GenBio-PathFM
GenBio-PathFM is a histopathology foundation (FM) model from GenBio AI.
At the time of release, GenBio-PathFM is the strongest open-weight histopathology FM and the only state-of-the-art histopathology FM trained exclusively on publicly available data.
Please see GitHub for the model implementation and inference examples.
For more details:
Usage
GenBio-PathFM can be loaded directly via HuggingFace AutoModel (tested on transformers==4.57.1):
from transformers import AutoModel
from torchvision import transforms
# Load model
model = AutoModel.from_pretrained("genbio-ai/genbio-pathfm", trust_remote_code=True)
model.eval()
# Transform
transform = transforms.Compose([
transforms.Resize((224, 224)),
transforms.ToTensor(),
transforms.Normalize(
mean=(0.697, 0.575, 0.728),
std=(0.188, 0.240, 0.187),
),
])
# Inference
import torch
from PIL import Image
image = Image.open("path/to/image.png").convert("RGB")
x = transform(image).unsqueeze(0) # [1, 3, 224, 224]
with torch.no_grad():
cls_features = model(x) # [1, 4608]
# Or with patch tokens:
cls_features, patch_features = model.forward_with_patches(x) # [1, 4608], [1, 196, 4608]
Abstract
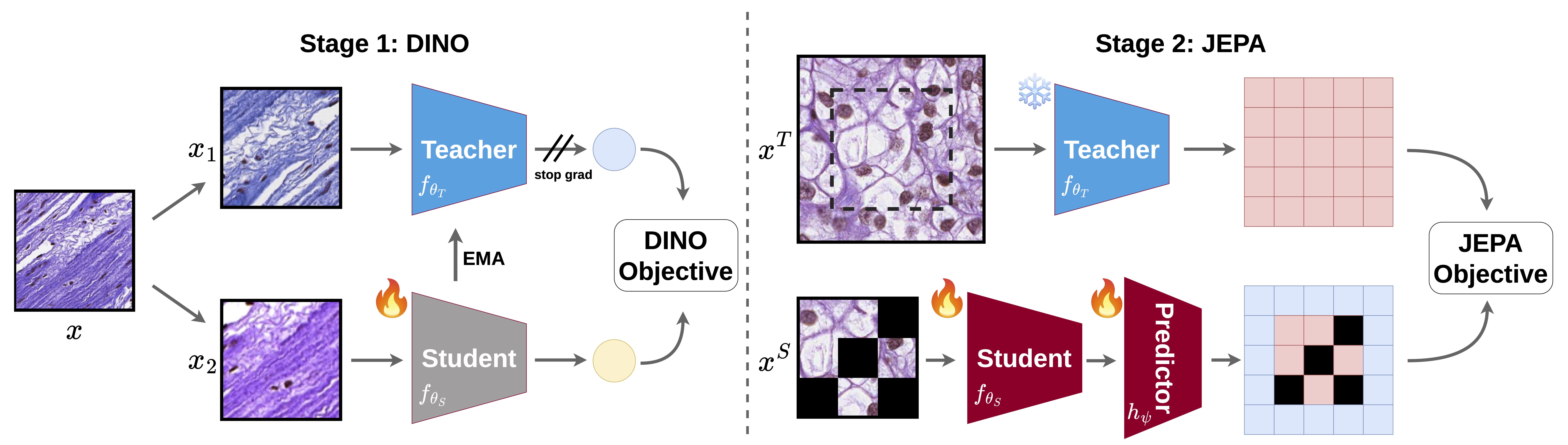
Recent advancements in histopathology foundation models (FMs) have largely been driven by scaling the training data, often utilizing massive proprietary datasets. However, the long-tailed distribution of morphological features in whole-slide images (WSIs) makes simple scaling inefficient, as common morphologies dominate the learning signal. We introduce GenBio-PathFM, a 1.1B-parameter FM that achieves state-of-the-art performance on public benchmarks while using a fraction of the training data required by current leading models. The efficiency of GenBio-PathFM is underpinned by two primary innovations: an automated data curation pipeline that prioritizes morphological diversity and a novel dual-stage learning strategy which we term JEDI (JEPA + DINO). Across the THUNDER, HEST, and PathoROB benchmarks, GenBio-PathFM demonstrates state-of-the-art accuracy and robustness. GenBio-PathFM is the strongest open-weight model to date and the only state-of-the-art model trained exclusively on public data.
License
GenBio-PathFM is available under the GenBio AI Community License.
Reference
If you find our work useful, consider citing our paper:
@article{kapse2026genbiopathfm,
title={GenBio-PathFM: A State-of-the-Art Foundation Model for Histopathology},
author={Kapse, Saarthak and Aygün, Mehmet and Cole, Elijah and Lundberg, Emma and Song, Le and Xing, Eric P.},
journal={bioRxiv},
year={2026}
}
- Downloads last month
- 306