| # Model Overview | |
| A pre-trained model for volumetric (3D) segmentation of brain tumor subregions from multimodal MRIs based on BraTS 2018 data. | |
| The model is trained to segment 3 nested subregions of primary brain tumors (gliomas): the "enhancing tumor" (ET), the "tumor core" (TC), the "whole tumor" (WT) based on 4 aligned input MRI scans (T1c, T1, T2, FLAIR). | |
| - The ET is described by areas that show hyper intensity in T1c when compared to T1, but also when compared to "healthy" white matter in T1c. | |
| - The TC describes the bulk of the tumor, which is what is typically resected. The TC entails the ET, as well as the necrotic (fluid-filled) and the non-enhancing (solid) parts of the tumor. | |
| - The WT describes the complete extent of the disease, as it entails the TC and the peritumoral edema (ED), which is typically depicted by hyper-intense signal in FLAIR. | |
|  | |
| ## Data | |
| The training data is from the [Multimodal Brain Tumor Segmentation Challenge (BraTS) 2018](https://www.med.upenn.edu/sbia/brats2018.html). | |
| - Target: 3 tumor subregions | |
| - Task: Segmentation | |
| - Modality: MRI | |
| - Size: 285 3D volumes (4 channels each) | |
| The provided labelled data was partitioned, based on our own split, into training (200 studies), validation (42 studies) and testing (43 studies) datasets. | |
| ### Preprocessing | |
| The data list/split can be created with the script `scripts/prepare_datalist.py`. | |
| ``` | |
| python scripts/prepare_datalist.py --path your-brats18-dataset-path | |
| ``` | |
| ## Training configuration | |
| This model utilized a similar approach described in 3D MRI brain tumor segmentation using autoencoder regularization, which was a winning method in BraTS2018 [1]. The training was performed with the following: | |
| - GPU: At least 16GB of GPU memory. | |
| - Actual Model Input: 224 x 224 x 144 | |
| - AMP: True | |
| - Optimizer: Adam | |
| - Learning Rate: 1e-4 | |
| - Loss: DiceLoss | |
| ## Input | |
| 4 channel aligned MRIs at 1 x 1 x 1 mm | |
| - T1c | |
| - T1 | |
| - T2 | |
| - FLAIR | |
| ## Output | |
| 3 channels | |
| - Label 0: TC tumor subregion | |
| - Label 1: WT tumor subregion | |
| - Label 2: ET tumor subregion | |
| ## Performance | |
| Dice score was used for evaluating the performance of the model. This model achieved Dice scores on the validation data of: | |
| - Tumor core (TC): 0.8559 | |
| - Whole tumor (WT): 0.9026 | |
| - Enhancing tumor (ET): 0.7905 | |
| - Average: 0.8518 | |
| Please note that this bundle is non-deterministic because of the trilinear interpolation used in the network. Therefore, reproducing the training process may not get exactly the same performance. | |
| Please refer to https://pytorch.org/docs/stable/notes/randomness.html#reproducibility for more details about reproducibility. | |
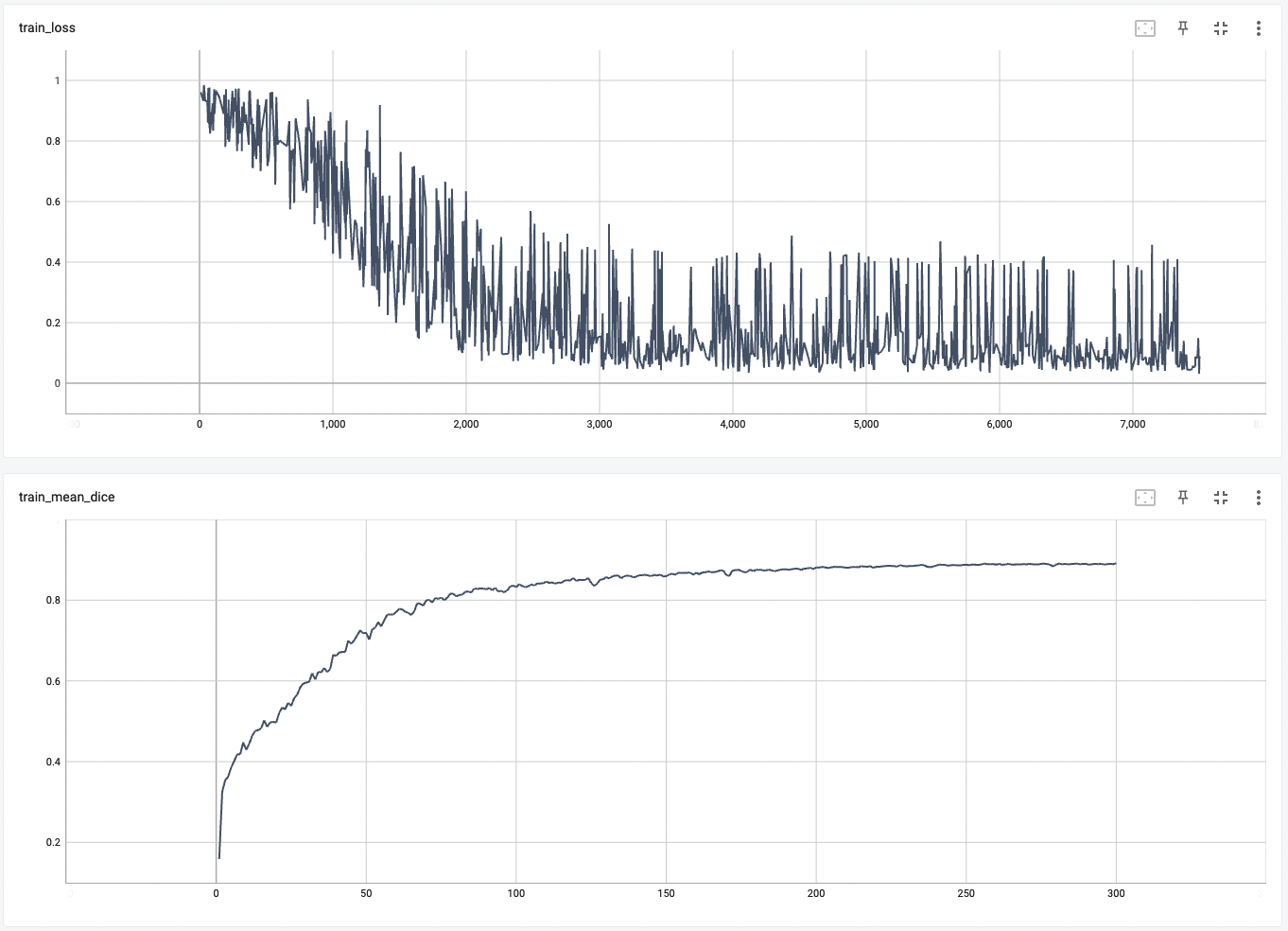
| #### Training Loss and Dice | |
|  | |
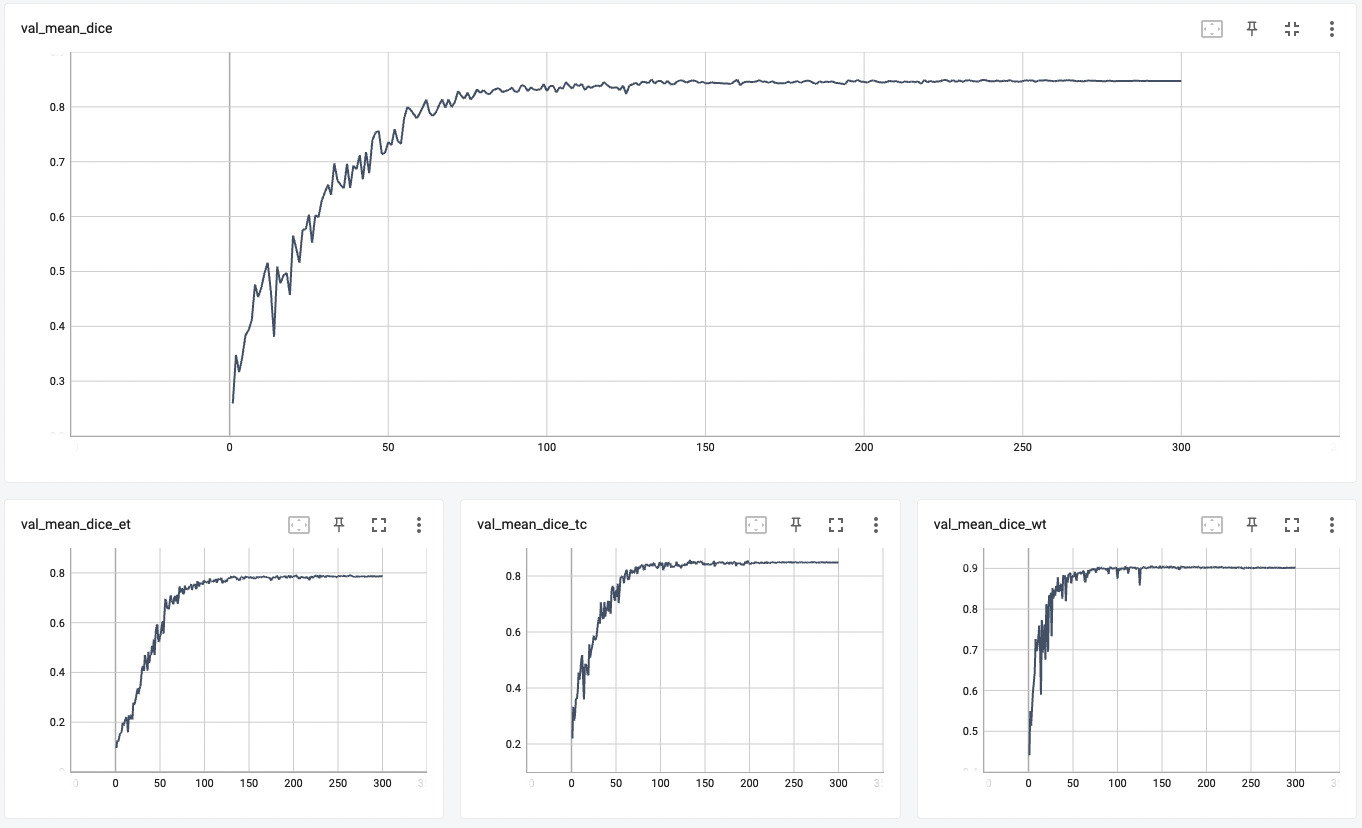
| #### Validation Dice | |
|  | |
| #### TensorRT speedup | |
| The `brats_mri_segmentation` bundle supports acceleration with TensorRT through the ONNX-TensorRT method. The table below displays the speedup ratios observed on an A100 80G GPU. | |
| | method | torch_fp32(ms) | torch_amp(ms) | trt_fp32(ms) | trt_fp16(ms) | speedup amp | speedup fp32 | speedup fp16 | amp vs fp16| | |
| | :---: | :---: | :---: | :---: | :---: | :---: | :---: | :---: | :---: | | |
| | model computation | 5.49 | 4.36 | 2.35 | 2.09 | 1.26 | 2.34 | 2.63 | 2.09 | | |
| | end2end | 592.01 | 434.59 | 395.73 | 394.93 | 1.36 | 1.50 | 1.50 | 1.10 | | |
| Where: | |
| - `model computation` means the speedup ratio of model's inference with a random input without preprocessing and postprocessing | |
| - `end2end` means run the bundle end-to-end with the TensorRT based model. | |
| - `torch_fp32` and `torch_amp` are for the PyTorch models with or without `amp` mode. | |
| - `trt_fp32` and `trt_fp16` are for the TensorRT based models converted in corresponding precision. | |
| - `speedup amp`, `speedup fp32` and `speedup fp16` are the speedup ratios of corresponding models versus the PyTorch float32 model | |
| - `amp vs fp16` is the speedup ratio between the PyTorch amp model and the TensorRT float16 based model. | |
| Currently, the only available method to accelerate this model is through ONNX-TensorRT. However, the Torch-TensorRT method is under development and will be available in the near future. | |
| This result is benchmarked under: | |
| - TensorRT: 8.5.3+cuda11.8 | |
| - Torch-TensorRT Version: 1.4.0 | |
| - CPU Architecture: x86-64 | |
| - OS: ubuntu 20.04 | |
| - Python version:3.8.10 | |
| - CUDA version: 12.0 | |
| - GPU models and configuration: A100 80G | |
| ## MONAI Bundle Commands | |
| In addition to the Pythonic APIs, a few command line interfaces (CLI) are provided to interact with the bundle. The CLI supports flexible use cases, such as overriding configs at runtime and predefining arguments in a file. | |
| For more details usage instructions, visit the [MONAI Bundle Configuration Page](https://docs.monai.io/en/latest/config_syntax.html). | |
| #### Execute training: | |
| ``` | |
| python -m monai.bundle run --config_file configs/train.json | |
| ``` | |
| Please note that if the default dataset path is not modified with the actual path in the bundle config files, you can also override it by using `--dataset_dir`: | |
| ``` | |
| python -m monai.bundle run --config_file configs/train.json --dataset_dir <actual dataset path> | |
| ``` | |
| #### Override the `train` config to execute multi-GPU training: | |
| ``` | |
| torchrun --standalone --nnodes=1 --nproc_per_node=8 -m monai.bundle run --config_file "['configs/train.json','configs/multi_gpu_train.json']" | |
| ``` | |
| Please note that the distributed training-related options depend on the actual running environment; thus, users may need to remove `--standalone`, modify `--nnodes`, or do some other necessary changes according to the machine used. For more details, please refer to [pytorch's official tutorial](https://pytorch.org/tutorials/intermediate/ddp_tutorial.html). | |
| #### Override the `train` config to execute evaluation with the trained model: | |
| ``` | |
| python -m monai.bundle run --config_file "['configs/train.json','configs/evaluate.json']" | |
| ``` | |
| #### Execute inference: | |
| ``` | |
| python -m monai.bundle run --config_file configs/inference.json | |
| ``` | |
| #### Export checkpoint to TensorRT based models with fp32 or fp16 precision: | |
| ```bash | |
| python -m monai.bundle trt_export --net_id network_def \ | |
| --filepath models/model_trt.ts --ckpt_file models/model.pt \ | |
| --meta_file configs/metadata.json --config_file configs/inference.json \ | |
| --precision <fp32/fp16> --input_shape "[1, 4, 240, 240, 160]" --use_onnx "True" \ | |
| --use_trace "True" | |
| ``` | |
| #### Execute inference with the TensorRT model: | |
| ``` | |
| python -m monai.bundle run --config_file "['configs/inference.json', 'configs/inference_trt.json']" | |
| ``` | |
| # References | |
| [1] Myronenko, Andriy. "3D MRI brain tumor segmentation using autoencoder regularization." International MICCAI Brainlesion Workshop. Springer, Cham, 2018. https://arxiv.org/abs/1810.11654. | |
| # License | |
| Copyright (c) MONAI Consortium | |
| Licensed under the Apache License, Version 2.0 (the "License"); | |
| you may not use this file except in compliance with the License. | |
| You may obtain a copy of the License at | |
| http://www.apache.org/licenses/LICENSE-2.0 | |
| Unless required by applicable law or agreed to in writing, software | |
| distributed under the License is distributed on an "AS IS" BASIS, | |
| WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. | |
| See the License for the specific language governing permissions and | |
| limitations under the License. | |