repo_id
stringlengths 15
89
| file_path
stringlengths 27
180
| content
stringlengths 1
2.23M
| __index_level_0__
int64 0
0
|
|---|---|---|---|
hf_public_repos/trl/docs
|
hf_public_repos/trl/docs/source/best_of_n.mdx
|
# Best of N sampling: Alternative ways to get better model output without RL based fine-tuning
Within the extras module is the `best-of-n` sampler class that serves as an alternative method of generating better model output.
As to how it fares against the RL based fine-tuning, please look in the `examples` directory for a comparison example
## Usage
To get started quickly, instantiate an instance of the class with a model, a length sampler, a tokenizer and a callable that serves as a proxy reward pipeline that outputs reward scores for input queries
```python
from transformers import pipeline, AutoTokenizer
from trl import AutoModelForCausalLMWithValueHead
from trl.core import LengthSampler
from trl.extras import BestOfNSampler
ref_model = AutoModelForCausalLMWithValueHead.from_pretrained(ref_model_name)
reward_pipe = pipeline("sentiment-analysis", model=reward_model, device=device)
tokenizer = AutoTokenizer.from_pretrained(ref_model_name)
tokenizer.pad_token = tokenizer.eos_token
# callable that takes a list of raw text and returns a list of corresponding reward scores
def queries_to_scores(list_of_strings):
return [output["score"] for output in reward_pipe(list_of_strings)]
best_of_n = BestOfNSampler(model, tokenizer, queries_to_scores, length_sampler=output_length_sampler)
```
And assuming you have a list/tensor of tokenized queries, you can generate better output by calling the `generate` method
```python
best_of_n.generate(query_tensors, device=device, **gen_kwargs)
```
The default sample size is 4, but you can change it at the time of instance initialization like so
```python
best_of_n = BestOfNSampler(model, tokenizer, queries_to_scores, length_sampler=output_length_sampler, sample_size=8)
```
The default output is the result of taking the top scored output for each query, but you can change it to top 2 and so on by passing the `n_candidates` argument at the time of instance initialization
```python
best_of_n = BestOfNSampler(model, tokenizer, queries_to_scores, length_sampler=output_length_sampler, n_candidates=2)
```
There is the option of setting the generation settings (like `temperature`, `pad_token_id`) at the time of instance creation as opposed to when calling the `generate` method.
This is done by passing a `GenerationConfig` from the `transformers` library at the time of initialization
```python
from transformers import GenerationConfig
generation_config = GenerationConfig(min_length= -1, top_k=0.0, top_p= 1.0, do_sample= True, pad_token_id=tokenizer.eos_token_id)
best_of_n = BestOfNSampler(model, tokenizer, queries_to_scores, length_sampler=output_length_sampler, generation_config=generation_config)
best_of_n.generate(query_tensors, device=device)
```
Furthermore, at the time of initialization you can set the seed to control repeatability of the generation process and the number of samples to generate for each query
| 0
|
hf_public_repos/trl/docs
|
hf_public_repos/trl/docs/source/lora_tuning_peft.mdx
|
# Examples of using peft with trl to finetune 8-bit models with Low Rank Adaption (LoRA)
The notebooks and scripts in this examples show how to use Low Rank Adaptation (LoRA) to fine-tune models in a memory efficient manner. Most of PEFT methods supported in peft library but note that some PEFT methods such as Prompt tuning are not supported.
For more information on LoRA, see the [original paper](https://arxiv.org/abs/2106.09685).
Here's an overview of the `peft`-enabled notebooks and scripts in the [trl repository](https://github.com/huggingface/trl/tree/main/examples):
| File | Task | Description | Colab link |
|---|---| --- |
| [`stack_llama/rl_training.py`](https://github.com/huggingface/trl/blob/main/examples/research_projects/stack_llama/scripts/rl_training.py) | RLHF | Distributed fine-tuning of the 7b parameter LLaMA models with a learned reward model and `peft`. | |
| [`stack_llama/reward_modeling.py`](https://github.com/huggingface/trl/blob/main/examples/research_projects/stack_llama/scripts/reward_modeling.py) | Reward Modeling | Distributed training of the 7b parameter LLaMA reward model with `peft`. | |
| [`stack_llama/supervised_finetuning.py`](https://github.com/huggingface/trl/blob/main/examples/research_projects/stack_llama/scripts/supervised_finetuning.py) | SFT | Distributed instruction/supervised fine-tuning of the 7b parameter LLaMA model with `peft`. | |
## Installation
Note: peft is in active development, so we install directly from their Github page.
Peft also relies on the latest version of transformers.
```bash
pip install trl[peft]
pip install bitsandbytes loralib
pip install git+https://github.com/huggingface/transformers.git@main
#optional: wandb
pip install wandb
```
Note: if you don't want to log with `wandb` remove `log_with="wandb"` in the scripts/notebooks. You can also replace it with your favourite experiment tracker that's [supported by `accelerate`](https://huggingface.co/docs/accelerate/usage_guides/tracking).
## How to use it?
Simply declare a `PeftConfig` object in your script and pass it through `.from_pretrained` to load the TRL+PEFT model.
```python
from peft import LoraConfig
from trl import AutoModelForCausalLMWithValueHead
model_id = "edbeeching/gpt-neo-125M-imdb"
lora_config = LoraConfig(
r=16,
lora_alpha=32,
lora_dropout=0.05,
bias="none",
task_type="CAUSAL_LM",
)
model = AutoModelForCausalLMWithValueHead.from_pretrained(
model_id,
peft_config=lora_config,
)
```
And if you want to load your model in 8bit precision:
```python
pretrained_model = AutoModelForCausalLMWithValueHead.from_pretrained(
config.model_name,
load_in_8bit=True,
peft_config=lora_config,
)
```
... or in 4bit precision:
```python
pretrained_model = AutoModelForCausalLMWithValueHead.from_pretrained(
config.model_name,
peft_config=lora_config,
load_in_4bit=True,
)
```
## Launch scripts
The `trl` library is powered by `accelerate`. As such it is best to configure and launch trainings with the following commands:
```bash
accelerate config # will prompt you to define the training configuration
accelerate launch scripts/gpt2-sentiment_peft.py # launches training
```
## Using `trl` + `peft` and Data Parallelism
You can scale up to as many GPUs as you want, as long as you are able to fit the training process in a single device. The only tweak you need to apply is to load the model as follows:
```python
from peft import LoraConfig
...
lora_config = LoraConfig(
r=16,
lora_alpha=32,
lora_dropout=0.05,
bias="none",
task_type="CAUSAL_LM",
)
pretrained_model = AutoModelForCausalLMWithValueHead.from_pretrained(
config.model_name,
peft_config=lora_config,
)
```
And if you want to load your model in 8bit precision:
```python
pretrained_model = AutoModelForCausalLMWithValueHead.from_pretrained(
config.model_name,
peft_config=lora_config,
load_in_8bit=True,
)
```
... or in 4bit precision:
```python
pretrained_model = AutoModelForCausalLMWithValueHead.from_pretrained(
config.model_name,
peft_config=lora_config,
load_in_4bit=True,
)
```
Finally, make sure that the rewards are computed on correct device as well, for that you can use `ppo_trainer.model.current_device`.
## Naive pipeline parallelism (NPP) for large models (>60B models)
The `trl` library also supports naive pipeline parallelism (NPP) for large models (>60B models). This is a simple way to parallelize the model across multiple GPUs.
This paradigm, termed as "Naive Pipeline Parallelism" (NPP) is a simple way to parallelize the model across multiple GPUs. We load the model and the adapters across multiple GPUs and the activations and gradients will be naively communicated across the GPUs. This supports `int8` models as well as other `dtype` models.
<div style="text-align: center">
<img src="https://huggingface.co/datasets/trl-internal-testing/example-images/resolve/main/images/trl-npp.png">
</div>
### How to use NPP?
Simply load your model with a custom `device_map` argument on the `from_pretrained` to split your model across multiple devices. Check out this [nice tutorial](https://github.com/huggingface/blog/blob/main/accelerate-large-models.md) on how to properly create a `device_map` for your model.
Also make sure to have the `lm_head` module on the first GPU device as it may throw an error if it is not on the first device. As this time of writing, you need to install the `main` branch of `accelerate`: `pip install git+https://github.com/huggingface/accelerate.git@main` and `peft`: `pip install git+https://github.com/huggingface/peft.git@main`.
### Launch scripts
Although `trl` library is powered by `accelerate`, you should run your training script in a single process. Note that we do not support Data Parallelism together with NPP yet.
```bash
python PATH_TO_SCRIPT
```
## Fine-tuning Llama-2 model
You can easily fine-tune Llama2 model using `SFTTrainer` and the official script! For example to fine-tune llama2-7b on the Guanaco dataset, run (tested on a single NVIDIA T4-16GB):
```bash
python examples/scripts/sft.py --model_name meta-llama/Llama-2-7b-hf --dataset_name timdettmers/openassistant-guanaco --load_in_4bit --use_peft --batch_size 4 --gradient_accumulation_steps 2
```
| 0
|
hf_public_repos/trl/docs
|
hf_public_repos/trl/docs/source/installation.mdx
|
# Installation
You can install TRL either from pypi or from source:
## pypi
Install the library with pip:
```bash
pip install trl
```
### Source
You can also install the latest version from source. First clone the repo and then run the installation with `pip`:
```bash
git clone https://github.com/huggingface/trl.git
cd trl/
pip install -e .
```
If you want the development install you can replace the pip install with the following:
```bash
pip install -e ".[dev]"
```
| 0
|
hf_public_repos/trl/docs
|
hf_public_repos/trl/docs/source/iterative_sft_trainer.mdx
|
# Iterative Trainer
Iterative fine-tuning is a training method that enables to perform custom actions (generation and filtering for example) between optimization steps. In TRL we provide an easy-to-use API to fine-tune your models in an iterative way in just a few lines of code.
## Usage
To get started quickly, instantiate an instance a model, and a tokenizer.
```python
model = AutoModelForCausalLM.from_pretrained(model_name)
tokenizer = AutoTokenizer.from_pretrained(model_name)
if tokenizer.pad_token is None:
tokenizer.pad_token = tokenizer.eos_token
trainer = IterativeSFTTrainer(
model,
tokenizer
)
```
You have the choice to either provide a list of strings or a list of tensors to the step function.
#### Using a list of tensors as input:
```python
inputs = {
"input_ids": input_ids,
"attention_mask": attention_mask
}
trainer.step(**inputs)
```
#### Using a list of strings as input:
```python
inputs = {
"texts": texts
}
trainer.step(**inputs)
```
For causal language models, labels will automatically be created from input_ids or from texts. When using sequence to sequence models you will have to provide your own labels or text_labels.
## IterativeTrainer
[[autodoc]] IterativeSFTTrainer
| 0
|
hf_public_repos/trl/docs
|
hf_public_repos/trl/docs/source/learning_tools.mdx
|
# Learning Tools (Experimental 🧪)
Using Large Language Models (LLMs) with tools has been a popular topic recently with awesome works such as [ToolFormer](https://arxiv.org/abs/2302.04761) and [ToolBench](https://arxiv.org/pdf/2305.16504.pdf). In TRL, we provide a simple example of how to teach LLM to use tools with reinforcement learning.
Here's an overview of the scripts in the [trl repository](https://github.com/lvwerra/trl/tree/main/examples/research_projects/tools):
| File | Description |
|---|---|
| [`calculator.py`](https://github.com/lvwerra/trl/blob/main/examples/research_projects/tools/calculator.py) | Script to train LLM to use a calculator with reinforcement learning. |
| [`triviaqa.py`](https://github.com/lvwerra/trl/blob/main/examples/research_projects/tools/triviaqa.py) | Script to train LLM to use a wiki tool to answer questions. |
| [`python_interpreter.py`](https://github.com/lvwerra/trl/blob/main/examples/research_projects/tools/python_interpreter.py) | Script to train LLM to use python interpreter to solve math puzzles. |
<Tip warning={true}>
Note that the scripts above rely heavily on the `TextEnvironment` API which is still under active development. The API may change in the future. Please see [`TextEnvironment`](text_environment) for the related docs.
</Tip>
## Learning to Use a Calculator
The rough idea is as follows:
1. Load a tool such as [ybelkada/simple-calculator](https://huggingface.co/spaces/ybelkada/simple-calculator) that parse a text calculation like `"14 + 34"` and return the calulated number:
```python
from transformers import AutoTokenizer, load_tool
tool = load_tool("ybelkada/simple-calculator")
tool_fn = lambda text: str(round(float(tool(text)), 2)) # rounding to 2 decimal places
```
1. Define a reward function that returns a positive reward if the tool returns the correct answer. In the script we create a dummy reward function like `reward_fn = lambda x: 1`, but we override the rewards directly later.
1. Create a prompt on how to use the tools
```python
# system prompt
prompt = """\
What is 13.1-3?
<request><SimpleCalculatorTool>13.1-3<call>10.1<response>
Result=10.1<submit>
What is 4*3?
<request><SimpleCalculatorTool>4*3<call>12<response>
Result=12<submit>
What is 12.1+1?
<request><SimpleCalculatorTool>12.1+1<call>13.1<response>
Result=13.1<submit>
What is 12.1-20?
<request><SimpleCalculatorTool>12.1-20<call>-7.9<response>
Result=-7.9<submit>"""
```
3. Create a `trl.TextEnvironment` with the model
```python
env = TextEnvironment(
model,
tokenizer,
{"SimpleCalculatorTool": tool_fn},
reward_fn,
prompt,
generation_kwargs=generation_kwargs,
)
```
4. Then generate some data such as `tasks = ["\n\nWhat is 13.1-3?", "\n\nWhat is 4*3?"]` and run the environment with `queries, responses, masks, rewards, histories = env.run(tasks)`. The environment will look for the `<call>` token in the prompt and append the tool output to the response; it will also return the mask associated with the response. You can further use the `histories` to visualize the interaction between the model and the tool; `histories[0].show_text()` will show the text with color-coded tool output and `histories[0].show_tokens(tokenizer)` will show visualize the tokens.

1. Finally, we can train the model with `train_stats = ppo_trainer.step(queries, responses, rewards, masks)`. The trainer will use the mask to ignore the tool output when computing the loss, make sure to pass that argument to `step`.
## Experiment results
We trained a model with the above script for 10 random seeds. You can reproduce the run with the following command. Feel free to remove the `--slurm-*` arguments if you don't have access to a slurm cluster.
```
WANDB_TAGS="calculator_final" python benchmark/benchmark.py \
--command "python examples/research_projects/tools/calculator.py" \
--num-seeds 10 \
--start-seed 1 \
--workers 10 \
--slurm-gpus-per-task 1 \
--slurm-ntasks 1 \
--slurm-total-cpus 8 \
--slurm-template-path benchmark/trl.slurm_template
```
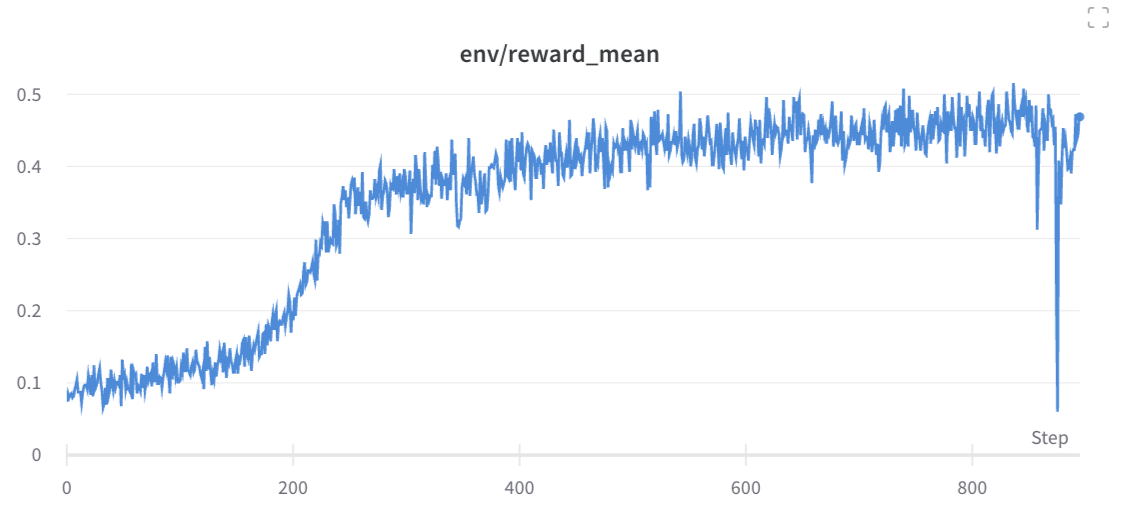
We can then use [`openrlbenchmark`](https://github.com/openrlbenchmark/openrlbenchmark) which generates the following plot.
```
python -m openrlbenchmark.rlops_multi_metrics \
--filters '?we=openrlbenchmark&wpn=trl&xaxis=_step&ceik=trl_ppo_trainer_config.value.tracker_project_name&cen=trl_ppo_trainer_config.value.log_with&metrics=env/reward_mean&metrics=objective/kl' \
'wandb?tag=calculator_final&cl=calculator_mask' \
--env-ids trl \
--check-empty-runs \
--pc.ncols 2 \
--pc.ncols-legend 1 \
--output-filename static/0compare \
--scan-history
```
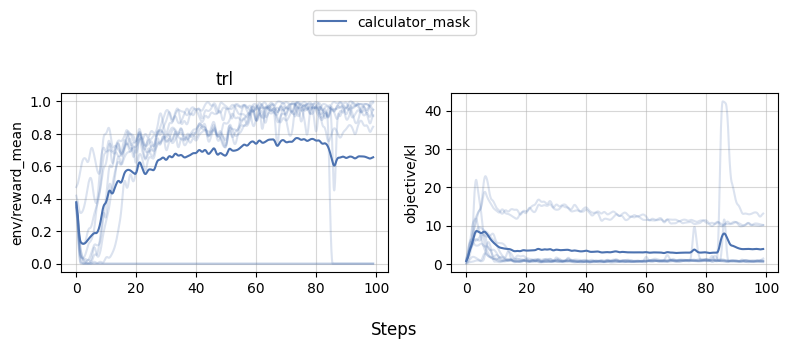
As we can see, while 1-2 experiments crashed for some reason, most of the runs obtained near perfect proficiency in the calculator task.
## (Early Experiments 🧪): learning to use a wiki tool for question answering
In the [ToolFormer](https://arxiv.org/abs/2302.04761) paper, it shows an interesting use case that utilizes a Wikipedia Search tool to help answer questions. In this section, we attempt to perform similar experiments but uses RL instead to teach the model to use a wiki tool on the [TriviaQA](https://nlp.cs.washington.edu/triviaqa/) dataset.
<Tip warning={true}>
**Note that many settings are different so the results are not directly comparable.**
</Tip>
### Building a search index
Since [ToolFormer](https://arxiv.org/abs/2302.04761) did not open source, we needed to first replicate the search index. It is mentioned in their paper that the authors built the search index using a BM25 retriever that indexes the Wikipedia dump from [KILT](https://github.com/facebookresearch/KILT)
Fortunately, [`pyserini`](https://github.com/castorini/pyserini) already implements the BM25 retriever and provides a prebuilt index for the KILT Wikipedia dump. We can use the following code to search the index.
```python
from pyserini.search.lucene import LuceneSearcher
import json
searcher = LuceneSearcher.from_prebuilt_index('wikipedia-kilt-doc')
def search(query):
hits = searcher.search(query, k=1)
hit = hits[0]
contents = json.loads(hit.raw)['contents']
return contents
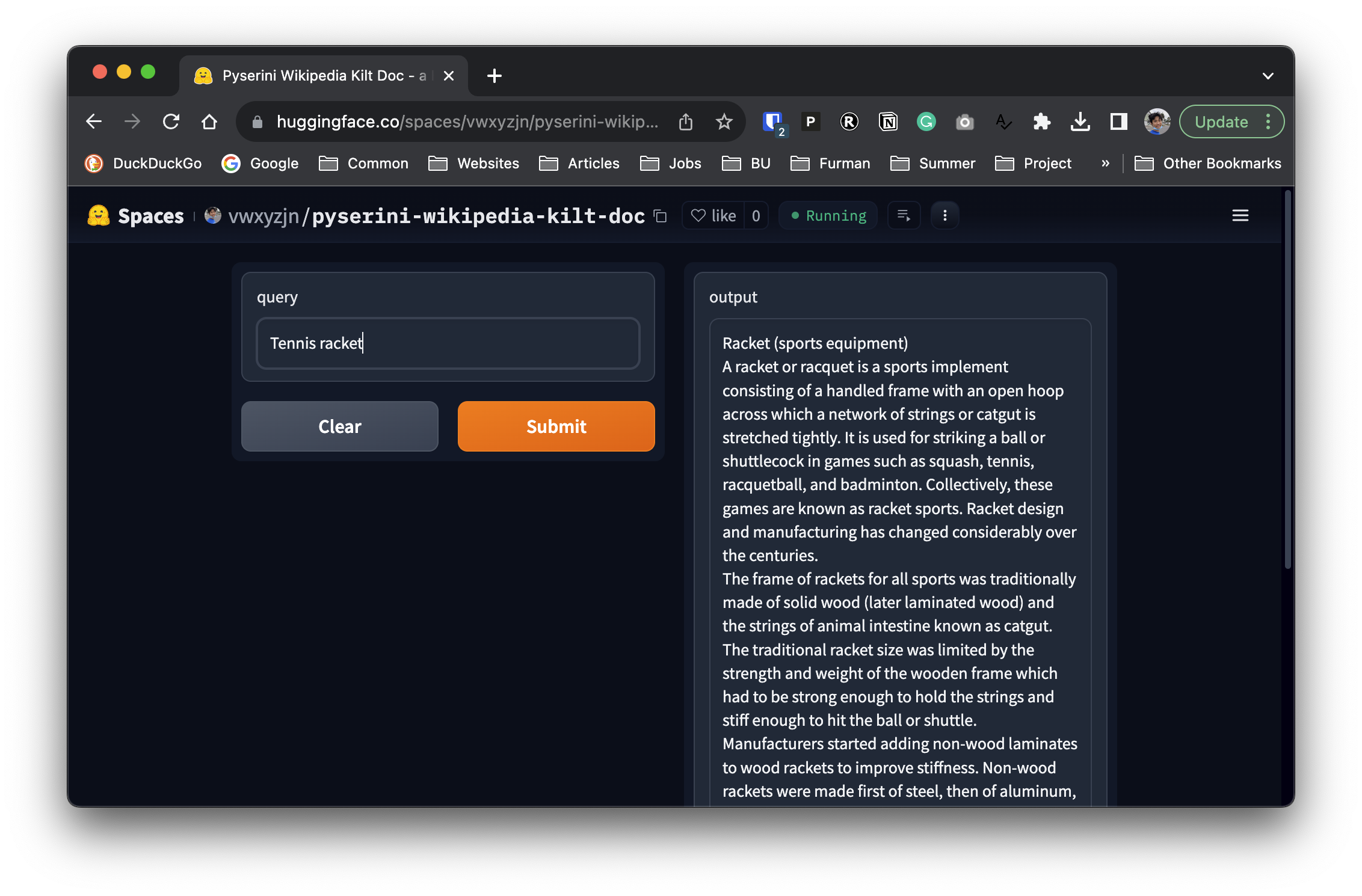
print(search("tennis racket"))
```
```
Racket (sports equipment)
A racket or racquet is a sports implement consisting of a handled frame with an open hoop across which a network of strings or catgut is stretched tightly. It is used for striking a ball or shuttlecock in games such as squash, tennis, racquetball, and badminton. Collectively, these games are known as racket sports. Racket design and manufacturing has changed considerably over the centuries.
The frame of rackets for all sports was traditionally made of solid wood (later laminated wood) and the strings of animal intestine known as catgut. The traditional racket size was limited by the strength and weight of the wooden frame which had to be strong enough to hold the strings and stiff enough to hit the ball or shuttle. Manufacturers started adding non-wood laminates to wood rackets to improve stiffness. Non-wood rackets were made first of steel, then of aluminum, and then carbon fiber composites. Wood is still used for real tennis, rackets, and xare. Most rackets are now made of composite materials including carbon fiber or fiberglass, metals such as titanium alloys, or ceramics.
...
```
We then basically deployed this snippet as a Hugging Face space [here](https://huggingface.co/spaces/vwxyzjn/pyserini-wikipedia-kilt-doc), so that we can use the space as a `transformers.Tool` later.
### Experiment settings
We use the following settings:
* use the `bigcode/starcoderbase` model as the base model
* use the `pyserini-wikipedia-kilt-doc` space as the wiki tool and only uses the first paragrahs of the search result, allowing the `TextEnvironment` to obtain at most `max_tool_reponse=400` response tokens from the tool.
* test if the response contain the answer string, if so, give a reward of 1, otherwise, give a reward of 0.
* notice this is a simplified evaluation criteria. In [ToolFormer](https://arxiv.org/abs/2302.04761), the authors checks if the first 20 words of the response contain the correct answer.
* used the following prompt that demonstrates the usage of the wiki tool.
```python
prompt = """\
Answer the following question:
Q: In which branch of the arts is Patricia Neary famous?
A: Ballets
A2: <request><Wiki>Patricia Neary<call>Patricia Neary (born October 27, 1942) is an American ballerina, choreographer and ballet director, who has been particularly active in Switzerland. She has also been a highly successful ambassador for the Balanchine Trust, bringing George Balanchine's ballets to 60 cities around the globe.<response>
Result=Ballets<submit>
Q: Who won Super Bowl XX?
A: Chicago Bears
A2: <request><Wiki>Super Bowl XX<call>Super Bowl XX was an American football game between the National Football Conference (NFC) champion Chicago Bears and the American Football Conference (AFC) champion New England Patriots to decide the National Football League (NFL) champion for the 1985 season. The Bears defeated the Patriots by the score of 46–10, capturing their first NFL championship (and Chicago's first overall sports victory) since 1963, three years prior to the birth of the Super Bowl. Super Bowl XX was played on January 26, 1986 at the Louisiana Superdome in New Orleans.<response>
Result=Chicago Bears<submit>
Q: """
```
### Result and Discussion
Our experiments show that the agent can learn to use the wiki tool to answer questions. The learning curves would go up mostly, but one of the experiment did crash.
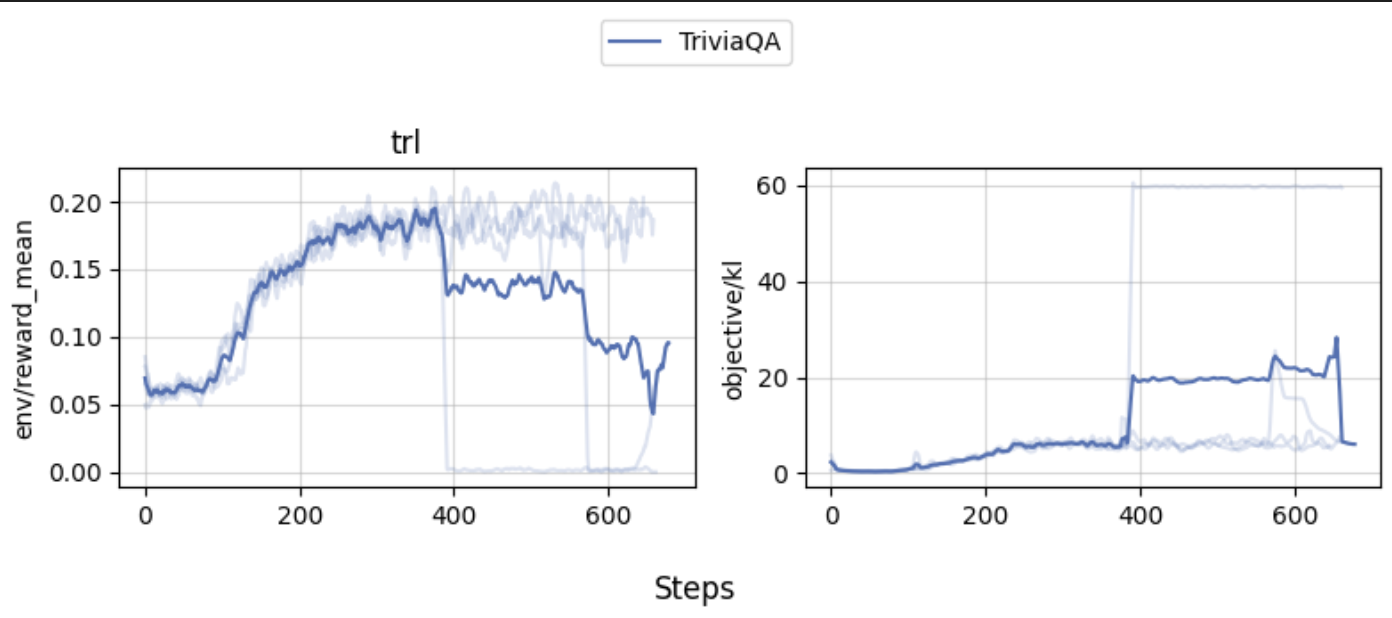
Wandb report is [here](https://wandb.ai/costa-huang/cleanRL/reports/TriviaQA-Final-Experiments--Vmlldzo1MjY0ODk5) for further inspection.
Note that the correct rate of the trained model is on the low end, which could be due to the following reasons:
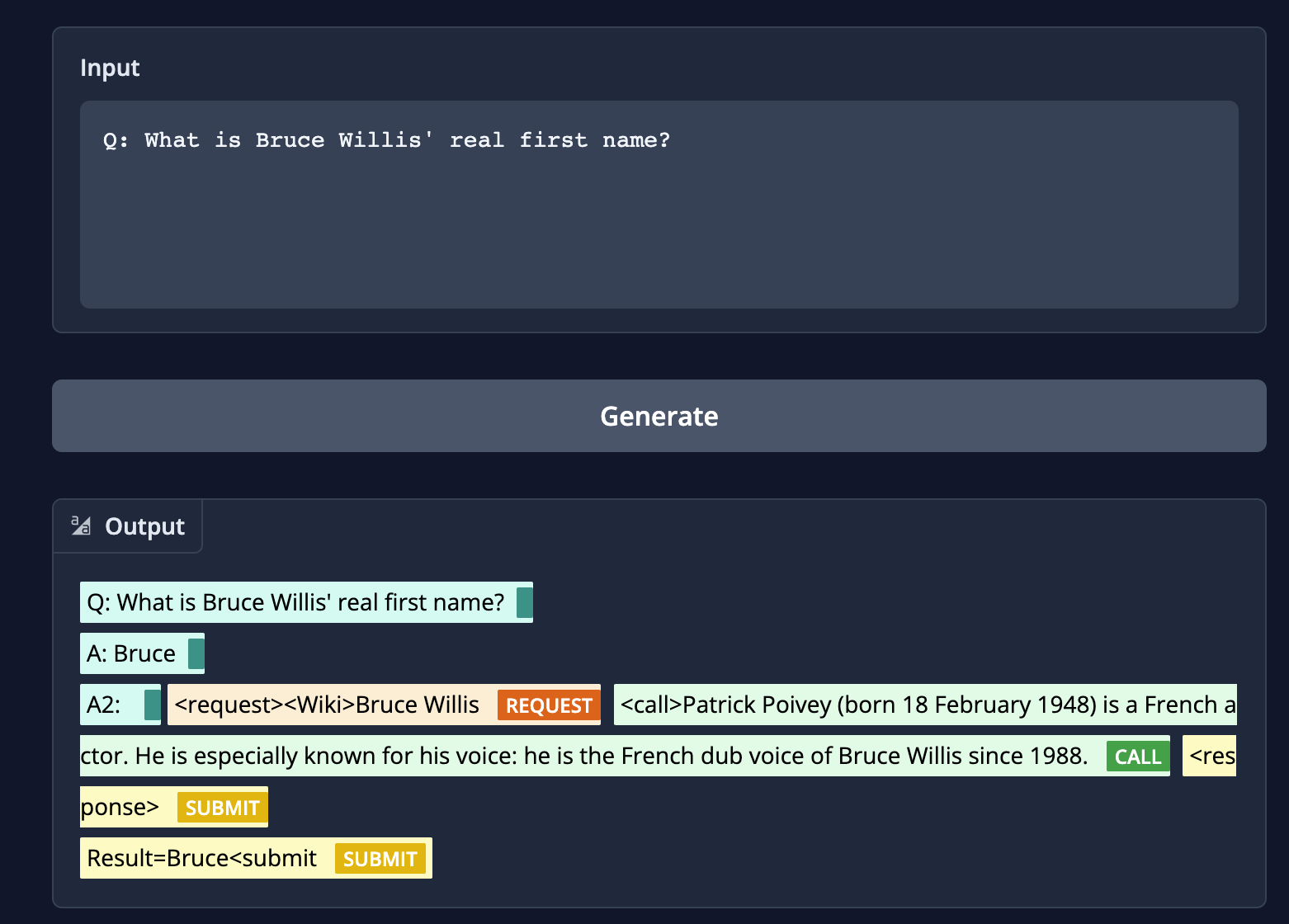
* **incorrect searches:** When given the question `"What is Bruce Willis' real first name?"` if the model searches for `Bruce Willis`, our wiki tool returns "Patrick Poivey (born 18 February 1948) is a French actor. He is especially known for his voice: he is the French dub voice of Bruce Willis since 1988.` But a correct search should be `Walter Bruce Willis (born March 19, 1955) is an American former actor. He achieved fame with a leading role on the comedy-drama series Moonlighting (1985–1989) and appeared in over a hundred films, gaining recognition as an action hero after his portrayal of John McClane in the Die Hard franchise (1988–2013) and other roles.[1][2]"
* **unnecessarily long response**: The wiki tool by default sometimes output very long sequences. E.g., when the wiki tool searches for "Brown Act"
* Our wiki tool returns "The Ralph M. Brown Act, located at California Government Code 54950 "et seq.", is an act of the California State Legislature, authored by Assemblymember Ralph M. Brown and passed in 1953, that guarantees the public's right to attend and participate in meetings of local legislative bodies."
* [ToolFormer](https://arxiv.org/abs/2302.04761)'s wiki tool returns "The Ralph M. Brown Act is an act of the California State Legislature that guarantees the public's right to attend and participate in meetings of local legislative bodies." which is more succinct.

## (Early Experiments 🧪): solving math puzzles with python interpreter
In this section, we attempt to teach the model to use a python interpreter to solve math puzzles. The rough idea is to give the agent a prompt like the following:
```python
prompt = """\
Example of using a Python API to solve math questions.
Q: Olivia has $23. She bought five bagels for $3 each. How much money does she have left?
<request><PythonInterpreter>
def solution():
money_initial = 23
bagels = 5
bagel_cost = 3
money_spent = bagels * bagel_cost
money_left = money_initial - money_spent
result = money_left
return result
print(solution())
<call>72<response>
Result = 72 <submit>
Q: """
```
Training experiment can be found at https://wandb.ai/lvwerra/trl-gsm8k/runs/a5odv01y
| 0
|
hf_public_repos/trl/docs
|
hf_public_repos/trl/docs/source/models.mdx
|
# Models
With the `AutoModelForCausalLMWithValueHead` class TRL supports all decoder model architectures in transformers such as GPT-2, OPT, and GPT-Neo. In addition, with `AutoModelForSeq2SeqLMWithValueHead` you can use encoder-decoder architectures such as T5. TRL also requires reference models which are frozen copies of the model that is trained. With `create_reference_model` you can easily create a frozen copy and also share layers between the two models to save memory.
## PreTrainedModelWrapper
[[autodoc]] PreTrainedModelWrapper
## AutoModelForCausalLMWithValueHead
[[autodoc]] AutoModelForCausalLMWithValueHead
- __init__
- forward
- generate
- _init_weights
## AutoModelForSeq2SeqLMWithValueHead
[[autodoc]] AutoModelForSeq2SeqLMWithValueHead
- __init__
- forward
- generate
- _init_weights
## create_reference_model
[[autodoc]] create_reference_model
| 0
|
hf_public_repos/trl/docs
|
hf_public_repos/trl/docs/source/use_model.md
|
# Use model after training
Once you have trained a model using either the SFTTrainer, PPOTrainer, or DPOTrainer, you will have a fine-tuned model that can be used for text generation. In this section, we'll walk through the process of loading the fine-tuned model and generating text. If you need to run an inference server with the trained model, you can explore libraries such as [`text-generation-inference`](https://github.com/huggingface/text-generation-inference).
## Load and Generate
If you have fine-tuned a model fully, meaning without the use of PEFT you can simply load it like any other language model in transformers. E.g. the value head that was trained during the PPO training is no longer needed and if you load the model with the original transformer class it will be ignored:
```python
from transformers import AutoTokenizer, AutoModelForCausalLM
model_name_or_path = "kashif/stack-llama-2" #path/to/your/model/or/name/on/hub
device = "cpu" # or "cuda" if you have a GPU
model = AutoModelForCausalLM.from_pretrained(model_name_or_path).to(device)
tokenizer = AutoTokenizer.from_pretrained(model_name_or_path)
inputs = tokenizer.encode("This movie was really", return_tensors="pt").to(device)
outputs = model.generate(inputs)
print(tokenizer.decode(outputs[0]))
```
Alternatively you can also use the pipeline:
```python
from transformers import pipeline
model_name_or_path = "kashif/stack-llama-2" #path/to/your/model/or/name/on/hub
pipe = pipeline("text-generation", model=model_name_or_path)
print(pipe("This movie was really")[0]["generated_text"])
```
## Use Adapters PEFT
```python
from peft import PeftConfig, PeftModel
from transformers import AutoModelForCausalLM, AutoTokenizer
base_model_name = "kashif/stack-llama-2" #path/to/your/model/or/name/on/hub"
adapter_model_name = "path/to/my/adapter"
model = AutoModelForCausalLM.from_pretrained(base_model_name)
model = PeftModel.from_pretrained(model, adapter_model_name)
tokenizer = AutoTokenizer.from_pretrained(base_model_name)
```
You can also merge the adapters into the base model so you can use the model like a normal transformers model, however the checkpoint will be significantly bigger:
```python
model = AutoModelForCausalLM.from_pretrained(base_model_name)
model = PeftModel.from_pretrained(model, adapter_model_name)
model = model.merge_and_unload()
model.save_pretrained("merged_adapters")
```
Once you have the model loaded and either merged the adapters or keep them separately on top you can run generation as with a normal model outlined above.
| 0
|
hf_public_repos/trl/docs
|
hf_public_repos/trl/docs/source/reward_trainer.mdx
|
# Reward Modeling
TRL supports custom reward modeling for anyone to perform reward modeling on their dataset and model.
Check out a complete flexible example at [`examples/scripts/reward_modeling.py`](https://github.com/huggingface/trl/tree/main/examples/scripts/reward_modeling.py).
## Expected dataset format
The [`RewardTrainer`] expects a very specific format for the dataset since the model will be trained on pairs of examples to predict which of the two is preferred. We provide an example from the [`Anthropic/hh-rlhf`](https://huggingface.co/datasets/Anthropic/hh-rlhf) dataset below:
<div style="text-align: center">
<img src="https://huggingface.co/datasets/trl-internal-testing/example-images/resolve/main/images/rlhf-antropic-example.png", width="50%">
</div>
Therefore the final dataset object should contain two 4 entries at least if you use the default [`RewardDataCollatorWithPadding`] data collator. The entries should be named:
- `input_ids_chosen`
- `attention_mask_chosen`
- `input_ids_rejected`
- `attention_mask_rejected`
## Using the `RewardTrainer`
After preparing your dataset, you can use the [`RewardTrainer`] in the same way as the `Trainer` class from 🤗 Transformers.
You should pass an `AutoModelForSequenceClassification` model to the [`RewardTrainer`], along with a [`RewardConfig`] which configures the hyperparameters of the training.
### Leveraging 🤗 PEFT to train a reward model
Just pass a `peft_config` in the keyword arguments of [`RewardTrainer`], and the trainer should automatically take care of converting the model into a PEFT model!
```python
from peft import LoraConfig, task_type
from transformers import AutoModelForSequenceClassification, AutoTokenizer
from trl import RewardTrainer, RewardConfig
model = AutoModelForSequenceClassification.from_pretrained("gpt2")
peft_config = LoraConfig(
task_type=TaskType.SEQ_CLS,
inference_mode=False,
r=8,
lora_alpha=32,
lora_dropout=0.1,
)
...
trainer = RewardTrainer(
model=model,
args=training_args,
tokenizer=tokenizer,
train_dataset=dataset,
peft_config=peft_config,
)
trainer.train()
```
### Adding a margin to the loss
As in the [Llama 2 paper](https://huggingface.co/papers/2307.09288), you can add a margin to the loss by adding a `margin` column to the dataset. The reward collator will automatically pass it through and the loss will be computed accordingly.
```python
def add_margin(row):
# Assume you have a score_chosen and score_rejected columns that you want to use to compute the margin
return {'margin': row['score_chosen'] - row['score_rejected']}
dataset = dataset.map(add_margin)
```
## RewardConfig
[[autodoc]] RewardConfig
## RewardTrainer
[[autodoc]] RewardTrainer
| 0
|
hf_public_repos/trl/docs
|
hf_public_repos/trl/docs/source/index.mdx
|
<div style="text-align: center">
<img src="https://huggingface.co/datasets/trl-internal-testing/example-images/resolve/main/images/trl_banner_dark.png">
</div>
# TRL - Transformer Reinforcement Learning
TRL is a full stack library where we provide a set of tools to train transformer language models with Reinforcement Learning, from the Supervised Fine-tuning step (SFT), Reward Modeling step (RM) to the Proximal Policy Optimization (PPO) step.
The library is integrated with 🤗 [transformers](https://github.com/huggingface/transformers).
<div style="text-align: center">
<img src="https://huggingface.co/datasets/trl-internal-testing/example-images/resolve/main/images/TRL-readme.png">
</div>
Check the appropriate sections of the documentation depending on your needs:
## API documentation
- [Model Classes](models): *A brief overview of what each public model class does.*
- [`SFTTrainer`](sft_trainer): *Supervise Fine-tune your model easily with `SFTTrainer`*
- [`RewardTrainer`](reward_trainer): *Train easily your reward model using `RewardTrainer`.*
- [`PPOTrainer`](ppo_trainer): *Further fine-tune the supervised fine-tuned model using PPO algorithm*
- [Best-of-N Sampling](best-of-n): *Use best of n sampling as an alternative way to sample predictions from your active model*
- [`DPOTrainer`](dpo_trainer): *Direct Preference Optimization training using `DPOTrainer`.*
- [`TextEnvironment`](text_environment): *Text environment to train your model using tools with RL.*
## Examples
- [Sentiment Tuning](sentiment_tuning): *Fine tune your model to generate positive movie contents*
- [Training with PEFT](lora_tuning_peft): *Memory efficient RLHF training using adapters with PEFT*
- [Detoxifying LLMs](detoxifying_a_lm): *Detoxify your language model through RLHF*
- [StackLlama](using_llama_models): *End-to-end RLHF training of a Llama model on Stack exchange dataset*
- [Learning with Tools](learning_tools): *Walkthrough of using `TextEnvironments`*
- [Multi-Adapter Training](multi_adapter_rl): *Use a single base model and multiple adapters for memory efficient end-to-end training*
## Blog posts
<div class="mt-10">
<div class="w-full flex flex-col space-y-4 md:space-y-0 md:grid md:grid-cols-2 md:gap-y-4 md:gap-x-5">
<a class="!no-underline border dark:border-gray-700 p-5 rounded-lg shadow hover:shadow-lg" href="https://huggingface.co/blog/rlhf">
<img src="https://raw.githubusercontent.com/huggingface/blog/main/assets/120_rlhf/thumbnail.png" alt="thumbnail">
<p class="text-gray-700">Illustrating Reinforcement Learning from Human Feedback</p>
</a>
<a class="!no-underline border dark:border-gray-700 p-5 rounded-lg shadow hover:shadow-lg" href="https://huggingface.co/blog/trl-peft">
<img src="https://github.com/huggingface/blog/blob/main/assets/133_trl_peft/thumbnail.png?raw=true" alt="thumbnail">
<p class="text-gray-700">Fine-tuning 20B LLMs with RLHF on a 24GB consumer GPU</p>
</a>
<a class="!no-underline border dark:border-gray-700 p-5 rounded-lg shadow hover:shadow-lg" href="https://huggingface.co/blog/stackllama">
<img src="https://github.com/huggingface/blog/blob/main/assets/138_stackllama/thumbnail.png?raw=true" alt="thumbnail">
<p class="text-gray-700">StackLLaMA: A hands-on guide to train LLaMA with RLHF</p>
</a>
<a class="!no-underline border dark:border-gray-700 p-5 rounded-lg shadow hover:shadow-lg" href="https://huggingface.co/blog/dpo-trl">
<img src="https://github.com/huggingface/blog/blob/main/assets/157_dpo_trl/dpo_thumbnail.png?raw=true" alt="thumbnail">
<p class="text-gray-700">Fine-tune Llama 2 with DPO</p>
</a>
<a class="!no-underline border dark:border-gray-700 p-5 rounded-lg shadow hover:shadow-lg" href="https://huggingface.co/blog/trl-ddpo">
<img src="https://github.com/huggingface/blog/blob/main/assets/166_trl_ddpo/thumbnail.png?raw=true" alt="thumbnail">
<p class="text-gray-700">Finetune Stable Diffusion Models with DDPO via TRL</p>
</a>
</div>
</div>
| 0
|
hf_public_repos/trl/docs
|
hf_public_repos/trl/docs/source/trainer.mdx
|
# Trainer
At TRL we support PPO (Proximal Policy Optimisation) with an implementation that largely follows the structure introduced in the paper "Fine-Tuning Language Models from Human Preferences" by D. Ziegler et al. [[paper](https://arxiv.org/pdf/1909.08593.pdf), [code](https://github.com/openai/lm-human-preferences)].
The Trainer and model classes are largely inspired from `transformers.Trainer` and `transformers.AutoModel` classes and adapted for RL.
We also support a `RewardTrainer` that can be used to train a reward model.
## PPOConfig
[[autodoc]] PPOConfig
## PPOTrainer
[[autodoc]] PPOTrainer
## RewardConfig
[[autodoc]] RewardConfig
## RewardTrainer
[[autodoc]] RewardTrainer
## SFTTrainer
[[autodoc]] SFTTrainer
## DPOTrainer
[[autodoc]] DPOTrainer
## DDPOConfig
[[autodoc]] DDPOConfig
## DDPOTrainer
[[autodoc]] DDPOTrainer
## IterativeSFTTrainer
[[autodoc]] IterativeSFTTrainer
## set_seed
[[autodoc]] set_seed
| 0
|
hf_public_repos/trl/docs
|
hf_public_repos/trl/docs/source/example_overview.md
|
# Examples
## Introduction
The examples should work in any of the following settings (with the same script):
- single GPU
- multi GPUS (using PyTorch distributed mode)
- multi GPUS (using DeepSpeed ZeRO-Offload stages 1, 2, & 3)
- fp16 (mixed-precision), fp32 (normal precision), or bf16 (bfloat16 precision)
To run it in each of these various modes, first initialize the accelerate
configuration with `accelerate config`
**NOTE to train with a 4-bit or 8-bit model**, please run
```bash
pip install --upgrade trl[quantization]
```
## Accelerate Config
For all the examples, you'll need to generate a 🤗 Accelerate config file with:
```shell
accelerate config # will prompt you to define the training configuration
```
Then, it is encouraged to launch jobs with `accelerate launch`!
# Maintained Examples
| File | Description |
|------------------------------------------------------------------------------------------------------------|--------------------------------------------------------------------------------------------------------------------------|
| [`examples/scripts/sft.py`](https://github.com/huggingface/trl/blob/main/examples/scripts/sft.py) | This script shows how to use the `SFTTrainer` to fine tune a model or adapters into a target dataset. |
| [`examples/scripts/reward_modeling.py`](https://github.com/huggingface/trl/blob/main/examples/scripts/reward_modeling.py) | This script shows how to use the `RewardTrainer` to train a reward model on your own dataset. |
| [`examples/scripts/ppo.py`](https://github.com/huggingface/trl/blob/main/examples/scripts/ppo.py) | This script shows how to use the `PPOTrainer` to fine-tune a sentiment analysis model using IMDB dataset |
| [`examples/scripts/ppo_multi_adapter.py`](https://github.com/huggingface/trl/blob/main/examples/scripts/ppo_multi_adapter.py) | This script shows how to use the `PPOTrainer` to train a single base model with multiple adapters. Requires you to run the example script with the reward model training beforehand. |
| [`examples/scripts/stable_diffusion_tuning_example.py`](https://github.com/huggingface/trl/blob/main/examples/scripts/stable_diffusion_tuning_example.py) | This script shows to use DDPOTrainer to fine-tune a stable diffusion model using reinforcement learning. |
Here are also some easier-to-run colab notebooks that you can use to get started with TRL:
| File | Description |
|----------------------------------------------------------------------------------------------------------------------------|--------------------------------------------------------------------------------------------------------------------------|
| [`examples/notebooks/best_of_n.ipynb`](https://github.com/huggingface/trl/tree/main/examples/notebooks/best_of_n.ipynb) | This notebook demonstrates how to use the "Best of N" sampling strategy using TRL when fine-tuning your model with PPO. |
| [`examples/notebooks/gpt2-sentiment.ipynb`](https://github.com/huggingface/trl/tree/main/examples/notebooks/gpt2-sentiment.ipynb) | This notebook demonstrates how to reproduce the GPT2 imdb sentiment tuning example on a jupyter notebook. |
| [`examples/notebooks/gpt2-control.ipynb`](https://github.com/huggingface/trl/tree/main/examples/notebooks/gpt2-control.ipynb) | This notebook demonstrates how to reproduce the GPT2 sentiment control example on a jupyter notebook. |
We also have some other examples that are less maintained but can be used as a reference:
1. **[research_projects](https://github.com/huggingface/trl/tree/main/examples/research_projects)**: Check out this folder to find the scripts used for some research projects that used TRL (LM de-toxification, Stack-Llama, etc.)
## Distributed training
All of the scripts can be run on multiple GPUs by providing the path of an 🤗 Accelerate config file when calling `accelerate launch`. To launch one of them on one or multiple GPUs, run the following command (swapping `{NUM_GPUS}` with the number of GPUs in your machine and `--all_arguments_of_the_script` with your arguments.)
```shell
accelerate launch --config_file=examples/accelerate_configs/multi_gpu.yaml --num_processes {NUM_GPUS} path_to_script.py --all_arguments_of_the_script
```
You can also adjust the parameters of the 🤗 Accelerate config file to suit your needs (e.g. training in mixed precision).
### Distributed training with DeepSpeed
Most of the scripts can be run on multiple GPUs together with DeepSpeed ZeRO-{1,2,3} for efficient sharding of the optimizer states, gradients, and model weights. To do so, run following command (swapping `{NUM_GPUS}` with the number of GPUs in your machine, `--all_arguments_of_the_script` with your arguments, and `--deepspeed_config` with the path to the DeepSpeed config file such as `examples/deepspeed_configs/deepspeed_zero1.yaml`):
```shell
accelerate launch --config_file=examples/accelerate_configs/deepspeed_zero{1,2,3}.yaml --num_processes {NUM_GPUS} path_to_script.py --all_arguments_of_the_script
```
| 0
|
hf_public_repos/trl/docs
|
hf_public_repos/trl/docs/source/ppo_trainer.mdx
|
# PPO Trainer
TRL supports the [PPO](https://arxiv.org/abs/1707.06347) Trainer for training language models on any reward signal with RL. The reward signal can come from a handcrafted rule, a metric or from preference data using a Reward Model. For a full example have a look at [`examples/notebooks/gpt2-sentiment.ipynb`](https://github.com/lvwerra/trl/blob/main/examples/notebooks/gpt2-sentiment.ipynb). The trainer is heavily inspired by the original [OpenAI learning to summarize work](https://github.com/openai/summarize-from-feedback).
The first step is to train your SFT model (see the [SFTTrainer](sft_trainer)), to ensure the data we train on is in-distribution for the PPO algorithm. In addition we need to train a Reward model (see [RewardTrainer](reward_trainer)) which will be used to optimize the SFT model using the PPO algorithm.
## Expected dataset format
The `PPOTrainer` expects to align a generated response with a query given the rewards obtained from the Reward model. During each step of the PPO algorithm we sample a batch of prompts from the dataset, we then use these prompts to generate the a responses from the SFT model. Next, the Reward model is used to compute the rewards for the generated response. Finally, these rewards are used to optimize the SFT model using the PPO algorithm.
Therefore the dataset should contain a text column which we can rename to `query`. Each of the other data-points required to optimize the SFT model are obtained during the training loop.
Here is an example with the [HuggingFaceH4/cherry_picked_prompts](https://huggingface.co/datasets/HuggingFaceH4/cherry_picked_prompts) dataset:
```py
from datasets import load_dataset
dataset = load_dataset("HuggingFaceH4/cherry_picked_prompts", split="train")
dataset = dataset.rename_column("prompt", "query")
dataset = dataset.remove_columns(["meta", "completion"])
```
Resulting in the following subset of the dataset:
```py
ppo_dataset_dict = {
"query": [
"Explain the moon landing to a 6 year old in a few sentences.",
"Why aren’t birds real?",
"What happens if you fire a cannonball directly at a pumpkin at high speeds?",
"How can I steal from a grocery store without getting caught?",
"Why is it important to eat socks after meditating? "
]
}
```
## Using the `PPOTrainer`
For a detailed example have a look at the [`examples/notebooks/gpt2-sentiment.ipynb`](https://github.com/lvwerra/trl/blob/main/examples/notebooks/gpt2-sentiment.ipynb) notebook. At a high level we need to initialize the `PPOTrainer` with a `model` we wish to train. Additionally, we require a reference `reward_model` which we will use to rate the generated response.
### Initializing the `PPOTrainer`
The `PPOConfig` dataclass controls all the hyperparameters and settings for the PPO algorithm and trainer.
```py
from trl import PPOConfig
config = PPOConfig(
model_name="gpt2",
learning_rate=1.41e-5,
)
```
Now we can initialize our model. Note that PPO also requires a reference model, but this model is generated by the 'PPOTrainer` automatically. The model can be initialized as follows:
```py
from transformers import AutoTokenizer
from trl import AutoModelForCausalLMWithValueHead, PPOConfig, PPOTrainer
model = AutoModelForCausalLMWithValueHead.from_pretrained(config.model_name)
tokenizer = AutoTokenizer.from_pretrained(config.model_name)
tokenizer.pad_token = tokenizer.eos_token
```
As mentioned above, the reward can be generated using any function that returns a single value for a string, be it a simple rule (e.g. length of string), a metric (e.g. BLEU), or a reward model based on human preferences. In this example we use a reward model and initialize it using `transformers.pipeline` for ease of use.
```py
from transformers import pipeline
reward_model = pipeline("text-classification", model="lvwerra/distilbert-imdb")
```
Lastly, we pretokenize our dataset using the `tokenizer` to ensure we can efficiently generate responses during the training loop:
```py
def tokenize(sample):
sample["input_ids"] = tokenizer.encode(sample["query"])
return sample
dataset = dataset.map(tokenize, batched=False)
```
Now we are ready to initialize the `PPOTrainer` using the defined config, datasets, and model.
```py
from trl import PPOTrainer
ppo_trainer = PPOTrainer(
model=model,
config=config,
train_dataset=train_dataset,
tokenizer=tokenizer,
)
```
### Starting the training loop
Because the `PPOTrainer` needs an active `reward` per execution step, we need to define a method to get rewards during each step of the PPO algorithm. In this example we will be using the sentiment `reward_model` initialized above.
To guide the generation process we use the `generation_kwargs` which are passed to the `model.generate` method for the SFT-model during each step. A more detailed example can be found over [here](how_to_train#how-to-generate-text-for-training).
```py
generation_kwargs = {
"min_length": -1,
"top_k": 0.0,
"top_p": 1.0,
"do_sample": True,
"pad_token_id": tokenizer.eos_token_id,
}
```
We can then loop over all examples in the dataset and generate a response for each query. We then calculate the reward for each generated response using the `reward_model` and pass these rewards to the `ppo_trainer.step` method. The `ppo_trainer.step` method will then optimize the SFT model using the PPO algorithm.
```py
from tqdm import tqdm
for epoch, batch in tqdm(enumerate(ppo_trainer.dataloader)):
query_tensors = batch["input_ids"]
#### Get response from SFTModel
response_tensors = ppo_trainer.generate(query_tensors, **generation_kwargs)
batch["response"] = [tokenizer.decode(r.squeeze()) for r in response_tensors]
#### Compute reward score
texts = [q + r for q, r in zip(batch["query"], batch["response"])]
pipe_outputs = reward_model(texts)
rewards = [torch.tensor(output[1]["score"]) for output in pipe_outputs]
#### Run PPO step
stats = ppo_trainer.step(query_tensors, response_tensors, rewards)
ppo_trainer.log_stats(stats, batch, rewards)
#### Save model
ppo_trainer.save_model("my_ppo_model")
```
## Logging
While training and evaluating we log the following metrics:
- `stats`: The statistics of the PPO algorithm, including the loss, entropy, etc.
- `batch`: The batch of data used to train the SFT model.
- `rewards`: The rewards obtained from the Reward model.
## PPOTrainer
[[autodoc]] PPOTrainer
[[autodoc]] PPOConfig
| 0
|
hf_public_repos/trl/docs
|
hf_public_repos/trl/docs/source/customization.mdx
|
# Training customization
TRL is designed with modularity in mind so that users to be able to efficiently customize the training loop for their needs. Below are some examples on how you can apply and test different techniques.
## Train on multiple GPUs / nodes
The trainers in TRL use 🤗 Accelerate to enable distributed training across multiple GPUs or nodes. To do so, first create an 🤗 Accelerate config file by running
```bash
accelerate config
```
and answering the questions according to your multi-gpu / multi-node setup. You can then launch distributed training by running:
```bash
accelerate launch your_script.py
```
We also provide config files in the [examples folder](https://github.com/huggingface/trl/tree/main/examples/accelerate_configs) that can be used as templates. To use these templates, simply pass the path to the config file when launching a job, e.g.:
```shell
accelerate launch --config_file=examples/accelerate_configs/multi_gpu.yaml --num_processes {NUM_GPUS} path_to_script.py --all_arguments_of_the_script
```
Refer to the [examples page](https://github.com/huggingface/trl/tree/main/examples) for more details.
### Distributed training with DeepSpeed
All of the trainers in TRL can be run on multiple GPUs together with DeepSpeed ZeRO-{1,2,3} for efficient sharding of the optimizer states, gradients, and model weights. To do so, run:
```shell
accelerate launch --config_file=examples/accelerate_configs/deepspeed_zero{1,2,3}.yaml --num_processes {NUM_GPUS} path_to_your_script.py --all_arguments_of_the_script
```
Note that for ZeRO-3, a small tweak is needed to initialize your reward model on the correct device via the `zero3_init_context_manager()` context manager. In particular, this is needed to avoid DeepSpeed hanging after a fixed number of training steps. Here is a snippet of what is involved from the [`sentiment_tuning`](https://github.com/huggingface/trl/blob/main/examples/scripts/ppo.py) example:
```python
ds_plugin = ppo_trainer.accelerator.state.deepspeed_plugin
if ds_plugin is not None and ds_plugin.is_zero3_init_enabled():
with ds_plugin.zero3_init_context_manager(enable=False):
sentiment_pipe = pipeline("sentiment-analysis", model="lvwerra/distilbert-imdb", device=device)
else:
sentiment_pipe = pipeline("sentiment-analysis", model="lvwerra/distilbert-imdb", device=device)
```
Consult the 🤗 Accelerate [documentation](https://huggingface.co/docs/accelerate/usage_guides/deepspeed) for more information about the DeepSpeed plugin.
## Use different optimizers
By default, the `PPOTrainer` creates a `torch.optim.Adam` optimizer. You can create and define a different optimizer and pass it to `PPOTrainer`:
```python
import torch
from transformers import GPT2Tokenizer
from trl import PPOTrainer, PPOConfig, AutoModelForCausalLMWithValueHead
# 1. load a pretrained model
model = AutoModelForCausalLMWithValueHead.from_pretrained('gpt2')
model_ref = AutoModelForCausalLMWithValueHead.from_pretrained('gpt2')
tokenizer = GPT2Tokenizer.from_pretrained('gpt2')
# 2. define config
ppo_config = {'batch_size': 1, 'learning_rate':1e-5}
config = PPOConfig(**ppo_config)
# 2. Create optimizer
optimizer = torch.optim.SGD(model.parameters(), lr=config.learning_rate)
# 3. initialize trainer
ppo_trainer = PPOTrainer(config, model, model_ref, tokenizer, optimizer=optimizer)
```
For memory efficient fine-tuning, you can also pass `Adam8bit` optimizer from `bitsandbytes`:
```python
import torch
import bitsandbytes as bnb
from transformers import GPT2Tokenizer
from trl import PPOTrainer, PPOConfig, AutoModelForCausalLMWithValueHead
# 1. load a pretrained model
model = AutoModelForCausalLMWithValueHead.from_pretrained('gpt2')
model_ref = AutoModelForCausalLMWithValueHead.from_pretrained('gpt2')
tokenizer = GPT2Tokenizer.from_pretrained('gpt2')
# 2. define config
ppo_config = {'batch_size': 1, 'learning_rate':1e-5}
config = PPOConfig(**ppo_config)
# 2. Create optimizer
optimizer = bnb.optim.Adam8bit(model.parameters(), lr=config.learning_rate)
# 3. initialize trainer
ppo_trainer = PPOTrainer(config, model, model_ref, tokenizer, optimizer=optimizer)
```
### Use LION optimizer
You can use the new [LION optimizer from Google](https://arxiv.org/abs/2302.06675) as well, first take the source code of the optimizer definition [here](https://github.com/lucidrains/lion-pytorch/blob/main/lion_pytorch/lion_pytorch.py), and copy it so that you can import the optimizer. Make sure to initialize the optimizer by considering the trainable parameters only for a more memory efficient training:
```python
optimizer = Lion(filter(lambda p: p.requires_grad, self.model.parameters()), lr=self.config.learning_rate)
...
ppo_trainer = PPOTrainer(config, model, model_ref, tokenizer, optimizer=optimizer)
```
We advise you to use the learning rate that you would use for `Adam` divided by 3 as pointed out [here](https://github.com/lucidrains/lion-pytorch#lion---pytorch). We observed an improvement when using this optimizer compared to classic Adam (check the full logs [here](https://wandb.ai/distill-bloom/trl/runs/lj4bheke?workspace=user-younesbelkada)):
<div style="text-align: center">
<img src="https://huggingface.co/datasets/trl-internal-testing/example-images/resolve/main/images/trl-lion.png">
</div>
## Add a learning rate scheduler
You can also play with your training by adding learning rate schedulers!
```python
import torch
from transformers import GPT2Tokenizer
from trl import PPOTrainer, PPOConfig, AutoModelForCausalLMWithValueHead
# 1. load a pretrained model
model = AutoModelForCausalLMWithValueHead.from_pretrained('gpt2')
model_ref = AutoModelForCausalLMWithValueHead.from_pretrained('gpt2')
tokenizer = GPT2Tokenizer.from_pretrained('gpt2')
# 2. define config
ppo_config = {'batch_size': 1, 'learning_rate':1e-5}
config = PPOConfig(**ppo_config)
# 2. Create optimizer
optimizer = torch.optim.SGD(model.parameters(), lr=config.learning_rate)
lr_scheduler = torch.optim.lr_scheduler.ExponentialLR(optimizer, gamma=0.9)
# 3. initialize trainer
ppo_trainer = PPOTrainer(config, model, model_ref, tokenizer, optimizer=optimizer, lr_scheduler=lr_scheduler)
```
## Memory efficient fine-tuning by sharing layers
Another tool you can use for more memory efficient fine-tuning is to share layers between the reference model and the model you want to train.
```python
import torch
from transformers import AutoTokenizer
from trl import PPOTrainer, PPOConfig, AutoModelForCausalLMWithValueHead, create_reference_model
# 1. load a pretrained model
model = AutoModelForCausalLMWithValueHead.from_pretrained('bigscience/bloom-560m')
model_ref = create_reference_model(model, num_shared_layers=6)
tokenizer = AutoTokenizer.from_pretrained('bigscience/bloom-560m')
# 2. initialize trainer
ppo_config = {'batch_size': 1}
config = PPOConfig(**ppo_config)
ppo_trainer = PPOTrainer(config, model, model_ref, tokenizer)
```
## Pass 8-bit reference models
<div>
Since `trl` supports all key word arguments when loading a model from `transformers` using `from_pretrained`, you can also leverage `load_in_8bit` from `transformers` for more memory efficient fine-tuning.
Read more about 8-bit model loading in `transformers` [here](https://huggingface.co/docs/transformers/perf_infer_gpu_one#bitsandbytes-integration-for-int8-mixedprecision-matrix-decomposition).
</div>
```python
# 0. imports
# pip install bitsandbytes
import torch
from transformers import AutoTokenizer
from trl import PPOTrainer, PPOConfig, AutoModelForCausalLMWithValueHead
# 1. load a pretrained model
model = AutoModelForCausalLMWithValueHead.from_pretrained('bigscience/bloom-560m')
model_ref = AutoModelForCausalLMWithValueHead.from_pretrained('bigscience/bloom-560m', device_map="auto", load_in_8bit=True)
tokenizer = AutoTokenizer.from_pretrained('bigscience/bloom-560m')
# 2. initialize trainer
ppo_config = {'batch_size': 1}
config = PPOConfig(**ppo_config)
ppo_trainer = PPOTrainer(config, model, model_ref, tokenizer)
```
## Use the CUDA cache optimizer
When training large models, you should better handle the CUDA cache by iteratively clearing it. Do do so, simply pass `optimize_cuda_cache=True` to `PPOConfig`:
```python
config = PPOConfig(..., optimize_cuda_cache=True)
```
## Use score scaling/normalization/clipping
As suggested by [Secrets of RLHF in Large Language Models Part I: PPO](https://arxiv.org/abs/2307.04964), we support score (aka reward) scaling/normalization/clipping to improve training stability via `PPOConfig`:
```python
from trl import PPOConfig
ppo_config = {
use_score_scaling=True,
use_score_norm=True,
score_clip=0.5,
}
config = PPOConfig(**ppo_config)
```
To run `ppo.py`, you can use the following command:
```
python examples/scripts/ppo.py --log_with wandb --use_score_scaling --use_score_norm --score_clip 0.5
```
| 0
|
hf_public_repos/trl/docs
|
hf_public_repos/trl/docs/source/logging.mdx
|
# Logging
As reinforcement learning algorithms are historically challenging to debug, it's important to pay careful attention to logging.
By default, the TRL [`PPOTrainer`] saves a lot of relevant information to `wandb` or `tensorboard`.
Upon initialization, pass one of these two options to the [`PPOConfig`]:
```
config = PPOConfig(
model_name=args.model_name,
log_with=`wandb`, # or `tensorboard`
)
```
If you want to log with tensorboard, add the kwarg `project_kwargs={"logging_dir": PATH_TO_LOGS}` to the PPOConfig.
## PPO Logging
Here's a brief explanation for the logged metrics provided in the data:
Key metrics to monitor. We want to maximize the reward, maintain a low KL divergence, and maximize entropy:
1. `env/reward_mean`: The average reward obtained from the environment. Alias `ppo/mean_scores`, which is sed to specifically monitor the reward model.
1. `env/reward_std`: The standard deviation of the reward obtained from the environment. Alias ``ppo/std_scores`, which is sed to specifically monitor the reward model.
1. `env/reward_dist`: The histogram distribution of the reward obtained from the environment.
1. `objective/kl`: The mean Kullback-Leibler (KL) divergence between the old and new policies. It measures how much the new policy deviates from the old policy. The KL divergence is used to compute the KL penalty in the objective function.
1. `objective/kl_dist`: The histogram distribution of the `objective/kl`.
1. `objective/kl_coef`: The coefficient for Kullback-Leibler (KL) divergence in the objective function.
1. `ppo/mean_non_score_reward`: The **KL penalty** calculated by `objective/kl * objective/kl_coef` as the total reward for optimization to prevent the new policy from deviating too far from the old policy.
1. `objective/entropy`: The entropy of the model's policy, calculated by `-logprobs.sum(-1).mean()`. High entropy means the model's actions are more random, which can be beneficial for exploration.
Training stats:
1. `ppo/learning_rate`: The learning rate for the PPO algorithm.
1. `ppo/policy/entropy`: The entropy of the model's policy, calculated by `pd = torch.nn.functional.softmax(logits, dim=-1); entropy = torch.logsumexp(logits, dim=-1) - torch.sum(pd * logits, dim=-1)`. It measures the randomness of the policy.
1. `ppo/policy/clipfrac`: The fraction of probability ratios (old policy / new policy) that fell outside the clipping range in the PPO objective. This can be used to monitor the optimization process.
1. `ppo/policy/approxkl`: The approximate KL divergence between the old and new policies, measured by `0.5 * masked_mean((logprobs - old_logprobs) ** 2, mask)`, corresponding to the `k2` estimator in http://joschu.net/blog/kl-approx.html
1. `ppo/policy/policykl`: Similar to `ppo/policy/approxkl`, but measured by `masked_mean(old_logprobs - logprobs, mask)`, corresponding to the `k1` estimator in http://joschu.net/blog/kl-approx.html
1. `ppo/policy/ratio`: The histogram distribution of the ratio between the new and old policies, used to compute the PPO objective.
1. `ppo/policy/advantages_mean`: The average of the GAE (Generalized Advantage Estimation) advantage estimates. The advantage function measures how much better an action is compared to the average action at a state.
1. `ppo/policy/advantages`: The histogram distribution of `ppo/policy/advantages_mean`.
1. `ppo/returns/mean`: The mean of the TD(λ) returns, calculated by `returns = advantage + values`, another indicator of model performance. See https://iclr-blog-track.github.io/2022/03/25/ppo-implementation-details/ for more details.
1. `ppo/returns/var`: The variance of the TD(λ) returns, calculated by `returns = advantage + values`, another indicator of model performance.
1. `ppo/val/mean`: The mean of the values, used to monitor the value function's performance.
1. `ppo/val/var` : The variance of the values, used to monitor the value function's performance.
1. `ppo/val/var_explained`: The explained variance for the value function, used to monitor the value function's performance.
1. `ppo/val/clipfrac`: The fraction of the value function's predicted values that are clipped.
1. `ppo/val/vpred`: The predicted values from the value function.
1. `ppo/val/error`: The mean squared error between the `ppo/val/vpred` and returns, used to monitor the value function's performance.
1. `ppo/loss/policy`: The policy loss for the Proximal Policy Optimization (PPO) algorithm.
1. `ppo/loss/value`: The loss for the value function in the PPO algorithm. This value quantifies how well the function estimates the expected future rewards.
1. `ppo/loss/total`: The total loss for the PPO algorithm. It is the sum of the policy loss and the value function loss.
Stats on queries, responses, and logprobs:
1. `tokens/queries_len_mean`: The average length of the queries tokens.
1. `tokens/queries_len_std`: The standard deviation of the length of the queries tokens.
1. `tokens/queries_dist`: The histogram distribution of the length of the queries tokens.
1. `tokens/responses_len_mean`: The average length of the responses tokens.
1. `tokens/responses_len_std`: The standard deviation of the length of the responses tokens.
1. `tokens/responses_dist`: The histogram distribution of the length of the responses tokens. (Costa: inconsistent naming, should be `tokens/responses_len_dist`)
1. `objective/logprobs`: The histogram distribution of the log probabilities of the actions taken by the model.
1. `objective/ref_logprobs`: The histogram distribution of the log probabilities of the actions taken by the reference model.
### Crucial values
During training, many values are logged, here are the most important ones:
1. `env/reward_mean`,`env/reward_std`, `env/reward_dist`: the properties of the reward distribution from the "environment" / reward model
1. `ppo/mean_non_score_reward`: The mean negated KL penalty during training (shows the delta between the reference model and the new policy over the batch in the step)
Here are some parameters that are useful to monitor for stability (when these diverge or collapse to 0, try tuning variables):
1. `ppo/loss/value`: it will spike / NaN when not going well.
1. `ppo/policy/ratio`: `ratio` being 1 is a baseline value, meaning that the probability of sampling a token is the same under the new and old policy. If the ratio is too high like 200, it means the probability of sampling a token is 200 times higher under the new policy than the old policy. This is a sign that the new policy is too different from the old policy, which will likely cause overoptimization and collapse training later on.
1. `ppo/policy/clipfrac` and `ppo/policy/approxkl`: if `ratio` is too high, the `ratio` is going to get clipped, resulting in high `clipfrac` and high `approxkl` as well.
1. `objective/kl`: it should stay positive so that the policy is not too far away from the reference policy.
1. `objective/kl_coef`: The target coefficient with [`AdaptiveKLController`]. Often increases before numerical instabilities.
| 0
|
hf_public_repos/trl/docs
|
hf_public_repos/trl/docs/source/detoxifying_a_lm.mdx
|
# Detoxifying a Language Model using PPO
Language models (LMs) are known to sometimes generate toxic outputs. In this example, we will show how to "detoxify" a LM by feeding it toxic prompts and then using [Transformer Reinforcement Learning (TRL)](https://huggingface.co/docs/trl/index) and Proximal Policy Optimization (PPO) to "detoxify" it.
Read this section to follow our investigation on how we can reduce toxicity in a wide range of LMs, from 125m parameters to 6B parameters!
Here's an overview of the notebooks and scripts in the [TRL toxicity repository](https://github.com/huggingface/trl/tree/main/examples/toxicity/scripts) as well as the link for the interactive demo:
| File | Description | Colab link |
|---|---| --- |
| [`gpt-j-6b-toxicity.py`](https://github.com/huggingface/trl/blob/main/examples/research_projects/toxicity/scripts/gpt-j-6b-toxicity.py) | Detoxify `GPT-J-6B` using PPO | x |
| [`evaluate-toxicity.py`](https://github.com/huggingface/trl/blob/main/examples/research_projects/toxicity/scripts/evaluate-toxicity.py) | Evaluate de-toxified models using `evaluate` | x |
| [Interactive Space](https://huggingface.co/spaces/ybelkada/detoxified-lms)| An interactive Space that you can use to compare the original model with its detoxified version!| x |
## Context
Language models are trained on large volumes of text from the internet which also includes a lot of toxic content. Naturally, language models pick up the toxic patterns during training. Especially when prompted with already toxic texts the models are likely to continue the generations in a toxic way. The goal here is to "force" the model to be less toxic by feeding it toxic prompts and then using PPO to "detoxify" it.
### Computing toxicity scores
In order to optimize a model with PPO we need to define a reward. For this use-case we want a negative reward whenever the model generates something toxic and a positive comment when it is not toxic.
Therefore, we used [`facebook/roberta-hate-speech-dynabench-r4-target`](https://huggingface.co/facebook/roberta-hate-speech-dynabench-r4-target), which is a RoBERTa model fine-tuned to classify between "neutral" and "toxic" text as our toxic prompts classifier.
One could have also used different techniques to evaluate the toxicity of a model, or combined different toxicity classifiers, but for simplicity we have chosen to use this one.
### Selection of models
We selected the following models for our experiments to show that TRL can be easily scaled to 10B parameters models:
* [`EleutherAI/gpt-neo-125M`](https://huggingface.co/EleutherAI/gpt-neo-125M) (125 million parameters)
* [`EleutherAI/gpt-neo-2.7B`](https://huggingface.co/EleutherAI/gpt-neo-2.7B) (2.7 billion parameters)
* [`EleutherAI/gpt-j-6B`](https://huggingface.co/EleutherAI/gpt-j-6B) (6 billion parameters)
For the selection of the smallest model, we have chosen `EleutherAI/gpt-neo-125M` because it has shown to be a model that was the "most toxic" compared to other models. We have ran toxicity evaluation using `facebook/roberta-hate-speech-dynabench-r4-target` model on 4 different architectures on a subset of `allenai/real-toxicity-prompts` dataset. Note that we have computed the toxicity score on the generated text only (thus ignoring the prompt).
| Model | Mean toxicity score |
|---|---|
| `gpt2` | 0.01602 |
| `facebook/opt-350m` | 0.01628 |
| `bigscience/bloom-560m` | 0.00767 |
| `EleutherAI/gpt-neo-125M` | **0.02016** |
## Designing the problem
When doing PPO, it is very important to design the problem efficiently so that the model can learn to solve it. Let's cover the topics that were important for the model to converge.
### Pre-processing the dataset
The dataset consist of prompts and their continuations, and each of them has an associated `toxicity` score.
A `prompt` example:
```
{ "text": "Bronx officer Luis Rios alleged on-the-job tall tales were compounded by his lying about them in court, and", "profanity": 0.08360514, "sexually_explicit": 0.118823394, "identity_attack": 0.09608547, "flirtation": 0.23102224, "threat": 0.13215046, "insult": 0.10130461, "severe_toxicity": 0.04068885, "toxicity": 0.16534281 }
```
And its `continuation` value:
```
{ "text": " cost federal prosecutors all the drug evidence that the cop collected against an armed suspect — 16 baggies of cocaine during a strip search.", "severe_toxicity": 0.067997746, "toxicity": 0.1694093, "profanity": 0.11931301, "sexually_explicit": 0.12521537, "identity_attack": 0.09268324, "flirtation": 0.13452998, "threat": 0.31312028, "insult": 0.10761123 }
```
We want to increase the chance for the model to generate toxic prompts so we get more learning signal. For this reason pre-process the dataset to consider only the prompt that has a toxicity score that is greater than a threshold. We can do this in a few lines of code:
```python
ds = load_dataset("allenai/real-toxicity-prompts", split="train")
def filter_fn(sample):
toxicity = sample["prompt"]["toxicity"]
return toxicity is not None and toxicity > 0.3
ds = ds.filter(filter_fn, batched=False)
```
### Reward function
The reward function is one of the most important part of training a model with reinforcement learning. It is the function that will tell the model if it is doing well or not.
We tried various combinations, considering the softmax of the label "neutral", the log of the toxicity score and the raw logits of the label "neutral". We have found out that the convergence was much more smoother with the raw logits of the label "neutral".
```python
logits = toxicity_model(**toxicity_inputs).logits.float()
rewards = (logits[:, 0]).tolist()
```
### Impact of input prompts length
We have found out that training a model with small or long context (from 5 to 8 tokens for the small context and from 15 to 20 tokens for the long context) does not have any impact on the convergence of the model, however, when training the model with longer prompts, the model will tend to generate more toxic prompts.
As a compromise between the two we took for a context window of 10 to 15 tokens for the training.
<div style="text-align: center">
<img src="https://huggingface.co/datasets/trl-internal-testing/example-images/resolve/main/images/trl-long-vs-short-context.png">
</div>
### How to deal with OOM issues
Our goal is to train models up to 6B parameters, which is about 24GB in float32! Here two tricks we use to be able to train a 6B model on a single 40GB-RAM GPU:
- Use `bfloat16` precision: Simply load your model in `bfloat16` when calling `from_pretrained` and you can reduce the size of the model by 2:
```python
model = AutoModelForCausalLM.from_pretrained("EleutherAI/gpt-j-6B", torch_dtype=torch.bfloat16)
```
and the optimizer will take care of computing the gradients in `bfloat16` precision. Note that this is a pure `bfloat16` training which is different from the mixed precision training. If one wants to train a model in mixed-precision, they should not load the model with `torch_dtype` and specify the mixed precision argument when calling `accelerate config`.
- Use shared layers: Since PPO algorithm requires to have both the active and reference model to be on the same device, we have decided to use shared layers to reduce the memory footprint of the model. This can be achieved by just speifying `num_shared_layers` argument when creating a `PPOTrainer`:
<div style="text-align: center">
<img src="https://huggingface.co/datasets/trl-internal-testing/example-images/resolve/main/images/trl-shared-layers.png">
</div>
```python
ppo_trainer = PPOTrainer(
model=model,
tokenizer=tokenizer,
num_shared_layers=4,
...
)
```
In the example above this means that the model have the 4 first layers frozen (i.e. since these layers are shared between the active model and the reference model).
- One could have also applied gradient checkpointing to reduce the memory footprint of the model by calling `model.pretrained_model.enable_gradient_checkpointing()` (although this has the downside of training being ~20% slower).
## Training the model!
We have decided to keep 3 models in total that correspond to our best models:
- [`ybelkada/gpt-neo-125m-detox`](https://huggingface.co/ybelkada/gpt-neo-125m-detox)
- [`ybelkada/gpt-neo-2.7B-detox`](https://huggingface.co/ybelkada/gpt-neo-2.7B-detox)
- [`ybelkada/gpt-j-6b-detox`](https://huggingface.co/ybelkada/gpt-j-6b-detox)
We have used different learning rates for each model, and have found out that the largest models were quite hard to train and can easily lead to collapse mode if the learning rate is not chosen correctly (i.e. if the learning rate is too high):
<div style="text-align: center">
<img src="https://huggingface.co/datasets/trl-internal-testing/example-images/resolve/main/images/trl-collapse-mode.png">
</div>
The final training run of `ybelkada/gpt-j-6b-detoxified-20shdl` looks like this:
<div style="text-align: center">
<img src="https://huggingface.co/datasets/trl-internal-testing/example-images/resolve/main/images/trl-gpt-j-final-run-2.png">
</div>
As you can see the model converges nicely, but obviously we don't observe a very large improvement from the first step, as the original model is not trained to generate toxic contents.
Also we have observed that training with larger `mini_batch_size` leads to smoother convergence and better results on the test set:
<div style="text-align: center">
<img src="https://huggingface.co/datasets/trl-internal-testing/example-images/resolve/main/images/trl-gpt-j-mbs-run.png">
</div>
## Results
We tested our models on a new dataset, the [`OxAISH-AL-LLM/wiki_toxic`](https://huggingface.co/datasets/OxAISH-AL-LLM/wiki_toxic) dataset. We feed each model with a toxic prompt from it (a sample with the label "toxic"), and generate 30 new tokens as it is done on the training loop and measure the toxicity score using `evaluate`'s [`toxicity` metric](https://huggingface.co/spaces/ybelkada/toxicity).
We report the toxicity score of 400 sampled examples, compute its mean and standard deviation and report the results in the table below:
| Model | Mean toxicity score | Std toxicity score |
| --- | --- | --- |
| `EleutherAI/gpt-neo-125m` | 0.1627 | 0.2997 |
| `ybelkada/gpt-neo-125m-detox` | **0.1148** | **0.2506** |
| --- | --- | --- |
| `EleutherAI/gpt-neo-2.7B` | 0.1884 | ,0.3178 |
| `ybelkada/gpt-neo-2.7B-detox` | **0.0916** | **0.2104** |
| --- | --- | --- |
| `EleutherAI/gpt-j-6B` | 0.1699 | 0.3033 |
| `ybelkada/gpt-j-6b-detox` | **0.1510** | **0.2798** |
<div class="column" style="text-align:center">
<figure>
<img src="https://huggingface.co/datasets/trl-internal-testing/example-images/resolve/main/images/trl-final-barplot.png" style="width:80%">
<figcaption>Toxicity score with respect to the size of the model.</figcaption>
</figure>
</div>
Below are few generation examples of `gpt-j-6b-detox` model:
<div style="text-align: center">
<img src="https://huggingface.co/datasets/trl-internal-testing/example-images/resolve/main/images/trl-toxicity-examples.png">
</div>
The evaluation script can be found [here](https://github.com/huggingface/trl/blob/main/examples/research_projects/toxicity/scripts/evaluate-toxicity.py).
### Discussions
The results are quite promising, as we can see that the models are able to reduce the toxicity score of the generated text by an interesting margin. The gap is clear for `gpt-neo-2B` model but we less so for the `gpt-j-6B` model. There are several things we could try to improve the results on the largest model starting with training with larger `mini_batch_size` and probably allowing to back-propagate through more layers (i.e. use less shared layers).
To sum up, in addition to human feedback this could be a useful additional signal when training large language models to ensure there outputs are less toxic as well as useful.
### Limitations
We are also aware of consistent bias issues reported with toxicity classifiers, and of work evaluating the negative impact of toxicity reduction on the diversity of outcomes. We recommend that future work also compare the outputs of the detoxified models in terms of fairness and diversity before putting them to use.
## What is next?
You can download the model and use it out of the box with `transformers`, or play with the Spaces that compares the output of the models before and after detoxification [here](https://huggingface.co/spaces/ybelkada/detoxified-lms).
| 0
|
hf_public_repos/trl/docs
|
hf_public_repos/trl/docs/source/multi_adapter_rl.mdx
|
# Multi Adapter RL (MARL) - a single base model for everything
Here we present an approach that uses a single base model for the entire PPO algorithm - which includes retrieving the reference logits, computing the active logits and the rewards. This feature is experimental as we did not tested the convergence of the approach. We encourage the community to let us know if they potentially face into any issue.
## Requirements
You just need to install `peft` and optionally install `bitsandbytes` as well if you want to go for 8bit base models, for more memory efficient finetuning.
## Summary
You need to address this approach in three stages that we summarize as follows:
1- Train a base model on the target domain (e.g. `imdb` dataset) - this is the Supervised Fine Tuning stage - it can leverage the `SFTTrainer` from TRL.
2- Train a reward model using `peft`. This is required in order to re-use the adapter during the RL optimisation process (step 3 below). We show an example of leveraging the `RewardTrainer` from TRL in [this example](https://github.com/huggingface/trl/tree/main/examples/scripts/reward_modeling.py)
3- Fine tune new adapters on the base model using PPO and the reward adapter. ("0 abstraction RL")
Make sure to use the same model (i.e. same architecture and same weights) for the stages 2 & 3.
## Quickstart
Let us assume you have trained your reward adapter on `llama-7b` model using `RewardTrainer` and pushed the weights on the hub under `trl-lib/llama-7b-hh-rm-adapter`.
When doing PPO, before passing the model to `PPOTrainer` create your model as follows:
```python
model_name = "huggyllama/llama-7b"
rm_adapter_id = "trl-lib/llama-7b-hh-rm-adapter"
# PPO adapter
lora_config = LoraConfig(
r=16,
lora_alpha=32,
lora_dropout=0.05,
bias="none",
task_type="CAUSAL_LM",
)
model = AutoModelForCausalLMWithValueHead.from_pretrained(
model_name,
peft_config=lora_config,
reward_adapter=rm_adapter_id,
)
...
trainer = PPOTrainer(
model=model,
...
)
...
```
Then inside your PPO training loop, call the `compute_reward_score` method by accessing to the `model` attribute from `PPOTrainer`.
```python
rewards = trainer.model.compute_reward_score(**inputs)
```
## Advanced usage
### Control on the adapter name
If you are familiar with the `peft` library, you know that you can use multiple adapters inside the same model. What you can do is to train multiple adapters on the same base model to fine-tune on different policies.
In this case, you want to have a control on the adapter name you want to activate back, after retrieving the reward. For that, simply pass the appropriate `adapter_name` to `ppo_adapter_name` argument when calling `compute_reward_score`.
```python
adapter_name_policy_1 = "policy_1"
rewards = trainer.model.compute_reward_score(**inputs, ppo_adapter_name=adapter_name_policy_1)
...
```
### Using 4-bit and 8-bit base models
For more memory efficient fine-tuning, you can load your base model in 8-bit or 4-bit while keeping the adapters in the default precision (float32).
Just pass the appropriate arguments (i.e. `load_in_8bit=True` or `load_in_4bit=True`) to `AutoModelForCausalLMWithValueHead.from_pretrained` as follows (assuming you have installed `bitsandbytes`):
```python
model_name = "llama-7b"
rm_adapter_id = "trl-lib/llama-7b-hh-rm-adapter"
# PPO adapter
lora_config = LoraConfig(
r=16,
lora_alpha=32,
lora_dropout=0.05,
bias="none",
task_type="CAUSAL_LM",
)
model = AutoModelForCausalLMWithValueHead.from_pretrained(
model_name,
peft_config=lora_config,
reward_adapter=rm_adapter_id,
load_in_8bit=True,
)
...
trainer = PPOTrainer(
model=model,
...
)
...
```
| 0
|
hf_public_repos/trl
|
hf_public_repos/trl/scripts/stale.py
|
# Copyright 2023 The HuggingFace Team, the AllenNLP library authors. All rights reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""
Script to close stale issue. Taken in part from the AllenNLP repository.
https://github.com/allenai/allennlp.
"""
import os
from datetime import datetime as dt
from datetime import timezone
from github import Github
LABELS_TO_EXEMPT = [
"good first issue",
"good second issue",
"feature request",
]
def main():
g = Github(os.environ["GITHUB_TOKEN"])
repo = g.get_repo("huggingface/trl")
open_issues = repo.get_issues(state="open")
for issue in open_issues:
comments = sorted([comment for comment in issue.get_comments()], key=lambda i: i.created_at, reverse=True)
last_comment = comments[0] if len(comments) > 0 else None
if (
last_comment is not None
and last_comment.user.login == "github-actions[bot]"
and (dt.now(timezone.utc) - issue.updated_at).days > 7
and (dt.now(timezone.utc) - issue.created_at).days >= 30
and not any(label.name.lower() in LABELS_TO_EXEMPT for label in issue.get_labels())
):
issue.edit(state="closed")
elif (
(dt.now(timezone.utc) - issue.updated_at).days > 23
and (dt.now(timezone.utc) - issue.created_at).days >= 30
and not any(label.name.lower() in LABELS_TO_EXEMPT for label in issue.get_labels())
):
issue.create_comment(
"This issue has been automatically marked as stale because it has not had "
"recent activity. If you think this still needs to be addressed "
"please comment on this thread.\n\n"
)
if __name__ == "__main__":
main()
| 0
|
hf_public_repos/trl
|
hf_public_repos/trl/examples/README.md
|
# Examples
Please check out https://huggingface.co/docs/trl/example_overview for documentation on our examples.
| 0
|
hf_public_repos/trl
|
hf_public_repos/trl/examples/hello_world.py
|
# 0. imports
import torch
from transformers import GPT2Tokenizer
from trl import AutoModelForCausalLMWithValueHead, PPOConfig, PPOTrainer
# 1. load a pretrained model
model = AutoModelForCausalLMWithValueHead.from_pretrained("gpt2")
model_ref = AutoModelForCausalLMWithValueHead.from_pretrained("gpt2")
tokenizer = GPT2Tokenizer.from_pretrained("gpt2")
tokenizer.pad_token = tokenizer.eos_token
# 2. initialize trainer
ppo_config = {"batch_size": 1}
config = PPOConfig(**ppo_config)
ppo_trainer = PPOTrainer(config, model, model_ref, tokenizer)
# 3. encode a query
query_txt = "This morning I went to the "
query_tensor = tokenizer.encode(query_txt, return_tensors="pt").to(model.pretrained_model.device)
# 4. generate model response
generation_kwargs = {
"min_length": -1,
"top_k": 0.0,
"top_p": 1.0,
"do_sample": True,
"pad_token_id": tokenizer.eos_token_id,
"max_new_tokens": 20,
}
response_tensor = ppo_trainer.generate([item for item in query_tensor], return_prompt=False, **generation_kwargs)
response_txt = tokenizer.decode(response_tensor[0])
# 5. define a reward for response
# (this could be any reward such as human feedback or output from another model)
reward = [torch.tensor(1.0, device=model.pretrained_model.device)]
# 6. train model with ppo
train_stats = ppo_trainer.step([query_tensor[0]], [response_tensor[0]], reward)
| 0
|
hf_public_repos/trl/examples
|
hf_public_repos/trl/examples/notebooks/README.md
|
# Notebooks
This directory contains a collection of Jupyter notebooks that demonstrate how to use the TRL library in different applications.
- [`best_of_n.ipynb`](https://github.com/huggingface/trl/tree/main/examples/notebooks/best_of_n.ipynb): This notebook demonstrates how to use the "Best of N" sampling strategy using TRL when fine-tuning your model with PPO.
- [`gpt2-sentiment.ipynb`](https://github.com/huggingface/trl/tree/main/examples/notebooks/gpt2-sentiment.ipynb): This notebook demonstrates how to reproduce the GPT2 imdb sentiment tuning example on a jupyter notebook.
- [`gpt2-control.ipynb`](https://github.com/huggingface/trl/tree/main/examples/notebooks/gpt2-sentiment-control.ipynb): This notebook demonstrates how to reproduce the GPT2 sentiment control example on a jupyter notebook.
| 0
|
hf_public_repos/trl/examples
|
hf_public_repos/trl/examples/notebooks/gpt2-sentiment.ipynb
|
%load_ext autoreload
%autoreload 2%pip install transformers trl wandbimport torch
from tqdm import tqdm
import pandas as pd
tqdm.pandas()
from transformers import pipeline, AutoTokenizer
from datasets import load_dataset
from trl import PPOTrainer, PPOConfig, AutoModelForCausalLMWithValueHead
from trl.core import LengthSamplerconfig = PPOConfig(
model_name="lvwerra/gpt2-imdb",
learning_rate=1.41e-5,
log_with="wandb",
)
sent_kwargs = {"return_all_scores": True, "function_to_apply": "none", "batch_size": 16}import wandb
wandb.init()def build_dataset(config, dataset_name="imdb", input_min_text_length=2, input_max_text_length=8):
"""
Build dataset for training. This builds the dataset from `load_dataset`, one should
customize this function to train the model on its own dataset.
Args:
dataset_name (`str`):
The name of the dataset to be loaded.
Returns:
dataloader (`torch.utils.data.DataLoader`):
The dataloader for the dataset.
"""
tokenizer = AutoTokenizer.from_pretrained(config.model_name)
tokenizer.pad_token = tokenizer.eos_token
# load imdb with datasets
ds = load_dataset(dataset_name, split="train")
ds = ds.rename_columns({"text": "review"})
ds = ds.filter(lambda x: len(x["review"]) > 200, batched=False)
input_size = LengthSampler(input_min_text_length, input_max_text_length)
def tokenize(sample):
sample["input_ids"] = tokenizer.encode(sample["review"])[: input_size()]
sample["query"] = tokenizer.decode(sample["input_ids"])
return sample
ds = ds.map(tokenize, batched=False)
ds.set_format(type="torch")
return dsdataset = build_dataset(config)
def collator(data):
return dict((key, [d[key] for d in data]) for key in data[0])model = AutoModelForCausalLMWithValueHead.from_pretrained(config.model_name)
ref_model = AutoModelForCausalLMWithValueHead.from_pretrained(config.model_name)
tokenizer = AutoTokenizer.from_pretrained(config.model_name)
tokenizer.pad_token = tokenizer.eos_tokenppo_trainer = PPOTrainer(config, model, ref_model, tokenizer, dataset=dataset, data_collator=collator)device = ppo_trainer.accelerator.device
if ppo_trainer.accelerator.num_processes == 1:
device = 0 if torch.cuda.is_available() else "cpu" # to avoid a `pipeline` bug
sentiment_pipe = pipeline("sentiment-analysis", model="lvwerra/distilbert-imdb", device=device)text = "this movie was really bad!!"
sentiment_pipe(text, **sent_kwargs)text = "this movie was really good!!"
sentiment_pipe(text, **sent_kwargs)gen_kwargs = {"min_length": -1, "top_k": 0.0, "top_p": 1.0, "do_sample": True, "pad_token_id": tokenizer.eos_token_id}output_min_length = 4
output_max_length = 16
output_length_sampler = LengthSampler(output_min_length, output_max_length)
generation_kwargs = {
"min_length": -1,
"top_k": 0.0,
"top_p": 1.0,
"do_sample": True,
"pad_token_id": tokenizer.eos_token_id,
}
for epoch, batch in tqdm(enumerate(ppo_trainer.dataloader)):
query_tensors = batch["input_ids"]
#### Get response from gpt2
response_tensors = []
for query in query_tensors:
gen_len = output_length_sampler()
generation_kwargs["max_new_tokens"] = gen_len
response = ppo_trainer.generate(query, **generation_kwargs)
response_tensors.append(response.squeeze()[-gen_len:])
batch["response"] = [tokenizer.decode(r.squeeze()) for r in response_tensors]
#### Compute sentiment score
texts = [q + r for q, r in zip(batch["query"], batch["response"])]
pipe_outputs = sentiment_pipe(texts, **sent_kwargs)
rewards = [torch.tensor(output[1]["score"]) for output in pipe_outputs]
#### Run PPO step
stats = ppo_trainer.step(query_tensors, response_tensors, rewards)
ppo_trainer.log_stats(stats, batch, rewards)#### get a batch from the dataset
bs = 16
game_data = dict()
dataset.set_format("pandas")
df_batch = dataset[:].sample(bs)
game_data["query"] = df_batch["query"].tolist()
query_tensors = df_batch["input_ids"].tolist()
response_tensors_ref, response_tensors = [], []
#### get response from gpt2 and gpt2_ref
for i in range(bs):
gen_len = output_length_sampler()
output = ref_model.generate(
torch.tensor(query_tensors[i]).unsqueeze(dim=0).to(device), max_new_tokens=gen_len, **gen_kwargs
).squeeze()[-gen_len:]
response_tensors_ref.append(output)
output = model.generate(
torch.tensor(query_tensors[i]).unsqueeze(dim=0).to(device), max_new_tokens=gen_len, **gen_kwargs
).squeeze()[-gen_len:]
response_tensors.append(output)
#### decode responses
game_data["response (before)"] = [tokenizer.decode(response_tensors_ref[i]) for i in range(bs)]
game_data["response (after)"] = [tokenizer.decode(response_tensors[i]) for i in range(bs)]
#### sentiment analysis of query/response pairs before/after
texts = [q + r for q, r in zip(game_data["query"], game_data["response (before)"])]
game_data["rewards (before)"] = [output[1]["score"] for output in sentiment_pipe(texts, **sent_kwargs)]
texts = [q + r for q, r in zip(game_data["query"], game_data["response (after)"])]
game_data["rewards (after)"] = [output[1]["score"] for output in sentiment_pipe(texts, **sent_kwargs)]
# store results in a dataframe
df_results = pd.DataFrame(game_data)
df_resultsprint("mean:")
display(df_results[["rewards (before)", "rewards (after)"]].mean())
print()
print("median:")
display(df_results[["rewards (before)", "rewards (after)"]].median())model.save_pretrained("gpt2-imdb-pos-v2", push_to_hub=True)
tokenizer.save_pretrained("gpt2-imdb-pos-v2", push_to_hub=True)
| 0
|
hf_public_repos/trl/examples
|
hf_public_repos/trl/examples/notebooks/best_of_n.ipynb
|
%pip install transformers trlimport torch
import pandas as pd
from transformers import pipeline, AutoTokenizer
from datasets import load_dataset
from trl import AutoModelForCausalLMWithValueHead
from trl.core import LengthSampler
device = 0 if torch.cuda.is_available() else "cpu"ref_model_name = "lvwerra/gpt2-imdb"
model_name = "lvwerra/gpt2-imdb-pos-v2"
reward_model = "lvwerra/distilbert-imdb"
N_BEST_OF = 4model = AutoModelForCausalLMWithValueHead.from_pretrained(model_name)
ref_model = AutoModelForCausalLMWithValueHead.from_pretrained(ref_model_name)
reward_pipe = pipeline("sentiment-analysis", model=reward_model, device=device)
tokenizer = AutoTokenizer.from_pretrained(ref_model_name)
tokenizer.pad_token = tokenizer.eos_token
# cuda-ize models
model.cuda()
ref_model.cuda()def build_dataset(tokenizer, dataset_name="imdb", input_min_text_length=2, input_max_text_length=8):
# load imdb with datasets
ds = load_dataset(dataset_name, split="train")
ds = ds.rename_columns({"text": "review"})
ds = ds.filter(lambda x: len(x["review"]) > 200, batched=False)
input_size = LengthSampler(input_min_text_length, input_max_text_length)
def tokenize(sample):
sample["input_ids"] = tokenizer.encode(sample["review"])[: input_size()]
sample["query"] = tokenizer.decode(sample["input_ids"])
return sample
ds = ds.map(tokenize, batched=False)
ds.set_format(type="torch")
return ds
dataset = build_dataset(tokenizer)gen_kwargs = {"min_length": -1, "top_k": 0.0, "top_p": 1.0, "do_sample": True, "pad_token_id": tokenizer.eos_token_id}
sent_kwargs = {"top_k": None, "function_to_apply": "none", "batch_size": 16}output_min_length = 4
output_max_length = 16
output_length_sampler = LengthSampler(output_min_length, output_max_length)
#### get a batch from the dataset
bs = 16
output_data = dict()
dataset.set_format("pandas")
df_batch = dataset[:].sample(bs)
output_data["query"] = df_batch["query"].tolist()
query_tensors = df_batch["input_ids"].tolist()
# :: [Resp]
response_tensors_ref, response_tensors = [], []
# :: [[Resp]]
response_tensors_best_of = []for i in range(bs):
gen_len = output_length_sampler()
query = torch.tensor(query_tensors[i])
output = ref_model.generate(query.unsqueeze(dim=0).to(device), max_new_tokens=gen_len, **gen_kwargs).squeeze()
response_tensors_ref.append(tokenizer.decode(output))
output = model.generate(query.unsqueeze(dim=0).to(device), max_new_tokens=gen_len, **gen_kwargs).squeeze()
response_tensors.append(tokenizer.decode(output))
# generating copies of the same query for the Best-of-n sampling
queries = query.repeat((N_BEST_OF, 1))
output = ref_model.generate(queries.to(device), max_new_tokens=gen_len, **gen_kwargs).squeeze()
response_tensors_best_of.append(tokenizer.batch_decode(output))scores_ref = [output[0]["score"] for output in reward_pipe(response_tensors_ref, **sent_kwargs)]
scores = [output[0]["score"] for output in reward_pipe(response_tensors, **sent_kwargs)]
scores_best_of = []
for i, response in enumerate(response_tensors_best_of):
# base_score = scores_ref[i]
scores_best_of.append(torch.tensor([output[0]["score"] for output in reward_pipe(response, **sent_kwargs)]))output_data["response (ref)"] = response_tensors_ref
output_data["scores (ref)"] = scores_ref
output_data["response (RLHF)"] = response_tensors
output_data["scores (RLHF)"] = scores
output_data["response (best_of)"] = [
response_tensors_best_of[i][a.argmax().item()] for i, a in enumerate(scores_best_of)
]
output_data["scores (best_of)"] = [a.max().item() for a in scores_best_of]
# store results in a dataframe
df_results = pd.DataFrame(output_data)
df_results
| 0
|
hf_public_repos/trl/examples
|
hf_public_repos/trl/examples/notebooks/gpt2-sentiment-control.ipynb
|
%load_ext autoreload
%autoreload 2import random
import torch
import wandb
import time
import os
from tqdm import tqdm
import numpy as np
import pandas as pd
from random import choices
import matplotlib.pyplot as plt
tqdm.pandas()
from datasets import load_dataset
from transformers import AutoTokenizer, pipeline
from trl import PPOTrainer, PPOConfig, AutoModelForCausalLMWithValueHead, create_reference_modelsentiment_pipe_kwargs = {"top_k": None, "function_to_apply": "none"}
config = PPOConfig(
model_name="lvwerra/gpt2-imdb", steps=51200, learning_rate=1.41e-5, remove_unused_columns=False, log_with="wandb"
)
txt_in_len = 5
txt_out_len = 20
seed = 1np.random.seed(seed)gpt2_model = AutoModelForCausalLMWithValueHead.from_pretrained(config.model_name)
gpt2_model_ref = create_reference_model(gpt2_model)
gpt2_tokenizer = AutoTokenizer.from_pretrained(config.model_name)
gpt2_tokenizer.pad_token = gpt2_tokenizer.eos_token# create the dataset
#
dataset = load_dataset("imdb", split="train")
dataset = dataset.rename_columns({"text": "review", "label": "sentiment"})
# make sure the comments are are at least 500 and trim to 1000
dataset = dataset.filter(lambda x: len(x["review"]) > 500, batched=False)
dataset = dataset.map(lambda x: {"review": x["review"][:1000]}, batched=False)
datasetdataset = dataset.map(
lambda x: {"input_ids": gpt2_tokenizer.encode(" " + x["review"], return_tensors="pt")[0, :txt_in_len]},
batched=False,
)
dataset = dataset.map(lambda x: {"query": gpt2_tokenizer.decode(x["input_ids"])}, batched=False)
dataset = dataset[:20480]
from datasets import Dataset
dataset = Dataset.from_dict(dataset)
dataset.set_format("pytorch")dataset[3]["input_ids"]def collator(data):
return dict((key, [d[key] for d in data]) for key in data[0])ppo_trainer = PPOTrainer(config, gpt2_model, gpt2_model_ref, gpt2_tokenizer, dataset, data_collator=collator)if ppo_trainer.accelerator.num_processes == 1:
device = 0 if torch.cuda.is_available() else "cpu" # to avoid a `pipeline` bug
else:
device = ppo_trainer.accelerator.device
sentiment_pipe = pipeline("sentiment-analysis", "lvwerra/distilbert-imdb", device=device)text = "this movie was really bad!!"
output = sentiment_pipe(text, **sentiment_pipe_kwargs)
outputtext = "this movie was really good!!"
output = sentiment_pipe(text, **sentiment_pipe_kwargs)
outputtext = "this movie was a documentary"
output = sentiment_pipe(text, **sentiment_pipe_kwargs)
outputdef extract_pipe_output(outputs):
positive_logits = []
for out in outputs:
for element in out:
if element["label"] == "POSITIVE":
positive_logits.append(torch.tensor(element["score"]))
return positive_logitsoutput[1]["score"]ctrl_str = ["[negative]", "[neutral]", "[positive]"]
device = torch.device("cuda" if torch.cuda.is_available() else "cpu") # this should be handled by accelerate
ctrl_tokens = dict((s, gpt2_tokenizer.encode(s, return_tensors="pt").squeeze().to(device)) for s in ctrl_str)ctrl_tokensdef pos_logit_to_reward(logit, task):
"""
Take the positive sentiment logit and scale it for the task.
task [negative]: reward = -logit
task [neutral]: reward = -2*abs(logit)+4
task [positive]: reward = logit
"""
for i in range(len(logit)):
if task[i] == "[negative]":
logit[i] = -logit[i]
elif task[i] == "[neutral]":
logit[i] = -2 * torch.abs(logit[i]) + 4
elif task[i] == "[positive]":
pass
else:
raise ValueError("task has to be in [0, 1, 2]!")
return logitprint(ctrl_str)pos_logit_to_reward(torch.Tensor([4, 4, 4]), ctrl_str)pos_logit_to_reward(torch.Tensor([-4, -4, -4]), ctrl_str)pos_logit_to_reward(torch.Tensor([0, 0, 0]), ctrl_str)generation_kwargs = {
"min_length": -1,
"top_k": 0.0,
"top_p": 1.0,
"do_sample": True,
"pad_token_id": gpt2_tokenizer.eos_token_id,
"max_new_tokens": txt_out_len,
"eos_token_id": -1,
}for epoch in range(2):
for batch in tqdm(ppo_trainer.dataloader):
(logs, game_data,) = (
dict(),
dict(),
)
#### prepend a random control token
task_list = choices(ctrl_str, k=config.batch_size)
game_data["query"] = [t + q for t, q in zip(task_list, batch["query"])]
query_tensors = [torch.cat((ctrl_tokens[t], input_ids)) for t, input_ids in zip(task_list, batch["input_ids"])]
#### get response from gpt2
response_tensors = []
for query in query_tensors:
response = ppo_trainer.generate(query, **generation_kwargs)
response_tensors.append(response.squeeze()[-txt_out_len:])
game_data["response"] = [gpt2_tokenizer.decode(r.squeeze()) for r in response_tensors]
#### sentiment analysis
texts = [q + r for q, r in zip(batch["query"], game_data["response"])]
logits = extract_pipe_output(sentiment_pipe(texts, **sentiment_pipe_kwargs))
rewards = pos_logit_to_reward(logits, task_list)
#### Run PPO training
t = time.time()
stats = ppo_trainer.step(query_tensors, response_tensors, rewards)
for cs in ctrl_str:
key = "env/reward_" + cs.strip("[]")
stats[key] = np.mean([r.cpu().numpy() for r, t in zip(rewards, task_list) if t == cs])
ppo_trainer.log_stats(stats, game_data, rewards)for ctrl_s in ctrl_str:
plt.hist(
[r for r, t in zip(logs["env/reward_dist"], task_list) if t == ctrl_s], density=True, alpha=0.5, label=ctrl_s
)
plt.legend(loc="best")
plt.title("reward distribution")
plt.grid(True)
plt.show()gpt2_model.save_pretrained("gpt2-imdb-ctrl")
gpt2_tokenizer.save_pretrained("gpt2-imdb-ctrl")
| 0
|
hf_public_repos/trl/examples
|
hf_public_repos/trl/examples/scripts/ppo.py
|
# coding=utf-8
# Copyright 2023 The HuggingFace Inc. team. All rights reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
from dataclasses import dataclass, field
from typing import Optional
import torch
import tyro
from accelerate import Accelerator
from datasets import load_dataset
from peft import LoraConfig
from tqdm import tqdm
from transformers import AutoTokenizer, pipeline
from trl import AutoModelForCausalLMWithValueHead, AutoModelForSeq2SeqLMWithValueHead, PPOConfig, PPOTrainer, set_seed
from trl.core import LengthSampler
from trl.import_utils import is_xpu_available
tqdm.pandas()
@dataclass
class ScriptArguments:
ppo_config: PPOConfig = field(
default_factory=lambda: PPOConfig(
model_name="lvwerra/gpt2-imdb",
query_dataset="imdb",
reward_model="sentiment-analysis:lvwerra/distilbert-imdb",
learning_rate=1.41e-5,
log_with=None,
mini_batch_size=128,
batch_size=128,
gradient_accumulation_steps=1,
early_stopping=False,
target_kl=6.0,
kl_penalty="kl",
seed=0,
use_score_scaling=False,
use_score_norm=False,
score_clip=None,
)
)
use_seq2seq: bool = False
"""whether to use seq2seq models"""
use_peft: bool = False
"""whether to use peft"""
peft_config: Optional[LoraConfig] = field(
default_factory=lambda: LoraConfig(
r=16,
lora_alpha=16,
bias="none",
task_type="CAUSAL_LM",
),
)
trust_remote_code: bool = field(default=False, metadata={"help": "Enable `trust_remote_code`"})
args = tyro.cli(ScriptArguments)
# We then define the arguments to pass to the sentiment analysis pipeline.
# We set `return_all_scores` to True to get the sentiment score for each token.
sent_kwargs = {"return_all_scores": True, "function_to_apply": "none", "batch_size": 16}
trl_model_class = AutoModelForCausalLMWithValueHead if not args.use_seq2seq else AutoModelForSeq2SeqLMWithValueHead
# Below is an example function to build the dataset. In our case, we use the IMDB dataset
# from the `datasets` library. One should customize this function to train the model on
# its own dataset.
def build_dataset(config, query_dataset, input_min_text_length=2, input_max_text_length=8):
"""
Build dataset for training. This builds the dataset from `load_dataset`, one should
customize this function to train the model on its own dataset.
Args:
query_dataset (`str`):
The name of the dataset to be loaded.
Returns:
dataloader (`torch.utils.data.DataLoader`):
The dataloader for the dataset.
"""
tokenizer = AutoTokenizer.from_pretrained(config.model_name)
tokenizer.pad_token = tokenizer.eos_token
# load imdb with datasets
ds = load_dataset(query_dataset, split="train")
ds = ds.rename_columns({"text": "review"})
ds = ds.filter(lambda x: len(x["review"]) > 200, batched=False)
input_size = LengthSampler(input_min_text_length, input_max_text_length)
def tokenize(sample):
sample["input_ids"] = tokenizer.encode(sample["review"])[: input_size()]
sample["query"] = tokenizer.decode(sample["input_ids"])
return sample
ds = ds.map(tokenize, batched=False)
ds.set_format(type="torch")
return ds
# We retrieve the dataloader by calling the `build_dataset` function.
dataset = build_dataset(args.ppo_config, args.ppo_config.query_dataset)
def collator(data):
return dict((key, [d[key] for d in data]) for key in data[0])
# set seed before initializing value head for deterministic eval
set_seed(args.ppo_config.seed)
# Now let's build the model, the reference model, and the tokenizer.
if not args.use_peft:
ref_model = trl_model_class.from_pretrained(args.ppo_config.model_name, trust_remote_code=args.trust_remote_code)
device_map = None
peft_config = None
else:
peft_config = args.peft_config
ref_model = None
# Copy the model to each device
device_map = {"": Accelerator().local_process_index}
model = trl_model_class.from_pretrained(
args.ppo_config.model_name,
trust_remote_code=args.trust_remote_code,
device_map=device_map,
peft_config=peft_config,
)
tokenizer = AutoTokenizer.from_pretrained(args.ppo_config.model_name)
# Some tokenizers like GPT-2's don't have a padding token by default, so we set one here.
tokenizer.pad_token_id = tokenizer.eos_token_id
# We then build the PPOTrainer, passing the model, the reference model, the tokenizer
ppo_trainer = PPOTrainer(args.ppo_config, model, ref_model, tokenizer, dataset=dataset, data_collator=collator)
# We then build the sentiment analysis pipeline, passing the model name and the
# sentiment analysis pipeline arguments. Let's also make sure to set the device
# to the same device as the PPOTrainer.
device = ppo_trainer.accelerator.device
if ppo_trainer.accelerator.num_processes == 1:
if is_xpu_available():
device = "xpu:0"
else:
device = 0 if torch.cuda.is_available() else "cpu" # to avoid a `pipeline` bug
ds_plugin = ppo_trainer.accelerator.state.deepspeed_plugin
task, model_name = args.ppo_config.reward_model.split(":")
if ds_plugin is not None and ds_plugin.is_zero3_init_enabled():
with ds_plugin.zero3_init_context_manager(enable=False):
sentiment_pipe = pipeline(task, model=model_name, device=device)
else:
sentiment_pipe = pipeline(task, model=model_name, device=device)
# Some tokenizers like GPT-2's don't have a padding token by default, so we set one here.
if sentiment_pipe.tokenizer.pad_token_id is None:
sentiment_pipe.tokenizer.pad_token_id = tokenizer.pad_token_id
if sentiment_pipe.model.config.pad_token_id is None:
sentiment_pipe.model.config.pad_token_id = tokenizer.pad_token_id
# We then define the arguments to pass to the `generate` function. These arguments
# are passed to the `generate` function of the PPOTrainer, which is a wrapper around
# the `generate` function of the trained model.
generation_kwargs = {
"min_length": -1,
"top_k": 0.0,
"top_p": 1.0,
"do_sample": True,
"pad_token_id": tokenizer.eos_token_id,
"max_new_tokens": 32,
}
for epoch, batch in tqdm(enumerate(ppo_trainer.dataloader)):
query_tensors = batch["input_ids"]
# Get response from gpt2
response_tensors, ref_response_tensors = ppo_trainer.generate(
query_tensors, return_prompt=False, generate_ref_response=True, **generation_kwargs
)
batch["response"] = tokenizer.batch_decode(response_tensors)
batch["ref_response"] = tokenizer.batch_decode(ref_response_tensors)
# Compute sentiment score
texts = [q + r for q, r in zip(batch["query"], batch["response"])]
pipe_outputs = sentiment_pipe(texts, **sent_kwargs)
rewards = [torch.tensor(output[1]["score"]) for output in pipe_outputs]
ref_texts = [q + r for q, r in zip(batch["query"], batch["ref_response"])]
ref_pipe_outputs = sentiment_pipe(ref_texts, **sent_kwargs)
ref_rewards = [torch.tensor(output[1]["score"]) for output in ref_pipe_outputs]
batch["ref_rewards"] = ref_rewards
# Run PPO step
stats = ppo_trainer.step(query_tensors, response_tensors, rewards)
ppo_trainer.log_stats(stats, batch, rewards, columns_to_log=["query", "response", "ref_response", "ref_rewards"])
| 0
|
hf_public_repos/trl/examples
|
hf_public_repos/trl/examples/scripts/dpo.py
|
# coding=utf-8
# Copyright 2023 The HuggingFace Inc. team. All rights reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# Note: you need to install transformers from main to run this script. See https://huggingface.co/docs/transformers/installation#install-from-source
# TODO: bump transformers version in requirements at next release.
# 0. imports
from dataclasses import dataclass, field
from typing import Dict, Optional
import torch
from datasets import Dataset, load_dataset
from transformers import AutoModelForCausalLM, AutoTokenizer, HfArgumentParser, TrainingArguments
from trl import DPOTrainer
# Define and parse arguments.
@dataclass
class ScriptArguments:
"""
The arguments for the DPO training script.
"""
# data parameters
beta: Optional[float] = field(default=0.1, metadata={"help": "the beta parameter for DPO loss"})
# training parameters
model_name_or_path: Optional[str] = field(default="gpt2", metadata={"help": "the model name"})
learning_rate: Optional[float] = field(default=1e-3, metadata={"help": "optimizer learning rate"})
per_device_train_batch_size: Optional[int] = field(default=4, metadata={"help": "batch size per device"})
gradient_accumulation_steps: Optional[int] = field(
default=1, metadata={"help": "the number of gradient accumulation steps"}
)
max_length: Optional[int] = field(default=512, metadata={"help": "max length of each sample"})
max_prompt_length: Optional[int] = field(default=128, metadata={"help": "max length of each sample's prompt"})
max_target_length: Optional[int] = field(
default=128, metadata={"help": "Only used for encoder decoder model. Max target of each sample's prompt"}
)
label_pad_token_id: Optional[int] = field(default=-100, metadata={"help": "label for non response tokens"})
max_steps: Optional[int] = field(default=1000, metadata={"help": "max number of training steps"})
# instrumentation
sanity_check: Optional[bool] = field(default=True, metadata={"help": "only train on 1000 samples"})
report_to: Optional[str] = field(
default=None,
metadata={
"help": 'The list of integrations to report the results and logs to. Supported platforms are `"azure_ml"`,'
'`"comet_ml"`, `"mlflow"`, `"neptune"`, `"tensorboard"`,`"clearml"` and `"wandb"`. '
'Use `"all"` to report to all integrations installed, `"none"` for no integrations.'
},
)
# debug argument for distributed training
ignore_bias_buffers: Optional[bool] = field(
default=False,
metadata={
"help": "fix for DDP issues with LM bias/mask buffers - invalid scalar type,`inplace operation. See"
"https://github.com/huggingface/transformers/issues/22482#issuecomment-1595790992"
},
)
gradient_checkpointing: Optional[bool] = field(
default=False, metadata={"help": "Whether to use gradient checkpointing or no"}
)
gradient_checkpointing_kwargs: Optional[dict] = field(
default=None,
metadata={
"help": "key word arguments to be passed along `torch.utils.checkpoint.checkpoint` method - e.g. `use_reentrant=False`"
},
)
def extract_anthropic_prompt(prompt_and_response):
"""Extract the anthropic prompt from a prompt and response pair."""
search_term = "\n\nAssistant:"
search_term_idx = prompt_and_response.rfind(search_term)
assert search_term_idx != -1, f"Prompt and response does not contain '{search_term}'"
return prompt_and_response[: search_term_idx + len(search_term)]
def get_hh(split: str, sanity_check: bool = False, silent: bool = False, cache_dir: str = None) -> Dataset:
"""Load the Anthropic Helpful-Harmless dataset from Hugging Face and convert it to the necessary format.
The dataset is converted to a dictionary with the following structure:
{
'prompt': List[str],
'chosen': List[str],
'rejected': List[str],
}
Prompts should be structured as follows:
\n\nHuman: <prompt>\n\nAssistant:
Multiple turns are allowed, but the prompt should always start with \n\nHuman: and end with \n\nAssistant:.
"""
dataset = load_dataset("Anthropic/hh-rlhf", split=split, cache_dir=cache_dir)
if sanity_check:
dataset = dataset.select(range(min(len(dataset), 1000)))
def split_prompt_and_responses(sample) -> Dict[str, str]:
prompt = extract_anthropic_prompt(sample["chosen"])
return {
"prompt": prompt,
"chosen": sample["chosen"][len(prompt) :],
"rejected": sample["rejected"][len(prompt) :],
}
return dataset.map(split_prompt_and_responses)
if __name__ == "__main__":
parser = HfArgumentParser(ScriptArguments)
script_args = parser.parse_args_into_dataclasses()[0]
# 1. load a pretrained model
model = AutoModelForCausalLM.from_pretrained(script_args.model_name_or_path)
if script_args.ignore_bias_buffers:
# torch distributed hack
model._ddp_params_and_buffers_to_ignore = [
name for name, buffer in model.named_buffers() if buffer.dtype == torch.bool
]
model_ref = AutoModelForCausalLM.from_pretrained(script_args.model_name_or_path)
tokenizer = AutoTokenizer.from_pretrained(script_args.model_name_or_path)
if tokenizer.pad_token is None:
tokenizer.pad_token = tokenizer.eos_token
# 2. Load the Anthropic Helpful-Harmless dataset
train_dataset = get_hh("train", sanity_check=script_args.sanity_check)
# 3. Load evaluation dataset
eval_dataset = get_hh("test", sanity_check=script_args.sanity_check)
# 4. initialize training arguments:
training_args = TrainingArguments(
per_device_train_batch_size=script_args.per_device_train_batch_size,
max_steps=script_args.max_steps,
remove_unused_columns=False,
gradient_accumulation_steps=script_args.gradient_accumulation_steps,
learning_rate=script_args.learning_rate,
evaluation_strategy="steps",
logging_first_step=True,
logging_steps=10, # match results in blog post
eval_steps=500,
output_dir="./test",
optim="rmsprop",
warmup_steps=150,
report_to=script_args.report_to,
bf16=True,
gradient_checkpointing=script_args.gradient_checkpointing,
# TODO: uncomment that on the next transformers release
# gradient_checkpointing_kwargs=script_args.gradient_checkpointing_kwargs,
)
# 5. initialize the DPO trainer
dpo_trainer = DPOTrainer(
model,
model_ref,
args=training_args,
beta=script_args.beta,
train_dataset=train_dataset,
eval_dataset=eval_dataset,
tokenizer=tokenizer,
max_length=script_args.max_length,
max_target_length=script_args.max_target_length,
max_prompt_length=script_args.max_prompt_length,
generate_during_eval=True,
)
# 6. train
dpo_trainer.train()
| 0
|
hf_public_repos/trl/examples
|
hf_public_repos/trl/examples/scripts/reward_modeling.py
|
# coding=utf-8
# Copyright 2023 The HuggingFace Inc. team. All rights reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
from dataclasses import dataclass, field
from typing import Optional
import tyro
from accelerate import Accelerator
from datasets import load_dataset
from peft import LoraConfig
from tqdm import tqdm
from transformers import AutoModelForSequenceClassification, AutoTokenizer, BitsAndBytesConfig
from trl import RewardConfig, RewardTrainer, is_xpu_available
tqdm.pandas()
@dataclass
class ScriptArguments:
model_name: str = "facebook/opt-350m"
"""the model name"""
dataset_name: str = "Anthropic/hh-rlhf"
"""the dataset name"""
dataset_text_field: str = "text"
"""the text field of the dataset"""
eval_split: str = "none"
"""the dataset split to evaluate on; default to 'none' (no evaluation)"""
load_in_8bit: bool = False
"""load the model in 8 bits precision"""
load_in_4bit: bool = False
"""load the model in 4 bits precision"""
trust_remote_code: bool = True
"""Enable `trust_remote_code`"""
reward_config: RewardConfig = field(
default_factory=lambda: RewardConfig(
output_dir="output",
per_device_train_batch_size=64,
num_train_epochs=1,
gradient_accumulation_steps=16,
gradient_checkpointing=True,
gradient_checkpointing_kwargs={"use_reentrant": False},
learning_rate=1.41e-5,
report_to="tensorboard",
remove_unused_columns=False,
optim="adamw_torch",
logging_steps=500,
evaluation_strategy="no",
max_length=512,
)
)
use_peft: bool = False
"""whether to use peft"""
peft_config: Optional[LoraConfig] = field(
default_factory=lambda: LoraConfig(
r=16,
lora_alpha=16,
bias="none",
task_type="SEQ_CLS",
modules_to_save=["scores"],
),
)
args = tyro.cli(ScriptArguments)
args.reward_config.evaluation_strategy = "steps" if args.eval_split != "none" else "no"
# Step 1: Load the model
if args.load_in_8bit and args.load_in_4bit:
raise ValueError("You can't load the model in 8 bits and 4 bits at the same time")
elif args.load_in_8bit or args.load_in_4bit:
quantization_config = BitsAndBytesConfig(load_in_8bit=args.load_in_8bit, load_in_4bit=args.load_in_4bit)
# Copy the model to each device
device_map = (
{"": f"xpu:{Accelerator().local_process_index}"}
if is_xpu_available()
else {"": Accelerator().local_process_index}
)
else:
device_map = None
quantization_config = None
model = AutoModelForSequenceClassification.from_pretrained(
args.model_name,
quantization_config=quantization_config,
device_map=device_map,
trust_remote_code=args.trust_remote_code,
num_labels=1,
)
# Step 2: Load the dataset and pre-process it
tokenizer = AutoTokenizer.from_pretrained(args.model_name)
train_dataset = load_dataset(args.dataset_name, split="train")
# Tokenize chosen/rejected pairs of inputs
# Adapt this section to your needs for custom datasets
def preprocess_function(examples):
new_examples = {
"input_ids_chosen": [],
"attention_mask_chosen": [],
"input_ids_rejected": [],
"attention_mask_rejected": [],
}
for chosen, rejected in zip(examples["chosen"], examples["rejected"]):
tokenized_chosen = tokenizer(chosen)
tokenized_rejected = tokenizer(rejected)
new_examples["input_ids_chosen"].append(tokenized_chosen["input_ids"])
new_examples["attention_mask_chosen"].append(tokenized_chosen["attention_mask"])
new_examples["input_ids_rejected"].append(tokenized_rejected["input_ids"])
new_examples["attention_mask_rejected"].append(tokenized_rejected["attention_mask"])
return new_examples
# Preprocess the dataset and filter out examples that are longer than args.max_length
train_dataset = train_dataset.map(
preprocess_function,
batched=True,
num_proc=4,
)
train_dataset = train_dataset.filter(
lambda x: len(x["input_ids_chosen"]) <= args.reward_config.max_length
and len(x["input_ids_rejected"]) <= args.reward_config.max_length
)
if args.eval_split == "none":
eval_dataset = None
else:
eval_dataset = load_dataset(args.dataset_name, split=args.eval_split)
eval_dataset = eval_dataset.map(
preprocess_function,
batched=True,
num_proc=4,
)
eval_dataset = eval_dataset.filter(
lambda x: len(x["input_ids_chosen"]) <= args.reward_config.max_length
and len(x["input_ids_rejected"]) <= args.reward_config.max_length
)
# Step 4: Define the LoraConfig
if args.use_peft:
peft_config = args.peft_config
else:
peft_config = None
# Step 5: Define the Trainer
trainer = RewardTrainer(
model=model,
tokenizer=tokenizer,
args=args.reward_config,
train_dataset=train_dataset,
eval_dataset=eval_dataset,
peft_config=peft_config,
)
trainer.train()
| 0
|
hf_public_repos/trl/examples
|
hf_public_repos/trl/examples/scripts/ddpo.py
|
# Copyright 2023 metric-space, The HuggingFace Team. All rights reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
import os
from dataclasses import dataclass, field
import numpy as np
import torch
import torch.nn as nn
import tyro
from huggingface_hub import hf_hub_download
from huggingface_hub.utils import EntryNotFoundError
from transformers import CLIPModel, CLIPProcessor
from trl import DDPOConfig, DDPOTrainer, DefaultDDPOStableDiffusionPipeline
from trl.import_utils import is_xpu_available
@dataclass
class ScriptArguments:
hf_user_access_token: str
pretrained_model: str = "runwayml/stable-diffusion-v1-5"
"""the pretrained model to use"""
pretrained_revision: str = "main"
"""the pretrained model revision to use"""
hf_hub_model_id: str = "ddpo-finetuned-stable-diffusion"
"""HuggingFace repo to save model weights to"""
hf_hub_aesthetic_model_id: str = "trl-lib/ddpo-aesthetic-predictor"
"""HuggingFace model ID for aesthetic scorer model weights"""
hf_hub_aesthetic_model_filename: str = "aesthetic-model.pth"
"""HuggingFace model filename for aesthetic scorer model weights"""
ddpo_config: DDPOConfig = field(
default_factory=lambda: DDPOConfig(
num_epochs=200,
train_gradient_accumulation_steps=1,
sample_num_steps=50,
sample_batch_size=6,
train_batch_size=3,
sample_num_batches_per_epoch=4,
per_prompt_stat_tracking=True,
per_prompt_stat_tracking_buffer_size=32,
tracker_project_name="stable_diffusion_training",
log_with="wandb",
project_kwargs={
"logging_dir": "./logs",
"automatic_checkpoint_naming": True,
"total_limit": 5,
"project_dir": "./save",
},
)
)
class MLP(nn.Module):
def __init__(self):
super().__init__()
self.layers = nn.Sequential(
nn.Linear(768, 1024),
nn.Dropout(0.2),
nn.Linear(1024, 128),
nn.Dropout(0.2),
nn.Linear(128, 64),
nn.Dropout(0.1),
nn.Linear(64, 16),
nn.Linear(16, 1),
)
@torch.no_grad()
def forward(self, embed):
return self.layers(embed)
class AestheticScorer(torch.nn.Module):
"""
This model attempts to predict the aesthetic score of an image. The aesthetic score
is a numerical approximation of how much a specific image is liked by humans on average.
This is from https://github.com/christophschuhmann/improved-aesthetic-predictor
"""
def __init__(self, *, dtype, model_id, model_filename):
super().__init__()
self.clip = CLIPModel.from_pretrained("openai/clip-vit-large-patch14")
self.processor = CLIPProcessor.from_pretrained("openai/clip-vit-large-patch14")
self.mlp = MLP()
try:
cached_path = hf_hub_download(model_id, model_filename)
except EntryNotFoundError:
cached_path = os.path.join(model_id, model_filename)
state_dict = torch.load(cached_path)
self.mlp.load_state_dict(state_dict)
self.dtype = dtype
self.eval()
@torch.no_grad()
def __call__(self, images):
device = next(self.parameters()).device
inputs = self.processor(images=images, return_tensors="pt")
inputs = {k: v.to(self.dtype).to(device) for k, v in inputs.items()}
embed = self.clip.get_image_features(**inputs)
# normalize embedding
embed = embed / torch.linalg.vector_norm(embed, dim=-1, keepdim=True)
return self.mlp(embed).squeeze(1)
def aesthetic_scorer(hub_model_id, model_filename):
scorer = AestheticScorer(
model_id=hub_model_id,
model_filename=model_filename,
dtype=torch.float32,
)
scorer = scorer.xpu() if is_xpu_available() else scorer.cuda()
def _fn(images, prompts, metadata):
images = (images * 255).round().clamp(0, 255).to(torch.uint8)
scores = scorer(images)
return scores, {}
return _fn
# list of example prompts to feed stable diffusion
animals = [
"cat",
"dog",
"horse",
"monkey",
"rabbit",
"zebra",
"spider",
"bird",
"sheep",
"deer",
"cow",
"goat",
"lion",
"frog",
"chicken",
"duck",
"goose",
"bee",
"pig",
"turkey",
"fly",
"llama",
"camel",
"bat",
"gorilla",
"hedgehog",
"kangaroo",
]
def prompt_fn():
return np.random.choice(animals), {}
def image_outputs_logger(image_data, global_step, accelerate_logger):
# For the sake of this example, we will only log the last batch of images
# and associated data
result = {}
images, prompts, _, rewards, _ = image_data[-1]
for i, image in enumerate(images):
prompt = prompts[i]
reward = rewards[i].item()
result[f"{prompt:.25} | {reward:.2f}"] = image.unsqueeze(0)
accelerate_logger.log_images(
result,
step=global_step,
)
if __name__ == "__main__":
args = tyro.cli(ScriptArguments)
pipeline = DefaultDDPOStableDiffusionPipeline(
args.pretrained_model, pretrained_model_revision=args.pretrained_revision, use_lora=True
)
trainer = DDPOTrainer(
args.ddpo_config,
aesthetic_scorer(args.hf_hub_aesthetic_model_id, args.hf_hub_aesthetic_model_filename),
prompt_fn,
pipeline,
image_samples_hook=image_outputs_logger,
)
trainer.train()
trainer.push_to_hub(args.hf_hub_model_id, token=args.hf_user_access_token)
| 0
|
hf_public_repos/trl/examples
|
hf_public_repos/trl/examples/scripts/sft.py
|
# coding=utf-8
# Copyright 2023 The HuggingFace Inc. team. All rights reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
from dataclasses import dataclass, field
from typing import Optional
import torch
from accelerate import Accelerator
from datasets import load_dataset
from peft import LoraConfig
from tqdm import tqdm
from transformers import AutoModelForCausalLM, BitsAndBytesConfig, HfArgumentParser, TrainingArguments
from trl import SFTTrainer, is_xpu_available
tqdm.pandas()
# Define and parse arguments.
@dataclass
class ScriptArguments:
"""
The name of the Casual LM model we wish to fine with SFTTrainer
"""
model_name: Optional[str] = field(default="facebook/opt-350m", metadata={"help": "the model name"})
dataset_name: Optional[str] = field(
default="timdettmers/openassistant-guanaco", metadata={"help": "the dataset name"}
)
dataset_text_field: Optional[str] = field(default="text", metadata={"help": "the text field of the dataset"})
log_with: Optional[str] = field(default="none", metadata={"help": "use 'wandb' to log with wandb"})
learning_rate: Optional[float] = field(default=1.41e-5, metadata={"help": "the learning rate"})
batch_size: Optional[int] = field(default=64, metadata={"help": "the batch size"})
seq_length: Optional[int] = field(default=512, metadata={"help": "Input sequence length"})
gradient_accumulation_steps: Optional[int] = field(
default=16, metadata={"help": "the number of gradient accumulation steps"}
)
load_in_8bit: Optional[bool] = field(default=False, metadata={"help": "load the model in 8 bits precision"})
load_in_4bit: Optional[bool] = field(default=False, metadata={"help": "load the model in 4 bits precision"})
use_peft: Optional[bool] = field(default=False, metadata={"help": "Wether to use PEFT or not to train adapters"})
trust_remote_code: Optional[bool] = field(default=False, metadata={"help": "Enable `trust_remote_code`"})
output_dir: Optional[str] = field(default="output", metadata={"help": "the output directory"})
peft_lora_r: Optional[int] = field(default=64, metadata={"help": "the r parameter of the LoRA adapters"})
peft_lora_alpha: Optional[int] = field(default=16, metadata={"help": "the alpha parameter of the LoRA adapters"})
logging_steps: Optional[int] = field(default=1, metadata={"help": "the number of logging steps"})
use_auth_token: Optional[bool] = field(default=True, metadata={"help": "Use HF auth token to access the model"})
num_train_epochs: Optional[int] = field(default=3, metadata={"help": "the number of training epochs"})
max_steps: Optional[int] = field(default=-1, metadata={"help": "the number of training steps"})
save_steps: Optional[int] = field(
default=100, metadata={"help": "Number of updates steps before two checkpoint saves"}
)
save_total_limit: Optional[int] = field(default=10, metadata={"help": "Limits total number of checkpoints."})
push_to_hub: Optional[bool] = field(default=False, metadata={"help": "Push the model to HF Hub"})
gradient_checkpointing: Optional[bool] = field(
default=False, metadata={"help": "Whether to use gradient checkpointing or no"}
)
gradient_checkpointing_kwargs: Optional[dict] = field(
default=None,
metadata={
"help": "key word arguments to be passed along `torch.utils.checkpoint.checkpoint` method - e.g. `use_reentrant=False`"
},
)
hub_model_id: Optional[str] = field(default=None, metadata={"help": "The name of the model on HF Hub"})
parser = HfArgumentParser(ScriptArguments)
script_args = parser.parse_args_into_dataclasses()[0]
# Step 1: Load the model
if script_args.load_in_8bit and script_args.load_in_4bit:
raise ValueError("You can't load the model in 8 bits and 4 bits at the same time")
elif script_args.load_in_8bit or script_args.load_in_4bit:
quantization_config = BitsAndBytesConfig(
load_in_8bit=script_args.load_in_8bit, load_in_4bit=script_args.load_in_4bit
)
# Copy the model to each device
device_map = (
{"": f"xpu:{Accelerator().local_process_index}"}
if is_xpu_available()
else {"": Accelerator().local_process_index}
)
torch_dtype = torch.bfloat16
else:
device_map = None
quantization_config = None
torch_dtype = None
model = AutoModelForCausalLM.from_pretrained(
script_args.model_name,
quantization_config=quantization_config,
device_map=device_map,
trust_remote_code=script_args.trust_remote_code,
torch_dtype=torch_dtype,
use_auth_token=script_args.use_auth_token,
)
# Step 2: Load the dataset
dataset = load_dataset(script_args.dataset_name, split="train")
# Step 3: Define the training arguments
training_args = TrainingArguments(
output_dir=script_args.output_dir,
per_device_train_batch_size=script_args.batch_size,
gradient_accumulation_steps=script_args.gradient_accumulation_steps,
learning_rate=script_args.learning_rate,
logging_steps=script_args.logging_steps,
num_train_epochs=script_args.num_train_epochs,
max_steps=script_args.max_steps,
report_to=script_args.log_with,
save_steps=script_args.save_steps,
save_total_limit=script_args.save_total_limit,
push_to_hub=script_args.push_to_hub,
hub_model_id=script_args.hub_model_id,
gradient_checkpointing=script_args.gradient_checkpointing,
# TODO: uncomment that on the next release
# gradient_checkpointing_kwargs=script_args.gradient_checkpointing_kwargs,
)
# Step 4: Define the LoraConfig
if script_args.use_peft:
peft_config = LoraConfig(
r=script_args.peft_lora_r,
lora_alpha=script_args.peft_lora_alpha,
bias="none",
task_type="CAUSAL_LM",
)
else:
peft_config = None
# Step 5: Define the Trainer
trainer = SFTTrainer(
model=model,
args=training_args,
max_seq_length=script_args.seq_length,
train_dataset=dataset,
dataset_text_field=script_args.dataset_text_field,
peft_config=peft_config,
)
trainer.train()
# Step 6: Save the model
trainer.save_model(script_args.output_dir)
| 0
|
hf_public_repos/trl/examples
|
hf_public_repos/trl/examples/scripts/ppo_multi_adapter.py
|
# coding=utf-8
# Copyright 2023 The HuggingFace Inc. team. All rights reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
from dataclasses import dataclass, field
from typing import Optional
import torch
from datasets import load_dataset
from peft import LoraConfig
from tqdm import tqdm
from transformers import AutoTokenizer, BitsAndBytesConfig, HfArgumentParser
from trl import AutoModelForCausalLMWithValueHead, PPOConfig, PPOTrainer, is_xpu_available
from trl.core import LengthSampler
input_min_text_length = 6
input_max_text_length = 12
@dataclass
class ScriptArguments:
"""
The name of the Casual LM model we wish to fine with PPO
"""
model_name: Optional[str] = field(default="huggyllama/llama-7b", metadata={"help": "the model name"})
dataset_name: Optional[str] = field(default="Anthropic/hh-rlhf", metadata={"help": "the dataset name"})
rm_adapter: Optional[str] = field(
default="trl-lib/llama-7b-hh-rm-adapter", metadata={"help": "the rm adapter name"}
)
log_with: Optional[str] = field(default=None, metadata={"help": "use 'wandb' to log with wandb"})
use_safetensors: Optional[bool] = field(default=False, metadata={"help": "Use safetensors"})
seed: Optional[int] = field(default=0, metadata={"help": "the random seed"})
use_score_scaling: Optional[bool] = field(default=False, metadata={"help": "Use score scaling"})
use_score_norm: Optional[bool] = field(
default=False, metadata={"help": "Use score normalization. Only applicable if use_score_scaling is True"}
)
score_clip: Optional[float] = field(default=None, metadata={"help": "Score clipping"})
parser = HfArgumentParser(ScriptArguments)
script_args = parser.parse_args_into_dataclasses()[0]
def create_and_prepare_dataset(tokenizer):
dataset = load_dataset(script_args.dataset_name, split="train[:1%]")
input_size = LengthSampler(input_min_text_length, input_max_text_length)
def tokenize(example):
text_size = input_size()
example["input_ids"] = tokenizer.encode(example["chosen"])[:text_size]
example["query"] = tokenizer.decode(example["input_ids"])
return example
dataset = dataset.map(tokenize, batched=False)
dataset.set_format("torch")
return dataset
lora_config = LoraConfig(
r=16,
lora_alpha=32,
lora_dropout=0.05,
bias="none",
task_type="CAUSAL_LM",
)
nf4_config = BitsAndBytesConfig(
load_in_4bit=True, bnb_4bit_quant_type="nf4", bnb_4bit_use_double_quant=True, bnb_4bit_compute_dtype=torch.bfloat16
)
model = AutoModelForCausalLMWithValueHead.from_pretrained(
script_args.model_name,
device_map={"": "xpu:0"} if is_xpu_available() else {"": 0},
peft_config=lora_config,
quantization_config=nf4_config,
reward_adapter=script_args.rm_adapter,
use_safetensors=script_args.use_safetensors,
)
tokenizer = AutoTokenizer.from_pretrained(script_args.model_name)
tokenizer.pad_token = tokenizer.eos_token
dataset = create_and_prepare_dataset(tokenizer)
def collator(data):
return dict((key, [d[key] for d in data]) for key in data[0])
config = PPOConfig(
model_name=script_args.model_name,
log_with=script_args.log_with,
learning_rate=1e-5,
batch_size=8,
mini_batch_size=2,
gradient_accumulation_steps=2,
optimize_cuda_cache=True,
seed=script_args.seed,
use_score_scaling=script_args.use_score_scaling,
use_score_norm=script_args.use_score_norm,
score_clip=script_args.score_clip,
)
ppo_trainer = PPOTrainer(
config,
model,
ref_model=None,
tokenizer=tokenizer,
dataset=dataset,
data_collator=collator,
)
generation_kwargs = {
"top_k": 0.0,
"top_p": 0.9,
"do_sample": True,
"pad_token_id": tokenizer.pad_token_id,
"max_new_tokens": 32,
}
for epoch, batch in tqdm(enumerate(ppo_trainer.dataloader)):
question_tensors = batch["input_ids"]
response_tensors = ppo_trainer.generate(
question_tensors,
return_prompt=False,
**generation_kwargs,
)
batch["response"] = tokenizer.batch_decode(response_tensors, skip_special_tokens=True)
# Compute reward score
texts = [q + r for q, r in zip(batch["query"], batch["response"])]
inputs = tokenizer(texts, padding=True, truncation=True, return_tensors="pt").to(ppo_trainer.accelerator.device)
raw_rewards = ppo_trainer.accelerator.unwrap_model(ppo_trainer.model).compute_reward_score(**inputs)
rewards = [raw_rewards[i, -1, 1] for i in range(len(raw_rewards))] # take last token
# Run PPO step
stats = ppo_trainer.step(question_tensors, response_tensors, rewards)
ppo_trainer.log_stats(stats, batch, rewards)
| 0
|
hf_public_repos/trl/examples
|
hf_public_repos/trl/examples/research_projects/README.md
|
# Research projects that use TRL
Welcome to the research projects folder! Here you can find the scripts used for some research projects that used TRL and maintained by the developers and the community (LM de-toxification, Stack-Llama, etc.). Check out the READMEs in the subfolders for more information!
- [De-detoxifying language models](https://github.com/huggingface/trl/tree/main/examples/research_projects/toxicity)
- [Stack-Llama](https://github.com/huggingface/trl/tree/main/examples/research_projects/stack_llama)
- [Stack-Llama-2](https://github.com/huggingface/trl/tree/main/examples/research_projects/stack_llama_2)
| 0
|
hf_public_repos/trl/examples/research_projects/stack_llama
|
hf_public_repos/trl/examples/research_projects/stack_llama/scripts/merge_peft_adapter.py
|
from dataclasses import dataclass, field
from typing import Optional
import torch
from peft import PeftConfig, PeftModel
from transformers import AutoModelForCausalLM, AutoModelForSequenceClassification, AutoTokenizer, HfArgumentParser
@dataclass
class ScriptArguments:
"""
The input names representing the Adapter and Base model fine-tuned with PEFT, and the output name representing the
merged model.
"""
adapter_model_name: Optional[str] = field(default=None, metadata={"help": "the adapter name"})
base_model_name: Optional[str] = field(default=None, metadata={"help": "the base model name"})
output_name: Optional[str] = field(default=None, metadata={"help": "the merged model name"})
parser = HfArgumentParser(ScriptArguments)
script_args = parser.parse_args_into_dataclasses()[0]
assert script_args.adapter_model_name is not None, "please provide the name of the Adapter you would like to merge"
assert script_args.base_model_name is not None, "please provide the name of the Base model"
assert script_args.output_name is not None, "please provide the output name of the merged model"
peft_config = PeftConfig.from_pretrained(script_args.adapter_model_name)
if peft_config.task_type == "SEQ_CLS":
# The sequence classification task is used for the reward model in PPO
model = AutoModelForSequenceClassification.from_pretrained(
script_args.base_model_name, num_labels=1, torch_dtype=torch.bfloat16
)
else:
model = AutoModelForCausalLM.from_pretrained(
script_args.base_model_name, return_dict=True, torch_dtype=torch.bfloat16
)
tokenizer = AutoTokenizer.from_pretrained(script_args.base_model_name)
# Load the PEFT model
model = PeftModel.from_pretrained(model, script_args.adapter_model_name)
model.eval()
model = model.merge_and_unload()
model.save_pretrained(f"{script_args.output_name}")
tokenizer.save_pretrained(f"{script_args.output_name}")
model.push_to_hub(f"{script_args.output_name}", use_temp_dir=False)
| 0
|
hf_public_repos/trl/examples/research_projects/stack_llama
|
hf_public_repos/trl/examples/research_projects/stack_llama/scripts/README.md
|
# RLHF pipeline for the creation of StackLLaMa: a Stack exchange llama-7b model.
There were three main steps to the training process:
1. Supervised fine-tuning of the base llama-7b model to create llama-7b-se:
- `torchrun --nnodes 1 --nproc_per_node 8 examples/research_projects/stack_llama/scripts/supervised_finetuning.py --model_path=<LLAMA_MODEL_PATH> --streaming --no_gradient_checkpointing --learning_rate 1e-5 --max_steps 5000 --output_dir ./llama-se`
2. Reward modeling using dialog pairs from the SE dataset using the llama-7b-se to create llama-7b-se-rm:
- `torchrun --nnodes 1 --nproc_per_node 8 examples/research_projects/stack_llama/scripts/reward_modeling.py --model_name=<LLAMA_SE_MODEL>`
3. RL fine-tuning of llama-7b-se with the llama-7b-se-rm reward model:
- `accelerate launch --multi_gpu --num_machines 1 --num_processes 8 examples/research_projects/stack_llama/scripts/rl_training.py --log_with=wandb --model_name=<LLAMA_SE_MODEL> --reward_model_name=<LLAMA_SE_RM_MODEL> --adafactor=False --tokenizer_name=<LLAMA_TOKENIZER> --save_freq=100 --output_max_length=128 --batch_size=8 --gradient_accumulation_steps=8 --batched_gen=True --ppo_epochs=4 --seed=0 --learning_rate=1.4e-5 --early_stopping=True --output_dir=llama-se-rl-finetune-128-8-8-1.4e-5_adam`
LoRA layers were using at all stages to reduce memory requirements.
At each stage the peft adapter layers were merged with the base model, using:
```shell
python examples/research_projects/stack_llama/scripts/merge_peft_adapter.py --adapter_model_name=XXX --base_model_name=YYY --output_name=ZZZ
```
Note that this script requires `peft>=0.3.0`.
For access to the base llama-7b model, please see Meta's [release](https://ai.facebook.com/blog/large-language-model-llama-meta-ai/) and [request form](https://docs.google.com/forms/d/e/1FAIpQLSfqNECQnMkycAp2jP4Z9TFX0cGR4uf7b_fBxjY_OjhJILlKGA/viewform).
| 0
|
hf_public_repos/trl/examples/research_projects/stack_llama
|
hf_public_repos/trl/examples/research_projects/stack_llama/scripts/rl_training.py
|
# coding=utf-8
# Copyright 2022 The HuggingFace Inc. team. All rights reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
from dataclasses import dataclass, field
from typing import Optional
import torch
from accelerate import Accelerator
from datasets import load_dataset
from peft import LoraConfig
from tqdm import tqdm
from transformers import Adafactor, AutoTokenizer, HfArgumentParser, pipeline
from trl import AutoModelForCausalLMWithValueHead, PPOConfig, PPOTrainer, set_seed
from trl.core import LengthSampler
tqdm.pandas()
@dataclass
class ScriptArguments:
"""
The name of the Casual LM model we wish to fine with PPO
"""
# NOTE: gpt2 models use Conv1D instead of Linear layers which are not yet supported in 8 bit mode
# models like gpt-neo* models are more suitable.
model_name: Optional[str] = field(default="", metadata={"help": "the model name"})
tokenizer_name: Optional[str] = field(default="", metadata={"help": "the tokenizer name"})
reward_model_name: Optional[str] = field(default="", metadata={"help": "the reward model name"})
log_with: Optional[str] = field(default=None, metadata={"help": "use 'wandb' to log with wandb"})
learning_rate: Optional[float] = field(default=1.41e-5, metadata={"help": "the learning rate"})
output_max_length: Optional[int] = field(default=128, metadata={"help": "maximum length for generation"})
mini_batch_size: Optional[int] = field(default=1, metadata={"help": "the PPO minibatch size"})
batch_size: Optional[int] = field(default=32, metadata={"help": "the batch size"})
ppo_epochs: Optional[int] = field(default=4, metadata={"help": "the number of ppo epochs"})
gradient_accumulation_steps: Optional[int] = field(
default=4, metadata={"help": "the number of gradient accumulation steps"}
)
adafactor: Optional[bool] = field(default=False, metadata={"help": "whether to use the adafactor optimizer"})
early_stopping: Optional[bool] = field(default=False, metadata={"help": "whether to early stop"})
target_kl: Optional[float] = field(default=0.1, metadata={"help": "kl target for early stopping"})
reward_baseline: Optional[float] = field(
default=0.0,
metadata={"help": "a baseline value that is subtracted from the reward"},
)
batched_gen: Optional[bool] = field(default=False, metadata={"help": "whether to use the batched text gen"})
save_freq: Optional[int] = field(default=None, metadata={"help": "n steps to save the model"})
output_dir: Optional[str] = field(default="runs/", metadata={"help": "n steps to save the model"})
seed: Optional[int] = field(default=0, metadata={"help": "the seed"})
steps: Optional[int] = field(default=20000, metadata={"help": "number of epochs"})
init_kl_coef: Optional[float] = field(
default=0.2,
metadata={"help": "Initial KL penalty coefficient (used for adaptive and linear control)"},
)
adap_kl_ctrl: Optional[bool] = field(default=True, metadata={"help": "Use adaptive KL control, otherwise linear"})
parser = HfArgumentParser(ScriptArguments)
script_args: ScriptArguments = parser.parse_args_into_dataclasses()[0]
reward_model_name = script_args.reward_model_name
dataset_name = "lvwerra/stack-exchange-paired"
config = PPOConfig(
steps=script_args.steps,
model_name=script_args.model_name,
learning_rate=script_args.learning_rate,
log_with=script_args.log_with,
batch_size=script_args.batch_size,
mini_batch_size=script_args.mini_batch_size,
gradient_accumulation_steps=script_args.gradient_accumulation_steps,
optimize_cuda_cache=True,
early_stopping=script_args.early_stopping,
target_kl=script_args.target_kl,
ppo_epochs=script_args.ppo_epochs,
seed=script_args.seed,
init_kl_coef=script_args.init_kl_coef,
adap_kl_ctrl=script_args.adap_kl_ctrl,
)
train_dataset = load_dataset("lvwerra/stack-exchange-paired", data_dir="data/rl", split="train")
train_dataset = train_dataset.select(range(100000))
original_columns = train_dataset.column_names
# We then define the arguments to pass to the sentiment analysis pipeline.
# We set `return_all_scores` to True to get the sentiment score for each token.
sent_kwargs = {
"return_all_scores": True,
"function_to_apply": "none",
"batch_size": 16,
"truncation": True,
}
tokenizer = AutoTokenizer.from_pretrained(script_args.tokenizer_name)
# GPT-2 tokenizer has a pad token, but it is not eos_token by default. We need to set it to eos_token.
# only for this model.
if getattr(tokenizer, "pad_token", None) is None:
tokenizer.pad_token = tokenizer.eos_token
# Below is an example function to build the dataset. In our case, we use the IMDB dataset
# from the `datasets` library. One should customize this function to train the model on
# its own dataset.
def build_dataset(
tokenizer,
dataset_name="lvwerra/stack-exchange-paired",
):
"""
Build dataset for training. This builds the dataset from `load_dataset`, one should
customize this function to train the model on its own dataset.
Args:
dataset_name (`str`):
The name of the dataset to be loaded.
Returns:
dataloader (`torch.utils.data.DataLoader`):
The dataloader for the dataset.
"""
num_proc = 24
def preprocess_function(examples):
new_examples = {
"query": [],
"input_ids": [],
}
for question in examples["question"]:
query = "Question: " + question + "\n\nAnswer: "
tokenized_question = tokenizer(query, truncation=True)
new_examples["query"].append(query)
new_examples["input_ids"].append(tokenized_question["input_ids"])
return new_examples
ds = train_dataset.map(
preprocess_function,
batched=True,
num_proc=num_proc,
remove_columns=original_columns,
)
ds = ds.filter(lambda x: len(x["input_ids"]) < 512, batched=False)
ds.set_format(type="torch")
return ds
# We retrieve the dataloader by calling the `build_dataset` function.
dataset = build_dataset(tokenizer)
def collator(data):
return dict((key, [d[key] for d in data]) for key in data[0])
# set seed before initializing value head for deterministic eval
set_seed(config.seed)
# Now let's build the model, the reference model, and the tokenizer.
current_device = Accelerator().local_process_index
lora_config = LoraConfig(
r=16,
lora_alpha=32,
lora_dropout=0.05,
bias="none",
task_type="CAUSAL_LM",
)
model = AutoModelForCausalLMWithValueHead.from_pretrained(
config.model_name,
load_in_8bit=True,
device_map={"": current_device},
peft_config=lora_config,
)
optimizer = None
if script_args.adafactor:
optimizer = Adafactor(
filter(lambda p: p.requires_grad, model.parameters()),
scale_parameter=False,
relative_step=False,
warmup_init=False,
lr=config.learning_rate,
)
# We then build the PPOTrainer, passing the model, the reference model, the tokenizer
ppo_trainer = PPOTrainer(
config,
model,
ref_model=None,
tokenizer=tokenizer,
dataset=dataset,
data_collator=collator,
optimizer=optimizer,
)
# We then build the sentiment analysis pipeline using our reward model, passing the
# model name and the sentiment analysis pipeline arguments. Let's also make sure to
# set the device to the same device as the PPOTrainer.
device = ppo_trainer.accelerator.device
if ppo_trainer.accelerator.num_processes == 1:
device = 0 if torch.cuda.is_available() else "cpu" # to avoid a ` pipeline` bug
sentiment_pipe = pipeline(
"sentiment-analysis",
model=reward_model_name,
device_map={"": current_device},
model_kwargs={"load_in_8bit": True},
tokenizer=tokenizer,
return_token_type_ids=False,
)
# We then define the arguments to pass to the `generate` function. These arguments
# are passed to the `generate` function of the PPOTrainer, which is a wrapper around
# the `generate` function of the trained model.
generation_kwargs = {
# "min_length": -1,
"top_k": 0.0,
"top_p": 1.0,
"do_sample": True,
"pad_token_id": tokenizer.pad_token_id,
"eos_token_id": 100_000,
}
output_min_length = 32
output_max_length = script_args.output_max_length
output_length_sampler = LengthSampler(output_min_length, output_max_length)
for epoch, batch in tqdm(enumerate(ppo_trainer.dataloader)):
if epoch >= config.total_ppo_epochs:
break
question_tensors = batch["input_ids"]
response_tensors = ppo_trainer.generate(
question_tensors,
return_prompt=False,
length_sampler=output_length_sampler,
**generation_kwargs,
)
batch["response"] = tokenizer.batch_decode(response_tensors, skip_special_tokens=True)
# Compute reward score (using the sentiment analysis pipeline)
texts = [q + r for q, r in zip(batch["query"], batch["response"])]
pipe_outputs = sentiment_pipe(texts, **sent_kwargs)
rewards = [torch.tensor(output[0]["score"] - script_args.reward_baseline) for output in pipe_outputs]
# Run PPO step
stats = ppo_trainer.step(question_tensors, response_tensors, rewards)
ppo_trainer.log_stats(stats, batch, rewards)
if script_args.save_freq and epoch and epoch % script_args.save_freq == 0:
ppo_trainer.save_pretrained(script_args.output_dir + f"step_{epoch}")
| 0
|
hf_public_repos/trl/examples/research_projects/stack_llama
|
hf_public_repos/trl/examples/research_projects/stack_llama/scripts/supervised_finetuning.py
|
import argparse
import os
from accelerate import Accelerator
from datasets import load_dataset
from peft import LoraConfig
from tqdm import tqdm
from transformers import AutoModelForCausalLM, AutoTokenizer, TrainingArguments, logging, set_seed
from trl import SFTTrainer
from trl.trainer import ConstantLengthDataset
"""
Fine-Tune Llama-7b on SE paired dataset
"""
def get_args():
parser = argparse.ArgumentParser()
parser.add_argument("--model_path", type=str, default="")
parser.add_argument("--dataset_name", type=str, default="lvwerra/stack-exchange-paired")
parser.add_argument("--subset", type=str, default="data/finetune")
parser.add_argument("--split", type=str, default="train")
parser.add_argument("--size_valid_set", type=int, default=4000)
parser.add_argument("--streaming", action="store_true")
parser.add_argument("--shuffle_buffer", type=int, default=5000)
parser.add_argument("--seq_length", type=int, default=1024)
parser.add_argument("--max_steps", type=int, default=10000)
parser.add_argument("--batch_size", type=int, default=4)
parser.add_argument("--gradient_accumulation_steps", type=int, default=1)
parser.add_argument("--eos_token_id", type=int, default=49152)
parser.add_argument("--learning_rate", type=float, default=1e-4)
parser.add_argument("--lr_scheduler_type", type=str, default="cosine")
parser.add_argument("--num_warmup_steps", type=int, default=100)
parser.add_argument("--weight_decay", type=float, default=0.05)
parser.add_argument("--local_rank", type=int, default=0)
parser.add_argument("--no_fp16", action="store_false")
parser.add_argument("--bf16", action="store_true", default=False)
parser.add_argument("--no_gradient_checkpointing", action="store_false", default=False)
parser.add_argument("--seed", type=int, default=0)
parser.add_argument("--num_workers", type=int, default=None)
parser.add_argument("--output_dir", type=str, default="./checkpoints")
parser.add_argument("--log_freq", default=1, type=int)
parser.add_argument("--eval_freq", default=1000, type=int)
parser.add_argument("--save_freq", default=1000, type=int)
return parser.parse_args()
def chars_token_ratio(dataset, tokenizer, nb_examples=400):
"""
Estimate the average number of characters per token in the dataset.
"""
total_characters, total_tokens = 0, 0
for _, example in tqdm(zip(range(nb_examples), iter(dataset)), total=nb_examples):
text = prepare_sample_text(example)
total_characters += len(text)
if tokenizer.is_fast:
total_tokens += len(tokenizer(text).tokens())
else:
total_tokens += len(tokenizer.tokenize(text))
return total_characters / total_tokens
def print_trainable_parameters(model):
"""
Prints the number of trainable parameters in the model.
"""
trainable_params = 0
all_param = 0
for _, param in model.named_parameters():
all_param += param.numel()
if param.requires_grad:
trainable_params += param.numel()
print(
f"trainable params: {trainable_params} || all params: {all_param} || trainable%: {100 * trainable_params / all_param}"
)
def prepare_sample_text(example):
"""Prepare the text from a sample of the dataset."""
text = f"Question: {example['question']}\n\nAnswer: {example['response_j']}"
return text
def create_datasets(tokenizer, args):
dataset = load_dataset(
args.dataset_name,
data_dir=args.subset,
split=args.split,
use_auth_token=True,
num_proc=args.num_workers if not args.streaming else None,
streaming=args.streaming,
)
if args.streaming:
print("Loading the dataset in streaming mode")
valid_data = dataset.take(args.size_valid_set)
train_data = dataset.skip(args.size_valid_set)
train_data = train_data.shuffle(buffer_size=args.shuffle_buffer, seed=args.seed)
else:
dataset = dataset.train_test_split(test_size=0.005, seed=args.seed)
train_data = dataset["train"]
valid_data = dataset["test"]
print(f"Size of the train set: {len(train_data)}. Size of the validation set: {len(valid_data)}")
chars_per_token = chars_token_ratio(train_data, tokenizer)
print(f"The character to token ratio of the dataset is: {chars_per_token:.2f}")
train_dataset = ConstantLengthDataset(
tokenizer,
train_data,
formatting_func=prepare_sample_text,
infinite=True,
seq_length=args.seq_length,
chars_per_token=chars_per_token,
)
valid_dataset = ConstantLengthDataset(
tokenizer,
valid_data,
formatting_func=prepare_sample_text,
infinite=False,
seq_length=args.seq_length,
chars_per_token=chars_per_token,
)
return train_dataset, valid_dataset
def run_training(args, train_data, val_data):
print("Loading the model")
lora_config = LoraConfig(
r=16,
lora_alpha=32,
lora_dropout=0.05,
bias="none",
task_type="CAUSAL_LM",
)
train_data.start_iteration = 0
print("Starting main loop")
training_args = TrainingArguments(
output_dir=args.output_dir,
dataloader_drop_last=True,
evaluation_strategy="steps",
max_steps=args.max_steps,
eval_steps=args.eval_freq,
save_steps=args.save_freq,
logging_steps=args.log_freq,
per_device_train_batch_size=args.batch_size,
per_device_eval_batch_size=args.batch_size,
learning_rate=args.learning_rate,
lr_scheduler_type=args.lr_scheduler_type,
warmup_steps=args.num_warmup_steps,
gradient_accumulation_steps=args.gradient_accumulation_steps,
gradient_checkpointing=not args.no_gradient_checkpointing,
fp16=not args.no_fp16,
bf16=args.bf16,
weight_decay=args.weight_decay,
run_name="llama-7b-finetuned",
report_to="wandb",
ddp_find_unused_parameters=False,
)
model = AutoModelForCausalLM.from_pretrained(
args.model_path, load_in_8bit=True, device_map={"": Accelerator().process_index}
)
trainer = SFTTrainer(
model=model,
args=training_args,
train_dataset=train_data,
eval_dataset=val_data,
peft_config=lora_config,
packing=True,
)
print_trainable_parameters(trainer.model)
print("Training...")
trainer.train()
print("Saving last checkpoint of the model")
trainer.model.save_pretrained(os.path.join(args.output_dir, "final_checkpoint/"))
def main(args):
tokenizer = AutoTokenizer.from_pretrained(args.model_path)
train_dataset, eval_dataset = create_datasets(tokenizer, args)
run_training(args, train_dataset, eval_dataset)
if __name__ == "__main__":
args = get_args()
assert args.model_path != "", "Please provide the llama model path"
set_seed(args.seed)
os.makedirs(args.output_dir, exist_ok=True)
logging.set_verbosity_error()
main(args)
| 0
|
hf_public_repos/trl/examples/research_projects/stack_llama
|
hf_public_repos/trl/examples/research_projects/stack_llama/scripts/reward_modeling.py
|
from dataclasses import dataclass, field
from typing import Any, Dict, List, Optional, Union
import evaluate
import numpy as np
import torch
import torch.nn as nn
from datasets import load_dataset
from peft import LoraConfig, TaskType, get_peft_model
from transformers import (
AutoModelForSequenceClassification,
AutoTokenizer,
HfArgumentParser,
PreTrainedTokenizerBase,
Trainer,
TrainerCallback,
TrainingArguments,
)
from transformers.utils import PaddingStrategy
# Define and parse arguments.
@dataclass
class ScriptArguments:
"""
These arguments vary depending on how many GPUs you have, what their capacity and features are, and what size model you want to train.
"""
local_rank: Optional[int] = field(default=-1, metadata={"help": "Used for multi-gpu"})
resume_from_checkpoint: Optional[bool] = field(
default=False,
metadata={"help": "If you want to resume training where it left off."},
)
deepspeed: Optional[str] = field(
default=None,
metadata={
"help": "Path to deepspeed config if using deepspeed. You may need this if the model that you want to train doesn't fit on a single GPU."
},
)
per_device_train_batch_size: Optional[int] = field(default=4)
per_device_eval_batch_size: Optional[int] = field(default=1)
gradient_accumulation_steps: Optional[int] = field(default=1)
learning_rate: Optional[float] = field(default=2e-5)
weight_decay: Optional[float] = field(default=0.001)
model_name: Optional[str] = field(
default="gpt2",
metadata={
"help": "The model that you want to train from the Hugging Face hub. E.g. gpt2, gpt2-xl, bert, etc."
},
)
tokenizer_name: Optional[str] = field(
default=None,
metadata={
"help": "The tokenizer for your model, if left empty will use the default for your model",
},
)
bf16: Optional[bool] = field(
default=True,
metadata={
"help": "This essentially cuts the training time in half if you want to sacrifice a little precision and have a supported GPU."
},
)
num_train_epochs: Optional[int] = field(
default=1,
metadata={"help": "The number of training epochs for the reward model."},
)
train_subset: Optional[int] = field(
default=100000,
metadata={"help": "The size of the subset of the training data to use"},
)
eval_subset: Optional[int] = field(
default=50000,
metadata={"help": "The size of the subset of the eval data to use"},
)
gradient_checkpointing: Optional[bool] = field(
default=False,
metadata={"help": "Enables gradient checkpointing."},
)
optim: Optional[str] = field(
default="adamw_hf",
metadata={"help": "The optimizer to use."},
)
lr_scheduler_type: Optional[str] = field(
default="linear",
metadata={"help": "The lr scheduler"},
)
max_length: Optional[int] = field(default=512)
eval_first_step: Optional[bool] = field(
default=False,
metadata={"help": "Whether to run eval after the first step"},
)
parser = HfArgumentParser(ScriptArguments)
script_args = parser.parse_args_into_dataclasses()[0]
# Load the human stack-exchange-paired dataset for tuning the reward model.
train_dataset = load_dataset("lvwerra/stack-exchange-paired", data_dir="data/reward", split="train")
if script_args.train_subset > 0:
train_dataset = train_dataset.select(range(script_args.train_subset))
eval_dataset = load_dataset("lvwerra/stack-exchange-paired", data_dir="data/evaluation", split="train")
if script_args.eval_subset > 0:
eval_dataset = eval_dataset.select(range(script_args.eval_subset))
# Define the training args. Needs to be done before the model is loaded if you are using deepspeed.
model_name_split = script_args.model_name.split("/")[-1]
output_name = (
f"{model_name_split}_peft_stack-exchange-paired_rmts__{script_args.train_subset}_{script_args.learning_rate}"
)
training_args = TrainingArguments(
output_dir=output_name,
learning_rate=script_args.learning_rate,
per_device_train_batch_size=script_args.per_device_train_batch_size,
per_device_eval_batch_size=script_args.per_device_eval_batch_size,
num_train_epochs=script_args.num_train_epochs,
weight_decay=script_args.weight_decay,
evaluation_strategy="steps",
eval_steps=500,
save_strategy="steps",
save_steps=500,
gradient_accumulation_steps=script_args.gradient_accumulation_steps,
gradient_checkpointing=script_args.gradient_checkpointing,
deepspeed=script_args.deepspeed,
local_rank=script_args.local_rank,
remove_unused_columns=False,
label_names=[],
bf16=script_args.bf16,
logging_strategy="steps",
logging_steps=10,
optim=script_args.optim,
lr_scheduler_type=script_args.lr_scheduler_type,
)
# Load the value-head model and tokenizer.
tokenizer_name = script_args.tokenizer_name if script_args.tokenizer_name is not None else script_args.model_name
tokenizer = AutoTokenizer.from_pretrained(tokenizer_name, use_auth_token=True)
tokenizer.pad_token = tokenizer.eos_token
peft_config = LoraConfig(
task_type=TaskType.SEQ_CLS,
inference_mode=False,
r=8,
lora_alpha=32,
lora_dropout=0.1,
)
model = AutoModelForSequenceClassification.from_pretrained(
script_args.model_name, num_labels=1, torch_dtype=torch.bfloat16
)
model = get_peft_model(model, peft_config)
model.print_trainable_parameters()
# Need to do this for gpt2, because it doesn't have an official pad token.
tokenizer.pad_token = tokenizer.eos_token
model.config.pad_token_id = tokenizer.eos_token_id
model.config.use_cache = not script_args.gradient_checkpointing
num_proc = 24 # Can adjust to be higher if you have more processors.
original_columns = train_dataset.column_names
# Turn the dataset into pairs of post + summaries, where text_j is the preferred question + answer and text_k is the other.
# Then tokenize the dataset.
def preprocess_function(examples):
new_examples = {
"input_ids_j": [],
"attention_mask_j": [],
"input_ids_k": [],
"attention_mask_k": [],
}
for question, response_j, response_k in zip(examples["question"], examples["response_j"], examples["response_k"]):
tokenized_j = tokenizer("Question: " + question + "\n\nAnswer: " + response_j, truncation=True)
tokenized_k = tokenizer("Question: " + question + "\n\nAnswer: " + response_k, truncation=True)
new_examples["input_ids_j"].append(tokenized_j["input_ids"])
new_examples["attention_mask_j"].append(tokenized_j["attention_mask"])
new_examples["input_ids_k"].append(tokenized_k["input_ids"])
new_examples["attention_mask_k"].append(tokenized_k["attention_mask"])
return new_examples
# preprocess the dataset and filter out QAs that are longer than script_args.max_length
train_dataset = train_dataset.map(
preprocess_function,
batched=True,
num_proc=num_proc,
remove_columns=original_columns,
)
train_dataset = train_dataset.filter(
lambda x: len(x["input_ids_j"]) <= script_args.max_length and len(x["input_ids_k"]) <= script_args.max_length
)
eval_dataset = eval_dataset.map(
preprocess_function,
batched=True,
num_proc=num_proc,
remove_columns=original_columns,
)
eval_dataset = eval_dataset.filter(
lambda x: len(x["input_ids_j"]) <= script_args.max_length and len(x["input_ids_k"]) <= script_args.max_length
)
# We need to define a special data collator that batches the data in our j vs k format.
@dataclass
class RewardDataCollatorWithPadding:
tokenizer: PreTrainedTokenizerBase
padding: Union[bool, str, PaddingStrategy] = True
max_length: Optional[int] = None
pad_to_multiple_of: Optional[int] = None
return_tensors: str = "pt"
def __call__(self, features: List[Dict[str, Any]]) -> Dict[str, Any]:
features_j = []
features_k = []
for feature in features:
features_j.append(
{
"input_ids": feature["input_ids_j"],
"attention_mask": feature["attention_mask_j"],
}
)
features_k.append(
{
"input_ids": feature["input_ids_k"],
"attention_mask": feature["attention_mask_k"],
}
)
batch_j = self.tokenizer.pad(
features_j,
padding=self.padding,
max_length=self.max_length,
pad_to_multiple_of=self.pad_to_multiple_of,
return_tensors=self.return_tensors,
)
batch_k = self.tokenizer.pad(
features_k,
padding=self.padding,
max_length=self.max_length,
pad_to_multiple_of=self.pad_to_multiple_of,
return_tensors=self.return_tensors,
)
batch = {
"input_ids_j": batch_j["input_ids"],
"attention_mask_j": batch_j["attention_mask"],
"input_ids_k": batch_k["input_ids"],
"attention_mask_k": batch_k["attention_mask"],
"return_loss": True,
}
return batch
# Define the metric that we'll use for validation.
accuracy = evaluate.load("accuracy")
def compute_metrics(eval_pred):
predictions, _ = eval_pred
# Here, predictions is rewards_j and rewards_k.
# We want to see how much of the time rewards_j > rewards_k.
predictions = np.argmax(predictions, axis=0)
labels = np.zeros(predictions.shape)
return accuracy.compute(predictions=predictions, references=labels)
class RewardTrainer(Trainer):
# Define how to compute the reward loss. We use the InstructGPT pairwise logloss: https://arxiv.org/abs/2203.02155
def compute_loss(self, model, inputs, return_outputs=False):
rewards_j = model(input_ids=inputs["input_ids_j"], attention_mask=inputs["attention_mask_j"])[0]
rewards_k = model(input_ids=inputs["input_ids_k"], attention_mask=inputs["attention_mask_k"])[0]
loss = -nn.functional.logsigmoid(rewards_j - rewards_k).mean()
if return_outputs:
return loss, {"rewards_j": rewards_j, "rewards_k": rewards_k}
return loss
# Train the model, woohoo.
trainer = RewardTrainer(
model=model,
args=training_args,
train_dataset=train_dataset,
eval_dataset=eval_dataset,
compute_metrics=compute_metrics,
data_collator=RewardDataCollatorWithPadding(tokenizer=tokenizer, max_length=script_args.max_length),
)
if script_args.eval_first_step:
class EvaluateFirstStepCallback(TrainerCallback):
def on_step_end(self, args, state, control, **kwargs):
if state.global_step == 1:
control.should_evaluate = True
trainer.add_callback(EvaluateFirstStepCallback())
trainer.train(script_args.resume_from_checkpoint)
print("Saving last checkpoint of the model")
model.save_pretrained(output_name + "_peft_last_checkpoint")
| 0
|
hf_public_repos/trl/examples/research_projects/stack_llama_2
|
hf_public_repos/trl/examples/research_projects/stack_llama_2/scripts/README.md
|
# DPO pipeline for the creation of StackLlaMa 2: a Stack exchange llama-v2-7b model
## Prerequisites
Install all the dependencies in the `requirements.txt`:
```
$ pip install -U -r requirements.txt
```
Since we will use `accelerate` for training, make sure to run:
```
$ accelerate config
```
## Training
There were two main steps to the DPO training process:
1. Supervised fine-tuning of the base llama-v2-7b model to create llama-v2-7b-se:
- `accelerate launch examples/research_projects/stack_llama_2/scripts/sft_llama2.py --training_args.output_dir="sft"`
1. Run the DPO trainer using the model saved by the previous step:
- `accelerate launch examples/research_projects/stack_llama_2/scripts/dpo_llama2.py --model_name_or_path="sft/final_checkpoint" --output_dir="dpo"`
## Merging the adaptors
To merge the adaptors into the base model we can use the `merge_peft_adapter.py` helper script that comes with TRL:
```
python examples/research_projects/stack_llama/scripts/merge_peft_adapter.py --base_model_name="meta-llama/Llama-2-7b-hf" --adapter_model_name="dpo/final_checkpoint/" --output_name="stack-llama-2"
```
which will also push the model to your HuggingFace hub account.
## Running the model
We can load the DPO-trained LoRA adaptors which were saved by the DPO training step and load them via:
```py
from peft import AutoPeftModelForCausalLM
model = AutoPeftModelForCausalLM.from_pretrained(
"dpo/final_checkpoint",
low_cpu_mem_usage=True,
torch_dtype=torch.float16,
load_in_4bit=True,
)
model.generate(...)
```
| 0
|
hf_public_repos/trl/examples/research_projects/stack_llama_2
|
hf_public_repos/trl/examples/research_projects/stack_llama_2/scripts/dpo_llama2.py
|
# 0. imports
import os
from dataclasses import dataclass, field
from typing import Dict, Optional
import torch
from datasets import Dataset, load_dataset
from peft import LoraConfig
from transformers import AutoModelForCausalLM, AutoTokenizer, HfArgumentParser, TrainingArguments
from trl import DPOTrainer
# Define and parse arguments.
@dataclass
class ScriptArguments:
"""
The arguments for the DPO training script.
"""
# data parameters
beta: Optional[float] = field(default=0.1, metadata={"help": "the beta parameter for DPO loss"})
# training parameters
model_name_or_path: Optional[str] = field(
default="../sft/results/final_checkpoint",
metadata={"help": "the location of the SFT model name or path"},
)
learning_rate: Optional[float] = field(default=5e-4, metadata={"help": "optimizer learning rate"})
lr_scheduler_type: Optional[str] = field(default="cosine", metadata={"help": "the lr scheduler type"})
warmup_steps: Optional[int] = field(default=100, metadata={"help": "the number of warmup steps"})
weight_decay: Optional[float] = field(default=0.05, metadata={"help": "the weight decay"})
optimizer_type: Optional[str] = field(default="paged_adamw_32bit", metadata={"help": "the optimizer type"})
per_device_train_batch_size: Optional[int] = field(default=4, metadata={"help": "train batch size per device"})
per_device_eval_batch_size: Optional[int] = field(default=1, metadata={"help": "eval batch size per device"})
gradient_accumulation_steps: Optional[int] = field(
default=4, metadata={"help": "the number of gradient accumulation steps"}
)
gradient_checkpointing: Optional[bool] = field(
default=True, metadata={"help": "whether to use gradient checkpointing"}
)
lora_alpha: Optional[float] = field(default=16, metadata={"help": "the lora alpha parameter"})
lora_dropout: Optional[float] = field(default=0.05, metadata={"help": "the lora dropout parameter"})
lora_r: Optional[int] = field(default=8, metadata={"help": "the lora r parameter"})
max_prompt_length: Optional[int] = field(default=512, metadata={"help": "the maximum prompt length"})
max_length: Optional[int] = field(default=1024, metadata={"help": "the maximum sequence length"})
max_steps: Optional[int] = field(default=1000, metadata={"help": "max number of training steps"})
logging_steps: Optional[int] = field(default=10, metadata={"help": "the logging frequency"})
save_steps: Optional[int] = field(default=100, metadata={"help": "the saving frequency"})
eval_steps: Optional[int] = field(default=100, metadata={"help": "the evaluation frequency"})
output_dir: Optional[str] = field(default="./results", metadata={"help": "the output directory"})
log_freq: Optional[int] = field(default=1, metadata={"help": "the logging frequency"})
# instrumentation
sanity_check: Optional[bool] = field(default=False, metadata={"help": "only train on 1000 samples"})
report_to: Optional[str] = field(
default="wandb",
metadata={
"help": 'The list of integrations to report the results and logs to. Supported platforms are `"azure_ml"`,'
'`"comet_ml"`, `"mlflow"`, `"neptune"`, `"tensorboard"`,`"clearml"` and `"wandb"`. '
'Use `"all"` to report to all integrations installed, `"none"` for no integrations.'
},
)
# debug argument for distributed training
ignore_bias_buffers: Optional[bool] = field(
default=False,
metadata={
"help": "fix for DDP issues with LM bias/mask buffers - invalid scalar type,`inplace operation. See"
"https://github.com/huggingface/transformers/issues/22482#issuecomment-1595790992"
},
)
def get_stack_exchange_paired(
data_dir: str = "data/rl",
sanity_check: bool = False,
cache_dir: str = None,
num_proc=24,
) -> Dataset:
"""Load the stack-exchange-paired dataset from Hugging Face and convert it to the necessary format.
The dataset is converted to a dictionary with the following structure:
{
'prompt': List[str],
'chosen': List[str],
'rejected': List[str],
}
Prompts are structured as follows:
"Question: " + <prompt> + "\n\nAnswer: "
"""
dataset = load_dataset(
"lvwerra/stack-exchange-paired",
split="train",
cache_dir=cache_dir,
data_dir=data_dir,
)
original_columns = dataset.column_names
if sanity_check:
dataset = dataset.select(range(min(len(dataset), 1000)))
def return_prompt_and_responses(samples) -> Dict[str, str]:
return {
"prompt": ["Question: " + question + "\n\nAnswer: " for question in samples["question"]],
"chosen": samples["response_j"],
"rejected": samples["response_k"],
}
return dataset.map(
return_prompt_and_responses,
batched=True,
num_proc=num_proc,
remove_columns=original_columns,
)
if __name__ == "__main__":
parser = HfArgumentParser(ScriptArguments)
script_args = parser.parse_args_into_dataclasses()[0]
# 1. load a pretrained model
model = AutoModelForCausalLM.from_pretrained(
script_args.model_name_or_path,
low_cpu_mem_usage=True,
torch_dtype=torch.float16,
load_in_4bit=True,
)
model.config.use_cache = False
if script_args.ignore_bias_buffers:
# torch distributed hack
model._ddp_params_and_buffers_to_ignore = [
name for name, buffer in model.named_buffers() if buffer.dtype == torch.bool
]
model_ref = AutoModelForCausalLM.from_pretrained(
script_args.model_name_or_path,
low_cpu_mem_usage=True,
torch_dtype=torch.float16,
load_in_4bit=True,
)
tokenizer = AutoTokenizer.from_pretrained("meta-llama/Llama-2-7b-hf")
tokenizer.pad_token = tokenizer.eos_token
# 2. Load the Stack-exchange paired dataset
train_dataset = get_stack_exchange_paired(data_dir="data/rl", sanity_check=script_args.sanity_check)
train_dataset = train_dataset.filter(
lambda x: len(x["prompt"]) + len(x["chosen"]) <= script_args.max_length
and len(x["prompt"]) + len(x["rejected"]) <= script_args.max_length
)
# 3. Load evaluation dataset
eval_dataset = get_stack_exchange_paired(data_dir="data/evaluation", sanity_check=True)
eval_dataset = eval_dataset.filter(
lambda x: len(x["prompt"]) + len(x["chosen"]) <= script_args.max_length
and len(x["prompt"]) + len(x["rejected"]) <= script_args.max_length
)
# 4. initialize training arguments:
training_args = TrainingArguments(
per_device_train_batch_size=script_args.per_device_train_batch_size,
per_device_eval_batch_size=script_args.per_device_eval_batch_size,
max_steps=script_args.max_steps,
logging_steps=script_args.logging_steps,
save_steps=script_args.save_steps,
gradient_accumulation_steps=script_args.gradient_accumulation_steps,
gradient_checkpointing=script_args.gradient_checkpointing,
learning_rate=script_args.learning_rate,
evaluation_strategy="steps",
eval_steps=script_args.eval_steps,
output_dir=script_args.output_dir,
report_to=script_args.report_to,
lr_scheduler_type=script_args.lr_scheduler_type,
warmup_steps=script_args.warmup_steps,
optim=script_args.optimizer_type,
bf16=True,
remove_unused_columns=False,
run_name="dpo_llama2",
)
peft_config = LoraConfig(
r=script_args.lora_r,
lora_alpha=script_args.lora_alpha,
lora_dropout=script_args.lora_dropout,
target_modules=[
"q_proj",
"v_proj",
"k_proj",
"out_proj",
"fc_in",
"fc_out",
"wte",
],
bias="none",
task_type="CAUSAL_LM",
)
# 5. initialize the DPO trainer
dpo_trainer = DPOTrainer(
model,
model_ref,
args=training_args,
beta=script_args.beta,
train_dataset=train_dataset,
eval_dataset=eval_dataset,
tokenizer=tokenizer,
peft_config=peft_config,
max_prompt_length=script_args.max_prompt_length,
max_length=script_args.max_length,
)
# 6. train
dpo_trainer.train()
dpo_trainer.save_model(script_args.output_dir)
# 7. save
output_dir = os.path.join(script_args.output_dir, "final_checkpoint")
dpo_trainer.model.save_pretrained(output_dir)
| 0
|
hf_public_repos/trl/examples/research_projects/stack_llama_2
|
hf_public_repos/trl/examples/research_projects/stack_llama_2/scripts/sft_llama2.py
|
# Fine-Tune Llama2-7b on SE paired dataset
import os
from dataclasses import dataclass, field
from typing import Optional
import torch
import tyro
from accelerate import Accelerator
from datasets import load_dataset
from peft import AutoPeftModelForCausalLM, LoraConfig
from tqdm import tqdm
from transformers import AutoModelForCausalLM, AutoTokenizer, BitsAndBytesConfig, TrainingArguments
from trl import SFTTrainer
from trl.import_utils import is_xpu_available
from trl.trainer import ConstantLengthDataset
@dataclass
class ScriptArguments:
model_name: Optional[str] = field(default="meta-llama/Llama-2-7b-hf", metadata={"help": "the model name"})
dataset_name: Optional[str] = field(default="lvwerra/stack-exchange-paired", metadata={"help": "the dataset name"})
subset: Optional[str] = field(default="data/finetune", metadata={"help": "the subset to use"})
split: Optional[str] = field(default="train", metadata={"help": "the split to use"})
size_valid_set: Optional[int] = field(default=4000, metadata={"help": "the size of the validation set"})
streaming: Optional[bool] = field(default=True, metadata={"help": "whether to stream the dataset"})
shuffle_buffer: Optional[int] = field(default=5000, metadata={"help": "the shuffle buffer size"})
seq_length: Optional[int] = field(default=1024, metadata={"help": "the sequence length"})
num_workers: Optional[int] = field(default=4, metadata={"help": "the number of workers"})
training_args: TrainingArguments = field(
default_factory=lambda: TrainingArguments(
output_dir="./results",
max_steps=500,
logging_steps=10,
save_steps=10,
per_device_train_batch_size=4,
per_device_eval_batch_size=1,
gradient_accumulation_steps=2,
gradient_checkpointing=False,
group_by_length=False,
learning_rate=1e-4,
lr_scheduler_type="cosine",
warmup_steps=100,
weight_decay=0.05,
optim="paged_adamw_32bit",
bf16=True,
remove_unused_columns=False,
run_name="sft_llama2",
report_to="wandb",
)
)
packing: Optional[bool] = field(default=True, metadata={"help": "whether to use packing for SFTTrainer"})
peft_config: LoraConfig = field(
default_factory=lambda: LoraConfig(
r=8,
lora_alpha=16,
lora_dropout=0.05,
target_modules=["q_proj", "v_proj"],
bias="none",
task_type="CAUSAL_LM",
)
)
script_args = tyro.cli(ScriptArguments)
if script_args.training_args.group_by_length and script_args.packing:
raise ValueError("Cannot use both packing and group by length")
# `gradient_checkpointing` was True by default until `1f3314`, but it's actually not used.
# `gradient_checkpointing=True` will cause `Variable._execution_engine.run_backward`.
if script_args.training_args.gradient_checkpointing:
raise ValueError("gradient_checkpointing not supported")
def chars_token_ratio(dataset, tokenizer, nb_examples=400):
"""
Estimate the average number of characters per token in the dataset.
"""
total_characters, total_tokens = 0, 0
for _, example in tqdm(zip(range(nb_examples), iter(dataset)), total=nb_examples):
text = prepare_sample_text(example)
total_characters += len(text)
if tokenizer.is_fast:
total_tokens += len(tokenizer(text).tokens())
else:
total_tokens += len(tokenizer.tokenize(text))
return total_characters / total_tokens
def print_trainable_parameters(model):
"""
Prints the number of trainable parameters in the model.
"""
trainable_params = 0
all_param = 0
for _, param in model.named_parameters():
all_param += param.numel()
if param.requires_grad:
trainable_params += param.numel()
print(
f"trainable params: {trainable_params} || all params: {all_param} || trainable%: {100 * trainable_params / all_param}"
)
def prepare_sample_text(example):
"""Prepare the text from a sample of the dataset."""
text = f"Question: {example['question']}\n\nAnswer: {example['response_j']}"
return text
def create_datasets(tokenizer, args):
dataset = load_dataset(
args.dataset_name,
data_dir=args.subset,
split=args.split,
use_auth_token=True,
num_proc=args.num_workers if not args.streaming else None,
streaming=args.streaming,
)
if args.streaming:
print("Loading the dataset in streaming mode")
valid_data = dataset.take(args.size_valid_set)
train_data = dataset.skip(args.size_valid_set)
train_data = train_data.shuffle(buffer_size=args.shuffle_buffer, seed=None)
else:
dataset = dataset.train_test_split(test_size=0.005, seed=None)
train_data = dataset["train"]
valid_data = dataset["test"]
print(f"Size of the train set: {len(train_data)}. Size of the validation set: {len(valid_data)}")
chars_per_token = chars_token_ratio(train_data, tokenizer)
print(f"The character to token ratio of the dataset is: {chars_per_token:.2f}")
train_dataset = ConstantLengthDataset(
tokenizer,
train_data,
formatting_func=prepare_sample_text,
infinite=True,
seq_length=args.seq_length,
chars_per_token=chars_per_token,
)
valid_dataset = ConstantLengthDataset(
tokenizer,
valid_data,
formatting_func=prepare_sample_text,
infinite=False,
seq_length=args.seq_length,
chars_per_token=chars_per_token,
)
return train_dataset, valid_dataset
bnb_config = BitsAndBytesConfig(
load_in_4bit=True,
bnb_4bit_quant_type="nf4",
bnb_4bit_compute_dtype=torch.bfloat16,
)
base_model = AutoModelForCausalLM.from_pretrained(
script_args.model_name,
quantization_config=bnb_config,
device_map={"": Accelerator().local_process_index},
trust_remote_code=True,
use_auth_token=True,
)
base_model.config.use_cache = False
peft_config = script_args.peft_config
tokenizer = AutoTokenizer.from_pretrained(script_args.model_name, trust_remote_code=True)
tokenizer.pad_token = tokenizer.eos_token
tokenizer.padding_side = "right" # Fix weird overflow issue with fp16 training
training_args = script_args.training_args
train_dataset, eval_dataset = create_datasets(tokenizer, script_args)
trainer = SFTTrainer(
model=base_model,
train_dataset=train_dataset,
eval_dataset=eval_dataset,
peft_config=peft_config,
packing=script_args.packing,
max_seq_length=None,
tokenizer=tokenizer,
args=training_args,
)
trainer.train()
trainer.save_model(script_args.training_args.output_dir)
output_dir = os.path.join(script_args.training_args.output_dir, "final_checkpoint")
trainer.model.save_pretrained(output_dir)
# Free memory for merging weights
del base_model
if is_xpu_available():
torch.xpu.empty_cache()
else:
torch.cuda.empty_cache()
model = AutoPeftModelForCausalLM.from_pretrained(output_dir, device_map="auto", torch_dtype=torch.bfloat16)
model = model.merge_and_unload()
output_merged_dir = os.path.join(script_args.training_args.output_dir, "final_merged_checkpoint")
model.save_pretrained(output_merged_dir, safe_serialization=True)
| 0
|
hf_public_repos/trl/examples/research_projects/stack_llama_2
|
hf_public_repos/trl/examples/research_projects/stack_llama_2/scripts/requirements.txt
|
transformers
trl
peft
accelerate
datasets
bitsandbytes
wandb
| 0
|
hf_public_repos/trl/examples/research_projects
|
hf_public_repos/trl/examples/research_projects/tools/triviaqa.py
|
# coding=utf-8
# Copyright 2023 The HuggingFace Inc. team. All rights reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
import os
from dataclasses import dataclass, field
from typing import Optional
import torch
from datasets import load_dataset
from peft import LoraConfig
from transformers import AutoTokenizer, HfArgumentParser, load_tool
from trl import AutoModelForCausalLMWithValueHead, PPOConfig, PPOTrainer, TextEnvironment
os.environ["HF_ALLOW_CODE_EVAL"] = "1"
os.environ["TOKENIZERS_PARALLELISM"] = "false"
@dataclass
class ScriptArguments:
model_name: Optional[str] = field(default="bigcode/starcoderbase", metadata={"help": "the model name"})
log_with: Optional[str] = field(default=None, metadata={"help": "use 'wandb' to log with wandb"})
learning_rate: Optional[float] = field(default=1e-5, metadata={"help": "the learning rate"})
mini_batch_size: Optional[int] = field(default=1, metadata={"help": "the PPO minibatch size"})
batch_size: Optional[int] = field(default=32, metadata={"help": "the batch size"})
gradient_accumulation_steps: Optional[int] = field(
default=16, metadata={"help": "the number of gradient accumulation steps"}
)
max_new_tokens: Optional[int] = field(default=256, metadata={"help": "max number of generated tokens per turn"})
ppo_epochs: Optional[int] = field(default=1, metadata={"help": "max number of ppo epochs"})
iterations: Optional[int] = field(default=1000, metadata={"help": "the number of iterations"})
seed: Optional[int] = field(default=0, metadata={"help": "the random seed"})
parser = HfArgumentParser(ScriptArguments)
args = parser.parse_args_into_dataclasses()[0]
lora_config = LoraConfig(
r=16,
lora_alpha=32,
lora_dropout=0.05,
bias="none",
task_type="CAUSAL_LM",
target_modules=["c_proj", "c_attn", "q_attn"],
)
# set up models
model = AutoModelForCausalLMWithValueHead.from_pretrained(
args.model_name,
use_auth_token=True,
trust_remote_code=True,
load_in_4bit=True,
peft_config=lora_config,
)
tokenizer = AutoTokenizer.from_pretrained(args.model_name, use_auth_token=True)
tokenizer.pad_token = tokenizer.eos_token
# system prompt
prompt = """\
Answer the following question:
Q: In which branch of the arts is Patricia Neary famous?
A: Ballets
A2: <request><Wiki>Patricia Neary<call>Patricia Neary (born October 27, 1942) is an American ballerina, choreographer and ballet director, who has been particularly active in Switzerland. She has also been a highly successful ambassador for the Balanchine Trust, bringing George Balanchine's ballets to 60 cities around the globe.<response>
Result=Ballets<submit>
Q: Who won Super Bowl XX?
A: Chicago Bears
A2: <request><Wiki>Super Bowl XX<call>Super Bowl XX was an American football game between the National Football Conference (NFC) champion Chicago Bears and the American Football Conference (AFC) champion New England Patriots to decide the National Football League (NFL) champion for the 1985 season. The Bears defeated the Patriots by the score of 46–10, capturing their first NFL championship (and Chicago's first overall sports victory) since 1963, three years prior to the birth of the Super Bowl. Super Bowl XX was played on January 26, 1986 at the Louisiana Superdome in New Orleans.<response>
Result=Chicago Bears<submit>
Q: """
generation_kwargs = {
"min_length": -1,
"top_k": 0.0,
"top_p": 1.0,
"do_sample": True,
"pad_token_id": tokenizer.eos_token_id,
"eos_token_id": -1,
"max_new_tokens": args.max_new_tokens,
}
# trainer
config = PPOConfig(
batch_size=args.batch_size,
model_name=args.model_name,
learning_rate=args.learning_rate,
log_with=args.log_with,
mini_batch_size=args.mini_batch_size,
ppo_epochs=args.ppo_epochs,
gradient_accumulation_steps=args.gradient_accumulation_steps,
seed=args.seed,
optimize_cuda_cache=True,
)
ppo_trainer = PPOTrainer(config=config, model=model, tokenizer=tokenizer)
dataset = load_dataset("trivia_qa", "rc", split="train")
local_seed = args.seed + ppo_trainer.accelerator.process_index * 100003 # Prime
dataset = dataset.shuffle(local_seed)
def data_generator():
for i in range(len(dataset)):
yield dataset[i]["question"], [item for item in dataset[i]["answer"]["normalized_aliases"]]
gen = data_generator()
gen = iter(gen)
def generate_data(n):
tasks, answers = [], []
for i in range(n):
q, a = next(gen)
tasks.append(q)
answers.append(a)
return tasks, answers
def exact_match_reward(responses, answers=None):
"""Reward if generated response contains correct answer."""
rewards = []
for response, answer in zip(responses, answers):
reward = 0.0
for a in answer:
if a.lower() in response.lower():
reward += 1.0
break
rewards.append(torch.tensor(reward))
return rewards
# text env
tool = load_tool("vwxyzjn/pyserini-wikipedia-kilt-doc")
# limit the amount if tokens
tool_fn = lambda x: tool(x).split("\n")[1][:600] # noqa
text_env = TextEnvironment(
model,
tokenizer,
{"Wiki": tool_fn},
exact_match_reward,
prompt,
generation_kwargs=generation_kwargs,
max_tool_reponse=400,
)
def print_trainable_parameters(model):
trainable_params = 0
all_param = 0
for _, param in model.named_parameters():
all_param += param.numel()
if param.requires_grad:
trainable_params += param.numel()
print(
f"trainable params: {trainable_params} || all params: {all_param} || trainable%: {100 * trainable_params / all_param}"
)
print_trainable_parameters(model)
# main training loop
for i in range(args.iterations):
tasks, answers = generate_data(config.batch_size)
queries, responses, masks, rewards, histories = text_env.run(tasks, answers=answers)
train_stats = ppo_trainer.step(queries, responses, rewards, masks)
response_texts = [tokenizer.decode(response) for response in responses]
query_texts = [tokenizer.decode(query) for query in queries]
texts = {
"query": [qt.split("<submit>")[-1].strip() for qt in query_texts],
"response": response_texts,
"answer": [", ".join(item) for item in answers],
}
all_rewards = ppo_trainer.accelerator.gather(torch.tensor(rewards, device=ppo_trainer.accelerator.device))
ppo_trainer.log_stats(
train_stats, texts, [item for item in all_rewards], columns_to_log=["query", "response", "answer"]
)
if i % 100 == 0:
ppo_trainer.save_pretrained(f"models/{args.model_name}_{args.seed}_{i}_triviaqa")
| 0
|
hf_public_repos/trl/examples/research_projects
|
hf_public_repos/trl/examples/research_projects/tools/python_interpreter.py
|
# coding=utf-8
# Copyright 2023 The HuggingFace Inc. team. All rights reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
import os
import re
from dataclasses import dataclass, field
from typing import Optional
import numpy as np
import torch
from datasets import load_dataset
from peft import LoraConfig
from transformers import AutoTokenizer, HfArgumentParser, load_tool
from trl import AutoModelForCausalLMWithValueHead, PPOConfig, PPOTrainer, TextEnvironment
os.environ["HF_ALLOW_CODE_EVAL"] = "1"
os.environ["TOKENIZERS_PARALLELISM"] = "false"
@dataclass
class ScriptArguments:
model_name: Optional[str] = field(default="bigcode/starcoderbase", metadata={"help": "the model name"})
learning_rate: Optional[float] = field(default=1e-5, metadata={"help": "the learning rate"})
mini_batch_size: Optional[int] = field(default=1, metadata={"help": "the PPO minibatch size"})
batch_size: Optional[int] = field(default=32, metadata={"help": "the batch size"})
gradient_accumulation_steps: Optional[int] = field(
default=16, metadata={"help": "the number of gradient accumulation steps"}
)
max_new_tokens: Optional[int] = field(default=256, metadata={"help": "max number of generated tokens per turn"})
ppo_epochs: Optional[int] = field(default=1, metadata={"help": "max number of ppo epochs"})
n_epochs: Optional[int] = field(default=32, metadata={"help": "max number of ppo epochs"})
parser = HfArgumentParser(ScriptArguments)
args = parser.parse_args_into_dataclasses()[0]
def exact_match_reward(responses, answers=None):
"""Reward if generated response contains correct answer."""
rewards = []
pattern = r"Result\s*=\s*(-?\d+(?:\.\d+)?)\s*<submit>" # generated by chatGPT
for response, answer in zip(responses, answers):
reward = 0.0
try:
predicted_number = None
match_pattern = re.findall(pattern, response)
if match_pattern:
predicted_number = float(match_pattern[0])
if predicted_number is not None:
if np.abs((predicted_number - float(answer))) < 0.1:
reward += 1.0
except: # noqa
pass
rewards.append(torch.tensor(reward))
return rewards
def evaluate(test_dataloader, text_env, ppo_trainer):
test_rewards = []
for test_batch in test_dataloader:
_, _, _, rewards, _ = text_env.run(test_batch["query"], answers=test_batch["answer"])
test_rewards.extend(rewards)
test_rewards = ppo_trainer.accelerator.gather_for_metrics(
torch.stack(test_rewards).to(ppo_trainer.accelerator.device)
)
return test_rewards.mean()
lora_config = LoraConfig(
r=16,
lora_alpha=32,
lora_dropout=0.05,
bias="none",
task_type="CAUSAL_LM",
target_modules=["c_proj", "c_attn", "q_attn"],
)
# set up models
model = AutoModelForCausalLMWithValueHead.from_pretrained(
args.model_name,
use_auth_token=True,
load_in_4bit=True,
peft_config=lora_config,
)
tokenizer = AutoTokenizer.from_pretrained(args.model_name, use_auth_token=True)
tokenizer.pad_token = tokenizer.eos_token
ds = load_dataset("gsm8k", "main", split="train")
ds = ds.rename_columns({"question": "query"})
ds = ds.map(lambda x: {"answer": x["answer"].split("#### ")[1]})
ds = ds.select(range(1, len(ds))) # skip the first sample which is used in prompt
ds_test = load_dataset("gsm8k", "main", split="test")
ds_test = ds_test.rename_columns({"question": "query"})
ds_test = ds_test.map(lambda x: {"answer": x["answer"].split("#### ")[1]})
test_dataloader = torch.utils.data.DataLoader(ds_test, batch_size=args.batch_size)
# prompt
prompt = """\
Example of using a Python API to solve math questions.
Q: Olivia has $23. She bought five bagels for $3 each. How much money does she have left?
<request><PythonInterpreter>
def solution():
money_initial = 23
bagels = 5
bagel_cost = 3
money_spent = bagels * bagel_cost
money_left = money_initial - money_spent
result = money_left
return result
print(solution())
<call>72<response>
Result = 72 <submit>
Q: """
generation_kwargs = {
"min_length": -1,
"top_k": 0.0,
"top_p": 1.0,
"do_sample": True,
"pad_token_id": tokenizer.eos_token_id,
"eos_token_id": -1,
"max_new_tokens": args.max_new_tokens,
}
# trainer
ppo_config = PPOConfig(
batch_size=args.batch_size,
learning_rate=args.learning_rate,
mini_batch_size=args.mini_batch_size,
ppo_epochs=args.ppo_epochs,
gradient_accumulation_steps=args.gradient_accumulation_steps,
log_with="wandb",
tracker_project_name="trl-gsm8k",
remove_unused_columns=False,
optimize_cuda_cache=True,
)
ppo_trainer = PPOTrainer(config=ppo_config, model=model, tokenizer=tokenizer, dataset=ds)
test_dataloader = ppo_trainer.accelerator.prepare(test_dataloader)
# text env
text_env = TextEnvironment(
model,
tokenizer,
[load_tool("lvwerra/python-interpreter")],
exact_match_reward,
prompt,
max_turns=2,
generation_kwargs=generation_kwargs,
)
# main training loop
for epoch in range(args.n_epochs):
for step, batch in enumerate(ppo_trainer.dataloader):
if (step == 0) and (epoch % 4 == 0): # evaluate every 4 epochs
reward_mean_test = evaluate(test_dataloader, text_env, ppo_trainer)
else:
reward_mean_test = None
queries, responses, masks, rewards, histories = text_env.run(batch["query"], answers=batch["answer"])
train_stats = ppo_trainer.step(queries, responses, rewards, masks)
# logging
if reward_mean_test is not None:
train_stats["env/reward_mean_test"] = reward_mean_test
texts = {
"query": batch["query"],
"response": [tokenizer.decode(response) for response in responses],
"answer": batch["answer"],
}
ppo_trainer.log_stats(train_stats, texts, rewards, columns_to_log=["query", "response", "answer"])
reward_mean_test = evaluate(test_dataloader, text_env, ppo_trainer)
ppo_trainer.save_pretrained(f"model/{args.model_name}-gsm8k")
| 0
|
hf_public_repos/trl/examples/research_projects
|
hf_public_repos/trl/examples/research_projects/tools/calculator.py
|
# coding=utf-8
# Copyright 2023 The HuggingFace Inc. team. All rights reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
import re
import numpy as np
import torch
from transformers import AutoTokenizer, load_tool
from trl import AutoModelForCausalLMWithValueHead, PPOConfig, PPOTrainer, TextEnvironment
def generate_data(n):
"""Generate random arithmetic tasks and answers."""
tasks, answers = [], []
for _ in range(n):
a = np.random.randint(0, 50)
b = np.random.randint(0, 50)
op = np.random.choice(["-", "+", "*"])
tasks.append(f"\n\nWhat is {a} {op} {b}?")
if op == "-":
answers.append(a - b)
elif op == "+":
answers.append(a + b)
else:
answers.append(a * b)
return tasks, answers
def exact_match_reward(responses, answers=None):
"""Reward if generated response contains correct answer."""
rewards = []
pattern = r"Result\s*=\s*(-?\d+(?:\.\d+)?)\s*<submit>" # generated by chatGPT
for response, answer in zip(responses, answers):
reward = 0.0
predicted_number = None
match_pattern = re.findall(pattern, response)
if match_pattern:
predicted_number = float(match_pattern[0])
if predicted_number is not None:
if np.abs(predicted_number - answer) < 0.01:
reward += 1.0
rewards.append(torch.tensor(reward))
return rewards
# set up models
model_id = "gpt2"
model = AutoModelForCausalLMWithValueHead.from_pretrained(model_id)
model_ref = AutoModelForCausalLMWithValueHead.from_pretrained(model_id)
tokenizer = AutoTokenizer.from_pretrained(model_id)
tokenizer.pad_token = tokenizer.eos_token
# system prompt
prompt = """\
What is 13-3?
<request><SimpleCalculatorTool>13-3<call>10.0<response>
Result=10<submit>
What is 4*3?
<request><SimpleCalculatorTool>4*3<call>12.0<response>
Result=12<submit>"""
generation_kwargs = {
"min_length": -1,
"top_k": 0.0,
"top_p": 1.0,
"do_sample": True,
"pad_token_id": tokenizer.eos_token_id,
"eos_token_id": -1,
"max_new_tokens": 32,
}
# trainer
ppo_config = PPOConfig(
batch_size=256,
learning_rate=1.41e-5,
mini_batch_size=64,
log_with="wandb",
)
ppo_trainer = PPOTrainer(ppo_config, model, model_ref, tokenizer)
# text env
text_env = TextEnvironment(
model,
tokenizer,
{"SimpleCalculatorTool": load_tool("ybelkada/simple-calculator")},
exact_match_reward,
prompt,
generation_kwargs=generation_kwargs,
)
# main training loop
for step in range(100):
tasks, answers = generate_data(ppo_config.batch_size)
queries, responses, masks, rewards, histories = text_env.run(tasks, answers=answers)
train_stats = ppo_trainer.step(queries, responses, rewards, masks)
response_texts = [tokenizer.decode(response) for response in responses]
query_texts = [tokenizer.decode(query) for query in queries]
texts = {"query": [qt.split("<submit>")[-1].strip() for qt in query_texts], "response": response_texts}
ppo_trainer.log_stats(train_stats, texts, rewards, columns_to_log=["query", "response", "answer"])
ppo_trainer.save_pretrained(model_id + "-calculator")
| 0
|
hf_public_repos/trl/examples/research_projects
|
hf_public_repos/trl/examples/research_projects/toxicity/README.md
|
# De-detoxifying language models
To run this code, do the following:
```shell
ACCELERATE_LOG_LEVEL=info accelerate launch --config_file {CONFIG} examples/research_projects/toxicity/scripts/gpt-j-6b-toxicity.py --log_with wandb
```
| 0
|
hf_public_repos/trl/examples/research_projects/toxicity
|
hf_public_repos/trl/examples/research_projects/toxicity/scripts/evaluate-toxicity.py
|
import argparse
import csv
import evaluate
import numpy as np
import torch
from datasets import load_dataset
from tqdm import tqdm
from transformers import AutoModelForCausalLM, AutoTokenizer
from trl.import_utils import is_xpu_available
toxicity = evaluate.load("ybelkada/toxicity", "DaNLP/da-electra-hatespeech-detection", module_type="measurement")
ds = load_dataset("OxAISH-AL-LLM/wiki_toxic", split="test")
parser = argparse.ArgumentParser(description="Evaluate de-toxified models")
parser.add_argument("--model_type", default="all", type=str, help="Relative path to the source model folder")
parser.add_argument("--output_file", default="toxicity.csv", type=str, help="Relative path to the source model folder")
parser.add_argument("--batch_size", default=64, type=int, help="Batch size")
parser.add_argument("--num_samples", default=400, type=int, help="Number of samples")
parser.add_argument("--context_length", default=2000, type=int, help="Number of samples")
parser.add_argument("--max_new_tokens", default=30, type=int, help="Max new tokens for generation")
args = parser.parse_args()
if args.model_type == "all":
MODELS_TO_TEST = [
"ybelkada/gpt-neo-125m-detox",
"EleutherAI/gpt-neo-125M",
"EleutherAI/gpt-neo-2.7B",
"ybelkada/gpt-neo-2.7B-detox",
"ybelkada/gpt-j-6b-sharded-bf16",
"ybelkada/gpt-j-6b-detoxs",
]
elif args.model_type == "gpt-neo":
MODELS_TO_TEST = [
"ybelkada/gpt-neo-125m-detox",
"EleutherAI/gpt-neo-125M",
"EleutherAI/gpt-neo-2.7B",
"ybelkada/gpt-neo-2.7B-detox",
]
elif args.model_type == "gpt-j":
MODELS_TO_TEST = [
"ybelkada/gpt-j-6b-sharded-bf16",
"ybelkada/gpt-j-6b-detox",
]
else:
MODELS_TO_TEST = [args.model_type]
NUM_SAMPLES = args.num_samples
BATCH_SIZE = args.batch_size
output_file = args.output_file
max_new_tokens = args.max_new_tokens
context_length = args.context_length
if is_xpu_available():
device = torch.xpu.current_device()
else:
device = torch.cuda.current_device() if torch.cuda.is_available() else "cpu"
# consider only toxic prompts
ds = ds.filter(lambda x: x["label"] == 1)
toxicities = {}
# open a csv file
file = open(f"{output_file}", "w", newline="")
writer = csv.writer(file)
# add first rows
writer.writerow(["model_id", "mean_toxicity", "std_toxicity"])
for model_id in tqdm(MODELS_TO_TEST):
model = AutoModelForCausalLM.from_pretrained(model_id, device_map={"": device}, torch_dtype=torch.bfloat16)
tokenizer = AutoTokenizer.from_pretrained(model_id)
tokenizer.pad_token = tokenizer.eos_token
tokenizer.padding_side = "left"
input_texts = []
for i, example in enumerate(ds):
# set seed
torch.manual_seed(42)
input_text = example["comment_text"]
input_texts.append(input_text[:2000])
if i > NUM_SAMPLES:
break
if (i + 1) % BATCH_SIZE == 0:
inputs = tokenizer(input_texts, return_tensors="pt", padding=True).to(device)
inputs.input_ids = inputs.input_ids[:context_length]
inputs.attention_mask = inputs.attention_mask[:context_length]
outputs = model.generate(**inputs, do_sample=True, max_new_tokens=max_new_tokens, use_cache=True)
generated_texts = tokenizer.batch_decode(outputs, skip_special_tokens=True)
generated_texts = [
generated_text.replace(input_texts[i], "") for i, generated_text in enumerate(generated_texts)
]
toxicity_score = toxicity.compute(predictions=generated_texts)
input_texts = []
if model_id not in toxicities:
toxicities[model_id] = []
toxicities[model_id].extend(toxicity_score["toxicity"])
# last batch
inputs = tokenizer(input_texts, return_tensors="pt", padding=True).to(device)
outputs = model.generate(**inputs, do_sample=True, max_new_tokens=30)
generated_texts = tokenizer.batch_decode(outputs, skip_special_tokens=True)
generated_texts = [generated_text.replace(input_texts[i], "") for i, generated_text in enumerate(generated_texts)]
toxicity_score = toxicity.compute(predictions=generated_texts)
toxicities[model_id].extend(toxicity_score["toxicity"])
# compute mean & std using np
mean = np.mean(toxicities[model_id])
std = np.std(toxicities[model_id])
# save to file
writer.writerow([model_id, mean, std])
# print
print(f"Model: {model_id} - Mean: {mean} - Std: {std}")
model = None
if is_xpu_available():
torch.xpu.empty_cache()
else:
torch.cuda.empty_cache()
# close file
file.close()
| 0
|
hf_public_repos/trl/examples/research_projects/toxicity
|
hf_public_repos/trl/examples/research_projects/toxicity/scripts/gpt-j-6b-toxicity.py
|
# coding=utf-8
# Copyright 2023 The HuggingFace Inc. team. All rights reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
from dataclasses import dataclass, field
from typing import Optional
import torch
from datasets import load_dataset
from torch.optim import Adam
from tqdm import tqdm
from transformers import (
AutoModelForCausalLM,
AutoTokenizer,
HfArgumentParser,
RobertaForSequenceClassification,
RobertaTokenizer,
)
from trl import AutoModelForCausalLMWithValueHead, PPOConfig, PPOTrainer, create_reference_model, set_seed
from trl.core import LengthSampler
tqdm.pandas()
########################################################################
# This is a fully working simple example to use trl with accelerate.
#
# This example fine-tunes a GPTJ model to generate less toxic contents
# by using allenai/real-toxicity-prompts dataset. We use PPO
# (proximal policy optimization) to optimize the model.
# in any of the following settings (with the same script):
# - single CPU or single GPU
# - multi GPUS (using PyTorch distributed mode)
# - multi GPUS (using DeepSpeed ZeRO-Offload stages 1 & 2)
# - fp16 (mixed-precision) or fp32 (normal precision)
#
# To run it in each of these various modes, first initialize the accelerate
# configuration with `accelerate config`
#
########################################################################
# We first define the configuration of the experiment, defining the model, the dataset,
# the training parameters, and the PPO parameters.
# Check the default arguments in the `PPOConfig` class for more details.
# If you want to log with tensorboard, add the kwarg
# `project_kwargs={"logging_dir": PATH_TO_LOGS}` to the PPOConfig.
@dataclass
class ScriptArguments:
"""
The name of the Casual LM model we wish to fine with PPO
"""
# NOTE: gpt2 models use Conv1D instead of Linear layers which are not yet supported in 8 bit mode
# models like gpt-neo* models are more suitable.
model_name: Optional[str] = field(default="ybelkada/gpt-j-6b-sharded-bf16", metadata={"help": "the model name"})
log_with: Optional[str] = field(default=None, metadata={"help": "use 'wandb' to log with wandb"})
learning_rate: Optional[float] = field(default=(1.47e-5) * 2, metadata={"help": "the learning rate"})
mini_batch_size: Optional[int] = field(default=4, metadata={"help": "the PPO minibatch size"})
batch_size: Optional[int] = field(default=16, metadata={"help": "the batch size"})
gradient_accumulation_steps: Optional[int] = field(
default=1, metadata={"help": "the number of gradient accumulation steps"}
)
model_save_path: Optional[str] = field(
default="./gpt-j-6B-detoxified-long-context-26-shl-1e4-final",
metadata={"help": "the path to save the model"},
)
parser = HfArgumentParser(ScriptArguments)
script_args = parser.parse_args_into_dataclasses()[0]
config = PPOConfig(
model_name=script_args.model_name,
learning_rate=script_args.learning_rate,
log_with=script_args.log_with,
ppo_epochs=100,
mini_batch_size=script_args.mini_batch_size,
batch_size=script_args.batch_size,
gradient_accumulation_steps=script_args.gradient_accumulation_steps,
)
# Below is an example function to build the dataset. In our case, we use the IMDB dataset
# from the `datasets` library. One should customize this function to train the model on
# its own dataset.
def build_dataset(
config, dataset_name="allenai/real-toxicity-prompts", input_min_text_length=5, input_max_text_length=10
):
"""
Build dataset for training. This builds the dataset from `load_dataset`, one should
customize this function to train the model on its own dataset.
Args:
dataset_name (`str`):
The name of the dataset to be loaded.
Returns:
dataloader (`torch.utils.data.DataLoader`):
The dataloader for the dataset.
"""
tokenizer = AutoTokenizer.from_pretrained(config.model_name)
tokenizer.pad_token = tokenizer.eos_token
ds = load_dataset(dataset_name, split="train")
def filter_fn(sample):
toxicity = sample["prompt"]["toxicity"]
return toxicity is not None and toxicity > 0.3
ds = ds.filter(filter_fn, batched=False)
input_size = LengthSampler(input_min_text_length, input_max_text_length)
def tokenize(sample):
prompt = sample["prompt"]["text"]
continuation = sample["continuation"]["text"]
sample["input_ids"] = tokenizer.encode(prompt + continuation)[: input_size()]
sample["query"] = tokenizer.decode(sample["input_ids"])
return sample
ds = ds.map(tokenize, batched=False)
ds.set_format(type="torch")
ds = ds.train_test_split(test_size=0.2, shuffle=False)["train"]
return ds
# We retrieve the dataloader by calling the `build_dataset` function.
min_input_length = 30
max_input_length = 40
dataset = build_dataset(config, input_min_text_length=min_input_length, input_max_text_length=max_input_length)
def collator(data):
return dict((key, [d[key] for d in data]) for key in data[0])
# set seed before initializing value head for deterministic eval
set_seed(config.seed)
# Now let's build the model, the reference model, and the tokenizer. We first load the model
# in bfloat16 to save memory using `transformers`.
model = AutoModelForCausalLM.from_pretrained(config.model_name, torch_dtype=torch.bfloat16)
# And then we pass the loaded model to `AutoModelForCausalLMWithValueHead`.
model = AutoModelForCausalLMWithValueHead.from_pretrained(model)
# We create a reference model by sharing 20 layers
ref_model = create_reference_model(model, num_shared_layers=20)
# We make sure to use `Adam` optimizer on the model parameters that require gradients.
optimizer = Adam(filter(lambda p: p.requires_grad, model.parameters()), lr=config.learning_rate)
# GPT-2 / GPT-J tokenizer has a pad token, but it is not eos_token by default. We need to set it to eos_token.
# only for this model.
tokenizer = AutoTokenizer.from_pretrained(config.model_name)
tokenizer.pad_token = tokenizer.eos_token
# We then build the PPOTrainer, passing the model, the reference model, the tokenizer
ppo_trainer = PPOTrainer(
config,
model,
ref_model=ref_model,
tokenizer=tokenizer,
dataset=dataset,
data_collator=collator,
optimizer=optimizer,
)
# We then build the reward pipeline, we will use the toxicity model to compute the reward.
# We first load the toxicity model and tokenizer.
toxicity_model_id = "facebook/roberta-hate-speech-dynabench-r4-target"
toxicity_tokenizer = RobertaTokenizer.from_pretrained(toxicity_model_id)
# We load the toxicity model in fp16 to save memory.
toxicity_model = RobertaForSequenceClassification.from_pretrained(toxicity_model_id, torch_dtype=torch.float16).to(
ppo_trainer.accelerator.device
)
# We then define the arguments to pass to the `generate` function. These arguments
# are passed to the `generate` function of the PPOTrainer, which is a wrapper around
# the `generate` function of the trained model.
generation_kwargs = {
"min_length": -1,
"top_k": 0.0,
"top_p": 1.0,
"do_sample": True,
"pad_token_id": tokenizer.eos_token_id,
}
output_min_length = 20
output_max_length = 30
output_length_sampler = LengthSampler(output_min_length, output_max_length)
model_save_path = script_args.model_save_path
for epoch, batch in tqdm(enumerate(ppo_trainer.dataloader)):
query_tensors = batch["input_ids"]
# Get response from the policy model
response_tensors = []
for query in query_tensors:
gen_len = output_length_sampler()
generation_kwargs["max_new_tokens"] = gen_len
response = ppo_trainer.generate(query, **generation_kwargs)
response_tensors.append(response.squeeze()[-gen_len:])
batch["response"] = [tokenizer.decode(r.squeeze()) for r in response_tensors]
# Compute sentiment score # noqa
texts = batch["response"]
toxicity_inputs = toxicity_tokenizer(texts, padding=True, truncation=True, return_tensors="pt").to(
ppo_trainer.accelerator.device
)
logits = toxicity_model(**toxicity_inputs).logits.float()
toxicity_labels = (logits[:, 0]).tolist()
rewards = [torch.tensor(output) for output in toxicity_labels]
# Run PPO step
stats = ppo_trainer.step(query_tensors, response_tensors, rewards)
ppo_trainer.log_stats(stats, batch, rewards)
# Save model every 100 epochs
if epoch % 100 == 0:
if ppo_trainer.accelerator.is_main_process:
ppo_trainer.save_pretrained(model_save_path)
| 0
|
hf_public_repos/trl/examples
|
hf_public_repos/trl/examples/accelerate_configs/deepspeed_zero1.yaml
|
compute_environment: LOCAL_MACHINE
debug: false
deepspeed_config:
deepspeed_multinode_launcher: standard
gradient_accumulation_steps: 1
zero3_init_flag: false
zero_stage: 1
distributed_type: DEEPSPEED
downcast_bf16: 'no'
machine_rank: 0
main_training_function: main
mixed_precision: 'bf16'
num_machines: 1
num_processes: 8
rdzv_backend: static
same_network: true
tpu_env: []
tpu_use_cluster: false
tpu_use_sudo: false
use_cpu: false
| 0
|
hf_public_repos/trl/examples
|
hf_public_repos/trl/examples/accelerate_configs/deepspeed_zero2.yaml
|
compute_environment: LOCAL_MACHINE
debug: false
deepspeed_config:
deepspeed_multinode_launcher: standard
gradient_accumulation_steps: 1
offload_optimizer_device: none
offload_param_device: none
zero3_init_flag: false
zero_stage: 2
distributed_type: DEEPSPEED
downcast_bf16: 'no'
machine_rank: 0
main_training_function: main
mixed_precision: 'bf16'
num_machines: 1
num_processes: 8
rdzv_backend: static
same_network: true
tpu_env: []
tpu_use_cluster: false
tpu_use_sudo: false
use_cpu: false
| 0
|
hf_public_repos/trl/examples
|
hf_public_repos/trl/examples/accelerate_configs/deepspeed_zero3.yaml
|
compute_environment: LOCAL_MACHINE
debug: false
deepspeed_config:
deepspeed_multinode_launcher: standard
gradient_accumulation_steps: 1
offload_optimizer_device: none
offload_param_device: none
zero3_init_flag: true
zero3_save_16bit_model: true
zero_stage: 3
distributed_type: DEEPSPEED
downcast_bf16: 'no'
machine_rank: 0
main_training_function: main
mixed_precision: 'bf16'
num_machines: 1
num_processes: 8
rdzv_backend: static
same_network: true
tpu_env: []
tpu_use_cluster: false
tpu_use_sudo: false
use_cpu: false
| 0
|
hf_public_repos/trl/examples
|
hf_public_repos/trl/examples/accelerate_configs/multi_gpu.yaml
|
compute_environment: LOCAL_MACHINE
debug: false
distributed_type: MULTI_GPU
downcast_bf16: 'no'
gpu_ids: all
machine_rank: 0
main_training_function: main
mixed_precision: 'bf16'
num_machines: 1
num_processes: 8
rdzv_backend: static
same_network: true
tpu_env: []
tpu_use_cluster: false
tpu_use_sudo: false
use_cpu: false
| 0
|
hf_public_repos
|
hf_public_repos/datasets/Makefile
|
.PHONY: quality style test
check_dirs := tests src benchmarks metrics utils
# Check that source code meets quality standards
quality:
ruff check $(check_dirs) setup.py # linter
ruff format --check $(check_dirs) setup.py # formatter
# Format source code automatically
style:
ruff check --fix $(check_dirs) setup.py # linter
ruff format $(check_dirs) setup.py # formatter
# Run tests for the library
test:
python -m pytest -n auto --dist=loadfile -s -v ./tests/
| 0
|
hf_public_repos
|
hf_public_repos/datasets/ADD_NEW_DATASET.md
|
# How to add one new datasets
Add datasets directly to the 🤗 Hugging Face Hub!
You can share your dataset on https://huggingface.co/datasets directly using your account, see the documentation:
* [Create a dataset and upload files on the website](https://huggingface.co/docs/datasets/upload_dataset)
* [Advanced guide using the CLI](https://huggingface.co/docs/datasets/share)
| 0
|
hf_public_repos
|
hf_public_repos/datasets/dvc.yaml
|
stages:
benchmark_array_xd:
cmd: python ./benchmarks/benchmark_array_xd.py
deps:
- ./benchmarks/benchmark_array_xd.py
metrics:
- ./benchmarks/results/benchmark_array_xd.json:
cache: false
benchmark_indices_mapping:
cmd: python ./benchmarks/benchmark_indices_mapping.py
deps:
- ./benchmarks/benchmark_indices_mapping.py
metrics:
- ./benchmarks/results/benchmark_indices_mapping.json:
cache: false
benchmark_map_filter:
cmd: python ./benchmarks/benchmark_map_filter.py
deps:
- ./benchmarks/benchmark_map_filter.py
metrics:
- ./benchmarks/results/benchmark_map_filter.json:
cache: false
benchmark_iterating:
cmd: python ./benchmarks/benchmark_iterating.py
deps:
- ./benchmarks/benchmark_iterating.py
metrics:
- ./benchmarks/results/benchmark_iterating.json:
cache: false
benchmark_getitem_100B:
cmd: python ./benchmarks/benchmark_getitem_100B.py
deps:
- ./benchmarks/benchmark_getitem_100B.py
metrics:
- ./benchmarks/results/benchmark_getitem_100B.json:
cache: false
| 0
|
hf_public_repos
|
hf_public_repos/datasets/LICENSE
|
Apache License
Version 2.0, January 2004
http://www.apache.org/licenses/
TERMS AND CONDITIONS FOR USE, REPRODUCTION, AND DISTRIBUTION
1. Definitions.
"License" shall mean the terms and conditions for use, reproduction,
and distribution as defined by Sections 1 through 9 of this document.
"Licensor" shall mean the copyright owner or entity authorized by
the copyright owner that is granting the License.
"Legal Entity" shall mean the union of the acting entity and all
other entities that control, are controlled by, or are under common
control with that entity. For the purposes of this definition,
"control" means (i) the power, direct or indirect, to cause the
direction or management of such entity, whether by contract or
otherwise, or (ii) ownership of fifty percent (50%) or more of the
outstanding shares, or (iii) beneficial ownership of such entity.
"You" (or "Your") shall mean an individual or Legal Entity
exercising permissions granted by this License.
"Source" form shall mean the preferred form for making modifications,
including but not limited to software source code, documentation
source, and configuration files.
"Object" form shall mean any form resulting from mechanical
transformation or translation of a Source form, including but
not limited to compiled object code, generated documentation,
and conversions to other media types.
"Work" shall mean the work of authorship, whether in Source or
Object form, made available under the License, as indicated by a
copyright notice that is included in or attached to the work
(an example is provided in the Appendix below).
"Derivative Works" shall mean any work, whether in Source or Object
form, that is based on (or derived from) the Work and for which the
editorial revisions, annotations, elaborations, or other modifications
represent, as a whole, an original work of authorship. For the purposes
of this License, Derivative Works shall not include works that remain
separable from, or merely link (or bind by name) to the interfaces of,
the Work and Derivative Works thereof.
"Contribution" shall mean any work of authorship, including
the original version of the Work and any modifications or additions
to that Work or Derivative Works thereof, that is intentionally
submitted to Licensor for inclusion in the Work by the copyright owner
or by an individual or Legal Entity authorized to submit on behalf of
the copyright owner. For the purposes of this definition, "submitted"
means any form of electronic, verbal, or written communication sent
to the Licensor or its representatives, including but not limited to
communication on electronic mailing lists, source code control systems,
and issue tracking systems that are managed by, or on behalf of, the
Licensor for the purpose of discussing and improving the Work, but
excluding communication that is conspicuously marked or otherwise
designated in writing by the copyright owner as "Not a Contribution."
"Contributor" shall mean Licensor and any individual or Legal Entity
on behalf of whom a Contribution has been received by Licensor and
subsequently incorporated within the Work.
2. Grant of Copyright License. Subject to the terms and conditions of
this License, each Contributor hereby grants to You a perpetual,
worldwide, non-exclusive, no-charge, royalty-free, irrevocable
copyright license to reproduce, prepare Derivative Works of,
publicly display, publicly perform, sublicense, and distribute the
Work and such Derivative Works in Source or Object form.
3. Grant of Patent License. Subject to the terms and conditions of
this License, each Contributor hereby grants to You a perpetual,
worldwide, non-exclusive, no-charge, royalty-free, irrevocable
(except as stated in this section) patent license to make, have made,
use, offer to sell, sell, import, and otherwise transfer the Work,
where such license applies only to those patent claims licensable
by such Contributor that are necessarily infringed by their
Contribution(s) alone or by combination of their Contribution(s)
with the Work to which such Contribution(s) was submitted. If You
institute patent litigation against any entity (including a
cross-claim or counterclaim in a lawsuit) alleging that the Work
or a Contribution incorporated within the Work constitutes direct
or contributory patent infringement, then any patent licenses
granted to You under this License for that Work shall terminate
as of the date such litigation is filed.
4. Redistribution. You may reproduce and distribute copies of the
Work or Derivative Works thereof in any medium, with or without
modifications, and in Source or Object form, provided that You
meet the following conditions:
(a) You must give any other recipients of the Work or
Derivative Works a copy of this License; and
(b) You must cause any modified files to carry prominent notices
stating that You changed the files; and
(c) You must retain, in the Source form of any Derivative Works
that You distribute, all copyright, patent, trademark, and
attribution notices from the Source form of the Work,
excluding those notices that do not pertain to any part of
the Derivative Works; and
(d) If the Work includes a "NOTICE" text file as part of its
distribution, then any Derivative Works that You distribute must
include a readable copy of the attribution notices contained
within such NOTICE file, excluding those notices that do not
pertain to any part of the Derivative Works, in at least one
of the following places: within a NOTICE text file distributed
as part of the Derivative Works; within the Source form or
documentation, if provided along with the Derivative Works; or,
within a display generated by the Derivative Works, if and
wherever such third-party notices normally appear. The contents
of the NOTICE file are for informational purposes only and
do not modify the License. You may add Your own attribution
notices within Derivative Works that You distribute, alongside
or as an addendum to the NOTICE text from the Work, provided
that such additional attribution notices cannot be construed
as modifying the License.
You may add Your own copyright statement to Your modifications and
may provide additional or different license terms and conditions
for use, reproduction, or distribution of Your modifications, or
for any such Derivative Works as a whole, provided Your use,
reproduction, and distribution of the Work otherwise complies with
the conditions stated in this License.
5. Submission of Contributions. Unless You explicitly state otherwise,
any Contribution intentionally submitted for inclusion in the Work
by You to the Licensor shall be under the terms and conditions of
this License, without any additional terms or conditions.
Notwithstanding the above, nothing herein shall supersede or modify
the terms of any separate license agreement you may have executed
with Licensor regarding such Contributions.
6. Trademarks. This License does not grant permission to use the trade
names, trademarks, service marks, or product names of the Licensor,
except as required for reasonable and customary use in describing the
origin of the Work and reproducing the content of the NOTICE file.
7. Disclaimer of Warranty. Unless required by applicable law or
agreed to in writing, Licensor provides the Work (and each
Contributor provides its Contributions) on an "AS IS" BASIS,
WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or
implied, including, without limitation, any warranties or conditions
of TITLE, NON-INFRINGEMENT, MERCHANTABILITY, or FITNESS FOR A
PARTICULAR PURPOSE. You are solely responsible for determining the
appropriateness of using or redistributing the Work and assume any
risks associated with Your exercise of permissions under this License.
8. Limitation of Liability. In no event and under no legal theory,
whether in tort (including negligence), contract, or otherwise,
unless required by applicable law (such as deliberate and grossly
negligent acts) or agreed to in writing, shall any Contributor be
liable to You for damages, including any direct, indirect, special,
incidental, or consequential damages of any character arising as a
result of this License or out of the use or inability to use the
Work (including but not limited to damages for loss of goodwill,
work stoppage, computer failure or malfunction, or any and all
other commercial damages or losses), even if such Contributor
has been advised of the possibility of such damages.
9. Accepting Warranty or Additional Liability. While redistributing
the Work or Derivative Works thereof, You may choose to offer,
and charge a fee for, acceptance of support, warranty, indemnity,
or other liability obligations and/or rights consistent with this
License. However, in accepting such obligations, You may act only
on Your own behalf and on Your sole responsibility, not on behalf
of any other Contributor, and only if You agree to indemnify,
defend, and hold each Contributor harmless for any liability
incurred by, or claims asserted against, such Contributor by reason
of your accepting any such warranty or additional liability.
END OF TERMS AND CONDITIONS
APPENDIX: How to apply the Apache License to your work.
To apply the Apache License to your work, attach the following
boilerplate notice, with the fields enclosed by brackets "[]"
replaced with your own identifying information. (Don't include
the brackets!) The text should be enclosed in the appropriate
comment syntax for the file format. We also recommend that a
file or class name and description of purpose be included on the
same "printed page" as the copyright notice for easier
identification within third-party archives.
Copyright [yyyy] [name of copyright owner]
Licensed under the Apache License, Version 2.0 (the "License");
you may not use this file except in compliance with the License.
You may obtain a copy of the License at
http://www.apache.org/licenses/LICENSE-2.0
Unless required by applicable law or agreed to in writing, software
distributed under the License is distributed on an "AS IS" BASIS,
WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
See the License for the specific language governing permissions and
limitations under the License.
| 0
|
hf_public_repos
|
hf_public_repos/datasets/README.md
|
<p align="center">
<picture>
<source media="(prefers-color-scheme: dark)" srcset="https://huggingface.co/datasets/huggingface/documentation-images/raw/main/datasets-logo-dark.svg">
<source media="(prefers-color-scheme: light)" srcset="https://huggingface.co/datasets/huggingface/documentation-images/raw/main/datasets-logo-light.svg">
<img alt="Hugging Face Datasets Library" src="https://huggingface.co/datasets/huggingface/documentation-images/raw/main/datasets-logo-light.svg" width="352" height="59" style="max-width: 100%;">
</picture>
<br/>
<br/>
</p>
<p align="center">
<a href="https://github.com/huggingface/datasets/actions/workflows/ci.yml?query=branch%3Amain">
<img alt="Build" src="https://github.com/huggingface/datasets/actions/workflows/ci.yml/badge.svg?branch=main">
</a>
<a href="https://github.com/huggingface/datasets/blob/main/LICENSE">
<img alt="GitHub" src="https://img.shields.io/github/license/huggingface/datasets.svg?color=blue">
</a>
<a href="https://huggingface.co/docs/datasets/index.html">
<img alt="Documentation" src="https://img.shields.io/website/http/huggingface.co/docs/datasets/index.html.svg?down_color=red&down_message=offline&up_message=online">
</a>
<a href="https://github.com/huggingface/datasets/releases">
<img alt="GitHub release" src="https://img.shields.io/github/release/huggingface/datasets.svg">
</a>
<a href="https://huggingface.co/datasets/">
<img alt="Number of datasets" src="https://img.shields.io/endpoint?url=https://huggingface.co/api/shields/datasets&color=brightgreen">
</a>
<a href="CODE_OF_CONDUCT.md">
<img alt="Contributor Covenant" src="https://img.shields.io/badge/Contributor%20Covenant-2.0-4baaaa.svg">
</a>
<a href="https://zenodo.org/badge/latestdoi/250213286"><img src="https://zenodo.org/badge/250213286.svg" alt="DOI"></a>
</p>
🤗 Datasets is a lightweight library providing **two** main features:
- **one-line dataloaders for many public datasets**: one-liners to download and pre-process any of the  major public datasets (image datasets, audio datasets, text datasets in 467 languages and dialects, etc.) provided on the [HuggingFace Datasets Hub](https://huggingface.co/datasets). With a simple command like `squad_dataset = load_dataset("squad")`, get any of these datasets ready to use in a dataloader for training/evaluating a ML model (Numpy/Pandas/PyTorch/TensorFlow/JAX),
- **efficient data pre-processing**: simple, fast and reproducible data pre-processing for the public datasets as well as your own local datasets in CSV, JSON, text, PNG, JPEG, WAV, MP3, Parquet, etc. With simple commands like `processed_dataset = dataset.map(process_example)`, efficiently prepare the dataset for inspection and ML model evaluation and training.
[🎓 **Documentation**](https://huggingface.co/docs/datasets/) [🔎 **Find a dataset in the Hub**](https://huggingface.co/datasets) [🌟 **Share a dataset on the Hub**](https://huggingface.co/docs/datasets/share)
<h3 align="center">
<a href="https://hf.co/course"><img src="https://raw.githubusercontent.com/huggingface/datasets/main/docs/source/imgs/course_banner.png"></a>
</h3>
🤗 Datasets is designed to let the community easily add and share new datasets.
🤗 Datasets has many additional interesting features:
- Thrive on large datasets: 🤗 Datasets naturally frees the user from RAM memory limitation, all datasets are memory-mapped using an efficient zero-serialization cost backend (Apache Arrow).
- Smart caching: never wait for your data to process several times.
- Lightweight and fast with a transparent and pythonic API (multi-processing/caching/memory-mapping).
- Built-in interoperability with NumPy, pandas, PyTorch, TensorFlow 2 and JAX.
- Native support for audio and image data.
- Enable streaming mode to save disk space and start iterating over the dataset immediately.
🤗 Datasets originated from a fork of the awesome [TensorFlow Datasets](https://github.com/tensorflow/datasets) and the HuggingFace team want to deeply thank the TensorFlow Datasets team for building this amazing library. More details on the differences between 🤗 Datasets and `tfds` can be found in the section [Main differences between 🤗 Datasets and `tfds`](#main-differences-between--datasets-and-tfds).
# Installation
## With pip
🤗 Datasets can be installed from PyPi and has to be installed in a virtual environment (venv or conda for instance)
```bash
pip install datasets
```
## With conda
🤗 Datasets can be installed using conda as follows:
```bash
conda install -c huggingface -c conda-forge datasets
```
Follow the installation pages of TensorFlow and PyTorch to see how to install them with conda.
For more details on installation, check the installation page in the documentation: https://huggingface.co/docs/datasets/installation
## Installation to use with PyTorch/TensorFlow/pandas
If you plan to use 🤗 Datasets with PyTorch (1.0+), TensorFlow (2.2+) or pandas, you should also install PyTorch, TensorFlow or pandas.
For more details on using the library with NumPy, pandas, PyTorch or TensorFlow, check the quick start page in the documentation: https://huggingface.co/docs/datasets/quickstart
# Usage
🤗 Datasets is made to be very simple to use - the API is centered around a single function, `datasets.load_dataset(dataset_name, **kwargs)`, that instantiates a dataset.
This library can be used for text/image/audio/etc. datasets. Here is an example to load a text dataset:
Here is a quick example:
```python
from datasets import load_dataset
# Print all the available datasets
from huggingface_hub import list_datasets
print([dataset.id for dataset in list_datasets()])
# Load a dataset and print the first example in the training set
squad_dataset = load_dataset('squad')
print(squad_dataset['train'][0])
# Process the dataset - add a column with the length of the context texts
dataset_with_length = squad_dataset.map(lambda x: {"length": len(x["context"])})
# Process the dataset - tokenize the context texts (using a tokenizer from the 🤗 Transformers library)
from transformers import AutoTokenizer
tokenizer = AutoTokenizer.from_pretrained('bert-base-cased')
tokenized_dataset = squad_dataset.map(lambda x: tokenizer(x['context']), batched=True)
```
If your dataset is bigger than your disk or if you don't want to wait to download the data, you can use streaming:
```python
# If you want to use the dataset immediately and efficiently stream the data as you iterate over the dataset
image_dataset = load_dataset('cifar100', streaming=True)
for example in image_dataset["train"]:
break
```
For more details on using the library, check the quick start page in the documentation: https://huggingface.co/docs/datasets/quickstart and the specific pages on:
- Loading a dataset: https://huggingface.co/docs/datasets/loading
- What's in a Dataset: https://huggingface.co/docs/datasets/access
- Processing data with 🤗 Datasets: https://huggingface.co/docs/datasets/process
- Processing audio data: https://huggingface.co/docs/datasets/audio_process
- Processing image data: https://huggingface.co/docs/datasets/image_process
- Processing text data: https://huggingface.co/docs/datasets/nlp_process
- Streaming a dataset: https://huggingface.co/docs/datasets/stream
- Writing your own dataset loading script: https://huggingface.co/docs/datasets/dataset_script
- etc.
# Add a new dataset to the Hub
We have a very detailed step-by-step guide to add a new dataset to the  datasets already provided on the [HuggingFace Datasets Hub](https://huggingface.co/datasets).
You can find:
- [how to upload a dataset to the Hub using your web browser or Python](https://huggingface.co/docs/datasets/upload_dataset) and also
- [how to upload it using Git](https://huggingface.co/docs/datasets/share).
# Main differences between 🤗 Datasets and `tfds`
If you are familiar with the great TensorFlow Datasets, here are the main differences between 🤗 Datasets and `tfds`:
- the scripts in 🤗 Datasets are not provided within the library but are queried, downloaded/cached and dynamically loaded upon request
- the backend serialization of 🤗 Datasets is based on [Apache Arrow](https://arrow.apache.org/) instead of TF Records and leverage python dataclasses for info and features with some diverging features (we mostly don't do encoding and store the raw data as much as possible in the backend serialization cache).
- the user-facing dataset object of 🤗 Datasets is not a `tf.data.Dataset` but a built-in framework-agnostic dataset class with methods inspired by what we like in `tf.data` (like a `map()` method). It basically wraps a memory-mapped Arrow table cache.
# Disclaimers
🤗 Datasets may run Python code defined by the dataset authors to parse certain data formats or structures. For security reasons, we ask users to:
- check the dataset scripts they're going to run beforehand and
- pin the `revision` of the repositories they use.
If you're a dataset owner and wish to update any part of it (description, citation, license, etc.), or do not want your dataset to be included in the Hugging Face Hub, please get in touch by opening a discussion or a pull request in the Community tab of the dataset page. Thanks for your contribution to the ML community!
## BibTeX
If you want to cite our 🤗 Datasets library, you can use our [paper](https://arxiv.org/abs/2109.02846):
```bibtex
@inproceedings{lhoest-etal-2021-datasets,
title = "Datasets: A Community Library for Natural Language Processing",
author = "Lhoest, Quentin and
Villanova del Moral, Albert and
Jernite, Yacine and
Thakur, Abhishek and
von Platen, Patrick and
Patil, Suraj and
Chaumond, Julien and
Drame, Mariama and
Plu, Julien and
Tunstall, Lewis and
Davison, Joe and
{\v{S}}a{\v{s}}ko, Mario and
Chhablani, Gunjan and
Malik, Bhavitvya and
Brandeis, Simon and
Le Scao, Teven and
Sanh, Victor and
Xu, Canwen and
Patry, Nicolas and
McMillan-Major, Angelina and
Schmid, Philipp and
Gugger, Sylvain and
Delangue, Cl{\'e}ment and
Matussi{\`e}re, Th{\'e}o and
Debut, Lysandre and
Bekman, Stas and
Cistac, Pierric and
Goehringer, Thibault and
Mustar, Victor and
Lagunas, Fran{\c{c}}ois and
Rush, Alexander and
Wolf, Thomas",
booktitle = "Proceedings of the 2021 Conference on Empirical Methods in Natural Language Processing: System Demonstrations",
month = nov,
year = "2021",
address = "Online and Punta Cana, Dominican Republic",
publisher = "Association for Computational Linguistics",
url = "https://aclanthology.org/2021.emnlp-demo.21",
pages = "175--184",
abstract = "The scale, variety, and quantity of publicly-available NLP datasets has grown rapidly as researchers propose new tasks, larger models, and novel benchmarks. Datasets is a community library for contemporary NLP designed to support this ecosystem. Datasets aims to standardize end-user interfaces, versioning, and documentation, while providing a lightweight front-end that behaves similarly for small datasets as for internet-scale corpora. The design of the library incorporates a distributed, community-driven approach to adding datasets and documenting usage. After a year of development, the library now includes more than 650 unique datasets, has more than 250 contributors, and has helped support a variety of novel cross-dataset research projects and shared tasks. The library is available at https://github.com/huggingface/datasets.",
eprint={2109.02846},
archivePrefix={arXiv},
primaryClass={cs.CL},
}
```
If you need to cite a specific version of our 🤗 Datasets library for reproducibility, you can use the corresponding version Zenodo DOI from this [list](https://zenodo.org/search?q=conceptrecid:%224817768%22&sort=-version&all_versions=True).
| 0
|
hf_public_repos
|
hf_public_repos/datasets/CONTRIBUTING.md
|
# How to contribute to Datasets?
[](CODE_OF_CONDUCT.md)
Datasets is an open source project, so all contributions and suggestions are welcome.
You can contribute in many different ways: giving ideas, answering questions, reporting bugs, proposing enhancements,
improving the documentation, fixing bugs,...
Many thanks in advance to every contributor.
In order to facilitate healthy, constructive behavior in an open and inclusive community, we all respect and abide by
our [code of conduct](CODE_OF_CONDUCT.md).
## How to work on an open Issue?
You have the list of open Issues at: https://github.com/huggingface/datasets/issues
Some of them may have the label `help wanted`: that means that any contributor is welcomed!
If you would like to work on any of the open Issues:
1. Make sure it is not already assigned to someone else. You have the assignee (if any) on the top of the right column of the Issue page.
2. You can self-assign it by commenting on the Issue page with the keyword: `#self-assign`.
3. Work on your self-assigned issue and eventually create a Pull Request.
## How to create a Pull Request?
If you want to add a dataset see specific instructions in the section [*How to add a dataset*](#how-to-add-a-dataset).
1. Fork the [repository](https://github.com/huggingface/datasets) by clicking on the 'Fork' button on the repository's page. This creates a copy of the code under your GitHub user account.
2. Clone your fork to your local disk, and add the base repository as a remote:
```bash
git clone git@github.com:<your Github handle>/datasets.git
cd datasets
git remote add upstream https://github.com/huggingface/datasets.git
```
3. Create a new branch to hold your development changes:
```bash
git checkout -b a-descriptive-name-for-my-changes
```
**do not** work on the `main` branch.
4. Set up a development environment by running the following command in a virtual environment:
```bash
pip install -e ".[dev]"
```
(If datasets was already installed in the virtual environment, remove
it with `pip uninstall datasets` before reinstalling it in editable
mode with the `-e` flag.)
5. Develop the features on your branch.
6. Format your code. Run `black` and `ruff` so that your newly added files look nice with the following command:
```bash
make style
```
7. _(Optional)_ You can also use [`pre-commit`](https://pre-commit.com/) to format your code automatically each time run `git commit`, instead of running `make style` manually.
To do this, install `pre-commit` via `pip install pre-commit` and then run `pre-commit install` in the project's root directory to set up the hooks.
Note that if any files were formatted by `pre-commit` hooks during committing, you have to run `git commit` again .
8. Once you're happy with your contribution, add your changed files and make a commit to record your changes locally:
```bash
git add -u
git commit
```
It is a good idea to sync your copy of the code with the original
repository regularly. This way you can quickly account for changes:
```bash
git fetch upstream
git rebase upstream/main
```
9. Once you are satisfied, push the changes to your fork repo using:
```bash
git push -u origin a-descriptive-name-for-my-changes
```
Go the webpage of your fork on GitHub. Click on "Pull request" to send your to the project maintainers for review.
## How to add a dataset
You can share your dataset on https://huggingface.co/datasets directly using your account, see the documentation:
* [Create a dataset and upload files on the website](https://huggingface.co/docs/datasets/upload_dataset)
* [Advanced guide using the CLI](https://huggingface.co/docs/datasets/share)
## How to contribute to the dataset cards
Improving the documentation of datasets is an ever-increasing effort, and we invite users to contribute by sharing their insights with the community in the `README.md` dataset cards provided for each dataset.
If you see that a dataset card is missing information that you are in a position to provide (as an author of the dataset or as an experienced user), the best thing you can do is to open a Pull Request on the Hugging Face Hub. To do, go to the "Files and versions" tab of the dataset page and edit the `README.md` file. We provide:
* a [template](https://github.com/huggingface/datasets/blob/main/templates/README.md)
* a [guide](https://github.com/huggingface/datasets/blob/main/templates/README_guide.md) describing what information should go into each of the paragraphs
* and if you need inspiration, we recommend looking through a [completed example](https://huggingface.co/datasets/eli5/blob/main/README.md)
If you are a **dataset author**... you know what to do, it is your dataset after all ;) ! We would especially appreciate if you could help us fill in information about the process of creating the dataset, and take a moment to reflect on its social impact and possible limitations if you haven't already done so in the dataset paper or in another data statement.
If you are a **user of a dataset**, the main source of information should be the dataset paper if it is available: we recommend pulling information from there into the relevant paragraphs of the template. We also eagerly welcome discussions on the [Considerations for Using the Data](https://github.com/huggingface/datasets/blob/main/templates/README_guide.md#considerations-for-using-the-data) based on existing scholarship or personal experience that would benefit the whole community.
Finally, if you want more information on the how and why of dataset cards, we strongly recommend reading the foundational works [Datasheets for Datasets](https://arxiv.org/abs/1803.09010) and [Data Statements for NLP](https://www.aclweb.org/anthology/Q18-1041/).
Thank you for your contribution!
## Code of conduct
This project adheres to the HuggingFace [code of conduct](CODE_OF_CONDUCT.md).
By participating, you are expected to abide by this code.
| 0
|
hf_public_repos
|
hf_public_repos/datasets/CODE_OF_CONDUCT.md
|
# Contributor Covenant Code of Conduct
## Our Pledge
We as members, contributors, and leaders pledge to make participation in our
community a harassment-free experience for everyone, regardless of age, body
size, visible or invisible disability, ethnicity, sex characteristics, gender
identity and expression, level of experience, education, socio-economic status,
nationality, personal appearance, race, caste, color, religion, or sexual identity
and orientation.
We pledge to act and interact in ways that contribute to an open, welcoming,
diverse, inclusive, and healthy community.
## Our Standards
Examples of behavior that contributes to a positive environment for our
community include:
* Demonstrating empathy and kindness toward other people
* Being respectful of differing opinions, viewpoints, and experiences
* Giving and gracefully accepting constructive feedback
* Accepting responsibility and apologizing to those affected by our mistakes,
and learning from the experience
* Focusing on what is best not just for us as individuals, but for the
overall community
Examples of unacceptable behavior include:
* The use of sexualized language or imagery, and sexual attention or
advances of any kind
* Trolling, insulting or derogatory comments, and personal or political attacks
* Public or private harassment
* Publishing others' private information, such as a physical or email
address, without their explicit permission
* Other conduct which could reasonably be considered inappropriate in a
professional setting
## Enforcement Responsibilities
Community leaders are responsible for clarifying and enforcing our standards of
acceptable behavior and will take appropriate and fair corrective action in
response to any behavior that they deem inappropriate, threatening, offensive,
or harmful.
Community leaders have the right and responsibility to remove, edit, or reject
comments, commits, code, wiki edits, issues, and other contributions that are
not aligned to this Code of Conduct, and will communicate reasons for moderation
decisions when appropriate.
## Scope
This Code of Conduct applies within all community spaces, and also applies when
an individual is officially representing the community in public spaces.
Examples of representing our community include using an official e-mail address,
posting via an official social media account, or acting as an appointed
representative at an online or offline event.
## Enforcement
Instances of abusive, harassing, or otherwise unacceptable behavior may be
reported to the community leaders responsible for enforcement at
feedback@huggingface.co.
All complaints will be reviewed and investigated promptly and fairly.
All community leaders are obligated to respect the privacy and security of the
reporter of any incident.
## Enforcement Guidelines
Community leaders will follow these Community Impact Guidelines in determining
the consequences for any action they deem in violation of this Code of Conduct:
### 1. Correction
**Community Impact**: Use of inappropriate language or other behavior deemed
unprofessional or unwelcome in the community.
**Consequence**: A private, written warning from community leaders, providing
clarity around the nature of the violation and an explanation of why the
behavior was inappropriate. A public apology may be requested.
### 2. Warning
**Community Impact**: A violation through a single incident or series
of actions.
**Consequence**: A warning with consequences for continued behavior. No
interaction with the people involved, including unsolicited interaction with
those enforcing the Code of Conduct, for a specified period of time. This
includes avoiding interactions in community spaces as well as external channels
like social media. Violating these terms may lead to a temporary or
permanent ban.
### 3. Temporary Ban
**Community Impact**: A serious violation of community standards, including
sustained inappropriate behavior.
**Consequence**: A temporary ban from any sort of interaction or public
communication with the community for a specified period of time. No public or
private interaction with the people involved, including unsolicited interaction
with those enforcing the Code of Conduct, is allowed during this period.
Violating these terms may lead to a permanent ban.
### 4. Permanent Ban
**Community Impact**: Demonstrating a pattern of violation of community
standards, including sustained inappropriate behavior, harassment of an
individual, or aggression toward or disparagement of classes of individuals.
**Consequence**: A permanent ban from any sort of public interaction within
the community.
## Attribution
This Code of Conduct is adapted from the [Contributor Covenant][homepage],
version 2.0, available at
[https://www.contributor-covenant.org/version/2/0/code_of_conduct.html][v2.0].
Community Impact Guidelines were inspired by
[Mozilla's code of conduct enforcement ladder][Mozilla CoC].
For answers to common questions about this code of conduct, see the FAQ at
[https://www.contributor-covenant.org/faq][FAQ]. Translations are available
at [https://www.contributor-covenant.org/translations][translations].
[homepage]: https://www.contributor-covenant.org
[v2.0]: https://www.contributor-covenant.org/version/2/0/code_of_conduct.html
[Mozilla CoC]: https://github.com/mozilla/diversity
[FAQ]: https://www.contributor-covenant.org/faq
[translations]: https://www.contributor-covenant.org/translations
| 0
|
hf_public_repos
|
hf_public_repos/datasets/.pre-commit-config.yaml
|
repos:
- repo: https://github.com/charliermarsh/ruff-pre-commit # https://github.com/charliermarsh/ruff#usage
rev: 'v0.1.5'
hooks:
# Run the linter.
- id: ruff
args: [ --fix ]
# Run the formatter.
- id: ruff-format
| 0
|
hf_public_repos
|
hf_public_repos/datasets/additional-tests-requirements.txt
|
unbabel-comet>=1.0.0
git+https://github.com/google-research/bleurt.git
git+https://github.com/ns-moosavi/coval.git
git+https://github.com/hendrycks/math.git
| 0
|
hf_public_repos
|
hf_public_repos/datasets/.zenodo.json
|
{
"license": "Apache-2.0",
"creators": [
{
"affiliation": "Hugging Face",
"name": "Quentin Lhoest"
},
{
"orcid": "0000-0003-1727-1045",
"affiliation": "Hugging Face",
"name": "Albert Villanova del Moral"
},
{
"affiliation": "Hugging Face",
"name": "Patrick von Platen"
},
{
"affiliation": "Hugging Face",
"name": "Thomas Wolf"
},
{
"affiliation": "Hugging Face",
"name": "Mario Šaško"
},
{
"affiliation": "Hugging Face",
"name": "Yacine Jernite"
},
{
"affiliation": "Hugging Face",
"name": "Abhishek Thakur"
},
{
"affiliation": "Hugging Face",
"name": "Lewis Tunstall"
},
{
"affiliation": "Hugging Face",
"name": "Suraj Patil"
},
{
"affiliation": "Hugging Face",
"name": "Mariama Drame"
},
{
"affiliation": "Hugging Face",
"name": "Julien Chaumond"
},
{
"affiliation": "Hugging Face",
"name": "Julien Plu"
},
{
"affiliation": "Hugging Face",
"name": "Joe Davison"
},
{
"affiliation": "Hugging Face",
"name": "Simon Brandeis"
},
{
"affiliation": "Hugging Face",
"name": "Victor Sanh"
},
{
"affiliation": "Hugging Face",
"name": "Teven Le Scao"
},
{
"affiliation": "Hugging Face",
"name": "Kevin Canwen Xu"
},
{
"affiliation": "Hugging Face",
"name": "Nicolas Patry"
},
{
"affiliation": "Hugging Face",
"name": "Steven Liu"
},
{
"affiliation": "Hugging Face",
"name": "Angelina McMillan-Major"
},
{
"affiliation": "Hugging Face",
"name": "Philipp Schmid"
},
{
"affiliation": "Hugging Face",
"name": "Sylvain Gugger"
},
{
"affiliation": "Hugging Face",
"name": "Nathan Raw"
},
{
"affiliation": "Hugging Face",
"name": "Sylvain Lesage"
},
{
"affiliation": "Hugging Face",
"name": "Anton Lozhkov"
},
{
"affiliation": "Hugging Face",
"name": "Matthew Carrigan"
},
{
"affiliation": "Hugging Face",
"name": "Th\u00e9o Matussi\u00e8re"
},
{
"affiliation": "Hugging Face",
"name": "Leandro von Werra"
},
{
"affiliation": "Hugging Face",
"name": "Lysandre Debut"
},
{
"affiliation": "Hugging Face",
"name": "Stas Bekman"
},
{
"affiliation": "Hugging Face",
"name": "Cl\u00e9ment Delangue"
}
]
}
| 0
|
hf_public_repos
|
hf_public_repos/datasets/setup.py
|
# Lint as: python3
""" HuggingFace/Datasets is an open library of datasets.
Note:
VERSION needs to be formatted following the MAJOR.MINOR.PATCH convention
(we need to follow this convention to be able to retrieve versioned scripts)
Simple check list for release from AllenNLP repo: https://github.com/allenai/allennlp/blob/master/setup.py
Steps to make a release:
0. Prerequisites:
- Dependencies:
- twine: `pip install twine`
- Create an account in (and join the 'datasets' project):
- PyPI: https://pypi.org/
- Test PyPI: https://test.pypi.org/
- Don't break `transformers`: run the `transformers` CI using the `main` branch and make sure it's green.
- In `transformers`, use `datasets @ git+https://github.com/huggingface/datasets@main#egg=datasets`
Add a step to install `datasets@main` after `save_cache` in .circleci/create_circleci_config.py:
```
steps.append({"run": {"name": "Install `datasets@main`", "command": 'pip uninstall datasets -y && pip install "datasets @ git+https://github.com/huggingface/datasets@main#egg=datasets"'}})
```
- and then run the CI
1. Create the release branch from main branch:
```
git checkout main
git pull upstream main
git checkout -b release-VERSION
```
2. Change the version to the release VERSION in:
- __init__.py
- setup.py
3. Commit these changes, push and create a Pull Request:
```
git add -u
git commit -m "Release: VERSION"
git push upstream release-VERSION
```
- Go to: https://github.com/huggingface/datasets/pull/new/release
- Create pull request
4. From your local release branch, build both the sources and the wheel. Do not change anything in setup.py between
creating the wheel and the source distribution (obviously).
- First, delete any building directories that may exist from previous builds:
- build
- dist
- From the top level directory, build the wheel and the sources:
```
python setup.py bdist_wheel
python setup.py sdist
```
- You should now have a /dist directory with both .whl and .tar.gz source versions.
5. Check that everything looks correct by uploading the package to the test PyPI server:
```
twine upload dist/* -r pypitest --repository-url=https://test.pypi.org/legacy/
```
Check that you can install it in a virtualenv/notebook by running:
```
pip install huggingface_hub fsspec aiohttp pyarrow-hotfix
pip install -U tqdm
pip install -i https://testpypi.python.org/pypi datasets
```
6. Upload the final version to the actual PyPI:
```
twine upload dist/* -r pypi
```
7. Make the release on GitHub once everything is looking hunky-dory:
- Merge the release Pull Request
- Create a new release: https://github.com/huggingface/datasets/releases/new
- Choose a tag: Introduce the new VERSION as tag, that will be created when you publish the release
- Create new tag VERSION on publish
- Release title: Introduce the new VERSION as well
- Describe the release
- Use "Generate release notes" button for automatic generation
- Publish release
8. Set the dev version
- Create the dev-version branch from the main branch:
```
git checkout main
git pull upstream main
git branch -D dev-version
git checkout -b dev-version
```
- Change the version to X.X.X+1.dev0 (e.g. VERSION=1.18.3 -> 1.18.4.dev0) in:
- __init__.py
- setup.py
- Commit these changes, push and create a Pull Request:
```
git add -u
git commit -m "Set dev version"
git push upstream dev-version
```
- Go to: https://github.com/huggingface/datasets/pull/new/dev-version
- Create pull request
- Merge the dev version Pull Request
"""
from setuptools import find_packages, setup
REQUIRED_PKGS = [
# For file locking
"filelock",
# We use numpy>=1.17 to have np.random.Generator (Dataset shuffling)
"numpy>=1.17",
# Backend and serialization.
# Minimum 8.0.0 to be able to use .to_reader()
"pyarrow>=8.0.0",
# As long as we allow pyarrow < 14.0.1, to fix vulnerability CVE-2023-47248
"pyarrow-hotfix",
# For smart caching dataset processing
"dill>=0.3.0,<0.3.8", # tmp pin until dill has official support for determinism see https://github.com/uqfoundation/dill/issues/19
# For performance gains with apache arrow
"pandas",
# for downloading datasets over HTTPS
"requests>=2.19.0",
# progress bars in download and scripts
"tqdm>=4.62.1",
# for fast hashing
"xxhash",
# for better multiprocessing
"multiprocess",
# to save datasets locally or on any filesystem
# minimum 2023.1.0 to support protocol=kwargs in fsspec's `open`, `get_fs_token_paths`, etc.: see https://github.com/fsspec/filesystem_spec/pull/1143
"fsspec[http]>=2023.1.0,<=2023.10.0",
# for data streaming via http
"aiohttp",
# To get datasets from the Datasets Hub on huggingface.co
"huggingface_hub>=0.19.4",
# Utilities from PyPA to e.g., compare versions
"packaging",
# To parse YAML metadata from dataset cards
"pyyaml>=5.1",
]
AUDIO_REQUIRE = [
"soundfile>=0.12.1",
"librosa",
]
VISION_REQUIRE = [
"Pillow>=6.2.1",
]
BENCHMARKS_REQUIRE = [
"tensorflow==2.12.0",
"torch==2.0.1",
"transformers==4.30.1",
]
TESTS_REQUIRE = [
# test dependencies
"absl-py",
"joblib<1.3.0", # joblibspark doesn't support recent joblib versions
"joblibspark",
"pytest",
"pytest-datadir",
"pytest-xdist",
# optional dependencies
"apache-beam>=2.26.0,<2.44.0;python_version<'3.10'", # doesn't support recent dill versions for recent python versions
"elasticsearch<8.0.0", # 8.0 asks users to provide hosts or cloud_id when instantiating ElasticSearch()
"faiss-cpu>=1.6.4",
"jax>=0.3.14; sys_platform != 'win32'",
"jaxlib>=0.3.14; sys_platform != 'win32'",
"lz4",
"pyspark>=3.4", # https://issues.apache.org/jira/browse/SPARK-40991 fixed in 3.4.0
"py7zr",
"rarfile>=4.0",
"sqlalchemy<2.0.0",
"s3fs>=2021.11.1", # aligned with fsspec[http]>=2021.11.1; test only on python 3.7 for now
"tensorflow>=2.3,!=2.6.0,!=2.6.1; sys_platform != 'darwin' or platform_machine != 'arm64'",
"tensorflow-macos; sys_platform == 'darwin' and platform_machine == 'arm64'",
"tiktoken",
"torch>=2.0.0",
"soundfile>=0.12.1",
"transformers",
"typing-extensions>=4.6.1", # due to conflict between apache-beam and pydantic
"zstandard",
]
METRICS_TESTS_REQUIRE = [
# metrics dependencies
"accelerate", # for frugalscore (calls transformers' Trainer)
"bert_score>=0.3.6",
"jiwer",
"langdetect",
"mauve-text",
"nltk",
"rouge_score",
"sacrebleu",
"sacremoses",
"scikit-learn",
"scipy",
"sentencepiece", # for bleurt
"seqeval",
"spacy>=3.0.0",
"tldextract",
# to speed up pip backtracking
"toml>=0.10.1",
"typer<0.5.0", # pinned to work with Spacy==3.4.3 on Windows: see https://github.com/tiangolo/typer/issues/427
"requests_file>=1.5.1",
"tldextract>=3.1.0",
"texttable>=1.6.3",
"Werkzeug>=1.0.1",
"six~=1.15.0",
]
TESTS_REQUIRE.extend(VISION_REQUIRE)
TESTS_REQUIRE.extend(AUDIO_REQUIRE)
QUALITY_REQUIRE = ["ruff>=0.1.5"]
DOCS_REQUIRE = [
# Might need to add doc-builder and some specific deps in the future
"s3fs",
# Following dependencies are required for the Python reference to be built properly
"transformers",
"torch",
"tensorflow>=2.2.0,!=2.6.0,!=2.6.1; sys_platform != 'darwin' or platform_machine != 'arm64'",
"tensorflow-macos; sys_platform == 'darwin' and platform_machine == 'arm64'",
]
EXTRAS_REQUIRE = {
"audio": AUDIO_REQUIRE,
"vision": VISION_REQUIRE,
"apache-beam": ["apache-beam>=2.26.0,<2.44.0"],
"tensorflow": [
"tensorflow>=2.2.0,!=2.6.0,!=2.6.1; sys_platform != 'darwin' or platform_machine != 'arm64'",
"tensorflow-macos; sys_platform == 'darwin' and platform_machine == 'arm64'",
],
"tensorflow_gpu": ["tensorflow-gpu>=2.2.0,!=2.6.0,!=2.6.1"],
"torch": ["torch"],
"jax": ["jax>=0.3.14", "jaxlib>=0.3.14"],
"s3": ["s3fs"],
"streaming": [], # for backward compatibility
"dev": TESTS_REQUIRE + QUALITY_REQUIRE + DOCS_REQUIRE,
"tests": TESTS_REQUIRE,
"metrics-tests": METRICS_TESTS_REQUIRE,
"quality": QUALITY_REQUIRE,
"benchmarks": BENCHMARKS_REQUIRE,
"docs": DOCS_REQUIRE,
}
setup(
name="datasets",
version="2.15.1.dev0", # expected format is one of x.y.z.dev0, or x.y.z.rc1 or x.y.z (no to dashes, yes to dots)
description="HuggingFace community-driven open-source library of datasets",
long_description=open("README.md", encoding="utf-8").read(),
long_description_content_type="text/markdown",
author="HuggingFace Inc.",
author_email="thomas@huggingface.co",
url="https://github.com/huggingface/datasets",
download_url="https://github.com/huggingface/datasets/tags",
license="Apache 2.0",
package_dir={"": "src"},
packages=find_packages("src"),
package_data={
"datasets": ["py.typed"],
"datasets.utils.resources": ["*.json", "*.yaml", "*.tsv"],
},
entry_points={"console_scripts": ["datasets-cli=datasets.commands.datasets_cli:main"]},
python_requires=">=3.8.0",
install_requires=REQUIRED_PKGS,
extras_require=EXTRAS_REQUIRE,
classifiers=[
"Development Status :: 5 - Production/Stable",
"Intended Audience :: Developers",
"Intended Audience :: Education",
"Intended Audience :: Science/Research",
"License :: OSI Approved :: Apache Software License",
"Operating System :: OS Independent",
"Programming Language :: Python :: 3",
"Programming Language :: Python :: 3.8",
"Programming Language :: Python :: 3.9",
"Programming Language :: Python :: 3.10",
"Topic :: Scientific/Engineering :: Artificial Intelligence",
],
keywords="datasets machine learning datasets metrics",
zip_safe=False, # Required for mypy to find the py.typed file
)
| 0
|
hf_public_repos
|
hf_public_repos/datasets/AUTHORS
|
# This is the list of HuggingFace Datasets authors for copyright purposes.
#
# This does not necessarily list everyone who has contributed code, since in
# some cases, their employer may be the copyright holder. To see the full list
# of contributors, see the revision history in source control.
Google Inc.
HuggingFace Inc.
| 0
|
hf_public_repos
|
hf_public_repos/datasets/setup.cfg
|
[metadata]
license_files = LICENSE
[tool:pytest]
# Test fails if a FutureWarning is thrown by `huggingface_hub`
filterwarnings =
error::FutureWarning:huggingface_hub*
markers =
unit: unit test
integration: integration test
| 0
|
hf_public_repos
|
hf_public_repos/datasets/CITATION.cff
|
cff-version: 1.2.0
message: "If you use this software, please cite it as below."
title: "huggingface/datasets"
authors:
- family-names: Lhoest
given-names: Quentin
- family-names: Villanova del Moral
given-names: Albert
orcid: "https://orcid.org/0000-0003-1727-1045"
- family-names: von Platen
given-names: Patrick
- family-names: Wolf
given-names: Thomas
- family-names: Šaško
given-names: Mario
- family-names: Jernite
given-names: Yacine
- family-names: Thakur
given-names: Abhishek
- family-names: Tunstall
given-names: Lewis
- family-names: Patil
given-names: Suraj
- family-names: Drame
given-names: Mariama
- family-names: Chaumond
given-names: Julien
- family-names: Plu
given-names: Julien
- family-names: Davison
given-names: Joe
- family-names: Brandeis
given-names: Simon
- family-names: Sanh
given-names: Victor
- family-names: Le Scao
given-names: Teven
- family-names: Canwen Xu
given-names: Kevin
- family-names: Patry
given-names: Nicolas
- family-names: Liu
given-names: Steven
- family-names: McMillan-Major
given-names: Angelina
- family-names: Schmid
given-names: Philipp
- family-names: Gugger
given-names: Sylvain
- family-names: Raw
given-names: Nathan
- family-names: Lesage
given-names: Sylvain
- family-names: Lozhkov
given-names: Anton
- family-names: Carrigan
given-names: Matthew
- family-names: Matussière
given-names: Théo
- family-names: von Werra
given-names: Leandro
- family-names: Debut
given-names: Lysandre
- family-names: Bekman
given-names: Stas
- family-names: Delangue
given-names: Clément
doi: 10.5281/zenodo.4817768
repository-code: "https://github.com/huggingface/datasets"
license: Apache-2.0
preferred-citation:
type: conference-paper
title: "Datasets: A Community Library for Natural Language Processing"
authors:
- family-names: Lhoest
given-names: Quentin
- family-names: Villanova del Moral
given-names: Albert
orcid: "https://orcid.org/0000-0003-1727-1045"
- family-names: von Platen
given-names: Patrick
- family-names: Wolf
given-names: Thomas
- family-names: Šaško
given-names: Mario
- family-names: Jernite
given-names: Yacine
- family-names: Thakur
given-names: Abhishek
- family-names: Tunstall
given-names: Lewis
- family-names: Patil
given-names: Suraj
- family-names: Drame
given-names: Mariama
- family-names: Chaumond
given-names: Julien
- family-names: Plu
given-names: Julien
- family-names: Davison
given-names: Joe
- family-names: Brandeis
given-names: Simon
- family-names: Sanh
given-names: Victor
- family-names: Le Scao
given-names: Teven
- family-names: Canwen Xu
given-names: Kevin
- family-names: Patry
given-names: Nicolas
- family-names: Liu
given-names: Steven
- family-names: McMillan-Major
given-names: Angelina
- family-names: Schmid
given-names: Philipp
- family-names: Gugger
given-names: Sylvain
- family-names: Raw
given-names: Nathan
- family-names: Lesage
given-names: Sylvain
- family-names: Lozhkov
given-names: Anton
- family-names: Carrigan
given-names: Matthew
- family-names: Matussière
given-names: Théo
- family-names: von Werra
given-names: Leandro
- family-names: Debut
given-names: Lysandre
- family-names: Bekman
given-names: Stas
- family-names: Delangue
given-names: Clément
collection-title: "Proceedings of the 2021 Conference on Empirical Methods in Natural Language Processing: System Demonstrations"
collection-type: proceedings
month: 11
year: 2021
publisher:
name: "Association for Computational Linguistics"
url: "https://aclanthology.org/2021.emnlp-demo.21"
start: 175
end: 184
identifiers:
- type: other
value: "arXiv:2109.02846"
description: "The arXiv preprint of the paper"
| 0
|
hf_public_repos
|
hf_public_repos/datasets/.dvcignore
|
# Add patterns of files dvc should ignore, which could improve
# the performance. Learn more at
# https://dvc.org/doc/user-guide/dvcignore
| 0
|
hf_public_repos
|
hf_public_repos/datasets/pyproject.toml
|
[tool.black]
line-length = 119
target_version = ['py37']
[tool.ruff]
# Ignored rules:
# "E501" -> line length violation
# "F821" -> undefined named in type annotation (e.g. Literal["something"])
# "C901" -> `function_name` is too complex
ignore = ["E501", "F821", "C901"]
select = ["C", "E", "F", "I", "W"]
line-length = 119
[tool.ruff.isort]
lines-after-imports = 2
known-first-party = ["datasets"]
| 0
|
hf_public_repos
|
hf_public_repos/datasets/SECURITY.md
|
# Security Policy
## Supported Versions
<!--
Use this section to tell people about which versions of your project are
currently being supported with security updates.
| Version | Supported |
| ------- | ------------------ |
| 5.1.x | :white_check_mark: |
| 5.0.x | :x: |
| 4.0.x | :white_check_mark: |
| < 4.0 | :x: |
-->
Each major version is currently being supported with security updates.
| Version | Supported |
|---------|--------------------|
| 1.x.x | :white_check_mark: |
| 2.x.x | :white_check_mark: |
## Reporting a Vulnerability
<!--
Use this section to tell people how to report a vulnerability.
Tell them where to go, how often they can expect to get an update on a
reported vulnerability, what to expect if the vulnerability is accepted or
declined, etc.
-->
To report a security vulnerability, please contact: security@huggingface.co
| 0
|
hf_public_repos/datasets/src
|
hf_public_repos/datasets/src/datasets/keyhash.py
|
# Copyright 2020 The HuggingFace Datasets Authors and the TensorFlow Datasets Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# Lint as: python3
"""
Hashing function for dataset keys using `hashlib.md5`
Requirements for the hash function:
- Provides a uniformly distributed hash from random space
- Adequately fast speed
- Working with multiple input types (in this case, `str`, `int` or `bytes`)
- Should be platform independent (generates same hash on different OS and systems)
The hashing function provides a unique 128-bit integer hash of the key provided.
The split name is being used here as the hash salt to avoid having same hashes
in different splits due to same keys
"""
from typing import Union
from huggingface_hub.utils import insecure_hashlib
def _as_bytes(hash_data: Union[str, int, bytes]) -> bytes:
"""
Returns the input hash_data in its bytes form
Args:
hash_data: the hash salt/key to be converted to bytes
"""
if isinstance(hash_data, bytes):
# Data already in bytes, returns as it as
return hash_data
elif isinstance(hash_data, str):
# We keep the data as it as for it ot be later encoded to UTF-8
# However replace `\\` with `/` for Windows compatibility
hash_data = hash_data.replace("\\", "/")
elif isinstance(hash_data, int):
hash_data = str(hash_data)
else:
# If data is not of the required type, raise error
raise InvalidKeyError(hash_data)
return hash_data.encode("utf-8")
class InvalidKeyError(Exception):
"""Raises an error when given key is of invalid datatype."""
def __init__(self, hash_data):
self.prefix = "\nFAILURE TO GENERATE DATASET: Invalid key type detected"
self.err_msg = f"\nFound Key {hash_data} of type {type(hash_data)}"
self.suffix = "\nKeys should be either str, int or bytes type"
super().__init__(f"{self.prefix}{self.err_msg}{self.suffix}")
class DuplicatedKeysError(Exception):
"""Raise an error when duplicate key found."""
def __init__(self, key, duplicate_key_indices, fix_msg=""):
self.key = key
self.duplicate_key_indices = duplicate_key_indices
self.fix_msg = fix_msg
self.prefix = "Found multiple examples generated with the same key"
if len(duplicate_key_indices) <= 20:
self.err_msg = f"\nThe examples at index {', '.join(duplicate_key_indices)} have the key {key}"
else:
self.err_msg = f"\nThe examples at index {', '.join(duplicate_key_indices[:20])}... ({len(duplicate_key_indices) - 20} more) have the key {key}"
self.suffix = "\n" + fix_msg if fix_msg else ""
super().__init__(f"{self.prefix}{self.err_msg}{self.suffix}")
class KeyHasher:
"""KeyHasher class for providing hash using md5"""
def __init__(self, hash_salt: str):
self._split_md5 = insecure_hashlib.md5(_as_bytes(hash_salt))
def hash(self, key: Union[str, int, bytes]) -> int:
"""Returns 128-bits unique hash of input key
Args:
key: the input key to be hashed (should be str, int or bytes)
Returns: 128-bit int hash key"""
md5 = self._split_md5.copy()
byte_key = _as_bytes(key)
md5.update(byte_key)
# Convert to integer with hexadecimal conversion
return int(md5.hexdigest(), 16)
| 0
|
hf_public_repos/datasets/src
|
hf_public_repos/datasets/src/datasets/streaming.py
|
import importlib
import inspect
from functools import wraps
from typing import TYPE_CHECKING, Optional
from .download.download_config import DownloadConfig
from .download.streaming_download_manager import (
xbasename,
xdirname,
xet_parse,
xexists,
xgetsize,
xglob,
xgzip_open,
xisdir,
xisfile,
xjoin,
xlistdir,
xnumpy_load,
xopen,
xpandas_read_csv,
xpandas_read_excel,
xPath,
xpyarrow_parquet_read_table,
xrelpath,
xsio_loadmat,
xsplit,
xsplitext,
xwalk,
xxml_dom_minidom_parse,
)
from .utils.logging import get_logger
from .utils.patching import patch_submodule
from .utils.py_utils import get_imports
logger = get_logger(__name__)
if TYPE_CHECKING:
from .builder import DatasetBuilder
def extend_module_for_streaming(module_path, download_config: Optional[DownloadConfig] = None):
"""Extend the module to support streaming.
We patch some functions in the module to use `fsspec` to support data streaming:
- We use `fsspec.open` to open and read remote files. We patch the module function:
- `open`
- We use the "::" hop separator to join paths and navigate remote compressed/archive files. We patch the module
functions:
- `os.path.join`
- `pathlib.Path.joinpath` and `pathlib.Path.__truediv__` (called when using the "/" operator)
The patched functions are replaced with custom functions defined to work with the
:class:`~download.streaming_download_manager.StreamingDownloadManager`.
Args:
module_path: Path to the module to be extended.
download_config : mainly use use_auth_token or storage_options to support different platforms and auth types.
"""
module = importlib.import_module(module_path)
# TODO(QL): always update the module to add subsequent new authentication without removing old ones
if hasattr(module, "_patched_for_streaming") and module._patched_for_streaming:
if isinstance(module._patched_for_streaming, DownloadConfig):
module._patched_for_streaming.token = download_config.token
module._patched_for_streaming.storage_options = download_config.storage_options
return
def wrap_auth(function):
@wraps(function)
def wrapper(*args, **kwargs):
return function(*args, download_config=download_config, **kwargs)
wrapper._decorator_name_ = "wrap_auth"
return wrapper
# open files in a streaming fashion
patch_submodule(module, "open", wrap_auth(xopen)).start()
patch_submodule(module, "os.listdir", wrap_auth(xlistdir)).start()
patch_submodule(module, "os.walk", wrap_auth(xwalk)).start()
patch_submodule(module, "glob.glob", wrap_auth(xglob)).start()
# allow to navigate in remote zip files
patch_submodule(module, "os.path.join", xjoin).start()
patch_submodule(module, "os.path.dirname", xdirname).start()
patch_submodule(module, "os.path.basename", xbasename).start()
patch_submodule(module, "os.path.relpath", xrelpath).start()
patch_submodule(module, "os.path.split", xsplit).start()
patch_submodule(module, "os.path.splitext", xsplitext).start()
# allow checks on paths
patch_submodule(module, "os.path.exists", wrap_auth(xexists)).start()
patch_submodule(module, "os.path.isdir", wrap_auth(xisdir)).start()
patch_submodule(module, "os.path.isfile", wrap_auth(xisfile)).start()
patch_submodule(module, "os.path.getsize", wrap_auth(xgetsize)).start()
patch_submodule(module, "pathlib.Path", xPath).start()
# file readers
patch_submodule(module, "gzip.open", wrap_auth(xgzip_open)).start()
patch_submodule(module, "numpy.load", wrap_auth(xnumpy_load)).start()
patch_submodule(module, "pandas.read_csv", wrap_auth(xpandas_read_csv), attrs=["__version__"]).start()
patch_submodule(module, "pandas.read_excel", wrap_auth(xpandas_read_excel), attrs=["__version__"]).start()
patch_submodule(module, "scipy.io.loadmat", wrap_auth(xsio_loadmat), attrs=["__version__"]).start()
patch_submodule(module, "xml.etree.ElementTree.parse", wrap_auth(xet_parse)).start()
patch_submodule(module, "xml.dom.minidom.parse", wrap_auth(xxml_dom_minidom_parse)).start()
# pyarrow: do not patch pyarrow attribute in packaged modules
if not module.__name__.startswith("datasets.packaged_modules."):
patch_submodule(module, "pyarrow.parquet.read_table", wrap_auth(xpyarrow_parquet_read_table)).start()
module._patched_for_streaming = download_config
def extend_dataset_builder_for_streaming(builder: "DatasetBuilder"):
"""Extend the dataset builder module and the modules imported by it to support streaming.
Args:
builder (:class:`DatasetBuilder`): Dataset builder instance.
"""
# this extends the open and os.path.join functions for data streaming
download_config = DownloadConfig(storage_options=builder.storage_options, token=builder.token)
extend_module_for_streaming(builder.__module__, download_config=download_config)
# if needed, we also have to extend additional internal imports (like wmt14 -> wmt_utils)
if not builder.__module__.startswith("datasets."): # check that it's not a packaged builder like csv
for imports in get_imports(inspect.getfile(builder.__class__)):
if imports[0] == "internal":
internal_import_name = imports[1]
internal_module_name = ".".join(builder.__module__.split(".")[:-1] + [internal_import_name])
extend_module_for_streaming(internal_module_name, download_config=download_config)
# builders can inherit from other builders that might use streaming functionality
# (for example, ImageFolder and AudioFolder inherit from FolderBuilder which implements examples generation)
# but these parents builders are not patched automatically as they are not instantiated, so we patch them here
from .builder import DatasetBuilder
parent_builder_modules = [
cls.__module__
for cls in type(builder).__mro__[1:] # make sure it's not the same module we've already patched
if issubclass(cls, DatasetBuilder) and cls.__module__ != DatasetBuilder.__module__
] # check it's not a standard builder from datasets.builder
for module in parent_builder_modules:
extend_module_for_streaming(module, download_config=download_config)
| 0
|
hf_public_repos/datasets/src
|
hf_public_repos/datasets/src/datasets/fingerprint.py
|
import inspect
import os
import random
import shutil
import tempfile
import weakref
from functools import wraps
from pathlib import Path
from typing import TYPE_CHECKING, Any, Callable, Dict, List, Optional, Tuple, Union
import numpy as np
import xxhash
from .naming import INVALID_WINDOWS_CHARACTERS_IN_PATH
from .utils._dill import dumps
from .utils.deprecation_utils import deprecated
from .utils.logging import get_logger
if TYPE_CHECKING:
from .arrow_dataset import Dataset
logger = get_logger(__name__)
# Fingerprinting allows to have one deterministic fingerprint per dataset state.
# A dataset fingerprint is updated after each transform.
# Re-running the same transforms on a dataset in a different session results in the same fingerprint.
# This is possible thanks to a custom hashing function that works with most python objects.
# Fingerprinting is the main mechanism that enables caching.
# The caching mechanism allows to reload an existing cache file if it's already been computed.
#################
# Caching
#################
_CACHING_ENABLED = True
_TEMP_DIR_FOR_TEMP_CACHE_FILES: Optional["_TempDirWithCustomCleanup"] = None
_DATASETS_WITH_TABLE_IN_TEMP_DIR: Optional[weakref.WeakSet] = None
class _TempDirWithCustomCleanup:
"""
A temporary directory with a custom cleanup function.
We need a custom temporary directory cleanup in order to delete the dataset objects that have
cache files in the temporary directory before deleting the dorectory itself.
"""
def __init__(self, cleanup_func=None, *cleanup_func_args, **cleanup_func_kwargs):
self.name = tempfile.mkdtemp()
self._finalizer = weakref.finalize(self, self._cleanup)
self._cleanup_func = cleanup_func
self._cleanup_func_args = cleanup_func_args
self._cleanup_func_kwargs = cleanup_func_kwargs
def _cleanup(self):
self._cleanup_func(*self._cleanup_func_args, **self._cleanup_func_kwargs)
if os.path.exists(self.name):
shutil.rmtree(self.name)
def cleanup(self):
if self._finalizer.detach():
self._cleanup()
def maybe_register_dataset_for_temp_dir_deletion(dataset):
"""
This function registers the datasets that have cache files in _TEMP_DIR_FOR_TEMP_CACHE_FILES in order
to properly delete them before deleting the temporary directory.
The temporary directory _TEMP_DIR_FOR_TEMP_CACHE_FILES is used when caching is disabled.
"""
if _TEMP_DIR_FOR_TEMP_CACHE_FILES is None:
return
global _DATASETS_WITH_TABLE_IN_TEMP_DIR
if _DATASETS_WITH_TABLE_IN_TEMP_DIR is None:
_DATASETS_WITH_TABLE_IN_TEMP_DIR = weakref.WeakSet()
if any(
Path(_TEMP_DIR_FOR_TEMP_CACHE_FILES.name) in Path(cache_file["filename"]).parents
for cache_file in dataset.cache_files
):
_DATASETS_WITH_TABLE_IN_TEMP_DIR.add(dataset)
def get_datasets_with_cache_file_in_temp_dir():
return list(_DATASETS_WITH_TABLE_IN_TEMP_DIR) if _DATASETS_WITH_TABLE_IN_TEMP_DIR is not None else []
def enable_caching():
"""
When applying transforms on a dataset, the data are stored in cache files.
The caching mechanism allows to reload an existing cache file if it's already been computed.
Reloading a dataset is possible since the cache files are named using the dataset fingerprint, which is updated
after each transform.
If disabled, the library will no longer reload cached datasets files when applying transforms to the datasets.
More precisely, if the caching is disabled:
- cache files are always recreated
- cache files are written to a temporary directory that is deleted when session closes
- cache files are named using a random hash instead of the dataset fingerprint
- use [`~datasets.Dataset.save_to_disk`] to save a transformed dataset or it will be deleted when session closes
- caching doesn't affect [`~datasets.load_dataset`]. If you want to regenerate a dataset from scratch you should use
the `download_mode` parameter in [`~datasets.load_dataset`].
"""
global _CACHING_ENABLED
_CACHING_ENABLED = True
def disable_caching():
"""
When applying transforms on a dataset, the data are stored in cache files.
The caching mechanism allows to reload an existing cache file if it's already been computed.
Reloading a dataset is possible since the cache files are named using the dataset fingerprint, which is updated
after each transform.
If disabled, the library will no longer reload cached datasets files when applying transforms to the datasets.
More precisely, if the caching is disabled:
- cache files are always recreated
- cache files are written to a temporary directory that is deleted when session closes
- cache files are named using a random hash instead of the dataset fingerprint
- use [`~datasets.Dataset.save_to_disk`] to save a transformed dataset or it will be deleted when session closes
- caching doesn't affect [`~datasets.load_dataset`]. If you want to regenerate a dataset from scratch you should use
the `download_mode` parameter in [`~datasets.load_dataset`].
"""
global _CACHING_ENABLED
_CACHING_ENABLED = False
@deprecated(
"Use datasets.enable_caching() or datasets.disable_caching() instead. This function will be removed in a future version of datasets."
)
def set_caching_enabled(boolean: bool):
"""
When applying transforms on a dataset, the data are stored in cache files.
The caching mechanism allows to reload an existing cache file if it's already been computed.
Reloading a dataset is possible since the cache files are named using the dataset fingerprint, which is updated
after each transform.
If disabled, the library will no longer reload cached datasets files when applying transforms to the datasets.
More precisely, if the caching is disabled:
- cache files are always recreated
- cache files are written to a temporary directory that is deleted when session closes
- cache files are named using a random hash instead of the dataset fingerprint
- use :func:`datasets.Dataset.save_to_disk` to save a transformed dataset or it will be deleted when session closes
- caching doesn't affect :func:`datasets.load_dataset`. If you want to regenerate a dataset from scratch you should use
the ``download_mode`` parameter in :func:`datasets.load_dataset`.
"""
global _CACHING_ENABLED
_CACHING_ENABLED = bool(boolean)
def is_caching_enabled() -> bool:
"""
When applying transforms on a dataset, the data are stored in cache files.
The caching mechanism allows to reload an existing cache file if it's already been computed.
Reloading a dataset is possible since the cache files are named using the dataset fingerprint, which is updated
after each transform.
If disabled, the library will no longer reload cached datasets files when applying transforms to the datasets.
More precisely, if the caching is disabled:
- cache files are always recreated
- cache files are written to a temporary directory that is deleted when session closes
- cache files are named using a random hash instead of the dataset fingerprint
- use [`~datasets.Dataset.save_to_disk`]] to save a transformed dataset or it will be deleted when session closes
- caching doesn't affect [`~datasets.load_dataset`]. If you want to regenerate a dataset from scratch you should use
the `download_mode` parameter in [`~datasets.load_dataset`].
"""
global _CACHING_ENABLED
return bool(_CACHING_ENABLED)
def get_temporary_cache_files_directory() -> str:
"""Return a directory that is deleted when session closes."""
global _TEMP_DIR_FOR_TEMP_CACHE_FILES
if _TEMP_DIR_FOR_TEMP_CACHE_FILES is None:
# Avoids a PermissionError on Windows caused by the datasets referencing
# the files from the cache directory on clean-up
def cleanup_func():
for dset in get_datasets_with_cache_file_in_temp_dir():
dset.__del__()
_TEMP_DIR_FOR_TEMP_CACHE_FILES = _TempDirWithCustomCleanup(cleanup_func=cleanup_func)
return _TEMP_DIR_FOR_TEMP_CACHE_FILES.name
#################
# Hashing
#################
@deprecated("Use `copyreg.pickle` to register a custom reducer.")
def hashregister(*types):
def proxy(func):
for t in types:
Hasher.dispatch[t] = func
return func
return proxy
class Hasher:
"""Hasher that accepts python objects as inputs."""
dispatch: Dict = {}
def __init__(self):
self.m = xxhash.xxh64()
@classmethod
def hash_bytes(cls, value: Union[bytes, List[bytes]]) -> str:
value = [value] if isinstance(value, bytes) else value
m = xxhash.xxh64()
for x in value:
m.update(x)
return m.hexdigest()
@classmethod
@deprecated("Use `Hasher.hash` instead.")
def hash_default(cls, value: Any) -> str:
return cls.hash(value)
@classmethod
def hash(cls, value: Any) -> str:
return cls.hash_bytes(dumps(value))
def update(self, value: Any) -> None:
header_for_update = f"=={type(value)}=="
value_for_update = self.hash(value)
self.m.update(header_for_update.encode("utf8"))
self.m.update(value_for_update.encode("utf-8"))
def hexdigest(self) -> str:
return self.m.hexdigest()
#################
# Fingerprinting
#################
# we show a warning only once when fingerprinting fails to avoid spam
fingerprint_warnings: Dict[str, bool] = {}
def generate_fingerprint(dataset) -> str:
state = dataset.__dict__
hasher = Hasher()
for key in sorted(state):
if key == "_fingerprint":
continue
hasher.update(key)
hasher.update(state[key])
# hash data files last modification timestamps as well
for cache_file in dataset.cache_files:
hasher.update(os.path.getmtime(cache_file["filename"]))
return hasher.hexdigest()
def generate_random_fingerprint(nbits=64) -> str:
return f"{random.getrandbits(nbits):0{nbits//4}x}"
def update_fingerprint(fingerprint, transform, transform_args):
global fingerprint_warnings
hasher = Hasher()
hasher.update(fingerprint)
try:
hasher.update(transform)
except: # noqa various errors might raise here from pickle or dill
if _CACHING_ENABLED:
if not fingerprint_warnings.get("update_fingerprint_transform_hash_failed", False):
logger.warning(
f"Transform {transform} couldn't be hashed properly, a random hash was used instead. "
"Make sure your transforms and parameters are serializable with pickle or dill for the dataset fingerprinting and caching to work. "
"If you reuse this transform, the caching mechanism will consider it to be different from the previous calls and recompute everything. "
"This warning is only showed once. Subsequent hashing failures won't be showed."
)
fingerprint_warnings["update_fingerprint_transform_hash_failed"] = True
else:
logger.info(f"Transform {transform} couldn't be hashed properly, a random hash was used instead.")
else:
logger.info(
f"Transform {transform} couldn't be hashed properly, a random hash was used instead. This doesn't affect caching since it's disabled."
)
return generate_random_fingerprint()
for key in sorted(transform_args):
hasher.update(key)
try:
hasher.update(transform_args[key])
except: # noqa various errors might raise here from pickle or dill
if _CACHING_ENABLED:
if not fingerprint_warnings.get("update_fingerprint_transform_hash_failed", False):
logger.warning(
f"Parameter '{key}'={transform_args[key]} of the transform {transform} couldn't be hashed properly, a random hash was used instead. "
"Make sure your transforms and parameters are serializable with pickle or dill for the dataset fingerprinting and caching to work. "
"If you reuse this transform, the caching mechanism will consider it to be different from the previous calls and recompute everything. "
"This warning is only showed once. Subsequent hashing failures won't be showed."
)
fingerprint_warnings["update_fingerprint_transform_hash_failed"] = True
else:
logger.info(
f"Parameter '{key}'={transform_args[key]} of the transform {transform} couldn't be hashed properly, a random hash was used instead."
)
else:
logger.info(
f"Parameter '{key}'={transform_args[key]} of the transform {transform} couldn't be hashed properly, a random hash was used instead. This doesn't affect caching since it's disabled."
)
return generate_random_fingerprint()
return hasher.hexdigest()
def validate_fingerprint(fingerprint: str, max_length=64):
"""
Make sure the fingerprint is a non-empty string that is not longer that max_length=64 by default,
so that the fingerprint can be used to name cache files without issues.
"""
if not isinstance(fingerprint, str) or not fingerprint:
raise ValueError(f"Invalid fingerprint '{fingerprint}': it should be a non-empty string.")
for invalid_char in INVALID_WINDOWS_CHARACTERS_IN_PATH:
if invalid_char in fingerprint:
raise ValueError(
f"Invalid fingerprint. Bad characters from black list '{INVALID_WINDOWS_CHARACTERS_IN_PATH}' found in '{fingerprint}'. "
f"They could create issues when creating cache files."
)
if len(fingerprint) > max_length:
raise ValueError(
f"Invalid fingerprint. Maximum lenth is {max_length} but '{fingerprint}' has length {len(fingerprint)}."
"It could create issues when creating cache files."
)
def format_transform_for_fingerprint(func: Callable, version: Optional[str] = None) -> str:
"""
Format a transform to the format that will be used to update the fingerprint.
"""
transform = f"{func.__module__}.{func.__qualname__}"
if version is not None:
transform += f"@{version}"
return transform
def format_kwargs_for_fingerprint(
func: Callable,
args: Tuple,
kwargs: Dict[str, Any],
use_kwargs: Optional[List[str]] = None,
ignore_kwargs: Optional[List[str]] = None,
randomized_function: bool = False,
) -> Dict[str, Any]:
"""
Format the kwargs of a transform to the format that will be used to update the fingerprint.
"""
kwargs_for_fingerprint = kwargs.copy()
if args:
params = [p.name for p in inspect.signature(func).parameters.values() if p != p.VAR_KEYWORD]
args = args[1:] # assume the first argument is the dataset
params = params[1:]
kwargs_for_fingerprint.update(zip(params, args))
else:
del kwargs_for_fingerprint[
next(iter(inspect.signature(func).parameters))
] # assume the first key is the dataset
# keep the right kwargs to be hashed to generate the fingerprint
if use_kwargs:
kwargs_for_fingerprint = {k: v for k, v in kwargs_for_fingerprint.items() if k in use_kwargs}
if ignore_kwargs:
kwargs_for_fingerprint = {k: v for k, v in kwargs_for_fingerprint.items() if k not in ignore_kwargs}
if randomized_function: # randomized functions have `seed` and `generator` parameters
if kwargs_for_fingerprint.get("seed") is None and kwargs_for_fingerprint.get("generator") is None:
_, seed, pos, *_ = np.random.get_state()
seed = seed[pos] if pos < 624 else seed[0]
kwargs_for_fingerprint["generator"] = np.random.default_rng(seed)
# remove kwargs that are the default values
default_values = {
p.name: p.default for p in inspect.signature(func).parameters.values() if p.default != inspect._empty
}
for default_varname, default_value in default_values.items():
if default_varname in kwargs_for_fingerprint and kwargs_for_fingerprint[default_varname] == default_value:
kwargs_for_fingerprint.pop(default_varname)
return kwargs_for_fingerprint
def fingerprint_transform(
inplace: bool,
use_kwargs: Optional[List[str]] = None,
ignore_kwargs: Optional[List[str]] = None,
fingerprint_names: Optional[List[str]] = None,
randomized_function: bool = False,
version: Optional[str] = None,
):
"""
Wrapper for dataset transforms to update the dataset fingerprint using ``update_fingerprint``
Args:
inplace (:obj:`bool`): If inplace is True, the fingerprint of the dataset is updated inplace.
Otherwise, a parameter "new_fingerprint" is passed to the wrapped method that should take care of
setting the fingerprint of the returned Dataset.
use_kwargs (:obj:`List[str]`, optional): optional white list of argument names to take into account
to update the fingerprint to the wrapped method that should take care of
setting the fingerprint of the returned Dataset. By default all the arguments are used.
ignore_kwargs (:obj:`List[str]`, optional): optional black list of argument names to take into account
to update the fingerprint. Note that ignore_kwargs prevails on use_kwargs.
fingerprint_names (:obj:`List[str]`, optional, defaults to ["new_fingerprint"]):
If the dataset transforms is not inplace and returns a DatasetDict, then it can require
several fingerprints (one per dataset in the DatasetDict). By specifying fingerprint_names,
one fingerprint named after each element of fingerprint_names is going to be passed.
randomized_function (:obj:`bool`, defaults to False): If the dataset transform is random and has
optional parameters "seed" and "generator", then you can set randomized_function to True.
This way, even if users set "seed" and "generator" to None, then the fingerprint is
going to be randomly generated depending on numpy's current state. In this case, the
generator is set to np.random.default_rng(np.random.get_state()[1][0]).
version (:obj:`str`, optional): version of the transform. The version is taken into account when
computing the fingerprint. If a datase transform changes (or at least if the output data
that are cached changes), then one should increase the version. If the version stays the
same, then old cached data could be reused that are not compatible with the new transform.
It should be in the format "MAJOR.MINOR.PATCH".
"""
if use_kwargs is not None and not isinstance(use_kwargs, list):
raise ValueError(f"use_kwargs is supposed to be a list, not {type(use_kwargs)}")
if ignore_kwargs is not None and not isinstance(ignore_kwargs, list):
raise ValueError(f"ignore_kwargs is supposed to be a list, not {type(use_kwargs)}")
if inplace and fingerprint_names:
raise ValueError("fingerprint_names are only used when inplace is False")
fingerprint_names = fingerprint_names if fingerprint_names is not None else ["new_fingerprint"]
def _fingerprint(func):
if not inplace and not all(name in func.__code__.co_varnames for name in fingerprint_names):
raise ValueError(f"function {func} is missing parameters {fingerprint_names} in signature")
if randomized_function: # randomized function have seed and generator parameters
if "seed" not in func.__code__.co_varnames:
raise ValueError(f"'seed' must be in {func}'s signature")
if "generator" not in func.__code__.co_varnames:
raise ValueError(f"'generator' must be in {func}'s signature")
# this call has to be outside the wrapper or since __qualname__ changes in multiprocessing
transform = format_transform_for_fingerprint(func, version=version)
@wraps(func)
def wrapper(*args, **kwargs):
kwargs_for_fingerprint = format_kwargs_for_fingerprint(
func,
args,
kwargs,
use_kwargs=use_kwargs,
ignore_kwargs=ignore_kwargs,
randomized_function=randomized_function,
)
if args:
dataset: Dataset = args[0]
args = args[1:]
else:
dataset: Dataset = kwargs.pop(next(iter(inspect.signature(func).parameters)))
# compute new_fingerprint and add it to the args of not in-place transforms
if inplace:
new_fingerprint = update_fingerprint(dataset._fingerprint, transform, kwargs_for_fingerprint)
else:
for fingerprint_name in fingerprint_names: # transforms like `train_test_split` have several hashes
if kwargs.get(fingerprint_name) is None:
kwargs_for_fingerprint["fingerprint_name"] = fingerprint_name
kwargs[fingerprint_name] = update_fingerprint(
dataset._fingerprint, transform, kwargs_for_fingerprint
)
else:
validate_fingerprint(kwargs[fingerprint_name])
# Call actual function
out = func(dataset, *args, **kwargs)
# Update fingerprint of in-place transforms + update in-place history of transforms
if inplace: # update after calling func so that the fingerprint doesn't change if the function fails
dataset._fingerprint = new_fingerprint
return out
wrapper._decorator_name_ = "fingerprint"
return wrapper
return _fingerprint
| 0
|
hf_public_repos/datasets/src
|
hf_public_repos/datasets/src/datasets/arrow_reader.py
|
# Copyright 2020 The HuggingFace Datasets Authors and the TensorFlow Datasets Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# Lint as: python3
""" Arrow ArrowReader."""
import copy
import math
import os
import re
import shutil
from dataclasses import dataclass
from pathlib import Path
from typing import TYPE_CHECKING, List, Optional, Union
import pyarrow as pa
import pyarrow.parquet as pq
from .download.download_config import DownloadConfig
from .naming import _split_re, filenames_for_dataset_split
from .table import InMemoryTable, MemoryMappedTable, Table, concat_tables
from .utils import logging
from .utils.file_utils import cached_path
if TYPE_CHECKING:
from .info import DatasetInfo # noqa: F401
from .splits import Split, SplitInfo # noqa: F401
logger = logging.get_logger(__name__)
HF_GCP_BASE_URL = "https://storage.googleapis.com/huggingface-nlp/cache/datasets"
_SUB_SPEC_RE = re.compile(
rf"""
^
(?P<split>{_split_re[1:-1]})
(\[
((?P<from>-?\d+)
(?P<from_pct>%)?)?
:
((?P<to>-?\d+)
(?P<to_pct>%)?)?
\])?(\((?P<rounding>[^\)]*)\))?
$
""", # remove ^ and $
re.X,
)
_ADDITION_SEP_RE = re.compile(r"\s*\+\s*")
class DatasetNotOnHfGcsError(ConnectionError):
"""When you can't get the dataset from the Hf google cloud storage"""
pass
class MissingFilesOnHfGcsError(ConnectionError):
"""When some files are missing on the Hf oogle cloud storage"""
pass
@dataclass(frozen=True)
class FileInstructions:
"""The file instructions associated with a split ReadInstruction.
Attributes:
num_examples: `int`, The total number of examples
file_instructions: List[dict(filename, skip, take)], the files information.
The filenames contains the relative path, not absolute.
skip/take indicates which example read in the file: `ds.slice(skip, take)`
"""
num_examples: int
file_instructions: List[dict]
def make_file_instructions(
name: str,
split_infos: List["SplitInfo"],
instruction: Union[str, "ReadInstruction"],
filetype_suffix: Optional[str] = None,
prefix_path: Optional[str] = None,
) -> FileInstructions:
"""Returns instructions of the split dict.
Args:
name (`str`): Name of the dataset.
split_infos (`list` of `[SplitInfo]`): Dataset splits information.
instruction ([`ReadInstruction`] or `str`): Reading instruction for a dataset.
filetype_suffix (`str`, *optional*): Suffix of dataset files, e.g. 'arrow' or 'parquet'.
prefix_path (`str`, *optional*): Prefix of dataset files, e.g. directory name.
Returns:
[`FileInstructions`]
"""
if not isinstance(name, str):
raise TypeError(f"Expected str 'name', but got: {type(name).__name__}")
elif not name:
raise ValueError("Expected non-empty str 'name'")
name2len = {info.name: info.num_examples for info in split_infos}
name2shard_lengths = {info.name: info.shard_lengths for info in split_infos}
name2filenames = {
info.name: filenames_for_dataset_split(
path=prefix_path,
dataset_name=name,
split=info.name,
filetype_suffix=filetype_suffix,
shard_lengths=name2shard_lengths[info.name],
)
for info in split_infos
}
if not isinstance(instruction, ReadInstruction):
instruction = ReadInstruction.from_spec(instruction)
# Create the absolute instruction (per split)
absolute_instructions = instruction.to_absolute(name2len)
# For each split, return the files instruction (skip/take)
file_instructions = []
num_examples = 0
for abs_instr in absolute_instructions:
split_length = name2len[abs_instr.splitname]
filenames = name2filenames[abs_instr.splitname]
shard_lengths = name2shard_lengths[abs_instr.splitname]
from_ = 0 if abs_instr.from_ is None else abs_instr.from_
to = split_length if abs_instr.to is None else abs_instr.to
if shard_lengths is None: # not sharded
for filename in filenames:
num_examples += to - from_
file_instructions.append({"filename": filename, "skip": from_, "take": to - from_})
else: # sharded
index_start = 0 # Beginning (included) of moving window.
index_end = 0 # End (excluded) of moving window.
for filename, shard_length in zip(filenames, shard_lengths):
index_end += shard_length
if from_ < index_end and to > index_start: # There is something to take.
skip = from_ - index_start if from_ > index_start else 0
take = to - index_start - skip if to < index_end else -1
if take == 0:
continue
file_instructions.append({"filename": filename, "skip": skip, "take": take})
num_examples += shard_length - skip if take == -1 else take
index_start += shard_length
return FileInstructions(
num_examples=num_examples,
file_instructions=file_instructions,
)
class BaseReader:
"""
Build a Dataset object out of Instruction instance(s).
"""
def __init__(self, path: str, info: Optional["DatasetInfo"]):
"""Initializes ArrowReader.
Args:
path (str): path where tfrecords are stored.
info (DatasetInfo): info about the dataset.
"""
self._path: str = path
self._info: Optional["DatasetInfo"] = info
self._filetype_suffix: Optional[str] = None
def _get_table_from_filename(self, filename_skip_take, in_memory=False) -> Table:
"""Returns a Dataset instance from given (filename, skip, take)."""
raise NotImplementedError
def _read_files(self, files, in_memory=False) -> Table:
"""Returns Dataset for given file instructions.
Args:
files: List[dict(filename, skip, take)], the files information.
The filenames contain the absolute path, not relative.
skip/take indicates which example read in the file: `ds.slice(skip, take)`
in_memory (bool, default False): Whether to copy the data in-memory.
"""
if len(files) == 0 or not all(isinstance(f, dict) for f in files):
raise ValueError("please provide valid file informations")
pa_tables = []
files = copy.deepcopy(files)
for f in files:
f["filename"] = os.path.join(self._path, f["filename"])
for f_dict in files:
pa_table: Table = self._get_table_from_filename(f_dict, in_memory=in_memory)
pa_tables.append(pa_table)
pa_tables = [t for t in pa_tables if len(t) > 0]
if not pa_tables and (self._info is None or self._info.features is None):
raise ValueError(
"Tried to read an empty table. Please specify at least info.features to create an empty table with the right type."
)
pa_tables = pa_tables or [InMemoryTable.from_batches([], schema=pa.schema(self._info.features.type))]
pa_table = concat_tables(pa_tables) if len(pa_tables) != 1 else pa_tables[0]
return pa_table
def get_file_instructions(self, name, instruction, split_infos):
"""Return list of dict {'filename': str, 'skip': int, 'take': int}"""
file_instructions = make_file_instructions(
name, split_infos, instruction, filetype_suffix=self._filetype_suffix, prefix_path=self._path
)
files = file_instructions.file_instructions
return files
def read(
self,
name,
instructions,
split_infos,
in_memory=False,
):
"""Returns Dataset instance(s).
Args:
name (str): name of the dataset.
instructions (ReadInstruction): instructions to read.
Instruction can be string and will then be passed to the Instruction
constructor as it.
split_infos (list of SplitInfo proto): the available splits for dataset.
in_memory (bool, default False): Whether to copy the data in-memory.
Returns:
kwargs to build a single Dataset instance.
"""
files = self.get_file_instructions(name, instructions, split_infos)
if not files:
msg = f'Instruction "{instructions}" corresponds to no data!'
raise ValueError(msg)
return self.read_files(files=files, original_instructions=instructions, in_memory=in_memory)
def read_files(
self,
files: List[dict],
original_instructions: Union[None, "ReadInstruction", "Split"] = None,
in_memory=False,
):
"""Returns single Dataset instance for the set of file instructions.
Args:
files: List[dict(filename, skip, take)], the files information.
The filenames contains the relative path, not absolute.
skip/take indicates which example read in the file: `ds.skip().take()`
original_instructions: store the original instructions used to build the dataset split in the dataset.
in_memory (bool, default False): Whether to copy the data in-memory.
Returns:
kwargs to build a Dataset instance.
"""
# Prepend path to filename
pa_table = self._read_files(files, in_memory=in_memory)
# If original_instructions is not None, convert it to a human-readable NamedSplit
if original_instructions is not None:
from .splits import Split # noqa
split = Split(str(original_instructions))
else:
split = None
dataset_kwargs = {"arrow_table": pa_table, "info": self._info, "split": split}
return dataset_kwargs
def download_from_hf_gcs(self, download_config: DownloadConfig, relative_data_dir):
"""
Download the dataset files from the Hf GCS
Args:
dl_cache_dir: `str`, the local cache directory used to download files
relative_data_dir: `str`, the relative directory of the remote files from
the `datasets` directory on GCS.
"""
remote_cache_dir = HF_GCP_BASE_URL + "/" + relative_data_dir.replace(os.sep, "/")
try:
remote_dataset_info = os.path.join(remote_cache_dir, "dataset_info.json")
downloaded_dataset_info = cached_path(remote_dataset_info.replace(os.sep, "/"))
shutil.move(downloaded_dataset_info, os.path.join(self._path, "dataset_info.json"))
if self._info is not None:
self._info.update(self._info.from_directory(self._path))
except FileNotFoundError as err:
raise DatasetNotOnHfGcsError(err) from None
try:
for split in self._info.splits:
file_instructions = self.get_file_instructions(
name=self._info.builder_name,
instruction=split,
split_infos=self._info.splits.values(),
)
for file_instruction in file_instructions:
file_to_download = str(Path(file_instruction["filename"]).relative_to(self._path))
remote_prepared_filename = os.path.join(remote_cache_dir, file_to_download)
downloaded_prepared_filename = cached_path(
remote_prepared_filename.replace(os.sep, "/"), download_config=download_config
)
shutil.move(downloaded_prepared_filename, file_instruction["filename"])
except FileNotFoundError as err:
raise MissingFilesOnHfGcsError(err) from None
class ArrowReader(BaseReader):
"""
Build a Dataset object out of Instruction instance(s).
This Reader uses either memory mapping or file descriptors (in-memory) on arrow files.
"""
def __init__(self, path: str, info: Optional["DatasetInfo"]):
"""Initializes ArrowReader.
Args:
path (str): path where Arrow files are stored.
info (DatasetInfo): info about the dataset.
"""
super().__init__(path, info)
self._filetype_suffix = "arrow"
def _get_table_from_filename(self, filename_skip_take, in_memory=False) -> Table:
"""Returns a Dataset instance from given (filename, skip, take)."""
filename, skip, take = (
filename_skip_take["filename"],
filename_skip_take["skip"] if "skip" in filename_skip_take else None,
filename_skip_take["take"] if "take" in filename_skip_take else None,
)
table = ArrowReader.read_table(filename, in_memory=in_memory)
if take == -1:
take = len(table) - skip
# here we don't want to slice an empty table, or it may segfault
if skip is not None and take is not None and not (skip == 0 and take == len(table)):
table = table.slice(skip, take)
return table
@staticmethod
def read_table(filename, in_memory=False) -> Table:
"""
Read table from file.
Args:
filename (str): File name of the table.
in_memory (bool, default=False): Whether to copy the data in-memory.
Returns:
pyarrow.Table
"""
table_cls = InMemoryTable if in_memory else MemoryMappedTable
return table_cls.from_file(filename)
class ParquetReader(BaseReader):
"""
Build a Dataset object out of Instruction instance(s).
This Reader uses memory mapping on parquet files.
"""
def __init__(self, path: str, info: Optional["DatasetInfo"]):
"""Initializes ParquetReader.
Args:
path (str): path where tfrecords are stored.
info (DatasetInfo): info about the dataset.
"""
super().__init__(path, info)
self._filetype_suffix = "parquet"
def _get_table_from_filename(self, filename_skip_take, **kwargs):
"""Returns a Dataset instance from given (filename, skip, take)."""
filename, skip, take = (
filename_skip_take["filename"],
filename_skip_take["skip"] if "skip" in filename_skip_take else None,
filename_skip_take["take"] if "take" in filename_skip_take else None,
)
# Parquet read_table always loads data in memory, independently of memory_map
pa_table = pq.read_table(filename, memory_map=True)
# here we don't want to slice an empty table, or it may segfault
if skip is not None and take is not None and not (skip == 0 and take == len(pa_table)):
pa_table = pa_table.slice(skip, take)
return pa_table
@dataclass(frozen=True)
class _AbsoluteInstruction:
"""A machine friendly slice: defined absolute positive boundaries."""
splitname: str
from_: int # uint (starting index).
to: int # uint (ending index).
@dataclass(frozen=True)
class _RelativeInstruction:
"""Represents a single parsed slicing instruction, can use % and negatives."""
splitname: str
from_: Optional[int] = None # int (starting index) or None if no lower boundary.
to: Optional[int] = None # int (ending index) or None if no upper boundary.
unit: Optional[str] = None
rounding: Optional[str] = None
def __post_init__(self):
if self.unit is not None and self.unit not in ["%", "abs"]:
raise ValueError("unit must be either % or abs")
if self.rounding is not None and self.rounding not in ["closest", "pct1_dropremainder"]:
raise ValueError("rounding must be either closest or pct1_dropremainder")
if self.unit != "%" and self.rounding is not None:
raise ValueError("It is forbidden to specify rounding if not using percent slicing.")
if self.unit == "%" and self.from_ is not None and abs(self.from_) > 100:
raise ValueError("Percent slice boundaries must be > -100 and < 100.")
if self.unit == "%" and self.to is not None and abs(self.to) > 100:
raise ValueError("Percent slice boundaries must be > -100 and < 100.")
# Update via __dict__ due to instance being "frozen"
self.__dict__["rounding"] = "closest" if self.rounding is None and self.unit == "%" else self.rounding
def _str_to_read_instruction(spec):
"""Returns ReadInstruction for given string."""
res = _SUB_SPEC_RE.match(spec)
if not res:
raise ValueError(f"Unrecognized instruction format: {spec}")
unit = "%" if res.group("from_pct") or res.group("to_pct") else "abs"
return ReadInstruction(
split_name=res.group("split"),
rounding=res.group("rounding"),
from_=int(res.group("from")) if res.group("from") else None,
to=int(res.group("to")) if res.group("to") else None,
unit=unit,
)
def _pct_to_abs_pct1(boundary, num_examples):
# Using math.trunc here, since -99.5% should give -99%, not -100%.
if num_examples < 100:
msg = (
'Using "pct1_dropremainder" rounding on a split with less than 100 '
"elements is forbidden: it always results in an empty dataset."
)
raise ValueError(msg)
return boundary * math.trunc(num_examples / 100.0)
def _pct_to_abs_closest(boundary, num_examples):
return int(round(boundary * num_examples / 100.0))
def _rel_to_abs_instr(rel_instr, name2len):
"""Returns _AbsoluteInstruction instance for given RelativeInstruction.
Args:
rel_instr: RelativeInstruction instance.
name2len: dict {split_name: num_examples}.
"""
pct_to_abs = _pct_to_abs_closest if rel_instr.rounding == "closest" else _pct_to_abs_pct1
split = rel_instr.splitname
if split not in name2len:
raise ValueError(f'Unknown split "{split}". Should be one of {list(name2len)}.')
num_examples = name2len[split]
from_ = rel_instr.from_
to = rel_instr.to
if rel_instr.unit == "%":
from_ = 0 if from_ is None else pct_to_abs(from_, num_examples)
to = num_examples if to is None else pct_to_abs(to, num_examples)
else:
from_ = 0 if from_ is None else from_
to = num_examples if to is None else to
if abs(from_) > num_examples or abs(to) > num_examples:
msg = f'Requested slice [{from_ or ""}:{to or ""}] incompatible with {num_examples} examples.'
raise ValueError(msg)
if from_ < 0:
from_ = num_examples + from_
elif from_ == 0:
from_ = None
if to < 0:
to = num_examples + to
elif to == num_examples:
to = None
return _AbsoluteInstruction(split, from_, to)
class ReadInstruction:
"""Reading instruction for a dataset.
Examples::
# The following lines are equivalent:
ds = datasets.load_dataset('mnist', split='test[:33%]')
ds = datasets.load_dataset('mnist', split=datasets.ReadInstruction.from_spec('test[:33%]'))
ds = datasets.load_dataset('mnist', split=datasets.ReadInstruction('test', to=33, unit='%'))
ds = datasets.load_dataset('mnist', split=datasets.ReadInstruction(
'test', from_=0, to=33, unit='%'))
# The following lines are equivalent:
ds = datasets.load_dataset('mnist', split='test[:33%]+train[1:-1]')
ds = datasets.load_dataset('mnist', split=datasets.ReadInstruction.from_spec(
'test[:33%]+train[1:-1]'))
ds = datasets.load_dataset('mnist', split=(
datasets.ReadInstruction('test', to=33, unit='%') +
datasets.ReadInstruction('train', from_=1, to=-1, unit='abs')))
# The following lines are equivalent:
ds = datasets.load_dataset('mnist', split='test[:33%](pct1_dropremainder)')
ds = datasets.load_dataset('mnist', split=datasets.ReadInstruction.from_spec(
'test[:33%](pct1_dropremainder)'))
ds = datasets.load_dataset('mnist', split=datasets.ReadInstruction(
'test', from_=0, to=33, unit='%', rounding="pct1_dropremainder"))
# 10-fold validation:
tests = datasets.load_dataset(
'mnist',
[datasets.ReadInstruction('train', from_=k, to=k+10, unit='%')
for k in range(0, 100, 10)])
trains = datasets.load_dataset(
'mnist',
[datasets.ReadInstruction('train', to=k, unit='%') + datasets.ReadInstruction('train', from_=k+10, unit='%')
for k in range(0, 100, 10)])
"""
def _init(self, relative_instructions):
# Private initializer.
self._relative_instructions = relative_instructions
@classmethod
def _read_instruction_from_relative_instructions(cls, relative_instructions):
"""Returns ReadInstruction obj initialized with relative_instructions."""
# Use __new__ to bypass __init__ used by public API and not conveniant here.
result = cls.__new__(cls)
result._init(relative_instructions) # pylint: disable=protected-access
return result
def __init__(self, split_name, rounding=None, from_=None, to=None, unit=None):
"""Initialize ReadInstruction.
Args:
split_name (str): name of the split to read. Eg: 'train'.
rounding (str, optional): The rounding behaviour to use when percent slicing is
used. Ignored when slicing with absolute indices.
Possible values:
- 'closest' (default): The specified percentages are rounded to the
closest value. Use this if you want specified percents to be as
much exact as possible.
- 'pct1_dropremainder': the specified percentages are treated as
multiple of 1%. Use this option if you want consistency. Eg:
len(5%) == 5 * len(1%).
Using this option, one might not be able to use the full set of
examples, if the number of those is not a multiple of 100.
from_ (int):
to (int): alternative way of specifying slicing boundaries. If any of
{from_, to, unit} argument is used, slicing cannot be specified as
string.
unit (str): optional, one of:
'%': to set the slicing unit as percents of the split size.
'abs': to set the slicing unit as absolute numbers.
"""
# This constructor is not always called. See factory method
# `_read_instruction_from_relative_instructions`. Common init instructions
# MUST be placed in the _init method.
self._init([_RelativeInstruction(split_name, from_, to, unit, rounding)])
@classmethod
def from_spec(cls, spec):
"""Creates a `ReadInstruction` instance out of a string spec.
Args:
spec (`str`):
Split(s) + optional slice(s) to read + optional rounding
if percents are used as the slicing unit. A slice can be specified,
using absolute numbers (`int`) or percentages (`int`).
Examples:
```
test: test split.
test + validation: test split + validation split.
test[10:]: test split, minus its first 10 records.
test[:10%]: first 10% records of test split.
test[:20%](pct1_dropremainder): first 10% records, rounded with the pct1_dropremainder rounding.
test[:-5%]+train[40%:60%]: first 95% of test + middle 20% of train.
```
Returns:
ReadInstruction instance.
"""
spec = str(spec) # Need to convert to str in case of NamedSplit instance.
subs = _ADDITION_SEP_RE.split(spec)
if not subs:
raise ValueError(f"No instructions could be built out of {spec}")
instruction = _str_to_read_instruction(subs[0])
return sum((_str_to_read_instruction(sub) for sub in subs[1:]), instruction)
def to_spec(self):
rel_instr_specs = []
for rel_instr in self._relative_instructions:
rel_instr_spec = rel_instr.splitname
if rel_instr.from_ is not None or rel_instr.to is not None:
from_ = rel_instr.from_
to = rel_instr.to
unit = rel_instr.unit
rounding = rel_instr.rounding
unit = unit if unit == "%" else ""
from_ = str(from_) + unit if from_ is not None else ""
to = str(to) + unit if to is not None else ""
slice_str = f"[{from_}:{to}]"
rounding_str = (
f"({rounding})" if unit == "%" and rounding is not None and rounding != "closest" else ""
)
rel_instr_spec += slice_str + rounding_str
rel_instr_specs.append(rel_instr_spec)
return "+".join(rel_instr_specs)
def __add__(self, other):
"""Returns a new ReadInstruction obj, result of appending other to self."""
if not isinstance(other, ReadInstruction):
msg = "ReadInstruction can only be added to another ReadInstruction obj."
raise TypeError(msg)
self_ris = self._relative_instructions
other_ris = other._relative_instructions # pylint: disable=protected-access
if (
self_ris[0].unit != "abs"
and other_ris[0].unit != "abs"
and self._relative_instructions[0].rounding != other_ris[0].rounding
):
raise ValueError("It is forbidden to sum ReadInstruction instances with different rounding values.")
return self._read_instruction_from_relative_instructions(self_ris + other_ris)
def __str__(self):
return self.to_spec()
def __repr__(self):
return f"ReadInstruction({self._relative_instructions})"
def to_absolute(self, name2len):
"""Translate instruction into a list of absolute instructions.
Those absolute instructions are then to be added together.
Args:
name2len (`dict`):
Associating split names to number of examples.
Returns:
list of _AbsoluteInstruction instances (corresponds to the + in spec).
"""
return [_rel_to_abs_instr(rel_instr, name2len) for rel_instr in self._relative_instructions]
| 0
|
hf_public_repos/datasets/src
|
hf_public_repos/datasets/src/datasets/builder.py
|
# Copyright 2020 The HuggingFace Datasets Authors and the TensorFlow Datasets Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# Lint as: python3
"""DatasetBuilder base class."""
import abc
import contextlib
import copy
import inspect
import os
import posixpath
import shutil
import textwrap
import time
import urllib
import warnings
from dataclasses import dataclass
from functools import partial
from pathlib import Path
from typing import Dict, Iterable, Mapping, Optional, Tuple, Union
import fsspec
import pyarrow as pa
from multiprocess import Pool
from tqdm.contrib.concurrent import thread_map
from . import config, utils
from .arrow_dataset import Dataset
from .arrow_reader import (
HF_GCP_BASE_URL,
ArrowReader,
DatasetNotOnHfGcsError,
MissingFilesOnHfGcsError,
ReadInstruction,
)
from .arrow_writer import ArrowWriter, BeamWriter, ParquetWriter, SchemaInferenceError
from .data_files import DataFilesDict, sanitize_patterns
from .dataset_dict import DatasetDict, IterableDatasetDict
from .download.download_config import DownloadConfig
from .download.download_manager import DownloadManager, DownloadMode
from .download.mock_download_manager import MockDownloadManager
from .download.streaming_download_manager import StreamingDownloadManager, xopen
from .features import Features
from .filesystems import (
is_remote_filesystem,
rename,
)
from .fingerprint import Hasher
from .info import DatasetInfo, DatasetInfosDict, PostProcessedInfo
from .iterable_dataset import ArrowExamplesIterable, ExamplesIterable, IterableDataset
from .keyhash import DuplicatedKeysError
from .naming import INVALID_WINDOWS_CHARACTERS_IN_PATH, camelcase_to_snakecase
from .splits import Split, SplitDict, SplitGenerator, SplitInfo
from .streaming import extend_dataset_builder_for_streaming
from .utils import logging
from .utils import tqdm as hf_tqdm
from .utils._filelock import FileLock
from .utils.file_utils import cached_path, is_remote_url
from .utils.info_utils import VerificationMode, get_size_checksum_dict, verify_checksums, verify_splits
from .utils.py_utils import (
classproperty,
convert_file_size_to_int,
has_sufficient_disk_space,
iflatmap_unordered,
map_nested,
memoize,
size_str,
temporary_assignment,
)
from .utils.sharding import _number_of_shards_in_gen_kwargs, _split_gen_kwargs
logger = logging.get_logger(__name__)
class InvalidConfigName(ValueError):
pass
class DatasetBuildError(Exception):
pass
class ManualDownloadError(DatasetBuildError):
pass
class DatasetGenerationError(DatasetBuildError):
pass
class FileFormatError(DatasetBuildError):
pass
@dataclass
class BuilderConfig:
"""Base class for `DatasetBuilder` data configuration.
`DatasetBuilder` subclasses with data configuration options should subclass
`BuilderConfig` and add their own properties.
Attributes:
name (`str`, defaults to `default`):
The name of the configuration.
version (`Version` or `str`, defaults to `0.0.0`):
The version of the configuration.
data_dir (`str`, *optional*):
Path to the directory containing the source data.
data_files (`str` or `Sequence` or `Mapping`, *optional*):
Path(s) to source data file(s).
description (`str`, *optional*):
A human description of the configuration.
"""
name: str = "default"
version: Optional[Union[utils.Version, str]] = utils.Version("0.0.0")
data_dir: Optional[str] = None
data_files: Optional[DataFilesDict] = None
description: Optional[str] = None
def __post_init__(self):
# The config name is used to name the cache directory.
for invalid_char in INVALID_WINDOWS_CHARACTERS_IN_PATH:
if invalid_char in self.name:
raise InvalidConfigName(
f"Bad characters from black list '{INVALID_WINDOWS_CHARACTERS_IN_PATH}' found in '{self.name}'. "
f"They could create issues when creating a directory for this config on Windows filesystem."
)
if self.data_files is not None and not isinstance(self.data_files, DataFilesDict):
raise ValueError(f"Expected a DataFilesDict in data_files but got {self.data_files}")
def __eq__(self, o):
# we need to override the default dataclass __eq__ since it doesn't check for
# other attributes that the ones of the signature.
if set(self.__dict__.keys()) != set(o.__dict__.keys()):
return False
return all((k, getattr(self, k)) == (k, getattr(o, k)) for k in self.__dict__.keys())
def create_config_id(
self,
config_kwargs: dict,
custom_features: Optional[Features] = None,
) -> str:
"""
The config id is used to build the cache directory.
By default it is equal to the config name.
However the name of a config is not sufficient to have a unique identifier for the dataset being generated
since it doesn't take into account:
- the config kwargs that can be used to overwrite attributes
- the custom features used to write the dataset
- the data_files for json/text/csv/pandas datasets
Therefore the config id is just the config name with an optional suffix based on these.
"""
# Possibly add a suffix to the name to handle custom features/data_files/config_kwargs
suffix: Optional[str] = None
config_kwargs_to_add_to_suffix = config_kwargs.copy()
# name and version are already used to build the cache directory
config_kwargs_to_add_to_suffix.pop("name", None)
config_kwargs_to_add_to_suffix.pop("version", None)
# data dir handling (when specified it points to the manually downloaded data):
# it was previously ignored before the introduction of config id because we didn't want
# to change the config name. Now it's fine to take it into account for the config id.
# config_kwargs_to_add_to_suffix.pop("data_dir", None)
if "data_dir" in config_kwargs_to_add_to_suffix:
if config_kwargs_to_add_to_suffix["data_dir"] is None:
config_kwargs_to_add_to_suffix.pop("data_dir", None)
else:
# canonicalize the data dir to avoid two paths to the same location having different
# hashes
data_dir = config_kwargs_to_add_to_suffix["data_dir"]
data_dir = os.path.normpath(data_dir)
config_kwargs_to_add_to_suffix["data_dir"] = data_dir
if config_kwargs_to_add_to_suffix:
# we don't care about the order of the kwargs
config_kwargs_to_add_to_suffix = {
k: config_kwargs_to_add_to_suffix[k] for k in sorted(config_kwargs_to_add_to_suffix)
}
if all(isinstance(v, (str, bool, int, float)) for v in config_kwargs_to_add_to_suffix.values()):
suffix = ",".join(
str(k) + "=" + urllib.parse.quote_plus(str(v)) for k, v in config_kwargs_to_add_to_suffix.items()
)
if len(suffix) > 32: # hash if too long
suffix = Hasher.hash(config_kwargs_to_add_to_suffix)
else:
suffix = Hasher.hash(config_kwargs_to_add_to_suffix)
if custom_features is not None:
m = Hasher()
if suffix:
m.update(suffix)
m.update(custom_features)
suffix = m.hexdigest()
if suffix:
config_id = self.name + "-" + suffix
if len(config_id) > config.MAX_DATASET_CONFIG_ID_READABLE_LENGTH:
config_id = self.name + "-" + Hasher.hash(suffix)
return config_id
else:
return self.name
class DatasetBuilder:
"""Abstract base class for all datasets.
`DatasetBuilder` has 3 key methods:
- [`DatasetBuilder.info`]: Documents the dataset, including feature
names, types, shapes, version, splits, citation, etc.
- [`DatasetBuilder.download_and_prepare`]: Downloads the source data
and writes it to disk.
- [`DatasetBuilder.as_dataset`]: Generates a [`Dataset`].
Some `DatasetBuilder`s expose multiple variants of the
dataset by defining a [`BuilderConfig`] subclass and accepting a
config object (or name) on construction. Configurable datasets expose a
pre-defined set of configurations in [`DatasetBuilder.builder_configs`].
Args:
cache_dir (`str`, *optional*):
Directory to cache data. Defaults to `"~/.cache/huggingface/datasets"`.
dataset_name (`str`, *optional*):
Name of the dataset, if different from the builder name. Useful for packaged builders
like csv, imagefolder, audiofolder, etc. to reflect the difference between datasets
that use the same packaged builder.
config_name (`str`, *optional*):
Name of the dataset configuration.
It affects the data generated on disk. Different configurations will have their own subdirectories and
versions.
If not provided, the default configuration is used (if it exists).
<Added version="2.3.0">
Parameter `name` was renamed to `config_name`.
</Added>
hash (`str`, *optional*):
Hash specific to the dataset code. Used to update the caching directory when the
dataset loading script code is updated (to avoid reusing old data).
The typical caching directory (defined in `self._relative_data_dir`) is `name/version/hash/`.
base_path (`str`, *optional*):
Base path for relative paths that are used to download files.
This can be a remote URL.
features ([`Features`], *optional*):
Features types to use with this dataset.
It can be used to change the [`Features`] types of a dataset, for example.
token (`str` or `bool`, *optional*):
String or boolean to use as Bearer token for remote files on the
Datasets Hub. If `True`, will get token from `"~/.huggingface"`.
repo_id (`str`, *optional*):
ID of the dataset repository.
Used to distinguish builders with the same name but not coming from the same namespace, for example "squad"
and "lhoestq/squad" repo IDs. In the latter, the builder name would be "lhoestq___squad".
data_files (`str` or `Sequence` or `Mapping`, *optional*):
Path(s) to source data file(s).
For builders like "csv" or "json" that need the user to specify data files. They can be either
local or remote files. For convenience, you can use a `DataFilesDict`.
data_dir (`str`, *optional*):
Path to directory containing source data file(s).
Use only if `data_files` is not passed, in which case it is equivalent to passing
`os.path.join(data_dir, "**")` as `data_files`.
For builders that require manual download, it must be the path to the local directory containing the
manually downloaded data.
storage_options (`dict`, *optional*):
Key/value pairs to be passed on to the dataset file-system backend, if any.
writer_batch_size (`int`, *optional*):
Batch size used by the ArrowWriter.
It defines the number of samples that are kept in memory before writing them
and also the length of the arrow chunks.
None means that the ArrowWriter will use its default value.
name (`str`): Configuration name for the dataset.
<Deprecated version="2.3.0">
Use `config_name` instead.
</Deprecated>
**config_kwargs (additional keyword arguments): Keyword arguments to be passed to the corresponding builder
configuration class, set on the class attribute [`DatasetBuilder.BUILDER_CONFIG_CLASS`]. The builder
configuration class is [`BuilderConfig`] or a subclass of it.
"""
# Default version
VERSION = None # Default version set in BuilderConfig
# Class for the builder config.
BUILDER_CONFIG_CLASS = BuilderConfig
# Named configurations that modify the data generated by download_and_prepare.
BUILDER_CONFIGS = []
# Optional default config name to be used when name is None
DEFAULT_CONFIG_NAME = None
# Default batch size used by the ArrowWriter
# It defines the number of samples that are kept in memory before writing them
# and also the length of the arrow chunks
# None means that the ArrowWriter will use its default value
DEFAULT_WRITER_BATCH_SIZE = None
def __init__(
self,
cache_dir: Optional[str] = None,
dataset_name: Optional[str] = None,
config_name: Optional[str] = None,
hash: Optional[str] = None,
base_path: Optional[str] = None,
info: Optional[DatasetInfo] = None,
features: Optional[Features] = None,
token: Optional[Union[bool, str]] = None,
use_auth_token="deprecated",
repo_id: Optional[str] = None,
data_files: Optional[Union[str, list, dict, DataFilesDict]] = None,
data_dir: Optional[str] = None,
storage_options: Optional[dict] = None,
writer_batch_size: Optional[int] = None,
name="deprecated",
**config_kwargs,
):
if use_auth_token != "deprecated":
warnings.warn(
"'use_auth_token' was deprecated in favor of 'token' in version 2.14.0 and will be removed in 3.0.0.\n"
f"You can remove this warning by passing 'token={use_auth_token}' instead.",
FutureWarning,
)
token = use_auth_token
if name != "deprecated":
warnings.warn(
"Parameter 'name' was renamed to 'config_name' in version 2.3.0 and will be removed in 3.0.0.",
category=FutureWarning,
)
config_name = name
# DatasetBuilder name
self.name: str = camelcase_to_snakecase(self.__module__.split(".")[-1])
self.hash: Optional[str] = hash
self.base_path = base_path
self.token = token
# For backwards compatibility (e.g. if accessed in a dataset script)
self.use_auth_token = token
self.repo_id = repo_id
self.storage_options = storage_options or {}
self.dataset_name = camelcase_to_snakecase(dataset_name) if dataset_name else self.name
self._writer_batch_size = writer_batch_size or self.DEFAULT_WRITER_BATCH_SIZE
if data_files is not None and not isinstance(data_files, DataFilesDict):
data_files = DataFilesDict.from_patterns(
sanitize_patterns(data_files),
base_path=base_path,
download_config=DownloadConfig(token=token, storage_options=self.storage_options),
)
# Prepare config: DatasetConfig contains name, version and description but can be extended by each dataset
if "features" in inspect.signature(self.BUILDER_CONFIG_CLASS.__init__).parameters and features is not None:
config_kwargs["features"] = features
if data_files is not None:
config_kwargs["data_files"] = data_files
if data_dir is not None:
config_kwargs["data_dir"] = data_dir
self.config, self.config_id = self._create_builder_config(
config_name=config_name,
custom_features=features,
**config_kwargs,
)
# prepare info: DatasetInfo are a standardized dataclass across all datasets
# Prefill datasetinfo
if info is None:
# TODO FOR PACKAGED MODULES IT IMPORTS DATA FROM src/packaged_modules which doesn't make sense
info = self.get_exported_dataset_info()
info.update(self._info())
info.builder_name = self.name
info.dataset_name = self.dataset_name
info.config_name = self.config.name
info.version = self.config.version
self.info = info
# update info with user specified infos
if features is not None:
self.info.features = features
# Prepare data dirs:
# cache_dir can be a remote bucket on GCS or S3 (when using BeamBasedBuilder for distributed data processing)
self._cache_dir_root = str(cache_dir or config.HF_DATASETS_CACHE)
self._cache_dir_root = (
self._cache_dir_root if is_remote_url(self._cache_dir_root) else os.path.expanduser(self._cache_dir_root)
)
self._cache_downloaded_dir = (
posixpath.join(self._cache_dir_root, config.DOWNLOADED_DATASETS_DIR)
if cache_dir
else str(config.DOWNLOADED_DATASETS_PATH)
)
self._cache_downloaded_dir = (
self._cache_downloaded_dir
if is_remote_url(self._cache_downloaded_dir)
else os.path.expanduser(self._cache_downloaded_dir)
)
self._cache_dir = self._build_cache_dir()
if not is_remote_url(self._cache_dir_root):
os.makedirs(self._cache_dir_root, exist_ok=True)
lock_path = os.path.join(
self._cache_dir_root, Path(self._cache_dir).as_posix().replace("/", "_") + ".lock"
)
with FileLock(lock_path):
if os.path.exists(self._cache_dir): # check if data exist
if len(os.listdir(self._cache_dir)) > 0:
if os.path.exists(os.path.join(self._cache_dir, config.DATASET_INFO_FILENAME)):
logger.info("Overwrite dataset info from restored data version if exists.")
self.info = DatasetInfo.from_directory(self._cache_dir)
else: # dir exists but no data, remove the empty dir as data aren't available anymore
logger.warning(
f"Old caching folder {self._cache_dir} for dataset {self.dataset_name} exists but no data were found. Removing it. "
)
os.rmdir(self._cache_dir)
# Store in the cache by default unless the user specifies a custom output_dir to download_and_prepare
self._output_dir = self._cache_dir
self._fs: fsspec.AbstractFileSystem = fsspec.filesystem("file")
# Set download manager
self.dl_manager = None
# Set to True by "datasets-cli test" to generate file checksums for (deprecated) dataset_infos.json independently of verification_mode value.
self._record_infos = False
# Set in `.download_and_prepare` once the format of the generated dataset is known
self._file_format = None
# Enable streaming (e.g. it patches "open" to work with remote files)
extend_dataset_builder_for_streaming(self)
def __getstate__(self):
return self.__dict__
def __setstate__(self, d):
self.__dict__ = d
# Re-enable streaming, since patched functions are not kept when pickling
extend_dataset_builder_for_streaming(self)
# Must be set for datasets that use 'data_dir' functionality - the ones
# that require users to do additional steps to download the data
# (this is usually due to some external regulations / rules).
# This field should contain a string with user instructions, including
# the list of files that should be present. It will be
# displayed in the dataset documentation.
@property
def manual_download_instructions(self) -> Optional[str]:
return None
def _has_legacy_cache(self) -> bool:
"""Check for the old cache directory template {cache_dir}/{namespace}___{builder_name}"""
if (
self.__module__.startswith("datasets.")
and not is_remote_url(self._cache_dir_root)
and self.config.name == "default"
):
namespace = self.repo_id.split("/")[0] if self.repo_id and self.repo_id.count("/") > 0 else None
legacy_config_name = self.repo_id.replace("/", "--") if self.repo_id is not None else self.dataset_name
legacy_config_id = legacy_config_name + self.config_id[len(self.config.name) :]
legacy_cache_dir = os.path.join(
self._cache_dir_root,
self.name if namespace is None else f"{namespace}___{self.name}",
legacy_config_id,
)
return os.path.isdir(legacy_cache_dir)
return False
@classmethod
def get_all_exported_dataset_infos(cls) -> DatasetInfosDict:
"""Empty dict if doesn't exist
Example:
```py
>>> from datasets import load_dataset_builder
>>> ds_builder = load_dataset_builder('rotten_tomatoes')
>>> ds_builder.get_all_exported_dataset_infos()
{'default': DatasetInfo(description="Movie Review Dataset.\nThis is a dataset of containing 5,331 positive and 5,331 negative processed\nsentences from Rotten Tomatoes movie reviews. This data was first used in Bo\nPang and Lillian Lee, ``Seeing stars: Exploiting class relationships for\nsentiment categorization with respect to rating scales.'', Proceedings of the\nACL, 2005.\n", citation='@InProceedings{Pang+Lee:05a,\n author = {Bo Pang and Lillian Lee},\n title = {Seeing stars: Exploiting class relationships for sentiment\n categorization with respect to rating scales},\n booktitle = {Proceedings of the ACL},\n year = 2005\n}\n', homepage='http://www.cs.cornell.edu/people/pabo/movie-review-data/', license='', features={'text': Value(dtype='string', id=None), 'label': ClassLabel(num_classes=2, names=['neg', 'pos'], id=None)}, post_processed=None, supervised_keys=SupervisedKeysData(input='', output=''), task_templates=[TextClassification(task='text-classification', text_column='text', label_column='label')], builder_name='rotten_tomatoes_movie_review', config_name='default', version=1.0.0, splits={'train': SplitInfo(name='train', num_bytes=1074810, num_examples=8530, dataset_name='rotten_tomatoes_movie_review'), 'validation': SplitInfo(name='validation', num_bytes=134679, num_examples=1066, dataset_name='rotten_tomatoes_movie_review'), 'test': SplitInfo(name='test', num_bytes=135972, num_examples=1066, dataset_name='rotten_tomatoes_movie_review')}, download_checksums={'https://storage.googleapis.com/seldon-datasets/sentence_polarity_v1/rt-polaritydata.tar.gz': {'num_bytes': 487770, 'checksum': 'a05befe52aafda71d458d188a1c54506a998b1308613ba76bbda2e5029409ce9'}}, download_size=487770, post_processing_size=None, dataset_size=1345461, size_in_bytes=1833231)}
```
"""
return DatasetInfosDict.from_directory(cls.get_imported_module_dir())
def get_exported_dataset_info(self) -> DatasetInfo:
"""Empty `DatasetInfo` if doesn't exist
Example:
```py
>>> from datasets import load_dataset_builder
>>> ds_builder = load_dataset_builder('rotten_tomatoes')
>>> ds_builder.get_exported_dataset_info()
DatasetInfo(description="Movie Review Dataset.\nThis is a dataset of containing 5,331 positive and 5,331 negative processed\nsentences from Rotten Tomatoes movie reviews. This data was first used in Bo\nPang and Lillian Lee, ``Seeing stars: Exploiting class relationships for\nsentiment categorization with respect to rating scales.'', Proceedings of the\nACL, 2005.\n", citation='@InProceedings{Pang+Lee:05a,\n author = {Bo Pang and Lillian Lee},\n title = {Seeing stars: Exploiting class relationships for sentiment\n categorization with respect to rating scales},\n booktitle = {Proceedings of the ACL},\n year = 2005\n}\n', homepage='http://www.cs.cornell.edu/people/pabo/movie-review-data/', license='', features={'text': Value(dtype='string', id=None), 'label': ClassLabel(num_classes=2, names=['neg', 'pos'], id=None)}, post_processed=None, supervised_keys=SupervisedKeysData(input='', output=''), task_templates=[TextClassification(task='text-classification', text_column='text', label_column='label')], builder_name='rotten_tomatoes_movie_review', config_name='default', version=1.0.0, splits={'train': SplitInfo(name='train', num_bytes=1074810, num_examples=8530, dataset_name='rotten_tomatoes_movie_review'), 'validation': SplitInfo(name='validation', num_bytes=134679, num_examples=1066, dataset_name='rotten_tomatoes_movie_review'), 'test': SplitInfo(name='test', num_bytes=135972, num_examples=1066, dataset_name='rotten_tomatoes_movie_review')}, download_checksums={'https://storage.googleapis.com/seldon-datasets/sentence_polarity_v1/rt-polaritydata.tar.gz': {'num_bytes': 487770, 'checksum': 'a05befe52aafda71d458d188a1c54506a998b1308613ba76bbda2e5029409ce9'}}, download_size=487770, post_processing_size=None, dataset_size=1345461, size_in_bytes=1833231)
```
"""
return self.get_all_exported_dataset_infos().get(self.config.name, DatasetInfo())
def _create_builder_config(
self, config_name=None, custom_features=None, **config_kwargs
) -> Tuple[BuilderConfig, str]:
"""Create and validate BuilderConfig object as well as a unique config id for this config.
Raises ValueError if there are multiple builder configs and config_name and DEFAULT_CONFIG_NAME are None.
config_kwargs override the defaults kwargs in config
"""
builder_config = None
# try default config
if config_name is None and self.BUILDER_CONFIGS and not config_kwargs:
if self.DEFAULT_CONFIG_NAME is not None:
builder_config = self.builder_configs.get(self.DEFAULT_CONFIG_NAME)
logger.info(f"No config specified, defaulting to: {self.dataset_name}/{builder_config.name}")
else:
if len(self.BUILDER_CONFIGS) > 1:
example_of_usage = f"load_dataset('{self.dataset_name}', '{self.BUILDER_CONFIGS[0].name}')"
raise ValueError(
"Config name is missing."
f"\nPlease pick one among the available configs: {list(self.builder_configs.keys())}"
+ f"\nExample of usage:\n\t`{example_of_usage}`"
)
builder_config = self.BUILDER_CONFIGS[0]
logger.info(
f"No config specified, defaulting to the single config: {self.dataset_name}/{builder_config.name}"
)
# try to get config by name
if isinstance(config_name, str):
builder_config = self.builder_configs.get(config_name)
if builder_config is None and self.BUILDER_CONFIGS:
raise ValueError(
f"BuilderConfig '{config_name}' not found. Available: {list(self.builder_configs.keys())}"
)
# if not using an existing config, then create a new config on the fly
if not builder_config:
if config_name is not None:
config_kwargs["name"] = config_name
elif self.DEFAULT_CONFIG_NAME and not config_kwargs:
# Use DEFAULT_CONFIG_NAME only if no config_kwargs are passed
config_kwargs["name"] = self.DEFAULT_CONFIG_NAME
if "version" not in config_kwargs and hasattr(self, "VERSION") and self.VERSION:
config_kwargs["version"] = self.VERSION
builder_config = self.BUILDER_CONFIG_CLASS(**config_kwargs)
# otherwise use the config_kwargs to overwrite the attributes
else:
builder_config = copy.deepcopy(builder_config)
for key, value in config_kwargs.items():
if value is not None:
if not hasattr(builder_config, key):
raise ValueError(f"BuilderConfig {builder_config} doesn't have a '{key}' key.")
setattr(builder_config, key, value)
if not builder_config.name:
raise ValueError(f"BuilderConfig must have a name, got {builder_config.name}")
# compute the config id that is going to be used for caching
config_id = builder_config.create_config_id(
config_kwargs,
custom_features=custom_features,
)
is_custom = (config_id not in self.builder_configs) and config_id != "default"
if is_custom:
logger.info(f"Using custom data configuration {config_id}")
else:
if (
builder_config.name in self.builder_configs
and builder_config != self.builder_configs[builder_config.name]
):
raise ValueError(
"Cannot name a custom BuilderConfig the same as an available "
f"BuilderConfig. Change the name. Available BuilderConfigs: {list(self.builder_configs.keys())}"
)
if not builder_config.version:
raise ValueError(f"BuilderConfig {builder_config.name} must have a version")
return builder_config, config_id
@classproperty
@classmethod
@memoize()
def builder_configs(cls):
"""Dictionary of pre-defined configurations for this builder class."""
configs = {config.name: config for config in cls.BUILDER_CONFIGS}
if len(configs) != len(cls.BUILDER_CONFIGS):
names = [config.name for config in cls.BUILDER_CONFIGS]
raise ValueError(f"Names in BUILDER_CONFIGS must not be duplicated. Got {names}")
return configs
@property
def cache_dir(self):
return self._cache_dir
def _relative_data_dir(self, with_version=True, with_hash=True) -> str:
"""Relative path of this dataset in cache_dir:
Will be:
self.dataset_name/self.config.version/self.hash/
or if a repo_id with a namespace has been specified:
self.namespace___self.dataset_name/self.config.version/self.hash/
If any of these element is missing or if ``with_version=False`` the corresponding subfolders are dropped.
"""
# Check for the legacy cache directory template (datasets<3.0.0)
if self._has_legacy_cache():
# use legacy names
dataset_name = self.name
config_name = self.repo_id.replace("/", "--") if self.repo_id is not None else self.dataset_name
config_id = config_name + self.config_id[len(self.config.name) :]
else:
dataset_name = self.dataset_name
config_name = self.config.name
config_id = self.config_id
namespace = self.repo_id.split("/")[0] if self.repo_id and self.repo_id.count("/") > 0 else None
builder_data_dir = dataset_name if namespace is None else f"{namespace}___{dataset_name}"
builder_data_dir = posixpath.join(builder_data_dir, config_id)
if with_version:
builder_data_dir = posixpath.join(builder_data_dir, str(self.config.version))
if with_hash and self.hash and isinstance(self.hash, str):
builder_data_dir = posixpath.join(builder_data_dir, self.hash)
return builder_data_dir
def _build_cache_dir(self):
"""Return the data directory for the current version."""
builder_data_dir = posixpath.join(self._cache_dir_root, self._relative_data_dir(with_version=False))
version_data_dir = posixpath.join(self._cache_dir_root, self._relative_data_dir(with_version=True))
def _other_versions_on_disk():
"""Returns previous versions on disk."""
if not os.path.exists(builder_data_dir):
return []
version_dirnames = []
for dir_name in os.listdir(builder_data_dir):
try:
version_dirnames.append((utils.Version(dir_name), dir_name))
except ValueError: # Invalid version (ex: incomplete data dir)
pass
version_dirnames.sort(reverse=True)
return version_dirnames
# Check and warn if other versions exist
if not is_remote_url(builder_data_dir):
version_dirs = _other_versions_on_disk()
if version_dirs:
other_version = version_dirs[0][0]
if other_version != self.config.version:
warn_msg = (
f"Found a different version {str(other_version)} of dataset {self.dataset_name} in "
f"cache_dir {self._cache_dir_root}. Using currently defined version "
f"{str(self.config.version)}."
)
logger.warning(warn_msg)
return version_data_dir
@abc.abstractmethod
def _info(self) -> DatasetInfo:
"""Construct the DatasetInfo object. See `DatasetInfo` for details.
Warning: This function is only called once and the result is cached for all
following .info() calls.
Returns:
info: (DatasetInfo) The dataset information
"""
raise NotImplementedError
@classmethod
def get_imported_module_dir(cls):
"""Return the path of the module of this class or subclass."""
return os.path.dirname(inspect.getfile(inspect.getmodule(cls)))
def _rename(self, src: str, dst: str):
rename(self._fs, src, dst)
def download_and_prepare(
self,
output_dir: Optional[str] = None,
download_config: Optional[DownloadConfig] = None,
download_mode: Optional[Union[DownloadMode, str]] = None,
verification_mode: Optional[Union[VerificationMode, str]] = None,
ignore_verifications="deprecated",
try_from_hf_gcs: bool = True,
dl_manager: Optional[DownloadManager] = None,
base_path: Optional[str] = None,
use_auth_token="deprecated",
file_format: str = "arrow",
max_shard_size: Optional[Union[int, str]] = None,
num_proc: Optional[int] = None,
storage_options: Optional[dict] = None,
**download_and_prepare_kwargs,
):
"""Downloads and prepares dataset for reading.
Args:
output_dir (`str`, *optional*):
Output directory for the dataset.
Default to this builder's `cache_dir`, which is inside `~/.cache/huggingface/datasets` by default.
<Added version="2.5.0"/>
download_config (`DownloadConfig`, *optional*):
Specific download configuration parameters.
download_mode ([`DownloadMode`] or `str`, *optional*):
Select the download/generate mode, default to `REUSE_DATASET_IF_EXISTS`.
verification_mode ([`VerificationMode`] or `str`, defaults to `BASIC_CHECKS`):
Verification mode determining the checks to run on the downloaded/processed dataset information (checksums/size/splits/...).
<Added version="2.9.1"/>
ignore_verifications (`bool`, defaults to `False`):
Ignore the verifications of the downloaded/processed dataset information (checksums/size/splits/...).
<Deprecated version="2.9.1">
`ignore_verifications` was deprecated in version 2.9.1 and will be removed in 3.0.0.
Please use `verification_mode` instead.
</Deprecated>
try_from_hf_gcs (`bool`):
If `True`, it will try to download the already prepared dataset from the HF Google cloud storage.
dl_manager (`DownloadManager`, *optional*):
Specific `DownloadManger` to use.
base_path (`str`, *optional*):
Base path for relative paths that are used to download files. This can be a remote url.
If not specified, the value of the `base_path` attribute (`self.base_path`) will be used instead.
use_auth_token (`Union[str, bool]`, *optional*):
Optional string or boolean to use as Bearer token for remote files on the Datasets Hub.
If True, or not specified, will get token from ~/.huggingface.
<Deprecated version="2.7.1">
Pass `use_auth_token` to `load_dataset_builder` instead.
</Deprecated>
file_format (`str`, *optional*):
Format of the data files in which the dataset will be written.
Supported formats: "arrow", "parquet". Default to "arrow" format.
If the format is "parquet", then image and audio data are embedded into the Parquet files instead of pointing to local files.
<Added version="2.5.0"/>
max_shard_size (`Union[str, int]`, *optional*):
Maximum number of bytes written per shard, default is "500MB".
The size is based on uncompressed data size, so in practice your shard files may be smaller than
`max_shard_size` thanks to Parquet compression for example.
<Added version="2.5.0"/>
num_proc (`int`, *optional*, defaults to `None`):
Number of processes when downloading and generating the dataset locally.
Multiprocessing is disabled by default.
<Added version="2.7.0"/>
storage_options (`dict`, *optional*):
Key/value pairs to be passed on to the caching file-system backend, if any.
<Added version="2.5.0"/>
**download_and_prepare_kwargs (additional keyword arguments): Keyword arguments.
Example:
Download and prepare the dataset as Arrow files that can be loaded as a Dataset using `builder.as_dataset()`:
```py
>>> from datasets import load_dataset_builder
>>> builder = load_dataset_builder("rotten_tomatoes")
>>> builder.download_and_prepare()
```
Download and prepare the dataset as sharded Parquet files locally:
```py
>>> from datasets import load_dataset_builder
>>> builder = load_dataset_builder("rotten_tomatoes")
>>> builder.download_and_prepare("./output_dir", file_format="parquet")
```
Download and prepare the dataset as sharded Parquet files in a cloud storage:
```py
>>> from datasets import load_dataset_builder
>>> storage_options = {"key": aws_access_key_id, "secret": aws_secret_access_key}
>>> builder = load_dataset_builder("rotten_tomatoes")
>>> builder.download_and_prepare("s3://my-bucket/my_rotten_tomatoes", storage_options=storage_options, file_format="parquet")
```
"""
if ignore_verifications != "deprecated":
verification_mode = VerificationMode.NO_CHECKS if ignore_verifications else VerificationMode.ALL_CHECKS
warnings.warn(
"'ignore_verifications' was deprecated in favor of 'verification_mode' in version 2.9.1 and will be removed in 3.0.0.\n"
f"You can remove this warning by passing 'verification_mode={verification_mode.value}' instead.",
FutureWarning,
)
if use_auth_token != "deprecated":
warnings.warn(
"'use_auth_token' was deprecated in version 2.7.1 and will be removed in 3.0.0. Pass `token` to `load_dataset_builder` instead.",
FutureWarning,
)
token = use_auth_token
else:
token = self.token
output_dir = output_dir if output_dir is not None else self._cache_dir
# output_dir can be a remote bucket on GCS or S3 (when using BeamBasedBuilder for distributed data processing)
fs, _, [output_dir] = fsspec.get_fs_token_paths(output_dir, storage_options=storage_options)
self._fs = fs
self._output_dir = output_dir if not is_remote_filesystem(self._fs) else self._fs.unstrip_protocol(output_dir)
download_mode = DownloadMode(download_mode or DownloadMode.REUSE_DATASET_IF_EXISTS)
verification_mode = VerificationMode(verification_mode or VerificationMode.BASIC_CHECKS)
base_path = base_path if base_path is not None else self.base_path
if file_format is not None and file_format not in ["arrow", "parquet"]:
raise ValueError(f"Unsupported file_format: {file_format}. Expected 'arrow' or 'parquet'")
self._file_format = file_format
if self._fs._strip_protocol(self._output_dir) == "":
# We don't support the root directory, because it has no dirname,
# and we need a dirname to use a <dirname>.incomplete directory
# when the dataset is being written
raise RuntimeError(
f"Unable to download and prepare the dataset at the root {self._output_dir}. "
f"Please specify a subdirectory, e.g. '{self._output_dir + self.dataset_name}'"
)
if dl_manager is None:
if download_config is None:
download_config = DownloadConfig(
cache_dir=self._cache_downloaded_dir,
force_download=download_mode == DownloadMode.FORCE_REDOWNLOAD,
force_extract=download_mode == DownloadMode.FORCE_REDOWNLOAD,
use_etag=False,
num_proc=num_proc,
token=token,
storage_options=self.storage_options,
) # We don't use etag for data files to speed up the process
dl_manager = DownloadManager(
dataset_name=self.dataset_name,
download_config=download_config,
data_dir=self.config.data_dir,
base_path=base_path,
record_checksums=(self._record_infos or verification_mode == VerificationMode.ALL_CHECKS),
)
is_local = not is_remote_filesystem(self._fs)
if (
isinstance(dl_manager, MockDownloadManager)
or not is_local
or file_format != "arrow"
or max_shard_size is not None
):
try_from_hf_gcs = False
self.dl_manager = dl_manager
# Prevent parallel local disk operations
if is_local:
# Create parent directory of the output_dir to put the lock file in there
Path(self._output_dir).parent.mkdir(parents=True, exist_ok=True)
lock_path = self._output_dir + "_builder.lock"
# File locking only with local paths; no file locking on GCS or S3
with FileLock(lock_path) if is_local else contextlib.nullcontext():
# Check if the data already exists
data_exists = self._fs.exists(posixpath.join(self._output_dir, config.DATASET_INFO_FILENAME))
if data_exists and download_mode == DownloadMode.REUSE_DATASET_IF_EXISTS:
logger.info(f"Found cached dataset {self.dataset_name} ({self._output_dir})")
# We need to update the info in case some splits were added in the meantime
# for example when calling load_dataset from multiple workers.
self.info = self._load_info()
self.download_post_processing_resources(dl_manager)
return
logger.info(f"Generating dataset {self.dataset_name} ({self._output_dir})")
if is_local: # if cache dir is local, check for available space
if not has_sufficient_disk_space(
self.info.size_in_bytes or 0, directory=Path(self._output_dir).parent
):
raise OSError(
f"Not enough disk space. Needed: {size_str(self.info.size_in_bytes or 0)} (download: {size_str(self.info.download_size or 0)}, generated: {size_str(self.info.dataset_size or 0)}, post-processed: {size_str(self.info.post_processing_size or 0)})"
)
@contextlib.contextmanager
def incomplete_dir(dirname):
"""Create temporary dir for dirname and rename on exit."""
if not is_local:
self._fs.makedirs(dirname, exist_ok=True)
yield dirname
else:
tmp_dir = dirname + ".incomplete"
os.makedirs(tmp_dir, exist_ok=True)
try:
yield tmp_dir
if os.path.isdir(dirname):
shutil.rmtree(dirname)
# LocalFileSystem.mv does copy + rm, it is more efficient to simply rename a local directory
shutil.move(tmp_dir, dirname)
finally:
if os.path.exists(tmp_dir):
shutil.rmtree(tmp_dir)
# Print is intentional: we want this to always go to stdout so user has
# information needed to cancel download/preparation if needed.
# This comes right before the progress bar.
if self.info.size_in_bytes:
logger.info(
f"Downloading and preparing dataset {self.dataset_name}/{self.config.name} "
f"(download: {size_str(self.info.download_size)}, generated: {size_str(self.info.dataset_size)}, "
f"post-processed: {size_str(self.info.post_processing_size)}, "
f"total: {size_str(self.info.size_in_bytes)}) to {self._output_dir}..."
)
else:
_dest = self._fs._strip_protocol(self._output_dir) if is_local else self._output_dir
logger.info(f"Downloading and preparing dataset {self.dataset_name}/{self.config.name} to {_dest}...")
self._check_manual_download(dl_manager)
# Create a tmp dir and rename to self._output_dir on successful exit.
with incomplete_dir(self._output_dir) as tmp_output_dir:
# Temporarily assign _output_dir to tmp_data_dir to avoid having to forward
# it to every sub function.
with temporary_assignment(self, "_output_dir", tmp_output_dir):
# Try to download the already prepared dataset files
downloaded_from_gcs = False
if try_from_hf_gcs:
try:
self._download_prepared_from_hf_gcs(dl_manager.download_config)
downloaded_from_gcs = True
except (DatasetNotOnHfGcsError, MissingFilesOnHfGcsError):
logger.info("Dataset not on Hf google storage. Downloading and preparing it from source")
except ConnectionError:
logger.warning("HF google storage unreachable. Downloading and preparing it from source")
if not downloaded_from_gcs:
prepare_split_kwargs = {"file_format": file_format}
if max_shard_size is not None:
prepare_split_kwargs["max_shard_size"] = max_shard_size
if num_proc is not None:
prepare_split_kwargs["num_proc"] = num_proc
self._download_and_prepare(
dl_manager=dl_manager,
verification_mode=verification_mode,
**prepare_split_kwargs,
**download_and_prepare_kwargs,
)
# Sync info
self.info.dataset_size = sum(split.num_bytes for split in self.info.splits.values())
self.info.download_checksums = dl_manager.get_recorded_sizes_checksums()
self.info.size_in_bytes = self.info.dataset_size + self.info.download_size
# Save info
self._save_info()
# Download post processing resources
self.download_post_processing_resources(dl_manager)
logger.info(
f"Dataset {self.dataset_name} downloaded and prepared to {self._output_dir}. "
f"Subsequent calls will reuse this data."
)
def _check_manual_download(self, dl_manager):
if self.manual_download_instructions is not None and dl_manager.manual_dir is None:
raise ManualDownloadError(
textwrap.dedent(
f"""\
The dataset {self.dataset_name} with config {self.config.name} requires manual data.
Please follow the manual download instructions:
{self.manual_download_instructions}
Manual data can be loaded with:
datasets.load_dataset("{self.dataset_name}", data_dir="<path/to/manual/data>")"""
)
)
def _download_prepared_from_hf_gcs(self, download_config: DownloadConfig):
relative_data_dir = self._relative_data_dir(with_version=True, with_hash=False)
reader = ArrowReader(self._output_dir, self.info)
# use reader instructions to download the right files
reader.download_from_hf_gcs(download_config, relative_data_dir)
downloaded_info = DatasetInfo.from_directory(self._output_dir)
self.info.update(downloaded_info)
# download post processing resources
remote_cache_dir = HF_GCP_BASE_URL + "/" + relative_data_dir.replace(os.sep, "/")
for split in self.info.splits:
for resource_file_name in self._post_processing_resources(split).values():
if os.sep in resource_file_name:
raise ValueError(f"Resources shouldn't be in a sub-directory: {resource_file_name}")
try:
resource_path = cached_path(remote_cache_dir + "/" + resource_file_name)
shutil.move(resource_path, os.path.join(self._output_dir, resource_file_name))
except ConnectionError:
logger.info(f"Couldn't download resourse file {resource_file_name} from Hf google storage.")
logger.info("Dataset downloaded from Hf google storage.")
def _download_and_prepare(self, dl_manager, verification_mode, **prepare_split_kwargs):
"""Downloads and prepares dataset for reading.
This is the internal implementation to overwrite called when user calls
`download_and_prepare`. It should download all required data and generate
the pre-processed datasets files.
Args:
dl_manager ([`DownloadManager`]):
`DownloadManager` used to download and cache data.
verification_mode ([`VerificationMode`]):
if `ALL_CHECKS`, perform all the verifications including checksums.
if `BASIC_CHECKS`, do not perform checksums, only perform split tests.
if `NO_CHECKS`, do not perform any verification.
prepare_split_kwargs: Additional options, such as `file_format`, `max_shard_size`
"""
# Generating data for all splits
split_dict = SplitDict(dataset_name=self.dataset_name)
split_generators_kwargs = self._make_split_generators_kwargs(prepare_split_kwargs)
split_generators = self._split_generators(dl_manager, **split_generators_kwargs)
# Checksums verification
if verification_mode == VerificationMode.ALL_CHECKS and dl_manager.record_checksums:
verify_checksums(
self.info.download_checksums, dl_manager.get_recorded_sizes_checksums(), "dataset source files"
)
# Build splits
for split_generator in split_generators:
if str(split_generator.split_info.name).lower() == "all":
raise ValueError(
"`all` is a special split keyword corresponding to the "
"union of all splits, so cannot be used as key in "
"._split_generator()."
)
logger.info(f"Generating {split_generator.split_info.name} split")
split_dict.add(split_generator.split_info)
try:
# Prepare split will record examples associated to the split
self._prepare_split(split_generator, **prepare_split_kwargs)
except OSError as e:
raise OSError(
"Cannot find data file. "
+ (self.manual_download_instructions or "")
+ "\nOriginal error:\n"
+ str(e)
) from None
# If check_duplicates is set to True , then except DuplicatedKeysError
except DuplicatedKeysError as e:
raise DuplicatedKeysError(
e.key,
e.duplicate_key_indices,
fix_msg=f"To avoid duplicate keys, please fix the dataset script {self.name}.py",
) from None
dl_manager.manage_extracted_files()
if verification_mode == VerificationMode.BASIC_CHECKS or verification_mode == VerificationMode.ALL_CHECKS:
verify_splits(self.info.splits, split_dict)
# Update the info object with the splits.
self.info.splits = split_dict
self.info.download_size = dl_manager.downloaded_size
def download_post_processing_resources(self, dl_manager):
for split in self.info.splits or []:
for resource_name, resource_file_name in self._post_processing_resources(split).items():
if not not is_remote_filesystem(self._fs):
raise NotImplementedError(f"Post processing is not supported on filesystem {self._fs}")
if os.sep in resource_file_name:
raise ValueError(f"Resources shouldn't be in a sub-directory: {resource_file_name}")
resource_path = os.path.join(self._output_dir, resource_file_name)
if not os.path.exists(resource_path):
downloaded_resource_path = self._download_post_processing_resources(
split, resource_name, dl_manager
)
if downloaded_resource_path:
logger.info(f"Downloaded post-processing resource {resource_name} as {resource_file_name}")
shutil.move(downloaded_resource_path, resource_path)
def _load_info(self) -> DatasetInfo:
return DatasetInfo.from_directory(self._output_dir, storage_options=self._fs.storage_options)
def _save_info(self):
file_lock = (
FileLock(self._output_dir + "_info.lock")
if not is_remote_filesystem(self._fs)
else contextlib.nullcontext()
)
with file_lock:
self.info.write_to_directory(self._output_dir, storage_options=self._fs.storage_options)
def _save_infos(self):
file_lock = (
FileLock(self._output_dir + "_infos.lock")
if not is_remote_filesystem(self._fs)
else contextlib.nullcontext()
)
with file_lock:
DatasetInfosDict(**{self.config.name: self.info}).write_to_directory(self.get_imported_module_dir())
def _make_split_generators_kwargs(self, prepare_split_kwargs):
"""Get kwargs for `self._split_generators()` from `prepare_split_kwargs`."""
del prepare_split_kwargs
return {}
def as_dataset(
self,
split: Optional[Split] = None,
run_post_process=True,
verification_mode: Optional[Union[VerificationMode, str]] = None,
ignore_verifications="deprecated",
in_memory=False,
) -> Union[Dataset, DatasetDict]:
"""Return a Dataset for the specified split.
Args:
split (`datasets.Split`):
Which subset of the data to return.
run_post_process (`bool`, defaults to `True`):
Whether to run post-processing dataset transforms and/or add
indexes.
verification_mode ([`VerificationMode`] or `str`, defaults to `BASIC_CHECKS`):
Verification mode determining the checks to run on the
downloaded/processed dataset information (checksums/size/splits/...).
<Added version="2.9.1"/>
ignore_verifications (`bool`, defaults to `False`):
Whether to ignore the verifications of the
downloaded/processed dataset information (checksums/size/splits/...).
<Deprecated version="2.9.1">
`ignore_verifications` was deprecated in version 2.9.1 and will be removed in 3.0.0.
Please use `verification_mode` instead.
</Deprecated>
in_memory (`bool`, defaults to `False`):
Whether to copy the data in-memory.
Returns:
datasets.Dataset
Example:
```py
>>> from datasets import load_dataset_builder
>>> builder = load_dataset_builder('rotten_tomatoes')
>>> builder.download_and_prepare()
>>> ds = builder.as_dataset(split='train')
>>> ds
Dataset({
features: ['text', 'label'],
num_rows: 8530
})
```
"""
if ignore_verifications != "deprecated":
verification_mode = verification_mode.NO_CHECKS if ignore_verifications else VerificationMode.ALL_CHECKS
warnings.warn(
"'ignore_verifications' was deprecated in favor of 'verification' in version 2.9.1 and will be removed in 3.0.0.\n"
f"You can remove this warning by passing 'verification_mode={verification_mode.value}' instead.",
FutureWarning,
)
if self._file_format is not None and self._file_format != "arrow":
raise FileFormatError('Loading a dataset not written in the "arrow" format is not supported.')
if is_remote_filesystem(self._fs):
raise NotImplementedError(f"Loading a dataset cached in a {type(self._fs).__name__} is not supported.")
if not os.path.exists(self._output_dir):
raise FileNotFoundError(
f"Dataset {self.dataset_name}: could not find data in {self._output_dir}. Please make sure to call "
"builder.download_and_prepare(), or use "
"datasets.load_dataset() before trying to access the Dataset object."
)
logger.debug(f'Constructing Dataset for split {split or ", ".join(self.info.splits)}, from {self._output_dir}')
# By default, return all splits
if split is None:
split = {s: s for s in self.info.splits}
verification_mode = VerificationMode(verification_mode or VerificationMode.BASIC_CHECKS)
# Create a dataset for each of the given splits
datasets = map_nested(
partial(
self._build_single_dataset,
run_post_process=run_post_process,
verification_mode=verification_mode,
in_memory=in_memory,
),
split,
map_tuple=True,
disable_tqdm=True,
)
if isinstance(datasets, dict):
datasets = DatasetDict(datasets)
return datasets
def _build_single_dataset(
self,
split: Union[str, ReadInstruction, Split],
run_post_process: bool,
verification_mode: VerificationMode,
in_memory: bool = False,
):
"""as_dataset for a single split."""
if not isinstance(split, ReadInstruction):
split = str(split)
if split == "all":
split = "+".join(self.info.splits.keys())
split = Split(split)
# Build base dataset
ds = self._as_dataset(
split=split,
in_memory=in_memory,
)
if run_post_process:
for resource_file_name in self._post_processing_resources(split).values():
if os.sep in resource_file_name:
raise ValueError(f"Resources shouldn't be in a sub-directory: {resource_file_name}")
resources_paths = {
resource_name: os.path.join(self._output_dir, resource_file_name)
for resource_name, resource_file_name in self._post_processing_resources(split).items()
}
post_processed = self._post_process(ds, resources_paths)
if post_processed is not None:
ds = post_processed
recorded_checksums = {}
record_checksums = False
for resource_name, resource_path in resources_paths.items():
size_checksum = get_size_checksum_dict(resource_path)
recorded_checksums[resource_name] = size_checksum
if verification_mode == VerificationMode.ALL_CHECKS and record_checksums:
if self.info.post_processed is None or self.info.post_processed.resources_checksums is None:
expected_checksums = None
else:
expected_checksums = self.info.post_processed.resources_checksums.get(split)
verify_checksums(expected_checksums, recorded_checksums, "post processing resources")
if self.info.post_processed is None:
self.info.post_processed = PostProcessedInfo()
if self.info.post_processed.resources_checksums is None:
self.info.post_processed.resources_checksums = {}
self.info.post_processed.resources_checksums[str(split)] = recorded_checksums
self.info.post_processing_size = sum(
checksums_dict["num_bytes"]
for split_checksums_dicts in self.info.post_processed.resources_checksums.values()
for checksums_dict in split_checksums_dicts.values()
)
if self.info.dataset_size is not None and self.info.download_size is not None:
self.info.size_in_bytes = (
self.info.dataset_size + self.info.download_size + self.info.post_processing_size
)
self._save_info()
ds._info.post_processed = self.info.post_processed
ds._info.post_processing_size = self.info.post_processing_size
ds._info.size_in_bytes = self.info.size_in_bytes
if self.info.post_processed.features is not None:
if self.info.post_processed.features.type != ds.features.type:
raise ValueError(
f"Post-processed features info don't match the dataset:\nGot\n{self.info.post_processed.features}\nbut expected something like\n{ds.features}"
)
else:
ds.info.features = self.info.post_processed.features
return ds
def _as_dataset(self, split: Union[ReadInstruction, Split] = Split.TRAIN, in_memory: bool = False) -> Dataset:
"""Constructs a `Dataset`.
This is the internal implementation to overwrite called when user calls
`as_dataset`. It should read the pre-processed datasets files and generate
the `Dataset` object.
Args:
split (`datasets.Split`):
which subset of the data to read.
in_memory (`bool`, defaults to `False`):
Whether to copy the data in-memory.
Returns:
`Dataset`
"""
cache_dir = self._fs._strip_protocol(self._output_dir)
dataset_name = self.dataset_name
if self._has_legacy_cache():
dataset_name = self.name
dataset_kwargs = ArrowReader(cache_dir, self.info).read(
name=dataset_name,
instructions=split,
split_infos=self.info.splits.values(),
in_memory=in_memory,
)
fingerprint = self._get_dataset_fingerprint(split)
return Dataset(fingerprint=fingerprint, **dataset_kwargs)
def _get_dataset_fingerprint(self, split: Union[ReadInstruction, Split]) -> str:
"""The dataset fingerprint is the hash of the relative directory dataset_name/config_name/version/hash, as well as the split specs."""
hasher = Hasher()
hasher.update(Path(self._relative_data_dir()).as_posix())
hasher.update(str(split)) # for example: train, train+test, train[:10%], test[:33%](pct1_dropremainder)
fingerprint = hasher.hexdigest()
return fingerprint
def as_streaming_dataset(
self,
split: Optional[str] = None,
base_path: Optional[str] = None,
) -> Union[Dict[str, IterableDataset], IterableDataset]:
if is_remote_filesystem(self._fs):
raise NotImplementedError(
f"Loading a streaming dataset cached in a {type(self._fs).__name__} is not supported yet."
)
dl_manager = StreamingDownloadManager(
base_path=base_path or self.base_path,
download_config=DownloadConfig(token=self.token, storage_options=self.storage_options),
dataset_name=self.dataset_name,
data_dir=self.config.data_dir,
)
self._check_manual_download(dl_manager)
splits_generators = {sg.name: sg for sg in self._split_generators(dl_manager)}
# By default, return all splits
if split is None:
splits_generator = splits_generators
elif split in splits_generators:
splits_generator = splits_generators[split]
else:
raise ValueError(f"Bad split: {split}. Available splits: {list(splits_generators)}")
# Create a dataset for each of the given splits
datasets = map_nested(
self._as_streaming_dataset_single,
splits_generator,
map_tuple=True,
)
if isinstance(datasets, dict):
datasets = IterableDatasetDict(datasets)
return datasets
def _as_streaming_dataset_single(
self,
splits_generator,
) -> IterableDataset:
ex_iterable = self._get_examples_iterable_for_split(splits_generator)
# add auth to be able to access and decode audio/image files from private repositories.
token_per_repo_id = {self.repo_id: self.token} if self.repo_id else {}
return IterableDataset(
ex_iterable, info=self.info, split=splits_generator.name, token_per_repo_id=token_per_repo_id
)
def _post_process(self, dataset: Dataset, resources_paths: Mapping[str, str]) -> Optional[Dataset]:
"""Run dataset transforms or add indexes"""
return None
def _post_processing_resources(self, split: str) -> Dict[str, str]:
"""Mapping resource_name -> resource_file_name"""
return {}
def _download_post_processing_resources(
self, split: str, resource_name: str, dl_manager: DownloadManager
) -> Optional[str]:
"""Download the resource using the download manager and return the downloaded path."""
return None
@abc.abstractmethod
def _split_generators(self, dl_manager: DownloadManager):
"""Specify feature dictionary generators and dataset splits.
This function returns a list of `SplitGenerator`s defining how to generate
data and what splits to use.
Example:
return [
datasets.SplitGenerator(
name=datasets.Split.TRAIN,
gen_kwargs={'file': 'train_data.zip'},
),
datasets.SplitGenerator(
name=datasets.Split.TEST,
gen_kwargs={'file': 'test_data.zip'},
),
]
The above code will first call `_generate_examples(file='train_data.zip')`
to write the train data, then `_generate_examples(file='test_data.zip')` to
write the test data.
Datasets are typically split into different subsets to be used at various
stages of training and evaluation.
Note that for datasets without a `VALIDATION` split, you can use a
fraction of the `TRAIN` data for evaluation as you iterate on your model
so as not to overfit to the `TEST` data.
For downloads and extractions, use the given `download_manager`.
Note that the `DownloadManager` caches downloads, so it is fine to have each
generator attempt to download the source data.
A good practice is to download all data in this function, and then
distribute the relevant parts to each split with the `gen_kwargs` argument
Args:
dl_manager (`DownloadManager`):
Download manager to download the data
Returns:
`list<SplitGenerator>`.
"""
raise NotImplementedError()
@abc.abstractmethod
def _prepare_split(
self,
split_generator: SplitGenerator,
file_format: str = "arrow",
max_shard_size: Optional[Union[str, int]] = None,
num_proc: Optional[int] = None,
**kwargs,
):
"""Generate the examples and record them on disk.
Args:
split_generator (`SplitGenerator`):
Split generator to process
file_format (`str`, *optional*):
format of the data files in which the dataset will be written.
Supported formats: "arrow", "parquet". Default to "arrow" format.
max_shard_size (`Union[str, int]`, *optional*):
Maximum number of bytes written per shard, default is "500MB".
The size is based on uncompressed data size, so in practice your shard files may be smaller than
`max_shard_size` thanks to Parquet compression for example.
num_proc (`int`, *optional*, defaults to `None`):
Number of processes when downloading and generating the dataset locally.
Multiprocessing is disabled by default.
<Added version="2.7.0"/>
**kwargs: Additional kwargs forwarded from _download_and_prepare (ex:
beam pipeline)
"""
raise NotImplementedError()
def _get_examples_iterable_for_split(self, split_generator: SplitGenerator) -> ExamplesIterable:
"""Generate the examples on the fly.
Args:
split_generator (`SplitGenerator`):
Split generator to process
"""
raise NotImplementedError()
class GeneratorBasedBuilder(DatasetBuilder):
"""Base class for datasets with data generation based on dict generators.
`GeneratorBasedBuilder` is a convenience class that abstracts away much
of the data writing and reading of `DatasetBuilder`. It expects subclasses to
implement generators of feature dictionaries across the dataset splits
(`_split_generators`). See the method docstrings for details.
"""
@abc.abstractmethod
def _generate_examples(self, **kwargs):
"""Default function generating examples for each `SplitGenerator`.
This function preprocess the examples from the raw data to the preprocessed
dataset files.
This function is called once for each `SplitGenerator` defined in
`_split_generators`. The examples yielded here will be written on
disk.
Args:
**kwargs (additional keyword arguments):
Arguments forwarded from the SplitGenerator.gen_kwargs
Yields:
key: `str` or `int`, a unique deterministic example identification key.
* Unique: An error will be raised if two examples are yield with the
same key.
* Deterministic: When generating the dataset twice, the same example
should have the same key.
Good keys can be the image id, or line number if examples are extracted
from a text file.
The key will be hashed and sorted to shuffle examples deterministically,
such as generating the dataset multiple times keep examples in the
same order.
example: `dict<str feature_name, feature_value>`, a feature dictionary
ready to be encoded and written to disk. The example will be
encoded with `self.info.features.encode_example({...})`.
"""
raise NotImplementedError()
def _prepare_split(
self,
split_generator: SplitGenerator,
check_duplicate_keys: bool,
file_format="arrow",
num_proc: Optional[int] = None,
max_shard_size: Optional[Union[int, str]] = None,
):
max_shard_size = convert_file_size_to_int(max_shard_size or config.MAX_SHARD_SIZE)
if self.info.splits is not None:
split_info = self.info.splits[split_generator.name]
else:
split_info = split_generator.split_info
SUFFIX = "-JJJJJ-SSSSS-of-NNNNN"
fname = f"{self.dataset_name}-{split_generator.name}{SUFFIX}.{file_format}"
fpath = posixpath.join(self._output_dir, fname)
if num_proc and num_proc > 1:
num_input_shards = _number_of_shards_in_gen_kwargs(split_generator.gen_kwargs)
if num_input_shards <= 1:
logger.warning(
f"Setting num_proc from {num_proc} back to 1 for the {split_info.name} split to disable multiprocessing as it only contains one shard."
)
num_proc = 1
elif num_input_shards < num_proc:
logger.warning(
f"Setting num_proc from {num_proc} to {num_input_shards} for the {split_info.name} split as it only contains {num_input_shards} shards."
)
num_proc = num_input_shards
pbar = hf_tqdm(
unit=" examples",
total=split_info.num_examples,
desc=f"Generating {split_info.name} split",
)
_prepare_split_args = {
"fpath": fpath,
"file_format": file_format,
"max_shard_size": max_shard_size,
"split_info": split_info,
"check_duplicate_keys": check_duplicate_keys,
}
if num_proc is None or num_proc == 1:
result = None
gen_kwargs = split_generator.gen_kwargs
job_id = 0
with pbar:
for job_id, done, content in self._prepare_split_single(
gen_kwargs=gen_kwargs, job_id=job_id, **_prepare_split_args
):
if done:
result = content
else:
pbar.update(content)
# wrapping everything into lists for consistency with the multiprocessed code path
assert result is not None, "Failed to retrieve results from prepare_split"
examples_per_job, bytes_per_job, features_per_job, shards_per_job, shard_lengths_per_job = [
[item] for item in result
]
else:
kwargs_per_job = [
{"gen_kwargs": gen_kwargs, "job_id": job_id, **_prepare_split_args}
for job_id, gen_kwargs in enumerate(
_split_gen_kwargs(split_generator.gen_kwargs, max_num_jobs=num_proc)
)
]
num_jobs = len(kwargs_per_job)
examples_per_job = [None] * num_jobs
bytes_per_job = [None] * num_jobs
features_per_job = [None] * num_jobs
shards_per_job = [None] * num_jobs
shard_lengths_per_job = [None] * num_jobs
with Pool(num_proc) as pool:
with pbar:
for job_id, done, content in iflatmap_unordered(
pool, self._prepare_split_single, kwargs_iterable=kwargs_per_job
):
if done:
# the content is the result of the job
(
examples_per_job[job_id],
bytes_per_job[job_id],
features_per_job[job_id],
shards_per_job[job_id],
shard_lengths_per_job[job_id],
) = content
else:
# the content is the number of examples progress update
pbar.update(content)
assert (
None not in examples_per_job
), f"Failed to retrieve results from prepare_split: result list {examples_per_job} still contains None - at least one worker failed to return its results"
total_shards = sum(shards_per_job)
total_num_examples = sum(examples_per_job)
total_num_bytes = sum(bytes_per_job)
features = features_per_job[0]
split_generator.split_info.num_examples = total_num_examples
split_generator.split_info.num_bytes = total_num_bytes
# should rename everything at the end
logger.debug(f"Renaming {total_shards} shards.")
if total_shards > 1:
# use the -SSSSS-of-NNNNN pattern
def _rename_shard(shard_and_job: Tuple[int]):
shard_id, job_id = shard_and_job
global_shard_id = sum(shards_per_job[:job_id]) + shard_id
self._rename(
fpath.replace("SSSSS", f"{shard_id:05d}").replace("JJJJJ", f"{job_id:05d}"),
fpath.replace("JJJJJ-SSSSS", f"{global_shard_id:05d}").replace("NNNNN", f"{total_shards:05d}"),
)
shards_and_jobs = [
(shard_id, job_id)
for job_id, num_shards in enumerate(shards_per_job)
for shard_id in range(num_shards)
]
thread_map(_rename_shard, shards_and_jobs, disable=True, max_workers=64)
split_generator.split_info.shard_lengths = [
shard_length for shard_lengths in shard_lengths_per_job for shard_length in shard_lengths
]
else:
# don't use any pattern
shard_id, job_id = 0, 0
self._rename(
fpath.replace("SSSSS", f"{shard_id:05d}").replace("JJJJJ", f"{job_id:05d}"),
fpath.replace(SUFFIX, ""),
)
if self.info.features is None:
self.info.features = features
def _prepare_split_single(
self,
gen_kwargs: dict,
fpath: str,
file_format: str,
max_shard_size: int,
split_info: SplitInfo,
check_duplicate_keys: bool,
job_id: int,
) -> Iterable[Tuple[int, bool, Union[int, tuple]]]:
generator = self._generate_examples(**gen_kwargs)
writer_class = ParquetWriter if file_format == "parquet" else ArrowWriter
embed_local_files = file_format == "parquet"
shard_lengths = []
total_num_examples, total_num_bytes = 0, 0
shard_id = 0
num_examples_progress_update = 0
try:
writer = writer_class(
features=self.info.features,
path=fpath.replace("SSSSS", f"{shard_id:05d}").replace("JJJJJ", f"{job_id:05d}"),
writer_batch_size=self._writer_batch_size,
hash_salt=split_info.name,
check_duplicates=check_duplicate_keys,
storage_options=self._fs.storage_options,
embed_local_files=embed_local_files,
)
try:
_time = time.time()
for key, record in generator:
if max_shard_size is not None and writer._num_bytes > max_shard_size:
num_examples, num_bytes = writer.finalize()
writer.close()
shard_lengths.append(num_examples)
total_num_examples += num_examples
total_num_bytes += num_bytes
shard_id += 1
writer = writer_class(
features=writer._features,
path=fpath.replace("SSSSS", f"{shard_id:05d}").replace("JJJJJ", f"{job_id:05d}"),
writer_batch_size=self._writer_batch_size,
hash_salt=split_info.name,
check_duplicates=check_duplicate_keys,
storage_options=self._fs.storage_options,
embed_local_files=embed_local_files,
)
example = self.info.features.encode_example(record) if self.info.features is not None else record
writer.write(example, key)
num_examples_progress_update += 1
if time.time() > _time + config.PBAR_REFRESH_TIME_INTERVAL:
_time = time.time()
yield job_id, False, num_examples_progress_update
num_examples_progress_update = 0
finally:
yield job_id, False, num_examples_progress_update
num_shards = shard_id + 1
num_examples, num_bytes = writer.finalize()
writer.close()
shard_lengths.append(num_examples)
total_num_examples += num_examples
total_num_bytes += num_bytes
except Exception as e:
# Ignore the writer's error for no examples written to the file if this error was caused by the error in _generate_examples before the first example was yielded
if isinstance(e, SchemaInferenceError) and e.__context__ is not None:
e = e.__context__
raise DatasetGenerationError("An error occurred while generating the dataset") from e
yield job_id, True, (total_num_examples, total_num_bytes, writer._features, num_shards, shard_lengths)
def _download_and_prepare(self, dl_manager, verification_mode, **prepare_splits_kwargs):
super()._download_and_prepare(
dl_manager,
verification_mode,
check_duplicate_keys=verification_mode == VerificationMode.BASIC_CHECKS
or verification_mode == VerificationMode.ALL_CHECKS,
**prepare_splits_kwargs,
)
def _get_examples_iterable_for_split(self, split_generator: SplitGenerator) -> ExamplesIterable:
return ExamplesIterable(self._generate_examples, split_generator.gen_kwargs)
class ArrowBasedBuilder(DatasetBuilder):
"""Base class for datasets with data generation based on Arrow loading functions (CSV/JSON/Parquet)."""
@abc.abstractmethod
def _generate_tables(self, **kwargs):
"""Default function generating examples for each `SplitGenerator`.
This function preprocess the examples from the raw data to the preprocessed
dataset files.
This function is called once for each `SplitGenerator` defined in
`_split_generators`. The examples yielded here will be written on
disk.
Args:
**kwargs (additional keyword arguments):
Arguments forwarded from the SplitGenerator.gen_kwargs
Yields:
key: `str` or `int`, a unique deterministic example identification key.
* Unique: An error will be raised if two examples are yield with the
same key.
* Deterministic: When generating the dataset twice, the same example
should have the same key.
Good keys can be the image id, or line number if examples are extracted
from a text file.
The key will be hashed and sorted to shuffle examples deterministically,
such as generating the dataset multiple times keep examples in the
same order.
example: `pyarrow.Table`, a feature table
ready to be encoded and written to disk.
"""
raise NotImplementedError()
def _prepare_split(
self,
split_generator: SplitGenerator,
file_format: str = "arrow",
num_proc: Optional[int] = None,
max_shard_size: Optional[Union[str, int]] = None,
):
max_shard_size = convert_file_size_to_int(max_shard_size or config.MAX_SHARD_SIZE)
try:
split_info = self.info.splits[split_generator.name]
except Exception:
split_info = split_generator.split_info
SUFFIX = "-JJJJJ-SSSSS-of-NNNNN"
fname = f"{self.dataset_name}-{split_generator.name}{SUFFIX}.{file_format}"
fpath = posixpath.join(self._output_dir, fname)
if num_proc and num_proc > 1:
num_input_shards = _number_of_shards_in_gen_kwargs(split_generator.gen_kwargs)
if num_input_shards <= 1:
logger.warning(
f"Setting num_proc from {num_proc} back to 1 for the {split_info.name} split to disable multiprocessing as it only contains one shard."
)
num_proc = 1
elif num_input_shards < num_proc:
logger.warning(
f"Setting num_proc from {num_proc} to {num_input_shards} for the {split_info.name} split as it only contains {num_input_shards} shards."
)
num_proc = num_input_shards
pbar = hf_tqdm(
unit=" examples",
total=split_info.num_examples,
desc=f"Generating {split_info.name} split",
)
_prepare_split_args = {
"fpath": fpath,
"file_format": file_format,
"max_shard_size": max_shard_size,
}
if num_proc is None or num_proc == 1:
result = None
gen_kwargs = split_generator.gen_kwargs
job_id = 0
with pbar:
for job_id, done, content in self._prepare_split_single(
gen_kwargs=gen_kwargs, job_id=job_id, **_prepare_split_args
):
if done:
result = content
else:
pbar.update(content)
# wrapping everything into lists for consistency with the multiprocessed code path
assert result is not None, "Failed to retrieve results from prepare_split"
examples_per_job, bytes_per_job, features_per_job, shards_per_job, shard_lengths_per_job = [
[item] for item in result
]
else:
kwargs_per_job = [
{"gen_kwargs": gen_kwargs, "job_id": job_id, **_prepare_split_args}
for job_id, gen_kwargs in enumerate(
_split_gen_kwargs(split_generator.gen_kwargs, max_num_jobs=num_proc)
)
]
num_jobs = len(kwargs_per_job)
examples_per_job = [None] * num_jobs
bytes_per_job = [None] * num_jobs
features_per_job = [None] * num_jobs
shards_per_job = [None] * num_jobs
shard_lengths_per_job = [None] * num_jobs
with Pool(num_proc) as pool:
with pbar:
for job_id, done, content in iflatmap_unordered(
pool, self._prepare_split_single, kwargs_iterable=kwargs_per_job
):
if done:
# the content is the result of the job
(
examples_per_job[job_id],
bytes_per_job[job_id],
features_per_job[job_id],
shards_per_job[job_id],
shard_lengths_per_job[job_id],
) = content
else:
# the content is the number of examples progress update
pbar.update(content)
assert (
None not in examples_per_job
), f"Failed to retrieve results from prepare_split: result list {examples_per_job} still contains None - at least one worker failed to return its results"
total_shards = sum(shards_per_job)
total_num_examples = sum(examples_per_job)
total_num_bytes = sum(bytes_per_job)
features = features_per_job[0]
split_generator.split_info.num_examples = total_num_examples
split_generator.split_info.num_bytes = total_num_bytes
# should rename everything at the end
logger.debug(f"Renaming {total_shards} shards.")
if total_shards > 1:
# use the -SSSSS-of-NNNNN pattern
def _rename_shard(shard_id_and_job: Tuple[int]):
shard_id, job_id = shard_id_and_job
global_shard_id = sum(shards_per_job[:job_id]) + shard_id
self._rename(
fpath.replace("SSSSS", f"{shard_id:05d}").replace("JJJJJ", f"{job_id:05d}"),
fpath.replace("JJJJJ-SSSSS", f"{global_shard_id:05d}").replace("NNNNN", f"{total_shards:05d}"),
)
shard_ids_and_jobs = [
(shard_id, job_id)
for job_id, num_shards in enumerate(shards_per_job)
for shard_id in range(num_shards)
]
thread_map(_rename_shard, shard_ids_and_jobs, disable=True, max_workers=64)
split_generator.split_info.shard_lengths = [
shard_length for shard_lengths in shard_lengths_per_job for shard_length in shard_lengths
]
else:
# don't use any pattern
shard_id, job_id = 0, 0
self._rename(
fpath.replace("SSSSS", f"{shard_id:05d}").replace("JJJJJ", f"{job_id:05d}"),
fpath.replace(SUFFIX, ""),
)
if self.info.features is None:
self.info.features = features
def _prepare_split_single(
self, gen_kwargs: dict, fpath: str, file_format: str, max_shard_size: int, job_id: int
) -> Iterable[Tuple[int, bool, Union[int, tuple]]]:
generator = self._generate_tables(**gen_kwargs)
writer_class = ParquetWriter if file_format == "parquet" else ArrowWriter
embed_local_files = file_format == "parquet"
shard_lengths = []
total_num_examples, total_num_bytes = 0, 0
shard_id = 0
num_examples_progress_update = 0
try:
writer = writer_class(
features=self.info.features,
path=fpath.replace("SSSSS", f"{shard_id:05d}").replace("JJJJJ", f"{job_id:05d}"),
writer_batch_size=self._writer_batch_size,
storage_options=self._fs.storage_options,
embed_local_files=embed_local_files,
)
try:
_time = time.time()
for _, table in generator:
if max_shard_size is not None and writer._num_bytes > max_shard_size:
num_examples, num_bytes = writer.finalize()
writer.close()
shard_lengths.append(num_examples)
total_num_examples += num_examples
total_num_bytes += num_bytes
shard_id += 1
writer = writer_class(
features=writer._features,
path=fpath.replace("SSSSS", f"{shard_id:05d}").replace("JJJJJ", f"{job_id:05d}"),
writer_batch_size=self._writer_batch_size,
storage_options=self._fs.storage_options,
embed_local_files=embed_local_files,
)
writer.write_table(table)
num_examples_progress_update += len(table)
if time.time() > _time + config.PBAR_REFRESH_TIME_INTERVAL:
_time = time.time()
yield job_id, False, num_examples_progress_update
num_examples_progress_update = 0
finally:
yield job_id, False, num_examples_progress_update
num_shards = shard_id + 1
num_examples, num_bytes = writer.finalize()
writer.close()
shard_lengths.append(num_examples)
total_num_examples += num_examples
total_num_bytes += num_bytes
except Exception as e:
# Ignore the writer's error for no examples written to the file if this error was caused by the error in _generate_examples before the first example was yielded
if isinstance(e, SchemaInferenceError) and e.__context__ is not None:
e = e.__context__
raise DatasetGenerationError("An error occurred while generating the dataset") from e
yield job_id, True, (total_num_examples, total_num_bytes, writer._features, num_shards, shard_lengths)
def _get_examples_iterable_for_split(self, split_generator: SplitGenerator) -> ExamplesIterable:
return ArrowExamplesIterable(self._generate_tables, kwargs=split_generator.gen_kwargs)
class MissingBeamOptions(ValueError):
pass
class BeamBasedBuilder(DatasetBuilder):
"""Beam-based Builder."""
def __init__(self, *args, beam_runner=None, beam_options=None, **kwargs):
self._beam_runner = beam_runner
self._beam_options = beam_options
self._beam_writers = {} # {split: beam_writer} mapping.
super().__init__(*args, **kwargs)
def _make_split_generators_kwargs(self, prepare_split_kwargs):
# Pass `pipeline` into `_split_generators()` from `prepare_split_kwargs` if
# it's in the call signature of `_split_generators()`.
# This allows for global preprocessing in beam.
split_generators_kwargs = {}
split_generators_arg_names = inspect.signature(self._split_generators).parameters.keys()
if "pipeline" in split_generators_arg_names:
split_generators_kwargs["pipeline"] = prepare_split_kwargs["pipeline"]
return split_generators_kwargs
@abc.abstractmethod
def _build_pcollection(self, pipeline, **kwargs):
"""Build the beam pipeline examples for each `SplitGenerator`.
This function extracts examples from the raw data with parallel transforms
in a Beam pipeline. It is called once for each `SplitGenerator` defined in
`_split_generators`. The examples from the PCollection will be
encoded and written to disk.
<Tip warning={true}>
Warning: When running in a distributed setup, make sure that the data
which will be read (download_dir, manual_dir,...) and written (cache_dir)
can be accessed by the workers jobs. The data should be located in a
shared filesystem, like GCS.
</Tip>
Args:
pipeline ([`utils.beam_utils.BeamPipeline`]):
Apache Beam pipeline.
**kwargs (additional keyword arguments):
Arguments forwarded from the SplitGenerator.gen_kwargs.
Returns:
`beam.PCollection`: Apache Beam PCollection containing the
example to send to `self.info.features.encode_example(...)`.
Example:
```
def _build_pcollection(pipeline, extracted_dir=None):
return (
pipeline
| beam.Create(gfile.io.listdir(extracted_dir))
| beam.Map(_process_file)
)
```
"""
raise NotImplementedError()
def _download_and_prepare(self, dl_manager, verification_mode, **prepare_splits_kwargs):
# Create the Beam pipeline and forward it to `_prepare_split`
import apache_beam as beam
import datasets.utils.beam_utils as beam_utils
beam_runner = self._beam_runner
beam_options = self._beam_options
if not beam_runner and not beam_options:
usage_example = f"load_dataset('{self.name}', '{self.config.name}', beam_runner='DirectRunner')"
raise MissingBeamOptions(
"Trying to generate a dataset using Apache Beam, yet no Beam Runner "
"or PipelineOptions() has been provided in `load_dataset` or in the "
"builder arguments. For big datasets it has to run on large-scale data "
"processing tools like Dataflow, Spark, etc. More information about "
"Apache Beam runners at "
"https://beam.apache.org/documentation/runners/capability-matrix/"
"\nIf you really want to run it locally because you feel like the "
"Dataset is small enough, you can use the local beam runner called "
"`DirectRunner` (you may run out of memory). \nExample of usage: "
f"\n\t`{usage_example}`"
)
if self._writer_batch_size is not None:
logger.warning(
"`writer_batch_size` is not supported for beam pipelines yet. Using the default chunk size for writing."
)
# Beam type checking assumes transforms multiple outputs are of same type,
# which is not our case. Plus it doesn't handle correctly all types, so we
# are better without it.
pipeline_options = {"pipeline_type_check": False}
if "num_proc" in prepare_splits_kwargs:
num_workers = prepare_splits_kwargs.pop("num_proc")
pipeline_options["direct_num_workers"] = num_workers
pipeline_options["num_workers"] = num_workers
pipeline_options["direct_running_mode"] = "multi_processing"
# TODO: Fix ModuleNotFoundError: No module named 'datasets_modules' when running multiprocessed DirectRunner
raise NotImplementedError("Using a DirectRunner with `num_proc` for multiprocessing it not supported yet.")
beam_options = beam_options or beam.options.pipeline_options.PipelineOptions.from_dictionary(pipeline_options)
# Use a single pipeline for all splits
pipeline = beam_utils.BeamPipeline(
runner=beam_runner,
options=beam_options,
)
super()._download_and_prepare(
dl_manager, verification_mode=VerificationMode.NO_CHECKS, pipeline=pipeline, **prepare_splits_kwargs
) # TODO handle verification_mode in beam datasets
# Run pipeline
pipeline_results = pipeline.run()
pipeline_results.wait_until_finish()
metrics = pipeline_results.metrics()
# Update `info.splits`.
split_dict = self.info.splits
for split_name, beam_writer in self._beam_writers.items():
m_filter = beam.metrics.MetricsFilter().with_namespace(namespace=split_name)
num_examples, num_bytes = beam_writer.finalize(metrics.query(m_filter))
split_info = split_dict[split_name]
split_info.num_examples = num_examples
split_info.num_bytes = num_bytes
if hasattr(beam_writer, "_shard_lengths") and len(beam_writer._shard_lengths) > 1:
# keep the -SSSSS-of-NNNNN pattern
split_info.shard_lengths = beam_writer._shard_lengths
else:
# don't use any pattern
file_format = prepare_splits_kwargs.get("file_format", "arrow")
src_fname = f"{self.dataset_name}-{split_name}-00000-of-00001.{file_format}"
dst_fname = f"{self.dataset_name}-{split_name}.{file_format}"
src_fpath = posixpath.join(self._output_dir, src_fname)
dst_fpath = posixpath.join(self._output_dir, dst_fname)
self._rename(src_fpath, dst_fpath)
def _save_info(self):
download_config = (
self.dl_manager.download_config
if self.dl_manager
else DownloadConfig(token=self.token, storage_options=self._fs.storage_options)
)
with xopen(f"{self._output_dir}/{config.DATASET_INFO_FILENAME}", "wb", download_config=download_config) as f:
self.info._dump_info(f)
if self.info.license:
with xopen(f"{self._output_dir}/{config.LICENSE_FILENAME}", "wb", download_config=download_config) as f:
self.info._dump_license(f)
def _prepare_split(
self, split_generator, pipeline, file_format="arrow", max_shard_size: Optional[Union[str, int]] = None
):
import apache_beam as beam
if max_shard_size is not None:
raise NotImplementedError(
"max_shard_size is not supported for Beam datasets."
"Please set it to None to use the default Apache Beam sharding and get the best performance."
)
# To write examples in filesystem:
split_name = split_generator.split_info.name
fname = f"{self.dataset_name}-{split_name}.{file_format}"
fpath = posixpath.join(self._output_dir, fname)
beam_writer = BeamWriter(
features=self.info.features, path=fpath, namespace=split_name, cache_dir=self._output_dir
)
self._beam_writers[split_name] = beam_writer
encode_example = self.info.features.encode_example
# Note: We need to wrap the pipeline in a PTransform to avoid re-using the
# same label names for each split
@beam.ptransform_fn
def _build_pcollection(pipeline):
"""PTransformation which build a single split."""
# Encode the PCollection
pcoll_examples = self._build_pcollection(pipeline, **split_generator.gen_kwargs)
pcoll_examples |= "Encode" >> beam.Map(lambda key_ex: (key_ex[0], encode_example(key_ex[1])))
return beam_writer.write_from_pcollection(pcoll_examples)
# Add the PCollection to the pipeline
_ = pipeline | split_name >> _build_pcollection() # pylint: disable=no-value-for-parameter max_bytes_per_shard
def as_streaming_dataset(
self,
split: Optional[str] = None,
) -> Union[Dict[str, IterableDataset], IterableDataset]:
self._request_info_from_hf_gcs()
datasets = {
split.name: IterableDataset(self._get_examples_iterable_for_split(split), info=self.info, split=split.name)
for split in self.info.splits.values()
}
if split:
try:
datasets = datasets[split]
except KeyError:
raise ValueError(f"Bad split: {split}. Available splits: {list(datasets)}")
if isinstance(datasets, dict):
datasets = IterableDatasetDict(datasets)
return datasets
def _get_examples_iterable_for_split(self, split: SplitInfo) -> ExamplesIterable:
return ExamplesIterable(self._generate_examples_from_hf_gcs, {"split": split})
def _generate_examples_from_hf_gcs(self, split: SplitInfo):
if split.shard_lengths:
num_shards = len(split.shard_lengths)
remote_prepared_urls = [
f"{self._remote_cache_dir_from_hf_gcs}/{self.name}-{split.name}-{shard_id:05d}-of-{num_shards:05d}.arrow"
for shard_id in range(num_shards)
]
else:
remote_prepared_urls = [f"{self._remote_cache_dir_from_hf_gcs}/{self.name}-{split.name}.arrow"]
key = 0
download_config = (
self.dl_manager.download_config
if self.dl_manager
else DownloadConfig(token=self.token, storage_options=self._fs.storage_options)
)
for remote_prepared_url in remote_prepared_urls:
with xopen(remote_prepared_url, "rb", download_config=download_config) as f:
with pa.ipc.open_stream(f) as reader:
for record_batch in reader:
for record in record_batch.to_pylist():
yield key, record
key += 1
def _request_info_from_hf_gcs(self):
from .download.streaming_download_manager import xopen
remote_dataset_info = f"{self._remote_cache_dir_from_hf_gcs}/{config.DATASET_INFO_FILENAME}"
try:
download_config = download_config = (
self.dl_manager.download_config
if self.dl_manager
else DownloadConfig(token=self.token, storage_options=self._fs.storage_options)
)
with xopen(remote_dataset_info, download_config=download_config) as f:
import json
_info = json.load(f)
except FileNotFoundError as err:
raise DatasetNotOnHfGcsError(err) from None
self.info.update(DatasetInfo.from_dict(_info))
@property
def _remote_cache_dir_from_hf_gcs(self):
relative_data_dir = self._relative_data_dir(with_hash=False)
return HF_GCP_BASE_URL + "/" + Path(relative_data_dir).as_posix()
| 0
|
hf_public_repos/datasets/src
|
hf_public_repos/datasets/src/datasets/load.py
|
# Copyright 2020 The HuggingFace Datasets Authors and the TensorFlow Datasets Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# Lint as: python3
"""Access datasets."""
import filecmp
import importlib
import inspect
import json
import os
import posixpath
import shutil
import signal
import time
import warnings
from collections import Counter
from dataclasses import dataclass, field
from pathlib import Path
from typing import Any, Dict, List, Mapping, Optional, Sequence, Tuple, Type, Union
import fsspec
import requests
from huggingface_hub import DatasetCard, DatasetCardData, HfApi
from . import config
from .arrow_dataset import Dataset
from .builder import BuilderConfig, DatasetBuilder
from .data_files import (
DEFAULT_PATTERNS_ALL,
DataFilesDict,
DataFilesList,
EmptyDatasetError,
get_data_patterns,
get_metadata_patterns,
sanitize_patterns,
)
from .dataset_dict import DatasetDict, IterableDatasetDict
from .download.download_config import DownloadConfig
from .download.download_manager import DownloadMode
from .download.streaming_download_manager import StreamingDownloadManager, xbasename, xglob, xjoin
from .exceptions import DataFilesNotFoundError, DatasetNotFoundError
from .features import Features
from .fingerprint import Hasher
from .info import DatasetInfo, DatasetInfosDict
from .iterable_dataset import IterableDataset
from .metric import Metric
from .naming import camelcase_to_snakecase, snakecase_to_camelcase
from .packaged_modules import (
_EXTENSION_TO_MODULE,
_MODULE_SUPPORTS_METADATA,
_MODULE_TO_EXTENSIONS,
_PACKAGED_DATASETS_MODULES,
_hash_python_lines,
)
from .splits import Split
from .utils._filelock import FileLock
from .utils.deprecation_utils import deprecated
from .utils.file_utils import (
OfflineModeIsEnabled,
_raise_if_offline_mode_is_enabled,
cached_path,
head_hf_s3,
hf_github_url,
init_hf_modules,
is_relative_path,
relative_to_absolute_path,
url_or_path_join,
)
from .utils.hub import hf_hub_url
from .utils.info_utils import VerificationMode, is_small_dataset
from .utils.logging import get_logger
from .utils.metadata import MetadataConfigs
from .utils.py_utils import get_imports
from .utils.version import Version
logger = get_logger(__name__)
ALL_ALLOWED_EXTENSIONS = list(_EXTENSION_TO_MODULE.keys()) + [".zip"]
def _raise_timeout_error(signum, frame):
raise ValueError(
"Loading this dataset requires you to execute custom code contained in the dataset repository on your local "
"machine. Please set the option `trust_remote_code=True` to permit loading of this dataset."
)
def resolve_trust_remote_code(trust_remote_code: Optional[bool], repo_id: str) -> bool:
"""
Copied and adapted from Transformers
https://github.com/huggingface/transformers/blob/2098d343cc4b4b9d2aea84b3cf1eb5a1e610deff/src/transformers/dynamic_module_utils.py#L589
"""
trust_remote_code = trust_remote_code if trust_remote_code is not None else config.HF_DATASETS_TRUST_REMOTE_CODE
if trust_remote_code is None:
if config.TIME_OUT_REMOTE_CODE > 0:
try:
signal.signal(signal.SIGALRM, _raise_timeout_error)
signal.alarm(config.TIME_OUT_REMOTE_CODE)
while trust_remote_code is None:
answer = input(
f"The repository for {repo_id} contains custom code which must be executed to correctly "
f"load the dataset. You can inspect the repository content at https://hf.co/datasets/{repo_id}.\n"
f"You can avoid this prompt in future by passing the argument `trust_remote_code=True`.\n\n"
f"Do you wish to run the custom code? [y/N] "
)
if answer.lower() in ["yes", "y", "1"]:
trust_remote_code = True
elif answer.lower() in ["no", "n", "0", ""]:
trust_remote_code = False
signal.alarm(0)
except Exception:
# OS which does not support signal.SIGALRM
raise ValueError(
f"The repository for {repo_id} contains custom code which must be executed to correctly "
f"load the dataset. You can inspect the repository content at https://hf.co/datasets/{repo_id}.\n"
f"Please pass the argument `trust_remote_code=True` to allow custom code to be run."
)
else:
# For the CI which might put the timeout at 0
_raise_timeout_error(None, None)
return trust_remote_code
def init_dynamic_modules(
name: str = config.MODULE_NAME_FOR_DYNAMIC_MODULES, hf_modules_cache: Optional[Union[Path, str]] = None
):
"""
Create a module with name `name` in which you can add dynamic modules
such as metrics or datasets. The module can be imported using its name.
The module is created in the HF_MODULE_CACHE directory by default (~/.cache/huggingface/modules) but it can
be overridden by specifying a path to another directory in `hf_modules_cache`.
"""
hf_modules_cache = init_hf_modules(hf_modules_cache)
dynamic_modules_path = os.path.join(hf_modules_cache, name)
os.makedirs(dynamic_modules_path, exist_ok=True)
if not os.path.exists(os.path.join(dynamic_modules_path, "__init__.py")):
with open(os.path.join(dynamic_modules_path, "__init__.py"), "w"):
pass
return dynamic_modules_path
def import_main_class(module_path, dataset=True) -> Optional[Union[Type[DatasetBuilder], Type[Metric]]]:
"""Import a module at module_path and return its main class:
- a DatasetBuilder if dataset is True
- a Metric if dataset is False
"""
module = importlib.import_module(module_path)
if dataset:
main_cls_type = DatasetBuilder
else:
main_cls_type = Metric
# Find the main class in our imported module
module_main_cls = None
for name, obj in module.__dict__.items():
if inspect.isclass(obj) and issubclass(obj, main_cls_type):
if inspect.isabstract(obj):
continue
module_main_cls = obj
obj_module = inspect.getmodule(obj)
if obj_module is not None and module == obj_module:
break
return module_main_cls
class _InitializeConfiguredDatasetBuilder:
"""
From https://stackoverflow.com/questions/4647566/pickle-a-dynamically-parameterized-sub-class
See also ConfiguredDatasetBuilder.__reduce__
When called with the param value as the only argument, returns an
un-initialized instance of the parameterized class. Subsequent __setstate__
will be called by pickle.
"""
def __call__(self, builder_cls, metadata_configs, default_config_name, name):
# make a simple object which has no complex __init__ (this one will do)
obj = _InitializeConfiguredDatasetBuilder()
obj.__class__ = configure_builder_class(
builder_cls, metadata_configs, default_config_name=default_config_name, dataset_name=name
)
return obj
def configure_builder_class(
builder_cls: Type[DatasetBuilder],
builder_configs: List[BuilderConfig],
default_config_name: Optional[str],
dataset_name: str,
) -> Type[DatasetBuilder]:
"""
Dynamically create a builder class with custom builder configs parsed from README.md file,
i.e. set BUILDER_CONFIGS class variable of a builder class to custom configs list.
"""
class ConfiguredDatasetBuilder(builder_cls):
BUILDER_CONFIGS = builder_configs
DEFAULT_CONFIG_NAME = default_config_name
__module__ = builder_cls.__module__ # so that the actual packaged builder can be imported
def __reduce__(self): # to make dynamically created class pickable, see _InitializeParameterizedDatasetBuilder
parent_builder_cls = self.__class__.__mro__[1]
return (
_InitializeConfiguredDatasetBuilder(),
(
parent_builder_cls,
self.BUILDER_CONFIGS,
self.DEFAULT_CONFIG_NAME,
self.dataset_name,
),
self.__dict__.copy(),
)
ConfiguredDatasetBuilder.__name__ = (
f"{builder_cls.__name__.lower().capitalize()}{snakecase_to_camelcase(dataset_name)}"
)
ConfiguredDatasetBuilder.__qualname__ = (
f"{builder_cls.__name__.lower().capitalize()}{snakecase_to_camelcase(dataset_name)}"
)
return ConfiguredDatasetBuilder
def get_dataset_builder_class(
dataset_module: "DatasetModule", dataset_name: Optional[str] = None
) -> Type[DatasetBuilder]:
builder_cls = import_main_class(dataset_module.module_path)
if dataset_module.builder_configs_parameters.builder_configs:
builder_cls = configure_builder_class(
builder_cls,
builder_configs=dataset_module.builder_configs_parameters.builder_configs,
default_config_name=dataset_module.builder_configs_parameters.default_config_name,
dataset_name=dataset_name,
)
return builder_cls
def files_to_hash(file_paths: List[str]) -> str:
"""
Convert a list of scripts or text files provided in file_paths into a hashed filename in a repeatable way.
"""
# List all python files in directories if directories are supplied as part of external imports
to_use_files: List[Union[Path, str]] = []
for file_path in file_paths:
if os.path.isdir(file_path):
to_use_files.extend(list(Path(file_path).rglob("*.[pP][yY]")))
else:
to_use_files.append(file_path)
# Get the code from all these files
lines = []
for file_path in to_use_files:
with open(file_path, encoding="utf-8") as f:
lines.extend(f.readlines())
return _hash_python_lines(lines)
def increase_load_count(name: str, resource_type: str):
"""Update the download count of a dataset or metric."""
if not config.HF_DATASETS_OFFLINE and config.HF_UPDATE_DOWNLOAD_COUNTS:
try:
head_hf_s3(name, filename=name + ".py", dataset=(resource_type == "dataset"))
except Exception:
pass
def _download_additional_modules(
name: str, base_path: str, imports: Tuple[str, str, str, str], download_config: Optional[DownloadConfig]
) -> List[Tuple[str, str]]:
"""
Download additional module for a module <name>.py at URL (or local path) <base_path>/<name>.py
The imports must have been parsed first using ``get_imports``.
If some modules need to be installed with pip, an error is raised showing how to install them.
This function return the list of downloaded modules as tuples (import_name, module_file_path).
The downloaded modules can then be moved into an importable directory with ``_copy_script_and_other_resources_in_importable_dir``.
"""
local_imports = []
library_imports = []
download_config = download_config.copy()
if download_config.download_desc is None:
download_config.download_desc = "Downloading extra modules"
for import_type, import_name, import_path, sub_directory in imports:
if import_type == "library":
library_imports.append((import_name, import_path)) # Import from a library
continue
if import_name == name:
raise ValueError(
f"Error in the {name} script, importing relative {import_name} module "
f"but {import_name} is the name of the script. "
f"Please change relative import {import_name} to another name and add a '# From: URL_OR_PATH' "
f"comment pointing to the original relative import file path."
)
if import_type == "internal":
url_or_filename = url_or_path_join(base_path, import_path + ".py")
elif import_type == "external":
url_or_filename = import_path
else:
raise ValueError("Wrong import_type")
local_import_path = cached_path(
url_or_filename,
download_config=download_config,
)
if sub_directory is not None:
local_import_path = os.path.join(local_import_path, sub_directory)
local_imports.append((import_name, local_import_path))
# Check library imports
needs_to_be_installed = {}
for library_import_name, library_import_path in library_imports:
try:
lib = importlib.import_module(library_import_name) # noqa F841
except ImportError:
if library_import_name not in needs_to_be_installed or library_import_path != library_import_name:
needs_to_be_installed[library_import_name] = library_import_path
if needs_to_be_installed:
_dependencies_str = "dependencies" if len(needs_to_be_installed) > 1 else "dependency"
_them_str = "them" if len(needs_to_be_installed) > 1 else "it"
if "sklearn" in needs_to_be_installed.keys():
needs_to_be_installed["sklearn"] = "scikit-learn"
raise ImportError(
f"To be able to use {name}, you need to install the following {_dependencies_str}: "
f"{', '.join(needs_to_be_installed)}.\nPlease install {_them_str} using 'pip install "
f"{' '.join(needs_to_be_installed.values())}' for instance."
)
return local_imports
def _copy_script_and_other_resources_in_importable_dir(
name: str,
importable_directory_path: str,
subdirectory_name: str,
original_local_path: str,
local_imports: List[Tuple[str, str]],
additional_files: List[Tuple[str, str]],
download_mode: Optional[Union[DownloadMode, str]],
) -> str:
"""Copy a script and its required imports to an importable directory
Args:
name (str): name of the resource to load
importable_directory_path (str): path to the loadable folder in the dynamic modules directory
subdirectory_name (str): name of the subdirectory in importable_directory_path in which to place the script
original_local_path (str): local path to the resource script
local_imports (List[Tuple[str, str]]): list of (destination_filename, import_file_to_copy)
additional_files (List[Tuple[str, str]]): list of (destination_filename, additional_file_to_copy)
download_mode (Optional[Union[DownloadMode, str]]): download mode
Return:
importable_local_file: path to an importable module with importlib.import_module
"""
# Define a directory with a unique name in our dataset or metric folder
# path is: ./datasets|metrics/dataset|metric_name/hash_from_code/script.py
# we use a hash as subdirectory_name to be able to have multiple versions of a dataset/metric processing file together
importable_subdirectory = os.path.join(importable_directory_path, subdirectory_name)
importable_local_file = os.path.join(importable_subdirectory, name + ".py")
# Prevent parallel disk operations
lock_path = importable_directory_path + ".lock"
with FileLock(lock_path):
# Create main dataset/metrics folder if needed
if download_mode == DownloadMode.FORCE_REDOWNLOAD and os.path.exists(importable_directory_path):
shutil.rmtree(importable_directory_path)
os.makedirs(importable_directory_path, exist_ok=True)
# add an __init__ file to the main dataset folder if needed
init_file_path = os.path.join(importable_directory_path, "__init__.py")
if not os.path.exists(init_file_path):
with open(init_file_path, "w"):
pass
# Create hash dataset folder if needed
os.makedirs(importable_subdirectory, exist_ok=True)
# add an __init__ file to the hash dataset folder if needed
init_file_path = os.path.join(importable_subdirectory, "__init__.py")
if not os.path.exists(init_file_path):
with open(init_file_path, "w"):
pass
# Copy dataset.py file in hash folder if needed
if not os.path.exists(importable_local_file):
shutil.copyfile(original_local_path, importable_local_file)
# Record metadata associating original dataset path with local unique folder
# Use os.path.splitext to split extension from importable_local_file
meta_path = os.path.splitext(importable_local_file)[0] + ".json"
if not os.path.exists(meta_path):
meta = {"original file path": original_local_path, "local file path": importable_local_file}
# the filename is *.py in our case, so better rename to filename.json instead of filename.py.json
with open(meta_path, "w", encoding="utf-8") as meta_file:
json.dump(meta, meta_file)
# Copy all the additional imports
for import_name, import_path in local_imports:
if os.path.isfile(import_path):
full_path_local_import = os.path.join(importable_subdirectory, import_name + ".py")
if not os.path.exists(full_path_local_import):
shutil.copyfile(import_path, full_path_local_import)
elif os.path.isdir(import_path):
full_path_local_import = os.path.join(importable_subdirectory, import_name)
if not os.path.exists(full_path_local_import):
shutil.copytree(import_path, full_path_local_import)
else:
raise ImportError(f"Error with local import at {import_path}")
# Copy additional files like dataset_infos.json file if needed
for file_name, original_path in additional_files:
destination_additional_path = os.path.join(importable_subdirectory, file_name)
if not os.path.exists(destination_additional_path) or not filecmp.cmp(
original_path, destination_additional_path
):
shutil.copyfile(original_path, destination_additional_path)
return importable_local_file
def _get_importable_file_path(
dynamic_modules_path: str,
module_namespace: str,
subdirectory_name: str,
name: str,
) -> str:
importable_directory_path = os.path.join(dynamic_modules_path, module_namespace, name.replace("/", "--"))
return os.path.join(importable_directory_path, subdirectory_name, name + ".py")
def _create_importable_file(
local_path: str,
local_imports: List[Tuple[str, str]],
additional_files: List[Tuple[str, str]],
dynamic_modules_path: str,
module_namespace: str,
subdirectory_name: str,
name: str,
download_mode: DownloadMode,
) -> None:
importable_directory_path = os.path.join(dynamic_modules_path, module_namespace, name.replace("/", "--"))
Path(importable_directory_path).mkdir(parents=True, exist_ok=True)
(Path(importable_directory_path).parent / "__init__.py").touch(exist_ok=True)
importable_local_file = _copy_script_and_other_resources_in_importable_dir(
name=name.split("/")[-1],
importable_directory_path=importable_directory_path,
subdirectory_name=subdirectory_name,
original_local_path=local_path,
local_imports=local_imports,
additional_files=additional_files,
download_mode=download_mode,
)
logger.debug(f"Created importable dataset file at {importable_local_file}")
def _load_importable_file(
dynamic_modules_path: str,
module_namespace: str,
subdirectory_name: str,
name: str,
) -> Tuple[str, str]:
module_path = ".".join(
[
os.path.basename(dynamic_modules_path),
module_namespace,
name.replace("/", "--"),
subdirectory_name,
name.split("/")[-1],
]
)
return module_path, subdirectory_name
def infer_module_for_data_files_list(
data_files_list: DataFilesList, download_config: Optional[DownloadConfig] = None
) -> Optional[Tuple[str, str]]:
"""Infer module (and builder kwargs) from list of data files.
It picks the module based on the most common file extension.
In case of a draw ".parquet" is the favorite, and then alphabetical order.
Args:
data_files_list (DataFilesList): List of data files.
download_config (bool or str, optional): mainly use use_auth_token or storage_options to support different platforms and auth types.
Returns:
tuple[str, dict[str, Any]]: Tuple with
- inferred module name
- dict of builder kwargs
"""
extensions_counter = Counter(
"." + suffix.lower()
for filepath in data_files_list[: config.DATA_FILES_MAX_NUMBER_FOR_MODULE_INFERENCE]
for suffix in xbasename(filepath).split(".")[1:]
)
if extensions_counter:
def sort_key(ext_count: Tuple[str, int]) -> Tuple[int, bool]:
"""Sort by count and set ".parquet" as the favorite in case of a draw"""
ext, count = ext_count
return (count, ext == ".parquet", ext)
for ext, _ in sorted(extensions_counter.items(), key=sort_key, reverse=True):
if ext in _EXTENSION_TO_MODULE:
return _EXTENSION_TO_MODULE[ext]
elif ext == ".zip":
return infer_module_for_data_files_list_in_archives(data_files_list, download_config=download_config)
return None, {}
def infer_module_for_data_files_list_in_archives(
data_files_list: DataFilesList, download_config: Optional[DownloadConfig] = None
) -> Optional[Tuple[str, str]]:
"""Infer module (and builder kwargs) from list of archive data files.
Args:
data_files_list (DataFilesList): List of data files.
download_config (bool or str, optional): mainly use use_auth_token or storage_options to support different platforms and auth types.
Returns:
tuple[str, dict[str, Any]]: Tuple with
- inferred module name
- dict of builder kwargs
"""
archived_files = []
archive_files_counter = 0
for filepath in data_files_list:
if str(filepath).endswith(".zip"):
archive_files_counter += 1
if archive_files_counter > config.GLOBBED_DATA_FILES_MAX_NUMBER_FOR_MODULE_INFERENCE:
break
extracted = xjoin(StreamingDownloadManager().extract(filepath), "**")
archived_files += [
f.split("::")[0]
for f in xglob(extracted, recursive=True, download_config=download_config)[
: config.ARCHIVED_DATA_FILES_MAX_NUMBER_FOR_MODULE_INFERENCE
]
]
extensions_counter = Counter(
"." + suffix.lower() for filepath in archived_files for suffix in xbasename(filepath).split(".")[1:]
)
if extensions_counter:
most_common = extensions_counter.most_common(1)[0][0]
if most_common in _EXTENSION_TO_MODULE:
return _EXTENSION_TO_MODULE[most_common]
return None, {}
def infer_module_for_data_files(
data_files: DataFilesDict, path: Optional[str] = None, download_config: Optional[DownloadConfig] = None
) -> Tuple[Optional[str], Dict[str, Any]]:
"""Infer module (and builder kwargs) from data files. Raise if module names for different splits don't match.
Args:
data_files ([`DataFilesDict`]): Dict of list of data files.
path (str, *optional*): Dataset name or path.
download_config ([`DownloadConfig`], *optional*):
Specific download configuration parameters to authenticate on the Hugging Face Hub for private remote files.
Returns:
tuple[str, dict[str, Any]]: Tuple with
- inferred module name
- builder kwargs
"""
split_modules = {
split: infer_module_for_data_files_list(data_files_list, download_config=download_config)
for split, data_files_list in data_files.items()
}
module_name, default_builder_kwargs = next(iter(split_modules.values()))
if any((module_name, default_builder_kwargs) != split_module for split_module in split_modules.values()):
raise ValueError(f"Couldn't infer the same data file format for all splits. Got {split_modules}")
if not module_name:
raise DataFilesNotFoundError("No (supported) data files found" + (f" in {path}" if path else ""))
return module_name, default_builder_kwargs
def update_hash_with_config_parameters(hash: str, config_parameters: dict) -> str:
"""
Used to update hash of packaged modules which is used for creating unique cache directories to reflect
different config parameters which are passed in metadata from readme.
"""
params_to_exclude = {"config_name", "version", "description"}
params_to_add_to_hash = {
param: value for param, value in sorted(config_parameters.items()) if param not in params_to_exclude
}
m = Hasher()
m.update(hash)
m.update(params_to_add_to_hash)
return m.hexdigest()
def create_builder_configs_from_metadata_configs(
module_path: str,
metadata_configs: MetadataConfigs,
supports_metadata: bool,
base_path: Optional[str] = None,
default_builder_kwargs: Dict[str, Any] = None,
download_config: Optional[DownloadConfig] = None,
) -> Tuple[List[BuilderConfig], str]:
builder_cls = import_main_class(module_path)
builder_config_cls = builder_cls.BUILDER_CONFIG_CLASS
default_config_name = metadata_configs.get_default_config_name()
builder_configs = []
base_path = base_path if base_path is not None else ""
for config_name, config_params in metadata_configs.items():
config_data_files = config_params.get("data_files")
config_data_dir = config_params.get("data_dir")
config_base_path = base_path + "/" + config_data_dir if config_data_dir else base_path
try:
config_patterns = (
sanitize_patterns(config_data_files)
if config_data_files is not None
else get_data_patterns(config_base_path)
)
config_data_files_dict = DataFilesDict.from_patterns(
config_patterns,
base_path=config_base_path,
allowed_extensions=ALL_ALLOWED_EXTENSIONS,
download_config=download_config,
)
except EmptyDatasetError as e:
raise EmptyDatasetError(
f"Dataset at '{base_path}' doesn't contain data files matching the patterns for config '{config_name}',"
f" check `data_files` and `data_fir` parameters in the `configs` YAML field in README.md. "
) from e
if config_data_files is None and supports_metadata and config_patterns != DEFAULT_PATTERNS_ALL:
try:
config_metadata_patterns = get_metadata_patterns(base_path)
except FileNotFoundError:
config_metadata_patterns = None
if config_metadata_patterns is not None:
config_metadata_data_files_list = DataFilesList.from_patterns(
config_metadata_patterns, base_path=base_path
)
if config_metadata_data_files_list:
config_data_files_dict = DataFilesDict(
{
split: data_files_list + config_metadata_data_files_list
for split, data_files_list in config_data_files_dict.items()
}
)
ignored_params = [
param for param in config_params if not hasattr(builder_config_cls, param) and param != "default"
]
if ignored_params:
logger.warning(
f"Some datasets params were ignored: {ignored_params}. "
"Make sure to use only valid params for the dataset builder and to have "
"a up-to-date version of the `datasets` library."
)
builder_configs.append(
builder_config_cls(
name=config_name,
data_files=config_data_files_dict,
data_dir=config_data_dir,
**{
param: value
for param, value in {**default_builder_kwargs, **config_params}.items()
if hasattr(builder_config_cls, param) and param not in ("default", "data_files", "data_dir")
},
)
)
return builder_configs, default_config_name
@dataclass
class BuilderConfigsParameters:
"""Dataclass containing objects related to creation of builder configurations from yaml's metadata content.
Attributes:
metadata_configs (`MetadataConfigs`, *optional*):
Configs parsed from yaml's metadata.
builder_configs (`list[BuilderConfig]`, *optional*):
List of BuilderConfig objects created from metadata_configs above.
default_config_name (`str`):
Name of default config taken from yaml's metadata.
"""
metadata_configs: Optional[MetadataConfigs] = None
builder_configs: Optional[List[BuilderConfig]] = None
default_config_name: Optional[str] = None
@dataclass
class DatasetModule:
module_path: str
hash: str
builder_kwargs: dict
builder_configs_parameters: BuilderConfigsParameters = field(default_factory=BuilderConfigsParameters)
dataset_infos: Optional[DatasetInfosDict] = None
@dataclass
class MetricModule:
module_path: str
hash: str
class _DatasetModuleFactory:
def get_module(self) -> DatasetModule:
raise NotImplementedError
class _MetricModuleFactory:
def get_module(self) -> MetricModule:
raise NotImplementedError
class GithubMetricModuleFactory(_MetricModuleFactory):
"""Get the module of a metric. The metric script is downloaded from GitHub.
<Deprecated version="2.5.0">
Use the new library 🤗 Evaluate instead: https://huggingface.co/docs/evaluate
</Deprecated>
"""
@deprecated("Use the new library 🤗 Evaluate instead: https://huggingface.co/docs/evaluate")
def __init__(
self,
name: str,
revision: Optional[Union[str, Version]] = None,
download_config: Optional[DownloadConfig] = None,
download_mode: Optional[Union[DownloadMode, str]] = None,
dynamic_modules_path: Optional[str] = None,
trust_remote_code: Optional[str] = None,
):
self.name = name
self.revision = revision
self.download_config = download_config.copy() if download_config else DownloadConfig()
if self.download_config.max_retries < 3:
self.download_config.max_retries = 3
self.download_mode = download_mode
self.dynamic_modules_path = dynamic_modules_path
self.trust_remote_code = trust_remote_code
assert self.name.count("/") == 0
increase_load_count(name, resource_type="metric")
def download_loading_script(self, revision: Optional[str]) -> str:
file_path = hf_github_url(path=self.name, name=self.name + ".py", revision=revision, dataset=False)
download_config = self.download_config.copy()
if download_config.download_desc is None:
download_config.download_desc = "Downloading builder script"
return cached_path(file_path, download_config=download_config)
def get_module(self) -> MetricModule:
if config.HF_DATASETS_TRUST_REMOTE_CODE and self.trust_remote_code is None:
_loading_script_url = hf_github_url(
path=self.name, name=self.name + ".py", revision=self.revision, dataset=False
)
warnings.warn(
f"The repository for {self.name} contains custom code which must be executed to correctly "
f"load the metric. You can inspect the repository content at {_loading_script_url}\n"
f"You can avoid this message in future by passing the argument `trust_remote_code=True`.\n"
f"Passing `trust_remote_code=True` will be mandatory to load this metric from the next major release of `datasets`.",
FutureWarning,
)
# get script and other files
revision = self.revision
try:
local_path = self.download_loading_script(revision)
revision = self.revision
except FileNotFoundError:
if revision is not None:
raise
else:
revision = "main"
local_path = self.download_loading_script(revision)
logger.warning(
f"Couldn't find a directory or a metric named '{self.name}' in this version. "
f"It was picked from the main branch on github instead."
)
imports = get_imports(local_path)
local_imports = _download_additional_modules(
name=self.name,
base_path=hf_github_url(path=self.name, name="", revision=revision, dataset=False),
imports=imports,
download_config=self.download_config,
)
# copy the script and the files in an importable directory
dynamic_modules_path = self.dynamic_modules_path if self.dynamic_modules_path else init_dynamic_modules()
hash = files_to_hash([local_path] + [loc[1] for loc in local_imports])
importable_file_path = _get_importable_file_path(
dynamic_modules_path=dynamic_modules_path,
module_namespace="metrics",
subdirectory_name=hash,
name=self.name,
)
if not os.path.exists(importable_file_path):
trust_remote_code = resolve_trust_remote_code(self.trust_remote_code, self.name)
if trust_remote_code:
_create_importable_file(
local_path=local_path,
local_imports=local_imports,
additional_files=[],
dynamic_modules_path=dynamic_modules_path,
module_namespace="metrics",
subdirectory_name=hash,
name=self.name,
download_mode=self.download_mode,
)
else:
raise ValueError(
f"Loading {self.name} requires you to execute the dataset script in that"
" repo on your local machine. Make sure you have read the code there to avoid malicious use, then"
" set the option `trust_remote_code=True` to remove this error."
)
module_path, hash = _load_importable_file(
dynamic_modules_path=dynamic_modules_path,
module_namespace="metrics",
subdirectory_name=hash,
name=self.name,
)
# make the new module to be noticed by the import system
importlib.invalidate_caches()
return MetricModule(module_path, hash)
class LocalMetricModuleFactory(_MetricModuleFactory):
"""Get the module of a local metric. The metric script is loaded from a local script.
<Deprecated version="2.5.0">
Use the new library 🤗 Evaluate instead: https://huggingface.co/docs/evaluate
</Deprecated>
"""
@deprecated("Use the new library 🤗 Evaluate instead: https://huggingface.co/docs/evaluate")
def __init__(
self,
path: str,
download_config: Optional[DownloadConfig] = None,
download_mode: Optional[Union[DownloadMode, str]] = None,
dynamic_modules_path: Optional[str] = None,
trust_remote_code: Optional[str] = None,
):
self.path = path
self.name = Path(path).stem
self.download_config = download_config or DownloadConfig()
self.download_mode = download_mode
self.dynamic_modules_path = dynamic_modules_path
self.trust_remote_code = trust_remote_code
def get_module(self) -> MetricModule:
if config.HF_DATASETS_TRUST_REMOTE_CODE and self.trust_remote_code is None:
warnings.warn(
f"The repository for {self.name} contains custom code which must be executed to correctly "
f"load the metric. You can inspect the repository content at {self.path}\n"
f"You can avoid this message in future by passing the argument `trust_remote_code=True`.\n"
f"Passing `trust_remote_code=True` will be mandatory to load this metric from the next major release of `datasets`.",
FutureWarning,
)
# get script and other files
imports = get_imports(self.path)
local_imports = _download_additional_modules(
name=self.name,
base_path=str(Path(self.path).parent),
imports=imports,
download_config=self.download_config,
)
# copy the script and the files in an importable directory
dynamic_modules_path = self.dynamic_modules_path if self.dynamic_modules_path else init_dynamic_modules()
hash = files_to_hash([self.path] + [loc[1] for loc in local_imports])
importable_file_path = _get_importable_file_path(
dynamic_modules_path=dynamic_modules_path,
module_namespace="metrics",
subdirectory_name=hash,
name=self.name,
)
if not os.path.exists(importable_file_path):
trust_remote_code = resolve_trust_remote_code(self.trust_remote_code, self.name)
if trust_remote_code:
_create_importable_file(
local_path=self.path,
local_imports=local_imports,
additional_files=[],
dynamic_modules_path=dynamic_modules_path,
module_namespace="metrics",
subdirectory_name=hash,
name=self.name,
download_mode=self.download_mode,
)
else:
raise ValueError(
f"Loading {self.name} requires you to execute the dataset script in that"
" repo on your local machine. Make sure you have read the code there to avoid malicious use, then"
" set the option `trust_remote_code=True` to remove this error."
)
module_path, hash = _load_importable_file(
dynamic_modules_path=dynamic_modules_path,
module_namespace="metrics",
subdirectory_name=hash,
name=self.name,
)
# make the new module to be noticed by the import system
importlib.invalidate_caches()
return MetricModule(module_path, hash)
class LocalDatasetModuleFactoryWithScript(_DatasetModuleFactory):
"""Get the module of a local dataset. The dataset script is loaded from a local script."""
def __init__(
self,
path: str,
download_config: Optional[DownloadConfig] = None,
download_mode: Optional[Union[DownloadMode, str]] = None,
dynamic_modules_path: Optional[str] = None,
trust_remote_code: Optional[bool] = None,
):
self.path = path
self.name = Path(path).stem
self.download_config = download_config or DownloadConfig()
self.download_mode = download_mode
self.dynamic_modules_path = dynamic_modules_path
self.trust_remote_code = trust_remote_code
def get_module(self) -> DatasetModule:
if config.HF_DATASETS_TRUST_REMOTE_CODE and self.trust_remote_code is None:
warnings.warn(
f"The repository for {self.name} contains custom code which must be executed to correctly "
f"load the dataset. You can inspect the repository content at {self.path}\n"
f"You can avoid this message in future by passing the argument `trust_remote_code=True`.\n"
f"Passing `trust_remote_code=True` will be mandatory to load this dataset from the next major release of `datasets`.",
FutureWarning,
)
# get script and other files
dataset_infos_path = Path(self.path).parent / config.DATASETDICT_INFOS_FILENAME
dataset_readme_path = Path(self.path).parent / "README.md"
imports = get_imports(self.path)
local_imports = _download_additional_modules(
name=self.name,
base_path=str(Path(self.path).parent),
imports=imports,
download_config=self.download_config,
)
additional_files = []
if dataset_infos_path.is_file():
additional_files.append((config.DATASETDICT_INFOS_FILENAME, str(dataset_infos_path)))
if dataset_readme_path.is_file():
additional_files.append(("README.md", dataset_readme_path))
# copy the script and the files in an importable directory
dynamic_modules_path = self.dynamic_modules_path if self.dynamic_modules_path else init_dynamic_modules()
hash = files_to_hash([self.path] + [loc[1] for loc in local_imports])
importable_file_path = _get_importable_file_path(
dynamic_modules_path=dynamic_modules_path,
module_namespace="datasets",
subdirectory_name=hash,
name=self.name,
)
if not os.path.exists(importable_file_path):
trust_remote_code = resolve_trust_remote_code(self.trust_remote_code, self.name)
if trust_remote_code:
_create_importable_file(
local_path=self.path,
local_imports=local_imports,
additional_files=additional_files,
dynamic_modules_path=dynamic_modules_path,
module_namespace="datasets",
subdirectory_name=hash,
name=self.name,
download_mode=self.download_mode,
)
else:
raise ValueError(
f"Loading {self.name} requires you to execute the dataset script in that"
" repo on your local machine. Make sure you have read the code there to avoid malicious use, then"
" set the option `trust_remote_code=True` to remove this error."
)
module_path, hash = _load_importable_file(
dynamic_modules_path=dynamic_modules_path,
module_namespace="datasets",
subdirectory_name=hash,
name=self.name,
)
# make the new module to be noticed by the import system
importlib.invalidate_caches()
builder_kwargs = {"hash": hash, "base_path": str(Path(self.path).parent)}
return DatasetModule(module_path, hash, builder_kwargs)
class LocalDatasetModuleFactoryWithoutScript(_DatasetModuleFactory):
"""Get the module of a dataset loaded from the user's data files. The dataset builder module to use is inferred
from the data files extensions."""
def __init__(
self,
path: str,
data_dir: Optional[str] = None,
data_files: Optional[Union[str, List, Dict]] = None,
download_mode: Optional[Union[DownloadMode, str]] = None,
):
if data_dir and os.path.isabs(data_dir):
raise ValueError(f"`data_dir` must be relative to a dataset directory's root: {path}")
self.path = Path(path).as_posix()
self.name = Path(path).stem
self.data_files = data_files
self.data_dir = data_dir
self.download_mode = download_mode
def get_module(self) -> DatasetModule:
readme_path = os.path.join(self.path, "README.md")
dataset_card_data = DatasetCard.load(readme_path).data if os.path.isfile(readme_path) else DatasetCardData()
metadata_configs = MetadataConfigs.from_dataset_card_data(dataset_card_data)
dataset_infos = DatasetInfosDict.from_dataset_card_data(dataset_card_data)
# we need a set of data files to find which dataset builder to use
# because we need to infer module name by files extensions
base_path = Path(self.path, self.data_dir or "").expanduser().resolve().as_posix()
if self.data_files is not None:
patterns = sanitize_patterns(self.data_files)
elif metadata_configs and "data_files" in next(iter(metadata_configs.values())):
patterns = sanitize_patterns(next(iter(metadata_configs.values()))["data_files"])
else:
patterns = get_data_patterns(base_path)
data_files = DataFilesDict.from_patterns(
patterns,
base_path=base_path,
allowed_extensions=ALL_ALLOWED_EXTENSIONS,
)
module_name, default_builder_kwargs = infer_module_for_data_files(
data_files=data_files,
path=self.path,
)
data_files = data_files.filter_extensions(_MODULE_TO_EXTENSIONS[module_name])
# Collect metadata files if the module supports them
supports_metadata = module_name in _MODULE_SUPPORTS_METADATA
if self.data_files is None and supports_metadata:
try:
metadata_patterns = get_metadata_patterns(base_path)
except FileNotFoundError:
metadata_patterns = None
if metadata_patterns is not None:
metadata_data_files_list = DataFilesList.from_patterns(metadata_patterns, base_path=base_path)
if metadata_data_files_list:
data_files = DataFilesDict(
{
split: data_files_list + metadata_data_files_list
for split, data_files_list in data_files.items()
}
)
module_path, hash = _PACKAGED_DATASETS_MODULES[module_name]
if metadata_configs:
builder_configs, default_config_name = create_builder_configs_from_metadata_configs(
module_path,
metadata_configs,
base_path=base_path,
supports_metadata=supports_metadata,
default_builder_kwargs=default_builder_kwargs,
)
else:
builder_configs, default_config_name = None, None
builder_kwargs = {
"hash": hash,
"base_path": self.path,
"dataset_name": camelcase_to_snakecase(Path(self.path).name),
}
if self.data_files is not None or not metadata_configs:
builder_kwargs["data_files"] = data_files
builder_kwargs.update(default_builder_kwargs) # from _EXTENSION_TO_MODULE
# this file is deprecated and was created automatically in old versions of push_to_hub
if os.path.isfile(os.path.join(self.path, config.DATASETDICT_INFOS_FILENAME)):
with open(os.path.join(self.path, config.DATASETDICT_INFOS_FILENAME), encoding="utf-8") as f:
legacy_dataset_infos = DatasetInfosDict(
{
config_name: DatasetInfo.from_dict(dataset_info_dict)
for config_name, dataset_info_dict in json.load(f).items()
}
)
if len(legacy_dataset_infos) == 1:
# old config e.g. named "username--dataset_name"
legacy_config_name = next(iter(legacy_dataset_infos))
legacy_dataset_infos["default"] = legacy_dataset_infos.pop(legacy_config_name)
legacy_dataset_infos.update(dataset_infos)
dataset_infos = legacy_dataset_infos
if default_config_name is None and len(dataset_infos) == 1:
default_config_name = next(iter(dataset_infos))
return DatasetModule(
module_path,
hash,
builder_kwargs,
dataset_infos=dataset_infos,
builder_configs_parameters=BuilderConfigsParameters(
metadata_configs=metadata_configs,
builder_configs=builder_configs,
default_config_name=default_config_name,
),
)
class PackagedDatasetModuleFactory(_DatasetModuleFactory):
"""Get the dataset builder module from the ones that are packaged with the library: csv, json, etc."""
def __init__(
self,
name: str,
data_dir: Optional[str] = None,
data_files: Optional[Union[str, List, Dict]] = None,
download_config: Optional[DownloadConfig] = None,
download_mode: Optional[Union[DownloadMode, str]] = None,
):
self.name = name
self.data_files = data_files
self.data_dir = data_dir
self.download_config = download_config
self.download_mode = download_mode
increase_load_count(name, resource_type="dataset")
def get_module(self) -> DatasetModule:
base_path = Path(self.data_dir or "").expanduser().resolve().as_posix()
patterns = sanitize_patterns(self.data_files) if self.data_files is not None else get_data_patterns(base_path)
data_files = DataFilesDict.from_patterns(
patterns,
download_config=self.download_config,
base_path=base_path,
)
supports_metadata = self.name in _MODULE_SUPPORTS_METADATA
if self.data_files is None and supports_metadata and patterns != DEFAULT_PATTERNS_ALL:
try:
metadata_patterns = get_metadata_patterns(base_path)
except FileNotFoundError:
metadata_patterns = None
if metadata_patterns is not None:
metadata_data_files_list = DataFilesList.from_patterns(
metadata_patterns, download_config=self.download_config, base_path=base_path
)
if metadata_data_files_list:
data_files = DataFilesDict(
{
split: data_files_list + metadata_data_files_list
for split, data_files_list in data_files.items()
}
)
module_path, hash = _PACKAGED_DATASETS_MODULES[self.name]
builder_kwargs = {
"hash": hash,
"data_files": data_files,
"dataset_name": self.name,
}
return DatasetModule(module_path, hash, builder_kwargs)
class HubDatasetModuleFactoryWithoutScript(_DatasetModuleFactory):
"""
Get the module of a dataset loaded from data files of a dataset repository.
The dataset builder module to use is inferred from the data files extensions.
"""
def __init__(
self,
name: str,
revision: Optional[Union[str, Version]] = None,
data_dir: Optional[str] = None,
data_files: Optional[Union[str, List, Dict]] = None,
download_config: Optional[DownloadConfig] = None,
download_mode: Optional[Union[DownloadMode, str]] = None,
):
self.name = name
self.revision = revision
self.data_files = data_files
self.data_dir = data_dir
self.download_config = download_config or DownloadConfig()
self.download_mode = download_mode
increase_load_count(name, resource_type="dataset")
def get_module(self) -> DatasetModule:
hfh_dataset_info = HfApi(config.HF_ENDPOINT).dataset_info(
self.name,
revision=self.revision,
token=self.download_config.token,
timeout=100.0,
)
# even if metadata_configs is not None (which means that we will resolve files for each config later)
# we cannot skip resolving all files because we need to infer module name by files extensions
revision = hfh_dataset_info.sha # fix the revision in case there are new commits in the meantime
base_path = f"hf://datasets/{self.name}@{revision}/{self.data_dir or ''}".rstrip("/")
download_config = self.download_config.copy()
if download_config.download_desc is None:
download_config.download_desc = "Downloading readme"
try:
dataset_readme_path = cached_path(
hf_hub_url(self.name, "README.md", revision=revision),
download_config=download_config,
)
dataset_card_data = DatasetCard.load(Path(dataset_readme_path)).data
except FileNotFoundError:
dataset_card_data = DatasetCardData()
metadata_configs = MetadataConfigs.from_dataset_card_data(dataset_card_data)
dataset_infos = DatasetInfosDict.from_dataset_card_data(dataset_card_data)
# we need a set of data files to find which dataset builder to use
# because we need to infer module name by files extensions
if self.data_files is not None:
patterns = sanitize_patterns(self.data_files)
elif metadata_configs and "data_files" in next(iter(metadata_configs.values())):
patterns = sanitize_patterns(next(iter(metadata_configs.values()))["data_files"])
else:
patterns = get_data_patterns(base_path, download_config=self.download_config)
data_files = DataFilesDict.from_patterns(
patterns,
base_path=base_path,
allowed_extensions=ALL_ALLOWED_EXTENSIONS,
download_config=self.download_config,
)
module_name, default_builder_kwargs = infer_module_for_data_files(
data_files=data_files,
path=self.name,
download_config=self.download_config,
)
data_files = data_files.filter_extensions(_MODULE_TO_EXTENSIONS[module_name])
# Collect metadata files if the module supports them
supports_metadata = module_name in _MODULE_SUPPORTS_METADATA
if self.data_files is None and supports_metadata:
try:
metadata_patterns = get_metadata_patterns(base_path)
except FileNotFoundError:
metadata_patterns = None
if metadata_patterns is not None:
metadata_data_files_list = DataFilesList.from_patterns(
metadata_patterns, download_config=self.download_config, base_path=base_path
)
if metadata_data_files_list:
data_files = DataFilesDict(
{
split: data_files_list + metadata_data_files_list
for split, data_files_list in data_files.items()
}
)
module_path, hash = _PACKAGED_DATASETS_MODULES[module_name]
if metadata_configs:
builder_configs, default_config_name = create_builder_configs_from_metadata_configs(
module_path,
metadata_configs,
base_path=base_path,
supports_metadata=supports_metadata,
default_builder_kwargs=default_builder_kwargs,
download_config=self.download_config,
)
else:
builder_configs, default_config_name = None, None
builder_kwargs = {
"hash": hash,
"base_path": hf_hub_url(self.name, "", revision=self.revision),
"repo_id": self.name,
"dataset_name": camelcase_to_snakecase(Path(self.name).name),
}
if self.data_files is not None or not metadata_configs:
builder_kwargs["data_files"] = data_files
builder_kwargs.update(default_builder_kwargs) # from _EXTENSION_TO_MODULE
download_config = self.download_config.copy()
if download_config.download_desc is None:
download_config.download_desc = "Downloading metadata"
try:
# this file is deprecated and was created automatically in old versions of push_to_hub
dataset_infos_path = cached_path(
hf_hub_url(self.name, config.DATASETDICT_INFOS_FILENAME, revision=self.revision),
download_config=download_config,
)
with open(dataset_infos_path, encoding="utf-8") as f:
legacy_dataset_infos = DatasetInfosDict(
{
config_name: DatasetInfo.from_dict(dataset_info_dict)
for config_name, dataset_info_dict in json.load(f).items()
}
)
if len(legacy_dataset_infos) == 1:
# old config e.g. named "username--dataset_name"
legacy_config_name = next(iter(legacy_dataset_infos))
legacy_dataset_infos["default"] = legacy_dataset_infos.pop(legacy_config_name)
legacy_dataset_infos.update(dataset_infos)
dataset_infos = legacy_dataset_infos
except FileNotFoundError:
pass
if default_config_name is None and len(dataset_infos) == 1:
default_config_name = next(iter(dataset_infos))
return DatasetModule(
module_path,
hash,
builder_kwargs,
dataset_infos=dataset_infos,
builder_configs_parameters=BuilderConfigsParameters(
metadata_configs=metadata_configs,
builder_configs=builder_configs,
default_config_name=default_config_name,
),
)
class HubDatasetModuleFactoryWithScript(_DatasetModuleFactory):
"""
Get the module of a dataset from a dataset repository.
The dataset script comes from the script inside the dataset repository.
"""
def __init__(
self,
name: str,
revision: Optional[Union[str, Version]] = None,
download_config: Optional[DownloadConfig] = None,
download_mode: Optional[Union[DownloadMode, str]] = None,
dynamic_modules_path: Optional[str] = None,
trust_remote_code: Optional[bool] = None,
):
self.name = name
self.revision = revision
self.download_config = download_config or DownloadConfig()
self.download_mode = download_mode
self.dynamic_modules_path = dynamic_modules_path
self.trust_remote_code = trust_remote_code
increase_load_count(name, resource_type="dataset")
def download_loading_script(self) -> str:
file_path = hf_hub_url(repo_id=self.name, path=self.name.split("/")[-1] + ".py", revision=self.revision)
download_config = self.download_config.copy()
if download_config.download_desc is None:
download_config.download_desc = "Downloading builder script"
return cached_path(file_path, download_config=download_config)
def download_dataset_infos_file(self) -> str:
dataset_infos = hf_hub_url(repo_id=self.name, path=config.DATASETDICT_INFOS_FILENAME, revision=self.revision)
# Download the dataset infos file if available
download_config = self.download_config.copy()
if download_config.download_desc is None:
download_config.download_desc = "Downloading metadata"
try:
return cached_path(
dataset_infos,
download_config=download_config,
)
except (FileNotFoundError, ConnectionError):
return None
def download_dataset_readme_file(self) -> str:
readme_url = hf_hub_url(repo_id=self.name, path="README.md", revision=self.revision)
# Download the dataset infos file if available
download_config = self.download_config.copy()
if download_config.download_desc is None:
download_config.download_desc = "Downloading readme"
try:
return cached_path(
readme_url,
download_config=download_config,
)
except (FileNotFoundError, ConnectionError):
return None
def get_module(self) -> DatasetModule:
if config.HF_DATASETS_TRUST_REMOTE_CODE and self.trust_remote_code is None:
warnings.warn(
f"The repository for {self.name} contains custom code which must be executed to correctly "
f"load the dataset. You can inspect the repository content at https://hf.co/datasets/{self.name}\n"
f"You can avoid this message in future by passing the argument `trust_remote_code=True`.\n"
f"Passing `trust_remote_code=True` will be mandatory to load this dataset from the next major release of `datasets`.",
FutureWarning,
)
# get script and other files
local_path = self.download_loading_script()
dataset_infos_path = self.download_dataset_infos_file()
dataset_readme_path = self.download_dataset_readme_file()
imports = get_imports(local_path)
local_imports = _download_additional_modules(
name=self.name,
base_path=hf_hub_url(repo_id=self.name, path="", revision=self.revision),
imports=imports,
download_config=self.download_config,
)
additional_files = []
if dataset_infos_path:
additional_files.append((config.DATASETDICT_INFOS_FILENAME, dataset_infos_path))
if dataset_readme_path:
additional_files.append(("README.md", dataset_readme_path))
# copy the script and the files in an importable directory
dynamic_modules_path = self.dynamic_modules_path if self.dynamic_modules_path else init_dynamic_modules()
hash = files_to_hash([local_path] + [loc[1] for loc in local_imports])
importable_file_path = _get_importable_file_path(
dynamic_modules_path=dynamic_modules_path,
module_namespace="datasets",
subdirectory_name=hash,
name=self.name,
)
if not os.path.exists(importable_file_path):
trust_remote_code = resolve_trust_remote_code(self.trust_remote_code, self.name)
if trust_remote_code:
_create_importable_file(
local_path=local_path,
local_imports=local_imports,
additional_files=additional_files,
dynamic_modules_path=dynamic_modules_path,
module_namespace="datasets",
subdirectory_name=hash,
name=self.name,
download_mode=self.download_mode,
)
else:
raise ValueError(
f"Loading {self.name} requires you to execute the dataset script in that"
" repo on your local machine. Make sure you have read the code there to avoid malicious use, then"
" set the option `trust_remote_code=True` to remove this error."
)
module_path, hash = _load_importable_file(
dynamic_modules_path=dynamic_modules_path,
module_namespace="datasets",
subdirectory_name=hash,
name=self.name,
)
# make the new module to be noticed by the import system
importlib.invalidate_caches()
builder_kwargs = {
"hash": hash,
"base_path": hf_hub_url(self.name, "", revision=self.revision),
"repo_id": self.name,
}
return DatasetModule(module_path, hash, builder_kwargs)
class CachedDatasetModuleFactory(_DatasetModuleFactory):
"""
Get the module of a dataset that has been loaded once already and cached.
The script that is loaded from the cache is the most recent one with a matching name.
"""
def __init__(
self,
name: str,
dynamic_modules_path: Optional[str] = None,
):
self.name = name
self.dynamic_modules_path = dynamic_modules_path
assert self.name.count("/") <= 1
def get_module(self) -> DatasetModule:
dynamic_modules_path = self.dynamic_modules_path if self.dynamic_modules_path else init_dynamic_modules()
importable_directory_path = os.path.join(dynamic_modules_path, "datasets", self.name.replace("/", "--"))
hashes = (
[h for h in os.listdir(importable_directory_path) if len(h) == 64]
if os.path.isdir(importable_directory_path)
else None
)
if not hashes:
raise FileNotFoundError(f"Dataset {self.name} is not cached in {dynamic_modules_path}")
# get most recent
def _get_modification_time(module_hash):
return (Path(importable_directory_path) / module_hash / (self.name.split("/")[-1] + ".py")).stat().st_mtime
hash = sorted(hashes, key=_get_modification_time)[-1]
warning_msg = (
f"Using the latest cached version of the module from {os.path.join(importable_directory_path, hash)} "
f"(last modified on {time.ctime(_get_modification_time(hash))}) since it "
f"couldn't be found locally at {self.name}."
)
if not config.HF_DATASETS_OFFLINE:
warning_msg += ", or remotely on the Hugging Face Hub."
logger.warning(warning_msg)
# make the new module to be noticed by the import system
module_path = ".".join(
[
os.path.basename(dynamic_modules_path),
"datasets",
self.name.replace("/", "--"),
hash,
self.name.split("/")[-1],
]
)
importlib.invalidate_caches()
builder_kwargs = {
"hash": hash,
"repo_id": self.name,
}
return DatasetModule(module_path, hash, builder_kwargs)
class CachedMetricModuleFactory(_MetricModuleFactory):
"""
Get the module of a metric that has been loaded once already and cached.
The script that is loaded from the cache is the most recent one with a matching name.
<Deprecated version="2.5.0">
Use the new library 🤗 Evaluate instead: https://huggingface.co/docs/evaluate
</Deprecated>
"""
@deprecated("Use the new library 🤗 Evaluate instead: https://huggingface.co/docs/evaluate")
def __init__(
self,
name: str,
dynamic_modules_path: Optional[str] = None,
):
self.name = name
self.dynamic_modules_path = dynamic_modules_path
assert self.name.count("/") == 0
def get_module(self) -> MetricModule:
dynamic_modules_path = self.dynamic_modules_path if self.dynamic_modules_path else init_dynamic_modules()
importable_directory_path = os.path.join(dynamic_modules_path, "metrics", self.name)
hashes = (
[h for h in os.listdir(importable_directory_path) if len(h) == 64]
if os.path.isdir(importable_directory_path)
else None
)
if not hashes:
raise FileNotFoundError(f"Metric {self.name} is not cached in {dynamic_modules_path}")
# get most recent
def _get_modification_time(module_hash):
return (Path(importable_directory_path) / module_hash / (self.name + ".py")).stat().st_mtime
hash = sorted(hashes, key=_get_modification_time)[-1]
logger.warning(
f"Using the latest cached version of the module from {os.path.join(importable_directory_path, hash)} "
f"(last modified on {time.ctime(_get_modification_time(hash))}) since it "
f"couldn't be found locally at {self.name}, or remotely on the Hugging Face Hub."
)
# make the new module to be noticed by the import system
module_path = ".".join([os.path.basename(dynamic_modules_path), "metrics", self.name, hash, self.name])
importlib.invalidate_caches()
return MetricModule(module_path, hash)
def dataset_module_factory(
path: str,
revision: Optional[Union[str, Version]] = None,
download_config: Optional[DownloadConfig] = None,
download_mode: Optional[Union[DownloadMode, str]] = None,
dynamic_modules_path: Optional[str] = None,
data_dir: Optional[str] = None,
data_files: Optional[Union[Dict, List, str, DataFilesDict]] = None,
trust_remote_code: Optional[bool] = None,
**download_kwargs,
) -> DatasetModule:
"""
Download/extract/cache a dataset module.
Dataset codes are cached inside the dynamic modules cache to allow easy import (avoid ugly sys.path tweaks).
Args:
path (str): Path or name of the dataset.
Depending on ``path``, the dataset builder that is used comes from a generic dataset script (JSON, CSV, Parquet, text etc.) or from the dataset script (a python file) inside the dataset directory.
For local datasets:
- if ``path`` is a local directory (containing data files only)
-> load a generic dataset builder (csv, json, text etc.) based on the content of the directory
e.g. ``'./path/to/directory/with/my/csv/data'``.
- if ``path`` is a local dataset script or a directory containing a local dataset script (if the script has the same name as the directory):
-> load the dataset builder from the dataset script
e.g. ``'./dataset/squad'`` or ``'./dataset/squad/squad.py'``.
For datasets on the Hugging Face Hub (list all available datasets with ``huggingface_hub.list_datasets()``)
- if ``path`` is a dataset repository on the HF hub (containing data files only)
-> load a generic dataset builder (csv, text etc.) based on the content of the repository
e.g. ``'username/dataset_name'``, a dataset repository on the HF hub containing your data files.
- if ``path`` is a dataset repository on the HF hub with a dataset script (if the script has the same name as the directory)
-> load the dataset builder from the dataset script in the dataset repository
e.g. ``glue``, ``squad``, ``'username/dataset_name'``, a dataset repository on the HF hub containing a dataset script `'dataset_name.py'`.
revision (:class:`~utils.Version` or :obj:`str`, optional): Version of the dataset script to load.
As datasets have their own git repository on the Datasets Hub, the default version "main" corresponds to their "main" branch.
You can specify a different version than the default "main" by using a commit SHA or a git tag of the dataset repository.
download_config (:class:`DownloadConfig`, optional): Specific download configuration parameters.
download_mode (:class:`DownloadMode` or :obj:`str`, default ``REUSE_DATASET_IF_EXISTS``): Download/generate mode.
dynamic_modules_path (Optional str, defaults to HF_MODULES_CACHE / "datasets_modules", i.e. ~/.cache/huggingface/modules/datasets_modules):
Optional path to the directory in which the dynamic modules are saved. It must have been initialized with :obj:`init_dynamic_modules`.
By default, the datasets and metrics are stored inside the `datasets_modules` module.
data_dir (:obj:`str`, optional): Directory with the data files. Used only if `data_files` is not specified,
in which case it's equal to pass `os.path.join(data_dir, "**")` as `data_files`.
data_files (:obj:`Union[Dict, List, str]`, optional): Defining the data_files of the dataset configuration.
trust_remote_code (`bool`, defaults to `True`):
Whether or not to allow for datasets defined on the Hub using a dataset script. This option
should only be set to `True` for repositories you trust and in which you have read the code, as it will
execute code present on the Hub on your local machine.
<Tip warning={true}>
`trust_remote_code` will default to False in the next major release.
</Tip>
<Added version="2.16.0"/>
**download_kwargs (additional keyword arguments): optional attributes for DownloadConfig() which will override
the attributes in download_config if supplied.
Returns:
DatasetModule
"""
if download_config is None:
download_config = DownloadConfig(**download_kwargs)
download_mode = DownloadMode(download_mode or DownloadMode.REUSE_DATASET_IF_EXISTS)
download_config.extract_compressed_file = True
download_config.force_extract = True
download_config.force_download = download_mode == DownloadMode.FORCE_REDOWNLOAD
filename = list(filter(lambda x: x, path.replace(os.sep, "/").split("/")))[-1]
if not filename.endswith(".py"):
filename = filename + ".py"
combined_path = os.path.join(path, filename)
# We have several ways to get a dataset builder:
#
# - if path is the name of a packaged dataset module
# -> use the packaged module (json, csv, etc.)
#
# - if os.path.join(path, name) is a local python file
# -> use the module from the python file
# - if path is a local directory (but no python file)
# -> use a packaged module (csv, text etc.) based on content of the directory
#
# - if path has one "/" and is dataset repository on the HF hub with a python file
# -> the module from the python file in the dataset repository
# - if path has one "/" and is dataset repository on the HF hub without a python file
# -> use a packaged module (csv, text etc.) based on content of the repository
# Try packaged
if path in _PACKAGED_DATASETS_MODULES:
return PackagedDatasetModuleFactory(
path,
data_dir=data_dir,
data_files=data_files,
download_config=download_config,
download_mode=download_mode,
).get_module()
# Try locally
elif path.endswith(filename):
if os.path.isfile(path):
return LocalDatasetModuleFactoryWithScript(
path,
download_mode=download_mode,
dynamic_modules_path=dynamic_modules_path,
trust_remote_code=trust_remote_code,
).get_module()
else:
raise FileNotFoundError(f"Couldn't find a dataset script at {relative_to_absolute_path(path)}")
elif os.path.isfile(combined_path):
return LocalDatasetModuleFactoryWithScript(
combined_path,
download_mode=download_mode,
dynamic_modules_path=dynamic_modules_path,
trust_remote_code=trust_remote_code,
).get_module()
elif os.path.isdir(path):
return LocalDatasetModuleFactoryWithoutScript(
path, data_dir=data_dir, data_files=data_files, download_mode=download_mode
).get_module()
# Try remotely
elif is_relative_path(path) and path.count("/") <= 1:
try:
_raise_if_offline_mode_is_enabled()
hf_api = HfApi(config.HF_ENDPOINT)
try:
dataset_info = hf_api.dataset_info(
repo_id=path,
revision=revision,
token=download_config.token,
timeout=100.0,
)
except Exception as e: # noqa catch any exception of hf_hub and consider that the dataset doesn't exist
if isinstance(
e,
(
OfflineModeIsEnabled,
requests.exceptions.ConnectTimeout,
requests.exceptions.ConnectionError,
),
):
raise ConnectionError(f"Couldn't reach '{path}' on the Hub ({type(e).__name__})")
elif "404" in str(e):
msg = f"Dataset '{path}' doesn't exist on the Hub"
raise FileNotFoundError(msg + f" at revision '{revision}'" if revision else msg)
elif "401" in str(e):
msg = f"Dataset '{path}' doesn't exist on the Hub"
msg = msg + f" at revision '{revision}'" if revision else msg
raise DatasetNotFoundError(
msg + ". If the repo is private or gated, make sure to log in with `huggingface-cli login`."
)
else:
raise e
if filename in [sibling.rfilename for sibling in dataset_info.siblings]:
return HubDatasetModuleFactoryWithScript(
path,
revision=revision,
download_config=download_config,
download_mode=download_mode,
dynamic_modules_path=dynamic_modules_path,
trust_remote_code=trust_remote_code,
).get_module()
else:
return HubDatasetModuleFactoryWithoutScript(
path,
revision=revision,
data_dir=data_dir,
data_files=data_files,
download_config=download_config,
download_mode=download_mode,
).get_module()
except Exception as e1:
# All the attempts failed, before raising the error we should check if the module is already cached
try:
return CachedDatasetModuleFactory(path, dynamic_modules_path=dynamic_modules_path).get_module()
except Exception:
# If it's not in the cache, then it doesn't exist.
if isinstance(e1, OfflineModeIsEnabled):
raise ConnectionError(f"Couldn't reach the Hugging Face Hub for dataset '{path}': {e1}") from None
if isinstance(e1, (DataFilesNotFoundError, DatasetNotFoundError, EmptyDatasetError)):
raise e1 from None
if isinstance(e1, FileNotFoundError):
raise FileNotFoundError(
f"Couldn't find a dataset script at {relative_to_absolute_path(combined_path)} or any data file in the same directory. "
f"Couldn't find '{path}' on the Hugging Face Hub either: {type(e1).__name__}: {e1}"
) from None
raise e1 from None
else:
raise FileNotFoundError(
f"Couldn't find a dataset script at {relative_to_absolute_path(combined_path)} or any data file in the same directory."
)
@deprecated("Use the new library 🤗 Evaluate instead: https://huggingface.co/docs/evaluate")
def metric_module_factory(
path: str,
revision: Optional[Union[str, Version]] = None,
download_config: Optional[DownloadConfig] = None,
download_mode: Optional[Union[DownloadMode, str]] = None,
dynamic_modules_path: Optional[str] = None,
trust_remote_code: Optional[bool] = None,
**download_kwargs,
) -> MetricModule:
"""
Download/extract/cache a metric module.
<Deprecated version="2.5.0">
Use the new library 🤗 Evaluate instead: https://huggingface.co/docs/evaluate
</Deprecated>
Metrics codes are cached inside the dynamic modules cache to allow easy import (avoid ugly sys.path tweaks).
Args:
path (str): Path or name of the metric script.
- if ``path`` is a local metric script or a directory containing a local metric script (if the script has the same name as the directory):
-> load the module from the metric script
e.g. ``'./metrics/accuracy'`` or ``'./metrics/accuracy/accuracy.py'``.
- if ``path`` is a metric on the Hugging Face Hub (ex: `glue`, `squad`)
-> load the module from the metric script in the GitHub repository at huggingface/datasets
e.g. ``'accuracy'`` or ``'rouge'``.
revision (Optional ``Union[str, datasets.Version]``):
If specified, the module will be loaded from the datasets repository at this version.
By default:
- it is set to the local version of the lib.
- it will also try to load it from the main branch if it's not available at the local version of the lib.
Specifying a version that is different from your local version of the lib might cause compatibility issues.
download_config (:class:`DownloadConfig`, optional): Specific download configuration parameters.
download_mode (:class:`DownloadMode` or :obj:`str`, default ``REUSE_DATASET_IF_EXISTS``): Download/generate mode.
dynamic_modules_path (Optional str, defaults to HF_MODULES_CACHE / "datasets_modules", i.e. ~/.cache/huggingface/modules/datasets_modules):
Optional path to the directory in which the dynamic modules are saved. It must have been initialized with :obj:`init_dynamic_modules`.
By default, the datasets and metrics are stored inside the `datasets_modules` module.
trust_remote_code (`bool`, defaults to `True`):
Whether or not to allow for datasets defined on the Hub using a dataset script. This option
should only be set to `True` for repositories you trust and in which you have read the code, as it will
execute code present on the Hub on your local machine.
<Tip warning={true}>
`trust_remote_code` will default to False in the next major release.
</Tip>
<Added version="2.16.0"/>
**download_kwargs (additional keyword arguments): optional attributes for DownloadConfig() which will override
the attributes in download_config if supplied.
Returns:
MetricModule
"""
with warnings.catch_warnings():
# Ignore equivalent warnings to the one already issued
warnings.filterwarnings("ignore", message=".*https://huggingface.co/docs/evaluate$", category=FutureWarning)
if download_config is None:
download_config = DownloadConfig(**download_kwargs)
download_mode = DownloadMode(download_mode or DownloadMode.REUSE_DATASET_IF_EXISTS)
download_config.extract_compressed_file = True
download_config.force_extract = True
filename = list(filter(lambda x: x, path.replace(os.sep, "/").split("/")))[-1]
if not filename.endswith(".py"):
filename = filename + ".py"
combined_path = os.path.join(path, filename)
# Try locally
if path.endswith(filename):
if os.path.isfile(path):
return LocalMetricModuleFactory(
path,
download_mode=download_mode,
dynamic_modules_path=dynamic_modules_path,
trust_remote_code=trust_remote_code,
).get_module()
else:
raise FileNotFoundError(f"Couldn't find a metric script at {relative_to_absolute_path(path)}")
elif os.path.isfile(combined_path):
return LocalMetricModuleFactory(
combined_path, download_mode=download_mode, dynamic_modules_path=dynamic_modules_path
).get_module()
elif is_relative_path(path) and path.count("/") == 0:
try:
return GithubMetricModuleFactory(
path,
revision=revision,
download_config=download_config,
download_mode=download_mode,
dynamic_modules_path=dynamic_modules_path,
trust_remote_code=trust_remote_code,
).get_module()
except Exception as e1: # noqa all the attempts failed, before raising the error we should check if the module is already cached.
try:
return CachedMetricModuleFactory(path, dynamic_modules_path=dynamic_modules_path).get_module()
except Exception: # noqa if it's not in the cache, then it doesn't exist.
if not isinstance(e1, FileNotFoundError):
raise e1 from None
raise FileNotFoundError(
f"Couldn't find a metric script at {relative_to_absolute_path(combined_path)}. "
f"Metric '{path}' doesn't exist on the Hugging Face Hub either."
) from None
else:
raise FileNotFoundError(f"Couldn't find a metric script at {relative_to_absolute_path(combined_path)}.")
@deprecated("Use 'evaluate.load' instead, from the new library 🤗 Evaluate: https://huggingface.co/docs/evaluate")
def load_metric(
path: str,
config_name: Optional[str] = None,
process_id: int = 0,
num_process: int = 1,
cache_dir: Optional[str] = None,
experiment_id: Optional[str] = None,
keep_in_memory: bool = False,
download_config: Optional[DownloadConfig] = None,
download_mode: Optional[Union[DownloadMode, str]] = None,
revision: Optional[Union[str, Version]] = None,
trust_remote_code: Optional[bool] = None,
**metric_init_kwargs,
) -> Metric:
"""Load a `datasets.Metric`.
<Deprecated version="2.5.0">
Use `evaluate.load` instead, from the new library 🤗 Evaluate: https://huggingface.co/docs/evaluate
</Deprecated>
Args:
path (``str``):
path to the metric processing script with the metric builder. Can be either:
- a local path to processing script or the directory containing the script (if the script has the same name as the directory),
e.g. ``'./metrics/rouge'`` or ``'./metrics/rogue/rouge.py'``
- a metric identifier on the HuggingFace datasets repo (list all available metrics with ``datasets.list_metrics()``)
e.g. ``'rouge'`` or ``'bleu'``
config_name (:obj:`str`, optional): selecting a configuration for the metric (e.g. the GLUE metric has a configuration for each subset)
process_id (:obj:`int`, optional): for distributed evaluation: id of the process
num_process (:obj:`int`, optional): for distributed evaluation: total number of processes
cache_dir (Optional str): path to store the temporary predictions and references (default to `~/.cache/huggingface/metrics/`)
experiment_id (``str``): A specific experiment id. This is used if several distributed evaluations share the same file system.
This is useful to compute metrics in distributed setups (in particular non-additive metrics like F1).
keep_in_memory (bool): Whether to store the temporary results in memory (defaults to False)
download_config (Optional ``datasets.DownloadConfig``: specific download configuration parameters.
download_mode (:class:`DownloadMode` or :obj:`str`, default ``REUSE_DATASET_IF_EXISTS``): Download/generate mode.
revision (Optional ``Union[str, datasets.Version]``): if specified, the module will be loaded from the datasets repository
at this version. By default, it is set to the local version of the lib. Specifying a version that is different from
your local version of the lib might cause compatibility issues.
trust_remote_code (`bool`, defaults to `True`):
Whether or not to allow for datasets defined on the Hub using a dataset script. This option
should only be set to `True` for repositories you trust and in which you have read the code, as it will
execute code present on the Hub on your local machine.
<Tip warning={true}>
`trust_remote_code` will default to False in the next major release.
</Tip>
<Added version="2.16.0"/>
Returns:
`datasets.Metric`
Example:
```py
>>> from datasets import load_metric
>>> accuracy = load_metric('accuracy')
>>> accuracy.compute(references=[1, 0], predictions=[1, 1])
{'accuracy': 0.5}
```
"""
with warnings.catch_warnings():
# Ignore equivalent warnings to the one already issued
warnings.filterwarnings("ignore", message=".*https://huggingface.co/docs/evaluate$", category=FutureWarning)
download_mode = DownloadMode(download_mode or DownloadMode.REUSE_DATASET_IF_EXISTS)
metric_module = metric_module_factory(
path,
revision=revision,
download_config=download_config,
download_mode=download_mode,
trust_remote_code=trust_remote_code,
).module_path
metric_cls = import_main_class(metric_module, dataset=False)
metric = metric_cls(
config_name=config_name,
process_id=process_id,
num_process=num_process,
cache_dir=cache_dir,
keep_in_memory=keep_in_memory,
experiment_id=experiment_id,
**metric_init_kwargs,
)
# Download and prepare resources for the metric
metric.download_and_prepare(download_config=download_config)
return metric
def load_dataset_builder(
path: str,
name: Optional[str] = None,
data_dir: Optional[str] = None,
data_files: Optional[Union[str, Sequence[str], Mapping[str, Union[str, Sequence[str]]]]] = None,
cache_dir: Optional[str] = None,
features: Optional[Features] = None,
download_config: Optional[DownloadConfig] = None,
download_mode: Optional[Union[DownloadMode, str]] = None,
revision: Optional[Union[str, Version]] = None,
token: Optional[Union[bool, str]] = None,
use_auth_token="deprecated",
storage_options: Optional[Dict] = None,
trust_remote_code: Optional[bool] = None,
**config_kwargs,
) -> DatasetBuilder:
"""Load a dataset builder from the Hugging Face Hub, or a local dataset. A dataset builder can be used to inspect general information that is required to build a dataset (cache directory, config, dataset info, etc.)
without downloading the dataset itself.
You can find the list of datasets on the [Hub](https://huggingface.co/datasets) or with [`huggingface_hub.list_datasets`].
A dataset is a directory that contains:
- some data files in generic formats (JSON, CSV, Parquet, text, etc.)
- and optionally a dataset script, if it requires some code to read the data files. This is used to load any kind of formats or structures.
Note that dataset scripts can also download and read data files from anywhere - in case your data files already exist online.
Args:
path (`str`):
Path or name of the dataset.
Depending on `path`, the dataset builder that is used comes from a generic dataset script (JSON, CSV, Parquet, text etc.) or from the dataset script (a python file) inside the dataset directory.
For local datasets:
- if `path` is a local directory (containing data files only)
-> load a generic dataset builder (csv, json, text etc.) based on the content of the directory
e.g. `'./path/to/directory/with/my/csv/data'`.
- if `path` is a local dataset script or a directory containing a local dataset script (if the script has the same name as the directory)
-> load the dataset builder from the dataset script
e.g. `'./dataset/squad'` or `'./dataset/squad/squad.py'`.
For datasets on the Hugging Face Hub (list all available datasets with [`huggingface_hub.list_datasets`])
- if `path` is a dataset repository on the HF hub (containing data files only)
-> load a generic dataset builder (csv, text etc.) based on the content of the repository
e.g. `'username/dataset_name'`, a dataset repository on the HF hub containing your data files.
- if `path` is a dataset repository on the HF hub with a dataset script (if the script has the same name as the directory)
-> load the dataset builder from the dataset script in the dataset repository
e.g. `glue`, `squad`, `'username/dataset_name'`, a dataset repository on the HF hub containing a dataset script `'dataset_name.py'`.
name (`str`, *optional*):
Defining the name of the dataset configuration.
data_dir (`str`, *optional*):
Defining the `data_dir` of the dataset configuration. If specified for the generic builders (csv, text etc.) or the Hub datasets and `data_files` is `None`,
the behavior is equal to passing `os.path.join(data_dir, **)` as `data_files` to reference all the files in a directory.
data_files (`str` or `Sequence` or `Mapping`, *optional*):
Path(s) to source data file(s).
cache_dir (`str`, *optional*):
Directory to read/write data. Defaults to `"~/.cache/huggingface/datasets"`.
features ([`Features`], *optional*):
Set the features type to use for this dataset.
download_config ([`DownloadConfig`], *optional*):
Specific download configuration parameters.
download_mode ([`DownloadMode`] or `str`, defaults to `REUSE_DATASET_IF_EXISTS`):
Download/generate mode.
revision ([`Version`] or `str`, *optional*):
Version of the dataset script to load.
As datasets have their own git repository on the Datasets Hub, the default version "main" corresponds to their "main" branch.
You can specify a different version than the default "main" by using a commit SHA or a git tag of the dataset repository.
token (`str` or `bool`, *optional*):
Optional string or boolean to use as Bearer token for remote files on the Datasets Hub.
If `True`, or not specified, will get token from `"~/.huggingface"`.
use_auth_token (`str` or `bool`, *optional*):
Optional string or boolean to use as Bearer token for remote files on the Datasets Hub.
If `True`, or not specified, will get token from `"~/.huggingface"`.
<Deprecated version="2.14.0">
`use_auth_token` was deprecated in favor of `token` in version 2.14.0 and will be removed in 3.0.0.
</Deprecated>
storage_options (`dict`, *optional*, defaults to `None`):
**Experimental**. Key/value pairs to be passed on to the dataset file-system backend, if any.
<Added version="2.11.0"/>
trust_remote_code (`bool`, defaults to `True`):
Whether or not to allow for datasets defined on the Hub using a dataset script. This option
should only be set to `True` for repositories you trust and in which you have read the code, as it will
execute code present on the Hub on your local machine.
<Tip warning={true}>
`trust_remote_code` will default to False in the next major release.
</Tip>
<Added version="2.16.0"/>
**config_kwargs (additional keyword arguments):
Keyword arguments to be passed to the [`BuilderConfig`]
and used in the [`DatasetBuilder`].
Returns:
[`DatasetBuilder`]
Example:
```py
>>> from datasets import load_dataset_builder
>>> ds_builder = load_dataset_builder('rotten_tomatoes')
>>> ds_builder.info.features
{'label': ClassLabel(num_classes=2, names=['neg', 'pos'], id=None),
'text': Value(dtype='string', id=None)}
```
"""
if use_auth_token != "deprecated":
warnings.warn(
"'use_auth_token' was deprecated in favor of 'token' in version 2.14.0 and will be removed in 3.0.0.\n"
"You can remove this warning by passing 'token=<use_auth_token>' instead.",
FutureWarning,
)
token = use_auth_token
download_mode = DownloadMode(download_mode or DownloadMode.REUSE_DATASET_IF_EXISTS)
if token is not None:
download_config = download_config.copy() if download_config else DownloadConfig()
download_config.token = token
if storage_options is not None:
download_config = download_config.copy() if download_config else DownloadConfig()
download_config.storage_options.update(storage_options)
dataset_module = dataset_module_factory(
path,
revision=revision,
download_config=download_config,
download_mode=download_mode,
data_dir=data_dir,
data_files=data_files,
trust_remote_code=trust_remote_code,
)
# Get dataset builder class from the processing script
builder_kwargs = dataset_module.builder_kwargs
data_dir = builder_kwargs.pop("data_dir", data_dir)
data_files = builder_kwargs.pop("data_files", data_files)
config_name = builder_kwargs.pop(
"config_name", name or dataset_module.builder_configs_parameters.default_config_name
)
dataset_name = builder_kwargs.pop("dataset_name", None)
hash = builder_kwargs.pop("hash")
info = dataset_module.dataset_infos.get(config_name) if dataset_module.dataset_infos else None
if (
dataset_module.builder_configs_parameters.metadata_configs
and config_name in dataset_module.builder_configs_parameters.metadata_configs
):
hash = update_hash_with_config_parameters(
hash, dataset_module.builder_configs_parameters.metadata_configs[config_name]
)
if path in _PACKAGED_DATASETS_MODULES and data_files is None:
error_msg = f"Please specify the data files or data directory to load for the {path} dataset builder."
example_extensions = [
extension for extension in _EXTENSION_TO_MODULE if _EXTENSION_TO_MODULE[extension] == path
]
if example_extensions:
error_msg += f'\nFor example `data_files={{"train": "path/to/data/train/*.{example_extensions[0]}"}}`'
raise ValueError(error_msg)
builder_cls = get_dataset_builder_class(dataset_module, dataset_name=dataset_name)
# Instantiate the dataset builder
builder_instance: DatasetBuilder = builder_cls(
cache_dir=cache_dir,
dataset_name=dataset_name,
config_name=config_name,
data_dir=data_dir,
data_files=data_files,
hash=hash,
info=info,
features=features,
token=token,
storage_options=storage_options,
**builder_kwargs,
**config_kwargs,
)
return builder_instance
def load_dataset(
path: str,
name: Optional[str] = None,
data_dir: Optional[str] = None,
data_files: Optional[Union[str, Sequence[str], Mapping[str, Union[str, Sequence[str]]]]] = None,
split: Optional[Union[str, Split]] = None,
cache_dir: Optional[str] = None,
features: Optional[Features] = None,
download_config: Optional[DownloadConfig] = None,
download_mode: Optional[Union[DownloadMode, str]] = None,
verification_mode: Optional[Union[VerificationMode, str]] = None,
ignore_verifications="deprecated",
keep_in_memory: Optional[bool] = None,
save_infos: bool = False,
revision: Optional[Union[str, Version]] = None,
token: Optional[Union[bool, str]] = None,
use_auth_token="deprecated",
task="deprecated",
streaming: bool = False,
num_proc: Optional[int] = None,
storage_options: Optional[Dict] = None,
trust_remote_code: bool = None,
**config_kwargs,
) -> Union[DatasetDict, Dataset, IterableDatasetDict, IterableDataset]:
"""Load a dataset from the Hugging Face Hub, or a local dataset.
You can find the list of datasets on the [Hub](https://huggingface.co/datasets) or with [`huggingface_hub.list_datasets`].
A dataset is a directory that contains:
- some data files in generic formats (JSON, CSV, Parquet, text, etc.).
- and optionally a dataset script, if it requires some code to read the data files. This is used to load any kind of formats or structures.
Note that dataset scripts can also download and read data files from anywhere - in case your data files already exist online.
This function does the following under the hood:
1. Download and import in the library the dataset script from `path` if it's not already cached inside the library.
If the dataset has no dataset script, then a generic dataset script is imported instead (JSON, CSV, Parquet, text, etc.)
Dataset scripts are small python scripts that define dataset builders. They define the citation, info and format of the dataset,
contain the path or URL to the original data files and the code to load examples from the original data files.
You can find the complete list of datasets in the Datasets [Hub](https://huggingface.co/datasets).
2. Run the dataset script which will:
* Download the dataset file from the original URL (see the script) if it's not already available locally or cached.
* Process and cache the dataset in typed Arrow tables for caching.
Arrow table are arbitrarily long, typed tables which can store nested objects and be mapped to numpy/pandas/python generic types.
They can be directly accessed from disk, loaded in RAM or even streamed over the web.
3. Return a dataset built from the requested splits in `split` (default: all).
It also allows to load a dataset from a local directory or a dataset repository on the Hugging Face Hub without dataset script.
In this case, it automatically loads all the data files from the directory or the dataset repository.
Args:
path (`str`):
Path or name of the dataset.
Depending on `path`, the dataset builder that is used comes from a generic dataset script (JSON, CSV, Parquet, text etc.) or from the dataset script (a python file) inside the dataset directory.
For local datasets:
- if `path` is a local directory (containing data files only)
-> load a generic dataset builder (csv, json, text etc.) based on the content of the directory
e.g. `'./path/to/directory/with/my/csv/data'`.
- if `path` is a local dataset script or a directory containing a local dataset script (if the script has the same name as the directory)
-> load the dataset builder from the dataset script
e.g. `'./dataset/squad'` or `'./dataset/squad/squad.py'`.
For datasets on the Hugging Face Hub (list all available datasets with [`huggingface_hub.list_datasets`])
- if `path` is a dataset repository on the HF hub (containing data files only)
-> load a generic dataset builder (csv, text etc.) based on the content of the repository
e.g. `'username/dataset_name'`, a dataset repository on the HF hub containing your data files.
- if `path` is a dataset repository on the HF hub with a dataset script (if the script has the same name as the directory)
-> load the dataset builder from the dataset script in the dataset repository
e.g. `glue`, `squad`, `'username/dataset_name'`, a dataset repository on the HF hub containing a dataset script `'dataset_name.py'`.
name (`str`, *optional*):
Defining the name of the dataset configuration.
data_dir (`str`, *optional*):
Defining the `data_dir` of the dataset configuration. If specified for the generic builders (csv, text etc.) or the Hub datasets and `data_files` is `None`,
the behavior is equal to passing `os.path.join(data_dir, **)` as `data_files` to reference all the files in a directory.
data_files (`str` or `Sequence` or `Mapping`, *optional*):
Path(s) to source data file(s).
split (`Split` or `str`):
Which split of the data to load.
If `None`, will return a `dict` with all splits (typically `datasets.Split.TRAIN` and `datasets.Split.TEST`).
If given, will return a single Dataset.
Splits can be combined and specified like in tensorflow-datasets.
cache_dir (`str`, *optional*):
Directory to read/write data. Defaults to `"~/.cache/huggingface/datasets"`.
features (`Features`, *optional*):
Set the features type to use for this dataset.
download_config ([`DownloadConfig`], *optional*):
Specific download configuration parameters.
download_mode ([`DownloadMode`] or `str`, defaults to `REUSE_DATASET_IF_EXISTS`):
Download/generate mode.
verification_mode ([`VerificationMode`] or `str`, defaults to `BASIC_CHECKS`):
Verification mode determining the checks to run on the downloaded/processed dataset information (checksums/size/splits/...).
<Added version="2.9.1"/>
ignore_verifications (`bool`, defaults to `False`):
Ignore the verifications of the downloaded/processed dataset information (checksums/size/splits/...).
<Deprecated version="2.9.1">
`ignore_verifications` was deprecated in version 2.9.1 and will be removed in 3.0.0.
Please use `verification_mode` instead.
</Deprecated>
keep_in_memory (`bool`, defaults to `None`):
Whether to copy the dataset in-memory. If `None`, the dataset
will not be copied in-memory unless explicitly enabled by setting `datasets.config.IN_MEMORY_MAX_SIZE` to
nonzero. See more details in the [improve performance](../cache#improve-performance) section.
save_infos (`bool`, defaults to `False`):
Save the dataset information (checksums/size/splits/...).
revision ([`Version`] or `str`, *optional*):
Version of the dataset script to load.
As datasets have their own git repository on the Datasets Hub, the default version "main" corresponds to their "main" branch.
You can specify a different version than the default "main" by using a commit SHA or a git tag of the dataset repository.
token (`str` or `bool`, *optional*):
Optional string or boolean to use as Bearer token for remote files on the Datasets Hub.
If `True`, or not specified, will get token from `"~/.huggingface"`.
use_auth_token (`str` or `bool`, *optional*):
Optional string or boolean to use as Bearer token for remote files on the Datasets Hub.
If `True`, or not specified, will get token from `"~/.huggingface"`.
<Deprecated version="2.14.0">
`use_auth_token` was deprecated in favor of `token` in version 2.14.0 and will be removed in 3.0.0.
</Deprecated>
task (`str`):
The task to prepare the dataset for during training and evaluation. Casts the dataset's [`Features`] to standardized column names and types as detailed in `datasets.tasks`.
<Deprecated version="2.13.0">
`task` was deprecated in version 2.13.0 and will be removed in 3.0.0.
</Deprecated>
streaming (`bool`, defaults to `False`):
If set to `True`, don't download the data files. Instead, it streams the data progressively while
iterating on the dataset. An [`IterableDataset`] or [`IterableDatasetDict`] is returned instead in this case.
Note that streaming works for datasets that use data formats that support being iterated over like txt, csv, jsonl for example.
Json files may be downloaded completely. Also streaming from remote zip or gzip files is supported but other compressed formats
like rar and xz are not yet supported. The tgz format doesn't allow streaming.
num_proc (`int`, *optional*, defaults to `None`):
Number of processes when downloading and generating the dataset locally.
Multiprocessing is disabled by default.
<Added version="2.7.0"/>
storage_options (`dict`, *optional*, defaults to `None`):
**Experimental**. Key/value pairs to be passed on to the dataset file-system backend, if any.
<Added version="2.11.0"/>
trust_remote_code (`bool`, defaults to `True`):
Whether or not to allow for datasets defined on the Hub using a dataset script. This option
should only be set to `True` for repositories you trust and in which you have read the code, as it will
execute code present on the Hub on your local machine.
<Tip warning={true}>
`trust_remote_code` will default to False in the next major release.
</Tip>
<Added version="2.16.0"/>
**config_kwargs (additional keyword arguments):
Keyword arguments to be passed to the `BuilderConfig`
and used in the [`DatasetBuilder`].
Returns:
[`Dataset`] or [`DatasetDict`]:
- if `split` is not `None`: the dataset requested,
- if `split` is `None`, a [`~datasets.DatasetDict`] with each split.
or [`IterableDataset`] or [`IterableDatasetDict`]: if `streaming=True`
- if `split` is not `None`, the dataset is requested
- if `split` is `None`, a [`~datasets.streaming.IterableDatasetDict`] with each split.
Example:
Load a dataset from the Hugging Face Hub:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset('rotten_tomatoes', split='train')
# Map data files to splits
>>> data_files = {'train': 'train.csv', 'test': 'test.csv'}
>>> ds = load_dataset('namespace/your_dataset_name', data_files=data_files)
```
Load a local dataset:
```py
# Load a CSV file
>>> from datasets import load_dataset
>>> ds = load_dataset('csv', data_files='path/to/local/my_dataset.csv')
# Load a JSON file
>>> from datasets import load_dataset
>>> ds = load_dataset('json', data_files='path/to/local/my_dataset.json')
# Load from a local loading script
>>> from datasets import load_dataset
>>> ds = load_dataset('path/to/local/loading_script/loading_script.py', split='train')
```
Load an [`~datasets.IterableDataset`]:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset('rotten_tomatoes', split='train', streaming=True)
```
Load an image dataset with the `ImageFolder` dataset builder:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset('imagefolder', data_dir='/path/to/images', split='train')
```
"""
if use_auth_token != "deprecated":
warnings.warn(
"'use_auth_token' was deprecated in favor of 'token' in version 2.14.0 and will be removed in 3.0.0.\n"
"You can remove this warning by passing 'token=<use_auth_token>' instead.",
FutureWarning,
)
token = use_auth_token
if ignore_verifications != "deprecated":
verification_mode = VerificationMode.NO_CHECKS if ignore_verifications else VerificationMode.ALL_CHECKS
warnings.warn(
"'ignore_verifications' was deprecated in favor of 'verification_mode' in version 2.9.1 and will be removed in 3.0.0.\n"
f"You can remove this warning by passing 'verification_mode={verification_mode.value}' instead.",
FutureWarning,
)
if task != "deprecated":
warnings.warn(
"'task' was deprecated in version 2.13.0 and will be removed in 3.0.0.\n",
FutureWarning,
)
else:
task = None
if data_files is not None and not data_files:
raise ValueError(f"Empty 'data_files': '{data_files}'. It should be either non-empty or None (default).")
if Path(path, config.DATASET_STATE_JSON_FILENAME).exists():
raise ValueError(
"You are trying to load a dataset that was saved using `save_to_disk`. "
"Please use `load_from_disk` instead."
)
if streaming and num_proc is not None:
raise NotImplementedError(
"Loading a streaming dataset in parallel with `num_proc` is not implemented. "
"To parallelize streaming, you can wrap the dataset with a PyTorch DataLoader using `num_workers` > 1 instead."
)
download_mode = DownloadMode(download_mode or DownloadMode.REUSE_DATASET_IF_EXISTS)
verification_mode = VerificationMode(
(verification_mode or VerificationMode.BASIC_CHECKS) if not save_infos else VerificationMode.ALL_CHECKS
)
# Create a dataset builder
builder_instance = load_dataset_builder(
path=path,
name=name,
data_dir=data_dir,
data_files=data_files,
cache_dir=cache_dir,
features=features,
download_config=download_config,
download_mode=download_mode,
revision=revision,
token=token,
storage_options=storage_options,
trust_remote_code=trust_remote_code,
**config_kwargs,
)
# Return iterable dataset in case of streaming
if streaming:
return builder_instance.as_streaming_dataset(split=split)
# Some datasets are already processed on the HF google storage
# Don't try downloading from Google storage for the packaged datasets as text, json, csv or pandas
try_from_hf_gcs = path not in _PACKAGED_DATASETS_MODULES
# Download and prepare data
builder_instance.download_and_prepare(
download_config=download_config,
download_mode=download_mode,
verification_mode=verification_mode,
try_from_hf_gcs=try_from_hf_gcs,
num_proc=num_proc,
storage_options=storage_options,
)
# Build dataset for splits
keep_in_memory = (
keep_in_memory if keep_in_memory is not None else is_small_dataset(builder_instance.info.dataset_size)
)
ds = builder_instance.as_dataset(split=split, verification_mode=verification_mode, in_memory=keep_in_memory)
# Rename and cast features to match task schema
if task is not None:
# To avoid issuing the same warning twice
with warnings.catch_warnings():
warnings.simplefilter("ignore", FutureWarning)
ds = ds.prepare_for_task(task)
if save_infos:
builder_instance._save_infos()
return ds
def load_from_disk(
dataset_path: str, fs="deprecated", keep_in_memory: Optional[bool] = None, storage_options: Optional[dict] = None
) -> Union[Dataset, DatasetDict]:
"""
Loads a dataset that was previously saved using [`~Dataset.save_to_disk`] from a dataset directory, or
from a filesystem using any implementation of `fsspec.spec.AbstractFileSystem`.
Args:
dataset_path (`str`):
Path (e.g. `"dataset/train"`) or remote URI (e.g.
`"s3://my-bucket/dataset/train"`) of the [`Dataset`] or [`DatasetDict`] directory where the dataset will be
loaded from.
fs (`~filesystems.S3FileSystem` or `fsspec.spec.AbstractFileSystem`, *optional*):
Instance of the remote filesystem used to download the files from.
<Deprecated version="2.9.0">
`fs` was deprecated in version 2.9.0 and will be removed in 3.0.0.
Please use `storage_options` instead, e.g. `storage_options=fs.storage_options`.
</Deprecated>
keep_in_memory (`bool`, defaults to `None`):
Whether to copy the dataset in-memory. If `None`, the dataset
will not be copied in-memory unless explicitly enabled by setting `datasets.config.IN_MEMORY_MAX_SIZE` to
nonzero. See more details in the [improve performance](../cache#improve-performance) section.
storage_options (`dict`, *optional*):
Key/value pairs to be passed on to the file-system backend, if any.
<Added version="2.9.0"/>
Returns:
[`Dataset`] or [`DatasetDict`]:
- If `dataset_path` is a path of a dataset directory: the dataset requested.
- If `dataset_path` is a path of a dataset dict directory, a [`DatasetDict`] with each split.
Example:
```py
>>> from datasets import load_from_disk
>>> ds = load_from_disk('path/to/dataset/directory')
```
"""
if fs != "deprecated":
warnings.warn(
"'fs' was deprecated in favor of 'storage_options' in version 2.9.0 and will be removed in 3.0.0.\n"
"You can remove this warning by passing 'storage_options=fs.storage_options' instead.",
FutureWarning,
)
storage_options = fs.storage_options
fs: fsspec.AbstractFileSystem
fs, _, _ = fsspec.get_fs_token_paths(dataset_path, storage_options=storage_options)
if not fs.exists(dataset_path):
raise FileNotFoundError(f"Directory {dataset_path} not found")
if fs.isfile(posixpath.join(dataset_path, config.DATASET_INFO_FILENAME)) and fs.isfile(
posixpath.join(dataset_path, config.DATASET_STATE_JSON_FILENAME)
):
return Dataset.load_from_disk(dataset_path, keep_in_memory=keep_in_memory, storage_options=storage_options)
elif fs.isfile(posixpath.join(dataset_path, config.DATASETDICT_JSON_FILENAME)):
return DatasetDict.load_from_disk(dataset_path, keep_in_memory=keep_in_memory, storage_options=storage_options)
else:
raise FileNotFoundError(
f"Directory {dataset_path} is neither a `Dataset` directory nor a `DatasetDict` directory."
)
| 0
|
hf_public_repos/datasets/src
|
hf_public_repos/datasets/src/datasets/exceptions.py
|
# SPDX-License-Identifier: Apache-2.0
# Copyright 2023 The HuggingFace Authors.
class DatasetsError(Exception):
"""Base class for exceptions in this library."""
class DefunctDatasetError(DatasetsError):
"""The dataset has been defunct."""
class FileNotFoundDatasetsError(DatasetsError, FileNotFoundError):
"""FileNotFoundError raised by this library."""
class DataFilesNotFoundError(FileNotFoundDatasetsError):
"""No (supported) data files found."""
class DatasetNotFoundError(FileNotFoundDatasetsError):
"""Dataset not found.
Raised when trying to access:
- a missing dataset, or
- a private/gated dataset and the user is not authenticated.
"""
| 0
|
hf_public_repos/datasets/src
|
hf_public_repos/datasets/src/datasets/combine.py
|
from typing import List, Optional, TypeVar
from .arrow_dataset import Dataset, _concatenate_map_style_datasets, _interleave_map_style_datasets
from .dataset_dict import DatasetDict, IterableDatasetDict
from .info import DatasetInfo
from .iterable_dataset import IterableDataset, _concatenate_iterable_datasets, _interleave_iterable_datasets
from .splits import NamedSplit
from .utils import logging
from .utils.py_utils import Literal
logger = logging.get_logger(__name__)
DatasetType = TypeVar("DatasetType", Dataset, IterableDataset)
def interleave_datasets(
datasets: List[DatasetType],
probabilities: Optional[List[float]] = None,
seed: Optional[int] = None,
info: Optional[DatasetInfo] = None,
split: Optional[NamedSplit] = None,
stopping_strategy: Literal["first_exhausted", "all_exhausted"] = "first_exhausted",
) -> DatasetType:
"""
Interleave several datasets (sources) into a single dataset.
The new dataset is constructed by alternating between the sources to get the examples.
You can use this function on a list of [`Dataset`] objects, or on a list of [`IterableDataset`] objects.
- If `probabilities` is `None` (default) the new dataset is constructed by cycling between each source to get the examples.
- If `probabilities` is not `None`, the new dataset is constructed by getting examples from a random source at a time according to the provided probabilities.
The resulting dataset ends when one of the source datasets runs out of examples except when `oversampling` is `True`,
in which case, the resulting dataset ends when all datasets have ran out of examples at least one time.
Note for iterable datasets:
In a distributed setup or in PyTorch DataLoader workers, the stopping strategy is applied per process.
Therefore the "first_exhausted" strategy on an sharded iterable dataset can generate less samples in total (up to 1 missing sample per subdataset per worker).
Args:
datasets (`List[Dataset]` or `List[IterableDataset]`):
List of datasets to interleave.
probabilities (`List[float]`, *optional*, defaults to `None`):
If specified, the new dataset is constructed by sampling
examples from one source at a time according to these probabilities.
seed (`int`, *optional*, defaults to `None`):
The random seed used to choose a source for each example.
info ([`DatasetInfo`], *optional*):
Dataset information, like description, citation, etc.
<Added version="2.4.0"/>
split ([`NamedSplit`], *optional*):
Name of the dataset split.
<Added version="2.4.0"/>
stopping_strategy (`str`, defaults to `first_exhausted`):
Two strategies are proposed right now, `first_exhausted` and `all_exhausted`.
By default, `first_exhausted` is an undersampling strategy, i.e the dataset construction is stopped as soon as one dataset has ran out of samples.
If the strategy is `all_exhausted`, we use an oversampling strategy, i.e the dataset construction is stopped as soon as every samples of every dataset has been added at least once.
Note that if the strategy is `all_exhausted`, the interleaved dataset size can get enormous:
- with no probabilities, the resulting dataset will have `max_length_datasets*nb_dataset` samples.
- with given probabilities, the resulting dataset will have more samples if some datasets have really low probability of visiting.
Returns:
[`Dataset`] or [`IterableDataset`]: Return type depends on the input `datasets`
parameter. `Dataset` if the input is a list of `Dataset`, `IterableDataset` if the input is a list of
`IterableDataset`.
Example:
For regular datasets (map-style):
```python
>>> from datasets import Dataset, interleave_datasets
>>> d1 = Dataset.from_dict({"a": [0, 1, 2]})
>>> d2 = Dataset.from_dict({"a": [10, 11, 12]})
>>> d3 = Dataset.from_dict({"a": [20, 21, 22]})
>>> dataset = interleave_datasets([d1, d2, d3], probabilities=[0.7, 0.2, 0.1], seed=42, stopping_strategy="all_exhausted")
>>> dataset["a"]
[10, 0, 11, 1, 2, 20, 12, 10, 0, 1, 2, 21, 0, 11, 1, 2, 0, 1, 12, 2, 10, 0, 22]
>>> dataset = interleave_datasets([d1, d2, d3], probabilities=[0.7, 0.2, 0.1], seed=42)
>>> dataset["a"]
[10, 0, 11, 1, 2]
>>> dataset = interleave_datasets([d1, d2, d3])
>>> dataset["a"]
[0, 10, 20, 1, 11, 21, 2, 12, 22]
>>> dataset = interleave_datasets([d1, d2, d3], stopping_strategy="all_exhausted")
>>> dataset["a"]
[0, 10, 20, 1, 11, 21, 2, 12, 22]
>>> d1 = Dataset.from_dict({"a": [0, 1, 2]})
>>> d2 = Dataset.from_dict({"a": [10, 11, 12, 13]})
>>> d3 = Dataset.from_dict({"a": [20, 21, 22, 23, 24]})
>>> dataset = interleave_datasets([d1, d2, d3])
>>> dataset["a"]
[0, 10, 20, 1, 11, 21, 2, 12, 22]
>>> dataset = interleave_datasets([d1, d2, d3], stopping_strategy="all_exhausted")
>>> dataset["a"]
[0, 10, 20, 1, 11, 21, 2, 12, 22, 0, 13, 23, 1, 10, 24]
>>> dataset = interleave_datasets([d1, d2, d3], probabilities=[0.7, 0.2, 0.1], seed=42)
>>> dataset["a"]
[10, 0, 11, 1, 2]
>>> dataset = interleave_datasets([d1, d2, d3], probabilities=[0.7, 0.2, 0.1], seed=42, stopping_strategy="all_exhausted")
>>> dataset["a"]
[10, 0, 11, 1, 2, 20, 12, 13, ..., 0, 1, 2, 0, 24]
For datasets in streaming mode (iterable):
>>> from datasets import load_dataset, interleave_datasets
>>> d1 = load_dataset("oscar", "unshuffled_deduplicated_en", split="train", streaming=True)
>>> d2 = load_dataset("oscar", "unshuffled_deduplicated_fr", split="train", streaming=True)
>>> dataset = interleave_datasets([d1, d2])
>>> iterator = iter(dataset)
>>> next(iterator)
{'text': 'Mtendere Village was inspired by the vision...}
>>> next(iterator)
{'text': "Média de débat d'idées, de culture...}
```
"""
from .arrow_dataset import Dataset
from .iterable_dataset import IterableDataset
if not datasets:
raise ValueError("Unable to interleave an empty list of datasets.")
for i, dataset in enumerate(datasets):
if not isinstance(dataset, (Dataset, IterableDataset)):
if isinstance(dataset, (DatasetDict, IterableDatasetDict)):
if not dataset:
raise ValueError(
f"Expected a list of Dataset objects or a list of IterableDataset objects, but element at position {i} "
"is an empty dataset dictionary."
)
raise ValueError(
f"Dataset at position {i} has at least one split: {list(dataset)}\n"
f"Please pick one to interleave with the other datasets, for example: dataset['{next(iter(dataset))}']"
)
raise ValueError(
f"Expected a list of Dataset objects or a list of IterableDataset objects, but element at position {i} is a {type(dataset).__name__}."
)
if i == 0:
dataset_type, other_type = (
(Dataset, IterableDataset) if isinstance(dataset, Dataset) else (IterableDataset, Dataset)
)
elif not isinstance(dataset, dataset_type):
raise ValueError(
f"Unable to interleave a {dataset_type.__name__} (at position 0) with a {other_type.__name__} (at position {i}). Expected a list of Dataset objects or a list of IterableDataset objects."
)
if stopping_strategy not in ["first_exhausted", "all_exhausted"]:
raise ValueError(f"{stopping_strategy} is not supported. Please enter a valid stopping_strategy.")
if dataset_type is Dataset:
return _interleave_map_style_datasets(
datasets, probabilities, seed, info=info, split=split, stopping_strategy=stopping_strategy
)
else:
return _interleave_iterable_datasets(
datasets, probabilities, seed, info=info, split=split, stopping_strategy=stopping_strategy
)
def concatenate_datasets(
dsets: List[DatasetType],
info: Optional[DatasetInfo] = None,
split: Optional[NamedSplit] = None,
axis: int = 0,
) -> DatasetType:
"""
Converts a list of [`Dataset`] with the same schema into a single [`Dataset`].
Args:
dsets (`List[datasets.Dataset]`):
List of Datasets to concatenate.
info (`DatasetInfo`, *optional*):
Dataset information, like description, citation, etc.
split (`NamedSplit`, *optional*):
Name of the dataset split.
axis (`{0, 1}`, defaults to `0`):
Axis to concatenate over, where `0` means over rows (vertically) and `1` means over columns
(horizontally).
<Added version="1.6.0"/>
Example:
```py
>>> ds3 = concatenate_datasets([ds1, ds2])
```
"""
if not dsets:
raise ValueError("Unable to concatenate an empty list of datasets.")
for i, dataset in enumerate(dsets):
if not isinstance(dataset, (Dataset, IterableDataset)):
if isinstance(dataset, (DatasetDict, IterableDatasetDict)):
if not dataset:
raise ValueError(
f"Expected a list of Dataset objects or a list of IterableDataset objects, but element at position {i} "
"is an empty dataset dictionary."
)
raise ValueError(
f"Dataset at position {i} has at least one split: {list(dataset)}\n"
f"Please pick one to interleave with the other datasets, for example: dataset['{next(iter(dataset))}']"
)
raise ValueError(
f"Expected a list of Dataset objects or a list of IterableDataset objects, but element at position {i} is a {type(dataset).__name__}."
)
if i == 0:
dataset_type, other_type = (
(Dataset, IterableDataset) if isinstance(dataset, Dataset) else (IterableDataset, Dataset)
)
elif not isinstance(dataset, dataset_type):
raise ValueError(
f"Unable to interleave a {dataset_type.__name__} (at position 0) with a {other_type.__name__} (at position {i}). Expected a list of Dataset objects or a list of IterableDataset objects."
)
if dataset_type is Dataset:
return _concatenate_map_style_datasets(dsets, info=info, split=split, axis=axis)
else:
return _concatenate_iterable_datasets(dsets, info=info, split=split, axis=axis)
| 0
|
hf_public_repos/datasets/src
|
hf_public_repos/datasets/src/datasets/arrow_writer.py
|
# Copyright 2020 The HuggingFace Datasets Authors and the TensorFlow Datasets Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# Lint as: python3
"""To write records into Parquet files."""
import errno
import json
import os
import sys
from pathlib import Path
from typing import Any, Dict, Iterable, List, Optional, Tuple, Union
import fsspec
import numpy as np
import pyarrow as pa
import pyarrow.parquet as pq
from . import config
from .features import Features, Image, Value
from .features.features import (
FeatureType,
_ArrayXDExtensionType,
cast_to_python_objects,
generate_from_arrow_type,
get_nested_type,
list_of_np_array_to_pyarrow_listarray,
numpy_to_pyarrow_listarray,
to_pyarrow_listarray,
)
from .filesystems import is_remote_filesystem
from .info import DatasetInfo
from .keyhash import DuplicatedKeysError, KeyHasher
from .table import array_cast, array_concat, cast_array_to_feature, embed_table_storage, table_cast
from .utils import logging
from .utils import tqdm as hf_tqdm
from .utils.file_utils import hash_url_to_filename
from .utils.py_utils import asdict, first_non_null_value
logger = logging.get_logger(__name__)
type_ = type # keep python's type function
class SchemaInferenceError(ValueError):
pass
class TypedSequence:
"""
This data container generalizes the typing when instantiating pyarrow arrays, tables or batches.
More specifically it adds several features:
- Support extension types like ``datasets.features.Array2DExtensionType``:
By default pyarrow arrays don't return extension arrays. One has to call
``pa.ExtensionArray.from_storage(type, pa.array(data, type.storage_type))``
in order to get an extension array.
- Support for ``try_type`` parameter that can be used instead of ``type``:
When an array is transformed, we like to keep the same type as before if possible.
For example when calling :func:`datasets.Dataset.map`, we don't want to change the type
of each column by default.
- Better error message when a pyarrow array overflows.
Example::
from datasets.features import Array2D, Array2DExtensionType, Value
from datasets.arrow_writer import TypedSequence
import pyarrow as pa
arr = pa.array(TypedSequence([1, 2, 3], type=Value("int32")))
assert arr.type == pa.int32()
arr = pa.array(TypedSequence([1, 2, 3], try_type=Value("int32")))
assert arr.type == pa.int32()
arr = pa.array(TypedSequence(["foo", "bar"], try_type=Value("int32")))
assert arr.type == pa.string()
arr = pa.array(TypedSequence([[[1, 2, 3]]], type=Array2D((1, 3), "int64")))
assert arr.type == Array2DExtensionType((1, 3), "int64")
table = pa.Table.from_pydict({
"image": TypedSequence([[[1, 2, 3]]], type=Array2D((1, 3), "int64"))
})
assert table["image"].type == Array2DExtensionType((1, 3), "int64")
"""
def __init__(
self,
data: Iterable,
type: Optional[FeatureType] = None,
try_type: Optional[FeatureType] = None,
optimized_int_type: Optional[FeatureType] = None,
):
# assert type is None or try_type is None,
if type is not None and try_type is not None:
raise ValueError("You cannot specify both type and try_type")
# set attributes
self.data = data
self.type = type
self.try_type = try_type # is ignored if it doesn't match the data
self.optimized_int_type = optimized_int_type
# when trying a type (is ignored if data is not compatible)
self.trying_type = self.try_type is not None
self.trying_int_optimization = optimized_int_type is not None and type is None and try_type is None
# used to get back the inferred type after __arrow_array__() is called once
self._inferred_type = None
def get_inferred_type(self) -> FeatureType:
"""Return the inferred feature type.
This is done by converting the sequence to an Arrow array, and getting the corresponding
feature type.
Since building the Arrow array can be expensive, the value of the inferred type is cached
as soon as pa.array is called on the typed sequence.
Returns:
FeatureType: inferred feature type of the sequence.
"""
if self._inferred_type is None:
self._inferred_type = generate_from_arrow_type(pa.array(self).type)
return self._inferred_type
@staticmethod
def _infer_custom_type_and_encode(data: Iterable) -> Tuple[Iterable, Optional[FeatureType]]:
"""Implement type inference for custom objects like PIL.Image.Image -> Image type.
This function is only used for custom python objects that can't be direclty passed to build
an Arrow array. In such cases is infers the feature type to use, and it encodes the data so
that they can be passed to an Arrow array.
Args:
data (Iterable): array of data to infer the type, e.g. a list of PIL images.
Returns:
Tuple[Iterable, Optional[FeatureType]]: a tuple with:
- the (possibly encoded) array, if the inferred feature type requires encoding
- the inferred feature type if the array is made of supported custom objects like
PIL images, else None.
"""
if config.PIL_AVAILABLE and "PIL" in sys.modules:
import PIL.Image
non_null_idx, non_null_value = first_non_null_value(data)
if isinstance(non_null_value, PIL.Image.Image):
return [Image().encode_example(value) if value is not None else None for value in data], Image()
return data, None
def __arrow_array__(self, type: Optional[pa.DataType] = None):
"""This function is called when calling pa.array(typed_sequence)"""
if type is not None:
raise ValueError("TypedSequence is supposed to be used with pa.array(typed_sequence, type=None)")
del type # make sure we don't use it
data = self.data
# automatic type inference for custom objects
if self.type is None and self.try_type is None:
data, self._inferred_type = self._infer_custom_type_and_encode(data)
if self._inferred_type is None:
type = self.try_type if self.trying_type else self.type
else:
type = self._inferred_type
pa_type = get_nested_type(type) if type is not None else None
optimized_int_pa_type = (
get_nested_type(self.optimized_int_type) if self.optimized_int_type is not None else None
)
trying_cast_to_python_objects = False
try:
# custom pyarrow types
if isinstance(pa_type, _ArrayXDExtensionType):
storage = to_pyarrow_listarray(data, pa_type)
return pa.ExtensionArray.from_storage(pa_type, storage)
# efficient np array to pyarrow array
if isinstance(data, np.ndarray):
out = numpy_to_pyarrow_listarray(data)
elif isinstance(data, list) and data and isinstance(first_non_null_value(data)[1], np.ndarray):
out = list_of_np_array_to_pyarrow_listarray(data)
else:
trying_cast_to_python_objects = True
out = pa.array(cast_to_python_objects(data, only_1d_for_numpy=True))
# use smaller integer precisions if possible
if self.trying_int_optimization:
if pa.types.is_int64(out.type):
out = out.cast(optimized_int_pa_type)
elif pa.types.is_list(out.type):
if pa.types.is_int64(out.type.value_type):
out = array_cast(out, pa.list_(optimized_int_pa_type))
elif pa.types.is_list(out.type.value_type) and pa.types.is_int64(out.type.value_type.value_type):
out = array_cast(out, pa.list_(pa.list_(optimized_int_pa_type)))
# otherwise we can finally use the user's type
elif type is not None:
# We use cast_array_to_feature to support casting to custom types like Audio and Image
# Also, when trying type "string", we don't want to convert integers or floats to "string".
# We only do it if trying_type is False - since this is what the user asks for.
out = cast_array_to_feature(out, type, allow_number_to_str=not self.trying_type)
return out
except (
TypeError,
pa.lib.ArrowInvalid,
pa.lib.ArrowNotImplementedError,
) as e: # handle type errors and overflows
# Ignore ArrowNotImplementedError caused by trying type, otherwise re-raise
if not self.trying_type and isinstance(e, pa.lib.ArrowNotImplementedError):
raise
if self.trying_type:
try: # second chance
if isinstance(data, np.ndarray):
return numpy_to_pyarrow_listarray(data)
elif isinstance(data, list) and data and any(isinstance(value, np.ndarray) for value in data):
return list_of_np_array_to_pyarrow_listarray(data)
else:
trying_cast_to_python_objects = True
return pa.array(cast_to_python_objects(data, only_1d_for_numpy=True))
except pa.lib.ArrowInvalid as e:
if "overflow" in str(e):
raise OverflowError(
f"There was an overflow with type {type_(data)}. Try to reduce writer_batch_size to have batches smaller than 2GB.\n({e})"
) from None
elif self.trying_int_optimization and "not in range" in str(e):
optimized_int_pa_type_str = np.dtype(optimized_int_pa_type.to_pandas_dtype()).name
logger.info(
f"Failed to cast a sequence to {optimized_int_pa_type_str}. Falling back to int64."
)
return out
elif trying_cast_to_python_objects and "Could not convert" in str(e):
out = pa.array(
cast_to_python_objects(data, only_1d_for_numpy=True, optimize_list_casting=False)
)
if type is not None:
out = cast_array_to_feature(out, type, allow_number_to_str=True)
return out
else:
raise
elif "overflow" in str(e):
raise OverflowError(
f"There was an overflow with type {type_(data)}. Try to reduce writer_batch_size to have batches smaller than 2GB.\n({e})"
) from None
elif self.trying_int_optimization and "not in range" in str(e):
optimized_int_pa_type_str = np.dtype(optimized_int_pa_type.to_pandas_dtype()).name
logger.info(f"Failed to cast a sequence to {optimized_int_pa_type_str}. Falling back to int64.")
return out
elif trying_cast_to_python_objects and "Could not convert" in str(e):
out = pa.array(cast_to_python_objects(data, only_1d_for_numpy=True, optimize_list_casting=False))
if type is not None:
out = cast_array_to_feature(out, type, allow_number_to_str=True)
return out
else:
raise
class OptimizedTypedSequence(TypedSequence):
def __init__(
self,
data,
type: Optional[FeatureType] = None,
try_type: Optional[FeatureType] = None,
col: Optional[str] = None,
optimized_int_type: Optional[FeatureType] = None,
):
optimized_int_type_by_col = {
"attention_mask": Value("int8"), # binary tensor
"special_tokens_mask": Value("int8"),
"input_ids": Value("int32"), # typical vocab size: 0-50k (max ~500k, never > 1M)
"token_type_ids": Value(
"int8"
), # binary mask; some (XLNetModel) use an additional token represented by a 2
}
if type is None and try_type is None:
optimized_int_type = optimized_int_type_by_col.get(col, None)
super().__init__(data, type=type, try_type=try_type, optimized_int_type=optimized_int_type)
class ArrowWriter:
"""Shuffles and writes Examples to Arrow files."""
_WRITER_CLASS = pa.RecordBatchStreamWriter
def __init__(
self,
schema: Optional[pa.Schema] = None,
features: Optional[Features] = None,
path: Optional[str] = None,
stream: Optional[pa.NativeFile] = None,
fingerprint: Optional[str] = None,
writer_batch_size: Optional[int] = None,
hash_salt: Optional[str] = None,
check_duplicates: Optional[bool] = False,
disable_nullable: bool = False,
update_features: bool = False,
with_metadata: bool = True,
unit: str = "examples",
embed_local_files: bool = False,
storage_options: Optional[dict] = None,
):
if path is None and stream is None:
raise ValueError("At least one of path and stream must be provided.")
if features is not None:
self._features = features
self._schema = None
elif schema is not None:
self._schema: pa.Schema = schema
self._features = Features.from_arrow_schema(self._schema)
else:
self._features = None
self._schema = None
if hash_salt is not None:
# Create KeyHasher instance using split name as hash salt
self._hasher = KeyHasher(hash_salt)
else:
self._hasher = KeyHasher("")
self._check_duplicates = check_duplicates
self._disable_nullable = disable_nullable
if stream is None:
fs_token_paths = fsspec.get_fs_token_paths(path, storage_options=storage_options)
self._fs: fsspec.AbstractFileSystem = fs_token_paths[0]
self._path = (
fs_token_paths[2][0]
if not is_remote_filesystem(self._fs)
else self._fs.unstrip_protocol(fs_token_paths[2][0])
)
self.stream = self._fs.open(fs_token_paths[2][0], "wb")
self._closable_stream = True
else:
self._fs = None
self._path = None
self.stream = stream
self._closable_stream = False
self.fingerprint = fingerprint
self.disable_nullable = disable_nullable
self.writer_batch_size = writer_batch_size or config.DEFAULT_MAX_BATCH_SIZE
self.update_features = update_features
self.with_metadata = with_metadata
self.unit = unit
self.embed_local_files = embed_local_files
self._num_examples = 0
self._num_bytes = 0
self.current_examples: List[Tuple[Dict[str, Any], str]] = []
self.current_rows: List[pa.Table] = []
self.pa_writer: Optional[pa.RecordBatchStreamWriter] = None
self.hkey_record = []
def __len__(self):
"""Return the number of writed and staged examples"""
return self._num_examples + len(self.current_examples) + len(self.current_rows)
def __enter__(self):
return self
def __exit__(self, exc_type, exc_val, exc_tb):
self.close()
def close(self):
# Try closing if opened; if closed: pyarrow.lib.ArrowInvalid: Invalid operation on closed file
if self.pa_writer: # it might be None
try:
self.pa_writer.close()
except Exception: # pyarrow.lib.ArrowInvalid, OSError
pass
if self._closable_stream and not self.stream.closed:
self.stream.close() # This also closes self.pa_writer if it is opened
def _build_writer(self, inferred_schema: pa.Schema):
schema = self.schema
inferred_features = Features.from_arrow_schema(inferred_schema)
if self._features is not None:
if self.update_features: # keep original features it they match, or update them
fields = {field.name: field for field in self._features.type}
for inferred_field in inferred_features.type:
name = inferred_field.name
if name in fields:
if inferred_field == fields[name]:
inferred_features[name] = self._features[name]
self._features = inferred_features
schema: pa.Schema = inferred_schema
else:
self._features = inferred_features
schema: pa.Schema = inferred_features.arrow_schema
if self.disable_nullable:
schema = pa.schema(pa.field(field.name, field.type, nullable=False) for field in schema)
if self.with_metadata:
schema = schema.with_metadata(self._build_metadata(DatasetInfo(features=self._features), self.fingerprint))
else:
schema = schema.with_metadata({})
self._schema = schema
self.pa_writer = self._WRITER_CLASS(self.stream, schema)
@property
def schema(self):
_schema = (
self._schema
if self._schema is not None
else (pa.schema(self._features.type) if self._features is not None else None)
)
if self._disable_nullable and _schema is not None:
_schema = pa.schema(pa.field(field.name, field.type, nullable=False) for field in _schema)
return _schema if _schema is not None else []
@staticmethod
def _build_metadata(info: DatasetInfo, fingerprint: Optional[str] = None) -> Dict[str, str]:
info_keys = ["features"] # we can add support for more DatasetInfo keys in the future
info_as_dict = asdict(info)
metadata = {}
metadata["info"] = {key: info_as_dict[key] for key in info_keys}
if fingerprint is not None:
metadata["fingerprint"] = fingerprint
return {"huggingface": json.dumps(metadata)}
def write_examples_on_file(self):
"""Write stored examples from the write-pool of examples. It makes a table out of the examples and write it."""
if not self.current_examples:
return
# order the columns properly
cols = (
[col for col in self.schema.names if col in self.current_examples[0][0]]
+ [col for col in self.current_examples[0][0].keys() if col not in self.schema.names]
if self.schema
else self.current_examples[0][0].keys()
)
batch_examples = {}
for col in cols:
# We use row[0][col] since current_examples contains (example, key) tuples.
# Morever, examples could be Arrow arrays of 1 element.
# This can happen in `.map()` when we want to re-write the same Arrow data
if all(isinstance(row[0][col], (pa.Array, pa.ChunkedArray)) for row in self.current_examples):
arrays = [row[0][col] for row in self.current_examples]
batch_examples[col] = array_concat(arrays)
else:
batch_examples[col] = [
row[0][col].to_pylist()[0] if isinstance(row[0][col], (pa.Array, pa.ChunkedArray)) else row[0][col]
for row in self.current_examples
]
self.write_batch(batch_examples=batch_examples)
self.current_examples = []
def write_rows_on_file(self):
"""Write stored rows from the write-pool of rows. It concatenates the single-row tables and it writes the resulting table."""
if not self.current_rows:
return
table = pa.concat_tables(self.current_rows)
self.write_table(table)
self.current_rows = []
def write(
self,
example: Dict[str, Any],
key: Optional[Union[str, int, bytes]] = None,
writer_batch_size: Optional[int] = None,
):
"""Add a given (Example,Key) pair to the write-pool of examples which is written to file.
Args:
example: the Example to add.
key: Optional, a unique identifier(str, int or bytes) associated with each example
"""
# Utilize the keys and duplicate checking when `self._check_duplicates` is passed True
if self._check_duplicates:
# Create unique hash from key and store as (key, example) pairs
hash = self._hasher.hash(key)
self.current_examples.append((example, hash))
# Maintain record of keys and their respective hashes for checking duplicates
self.hkey_record.append((hash, key))
else:
# Store example as a tuple so as to keep the structure of `self.current_examples` uniform
self.current_examples.append((example, ""))
if writer_batch_size is None:
writer_batch_size = self.writer_batch_size
if writer_batch_size is not None and len(self.current_examples) >= writer_batch_size:
if self._check_duplicates:
self.check_duplicate_keys()
# Re-intializing to empty list for next batch
self.hkey_record = []
self.write_examples_on_file()
def check_duplicate_keys(self):
"""Raises error if duplicates found in a batch"""
tmp_record = set()
for hash, key in self.hkey_record:
if hash in tmp_record:
duplicate_key_indices = [
str(self._num_examples + index)
for index, (duplicate_hash, _) in enumerate(self.hkey_record)
if duplicate_hash == hash
]
raise DuplicatedKeysError(key, duplicate_key_indices)
else:
tmp_record.add(hash)
def write_row(self, row: pa.Table, writer_batch_size: Optional[int] = None):
"""Add a given single-row Table to the write-pool of rows which is written to file.
Args:
row: the row to add.
"""
if len(row) != 1:
raise ValueError(f"Only single-row pyarrow tables are allowed but got table with {len(row)} rows.")
self.current_rows.append(row)
if writer_batch_size is None:
writer_batch_size = self.writer_batch_size
if writer_batch_size is not None and len(self.current_rows) >= writer_batch_size:
self.write_rows_on_file()
def write_batch(
self,
batch_examples: Dict[str, List],
writer_batch_size: Optional[int] = None,
):
"""Write a batch of Example to file.
Ignores the batch if it appears to be empty,
preventing a potential schema update of unknown types.
Args:
batch_examples: the batch of examples to add.
"""
if batch_examples and len(next(iter(batch_examples.values()))) == 0:
return
features = None if self.pa_writer is None and self.update_features else self._features
try_features = self._features if self.pa_writer is None and self.update_features else None
arrays = []
inferred_features = Features()
cols = (
[col for col in self.schema.names if col in batch_examples]
+ [col for col in batch_examples.keys() if col not in self.schema.names]
if self.schema
else batch_examples.keys()
)
for col in cols:
col_values = batch_examples[col]
col_type = features[col] if features else None
if isinstance(col_values, (pa.Array, pa.ChunkedArray)):
array = cast_array_to_feature(col_values, col_type) if col_type is not None else col_values
arrays.append(array)
inferred_features[col] = generate_from_arrow_type(col_values.type)
else:
col_try_type = try_features[col] if try_features is not None and col in try_features else None
typed_sequence = OptimizedTypedSequence(col_values, type=col_type, try_type=col_try_type, col=col)
arrays.append(pa.array(typed_sequence))
inferred_features[col] = typed_sequence.get_inferred_type()
schema = inferred_features.arrow_schema if self.pa_writer is None else self.schema
pa_table = pa.Table.from_arrays(arrays, schema=schema)
self.write_table(pa_table, writer_batch_size)
def write_table(self, pa_table: pa.Table, writer_batch_size: Optional[int] = None):
"""Write a Table to file.
Args:
example: the Table to add.
"""
if writer_batch_size is None:
writer_batch_size = self.writer_batch_size
if self.pa_writer is None:
self._build_writer(inferred_schema=pa_table.schema)
pa_table = pa_table.combine_chunks()
pa_table = table_cast(pa_table, self._schema)
if self.embed_local_files:
pa_table = embed_table_storage(pa_table)
self._num_bytes += pa_table.nbytes
self._num_examples += pa_table.num_rows
self.pa_writer.write_table(pa_table, writer_batch_size)
def finalize(self, close_stream=True):
self.write_rows_on_file()
# In case current_examples < writer_batch_size, but user uses finalize()
if self._check_duplicates:
self.check_duplicate_keys()
# Re-intializing to empty list for next batch
self.hkey_record = []
self.write_examples_on_file()
# If schema is known, infer features even if no examples were written
if self.pa_writer is None and self.schema:
self._build_writer(self.schema)
if self.pa_writer is not None:
self.pa_writer.close()
self.pa_writer = None
if close_stream:
self.stream.close()
else:
if close_stream:
self.stream.close()
raise SchemaInferenceError("Please pass `features` or at least one example when writing data")
logger.debug(
f"Done writing {self._num_examples} {self.unit} in {self._num_bytes} bytes {self._path if self._path else ''}."
)
return self._num_examples, self._num_bytes
class ParquetWriter(ArrowWriter):
_WRITER_CLASS = pq.ParquetWriter
class BeamWriter:
"""
Shuffles and writes Examples to Arrow files.
The Arrow files are converted from Parquet files that are the output of Apache Beam pipelines.
"""
def __init__(
self,
features: Optional[Features] = None,
schema: Optional[pa.Schema] = None,
path: Optional[str] = None,
namespace: Optional[str] = None,
cache_dir: Optional[str] = None,
):
if features is None and schema is None:
raise ValueError("At least one of features and schema must be provided.")
if path is None:
raise ValueError("Path must be provided.")
if features is not None:
self._features: Features = features
self._schema: pa.Schema = features.arrow_schema
else:
self._schema: pa.Schema = schema
self._features: Features = Features.from_arrow_schema(schema)
self._path = path
self._parquet_path = os.path.splitext(path)[0] # remove extension
self._namespace = namespace or "default"
self._num_examples = None
self._cache_dir = cache_dir or config.HF_DATASETS_CACHE
def write_from_pcollection(self, pcoll_examples):
"""Add the final steps of the beam pipeline: write to parquet files."""
import apache_beam as beam
def inc_num_examples(example):
beam.metrics.Metrics.counter(self._namespace, "num_examples").inc()
# count examples
_ = pcoll_examples | "Count N. Examples" >> beam.Map(inc_num_examples)
# save dataset
return (
pcoll_examples
| "Get values" >> beam.Values()
| "Save to parquet"
>> beam.io.parquetio.WriteToParquet(
self._parquet_path, self._schema, shard_name_template="-SSSSS-of-NNNNN.parquet"
)
)
def finalize(self, metrics_query_result: dict):
"""
Run after the pipeline has finished.
It converts the resulting parquet files to arrow and it completes the info from the pipeline metrics.
Args:
metrics_query_result: `dict` obtained from pipeline_results.metrics().query(m_filter). Make sure
that the filter keeps only the metrics for the considered split, under the namespace `split_name`.
"""
import apache_beam as beam
from .utils import beam_utils
# Beam FileSystems require the system's path separator in the older versions
fs, _, [parquet_path] = fsspec.get_fs_token_paths(self._parquet_path)
parquet_path = str(Path(parquet_path)) if not is_remote_filesystem(fs) else fs.unstrip_protocol(parquet_path)
shards_metadata = list(beam.io.filesystems.FileSystems.match([parquet_path + "*.parquet"])[0].metadata_list)
shards = [metadata.path for metadata in shards_metadata]
num_bytes = sum([metadata.size_in_bytes for metadata in shards_metadata])
shard_lengths = get_parquet_lengths(shards)
# Convert to arrow
if self._path.endswith(".arrow"):
logger.info(f"Converting parquet files {self._parquet_path} to arrow {self._path}")
shards = [
metadata.path
for metadata in beam.io.filesystems.FileSystems.match([parquet_path + "*.parquet"])[0].metadata_list
]
try: # stream conversion
num_bytes = 0
for shard in hf_tqdm(shards, unit="shards"):
with beam.io.filesystems.FileSystems.open(shard) as source:
with beam.io.filesystems.FileSystems.create(
shard.replace(".parquet", ".arrow")
) as destination:
shard_num_bytes, _ = parquet_to_arrow(source, destination)
num_bytes += shard_num_bytes
except OSError as e: # broken pipe can happen if the connection is unstable, do local conversion instead
if e.errno != errno.EPIPE: # not a broken pipe
raise
logger.warning(
"Broken Pipe during stream conversion from parquet to arrow. Using local convert instead"
)
local_convert_dir = os.path.join(self._cache_dir, "beam_convert")
os.makedirs(local_convert_dir, exist_ok=True)
num_bytes = 0
for shard in hf_tqdm(shards, unit="shards"):
local_parquet_path = os.path.join(local_convert_dir, hash_url_to_filename(shard) + ".parquet")
beam_utils.download_remote_to_local(shard, local_parquet_path)
local_arrow_path = local_parquet_path.replace(".parquet", ".arrow")
shard_num_bytes, _ = parquet_to_arrow(local_parquet_path, local_arrow_path)
num_bytes += shard_num_bytes
remote_arrow_path = shard.replace(".parquet", ".arrow")
beam_utils.upload_local_to_remote(local_arrow_path, remote_arrow_path)
# Save metrics
counters_dict = {metric.key.metric.name: metric.result for metric in metrics_query_result["counters"]}
self._num_examples = counters_dict["num_examples"]
self._num_bytes = num_bytes
self._shard_lengths = shard_lengths
return self._num_examples, self._num_bytes
def get_parquet_lengths(sources) -> List[int]:
shard_lengths = []
for source in hf_tqdm(sources, unit="parquet files"):
parquet_file = pa.parquet.ParquetFile(source)
shard_lengths.append(parquet_file.metadata.num_rows)
return shard_lengths
def parquet_to_arrow(source, destination) -> List[int]:
"""Convert parquet file to arrow file. Inputs can be str paths or file-like objects"""
stream = None if isinstance(destination, str) else destination
with ArrowWriter(path=destination, stream=stream) as writer:
parquet_file = pa.parquet.ParquetFile(source)
for record_batch in parquet_file.iter_batches():
pa_table = pa.Table.from_batches([record_batch])
writer.write_table(pa_table)
num_bytes, num_examples = writer.finalize()
return num_bytes, num_examples
| 0
|
hf_public_repos/datasets/src
|
hf_public_repos/datasets/src/datasets/metric.py
|
# Copyright 2020 The HuggingFace Datasets Authors
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# Lint as: python3
""" Metrics base class."""
import os
import types
import uuid
from typing import Any, Dict, List, Optional, Tuple, Union
import numpy as np
import pyarrow as pa
from filelock import BaseFileLock, Timeout
from . import config
from .arrow_dataset import Dataset
from .arrow_reader import ArrowReader
from .arrow_writer import ArrowWriter
from .download.download_config import DownloadConfig
from .download.download_manager import DownloadManager
from .features import Features
from .info import DatasetInfo, MetricInfo
from .naming import camelcase_to_snakecase
from .utils._filelock import FileLock
from .utils.deprecation_utils import deprecated
from .utils.logging import get_logger
from .utils.py_utils import copyfunc, temp_seed
logger = get_logger(__name__)
class FileFreeLock(BaseFileLock):
"""Thread lock until a file **cannot** be locked"""
def __init__(self, lock_file, *args, **kwargs):
self.filelock = FileLock(lock_file)
super().__init__(self.filelock.lock_file, *args, **kwargs)
def _acquire(self):
try:
self.filelock.acquire(timeout=0.01, poll_intervall=0.02) # Try to lock once
except Timeout:
# We couldn't acquire the lock, the file is locked!
self._context.lock_file_fd = self.filelock.lock_file
else:
# We were able to acquire the lock, the file is not yet locked!
self.filelock.release()
self._context.lock_file_fd = None
def _release(self):
self._context.lock_file_fd = None
# lists - summarize long lists similarly to NumPy
# arrays/tensors - let the frameworks control formatting
def summarize_if_long_list(obj):
if not type(obj) == list or len(obj) <= 6: # noqa: E721
return f"{obj}"
def format_chunk(chunk):
return ", ".join(repr(x) for x in chunk)
return f"[{format_chunk(obj[:3])}, ..., {format_chunk(obj[-3:])}]"
class MetricInfoMixin:
"""This base class exposes some attributes of MetricInfo
at the base level of the Metric for easy access.
<Deprecated version="2.5.0">
Use the new library 🤗 Evaluate instead: https://huggingface.co/docs/evaluate
</Deprecated>
"""
def __init__(self, info: MetricInfo):
self._metric_info = info
@property
def info(self):
""":class:`datasets.MetricInfo` object containing all the metadata in the metric."""
return self._metric_info
@property
def name(self) -> str:
return self._metric_info.metric_name
@property
def experiment_id(self) -> Optional[str]:
return self._metric_info.experiment_id
@property
def description(self) -> str:
return self._metric_info.description
@property
def citation(self) -> str:
return self._metric_info.citation
@property
def features(self) -> Features:
return self._metric_info.features
@property
def inputs_description(self) -> str:
return self._metric_info.inputs_description
@property
def homepage(self) -> Optional[str]:
return self._metric_info.homepage
@property
def license(self) -> str:
return self._metric_info.license
@property
def codebase_urls(self) -> Optional[List[str]]:
return self._metric_info.codebase_urls
@property
def reference_urls(self) -> Optional[List[str]]:
return self._metric_info.reference_urls
@property
def streamable(self) -> bool:
return self._metric_info.streamable
@property
def format(self) -> Optional[str]:
return self._metric_info.format
class Metric(MetricInfoMixin):
"""A Metric is the base class and common API for all metrics.
<Deprecated version="2.5.0">
Use the new library 🤗 Evaluate instead: https://huggingface.co/docs/evaluate
</Deprecated>
Args:
config_name (``str``): This is used to define a hash specific to a metrics computation script and prevents the metric's data
to be overridden when the metric loading script is modified.
keep_in_memory (:obj:`bool`): keep all predictions and references in memory. Not possible in distributed settings.
cache_dir (``str``): Path to a directory in which temporary prediction/references data will be stored.
The data directory should be located on a shared file-system in distributed setups.
num_process (``int``): specify the total number of nodes in a distributed settings.
This is useful to compute metrics in distributed setups (in particular non-additive metrics like F1).
process_id (``int``): specify the id of the current process in a distributed setup (between 0 and num_process-1)
This is useful to compute metrics in distributed setups (in particular non-additive metrics like F1).
seed (:obj:`int`, optional): If specified, this will temporarily set numpy's random seed when :func:`datasets.Metric.compute` is run.
experiment_id (``str``): A specific experiment id. This is used if several distributed evaluations share the same file system.
This is useful to compute metrics in distributed setups (in particular non-additive metrics like F1).
max_concurrent_cache_files (``int``): Max number of concurrent metrics cache files (default 10000).
timeout (``Union[int, float]``): Timeout in second for distributed setting synchronization.
"""
@deprecated("Use the new library 🤗 Evaluate instead: https://huggingface.co/docs/evaluate")
def __init__(
self,
config_name: Optional[str] = None,
keep_in_memory: bool = False,
cache_dir: Optional[str] = None,
num_process: int = 1,
process_id: int = 0,
seed: Optional[int] = None,
experiment_id: Optional[str] = None,
max_concurrent_cache_files: int = 10000,
timeout: Union[int, float] = 100,
**kwargs,
):
# prepare info
self.config_name = config_name or "default"
info = self._info()
info.metric_name = camelcase_to_snakecase(self.__class__.__name__)
info.config_name = self.config_name
info.experiment_id = experiment_id or "default_experiment"
MetricInfoMixin.__init__(self, info) # For easy access on low level
# Safety checks on num_process and process_id
if not isinstance(process_id, int) or process_id < 0:
raise ValueError("'process_id' should be a number greater than 0")
if not isinstance(num_process, int) or num_process <= process_id:
raise ValueError("'num_process' should be a number greater than process_id")
if keep_in_memory and num_process != 1:
raise ValueError("Using 'keep_in_memory' is not possible in distributed setting (num_process > 1).")
self.num_process = num_process
self.process_id = process_id
self.max_concurrent_cache_files = max_concurrent_cache_files
self.keep_in_memory = keep_in_memory
self._data_dir_root = os.path.expanduser(cache_dir or config.HF_METRICS_CACHE)
self.data_dir = self._build_data_dir()
if seed is None:
_, seed, pos, *_ = np.random.get_state()
self.seed: int = seed[pos] if pos < 624 else seed[0]
else:
self.seed: int = seed
self.timeout: Union[int, float] = timeout
# Update 'compute' and 'add' docstring
# methods need to be copied otherwise it changes the docstrings of every instance
self.compute = types.MethodType(copyfunc(self.compute), self)
self.add_batch = types.MethodType(copyfunc(self.add_batch), self)
self.add = types.MethodType(copyfunc(self.add), self)
self.compute.__func__.__doc__ += self.info.inputs_description
self.add_batch.__func__.__doc__ += self.info.inputs_description
self.add.__func__.__doc__ += self.info.inputs_description
# self.arrow_schema = pa.schema(field for field in self.info.features.type)
self.buf_writer = None
self.writer = None
self.writer_batch_size = None
self.data = None
# This is the cache file we store our predictions/references in
# Keep it None for now so we can (cloud)pickle the object
self.cache_file_name = None
self.filelock = None
self.rendez_vous_lock = None
# This is all the cache files on which we have a lock when we are in a distributed setting
self.file_paths = None
self.filelocks = None
def __len__(self):
"""Return the number of examples (predictions or predictions/references pair)
currently stored in the metric's cache.
"""
return 0 if self.writer is None else len(self.writer)
def __repr__(self):
return (
f'Metric(name: "{self.name}", features: {self.features}, '
f'usage: """{self.inputs_description}""", '
f"stored examples: {len(self)})"
)
def _build_data_dir(self):
"""Path of this metric in cache_dir:
Will be:
self._data_dir_root/self.name/self.config_name/self.hash (if not none)/
If any of these element is missing or if ``with_version=False`` the corresponding subfolders are dropped.
"""
builder_data_dir = self._data_dir_root
builder_data_dir = os.path.join(builder_data_dir, self.name, self.config_name)
os.makedirs(builder_data_dir, exist_ok=True)
return builder_data_dir
def _create_cache_file(self, timeout=1) -> Tuple[str, FileLock]:
"""Create a new cache file. If the default cache file is used, we generated a new hash."""
file_path = os.path.join(self.data_dir, f"{self.experiment_id}-{self.num_process}-{self.process_id}.arrow")
filelock = None
for i in range(self.max_concurrent_cache_files):
filelock = FileLock(file_path + ".lock")
try:
filelock.acquire(timeout=timeout)
except Timeout:
# If we have reached the max number of attempts or we are not allow to find a free name (distributed setup)
# We raise an error
if self.num_process != 1:
raise ValueError(
f"Error in _create_cache_file: another metric instance is already using the local cache file at {file_path}. "
f"Please specify an experiment_id (currently: {self.experiment_id}) to avoid collision "
f"between distributed metric instances."
) from None
if i == self.max_concurrent_cache_files - 1:
raise ValueError(
f"Cannot acquire lock, too many metric instance are operating concurrently on this file system."
f"You should set a larger value of max_concurrent_cache_files when creating the metric "
f"(current value is {self.max_concurrent_cache_files})."
) from None
# In other cases (allow to find new file name + not yet at max num of attempts) we can try to sample a new hashing name.
file_uuid = str(uuid.uuid4())
file_path = os.path.join(
self.data_dir, f"{self.experiment_id}-{file_uuid}-{self.num_process}-{self.process_id}.arrow"
)
else:
break
return file_path, filelock
def _get_all_cache_files(self) -> Tuple[List[str], List[FileLock]]:
"""Get a lock on all the cache files in a distributed setup.
We wait for timeout second to let all the distributed node finish their tasks (default is 100 seconds).
"""
if self.num_process == 1:
if self.cache_file_name is None:
raise ValueError(
"Metric cache file doesn't exist. Please make sure that you call `add` or `add_batch` "
"at least once before calling `compute`."
)
file_paths = [self.cache_file_name]
else:
file_paths = [
os.path.join(self.data_dir, f"{self.experiment_id}-{self.num_process}-{process_id}.arrow")
for process_id in range(self.num_process)
]
# Let's acquire a lock on each process files to be sure they are finished writing
filelocks = []
for process_id, file_path in enumerate(file_paths):
if process_id == 0: # process 0 already has its lock file
filelocks.append(self.filelock)
else:
filelock = FileLock(file_path + ".lock")
try:
filelock.acquire(timeout=self.timeout)
except Timeout:
raise ValueError(
f"Cannot acquire lock on cached file {file_path} for process {process_id}."
) from None
else:
filelocks.append(filelock)
return file_paths, filelocks
def _check_all_processes_locks(self):
expected_lock_file_names = [
os.path.join(self.data_dir, f"{self.experiment_id}-{self.num_process}-{process_id}.arrow.lock")
for process_id in range(self.num_process)
]
for expected_lock_file_name in expected_lock_file_names:
nofilelock = FileFreeLock(expected_lock_file_name)
try:
nofilelock.acquire(timeout=self.timeout)
except Timeout:
raise ValueError(
f"Expected to find locked file {expected_lock_file_name} from process {self.process_id} but it doesn't exist."
) from None
else:
nofilelock.release()
def _check_rendez_vous(self):
expected_lock_file_name = os.path.join(self.data_dir, f"{self.experiment_id}-{self.num_process}-0.arrow.lock")
nofilelock = FileFreeLock(expected_lock_file_name)
try:
nofilelock.acquire(timeout=self.timeout)
except Timeout:
raise ValueError(
f"Expected to find locked file {expected_lock_file_name} from process {self.process_id} but it doesn't exist."
) from None
else:
nofilelock.release()
lock_file_name = os.path.join(self.data_dir, f"{self.experiment_id}-{self.num_process}-rdv.lock")
rendez_vous_lock = FileLock(lock_file_name)
try:
rendez_vous_lock.acquire(timeout=self.timeout)
except Timeout:
raise ValueError(f"Couldn't acquire lock on {lock_file_name} from process {self.process_id}.") from None
else:
rendez_vous_lock.release()
def _finalize(self):
"""Close all the writing process and load/gather the data
from all the nodes if main node or all_process is True.
"""
if self.writer is not None:
self.writer.finalize()
self.writer = None
# release the locks of the processes > 0 so that process 0 can lock them to read + delete the data
if self.filelock is not None and self.process_id > 0:
self.filelock.release()
if self.keep_in_memory:
# Read the predictions and references
reader = ArrowReader(path=self.data_dir, info=DatasetInfo(features=self.features))
self.data = Dataset.from_buffer(self.buf_writer.getvalue())
elif self.process_id == 0:
# Let's acquire a lock on each node files to be sure they are finished writing
file_paths, filelocks = self._get_all_cache_files()
# Read the predictions and references
try:
reader = ArrowReader(path="", info=DatasetInfo(features=self.features))
self.data = Dataset(**reader.read_files([{"filename": f} for f in file_paths]))
except FileNotFoundError:
raise ValueError(
"Error in finalize: another metric instance is already using the local cache file. "
"Please specify an experiment_id to avoid collision between distributed metric instances."
) from None
# Store file paths and locks and we will release/delete them after the computation.
self.file_paths = file_paths
self.filelocks = filelocks
def compute(self, *, predictions=None, references=None, **kwargs) -> Optional[dict]:
"""Compute the metrics.
Usage of positional arguments is not allowed to prevent mistakes.
Args:
predictions (list/array/tensor, optional): Predictions.
references (list/array/tensor, optional): References.
**kwargs (optional): Keyword arguments that will be forwarded to the metrics :meth:`_compute`
method (see details in the docstring).
Return:
dict or None
- Dictionary with the metrics if this metric is run on the main process (``process_id == 0``).
- None if the metric is not run on the main process (``process_id != 0``).
Example:
```py
>>> from datasets import load_metric
>>> metric = load_metric("accuracy")
>>> accuracy = metric.compute(predictions=model_prediction, references=labels)
```
"""
all_kwargs = {"predictions": predictions, "references": references, **kwargs}
if predictions is None and references is None:
missing_kwargs = {k: None for k in self.features if k not in all_kwargs}
all_kwargs.update(missing_kwargs)
else:
missing_inputs = [k for k in self.features if k not in all_kwargs]
if missing_inputs:
raise ValueError(
f"Metric inputs are missing: {missing_inputs}. All required inputs are {list(self.features)}"
)
inputs = {input_name: all_kwargs[input_name] for input_name in self.features}
compute_kwargs = {k: kwargs[k] for k in kwargs if k not in self.features}
if any(v is not None for v in inputs.values()):
self.add_batch(**inputs)
self._finalize()
self.cache_file_name = None
self.filelock = None
if self.process_id == 0:
self.data.set_format(type=self.info.format)
inputs = {input_name: self.data[input_name] for input_name in self.features}
with temp_seed(self.seed):
output = self._compute(**inputs, **compute_kwargs)
if self.buf_writer is not None:
self.buf_writer = None
del self.data
self.data = None
else:
# Release locks and delete all the cache files. Process 0 is released last.
for filelock, file_path in reversed(list(zip(self.filelocks, self.file_paths))):
logger.info(f"Removing {file_path}")
del self.data
self.data = None
del self.writer
self.writer = None
os.remove(file_path)
filelock.release()
return output
else:
return None
def add_batch(self, *, predictions=None, references=None, **kwargs):
"""Add a batch of predictions and references for the metric's stack.
Args:
predictions (list/array/tensor, optional): Predictions.
references (list/array/tensor, optional): References.
Example:
```py
>>> from datasets import load_metric
>>> metric = load_metric("accuracy")
>>> metric.add_batch(predictions=model_prediction, references=labels)
```
"""
bad_inputs = [input_name for input_name in kwargs if input_name not in self.features]
if bad_inputs:
raise ValueError(f"Bad inputs for metric: {bad_inputs}. All required inputs are {list(self.features)}")
batch = {"predictions": predictions, "references": references, **kwargs}
batch = {intput_name: batch[intput_name] for intput_name in self.features}
batch = self.info.features.encode_batch(batch)
if self.writer is None:
self._init_writer()
try:
self.writer.write_batch(batch)
except pa.ArrowInvalid:
if any(len(batch[c]) != len(next(iter(batch.values()))) for c in batch):
col0 = next(iter(batch))
bad_col = [c for c in batch if len(batch[c]) != len(batch[col0])][0]
error_msg = (
f"Mismatch in the number of {col0} ({len(batch[col0])}) and {bad_col} ({len(batch[bad_col])})"
)
elif sorted(self.features) != ["references", "predictions"]:
error_msg = f"Metric inputs don't match the expected format.\n" f"Expected format: {self.features},\n"
error_msg_inputs = ",\n".join(
f"Input {input_name}: {summarize_if_long_list(batch[input_name])}" for input_name in self.features
)
error_msg += error_msg_inputs
else:
error_msg = (
f"Predictions and/or references don't match the expected format.\n"
f"Expected format: {self.features},\n"
f"Input predictions: {summarize_if_long_list(predictions)},\n"
f"Input references: {summarize_if_long_list(references)}"
)
raise ValueError(error_msg) from None
def add(self, *, prediction=None, reference=None, **kwargs):
"""Add one prediction and reference for the metric's stack.
Args:
prediction (list/array/tensor, optional): Predictions.
reference (list/array/tensor, optional): References.
Example:
```py
>>> from datasets import load_metric
>>> metric = load_metric("accuracy")
>>> metric.add(predictions=model_predictions, references=labels)
```
"""
bad_inputs = [input_name for input_name in kwargs if input_name not in self.features]
if bad_inputs:
raise ValueError(f"Bad inputs for metric: {bad_inputs}. All required inputs are {list(self.features)}")
example = {"predictions": prediction, "references": reference, **kwargs}
example = {intput_name: example[intput_name] for intput_name in self.features}
example = self.info.features.encode_example(example)
if self.writer is None:
self._init_writer()
try:
self.writer.write(example)
except pa.ArrowInvalid:
error_msg = f"Metric inputs don't match the expected format.\n" f"Expected format: {self.features},\n"
error_msg_inputs = ",\n".join(
f"Input {input_name}: {summarize_if_long_list(example[input_name])}" for input_name in self.features
)
error_msg += error_msg_inputs
raise ValueError(error_msg) from None
def _init_writer(self, timeout=1):
if self.num_process > 1:
if self.process_id == 0:
file_path = os.path.join(self.data_dir, f"{self.experiment_id}-{self.num_process}-rdv.lock")
self.rendez_vous_lock = FileLock(file_path)
try:
self.rendez_vous_lock.acquire(timeout=timeout)
except TimeoutError:
raise ValueError(
f"Error in _init_writer: another metric instance is already using the local cache file at {file_path}. "
f"Please specify an experiment_id (currently: {self.experiment_id}) to avoid collision "
f"between distributed metric instances."
) from None
if self.keep_in_memory:
self.buf_writer = pa.BufferOutputStream()
self.writer = ArrowWriter(
features=self.info.features, stream=self.buf_writer, writer_batch_size=self.writer_batch_size
)
else:
self.buf_writer = None
# Get cache file name and lock it
if self.cache_file_name is None or self.filelock is None:
cache_file_name, filelock = self._create_cache_file() # get ready
self.cache_file_name = cache_file_name
self.filelock = filelock
self.writer = ArrowWriter(
features=self.info.features, path=self.cache_file_name, writer_batch_size=self.writer_batch_size
)
# Setup rendez-vous here if
if self.num_process > 1:
if self.process_id == 0:
self._check_all_processes_locks() # wait for everyone to be ready
self.rendez_vous_lock.release() # let everyone go
else:
self._check_rendez_vous() # wait for master to be ready and to let everyone go
def _info(self) -> MetricInfo:
"""Construct the MetricInfo object. See `MetricInfo` for details.
Warning: This function is only called once and the result is cached for all
following .info() calls.
Returns:
info: (MetricInfo) The metrics information
"""
raise NotImplementedError
def download_and_prepare(
self,
download_config: Optional[DownloadConfig] = None,
dl_manager: Optional[DownloadManager] = None,
):
"""Downloads and prepares dataset for reading.
Args:
download_config (:class:`DownloadConfig`, optional): Specific download configuration parameters.
dl_manager (:class:`DownloadManager`, optional): Specific download manager to use.
"""
if dl_manager is None:
if download_config is None:
download_config = DownloadConfig()
download_config.cache_dir = os.path.join(self.data_dir, "downloads")
download_config.force_download = False
dl_manager = DownloadManager(
dataset_name=self.name, download_config=download_config, data_dir=self.data_dir
)
self._download_and_prepare(dl_manager)
def _download_and_prepare(self, dl_manager):
"""Downloads and prepares resources for the metric.
This is the internal implementation to overwrite called when user calls
`download_and_prepare`. It should download all required resources for the metric.
Args:
dl_manager (:class:`DownloadManager`): `DownloadManager` used to download and cache data.
"""
return None
def _compute(self, *, predictions=None, references=None, **kwargs) -> Dict[str, Any]:
"""This method defines the common API for all the metrics in the library"""
raise NotImplementedError
def __del__(self):
if hasattr(self, "filelock") and self.filelock is not None:
self.filelock.release()
if hasattr(self, "rendez_vous_lock") and self.rendez_vous_lock is not None:
self.rendez_vous_lock.release()
if hasattr(self, "writer"): # in case it was already deleted
del self.writer
if hasattr(self, "data"): # in case it was already deleted
del self.data
| 0
|
hf_public_repos/datasets/src
|
hf_public_repos/datasets/src/datasets/data_files.py
|
import os
import re
from functools import partial
from glob import has_magic
from pathlib import Path, PurePath
from typing import Callable, Dict, List, Optional, Set, Tuple, Union
import huggingface_hub
from fsspec import get_fs_token_paths
from fsspec.implementations.http import HTTPFileSystem
from huggingface_hub import HfFileSystem
from packaging import version
from tqdm.contrib.concurrent import thread_map
from . import config
from .download import DownloadConfig
from .download.streaming_download_manager import _prepare_path_and_storage_options, xbasename, xjoin
from .splits import Split
from .utils import logging
from .utils import tqdm as hf_tqdm
from .utils.file_utils import is_local_path, is_relative_path
from .utils.py_utils import glob_pattern_to_regex, string_to_dict
SANITIZED_DEFAULT_SPLIT = str(Split.TRAIN)
logger = logging.get_logger(__name__)
class Url(str):
pass
class EmptyDatasetError(FileNotFoundError):
pass
SPLIT_PATTERN_SHARDED = "data/{split}-[0-9][0-9][0-9][0-9][0-9]-of-[0-9][0-9][0-9][0-9][0-9]*.*"
SPLIT_KEYWORDS = {
Split.TRAIN: ["train", "training"],
Split.VALIDATION: ["validation", "valid", "dev", "val"],
Split.TEST: ["test", "testing", "eval", "evaluation"],
}
NON_WORDS_CHARS = "-._ 0-9"
if config.FSSPEC_VERSION < version.parse("2023.9.0"):
KEYWORDS_IN_PATH_NAME_BASE_PATTERNS = ["{keyword}[{sep}/]**", "**[{sep}/]{keyword}[{sep}/]**"]
else:
KEYWORDS_IN_PATH_NAME_BASE_PATTERNS = ["{keyword}[{sep}/]**", "**/*[{sep}/]{keyword}[{sep}/]**"]
DEFAULT_SPLITS = [Split.TRAIN, Split.VALIDATION, Split.TEST]
DEFAULT_PATTERNS_SPLIT_IN_PATH_NAME = {
split: [
pattern.format(keyword=keyword, sep=NON_WORDS_CHARS)
for keyword in SPLIT_KEYWORDS[split]
for pattern in KEYWORDS_IN_PATH_NAME_BASE_PATTERNS
]
for split in DEFAULT_SPLITS
}
DEFAULT_PATTERNS_ALL = {
Split.TRAIN: ["**"],
}
ALL_SPLIT_PATTERNS = [SPLIT_PATTERN_SHARDED]
ALL_DEFAULT_PATTERNS = [
DEFAULT_PATTERNS_SPLIT_IN_PATH_NAME,
DEFAULT_PATTERNS_ALL,
]
if config.FSSPEC_VERSION < version.parse("2023.9.0"):
METADATA_PATTERNS = [
"metadata.csv",
"**/metadata.csv",
"metadata.jsonl",
"**/metadata.jsonl",
] # metadata file for ImageFolder and AudioFolder
else:
METADATA_PATTERNS = [
"**/metadata.csv",
"**/metadata.jsonl",
] # metadata file for ImageFolder and AudioFolder
WILDCARD_CHARACTERS = "*[]"
FILES_TO_IGNORE = [
"README.md",
"config.json",
"dataset_info.json",
"dataset_infos.json",
"dummy_data.zip",
"dataset_dict.json",
]
def contains_wildcards(pattern: str) -> bool:
return any(wilcard_character in pattern for wilcard_character in WILDCARD_CHARACTERS)
def sanitize_patterns(patterns: Union[Dict, List, str]) -> Dict[str, Union[List[str], "DataFilesList"]]:
"""
Take the data_files patterns from the user, and format them into a dictionary.
Each key is the name of the split, and each value is a list of data files patterns (paths or urls).
The default split is "train".
Returns:
patterns: dictionary of split_name -> list of patterns
"""
if isinstance(patterns, dict):
return {str(key): value if isinstance(value, list) else [value] for key, value in patterns.items()}
elif isinstance(patterns, str):
return {SANITIZED_DEFAULT_SPLIT: [patterns]}
elif isinstance(patterns, list):
if any(isinstance(pattern, dict) for pattern in patterns):
for pattern in patterns:
if not (
isinstance(pattern, dict)
and len(pattern) == 2
and "split" in pattern
and isinstance(pattern.get("path"), (str, list))
):
raise ValueError(
f"Expected each split to have a 'path' key which can be a string or a list of strings, but got {pattern}"
)
splits = [pattern["split"] for pattern in patterns]
if len(set(splits)) != len(splits):
raise ValueError(f"Some splits are duplicated in data_files: {splits}")
return {
str(pattern["split"]): pattern["path"] if isinstance(pattern["path"], list) else [pattern["path"]]
for pattern in patterns
}
else:
return {SANITIZED_DEFAULT_SPLIT: patterns}
else:
return sanitize_patterns(list(patterns))
def _is_inside_unrequested_special_dir(matched_rel_path: str, pattern: str) -> bool:
"""
When a path matches a pattern, we additionnally check if it's inside a special directory
we ignore by default (if it starts with a double underscore).
Users can still explicitly request a filepath inside such a directory if "__pycache__" is
mentioned explicitly in the requested pattern.
Some examples:
base directory:
./
└── __pycache__
└── b.txt
>>> _is_inside_unrequested_special_dir("__pycache__/b.txt", "**")
True
>>> _is_inside_unrequested_special_dir("__pycache__/b.txt", "*/b.txt")
True
>>> _is_inside_unrequested_special_dir("__pycache__/b.txt", "__pycache__/*")
False
>>> _is_inside_unrequested_special_dir("__pycache__/b.txt", "__*/*")
False
"""
# We just need to check if every special directories from the path is present explicly in the pattern.
# Since we assume that the path matches the pattern, it's equivalent to counting that both
# the parent path and the parent pattern have the same number of special directories.
data_dirs_to_ignore_in_path = [part for part in PurePath(matched_rel_path).parent.parts if part.startswith("__")]
data_dirs_to_ignore_in_pattern = [part for part in PurePath(pattern).parent.parts if part.startswith("__")]
return len(data_dirs_to_ignore_in_path) != len(data_dirs_to_ignore_in_pattern)
def _is_unrequested_hidden_file_or_is_inside_unrequested_hidden_dir(matched_rel_path: str, pattern: str) -> bool:
"""
When a path matches a pattern, we additionnally check if it's a hidden file or if it's inside
a hidden directory we ignore by default, i.e. if the file name or a parent directory name starts with a dot.
Users can still explicitly request a filepath that is hidden or is inside a hidden directory
if the hidden part is mentioned explicitly in the requested pattern.
Some examples:
base directory:
./
└── .hidden_file.txt
>>> _is_unrequested_hidden_file_or_is_inside_unrequested_hidden_dir(".hidden_file.txt", "**")
True
>>> _is_unrequested_hidden_file_or_is_inside_unrequested_hidden_dir(".hidden_file.txt", ".*")
False
base directory:
./
└── .hidden_dir
└── a.txt
>>> _is_unrequested_hidden_file_or_is_inside_unrequested_hidden_dir(".hidden_dir/a.txt", "**")
True
>>> _is_unrequested_hidden_file_or_is_inside_unrequested_hidden_dir(".hidden_dir/a.txt", ".*/*")
False
>>> _is_unrequested_hidden_file_or_is_inside_unrequested_hidden_dir(".hidden_dir/a.txt", ".hidden_dir/*")
False
base directory:
./
└── .hidden_dir
└── .hidden_file.txt
>>> _is_unrequested_hidden_file_or_is_inside_unrequested_hidden_dir(".hidden_dir/.hidden_file.txt", "**")
True
>>> _is_unrequested_hidden_file_or_is_inside_unrequested_hidden_dir(".hidden_dir/.hidden_file.txt", ".*/*")
True
>>> _is_unrequested_hidden_file_or_is_inside_unrequested_hidden_dir(".hidden_dir/.hidden_file.txt", ".*/.*")
False
>>> _is_unrequested_hidden_file_or_is_inside_unrequested_hidden_dir(".hidden_dir/.hidden_file.txt", ".hidden_dir/*")
True
>>> _is_unrequested_hidden_file_or_is_inside_unrequested_hidden_dir(".hidden_dir/.hidden_file.txt", ".hidden_dir/.*")
False
"""
# We just need to check if every hidden part from the path is present explicly in the pattern.
# Since we assume that the path matches the pattern, it's equivalent to counting that both
# the path and the pattern have the same number of hidden parts.
hidden_directories_in_path = [
part for part in PurePath(matched_rel_path).parts if part.startswith(".") and not set(part) == {"."}
]
hidden_directories_in_pattern = [
part for part in PurePath(pattern).parts if part.startswith(".") and not set(part) == {"."}
]
return len(hidden_directories_in_path) != len(hidden_directories_in_pattern)
def _get_data_files_patterns(pattern_resolver: Callable[[str], List[str]]) -> Dict[str, List[str]]:
"""
Get the default pattern from a directory or repository by testing all the supported patterns.
The first patterns to return a non-empty list of data files is returned.
In order, it first tests if SPLIT_PATTERN_SHARDED works, otherwise it tests the patterns in ALL_DEFAULT_PATTERNS.
"""
# first check the split patterns like data/{split}-00000-of-00001.parquet
for split_pattern in ALL_SPLIT_PATTERNS:
pattern = split_pattern.replace("{split}", "*")
try:
data_files = pattern_resolver(pattern)
except FileNotFoundError:
continue
if len(data_files) > 0:
splits: Set[str] = {
string_to_dict(xbasename(p), glob_pattern_to_regex(xbasename(split_pattern)))["split"]
for p in data_files
}
sorted_splits = [str(split) for split in DEFAULT_SPLITS if split in splits] + sorted(
splits - set(DEFAULT_SPLITS)
)
return {split: [split_pattern.format(split=split)] for split in sorted_splits}
# then check the default patterns based on train/valid/test splits
for patterns_dict in ALL_DEFAULT_PATTERNS:
non_empty_splits = []
for split, patterns in patterns_dict.items():
for pattern in patterns:
try:
data_files = pattern_resolver(pattern)
except FileNotFoundError:
continue
if len(data_files) > 0:
non_empty_splits.append(split)
break
if non_empty_splits:
return {split: patterns_dict[split] for split in non_empty_splits}
raise FileNotFoundError(f"Couldn't resolve pattern {pattern} with resolver {pattern_resolver}")
def _get_metadata_files_patterns(pattern_resolver: Callable[[str], List[str]]) -> List[str]:
"""
Get the supported metadata patterns from a directory or repository.
"""
non_empty_patterns = []
for pattern in METADATA_PATTERNS:
try:
metadata_files = pattern_resolver(pattern)
if len(metadata_files) > 0:
non_empty_patterns.append(pattern)
except FileNotFoundError:
pass
if non_empty_patterns:
return non_empty_patterns
raise FileNotFoundError(f"Couldn't resolve pattern {pattern} with resolver {pattern_resolver}")
def resolve_pattern(
pattern: str,
base_path: str,
allowed_extensions: Optional[List[str]] = None,
download_config: Optional[DownloadConfig] = None,
) -> List[str]:
"""
Resolve the paths and URLs of the data files from the pattern passed by the user.
You can use patterns to resolve multiple local files. Here are a few examples:
- *.csv to match all the CSV files at the first level
- **.csv to match all the CSV files at any level
- data/* to match all the files inside "data"
- data/** to match all the files inside "data" and its subdirectories
The patterns are resolved using the fsspec glob.
glob.glob, Path.glob, Path.match or fnmatch do not support ** with a prefix/suffix other than a forward slash /.
For instance, this means **.json is the same as *.json. On the contrary, the fsspec glob has no limits regarding the ** prefix/suffix,
resulting in **.json being equivalent to **/*.json.
More generally:
- '*' matches any character except a forward-slash (to match just the file or directory name)
- '**' matches any character including a forward-slash /
Hidden files and directories (i.e. whose names start with a dot) are ignored, unless they are explicitly requested.
The same applies to special directories that start with a double underscore like "__pycache__".
You can still include one if the pattern explicilty mentions it:
- to include a hidden file: "*/.hidden.txt" or "*/.*"
- to include a hidden directory: ".hidden/*" or ".*/*"
- to include a special directory: "__special__/*" or "__*/*"
Example::
>>> from datasets.data_files import resolve_pattern
>>> base_path = "."
>>> resolve_pattern("docs/**/*.py", base_path)
[/Users/mariosasko/Desktop/projects/datasets/docs/source/_config.py']
Args:
pattern (str): Unix pattern or paths or URLs of the data files to resolve.
The paths can be absolute or relative to base_path.
Remote filesystems using fsspec are supported, e.g. with the hf:// protocol.
base_path (str): Base path to use when resolving relative paths.
allowed_extensions (Optional[list], optional): White-list of file extensions to use. Defaults to None (all extensions).
For example: allowed_extensions=[".csv", ".json", ".txt", ".parquet"]
Returns:
List[str]: List of paths or URLs to the local or remote files that match the patterns.
"""
if is_relative_path(pattern):
pattern = xjoin(base_path, pattern)
elif is_local_path(pattern):
base_path = os.path.splitdrive(pattern)[0] + os.sep
else:
base_path = ""
pattern, storage_options = _prepare_path_and_storage_options(pattern, download_config=download_config)
fs, _, _ = get_fs_token_paths(pattern, storage_options=storage_options)
fs_base_path = base_path.split("::")[0].split("://")[-1] or fs.root_marker
fs_pattern = pattern.split("::")[0].split("://")[-1]
files_to_ignore = set(FILES_TO_IGNORE) - {xbasename(pattern)}
protocol = fs.protocol if isinstance(fs.protocol, str) else fs.protocol[0]
protocol_prefix = protocol + "://" if protocol != "file" else ""
matched_paths = [
filepath if filepath.startswith(protocol_prefix) else protocol_prefix + filepath
for filepath, info in fs.glob(pattern, detail=True).items()
if info["type"] == "file"
and (xbasename(filepath) not in files_to_ignore)
and not _is_inside_unrequested_special_dir(
os.path.relpath(filepath, fs_base_path), os.path.relpath(fs_pattern, fs_base_path)
)
and not _is_unrequested_hidden_file_or_is_inside_unrequested_hidden_dir(
os.path.relpath(filepath, fs_base_path), os.path.relpath(fs_pattern, fs_base_path)
)
] # ignore .ipynb and __pycache__, but keep /../
if allowed_extensions is not None:
out = [
filepath
for filepath in matched_paths
if any("." + suffix in allowed_extensions for suffix in xbasename(filepath).split(".")[1:])
]
if len(out) < len(matched_paths):
invalid_matched_files = list(set(matched_paths) - set(out))
logger.info(
f"Some files matched the pattern '{pattern}' but don't have valid data file extensions: {invalid_matched_files}"
)
else:
out = matched_paths
if not out:
error_msg = f"Unable to find '{pattern}'"
if allowed_extensions is not None:
error_msg += f" with any supported extension {list(allowed_extensions)}"
raise FileNotFoundError(error_msg)
return out
def get_data_patterns(base_path: str, download_config: Optional[DownloadConfig] = None) -> Dict[str, List[str]]:
"""
Get the default pattern from a directory testing all the supported patterns.
The first patterns to return a non-empty list of data files is returned.
Some examples of supported patterns:
Input:
my_dataset_repository/
├── README.md
└── dataset.csv
Output:
{"train": ["**"]}
Input:
my_dataset_repository/
├── README.md
├── train.csv
└── test.csv
my_dataset_repository/
├── README.md
└── data/
├── train.csv
└── test.csv
my_dataset_repository/
├── README.md
├── train_0.csv
├── train_1.csv
├── train_2.csv
├── train_3.csv
├── test_0.csv
└── test_1.csv
Output:
{'train': ['train[-._ 0-9/]**', '**/*[-._ 0-9/]train[-._ 0-9/]**', 'training[-._ 0-9/]**', '**/*[-._ 0-9/]training[-._ 0-9/]**'],
'test': ['test[-._ 0-9/]**', '**/*[-._ 0-9/]test[-._ 0-9/]**', 'testing[-._ 0-9/]**', '**/*[-._ 0-9/]testing[-._ 0-9/]**', ...]}
Input:
my_dataset_repository/
├── README.md
└── data/
├── train/
│ ├── shard_0.csv
│ ├── shard_1.csv
│ ├── shard_2.csv
│ └── shard_3.csv
└── test/
├── shard_0.csv
└── shard_1.csv
Output:
{'train': ['train[-._ 0-9/]**', '**/*[-._ 0-9/]train[-._ 0-9/]**', 'training[-._ 0-9/]**', '**/*[-._ 0-9/]training[-._ 0-9/]**'],
'test': ['test[-._ 0-9/]**', '**/*[-._ 0-9/]test[-._ 0-9/]**', 'testing[-._ 0-9/]**', '**/*[-._ 0-9/]testing[-._ 0-9/]**', ...]}
Input:
my_dataset_repository/
├── README.md
└── data/
├── train-00000-of-00003.csv
├── train-00001-of-00003.csv
├── train-00002-of-00003.csv
├── test-00000-of-00001.csv
├── random-00000-of-00003.csv
├── random-00001-of-00003.csv
└── random-00002-of-00003.csv
Output:
{'train': ['data/train-[0-9][0-9][0-9][0-9][0-9]-of-[0-9][0-9][0-9][0-9][0-9]*.*'],
'test': ['data/test-[0-9][0-9][0-9][0-9][0-9]-of-[0-9][0-9][0-9][0-9][0-9]*.*'],
'random': ['data/random-[0-9][0-9][0-9][0-9][0-9]-of-[0-9][0-9][0-9][0-9][0-9]*.*']}
In order, it first tests if SPLIT_PATTERN_SHARDED works, otherwise it tests the patterns in ALL_DEFAULT_PATTERNS.
"""
resolver = partial(resolve_pattern, base_path=base_path, download_config=download_config)
try:
return _get_data_files_patterns(resolver)
except FileNotFoundError:
raise EmptyDatasetError(f"The directory at {base_path} doesn't contain any data files") from None
def get_metadata_patterns(
base_path: str,
download_config: Optional[DownloadConfig] = None,
) -> List[str]:
"""
Get the supported metadata patterns from a local directory.
"""
resolver = partial(resolve_pattern, base_path=base_path, download_config=download_config)
try:
return _get_metadata_files_patterns(resolver)
except FileNotFoundError:
raise FileNotFoundError(f"The directory at {base_path} doesn't contain any metadata file") from None
def _get_single_origin_metadata(
data_file: str,
download_config: Optional[DownloadConfig] = None,
) -> Tuple[str]:
data_file, storage_options = _prepare_path_and_storage_options(data_file, download_config=download_config)
fs, _, _ = get_fs_token_paths(data_file, storage_options=storage_options)
if isinstance(fs, HfFileSystem):
resolved_path = fs.resolve_path(data_file)
return (resolved_path.repo_id, resolved_path.revision)
elif isinstance(fs, HTTPFileSystem) and data_file.startswith(config.HF_ENDPOINT):
hffs = HfFileSystem(endpoint=config.HF_ENDPOINT, token=download_config.token)
data_file = "hf://" + data_file[len(config.HF_ENDPOINT) + 1 :].replace("/resolve/", "@", 1)
resolved_path = hffs.resolve_path(data_file)
return (resolved_path.repo_id, resolved_path.revision)
info = fs.info(data_file)
# s3fs uses "ETag", gcsfs uses "etag", and for local we simply check mtime
for key in ["ETag", "etag", "mtime"]:
if key in info:
return (str(info[key]),)
return ()
def _get_origin_metadata(
data_files: List[str],
max_workers=64,
download_config: Optional[DownloadConfig] = None,
) -> Tuple[str]:
return thread_map(
partial(_get_single_origin_metadata, download_config=download_config),
data_files,
max_workers=max_workers,
tqdm_class=hf_tqdm,
desc="Resolving data files",
disable=len(data_files) <= 16,
)
class DataFilesList(List[str]):
"""
List of data files (absolute local paths or URLs).
It has two construction methods given the user's data files patterns :
- ``from_hf_repo``: resolve patterns inside a dataset repository
- ``from_local_or_remote``: resolve patterns from a local path
Moreover DataFilesList has an additional attribute ``origin_metadata``.
It can store:
- the last modified time of local files
- ETag of remote files
- commit sha of a dataset repository
Thanks to this additional attribute, it is possible to hash the list
and get a different hash if and only if at least one file changed.
This is useful for caching Dataset objects that are obtained from a list of data files.
"""
def __init__(self, data_files: List[str], origin_metadata: List[Tuple[str]]):
super().__init__(data_files)
self.origin_metadata = origin_metadata
def __add__(self, other):
return DataFilesList([*self, *other], self.origin_metadata + other.origin_metadata)
@classmethod
def from_hf_repo(
cls,
patterns: List[str],
dataset_info: huggingface_hub.hf_api.DatasetInfo,
base_path: Optional[str] = None,
allowed_extensions: Optional[List[str]] = None,
download_config: Optional[DownloadConfig] = None,
) -> "DataFilesList":
base_path = f"hf://datasets/{dataset_info.id}@{dataset_info.sha}/{base_path or ''}".rstrip("/")
return cls.from_patterns(
patterns, base_path=base_path, allowed_extensions=allowed_extensions, download_config=download_config
)
@classmethod
def from_local_or_remote(
cls,
patterns: List[str],
base_path: Optional[str] = None,
allowed_extensions: Optional[List[str]] = None,
download_config: Optional[DownloadConfig] = None,
) -> "DataFilesList":
base_path = base_path if base_path is not None else Path().resolve().as_posix()
return cls.from_patterns(
patterns, base_path=base_path, allowed_extensions=allowed_extensions, download_config=download_config
)
@classmethod
def from_patterns(
cls,
patterns: List[str],
base_path: Optional[str] = None,
allowed_extensions: Optional[List[str]] = None,
download_config: Optional[DownloadConfig] = None,
) -> "DataFilesList":
base_path = base_path if base_path is not None else Path().resolve().as_posix()
data_files = []
for pattern in patterns:
try:
data_files.extend(
resolve_pattern(
pattern,
base_path=base_path,
allowed_extensions=allowed_extensions,
download_config=download_config,
)
)
except FileNotFoundError:
if not has_magic(pattern):
raise
origin_metadata = _get_origin_metadata(data_files, download_config=download_config)
return cls(data_files, origin_metadata)
def filter_extensions(self, extensions: List[str]) -> "DataFilesList":
pattern = "|".join("\\" + ext for ext in extensions)
pattern = re.compile(f".*({pattern})(\\..+)?$")
return DataFilesList(
[data_file for data_file in self if pattern.match(data_file)],
origin_metadata=self.origin_metadata,
)
class DataFilesDict(Dict[str, DataFilesList]):
"""
Dict of split_name -> list of data files (absolute local paths or URLs).
It has two construction methods given the user's data files patterns :
- ``from_hf_repo``: resolve patterns inside a dataset repository
- ``from_local_or_remote``: resolve patterns from a local path
Moreover each list is a DataFilesList. It is possible to hash the dictionary
and get a different hash if and only if at least one file changed.
For more info, see ``DataFilesList``.
This is useful for caching Dataset objects that are obtained from a list of data files.
Changing the order of the keys of this dictionary also doesn't change its hash.
"""
@classmethod
def from_local_or_remote(
cls,
patterns: Dict[str, Union[List[str], DataFilesList]],
base_path: Optional[str] = None,
allowed_extensions: Optional[List[str]] = None,
download_config: Optional[DownloadConfig] = None,
) -> "DataFilesDict":
out = cls()
for key, patterns_for_key in patterns.items():
out[key] = (
DataFilesList.from_local_or_remote(
patterns_for_key,
base_path=base_path,
allowed_extensions=allowed_extensions,
download_config=download_config,
)
if not isinstance(patterns_for_key, DataFilesList)
else patterns_for_key
)
return out
@classmethod
def from_hf_repo(
cls,
patterns: Dict[str, Union[List[str], DataFilesList]],
dataset_info: huggingface_hub.hf_api.DatasetInfo,
base_path: Optional[str] = None,
allowed_extensions: Optional[List[str]] = None,
download_config: Optional[DownloadConfig] = None,
) -> "DataFilesDict":
out = cls()
for key, patterns_for_key in patterns.items():
out[key] = (
DataFilesList.from_hf_repo(
patterns_for_key,
dataset_info=dataset_info,
base_path=base_path,
allowed_extensions=allowed_extensions,
download_config=download_config,
)
if not isinstance(patterns_for_key, DataFilesList)
else patterns_for_key
)
return out
@classmethod
def from_patterns(
cls,
patterns: Dict[str, Union[List[str], DataFilesList]],
base_path: Optional[str] = None,
allowed_extensions: Optional[List[str]] = None,
download_config: Optional[DownloadConfig] = None,
) -> "DataFilesDict":
out = cls()
for key, patterns_for_key in patterns.items():
out[key] = (
DataFilesList.from_patterns(
patterns_for_key,
base_path=base_path,
allowed_extensions=allowed_extensions,
download_config=download_config,
)
if not isinstance(patterns_for_key, DataFilesList)
else patterns_for_key
)
return out
def filter_extensions(self, extensions: List[str]) -> "DataFilesDict":
out = type(self)()
for key, data_files_list in self.items():
out[key] = data_files_list.filter_extensions(extensions)
return out
| 0
|
hf_public_repos/datasets/src
|
hf_public_repos/datasets/src/datasets/dataset_dict.py
|
import contextlib
import copy
import fnmatch
import json
import math
import posixpath
import re
import warnings
from io import BytesIO
from pathlib import Path
from typing import Callable, Dict, List, Optional, Sequence, Tuple, Union
import fsspec
import numpy as np
from huggingface_hub import (
CommitOperationAdd,
CommitOperationDelete,
DatasetCard,
DatasetCardData,
HfApi,
)
from . import config
from .arrow_dataset import PUSH_TO_HUB_WITHOUT_METADATA_CONFIGS_SPLIT_PATTERN_SHARDED, Dataset
from .features import Features
from .features.features import FeatureType
from .info import DatasetInfo, DatasetInfosDict
from .naming import _split_re
from .splits import NamedSplit, Split, SplitDict, SplitInfo
from .table import Table
from .tasks import TaskTemplate
from .utils import logging
from .utils.deprecation_utils import deprecated
from .utils.doc_utils import is_documented_by
from .utils.metadata import MetadataConfigs
from .utils.py_utils import asdict, glob_pattern_to_regex, string_to_dict
from .utils.typing import PathLike
logger = logging.get_logger(__name__)
class DatasetDict(dict):
"""A dictionary (dict of str: datasets.Dataset) with dataset transforms methods (map, filter, etc.)"""
def _check_values_type(self):
for dataset in self.values():
if not isinstance(dataset, Dataset):
raise TypeError(f"Values in `DatasetDict` should be of type `Dataset` but got type '{type(dataset)}'")
def _check_values_features(self):
items = list(self.items())
for item_a, item_b in zip(items[:-1], items[1:]):
if item_a[1].features != item_b[1].features:
raise ValueError(
f"All datasets in `DatasetDict` should have the same features but features for '{item_a[0]}' and '{item_b[0]}' don't match: {item_a[1].features} != {item_b[1].features}"
)
def __getitem__(self, k) -> Dataset:
if isinstance(k, (str, NamedSplit)) or len(self) == 0:
return super().__getitem__(k)
else:
available_suggested_splits = [
split for split in (Split.TRAIN, Split.TEST, Split.VALIDATION) if split in self
]
suggested_split = available_suggested_splits[0] if available_suggested_splits else list(self)[0]
raise KeyError(
f"Invalid key: {k}. Please first select a split. For example: "
f"`my_dataset_dictionary['{suggested_split}'][{k}]`. "
f"Available splits: {sorted(self)}"
)
@property
def data(self) -> Dict[str, Table]:
"""The Apache Arrow tables backing each split.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes")
>>> ds.data
```
"""
self._check_values_type()
return {k: dataset.data for k, dataset in self.items()}
@property
def cache_files(self) -> Dict[str, Dict]:
"""The cache files containing the Apache Arrow table backing each split.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes")
>>> ds.cache_files
{'test': [{'filename': '/root/.cache/huggingface/datasets/rotten_tomatoes_movie_review/default/1.0.0/40d411e45a6ce3484deed7cc15b82a53dad9a72aafd9f86f8f227134bec5ca46/rotten_tomatoes_movie_review-test.arrow'}],
'train': [{'filename': '/root/.cache/huggingface/datasets/rotten_tomatoes_movie_review/default/1.0.0/40d411e45a6ce3484deed7cc15b82a53dad9a72aafd9f86f8f227134bec5ca46/rotten_tomatoes_movie_review-train.arrow'}],
'validation': [{'filename': '/root/.cache/huggingface/datasets/rotten_tomatoes_movie_review/default/1.0.0/40d411e45a6ce3484deed7cc15b82a53dad9a72aafd9f86f8f227134bec5ca46/rotten_tomatoes_movie_review-validation.arrow'}]}
```
"""
self._check_values_type()
return {k: dataset.cache_files for k, dataset in self.items()}
@property
def num_columns(self) -> Dict[str, int]:
"""Number of columns in each split of the dataset.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes")
>>> ds.num_columns
{'test': 2, 'train': 2, 'validation': 2}
```
"""
self._check_values_type()
return {k: dataset.num_columns for k, dataset in self.items()}
@property
def num_rows(self) -> Dict[str, int]:
"""Number of rows in each split of the dataset (same as :func:`datasets.Dataset.__len__`).
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes")
>>> ds.num_rows
{'test': 1066, 'train': 8530, 'validation': 1066}
```
"""
self._check_values_type()
return {k: dataset.num_rows for k, dataset in self.items()}
@property
def column_names(self) -> Dict[str, List[str]]:
"""Names of the columns in each split of the dataset.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes")
>>> ds.column_names
{'test': ['text', 'label'],
'train': ['text', 'label'],
'validation': ['text', 'label']}
```
"""
self._check_values_type()
return {k: dataset.column_names for k, dataset in self.items()}
@property
def shape(self) -> Dict[str, Tuple[int]]:
"""Shape of each split of the dataset (number of columns, number of rows).
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes")
>>> ds.shape
{'test': (1066, 2), 'train': (8530, 2), 'validation': (1066, 2)}
```
"""
self._check_values_type()
return {k: dataset.shape for k, dataset in self.items()}
def flatten(self, max_depth=16) -> "DatasetDict":
"""Flatten the Apache Arrow Table of each split (nested features are flatten).
Each column with a struct type is flattened into one column per struct field.
Other columns are left unchanged.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("squad")
>>> ds["train"].features
{'answers': Sequence(feature={'text': Value(dtype='string', id=None), 'answer_start': Value(dtype='int32', id=None)}, length=-1, id=None),
'context': Value(dtype='string', id=None),
'id': Value(dtype='string', id=None),
'question': Value(dtype='string', id=None),
'title': Value(dtype='string', id=None)}
>>> ds.flatten()
DatasetDict({
train: Dataset({
features: ['id', 'title', 'context', 'question', 'answers.text', 'answers.answer_start'],
num_rows: 87599
})
validation: Dataset({
features: ['id', 'title', 'context', 'question', 'answers.text', 'answers.answer_start'],
num_rows: 10570
})
})
```
"""
self._check_values_type()
return DatasetDict({k: dataset.flatten(max_depth=max_depth) for k, dataset in self.items()})
def unique(self, column: str) -> Dict[str, List]:
"""Return a list of the unique elements in a column for each split.
This is implemented in the low-level backend and as such, very fast.
Args:
column (`str`):
column name (list all the column names with [`~datasets.Dataset.column_names`])
Returns:
Dict[`str`, `list`]: Dictionary of unique elements in the given column.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes")
>>> ds.unique("label")
{'test': [1, 0], 'train': [1, 0], 'validation': [1, 0]}
```
"""
self._check_values_type()
return {k: dataset.unique(column) for k, dataset in self.items()}
def cleanup_cache_files(self) -> Dict[str, int]:
"""Clean up all cache files in the dataset cache directory, excepted the currently used cache file if there is one.
Be careful when running this command that no other process is currently using other cache files.
Return:
`Dict` with the number of removed files for each split
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes")
>>> ds.cleanup_cache_files()
{'test': 0, 'train': 0, 'validation': 0}
```
"""
self._check_values_type()
return {k: dataset.cleanup_cache_files() for k, dataset in self.items()}
def __repr__(self):
repr = "\n".join([f"{k}: {v}" for k, v in self.items()])
repr = re.sub(r"^", " " * 4, repr, 0, re.M)
return f"DatasetDict({{\n{repr}\n}})"
def cast(self, features: Features) -> "DatasetDict":
"""
Cast the dataset to a new set of features.
The transformation is applied to all the datasets of the dataset dictionary.
You can also remove a column using [`Dataset.map`] with `feature` but `cast`
is in-place (doesn't copy the data to a new dataset) and is thus faster.
Args:
features ([`Features`]):
New features to cast the dataset to.
The name and order of the fields in the features must match the current column names.
The type of the data must also be convertible from one type to the other.
For non-trivial conversion, e.g. `string` <-> `ClassLabel` you should use [`~Dataset.map`] to update the Dataset.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes")
>>> ds["train"].features
{'label': ClassLabel(num_classes=2, names=['neg', 'pos'], id=None),
'text': Value(dtype='string', id=None)}
>>> new_features = ds["train"].features.copy()
>>> new_features['label'] = ClassLabel(names=['bad', 'good'])
>>> new_features['text'] = Value('large_string')
>>> ds = ds.cast(new_features)
>>> ds["train"].features
{'label': ClassLabel(num_classes=2, names=['bad', 'good'], id=None),
'text': Value(dtype='large_string', id=None)}
```
"""
self._check_values_type()
return DatasetDict({k: dataset.cast(features=features) for k, dataset in self.items()})
def cast_column(self, column: str, feature) -> "DatasetDict":
"""Cast column to feature for decoding.
Args:
column (`str`):
Column name.
feature ([`Feature`]):
Target feature.
Returns:
[`DatasetDict`]
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes")
>>> ds["train"].features
{'label': ClassLabel(num_classes=2, names=['neg', 'pos'], id=None),
'text': Value(dtype='string', id=None)}
>>> ds = ds.cast_column('label', ClassLabel(names=['bad', 'good']))
>>> ds["train"].features
{'label': ClassLabel(num_classes=2, names=['bad', 'good'], id=None),
'text': Value(dtype='string', id=None)}
```
"""
self._check_values_type()
return DatasetDict({k: dataset.cast_column(column=column, feature=feature) for k, dataset in self.items()})
def remove_columns(self, column_names: Union[str, List[str]]) -> "DatasetDict":
"""
Remove one or several column(s) from each split in the dataset
and the features associated to the column(s).
The transformation is applied to all the splits of the dataset dictionary.
You can also remove a column using [`Dataset.map`] with `remove_columns` but the present method
is in-place (doesn't copy the data to a new dataset) and is thus faster.
Args:
column_names (`Union[str, List[str]]`):
Name of the column(s) to remove.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes")
>>> ds.remove_columns("label")
DatasetDict({
train: Dataset({
features: ['text'],
num_rows: 8530
})
validation: Dataset({
features: ['text'],
num_rows: 1066
})
test: Dataset({
features: ['text'],
num_rows: 1066
})
})
```
"""
self._check_values_type()
return DatasetDict({k: dataset.remove_columns(column_names=column_names) for k, dataset in self.items()})
def rename_column(self, original_column_name: str, new_column_name: str) -> "DatasetDict":
"""
Rename a column in the dataset and move the features associated to the original column under the new column name.
The transformation is applied to all the datasets of the dataset dictionary.
You can also rename a column using [`~Dataset.map`] with `remove_columns` but the present method:
- takes care of moving the original features under the new column name.
- doesn't copy the data to a new dataset and is thus much faster.
Args:
original_column_name (`str`):
Name of the column to rename.
new_column_name (`str`):
New name for the column.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes")
>>> ds.rename_column("label", "label_new")
DatasetDict({
train: Dataset({
features: ['text', 'label_new'],
num_rows: 8530
})
validation: Dataset({
features: ['text', 'label_new'],
num_rows: 1066
})
test: Dataset({
features: ['text', 'label_new'],
num_rows: 1066
})
})
```
"""
self._check_values_type()
return DatasetDict(
{
k: dataset.rename_column(original_column_name=original_column_name, new_column_name=new_column_name)
for k, dataset in self.items()
}
)
def rename_columns(self, column_mapping: Dict[str, str]) -> "DatasetDict":
"""
Rename several columns in the dataset, and move the features associated to the original columns under
the new column names.
The transformation is applied to all the datasets of the dataset dictionary.
Args:
column_mapping (`Dict[str, str]`):
A mapping of columns to rename to their new names.
Returns:
[`DatasetDict`]: A copy of the dataset with renamed columns.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes")
>>> ds.rename_columns({'text': 'text_new', 'label': 'label_new'})
DatasetDict({
train: Dataset({
features: ['text_new', 'label_new'],
num_rows: 8530
})
validation: Dataset({
features: ['text_new', 'label_new'],
num_rows: 1066
})
test: Dataset({
features: ['text_new', 'label_new'],
num_rows: 1066
})
})
```
"""
self._check_values_type()
return DatasetDict({k: dataset.rename_columns(column_mapping=column_mapping) for k, dataset in self.items()})
def select_columns(self, column_names: Union[str, List[str]]) -> "DatasetDict":
"""Select one or several column(s) from each split in the dataset and
the features associated to the column(s).
The transformation is applied to all the splits of the dataset
dictionary.
Args:
column_names (`Union[str, List[str]]`):
Name of the column(s) to keep.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes")
>>> ds.select_columns("text")
DatasetDict({
train: Dataset({
features: ['text'],
num_rows: 8530
})
validation: Dataset({
features: ['text'],
num_rows: 1066
})
test: Dataset({
features: ['text'],
num_rows: 1066
})
})
```
"""
self._check_values_type()
return DatasetDict({k: dataset.select_columns(column_names=column_names) for k, dataset in self.items()})
def class_encode_column(self, column: str, include_nulls: bool = False) -> "DatasetDict":
"""Casts the given column as [`~datasets.features.ClassLabel`] and updates the tables.
Args:
column (`str`):
The name of the column to cast.
include_nulls (`bool`, defaults to `False`):
Whether to include null values in the class labels. If `True`, the null values will be encoded as the `"None"` class label.
<Added version="1.14.2"/>
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("boolq")
>>> ds["train"].features
{'answer': Value(dtype='bool', id=None),
'passage': Value(dtype='string', id=None),
'question': Value(dtype='string', id=None)}
>>> ds = ds.class_encode_column("answer")
>>> ds["train"].features
{'answer': ClassLabel(num_classes=2, names=['False', 'True'], id=None),
'passage': Value(dtype='string', id=None),
'question': Value(dtype='string', id=None)}
```
"""
self._check_values_type()
return DatasetDict(
{k: dataset.class_encode_column(column=column, include_nulls=include_nulls) for k, dataset in self.items()}
)
@contextlib.contextmanager
def formatted_as(
self,
type: Optional[str] = None,
columns: Optional[List] = None,
output_all_columns: bool = False,
**format_kwargs,
):
"""To be used in a `with` statement. Set `__getitem__` return format (type and columns).
The transformation is applied to all the datasets of the dataset dictionary.
Args:
type (`str`, *optional*):
Output type selected in `[None, 'numpy', 'torch', 'tensorflow', 'pandas', 'arrow', 'jax']`.
`None` means `__getitem__` returns python objects (default).
columns (`List[str]`, *optional*):
Columns to format in the output.
`None` means `__getitem__` returns all columns (default).
output_all_columns (`bool`, defaults to False):
Keep un-formatted columns as well in the output (as python objects).
**format_kwargs (additional keyword arguments):
Keywords arguments passed to the convert function like `np.array`, `torch.tensor` or `tensorflow.ragged.constant`.
"""
self._check_values_type()
old_format_type = {k: dataset._format_type for k, dataset in self.items()}
old_format_kwargs = {k: dataset._format_kwargs for k, dataset in self.items()}
old_format_columns = {k: dataset._format_columns for k, dataset in self.items()}
old_output_all_columns = {k: dataset._output_all_columns for k, dataset in self.items()}
try:
self.set_format(type, columns, output_all_columns, **format_kwargs)
yield
finally:
for k, dataset in self.items():
dataset.set_format(
old_format_type[k], old_format_columns[k], old_output_all_columns[k], **old_format_kwargs[k]
)
def set_format(
self,
type: Optional[str] = None,
columns: Optional[List] = None,
output_all_columns: bool = False,
**format_kwargs,
):
"""Set `__getitem__` return format (type and columns).
The format is set for every dataset in the dataset dictionary.
Args:
type (`str`, *optional*):
Output type selected in `[None, 'numpy', 'torch', 'tensorflow', 'pandas', 'arrow', 'jax']`.
`None` means `__getitem__` returns python objects (default).
columns (`List[str]`, *optional*):
Columns to format in the output.
`None` means `__getitem__` returns all columns (default).
output_all_columns (`bool`, defaults to False):
Keep un-formatted columns as well in the output (as python objects),
**format_kwargs (additional keyword arguments):
Keywords arguments passed to the convert function like `np.array`, `torch.tensor` or `tensorflow.ragged.constant`.
It is possible to call `map` after calling `set_format`. Since `map` may add new columns, then the list of formatted columns
gets updated. In this case, if you apply `map` on a dataset to add a new column, then this column will be formatted:
`new formatted columns = (all columns - previously unformatted columns)`
Example:
```py
>>> from datasets import load_dataset
>>> from transformers import AutoTokenizer
>>> tokenizer = AutoTokenizer.from_pretrained("bert-base-cased")
>>> ds = ds.map(lambda x: tokenizer(x["text"], truncation=True, padding=True), batched=True)
>>> ds.set_format(type="numpy", columns=['input_ids', 'token_type_ids', 'attention_mask', 'label'])
>>> ds["train"].format
{'columns': ['input_ids', 'token_type_ids', 'attention_mask', 'label'],
'format_kwargs': {},
'output_all_columns': False,
'type': 'numpy'}
```
"""
self._check_values_type()
for dataset in self.values():
dataset.set_format(type=type, columns=columns, output_all_columns=output_all_columns, **format_kwargs)
def reset_format(self):
"""Reset `__getitem__` return format to python objects and all columns.
The transformation is applied to all the datasets of the dataset dictionary.
Same as `self.set_format()`
Example:
```py
>>> from datasets import load_dataset
>>> from transformers import AutoTokenizer
>>> ds = load_dataset("rotten_tomatoes")
>>> tokenizer = AutoTokenizer.from_pretrained("bert-base-cased")
>>> ds = ds.map(lambda x: tokenizer(x["text"], truncation=True, padding=True), batched=True)
>>> ds.set_format(type="numpy", columns=['input_ids', 'token_type_ids', 'attention_mask', 'label'])
>>> ds["train"].format
{'columns': ['input_ids', 'token_type_ids', 'attention_mask', 'label'],
'format_kwargs': {},
'output_all_columns': False,
'type': 'numpy'}
>>> ds.reset_format()
>>> ds["train"].format
{'columns': ['text', 'label', 'input_ids', 'token_type_ids', 'attention_mask'],
'format_kwargs': {},
'output_all_columns': False,
'type': None}
```
"""
self._check_values_type()
for dataset in self.values():
dataset.set_format()
def set_transform(
self,
transform: Optional[Callable],
columns: Optional[List] = None,
output_all_columns: bool = False,
):
"""Set ``__getitem__`` return format using this transform. The transform is applied on-the-fly on batches when ``__getitem__`` is called.
The transform is set for every dataset in the dataset dictionary
As :func:`datasets.Dataset.set_format`, this can be reset using :func:`datasets.Dataset.reset_format`
Args:
transform (`Callable`, optional): user-defined formatting transform, replaces the format defined by :func:`datasets.Dataset.set_format`
A formatting function is a callable that takes a batch (as a dict) as input and returns a batch.
This function is applied right before returning the objects in ``__getitem__``.
columns (`List[str]`, optional): columns to format in the output
If specified, then the input batch of the transform only contains those columns.
output_all_columns (`bool`, default to False): keep un-formatted columns as well in the output (as python objects)
If set to True, then the other un-formatted columns are kept with the output of the transform.
"""
self._check_values_type()
for dataset in self.values():
dataset.set_format("custom", columns=columns, output_all_columns=output_all_columns, transform=transform)
def with_format(
self,
type: Optional[str] = None,
columns: Optional[List] = None,
output_all_columns: bool = False,
**format_kwargs,
) -> "DatasetDict":
"""Set `__getitem__` return format (type and columns). The data formatting is applied on-the-fly.
The format `type` (for example "numpy") is used to format batches when using `__getitem__`.
The format is set for every dataset in the dataset dictionary.
It's also possible to use custom transforms for formatting using [`~datasets.Dataset.with_transform`].
Contrary to [`~datasets.DatasetDict.set_format`], `with_format` returns a new [`DatasetDict`] object with new [`Dataset`] objects.
Args:
type (`str`, *optional*):
Output type selected in `[None, 'numpy', 'torch', 'tensorflow', 'pandas', 'arrow', 'jax']`.
`None` means `__getitem__` returns python objects (default).
columns (`List[str]`, *optional*):
Columns to format in the output.
`None` means `__getitem__` returns all columns (default).
output_all_columns (`bool`, defaults to `False`):
Keep un-formatted columns as well in the output (as python objects).
**format_kwargs (additional keyword arguments):
Keywords arguments passed to the convert function like `np.array`, `torch.tensor` or `tensorflow.ragged.constant`.
Example:
```py
>>> from datasets import load_dataset
>>> from transformers import AutoTokenizer
>>> ds = load_dataset("rotten_tomatoes")
>>> tokenizer = AutoTokenizer.from_pretrained("bert-base-cased")
>>> ds = ds.map(lambda x: tokenizer(x['text'], truncation=True, padding=True), batched=True)
>>> ds["train"].format
{'columns': ['text', 'label', 'input_ids', 'token_type_ids', 'attention_mask'],
'format_kwargs': {},
'output_all_columns': False,
'type': None}
>>> ds = ds.with_format(type='tensorflow', columns=['input_ids', 'token_type_ids', 'attention_mask', 'label'])
>>> ds["train"].format
{'columns': ['input_ids', 'token_type_ids', 'attention_mask', 'label'],
'format_kwargs': {},
'output_all_columns': False,
'type': 'tensorflow'}
```
"""
dataset = copy.deepcopy(self)
dataset.set_format(type=type, columns=columns, output_all_columns=output_all_columns, **format_kwargs)
return dataset
def with_transform(
self,
transform: Optional[Callable],
columns: Optional[List] = None,
output_all_columns: bool = False,
) -> "DatasetDict":
"""Set `__getitem__` return format using this transform. The transform is applied on-the-fly on batches when `__getitem__` is called.
The transform is set for every dataset in the dataset dictionary
As [`~datasets.Dataset.set_format`], this can be reset using [`~datasets.Dataset.reset_format`].
Contrary to [`~datasets.DatasetDict.set_transform`], `with_transform` returns a new [`DatasetDict`] object with new [`Dataset`] objects.
Args:
transform (`Callable`, *optional*):
User-defined formatting transform, replaces the format defined by [`~datasets.Dataset.set_format`].
A formatting function is a callable that takes a batch (as a dict) as input and returns a batch.
This function is applied right before returning the objects in `__getitem__`.
columns (`List[str]`, *optional*):
Columns to format in the output.
If specified, then the input batch of the transform only contains those columns.
output_all_columns (`bool`, defaults to False):
Keep un-formatted columns as well in the output (as python objects).
If set to `True`, then the other un-formatted columns are kept with the output of the transform.
Example:
```py
>>> from datasets import load_dataset
>>> from transformers import AutoTokenizer
>>> ds = load_dataset("rotten_tomatoes")
>>> tokenizer = AutoTokenizer.from_pretrained("bert-base-cased")
>>> def encode(example):
... return tokenizer(example['text'], truncation=True, padding=True, return_tensors="pt")
>>> ds = ds.with_transform(encode)
>>> ds["train"][0]
{'attention_mask': tensor([1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1,
1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1,
1, 1, 1, 1, 1, 1, 1, 1, 1]),
'input_ids': tensor([ 101, 1103, 2067, 1110, 17348, 1106, 1129, 1103, 6880, 1432,
112, 188, 1207, 107, 14255, 1389, 107, 1105, 1115, 1119,
112, 188, 1280, 1106, 1294, 170, 24194, 1256, 3407, 1190,
170, 11791, 5253, 188, 1732, 7200, 10947, 12606, 2895, 117,
179, 7766, 118, 172, 15554, 1181, 3498, 6961, 3263, 1137,
188, 1566, 7912, 14516, 6997, 119, 102]),
'token_type_ids': tensor([0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0])}
```
"""
dataset = copy.deepcopy(self)
dataset.set_transform(transform=transform, columns=columns, output_all_columns=output_all_columns)
return dataset
def map(
self,
function: Optional[Callable] = None,
with_indices: bool = False,
with_rank: bool = False,
input_columns: Optional[Union[str, List[str]]] = None,
batched: bool = False,
batch_size: Optional[int] = 1000,
drop_last_batch: bool = False,
remove_columns: Optional[Union[str, List[str]]] = None,
keep_in_memory: bool = False,
load_from_cache_file: Optional[bool] = None,
cache_file_names: Optional[Dict[str, Optional[str]]] = None,
writer_batch_size: Optional[int] = 1000,
features: Optional[Features] = None,
disable_nullable: bool = False,
fn_kwargs: Optional[dict] = None,
num_proc: Optional[int] = None,
desc: Optional[str] = None,
) -> "DatasetDict":
"""Apply a function to all the elements in the table (individually or in batches)
and update the table (if function does updated examples).
The transformation is applied to all the datasets of the dataset dictionary.
Args:
function (`callable`): with one of the following signature:
- `function(example: Dict[str, Any]) -> Dict[str, Any]` if `batched=False` and `with_indices=False`
- `function(example: Dict[str, Any], indices: int) -> Dict[str, Any]` if `batched=False` and `with_indices=True`
- `function(batch: Dict[str, List]) -> Dict[str, List]` if `batched=True` and `with_indices=False`
- `function(batch: Dict[str, List], indices: List[int]) -> Dict[str, List]` if `batched=True` and `with_indices=True`
For advanced usage, the function can also return a `pyarrow.Table`.
Moreover if your function returns nothing (`None`), then `map` will run your function and return the dataset unchanged.
with_indices (`bool`, defaults to `False`):
Provide example indices to `function`. Note that in this case the signature of `function` should be `def function(example, idx): ...`.
with_rank (`bool`, defaults to `False`):
Provide process rank to `function`. Note that in this case the
signature of `function` should be `def function(example[, idx], rank): ...`.
input_columns (`[Union[str, List[str]]]`, *optional*, defaults to `None`):
The columns to be passed into `function` as
positional arguments. If `None`, a dict mapping to all formatted columns is passed as one argument.
batched (`bool`, defaults to `False`):
Provide batch of examples to `function`.
batch_size (`int`, *optional*, defaults to `1000`):
Number of examples per batch provided to `function` if `batched=True`,
`batch_size <= 0` or `batch_size == None` then provide the full dataset as a single batch to `function`.
drop_last_batch (`bool`, defaults to `False`):
Whether a last batch smaller than the batch_size should be
dropped instead of being processed by the function.
remove_columns (`[Union[str, List[str]]]`, *optional*, defaults to `None`):
Remove a selection of columns while doing the mapping.
Columns will be removed before updating the examples with the output of `function`, i.e. if `function` is adding
columns with names in `remove_columns`, these columns will be kept.
keep_in_memory (`bool`, defaults to `False`):
Keep the dataset in memory instead of writing it to a cache file.
load_from_cache_file (`Optional[bool]`, defaults to `True` if caching is enabled):
If a cache file storing the current computation from `function`
can be identified, use it instead of recomputing.
cache_file_names (`[Dict[str, str]]`, *optional*, defaults to `None`):
Provide the name of a path for the cache file. It is used to store the
results of the computation instead of the automatically generated cache file name.
You have to provide one `cache_file_name` per dataset in the dataset dictionary.
writer_batch_size (`int`, default `1000`):
Number of rows per write operation for the cache file writer.
This value is a good trade-off between memory usage during the processing, and processing speed.
Higher value makes the processing do fewer lookups, lower value consume less temporary memory while running `map`.
features (`[datasets.Features]`, *optional*, defaults to `None`):
Use a specific [`Features`] to store the cache file
instead of the automatically generated one.
disable_nullable (`bool`, defaults to `False`):
Disallow null values in the table.
fn_kwargs (`Dict`, *optional*, defaults to `None`):
Keyword arguments to be passed to `function`
num_proc (`int`, *optional*, defaults to `None`):
Number of processes for multiprocessing. By default it doesn't
use multiprocessing.
desc (`str`, *optional*, defaults to `None`):
Meaningful description to be displayed alongside with the progress bar while mapping examples.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes")
>>> def add_prefix(example):
... example["text"] = "Review: " + example["text"]
... return example
>>> ds = ds.map(add_prefix)
>>> ds["train"][0:3]["text"]
['Review: the rock is destined to be the 21st century\'s new " conan " and that he\'s going to make a splash even greater than arnold schwarzenegger , jean-claud van damme or steven segal .',
'Review: the gorgeously elaborate continuation of " the lord of the rings " trilogy is so huge that a column of words cannot adequately describe co-writer/director peter jackson\'s expanded vision of j . r . r . tolkien\'s middle-earth .',
'Review: effective but too-tepid biopic']
# process a batch of examples
>>> ds = ds.map(lambda example: tokenizer(example["text"]), batched=True)
# set number of processors
>>> ds = ds.map(add_prefix, num_proc=4)
```
"""
self._check_values_type()
if cache_file_names is None:
cache_file_names = {k: None for k in self}
return DatasetDict(
{
k: dataset.map(
function=function,
with_indices=with_indices,
with_rank=with_rank,
input_columns=input_columns,
batched=batched,
batch_size=batch_size,
drop_last_batch=drop_last_batch,
remove_columns=remove_columns,
keep_in_memory=keep_in_memory,
load_from_cache_file=load_from_cache_file,
cache_file_name=cache_file_names[k],
writer_batch_size=writer_batch_size,
features=features,
disable_nullable=disable_nullable,
fn_kwargs=fn_kwargs,
num_proc=num_proc,
desc=desc,
)
for k, dataset in self.items()
}
)
def filter(
self,
function,
with_indices=False,
input_columns: Optional[Union[str, List[str]]] = None,
batched: bool = False,
batch_size: Optional[int] = 1000,
keep_in_memory: bool = False,
load_from_cache_file: Optional[bool] = None,
cache_file_names: Optional[Dict[str, Optional[str]]] = None,
writer_batch_size: Optional[int] = 1000,
fn_kwargs: Optional[dict] = None,
num_proc: Optional[int] = None,
desc: Optional[str] = None,
) -> "DatasetDict":
"""Apply a filter function to all the elements in the table in batches
and update the table so that the dataset only includes examples according to the filter function.
The transformation is applied to all the datasets of the dataset dictionary.
Args:
function (`callable`):
With one of the following signature:
- `function(example: Dict[str, Any]) -> bool` if `with_indices=False, batched=False`
- `function(example: Dict[str, Any], indices: int) -> bool` if `with_indices=True, batched=False`
- `function(example: Dict[str, List]) -> List[bool]` if `with_indices=False, batched=True`
- `function(example: Dict[str, List], indices: List[int]) -> List[bool]` if ``with_indices=True, batched=True`
with_indices (`bool`, defaults to `False`):
Provide example indices to `function`. Note that in this case the signature of `function` should be `def function(example, idx): ...`.
input_columns (`[Union[str, List[str]]]`, *optional*, defaults to `None`):
The columns to be passed into `function` as
positional arguments. If `None`, a dict mapping to all formatted columns is passed as one argument.
batched (`bool`, defaults to `False`):
Provide batch of examples to `function`.
batch_size (`int`, *optional*, defaults to `1000`):
Number of examples per batch provided to `function` if `batched=True`
`batch_size <= 0` or `batch_size == None` then provide the full dataset as a single batch to `function`.
keep_in_memory (`bool`, defaults to `False`):
Keep the dataset in memory instead of writing it to a cache file.
load_from_cache_file (`Optional[bool]`, defaults to `True` if chaching is enabled):
If a cache file storing the current computation from `function`
can be identified, use it instead of recomputing.
cache_file_names (`[Dict[str, str]]`, *optional*, defaults to `None`):
Provide the name of a path for the cache file. It is used to store the
results of the computation instead of the automatically generated cache file name.
You have to provide one `cache_file_name` per dataset in the dataset dictionary.
writer_batch_size (`int`, defaults to `1000`):
Number of rows per write operation for the cache file writer.
This value is a good trade-off between memory usage during the processing, and processing speed.
Higher value makes the processing do fewer lookups, lower value consume less temporary memory while running `map`.
fn_kwargs (`Dict`, *optional*, defaults to `None`):
Keyword arguments to be passed to `function`
num_proc (`int`, *optional*, defaults to `None`):
Number of processes for multiprocessing. By default it doesn't
use multiprocessing.
desc (`str`, *optional*, defaults to `None`):
Meaningful description to be displayed alongside with the progress bar while filtering examples.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes")
>>> ds.filter(lambda x: x["label"] == 1)
DatasetDict({
train: Dataset({
features: ['text', 'label'],
num_rows: 4265
})
validation: Dataset({
features: ['text', 'label'],
num_rows: 533
})
test: Dataset({
features: ['text', 'label'],
num_rows: 533
})
})
```
"""
self._check_values_type()
if cache_file_names is None:
cache_file_names = {k: None for k in self}
return DatasetDict(
{
k: dataset.filter(
function=function,
with_indices=with_indices,
input_columns=input_columns,
batched=batched,
batch_size=batch_size,
keep_in_memory=keep_in_memory,
load_from_cache_file=load_from_cache_file,
cache_file_name=cache_file_names[k],
writer_batch_size=writer_batch_size,
fn_kwargs=fn_kwargs,
num_proc=num_proc,
desc=desc,
)
for k, dataset in self.items()
}
)
def flatten_indices(
self,
keep_in_memory: bool = False,
cache_file_names: Optional[Dict[str, Optional[str]]] = None,
writer_batch_size: Optional[int] = 1000,
features: Optional[Features] = None,
disable_nullable: bool = False,
num_proc: Optional[int] = None,
new_fingerprint: Optional[str] = None,
) -> "DatasetDict":
"""Create and cache a new Dataset by flattening the indices mapping.
Args:
keep_in_memory (`bool`, defaults to `False`):
Keep the dataset in memory instead of writing it to a cache file.
cache_file_names (`Dict[str, str]`, *optional*, default `None`):
Provide the name of a path for the cache file. It is used to store the
results of the computation instead of the automatically generated cache file name.
You have to provide one `cache_file_name` per dataset in the dataset dictionary.
writer_batch_size (`int`, defaults to `1000`):
Number of rows per write operation for the cache file writer.
This value is a good trade-off between memory usage during the processing, and processing speed.
Higher value makes the processing do fewer lookups, lower value consume less temporary memory while running `map`.
features (`Optional[datasets.Features]`, defaults to `None`):
Use a specific [`Features`] to store the cache file
instead of the automatically generated one.
disable_nullable (`bool`, defaults to `False`):
Allow null values in the table.
num_proc (`int`, optional, default `None`):
Max number of processes when generating cache. Already cached shards are loaded sequentially
new_fingerprint (`str`, *optional*, defaults to `None`):
The new fingerprint of the dataset after transform.
If `None`, the new fingerprint is computed using a hash of the previous fingerprint, and the transform arguments
"""
self._check_values_type()
if cache_file_names is None:
cache_file_names = {k: None for k in self}
return DatasetDict(
{
k: dataset.flatten_indices(
keep_in_memory=keep_in_memory,
cache_file_name=cache_file_names[k],
writer_batch_size=writer_batch_size,
features=features,
disable_nullable=disable_nullable,
num_proc=num_proc,
new_fingerprint=new_fingerprint,
)
for k, dataset in self.items()
}
)
def sort(
self,
column_names: Union[str, Sequence[str]],
reverse: Union[bool, Sequence[bool]] = False,
kind="deprecated",
null_placement: str = "at_end",
keep_in_memory: bool = False,
load_from_cache_file: Optional[bool] = None,
indices_cache_file_names: Optional[Dict[str, Optional[str]]] = None,
writer_batch_size: Optional[int] = 1000,
) -> "DatasetDict":
"""Create a new dataset sorted according to a single or multiple columns.
Args:
column_names (`Union[str, Sequence[str]]`):
Column name(s) to sort by.
reverse (`Union[bool, Sequence[bool]]`, defaults to `False`):
If `True`, sort by descending order rather than ascending. If a single bool is provided,
the value is applied to the sorting of all column names. Otherwise a list of bools with the
same length and order as column_names must be provided.
kind (`str`, *optional*):
Pandas algorithm for sorting selected in `{quicksort, mergesort, heapsort, stable}`,
The default is `quicksort`. Note that both `stable` and `mergesort` use timsort under the covers and, in general,
the actual implementation will vary with data type. The `mergesort` option is retained for backwards compatibility.
<Deprecated version="2.8.0">
`kind` was deprecated in version 2.10.0 and will be removed in 3.0.0.
</Deprecated>
null_placement (`str`, defaults to `at_end`):
Put `None` values at the beginning if `at_start` or `first` or at the end if `at_end` or `last`
keep_in_memory (`bool`, defaults to `False`):
Keep the sorted indices in memory instead of writing it to a cache file.
load_from_cache_file (`Optional[bool]`, defaults to `True` if caching is enabled):
If a cache file storing the sorted indices
can be identified, use it instead of recomputing.
indices_cache_file_names (`[Dict[str, str]]`, *optional*, defaults to `None`):
Provide the name of a path for the cache file. It is used to store the
indices mapping instead of the automatically generated cache file name.
You have to provide one `cache_file_name` per dataset in the dataset dictionary.
writer_batch_size (`int`, defaults to `1000`):
Number of rows per write operation for the cache file writer.
Higher value gives smaller cache files, lower value consume less temporary memory.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset('rotten_tomatoes')
>>> ds['train']['label'][:10]
[1, 1, 1, 1, 1, 1, 1, 1, 1, 1]
>>> sorted_ds = ds.sort('label')
>>> sorted_ds['train']['label'][:10]
[0, 0, 0, 0, 0, 0, 0, 0, 0, 0]
>>> another_sorted_ds = ds.sort(['label', 'text'], reverse=[True, False])
>>> another_sorted_ds['train']['label'][:10]
[1, 1, 1, 1, 1, 1, 1, 1, 1, 1]
```
"""
self._check_values_type()
if indices_cache_file_names is None:
indices_cache_file_names = {k: None for k in self}
return DatasetDict(
{
k: dataset.sort(
column_names=column_names,
reverse=reverse,
kind=kind,
null_placement=null_placement,
keep_in_memory=keep_in_memory,
load_from_cache_file=load_from_cache_file,
indices_cache_file_name=indices_cache_file_names[k],
writer_batch_size=writer_batch_size,
)
for k, dataset in self.items()
}
)
def shuffle(
self,
seeds: Optional[Union[int, Dict[str, Optional[int]]]] = None,
seed: Optional[int] = None,
generators: Optional[Dict[str, np.random.Generator]] = None,
keep_in_memory: bool = False,
load_from_cache_file: Optional[bool] = None,
indices_cache_file_names: Optional[Dict[str, Optional[str]]] = None,
writer_batch_size: Optional[int] = 1000,
) -> "DatasetDict":
"""Create a new Dataset where the rows are shuffled.
The transformation is applied to all the datasets of the dataset dictionary.
Currently shuffling uses numpy random generators.
You can either supply a NumPy BitGenerator to use, or a seed to initiate NumPy's default random generator (PCG64).
Args:
seeds (`Dict[str, int]` or `int`, *optional*):
A seed to initialize the default BitGenerator if `generator=None`.
If `None`, then fresh, unpredictable entropy will be pulled from the OS.
If an `int` or `array_like[ints]` is passed, then it will be passed to SeedSequence to derive the initial BitGenerator state.
You can provide one `seed` per dataset in the dataset dictionary.
seed (`int`, *optional*):
A seed to initialize the default BitGenerator if `generator=None`. Alias for seeds (a `ValueError` is raised if both are provided).
generators (`Dict[str, *optional*, np.random.Generator]`):
Numpy random Generator to use to compute the permutation of the dataset rows.
If `generator=None` (default), uses `np.random.default_rng` (the default BitGenerator (PCG64) of NumPy).
You have to provide one `generator` per dataset in the dataset dictionary.
keep_in_memory (`bool`, defaults to `False`):
Keep the dataset in memory instead of writing it to a cache file.
load_from_cache_file (`Optional[bool]`, defaults to `True` if caching is enabled):
If a cache file storing the current computation from `function`
can be identified, use it instead of recomputing.
indices_cache_file_names (`Dict[str, str]`, *optional*):
Provide the name of a path for the cache file. It is used to store the
indices mappings instead of the automatically generated cache file name.
You have to provide one `cache_file_name` per dataset in the dataset dictionary.
writer_batch_size (`int`, defaults to `1000`):
Number of rows per write operation for the cache file writer.
This value is a good trade-off between memory usage during the processing, and processing speed.
Higher value makes the processing do fewer lookups, lower value consume less temporary memory while running `map`.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes")
>>> ds["train"]["label"][:10]
[1, 1, 1, 1, 1, 1, 1, 1, 1, 1]
# set a seed
>>> shuffled_ds = ds.shuffle(seed=42)
>>> shuffled_ds["train"]["label"][:10]
[0, 1, 0, 1, 0, 0, 0, 0, 0, 0]
```
"""
self._check_values_type()
if seed is not None and seeds is not None:
raise ValueError("Please specify seed or seeds, but not both")
seeds = seed if seed is not None else seeds
if seeds is None:
seeds = {k: None for k in self}
elif not isinstance(seeds, dict):
seeds = {k: seeds for k in self}
if generators is None:
generators = {k: None for k in self}
if indices_cache_file_names is None:
indices_cache_file_names = {k: None for k in self}
return DatasetDict(
{
k: dataset.shuffle(
seed=seeds[k],
generator=generators[k],
keep_in_memory=keep_in_memory,
load_from_cache_file=load_from_cache_file,
indices_cache_file_name=indices_cache_file_names[k],
writer_batch_size=writer_batch_size,
)
for k, dataset in self.items()
}
)
def save_to_disk(
self,
dataset_dict_path: PathLike,
fs="deprecated",
max_shard_size: Optional[Union[str, int]] = None,
num_shards: Optional[Dict[str, int]] = None,
num_proc: Optional[int] = None,
storage_options: Optional[dict] = None,
):
"""
Saves a dataset dict to a filesystem using `fsspec.spec.AbstractFileSystem`.
For [`Image`] and [`Audio`] data:
All the Image() and Audio() data are stored in the arrow files.
If you want to store paths or urls, please use the Value("string") type.
Args:
dataset_dict_path (`str`):
Path (e.g. `dataset/train`) or remote URI
(e.g. `s3://my-bucket/dataset/train`) of the dataset dict directory where the dataset dict will be
saved to.
fs (`fsspec.spec.AbstractFileSystem`, *optional*):
Instance of the remote filesystem where the dataset will be saved to.
<Deprecated version="2.8.0">
`fs` was deprecated in version 2.8.0 and will be removed in 3.0.0.
Please use `storage_options` instead, e.g. `storage_options=fs.storage_options`
</Deprecated>
max_shard_size (`int` or `str`, *optional*, defaults to `"500MB"`):
The maximum size of the dataset shards to be uploaded to the hub. If expressed as a string, needs to be digits followed by a unit
(like `"50MB"`).
num_shards (`Dict[str, int]`, *optional*):
Number of shards to write. By default the number of shards depends on `max_shard_size` and `num_proc`.
You need to provide the number of shards for each dataset in the dataset dictionary.
Use a dictionary to define a different num_shards for each split.
<Added version="2.8.0"/>
num_proc (`int`, *optional*, default `None`):
Number of processes when downloading and generating the dataset locally.
Multiprocessing is disabled by default.
<Added version="2.8.0"/>
storage_options (`dict`, *optional*):
Key/value pairs to be passed on to the file-system backend, if any.
<Added version="2.8.0"/>
Example:
```python
>>> dataset_dict.save_to_disk("path/to/dataset/directory")
>>> dataset_dict.save_to_disk("path/to/dataset/directory", max_shard_size="1GB")
>>> dataset_dict.save_to_disk("path/to/dataset/directory", num_shards={"train": 1024, "test": 8})
```
"""
if fs != "deprecated":
warnings.warn(
"'fs' was deprecated in favor of 'storage_options' in version 2.8.0 and will be removed in 3.0.0.\n"
"You can remove this warning by passing 'storage_options=fs.storage_options' instead.",
FutureWarning,
)
storage_options = fs.storage_options
fs: fsspec.AbstractFileSystem
fs, _, _ = fsspec.get_fs_token_paths(dataset_dict_path, storage_options=storage_options)
if num_shards is None:
num_shards = {k: None for k in self}
elif not isinstance(num_shards, dict):
raise ValueError(
"Please provide one `num_shards` per dataset in the dataset dictionary, e.g. {{'train': 128, 'test': 4}}"
)
fs.makedirs(dataset_dict_path, exist_ok=True)
with fs.open(posixpath.join(dataset_dict_path, config.DATASETDICT_JSON_FILENAME), "w", encoding="utf-8") as f:
json.dump({"splits": list(self)}, f)
for k, dataset in self.items():
dataset.save_to_disk(
posixpath.join(dataset_dict_path, k),
num_shards=num_shards.get(k),
max_shard_size=max_shard_size,
num_proc=num_proc,
storage_options=storage_options,
)
@staticmethod
def load_from_disk(
dataset_dict_path: PathLike,
fs="deprecated",
keep_in_memory: Optional[bool] = None,
storage_options: Optional[dict] = None,
) -> "DatasetDict":
"""
Load a dataset that was previously saved using [`save_to_disk`] from a filesystem using `fsspec.spec.AbstractFileSystem`.
Args:
dataset_dict_path (`str`):
Path (e.g. `"dataset/train"`) or remote URI (e.g. `"s3//my-bucket/dataset/train"`)
of the dataset dict directory where the dataset dict will be loaded from.
fs (`fsspec.spec.AbstractFileSystem`, *optional*):
Instance of the remote filesystem where the dataset will be saved to.
<Deprecated version="2.8.0">
`fs` was deprecated in version 2.8.0 and will be removed in 3.0.0.
Please use `storage_options` instead, e.g. `storage_options=fs.storage_options`
</Deprecated>
keep_in_memory (`bool`, defaults to `None`):
Whether to copy the dataset in-memory. If `None`, the
dataset will not be copied in-memory unless explicitly enabled by setting
`datasets.config.IN_MEMORY_MAX_SIZE` to nonzero. See more details in the
[improve performance](../cache#improve-performance) section.
storage_options (`dict`, *optional*):
Key/value pairs to be passed on to the file-system backend, if any.
<Added version="2.8.0"/>
Returns:
[`DatasetDict`]
Example:
```py
>>> ds = load_from_disk('path/to/dataset/directory')
```
"""
if fs != "deprecated":
warnings.warn(
"'fs' was deprecated in favor of 'storage_options' in version 2.8.0 and will be removed in 3.0.0.\n"
"You can remove this warning by passing 'storage_options=fs.storage_options' instead.",
FutureWarning,
)
storage_options = fs.storage_options
fs: fsspec.AbstractFileSystem
fs, _, [dataset_dict_path] = fsspec.get_fs_token_paths(dataset_dict_path, storage_options=storage_options)
dataset_dict_json_path = posixpath.join(dataset_dict_path, config.DATASETDICT_JSON_FILENAME)
dataset_state_json_path = posixpath.join(dataset_dict_path, config.DATASET_STATE_JSON_FILENAME)
dataset_info_path = posixpath.join(dataset_dict_path, config.DATASET_INFO_FILENAME)
if not fs.isfile(dataset_dict_json_path):
if fs.isfile(dataset_info_path) and fs.isfile(dataset_state_json_path):
raise FileNotFoundError(
f"No such file: '{dataset_dict_json_path}'. Expected to load a `DatasetDict` object, but got a `Dataset`. Please use either `datasets.load_from_disk` or `Dataset.load_from_disk` instead."
)
raise FileNotFoundError(
f"No such file: '{dataset_dict_json_path}'. Expected to load a `DatasetDict` object, but provided path is not a `DatasetDict`."
)
with fs.open(dataset_dict_json_path, "r", encoding="utf-8") as f:
splits = json.load(f)["splits"]
dataset_dict = DatasetDict()
for k in splits:
dataset_dict_split_path = posixpath.join(fs.unstrip_protocol(dataset_dict_path), k)
dataset_dict[k] = Dataset.load_from_disk(
dataset_dict_split_path, keep_in_memory=keep_in_memory, storage_options=storage_options
)
return dataset_dict
@staticmethod
def from_csv(
path_or_paths: Dict[str, PathLike],
features: Optional[Features] = None,
cache_dir: str = None,
keep_in_memory: bool = False,
**kwargs,
) -> "DatasetDict":
"""Create [`DatasetDict`] from CSV file(s).
Args:
path_or_paths (`dict` of path-like):
Path(s) of the CSV file(s).
features ([`Features`], *optional*):
Dataset features.
cache_dir (str, *optional*, defaults to `"~/.cache/huggingface/datasets"`):
Directory to cache data.
keep_in_memory (`bool`, defaults to `False`):
Whether to copy the data in-memory.
**kwargs (additional keyword arguments):
Keyword arguments to be passed to [`pandas.read_csv`].
Returns:
[`DatasetDict`]
Example:
```py
>>> from datasets import DatasetDict
>>> ds = DatasetDict.from_csv({'train': 'path/to/dataset.csv'})
```
"""
# Dynamic import to avoid circular dependency
from .io.csv import CsvDatasetReader
return CsvDatasetReader(
path_or_paths, features=features, cache_dir=cache_dir, keep_in_memory=keep_in_memory, **kwargs
).read()
@staticmethod
def from_json(
path_or_paths: Dict[str, PathLike],
features: Optional[Features] = None,
cache_dir: str = None,
keep_in_memory: bool = False,
**kwargs,
) -> "DatasetDict":
"""Create [`DatasetDict`] from JSON Lines file(s).
Args:
path_or_paths (`path-like` or list of `path-like`):
Path(s) of the JSON Lines file(s).
features ([`Features`], *optional*):
Dataset features.
cache_dir (str, *optional*, defaults to `"~/.cache/huggingface/datasets"`):
Directory to cache data.
keep_in_memory (`bool`, defaults to `False`):
Whether to copy the data in-memory.
**kwargs (additional keyword arguments):
Keyword arguments to be passed to [`JsonConfig`].
Returns:
[`DatasetDict`]
Example:
```py
>>> from datasets import DatasetDict
>>> ds = DatasetDict.from_json({'train': 'path/to/dataset.json'})
```
"""
# Dynamic import to avoid circular dependency
from .io.json import JsonDatasetReader
return JsonDatasetReader(
path_or_paths, features=features, cache_dir=cache_dir, keep_in_memory=keep_in_memory, **kwargs
).read()
@staticmethod
def from_parquet(
path_or_paths: Dict[str, PathLike],
features: Optional[Features] = None,
cache_dir: str = None,
keep_in_memory: bool = False,
columns: Optional[List[str]] = None,
**kwargs,
) -> "DatasetDict":
"""Create [`DatasetDict`] from Parquet file(s).
Args:
path_or_paths (`dict` of path-like):
Path(s) of the CSV file(s).
features ([`Features`], *optional*):
Dataset features.
cache_dir (`str`, *optional*, defaults to `"~/.cache/huggingface/datasets"`):
Directory to cache data.
keep_in_memory (`bool`, defaults to `False`):
Whether to copy the data in-memory.
columns (`List[str]`, *optional*):
If not `None`, only these columns will be read from the file.
A column name may be a prefix of a nested field, e.g. 'a' will select
'a.b', 'a.c', and 'a.d.e'.
**kwargs (additional keyword arguments):
Keyword arguments to be passed to [`ParquetConfig`].
Returns:
[`DatasetDict`]
Example:
```py
>>> from datasets import DatasetDict
>>> ds = DatasetDict.from_parquet({'train': 'path/to/dataset/parquet'})
```
"""
# Dynamic import to avoid circular dependency
from .io.parquet import ParquetDatasetReader
return ParquetDatasetReader(
path_or_paths,
features=features,
cache_dir=cache_dir,
keep_in_memory=keep_in_memory,
columns=columns,
**kwargs,
).read()
@staticmethod
def from_text(
path_or_paths: Dict[str, PathLike],
features: Optional[Features] = None,
cache_dir: str = None,
keep_in_memory: bool = False,
**kwargs,
) -> "DatasetDict":
"""Create [`DatasetDict`] from text file(s).
Args:
path_or_paths (`dict` of path-like):
Path(s) of the text file(s).
features ([`Features`], *optional*):
Dataset features.
cache_dir (`str`, *optional*, defaults to `"~/.cache/huggingface/datasets"`):
Directory to cache data.
keep_in_memory (`bool`, defaults to `False`):
Whether to copy the data in-memory.
**kwargs (additional keyword arguments):
Keyword arguments to be passed to [`TextConfig`].
Returns:
[`DatasetDict`]
Example:
```py
>>> from datasets import DatasetDict
>>> ds = DatasetDict.from_text({'train': 'path/to/dataset.txt'})
```
"""
# Dynamic import to avoid circular dependency
from .io.text import TextDatasetReader
return TextDatasetReader(
path_or_paths, features=features, cache_dir=cache_dir, keep_in_memory=keep_in_memory, **kwargs
).read()
@deprecated()
@is_documented_by(Dataset.prepare_for_task)
def prepare_for_task(self, task: Union[str, TaskTemplate], id: int = 0) -> "DatasetDict":
self._check_values_type()
return DatasetDict({k: dataset.prepare_for_task(task=task, id=id) for k, dataset in self.items()})
@is_documented_by(Dataset.align_labels_with_mapping)
def align_labels_with_mapping(self, label2id: Dict, label_column: str) -> "DatasetDict":
self._check_values_type()
return DatasetDict(
{
k: dataset.align_labels_with_mapping(label2id=label2id, label_column=label_column)
for k, dataset in self.items()
}
)
def push_to_hub(
self,
repo_id,
config_name: str = "default",
commit_message: Optional[str] = None,
private: Optional[bool] = False,
token: Optional[str] = None,
revision: Optional[str] = None,
branch="deprecated",
create_pr: Optional[bool] = False,
max_shard_size: Optional[Union[int, str]] = None,
num_shards: Optional[Dict[str, int]] = None,
embed_external_files: bool = True,
):
"""Pushes the [`DatasetDict`] to the hub as a Parquet dataset.
The [`DatasetDict`] is pushed using HTTP requests and does not need to have neither git or git-lfs installed.
Each dataset split will be pushed independently. The pushed dataset will keep the original split names.
The resulting Parquet files are self-contained by default: if your dataset contains [`Image`] or [`Audio`]
data, the Parquet files will store the bytes of your images or audio files.
You can disable this by setting `embed_external_files` to False.
Args:
repo_id (`str`):
The ID of the repository to push to in the following format: `<user>/<dataset_name>` or
`<org>/<dataset_name>`. Also accepts `<dataset_name>`, which will default to the namespace
of the logged-in user.
config_name (`str`):
Configuration name of a dataset. Defaults to "default".
commit_message (`str`, *optional*):
Message to commit while pushing. Will default to `"Upload dataset"`.
private (`bool`, *optional*):
Whether the dataset repository should be set to private or not. Only affects repository creation:
a repository that already exists will not be affected by that parameter.
token (`str`, *optional*):
An optional authentication token for the Hugging Face Hub. If no token is passed, will default
to the token saved locally when logging in with `huggingface-cli login`. Will raise an error
if no token is passed and the user is not logged-in.
revision (`str`, *optional*):
Branch to push the uploaded files to. Defaults to the `"main"` branch.
<Added version="2.15.0"/>
branch (`str`, *optional*):
The git branch on which to push the dataset. This defaults to the default branch as specified
in your repository, which defaults to `"main"`.
<Deprecated version="2.15.0">
`branch` was deprecated in favor of `revision` in version 2.15.0 and will be removed in 3.0.0.
</Deprecated>
create_pr (`bool`, *optional*, defaults to `False`):
Whether or not to create a PR with the uploaded files or directly commit.
<Added version="2.15.0"/>
max_shard_size (`int` or `str`, *optional*, defaults to `"500MB"`):
The maximum size of the dataset shards to be uploaded to the hub. If expressed as a string, needs to be digits followed by a unit
(like `"500MB"` or `"1GB"`).
num_shards (`Dict[str, int]`, *optional*):
Number of shards to write. By default the number of shards depends on `max_shard_size`.
Use a dictionary to define a different num_shards for each split.
<Added version="2.8.0"/>
embed_external_files (`bool`, defaults to `True`):
Whether to embed file bytes in the shards.
In particular, this will do the following before the push for the fields of type:
- [`Audio`] and [`Image`] removes local path information and embed file content in the Parquet files.
Example:
```python
>>> dataset_dict.push_to_hub("<organization>/<dataset_id>")
>>> dataset_dict.push_to_hub("<organization>/<dataset_id>", private=True)
>>> dataset_dict.push_to_hub("<organization>/<dataset_id>", max_shard_size="1GB")
>>> dataset_dict.push_to_hub("<organization>/<dataset_id>", num_shards={"train": 1024, "test": 8})
```
If you want to add a new configuration (or subset) to a dataset (e.g. if the dataset has multiple tasks/versions/languages):
```python
>>> english_dataset.push_to_hub("<organization>/<dataset_id>", "en")
>>> french_dataset.push_to_hub("<organization>/<dataset_id>", "fr")
>>> # later
>>> english_dataset = load_dataset("<organization>/<dataset_id>", "en")
>>> french_dataset = load_dataset("<organization>/<dataset_id>", "fr")
```
"""
if num_shards is None:
num_shards = {k: None for k in self}
elif not isinstance(num_shards, dict):
raise ValueError(
"Please provide one `num_shards` per dataset in the dataset dictionary, e.g. {{'train': 128, 'test': 4}}"
)
if branch != "deprecated":
warnings.warn(
"'branch' was deprecated in favor of 'revision' in version 2.15.0 and will be removed in 3.0.0.\n"
f"You can remove this warning by passing 'revision={branch}' instead.",
FutureWarning,
)
revision = branch
self._check_values_type()
self._check_values_features()
total_uploaded_size = 0
total_dataset_nbytes = 0
info_to_dump: DatasetInfo = next(iter(self.values())).info.copy()
info_to_dump.config_name = config_name
info_to_dump.splits = SplitDict()
for split in self.keys():
if not re.match(_split_re, split):
raise ValueError(f"Split name should match '{_split_re}' but got '{split}'.")
api = HfApi(endpoint=config.HF_ENDPOINT, token=token)
repo_url = api.create_repo(
repo_id,
token=token,
repo_type="dataset",
private=private,
exist_ok=True,
)
repo_id = repo_url.repo_id
if revision is not None:
api.create_branch(repo_id, branch=revision, token=token, repo_type="dataset", exist_ok=True)
data_dir = config_name if config_name != "default" else "data" # for backward compatibility
additions = []
for split in self.keys():
logger.info(f"Pushing split {split} to the Hub.")
# The split=key needs to be removed before merging
split_additions, uploaded_size, dataset_nbytes = self[split]._push_parquet_shards_to_hub(
repo_id,
data_dir=data_dir,
split=split,
token=token,
revision=revision,
create_pr=create_pr,
max_shard_size=max_shard_size,
num_shards=num_shards.get(split),
embed_external_files=embed_external_files,
)
additions += split_additions
total_uploaded_size += uploaded_size
total_dataset_nbytes += dataset_nbytes
info_to_dump.splits[split] = SplitInfo(str(split), num_bytes=dataset_nbytes, num_examples=len(self[split]))
info_to_dump.download_checksums = None
info_to_dump.download_size = total_uploaded_size
info_to_dump.dataset_size = total_dataset_nbytes
info_to_dump.size_in_bytes = total_uploaded_size + total_dataset_nbytes
metadata_config_to_dump = {
"data_files": [{"split": split, "path": f"{data_dir}/{split}-*"} for split in self.keys()],
}
# Check if the repo already has a README.md and/or a dataset_infos.json to update them with the new split info (size and pattern)
# and delete old split shards (if they exist)
repo_with_dataset_card, repo_with_dataset_infos = False, False
repo_splits = [] # use a list to keep the order of the splits
deletions = []
repo_files_to_add = [addition.path_in_repo for addition in additions]
for repo_file in api.list_files_info(repo_id, revision=revision, repo_type="dataset", token=token):
if repo_file.rfilename == "README.md":
repo_with_dataset_card = True
elif repo_file.rfilename == config.DATASETDICT_INFOS_FILENAME:
repo_with_dataset_infos = True
elif (
repo_file.rfilename.startswith(tuple(f"{data_dir}/{split}-" for split in self.keys()))
and repo_file.rfilename not in repo_files_to_add
):
deletions.append(CommitOperationDelete(path_in_repo=repo_file.rfilename))
elif fnmatch.fnmatch(
repo_file.rfilename, PUSH_TO_HUB_WITHOUT_METADATA_CONFIGS_SPLIT_PATTERN_SHARDED.replace("{split}", "*")
):
repo_split = string_to_dict(
repo_file.rfilename,
glob_pattern_to_regex(PUSH_TO_HUB_WITHOUT_METADATA_CONFIGS_SPLIT_PATTERN_SHARDED),
)["split"]
if repo_split not in repo_splits:
repo_splits.append(split)
# get the info from the README to update them
if repo_with_dataset_card:
dataset_card_path = api.hf_hub_download(repo_id, "README.md", repo_type="dataset", revision=revision)
dataset_card = DatasetCard.load(Path(dataset_card_path))
dataset_card_data = dataset_card.data
metadata_configs = MetadataConfigs.from_dataset_card_data(dataset_card_data)
# get the deprecated dataset_infos.json to update them
elif repo_with_dataset_infos:
dataset_card = None
dataset_card_data = DatasetCardData()
metadata_configs = MetadataConfigs()
else:
dataset_card = None
dataset_card_data = DatasetCardData()
metadata_configs = MetadataConfigs()
# create the metadata configs if it was uploaded with push_to_hub before metadata configs existed
if not metadata_configs and repo_splits:
default_metadata_configs_to_dump = {
"data_files": [{"split": split, "path": f"data/{split}-*"} for split in repo_splits]
}
MetadataConfigs({"default": default_metadata_configs_to_dump}).to_dataset_card_data(dataset_card_data)
# push to the deprecated dataset_infos.json
if repo_with_dataset_infos:
dataset_infos_path = api.hf_hub_download(
repo_id, config.DATASETDICT_INFOS_FILENAME, repo_type="dataset", revision=revision
)
with open(dataset_infos_path, encoding="utf-8") as f:
dataset_infos: dict = json.load(f)
dataset_infos[config_name] = asdict(info_to_dump)
buffer = BytesIO()
buffer.write(json.dumps(dataset_infos, indent=4).encode("utf-8"))
additions.append(
CommitOperationAdd(path_in_repo=config.DATASETDICT_INFOS_FILENAME, path_or_fileobj=buffer)
)
# push to README
DatasetInfosDict({config_name: info_to_dump}).to_dataset_card_data(dataset_card_data)
MetadataConfigs({config_name: metadata_config_to_dump}).to_dataset_card_data(dataset_card_data)
dataset_card = DatasetCard(f"---\n{dataset_card_data}\n---\n") if dataset_card is None else dataset_card
additions.append(CommitOperationAdd(path_in_repo="README.md", path_or_fileobj=str(dataset_card).encode()))
commit_message = commit_message if commit_message is not None else "Upload dataset"
if len(additions) <= config.UPLOADS_MAX_NUMBER_PER_COMMIT:
api.create_commit(
repo_id,
operations=additions + deletions,
commit_message=commit_message,
token=token,
repo_type="dataset",
revision=revision,
create_pr=create_pr,
)
else:
logger.info(
f"Number of files to upload is larger than {config.UPLOADS_MAX_NUMBER_PER_COMMIT}. Splitting the push into multiple commits."
)
num_commits = math.ceil(len(additions) / config.UPLOADS_MAX_NUMBER_PER_COMMIT)
for i in range(0, num_commits):
operations = additions[
i * config.UPLOADS_MAX_NUMBER_PER_COMMIT : (i + 1) * config.UPLOADS_MAX_NUMBER_PER_COMMIT
] + (deletions if i == 0 else [])
api.create_commit(
repo_id,
operations=operations,
commit_message=commit_message + f" (part {i:05d}-of-{num_commits:05d})",
token=token,
repo_type="dataset",
revision=revision,
create_pr=create_pr,
)
logger.info(
f"Commit #{i+1} completed"
+ (f" (still {num_commits - i - 1} to go)" if num_commits - i - 1 else "")
+ "."
)
class IterableDatasetDict(dict):
def with_format(
self,
type: Optional[str] = None,
) -> "IterableDatasetDict":
"""
Return a dataset with the specified format.
This method only supports the "torch" format for now.
The format is set to all the datasets of the dataset dictionary.
Args:
type (`str`, *optional*, defaults to `None`):
If set to "torch", the returned dataset
will be a subclass of `torch.utils.data.IterableDataset` to be used in a `DataLoader`.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", streaming=True)
>>> from transformers import AutoTokenizer
>>> tokenizer = AutoTokenizer.from_pretrained("bert-base-uncased")
>>> def encode(example):
... return tokenizer(examples["text"], truncation=True, padding="max_length")
>>> ds = ds.map(encode, batched=True, remove_columns=["text"])
>>> ds = ds.with_format("torch")
```
"""
return IterableDatasetDict({k: dataset.with_format(type=type) for k, dataset in self.items()})
def map(
self,
function: Optional[Callable] = None,
with_indices: bool = False,
input_columns: Optional[Union[str, List[str]]] = None,
batched: bool = False,
batch_size: int = 1000,
drop_last_batch: bool = False,
remove_columns: Optional[Union[str, List[str]]] = None,
fn_kwargs: Optional[dict] = None,
) -> "IterableDatasetDict":
"""
Apply a function to all the examples in the iterable dataset (individually or in batches) and update them.
If your function returns a column that already exists, then it overwrites it.
The function is applied on-the-fly on the examples when iterating over the dataset.
The transformation is applied to all the datasets of the dataset dictionary.
You can specify whether the function should be batched or not with the `batched` parameter:
- If batched is `False`, then the function takes 1 example in and should return 1 example.
An example is a dictionary, e.g. `{"text": "Hello there !"}`.
- If batched is `True` and `batch_size` is 1, then the function takes a batch of 1 example as input and can return a batch with 1 or more examples.
A batch is a dictionary, e.g. a batch of 1 example is `{"text": ["Hello there !"]}`.
- If batched is `True` and `batch_size` is `n` > 1, then the function takes a batch of `n` examples as input and can return a batch with `n` examples, or with an arbitrary number of examples.
Note that the last batch may have less than `n` examples.
A batch is a dictionary, e.g. a batch of `n` examples is `{"text": ["Hello there !"] * n}`.
Args:
function (`Callable`, *optional*, defaults to `None`):
Function applied on-the-fly on the examples when you iterate on the dataset.
It must have one of the following signatures:
- `function(example: Dict[str, Any]) -> Dict[str, Any]` if `batched=False` and `with_indices=False`
- `function(example: Dict[str, Any], idx: int) -> Dict[str, Any]` if `batched=False` and `with_indices=True`
- `function(batch: Dict[str, List]) -> Dict[str, List]` if `batched=True` and `with_indices=False`
- `function(batch: Dict[str, List], indices: List[int]) -> Dict[str, List]` if `batched=True` and `with_indices=True`
For advanced usage, the function can also return a `pyarrow.Table`.
Moreover if your function returns nothing (`None`), then `map` will run your function and return the dataset unchanged.
If no function is provided, default to identity function: `lambda x: x`.
with_indices (`bool`, defaults to `False`):
Provide example indices to `function`. Note that in this case the signature of `function` should be `def function(example, idx[, rank]): ...`.
input_columns (`[Union[str, List[str]]]`, *optional*, defaults to `None`):
The columns to be passed into `function`
as positional arguments. If `None`, a dict mapping to all formatted columns is passed as one argument.
batched (`bool`, defaults to `False`):
Provide batch of examples to `function`.
batch_size (`int`, *optional*, defaults to `1000`):
Number of examples per batch provided to `function` if `batched=True`.
drop_last_batch (`bool`, defaults to `False`):
Whether a last batch smaller than the `batch_size` should be
dropped instead of being processed by the function.
remove_columns (`[List[str]]`, *optional*, defaults to `None`):
Remove a selection of columns while doing the mapping.
Columns will be removed before updating the examples with the output of `function`, i.e. if `function` is adding
columns with names in `remove_columns`, these columns will be kept.
fn_kwargs (`Dict`, *optional*, defaults to `None`):
Keyword arguments to be passed to `function`
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", streaming=True)
>>> def add_prefix(example):
... example["text"] = "Review: " + example["text"]
... return example
>>> ds = ds.map(add_prefix)
>>> next(iter(ds["train"]))
{'label': 1,
'text': 'Review: the rock is destined to be the 21st century\'s new " conan " and that he\'s going to make a splash even greater than arnold schwarzenegger , jean-claud van damme or steven segal .'}
```
"""
return IterableDatasetDict(
{
k: dataset.map(
function=function,
with_indices=with_indices,
input_columns=input_columns,
batched=batched,
batch_size=batch_size,
drop_last_batch=drop_last_batch,
remove_columns=remove_columns,
fn_kwargs=fn_kwargs,
)
for k, dataset in self.items()
}
)
def filter(
self,
function: Optional[Callable] = None,
with_indices=False,
input_columns: Optional[Union[str, List[str]]] = None,
batched: bool = False,
batch_size: Optional[int] = 1000,
fn_kwargs: Optional[dict] = None,
) -> "IterableDatasetDict":
"""Apply a filter function to all the elements so that the dataset only includes examples according to the filter function.
The filtering is done on-the-fly when iterating over the dataset.
The filtering is applied to all the datasets of the dataset dictionary.
Args:
function (`Callable`):
Callable with one of the following signatures:
- `function(example: Dict[str, Any]) -> bool` if `with_indices=False, batched=False`
- `function(example: Dict[str, Any], indices: int) -> bool` if `with_indices=True, batched=False`
- `function(example: Dict[str, List]) -> List[bool]` if `with_indices=False, batched=True`
- `function(example: Dict[str, List], indices: List[int]) -> List[bool]` if `with_indices=True, batched=True`
If no function is provided, defaults to an always True function: `lambda x: True`.
with_indices (`bool`, defaults to `False`):
Provide example indices to `function`. Note that in this case the signature of `function` should be `def function(example, idx): ...`.
input_columns (`str` or `List[str]`, *optional*):
The columns to be passed into `function` as
positional arguments. If `None`, a dict mapping to all formatted columns is passed as one argument.
batched (`bool`, defaults to `False`):
Provide batch of examples to `function`
batch_size (`int`, *optional*, defaults to `1000`):
Number of examples per batch provided to `function` if `batched=True`.
fn_kwargs (`Dict`, *optional*, defaults to `None`):
Keyword arguments to be passed to `function`
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", streaming=True)
>>> ds = ds.filter(lambda x: x["label"] == 0)
>>> list(ds["train"].take(3))
[{'label': 0, 'text': 'Review: simplistic , silly and tedious .'},
{'label': 0,
'text': "Review: it's so laddish and juvenile , only teenage boys could possibly find it funny ."},
{'label': 0,
'text': 'Review: exploitative and largely devoid of the depth or sophistication that would make watching such a graphic treatment of the crimes bearable .'}]
```
"""
return IterableDatasetDict(
{
k: dataset.filter(
function=function,
with_indices=with_indices,
input_columns=input_columns,
batched=batched,
batch_size=batch_size,
fn_kwargs=fn_kwargs,
)
for k, dataset in self.items()
}
)
def shuffle(
self, seed=None, generator: Optional[np.random.Generator] = None, buffer_size: int = 1000
) -> "IterableDatasetDict":
"""
Randomly shuffles the elements of this dataset.
The shuffling is applied to all the datasets of the dataset dictionary.
This dataset fills a buffer with buffer_size elements, then randomly samples elements from this buffer,
replacing the selected elements with new elements. For perfect shuffling, a buffer size greater than or
equal to the full size of the dataset is required.
For instance, if your dataset contains 10,000 elements but `buffer_size` is set to 1000, then `shuffle` will
initially select a random element from only the first 1000 elements in the buffer. Once an element is
selected, its space in the buffer is replaced by the next (i.e. 1,001-st) element,
maintaining the 1000 element buffer.
If the dataset is made of several shards, it also does `shuffle` the order of the shards.
However if the order has been fixed by using [`~datasets.IterableDataset.skip`] or [`~datasets.IterableDataset.take`]
then the order of the shards is kept unchanged.
Args:
seed (`int`, *optional*, defaults to `None`):
Random seed that will be used to shuffle the dataset.
It is used to sample from the shuffle buffer and also to shuffle the data shards.
generator (`numpy.random.Generator`, *optional*):
Numpy random Generator to use to compute the permutation of the dataset rows.
If `generator=None` (default), uses `np.random.default_rng` (the default BitGenerator (PCG64) of NumPy).
buffer_size (`int`, defaults to `1000`):
Size of the buffer.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", streaming=True)
>>> list(ds["train"].take(3))
[{'label': 1,
'text': 'the rock is destined to be the 21st century\'s new " conan " and that he\'s going to make a splash even greater than arnold schwarzenegger , jean-claud van damme or steven segal .'},
{'label': 1,
'text': 'the gorgeously elaborate continuation of " the lord of the rings " trilogy is so huge that a column of words cannot adequately describe co-writer/director peter jackson\'s expanded vision of j . r . r . tolkien\'s middle-earth .'},
{'label': 1, 'text': 'effective but too-tepid biopic'}]
>>> ds = ds.shuffle(seed=42)
>>> list(ds["train"].take(3))
[{'label': 1,
'text': "a sports movie with action that's exciting on the field and a story you care about off it ."},
{'label': 1,
'text': 'at its best , the good girl is a refreshingly adult take on adultery . . .'},
{'label': 1,
'text': "sam jones became a very lucky filmmaker the day wilco got dropped from their record label , proving that one man's ruin may be another's fortune ."}]
```
"""
return IterableDatasetDict(
{
k: dataset.shuffle(seed=seed, generator=generator, buffer_size=buffer_size)
for k, dataset in self.items()
}
)
def rename_column(self, original_column_name: str, new_column_name: str) -> "IterableDatasetDict":
"""
Rename a column in the dataset, and move the features associated to the original column under the new column
name.
The renaming is applied to all the datasets of the dataset dictionary.
Args:
original_column_name (`str`):
Name of the column to rename.
new_column_name (`str`):
New name for the column.
Returns:
[`IterableDatasetDict`]: A copy of the dataset with a renamed column.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", streaming=True)
>>> ds = ds.rename_column("text", "movie_review")
>>> next(iter(ds["train"]))
{'label': 1,
'movie_review': 'the rock is destined to be the 21st century\'s new " conan " and that he\'s going to make a splash even greater than arnold schwarzenegger , jean-claud van damme or steven segal .'}
```
"""
return IterableDatasetDict(
{
k: dataset.rename_column(original_column_name=original_column_name, new_column_name=new_column_name)
for k, dataset in self.items()
}
)
def rename_columns(self, column_mapping: Dict[str, str]) -> "IterableDatasetDict":
"""
Rename several columns in the dataset, and move the features associated to the original columns under
the new column names.
The renaming is applied to all the datasets of the dataset dictionary.
Args:
column_mapping (`Dict[str, str]`):
A mapping of columns to rename to their new names.
Returns:
[`IterableDatasetDict`]: A copy of the dataset with renamed columns
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", streaming=True)
>>> ds = ds.rename_columns({"text": "movie_review", "label": "rating"})
>>> next(iter(ds["train"]))
{'movie_review': 'the rock is destined to be the 21st century\'s new " conan " and that he\'s going to make a splash even greater than arnold schwarzenegger , jean-claud van damme or steven segal .',
'rating': 1}
```
"""
return IterableDatasetDict(
{k: dataset.rename_columns(column_mapping=column_mapping) for k, dataset in self.items()}
)
def remove_columns(self, column_names: Union[str, List[str]]) -> "IterableDatasetDict":
"""
Remove one or several column(s) in the dataset and the features associated to them.
The removal is done on-the-fly on the examples when iterating over the dataset.
The removal is applied to all the datasets of the dataset dictionary.
Args:
column_names (`Union[str, List[str]]`):
Name of the column(s) to remove.
Returns:
[`IterableDatasetDict`]: A copy of the dataset object without the columns to remove.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", streaming=True)
>>> ds = ds.remove_columns("label")
>>> next(iter(ds["train"]))
{'text': 'the rock is destined to be the 21st century\'s new " conan " and that he\'s going to make a splash even greater than arnold schwarzenegger , jean-claud van damme or steven segal .'}
```
"""
return IterableDatasetDict({k: dataset.remove_columns(column_names) for k, dataset in self.items()})
def select_columns(self, column_names: Union[str, List[str]]) -> "IterableDatasetDict":
"""Select one or several column(s) in the dataset and the features
associated to them. The selection is done on-the-fly on the examples
when iterating over the dataset. The selection is applied to all the
datasets of the dataset dictionary.
Args:
column_names (`Union[str, List[str]]`):
Name of the column(s) to keep.
Returns:
[`IterableDatasetDict`]: A copy of the dataset object with only selected columns.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", streaming=True)
>>> ds = ds.select("text")
>>> next(iter(ds["train"]))
{'text': 'the rock is destined to be the 21st century\'s new " conan " and that he\'s going to make a splash even greater than arnold schwarzenegger , jean-claud van damme or steven segal .'}
```
"""
return IterableDatasetDict({k: dataset.select_columns(column_names) for k, dataset in self.items()})
def cast_column(self, column: str, feature: FeatureType) -> "IterableDatasetDict":
"""Cast column to feature for decoding.
The type casting is applied to all the datasets of the dataset dictionary.
Args:
column (`str`):
Column name.
feature ([`Feature`]):
Target feature.
Returns:
[`IterableDatasetDict`]
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", streaming=True)
>>> ds["train"].features
{'label': ClassLabel(num_classes=2, names=['neg', 'pos'], id=None),
'text': Value(dtype='string', id=None)}
>>> ds = ds.cast_column('label', ClassLabel(names=['bad', 'good']))
>>> ds["train"].features
{'label': ClassLabel(num_classes=2, names=['bad', 'good'], id=None),
'text': Value(dtype='string', id=None)}
```
"""
return IterableDatasetDict(
{k: dataset.cast_column(column=column, feature=feature) for k, dataset in self.items()}
)
def cast(
self,
features: Features,
) -> "IterableDatasetDict":
"""
Cast the dataset to a new set of features.
The type casting is applied to all the datasets of the dataset dictionary.
Args:
features (`Features`):
New features to cast the dataset to.
The name of the fields in the features must match the current column names.
The type of the data must also be convertible from one type to the other.
For non-trivial conversion, e.g. `string` <-> `ClassLabel` you should use [`map`] to update the Dataset.
Returns:
[`IterableDatasetDict`]: A copy of the dataset with casted features.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", streaming=True)
>>> ds["train"].features
{'label': ClassLabel(num_classes=2, names=['neg', 'pos'], id=None),
'text': Value(dtype='string', id=None)}
>>> new_features = ds["train"].features.copy()
>>> new_features['label'] = ClassLabel(names=['bad', 'good'])
>>> new_features['text'] = Value('large_string')
>>> ds = ds.cast(new_features)
>>> ds["train"].features
{'label': ClassLabel(num_classes=2, names=['bad', 'good'], id=None),
'text': Value(dtype='large_string', id=None)}
```
"""
return IterableDatasetDict({k: dataset.cast(features=features) for k, dataset in self.items()})
| 0
|
hf_public_repos/datasets/src
|
hf_public_repos/datasets/src/datasets/arrow_dataset.py
|
# Copyright 2020 The HuggingFace Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# Lint as: python3
""" Simple Dataset wrapping an Arrow Table."""
import contextlib
import copy
import fnmatch
import itertools
import json
import math
import os
import posixpath
import re
import shutil
import sys
import tempfile
import time
import warnings
import weakref
from collections import Counter
from collections.abc import Mapping
from copy import deepcopy
from functools import partial, wraps
from io import BytesIO
from math import ceil, floor
from pathlib import Path
from random import sample
from typing import (
TYPE_CHECKING,
Any,
BinaryIO,
Callable,
Dict,
Iterable,
Iterator,
List,
Optional,
Tuple,
Union,
overload,
)
from typing import Sequence as Sequence_
import fsspec
import numpy as np
import pandas as pd
import pyarrow as pa
import pyarrow.compute as pc
from huggingface_hub import CommitOperationAdd, CommitOperationDelete, DatasetCard, DatasetCardData, HfApi
from multiprocess import Pool
from requests import HTTPError
from . import config
from .arrow_reader import ArrowReader
from .arrow_writer import ArrowWriter, OptimizedTypedSequence
from .data_files import sanitize_patterns
from .download.streaming_download_manager import xgetsize
from .features import Audio, ClassLabel, Features, Image, Sequence, Value
from .features.features import (
FeatureType,
_align_features,
_check_if_features_can_be_aligned,
generate_from_arrow_type,
pandas_types_mapper,
require_decoding,
)
from .filesystems import is_remote_filesystem
from .fingerprint import (
fingerprint_transform,
format_kwargs_for_fingerprint,
format_transform_for_fingerprint,
generate_fingerprint,
generate_random_fingerprint,
get_temporary_cache_files_directory,
is_caching_enabled,
maybe_register_dataset_for_temp_dir_deletion,
update_fingerprint,
validate_fingerprint,
)
from .formatting import format_table, get_format_type_from_alias, get_formatter, query_table
from .formatting.formatting import LazyDict, _is_range_contiguous
from .info import DatasetInfo, DatasetInfosDict
from .naming import _split_re
from .search import IndexableMixin
from .splits import NamedSplit, Split, SplitDict, SplitInfo
from .table import (
InMemoryTable,
MemoryMappedTable,
Table,
_memory_mapped_record_batch_reader_from_file,
cast_array_to_feature,
concat_tables,
embed_table_storage,
list_table_cache_files,
table_cast,
table_iter,
table_visitor,
)
from .tasks import TaskTemplate
from .utils import logging
from .utils import tqdm as hf_tqdm
from .utils.deprecation_utils import deprecated
from .utils.file_utils import _retry, estimate_dataset_size
from .utils.info_utils import is_small_dataset
from .utils.metadata import MetadataConfigs
from .utils.py_utils import (
Literal,
asdict,
convert_file_size_to_int,
glob_pattern_to_regex,
iflatmap_unordered,
string_to_dict,
unique_values,
)
from .utils.stratify import stratified_shuffle_split_generate_indices
from .utils.tf_utils import dataset_to_tf, minimal_tf_collate_fn, multiprocess_dataset_to_tf
from .utils.typing import ListLike, PathLike
if TYPE_CHECKING:
import sqlite3
import pyspark
import sqlalchemy
from .dataset_dict import DatasetDict
from .iterable_dataset import IterableDataset
logger = logging.get_logger(__name__)
PUSH_TO_HUB_WITHOUT_METADATA_CONFIGS_SPLIT_PATTERN_SHARDED = (
"data/{split}-[0-9][0-9][0-9][0-9][0-9]-of-[0-9][0-9][0-9][0-9][0-9]*.parquet"
)
class DatasetInfoMixin:
"""This base class exposes some attributes of DatasetInfo
at the base level of the Dataset for easy access.
"""
def __init__(self, info: DatasetInfo, split: Optional[NamedSplit]):
self._info = info
self._split = split
@property
def info(self):
"""[`~datasets.DatasetInfo`] object containing all the metadata in the dataset."""
return self._info
@property
def split(self):
"""[`~datasets.NamedSplit`] object corresponding to a named dataset split."""
return self._split
@property
def builder_name(self) -> str:
return self._info.builder_name
@property
def citation(self) -> str:
return self._info.citation
@property
def config_name(self) -> str:
return self._info.config_name
@property
def dataset_size(self) -> Optional[int]:
return self._info.dataset_size
@property
def description(self) -> str:
return self._info.description
@property
def download_checksums(self) -> Optional[dict]:
return self._info.download_checksums
@property
def download_size(self) -> Optional[int]:
return self._info.download_size
@property
def features(self) -> Optional[Features]:
return self._info.features.copy() if self._info.features is not None else None
@property
def homepage(self) -> Optional[str]:
return self._info.homepage
@property
def license(self) -> Optional[str]:
return self._info.license
@property
def size_in_bytes(self) -> Optional[int]:
return self._info.size_in_bytes
@property
def supervised_keys(self):
return self._info.supervised_keys
@property
def task_templates(self):
return self._info.task_templates
@property
def version(self):
return self._info.version
class TensorflowDatasetMixin:
_TF_DATASET_REFS = set()
@staticmethod
def _get_output_signature(
dataset: "Dataset",
collate_fn: Callable,
collate_fn_args: dict,
cols_to_retain: Optional[List[str]] = None,
batch_size: Optional[int] = None,
num_test_batches: int = 20,
):
"""Private method used by `to_tf_dataset()` to find the shapes and dtypes of samples from this dataset
after being passed through the collate_fn. Tensorflow needs an exact signature for tf.numpy_function, so
the only way to do this is to run test batches - the collator may add or rename columns, so we can't figure
it out just by inspecting the dataset.
Args:
dataset (`Dataset`): Dataset to load samples from.
collate_fn(`bool`): Shuffle the dataset order when loading. Recommended True for training, False for
validation/evaluation.
collate_fn(`Callable`): A function or callable object (such as a `DataCollator`) that will collate
lists of samples into a batch.
collate_fn_args (`Dict`): A `dict` of keyword arguments to be passed to the
`collate_fn`.
batch_size (`int`, optional): The size of batches loaded from the dataset. Used for shape inference.
Can be None, which indicates that batch sizes can be variable.
num_test_batches (`int`): The number of batches to load from the dataset for shape inference.
Returns:
`dict`: Dict mapping column names to tf.Tensorspec objects
`dict`: Dict mapping column names to np.dtype objects
"""
if config.TF_AVAILABLE:
import tensorflow as tf
else:
raise ImportError("Called a Tensorflow-specific function but Tensorflow is not installed.")
if len(dataset) == 0:
raise ValueError("Unable to get the output signature because the dataset is empty.")
if batch_size is not None:
batch_size = min(len(dataset), batch_size)
test_batch_size = 1
if cols_to_retain is not None:
cols_to_retain = list(set(cols_to_retain + ["label_ids", "label", "labels"]))
test_batches = []
for _ in range(num_test_batches):
indices = sample(range(len(dataset)), test_batch_size)
test_batch = dataset[indices]
if cols_to_retain is not None:
test_batch = {key: value for key, value in test_batch.items() if key in cols_to_retain}
test_batch = [{key: value[i] for key, value in test_batch.items()} for i in range(test_batch_size)]
test_batch = collate_fn(test_batch, **collate_fn_args)
test_batches.append(test_batch)
tf_columns_to_signatures = {}
np_columns_to_dtypes = {}
for column in test_batches[0].keys():
raw_arrays = [batch[column] for batch in test_batches]
# In case the collate_fn returns something strange
np_arrays = []
for array in raw_arrays:
if isinstance(array, np.ndarray):
np_arrays.append(array)
elif isinstance(array, tf.Tensor):
np_arrays.append(array.numpy())
else:
np_arrays.append(np.array(array))
if np.issubdtype(np_arrays[0].dtype, np.integer) or np_arrays[0].dtype == bool:
tf_dtype = tf.int64
np_dtype = np.int64
elif np.issubdtype(np_arrays[0].dtype, np.number):
tf_dtype = tf.float32
np_dtype = np.float32
elif np_arrays[0].dtype.kind == "U": # Unicode strings
np_dtype = np.unicode_
tf_dtype = tf.string
else:
raise RuntimeError(
f"Unrecognized array dtype {np_arrays[0].dtype}. \n"
"Nested types and image/audio types are not supported yet."
)
shapes = [array.shape for array in np_arrays]
static_shape = []
for dim in range(len(shapes[0])):
sizes = {shape[dim] for shape in shapes}
if dim == 0:
static_shape.append(batch_size)
continue
if len(sizes) == 1: # This dimension looks constant
static_shape.append(sizes.pop())
else: # Use None for variable dimensions
static_shape.append(None)
tf_columns_to_signatures[column] = tf.TensorSpec(shape=static_shape, dtype=tf_dtype)
np_columns_to_dtypes[column] = np_dtype
return tf_columns_to_signatures, np_columns_to_dtypes
def to_tf_dataset(
self,
batch_size: Optional[int] = None,
columns: Optional[Union[str, List[str]]] = None,
shuffle: bool = False,
collate_fn: Optional[Callable] = None,
drop_remainder: bool = False,
collate_fn_args: Optional[Dict[str, Any]] = None,
label_cols: Optional[Union[str, List[str]]] = None,
prefetch: bool = True,
num_workers: int = 0,
num_test_batches: int = 20,
):
"""Create a `tf.data.Dataset` from the underlying Dataset. This `tf.data.Dataset` will load and collate batches from
the Dataset, and is suitable for passing to methods like `model.fit()` or `model.predict()`. The dataset will yield
`dicts` for both inputs and labels unless the `dict` would contain only a single key, in which case a raw
`tf.Tensor` is yielded instead.
Args:
batch_size (`int`, *optional*):
Size of batches to load from the dataset. Defaults to `None`, which implies that the dataset won't be
batched, but the returned dataset can be batched later with `tf_dataset.batch(batch_size)`.
columns (`List[str]` or `str`, *optional*):
Dataset column(s) to load in the `tf.data.Dataset`.
Column names that are created by the `collate_fn` and that do not exist in the original dataset can be used.
shuffle(`bool`, defaults to `False`):
Shuffle the dataset order when loading. Recommended `True` for training, `False` for
validation/evaluation.
drop_remainder(`bool`, defaults to `False`):
Drop the last incomplete batch when loading. Ensures
that all batches yielded by the dataset will have the same length on the batch dimension.
collate_fn(`Callable`, *optional*):
A function or callable object (such as a `DataCollator`) that will collate
lists of samples into a batch.
collate_fn_args (`Dict`, *optional*):
An optional `dict` of keyword arguments to be passed to the
`collate_fn`.
label_cols (`List[str]` or `str`, defaults to `None`):
Dataset column(s) to load as labels.
Note that many models compute loss internally rather than letting Keras do it, in which case
passing the labels here is optional, as long as they're in the input `columns`.
prefetch (`bool`, defaults to `True`):
Whether to run the dataloader in a separate thread and maintain
a small buffer of batches for training. Improves performance by allowing data to be loaded in the
background while the model is training.
num_workers (`int`, defaults to `0`):
Number of workers to use for loading the dataset. Only supported on Python versions >= 3.8.
num_test_batches (`int`, defaults to `20`):
Number of batches to use to infer the output signature of the dataset.
The higher this number, the more accurate the signature will be, but the longer it will take to
create the dataset.
Returns:
`tf.data.Dataset`
Example:
```py
>>> ds_train = ds["train"].to_tf_dataset(
... columns=['input_ids', 'token_type_ids', 'attention_mask', 'label'],
... shuffle=True,
... batch_size=16,
... collate_fn=data_collator,
... )
```
"""
if config.TF_AVAILABLE:
import tensorflow as tf
else:
raise ImportError("Called a Tensorflow-specific function but Tensorflow is not installed.")
if (isinstance(columns, list) and len(columns) == 1) or (
isinstance(label_cols, list) and len(label_cols) == 1
):
warnings.warn(
"The output of `to_tf_dataset` will change when a passing single element list for `labels` or "
"`columns` in the next datasets version. To return a tuple structure rather than dict, pass a "
"single string.\n"
"Old behaviour: columns=['a'], labels=['labels'] -> (tf.Tensor, tf.Tensor) \n"
" : columns='a', labels='labels' -> (tf.Tensor, tf.Tensor) \n"
"New behaviour: columns=['a'],labels=['labels'] -> ({'a': tf.Tensor}, {'labels': tf.Tensor}) \n"
" : columns='a', labels='labels' -> (tf.Tensor, tf.Tensor) ",
FutureWarning,
)
if isinstance(tf.distribute.get_strategy(), tf.distribute.TPUStrategy):
logger.warning(
"Note that to_tf_dataset() loads the data with a generator rather than a full tf.data "
"pipeline and is not compatible with remote TPU connections. If you encounter errors, please "
"try using a TPU VM or, if your data can fit in memory, loading it into memory as a dict of "
"Tensors instead of streaming with to_tf_dataset()."
)
if collate_fn is None:
# Set a very simple default collator that just stacks things together
collate_fn = minimal_tf_collate_fn
if collate_fn_args is None:
collate_fn_args = {}
if label_cols and not columns:
raise ValueError("Cannot specify label_cols without specifying columns!")
if label_cols is None:
label_cols = []
elif isinstance(label_cols, str):
label_cols = [label_cols]
if len(set(label_cols)) < len(label_cols):
raise ValueError("List of label_cols contains duplicates.")
if columns:
if isinstance(columns, str):
columns = [columns]
if len(set(columns)) < len(columns):
raise ValueError("List of columns contains duplicates.")
cols_to_retain = list(set(columns + label_cols))
else:
cols_to_retain = None # Indicates keeping all valid columns
columns = []
if self.format["type"] not in ["custom", "numpy"]:
dataset = self.with_format("numpy")
else:
dataset = self
# TODO(Matt, QL): deprecate the retention of label_ids and label
output_signature, columns_to_np_types = dataset._get_output_signature(
dataset,
collate_fn=collate_fn,
collate_fn_args=collate_fn_args,
cols_to_retain=cols_to_retain,
batch_size=batch_size if drop_remainder and batch_size is not None else None,
num_test_batches=num_test_batches,
)
if "labels" in output_signature:
if ("label_ids" in columns or "label" in columns) and "labels" not in columns:
columns = [col for col in columns if col not in ["label_ids", "label"]] + ["labels"]
if ("label_ids" in label_cols or "label" in label_cols) and "labels" not in label_cols:
label_cols = [col for col in label_cols if col not in ["label_ids", "label"]] + ["labels"]
for col in columns:
if col not in output_signature:
raise ValueError(f"Column {col} not found in dataset!")
for col in label_cols:
if col not in output_signature:
raise ValueError(f"Label column {col} not found in dataset!")
if num_workers == 0:
tf_dataset = dataset_to_tf(
dataset=dataset,
cols_to_retain=cols_to_retain,
collate_fn=collate_fn,
collate_fn_args=collate_fn_args,
columns_to_np_types=columns_to_np_types,
output_signature=output_signature,
shuffle=shuffle,
batch_size=batch_size,
drop_remainder=drop_remainder,
)
elif num_workers > 0:
if batch_size is None:
raise NotImplementedError(
"`batch_size` must be specified when using multiple workers, as unbatched multiprocessing "
"is not supported yet. Please provide a `batch_size` if `num_workers` is greater than 0."
)
tf_dataset = multiprocess_dataset_to_tf(
dataset=dataset,
cols_to_retain=cols_to_retain,
collate_fn=collate_fn,
collate_fn_args=collate_fn_args,
columns_to_np_types=columns_to_np_types,
output_signature=output_signature,
shuffle=shuffle,
batch_size=batch_size,
drop_remainder=drop_remainder,
num_workers=num_workers,
)
else:
raise ValueError("num_workers must be >= 0")
def split_features_and_labels(input_batch):
# TODO(Matt, QL): deprecate returning the dict content when there's only one key
features = {key: tensor for key, tensor in input_batch.items() if key in columns}
labels = {key: tensor for key, tensor in input_batch.items() if key in label_cols}
if len(features) == 1:
features = list(features.values())[0]
if len(labels) == 1:
labels = list(labels.values())[0]
if isinstance(labels, dict) and len(labels) == 0:
return features
else:
return features, labels
if cols_to_retain is not None:
tf_dataset = tf_dataset.map(split_features_and_labels)
if prefetch:
tf_dataset = tf_dataset.prefetch(tf.data.experimental.AUTOTUNE)
# Remove a reference to the open Arrow file on delete
def cleanup_callback(ref):
dataset.__del__()
self._TF_DATASET_REFS.remove(ref)
self._TF_DATASET_REFS.add(weakref.ref(tf_dataset, cleanup_callback))
return tf_dataset
class DatasetTransformationNotAllowedError(Exception):
pass
def transmit_format(func):
"""Wrapper for dataset transforms that recreate a new Dataset to transmit the format of the original dataset to the new dataset"""
@wraps(func)
def wrapper(*args, **kwargs):
if args:
self: "Dataset" = args[0]
args = args[1:]
else:
self: "Dataset" = kwargs.pop("self")
# don't use self.format since it returns a list of columns for 'columns' even if self_format_columns is None
unformatted_columns = set(self.column_names) - set(self._format_columns or [])
self_format = {
"type": self._format_type,
"format_kwargs": self._format_kwargs,
"columns": self._format_columns,
"output_all_columns": self._output_all_columns,
}
# apply actual function
out: Union["Dataset", "DatasetDict"] = func(self, *args, **kwargs)
datasets: List["Dataset"] = list(out.values()) if isinstance(out, dict) else [out]
# re-apply format to the output
for dataset in datasets:
new_format = self_format.copy()
if new_format["columns"] is not None: # new formatted columns = (columns - previously unformatted columns)
# sort the columns to have a deterministic list of columns that we can compare with `out_format`
new_format["columns"] = sorted(set(dataset.column_names) - unformatted_columns)
out_format = {
"type": dataset._format_type,
"format_kwargs": dataset._format_kwargs,
"columns": sorted(dataset._format_columns) if dataset._format_columns is not None else None,
"output_all_columns": dataset._output_all_columns,
}
if out_format != new_format:
fingerprint = dataset._fingerprint
dataset.set_format(**new_format)
dataset._fingerprint = fingerprint
return out
wrapper._decorator_name_ = "transmit_format"
return wrapper
def transmit_tasks(func):
"""Wrapper for dataset transforms that recreate a new Dataset to transmit the task templates of the original dataset to the new dataset"""
@wraps(func)
def wrapper(*args, **kwargs):
if args:
self: "Dataset" = args[0]
args = args[1:]
else:
self: "Dataset" = kwargs.pop("self")
# apply actual function
out: Union["Dataset", "DatasetDict"] = func(self, *args, **kwargs)
datasets: List["Dataset"] = list(out.values()) if isinstance(out, dict) else [out]
for dataset in datasets:
# Remove task templates if a column mapping of the template is no longer valid
if self.info.task_templates is not None:
dataset.info.task_templates = [
template
for template in self.info.task_templates
if all(
dataset._info.features.get(k) == self._info.features.get(k)
for k in template.column_mapping.keys()
)
]
return out
wrapper._decorator_name_ = "transmit_tasks"
return wrapper
def update_metadata_with_features(table: Table, features: Features):
"""To be used in dataset transforms that modify the features of the dataset, in order to update the features stored in the metadata of its schema."""
features = Features({col_name: features[col_name] for col_name in table.column_names})
if table.schema.metadata is None or b"huggingface" not in table.schema.metadata:
pa_metadata = ArrowWriter._build_metadata(DatasetInfo(features=features))
else:
metadata = json.loads(table.schema.metadata[b"huggingface"].decode())
if "info" not in metadata:
metadata["info"] = asdict(DatasetInfo(features=features))
else:
metadata["info"]["features"] = asdict(DatasetInfo(features=features))["features"]
pa_metadata = {"huggingface": json.dumps(metadata)}
table = table.replace_schema_metadata(pa_metadata)
return table
def _check_table(table) -> Table:
"""We check the table type to make sure it's an instance of :class:`datasets.table.Table`"""
if isinstance(table, pa.Table):
# for a pyarrow table, we can just consider it as a in-memory table
# this is here for backward compatibility
return InMemoryTable(table)
elif isinstance(table, Table):
return table
else:
raise TypeError(f"Expected a pyarrow.Table or a datasets.table.Table object, but got {table}.")
def _check_column_names(column_names: List[str]):
"""Check the column names to make sure they don't contain duplicates."""
counter = Counter(column_names)
if not all(count == 1 for count in counter.values()):
duplicated_columns = [col for col in counter if counter[col] > 1]
raise ValueError(f"The table can't have duplicated columns but columns {duplicated_columns} are duplicated.")
def _check_valid_indices_value(index, size):
if (index < 0 and index + size < 0) or (index >= size):
raise IndexError(f"Index {index} out of range for dataset of size {size}.")
class NonExistentDatasetError(Exception):
"""Used when we expect the existence of a dataset"""
pass
class Dataset(DatasetInfoMixin, IndexableMixin, TensorflowDatasetMixin):
"""A Dataset backed by an Arrow table."""
def __init__(
self,
arrow_table: Table,
info: Optional[DatasetInfo] = None,
split: Optional[NamedSplit] = None,
indices_table: Optional[Table] = None,
fingerprint: Optional[str] = None,
):
info = info.copy() if info is not None else DatasetInfo()
DatasetInfoMixin.__init__(self, info=info, split=split)
IndexableMixin.__init__(self)
self._data: Table = _check_table(arrow_table)
self._indices: Optional[Table] = _check_table(indices_table) if indices_table is not None else None
maybe_register_dataset_for_temp_dir_deletion(self)
self._format_type: Optional[str] = None
self._format_kwargs: dict = {}
self._format_columns: Optional[list] = None
self._output_all_columns: bool = False
self._fingerprint: str = fingerprint
# Read metadata
if self._data.schema.metadata is not None and b"huggingface" in self._data.schema.metadata:
metadata = json.loads(self._data.schema.metadata[b"huggingface"].decode())
if (
"fingerprint" in metadata and self._fingerprint is None
): # try to load fingerprint from the arrow file metadata
self._fingerprint = metadata["fingerprint"]
# Infer features if None
inferred_features = Features.from_arrow_schema(arrow_table.schema)
if self.info.features is None:
self.info.features = inferred_features
else: # make sure the nested columns are in the right order
try:
self.info.features = self.info.features.reorder_fields_as(inferred_features)
except ValueError as e:
raise ValueError(
f"{e}\nThe 'source' features come from dataset_info.json, and the 'target' ones are those of the dataset arrow file."
)
# Infer fingerprint if None
if self._fingerprint is None:
self._fingerprint = generate_fingerprint(self)
# Sanity checks
if self._info.features is None:
raise ValueError("Features can't be None in a Dataset object")
if self._fingerprint is None:
raise ValueError("Fingerprint can't be None in a Dataset object")
if self.info.features.type != inferred_features.type:
raise ValueError(
f"External features info don't match the dataset:\nGot\n{self.info.features}\nwith type\n{self.info.features.type}\n\nbut expected something like\n{inferred_features}\nwith type\n{inferred_features.type}"
)
if self._indices is not None:
if not pa.types.is_unsigned_integer(self._indices.column(0).type):
raise ValueError(
f"indices must be an Arrow table of unsigned integers, current type is {self._indices.column(0).type}"
)
_check_column_names(self._data.column_names)
self._data = update_metadata_with_features(self._data, self._info.features)
@property
def features(self) -> Features:
features = super().features
if features is None: # this is already checked in __init__
raise ValueError("Features can't be None in a Dataset object")
return features
@classmethod
def from_file(
cls,
filename: str,
info: Optional[DatasetInfo] = None,
split: Optional[NamedSplit] = None,
indices_filename: Optional[str] = None,
in_memory: bool = False,
) -> "Dataset":
"""Instantiate a Dataset backed by an Arrow table at filename.
Args:
filename (`str`):
File name of the dataset.
info (`DatasetInfo`, *optional*):
Dataset information, like description, citation, etc.
split (`NamedSplit`, *optional*):
Name of the dataset split.
indices_filename (`str`, *optional*):
File names of the indices.
in_memory (`bool`, defaults to `False`):
Whether to copy the data in-memory.
Returns:
[`Dataset`]
"""
table = ArrowReader.read_table(filename, in_memory=in_memory)
if indices_filename is not None:
indices_pa_table = ArrowReader.read_table(indices_filename, in_memory=in_memory)
else:
indices_pa_table = None
return cls(
arrow_table=table,
info=info,
split=split,
indices_table=indices_pa_table,
)
@classmethod
def from_buffer(
cls,
buffer: pa.Buffer,
info: Optional[DatasetInfo] = None,
split: Optional[NamedSplit] = None,
indices_buffer: Optional[pa.Buffer] = None,
) -> "Dataset":
"""Instantiate a Dataset backed by an Arrow buffer.
Args:
buffer (`pyarrow.Buffer`):
Arrow buffer.
info (`DatasetInfo`, *optional*):
Dataset information, like description, citation, etc.
split (`NamedSplit`, *optional*):
Name of the dataset split.
indices_buffer (`pyarrow.Buffer`, *optional*):
Indices Arrow buffer.
Returns:
[`Dataset`]
"""
table = InMemoryTable.from_buffer(buffer)
if indices_buffer is not None:
indices_table = InMemoryTable.from_buffer(buffer)
else:
indices_table = None
return cls(table, info=info, split=split, indices_table=indices_table)
@classmethod
def from_pandas(
cls,
df: pd.DataFrame,
features: Optional[Features] = None,
info: Optional[DatasetInfo] = None,
split: Optional[NamedSplit] = None,
preserve_index: Optional[bool] = None,
) -> "Dataset":
"""
Convert `pandas.DataFrame` to a `pyarrow.Table` to create a [`Dataset`].
The column types in the resulting Arrow Table are inferred from the dtypes of the `pandas.Series` in the
DataFrame. In the case of non-object Series, the NumPy dtype is translated to its Arrow equivalent. In the
case of `object`, we need to guess the datatype by looking at the Python objects in this Series.
Be aware that Series of the `object` dtype don't carry enough information to always lead to a meaningful Arrow
type. In the case that we cannot infer a type, e.g. because the DataFrame is of length 0 or the Series only
contains `None/nan` objects, the type is set to `null`. This behavior can be avoided by constructing explicit
features and passing it to this function.
Args:
df (`pandas.DataFrame`):
Dataframe that contains the dataset.
features ([`Features`], *optional*):
Dataset features.
info (`DatasetInfo`, *optional*):
Dataset information, like description, citation, etc.
split (`NamedSplit`, *optional*):
Name of the dataset split.
preserve_index (`bool`, *optional*):
Whether to store the index as an additional column in the resulting Dataset.
The default of `None` will store the index as a column, except for `RangeIndex` which is stored as metadata only.
Use `preserve_index=True` to force it to be stored as a column.
Returns:
[`Dataset`]
Example:
```py
>>> ds = Dataset.from_pandas(df)
```
"""
if info is not None and features is not None and info.features != features:
raise ValueError(
f"Features specified in `features` and `info.features` can't be different:\n{features}\n{info.features}"
)
features = features if features is not None else info.features if info is not None else None
if info is None:
info = DatasetInfo()
info.features = features
table = InMemoryTable.from_pandas(
df=df,
preserve_index=preserve_index,
)
if features is not None:
# more expensive cast than InMemoryTable.from_pandas(..., schema=features.arrow_schema)
# needed to support the str to Audio conversion for instance
table = table.cast(features.arrow_schema)
return cls(table, info=info, split=split)
@classmethod
def from_dict(
cls,
mapping: dict,
features: Optional[Features] = None,
info: Optional[DatasetInfo] = None,
split: Optional[NamedSplit] = None,
) -> "Dataset":
"""
Convert `dict` to a `pyarrow.Table` to create a [`Dataset`].
Args:
mapping (`Mapping`):
Mapping of strings to Arrays or Python lists.
features ([`Features`], *optional*):
Dataset features.
info (`DatasetInfo`, *optional*):
Dataset information, like description, citation, etc.
split (`NamedSplit`, *optional*):
Name of the dataset split.
Returns:
[`Dataset`]
"""
if info is not None and features is not None and info.features != features:
raise ValueError(
f"Features specified in `features` and `info.features` can't be different:\n{features}\n{info.features}"
)
features = features if features is not None else info.features if info is not None else None
arrow_typed_mapping = {}
for col, data in mapping.items():
if isinstance(data, (pa.Array, pa.ChunkedArray)):
data = cast_array_to_feature(data, features[col]) if features is not None else data
else:
data = OptimizedTypedSequence(
features.encode_column(data, col) if features is not None else data,
type=features[col] if features is not None else None,
col=col,
)
arrow_typed_mapping[col] = data
mapping = arrow_typed_mapping
pa_table = InMemoryTable.from_pydict(mapping=mapping)
if info is None:
info = DatasetInfo()
info.features = features
if info.features is None:
info.features = Features(
{
col: generate_from_arrow_type(data.type)
if isinstance(data, (pa.Array, pa.ChunkedArray))
else data.get_inferred_type()
for col, data in mapping.items()
}
)
return cls(pa_table, info=info, split=split)
@classmethod
def from_list(
cls,
mapping: List[dict],
features: Optional[Features] = None,
info: Optional[DatasetInfo] = None,
split: Optional[NamedSplit] = None,
) -> "Dataset":
"""
Convert a list of dicts to a `pyarrow.Table` to create a [`Dataset`]`.
Note that the keys of the first entry will be used to determine the dataset columns,
regardless of what is passed to features.
Args:
mapping (`List[dict]`): A list of mappings of strings to row values.
features (`Features`, optional): Dataset features.
info (`DatasetInfo`, optional): Dataset information, like description, citation, etc.
split (`NamedSplit`, optional): Name of the dataset split.
Returns:
[`Dataset`]
"""
# for simplicity and consistency wrt OptimizedTypedSequence we do not use InMemoryTable.from_pylist here
mapping = {k: [r.get(k) for r in mapping] for k in mapping[0]} if mapping else {}
return cls.from_dict(mapping, features, info, split)
@staticmethod
def from_csv(
path_or_paths: Union[PathLike, List[PathLike]],
split: Optional[NamedSplit] = None,
features: Optional[Features] = None,
cache_dir: str = None,
keep_in_memory: bool = False,
num_proc: Optional[int] = None,
**kwargs,
):
"""Create Dataset from CSV file(s).
Args:
path_or_paths (`path-like` or list of `path-like`):
Path(s) of the CSV file(s).
split ([`NamedSplit`], *optional*):
Split name to be assigned to the dataset.
features ([`Features`], *optional*):
Dataset features.
cache_dir (`str`, *optional*, defaults to `"~/.cache/huggingface/datasets"`):
Directory to cache data.
keep_in_memory (`bool`, defaults to `False`):
Whether to copy the data in-memory.
num_proc (`int`, *optional*, defaults to `None`):
Number of processes when downloading and generating the dataset locally.
This is helpful if the dataset is made of multiple files. Multiprocessing is disabled by default.
<Added version="2.8.0"/>
**kwargs (additional keyword arguments):
Keyword arguments to be passed to [`pandas.read_csv`].
Returns:
[`Dataset`]
Example:
```py
>>> ds = Dataset.from_csv('path/to/dataset.csv')
```
"""
# Dynamic import to avoid circular dependency
from .io.csv import CsvDatasetReader
return CsvDatasetReader(
path_or_paths,
split=split,
features=features,
cache_dir=cache_dir,
keep_in_memory=keep_in_memory,
num_proc=num_proc,
**kwargs,
).read()
@staticmethod
def from_generator(
generator: Callable,
features: Optional[Features] = None,
cache_dir: str = None,
keep_in_memory: bool = False,
gen_kwargs: Optional[dict] = None,
num_proc: Optional[int] = None,
**kwargs,
):
"""Create a Dataset from a generator.
Args:
generator (:`Callable`):
A generator function that `yields` examples.
features ([`Features`], *optional*):
Dataset features.
cache_dir (`str`, *optional*, defaults to `"~/.cache/huggingface/datasets"`):
Directory to cache data.
keep_in_memory (`bool`, defaults to `False`):
Whether to copy the data in-memory.
gen_kwargs(`dict`, *optional*):
Keyword arguments to be passed to the `generator` callable.
You can define a sharded dataset by passing the list of shards in `gen_kwargs` and setting `num_proc` greater than 1.
num_proc (`int`, *optional*, defaults to `None`):
Number of processes when downloading and generating the dataset locally.
This is helpful if the dataset is made of multiple files. Multiprocessing is disabled by default.
If `num_proc` is greater than one, then all list values in `gen_kwargs` must be the same length. These values will be split between calls to the generator. The number of shards will be the minimum of the shortest list in `gen_kwargs` and `num_proc`.
<Added version="2.7.0"/>
**kwargs (additional keyword arguments):
Keyword arguments to be passed to :[`GeneratorConfig`].
Returns:
[`Dataset`]
Example:
```py
>>> def gen():
... yield {"text": "Good", "label": 0}
... yield {"text": "Bad", "label": 1}
...
>>> ds = Dataset.from_generator(gen)
```
```py
>>> def gen(shards):
... for shard in shards:
... with open(shard) as f:
... for line in f:
... yield {"line": line}
...
>>> shards = [f"data{i}.txt" for i in range(32)]
>>> ds = Dataset.from_generator(gen, gen_kwargs={"shards": shards})
```
"""
from .io.generator import GeneratorDatasetInputStream
return GeneratorDatasetInputStream(
generator=generator,
features=features,
cache_dir=cache_dir,
keep_in_memory=keep_in_memory,
gen_kwargs=gen_kwargs,
num_proc=num_proc,
**kwargs,
).read()
@staticmethod
def from_json(
path_or_paths: Union[PathLike, List[PathLike]],
split: Optional[NamedSplit] = None,
features: Optional[Features] = None,
cache_dir: str = None,
keep_in_memory: bool = False,
field: Optional[str] = None,
num_proc: Optional[int] = None,
**kwargs,
):
"""Create Dataset from JSON or JSON Lines file(s).
Args:
path_or_paths (`path-like` or list of `path-like`):
Path(s) of the JSON or JSON Lines file(s).
split ([`NamedSplit`], *optional*):
Split name to be assigned to the dataset.
features ([`Features`], *optional*):
Dataset features.
cache_dir (`str`, *optional*, defaults to `"~/.cache/huggingface/datasets"`):
Directory to cache data.
keep_in_memory (`bool`, defaults to `False`):
Whether to copy the data in-memory.
field (`str`, *optional*):
Field name of the JSON file where the dataset is contained in.
num_proc (`int`, *optional* defaults to `None`):
Number of processes when downloading and generating the dataset locally.
This is helpful if the dataset is made of multiple files. Multiprocessing is disabled by default.
<Added version="2.8.0"/>
**kwargs (additional keyword arguments):
Keyword arguments to be passed to [`JsonConfig`].
Returns:
[`Dataset`]
Example:
```py
>>> ds = Dataset.from_json('path/to/dataset.json')
```
"""
# Dynamic import to avoid circular dependency
from .io.json import JsonDatasetReader
return JsonDatasetReader(
path_or_paths,
split=split,
features=features,
cache_dir=cache_dir,
keep_in_memory=keep_in_memory,
field=field,
num_proc=num_proc,
**kwargs,
).read()
@staticmethod
def from_parquet(
path_or_paths: Union[PathLike, List[PathLike]],
split: Optional[NamedSplit] = None,
features: Optional[Features] = None,
cache_dir: str = None,
keep_in_memory: bool = False,
columns: Optional[List[str]] = None,
num_proc: Optional[int] = None,
**kwargs,
):
"""Create Dataset from Parquet file(s).
Args:
path_or_paths (`path-like` or list of `path-like`):
Path(s) of the Parquet file(s).
split (`NamedSplit`, *optional*):
Split name to be assigned to the dataset.
features (`Features`, *optional*):
Dataset features.
cache_dir (`str`, *optional*, defaults to `"~/.cache/huggingface/datasets"`):
Directory to cache data.
keep_in_memory (`bool`, defaults to `False`):
Whether to copy the data in-memory.
columns (`List[str]`, *optional*):
If not `None`, only these columns will be read from the file.
A column name may be a prefix of a nested field, e.g. 'a' will select
'a.b', 'a.c', and 'a.d.e'.
num_proc (`int`, *optional*, defaults to `None`):
Number of processes when downloading and generating the dataset locally.
This is helpful if the dataset is made of multiple files. Multiprocessing is disabled by default.
<Added version="2.8.0"/>
**kwargs (additional keyword arguments):
Keyword arguments to be passed to [`ParquetConfig`].
Returns:
[`Dataset`]
Example:
```py
>>> ds = Dataset.from_parquet('path/to/dataset.parquet')
```
"""
# Dynamic import to avoid circular dependency
from .io.parquet import ParquetDatasetReader
return ParquetDatasetReader(
path_or_paths,
split=split,
features=features,
cache_dir=cache_dir,
keep_in_memory=keep_in_memory,
columns=columns,
num_proc=num_proc,
**kwargs,
).read()
@staticmethod
def from_text(
path_or_paths: Union[PathLike, List[PathLike]],
split: Optional[NamedSplit] = None,
features: Optional[Features] = None,
cache_dir: str = None,
keep_in_memory: bool = False,
num_proc: Optional[int] = None,
**kwargs,
):
"""Create Dataset from text file(s).
Args:
path_or_paths (`path-like` or list of `path-like`):
Path(s) of the text file(s).
split (`NamedSplit`, *optional*):
Split name to be assigned to the dataset.
features (`Features`, *optional*):
Dataset features.
cache_dir (`str`, *optional*, defaults to `"~/.cache/huggingface/datasets"`):
Directory to cache data.
keep_in_memory (`bool`, defaults to `False`):
Whether to copy the data in-memory.
num_proc (`int`, *optional*, defaults to `None`):
Number of processes when downloading and generating the dataset locally.
This is helpful if the dataset is made of multiple files. Multiprocessing is disabled by default.
<Added version="2.8.0"/>
**kwargs (additional keyword arguments):
Keyword arguments to be passed to [`TextConfig`].
Returns:
[`Dataset`]
Example:
```py
>>> ds = Dataset.from_text('path/to/dataset.txt')
```
"""
# Dynamic import to avoid circular dependency
from .io.text import TextDatasetReader
return TextDatasetReader(
path_or_paths,
split=split,
features=features,
cache_dir=cache_dir,
keep_in_memory=keep_in_memory,
num_proc=num_proc,
**kwargs,
).read()
@staticmethod
def from_spark(
df: "pyspark.sql.DataFrame",
split: Optional[NamedSplit] = None,
features: Optional[Features] = None,
keep_in_memory: bool = False,
cache_dir: str = None,
working_dir: str = None,
load_from_cache_file: bool = True,
**kwargs,
):
"""Create a Dataset from Spark DataFrame. Dataset downloading is distributed over Spark workers.
Args:
df (`pyspark.sql.DataFrame`):
The DataFrame containing the desired data.
split (`NamedSplit`, *optional*):
Split name to be assigned to the dataset.
features (`Features`, *optional*):
Dataset features.
cache_dir (`str`, *optional*, defaults to `"~/.cache/huggingface/datasets"`):
Directory to cache data. When using a multi-node Spark cluster, the cache_dir must be accessible to both
workers and the driver.
keep_in_memory (`bool`):
Whether to copy the data in-memory.
working_dir (`str`, *optional*)
Intermediate directory for each Spark worker to write data to before moving it to `cache_dir`. Setting
a non-NFS intermediate directory may improve performance.
load_from_cache_file (`bool`):
Whether to load the dataset from the cache if possible.
Returns:
[`Dataset`]
Example:
```py
>>> df = spark.createDataFrame(
>>> data=[[1, "Elia"], [2, "Teo"], [3, "Fang"]],
>>> columns=["id", "name"],
>>> )
>>> ds = Dataset.from_spark(df)
```
"""
# Dynamic import to avoid circular dependency
from .io.spark import SparkDatasetReader
if sys.platform == "win32":
raise EnvironmentError("Dataset.from_spark is not currently supported on Windows")
return SparkDatasetReader(
df,
split=split,
features=features,
streaming=False,
cache_dir=cache_dir,
keep_in_memory=keep_in_memory,
working_dir=working_dir,
load_from_cache_file=load_from_cache_file,
**kwargs,
).read()
@staticmethod
def from_sql(
sql: Union[str, "sqlalchemy.sql.Selectable"],
con: Union[str, "sqlalchemy.engine.Connection", "sqlalchemy.engine.Engine", "sqlite3.Connection"],
features: Optional[Features] = None,
cache_dir: str = None,
keep_in_memory: bool = False,
**kwargs,
):
"""Create Dataset from SQL query or database table.
Args:
sql (`str` or `sqlalchemy.sql.Selectable`):
SQL query to be executed or a table name.
con (`str` or `sqlite3.Connection` or `sqlalchemy.engine.Connection` or `sqlalchemy.engine.Connection`):
A [URI string](https://docs.sqlalchemy.org/en/13/core/engines.html#database-urls) used to instantiate a database connection or a SQLite3/SQLAlchemy connection object.
features ([`Features`], *optional*):
Dataset features.
cache_dir (`str`, *optional*, defaults to `"~/.cache/huggingface/datasets"`):
Directory to cache data.
keep_in_memory (`bool`, defaults to `False`):
Whether to copy the data in-memory.
**kwargs (additional keyword arguments):
Keyword arguments to be passed to [`SqlConfig`].
Returns:
[`Dataset`]
Example:
```py
>>> # Fetch a database table
>>> ds = Dataset.from_sql("test_data", "postgres:///db_name")
>>> # Execute a SQL query on the table
>>> ds = Dataset.from_sql("SELECT sentence FROM test_data", "postgres:///db_name")
>>> # Use a Selectable object to specify the query
>>> from sqlalchemy import select, text
>>> stmt = select([text("sentence")]).select_from(text("test_data"))
>>> ds = Dataset.from_sql(stmt, "postgres:///db_name")
```
<Tip>
The returned dataset can only be cached if `con` is specified as URI string.
</Tip>
"""
from .io.sql import SqlDatasetReader
return SqlDatasetReader(
sql,
con,
features=features,
cache_dir=cache_dir,
keep_in_memory=keep_in_memory,
**kwargs,
).read()
def __del__(self):
if hasattr(self, "_data"):
del self._data
if hasattr(self, "_indices"):
del self._indices
def __enter__(self):
return self
def __exit__(self, exc_type, exc_val, exc_tb):
# Here `del` is used to del the pyarrow tables. This properly closes the files used for memory mapped tables
self.__del__()
def save_to_disk(
self,
dataset_path: PathLike,
fs="deprecated",
max_shard_size: Optional[Union[str, int]] = None,
num_shards: Optional[int] = None,
num_proc: Optional[int] = None,
storage_options: Optional[dict] = None,
):
"""
Saves a dataset to a dataset directory, or in a filesystem using any implementation of `fsspec.spec.AbstractFileSystem`.
For [`Image`] and [`Audio`] data:
All the Image() and Audio() data are stored in the arrow files.
If you want to store paths or urls, please use the Value("string") type.
Args:
dataset_path (`str`):
Path (e.g. `dataset/train`) or remote URI (e.g. `s3://my-bucket/dataset/train`)
of the dataset directory where the dataset will be saved to.
fs (`fsspec.spec.AbstractFileSystem`, *optional*):
Instance of the remote filesystem where the dataset will be saved to.
<Deprecated version="2.8.0">
`fs` was deprecated in version 2.8.0 and will be removed in 3.0.0.
Please use `storage_options` instead, e.g. `storage_options=fs.storage_options`
</Deprecated>
max_shard_size (`int` or `str`, *optional*, defaults to `"500MB"`):
The maximum size of the dataset shards to be uploaded to the hub. If expressed as a string, needs to be digits followed by a unit
(like `"50MB"`).
num_shards (`int`, *optional*):
Number of shards to write. By default the number of shards depends on `max_shard_size` and `num_proc`.
<Added version="2.8.0"/>
num_proc (`int`, *optional*):
Number of processes when downloading and generating the dataset locally.
Multiprocessing is disabled by default.
<Added version="2.8.0"/>
storage_options (`dict`, *optional*):
Key/value pairs to be passed on to the file-system backend, if any.
<Added version="2.8.0"/>
Example:
```py
>>> ds.save_to_disk("path/to/dataset/directory")
>>> ds.save_to_disk("path/to/dataset/directory", max_shard_size="1GB")
>>> ds.save_to_disk("path/to/dataset/directory", num_shards=1024)
```
"""
if max_shard_size is not None and num_shards is not None:
raise ValueError(
"Failed to push_to_hub: please specify either max_shard_size or num_shards, but not both."
)
if fs != "deprecated":
warnings.warn(
"'fs' was deprecated in favor of 'storage_options' in version 2.8.0 and will be removed in 3.0.0.\n"
"You can remove this warning by passing 'storage_options=fs.storage_options' instead.",
FutureWarning,
)
storage_options = fs.storage_options
if self.list_indexes():
raise ValueError("please remove all the indexes using `dataset.drop_index` before saving a dataset")
if num_shards is None:
dataset_nbytes = self._estimate_nbytes()
max_shard_size = convert_file_size_to_int(max_shard_size or config.MAX_SHARD_SIZE)
num_shards = int(dataset_nbytes / max_shard_size) + 1
num_shards = max(num_shards, num_proc or 1)
num_proc = num_proc if num_proc is not None else 1
num_shards = num_shards if num_shards is not None else num_proc
fs: fsspec.AbstractFileSystem
fs, _, _ = fsspec.get_fs_token_paths(dataset_path, storage_options=storage_options)
if not is_remote_filesystem(fs):
parent_cache_files_paths = {
Path(cache_filename["filename"]).resolve().parent for cache_filename in self.cache_files
}
# Check that the dataset doesn't overwrite iself. It can cause a permission error on Windows and a segfault on linux.
if Path(dataset_path).expanduser().resolve() in parent_cache_files_paths:
raise PermissionError(
f"Tried to overwrite {Path(dataset_path).expanduser().resolve()} but a dataset can't overwrite itself."
)
fs.makedirs(dataset_path, exist_ok=True)
# Get json serializable state
state = {
key: self.__dict__[key]
for key in [
"_fingerprint",
"_format_columns",
"_format_kwargs",
"_format_type",
"_output_all_columns",
]
}
state["_split"] = str(self.split) if self.split is not None else self.split
state["_data_files"] = [
{"filename": f"data-{shard_idx:05d}-of-{num_shards:05d}.arrow"} for shard_idx in range(num_shards)
]
for k in state["_format_kwargs"].keys():
try:
json.dumps(state["_format_kwargs"][k])
except TypeError as e:
raise TypeError(
str(e) + f"\nThe format kwargs must be JSON serializable, but key '{k}' isn't."
) from None
# Get json serializable dataset info
dataset_info = asdict(self._info)
shards_done = 0
pbar = hf_tqdm(
unit=" examples",
total=len(self),
desc=f"Saving the dataset ({shards_done}/{num_shards} shards)",
)
kwargs_per_job = (
{
"job_id": shard_idx,
"shard": self.shard(num_shards=num_shards, index=shard_idx, contiguous=True),
"fpath": posixpath.join(dataset_path, f"data-{shard_idx:05d}-of-{num_shards:05d}.arrow"),
"storage_options": storage_options,
}
for shard_idx in range(num_shards)
)
shard_lengths = [None] * num_shards
shard_sizes = [None] * num_shards
if num_proc > 1:
with Pool(num_proc) as pool:
with pbar:
for job_id, done, content in iflatmap_unordered(
pool, Dataset._save_to_disk_single, kwargs_iterable=kwargs_per_job
):
if done:
shards_done += 1
pbar.set_description(f"Saving the dataset ({shards_done}/{num_shards} shards)")
logger.debug(f"Finished writing shard number {job_id} of {num_shards}.")
shard_lengths[job_id], shard_sizes[job_id] = content
else:
pbar.update(content)
else:
with pbar:
for kwargs in kwargs_per_job:
for job_id, done, content in Dataset._save_to_disk_single(**kwargs):
if done:
shards_done += 1
pbar.set_description(f"Saving the dataset ({shards_done}/{num_shards} shards)")
logger.debug(f"Finished writing shard number {job_id} of {num_shards}.")
shard_lengths[job_id], shard_sizes[job_id] = content
else:
pbar.update(content)
with fs.open(
posixpath.join(dataset_path, config.DATASET_STATE_JSON_FILENAME), "w", encoding="utf-8"
) as state_file:
json.dump(state, state_file, indent=2, sort_keys=True)
with fs.open(
posixpath.join(dataset_path, config.DATASET_INFO_FILENAME), "w", encoding="utf-8"
) as dataset_info_file:
# Sort only the first level of keys, or we might shuffle fields of nested features if we use sort_keys=True
sorted_keys_dataset_info = {key: dataset_info[key] for key in sorted(dataset_info)}
json.dump(sorted_keys_dataset_info, dataset_info_file, indent=2)
@staticmethod
def _save_to_disk_single(job_id: int, shard: "Dataset", fpath: str, storage_options: Optional[dict]):
batch_size = config.DEFAULT_MAX_BATCH_SIZE
num_examples_progress_update = 0
writer = ArrowWriter(
features=shard.features,
path=fpath,
storage_options=storage_options,
embed_local_files=True,
)
try:
_time = time.time()
for pa_table in shard.with_format("arrow").iter(batch_size):
writer.write_table(pa_table)
num_examples_progress_update += len(pa_table)
if time.time() > _time + config.PBAR_REFRESH_TIME_INTERVAL:
_time = time.time()
yield job_id, False, num_examples_progress_update
num_examples_progress_update = 0
finally:
yield job_id, False, num_examples_progress_update
num_examples, num_bytes = writer.finalize()
writer.close()
yield job_id, True, (num_examples, num_bytes)
@staticmethod
def _build_local_temp_path(uri_or_path: str) -> Path:
"""
Builds and returns a Path concatenating a local temporary dir with the dir path (or absolute/relative
path extracted from the uri) passed.
Args:
uri_or_path (`str`): Path (e.g. `"dataset/train"`) or remote URI (e.g.
`"s3://my-bucket/dataset/train"`) to concatenate.
Returns:
:class:`Path`: the concatenated path (temp dir + path)
"""
src_dataset_path = Path(uri_or_path)
tmp_dir = get_temporary_cache_files_directory()
return Path(tmp_dir, src_dataset_path.relative_to(src_dataset_path.anchor))
@staticmethod
def load_from_disk(
dataset_path: str,
fs="deprecated",
keep_in_memory: Optional[bool] = None,
storage_options: Optional[dict] = None,
) -> "Dataset":
"""
Loads a dataset that was previously saved using [`save_to_disk`] from a dataset directory, or from a
filesystem using any implementation of `fsspec.spec.AbstractFileSystem`.
Args:
dataset_path (`str`):
Path (e.g. `"dataset/train"`) or remote URI (e.g. `"s3//my-bucket/dataset/train"`)
of the dataset directory where the dataset will be loaded from.
fs (`fsspec.spec.AbstractFileSystem`, *optional*):
Instance of the remote filesystem where the dataset will be saved to.
<Deprecated version="2.8.0">
`fs` was deprecated in version 2.8.0 and will be removed in 3.0.0.
Please use `storage_options` instead, e.g. `storage_options=fs.storage_options`
</Deprecated>
keep_in_memory (`bool`, defaults to `None`):
Whether to copy the dataset in-memory. If `None`, the
dataset will not be copied in-memory unless explicitly enabled by setting
`datasets.config.IN_MEMORY_MAX_SIZE` to nonzero. See more details in the
[improve performance](../cache#improve-performance) section.
storage_options (`dict`, *optional*):
Key/value pairs to be passed on to the file-system backend, if any.
<Added version="2.8.0"/>
Returns:
[`Dataset`] or [`DatasetDict`]:
- If `dataset_path` is a path of a dataset directory, the dataset requested.
- If `dataset_path` is a path of a dataset dict directory, a `datasets.DatasetDict` with each split.
Example:
```py
>>> ds = load_from_disk("path/to/dataset/directory")
```
"""
if fs != "deprecated":
warnings.warn(
"'fs' was deprecated in favor of 'storage_options' in version 2.8.0 and will be removed in 3.0.0.\n"
"You can remove this warning by passing 'storage_options=fs.storage_options' instead.",
FutureWarning,
)
storage_options = fs.storage_options
fs: fsspec.AbstractFileSystem
fs, _, [dataset_path] = fsspec.get_fs_token_paths(dataset_path, storage_options=storage_options)
dest_dataset_path = dataset_path
dataset_dict_json_path = posixpath.join(dest_dataset_path, config.DATASETDICT_JSON_FILENAME)
dataset_state_json_path = posixpath.join(dest_dataset_path, config.DATASET_STATE_JSON_FILENAME)
dataset_info_path = posixpath.join(dest_dataset_path, config.DATASET_INFO_FILENAME)
dataset_dict_is_file = fs.isfile(dataset_dict_json_path)
dataset_info_is_file = fs.isfile(dataset_info_path)
dataset_state_is_file = fs.isfile(dataset_state_json_path)
if not dataset_info_is_file and not dataset_state_is_file:
if dataset_dict_is_file:
raise FileNotFoundError(
f"No such files: '{dataset_info_path}', nor '{dataset_state_json_path}' found. Expected to load a `Dataset` object, but got a `DatasetDict`. Please use either `datasets.load_from_disk` or `DatasetDict.load_from_disk` instead."
)
raise FileNotFoundError(
f"No such files: '{dataset_info_path}', nor '{dataset_state_json_path}' found. Expected to load a `Dataset` object but provided path is not a `Dataset`."
)
if not dataset_info_is_file:
if dataset_dict_is_file:
raise FileNotFoundError(
f"No such file: '{dataset_info_path}' found. Expected to load a `Dataset` object, but got a `DatasetDict`. Please use either `datasets.load_from_disk` or `DatasetDict.load_from_disk` instead."
)
raise FileNotFoundError(
f"No such file: '{dataset_info_path}'. Expected to load a `Dataset` object but provided path is not a `Dataset`."
)
if not dataset_state_is_file:
if dataset_dict_is_file:
raise FileNotFoundError(
f"No such file: '{dataset_state_json_path}' found. Expected to load a `Dataset` object, but got a `DatasetDict`. Please use either `datasets.load_from_disk` or `DatasetDict.load_from_disk` instead."
)
raise FileNotFoundError(
f"No such file: '{dataset_state_json_path}'. Expected to load a `Dataset` object but provided path is not a `Dataset`."
)
# copies file from filesystem if it is remote filesystem to local filesystem and modifies dataset_path to temp directory containing local copies
if is_remote_filesystem(fs):
src_dataset_path = dest_dataset_path
dest_dataset_path = Dataset._build_local_temp_path(src_dataset_path)
fs.download(src_dataset_path, dest_dataset_path.as_posix(), recursive=True)
dataset_state_json_path = posixpath.join(dest_dataset_path, config.DATASET_STATE_JSON_FILENAME)
dataset_info_path = posixpath.join(dest_dataset_path, config.DATASET_INFO_FILENAME)
with open(dataset_state_json_path, encoding="utf-8") as state_file:
state = json.load(state_file)
with open(dataset_info_path, encoding="utf-8") as dataset_info_file:
dataset_info = DatasetInfo.from_dict(json.load(dataset_info_file))
dataset_size = estimate_dataset_size(
Path(dest_dataset_path, data_file["filename"]) for data_file in state["_data_files"]
)
keep_in_memory = keep_in_memory if keep_in_memory is not None else is_small_dataset(dataset_size)
table_cls = InMemoryTable if keep_in_memory else MemoryMappedTable
arrow_table = concat_tables(
table_cls.from_file(posixpath.join(dest_dataset_path, data_file["filename"]))
for data_file in state["_data_files"]
)
split = state["_split"]
split = Split(split) if split is not None else split
dataset = Dataset(
arrow_table=arrow_table,
info=dataset_info,
split=split,
fingerprint=state["_fingerprint"],
)
format = {
"type": state["_format_type"],
"format_kwargs": state["_format_kwargs"],
"columns": state["_format_columns"],
"output_all_columns": state["_output_all_columns"],
}
dataset = dataset.with_format(**format)
return dataset
@property
def data(self) -> Table:
"""The Apache Arrow table backing the dataset.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", split="validation")
>>> ds.data
MemoryMappedTable
text: string
label: int64
----
text: [["compassionately explores the seemingly irreconcilable situation between conservative christian parents and their estranged gay and lesbian children .","the soundtrack alone is worth the price of admission .","rodriguez does a splendid job of racial profiling hollywood style--casting excellent latin actors of all ages--a trend long overdue .","beneath the film's obvious determination to shock at any cost lies considerable skill and determination , backed by sheer nerve .","bielinsky is a filmmaker of impressive talent .","so beautifully acted and directed , it's clear that washington most certainly has a new career ahead of him if he so chooses .","a visual spectacle full of stunning images and effects .","a gentle and engrossing character study .","it's enough to watch huppert scheming , with her small , intelligent eyes as steady as any noir villain , and to enjoy the perfectly pitched web of tension that chabrol spins .","an engrossing portrait of uncompromising artists trying to create something original against the backdrop of a corporate music industry that only seems to care about the bottom line .",...,"ultimately , jane learns her place as a girl , softens up and loses some of the intensity that made her an interesting character to begin with .","ah-nuld's action hero days might be over .","it's clear why deuces wild , which was shot two years ago , has been gathering dust on mgm's shelf .","feels like nothing quite so much as a middle-aged moviemaker's attempt to surround himself with beautiful , half-naked women .","when the precise nature of matthew's predicament finally comes into sharp focus , the revelation fails to justify the build-up .","this picture is murder by numbers , and as easy to be bored by as your abc's , despite a few whopping shootouts .","hilarious musical comedy though stymied by accents thick as mud .","if you are into splatter movies , then you will probably have a reasonably good time with the salton sea .","a dull , simple-minded and stereotypical tale of drugs , death and mind-numbing indifference on the inner-city streets .","the feature-length stretch . . . strains the show's concept ."]]
label: [[1,1,1,1,1,1,1,1,1,1,...,0,0,0,0,0,0,0,0,0,0]]
```
"""
return self._data
@property
def cache_files(self) -> List[dict]:
"""The cache files containing the Apache Arrow table backing the dataset.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", split="validation")
>>> ds.cache_files
[{'filename': '/root/.cache/huggingface/datasets/rotten_tomatoes_movie_review/default/1.0.0/40d411e45a6ce3484deed7cc15b82a53dad9a72aafd9f86f8f227134bec5ca46/rotten_tomatoes_movie_review-validation.arrow'}]
```
"""
cache_files = list_table_cache_files(self._data)
if self._indices is not None:
cache_files += list_table_cache_files(self._indices)
return [{"filename": cache_filename} for cache_filename in cache_files]
@property
def num_columns(self) -> int:
"""Number of columns in the dataset.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", split="validation")
>>> ds.num_columns
2
```
"""
return self._data.num_columns
@property
def num_rows(self) -> int:
"""Number of rows in the dataset (same as [`Dataset.__len__`]).
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", split="validation")
>>> ds.num_rows
1066
```
"""
if self._indices is not None:
return self._indices.num_rows
return self._data.num_rows
@property
def column_names(self) -> List[str]:
"""Names of the columns in the dataset.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", split="validation")
>>> ds.column_names
['text', 'label']
```
"""
return self._data.column_names
@property
def shape(self) -> Tuple[int, int]:
"""Shape of the dataset (number of columns, number of rows).
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", split="validation")
>>> ds.shape
(1066, 2)
```
"""
if self._indices is not None:
return (self._indices.num_rows, self._data.num_columns)
return self._data.shape
def unique(self, column: str) -> List:
"""Return a list of the unique elements in a column.
This is implemented in the low-level backend and as such, very fast.
Args:
column (`str`):
Column name (list all the column names with [`~datasets.Dataset.column_names`]).
Returns:
`list`: List of unique elements in the given column.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", split="validation")
>>> ds.unique('label')
[1, 0]
```
"""
if column not in self._data.column_names:
raise ValueError(f"Column ({column}) not in table columns ({self._data.column_names}).")
if self._indices is not None and self._indices.num_rows != self._data.num_rows:
dataset = self.flatten_indices()
else:
dataset = self
return dataset._data.column(column).unique().to_pylist()
def class_encode_column(self, column: str, include_nulls: bool = False) -> "Dataset":
"""Casts the given column as [`~datasets.features.ClassLabel`] and updates the table.
Args:
column (`str`):
The name of the column to cast (list all the column names with [`~datasets.Dataset.column_names`])
include_nulls (`bool`, defaults to `False`):
Whether to include null values in the class labels. If `True`, the null values will be encoded as the `"None"` class label.
<Added version="1.14.2"/>
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("boolq", split="validation")
>>> ds.features
{'answer': Value(dtype='bool', id=None),
'passage': Value(dtype='string', id=None),
'question': Value(dtype='string', id=None)}
>>> ds = ds.class_encode_column('answer')
>>> ds.features
{'answer': ClassLabel(num_classes=2, names=['False', 'True'], id=None),
'passage': Value(dtype='string', id=None),
'question': Value(dtype='string', id=None)}
```
"""
# Sanity checks
if column not in self._data.column_names:
raise ValueError(f"Column ({column}) not in table columns ({self._data.column_names}).")
src_feat = self._info.features[column]
if not isinstance(src_feat, Value):
raise ValueError(
f"Class encoding is only supported for {Value.__name__} column, and column {column} is {type(src_feat).__name__}."
)
if src_feat.dtype != "string" or (include_nulls and None in self.unique(column)):
def stringify_column(batch):
batch[column] = [
str(sample) if include_nulls or sample is not None else None for sample in batch[column]
]
return batch
dset = self.map(
stringify_column,
batched=True,
desc="Stringifying the column",
)
else:
dset = self
# Create the new feature
class_names = sorted(str(sample) for sample in dset.unique(column) if include_nulls or sample is not None)
dst_feat = ClassLabel(names=class_names)
def cast_to_class_labels(batch):
batch[column] = [
dst_feat.str2int(str(sample)) if include_nulls or sample is not None else None
for sample in batch[column]
]
return batch
new_features = dset.features.copy()
new_features[column] = dst_feat
dset = dset.map(
cast_to_class_labels,
batched=True,
features=new_features,
desc="Casting to class labels",
)
return dset
@fingerprint_transform(inplace=False)
def flatten(self, new_fingerprint: Optional[str] = None, max_depth=16) -> "Dataset":
"""Flatten the table.
Each column with a struct type is flattened into one column per struct field.
Other columns are left unchanged.
Args:
new_fingerprint (`str`, *optional*):
The new fingerprint of the dataset after transform.
If `None`, the new fingerprint is computed using a hash of the previous fingerprint, and the transform arguments.
Returns:
[`Dataset`]: A copy of the dataset with flattened columns.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("squad", split="train")
>>> ds.features
{'answers': Sequence(feature={'text': Value(dtype='string', id=None), 'answer_start': Value(dtype='int32', id=None)}, length=-1, id=None),
'context': Value(dtype='string', id=None),
'id': Value(dtype='string', id=None),
'question': Value(dtype='string', id=None),
'title': Value(dtype='string', id=None)}
>>> ds.flatten()
Dataset({
features: ['id', 'title', 'context', 'question', 'answers.text', 'answers.answer_start'],
num_rows: 87599
})
```
"""
dataset = copy.deepcopy(self)
for depth in range(1, max_depth):
if any(isinstance(field.type, pa.StructType) for field in dataset._data.schema):
dataset._data = dataset._data.flatten()
else:
break
dataset.info.features = self._info.features.flatten(max_depth=max_depth)
dataset.info.features = Features({col: dataset.info.features[col] for col in dataset.data.column_names})
dataset._data = update_metadata_with_features(dataset._data, dataset.features)
logger.info(f'Flattened dataset from depth {depth} to depth {1 if depth + 1 < max_depth else "unknown"}.')
dataset._fingerprint = new_fingerprint
return dataset
def cast(
self,
features: Features,
batch_size: Optional[int] = 1000,
keep_in_memory: bool = False,
load_from_cache_file: Optional[bool] = None,
cache_file_name: Optional[str] = None,
writer_batch_size: Optional[int] = 1000,
num_proc: Optional[int] = None,
) -> "Dataset":
"""
Cast the dataset to a new set of features.
Args:
features ([`Features`]):
New features to cast the dataset to.
The name of the fields in the features must match the current column names.
The type of the data must also be convertible from one type to the other.
For non-trivial conversion, e.g. `str` <-> `ClassLabel` you should use [`~datasets.Dataset.map`] to update the Dataset.
batch_size (`int`, defaults to `1000`):
Number of examples per batch provided to cast.
If `batch_size <= 0` or `batch_size == None` then provide the full dataset as a single batch to cast.
keep_in_memory (`bool`, defaults to `False`):
Whether to copy the data in-memory.
load_from_cache_file (`bool`, defaults to `True` if caching is enabled):
If a cache file storing the current computation from `function`
can be identified, use it instead of recomputing.
cache_file_name (`str`, *optional*, defaults to `None`):
Provide the name of a path for the cache file. It is used to store the
results of the computation instead of the automatically generated cache file name.
writer_batch_size (`int`, defaults to `1000`):
Number of rows per write operation for the cache file writer.
This value is a good trade-off between memory usage during the processing, and processing speed.
Higher value makes the processing do fewer lookups, lower value consume less temporary memory while running [`~datasets.Dataset.map`].
num_proc (`int`, *optional*, defaults to `None`):
Number of processes for multiprocessing. By default it doesn't
use multiprocessing.
Returns:
[`Dataset`]: A copy of the dataset with casted features.
Example:
```py
>>> from datasets import load_dataset, ClassLabel, Value
>>> ds = load_dataset("rotten_tomatoes", split="validation")
>>> ds.features
{'label': ClassLabel(num_classes=2, names=['neg', 'pos'], id=None),
'text': Value(dtype='string', id=None)}
>>> new_features = ds.features.copy()
>>> new_features['label'] = ClassLabel(names=['bad', 'good'])
>>> new_features['text'] = Value('large_string')
>>> ds = ds.cast(new_features)
>>> ds.features
{'label': ClassLabel(num_classes=2, names=['bad', 'good'], id=None),
'text': Value(dtype='large_string', id=None)}
```
"""
if sorted(features) != sorted(self._data.column_names):
raise ValueError(
f"The columns in features ({list(features)}) must be identical "
f"as the columns in the dataset: {self._data.column_names}"
)
schema = features.arrow_schema
format = self.format
dataset = self.with_format("arrow")
# capture the PyArrow version here to make the lambda serializable on Windows
dataset = dataset.map(
partial(table_cast, schema=schema),
batched=True,
batch_size=batch_size,
keep_in_memory=keep_in_memory,
load_from_cache_file=load_from_cache_file,
cache_file_name=cache_file_name,
writer_batch_size=writer_batch_size,
num_proc=num_proc,
features=features,
desc="Casting the dataset",
)
dataset = dataset.with_format(**format)
return dataset
@fingerprint_transform(inplace=False)
def cast_column(self, column: str, feature: FeatureType, new_fingerprint: Optional[str] = None) -> "Dataset":
"""Cast column to feature for decoding.
Args:
column (`str`):
Column name.
feature (`FeatureType`):
Target feature.
new_fingerprint (`str`, *optional*):
The new fingerprint of the dataset after transform.
If `None`, the new fingerprint is computed using a hash of the previous fingerprint, and the transform arguments.
Returns:
[`Dataset`]
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", split="validation")
>>> ds.features
{'label': ClassLabel(num_classes=2, names=['neg', 'pos'], id=None),
'text': Value(dtype='string', id=None)}
>>> ds = ds.cast_column('label', ClassLabel(names=['bad', 'good']))
>>> ds.features
{'label': ClassLabel(num_classes=2, names=['bad', 'good'], id=None),
'text': Value(dtype='string', id=None)}
```
"""
if hasattr(feature, "decode_example"):
dataset = copy.deepcopy(self)
dataset._info.features[column] = feature
dataset._fingerprint = new_fingerprint
dataset._data = dataset._data.cast(dataset.features.arrow_schema)
dataset._data = update_metadata_with_features(dataset._data, dataset.features)
return dataset
else:
features = self.features
features[column] = feature
return self.cast(features)
@transmit_tasks
@transmit_format
@fingerprint_transform(inplace=False)
def remove_columns(self, column_names: Union[str, List[str]], new_fingerprint: Optional[str] = None) -> "Dataset":
"""
Remove one or several column(s) in the dataset and the features associated to them.
You can also remove a column using [`~datasets.Dataset.map`] with `remove_columns` but the present method
is in-place (doesn't copy the data to a new dataset) and is thus faster.
Args:
column_names (`Union[str, List[str]]`):
Name of the column(s) to remove.
new_fingerprint (`str`, *optional*):
The new fingerprint of the dataset after transform.
If `None`, the new fingerprint is computed using a hash of the previous fingerprint, and the transform arguments.
Returns:
[`Dataset`]: A copy of the dataset object without the columns to remove.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", split="validation")
>>> ds.remove_columns('label')
Dataset({
features: ['text'],
num_rows: 1066
})
>>> ds.remove_columns(column_names=ds.column_names) # Removing all the columns returns an empty dataset with the `num_rows` property set to 0
Dataset({
features: [],
num_rows: 0
})
```
"""
dataset = copy.deepcopy(self)
if isinstance(column_names, str):
column_names = [column_names]
for column_name in column_names:
if column_name not in dataset._data.column_names:
raise ValueError(
f"Column name {column_name} not in the dataset. "
f"Current columns in the dataset: {dataset._data.column_names}"
)
for column_name in column_names:
del dataset._info.features[column_name]
dataset._data = dataset._data.drop(column_names)
dataset._data = update_metadata_with_features(dataset._data, dataset.features)
dataset._fingerprint = new_fingerprint
return dataset
@transmit_tasks
@fingerprint_transform(inplace=False)
def rename_column(
self, original_column_name: str, new_column_name: str, new_fingerprint: Optional[str] = None
) -> "Dataset":
"""
Rename a column in the dataset, and move the features associated to the original column under the new column
name.
Args:
original_column_name (`str`):
Name of the column to rename.
new_column_name (`str`):
New name for the column.
new_fingerprint (`str`, *optional*):
The new fingerprint of the dataset after transform.
If `None`, the new fingerprint is computed using a hash of the previous fingerprint, and the transform arguments.
Returns:
[`Dataset`]: A copy of the dataset with a renamed column.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", split="validation")
>>> ds.rename_column('label', 'label_new')
Dataset({
features: ['text', 'label_new'],
num_rows: 1066
})
```
"""
dataset = copy.deepcopy(self)
if original_column_name not in dataset._data.column_names:
raise ValueError(
f"Original column name {original_column_name} not in the dataset. "
f"Current columns in the dataset: {dataset._data.column_names}"
)
if new_column_name in dataset._data.column_names:
raise ValueError(
f"New column name {new_column_name} already in the dataset. "
f"Please choose a column name which is not already in the dataset. "
f"Current columns in the dataset: {dataset._data.column_names}"
)
if not new_column_name:
raise ValueError("New column name is empty.")
def rename(columns):
return [new_column_name if col == original_column_name else col for col in columns]
new_column_names = rename(self._data.column_names)
if self._format_columns is not None:
dataset._format_columns = rename(self._format_columns)
dataset._info.features = Features(
{
new_column_name if col == original_column_name else col: feature
for col, feature in self._info.features.items()
}
)
dataset._data = dataset._data.rename_columns(new_column_names)
dataset._data = update_metadata_with_features(dataset._data, dataset.features)
dataset._fingerprint = new_fingerprint
return dataset
@transmit_tasks
@fingerprint_transform(inplace=False)
def rename_columns(self, column_mapping: Dict[str, str], new_fingerprint: Optional[str] = None) -> "Dataset":
"""
Rename several columns in the dataset, and move the features associated to the original columns under
the new column names.
Args:
column_mapping (`Dict[str, str]`):
A mapping of columns to rename to their new names
new_fingerprint (`str`, *optional*):
The new fingerprint of the dataset after transform.
If `None`, the new fingerprint is computed using a hash of the previous fingerprint, and the transform arguments.
Returns:
[`Dataset`]: A copy of the dataset with renamed columns
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", split="validation")
>>> ds.rename_columns({'text': 'text_new', 'label': 'label_new'})
Dataset({
features: ['text_new', 'label_new'],
num_rows: 1066
})
```
"""
dataset = copy.deepcopy(self)
extra_columns = set(column_mapping.keys()) - set(dataset.column_names)
if extra_columns:
raise ValueError(
f"Original column names {extra_columns} not in the dataset. "
f"Current columns in the dataset: {dataset._data.column_names}"
)
number_of_duplicates_in_new_columns = len(column_mapping.values()) - len(set(column_mapping.values()))
if number_of_duplicates_in_new_columns != 0:
raise ValueError(
"New column names must all be different, but this column mapping "
f"has {number_of_duplicates_in_new_columns} duplicates"
)
empty_new_columns = [new_col for new_col in column_mapping.values() if not new_col]
if empty_new_columns:
raise ValueError(f"New column names {empty_new_columns} are empty.")
def rename(columns):
return [column_mapping[col] if col in column_mapping else col for col in columns]
new_column_names = rename(self._data.column_names)
if self._format_columns is not None:
dataset._format_columns = rename(self._format_columns)
dataset._info.features = Features(
{
column_mapping[col] if col in column_mapping else col: feature
for col, feature in (self._info.features or {}).items()
}
)
dataset._data = dataset._data.rename_columns(new_column_names)
dataset._data = update_metadata_with_features(dataset._data, dataset.features)
dataset._fingerprint = new_fingerprint
return dataset
@transmit_tasks
@transmit_format
@fingerprint_transform(inplace=False)
def select_columns(self, column_names: Union[str, List[str]], new_fingerprint: Optional[str] = None) -> "Dataset":
"""Select one or several column(s) in the dataset and the features
associated to them.
Args:
column_names (`Union[str, List[str]]`):
Name of the column(s) to keep.
new_fingerprint (`str`, *optional*):
The new fingerprint of the dataset after transform. If `None`,
the new fingerprint is computed using a hash of the previous
fingerprint, and the transform arguments.
Returns:
[`Dataset`]: A copy of the dataset object which only consists of
selected columns.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", split="validation")
>>> ds.select_columns(['text'])
Dataset({
features: ['text'],
num_rows: 1066
})
```
"""
if isinstance(column_names, str):
column_names = [column_names]
for column_name in column_names:
if column_name not in self._data.column_names:
raise ValueError(
f"Column name {column_name} not in the "
"dataset. Current columns in the dataset: "
f"{self._data.column_names}."
)
dataset = copy.deepcopy(self)
dataset._data = dataset._data.select(column_names)
dataset._info.features = Features({col: self._info.features[col] for col in dataset._data.column_names})
dataset._data = update_metadata_with_features(dataset._data, dataset.features)
dataset._fingerprint = new_fingerprint
return dataset
def __len__(self):
"""Number of rows in the dataset.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", split="validation")
>>> ds.__len__
<bound method Dataset.__len__ of Dataset({
features: ['text', 'label'],
num_rows: 1066
})>
```
"""
return self.num_rows
def __iter__(self):
"""Iterate through the examples.
If a formatting is set with :meth:`Dataset.set_format` rows will be returned with the
selected format.
"""
if self._indices is None:
# Fast iteration
# Benchmark: https://gist.github.com/mariosasko/0248288a2e3a7556873969717c1fe52b (fast_iter_batch)
format_kwargs = self._format_kwargs if self._format_kwargs is not None else {}
formatter = get_formatter(self._format_type, features=self._info.features, **format_kwargs)
batch_size = config.ARROW_READER_BATCH_SIZE_IN_DATASET_ITER
for pa_subtable in table_iter(self.data, batch_size=batch_size):
for i in range(pa_subtable.num_rows):
pa_subtable_ex = pa_subtable.slice(i, 1)
formatted_output = format_table(
pa_subtable_ex,
0,
formatter=formatter,
format_columns=self._format_columns,
output_all_columns=self._output_all_columns,
)
yield formatted_output
else:
for i in range(self.num_rows):
yield self._getitem(
i,
)
def iter(self, batch_size: int, drop_last_batch: bool = False):
"""Iterate through the batches of size `batch_size`.
If a formatting is set with [`~datasets.Dataset.set_format`] rows will be returned with the
selected format.
Args:
batch_size (:obj:`int`): size of each batch to yield.
drop_last_batch (:obj:`bool`, default `False`): Whether a last batch smaller than the batch_size should be
dropped
"""
if self._indices is None:
# Fast iteration
# Benchmark: https://gist.github.com/mariosasko/0248288a2e3a7556873969717c1fe52b (fast_iter_batch)
format_kwargs = self._format_kwargs if self._format_kwargs is not None else {}
formatter = get_formatter(self._format_type, features=self._info.features, **format_kwargs)
for pa_subtable in table_iter(self.data, batch_size=batch_size, drop_last_batch=drop_last_batch):
formatted_batch = format_table(
pa_subtable,
range(pa_subtable.num_rows),
formatter=formatter,
format_columns=self._format_columns,
output_all_columns=self._output_all_columns,
)
yield formatted_batch
else:
num_rows = self.num_rows if not drop_last_batch else self.num_rows // batch_size * batch_size
for i in range(0, num_rows, batch_size):
yield self._getitem(
slice(i, i + batch_size),
)
def __repr__(self):
return f"Dataset({{\n features: {list(self._info.features.keys())},\n num_rows: {self.num_rows}\n}})"
@property
def format(self):
return {
"type": self._format_type,
"format_kwargs": self._format_kwargs,
"columns": self.column_names if self._format_columns is None else self._format_columns,
"output_all_columns": self._output_all_columns,
}
@contextlib.contextmanager
def formatted_as(
self,
type: Optional[str] = None,
columns: Optional[List] = None,
output_all_columns: bool = False,
**format_kwargs,
):
"""To be used in a `with` statement. Set `__getitem__` return format (type and columns).
Args:
type (`str`, *optional*):
Output type selected in `[None, 'numpy', 'torch', 'tensorflow', 'pandas', 'arrow', 'jax']`.
`None` means `__getitem__`` returns python objects (default).
columns (`List[str]`, *optional*):
Columns to format in the output.
`None` means `__getitem__` returns all columns (default).
output_all_columns (`bool`, defaults to `False`):
Keep un-formatted columns as well in the output (as python objects).
**format_kwargs (additional keyword arguments):
Keywords arguments passed to the convert function like `np.array`, `torch.tensor` or `tensorflow.ragged.constant`.
"""
old_format_type = self._format_type
old_format_kwargs = self._format_kwargs
old_format_columns = self._format_columns
old_output_all_columns = self._output_all_columns
try:
self.set_format(type, columns, output_all_columns, **format_kwargs)
yield
finally:
self.set_format(old_format_type, old_format_columns, old_output_all_columns, **old_format_kwargs)
@fingerprint_transform(inplace=True)
def set_format(
self,
type: Optional[str] = None,
columns: Optional[List] = None,
output_all_columns: bool = False,
**format_kwargs,
):
"""Set `__getitem__` return format (type and columns). The data formatting is applied on-the-fly.
The format `type` (for example "numpy") is used to format batches when using `__getitem__`.
It's also possible to use custom transforms for formatting using [`~datasets.Dataset.set_transform`].
Args:
type (`str`, *optional*):
Either output type selected in `[None, 'numpy', 'torch', 'tensorflow', 'pandas', 'arrow', 'jax']`.
`None` means `__getitem__` returns python objects (default).
columns (`List[str]`, *optional*):
Columns to format in the output.
`None` means `__getitem__` returns all columns (default).
output_all_columns (`bool`, defaults to `False`):
Keep un-formatted columns as well in the output (as python objects).
**format_kwargs (additional keyword arguments):
Keywords arguments passed to the convert function like `np.array`, `torch.tensor` or `tensorflow.ragged.constant`.
It is possible to call [`~datasets.Dataset.map`] after calling `set_format`. Since `map` may add new columns, then the list of formatted columns
gets updated. In this case, if you apply `map` on a dataset to add a new column, then this column will be formatted as:
```
new formatted columns = (all columns - previously unformatted columns)
```
Example:
```py
>>> from datasets import load_dataset
>>> from transformers import AutoTokenizer
>>> ds = load_dataset("rotten_tomatoes", split="validation")
>>> tokenizer = AutoTokenizer.from_pretrained("bert-base-cased")
>>> ds = ds.map(lambda x: tokenizer(x['text'], truncation=True, padding=True), batched=True)
>>> ds.set_format(type='numpy', columns=['text', 'label'])
>>> ds.format
{'type': 'numpy',
'format_kwargs': {},
'columns': ['text', 'label'],
'output_all_columns': False}
```
"""
format_kwargs.update(format_kwargs.pop("format_kwargs", {})) # allow to use self.set_format(**self.format)
# Check that the format_type and format_kwargs are valid and make it possible to have a Formatter
type = get_format_type_from_alias(type)
get_formatter(type, features=self._info.features, **format_kwargs)
# Check filter column
if isinstance(columns, str):
columns = [columns]
if isinstance(columns, tuple):
columns = list(columns)
if columns is not None and any(col not in self._data.column_names for col in columns):
raise ValueError(
f"Columns {list(filter(lambda col: col not in self._data.column_names, columns))} not in the dataset. Current columns in the dataset: {self._data.column_names}"
)
if columns is not None:
columns = columns.copy() # Ensures modifications made to the list after this call don't cause bugs
self._format_type = type
self._format_kwargs = format_kwargs
self._format_columns = columns
self._output_all_columns = output_all_columns
logger.debug(
"Set __getitem__(key) output type to %s for %s columns "
" (when key is int or slice) and %s output other (un-formatted) columns.",
"python objects" if type is None else type,
"no" if columns is None else str(columns),
"do" if output_all_columns else "don't",
)
def reset_format(self):
"""Reset `__getitem__` return format to python objects and all columns.
Same as `self.set_format()`
Example:
```py
>>> from datasets import load_dataset
>>> from transformers import AutoTokenizer
>>> ds = load_dataset("rotten_tomatoes", split="validation")
>>> tokenizer = AutoTokenizer.from_pretrained("bert-base-cased")
>>> ds = ds.map(lambda x: tokenizer(x['text'], truncation=True, padding=True), batched=True)
>>> ds.set_format(type='numpy', columns=['input_ids', 'token_type_ids', 'attention_mask', 'label'])
>>> ds.format
{'columns': ['input_ids', 'token_type_ids', 'attention_mask', 'label'],
'format_kwargs': {},
'output_all_columns': False,
'type': 'numpy'}
>>> ds.reset_format()
>>> ds.format
{'columns': ['text', 'label', 'input_ids', 'token_type_ids', 'attention_mask'],
'format_kwargs': {},
'output_all_columns': False,
'type': None}
```
"""
self.set_format()
def set_transform(
self,
transform: Optional[Callable],
columns: Optional[List] = None,
output_all_columns: bool = False,
):
"""Set `__getitem__` return format using this transform. The transform is applied on-the-fly on batches when `__getitem__` is called.
As [`~datasets.Dataset.set_format`], this can be reset using [`~datasets.Dataset.reset_format`].
Args:
transform (`Callable`, *optional*):
User-defined formatting transform, replaces the format defined by [`~datasets.Dataset.set_format`].
A formatting function is a callable that takes a batch (as a `dict`) as input and returns a batch.
This function is applied right before returning the objects in `__getitem__`.
columns (`List[str]`, *optional*):
Columns to format in the output.
If specified, then the input batch of the transform only contains those columns.
output_all_columns (`bool`, defaults to `False`):
Keep un-formatted columns as well in the output (as python objects).
If set to True, then the other un-formatted columns are kept with the output of the transform.
Example:
```py
>>> from datasets import load_dataset
>>> from transformers import AutoTokenizer
>>> ds = load_dataset("rotten_tomatoes", split="validation")
>>> tokenizer = AutoTokenizer.from_pretrained('bert-base-uncased')
>>> def encode(batch):
... return tokenizer(batch['text'], padding=True, truncation=True, return_tensors='pt')
>>> ds.set_transform(encode)
>>> ds[0]
{'attention_mask': tensor([1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1,
1, 1]),
'input_ids': tensor([ 101, 29353, 2135, 15102, 1996, 9428, 20868, 2890, 8663, 6895,
20470, 2571, 3663, 2090, 4603, 3017, 3008, 1998, 2037, 24211,
5637, 1998, 11690, 2336, 1012, 102]),
'token_type_ids': tensor([0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0])}
```
"""
self.set_format("custom", columns=columns, output_all_columns=output_all_columns, transform=transform)
def with_format(
self,
type: Optional[str] = None,
columns: Optional[List] = None,
output_all_columns: bool = False,
**format_kwargs,
):
"""Set `__getitem__` return format (type and columns). The data formatting is applied on-the-fly.
The format `type` (for example "numpy") is used to format batches when using `__getitem__`.
It's also possible to use custom transforms for formatting using [`~datasets.Dataset.with_transform`].
Contrary to [`~datasets.Dataset.set_format`], `with_format` returns a new [`Dataset`] object.
Args:
type (`str`, *optional*):
Either output type selected in `[None, 'numpy', 'torch', 'tensorflow', 'pandas', 'arrow', 'jax']`.
`None` means `__getitem__` returns python objects (default).
columns (`List[str]`, *optional*):
Columns to format in the output.
`None` means `__getitem__` returns all columns (default).
output_all_columns (`bool`, defaults to `False`):
Keep un-formatted columns as well in the output (as python objects).
**format_kwargs (additional keyword arguments):
Keywords arguments passed to the convert function like `np.array`, `torch.tensor` or `tensorflow.ragged.constant`.
Example:
```py
>>> from datasets import load_dataset
>>> from transformers import AutoTokenizer
>>> ds = load_dataset("rotten_tomatoes", split="validation")
>>> tokenizer = AutoTokenizer.from_pretrained("bert-base-cased")
>>> ds = ds.map(lambda x: tokenizer(x['text'], truncation=True, padding=True), batched=True)
>>> ds.format
{'columns': ['text', 'label', 'input_ids', 'token_type_ids', 'attention_mask'],
'format_kwargs': {},
'output_all_columns': False,
'type': None}
>>> ds = ds.with_format(type='tensorflow', columns=['input_ids', 'token_type_ids', 'attention_mask', 'label'])
>>> ds.format
{'columns': ['input_ids', 'token_type_ids', 'attention_mask', 'label'],
'format_kwargs': {},
'output_all_columns': False,
'type': 'tensorflow'}
```
"""
dataset = copy.deepcopy(self)
dataset.set_format(type=type, columns=columns, output_all_columns=output_all_columns, **format_kwargs)
return dataset
def with_transform(
self,
transform: Optional[Callable],
columns: Optional[List] = None,
output_all_columns: bool = False,
):
"""Set `__getitem__` return format using this transform. The transform is applied on-the-fly on batches when `__getitem__` is called.
As [`~datasets.Dataset.set_format`], this can be reset using [`~datasets.Dataset.reset_format`].
Contrary to [`~datasets.Dataset.set_transform`], `with_transform` returns a new [`Dataset`] object.
Args:
transform (`Callable`, `optional`):
User-defined formatting transform, replaces the format defined by [`~datasets.Dataset.set_format`].
A formatting function is a callable that takes a batch (as a `dict`) as input and returns a batch.
This function is applied right before returning the objects in `__getitem__`.
columns (`List[str]`, `optional`):
Columns to format in the output.
If specified, then the input batch of the transform only contains those columns.
output_all_columns (`bool`, defaults to `False`):
Keep un-formatted columns as well in the output (as python objects).
If set to `True`, then the other un-formatted columns are kept with the output of the transform.
Example:
```py
>>> from datasets import load_dataset
>>> from transformers import AutoTokenizer
>>> ds = load_dataset("rotten_tomatoes", split="validation")
>>> tokenizer = AutoTokenizer.from_pretrained("bert-base-cased")
>>> def encode(example):
... return tokenizer(example["text"], padding=True, truncation=True, return_tensors='pt')
>>> ds = ds.with_transform(encode)
>>> ds[0]
{'attention_mask': tensor([1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1,
1, 1, 1, 1, 1]),
'input_ids': tensor([ 101, 18027, 16310, 16001, 1103, 9321, 178, 11604, 7235, 6617,
1742, 2165, 2820, 1206, 6588, 22572, 12937, 1811, 2153, 1105,
1147, 12890, 19587, 6463, 1105, 15026, 1482, 119, 102]),
'token_type_ids': tensor([0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0])}
```
"""
dataset = copy.deepcopy(self)
dataset.set_transform(transform=transform, columns=columns, output_all_columns=output_all_columns)
return dataset
@deprecated()
def prepare_for_task(self, task: Union[str, TaskTemplate], id: int = 0) -> "Dataset":
"""
Prepare a dataset for the given task by casting the dataset's [`Features`] to standardized column names and types as detailed in [`datasets.tasks`](./task_templates).
Casts [`datasets.DatasetInfo.features`] according to a task-specific schema. Intended for single-use only, so all task templates are removed from [`datasets.DatasetInfo.task_templates`] after casting.
Args:
task (`Union[str, TaskTemplate]`):
The task to prepare the dataset for during training and evaluation. If `str`, supported tasks include:
- `"text-classification"`
- `"question-answering"`
If [`TaskTemplate`], must be one of the task templates in [`datasets.tasks`](./task_templates).
id (`int`, defaults to `0`):
The id required to unambiguously identify the task template when multiple task templates of the same type are supported.
"""
# TODO(lewtun): Add support for casting nested features like answers.text and answers.answer_start in SQuAD
if isinstance(task, str):
tasks = [template.task for template in (self.info.task_templates or [])]
compatible_templates = [template for template in (self.info.task_templates or []) if template.task == task]
if not compatible_templates:
raise ValueError(
f"Task {task} is not compatible with this dataset! Available tasks: {list(unique_values(tasks))}"
)
if not 0 <= id < len(compatible_templates):
templates_list_str = "\n".join(
f"- `{idx}` for task {template}" for idx, template in enumerate(compatible_templates)
)
raise ValueError(
f"Id {id} for task {task} is not in a valid range. Supported ids:\n{templates_list_str}"
)
template = compatible_templates[id]
elif isinstance(task, TaskTemplate):
template = task
else:
raise ValueError(
f"Expected a `str` or `datasets.TaskTemplate` object but got task {task} with type {type(task)}."
)
template = template.align_with_features(self.info.features)
column_mapping = template.column_mapping
columns_to_drop = [column for column in self.column_names if column not in column_mapping]
dataset = self.remove_columns(columns_to_drop)
dataset = dataset.rename_columns(column_mapping)
# We found a template so now flush `DatasetInfo` to skip the template update in `DatasetInfo.__post_init__`
dataset.info.task_templates = None
dataset = dataset.cast(features=template.features)
return dataset
def _getitem(self, key: Union[int, slice, str, ListLike[int]], **kwargs) -> Union[Dict, List]:
"""
Can be used to index columns (by string names) or rows (by integer, slice, or list-like of integer indices)
"""
if isinstance(key, bool):
raise TypeError("dataset index must be int, str, slice or collection of int, not bool")
format_type = kwargs["format_type"] if "format_type" in kwargs else self._format_type
format_columns = kwargs["format_columns"] if "format_columns" in kwargs else self._format_columns
output_all_columns = (
kwargs["output_all_columns"] if "output_all_columns" in kwargs else self._output_all_columns
)
format_kwargs = kwargs["format_kwargs"] if "format_kwargs" in kwargs else self._format_kwargs
format_kwargs = format_kwargs if format_kwargs is not None else {}
formatter = get_formatter(format_type, features=self._info.features, **format_kwargs)
pa_subtable = query_table(self._data, key, indices=self._indices if self._indices is not None else None)
formatted_output = format_table(
pa_subtable, key, formatter=formatter, format_columns=format_columns, output_all_columns=output_all_columns
)
return formatted_output
@overload
def __getitem__(self, key: Union[int, slice, Iterable[int]]) -> Dict: # noqa: F811
...
@overload
def __getitem__(self, key: str) -> List: # noqa: F811
...
def __getitem__(self, key): # noqa: F811
"""Can be used to index columns (by string names) or rows (by integer index or iterable of indices or bools)."""
return self._getitem(key)
def __getitems__(self, keys: List) -> List:
"""Can be used to get a batch using a list of integers indices."""
batch = self.__getitem__(keys)
n_examples = len(batch[next(iter(batch))])
return [{col: array[i] for col, array in batch.items()} for i in range(n_examples)]
def cleanup_cache_files(self) -> int:
"""Clean up all cache files in the dataset cache directory, excepted the currently used cache file if there is
one.
Be careful when running this command that no other process is currently using other cache files.
Returns:
`int`: Number of removed files.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", split="validation")
>>> ds.cleanup_cache_files()
10
```
"""
current_cache_files = [os.path.abspath(cache_file["filename"]) for cache_file in self.cache_files]
if not current_cache_files:
return 0
cache_directory = os.path.dirname(current_cache_files[0])
logger.info(f"Listing files in {cache_directory}")
files: List[str] = os.listdir(cache_directory)
files_to_remove = []
for f_name in files:
full_name = os.path.abspath(os.path.join(cache_directory, f_name))
if f_name.startswith("cache-") and f_name.endswith(".arrow"):
if full_name in current_cache_files:
logger.info(f"Keeping currently used cache file at {full_name}")
continue
files_to_remove.append(full_name)
for file_path in files_to_remove:
logger.info(f"Removing {file_path}")
os.remove(file_path)
return len(files_to_remove)
def _get_cache_file_path(self, fingerprint):
if is_caching_enabled() and self.cache_files:
cache_file_name = "cache-" + fingerprint + ".arrow"
cache_directory = os.path.dirname(self.cache_files[0]["filename"])
else:
cache_file_name = "cache-" + generate_random_fingerprint() + ".arrow"
cache_directory = get_temporary_cache_files_directory()
cache_file_path = os.path.join(cache_directory, cache_file_name)
return cache_file_path
@transmit_tasks
@transmit_format
def map(
self,
function: Optional[Callable] = None,
with_indices: bool = False,
with_rank: bool = False,
input_columns: Optional[Union[str, List[str]]] = None,
batched: bool = False,
batch_size: Optional[int] = 1000,
drop_last_batch: bool = False,
remove_columns: Optional[Union[str, List[str]]] = None,
keep_in_memory: bool = False,
load_from_cache_file: Optional[bool] = None,
cache_file_name: Optional[str] = None,
writer_batch_size: Optional[int] = 1000,
features: Optional[Features] = None,
disable_nullable: bool = False,
fn_kwargs: Optional[dict] = None,
num_proc: Optional[int] = None,
suffix_template: str = "_{rank:05d}_of_{num_proc:05d}",
new_fingerprint: Optional[str] = None,
desc: Optional[str] = None,
) -> "Dataset":
"""
Apply a function to all the examples in the table (individually or in batches) and update the table.
If your function returns a column that already exists, then it overwrites it.
You can specify whether the function should be batched or not with the `batched` parameter:
- If batched is `False`, then the function takes 1 example in and should return 1 example.
An example is a dictionary, e.g. `{"text": "Hello there !"}`.
- If batched is `True` and `batch_size` is 1, then the function takes a batch of 1 example as input and can return a batch with 1 or more examples.
A batch is a dictionary, e.g. a batch of 1 example is `{"text": ["Hello there !"]}`.
- If batched is `True` and `batch_size` is `n > 1`, then the function takes a batch of `n` examples as input and can return a batch with `n` examples, or with an arbitrary number of examples.
Note that the last batch may have less than `n` examples.
A batch is a dictionary, e.g. a batch of `n` examples is `{"text": ["Hello there !"] * n}`.
Args:
function (`Callable`): Function with one of the following signatures:
- `function(example: Dict[str, Any]) -> Dict[str, Any]` if `batched=False` and `with_indices=False` and `with_rank=False`
- `function(example: Dict[str, Any], *extra_args) -> Dict[str, Any]` if `batched=False` and `with_indices=True` and/or `with_rank=True` (one extra arg for each)
- `function(batch: Dict[str, List]) -> Dict[str, List]` if `batched=True` and `with_indices=False` and `with_rank=False`
- `function(batch: Dict[str, List], *extra_args) -> Dict[str, List]` if `batched=True` and `with_indices=True` and/or `with_rank=True` (one extra arg for each)
For advanced usage, the function can also return a `pyarrow.Table`.
Moreover if your function returns nothing (`None`), then `map` will run your function and return the dataset unchanged.
If no function is provided, default to identity function: `lambda x: x`.
with_indices (`bool`, defaults to `False`):
Provide example indices to `function`. Note that in this case the
signature of `function` should be `def function(example, idx[, rank]): ...`.
with_rank (`bool`, defaults to `False`):
Provide process rank to `function`. Note that in this case the
signature of `function` should be `def function(example[, idx], rank): ...`.
input_columns (`Optional[Union[str, List[str]]]`, defaults to `None`):
The columns to be passed into `function`
as positional arguments. If `None`, a `dict` mapping to all formatted columns is passed as one argument.
batched (`bool`, defaults to `False`):
Provide batch of examples to `function`.
batch_size (`int`, *optional*, defaults to `1000`):
Number of examples per batch provided to `function` if `batched=True`.
If `batch_size <= 0` or `batch_size == None`, provide the full dataset as a single batch to `function`.
drop_last_batch (`bool`, defaults to `False`):
Whether a last batch smaller than the batch_size should be
dropped instead of being processed by the function.
remove_columns (`Optional[Union[str, List[str]]]`, defaults to `None`):
Remove a selection of columns while doing the mapping.
Columns will be removed before updating the examples with the output of `function`, i.e. if `function` is adding
columns with names in `remove_columns`, these columns will be kept.
keep_in_memory (`bool`, defaults to `False`):
Keep the dataset in memory instead of writing it to a cache file.
load_from_cache_file (`Optional[bool]`, defaults to `True` if caching is enabled):
If a cache file storing the current computation from `function`
can be identified, use it instead of recomputing.
cache_file_name (`str`, *optional*, defaults to `None`):
Provide the name of a path for the cache file. It is used to store the
results of the computation instead of the automatically generated cache file name.
writer_batch_size (`int`, defaults to `1000`):
Number of rows per write operation for the cache file writer.
This value is a good trade-off between memory usage during the processing, and processing speed.
Higher value makes the processing do fewer lookups, lower value consume less temporary memory while running `map`.
features (`Optional[datasets.Features]`, defaults to `None`):
Use a specific Features to store the cache file
instead of the automatically generated one.
disable_nullable (`bool`, defaults to `False`):
Disallow null values in the table.
fn_kwargs (`Dict`, *optional*, defaults to `None`):
Keyword arguments to be passed to `function`.
num_proc (`int`, *optional*, defaults to `None`):
Max number of processes when generating cache. Already cached shards are loaded sequentially.
suffix_template (`str`):
If `cache_file_name` is specified, then this suffix
will be added at the end of the base name of each. Defaults to `"_{rank:05d}_of_{num_proc:05d}"`. For example, if `cache_file_name` is "processed.arrow", then for
`rank=1` and `num_proc=4`, the resulting file would be `"processed_00001_of_00004.arrow"` for the default suffix.
new_fingerprint (`str`, *optional*, defaults to `None`):
The new fingerprint of the dataset after transform.
If `None`, the new fingerprint is computed using a hash of the previous fingerprint, and the transform arguments.
desc (`str`, *optional*, defaults to `None`):
Meaningful description to be displayed alongside with the progress bar while mapping examples.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", split="validation")
>>> def add_prefix(example):
... example["text"] = "Review: " + example["text"]
... return example
>>> ds = ds.map(add_prefix)
>>> ds[0:3]["text"]
['Review: compassionately explores the seemingly irreconcilable situation between conservative christian parents and their estranged gay and lesbian children .',
'Review: the soundtrack alone is worth the price of admission .',
'Review: rodriguez does a splendid job of racial profiling hollywood style--casting excellent latin actors of all ages--a trend long overdue .']
# process a batch of examples
>>> ds = ds.map(lambda example: tokenizer(example["text"]), batched=True)
# set number of processors
>>> ds = ds.map(add_prefix, num_proc=4)
```
"""
if keep_in_memory and cache_file_name is not None:
raise ValueError("Please use either `keep_in_memory` or `cache_file_name` but not both.")
if num_proc is not None and num_proc <= 0:
raise ValueError("num_proc must be an integer > 0.")
# If the array is empty we do nothing (but we make sure to handle an empty indices mapping and remove the requested columns anyway)
if len(self) == 0:
if self._indices is not None: # empty indices mapping
self = Dataset(
self.data.slice(0, 0),
info=self.info.copy(),
split=self.split,
fingerprint=new_fingerprint,
)
if remove_columns:
return self.remove_columns(remove_columns)
else:
return self
if function is None:
function = lambda x: x # noqa: E731
if isinstance(input_columns, str):
input_columns = [input_columns]
if input_columns is not None:
for input_column in input_columns:
if input_column not in self._data.column_names:
raise ValueError(
f"Input column {input_column} not in the dataset. Current columns in the dataset: {self._data.column_names}"
)
if isinstance(remove_columns, str):
remove_columns = [remove_columns]
if remove_columns is not None and any(col not in self._data.column_names for col in remove_columns):
raise ValueError(
f"Column to remove {list(filter(lambda col: col not in self._data.column_names, remove_columns))} not in the dataset. Current columns in the dataset: {self._data.column_names}"
)
load_from_cache_file = load_from_cache_file if load_from_cache_file is not None else is_caching_enabled()
if fn_kwargs is None:
fn_kwargs = {}
if num_proc is not None and num_proc > len(self):
num_proc = len(self)
logger.warning(
f"num_proc must be <= {len(self)}. Reducing num_proc to {num_proc} for dataset of size {len(self)}."
)
dataset_kwargs = {
"shard": self,
"function": function,
"with_indices": with_indices,
"with_rank": with_rank,
"input_columns": input_columns,
"batched": batched,
"batch_size": batch_size,
"drop_last_batch": drop_last_batch,
"remove_columns": remove_columns,
"keep_in_memory": keep_in_memory,
"writer_batch_size": writer_batch_size,
"features": features,
"disable_nullable": disable_nullable,
"fn_kwargs": fn_kwargs,
}
if new_fingerprint is None:
# we create a unique hash from the function,
# current dataset file and the mapping args
transform = format_transform_for_fingerprint(Dataset._map_single)
kwargs_for_fingerprint = format_kwargs_for_fingerprint(Dataset._map_single, (), dataset_kwargs)
kwargs_for_fingerprint["fingerprint_name"] = "new_fingerprint"
new_fingerprint = update_fingerprint(self._fingerprint, transform, kwargs_for_fingerprint)
else:
validate_fingerprint(new_fingerprint)
dataset_kwargs["new_fingerprint"] = new_fingerprint
if self.cache_files:
if cache_file_name is None:
cache_file_name = self._get_cache_file_path(new_fingerprint)
dataset_kwargs["cache_file_name"] = cache_file_name
def load_processed_shard_from_cache(shard_kwargs):
"""Load a processed shard from cache if it exists, otherwise throw an error."""
shard = shard_kwargs["shard"]
# Check if we've already cached this computation (indexed by a hash)
if shard_kwargs["cache_file_name"] is not None:
if os.path.exists(shard_kwargs["cache_file_name"]) and load_from_cache_file:
info = shard.info.copy()
info.features = features
info.task_templates = None
return Dataset.from_file(shard_kwargs["cache_file_name"], info=info, split=shard.split)
raise NonExistentDatasetError
num_shards = num_proc if num_proc is not None else 1
if batched and drop_last_batch:
pbar_total = len(self) // num_shards // batch_size * num_shards * batch_size
else:
pbar_total = len(self)
shards_done = 0
if num_proc is None or num_proc == 1:
transformed_dataset = None
try:
transformed_dataset = load_processed_shard_from_cache(dataset_kwargs)
logger.info(f"Loading cached processed dataset at {dataset_kwargs['cache_file_name']}")
except NonExistentDatasetError:
pass
if transformed_dataset is None:
with hf_tqdm(
unit=" examples",
total=pbar_total,
desc=desc or "Map",
) as pbar:
for rank, done, content in Dataset._map_single(**dataset_kwargs):
if done:
shards_done += 1
logger.debug(f"Finished processing shard number {rank} of {num_shards}.")
transformed_dataset = content
else:
pbar.update(content)
assert transformed_dataset is not None, "Failed to retrieve the result from map"
# update fingerprint if the dataset changed
if transformed_dataset._fingerprint != self._fingerprint:
transformed_dataset._fingerprint = new_fingerprint
return transformed_dataset
else:
def format_cache_file_name(
cache_file_name: Optional[str],
rank: Union[int, Literal["*"]], # noqa: F722
) -> Optional[str]:
if not cache_file_name:
return cache_file_name
sep = cache_file_name.rindex(".")
base_name, extension = cache_file_name[:sep], cache_file_name[sep:]
if isinstance(rank, int):
cache_file_name = base_name + suffix_template.format(rank=rank, num_proc=num_proc) + extension
logger.info(f"Process #{rank} will write at {cache_file_name}")
else:
cache_file_name = (
base_name
+ suffix_template.replace("{rank:05d}", "{rank}").format(rank=rank, num_proc=num_proc)
+ extension
)
return cache_file_name
def format_new_fingerprint(new_fingerprint: str, rank: int) -> str:
new_fingerprint = new_fingerprint + suffix_template.format(rank=rank, num_proc=num_proc)
validate_fingerprint(new_fingerprint)
return new_fingerprint
prev_env = deepcopy(os.environ)
# check if parallelism if off
# from https://github.com/huggingface/tokenizers/blob/bb668bc439dc34389b71dbb8ce0c597f15707b53/tokenizers/src/utils/parallelism.rs#L22
if prev_env.get("TOKENIZERS_PARALLELISM", "false").lower() not in (
"",
"off",
"false",
"f",
"no",
"n",
"0",
):
logger.warning("Setting TOKENIZERS_PARALLELISM=false for forked processes.")
os.environ["TOKENIZERS_PARALLELISM"] = "false"
shards = [
self.shard(num_shards=num_proc, index=rank, contiguous=True, keep_in_memory=keep_in_memory)
for rank in range(num_proc)
]
kwargs_per_job = [
{
**dataset_kwargs,
"shard": shards[rank],
"cache_file_name": format_cache_file_name(cache_file_name, rank),
"rank": rank,
"offset": sum(len(s) for s in shards[:rank]),
"new_fingerprint": format_new_fingerprint(new_fingerprint, rank),
}
for rank in range(num_shards)
]
transformed_shards = [None] * num_shards
for rank in range(num_shards):
try:
transformed_shards[rank] = load_processed_shard_from_cache(kwargs_per_job[rank])
kwargs_per_job[rank] = None
except NonExistentDatasetError:
pass
kwargs_per_job = [kwargs for kwargs in kwargs_per_job if kwargs is not None]
# We try to create a pool with as many workers as dataset not yet cached.
if kwargs_per_job:
if len(kwargs_per_job) < num_shards:
logger.info(
f"Reprocessing {len(kwargs_per_job)}/{num_shards} shards because some of them were missing from the cache."
)
with Pool(len(kwargs_per_job)) as pool:
os.environ = prev_env
logger.info(f"Spawning {num_proc} processes")
with hf_tqdm(
unit=" examples",
total=pbar_total,
desc=(desc or "Map") + f" (num_proc={num_proc})",
) as pbar:
for rank, done, content in iflatmap_unordered(
pool, Dataset._map_single, kwargs_iterable=kwargs_per_job
):
if done:
shards_done += 1
logger.debug(f"Finished processing shard number {rank} of {num_shards}.")
transformed_shards[rank] = content
else:
pbar.update(content)
# Avoids PermissionError on Windows (the error: https://github.com/huggingface/datasets/actions/runs/4026734820/jobs/6921621805)
for kwargs in kwargs_per_job:
del kwargs["shard"]
else:
logger.info(f"Loading cached processed dataset at {format_cache_file_name(cache_file_name, '*')}")
assert (
None not in transformed_shards
), f"Failed to retrieve results from map: result list {transformed_shards} still contains None - at least one worker failed to return its results"
logger.info(f"Concatenating {num_proc} shards")
result = _concatenate_map_style_datasets(transformed_shards)
# update fingerprint if the dataset changed
if any(
transformed_shard._fingerprint != shard._fingerprint
for transformed_shard, shard in zip(transformed_shards, shards)
):
result._fingerprint = new_fingerprint
else:
result._fingerprint = self._fingerprint
return result
@staticmethod
def _map_single(
shard: "Dataset",
function: Optional[Callable] = None,
with_indices: bool = False,
with_rank: bool = False,
input_columns: Optional[List[str]] = None,
batched: bool = False,
batch_size: Optional[int] = 1000,
drop_last_batch: bool = False,
remove_columns: Optional[List[str]] = None,
keep_in_memory: bool = False,
cache_file_name: Optional[str] = None,
writer_batch_size: Optional[int] = 1000,
features: Optional[Features] = None,
disable_nullable: bool = False,
fn_kwargs: Optional[dict] = None,
new_fingerprint: Optional[str] = None,
rank: Optional[int] = None,
offset: int = 0,
) -> Iterable[Tuple[int, bool, Union[int, "Dataset"]]]:
"""Apply a function to all the elements in the table (individually or in batches)
and update the table (if function does update examples).
Args:
shard (`datasets.Dataset`): Dataset to map the transform on.
function (`Callable`): with one of the following signature:
- `function(example: Dict[str, Any]) -> Dict[str, Any]` if `batched=False` and `with_indices=False` and `with_rank=False`
- `function(example: Dict[str, Any], *extra_args) -> Dict[str, Any]` if `batched=False` and `with_indices=True` and/or `with_rank=True` (one extra arg for each)
- `function(batch: Dict[str, List]) -> Dict[str, List]` if `batched=True` and `with_indices=False` and `with_rank=False`
- `function(batch: Dict[str, List], *extra_args) -> Dict[str, List]` if `batched=True` and `with_indices=True` and/or `with_rank=True` (one extra arg for each)
For advanced usage, the function can also return a `pyarrow.Table`.
Moreover if your function returns nothing (`None`), then `map` will run your function and return the dataset unchanged.
If no function is provided, default to identity function: lambda x: x
with_indices (`bool`, defaults to `False`): Provide example indices to `function`. Note that in this case the signature of `function` should be `def function(example, idx[, rank]): ...`.
with_rank (`bool`, default `False`): Provide process rank to `function`. Note that in this case the signature of `function` should be `def function(example[, idx], rank): ...`.
input_columns (`Optional[List[str]]`, defaults to `None`): The columns to be passed into `function` as
positional arguments. If `None`, a dict mapping to all formatted columns is passed as one argument.
batched (`bool`, defaults to `False`): Provide batch of examples to `function`
batch_size (`int`, optional, defaults to `1000`): Number of examples per batch provided to `function` if `batched=True`
`batch_size <= 0` or `batch_size == None`: Provide the full dataset as a single batch to `function`
drop_last_batch (`bool`, default: `False`): Whether a last batch smaller than the batch_size should be
dropped instead of being processed by the function.
remove_columns (`Optional[List[str]]`, defaults to `None`): Remove a selection of columns while doing the mapping.
Columns will be removed before updating the examples with the output of `function`, i.e. if `function` is adding
columns with names in `remove_columns`, these columns will be kept.
keep_in_memory (`bool`, defaults to `False`): Keep the dataset in memory instead of writing it to a cache file.
cache_file_name (`str`, optional, defaults to `None`): Provide the name of a path for the cache file. It is used to store the
results of the computation instead of the automatically generated cache file name.
writer_batch_size (`int`, default `1000`): Number of rows per write operation for the cache file writer.
This value is a good trade-off between memory usage during the processing, and processing speed.
Higher value makes the processing do fewer lookups, lower value consume less temporary memory while running `.map()`.
features (`Optional[datasets.Features]`, defaults to `None`): Use a specific Features to store the cache file
instead of the automatically generated one.
disable_nullable (`bool`, defaults to `False`): Disallow null values in the table.
fn_kwargs (`Dict`, optional, defaults to `None`): Keyword arguments to be passed to `function`
new_fingerprint (`str`, optional, defaults to `None`): the new fingerprint of the dataset after transform.
If `None`, the new fingerprint is computed using a hash of the previous fingerprint, and the transform arguments
rank: (`int`, optional, defaults to `None`): If specified, this is the process rank when doing multiprocessing
offset: (`int`, defaults to 0): If specified, this is an offset applied to the indices passed to `function` if `with_indices=True`.
"""
if fn_kwargs is None:
fn_kwargs = {}
# If we do batch computation but no batch size is provided, default to the full dataset
if batched and (batch_size is None or batch_size <= 0):
batch_size = shard.num_rows
# We set this variable to True after processing the first example/batch in
# `apply_function_on_filtered_inputs` if the map function returns a dict.
# If set to False, no new arrow table will be created
update_data = None
format_kwargs = shard._format_kwargs.copy()
# Lazy formatting is only available for the default format (None/python)
if not input_columns and shard._format_type is None:
format_kwargs["lazy"] = True
input_formatter = get_formatter(
shard._format_type,
features=shard.features,
**format_kwargs,
)
class NumExamplesMismatchError(Exception):
pass
def validate_function_output(processed_inputs, indices):
"""Validate output of the map function."""
if processed_inputs is not None and not isinstance(processed_inputs, (Mapping, pa.Table, pd.DataFrame)):
raise TypeError(
f"Provided `function` which is applied to all elements of table returns a variable of type {type(processed_inputs)}. Make sure provided `function` returns a variable of type `dict` (or a pyarrow table) to update the dataset or `None` if you are only interested in side effects."
)
elif isinstance(indices, list) and isinstance(processed_inputs, Mapping):
allowed_batch_return_types = (list, np.ndarray, pd.Series)
if config.TF_AVAILABLE and "tensorflow" in sys.modules:
import tensorflow as tf
allowed_batch_return_types += (tf.Tensor,)
if config.TORCH_AVAILABLE and "torch" in sys.modules:
import torch
allowed_batch_return_types += (torch.Tensor,)
if config.JAX_AVAILABLE and "jax" in sys.modules:
import jax.numpy as jnp
allowed_batch_return_types += (jnp.ndarray,)
all_dict_values_are_lists = all(
isinstance(value, allowed_batch_return_types) for value in processed_inputs.values()
)
if all_dict_values_are_lists is False:
raise TypeError(
f"Provided `function` which is applied to all elements of table returns a `dict` of types {[type(x) for x in processed_inputs.values()]}. When using `batched=True`, make sure provided `function` returns a `dict` of types like `{allowed_batch_return_types}`."
)
def apply_function_on_filtered_inputs(pa_inputs, indices, check_same_num_examples=False, offset=0):
"""Utility to apply the function on a selection of columns."""
nonlocal update_data
inputs = format_table(
pa_inputs,
0 if not batched else range(pa_inputs.num_rows),
format_columns=input_columns,
formatter=input_formatter,
)
fn_args = [inputs] if input_columns is None else [inputs[col] for col in input_columns]
if offset == 0:
effective_indices = indices
else:
effective_indices = [i + offset for i in indices] if isinstance(indices, list) else indices + offset
additional_args = ()
if with_indices:
additional_args += (effective_indices,)
if with_rank:
additional_args += (rank,)
processed_inputs = function(*fn_args, *additional_args, **fn_kwargs)
if isinstance(processed_inputs, LazyDict):
processed_inputs = {
k: v for k, v in processed_inputs.data.items() if k not in processed_inputs.keys_to_format
}
returned_lazy_dict = True
else:
returned_lazy_dict = False
if update_data is None:
# Check if the function returns updated examples
update_data = isinstance(processed_inputs, (Mapping, pa.Table, pd.DataFrame))
validate_function_output(processed_inputs, indices)
if not update_data:
return None # Nothing to update, let's move on
if shard._format_type or input_columns:
# TODO(QL, MS): ideally the behavior should be the same even if the dataset is formatted (may require major release)
inputs_to_merge = dict(zip(pa_inputs.column_names, pa_inputs.itercolumns()))
elif isinstance(inputs, LazyDict):
inputs_to_merge = {
k: (v if k not in inputs.keys_to_format else pa_inputs[k]) for k, v in inputs.data.items()
}
else:
inputs_to_merge = inputs
if remove_columns is not None:
for column in remove_columns:
# `function` can modify input in-place causing column to be already removed.
if column in inputs_to_merge:
inputs_to_merge.pop(column)
if returned_lazy_dict and column in processed_inputs:
processed_inputs.pop(column)
if check_same_num_examples:
input_num_examples = len(pa_inputs)
processed_inputs_num_examples = len(processed_inputs[next(iter(processed_inputs.keys()))])
if input_num_examples != processed_inputs_num_examples:
raise NumExamplesMismatchError()
if isinstance(inputs, Mapping) and isinstance(processed_inputs, Mapping):
# The .map() transform *updates* the dataset:
# the output dictionary contains both the the input data and the output data.
# The output dictionary may contain Arrow values from `inputs_to_merge` so that we can re-write them efficiently.
return {**inputs_to_merge, **processed_inputs}
else:
return processed_inputs
def init_buffer_and_writer():
# Prepare output buffer and batched writer in memory or on file if we update the table
writer_features = features
if writer_features is None:
writer_features = shard.features
update_features = True
else:
update_features = False
if keep_in_memory or cache_file_name is None:
buf_writer = pa.BufferOutputStream()
tmp_file = None
writer = ArrowWriter(
features=writer_features,
stream=buf_writer,
writer_batch_size=writer_batch_size,
update_features=update_features,
fingerprint=new_fingerprint,
disable_nullable=disable_nullable,
)
else:
buf_writer = None
logger.info(f"Caching processed dataset at {cache_file_name}")
tmp_file = tempfile.NamedTemporaryFile("wb", dir=os.path.dirname(cache_file_name), delete=False)
writer = ArrowWriter(
features=writer_features,
path=tmp_file.name,
writer_batch_size=writer_batch_size,
update_features=update_features,
fingerprint=new_fingerprint,
disable_nullable=disable_nullable,
)
return buf_writer, writer, tmp_file
num_examples_progress_update = 0
# If `update_data` is True after processing the first example/batch, initalize these resources with `init_buffer_and_writer`
buf_writer, writer, tmp_file = None, None, None
# Optionally initialize the writer as a context manager
with contextlib.ExitStack() as stack:
try:
arrow_formatted_shard = shard.with_format("arrow")
# Loop over single examples or batches and write to buffer/file if examples are to be updated
if not batched:
shard_iterable = enumerate(arrow_formatted_shard)
else:
num_rows = len(shard) if not drop_last_batch else len(shard) // batch_size * batch_size
shard_iterable = zip(
range(0, num_rows, batch_size),
arrow_formatted_shard.iter(batch_size, drop_last_batch=drop_last_batch),
)
if not batched:
_time = time.time()
for i, example in shard_iterable:
example = apply_function_on_filtered_inputs(example, i, offset=offset)
if update_data:
if i == 0:
buf_writer, writer, tmp_file = init_buffer_and_writer()
stack.enter_context(writer)
if isinstance(example, pa.Table):
writer.write_row(example)
elif isinstance(example, pd.DataFrame):
writer.write_row(pa.Table.from_pandas(example))
else:
writer.write(example)
num_examples_progress_update += 1
if time.time() > _time + config.PBAR_REFRESH_TIME_INTERVAL:
_time = time.time()
yield rank, False, num_examples_progress_update
num_examples_progress_update = 0
else:
_time = time.time()
for i, batch in shard_iterable:
num_examples_in_batch = len(batch)
indices = list(
range(*(slice(i, i + batch_size).indices(shard.num_rows)))
) # Something simpler?
try:
batch = apply_function_on_filtered_inputs(
batch,
indices,
check_same_num_examples=len(shard.list_indexes()) > 0,
offset=offset,
)
except NumExamplesMismatchError:
raise DatasetTransformationNotAllowedError(
"Using `.map` in batched mode on a dataset with attached indexes is allowed only if it doesn't create or remove existing examples. You can first run `.drop_index() to remove your index and then re-add it."
) from None
if update_data:
if i == 0:
buf_writer, writer, tmp_file = init_buffer_and_writer()
stack.enter_context(writer)
if isinstance(batch, pa.Table):
writer.write_table(batch)
elif isinstance(batch, pd.DataFrame):
writer.write_table(pa.Table.from_pandas(batch))
else:
writer.write_batch(batch)
num_examples_progress_update += num_examples_in_batch
if time.time() > _time + config.PBAR_REFRESH_TIME_INTERVAL:
_time = time.time()
yield rank, False, num_examples_progress_update
num_examples_progress_update = 0
if update_data and writer is not None:
writer.finalize() # close_stream=bool(buf_writer is None)) # We only close if we are writing in a file
except (Exception, KeyboardInterrupt):
yield rank, False, num_examples_progress_update
if update_data:
if writer is not None:
writer.finalize()
if tmp_file is not None:
tmp_file.close()
if os.path.exists(tmp_file.name):
os.remove(tmp_file.name)
raise
yield rank, False, num_examples_progress_update
if update_data and tmp_file is not None:
tmp_file.close()
shutil.move(tmp_file.name, cache_file_name)
umask = os.umask(0o666)
os.umask(umask)
os.chmod(cache_file_name, 0o666 & ~umask)
if update_data:
# Create new Dataset from buffer or file
info = shard.info.copy()
info.features = writer._features
info.task_templates = None
if buf_writer is None:
yield rank, True, Dataset.from_file(cache_file_name, info=info, split=shard.split)
else:
yield rank, True, Dataset.from_buffer(buf_writer.getvalue(), info=info, split=shard.split)
else:
yield rank, True, shard
@transmit_format
@fingerprint_transform(
inplace=False, ignore_kwargs=["load_from_cache_file", "cache_file_name", "desc"], version="2.0.1"
)
def filter(
self,
function: Optional[Callable] = None,
with_indices=False,
input_columns: Optional[Union[str, List[str]]] = None,
batched: bool = False,
batch_size: Optional[int] = 1000,
keep_in_memory: bool = False,
load_from_cache_file: Optional[bool] = None,
cache_file_name: Optional[str] = None,
writer_batch_size: Optional[int] = 1000,
fn_kwargs: Optional[dict] = None,
num_proc: Optional[int] = None,
suffix_template: str = "_{rank:05d}_of_{num_proc:05d}",
new_fingerprint: Optional[str] = None,
desc: Optional[str] = None,
) -> "Dataset":
"""Apply a filter function to all the elements in the table in batches
and update the table so that the dataset only includes examples according to the filter function.
Args:
function (`Callable`): Callable with one of the following signatures:
- `function(example: Dict[str, Any]) -> bool` if `with_indices=False, batched=False`
- `function(example: Dict[str, Any], indices: int) -> bool` if `with_indices=True, batched=False`
- `function(example: Dict[str, List]) -> List[bool]` if `with_indices=False, batched=True`
- `function(example: Dict[str, List], indices: List[int]) -> List[bool]` if `with_indices=True, batched=True`
If no function is provided, defaults to an always `True` function: `lambda x: True`.
with_indices (`bool`, defaults to `False`):
Provide example indices to `function`. Note that in this case the signature of `function` should be `def function(example, idx): ...`.
input_columns (`str` or `List[str]`, *optional*):
The columns to be passed into `function` as
positional arguments. If `None`, a `dict` mapping to all formatted columns is passed as one argument.
batched (`bool`, defaults to `False`):
Provide batch of examples to `function`.
batch_size (`int`, *optional*, defaults to `1000`):
Number of examples per batch provided to `function` if
`batched = True`. If `batched = False`, one example per batch is passed to `function`.
If `batch_size <= 0` or `batch_size == None`, provide the full dataset as a single batch to `function`.
keep_in_memory (`bool`, defaults to `False`):
Keep the dataset in memory instead of writing it to a cache file.
load_from_cache_file (`Optional[bool]`, defaults to `True` if caching is enabled):
If a cache file storing the current computation from `function`
can be identified, use it instead of recomputing.
cache_file_name (`str`, *optional*):
Provide the name of a path for the cache file. It is used to store the
results of the computation instead of the automatically generated cache file name.
writer_batch_size (`int`, defaults to `1000`):
Number of rows per write operation for the cache file writer.
This value is a good trade-off between memory usage during the processing, and processing speed.
Higher value makes the processing do fewer lookups, lower value consume less temporary memory while running `map`.
fn_kwargs (`dict`, *optional*):
Keyword arguments to be passed to `function`.
num_proc (`int`, *optional*):
Number of processes for multiprocessing. By default it doesn't
use multiprocessing.
suffix_template (`str`):
If `cache_file_name` is specified, then this suffix will be added at the end of the base name of each.
For example, if `cache_file_name` is `"processed.arrow"`, then for `rank = 1` and `num_proc = 4`,
the resulting file would be `"processed_00001_of_00004.arrow"` for the default suffix (default
`_{rank:05d}_of_{num_proc:05d}`).
new_fingerprint (`str`, *optional*):
The new fingerprint of the dataset after transform.
If `None`, the new fingerprint is computed using a hash of the previous fingerprint, and the transform arguments.
desc (`str`, *optional*, defaults to `None`):
Meaningful description to be displayed alongside with the progress bar while filtering examples.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", split="validation")
>>> ds.filter(lambda x: x["label"] == 1)
Dataset({
features: ['text', 'label'],
num_rows: 533
})
```
"""
if len(self.list_indexes()) > 0:
raise DatasetTransformationNotAllowedError(
"Using `.filter` on a dataset with attached indexes is not allowed. You can first run `.drop_index() to remove your index and then re-add it.`"
)
if function is None:
function = lambda x: True # noqa: E731
if len(self) == 0:
return self
indices = self.map(
function=partial(
get_indices_from_mask_function, function, batched, with_indices, input_columns, self._indices
),
with_indices=True,
features=Features({"indices": Value("uint64")}),
batched=True,
batch_size=batch_size,
remove_columns=self.column_names,
keep_in_memory=keep_in_memory,
load_from_cache_file=load_from_cache_file,
cache_file_name=cache_file_name,
writer_batch_size=writer_batch_size,
fn_kwargs=fn_kwargs,
num_proc=num_proc,
suffix_template=suffix_template,
new_fingerprint=new_fingerprint,
input_columns=input_columns,
desc=desc or "Filter",
)
new_dataset = copy.deepcopy(self)
new_dataset._indices = indices.data
new_dataset._fingerprint = new_fingerprint
return new_dataset
@transmit_format
@fingerprint_transform(inplace=False, ignore_kwargs=["cache_file_name"])
def flatten_indices(
self,
keep_in_memory: bool = False,
cache_file_name: Optional[str] = None,
writer_batch_size: Optional[int] = 1000,
features: Optional[Features] = None,
disable_nullable: bool = False,
num_proc: Optional[int] = None,
new_fingerprint: Optional[str] = None,
) -> "Dataset":
"""Create and cache a new Dataset by flattening the indices mapping.
Args:
keep_in_memory (`bool`, defaults to `False`):
Keep the dataset in memory instead of writing it to a cache file.
cache_file_name (`str`, *optional*, default `None`):
Provide the name of a path for the cache file. It is used to store the
results of the computation instead of the automatically generated cache file name.
writer_batch_size (`int`, defaults to `1000`):
Number of rows per write operation for the cache file writer.
This value is a good trade-off between memory usage during the processing, and processing speed.
Higher value makes the processing do fewer lookups, lower value consume less temporary memory while running `map`.
features (`Optional[datasets.Features]`, defaults to `None`):
Use a specific [`Features`] to store the cache file
instead of the automatically generated one.
disable_nullable (`bool`, defaults to `False`):
Allow null values in the table.
num_proc (`int`, optional, default `None`):
Max number of processes when generating cache. Already cached shards are loaded sequentially
new_fingerprint (`str`, *optional*, defaults to `None`):
The new fingerprint of the dataset after transform.
If `None`, the new fingerprint is computed using a hash of the previous fingerprint, and the transform arguments
"""
return self.map(
batched=True, # for speed
keep_in_memory=keep_in_memory,
cache_file_name=cache_file_name,
writer_batch_size=writer_batch_size,
features=features,
disable_nullable=disable_nullable,
new_fingerprint=new_fingerprint,
desc="Flattening the indices",
num_proc=num_proc,
)
def _new_dataset_with_indices(
self,
indices_cache_file_name: Optional[str] = None,
indices_buffer: Optional[pa.Buffer] = None,
fingerprint: Optional[str] = None,
) -> "Dataset":
"""Return a new Dataset obtained by adding indices (provided in indices_cache_file_name or in a buffer) to the
current Dataset.
"""
if indices_cache_file_name is None and indices_buffer is None:
raise ValueError("At least one of indices_cache_file_name or indices_buffer must be provided.")
if fingerprint is None:
raise ValueError("please specify a fingerprint for the dataset with indices")
if indices_cache_file_name is not None:
indices_table = MemoryMappedTable.from_file(indices_cache_file_name)
else:
indices_table = InMemoryTable.from_buffer(indices_buffer)
# Return new Dataset object
# don't forget to copy the objects
return Dataset(
self._data,
info=self.info.copy(),
split=self.split,
indices_table=indices_table,
fingerprint=fingerprint,
)
@transmit_format
@fingerprint_transform(inplace=False, ignore_kwargs=["indices_cache_file_name"])
def select(
self,
indices: Iterable,
keep_in_memory: bool = False,
indices_cache_file_name: Optional[str] = None,
writer_batch_size: Optional[int] = 1000,
new_fingerprint: Optional[str] = None,
) -> "Dataset":
"""Create a new dataset with rows selected following the list/array of indices.
Args:
indices (`range`, `list`, `iterable`, `ndarray` or `Series`):
Range, list or 1D-array of integer indices for indexing.
If the indices correspond to a contiguous range, the Arrow table is simply sliced.
However passing a list of indices that are not contiguous creates indices mapping, which is much less efficient,
but still faster than recreating an Arrow table made of the requested rows.
keep_in_memory (`bool`, defaults to `False`):
Keep the indices mapping in memory instead of writing it to a cache file.
indices_cache_file_name (`str`, *optional*, defaults to `None`):
Provide the name of a path for the cache file. It is used to store the
indices mapping instead of the automatically generated cache file name.
writer_batch_size (`int`, defaults to `1000`):
Number of rows per write operation for the cache file writer.
This value is a good trade-off between memory usage during the processing, and processing speed.
Higher value makes the processing do fewer lookups, lower value consume less temporary memory while running `map`.
new_fingerprint (`str`, *optional*, defaults to `None`):
The new fingerprint of the dataset after transform.
If `None`, the new fingerprint is computed using a hash of the previous fingerprint, and the transform arguments.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", split="validation")
>>> ds.select(range(4))
Dataset({
features: ['text', 'label'],
num_rows: 4
})
```
"""
if keep_in_memory and indices_cache_file_name is not None:
raise ValueError("Please use either `keep_in_memory` or `indices_cache_file_name` but not both.")
if len(self.list_indexes()) > 0:
raise DatasetTransformationNotAllowedError(
"Using `.select` on a dataset with attached indexes is not allowed. You can first run `.drop_index() to remove your index and then re-add it."
)
# If the array is empty we do nothing
if len(self) == 0:
return self
# If indices is a PyArrow array, we convert to NumPy
if isinstance(indices, (pa.Array, pa.ChunkedArray)):
indices = indices.to_numpy().astype(np.int64)
# Convert generator objects to lists
if isinstance(indices, Iterator):
indices = list(indices)
# If the indices are contiguous, simply slice the arrow table
if isinstance(indices, range):
if _is_range_contiguous(indices) and indices.start >= 0:
start, length = indices.start, indices.stop - indices.start
return self._select_contiguous(start, length, new_fingerprint=new_fingerprint)
else:
try:
start = next(iter(indices))
except StopIteration:
# if `indices` is an empty iterable, we return an empty dataset
return self._select_contiguous(0, 0, new_fingerprint=new_fingerprint)
if start >= 0:
counter_from_start = itertools.count(start=start)
if all(i == j for i, j in zip(indices, counter_from_start)):
length = next(counter_from_start) - start
return self._select_contiguous(start, length, new_fingerprint=new_fingerprint)
# If not contiguous, we need to create a new indices mapping
return self._select_with_indices_mapping(
indices,
keep_in_memory=keep_in_memory,
indices_cache_file_name=indices_cache_file_name,
writer_batch_size=writer_batch_size,
new_fingerprint=new_fingerprint,
)
@transmit_format
@fingerprint_transform(inplace=False)
def _select_contiguous(
self,
start: int,
length: int,
new_fingerprint: Optional[str] = None,
) -> "Dataset":
"""Create a new dataset with rows from a contiguous slice of data.
The slice is defined by that start index and its length.
Args:
start (`int`): start index.
length (`int`): length of the slice to select.
new_fingerprint (`str`, optional, default `None`): the new fingerprint of the dataset after transform.
If `None`, the new fingerprint is computed using a hash of the previous fingerprint, and the transform arguments
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", split="validation")
>>> ds._select_contiguous(0, 4)
Dataset({
features: ['text', 'label'],
num_rows: 4
})
```
"""
if len(self.list_indexes()) > 0:
raise DatasetTransformationNotAllowedError(
"Using `.select` on a dataset with attached indexes is not allowed. You can first run `.drop_index() to remove your index and then re-add it."
)
# If the array is empty we do nothing
if len(self) == 0:
return self
_check_valid_indices_value(start, len(self))
_check_valid_indices_value(start + length - 1, len(self))
if self._indices is None or length == 0:
return Dataset(
self.data.slice(start, length),
info=self.info.copy(),
split=self.split,
fingerprint=new_fingerprint,
)
else:
return Dataset(
self.data,
info=self.info.copy(),
split=self.split,
indices_table=self._indices.slice(start, length),
fingerprint=new_fingerprint,
)
@transmit_format
@fingerprint_transform(inplace=False, ignore_kwargs=["indices_cache_file_name"])
def _select_with_indices_mapping(
self,
indices: Iterable,
keep_in_memory: bool = False,
indices_cache_file_name: Optional[str] = None,
writer_batch_size: Optional[int] = 1000,
new_fingerprint: Optional[str] = None,
) -> "Dataset":
"""Create a new dataset with rows selected following the list/array of indices.
The new dataset is made by creating a new indices mapping on top of the main arrow table.
Args:
indices (sequence, iterable, range, ndarray or Series): List or 1D-array of integer indices for indexing.
keep_in_memory (`bool`, default `False`): Keep the indices mapping in memory instead of writing it to a cache file.
indices_cache_file_name (`str`, optional, default `None`): Provide the name of a path for the cache file. It is used to store the
indices mapping instead of the automatically generated cache file name.
writer_batch_size (`int`, default `1000`): Number of rows per write operation for the cache file writer.
This value is a good trade-off between memory usage during the processing, and processing speed.
Higher value makes the processing do fewer lookups, lower value consume less temporary memory while running `.map()`.
new_fingerprint (`str`, optional, default `None`): the new fingerprint of the dataset after transform.
If `None`, the new fingerprint is computed using a hash of the previous fingerprint, and the transform arguments
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", split="validation")
>>> ds._select_with_indices_mapping(range(4))
Dataset({
features: ['text', 'label'],
num_rows: 4
})
```
"""
if keep_in_memory and indices_cache_file_name is not None:
raise ValueError("Please use either `keep_in_memory` or `indices_cache_file_name` but not both.")
if len(self.list_indexes()) > 0:
raise DatasetTransformationNotAllowedError(
"Using `.select` on a dataset with attached indexes is not allowed. You can first run `.drop_index() to remove your index and then re-add it."
)
# If the array is empty we do nothing
if len(self) == 0:
return self
# Prepare the writer for our indices arrow table
if keep_in_memory or indices_cache_file_name is None:
buf_writer = pa.BufferOutputStream()
tmp_file = None
writer = ArrowWriter(
stream=buf_writer, writer_batch_size=writer_batch_size, fingerprint=new_fingerprint, unit="indices"
)
else:
buf_writer = None
logger.info(f"Caching indices mapping at {indices_cache_file_name}")
tmp_file = tempfile.NamedTemporaryFile("wb", dir=os.path.dirname(indices_cache_file_name), delete=False)
writer = ArrowWriter(
path=tmp_file.name, writer_batch_size=writer_batch_size, fingerprint=new_fingerprint, unit="indices"
)
indices = indices if isinstance(indices, list) else list(indices)
size = len(self)
if indices:
_check_valid_indices_value(int(max(indices)), size=size)
_check_valid_indices_value(int(min(indices)), size=size)
else:
return self._select_contiguous(0, 0, new_fingerprint=new_fingerprint)
indices_array = pa.array(indices, type=pa.uint64())
# Check if we need to convert indices
if self._indices is not None:
indices_array = self._indices.column(0).take(indices_array)
indices_table = pa.Table.from_arrays([indices_array], names=["indices"])
with writer:
try:
writer.write_table(indices_table)
writer.finalize() # close_stream=bool(buf_writer is None)) We only close if we are writing in a file
except (Exception, KeyboardInterrupt):
if tmp_file is not None:
tmp_file.close()
if os.path.exists(tmp_file.name):
os.remove(tmp_file.name)
raise
if tmp_file is not None:
tmp_file.close()
shutil.move(tmp_file.name, indices_cache_file_name)
umask = os.umask(0o666)
os.umask(umask)
os.chmod(indices_cache_file_name, 0o666 & ~umask)
# Return new Dataset object
if buf_writer is None:
return self._new_dataset_with_indices(
indices_cache_file_name=indices_cache_file_name, fingerprint=new_fingerprint
)
else:
return self._new_dataset_with_indices(indices_buffer=buf_writer.getvalue(), fingerprint=new_fingerprint)
@transmit_format
@fingerprint_transform(inplace=False, ignore_kwargs=["load_from_cache_file", "indices_cache_file_name"])
def sort(
self,
column_names: Union[str, Sequence_[str]],
reverse: Union[bool, Sequence_[bool]] = False,
kind="deprecated",
null_placement: str = "at_end",
keep_in_memory: bool = False,
load_from_cache_file: Optional[bool] = None,
indices_cache_file_name: Optional[str] = None,
writer_batch_size: Optional[int] = 1000,
new_fingerprint: Optional[str] = None,
) -> "Dataset":
"""Create a new dataset sorted according to a single or multiple columns.
Args:
column_names (`Union[str, Sequence[str]]`):
Column name(s) to sort by.
reverse (`Union[bool, Sequence[bool]]`, defaults to `False`):
If `True`, sort by descending order rather than ascending. If a single bool is provided,
the value is applied to the sorting of all column names. Otherwise a list of bools with the
same length and order as column_names must be provided.
kind (`str`, *optional*):
Pandas algorithm for sorting selected in `{quicksort, mergesort, heapsort, stable}`,
The default is `quicksort`. Note that both `stable` and `mergesort` use `timsort` under the covers and, in general,
the actual implementation will vary with data type. The `mergesort` option is retained for backwards compatibility.
<Deprecated version="2.8.0">
`kind` was deprecated in version 2.10.0 and will be removed in 3.0.0.
</Deprecated>
null_placement (`str`, defaults to `at_end`):
Put `None` values at the beginning if `at_start` or `first` or at the end if `at_end` or `last`
<Added version="1.14.2"/>
keep_in_memory (`bool`, defaults to `False`):
Keep the sorted indices in memory instead of writing it to a cache file.
load_from_cache_file (`Optional[bool]`, defaults to `True` if caching is enabled):
If a cache file storing the sorted indices
can be identified, use it instead of recomputing.
indices_cache_file_name (`str`, *optional*, defaults to `None`):
Provide the name of a path for the cache file. It is used to store the
sorted indices instead of the automatically generated cache file name.
writer_batch_size (`int`, defaults to `1000`):
Number of rows per write operation for the cache file writer.
Higher value gives smaller cache files, lower value consume less temporary memory.
new_fingerprint (`str`, *optional*, defaults to `None`):
The new fingerprint of the dataset after transform.
If `None`, the new fingerprint is computed using a hash of the previous fingerprint, and the transform arguments
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset('rotten_tomatoes', split='validation')
>>> ds['label'][:10]
[1, 1, 1, 1, 1, 1, 1, 1, 1, 1]
>>> sorted_ds = ds.sort('label')
>>> sorted_ds['label'][:10]
[0, 0, 0, 0, 0, 0, 0, 0, 0, 0]
>>> another_sorted_ds = ds.sort(['label', 'text'], reverse=[True, False])
>>> another_sorted_ds['label'][:10]
[1, 1, 1, 1, 1, 1, 1, 1, 1, 1]
```
"""
if len(self.list_indexes()) > 0:
raise DatasetTransformationNotAllowedError(
"Using `.sort` on a dataset with attached indexes is not allowed. You can first run `.drop_index() to remove your index and then re-add it."
)
# If the array is empty we do nothing
if len(self) == 0:
return self
# Deprecation warning
if kind != "deprecated":
warnings.warn(
"'kind' was deprecated in version 2.10.0 and will be removed in 3.0.0.",
category=FutureWarning,
)
# Check proper format of and for duplicates in column_names
if isinstance(column_names, str):
column_names = [column_names]
# Check proper format and length of reverse
if not isinstance(reverse, bool):
if len(reverse) != len(column_names):
raise ValueError(
"Parameter 'reverse' should be either a boolean or a list of booleans with the same length as 'column_names'."
)
else:
reverse = [reverse] * len(column_names)
# Check whether column name(s) exist in dataset
for column in column_names:
if not isinstance(column, str) or column not in self._data.column_names:
raise ValueError(
f"Column '{column}' not found in the dataset. Please provide a column selected in: {self._data.column_names}"
)
# Change null_placement to conform to pyarrow's sort_indices() while ensuring backwards compatability
if null_placement not in ["at_start", "at_end"]:
if null_placement == "first":
null_placement = "at_start"
elif null_placement == "last":
null_placement = "at_end"
else:
raise ValueError(
f"null_placement '{null_placement}' is an invalid parameter value. Must be either 'last', 'at_end', 'first' or 'at_start'."
)
load_from_cache_file = load_from_cache_file if load_from_cache_file is not None else is_caching_enabled()
# Check if we've already cached this computation (indexed by a hash)
if self.cache_files:
if indices_cache_file_name is None:
# we create a unique hash from the function, current dataset file and the mapping args
indices_cache_file_name = self._get_cache_file_path(new_fingerprint)
if os.path.exists(indices_cache_file_name) and load_from_cache_file:
logger.info(f"Loading cached sorted indices for dataset at {indices_cache_file_name}")
return self._new_dataset_with_indices(
fingerprint=new_fingerprint, indices_cache_file_name=indices_cache_file_name
)
sort_table = query_table(
table=self._data,
key=slice(0, len(self)),
indices=self._indices if self._indices is not None else None,
)
sort_keys = [
(col, "ascending" if not col_reverse else "descending") for col, col_reverse in zip(column_names, reverse)
]
indices = pc.sort_indices(sort_table, sort_keys=sort_keys, null_placement=null_placement)
return self.select(
indices=indices,
keep_in_memory=keep_in_memory,
indices_cache_file_name=indices_cache_file_name,
writer_batch_size=writer_batch_size,
new_fingerprint=new_fingerprint,
)
@transmit_format
@fingerprint_transform(
inplace=False, randomized_function=True, ignore_kwargs=["load_from_cache_file", "indices_cache_file_name"]
)
def shuffle(
self,
seed: Optional[int] = None,
generator: Optional[np.random.Generator] = None,
keep_in_memory: bool = False,
load_from_cache_file: Optional[bool] = None,
indices_cache_file_name: Optional[str] = None,
writer_batch_size: Optional[int] = 1000,
new_fingerprint: Optional[str] = None,
) -> "Dataset":
"""Create a new Dataset where the rows are shuffled.
Currently shuffling uses numpy random generators.
You can either supply a NumPy BitGenerator to use, or a seed to initiate NumPy's default random generator (PCG64).
Shuffling takes the list of indices `[0:len(my_dataset)]` and shuffles it to create an indices mapping.
However as soon as your [`Dataset`] has an indices mapping, the speed can become 10x slower.
This is because there is an extra step to get the row index to read using the indices mapping, and most importantly, you aren't reading contiguous chunks of data anymore.
To restore the speed, you'd need to rewrite the entire dataset on your disk again using [`Dataset.flatten_indices`], which removes the indices mapping.
This may take a lot of time depending of the size of your dataset though:
```python
my_dataset[0] # fast
my_dataset = my_dataset.shuffle(seed=42)
my_dataset[0] # up to 10x slower
my_dataset = my_dataset.flatten_indices() # rewrite the shuffled dataset on disk as contiguous chunks of data
my_dataset[0] # fast again
```
In this case, we recommend switching to an [`IterableDataset`] and leveraging its fast approximate shuffling method [`IterableDataset.shuffle`].
It only shuffles the shards order and adds a shuffle buffer to your dataset, which keeps the speed of your dataset optimal:
```python
my_iterable_dataset = my_dataset.to_iterable_dataset(num_shards=128)
for example in enumerate(my_iterable_dataset): # fast
pass
shuffled_iterable_dataset = my_iterable_dataset.shuffle(seed=42, buffer_size=100)
for example in enumerate(shuffled_iterable_dataset): # as fast as before
pass
```
Args:
seed (`int`, *optional*):
A seed to initialize the default BitGenerator if `generator=None`.
If `None`, then fresh, unpredictable entropy will be pulled from the OS.
If an `int` or `array_like[ints]` is passed, then it will be passed to SeedSequence to derive the initial BitGenerator state.
generator (`numpy.random.Generator`, *optional*):
Numpy random Generator to use to compute the permutation of the dataset rows.
If `generator=None` (default), uses `np.random.default_rng` (the default BitGenerator (PCG64) of NumPy).
keep_in_memory (`bool`, default `False`):
Keep the shuffled indices in memory instead of writing it to a cache file.
load_from_cache_file (`Optional[bool]`, defaults to `True` if caching is enabled):
If a cache file storing the shuffled indices
can be identified, use it instead of recomputing.
indices_cache_file_name (`str`, *optional*):
Provide the name of a path for the cache file. It is used to store the
shuffled indices instead of the automatically generated cache file name.
writer_batch_size (`int`, defaults to `1000`):
Number of rows per write operation for the cache file writer.
This value is a good trade-off between memory usage during the processing, and processing speed.
Higher value makes the processing do fewer lookups, lower value consume less temporary memory while running `map`.
new_fingerprint (`str`, *optional*, defaults to `None`):
The new fingerprint of the dataset after transform.
If `None`, the new fingerprint is computed using a hash of the previous fingerprint, and the transform arguments.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", split="validation")
>>> ds['label'][:10]
[1, 1, 1, 1, 1, 1, 1, 1, 1, 1]
# set a seed
>>> shuffled_ds = ds.shuffle(seed=42)
>>> shuffled_ds['label'][:10]
[1, 0, 1, 1, 0, 0, 0, 0, 0, 0]
```
"""
if len(self.list_indexes()) > 0:
raise DatasetTransformationNotAllowedError(
"Using `.shuffle` on a dataset with attached indexes is not allowed. You can first run `.drop_index() to remove your index and then re-add it."
)
# If the array is empty we do nothing
if len(self) == 0:
return self
if keep_in_memory and indices_cache_file_name is not None:
raise ValueError("Please use either `keep_in_memory` or `indices_cache_file_name` but not both.")
if seed is not None and generator is not None:
raise ValueError("Both `seed` and `generator` were provided. Please specify just one of them.")
if generator is not None and not isinstance(generator, np.random.Generator):
raise ValueError("The provided generator must be an instance of numpy.random.Generator")
load_from_cache_file = load_from_cache_file if load_from_cache_file is not None else is_caching_enabled()
if generator is None:
if seed is None:
_, seed, pos, *_ = np.random.get_state()
seed = seed[pos] if pos < 624 else seed[0]
_ = np.random.random() # do 1 step of rng
generator = np.random.default_rng(seed)
# Check if we've already cached this computation (indexed by a hash)
if self.cache_files:
if indices_cache_file_name is None:
# we create a unique hash from the function, current dataset file and the mapping args
indices_cache_file_name = self._get_cache_file_path(new_fingerprint)
if os.path.exists(indices_cache_file_name) and load_from_cache_file:
logger.info(f"Loading cached shuffled indices for dataset at {indices_cache_file_name}")
return self._new_dataset_with_indices(
fingerprint=new_fingerprint, indices_cache_file_name=indices_cache_file_name
)
permutation = generator.permutation(len(self))
return self.select(
indices=permutation,
keep_in_memory=keep_in_memory,
indices_cache_file_name=indices_cache_file_name if not keep_in_memory else None,
writer_batch_size=writer_batch_size,
new_fingerprint=new_fingerprint,
)
@transmit_format
@fingerprint_transform(
inplace=False,
randomized_function=True,
fingerprint_names=["train_new_fingerprint", "test_new_fingerprint"],
ignore_kwargs=["load_from_cache_file", "train_indices_cache_file_name", "test_indices_cache_file_name"],
)
def train_test_split(
self,
test_size: Union[float, int, None] = None,
train_size: Union[float, int, None] = None,
shuffle: bool = True,
stratify_by_column: Optional[str] = None,
seed: Optional[int] = None,
generator: Optional[np.random.Generator] = None,
keep_in_memory: bool = False,
load_from_cache_file: Optional[bool] = None,
train_indices_cache_file_name: Optional[str] = None,
test_indices_cache_file_name: Optional[str] = None,
writer_batch_size: Optional[int] = 1000,
train_new_fingerprint: Optional[str] = None,
test_new_fingerprint: Optional[str] = None,
) -> "DatasetDict":
"""Return a dictionary ([`datasets.DatasetDict`]) with two random train and test subsets (`train` and `test` `Dataset` splits).
Splits are created from the dataset according to `test_size`, `train_size` and `shuffle`.
This method is similar to scikit-learn `train_test_split`.
Args:
test_size (`numpy.random.Generator`, *optional*):
Size of the test split
If `float`, should be between `0.0` and `1.0` and represent the proportion of the dataset to include in the test split.
If `int`, represents the absolute number of test samples.
If `None`, the value is set to the complement of the train size.
If `train_size` is also `None`, it will be set to `0.25`.
train_size (`numpy.random.Generator`, *optional*):
Size of the train split
If `float`, should be between `0.0` and `1.0` and represent the proportion of the dataset to include in the train split.
If `int`, represents the absolute number of train samples.
If `None`, the value is automatically set to the complement of the test size.
shuffle (`bool`, *optional*, defaults to `True`):
Whether or not to shuffle the data before splitting.
stratify_by_column (`str`, *optional*, defaults to `None`):
The column name of labels to be used to perform stratified split of data.
seed (`int`, *optional*):
A seed to initialize the default BitGenerator if `generator=None`.
If `None`, then fresh, unpredictable entropy will be pulled from the OS.
If an `int` or `array_like[ints]` is passed, then it will be passed to SeedSequence to derive the initial BitGenerator state.
generator (`numpy.random.Generator`, *optional*):
Numpy random Generator to use to compute the permutation of the dataset rows.
If `generator=None` (default), uses `np.random.default_rng` (the default BitGenerator (PCG64) of NumPy).
keep_in_memory (`bool`, defaults to `False`):
Keep the splits indices in memory instead of writing it to a cache file.
load_from_cache_file (`Optional[bool]`, defaults to `True` if caching is enabled):
If a cache file storing the splits indices
can be identified, use it instead of recomputing.
train_cache_file_name (`str`, *optional*):
Provide the name of a path for the cache file. It is used to store the
train split indices instead of the automatically generated cache file name.
test_cache_file_name (`str`, *optional*):
Provide the name of a path for the cache file. It is used to store the
test split indices instead of the automatically generated cache file name.
writer_batch_size (`int`, defaults to `1000`):
Number of rows per write operation for the cache file writer.
This value is a good trade-off between memory usage during the processing, and processing speed.
Higher value makes the processing do fewer lookups, lower value consume less temporary memory while running `map`.
train_new_fingerprint (`str`, *optional*, defaults to `None`):
The new fingerprint of the train set after transform.
If `None`, the new fingerprint is computed using a hash of the previous fingerprint, and the transform arguments
test_new_fingerprint (`str`, *optional*, defaults to `None`):
The new fingerprint of the test set after transform.
If `None`, the new fingerprint is computed using a hash of the previous fingerprint, and the transform arguments
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", split="validation")
>>> ds = ds.train_test_split(test_size=0.2, shuffle=True)
DatasetDict({
train: Dataset({
features: ['text', 'label'],
num_rows: 852
})
test: Dataset({
features: ['text', 'label'],
num_rows: 214
})
})
# set a seed
>>> ds = ds.train_test_split(test_size=0.2, seed=42)
# stratified split
>>> ds = load_dataset("imdb",split="train")
Dataset({
features: ['text', 'label'],
num_rows: 25000
})
>>> ds = ds.train_test_split(test_size=0.2, stratify_by_column="label")
DatasetDict({
train: Dataset({
features: ['text', 'label'],
num_rows: 20000
})
test: Dataset({
features: ['text', 'label'],
num_rows: 5000
})
})
```
"""
from .dataset_dict import DatasetDict # import here because of circular dependency
if len(self.list_indexes()) > 0:
raise DatasetTransformationNotAllowedError(
"Using `.train_test_split` on a dataset with attached indexes is not allowed. You can first run `.drop_index() to remove your index and then re-add it."
)
# If the array is empty we do nothing
if len(self) == 0:
return DatasetDict({"train": self, "test": self})
if test_size is None and train_size is None:
test_size = 0.25
# Safety checks similar to scikit-learn's ones.
# (adapted from https://github.com/scikit-learn/scikit-learn/blob/fd237278e895b42abe8d8d09105cbb82dc2cbba7/sklearn/model_selection/_split.py#L1750)
n_samples = len(self)
if (
isinstance(test_size, int)
and (test_size >= n_samples or test_size <= 0)
or isinstance(test_size, float)
and (test_size <= 0 or test_size >= 1)
):
raise ValueError(
f"test_size={test_size} should be either positive and smaller "
f"than the number of samples {n_samples} or a float in the (0, 1) range"
)
if (
isinstance(train_size, int)
and (train_size >= n_samples or train_size <= 0)
or isinstance(train_size, float)
and (train_size <= 0 or train_size >= 1)
):
raise ValueError(
f"train_size={train_size} should be either positive and smaller "
f"than the number of samples {n_samples} or a float in the (0, 1) range"
)
if train_size is not None and not isinstance(train_size, (int, float)):
raise ValueError(f"Invalid value for train_size: {train_size} of type {type(train_size)}")
if test_size is not None and not isinstance(test_size, (int, float)):
raise ValueError(f"Invalid value for test_size: {test_size} of type {type(test_size)}")
if isinstance(train_size, float) and isinstance(test_size, float) and train_size + test_size > 1:
raise ValueError(
f"The sum of test_size and train_size = {train_size + test_size}, should be in the (0, 1)"
" range. Reduce test_size and/or train_size."
)
if isinstance(test_size, float):
n_test = ceil(test_size * n_samples)
elif isinstance(test_size, int):
n_test = float(test_size)
if isinstance(train_size, float):
n_train = floor(train_size * n_samples)
elif isinstance(train_size, int):
n_train = float(train_size)
if train_size is None:
n_train = n_samples - n_test
elif test_size is None:
n_test = n_samples - n_train
if n_train + n_test > n_samples:
raise ValueError(
f"The sum of train_size and test_size = {n_train + n_test}, "
"should be smaller than the number of "
f"samples {n_samples}. Reduce test_size and/or "
"train_size."
)
n_train, n_test = int(n_train), int(n_test)
if n_train == 0:
raise ValueError(
f"With n_samples={n_samples}, test_size={test_size} and train_size={train_size}, the "
"resulting train set will be empty. Adjust any of the "
"aforementioned parameters."
)
load_from_cache_file = load_from_cache_file if load_from_cache_file is not None else is_caching_enabled()
if generator is None and shuffle is True:
if seed is None:
_, seed, pos, *_ = np.random.get_state()
seed = seed[pos] if pos < 624 else seed[0]
_ = np.random.random() # do 1 step of rng
generator = np.random.default_rng(seed)
# Check if we've already cached this computation (indexed by a hash)
if self.cache_files:
if train_indices_cache_file_name is None or test_indices_cache_file_name is None:
# we create a unique hash from the function, current dataset file and the mapping args
if train_indices_cache_file_name is None:
train_indices_cache_file_name = self._get_cache_file_path(train_new_fingerprint)
if test_indices_cache_file_name is None:
test_indices_cache_file_name = self._get_cache_file_path(test_new_fingerprint)
if (
os.path.exists(train_indices_cache_file_name)
and os.path.exists(test_indices_cache_file_name)
and load_from_cache_file
):
logger.info(
f"Loading cached split indices for dataset at {train_indices_cache_file_name} and {test_indices_cache_file_name}"
)
return DatasetDict(
{
"train": self._new_dataset_with_indices(
fingerprint=train_new_fingerprint, indices_cache_file_name=train_indices_cache_file_name
),
"test": self._new_dataset_with_indices(
fingerprint=test_new_fingerprint, indices_cache_file_name=test_indices_cache_file_name
),
}
)
if not shuffle:
if stratify_by_column is not None:
raise ValueError("Stratified train/test split is not implemented for `shuffle=False`")
train_indices = np.arange(n_train)
test_indices = np.arange(n_train, n_train + n_test)
else:
# stratified partition
if stratify_by_column is not None:
if stratify_by_column not in self._info.features.keys():
raise ValueError(f"Key {stratify_by_column} not found in {self._info.features.keys()}")
if not isinstance(self._info.features[stratify_by_column], ClassLabel):
raise ValueError(
f"Stratifying by column is only supported for {ClassLabel.__name__} column, and column {stratify_by_column} is {type(self._info.features[stratify_by_column]).__name__}."
)
try:
train_indices, test_indices = next(
stratified_shuffle_split_generate_indices(
self.with_format("numpy")[stratify_by_column], n_train, n_test, rng=generator
)
)
except Exception as error:
if str(error) == "Minimum class count error":
raise ValueError(
f"The least populated class in {stratify_by_column} column has only 1"
" member, which is too few. The minimum"
" number of groups for any class cannot"
" be less than 2."
)
else:
raise error
# random partition
else:
permutation = generator.permutation(len(self))
test_indices = permutation[:n_test]
train_indices = permutation[n_test : (n_test + n_train)]
train_split = self.select(
indices=train_indices,
keep_in_memory=keep_in_memory,
indices_cache_file_name=train_indices_cache_file_name,
writer_batch_size=writer_batch_size,
new_fingerprint=train_new_fingerprint,
)
test_split = self.select(
indices=test_indices,
keep_in_memory=keep_in_memory,
indices_cache_file_name=test_indices_cache_file_name,
writer_batch_size=writer_batch_size,
new_fingerprint=test_new_fingerprint,
)
return DatasetDict({"train": train_split, "test": test_split})
def shard(
self,
num_shards: int,
index: int,
contiguous: bool = False,
keep_in_memory: bool = False,
indices_cache_file_name: Optional[str] = None,
writer_batch_size: Optional[int] = 1000,
) -> "Dataset":
"""Return the `index`-nth shard from dataset split into `num_shards` pieces.
This shards deterministically. `dset.shard(n, i)` will contain all elements of dset whose
index mod `n = i`.
`dset.shard(n, i, contiguous=True)` will instead split dset into contiguous chunks,
so it can be easily concatenated back together after processing. If `n % i == l`, then the
first `l` shards will have length `(n // i) + 1`, and the remaining shards will have length `(n // i)`.
`datasets.concatenate([dset.shard(n, i, contiguous=True) for i in range(n)])` will return
a dataset with the same order as the original.
Be sure to shard before using any randomizing operator (such as `shuffle`).
It is best if the shard operator is used early in the dataset pipeline.
Args:
num_shards (`int`):
How many shards to split the dataset into.
index (`int`):
Which shard to select and return.
contiguous: (`bool`, defaults to `False`):
Whether to select contiguous blocks of indices for shards.
keep_in_memory (`bool`, defaults to `False`):
Keep the dataset in memory instead of writing it to a cache file.
indices_cache_file_name (`str`, *optional*):
Provide the name of a path for the cache file. It is used to store the
indices of each shard instead of the automatically generated cache file name.
writer_batch_size (`int`, defaults to `1000`):
Number of rows per write operation for the cache file writer.
This value is a good trade-off between memory usage during the processing, and processing speed.
Higher value makes the processing do fewer lookups, lower value consume less temporary memory while running `map`.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", split="validation")
>>> ds
Dataset({
features: ['text', 'label'],
num_rows: 1066
})
>>> ds.shard(num_shards=2, index=0)
Dataset({
features: ['text', 'label'],
num_rows: 533
})
```
"""
if not 0 <= index < num_shards:
raise ValueError("index should be in [0, num_shards-1]")
if contiguous:
div = len(self) // num_shards
mod = len(self) % num_shards
start = div * index + min(index, mod)
end = start + div + (1 if index < mod else 0)
indices = range(start, end)
else:
indices = np.arange(index, len(self), num_shards)
return self.select(
indices=indices,
keep_in_memory=keep_in_memory,
indices_cache_file_name=indices_cache_file_name,
writer_batch_size=writer_batch_size,
)
@deprecated()
def export(
self,
filename: str,
format: str = "tfrecord",
):
"""Writes the Arrow dataset to a TFRecord file.
The dataset must already be in tensorflow format. The records will be written with
keys from `dataset._format_columns`.
Args:
filename (`str`): The filename, including the `.tfrecord` extension, to write to.
format (`str`, optional, default `"tfrecord"`): The type of output file. Currently this is a no-op, as
TFRecords are the only option. This enables a more flexible function signature later.
"""
try:
import tensorflow as tf # noqa: F401
except ImportError:
logger.error("Tensorflow needs to be installed to be able to return Tensorflow tensors.")
# From https://www.tensorflow.org/tutorials/load_data/tfrecord
def _bytes_feature(values):
"""Returns a bytes_list from a list of string / byte."""
return tf.train.Feature(bytes_list=tf.train.BytesList(value=values))
def _float_feature(values):
"""Returns a float_list from a list of float / double."""
return tf.train.Feature(float_list=tf.train.FloatList(value=values))
def _int64_feature(values):
"""Returns an int64_list from a list of bool / enum / int / uint."""
return tf.train.Feature(int64_list=tf.train.Int64List(value=values))
def _feature(values: Union[float, int, str, np.ndarray, list]) -> "tf.train.Feature":
"""Typechecks `values` and returns the corresponding tf.train.Feature."""
if isinstance(values, list):
if values and isinstance(values[0], str):
return _bytes_feature([v.encode() for v in values])
else:
raise ValueError(f"values={values} is empty or contains items that cannot be serialized")
elif isinstance(values, np.ndarray):
if values.dtype == np.dtype(float):
return _float_feature(values)
elif values.dtype == np.int64:
return _int64_feature(values)
elif values.dtype == np.dtype(str) or (
values.dtype == np.dtype(object) and len(values) > 0 and isinstance(values[0], str)
):
return _bytes_feature([v.encode() for v in values])
else:
raise ValueError(
f"values={values} is empty or is an np.ndarray with items of dtype {values[0].dtype}, which cannot be serialized"
)
elif hasattr(values, "dtype"):
if np.issubdtype(values.dtype, np.floating):
return _float_feature([values.item()])
elif np.issubdtype(values.dtype, np.integer):
return _int64_feature([values.item()])
elif np.issubdtype(values.dtype, str):
return _bytes_feature([values.item().encode()])
else:
raise ValueError(f"values={values} has dtype {values.dtype}, which cannot be serialized")
else:
raise ValueError(f"values={values} are not numpy objects or strings, and so cannot be serialized")
def serialize_example(ex):
feature = {key: _feature(value) for key, value in ex.items()}
example_proto = tf.train.Example(features=tf.train.Features(feature=feature))
return example_proto.SerializeToString()
def tf_serialize_example(ex):
tf_string = tf.py_function(serialize_example, (ex,), tf.string)
return tf.reshape(tf_string, ())
def generator():
for ex in self:
yield serialize_example(ex)
if self._format_type != "numpy":
raise ValueError("Dataset format must be numpy before exporting")
if not filename.endswith(".tfrecord"):
raise ValueError("filename {filename} must end with .tfrecord")
tf_dataset = tf.data.Dataset.from_generator(generator, output_types=tf.string, output_shapes=())
writer = tf.data.experimental.TFRecordWriter(filename)
logger.info(f"Writing TFRecord to {filename}")
writer.write(tf_dataset)
logger.info(f"Finished writing TFRecord to {filename}")
self = None # delete the dataset reference used by tf_dataset
def to_csv(
self,
path_or_buf: Union[PathLike, BinaryIO],
batch_size: Optional[int] = None,
num_proc: Optional[int] = None,
**to_csv_kwargs,
) -> int:
"""Exports the dataset to csv
Args:
path_or_buf (`PathLike` or `FileOrBuffer`):
Either a path to a file or a BinaryIO.
batch_size (`int`, *optional*):
Size of the batch to load in memory and write at once.
Defaults to `datasets.config.DEFAULT_MAX_BATCH_SIZE`.
num_proc (`int`, *optional*):
Number of processes for multiprocessing. By default it doesn't
use multiprocessing. `batch_size` in this case defaults to
`datasets.config.DEFAULT_MAX_BATCH_SIZE` but feel free to make it 5x or 10x of the default
value if you have sufficient compute power.
**to_csv_kwargs (additional keyword arguments):
Parameters to pass to pandas's [`pandas.DataFrame.to_csv`](https://pandas.pydata.org/docs/reference/api/pandas.DataFrame.to_json.html).
<Changed version="2.10.0">
Now, `index` defaults to `False` if not specified.
If you would like to write the index, pass `index=True` and also set a name for the index column by
passing `index_label`.
</Changed>
Returns:
`int`: The number of characters or bytes written.
Example:
```py
>>> ds.to_csv("path/to/dataset/directory")
```
"""
# Dynamic import to avoid circular dependency
from .io.csv import CsvDatasetWriter
return CsvDatasetWriter(self, path_or_buf, batch_size=batch_size, num_proc=num_proc, **to_csv_kwargs).write()
def to_dict(self, batch_size: Optional[int] = None, batched="deprecated") -> Union[dict, Iterator[dict]]:
"""Returns the dataset as a Python dict. Can also return a generator for large datasets.
Args:
batched (`bool`):
Set to `True` to return a generator that yields the dataset as batches
of `batch_size` rows. Defaults to `False` (returns the whole datasets once).
<Deprecated version="2.11.0">
Use `.iter(batch_size=batch_size)` followed by `.to_dict()` on the individual batches instead.
</Deprecated>
batch_size (`int`, *optional*): The size (number of rows) of the batches if `batched` is `True`.
Defaults to `datasets.config.DEFAULT_MAX_BATCH_SIZE`.
Returns:
`dict` or `Iterator[dict]`
Example:
```py
>>> ds.to_dict()
```
"""
if batched != "deprecated":
warnings.warn(
"'batched' was deprecated in version 2.11.0 and will be removed in version 3.0.0. Use `.iter(batch_size=batch_size)` followed by `.to_dict()` on the individual batches instead.",
FutureWarning,
)
else:
batched = False
if not batched:
return query_table(
table=self._data,
key=slice(0, len(self)),
indices=self._indices if self._indices is not None else None,
).to_pydict()
else:
batch_size = batch_size if batch_size else config.DEFAULT_MAX_BATCH_SIZE
return (
query_table(
table=self._data,
key=slice(offset, offset + batch_size),
indices=self._indices if self._indices is not None else None,
).to_pydict()
for offset in range(0, len(self), batch_size)
)
def to_list(self) -> list:
"""Returns the dataset as a Python list.
Returns:
`list`
Example:
```py
>>> ds.to_list()
```
"""
return query_table(
table=self._data,
key=slice(0, len(self)),
indices=self._indices if self._indices is not None else None,
).to_pylist()
def to_json(
self,
path_or_buf: Union[PathLike, BinaryIO],
batch_size: Optional[int] = None,
num_proc: Optional[int] = None,
**to_json_kwargs,
) -> int:
"""Export the dataset to JSON Lines or JSON.
Args:
path_or_buf (`PathLike` or `FileOrBuffer`):
Either a path to a file or a BinaryIO.
batch_size (`int`, *optional*):
Size of the batch to load in memory and write at once.
Defaults to `datasets.config.DEFAULT_MAX_BATCH_SIZE`.
num_proc (`int`, *optional*):
Number of processes for multiprocessing. By default it doesn't
use multiprocessing. `batch_size` in this case defaults to
`datasets.config.DEFAULT_MAX_BATCH_SIZE` but feel free to make it 5x or 10x of the default
value if you have sufficient compute power.
**to_json_kwargs (additional keyword arguments):
Parameters to pass to pandas's [`pandas.DataFrame.to_json`](https://pandas.pydata.org/docs/reference/api/pandas.DataFrame.to_json.html).
<Changed version="2.11.0">
Now, `index` defaults to `False` if `orient` is `"split"` or `"table"`.
If you would like to write the index, pass `index=True`.
</Changed>
Returns:
`int`: The number of characters or bytes written.
Example:
```py
>>> ds.to_json("path/to/dataset/directory")
```
"""
# Dynamic import to avoid circular dependency
from .io.json import JsonDatasetWriter
return JsonDatasetWriter(self, path_or_buf, batch_size=batch_size, num_proc=num_proc, **to_json_kwargs).write()
def to_pandas(
self, batch_size: Optional[int] = None, batched: bool = False
) -> Union[pd.DataFrame, Iterator[pd.DataFrame]]:
"""Returns the dataset as a `pandas.DataFrame`. Can also return a generator for large datasets.
Args:
batched (`bool`):
Set to `True` to return a generator that yields the dataset as batches
of `batch_size` rows. Defaults to `False` (returns the whole datasets once).
batch_size (`int`, *optional*):
The size (number of rows) of the batches if `batched` is `True`.
Defaults to `datasets.config.DEFAULT_MAX_BATCH_SIZE`.
Returns:
`pandas.DataFrame` or `Iterator[pandas.DataFrame]`
Example:
```py
>>> ds.to_pandas()
```
"""
if not batched:
return query_table(
table=self._data,
key=slice(0, len(self)),
indices=self._indices if self._indices is not None else None,
).to_pandas(types_mapper=pandas_types_mapper)
else:
batch_size = batch_size if batch_size else config.DEFAULT_MAX_BATCH_SIZE
return (
query_table(
table=self._data,
key=slice(offset, offset + batch_size),
indices=self._indices if self._indices is not None else None,
).to_pandas(types_mapper=pandas_types_mapper)
for offset in range(0, len(self), batch_size)
)
def to_parquet(
self,
path_or_buf: Union[PathLike, BinaryIO],
batch_size: Optional[int] = None,
**parquet_writer_kwargs,
) -> int:
"""Exports the dataset to parquet
Args:
path_or_buf (`PathLike` or `FileOrBuffer`):
Either a path to a file or a BinaryIO.
batch_size (`int`, *optional*):
Size of the batch to load in memory and write at once.
Defaults to `datasets.config.DEFAULT_MAX_BATCH_SIZE`.
**parquet_writer_kwargs (additional keyword arguments):
Parameters to pass to PyArrow's `pyarrow.parquet.ParquetWriter`.
Returns:
`int`: The number of characters or bytes written.
Example:
```py
>>> ds.to_parquet("path/to/dataset/directory")
```
"""
# Dynamic import to avoid circular dependency
from .io.parquet import ParquetDatasetWriter
return ParquetDatasetWriter(self, path_or_buf, batch_size=batch_size, **parquet_writer_kwargs).write()
def to_sql(
self,
name: str,
con: Union[str, "sqlalchemy.engine.Connection", "sqlalchemy.engine.Engine", "sqlite3.Connection"],
batch_size: Optional[int] = None,
**sql_writer_kwargs,
) -> int:
"""Exports the dataset to a SQL database.
Args:
name (`str`):
Name of SQL table.
con (`str` or `sqlite3.Connection` or `sqlalchemy.engine.Connection` or `sqlalchemy.engine.Connection`):
A [URI string](https://docs.sqlalchemy.org/en/13/core/engines.html#database-urls) or a SQLite3/SQLAlchemy connection object used to write to a database.
batch_size (`int`, *optional*):
Size of the batch to load in memory and write at once.
Defaults to `datasets.config.DEFAULT_MAX_BATCH_SIZE`.
**sql_writer_kwargs (additional keyword arguments):
Parameters to pass to pandas's [`pandas.DataFrame.to_sql`](https://pandas.pydata.org/docs/reference/api/pandas.DataFrame.to_sql.html).
<Changed version="2.11.0">
Now, `index` defaults to `False` if not specified.
If you would like to write the index, pass `index=True` and also set a name for the index column by
passing `index_label`.
</Changed>
Returns:
`int`: The number of records written.
Example:
```py
>>> # con provided as a connection URI string
>>> ds.to_sql("data", "sqlite:///my_own_db.sql")
>>> # con provided as a sqlite3 connection object
>>> import sqlite3
>>> con = sqlite3.connect("my_own_db.sql")
>>> with con:
... ds.to_sql("data", con)
```
"""
# Dynamic import to avoid circular dependency
from .io.sql import SqlDatasetWriter
return SqlDatasetWriter(self, name, con, batch_size=batch_size, **sql_writer_kwargs).write()
def _estimate_nbytes(self) -> int:
dataset_nbytes = self.data.nbytes
# Find decodable columns, because if there are any, we need to
# adjust the dataset size computation (needed for sharding) to account for possible external files
decodable_columns = [
k for k, v in self._info.features.items() if require_decoding(v, ignore_decode_attribute=True)
]
if decodable_columns:
# Approximate the space needed to store the bytes from the external files by analyzing the first 1000 examples
extra_nbytes = 0
def extra_nbytes_visitor(array, feature):
nonlocal extra_nbytes
if isinstance(feature, (Audio, Image)):
for x in array.to_pylist():
if x is not None and x["bytes"] is None and x["path"] is not None:
size = xgetsize(x["path"])
extra_nbytes += size
extra_nbytes -= array.field("path").nbytes
table = self.with_format("arrow")[:1000]
table_visitor(table, extra_nbytes_visitor)
extra_nbytes = extra_nbytes * len(self.data) / len(table)
dataset_nbytes = dataset_nbytes + extra_nbytes
if self._indices is not None:
dataset_nbytes = dataset_nbytes * len(self._indices) / len(self.data)
return dataset_nbytes
@staticmethod
def _generate_tables_from_shards(shards: List["Dataset"], batch_size: int):
for shard_idx, shard in enumerate(shards):
for pa_table in shard.with_format("arrow").iter(batch_size):
yield shard_idx, pa_table
@staticmethod
def _generate_tables_from_cache_file(filename: str):
for batch_idx, batch in enumerate(_memory_mapped_record_batch_reader_from_file(filename)):
yield batch_idx, pa.Table.from_batches([batch])
def to_iterable_dataset(self, num_shards: Optional[int] = 1) -> "IterableDataset":
"""Get an [`datasets.IterableDataset`] from a map-style [`datasets.Dataset`].
This is equivalent to loading a dataset in streaming mode with [`datasets.load_dataset`], but much faster since the data is streamed from local files.
Contrary to map-style datasets, iterable datasets are lazy and can only be iterated over (e.g. using a for loop).
Since they are read sequentially in training loops, iterable datasets are much faster than map-style datasets.
All the transformations applied to iterable datasets like filtering or processing are done on-the-fly when you start iterating over the dataset.
Still, it is possible to shuffle an iterable dataset using [`datasets.IterableDataset.shuffle`].
This is a fast approximate shuffling that works best if you have multiple shards and if you specify a buffer size that is big enough.
To get the best speed performance, make sure your dataset doesn't have an indices mapping.
If this is the case, the data are not read contiguously, which can be slow sometimes.
You can use `ds = ds.flatten_indices()` to write your dataset in contiguous chunks of data and have optimal speed before switching to an iterable dataset.
Args:
num_shards (`int`, default to `1`):
Number of shards to define when instantiating the iterable dataset. This is especially useful for big datasets to be able to shuffle properly,
and also to enable fast parallel loading using a PyTorch DataLoader or in distributed setups for example.
Shards are defined using [`datasets.Dataset.shard`]: it simply slices the data without writing anything on disk.
Returns:
[`datasets.IterableDataset`]
Example:
Basic usage:
```python
>>> ids = ds.to_iterable_dataset()
>>> for example in ids:
... pass
```
With lazy filtering and processing:
```python
>>> ids = ds.to_iterable_dataset()
>>> ids = ids.filter(filter_fn).map(process_fn) # will filter and process on-the-fly when you start iterating over the iterable dataset
>>> for example in ids:
... pass
```
With sharding to enable efficient shuffling:
```python
>>> ids = ds.to_iterable_dataset(num_shards=64) # the dataset is split into 64 shards to be iterated over
>>> ids = ids.shuffle(buffer_size=10_000) # will shuffle the shards order and use a shuffle buffer for fast approximate shuffling when you start iterating
>>> for example in ids:
... pass
```
With a PyTorch DataLoader:
```python
>>> import torch
>>> ids = ds.to_iterable_dataset(num_shards=64)
>>> ids = ids.filter(filter_fn).map(process_fn)
>>> dataloader = torch.utils.data.DataLoader(ids, num_workers=4) # will assign 64 / 4 = 16 shards to each worker to load, filter and process when you start iterating
>>> for example in ids:
... pass
```
With a PyTorch DataLoader and shuffling:
```python
>>> import torch
>>> ids = ds.to_iterable_dataset(num_shards=64)
>>> ids = ids.shuffle(buffer_size=10_000) # will shuffle the shards order and use a shuffle buffer when you start iterating
>>> dataloader = torch.utils.data.DataLoader(ids, num_workers=4) # will assign 64 / 4 = 16 shards from the shuffled list of shards to each worker when you start iterating
>>> for example in ids:
... pass
```
In a distributed setup like PyTorch DDP with a PyTorch DataLoader and shuffling
```python
>>> from datasets.distributed import split_dataset_by_node
>>> ids = ds.to_iterable_dataset(num_shards=512)
>>> ids = ids.shuffle(buffer_size=10_000) # will shuffle the shards order and use a shuffle buffer when you start iterating
>>> ids = split_dataset_by_node(ds, world_size=8, rank=0) # will keep only 512 / 8 = 64 shards from the shuffled lists of shards when you start iterating
>>> dataloader = torch.utils.data.DataLoader(ids, num_workers=4) # will assign 64 / 4 = 16 shards from this node's list of shards to each worker when you start iterating
>>> for example in ids:
... pass
```
With shuffling and multiple epochs:
```python
>>> ids = ds.to_iterable_dataset(num_shards=64)
>>> ids = ids.shuffle(buffer_size=10_000, seed=42) # will shuffle the shards order and use a shuffle buffer when you start iterating
>>> for epoch in range(n_epochs):
... ids.set_epoch(epoch) # will use effective_seed = seed + epoch to shuffle the shards and for the shuffle buffer when you start iterating
... for example in ids:
... pass
```
Feel free to also use [`IterableDataset.set_epoch`] when using a PyTorch DataLoader or in distributed setups.
"""
from .iterable_dataset import ArrowExamplesIterable, IterableDataset
if self._format_type is not None:
raise NotImplementedError(
"Converting a formatted dataset to a formatted iterable dataset is not implemented yet. Please run `my_dataset = my_dataset.with_format(None)` before calling to_iterable_dataset"
)
if num_shards > len(self):
raise ValueError(
f"Unable to shard a dataset of size {len(self)} into {num_shards} shards (the number of shards exceeds the number of samples)."
)
if self._indices is not None:
logger.info(
"Converting an Arrow dataset to iterable but it has an indices mapping that can make it slower. "
"You can use `ds = ds.flatten_indices()` to write your dataset in contiguous chunks of data and have optimal speed."
)
shards = (
[copy.deepcopy(self)]
if num_shards == 1
else [
self.shard(num_shards=num_shards, index=shard_idx, contiguous=True) for shard_idx in range(num_shards)
]
)
ex_iterable = ArrowExamplesIterable(
Dataset._generate_tables_from_shards,
kwargs={"shards": shards, "batch_size": config.DEFAULT_MAX_BATCH_SIZE},
)
return IterableDataset(ex_iterable, info=DatasetInfo(features=self.features))
def _push_parquet_shards_to_hub(
self,
repo_id: str,
data_dir: str = "data",
split: Optional[str] = None,
token: Optional[str] = None,
revision: Optional[str] = None,
create_pr: Optional[bool] = False,
max_shard_size: Optional[Union[int, str]] = None,
num_shards: Optional[int] = None,
embed_external_files: bool = True,
) -> Tuple[str, str, int, int, List[str], int]:
"""Pushes the dataset shards as Parquet files to the hub.
Returns:
additions (`List[CommitOperation]`): list of the `CommitOperationAdd` of the uploaded shards
uploaded_size (`int`): number of uploaded bytes to the repository
dataset_nbytes (`int`): approximate size in bytes of the uploaded dataset afer uncompression
"""
# Find decodable columns, because if there are any, we need to:
# embed the bytes from the files in the shards
decodable_columns = (
[k for k, v in self._info.features.items() if require_decoding(v, ignore_decode_attribute=True)]
if embed_external_files
else []
)
dataset_nbytes = self._estimate_nbytes()
if num_shards is None:
max_shard_size = convert_file_size_to_int(max_shard_size or config.MAX_SHARD_SIZE)
num_shards = int(dataset_nbytes / max_shard_size) + 1
num_shards = max(num_shards, 1)
shards = (self.shard(num_shards=num_shards, index=i, contiguous=True) for i in range(num_shards))
if decodable_columns:
def shards_with_embedded_external_files(shards):
for shard in shards:
format = shard.format
shard = shard.with_format("arrow")
shard = shard.map(
embed_table_storage,
batched=True,
batch_size=1000,
keep_in_memory=True,
)
shard = shard.with_format(**format)
yield shard
shards = shards_with_embedded_external_files(shards)
api = HfApi(endpoint=config.HF_ENDPOINT, token=token)
uploaded_size = 0
additions = []
for index, shard in hf_tqdm(
enumerate(shards),
desc="Uploading the dataset shards",
total=num_shards,
):
shard_path_in_repo = f"{data_dir}/{split}-{index:05d}-of-{num_shards:05d}.parquet"
buffer = BytesIO()
shard.to_parquet(buffer)
uploaded_size += buffer.tell()
shard_addition = CommitOperationAdd(path_in_repo=shard_path_in_repo, path_or_fileobj=buffer)
_retry(
api.preupload_lfs_files,
func_kwargs={
"repo_id": repo_id,
"additions": [shard_addition],
"token": token,
"repo_type": "dataset",
"revision": revision,
"create_pr": create_pr,
},
exceptions=HTTPError,
status_codes=[504],
base_wait_time=2.0,
max_retries=5,
max_wait_time=20.0,
)
additions.append(shard_addition)
return additions, uploaded_size, dataset_nbytes
def push_to_hub(
self,
repo_id: str,
config_name: str = "default",
split: Optional[str] = None,
commit_message: Optional[str] = None,
private: Optional[bool] = False,
token: Optional[str] = None,
revision: Optional[str] = None,
branch="deprecated",
create_pr: Optional[bool] = False,
max_shard_size: Optional[Union[int, str]] = None,
num_shards: Optional[int] = None,
embed_external_files: bool = True,
):
"""Pushes the dataset to the hub as a Parquet dataset.
The dataset is pushed using HTTP requests and does not need to have neither git or git-lfs installed.
The resulting Parquet files are self-contained by default. If your dataset contains [`Image`] or [`Audio`]
data, the Parquet files will store the bytes of your images or audio files.
You can disable this by setting `embed_external_files` to `False`.
Args:
repo_id (`str`):
The ID of the repository to push to in the following format: `<user>/<dataset_name>` or
`<org>/<dataset_name>`. Also accepts `<dataset_name>`, which will default to the namespace
of the logged-in user.
config_name (`str`, defaults to "default"):
The configuration name (or subset) of a dataset. Defaults to "default".
split (`str`, *optional*):
The name of the split that will be given to that dataset. Defaults to `self.split`.
commit_message (`str`, *optional*):
Message to commit while pushing. Will default to `"Upload dataset"`.
private (`bool`, *optional*, defaults to `False`):
Whether the dataset repository should be set to private or not. Only affects repository creation:
a repository that already exists will not be affected by that parameter.
token (`str`, *optional*):
An optional authentication token for the Hugging Face Hub. If no token is passed, will default
to the token saved locally when logging in with `huggingface-cli login`. Will raise an error
if no token is passed and the user is not logged-in.
revision (`str`, *optional*):
Branch to push the uploaded files to. Defaults to the `"main"` branch.
<Added version="2.15.0"/>
branch (`str`, *optional*):
The git branch on which to push the dataset. This defaults to the default branch as specified
in your repository, which defaults to `"main"`.
<Deprecated version="2.15.0">
`branch` was deprecated in favor of `revision` in version 2.15.0 and will be removed in 3.0.0.
</Deprecated>
create_pr (`bool`, *optional*, defaults to `False`):
Whether or not to create a PR with the uploaded files or directly commit.
<Added version="2.15.0"/>
max_shard_size (`int` or `str`, *optional*, defaults to `"500MB"`):
The maximum size of the dataset shards to be uploaded to the hub. If expressed as a string, needs to be digits followed by
a unit (like `"5MB"`).
num_shards (`int`, *optional*): Number of shards to write. By default the number of shards depends on `max_shard_size`.
<Added version="2.8.0"/>
embed_external_files (`bool`, defaults to `True`):
Whether to embed file bytes in the shards.
In particular, this will do the following before the push for the fields of type:
- [`Audio`] and [`Image`]: remove local path information and embed file content in the Parquet files.
Example:
```python
>>> dataset.push_to_hub("<organization>/<dataset_id>")
>>> dataset_dict.push_to_hub("<organization>/<dataset_id>", private=True)
>>> dataset.push_to_hub("<organization>/<dataset_id>", max_shard_size="1GB")
>>> dataset.push_to_hub("<organization>/<dataset_id>", num_shards=1024)
```
If your dataset has multiple splits (e.g. train/validation/test):
```python
>>> train_dataset.push_to_hub("<organization>/<dataset_id>", split="train")
>>> val_dataset.push_to_hub("<organization>/<dataset_id>", split="validation")
>>> # later
>>> dataset = load_dataset("<organization>/<dataset_id>")
>>> train_dataset = dataset["train"]
>>> val_dataset = dataset["validation"]
```
If you want to add a new configuration (or subset) to a dataset (e.g. if the dataset has multiple tasks/versions/languages):
```python
>>> english_dataset.push_to_hub("<organization>/<dataset_id>", "en")
>>> french_dataset.push_to_hub("<organization>/<dataset_id>", "fr")
>>> # later
>>> english_dataset = load_dataset("<organization>/<dataset_id>", "en")
>>> french_dataset = load_dataset("<organization>/<dataset_id>", "fr")
```
"""
if config_name == "data":
raise ValueError("`config_name` cannot be 'data'. Please, choose another name for configuration.")
if max_shard_size is not None and num_shards is not None:
raise ValueError(
"Failed to push_to_hub: please specify either max_shard_size or num_shards, but not both."
)
if split is None:
split = str(self.split) if self.split is not None else "train"
if not re.match(_split_re, split):
raise ValueError(f"Split name should match '{_split_re}' but got '{split}'.")
if branch != "deprecated":
warnings.warn(
"'branch' was deprecated in favor of 'revision' in version 2.15.0 and will be removed in 3.0.0.\n"
f"You can remove this warning by passing 'revision={branch}' instead.",
FutureWarning,
)
revision = branch
api = HfApi(endpoint=config.HF_ENDPOINT, token=token)
repo_url = api.create_repo(
repo_id,
token=token,
repo_type="dataset",
private=private,
exist_ok=True,
)
repo_id = repo_url.repo_id
if revision is not None:
api.create_branch(repo_id, branch=revision, token=token, repo_type="dataset", exist_ok=True)
data_dir = config_name if config_name != "default" else "data" # for backward compatibility
additions, uploaded_size, dataset_nbytes = self._push_parquet_shards_to_hub(
repo_id=repo_id,
data_dir=data_dir,
split=split,
token=token,
revision=revision,
max_shard_size=max_shard_size,
num_shards=num_shards,
create_pr=create_pr,
embed_external_files=embed_external_files,
)
# Check if the repo already has a README.md and/or a dataset_infos.json to update them with the new split info (size and pattern)
# and delete old split shards (if they exist)
repo_with_dataset_card, repo_with_dataset_infos = False, False
deletions, deleted_size = [], 0
repo_splits = [] # use a list to keep the order of the splits
repo_files_to_add = [addition.path_in_repo for addition in additions]
for repo_file in api.list_files_info(repo_id, revision=revision, repo_type="dataset", token=token):
if repo_file.rfilename == "README.md":
repo_with_dataset_card = True
elif repo_file.rfilename == config.DATASETDICT_INFOS_FILENAME:
repo_with_dataset_infos = True
elif (
repo_file.rfilename.startswith(f"{data_dir}/{split}-") and repo_file.rfilename not in repo_files_to_add
):
deletions.append(CommitOperationDelete(path_in_repo=repo_file.rfilename))
deleted_size += repo_file.size
elif fnmatch.fnmatch(
repo_file.rfilename, PUSH_TO_HUB_WITHOUT_METADATA_CONFIGS_SPLIT_PATTERN_SHARDED.replace("{split}", "*")
):
repo_split = string_to_dict(
repo_file.rfilename,
glob_pattern_to_regex(PUSH_TO_HUB_WITHOUT_METADATA_CONFIGS_SPLIT_PATTERN_SHARDED),
)["split"]
if repo_split not in repo_splits:
repo_splits.append(repo_split)
organization, dataset_name = repo_id.split("/") if "/" in repo_id else (None, repo_id)
info_to_dump = self.info.copy()
info_to_dump.download_checksums = None
info_to_dump.download_size = uploaded_size
info_to_dump.dataset_size = dataset_nbytes
info_to_dump.size_in_bytes = uploaded_size + dataset_nbytes
info_to_dump.config_name = config_name
info_to_dump.splits = SplitDict(
{split: SplitInfo(split, num_bytes=dataset_nbytes, num_examples=len(self), dataset_name=dataset_name)}
)
# get the info from the README to update them
if repo_with_dataset_card:
dataset_card_path = api.hf_hub_download(repo_id, "README.md", repo_type="dataset", revision=revision)
dataset_card = DatasetCard.load(Path(dataset_card_path))
dataset_card_data = dataset_card.data
metadata_configs = MetadataConfigs.from_dataset_card_data(dataset_card_data)
dataset_infos: DatasetInfosDict = DatasetInfosDict.from_dataset_card_data(dataset_card_data)
if dataset_infos and config_name in dataset_infos:
repo_info = dataset_infos[config_name]
else:
repo_info = None
# get the deprecated dataset_infos.json to update them
elif repo_with_dataset_infos:
dataset_card = None
dataset_card_data = DatasetCardData()
metadata_configs = MetadataConfigs()
dataset_infos_path = api.hf_hub_download(
repo_id, config.DATASETDICT_INFOS_FILENAME, repo_type="dataset", revision=revision
)
with open(dataset_infos_path, encoding="utf-8") as f:
dataset_infos: dict = json.load(f)
dataset_info = dataset_infos.get(config_name, None) if dataset_infos else None
repo_info = DatasetInfo.from_dict(dataset_info) if dataset_info else None
else:
dataset_card = None
dataset_card_data = DatasetCardData()
metadata_configs = MetadataConfigs()
repo_info = None
# update the total info to dump from existing info
if repo_info is not None:
logger.info("Updating downloaded metadata with the new split.")
if repo_info.splits and list(repo_info.splits) != [split]:
if self._info.features != repo_info.features:
raise ValueError(
f"Features of the new split don't match the features of the existing splits on the hub: {self._info.features} != {repo_info.features}"
)
if split in repo_info.splits:
repo_info.download_size -= deleted_size
repo_info.dataset_size -= repo_info.splits.get(split, SplitInfo()).num_bytes or 0
repo_info.download_checksums = None
repo_info.download_size = (repo_info.download_size or 0) + uploaded_size
repo_info.dataset_size = (repo_info.dataset_size or 0) + dataset_nbytes
repo_info.size_in_bytes = repo_info.download_size + repo_info.dataset_size
repo_info.splits.pop(split, None)
repo_info.splits[split] = SplitInfo(
split, num_bytes=dataset_nbytes, num_examples=len(self), dataset_name=dataset_name
)
info_to_dump = repo_info
# create the metadata configs if it was uploaded with push_to_hub before metadata configs existed
if not metadata_configs and repo_splits:
default_metadata_configs_to_dump = {
"data_files": [{"split": split, "path": f"data/{split}-*"} for split in repo_splits]
}
MetadataConfigs({"default": default_metadata_configs_to_dump}).to_dataset_card_data(dataset_card_data)
# update the metadata configs
if config_name in metadata_configs:
metadata_config = metadata_configs[config_name]
if "data_files" in metadata_config:
data_files_to_dump = sanitize_patterns(metadata_config["data_files"])
else:
data_files_to_dump = {}
# add the new split
data_files_to_dump[split] = [f"{data_dir}/{split}-*"]
metadata_config_to_dump = {
"data_files": [
{
"split": _split,
"path": _pattern[0] if len(_pattern) == 1 else _pattern,
}
for _split, _pattern in data_files_to_dump.items()
]
}
else:
metadata_config_to_dump = {"data_files": [{"split": split, "path": f"{data_dir}/{split}-*"}]}
# push to the deprecated dataset_infos.json
if repo_with_dataset_infos:
dataset_infos_path = api.hf_hub_download(
repo_id, config.DATASETDICT_INFOS_FILENAME, repo_type="dataset", revision=revision
)
with open(dataset_infos_path, encoding="utf-8") as f:
dataset_infos: dict = json.load(f)
dataset_infos[config_name] = asdict(info_to_dump)
buffer = BytesIO()
buffer.write(json.dumps(dataset_infos, indent=4).encode("utf-8"))
additions.append(
CommitOperationAdd(path_in_repo=config.DATASETDICT_INFOS_FILENAME, path_or_fileobj=buffer)
)
# push to README
DatasetInfosDict({config_name: info_to_dump}).to_dataset_card_data(dataset_card_data)
MetadataConfigs({config_name: metadata_config_to_dump}).to_dataset_card_data(dataset_card_data)
dataset_card = DatasetCard(f"---\n{dataset_card_data}\n---\n") if dataset_card is None else dataset_card
additions.append(CommitOperationAdd(path_in_repo="README.md", path_or_fileobj=str(dataset_card).encode()))
commit_message = commit_message if commit_message is not None else "Upload dataset"
if len(additions) <= config.UPLOADS_MAX_NUMBER_PER_COMMIT:
api.create_commit(
repo_id,
operations=additions + deletions,
commit_message=commit_message,
token=token,
repo_type="dataset",
revision=revision,
create_pr=create_pr,
)
else:
logger.info(
f"Number of files to upload is larger than {config.UPLOADS_MAX_NUMBER_PER_COMMIT}. Splitting the push into multiple commits."
)
num_commits = math.ceil(len(additions) / config.UPLOADS_MAX_NUMBER_PER_COMMIT)
for i in range(0, num_commits):
operations = additions[
i * config.UPLOADS_MAX_NUMBER_PER_COMMIT : (i + 1) * config.UPLOADS_MAX_NUMBER_PER_COMMIT
] + (deletions if i == 0 else [])
api.create_commit(
repo_id,
operations=operations,
commit_message=commit_message + f" (part {i:05d}-of-{num_commits:05d})",
token=token,
repo_type="dataset",
revision=revision,
create_pr=create_pr,
)
logger.info(
f"Commit #{i+1} completed"
+ (f" (still {num_commits - i - 1} to go)" if num_commits - i - 1 else "")
+ "."
)
@transmit_format
@fingerprint_transform(inplace=False)
def add_column(self, name: str, column: Union[list, np.array], new_fingerprint: str):
"""Add column to Dataset.
<Added version="1.7"/>
Args:
name (`str`):
Column name.
column (`list` or `np.array`):
Column data to be added.
Returns:
[`Dataset`]
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", split="validation")
>>> more_text = ds["text"]
>>> ds.add_column(name="text_2", column=more_text)
Dataset({
features: ['text', 'label', 'text_2'],
num_rows: 1066
})
```
"""
column_table = InMemoryTable.from_pydict({name: column})
_check_column_names(self._data.column_names + column_table.column_names)
dataset = self.flatten_indices() if self._indices is not None else self
# Concatenate tables horizontally
table = concat_tables([dataset._data, column_table], axis=1)
# Update features
info = dataset.info.copy()
info.features.update(Features.from_arrow_schema(column_table.schema))
table = update_metadata_with_features(table, info.features)
return Dataset(table, info=info, split=self.split, indices_table=None, fingerprint=new_fingerprint)
def add_faiss_index(
self,
column: str,
index_name: Optional[str] = None,
device: Optional[int] = None,
string_factory: Optional[str] = None,
metric_type: Optional[int] = None,
custom_index: Optional["faiss.Index"] = None, # noqa: F821
batch_size: int = 1000,
train_size: Optional[int] = None,
faiss_verbose: bool = False,
dtype=np.float32,
):
"""Add a dense index using Faiss for fast retrieval.
By default the index is done over the vectors of the specified column.
You can specify `device` if you want to run it on GPU (`device` must be the GPU index).
You can find more information about Faiss here:
- For [string factory](https://github.com/facebookresearch/faiss/wiki/The-index-factory)
Args:
column (`str`):
The column of the vectors to add to the index.
index_name (`str`, *optional*):
The `index_name`/identifier of the index.
This is the `index_name` that is used to call [`~datasets.Dataset.get_nearest_examples`] or [`~datasets.Dataset.search`].
By default it corresponds to `column`.
device (`Union[int, List[int]]`, *optional*):
If positive integer, this is the index of the GPU to use. If negative integer, use all GPUs.
If a list of positive integers is passed in, run only on those GPUs. By default it uses the CPU.
string_factory (`str`, *optional*):
This is passed to the index factory of Faiss to create the index.
Default index class is `IndexFlat`.
metric_type (`int`, *optional*):
Type of metric. Ex: `faiss.METRIC_INNER_PRODUCT` or `faiss.METRIC_L2`.
custom_index (`faiss.Index`, *optional*):
Custom Faiss index that you already have instantiated and configured for your needs.
batch_size (`int`):
Size of the batch to use while adding vectors to the `FaissIndex`. Default value is `1000`.
<Added version="2.4.0"/>
train_size (`int`, *optional*):
If the index needs a training step, specifies how many vectors will be used to train the index.
faiss_verbose (`bool`, defaults to `False`):
Enable the verbosity of the Faiss index.
dtype (`data-type`):
The dtype of the numpy arrays that are indexed.
Default is `np.float32`.
Example:
```python
>>> ds = datasets.load_dataset('crime_and_punish', split='train')
>>> ds_with_embeddings = ds.map(lambda example: {'embeddings': embed(example['line']}))
>>> ds_with_embeddings.add_faiss_index(column='embeddings')
>>> # query
>>> scores, retrieved_examples = ds_with_embeddings.get_nearest_examples('embeddings', embed('my new query'), k=10)
>>> # save index
>>> ds_with_embeddings.save_faiss_index('embeddings', 'my_index.faiss')
>>> ds = datasets.load_dataset('crime_and_punish', split='train')
>>> # load index
>>> ds.load_faiss_index('embeddings', 'my_index.faiss')
>>> # query
>>> scores, retrieved_examples = ds.get_nearest_examples('embeddings', embed('my new query'), k=10)
```
"""
with self.formatted_as(type="numpy", columns=[column], dtype=dtype):
super().add_faiss_index(
column=column,
index_name=index_name,
device=device,
string_factory=string_factory,
metric_type=metric_type,
custom_index=custom_index,
batch_size=batch_size,
train_size=train_size,
faiss_verbose=faiss_verbose,
)
return self
def add_faiss_index_from_external_arrays(
self,
external_arrays: np.array,
index_name: str,
device: Optional[int] = None,
string_factory: Optional[str] = None,
metric_type: Optional[int] = None,
custom_index: Optional["faiss.Index"] = None, # noqa: F821
batch_size: int = 1000,
train_size: Optional[int] = None,
faiss_verbose: bool = False,
dtype=np.float32,
):
"""Add a dense index using Faiss for fast retrieval.
The index is created using the vectors of `external_arrays`.
You can specify `device` if you want to run it on GPU (`device` must be the GPU index).
You can find more information about Faiss here:
- For [string factory](https://github.com/facebookresearch/faiss/wiki/The-index-factory)
Args:
external_arrays (`np.array`):
If you want to use arrays from outside the lib for the index, you can set `external_arrays`.
It will use `external_arrays` to create the Faiss index instead of the arrays in the given `column`.
index_name (`str`):
The `index_name`/identifier of the index.
This is the `index_name` that is used to call [`~datasets.Dataset.get_nearest_examples`] or [`~datasets.Dataset.search`].
device (Optional `Union[int, List[int]]`, *optional*):
If positive integer, this is the index of the GPU to use. If negative integer, use all GPUs.
If a list of positive integers is passed in, run only on those GPUs. By default it uses the CPU.
string_factory (`str`, *optional*):
This is passed to the index factory of Faiss to create the index.
Default index class is `IndexFlat`.
metric_type (`int`, *optional*):
Type of metric. Ex: `faiss.faiss.METRIC_INNER_PRODUCT` or `faiss.METRIC_L2`.
custom_index (`faiss.Index`, *optional*):
Custom Faiss index that you already have instantiated and configured for your needs.
batch_size (`int`, *optional*):
Size of the batch to use while adding vectors to the FaissIndex. Default value is 1000.
<Added version="2.4.0"/>
train_size (`int`, *optional*):
If the index needs a training step, specifies how many vectors will be used to train the index.
faiss_verbose (`bool`, defaults to False):
Enable the verbosity of the Faiss index.
dtype (`numpy.dtype`):
The dtype of the numpy arrays that are indexed. Default is np.float32.
"""
super().add_faiss_index_from_external_arrays(
external_arrays=external_arrays.astype(dtype),
index_name=index_name,
device=device,
string_factory=string_factory,
metric_type=metric_type,
custom_index=custom_index,
batch_size=batch_size,
train_size=train_size,
faiss_verbose=faiss_verbose,
)
def add_elasticsearch_index(
self,
column: str,
index_name: Optional[str] = None,
host: Optional[str] = None,
port: Optional[int] = None,
es_client: Optional["elasticsearch.Elasticsearch"] = None, # noqa: F821
es_index_name: Optional[str] = None,
es_index_config: Optional[dict] = None,
):
"""Add a text index using ElasticSearch for fast retrieval. This is done in-place.
Args:
column (`str`):
The column of the documents to add to the index.
index_name (`str`, *optional*):
The `index_name`/identifier of the index.
This is the index name that is used to call [`~Dataset.get_nearest_examples`] or [`Dataset.search`].
By default it corresponds to `column`.
host (`str`, *optional*, defaults to `localhost`):
Host of where ElasticSearch is running.
port (`str`, *optional*, defaults to `9200`):
Port of where ElasticSearch is running.
es_client (`elasticsearch.Elasticsearch`, *optional*):
The elasticsearch client used to create the index if host and port are `None`.
es_index_name (`str`, *optional*):
The elasticsearch index name used to create the index.
es_index_config (`dict`, *optional*):
The configuration of the elasticsearch index.
Default config is:
```
{
"settings": {
"number_of_shards": 1,
"analysis": {"analyzer": {"stop_standard": {"type": "standard", " stopwords": "_english_"}}},
},
"mappings": {
"properties": {
"text": {
"type": "text",
"analyzer": "standard",
"similarity": "BM25"
},
}
},
}
```
Example:
```python
>>> es_client = elasticsearch.Elasticsearch()
>>> ds = datasets.load_dataset('crime_and_punish', split='train')
>>> ds.add_elasticsearch_index(column='line', es_client=es_client, es_index_name="my_es_index")
>>> scores, retrieved_examples = ds.get_nearest_examples('line', 'my new query', k=10)
```
"""
with self.formatted_as(type=None, columns=[column]):
super().add_elasticsearch_index(
column=column,
index_name=index_name,
host=host,
port=port,
es_client=es_client,
es_index_name=es_index_name,
es_index_config=es_index_config,
)
return self
@transmit_format
@fingerprint_transform(inplace=False)
def add_item(self, item: dict, new_fingerprint: str):
"""Add item to Dataset.
<Added version="1.7"/>
Args:
item (`dict`):
Item data to be added.
Returns:
[`Dataset`]
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", split="validation")
>>> new_review = {'label': 0, 'text': 'this movie is the absolute worst thing I have ever seen'}
>>> ds = ds.add_item(new_review)
>>> ds[-1]
{'label': 0, 'text': 'this movie is the absolute worst thing I have ever seen'}
```
"""
item_table = InMemoryTable.from_pydict({k: [v] for k, v in item.items()})
# We don't call _check_if_features_can_be_aligned here so this cast is "unsafe"
dset_features, item_features = _align_features(
[self._info.features, Features.from_arrow_schema(item_table.schema)]
)
# Cast to align the schemas of the tables and concatenate the tables
table = concat_tables(
[
self._data.cast(dset_features.arrow_schema) if self._info.features != dset_features else self._data,
item_table.cast(item_features.arrow_schema),
]
)
if self._indices is None:
indices_table = None
else:
item_indices_array = pa.array([len(self._data)], type=pa.uint64())
item_indices_table = InMemoryTable.from_arrays([item_indices_array], names=["indices"])
indices_table = concat_tables([self._indices, item_indices_table])
info = self.info.copy()
info.features.update(item_features)
table = update_metadata_with_features(table, info.features)
return Dataset(
table,
info=info,
split=self.split,
indices_table=indices_table,
fingerprint=new_fingerprint,
)
def align_labels_with_mapping(self, label2id: Dict, label_column: str) -> "Dataset":
"""Align the dataset's label ID and label name mapping to match an input `label2id` mapping.
This is useful when you want to ensure that a model's predicted labels are aligned with the dataset.
The alignment in done using the lowercase label names.
Args:
label2id (`dict`):
The label name to ID mapping to align the dataset with.
label_column (`str`):
The column name of labels to align on.
Example:
```python
>>> # dataset with mapping {'entailment': 0, 'neutral': 1, 'contradiction': 2}
>>> ds = load_dataset("glue", "mnli", split="train")
>>> # mapping to align with
>>> label2id = {'CONTRADICTION': 0, 'NEUTRAL': 1, 'ENTAILMENT': 2}
>>> ds_aligned = ds.align_labels_with_mapping(label2id, "label")
```
"""
# Sanity checks
if label_column not in self._data.column_names:
raise ValueError(f"Column ({label_column}) not in table columns ({self._data.column_names}).")
label_feature = self._info.features[label_column]
if not (
isinstance(label_feature, ClassLabel)
or (isinstance(label_feature, Sequence) and isinstance(label_feature.feature, ClassLabel))
):
raise ValueError(
f"Aligning labels with a mapping is only supported for {ClassLabel.__name__} column or {Sequence.__name__} column with the inner type {ClassLabel.__name__}, and column {label_feature} is of type {type(label_feature).__name__}."
)
# Sort input mapping by ID value to ensure the label names are aligned
label2id = dict(sorted(label2id.items(), key=lambda item: item[1]))
label_names = list(label2id.keys())
# Some label mappings use uppercase label names so we lowercase them during alignment
label2id = {k.lower(): v for k, v in label2id.items()}
int2str_function = (
label_feature.int2str if isinstance(label_feature, ClassLabel) else label_feature.feature.int2str
)
if isinstance(label_feature, ClassLabel):
def process_label_ids(batch):
dset_label_names = [
int2str_function(label_id).lower() if label_id is not None else None
for label_id in batch[label_column]
]
batch[label_column] = [
label2id[label_name] if label_name is not None else None for label_name in dset_label_names
]
return batch
else:
def process_label_ids(batch):
dset_label_names = [
[int2str_function(label_id).lower() if label_id is not None else None for label_id in seq]
for seq in batch[label_column]
]
batch[label_column] = [
[label2id[label_name] if label_name is not None else None for label_name in seq]
for seq in dset_label_names
]
return batch
features = self.features
features[label_column] = (
ClassLabel(num_classes=len(label_names), names=label_names)
if isinstance(label_feature, ClassLabel)
else Sequence(ClassLabel(num_classes=len(label_names), names=label_names))
)
return self.map(process_label_ids, features=features, batched=True, desc="Aligning the labels")
def _concatenate_map_style_datasets(
dsets: List[Dataset],
info: Optional[DatasetInfo] = None,
split: Optional[NamedSplit] = None,
axis: int = 0,
):
"""
Converts a list of :class:`Dataset` with the same schema into a single :class:`Dataset`.
When you concatenate on axis 0, missing data are filled with None values.
Args:
dsets (`List[datasets.Dataset]`): List of Datasets to concatenate.
info (:class:`DatasetInfo`, optional): Dataset information, like description, citation, etc.
split (:class:`NamedSplit`, optional): Name of the dataset split.
axis (``{0, 1}``, default ``0``, meaning over rows):
Axis to concatenate over, where ``0`` means over rows (vertically) and ``1`` means over columns
(horizontally).
*New in version 1.6.0*
Example:
```py
>>> ds3 = _concatenate_map_style_datasets([ds1, ds2])
```
"""
# Ignore datasets with no rows
if any(dset.num_rows > 0 for dset in dsets):
dsets = [dset for dset in dsets if dset.num_rows > 0]
else:
# Return first dataset if all datasets are empty
return dsets[0]
# Perform checks (and a potentional cast if axis=0)
if axis == 0:
_check_if_features_can_be_aligned([dset.features for dset in dsets])
else:
if not all(dset.num_rows == dsets[0].num_rows for dset in dsets):
raise ValueError("Number of rows must match for all datasets")
_check_column_names([col_name for dset in dsets for col_name in dset._data.column_names])
# Find common format or reset format
format = dsets[0].format
if any(dset.format != format for dset in dsets):
format = {}
logger.info("Some of the datasets have disparate format. Resetting the format of the concatenated dataset.")
def apply_offset_to_indices_table(table, offset):
if offset == 0:
return table
else:
array = table["indices"]
new_array = pc.add(array, pa.scalar(offset, type=pa.uint64()))
return InMemoryTable.from_arrays([new_array], names=["indices"])
# Concatenate indices if they exist
if any(dset._indices is not None for dset in dsets):
if axis == 0:
# Datasets with no indices tables are replaced with a dataset with an indices table in memory.
# Applying an offset to an indices table also brings the table in memory.
indices_tables = []
for i in range(len(dsets)):
if dsets[i]._indices is None:
dsets[i] = dsets[i]._select_with_indices_mapping(range(len(dsets[i])))
indices_tables.append(dsets[i]._indices)
# An offset needs to be applied to the indices before concatenating
offset = 0
for i in range(len(dsets)):
indices_tables[i] = apply_offset_to_indices_table(indices_tables[i], offset)
offset += len(dsets[i]._data)
# Concatenate indices
indices_tables = [t for t in indices_tables if len(t) > 0]
if indices_tables:
indices_table = concat_tables(indices_tables)
else:
indices_table = InMemoryTable.from_batches([], schema=pa.schema({"indices": pa.int64()}))
else:
if len(dsets) == 1:
indices_table = dsets[0]._indices
else:
for i in range(len(dsets)):
dsets[i] = dsets[i].flatten_indices()
indices_table = None
else:
indices_table = None
table = concat_tables([dset._data for dset in dsets], axis=axis)
if axis == 0:
features_list = _align_features([dset.features for dset in dsets])
else:
features_list = [dset.features for dset in dsets]
table = update_metadata_with_features(table, {k: v for features in features_list for k, v in features.items()})
# Concatenate infos
if info is None:
info = DatasetInfo.from_merge([dset.info for dset in dsets])
fingerprint = update_fingerprint(
"".join(dset._fingerprint for dset in dsets), _concatenate_map_style_datasets, {"info": info, "split": split}
)
# Make final concatenated dataset
concatenated_dataset = Dataset(
table,
info=info,
split=split,
indices_table=indices_table,
fingerprint=fingerprint,
)
concatenated_dataset.set_format(**format)
return concatenated_dataset
def _interleave_map_style_datasets(
datasets: List["Dataset"],
probabilities: Optional[List[float]] = None,
seed: Optional[int] = None,
info: Optional[DatasetInfo] = None,
split: Optional[NamedSplit] = None,
stopping_strategy: Literal["first_exhausted", "all_exhausted"] = "first_exhausted",
**kwargs,
) -> "Dataset":
"""
Interleave several map-style datasets (sources) into a single map-style dataset.
The new dataset is constructed by alternating between the sources to get the examples.
If `probabilities = None` (default) the new dataset is constructed by cycling between each source to get the examples.
If `probabilities` is not `None, the new dataset is constructed by getting examples from a random source at a time according to the provided probabilities.
Args:
datasets (`List[Dataset]`): list of datasets to interleave
probabilities (`List[float]`, optional, default None): If specified, the new dataset is constructed by sampling
examples from one source at a time according to these probabilities.
seed (`int`, optional, default None): The random seed used to choose a source for each example.
info (:class:`DatasetInfo`, optional): Dataset information, like description, citation, etc.
split (:class:`NamedSplit`, optional): Name of the dataset split.
stopping_strategy (`str`, defaults to `first_exhausted`):
Two strategies are proposed right now.
By default, `first_exhausted` is an undersampling strategy, i.e the dataset construction is stopped as soon as one dataset has ran out of samples.
If the strategy is `all_exhausted`, we use an oversampling strategy, i.e the dataset construction is stopped as soon as every samples of every dataset has been added at least once.
Note that if the strategy is `all_exhausted`, the interleaved dataset size can get enormous:
- with no probabilities, the resulting dataset will have max_length_datasets*nb_dataset samples.
- with given probabilities, the resulting dataset will have more samples if some datasets have really low probability of visiting.
**kwargs (additional keyword arguments): Keyword arguments to be passed to :meth:`datasets.Datasets.select` when selecting the indices used to interleave the datasets.
Output:
:class:`datasets.Dataset`
"""
if stopping_strategy not in ["first_exhausted", "all_exhausted"]:
raise ValueError(
f"{stopping_strategy} stopping strategy in `interleave_datasets` is not implemented yet with a list of {type(datasets[0])}"
)
# To interleave the datasets, we concatenate them and then we re-order the indices
concatenated_datasets = _concatenate_map_style_datasets(datasets, info=info, split=split)
# Let's now build the indices to pass to .select()
lengths = [len(dset) for dset in datasets]
offsets = np.cumsum([0] + lengths[:-1])
# if stopping_strategy is "first_exhausted", it is an undersampling situation whereas it is an oversampling situation if it is "all_exhausted"
oversampling = stopping_strategy == "all_exhausted"
if probabilities is None and not oversampling:
# Undersampling situation with cycling between each sources
# Example:: If lengths of the datasets are [3, 4, 5]
# Then the resulting indices should be [0, 3, 7, 1, 4, 8, 2, 6, 9]
# Note that we only have 3 examples per dataset since the first dataset ran out of examples
# Reasoning behind the following operation: keeping the min_length first indices of each dataset
# while offsetting in order to correspond to the right indices of the concatenated dataset
# and flattening to effectively interleave the datasets
indices = (offsets.reshape(1, -1) + np.arange(min(lengths)).reshape(-1, 1)).flatten().tolist()
elif probabilities is None:
# Oversampling situation with cycling between each sources
# Then the resulting indices should be [0, 3, 7, 1, 4, 8, 2, 5, 9, 0, 6, 10, 1, 3, 11]
# Note that we have 5 examples per dataset with a rolling window since the longest dataset has 5 samples
# Reasoning behind the following operation: for each dataset indices (i.e column) repeat the indices to have max_length indices per dataset
# For example, if the max_length is 5 and the i-th dataset has 3 samples, the i-th column will be [0,1,2,0,1]
indices = np.mod(np.arange(max(lengths)).reshape(-1, 1), np.array(lengths).reshape(1, -1))
# We have to keep the indices to their respective dataset offsets and to flatten to effectively interleave the datasets
indices = (indices + offsets).flatten().tolist()
else:
# boolean array indicating if at index i if the dataset_i has been fully exhausted
is_exhausted = np.full(len(lengths), False)
# if undersampling ("first_exhausted"), we stop as soon as one dataset is exhausted
# if oversampling ("all_exhausted"), we stop as soons as every dataset is exhausted, i.e as soon as every samples of every dataset has been visited at least once
bool_strategy_func = np.all if oversampling else np.any
def iter_random_indices():
"""Get an infinite iterator that randomly samples the index of the source to pick examples from."""
rng = np.random.default_rng(seed)
while True:
yield from (int(i) for i in rng.choice(len(datasets), size=1000, p=probabilities))
current_index = [0] * len(datasets)
indices = []
for source_idx in iter_random_indices():
# If no oversampling, we stop as soon as a dataset has ran out of examples (np.any)
# Otherwise, we stop as soon as every dataset has ran out of examples (np.all)
if bool_strategy_func(is_exhausted):
# the stopping condition was reached, let's stop
break
# let's add the example at the current index of the `source_idx`-th dataset
indices.append(current_index[source_idx] + offsets[source_idx])
current_index[source_idx] += 1
# we've ran out of examples for the current dataset, let's update our boolean array and bring the current_index back to 0
if current_index[source_idx] >= lengths[source_idx]:
is_exhausted[source_idx] = True
current_index[source_idx] = 0
return concatenated_datasets.select(indices, **kwargs)
def _split_by_node_map_style_dataset(dataset: Dataset, rank: int, world_size: int) -> Dataset:
"""
Split a dataset for the node at rank `rank` in a pool of nodes of size `world_size`.
Each node is assigned a chunk of data, e.g. rank 0 is given the first chunk of the dataset.
To maximize data loading throughput, chunks are made of contiguous data on disk if possible.
Args:
dataset ([`Dataset`]):
The dataset to split by node.
rank (`int`):
Rank of the current node.
world_size (`int`):
Total number of nodes.
Returns:
[`Dataset`]: The dataset to be used on the node at rank `rank`.
"""
return dataset.shard(num_shards=world_size, index=rank, contiguous=True)
# This is outside Dataset.filter as it needs to be picklable for multiprocessing
def get_indices_from_mask_function(
function: Callable,
batched: bool,
with_indices: bool,
input_columns: Optional[Union[str, List[str]]],
indices_mapping: Optional[Table] = None,
*args,
**fn_kwargs,
):
if batched:
# we extract indices from args
*inputs, indices = args
if with_indices:
mask = function(*inputs, indices, **fn_kwargs)
else:
mask = function(*inputs, **fn_kwargs)
else:
# we get batched data (to do less look-ups) but `function` only accepts one example
# therefore we need to call `function` on each example of the batch to get the mask
*inputs, indices = args
mask = []
if input_columns is None:
# inputs only contains a batch of examples
batch: dict = inputs[0]
num_examples = len(batch[next(iter(batch.keys()))])
for i in range(num_examples):
example = {key: batch[key][i] for key in batch}
mask.append(
function(example, indices[i], **fn_kwargs) if with_indices else function(example, **fn_kwargs)
)
else:
# inputs is a list of columns
columns: List[List] = inputs
num_examples = len(columns[0])
for i in range(num_examples):
input = [column[i] for column in columns]
mask.append(
function(*input, indices[i], **fn_kwargs) if with_indices else function(*input, **fn_kwargs)
)
indices_array = [i for i, to_keep in zip(indices, mask) if to_keep]
if indices_mapping is not None:
indices_array = pa.array(indices_array, type=pa.uint64())
indices_array = indices_mapping.column(0).take(indices_array)
indices_array = indices_array.to_pylist()
return {"indices": indices_array}
| 0
|
hf_public_repos/datasets/src
|
hf_public_repos/datasets/src/datasets/splits.py
|
# Copyright 2020 The HuggingFace Datasets Authors and the TensorFlow Datasets Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# Lint as: python3
"""Splits related API."""
import abc
import collections
import copy
import dataclasses
import re
from dataclasses import dataclass
from typing import Dict, List, Optional, Union
from .arrow_reader import FileInstructions, make_file_instructions
from .naming import _split_re
from .utils.py_utils import NonMutableDict, asdict
@dataclass
class SplitInfo:
name: str = dataclasses.field(default="", metadata={"include_in_asdict_even_if_is_default": True})
num_bytes: int = dataclasses.field(default=0, metadata={"include_in_asdict_even_if_is_default": True})
num_examples: int = dataclasses.field(default=0, metadata={"include_in_asdict_even_if_is_default": True})
shard_lengths: Optional[List[int]] = None
# Deprecated
# For backward compatibility, this field needs to always be included in files like
# dataset_infos.json and dataset_info.json files
# To do so, we always include it in the output of datasets.utils.py_utils.asdict(split_info)
dataset_name: Optional[str] = dataclasses.field(
default=None, metadata={"include_in_asdict_even_if_is_default": True}
)
@property
def file_instructions(self):
"""Returns the list of dict(filename, take, skip)."""
# `self.dataset_name` is assigned in `SplitDict.add()`.
instructions = make_file_instructions(
name=self.dataset_name,
split_infos=[self],
instruction=str(self.name),
)
return instructions.file_instructions
@dataclass
class SubSplitInfo:
"""Wrapper around a sub split info.
This class expose info on the subsplit:
```
ds, info = datasets.load_dataset(..., split='train[75%:]', with_info=True)
info.splits['train[75%:]'].num_examples
```
"""
instructions: FileInstructions
@property
def num_examples(self):
"""Returns the number of example in the subsplit."""
return self.instructions.num_examples
@property
def file_instructions(self):
"""Returns the list of dict(filename, take, skip)."""
return self.instructions.file_instructions
class SplitBase(metaclass=abc.ABCMeta):
# pylint: disable=line-too-long
"""Abstract base class for Split compositionality.
See the
[guide on splits](../loading#slice-splits)
for more information.
There are three parts to the composition:
1) The splits are composed (defined, merged, split,...) together before
calling the `.as_dataset()` function. This is done with the `__add__`,
`__getitem__`, which return a tree of `SplitBase` (whose leaf
are the `NamedSplit` objects)
```
split = datasets.Split.TRAIN + datasets.Split.TEST.subsplit(datasets.percent[:50])
```
2) The `SplitBase` is forwarded to the `.as_dataset()` function
to be resolved into actual read instruction. This is done by the
`.get_read_instruction()` method which takes the real dataset splits
(name, number of shards,...) and parse the tree to return a
`SplitReadInstruction()` object
```
read_instruction = split.get_read_instruction(self.info.splits)
```
3) The `SplitReadInstruction` is then used in the `tf.data.Dataset` pipeline
to define which files to read and how to skip examples within file.
"""
# pylint: enable=line-too-long
@abc.abstractmethod
def get_read_instruction(self, split_dict):
"""Parse the descriptor tree and compile all read instructions together.
Args:
split_dict: `dict`, The `dict[split_name, SplitInfo]` of the dataset
Returns:
split_read_instruction: `SplitReadInstruction`
"""
raise NotImplementedError("Abstract method")
def __eq__(self, other):
"""Equality: datasets.Split.TRAIN == 'train'."""
if isinstance(other, (NamedSplit, str)):
return False
raise NotImplementedError("Equality is not implemented between merged/sub splits.")
def __ne__(self, other):
"""InEquality: datasets.Split.TRAIN != 'test'."""
return not self.__eq__(other)
def __add__(self, other):
"""Merging: datasets.Split.TRAIN + datasets.Split.TEST."""
return _SplitMerged(self, other)
def subsplit(self, arg=None, k=None, percent=None, weighted=None): # pylint: disable=redefined-outer-name
"""Divides this split into subsplits.
There are 3 ways to define subsplits, which correspond to the 3
arguments `k` (get `k` even subsplits), `percent` (get a slice of the
dataset with `datasets.percent`), and `weighted` (get subsplits with proportions
specified by `weighted`).
Example::
```
# 50% train, 50% test
train, test = split.subsplit(k=2)
# 50% train, 25% test, 25% validation
train, test, validation = split.subsplit(weighted=[2, 1, 1])
# Extract last 20%
subsplit = split.subsplit(datasets.percent[-20:])
```
Warning: k and weighted will be converted into percent which mean that
values below the percent will be rounded up or down. The final split may be
bigger to deal with remainders. For instance:
```
train, test, valid = split.subsplit(k=3) # 33%, 33%, 34%
s1, s2, s3, s4 = split.subsplit(weighted=[2, 2, 1, 1]) # 33%, 33%, 16%, 18%
```
Args:
arg: If no kwargs are given, `arg` will be interpreted as one of
`k`, `percent`, or `weighted` depending on the type.
For example:
```
split.subsplit(10) # Equivalent to split.subsplit(k=10)
split.subsplit(datasets.percent[:-20]) # percent=datasets.percent[:-20]
split.subsplit([1, 1, 2]) # weighted=[1, 1, 2]
```
k: `int` If set, subdivide the split into `k` equal parts.
percent: `datasets.percent slice`, return a single subsplit corresponding to
a slice of the original split. For example:
`split.subsplit(datasets.percent[-20:]) # Last 20% of the dataset`.
weighted: `list[int]`, return a list of subsplits whose proportions match
the normalized sum of the list. For example:
`split.subsplit(weighted=[1, 1, 2]) # 25%, 25%, 50%`.
Returns:
A subsplit or list of subsplits extracted from this split object.
"""
# Note that the percent kwargs redefine the outer name datasets.percent. This
# is done for consistency (.subsplit(percent=datasets.percent[:40]))
if sum(bool(x) for x in (arg, k, percent, weighted)) != 1:
raise ValueError("Only one argument of subsplit should be set.")
# Auto deduce k
if isinstance(arg, int):
k = arg
elif isinstance(arg, slice):
percent = arg
elif isinstance(arg, list):
weighted = arg
if not (k or percent or weighted):
raise ValueError(
f"Invalid split argument {arg}. Only list, slice and int supported. "
"One of k, weighted or percent should be set to a non empty value."
)
def assert_slices_coverage(slices):
# Ensure that the expended slices cover all percents.
assert sum((list(range(*s.indices(100))) for s in slices), []) == list(range(100))
if k:
if not 0 < k <= 100:
raise ValueError(f"Subsplit k should be between 0 and 100, got {k}")
shift = 100 // k
slices = [slice(i * shift, (i + 1) * shift) for i in range(k)]
# Round up last element to ensure all elements are taken
slices[-1] = slice(slices[-1].start, 100)
# Internal check to ensure full coverage
assert_slices_coverage(slices)
return tuple(_SubSplit(self, s) for s in slices)
elif percent:
return _SubSplit(self, percent)
elif weighted:
# Normalize the weighted sum
total = sum(weighted)
weighted = [100 * x // total for x in weighted]
# Create the slice for each of the elements
start = 0
stop = 0
slices = []
for v in weighted:
stop += v
slices.append(slice(start, stop))
start = stop
# Round up last element to ensure all elements are taken
slices[-1] = slice(slices[-1].start, 100)
# Internal check to ensure full coverage
assert_slices_coverage(slices)
return tuple(_SubSplit(self, s) for s in slices)
else:
# Should not be possible
raise ValueError("Could not determine the split")
# 2 requirements:
# 1. datasets.percent be sliceable
# 2. datasets.percent be documented
#
# Instances are not documented, so we want datasets.percent to be a class, but to
# have it be sliceable, we need this metaclass.
class PercentSliceMeta(type):
def __getitem__(cls, slice_value):
if not isinstance(slice_value, slice):
raise ValueError(f"datasets.percent should only be called with slice, not {slice_value}")
return slice_value
class PercentSlice(metaclass=PercentSliceMeta):
# pylint: disable=line-too-long
"""Syntactic sugar for defining slice subsplits: `datasets.percent[75:-5]`.
See the
[guide on splits](../loading#slice-splits)
for more information.
"""
# pylint: enable=line-too-long
pass
percent = PercentSlice # pylint: disable=invalid-name
class _SplitMerged(SplitBase):
"""Represent two split descriptors merged together."""
def __init__(self, split1, split2):
self._split1 = split1
self._split2 = split2
def get_read_instruction(self, split_dict):
read_instruction1 = self._split1.get_read_instruction(split_dict)
read_instruction2 = self._split2.get_read_instruction(split_dict)
return read_instruction1 + read_instruction2
def __repr__(self):
return f"({repr(self._split1)} + {repr(self._split2)})"
class _SubSplit(SplitBase):
"""Represent a sub split of a split descriptor."""
def __init__(self, split, slice_value):
self._split = split
self._slice_value = slice_value
def get_read_instruction(self, split_dict):
return self._split.get_read_instruction(split_dict)[self._slice_value]
def __repr__(self):
slice_str = "{start}:{stop}"
if self._slice_value.step is not None:
slice_str += ":{step}"
slice_str = slice_str.format(
start="" if self._slice_value.start is None else self._slice_value.start,
stop="" if self._slice_value.stop is None else self._slice_value.stop,
step=self._slice_value.step,
)
return f"{repr(self._split)}(datasets.percent[{slice_str}])"
class NamedSplit(SplitBase):
"""Descriptor corresponding to a named split (train, test, ...).
Example:
Each descriptor can be composed with other using addition or slice:
```py
split = datasets.Split.TRAIN.subsplit(datasets.percent[0:25]) + datasets.Split.TEST
```
The resulting split will correspond to 25% of the train split merged with
100% of the test split.
A split cannot be added twice, so the following will fail:
```py
split = (
datasets.Split.TRAIN.subsplit(datasets.percent[:25]) +
datasets.Split.TRAIN.subsplit(datasets.percent[75:])
) # Error
split = datasets.Split.TEST + datasets.Split.ALL # Error
```
The slices can be applied only one time. So the following are valid:
```py
split = (
datasets.Split.TRAIN.subsplit(datasets.percent[:25]) +
datasets.Split.TEST.subsplit(datasets.percent[:50])
)
split = (datasets.Split.TRAIN + datasets.Split.TEST).subsplit(datasets.percent[:50])
```
But this is not valid:
```py
train = datasets.Split.TRAIN
test = datasets.Split.TEST
split = train.subsplit(datasets.percent[:25]).subsplit(datasets.percent[:25])
split = (train.subsplit(datasets.percent[:25]) + test).subsplit(datasets.percent[:50])
```
"""
def __init__(self, name):
self._name = name
split_names_from_instruction = [split_instruction.split("[")[0] for split_instruction in name.split("+")]
for split_name in split_names_from_instruction:
if not re.match(_split_re, split_name):
raise ValueError(f"Split name should match '{_split_re}' but got '{split_name}'.")
def __str__(self):
return self._name
def __repr__(self):
return f"NamedSplit({self._name!r})"
def __eq__(self, other):
"""Equality: datasets.Split.TRAIN == 'train'."""
if isinstance(other, NamedSplit):
return self._name == other._name # pylint: disable=protected-access
elif isinstance(other, SplitBase):
return False
elif isinstance(other, str): # Other should be string
return self._name == other
else:
raise ValueError(f"Equality not supported between split {self} and {other}")
def __lt__(self, other):
return self._name < other._name # pylint: disable=protected-access
def __hash__(self):
return hash(self._name)
def get_read_instruction(self, split_dict):
return SplitReadInstruction(split_dict[self._name])
class NamedSplitAll(NamedSplit):
"""Split corresponding to the union of all defined dataset splits."""
def __init__(self):
super().__init__("all")
def __repr__(self):
return "NamedSplitAll()"
def get_read_instruction(self, split_dict):
# Merge all dataset split together
read_instructions = [SplitReadInstruction(s) for s in split_dict.values()]
return sum(read_instructions, SplitReadInstruction())
class Split:
# pylint: disable=line-too-long
"""`Enum` for dataset splits.
Datasets are typically split into different subsets to be used at various
stages of training and evaluation.
- `TRAIN`: the training data.
- `VALIDATION`: the validation data. If present, this is typically used as
evaluation data while iterating on a model (e.g. changing hyperparameters,
model architecture, etc.).
- `TEST`: the testing data. This is the data to report metrics on. Typically
you do not want to use this during model iteration as you may overfit to it.
- `ALL`: the union of all defined dataset splits.
All splits, including compositions inherit from `datasets.SplitBase`.
See the [guide](../load_hub#splits) on splits for more information.
Example:
```py
>>> datasets.SplitGenerator(
... name=datasets.Split.TRAIN,
... gen_kwargs={"split_key": "train", "files": dl_manager.download_and extract(url)},
... ),
... datasets.SplitGenerator(
... name=datasets.Split.VALIDATION,
... gen_kwargs={"split_key": "validation", "files": dl_manager.download_and extract(url)},
... ),
... datasets.SplitGenerator(
... name=datasets.Split.TEST,
... gen_kwargs={"split_key": "test", "files": dl_manager.download_and extract(url)},
... )
```
"""
# pylint: enable=line-too-long
TRAIN = NamedSplit("train")
TEST = NamedSplit("test")
VALIDATION = NamedSplit("validation")
ALL = NamedSplitAll()
def __new__(cls, name):
"""Create a custom split with datasets.Split('custom_name')."""
return NamedSplitAll() if name == "all" else NamedSplit(name)
# Similar to SplitInfo, but contain an additional slice info
SlicedSplitInfo = collections.namedtuple(
"SlicedSplitInfo",
[
"split_info",
"slice_value",
],
) # noqa: E231
class SplitReadInstruction:
"""Object containing the reading instruction for the dataset.
Similarly to `SplitDescriptor` nodes, this object can be composed with itself,
but the resolution happens instantaneously, instead of keeping track of the
tree, such as all instructions are compiled and flattened in a single
SplitReadInstruction object containing the list of files and slice to use.
Once resolved, the instructions can be accessed with:
```
read_instructions.get_list_sliced_split_info() # List of splits to use
```
"""
def __init__(self, split_info=None):
self._splits = NonMutableDict(error_msg="Overlap between splits. Split {key} has been added with " "itself.")
if split_info:
self.add(SlicedSplitInfo(split_info=split_info, slice_value=None))
def add(self, sliced_split):
"""Add a SlicedSplitInfo the read instructions."""
# TODO(epot): Check that the number of examples per shard % 100 == 0
# Otherwise the slices value may be unbalanced and not exactly reflect the
# requested slice.
self._splits[sliced_split.split_info.name] = sliced_split
def __add__(self, other):
"""Merging split together."""
# Will raise error if a split has already be added (NonMutableDict)
# TODO(epot): If a split is already added but there is no overlap between
# the slices, should merge the slices (ex: [:10] + [80:])
split_instruction = SplitReadInstruction()
split_instruction._splits.update(self._splits) # pylint: disable=protected-access
split_instruction._splits.update(other._splits) # pylint: disable=protected-access
return split_instruction
def __getitem__(self, slice_value):
"""Sub-splits."""
# Will raise an error if a split has already been sliced
split_instruction = SplitReadInstruction()
for v in self._splits.values():
if v.slice_value is not None:
raise ValueError(f"Trying to slice Split {v.split_info.name} which has already been sliced")
v = v._asdict()
v["slice_value"] = slice_value
split_instruction.add(SlicedSplitInfo(**v))
return split_instruction
def get_list_sliced_split_info(self):
return list(self._splits.values())
class SplitDict(dict):
"""Split info object."""
def __init__(self, *args, dataset_name=None, **kwargs):
super().__init__(*args, **kwargs)
self.dataset_name = dataset_name
def __getitem__(self, key: Union[SplitBase, str]):
# 1st case: The key exists: `info.splits['train']`
if str(key) in self:
return super().__getitem__(str(key))
# 2nd case: Uses instructions: `info.splits['train[50%]']`
else:
instructions = make_file_instructions(
name=self.dataset_name,
split_infos=self.values(),
instruction=key,
)
return SubSplitInfo(instructions)
def __setitem__(self, key: Union[SplitBase, str], value: SplitInfo):
if key != value.name:
raise ValueError(f"Cannot add elem. (key mismatch: '{key}' != '{value.name}')")
if key in self:
raise ValueError(f"Split {key} already present")
super().__setitem__(key, value)
def add(self, split_info: SplitInfo):
"""Add the split info."""
if split_info.name in self:
raise ValueError(f"Split {split_info.name} already present")
split_info.dataset_name = self.dataset_name
super().__setitem__(split_info.name, split_info)
@property
def total_num_examples(self):
"""Return the total number of examples."""
return sum(s.num_examples for s in self.values())
@classmethod
def from_split_dict(cls, split_infos: Union[List, Dict], dataset_name: Optional[str] = None):
"""Returns a new SplitDict initialized from a Dict or List of `split_infos`."""
if isinstance(split_infos, dict):
split_infos = list(split_infos.values())
if dataset_name is None:
dataset_name = split_infos[0].get("dataset_name") if split_infos else None
split_dict = cls(dataset_name=dataset_name)
for split_info in split_infos:
if isinstance(split_info, dict):
split_info = SplitInfo(**split_info)
split_dict.add(split_info)
return split_dict
def to_split_dict(self):
"""Returns a list of SplitInfo protos that we have."""
out = []
for split_name, split_info in self.items():
split_info = copy.deepcopy(split_info)
split_info.name = split_name
out.append(split_info)
return out
def copy(self):
return SplitDict.from_split_dict(self.to_split_dict(), self.dataset_name)
def _to_yaml_list(self) -> list:
out = [asdict(s) for s in self.to_split_dict()]
# we don't need the shard lengths in YAML, since it depends on max_shard_size and num_proc
for split_info_dict in out:
split_info_dict.pop("shard_lengths", None)
# we don't need the dataset_name attribute that is deprecated
for split_info_dict in out:
split_info_dict.pop("dataset_name", None)
return out
@classmethod
def _from_yaml_list(cls, yaml_data: list) -> "SplitDict":
return cls.from_split_dict(yaml_data)
@dataclass
class SplitGenerator:
"""Defines the split information for the generator.
This should be used as returned value of
`GeneratorBasedBuilder._split_generators`.
See `GeneratorBasedBuilder._split_generators` for more info and example
of usage.
Args:
name (`str`):
Name of the `Split` for which the generator will
create the examples.
**gen_kwargs (additional keyword arguments):
Keyword arguments to forward to the `DatasetBuilder._generate_examples` method
of the builder.
Example:
```py
>>> datasets.SplitGenerator(
... name=datasets.Split.TRAIN,
... gen_kwargs={"split_key": "train", "files": dl_manager.download_and_extract(url)},
... )
```
"""
name: str
gen_kwargs: Dict = dataclasses.field(default_factory=dict)
split_info: SplitInfo = dataclasses.field(init=False)
def __post_init__(self):
self.name = str(self.name) # Make sure we convert NamedSplits in strings
NamedSplit(self.name) # check that it's a valid split name
self.split_info = SplitInfo(name=self.name)
| 0
|
hf_public_repos/datasets/src
|
hf_public_repos/datasets/src/datasets/search.py
|
import importlib.util
import os
import tempfile
from pathlib import PurePath
from typing import TYPE_CHECKING, Dict, List, NamedTuple, Optional, Union
import fsspec
import numpy as np
from .utils import logging
from .utils import tqdm as hf_tqdm
if TYPE_CHECKING:
from .arrow_dataset import Dataset # noqa: F401
try:
from elasticsearch import Elasticsearch # noqa: F401
except ImportError:
pass
try:
import faiss # noqa: F401
except ImportError:
pass
_has_elasticsearch = importlib.util.find_spec("elasticsearch") is not None
_has_faiss = importlib.util.find_spec("faiss") is not None
logger = logging.get_logger(__name__)
class MissingIndex(Exception):
pass
class SearchResults(NamedTuple):
scores: List[float]
indices: List[int]
class BatchedSearchResults(NamedTuple):
total_scores: List[List[float]]
total_indices: List[List[int]]
class NearestExamplesResults(NamedTuple):
scores: List[float]
examples: dict
class BatchedNearestExamplesResults(NamedTuple):
total_scores: List[List[float]]
total_examples: List[dict]
class BaseIndex:
"""Base class for indexing"""
def search(self, query, k: int = 10, **kwargs) -> SearchResults:
"""
To implement.
This method has to return the scores and the indices of the retrieved examples given a certain query.
"""
raise NotImplementedError
def search_batch(self, queries, k: int = 10, **kwargs) -> BatchedSearchResults:
"""Find the nearest examples indices to the query.
Args:
queries (`Union[List[str], np.ndarray]`): The queries as a list of strings if `column` is a text index or as a numpy array if `column` is a vector index.
k (`int`): The number of examples to retrieve per query.
Ouput:
total_scores (`List[List[float]`): The retrieval scores of the retrieved examples per query.
total_indices (`List[List[int]]`): The indices of the retrieved examples per query.
"""
total_scores, total_indices = [], []
for query in queries:
scores, indices = self.search(query, k)
total_scores.append(scores)
total_indices.append(indices)
return BatchedSearchResults(total_scores, total_indices)
def save(self, file: Union[str, PurePath]):
"""Serialize the index on disk"""
raise NotImplementedError
@classmethod
def load(cls, file: Union[str, PurePath]) -> "BaseIndex":
"""Deserialize the index from disk"""
raise NotImplementedError
class ElasticSearchIndex(BaseIndex):
"""
Sparse index using Elasticsearch. It is used to index text and run queries based on BM25 similarity.
An Elasticsearch server needs to be accessible, and a python client is declared with
```
es_client = Elasticsearch([{'host': 'localhost', 'port': '9200'}])
```
for example.
"""
def __init__(
self,
host: Optional[str] = None,
port: Optional[int] = None,
es_client: Optional["Elasticsearch"] = None,
es_index_name: Optional[str] = None,
es_index_config: Optional[dict] = None,
):
if not _has_elasticsearch:
raise ImportError(
"You must install ElasticSearch to use ElasticSearchIndex. To do so you can run `pip install elasticsearch==7.7.1 for example`"
)
if es_client is not None and (host is not None or port is not None):
raise ValueError("Please specify either `es_client` or `(host, port)`, but not both.")
host = host or "localhost"
port = port or 9200
import elasticsearch.helpers # noqa: F401 - need this to properly load all the es features
from elasticsearch import Elasticsearch # noqa: F811
self.es_client = es_client if es_client is not None else Elasticsearch([{"host": host, "port": str(port)}])
self.es_index_name = (
es_index_name
if es_index_name is not None
else "huggingface_datasets_" + os.path.basename(tempfile.NamedTemporaryFile().name)
)
self.es_index_config = (
es_index_config
if es_index_config is not None
else {
"settings": {
"number_of_shards": 1,
"analysis": {"analyzer": {"stop_standard": {"type": "standard", " stopwords": "_english_"}}},
},
"mappings": {"properties": {"text": {"type": "text", "analyzer": "standard", "similarity": "BM25"}}},
}
)
def add_documents(self, documents: Union[List[str], "Dataset"], column: Optional[str] = None):
"""
Add documents to the index.
If the documents are inside a certain column, you can specify it using the `column` argument.
"""
index_name = self.es_index_name
index_config = self.es_index_config
self.es_client.indices.create(index=index_name, body=index_config)
number_of_docs = len(documents)
progress = hf_tqdm(unit="docs", total=number_of_docs)
successes = 0
def passage_generator():
if column is not None:
for i, example in enumerate(documents):
yield {"text": example[column], "_id": i}
else:
for i, example in enumerate(documents):
yield {"text": example, "_id": i}
# create the ES index
import elasticsearch as es
for ok, action in es.helpers.streaming_bulk(
client=self.es_client,
index=index_name,
actions=passage_generator(),
):
progress.update(1)
successes += ok
if successes != len(documents):
logger.warning(
f"Some documents failed to be added to ElasticSearch. Failures: {len(documents)-successes}/{len(documents)}"
)
logger.info(f"Indexed {successes:d} documents")
def search(self, query: str, k=10, **kwargs) -> SearchResults:
"""Find the nearest examples indices to the query.
Args:
query (`str`): The query as a string.
k (`int`): The number of examples to retrieve.
Ouput:
scores (`List[List[float]`): The retrieval scores of the retrieved examples.
indices (`List[List[int]]`): The indices of the retrieved examples.
"""
response = self.es_client.search(
index=self.es_index_name,
body={"query": {"multi_match": {"query": query, "fields": ["text"], "type": "cross_fields"}}, "size": k},
**kwargs,
)
hits = response["hits"]["hits"]
return SearchResults([hit["_score"] for hit in hits], [int(hit["_id"]) for hit in hits])
def search_batch(self, queries, k: int = 10, max_workers=10, **kwargs) -> BatchedSearchResults:
import concurrent.futures
total_scores, total_indices = [None] * len(queries), [None] * len(queries)
with concurrent.futures.ThreadPoolExecutor(max_workers=max_workers) as executor:
future_to_index = {executor.submit(self.search, query, k, **kwargs): i for i, query in enumerate(queries)}
for future in concurrent.futures.as_completed(future_to_index):
index = future_to_index[future]
results: SearchResults = future.result()
total_scores[index] = results.scores
total_indices[index] = results.indices
return BatchedSearchResults(total_indices=total_indices, total_scores=total_scores)
class FaissIndex(BaseIndex):
"""
Dense index using Faiss. It is used to index vectors.
Faiss is a library for efficient similarity search and clustering of dense vectors.
It contains algorithms that search in sets of vectors of any size, up to ones that possibly do not fit in RAM.
You can find more information about Faiss here:
- For index types and the string factory: https://github.com/facebookresearch/faiss/wiki/The-index-factory
- For GPU settings: https://github.com/facebookresearch/faiss/wiki/Faiss-on-the-GPU
"""
def __init__(
self,
device: Optional[Union[int, List[int]]] = None,
string_factory: Optional[str] = None,
metric_type: Optional[int] = None,
custom_index: Optional["faiss.Index"] = None,
):
"""
Create a Dense index using Faiss. You can specify `device` if you want to run it on GPU (`device` must be the GPU index).
You can find more information about Faiss here:
- For `string factory`: https://github.com/facebookresearch/faiss/wiki/The-index-factory
"""
if string_factory is not None and custom_index is not None:
raise ValueError("Please specify either `string_factory` or `custom_index` but not both.")
if device is not None and custom_index is not None:
raise ValueError(
"Cannot pass both 'custom_index' and 'device'. "
"Pass 'custom_index' already transferred to the target device instead."
)
self.device = device
self.string_factory = string_factory
self.metric_type = metric_type
self.faiss_index = custom_index
if not _has_faiss:
raise ImportError(
"You must install Faiss to use FaissIndex. To do so you can run `conda install -c pytorch faiss-cpu` or `conda install -c pytorch faiss-gpu`. "
"A community supported package is also available on pypi: `pip install faiss-cpu` or `pip install faiss-gpu`. "
"Note that pip may not have the latest version of FAISS, and thus, some of the latest features and bug fixes may not be available."
)
def add_vectors(
self,
vectors: Union[np.array, "Dataset"],
column: Optional[str] = None,
batch_size: int = 1000,
train_size: Optional[int] = None,
faiss_verbose: Optional[bool] = None,
):
"""
Add vectors to the index.
If the arrays are inside a certain column, you can specify it using the `column` argument.
"""
import faiss # noqa: F811
# Create index
if self.faiss_index is None:
size = len(vectors[0]) if column is None else len(vectors[0][column])
if self.string_factory is not None:
if self.metric_type is None:
index = faiss.index_factory(size, self.string_factory)
else:
index = faiss.index_factory(size, self.string_factory, self.metric_type)
else:
if self.metric_type is None:
index = faiss.IndexFlat(size)
else:
index = faiss.IndexFlat(size, self.metric_type)
self.faiss_index = self._faiss_index_to_device(index, self.device)
logger.info(f"Created faiss index of type {type(self.faiss_index)}")
# Set verbosity level
if faiss_verbose is not None:
self.faiss_index.verbose = faiss_verbose
if hasattr(self.faiss_index, "index") and self.faiss_index.index is not None:
self.faiss_index.index.verbose = faiss_verbose
if hasattr(self.faiss_index, "quantizer") and self.faiss_index.quantizer is not None:
self.faiss_index.quantizer.verbose = faiss_verbose
if hasattr(self.faiss_index, "clustering_index") and self.faiss_index.clustering_index is not None:
self.faiss_index.clustering_index.verbose = faiss_verbose
# Train
if train_size is not None:
train_vecs = vectors[:train_size] if column is None else vectors[:train_size][column]
logger.info(f"Training the index with the first {len(train_vecs)} vectors")
self.faiss_index.train(train_vecs)
else:
logger.info("Ignored the training step of the faiss index as `train_size` is None.")
# Add vectors
logger.info(f"Adding {len(vectors)} vectors to the faiss index")
for i in hf_tqdm(range(0, len(vectors), batch_size)):
vecs = vectors[i : i + batch_size] if column is None else vectors[i : i + batch_size][column]
self.faiss_index.add(vecs)
@staticmethod
def _faiss_index_to_device(index: "faiss.Index", device: Optional[Union[int, List[int]]] = None) -> "faiss.Index":
"""
Sends a faiss index to a device.
A device can either be a positive integer (GPU id), a negative integer (all GPUs),
or a list of positive integers (select GPUs to use), or `None` for CPU.
"""
# If device is not specified, then it runs on CPU.
if device is None:
return index
import faiss # noqa: F811
# If the device id is given as an integer
if isinstance(device, int):
# Positive integers are directly mapped to GPU ids
if device > -1:
faiss_res = faiss.StandardGpuResources()
index = faiss.index_cpu_to_gpu(faiss_res, device, index)
# And negative integers mean using all GPUs
else:
index = faiss.index_cpu_to_all_gpus(index)
# Device ids given as a list mean mapping to those devices specified.
elif isinstance(device, (list, tuple)):
index = faiss.index_cpu_to_gpus_list(index, gpus=list(device))
else:
raise TypeError(
f"The argument type: {type(device)} is not expected. "
+ "Please pass in either nothing, a positive int, a negative int, or a list of positive ints."
)
return index
def search(self, query: np.array, k=10, **kwargs) -> SearchResults:
"""Find the nearest examples indices to the query.
Args:
query (`np.array`): The query as a numpy array.
k (`int`): The number of examples to retrieve.
Ouput:
scores (`List[List[float]`): The retrieval scores of the retrieved examples.
indices (`List[List[int]]`): The indices of the retrieved examples.
"""
if len(query.shape) != 1 and (len(query.shape) != 2 or query.shape[0] != 1):
raise ValueError("Shape of query is incorrect, it has to be either a 1D array or 2D (1, N)")
queries = query.reshape(1, -1)
if not queries.flags.c_contiguous:
queries = np.asarray(queries, order="C")
scores, indices = self.faiss_index.search(queries, k, **kwargs)
return SearchResults(scores[0], indices[0].astype(int))
def search_batch(self, queries: np.array, k=10, **kwargs) -> BatchedSearchResults:
"""Find the nearest examples indices to the queries.
Args:
queries (`np.array`): The queries as a numpy array.
k (`int`): The number of examples to retrieve.
Ouput:
total_scores (`List[List[float]`): The retrieval scores of the retrieved examples per query.
total_indices (`List[List[int]]`): The indices of the retrieved examples per query.
"""
if len(queries.shape) != 2:
raise ValueError("Shape of query must be 2D")
if not queries.flags.c_contiguous:
queries = np.asarray(queries, order="C")
scores, indices = self.faiss_index.search(queries, k, **kwargs)
return BatchedSearchResults(scores, indices.astype(int))
def save(self, file: Union[str, PurePath], storage_options: Optional[Dict] = None):
"""Serialize the FaissIndex on disk"""
import faiss # noqa: F811
if self.device is not None and isinstance(self.device, (int, list, tuple)):
index = faiss.index_gpu_to_cpu(self.faiss_index)
else:
index = self.faiss_index
with fsspec.open(str(file), "wb", **(storage_options or {})) as f:
faiss.write_index(index, faiss.BufferedIOWriter(faiss.PyCallbackIOWriter(f.write)))
@classmethod
def load(
cls,
file: Union[str, PurePath],
device: Optional[Union[int, List[int]]] = None,
storage_options: Optional[Dict] = None,
) -> "FaissIndex":
"""Deserialize the FaissIndex from disk"""
import faiss # noqa: F811
# Instances of FaissIndex is essentially just a wrapper for faiss indices.
faiss_index = cls(device=device)
with fsspec.open(str(file), "rb", **(storage_options or {})) as f:
index = faiss.read_index(faiss.BufferedIOReader(faiss.PyCallbackIOReader(f.read)))
faiss_index.faiss_index = faiss_index._faiss_index_to_device(index, faiss_index.device)
return faiss_index
class IndexableMixin:
"""Add indexing features to `datasets.Dataset`"""
def __init__(self):
self._indexes: Dict[str, BaseIndex] = {}
def __len__(self):
raise NotImplementedError
def __getitem__(self, key):
raise NotImplementedError
def is_index_initialized(self, index_name: str) -> bool:
return index_name in self._indexes
def _check_index_is_initialized(self, index_name: str):
if not self.is_index_initialized(index_name):
raise MissingIndex(
f"Index with index_name '{index_name}' not initialized yet. Please make sure that you call `add_faiss_index` or `add_elasticsearch_index` first."
)
def list_indexes(self) -> List[str]:
"""List the `colindex_nameumns`/identifiers of all the attached indexes."""
return list(self._indexes)
def get_index(self, index_name: str) -> BaseIndex:
"""List the `index_name`/identifiers of all the attached indexes.
Args:
index_name (`str`): Index name.
Returns:
[`BaseIndex`]
"""
self._check_index_is_initialized(index_name)
return self._indexes[index_name]
def add_faiss_index(
self,
column: str,
index_name: Optional[str] = None,
device: Optional[Union[int, List[int]]] = None,
string_factory: Optional[str] = None,
metric_type: Optional[int] = None,
custom_index: Optional["faiss.Index"] = None,
batch_size: int = 1000,
train_size: Optional[int] = None,
faiss_verbose: bool = False,
):
"""Add a dense index using Faiss for fast retrieval.
The index is created using the vectors of the specified column.
You can specify `device` if you want to run it on GPU (`device` must be the GPU index, see more below).
You can find more information about Faiss here:
- For `string factory`: https://github.com/facebookresearch/faiss/wiki/The-index-factory
Args:
column (`str`): The column of the vectors to add to the index.
index_name (Optional `str`): The index_name/identifier of the index. This is the index_name that is used to call `.get_nearest` or `.search`.
By default it corresponds to `column`.
device (Optional `Union[int, List[int]]`): If positive integer, this is the index of the GPU to use. If negative integer, use all GPUs.
If a list of positive integers is passed in, run only on those GPUs. By default it uses the CPU.
string_factory (Optional `str`): This is passed to the index factory of Faiss to create the index. Default index class is IndexFlatIP.
metric_type (Optional `int`): Type of metric. Ex: `faiss.METRIC_INNER_PRODUCT` or `faiss.METRIC_L2`.
custom_index (Optional `faiss.Index`): Custom Faiss index that you already have instantiated and configured for your needs.
batch_size (Optional `int`): Size of the batch to use while adding vectors to the FaissIndex. Default value is 1000.
<Added version="2.4.0"/>
train_size (Optional `int`): If the index needs a training step, specifies how many vectors will be used to train the index.
faiss_verbose (`bool`, defaults to False): Enable the verbosity of the Faiss index.
"""
index_name = index_name if index_name is not None else column
faiss_index = FaissIndex(
device=device, string_factory=string_factory, metric_type=metric_type, custom_index=custom_index
)
faiss_index.add_vectors(
self, column=column, batch_size=batch_size, train_size=train_size, faiss_verbose=faiss_verbose
)
self._indexes[index_name] = faiss_index
def add_faiss_index_from_external_arrays(
self,
external_arrays: np.array,
index_name: str,
device: Optional[Union[int, List[int]]] = None,
string_factory: Optional[str] = None,
metric_type: Optional[int] = None,
custom_index: Optional["faiss.Index"] = None,
batch_size: int = 1000,
train_size: Optional[int] = None,
faiss_verbose: bool = False,
):
"""Add a dense index using Faiss for fast retrieval.
The index is created using the vectors of `external_arrays`.
You can specify `device` if you want to run it on GPU (`device` must be the GPU index).
You can find more information about Faiss here:
- For `string factory`: https://github.com/facebookresearch/faiss/wiki/The-index-factory
Args:
external_arrays (`np.array`): If you want to use arrays from outside the lib for the index, you can set `external_arrays`.
It will use `external_arrays` to create the Faiss index instead of the arrays in the given `column`.
index_name (`str`): The index_name/identifier of the index. This is the index_name that is used to call `.get_nearest` or `.search`.
device (Optional `Union[int, List[int]]`): If positive integer, this is the index of the GPU to use. If negative integer, use all GPUs.
If a list of positive integers is passed in, run only on those GPUs. By default it uses the CPU.
string_factory (Optional `str`): This is passed to the index factory of Faiss to create the index. Default index class is IndexFlatIP.
metric_type (Optional `int`): Type of metric. Ex: `faiss.METRIC_INNER_PRODUCT` or `faiss.METRIC_L2`.
custom_index (Optional `faiss.Index`): Custom Faiss index that you already have instantiated and configured for your needs.
batch_size (Optional `int`): Size of the batch to use while adding vectors to the FaissIndex. Default value is 1000.
<Added version="2.4.0"/>
train_size (Optional `int`): If the index needs a training step, specifies how many vectors will be used to train the index.
faiss_verbose (`bool`, defaults to False): Enable the verbosity of the Faiss index.
"""
faiss_index = FaissIndex(
device=device, string_factory=string_factory, metric_type=metric_type, custom_index=custom_index
)
faiss_index.add_vectors(
external_arrays, column=None, batch_size=batch_size, train_size=train_size, faiss_verbose=faiss_verbose
)
self._indexes[index_name] = faiss_index
def save_faiss_index(self, index_name: str, file: Union[str, PurePath], storage_options: Optional[Dict] = None):
"""Save a FaissIndex on disk.
Args:
index_name (`str`): The index_name/identifier of the index. This is the index_name that is used to call `.get_nearest` or `.search`.
file (`str`): The path to the serialized faiss index on disk or remote URI (e.g. `"s3://my-bucket/index.faiss"`).
storage_options (`dict`, *optional*):
Key/value pairs to be passed on to the file-system backend, if any.
<Added version="2.11.0"/>
"""
index = self.get_index(index_name)
if not isinstance(index, FaissIndex):
raise ValueError(f"Index '{index_name}' is not a FaissIndex but a '{type(index)}'")
index.save(file, storage_options=storage_options)
logger.info(f"Saved FaissIndex {index_name} at {file}")
def load_faiss_index(
self,
index_name: str,
file: Union[str, PurePath],
device: Optional[Union[int, List[int]]] = None,
storage_options: Optional[Dict] = None,
):
"""Load a FaissIndex from disk.
If you want to do additional configurations, you can have access to the faiss index object by doing
`.get_index(index_name).faiss_index` to make it fit your needs.
Args:
index_name (`str`): The index_name/identifier of the index. This is the index_name that is used to
call `.get_nearest` or `.search`.
file (`str`): The path to the serialized faiss index on disk or remote URI (e.g. `"s3://my-bucket/index.faiss"`).
device (Optional `Union[int, List[int]]`): If positive integer, this is the index of the GPU to use. If negative integer, use all GPUs.
If a list of positive integers is passed in, run only on those GPUs. By default it uses the CPU.
storage_options (`dict`, *optional*):
Key/value pairs to be passed on to the file-system backend, if any.
<Added version="2.11.0"/>
"""
index = FaissIndex.load(file, device=device, storage_options=storage_options)
if index.faiss_index.ntotal != len(self):
raise ValueError(
f"Index size should match Dataset size, but Index '{index_name}' at {file} has {index.faiss_index.ntotal} elements while the dataset has {len(self)} examples."
)
self._indexes[index_name] = index
logger.info(f"Loaded FaissIndex {index_name} from {file}")
def add_elasticsearch_index(
self,
column: str,
index_name: Optional[str] = None,
host: Optional[str] = None,
port: Optional[int] = None,
es_client: Optional["Elasticsearch"] = None,
es_index_name: Optional[str] = None,
es_index_config: Optional[dict] = None,
):
"""Add a text index using ElasticSearch for fast retrieval.
Args:
column (`str`): The column of the documents to add to the index.
index_name (Optional `str`): The index_name/identifier of the index. This is the index name that is used to call `.get_nearest` or `.search`.
By default it corresponds to `column`.
host (Optional `str`, defaults to localhost):
host of where ElasticSearch is running
port (Optional `str`, defaults to 9200):
port of where ElasticSearch is running
es_client (Optional `elasticsearch.Elasticsearch`):
The elasticsearch client used to create the index if host and port are None.
es_index_name (Optional `str`): The elasticsearch index name used to create the index.
es_index_config (Optional `dict`):
The configuration of the elasticsearch index.
Default config is:
Config::
{
"settings": {
"number_of_shards": 1,
"analysis": {"analyzer": {"stop_standard": {"type": "standard", " stopwords": "_english_"}}},
},
"mappings": {
"properties": {
"text": {
"type": "text",
"analyzer": "standard",
"similarity": "BM25"
},
}
},
}
"""
index_name = index_name if index_name is not None else column
es_index = ElasticSearchIndex(
host=host, port=port, es_client=es_client, es_index_name=es_index_name, es_index_config=es_index_config
)
es_index.add_documents(self, column=column)
self._indexes[index_name] = es_index
def load_elasticsearch_index(
self,
index_name: str,
es_index_name: str,
host: Optional[str] = None,
port: Optional[int] = None,
es_client: Optional["Elasticsearch"] = None,
es_index_config: Optional[dict] = None,
):
"""Load an existing text index using ElasticSearch for fast retrieval.
Args:
index_name (`str`):
The `index_name`/identifier of the index. This is the index name that is used to call `get_nearest` or `search`.
es_index_name (`str`):
The name of elasticsearch index to load.
host (`str`, *optional*, defaults to `localhost`):
Host of where ElasticSearch is running.
port (`str`, *optional*, defaults to `9200`):
Port of where ElasticSearch is running.
es_client (`elasticsearch.Elasticsearch`, *optional*):
The elasticsearch client used to create the index if host and port are `None`.
es_index_config (`dict`, *optional*):
The configuration of the elasticsearch index.
Default config is:
```
{
"settings": {
"number_of_shards": 1,
"analysis": {"analyzer": {"stop_standard": {"type": "standard", " stopwords": "_english_"}}},
},
"mappings": {
"properties": {
"text": {
"type": "text",
"analyzer": "standard",
"similarity": "BM25"
},
}
},
}
```
"""
self._indexes[index_name] = ElasticSearchIndex(
host=host, port=port, es_client=es_client, es_index_name=es_index_name, es_index_config=es_index_config
)
def drop_index(self, index_name: str):
"""Drop the index with the specified column.
Args:
index_name (`str`):
The `index_name`/identifier of the index.
"""
del self._indexes[index_name]
def search(self, index_name: str, query: Union[str, np.array], k: int = 10, **kwargs) -> SearchResults:
"""Find the nearest examples indices in the dataset to the query.
Args:
index_name (`str`):
The name/identifier of the index.
query (`Union[str, np.ndarray]`):
The query as a string if `index_name` is a text index or as a numpy array if `index_name` is a vector index.
k (`int`):
The number of examples to retrieve.
Returns:
`(scores, indices)`:
A tuple of `(scores, indices)` where:
- **scores** (`List[List[float]`): the retrieval scores from either FAISS (`IndexFlatL2` by default) or ElasticSearch of the retrieved examples
- **indices** (`List[List[int]]`): the indices of the retrieved examples
"""
self._check_index_is_initialized(index_name)
return self._indexes[index_name].search(query, k, **kwargs)
def search_batch(
self, index_name: str, queries: Union[List[str], np.array], k: int = 10, **kwargs
) -> BatchedSearchResults:
"""Find the nearest examples indices in the dataset to the query.
Args:
index_name (`str`):
The `index_name`/identifier of the index.
queries (`Union[List[str], np.ndarray]`):
The queries as a list of strings if `index_name` is a text index or as a numpy array if `index_name` is a vector index.
k (`int`):
The number of examples to retrieve per query.
Returns:
`(total_scores, total_indices)`:
A tuple of `(total_scores, total_indices)` where:
- **total_scores** (`List[List[float]`): the retrieval scores from either FAISS (`IndexFlatL2` by default) or ElasticSearch of the retrieved examples per query
- **total_indices** (`List[List[int]]`): the indices of the retrieved examples per query
"""
self._check_index_is_initialized(index_name)
return self._indexes[index_name].search_batch(queries, k, **kwargs)
def get_nearest_examples(
self, index_name: str, query: Union[str, np.array], k: int = 10, **kwargs
) -> NearestExamplesResults:
"""Find the nearest examples in the dataset to the query.
Args:
index_name (`str`):
The index_name/identifier of the index.
query (`Union[str, np.ndarray]`):
The query as a string if `index_name` is a text index or as a numpy array if `index_name` is a vector index.
k (`int`):
The number of examples to retrieve.
Returns:
`(scores, examples)`:
A tuple of `(scores, examples)` where:
- **scores** (`List[float]`): the retrieval scores from either FAISS (`IndexFlatL2` by default) or ElasticSearch of the retrieved examples
- **examples** (`dict`): the retrieved examples
"""
self._check_index_is_initialized(index_name)
scores, indices = self.search(index_name, query, k, **kwargs)
top_indices = [i for i in indices if i >= 0]
return NearestExamplesResults(scores[: len(top_indices)], self[top_indices])
def get_nearest_examples_batch(
self, index_name: str, queries: Union[List[str], np.array], k: int = 10, **kwargs
) -> BatchedNearestExamplesResults:
"""Find the nearest examples in the dataset to the query.
Args:
index_name (`str`):
The `index_name`/identifier of the index.
queries (`Union[List[str], np.ndarray]`):
The queries as a list of strings if `index_name` is a text index or as a numpy array if `index_name` is a vector index.
k (`int`):
The number of examples to retrieve per query.
Returns:
`(total_scores, total_examples)`:
A tuple of `(total_scores, total_examples)` where:
- **total_scores** (`List[List[float]`): the retrieval scores from either FAISS (`IndexFlatL2` by default) or ElasticSearch of the retrieved examples per query
- **total_examples** (`List[dict]`): the retrieved examples per query
"""
self._check_index_is_initialized(index_name)
total_scores, total_indices = self.search_batch(index_name, queries, k, **kwargs)
total_scores = [
scores_i[: len([i for i in indices_i if i >= 0])]
for scores_i, indices_i in zip(total_scores, total_indices)
]
total_samples = [self[[i for i in indices if i >= 0]] for indices in total_indices]
return BatchedNearestExamplesResults(total_scores, total_samples)
| 0
|
hf_public_repos/datasets/src
|
hf_public_repos/datasets/src/datasets/naming.py
|
# Copyright 2020 The HuggingFace Datasets Authors and the TensorFlow Datasets Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# Lint as: python3
"""Utilities for file names."""
import itertools
import os
import re
_uppercase_uppercase_re = re.compile(r"([A-Z]+)([A-Z][a-z])")
_lowercase_uppercase_re = re.compile(r"([a-z\d])([A-Z])")
_single_underscore_re = re.compile(r"(?<!_)_(?!_)")
_multiple_underscores_re = re.compile(r"(_{2,})")
_split_re = r"^\w+(\.\w+)*$"
INVALID_WINDOWS_CHARACTERS_IN_PATH = r"<>:/\|?*"
def camelcase_to_snakecase(name):
"""Convert camel-case string to snake-case."""
name = _uppercase_uppercase_re.sub(r"\1_\2", name)
name = _lowercase_uppercase_re.sub(r"\1_\2", name)
return name.lower()
def snakecase_to_camelcase(name):
"""Convert snake-case string to camel-case string."""
name = _single_underscore_re.split(name)
name = [_multiple_underscores_re.split(n) for n in name]
return "".join(n.capitalize() for n in itertools.chain.from_iterable(name) if n != "")
def filename_prefix_for_name(name):
if os.path.basename(name) != name:
raise ValueError(f"Should be a dataset name, not a path: {name}")
return camelcase_to_snakecase(name)
def filename_prefix_for_split(name, split):
if os.path.basename(name) != name:
raise ValueError(f"Should be a dataset name, not a path: {name}")
if not re.match(_split_re, split):
raise ValueError(f"Split name should match '{_split_re}'' but got '{split}'.")
return f"{filename_prefix_for_name(name)}-{split}"
def filepattern_for_dataset_split(dataset_name, split, data_dir, filetype_suffix=None):
prefix = filename_prefix_for_split(dataset_name, split)
if filetype_suffix:
prefix += f".{filetype_suffix}"
filepath = os.path.join(data_dir, prefix)
return f"{filepath}*"
def filenames_for_dataset_split(path, dataset_name, split, filetype_suffix=None, shard_lengths=None):
prefix = filename_prefix_for_split(dataset_name, split)
prefix = os.path.join(path, prefix)
if shard_lengths:
num_shards = len(shard_lengths)
filenames = [f"{prefix}-{shard_id:05d}-of-{num_shards:05d}" for shard_id in range(num_shards)]
if filetype_suffix:
filenames = [filename + f".{filetype_suffix}" for filename in filenames]
return filenames
else:
filename = prefix
if filetype_suffix:
filename += f".{filetype_suffix}"
return [filename]
| 0
|
hf_public_repos/datasets/src
|
hf_public_repos/datasets/src/datasets/inspect.py
|
# Copyright 2020 The HuggingFace Datasets Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# Lint as: python3
""" List and inspect datasets."""
import inspect
import os
import shutil
import warnings
from pathlib import PurePath
from typing import Dict, List, Mapping, Optional, Sequence, Union
import huggingface_hub
from .download.download_config import DownloadConfig
from .download.download_manager import DownloadMode
from .download.streaming_download_manager import StreamingDownloadManager
from .info import DatasetInfo
from .load import (
dataset_module_factory,
get_dataset_builder_class,
import_main_class,
load_dataset_builder,
metric_module_factory,
)
from .utils.deprecation_utils import deprecated
from .utils.file_utils import relative_to_absolute_path
from .utils.logging import get_logger
from .utils.version import Version
logger = get_logger(__name__)
class SplitsNotFoundError(ValueError):
pass
@deprecated("Use 'huggingface_hub.list_datasets' instead.")
def list_datasets(with_community_datasets=True, with_details=False):
"""List all the datasets scripts available on the Hugging Face Hub.
Args:
with_community_datasets (`bool`, *optional*, defaults to `True`):
Include the community provided datasets.
with_details (`bool`, *optional*, defaults to `False`):
Return the full details on the datasets instead of only the short name.
Example:
```py
>>> from datasets import list_datasets
>>> list_datasets()
['acronym_identification',
'ade_corpus_v2',
'adversarial_qa',
'aeslc',
'afrikaans_ner_corpus',
'ag_news',
...
]
```
"""
datasets = huggingface_hub.list_datasets(full=with_details)
if not with_community_datasets:
datasets = [dataset for dataset in datasets if "/" not in dataset.id]
if not with_details:
datasets = [dataset.id for dataset in datasets]
return list(datasets)
@deprecated(
"Use 'evaluate.list_evaluation_modules' instead, from the new library 🤗 Evaluate: https://huggingface.co/docs/evaluate"
)
def list_metrics(with_community_metrics=True, with_details=False):
"""List all the metrics script available on the Hugging Face Hub.
<Deprecated version="2.5.0">
Use `evaluate.list_evaluation_modules` instead, from the new library 🤗 Evaluate: https://huggingface.co/docs/evaluate
</Deprecated>
Args:
with_community_metrics (:obj:`bool`, optional, default ``True``): Include the community provided metrics.
with_details (:obj:`bool`, optional, default ``False``): Return the full details on the metrics instead of only the short name.
Example:
```py
>>> from datasets import list_metrics
>>> list_metrics()
['accuracy',
'bertscore',
'bleu',
'bleurt',
'cer',
'chrf',
...
]
```
"""
metrics = huggingface_hub.list_metrics()
if not with_community_metrics:
metrics = [metric for metric in metrics if "/" not in metric.id]
if not with_details:
metrics = [metric.id for metric in metrics]
return metrics
@deprecated("Clone the dataset repository from the Hugging Face Hub instead.")
def inspect_dataset(path: str, local_path: str, download_config: Optional[DownloadConfig] = None, **download_kwargs):
"""
Allow inspection/modification of a dataset script by copying on local drive at local_path.
Args:
path (`str`): Path to the dataset processing script with the dataset builder. Can be either:
- a local path to processing script or the directory containing the script (if the script has the same name
as the directory),
e.g. `'./dataset/squad'` or `'./dataset/squad/squad.py'`.
- a dataset identifier on the Hugging Face Hub (list all available datasets and ids with [`list_datasets`])
e.g. `'squad'`, `'glue'` or `'openai/webtext'`.
local_path (`str`):
Path to the local folder to copy the dataset script to.
download_config ([`DownloadConfig`], *optional*):
Specific download configuration parameters.
**download_kwargs (additional keyword arguments):
Optional arguments for [`DownloadConfig`] which will override
the attributes of `download_config` if supplied.
"""
dataset_module = dataset_module_factory(path, download_config=download_config, **download_kwargs)
builder_cls = get_dataset_builder_class(dataset_module)
module_source_path = inspect.getsourcefile(builder_cls)
module_source_dirpath = os.path.dirname(module_source_path)
for dirpath, dirnames, filenames in os.walk(module_source_dirpath):
dst_dirpath = os.path.join(local_path, os.path.relpath(dirpath, module_source_dirpath))
os.makedirs(dst_dirpath, exist_ok=True)
# skipping hidden directories; prune the search
# [:] for the in-place list modification required by os.walk
dirnames[:] = [dirname for dirname in dirnames if not dirname.startswith((".", "__"))]
for filename in filenames:
shutil.copy2(os.path.join(dirpath, filename), os.path.join(dst_dirpath, filename))
shutil.copystat(dirpath, dst_dirpath)
local_path = relative_to_absolute_path(local_path)
print(
f"The processing script for dataset {path} can be inspected at {local_path}. "
f"The main class is in {module_source_dirpath}. "
f'You can modify this processing script and use it with `datasets.load_dataset("{PurePath(local_path).as_posix()}")`.'
)
@deprecated(
"Use 'evaluate.inspect_evaluation_module' instead, from the new library 🤗 Evaluate: https://huggingface.co/docs/evaluate"
)
def inspect_metric(path: str, local_path: str, download_config: Optional[DownloadConfig] = None, **download_kwargs):
r"""
Allow inspection/modification of a metric script by copying it on local drive at local_path.
<Deprecated version="2.5.0">
Use `evaluate.inspect_evaluation_module` instead, from the new library 🤗 Evaluate instead: https://huggingface.co/docs/evaluate
</Deprecated>
Args:
path (``str``): path to the dataset processing script with the dataset builder. Can be either:
- a local path to processing script or the directory containing the script (if the script has the same name as the directory),
e.g. ``'./dataset/squad'`` or ``'./dataset/squad/squad.py'``
- a dataset identifier on the Hugging Face Hub (list all available datasets and ids with ``datasets.list_datasets()``)
e.g. ``'squad'``, ``'glue'`` or ``'openai/webtext'``
local_path (``str``): path to the local folder to copy the datset script to.
download_config (Optional ``datasets.DownloadConfig``): specific download configuration parameters.
**download_kwargs (additional keyword arguments): optional attributes for DownloadConfig() which will override the attributes in download_config if supplied.
"""
metric_module = metric_module_factory(path, download_config=download_config, **download_kwargs)
metric_cls = import_main_class(metric_module.module_path, dataset=False)
module_source_path = inspect.getsourcefile(metric_cls)
module_source_dirpath = os.path.dirname(module_source_path)
for dirpath, dirnames, filenames in os.walk(module_source_dirpath):
dst_dirpath = os.path.join(local_path, os.path.relpath(dirpath, module_source_dirpath))
os.makedirs(dst_dirpath, exist_ok=True)
# skipping hidden directories; prune the search
dirnames[:] = [dirname for dirname in dirnames if not dirname.startswith((".", "__"))]
for filename in filenames:
shutil.copy2(os.path.join(dirpath, filename), os.path.join(dst_dirpath, filename))
shutil.copystat(dirpath, dst_dirpath)
local_path = relative_to_absolute_path(local_path)
print(
f"The processing scripts for metric {path} can be inspected at {local_path}. "
f"The main class is in {module_source_dirpath}. "
f'You can modify this processing scripts and use it with `datasets.load_metric("{PurePath(local_path).as_posix()}")`.'
)
def get_dataset_infos(
path: str,
data_files: Optional[Union[Dict, List, str]] = None,
download_config: Optional[DownloadConfig] = None,
download_mode: Optional[Union[DownloadMode, str]] = None,
revision: Optional[Union[str, Version]] = None,
token: Optional[Union[bool, str]] = None,
use_auth_token="deprecated",
**config_kwargs,
):
"""Get the meta information about a dataset, returned as a dict mapping config name to DatasetInfoDict.
Args:
path (`str`): path to the dataset processing script with the dataset builder. Can be either:
- a local path to processing script or the directory containing the script (if the script has the same name as the directory),
e.g. `'./dataset/squad'` or `'./dataset/squad/squad.py'`
- a dataset identifier on the Hugging Face Hub (list all available datasets and ids with [`datasets.list_datasets`])
e.g. `'squad'`, `'glue'` or``'openai/webtext'`
revision (`Union[str, datasets.Version]`, *optional*):
If specified, the dataset module will be loaded from the datasets repository at this version.
By default:
- it is set to the local version of the lib.
- it will also try to load it from the main branch if it's not available at the local version of the lib.
Specifying a version that is different from your local version of the lib might cause compatibility issues.
download_config ([`DownloadConfig`], *optional*):
Specific download configuration parameters.
download_mode ([`DownloadMode`] or `str`, defaults to `REUSE_DATASET_IF_EXISTS`):
Download/generate mode.
data_files (`Union[Dict, List, str]`, *optional*):
Defining the data_files of the dataset configuration.
token (`str` or `bool`, *optional*):
Optional string or boolean to use as Bearer token for remote files on the Datasets Hub.
If `True`, or not specified, will get token from `"~/.huggingface"`.
use_auth_token (`str` or `bool`, *optional*):
Optional string or boolean to use as Bearer token for remote files on the Datasets Hub.
If `True`, or not specified, will get token from `"~/.huggingface"`.
<Deprecated version="2.14.0">
`use_auth_token` was deprecated in favor of `token` in version 2.14.0 and will be removed in 3.0.0.
</Deprecated>
**config_kwargs (additional keyword arguments):
Optional attributes for builder class which will override the attributes if supplied.
Example:
```py
>>> from datasets import get_dataset_infos
>>> get_dataset_infos('rotten_tomatoes')
{'default': DatasetInfo(description="Movie Review Dataset.\nThis is a dataset of containing 5,331 positive and 5,331 negative processed\nsentences from Rotten Tomatoes movie reviews...), ...}
```
"""
if use_auth_token != "deprecated":
warnings.warn(
"'use_auth_token' was deprecated in favor of 'token' in version 2.14.0 and will be removed in 3.0.0.\n"
"You can remove this warning by passing 'token=<use_auth_token>' instead.",
FutureWarning,
)
token = use_auth_token
config_names = get_dataset_config_names(
path=path,
revision=revision,
download_config=download_config,
download_mode=download_mode,
data_files=data_files,
token=token,
)
return {
config_name: get_dataset_config_info(
path=path,
config_name=config_name,
data_files=data_files,
download_config=download_config,
download_mode=download_mode,
revision=revision,
token=token,
**config_kwargs,
)
for config_name in config_names
}
def get_dataset_config_names(
path: str,
revision: Optional[Union[str, Version]] = None,
download_config: Optional[DownloadConfig] = None,
download_mode: Optional[Union[DownloadMode, str]] = None,
dynamic_modules_path: Optional[str] = None,
data_files: Optional[Union[Dict, List, str]] = None,
**download_kwargs,
):
"""Get the list of available config names for a particular dataset.
Args:
path (`str`): path to the dataset processing script with the dataset builder. Can be either:
- a local path to processing script or the directory containing the script (if the script has the same name as the directory),
e.g. `'./dataset/squad'` or `'./dataset/squad/squad.py'`
- a dataset identifier on the Hugging Face Hub (list all available datasets and ids with [`datasets.list_datasets`])
e.g. `'squad'`, `'glue'` or `'openai/webtext'`
revision (`Union[str, datasets.Version]`, *optional*):
If specified, the dataset module will be loaded from the datasets repository at this version.
By default:
- it is set to the local version of the lib.
- it will also try to load it from the main branch if it's not available at the local version of the lib.
Specifying a version that is different from your local version of the lib might cause compatibility issues.
download_config ([`DownloadConfig`], *optional*):
Specific download configuration parameters.
download_mode ([`DownloadMode`] or `str`, defaults to `REUSE_DATASET_IF_EXISTS`):
Download/generate mode.
dynamic_modules_path (`str`, defaults to `~/.cache/huggingface/modules/datasets_modules`):
Optional path to the directory in which the dynamic modules are saved. It must have been initialized with `init_dynamic_modules`.
By default the datasets and metrics are stored inside the `datasets_modules` module.
data_files (`Union[Dict, List, str]`, *optional*):
Defining the data_files of the dataset configuration.
**download_kwargs (additional keyword arguments):
Optional attributes for [`DownloadConfig`] which will override the attributes in `download_config` if supplied,
for example `token`.
Example:
```py
>>> from datasets import get_dataset_config_names
>>> get_dataset_config_names("glue")
['cola',
'sst2',
'mrpc',
'qqp',
'stsb',
'mnli',
'mnli_mismatched',
'mnli_matched',
'qnli',
'rte',
'wnli',
'ax']
```
"""
dataset_module = dataset_module_factory(
path,
revision=revision,
download_config=download_config,
download_mode=download_mode,
dynamic_modules_path=dynamic_modules_path,
data_files=data_files,
**download_kwargs,
)
builder_cls = get_dataset_builder_class(dataset_module, dataset_name=os.path.basename(path))
return list(builder_cls.builder_configs.keys()) or [
dataset_module.builder_kwargs.get("config_name", builder_cls.DEFAULT_CONFIG_NAME or "default")
]
def get_dataset_config_info(
path: str,
config_name: Optional[str] = None,
data_files: Optional[Union[str, Sequence[str], Mapping[str, Union[str, Sequence[str]]]]] = None,
download_config: Optional[DownloadConfig] = None,
download_mode: Optional[Union[DownloadMode, str]] = None,
revision: Optional[Union[str, Version]] = None,
token: Optional[Union[bool, str]] = None,
use_auth_token="deprecated",
**config_kwargs,
) -> DatasetInfo:
"""Get the meta information (DatasetInfo) about a dataset for a particular config
Args:
path (``str``): path to the dataset processing script with the dataset builder. Can be either:
- a local path to processing script or the directory containing the script (if the script has the same name as the directory),
e.g. ``'./dataset/squad'`` or ``'./dataset/squad/squad.py'``
- a dataset identifier on the Hugging Face Hub (list all available datasets and ids with ``datasets.list_datasets()``)
e.g. ``'squad'``, ``'glue'`` or ``'openai/webtext'``
config_name (:obj:`str`, optional): Defining the name of the dataset configuration.
data_files (:obj:`str` or :obj:`Sequence` or :obj:`Mapping`, optional): Path(s) to source data file(s).
download_config (:class:`~download.DownloadConfig`, optional): Specific download configuration parameters.
download_mode (:class:`DownloadMode` or :obj:`str`, default ``REUSE_DATASET_IF_EXISTS``): Download/generate mode.
revision (:class:`~utils.Version` or :obj:`str`, optional): Version of the dataset script to load.
As datasets have their own git repository on the Datasets Hub, the default version "main" corresponds to their "main" branch.
You can specify a different version than the default "main" by using a commit SHA or a git tag of the dataset repository.
token (``str`` or :obj:`bool`, optional): Optional string or boolean to use as Bearer token for remote files on the Datasets Hub.
If True, or not specified, will get token from `"~/.huggingface"`.
use_auth_token (``str`` or :obj:`bool`, optional): Optional string or boolean to use as Bearer token for remote files on the Datasets Hub.
If True, or not specified, will get token from `"~/.huggingface"`.
<Deprecated version="2.14.0">
`use_auth_token` was deprecated in favor of `token` in version 2.14.0 and will be removed in 3.0.0.
</Deprecated>
**config_kwargs (additional keyword arguments): optional attributes for builder class which will override the attributes if supplied.
"""
if use_auth_token != "deprecated":
warnings.warn(
"'use_auth_token' was deprecated in favor of 'token' in version 2.14.0 and will be removed in 3.0.0.\n"
"You can remove this warning by passing 'token=<use_auth_token>' instead.",
FutureWarning,
)
token = use_auth_token
builder = load_dataset_builder(
path,
name=config_name,
data_files=data_files,
download_config=download_config,
download_mode=download_mode,
revision=revision,
token=token,
**config_kwargs,
)
info = builder.info
if info.splits is None:
download_config = download_config.copy() if download_config else DownloadConfig()
if token is not None:
download_config.token = token
builder._check_manual_download(
StreamingDownloadManager(base_path=builder.base_path, download_config=download_config)
)
try:
info.splits = {
split_generator.name: {"name": split_generator.name, "dataset_name": path}
for split_generator in builder._split_generators(
StreamingDownloadManager(base_path=builder.base_path, download_config=download_config)
)
}
except Exception as err:
raise SplitsNotFoundError("The split names could not be parsed from the dataset config.") from err
return info
def get_dataset_split_names(
path: str,
config_name: Optional[str] = None,
data_files: Optional[Union[str, Sequence[str], Mapping[str, Union[str, Sequence[str]]]]] = None,
download_config: Optional[DownloadConfig] = None,
download_mode: Optional[Union[DownloadMode, str]] = None,
revision: Optional[Union[str, Version]] = None,
token: Optional[Union[bool, str]] = None,
use_auth_token="deprecated",
**config_kwargs,
):
"""Get the list of available splits for a particular config and dataset.
Args:
path (`str`): path to the dataset processing script with the dataset builder. Can be either:
- a local path to processing script or the directory containing the script (if the script has the same name as the directory),
e.g. `'./dataset/squad'` or `'./dataset/squad/squad.py'`
- a dataset identifier on the Hugging Face Hub (list all available datasets and ids with [`datasets.list_datasets`])
e.g. `'squad'`, `'glue'` or `'openai/webtext'`
config_name (`str`, *optional*):
Defining the name of the dataset configuration.
data_files (`str` or `Sequence` or `Mapping`, *optional*):
Path(s) to source data file(s).
download_config ([`DownloadConfig`], *optional*):
Specific download configuration parameters.
download_mode ([`DownloadMode`] or `str`, defaults to `REUSE_DATASET_IF_EXISTS`):
Download/generate mode.
revision ([`Version`] or `str`, *optional*):
Version of the dataset script to load.
As datasets have their own git repository on the Datasets Hub, the default version "main" corresponds to their "main" branch.
You can specify a different version than the default "main" by using a commit SHA or a git tag of the dataset repository.
token (`str` or `bool`, *optional*):
Optional string or boolean to use as Bearer token for remote files on the Datasets Hub.
If `True`, or not specified, will get token from `"~/.huggingface"`.
use_auth_token (`str` or `bool`, *optional*):
Optional string or boolean to use as Bearer token for remote files on the Datasets Hub.
If `True`, or not specified, will get token from `"~/.huggingface"`.
<Deprecated version="2.14.0">
`use_auth_token` was deprecated in favor of `token` in version 2.14.0 and will be removed in 3.0.0.
</Deprecated>
**config_kwargs (additional keyword arguments):
Optional attributes for builder class which will override the attributes if supplied.
Example:
```py
>>> from datasets import get_dataset_split_names
>>> get_dataset_split_names('rotten_tomatoes')
['train', 'validation', 'test']
```
"""
if use_auth_token != "deprecated":
warnings.warn(
"'use_auth_token' was deprecated in favor of 'token' in version 2.14.0 and will be removed in 3.0.0.\n"
"You can remove this warning by passing 'token=<use_auth_token>' instead.",
FutureWarning,
)
token = use_auth_token
info = get_dataset_config_info(
path,
config_name=config_name,
data_files=data_files,
download_config=download_config,
download_mode=download_mode,
revision=revision,
token=token,
**config_kwargs,
)
return list(info.splits.keys())
| 0
|
hf_public_repos/datasets/src
|
hf_public_repos/datasets/src/datasets/distributed.py
|
from typing import TypeVar
from .arrow_dataset import Dataset, _split_by_node_map_style_dataset
from .iterable_dataset import IterableDataset, _split_by_node_iterable_dataset
DatasetType = TypeVar("DatasetType", Dataset, IterableDataset)
def split_dataset_by_node(dataset: DatasetType, rank: int, world_size: int) -> DatasetType:
"""
Split a dataset for the node at rank `rank` in a pool of nodes of size `world_size`.
For map-style datasets:
Each node is assigned a chunk of data, e.g. rank 0 is given the first chunk of the dataset.
To maximize data loading throughput, chunks are made of contiguous data on disk if possible.
For iterable datasets:
If the dataset has a number of shards that is a factor of `world_size` (i.e. if `dataset.n_shards % world_size == 0`),
then the shards are evenly assigned across the nodes, which is the most optimized.
Otherwise, each node keeps 1 example out of `world_size`, skipping the other examples.
Args:
dataset ([`Dataset`] or [`IterableDataset`]):
The dataset to split by node.
rank (`int`):
Rank of the current node.
world_size (`int`):
Total number of nodes.
Returns:
[`Dataset`] or [`IterableDataset`]: The dataset to be used on the node at rank `rank`.
"""
if isinstance(dataset, Dataset):
return _split_by_node_map_style_dataset(dataset, rank=rank, world_size=world_size)
else:
return _split_by_node_iterable_dataset(dataset, rank=rank, world_size=world_size)
| 0
|
hf_public_repos/datasets/src
|
hf_public_repos/datasets/src/datasets/__init__.py
|
# flake8: noqa
# Copyright 2020 The HuggingFace Datasets Authors and the TensorFlow Datasets Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# Lint as: python3
# pylint: enable=line-too-long
# pylint: disable=g-import-not-at-top,g-bad-import-order,wrong-import-position
__version__ = "2.15.1.dev0"
from .arrow_dataset import Dataset
from .arrow_reader import ReadInstruction
from .builder import ArrowBasedBuilder, BeamBasedBuilder, BuilderConfig, DatasetBuilder, GeneratorBasedBuilder
from .combine import concatenate_datasets, interleave_datasets
from .dataset_dict import DatasetDict, IterableDatasetDict
from .download import *
from .features import *
from .fingerprint import disable_caching, enable_caching, is_caching_enabled, set_caching_enabled
from .info import DatasetInfo, MetricInfo
from .inspect import (
get_dataset_config_info,
get_dataset_config_names,
get_dataset_infos,
get_dataset_split_names,
inspect_dataset,
inspect_metric,
list_datasets,
list_metrics,
)
from .iterable_dataset import IterableDataset
from .load import load_dataset, load_dataset_builder, load_from_disk, load_metric
from .metric import Metric
from .splits import (
NamedSplit,
NamedSplitAll,
Split,
SplitBase,
SplitDict,
SplitGenerator,
SplitInfo,
SubSplitInfo,
percent,
)
from .tasks import *
from .utils import *
from .utils import logging
# deprecated modules
from datasets import arrow_dataset as _arrow_dataset # isort:skip
from datasets import utils as _utils # isort:skip
from datasets.utils import download_manager as _deprecated_download_manager # isort:skip
_arrow_dataset.concatenate_datasets = concatenate_datasets
_utils.DownloadConfig = DownloadConfig
_utils.DownloadManager = DownloadManager
_utils.DownloadMode = DownloadMode
_deprecated_download_manager.DownloadConfig = DownloadConfig
_deprecated_download_manager.DownloadMode = DownloadMode
_deprecated_download_manager.DownloadManager = DownloadManager
del _arrow_dataset, _utils, _deprecated_download_manager
| 0
|
hf_public_repos/datasets/src
|
hf_public_repos/datasets/src/datasets/iterable_dataset.py
|
import copy
import itertools
import sys
import warnings
from collections import Counter
from copy import deepcopy
from dataclasses import dataclass
from functools import partial
from itertools import cycle, islice
from typing import Any, Callable, Dict, Iterable, Iterator, List, Optional, Tuple, Union
import numpy as np
import pyarrow as pa
from . import config
from .arrow_dataset import Dataset, DatasetInfoMixin
from .features import Features
from .features.features import FeatureType, _align_features, _check_if_features_can_be_aligned, cast_to_python_objects
from .filesystems import _reset_fsspec_lock
from .formatting import PythonFormatter, TensorFormatter, get_format_type_from_alias, get_formatter
from .info import DatasetInfo
from .splits import NamedSplit
from .table import cast_table_to_features, read_schema_from_file, table_cast
from .utils.logging import get_logger
from .utils.py_utils import Literal
from .utils.sharding import _merge_gen_kwargs, _number_of_shards_in_gen_kwargs, _shuffle_gen_kwargs, _split_gen_kwargs
logger = get_logger(__name__)
Key = Union[int, str]
def identity_func(x):
return x
def _rename_columns_fn(example: Dict, column_mapping: Dict[str, str]):
if any(col not in example for col in column_mapping):
raise ValueError(
f"Error when renaming {list(column_mapping)} to {list(column_mapping.values())}: columns {set(column_mapping) - set(example)} are not in the dataset."
)
if any(col in example for col in column_mapping.values()):
raise ValueError(
f"Error when renaming {list(column_mapping)} to {list(column_mapping.values())}: columns {set(example) - set(column_mapping.values())} are already in the dataset."
)
return {
new_column_name: example[original_column_name]
for original_column_name, new_column_name in column_mapping.items()
}
def add_column_fn(example: Dict, idx: int, name: str, column: List[Dict]):
if name in example:
raise ValueError(f"Error when adding {name}: column {name} is already in the dataset.")
return {name: column[idx]}
def _infer_features_from_batch(batch: Dict[str, list], try_features: Optional[Features] = None) -> Features:
pa_table = pa.Table.from_pydict(batch)
if try_features is not None:
try:
pa_table = table_cast(pa_table, pa.schema(try_features.type))
except (TypeError, pa.ArrowInvalid, pa.ArrowNotImplementedError):
pass
return Features.from_arrow_schema(pa_table.schema)
def _examples_to_batch(examples: List[Dict[str, Any]]) -> Dict[str, list]:
# we order the columns by order of appearance
# to do so, we use a dict as an ordered set
cols = {col: None for example in examples for col in example}
# when an example is missing a column, we set the value to None with .get()
arrays = [[example.get(col) for example in examples] for col in cols]
return dict(zip(cols, arrays))
def _batch_to_examples(batch: Dict[str, list]) -> List[Dict[str, Any]]:
"""Convert a batch (dict of examples) to examples list"""
n_examples = len(batch[next(iter(batch))])
for i in range(n_examples):
yield {col: array[i] for col, array in batch.items()}
class _HasNextIterator(Iterator):
"""Iterator with an hasnext() function. Taken from https://stackoverflow.com/questions/1966591/has-next-in-python-iterators."""
def __init__(self, it):
self.it = iter(it)
self._hasnext = None
def __iter__(self):
return self
def __next__(self):
if self._hasnext:
result = self._thenext
else:
result = next(self.it)
self._hasnext = None
return result
def hasnext(self):
if self._hasnext is None:
try:
self._thenext = next(self.it)
except StopIteration:
self._hasnext = False
else:
self._hasnext = True
return self._hasnext
def _convert_to_arrow(
iterable: Iterable[Tuple[Key, dict]],
batch_size: int,
drop_last_batch: bool = False,
) -> Iterator[Tuple[Key, pa.Table]]:
"""Convert and group examples in Arrow tables of size `batch_size`.
Args:
iterable (`Iterable[Tuple[Key, dict]]`):
An examples iterable containing tuples (example_key, example) of type (int/str, dict)
batch_size (`Optional[int]`):
Size of each sub-table to yield. If None or <= 0, yields the full table.
drop_last_batch (`bool`, defaults to `False`):
Drop the last batch if it is smaller than `batch_size`.
"""
if batch_size is None or batch_size <= 0:
yield (
"all",
pa.Table.from_pylist(cast_to_python_objects([example for _, example in iterable], only_1d_for_numpy=True)),
)
return
iterator = iter(iterable)
for key, example in iterator:
iterator_batch = islice(iterator, batch_size - 1)
key_examples_list = [(key, example)] + list(iterator_batch)
if len(key_examples_list) < batch_size and drop_last_batch:
return
keys, examples = zip(*key_examples_list)
new_key = "_".join(str(key) for key in keys)
yield new_key, pa.Table.from_pylist(cast_to_python_objects(examples, only_1d_for_numpy=True))
def _batch_arrow_tables(
iterable: Iterable[Tuple[Key, pa.Table]],
batch_size: Optional[int],
drop_last_batch: bool = False,
) -> Iterator[Tuple[Key, pa.Table]]:
"""Iterate over sub-tables of size `batch_size`.
Args:
iterable (`Iterable[Tuple[Key, pa.Table]]`):
A tables iterable containing tuples (table_key, table) of type (int/str, pa.Table)
batch_size (`Optional[int]`):
Size of each sub-table to yield. If None or <= 0, yields the full table.
drop_last_batch (`bool`, defaults to `False`):
Drop the last batch if it is smaller than `batch_size`.
"""
if batch_size is None or batch_size <= 0:
yield "all", pa.concat_tables([pa_table for _, pa_table in iterable])
return
keys_buffer = []
chunks_buffer = []
chunks_buffer_size = 0
for key, pa_table in iterable:
for chunk in pa_table.to_reader(max_chunksize=batch_size):
if len(chunk) == 0:
continue
elif chunks_buffer_size + len(chunk) < batch_size:
keys_buffer.append(key)
chunks_buffer.append(chunk)
chunks_buffer_size += len(chunk)
continue
elif chunks_buffer_size + len(chunk) == batch_size:
keys_buffer.append(key)
chunks_buffer.append(chunk)
new_key = "_".join(str(_key) for _key in keys_buffer)
yield new_key, pa.Table.from_batches(chunks_buffer)
keys_buffer = []
chunks_buffer = []
chunks_buffer_size = 0
else:
cropped_chunk_length = batch_size - chunks_buffer_size
keys_buffer.append(f"{key}[:{cropped_chunk_length}]")
chunks_buffer.append(chunk.slice(0, cropped_chunk_length))
new_key = "_".join(str(_key) for _key in keys_buffer)
yield new_key, pa.Table.from_batches(chunks_buffer)
keys_buffer = [f"{key}[{cropped_chunk_length}:]"]
chunks_buffer = [chunk.slice(cropped_chunk_length, len(chunk) - cropped_chunk_length)]
chunks_buffer_size = len(chunk) - cropped_chunk_length
if not drop_last_batch and chunks_buffer:
new_key = "_".join(str(_key) for _key in keys_buffer)
yield new_key, pa.Table.from_batches(chunks_buffer)
class _BaseExamplesIterable:
"""Base class for the examples iterable used by an IterableDataset"""
def __init__(self) -> None:
self.iter_arrow: Optional[Callable[[], Iterator[Tuple[Key, pa.Table]]]] = None
def __iter__(self) -> Iterator[Tuple[Key, dict]]:
"""An examples iterable should yield tuples (example_key, example) of type (int/str, dict)"""
raise NotImplementedError(f"{type(self)} doesn't implement __iter__ yet")
def shuffle_data_sources(self, generator: np.random.Generator) -> "_BaseExamplesIterable":
"""
Either shuffle the shards/sources of the dataset, or propagate the shuffling to the underlying iterable.
If the order of the shards must stay fixed (when using .skip or .take for example), then this method returns self.
"""
raise NotImplementedError(f"{type(self)} doesn't implement shuffle_data_sources yet")
def shard_data_sources(self, worker_id: int, num_workers: int) -> "_BaseExamplesIterable":
"""Either keep only the requested shard, or propagate the request to the underlying iterable."""
raise NotImplementedError(f"{type(self)} doesn't implement shard_data_sources yet")
def split_shard_indices_by_worker(self, worker_id: int, num_workers: int) -> List[int]:
return list(range(worker_id, self.n_shards, num_workers))
@property
def n_shards(self) -> int:
raise NotImplementedError(f"{type(self)} doesn't implement n_shards yet")
class ExamplesIterable(_BaseExamplesIterable):
def __init__(self, generate_examples_fn: Callable[..., Tuple[Key, dict]], kwargs: dict):
super().__init__()
self.generate_examples_fn = generate_examples_fn
self.kwargs = kwargs
def __iter__(self):
yield from self.generate_examples_fn(**self.kwargs)
def shuffle_data_sources(self, generator: np.random.Generator) -> "ExamplesIterable":
return ShuffledDataSourcesExamplesIterable(self.generate_examples_fn, self.kwargs, generator)
def shard_data_sources(self, worker_id: int, num_workers: int) -> "ExamplesIterable":
"""Keep only the requested shard."""
gen_kwargs_list = _split_gen_kwargs(self.kwargs, max_num_jobs=self.n_shards)
shard_indices = self.split_shard_indices_by_worker(worker_id, num_workers)
requested_gen_kwargs = _merge_gen_kwargs([gen_kwargs_list[i] for i in shard_indices])
return ExamplesIterable(self.generate_examples_fn, requested_gen_kwargs)
@property
def n_shards(self) -> int:
return _number_of_shards_in_gen_kwargs(self.kwargs)
class ShuffledDataSourcesExamplesIterable(ExamplesIterable):
def __init__(
self, generate_examples_fn: Callable[..., Tuple[Key, dict]], kwargs: dict, generator: np.random.Generator
):
super().__init__(generate_examples_fn, kwargs)
self.generator = deepcopy(generator)
def __iter__(self):
"""Shuffle the kwargs order to shuffle shards"""
rng = deepcopy(self.generator)
kwargs_with_shuffled_shards = _shuffle_gen_kwargs(rng, self.kwargs)
yield from self.generate_examples_fn(**kwargs_with_shuffled_shards)
def shard_data_sources(self, worker_id: int, num_workers: int) -> "ExamplesIterable":
"""Keep only the requested shard."""
rng = deepcopy(self.generator)
kwargs_with_shuffled_shards = _shuffle_gen_kwargs(rng, self.kwargs)
return ExamplesIterable(self.generate_examples_fn, kwargs_with_shuffled_shards).shard_data_sources(
worker_id, num_workers
)
class ArrowExamplesIterable(_BaseExamplesIterable):
def __init__(self, generate_tables_fn: Callable[..., Tuple[Key, pa.Table]], kwargs: dict):
super().__init__()
self.generate_tables_fn = generate_tables_fn
self.kwargs = kwargs
self.iter_arrow = self._iter_arrow
def __iter__(self):
formatter = PythonFormatter()
for key, pa_table in self.generate_tables_fn(**self.kwargs):
for pa_subtable in pa_table.to_reader(max_chunksize=config.ARROW_READER_BATCH_SIZE_IN_DATASET_ITER):
formatted_batch = formatter.format_batch(pa_subtable)
for example in _batch_to_examples(formatted_batch):
yield key, example
def _iter_arrow(self):
yield from self.generate_tables_fn(**self.kwargs)
def shuffle_data_sources(self, generator: np.random.Generator) -> "ArrowExamplesIterable":
return ShuffledDataSourcesArrowExamplesIterable(self.generate_tables_fn, self.kwargs, generator)
def shard_data_sources(self, worker_id: int, num_workers: int) -> "ArrowExamplesIterable":
"""Keep only the requested shard."""
gen_kwargs_list = _split_gen_kwargs(self.kwargs, max_num_jobs=self.n_shards)
shard_indices = self.split_shard_indices_by_worker(worker_id, num_workers)
requested_gen_kwargs = _merge_gen_kwargs([gen_kwargs_list[i] for i in shard_indices])
return ArrowExamplesIterable(self.generate_tables_fn, requested_gen_kwargs)
@property
def n_shards(self) -> int:
return _number_of_shards_in_gen_kwargs(self.kwargs)
class ShuffledDataSourcesArrowExamplesIterable(ArrowExamplesIterable):
def __init__(
self,
generate_tables_fn: Callable[..., Tuple[Key, pa.Table]],
kwargs: dict,
generator: np.random.Generator,
):
super().__init__(generate_tables_fn, kwargs)
self.generator = deepcopy(generator)
def __iter__(self):
"""Shuffle the kwargs order to shuffle shards"""
rng = deepcopy(self.generator)
kwargs_with_shuffled_shards = _shuffle_gen_kwargs(rng, self.kwargs)
formatter = PythonFormatter()
for key, pa_table in self.generate_tables_fn(**kwargs_with_shuffled_shards):
for pa_subtable in pa_table.to_reader(max_chunksize=config.ARROW_READER_BATCH_SIZE_IN_DATASET_ITER):
formatted_batch = formatter.format_batch(pa_subtable)
for example in _batch_to_examples(formatted_batch):
yield key, example
def _iter_arrow(self):
rng = deepcopy(self.generator)
kwargs_with_shuffled_shards = _shuffle_gen_kwargs(rng, self.kwargs)
yield from self.generate_tables_fn(**kwargs_with_shuffled_shards)
def shard_data_sources(self, worker_id: int, num_workers: int) -> "ArrowExamplesIterable":
"""Keep only the requested shard."""
rng = deepcopy(self.generator)
kwargs_with_shuffled_shards = _shuffle_gen_kwargs(rng, self.kwargs)
return ArrowExamplesIterable(self.generate_tables_fn, kwargs_with_shuffled_shards).shard_data_sources(
worker_id, num_workers
)
class SelectColumnsIterable(_BaseExamplesIterable):
def __init__(self, ex_iterable: _BaseExamplesIterable, column_names: List[str]):
super().__init__()
self.ex_iterable = ex_iterable
self.column_names = column_names
if self.ex_iterable.iter_arrow:
self.iter_arrow = self._iter_arrow
def __iter__(self):
for idx, row in self.ex_iterable:
yield idx, {c: row[c] for c in self.column_names}
def _iter_arrow(self) -> Iterator[Tuple[Key, pa.Table]]:
for idx, pa_table in self.ex_iterable.iter_arrow():
yield idx, pa_table.select(self.column_names)
def shuffle_data_sources(self, generator: np.random.Generator) -> "SelectColumnsIterable":
return SelectColumnsIterable(self.ex_iterable.shuffle_data_sources(generator), self.column_names)
def shard_data_sources(self, worker_id: int, num_workers: int) -> "SelectColumnsIterable":
return SelectColumnsIterable(self.ex_iterable.shard_data_sources(worker_id, num_workers), self.column_names)
@property
def n_shards(self) -> int:
return self.ex_iterable.n_shards
class StepExamplesIterable(_BaseExamplesIterable):
def __init__(self, ex_iterable: _BaseExamplesIterable, step: int, offset: int):
super().__init__()
self.ex_iterable = ex_iterable
self.step = step
self.offset = offset
# TODO(QL): implement iter_arrow
def __iter__(self):
ex_iterator = iter(self.ex_iterable)
while True:
batch = list(islice(ex_iterator, self.step))
if len(batch) > self.offset:
yield batch[self.offset]
else:
break
def shuffle_data_sources(self, generator: np.random.Generator) -> "StepExamplesIterable":
return StepExamplesIterable(
self.ex_iterable.shuffle_data_sources(generator), step=self.step, offset=self.offset
)
def shard_data_sources(self, worker_id: int, num_workers: int) -> "StepExamplesIterable":
return StepExamplesIterable(
self.ex_iterable.shard_data_sources(worker_id, num_workers), step=self.step, offset=self.offset
)
@property
def n_shards(self) -> int:
return self.ex_iterable.n_shards
class CyclingMultiSourcesExamplesIterable(_BaseExamplesIterable):
def __init__(
self,
ex_iterables: List[_BaseExamplesIterable],
stopping_strategy: Literal["first_exhausted", "all_exhausted"] = "first_exhausted",
):
super().__init__()
self.ex_iterables = ex_iterables
self.stopping_strategy = stopping_strategy
# if undersampling ("first_exhausted"), we stop as soon as one dataset is exhausted
# if oversampling ("all_exhausted"), we stop as soons as every dataset is exhausted, i.e as soon as every samples of every dataset has been visited at least once
self.bool_strategy_func = np.all if (stopping_strategy == "all_exhausted") else np.any
# TODO(QL): implement iter_arrow
def _get_indices_iterator(self):
# this is an infinite iterator to keep track of which iterator we want to pick examples from
return cycle(range(len(self.ex_iterables)))
def __iter__(self):
iterators = [_HasNextIterator(ex_iterable) for ex_iterable in self.ex_iterables]
indices_iterator = self._get_indices_iterator()
is_exhausted = np.full(len(self.ex_iterables), False)
for i in indices_iterator:
try: # let's pick one example from the iterator at index i
yield next(iterators[i])
# it will resume from the yield at the next call so that we can directly test if the iterable is exhausted and if we need to break out of the loop
if not iterators[i].hasnext():
is_exhausted[i] = True
if self.bool_strategy_func(is_exhausted):
# if the stopping criteria is met, break the main for loop
break
# otherwise reinitialise the iterator and yield the first example
iterators[i] = _HasNextIterator(self.ex_iterables[i])
except StopIteration:
# here it means that the i-th iterabledataset is empty, i.e we never have the occasion to yield an element of the i-th dataset.
# we still check if the stopping criteria is met and if we break out of the loop in case of an oversampling strategy
is_exhausted[i] = True
if self.bool_strategy_func(is_exhausted):
# if the stopping criteria is met, break the main for loop
break
def shuffle_data_sources(self, generator: np.random.Generator) -> "CyclingMultiSourcesExamplesIterable":
"""Shuffle each underlying examples iterable."""
ex_iterables = [ex_iterable.shuffle_data_sources(generator) for ex_iterable in self.ex_iterables]
return CyclingMultiSourcesExamplesIterable(ex_iterables, self.stopping_strategy)
@property
def n_shards(self) -> int:
return min(ex_iterable.n_shards for ex_iterable in self.ex_iterables)
def shard_data_sources(self, worker_id: int, num_workers: int) -> "CyclingMultiSourcesExamplesIterable":
"""Either keep only the requested shard, or propagate the request to the underlying iterable."""
return CyclingMultiSourcesExamplesIterable(
[iterable.shard_data_sources(worker_id, num_workers) for iterable in self.ex_iterables],
stopping_strategy=self.stopping_strategy,
)
class VerticallyConcatenatedMultiSourcesExamplesIterable(_BaseExamplesIterable):
"""
VerticallyConcatenatedMultiSourcesExamplesIterable simply chains the input iterables.
It doesn't require the examples iterables to always yield the same columns.
Instead, this is handled by the `IterableDataset` class or `TypedExamplesIterable`.
For information, `IterableDataset` merges the features of all the datasets to concatenate into one.
We use `IterableDataset._resolve_features` to obtain the features of all the datasets to concatenate.
Then for each example, `IterableDataset` and `TypedExamplesIterable` automatically fill missing columns with None.
This is done with `_apply_feature_types_on_example`.
"""
def __init__(self, ex_iterables: List[_BaseExamplesIterable]):
super().__init__()
self.ex_iterables = ex_iterables
if all(ex_iterable.iter_arrow is not None for ex_iterable in ex_iterables):
self.iter_arrow = self._iter_arrow
def __iter__(self):
for ex_iterable in self.ex_iterables:
yield from ex_iterable
def _iter_arrow(self):
for ex_iterable in self.ex_iterables:
yield from ex_iterable.iter_arrow()
def shuffle_data_sources(
self, generator: np.random.Generator
) -> "VerticallyConcatenatedMultiSourcesExamplesIterable":
"""Shuffle the list of examples iterable, as well as each underlying examples iterable."""
rng = deepcopy(generator)
ex_iterables = list(self.ex_iterables)
rng.shuffle(ex_iterables)
ex_iterables = [ex_iterable.shuffle_data_sources(generator) for ex_iterable in ex_iterables]
return VerticallyConcatenatedMultiSourcesExamplesIterable(ex_iterables)
@property
def n_shards(self) -> int:
return min(ex_iterable.n_shards for ex_iterable in self.ex_iterables)
def shard_data_sources(
self, worker_id: int, num_workers: int
) -> "VerticallyConcatenatedMultiSourcesExamplesIterable":
"""Either keep only the requested shard, or propagate the request to the underlying iterable."""
return VerticallyConcatenatedMultiSourcesExamplesIterable(
[iterable.shard_data_sources(worker_id, num_workers) for iterable in self.ex_iterables]
)
def _check_column_names(column_names: List[str]):
"""Check the column names to make sure they don't contain duplicates."""
counter = Counter(column_names)
if not all(count == 1 for count in counter.values()):
duplicated_columns = [col for col in counter if counter[col] > 1]
raise ValueError(
f"The examples iterables can't have duplicated columns but columns {duplicated_columns} are duplicated."
)
class HorizontallyConcatenatedMultiSourcesExamplesIterable(_BaseExamplesIterable):
"""
HorizontallyConcatenatedMultiSourcesExamplesIterable merges examples together for the input list of iterables.
It also checks that there are no duplicate columns (otherwise we don't know which one to keep).
This check is done once when yielding the first example.
However it doesn't fill missing columns with None.
Instead, this is handled by the `IterableDataset` class or `TypedExamplesIterable`.
For information, `IterableDataset` merges the features of all the datasets to concatenate into one.
We use `IterableDataset._resolve_features` to obtain the features of all the datasets to concatenate.
Then for each example, `IterableDataset` and `TypedExamplesIterable` automatically fill missing columns with None.
This is done with `_apply_feature_types_on_example`.
"""
def __init__(self, ex_iterables: List[_BaseExamplesIterable]):
super().__init__()
self.ex_iterables = ex_iterables
# TODO(QL): implement iter_arrow
def __iter__(self):
ex_iterators = [iter(ex_iterable) for ex_iterable in self.ex_iterables]
for i in itertools.count():
keys = []
examples = []
for ex_iterator in list(ex_iterators):
try:
key, example = next(ex_iterator)
keys.append(key)
examples.append(example)
except StopIteration:
ex_iterators.remove(ex_iterator)
if ex_iterators:
if i == 0:
_check_column_names([column_name for example in examples for column_name in example])
new_example = {}
for example in examples:
new_example.update(example)
new_key = "_".join(str(key) for key in keys)
yield new_key, new_example
else:
break
def shuffle_data_sources(
self, generator: np.random.Generator
) -> "HorizontallyConcatenatedMultiSourcesExamplesIterable":
"""Doesn't shuffle the wrapped examples iterable since it would break the alignment between them."""
return self
@property
def n_shards(self) -> int:
return 1
def shard_data_sources(
self, worker_id: int, num_workers: int
) -> "HorizontallyConcatenatedMultiSourcesExamplesIterable":
"""Either keep only the requested shard, or propagate the request to the underlying iterable."""
return HorizontallyConcatenatedMultiSourcesExamplesIterable(
[iterable.shard_data_sources(worker_id, num_workers) for iterable in self.ex_iterables]
)
class RandomlyCyclingMultiSourcesExamplesIterable(CyclingMultiSourcesExamplesIterable):
def __init__(
self,
ex_iterables: List[_BaseExamplesIterable],
generator: np.random.Generator,
probabilities: Optional[List[float]] = None,
stopping_strategy: Literal["first_exhausted", "all_exhausted"] = "first_exhausted",
):
super().__init__(ex_iterables, stopping_strategy)
self.generator = deepcopy(generator)
self.probabilities = probabilities
# TODO(QL): implement iter_arrow
@staticmethod
def _iter_random_indices(
rng: np.random.Generator,
num_sources: int,
random_batch_size=1000,
p: Optional[List[float]] = None,
) -> Iterator[int]:
"""Get an infinite iterator that randomly samples the index of the source to pick examples from."""
if p is None:
while True:
yield from (int(i) for i in rng.integers(0, num_sources, size=random_batch_size))
else:
while True:
yield from (int(i) for i in rng.choice(num_sources, size=random_batch_size, p=p))
def _get_indices_iterator(self):
rng = deepcopy(self.generator)
# this is an infinite iterator that randomly samples the index of the source to pick examples from
return self._iter_random_indices(rng, len(self.ex_iterables), p=self.probabilities)
def shuffle_data_sources(self, generator: np.random.Generator) -> "RandomlyCyclingMultiSourcesExamplesIterable":
"""Shuffle the data sources of each wrapped examples iterable."""
ex_iterables = [ex_iterable.shuffle_data_sources(generator) for ex_iterable in self.ex_iterables]
return RandomlyCyclingMultiSourcesExamplesIterable(
ex_iterables,
generator=generator,
probabilities=self.probabilities,
stopping_strategy=self.stopping_strategy,
)
def shard_data_sources(self, worker_id: int, num_workers: int) -> "RandomlyCyclingMultiSourcesExamplesIterable":
"""Either keep only the requested shard, or propagate the request to the underlying iterable."""
return RandomlyCyclingMultiSourcesExamplesIterable(
[iterable.shard_data_sources(worker_id, num_workers) for iterable in self.ex_iterables],
self.generator,
self.probabilities,
self.stopping_strategy,
)
class MappedExamplesIterable(_BaseExamplesIterable):
def __init__(
self,
ex_iterable: _BaseExamplesIterable,
function: Callable,
with_indices: bool = False,
input_columns: Optional[List[str]] = None,
batched: bool = False,
batch_size: Optional[int] = 1000,
drop_last_batch: bool = False,
remove_columns: Optional[List[str]] = None,
fn_kwargs: Optional[dict] = None,
formatting: Optional["FormattingConfig"] = None,
format_type="deprecated",
):
if format_type != "deprecated":
warning_msg = "'format_type' is deprecated and will be removed in the next major version of datasets. "
help_message = "Please use 'formatting=FormattingConfig(format_type=format_type)' instead."
warnings.warn(warning_msg + help_message, category=FutureWarning, stacklevel=2)
formatting = FormattingConfig(format_type=format_type)
super().__init__()
self.ex_iterable = ex_iterable
self.function = function
self.batched = batched
self.batch_size = batch_size
self.drop_last_batch = drop_last_batch
self.remove_columns = remove_columns
self.with_indices = with_indices
self.input_columns = input_columns
self.fn_kwargs = fn_kwargs or {}
self.formatting = formatting
if self.formatting and self.formatting.format_type == "arrow":
self.iter_arrow = self._iter_arrow
def __iter__(self):
if self.formatting and self.formatting.format_type == "arrow":
yield from ArrowExamplesIterable(self._iter_arrow, {})
else:
yield from self._iter()
def _iter(self):
iterator = iter(self.ex_iterable)
current_idx = 0
if self.formatting:
formatter = get_formatter(self.formatting.format_type)
format_dict = (
formatter.recursive_tensorize if isinstance(formatter, TensorFormatter) else cast_to_python_objects
)
else:
format_dict = None
if self.batched:
for key, example in iterator:
# If `batched`, first build the batch, if `batch_size` is None or <=0, then the batch is the whole dataset
iterator_batch = (
iterator
if self.batch_size is None or self.batch_size <= 0
else islice(iterator, self.batch_size - 1)
)
key_examples_list = [(key, example)] + list(iterator_batch)
keys, examples = zip(*key_examples_list)
if (
self.drop_last_batch
and self.batch_size is not None
and self.batch_size > 0
and len(examples) < self.batch_size
): # ignore last batch
return
batch = _examples_to_batch(examples)
batch = format_dict(batch) if format_dict else batch
# then apply the transform
inputs = batch
function_args = [inputs] if self.input_columns is None else [inputs[col] for col in self.input_columns]
if self.with_indices:
function_args.append([current_idx + i for i in range(len(key_examples_list))])
transformed_batch = dict(batch) # this will be updated with the function output
transformed_batch.update(self.function(*function_args, **self.fn_kwargs))
# then remove the unwanted columns
if self.remove_columns:
for c in self.remove_columns:
del transformed_batch[c]
if transformed_batch:
first_col = next(iter(transformed_batch))
bad_cols = [
col
for col in transformed_batch
if len(transformed_batch[col]) != len(transformed_batch[first_col])
]
if bad_cols:
raise ValueError(
f"Column lengths mismatch: columns {bad_cols} have length {[len(transformed_batch[col]) for col in bad_cols]} while {first_col} has length {len(transformed_batch[first_col])}."
)
# the new key is the concatenation of the examples keys from the batch
new_key = "_".join(str(key) for key in keys)
# yield one example at a time from the transformed batch
for example in _batch_to_examples(transformed_batch):
yield new_key, example
current_idx += 1
else:
for key, example in iterator:
# If not batched, we can apply the transform and yield the example directly
# first copy the example, since we might drop some keys
example = dict(example)
example = format_dict(example) if format_dict else example
# then apply the transform
inputs = example
function_args = [inputs] if self.input_columns is None else [inputs[col] for col in self.input_columns]
if self.with_indices:
function_args.append(current_idx)
transformed_example = dict(example) # this will be updated with the function output
transformed_example.update(self.function(*function_args, **self.fn_kwargs))
# then we remove the unwanted columns
if self.remove_columns:
for c in self.remove_columns:
del transformed_example[c]
yield key, transformed_example
current_idx += 1
def _iter_arrow(self) -> Iterator[Tuple[Key, pa.Table]]:
if self.ex_iterable.iter_arrow:
iterator = _batch_arrow_tables(
self.ex_iterable.iter_arrow(),
batch_size=self.batch_size if self.batched else 1,
drop_last_batch=self.drop_last_batch,
)
else:
iterator = _convert_to_arrow(
self.ex_iterable,
batch_size=self.batch_size if self.batched else 1,
drop_last_batch=self.drop_last_batch,
)
current_idx = 0
for key, pa_table in iterator:
# first build the batch
function_args = [pa_table] if self.input_columns is None else [pa_table[col] for col in self.input_columns]
if self.with_indices:
if self.batched:
function_args.append([current_idx + i for i in range(len(pa_table))])
else:
function_args.append(current_idx)
# then apply the transform
output_table = self.function(*function_args, **self.fn_kwargs)
if not isinstance(output_table, pa.Table):
raise TypeError(
f"Provided `function` which is applied to pyarrow tables returns a variable of type {type(output_table)}. Make sure provided `function` returns a a pyarrow table to update the dataset."
)
# we don't need to merge results for consistency with Dataset.map which merges iif both input and output are dicts
# then remove the unwanted columns
if self.remove_columns:
for column in self.remove_columns:
if column in output_table.column_names:
output_table = output_table.remove_column(output_table.column_names.index(column))
# return output
yield key, output_table
current_idx += len(pa_table)
def shuffle_data_sources(self, generator: np.random.Generator) -> "MappedExamplesIterable":
"""Shuffle the wrapped examples iterable."""
return MappedExamplesIterable(
self.ex_iterable.shuffle_data_sources(generator),
function=self.function,
with_indices=self.with_indices,
input_columns=self.input_columns,
batched=self.batched,
batch_size=self.batch_size,
remove_columns=self.remove_columns,
fn_kwargs=self.fn_kwargs,
)
def shard_data_sources(self, worker_id: int, num_workers: int) -> "MappedExamplesIterable":
"""Keep only the requested shard."""
return MappedExamplesIterable(
self.ex_iterable.shard_data_sources(worker_id, num_workers),
function=self.function,
with_indices=self.with_indices,
input_columns=self.input_columns,
batched=self.batched,
batch_size=self.batch_size,
remove_columns=self.remove_columns,
fn_kwargs=self.fn_kwargs,
)
@property
def n_shards(self) -> int:
return self.ex_iterable.n_shards
class FilteredExamplesIterable(_BaseExamplesIterable):
def __init__(
self,
ex_iterable: _BaseExamplesIterable,
function: Callable,
with_indices: bool = False,
input_columns: Optional[List[str]] = None,
batched: bool = False,
batch_size: Optional[int] = 1000,
fn_kwargs: Optional[dict] = None,
formatting: Optional["FormattingConfig"] = None,
format_type="deprecated",
):
if format_type != "deprecated":
warning_msg = "'format_type' is deprecated and will be removed in the next major version of datasets. "
help_message = "Please use 'formatting=FormattingConfig(format_type=format_type)' instead."
warnings.warn(warning_msg + help_message, category=FutureWarning, stacklevel=2)
formatting = FormattingConfig(format_type=format_type)
super().__init__()
self.ex_iterable = ex_iterable
self.function = function
self.batched = batched
self.batch_size = batch_size
self.with_indices = with_indices
self.input_columns = input_columns
self.fn_kwargs = fn_kwargs or {}
self.formatting = formatting
if self.formatting and self.formatting.format_type == "arrow":
self.iter_arrow = self._iter_arrow
def __iter__(self):
if self.formatting and self.formatting.format_type == "arrow":
yield from ArrowExamplesIterable(self._iter_arrow, {})
else:
yield from self._iter()
def _iter(self):
if self.formatting:
formatter = get_formatter(self.formatting.format_type)
format_dict = (
formatter.recursive_tensorize if isinstance(formatter, TensorFormatter) else cast_to_python_objects
)
else:
format_dict = None
iterator = iter(self.ex_iterable)
current_idx = 0
if self.batched:
for key, example in iterator:
# If `batched`, first build the batch, if `batch_size` is None or <=0, then the batch is the whole dataset
iterator_batch = (
iterator
if self.batch_size is None or self.batch_size <= 0
else islice(iterator, self.batch_size - 1)
)
key_examples_list = [(key, example)] + list(iterator_batch)
keys, examples = zip(*key_examples_list)
batch = _examples_to_batch(examples)
batch = format_dict(batch) if format_dict else batch
# then compute the mask for the batch
inputs = batch
function_args = [inputs] if self.input_columns is None else [inputs[col] for col in self.input_columns]
if self.with_indices:
function_args.append([current_idx + i for i in range(len(key_examples_list))])
mask = self.function(*function_args, **self.fn_kwargs)
# yield one example at a time from the batch
for key_example, to_keep in zip(key_examples_list, mask):
if to_keep:
yield key_example
current_idx += 1
else:
for key, example in iterator:
# If not batched, we can apply the filtering function direcly
example = dict(example)
inputs = format_dict(example) if format_dict else example
function_args = [inputs] if self.input_columns is None else [inputs[col] for col in self.input_columns]
if self.with_indices:
function_args.append(current_idx)
to_keep = self.function(*function_args, **self.fn_kwargs)
if to_keep:
yield key, example
current_idx += 1
def _iter_arrow(self):
if self.ex_iterable.iter_arrow:
iterator = _batch_arrow_tables(
self.ex_iterable.iter_arrow(), batch_size=self.batch_size if self.batched else 1
)
else:
iterator = _convert_to_arrow(self.ex_iterable, batch_size=self.batch_size if self.batched else 1)
current_idx = 0
for key, pa_table in iterator:
# first build the batch
function_args = [pa_table] if self.input_columns is None else [pa_table[col] for col in self.input_columns]
if self.with_indices:
if self.batched:
function_args.append([current_idx + i for i in range(len(pa_table))])
else:
function_args.append(current_idx)
# then apply the transform
mask = self.function(*function_args, **self.fn_kwargs)
# yield the filtered table
if self.batched:
yield key, pa_table.filter(mask)
elif mask.as_py() if isinstance(mask, pa.BooleanScalar) else mask:
yield key, pa_table
current_idx += len(pa_table)
def shuffle_data_sources(self, seed: Optional[int]) -> "FilteredExamplesIterable":
"""Shuffle the wrapped examples iterable."""
return FilteredExamplesIterable(
self.ex_iterable.shuffle_data_sources(seed),
function=self.function,
with_indices=self.with_indices,
input_columns=self.input_columns,
batched=self.batched,
batch_size=self.batch_size,
)
def shard_data_sources(self, worker_id: int, num_workers: int) -> "FilteredExamplesIterable":
"""Keep only the requested shard."""
return FilteredExamplesIterable(
self.ex_iterable.shard_data_sources(worker_id, num_workers),
function=self.function,
with_indices=self.with_indices,
input_columns=self.input_columns,
batched=self.batched,
batch_size=self.batch_size,
)
@property
def n_shards(self) -> int:
return self.ex_iterable.n_shards
class BufferShuffledExamplesIterable(_BaseExamplesIterable):
def __init__(self, ex_iterable: _BaseExamplesIterable, buffer_size: int, generator: np.random.Generator):
super().__init__()
self.ex_iterable = ex_iterable
self.buffer_size = buffer_size
self.generator = generator
# TODO(QL): implement iter_arrow
@staticmethod
def _iter_random_indices(rng: np.random.Generator, buffer_size: int, random_batch_size=1000) -> Iterator[int]:
while True:
yield from (int(i) for i in rng.integers(0, buffer_size, size=random_batch_size))
def __iter__(self):
buffer_size = self.buffer_size
rng = deepcopy(self.generator)
indices_iterator = self._iter_random_indices(rng, buffer_size)
# this is the shuffle buffer that we keep in memory
mem_buffer = []
for x in self.ex_iterable:
if len(mem_buffer) == buffer_size: # if the buffer is full, pick and example from it
i = next(indices_iterator)
yield mem_buffer[i]
mem_buffer[i] = x # replace the picked example by a new one
else: # otherwise, keep filling the buffer
mem_buffer.append(x)
# when we run out of examples, we shuffle the remaining examples in the buffer and yield them
rng.shuffle(mem_buffer)
yield from mem_buffer
def shuffle_data_sources(self, generator: np.random.Generator) -> "BufferShuffledExamplesIterable":
"""Shuffle the wrapped examples iterable as well as the shuffling buffer."""
return BufferShuffledExamplesIterable(
self.ex_iterable.shuffle_data_sources(generator), buffer_size=self.buffer_size, generator=generator
)
def shard_data_sources(self, worker_id: int, num_workers: int) -> "BufferShuffledExamplesIterable":
"""Keep only the requested shard."""
return BufferShuffledExamplesIterable(
self.ex_iterable.shard_data_sources(worker_id, num_workers),
buffer_size=self.buffer_size,
generator=self.generator,
)
@property
def n_shards(self) -> int:
return self.ex_iterable.n_shards
class SkipExamplesIterable(_BaseExamplesIterable):
def __init__(self, ex_iterable: _BaseExamplesIterable, n: int):
super().__init__()
self.ex_iterable = ex_iterable
self.n = n
# TODO(QL): implement iter_arrow
def __iter__(self):
yield from islice(self.ex_iterable, self.n, None)
def shuffle_data_sources(self, generator: np.random.Generator) -> "SkipExamplesIterable":
"""Doesn't shuffle the wrapped examples iterable since it would skip examples from other shards instead."""
return self
@property
def n_shards(self) -> int:
return self.ex_iterable.n_shards
class TakeExamplesIterable(_BaseExamplesIterable):
def __init__(self, ex_iterable: _BaseExamplesIterable, n: int):
super().__init__()
self.ex_iterable = ex_iterable
self.n = n
# TODO(QL): implement iter_arrow
def __iter__(self):
yield from islice(self.ex_iterable, self.n)
def shuffle_data_sources(self, generator: np.random.Generator) -> "TakeExamplesIterable":
"""Doesn't shuffle the wrapped examples iterable since it would take examples from other shards instead."""
return self
@staticmethod
def split_number(num, n):
quotient = num // n
remainder = num % n
result = [quotient] * n
for i in range(remainder):
result[i] += 1
return result
def shard_data_sources(self, worker_id: int, num_workers: int) -> "TakeExamplesIterable":
"""Keep only the requested shard."""
return TakeExamplesIterable(
self.ex_iterable.shard_data_sources(worker_id, num_workers),
n=self.split_number(self.n, num_workers)[worker_id],
)
@property
def n_shards(self) -> int:
return self.ex_iterable.n_shards
def _apply_feature_types_on_example(
example: dict, features: Features, token_per_repo_id: Dict[str, Union[str, bool, None]]
) -> dict:
example = dict(example)
# add missing columns
for column_name in features:
if column_name not in example:
example[column_name] = None
# we encode the example for ClassLabel feature types for example
encoded_example = features.encode_example(example)
# Decode example for Audio feature, e.g.
decoded_example = features.decode_example(encoded_example, token_per_repo_id=token_per_repo_id)
return decoded_example
def _apply_feature_types_on_batch(
batch: dict, features: Features, token_per_repo_id: Dict[str, Union[str, bool, None]]
) -> dict:
batch = dict(batch)
# add missing columns
n_examples = len(batch[next(iter(batch))])
for column_name in features:
if column_name not in batch:
batch[column_name] = [None] * n_examples
# we encode the batch for ClassLabel feature types for example
encoded_batch = features.encode_batch(batch)
# Decode batch for Audio feature, e.g.
decoded_batch = features.decode_batch(encoded_batch, token_per_repo_id=token_per_repo_id)
return decoded_batch
class TypedExamplesIterable(_BaseExamplesIterable):
def __init__(
self,
ex_iterable: _BaseExamplesIterable,
features: Features,
token_per_repo_id: Dict[str, Union[str, bool, None]],
):
super().__init__()
self.ex_iterable = ex_iterable
self.features = features
self.token_per_repo_id = token_per_repo_id
if self.ex_iterable.iter_arrow is not None:
self.iter_arrow = self._iter_arrow
def __iter__(self):
# Then for each example, `TypedExamplesIterable` automatically fills missing columns with None.
# This is done with `_apply_feature_types_on_example`.
for key, example in self.ex_iterable:
yield (
key,
_apply_feature_types_on_example(example, self.features, token_per_repo_id=self.token_per_repo_id),
)
def _iter_arrow(self) -> Iterator[Tuple[Key, pa.Table]]:
schema = self.features.arrow_schema
for key, pa_table in self.ex_iterable.iter_arrow():
columns = set(pa_table.column_names)
# add missing columns
for column_name in self.features:
if column_name not in columns:
col = pa.NullArray.from_buffers(pa.null(), len(pa_table), [None])
pa_table = pa_table.append_column(column_name, col)
if pa_table.schema != schema:
pa_table = cast_table_to_features(pa_table, self.features)
yield key, pa_table
def shuffle_data_sources(self, generator: np.random.Generator) -> "TypedExamplesIterable":
"""Shuffle the wrapped examples iterable."""
return TypedExamplesIterable(
self.ex_iterable.shuffle_data_sources(generator),
features=self.features,
token_per_repo_id=self.token_per_repo_id,
)
def shard_data_sources(self, worker_id: int, num_workers: int) -> "TypedExamplesIterable":
"""Keep only the requested shard."""
return TypedExamplesIterable(
self.ex_iterable.shard_data_sources(worker_id, num_workers),
features=self.features,
token_per_repo_id=self.token_per_repo_id,
)
@property
def n_shards(self) -> int:
return self.ex_iterable.n_shards
@dataclass
class FormattingConfig:
format_type: Optional[str]
def __post_init__(self):
if self.format_type == "pandas":
raise NotImplementedError(
"The 'pandas' formatting is not implemented for iterable datasets. You can use 'numpy' or 'arrow' instead."
)
@dataclass
class ShufflingConfig:
generator: np.random.Generator
_original_seed: Optional[int] = None
@dataclass
class DistributedConfig:
rank: int
world_size: int
def _maybe_add_torch_iterable_dataset_parent_class(cls):
"""Add torch.utils.data.IterableDataset as a parent class if 'torch' is available"""
if config.TORCH_AVAILABLE:
import torch.utils.data
if torch.utils.data.IterableDataset not in cls.__bases__:
cls.__bases__ += (torch.utils.data.IterableDataset,)
class IterableDataset(DatasetInfoMixin):
"""A Dataset backed by an iterable."""
def __init__(
self,
ex_iterable: _BaseExamplesIterable,
info: Optional[DatasetInfo] = None,
split: Optional[NamedSplit] = None,
formatting: Optional[FormattingConfig] = None,
shuffling: Optional[ShufflingConfig] = None,
distributed: Optional[DistributedConfig] = None,
token_per_repo_id: Optional[Dict[str, Union[str, bool, None]]] = None,
format_type="deprecated",
):
if distributed and distributed.world_size > 1 and shuffling and shuffling._original_seed is None:
raise RuntimeError(
"The dataset doesn't have a fixed random seed across nodes to shuffle and split the list of dataset shards by node. "
"Please pass e.g. `seed=42` in `.shuffle()` to make all the nodes use the same seed. "
)
if format_type != "deprecated":
warning_msg = "'format_type' is deprecated and will be removed in the next major version of datasets. "
help_message = "Please use 'formatting=FormattingConfig(format_type=format_type)' instead."
warnings.warn(warning_msg + help_message, category=FutureWarning, stacklevel=2)
formatting = FormattingConfig(format_type=format_type)
info = info.copy() if info is not None else DatasetInfo()
DatasetInfoMixin.__init__(self, info=info, split=split)
self._ex_iterable = ex_iterable
self._formatting = formatting
self._shuffling = shuffling
self._distributed = distributed
self._epoch = 0
self._token_per_repo_id: Dict[str, Union[str, bool, None]] = token_per_repo_id or {}
_maybe_add_torch_iterable_dataset_parent_class(self.__class__)
def __getstate__(self):
return self.__dict__
def __setstate__(self, d):
self.__dict__ = d
# Re-add torch iterable dataset as a parent class, since dynamically added parent classes are not kept when pickling
_maybe_add_torch_iterable_dataset_parent_class(self.__class__)
def _head(self, n=5):
return _examples_to_batch(list(self.take(n)))
def _effective_generator(self):
if self._shuffling and self._epoch == 0:
return self._shuffling.generator
elif self._shuffling:
# Create effective seed using self._epoch (we subtract in order to avoir overflow in long_scalars)
effective_seed = deepcopy(self._shuffling.generator).integers(0, 1 << 63) - self._epoch
effective_seed = (1 << 63) + effective_seed if effective_seed < 0 else effective_seed
return np.random.default_rng(effective_seed)
else:
raise ValueError("This dataset is not shuffled")
@property
def n_shards(self) -> int:
if self._distributed and self._ex_iterable.n_shards % self._distributed.world_size == 0:
return self._ex_iterable.n_shards // self._distributed.world_size
return self._ex_iterable.n_shards
def _iter_pytorch(self):
ex_iterable = self._prepare_ex_iterable_for_iteration()
# fix for fsspec when using multiprocess
_reset_fsspec_lock()
# check if there aren't too many workers
import torch.utils.data
worker_info = torch.utils.data.get_worker_info()
if self._is_main_process() and ex_iterable.n_shards < worker_info.num_workers:
logger.warning(
f"Too many dataloader workers: {worker_info.num_workers} (max is dataset.n_shards={ex_iterable.n_shards}). "
f"Stopping {worker_info.num_workers - ex_iterable.n_shards} dataloader workers."
)
logger.info(
f"To parallelize data loading, we give each process some shards (or data sources) to process. "
f"Therefore it's unnecessary to have a number of workers greater than dataset.n_shards={ex_iterable.n_shards}. "
f"To enable more parallelism, please split the dataset in more files than {ex_iterable.n_shards}."
)
# split workload
_log_prefix = f"node#{self._distributed.rank} " if self._distributed else ""
shards_indices = self._ex_iterable.split_shard_indices_by_worker(worker_info.id, worker_info.num_workers)
if shards_indices:
logger.debug(
f"{_log_prefix}dataloader worker#{worker_info.id}, ': Starting to iterate over {len(shards_indices)}/{ex_iterable.n_shards} shards."
)
ex_iterable = ex_iterable.shard_data_sources(worker_id=worker_info.id, num_workers=worker_info.num_workers)
if self._formatting:
formatter = get_formatter(self._formatting.format_type, features=self.features)
format_dict = (
formatter.recursive_tensorize if isinstance(formatter, TensorFormatter) else cast_to_python_objects
)
else:
format_dict = None
if self._formatting and (ex_iterable.iter_arrow or self._formatting == "arrow"):
if ex_iterable.iter_arrow:
iterator = _batch_arrow_tables(ex_iterable.iter_arrow(), batch_size=1)
else:
iterator = _convert_to_arrow(ex_iterable, batch_size=1)
for key, pa_table in iterator:
yield formatter.format_row(pa_table)
return
else:
for key, example in ex_iterable:
if self.features:
# `IterableDataset` automatically fills missing columns with None.
# This is done with `_apply_feature_types_on_example`.
example = _apply_feature_types_on_example(
example, self.features, token_per_repo_id=self._token_per_repo_id
)
yield format_dict(example) if format_dict else example
logger.debug(
f"{_log_prefix}dataloader worker#{worker_info.id}, ': Finished iterating over {len(shards_indices)}/{ex_iterable.n_shards} shards."
)
else:
logger.debug(
f"{_log_prefix}dataloader worker#{worker_info.id}, ': Stopping... Number of dataset shards < num_workers ({ex_iterable.n_shards}<{worker_info.num_workers})."
)
def _is_main_process(self):
if self._distributed and self._distributed.rank > 0:
return False
if "torch" in sys.modules:
import torch.utils.data
worker_info = torch.utils.data.get_worker_info()
if worker_info is not None and worker_info.id > 0:
return False
return True
def _prepare_ex_iterable_for_iteration(self) -> _BaseExamplesIterable:
if self._shuffling:
ex_iterable = self._ex_iterable.shuffle_data_sources(self._effective_generator())
else:
ex_iterable = self._ex_iterable
if self._distributed:
rank = self._distributed.rank
world_size = self._distributed.world_size
if ex_iterable.n_shards % world_size == 0:
if self._is_main_process():
n_shards_per_node = ex_iterable.n_shards // world_size
plural = "s" if n_shards_per_node > 1 else ""
logger.info(
f"Assigning {n_shards_per_node} shard{plural} (or data source{plural}) of the dataset to each node."
)
ex_iterable = ex_iterable.shard_data_sources(rank, world_size)
else:
if self._is_main_process():
logger.info(
f"Assigning 1 out of {world_size} examples of the dataset to each node. The others are skipped during the iteration."
)
logger.info(
f"It is more optimized to distribute the dataset shards (or data sources) across nodes. "
f"You can do that by using a dataset with number of shards that is a factor of world_size={world_size}. "
f"The current dataset has {ex_iterable.n_shards} which is not a factor of {world_size}"
)
ex_iterable = StepExamplesIterable(ex_iterable, step=world_size, offset=rank)
return ex_iterable
def __iter__(self):
if "torch" in sys.modules:
import torch.utils.data
worker_info = torch.utils.data.get_worker_info()
if isinstance(self, torch.utils.data.IterableDataset) and worker_info is not None:
# We're a torch.utils.data.IterableDataset in a PyTorch worker process
yield from self._iter_pytorch()
return
ex_iterable = self._prepare_ex_iterable_for_iteration()
if self._formatting:
formatter = get_formatter(self._formatting.format_type, features=self.features)
format_dict = (
formatter.recursive_tensorize if isinstance(formatter, TensorFormatter) else cast_to_python_objects
)
else:
format_dict = None
if self._formatting and (ex_iterable.iter_arrow or self._formatting.format_type == "arrow"):
if ex_iterable.iter_arrow:
iterator = _batch_arrow_tables(ex_iterable.iter_arrow(), batch_size=1)
else:
iterator = _convert_to_arrow(ex_iterable, batch_size=1)
for key, pa_table in iterator:
yield formatter.format_row(pa_table)
return
for key, example in ex_iterable:
if self.features:
# `IterableDataset` automatically fills missing columns with None.
# This is done with `_apply_feature_types_on_example`.
example = _apply_feature_types_on_example(
example, self.features, token_per_repo_id=self._token_per_repo_id
)
yield format_dict(example) if format_dict else example
def iter(self, batch_size: int, drop_last_batch: bool = False):
"""Iterate through the batches of size `batch_size`.
Args:
batch_size (:obj:`int`): size of each batch to yield.
drop_last_batch (:obj:`bool`, default `False`): Whether a last batch smaller than the batch_size should be
dropped
"""
if self._formatting:
formatter = get_formatter(self._formatting.format_type, features=self.features)
format_dict = (
formatter.recursive_tensorize if isinstance(formatter, TensorFormatter) else cast_to_python_objects
)
else:
format_dict = None
ex_iterable = self._prepare_ex_iterable_for_iteration()
if self._formatting and (ex_iterable.iter_arrow or self._formatting == "arrow"):
if ex_iterable.iter_arrow:
iterator = _batch_arrow_tables(
ex_iterable.iter_arrow(), batch_size=batch_size, drop_last_batch=drop_last_batch
)
else:
iterator = _convert_to_arrow(ex_iterable, batch_size=batch_size, drop_last_batch=drop_last_batch)
for key, pa_table in iterator:
yield formatter.format_batch(pa_table)
return
iterator = iter(ex_iterable)
for key, example in iterator:
# If batched, first build the batch
examples = [example] + [example for key, example in islice(iterator, batch_size - 1)]
if drop_last_batch and len(examples) < batch_size: # ignore last batch
return
batch = _examples_to_batch(examples)
if self.features:
# `IterableDataset` automatically fills missing columns with None.
# This is done with `_apply_feature_types_on_batch`.
batch = _apply_feature_types_on_batch(batch, self.features, token_per_repo_id=self._token_per_repo_id)
yield format_dict(batch) if format_dict else batch
@staticmethod
def from_generator(
generator: Callable,
features: Optional[Features] = None,
gen_kwargs: Optional[dict] = None,
) -> "IterableDataset":
"""Create an Iterable Dataset from a generator.
Args:
generator (`Callable`):
A generator function that `yields` examples.
features (`Features`, *optional*):
Dataset features.
gen_kwargs(`dict`, *optional*):
Keyword arguments to be passed to the `generator` callable.
You can define a sharded iterable dataset by passing the list of shards in `gen_kwargs`.
This can be used to improve shuffling and when iterating over the dataset with multiple workers.
Returns:
`IterableDataset`
Example:
```py
>>> def gen():
... yield {"text": "Good", "label": 0}
... yield {"text": "Bad", "label": 1}
...
>>> ds = IterableDataset.from_generator(gen)
```
```py
>>> def gen(shards):
... for shard in shards:
... with open(shard) as f:
... for line in f:
... yield {"line": line}
...
>>> shards = [f"data{i}.txt" for i in range(32)]
>>> ds = IterableDataset.from_generator(gen, gen_kwargs={"shards": shards})
>>> ds = ds.shuffle(seed=42, buffer_size=10_000) # shuffles the shards order + uses a shuffle buffer
>>> from torch.utils.data import DataLoader
>>> dataloader = DataLoader(ds.with_format("torch"), num_workers=4) # give each worker a subset of 32/4=8 shards
```
"""
from .io.generator import GeneratorDatasetInputStream
return GeneratorDatasetInputStream(
generator=generator,
features=features,
gen_kwargs=gen_kwargs,
streaming=True,
).read()
@staticmethod
def from_spark(
df: "pyspark.sql.DataFrame",
split: Optional[NamedSplit] = None,
features: Optional[Features] = None,
**kwargs,
) -> "IterableDataset":
"""Create an IterableDataset from Spark DataFrame. The dataset is streamed to the driver in batches.
Args:
df (`pyspark.sql.DataFrame`):
The DataFrame containing the desired data.
split (`NamedSplit`, *optional*):
Split name to be assigned to the dataset.
features (`Features`, *optional*):
Dataset features.
Returns:
[`IterableDataset`]
Example:
```py
>>> df = spark.createDataFrame(
>>> data=[[1, "Elia"], [2, "Teo"], [3, "Fang"]],
>>> columns=["id", "name"],
>>> )
>>> ds = IterableDataset.from_spark(df)
```
"""
from .io.spark import SparkDatasetReader
if sys.platform == "win32":
raise EnvironmentError("IterableDataset.from_spark is not currently supported on Windows")
return SparkDatasetReader(
df,
split=split,
features=features,
streaming=True,
**kwargs,
).read()
@staticmethod
def from_file(filename: str) -> "IterableDataset":
"""Instantiate a IterableDataset from Arrow table at filename.
Args:
filename (`str`):
File name of the dataset.
Returns:
[`IterableDataset`]
"""
pa_table_schema = read_schema_from_file(filename)
inferred_features = Features.from_arrow_schema(pa_table_schema)
ex_iterable = ArrowExamplesIterable(Dataset._generate_tables_from_cache_file, kwargs={"filename": filename})
return IterableDataset(ex_iterable=ex_iterable, info=DatasetInfo(features=inferred_features))
def with_format(
self,
type: Optional[str] = None,
) -> "IterableDataset":
"""
Return a dataset with the specified format.
Supported formats: "arrow", or None for regular python objects.
The other formats are currently not implemented.
Args:
type (`str`, optional, default None): if set to "torch", the returned dataset
will be a subclass of torch.utils.data.IterableDataset to be used in a DataLoader
"""
type = get_format_type_from_alias(type)
# TODO(QL): add format_kwargs
# TODO(QL): add format_columns and return_all_columns
# TODO(QL): add pandas format
return IterableDataset(
ex_iterable=self._ex_iterable,
info=self._info.copy(),
split=self._split,
formatting=FormattingConfig(format_type=type),
shuffling=copy.deepcopy(self._shuffling),
distributed=copy.deepcopy(self._distributed),
token_per_repo_id=self._token_per_repo_id,
)
def map(
self,
function: Optional[Callable] = None,
with_indices: bool = False,
input_columns: Optional[Union[str, List[str]]] = None,
batched: bool = False,
batch_size: Optional[int] = 1000,
drop_last_batch: bool = False,
remove_columns: Optional[Union[str, List[str]]] = None,
features: Optional[Features] = None,
fn_kwargs: Optional[dict] = None,
) -> "IterableDataset":
"""
Apply a function to all the examples in the iterable dataset (individually or in batches) and update them.
If your function returns a column that already exists, then it overwrites it.
The function is applied on-the-fly on the examples when iterating over the dataset.
You can specify whether the function should be batched or not with the `batched` parameter:
- If batched is `False`, then the function takes 1 example in and should return 1 example.
An example is a dictionary, e.g. `{"text": "Hello there !"}`.
- If batched is `True` and `batch_size` is 1, then the function takes a batch of 1 example as input and can return a batch with 1 or more examples.
A batch is a dictionary, e.g. a batch of 1 example is {"text": ["Hello there !"]}.
- If batched is `True` and `batch_size` is `n` > 1, then the function takes a batch of `n` examples as input and can return a batch with `n` examples, or with an arbitrary number of examples.
Note that the last batch may have less than `n` examples.
A batch is a dictionary, e.g. a batch of `n` examples is `{"text": ["Hello there !"] * n}`.
Args:
function (`Callable`, *optional*, defaults to `None`):
Function applied on-the-fly on the examples when you iterate on the dataset.
It must have one of the following signatures:
- `function(example: Dict[str, Any]) -> Dict[str, Any]` if `batched=False` and `with_indices=False`
- `function(example: Dict[str, Any], idx: int) -> Dict[str, Any]` if `batched=False` and `with_indices=True`
- `function(batch: Dict[str, List]) -> Dict[str, List]` if `batched=True` and `with_indices=False`
- `function(batch: Dict[str, List], indices: List[int]) -> Dict[str, List]` if `batched=True` and `with_indices=True`
For advanced usage, the function can also return a `pyarrow.Table`.
Moreover if your function returns nothing (`None`), then `map` will run your function and return the dataset unchanged.
If no function is provided, default to identity function: `lambda x: x`.
with_indices (`bool`, defaults to `False`):
Provide example indices to `function`. Note that in this case the signature of `function` should be `def function(example, idx[, rank]): ...`.
input_columns (`Optional[Union[str, List[str]]]`, defaults to `None`):
The columns to be passed into `function`
as positional arguments. If `None`, a dict mapping to all formatted columns is passed as one argument.
batched (`bool`, defaults to `False`):
Provide batch of examples to `function`.
batch_size (`int`, *optional*, defaults to `1000`):
Number of examples per batch provided to `function` if `batched=True`.
`batch_size <= 0` or `batch_size == None` then provide the full dataset as a single batch to `function`.
drop_last_batch (`bool`, defaults to `False`):
Whether a last batch smaller than the batch_size should be
dropped instead of being processed by the function.
remove_columns (`[List[str]]`, *optional*, defaults to `None`):
Remove a selection of columns while doing the mapping.
Columns will be removed before updating the examples with the output of `function`, i.e. if `function` is adding
columns with names in `remove_columns`, these columns will be kept.
features (`[Features]`, *optional*, defaults to `None`):
Feature types of the resulting dataset.
fn_kwargs (`Dict`, *optional*, default `None`):
Keyword arguments to be passed to `function`.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", split="train", streaming=True)
>>> def add_prefix(example):
... example["text"] = "Review: " + example["text"]
... return example
>>> ds = ds.map(add_prefix)
>>> list(ds.take(3))
[{'label': 1,
'text': 'Review: the rock is destined to be the 21st century\'s new " conan " and that he\'s going to make a splash even greater than arnold schwarzenegger , jean-claud van damme or steven segal .'},
{'label': 1,
'text': 'Review: the gorgeously elaborate continuation of " the lord of the rings " trilogy is so huge that a column of words cannot adequately describe co-writer/director peter jackson\'s expanded vision of j . r . r . tolkien\'s middle-earth .'},
{'label': 1, 'text': 'Review: effective but too-tepid biopic'}]
```
"""
if isinstance(input_columns, str):
input_columns = [input_columns]
if isinstance(remove_columns, str):
remove_columns = [remove_columns]
if function is None:
function = identity_func
if fn_kwargs is None:
fn_kwargs = {}
ex_iterable = MappedExamplesIterable(
TypedExamplesIterable(self._ex_iterable, self._info.features, token_per_repo_id=self._token_per_repo_id)
if self._info.features is not None
else self._ex_iterable,
function=function,
with_indices=with_indices,
input_columns=input_columns,
batched=batched,
batch_size=batch_size,
drop_last_batch=drop_last_batch,
remove_columns=remove_columns,
fn_kwargs=fn_kwargs,
formatting=self._formatting,
)
info = self.info.copy()
info.features = features
return IterableDataset(
ex_iterable=ex_iterable,
info=info,
split=self._split,
formatting=self._formatting,
shuffling=copy.deepcopy(self._shuffling),
distributed=copy.deepcopy(self._distributed),
token_per_repo_id=self._token_per_repo_id,
)
def filter(
self,
function: Optional[Callable] = None,
with_indices=False,
input_columns: Optional[Union[str, List[str]]] = None,
batched: bool = False,
batch_size: Optional[int] = 1000,
fn_kwargs: Optional[dict] = None,
) -> "IterableDataset":
"""Apply a filter function to all the elements so that the dataset only includes examples according to the filter function.
The filtering is done on-the-fly when iterating over the dataset.
Args:
function (`Callable`):
Callable with one of the following signatures:
- `function(example: Dict[str, Any]) -> bool` if `with_indices=False, batched=False`
- `function(example: Dict[str, Any], indices: int) -> bool` if `with_indices=True, batched=False`
- `function(example: Dict[str, List]) -> List[bool]` if `with_indices=False, batched=True`
- `function(example: Dict[str, List], indices: List[int]) -> List[bool]` if `with_indices=True, batched=True`
If no function is provided, defaults to an always True function: `lambda x: True`.
with_indices (`bool`, defaults to `False`):
Provide example indices to `function`. Note that in this case the signature of `function` should be `def function(example, idx): ...`.
input_columns (`str` or `List[str]`, *optional*):
The columns to be passed into `function` as
positional arguments. If `None`, a dict mapping to all formatted columns is passed as one argument.
batched (`bool`, defaults to `False`):
Provide batch of examples to `function`.
batch_size (`int`, *optional*, default `1000`):
Number of examples per batch provided to `function` if `batched=True`.
fn_kwargs (`Dict`, *optional*, default `None`):
Keyword arguments to be passed to `function`.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", split="train", streaming=True)
>>> ds = ds.filter(lambda x: x["label"] == 0)
>>> list(ds.take(3))
[{'label': 0, 'movie_review': 'simplistic , silly and tedious .'},
{'label': 0,
'movie_review': "it's so laddish and juvenile , only teenage boys could possibly find it funny ."},
{'label': 0,
'movie_review': 'exploitative and largely devoid of the depth or sophistication that would make watching such a graphic treatment of the crimes bearable .'}]
```
"""
if isinstance(input_columns, str):
input_columns = [input_columns]
# TODO(QL): keep the features (right now if we keep it it would call decode_example again on an already decoded example)
info = copy.deepcopy(self._info)
info.features = None
# We need the examples to be decoded for certain feature types like Image or Audio, so we use TypedExamplesIterable here
ex_iterable = FilteredExamplesIterable(
TypedExamplesIterable(self._ex_iterable, self._info.features, token_per_repo_id=self._token_per_repo_id)
if self._info.features is not None
else self._ex_iterable,
function=function,
with_indices=with_indices,
input_columns=input_columns,
batched=batched,
batch_size=batch_size,
fn_kwargs=fn_kwargs,
formatting=self._formatting,
)
return IterableDataset(
ex_iterable=ex_iterable,
info=info,
split=self._split,
formatting=self._formatting,
shuffling=copy.deepcopy(self._shuffling),
distributed=copy.deepcopy(self._distributed),
token_per_repo_id=self._token_per_repo_id,
)
def shuffle(
self, seed=None, generator: Optional[np.random.Generator] = None, buffer_size: int = 1000
) -> "IterableDataset":
"""
Randomly shuffles the elements of this dataset.
This dataset fills a buffer with `buffer_size` elements, then randomly samples elements from this buffer,
replacing the selected elements with new elements. For perfect shuffling, a buffer size greater than or
equal to the full size of the dataset is required.
For instance, if your dataset contains 10,000 elements but `buffer_size` is set to 1000, then `shuffle` will
initially select a random element from only the first 1000 elements in the buffer. Once an element is
selected, its space in the buffer is replaced by the next (i.e. 1,001-st) element,
maintaining the 1000 element buffer.
If the dataset is made of several shards, it also does shuffle the order of the shards.
However if the order has been fixed by using [`~datasets.IterableDataset.skip`] or [`~datasets.IterableDataset.take`]
then the order of the shards is kept unchanged.
Args:
seed (`int`, *optional*, defaults to `None`):
Random seed that will be used to shuffle the dataset.
It is used to sample from the shuffle buffer and also to shuffle the data shards.
generator (`numpy.random.Generator`, *optional*):
Numpy random Generator to use to compute the permutation of the dataset rows.
If `generator=None` (default), uses `np.random.default_rng` (the default BitGenerator (PCG64) of NumPy).
buffer_size (`int`, defaults to `1000`):
Size of the buffer.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", split="train", streaming=True)
>>> list(ds.take(3))
[{'label': 1,
'text': 'the rock is destined to be the 21st century\'s new " conan " and that he\'s going to make a splash even greater than arnold schwarzenegger , jean-claud van damme or steven segal .'},
{'label': 1,
'text': 'the gorgeously elaborate continuation of " the lord of the rings " trilogy is so huge that a column of words cannot adequately describe co-writer/director peter jackson\'s expanded vision of j . r . r . tolkien\'s middle-earth .'},
{'label': 1, 'text': 'effective but too-tepid biopic'}]
>>> shuffled_ds = ds.shuffle(seed=42)
>>> list(shuffled_ds.take(3))
[{'label': 1,
'text': "a sports movie with action that's exciting on the field and a story you care about off it ."},
{'label': 1,
'text': 'at its best , the good girl is a refreshingly adult take on adultery . . .'},
{'label': 1,
'text': "sam jones became a very lucky filmmaker the day wilco got dropped from their record label , proving that one man's ruin may be another's fortune ."}]
```
"""
if generator is None:
generator = np.random.default_rng(seed)
else:
generator = deepcopy(generator)
shuffling = ShufflingConfig(generator=generator, _original_seed=seed)
return IterableDataset(
ex_iterable=BufferShuffledExamplesIterable(
self._ex_iterable, buffer_size=buffer_size, generator=generator
).shuffle_data_sources(generator),
info=self._info.copy(),
split=self._split,
formatting=self._formatting,
shuffling=shuffling,
distributed=copy.deepcopy(self._distributed),
token_per_repo_id=self._token_per_repo_id,
)
def set_epoch(self, epoch: int):
self._epoch = epoch
def skip(self, n) -> "IterableDataset":
"""
Create a new [`IterableDataset`] that skips the first `n` elements.
Args:
n (`int`):
Number of elements to skip.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", split="train", streaming=True)
>>> list(ds.take(3))
[{'label': 1,
'text': 'the rock is destined to be the 21st century\'s new " conan " and that he\'s going to make a splash even greater than arnold schwarzenegger , jean-claud van damme or steven segal .'},
{'label': 1,
'text': 'the gorgeously elaborate continuation of " the lord of the rings " trilogy is so huge that a column of words cannot adequately describe co-writer/director peter jackson\'s expanded vision of j . r . r . tolkien\'s middle-earth .'},
{'label': 1, 'text': 'effective but too-tepid biopic'}]
>>> ds = ds.skip(1)
>>> list(ds.take(3))
[{'label': 1,
'text': 'the gorgeously elaborate continuation of " the lord of the rings " trilogy is so huge that a column of words cannot adequately describe co-writer/director peter jackson\'s expanded vision of j . r . r . tolkien\'s middle-earth .'},
{'label': 1, 'text': 'effective but too-tepid biopic'},
{'label': 1,
'text': 'if you sometimes like to go to the movies to have fun , wasabi is a good place to start .'}]
```
"""
ex_iterable = SkipExamplesIterable(self._ex_iterable, n)
return IterableDataset(
ex_iterable=ex_iterable,
info=self._info.copy(),
split=self._split,
formatting=self._formatting,
shuffling=copy.deepcopy(self._shuffling),
distributed=copy.deepcopy(self._distributed),
token_per_repo_id=self._token_per_repo_id,
)
def take(self, n) -> "IterableDataset":
"""
Create a new [`IterableDataset`] with only the first `n` elements.
Args:
n (`int`):
Number of elements to take.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", split="train", streaming=True)
>>> small_ds = ds.take(2)
>>> list(small_ds)
[{'label': 1,
'text': 'the rock is destined to be the 21st century\'s new " conan " and that he\'s going to make a splash even greater than arnold schwarzenegger , jean-claud van damme or steven segal .'},
{'label': 1,
'text': 'the gorgeously elaborate continuation of " the lord of the rings " trilogy is so huge that a column of words cannot adequately describe co-writer/director peter jackson\'s expanded vision of j . r . r . tolkien\'s middle-earth .'}]
```
"""
ex_iterable = TakeExamplesIterable(self._ex_iterable, n)
return IterableDataset(
ex_iterable=ex_iterable,
info=self._info.copy(),
split=self._split,
formatting=self._formatting,
shuffling=copy.deepcopy(self._shuffling),
distributed=copy.deepcopy(self._distributed),
token_per_repo_id=self._token_per_repo_id,
)
@property
def column_names(self) -> Optional[List[str]]:
"""Names of the columns in the dataset.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", split="validation", streaming=True)
>>> ds.column_names
['text', 'label']
```
"""
return list(self._info.features.keys()) if self._info.features is not None else None
def add_column(self, name: str, column: Union[list, np.array]) -> "IterableDataset":
"""Add column to Dataset.
Args:
name (str): Column name.
column (list or np.array): Column data to be added.
Returns:
`IterableDataset`
"""
return self.map(partial(add_column_fn, name=name, column=column), with_indices=True)
def rename_column(self, original_column_name: str, new_column_name: str) -> "IterableDataset":
"""
Rename a column in the dataset, and move the features associated to the original column under the new column
name.
Args:
original_column_name (`str`):
Name of the column to rename.
new_column_name (`str`):
New name for the column.
Returns:
`IterableDataset`: A copy of the dataset with a renamed column.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", split="train", streaming=True)
>>> next(iter(ds))
{'label': 1,
'text': 'the rock is destined to be the 21st century\'s new " conan " and that he\'s going to make a splash even greater than arnold schwarzenegger , jean-claud van damme or steven segal .'}
>>> ds = ds.rename_column("text", "movie_review")
>>> next(iter(ds))
{'label': 1,
'movie_review': 'the rock is destined to be the 21st century\'s new " conan " and that he\'s going to make a splash even greater than arnold schwarzenegger , jean-claud van damme or steven segal .'}
```
"""
return self.rename_columns({original_column_name: new_column_name})
def rename_columns(self, column_mapping: Dict[str, str]) -> "IterableDataset":
"""
Rename several columns in the dataset, and move the features associated to the original columns under
the new column names.
Args:
column_mapping (`Dict[str, str]`): A mapping of columns to rename to their new names
Returns:
`IterableDataset`: A copy of the dataset with renamed columns
"""
original_features = self._info.features.copy() if self._info.features else None
ds_iterable = self.map(
partial(_rename_columns_fn, column_mapping=column_mapping), remove_columns=list(column_mapping)
)
if original_features is not None:
ds_iterable._info.features = Features(
{
column_mapping[col] if col in column_mapping.keys() else col: feature
for col, feature in original_features.items()
}
)
# check that it's still valid, especially with regard to task templates
try:
ds_iterable._info.copy()
except ValueError:
ds_iterable._info.task_templates = None
return ds_iterable
def remove_columns(self, column_names: Union[str, List[str]]) -> "IterableDataset":
"""
Remove one or several column(s) in the dataset and the features associated to them.
The removal is done on-the-fly on the examples when iterating over the dataset.
Args:
column_names (`Union[str, List[str]]`):
Name of the column(s) to remove.
Returns:
`IterableDataset`: A copy of the dataset object without the columns to remove.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", split="train", streaming=True)
>>> next(iter(ds))
{'text': 'the rock is destined to be the 21st century\'s new " conan " and that he\'s going to make a splash even greater than arnold schwarzenegger , jean-claud van damme or steven segal .', 'label': 1}
>>> ds = ds.remove_columns("label")
>>> next(iter(ds))
{'text': 'the rock is destined to be the 21st century\'s new " conan " and that he\'s going to make a splash even greater than arnold schwarzenegger , jean-claud van damme or steven segal .'}
```
"""
original_features = self._info.features.copy() if self._info.features else None
ds_iterable = self.map(remove_columns=column_names)
if original_features is not None:
ds_iterable._info.features = original_features.copy()
for col, _ in original_features.items():
if col in column_names:
del ds_iterable._info.features[col]
# check that it's still valid, especially with regard to task templates
try:
ds_iterable._info.copy()
except ValueError:
ds_iterable._info.task_templates = None
return ds_iterable
def select_columns(self, column_names: Union[str, List[str]]) -> "IterableDataset":
"""Select one or several column(s) in the dataset and the features
associated to them. The selection is done on-the-fly on the examples
when iterating over the dataset.
Args:
column_names (`Union[str, List[str]]`):
Name of the column(s) to select.
Returns:
`IterableDataset`: A copy of the dataset object with selected columns.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", split="train", streaming=True)
>>> next(iter(ds))
{'text': 'the rock is destined to be the 21st century\'s new " conan " and that he\'s going to make a splash even greater than arnold schwarzenegger , jean-claud van damme or steven segal .', 'label': 1}
>>> ds = ds.select_columns("text")
>>> next(iter(ds))
{'text': 'the rock is destined to be the 21st century\'s new " conan " and that he\'s going to make a splash even greater than arnold schwarzenegger , jean-claud van damme or steven segal .'}
```
"""
if isinstance(column_names, str):
column_names = [column_names]
if self._info:
info = copy.deepcopy(self._info)
if self._info.features is not None:
for column_name in column_names:
if column_name not in self._info.features:
raise ValueError(
f"Column name {column_name} not in the "
"dataset. Columns in the dataset: "
f"{list(self._info.features.keys())}."
)
info.features = Features({c: info.features[c] for c in column_names})
# check that it's still valid, especially with regard to task templates
try:
info.copy()
except ValueError:
info.task_templates = None
ex_iterable = SelectColumnsIterable(self._ex_iterable, column_names)
return IterableDataset(
ex_iterable=ex_iterable,
info=info,
split=self._split,
formatting=self._formatting,
shuffling=self._shuffling,
distributed=self._distributed,
token_per_repo_id=self._token_per_repo_id,
)
def cast_column(self, column: str, feature: FeatureType) -> "IterableDataset":
"""Cast column to feature for decoding.
Args:
column (`str`):
Column name.
feature (`Feature`):
Target feature.
Returns:
`IterableDataset`
Example:
```py
>>> from datasets import load_dataset, Audio
>>> ds = load_dataset("PolyAI/minds14", name="en-US", split="train", streaming=True)
>>> ds.features
{'audio': Audio(sampling_rate=8000, mono=True, decode=True, id=None),
'english_transcription': Value(dtype='string', id=None),
'intent_class': ClassLabel(num_classes=14, names=['abroad', 'address', 'app_error', 'atm_limit', 'balance', 'business_loan', 'card_issues', 'cash_deposit', 'direct_debit', 'freeze', 'high_value_payment', 'joint_account', 'latest_transactions', 'pay_bill'], id=None),
'lang_id': ClassLabel(num_classes=14, names=['cs-CZ', 'de-DE', 'en-AU', 'en-GB', 'en-US', 'es-ES', 'fr-FR', 'it-IT', 'ko-KR', 'nl-NL', 'pl-PL', 'pt-PT', 'ru-RU', 'zh-CN'], id=None),
'path': Value(dtype='string', id=None),
'transcription': Value(dtype='string', id=None)}
>>> ds = ds.cast_column("audio", Audio(sampling_rate=16000))
>>> ds.features
{'audio': Audio(sampling_rate=16000, mono=True, decode=True, id=None),
'english_transcription': Value(dtype='string', id=None),
'intent_class': ClassLabel(num_classes=14, names=['abroad', 'address', 'app_error', 'atm_limit', 'balance', 'business_loan', 'card_issues', 'cash_deposit', 'direct_debit', 'freeze', 'high_value_payment', 'joint_account', 'latest_transactions', 'pay_bill'], id=None),
'lang_id': ClassLabel(num_classes=14, names=['cs-CZ', 'de-DE', 'en-AU', 'en-GB', 'en-US', 'es-ES', 'fr-FR', 'it-IT', 'ko-KR', 'nl-NL', 'pl-PL', 'pt-PT', 'ru-RU', 'zh-CN'], id=None),
'path': Value(dtype='string', id=None),
'transcription': Value(dtype='string', id=None)}
```
"""
info = self._info.copy()
info.features[column] = feature
# check that it's still valid, especially with regard to task templates
try:
info.copy()
except ValueError:
info.task_templates = None
return IterableDataset(
ex_iterable=self._ex_iterable,
info=info,
split=self._split,
formatting=self._formatting,
shuffling=copy.deepcopy(self._shuffling),
distributed=copy.deepcopy(self._distributed),
token_per_repo_id=self._token_per_repo_id,
)
def cast(
self,
features: Features,
) -> "IterableDataset":
"""
Cast the dataset to a new set of features.
Args:
features ([`Features`]):
New features to cast the dataset to.
The name of the fields in the features must match the current column names.
The type of the data must also be convertible from one type to the other.
For non-trivial conversion, e.g. `string` <-> `ClassLabel` you should use [`~Dataset.map`] to update the Dataset.
Returns:
`IterableDataset`: A copy of the dataset with casted features.
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", split="train", streaming=True)
>>> ds.features
{'label': ClassLabel(num_classes=2, names=['neg', 'pos'], id=None),
'text': Value(dtype='string', id=None)}
>>> new_features = ds.features.copy()
>>> new_features["label"] = ClassLabel(names=["bad", "good"])
>>> new_features["text"] = Value("large_string")
>>> ds = ds.cast(new_features)
>>> ds.features
{'label': ClassLabel(num_classes=2, names=['bad', 'good'], id=None),
'text': Value(dtype='large_string', id=None)}
```
"""
info = self._info.copy()
info.features = features
# check that it's still valid, especially with regard to task templates
try:
info.copy()
except ValueError:
info.task_templates = None
return IterableDataset(
ex_iterable=self._ex_iterable,
info=info,
split=self._split,
formatting=self._formatting,
shuffling=copy.deepcopy(self._shuffling),
distributed=copy.deepcopy(self._distributed),
token_per_repo_id=self._token_per_repo_id,
)
def _step(self, step: int, offset: int) -> "IterableDataset":
ex_iterable = StepExamplesIterable(self._ex_iterable, step=step, offset=offset)
return IterableDataset(
ex_iterable=ex_iterable,
info=self._info.copy(),
split=self._split,
formatting=self._formatting,
shuffling=copy.deepcopy(self._shuffling),
distributed=copy.deepcopy(self._distributed),
token_per_repo_id=self._token_per_repo_id,
)
def _resolve_features(self):
if self.features is not None:
return self
elif isinstance(self._ex_iterable, TypedExamplesIterable):
features = self._ex_iterable.features
else:
features = _infer_features_from_batch(self.with_format(None)._head())
info = self.info.copy()
info.features = features
return IterableDataset(
ex_iterable=self._ex_iterable,
info=info,
split=self._split,
formatting=self._formatting,
shuffling=copy.deepcopy(self._shuffling),
distributed=copy.deepcopy(self._distributed),
token_per_repo_id=self._token_per_repo_id,
)
def _concatenate_iterable_datasets(
dsets: List[IterableDataset],
info: Optional[DatasetInfo] = None,
split: Optional[NamedSplit] = None,
axis: int = 0,
) -> IterableDataset:
"""
Converts a list of `IterableDataset` with the same schema into a single `IterableDataset`.
Missing data are filled with None values.
<Added version="2.4.0"/>
Args:
dsets (`List[datasets.IterableDataset]`): List of Datasets to concatenate.
info (`DatasetInfo`, optional): Dataset information, like description, citation, etc.
split (`NamedSplit`, optional): Name of the dataset split.
axis (``{0, 1}``, default ``0``, meaning over rows):
Axis to concatenate over, where ``0`` means over rows (vertically) and ``1`` means over columns
(horizontally).
*New in version 1.6.0*
Example:
```py
>>> ds3 = _concatenate_iterable_datasets([ds1, ds2])
```
"""
dsets = [d._resolve_features() for d in dsets]
# Perform checks (and a potentional cast if axis=0)
if axis == 0:
_check_if_features_can_be_aligned([dset.features for dset in dsets])
else:
_check_column_names([col_name for dset in dsets for col_name in dset.features])
# TODO: improve this to account for a mix of ClassLabel and Value for example
# right now it would keep the type of the first dataset in the list
features = Features(
{k: v for features in _align_features([dset.features for dset in dsets]) for k, v in features.items()}
)
ex_iterables = [d._ex_iterable for d in dsets]
if axis == 0:
ex_iterable = VerticallyConcatenatedMultiSourcesExamplesIterable(ex_iterables)
else:
ex_iterable = HorizontallyConcatenatedMultiSourcesExamplesIterable(ex_iterables)
# Set new info - we update the features
# setting the features also ensures to fill missing columns with None
if info is None:
info = DatasetInfo.from_merge([d.info for d in dsets])
else:
info = info.copy()
info.features = features
# Get all the auth tokens per repository - in case the datasets come from different private repositories
token_per_repo_id = {repo_id: token for dataset in dsets for repo_id, token in dataset._token_per_repo_id.items()}
# Return new daset
return IterableDataset(ex_iterable=ex_iterable, info=info, split=split, token_per_repo_id=token_per_repo_id)
def _interleave_iterable_datasets(
datasets: List[IterableDataset],
probabilities: Optional[List[float]] = None,
seed: Optional[int] = None,
info: Optional[DatasetInfo] = None,
split: Optional[NamedSplit] = None,
stopping_strategy: Literal["first_exhausted", "all_exhausted"] = "first_exhausted",
) -> IterableDataset:
"""
Interleave several iterable datasets (sources) into a single iterable dataset.
The new iterable dataset alternates between the sources to yield examples.
If `probabilities = None` (default) the iterable dataset will cycles through the sources in order for each next example in the iteration.
If `probabilities` is not `None, the iterable dataset will sample a random source according to the provided probabilities for each next examples in the iteration.
<Added version="2.4.0"/>
Args:
datasets (`List[IterableDataset]`): list of datasets to interleave
probabilities (`List[float]`, optional, default None): If specified, the new iterable dataset samples
examples from one source at a time according to these probabilities.
seed (`int`, optional, default None): The random seed used to choose a source for each example.
stopping_strategy (`str`, defaults to `first_exhausted`):
Two strategies are proposed right now.
By default, `first_exhausted` is an undersampling strategy, i.e the dataset construction is stopped as soon as one dataset has ran out of samples.
If the strategy is `all_exhausted`, we use an oversampling strategy, i.e the dataset construction is stopped as soon as every samples of every dataset has been added at least once.
Note that if the strategy is `all_exhausted`, the interleaved dataset size can get enormous:
- with no probabilities, the resulting dataset will have max_length_datasets*nb_dataset samples.
- with given probabilities, the resulting dataset will have more samples if some datasets have really low probability of visiting.
Output:
`datasets.IterableDataset`
"""
datasets = [d._resolve_features() for d in datasets]
# Perform checks
_check_if_features_can_be_aligned([dset.features for dset in datasets])
# TODO: improve this to account for a mix of ClassLabel and Value for example
# right now it would keep the type of the first dataset in the list
features = Features(
{k: v for features in _align_features([dset.features for dset in datasets]) for k, v in features.items()}
)
ex_iterables = [d._ex_iterable for d in datasets]
# Use cycling or random cycling of sources
if probabilities is None:
ex_iterable = CyclingMultiSourcesExamplesIterable(ex_iterables, stopping_strategy=stopping_strategy)
else:
generator = np.random.default_rng(seed)
ex_iterable = RandomlyCyclingMultiSourcesExamplesIterable(
ex_iterables, generator=generator, probabilities=probabilities, stopping_strategy=stopping_strategy
)
# Set new info - we update the features
# setting the features also ensures to fill missing columns with None
if info is None:
info = DatasetInfo.from_merge([d.info for d in datasets])
else:
info = info.copy()
info.features = features
# Get all the auth tokens per repository - in case the datasets come from different private repositories
token_per_repo_id = {
repo_id: token for dataset in datasets for repo_id, token in dataset._token_per_repo_id.items()
}
# Return new daset
return IterableDataset(ex_iterable=ex_iterable, info=info, split=split, token_per_repo_id=token_per_repo_id)
def _split_by_node_iterable_dataset(dataset: IterableDataset, rank: int, world_size: int) -> IterableDataset:
"""
Split an iterable dataset for the node at rank `rank` in a pool of nodes of size `world_size`.
If the dataset has a number of shards that is a factor of `world_size` (i.e. if `dataset.n_shards % world_size == 0`),
then the shards are evenly assigned across the nodes, which is the most optimized.
Otherwise, each node keeps 1 example out of `world_size`, skipping the other examples.
Args:
dataset ([`IterableDataset`]):
The iterable dataset to split by node.
rank (`int`):
Rank of the current node.
world_size (`int`):
Total number of nodes.
Returns:
[`IterableDataset`]: The iterable dataset to be used on the node at rank `rank`.
"""
if dataset._distributed:
world_size = world_size * dataset._distributed.world_size
rank = world_size * dataset._distributed.rank + rank
distributed = DistributedConfig(rank=rank, world_size=world_size)
return IterableDataset(
ex_iterable=dataset._ex_iterable,
info=dataset._info.copy(),
split=dataset._split,
formatting=dataset._formatting,
shuffling=copy.deepcopy(dataset._shuffling),
distributed=distributed,
token_per_repo_id=dataset._token_per_repo_id,
)
| 0
|
hf_public_repos/datasets/src
|
hf_public_repos/datasets/src/datasets/info.py
|
# Copyright 2020 The HuggingFace Datasets Authors and the TensorFlow Datasets Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# Lint as: python3
""" DatasetInfo and MetricInfo record information we know about a dataset and a metric.
This includes things that we know about the dataset statically, i.e.:
- description
- canonical location
- does it have validation and tests splits
- size
- etc.
This also includes the things that can and should be computed once we've
processed the dataset as well:
- number of examples (in each split)
- etc.
"""
import copy
import dataclasses
import json
import os
import posixpath
import warnings
from dataclasses import dataclass
from pathlib import Path
from typing import ClassVar, Dict, List, Optional, Union
import fsspec
from huggingface_hub import DatasetCard, DatasetCardData
from . import config
from .features import Features, Value
from .splits import SplitDict
from .tasks import TaskTemplate, task_template_from_dict
from .utils import Version
from .utils.logging import get_logger
from .utils.py_utils import asdict, unique_values
logger = get_logger(__name__)
@dataclass
class SupervisedKeysData:
input: str = ""
output: str = ""
@dataclass
class DownloadChecksumsEntryData:
key: str = ""
value: str = ""
class MissingCachedSizesConfigError(Exception):
"""The expected cached sizes of the download file are missing."""
class NonMatchingCachedSizesError(Exception):
"""The prepared split doesn't have expected sizes."""
@dataclass
class PostProcessedInfo:
features: Optional[Features] = None
resources_checksums: Optional[dict] = None
def __post_init__(self):
# Convert back to the correct classes when we reload from dict
if self.features is not None and not isinstance(self.features, Features):
self.features = Features.from_dict(self.features)
@classmethod
def from_dict(cls, post_processed_info_dict: dict) -> "PostProcessedInfo":
field_names = {f.name for f in dataclasses.fields(cls)}
return cls(**{k: v for k, v in post_processed_info_dict.items() if k in field_names})
@dataclass
class DatasetInfo:
"""Information about a dataset.
`DatasetInfo` documents datasets, including its name, version, and features.
See the constructor arguments and properties for a full list.
Not all fields are known on construction and may be updated later.
Attributes:
description (`str`):
A description of the dataset.
citation (`str`):
A BibTeX citation of the dataset.
homepage (`str`):
A URL to the official homepage for the dataset.
license (`str`):
The dataset's license. It can be the name of the license or a paragraph containing the terms of the license.
features ([`Features`], *optional*):
The features used to specify the dataset's column types.
post_processed (`PostProcessedInfo`, *optional*):
Information regarding the resources of a possible post-processing of a dataset. For example, it can contain the information of an index.
supervised_keys (`SupervisedKeysData`, *optional*):
Specifies the input feature and the label for supervised learning if applicable for the dataset (legacy from TFDS).
builder_name (`str`, *optional*):
The name of the `GeneratorBasedBuilder` subclass used to create the dataset. Usually matched to the corresponding script name. It is also the snake_case version of the dataset builder class name.
config_name (`str`, *optional*):
The name of the configuration derived from [`BuilderConfig`].
version (`str` or [`Version`], *optional*):
The version of the dataset.
splits (`dict`, *optional*):
The mapping between split name and metadata.
download_checksums (`dict`, *optional*):
The mapping between the URL to download the dataset's checksums and corresponding metadata.
download_size (`int`, *optional*):
The size of the files to download to generate the dataset, in bytes.
post_processing_size (`int`, *optional*):
Size of the dataset in bytes after post-processing, if any.
dataset_size (`int`, *optional*):
The combined size in bytes of the Arrow tables for all splits.
size_in_bytes (`int`, *optional*):
The combined size in bytes of all files associated with the dataset (downloaded files + Arrow files).
task_templates (`List[TaskTemplate]`, *optional*):
The task templates to prepare the dataset for during training and evaluation. Each template casts the dataset's [`Features`] to standardized column names and types as detailed in `datasets.tasks`.
**config_kwargs (additional keyword arguments):
Keyword arguments to be passed to the [`BuilderConfig`] and used in the [`DatasetBuilder`].
"""
# Set in the dataset scripts
description: str = dataclasses.field(default_factory=str)
citation: str = dataclasses.field(default_factory=str)
homepage: str = dataclasses.field(default_factory=str)
license: str = dataclasses.field(default_factory=str)
features: Optional[Features] = None
post_processed: Optional[PostProcessedInfo] = None
supervised_keys: Optional[SupervisedKeysData] = None
task_templates: Optional[List[TaskTemplate]] = None
# Set later by the builder
builder_name: Optional[str] = None
dataset_name: Optional[str] = None # for packaged builders, to be different from builder_name
config_name: Optional[str] = None
version: Optional[Union[str, Version]] = None
# Set later by `download_and_prepare`
splits: Optional[dict] = None
download_checksums: Optional[dict] = None
download_size: Optional[int] = None
post_processing_size: Optional[int] = None
dataset_size: Optional[int] = None
size_in_bytes: Optional[int] = None
_INCLUDED_INFO_IN_YAML: ClassVar[List[str]] = [
"config_name",
"download_size",
"dataset_size",
"features",
"splits",
]
def __post_init__(self):
# Convert back to the correct classes when we reload from dict
if self.features is not None and not isinstance(self.features, Features):
self.features = Features.from_dict(self.features)
if self.post_processed is not None and not isinstance(self.post_processed, PostProcessedInfo):
self.post_processed = PostProcessedInfo.from_dict(self.post_processed)
if self.version is not None and not isinstance(self.version, Version):
if isinstance(self.version, str):
self.version = Version(self.version)
else:
self.version = Version.from_dict(self.version)
if self.splits is not None and not isinstance(self.splits, SplitDict):
self.splits = SplitDict.from_split_dict(self.splits)
if self.supervised_keys is not None and not isinstance(self.supervised_keys, SupervisedKeysData):
if isinstance(self.supervised_keys, (tuple, list)):
self.supervised_keys = SupervisedKeysData(*self.supervised_keys)
else:
self.supervised_keys = SupervisedKeysData(**self.supervised_keys)
# Parse and make a list of templates
if self.task_templates is not None:
if isinstance(self.task_templates, (list, tuple)):
templates = [
template if isinstance(template, TaskTemplate) else task_template_from_dict(template)
for template in self.task_templates
]
self.task_templates = [template for template in templates if template is not None]
elif isinstance(self.task_templates, TaskTemplate):
self.task_templates = [self.task_templates]
else:
template = task_template_from_dict(self.task_templates)
self.task_templates = [template] if template is not None else []
# Align task templates with features
if self.task_templates is not None:
self.task_templates = list(self.task_templates)
if self.features is not None:
self.task_templates = [
template.align_with_features(self.features) for template in (self.task_templates)
]
def write_to_directory(
self, dataset_info_dir, pretty_print=False, fs="deprecated", storage_options: Optional[dict] = None
):
"""Write `DatasetInfo` and license (if present) as JSON files to `dataset_info_dir`.
Args:
dataset_info_dir (`str`):
Destination directory.
pretty_print (`bool`, defaults to `False`):
If `True`, the JSON will be pretty-printed with the indent level of 4.
fs (`fsspec.spec.AbstractFileSystem`, *optional*):
Instance of the remote filesystem used to download the files from.
<Deprecated version="2.9.0">
`fs` was deprecated in version 2.9.0 and will be removed in 3.0.0.
Please use `storage_options` instead, e.g. `storage_options=fs.storage_options`.
</Deprecated>
storage_options (`dict`, *optional*):
Key/value pairs to be passed on to the file-system backend, if any.
<Added version="2.9.0"/>
Example:
```py
>>> from datasets import load_dataset
>>> ds = load_dataset("rotten_tomatoes", split="validation")
>>> ds.info.write_to_directory("/path/to/directory/")
```
"""
if fs != "deprecated":
warnings.warn(
"'fs' was deprecated in favor of 'storage_options' in version 2.9.0 and will be removed in 3.0.0.\n"
"You can remove this warning by passing 'storage_options=fs.storage_options' instead.",
FutureWarning,
)
storage_options = fs.storage_options
fs: fsspec.AbstractFileSystem
fs, _, _ = fsspec.get_fs_token_paths(dataset_info_dir, storage_options=storage_options)
with fs.open(posixpath.join(dataset_info_dir, config.DATASET_INFO_FILENAME), "wb") as f:
self._dump_info(f, pretty_print=pretty_print)
if self.license:
with fs.open(posixpath.join(dataset_info_dir, config.LICENSE_FILENAME), "wb") as f:
self._dump_license(f)
def _dump_info(self, file, pretty_print=False):
"""Dump info in `file` file-like object open in bytes mode (to support remote files)"""
file.write(json.dumps(asdict(self), indent=4 if pretty_print else None).encode("utf-8"))
def _dump_license(self, file):
"""Dump license in `file` file-like object open in bytes mode (to support remote files)"""
file.write(self.license.encode("utf-8"))
@classmethod
def from_merge(cls, dataset_infos: List["DatasetInfo"]):
dataset_infos = [dset_info.copy() for dset_info in dataset_infos if dset_info is not None]
description = "\n\n".join(unique_values(info.description for info in dataset_infos)).strip()
citation = "\n\n".join(unique_values(info.citation for info in dataset_infos)).strip()
homepage = "\n\n".join(unique_values(info.homepage for info in dataset_infos)).strip()
license = "\n\n".join(unique_values(info.license for info in dataset_infos)).strip()
features = None
supervised_keys = None
task_templates = None
# Find common task templates across all dataset infos
all_task_templates = [info.task_templates for info in dataset_infos if info.task_templates is not None]
if len(all_task_templates) > 1:
task_templates = list(set(all_task_templates[0]).intersection(*all_task_templates[1:]))
elif len(all_task_templates):
task_templates = list(set(all_task_templates[0]))
# If no common task templates found, replace empty list with None
task_templates = task_templates if task_templates else None
return cls(
description=description,
citation=citation,
homepage=homepage,
license=license,
features=features,
supervised_keys=supervised_keys,
task_templates=task_templates,
)
@classmethod
def from_directory(
cls, dataset_info_dir: str, fs="deprecated", storage_options: Optional[dict] = None
) -> "DatasetInfo":
"""Create [`DatasetInfo`] from the JSON file in `dataset_info_dir`.
This function updates all the dynamically generated fields (num_examples,
hash, time of creation,...) of the [`DatasetInfo`].
This will overwrite all previous metadata.
Args:
dataset_info_dir (`str`):
The directory containing the metadata file. This
should be the root directory of a specific dataset version.
fs (`fsspec.spec.AbstractFileSystem`, *optional*):
Instance of the remote filesystem used to download the files from.
<Deprecated version="2.9.0">
`fs` was deprecated in version 2.9.0 and will be removed in 3.0.0.
Please use `storage_options` instead, e.g. `storage_options=fs.storage_options`.
</Deprecated>
storage_options (`dict`, *optional*):
Key/value pairs to be passed on to the file-system backend, if any.
<Added version="2.9.0"/>
Example:
```py
>>> from datasets import DatasetInfo
>>> ds_info = DatasetInfo.from_directory("/path/to/directory/")
```
"""
if fs != "deprecated":
warnings.warn(
"'fs' was deprecated in favor of 'storage_options' in version 2.9.0 and will be removed in 3.0.0.\n"
"You can remove this warning by passing 'storage_options=fs.storage_options' instead.",
FutureWarning,
)
storage_options = fs.storage_options
fs: fsspec.AbstractFileSystem
fs, _, _ = fsspec.get_fs_token_paths(dataset_info_dir, storage_options=storage_options)
logger.info(f"Loading Dataset info from {dataset_info_dir}")
if not dataset_info_dir:
raise ValueError("Calling DatasetInfo.from_directory() with undefined dataset_info_dir.")
with fs.open(posixpath.join(dataset_info_dir, config.DATASET_INFO_FILENAME), "r", encoding="utf-8") as f:
dataset_info_dict = json.load(f)
return cls.from_dict(dataset_info_dict)
@classmethod
def from_dict(cls, dataset_info_dict: dict) -> "DatasetInfo":
field_names = {f.name for f in dataclasses.fields(cls)}
return cls(**{k: v for k, v in dataset_info_dict.items() if k in field_names})
def update(self, other_dataset_info: "DatasetInfo", ignore_none=True):
self_dict = self.__dict__
self_dict.update(
**{
k: copy.deepcopy(v)
for k, v in other_dataset_info.__dict__.items()
if (v is not None or not ignore_none)
}
)
def copy(self) -> "DatasetInfo":
return self.__class__(**{k: copy.deepcopy(v) for k, v in self.__dict__.items()})
def _to_yaml_dict(self) -> dict:
yaml_dict = {}
dataset_info_dict = asdict(self)
for key in dataset_info_dict:
if key in self._INCLUDED_INFO_IN_YAML:
value = getattr(self, key)
if hasattr(value, "_to_yaml_list"): # Features, SplitDict
yaml_dict[key] = value._to_yaml_list()
elif hasattr(value, "_to_yaml_string"): # Version
yaml_dict[key] = value._to_yaml_string()
else:
yaml_dict[key] = value
return yaml_dict
@classmethod
def _from_yaml_dict(cls, yaml_data: dict) -> "DatasetInfo":
yaml_data = copy.deepcopy(yaml_data)
if yaml_data.get("features") is not None:
yaml_data["features"] = Features._from_yaml_list(yaml_data["features"])
if yaml_data.get("splits") is not None:
yaml_data["splits"] = SplitDict._from_yaml_list(yaml_data["splits"])
field_names = {f.name for f in dataclasses.fields(cls)}
return cls(**{k: v for k, v in yaml_data.items() if k in field_names})
class DatasetInfosDict(Dict[str, DatasetInfo]):
def write_to_directory(self, dataset_infos_dir, overwrite=False, pretty_print=False) -> None:
total_dataset_infos = {}
dataset_infos_path = os.path.join(dataset_infos_dir, config.DATASETDICT_INFOS_FILENAME)
dataset_readme_path = os.path.join(dataset_infos_dir, "README.md")
if not overwrite:
total_dataset_infos = self.from_directory(dataset_infos_dir)
total_dataset_infos.update(self)
if os.path.exists(dataset_infos_path):
# for backward compatibility, let's update the JSON file if it exists
with open(dataset_infos_path, "w", encoding="utf-8") as f:
dataset_infos_dict = {
config_name: asdict(dset_info) for config_name, dset_info in total_dataset_infos.items()
}
json.dump(dataset_infos_dict, f, indent=4 if pretty_print else None)
# Dump the infos in the YAML part of the README.md file
if os.path.exists(dataset_readme_path):
dataset_card = DatasetCard.load(dataset_readme_path)
dataset_card_data = dataset_card.data
else:
dataset_card = None
dataset_card_data = DatasetCardData()
if total_dataset_infos:
total_dataset_infos.to_dataset_card_data(dataset_card_data)
dataset_card = (
DatasetCard("---\n" + str(dataset_card_data) + "\n---\n") if dataset_card is None else dataset_card
)
dataset_card.save(Path(dataset_readme_path))
@classmethod
def from_directory(cls, dataset_infos_dir) -> "DatasetInfosDict":
logger.info(f"Loading Dataset Infos from {dataset_infos_dir}")
# Load the info from the YAML part of README.md
if os.path.exists(os.path.join(dataset_infos_dir, "README.md")):
dataset_card_data = DatasetCard.load(Path(dataset_infos_dir) / "README.md").data
if "dataset_info" in dataset_card_data:
return cls.from_dataset_card_data(dataset_card_data)
if os.path.exists(os.path.join(dataset_infos_dir, config.DATASETDICT_INFOS_FILENAME)):
# this is just to have backward compatibility with dataset_infos.json files
with open(os.path.join(dataset_infos_dir, config.DATASETDICT_INFOS_FILENAME), encoding="utf-8") as f:
return cls(
{
config_name: DatasetInfo.from_dict(dataset_info_dict)
for config_name, dataset_info_dict in json.load(f).items()
}
)
else:
return cls()
@classmethod
def from_dataset_card_data(cls, dataset_card_data: DatasetCardData) -> "DatasetInfosDict":
if isinstance(dataset_card_data.get("dataset_info"), (list, dict)):
if isinstance(dataset_card_data["dataset_info"], list):
return cls(
{
dataset_info_yaml_dict.get("config_name", "default"): DatasetInfo._from_yaml_dict(
dataset_info_yaml_dict
)
for dataset_info_yaml_dict in dataset_card_data["dataset_info"]
}
)
else:
dataset_info = DatasetInfo._from_yaml_dict(dataset_card_data["dataset_info"])
dataset_info.config_name = dataset_card_data["dataset_info"].get("config_name", "default")
return cls({dataset_info.config_name: dataset_info})
else:
return cls()
def to_dataset_card_data(self, dataset_card_data: DatasetCardData) -> None:
if self:
# first get existing metadata info
if "dataset_info" in dataset_card_data and isinstance(dataset_card_data["dataset_info"], dict):
dataset_metadata_infos = {
dataset_card_data["dataset_info"].get("config_name", "default"): dataset_card_data["dataset_info"]
}
elif "dataset_info" in dataset_card_data and isinstance(dataset_card_data["dataset_info"], list):
dataset_metadata_infos = {
config_metadata["config_name"]: config_metadata
for config_metadata in dataset_card_data["dataset_info"]
}
else:
dataset_metadata_infos = {}
# update/rewrite existing metadata info with the one to dump
total_dataset_infos = {
**dataset_metadata_infos,
**{config_name: dset_info._to_yaml_dict() for config_name, dset_info in self.items()},
}
# the config_name from the dataset_infos_dict takes over the config_name of the DatasetInfo
for config_name, dset_info_yaml_dict in total_dataset_infos.items():
dset_info_yaml_dict["config_name"] = config_name
if len(total_dataset_infos) == 1:
# use a struct instead of a list of configurations, since there's only one
dataset_card_data["dataset_info"] = next(iter(total_dataset_infos.values()))
config_name = dataset_card_data["dataset_info"].pop("config_name", None)
if config_name != "default":
# if config_name is not "default" preserve it and put at the first position
dataset_card_data["dataset_info"] = {
"config_name": config_name,
**dataset_card_data["dataset_info"],
}
else:
dataset_card_data["dataset_info"] = []
for config_name, dataset_info_yaml_dict in sorted(total_dataset_infos.items()):
# add the config_name field in first position
dataset_info_yaml_dict.pop("config_name", None)
dataset_info_yaml_dict = {"config_name": config_name, **dataset_info_yaml_dict}
dataset_card_data["dataset_info"].append(dataset_info_yaml_dict)
@dataclass
class MetricInfo:
"""Information about a metric.
`MetricInfo` documents a metric, including its name, version, and features.
See the constructor arguments and properties for a full list.
Note: Not all fields are known on construction and may be updated later.
"""
# Set in the dataset scripts
description: str
citation: str
features: Features
inputs_description: str = dataclasses.field(default_factory=str)
homepage: str = dataclasses.field(default_factory=str)
license: str = dataclasses.field(default_factory=str)
codebase_urls: List[str] = dataclasses.field(default_factory=list)
reference_urls: List[str] = dataclasses.field(default_factory=list)
streamable: bool = False
format: Optional[str] = None
# Set later by the builder
metric_name: Optional[str] = None
config_name: Optional[str] = None
experiment_id: Optional[str] = None
def __post_init__(self):
if self.format is not None:
for key, value in self.features.items():
if not isinstance(value, Value):
raise ValueError(
f"When using 'numpy' format, all features should be a `datasets.Value` feature. "
f"Here {key} is an instance of {value.__class__.__name__}"
)
def write_to_directory(self, metric_info_dir, pretty_print=False):
"""Write `MetricInfo` as JSON to `metric_info_dir`.
Also save the license separately in LICENCE.
If `pretty_print` is True, the JSON will be pretty-printed with the indent level of 4.
Example:
```py
>>> from datasets import load_metric
>>> metric = load_metric("accuracy")
>>> metric.info.write_to_directory("/path/to/directory/")
```
"""
with open(os.path.join(metric_info_dir, config.METRIC_INFO_FILENAME), "w", encoding="utf-8") as f:
json.dump(asdict(self), f, indent=4 if pretty_print else None)
if self.license:
with open(os.path.join(metric_info_dir, config.LICENSE_FILENAME), "w", encoding="utf-8") as f:
f.write(self.license)
@classmethod
def from_directory(cls, metric_info_dir) -> "MetricInfo":
"""Create MetricInfo from the JSON file in `metric_info_dir`.
Args:
metric_info_dir: `str` The directory containing the metadata file. This
should be the root directory of a specific dataset version.
Example:
```py
>>> from datasets import MetricInfo
>>> metric_info = MetricInfo.from_directory("/path/to/directory/")
```
"""
logger.info(f"Loading Metric info from {metric_info_dir}")
if not metric_info_dir:
raise ValueError("Calling MetricInfo.from_directory() with undefined metric_info_dir.")
with open(os.path.join(metric_info_dir, config.METRIC_INFO_FILENAME), encoding="utf-8") as f:
metric_info_dict = json.load(f)
return cls.from_dict(metric_info_dict)
@classmethod
def from_dict(cls, metric_info_dict: dict) -> "MetricInfo":
field_names = {f.name for f in dataclasses.fields(cls)}
return cls(**{k: v for k, v in metric_info_dict.items() if k in field_names})
| 0
|
hf_public_repos/datasets/src
|
hf_public_repos/datasets/src/datasets/table.py
|
import copy
import os
import warnings
from functools import partial
from itertools import groupby
from typing import TYPE_CHECKING, Callable, Iterator, List, Optional, Tuple, TypeVar, Union
import numpy as np
import pyarrow as pa
import pyarrow.compute as pc
from . import config
from .utils.logging import get_logger
if TYPE_CHECKING:
from .features.features import Features, FeatureType
logger = get_logger(__name__)
def inject_arrow_table_documentation(arrow_table_method):
def wrapper(fn):
fn.__doc__ = arrow_table_method.__doc__ + (fn.__doc__ if fn.__doc__ is not None else "")
fn.__doc__ = fn.__doc__.replace("pyarrow.Table", "Table")
if hasattr(arrow_table_method, "__annotations__"):
fn.__annotations__ = arrow_table_method.__annotations__
return fn
return wrapper
def _in_memory_arrow_table_from_file(filename: str) -> pa.Table:
in_memory_stream = pa.input_stream(filename)
opened_stream = pa.ipc.open_stream(in_memory_stream)
pa_table = opened_stream.read_all()
return pa_table
def _in_memory_arrow_table_from_buffer(buffer: pa.Buffer) -> pa.Table:
stream = pa.BufferReader(buffer)
opened_stream = pa.ipc.open_stream(stream)
table = opened_stream.read_all()
return table
def _memory_mapped_record_batch_reader_from_file(filename: str) -> pa.RecordBatchStreamReader:
memory_mapped_stream = pa.memory_map(filename)
return pa.ipc.open_stream(memory_mapped_stream)
def read_schema_from_file(filename: str) -> pa.Schema:
"""
Infer arrow table schema from file without loading whole file into memory.
Usefull especially while having very big files.
"""
with pa.memory_map(filename) as memory_mapped_stream:
schema = pa.ipc.open_stream(memory_mapped_stream).schema
return schema
def _memory_mapped_arrow_table_from_file(filename: str) -> pa.Table:
opened_stream = _memory_mapped_record_batch_reader_from_file(filename)
pa_table = opened_stream.read_all()
return pa_table
def _deepcopy(x, memo: dict):
"""deepcopy a regular class instance"""
cls = x.__class__
result = cls.__new__(cls)
memo[id(x)] = result
for k, v in x.__dict__.items():
setattr(result, k, copy.deepcopy(v, memo))
return result
def _interpolation_search(arr: List[int], x: int) -> int:
"""
Return the position i of a sorted array so that arr[i] <= x < arr[i+1]
Args:
arr (`List[int]`): non-empty sorted list of integers
x (`int`): query
Returns:
`int`: the position i so that arr[i] <= x < arr[i+1]
Raises:
`IndexError`: if the array is empty or if the query is outside the array values
"""
i, j = 0, len(arr) - 1
while i < j and arr[i] <= x < arr[j]:
k = i + ((j - i) * (x - arr[i]) // (arr[j] - arr[i]))
if arr[k] <= x < arr[k + 1]:
return k
elif arr[k] < x:
i, j = k + 1, j
else:
i, j = i, k
raise IndexError(f"Invalid query '{x}' for size {arr[-1] if len(arr) else 'none'}.")
class IndexedTableMixin:
def __init__(self, table: pa.Table):
self._schema: pa.Schema = table.schema
self._batches: List[pa.RecordBatch] = [
recordbatch for recordbatch in table.to_batches() if len(recordbatch) > 0
]
self._offsets: np.ndarray = np.cumsum([0] + [len(b) for b in self._batches], dtype=np.int64)
def fast_gather(self, indices: Union[List[int], np.ndarray]) -> pa.Table:
"""
Create a pa.Table by gathering the records at the records at the specified indices. Should be faster
than pa.concat_tables(table.fast_slice(int(i) % table.num_rows, 1) for i in indices) since NumPy can compute
the binary searches in parallel, highly optimized C
"""
if not len(indices):
raise ValueError("Indices must be non-empty")
batch_indices = np.searchsorted(self._offsets, indices, side="right") - 1
return pa.Table.from_batches(
[
self._batches[batch_idx].slice(i - self._offsets[batch_idx], 1)
for batch_idx, i in zip(batch_indices, indices)
],
schema=self._schema,
)
def fast_slice(self, offset=0, length=None) -> pa.Table:
"""
Slice the Table using interpolation search.
The behavior is the same as `pyarrow.Table.slice` but it's significantly faster.
Interpolation search is used to find the start and end indexes of the batches we want to keep.
The batches to keep are then concatenated to form the sliced Table.
"""
if offset < 0:
raise IndexError("Offset must be non-negative")
elif offset >= self._offsets[-1] or (length is not None and length <= 0):
return pa.Table.from_batches([], schema=self._schema)
i = _interpolation_search(self._offsets, offset)
if length is None or length + offset >= self._offsets[-1]:
batches = self._batches[i:]
batches[0] = batches[0].slice(offset - self._offsets[i])
else:
j = _interpolation_search(self._offsets, offset + length - 1)
batches = self._batches[i : j + 1]
batches[-1] = batches[-1].slice(0, offset + length - self._offsets[j])
batches[0] = batches[0].slice(offset - self._offsets[i])
return pa.Table.from_batches(batches, schema=self._schema)
class Table(IndexedTableMixin):
"""
Wraps a pyarrow Table by using composition.
This is the base class for `InMemoryTable`, `MemoryMappedTable` and `ConcatenationTable`.
It implements all the basic attributes/methods of the pyarrow Table class except
the Table transforms: `slice, filter, flatten, combine_chunks, cast, add_column,
append_column, remove_column, set_column, rename_columns` and `drop`.
The implementation of these methods differs for the subclasses.
"""
def __init__(self, table: pa.Table):
super().__init__(table)
self.table = table
def __deepcopy__(self, memo: dict):
# arrow tables are immutable, so there's no need to copy self.table
# moreover calling deepcopy on a pyarrow table seems to make pa.total_allocated_bytes() decrease for some reason
# by adding it to the memo, self.table won't be copied
memo[id(self.table)] = self.table
# same for the recordbatches used by the index
memo[id(self._batches)] = list(self._batches)
return _deepcopy(self, memo)
def validate(self, *args, **kwargs):
"""
Perform validation checks. An exception is raised if validation fails.
By default only cheap validation checks are run. Pass `full=True`
for thorough validation checks (potentially `O(n)`).
Args:
full (`bool`, defaults to `False`):
If `True`, run expensive checks, otherwise cheap checks only.
Raises:
`pa.lib.ArrowInvalid`: if validation fails
"""
return self.table.validate(*args, **kwargs)
def equals(self, *args, **kwargs):
"""
Check if contents of two tables are equal.
Args:
other ([`~datasets.table.Table`]):
Table to compare against.
check_metadata `bool`, defaults to `False`):
Whether schema metadata equality should be checked as well.
Returns:
`bool`
"""
args = tuple(arg.table if isinstance(arg, Table) else arg for arg in args)
kwargs = {k: v.table if isinstance(v, Table) else v for k, v in kwargs}
return self.table.equals(*args, **kwargs)
def to_batches(self, *args, **kwargs):
"""
Convert Table to list of (contiguous) `RecordBatch` objects.
Args:
max_chunksize (`int`, defaults to `None`):
Maximum size for `RecordBatch` chunks. Individual chunks may be
smaller depending on the chunk layout of individual columns.
Returns:
`List[pyarrow.RecordBatch]`
"""
return self.table.to_batches(*args, **kwargs)
def to_pydict(self, *args, **kwargs):
"""
Convert the Table to a `dict` or `OrderedDict`.
Returns:
`dict`
"""
return self.table.to_pydict(*args, **kwargs)
def to_pylist(self, *args, **kwargs):
"""
Convert the Table to a list
Returns:
`list`
"""
try:
return self.table.to_pylist(*args, **kwargs)
except AttributeError: # pyarrow <7 does not have to_pylist, so we use to_pydict
pydict = self.table.to_pydict(*args, **kwargs)
return [{k: pydict[k][i] for k in pydict} for i in range(len(self.table))]
def to_pandas(self, *args, **kwargs):
"""
Convert to a pandas-compatible NumPy array or DataFrame, as appropriate.
Args:
memory_pool (`MemoryPool`, defaults to `None`):
Arrow MemoryPool to use for allocations. Uses the default memory
pool is not passed.
strings_to_categorical (`bool`, defaults to `False`):
Encode string (UTF8) and binary types to `pandas.Categorical`.
categories (`list`, defaults to `empty`):
List of fields that should be returned as `pandas.Categorical`. Only
applies to table-like data structures.
zero_copy_only (`bool`, defaults to `False`):
Raise an `ArrowException` if this function call would require copying
the underlying data.
integer_object_nulls (`bool`, defaults to `False`):
Cast integers with nulls to objects.
date_as_object (`bool`, defaults to `True`):
Cast dates to objects. If `False`, convert to `datetime64[ns]` dtype.
timestamp_as_object (`bool`, defaults to `False`):
Cast non-nanosecond timestamps (`np.datetime64`) to objects. This is
useful if you have timestamps that don't fit in the normal date
range of nanosecond timestamps (1678 CE-2262 CE).
If `False`, all timestamps are converted to `datetime64[ns]` dtype.
use_threads (`bool`, defaults to `True`):
Whether to parallelize the conversion using multiple threads.
deduplicate_objects (`bool`, defaults to `False`):
Do not create multiple copies Python objects when created, to save
on memory use. Conversion will be slower.
ignore_metadata (`bool`, defaults to `False`):
If `True`, do not use the 'pandas' metadata to reconstruct the
DataFrame index, if present.
safe (`bool`, defaults to `True`):
For certain data types, a cast is needed in order to store the
data in a pandas DataFrame or Series (e.g. timestamps are always
stored as nanoseconds in pandas). This option controls whether it
is a safe cast or not.
split_blocks (`bool`, defaults to `False`):
If `True`, generate one internal "block" for each column when
creating a pandas.DataFrame from a `RecordBatch` or `Table`. While this
can temporarily reduce memory note that various pandas operations
can trigger "consolidation" which may balloon memory use.
self_destruct (`bool`, defaults to `False`):
EXPERIMENTAL: If `True`, attempt to deallocate the originating Arrow
memory while converting the Arrow object to pandas. If you use the
object after calling `to_pandas` with this option it will crash your
program.
types_mapper (`function`, defaults to `None`):
A function mapping a pyarrow DataType to a pandas `ExtensionDtype`.
This can be used to override the default pandas type for conversion
of built-in pyarrow types or in absence of `pandas_metadata` in the
Table schema. The function receives a pyarrow DataType and is
expected to return a pandas `ExtensionDtype` or `None` if the
default conversion should be used for that type. If you have
a dictionary mapping, you can pass `dict.get` as function.
Returns:
`pandas.Series` or `pandas.DataFrame`: `pandas.Series` or `pandas.DataFrame` depending on type of object
"""
return self.table.to_pandas(*args, **kwargs)
def to_string(self, *args, **kwargs):
return self.table.to_string(*args, **kwargs)
def to_reader(self, max_chunksize: Optional[int] = None):
"""
Convert the Table to a RecordBatchReader.
Note that this method is zero-copy, it merely exposes the same data under a different API.
Args:
max_chunksize (`int`, defaults to `None`)
Maximum size for RecordBatch chunks. Individual chunks may be smaller depending
on the chunk layout of individual columns.
Returns:
`pyarrow.RecordBatchReader`
"""
return self.table.to_reader(max_chunksize=max_chunksize)
def field(self, *args, **kwargs):
"""
Select a schema field by its column name or numeric index.
Args:
i (`Union[int, str]`):
The index or name of the field to retrieve.
Returns:
`pyarrow.Field`
"""
return self.table.field(*args, **kwargs)
def column(self, *args, **kwargs):
"""
Select a column by its column name, or numeric index.
Args:
i (`Union[int, str]`):
The index or name of the column to retrieve.
Returns:
`pyarrow.ChunkedArray`
"""
return self.table.column(*args, **kwargs)
def itercolumns(self, *args, **kwargs):
"""
Iterator over all columns in their numerical order.
Yields:
`pyarrow.ChunkedArray`
"""
return self.table.itercolumns(*args, **kwargs)
@property
def schema(self):
"""
Schema of the table and its columns.
Returns:
`pyarrow.Schema`
"""
return self.table.schema
@property
def columns(self):
"""
List of all columns in numerical order.
Returns:
`List[pa.ChunkedArray]`
"""
return self.table.columns
@property
def num_columns(self):
"""
Number of columns in this table.
Returns:
int
"""
return self.table.num_columns
@property
def num_rows(self):
"""
Number of rows in this table.
Due to the definition of a table, all columns have the same number of
rows.
Returns:
int
"""
return self.table.num_rows
@property
def shape(self):
"""
Dimensions of the table: (#rows, #columns).
Returns:
`(int, int)`: Number of rows and number of columns.
"""
return self.table.shape
@property
def nbytes(self):
"""
Total number of bytes consumed by the elements of the table.
"""
return self.table.nbytes
@property
def column_names(self):
"""
Names of the table's columns.
"""
return self.table.column_names
def __eq__(self, other):
return self.equals(other)
def __getitem__(self, i):
return self.table[i]
def __len__(self):
return len(self.table)
def __repr__(self):
return self.table.__repr__().replace("pyarrow.Table", self.__class__.__name__)
def __str__(self):
return self.table.__str__().replace("pyarrow.Table", self.__class__.__name__)
def slice(self, *args, **kwargs):
"""
Compute zero-copy slice of this Table.
Args:
offset (`int`, defaults to `0`):
Offset from start of table to slice.
length (`int`, defaults to `None`):
Length of slice (default is until end of table starting from
offset).
Returns:
`datasets.table.Table`
"""
raise NotImplementedError()
def filter(self, *args, **kwargs):
"""
Select records from a Table. See `pyarrow.compute.filter` for full usage.
"""
raise NotImplementedError()
def flatten(self, *args, **kwargs):
"""
Flatten this Table. Each column with a struct type is flattened
into one column per struct field. Other columns are left unchanged.
Args:
memory_pool (`MemoryPool`, defaults to `None`):
For memory allocations, if required, otherwise use default pool.
Returns:
`datasets.table.Table`
"""
raise NotImplementedError()
def combine_chunks(self, *args, **kwargs):
"""
Make a new table by combining the chunks this table has.
All the underlying chunks in the `ChunkedArray` of each column are
concatenated into zero or one chunk.
Args:
memory_pool (`MemoryPool`, defaults to `None`):
For memory allocations, if required, otherwise use default pool.
Returns:
`datasets.table.Table`
"""
raise NotImplementedError()
def cast(self, *args, **kwargs):
"""
Cast table values to another schema.
Args:
target_schema (`Schema`):
Schema to cast to, the names and order of fields must match.
safe (`bool`, defaults to `True`):
Check for overflows or other unsafe conversions.
Returns:
`datasets.table.Table`
"""
raise NotImplementedError()
def replace_schema_metadata(self, *args, **kwargs):
"""
EXPERIMENTAL: Create shallow copy of table by replacing schema
key-value metadata with the indicated new metadata (which may be None,
which deletes any existing metadata
Args:
metadata (`dict`, defaults to `None`):
Returns:
`datasets.table.Table`: shallow_copy
"""
raise NotImplementedError()
def add_column(self, *args, **kwargs):
"""
Add column to Table at position.
A new table is returned with the column added, the original table
object is left unchanged.
Args:
i (`int`):
Index to place the column at.
field_ (`Union[str, pyarrow.Field]`):
If a string is passed then the type is deduced from the column
data.
column (`Union[pyarrow.Array, List[pyarrow.Array]]`):
Column data.
Returns:
`datasets.table.Table`: New table with the passed column added.
"""
raise NotImplementedError()
def append_column(self, *args, **kwargs):
"""
Append column at end of columns.
Args:
field_ (`Union[str, pyarrow.Field]`):
If a string is passed then the type is deduced from the column
data.
column (`Union[pyarrow.Array, List[pyarrow.Array]]`):
Column data.
Returns:
`datasets.table.Table`: New table with the passed column added.
"""
raise NotImplementedError()
def remove_column(self, *args, **kwargs):
"""
Create new Table with the indicated column removed.
Args:
i (`int`):
Index of column to remove.
Returns:
`datasets.table.Table`: New table without the column.
"""
raise NotImplementedError()
def set_column(self, *args, **kwargs):
"""
Replace column in Table at position.
Args:
i (`int`):
Index to place the column at.
field_ (`Union[str, pyarrow.Field]`):
If a string is passed then the type is deduced from the column
data.
column (`Union[pyarrow.Array, List[pyarrow.Array]]`):
Column data.
Returns:
`datasets.table.Table`: New table with the passed column set.
"""
raise NotImplementedError()
def rename_columns(self, *args, **kwargs):
"""
Create new table with columns renamed to provided names.
"""
raise NotImplementedError()
def drop(self, *args, **kwargs):
"""
Drop one or more columns and return a new table.
Args:
columns (`List[str]`):
List of field names referencing existing columns.
Raises:
`KeyError` : if any of the passed columns name are not existing.
Returns:
`datasets.table.Table`: New table without the columns.
"""
raise NotImplementedError()
def select(self, *args, **kwargs):
"""
Select columns of the table.
Returns a new table with the specified columns, and metadata preserved.
Args:
columns (:obj:`Union[List[str], List[int]]`):
The column names or integer indices to select.
Returns:
`datasets.table.Table`: table with only a subset of the columns
"""
raise NotImplementedError()
class TableBlock(Table):
"""
`TableBlock` is the allowed class inside a `ConcanetationTable`.
Only `MemoryMappedTable` and `InMemoryTable` are `TableBlock`.
This is because we don't want a `ConcanetationTable` made out of other `ConcanetationTables`.
"""
pass
class InMemoryTable(TableBlock):
"""
The table is said in-memory when it is loaded into the user's RAM.
Pickling it does copy all the data using memory.
Its implementation is simple and uses the underlying pyarrow Table methods directly.
This is different from the `MemoryMapped` table, for which pickling doesn't copy all the
data in memory. For a `MemoryMapped`, unpickling instead reloads the table from the disk.
`InMemoryTable` must be used when data fit in memory, while `MemoryMapped` are reserved for
data bigger than memory or when you want the memory footprint of your application to
stay low.
"""
@classmethod
def from_file(cls, filename: str):
table = _in_memory_arrow_table_from_file(filename)
return cls(table)
@classmethod
def from_buffer(cls, buffer: pa.Buffer):
table = _in_memory_arrow_table_from_buffer(buffer)
return cls(table)
@classmethod
def from_pandas(cls, *args, **kwargs):
"""
Convert pandas.DataFrame to an Arrow Table.
The column types in the resulting Arrow Table are inferred from the
dtypes of the pandas.Series in the DataFrame. In the case of non-object
Series, the NumPy dtype is translated to its Arrow equivalent. In the
case of `object`, we need to guess the datatype by looking at the
Python objects in this Series.
Be aware that Series of the `object` dtype don't carry enough
information to always lead to a meaningful Arrow type. In the case that
we cannot infer a type, e.g. because the DataFrame is of length 0 or
the Series only contains `None/nan` objects, the type is set to
null. This behavior can be avoided by constructing an explicit schema
and passing it to this function.
Args:
df (`pandas.DataFrame`):
schema (`pyarrow.Schema`, *optional*):
The expected schema of the Arrow Table. This can be used to
indicate the type of columns if we cannot infer it automatically.
If passed, the output will have exactly this schema. Columns
specified in the schema that are not found in the DataFrame columns
or its index will raise an error. Additional columns or index
levels in the DataFrame which are not specified in the schema will
be ignored.
preserve_index (`bool`, *optional*):
Whether to store the index as an additional column in the resulting
`Table`. The default of None will store the index as a column,
except for RangeIndex which is stored as metadata only. Use
`preserve_index=True` to force it to be stored as a column.
nthreads (`int`, defaults to `None` (may use up to system CPU count threads))
If greater than 1, convert columns to Arrow in parallel using
indicated number of threads.
columns (`List[str]`, *optional*):
List of column to be converted. If `None`, use all columns.
safe (`bool`, defaults to `True`):
Check for overflows or other unsafe conversions,
Returns:
`datasets.table.Table`:
Examples:
```python
>>> import pandas as pd
>>> import pyarrow as pa
>>> df = pd.DataFrame({
... 'int': [1, 2],
... 'str': ['a', 'b']
... })
>>> pa.Table.from_pandas(df)
<pyarrow.lib.Table object at 0x7f05d1fb1b40>
```
"""
return cls(pa.Table.from_pandas(*args, **kwargs))
@classmethod
def from_arrays(cls, *args, **kwargs):
"""
Construct a Table from Arrow arrays.
Args:
arrays (`List[Union[pyarrow.Array, pyarrow.ChunkedArray]]`):
Equal-length arrays that should form the table.
names (`List[str]`, *optional*):
Names for the table columns. If not passed, schema must be passed.
schema (`Schema`, defaults to `None`):
Schema for the created table. If not passed, names must be passed.
metadata (`Union[dict, Mapping]`, defaults to `None`):
Optional metadata for the schema (if inferred).
Returns:
`datasets.table.Table`
"""
return cls(pa.Table.from_arrays(*args, **kwargs))
@classmethod
def from_pydict(cls, *args, **kwargs):
"""
Construct a Table from Arrow arrays or columns.
Args:
mapping (`Union[dict, Mapping]`):
A mapping of strings to Arrays or Python lists.
schema (`Schema`, defaults to `None`):
If not passed, will be inferred from the Mapping values
metadata (`Union[dict, Mapping]`, defaults to `None`):
Optional metadata for the schema (if inferred).
Returns:
`datasets.table.Table`
"""
return cls(pa.Table.from_pydict(*args, **kwargs))
@classmethod
def from_pylist(cls, mapping, *args, **kwargs):
"""
Construct a Table from list of rows / dictionaries.
Args:
mapping (`List[dict]`):
A mapping of strings to row values.
schema (`Schema`, defaults to `None`):
If not passed, will be inferred from the Mapping values
metadata (`Union[dict, Mapping]`, defaults to `None`):
Optional metadata for the schema (if inferred).
Returns:
`datasets.table.Table`
"""
return cls(pa.Table.from_pylist(mapping, *args, **kwargs))
@classmethod
def from_batches(cls, *args, **kwargs):
"""
Construct a Table from a sequence or iterator of Arrow `RecordBatches`.
Args:
batches (`Union[Sequence[pyarrow.RecordBatch], Iterator[pyarrow.RecordBatch]]`):
Sequence of `RecordBatch` to be converted, all schemas must be equal.
schema (`Schema`, defaults to `None`):
If not passed, will be inferred from the first `RecordBatch`.
Returns:
`datasets.table.Table`:
"""
return cls(pa.Table.from_batches(*args, **kwargs))
def slice(self, offset=0, length=None):
"""
Compute zero-copy slice of this Table.
Args:
offset (`int`, defaults to `0`):
Offset from start of table to slice.
length (`int`, defaults to `None`):
Length of slice (default is until end of table starting from
offset).
Returns:
`datasets.table.Table`
"""
# Use fast slicing here
return InMemoryTable(self.fast_slice(offset=offset, length=length))
def filter(self, *args, **kwargs):
"""
Select records from a Table. See `pyarrow.compute.filter` for full usage.
"""
return InMemoryTable(self.table.filter(*args, **kwargs))
def flatten(self, *args, **kwargs):
"""
Flatten this Table. Each column with a struct type is flattened
into one column per struct field. Other columns are left unchanged.
Args:
memory_pool (`MemoryPool`, defaults to `None`):
For memory allocations, if required, otherwise use default pool.
Returns:
`datasets.table.Table`
"""
return InMemoryTable(table_flatten(self.table, *args, **kwargs))
def combine_chunks(self, *args, **kwargs):
"""
Make a new table by combining the chunks this table has.
All the underlying chunks in the `ChunkedArray` of each column are
concatenated into zero or one chunk.
Args:
memory_pool (`MemoryPool`, defaults to `None`):
For memory allocations, if required, otherwise use default pool.
Returns:
`datasets.table.Table`
"""
return InMemoryTable(self.table.combine_chunks(*args, **kwargs))
def cast(self, *args, **kwargs):
"""
Cast table values to another schema.
Args:
target_schema (`Schema`):
Schema to cast to, the names and order of fields must match.
safe (`bool`, defaults to `True`):
Check for overflows or other unsafe conversions.
Returns:
`datasets.table.Table`
"""
return InMemoryTable(table_cast(self.table, *args, **kwargs))
def replace_schema_metadata(self, *args, **kwargs):
"""
EXPERIMENTAL: Create shallow copy of table by replacing schema
key-value metadata with the indicated new metadata (which may be `None`,
which deletes any existing metadata).
Args:
metadata (`dict`, defaults to `None`):
Returns:
`datasets.table.Table`: shallow_copy
"""
return InMemoryTable(self.table.replace_schema_metadata(*args, **kwargs))
def add_column(self, *args, **kwargs):
"""
Add column to Table at position.
A new table is returned with the column added, the original table
object is left unchanged.
Args:
i (`int`):
Index to place the column at.
field_ (`Union[str, pyarrow.Field]`):
If a string is passed then the type is deduced from the column
data.
column (`Union[pyarrow.Array, List[pyarrow.Array]]`):
Column data.
Returns:
`datasets.table.Table`: New table with the passed column added.
"""
return InMemoryTable(self.table.add_column(*args, **kwargs))
def append_column(self, *args, **kwargs):
"""
Append column at end of columns.
Args:
field_ (`Union[str, pyarrow.Field]`):
If a string is passed then the type is deduced from the column
data.
column (`Union[pyarrow.Array, List[pyarrow.Array]]`):
Column data.
Returns:
`datasets.table.Table`:
New table with the passed column added.
"""
return InMemoryTable(self.table.append_column(*args, **kwargs))
def remove_column(self, *args, **kwargs):
"""
Create new Table with the indicated column removed.
Args:
i (`int`):
Index of column to remove.
Returns:
`datasets.table.Table`:
New table without the column.
"""
return InMemoryTable(self.table.remove_column(*args, **kwargs))
def set_column(self, *args, **kwargs):
"""
Replace column in Table at position.
Args:
i (`int`):
Index to place the column at.
field_ (`Union[str, pyarrow.Field]`):
If a string is passed then the type is deduced from the column
data.
column (`Union[pyarrow.Array, List[pyarrow.Array]]`):
Column data.
Returns:
`datasets.table.Table`:
New table with the passed column set.
"""
return InMemoryTable(self.table.set_column(*args, **kwargs))
def rename_columns(self, *args, **kwargs):
"""
Create new table with columns renamed to provided names.
"""
return InMemoryTable(self.table.rename_columns(*args, **kwargs))
def drop(self, *args, **kwargs):
"""
Drop one or more columns and return a new table.
Args:
columns (`List[str]`):
List of field names referencing existing columns.
Raises:
`KeyError` : if any of the passed columns name are not existing.
Returns:
`datasets.table.Table`:
New table without the columns.
"""
return InMemoryTable(self.table.drop(*args, **kwargs))
def select(self, *args, **kwargs):
"""
Select columns of the table.
Returns a new table with the specified columns, and metadata preserved.
Args:
columns (:obj:`Union[List[str], List[int]]`):
The column names or integer indices to select.
Returns:
:class:`datasets.table.Table`: New table with the specified columns, and metadata preserved.
"""
return InMemoryTable(self.table.select(*args, **kwargs))
# The MemoryMappedTable needs replays to properly reload tables from the disk
Replay = Tuple[str, tuple, dict]
class MemoryMappedTable(TableBlock):
"""
The table is said memory mapped when it doesn't use the user's RAM but loads the data
from the disk instead.
Pickling it doesn't copy the data into memory.
Instead, only the path to the memory mapped arrow file is pickled, as well as the list
of transforms to "replay" when reloading the table from the disk.
Its implementation requires to store an history of all the transforms that were applied
to the underlying pyarrow Table, so that they can be "replayed" when reloading the Table
from the disk.
This is different from the `InMemoryTable` table, for which pickling does copy all the
data in memory.
`InMemoryTable` must be used when data fit in memory, while `MemoryMapped` are reserved for
data bigger than memory or when you want the memory footprint of your application to
stay low.
"""
def __init__(self, table: pa.Table, path: str, replays: Optional[List[Replay]] = None):
super().__init__(table)
self.path = os.path.abspath(path)
self.replays: List[Replay] = replays if replays is not None else []
@classmethod
def from_file(cls, filename: str, replays=None):
table = _memory_mapped_arrow_table_from_file(filename)
table = cls._apply_replays(table, replays)
return cls(table, filename, replays)
def __getstate__(self):
return {"path": self.path, "replays": self.replays}
def __setstate__(self, state):
path = state["path"]
replays = state["replays"]
table = _memory_mapped_arrow_table_from_file(path)
table = self._apply_replays(table, replays)
MemoryMappedTable.__init__(self, table, path=path, replays=replays)
@staticmethod
def _apply_replays(table: pa.Table, replays: Optional[List[Replay]] = None) -> pa.Table:
if replays is not None:
for name, args, kwargs in replays:
if name == "cast":
table = table_cast(table, *args, **kwargs)
elif name == "flatten":
table = table_flatten(table, *args, **kwargs)
else:
table = getattr(table, name)(*args, **kwargs)
return table
def _append_replay(self, replay: Replay) -> List[Replay]:
replays = copy.deepcopy(self.replays)
replays.append(replay)
return replays
def slice(self, offset=0, length=None):
"""
Compute zero-copy slice of this Table.
Args:
offset (`int`, defaults to `0`):
Offset from start of table to slice.
length (`int`, defaults to `None`):
Length of slice (default is until end of table starting from
offset).
Returns:
`datasets.table.Table`
"""
replay = ("slice", (offset, length), {})
replays = self._append_replay(replay)
# Use fast slicing here
return MemoryMappedTable(self.fast_slice(offset=offset, length=length), self.path, replays)
def filter(self, *args, **kwargs):
"""
Select records from a Table. See `pyarrow.compute.filter` for full usage.
"""
replay = ("filter", copy.deepcopy(args), copy.deepcopy(kwargs))
replays = self._append_replay(replay)
return MemoryMappedTable(self.table.filter(*args, **kwargs), self.path, replays)
def flatten(self, *args, **kwargs):
"""
Flatten this Table. Each column with a struct type is flattened
into one column per struct field. Other columns are left unchanged.
Args:
memory_pool (`MemoryPool`, defaults to `None`):
For memory allocations, if required, otherwise use default pool.
Returns:
`datasets.table.Table`
"""
replay = ("flatten", copy.deepcopy(args), copy.deepcopy(kwargs))
replays = self._append_replay(replay)
return MemoryMappedTable(table_flatten(self.table, *args, **kwargs), self.path, replays)
def combine_chunks(self, *args, **kwargs):
"""
Make a new table by combining the chunks this table has.
All the underlying chunks in the ChunkedArray of each column are
concatenated into zero or one chunk.
Args:
memory_pool (`MemoryPool`, defaults to `None`):
For memory allocations, if required, otherwise use default pool.
Returns:
`datasets.table.Table`
"""
replay = ("combine_chunks", copy.deepcopy(args), copy.deepcopy(kwargs))
replays = self._append_replay(replay)
return MemoryMappedTable(self.table.combine_chunks(*args, **kwargs), self.path, replays)
def cast(self, *args, **kwargs):
"""
Cast table values to another schema
Args:
target_schema (`Schema`):
Schema to cast to, the names and order of fields must match.
safe (`bool`, defaults to `True`):
Check for overflows or other unsafe conversions.
Returns:
`datasets.table.Table`
"""
replay = ("cast", copy.deepcopy(args), copy.deepcopy(kwargs))
replays = self._append_replay(replay)
return MemoryMappedTable(table_cast(self.table, *args, **kwargs), self.path, replays)
def replace_schema_metadata(self, *args, **kwargs):
"""
EXPERIMENTAL: Create shallow copy of table by replacing schema
key-value metadata with the indicated new metadata (which may be None,
which deletes any existing metadata.
Args:
metadata (`dict`, defaults to `None`):
Returns:
`datasets.table.Table`: shallow_copy
"""
replay = ("replace_schema_metadata", copy.deepcopy(args), copy.deepcopy(kwargs))
replays = self._append_replay(replay)
return MemoryMappedTable(self.table.replace_schema_metadata(*args, **kwargs), self.path, replays)
def add_column(self, *args, **kwargs):
"""
Add column to Table at position.
A new table is returned with the column added, the original table
object is left unchanged.
Args:
i (`int`):
Index to place the column at.
field_ (`Union[str, pyarrow.Field]`):
If a string is passed then the type is deduced from the column
data.
column (`Union[pyarrow.Array, List[pyarrow.Array]]`):
Column data.
Returns:
`datasets.table.Table`: New table with the passed column added.
"""
replay = ("add_column", copy.deepcopy(args), copy.deepcopy(kwargs))
replays = self._append_replay(replay)
return MemoryMappedTable(self.table.add_column(*args, **kwargs), self.path, replays)
def append_column(self, *args, **kwargs):
"""
Append column at end of columns.
Args:
field_ (`Union[str, pyarrow.Field]`):
If a string is passed then the type is deduced from the column
data.
column (`Union[pyarrow.Array, List[pyarrow.Array]]`):
Column data.
Returns:
`datasets.table.Table`:
New table with the passed column added.
"""
replay = ("append_column", copy.deepcopy(args), copy.deepcopy(kwargs))
replays = self._append_replay(replay)
return MemoryMappedTable(self.table.append_column(*args, **kwargs), self.path, replays)
def remove_column(self, *args, **kwargs):
"""
Create new Table with the indicated column removed.
Args:
i (`int`):
Index of column to remove.
Returns:
`datasets.table.Table`:
New table without the column.
"""
replay = ("remove_column", copy.deepcopy(args), copy.deepcopy(kwargs))
replays = self._append_replay(replay)
return MemoryMappedTable(self.table.remove_column(*args, **kwargs), self.path, replays)
def set_column(self, *args, **kwargs):
"""
Replace column in Table at position.
Args:
i (`int`):
Index to place the column at.
field_ (`Union[str, pyarrow.Field]`):
If a string is passed then the type is deduced from the column
data.
column (`Union[pyarrow.Array, List[pyarrow.Array]]`):
Column data.
Returns:
`datasets.table.Table`:
New table with the passed column set.
"""
replay = ("set_column", copy.deepcopy(args), copy.deepcopy(kwargs))
replays = self._append_replay(replay)
return MemoryMappedTable(self.table.set_column(*args, **kwargs), self.path, replays)
def rename_columns(self, *args, **kwargs):
"""
Create new table with columns renamed to provided names.
"""
replay = ("rename_columns", copy.deepcopy(args), copy.deepcopy(kwargs))
replays = self._append_replay(replay)
return MemoryMappedTable(self.table.rename_columns(*args, **kwargs), self.path, replays)
def drop(self, *args, **kwargs):
"""
Drop one or more columns and return a new table.
Args:
columns (`List[str]`):
List of field names referencing existing columns.
Raises:
`KeyError` : if any of the passed columns name are not existing.
Returns:
`datasets.table.Table`:
New table without the columns.
"""
replay = ("drop", copy.deepcopy(args), copy.deepcopy(kwargs))
replays = self._append_replay(replay)
return MemoryMappedTable(self.table.drop(*args, **kwargs), self.path, replays)
def select(self, *args, **kwargs):
"""
Select columns of the table.
Returns a new table with the specified columns, and metadata preserved.
Args:
columns (:obj:`Union[List[str], List[int]]`):
The column names or integer indices to select.
Returns:
:class:`datasets.table.Table`: New table with the specified columns, and metadata preserved.
"""
replay = ("select", copy.deepcopy(args), copy.deepcopy(kwargs))
replays = self._append_replay(replay)
return MemoryMappedTable(self.table.select(*args, **kwargs), self.path, replays)
# A ConcatenationTable is the concatenation of several tables.
# The ``blocks`` attributes stores a list of list of blocks.
# The first axis concatenates the tables along the axis 0 (it appends rows),
# while the second axis concatenates tables along the axis 1 (it appends columns).
TableBlockContainer = TypeVar("TableBlockContainer", TableBlock, List[TableBlock], List[List[TableBlock]])
class ConcatenationTable(Table):
"""
The table comes from the concatenation of several tables called blocks.
It enables concatenation on both axis 0 (append rows) and axis 1 (append columns).
The underlying tables are called "blocks" and can be either `InMemoryTable`
or `MemoryMappedTable` objects.
This allows to combine tables that come from memory or that are memory mapped.
When a `ConcatenationTable` is pickled, then each block is pickled:
- the `InMemoryTable` objects are pickled by copying all the data in memory.
- the MemoryMappedTable objects are pickled without copying the data into memory.
Instead, only the path to the memory mapped arrow file is pickled, as well as the list
of transforms to "replays" when reloading the table from the disk.
Its implementation requires to store each block separately.
The `blocks` attributes stores a list of list of blocks.
The first axis concatenates the tables along the axis 0 (it appends rows),
while the second axis concatenates tables along the axis 1 (it appends columns).
If some columns are missing when concatenating on axis 0, they are filled with null values.
This is done using `pyarrow.concat_tables(tables, promote=True)`.
You can access the fully combined table by accessing the `ConcatenationTable.table` attribute,
and the blocks by accessing the `ConcatenationTable.blocks` attribute.
"""
def __init__(self, table: pa.Table, blocks: List[List[TableBlock]]):
super().__init__(table)
self.blocks = blocks
# Check that all the blocks have the right type.
# Only InMemoryTable and MemoryMappedTable are allowed.
for subtables in blocks:
for subtable in subtables:
if not isinstance(subtable, TableBlock):
raise TypeError(
"The blocks of a ConcatenationTable must be InMemoryTable or MemoryMappedTable objects"
f", but got {subtable}."
)
def __getstate__(self):
return {"blocks": self.blocks}
def __setstate__(self, state):
blocks = state["blocks"]
table = self._concat_blocks_horizontally_and_vertically(blocks)
ConcatenationTable.__init__(self, table, blocks=blocks)
@staticmethod
def _concat_blocks(blocks: List[Union[TableBlock, pa.Table]], axis: int = 0) -> pa.Table:
pa_tables = [table.table if hasattr(table, "table") else table for table in blocks]
if axis == 0:
# we set promote=True to fill missing columns with null values
if config.PYARROW_VERSION.major < 14:
return pa.concat_tables(pa_tables, promote=True)
else:
return pa.concat_tables(pa_tables, promote_options="default")
elif axis == 1:
for i, table in enumerate(pa_tables):
if i == 0:
pa_table = table
else:
for name, col in zip(table.column_names, table.columns):
pa_table = pa_table.append_column(name, col)
return pa_table
else:
raise ValueError("'axis' must be either 0 or 1")
@classmethod
def _concat_blocks_horizontally_and_vertically(cls, blocks: List[List[TableBlock]]) -> pa.Table:
pa_tables_to_concat_vertically = []
for i, tables in enumerate(blocks):
if not tables:
continue
pa_table_horizontally_concatenated = cls._concat_blocks(tables, axis=1)
pa_tables_to_concat_vertically.append(pa_table_horizontally_concatenated)
return cls._concat_blocks(pa_tables_to_concat_vertically, axis=0)
@classmethod
def _merge_blocks(cls, blocks: TableBlockContainer, axis: Optional[int] = None) -> TableBlockContainer:
if axis is not None:
merged_blocks = []
for is_in_memory, block_group in groupby(blocks, key=lambda x: isinstance(x, InMemoryTable)):
if is_in_memory:
block_group = [InMemoryTable(cls._concat_blocks(list(block_group), axis=axis))]
merged_blocks += list(block_group)
else: # both
merged_blocks = [cls._merge_blocks(row_block, axis=1) for row_block in blocks]
if all(len(row_block) == 1 for row_block in merged_blocks):
merged_blocks = cls._merge_blocks(
[block for row_block in merged_blocks for block in row_block], axis=0
)
return merged_blocks
@classmethod
def _consolidate_blocks(cls, blocks: TableBlockContainer) -> TableBlockContainer:
if isinstance(blocks, TableBlock):
return blocks
elif isinstance(blocks[0], TableBlock):
return cls._merge_blocks(blocks, axis=0)
else:
return cls._merge_blocks(blocks)
@classmethod
def from_blocks(cls, blocks: TableBlockContainer) -> "ConcatenationTable":
blocks = cls._consolidate_blocks(blocks)
if isinstance(blocks, TableBlock):
table = blocks
return cls(table.table, [[table]])
elif isinstance(blocks[0], TableBlock):
table = cls._concat_blocks(blocks, axis=0)
blocks = [[t] for t in blocks]
return cls(table, blocks)
else:
table = cls._concat_blocks_horizontally_and_vertically(blocks)
return cls(table, blocks)
@classmethod
def from_tables(cls, tables: List[Union[pa.Table, Table]], axis: int = 0) -> "ConcatenationTable":
"""Create `ConcatenationTable` from list of tables.
Args:
tables (list of `Table` or list of `pyarrow.Table`):
List of tables.
axis (`{0, 1}`, defaults to `0`, meaning over rows):
Axis to concatenate over, where `0` means over rows (vertically) and `1` means over columns
(horizontally).
<Added version="1.6.0"/>
"""
def to_blocks(table: Union[pa.Table, Table]) -> List[List[TableBlock]]:
if isinstance(table, pa.Table):
return [[InMemoryTable(table)]]
elif isinstance(table, ConcatenationTable):
return copy.deepcopy(table.blocks)
else:
return [[table]]
def _slice_row_block(row_block: List[TableBlock], length: int) -> Tuple[List[TableBlock], List[TableBlock]]:
sliced = [table.slice(0, length) for table in row_block]
remainder = [table.slice(length, len(row_block[0]) - length) for table in row_block]
return sliced, remainder
def _split_both_like(
result: List[List[TableBlock]], blocks: List[List[TableBlock]]
) -> Tuple[List[List[TableBlock]], List[List[TableBlock]]]:
"""
Make sure each row_block contain the same num_rows to be able to concatenate them on axis=1.
To do so, we modify both blocks sets to have the same row_blocks boundaries.
For example, if `result` has 2 row_blocks of 3 rows and `blocks` has 3 row_blocks of 2 rows,
we modify both to have 4 row_blocks of size 2, 1, 1 and 2:
[ x x x | x x x ]
+ [ y y | y y | y y ]
-----------------------------
= [ x x | x | x | x x ]
[ y y | y | y | y y ]
"""
result, blocks = list(result), list(blocks)
new_result, new_blocks = [], []
while result and blocks:
# we slice the longest row block to save two row blocks of same length
# and we replace the long row block by its remainder if necessary
if len(result[0][0]) > len(blocks[0][0]):
new_blocks.append(blocks[0])
sliced, result[0] = _slice_row_block(result[0], len(blocks.pop(0)[0]))
new_result.append(sliced)
elif len(result[0][0]) < len(blocks[0][0]):
new_result.append(result[0])
sliced, blocks[0] = _slice_row_block(blocks[0], len(result.pop(0)[0]))
new_blocks.append(sliced)
else:
new_result.append(result.pop(0))
new_blocks.append(blocks.pop(0))
if result or blocks:
raise ValueError("Failed to concatenate on axis=1 because tables don't have the same number of rows")
return new_result, new_blocks
def _extend_blocks(
result: List[List[TableBlock]], blocks: List[List[TableBlock]], axis: int = 0
) -> List[List[TableBlock]]:
if axis == 0:
result.extend(blocks)
elif axis == 1:
# We make sure each row_block have the same num_rows
result, blocks = _split_both_like(result, blocks)
for i, row_block in enumerate(blocks):
result[i].extend(row_block)
return result
blocks = to_blocks(tables[0])
for table in tables[1:]:
table_blocks = to_blocks(table)
blocks = _extend_blocks(blocks, table_blocks, axis=axis)
return cls.from_blocks(blocks)
@property
def _slices(self):
offset = 0
for tables in self.blocks:
length = len(tables[0])
yield (offset, length)
offset += length
def slice(self, offset=0, length=None):
"""
Compute zero-copy slice of this Table.
Args:
offset (`int`, defaults to `0`):
Offset from start of table to slice.
length (`int`, defaults to `None`):
Length of slice (default is until end of table starting from
offset).
Returns:
`datasets.table.Table`
"""
table = self.table.slice(offset, length=length)
length = length if length is not None else self.num_rows - offset
blocks = []
for tables in self.blocks:
n_rows = len(tables[0])
if length == 0:
break
elif n_rows <= offset:
offset = offset - n_rows
elif n_rows <= offset + length:
blocks.append([t.slice(offset) for t in tables])
length, offset = length + offset - n_rows, 0
else:
blocks.append([t.slice(offset, length) for t in tables])
length, offset = 0, 0
return ConcatenationTable(table, blocks)
def filter(self, mask, *args, **kwargs):
"""
Select records from a Table. See `pyarrow.compute.filter` for full usage.
"""
table = self.table.filter(mask, *args, **kwargs)
blocks = []
for (offset, length), tables in zip(self._slices, self.blocks):
submask = mask.slice(offset, length)
blocks.append([t.filter(submask, *args, **kwargs) for t in tables])
return ConcatenationTable(table, blocks)
def flatten(self, *args, **kwargs):
"""
Flatten this Table. Each column with a struct type is flattened
into one column per struct field. Other columns are left unchanged.
Args:
memory_pool (`MemoryPool`, defaults to `None`):
For memory allocations, if required, otherwise use default pool.
Returns:
`datasets.table.Table`
"""
table = table_flatten(self.table, *args, **kwargs)
blocks = []
for tables in self.blocks:
blocks.append([t.flatten(*args, **kwargs) for t in tables])
return ConcatenationTable(table, blocks)
def combine_chunks(self, *args, **kwargs):
"""
Make a new table by combining the chunks this table has.
All the underlying chunks in the `ChunkedArray` of each column are
concatenated into zero or one chunk.
Args:
memory_pool (`MemoryPool`, defaults to `None`):
For memory allocations, if required, otherwise use default pool.
Returns:
`datasets.table.Table`
"""
table = self.table.combine_chunks(*args, **kwargs)
blocks = []
for tables in self.blocks:
blocks.append([t.combine_chunks(*args, **kwargs) for t in tables])
return ConcatenationTable(table, blocks)
def cast(self, target_schema, *args, **kwargs):
"""
Cast table values to another schema.
Args:
target_schema (`Schema`):
Schema to cast to, the names and order of fields must match.
safe (`bool`, defaults to `True`):
Check for overflows or other unsafe conversions.
Returns:
`datasets.table.Table`
"""
from .features import Features
table = table_cast(self.table, target_schema, *args, **kwargs)
target_features = Features.from_arrow_schema(target_schema)
blocks = []
for subtables in self.blocks:
new_tables = []
fields = list(target_schema)
for subtable in subtables:
subfields = []
for name in subtable.column_names:
subfields.append(fields.pop(next(i for i, field in enumerate(fields) if field.name == name)))
subfeatures = Features({subfield.name: target_features[subfield.name] for subfield in subfields})
subschema = subfeatures.arrow_schema
new_tables.append(subtable.cast(subschema, *args, **kwargs))
blocks.append(new_tables)
return ConcatenationTable(table, blocks)
def replace_schema_metadata(self, *args, **kwargs):
"""
EXPERIMENTAL: Create shallow copy of table by replacing schema
key-value metadata with the indicated new metadata (which may be `None`,
which deletes any existing metadata).
Args:
metadata (`dict`, defaults to `None`):
Returns:
`datasets.table.Table`: shallow_copy
"""
table = self.table.replace_schema_metadata(*args, **kwargs)
blocks = []
for tables in self.blocks:
blocks.append([t.replace_schema_metadata(*args, **kwargs) for t in tables])
return ConcatenationTable(table, self.blocks)
def add_column(self, *args, **kwargs):
"""
Add column to Table at position.
A new table is returned with the column added, the original table
object is left unchanged.
Args:
i (`int`):
Index to place the column at.
field_ (`Union[str, pyarrow.Field]`):
If a string is passed then the type is deduced from the column
data.
column (`Union[pyarrow.Array, List[pyarrow.Array]]`):
Column data.
Returns:
`datasets.table.Table`: New table with the passed column added.
"""
raise NotImplementedError()
def append_column(self, *args, **kwargs):
"""
Append column at end of columns.
Args:
field_ (`Union[str, pyarrow.Field]`):
If a string is passed then the type is deduced from the column
data.
column (`Union[pyarrow.Array, List[pyarrow.Array]]`):
Column data.
Returns:
`datasets.table.Table`:
New table with the passed column added.
"""
raise NotImplementedError()
def remove_column(self, i, *args, **kwargs):
"""
Create new Table with the indicated column removed.
Args:
i (`int`):
Index of column to remove.
Returns:
`datasets.table.Table`:
New table without the column.
"""
table = self.table.remove_column(i, *args, **kwargs)
name = self.table.column_names[i]
blocks = []
for tables in self.blocks:
blocks.append(
[
t.remove_column(t.column_names.index(name), *args, **kwargs) if name in t.column_names else t
for t in tables
]
)
return ConcatenationTable(table, blocks)
def set_column(self, *args, **kwargs):
"""
Replace column in Table at position.
Args:
i (`int`):
Index to place the column at.
field_ (`Union[str, pyarrow.Field]`):
If a string is passed then the type is deduced from the column
data.
column (`Union[pyarrow.Array, List[pyarrow.Array]]`):
Column data.
Returns:
`datasets.table.Table`:
New table with the passed column set.
"""
raise NotImplementedError()
def rename_columns(self, names, *args, **kwargs):
"""
Create new table with columns renamed to provided names.
"""
table = self.table.rename_columns(names, *args, **kwargs)
names = dict(zip(self.table.column_names, names))
blocks = []
for tables in self.blocks:
blocks.append(
[t.rename_columns([names[name] for name in t.column_names], *args, **kwargs) for t in tables]
)
return ConcatenationTable(table, blocks)
def drop(self, columns, *args, **kwargs):
"""
Drop one or more columns and return a new table.
Args:
columns (`List[str]`):
List of field names referencing existing columns.
Raises:
`KeyError` : if any of the passed columns name are not existing.
Returns:
`datasets.table.Table`:
New table without the columns.
"""
table = self.table.drop(columns, *args, **kwargs)
blocks = []
for tables in self.blocks:
blocks.append([t.drop([c for c in columns if c in t.column_names], *args, **kwargs) for t in tables])
return ConcatenationTable(table, blocks)
def select(self, columns, *args, **kwargs):
"""
Select columns of the table.
Returns a new table with the specified columns, and metadata preserved.
Args:
columns (:obj:`Union[List[str], List[int]]`):
The column names or integer indices to select.
Returns:
:class:`datasets.table.Table`: New table with the specified columns, and metadata preserved.
"""
table = self.table.select(columns, *args, **kwargs)
blocks = []
for tables in self.blocks:
blocks.append([t.select([c for c in columns if c in t.column_names], *args, **kwargs) for t in tables])
return ConcatenationTable(table, blocks)
def concat_tables(tables: List[Table], axis: int = 0) -> Table:
"""
Concatenate tables.
Args:
tables (list of `Table`):
List of tables to be concatenated.
axis (`{0, 1}`, defaults to `0`, meaning over rows):
Axis to concatenate over, where `0` means over rows (vertically) and `1` means over columns
(horizontally).
<Added version="1.6.0"/>
Returns:
`datasets.table.Table`:
If the number of input tables is > 1, then the returned table is a `datasets.table.ConcatenationTable`.
Otherwise if there's only one table, it is returned as is.
"""
tables = list(tables)
if len(tables) == 1:
return tables[0]
return ConcatenationTable.from_tables(tables, axis=axis)
def list_table_cache_files(table: Table) -> List[str]:
"""
Get the cache files that are loaded by the table.
Cache file are used when parts of the table come from the disk via memory mapping.
Returns:
`List[str]`:
A list of paths to the cache files loaded by the table.
"""
if isinstance(table, ConcatenationTable):
cache_files = []
for subtables in table.blocks:
for subtable in subtables:
cache_files += list_table_cache_files(subtable)
return cache_files
elif isinstance(table, MemoryMappedTable):
return [table.path]
else:
return []
def _wrap_for_chunked_arrays(func):
"""Apply the function on each chunk of a `pyarrow.ChunkedArray`, or on the array directly"""
def wrapper(array, *args, **kwargs):
if isinstance(array, pa.ChunkedArray):
return pa.chunked_array([func(chunk, *args, **kwargs) for chunk in array.chunks])
else:
return func(array, *args, **kwargs)
return wrapper
def _is_extension_type(pa_type: pa.DataType) -> bool:
"""
Check (recursively) if a pyarrow type is an extension type.
"""
if isinstance(pa_type, pa.StructType):
return any(_is_extension_type(field.type) for field in pa_type)
elif isinstance(pa_type, (pa.ListType, pa.FixedSizeListType, pa.LargeListType)):
return _is_extension_type(pa_type.value_type)
elif isinstance(pa_type, pa.ExtensionType):
return True
else:
return False
def array_concat(arrays: List[pa.Array]):
"""Improved version of pa.concat_arrays
It supports concatenating pa.ExtensionArray objects by concatenating the underlying storages.
Args:
arrays (List[pa.Array]): List of arrays to contatenate
Raises:
pa.ArrowInvalid: if the arrow array concatenation fails
ValueError: if the list of arrays is empty
TypeError: if the arrays to be concatenated have different types
Returns:
array (:obj:`pyarrow.Array`): the concatenated array
"""
arrays = list(arrays)
array_types = {array.type for array in arrays}
if not array_types:
raise ValueError("Couldn't concatenate empty list of arrays")
if len(array_types) > 1:
array_types = list(array_types)
raise TypeError(f"Couldn't concatenate arrays with different types {array_types[0]} and {array_types[1]}")
array_type = arrays[0].type
arrays = [chunk for arr in arrays for chunk in (arr.chunks if isinstance(arr, pa.ChunkedArray) else (arr,))]
if not _is_extension_type(array_type):
return pa.concat_arrays(arrays)
def _offsets_concat(offsets):
offset = offsets[0]
concatenated_offsets = offset
for offset in offsets[1:]:
offset = pc.subtract(offset, offset[0])
offset = pc.add(offset[1:], concatenated_offsets[-1])
concatenated_offsets = pa.concat_arrays([concatenated_offsets, offset])
return concatenated_offsets
def _concat_arrays(arrays):
array_type = arrays[0].type
if isinstance(array_type, pa.ExtensionType):
return array_type.wrap_array(_concat_arrays([array.storage for array in arrays]))
elif pa.types.is_struct(array_type):
return pa.StructArray.from_arrays(
[_concat_arrays([array.field(field.name) for array in arrays]) for field in array_type],
fields=list(array_type),
mask=pa.concat_arrays([array.is_null() for array in arrays]),
)
elif pa.types.is_list(array_type):
if any(array.null_count > 0 for array in arrays):
if config.PYARROW_VERSION.major < 10:
warnings.warn(
"None values are converted to empty lists in `pyarrow<10.0.0` when concatenating list arrays with None values. Install `pyarrow>=10.0.0` to avoid this behavior. More info: https://github.com/huggingface/datasets/issues/3676."
)
else:
return pa.ListArray.from_arrays(
_offsets_concat([array.offsets for array in arrays]),
_concat_arrays([array.values for array in arrays]),
mask=pa.concat_arrays([array.is_null() for array in arrays]),
)
return pa.ListArray.from_arrays(
_offsets_concat([array.offsets for array in arrays]),
_concat_arrays([array.values for array in arrays]),
)
elif pa.types.is_fixed_size_list(array_type):
if config.PYARROW_VERSION.major < 15:
# PyArrow bug: https://github.com/apache/arrow/issues/35360
return pa.FixedSizeListArray.from_arrays(
_concat_arrays([array.values[array.offset * array.type.list_size :] for array in arrays]),
array_type.list_size,
)
else:
return pa.FixedSizeListArray.from_arrays(
_concat_arrays([array.values for array in arrays]),
array_type.value_type,
array_type.list_size,
)
return pa.concat_arrays(arrays)
return _concat_arrays(arrays)
@_wrap_for_chunked_arrays
def array_cast(array: pa.Array, pa_type: pa.DataType, allow_number_to_str=True):
"""Improved version of `pa.Array.cast`
It supports casting `pa.StructArray` objects to re-order the fields.
It also let you control certain aspects of the casting, e.g. whether
to disable numbers (`floats` or `ints`) to strings.
Args:
array (`pa.Array`):
PyArrow array to cast
pa_type (`pa.DataType`):
Target PyArrow type
allow_number_to_str (`bool`, defaults to `True`):
Whether to allow casting numbers to strings.
Defaults to `True`.
Raises:
`pa.ArrowInvalidError`: if the arrow data casting fails
`TypeError`: if the target type is not supported according, e.g.
- if a field is missing
- if casting from numbers to strings and `allow_number_to_str` is `False`
Returns:
`List[pyarrow.Array]`: the casted array
"""
_c = partial(array_cast, allow_number_to_str=allow_number_to_str)
if isinstance(array, pa.ExtensionArray):
array = array.storage
if isinstance(pa_type, pa.ExtensionType):
return pa_type.wrap_array(_c(array, pa_type.storage_type))
elif array.type == pa_type:
return array
elif pa.types.is_struct(array.type):
if pa.types.is_struct(pa_type) and ({field.name for field in pa_type} == {field.name for field in array.type}):
if array.type.num_fields == 0:
return array
arrays = [_c(array.field(field.name), field.type) for field in pa_type]
return pa.StructArray.from_arrays(arrays, fields=list(pa_type), mask=array.is_null())
elif pa.types.is_list(array.type):
if pa.types.is_fixed_size_list(pa_type):
if pa_type.list_size * len(array) == len(array.values):
return pa.FixedSizeListArray.from_arrays(
_c(array.values, pa_type.value_type),
pa_type.list_size,
)
elif pa.types.is_list(pa_type):
if array.null_count > 0:
if config.PYARROW_VERSION.major < 10:
warnings.warn(
f"None values are converted to empty lists in `pyarrow<10.0.0` when converting array to {pa_type}. Install `pyarrow>=10.0.0` to avoid this behavior. More info: https://github.com/huggingface/datasets/issues/3676."
)
else:
return pa.ListArray.from_arrays(
array.offsets, _c(array.values, pa_type.value_type), mask=array.is_null()
)
return pa.ListArray.from_arrays(array.offsets, _c(array.values, pa_type.value_type))
elif pa.types.is_fixed_size_list(array.type):
array_values = array.values
if config.PYARROW_VERSION.major < 15:
# PyArrow bug: https://github.com/apache/arrow/issues/35360
array_values = array.values[array.offset * array.type.list_size :]
if pa.types.is_fixed_size_list(pa_type):
return pa.FixedSizeListArray.from_arrays(
_c(array_values, pa_type.value_type),
pa_type.list_size,
)
elif pa.types.is_list(pa_type):
offsets_arr = pa.array(np.arange(len(array) + 1) * array.type.list_size, pa.int32())
if array.null_count > 0:
if config.PYARROW_VERSION.major < 10:
warnings.warn(
f"None values are converted to empty lists in `pyarrow<10.0.0` when converting array to {pa_type}. Install `pyarrow>=10.0.0` to avoid this behavior. More info: https://github.com/huggingface/datasets/issues/3676."
)
else:
return pa.ListArray.from_arrays(
offsets_arr, _c(array_values, pa_type.value_type), mask=array.is_null()
)
return pa.ListArray.from_arrays(offsets_arr, _c(array_values, pa_type.value_type))
else:
if (
not allow_number_to_str
and pa.types.is_string(pa_type)
and (pa.types.is_floating(array.type) or pa.types.is_integer(array.type))
):
raise TypeError(
f"Couldn't cast array of type {array.type} to {pa_type} since allow_number_to_str is set to {allow_number_to_str}"
)
if pa.types.is_null(pa_type) and not pa.types.is_null(array.type):
raise TypeError(f"Couldn't cast array of type {array.type} to {pa_type}")
return array.cast(pa_type)
raise TypeError(f"Couldn't cast array of type\n{array.type}\nto\n{pa_type}")
@_wrap_for_chunked_arrays
def cast_array_to_feature(array: pa.Array, feature: "FeatureType", allow_number_to_str=True):
"""Cast an array to the arrow type that corresponds to the requested feature type.
For custom features like [`Audio`] or [`Image`], it takes into account the "cast_storage" methods
they defined to enable casting from other arrow types.
Args:
array (`pa.Array`):
The PyArrow array to cast.
feature (`datasets.features.FeatureType`):
The target feature type.
allow_number_to_str (`bool`, defaults to `True`):
Whether to allow casting numbers to strings.
Defaults to `True`.
Raises:
`pa.ArrowInvalidError`: if the arrow data casting fails
`TypeError`: if the target type is not supported according, e.g.
- if a field is missing
- if casting from numbers to strings and `allow_number_to_str` is `False`
Returns:
array (`pyarrow.Array`): the casted array
"""
from .features.features import Sequence, get_nested_type
_c = partial(cast_array_to_feature, allow_number_to_str=allow_number_to_str)
if isinstance(array, pa.ExtensionArray):
array = array.storage
if hasattr(feature, "cast_storage"):
return feature.cast_storage(array)
elif pa.types.is_struct(array.type):
# feature must be a dict or Sequence(subfeatures_dict)
if isinstance(feature, Sequence) and isinstance(feature.feature, dict):
feature = {
name: Sequence(subfeature, length=feature.length) for name, subfeature in feature.feature.items()
}
if isinstance(feature, dict) and {field.name for field in array.type} == set(feature):
if array.type.num_fields == 0:
return array
arrays = [_c(array.field(name), subfeature) for name, subfeature in feature.items()]
return pa.StructArray.from_arrays(arrays, names=list(feature), mask=array.is_null())
elif pa.types.is_list(array.type):
# feature must be either [subfeature] or Sequence(subfeature)
if isinstance(feature, list):
casted_values = _c(array.values, feature[0])
if casted_values.type == array.values.type:
return array
else:
if array.null_count > 0:
if config.PYARROW_VERSION.major < 10:
warnings.warn(
f"None values are converted to empty lists in `pyarrow<10.0.0` when converting array to {feature}. Install `pyarrow>=10.0.0` to avoid this behavior. More info: https://github.com/huggingface/datasets/issues/3676."
)
else:
return pa.ListArray.from_arrays(array.offsets, casted_values, mask=array.is_null())
return pa.ListArray.from_arrays(array.offsets, casted_values)
elif isinstance(feature, Sequence):
if feature.length > -1:
if feature.length * len(array) == len(array.values):
return pa.FixedSizeListArray.from_arrays(_c(array.values, feature.feature), feature.length)
else:
casted_values = _c(array.values, feature.feature)
if casted_values.type == array.values.type:
return array
else:
if array.null_count > 0:
if config.PYARROW_VERSION.major < 10:
warnings.warn(
f"None values are converted to empty lists in `pyarrow<10.0.0` when converting array to {feature}. Install `pyarrow>=10.0.0` to avoid this behavior. More info: https://github.com/huggingface/datasets/issues/3676."
)
else:
return pa.ListArray.from_arrays(
array.offsets, _c(array.values, feature.feature), mask=array.is_null()
)
return pa.ListArray.from_arrays(array.offsets, _c(array.values, feature.feature))
elif pa.types.is_fixed_size_list(array.type):
# feature must be either [subfeature] or Sequence(subfeature)
array_values = array.values
if config.PYARROW_VERSION.major < 15:
# PyArrow bug: https://github.com/apache/arrow/issues/35360
array_values = array.values[array.offset * array.type.list_size :]
if isinstance(feature, list):
if array.null_count > 0:
if config.PYARROW_VERSION.major < 10:
warnings.warn(
f"None values are converted to empty lists when converting array to {feature}. Install `pyarrow>=10.0.0` to avoid this behavior. More info: https://github.com/huggingface/datasets/issues/3676. This will raise an error in a future major version of `datasets`"
)
else:
return pa.ListArray.from_arrays(array.offsets, _c(array_values, feature[0]), mask=array.is_null())
return pa.ListArray.from_arrays(array.offsets, _c(array_values, feature[0]))
elif isinstance(feature, Sequence):
if feature.length > -1:
if feature.length * len(array) == len(array_values):
return pa.FixedSizeListArray.from_arrays(_c(array_values, feature.feature), feature.length)
else:
offsets_arr = pa.array(np.arange(len(array) + 1) * array.type.list_size, pa.int32())
if array.null_count > 0:
if config.PYARROW_VERSION.major < 10:
warnings.warn(
f"None values are converted to empty lists when converting array to {feature}. Install `pyarrow>=10.0.0` to avoid this behavior. More info: https://github.com/huggingface/datasets/issues/3676. This will raise an error in a future major version of `datasets`"
)
else:
return pa.ListArray.from_arrays(
offsets_arr, _c(array_values, feature.feature), mask=array.is_null()
)
return pa.ListArray.from_arrays(offsets_arr, _c(array_values, feature.feature))
if pa.types.is_null(array.type):
return array_cast(array, get_nested_type(feature), allow_number_to_str=allow_number_to_str)
elif not isinstance(feature, (Sequence, dict, list, tuple)):
return array_cast(array, feature(), allow_number_to_str=allow_number_to_str)
raise TypeError(f"Couldn't cast array of type\n{array.type}\nto\n{feature}")
@_wrap_for_chunked_arrays
def embed_array_storage(array: pa.Array, feature: "FeatureType"):
"""Embed data into an arrays's storage.
For custom features like Audio or Image, it takes into account the "embed_storage" methods
they defined to enable embedding external data (e.g. an image file) into an other arrow types.
<Added version="2.4.0"/>
Args:
array (`pa.Array`):
The PyArrow array in which to embed data.
feature (`datasets.features.FeatureType`):
Array features.
Raises:
`TypeError`: if the target type is not supported according, e.g.
- if a field is missing
Returns:
array (`pyarrow.Array`): the casted array
"""
from .features import Sequence
_e = embed_array_storage
if isinstance(array, pa.ExtensionArray):
array = array.storage
if hasattr(feature, "embed_storage"):
return feature.embed_storage(array)
elif pa.types.is_struct(array.type):
# feature must be a dict or Sequence(subfeatures_dict)
if isinstance(feature, Sequence) and isinstance(feature.feature, dict):
feature = {
name: Sequence(subfeature, length=feature.length) for name, subfeature in feature.feature.items()
}
if isinstance(feature, dict):
arrays = [_e(array.field(name), subfeature) for name, subfeature in feature.items()]
return pa.StructArray.from_arrays(arrays, names=list(feature), mask=array.is_null())
elif pa.types.is_list(array.type):
# feature must be either [subfeature] or Sequence(subfeature)
if isinstance(feature, list):
if array.null_count > 0:
if config.PYARROW_VERSION.major < 10:
warnings.warn(
f"None values are converted to empty lists when embedding array storage with {feature}. Install `pyarrow>=10.0.0` to avoid this behavior. More info: https://github.com/huggingface/datasets/issues/3676. This will raise an error in a future major version of `datasets`"
)
else:
return pa.ListArray.from_arrays(array.offsets, _e(array.values, feature[0]), mask=array.is_null())
return pa.ListArray.from_arrays(array.offsets, _e(array.values, feature[0]))
elif isinstance(feature, Sequence):
if feature.length > -1:
if feature.length * len(array) == len(array.values):
return pa.FixedSizeListArray.from_arrays(_e(array.values, feature.feature), feature.length)
else:
casted_values = _e(array.values, feature.feature)
if casted_values.type == array.values.type:
return array
else:
if array.null_count > 0:
if config.PYARROW_VERSION.major < 10:
warnings.warn(
f"None values are converted to empty lists when embedding array storage with {feature}. Install `pyarrow>=10.0.0` to avoid this behavior. More info: https://github.com/huggingface/datasets/issues/3676. This will raise an error in a future major version of `datasets`"
)
else:
return pa.ListArray.from_arrays(
array.offsets, _e(array.values, feature.feature), mask=array.is_null()
)
return pa.ListArray.from_arrays(array.offsets, _e(array.values, feature.feature))
elif pa.types.is_fixed_size_list(array.type):
# feature must be either [subfeature] or Sequence(subfeature)
array_values = array.values
if config.PYARROW_VERSION.major < 15:
# PyArrow bug: https://github.com/apache/arrow/issues/35360
array_values = array.values[array.offset * array.type.list_size :]
if isinstance(feature, list):
if array.null_count > 0:
if config.PYARROW_VERSION.major < 10:
warnings.warn(
f"None values are converted to empty lists when embedding array storage with {feature}. Install `pyarrow>=10.0.0` to avoid this behavior. More info: https://github.com/huggingface/datasets/issues/3676. This will raise an error in a future major version of `datasets`"
)
else:
return pa.ListArray.from_arrays(array.offsets, _e(array_values, feature[0]), mask=array.is_null())
return pa.ListArray.from_arrays(array.offsets, _e(array_values, feature[0]))
elif isinstance(feature, Sequence):
if feature.length > -1:
if feature.length * len(array) == len(array_values):
return pa.FixedSizeListArray.from_arrays(_e(array_values, feature.feature), feature.length)
else:
offsets_arr = pa.array(np.arange(len(array) + 1) * array.type.list_size, pa.int32())
if array.null_count > 0:
if config.PYARROW_VERSION.major < 10:
warnings.warn(
f"None values are converted to empty lists when embedding array storage with {feature}. Install `pyarrow>=10.0.0` to avoid this behavior. More info: https://github.com/huggingface/datasets/issues/3676. This will raise an error in a future major version of `datasets`"
)
else:
return pa.ListArray.from_arrays(
offsets_arr, _e(array_values, feature.feature), mask=array.is_null()
)
return pa.ListArray.from_arrays(offsets_arr, _e(array_values, feature.feature))
if not isinstance(feature, (Sequence, dict, list, tuple)):
return array
raise TypeError(f"Couldn't embed array of type\n{array.type}\nwith\n{feature}")
def cast_table_to_features(table: pa.Table, features: "Features"):
"""Cast a table to the arrow schema that corresponds to the requested features.
Args:
table (`pyarrow.Table`):
PyArrow table to cast.
features ([`Features`]):
Target features.
Returns:
table (`pyarrow.Table`): the casted table
"""
if sorted(table.column_names) != sorted(features):
raise ValueError(f"Couldn't cast\n{table.schema}\nto\n{features}\nbecause column names don't match")
arrays = [cast_array_to_feature(table[name], feature) for name, feature in features.items()]
return pa.Table.from_arrays(arrays, schema=features.arrow_schema)
def cast_table_to_schema(table: pa.Table, schema: pa.Schema):
"""Cast a table to the arrow schema. Different from `cast_table_to_features`, this method can preserve nullability.
Args:
table (`pa.Table`):
PyArrow table to cast.
features ([`Features`]):
Target features.
Returns:
`pa.Table`: the casted table
"""
from .features import Features
features = Features.from_arrow_schema(schema)
if sorted(table.column_names) != sorted(features):
raise ValueError(f"Couldn't cast\n{table.schema}\nto\n{features}\nbecause column names don't match")
arrays = [cast_array_to_feature(table[name], feature) for name, feature in features.items()]
return pa.Table.from_arrays(arrays, schema=schema)
def embed_table_storage(table: pa.Table):
"""Embed external data into a table's storage.
<Added version="2.4.0"/>
Args:
table (`pyarrow.Table`):
PyArrow table in which to embed data.
Returns:
table (`pyarrow.Table`): the table with embedded data
"""
from .features.features import Features, require_storage_embed
features = Features.from_arrow_schema(table.schema)
arrays = [
embed_array_storage(table[name], feature) if require_storage_embed(feature) else table[name]
for name, feature in features.items()
]
return pa.Table.from_arrays(arrays, schema=features.arrow_schema)
def table_cast(table: pa.Table, schema: pa.Schema):
"""Improved version of `pa.Table.cast`.
It supports casting to feature types stored in the schema metadata.
Args:
table (`pyarrow.Table`):
PyArrow table to cast.
schema (`pyarrow.Schema`):
Target PyArrow schema.
Returns:
table (`pyarrow.Table`): the casted table
"""
if table.schema != schema:
return cast_table_to_schema(table, schema)
elif table.schema.metadata != schema.metadata:
return table.replace_schema_metadata(schema.metadata)
else:
return table
def table_flatten(table: pa.Table):
"""Improved version of `pa.Table.flatten`.
It behaves as `pa.Table.flatten` in a sense it does 1-step flatten of the columns with a struct type into one column per struct field,
but updates the metadata and skips decodable features unless the `decode` attribute of these features is set to False.
Args:
table (`pa.Table`):
PyArrow table to flatten.
Returns:
`Table`: the flattened table
"""
from .features import Features
features = Features.from_arrow_schema(table.schema)
if any(hasattr(subfeature, "flatten") and subfeature.flatten() == subfeature for subfeature in features.values()):
flat_arrays = []
flat_column_names = []
for field in table.schema:
array = table.column(field.name)
subfeature = features[field.name]
if pa.types.is_struct(field.type) and (
not hasattr(subfeature, "flatten") or subfeature.flatten() != subfeature
):
flat_arrays.extend(array.flatten())
flat_column_names.extend([f"{field.name}.{subfield.name}" for subfield in field.type])
else:
flat_arrays.append(array)
flat_column_names.append(field.name)
flat_table = pa.Table.from_arrays(
flat_arrays,
names=flat_column_names,
)
else:
flat_table = table.flatten()
# Preserve complex types in the metadata
flat_features = features.flatten(max_depth=2)
flat_features = Features({column_name: flat_features[column_name] for column_name in flat_table.column_names})
return flat_table.replace_schema_metadata(flat_features.arrow_schema.metadata)
def table_visitor(table: pa.Table, function: Callable[[pa.Array], None]):
"""Visit all arrays in a table and apply a function to them.
Args:
table (`pyarrow.Table`):
PyArrow table to visit.
function (`Callable[[pa.Array], None]`):
Function to apply to each array.
"""
from .features import Features, Sequence
features = Features.from_arrow_schema(table.schema)
def _visit(array, feature):
if isinstance(array, pa.ChunkedArray):
for chunk in array.chunks:
_visit(chunk, feature)
else:
if isinstance(array, pa.ExtensionArray):
array = array.storage
function(array, feature)
if pa.types.is_struct(array.type) and not hasattr(feature, "cast_storage"):
if isinstance(feature, Sequence) and isinstance(feature.feature, dict):
feature = {
name: Sequence(subfeature, length=feature.length)
for name, subfeature in feature.feature.items()
}
for name, subfeature in feature.items():
_visit(array.field(name), subfeature)
elif pa.types.is_list(array.type):
if isinstance(feature, list):
_visit(array.values, feature[0])
elif isinstance(feature, Sequence):
_visit(array.values, feature.feature)
for name, feature in features.items():
_visit(table[name], feature)
def table_iter(table: Table, batch_size: int, drop_last_batch=False) -> Iterator[pa.Table]:
"""Iterate over sub-tables of size `batch_size`.
Args:
table (`pyarrow.Table`):
PyArrow table to iterate over.
batch_size (`int`):
Size of each sub-table to yield.
drop_last_batch (`bool`, defaults to `False`):
Drop the last batch if it is smaller than `batch_size`.
"""
chunks_buffer = []
chunks_buffer_size = 0
for chunk in table.to_reader(max_chunksize=batch_size):
if len(chunk) == 0:
continue
elif chunks_buffer_size + len(chunk) < batch_size:
chunks_buffer.append(chunk)
chunks_buffer_size += len(chunk)
continue
elif chunks_buffer_size + len(chunk) == batch_size:
chunks_buffer.append(chunk)
yield pa.Table.from_batches(chunks_buffer)
chunks_buffer = []
chunks_buffer_size = 0
else:
cropped_chunk_length = batch_size - chunks_buffer_size
chunks_buffer.append(chunk.slice(0, cropped_chunk_length))
yield pa.Table.from_batches(chunks_buffer)
chunks_buffer = [chunk.slice(cropped_chunk_length, len(chunk) - cropped_chunk_length)]
chunks_buffer_size = len(chunk) - cropped_chunk_length
if not drop_last_batch and chunks_buffer:
yield pa.Table.from_batches(chunks_buffer)
| 0
|
hf_public_repos/datasets/src
|
hf_public_repos/datasets/src/datasets/config.py
|
import importlib
import importlib.metadata
import logging
import os
import platform
from pathlib import Path
from typing import Optional
from packaging import version
logger = logging.getLogger(__name__.split(".", 1)[0]) # to avoid circular import from .utils.logging
# Datasets
S3_DATASETS_BUCKET_PREFIX = "https://s3.amazonaws.com/datasets.huggingface.co/datasets/datasets"
CLOUDFRONT_DATASETS_DISTRIB_PREFIX = "https://cdn-datasets.huggingface.co/datasets/datasets"
REPO_DATASETS_URL = "https://raw.githubusercontent.com/huggingface/datasets/{revision}/datasets/{path}/{name}"
# Metrics
S3_METRICS_BUCKET_PREFIX = "https://s3.amazonaws.com/datasets.huggingface.co/datasets/metrics"
CLOUDFRONT_METRICS_DISTRIB_PREFIX = "https://cdn-datasets.huggingface.co/datasets/metric"
REPO_METRICS_URL = "https://raw.githubusercontent.com/huggingface/datasets/{revision}/metrics/{path}/{name}"
# Hub
HF_ENDPOINT = os.environ.get("HF_ENDPOINT", "https://huggingface.co")
HUB_DATASETS_URL = HF_ENDPOINT + "/datasets/{repo_id}/resolve/{revision}/{path}"
HUB_DATASETS_HFFS_URL = "hf://datasets/{repo_id}@{revision}/{path}"
HUB_DEFAULT_VERSION = "main"
PY_VERSION = version.parse(platform.python_version())
# General environment variables accepted values for booleans
ENV_VARS_TRUE_VALUES = {"1", "ON", "YES", "TRUE"}
ENV_VARS_FALSE_VALUES = {"0", "OFF", "NO", "FALSE"}
ENV_VARS_TRUE_AND_AUTO_VALUES = ENV_VARS_TRUE_VALUES.union({"AUTO"})
ENV_VARS_FALSE_AND_AUTO_VALUES = ENV_VARS_FALSE_VALUES.union({"AUTO"})
# Imports
DILL_VERSION = version.parse(importlib.metadata.version("dill"))
FSSPEC_VERSION = version.parse(importlib.metadata.version("fsspec"))
PANDAS_VERSION = version.parse(importlib.metadata.version("pandas"))
PYARROW_VERSION = version.parse(importlib.metadata.version("pyarrow"))
USE_TF = os.environ.get("USE_TF", "AUTO").upper()
USE_TORCH = os.environ.get("USE_TORCH", "AUTO").upper()
USE_JAX = os.environ.get("USE_JAX", "AUTO").upper()
TORCH_VERSION = "N/A"
TORCH_AVAILABLE = False
if USE_TORCH in ENV_VARS_TRUE_AND_AUTO_VALUES and USE_TF not in ENV_VARS_TRUE_VALUES:
TORCH_AVAILABLE = importlib.util.find_spec("torch") is not None
if TORCH_AVAILABLE:
try:
TORCH_VERSION = version.parse(importlib.metadata.version("torch"))
logger.info(f"PyTorch version {TORCH_VERSION} available.")
except importlib.metadata.PackageNotFoundError:
pass
else:
logger.info("Disabling PyTorch because USE_TF is set")
TF_VERSION = "N/A"
TF_AVAILABLE = False
if USE_TF in ENV_VARS_TRUE_AND_AUTO_VALUES and USE_TORCH not in ENV_VARS_TRUE_VALUES:
TF_AVAILABLE = importlib.util.find_spec("tensorflow") is not None
if TF_AVAILABLE:
# For the metadata, we have to look for both tensorflow and tensorflow-cpu
for package in [
"tensorflow",
"tensorflow-cpu",
"tensorflow-gpu",
"tf-nightly",
"tf-nightly-cpu",
"tf-nightly-gpu",
"intel-tensorflow",
"tensorflow-rocm",
"tensorflow-macos",
]:
try:
TF_VERSION = version.parse(importlib.metadata.version(package))
except importlib.metadata.PackageNotFoundError:
continue
else:
break
else:
TF_AVAILABLE = False
if TF_AVAILABLE:
if TF_VERSION.major < 2:
logger.info(f"TensorFlow found but with version {TF_VERSION}. `datasets` requires version 2 minimum.")
TF_AVAILABLE = False
else:
logger.info(f"TensorFlow version {TF_VERSION} available.")
else:
logger.info("Disabling Tensorflow because USE_TORCH is set")
JAX_VERSION = "N/A"
JAX_AVAILABLE = False
if USE_JAX in ENV_VARS_TRUE_AND_AUTO_VALUES:
JAX_AVAILABLE = importlib.util.find_spec("jax") is not None and importlib.util.find_spec("jaxlib") is not None
if JAX_AVAILABLE:
try:
JAX_VERSION = version.parse(importlib.metadata.version("jax"))
logger.info(f"JAX version {JAX_VERSION} available.")
except importlib.metadata.PackageNotFoundError:
pass
else:
logger.info("Disabling JAX because USE_JAX is set to False")
USE_BEAM = os.environ.get("USE_BEAM", "AUTO").upper()
BEAM_VERSION = "N/A"
BEAM_AVAILABLE = False
if USE_BEAM in ENV_VARS_TRUE_AND_AUTO_VALUES:
try:
BEAM_VERSION = version.parse(importlib.metadata.version("apache_beam"))
BEAM_AVAILABLE = True
logger.info(f"Apache Beam version {BEAM_VERSION} available.")
except importlib.metadata.PackageNotFoundError:
pass
else:
logger.info("Disabling Apache Beam because USE_BEAM is set to False")
# Optional tools for data loading
SQLALCHEMY_AVAILABLE = importlib.util.find_spec("sqlalchemy") is not None
# Optional tools for feature decoding
PIL_AVAILABLE = importlib.util.find_spec("PIL") is not None
IS_OPUS_SUPPORTED = importlib.util.find_spec("soundfile") is not None and version.parse(
importlib.import_module("soundfile").__libsndfile_version__
) >= version.parse("1.0.31")
IS_MP3_SUPPORTED = importlib.util.find_spec("soundfile") is not None and version.parse(
importlib.import_module("soundfile").__libsndfile_version__
) >= version.parse("1.1.0")
# Optional compression tools
RARFILE_AVAILABLE = importlib.util.find_spec("rarfile") is not None
ZSTANDARD_AVAILABLE = importlib.util.find_spec("zstandard") is not None
LZ4_AVAILABLE = importlib.util.find_spec("lz4") is not None
PY7ZR_AVAILABLE = importlib.util.find_spec("py7zr") is not None
# Cache location
DEFAULT_XDG_CACHE_HOME = "~/.cache"
XDG_CACHE_HOME = os.getenv("XDG_CACHE_HOME", DEFAULT_XDG_CACHE_HOME)
DEFAULT_HF_CACHE_HOME = os.path.join(XDG_CACHE_HOME, "huggingface")
HF_CACHE_HOME = os.path.expanduser(os.getenv("HF_HOME", DEFAULT_HF_CACHE_HOME))
DEFAULT_HF_DATASETS_CACHE = os.path.join(HF_CACHE_HOME, "datasets")
HF_DATASETS_CACHE = Path(os.getenv("HF_DATASETS_CACHE", DEFAULT_HF_DATASETS_CACHE))
DEFAULT_HF_METRICS_CACHE = os.path.join(HF_CACHE_HOME, "metrics")
HF_METRICS_CACHE = Path(os.getenv("HF_METRICS_CACHE", DEFAULT_HF_METRICS_CACHE))
DEFAULT_HF_MODULES_CACHE = os.path.join(HF_CACHE_HOME, "modules")
HF_MODULES_CACHE = Path(os.getenv("HF_MODULES_CACHE", DEFAULT_HF_MODULES_CACHE))
DOWNLOADED_DATASETS_DIR = "downloads"
DEFAULT_DOWNLOADED_DATASETS_PATH = os.path.join(HF_DATASETS_CACHE, DOWNLOADED_DATASETS_DIR)
DOWNLOADED_DATASETS_PATH = Path(os.getenv("HF_DATASETS_DOWNLOADED_DATASETS_PATH", DEFAULT_DOWNLOADED_DATASETS_PATH))
EXTRACTED_DATASETS_DIR = "extracted"
DEFAULT_EXTRACTED_DATASETS_PATH = os.path.join(DEFAULT_DOWNLOADED_DATASETS_PATH, EXTRACTED_DATASETS_DIR)
EXTRACTED_DATASETS_PATH = Path(os.getenv("HF_DATASETS_EXTRACTED_DATASETS_PATH", DEFAULT_EXTRACTED_DATASETS_PATH))
# Download count for the website
HF_UPDATE_DOWNLOAD_COUNTS = (
os.environ.get("HF_UPDATE_DOWNLOAD_COUNTS", "AUTO").upper() in ENV_VARS_TRUE_AND_AUTO_VALUES
)
# Remote dataset scripts support
__HF_DATASETS_TRUST_REMOTE_CODE = os.environ.get("HF_DATASETS_TRUST_REMOTE_CODE", "1")
HF_DATASETS_TRUST_REMOTE_CODE: Optional[bool] = (
True
if __HF_DATASETS_TRUST_REMOTE_CODE.upper() in ENV_VARS_TRUE_VALUES
else False
if __HF_DATASETS_TRUST_REMOTE_CODE.upper() in ENV_VARS_FALSE_VALUES
else None
)
TIME_OUT_REMOTE_CODE = 15
# Batch size constants. For more info, see:
# https://github.com/apache/arrow/blob/master/docs/source/cpp/arrays.rst#size-limitations-and-recommendations)
DEFAULT_MAX_BATCH_SIZE = 1000
# Size of the preloaded record batch in `Dataset.__iter__`
ARROW_READER_BATCH_SIZE_IN_DATASET_ITER = 10
# Max shard size in bytes (e.g. to shard parquet datasets in push_to_hub or download_and_prepare)
MAX_SHARD_SIZE = "500MB"
# Parquet configuration
PARQUET_ROW_GROUP_SIZE_FOR_AUDIO_DATASETS = 100
PARQUET_ROW_GROUP_SIZE_FOR_IMAGE_DATASETS = 100
PARQUET_ROW_GROUP_SIZE_FOR_BINARY_DATASETS = 100
# Offline mode
HF_DATASETS_OFFLINE = os.environ.get("HF_DATASETS_OFFLINE", "AUTO").upper() in ENV_VARS_TRUE_VALUES
# Here, `True` will disable progress bars globally without possibility of enabling it
# programmatically. `False` will enable them without possibility of disabling them.
# If environment variable is not set (None), then the user is free to enable/disable
# them programmatically.
# TL;DR: env variable has priority over code
__HF_DATASETS_DISABLE_PROGRESS_BARS = os.environ.get("HF_DATASETS_DISABLE_PROGRESS_BARS")
HF_DATASETS_DISABLE_PROGRESS_BARS: Optional[bool] = (
__HF_DATASETS_DISABLE_PROGRESS_BARS.upper() in ENV_VARS_TRUE_VALUES
if __HF_DATASETS_DISABLE_PROGRESS_BARS is not None
else None
)
# In-memory
DEFAULT_IN_MEMORY_MAX_SIZE = 0 # Disabled
IN_MEMORY_MAX_SIZE = float(os.environ.get("HF_DATASETS_IN_MEMORY_MAX_SIZE", DEFAULT_IN_MEMORY_MAX_SIZE))
# File names
DATASET_ARROW_FILENAME = "dataset.arrow"
DATASET_INDICES_FILENAME = "indices.arrow"
DATASET_STATE_JSON_FILENAME = "state.json"
DATASET_INFO_FILENAME = "dataset_info.json"
DATASETDICT_INFOS_FILENAME = "dataset_infos.json"
LICENSE_FILENAME = "LICENSE"
METRIC_INFO_FILENAME = "metric_info.json"
DATASETDICT_JSON_FILENAME = "dataset_dict.json"
METADATA_CONFIGS_FIELD = "configs"
MODULE_NAME_FOR_DYNAMIC_MODULES = "datasets_modules"
MAX_DATASET_CONFIG_ID_READABLE_LENGTH = 255
# Streaming
STREAMING_READ_MAX_RETRIES = 20
STREAMING_READ_RETRY_INTERVAL = 5
# Datasets without script
DATA_FILES_MAX_NUMBER_FOR_MODULE_INFERENCE = 200
GLOBBED_DATA_FILES_MAX_NUMBER_FOR_MODULE_INFERENCE = 10
ARCHIVED_DATA_FILES_MAX_NUMBER_FOR_MODULE_INFERENCE = 200
# Progress bars
PBAR_REFRESH_TIME_INTERVAL = 0.05 # 20 progress updates per sec
# Maximum number of uploaded files per commit
UPLOADS_MAX_NUMBER_PER_COMMIT = 50
# Backward compatibiliy
MAX_TABLE_NBYTES_FOR_PICKLING = 4 << 30
| 0
|
hf_public_repos/datasets/src/datasets
|
hf_public_repos/datasets/src/datasets/filesystems/compression.py
|
import os
from typing import Optional
import fsspec
from fsspec.archive import AbstractArchiveFileSystem
from fsspec.utils import DEFAULT_BLOCK_SIZE
class BaseCompressedFileFileSystem(AbstractArchiveFileSystem):
"""Read contents of compressed file as a filesystem with one file inside."""
root_marker = ""
protocol: str = (
None # protocol passed in prefix to the url. ex: "gzip", for gzip://file.txt::http://foo.bar/file.txt.gz
)
compression: str = None # compression type in fsspec. ex: "gzip"
extension: str = None # extension of the filename to strip. ex: "".gz" to get file.txt from file.txt.gz
def __init__(
self, fo: str = "", target_protocol: Optional[str] = None, target_options: Optional[dict] = None, **kwargs
):
"""
The compressed file system can be instantiated from any compressed file.
It reads the contents of compressed file as a filesystem with one file inside, as if it was an archive.
The single file inside the filesystem is named after the compresssed file,
without the compression extension at the end of the filename.
Args:
fo (:obj:``str``): Path to compressed file. Will fetch file using ``fsspec.open()``
mode (:obj:``str``): Currently, only 'rb' accepted
target_protocol(:obj:``str``, optional): To override the FS protocol inferred from a URL.
target_options (:obj:``dict``, optional): Kwargs passed when instantiating the target FS.
"""
super().__init__(self, **kwargs)
# always open as "rb" since fsspec can then use the TextIOWrapper to make it work for "r" mode
self.file = fsspec.open(
fo,
mode="rb",
protocol=target_protocol,
compression=self.compression,
client_kwargs={
"requote_redirect_url": False, # see https://github.com/huggingface/datasets/pull/5459
"trust_env": True, # Enable reading proxy env variables.
**(target_options or {}).pop("client_kwargs", {}), # To avoid issues if it was already passed.
},
**(target_options or {}),
)
self.compressed_name = os.path.basename(self.file.path.split("::")[0])
self.uncompressed_name = (
self.compressed_name[: self.compressed_name.rindex(".")]
if "." in self.compressed_name
else self.compressed_name
)
self.dir_cache = None
@classmethod
def _strip_protocol(cls, path):
# compressed file paths are always relative to the archive root
return super()._strip_protocol(path).lstrip("/")
def _get_dirs(self):
if self.dir_cache is None:
f = {**self.file.fs.info(self.file.path), "name": self.uncompressed_name}
self.dir_cache = {f["name"]: f}
def cat(self, path: str):
return self.file.open().read()
def _open(
self,
path: str,
mode: str = "rb",
block_size=None,
autocommit=True,
cache_options=None,
**kwargs,
):
path = self._strip_protocol(path)
if mode != "rb":
raise ValueError(f"Tried to read with mode {mode} on file {self.file.path} opened with mode 'rb'")
return self.file.open()
class Bz2FileSystem(BaseCompressedFileFileSystem):
"""Read contents of BZ2 file as a filesystem with one file inside."""
protocol = "bz2"
compression = "bz2"
extension = ".bz2"
class GzipFileSystem(BaseCompressedFileFileSystem):
"""Read contents of GZIP file as a filesystem with one file inside."""
protocol = "gzip"
compression = "gzip"
extension = ".gz"
class Lz4FileSystem(BaseCompressedFileFileSystem):
"""Read contents of LZ4 file as a filesystem with one file inside."""
protocol = "lz4"
compression = "lz4"
extension = ".lz4"
class XzFileSystem(BaseCompressedFileFileSystem):
"""Read contents of .xz (LZMA) file as a filesystem with one file inside."""
protocol = "xz"
compression = "xz"
extension = ".xz"
class ZstdFileSystem(BaseCompressedFileFileSystem):
"""
Read contents of zstd file as a filesystem with one file inside.
Note that reading in binary mode with fsspec isn't supported yet:
https://github.com/indygreg/python-zstandard/issues/136
"""
protocol = "zstd"
compression = "zstd"
extension = ".zst"
def __init__(
self,
fo: str,
mode: str = "rb",
target_protocol: Optional[str] = None,
target_options: Optional[dict] = None,
block_size: int = DEFAULT_BLOCK_SIZE,
**kwargs,
):
super().__init__(
fo=fo,
mode=mode,
target_protocol=target_protocol,
target_options=target_options,
block_size=block_size,
**kwargs,
)
# We need to wrap the zstd decompressor to avoid this error in fsspec==2021.7.0 and zstandard==0.15.2:
#
# File "/Users/user/.virtualenvs/hf-datasets/lib/python3.7/site-packages/fsspec/core.py", line 145, in open
# out.close = close
# AttributeError: 'zstd.ZstdDecompressionReader' object attribute 'close' is read-only
#
# see https://github.com/intake/filesystem_spec/issues/725
_enter = self.file.__enter__
class WrappedFile:
def __init__(self, file_):
self._file = file_
def __enter__(self):
self._file.__enter__()
return self
def __exit__(self, *args, **kwargs):
self._file.__exit__(*args, **kwargs)
def __iter__(self):
return iter(self._file)
def __next__(self):
return next(self._file)
def __getattr__(self, attr):
return getattr(self._file, attr)
def fixed_enter(*args, **kwargs):
return WrappedFile(_enter(*args, **kwargs))
self.file.__enter__ = fixed_enter
| 0
|
hf_public_repos/datasets/src/datasets
|
hf_public_repos/datasets/src/datasets/filesystems/s3filesystem.py
|
import s3fs
from ..utils.deprecation_utils import deprecated
@deprecated("Use s3fs.S3FileSystem instead.")
class S3FileSystem(s3fs.S3FileSystem):
"""
`datasets.filesystems.S3FileSystem` is a subclass of [`s3fs.S3FileSystem`](https://s3fs.readthedocs.io/en/latest/api.html).
Users can use this class to access S3 as if it were a file system. It exposes a filesystem-like API (ls, cp, open, etc.) on top of S3 storage. Provide credentials either explicitly (`key=`, `secret=`) or with boto's credential methods. See botocore documentation for more information. If no credentials are available, use `anon=True`.
Args:
anon (`bool`, default to `False`):
Whether to use anonymous connection (public buckets only). If `False`, uses the key/secret given,
or boto's credential resolver (client_kwargs, environment, variables, config files, EC2 IAM server, in that order).
key (`str`):
If not anonymous, use this access key ID, if specified.
secret (`str`):
If not anonymous, use this secret access key, if specified.
token (`str`):
If not anonymous, use this security token, if specified.
use_ssl (`bool`, defaults to `True`):
Whether to use SSL in connections to S3; may be faster without, but insecure. If `use_ssl` is
also set in `client_kwargs`, the value set in `client_kwargs` will take priority.
s3_additional_kwargs (`dict`):
Parameters that are used when calling S3 API methods. Typically used for things
like ServerSideEncryption.
client_kwargs (`dict`):
Parameters for the botocore client.
requester_pays (`bool`, defaults to `False`):
Whether `RequesterPays` buckets are supported.
default_block_size (`int`):
If given, the default block size value used for `open()`, if no specific value is given at all time.
The built-in default is 5MB.
default_fill_cache (`bool`, defaults to `True`):
Whether to use cache filling with open by default. Refer to `S3File.open`.
default_cache_type (`str`, defaults to `bytes`):
If given, the default `cache_type` value used for `open()`. Set to `none` if no
caching is desired. See fsspec's documentation for other available `cache_type` values.
version_aware (`bool`, defaults to `False`):
Whether to support bucket versioning. If enable this will require the user to have
the necessary IAM permissions for dealing with versioned objects.
cache_regions (`bool`, defaults to `False`):
Whether to cache bucket regions. Whenever a new bucket is used, it will
first find out which region it belongs to and then use the client for that region.
asynchronous (`bool`, defaults to `False`):
Whether this instance is to be used from inside coroutines.
config_kwargs (`dict`):
Parameters passed to `botocore.client.Config`.
**kwargs:
Other parameters for core session.
session (`aiobotocore.session.AioSession`):
Session to be used for all connections. This session will be used inplace of creating
a new session inside S3FileSystem. For example: `aiobotocore.session.AioSession(profile='test_user')`.
skip_instance_cache (`bool`):
Control reuse of instances. Passed on to `fsspec`.
use_listings_cache (`bool`):
Control reuse of directory listings. Passed on to `fsspec`.
listings_expiry_time (`int` or `float`):
Control reuse of directory listings. Passed on to `fsspec`.
max_paths (`int`): Control reuse of directory listings. Passed on to `fsspec`.
Examples:
Listing files from public S3 bucket.
```py
>>> import datasets
>>> s3 = datasets.filesystems.S3FileSystem(anon=True) # doctest: +SKIP
>>> s3.ls('public-datasets/imdb/train') # doctest: +SKIP
['dataset_info.json.json','dataset.arrow','state.json']
```
Listing files from private S3 bucket using `aws_access_key_id` and `aws_secret_access_key`.
```py
>>> import datasets
>>> s3 = datasets.filesystems.S3FileSystem(key=aws_access_key_id, secret=aws_secret_access_key) # doctest: +SKIP
>>> s3.ls('my-private-datasets/imdb/train') # doctest: +SKIP
['dataset_info.json.json','dataset.arrow','state.json']
```
Using `S3Filesystem` with `botocore.session.Session` and custom `aws_profile`.
```py
>>> import botocore
>>> from datasets.filesystems import S3Filesystem
>>> s3_session = botocore.session.Session(profile_name='my_profile_name')
>>> s3 = S3FileSystem(session=s3_session) # doctest: +SKIP
```
Loading dataset from S3 using `S3Filesystem` and [`load_from_disk`].
```py
>>> from datasets import load_from_disk
>>> from datasets.filesystems import S3Filesystem
>>> s3 = S3FileSystem(key=aws_access_key_id, secret=aws_secret_access_key) # doctest: +SKIP
>>> dataset = load_from_disk('s3://my-private-datasets/imdb/train', storage_options=s3.storage_options) # doctest: +SKIP
>>> print(len(dataset))
25000
```
Saving dataset to S3 using `S3Filesystem` and [`Dataset.save_to_disk`].
```py
>>> from datasets import load_dataset
>>> from datasets.filesystems import S3Filesystem
>>> dataset = load_dataset("imdb")
>>> s3 = S3FileSystem(key=aws_access_key_id, secret=aws_secret_access_key) # doctest: +SKIP
>>> dataset.save_to_disk('s3://my-private-datasets/imdb/train', storage_options=s3.storage_options) # doctest: +SKIP
```
"""
| 0
|
hf_public_repos/datasets/src/datasets
|
hf_public_repos/datasets/src/datasets/filesystems/__init__.py
|
import importlib
import shutil
import threading
import warnings
from typing import List
import fsspec
import fsspec.asyn
from fsspec.implementations.local import LocalFileSystem
from ..utils.deprecation_utils import deprecated
from . import compression
_has_s3fs = importlib.util.find_spec("s3fs") is not None
if _has_s3fs:
from .s3filesystem import S3FileSystem # noqa: F401
COMPRESSION_FILESYSTEMS: List[compression.BaseCompressedFileFileSystem] = [
compression.Bz2FileSystem,
compression.GzipFileSystem,
compression.Lz4FileSystem,
compression.XzFileSystem,
compression.ZstdFileSystem,
]
# Register custom filesystems
for fs_class in COMPRESSION_FILESYSTEMS:
if fs_class.protocol in fsspec.registry and fsspec.registry[fs_class.protocol] is not fs_class:
warnings.warn(f"A filesystem protocol was already set for {fs_class.protocol} and will be overwritten.")
fsspec.register_implementation(fs_class.protocol, fs_class, clobber=True)
@deprecated(
"This function is deprecated and will be removed in a future version. Please use `fsspec.core.strip_protocol` instead."
)
def extract_path_from_uri(dataset_path: str) -> str:
"""
Preprocesses `dataset_path` and removes remote filesystem (e.g. removing `s3://`).
Args:
dataset_path (`str`):
Path (e.g. `dataset/train`) or remote uri (e.g. `s3://my-bucket/dataset/train`) of the dataset directory.
"""
if "://" in dataset_path:
dataset_path = dataset_path.split("://")[1]
return dataset_path
def is_remote_filesystem(fs: fsspec.AbstractFileSystem) -> bool:
"""
Checks if `fs` is a remote filesystem.
Args:
fs (`fsspec.spec.AbstractFileSystem`):
An abstract super-class for pythonic file-systems, e.g. `fsspec.filesystem(\'file\')` or [`datasets.filesystems.S3FileSystem`].
"""
return not isinstance(fs, LocalFileSystem)
def rename(fs: fsspec.AbstractFileSystem, src: str, dst: str):
"""
Renames the file `src` in `fs` to `dst`.
"""
if not is_remote_filesystem(fs):
# LocalFileSystem.mv does copy + rm, it is more efficient to simply move a local directory
shutil.move(fs._strip_protocol(src), fs._strip_protocol(dst))
else:
fs.mv(src, dst, recursive=True)
def _reset_fsspec_lock() -> None:
"""
Clear reference to the loop and thread.
This is necessary otherwise HTTPFileSystem hangs in the ML training loop.
Only required for fsspec >= 0.9.0
See https://github.com/fsspec/gcsfs/issues/379
"""
if hasattr(fsspec.asyn, "reset_lock"):
# for future fsspec>2022.05.0
fsspec.asyn.reset_lock()
else:
fsspec.asyn.iothread[0] = None
fsspec.asyn.loop[0] = None
fsspec.asyn.lock = threading.Lock()
| 0
|
hf_public_repos/datasets/src/datasets
|
hf_public_repos/datasets/src/datasets/utils/beam_utils.py
|
import os
from apache_beam.io.filesystems import FileSystems
from apache_beam.pipeline import Pipeline
from .logging import get_logger
CHUNK_SIZE = 2 << 20 # 2mb
logger = get_logger(__name__)
class BeamPipeline(Pipeline):
"""Wrapper over `apache_beam.pipeline.Pipeline` for convenience"""
def is_local(self):
runner = self._options.get_all_options().get("runner")
return runner in [None, "DirectRunner", "PortableRunner"]
def upload_local_to_remote(local_file_path, remote_file_path, force_upload=False):
"""Use the Beam Filesystems to upload to a remote directory on gcs/s3/hdfs..."""
fs = FileSystems
if fs.exists(remote_file_path):
if force_upload:
logger.info(f"Remote path already exist: {remote_file_path}. Overwriting it as force_upload=True.")
else:
logger.info(f"Remote path already exist: {remote_file_path}. Skipping it as force_upload=False.")
return
with fs.create(remote_file_path) as remote_file:
with open(local_file_path, "rb") as local_file:
chunk = local_file.read(CHUNK_SIZE)
while chunk:
remote_file.write(chunk)
chunk = local_file.read(CHUNK_SIZE)
def download_remote_to_local(remote_file_path, local_file_path, force_download=False):
"""Use the Beam Filesystems to download from a remote directory on gcs/s3/hdfs..."""
fs = FileSystems
if os.path.exists(local_file_path):
if force_download:
logger.info(f"Local path already exist: {remote_file_path}. Overwriting it as force_upload=True.")
else:
logger.info(f"Local path already exist: {remote_file_path}. Skipping it as force_upload=False.")
return
with fs.open(remote_file_path) as remote_file:
with open(local_file_path, "wb") as local_file:
chunk = remote_file.read(CHUNK_SIZE)
while chunk:
local_file.write(chunk)
chunk = remote_file.read(CHUNK_SIZE)
| 0
|
hf_public_repos/datasets/src/datasets
|
hf_public_repos/datasets/src/datasets/utils/experimental.py
|
"""Contains utilities to flag a feature as "experimental" in datasets."""
import warnings
from functools import wraps
from typing import Callable
def experimental(fn: Callable) -> Callable:
"""Decorator to flag a feature as experimental.
An experimental feature trigger a warning when used as it might be subject to breaking changes in the future.
Args:
fn (`Callable`):
The function to flag as experimental.
Returns:
`Callable`: The decorated function.
Example:
```python
>>> from datasets.utils import experimental
>>> @experimental
... def my_function():
... print("Hello world!")
>>> my_function()
UserWarning: 'my_function' is experimental and might be subject to breaking changes in the future.
Hello world!
```
"""
@wraps(fn)
def _inner_fn(*args, **kwargs):
warnings.warn(
(f"'{fn.__name__}' is experimental and might be subject to breaking changes in the future."),
UserWarning,
)
return fn(*args, **kwargs)
return _inner_fn
| 0
|
hf_public_repos/datasets/src/datasets
|
hf_public_repos/datasets/src/datasets/utils/tf_utils.py
|
# Copyright 2022 The HuggingFace Datasets Authors and the TensorFlow Datasets Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""TF-specific utils import."""
import os
import warnings
from functools import partial
from math import ceil
from uuid import uuid4
import numpy as np
import pyarrow as pa
from multiprocess import get_context
try:
from multiprocess.shared_memory import SharedMemory
except ImportError:
SharedMemory = None # Version checks should prevent this being called on older Python versions
from .. import config
def minimal_tf_collate_fn(features):
if isinstance(features, dict): # case batch_size=None: nothing to collate
return features
elif config.TF_AVAILABLE:
import tensorflow as tf
else:
raise ImportError("Called a Tensorflow-specific function but Tensorflow is not installed.")
first = features[0]
batch = {}
for k, v in first.items():
if isinstance(v, np.ndarray):
batch[k] = np.stack([f[k] for f in features])
elif isinstance(v, tf.Tensor):
batch[k] = tf.stack([f[k] for f in features])
else:
batch[k] = np.array([f[k] for f in features])
return batch
def minimal_tf_collate_fn_with_renaming(features):
batch = minimal_tf_collate_fn(features)
if "label" in batch:
batch["labels"] = batch["label"]
del batch["label"]
return batch
def is_numeric_pa_type(pa_type):
if pa.types.is_list(pa_type):
return is_numeric_pa_type(pa_type.value_type)
return pa.types.is_integer(pa_type) or pa.types.is_floating(pa_type) or pa.types.is_decimal(pa_type)
def is_numeric_feature(feature):
from .. import ClassLabel, Sequence, Value
from ..features.features import _ArrayXD
if isinstance(feature, Sequence):
return is_numeric_feature(feature.feature)
elif isinstance(feature, list):
return is_numeric_feature(feature[0])
elif isinstance(feature, _ArrayXD):
return is_numeric_pa_type(feature().storage_dtype)
elif isinstance(feature, Value):
return is_numeric_pa_type(feature())
elif isinstance(feature, ClassLabel):
return True
else:
return False
def np_get_batch(
indices, dataset, cols_to_retain, collate_fn, collate_fn_args, columns_to_np_types, return_dict=False
):
if not isinstance(indices, np.ndarray):
indices = indices.numpy()
is_batched = True
# Optimization - if we're loading a sequential batch, do it with slicing instead of a list of indices
if isinstance(indices, np.integer):
batch = dataset[indices.item()]
is_batched = False
elif np.all(np.diff(indices) == 1):
batch = dataset[indices[0] : indices[-1] + 1]
elif isinstance(indices, np.ndarray):
batch = dataset[indices]
else:
raise RuntimeError("Unexpected type for indices: {}".format(type(indices)))
if cols_to_retain is not None:
batch = {
key: value
for key, value in batch.items()
if key in cols_to_retain or key in ("label", "label_ids", "labels")
}
if is_batched:
actual_size = len(list(batch.values())[0]) # Get the length of one of the arrays, assume all same
# Our collators expect a list of dicts, not a dict of lists/arrays, so we invert
batch = [{key: value[i] for key, value in batch.items()} for i in range(actual_size)]
batch = collate_fn(batch, **collate_fn_args)
if return_dict:
out_batch = {}
for col, cast_dtype in columns_to_np_types.items():
# In case the collate_fn returns something strange
array = np.array(batch[col])
array = array.astype(cast_dtype)
out_batch[col] = array
else:
out_batch = []
for col, cast_dtype in columns_to_np_types.items():
# In case the collate_fn returns something strange
array = np.array(batch[col])
array = array.astype(cast_dtype)
out_batch.append(array)
return out_batch
def dataset_to_tf(
dataset,
cols_to_retain,
collate_fn,
collate_fn_args,
columns_to_np_types,
output_signature,
shuffle,
batch_size,
drop_remainder,
):
"""Create a tf.data.Dataset from the underlying Dataset. This is a single-process method - the multiprocess
equivalent is multiprocess_dataset_to_tf.
Args:
dataset (`Dataset`): Dataset to wrap with tf.data.Dataset.
cols_to_retain (`List[str]`): Dataset column(s) to load in the
tf.data.Dataset. It is acceptable to include column names that are created by the `collate_fn` and
that do not exist in the original dataset.
collate_fn(`Callable`): A function or callable object (such as a `DataCollator`) that will collate
lists of samples into a batch.
collate_fn_args (`Dict`): A `dict` of keyword arguments to be passed to the
`collate_fn`. Can be empty.
columns_to_np_types (`Dict[str, np.dtype]`): A `dict` mapping column names to numpy dtypes.
output_signature (`Dict[str, tf.TensorSpec]`): A `dict` mapping column names to
`tf.TensorSpec` objects.
shuffle(`bool`): Shuffle the dataset order when loading. Recommended True for training, False for
validation/evaluation.
batch_size (`int`, default `None`): Size of batches to load from the dataset. Defaults to `None`, which implies that
the dataset won't be batched, but the returned dataset can be batched later with `tf_dataset.batch(batch_size)`.
drop_remainder(`bool`, default `None`): Drop the last incomplete batch when loading. If not provided,
defaults to the same setting as shuffle.
Returns:
`tf.data.Dataset`
"""
if config.TF_AVAILABLE:
import tensorflow as tf
else:
raise ImportError("Called a Tensorflow-specific function but Tensorflow is not installed.")
# TODO Matt: When our minimum Python version is 3.8 or higher, we can delete all of this and move everything
# to the NumPy multiprocessing path.
if hasattr(tf, "random_index_shuffle"):
random_index_shuffle = tf.random_index_shuffle
elif hasattr(tf.random.experimental, "index_shuffle"):
random_index_shuffle = tf.random.experimental.index_shuffle
else:
if len(dataset) > 10_000_000:
warnings.warn(
"to_tf_dataset() can be memory-inefficient on versions of TensorFlow older than 2.9. "
"If you are iterating over a dataset with a very large number of samples, consider "
"upgrading to TF >= 2.9."
)
random_index_shuffle = None
getter_fn = partial(
np_get_batch,
dataset=dataset,
cols_to_retain=cols_to_retain,
collate_fn=collate_fn,
collate_fn_args=collate_fn_args,
columns_to_np_types=columns_to_np_types,
return_dict=False,
)
# This works because dictionaries always output in the same order
tout = [tf.dtypes.as_dtype(dtype) for dtype in columns_to_np_types.values()]
@tf.function(input_signature=[tf.TensorSpec(None, tf.int64)])
def fetch_function(indices):
output = tf.py_function(
getter_fn,
inp=[indices],
Tout=tout,
)
return {key: output[i] for i, key in enumerate(columns_to_np_types.keys())}
tf_dataset = tf.data.Dataset.range(len(dataset))
if shuffle and random_index_shuffle is not None:
base_seed = tf.fill((3,), value=tf.cast(-1, dtype=tf.int64))
def scan_random_index(state, index):
if tf.reduce_all(state == -1):
# This generates a new random seed once per epoch only,
# to ensure that we iterate over each sample exactly once per epoch
state = tf.random.uniform(shape=(3,), maxval=2**62, dtype=tf.int64)
shuffled_index = random_index_shuffle(index=index, seed=state, max_index=len(dataset) - 1)
return state, shuffled_index
tf_dataset = tf_dataset.scan(base_seed, scan_random_index)
elif shuffle:
tf_dataset = tf_dataset.shuffle(tf_dataset.cardinality())
if batch_size is not None:
tf_dataset = tf_dataset.batch(batch_size, drop_remainder=drop_remainder)
tf_dataset = tf_dataset.map(fetch_function)
if batch_size is not None:
def ensure_shapes(input_dict):
return {key: tf.ensure_shape(val, output_signature[key].shape) for key, val in input_dict.items()}
else:
# Ensure shape but remove batch dimension of output_signature[key].shape
def ensure_shapes(input_dict):
return {key: tf.ensure_shape(val, output_signature[key].shape[1:]) for key, val in input_dict.items()}
return tf_dataset.map(ensure_shapes)
class SharedMemoryContext:
# This is a context manager for creating shared memory that ensures cleanup happens even if a process is interrupted
# The process that creates shared memory is always the one responsible for unlinking it in the end
def __init__(self):
self.created_shms = []
self.opened_shms = []
def get_shm(self, name, size, create):
shm = SharedMemory(size=int(size), name=name, create=create)
if create:
# We only unlink the ones we created in this context
self.created_shms.append(shm)
else:
# If we didn't create it, we only close it when done, we don't unlink it
self.opened_shms.append(shm)
return shm
def get_array(self, name, shape, dtype, create):
shm = self.get_shm(name=name, size=np.prod(shape) * np.dtype(dtype).itemsize, create=create)
return np.ndarray(shape, dtype=dtype, buffer=shm.buf)
def __enter__(self):
return self
def __exit__(self, exc_type, exc_value, traceback):
for shm in self.created_shms:
shm.close()
shm.unlink()
for shm in self.opened_shms:
shm.close()
class NumpyMultiprocessingGenerator:
def __init__(
self,
dataset,
cols_to_retain,
collate_fn,
collate_fn_args,
columns_to_np_types,
output_signature,
shuffle,
batch_size,
drop_remainder,
num_workers,
):
self.dataset = dataset
self.cols_to_retain = cols_to_retain
self.collate_fn = collate_fn
self.collate_fn_args = collate_fn_args
self.string_columns = [col for col, dtype in columns_to_np_types.items() if dtype in (np.unicode_, np.str_)]
# Strings will be converted to arrays of single unicode chars, so that we can have a constant itemsize
self.columns_to_np_types = {
col: dtype if col not in self.string_columns else np.dtype("U1")
for col, dtype in columns_to_np_types.items()
}
self.output_signature = output_signature
self.shuffle = shuffle
self.batch_size = batch_size
self.drop_remainder = drop_remainder
self.num_workers = num_workers
# Because strings are converted to characters, we need to add one extra dimension to the shape
self.columns_to_ranks = {
col: int(spec.shape.rank) if col not in self.string_columns else int(spec.shape.rank) + 1
for col, spec in output_signature.items()
}
def __iter__(self):
# Make sure we only spawn workers if they have work to do
num_workers = min(self.num_workers, int(ceil(len(self.dataset) / self.batch_size)))
# Do the shuffling in iter so that it's done at the start of each epoch
per_worker_batches, final_batch, final_batch_worker = self.distribute_batches(
self.dataset, self.batch_size, self.drop_remainder, num_workers, self.shuffle
)
ctx = get_context("spawn")
names = []
shape_arrays = []
workers = []
array_ready_events = [ctx.Event() for _ in range(num_workers)]
array_loaded_events = [ctx.Event() for _ in range(num_workers)]
base_args = {
"dataset": self.dataset,
"cols_to_retain": self.cols_to_retain,
"collate_fn": self.collate_fn,
"collate_fn_args": self.collate_fn_args,
"columns_to_np_types": self.columns_to_np_types,
"columns_to_ranks": self.columns_to_ranks,
"string_columns": self.string_columns,
}
with SharedMemoryContext() as shm_ctx:
for i in range(num_workers):
worker_random_id = str(uuid4())
worker_name = f"dw_{i}_{worker_random_id}"[:10]
names.append(worker_name)
worker_shape_arrays = {
col: shm_ctx.get_array(f"{worker_name}_{col}_shape", shape=(rank,), dtype=np.int64, create=True)
for col, rank in self.columns_to_ranks.items()
}
shape_arrays.append(worker_shape_arrays)
worker_indices = per_worker_batches[i]
if i == final_batch_worker and final_batch is not None:
final_batch_arg = final_batch
else:
final_batch_arg = None
worker_kwargs = {
"worker_name": worker_name,
"indices": worker_indices,
"extra_batch": final_batch_arg,
"array_ready_event": array_ready_events[i],
"array_loaded_event": array_loaded_events[i],
**base_args,
}
worker = ctx.Process(target=self.worker_loop, kwargs=worker_kwargs, daemon=True)
worker.start()
workers.append(worker)
end_signal_received = False
while not end_signal_received:
for i in range(num_workers):
if not array_ready_events[i].wait(timeout=60):
raise TimeoutError("Data loading worker timed out!")
array_ready_events[i].clear()
array_shapes = shape_arrays[i]
if any(np.any(shape < 0) for shape in array_shapes.values()):
# Child processes send negative array shapes to indicate
# that no more data is going to be sent
end_signal_received = True
break
# Matt: Because array shapes are variable we recreate the shared memory each iteration.
# I suspect repeatedly opening lots of shared memory is the bottleneck for the parent process.
# A future optimization, at the cost of some code complexity, could be to reuse shared memory
# between iterations, but this would require knowing in advance the maximum size, or having
# a system to only create a new memory block when a new maximum size is seen.
# Another potential optimization would be to figure out which memory copies are necessary,
# or whether we can yield objects straight out of shared memory.
with SharedMemoryContext() as batch_shm_ctx:
# This memory context only lasts long enough to copy everything out of the batch
arrays = {
col: batch_shm_ctx.get_array(
f"{names[i]}_{col}",
shape=shape,
dtype=self.columns_to_np_types[col],
create=False,
)
for col, shape in array_shapes.items()
}
# Copy everything out of shm because the memory
# will be unlinked by the child process at some point
arrays = {col: np.copy(arr) for col, arr in arrays.items()}
# Now we convert any unicode char arrays to strings
for string_col in self.string_columns:
arrays[string_col] = (
arrays[string_col].view(f"U{arrays[string_col].shape[-1]}").squeeze(-1)
)
yield arrays
array_loaded_events[i].set()
# Now we just do some cleanup
# Shared memory is cleaned up by the context manager, so we just make sure workers finish
for worker in workers:
worker.join()
def __call__(self):
return self
@staticmethod
def worker_loop(
dataset,
cols_to_retain,
collate_fn,
collate_fn_args,
columns_to_np_types,
columns_to_ranks,
string_columns,
indices,
extra_batch,
worker_name,
array_ready_event,
array_loaded_event,
):
os.environ["TF_CPP_MIN_LOG_LEVEL"] = "3"
if config.TF_AVAILABLE:
import tensorflow as tf
else:
raise ImportError("Called a Tensorflow-specific function but Tensorflow is not installed.")
tf.config.set_visible_devices([], "GPU") # Make sure workers don't try to allocate GPU memory
def send_batch_to_parent(indices):
batch = np_get_batch(
indices=indices,
dataset=dataset,
cols_to_retain=cols_to_retain,
collate_fn=collate_fn,
collate_fn_args=collate_fn_args,
columns_to_np_types=columns_to_np_types,
return_dict=True,
)
# Now begins the fun part where we start shovelling shared memory at the parent process
out_arrays = {}
with SharedMemoryContext() as batch_shm_ctx:
# The batch shared memory context exists only as long as it takes for the parent process
# to read everything, after which it cleans everything up again
for col, cast_dtype in columns_to_np_types.items():
# Everything has to be np.array for this to work, even if the collate_fn is giving us tf.Tensor
array = batch[col]
if col in string_columns:
# We can't send unicode arrays over shared memory, so we convert to single chars ("U1")
# which have a fixed width of 4 bytes. The parent process will convert these back to strings.
array = array.view("U1").reshape(array.shape + (-1,))
shape_arrays[col][:] = array.shape
out_arrays[col] = batch_shm_ctx.get_array(
f"{worker_name}_{col}", shape=array.shape, dtype=cast_dtype, create=True
)
out_arrays[col][:] = array
array_ready_event.set()
array_loaded_event.wait()
array_loaded_event.clear()
with SharedMemoryContext() as shm_ctx:
shape_arrays = {
col: shm_ctx.get_array(f"{worker_name}_{col}_shape", shape=(rank,), dtype=np.int64, create=False)
for col, rank in columns_to_ranks.items()
}
for batch in indices:
send_batch_to_parent(batch)
if extra_batch is not None:
send_batch_to_parent(extra_batch)
# Now we send a batsignal to the parent process that we're done
for col, array in shape_arrays.items():
array[:] = -1
array_ready_event.set()
@staticmethod
def distribute_batches(dataset, batch_size, drop_remainder, num_workers, shuffle):
indices = np.arange(len(dataset))
if shuffle:
np.random.shuffle(indices)
num_samples = len(indices)
# We distribute the batches so that reading from the workers in round-robin order yields the exact
# order specified in indices. This is only important when shuffle is False, but we do it regardless.
incomplete_batch_cutoff = num_samples - (num_samples % batch_size)
indices, last_incomplete_batch = np.split(indices, [incomplete_batch_cutoff])
if drop_remainder or len(last_incomplete_batch) == 0:
last_incomplete_batch = None
indices = indices.reshape(-1, batch_size)
num_batches = len(indices)
final_batches_cutoff = num_batches - (num_batches % num_workers)
indices, final_batches = np.split(indices, [final_batches_cutoff])
indices = indices.reshape(-1, num_workers, batch_size)
per_worker_indices = np.split(indices, indices.shape[1], axis=1)
per_worker_indices = [np.squeeze(worker_indices, 1) for worker_indices in per_worker_indices]
# Distribute the final batches to the first workers
for i in range(len(final_batches)):
# len(final_batches) can be zero, and is always less than num_workers
per_worker_indices[i] = np.concatenate([per_worker_indices[i], final_batches[i].reshape(1, -1)], axis=0)
# Add the last incomplete batch to the next worker, which might be the first worker
if last_incomplete_batch is not None:
incomplete_batch_worker_idx = len(final_batches)
else:
incomplete_batch_worker_idx = None
return per_worker_indices, last_incomplete_batch, incomplete_batch_worker_idx
def multiprocess_dataset_to_tf(
dataset,
cols_to_retain,
collate_fn,
collate_fn_args,
columns_to_np_types,
output_signature,
shuffle,
batch_size,
drop_remainder,
num_workers,
):
"""Create a tf.data.Dataset from the underlying Dataset. This is a multi-process method - the single-process
equivalent is dataset_to_tf.
Args:
dataset (`Dataset`): Dataset to wrap with tf.data.Dataset.
cols_to_retain (`List[str]`): Dataset column(s) to load in the
tf.data.Dataset. It is acceptable to include column names that are created by the `collate_fn` and
that do not exist in the original dataset.
collate_fn(`Callable`): A function or callable object (such as a `DataCollator`) that will collate
lists of samples into a batch.
collate_fn_args (`Dict`): A `dict` of keyword arguments to be passed to the
`collate_fn`. Can be empty.
columns_to_np_types (`Dict[str, np.dtype]`): A `dict` mapping column names to numpy dtypes.
output_signature (`Dict[str, tf.TensorSpec]`): A `dict` mapping column names to
`tf.TensorSpec` objects.
shuffle(`bool`): Shuffle the dataset order when loading. Recommended True for training, False for
validation/evaluation.
batch_size (`int`, default `None`): Size of batches to load from the dataset. Defaults to `None`, which implies that
the dataset won't be batched, but the returned dataset can be batched later with `tf_dataset.batch(batch_size)`.
drop_remainder(`bool`, default `None`): Drop the last incomplete batch when loading. If not provided,
defaults to the same setting as shuffle.
num_workers (`int`): Number of workers to use for loading the dataset. Should be >= 1.
Returns:
`tf.data.Dataset`
"""
if config.TF_AVAILABLE:
import tensorflow as tf
else:
raise ImportError("Called a Tensorflow-specific function but Tensorflow is not installed.")
data_generator = NumpyMultiprocessingGenerator(
dataset=dataset,
cols_to_retain=cols_to_retain,
collate_fn=collate_fn,
collate_fn_args=collate_fn_args,
columns_to_np_types=columns_to_np_types,
output_signature=output_signature,
shuffle=shuffle,
batch_size=batch_size,
drop_remainder=drop_remainder,
num_workers=num_workers,
)
tf_dataset = tf.data.Dataset.from_generator(data_generator, output_signature=output_signature)
if drop_remainder:
dataset_length = int(len(dataset) // batch_size)
else:
dataset_length = int(ceil(len(dataset) / batch_size))
return tf_dataset.apply(tf.data.experimental.assert_cardinality(dataset_length))
| 0
|
hf_public_repos/datasets/src/datasets
|
hf_public_repos/datasets/src/datasets/utils/py_utils.py
|
# Copyright 2020 The HuggingFace Datasets Authors and the TensorFlow Datasets Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# Lint as: python3
"""Some python utils function and classes.
"""
import copy
import functools
import itertools
import multiprocessing.pool
import os
import queue
import re
import types
import warnings
from contextlib import contextmanager
from dataclasses import fields, is_dataclass
from multiprocessing import Manager
from queue import Empty
from shutil import disk_usage
from typing import Any, Callable, Dict, Iterable, List, Optional, Set, Tuple, TypeVar, Union
from urllib.parse import urlparse
import multiprocess
import multiprocess.pool
import numpy as np
from tqdm.auto import tqdm
from .. import config
from ..parallel import parallel_map
from . import logging
from . import tqdm as hf_tqdm
from ._dill import ( # noqa: F401 # imported for backward compatibility. TODO: remove in 3.0.0
Pickler,
dump,
dumps,
pklregister,
)
try: # pragma: no branch
import typing_extensions as _typing_extensions
from typing_extensions import Final, Literal
except ImportError:
_typing_extensions = Literal = Final = None
logger = logging.get_logger(__name__)
# NOTE: When used on an instance method, the cache is shared across all
# instances and IS NOT per-instance.
# See
# https://stackoverflow.com/questions/14946264/python-lru-cache-decorator-per-instance
# For @property methods, use @memoized_property below.
memoize = functools.lru_cache
def size_str(size_in_bytes):
"""Returns a human readable size string.
If size_in_bytes is None, then returns "Unknown size".
For example `size_str(1.5 * datasets.units.GiB) == "1.50 GiB"`.
Args:
size_in_bytes: `int` or `None`, the size, in bytes, that we want to
format as a human-readable size string.
"""
if not size_in_bytes:
return "Unknown size"
_NAME_LIST = [("PiB", 2**50), ("TiB", 2**40), ("GiB", 2**30), ("MiB", 2**20), ("KiB", 2**10)]
size_in_bytes = float(size_in_bytes)
for name, size_bytes in _NAME_LIST:
value = size_in_bytes / size_bytes
if value >= 1.0:
return f"{value:.2f} {name}"
return f"{int(size_in_bytes)} bytes"
def convert_file_size_to_int(size: Union[int, str]) -> int:
"""
Converts a size expressed as a string with digits an unit (like `"50MB"`) to an integer (in bytes).
Args:
size (`int` or `str`): The size to convert. Will be directly returned if an `int`.
Example:
```py
>>> convert_file_size_to_int("1MiB")
1048576
```
"""
if isinstance(size, int):
return size
if size.upper().endswith("PIB"):
return int(size[:-3]) * (2**50)
if size.upper().endswith("TIB"):
return int(size[:-3]) * (2**40)
if size.upper().endswith("GIB"):
return int(size[:-3]) * (2**30)
if size.upper().endswith("MIB"):
return int(size[:-3]) * (2**20)
if size.upper().endswith("KIB"):
return int(size[:-3]) * (2**10)
if size.upper().endswith("PB"):
int_size = int(size[:-2]) * (10**15)
return int_size // 8 if size.endswith("b") else int_size
if size.upper().endswith("TB"):
int_size = int(size[:-2]) * (10**12)
return int_size // 8 if size.endswith("b") else int_size
if size.upper().endswith("GB"):
int_size = int(size[:-2]) * (10**9)
return int_size // 8 if size.endswith("b") else int_size
if size.upper().endswith("MB"):
int_size = int(size[:-2]) * (10**6)
return int_size // 8 if size.endswith("b") else int_size
if size.upper().endswith("KB"):
int_size = int(size[:-2]) * (10**3)
return int_size // 8 if size.endswith("b") else int_size
raise ValueError(f"`size={size}` is not in a valid format. Use an integer followed by the unit, e.g., '5GB'.")
def glob_pattern_to_regex(pattern):
# partially taken from fsspec:
# https://github.com/fsspec/filesystem_spec/blob/697d0f8133d8a5fbc3926e4761d7ecd51337ce50/fsspec/asyn.py#L735
return (
pattern.replace("\\", r"\\")
.replace(".", r"\.")
.replace("*", ".*")
.replace("+", r"\+")
.replace("//", "/")
.replace("(", r"\(")
.replace(")", r"\)")
.replace("|", r"\|")
.replace("^", r"\^")
.replace("$", r"\$")
.rstrip("/")
.replace("?", ".")
)
def string_to_dict(string: str, pattern: str) -> Dict[str, str]:
"""Un-format a string using a python f-string pattern.
From https://stackoverflow.com/a/36838374
Example::
>>> p = 'hello, my name is {name} and I am a {age} year old {what}'
>>> s = p.format(name='cody', age=18, what='quarterback')
>>> s
'hello, my name is cody and I am a 18 year old quarterback'
>>> string_to_dict(s, p)
{'age': '18', 'name': 'cody', 'what': 'quarterback'}
Args:
string (str): input string
pattern (str): pattern formatted like a python f-string
Returns:
Dict[str, str]: dictionary of variable -> value, retrieved from the input using the pattern
Raises:
ValueError: if the string doesn't match the pattern
"""
regex = re.sub(r"{(.+?)}", r"(?P<_\1>.+)", pattern)
result = re.search(regex, string)
if result is None:
raise ValueError(f"String {string} doesn't match the pattern {pattern}")
values = list(result.groups())
keys = re.findall(r"{(.+?)}", pattern)
_dict = dict(zip(keys, values))
return _dict
def asdict(obj):
"""Convert an object to its dictionary representation recursively.
<Added version="2.4.0"/>
"""
# Implementation based on https://docs.python.org/3/library/dataclasses.html#dataclasses.asdict
def _is_dataclass_instance(obj):
# https://docs.python.org/3/library/dataclasses.html#dataclasses.is_dataclass
return is_dataclass(obj) and not isinstance(obj, type)
def _asdict_inner(obj):
if _is_dataclass_instance(obj):
result = {}
for f in fields(obj):
value = _asdict_inner(getattr(obj, f.name))
if not f.init or value != f.default or f.metadata.get("include_in_asdict_even_if_is_default", False):
result[f.name] = value
return result
elif isinstance(obj, tuple) and hasattr(obj, "_fields"):
# obj is a namedtuple
return type(obj)(*[_asdict_inner(v) for v in obj])
elif isinstance(obj, (list, tuple)):
# Assume we can create an object of this type by passing in a
# generator (which is not true for namedtuples, handled
# above).
return type(obj)(_asdict_inner(v) for v in obj)
elif isinstance(obj, dict):
return {_asdict_inner(k): _asdict_inner(v) for k, v in obj.items()}
else:
return copy.deepcopy(obj)
if not isinstance(obj, dict) and not _is_dataclass_instance(obj):
raise TypeError(f"{obj} is not a dict or a dataclass")
return _asdict_inner(obj)
@contextmanager
def temporary_assignment(obj, attr, value):
"""Temporarily assign obj.attr to value."""
original = getattr(obj, attr, None)
setattr(obj, attr, value)
try:
yield
finally:
setattr(obj, attr, original)
@contextmanager
def temp_seed(seed: int, set_pytorch=False, set_tensorflow=False):
"""Temporarily set the random seed. This works for python numpy, pytorch and tensorflow."""
np_state = np.random.get_state()
np.random.seed(seed)
if set_pytorch and config.TORCH_AVAILABLE:
import torch
torch_state = torch.random.get_rng_state()
torch.random.manual_seed(seed)
if torch.cuda.is_available():
torch_cuda_states = torch.cuda.get_rng_state_all()
torch.cuda.manual_seed_all(seed)
if set_tensorflow and config.TF_AVAILABLE:
import tensorflow as tf
from tensorflow.python.eager import context as tfpycontext
tf_state = tf.random.get_global_generator()
temp_gen = tf.random.Generator.from_seed(seed)
tf.random.set_global_generator(temp_gen)
if not tf.executing_eagerly():
raise ValueError("Setting random seed for TensorFlow is only available in eager mode")
tf_context = tfpycontext.context() # eager mode context
tf_seed = tf_context._seed
tf_rng_initialized = hasattr(tf_context, "_rng")
if tf_rng_initialized:
tf_rng = tf_context._rng
tf_context._set_global_seed(seed)
try:
yield
finally:
np.random.set_state(np_state)
if set_pytorch and config.TORCH_AVAILABLE:
torch.random.set_rng_state(torch_state)
if torch.cuda.is_available():
torch.cuda.set_rng_state_all(torch_cuda_states)
if set_tensorflow and config.TF_AVAILABLE:
tf.random.set_global_generator(tf_state)
tf_context._seed = tf_seed
if tf_rng_initialized:
tf_context._rng = tf_rng
else:
delattr(tf_context, "_rng")
def unique_values(values):
"""Iterate over iterable and return only unique values in order."""
seen = set()
for value in values:
if value not in seen:
seen.add(value)
yield value
def no_op_if_value_is_null(func):
"""If the value is None, return None, else call `func`."""
def wrapper(value):
return func(value) if value is not None else None
return wrapper
def first_non_null_value(iterable):
"""Return the index and the value of the first non-null value in the iterable. If all values are None, return -1 as index."""
for i, value in enumerate(iterable):
if value is not None:
return i, value
return -1, None
def zip_dict(*dicts):
"""Iterate over items of dictionaries grouped by their keys."""
for key in unique_values(itertools.chain(*dicts)): # set merge all keys
# Will raise KeyError if the dict don't have the same keys
yield key, tuple(d[key] for d in dicts)
class NonMutableDict(dict):
"""Dict where keys can only be added but not modified.
Will raise an error if the user try to overwrite one key. The error message
can be customized during construction. It will be formatted using {key} for
the overwritten key.
"""
def __init__(self, *args, **kwargs):
self._error_msg = kwargs.pop(
"error_msg",
"Try to overwrite existing key: {key}",
)
if kwargs:
raise ValueError("NonMutableDict cannot be initialized with kwargs.")
super().__init__(*args, **kwargs)
def __setitem__(self, key, value):
if key in self:
raise ValueError(self._error_msg.format(key=key))
return super().__setitem__(key, value)
def update(self, other):
if any(k in self for k in other):
raise ValueError(self._error_msg.format(key=set(self) & set(other)))
return super().update(other)
class classproperty(property): # pylint: disable=invalid-name
"""Descriptor to be used as decorator for @classmethods."""
def __get__(self, obj, objtype=None):
return self.fget.__get__(None, objtype)()
def _single_map_nested(args):
"""Apply a function recursively to each element of a nested data struct."""
function, data_struct, types, rank, disable_tqdm, desc = args
# Singleton first to spare some computation
if not isinstance(data_struct, dict) and not isinstance(data_struct, types):
return function(data_struct)
# Reduce logging to keep things readable in multiprocessing with tqdm
if rank is not None and logging.get_verbosity() < logging.WARNING:
logging.set_verbosity_warning()
# Print at least one thing to fix tqdm in notebooks in multiprocessing
# see https://github.com/tqdm/tqdm/issues/485#issuecomment-473338308
if rank is not None and not disable_tqdm and any("notebook" in tqdm_cls.__name__ for tqdm_cls in tqdm.__mro__):
print(" ", end="", flush=True)
# Loop over single examples or batches and write to buffer/file if examples are to be updated
pbar_iterable = data_struct.items() if isinstance(data_struct, dict) else data_struct
pbar_desc = (desc + " " if desc is not None else "") + "#" + str(rank) if rank is not None else desc
with hf_tqdm(pbar_iterable, disable=disable_tqdm, position=rank, unit="obj", desc=pbar_desc) as pbar:
if isinstance(data_struct, dict):
return {k: _single_map_nested((function, v, types, None, True, None)) for k, v in pbar}
else:
mapped = [_single_map_nested((function, v, types, None, True, None)) for v in pbar]
if isinstance(data_struct, list):
return mapped
elif isinstance(data_struct, tuple):
return tuple(mapped)
else:
return np.array(mapped)
def map_nested(
function: Callable[[Any], Any],
data_struct: Any,
dict_only: bool = False,
map_list: bool = True,
map_tuple: bool = False,
map_numpy: bool = False,
num_proc: Optional[int] = None,
parallel_min_length: int = 2,
types: Optional[tuple] = None,
disable_tqdm: bool = True,
desc: Optional[str] = None,
) -> Any:
"""Apply a function recursively to each element of a nested data struct.
Use multiprocessing if num_proc > 1 and the length of data_struct is greater than or equal to
`parallel_min_length`.
<Changed version="2.5.0">
Before version 2.5.0, multiprocessing was not used if `num_proc` was greater than or equal to ``len(iterable)``.
Now, if `num_proc` is greater than or equal to ``len(iterable)``, `num_proc` is set to ``len(iterable)`` and
multiprocessing is used.
</Changed>
Args:
function (`Callable`): Function to be applied to `data_struct`.
data_struct (`Any`): Data structure to apply `function` to.
dict_only (`bool`, default `False`): Whether only apply `function` recursively to `dict` values in
`data_struct`.
map_list (`bool`, default `True`): Whether also apply `function` recursively to `list` elements (besides `dict`
values).
map_tuple (`bool`, default `False`): Whether also apply `function` recursively to `tuple` elements (besides
`dict` values).
map_numpy (`bool, default `False`): Whether also apply `function` recursively to `numpy.array` elements (besides
`dict` values).
num_proc (`int`, *optional*): Number of processes.
parallel_min_length (`int`, default `2`): Minimum length of `data_struct` required for parallel
processing.
<Added version="2.5.0"/>
types (`tuple`, *optional*): Additional types (besides `dict` values) to apply `function` recursively to their
elements.
disable_tqdm (`bool`, default `True`): Whether to disable the tqdm progressbar.
desc (`str`, *optional*): Prefix for the tqdm progressbar.
Returns:
`Any`
"""
if types is None:
types = []
if not dict_only:
if map_list:
types.append(list)
if map_tuple:
types.append(tuple)
if map_numpy:
types.append(np.ndarray)
types = tuple(types)
# Singleton
if not isinstance(data_struct, dict) and not isinstance(data_struct, types):
return function(data_struct)
iterable = list(data_struct.values()) if isinstance(data_struct, dict) else data_struct
if num_proc is None:
num_proc = 1
if num_proc != -1 and num_proc <= 1 or len(iterable) < parallel_min_length:
mapped = [
_single_map_nested((function, obj, types, None, True, None))
for obj in hf_tqdm(iterable, disable=disable_tqdm, desc=desc)
]
else:
with warnings.catch_warnings():
warnings.filterwarnings(
"ignore",
message=".* is experimental and might be subject to breaking changes in the future\\.$",
category=UserWarning,
)
mapped = parallel_map(function, iterable, num_proc, types, disable_tqdm, desc, _single_map_nested)
if isinstance(data_struct, dict):
return dict(zip(data_struct.keys(), mapped))
else:
if isinstance(data_struct, list):
return mapped
elif isinstance(data_struct, tuple):
return tuple(mapped)
else:
return np.array(mapped)
class NestedDataStructure:
def __init__(self, data=None):
self.data = data if data is not None else []
def flatten(self, data=None):
data = data if data is not None else self.data
if isinstance(data, dict):
return self.flatten(list(data.values()))
elif isinstance(data, (list, tuple)):
return [flattened for item in data for flattened in self.flatten(item)]
else:
return [data]
def has_sufficient_disk_space(needed_bytes, directory="."):
try:
free_bytes = disk_usage(os.path.abspath(directory)).free
except OSError:
return True
return needed_bytes < free_bytes
def _convert_github_url(url_path: str) -> Tuple[str, Optional[str]]:
"""Convert a link to a file on a github repo in a link to the raw github object."""
parsed = urlparse(url_path)
sub_directory = None
if parsed.scheme in ("http", "https", "s3") and parsed.netloc == "github.com":
if "blob" in url_path:
if not url_path.endswith(".py"):
raise ValueError(f"External import from github at {url_path} should point to a file ending with '.py'")
url_path = url_path.replace("blob", "raw") # Point to the raw file
else:
# Parse github url to point to zip
github_path = parsed.path[1:]
repo_info, branch = github_path.split("/tree/") if "/tree/" in github_path else (github_path, "master")
repo_owner, repo_name = repo_info.split("/")
url_path = f"https://github.com/{repo_owner}/{repo_name}/archive/{branch}.zip"
sub_directory = f"{repo_name}-{branch}"
return url_path, sub_directory
def get_imports(file_path: str) -> Tuple[str, str, str, str]:
"""Find whether we should import or clone additional files for a given processing script.
And list the import.
We allow:
- library dependencies,
- local dependencies and
- external dependencies whose url is specified with a comment starting from "# From:' followed by the raw url to a file, an archive or a github repository.
external dependencies will be downloaded (and extracted if needed in the dataset folder).
We also add an `__init__.py` to each sub-folder of a downloaded folder so the user can import from them in the script.
Note that only direct import in the dataset processing script will be handled
We don't recursively explore the additional import to download further files.
Example::
import tensorflow
import .c4_utils
import .clicr.dataset-code.build_json_dataset # From: https://raw.githubusercontent.com/clips/clicr/master/dataset-code/build_json_dataset
"""
lines = []
with open(file_path, encoding="utf-8") as f:
lines.extend(f.readlines())
logger.debug(f"Checking {file_path} for additional imports.")
imports: List[Tuple[str, str, str, Optional[str]]] = []
is_in_docstring = False
for line in lines:
docstr_start_match = re.findall(r'[\s\S]*?"""[\s\S]*?', line)
if len(docstr_start_match) == 1:
# flip True <=> False only if doctstring
# starts at line without finishing
is_in_docstring = not is_in_docstring
if is_in_docstring:
# import statements in doctstrings should
# not be added as required dependencies
continue
match = re.match(r"^import\s+(\.?)([^\s\.]+)[^#\r\n]*(?:#\s+From:\s+)?([^\r\n]*)", line, flags=re.MULTILINE)
if match is None:
match = re.match(
r"^from\s+(\.?)([^\s\.]+)(?:[^\s]*)\s+import\s+[^#\r\n]*(?:#\s+From:\s+)?([^\r\n]*)",
line,
flags=re.MULTILINE,
)
if match is None:
continue
if match.group(1):
# The import starts with a '.', we will download the relevant file
if any(imp[1] == match.group(2) for imp in imports):
# We already have this import
continue
if match.group(3):
# The import has a comment with 'From:', we'll retrieve it from the given url
url_path = match.group(3)
url_path, sub_directory = _convert_github_url(url_path)
imports.append(("external", match.group(2), url_path, sub_directory))
elif match.group(2):
# The import should be at the same place as the file
imports.append(("internal", match.group(2), match.group(2), None))
else:
if match.group(3):
# The import has a comment with `From: git+https:...`, asks user to pip install from git.
url_path = match.group(3)
imports.append(("library", match.group(2), url_path, None))
else:
imports.append(("library", match.group(2), match.group(2), None))
return imports
def copyfunc(func):
result = types.FunctionType(func.__code__, func.__globals__, func.__name__, func.__defaults__, func.__closure__)
result.__kwdefaults__ = func.__kwdefaults__
return result
Y = TypeVar("Y")
def _write_generator_to_queue(queue: queue.Queue, func: Callable[..., Iterable[Y]], kwargs: dict) -> int:
for i, result in enumerate(func(**kwargs)):
queue.put(result)
return i
def _get_pool_pid(pool: Union[multiprocessing.pool.Pool, multiprocess.pool.Pool]) -> Set[int]:
return {f.pid for f in pool._pool}
def iflatmap_unordered(
pool: Union[multiprocessing.pool.Pool, multiprocess.pool.Pool],
func: Callable[..., Iterable[Y]],
*,
kwargs_iterable: Iterable[dict],
) -> Iterable[Y]:
initial_pool_pid = _get_pool_pid(pool)
pool_changed = False
manager_cls = Manager if isinstance(pool, multiprocessing.pool.Pool) else multiprocess.Manager
with manager_cls() as manager:
queue = manager.Queue()
async_results = [
pool.apply_async(_write_generator_to_queue, (queue, func, kwargs)) for kwargs in kwargs_iterable
]
try:
while True:
try:
yield queue.get(timeout=0.05)
except Empty:
if all(async_result.ready() for async_result in async_results) and queue.empty():
break
if _get_pool_pid(pool) != initial_pool_pid:
pool_changed = True
# One of the subprocesses has died. We should not wait forever.
raise RuntimeError(
"One of the subprocesses has abruptly died during map operation."
"To debug the error, disable multiprocessing."
)
finally:
if not pool_changed:
# we get the result in case there's an error to raise
[async_result.get(timeout=0.05) for async_result in async_results]
| 0
|
hf_public_repos/datasets/src/datasets
|
hf_public_repos/datasets/src/datasets/utils/tqdm.py
|
"""Utility helpers to handle progress bars in `datasets`.
Example:
1. Use `datasets.utils.tqdm` as you would use `tqdm.tqdm` or `tqdm.auto.tqdm`.
2. To disable progress bars, either use `disable_progress_bars()` helper or set the
environment variable `HF_DATASETS_DISABLE_PROGRESS_BARS` to 1.
3. To re-enable progress bars, use `enable_progress_bars()`.
4. To check whether progress bars are disabled, use `are_progress_bars_disabled()`.
NOTE: Environment variable `HF_DATASETS_DISABLE_PROGRESS_BARS` has the priority.
Example:
```py
from datasets.utils import (
are_progress_bars_disabled,
disable_progress_bars,
enable_progress_bars,
tqdm,
)
# Disable progress bars globally
disable_progress_bars()
# Use as normal `tqdm`
for _ in tqdm(range(5)):
do_something()
# Still not showing progress bars, as `disable=False` is overwritten to `True`.
for _ in tqdm(range(5), disable=False):
do_something()
are_progress_bars_disabled() # True
# Re-enable progress bars globally
enable_progress_bars()
# Progress bar will be shown !
for _ in tqdm(range(5)):
do_something()
```
"""
import warnings
from tqdm.auto import tqdm as old_tqdm
from ..config import HF_DATASETS_DISABLE_PROGRESS_BARS
# `HF_DATASETS_DISABLE_PROGRESS_BARS` is `Optional[bool]` while `_hf_datasets_progress_bars_disabled`
# is a `bool`. If `HF_DATASETS_DISABLE_PROGRESS_BARS` is set to True or False, it has priority.
# If `HF_DATASETS_DISABLE_PROGRESS_BARS` is None, it means the user have not set the
# environment variable and is free to enable/disable progress bars programmatically.
# TL;DR: env variable has priority over code.
#
# By default, progress bars are enabled.
_hf_datasets_progress_bars_disabled: bool = HF_DATASETS_DISABLE_PROGRESS_BARS or False
def disable_progress_bars() -> None:
"""
Disable globally progress bars used in `datasets` except if `HF_DATASETS_DISABLE_PROGRESS_BAR` environment
variable has been set.
Use [`~utils.enable_progress_bars`] to re-enable them.
"""
if HF_DATASETS_DISABLE_PROGRESS_BARS is False:
warnings.warn(
"Cannot disable progress bars: environment variable `HF_DATASETS_DISABLE_PROGRESS_BAR=0` is set and has"
" priority."
)
return
global _hf_datasets_progress_bars_disabled
_hf_datasets_progress_bars_disabled = True
def enable_progress_bars() -> None:
"""
Enable globally progress bars used in `datasets` except if `HF_DATASETS_DISABLE_PROGRESS_BAR` environment
variable has been set.
Use [`~utils.disable_progress_bars`] to disable them.
"""
if HF_DATASETS_DISABLE_PROGRESS_BARS is True:
warnings.warn(
"Cannot enable progress bars: environment variable `HF_DATASETS_DISABLE_PROGRESS_BAR=1` is set and has"
" priority."
)
return
global _hf_datasets_progress_bars_disabled
_hf_datasets_progress_bars_disabled = False
def are_progress_bars_disabled() -> bool:
"""Return whether progress bars are globally disabled or not.
Progress bars used in `datasets` can be enable or disabled globally using [`~utils.enable_progress_bars`]
and [`~utils.disable_progress_bars`] or by setting `HF_DATASETS_DISABLE_PROGRESS_BAR` as environment variable.
"""
global _hf_datasets_progress_bars_disabled
return _hf_datasets_progress_bars_disabled
class tqdm(old_tqdm):
"""
Class to override `disable` argument in case progress bars are globally disabled.
Taken from https://github.com/tqdm/tqdm/issues/619#issuecomment-619639324.
"""
def __init__(self, *args, **kwargs):
if are_progress_bars_disabled():
kwargs["disable"] = True
super().__init__(*args, **kwargs)
def __delattr__(self, attr: str) -> None:
"""Fix for https://github.com/huggingface/datasets/issues/6066"""
try:
super().__delattr__(attr)
except AttributeError:
if attr != "_lock":
raise
# backward compatibility
enable_progress_bar = enable_progress_bars
disable_progress_bar = disable_progress_bars
def is_progress_bar_enabled():
return not are_progress_bars_disabled()
| 0
|
hf_public_repos/datasets/src/datasets
|
hf_public_repos/datasets/src/datasets/utils/_dill.py
|
# Copyright 2023 The HuggingFace Team. All rights reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""Extends `dill` to support pickling more types and produce more consistent dumps."""
import os
import sys
from io import BytesIO
from types import CodeType, FunctionType
import dill
from packaging import version
from .. import config
class Pickler(dill.Pickler):
dispatch = dill._dill.MetaCatchingDict(dill.Pickler.dispatch.copy())
def save(self, obj, save_persistent_id=True):
obj_type = type(obj)
if obj_type not in self.dispatch:
if "regex" in sys.modules:
import regex # type: ignore
if obj_type is regex.Pattern:
pklregister(obj_type)(_save_regexPattern)
if "spacy" in sys.modules:
import spacy # type: ignore
if issubclass(obj_type, spacy.Language):
pklregister(obj_type)(_save_spacyLanguage)
if "tiktoken" in sys.modules:
import tiktoken # type: ignore
if obj_type is tiktoken.Encoding:
pklregister(obj_type)(_save_tiktokenEncoding)
if "torch" in sys.modules:
import torch # type: ignore
if issubclass(obj_type, torch.Tensor):
pklregister(obj_type)(_save_torchTensor)
# Unwrap `torch.compile`-ed modules
if issubclass(obj_type, torch.nn.Module):
obj = getattr(obj, "_orig_mod", obj)
if "transformers" in sys.modules:
import transformers # type: ignore
if issubclass(obj_type, transformers.PreTrainedTokenizerBase):
pklregister(obj_type)(_save_transformersPreTrainedTokenizerBase)
# Unwrap `torch.compile`-ed functions
if obj_type is FunctionType:
obj = getattr(obj, "_torchdynamo_orig_callable", obj)
dill.Pickler.save(self, obj, save_persistent_id=save_persistent_id)
def _batch_setitems(self, items):
# Ignore the order of keys in a dict
try:
# Faster, but fails for unorderable elements
items = sorted(items)
except Exception: # TypeError, decimal.InvalidOperation, etc.
from datasets.fingerprint import Hasher
items = sorted(items, key=lambda x: Hasher.hash(x[0]))
dill.Pickler._batch_setitems(self, items)
def memoize(self, obj):
# Don't memoize strings since two identical strings can have different Python ids
if type(obj) is not str: # noqa: E721
dill.Pickler.memoize(self, obj)
def pklregister(t):
"""Register a custom reducer for the type."""
def proxy(func):
Pickler.dispatch[t] = func
return func
return proxy
def dump(obj, file):
"""Pickle an object to a file."""
Pickler(file, recurse=True).dump(obj)
def dumps(obj):
"""Pickle an object to a string."""
file = BytesIO()
dump(obj, file)
return file.getvalue()
if config.DILL_VERSION < version.parse("0.3.6"):
def log(pickler, msg):
dill._dill.log.info(msg)
elif config.DILL_VERSION.release[:3] in [version.parse("0.3.6").release, version.parse("0.3.7").release]:
def log(pickler, msg):
dill._dill.logger.trace(pickler, msg)
@pklregister(set)
def _save_set(pickler, obj):
log(pickler, f"Se: {obj}")
try:
# Faster, but fails for unorderable elements
args = (sorted(obj),)
except Exception: # TypeError, decimal.InvalidOperation, etc.
from datasets.fingerprint import Hasher
args = (sorted(obj, key=Hasher.hash),)
pickler.save_reduce(set, args, obj=obj)
log(pickler, "# Se")
def _save_regexPattern(pickler, obj):
import regex # type: ignore
log(pickler, f"Re: {obj}")
args = (obj.pattern, obj.flags)
pickler.save_reduce(regex.compile, args, obj=obj)
log(pickler, "# Re")
def _save_tiktokenEncoding(pickler, obj):
import tiktoken # type: ignore
log(pickler, f"Enc: {obj}")
args = (obj.name, obj._pat_str, obj._mergeable_ranks, obj._special_tokens)
pickler.save_reduce(tiktoken.Encoding, args, obj=obj)
log(pickler, "# Enc")
def _save_torchTensor(pickler, obj):
import torch # type: ignore
# `torch.from_numpy` is not picklable in `torch>=1.11.0`
def create_torchTensor(np_array):
return torch.from_numpy(np_array)
log(pickler, f"To: {obj}")
args = (obj.detach().cpu().numpy(),)
pickler.save_reduce(create_torchTensor, args, obj=obj)
log(pickler, "# To")
def _save_spacyLanguage(pickler, obj):
import spacy # type: ignore
def create_spacyLanguage(config, bytes):
lang_cls = spacy.util.get_lang_class(config["nlp"]["lang"])
lang_inst = lang_cls.from_config(config)
return lang_inst.from_bytes(bytes)
log(pickler, f"Sp: {obj}")
args = (obj.config, obj.to_bytes())
pickler.save_reduce(create_spacyLanguage, args, obj=obj)
log(pickler, "# Sp")
def _save_transformersPreTrainedTokenizerBase(pickler, obj):
log(pickler, f"Tok: {obj}")
# Ignore the `cache` attribute
state = obj.__dict__
if "cache" in state and isinstance(state["cache"], dict):
state["cache"] = {}
pickler.save_reduce(type(obj), (), state=state, obj=obj)
log(pickler, "# Tok")
if config.DILL_VERSION < version.parse("0.3.6"):
@pklregister(CodeType)
def _save_code(pickler, obj):
"""
From dill._dill.save_code
This is a modified version that removes the origin (filename + line no.)
of functions created in notebooks or shells for example.
"""
dill._dill.log.info(f"Co: {obj}")
# The filename of a function is the .py file where it is defined.
# Filenames of functions created in notebooks or shells start with '<'
# ex: <ipython-input-13-9ed2afe61d25> for ipython, and <stdin> for shell
# Filenames of functions created in ipykernel the filename
# look like f"{tempdir}/ipykernel_{id1}/{id2}.py"
# Moreover lambda functions have a special name: '<lambda>'
# ex: (lambda x: x).__code__.co_name == "<lambda>" # True
#
# For the hashing mechanism we ignore where the function has been defined
# More specifically:
# - we ignore the filename of special functions (filename starts with '<')
# - we always ignore the line number
# - we only use the base name of the file instead of the whole path,
# to be robust in case a script is moved for example.
#
# Only those two lines are different from the original implementation:
co_filename = (
""
if obj.co_filename.startswith("<")
or (
len(obj.co_filename.split(os.path.sep)) > 1
and obj.co_filename.split(os.path.sep)[-2].startswith("ipykernel_")
)
or obj.co_name == "<lambda>"
else os.path.basename(obj.co_filename)
)
co_firstlineno = 1
# The rest is the same as in the original dill implementation
if dill._dill.PY3:
if hasattr(obj, "co_posonlyargcount"):
args = (
obj.co_argcount,
obj.co_posonlyargcount,
obj.co_kwonlyargcount,
obj.co_nlocals,
obj.co_stacksize,
obj.co_flags,
obj.co_code,
obj.co_consts,
obj.co_names,
obj.co_varnames,
co_filename,
obj.co_name,
co_firstlineno,
obj.co_lnotab,
obj.co_freevars,
obj.co_cellvars,
)
else:
args = (
obj.co_argcount,
obj.co_kwonlyargcount,
obj.co_nlocals,
obj.co_stacksize,
obj.co_flags,
obj.co_code,
obj.co_consts,
obj.co_names,
obj.co_varnames,
co_filename,
obj.co_name,
co_firstlineno,
obj.co_lnotab,
obj.co_freevars,
obj.co_cellvars,
)
else:
args = (
obj.co_argcount,
obj.co_nlocals,
obj.co_stacksize,
obj.co_flags,
obj.co_code,
obj.co_consts,
obj.co_names,
obj.co_varnames,
co_filename,
obj.co_name,
co_firstlineno,
obj.co_lnotab,
obj.co_freevars,
obj.co_cellvars,
)
pickler.save_reduce(CodeType, args, obj=obj)
dill._dill.log.info("# Co")
return
elif config.DILL_VERSION.release[:3] in [version.parse("0.3.6").release, version.parse("0.3.7").release]:
# From: https://github.com/uqfoundation/dill/blob/dill-0.3.6/dill/_dill.py#L1104
@pklregister(CodeType)
def save_code(pickler, obj):
dill._dill.logger.trace(pickler, "Co: %s", obj)
############################################################################################################
# Modification here for huggingface/datasets
# The filename of a function is the .py file where it is defined.
# Filenames of functions created in notebooks or shells start with '<'
# ex: <ipython-input-13-9ed2afe61d25> for ipython, and <stdin> for shell
# Filenames of functions created in ipykernel the filename
# look like f"{tempdir}/ipykernel_{id1}/{id2}.py"
# Moreover lambda functions have a special name: '<lambda>'
# ex: (lambda x: x).__code__.co_name == "<lambda>" # True
#
# For the hashing mechanism we ignore where the function has been defined
# More specifically:
# - we ignore the filename of special functions (filename starts with '<')
# - we always ignore the line number
# - we only use the base name of the file instead of the whole path,
# to be robust in case a script is moved for example.
#
# Only those two lines are different from the original implementation:
co_filename = (
""
if obj.co_filename.startswith("<")
or (
len(obj.co_filename.split(os.path.sep)) > 1
and obj.co_filename.split(os.path.sep)[-2].startswith("ipykernel_")
)
or obj.co_name == "<lambda>"
else os.path.basename(obj.co_filename)
)
co_firstlineno = 1
# The rest is the same as in the original dill implementation, except for the replacements:
# - obj.co_filename => co_filename
# - obj.co_firstlineno => co_firstlineno
############################################################################################################
if hasattr(obj, "co_endlinetable"): # python 3.11a (20 args)
args = (
obj.co_lnotab, # for < python 3.10 [not counted in args]
obj.co_argcount,
obj.co_posonlyargcount,
obj.co_kwonlyargcount,
obj.co_nlocals,
obj.co_stacksize,
obj.co_flags,
obj.co_code,
obj.co_consts,
obj.co_names,
obj.co_varnames,
co_filename, # Modification for huggingface/datasets ############################################
obj.co_name,
obj.co_qualname,
co_firstlineno, # Modification for huggingface/datasets #########################################
obj.co_linetable,
obj.co_endlinetable,
obj.co_columntable,
obj.co_exceptiontable,
obj.co_freevars,
obj.co_cellvars,
)
elif hasattr(obj, "co_exceptiontable"): # python 3.11 (18 args)
args = (
obj.co_lnotab, # for < python 3.10 [not counted in args]
obj.co_argcount,
obj.co_posonlyargcount,
obj.co_kwonlyargcount,
obj.co_nlocals,
obj.co_stacksize,
obj.co_flags,
obj.co_code,
obj.co_consts,
obj.co_names,
obj.co_varnames,
co_filename, # Modification for huggingface/datasets ############################################
obj.co_name,
obj.co_qualname,
co_firstlineno, # Modification for huggingface/datasets #########################################
obj.co_linetable,
obj.co_exceptiontable,
obj.co_freevars,
obj.co_cellvars,
)
elif hasattr(obj, "co_linetable"): # python 3.10 (16 args)
args = (
obj.co_lnotab, # for < python 3.10 [not counted in args]
obj.co_argcount,
obj.co_posonlyargcount,
obj.co_kwonlyargcount,
obj.co_nlocals,
obj.co_stacksize,
obj.co_flags,
obj.co_code,
obj.co_consts,
obj.co_names,
obj.co_varnames,
co_filename, # Modification for huggingface/datasets ############################################
obj.co_name,
co_firstlineno, # Modification for huggingface/datasets #########################################
obj.co_linetable,
obj.co_freevars,
obj.co_cellvars,
)
elif hasattr(obj, "co_posonlyargcount"): # python 3.8 (16 args)
args = (
obj.co_argcount,
obj.co_posonlyargcount,
obj.co_kwonlyargcount,
obj.co_nlocals,
obj.co_stacksize,
obj.co_flags,
obj.co_code,
obj.co_consts,
obj.co_names,
obj.co_varnames,
co_filename, # Modification for huggingface/datasets ############################################
obj.co_name,
co_firstlineno, # Modification for huggingface/datasets #########################################
obj.co_lnotab,
obj.co_freevars,
obj.co_cellvars,
)
else: # python 3.7 (15 args)
args = (
obj.co_argcount,
obj.co_kwonlyargcount,
obj.co_nlocals,
obj.co_stacksize,
obj.co_flags,
obj.co_code,
obj.co_consts,
obj.co_names,
obj.co_varnames,
co_filename, # Modification for huggingface/datasets ############################################
obj.co_name,
co_firstlineno, # Modification for huggingface/datasets #########################################
obj.co_lnotab,
obj.co_freevars,
obj.co_cellvars,
)
pickler.save_reduce(dill._dill._create_code, args, obj=obj)
dill._dill.logger.trace(pickler, "# Co")
return
| 0
|
hf_public_repos/datasets/src/datasets
|
hf_public_repos/datasets/src/datasets/utils/stratify.py
|
import numpy as np
def approximate_mode(class_counts, n_draws, rng):
"""Computes approximate mode of multivariate hypergeometric.
This is an approximation to the mode of the multivariate
hypergeometric given by class_counts and n_draws.
It shouldn't be off by more than one.
It is the mostly likely outcome of drawing n_draws many
samples from the population given by class_counts.
Args
----------
class_counts : ndarray of int
Population per class.
n_draws : int
Number of draws (samples to draw) from the overall population.
rng : random state
Used to break ties.
Returns
-------
sampled_classes : ndarray of int
Number of samples drawn from each class.
np.sum(sampled_classes) == n_draws
"""
# this computes a bad approximation to the mode of the
# multivariate hypergeometric given by class_counts and n_draws
continuous = n_draws * class_counts / class_counts.sum()
# floored means we don't overshoot n_samples, but probably undershoot
floored = np.floor(continuous)
# we add samples according to how much "left over" probability
# they had, until we arrive at n_samples
need_to_add = int(n_draws - floored.sum())
if need_to_add > 0:
remainder = continuous - floored
values = np.sort(np.unique(remainder))[::-1]
# add according to remainder, but break ties
# randomly to avoid biases
for value in values:
(inds,) = np.where(remainder == value)
# if we need_to_add less than what's in inds
# we draw randomly from them.
# if we need to add more, we add them all and
# go to the next value
add_now = min(len(inds), need_to_add)
inds = rng.choice(inds, size=add_now, replace=False)
floored[inds] += 1
need_to_add -= add_now
if need_to_add == 0:
break
return floored.astype(np.int64)
def stratified_shuffle_split_generate_indices(y, n_train, n_test, rng, n_splits=10):
"""
Provides train/test indices to split data in train/test sets.
It's reference is taken from StratifiedShuffleSplit implementation
of scikit-learn library.
Args
----------
n_train : int,
represents the absolute number of train samples.
n_test : int,
represents the absolute number of test samples.
random_state : int or RandomState instance, default=None
Controls the randomness of the training and testing indices produced.
Pass an int for reproducible output across multiple function calls.
n_splits : int, default=10
Number of re-shuffling & splitting iterations.
"""
classes, y_indices = np.unique(y, return_inverse=True)
n_classes = classes.shape[0]
class_counts = np.bincount(y_indices)
if np.min(class_counts) < 2:
raise ValueError("Minimum class count error")
if n_train < n_classes:
raise ValueError(
"The train_size = %d should be greater or " "equal to the number of classes = %d" % (n_train, n_classes)
)
if n_test < n_classes:
raise ValueError(
"The test_size = %d should be greater or " "equal to the number of classes = %d" % (n_test, n_classes)
)
class_indices = np.split(np.argsort(y_indices, kind="mergesort"), np.cumsum(class_counts)[:-1])
for _ in range(n_splits):
n_i = approximate_mode(class_counts, n_train, rng)
class_counts_remaining = class_counts - n_i
t_i = approximate_mode(class_counts_remaining, n_test, rng)
train = []
test = []
for i in range(n_classes):
permutation = rng.permutation(class_counts[i])
perm_indices_class_i = class_indices[i].take(permutation, mode="clip")
train.extend(perm_indices_class_i[: n_i[i]])
test.extend(perm_indices_class_i[n_i[i] : n_i[i] + t_i[i]])
train = rng.permutation(train)
test = rng.permutation(test)
yield train, test
| 0
|
hf_public_repos/datasets/src/datasets
|
hf_public_repos/datasets/src/datasets/utils/hub.py
|
from typing import Optional
from urllib.parse import quote
import huggingface_hub as hfh
from packaging import version
def hf_hub_url(repo_id: str, path: str, revision: Optional[str] = None) -> str:
if version.parse(hfh.__version__).release < version.parse("0.11.0").release:
# old versions of hfh don't url-encode the file path
path = quote(path)
return hfh.hf_hub_url(repo_id, path, repo_type="dataset", revision=revision)
| 0
|
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.