MadriMed-VL-2B
A 2B-parameter multimodal medical vision-language model**, trained for medical image understanding, radiology report assistance, clinical visual question answering, and medical text reasoning.
📊 Benchmarks
Vision-Language (Medical Image Dataset evaluation)
| Benchmark | MadriMed-VL (2B) | MedGemma (4B) | Gap |
|---|---|---|---|
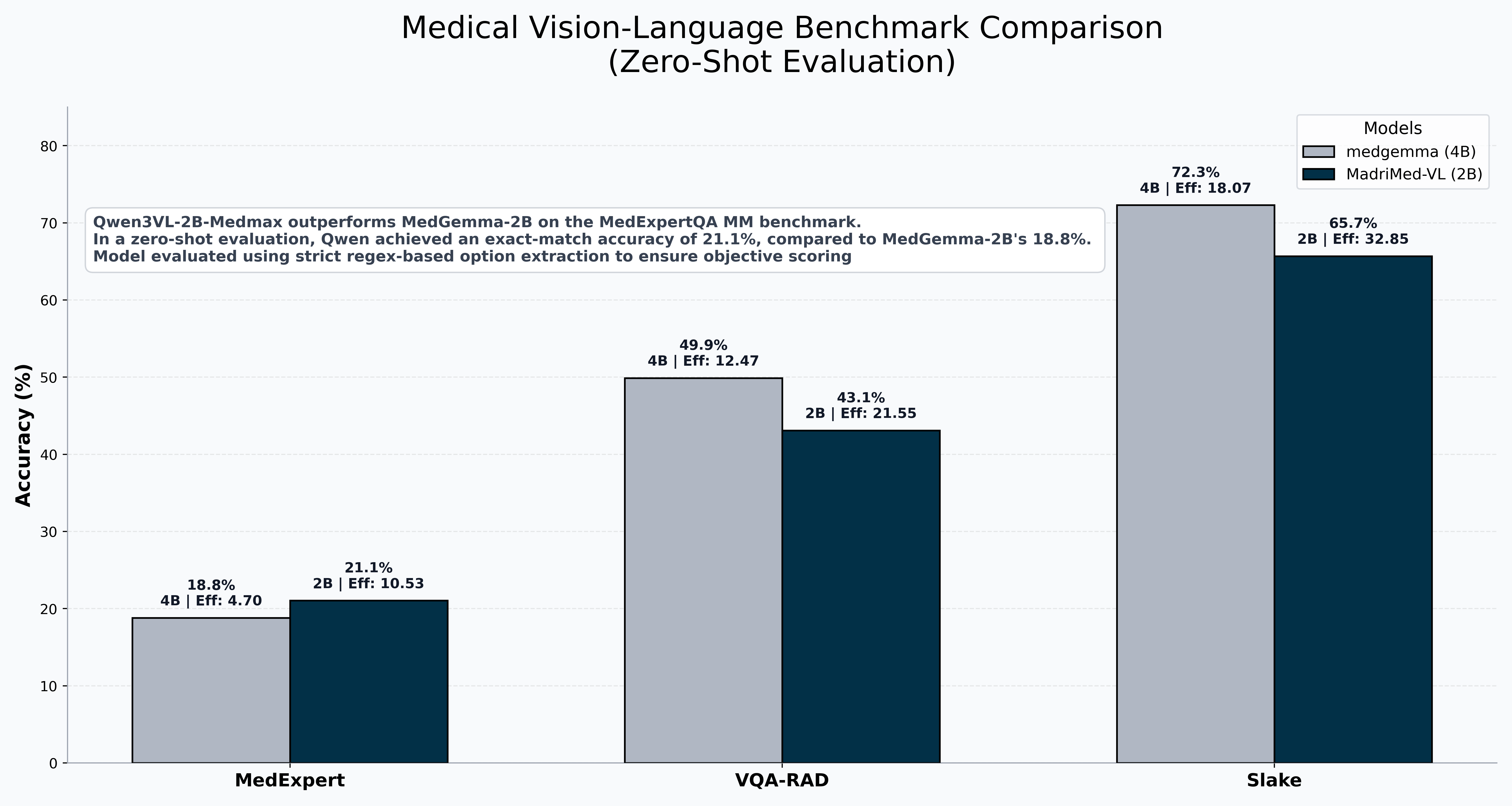
| SLAKE | 65.7% | 72.3% | -6.6 |
| VQA-RAD | 43.09% | 49.9% | -6.81 |
| MedXpertQA-MM | 21.05% | 18.8% | +2.25 |
Key Result: On MedXpertQA-MM — a USMLE-level multimodal benchmark — MadriMed-VL-2B outperforms Google's 4B MedGemma in come some cases despite being half the size. This demonstrates strong clinical reasoning transfer from focused radiology VQA fine-tuning.
Text-Only (Zero-Shot)
⚠️ Important: MadriMed-VL-2B was not trained on text-only medical datasets. These scores reflect zero-shot generalization from visual fine-tuning only.
| Benchmark | MadriMed-VL (2B) | OpenBioLLM (8B) | MedGemma (4B) | Meditron (2B) |
|---|---|---|---|---|
| PubMedQA | 59.90% | 74.1% | 73.4% | 58.2% |
| USMLE-4 (MedQA) | 45.40% | 57.7% | 49.8% | 34.1% |
| MedXpertQA (Text) | 11.02% | 10.7% | 14.2% | N/A |
🎯 What This Model Does
MadriMed-VL-2B is optimized on medical image like Xrays, CTscan, MRI, etc.:
- 🔬 Radiology VQA — answer questions about medical images
- 🫁 Modality/Plane Detection — identify imaging type and orientation
- 🏥 Organ/Position Recognition — localize anatomical structures
- ⚕️ Binary Screening — yes/no presence/absence questions
What It Does NOT Do (Safely)
- ❌ Diagnosis — do not use for patient diagnosis
- ❌ Treatment Recommendations — not trained on clinical guidelines
- ❌ Pathology Detection — abnormality recognition is limited
🚀 Quick Start
Installation
pip install transformers torch Pillow
Run the model directly
import torch
import re
from transformers import AutoProcessor, Qwen3VLForConditionalGeneration
MODEL_PATH = "madrisight/MadriMed-VL-2B"
print(f"🚀 Loading {MODEL_PATH} for MedQA inference...")
# -----------------------
# Load model + processor
# -----------------------
processor = AutoProcessor.from_pretrained(MODEL_PATH, trust_remote_code=True)
model = Qwen3VLForConditionalGeneration.from_pretrained(
MODEL_PATH,
device_map="mps",
trust_remote_code=True
)
model.eval()
# -----------------------
# Output cleaner
# -----------------------
def clean_output(text: str) -> str:
text = re.sub(r"</?think>", "", text)
text = re.sub(r"<\|?image\|?>", "", text)
return text.strip()
prompt = (
"You are a medical reasoning classification system.\n\n"
"Your task is to classify each question into the SINGLE most appropriate underlying immune mechanism.\n\n"
"RULES:\n"
"- Output ONLY one standardized category label\n"
"- Choose the most specific textbook mechanism\n"
"- If multiple are possible, choose the most primary cause\n\n"
"ALLOWED OUTPUT CATEGORIES (use exact wording):\n"
"- CD4+ T-cell deficiency\n"
"- CD8+ T-cell dysfunction\n"
"- Neutropenia\n"
"- Defective phagocytic oxidative burst\n"
"- Humoral immunity deficiency (B-cell dysfunction)\n"
"- Complement deficiency\n"
"- Bone marrow suppression\n"
"- Cytokine signaling defect\n"
"- Hypersensitivity reaction Type I\n"
"- Hypersensitivity reaction Type II\n"
"- Hypersensitivity reaction Type III\n"
"- Hypersensitivity reaction Type IV\n"
"- General immunosuppression (only if no specific mechanism applies)\n\n"
"OUTPUT FORMAT:\n"
"Reasoning: <brief reasoning>\n"
"Final Answer: <category>\n\n"
"Question:\n"
"A patient with HIV develops progressive loss of helper T-cell function leading to opportunistic infections."
)
messages = [
{
"role": "system",
"content": "You are an expert medical AI. You must deeply analyze the question and must provide your final answer."
},
{
"role": "user",
"content": [{"type": "text", "text": prompt}]
}
]
# -----------------------
# Inference
# -----------------------
with torch.inference_mode():
text = processor.apply_chat_template(
messages,
tokenize=False,
add_generation_prompt=True
)
inputs = processor(
text=text,
# images=image,
return_tensors="pt",
padding=True,
truncation=True,
).to("mps")
generated_ids = model.generate(
**inputs,
max_new_tokens=512,
do_sample=False,
repetition_penalty=1.05
)
output_text = processor.batch_decode(
generated_ids[:, inputs.input_ids.shape[1]:],
skip_special_tokens=True
)[0]
output_text = clean_output(output_text)
print(output_text)
Reasoning: The loss of helper T-cell function in HIV leads to impaired immune response, particularly affecting CD4+ T-cells which are crucial for orchestrating immune responses. This aligns with CD4+ T-cell deficiency as it directly impacts the ability of these cells to support other immune cells.
Final Answer: CD4+ T-cell deficiency
⚠️ Limitations & Safety
Known Issues
| Issue | Severity | Details |
|---|---|---|
| Output corruption | 🔴 High | Rare tokenization artifacts (non-ASCII) on very long sequences |
| Hallucination | 🟡 Medium | Invented findings |
| Positive bias | 🟡 Medium | Tendency to answer "yes" on presence questions |
| Pathology substitution | 🟡 Medium | Confuses similar conditions (subdural ↔ subarachnoid) |
Safety Guidelines
- NOT for clinical diagnosis
- NOT for treatment decisions
- Use as educational aid only
- Always verify binary answers
- Check for corruption
🔬 Technical Details
Training Configuration
| Parameter | Value |
|---|---|
| Base model | Qwen/Qwen3-VL-2B-Thinking |
| Training data | medmax |
| Fine-tuning type | Lora SFTP |
| Precision | bfloat16 |
| Hardware | Single Apple MPS (M4 Pro) on heavily optimized Trl (https://github.com/krrish-v/trlmps) |
🙏 Acknowledgments
- Base model: Qwen3-VL-2B-Instruct by Alibaba Cloud
- Training data: mint-medmax/medmax_data
📄 Citation
If you use this model in research, please cite:
@software{madrimedvl2b,
title = {MadriMed-VL-2B: A Compact Multimodal Medical Vision-Language Model},
author = {Madrisight},
year = {2026},
url = {https://huggingface.co/madrisight/MadriMed-VL-2B}
}
Disclaimer: This model is provided for research and educational purposes only. It is not FDA-approved, not clinically validated, and must not be used for patient care without expert human oversight. The authors assume no liability for clinical use.
- Downloads last month
- 37