.path.join(Ccodesdir,"GiRaFFE_NRPy_standalone.c"),
os.path.join(Ccodesdir,"output","GiRaFFE_NRPy_standalone"),compile_mode="safe")
# # !gcc -g -O2 -fopenmp GiRaFFE_standalone_Ccodes/GiRaFFE_NRPy_standalone.c -o GiRaFFE_NRPy_standalone -lm
# Change to output directory
os.chdir(outdir)
# Clean up existing output files
cmd.delete_existing_files("out*.txt")
cmd.delete_existing_files("out*.png")
# cmd.Execute(os.path.join(Ccodesdir,"output","GiRaFFE_NRPy_standalone"), "640 16 16", os.path.join(outdir,"out640.txt"))
cmd.Execute("GiRaFFE_NRPy_standalone", "119 119 119","out119.txt")
# cmd.Execute("GiRaFFE_NRPy_standalone", "239 15 15","out239.txt")
# # !OMP_NUM_THREADS=1 valgrind --track-origins=yes -v ./GiRaFFE_NRPy_standalone 1280 32 32
# Return to root directory
os.chdir(os.path.join("../../"))
# -
# Now, we will load the data generated by the simulation and plot it in order to test for convergence.
# +
import numpy as np
import matplotlib.pyplot as plt
Data_numer = np.loadtxt(os.path.join("GiRaFFE_standalone_Ccodes","output","out119-00000040.txt"))
# Data_num_2 = np.loadtxt(os.path.join("GiRaFFE_standalone_Ccodes","output","out239-00000080.txt"))
# Data_old = np.loadtxt("/home/penelson/OldCactus/Cactus/exe/ABE-GiRaFFEfood_1D_AlfvenWave/giraffe-grmhd_primitives_bi.x.asc")
# Data_o_2 = np.loadtxt("/home/penelson/OldCactus/Cactus/exe/ABE-GiRaFFEfood_1D_AlfvenWave_2/giraffe-grmhd_primitives_bi.x.asc")
# Data_numer = Data_old[5000:5125,11:15] # The column range is chosen for compatibility with the plotting script.
# Data_num_2 = Data_o_2[19600:19845,11:15] # The column range is chosen for compatibility with the plotting script.
Data_exact = np.loadtxt(os.path.join("GiRaFFE_standalone_Ccodes","output","out119-00000040_exact.txt"))
# Data_exa_2 = np.loadtxt(os.path.join("GiRaFFE_standalone_Ccodes","output","out239-00000080_exact.txt"))
predicted_order = 2.0
column = 3
plt.figure()
# # plt.plot(Data_exact[2:-2,0],np.log2(np.absolute((Data_numer[2:-2,column]-Data_exact[2:-2,column])/\
# # (Data_num_2[2:-2:2,column]-Data_exa_2[2:-2:2,column]))),'.')
plt.plot(Data_exact[:,0],Data_exact[:,column])
plt.plot(Data_exact[:,0],Data_numer[:,column],'.')
# plt.xlim(-0.0,1.0)
# # plt.ylim(-1.0,5.0)
# # plt.ylim(-0.0005,0.0005)
# plt.xlabel("x")
# plt.ylabel("BU2")
plt.show()
# # 0 1 2 3 4 5 6 7 8 9 10 11 12 13
# labels = ["x","BU0","BU1","BU2","AD0","AD1","AD2","StildeD0","StildeD1","StildeD2","ValenciavU0","ValenciavU1","ValenciavU2", "psi6Phi"]
# old_files = ["",
# "giraffe-grmhd_primitives_bi.x.asc","giraffe-grmhd_primitives_bi.x.asc","giraffe-grmhd_primitives_bi.x.asc",
# # "giraffe-em_ax.x.asc","giraffe-em_ay.x.asc","giraffe-em_az.x.asc",
# "cell_centered_Ai.txt","cell_centered_Ai.txt","cell_centered_Ai.txt",
# "giraffe-grmhd_conservatives.x.asc","giraffe-grmhd_conservatives.x.asc","giraffe-grmhd_conservatives.x.asc",
# "giraffe-grmhd_primitives_allbutbi.x.asc","giraffe-grmhd_primitives_allbutbi.x.asc","giraffe-grmhd_primitives_allbutbi.x.asc",
# "giraffe-em_psi6phi.x.asc"]
# column = 5
# column_old = [0,12,13,14,0,1,2,12,13,14,12,13,14,12]
# old_path = "/home/penelson/OldCactus/Cactus/exe/ABE-GiRaFFEfood_1D_AlfvenWave"
# new_path = os.path.join("GiRaFFE_standalone_Ccodes","output")
# data_old = np.loadtxt(os.path.join(old_path,old_files[column]))
# # data_old = data_old[250:375,:]# Select only the second timestep
# # data_old = data_old[125:250,:]# Select only the first timestep
# # data_old = data_old[0:125,:]# Select only the zeroth timestep
# data_new = np.loadtxt(os.path.join(new_path,"out119-00000001.txt"))
# deltaA_old = data_old[125:250,:] - data_old[0:125,:]
# data_new_t0 = np.loadtxt(os.path.join(new_path,"out119-00000000.txt"))
# deltaA_new = data_new[:,:] - data_new_t0[:,:]
# plt.figure()
# # plt.plot(data_new[3:-3,0],data_new[3:-3,column]-data_old[3:-3,column_old[column]])
# # plt.plot(data_new[:,0],data_new[:,column]-((3*np.sin(5*np.pi*data_new[:,0]/np.sqrt(1 - (-0.5)**2))/20 + 23/20)*(data_new[:,0]/2 + np.sqrt(1 - (-0.5)**2)/20 + np.absolute(data_new[:,0] + np.sqrt(1 - (-0.5)**2)/10)/2)*(-1e-100/2 + data_new[:,0]/2 - np.sqrt(1 - (-0.5)**2)/20 - np.absolute(-1e-100 + data_new[:,0] - np.sqrt(1 - (-0.5)**2)/10)/2)/((-1e-100 + data_new[:,0] - np.sqrt(1 - (-0.5)**2)/10)*(1e-100 + data_new[:,0] + np.sqrt(1 - (-0.5)**2)/10)) + 13*(data_new[:,0]/2 - np.sqrt(1 - (-0.5)**2)/20 + np.absolute(data_new[:,0] - np.sqrt(1 - (-0.5)**2)/10)/2)/(10*(1e-100 + data_new[:,0] - np.sqrt(1 - (-0.5)**2)/10)) + (-1e-100/2 + data_new[:,0]/2 + np.sqrt(1 - (-0.5)**2)/20 - np.absolute(-1e-100 + data_new[:,0] + np.sqrt(1 - (-0.5)**2)/10)/2)/(-1e-100 + data_new[:,0] + np.sqrt(1 - (-0.5)**2)/10))/np.sqrt(1 - (-0.5)**2))
# # plt.plot(data_new[1:,0]-(data_new[0,0]-data_new[1,0])/2.0,(data_new[0:-1,column]+data_new[1:,column])/2,'.',label="GiRaFFE_NRPy+injected BU")
# # plt.plot(data_new[1:,0]-(data_new[0,0]-data_new[1,0])/2.0,data_old[1:,column_old[column]],label="old GiRaFFE")
# # -(data_old[0,9]-data_old[1,9])/2.0
# # plt.plot(data_new[3:-3,0],deltaA_new[3:-3,column],'.')
# plt.plot(data_new[3:-3,0],deltaA_old[3:-3,column_old[column]]-deltaA_new[3:-3,column])
# # plt.xlim(-0.1,0.1)
# # plt.ylim(-0.2,0.2)
# plt.legend()
# plt.xlabel(labels[0])
# plt.ylabel(labels[column])
# plt.show()
# # print(np.argmin(deltaA_old[3:-3,column_old[column]]-deltaA_new[3:-3,column]))
# -
# This code will create an animation of the wave over time.
# +
# import matplotlib.pyplot as plt
from matplotlib.pyplot import savefig
from IPython.display import HTML
import matplotlib.image as mgimg
import glob
import sys
from matplotlib import animation
cmd.delete_existing_files("out119-00*.png")
globby = glob.glob(os.path.join('GiRaFFE_standalone_Ccodes','output','out119-00*.txt'))
file_list = []
for x in sorted(globby):
file_list.append(x)
number_of_files = int(len(file_list)/2)
for timestep in range(number_of_files):
fig = plt.figure()
numer_filename = file_list[2*timestep]
exact_filename = file_list[2*timestep+1]
Numer = np.loadtxt(numer_filename)
Exact = np.loadtxt(exact_filename)
plt.title("Alfven Wave")
plt.xlabel("x")
plt.ylabel("BU2")
plt.xlim(-0.5,0.5)
plt.ylim(1.0,1.7)
plt.plot(Numer[3:-3,0],Numer[3:-3,3],'.',label="Numerical")
plt.plot(Exact[3:-3,0],Exact[3:-3,3],label="Exact")
plt.legend()
savefig(numer_filename+".png",dpi=150)
plt.close(fig)
sys.stdout.write("%c[2K" % 27)
sys.stdout.write("Processing file "+numer_filename+"\r")
sys.stdout.flush()
# +
## VISUALIZATION ANIMATION, PART 2: Combine PNGs to generate movie ##
# https://stackoverflow.com/questions/14908576/how-to-remove-frame-from-matplotlib-pyplot-figure-vs-matplotlib-figure-frame
# https://stackoverflow.com/questions/23176161/animating-pngs-in-matplotlib-using-artistanimation
# # !rm -f GiRaFFE_NRPy-1D_tests.mp4
cmd.delete_existing_files("GiRaFFE_NRPy-1D_tests.mp4")
fig = plt.figure(frameon=False)
ax = fig.add_axes([0, 0, 1, 1])
ax.axis('off')
myimages = []
for i in range(number_of_files):
img = mgimg.imread(file_list[2*i]+".png")
imgplot = plt.imshow(img)
myimages.append([imgplot])
ani = animation.ArtistAnimation(fig, myimages, interval=100, repeat_delay=1000)
plt.close()
ani.save('GiRaFFE_NRPy-1D_tests.mp4', fps=5,dpi=150)
# -
# %%HTML
<video width="480" height="360" controls>
<source src="GiRaFFE_NRPy-1D_tests.mp4" type="video/mp4">
</video>
import cmdline_helper as cmd # NRPy+: Multi-platform Python command-line interface
cmd.output_Jupyter_notebook_to_LaTeXed_PDF("Tutorial-GiRaFFE_NRPy_Main_Driver",location_of_template_file=os.path.join(".."))
|
in_progress/Tutorial-Start_to_Finish-GiRaFFE_NRPy-3D_tests-unstaggered.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# 
#
#
# <h2 align='center'>Data Literacy through Sports Analytics</h2>
# <h3 align='center'>Southern Alberta Teachers' Convention 2021</h3>
#
# <h3 align='center'><NAME> (Cybera)<br>
# <NAME> (University of Calgary)</h3><br>
#
# <h4 align='center'> Slides at: https://tinyurl.com/pyryclxf </h4>
#
# 
# + [markdown] slideshow={"slide_type": "skip"}
# <center><img src='./images/ccby.png' alt="CC BY logo" width='300' /></center>
#
# <p><center><a href='https://creativecommons.org/licenses/by/4.0/'>CC BY</a>:<br>
# This license allows reusers to distribute, remix, adapt, and build upon the material in any medium or format,<br>
# so long as attribution is given to the creator.
# </center></p>
# + slideshow={"slide_type": "skip"}
import numpy as np
import pandas as pd
from pandas import read_csv
import plotly.graph_objects as go
import plotly.express as px
from plotly import subplots
from plotly.offline import download_plotlyjs, plot,iplot
import cufflinks as cf
cf.go_offline()
from IPython.display import YouTubeVideo
from ipysheet import sheet, cell, cell_range
# %matplotlib inline
# + [markdown] slideshow={"slide_type": "slide"}
# # Overview
#
# - Data literacy via sports
# - The learning progression
# - Examples of learning and data analysis
# - Professional soccer
# - Ice hockey
# - Field hockey
# - Python, Jupyter, and Callysto
# + [markdown] slideshow={"slide_type": "slide"}
# <center><img src='./images/data_literacy.png' alt='data literacy' width='85%' /></center>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Example: professional soccer event data
# Data source: https://github.com/metrica-sports/sample-data
# + slideshow={"slide_type": "-"}
df_soccer = pd.read_csv("https://raw.githubusercontent.com/metrica-sports/sample-data/master/data/Sample_Game_1/Sample_Game_1_RawEventsData.csv"); df_soccer
# + [markdown] slideshow={"slide_type": "slide"}
# **Home team passes, second half**
# + slideshow={"slide_type": "-"}
df_soccer.loc[lambda df: (df['Team'] == 'Home') & (df['Period'] == 2) & (df['Type'] == 'PASS'), :] \
.iplot(kind="scatter",x = "Start X", y = "Start Y", mode = "markers")
# + [markdown] slideshow={"slide_type": "slide"}
# ## Bridging expert to novice
# + [markdown] slideshow={"slide_type": "slide"}
# ## Data visualization learning progression
# <img src='./images/creating_scaffolding.png' alt='scaffolding' width='95%' />
# + [markdown] slideshow={"slide_type": "slide"}
# ## Data visualization learning progression
# <img src='./images/creating_adapting.png' alt='adapting' width='95%' />
# + [markdown] slideshow={"slide_type": "slide"}
# ## Data gathering learning progression
# <br>
# <center><img src='./images/data_gathering.png' alt='data gathering' width='85%' /></center>
# <br><br><br>Source: <a href='http://oceansofdata.org/sites/oceansofdata.org/files/pervasive-and-persistent-understandings-01-14.pdf'>Pervasive and Persistent Understandings about Data</a>, <br>Kastens (2014)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Authentic learning approach
#
# - Learning design based on interdisciplinary<br>
# connections and real-world examples
# - Industry-aligned data science analysis process
# - Python, an all-purpose programming language
# - Jupyter notebook, a free industry-standard tool for data scientists
# - CallystoHub, free cloud computing
# + [markdown] slideshow={"slide_type": "slide"}
# ## Athlete development
#
# ### U15 training to train
# - Promotes tactical strategies for in-game decision making, reading the situation and inferring
# - Focuses on the team and the process
# - Situates personal goals within a team approach
#
# ### U18 training to compete
# - Emphasizes individual technical and position-specific training
# + [markdown] slideshow={"slide_type": "slide"}
# ## Youth sports analytics
#
# Online communication,<br>
# sometimes through shared video analysis spaces
#
# Video replay during games and training
#
# Post–game video analysis, limitted statistics
# + [markdown] slideshow={"slide_type": "slide"}
# ## Learning design and flexibility
# <br>
# <img src='./images/flexibility.png' alt='adapting' width='90%' />
# + [markdown] slideshow={"slide_type": "slide"}
# ## Two data examples
#
# 1. Import a csv file and use a Python spreadsheet<br>to create shot maps (ice hockey)
# 2. Gather data from video to analyze and make decisions (field hockey)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Data example 1:
# ## Using IIHF World Junior Championship data to create graphs and a shot map
# + [markdown] slideshow={"slide_type": "slide"}
# ## Defining ice hockey net zones:<br> What factors can lead to scoring?
# <!--USA Hockey Goaltender Basics https://www.usahockeygoaltending.com/page/show/890039-stance-->
# ||
# |-|-|
# |<img src='./images/hockey_net_zones.png' width='100%'/>|<img src='https://cdn.hockeycanada.ca/hockey-canada/Team-Canada/Men/Under-18/2014-15/2014-15_goalie_camp.jpg?q=60' />|
# ||<a href='https://www.hockeycanada.ca/en-ca/news/34-goaltenders-invited-to-2014-poe-camp'>Image source: Hockey Canada</a>|
# + slideshow={"slide_type": "slide"} language="html"
# <h2>Data source IIHF: Shot charts</h2><br>
# <iframe width="1200" height="600" src="https://www.iihf.com/pdf/503/ihm503a13_77a_3_0" frameborder="0" ></iframe>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Tally chart
# <img src='./images/hockey_tally.png' alt='tally chart' width='85%' />
# + [markdown] slideshow={"slide_type": "slide"}
# ## Generating a csv file
#
# Zone,Austria,Canada,Czech_Republic,Finland,Germany,Russia,Switzerland,Slovakia,Sweden,USA,Total<br>
# one,0,7,0,3,2,0,0,0,3,3,18<br>
# two,0,1,1,0,1,0,0,0,0,0,3<br>
# three,0,5,0,2,2,4,1,0,3,6,23<br>
# four,0,4,3,2,1,1,0,1,0,3,15<br>
# five,0,1,0,2,1,0,0,0,0,0,4<br>
# six,1,1,2,4,0,2,0,1,0,2,13<br>
# seven,0,6,0,1,3,3,1,1,0,9,24<br>
# eight,0,5,1,2,2,3,1,2,3,2,21<br>
# nine,0,3,3,0,2,3,2,0,5,0,18<br>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Exploring scoring on net zones
# + slideshow={"slide_type": "-"}
hockey_goals_df = pd.read_csv('./data/hockey_goals.csv')
hockey_goals_df.head(9)
# + [markdown] slideshow={"slide_type": "slide"}
# ### What do measures of central tendency<br>tell us about the total goals per net zone?
# + slideshow={"slide_type": "-"}
hockey_goals_df['Total'].sum()
# + slideshow={"slide_type": "-"}
hockey_goals_df['Total'].min()
# + slideshow={"slide_type": "-"}
hockey_goals_df['Total'].max()
# + slideshow={"slide_type": "slide"}
scatter_hockey_goals_df = px.scatter(hockey_goals_df,x="Zone",y="Total",title="Total goals per net zone")
scatter_hockey_goals_df.show()
# + slideshow={"slide_type": "slide"}
hockey_goals_df['Total'].mean()
# + slideshow={"slide_type": "-"}
hockey_goals_df['Total'].median()
# + slideshow={"slide_type": "-"}
hockey_goals_df['Total'].mode()
# + [markdown] slideshow={"slide_type": "slide"}
# ### Which net zones score above the median?
# + slideshow={"slide_type": "-"}
hockey_goals_df = hockey_goals_df.sort_values('Total', ascending=False)
hockey_goals_df
# + slideshow={"slide_type": "slide"}
bar_hockey_goals_df = px.bar(hockey_goals_df,
x="Zone", y="Total", title="Goals by net zone")
bar_hockey_goals_df.update_layout(title_text='Total goals by net zone')
# + [markdown] slideshow={"slide_type": "slide"}
# ### What connections exist between<br>goalie position and scoring?
# + slideshow={"slide_type": "skip"}
hockey_goals_df = pd.read_csv('./data/hockey_goals.csv')
hockey_goals_df.Total
# + slideshow={"slide_type": "skip"}
spread_sheet_hockey_net = sheet(rows=3, columns=3)
my_cells_net = cell_range([[18,3,23],[15,4,13],[24,21,18]],row_start=0,col_start=0,numeric_format="int")
figure_hockey_net = go.Figure(data=go.Heatmap(
z =list(reversed(my_cells_net.value)),
type = 'heatmap',
colorscale = 'greys',opacity = 1.0))
axis_template = dict(range = [0,5], autorange = True,
showgrid = False, zeroline = False,
showticklabels = False,
ticks = '' )
figure_hockey_net.update_layout(margin = dict(t=50,r=200,b=200,l=200),
xaxis = axis_template,
yaxis = axis_template,
showlegend = False,
width = 800, height = 500, title="Ice hockey net zones",
autosize = True )
# Add image in the background
nLanes = 3
nZones = 3
figure_hockey_net.add_layout_image(
dict(
source="images/hockey_net.png",
xref="x",
yref="y",
x=-0.5,
y=-.5 + nLanes, #this adjusts the placement of the image
sizex=nZones,
sizey=nLanes,
sizing="fill",
opacity=1.0,
layer="above")
)
# changes in my_cells should trigger this function
def calculate(change):
figure_hockey_net.update_traces(z=list(reversed(my_cells_net.value)))
my_cells_net.observe(calculate, 'value')
# + slideshow={"slide_type": "-"}
spread_sheet_hockey_net
# + slideshow={"slide_type": "-"}
139
# + slideshow={"slide_type": "-"}
figure_hockey_net.update() # Click the keys "Shift-return" to update the figure
# + [markdown] slideshow={"slide_type": "slide"}
# ## Data example 2:
#
# ## Analyzing youth field hockey data to make decisions
# + [markdown] slideshow={"slide_type": "slide"}
# <center><img src='./images/learning_cycle1.png' alt="Learning design and context" width='90%' /></center>
# + [markdown] slideshow={"slide_type": "slide"}
# <center><img src='./images/learning_cycle5.png' alt="Learning cycle" width='90%' /></center>
# + [markdown] slideshow={"slide_type": "slide"}
# <center><img src='./images/computational_thinking.png' alt="Computationl thinking" width='90%' /></center>
# + [markdown] slideshow={"slide_type": "slide"}
# <center><img src='./images/analysis_process.png' alt="Data science analysis process" width='90%' /></center>
# + [markdown] slideshow={"slide_type": "slide"}
# <center><img src='./images/collective.png' alt="Collective decision making" width='90%' /></center>
# + [markdown] slideshow={"slide_type": "slide"}
# <center><img src='./images/field_hockey_game.png' alt="Field hockey" width='90%' /></center>
# + [markdown] slideshow={"slide_type": "slide"}
# <center><img src='./images/understand1.png' alt="Understand actions" width='90%' /></center>
# + [markdown] slideshow={"slide_type": "slide"}
# <center><img src='./images/actions.png' alt="Understand viewpoints" width='90%' /></center>
# + slideshow={"slide_type": "slide"}
print ('Passes received')
YouTubeVideo('mIwiiJO7Rk4?start=2893&end=2915', width='600', height='355')
# + [markdown] slideshow={"slide_type": "slide"}
# <center><img src='./images/gather4.png' alt="Gather" width='90%' /></center>
# + [markdown] slideshow={"slide_type": "slide"}
# <center><img src='./images/collection_passing.png' alt="Passing" width='90%' /></center>
# + [markdown] slideshow={"slide_type": "slide"}
# ## 3. Organize
# + slideshow={"slide_type": "-"}
possession_time_df = read_csv('data/field_hockey_possession_time.csv')
possession_time_df.head(8)
# + [markdown] slideshow={"slide_type": "slide"}
# ## 4. Explore:
# How does ball possession affect outcomes?
# + [markdown] slideshow={"slide_type": "skip"}
# Home 2 shots on net (Q3); Away 1 shot on net (Q1)
# + slideshow={"slide_type": "-"}
bar_possession_time_df = px.bar(possession_time_df,x="Possession Time (seconds)",y="Quarter",title="Possession per quarter",color="Team")
bar_possession_time_df.update_layout(autosize=False, width=600, height=400)
# + slideshow={"slide_type": "skip"}
lanes_home_passes_df = read_csv('data/field_hockey_lanes_home_passes.csv')
lanes_home_passes_df.head()
# + slideshow={"slide_type": "slide"}
lanes_home_passes_df.iplot(kind='pie',values="Count",labels="Action",
title="Passes received, intercepted, and missed for Home team")
# + [markdown] slideshow={"slide_type": "slide"}
# ## 4. Explore passes received:
# What stays the same and what changes?
# + slideshow={"slide_type": "skip"}
df_passes_home = pd.read_csv('data/field_hockey_home_passes.csv'); df_passes_home
# + slideshow={"slide_type": "skip"}
df_temp_1 = df_passes_home.loc[lambda df: (df['Phase of Play'] == 'attack') &(df['Quarter'] == 'first') ];
df_temp_2 = df_passes_home.loc[lambda df: (df['Phase of Play'] == 'attack') &(df['Quarter'] == 'second') ];
df_temp_3 = df_passes_home.loc[lambda df: (df['Phase of Play'] == 'attack') &(df['Quarter'] == 'third') ];
df_temp_4 = df_passes_home.loc[lambda df: (df['Phase of Play'] == 'attack') &(df['Quarter'] == 'fourth') ];
# + slideshow={"slide_type": "skip"}
#import plotly.tools as tls
fig_all = subplots.make_subplots(rows=1, cols=4)
fig_1 = df_temp_1.iplot(kind='heatmap', colorscale='blues', x='Lane', y='Zone', z='Count' , asFigure=True)
fig_2 = df_temp_2.iplot(kind='heatmap', colorscale='blues', x='Lane', y='Zone', z='Count' , asFigure=True)
fig_3 = df_temp_3.iplot(kind='heatmap', colorscale='blues', x='Lane', y='Zone', z='Count' , asFigure=True)
fig_4 = df_temp_4.iplot(kind='heatmap', colorscale='blues', x='Lane', y='Zone', z='Count' , asFigure=True)
fig_all.append_trace(fig_1['data'][0], 1, 1)
fig_all.append_trace(fig_2['data'][0], 1, 2)
fig_all.append_trace(fig_3['data'][0], 1, 3)
fig_all.append_trace(fig_4['data'][0], 1, 4)
# + slideshow={"slide_type": "-"}
fig_all.update_xaxes(showticklabels = False, linecolor='black')
fig_all.update_yaxes(showticklabels = False, linecolor='black')
iplot(fig_all)
# + slideshow={"slide_type": "skip"}
df_passes_home.loc[lambda df: (df['Lane'] == 1) &(df['Phase of Play'] == 'attack') &(df['Quarter'].isin(['first', 'second'])) ].sum()
# + slideshow={"slide_type": "skip"}
df_passes_home.loc[lambda df: (df['Lane'] == 1) &(df['Phase of Play'] == 'attack') &(df['Quarter']== 'first') ].sum()
# + slideshow={"slide_type": "skip"}
df_passes_home.loc[lambda df: (df['Lane'] == 1) &(df['Phase of Play'] == 'attack') &(df['Quarter']== 'second') ].sum()
# + slideshow={"slide_type": "skip"}
df_passes_home.loc[lambda df: (df['Lane'] == 5) &(df['Phase of Play'] == 'attack') &(df['Quarter'].isin(['third', 'fourth'])) ].sum()
# + slideshow={"slide_type": "skip"}
df_passes_home.loc[lambda df: (df['Lane'] == 5) &(df['Phase of Play'] == 'attack') &(df['Quarter']== 'third') ].sum()
# + slideshow={"slide_type": "skip"}
df_passes_home.loc[lambda df: (df['Lane'] == 5) &(df['Phase of Play'] == 'attack') &(df['Quarter']== 'fourth') ].sum()
# + [markdown] slideshow={"slide_type": "-"}
# - The most passes in a zone occur toward the left outside lane of the opponent's net across quarters: first quarter, 14/49 (29%); second quarter, 13/32 (41%); thrid quarter, 16/42 (38%); fourth quarter, 8/29 (28%).
# - In the second half, defence keeps passes out of the zone of the net. 0 passes occur in that top horizontal zone.
# + slideshow={"slide_type": "skip"}
8/29
# + [markdown] slideshow={"slide_type": "slide"}
# ## 5. Interpret<br> How can the data exploration inform decision making?
#
# > - Keeping ball possession by carrying the ball
# > - Dominating play by keeping the ball out of the zone near the net
# > - Attacking on the outer lanes, especially toward the left side of the opponent's net
# + [markdown] slideshow={"slide_type": "slide"}
# # The technology in this talk
#
# - **Jupyter** notebooks, **Python** programming, **Pandas** for data
# - Free to teachers and students
# - **Callysto.ca** project (CanCode, Cybera, PIMS)
# - This slideshow **IS** a Jupyter notebook! (take a tour)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Callysto resources
#
# - <a href="https://www.callysto.ca/starter-kit/">Callysto starter kit</a> Getting started
# - <a href="https://courses.callysto.ca">courses.callysto.ca</a> Online courses
# - <a href="https://www.callysto.ca/weekly-data-visualization/">Weekly data visualizations</a> Quick activities
# - <a href="https://app.lucidchart.com/documents/view/8e3186f7-bdfe-46af-9c7f-9c426b80d083">Connecting data literacy and sports</a> About sports analytics
# + [markdown] slideshow={"slide_type": "slide"}
# <center><a href='https://www.callysto.ca/learning-modules/'><img src='./images/learning_modules.png' target='_blank' alt="Callysto learning modules" width='90%' /></a></center>
# <center>All free, all open source, aimed at teachers and students</center>
# + [markdown] slideshow={"slide_type": "slide"}
# <center><h3>Enter to win a $50 Amazon gift card by completing a Callysto feedback survey </h3></center>
# <center><h3>http://bit.ly/callysto-feedback</h3></center>
#
# <br>
# <p><center>Contact us at <a href="mailto:<EMAIL>"><EMAIL></a><br>
# for in-class workshops, virtual hackathons...<br>
# <a href="https://twitter.com/callysto_canada">@callysto_canada</a><br>
# <a href="https://callysto.ca">callysto.ca</a><br>
# <a href="https://www.youtube.com/channel/UCPdq1SYKA42EZBvUlNQUAng">YouTube</a>
# </center></p>
# + [markdown] slideshow={"slide_type": "slide"}
# <center><img src='./images/callysto_logo.png' alt="Callysto logo" width='80%' /></center>
# <center><img src='./images/callysto_partners2.png' alt='Callysto partners' width='80%' /></center>
# + [markdown] slideshow={"slide_type": "skip"}
# Alberta Education. (2017). *Career and Technology Foundations* [Program of Studies]. https://education.alberta.ca/media/3795641/ctf-program-of-studies-jan-4-2019.pdf
#
# Alberta Education. (2007, updated 2016). *Mathematics Kindergarten to Grade 9* [Program of Studies]. https://education.alberta.ca/media/3115252/2016_k_to_9_math_pos.pdf
#
# Alberta Education. (2000). *Physical Education* [Program of Studies]. https://education.alberta.ca/media/160191/phys2000.pdf
#
# BC Ministry of Education. (2020). *BC's Digital Literacy Framework*. https://www2.gov.bc.ca/assets/gov/education/kindergarten-to-grade-12/teach/teaching-tools/digital-literacy-framework.pdf
#
# <NAME>., & <NAME>. (2020). *Investigating “Problem-Solving With Datasets” as an Implementation of Computational Thinking: A Literature Review*. Journal of Educational Computing Research, 58(2), 502–534. https://doi.org/10.1177/0735633119845694
#
# <NAME>. (2006). *Theoretical and empirical differentiations of phases in the modeling process*. Zentralblatt fuer Didaktik der Mathematik, 38(2), 86-95. ZDM. 38. 86-95. 10.1007/BF02655883.
#
# <NAME>. & <NAME>. (1995). *Feedback and Self-Regulated Learning: A Theoretical Synthesis*. Review of Educational Research - REV EDUC RES. 65. 245-281. 10.2307/1170684.
#
# <NAME>. & <NAME>. (2001). *Self-Regulation Differences during Athletic Practice by Experts, Non-Experts, and Novices*. Journal of Applied Sport Psychology. 13. 185-206. 10.1080/104132001753149883.
#
# Conseil supérieur de l’éducation (2020). Éduquer au numérique, Rapport sur l’état et les besoins de
# l’éducation 2018-2020, Québec, Le Conseil, 96 p.
#
# Field Hockey Alberta. (2020). *Tactical Seminars*. http://www.fieldhockey.ab.ca/content/tactical-seminars
#
# Field Hockey Canada. (2020). *Ahead of the Game*. http://www.fieldhockey.ca/ahead-of-the-game-field-hockey-canada-webinar-series/
#
# <NAME>. (2020, September 2). *Shifting from Computational Thinking to Computational Modelling in Math Education* [Online plenary talk]. Changing the Culture 2020, Pacific Institute for the Mathematical Sciences.
#
# <NAME>. & <NAME>. (2019). *Assessment of Informal and Formal Inferential Reasoning: A Critical Research Review*. Statistics Education Research Journal, 18, 8-25. https://www.researchgate.net/publication/335057564_ASSESSMENT_OF_INFORMAL_AND_FORMAL_INFERENTIAL_REASONING_A_CRITICAL_RESEARCH_REVIEW
#
# <NAME>., <NAME>., & <NAME>. (2018). Self-regulation, co-regulation, and shared regulation in collaborative learning environments. In <NAME>, & <NAME>, (Eds.). *Handbook of self-regulation of learning and performance* (2nd ed.). New York: Routledge. https://www.researchgate.net/publication/313369294_Self-regulation_co-regulation_and_shared_regulation_in_collaborative_learning_environments
#
#
# <NAME>. (2014). *Pervasive and Persistent Understandings about Data*. Oceans of Data Institute. http://oceansofdata.org/sites/oceansofdata.org/files/pervasive-and-persistent-understandings-01-14.pdf
#
# <NAME>., & <NAME>. (2001, May). *Analyzing logfile data to produce Navigation Profiles of Studying as Self-regulated learning*. Paper presented at the annual meeting of the Canadian Society for the Study of Education, Quebec City, Quebec, Canada.
#
# Manitoba Education and Training (2020). *Literacy with ICT across the Curriculum: A Model for 21st Century Learning from K-12*. https://www.edu.gov.mb.ca/k12/tech/lict/index.html
#
# Ontario Ministry of Education. (2020). *The Ontario curriculum grades 1‐8: Mathematics* [Program of Studies]. https://www.dcp.edu.gov.on.ca/en/curriculum/elementary-mathematics
|
SATC-2021-sports/sports.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import os
os.chdir('..')
from app.ui.debug_view import DebugView
from app.overpass.location import Location
from app.tour_optimizer.poi import Poi
view = DebugView()
# +
a = Location(13.3587658, 52.4857809)
b = Location(13.412003, 52.522625)
view.display_path(a, b)
# +
neukoelln = Poi(0, 'Neukölln', Location(13.434469, 52.481374))
schlesi = Poi(1, 'Schlesi', Location(13.441840, 52.500931))
alex = Poi(2, 'Alex', Location(13.412937, 52.521811))
charlie = Poi(3, 'Charlie', Location(13.390779, 52.507484))
kotti = Poi(4, 'Kotti', Location(13.418210, 52.499224))
view.display_tour([neukoelln, schlesi, alex, charlie, kotti])
# -
|
notebooks/debug_view.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
from os.path import join, exists
from time import clock
datapath = '/cluster/work/grlab/clinical/Inselspital/DataReleases/01-19-2017/InselSpital/'
data_version = '180822'
# -
consent_patient = get_consent_patient_ids()
len(consent_patient)
# cd
# +
def get_datapath():
return datapath
def get_table_names(tbl_type='all'):
tbl_list = ['monvals', 'comprvals', 'dervals', 'observrec', 'labres', 'pharmarec']
if tbl_type == 'all':
return tbl_list
elif tbl_type == 'pharma':
return ['pharmarec']
elif tbl_type == 'non-pharma':
return tbl_list[:-1]
def get_status_str(statusID, tbl_name):
'''
Get the meaning of a status
'''
status_str = []
if tbl_name == 'pharmarec':
statusID_bin_str = '{0:10b}'.format(statusID)[::-1]
if statusID_bin_str[1] == '1':
status_str.append('invalidated')
if statusID_bin_str[2] == '1':
status_str.append('start')
if statusID_bin_str[3] == '1':
status_str.append('caused by event')
if statusID_bin_str[5] == '1':
status_str.append('notified')
if statusID_bin_str[8] == '1':
status_str.append('stop')
if statusID_bin_str[9] == '1':
status_str.append('included in record reports')
else:
statusID_bin_str = '{0:11b}'.format(statusID)[::-1]
if statusID_bin_str[0] == '1':
status_str.append('out of range')
if statusID_bin_str[1] == '1':
status_str.append('invalidated')
if statusID_bin_str[2] == '1':
status_str.append('first of connection')
if statusID_bin_str[3] == '1':
status_str.append('caused by event')
if statusID_bin_str[4] == '1':
status_str.append('compressed')
if statusID_bin_str[5] == '1':
status_str.append('notified but not measured')
if statusID_bin_str[6] == '1':
status_str.append('bigger than')
if statusID_bin_str[7] == '1':
status_str.append('smaller than')
if statusID_bin_str[10] == '1':
status_str.append('mandatory')
return ', '.join(status_str)
def voi_id_name_mapping(tbl_name, replace_name=False, include_all=False, use_ref='excel', version='v6'):
"""
Load the mapping between variableID and variableName
Parameters:
tbl_name: the name of the table (string)
replace_name: bool; if True, the variableNames are the medical terms; otherwise, the varialbeName
are the 'v%s'%variableIDs.
include_all: bool; if True, all the variables in the table are variables of interest; otherwise,
only variables can be found in the ref_excel files are the variables of interest.
Returns:
voi: dataframe; include the mapping between variableIDs and variableNames of the variables of
interest
"""
if use_ref not in ['excel', 'expot']:
raise Exception('use_ref can only be "excel" or "expot"')
id_list_path = join(datapath, 'misc_derived', 'id_lists')
excelref_path = join(datapath, 'misc_derived', 'ref_excel')
expotref_path = join(datapath, '0_csv_exports')
vid_set = pd.read_csv(join(id_list_path, 'vID_%s.csv'%tbl_name), sep=',',
dtype={'VariableID': int}).VariableID.unique()
# The head of the variable name is 'v' is the variable is non-pharma variable;
# otherwise the variable name head is 'p'
vname_head = 'p' if tbl_name == 'pharmarec' else 'v'
if include_all and not replace_name:
ref_voi = pd.DataFrame([[x, '%s%d'%(vname_head, x)] for x in vid_set],
columns=['VariableID', 'VariableName'])
else:
excelref_filename = 'labref_excel_%s.tsv'%version if tbl_name == 'labres' else 'varref_excel_%s.tsv'%version
excelref = pd.read_csv(join(excelref_path, excelref_filename), sep='\t', encoding='cp1252')
# Avoid repeat VariableIDs in 'pharmarec' and other tables
if tbl_name == 'pharmarec':
excelref = excelref[excelref.Type=='Pharma']
elif tbl_name != 'labres':
excelref = excelref[excelref.Type!='Pharma']
try:
excelref['VariableID'] = excelref['VariableID'].astype(int)
except:
excelref['VariableID'] = excelref.VariableID.apply(lambda x: float('NaN') if x=='???' else float(x))
if not include_all and not replace_name:
vid_voi = set(excelref.VariableID) & set(vid_set)
ref_voi = pd.DataFrame([[x, '%s%d'%(vname_head, x)] for x in vid_voi],
columns=['VariableID', 'VariableName'])
else:
if tbl_name == 'pharmarec':
expotref = pd.read_csv(join(expotref_path, 'expot-pharmaref.csv'), sep='\t', encoding='cp1252',
usecols=['PharmaID', 'PharmaName']).drop_duplicates('PharmaID', keep='last')
expotref.rename(columns={'PharmaID': 'VariableID', 'PharmaName': 'VariableName'}, inplace=True)
else:
expotref = pd.read_csv(join(expotref_path, 'expot-varref.csv'), sep='\t', encoding='cp1252',
usecols=['VariableID', 'Abbreviation']).drop_duplicates('VariableID', keep='last')
expotref.rename(columns={'Abbreviation': 'VariableName'}, inplace=True)
if include_all:
ref_voi = expotref[expotref.VariableID.isin(vid_set)].copy()
if use_ref == 'excel':
print('Could not use the excel reference table when include_all=True, because the excel reference table is incomplete.')
else:
vid_voi = set(excelref.VariableID) & set(vid_set)
if use_ref == 'excel':
ref_voi = excelref[excelref.VariableID.isin(vid_voi)].copy()
else:
ref_voi = expotref[excelref.VariableID.isin(vid_voi)].copy()
ref_voi.VariableID = ref_voi.VariableID.astype(int)
ref_voi.VariableName = ref_voi.VariableName.astype(str)
ref_voi.set_index('VariableID', inplace=True)
return ref_voi
def get_consent_patient_ids(recompute=False, replace=False):
"""
Return the list of patientID of patients with consent, i.e. those whose GeneralConsent
value smaller than 4 or not recorded in the observrec table, and not in the
PID_Exclusion_GeneralConsent (sent by Martin) list
"""
id_list_path = join(datapath, 'misc_derived', 'id_lists')
filepath = join(id_list_path, 'PID_WithConsent_not_on_ECMO.csv')
if recompute:
df_gc = pd.read_csv(join(datapath, '0_csv_exports', 'expot-observrec.csv'), sep=';',
usecols=['PatientID', 'VariableID', 'Value'])
pid_exclude = set(pd.read_csv(join(id_list_path, 'PID_Exclusion_GeneralConsent.csv')).PatientID) | set(pd.read_csv(join(id_list_path, 'PID_on_ECMO.csv')).PatientID)
df_gc = df_gc[df_gc.VariableID == 15004651] # extract time series of GeneralConsent
df_gc.loc[:,'Value'] = df_gc.Value.astype(float)
pid_noconsent = set(df_gc[df_gc.Value >= 4].PatientID)
pid_gd = set(pd.read_csv(join(datapath, '0_csv_exports', 'expot-generaldata.csv'), sep=';',
usecols=['PatientID']).PatientID)
pid_any_but_gd = set()
for tbl in get_table_names():
pid_tbl_filepath = join(id_list_path, 'PID_%s.csv'%tbl)
if not exists(pid_tbl_filepath):
import ipdb
ipdb.set_trace()
if tbl in ['monvals', 'comprvals']:
csv_iters = pd.read_csv(join(datapath, '0_csv_exports', 'expot-%s.csv'%tbl), sep=';',
usecols=['PatientID'], chunksize=10**7)
pid_tbl = set()
for i, chunk in enumerate(csv_iters):
pid_tbl = pid_tbl | set(chunk.PatientID)
else:
df_tmp = pd.read_csv(join(datapath, '0_csv_exports', 'expot-%s.csv'%tbl),
usecols=['PatientID'], sep=';')
pid_tbl = set(df_tmp.PatientID)
df_pid_tbl = pd.DataFrame(list(pid_tbl), columns=['PatientID'])
df_pid_tbl.to_csv(pid_tbl_filepath, index=False)
else:
pid_tbl = set(pd.read_csv(pid_tbl_filepath).PatientID)
print('# patients in %s: %d'%(tbl, len(pid_tbl)))
pid_any_but_gd = pid_any_but_gd | pid_tbl
pid_any_and_gd = pid_any_but_gd & pid_gd
pid_gd_include = pid_gd - (pid_exclude | pid_noconsent)
pid_any_but_gd_include = pid_any_but_gd - (pid_exclude | pid_noconsent)
pid_include = list((pid_any_but_gd | pid_gd) - (pid_gd - pid_any_but_gd) - (pid_exclude | pid_noconsent))
print('# patients in any measurement tables but generaldata: %d'%(len(pid_any_but_gd)))
print('# patients in generaldata: %d'%(len(pid_gd)))
print('# patients in any measurement tables but also in generaldata: %d'%(len(pid_any_and_gd)))
print('# patients in generaldata but have no measurement: %d'%(len(pid_gd - pid_any_but_gd)))
print('# patients have measurement but not in generaldata: %d'%(len(pid_any_but_gd - pid_gd)))
print('# patients in "PID_Exclusion_GeneralConsent.csv": %d'%(len(pid_exclude)))
print('# patients in "PID_Exclusion_GeneralConsent.csv" but not in general data: %d'%(len(pid_exclude - pid_gd)))
print('# patients whose GeneralConsent value >= 4: %d'%(len(pid_noconsent)))
print('# patients whose GeneralConsent >=4 and are also in "PID_Exclusion_GeneralConsent.csv": %d'%len(pid_exclude & pid_noconsent))
print('# patients in generaldata who give consent: %d'%(len(pid_gd_include)))
print('# patients in any measurement table but general who give consent: %d'%(len(pid_any_but_gd_include)))
print('# patients who give consent and have measurement data: %d'%(len(pid_include)))
df_pid_include = pd.DataFrame(pid_include, columns=['PatientID'])
if replace:
df_pid_include.to_csv(filepath, index=False)
else:
pid_include = pd.read_csv(filepath, sep=',').PatientID.unique()
pid_include = np.sort(pid_include)
return pid_include
def get_patients_admitted_later(year=2008, rewrite=False):
pid_list_file = join(datapath, 'misc_derived', 'id_lists', 'PID_WithConsent_AdmittedAfter%d_not_on_ECMO.csv'%year)
if not exists(pid_list_file) or rewrite:
static = pd.read_csv(join(datapath, '0_csv_exports', 'expot-generaldata.csv'), usecols=['PatientID', 'AdmissionTime'], sep=';')
pids_with_consent = get_consent_patient_ids()
static = static[static.PatientID.isin(pids_with_consent)].copy()
static['AdmissionTime'] = pd.to_datetime(static['AdmissionTime'])
pid_list = static[static.AdmissionTime.apply(lambda x: x.year >= year)].PatientID.unique()
df = pd.DataFrame(pid_list.reshape((-1,1)), columns=['PatientID'])
df.to_csv(pid_list_file, index=False)
else:
pid_list = np.array(pd.read_csv(pid_list_file).PatientID)
pid_list = np.sort(pid_list)
return pid_list
def load_chunkfile_index(num_chunks=50, rewrite=False):
chunkfile_index_file = join(datapath, 'misc_derived', 'id_lists', 'PID_%dchunkfile_index_not_on_ECMO.csv'%num_chunks)
if not exists(chunkfile_index_file) or rewrite:
pid_list = get_patients_admitted_later()
chunksize = int(np.ceil( len(pid_list) / num_chunks))
chunkfile_index = []
for i in range(num_chunks):
idx_start = i*chunksize
idx_stop = min((i+1)*chunksize, len(pid_list))
pid_list_tmp = pid_list[idx_start:idx_stop]
chunkfile_index.extend([[pid, i] for pid in pid_list_tmp])
chunkfile_index = pd.DataFrame(np.array(chunkfile_index), columns=['PatientID', 'ChunkfileIndex'])
chunkfile_index.to_csv(chunkfile_index_file, index=False)
else:
chunkfile_index = pd.read_csv(chunkfile_index_file)
chunkfile_index = chunkfile_index.set_index('PatientID')
return chunkfile_index
def get_subset_patient_ids():
"""
Return the list of patientID of 5% of the patients with consent
"""
id_list_path = join(datapath, 'misc_derived', 'id_lists')
filepath = join(id_list_path, 'PID_Subset.csv')
if not exists(filepath):
GetValidPidList()
np.random.seed(0)
tmp = np.random.rand(len(pID_set))
pID_subset = pID_set[tmp <= .05] # choose only 5% of the patient
df_pID_subset = pd.DataFrame(pID_subset, columns=['PatientID'])
df_pID_subset.to_csv(filepath, index=False)
else:
pID_subset = pd.read_csv(filepath).PatientID.unique()
return pID_subset
def time_difference(t_early, t_later):
"""
Compute the time difference between t_early and t_later
Parameters:
t_early: np.datetime64, list or pandas series.
t_later: np.datetime64, list or pandas series.
"""
if type(t_early) == list:
t1 = np.array(t_early)
elif type(t_early) == pd.Series:
t1 = np.array(t_early.tolist())
else:
t1 = np.array([t_early])
if type(t_later) == list:
t2 = np.array(t_later)
elif type(t_later) == pd.Series:
t2 = np.array(t_later.tolist())
else:
t2 = np.array([t_later])
timedelta2float = np.vectorize(lambda x: x / np.timedelta64(3600, 's'))
t_diff = timedelta2float(t2 - t1)
return t_diff
def get_all_var_names():
tbl_list = ['monvals', 'comprvals', 'dervals', 'observrec', 'pharmarec', 'labres']
vname_list = []
for tbl in tbl_list:
ref_voi = voi_id_name_mapping(tbl)
vname_list.append(ref_voi.VariableName.unique())
vname_list = np.unique(np.concatenate(tuple(vname_list)))
return vname_list
def read_single_patient_from_merged(patient_id, vname_list=None, verbose=False):
readpath = join(datapath, '3_merged', 'fmat_170327')
t = clock()
if vname_list is None:
vname_list = get_all_var_names()
filename = 'p%s.h5'%patient_id
filepath = join(readpath, filename)
df = [pd.read_hdf(filepath, 'Datetime')]
for var_name in vname_list:
df.append(pd.read_hdf(filepath, var_name))
if verbose:
print('Time to read patient_id=%d: %g sec'%(patient_id, (clock()-t)))
t = clock()
df = pd.concat(df, axis=1, join_axes=[df[0].index])
if verbose:
print('Time to concate patient_id=%d: %g sec'%(patient_id, (clock()-t)))
return df
def generate_id2string():
tbl_list = ['monvals', 'comprvals', 'dervals', 'observrec', 'pharmarec', 'labres']
ref_list = []
for tbl in tbl_list:
ref = voi_id_name_mapping(tbl)
ref_string = voi_id_name_mapping(tbl, replace_name=True)
ref_string.columns = ['string']
ref['string'] = ref_string['string']
ref_list.append(ref)
ref = pd.concat(ref_list)
ref.reset_index(drop=True, inplace=True)
ref.columns = ['id', 'string']
ref.drop_duplicates(inplace=True)
ref.to_csv('id2string.csv', index=False)
return ref
# -
|
utils/Untitled.ipynb
|