upload intellifold_v2.py
#2
by Henry1112 - opened
- .gitattributes +0 -1
- README.md +8 -30
- ccd_v2.pkl +0 -3
- intellifold_v2.pt +0 -3
- pdb_seqres_2022_09_28.fasta +0 -3
.gitattributes
CHANGED
|
@@ -35,4 +35,3 @@ saved_model/**/* filter=lfs diff=lfs merge=lfs -text
|
|
| 35 |
*tfevents* filter=lfs diff=lfs merge=lfs -text
|
| 36 |
protein_id_groups.json filter=lfs diff=lfs merge=lfs -text
|
| 37 |
unique_protein_sequences.fasta filter=lfs diff=lfs merge=lfs -text
|
| 38 |
-
pdb_seqres_2022_09_28.fasta filter=lfs diff=lfs merge=lfs -text
|
|
|
|
| 35 |
*tfevents* filter=lfs diff=lfs merge=lfs -text
|
| 36 |
protein_id_groups.json filter=lfs diff=lfs merge=lfs -text
|
| 37 |
unique_protein_sequences.fasta filter=lfs diff=lfs merge=lfs -text
|
|
|
README.md
CHANGED
|
@@ -5,7 +5,6 @@ tags:
|
|
| 5 |
- chemistry
|
| 6 |
- biomolecular-structure-prediction
|
| 7 |
- IntelliFold
|
| 8 |
-
library_name: intellifold
|
| 9 |
---
|
| 10 |
|
| 11 |

|
|
@@ -19,27 +18,12 @@ library_name: intellifold
|
|
| 19 |
|
| 20 |
<div align="center" style="margin: 20px 0;">
|
| 21 |
<span style="margin: 0 10px;">⚡ <a href="https://server.intfold.com">IntelliFold Server</a></span>
|
| 22 |
-
• <span style="margin: 0 10px;">📄 <a href="https://
|
| 23 |
-
|
| 24 |
</div>
|
| 25 |
|
| 26 |
|
| 27 |
|
| 28 |
|
| 29 |
-
## 🎉 New Model Release
|
| 30 |
-
|
| 31 |
-
- **2026-02-07**: We are excited to present [[IntelliFold 2]](assets/Intellifold_v2_release_note.pdf). This version represents a
|
| 32 |
-
major architectural update and is one of the first open-source models to outperform AlphaFold3 on
|
| 33 |
-
Foldbench.
|
| 34 |
-
|
| 35 |
-
|
| 36 |
-
## 📊 Benchmarking
|
| 37 |
-
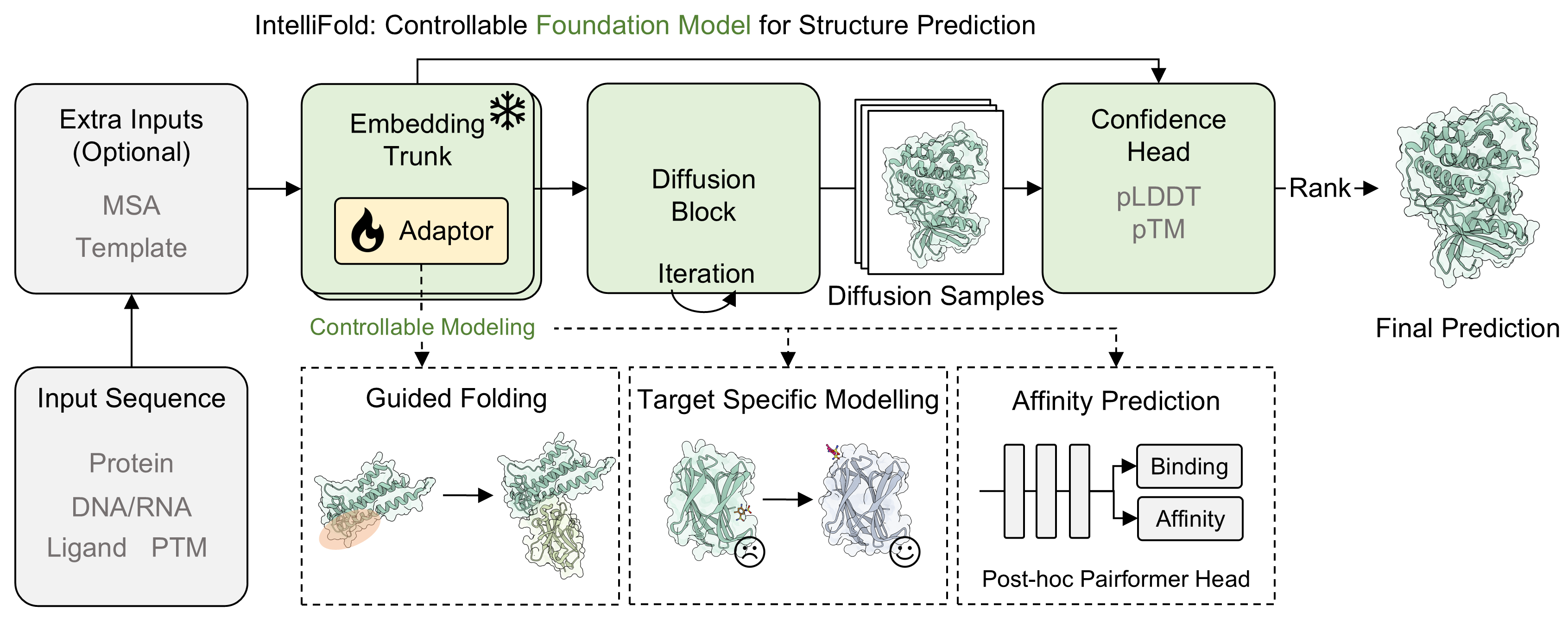
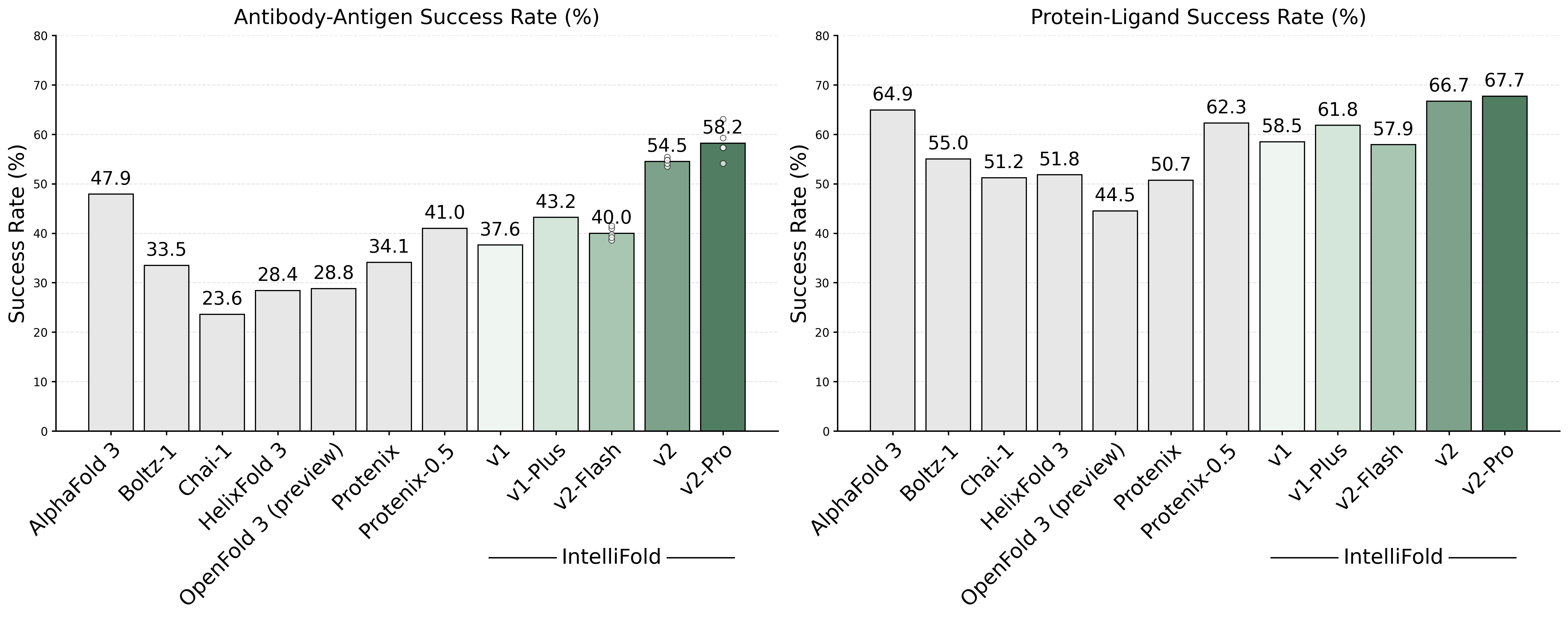
To comprehensively evaluate the performance of IntelliFold 2, we conducted a rigorous evaluation on [FoldBench](https://github.com/BEAM-Labs/FoldBench). We compared IntelliFold against several leading methods, including [Boltz-1,2](https://github.com/jwohlwend/boltz), [Chai-1](https://github.com/chaidiscovery/chai-lab), [Protenix](https://github.com/bytedance/Protenix) and [Alphafold3](https://github.com/google-deepmind/alphafold3).
|
| 38 |
-
|
| 39 |
-
For more details on the benchmarking process and results, please refer to our release note [IntelliFold 2 Release Note](https://raw.githubusercontent.com/IntelliGen-AI/IntelliFold/main/assets/Intellifold_v2_release_note.pdf) and [IntelliFold Technical Report](https://arxiv.org/abs/2507.02025).
|
| 40 |
-
|
| 41 |
-
|
| 42 |
-
|
| 43 |
|
| 44 |
## 🚀 Quick Start
|
| 45 |
|
|
@@ -65,11 +49,6 @@ To more complete installation instructions and usage, please refer to the [Insta
|
|
| 65 |
intellifold predict your_input.yaml --out_dir ./results
|
| 66 |
```
|
| 67 |
|
| 68 |
-
IntelliFold v2-Flash will be used by default, you can also use IntelliFold v2 by specifying the model name:
|
| 69 |
-
```bash
|
| 70 |
-
intellifold predict your_input.yaml --out_dir ./results --model v2
|
| 71 |
-
```
|
| 72 |
-
|
| 73 |
3. **Check Results**: Find predicted structures and confidence scores in the output directory, you can also check the section of **output format** in [output documentation](https://github.com/IntelliGen-AI/IntelliFold/blob/main/docs/input_yaml_format.md#output-format).
|
| 74 |
|
| 75 |
4. **Optional Optimization**: Enable [custom kernels](https://github.com/IntelliGen-AI/IntelliFold/blob/main/docs/kernels.md) for faster inference and reduced memory usage
|
|
@@ -77,6 +56,13 @@ To more complete installation instructions and usage, please refer to the [Insta
|
|
| 77 |
For comprehensive usage instructions and examples, refer to the [Usage Guide](https://github.com/IntelliGen-AI/IntelliFold/blob/main/docs/usage.md).
|
| 78 |
|
| 79 |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 80 |
|
| 81 |
|
| 82 |
## 🌐 IntelliFold Server
|
|
@@ -91,14 +77,6 @@ For comprehensive usage instructions and examples, refer to the [Usage Guide](ht
|
|
| 91 |
If you use IntelliFold in your research, please cite our paper:
|
| 92 |
|
| 93 |
```
|
| 94 |
-
@techreport{qiao2026intellifold,
|
| 95 |
-
title={{IntelliFold 2: Surpassing AlphaFold 3 via Architectural Refinement and Structural Consistency}},
|
| 96 |
-
author={Lifeng Qiao and He Yan and Gary Liu and Gaoxing Guo and Siqi Sun},
|
| 97 |
-
year={2026},
|
| 98 |
-
institution={IntelliGen-AI},
|
| 99 |
-
type={Release Note},
|
| 100 |
-
url={https://raw.githubusercontent.com/IntelliGen-AI/IntelliFold/main/assets/Intellifold_v2_release_note.pdf}
|
| 101 |
-
}
|
| 102 |
@misc{theintfoldteam2025intfoldcontrollablefoundationmodel,
|
| 103 |
title={IntFold: A Controllable Foundation Model for General and Specialized Biomolecular Structure Prediction},
|
| 104 |
author={The IntFold Team and Leon Qiao and Wayne Bai and He Yan and Gary Liu and Nova Xi and Xiang Zhang},
|
|
|
|
| 5 |
- chemistry
|
| 6 |
- biomolecular-structure-prediction
|
| 7 |
- IntelliFold
|
|
|
|
| 8 |
---
|
| 9 |
|
| 10 |

|
|
|
|
| 18 |
|
| 19 |
<div align="center" style="margin: 20px 0;">
|
| 20 |
<span style="margin: 0 10px;">⚡ <a href="https://server.intfold.com">IntelliFold Server</a></span>
|
| 21 |
+
• <span style="margin: 0 10px;">📄 <a href="https://arxiv.org/abs/2507.02025">Technical Report</a></span>
|
|
|
|
| 22 |
</div>
|
| 23 |
|
| 24 |
|
| 25 |

|
| 26 |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 27 |
|
| 28 |
## 🚀 Quick Start
|
| 29 |
|
|
|
|
| 49 |
intellifold predict your_input.yaml --out_dir ./results
|
| 50 |
```
|
| 51 |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 52 |
3. **Check Results**: Find predicted structures and confidence scores in the output directory, you can also check the section of **output format** in [output documentation](https://github.com/IntelliGen-AI/IntelliFold/blob/main/docs/input_yaml_format.md#output-format).
|
| 53 |
|
| 54 |
4. **Optional Optimization**: Enable [custom kernels](https://github.com/IntelliGen-AI/IntelliFold/blob/main/docs/kernels.md) for faster inference and reduced memory usage
|
|
|
|
| 56 |
For comprehensive usage instructions and examples, refer to the [Usage Guide](https://github.com/IntelliGen-AI/IntelliFold/blob/main/docs/usage.md).
|
| 57 |
|
| 58 |
|
| 59 |
+
## 📊 Benchmarking
|
| 60 |
+
To comprehensively evaluate the performance of To quickly get started with IntelliFold, you can use the following commands:
|
| 61 |
+
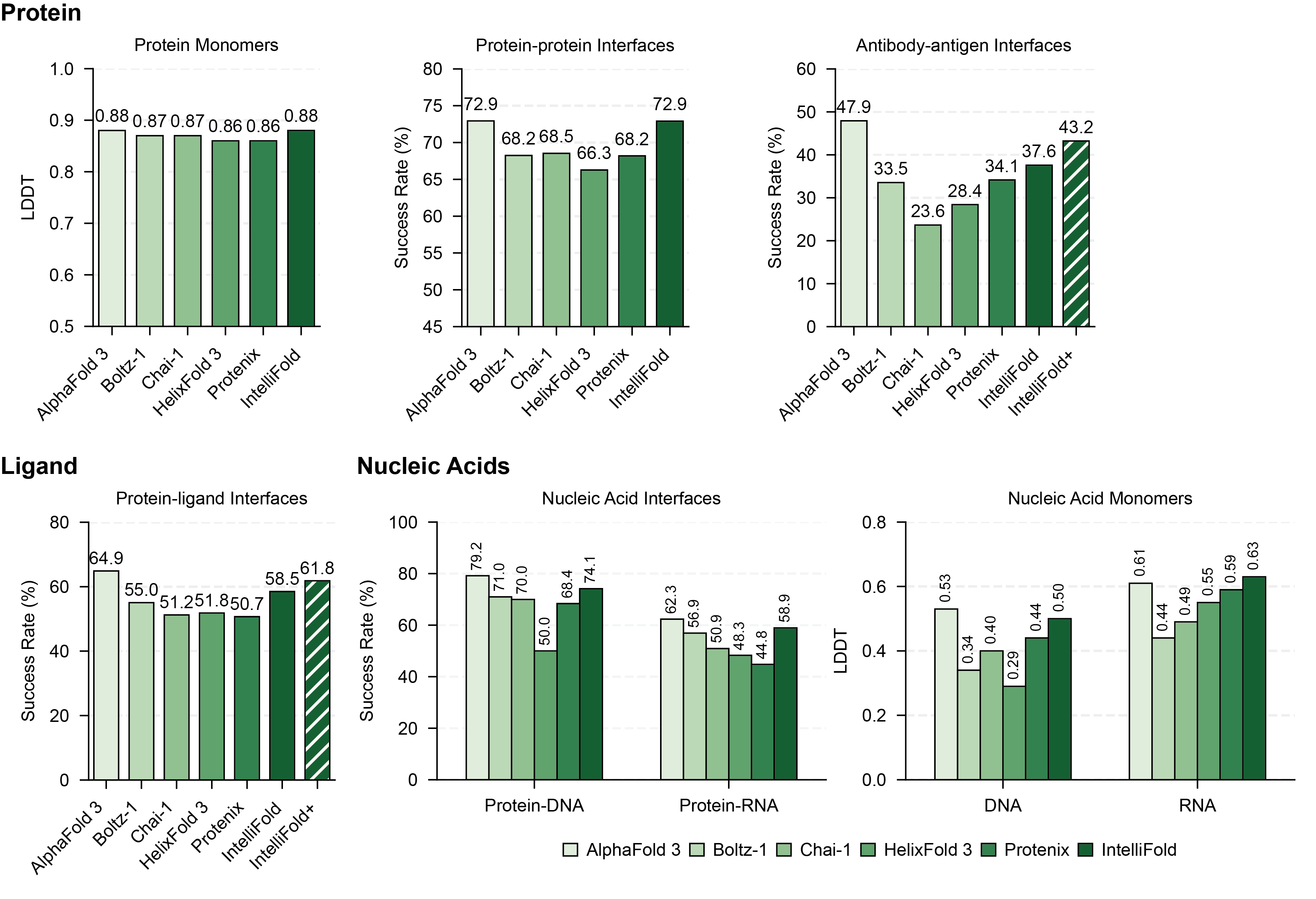
, we conducted a rigorous evaluation on [FoldBench](https://github.com/BEAM-Labs/FoldBench). We compared IntelliFold against several leading methods, including [Boltz-1,2](https://github.com/jwohlwend/boltz), [Chai-1](https://github.com/chaidiscovery/chai-lab), [Protenix](https://github.com/bytedance/Protenix) and [Alphafold3](https://github.com/google-deepmind/alphafold3).
|
| 62 |
+
|
| 63 |
+
For more details on the benchmarking process and results, please refer to our [Technical Report](https://arxiv.org/abs/2507.02025).
|
| 64 |
+
|
| 65 |
+
|
| 66 |
|
| 67 |
|
| 68 |
## 🌐 IntelliFold Server
|
|
|
|
| 77 |
If you use IntelliFold in your research, please cite our paper:
|
| 78 |
|
| 79 |
```
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 80 |
@misc{theintfoldteam2025intfoldcontrollablefoundationmodel,
|
| 81 |
title={IntFold: A Controllable Foundation Model for General and Specialized Biomolecular Structure Prediction},
|
| 82 |
author={The IntFold Team and Leon Qiao and Wayne Bai and He Yan and Gary Liu and Nova Xi and Xiang Zhang},
|
ccd_v2.pkl
DELETED
|
@@ -1,3 +0,0 @@
|
|
| 1 |
-
version https://git-lfs.github.com/spec/v1
|
| 2 |
-
oid sha256:8766edb6a88e01461a123e8a4e2d5e33846821b808444737cd82e441998801f8
|
| 3 |
-
size 413033419
|
|
|
|
|
|
|
|
|
|
|
|
intellifold_v2.pt
DELETED
|
@@ -1,3 +0,0 @@
|
|
| 1 |
-
version https://git-lfs.github.com/spec/v1
|
| 2 |
-
oid sha256:8ee1c03344a94c8d3408f9579b3869f791701b7945e23255331d44fb7cc41aaa
|
| 3 |
-
size 3401386544
|
|
|
|
|
|
|
|
|
|
|
|
pdb_seqres_2022_09_28.fasta
DELETED
|
@@ -1,3 +0,0 @@
|
|
| 1 |
-
version https://git-lfs.github.com/spec/v1
|
| 2 |
-
oid sha256:1b3bc853322c32f2eea818065b8f569a18d25a52326a8d2c2c3de85752e55fe1
|
| 3 |
-
size 232899463
|
|
|
|
|
|
|
|
|
|
|
|