license: cc-by-nc-4.0
tags:
- computational-pathology
- survival-analysis
- whole-slide-imaging
- gene-expression
- oncology
- histopathology
extra_gated_prompt: |
These weights are released under CC BY-NC 4.0 — strictly non-commercial,
research and educational use only. By requesting access you agree to:
1. Use the weights only for non-commercial research.
2. Cite the SPARC paper in any derived publication.
3. Not redistribute the weights to third parties.
extra_gated_fields:
Name: text
Affiliation: text
Email: text
Intended use: text
I agree to the non-commercial license: checkbox
SPARC
Gene-program-aware survival modelling from H&E whole-slide images.
This repository hosts the trained model weights for the SPARC paper (Ayed, Cohn, et al.). Code, configs, training scripts, and figure-regeneration notebooks live at github.com/aziz-ayed/SPARC.
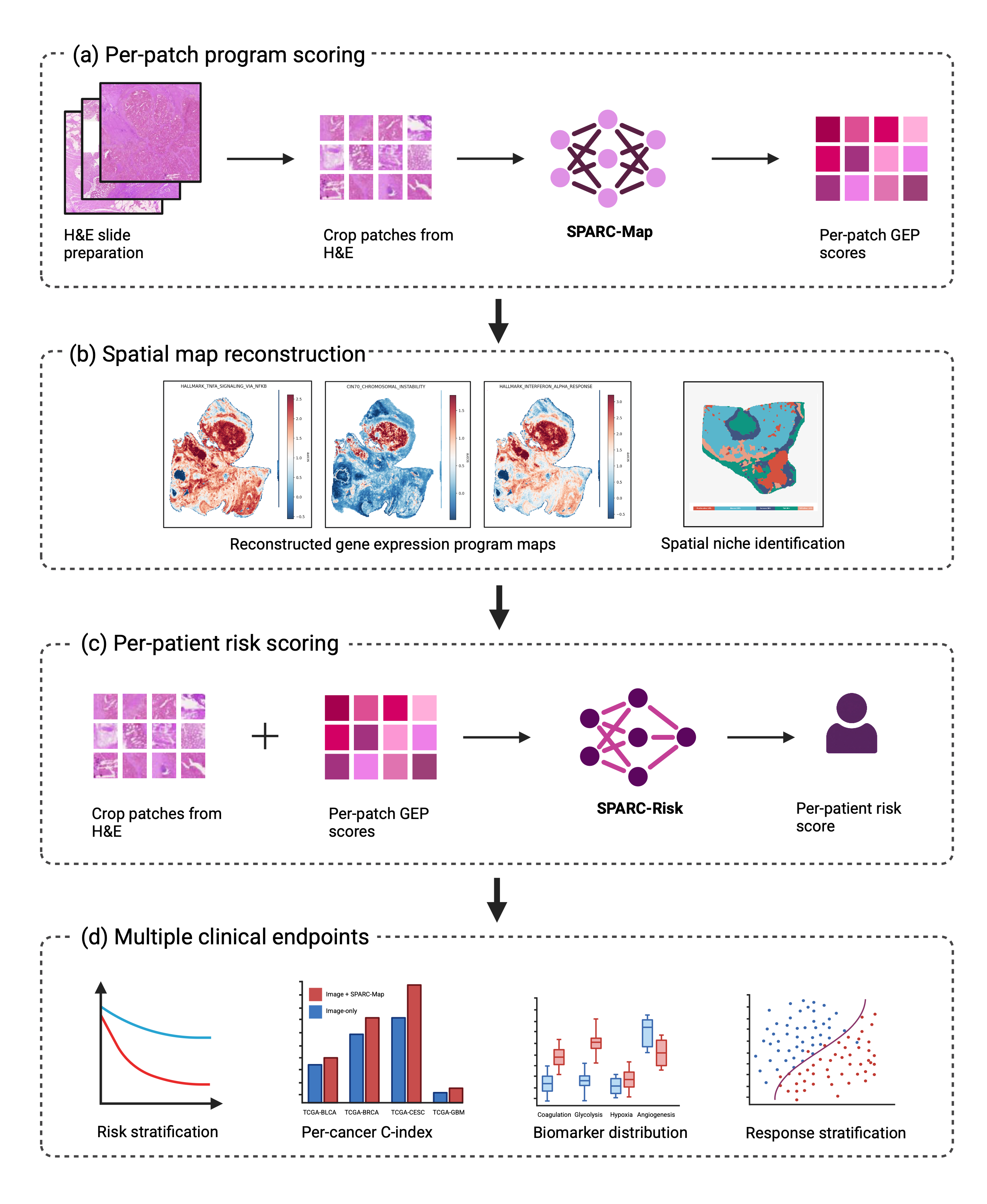
SPARC is a two-stage pipeline:
- SPARC-Map predicts 40 hallmark gene-expression-program (GEP) scores per H&E patch, recovering a spatial molecular map of each slide.
- SPARC-Risk fuses those per-patch GEP scores with the same H&E features through a signature-query attention head and a cancer-aware gate, producing a single per-patient risk score.
These weights cover the SPARC-Risk model and the image-only baseline used for ablations.
What you get
| Folder | Model | Description |
|---|---|---|
sparc_risk/ |
SPARC-Risk (canonical) | Signature-query fusion + H&E. The model reported throughout the paper. |
image_only/ |
Image-only baseline | Same backbone, GEP pathway disabled. Use for direct ablation against SPARC-Risk. |
Each folder contains 5 checkpoints — fold_0_best.pt through
fold_4_best.pt — corresponding to the 5-fold cross-validation splits
described in the paper and in
data/mmp_hybrid_splits_v2_20cancer.csv.
Every .pt carries both model_state_dict and the original training
config, so the model can be rebuilt with one line:
import torch
from sparc.models.factory import build_model
ckpt = torch.load("sparc_risk/fold_0_best.pt", map_location="cpu", weights_only=False)
model = build_model(ckpt["config"])
model.load_state_dict(ckpt["model_state_dict"])
model.eval()
Quick start
# 1. Install the SPARC package
git clone https://github.com/aziz-ayed/SPARC.git && cd SPARC
conda env create -f environment.yml
conda activate sparc
# 2. Accept the license on https://huggingface.co/azizayed/SPARC, then:
pip install -U "huggingface_hub[cli]"
hf auth login
hf download azizayed/SPARC --local-dir checkpoints
# 3. Inference on an external cohort (e.g. NLST lung)
python -m inference.run \
--cohort nlst \
--checkpoint_dir checkpoints/sparc_risk \
--gpus 0,1,2,3
The download produces:
checkpoints/
├── sparc_risk/ fold_{0..4}_best.pt
└── image_only/ fold_{0..4}_best.pt
Architecture (SPARC-Risk)
| Component | Setting |
|---|---|
| Image backbone | H-optimus-1 (1536-dim) |
| Patch size / magnification | 224 px @ 20× |
| Max patches per slide | 4096 |
| Fusion | Signature-query cross-attention (64-NN, 4 heads) |
| Aggregator | Gated attention MIL |
| Head | Discrete-time NLL survival, 4 bins |
| Cancer conditioning | Per-cancer learned gate |
| Hidden dim | 256 |
| Trainable params | ≈ 2.6 M |
| Optimiser / schedule | Adam, lr 1 × 10⁻⁴, cosine T_max 20 |
| Random seed | 1337 |
Full config + reproduction recipe: configs/sparc_risk.yaml.
Training data
5-fold patient-level cross-validation over 20 TCGA cancer types
(BLCA, BRCA, CESC, COAD, ESCA, GBM, HNSC, KIRC, KIRP, LGG, LIHC, LUAD,
LUSC, PAAD, READ, SARC, SKCM, STAD, UCEC, plus a held-out evaluation
split). Splits derive from the MMP hybrid scheme of Mahmood et al. and
are released alongside the code at
data/mmp_hybrid_splits_v2_20cancer.csv.
External validation cohorts (not used for training) — NLST lung, SurGen CRC, Yale breast, ovarian — are described in the paper.
Intended use
These weights are intended for non-commercial biomedical research and education only. Acceptable uses include:
- Reproducing the SPARC paper's results.
- Benchmarking against SPARC-Risk in computational-pathology research.
- Methodological extensions (new fusion designs, additional cohorts, ablation studies).
Citation
The SPARC paper is currently under review. Once a preprint or accepted version is available, a BibTeX entry will be added here. In the meantime, if you use these weights, please link back to github.com/aziz-ayed/SPARC and contact the corresponding author at azizayed@mit.edu.
License
These weights are released under the Creative Commons Attribution-NonCommercial 4.0 International License (CC BY-NC 4.0). For commercial licensing, please contact the authors via the corresponding GitHub issues page.