Spaces:
Sleeping
A newer version of the Gradio SDK is available:
6.3.0
title: Lungs Segmentation Web App
emoji: 🖥️
colorFrom: indigo
colorTo: green
sdk: gradio
sdk_version: 5.49.1
app_file: app.py
pinned: false
🖥️ Lungs segmentation web application
A web-based application for automated lung segmentation using deep learning, powered by Gradio and PyTorch. This tool allows users to upload lung images and obtain segmented outputs efficiently.
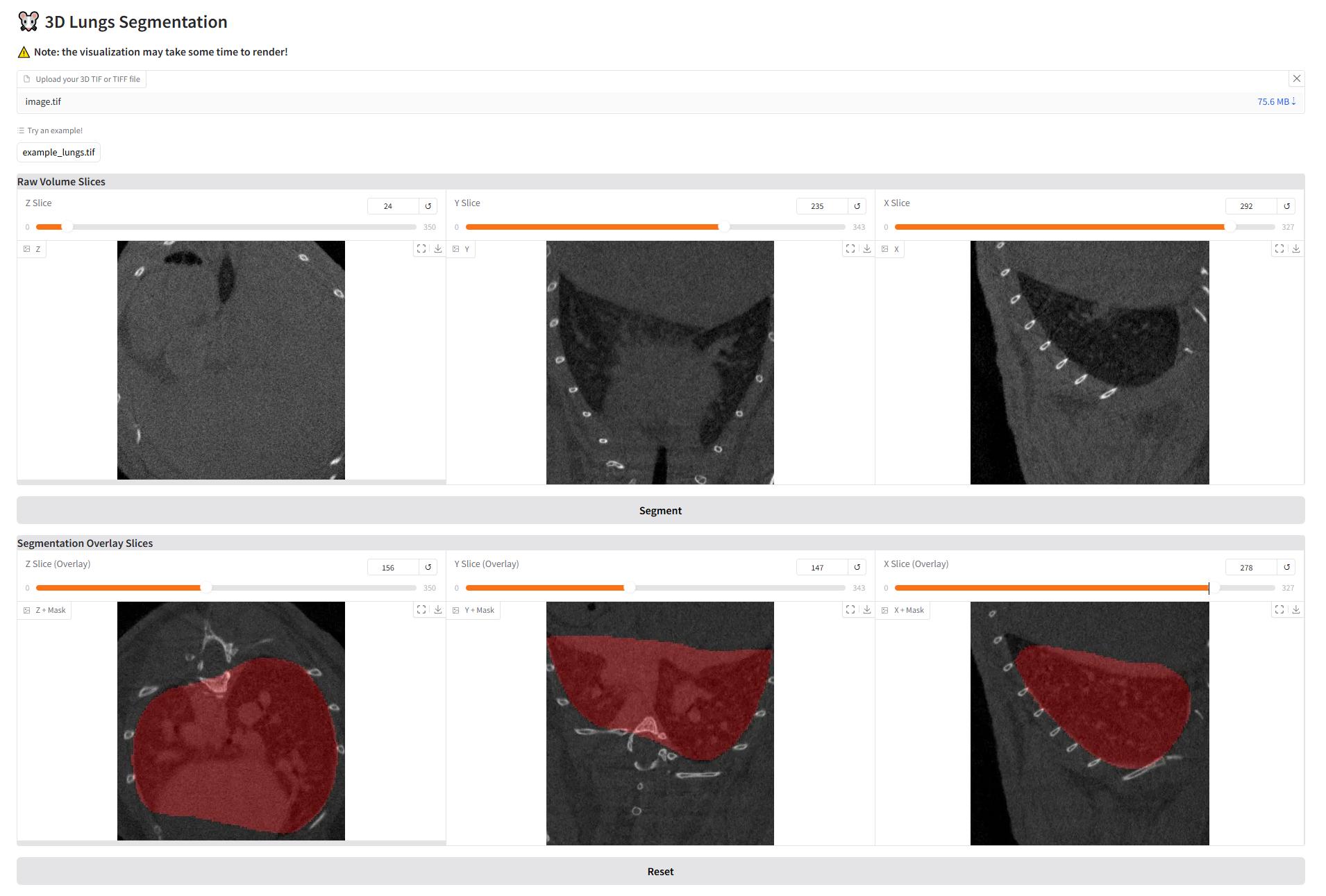
Try the app
The application is running on Hugging Face, try it using this link!
Example File
If you don't have your own .tif image, the app includes a built-in example file that can be used directly from the UI by clicking "Try an example!".
Load from URL (file_url parameter)
You can also provide a .tif file hosted online using a URL parameter.
To do so, simply append ?file_url=... to your app's URL.
Example (hosted on Hugging Face):
https://huggingface.co/spaces/qchapp/3d-lungs-segmentation/?file_url=https://zenodo.org/record/8099852/files/lungs_ct.tif
The application will automatically download the file and load it into the viewer (the operation can take some time).
Installation
We recommend performing the installation in a clean Python environment.
The code requires python>=3.10, as well as pytorch>=2.0. Please install Pytorch first and separately following the instructions for your platform on pytorch.org.
After that please run the following command:
pip install -r requirements.txt
Usage
Run:
python app.py
And go to the indicated local URL.
Usage as an API
Install gradio_client and run the following Python code:
from pathlib import Path
import shutil
from gradio_client import Client, handle_file
client = Client("qchapp/3d-lungs-segmentation")
result_path = client.predict(
file_obj=handle_file("https://zenodo.org/record/8099852/files/lungs_ct.tif?download=1"),
api_name="/segment",
)
dest = Path("outputs/mask.tif")
dest.parent.mkdir(parents=True, exist_ok=True)
shutil.copy(result_path, dest)
print("Saved the mask in:", dest.resolve())
About Lungs Segmentation
If you are interested in the package used for segmentation please check the following GitHub repository!