YAML Metadata Warning:empty or missing yaml metadata in repo card
Check out the documentation for more information.
SegFormer3D
SegFormer3D is a novel and efficient transformer-based architecture designed specifically for 3D medical image segmentation. It extends the successful 2D SegFormer architecture to handle volumetric medical data while maintaining computational efficiency and strong segmentation performance.
Model Description
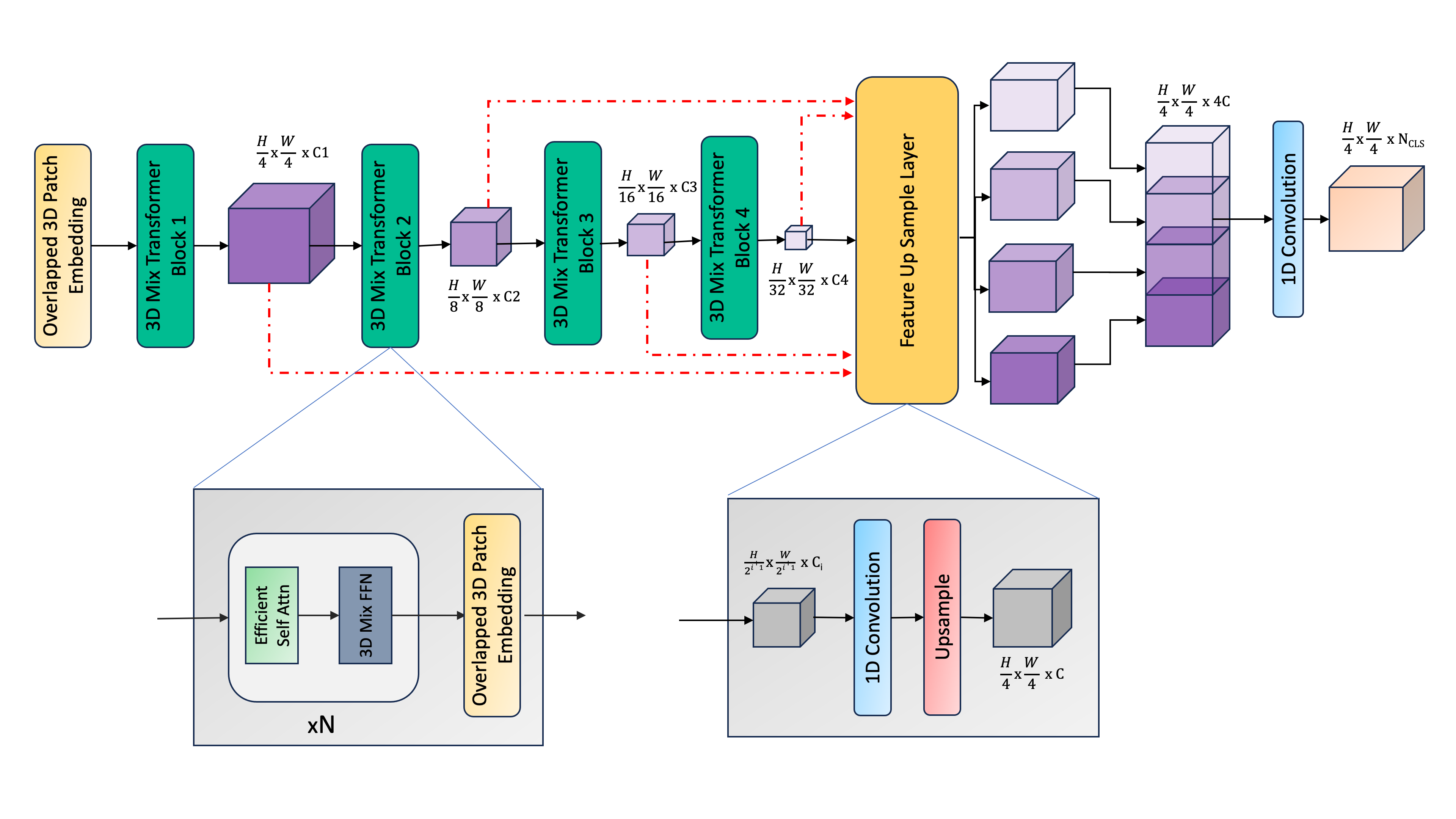
SegFormer3D introduces several key innovations for efficient 3D medical image segmentation:
- Hierarchical 3D Feature Learning: Uses a multi-scale transformer encoder with progressively reduced sequence lengths for efficient processing of volumetric data
- Efficient Self-Attention: Implements spatially-reduced attention mechanism adapted for 3D, reducing computational complexity while maintaining performance
- All-MLP 3D Decoder: Lightweight decoder that effectively fuses multi-scale features through simple MLP layers
- Memory-Efficient Design: Optimized architecture that can process full 3D volumes without patch-based inference
The model achieves state-of-the-art performance on multiple 3D medical segmentation benchmarks while being computationally efficient.
Training and Evaluation
The model was trained and evaluated on several medical imaging datasets:
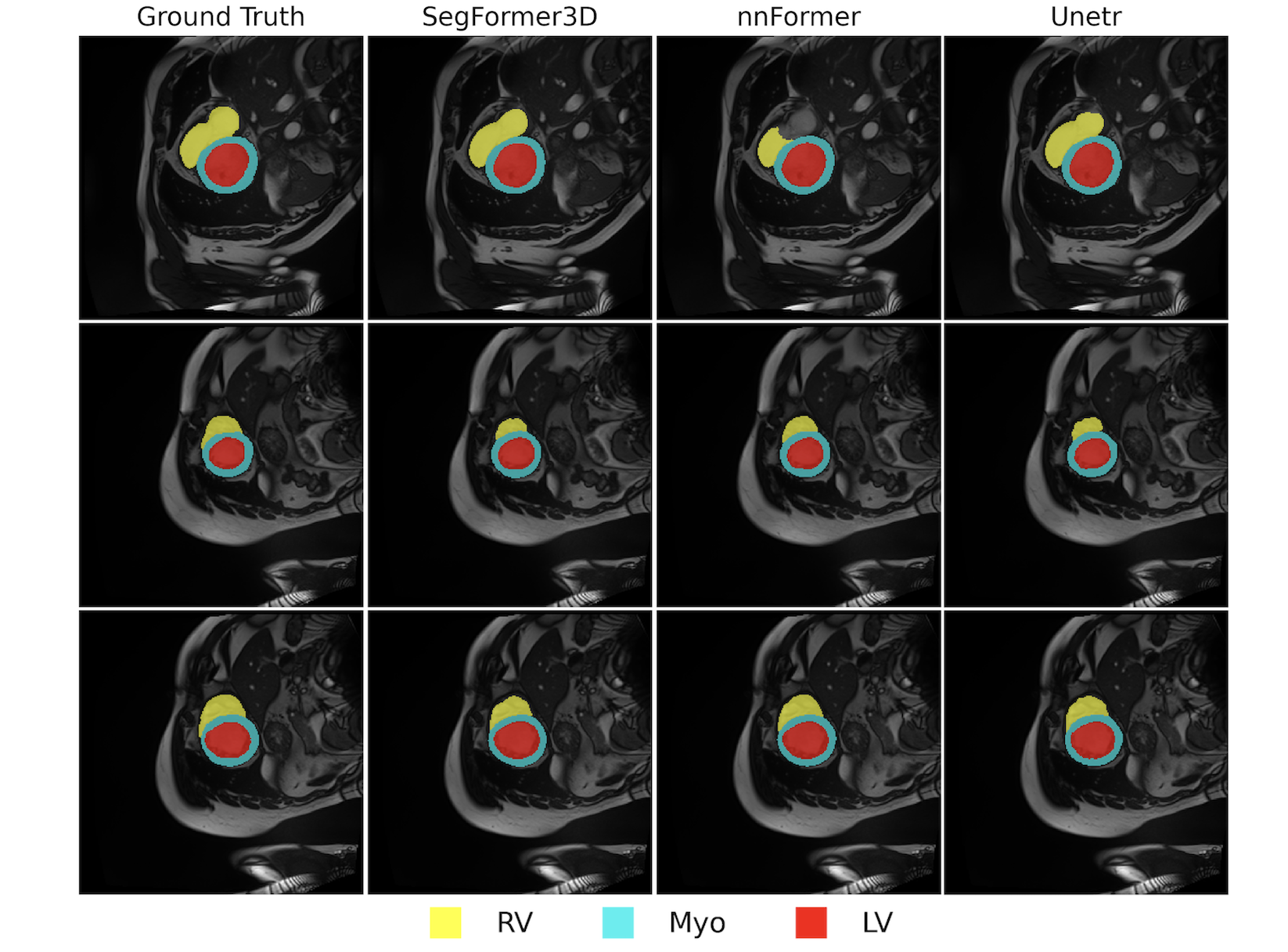
ACDC Dataset
- Task: Cardiac MRI segmentation
- Classes: Left ventricle, right ventricle, and myocardium
- Performance:
- Dice Score: 90.96%
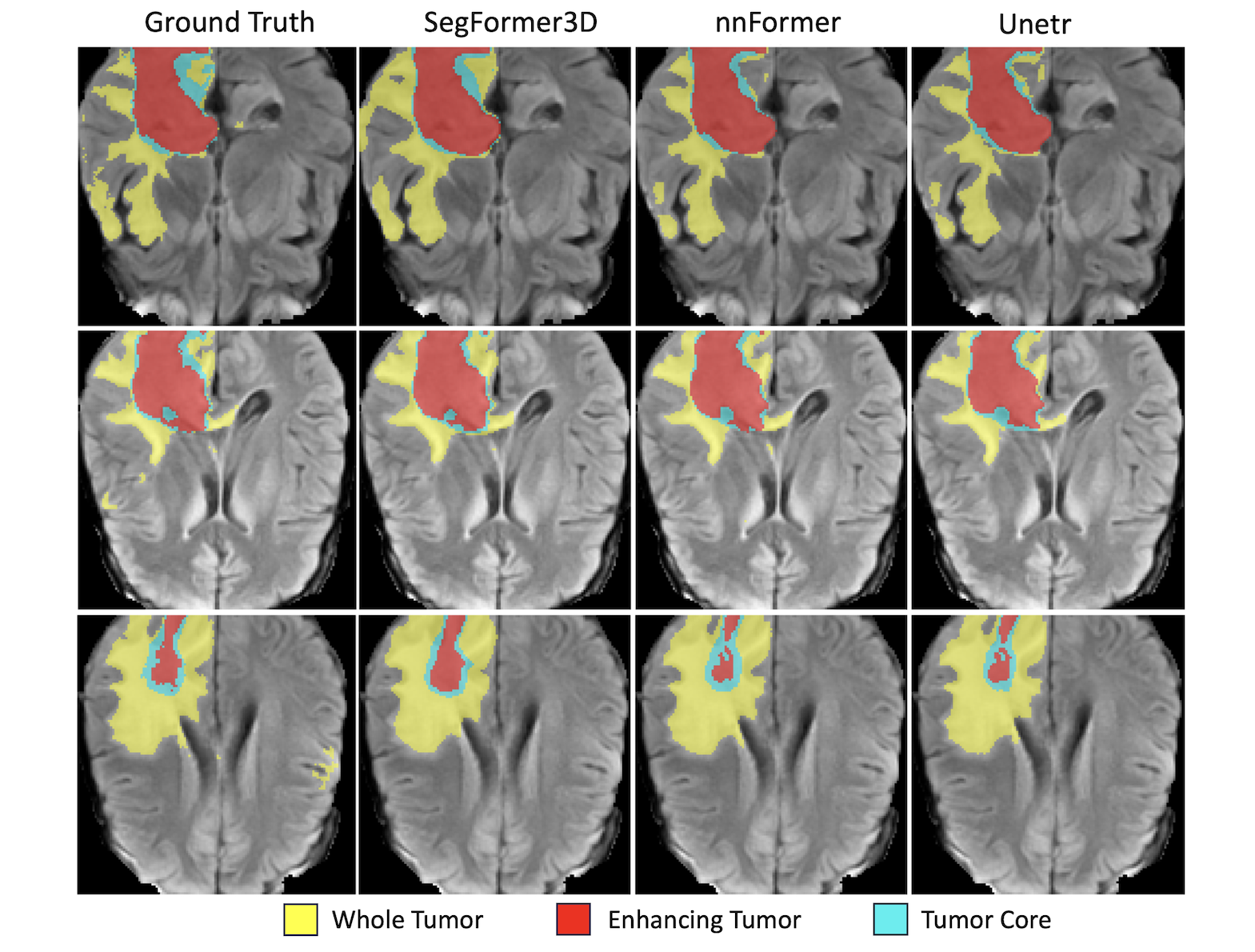
BraTS 2017
- Task: Brain tumor segmentation
- Classes: Enhancing tumor, tumor core, and whole tumor
Performance: - Average Dice: 82.1%
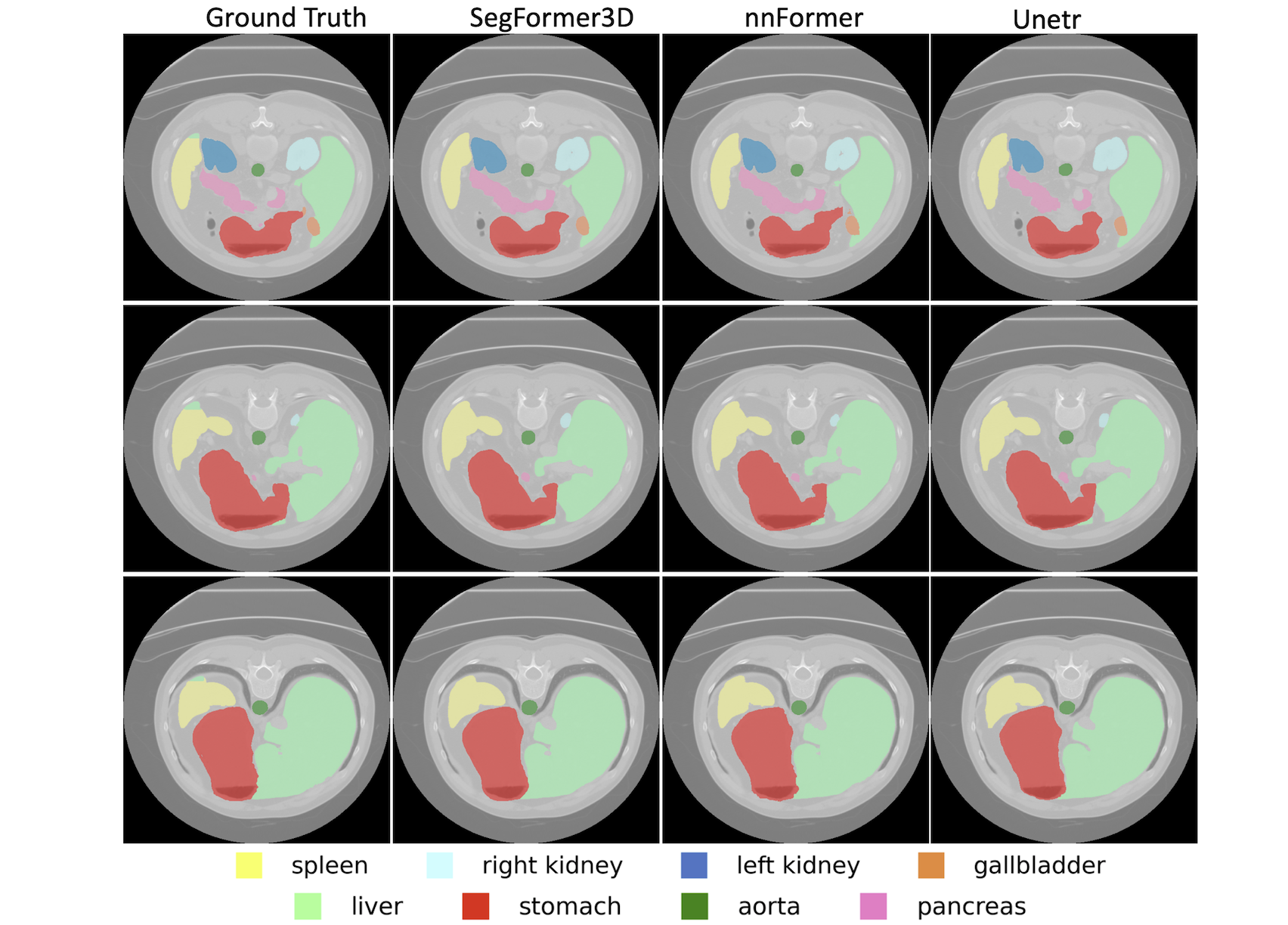
Synapse Dataset
- Task: Multi-organ segmentation
- Classes: 8 abdominal organs
- Performance:
- Average Dice: 82.15%
Usage
from transformers import SegFormer3DConfig, SegFormer3DModel
import torch
# Initialize configuration
config = SegFormer3DConfig(
in_channels=4, # Number of input channels
num_classes=3, # Number of segmentation classes
# Model architecture parameters
embed_dims=[32, 64, 160, 256],
num_heads=[1, 2, 5, 8],
depths=[2, 2, 2, 2],
sr_ratios=[4, 2, 1, 1]
)
# Initialize model
model = SegFormer3DModel(config)
# Example forward pass
batch_size = 1
depth, height, width = 128, 128, 128 # Example input dimensions
x = torch.randn(batch_size, config.in_channels, depth, height, width)
outputs = model(x)
# Get segmentation logits
logits = outputs.logits # Shape: (batch_size, num_classes, D, H, W)
Limitations
- Input dimensions must be properly configured to ensure valid spatial dimensions after each stage
- Memory requirements increase with input volume size
- Performance may vary on different medical imaging modalities
- Assumes consistent voxel spacing in input volumes
Training Tips
Input Preprocessing:
- Normalize input volumes to [0, 1] range
- Consider standardization per modality
- Use appropriate data augmentation for medical volumes
Training Strategy:
- Start with a smaller learning rate (1e-4 recommended)
- Use gradient clipping to stabilize training
- Consider mixed precision training for memory efficiency
Memory Management:
- Adjust batch size based on available GPU memory
- Use gradient checkpointing if needed
- Consider input volume size vs. model depth trade-offs
Citation
@InProceedings{Perera_2024_CVPR,
title={Segformer3d: an efficient transformer for 3d medical image segmentation},
author={Perera, Shehan and Navard, Pouyan and Yilmaz, Alper},
booktitle={Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition},
pages={4981--4988},
year={2024}
}
Acknowledgements
This implementation is based on the original SegFormer architecture by Xie et al. and extends it to efficient 3D medical image segmentation. We thank the authors for their valuable contributions to the field.
