| # Model Overview | |
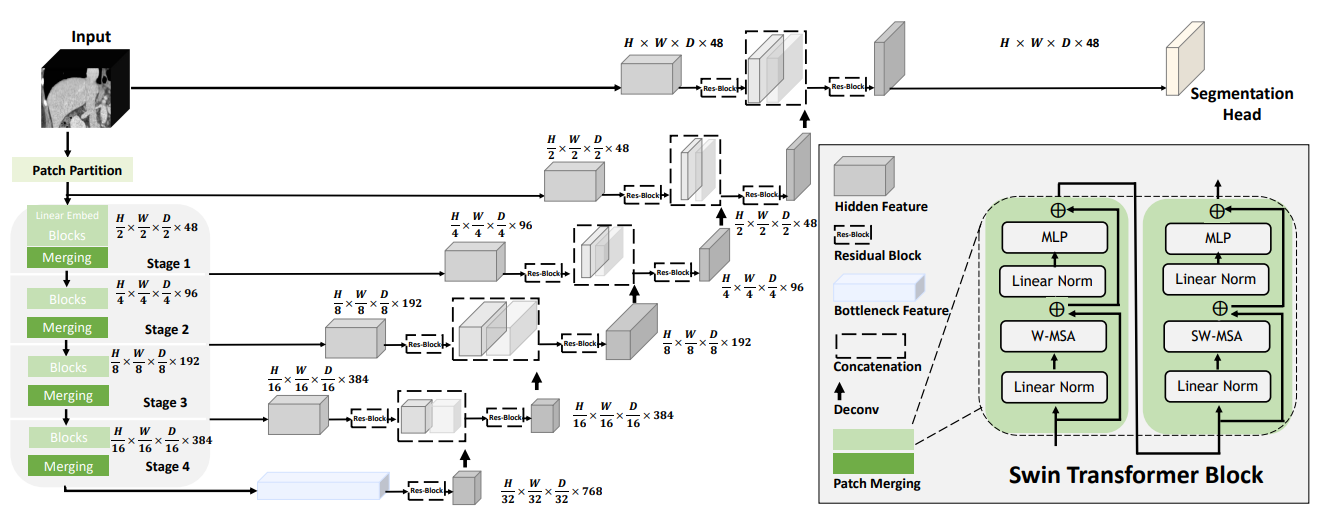
| A pre-trained Swin UNETR [1,2] for volumetric (3D) multi-organ segmentation using CT images from Beyond the Cranial Vault (BTCV) Segmentation Challenge dataset [3]. | |
|  | |
| ## Data | |
| The training data is from the [BTCV dataset](https://www.synapse.org/#!Synapse:syn3193805/wiki/89480/) (Register through `Synapse` and download the `Abdomen/RawData.zip`). | |
| - Target: Multi-organs | |
| - Task: Segmentation | |
| - Modality: CT | |
| - Size: 30 3D volumes (24 Training + 6 Testing) | |
| ### Preprocessing | |
| The dataset format needs to be redefined using the following commands: | |
| ``` | |
| unzip RawData.zip | |
| mv RawData/Training/img/ RawData/imagesTr | |
| mv RawData/Training/label/ RawData/labelsTr | |
| mv RawData/Testing/img/ RawData/imagesTs | |
| ``` | |
| ## Training configuration | |
| The training as performed with the following: | |
| - GPU: At least 32GB of GPU memory | |
| - Actual Model Input: 96 x 96 x 96 | |
| - AMP: True | |
| - Optimizer: Adam | |
| - Learning Rate: 2e-4 | |
| ### Memory Consumption | |
| - Dataset Manager: CacheDataset | |
| - Data Size: 30 samples | |
| - Cache Rate: 1.0 | |
| - Single GPU - System RAM Usage: 5.8G | |
| ### Memory Consumption Warning | |
| If you face memory issues with CacheDataset, you can either switch to a regular Dataset class or lower the caching rate `cache_rate` in the configurations within range [0, 1] to minimize the System RAM requirements. | |
| ### Input | |
| 1 channel | |
| - CT image | |
| ### Output | |
| 14 channels: | |
| - 0: Background | |
| - 1: Spleen | |
| - 2: Right Kidney | |
| - 3: Left Kideny | |
| - 4: Gallbladder | |
| - 5: Esophagus | |
| - 6: Liver | |
| - 7: Stomach | |
| - 8: Aorta | |
| - 9: IVC | |
| - 10: Portal and Splenic Veins | |
| - 11: Pancreas | |
| - 12: Right adrenal gland | |
| - 13: Left adrenal gland | |
| ## Performance | |
| Dice score was used for evaluating the performance of the model. This model achieves a mean dice score of 0.82 | |
| #### Training Loss | |
| 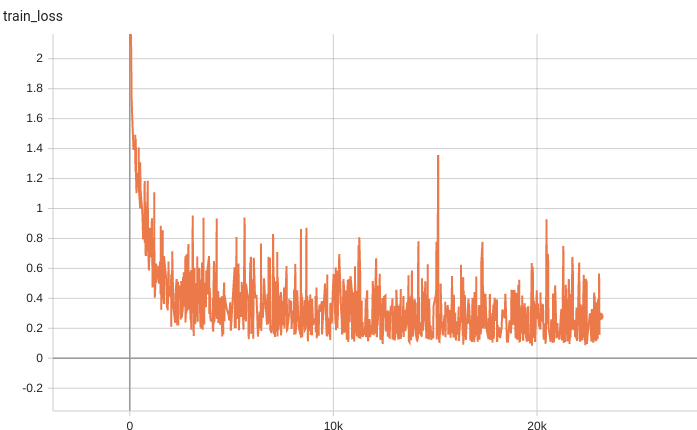 | |
| #### Validation Dice | |
| 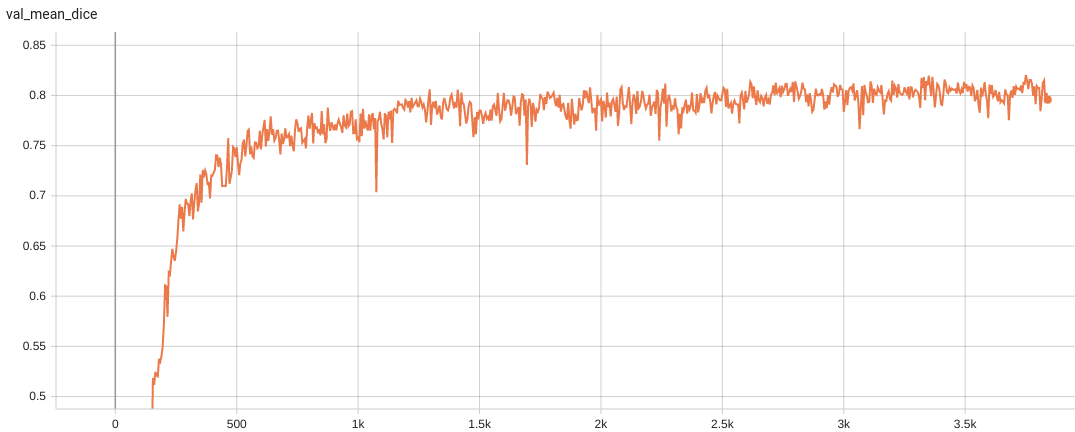 | |
| ## MONAI Bundle Commands | |
| In addition to the Pythonic APIs, a few command line interfaces (CLI) are provided to interact with the bundle. The CLI supports flexible use cases, such as overriding configs at runtime and predefining arguments in a file. | |
| For more details usage instructions, visit the [MONAI Bundle Configuration Page](https://docs.monai.io/en/latest/config_syntax.html). | |
| #### Execute training: | |
| ``` | |
| python -m monai.bundle run --config_file configs/train.json | |
| ``` | |
| Please note that if the default dataset path is not modified with the actual path in the bundle config files, you can also override it by using `--dataset_dir`: | |
| ``` | |
| python -m monai.bundle run --config_file configs/train.json --dataset_dir <actual dataset path> | |
| ``` | |
| #### Override the `train` config to execute multi-GPU training: | |
| ``` | |
| torchrun --standalone --nnodes=1 --nproc_per_node=2 -m monai.bundle run --config_file "['configs/train.json','configs/multi_gpu_train.json']" | |
| ``` | |
| Please note that the distributed training-related options depend on the actual running environment; thus, users may need to remove `--standalone`, modify `--nnodes`, or do some other necessary changes according to the machine used. For more details, please refer to [pytorch's official tutorial](https://pytorch.org/tutorials/intermediate/ddp_tutorial.html). | |
| #### Override the `train` config to execute evaluation with the trained model: | |
| ``` | |
| python -m monai.bundle run --config_file "['configs/train.json','configs/evaluate.json']" | |
| ``` | |
| #### Execute inference: | |
| ``` | |
| python -m monai.bundle run --config_file configs/inference.json | |
| ``` | |
| #### Export checkpoint to TorchScript file: | |
| TorchScript conversion is currently not supported. | |
| # References | |
| [1] Hatamizadeh, Ali, et al. "Swin UNETR: Swin Transformers for Semantic Segmentation of Brain Tumors in MRI Images." arXiv preprint arXiv:2201.01266 (2022). https://arxiv.org/abs/2201.01266. | |
| [2] Tang, Yucheng, et al. "Self-supervised pre-training of swin transformers for 3d medical image analysis." arXiv preprint arXiv:2111.14791 (2021). https://arxiv.org/abs/2111.14791. | |
| [3] Landman B, et al. "MICCAI multi-atlas labeling beyond the cranial vault–workshop and challenge." In Proc. of the MICCAI Multi-Atlas Labeling Beyond Cranial Vault—Workshop Challenge 2015 Oct (Vol. 5, p. 12). | |
| # License | |
| Copyright (c) MONAI Consortium | |
| Licensed under the Apache License, Version 2.0 (the "License"); | |
| you may not use this file except in compliance with the License. | |
| You may obtain a copy of the License at | |
| http://www.apache.org/licenses/LICENSE-2.0 | |
| Unless required by applicable law or agreed to in writing, software | |
| distributed under the License is distributed on an "AS IS" BASIS, | |
| WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. | |
| See the License for the specific language governing permissions and | |
| limitations under the License. | |