repo_id
stringlengths 15
89
| file_path
stringlengths 27
180
| content
stringlengths 1
2.23M
| __index_level_0__
int64 0
0
|
|---|---|---|---|
hf_public_repos/datasets
|
hf_public_repos/datasets/tests/test_experimental.py
|
import unittest
import warnings
from datasets.utils import experimental
@experimental
def dummy_function():
return "success"
class TestExperimentalFlag(unittest.TestCase):
def test_experimental_warning(self):
with warnings.catch_warnings(record=True) as w:
warnings.simplefilter("always")
self.assertEqual(dummy_function(), "success")
self.assertEqual(len(w), 1)
| 0
|
hf_public_repos/datasets
|
hf_public_repos/datasets/tests/test_distributed.py
|
import os
import sys
from pathlib import Path
import pytest
from datasets import Dataset, IterableDataset
from datasets.distributed import split_dataset_by_node
from .utils import execute_subprocess_async, get_torch_dist_unique_port, require_torch
def test_split_dataset_by_node_map_style():
full_ds = Dataset.from_dict({"i": range(17)})
full_size = len(full_ds)
world_size = 3
datasets_per_rank = [
split_dataset_by_node(full_ds, rank=rank, world_size=world_size) for rank in range(world_size)
]
assert sum(len(ds) for ds in datasets_per_rank) == full_size
assert len({tuple(x.values()) for ds in datasets_per_rank for x in ds}) == full_size
def test_split_dataset_by_node_iterable():
def gen():
return ({"i": i} for i in range(17))
world_size = 3
full_ds = IterableDataset.from_generator(gen)
full_size = len(list(full_ds))
datasets_per_rank = [
split_dataset_by_node(full_ds, rank=rank, world_size=world_size) for rank in range(world_size)
]
assert sum(len(list(ds)) for ds in datasets_per_rank) == full_size
assert len({tuple(x.values()) for ds in datasets_per_rank for x in ds}) == full_size
@pytest.mark.parametrize("shards_per_node", [1, 2, 3])
def test_split_dataset_by_node_iterable_sharded(shards_per_node):
def gen(shards):
for shard in shards:
yield from ({"i": i, "shard": shard} for i in range(17))
world_size = 3
num_shards = shards_per_node * world_size
gen_kwargs = {"shards": [f"shard_{shard_idx}.txt" for shard_idx in range(num_shards)]}
full_ds = IterableDataset.from_generator(gen, gen_kwargs=gen_kwargs)
full_size = len(list(full_ds))
assert full_ds.n_shards == world_size * shards_per_node
datasets_per_rank = [
split_dataset_by_node(full_ds, rank=rank, world_size=world_size) for rank in range(world_size)
]
assert [ds.n_shards for ds in datasets_per_rank] == [shards_per_node] * world_size
assert sum(len(list(ds)) for ds in datasets_per_rank) == full_size
assert len({tuple(x.values()) for ds in datasets_per_rank for x in ds}) == full_size
def test_distributed_shuffle_iterable():
def gen():
return ({"i": i} for i in range(17))
world_size = 2
full_ds = IterableDataset.from_generator(gen)
full_size = len(list(full_ds))
ds_rank0 = split_dataset_by_node(full_ds, rank=0, world_size=world_size).shuffle(seed=42)
assert len(list(ds_rank0)) == 1 + full_size // world_size
with pytest.raises(RuntimeError):
split_dataset_by_node(full_ds, rank=0, world_size=world_size).shuffle()
ds_rank0 = split_dataset_by_node(full_ds.shuffle(seed=42), rank=0, world_size=world_size)
assert len(list(ds_rank0)) == 1 + full_size // world_size
with pytest.raises(RuntimeError):
split_dataset_by_node(full_ds.shuffle(), rank=0, world_size=world_size)
@pytest.mark.parametrize("streaming", [False, True])
@require_torch
@pytest.mark.skipif(os.name == "nt", reason="execute_subprocess_async doesn't support windows")
@pytest.mark.integration
def test_torch_distributed_run(streaming):
nproc_per_node = 2
master_port = get_torch_dist_unique_port()
test_script = Path(__file__).resolve().parent / "distributed_scripts" / "run_torch_distributed.py"
distributed_args = f"""
-m torch.distributed.run
--nproc_per_node={nproc_per_node}
--master_port={master_port}
{test_script}
""".split()
args = f"""
--streaming={streaming}
""".split()
cmd = [sys.executable] + distributed_args + args
execute_subprocess_async(cmd, env=os.environ.copy())
@pytest.mark.parametrize(
"nproc_per_node, num_workers",
[
(2, 2), # each node has 2 shards and each worker has 1 shards
(3, 2), # each node uses all the shards but skips examples, and each worker has 2 shards
],
)
@require_torch
@pytest.mark.skipif(os.name == "nt", reason="execute_subprocess_async doesn't support windows")
@pytest.mark.integration
def test_torch_distributed_run_streaming_with_num_workers(nproc_per_node, num_workers):
streaming = True
master_port = get_torch_dist_unique_port()
test_script = Path(__file__).resolve().parent / "distributed_scripts" / "run_torch_distributed.py"
distributed_args = f"""
-m torch.distributed.run
--nproc_per_node={nproc_per_node}
--master_port={master_port}
{test_script}
""".split()
args = f"""
--streaming={streaming}
--num_workers={num_workers}
""".split()
cmd = [sys.executable] + distributed_args + args
execute_subprocess_async(cmd, env=os.environ.copy())
| 0
|
hf_public_repos/datasets
|
hf_public_repos/datasets/tests/test_iterable_dataset.py
|
import pickle
from copy import deepcopy
from itertools import chain, islice
import numpy as np
import pandas as pd
import pyarrow as pa
import pyarrow.compute as pc
import pytest
from datasets import Dataset, load_dataset
from datasets.combine import concatenate_datasets, interleave_datasets
from datasets.features import (
ClassLabel,
Features,
Image,
Value,
)
from datasets.formatting import get_format_type_from_alias
from datasets.info import DatasetInfo
from datasets.iterable_dataset import (
ArrowExamplesIterable,
BufferShuffledExamplesIterable,
CyclingMultiSourcesExamplesIterable,
ExamplesIterable,
FilteredExamplesIterable,
FormattingConfig,
HorizontallyConcatenatedMultiSourcesExamplesIterable,
IterableDataset,
MappedExamplesIterable,
RandomlyCyclingMultiSourcesExamplesIterable,
SelectColumnsIterable,
ShuffledDataSourcesArrowExamplesIterable,
ShuffledDataSourcesExamplesIterable,
ShufflingConfig,
SkipExamplesIterable,
StepExamplesIterable,
TakeExamplesIterable,
TypedExamplesIterable,
VerticallyConcatenatedMultiSourcesExamplesIterable,
_BaseExamplesIterable,
_batch_arrow_tables,
_batch_to_examples,
_convert_to_arrow,
_examples_to_batch,
)
from .utils import (
assert_arrow_memory_doesnt_increase,
is_rng_equal,
require_dill_gt_0_3_2,
require_not_windows,
require_pyspark,
require_tf,
require_torch,
)
DEFAULT_N_EXAMPLES = 20
DEFAULT_BATCH_SIZE = 4
DEFAULT_FILEPATH = "file.txt"
SAMPLE_DATASET_IDENTIFIER = "hf-internal-testing/dataset_with_script" # has dataset script
def generate_examples_fn(**kwargs):
kwargs = kwargs.copy()
n = kwargs.pop("n", DEFAULT_N_EXAMPLES)
filepaths = kwargs.pop("filepaths", None)
for filepath in filepaths or [DEFAULT_FILEPATH]:
if filepaths is not None:
kwargs["filepath"] = filepath
for i in range(n):
yield f"{filepath}_{i}", {"id": i, **kwargs}
def generate_tables_fn(**kwargs):
kwargs = kwargs.copy()
n = kwargs.pop("n", DEFAULT_N_EXAMPLES)
batch_size = kwargs.pop("batch_size", DEFAULT_BATCH_SIZE)
filepaths = kwargs.pop("filepaths", None)
for filepath in filepaths or [DEFAULT_FILEPATH]:
buffer = []
batch_idx = 0
if filepaths is not None:
kwargs["filepath"] = filepath
for i in range(n):
buffer.append({"id": i, **kwargs})
if len(buffer) == batch_size:
yield f"{filepath}_{batch_idx}", pa.Table.from_pylist(buffer)
buffer = []
batch_idx += 1
yield batch_idx, pa.Table.from_pylist(buffer)
@pytest.fixture
def dataset():
ex_iterable = ExamplesIterable(generate_examples_fn, {})
return IterableDataset(ex_iterable, info=DatasetInfo(description="dummy"), split="train")
@pytest.fixture
def dataset_with_several_columns():
ex_iterable = ExamplesIterable(
generate_examples_fn,
{"filepath": ["data0.txt", "data1.txt", "data2.txt"], "metadata": {"sources": ["https://foo.bar"]}},
)
return IterableDataset(ex_iterable, info=DatasetInfo(description="dummy"), split="train")
@pytest.fixture
def arrow_file(tmp_path_factory, dataset: IterableDataset):
filename = str(tmp_path_factory.mktemp("data") / "file.arrow")
Dataset.from_generator(dataset.__iter__).map(cache_file_name=filename)
return filename
################################
#
# Utilities tests
#
################################
@pytest.mark.parametrize("batch_size", [1, 2, 3, 9, 10, 11, 20])
@pytest.mark.parametrize("drop_last_batch", [False, True])
def test_convert_to_arrow(batch_size, drop_last_batch):
examples = [{"foo": i} for i in range(10)]
full_table = pa.Table.from_pylist(examples)
num_rows = len(full_table) if not drop_last_batch else len(full_table) // batch_size * batch_size
num_batches = (num_rows // batch_size) + 1 if num_rows % batch_size else num_rows // batch_size
subtables = list(
_convert_to_arrow(
list(enumerate(examples)),
batch_size=batch_size,
drop_last_batch=drop_last_batch,
)
)
assert len(subtables) == num_batches
if drop_last_batch:
assert all(len(subtable) == batch_size for _, subtable in subtables)
else:
assert all(len(subtable) == batch_size for _, subtable in subtables[:-1])
assert len(subtables[-1][1]) <= batch_size
if num_rows > 0:
reloaded = pa.concat_tables([subtable for _, subtable in subtables])
assert full_table.slice(0, num_rows).to_pydict() == reloaded.to_pydict()
@pytest.mark.parametrize(
"tables",
[
[pa.table({"foo": range(10)})],
[pa.table({"foo": range(0, 5)}), pa.table({"foo": range(5, 10)})],
[pa.table({"foo": [i]}) for i in range(10)],
],
)
@pytest.mark.parametrize("batch_size", [1, 2, 3, 9, 10, 11, 20])
@pytest.mark.parametrize("drop_last_batch", [False, True])
def test_batch_arrow_tables(tables, batch_size, drop_last_batch):
full_table = pa.concat_tables(tables)
num_rows = len(full_table) if not drop_last_batch else len(full_table) // batch_size * batch_size
num_batches = (num_rows // batch_size) + 1 if num_rows % batch_size else num_rows // batch_size
subtables = list(
_batch_arrow_tables(list(enumerate(tables)), batch_size=batch_size, drop_last_batch=drop_last_batch)
)
assert len(subtables) == num_batches
if drop_last_batch:
assert all(len(subtable) == batch_size for _, subtable in subtables)
else:
assert all(len(subtable) == batch_size for _, subtable in subtables[:-1])
assert len(subtables[-1][1]) <= batch_size
if num_rows > 0:
reloaded = pa.concat_tables([subtable for _, subtable in subtables])
assert full_table.slice(0, num_rows).to_pydict() == reloaded.to_pydict()
################################
#
# _BaseExampleIterable tests
#
################################
def test_examples_iterable():
ex_iterable = ExamplesIterable(generate_examples_fn, {})
expected = list(generate_examples_fn())
assert next(iter(ex_iterable)) == expected[0]
assert list(ex_iterable) == expected
assert ex_iterable.iter_arrow is None
def test_examples_iterable_with_kwargs():
ex_iterable = ExamplesIterable(generate_examples_fn, {"filepaths": ["0.txt", "1.txt"], "split": "train"})
expected = list(generate_examples_fn(filepaths=["0.txt", "1.txt"], split="train"))
assert list(ex_iterable) == expected
assert all("split" in ex for _, ex in ex_iterable)
assert sorted({ex["filepath"] for _, ex in ex_iterable}) == ["0.txt", "1.txt"]
def test_examples_iterable_shuffle_data_sources():
ex_iterable = ExamplesIterable(generate_examples_fn, {"filepaths": ["0.txt", "1.txt"]})
ex_iterable = ex_iterable.shuffle_data_sources(np.random.default_rng(40))
expected = list(generate_examples_fn(filepaths=["1.txt", "0.txt"])) # shuffle the filepaths
assert list(ex_iterable) == expected
def test_examples_iterable_shuffle_shards_and_metadata():
def gen(filepaths, all_metadata):
for i, (filepath, metadata) in enumerate(zip(filepaths, all_metadata)):
yield i, {"filepath": filepath, "metadata": metadata}
ex_iterable = ExamplesIterable(
gen,
{
"filepaths": [f"{i}.txt" for i in range(100)],
"all_metadata": [{"id": str(i)} for i in range(100)],
},
)
ex_iterable = ex_iterable.shuffle_data_sources(np.random.default_rng(42))
out = list(ex_iterable)
filepaths_ids = [x["filepath"].split(".")[0] for _, x in out]
metadata_ids = [x["metadata"]["id"] for _, x in out]
assert filepaths_ids == metadata_ids, "entangled lists of shards/metadata should be shuffled the same way"
def test_arrow_examples_iterable():
ex_iterable = ArrowExamplesIterable(generate_tables_fn, {})
expected = sum([pa_table.to_pylist() for _, pa_table in generate_tables_fn()], [])
assert next(iter(ex_iterable))[1] == expected[0]
assert [example for _, example in ex_iterable] == expected
expected = list(generate_tables_fn())
assert list(ex_iterable.iter_arrow()) == expected
def test_arrow_examples_iterable_with_kwargs():
ex_iterable = ArrowExamplesIterable(generate_tables_fn, {"filepaths": ["0.txt", "1.txt"], "split": "train"})
expected = sum(
[pa_table.to_pylist() for _, pa_table in generate_tables_fn(filepaths=["0.txt", "1.txt"], split="train")], []
)
assert [example for _, example in ex_iterable] == expected
assert all("split" in ex for _, ex in ex_iterable)
assert sorted({ex["filepath"] for _, ex in ex_iterable}) == ["0.txt", "1.txt"]
expected = list(generate_tables_fn(filepaths=["0.txt", "1.txt"], split="train"))
assert list(ex_iterable.iter_arrow()) == expected
def test_arrow_examples_iterable_shuffle_data_sources():
ex_iterable = ArrowExamplesIterable(generate_tables_fn, {"filepaths": ["0.txt", "1.txt"]})
ex_iterable = ex_iterable.shuffle_data_sources(np.random.default_rng(40))
expected = sum(
[pa_table.to_pylist() for _, pa_table in generate_tables_fn(filepaths=["1.txt", "0.txt"])], []
) # shuffle the filepaths
assert [example for _, example in ex_iterable] == expected
expected = list(generate_tables_fn(filepaths=["1.txt", "0.txt"]))
assert list(ex_iterable.iter_arrow()) == expected
@pytest.mark.parametrize("seed", [42, 1337, 101010, 123456])
def test_buffer_shuffled_examples_iterable(seed):
n, buffer_size = 100, 30
generator = np.random.default_rng(seed)
base_ex_iterable = ExamplesIterable(generate_examples_fn, {"n": n})
ex_iterable = BufferShuffledExamplesIterable(base_ex_iterable, buffer_size=buffer_size, generator=generator)
rng = deepcopy(generator)
expected_indices_used_for_shuffling = list(
islice(BufferShuffledExamplesIterable._iter_random_indices(rng, buffer_size=buffer_size), n - buffer_size)
)
# indices to pick in the shuffle buffer should all be in the right range
assert all(0 <= index_to_pick < buffer_size for index_to_pick in expected_indices_used_for_shuffling)
# it should be random indices
assert expected_indices_used_for_shuffling != list(range(buffer_size))
# The final order of examples is the result of a shuffle buffer.
all_examples = list(generate_examples_fn(n=n))
# We create a buffer and we pick random examples from it.
buffer, rest = all_examples[:buffer_size], all_examples[buffer_size:]
expected = []
for i, index_to_pick in enumerate(expected_indices_used_for_shuffling):
expected.append(buffer[index_to_pick])
# The picked examples are directly replaced by the next examples from the iterable.
buffer[index_to_pick] = rest.pop(0)
# Once we have reached the end of the iterable, we shuffle the buffer and return the remaining examples.
rng.shuffle(buffer)
expected += buffer
assert next(iter(ex_iterable)) == expected[0]
assert list(ex_iterable) == expected
assert sorted(ex_iterable) == sorted(all_examples)
def test_cycling_multi_sources_examples_iterable():
ex_iterable1 = ExamplesIterable(generate_examples_fn, {"text": "foo"})
ex_iterable2 = ExamplesIterable(generate_examples_fn, {"text": "bar"})
ex_iterable = CyclingMultiSourcesExamplesIterable([ex_iterable1, ex_iterable2])
expected = list(chain(*zip(generate_examples_fn(text="foo"), generate_examples_fn(text="bar"))))
# The cycling stops as soon as one iterable is out of examples (here ex_iterable1), so the last sample from ex_iterable2 is unecessary
expected = expected[:-1]
assert next(iter(ex_iterable)) == expected[0]
assert list(ex_iterable) == expected
assert all((x["id"], x["text"]) == (i // 2, "bar" if i % 2 else "foo") for i, (_, x) in enumerate(ex_iterable))
@pytest.mark.parametrize("probabilities", [None, (0.5, 0.5), (0.9, 0.1)])
def test_randomly_cycling_multi_sources_examples_iterable(probabilities):
seed = 42
generator = np.random.default_rng(seed)
ex_iterable1 = ExamplesIterable(generate_examples_fn, {"text": "foo"})
ex_iterable2 = ExamplesIterable(generate_examples_fn, {"text": "bar"})
ex_iterable = RandomlyCyclingMultiSourcesExamplesIterable(
[ex_iterable1, ex_iterable2], generator=generator, probabilities=probabilities
)
# The source used randomly changes at each example. It stops when one of the iterators is empty.
rng = deepcopy(generator)
iterators = (generate_examples_fn(text="foo"), generate_examples_fn(text="bar"))
indices_iterator = RandomlyCyclingMultiSourcesExamplesIterable._iter_random_indices(
rng, len(iterators), p=probabilities
)
expected = []
lengths = [len(list(ex_iterable1)), len(list(ex_iterable2))]
for i in indices_iterator:
if lengths[0] == 0 or lengths[1] == 0:
break
for key, example in iterators[i]:
expected.append((key, example))
lengths[i] -= 1
break
else:
break
assert next(iter(ex_iterable)) == expected[0]
assert list(ex_iterable) == expected
@pytest.mark.parametrize(
"n, func, batched, batch_size",
[
(3, lambda x: {"id+1": x["id"] + 1}, False, None), # just add 1 to the id
(3, lambda x: {"id+1": [x["id"][0] + 1]}, True, 1), # same with bs=1
(5, lambda x: {"id+1": [i + 1 for i in x["id"]]}, True, 10), # same with bs=10
(25, lambda x: {"id+1": [i + 1 for i in x["id"]]}, True, 10), # same with bs=10
(5, lambda x: {"id+1": [i + 1 for i in x["id"]]}, True, None), # same with bs=None
(5, lambda x: {"id+1": [i + 1 for i in x["id"]]}, True, -1), # same with bs<=0
(3, lambda x: {k: v * 2 for k, v in x.items()}, True, 1), # make a duplicate of each example
],
)
def test_mapped_examples_iterable(n, func, batched, batch_size):
base_ex_iterable = ExamplesIterable(generate_examples_fn, {"n": n})
ex_iterable = MappedExamplesIterable(base_ex_iterable, func, batched=batched, batch_size=batch_size)
all_examples = [x for _, x in generate_examples_fn(n=n)]
if batched is False:
expected = [{**x, **func(x)} for x in all_examples]
else:
# For batched map we have to format the examples as a batch (i.e. in one single dictionary) to pass the batch to the function
all_transformed_examples = []
# If batch_size is None or <=0, we use the whole dataset as a single batch
if batch_size is None or batch_size <= 0:
batch_size = len(all_examples)
for batch_offset in range(0, len(all_examples), batch_size):
examples = all_examples[batch_offset : batch_offset + batch_size]
batch = _examples_to_batch(examples)
transformed_batch = func(batch)
all_transformed_examples.extend(_batch_to_examples(transformed_batch))
expected = _examples_to_batch(all_examples)
expected.update(_examples_to_batch(all_transformed_examples))
expected = list(_batch_to_examples(expected))
assert next(iter(ex_iterable))[1] == expected[0]
assert [x for _, x in ex_iterable] == expected
@pytest.mark.parametrize(
"n, func, batched, batch_size",
[
(3, lambda x: {"id+1": x["id"] + 1}, False, None), # just add 1 to the id
(3, lambda x: {"id+1": [x["id"][0] + 1]}, True, 1), # same with bs=1
(5, lambda x: {"id+1": [i + 1 for i in x["id"]]}, True, 10), # same with bs=10
(25, lambda x: {"id+1": [i + 1 for i in x["id"]]}, True, 10), # same with bs=10
(5, lambda x: {"id+1": [i + 1 for i in x["id"]]}, True, None), # same with bs=None
(5, lambda x: {"id+1": [i + 1 for i in x["id"]]}, True, -1), # same with bs<=0
(3, lambda x: {k: v * 2 for k, v in x.items()}, True, 1), # make a duplicate of each example
],
)
def test_mapped_examples_iterable_drop_last_batch(n, func, batched, batch_size):
base_ex_iterable = ExamplesIterable(generate_examples_fn, {"n": n})
ex_iterable = MappedExamplesIterable(
base_ex_iterable, func, batched=batched, batch_size=batch_size, drop_last_batch=True
)
all_examples = [x for _, x in generate_examples_fn(n=n)]
is_empty = False
if batched is False:
# `drop_last_batch` has no effect here
expected = [{**x, **func(x)} for x in all_examples]
else:
# For batched map we have to format the examples as a batch (i.e. in one single dictionary) to pass the batch to the function
all_transformed_examples = []
# If batch_size is None or <=0, we use the whole dataset as a single batch
if batch_size is None or batch_size <= 0:
batch_size = len(all_examples)
for batch_offset in range(0, len(all_examples), batch_size):
examples = all_examples[batch_offset : batch_offset + batch_size]
if len(examples) < batch_size: # ignore last batch
break
batch = _examples_to_batch(examples)
transformed_batch = func(batch)
all_transformed_examples.extend(_batch_to_examples(transformed_batch))
all_examples = all_examples if n % batch_size == 0 else all_examples[: n // batch_size * batch_size]
if all_examples:
expected = _examples_to_batch(all_examples)
expected.update(_examples_to_batch(all_transformed_examples))
expected = list(_batch_to_examples(expected))
else:
is_empty = True
if not is_empty:
assert next(iter(ex_iterable))[1] == expected[0]
assert [x for _, x in ex_iterable] == expected
else:
with pytest.raises(StopIteration):
next(iter(ex_iterable))
@pytest.mark.parametrize(
"n, func, batched, batch_size",
[
(3, lambda x, index: {"id+idx": x["id"] + index}, False, None), # add the index to the id
(
25,
lambda x, indices: {"id+idx": [i + j for i, j in zip(x["id"], indices)]},
True,
10,
), # add the index to the id
(5, lambda x, indices: {"id+idx": [i + j for i, j in zip(x["id"], indices)]}, True, None), # same with bs=None
(5, lambda x, indices: {"id+idx": [i + j for i, j in zip(x["id"], indices)]}, True, -1), # same with bs<=0
],
)
def test_mapped_examples_iterable_with_indices(n, func, batched, batch_size):
base_ex_iterable = ExamplesIterable(generate_examples_fn, {"n": n})
ex_iterable = MappedExamplesIterable(
base_ex_iterable, func, batched=batched, batch_size=batch_size, with_indices=True
)
all_examples = [x for _, x in generate_examples_fn(n=n)]
if batched is False:
expected = [{**x, **func(x, idx)} for idx, x in enumerate(all_examples)]
else:
# For batched map we have to format the examples as a batch (i.e. in one single dictionary) to pass the batch to the function
all_transformed_examples = []
# If batch_size is None or <=0, we use the whole dataset as a single batch
if batch_size is None or batch_size <= 0:
batch_size = len(all_examples)
for batch_offset in range(0, len(all_examples), batch_size):
examples = all_examples[batch_offset : batch_offset + batch_size]
batch = _examples_to_batch(examples)
indices = list(range(batch_offset, batch_offset + len(examples)))
transformed_batch = func(batch, indices)
all_transformed_examples.extend(_batch_to_examples(transformed_batch))
expected = _examples_to_batch(all_examples)
expected.update(_examples_to_batch(all_transformed_examples))
expected = list(_batch_to_examples(expected))
assert next(iter(ex_iterable))[1] == expected[0]
assert [x for _, x in ex_iterable] == expected
@pytest.mark.parametrize(
"n, func, batched, batch_size, remove_columns",
[
(3, lambda x: {"id+1": x["id"] + 1}, False, None, ["extra_column"]), # just add 1 to the id
(25, lambda x: {"id+1": [i + 1 for i in x["id"]]}, True, 10, ["extra_column"]), # same with bs=10
(
50,
lambda x: {"foo": ["bar"] * np.random.default_rng(x["id"][0]).integers(0, 10)},
True,
8,
["extra_column", "id"],
), # make a duplicate of each example
(5, lambda x: {"id+1": [i + 1 for i in x["id"]]}, True, None, ["extra_column"]), # same with bs=None
(5, lambda x: {"id+1": [i + 1 for i in x["id"]]}, True, -1, ["extra_column"]), # same with bs<=0
],
)
def test_mapped_examples_iterable_remove_columns(n, func, batched, batch_size, remove_columns):
base_ex_iterable = ExamplesIterable(generate_examples_fn, {"n": n, "extra_column": "foo"})
ex_iterable = MappedExamplesIterable(
base_ex_iterable, func, batched=batched, batch_size=batch_size, remove_columns=remove_columns
)
all_examples = [x for _, x in generate_examples_fn(n=n)]
columns_to_remove = remove_columns if isinstance(remove_columns, list) else [remove_columns]
if batched is False:
expected = [{**{k: v for k, v in x.items() if k not in columns_to_remove}, **func(x)} for x in all_examples]
else:
# For batched map we have to format the examples as a batch (i.e. in one single dictionary) to pass the batch to the function
all_transformed_examples = []
# If batch_size is None or <=0, we use the whole dataset as a single batch
if batch_size is None or batch_size <= 0:
batch_size = len(all_examples)
for batch_offset in range(0, len(all_examples), batch_size):
examples = all_examples[batch_offset : batch_offset + batch_size]
batch = _examples_to_batch(examples)
transformed_batch = func(batch)
all_transformed_examples.extend(_batch_to_examples(transformed_batch))
expected = {k: v for k, v in _examples_to_batch(all_examples).items() if k not in columns_to_remove}
expected.update(_examples_to_batch(all_transformed_examples))
expected = list(_batch_to_examples(expected))
assert next(iter(ex_iterable))[1] == expected[0]
assert [x for _, x in ex_iterable] == expected
@pytest.mark.parametrize(
"n, func, batched, batch_size, fn_kwargs",
[
(3, lambda x, y=0: {"id+y": x["id"] + y}, False, None, None),
(3, lambda x, y=0: {"id+y": x["id"] + y}, False, None, {"y": 3}),
(25, lambda x, y=0: {"id+y": [i + y for i in x["id"]]}, True, 10, {"y": 3}),
(5, lambda x, y=0: {"id+y": [i + y for i in x["id"]]}, True, None, {"y": 3}), # same with bs=None
(5, lambda x, y=0: {"id+y": [i + y for i in x["id"]]}, True, -1, {"y": 3}), # same with bs<=0
],
)
def test_mapped_examples_iterable_fn_kwargs(n, func, batched, batch_size, fn_kwargs):
base_ex_iterable = ExamplesIterable(generate_examples_fn, {"n": n})
ex_iterable = MappedExamplesIterable(
base_ex_iterable, func, batched=batched, batch_size=batch_size, fn_kwargs=fn_kwargs
)
all_examples = [x for _, x in generate_examples_fn(n=n)]
if fn_kwargs is None:
fn_kwargs = {}
if batched is False:
expected = [{**x, **func(x, **fn_kwargs)} for x in all_examples]
else:
# For batched map we have to format the examples as a batch (i.e. in one single dictionary) to pass the batch to the function
all_transformed_examples = []
# If batch_size is None or <=0, we use the whole dataset as a single batch
if batch_size is None or batch_size <= 0:
batch_size = len(all_examples)
for batch_offset in range(0, len(all_examples), batch_size):
examples = all_examples[batch_offset : batch_offset + batch_size]
batch = _examples_to_batch(examples)
transformed_batch = func(batch, **fn_kwargs)
all_transformed_examples.extend(_batch_to_examples(transformed_batch))
expected = _examples_to_batch(all_examples)
expected.update(_examples_to_batch(all_transformed_examples))
expected = list(_batch_to_examples(expected))
assert next(iter(ex_iterable))[1] == expected[0]
assert [x for _, x in ex_iterable] == expected
@pytest.mark.parametrize(
"n, func, batched, batch_size, input_columns",
[
(3, lambda id_: {"id+1": id_ + 1}, False, None, ["id"]), # just add 1 to the id
(25, lambda ids_: {"id+1": [i + 1 for i in ids_]}, True, 10, ["id"]), # same with bs=10
(5, lambda ids_: {"id+1": [i + 1 for i in ids_]}, True, None, ["id"]), # same with bs=None
(5, lambda ids_: {"id+1": [i + 1 for i in ids_]}, True, -1, ["id"]), # same with bs<=0
],
)
def test_mapped_examples_iterable_input_columns(n, func, batched, batch_size, input_columns):
base_ex_iterable = ExamplesIterable(generate_examples_fn, {"n": n})
ex_iterable = MappedExamplesIterable(
base_ex_iterable, func, batched=batched, batch_size=batch_size, input_columns=input_columns
)
all_examples = [x for _, x in generate_examples_fn(n=n)]
columns_to_input = input_columns if isinstance(input_columns, list) else [input_columns]
if batched is False:
expected = [{**x, **func(*[x[col] for col in columns_to_input])} for x in all_examples]
else:
# For batched map we have to format the examples as a batch (i.e. in one single dictionary) to pass the batch to the function
all_transformed_examples = []
# If batch_size is None or <=0, we use the whole dataset as a single batch
if batch_size is None or batch_size <= 0:
batch_size = len(all_examples)
for batch_offset in range(0, len(all_examples), batch_size):
examples = all_examples[batch_offset : batch_offset + batch_size]
batch = _examples_to_batch(examples)
transformed_batch = func(*[batch[col] for col in columns_to_input])
all_transformed_examples.extend(_batch_to_examples(transformed_batch))
expected = _examples_to_batch(all_examples)
expected.update(_examples_to_batch(all_transformed_examples))
expected = list(_batch_to_examples(expected))
assert next(iter(ex_iterable))[1] == expected[0]
assert [x for _, x in ex_iterable] == expected
@pytest.mark.parametrize(
"n, func, batched, batch_size",
[
(3, lambda t: t.append_column("id+1", pc.add(t["id"], 1)), False, None), # just add 1 to the id
(3, lambda t: t.append_column("id+1", pc.add(t["id"], 1)), True, 1), # same with bs=1
(5, lambda t: t.append_column("id+1", pc.add(t["id"], 1)), True, 10), # same with bs=10
(25, lambda t: t.append_column("id+1", pc.add(t["id"], 1)), True, 10), # same with bs=10
(5, lambda t: t.append_column("id+1", pc.add(t["id"], 1)), True, None), # same with bs=None
(5, lambda t: t.append_column("id+1", pc.add(t["id"], 1)), True, -1), # same with bs<=0
(3, lambda t: pa.concat_tables([t] * 2), True, 1), # make a duplicate of each example
],
)
def test_mapped_examples_iterable_arrow_format(n, func, batched, batch_size):
base_ex_iterable = ExamplesIterable(generate_examples_fn, {"n": n})
ex_iterable = MappedExamplesIterable(
base_ex_iterable,
func,
batched=batched,
batch_size=batch_size,
formatting=FormattingConfig(format_type="arrow"),
)
all_examples = [x for _, x in generate_examples_fn(n=n)]
if batched is False:
expected = [func(pa.Table.from_pylist([x])).to_pylist()[0] for x in all_examples]
else:
expected = []
# If batch_size is None or <=0, we use the whole dataset as a single batch
if batch_size is None or batch_size <= 0:
batch_size = len(all_examples)
for batch_offset in range(0, len(all_examples), batch_size):
examples = all_examples[batch_offset : batch_offset + batch_size]
batch = pa.Table.from_pylist(examples)
expected.extend(func(batch).to_pylist())
assert next(iter(ex_iterable))[1] == expected[0]
assert [x for _, x in ex_iterable] == expected
@pytest.mark.parametrize(
"n, func, batched, batch_size",
[
(3, lambda t: t.append_column("id+1", pc.add(t["id"], 1)), False, None), # just add 1 to the id
(3, lambda t: t.append_column("id+1", pc.add(t["id"], 1)), True, 1), # same with bs=1
(5, lambda t: t.append_column("id+1", pc.add(t["id"], 1)), True, 10), # same with bs=10
(25, lambda t: t.append_column("id+1", pc.add(t["id"], 1)), True, 10), # same with bs=10
(5, lambda t: t.append_column("id+1", pc.add(t["id"], 1)), True, None), # same with bs=None
(5, lambda t: t.append_column("id+1", pc.add(t["id"], 1)), True, -1), # same with bs<=0
(3, lambda t: pa.concat_tables([t] * 2), True, 1), # make a duplicate of each example
],
)
def test_mapped_examples_iterable_drop_last_batch_and_arrow_format(n, func, batched, batch_size):
base_ex_iterable = ExamplesIterable(generate_examples_fn, {"n": n})
ex_iterable = MappedExamplesIterable(
base_ex_iterable,
func,
batched=batched,
batch_size=batch_size,
drop_last_batch=True,
formatting=FormattingConfig(format_type="arrow"),
)
all_examples = [x for _, x in generate_examples_fn(n=n)]
is_empty = False
if batched is False:
# `drop_last_batch` has no effect here
expected = [func(pa.Table.from_pylist([x])).to_pylist()[0] for x in all_examples]
else:
all_transformed_examples = []
# If batch_size is None or <=0, we use the whole dataset as a single batch
if batch_size is None or batch_size <= 0:
batch_size = len(all_examples)
for batch_offset in range(0, len(all_examples), batch_size):
examples = all_examples[batch_offset : batch_offset + batch_size]
if len(examples) < batch_size: # ignore last batch
break
batch = pa.Table.from_pylist(examples)
out = func(batch)
all_transformed_examples.extend(
out.to_pylist()
) # we don't merge with input since they're arrow tables and not dictionaries
all_examples = all_examples if n % batch_size == 0 else all_examples[: n // batch_size * batch_size]
if all_examples:
expected = all_transformed_examples
else:
is_empty = True
if not is_empty:
assert next(iter(ex_iterable))[1] == expected[0]
assert [x for _, x in ex_iterable] == expected
else:
with pytest.raises(StopIteration):
next(iter(ex_iterable))
@pytest.mark.parametrize(
"n, func, batched, batch_size",
[
(
3,
lambda t, index: t.append_column("id+idx", pc.add(t["id"], index)),
False,
None,
), # add the index to the id
(
25,
lambda t, indices: t.append_column("id+idx", pc.add(t["id"], indices)),
True,
10,
), # add the index to the id
(5, lambda t, indices: t.append_column("id+idx", pc.add(t["id"], indices)), True, None), # same with bs=None
(5, lambda t, indices: t.append_column("id+idx", pc.add(t["id"], indices)), True, -1), # same with bs<=0
],
)
def test_mapped_examples_iterable_with_indices_and_arrow_format(n, func, batched, batch_size):
base_ex_iterable = ExamplesIterable(generate_examples_fn, {"n": n})
ex_iterable = MappedExamplesIterable(
base_ex_iterable,
func,
batched=batched,
batch_size=batch_size,
with_indices=True,
formatting=FormattingConfig(format_type="arrow"),
)
all_examples = [x for _, x in generate_examples_fn(n=n)]
if batched is False:
expected = [func(pa.Table.from_pylist([x]), i).to_pylist()[0] for i, x in enumerate(all_examples)]
else:
expected = []
# If batch_size is None or <=0, we use the whole dataset as a single batch
if batch_size is None or batch_size <= 0:
batch_size = len(all_examples)
for batch_offset in range(0, len(all_examples), batch_size):
examples = all_examples[batch_offset : batch_offset + batch_size]
batch = pa.Table.from_pylist(examples)
expected.extend(func(batch, list(range(batch_offset, batch_offset + len(batch)))).to_pylist())
assert next(iter(ex_iterable))[1] == expected[0]
assert [x for _, x in ex_iterable] == expected
@pytest.mark.parametrize(
"n, func, batched, batch_size, remove_columns",
[
(
3,
lambda t: t.append_column("id+1", pc.add(t["id"], 1)),
False,
None,
["extra_column"],
), # just add 1 to the id
(25, lambda t: t.append_column("id+1", pc.add(t["id"], 1)), True, 10, ["extra_column"]), # same with bs=10
(
50,
lambda t: pa.table({"foo": ["bar"] * np.random.default_rng(t["id"][0].as_py()).integers(0, 10)}),
True,
8,
["extra_column", "id"],
), # make a duplicate of each example
(5, lambda t: t.append_column("id+1", pc.add(t["id"], 1)), True, None, ["extra_column"]), # same with bs=None
(5, lambda t: t.append_column("id+1", pc.add(t["id"], 1)), True, -1, ["extra_column"]), # same with bs<=0
],
)
def test_mapped_examples_iterable_remove_columns_arrow_format(n, func, batched, batch_size, remove_columns):
base_ex_iterable = ExamplesIterable(generate_examples_fn, {"n": n, "extra_column": "foo"})
ex_iterable = MappedExamplesIterable(
base_ex_iterable,
func,
batched=batched,
batch_size=batch_size,
remove_columns=remove_columns,
formatting=FormattingConfig(format_type="arrow"),
)
all_examples = [x for _, x in generate_examples_fn(n=n)]
columns_to_remove = remove_columns if isinstance(remove_columns, list) else [remove_columns]
if batched is False:
expected = [
{**{k: v for k, v in func(pa.Table.from_pylist([x])).to_pylist()[0].items() if k not in columns_to_remove}}
for x in all_examples
]
else:
expected = []
# If batch_size is None or <=0, we use the whole dataset as a single batch
if batch_size is None or batch_size <= 0:
batch_size = len(all_examples)
for batch_offset in range(0, len(all_examples), batch_size):
examples = all_examples[batch_offset : batch_offset + batch_size]
batch = pa.Table.from_pylist(examples)
expected.extend(
[{k: v for k, v in x.items() if k not in columns_to_remove} for x in func(batch).to_pylist()]
)
assert next(iter(ex_iterable))[1] == expected[0]
assert [x for _, x in ex_iterable] == expected
@pytest.mark.parametrize(
"n, func, batched, batch_size, fn_kwargs",
[
(3, lambda t, y=0: t.append_column("id+idx", pc.add(t["id"], y)), False, None, None),
(3, lambda t, y=0: t.append_column("id+idx", pc.add(t["id"], y)), False, None, {"y": 3}),
(25, lambda t, y=0: t.append_column("id+idx", pc.add(t["id"], y)), True, 10, {"y": 3}),
(5, lambda t, y=0: t.append_column("id+idx", pc.add(t["id"], y)), True, None, {"y": 3}), # same with bs=None
(5, lambda t, y=0: t.append_column("id+idx", pc.add(t["id"], y)), True, -1, {"y": 3}), # same with bs<=0
],
)
def test_mapped_examples_iterable_fn_kwargs_and_arrow_format(n, func, batched, batch_size, fn_kwargs):
base_ex_iterable = ExamplesIterable(generate_examples_fn, {"n": n})
ex_iterable = MappedExamplesIterable(
base_ex_iterable,
func,
batched=batched,
batch_size=batch_size,
fn_kwargs=fn_kwargs,
formatting=FormattingConfig(format_type="arrow"),
)
all_examples = [x for _, x in generate_examples_fn(n=n)]
if fn_kwargs is None:
fn_kwargs = {}
if batched is False:
expected = [func(pa.Table.from_pylist([x]), **fn_kwargs).to_pylist()[0] for x in all_examples]
else:
expected = []
# If batch_size is None or <=0, we use the whole dataset as a single batch
if batch_size is None or batch_size <= 0:
batch_size = len(all_examples)
for batch_offset in range(0, len(all_examples), batch_size):
examples = all_examples[batch_offset : batch_offset + batch_size]
batch = pa.Table.from_pylist(examples)
expected.extend(func(batch, **fn_kwargs).to_pylist())
assert next(iter(ex_iterable))[1] == expected[0]
assert [x for _, x in ex_iterable] == expected
@pytest.mark.parametrize(
"n, func, batched, batch_size, input_columns",
[
(3, lambda id_: pa.table({"id+1": pc.add(id_, 1)}), False, None, ["id"]), # just add 1 to the id
(25, lambda ids_: pa.table({"id+1": pc.add(ids_, 1)}), True, 10, ["id"]), # same with bs=10
(5, lambda ids_: pa.table({"id+1": pc.add(ids_, 1)}), True, None, ["id"]), # same with bs=None
(5, lambda ids_: pa.table({"id+1": pc.add(ids_, 1)}), True, -1, ["id"]), # same with bs<=0
],
)
def test_mapped_examples_iterable_input_columns_and_arrow_format(n, func, batched, batch_size, input_columns):
base_ex_iterable = ExamplesIterable(generate_examples_fn, {"n": n})
ex_iterable = MappedExamplesIterable(
base_ex_iterable,
func,
batched=batched,
batch_size=batch_size,
input_columns=input_columns,
formatting=FormattingConfig(format_type="arrow"),
)
all_examples = [x for _, x in generate_examples_fn(n=n)]
columns_to_input = input_columns if isinstance(input_columns, list) else [input_columns]
if batched is False:
expected = [
func(*[pa.Table.from_pylist([x])[col] for col in columns_to_input]).to_pylist()[0] for x in all_examples
]
else:
expected = []
# If batch_size is None or <=0, we use the whole dataset as a single batch
if batch_size is None or batch_size <= 0:
batch_size = len(all_examples)
for batch_offset in range(0, len(all_examples), batch_size):
examples = all_examples[batch_offset : batch_offset + batch_size]
batch = pa.Table.from_pylist(examples)
expected.extend(func(*[batch[col] for col in columns_to_input]).to_pylist())
assert next(iter(ex_iterable))[1] == expected[0]
assert [x for _, x in ex_iterable] == expected
@pytest.mark.parametrize(
"n, func, batched, batch_size",
[
(3, lambda x: x["id"] % 2 == 0, False, None), # keep even number
(3, lambda x: [x["id"][0] % 2 == 0], True, 1), # same with bs=1
(25, lambda x: [i % 2 == 0 for i in x["id"]], True, 10), # same with bs=10
(5, lambda x: [i % 2 == 0 for i in x["id"]], True, None), # same with bs=None
(5, lambda x: [i % 2 == 0 for i in x["id"]], True, -1), # same with bs<=0
(3, lambda x: False, False, None), # return 0 examples
(3, lambda x: [False] * len(x["id"]), True, 10), # same with bs=10
],
)
def test_filtered_examples_iterable(n, func, batched, batch_size):
base_ex_iterable = ExamplesIterable(generate_examples_fn, {"n": n})
ex_iterable = FilteredExamplesIterable(base_ex_iterable, func, batched=batched, batch_size=batch_size)
all_examples = [x for _, x in generate_examples_fn(n=n)]
if batched is False:
expected = [x for x in all_examples if func(x)]
else:
# For batched filter we have to format the examples as a batch (i.e. in one single dictionary) to pass the batch to the function
expected = []
# If batch_size is None or <=0, we use the whole dataset as a single batch
if batch_size is None or batch_size <= 0:
batch_size = len(all_examples)
for batch_offset in range(0, len(all_examples), batch_size):
examples = all_examples[batch_offset : batch_offset + batch_size]
batch = _examples_to_batch(examples)
mask = func(batch)
expected.extend([x for x, to_keep in zip(examples, mask) if to_keep])
if expected:
assert next(iter(ex_iterable))[1] == expected[0]
assert [x for _, x in ex_iterable] == expected
@pytest.mark.parametrize(
"n, func, batched, batch_size",
[
(3, lambda x, index: index % 2 == 0, False, None), # keep even number
(25, lambda x, indices: [idx % 2 == 0 for idx in indices], True, 10), # same with bs=10
(5, lambda x, indices: [idx % 2 == 0 for idx in indices], True, None), # same with bs=None
(5, lambda x, indices: [idx % 2 == 0 for idx in indices], True, -1), # same with bs<=0
],
)
def test_filtered_examples_iterable_with_indices(n, func, batched, batch_size):
base_ex_iterable = ExamplesIterable(generate_examples_fn, {"n": n})
ex_iterable = FilteredExamplesIterable(
base_ex_iterable, func, batched=batched, batch_size=batch_size, with_indices=True
)
all_examples = [x for _, x in generate_examples_fn(n=n)]
if batched is False:
expected = [x for idx, x in enumerate(all_examples) if func(x, idx)]
else:
# For batched filter we have to format the examples as a batch (i.e. in one single dictionary) to pass the batch to the function
expected = []
# If batch_size is None or <=0, we use the whole dataset as a single batch
if batch_size is None or batch_size <= 0:
batch_size = len(all_examples)
for batch_offset in range(0, len(all_examples), batch_size):
examples = all_examples[batch_offset : batch_offset + batch_size]
batch = _examples_to_batch(examples)
indices = list(range(batch_offset, batch_offset + len(examples)))
mask = func(batch, indices)
expected.extend([x for x, to_keep in zip(examples, mask) if to_keep])
assert next(iter(ex_iterable))[1] == expected[0]
assert [x for _, x in ex_iterable] == expected
@pytest.mark.parametrize(
"n, func, batched, batch_size, input_columns",
[
(3, lambda id_: id_ % 2 == 0, False, None, ["id"]), # keep even number
(25, lambda ids_: [i % 2 == 0 for i in ids_], True, 10, ["id"]), # same with bs=10
(3, lambda ids_: [i % 2 == 0 for i in ids_], True, None, ["id"]), # same with bs=None
(3, lambda ids_: [i % 2 == 0 for i in ids_], True, None, ["id"]), # same with bs=None
],
)
def test_filtered_examples_iterable_input_columns(n, func, batched, batch_size, input_columns):
base_ex_iterable = ExamplesIterable(generate_examples_fn, {"n": n})
ex_iterable = FilteredExamplesIterable(
base_ex_iterable, func, batched=batched, batch_size=batch_size, input_columns=input_columns
)
all_examples = [x for _, x in generate_examples_fn(n=n)]
columns_to_input = input_columns if isinstance(input_columns, list) else [input_columns]
if batched is False:
expected = [x for x in all_examples if func(*[x[col] for col in columns_to_input])]
else:
# For batched filter we have to format the examples as a batch (i.e. in one single dictionary) to pass the batch to the function
expected = []
# If batch_size is None or <=0, we use the whole dataset as a single batch
if batch_size is None or batch_size <= 0:
batch_size = len(all_examples)
for batch_offset in range(0, len(all_examples), batch_size):
examples = all_examples[batch_offset : batch_offset + batch_size]
batch = _examples_to_batch(examples)
mask = func(*[batch[col] for col in columns_to_input])
expected.extend([x for x, to_keep in zip(examples, mask) if to_keep])
assert next(iter(ex_iterable))[1] == expected[0]
assert [x for _, x in ex_iterable] == expected
def test_skip_examples_iterable():
total, count = 10, 2
base_ex_iterable = ExamplesIterable(generate_examples_fn, {"n": total})
skip_ex_iterable = SkipExamplesIterable(base_ex_iterable, n=count)
expected = list(generate_examples_fn(n=total))[count:]
assert list(skip_ex_iterable) == expected
assert (
skip_ex_iterable.shuffle_data_sources(np.random.default_rng(42)) is skip_ex_iterable
), "skip examples makes the shards order fixed"
def test_take_examples_iterable():
total, count = 10, 2
base_ex_iterable = ExamplesIterable(generate_examples_fn, {"n": total})
take_ex_iterable = TakeExamplesIterable(base_ex_iterable, n=count)
expected = list(generate_examples_fn(n=total))[:count]
assert list(take_ex_iterable) == expected
assert (
take_ex_iterable.shuffle_data_sources(np.random.default_rng(42)) is take_ex_iterable
), "skip examples makes the shards order fixed"
def test_vertically_concatenated_examples_iterable():
ex_iterable1 = ExamplesIterable(generate_examples_fn, {"label": 10})
ex_iterable2 = ExamplesIterable(generate_examples_fn, {"label": 5})
concatenated_ex_iterable = VerticallyConcatenatedMultiSourcesExamplesIterable([ex_iterable1, ex_iterable2])
expected = [x for _, x in ex_iterable1] + [x for _, x in ex_iterable2]
assert [x for _, x in concatenated_ex_iterable] == expected
def test_vertically_concatenated_examples_iterable_with_different_columns():
# having different columns is supported
# Though iterable datasets fill the missing data with nulls
ex_iterable1 = ExamplesIterable(generate_examples_fn, {"label": 10})
ex_iterable2 = ExamplesIterable(generate_examples_fn, {})
concatenated_ex_iterable = VerticallyConcatenatedMultiSourcesExamplesIterable([ex_iterable1, ex_iterable2])
expected = [x for _, x in ex_iterable1] + [x for _, x in ex_iterable2]
assert [x for _, x in concatenated_ex_iterable] == expected
def test_vertically_concatenated_examples_iterable_shuffle_data_sources():
ex_iterable1 = ExamplesIterable(generate_examples_fn, {"label": 10})
ex_iterable2 = ExamplesIterable(generate_examples_fn, {"label": 5})
concatenated_ex_iterable = VerticallyConcatenatedMultiSourcesExamplesIterable([ex_iterable1, ex_iterable2])
rng = np.random.default_rng(42)
shuffled_ex_iterable = concatenated_ex_iterable.shuffle_data_sources(rng)
# make sure the list of examples iterables is shuffled, and each examples iterable is shuffled
expected = [x for _, x in ex_iterable2.shuffle_data_sources(rng)] + [
x for _, x in ex_iterable1.shuffle_data_sources(rng)
]
assert [x for _, x in shuffled_ex_iterable] == expected
def test_horizontally_concatenated_examples_iterable():
ex_iterable1 = ExamplesIterable(generate_examples_fn, {"label1": 10})
ex_iterable2 = ExamplesIterable(generate_examples_fn, {"label2": 5})
concatenated_ex_iterable = HorizontallyConcatenatedMultiSourcesExamplesIterable([ex_iterable1, ex_iterable2])
with pytest.raises(ValueError): # column "id" is duplicated -> raise an error
list(concatenated_ex_iterable)
ex_iterable2 = MappedExamplesIterable(ex_iterable2, lambda x: x, remove_columns=["id"])
concatenated_ex_iterable = HorizontallyConcatenatedMultiSourcesExamplesIterable([ex_iterable1, ex_iterable2])
expected = [{**x, **y} for (_, x), (_, y) in zip(ex_iterable1, ex_iterable2)]
assert [x for _, x in concatenated_ex_iterable] == expected
assert (
concatenated_ex_iterable.shuffle_data_sources(np.random.default_rng(42)) is concatenated_ex_iterable
), "horizontally concatenated examples makes the shards order fixed"
@pytest.mark.parametrize(
"ex_iterable",
[
ExamplesIterable(generate_examples_fn, {}),
ShuffledDataSourcesExamplesIterable(generate_examples_fn, {}, np.random.default_rng(42)),
SelectColumnsIterable(ExamplesIterable(generate_examples_fn, {}), ["id"]),
StepExamplesIterable(ExamplesIterable(generate_examples_fn, {}), 2, 0),
CyclingMultiSourcesExamplesIterable([ExamplesIterable(generate_examples_fn, {})]),
VerticallyConcatenatedMultiSourcesExamplesIterable([ExamplesIterable(generate_examples_fn, {})]),
HorizontallyConcatenatedMultiSourcesExamplesIterable([ExamplesIterable(generate_examples_fn, {})]),
RandomlyCyclingMultiSourcesExamplesIterable(
[ExamplesIterable(generate_examples_fn, {})], np.random.default_rng(42)
),
MappedExamplesIterable(ExamplesIterable(generate_examples_fn, {}), lambda x: x),
MappedExamplesIterable(ArrowExamplesIterable(generate_tables_fn, {}), lambda x: x),
FilteredExamplesIterable(ExamplesIterable(generate_examples_fn, {}), lambda x: True),
FilteredExamplesIterable(ArrowExamplesIterable(generate_tables_fn, {}), lambda x: True),
BufferShuffledExamplesIterable(ExamplesIterable(generate_examples_fn, {}), 10, np.random.default_rng(42)),
SkipExamplesIterable(ExamplesIterable(generate_examples_fn, {}), 10),
TakeExamplesIterable(ExamplesIterable(generate_examples_fn, {}), 10),
TypedExamplesIterable(
ExamplesIterable(generate_examples_fn, {}), Features({"id": Value("int32")}), token_per_repo_id={}
),
],
)
def test_no_iter_arrow(ex_iterable: _BaseExamplesIterable):
assert ex_iterable.iter_arrow is None
@pytest.mark.parametrize(
"ex_iterable",
[
ArrowExamplesIterable(generate_tables_fn, {}),
ShuffledDataSourcesArrowExamplesIterable(generate_tables_fn, {}, np.random.default_rng(42)),
SelectColumnsIterable(ArrowExamplesIterable(generate_tables_fn, {}), ["id"]),
# StepExamplesIterable(ArrowExamplesIterable(generate_tables_fn, {}), 2, 0), # not implemented
# CyclingMultiSourcesExamplesIterable([ArrowExamplesIterable(generate_tables_fn, {})]), # not implemented
VerticallyConcatenatedMultiSourcesExamplesIterable([ArrowExamplesIterable(generate_tables_fn, {})]),
# HorizontallyConcatenatedMultiSourcesExamplesIterable([ArrowExamplesIterable(generate_tables_fn, {})]), # not implemented
# RandomlyCyclingMultiSourcesExamplesIterable([ArrowExamplesIterable(generate_tables_fn, {})], np.random.default_rng(42)), # not implemented
MappedExamplesIterable(
ExamplesIterable(generate_examples_fn, {}), lambda t: t, formatting=FormattingConfig(format_type="arrow")
),
MappedExamplesIterable(
ArrowExamplesIterable(generate_tables_fn, {}),
lambda t: t,
formatting=FormattingConfig(format_type="arrow"),
),
FilteredExamplesIterable(
ExamplesIterable(generate_examples_fn, {}),
lambda t: True,
formatting=FormattingConfig(format_type="arrow"),
),
FilteredExamplesIterable(
ArrowExamplesIterable(generate_tables_fn, {}),
lambda t: True,
formatting=FormattingConfig(format_type="arrow"),
),
# BufferShuffledExamplesIterable(ArrowExamplesIterable(generate_tables_fn, {}), 10, np.random.default_rng(42)), # not implemented
# SkipExamplesIterable(ArrowExamplesIterable(generate_tables_fn, {}), 10), # not implemented
# TakeExamplesIterable(ArrowExamplesIterable(generate_tables_fn, {}), 10), # not implemented
TypedExamplesIterable(
ArrowExamplesIterable(generate_tables_fn, {}), Features({"id": Value("int32")}), token_per_repo_id={}
),
],
)
def test_iter_arrow(ex_iterable: _BaseExamplesIterable):
assert ex_iterable.iter_arrow is not None
key, pa_table = next(ex_iterable.iter_arrow())
assert isinstance(pa_table, pa.Table)
############################
#
# IterableDataset tests
#
############################
def test_iterable_dataset():
dataset = IterableDataset(ExamplesIterable(generate_examples_fn, {}))
expected = [x for _, x in generate_examples_fn()]
assert next(iter(dataset)) == expected[0]
assert list(dataset) == expected
def test_iterable_dataset_from_generator():
data = [
{"col_1": "0", "col_2": 0, "col_3": 0.0},
{"col_1": "1", "col_2": 1, "col_3": 1.0},
{"col_1": "2", "col_2": 2, "col_3": 2.0},
{"col_1": "3", "col_2": 3, "col_3": 3.0},
]
def gen():
yield from data
dataset = IterableDataset.from_generator(gen)
assert isinstance(dataset, IterableDataset)
assert list(dataset) == data
def test_iterable_dataset_from_generator_with_shards():
def gen(shard_names):
for shard_name in shard_names:
for i in range(10):
yield {"shard_name": shard_name, "i": i}
shard_names = [f"data{shard_idx}.txt" for shard_idx in range(4)]
dataset = IterableDataset.from_generator(gen, gen_kwargs={"shard_names": shard_names})
assert isinstance(dataset, IterableDataset)
assert dataset.n_shards == len(shard_names)
def test_iterable_dataset_from_file(dataset: IterableDataset, arrow_file: str):
with assert_arrow_memory_doesnt_increase():
dataset_from_file = IterableDataset.from_file(arrow_file)
expected_features = dataset._resolve_features().features
assert dataset_from_file.features.type == expected_features.type
assert dataset_from_file.features == expected_features
assert isinstance(dataset_from_file, IterableDataset)
assert list(dataset_from_file) == list(dataset)
@require_not_windows
@require_dill_gt_0_3_2
@require_pyspark
def test_from_spark_streaming():
import pyspark
spark = pyspark.sql.SparkSession.builder.master("local[*]").appName("pyspark").getOrCreate()
data = [
("0", 0, 0.0),
("1", 1, 1.0),
("2", 2, 2.0),
("3", 3, 3.0),
]
df = spark.createDataFrame(data, "col_1: string, col_2: int, col_3: float")
dataset = IterableDataset.from_spark(df)
assert isinstance(dataset, IterableDataset)
results = []
for ex in dataset:
results.append(ex)
assert results == [
{"col_1": "0", "col_2": 0, "col_3": 0.0},
{"col_1": "1", "col_2": 1, "col_3": 1.0},
{"col_1": "2", "col_2": 2, "col_3": 2.0},
{"col_1": "3", "col_2": 3, "col_3": 3.0},
]
@require_not_windows
@require_dill_gt_0_3_2
@require_pyspark
def test_from_spark_streaming_features():
import PIL.Image
import pyspark
spark = pyspark.sql.SparkSession.builder.master("local[*]").appName("pyspark").getOrCreate()
data = [(0, np.arange(4 * 4 * 3).reshape(4, 4, 3).tolist())]
df = spark.createDataFrame(data, "idx: int, image: array<array<array<int>>>")
features = Features({"idx": Value("int64"), "image": Image()})
dataset = IterableDataset.from_spark(
df,
features=features,
)
assert isinstance(dataset, IterableDataset)
results = []
for ex in dataset:
results.append(ex)
assert len(results) == 1
isinstance(results[0]["image"], PIL.Image.Image)
@require_torch
def test_iterable_dataset_torch_integration():
ex_iterable = ExamplesIterable(generate_examples_fn, {})
dataset = IterableDataset(ex_iterable)
import torch.utils.data
assert isinstance(dataset, torch.utils.data.IterableDataset)
assert isinstance(dataset, IterableDataset)
assert dataset._ex_iterable is ex_iterable
@require_torch
def test_iterable_dataset_torch_picklable():
import pickle
ex_iterable = ExamplesIterable(generate_examples_fn, {})
dataset = IterableDataset(ex_iterable, formatting=FormattingConfig(format_type="torch"))
reloaded_dataset = pickle.loads(pickle.dumps(dataset))
import torch.utils.data
assert isinstance(reloaded_dataset, IterableDataset)
assert isinstance(reloaded_dataset, torch.utils.data.IterableDataset)
assert reloaded_dataset._formatting.format_type == "torch"
assert len(list(dataset)) == len(list(reloaded_dataset))
@require_torch
def test_iterable_dataset_with_format_torch():
ex_iterable = ExamplesIterable(generate_examples_fn, {})
dataset = IterableDataset(ex_iterable)
from torch.utils.data import DataLoader
dataloader = DataLoader(dataset)
assert len(list(dataloader)) == len(list(ex_iterable))
@require_torch
def test_iterable_dataset_torch_dataloader_parallel():
from torch.utils.data import DataLoader
ex_iterable = ExamplesIterable(generate_examples_fn, {})
dataset = IterableDataset(ex_iterable)
dataloader = DataLoader(dataset, num_workers=2, batch_size=None)
result = list(dataloader)
expected = [example for _, example in ex_iterable]
assert len(result) == len(expected)
assert {str(x) for x in result} == {str(x) for x in expected}
@require_torch
@pytest.mark.filterwarnings("ignore:This DataLoader will create:UserWarning")
@pytest.mark.parametrize("n_shards, num_workers", [(2, 1), (2, 2), (3, 2), (2, 3)])
def test_sharded_iterable_dataset_torch_dataloader_parallel(n_shards, num_workers):
from torch.utils.data import DataLoader
ex_iterable = ExamplesIterable(generate_examples_fn, {"filepaths": [f"{i}.txt" for i in range(n_shards)]})
dataset = IterableDataset(ex_iterable)
dataloader = DataLoader(dataset, batch_size=None, num_workers=num_workers)
result = list(dataloader)
expected = [example for _, example in ex_iterable]
assert len(result) == len(expected)
assert {str(x) for x in result} == {str(x) for x in expected}
@require_torch
@pytest.mark.integration
@pytest.mark.parametrize("num_workers", [1, 2])
def test_iterable_dataset_from_hub_torch_dataloader_parallel(num_workers, tmp_path):
from torch.utils.data import DataLoader
dataset = load_dataset(SAMPLE_DATASET_IDENTIFIER, cache_dir=str(tmp_path), streaming=True, split="train")
dataloader = DataLoader(dataset, batch_size=None, num_workers=num_workers)
result = list(dataloader)
assert len(result) == 2
@pytest.mark.parametrize("batch_size", [4, 5])
@pytest.mark.parametrize("drop_last_batch", [False, True])
def test_iterable_dataset_iter_batch(batch_size, drop_last_batch):
n = 25
dataset = IterableDataset(ExamplesIterable(generate_examples_fn, {"n": n}))
all_examples = [ex for _, ex in generate_examples_fn(n=n)]
expected = []
for i in range(0, len(all_examples), batch_size):
if len(all_examples[i : i + batch_size]) < batch_size and drop_last_batch:
continue
expected.append(_examples_to_batch(all_examples[i : i + batch_size]))
assert next(iter(dataset.iter(batch_size, drop_last_batch=drop_last_batch))) == expected[0]
assert list(dataset.iter(batch_size, drop_last_batch=drop_last_batch)) == expected
def test_iterable_dataset_info():
info = DatasetInfo(description="desc", citation="@article{}", size_in_bytes=42)
ex_iterable = ExamplesIterable(generate_examples_fn, {})
dataset = IterableDataset(ex_iterable, info=info)
assert dataset.info == info
assert dataset.description == info.description
assert dataset.citation == info.citation
assert dataset.size_in_bytes == info.size_in_bytes
def test_iterable_dataset_set_epoch(dataset: IterableDataset):
assert dataset._epoch == 0
dataset.set_epoch(42)
assert dataset._epoch == 42
@pytest.mark.parametrize("seed", [None, 42, 1337])
@pytest.mark.parametrize("epoch", [None, 0, 1, 10])
def test_iterable_dataset_set_epoch_of_shuffled_dataset(dataset: IterableDataset, seed, epoch):
buffer_size = 10
shuffled_dataset = dataset.shuffle(seed, buffer_size=buffer_size)
base_generator = shuffled_dataset._shuffling.generator
if epoch is not None:
shuffled_dataset.set_epoch(epoch)
effective_generator = shuffled_dataset._effective_generator()
assert effective_generator is not None
if epoch is None or epoch == 0:
assert is_rng_equal(base_generator, shuffled_dataset._effective_generator())
else:
assert not is_rng_equal(base_generator, shuffled_dataset._effective_generator())
effective_seed = deepcopy(base_generator).integers(0, 1 << 63) - epoch
assert is_rng_equal(np.random.default_rng(effective_seed), shuffled_dataset._effective_generator())
def test_iterable_dataset_map(
dataset: IterableDataset,
):
func = lambda x: {"id+1": x["id"] + 1} # noqa: E731
mapped_dataset = dataset.map(func)
assert isinstance(mapped_dataset._ex_iterable, MappedExamplesIterable)
assert mapped_dataset._ex_iterable.function is func
assert mapped_dataset._ex_iterable.batched is False
assert next(iter(mapped_dataset)) == {**next(iter(dataset)), **func(next(iter(generate_examples_fn()))[1])}
def test_iterable_dataset_map_batched(
dataset: IterableDataset,
):
func = lambda x: {"id+1": [i + 1 for i in x["id"]]} # noqa: E731
batch_size = 3
dataset = dataset.map(func, batched=True, batch_size=batch_size)
assert isinstance(dataset._ex_iterable, MappedExamplesIterable)
assert dataset._ex_iterable.function is func
assert dataset._ex_iterable.batch_size == batch_size
assert next(iter(dataset)) == {"id": 0, "id+1": 1}
def test_iterable_dataset_map_complex_features(
dataset: IterableDataset,
):
# https://github.com/huggingface/datasets/issues/3505
ex_iterable = ExamplesIterable(generate_examples_fn, {"label": "positive"})
features = Features(
{
"id": Value("int64"),
"label": Value("string"),
}
)
dataset = IterableDataset(ex_iterable, info=DatasetInfo(features=features))
dataset = dataset.cast_column("label", ClassLabel(names=["negative", "positive"]))
dataset = dataset.map(lambda x: {"id+1": x["id"] + 1, **x})
assert isinstance(dataset._ex_iterable, MappedExamplesIterable)
features["label"] = ClassLabel(names=["negative", "positive"])
assert [{k: v for k, v in ex.items() if k != "id+1"} for ex in dataset] == [
features.encode_example(ex) for _, ex in ex_iterable
]
def test_iterable_dataset_map_with_features(dataset: IterableDataset) -> None:
# https://github.com/huggingface/datasets/issues/3888
ex_iterable = ExamplesIterable(generate_examples_fn, {"label": "positive"})
features_before_map = Features(
{
"id": Value("int64"),
"label": Value("string"),
}
)
dataset = IterableDataset(ex_iterable, info=DatasetInfo(features=features_before_map))
assert dataset.info.features is not None
assert dataset.info.features == features_before_map
features_after_map = Features(
{
"id": Value("int64"),
"label": Value("string"),
"target": Value("string"),
}
)
dataset = dataset.map(lambda x: {"target": x["label"]}, features=features_after_map)
assert dataset.info.features is not None
assert dataset.info.features == features_after_map
def test_iterable_dataset_map_with_fn_kwargs(dataset: IterableDataset) -> None:
fn_kwargs = {"y": 1}
mapped_dataset = dataset.map(lambda x, y: {"id+y": x["id"] + y}, fn_kwargs=fn_kwargs)
assert mapped_dataset._ex_iterable.batched is False
assert next(iter(mapped_dataset)) == {"id": 0, "id+y": 1}
batch_size = 3
mapped_dataset = dataset.map(
lambda x, y: {"id+y": [i + y for i in x["id"]]}, batched=True, batch_size=batch_size, fn_kwargs=fn_kwargs
)
assert isinstance(mapped_dataset._ex_iterable, MappedExamplesIterable)
assert mapped_dataset._ex_iterable.batch_size == batch_size
assert next(iter(mapped_dataset)) == {"id": 0, "id+y": 1}
def test_iterable_dataset_filter(dataset: IterableDataset) -> None:
fn_kwargs = {"y": 1}
filtered_dataset = dataset.filter(lambda x, y: x["id"] == y, fn_kwargs=fn_kwargs)
assert filtered_dataset._ex_iterable.batched is False
assert next(iter(filtered_dataset)) == {"id": 1}
@pytest.mark.parametrize("seed", [42, 1337, 101010, 123456])
@pytest.mark.parametrize("epoch", [None, 0, 1])
def test_iterable_dataset_shuffle(dataset: IterableDataset, seed, epoch):
buffer_size = 3
dataset = deepcopy(dataset)
dataset._ex_iterable.kwargs["filepaths"] = ["0.txt", "1.txt"]
dataset = dataset.shuffle(seed, buffer_size=buffer_size)
assert isinstance(dataset._shuffling, ShufflingConfig)
assert isinstance(dataset._shuffling.generator, np.random.Generator)
assert is_rng_equal(dataset._shuffling.generator, np.random.default_rng(seed))
# Effective seed is sum of seed and epoch
if epoch is None or epoch == 0:
effective_seed = seed
else:
dataset.set_epoch(epoch)
effective_seed = np.random.default_rng(seed).integers(0, 1 << 63) - epoch
# Shuffling adds a shuffle buffer
expected_first_example_index = next(
iter(BufferShuffledExamplesIterable._iter_random_indices(np.random.default_rng(effective_seed), buffer_size))
)
assert isinstance(dataset._ex_iterable, BufferShuffledExamplesIterable)
# It also shuffles the underlying examples iterable
expected_ex_iterable = ExamplesIterable(
generate_examples_fn, {"filepaths": ["0.txt", "1.txt"]}
).shuffle_data_sources(np.random.default_rng(effective_seed))
assert isinstance(dataset._ex_iterable.ex_iterable, ExamplesIterable)
assert next(iter(dataset)) == list(islice(expected_ex_iterable, expected_first_example_index + 1))[-1][1]
@pytest.mark.parametrize(
"features",
[
None,
Features(
{
"id": Value("int64"),
"label": Value("int64"),
}
),
Features(
{
"id": Value("int64"),
"label": ClassLabel(names=["negative", "positive"]),
}
),
],
)
def test_iterable_dataset_features(features):
ex_iterable = ExamplesIterable(generate_examples_fn, {"label": 0})
dataset = IterableDataset(ex_iterable, info=DatasetInfo(features=features))
if features:
expected = [features.encode_example(x) for _, x in ex_iterable]
else:
expected = [x for _, x in ex_iterable]
assert list(dataset) == expected
def test_iterable_dataset_features_cast_to_python():
ex_iterable = ExamplesIterable(
generate_examples_fn, {"timestamp": pd.Timestamp(2020, 1, 1), "array": np.ones(5), "n": 1}
)
features = Features(
{
"id": Value("int64"),
"timestamp": Value("timestamp[us]"),
"array": [Value("int64")],
}
)
dataset = IterableDataset(ex_iterable, info=DatasetInfo(features=features))
assert list(dataset) == [{"timestamp": pd.Timestamp(2020, 1, 1).to_pydatetime(), "array": [1] * 5, "id": 0}]
@pytest.mark.parametrize("format_type", [None, "torch", "python", "tf", "tensorflow", "np", "numpy", "jax"])
def test_iterable_dataset_with_format(dataset: IterableDataset, format_type):
formatted_dataset = dataset.with_format(format_type)
assert formatted_dataset._formatting.format_type == get_format_type_from_alias(format_type)
@require_torch
def test_iterable_dataset_is_torch_iterable_dataset(dataset: IterableDataset):
from torch.utils.data import DataLoader, _DatasetKind
dataloader = DataLoader(dataset)
assert dataloader._dataset_kind == _DatasetKind.Iterable
out = list(dataloader)
assert len(out) == DEFAULT_N_EXAMPLES
@pytest.mark.parametrize("n", [0, 2, int(1e10)])
def test_iterable_dataset_skip(dataset: IterableDataset, n):
skip_dataset = dataset.skip(n)
assert isinstance(skip_dataset._ex_iterable, SkipExamplesIterable)
assert skip_dataset._ex_iterable.n == n
assert list(skip_dataset) == list(dataset)[n:]
@pytest.mark.parametrize("n", [0, 2, int(1e10)])
def test_iterable_dataset_take(dataset: IterableDataset, n):
take_dataset = dataset.take(n)
assert isinstance(take_dataset._ex_iterable, TakeExamplesIterable)
assert take_dataset._ex_iterable.n == n
assert list(take_dataset) == list(dataset)[:n]
@pytest.mark.parametrize("method", ["skip", "take"])
def test_iterable_dataset_shuffle_after_skip_or_take(method):
seed = 42
n, n_shards = 3, 10
count = 7
ex_iterable = ExamplesIterable(generate_examples_fn, {"n": n, "filepaths": [f"{i}.txt" for i in range(n_shards)]})
dataset = IterableDataset(ex_iterable)
dataset = dataset.skip(n) if method == "skip" else dataset.take(count)
shuffled_dataset = dataset.shuffle(seed, buffer_size=DEFAULT_N_EXAMPLES)
# shuffling a skip/take dataset should keep the same examples and don't shuffle the shards
key = lambda x: f"{x['filepath']}_{x['id']}" # noqa: E731
assert sorted(dataset, key=key) == sorted(shuffled_dataset, key=key)
def test_iterable_dataset_add_column(dataset_with_several_columns):
new_column = list(range(DEFAULT_N_EXAMPLES))
new_dataset = dataset_with_several_columns.add_column("new_column", new_column)
assert list(new_dataset) == [
{**example, "new_column": idx} for idx, example in enumerate(dataset_with_several_columns)
]
new_dataset = new_dataset._resolve_features()
assert "new_column" in new_dataset.column_names
def test_iterable_dataset_rename_column(dataset_with_several_columns):
new_dataset = dataset_with_several_columns.rename_column("id", "new_id")
assert list(new_dataset) == [
{("new_id" if k == "id" else k): v for k, v in example.items()} for example in dataset_with_several_columns
]
assert new_dataset.features is None
assert new_dataset.column_names is None
# rename the column if ds.features was not None
new_dataset = dataset_with_several_columns._resolve_features().rename_column("id", "new_id")
assert new_dataset.features is not None
assert new_dataset.column_names is not None
assert "id" not in new_dataset.column_names
assert "new_id" in new_dataset.column_names
def test_iterable_dataset_rename_columns(dataset_with_several_columns):
column_mapping = {"id": "new_id", "filepath": "filename"}
new_dataset = dataset_with_several_columns.rename_columns(column_mapping)
assert list(new_dataset) == [
{column_mapping.get(k, k): v for k, v in example.items()} for example in dataset_with_several_columns
]
assert new_dataset.features is None
assert new_dataset.column_names is None
# rename the columns if ds.features was not None
new_dataset = dataset_with_several_columns._resolve_features().rename_columns(column_mapping)
assert new_dataset.features is not None
assert new_dataset.column_names is not None
assert all(c not in new_dataset.column_names for c in ["id", "filepath"])
assert all(c in new_dataset.column_names for c in ["new_id", "filename"])
def test_iterable_dataset_remove_columns(dataset_with_several_columns):
new_dataset = dataset_with_several_columns.remove_columns("id")
assert list(new_dataset) == [
{k: v for k, v in example.items() if k != "id"} for example in dataset_with_several_columns
]
assert new_dataset.features is None
new_dataset = dataset_with_several_columns.remove_columns(["id", "filepath"])
assert list(new_dataset) == [
{k: v for k, v in example.items() if k != "id" and k != "filepath"} for example in dataset_with_several_columns
]
assert new_dataset.features is None
assert new_dataset.column_names is None
# remove the columns if ds.features was not None
new_dataset = dataset_with_several_columns._resolve_features().remove_columns(["id", "filepath"])
assert new_dataset.features is not None
assert new_dataset.column_names is not None
assert all(c not in new_dataset.features for c in ["id", "filepath"])
assert all(c not in new_dataset.column_names for c in ["id", "filepath"])
def test_iterable_dataset_select_columns(dataset_with_several_columns):
new_dataset = dataset_with_several_columns.select_columns("id")
assert list(new_dataset) == [
{k: v for k, v in example.items() if k == "id"} for example in dataset_with_several_columns
]
assert new_dataset.features is None
new_dataset = dataset_with_several_columns.select_columns(["id", "filepath"])
assert list(new_dataset) == [
{k: v for k, v in example.items() if k in ("id", "filepath")} for example in dataset_with_several_columns
]
assert new_dataset.features is None
# select the columns if ds.features was not None
new_dataset = dataset_with_several_columns._resolve_features().select_columns(["id", "filepath"])
assert new_dataset.features is not None
assert new_dataset.column_names is not None
assert all(c in new_dataset.features for c in ["id", "filepath"])
assert all(c in new_dataset.column_names for c in ["id", "filepath"])
def test_iterable_dataset_cast_column():
ex_iterable = ExamplesIterable(generate_examples_fn, {"label": 10})
features = Features({"id": Value("int64"), "label": Value("int64")})
dataset = IterableDataset(ex_iterable, info=DatasetInfo(features=features))
casted_dataset = dataset.cast_column("label", Value("bool"))
casted_features = features.copy()
casted_features["label"] = Value("bool")
assert list(casted_dataset) == [casted_features.encode_example(ex) for _, ex in ex_iterable]
def test_iterable_dataset_cast():
ex_iterable = ExamplesIterable(generate_examples_fn, {"label": 10})
features = Features({"id": Value("int64"), "label": Value("int64")})
dataset = IterableDataset(ex_iterable, info=DatasetInfo(features=features))
new_features = Features({"id": Value("int64"), "label": Value("bool")})
casted_dataset = dataset.cast(new_features)
assert list(casted_dataset) == [new_features.encode_example(ex) for _, ex in ex_iterable]
def test_iterable_dataset_resolve_features():
ex_iterable = ExamplesIterable(generate_examples_fn, {})
dataset = IterableDataset(ex_iterable)
assert dataset.features is None
assert dataset.column_names is None
dataset = dataset._resolve_features()
assert dataset.features == Features(
{
"id": Value("int64"),
}
)
assert dataset.column_names == ["id"]
def test_iterable_dataset_resolve_features_keep_order():
def gen():
yield from zip(range(3), [{"a": 1}, {"c": 1}, {"b": 1}])
ex_iterable = ExamplesIterable(gen, {})
dataset = IterableDataset(ex_iterable)._resolve_features()
# columns appear in order of appearance in the dataset
assert list(dataset.features) == ["a", "c", "b"]
assert dataset.column_names == ["a", "c", "b"]
def test_iterable_dataset_with_features_fill_with_none():
def gen():
yield from zip(range(2), [{"a": 1}, {"b": 1}])
ex_iterable = ExamplesIterable(gen, {})
info = DatasetInfo(features=Features({"a": Value("int32"), "b": Value("int32")}))
dataset = IterableDataset(ex_iterable, info=info)
assert list(dataset) == [{"a": 1, "b": None}, {"b": 1, "a": None}]
def test_concatenate_datasets():
ex_iterable1 = ExamplesIterable(generate_examples_fn, {"label": 10})
dataset1 = IterableDataset(ex_iterable1)
ex_iterable2 = ExamplesIterable(generate_examples_fn, {"label": 5})
dataset2 = IterableDataset(ex_iterable2)
concatenated_dataset = concatenate_datasets([dataset1, dataset2])
assert list(concatenated_dataset) == list(dataset1) + list(dataset2)
def test_concatenate_datasets_resolves_features():
ex_iterable1 = ExamplesIterable(generate_examples_fn, {"label": 10})
dataset1 = IterableDataset(ex_iterable1)
ex_iterable2 = ExamplesIterable(generate_examples_fn, {"label": 5})
dataset2 = IterableDataset(ex_iterable2)
concatenated_dataset = concatenate_datasets([dataset1, dataset2])
assert concatenated_dataset.features is not None
assert sorted(concatenated_dataset.features) == ["id", "label"]
def test_concatenate_datasets_with_different_columns():
ex_iterable1 = ExamplesIterable(generate_examples_fn, {"label": 10})
dataset1 = IterableDataset(ex_iterable1)
ex_iterable2 = ExamplesIterable(generate_examples_fn, {})
dataset2 = IterableDataset(ex_iterable2)
# missing column "label" -> it should be replaced with nulls
extended_dataset2_list = [{"label": None, **x} for x in dataset2]
concatenated_dataset = concatenate_datasets([dataset1, dataset2])
assert list(concatenated_dataset) == list(dataset1) + extended_dataset2_list
# change order
concatenated_dataset = concatenate_datasets([dataset2, dataset1])
assert list(concatenated_dataset) == extended_dataset2_list + list(dataset1)
def test_concatenate_datasets_axis_1():
ex_iterable1 = ExamplesIterable(generate_examples_fn, {"label1": 10})
dataset1 = IterableDataset(ex_iterable1)
ex_iterable2 = ExamplesIterable(generate_examples_fn, {"label2": 5})
dataset2 = IterableDataset(ex_iterable2)
with pytest.raises(ValueError): # column "id" is duplicated -> raise an error
concatenate_datasets([dataset1, dataset2], axis=1)
concatenated_dataset = concatenate_datasets([dataset1, dataset2.remove_columns("id")], axis=1)
assert list(concatenated_dataset) == [{**x, **y} for x, y in zip(dataset1, dataset2)]
def test_concatenate_datasets_axis_1_resolves_features():
ex_iterable1 = ExamplesIterable(generate_examples_fn, {"label1": 10})
dataset1 = IterableDataset(ex_iterable1)
ex_iterable2 = ExamplesIterable(generate_examples_fn, {"label2": 5})
dataset2 = IterableDataset(ex_iterable2).remove_columns("id")
concatenated_dataset = concatenate_datasets([dataset1, dataset2], axis=1)
assert concatenated_dataset.features is not None
assert sorted(concatenated_dataset.features) == ["id", "label1", "label2"]
def test_concatenate_datasets_axis_1_with_different_lengths():
n1 = 10
ex_iterable1 = ExamplesIterable(generate_examples_fn, {"label1": 10, "n": n1})
dataset1 = IterableDataset(ex_iterable1)
n2 = 5
ex_iterable2 = ExamplesIterable(generate_examples_fn, {"label2": 5, "n": n2})
dataset2 = IterableDataset(ex_iterable2).remove_columns("id")
# missing rows -> they should be replaced with nulls
extended_dataset2_list = list(dataset2) + [{"label2": None}] * (n1 - n2)
concatenated_dataset = concatenate_datasets([dataset1, dataset2], axis=1)
assert list(concatenated_dataset) == [{**x, **y} for x, y in zip(dataset1, extended_dataset2_list)]
# change order
concatenated_dataset = concatenate_datasets([dataset2, dataset1], axis=1)
assert list(concatenated_dataset) == [{**x, **y} for x, y in zip(extended_dataset2_list, dataset1)]
@pytest.mark.parametrize(
"probas, seed, expected_length, stopping_strategy",
[
(None, None, 3 * (DEFAULT_N_EXAMPLES - 1) + 1, "first_exhausted"),
([1, 0, 0], None, DEFAULT_N_EXAMPLES, "first_exhausted"),
([0, 1, 0], None, DEFAULT_N_EXAMPLES, "first_exhausted"),
([0.2, 0.5, 0.3], 42, None, "first_exhausted"),
([0.1, 0.1, 0.8], 1337, None, "first_exhausted"),
([0.5, 0.2, 0.3], 101010, None, "first_exhausted"),
(None, None, 3 * DEFAULT_N_EXAMPLES, "all_exhausted"),
([0.2, 0.5, 0.3], 42, None, "all_exhausted"),
([0.1, 0.1, 0.8], 1337, None, "all_exhausted"),
([0.5, 0.2, 0.3], 101010, None, "all_exhausted"),
],
)
def test_interleave_datasets(dataset: IterableDataset, probas, seed, expected_length, stopping_strategy):
d1 = dataset
d2 = dataset.map(lambda x: {"id+1": x["id"] + 1, **x})
d3 = dataset.with_format("python")
datasets = [d1, d2, d3]
merged_dataset = interleave_datasets(
datasets, probabilities=probas, seed=seed, stopping_strategy=stopping_strategy
)
def fill_default(example):
return {"id": None, "id+1": None, **example}
# Check the examples iterable
assert isinstance(
merged_dataset._ex_iterable, (CyclingMultiSourcesExamplesIterable, RandomlyCyclingMultiSourcesExamplesIterable)
)
# Check that it is deterministic
if seed is not None:
merged_dataset2 = interleave_datasets(
[d1, d2, d3], probabilities=probas, seed=seed, stopping_strategy=stopping_strategy
)
assert list(merged_dataset) == list(merged_dataset2)
# Check features
assert merged_dataset.features == Features({"id": Value("int64"), "id+1": Value("int64")})
# Check first example
if seed is not None:
rng = np.random.default_rng(seed)
i = next(iter(RandomlyCyclingMultiSourcesExamplesIterable._iter_random_indices(rng, len(datasets), p=probas)))
assert next(iter(merged_dataset)) == fill_default(next(iter(datasets[i])))
else:
assert any(next(iter(merged_dataset)) == fill_default(next(iter(dataset))) for dataset in datasets)
# Compute length it case it's random
if expected_length is None:
expected_length = 0
counts = np.array([len(list(d)) for d in datasets])
bool_strategy_func = np.all if stopping_strategy == "all_exhausted" else np.any
rng = np.random.default_rng(seed)
for i in RandomlyCyclingMultiSourcesExamplesIterable._iter_random_indices(rng, len(datasets), p=probas):
counts[i] -= 1
expected_length += 1
if bool_strategy_func(counts <= 0):
break
# Check length
assert len(list(merged_dataset)) == expected_length
def test_interleave_datasets_with_features(
dataset: IterableDataset,
):
features = Features(
{
"id": Value("int64"),
"label": ClassLabel(names=["negative", "positive"]),
}
)
ex_iterable = ExamplesIterable(generate_examples_fn, {"label": 0})
dataset_with_features = IterableDataset(ex_iterable, info=DatasetInfo(features=features))
merged_dataset = interleave_datasets([dataset, dataset_with_features])
assert merged_dataset.features == features
def test_interleave_datasets_with_oversampling():
# Test hardcoded results
d1 = IterableDataset(ExamplesIterable((lambda: (yield from [(i, {"a": i}) for i in [0, 1, 2]])), {}))
d2 = IterableDataset(ExamplesIterable((lambda: (yield from [(i, {"a": i}) for i in [10, 11, 12, 13]])), {}))
d3 = IterableDataset(ExamplesIterable((lambda: (yield from [(i, {"a": i}) for i in [20, 21, 22, 23, 24]])), {}))
expected_values = [0, 10, 20, 1, 11, 21, 2, 12, 22, 0, 13, 23, 1, 10, 24]
# Check oversampling strategy without probabilities
assert [x["a"] for x in interleave_datasets([d1, d2, d3], stopping_strategy="all_exhausted")] == expected_values
# Check oversampling strategy with probabilities
expected_values = [20, 0, 21, 10, 1, 22, 23, 24, 2, 0, 1, 20, 11, 21, 2, 0, 12, 1, 22, 13]
values = [
x["a"]
for x in interleave_datasets(
[d1, d2, d3], probabilities=[0.5, 0.2, 0.3], seed=42, stopping_strategy="all_exhausted"
)
]
assert values == expected_values
@require_torch
def test_with_format_torch(dataset_with_several_columns: IterableDataset):
import torch
dset = dataset_with_several_columns.with_format(type="torch")
example = next(iter(dset))
batch = next(iter(dset.iter(batch_size=3)))
assert len(example) == 3
assert isinstance(example["id"], torch.Tensor)
assert list(example["id"].shape) == []
assert example["id"].item() == 0
assert isinstance(batch["id"], torch.Tensor)
assert isinstance(example["filepath"], list)
assert isinstance(example["filepath"][0], str)
assert example["filepath"][0] == "data0.txt"
assert isinstance(batch["filepath"], list)
assert isinstance(example["metadata"], dict)
assert isinstance(example["metadata"]["sources"], list)
assert isinstance(example["metadata"]["sources"][0], str)
assert isinstance(batch["metadata"], list)
@require_tf
def test_with_format_tf(dataset_with_several_columns: IterableDataset):
import tensorflow as tf
dset = dataset_with_several_columns.with_format(type="tensorflow")
example = next(iter(dset))
batch = next(iter(dset.iter(batch_size=3)))
assert isinstance(example["id"], tf.Tensor)
assert list(example["id"].shape) == []
assert example["id"].numpy().item() == 0
assert isinstance(batch["id"], tf.Tensor)
assert isinstance(example["filepath"], tf.Tensor)
assert example["filepath"][0] == b"data0.txt"
assert isinstance(batch["filepath"], tf.Tensor)
assert isinstance(example["metadata"], dict)
assert isinstance(example["metadata"]["sources"], tf.Tensor)
assert isinstance(batch["metadata"], list)
def test_map_array_are_not_converted_back_to_lists(dataset: IterableDataset):
def func(example):
return {"array": np.array([1, 2, 3])}
dset_test = dataset.map(func)
example = next(iter(dset_test))
# not aligned with Dataset.map because we don't convert back to lists after map()
assert isinstance(example["array"], np.ndarray)
def test_formatted_map(dataset: IterableDataset):
dataset = dataset.with_format("np")
assert isinstance(next(dataset.iter(batch_size=3))["id"], np.ndarray)
dataset = dataset.with_format(None)
assert isinstance(next(dataset.iter(batch_size=3))["id"], list)
def add_one_numpy(example):
assert isinstance(example["id"], np.ndarray)
return {"id": example["id"] + 1}
dataset = dataset.with_format("np")
dataset = dataset.map(add_one_numpy, batched=True)
assert isinstance(next(dataset.iter(batch_size=3))["id"], np.ndarray)
dataset = dataset.with_format(None)
assert isinstance(next(dataset.iter(batch_size=3))["id"], list)
@pytest.mark.parametrize("n_shards1, n_shards2, num_workers", [(2, 1, 1), (2, 2, 2), (1, 3, 1), (4, 3, 3)])
def test_interleave_dataset_with_sharding(n_shards1, n_shards2, num_workers):
from torch.utils.data import DataLoader
ex_iterable1 = ExamplesIterable(generate_examples_fn, {"filepaths": [f"{i}-1.txt" for i in range(n_shards1)]})
dataset1 = IterableDataset(ex_iterable1).with_format("torch")
ex_iterable2 = ExamplesIterable(generate_examples_fn, {"filepaths": [f"{i}-2.txt" for i in range(n_shards2)]})
dataset2 = IterableDataset(ex_iterable2).with_format("torch")
dataset_merged = interleave_datasets([dataset1, dataset2], stopping_strategy="first_exhausted")
assert dataset_merged.n_shards == min(n_shards1, n_shards2)
dataloader = DataLoader(dataset_merged, batch_size=None, num_workers=num_workers)
result = list(dataloader)
expected_length = 2 * min(
len([example for _, example in ex_iterable1]), len([example for _, example in ex_iterable2])
)
# some samples may be missing because the stopping strategy is applied per process
assert expected_length - num_workers <= len(result) <= expected_length
assert len(result) == len({str(x) for x in result})
def filter_func(batch):
return batch["id"] == 4
def map_func(batch):
batch["id"] *= 2
return batch
def test_pickle_after_many_transforms(dataset_with_several_columns):
dataset = dataset_with_several_columns
dataset = dataset.remove_columns(["filepath"])
dataset = dataset.take(5)
dataset = dataset.map(map_func)
dataset = dataset.shuffle()
dataset = dataset.skip(1)
dataset = dataset.filter(filter_func)
dataset = dataset.add_column("additional_col", ["something"])
dataset = dataset.rename_column("metadata", "metadata1")
dataset = dataset.rename_columns({"id": "id1", "metadata1": "metadata2"})
dataset = dataset.select_columns(["id1", "additional_col"])
unpickled_dataset = pickle.loads(pickle.dumps(dataset))
assert list(unpickled_dataset) == list(dataset)
| 0
|
hf_public_repos/datasets
|
hf_public_repos/datasets/tests/test_splits.py
|
import pytest
from datasets.splits import SplitDict, SplitInfo
from datasets.utils.py_utils import asdict
@pytest.mark.parametrize(
"split_dict",
[
SplitDict(),
SplitDict({"train": SplitInfo(name="train", num_bytes=1337, num_examples=42, dataset_name="my_dataset")}),
SplitDict({"train": SplitInfo(name="train", num_bytes=1337, num_examples=42)}),
SplitDict({"train": SplitInfo()}),
],
)
def test_split_dict_to_yaml_list(split_dict: SplitDict):
split_dict_yaml_list = split_dict._to_yaml_list()
assert len(split_dict_yaml_list) == len(split_dict)
reloaded = SplitDict._from_yaml_list(split_dict_yaml_list)
for split_name, split_info in split_dict.items():
# dataset_name field is deprecated, and is therefore not part of the YAML dump
split_info.dataset_name = None
# the split name of split_dict takes over the name of the split info object
split_info.name = split_name
assert split_dict == reloaded
@pytest.mark.parametrize(
"split_info", [SplitInfo(), SplitInfo(dataset_name=None), SplitInfo(dataset_name="my_dataset")]
)
def test_split_dict_asdict_has_dataset_name(split_info):
# For backward compatibility, we need asdict(split_dict) to return split info dictrionaries with the "dataset_name"
# field even if it's deprecated. This way old versionso of `datasets` can still reload dataset_infos.json files
split_dict_asdict = asdict(SplitDict({"train": split_info}))
assert "dataset_name" in split_dict_asdict["train"]
assert split_dict_asdict["train"]["dataset_name"] == split_info.dataset_name
| 0
|
hf_public_repos/datasets
|
hf_public_repos/datasets/tests/test_metric.py
|
import os
import pickle
import tempfile
import time
from multiprocessing import Pool
from unittest import TestCase
import pytest
from datasets.features import Features, Sequence, Value
from datasets.metric import Metric, MetricInfo
from .utils import require_tf, require_torch
class DummyMetric(Metric):
def _info(self):
return MetricInfo(
description="dummy metric for tests",
citation="insert citation here",
features=Features({"predictions": Value("int64"), "references": Value("int64")}),
)
def _compute(self, predictions, references):
return (
{
"accuracy": sum(i == j for i, j in zip(predictions, references)) / len(predictions),
"set_equality": set(predictions) == set(references),
}
if predictions
else {}
)
@classmethod
def predictions_and_references(cls):
return ([1, 2, 3, 4], [1, 2, 4, 3])
@classmethod
def expected_results(cls):
return {"accuracy": 0.5, "set_equality": True}
@classmethod
def other_predictions_and_references(cls):
return ([1, 3, 4, 5], [1, 2, 3, 4])
@classmethod
def other_expected_results(cls):
return {"accuracy": 0.25, "set_equality": False}
@classmethod
def distributed_predictions_and_references(cls):
return ([1, 2, 3, 4], [1, 2, 3, 4]), ([1, 2, 4, 5], [1, 2, 3, 4])
@classmethod
def distributed_expected_results(cls):
return {"accuracy": 0.75, "set_equality": False}
@classmethod
def separate_predictions_and_references(cls):
return ([1, 2, 3, 4], [1, 2, 3, 4]), ([1, 2, 4, 5], [1, 2, 3, 4])
@classmethod
def separate_expected_results(cls):
return [{"accuracy": 1.0, "set_equality": True}, {"accuracy": 0.5, "set_equality": False}]
def properly_del_metric(metric):
"""properly delete a metric on windows if the process is killed during multiprocessing"""
if metric is not None:
if metric.filelock is not None:
metric.filelock.release()
if metric.rendez_vous_lock is not None:
metric.rendez_vous_lock.release()
del metric.writer
del metric.data
del metric
def metric_compute(arg):
"""Thread worker function for distributed evaluation testing.
On base level to be pickable.
"""
metric = None
try:
num_process, process_id, preds, refs, exp_id, cache_dir, wait = arg
metric = DummyMetric(
num_process=num_process, process_id=process_id, experiment_id=exp_id, cache_dir=cache_dir, timeout=5
)
time.sleep(wait)
results = metric.compute(predictions=preds, references=refs)
return results
finally:
properly_del_metric(metric)
def metric_add_batch_and_compute(arg):
"""Thread worker function for distributed evaluation testing.
On base level to be pickable.
"""
metric = None
try:
num_process, process_id, preds, refs, exp_id, cache_dir, wait = arg
metric = DummyMetric(
num_process=num_process, process_id=process_id, experiment_id=exp_id, cache_dir=cache_dir, timeout=5
)
metric.add_batch(predictions=preds, references=refs)
time.sleep(wait)
results = metric.compute()
return results
finally:
properly_del_metric(metric)
def metric_add_and_compute(arg):
"""Thread worker function for distributed evaluation testing.
On base level to be pickable.
"""
metric = None
try:
num_process, process_id, preds, refs, exp_id, cache_dir, wait = arg
metric = DummyMetric(
num_process=num_process, process_id=process_id, experiment_id=exp_id, cache_dir=cache_dir, timeout=5
)
for pred, ref in zip(preds, refs):
metric.add(prediction=pred, reference=ref)
time.sleep(wait)
results = metric.compute()
return results
finally:
properly_del_metric(metric)
@pytest.mark.filterwarnings("ignore:Metric is deprecated:FutureWarning")
class TestMetric(TestCase):
def test_dummy_metric(self):
preds, refs = DummyMetric.predictions_and_references()
expected_results = DummyMetric.expected_results()
metric = DummyMetric(experiment_id="test_dummy_metric")
self.assertDictEqual(expected_results, metric.compute(predictions=preds, references=refs))
del metric
metric = DummyMetric(experiment_id="test_dummy_metric")
metric.add_batch(predictions=preds, references=refs)
self.assertDictEqual(expected_results, metric.compute())
del metric
metric = DummyMetric(experiment_id="test_dummy_metric")
for pred, ref in zip(preds, refs):
metric.add(prediction=pred, reference=ref)
self.assertDictEqual(expected_results, metric.compute())
del metric
# With keep_in_memory
metric = DummyMetric(keep_in_memory=True, experiment_id="test_dummy_metric")
self.assertDictEqual(expected_results, metric.compute(predictions=preds, references=refs))
del metric
metric = DummyMetric(keep_in_memory=True, experiment_id="test_dummy_metric")
metric.add_batch(predictions=preds, references=refs)
self.assertDictEqual(expected_results, metric.compute())
del metric
metric = DummyMetric(keep_in_memory=True, experiment_id="test_dummy_metric")
for pred, ref in zip(preds, refs):
metric.add(prediction=pred, reference=ref)
self.assertDictEqual(expected_results, metric.compute())
del metric
metric = DummyMetric(keep_in_memory=True, experiment_id="test_dummy_metric")
self.assertDictEqual({}, metric.compute(predictions=[], references=[]))
del metric
metric = DummyMetric(keep_in_memory=True, experiment_id="test_dummy_metric")
with self.assertRaisesRegex(ValueError, "Mismatch in the number"):
metric.add_batch(predictions=[1, 2, 3], references=[1, 2, 3, 4])
del metric
def test_metric_with_cache_dir(self):
preds, refs = DummyMetric.predictions_and_references()
expected_results = DummyMetric.expected_results()
with tempfile.TemporaryDirectory() as tmp_dir:
metric = DummyMetric(experiment_id="test_dummy_metric", cache_dir=tmp_dir)
self.assertDictEqual(expected_results, metric.compute(predictions=preds, references=refs))
del metric
def test_concurrent_metrics(self):
preds, refs = DummyMetric.predictions_and_references()
other_preds, other_refs = DummyMetric.other_predictions_and_references()
expected_results = DummyMetric.expected_results()
other_expected_results = DummyMetric.other_expected_results()
metric = DummyMetric(experiment_id="test_concurrent_metrics")
other_metric = DummyMetric(
experiment_id="test_concurrent_metrics",
)
self.assertDictEqual(expected_results, metric.compute(predictions=preds, references=refs))
self.assertDictEqual(
other_expected_results, other_metric.compute(predictions=other_preds, references=other_refs)
)
del metric, other_metric
metric = DummyMetric(
experiment_id="test_concurrent_metrics",
)
other_metric = DummyMetric(
experiment_id="test_concurrent_metrics",
)
metric.add_batch(predictions=preds, references=refs)
other_metric.add_batch(predictions=other_preds, references=other_refs)
self.assertDictEqual(expected_results, metric.compute())
self.assertDictEqual(other_expected_results, other_metric.compute())
for pred, ref, other_pred, other_ref in zip(preds, refs, other_preds, other_refs):
metric.add(prediction=pred, reference=ref)
other_metric.add(prediction=other_pred, reference=other_ref)
self.assertDictEqual(expected_results, metric.compute())
self.assertDictEqual(other_expected_results, other_metric.compute())
del metric, other_metric
# With keep_in_memory
metric = DummyMetric(experiment_id="test_concurrent_metrics", keep_in_memory=True)
other_metric = DummyMetric(experiment_id="test_concurrent_metrics", keep_in_memory=True)
self.assertDictEqual(expected_results, metric.compute(predictions=preds, references=refs))
self.assertDictEqual(
other_expected_results, other_metric.compute(predictions=other_preds, references=other_refs)
)
metric = DummyMetric(experiment_id="test_concurrent_metrics", keep_in_memory=True)
other_metric = DummyMetric(experiment_id="test_concurrent_metrics", keep_in_memory=True)
metric.add_batch(predictions=preds, references=refs)
other_metric.add_batch(predictions=other_preds, references=other_refs)
self.assertDictEqual(expected_results, metric.compute())
self.assertDictEqual(other_expected_results, other_metric.compute())
for pred, ref, other_pred, other_ref in zip(preds, refs, other_preds, other_refs):
metric.add(prediction=pred, reference=ref)
other_metric.add(prediction=other_pred, reference=other_ref)
self.assertDictEqual(expected_results, metric.compute())
self.assertDictEqual(other_expected_results, other_metric.compute())
del metric, other_metric
def test_separate_experiments_in_parallel(self):
with tempfile.TemporaryDirectory() as tmp_dir:
(preds_0, refs_0), (preds_1, refs_1) = DummyMetric.separate_predictions_and_references()
expected_results = DummyMetric.separate_expected_results()
pool = Pool(processes=4)
results = pool.map(
metric_compute,
[
(1, 0, preds_0, refs_0, None, tmp_dir, 0),
(1, 0, preds_1, refs_1, None, tmp_dir, 0),
],
)
self.assertDictEqual(expected_results[0], results[0])
self.assertDictEqual(expected_results[1], results[1])
del results
# more than one sec of waiting so that the second metric has to sample a new hashing name
results = pool.map(
metric_compute,
[
(1, 0, preds_0, refs_0, None, tmp_dir, 2),
(1, 0, preds_1, refs_1, None, tmp_dir, 2),
],
)
self.assertDictEqual(expected_results[0], results[0])
self.assertDictEqual(expected_results[1], results[1])
del results
results = pool.map(
metric_add_and_compute,
[
(1, 0, preds_0, refs_0, None, tmp_dir, 0),
(1, 0, preds_1, refs_1, None, tmp_dir, 0),
],
)
self.assertDictEqual(expected_results[0], results[0])
self.assertDictEqual(expected_results[1], results[1])
del results
results = pool.map(
metric_add_batch_and_compute,
[
(1, 0, preds_0, refs_0, None, tmp_dir, 0),
(1, 0, preds_1, refs_1, None, tmp_dir, 0),
],
)
self.assertDictEqual(expected_results[0], results[0])
self.assertDictEqual(expected_results[1], results[1])
del results
def test_distributed_metrics(self):
with tempfile.TemporaryDirectory() as tmp_dir:
(preds_0, refs_0), (preds_1, refs_1) = DummyMetric.distributed_predictions_and_references()
expected_results = DummyMetric.distributed_expected_results()
pool = Pool(processes=4)
results = pool.map(
metric_compute,
[
(2, 0, preds_0, refs_0, "test_distributed_metrics_0", tmp_dir, 0),
(2, 1, preds_1, refs_1, "test_distributed_metrics_0", tmp_dir, 0.5),
],
)
self.assertDictEqual(expected_results, results[0])
self.assertIsNone(results[1])
del results
results = pool.map(
metric_compute,
[
(2, 0, preds_0, refs_0, "test_distributed_metrics_0", tmp_dir, 0.5),
(2, 1, preds_1, refs_1, "test_distributed_metrics_0", tmp_dir, 0),
],
)
self.assertDictEqual(expected_results, results[0])
self.assertIsNone(results[1])
del results
results = pool.map(
metric_add_and_compute,
[
(2, 0, preds_0, refs_0, "test_distributed_metrics_1", tmp_dir, 0),
(2, 1, preds_1, refs_1, "test_distributed_metrics_1", tmp_dir, 0),
],
)
self.assertDictEqual(expected_results, results[0])
self.assertIsNone(results[1])
del results
results = pool.map(
metric_add_batch_and_compute,
[
(2, 0, preds_0, refs_0, "test_distributed_metrics_2", tmp_dir, 0),
(2, 1, preds_1, refs_1, "test_distributed_metrics_2", tmp_dir, 0),
],
)
self.assertDictEqual(expected_results, results[0])
self.assertIsNone(results[1])
del results
# To use several distributed metrics on the same local file system, need to specify an experiment_id
try:
results = pool.map(
metric_add_and_compute,
[
(2, 0, preds_0, refs_0, "test_distributed_metrics_3", tmp_dir, 0),
(2, 1, preds_1, refs_1, "test_distributed_metrics_3", tmp_dir, 0),
(2, 0, preds_0, refs_0, "test_distributed_metrics_3", tmp_dir, 0),
(2, 1, preds_1, refs_1, "test_distributed_metrics_3", tmp_dir, 0),
],
)
except ValueError:
# We are fine with either raising a ValueError or computing well the metric
# Being sure we raise the error would means making the dummy dataset bigger
# and the test longer...
pass
else:
self.assertDictEqual(expected_results, results[0])
self.assertDictEqual(expected_results, results[2])
self.assertIsNone(results[1])
self.assertIsNone(results[3])
del results
results = pool.map(
metric_add_and_compute,
[
(2, 0, preds_0, refs_0, "exp_0", tmp_dir, 0),
(2, 1, preds_1, refs_1, "exp_0", tmp_dir, 0),
(2, 0, preds_0, refs_0, "exp_1", tmp_dir, 0),
(2, 1, preds_1, refs_1, "exp_1", tmp_dir, 0),
],
)
self.assertDictEqual(expected_results, results[0])
self.assertDictEqual(expected_results, results[2])
self.assertIsNone(results[1])
self.assertIsNone(results[3])
del results
# With keep_in_memory is not allowed
with self.assertRaises(ValueError):
DummyMetric(
experiment_id="test_distributed_metrics_4",
keep_in_memory=True,
num_process=2,
process_id=0,
cache_dir=tmp_dir,
)
def test_dummy_metric_pickle(self):
with tempfile.TemporaryDirectory() as tmp_dir:
tmp_file = os.path.join(tmp_dir, "metric.pt")
preds, refs = DummyMetric.predictions_and_references()
expected_results = DummyMetric.expected_results()
metric = DummyMetric(experiment_id="test_dummy_metric_pickle")
with open(tmp_file, "wb") as f:
pickle.dump(metric, f)
del metric
with open(tmp_file, "rb") as f:
metric = pickle.load(f)
self.assertDictEqual(expected_results, metric.compute(predictions=preds, references=refs))
del metric
def test_input_numpy(self):
import numpy as np
preds, refs = DummyMetric.predictions_and_references()
expected_results = DummyMetric.expected_results()
preds, refs = np.array(preds), np.array(refs)
metric = DummyMetric(experiment_id="test_input_numpy")
self.assertDictEqual(expected_results, metric.compute(predictions=preds, references=refs))
del metric
metric = DummyMetric(experiment_id="test_input_numpy")
metric.add_batch(predictions=preds, references=refs)
self.assertDictEqual(expected_results, metric.compute())
del metric
metric = DummyMetric(experiment_id="test_input_numpy")
for pred, ref in zip(preds, refs):
metric.add(prediction=pred, reference=ref)
self.assertDictEqual(expected_results, metric.compute())
del metric
@require_torch
def test_input_torch(self):
import torch
preds, refs = DummyMetric.predictions_and_references()
expected_results = DummyMetric.expected_results()
preds, refs = torch.tensor(preds), torch.tensor(refs)
metric = DummyMetric(experiment_id="test_input_torch")
self.assertDictEqual(expected_results, metric.compute(predictions=preds, references=refs))
del metric
metric = DummyMetric(experiment_id="test_input_torch")
metric.add_batch(predictions=preds, references=refs)
self.assertDictEqual(expected_results, metric.compute())
del metric
metric = DummyMetric(experiment_id="test_input_torch")
for pred, ref in zip(preds, refs):
metric.add(prediction=pred, reference=ref)
self.assertDictEqual(expected_results, metric.compute())
del metric
@require_tf
def test_input_tf(self):
import tensorflow as tf
preds, refs = DummyMetric.predictions_and_references()
expected_results = DummyMetric.expected_results()
preds, refs = tf.constant(preds), tf.constant(refs)
metric = DummyMetric(experiment_id="test_input_tf")
self.assertDictEqual(expected_results, metric.compute(predictions=preds, references=refs))
del metric
metric = DummyMetric(experiment_id="test_input_tf")
metric.add_batch(predictions=preds, references=refs)
self.assertDictEqual(expected_results, metric.compute())
del metric
metric = DummyMetric(experiment_id="test_input_tf")
for pred, ref in zip(preds, refs):
metric.add(prediction=pred, reference=ref)
self.assertDictEqual(expected_results, metric.compute())
del metric
class MetricWithMultiLabel(Metric):
def _info(self):
return MetricInfo(
description="dummy metric for tests",
citation="insert citation here",
features=Features(
{"predictions": Sequence(Value("int64")), "references": Sequence(Value("int64"))}
if self.config_name == "multilabel"
else {"predictions": Value("int64"), "references": Value("int64")}
),
)
def _compute(self, predictions=None, references=None):
return (
{
"accuracy": sum(i == j for i, j in zip(predictions, references)) / len(predictions),
}
if predictions
else {}
)
@pytest.mark.parametrize(
"config_name, predictions, references, expected",
[
(None, [1, 2, 3, 4], [1, 2, 4, 3], 0.5), # Multiclass: Value("int64")
(
"multilabel",
[[1, 0], [1, 0], [1, 0], [1, 0]],
[[1, 0], [0, 1], [1, 1], [0, 0]],
0.25,
), # Multilabel: Sequence(Value("int64"))
],
)
def test_metric_with_multilabel(config_name, predictions, references, expected, tmp_path):
cache_dir = tmp_path / "cache"
metric = MetricWithMultiLabel(config_name, cache_dir=cache_dir)
results = metric.compute(predictions=predictions, references=references)
assert results["accuracy"] == expected
def test_safety_checks_process_vars():
with pytest.raises(ValueError):
_ = DummyMetric(process_id=-2)
with pytest.raises(ValueError):
_ = DummyMetric(num_process=2, process_id=3)
class AccuracyWithNonStandardFeatureNames(Metric):
def _info(self):
return MetricInfo(
description="dummy metric for tests",
citation="insert citation here",
features=Features({"inputs": Value("int64"), "targets": Value("int64")}),
)
def _compute(self, inputs, targets):
return (
{
"accuracy": sum(i == j for i, j in zip(inputs, targets)) / len(targets),
}
if targets
else {}
)
@classmethod
def inputs_and_targets(cls):
return ([1, 2, 3, 4], [1, 2, 4, 3])
@classmethod
def expected_results(cls):
return {"accuracy": 0.5}
def test_metric_with_non_standard_feature_names_add(tmp_path):
cache_dir = tmp_path / "cache"
inputs, targets = AccuracyWithNonStandardFeatureNames.inputs_and_targets()
metric = AccuracyWithNonStandardFeatureNames(cache_dir=cache_dir)
for input, target in zip(inputs, targets):
metric.add(inputs=input, targets=target)
results = metric.compute()
assert results == AccuracyWithNonStandardFeatureNames.expected_results()
def test_metric_with_non_standard_feature_names_add_batch(tmp_path):
cache_dir = tmp_path / "cache"
inputs, targets = AccuracyWithNonStandardFeatureNames.inputs_and_targets()
metric = AccuracyWithNonStandardFeatureNames(cache_dir=cache_dir)
metric.add_batch(inputs=inputs, targets=targets)
results = metric.compute()
assert results == AccuracyWithNonStandardFeatureNames.expected_results()
def test_metric_with_non_standard_feature_names_compute(tmp_path):
cache_dir = tmp_path / "cache"
inputs, targets = AccuracyWithNonStandardFeatureNames.inputs_and_targets()
metric = AccuracyWithNonStandardFeatureNames(cache_dir=cache_dir)
results = metric.compute(inputs=inputs, targets=targets)
assert results == AccuracyWithNonStandardFeatureNames.expected_results()
| 0
|
hf_public_repos/datasets
|
hf_public_repos/datasets/tests/test_filelock.py
|
import os
from datasets.utils._filelock import FileLock
def test_long_path(tmpdir):
filename = "a" * 1000 + ".lock"
lock1 = FileLock(str(tmpdir / filename))
assert lock1.lock_file.endswith(".lock")
assert not lock1.lock_file.endswith(filename)
assert len(os.path.basename(lock1.lock_file)) <= 255
| 0
|
hf_public_repos/datasets
|
hf_public_repos/datasets/tests/test_sharding_utils.py
|
import pytest
from datasets.utils.sharding import _distribute_shards, _number_of_shards_in_gen_kwargs, _split_gen_kwargs
@pytest.mark.parametrize(
"kwargs, expected",
[
({"num_shards": 0, "max_num_jobs": 1}, []),
({"num_shards": 10, "max_num_jobs": 1}, [range(10)]),
({"num_shards": 10, "max_num_jobs": 10}, [range(i, i + 1) for i in range(10)]),
({"num_shards": 1, "max_num_jobs": 10}, [range(1)]),
({"num_shards": 10, "max_num_jobs": 3}, [range(0, 4), range(4, 7), range(7, 10)]),
({"num_shards": 3, "max_num_jobs": 10}, [range(0, 1), range(1, 2), range(2, 3)]),
],
)
def test_distribute_shards(kwargs, expected):
out = _distribute_shards(**kwargs)
assert out == expected
@pytest.mark.parametrize(
"gen_kwargs, max_num_jobs, expected",
[
({"foo": 0}, 10, [{"foo": 0}]),
({"shards": [0, 1, 2, 3]}, 1, [{"shards": [0, 1, 2, 3]}]),
({"shards": [0, 1, 2, 3]}, 4, [{"shards": [0]}, {"shards": [1]}, {"shards": [2]}, {"shards": [3]}]),
({"shards": [0, 1]}, 4, [{"shards": [0]}, {"shards": [1]}]),
({"shards": [0, 1, 2, 3]}, 2, [{"shards": [0, 1]}, {"shards": [2, 3]}]),
],
)
def test_split_gen_kwargs(gen_kwargs, max_num_jobs, expected):
out = _split_gen_kwargs(gen_kwargs, max_num_jobs)
assert out == expected
@pytest.mark.parametrize(
"gen_kwargs, expected",
[
({"foo": 0}, 1),
({"shards": [0]}, 1),
({"shards": [0, 1, 2, 3]}, 4),
({"shards": [0, 1, 2, 3], "foo": 0}, 4),
({"shards": [0, 1, 2, 3], "other": (0, 1)}, 4),
({"shards": [0, 1, 2, 3], "shards2": [0, 1]}, RuntimeError),
],
)
def test_number_of_shards_in_gen_kwargs(gen_kwargs, expected):
if expected is RuntimeError:
with pytest.raises(expected):
_number_of_shards_in_gen_kwargs(gen_kwargs)
else:
out = _number_of_shards_in_gen_kwargs(gen_kwargs)
assert out == expected
| 0
|
hf_public_repos/datasets
|
hf_public_repos/datasets/tests/test_upstream_hub.py
|
import fnmatch
import gc
import os
import shutil
import tempfile
import textwrap
import time
import unittest
from io import BytesIO
from pathlib import Path
from unittest.mock import patch
import numpy as np
import pytest
from huggingface_hub import DatasetCard, HfApi
from datasets import (
Audio,
ClassLabel,
Dataset,
DatasetDict,
DownloadManager,
Features,
Image,
Value,
load_dataset,
load_dataset_builder,
)
from datasets.config import METADATA_CONFIGS_FIELD
from datasets.data_files import get_data_patterns
from datasets.packaged_modules.folder_based_builder.folder_based_builder import (
FolderBasedBuilder,
FolderBasedBuilderConfig,
)
from datasets.utils.file_utils import cached_path
from datasets.utils.hub import hf_hub_url
from tests.fixtures.hub import CI_HUB_ENDPOINT, CI_HUB_USER, CI_HUB_USER_TOKEN
from tests.utils import for_all_test_methods, require_pil, require_sndfile, xfail_if_500_502_http_error
pytestmark = pytest.mark.integration
@for_all_test_methods(xfail_if_500_502_http_error)
@pytest.mark.usefixtures("ci_hub_config", "ci_hfh_hf_hub_url")
class TestPushToHub:
_api = HfApi(endpoint=CI_HUB_ENDPOINT)
_token = CI_HUB_USER_TOKEN
def test_push_dataset_dict_to_hub_no_token(self, temporary_repo, set_ci_hub_access_token):
ds = Dataset.from_dict({"x": [1, 2, 3], "y": [4, 5, 6]})
local_ds = DatasetDict({"train": ds})
with temporary_repo() as ds_name:
local_ds.push_to_hub(ds_name)
hub_ds = load_dataset(ds_name, download_mode="force_redownload")
assert local_ds.column_names == hub_ds.column_names
assert list(local_ds["train"].features.keys()) == list(hub_ds["train"].features.keys())
assert local_ds["train"].features == hub_ds["train"].features
# Ensure that there is a single file on the repository that has the correct name
files = sorted(self._api.list_repo_files(ds_name, repo_type="dataset"))
assert files == [".gitattributes", "README.md", "data/train-00000-of-00001.parquet"]
def test_push_dataset_dict_to_hub_name_without_namespace(self, temporary_repo):
ds = Dataset.from_dict({"x": [1, 2, 3], "y": [4, 5, 6]})
local_ds = DatasetDict({"train": ds})
with temporary_repo() as ds_name:
local_ds.push_to_hub(ds_name.split("/")[-1], token=self._token)
hub_ds = load_dataset(ds_name, download_mode="force_redownload")
assert local_ds.column_names == hub_ds.column_names
assert list(local_ds["train"].features.keys()) == list(hub_ds["train"].features.keys())
assert local_ds["train"].features == hub_ds["train"].features
# Ensure that there is a single file on the repository that has the correct name
files = sorted(self._api.list_repo_files(ds_name, repo_type="dataset"))
assert files == [".gitattributes", "README.md", "data/train-00000-of-00001.parquet"]
def test_push_dataset_dict_to_hub_datasets_with_different_features(self, cleanup_repo):
ds_train = Dataset.from_dict({"x": [1, 2, 3], "y": [4, 5, 6]})
ds_test = Dataset.from_dict({"x": [True, False, True], "y": ["a", "b", "c"]})
local_ds = DatasetDict({"train": ds_train, "test": ds_test})
ds_name = f"{CI_HUB_USER}/test-{int(time.time() * 10e6)}"
try:
with pytest.raises(ValueError):
local_ds.push_to_hub(ds_name.split("/")[-1], token=self._token)
except AssertionError:
cleanup_repo(ds_name)
raise
def test_push_dataset_dict_to_hub_private(self, temporary_repo):
ds = Dataset.from_dict({"x": [1, 2, 3], "y": [4, 5, 6]})
local_ds = DatasetDict({"train": ds})
with temporary_repo() as ds_name:
local_ds.push_to_hub(ds_name, token=self._token, private=True)
hub_ds = load_dataset(ds_name, download_mode="force_redownload", token=self._token)
assert local_ds.column_names == hub_ds.column_names
assert list(local_ds["train"].features.keys()) == list(hub_ds["train"].features.keys())
assert local_ds["train"].features == hub_ds["train"].features
# Ensure that there is a single file on the repository that has the correct name
files = sorted(self._api.list_repo_files(ds_name, repo_type="dataset", token=self._token))
assert files == [".gitattributes", "README.md", "data/train-00000-of-00001.parquet"]
def test_push_dataset_dict_to_hub(self, temporary_repo):
ds = Dataset.from_dict({"x": [1, 2, 3], "y": [4, 5, 6]})
local_ds = DatasetDict({"train": ds})
with temporary_repo() as ds_name:
local_ds.push_to_hub(ds_name, token=self._token)
hub_ds = load_dataset(ds_name, download_mode="force_redownload")
assert local_ds.column_names == hub_ds.column_names
assert list(local_ds["train"].features.keys()) == list(hub_ds["train"].features.keys())
assert local_ds["train"].features == hub_ds["train"].features
# Ensure that there is a single file on the repository that has the correct name
files = sorted(self._api.list_repo_files(ds_name, repo_type="dataset", token=self._token))
assert files == [".gitattributes", "README.md", "data/train-00000-of-00001.parquet"]
def test_push_dataset_dict_to_hub_with_pull_request(self, temporary_repo):
ds = Dataset.from_dict({"x": [1, 2, 3], "y": [4, 5, 6]})
local_ds = DatasetDict({"train": ds})
with temporary_repo() as ds_name:
local_ds.push_to_hub(ds_name, token=self._token, create_pr=True)
hub_ds = load_dataset(ds_name, revision="refs/pr/1", download_mode="force_redownload")
assert local_ds["train"].features == hub_ds["train"].features
assert list(local_ds.keys()) == list(hub_ds.keys())
assert local_ds["train"].features == hub_ds["train"].features
# Ensure that there is a single file on the repository that has the correct name
files = sorted(
self._api.list_repo_files(ds_name, revision="refs/pr/1", repo_type="dataset", token=self._token)
)
assert files == [".gitattributes", "README.md", "data/train-00000-of-00001.parquet"]
def test_push_dataset_dict_to_hub_with_revision(self, temporary_repo):
ds = Dataset.from_dict({"x": [1, 2, 3], "y": [4, 5, 6]})
local_ds = DatasetDict({"train": ds})
with temporary_repo() as ds_name:
local_ds.push_to_hub(ds_name, token=self._token, revision="dev")
hub_ds = load_dataset(ds_name, revision="dev", download_mode="force_redownload")
assert local_ds["train"].features == hub_ds["train"].features
assert list(local_ds.keys()) == list(hub_ds.keys())
assert local_ds["train"].features == hub_ds["train"].features
# Ensure that there is a single file on the repository that has the correct name
files = sorted(self._api.list_repo_files(ds_name, revision="dev", repo_type="dataset", token=self._token))
assert files == [".gitattributes", "README.md", "data/train-00000-of-00001.parquet"]
def test_push_dataset_dict_to_hub_multiple_files(self, temporary_repo):
ds = Dataset.from_dict({"x": list(range(1000)), "y": list(range(1000))})
local_ds = DatasetDict({"train": ds})
with temporary_repo() as ds_name:
with patch("datasets.config.MAX_SHARD_SIZE", "16KB"):
local_ds.push_to_hub(ds_name, token=self._token)
hub_ds = load_dataset(ds_name, download_mode="force_redownload")
assert local_ds.column_names == hub_ds.column_names
assert list(local_ds["train"].features.keys()) == list(hub_ds["train"].features.keys())
assert local_ds["train"].features == hub_ds["train"].features
# Ensure that there are two files on the repository that have the correct name
files = sorted(self._api.list_repo_files(ds_name, repo_type="dataset", token=self._token))
assert files == [
".gitattributes",
"README.md",
"data/train-00000-of-00002.parquet",
"data/train-00001-of-00002.parquet",
]
def test_push_dataset_dict_to_hub_multiple_files_with_max_shard_size(self, temporary_repo):
ds = Dataset.from_dict({"x": list(range(1000)), "y": list(range(1000))})
local_ds = DatasetDict({"train": ds})
with temporary_repo() as ds_name:
local_ds.push_to_hub(ds_name, token=self._token, max_shard_size="16KB")
hub_ds = load_dataset(ds_name, download_mode="force_redownload")
assert local_ds.column_names == hub_ds.column_names
assert list(local_ds["train"].features.keys()) == list(hub_ds["train"].features.keys())
assert local_ds["train"].features == hub_ds["train"].features
# Ensure that there are two files on the repository that have the correct name
files = sorted(self._api.list_repo_files(ds_name, repo_type="dataset", token=self._token))
assert files == [
".gitattributes",
"README.md",
"data/train-00000-of-00002.parquet",
"data/train-00001-of-00002.parquet",
]
def test_push_dataset_dict_to_hub_multiple_files_with_num_shards(self, temporary_repo):
ds = Dataset.from_dict({"x": list(range(1000)), "y": list(range(1000))})
local_ds = DatasetDict({"train": ds})
with temporary_repo() as ds_name:
local_ds.push_to_hub(ds_name, token=self._token, num_shards={"train": 2})
hub_ds = load_dataset(ds_name, download_mode="force_redownload")
assert local_ds.column_names == hub_ds.column_names
assert list(local_ds["train"].features.keys()) == list(hub_ds["train"].features.keys())
assert local_ds["train"].features == hub_ds["train"].features
# Ensure that there are two files on the repository that have the correct name
files = sorted(self._api.list_repo_files(ds_name, repo_type="dataset", token=self._token))
assert files == [
".gitattributes",
"README.md",
"data/train-00000-of-00002.parquet",
"data/train-00001-of-00002.parquet",
]
def test_push_dataset_dict_to_hub_with_multiple_commits(self, temporary_repo):
ds = Dataset.from_dict({"x": list(range(1000)), "y": list(range(1000))})
local_ds = DatasetDict({"train": ds})
with temporary_repo() as ds_name:
self._api.create_repo(ds_name, token=self._token, repo_type="dataset")
num_commits_before_push = len(self._api.list_repo_commits(ds_name, repo_type="dataset", token=self._token))
with patch("datasets.config.MAX_SHARD_SIZE", "16KB"), patch(
"datasets.config.UPLOADS_MAX_NUMBER_PER_COMMIT", 1
):
local_ds.push_to_hub(ds_name, token=self._token)
hub_ds = load_dataset(ds_name, download_mode="force_redownload")
assert local_ds.column_names == hub_ds.column_names
assert list(local_ds["train"].features.keys()) == list(hub_ds["train"].features.keys())
assert local_ds["train"].features == hub_ds["train"].features
# Ensure that there are two files on the repository that have the correct name
files = sorted(self._api.list_repo_files(ds_name, repo_type="dataset", token=self._token))
assert files == [
".gitattributes",
"README.md",
"data/train-00000-of-00002.parquet",
"data/train-00001-of-00002.parquet",
]
num_commits_after_push = len(self._api.list_repo_commits(ds_name, repo_type="dataset", token=self._token))
assert num_commits_after_push - num_commits_before_push > 1
def test_push_dataset_dict_to_hub_overwrite_files(self, temporary_repo):
ds = Dataset.from_dict({"x": list(range(1000)), "y": list(range(1000))})
ds2 = Dataset.from_dict({"x": list(range(100)), "y": list(range(100))})
local_ds = DatasetDict({"train": ds, "random": ds2})
# Push to hub two times, but the second time with a larger amount of files.
# Verify that the new files contain the correct dataset.
with temporary_repo() as ds_name:
local_ds.push_to_hub(ds_name, token=self._token)
with tempfile.TemporaryDirectory() as tmp:
# Add a file starting with "data" to ensure it doesn't get deleted.
path = Path(tmp) / "datafile.txt"
with open(path, "w") as f:
f.write("Bogus file")
self._api.upload_file(
path_or_fileobj=str(path),
path_in_repo="datafile.txt",
repo_id=ds_name,
repo_type="dataset",
token=self._token,
)
local_ds.push_to_hub(ds_name, token=self._token, max_shard_size=500 << 5)
# Ensure that there are two files on the repository that have the correct name
files = sorted(self._api.list_repo_files(ds_name, repo_type="dataset", token=self._token))
assert files == [
".gitattributes",
"README.md",
"data/random-00000-of-00001.parquet",
"data/train-00000-of-00002.parquet",
"data/train-00001-of-00002.parquet",
"datafile.txt",
]
self._api.delete_file("datafile.txt", repo_id=ds_name, repo_type="dataset", token=self._token)
hub_ds = load_dataset(ds_name, download_mode="force_redownload")
assert local_ds.column_names == hub_ds.column_names
assert list(local_ds["train"].features.keys()) == list(hub_ds["train"].features.keys())
assert local_ds["train"].features == hub_ds["train"].features
del hub_ds
# To ensure the reference to the memory-mapped Arrow file is dropped to avoid the PermissionError on Windows
gc.collect()
# Push to hub two times, but the second time with fewer files.
# Verify that the new files contain the correct dataset and that non-necessary files have been deleted.
with temporary_repo(ds_name):
local_ds.push_to_hub(ds_name, token=self._token, max_shard_size=500 << 5)
with tempfile.TemporaryDirectory() as tmp:
# Add a file starting with "data" to ensure it doesn't get deleted.
path = Path(tmp) / "datafile.txt"
with open(path, "w") as f:
f.write("Bogus file")
self._api.upload_file(
path_or_fileobj=str(path),
path_in_repo="datafile.txt",
repo_id=ds_name,
repo_type="dataset",
token=self._token,
)
local_ds.push_to_hub(ds_name, token=self._token)
# Ensure that there are two files on the repository that have the correct name
files = sorted(self._api.list_repo_files(ds_name, repo_type="dataset", token=self._token))
assert files == [
".gitattributes",
"README.md",
"data/random-00000-of-00001.parquet",
"data/train-00000-of-00001.parquet",
"datafile.txt",
]
# Keeping the "datafile.txt" breaks the load_dataset to think it's a text-based dataset
self._api.delete_file("datafile.txt", repo_id=ds_name, repo_type="dataset", token=self._token)
hub_ds = load_dataset(ds_name, download_mode="force_redownload")
assert local_ds.column_names == hub_ds.column_names
assert list(local_ds["train"].features.keys()) == list(hub_ds["train"].features.keys())
assert local_ds["train"].features == hub_ds["train"].features
def test_push_dataset_to_hub(self, temporary_repo):
local_ds = Dataset.from_dict({"x": [1, 2, 3], "y": [4, 5, 6]})
with temporary_repo() as ds_name:
local_ds.push_to_hub(ds_name, split="train", token=self._token)
local_ds_dict = {"train": local_ds}
hub_ds_dict = load_dataset(ds_name, download_mode="force_redownload")
assert list(local_ds_dict.keys()) == list(hub_ds_dict.keys())
for ds_split_name in local_ds_dict.keys():
local_ds = local_ds_dict[ds_split_name]
hub_ds = hub_ds_dict[ds_split_name]
assert local_ds.column_names == hub_ds.column_names
assert list(local_ds.features.keys()) == list(hub_ds.features.keys())
assert local_ds.features == hub_ds.features
def test_push_dataset_to_hub_custom_features(self, temporary_repo):
features = Features({"x": Value("int64"), "y": ClassLabel(names=["neg", "pos"])})
ds = Dataset.from_dict({"x": [1, 2, 3], "y": [0, 0, 1]}, features=features)
with temporary_repo() as ds_name:
ds.push_to_hub(ds_name, token=self._token)
hub_ds = load_dataset(ds_name, split="train", download_mode="force_redownload")
assert ds.column_names == hub_ds.column_names
assert list(ds.features.keys()) == list(hub_ds.features.keys())
assert ds.features == hub_ds.features
assert ds[:] == hub_ds[:]
@require_sndfile
def test_push_dataset_to_hub_custom_features_audio(self, temporary_repo):
audio_path = os.path.join(os.path.dirname(__file__), "features", "data", "test_audio_44100.wav")
data = {"x": [audio_path, None], "y": [0, -1]}
features = Features({"x": Audio(), "y": Value("int32")})
ds = Dataset.from_dict(data, features=features)
for embed_external_files in [True, False]:
with temporary_repo() as ds_name:
ds.push_to_hub(ds_name, embed_external_files=embed_external_files, token=self._token)
hub_ds = load_dataset(ds_name, split="train", download_mode="force_redownload")
assert ds.column_names == hub_ds.column_names
assert list(ds.features.keys()) == list(hub_ds.features.keys())
assert ds.features == hub_ds.features
np.testing.assert_equal(ds[0]["x"]["array"], hub_ds[0]["x"]["array"])
assert ds[1] == hub_ds[1] # don't test hub_ds[0] since audio decoding might be slightly different
hub_ds = hub_ds.cast_column("x", Audio(decode=False))
elem = hub_ds[0]["x"]
path, bytes_ = elem["path"], elem["bytes"]
assert isinstance(path, str)
assert os.path.basename(path) == "test_audio_44100.wav"
assert bool(bytes_) == embed_external_files
@require_pil
def test_push_dataset_to_hub_custom_features_image(self, temporary_repo):
image_path = os.path.join(os.path.dirname(__file__), "features", "data", "test_image_rgb.jpg")
data = {"x": [image_path, None], "y": [0, -1]}
features = Features({"x": Image(), "y": Value("int32")})
ds = Dataset.from_dict(data, features=features)
for embed_external_files in [True, False]:
with temporary_repo() as ds_name:
ds.push_to_hub(ds_name, embed_external_files=embed_external_files, token=self._token)
hub_ds = load_dataset(ds_name, split="train", download_mode="force_redownload")
assert ds.column_names == hub_ds.column_names
assert list(ds.features.keys()) == list(hub_ds.features.keys())
assert ds.features == hub_ds.features
assert ds[:] == hub_ds[:]
hub_ds = hub_ds.cast_column("x", Image(decode=False))
elem = hub_ds[0]["x"]
path, bytes_ = elem["path"], elem["bytes"]
assert isinstance(path, str)
assert bool(bytes_) == embed_external_files
@require_pil
def test_push_dataset_to_hub_custom_features_image_list(self, temporary_repo):
image_path = os.path.join(os.path.dirname(__file__), "features", "data", "test_image_rgb.jpg")
data = {"x": [[image_path], [image_path, image_path]], "y": [0, -1]}
features = Features({"x": [Image()], "y": Value("int32")})
ds = Dataset.from_dict(data, features=features)
for embed_external_files in [True, False]:
with temporary_repo() as ds_name:
ds.push_to_hub(ds_name, embed_external_files=embed_external_files, token=self._token)
hub_ds = load_dataset(ds_name, split="train", download_mode="force_redownload")
assert ds.column_names == hub_ds.column_names
assert list(ds.features.keys()) == list(hub_ds.features.keys())
assert ds.features == hub_ds.features
assert ds[:] == hub_ds[:]
hub_ds = hub_ds.cast_column("x", [Image(decode=False)])
elem = hub_ds[0]["x"][0]
path, bytes_ = elem["path"], elem["bytes"]
assert isinstance(path, str)
assert bool(bytes_) == embed_external_files
def test_push_dataset_dict_to_hub_custom_features(self, temporary_repo):
features = Features({"x": Value("int64"), "y": ClassLabel(names=["neg", "pos"])})
ds = Dataset.from_dict({"x": [1, 2, 3], "y": [0, 0, 1]}, features=features)
local_ds = DatasetDict({"test": ds})
with temporary_repo() as ds_name:
local_ds.push_to_hub(ds_name, token=self._token)
hub_ds = load_dataset(ds_name, download_mode="force_redownload")
assert local_ds.column_names == hub_ds.column_names
assert list(local_ds["test"].features.keys()) == list(hub_ds["test"].features.keys())
assert local_ds["test"].features == hub_ds["test"].features
def test_push_dataset_to_hub_custom_splits(self, temporary_repo):
ds = Dataset.from_dict({"x": [1, 2, 3], "y": [4, 5, 6]})
with temporary_repo() as ds_name:
ds.push_to_hub(ds_name, split="random", token=self._token)
hub_ds = load_dataset(ds_name, download_mode="force_redownload")
assert ds.column_names == hub_ds["random"].column_names
assert list(ds.features.keys()) == list(hub_ds["random"].features.keys())
assert ds.features == hub_ds["random"].features
def test_push_dataset_to_hub_multiple_splits_one_by_one(self, temporary_repo):
ds = Dataset.from_dict({"x": [1, 2, 3], "y": [4, 5, 6]})
with temporary_repo() as ds_name:
ds.push_to_hub(ds_name, split="train", token=self._token)
ds.push_to_hub(ds_name, split="test", token=self._token)
hub_ds = load_dataset(ds_name, download_mode="force_redownload")
assert sorted(hub_ds) == ["test", "train"]
assert ds.column_names == hub_ds["train"].column_names
assert list(ds.features.keys()) == list(hub_ds["train"].features.keys())
assert ds.features == hub_ds["train"].features
def test_push_dataset_dict_to_hub_custom_splits(self, temporary_repo):
ds = Dataset.from_dict({"x": [1, 2, 3], "y": [4, 5, 6]})
local_ds = DatasetDict({"random": ds})
with temporary_repo() as ds_name:
local_ds.push_to_hub(ds_name, token=self._token)
hub_ds = load_dataset(ds_name, download_mode="force_redownload")
assert local_ds.column_names == hub_ds.column_names
assert list(local_ds["random"].features.keys()) == list(hub_ds["random"].features.keys())
assert local_ds["random"].features == hub_ds["random"].features
@unittest.skip("This test cannot pass until iterable datasets have push to hub")
def test_push_streaming_dataset_dict_to_hub(self, temporary_repo):
ds = Dataset.from_dict({"x": [1, 2, 3], "y": [4, 5, 6]})
local_ds = DatasetDict({"train": ds})
with tempfile.TemporaryDirectory() as tmp:
local_ds.save_to_disk(tmp)
local_ds = load_dataset(tmp, streaming=True)
with temporary_repo() as ds_name:
local_ds.push_to_hub(ds_name, token=self._token)
hub_ds = load_dataset(ds_name, download_mode="force_redownload")
assert local_ds.column_names == hub_ds.column_names
assert list(local_ds["train"].features.keys()) == list(hub_ds["train"].features.keys())
assert local_ds["train"].features == hub_ds["train"].features
def test_push_multiple_dataset_configs_to_hub_load_dataset_builder(self, temporary_repo):
ds_default = Dataset.from_dict({"a": [0], "b": [1]})
ds_config1 = Dataset.from_dict({"x": [1, 2, 3], "y": [4, 5, 6]})
ds_config2 = Dataset.from_dict({"foo": [1, 2], "bar": [4, 5]})
with temporary_repo() as ds_name:
ds_default.push_to_hub(ds_name, token=self._token)
ds_config1.push_to_hub(ds_name, "config1", token=self._token)
ds_config2.push_to_hub(ds_name, "config2", token=self._token)
ds_builder_default = load_dataset_builder(ds_name, download_mode="force_redownload") # default config
assert len(ds_builder_default.BUILDER_CONFIGS) == 3
assert len(ds_builder_default.config.data_files["train"]) == 1
assert fnmatch.fnmatch(
ds_builder_default.config.data_files["train"][0],
"*/data/train-*",
)
ds_builder_config1 = load_dataset_builder(ds_name, "config1", download_mode="force_redownload")
assert len(ds_builder_config1.BUILDER_CONFIGS) == 3
assert len(ds_builder_config1.config.data_files["train"]) == 1
assert fnmatch.fnmatch(
ds_builder_config1.config.data_files["train"][0],
"*/config1/train-*",
)
ds_builder_config2 = load_dataset_builder(ds_name, "config2", download_mode="force_redownload")
assert len(ds_builder_config2.BUILDER_CONFIGS) == 3
assert len(ds_builder_config2.config.data_files["train"]) == 1
assert fnmatch.fnmatch(
ds_builder_config2.config.data_files["train"][0],
"*/config2/train-*",
)
with pytest.raises(ValueError): # no config 'config3'
load_dataset_builder(ds_name, "config3", download_mode="force_redownload")
def test_push_multiple_dataset_configs_to_hub_load_dataset(self, temporary_repo):
ds_default = Dataset.from_dict({"a": [0], "b": [1]})
ds_config1 = Dataset.from_dict({"x": [1, 2, 3], "y": [4, 5, 6]})
ds_config2 = Dataset.from_dict({"foo": [1, 2], "bar": [4, 5]})
with temporary_repo() as ds_name:
ds_default.push_to_hub(ds_name, token=self._token)
ds_config1.push_to_hub(ds_name, "config1", token=self._token)
ds_config2.push_to_hub(ds_name, "config2", token=self._token)
files = sorted(self._api.list_repo_files(ds_name, repo_type="dataset"))
assert files == [
".gitattributes",
"README.md",
"config1/train-00000-of-00001.parquet",
"config2/train-00000-of-00001.parquet",
"data/train-00000-of-00001.parquet",
]
hub_ds_default = load_dataset(ds_name, download_mode="force_redownload")
hub_ds_config1 = load_dataset(ds_name, "config1", download_mode="force_redownload")
hub_ds_config2 = load_dataset(ds_name, "config2", download_mode="force_redownload")
# only "train" split
assert len(hub_ds_default) == len(hub_ds_config1) == len(hub_ds_config2) == 1
assert ds_default.column_names == hub_ds_default["train"].column_names == ["a", "b"]
assert ds_config1.column_names == hub_ds_config1["train"].column_names == ["x", "y"]
assert ds_config2.column_names == hub_ds_config2["train"].column_names == ["foo", "bar"]
assert ds_default.features == hub_ds_default["train"].features
assert ds_config1.features == hub_ds_config1["train"].features
assert ds_config2.features == hub_ds_config2["train"].features
assert ds_default.num_rows == hub_ds_default["train"].num_rows == 1
assert ds_config1.num_rows == hub_ds_config1["train"].num_rows == 3
assert ds_config2.num_rows == hub_ds_config2["train"].num_rows == 2
with pytest.raises(ValueError): # no config 'config3'
load_dataset(ds_name, "config3", download_mode="force_redownload")
def test_push_multiple_dataset_configs_to_hub_readme_metadata_content(self, temporary_repo):
ds_default = Dataset.from_dict({"a": [0], "b": [2]})
ds_config1 = Dataset.from_dict({"x": [1, 2, 3], "y": [4, 5, 6]})
ds_config2 = Dataset.from_dict({"foo": [1, 2], "bar": [4, 5]})
with temporary_repo() as ds_name:
ds_default.push_to_hub(ds_name, token=self._token)
ds_config1.push_to_hub(ds_name, "config1", token=self._token)
ds_config2.push_to_hub(ds_name, "config2", token=self._token)
# check that configs args was correctly pushed to README.md
ds_readme_path = cached_path(hf_hub_url(ds_name, "README.md"))
dataset_card_data = DatasetCard.load(ds_readme_path).data
assert METADATA_CONFIGS_FIELD in dataset_card_data
assert isinstance(dataset_card_data[METADATA_CONFIGS_FIELD], list)
assert sorted(dataset_card_data[METADATA_CONFIGS_FIELD], key=lambda x: x["config_name"]) == [
{
"config_name": "config1",
"data_files": [
{"split": "train", "path": "config1/train-*"},
],
},
{
"config_name": "config2",
"data_files": [
{"split": "train", "path": "config2/train-*"},
],
},
{
"config_name": "default",
"data_files": [
{"split": "train", "path": "data/train-*"},
],
},
]
def test_push_multiple_dataset_dict_configs_to_hub_load_dataset_builder(self, temporary_repo):
ds_default = Dataset.from_dict({"a": [0], "b": [1]})
ds_config1 = Dataset.from_dict({"x": [1, 2, 3], "y": [4, 5, 6]})
ds_config2 = Dataset.from_dict({"foo": [1, 2], "bar": [4, 5]})
ds_default = DatasetDict({"random": ds_default})
ds_config1 = DatasetDict({"random": ds_config1})
ds_config2 = DatasetDict({"random": ds_config2})
with temporary_repo() as ds_name:
ds_default.push_to_hub(ds_name, token=self._token)
ds_config1.push_to_hub(ds_name, "config1", token=self._token)
ds_config2.push_to_hub(ds_name, "config2", token=self._token)
ds_builder_default = load_dataset_builder(ds_name, download_mode="force_redownload") # default config
assert len(ds_builder_default.BUILDER_CONFIGS) == 3
assert len(ds_builder_default.config.data_files["random"]) == 1
assert fnmatch.fnmatch(
ds_builder_default.config.data_files["random"][0],
"*/data/random-*",
)
ds_builder_config1 = load_dataset_builder(ds_name, "config1", download_mode="force_redownload")
assert len(ds_builder_config1.BUILDER_CONFIGS) == 3
assert len(ds_builder_config1.config.data_files["random"]) == 1
assert fnmatch.fnmatch(
ds_builder_config1.config.data_files["random"][0],
"*/config1/random-*",
)
ds_builder_config2 = load_dataset_builder(ds_name, "config2", download_mode="force_redownload")
assert len(ds_builder_config2.BUILDER_CONFIGS) == 3
assert len(ds_builder_config2.config.data_files["random"]) == 1
assert fnmatch.fnmatch(
ds_builder_config2.config.data_files["random"][0],
"*/config2/random-*",
)
with pytest.raises(ValueError): # no config named 'config3'
load_dataset_builder(ds_name, "config3", download_mode="force_redownload")
def test_push_multiple_dataset_dict_configs_to_hub_load_dataset(self, temporary_repo):
ds_default = Dataset.from_dict({"a": [0], "b": [1]})
ds_config1 = Dataset.from_dict({"x": [1, 2, 3], "y": [4, 5, 6]})
ds_config2 = Dataset.from_dict({"foo": [1, 2], "bar": [4, 5]})
ds_default = DatasetDict({"train": ds_default, "random": ds_default})
ds_config1 = DatasetDict({"train": ds_config1, "random": ds_config1})
ds_config2 = DatasetDict({"train": ds_config2, "random": ds_config2})
with temporary_repo() as ds_name:
ds_default.push_to_hub(ds_name, token=self._token)
ds_config1.push_to_hub(ds_name, "config1", token=self._token)
ds_config2.push_to_hub(ds_name, "config2", token=self._token)
files = sorted(self._api.list_repo_files(ds_name, repo_type="dataset"))
assert files == [
".gitattributes",
"README.md",
"config1/random-00000-of-00001.parquet",
"config1/train-00000-of-00001.parquet",
"config2/random-00000-of-00001.parquet",
"config2/train-00000-of-00001.parquet",
"data/random-00000-of-00001.parquet",
"data/train-00000-of-00001.parquet",
]
hub_ds_default = load_dataset(ds_name, download_mode="force_redownload")
hub_ds_config1 = load_dataset(ds_name, "config1", download_mode="force_redownload")
hub_ds_config2 = load_dataset(ds_name, "config2", download_mode="force_redownload")
# two splits
expected_splits = ["random", "train"]
assert len(hub_ds_default) == len(hub_ds_config1) == len(hub_ds_config2) == 2
assert sorted(hub_ds_default) == sorted(hub_ds_config1) == sorted(hub_ds_config2) == expected_splits
for split in expected_splits:
assert ds_default[split].column_names == hub_ds_default[split].column_names == ["a", "b"]
assert ds_config1[split].column_names == hub_ds_config1[split].column_names == ["x", "y"]
assert ds_config2[split].column_names == hub_ds_config2[split].column_names == ["foo", "bar"]
assert ds_default[split].features == hub_ds_default[split].features
assert ds_config1[split].features == hub_ds_config1[split].features
assert ds_config2[split].features == hub_ds_config2["train"].features
assert ds_default[split].num_rows == hub_ds_default[split].num_rows == 1
assert ds_config1[split].num_rows == hub_ds_config1[split].num_rows == 3
assert ds_config2[split].num_rows == hub_ds_config2[split].num_rows == 2
with pytest.raises(ValueError): # no config 'config3'
load_dataset(ds_name, "config3", download_mode="force_redownload")
def test_push_multiple_dataset_dict_configs_to_hub_readme_metadata_content(self, temporary_repo):
ds_default = Dataset.from_dict({"a": [0], "b": [1]})
ds_config1 = Dataset.from_dict({"x": [1, 2, 3], "y": [4, 5, 6]})
ds_config2 = Dataset.from_dict({"foo": [1, 2], "bar": [4, 5]})
ds_default = DatasetDict({"train": ds_default, "random": ds_default})
ds_config1 = DatasetDict({"train": ds_config1, "random": ds_config1})
ds_config2 = DatasetDict({"train": ds_config2, "random": ds_config2})
with temporary_repo() as ds_name:
ds_default.push_to_hub(ds_name, token=self._token)
ds_config1.push_to_hub(ds_name, "config1", token=self._token)
ds_config2.push_to_hub(ds_name, "config2", token=self._token)
# check that configs args was correctly pushed to README.md
ds_readme_path = cached_path(hf_hub_url(ds_name, "README.md"))
dataset_card_data = DatasetCard.load(ds_readme_path).data
assert METADATA_CONFIGS_FIELD in dataset_card_data
assert isinstance(dataset_card_data[METADATA_CONFIGS_FIELD], list)
assert sorted(dataset_card_data[METADATA_CONFIGS_FIELD], key=lambda x: x["config_name"]) == [
{
"config_name": "config1",
"data_files": [
{"split": "train", "path": "config1/train-*"},
{"split": "random", "path": "config1/random-*"},
],
},
{
"config_name": "config2",
"data_files": [
{"split": "train", "path": "config2/train-*"},
{"split": "random", "path": "config2/random-*"},
],
},
{
"config_name": "default",
"data_files": [
{"split": "train", "path": "data/train-*"},
{"split": "random", "path": "data/random-*"},
],
},
]
def test_push_dataset_to_hub_with_config_no_metadata_configs(self, temporary_repo):
ds = Dataset.from_dict({"x": [1, 2, 3], "y": [4, 5, 6]})
ds_another_config = Dataset.from_dict({"foo": [1, 2], "bar": [4, 5]})
parquet_buf = BytesIO()
ds.to_parquet(parquet_buf)
parquet_content = parquet_buf.getvalue()
with temporary_repo() as ds_name:
self._api.create_repo(ds_name, token=self._token, repo_type="dataset")
# old push_to_hub was uploading the parquet files only - without metadata configs
self._api.upload_file(
path_or_fileobj=parquet_content,
path_in_repo="data/train-00000-of-00001.parquet",
repo_id=ds_name,
repo_type="dataset",
token=self._token,
)
ds_another_config.push_to_hub(ds_name, "another_config", token=self._token)
ds_builder = load_dataset_builder(ds_name, download_mode="force_redownload")
assert len(ds_builder.config.data_files) == 1
assert len(ds_builder.config.data_files["train"]) == 1
assert fnmatch.fnmatch(ds_builder.config.data_files["train"][0], "*/data/train-00000-of-00001.parquet")
ds_another_config_builder = load_dataset_builder(
ds_name, "another_config", download_mode="force_redownload"
)
assert len(ds_another_config_builder.config.data_files) == 1
assert len(ds_another_config_builder.config.data_files["train"]) == 1
assert fnmatch.fnmatch(
ds_another_config_builder.config.data_files["train"][0],
"*/another_config/train-00000-of-00001.parquet",
)
def test_push_dataset_dict_to_hub_with_config_no_metadata_configs(self, temporary_repo):
ds = Dataset.from_dict({"x": [1, 2, 3], "y": [4, 5, 6]})
ds_another_config = Dataset.from_dict({"foo": [1, 2], "bar": [4, 5]})
parquet_buf = BytesIO()
ds.to_parquet(parquet_buf)
parquet_content = parquet_buf.getvalue()
local_ds_another_config = DatasetDict({"random": ds_another_config})
with temporary_repo() as ds_name:
self._api.create_repo(ds_name, token=self._token, repo_type="dataset")
# old push_to_hub was uploading the parquet files only - without metadata configs
self._api.upload_file(
path_or_fileobj=parquet_content,
path_in_repo="data/random-00000-of-00001.parquet",
repo_id=ds_name,
repo_type="dataset",
token=self._token,
)
local_ds_another_config.push_to_hub(ds_name, "another_config", token=self._token)
ds_builder = load_dataset_builder(ds_name, download_mode="force_redownload")
assert len(ds_builder.config.data_files) == 1
assert len(ds_builder.config.data_files["random"]) == 1
assert fnmatch.fnmatch(ds_builder.config.data_files["random"][0], "*/data/random-00000-of-00001.parquet")
ds_another_config_builder = load_dataset_builder(
ds_name, "another_config", download_mode="force_redownload"
)
assert len(ds_another_config_builder.config.data_files) == 1
assert len(ds_another_config_builder.config.data_files["random"]) == 1
assert fnmatch.fnmatch(
ds_another_config_builder.config.data_files["random"][0],
"*/another_config/random-00000-of-00001.parquet",
)
class DummyFolderBasedBuilder(FolderBasedBuilder):
BASE_FEATURE = dict
BASE_COLUMN_NAME = "base"
BUILDER_CONFIG_CLASS = FolderBasedBuilderConfig
EXTENSIONS = [".txt"]
# CLASSIFICATION_TASK = TextClassification(text_column="base", label_column="label")
@pytest.fixture(params=[".jsonl", ".csv"])
def text_file_with_metadata(request, tmp_path, text_file):
metadata_filename_extension = request.param
data_dir = tmp_path / "data_dir"
data_dir.mkdir()
text_file_path = data_dir / "file.txt"
shutil.copyfile(text_file, text_file_path)
metadata_file_path = data_dir / f"metadata{metadata_filename_extension}"
metadata = textwrap.dedent(
"""\
{"file_name": "file.txt", "additional_feature": "Dummy file"}
"""
if metadata_filename_extension == ".jsonl"
else """\
file_name,additional_feature
file.txt,Dummy file
"""
)
with open(metadata_file_path, "w", encoding="utf-8") as f:
f.write(metadata)
return text_file_path, metadata_file_path
@for_all_test_methods(xfail_if_500_502_http_error)
@pytest.mark.usefixtures("ci_hub_config", "ci_hfh_hf_hub_url")
class TestLoadFromHub:
_api = HfApi(endpoint=CI_HUB_ENDPOINT)
_token = CI_HUB_USER_TOKEN
def test_load_dataset_with_metadata_file(self, temporary_repo, text_file_with_metadata, tmp_path):
text_file_path, metadata_file_path = text_file_with_metadata
data_dir_path = text_file_path.parent
cache_dir_path = tmp_path / ".cache"
cache_dir_path.mkdir()
with temporary_repo() as repo_id:
self._api.create_repo(repo_id, token=self._token, repo_type="dataset")
self._api.upload_folder(
folder_path=str(data_dir_path),
repo_id=repo_id,
repo_type="dataset",
token=self._token,
)
data_files = [
f"hf://datasets/{repo_id}/{text_file_path.name}",
f"hf://datasets/{repo_id}/{metadata_file_path.name}",
]
builder = DummyFolderBasedBuilder(
dataset_name=repo_id.split("/")[-1], data_files=data_files, cache_dir=str(cache_dir_path)
)
download_manager = DownloadManager()
gen_kwargs = builder._split_generators(download_manager)[0].gen_kwargs
generator = builder._generate_examples(**gen_kwargs)
result = [example for _, example in generator]
assert len(result) == 1
def test_get_data_patterns(self, temporary_repo, tmp_path):
repo_dir = tmp_path / "test_get_data_patterns"
data_dir = repo_dir / "data"
data_dir.mkdir(parents=True)
data_file = data_dir / "train-00001-of-00009.parquet"
data_file.touch()
with temporary_repo() as repo_id:
self._api.create_repo(repo_id, token=self._token, repo_type="dataset")
self._api.upload_folder(
folder_path=str(repo_dir),
repo_id=repo_id,
repo_type="dataset",
token=self._token,
)
data_file_patterns = get_data_patterns(f"hf://datasets/{repo_id}")
assert data_file_patterns == {
"train": ["data/train-[0-9][0-9][0-9][0-9][0-9]-of-[0-9][0-9][0-9][0-9][0-9]*.*"]
}
| 0
|
hf_public_repos/datasets
|
hf_public_repos/datasets/tests/test_arrow_dataset.py
|
import contextlib
import copy
import itertools
import json
import os
import pickle
import re
import sys
import tempfile
from functools import partial
from pathlib import Path
from unittest import TestCase
from unittest.mock import MagicMock, patch
import numpy as np
import numpy.testing as npt
import pandas as pd
import pyarrow as pa
import pytest
from absl.testing import parameterized
from fsspec.core import strip_protocol
from packaging import version
import datasets.arrow_dataset
from datasets import concatenate_datasets, interleave_datasets, load_from_disk
from datasets.arrow_dataset import Dataset, transmit_format, update_metadata_with_features
from datasets.dataset_dict import DatasetDict
from datasets.features import (
Array2D,
Array3D,
Audio,
ClassLabel,
Features,
Image,
Sequence,
Translation,
TranslationVariableLanguages,
Value,
)
from datasets.info import DatasetInfo
from datasets.iterable_dataset import IterableDataset
from datasets.splits import NamedSplit
from datasets.table import ConcatenationTable, InMemoryTable, MemoryMappedTable
from datasets.tasks import (
AutomaticSpeechRecognition,
LanguageModeling,
QuestionAnsweringExtractive,
Summarization,
TextClassification,
)
from datasets.utils.logging import INFO, get_logger
from datasets.utils.py_utils import temp_seed
from .utils import (
assert_arrow_memory_doesnt_increase,
assert_arrow_memory_increases,
require_dill_gt_0_3_2,
require_jax,
require_not_windows,
require_pil,
require_pyspark,
require_sqlalchemy,
require_tf,
require_torch,
require_transformers,
set_current_working_directory_to_temp_dir,
)
class PickableMagicMock(MagicMock):
def __reduce__(self):
return MagicMock, ()
class Unpicklable:
def __getstate__(self):
raise pickle.PicklingError()
def picklable_map_function(x):
return {"id": int(x["filename"].split("_")[-1])}
def picklable_map_function_with_indices(x, i):
return {"id": i}
def picklable_map_function_with_rank(x, r):
return {"rank": r}
def picklable_map_function_with_indices_and_rank(x, i, r):
return {"id": i, "rank": r}
def picklable_filter_function(x):
return int(x["filename"].split("_")[-1]) < 10
def assert_arrow_metadata_are_synced_with_dataset_features(dataset: Dataset):
assert dataset.data.schema.metadata is not None
assert b"huggingface" in dataset.data.schema.metadata
metadata = json.loads(dataset.data.schema.metadata[b"huggingface"].decode())
assert "info" in metadata
features = DatasetInfo.from_dict(metadata["info"]).features
assert features is not None
assert features == dataset.features
assert features == Features.from_arrow_schema(dataset.data.schema)
assert list(features) == dataset.data.column_names
assert list(features) == list(dataset.features)
IN_MEMORY_PARAMETERS = [
{"testcase_name": name, "in_memory": im} for im, name in [(True, "in_memory"), (False, "on_disk")]
]
@parameterized.named_parameters(IN_MEMORY_PARAMETERS)
class BaseDatasetTest(TestCase):
@pytest.fixture(autouse=True)
def inject_fixtures(self, caplog, set_sqlalchemy_silence_uber_warning):
self._caplog = caplog
def _create_dummy_dataset(
self, in_memory: bool, tmp_dir: str, multiple_columns=False, array_features=False, nested_features=False
) -> Dataset:
assert int(multiple_columns) + int(array_features) + int(nested_features) < 2
if multiple_columns:
data = {"col_1": [3, 2, 1, 0], "col_2": ["a", "b", "c", "d"], "col_3": [False, True, False, True]}
dset = Dataset.from_dict(data)
elif array_features:
data = {
"col_1": [[[True, False], [False, True]]] * 4, # 2D
"col_2": [[[["a", "b"], ["c", "d"]], [["e", "f"], ["g", "h"]]]] * 4, # 3D array
"col_3": [[3, 2, 1, 0]] * 4, # Sequence
}
features = Features(
{
"col_1": Array2D(shape=(2, 2), dtype="bool"),
"col_2": Array3D(shape=(2, 2, 2), dtype="string"),
"col_3": Sequence(feature=Value("int64")),
}
)
dset = Dataset.from_dict(data, features=features)
elif nested_features:
data = {"nested": [{"a": i, "x": i * 10, "c": i * 100} for i in range(1, 11)]}
features = Features({"nested": {"a": Value("int64"), "x": Value("int64"), "c": Value("int64")}})
dset = Dataset.from_dict(data, features=features)
else:
dset = Dataset.from_dict({"filename": ["my_name-train" + "_" + str(x) for x in np.arange(30).tolist()]})
if not in_memory:
dset = self._to(in_memory, tmp_dir, dset)
return dset
def _to(self, in_memory, tmp_dir, *datasets):
if in_memory:
datasets = [dataset.map(keep_in_memory=True) for dataset in datasets]
else:
start = 0
while os.path.isfile(os.path.join(tmp_dir, f"dataset{start}.arrow")):
start += 1
datasets = [
dataset.map(cache_file_name=os.path.join(tmp_dir, f"dataset{start + i}.arrow"))
for i, dataset in enumerate(datasets)
]
return datasets if len(datasets) > 1 else datasets[0]
def test_dummy_dataset(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertEqual(dset[0]["filename"], "my_name-train_0")
self.assertEqual(dset["filename"][0], "my_name-train_0")
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
self.assertDictEqual(
dset.features,
Features({"col_1": Value("int64"), "col_2": Value("string"), "col_3": Value("bool")}),
)
self.assertEqual(dset[0]["col_1"], 3)
self.assertEqual(dset["col_1"][0], 3)
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir, array_features=True) as dset:
self.assertDictEqual(
dset.features,
Features(
{
"col_1": Array2D(shape=(2, 2), dtype="bool"),
"col_2": Array3D(shape=(2, 2, 2), dtype="string"),
"col_3": Sequence(feature=Value("int64")),
}
),
)
self.assertEqual(dset[0]["col_2"], [[["a", "b"], ["c", "d"]], [["e", "f"], ["g", "h"]]])
self.assertEqual(dset["col_2"][0], [[["a", "b"], ["c", "d"]], [["e", "f"], ["g", "h"]]])
def test_dataset_getitem(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
self.assertEqual(dset[0]["filename"], "my_name-train_0")
self.assertEqual(dset["filename"][0], "my_name-train_0")
self.assertEqual(dset[-1]["filename"], "my_name-train_29")
self.assertEqual(dset["filename"][-1], "my_name-train_29")
self.assertListEqual(dset[:2]["filename"], ["my_name-train_0", "my_name-train_1"])
self.assertListEqual(dset["filename"][:2], ["my_name-train_0", "my_name-train_1"])
self.assertEqual(dset[:-1]["filename"][-1], "my_name-train_28")
self.assertEqual(dset["filename"][:-1][-1], "my_name-train_28")
self.assertListEqual(dset[[0, -1]]["filename"], ["my_name-train_0", "my_name-train_29"])
self.assertListEqual(dset[range(0, -2, -1)]["filename"], ["my_name-train_0", "my_name-train_29"])
self.assertListEqual(dset[np.array([0, -1])]["filename"], ["my_name-train_0", "my_name-train_29"])
self.assertListEqual(dset[pd.Series([0, -1])]["filename"], ["my_name-train_0", "my_name-train_29"])
with dset.select(range(2)) as dset_subset:
self.assertListEqual(dset_subset[-1:]["filename"], ["my_name-train_1"])
self.assertListEqual(dset_subset["filename"][-1:], ["my_name-train_1"])
def test_dummy_dataset_deepcopy(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir).select(range(10)) as dset:
with assert_arrow_memory_doesnt_increase():
dset2 = copy.deepcopy(dset)
# don't copy the underlying arrow data using memory
self.assertEqual(len(dset2), 10)
self.assertDictEqual(dset2.features, Features({"filename": Value("string")}))
self.assertEqual(dset2[0]["filename"], "my_name-train_0")
self.assertEqual(dset2["filename"][0], "my_name-train_0")
del dset2
def test_dummy_dataset_pickle(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
tmp_file = os.path.join(tmp_dir, "dset.pt")
with self._create_dummy_dataset(in_memory, tmp_dir).select(range(0, 10, 2)) as dset:
with open(tmp_file, "wb") as f:
pickle.dump(dset, f)
with open(tmp_file, "rb") as f:
with pickle.load(f) as dset:
self.assertEqual(len(dset), 5)
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertEqual(dset[0]["filename"], "my_name-train_0")
self.assertEqual(dset["filename"][0], "my_name-train_0")
with self._create_dummy_dataset(in_memory, tmp_dir).select(
range(0, 10, 2), indices_cache_file_name=os.path.join(tmp_dir, "ind.arrow")
) as dset:
if not in_memory:
dset._data.table = Unpicklable()
dset._indices.table = Unpicklable()
with open(tmp_file, "wb") as f:
pickle.dump(dset, f)
with open(tmp_file, "rb") as f:
with pickle.load(f) as dset:
self.assertEqual(len(dset), 5)
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertEqual(dset[0]["filename"], "my_name-train_0")
self.assertEqual(dset["filename"][0], "my_name-train_0")
def test_dummy_dataset_serialize(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
with set_current_working_directory_to_temp_dir():
with self._create_dummy_dataset(in_memory, tmp_dir).select(range(10)) as dset:
dataset_path = "my_dataset" # rel path
dset.save_to_disk(dataset_path)
with Dataset.load_from_disk(dataset_path) as dset:
self.assertEqual(len(dset), 10)
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertEqual(dset[0]["filename"], "my_name-train_0")
self.assertEqual(dset["filename"][0], "my_name-train_0")
expected = dset.to_dict()
with self._create_dummy_dataset(in_memory, tmp_dir).select(range(10)) as dset:
dataset_path = os.path.join(tmp_dir, "my_dataset") # abs path
dset.save_to_disk(dataset_path)
with Dataset.load_from_disk(dataset_path) as dset:
self.assertEqual(len(dset), 10)
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertEqual(dset[0]["filename"], "my_name-train_0")
self.assertEqual(dset["filename"][0], "my_name-train_0")
with self._create_dummy_dataset(in_memory, tmp_dir).select(
range(10), indices_cache_file_name=os.path.join(tmp_dir, "ind.arrow")
) as dset:
with assert_arrow_memory_doesnt_increase():
dset.save_to_disk(dataset_path)
with Dataset.load_from_disk(dataset_path) as dset:
self.assertEqual(len(dset), 10)
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertEqual(dset[0]["filename"], "my_name-train_0")
self.assertEqual(dset["filename"][0], "my_name-train_0")
with self._create_dummy_dataset(in_memory, tmp_dir, nested_features=True) as dset:
with assert_arrow_memory_doesnt_increase():
dset.save_to_disk(dataset_path)
with Dataset.load_from_disk(dataset_path) as dset:
self.assertEqual(len(dset), 10)
self.assertDictEqual(
dset.features,
Features({"nested": {"a": Value("int64"), "x": Value("int64"), "c": Value("int64")}}),
)
self.assertDictEqual(dset[0]["nested"], {"a": 1, "c": 100, "x": 10})
self.assertDictEqual(dset["nested"][0], {"a": 1, "c": 100, "x": 10})
with self._create_dummy_dataset(in_memory, tmp_dir).select(range(10)) as dset:
with assert_arrow_memory_doesnt_increase():
dset.save_to_disk(dataset_path, num_shards=4)
with Dataset.load_from_disk(dataset_path) as dset:
self.assertEqual(len(dset), 10)
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertDictEqual(dset.to_dict(), expected)
self.assertEqual(len(dset.cache_files), 4)
with self._create_dummy_dataset(in_memory, tmp_dir).select(range(10)) as dset:
with assert_arrow_memory_doesnt_increase():
dset.save_to_disk(dataset_path, num_proc=2)
with Dataset.load_from_disk(dataset_path) as dset:
self.assertEqual(len(dset), 10)
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertDictEqual(dset.to_dict(), expected)
self.assertEqual(len(dset.cache_files), 2)
with self._create_dummy_dataset(in_memory, tmp_dir).select(range(10)) as dset:
with assert_arrow_memory_doesnt_increase():
dset.save_to_disk(dataset_path, num_shards=7, num_proc=2)
with Dataset.load_from_disk(dataset_path) as dset:
self.assertEqual(len(dset), 10)
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertDictEqual(dset.to_dict(), expected)
self.assertEqual(len(dset.cache_files), 7)
with self._create_dummy_dataset(in_memory, tmp_dir).select(range(10)) as dset:
with assert_arrow_memory_doesnt_increase():
max_shard_size = dset._estimate_nbytes() // 2 + 1
dset.save_to_disk(dataset_path, max_shard_size=max_shard_size)
with Dataset.load_from_disk(dataset_path) as dset:
self.assertEqual(len(dset), 10)
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertDictEqual(dset.to_dict(), expected)
self.assertEqual(len(dset.cache_files), 2)
def test_dummy_dataset_load_from_disk(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir).select(range(10)) as dset:
dataset_path = os.path.join(tmp_dir, "my_dataset")
dset.save_to_disk(dataset_path)
with load_from_disk(dataset_path) as dset:
self.assertEqual(len(dset), 10)
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertEqual(dset[0]["filename"], "my_name-train_0")
self.assertEqual(dset["filename"][0], "my_name-train_0")
def test_restore_saved_format(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
dset.set_format(type="numpy", columns=["col_1"], output_all_columns=True)
dataset_path = os.path.join(tmp_dir, "my_dataset")
dset.save_to_disk(dataset_path)
with load_from_disk(dataset_path) as loaded_dset:
self.assertEqual(dset.format, loaded_dset.format)
def test_set_format_numpy_multiple_columns(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
fingerprint = dset._fingerprint
dset.set_format(type="numpy", columns=["col_1"])
self.assertEqual(len(dset[0]), 1)
self.assertIsInstance(dset[0]["col_1"], np.int64)
self.assertEqual(dset[0]["col_1"].item(), 3)
self.assertIsInstance(dset["col_1"], np.ndarray)
self.assertListEqual(list(dset["col_1"].shape), [4])
np.testing.assert_array_equal(dset["col_1"], np.array([3, 2, 1, 0]))
self.assertNotEqual(dset._fingerprint, fingerprint)
dset.reset_format()
with dset.formatted_as(type="numpy", columns=["col_1"]):
self.assertEqual(len(dset[0]), 1)
self.assertIsInstance(dset[0]["col_1"], np.int64)
self.assertEqual(dset[0]["col_1"].item(), 3)
self.assertIsInstance(dset["col_1"], np.ndarray)
self.assertListEqual(list(dset["col_1"].shape), [4])
np.testing.assert_array_equal(dset["col_1"], np.array([3, 2, 1, 0]))
self.assertEqual(dset.format["type"], None)
self.assertEqual(dset.format["format_kwargs"], {})
self.assertEqual(dset.format["columns"], dset.column_names)
self.assertEqual(dset.format["output_all_columns"], False)
dset.set_format(type="numpy", columns=["col_1"], output_all_columns=True)
self.assertEqual(len(dset[0]), 3)
self.assertIsInstance(dset[0]["col_2"], str)
self.assertEqual(dset[0]["col_2"], "a")
dset.set_format(type="numpy", columns=["col_1", "col_2"])
self.assertEqual(len(dset[0]), 2)
self.assertIsInstance(dset[0]["col_2"], np.str_)
self.assertEqual(dset[0]["col_2"].item(), "a")
@require_torch
def test_set_format_torch(self, in_memory):
import torch
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
dset.set_format(type="torch", columns=["col_1"])
self.assertEqual(len(dset[0]), 1)
self.assertIsInstance(dset[0]["col_1"], torch.Tensor)
self.assertIsInstance(dset["col_1"], torch.Tensor)
self.assertListEqual(list(dset[0]["col_1"].shape), [])
self.assertEqual(dset[0]["col_1"].item(), 3)
dset.set_format(type="torch", columns=["col_1"], output_all_columns=True)
self.assertEqual(len(dset[0]), 3)
self.assertIsInstance(dset[0]["col_2"], str)
self.assertEqual(dset[0]["col_2"], "a")
dset.set_format(type="torch")
self.assertEqual(len(dset[0]), 3)
self.assertIsInstance(dset[0]["col_1"], torch.Tensor)
self.assertIsInstance(dset["col_1"], torch.Tensor)
self.assertListEqual(list(dset[0]["col_1"].shape), [])
self.assertEqual(dset[0]["col_1"].item(), 3)
self.assertIsInstance(dset[0]["col_2"], str)
self.assertEqual(dset[0]["col_2"], "a")
self.assertIsInstance(dset[0]["col_3"], torch.Tensor)
self.assertIsInstance(dset["col_3"], torch.Tensor)
self.assertListEqual(list(dset[0]["col_3"].shape), [])
@require_tf
def test_set_format_tf(self, in_memory):
import tensorflow as tf
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
dset.set_format(type="tensorflow", columns=["col_1"])
self.assertEqual(len(dset[0]), 1)
self.assertIsInstance(dset[0]["col_1"], tf.Tensor)
self.assertListEqual(list(dset[0]["col_1"].shape), [])
self.assertEqual(dset[0]["col_1"].numpy().item(), 3)
dset.set_format(type="tensorflow", columns=["col_1"], output_all_columns=True)
self.assertEqual(len(dset[0]), 3)
self.assertIsInstance(dset[0]["col_2"], str)
self.assertEqual(dset[0]["col_2"], "a")
dset.set_format(type="tensorflow", columns=["col_1", "col_2"])
self.assertEqual(len(dset[0]), 2)
self.assertEqual(dset[0]["col_2"].numpy().decode("utf-8"), "a")
def test_set_format_pandas(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
dset.set_format(type="pandas", columns=["col_1"])
self.assertEqual(len(dset[0].columns), 1)
self.assertIsInstance(dset[0], pd.DataFrame)
self.assertListEqual(list(dset[0].shape), [1, 1])
self.assertEqual(dset[0]["col_1"].item(), 3)
dset.set_format(type="pandas", columns=["col_1", "col_2"])
self.assertEqual(len(dset[0].columns), 2)
self.assertEqual(dset[0]["col_2"].item(), "a")
def test_set_transform(self, in_memory):
def transform(batch):
return {k: [str(i).upper() for i in v] for k, v in batch.items()}
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
dset.set_transform(transform=transform, columns=["col_1"])
self.assertEqual(dset.format["type"], "custom")
self.assertEqual(len(dset[0].keys()), 1)
self.assertEqual(dset[0]["col_1"], "3")
self.assertEqual(dset[:2]["col_1"], ["3", "2"])
self.assertEqual(dset["col_1"][:2], ["3", "2"])
prev_format = dset.format
dset.set_format(**dset.format)
self.assertEqual(prev_format, dset.format)
dset.set_transform(transform=transform, columns=["col_1", "col_2"])
self.assertEqual(len(dset[0].keys()), 2)
self.assertEqual(dset[0]["col_2"], "A")
def test_transmit_format(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
transform = datasets.arrow_dataset.transmit_format(lambda x: x)
# make sure identity transform doesn't apply unnecessary format
self.assertEqual(dset._fingerprint, transform(dset)._fingerprint)
dset.set_format(**dset.format)
self.assertEqual(dset._fingerprint, transform(dset)._fingerprint)
# check lists comparisons
dset.set_format(columns=["col_1"])
self.assertEqual(dset._fingerprint, transform(dset)._fingerprint)
dset.set_format(columns=["col_1", "col_2"])
self.assertEqual(dset._fingerprint, transform(dset)._fingerprint)
dset.set_format("numpy", columns=["col_1", "col_2"])
self.assertEqual(dset._fingerprint, transform(dset)._fingerprint)
def test_cast(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
features = dset.features
features["col_1"] = Value("float64")
features = Features({k: features[k] for k in list(features)[::-1]})
fingerprint = dset._fingerprint
# TODO: with assert_arrow_memory_increases() if in_memory else assert_arrow_memory_doesnt_increase():
with dset.cast(features) as casted_dset:
self.assertEqual(casted_dset.num_columns, 3)
self.assertEqual(casted_dset.features["col_1"], Value("float64"))
self.assertIsInstance(casted_dset[0]["col_1"], float)
self.assertNotEqual(casted_dset._fingerprint, fingerprint)
self.assertNotEqual(casted_dset, dset)
assert_arrow_metadata_are_synced_with_dataset_features(casted_dset)
def test_class_encode_column(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
with self.assertRaises(ValueError):
dset.class_encode_column(column="does not exist")
with dset.class_encode_column("col_1") as casted_dset:
self.assertIsInstance(casted_dset.features["col_1"], ClassLabel)
self.assertListEqual(casted_dset.features["col_1"].names, ["0", "1", "2", "3"])
self.assertListEqual(casted_dset["col_1"], [3, 2, 1, 0])
self.assertNotEqual(casted_dset._fingerprint, dset._fingerprint)
self.assertNotEqual(casted_dset, dset)
assert_arrow_metadata_are_synced_with_dataset_features(casted_dset)
with dset.class_encode_column("col_2") as casted_dset:
self.assertIsInstance(casted_dset.features["col_2"], ClassLabel)
self.assertListEqual(casted_dset.features["col_2"].names, ["a", "b", "c", "d"])
self.assertListEqual(casted_dset["col_2"], [0, 1, 2, 3])
self.assertNotEqual(casted_dset._fingerprint, dset._fingerprint)
self.assertNotEqual(casted_dset, dset)
assert_arrow_metadata_are_synced_with_dataset_features(casted_dset)
with dset.class_encode_column("col_3") as casted_dset:
self.assertIsInstance(casted_dset.features["col_3"], ClassLabel)
self.assertListEqual(casted_dset.features["col_3"].names, ["False", "True"])
self.assertListEqual(casted_dset["col_3"], [0, 1, 0, 1])
self.assertNotEqual(casted_dset._fingerprint, dset._fingerprint)
self.assertNotEqual(casted_dset, dset)
assert_arrow_metadata_are_synced_with_dataset_features(casted_dset)
# Test raises if feature is an array / sequence
with self._create_dummy_dataset(in_memory, tmp_dir, array_features=True) as dset:
for column in dset.column_names:
with self.assertRaises(ValueError):
dset.class_encode_column(column)
def test_remove_columns(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
fingerprint = dset._fingerprint
with dset.remove_columns(column_names="col_1") as new_dset:
self.assertEqual(new_dset.num_columns, 2)
self.assertListEqual(list(new_dset.column_names), ["col_2", "col_3"])
self.assertNotEqual(new_dset._fingerprint, fingerprint)
assert_arrow_metadata_are_synced_with_dataset_features(new_dset)
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
with dset.remove_columns(column_names=["col_1", "col_2", "col_3"]) as new_dset:
self.assertEqual(new_dset.num_columns, 0)
self.assertNotEqual(new_dset._fingerprint, fingerprint)
assert_arrow_metadata_are_synced_with_dataset_features(new_dset)
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
dset._format_columns = ["col_1", "col_2", "col_3"]
with dset.remove_columns(column_names=["col_1"]) as new_dset:
self.assertListEqual(new_dset._format_columns, ["col_2", "col_3"])
self.assertEqual(new_dset.num_columns, 2)
self.assertListEqual(list(new_dset.column_names), ["col_2", "col_3"])
self.assertNotEqual(new_dset._fingerprint, fingerprint)
assert_arrow_metadata_are_synced_with_dataset_features(new_dset)
def test_rename_column(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
fingerprint = dset._fingerprint
with dset.rename_column(original_column_name="col_1", new_column_name="new_name") as new_dset:
self.assertEqual(new_dset.num_columns, 3)
self.assertListEqual(list(new_dset.column_names), ["new_name", "col_2", "col_3"])
self.assertListEqual(list(dset.column_names), ["col_1", "col_2", "col_3"])
self.assertNotEqual(new_dset._fingerprint, fingerprint)
assert_arrow_metadata_are_synced_with_dataset_features(new_dset)
def test_rename_columns(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
fingerprint = dset._fingerprint
with dset.rename_columns({"col_1": "new_name"}) as new_dset:
self.assertEqual(new_dset.num_columns, 3)
self.assertListEqual(list(new_dset.column_names), ["new_name", "col_2", "col_3"])
self.assertListEqual(list(dset.column_names), ["col_1", "col_2", "col_3"])
self.assertNotEqual(new_dset._fingerprint, fingerprint)
with dset.rename_columns({"col_1": "new_name", "col_2": "new_name2"}) as new_dset:
self.assertEqual(new_dset.num_columns, 3)
self.assertListEqual(list(new_dset.column_names), ["new_name", "new_name2", "col_3"])
self.assertListEqual(list(dset.column_names), ["col_1", "col_2", "col_3"])
self.assertNotEqual(new_dset._fingerprint, fingerprint)
# Original column not in dataset
with self.assertRaises(ValueError):
dset.rename_columns({"not_there": "new_name"})
# Empty new name
with self.assertRaises(ValueError):
dset.rename_columns({"col_1": ""})
# Duplicates
with self.assertRaises(ValueError):
dset.rename_columns({"col_1": "new_name", "col_2": "new_name"})
def test_select_columns(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
fingerprint = dset._fingerprint
with dset.select_columns(column_names=[]) as new_dset:
self.assertEqual(new_dset.num_columns, 0)
self.assertListEqual(list(new_dset.column_names), [])
self.assertNotEqual(new_dset._fingerprint, fingerprint)
assert_arrow_metadata_are_synced_with_dataset_features(new_dset)
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
fingerprint = dset._fingerprint
with dset.select_columns(column_names="col_1") as new_dset:
self.assertEqual(new_dset.num_columns, 1)
self.assertListEqual(list(new_dset.column_names), ["col_1"])
self.assertNotEqual(new_dset._fingerprint, fingerprint)
assert_arrow_metadata_are_synced_with_dataset_features(new_dset)
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
with dset.select_columns(column_names=["col_1", "col_2", "col_3"]) as new_dset:
self.assertEqual(new_dset.num_columns, 3)
self.assertListEqual(list(new_dset.column_names), ["col_1", "col_2", "col_3"])
self.assertNotEqual(new_dset._fingerprint, fingerprint)
assert_arrow_metadata_are_synced_with_dataset_features(new_dset)
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
with dset.select_columns(column_names=["col_3", "col_2", "col_1"]) as new_dset:
self.assertEqual(new_dset.num_columns, 3)
self.assertListEqual(list(new_dset.column_names), ["col_3", "col_2", "col_1"])
self.assertNotEqual(new_dset._fingerprint, fingerprint)
assert_arrow_metadata_are_synced_with_dataset_features(new_dset)
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
dset._format_columns = ["col_1", "col_2", "col_3"]
with dset.select_columns(column_names=["col_1"]) as new_dset:
self.assertListEqual(new_dset._format_columns, ["col_1"])
self.assertEqual(new_dset.num_columns, 1)
self.assertListEqual(list(new_dset.column_names), ["col_1"])
self.assertNotEqual(new_dset._fingerprint, fingerprint)
assert_arrow_metadata_are_synced_with_dataset_features(new_dset)
def test_concatenate(self, in_memory):
data1, data2, data3 = {"id": [0, 1, 2]}, {"id": [3, 4, 5]}, {"id": [6, 7]}
info1 = DatasetInfo(description="Dataset1")
info2 = DatasetInfo(description="Dataset2")
with tempfile.TemporaryDirectory() as tmp_dir:
dset1, dset2, dset3 = (
Dataset.from_dict(data1, info=info1),
Dataset.from_dict(data2, info=info2),
Dataset.from_dict(data3),
)
dset1, dset2, dset3 = self._to(in_memory, tmp_dir, dset1, dset2, dset3)
with concatenate_datasets([dset1, dset2, dset3]) as dset_concat:
self.assertTupleEqual((len(dset1), len(dset2), len(dset3)), (3, 3, 2))
self.assertEqual(len(dset_concat), len(dset1) + len(dset2) + len(dset3))
self.assertListEqual(dset_concat["id"], [0, 1, 2, 3, 4, 5, 6, 7])
self.assertEqual(len(dset_concat.cache_files), 0 if in_memory else 3)
self.assertEqual(dset_concat.info.description, "Dataset1\n\nDataset2")
del dset1, dset2, dset3
def test_concatenate_formatted(self, in_memory):
data1, data2, data3 = {"id": [0, 1, 2]}, {"id": [3, 4, 5]}, {"id": [6, 7]}
info1 = DatasetInfo(description="Dataset1")
info2 = DatasetInfo(description="Dataset2")
with tempfile.TemporaryDirectory() as tmp_dir:
dset1, dset2, dset3 = (
Dataset.from_dict(data1, info=info1),
Dataset.from_dict(data2, info=info2),
Dataset.from_dict(data3),
)
dset1, dset2, dset3 = self._to(in_memory, tmp_dir, dset1, dset2, dset3)
dset1.set_format("numpy")
with concatenate_datasets([dset1, dset2, dset3]) as dset_concat:
self.assertEqual(dset_concat.format["type"], None)
dset2.set_format("numpy")
dset3.set_format("numpy")
with concatenate_datasets([dset1, dset2, dset3]) as dset_concat:
self.assertEqual(dset_concat.format["type"], "numpy")
del dset1, dset2, dset3
def test_concatenate_with_indices(self, in_memory):
data1, data2, data3 = {"id": [0, 1, 2] * 2}, {"id": [3, 4, 5] * 2}, {"id": [6, 7, 8]}
info1 = DatasetInfo(description="Dataset1")
info2 = DatasetInfo(description="Dataset2")
with tempfile.TemporaryDirectory() as tmp_dir:
dset1, dset2, dset3 = (
Dataset.from_dict(data1, info=info1),
Dataset.from_dict(data2, info=info2),
Dataset.from_dict(data3),
)
dset1, dset2, dset3 = self._to(in_memory, tmp_dir, dset1, dset2, dset3)
dset1, dset2, dset3 = dset1.select([2, 1, 0]), dset2.select([2, 1, 0]), dset3
with concatenate_datasets([dset3, dset2, dset1]) as dset_concat:
self.assertTupleEqual((len(dset1), len(dset2), len(dset3)), (3, 3, 3))
self.assertEqual(len(dset_concat), len(dset1) + len(dset2) + len(dset3))
self.assertListEqual(dset_concat["id"], [6, 7, 8, 5, 4, 3, 2, 1, 0])
# in_memory = False:
# 3 cache files for the dset_concat._data table
# no cache file for the indices because it's in memory
# in_memory = True:
# no cache files since both dset_concat._data and dset_concat._indices are in memory
self.assertEqual(len(dset_concat.cache_files), 0 if in_memory else 3)
self.assertEqual(dset_concat.info.description, "Dataset2\n\nDataset1")
dset1 = dset1.rename_columns({"id": "id1"})
dset2 = dset2.rename_columns({"id": "id2"})
dset3 = dset3.rename_columns({"id": "id3"})
with concatenate_datasets([dset1, dset2, dset3], axis=1) as dset_concat:
self.assertTupleEqual((len(dset1), len(dset2), len(dset3)), (3, 3, 3))
self.assertEqual(len(dset_concat), len(dset1))
self.assertListEqual(dset_concat["id1"], [2, 1, 0])
self.assertListEqual(dset_concat["id2"], [5, 4, 3])
self.assertListEqual(dset_concat["id3"], [6, 7, 8])
# in_memory = False:
# 3 cache files for the dset_concat._data table
# no cache file for the indices because it's None
# in_memory = True:
# no cache files since dset_concat._data is in memory and dset_concat._indices is None
self.assertEqual(len(dset_concat.cache_files), 0 if in_memory else 3)
self.assertIsNone(dset_concat._indices)
self.assertEqual(dset_concat.info.description, "Dataset1\n\nDataset2")
with concatenate_datasets([dset1], axis=1) as dset_concat:
self.assertEqual(len(dset_concat), len(dset1))
self.assertListEqual(dset_concat["id1"], [2, 1, 0])
# in_memory = False:
# 1 cache file for the dset_concat._data table
# no cache file for the indices because it's in memory
# in_memory = True:
# no cache files since both dset_concat._data and dset_concat._indices are in memory
self.assertEqual(len(dset_concat.cache_files), 0 if in_memory else 1)
self.assertTrue(dset_concat._indices == dset1._indices)
self.assertEqual(dset_concat.info.description, "Dataset1")
del dset1, dset2, dset3
def test_concatenate_with_indices_from_disk(self, in_memory):
data1, data2, data3 = {"id": [0, 1, 2] * 2}, {"id": [3, 4, 5] * 2}, {"id": [6, 7]}
info1 = DatasetInfo(description="Dataset1")
info2 = DatasetInfo(description="Dataset2")
with tempfile.TemporaryDirectory() as tmp_dir:
dset1, dset2, dset3 = (
Dataset.from_dict(data1, info=info1),
Dataset.from_dict(data2, info=info2),
Dataset.from_dict(data3),
)
dset1, dset2, dset3 = self._to(in_memory, tmp_dir, dset1, dset2, dset3)
dset1, dset2, dset3 = (
dset1.select([2, 1, 0], indices_cache_file_name=os.path.join(tmp_dir, "i1.arrow")),
dset2.select([2, 1, 0], indices_cache_file_name=os.path.join(tmp_dir, "i2.arrow")),
dset3.select([1, 0], indices_cache_file_name=os.path.join(tmp_dir, "i3.arrow")),
)
with concatenate_datasets([dset3, dset2, dset1]) as dset_concat:
self.assertTupleEqual((len(dset1), len(dset2), len(dset3)), (3, 3, 2))
self.assertEqual(len(dset_concat), len(dset1) + len(dset2) + len(dset3))
self.assertListEqual(dset_concat["id"], [7, 6, 5, 4, 3, 2, 1, 0])
# in_memory = False:
# 3 cache files for the dset_concat._data table, and 1 for the dset_concat._indices_table
# There is only 1 for the indices tables (i1.arrow)
# Indeed, the others are brought to memory since an offset is applied to them.
# in_memory = True:
# 1 cache file for i1.arrow since both dset_concat._data and dset_concat._indices are in memory
self.assertEqual(len(dset_concat.cache_files), 1 if in_memory else 3 + 1)
self.assertEqual(dset_concat.info.description, "Dataset2\n\nDataset1")
del dset1, dset2, dset3
def test_concatenate_pickle(self, in_memory):
data1, data2, data3 = {"id": [0, 1, 2] * 2}, {"id": [3, 4, 5] * 2}, {"id": [6, 7], "foo": ["bar", "bar"]}
info1 = DatasetInfo(description="Dataset1")
info2 = DatasetInfo(description="Dataset2")
with tempfile.TemporaryDirectory() as tmp_dir:
dset1, dset2, dset3 = (
Dataset.from_dict(data1, info=info1),
Dataset.from_dict(data2, info=info2),
Dataset.from_dict(data3),
)
# mix from in-memory and on-disk datasets
dset1, dset2 = self._to(in_memory, tmp_dir, dset1, dset2)
dset3 = self._to(not in_memory, tmp_dir, dset3)
dset1, dset2, dset3 = (
dset1.select(
[2, 1, 0],
keep_in_memory=in_memory,
indices_cache_file_name=os.path.join(tmp_dir, "i1.arrow") if not in_memory else None,
),
dset2.select(
[2, 1, 0],
keep_in_memory=in_memory,
indices_cache_file_name=os.path.join(tmp_dir, "i2.arrow") if not in_memory else None,
),
dset3.select(
[1, 0],
keep_in_memory=in_memory,
indices_cache_file_name=os.path.join(tmp_dir, "i3.arrow") if not in_memory else None,
),
)
dset3 = dset3.rename_column("foo", "new_foo")
dset3 = dset3.remove_columns("new_foo")
if in_memory:
dset3._data.table = Unpicklable()
else:
dset1._data.table, dset2._data.table = Unpicklable(), Unpicklable()
dset1, dset2, dset3 = (pickle.loads(pickle.dumps(d)) for d in (dset1, dset2, dset3))
with concatenate_datasets([dset3, dset2, dset1]) as dset_concat:
if not in_memory:
dset_concat._data.table = Unpicklable()
with pickle.loads(pickle.dumps(dset_concat)) as dset_concat:
self.assertTupleEqual((len(dset1), len(dset2), len(dset3)), (3, 3, 2))
self.assertEqual(len(dset_concat), len(dset1) + len(dset2) + len(dset3))
self.assertListEqual(dset_concat["id"], [7, 6, 5, 4, 3, 2, 1, 0])
# in_memory = True: 1 cache file for dset3
# in_memory = False: 2 caches files for dset1 and dset2, and 1 cache file for i1.arrow
self.assertEqual(len(dset_concat.cache_files), 1 if in_memory else 2 + 1)
self.assertEqual(dset_concat.info.description, "Dataset2\n\nDataset1")
del dset1, dset2, dset3
def test_flatten(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
with Dataset.from_dict(
{"a": [{"b": {"c": ["text"]}}] * 10, "foo": [1] * 10},
features=Features({"a": {"b": Sequence({"c": Value("string")})}, "foo": Value("int64")}),
) as dset:
with self._to(in_memory, tmp_dir, dset) as dset:
fingerprint = dset._fingerprint
with dset.flatten() as dset:
self.assertListEqual(sorted(dset.column_names), ["a.b.c", "foo"])
self.assertListEqual(sorted(dset.features.keys()), ["a.b.c", "foo"])
self.assertDictEqual(
dset.features, Features({"a.b.c": Sequence(Value("string")), "foo": Value("int64")})
)
self.assertNotEqual(dset._fingerprint, fingerprint)
assert_arrow_metadata_are_synced_with_dataset_features(dset)
with tempfile.TemporaryDirectory() as tmp_dir:
with Dataset.from_dict(
{"a": [{"en": "Thank you", "fr": "Merci"}] * 10, "foo": [1] * 10},
features=Features({"a": Translation(languages=["en", "fr"]), "foo": Value("int64")}),
) as dset:
with self._to(in_memory, tmp_dir, dset) as dset:
fingerprint = dset._fingerprint
with dset.flatten() as dset:
self.assertListEqual(sorted(dset.column_names), ["a.en", "a.fr", "foo"])
self.assertListEqual(sorted(dset.features.keys()), ["a.en", "a.fr", "foo"])
self.assertDictEqual(
dset.features,
Features({"a.en": Value("string"), "a.fr": Value("string"), "foo": Value("int64")}),
)
self.assertNotEqual(dset._fingerprint, fingerprint)
assert_arrow_metadata_are_synced_with_dataset_features(dset)
with tempfile.TemporaryDirectory() as tmp_dir:
with Dataset.from_dict(
{"a": [{"en": "the cat", "fr": ["le chat", "la chatte"], "de": "die katze"}] * 10, "foo": [1] * 10},
features=Features(
{"a": TranslationVariableLanguages(languages=["en", "fr", "de"]), "foo": Value("int64")}
),
) as dset:
with self._to(in_memory, tmp_dir, dset) as dset:
fingerprint = dset._fingerprint
with dset.flatten() as dset:
self.assertListEqual(sorted(dset.column_names), ["a.language", "a.translation", "foo"])
self.assertListEqual(sorted(dset.features.keys()), ["a.language", "a.translation", "foo"])
self.assertDictEqual(
dset.features,
Features(
{
"a.language": Sequence(Value("string")),
"a.translation": Sequence(Value("string")),
"foo": Value("int64"),
}
),
)
self.assertNotEqual(dset._fingerprint, fingerprint)
assert_arrow_metadata_are_synced_with_dataset_features(dset)
@require_pil
def test_flatten_complex_image(self, in_memory):
# decoding turned on
with tempfile.TemporaryDirectory() as tmp_dir:
with Dataset.from_dict(
{"a": [np.arange(4 * 4 * 3, dtype=np.uint8).reshape(4, 4, 3)] * 10, "foo": [1] * 10},
features=Features({"a": Image(), "foo": Value("int64")}),
) as dset:
with self._to(in_memory, tmp_dir, dset) as dset:
fingerprint = dset._fingerprint
with dset.flatten() as dset:
self.assertListEqual(sorted(dset.column_names), ["a", "foo"])
self.assertListEqual(sorted(dset.features.keys()), ["a", "foo"])
self.assertDictEqual(dset.features, Features({"a": Image(), "foo": Value("int64")}))
self.assertNotEqual(dset._fingerprint, fingerprint)
assert_arrow_metadata_are_synced_with_dataset_features(dset)
# decoding turned on + nesting
with tempfile.TemporaryDirectory() as tmp_dir:
with Dataset.from_dict(
{"a": [{"b": np.arange(4 * 4 * 3, dtype=np.uint8).reshape(4, 4, 3)}] * 10, "foo": [1] * 10},
features=Features({"a": {"b": Image()}, "foo": Value("int64")}),
) as dset:
with self._to(in_memory, tmp_dir, dset) as dset:
fingerprint = dset._fingerprint
with dset.flatten() as dset:
self.assertListEqual(sorted(dset.column_names), ["a.b", "foo"])
self.assertListEqual(sorted(dset.features.keys()), ["a.b", "foo"])
self.assertDictEqual(dset.features, Features({"a.b": Image(), "foo": Value("int64")}))
self.assertNotEqual(dset._fingerprint, fingerprint)
assert_arrow_metadata_are_synced_with_dataset_features(dset)
# decoding turned off
with tempfile.TemporaryDirectory() as tmp_dir:
with Dataset.from_dict(
{"a": [np.arange(4 * 4 * 3, dtype=np.uint8).reshape(4, 4, 3)] * 10, "foo": [1] * 10},
features=Features({"a": Image(decode=False), "foo": Value("int64")}),
) as dset:
with self._to(in_memory, tmp_dir, dset) as dset:
fingerprint = dset._fingerprint
with dset.flatten() as dset:
self.assertListEqual(sorted(dset.column_names), ["a.bytes", "a.path", "foo"])
self.assertListEqual(sorted(dset.features.keys()), ["a.bytes", "a.path", "foo"])
self.assertDictEqual(
dset.features,
Features({"a.bytes": Value("binary"), "a.path": Value("string"), "foo": Value("int64")}),
)
self.assertNotEqual(dset._fingerprint, fingerprint)
assert_arrow_metadata_are_synced_with_dataset_features(dset)
# decoding turned off + nesting
with tempfile.TemporaryDirectory() as tmp_dir:
with Dataset.from_dict(
{"a": [{"b": np.arange(4 * 4 * 3, dtype=np.uint8).reshape(4, 4, 3)}] * 10, "foo": [1] * 10},
features=Features({"a": {"b": Image(decode=False)}, "foo": Value("int64")}),
) as dset:
with self._to(in_memory, tmp_dir, dset) as dset:
fingerprint = dset._fingerprint
with dset.flatten() as dset:
self.assertListEqual(sorted(dset.column_names), ["a.b.bytes", "a.b.path", "foo"])
self.assertListEqual(sorted(dset.features.keys()), ["a.b.bytes", "a.b.path", "foo"])
self.assertDictEqual(
dset.features,
Features(
{"a.b.bytes": Value("binary"), "a.b.path": Value("string"), "foo": Value("int64")}
),
)
self.assertNotEqual(dset._fingerprint, fingerprint)
assert_arrow_metadata_are_synced_with_dataset_features(dset)
def test_map(self, in_memory):
# standard
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
fingerprint = dset._fingerprint
with dset.map(
lambda x: {"name": x["filename"][:-2], "id": int(x["filename"].split("_")[-1])}
) as dset_test:
self.assertEqual(len(dset_test), 30)
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertDictEqual(
dset_test.features,
Features({"filename": Value("string"), "name": Value("string"), "id": Value("int64")}),
)
self.assertListEqual(dset_test["id"], list(range(30)))
self.assertNotEqual(dset_test._fingerprint, fingerprint)
assert_arrow_metadata_are_synced_with_dataset_features(dset_test)
# no transform
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
fingerprint = dset._fingerprint
with dset.map(lambda x: None) as dset_test:
self.assertEqual(len(dset_test), 30)
self.assertEqual(dset_test._fingerprint, fingerprint)
assert_arrow_metadata_are_synced_with_dataset_features(dset_test)
# with indices
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
with dset.map(
lambda x, i: {"name": x["filename"][:-2], "id": i}, with_indices=True
) as dset_test_with_indices:
self.assertEqual(len(dset_test_with_indices), 30)
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertDictEqual(
dset_test_with_indices.features,
Features({"filename": Value("string"), "name": Value("string"), "id": Value("int64")}),
)
self.assertListEqual(dset_test_with_indices["id"], list(range(30)))
assert_arrow_metadata_are_synced_with_dataset_features(dset_test_with_indices)
# interrupted
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
def func(x, i):
if i == 4:
raise KeyboardInterrupt()
return {"name": x["filename"][:-2], "id": i}
tmp_file = os.path.join(tmp_dir, "test.arrow")
self.assertRaises(
KeyboardInterrupt,
dset.map,
function=func,
with_indices=True,
cache_file_name=tmp_file,
writer_batch_size=2,
)
self.assertFalse(os.path.exists(tmp_file))
with dset.map(
lambda x, i: {"name": x["filename"][:-2], "id": i},
with_indices=True,
cache_file_name=tmp_file,
writer_batch_size=2,
) as dset_test_with_indices:
self.assertTrue(os.path.exists(tmp_file))
self.assertEqual(len(dset_test_with_indices), 30)
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertDictEqual(
dset_test_with_indices.features,
Features({"filename": Value("string"), "name": Value("string"), "id": Value("int64")}),
)
self.assertListEqual(dset_test_with_indices["id"], list(range(30)))
assert_arrow_metadata_are_synced_with_dataset_features(dset_test_with_indices)
# formatted
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
dset.set_format("numpy", columns=["col_1"])
with dset.map(lambda x: {"col_1_plus_one": x["col_1"] + 1}) as dset_test:
self.assertEqual(len(dset_test), 4)
self.assertEqual(dset_test.format["type"], "numpy")
self.assertIsInstance(dset_test["col_1"], np.ndarray)
self.assertIsInstance(dset_test["col_1_plus_one"], np.ndarray)
self.assertListEqual(sorted(dset_test[0].keys()), ["col_1", "col_1_plus_one"])
self.assertListEqual(sorted(dset_test.column_names), ["col_1", "col_1_plus_one", "col_2", "col_3"])
assert_arrow_metadata_are_synced_with_dataset_features(dset_test)
def test_map_multiprocessing(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir: # standard
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
fingerprint = dset._fingerprint
with dset.map(picklable_map_function, num_proc=2) as dset_test:
self.assertEqual(len(dset_test), 30)
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertDictEqual(
dset_test.features,
Features({"filename": Value("string"), "id": Value("int64")}),
)
self.assertEqual(len(dset_test.cache_files), 0 if in_memory else 2)
if not in_memory:
self.assertIn("_of_00002.arrow", dset_test.cache_files[0]["filename"])
self.assertListEqual(dset_test["id"], list(range(30)))
self.assertNotEqual(dset_test._fingerprint, fingerprint)
assert_arrow_metadata_are_synced_with_dataset_features(dset_test)
with tempfile.TemporaryDirectory() as tmp_dir: # num_proc > num rows
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
fingerprint = dset._fingerprint
with dset.select([0, 1], keep_in_memory=True).map(picklable_map_function, num_proc=10) as dset_test:
self.assertEqual(len(dset_test), 2)
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertDictEqual(
dset_test.features,
Features({"filename": Value("string"), "id": Value("int64")}),
)
self.assertEqual(len(dset_test.cache_files), 0 if in_memory else 2)
self.assertListEqual(dset_test["id"], list(range(2)))
self.assertNotEqual(dset_test._fingerprint, fingerprint)
assert_arrow_metadata_are_synced_with_dataset_features(dset_test)
with tempfile.TemporaryDirectory() as tmp_dir: # with_indices
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
fingerprint = dset._fingerprint
with dset.map(picklable_map_function_with_indices, num_proc=3, with_indices=True) as dset_test:
self.assertEqual(len(dset_test), 30)
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertDictEqual(
dset_test.features,
Features({"filename": Value("string"), "id": Value("int64")}),
)
self.assertEqual(len(dset_test.cache_files), 0 if in_memory else 3)
self.assertListEqual(dset_test["id"], list(range(30)))
self.assertNotEqual(dset_test._fingerprint, fingerprint)
assert_arrow_metadata_are_synced_with_dataset_features(dset_test)
with tempfile.TemporaryDirectory() as tmp_dir: # with_rank
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
fingerprint = dset._fingerprint
with dset.map(picklable_map_function_with_rank, num_proc=3, with_rank=True) as dset_test:
self.assertEqual(len(dset_test), 30)
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertDictEqual(
dset_test.features,
Features({"filename": Value("string"), "rank": Value("int64")}),
)
self.assertEqual(len(dset_test.cache_files), 0 if in_memory else 3)
self.assertListEqual(dset_test["rank"], [0] * 10 + [1] * 10 + [2] * 10)
self.assertNotEqual(dset_test._fingerprint, fingerprint)
assert_arrow_metadata_are_synced_with_dataset_features(dset_test)
with tempfile.TemporaryDirectory() as tmp_dir: # with_indices AND with_rank
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
fingerprint = dset._fingerprint
with dset.map(
picklable_map_function_with_indices_and_rank, num_proc=3, with_indices=True, with_rank=True
) as dset_test:
self.assertEqual(len(dset_test), 30)
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertDictEqual(
dset_test.features,
Features({"filename": Value("string"), "id": Value("int64"), "rank": Value("int64")}),
)
self.assertEqual(len(dset_test.cache_files), 0 if in_memory else 3)
self.assertListEqual(dset_test["id"], list(range(30)))
self.assertListEqual(dset_test["rank"], [0] * 10 + [1] * 10 + [2] * 10)
self.assertNotEqual(dset_test._fingerprint, fingerprint)
assert_arrow_metadata_are_synced_with_dataset_features(dset_test)
with tempfile.TemporaryDirectory() as tmp_dir: # new_fingerprint
new_fingerprint = "foobar"
invalid_new_fingerprint = "foobar/hey"
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
fingerprint = dset._fingerprint
self.assertRaises(
ValueError, dset.map, picklable_map_function, num_proc=2, new_fingerprint=invalid_new_fingerprint
)
with dset.map(picklable_map_function, num_proc=2, new_fingerprint=new_fingerprint) as dset_test:
self.assertEqual(len(dset_test), 30)
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertDictEqual(
dset_test.features,
Features({"filename": Value("string"), "id": Value("int64")}),
)
self.assertEqual(len(dset_test.cache_files), 0 if in_memory else 2)
self.assertListEqual(dset_test["id"], list(range(30)))
self.assertNotEqual(dset_test._fingerprint, fingerprint)
self.assertEqual(dset_test._fingerprint, new_fingerprint)
assert_arrow_metadata_are_synced_with_dataset_features(dset_test)
file_names = sorted(Path(cache_file["filename"]).name for cache_file in dset_test.cache_files)
for i, file_name in enumerate(file_names):
self.assertIn(new_fingerprint + f"_{i:05d}", file_name)
with tempfile.TemporaryDirectory() as tmp_dir: # lambda (requires multiprocess from pathos)
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
fingerprint = dset._fingerprint
with dset.map(lambda x: {"id": int(x["filename"].split("_")[-1])}, num_proc=2) as dset_test:
self.assertEqual(len(dset_test), 30)
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertDictEqual(
dset_test.features,
Features({"filename": Value("string"), "id": Value("int64")}),
)
self.assertEqual(len(dset_test.cache_files), 0 if in_memory else 2)
self.assertListEqual(dset_test["id"], list(range(30)))
self.assertNotEqual(dset_test._fingerprint, fingerprint)
assert_arrow_metadata_are_synced_with_dataset_features(dset_test)
def test_map_new_features(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
features = Features({"filename": Value("string"), "label": ClassLabel(names=["positive", "negative"])})
with dset.map(
lambda x, i: {"label": i % 2}, with_indices=True, features=features
) as dset_test_with_indices:
self.assertEqual(len(dset_test_with_indices), 30)
self.assertDictEqual(
dset_test_with_indices.features,
features,
)
assert_arrow_metadata_are_synced_with_dataset_features(dset_test_with_indices)
def test_map_batched(self, in_memory):
def map_batched(example):
return {"filename_new": [x + "_extension" for x in example["filename"]]}
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
with dset.map(map_batched, batched=True) as dset_test_batched:
self.assertEqual(len(dset_test_batched), 30)
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertDictEqual(
dset_test_batched.features,
Features({"filename": Value("string"), "filename_new": Value("string")}),
)
assert_arrow_metadata_are_synced_with_dataset_features(dset_test_batched)
# change batch size and drop the last batch
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
batch_size = 4
with dset.map(
map_batched, batched=True, batch_size=batch_size, drop_last_batch=True
) as dset_test_batched:
self.assertEqual(len(dset_test_batched), 30 // batch_size * batch_size)
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertDictEqual(
dset_test_batched.features,
Features({"filename": Value("string"), "filename_new": Value("string")}),
)
assert_arrow_metadata_are_synced_with_dataset_features(dset_test_batched)
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
with dset.formatted_as("numpy", columns=["filename"]):
with dset.map(map_batched, batched=True) as dset_test_batched:
self.assertEqual(len(dset_test_batched), 30)
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertDictEqual(
dset_test_batched.features,
Features({"filename": Value("string"), "filename_new": Value("string")}),
)
assert_arrow_metadata_are_synced_with_dataset_features(dset_test_batched)
def map_batched_with_indices(example, idx):
return {"filename_new": [x + "_extension_" + str(idx) for x in example["filename"]]}
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
with dset.map(
map_batched_with_indices, batched=True, with_indices=True
) as dset_test_with_indices_batched:
self.assertEqual(len(dset_test_with_indices_batched), 30)
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertDictEqual(
dset_test_with_indices_batched.features,
Features({"filename": Value("string"), "filename_new": Value("string")}),
)
assert_arrow_metadata_are_synced_with_dataset_features(dset_test_with_indices_batched)
# check remove columns for even if the function modifies input in-place
def map_batched_modifying_inputs_inplace(example):
result = {"filename_new": [x + "_extension" for x in example["filename"]]}
del example["filename"]
return result
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
with dset.map(
map_batched_modifying_inputs_inplace, batched=True, remove_columns="filename"
) as dset_test_modifying_inputs_inplace:
self.assertEqual(len(dset_test_modifying_inputs_inplace), 30)
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertDictEqual(
dset_test_modifying_inputs_inplace.features,
Features({"filename_new": Value("string")}),
)
assert_arrow_metadata_are_synced_with_dataset_features(dset_test_modifying_inputs_inplace)
def test_map_nested(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
with Dataset.from_dict({"field": ["a", "b"]}) as dset:
with self._to(in_memory, tmp_dir, dset) as dset:
with dset.map(lambda example: {"otherfield": {"capital": example["field"].capitalize()}}) as dset:
with dset.map(lambda example: {"otherfield": {"append_x": example["field"] + "x"}}) as dset:
self.assertEqual(dset[0], {"field": "a", "otherfield": {"append_x": "ax"}})
def test_map_return_example_as_dict_value(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
with Dataset.from_dict({"en": ["aa", "bb"], "fr": ["cc", "dd"]}) as dset:
with self._to(in_memory, tmp_dir, dset) as dset:
with dset.map(lambda example: {"translation": example}) as dset:
self.assertEqual(dset[0], {"en": "aa", "fr": "cc", "translation": {"en": "aa", "fr": "cc"}})
def test_map_fn_kwargs(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
with Dataset.from_dict({"id": range(10)}) as dset:
with self._to(in_memory, tmp_dir, dset) as dset:
fn_kwargs = {"offset": 3}
with dset.map(
lambda example, offset: {"id+offset": example["id"] + offset}, fn_kwargs=fn_kwargs
) as mapped_dset:
assert mapped_dset["id+offset"] == list(range(3, 13))
with dset.map(
lambda id, offset: {"id+offset": id + offset}, fn_kwargs=fn_kwargs, input_columns="id"
) as mapped_dset:
assert mapped_dset["id+offset"] == list(range(3, 13))
with dset.map(
lambda id, i, offset: {"id+offset": i + offset},
fn_kwargs=fn_kwargs,
input_columns="id",
with_indices=True,
) as mapped_dset:
assert mapped_dset["id+offset"] == list(range(3, 13))
def test_map_caching(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
self._caplog.clear()
with self._caplog.at_level(INFO, logger=get_logger().name):
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
with patch(
"datasets.arrow_dataset.Dataset._map_single",
autospec=Dataset._map_single,
side_effect=Dataset._map_single,
) as mock_map_single:
with dset.map(lambda x: {"foo": "bar"}) as dset_test1:
dset_test1_data_files = list(dset_test1.cache_files)
self.assertEqual(mock_map_single.call_count, 1)
with dset.map(lambda x: {"foo": "bar"}) as dset_test2:
self.assertEqual(dset_test1_data_files, dset_test2.cache_files)
self.assertEqual(len(dset_test2.cache_files), 1 - int(in_memory))
self.assertTrue(("Loading cached processed dataset" in self._caplog.text) ^ in_memory)
self.assertEqual(mock_map_single.call_count, 2 if in_memory else 1)
with tempfile.TemporaryDirectory() as tmp_dir:
self._caplog.clear()
with self._caplog.at_level(INFO, logger=get_logger().name):
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
with dset.map(lambda x: {"foo": "bar"}) as dset_test1:
dset_test1_data_files = list(dset_test1.cache_files)
with dset.map(lambda x: {"foo": "bar"}, load_from_cache_file=False) as dset_test2:
self.assertEqual(dset_test1_data_files, dset_test2.cache_files)
self.assertEqual(len(dset_test2.cache_files), 1 - int(in_memory))
self.assertNotIn("Loading cached processed dataset", self._caplog.text)
with tempfile.TemporaryDirectory() as tmp_dir:
self._caplog.clear()
with self._caplog.at_level(INFO, logger=get_logger().name):
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
with patch(
"datasets.arrow_dataset.Pool",
new_callable=PickableMagicMock,
side_effect=datasets.arrow_dataset.Pool,
) as mock_pool:
with dset.map(lambda x: {"foo": "bar"}, num_proc=2) as dset_test1:
dset_test1_data_files = list(dset_test1.cache_files)
self.assertEqual(mock_pool.call_count, 1)
with dset.map(lambda x: {"foo": "bar"}, num_proc=2) as dset_test2:
self.assertEqual(dset_test1_data_files, dset_test2.cache_files)
self.assertTrue(
(len(re.findall("Loading cached processed dataset", self._caplog.text)) == 1)
^ in_memory
)
self.assertEqual(mock_pool.call_count, 2 if in_memory else 1)
with tempfile.TemporaryDirectory() as tmp_dir:
self._caplog.clear()
with self._caplog.at_level(INFO, logger=get_logger().name):
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
with dset.map(lambda x: {"foo": "bar"}, num_proc=2) as dset_test1:
dset_test1_data_files = list(dset_test1.cache_files)
with dset.map(lambda x: {"foo": "bar"}, num_proc=2, load_from_cache_file=False) as dset_test2:
self.assertEqual(dset_test1_data_files, dset_test2.cache_files)
self.assertEqual(len(dset_test2.cache_files), (1 - int(in_memory)) * 2)
self.assertNotIn("Loading cached processed dataset", self._caplog.text)
if not in_memory:
try:
self._caplog.clear()
with tempfile.TemporaryDirectory() as tmp_dir:
with self._caplog.at_level(INFO, logger=get_logger().name):
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
datasets.disable_caching()
with dset.map(lambda x: {"foo": "bar"}) as dset_test1:
with dset.map(lambda x: {"foo": "bar"}) as dset_test2:
self.assertNotEqual(dset_test1.cache_files, dset_test2.cache_files)
self.assertEqual(len(dset_test1.cache_files), 1)
self.assertEqual(len(dset_test2.cache_files), 1)
self.assertNotIn("Loading cached processed dataset", self._caplog.text)
# make sure the arrow files are going to be removed
self.assertIn("tmp", dset_test1.cache_files[0]["filename"])
self.assertIn("tmp", dset_test2.cache_files[0]["filename"])
finally:
datasets.enable_caching()
def test_map_return_pa_table(self, in_memory):
def func_return_single_row_pa_table(x):
return pa.table({"id": [0], "text": ["a"]})
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
with dset.map(func_return_single_row_pa_table) as dset_test:
self.assertEqual(len(dset_test), 30)
self.assertDictEqual(
dset_test.features,
Features({"id": Value("int64"), "text": Value("string")}),
)
self.assertEqual(dset_test[0]["id"], 0)
self.assertEqual(dset_test[0]["text"], "a")
# Batched
def func_return_single_row_pa_table_batched(x):
batch_size = len(x[next(iter(x))])
return pa.table({"id": [0] * batch_size, "text": ["a"] * batch_size})
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
with dset.map(func_return_single_row_pa_table_batched, batched=True) as dset_test:
self.assertEqual(len(dset_test), 30)
self.assertDictEqual(
dset_test.features,
Features({"id": Value("int64"), "text": Value("string")}),
)
self.assertEqual(dset_test[0]["id"], 0)
self.assertEqual(dset_test[0]["text"], "a")
# Error when returning a table with more than one row in the non-batched mode
def func_return_multi_row_pa_table(x):
return pa.table({"id": [0, 1], "text": ["a", "b"]})
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
self.assertRaises(ValueError, dset.map, func_return_multi_row_pa_table)
def test_map_return_pd_dataframe(self, in_memory):
def func_return_single_row_pd_dataframe(x):
return pd.DataFrame({"id": [0], "text": ["a"]})
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
with dset.map(func_return_single_row_pd_dataframe) as dset_test:
self.assertEqual(len(dset_test), 30)
self.assertDictEqual(
dset_test.features,
Features({"id": Value("int64"), "text": Value("string")}),
)
self.assertEqual(dset_test[0]["id"], 0)
self.assertEqual(dset_test[0]["text"], "a")
# Batched
def func_return_single_row_pd_dataframe_batched(x):
batch_size = len(x[next(iter(x))])
return pd.DataFrame({"id": [0] * batch_size, "text": ["a"] * batch_size})
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
with dset.map(func_return_single_row_pd_dataframe_batched, batched=True) as dset_test:
self.assertEqual(len(dset_test), 30)
self.assertDictEqual(
dset_test.features,
Features({"id": Value("int64"), "text": Value("string")}),
)
self.assertEqual(dset_test[0]["id"], 0)
self.assertEqual(dset_test[0]["text"], "a")
# Error when returning a table with more than one row in the non-batched mode
def func_return_multi_row_pd_dataframe(x):
return pd.DataFrame({"id": [0, 1], "text": ["a", "b"]})
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
self.assertRaises(ValueError, dset.map, func_return_multi_row_pd_dataframe)
@require_torch
def test_map_torch(self, in_memory):
import torch
def func(example):
return {"tensor": torch.tensor([1.0, 2, 3])}
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
with dset.map(func) as dset_test:
self.assertEqual(len(dset_test), 30)
self.assertDictEqual(
dset_test.features,
Features({"filename": Value("string"), "tensor": Sequence(Value("float32"))}),
)
self.assertListEqual(dset_test[0]["tensor"], [1, 2, 3])
@require_tf
def test_map_tf(self, in_memory):
import tensorflow as tf
def func(example):
return {"tensor": tf.constant([1.0, 2, 3])}
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
with dset.map(func) as dset_test:
self.assertEqual(len(dset_test), 30)
self.assertDictEqual(
dset_test.features,
Features({"filename": Value("string"), "tensor": Sequence(Value("float32"))}),
)
self.assertListEqual(dset_test[0]["tensor"], [1, 2, 3])
@require_jax
def test_map_jax(self, in_memory):
import jax.numpy as jnp
def func(example):
return {"tensor": jnp.asarray([1.0, 2, 3])}
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
with dset.map(func) as dset_test:
self.assertEqual(len(dset_test), 30)
self.assertDictEqual(
dset_test.features,
Features({"filename": Value("string"), "tensor": Sequence(Value("float32"))}),
)
self.assertListEqual(dset_test[0]["tensor"], [1, 2, 3])
def test_map_numpy(self, in_memory):
def func(example):
return {"tensor": np.array([1.0, 2, 3])}
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
with dset.map(func) as dset_test:
self.assertEqual(len(dset_test), 30)
self.assertDictEqual(
dset_test.features,
Features({"filename": Value("string"), "tensor": Sequence(Value("float64"))}),
)
self.assertListEqual(dset_test[0]["tensor"], [1, 2, 3])
@require_torch
def test_map_tensor_batched(self, in_memory):
import torch
def func(batch):
return {"tensor": torch.tensor([[1.0, 2, 3]] * len(batch["filename"]))}
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
with dset.map(func, batched=True) as dset_test:
self.assertEqual(len(dset_test), 30)
self.assertDictEqual(
dset_test.features,
Features({"filename": Value("string"), "tensor": Sequence(Value("float32"))}),
)
self.assertListEqual(dset_test[0]["tensor"], [1, 2, 3])
def test_map_input_columns(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
with dset.map(lambda col_1: {"label": col_1 % 2}, input_columns="col_1") as mapped_dset:
self.assertEqual(mapped_dset[0].keys(), {"col_1", "col_2", "col_3", "label"})
self.assertEqual(
mapped_dset.features,
Features(
{
"col_1": Value("int64"),
"col_2": Value("string"),
"col_3": Value("bool"),
"label": Value("int64"),
}
),
)
def test_map_remove_columns(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
with dset.map(lambda x, i: {"name": x["filename"][:-2], "id": i}, with_indices=True) as dset:
self.assertTrue("id" in dset[0])
self.assertDictEqual(
dset.features,
Features({"filename": Value("string"), "name": Value("string"), "id": Value("int64")}),
)
assert_arrow_metadata_are_synced_with_dataset_features(dset)
with dset.map(lambda x: x, remove_columns=["id"]) as mapped_dset:
self.assertTrue("id" not in mapped_dset[0])
self.assertDictEqual(
mapped_dset.features, Features({"filename": Value("string"), "name": Value("string")})
)
assert_arrow_metadata_are_synced_with_dataset_features(mapped_dset)
with mapped_dset.with_format("numpy", columns=mapped_dset.column_names) as mapped_dset:
with mapped_dset.map(
lambda x: {"name": 1}, remove_columns=mapped_dset.column_names
) as mapped_dset:
self.assertTrue("filename" not in mapped_dset[0])
self.assertTrue("name" in mapped_dset[0])
self.assertDictEqual(mapped_dset.features, Features({"name": Value(dtype="int64")}))
assert_arrow_metadata_are_synced_with_dataset_features(mapped_dset)
# empty dataset
columns_names = dset.column_names
with dset.select([]) as empty_dset:
self.assertEqual(len(empty_dset), 0)
with empty_dset.map(lambda x: {}, remove_columns=columns_names[0]) as mapped_dset:
self.assertListEqual(columns_names[1:], mapped_dset.column_names)
assert_arrow_metadata_are_synced_with_dataset_features(mapped_dset)
def test_map_stateful_callable(self, in_memory):
# be sure that the state of the map callable is unaffected
# before processing the dataset examples
class ExampleCounter:
def __init__(self, batched=False):
self.batched = batched
# state
self.cnt = 0
def __call__(self, example):
if self.batched:
self.cnt += len(example)
else:
self.cnt += 1
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
ex_cnt = ExampleCounter()
dset.map(ex_cnt)
self.assertEqual(ex_cnt.cnt, len(dset))
ex_cnt = ExampleCounter(batched=True)
dset.map(ex_cnt)
self.assertEqual(ex_cnt.cnt, len(dset))
@require_not_windows
def test_map_crash_subprocess(self, in_memory):
# be sure that a crash in one of the subprocess will not
# hang dataset.map() call forever
def do_crash(row):
import os
os.kill(os.getpid(), 9)
return row
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
with pytest.raises(RuntimeError) as excinfo:
dset.map(do_crash, num_proc=2)
assert str(excinfo.value) == (
"One of the subprocesses has abruptly died during map operation."
"To debug the error, disable multiprocessing."
)
def test_filter(self, in_memory):
# keep only first five examples
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
fingerprint = dset._fingerprint
with dset.filter(lambda x, i: i < 5, with_indices=True) as dset_filter_first_five:
self.assertEqual(len(dset_filter_first_five), 5)
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertDictEqual(dset_filter_first_five.features, Features({"filename": Value("string")}))
self.assertNotEqual(dset_filter_first_five._fingerprint, fingerprint)
# filter filenames with even id at the end + formatted
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
dset.set_format("numpy")
fingerprint = dset._fingerprint
with dset.filter(lambda x: (int(x["filename"][-1]) % 2 == 0)) as dset_filter_even_num:
self.assertEqual(len(dset_filter_even_num), 15)
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertDictEqual(dset_filter_even_num.features, Features({"filename": Value("string")}))
self.assertNotEqual(dset_filter_even_num._fingerprint, fingerprint)
self.assertEqual(dset_filter_even_num.format["type"], "numpy")
def test_filter_with_indices_mapping(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
dset = Dataset.from_dict({"col": [0, 1, 2]})
with self._to(in_memory, tmp_dir, dset) as dset:
with dset.filter(lambda x: x["col"] > 0) as dset:
self.assertListEqual(dset["col"], [1, 2])
with dset.filter(lambda x: x["col"] < 2) as dset:
self.assertListEqual(dset["col"], [1])
def test_filter_empty(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
self.assertIsNone(dset._indices, None)
tmp_file = os.path.join(tmp_dir, "test.arrow")
with dset.filter(lambda _: False, cache_file_name=tmp_file) as dset:
self.assertEqual(len(dset), 0)
self.assertIsNotNone(dset._indices, None)
tmp_file_2 = os.path.join(tmp_dir, "test_2.arrow")
with dset.filter(lambda _: False, cache_file_name=tmp_file_2) as dset2:
self.assertEqual(len(dset2), 0)
self.assertEqual(dset._indices, dset2._indices)
def test_filter_batched(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
dset = Dataset.from_dict({"col": [0, 1, 2]})
with self._to(in_memory, tmp_dir, dset) as dset:
with dset.filter(lambda x: [i > 0 for i in x["col"]], batched=True) as dset:
self.assertListEqual(dset["col"], [1, 2])
with dset.filter(lambda x: [i < 2 for i in x["col"]], batched=True) as dset:
self.assertListEqual(dset["col"], [1])
def test_filter_input_columns(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
dset = Dataset.from_dict({"col_1": [0, 1, 2], "col_2": ["a", "b", "c"]})
with self._to(in_memory, tmp_dir, dset) as dset:
with dset.filter(lambda x: x > 0, input_columns=["col_1"]) as filtered_dset:
self.assertListEqual(filtered_dset.column_names, dset.column_names)
self.assertListEqual(filtered_dset["col_1"], [1, 2])
self.assertListEqual(filtered_dset["col_2"], ["b", "c"])
def test_filter_fn_kwargs(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
with Dataset.from_dict({"id": range(10)}) as dset:
with self._to(in_memory, tmp_dir, dset) as dset:
fn_kwargs = {"max_offset": 3}
with dset.filter(
lambda example, max_offset: example["id"] < max_offset, fn_kwargs=fn_kwargs
) as filtered_dset:
assert len(filtered_dset) == 3
with dset.filter(
lambda id, max_offset: id < max_offset, fn_kwargs=fn_kwargs, input_columns="id"
) as filtered_dset:
assert len(filtered_dset) == 3
with dset.filter(
lambda id, i, max_offset: i < max_offset,
fn_kwargs=fn_kwargs,
input_columns="id",
with_indices=True,
) as filtered_dset:
assert len(filtered_dset) == 3
def test_filter_multiprocessing(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
fingerprint = dset._fingerprint
with dset.filter(picklable_filter_function, num_proc=2) as dset_filter_first_ten:
self.assertEqual(len(dset_filter_first_ten), 10)
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertDictEqual(dset_filter_first_ten.features, Features({"filename": Value("string")}))
self.assertEqual(len(dset_filter_first_ten.cache_files), 0 if in_memory else 2)
self.assertNotEqual(dset_filter_first_ten._fingerprint, fingerprint)
def test_filter_caching(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
self._caplog.clear()
with self._caplog.at_level(INFO, logger=get_logger().name):
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
with dset.filter(lambda x, i: i < 5, with_indices=True) as dset_filter_first_five1:
dset_test1_data_files = list(dset_filter_first_five1.cache_files)
with dset.filter(lambda x, i: i < 5, with_indices=True) as dset_filter_first_five2:
self.assertEqual(dset_test1_data_files, dset_filter_first_five2.cache_files)
self.assertEqual(len(dset_filter_first_five2.cache_files), 0 if in_memory else 2)
self.assertTrue(("Loading cached processed dataset" in self._caplog.text) ^ in_memory)
def test_keep_features_after_transform_specified(self, in_memory):
features = Features(
{"tokens": Sequence(Value("string")), "labels": Sequence(ClassLabel(names=["negative", "positive"]))}
)
def invert_labels(x):
return {"labels": [(1 - label) for label in x["labels"]]}
with tempfile.TemporaryDirectory() as tmp_dir:
with Dataset.from_dict(
{"tokens": [["foo"] * 5] * 10, "labels": [[1] * 5] * 10}, features=features
) as dset:
with self._to(in_memory, tmp_dir, dset) as dset:
with dset.map(invert_labels, features=features) as inverted_dset:
self.assertEqual(inverted_dset.features.type, features.type)
self.assertDictEqual(inverted_dset.features, features)
assert_arrow_metadata_are_synced_with_dataset_features(inverted_dset)
def test_keep_features_after_transform_unspecified(self, in_memory):
features = Features(
{"tokens": Sequence(Value("string")), "labels": Sequence(ClassLabel(names=["negative", "positive"]))}
)
def invert_labels(x):
return {"labels": [(1 - label) for label in x["labels"]]}
with tempfile.TemporaryDirectory() as tmp_dir:
with Dataset.from_dict(
{"tokens": [["foo"] * 5] * 10, "labels": [[1] * 5] * 10}, features=features
) as dset:
with self._to(in_memory, tmp_dir, dset) as dset:
with dset.map(invert_labels) as inverted_dset:
self.assertEqual(inverted_dset.features.type, features.type)
self.assertDictEqual(inverted_dset.features, features)
assert_arrow_metadata_are_synced_with_dataset_features(inverted_dset)
def test_keep_features_after_transform_to_file(self, in_memory):
features = Features(
{"tokens": Sequence(Value("string")), "labels": Sequence(ClassLabel(names=["negative", "positive"]))}
)
def invert_labels(x):
return {"labels": [(1 - label) for label in x["labels"]]}
with tempfile.TemporaryDirectory() as tmp_dir:
with Dataset.from_dict(
{"tokens": [["foo"] * 5] * 10, "labels": [[1] * 5] * 10}, features=features
) as dset:
with self._to(in_memory, tmp_dir, dset) as dset:
tmp_file = os.path.join(tmp_dir, "test.arrow")
dset.map(invert_labels, cache_file_name=tmp_file)
with Dataset.from_file(tmp_file) as inverted_dset:
self.assertEqual(inverted_dset.features.type, features.type)
self.assertDictEqual(inverted_dset.features, features)
def test_keep_features_after_transform_to_memory(self, in_memory):
features = Features(
{"tokens": Sequence(Value("string")), "labels": Sequence(ClassLabel(names=["negative", "positive"]))}
)
def invert_labels(x):
return {"labels": [(1 - label) for label in x["labels"]]}
with tempfile.TemporaryDirectory() as tmp_dir:
with Dataset.from_dict(
{"tokens": [["foo"] * 5] * 10, "labels": [[1] * 5] * 10}, features=features
) as dset:
with self._to(in_memory, tmp_dir, dset) as dset:
with dset.map(invert_labels, keep_in_memory=True) as inverted_dset:
self.assertEqual(inverted_dset.features.type, features.type)
self.assertDictEqual(inverted_dset.features, features)
def test_keep_features_after_loading_from_cache(self, in_memory):
features = Features(
{"tokens": Sequence(Value("string")), "labels": Sequence(ClassLabel(names=["negative", "positive"]))}
)
def invert_labels(x):
return {"labels": [(1 - label) for label in x["labels"]]}
with tempfile.TemporaryDirectory() as tmp_dir:
with Dataset.from_dict(
{"tokens": [["foo"] * 5] * 10, "labels": [[1] * 5] * 10}, features=features
) as dset:
with self._to(in_memory, tmp_dir, dset) as dset:
tmp_file1 = os.path.join(tmp_dir, "test1.arrow")
tmp_file2 = os.path.join(tmp_dir, "test2.arrow")
# TODO: Why mapped twice?
inverted_dset = dset.map(invert_labels, cache_file_name=tmp_file1)
inverted_dset = dset.map(invert_labels, cache_file_name=tmp_file2)
self.assertGreater(len(inverted_dset.cache_files), 0)
self.assertEqual(inverted_dset.features.type, features.type)
self.assertDictEqual(inverted_dset.features, features)
del inverted_dset
def test_keep_features_with_new_features(self, in_memory):
features = Features(
{"tokens": Sequence(Value("string")), "labels": Sequence(ClassLabel(names=["negative", "positive"]))}
)
def invert_labels(x):
return {"labels": [(1 - label) for label in x["labels"]], "labels2": x["labels"]}
expected_features = Features(
{
"tokens": Sequence(Value("string")),
"labels": Sequence(ClassLabel(names=["negative", "positive"])),
"labels2": Sequence(Value("int64")),
}
)
with tempfile.TemporaryDirectory() as tmp_dir:
with Dataset.from_dict(
{"tokens": [["foo"] * 5] * 10, "labels": [[1] * 5] * 10}, features=features
) as dset:
with self._to(in_memory, tmp_dir, dset) as dset:
with dset.map(invert_labels) as inverted_dset:
self.assertEqual(inverted_dset.features.type, expected_features.type)
self.assertDictEqual(inverted_dset.features, expected_features)
assert_arrow_metadata_are_synced_with_dataset_features(inverted_dset)
def test_select(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
# select every two example
indices = list(range(0, len(dset), 2))
tmp_file = os.path.join(tmp_dir, "test.arrow")
fingerprint = dset._fingerprint
with dset.select(indices, indices_cache_file_name=tmp_file) as dset_select_even:
self.assertIsNotNone(dset_select_even._indices) # an indices mapping is created
self.assertTrue(os.path.exists(tmp_file))
self.assertEqual(len(dset_select_even), 15)
for row in dset_select_even:
self.assertEqual(int(row["filename"][-1]) % 2, 0)
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertDictEqual(dset_select_even.features, Features({"filename": Value("string")}))
self.assertNotEqual(dset_select_even._fingerprint, fingerprint)
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
indices = list(range(0, len(dset)))
with dset.select(indices) as dset_select_all:
# no indices mapping, since the indices are contiguous
# (in this case the arrow table is simply sliced, which is more efficient)
self.assertIsNone(dset_select_all._indices)
self.assertEqual(len(dset_select_all), len(dset))
self.assertListEqual(list(dset_select_all), list(dset))
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertDictEqual(dset_select_all.features, Features({"filename": Value("string")}))
self.assertNotEqual(dset_select_all._fingerprint, fingerprint)
indices = range(0, len(dset))
with dset.select(indices) as dset_select_all:
# same but with range
self.assertIsNone(dset_select_all._indices)
self.assertEqual(len(dset_select_all), len(dset))
self.assertListEqual(list(dset_select_all), list(dset))
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertDictEqual(dset_select_all.features, Features({"filename": Value("string")}))
self.assertNotEqual(dset_select_all._fingerprint, fingerprint)
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
bad_indices = list(range(5))
bad_indices[-1] = len(dset) + 10 # out of bounds
tmp_file = os.path.join(tmp_dir, "test.arrow")
self.assertRaises(
Exception,
dset.select,
indices=bad_indices,
indices_cache_file_name=tmp_file,
writer_batch_size=2,
)
self.assertFalse(os.path.exists(tmp_file))
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
indices = iter(range(len(dset))) # iterator of contiguous indices
with dset.select(indices) as dset_select_all:
# no indices mapping, since the indices are contiguous
self.assertIsNone(dset_select_all._indices)
self.assertEqual(len(dset_select_all), len(dset))
indices = reversed(range(len(dset))) # iterator of not contiguous indices
tmp_file = os.path.join(tmp_dir, "test.arrow")
with dset.select(indices, indices_cache_file_name=tmp_file) as dset_select_all:
# new indices mapping, since the indices are not contiguous
self.assertIsNotNone(dset_select_all._indices)
self.assertEqual(len(dset_select_all), len(dset))
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
bad_indices = list(range(5))
bad_indices[3] = "foo" # wrong type
tmp_file = os.path.join(tmp_dir, "test.arrow")
self.assertRaises(
Exception,
dset.select,
indices=bad_indices,
indices_cache_file_name=tmp_file,
writer_batch_size=2,
)
self.assertFalse(os.path.exists(tmp_file))
dset.set_format("numpy")
with dset.select(
range(5),
indices_cache_file_name=tmp_file,
writer_batch_size=2,
) as dset_select_five:
self.assertIsNone(dset_select_five._indices)
self.assertEqual(len(dset_select_five), 5)
self.assertEqual(dset_select_five.format["type"], "numpy")
for i, row in enumerate(dset_select_five):
self.assertEqual(int(row["filename"][-1]), i)
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertDictEqual(dset_select_five.features, Features({"filename": Value("string")}))
def test_select_then_map(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
with dset.select([0]) as d1:
with d1.map(lambda x: {"id": int(x["filename"].split("_")[-1])}) as d1:
self.assertEqual(d1[0]["id"], 0)
with dset.select([1]) as d2:
with d2.map(lambda x: {"id": int(x["filename"].split("_")[-1])}) as d2:
self.assertEqual(d2[0]["id"], 1)
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
with dset.select([0], indices_cache_file_name=os.path.join(tmp_dir, "i1.arrow")) as d1:
with d1.map(lambda x: {"id": int(x["filename"].split("_")[-1])}) as d1:
self.assertEqual(d1[0]["id"], 0)
with dset.select([1], indices_cache_file_name=os.path.join(tmp_dir, "i2.arrow")) as d2:
with d2.map(lambda x: {"id": int(x["filename"].split("_")[-1])}) as d2:
self.assertEqual(d2[0]["id"], 1)
def test_pickle_after_many_transforms_on_disk(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
self.assertEqual(len(dset.cache_files), 0 if in_memory else 1)
with dset.rename_column("filename", "file") as dset:
self.assertListEqual(dset.column_names, ["file"])
with dset.select(range(5)) as dset:
self.assertEqual(len(dset), 5)
with dset.map(lambda x: {"id": int(x["file"][-1])}) as dset:
self.assertListEqual(sorted(dset.column_names), ["file", "id"])
with dset.rename_column("id", "number") as dset:
self.assertListEqual(sorted(dset.column_names), ["file", "number"])
with dset.select([1, 0]) as dset:
self.assertEqual(dset[0]["file"], "my_name-train_1")
self.assertEqual(dset[0]["number"], 1)
self.assertEqual(dset._indices["indices"].to_pylist(), [1, 0])
if not in_memory:
self.assertIn(
("rename_columns", (["file", "number"],), {}),
dset._data.replays,
)
if not in_memory:
dset._data.table = Unpicklable() # check that we don't pickle the entire table
pickled = pickle.dumps(dset)
with pickle.loads(pickled) as loaded:
self.assertEqual(loaded[0]["file"], "my_name-train_1")
self.assertEqual(loaded[0]["number"], 1)
def test_shuffle(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
tmp_file = os.path.join(tmp_dir, "test.arrow")
fingerprint = dset._fingerprint
with dset.shuffle(seed=1234, keep_in_memory=True) as dset_shuffled:
self.assertEqual(len(dset_shuffled), 30)
self.assertEqual(dset_shuffled[0]["filename"], "my_name-train_28")
self.assertEqual(dset_shuffled[2]["filename"], "my_name-train_10")
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertDictEqual(dset_shuffled.features, Features({"filename": Value("string")}))
self.assertNotEqual(dset_shuffled._fingerprint, fingerprint)
with dset.shuffle(seed=1234, indices_cache_file_name=tmp_file) as dset_shuffled:
self.assertEqual(len(dset_shuffled), 30)
self.assertEqual(dset_shuffled[0]["filename"], "my_name-train_28")
self.assertEqual(dset_shuffled[2]["filename"], "my_name-train_10")
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertDictEqual(dset_shuffled.features, Features({"filename": Value("string")}))
self.assertNotEqual(dset_shuffled._fingerprint, fingerprint)
# Reproducibility
tmp_file = os.path.join(tmp_dir, "test_2.arrow")
with dset.shuffle(seed=1234, indices_cache_file_name=tmp_file) as dset_shuffled_2:
self.assertListEqual(dset_shuffled["filename"], dset_shuffled_2["filename"])
# Compatible with temp_seed
with temp_seed(42), dset.shuffle() as d1:
with temp_seed(42), dset.shuffle() as d2, dset.shuffle() as d3:
self.assertListEqual(d1["filename"], d2["filename"])
self.assertEqual(d1._fingerprint, d2._fingerprint)
self.assertNotEqual(d3["filename"], d2["filename"])
self.assertNotEqual(d3._fingerprint, d2._fingerprint)
def test_sort(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
# Sort on a single key
with self._create_dummy_dataset(in_memory=in_memory, tmp_dir=tmp_dir) as dset:
# Keep only 10 examples
tmp_file = os.path.join(tmp_dir, "test.arrow")
with dset.select(range(10), indices_cache_file_name=tmp_file) as dset:
tmp_file = os.path.join(tmp_dir, "test_2.arrow")
with dset.shuffle(seed=1234, indices_cache_file_name=tmp_file) as dset:
self.assertEqual(len(dset), 10)
self.assertEqual(dset[0]["filename"], "my_name-train_8")
self.assertEqual(dset[1]["filename"], "my_name-train_9")
# Sort
tmp_file = os.path.join(tmp_dir, "test_3.arrow")
fingerprint = dset._fingerprint
with dset.sort("filename", indices_cache_file_name=tmp_file) as dset_sorted:
for i, row in enumerate(dset_sorted):
self.assertEqual(int(row["filename"][-1]), i)
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertDictEqual(dset_sorted.features, Features({"filename": Value("string")}))
self.assertNotEqual(dset_sorted._fingerprint, fingerprint)
# Sort reversed
tmp_file = os.path.join(tmp_dir, "test_4.arrow")
fingerprint = dset._fingerprint
with dset.sort("filename", indices_cache_file_name=tmp_file, reverse=True) as dset_sorted:
for i, row in enumerate(dset_sorted):
self.assertEqual(int(row["filename"][-1]), len(dset_sorted) - 1 - i)
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertDictEqual(dset_sorted.features, Features({"filename": Value("string")}))
self.assertNotEqual(dset_sorted._fingerprint, fingerprint)
# formatted
dset.set_format("numpy")
with dset.sort("filename") as dset_sorted_formatted:
self.assertEqual(dset_sorted_formatted.format["type"], "numpy")
# Sort on multiple keys
with self._create_dummy_dataset(in_memory=in_memory, tmp_dir=tmp_dir, multiple_columns=True) as dset:
tmp_file = os.path.join(tmp_dir, "test_5.arrow")
fingerprint = dset._fingerprint
# Throw error when reverse is a list of bools that does not match the length of column_names
with pytest.raises(ValueError):
dset.sort(["col_1", "col_2", "col_3"], reverse=[False])
with dset.shuffle(seed=1234, indices_cache_file_name=tmp_file) as dset:
# Sort
with dset.sort(["col_1", "col_2", "col_3"], reverse=[False, True, False]) as dset_sorted:
for i, row in enumerate(dset_sorted):
self.assertEqual(row["col_1"], i)
self.assertDictEqual(
dset.features,
Features(
{
"col_1": Value("int64"),
"col_2": Value("string"),
"col_3": Value("bool"),
}
),
)
self.assertDictEqual(
dset_sorted.features,
Features(
{
"col_1": Value("int64"),
"col_2": Value("string"),
"col_3": Value("bool"),
}
),
)
self.assertNotEqual(dset_sorted._fingerprint, fingerprint)
# Sort reversed
with dset.sort(["col_1", "col_2", "col_3"], reverse=[True, False, True]) as dset_sorted:
for i, row in enumerate(dset_sorted):
self.assertEqual(row["col_1"], len(dset_sorted) - 1 - i)
self.assertDictEqual(
dset.features,
Features(
{
"col_1": Value("int64"),
"col_2": Value("string"),
"col_3": Value("bool"),
}
),
)
self.assertDictEqual(
dset_sorted.features,
Features(
{
"col_1": Value("int64"),
"col_2": Value("string"),
"col_3": Value("bool"),
}
),
)
self.assertNotEqual(dset_sorted._fingerprint, fingerprint)
# formatted
dset.set_format("numpy")
with dset.sort(
["col_1", "col_2", "col_3"], reverse=[False, True, False]
) as dset_sorted_formatted:
self.assertEqual(dset_sorted_formatted.format["type"], "numpy")
@require_tf
def test_export(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
# Export the data
tfrecord_path = os.path.join(tmp_dir, "test.tfrecord")
with dset.map(
lambda ex, i: {
"id": i,
"question": f"Question {i}",
"answers": {"text": [f"Answer {i}-0", f"Answer {i}-1"], "answer_start": [0, 1]},
},
with_indices=True,
remove_columns=["filename"],
) as formatted_dset:
with formatted_dset.flatten() as formatted_dset:
formatted_dset.set_format("numpy")
formatted_dset.export(filename=tfrecord_path, format="tfrecord")
# Import the data
import tensorflow as tf
tf_dset = tf.data.TFRecordDataset([tfrecord_path])
feature_description = {
"id": tf.io.FixedLenFeature([], tf.int64),
"question": tf.io.FixedLenFeature([], tf.string),
"answers.text": tf.io.VarLenFeature(tf.string),
"answers.answer_start": tf.io.VarLenFeature(tf.int64),
}
tf_parsed_dset = tf_dset.map(
lambda example_proto: tf.io.parse_single_example(example_proto, feature_description)
)
# Test that keys match original dataset
for i, ex in enumerate(tf_parsed_dset):
self.assertEqual(ex.keys(), formatted_dset[i].keys())
# Test for equal number of elements
self.assertEqual(i, len(formatted_dset) - 1)
def test_to_csv(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
# File path argument
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
file_path = os.path.join(tmp_dir, "test_path.csv")
bytes_written = dset.to_csv(path_or_buf=file_path)
self.assertTrue(os.path.isfile(file_path))
self.assertEqual(bytes_written, os.path.getsize(file_path))
csv_dset = pd.read_csv(file_path)
self.assertEqual(csv_dset.shape, dset.shape)
self.assertListEqual(list(csv_dset.columns), list(dset.column_names))
# File buffer argument
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
file_path = os.path.join(tmp_dir, "test_buffer.csv")
with open(file_path, "wb+") as buffer:
bytes_written = dset.to_csv(path_or_buf=buffer)
self.assertTrue(os.path.isfile(file_path))
self.assertEqual(bytes_written, os.path.getsize(file_path))
csv_dset = pd.read_csv(file_path)
self.assertEqual(csv_dset.shape, dset.shape)
self.assertListEqual(list(csv_dset.columns), list(dset.column_names))
# After a select/shuffle transform
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
dset = dset.select(range(0, len(dset), 2)).shuffle()
file_path = os.path.join(tmp_dir, "test_path.csv")
bytes_written = dset.to_csv(path_or_buf=file_path)
self.assertTrue(os.path.isfile(file_path))
self.assertEqual(bytes_written, os.path.getsize(file_path))
csv_dset = pd.read_csv(file_path)
self.assertEqual(csv_dset.shape, dset.shape)
self.assertListEqual(list(csv_dset.columns), list(dset.column_names))
# With array features
with self._create_dummy_dataset(in_memory, tmp_dir, array_features=True) as dset:
file_path = os.path.join(tmp_dir, "test_path.csv")
bytes_written = dset.to_csv(path_or_buf=file_path)
self.assertTrue(os.path.isfile(file_path))
self.assertEqual(bytes_written, os.path.getsize(file_path))
csv_dset = pd.read_csv(file_path)
self.assertEqual(csv_dset.shape, dset.shape)
self.assertListEqual(list(csv_dset.columns), list(dset.column_names))
def test_to_dict(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
# Full
dset_to_dict = dset.to_dict()
self.assertIsInstance(dset_to_dict, dict)
self.assertListEqual(sorted(dset_to_dict.keys()), sorted(dset.column_names))
for col_name in dset.column_names:
self.assertLessEqual(len(dset_to_dict[col_name]), len(dset))
# With index mapping
with dset.select([1, 0, 3]) as dset:
dset_to_dict = dset.to_dict()
self.assertIsInstance(dset_to_dict, dict)
self.assertEqual(len(dset_to_dict), 3)
self.assertListEqual(sorted(dset_to_dict.keys()), sorted(dset.column_names))
for col_name in dset.column_names:
self.assertIsInstance(dset_to_dict[col_name], list)
self.assertEqual(len(dset_to_dict[col_name]), len(dset))
def test_to_list(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
dset_to_list = dset.to_list()
self.assertIsInstance(dset_to_list, list)
for row in dset_to_list:
self.assertIsInstance(row, dict)
self.assertListEqual(sorted(row.keys()), sorted(dset.column_names))
# With index mapping
with dset.select([1, 0, 3]) as dset:
dset_to_list = dset.to_list()
self.assertIsInstance(dset_to_list, list)
self.assertEqual(len(dset_to_list), 3)
for row in dset_to_list:
self.assertIsInstance(row, dict)
self.assertListEqual(sorted(row.keys()), sorted(dset.column_names))
def test_to_pandas(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
# Batched
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
batch_size = dset.num_rows - 1
to_pandas_generator = dset.to_pandas(batched=True, batch_size=batch_size)
for batch in to_pandas_generator:
self.assertIsInstance(batch, pd.DataFrame)
self.assertListEqual(sorted(batch.columns), sorted(dset.column_names))
for col_name in dset.column_names:
self.assertLessEqual(len(batch[col_name]), batch_size)
# Full
dset_to_pandas = dset.to_pandas()
self.assertIsInstance(dset_to_pandas, pd.DataFrame)
self.assertListEqual(sorted(dset_to_pandas.columns), sorted(dset.column_names))
for col_name in dset.column_names:
self.assertEqual(len(dset_to_pandas[col_name]), len(dset))
# With index mapping
with dset.select([1, 0, 3]) as dset:
dset_to_pandas = dset.to_pandas()
self.assertIsInstance(dset_to_pandas, pd.DataFrame)
self.assertEqual(len(dset_to_pandas), 3)
self.assertListEqual(sorted(dset_to_pandas.columns), sorted(dset.column_names))
for col_name in dset.column_names:
self.assertEqual(len(dset_to_pandas[col_name]), dset.num_rows)
def test_to_parquet(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
# File path argument
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
file_path = os.path.join(tmp_dir, "test_path.parquet")
dset.to_parquet(path_or_buf=file_path)
self.assertTrue(os.path.isfile(file_path))
# self.assertEqual(bytes_written, os.path.getsize(file_path)) # because of compression, the number of bytes doesn't match
parquet_dset = pd.read_parquet(file_path)
self.assertEqual(parquet_dset.shape, dset.shape)
self.assertListEqual(list(parquet_dset.columns), list(dset.column_names))
# File buffer argument
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
file_path = os.path.join(tmp_dir, "test_buffer.parquet")
with open(file_path, "wb+") as buffer:
dset.to_parquet(path_or_buf=buffer)
self.assertTrue(os.path.isfile(file_path))
# self.assertEqual(bytes_written, os.path.getsize(file_path)) # because of compression, the number of bytes doesn't match
parquet_dset = pd.read_parquet(file_path)
self.assertEqual(parquet_dset.shape, dset.shape)
self.assertListEqual(list(parquet_dset.columns), list(dset.column_names))
# After a select/shuffle transform
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
dset = dset.select(range(0, len(dset), 2)).shuffle()
file_path = os.path.join(tmp_dir, "test_path.parquet")
dset.to_parquet(path_or_buf=file_path)
self.assertTrue(os.path.isfile(file_path))
# self.assertEqual(bytes_written, os.path.getsize(file_path)) # because of compression, the number of bytes doesn't match
parquet_dset = pd.read_parquet(file_path)
self.assertEqual(parquet_dset.shape, dset.shape)
self.assertListEqual(list(parquet_dset.columns), list(dset.column_names))
# With array features
with self._create_dummy_dataset(in_memory, tmp_dir, array_features=True) as dset:
file_path = os.path.join(tmp_dir, "test_path.parquet")
dset.to_parquet(path_or_buf=file_path)
self.assertTrue(os.path.isfile(file_path))
# self.assertEqual(bytes_written, os.path.getsize(file_path)) # because of compression, the number of bytes doesn't match
parquet_dset = pd.read_parquet(file_path)
self.assertEqual(parquet_dset.shape, dset.shape)
self.assertListEqual(list(parquet_dset.columns), list(dset.column_names))
@require_sqlalchemy
def test_to_sql(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
# Destionation specified as database URI string
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
file_path = os.path.join(tmp_dir, "test_path.sqlite")
_ = dset.to_sql("data", "sqlite:///" + file_path)
self.assertTrue(os.path.isfile(file_path))
sql_dset = pd.read_sql("data", "sqlite:///" + file_path)
self.assertEqual(sql_dset.shape, dset.shape)
self.assertListEqual(list(sql_dset.columns), list(dset.column_names))
# Destionation specified as sqlite3 connection
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
import sqlite3
file_path = os.path.join(tmp_dir, "test_path.sqlite")
with contextlib.closing(sqlite3.connect(file_path)) as con:
_ = dset.to_sql("data", con, if_exists="replace")
self.assertTrue(os.path.isfile(file_path))
sql_dset = pd.read_sql("data", "sqlite:///" + file_path)
self.assertEqual(sql_dset.shape, dset.shape)
self.assertListEqual(list(sql_dset.columns), list(dset.column_names))
# Test writing to a database in chunks
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
file_path = os.path.join(tmp_dir, "test_path.sqlite")
_ = dset.to_sql("data", "sqlite:///" + file_path, batch_size=1, if_exists="replace")
self.assertTrue(os.path.isfile(file_path))
sql_dset = pd.read_sql("data", "sqlite:///" + file_path)
self.assertEqual(sql_dset.shape, dset.shape)
self.assertListEqual(list(sql_dset.columns), list(dset.column_names))
# After a select/shuffle transform
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
dset = dset.select(range(0, len(dset), 2)).shuffle()
file_path = os.path.join(tmp_dir, "test_path.sqlite")
_ = dset.to_sql("data", "sqlite:///" + file_path, if_exists="replace")
self.assertTrue(os.path.isfile(file_path))
sql_dset = pd.read_sql("data", "sqlite:///" + file_path)
self.assertEqual(sql_dset.shape, dset.shape)
self.assertListEqual(list(sql_dset.columns), list(dset.column_names))
# With array features
with self._create_dummy_dataset(in_memory, tmp_dir, array_features=True) as dset:
file_path = os.path.join(tmp_dir, "test_path.sqlite")
_ = dset.to_sql("data", "sqlite:///" + file_path, if_exists="replace")
self.assertTrue(os.path.isfile(file_path))
sql_dset = pd.read_sql("data", "sqlite:///" + file_path)
self.assertEqual(sql_dset.shape, dset.shape)
self.assertListEqual(list(sql_dset.columns), list(dset.column_names))
def test_train_test_split(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
fingerprint = dset._fingerprint
dset_dict = dset.train_test_split(test_size=10, shuffle=False)
self.assertListEqual(list(dset_dict.keys()), ["train", "test"])
dset_train = dset_dict["train"]
dset_test = dset_dict["test"]
self.assertEqual(len(dset_train), 20)
self.assertEqual(len(dset_test), 10)
self.assertEqual(dset_train[0]["filename"], "my_name-train_0")
self.assertEqual(dset_train[-1]["filename"], "my_name-train_19")
self.assertEqual(dset_test[0]["filename"], "my_name-train_20")
self.assertEqual(dset_test[-1]["filename"], "my_name-train_29")
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertDictEqual(dset_train.features, Features({"filename": Value("string")}))
self.assertDictEqual(dset_test.features, Features({"filename": Value("string")}))
self.assertNotEqual(dset_train._fingerprint, fingerprint)
self.assertNotEqual(dset_test._fingerprint, fingerprint)
self.assertNotEqual(dset_train._fingerprint, dset_test._fingerprint)
dset_dict = dset.train_test_split(test_size=0.5, shuffle=False)
self.assertListEqual(list(dset_dict.keys()), ["train", "test"])
dset_train = dset_dict["train"]
dset_test = dset_dict["test"]
self.assertEqual(len(dset_train), 15)
self.assertEqual(len(dset_test), 15)
self.assertEqual(dset_train[0]["filename"], "my_name-train_0")
self.assertEqual(dset_train[-1]["filename"], "my_name-train_14")
self.assertEqual(dset_test[0]["filename"], "my_name-train_15")
self.assertEqual(dset_test[-1]["filename"], "my_name-train_29")
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertDictEqual(dset_train.features, Features({"filename": Value("string")}))
self.assertDictEqual(dset_test.features, Features({"filename": Value("string")}))
dset_dict = dset.train_test_split(train_size=10, shuffle=False)
self.assertListEqual(list(dset_dict.keys()), ["train", "test"])
dset_train = dset_dict["train"]
dset_test = dset_dict["test"]
self.assertEqual(len(dset_train), 10)
self.assertEqual(len(dset_test), 20)
self.assertEqual(dset_train[0]["filename"], "my_name-train_0")
self.assertEqual(dset_train[-1]["filename"], "my_name-train_9")
self.assertEqual(dset_test[0]["filename"], "my_name-train_10")
self.assertEqual(dset_test[-1]["filename"], "my_name-train_29")
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertDictEqual(dset_train.features, Features({"filename": Value("string")}))
self.assertDictEqual(dset_test.features, Features({"filename": Value("string")}))
dset.set_format("numpy")
dset_dict = dset.train_test_split(train_size=10, seed=42)
self.assertListEqual(list(dset_dict.keys()), ["train", "test"])
dset_train = dset_dict["train"]
dset_test = dset_dict["test"]
self.assertEqual(len(dset_train), 10)
self.assertEqual(len(dset_test), 20)
self.assertEqual(dset_train.format["type"], "numpy")
self.assertEqual(dset_test.format["type"], "numpy")
self.assertNotEqual(dset_train[0]["filename"].item(), "my_name-train_0")
self.assertNotEqual(dset_train[-1]["filename"].item(), "my_name-train_9")
self.assertNotEqual(dset_test[0]["filename"].item(), "my_name-train_10")
self.assertNotEqual(dset_test[-1]["filename"].item(), "my_name-train_29")
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertDictEqual(dset_train.features, Features({"filename": Value("string")}))
self.assertDictEqual(dset_test.features, Features({"filename": Value("string")}))
del dset_test, dset_train, dset_dict # DatasetDict
def test_shard(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir, self._create_dummy_dataset(in_memory, tmp_dir) as dset:
tmp_file = os.path.join(tmp_dir, "test.arrow")
with dset.select(range(10), indices_cache_file_name=tmp_file) as dset:
self.assertEqual(len(dset), 10)
# Shard
tmp_file_1 = os.path.join(tmp_dir, "test_1.arrow")
fingerprint = dset._fingerprint
with dset.shard(num_shards=8, index=1, indices_cache_file_name=tmp_file_1) as dset_sharded:
self.assertEqual(2, len(dset_sharded))
self.assertEqual(["my_name-train_1", "my_name-train_9"], dset_sharded["filename"])
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertDictEqual(dset_sharded.features, Features({"filename": Value("string")}))
self.assertNotEqual(dset_sharded._fingerprint, fingerprint)
# Shard contiguous
tmp_file_2 = os.path.join(tmp_dir, "test_2.arrow")
with dset.shard(
num_shards=3, index=0, contiguous=True, indices_cache_file_name=tmp_file_2
) as dset_sharded_contiguous:
self.assertEqual([f"my_name-train_{i}" for i in (0, 1, 2, 3)], dset_sharded_contiguous["filename"])
self.assertDictEqual(dset.features, Features({"filename": Value("string")}))
self.assertDictEqual(dset_sharded_contiguous.features, Features({"filename": Value("string")}))
# Test lengths of sharded contiguous
self.assertEqual(
[4, 3, 3],
[
len(dset.shard(3, index=i, contiguous=True, indices_cache_file_name=tmp_file_2 + str(i)))
for i in range(3)
],
)
# formatted
dset.set_format("numpy")
with dset.shard(num_shards=3, index=0) as dset_sharded_formatted:
self.assertEqual(dset_sharded_formatted.format["type"], "numpy")
def test_flatten_indices(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
self.assertIsNone(dset._indices)
tmp_file = os.path.join(tmp_dir, "test.arrow")
with dset.select(range(0, 10, 2), indices_cache_file_name=tmp_file) as dset:
self.assertEqual(len(dset), 5)
self.assertIsNotNone(dset._indices)
tmp_file_2 = os.path.join(tmp_dir, "test_2.arrow")
fingerprint = dset._fingerprint
dset.set_format("numpy")
with dset.flatten_indices(cache_file_name=tmp_file_2) as dset:
self.assertEqual(len(dset), 5)
self.assertEqual(len(dset.data), len(dset))
self.assertIsNone(dset._indices)
self.assertNotEqual(dset._fingerprint, fingerprint)
self.assertEqual(dset.format["type"], "numpy")
# Test unique works
dset.unique(dset.column_names[0])
assert_arrow_metadata_are_synced_with_dataset_features(dset)
# Empty indices mapping
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir) as dset:
self.assertIsNone(dset._indices, None)
tmp_file = os.path.join(tmp_dir, "test.arrow")
with dset.filter(lambda _: False, cache_file_name=tmp_file) as dset:
self.assertEqual(len(dset), 0)
self.assertIsNotNone(dset._indices, None)
tmp_file_2 = os.path.join(tmp_dir, "test_2.arrow")
fingerprint = dset._fingerprint
dset.set_format("numpy")
with dset.flatten_indices(cache_file_name=tmp_file_2) as dset:
self.assertEqual(len(dset), 0)
self.assertEqual(len(dset.data), len(dset))
self.assertIsNone(dset._indices, None)
self.assertNotEqual(dset._fingerprint, fingerprint)
self.assertEqual(dset.format["type"], "numpy")
# Test unique works
dset.unique(dset.column_names[0])
assert_arrow_metadata_are_synced_with_dataset_features(dset)
@require_tf
@require_torch
def test_format_vectors(self, in_memory):
import numpy as np
import tensorflow as tf
import torch
with tempfile.TemporaryDirectory() as tmp_dir, self._create_dummy_dataset(
in_memory, tmp_dir
) as dset, dset.map(lambda ex, i: {"vec": np.ones(3) * i}, with_indices=True) as dset:
columns = dset.column_names
self.assertIsNotNone(dset[0])
self.assertIsNotNone(dset[:2])
for col in columns:
self.assertIsInstance(dset[0][col], (str, list))
self.assertIsInstance(dset[:2][col], list)
self.assertDictEqual(
dset.features, Features({"filename": Value("string"), "vec": Sequence(Value("float64"))})
)
dset.set_format("tensorflow")
self.assertIsNotNone(dset[0])
self.assertIsNotNone(dset[:2])
for col in columns:
self.assertIsInstance(dset[0][col], (tf.Tensor, tf.RaggedTensor))
self.assertIsInstance(dset[:2][col], (tf.Tensor, tf.RaggedTensor))
self.assertIsInstance(dset[col], (tf.Tensor, tf.RaggedTensor))
self.assertTupleEqual(tuple(dset[:2]["vec"].shape), (2, 3))
self.assertTupleEqual(tuple(dset["vec"][:2].shape), (2, 3))
dset.set_format("numpy")
self.assertIsNotNone(dset[0])
self.assertIsNotNone(dset[:2])
self.assertIsInstance(dset[0]["filename"], np.str_)
self.assertIsInstance(dset[:2]["filename"], np.ndarray)
self.assertIsInstance(dset["filename"], np.ndarray)
self.assertIsInstance(dset[0]["vec"], np.ndarray)
self.assertIsInstance(dset[:2]["vec"], np.ndarray)
self.assertIsInstance(dset["vec"], np.ndarray)
self.assertTupleEqual(dset[:2]["vec"].shape, (2, 3))
self.assertTupleEqual(dset["vec"][:2].shape, (2, 3))
dset.set_format("torch", columns=["vec"])
self.assertIsNotNone(dset[0])
self.assertIsNotNone(dset[:2])
# torch.Tensor is only for numerical columns
self.assertIsInstance(dset[0]["vec"], torch.Tensor)
self.assertIsInstance(dset[:2]["vec"], torch.Tensor)
self.assertIsInstance(dset["vec"][:2], torch.Tensor)
self.assertTupleEqual(dset[:2]["vec"].shape, (2, 3))
self.assertTupleEqual(dset["vec"][:2].shape, (2, 3))
@require_tf
@require_torch
def test_format_ragged_vectors(self, in_memory):
import numpy as np
import tensorflow as tf
import torch
with tempfile.TemporaryDirectory() as tmp_dir, self._create_dummy_dataset(
in_memory, tmp_dir
) as dset, dset.map(lambda ex, i: {"vec": np.ones(3 + i) * i}, with_indices=True) as dset:
columns = dset.column_names
self.assertIsNotNone(dset[0])
self.assertIsNotNone(dset[:2])
for col in columns:
self.assertIsInstance(dset[0][col], (str, list))
self.assertIsInstance(dset[:2][col], list)
self.assertDictEqual(
dset.features, Features({"filename": Value("string"), "vec": Sequence(Value("float64"))})
)
dset.set_format("tensorflow")
self.assertIsNotNone(dset[0])
self.assertIsNotNone(dset[:2])
for col in columns:
self.assertIsInstance(dset[0][col], tf.Tensor)
self.assertIsInstance(dset[:2][col], tf.RaggedTensor if col == "vec" else tf.Tensor)
self.assertIsInstance(dset[col], tf.RaggedTensor if col == "vec" else tf.Tensor)
# dim is None for ragged vectors in tensorflow
self.assertListEqual(dset[:2]["vec"].shape.as_list(), [2, None])
self.assertListEqual(dset["vec"][:2].shape.as_list(), [2, None])
dset.set_format("numpy")
self.assertIsNotNone(dset[0])
self.assertIsNotNone(dset[:2])
self.assertIsInstance(dset[0]["filename"], np.str_)
self.assertIsInstance(dset[:2]["filename"], np.ndarray)
self.assertIsInstance(dset["filename"], np.ndarray)
self.assertIsInstance(dset[0]["vec"], np.ndarray)
self.assertIsInstance(dset[:2]["vec"], np.ndarray)
self.assertIsInstance(dset["vec"], np.ndarray)
# array is flat for ragged vectors in numpy
self.assertTupleEqual(dset[:2]["vec"].shape, (2,))
self.assertTupleEqual(dset["vec"][:2].shape, (2,))
dset.set_format("torch")
self.assertIsNotNone(dset[0])
self.assertIsNotNone(dset[:2])
self.assertIsInstance(dset[0]["filename"], str)
self.assertIsInstance(dset[:2]["filename"], list)
self.assertIsInstance(dset["filename"], list)
self.assertIsInstance(dset[0]["vec"], torch.Tensor)
self.assertIsInstance(dset[:2]["vec"][0], torch.Tensor)
self.assertIsInstance(dset["vec"][0], torch.Tensor)
# pytorch doesn't support ragged tensors, so we should have lists
self.assertIsInstance(dset[:2]["vec"], list)
self.assertIsInstance(dset[:2]["vec"][0], torch.Tensor)
self.assertIsInstance(dset["vec"][:2], list)
self.assertIsInstance(dset["vec"][0], torch.Tensor)
@require_tf
@require_torch
def test_format_nested(self, in_memory):
import numpy as np
import tensorflow as tf
import torch
with tempfile.TemporaryDirectory() as tmp_dir, self._create_dummy_dataset(
in_memory, tmp_dir
) as dset, dset.map(lambda ex: {"nested": [{"foo": np.ones(3)}] * len(ex["filename"])}, batched=True) as dset:
self.assertDictEqual(
dset.features, Features({"filename": Value("string"), "nested": {"foo": Sequence(Value("float64"))}})
)
dset.set_format("tensorflow")
self.assertIsNotNone(dset[0])
self.assertIsInstance(dset[0]["nested"]["foo"], (tf.Tensor, tf.RaggedTensor))
self.assertIsNotNone(dset[:2])
self.assertIsInstance(dset[:2]["nested"][0]["foo"], (tf.Tensor, tf.RaggedTensor))
self.assertIsInstance(dset["nested"][0]["foo"], (tf.Tensor, tf.RaggedTensor))
dset.set_format("numpy")
self.assertIsNotNone(dset[0])
self.assertIsInstance(dset[0]["nested"]["foo"], np.ndarray)
self.assertIsNotNone(dset[:2])
self.assertIsInstance(dset[:2]["nested"][0]["foo"], np.ndarray)
self.assertIsInstance(dset["nested"][0]["foo"], np.ndarray)
dset.set_format("torch", columns="nested")
self.assertIsNotNone(dset[0])
self.assertIsInstance(dset[0]["nested"]["foo"], torch.Tensor)
self.assertIsNotNone(dset[:2])
self.assertIsInstance(dset[:2]["nested"][0]["foo"], torch.Tensor)
self.assertIsInstance(dset["nested"][0]["foo"], torch.Tensor)
def test_format_pandas(self, in_memory):
import pandas as pd
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
dset.set_format("pandas")
self.assertIsInstance(dset[0], pd.DataFrame)
self.assertIsInstance(dset[:2], pd.DataFrame)
self.assertIsInstance(dset["col_1"], pd.Series)
def test_transmit_format_single(self, in_memory):
@transmit_format
def my_single_transform(self, return_factory, *args, **kwargs):
return return_factory()
with tempfile.TemporaryDirectory() as tmp_dir:
return_factory = partial(
self._create_dummy_dataset, in_memory=in_memory, tmp_dir=tmp_dir, multiple_columns=True
)
with return_factory() as dset:
dset.set_format("numpy", columns=["col_1"])
prev_format = dset.format
with my_single_transform(dset, return_factory) as transformed_dset:
self.assertDictEqual(transformed_dset.format, prev_format)
def test_transmit_format_dict(self, in_memory):
@transmit_format
def my_split_transform(self, return_factory, *args, **kwargs):
return DatasetDict({"train": return_factory()})
with tempfile.TemporaryDirectory() as tmp_dir:
return_factory = partial(
self._create_dummy_dataset, in_memory=in_memory, tmp_dir=tmp_dir, multiple_columns=True
)
with return_factory() as dset:
dset.set_format("numpy", columns=["col_1"])
prev_format = dset.format
transformed_dset = my_split_transform(dset, return_factory)["train"]
self.assertDictEqual(transformed_dset.format, prev_format)
del transformed_dset # DatasetDict
def test_with_format(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
with dset.with_format("numpy", columns=["col_1"]) as dset2:
dset.set_format("numpy", columns=["col_1"])
self.assertDictEqual(dset.format, dset2.format)
self.assertEqual(dset._fingerprint, dset2._fingerprint)
# dset.reset_format()
# self.assertNotEqual(dset.format, dset2.format)
# self.assertNotEqual(dset._fingerprint, dset2._fingerprint)
def test_with_transform(self, in_memory):
with tempfile.TemporaryDirectory() as tmp_dir:
with self._create_dummy_dataset(in_memory, tmp_dir, multiple_columns=True) as dset:
transform = lambda x: {"foo": x["col_1"]} # noqa: E731
with dset.with_transform(transform, columns=["col_1"]) as dset2:
dset.set_transform(transform, columns=["col_1"])
self.assertDictEqual(dset.format, dset2.format)
self.assertEqual(dset._fingerprint, dset2._fingerprint)
dset.reset_format()
self.assertNotEqual(dset.format, dset2.format)
self.assertNotEqual(dset._fingerprint, dset2._fingerprint)
@require_tf
def test_tf_dataset_conversion(self, in_memory):
tmp_dir = tempfile.TemporaryDirectory()
for num_workers in [0, 1, 2]:
if num_workers > 0 and sys.platform == "win32" and not in_memory:
continue # This test hangs on the Py3.10 test worker, but it runs fine locally on my Windows machine
with self._create_dummy_dataset(in_memory, tmp_dir.name, array_features=True) as dset:
tf_dataset = dset.to_tf_dataset(columns="col_3", batch_size=2, num_workers=num_workers)
batch = next(iter(tf_dataset))
self.assertEqual(batch.shape.as_list(), [2, 4])
self.assertEqual(batch.dtype.name, "int64")
with self._create_dummy_dataset(in_memory, tmp_dir.name, multiple_columns=True) as dset:
tf_dataset = dset.to_tf_dataset(columns="col_1", batch_size=2, num_workers=num_workers)
batch = next(iter(tf_dataset))
self.assertEqual(batch.shape.as_list(), [2])
self.assertEqual(batch.dtype.name, "int64")
with self._create_dummy_dataset(in_memory, tmp_dir.name, multiple_columns=True) as dset:
# Check that it works with all default options (except batch_size because the dummy dataset only has 4)
tf_dataset = dset.to_tf_dataset(batch_size=2, num_workers=num_workers)
batch = next(iter(tf_dataset))
self.assertEqual(batch["col_1"].shape.as_list(), [2])
self.assertEqual(batch["col_2"].shape.as_list(), [2])
self.assertEqual(batch["col_1"].dtype.name, "int64")
self.assertEqual(batch["col_2"].dtype.name, "string") # Assert that we're converting strings properly
with self._create_dummy_dataset(in_memory, tmp_dir.name, multiple_columns=True) as dset:
# Check that when we use a transform that creates a new column from existing column values
# but don't load the old columns that the new column depends on in the final dataset,
# that they're still kept around long enough to be used in the transform
transform_dset = dset.with_transform(
lambda x: {"new_col": [val * 2 for val in x["col_1"]], "col_1": x["col_1"]}
)
tf_dataset = transform_dset.to_tf_dataset(columns="new_col", batch_size=2, num_workers=num_workers)
batch = next(iter(tf_dataset))
self.assertEqual(batch.shape.as_list(), [2])
self.assertEqual(batch.dtype.name, "int64")
del transform_dset
del tf_dataset # For correct cleanup
@require_tf
def test_tf_index_reshuffling(self, in_memory):
# This test checks that when we do two epochs over a tf.data.Dataset from to_tf_dataset
# that we get a different shuffle order each time
# It also checks that when we aren't shuffling, that the dataset order is fully preserved
# even when loading is split across multiple workers
data = {"col_1": list(range(20))}
for num_workers in [0, 1, 2, 3]:
with Dataset.from_dict(data) as dset:
tf_dataset = dset.to_tf_dataset(batch_size=10, shuffle=True, num_workers=num_workers)
indices = []
for batch in tf_dataset:
indices.append(batch["col_1"])
indices = np.concatenate([arr.numpy() for arr in indices])
second_indices = []
for batch in tf_dataset:
second_indices.append(batch["col_1"])
second_indices = np.concatenate([arr.numpy() for arr in second_indices])
self.assertFalse(np.array_equal(indices, second_indices))
self.assertEqual(len(indices), len(np.unique(indices)))
self.assertEqual(len(second_indices), len(np.unique(second_indices)))
tf_dataset = dset.to_tf_dataset(batch_size=1, shuffle=False, num_workers=num_workers)
for i, batch in enumerate(tf_dataset):
# Assert that the unshuffled order is fully preserved even when multiprocessing
self.assertEqual(i, batch["col_1"].numpy())
@require_tf
def test_tf_label_renaming(self, in_memory):
# Protect TF-specific imports in here
import tensorflow as tf
from datasets.utils.tf_utils import minimal_tf_collate_fn_with_renaming
tmp_dir = tempfile.TemporaryDirectory()
with self._create_dummy_dataset(in_memory, tmp_dir.name, multiple_columns=True) as dset:
with dset.rename_columns({"col_1": "features", "col_2": "label"}) as new_dset:
tf_dataset = new_dset.to_tf_dataset(collate_fn=minimal_tf_collate_fn_with_renaming, batch_size=4)
batch = next(iter(tf_dataset))
self.assertTrue("labels" in batch and "features" in batch)
tf_dataset = new_dset.to_tf_dataset(
columns=["features", "labels"], collate_fn=minimal_tf_collate_fn_with_renaming, batch_size=4
)
batch = next(iter(tf_dataset))
self.assertTrue("labels" in batch and "features" in batch)
tf_dataset = new_dset.to_tf_dataset(
columns=["features", "label"], collate_fn=minimal_tf_collate_fn_with_renaming, batch_size=4
)
batch = next(iter(tf_dataset))
self.assertTrue("labels" in batch and "features" in batch) # Assert renaming was handled correctly
tf_dataset = new_dset.to_tf_dataset(
columns=["features"],
label_cols=["labels"],
collate_fn=minimal_tf_collate_fn_with_renaming,
batch_size=4,
)
batch = next(iter(tf_dataset))
self.assertEqual(len(batch), 2)
# Assert that we don't have any empty entries here
self.assertTrue(isinstance(batch[0], tf.Tensor) and isinstance(batch[1], tf.Tensor))
tf_dataset = new_dset.to_tf_dataset(
columns=["features"],
label_cols=["label"],
collate_fn=minimal_tf_collate_fn_with_renaming,
batch_size=4,
)
batch = next(iter(tf_dataset))
self.assertEqual(len(batch), 2)
# Assert that we don't have any empty entries here
self.assertTrue(isinstance(batch[0], tf.Tensor) and isinstance(batch[1], tf.Tensor))
tf_dataset = new_dset.to_tf_dataset(
columns=["features"],
collate_fn=minimal_tf_collate_fn_with_renaming,
batch_size=4,
)
batch = next(iter(tf_dataset))
# Assert that labels didn't creep in when we don't ask for them
# just because the collate_fn added them
self.assertTrue(isinstance(batch, tf.Tensor))
del tf_dataset # For correct cleanup
@require_tf
def test_tf_dataset_options(self, in_memory):
tmp_dir = tempfile.TemporaryDirectory()
# Test that batch_size option works as expected
with self._create_dummy_dataset(in_memory, tmp_dir.name, array_features=True) as dset:
tf_dataset = dset.to_tf_dataset(columns="col_3", batch_size=2)
batch = next(iter(tf_dataset))
self.assertEqual(batch.shape.as_list(), [2, 4])
self.assertEqual(batch.dtype.name, "int64")
# Test that batch_size=None (optional) works as expected
with self._create_dummy_dataset(in_memory, tmp_dir.name, multiple_columns=True) as dset:
tf_dataset = dset.to_tf_dataset(columns="col_3", batch_size=None)
single_example = next(iter(tf_dataset))
self.assertEqual(single_example.shape.as_list(), [])
self.assertEqual(single_example.dtype.name, "int64")
# Assert that we can batch it with `tf.data.Dataset.batch` method
batched_dataset = tf_dataset.batch(batch_size=2)
batch = next(iter(batched_dataset))
self.assertEqual(batch.shape.as_list(), [2])
self.assertEqual(batch.dtype.name, "int64")
# Test that batching a batch_size=None dataset produces the same results as using batch_size arg
with self._create_dummy_dataset(in_memory, tmp_dir.name, multiple_columns=True) as dset:
batch_size = 2
tf_dataset_no_batch = dset.to_tf_dataset(columns="col_3")
tf_dataset_batch = dset.to_tf_dataset(columns="col_3", batch_size=batch_size)
self.assertEqual(tf_dataset_no_batch.element_spec, tf_dataset_batch.unbatch().element_spec)
self.assertEqual(tf_dataset_no_batch.cardinality(), tf_dataset_batch.cardinality() * batch_size)
for batch_1, batch_2 in zip(tf_dataset_no_batch.batch(batch_size=batch_size), tf_dataset_batch):
self.assertEqual(batch_1.shape, batch_2.shape)
self.assertEqual(batch_1.dtype, batch_2.dtype)
self.assertListEqual(batch_1.numpy().tolist(), batch_2.numpy().tolist())
# Test that requesting label_cols works as expected
with self._create_dummy_dataset(in_memory, tmp_dir.name, multiple_columns=True) as dset:
tf_dataset = dset.to_tf_dataset(columns="col_1", label_cols=["col_2", "col_3"], batch_size=4)
batch = next(iter(tf_dataset))
self.assertEqual(len(batch), 2)
self.assertEqual(set(batch[1].keys()), {"col_2", "col_3"})
self.assertEqual(batch[0].dtype.name, "int64")
# Assert data comes out as expected and isn't shuffled
self.assertEqual(batch[0].numpy().tolist(), [3, 2, 1, 0])
self.assertEqual(batch[1]["col_2"].numpy().tolist(), [b"a", b"b", b"c", b"d"])
self.assertEqual(batch[1]["col_3"].numpy().tolist(), [0, 1, 0, 1])
# Check that incomplete batches are dropped if requested
with self._create_dummy_dataset(in_memory, tmp_dir.name, multiple_columns=True) as dset:
tf_dataset = dset.to_tf_dataset(columns="col_1", batch_size=3)
tf_dataset_with_drop = dset.to_tf_dataset(columns="col_1", batch_size=3, drop_remainder=True)
self.assertEqual(len(tf_dataset), 2) # One batch of 3 and one batch of 1
self.assertEqual(len(tf_dataset_with_drop), 1) # Incomplete batch of 1 is dropped
# Test that `NotImplementedError` is raised `batch_size` is None and `num_workers` is > 0
if sys.version_info >= (3, 8):
with self._create_dummy_dataset(in_memory, tmp_dir.name, multiple_columns=True) as dset:
with self.assertRaisesRegex(
NotImplementedError, "`batch_size` must be specified when using multiple workers"
):
dset.to_tf_dataset(columns="col_1", batch_size=None, num_workers=2)
del tf_dataset # For correct cleanup
del tf_dataset_with_drop
class MiscellaneousDatasetTest(TestCase):
def test_from_pandas(self):
data = {"col_1": [3, 2, 1, 0], "col_2": ["a", "b", "c", "d"]}
df = pd.DataFrame.from_dict(data)
with Dataset.from_pandas(df) as dset:
self.assertListEqual(dset["col_1"], data["col_1"])
self.assertListEqual(dset["col_2"], data["col_2"])
self.assertListEqual(list(dset.features.keys()), ["col_1", "col_2"])
self.assertDictEqual(dset.features, Features({"col_1": Value("int64"), "col_2": Value("string")}))
features = Features({"col_1": Value("int64"), "col_2": Value("string")})
with Dataset.from_pandas(df, features=features) as dset:
self.assertListEqual(dset["col_1"], data["col_1"])
self.assertListEqual(dset["col_2"], data["col_2"])
self.assertListEqual(list(dset.features.keys()), ["col_1", "col_2"])
self.assertDictEqual(dset.features, Features({"col_1": Value("int64"), "col_2": Value("string")}))
features = Features({"col_1": Value("int64"), "col_2": Value("string")})
with Dataset.from_pandas(df, features=features, info=DatasetInfo(features=features)) as dset:
self.assertListEqual(dset["col_1"], data["col_1"])
self.assertListEqual(dset["col_2"], data["col_2"])
self.assertListEqual(list(dset.features.keys()), ["col_1", "col_2"])
self.assertDictEqual(dset.features, Features({"col_1": Value("int64"), "col_2": Value("string")}))
features = Features({"col_1": Sequence(Value("string")), "col_2": Value("string")})
self.assertRaises(TypeError, Dataset.from_pandas, df, features=features)
def test_from_dict(self):
data = {"col_1": [3, 2, 1, 0], "col_2": ["a", "b", "c", "d"], "col_3": pa.array([True, False, True, False])}
with Dataset.from_dict(data) as dset:
self.assertListEqual(dset["col_1"], data["col_1"])
self.assertListEqual(dset["col_2"], data["col_2"])
self.assertListEqual(dset["col_3"], data["col_3"].to_pylist())
self.assertListEqual(list(dset.features.keys()), ["col_1", "col_2", "col_3"])
self.assertDictEqual(
dset.features, Features({"col_1": Value("int64"), "col_2": Value("string"), "col_3": Value("bool")})
)
features = Features({"col_1": Value("int64"), "col_2": Value("string"), "col_3": Value("bool")})
with Dataset.from_dict(data, features=features) as dset:
self.assertListEqual(dset["col_1"], data["col_1"])
self.assertListEqual(dset["col_2"], data["col_2"])
self.assertListEqual(dset["col_3"], data["col_3"].to_pylist())
self.assertListEqual(list(dset.features.keys()), ["col_1", "col_2", "col_3"])
self.assertDictEqual(
dset.features, Features({"col_1": Value("int64"), "col_2": Value("string"), "col_3": Value("bool")})
)
features = Features({"col_1": Value("int64"), "col_2": Value("string"), "col_3": Value("bool")})
with Dataset.from_dict(data, features=features, info=DatasetInfo(features=features)) as dset:
self.assertListEqual(dset["col_1"], data["col_1"])
self.assertListEqual(dset["col_2"], data["col_2"])
self.assertListEqual(dset["col_3"], data["col_3"].to_pylist())
self.assertListEqual(list(dset.features.keys()), ["col_1", "col_2", "col_3"])
self.assertDictEqual(
dset.features, Features({"col_1": Value("int64"), "col_2": Value("string"), "col_3": Value("bool")})
)
features = Features({"col_1": Value("string"), "col_2": Value("string"), "col_3": Value("int32")})
with Dataset.from_dict(data, features=features) as dset:
# the integers are converted to strings
self.assertListEqual(dset["col_1"], [str(x) for x in data["col_1"]])
self.assertListEqual(dset["col_2"], data["col_2"])
self.assertListEqual(dset["col_3"], [int(x) for x in data["col_3"].to_pylist()])
self.assertListEqual(list(dset.features.keys()), ["col_1", "col_2", "col_3"])
self.assertDictEqual(
dset.features, Features({"col_1": Value("string"), "col_2": Value("string"), "col_3": Value("int32")})
)
features = Features({"col_1": Value("int64"), "col_2": Value("int64"), "col_3": Value("bool")})
self.assertRaises(ValueError, Dataset.from_dict, data, features=features)
def test_concatenate_mixed_memory_and_disk(self):
data1, data2, data3 = {"id": [0, 1, 2]}, {"id": [3, 4, 5]}, {"id": [6, 7]}
info1 = DatasetInfo(description="Dataset1")
info2 = DatasetInfo(description="Dataset2")
with tempfile.TemporaryDirectory() as tmp_dir:
with Dataset.from_dict(data1, info=info1).map(
cache_file_name=os.path.join(tmp_dir, "d1.arrow")
) as dset1, Dataset.from_dict(data2, info=info2).map(
cache_file_name=os.path.join(tmp_dir, "d2.arrow")
) as dset2, Dataset.from_dict(data3) as dset3:
with concatenate_datasets([dset1, dset2, dset3]) as concatenated_dset:
self.assertEqual(len(concatenated_dset), len(dset1) + len(dset2) + len(dset3))
self.assertListEqual(concatenated_dset["id"], dset1["id"] + dset2["id"] + dset3["id"])
@require_transformers
@pytest.mark.integration
def test_set_format_encode(self):
from transformers import BertTokenizer
tokenizer = BertTokenizer.from_pretrained("bert-base-uncased")
def encode(batch):
return tokenizer(batch["text"], padding="longest", return_tensors="np")
with Dataset.from_dict({"text": ["hello there", "foo"]}) as dset:
dset.set_transform(transform=encode)
self.assertEqual(str(dset[:2]), str(encode({"text": ["hello there", "foo"]})))
@require_tf
def test_tf_string_encoding(self):
data = {"col_1": ["á", "é", "í", "ó", "ú"], "col_2": ["à", "è", "ì", "ò", "ù"]}
with Dataset.from_dict(data) as dset:
tf_dset_wo_batch = dset.to_tf_dataset(columns=["col_1", "col_2"])
for tf_row, row in zip(tf_dset_wo_batch, dset):
self.assertEqual(tf_row["col_1"].numpy().decode("utf-8"), row["col_1"])
self.assertEqual(tf_row["col_2"].numpy().decode("utf-8"), row["col_2"])
tf_dset_w_batch = dset.to_tf_dataset(columns=["col_1", "col_2"], batch_size=2)
for tf_row, row in zip(tf_dset_w_batch.unbatch(), dset):
self.assertEqual(tf_row["col_1"].numpy().decode("utf-8"), row["col_1"])
self.assertEqual(tf_row["col_2"].numpy().decode("utf-8"), row["col_2"])
self.assertEqual(tf_dset_w_batch.unbatch().element_spec, tf_dset_wo_batch.element_spec)
self.assertEqual(tf_dset_w_batch.element_spec, tf_dset_wo_batch.batch(2).element_spec)
def test_cast_with_sliced_list():
old_features = Features({"foo": Sequence(Value("int64"))})
new_features = Features({"foo": Sequence(Value("int32"))})
dataset = Dataset.from_dict({"foo": [[i] * (i % 3) for i in range(20)]}, features=old_features)
casted_dataset = dataset.cast(new_features, batch_size=2) # small batch size to slice the ListArray
assert dataset["foo"] == casted_dataset["foo"]
assert casted_dataset.features == new_features
@pytest.mark.parametrize("include_nulls", [False, True])
def test_class_encode_column_with_none(include_nulls):
dataset = Dataset.from_dict({"col_1": ["a", "b", "c", None, "d", None]})
dataset = dataset.class_encode_column("col_1", include_nulls=include_nulls)
class_names = ["a", "b", "c", "d"]
if include_nulls:
class_names += ["None"]
assert isinstance(dataset.features["col_1"], ClassLabel)
assert set(dataset.features["col_1"].names) == set(class_names)
assert (None in dataset.unique("col_1")) == (not include_nulls)
@pytest.mark.parametrize("null_placement", ["first", "last"])
def test_sort_with_none(null_placement):
dataset = Dataset.from_dict({"col_1": ["item_2", "item_3", "item_1", None, "item_4", None]})
dataset = dataset.sort("col_1", null_placement=null_placement)
if null_placement == "first":
assert dataset["col_1"] == [None, None, "item_1", "item_2", "item_3", "item_4"]
else:
assert dataset["col_1"] == ["item_1", "item_2", "item_3", "item_4", None, None]
def test_update_metadata_with_features(dataset_dict):
table1 = pa.Table.from_pydict(dataset_dict)
features1 = Features.from_arrow_schema(table1.schema)
features2 = features1.copy()
features2["col_2"] = ClassLabel(num_classes=len(table1))
assert features1 != features2
table2 = update_metadata_with_features(table1, features2)
metadata = json.loads(table2.schema.metadata[b"huggingface"].decode())
assert features2 == Features.from_dict(metadata["info"]["features"])
with Dataset(table1) as dset1, Dataset(table2) as dset2:
assert dset1.features == features1
assert dset2.features == features2
@pytest.mark.parametrize("dataset_type", ["in_memory", "memory_mapped", "mixed"])
@pytest.mark.parametrize("axis, expected_shape", [(0, (4, 3)), (1, (2, 6))])
def test_concatenate_datasets(dataset_type, axis, expected_shape, dataset_dict, arrow_path):
table = {
"in_memory": InMemoryTable.from_pydict(dataset_dict),
"memory_mapped": MemoryMappedTable.from_file(arrow_path),
}
tables = [
table[dataset_type if dataset_type != "mixed" else "memory_mapped"].slice(0, 2), # shape = (2, 3)
table[dataset_type if dataset_type != "mixed" else "in_memory"].slice(2, 4), # shape = (2, 3)
]
if axis == 1: # don't duplicate columns
tables[1] = tables[1].rename_columns([col + "_bis" for col in tables[1].column_names])
datasets = [Dataset(table) for table in tables]
dataset = concatenate_datasets(datasets, axis=axis)
assert dataset.shape == expected_shape
assert_arrow_metadata_are_synced_with_dataset_features(dataset)
def test_concatenate_datasets_new_columns():
dataset1 = Dataset.from_dict({"col_1": ["a", "b", "c"]})
dataset2 = Dataset.from_dict({"col_1": ["d", "e", "f"], "col_2": [True, False, True]})
dataset = concatenate_datasets([dataset1, dataset2])
assert dataset.data.shape == (6, 2)
assert dataset.features == Features({"col_1": Value("string"), "col_2": Value("bool")})
assert dataset[:] == {"col_1": ["a", "b", "c", "d", "e", "f"], "col_2": [None, None, None, True, False, True]}
dataset3 = Dataset.from_dict({"col_3": ["a_1"]})
dataset = concatenate_datasets([dataset, dataset3])
assert dataset.data.shape == (7, 3)
assert dataset.features == Features({"col_1": Value("string"), "col_2": Value("bool"), "col_3": Value("string")})
assert dataset[:] == {
"col_1": ["a", "b", "c", "d", "e", "f", None],
"col_2": [None, None, None, True, False, True, None],
"col_3": [None, None, None, None, None, None, "a_1"],
}
@pytest.mark.parametrize("axis", [0, 1])
def test_concatenate_datasets_complex_features(axis):
n = 5
dataset1 = Dataset.from_dict(
{"col_1": [0] * n, "col_2": list(range(n))},
features=Features({"col_1": Value("int32"), "col_2": ClassLabel(num_classes=n)}),
)
if axis == 1:
dataset2 = dataset1.rename_columns({col: col + "_" for col in dataset1.column_names})
expected_features = Features({**dataset1.features, **dataset2.features})
else:
dataset2 = dataset1
expected_features = dataset1.features
assert concatenate_datasets([dataset1, dataset2], axis=axis).features == expected_features
@pytest.mark.parametrize("other_dataset_type", ["in_memory", "memory_mapped", "concatenation"])
@pytest.mark.parametrize("axis, expected_shape", [(0, (8, 3)), (1, (4, 6))])
def test_concatenate_datasets_with_concatenation_tables(
axis, expected_shape, other_dataset_type, dataset_dict, arrow_path
):
def _create_concatenation_table(axis):
if axis == 0: # shape: (4, 3) = (4, 1) + (4, 2)
concatenation_table = ConcatenationTable.from_blocks(
[
[
InMemoryTable.from_pydict({"col_1": dataset_dict["col_1"]}),
MemoryMappedTable.from_file(arrow_path).remove_column(0),
]
]
)
elif axis == 1: # shape: (4, 3) = (1, 3) + (3, 3)
concatenation_table = ConcatenationTable.from_blocks(
[
[InMemoryTable.from_pydict(dataset_dict).slice(0, 1)],
[MemoryMappedTable.from_file(arrow_path).slice(1, 4)],
]
)
return concatenation_table
concatenation_table = _create_concatenation_table(axis)
assert concatenation_table.shape == (4, 3)
if other_dataset_type == "in_memory":
other_table = InMemoryTable.from_pydict(dataset_dict)
elif other_dataset_type == "memory_mapped":
other_table = MemoryMappedTable.from_file(arrow_path)
elif other_dataset_type == "concatenation":
other_table = _create_concatenation_table(axis)
assert other_table.shape == (4, 3)
tables = [concatenation_table, other_table]
if axis == 1: # don't duplicate columns
tables[1] = tables[1].rename_columns([col + "_bis" for col in tables[1].column_names])
for tables in [tables, reversed(tables)]:
datasets = [Dataset(table) for table in tables]
dataset = concatenate_datasets(datasets, axis=axis)
assert dataset.shape == expected_shape
def test_concatenate_datasets_duplicate_columns(dataset):
with pytest.raises(ValueError) as excinfo:
concatenate_datasets([dataset, dataset], axis=1)
assert "duplicated" in str(excinfo.value)
def test_interleave_datasets():
d1 = Dataset.from_dict({"a": [0, 1, 2]})
d2 = Dataset.from_dict({"a": [10, 11, 12, 13]})
d3 = Dataset.from_dict({"a": [22, 21, 20]}).select([2, 1, 0])
dataset = interleave_datasets([d1, d2, d3])
expected_length = 3 * min(len(d1), len(d2), len(d3))
expected_values = [x["a"] for x in itertools.chain(*zip(d1, d2, d3))]
assert isinstance(dataset, Dataset)
assert len(dataset) == expected_length
assert dataset["a"] == expected_values
assert dataset._fingerprint == interleave_datasets([d1, d2, d3])._fingerprint
def test_interleave_datasets_probabilities():
seed = 42
probabilities = [0.3, 0.5, 0.2]
d1 = Dataset.from_dict({"a": [0, 1, 2]})
d2 = Dataset.from_dict({"a": [10, 11, 12, 13]})
d3 = Dataset.from_dict({"a": [22, 21, 20]}).select([2, 1, 0])
dataset = interleave_datasets([d1, d2, d3], probabilities=probabilities, seed=seed)
expected_length = 7 # hardcoded
expected_values = [10, 11, 20, 12, 0, 21, 13] # hardcoded
assert isinstance(dataset, Dataset)
assert len(dataset) == expected_length
assert dataset["a"] == expected_values
assert (
dataset._fingerprint == interleave_datasets([d1, d2, d3], probabilities=probabilities, seed=seed)._fingerprint
)
def test_interleave_datasets_oversampling_strategy():
d1 = Dataset.from_dict({"a": [0, 1, 2]})
d2 = Dataset.from_dict({"a": [10, 11, 12, 13]})
d3 = Dataset.from_dict({"a": [22, 21, 20]}).select([2, 1, 0])
dataset = interleave_datasets([d1, d2, d3], stopping_strategy="all_exhausted")
expected_length = 3 * max(len(d1), len(d2), len(d3))
expected_values = [0, 10, 20, 1, 11, 21, 2, 12, 22, 0, 13, 20] # hardcoded
assert isinstance(dataset, Dataset)
assert len(dataset) == expected_length
assert dataset["a"] == expected_values
assert dataset._fingerprint == interleave_datasets([d1, d2, d3], stopping_strategy="all_exhausted")._fingerprint
def test_interleave_datasets_probabilities_oversampling_strategy():
seed = 42
probabilities = [0.3, 0.5, 0.2]
d1 = Dataset.from_dict({"a": [0, 1, 2]})
d2 = Dataset.from_dict({"a": [10, 11, 12, 13]})
d3 = Dataset.from_dict({"a": [22, 21, 20]}).select([2, 1, 0])
dataset = interleave_datasets(
[d1, d2, d3], stopping_strategy="all_exhausted", probabilities=probabilities, seed=seed
)
expected_length = 16 # hardcoded
expected_values = [10, 11, 20, 12, 0, 21, 13, 10, 1, 11, 12, 22, 13, 20, 10, 2] # hardcoded
assert isinstance(dataset, Dataset)
assert len(dataset) == expected_length
assert dataset["a"] == expected_values
assert (
dataset._fingerprint
== interleave_datasets(
[d1, d2, d3], stopping_strategy="all_exhausted", probabilities=probabilities, seed=seed
)._fingerprint
)
@pytest.mark.parametrize("batch_size", [4, 5])
@pytest.mark.parametrize("drop_last_batch", [False, True])
def test_dataset_iter_batch(batch_size, drop_last_batch):
n = 25
dset = Dataset.from_dict({"i": list(range(n))})
all_col_values = list(range(n))
batches = []
for i, batch in enumerate(dset.iter(batch_size, drop_last_batch=drop_last_batch)):
assert batch == {"i": all_col_values[i * batch_size : (i + 1) * batch_size]}
batches.append(batch)
if drop_last_batch:
assert all(len(batch["i"]) == batch_size for batch in batches)
else:
assert all(len(batch["i"]) == batch_size for batch in batches[:-1])
assert len(batches[-1]["i"]) <= batch_size
@pytest.mark.parametrize(
"column, expected_dtype",
[(["a", "b", "c", "d"], "string"), ([1, 2, 3, 4], "int64"), ([1.0, 2.0, 3.0, 4.0], "float64")],
)
@pytest.mark.parametrize("in_memory", [False, True])
@pytest.mark.parametrize(
"transform",
[
None,
("shuffle", (42,), {}),
("with_format", ("pandas",), {}),
("class_encode_column", ("col_2",), {}),
("select", (range(3),), {}),
],
)
def test_dataset_add_column(column, expected_dtype, in_memory, transform, dataset_dict, arrow_path):
column_name = "col_4"
original_dataset = (
Dataset(InMemoryTable.from_pydict(dataset_dict))
if in_memory
else Dataset(MemoryMappedTable.from_file(arrow_path))
)
if transform is not None:
transform_name, args, kwargs = transform
original_dataset: Dataset = getattr(original_dataset, transform_name)(*args, **kwargs)
column = column[:3] if transform is not None and transform_name == "select" else column
dataset = original_dataset.add_column(column_name, column)
assert dataset.data.shape == (3, 4) if transform is not None and transform_name == "select" else (4, 4)
expected_features = {"col_1": "string", "col_2": "int64", "col_3": "float64"}
# Sort expected features as in the original dataset
expected_features = {feature: expected_features[feature] for feature in original_dataset.features}
# Add new column feature
expected_features[column_name] = expected_dtype
assert dataset.data.column_names == list(expected_features.keys())
for feature, expected_dtype in expected_features.items():
assert dataset.features[feature].dtype == expected_dtype
assert len(dataset.data.blocks) == 1 if in_memory else 2 # multiple InMemoryTables are consolidated as one
assert dataset.format["type"] == original_dataset.format["type"]
assert dataset._fingerprint != original_dataset._fingerprint
dataset.reset_format()
original_dataset.reset_format()
assert all(dataset[col] == original_dataset[col] for col in original_dataset.column_names)
assert set(dataset["col_4"]) == set(column)
if dataset._indices is not None:
dataset_indices = dataset._indices["indices"].to_pylist()
expected_dataset_indices = original_dataset._indices["indices"].to_pylist()
assert dataset_indices == expected_dataset_indices
assert_arrow_metadata_are_synced_with_dataset_features(dataset)
@pytest.mark.parametrize(
"transform",
[None, ("shuffle", (42,), {}), ("with_format", ("pandas",), {}), ("class_encode_column", ("col_2",), {})],
)
@pytest.mark.parametrize("in_memory", [False, True])
@pytest.mark.parametrize(
"item",
[
{"col_1": "2", "col_2": 2, "col_3": 2.0},
{"col_1": "2", "col_2": "2", "col_3": "2"},
{"col_1": 2, "col_2": 2, "col_3": 2},
{"col_1": 2.0, "col_2": 2.0, "col_3": 2.0},
],
)
def test_dataset_add_item(item, in_memory, dataset_dict, arrow_path, transform):
dataset_to_test = (
Dataset(InMemoryTable.from_pydict(dataset_dict))
if in_memory
else Dataset(MemoryMappedTable.from_file(arrow_path))
)
if transform is not None:
transform_name, args, kwargs = transform
dataset_to_test: Dataset = getattr(dataset_to_test, transform_name)(*args, **kwargs)
dataset = dataset_to_test.add_item(item)
assert dataset.data.shape == (5, 3)
expected_features = dataset_to_test.features
assert sorted(dataset.data.column_names) == sorted(expected_features.keys())
for feature, expected_dtype in expected_features.items():
assert dataset.features[feature] == expected_dtype
assert len(dataset.data.blocks) == 1 if in_memory else 2 # multiple InMemoryTables are consolidated as one
assert dataset.format["type"] == dataset_to_test.format["type"]
assert dataset._fingerprint != dataset_to_test._fingerprint
dataset.reset_format()
dataset_to_test.reset_format()
assert dataset[:-1] == dataset_to_test[:]
assert {k: int(v) for k, v in dataset[-1].items()} == {k: int(v) for k, v in item.items()}
if dataset._indices is not None:
dataset_indices = dataset._indices["indices"].to_pylist()
dataset_to_test_indices = dataset_to_test._indices["indices"].to_pylist()
assert dataset_indices == dataset_to_test_indices + [len(dataset_to_test._data)]
def test_dataset_add_item_new_columns():
dataset = Dataset.from_dict({"col_1": [0, 1, 2]}, features=Features({"col_1": Value("uint8")}))
dataset = dataset.add_item({"col_1": 3, "col_2": "a"})
assert dataset.data.shape == (4, 2)
assert dataset.features == Features({"col_1": Value("uint8"), "col_2": Value("string")})
assert dataset[:] == {"col_1": [0, 1, 2, 3], "col_2": [None, None, None, "a"]}
dataset = dataset.add_item({"col_3": True})
assert dataset.data.shape == (5, 3)
assert dataset.features == Features({"col_1": Value("uint8"), "col_2": Value("string"), "col_3": Value("bool")})
assert dataset[:] == {
"col_1": [0, 1, 2, 3, None],
"col_2": [None, None, None, "a", None],
"col_3": [None, None, None, None, True],
}
def test_dataset_add_item_introduce_feature_type():
dataset = Dataset.from_dict({"col_1": [None, None, None]})
dataset = dataset.add_item({"col_1": "a"})
assert dataset.data.shape == (4, 1)
assert dataset.features == Features({"col_1": Value("string")})
assert dataset[:] == {"col_1": [None, None, None, "a"]}
def test_dataset_filter_batched_indices():
ds = Dataset.from_dict({"num": [0, 1, 2, 3]})
ds = ds.filter(lambda num: num % 2 == 0, input_columns="num", batch_size=2)
assert all(item["num"] % 2 == 0 for item in ds)
@pytest.mark.parametrize("in_memory", [False, True])
def test_dataset_from_file(in_memory, dataset, arrow_file):
filename = arrow_file
with assert_arrow_memory_increases() if in_memory else assert_arrow_memory_doesnt_increase():
dataset_from_file = Dataset.from_file(filename, in_memory=in_memory)
assert dataset_from_file.features.type == dataset.features.type
assert dataset_from_file.features == dataset.features
assert dataset_from_file.cache_files == ([{"filename": filename}] if not in_memory else [])
def _check_csv_dataset(dataset, expected_features):
assert isinstance(dataset, Dataset)
assert dataset.num_rows == 4
assert dataset.num_columns == 3
assert dataset.column_names == ["col_1", "col_2", "col_3"]
for feature, expected_dtype in expected_features.items():
assert dataset.features[feature].dtype == expected_dtype
@pytest.mark.parametrize("keep_in_memory", [False, True])
def test_dataset_from_csv_keep_in_memory(keep_in_memory, csv_path, tmp_path):
cache_dir = tmp_path / "cache"
expected_features = {"col_1": "int64", "col_2": "int64", "col_3": "float64"}
with assert_arrow_memory_increases() if keep_in_memory else assert_arrow_memory_doesnt_increase():
dataset = Dataset.from_csv(csv_path, cache_dir=cache_dir, keep_in_memory=keep_in_memory)
_check_csv_dataset(dataset, expected_features)
@pytest.mark.parametrize(
"features",
[
None,
{"col_1": "string", "col_2": "int64", "col_3": "float64"},
{"col_1": "string", "col_2": "string", "col_3": "string"},
{"col_1": "int32", "col_2": "int32", "col_3": "int32"},
{"col_1": "float32", "col_2": "float32", "col_3": "float32"},
],
)
def test_dataset_from_csv_features(features, csv_path, tmp_path):
cache_dir = tmp_path / "cache"
# CSV file loses col_1 string dtype information: default now is "int64" instead of "string"
default_expected_features = {"col_1": "int64", "col_2": "int64", "col_3": "float64"}
expected_features = features.copy() if features else default_expected_features
features = (
Features({feature: Value(dtype) for feature, dtype in features.items()}) if features is not None else None
)
dataset = Dataset.from_csv(csv_path, features=features, cache_dir=cache_dir)
_check_csv_dataset(dataset, expected_features)
@pytest.mark.parametrize("split", [None, NamedSplit("train"), "train", "test"])
def test_dataset_from_csv_split(split, csv_path, tmp_path):
cache_dir = tmp_path / "cache"
expected_features = {"col_1": "int64", "col_2": "int64", "col_3": "float64"}
dataset = Dataset.from_csv(csv_path, cache_dir=cache_dir, split=split)
_check_csv_dataset(dataset, expected_features)
assert dataset.split == split if split else "train"
@pytest.mark.parametrize("path_type", [str, list])
def test_dataset_from_csv_path_type(path_type, csv_path, tmp_path):
if issubclass(path_type, str):
path = csv_path
elif issubclass(path_type, list):
path = [csv_path]
cache_dir = tmp_path / "cache"
expected_features = {"col_1": "int64", "col_2": "int64", "col_3": "float64"}
dataset = Dataset.from_csv(path, cache_dir=cache_dir)
_check_csv_dataset(dataset, expected_features)
def _check_json_dataset(dataset, expected_features):
assert isinstance(dataset, Dataset)
assert dataset.num_rows == 4
assert dataset.num_columns == 3
assert dataset.column_names == ["col_1", "col_2", "col_3"]
for feature, expected_dtype in expected_features.items():
assert dataset.features[feature].dtype == expected_dtype
@pytest.mark.parametrize("keep_in_memory", [False, True])
def test_dataset_from_json_keep_in_memory(keep_in_memory, jsonl_path, tmp_path):
cache_dir = tmp_path / "cache"
expected_features = {"col_1": "string", "col_2": "int64", "col_3": "float64"}
with assert_arrow_memory_increases() if keep_in_memory else assert_arrow_memory_doesnt_increase():
dataset = Dataset.from_json(jsonl_path, cache_dir=cache_dir, keep_in_memory=keep_in_memory)
_check_json_dataset(dataset, expected_features)
@pytest.mark.parametrize(
"features",
[
None,
{"col_1": "string", "col_2": "int64", "col_3": "float64"},
{"col_1": "string", "col_2": "string", "col_3": "string"},
{"col_1": "int32", "col_2": "int32", "col_3": "int32"},
{"col_1": "float32", "col_2": "float32", "col_3": "float32"},
],
)
def test_dataset_from_json_features(features, jsonl_path, tmp_path):
cache_dir = tmp_path / "cache"
default_expected_features = {"col_1": "string", "col_2": "int64", "col_3": "float64"}
expected_features = features.copy() if features else default_expected_features
features = (
Features({feature: Value(dtype) for feature, dtype in features.items()}) if features is not None else None
)
dataset = Dataset.from_json(jsonl_path, features=features, cache_dir=cache_dir)
_check_json_dataset(dataset, expected_features)
def test_dataset_from_json_with_class_label_feature(jsonl_str_path, tmp_path):
features = Features(
{"col_1": ClassLabel(names=["s0", "s1", "s2", "s3"]), "col_2": Value("int64"), "col_3": Value("float64")}
)
cache_dir = tmp_path / "cache"
dataset = Dataset.from_json(jsonl_str_path, features=features, cache_dir=cache_dir)
assert dataset.features["col_1"].dtype == "int64"
@pytest.mark.parametrize("split", [None, NamedSplit("train"), "train", "test"])
def test_dataset_from_json_split(split, jsonl_path, tmp_path):
cache_dir = tmp_path / "cache"
expected_features = {"col_1": "string", "col_2": "int64", "col_3": "float64"}
dataset = Dataset.from_json(jsonl_path, cache_dir=cache_dir, split=split)
_check_json_dataset(dataset, expected_features)
assert dataset.split == split if split else "train"
@pytest.mark.parametrize("path_type", [str, list])
def test_dataset_from_json_path_type(path_type, jsonl_path, tmp_path):
if issubclass(path_type, str):
path = jsonl_path
elif issubclass(path_type, list):
path = [jsonl_path]
cache_dir = tmp_path / "cache"
expected_features = {"col_1": "string", "col_2": "int64", "col_3": "float64"}
dataset = Dataset.from_json(path, cache_dir=cache_dir)
_check_json_dataset(dataset, expected_features)
def _check_parquet_dataset(dataset, expected_features):
assert isinstance(dataset, Dataset)
assert dataset.num_rows == 4
assert dataset.num_columns == 3
assert dataset.column_names == ["col_1", "col_2", "col_3"]
for feature, expected_dtype in expected_features.items():
assert dataset.features[feature].dtype == expected_dtype
@pytest.mark.parametrize("keep_in_memory", [False, True])
def test_dataset_from_parquet_keep_in_memory(keep_in_memory, parquet_path, tmp_path):
cache_dir = tmp_path / "cache"
expected_features = {"col_1": "string", "col_2": "int64", "col_3": "float64"}
with assert_arrow_memory_increases() if keep_in_memory else assert_arrow_memory_doesnt_increase():
dataset = Dataset.from_parquet(parquet_path, cache_dir=cache_dir, keep_in_memory=keep_in_memory)
_check_parquet_dataset(dataset, expected_features)
@pytest.mark.parametrize(
"features",
[
None,
{"col_1": "string", "col_2": "int64", "col_3": "float64"},
{"col_1": "string", "col_2": "string", "col_3": "string"},
{"col_1": "int32", "col_2": "int32", "col_3": "int32"},
{"col_1": "float32", "col_2": "float32", "col_3": "float32"},
],
)
def test_dataset_from_parquet_features(features, parquet_path, tmp_path):
cache_dir = tmp_path / "cache"
default_expected_features = {"col_1": "string", "col_2": "int64", "col_3": "float64"}
expected_features = features.copy() if features else default_expected_features
features = (
Features({feature: Value(dtype) for feature, dtype in features.items()}) if features is not None else None
)
dataset = Dataset.from_parquet(parquet_path, features=features, cache_dir=cache_dir)
_check_parquet_dataset(dataset, expected_features)
@pytest.mark.parametrize("split", [None, NamedSplit("train"), "train", "test"])
def test_dataset_from_parquet_split(split, parquet_path, tmp_path):
cache_dir = tmp_path / "cache"
expected_features = {"col_1": "string", "col_2": "int64", "col_3": "float64"}
dataset = Dataset.from_parquet(parquet_path, cache_dir=cache_dir, split=split)
_check_parquet_dataset(dataset, expected_features)
assert dataset.split == split if split else "train"
@pytest.mark.parametrize("path_type", [str, list])
def test_dataset_from_parquet_path_type(path_type, parquet_path, tmp_path):
if issubclass(path_type, str):
path = parquet_path
elif issubclass(path_type, list):
path = [parquet_path]
cache_dir = tmp_path / "cache"
expected_features = {"col_1": "string", "col_2": "int64", "col_3": "float64"}
dataset = Dataset.from_parquet(path, cache_dir=cache_dir)
_check_parquet_dataset(dataset, expected_features)
def _check_text_dataset(dataset, expected_features):
assert isinstance(dataset, Dataset)
assert dataset.num_rows == 4
assert dataset.num_columns == 1
assert dataset.column_names == ["text"]
for feature, expected_dtype in expected_features.items():
assert dataset.features[feature].dtype == expected_dtype
@pytest.mark.parametrize("keep_in_memory", [False, True])
def test_dataset_from_text_keep_in_memory(keep_in_memory, text_path, tmp_path):
cache_dir = tmp_path / "cache"
expected_features = {"text": "string"}
with assert_arrow_memory_increases() if keep_in_memory else assert_arrow_memory_doesnt_increase():
dataset = Dataset.from_text(text_path, cache_dir=cache_dir, keep_in_memory=keep_in_memory)
_check_text_dataset(dataset, expected_features)
@pytest.mark.parametrize(
"features",
[
None,
{"text": "string"},
{"text": "int32"},
{"text": "float32"},
],
)
def test_dataset_from_text_features(features, text_path, tmp_path):
cache_dir = tmp_path / "cache"
default_expected_features = {"text": "string"}
expected_features = features.copy() if features else default_expected_features
features = (
Features({feature: Value(dtype) for feature, dtype in features.items()}) if features is not None else None
)
dataset = Dataset.from_text(text_path, features=features, cache_dir=cache_dir)
_check_text_dataset(dataset, expected_features)
@pytest.mark.parametrize("split", [None, NamedSplit("train"), "train", "test"])
def test_dataset_from_text_split(split, text_path, tmp_path):
cache_dir = tmp_path / "cache"
expected_features = {"text": "string"}
dataset = Dataset.from_text(text_path, cache_dir=cache_dir, split=split)
_check_text_dataset(dataset, expected_features)
assert dataset.split == split if split else "train"
@pytest.mark.parametrize("path_type", [str, list])
def test_dataset_from_text_path_type(path_type, text_path, tmp_path):
if issubclass(path_type, str):
path = text_path
elif issubclass(path_type, list):
path = [text_path]
cache_dir = tmp_path / "cache"
expected_features = {"text": "string"}
dataset = Dataset.from_text(path, cache_dir=cache_dir)
_check_text_dataset(dataset, expected_features)
@pytest.fixture
def data_generator():
def _gen():
data = [
{"col_1": "0", "col_2": 0, "col_3": 0.0},
{"col_1": "1", "col_2": 1, "col_3": 1.0},
{"col_1": "2", "col_2": 2, "col_3": 2.0},
{"col_1": "3", "col_2": 3, "col_3": 3.0},
]
for item in data:
yield item
return _gen
def _check_generator_dataset(dataset, expected_features):
assert isinstance(dataset, Dataset)
assert dataset.num_rows == 4
assert dataset.num_columns == 3
assert dataset.column_names == ["col_1", "col_2", "col_3"]
for feature, expected_dtype in expected_features.items():
assert dataset.features[feature].dtype == expected_dtype
@pytest.mark.parametrize("keep_in_memory", [False, True])
def test_dataset_from_generator_keep_in_memory(keep_in_memory, data_generator, tmp_path):
cache_dir = tmp_path / "cache"
expected_features = {"col_1": "string", "col_2": "int64", "col_3": "float64"}
with assert_arrow_memory_increases() if keep_in_memory else assert_arrow_memory_doesnt_increase():
dataset = Dataset.from_generator(data_generator, cache_dir=cache_dir, keep_in_memory=keep_in_memory)
_check_generator_dataset(dataset, expected_features)
@pytest.mark.parametrize(
"features",
[
None,
{"col_1": "string", "col_2": "int64", "col_3": "float64"},
{"col_1": "string", "col_2": "string", "col_3": "string"},
{"col_1": "int32", "col_2": "int32", "col_3": "int32"},
{"col_1": "float32", "col_2": "float32", "col_3": "float32"},
],
)
def test_dataset_from_generator_features(features, data_generator, tmp_path):
cache_dir = tmp_path / "cache"
default_expected_features = {"col_1": "string", "col_2": "int64", "col_3": "float64"}
expected_features = features.copy() if features else default_expected_features
features = (
Features({feature: Value(dtype) for feature, dtype in features.items()}) if features is not None else None
)
dataset = Dataset.from_generator(data_generator, features=features, cache_dir=cache_dir)
_check_generator_dataset(dataset, expected_features)
@require_not_windows
@require_dill_gt_0_3_2
@require_pyspark
def test_from_spark():
import pyspark
spark = pyspark.sql.SparkSession.builder.master("local[*]").appName("pyspark").getOrCreate()
data = [
("0", 0, 0.0),
("1", 1, 1.0),
("2", 2, 2.0),
("3", 3, 3.0),
]
df = spark.createDataFrame(data, "col_1: string, col_2: int, col_3: float")
dataset = Dataset.from_spark(df)
assert isinstance(dataset, Dataset)
assert dataset.num_rows == 4
assert dataset.num_columns == 3
assert dataset.column_names == ["col_1", "col_2", "col_3"]
@require_not_windows
@require_dill_gt_0_3_2
@require_pyspark
def test_from_spark_features():
import PIL.Image
import pyspark
spark = pyspark.sql.SparkSession.builder.master("local[*]").appName("pyspark").getOrCreate()
data = [(0, np.arange(4 * 4 * 3).reshape(4, 4, 3).tolist())]
df = spark.createDataFrame(data, "idx: int, image: array<array<array<int>>>")
features = Features({"idx": Value("int64"), "image": Image()})
dataset = Dataset.from_spark(
df,
features=features,
)
assert isinstance(dataset, Dataset)
assert dataset.num_rows == 1
assert dataset.num_columns == 2
assert dataset.column_names == ["idx", "image"]
assert isinstance(dataset[0]["image"], PIL.Image.Image)
assert dataset.features == features
assert_arrow_metadata_are_synced_with_dataset_features(dataset)
@require_not_windows
@require_dill_gt_0_3_2
@require_pyspark
def test_from_spark_different_cache():
import pyspark
spark = pyspark.sql.SparkSession.builder.master("local[*]").appName("pyspark").getOrCreate()
df = spark.createDataFrame([("0", 0)], "col_1: string, col_2: int")
dataset = Dataset.from_spark(df)
assert isinstance(dataset, Dataset)
different_df = spark.createDataFrame([("1", 1)], "col_1: string, col_2: int")
different_dataset = Dataset.from_spark(different_df)
assert isinstance(different_dataset, Dataset)
assert dataset[0]["col_1"] == "0"
# Check to make sure that the second dataset wasn't read from the cache.
assert different_dataset[0]["col_1"] == "1"
def _check_sql_dataset(dataset, expected_features):
assert isinstance(dataset, Dataset)
assert dataset.num_rows == 4
assert dataset.num_columns == 3
assert dataset.column_names == ["col_1", "col_2", "col_3"]
for feature, expected_dtype in expected_features.items():
assert dataset.features[feature].dtype == expected_dtype
@require_sqlalchemy
@pytest.mark.parametrize("con_type", ["string", "engine"])
def test_dataset_from_sql_con_type(con_type, sqlite_path, tmp_path, set_sqlalchemy_silence_uber_warning):
cache_dir = tmp_path / "cache"
expected_features = {"col_1": "string", "col_2": "int64", "col_3": "float64"}
if con_type == "string":
con = "sqlite:///" + sqlite_path
elif con_type == "engine":
import sqlalchemy
con = sqlalchemy.create_engine("sqlite:///" + sqlite_path)
# # https://github.com/huggingface/datasets/issues/2832 needs to be fixed first for this to work
# with caplog.at_level(INFO):
# dataset = Dataset.from_sql(
# "dataset",
# con,
# cache_dir=cache_dir,
# )
# if con_type == "string":
# assert "couldn't be hashed properly" not in caplog.text
# elif con_type == "engine":
# assert "couldn't be hashed properly" in caplog.text
dataset = Dataset.from_sql(
"dataset",
con,
cache_dir=cache_dir,
)
_check_sql_dataset(dataset, expected_features)
@require_sqlalchemy
@pytest.mark.parametrize(
"features",
[
None,
{"col_1": "string", "col_2": "int64", "col_3": "float64"},
{"col_1": "string", "col_2": "string", "col_3": "string"},
{"col_1": "int32", "col_2": "int32", "col_3": "int32"},
{"col_1": "float32", "col_2": "float32", "col_3": "float32"},
],
)
def test_dataset_from_sql_features(features, sqlite_path, tmp_path, set_sqlalchemy_silence_uber_warning):
cache_dir = tmp_path / "cache"
default_expected_features = {"col_1": "string", "col_2": "int64", "col_3": "float64"}
expected_features = features.copy() if features else default_expected_features
features = (
Features({feature: Value(dtype) for feature, dtype in features.items()}) if features is not None else None
)
dataset = Dataset.from_sql("dataset", "sqlite:///" + sqlite_path, features=features, cache_dir=cache_dir)
_check_sql_dataset(dataset, expected_features)
@require_sqlalchemy
@pytest.mark.parametrize("keep_in_memory", [False, True])
def test_dataset_from_sql_keep_in_memory(keep_in_memory, sqlite_path, tmp_path, set_sqlalchemy_silence_uber_warning):
cache_dir = tmp_path / "cache"
expected_features = {"col_1": "string", "col_2": "int64", "col_3": "float64"}
with assert_arrow_memory_increases() if keep_in_memory else assert_arrow_memory_doesnt_increase():
dataset = Dataset.from_sql(
"dataset", "sqlite:///" + sqlite_path, cache_dir=cache_dir, keep_in_memory=keep_in_memory
)
_check_sql_dataset(dataset, expected_features)
def test_dataset_to_json(dataset, tmp_path):
file_path = tmp_path / "test_path.jsonl"
bytes_written = dataset.to_json(path_or_buf=file_path)
assert file_path.is_file()
assert bytes_written == file_path.stat().st_size
df = pd.read_json(file_path, orient="records", lines=True)
assert df.shape == dataset.shape
assert list(df.columns) == list(dataset.column_names)
@pytest.mark.parametrize("in_memory", [False, True])
@pytest.mark.parametrize(
"method_and_params",
[
("rename_column", (), {"original_column_name": "labels", "new_column_name": "label"}),
("remove_columns", (), {"column_names": "labels"}),
(
"cast",
(),
{
"features": Features(
{
"tokens": Sequence(Value("string")),
"labels": Sequence(Value("int16")),
"answers": Sequence(
{
"text": Value("string"),
"answer_start": Value("int32"),
}
),
"id": Value("int32"),
}
)
},
),
("flatten", (), {}),
],
)
def test_pickle_dataset_after_transforming_the_table(in_memory, method_and_params, arrow_file):
method, args, kwargs = method_and_params
with Dataset.from_file(arrow_file, in_memory=in_memory) as dataset, Dataset.from_file(
arrow_file, in_memory=in_memory
) as reference_dataset:
out = getattr(dataset, method)(*args, **kwargs)
dataset = out if out is not None else dataset
pickled_dataset = pickle.dumps(dataset)
reloaded_dataset = pickle.loads(pickled_dataset)
assert dataset._data != reference_dataset._data
assert dataset._data.table == reloaded_dataset._data.table
def test_dummy_dataset_serialize_fs(dataset, mockfs):
dataset_path = "mock://my_dataset"
dataset.save_to_disk(dataset_path, storage_options=mockfs.storage_options)
assert mockfs.isdir(dataset_path)
assert mockfs.glob(dataset_path + "/*")
reloaded = dataset.load_from_disk(dataset_path, storage_options=mockfs.storage_options)
assert len(reloaded) == len(dataset)
assert reloaded.features == dataset.features
assert reloaded.to_dict() == dataset.to_dict()
@pytest.mark.parametrize(
"uri_or_path",
[
"relative/path",
"/absolute/path",
"s3://bucket/relative/path",
"hdfs://relative/path",
"hdfs:///absolute/path",
],
)
def test_build_local_temp_path(uri_or_path):
extracted_path = strip_protocol(uri_or_path)
local_temp_path = Dataset._build_local_temp_path(extracted_path).as_posix()
extracted_path_without_anchor = Path(extracted_path).relative_to(Path(extracted_path).anchor).as_posix()
path_relative_to_tmp_dir = local_temp_path.split("tmp")[-1].split("/", 1)[1]
assert (
"tmp" in local_temp_path
and "hdfs" not in path_relative_to_tmp_dir
and "s3" not in path_relative_to_tmp_dir
and not local_temp_path.startswith(extracted_path_without_anchor)
and local_temp_path.endswith(extracted_path_without_anchor)
), f"Local temp path: {local_temp_path}"
class TaskTemplatesTest(TestCase):
def test_task_text_classification(self):
labels = sorted(["pos", "neg"])
features_before_cast = Features(
{
"input_text": Value("string"),
"input_labels": ClassLabel(names=labels),
}
)
# Labels are cast to tuple during `TextClassification.__post_init_`, so we do the same here
features_after_cast = Features(
{
"text": Value("string"),
"labels": ClassLabel(names=labels),
}
)
# Label names are added in `DatasetInfo.__post_init__` so not needed here
task_without_labels = TextClassification(text_column="input_text", label_column="input_labels")
info1 = DatasetInfo(
features=features_before_cast,
task_templates=task_without_labels,
)
# Label names are required when passing a TextClassification template directly to `Dataset.prepare_for_task`
# However they also can be used to define `DatasetInfo` so we include a test for this too
task_with_labels = TextClassification(text_column="input_text", label_column="input_labels")
info2 = DatasetInfo(
features=features_before_cast,
task_templates=task_with_labels,
)
data = {"input_text": ["i love transformers!"], "input_labels": [1]}
# Test we can load from task name when label names not included in template (default behaviour)
with Dataset.from_dict(data, info=info1) as dset:
self.assertSetEqual({"input_text", "input_labels"}, set(dset.column_names))
self.assertDictEqual(features_before_cast, dset.features)
with dset.prepare_for_task(task="text-classification") as dset:
self.assertSetEqual({"labels", "text"}, set(dset.column_names))
self.assertDictEqual(features_after_cast, dset.features)
# Test we can load from task name when label names included in template
with Dataset.from_dict(data, info=info2) as dset:
self.assertSetEqual({"input_text", "input_labels"}, set(dset.column_names))
self.assertDictEqual(features_before_cast, dset.features)
with dset.prepare_for_task(task="text-classification") as dset:
self.assertSetEqual({"labels", "text"}, set(dset.column_names))
self.assertDictEqual(features_after_cast, dset.features)
# Test we can load from TextClassification template
info1.task_templates = None
with Dataset.from_dict(data, info=info1) as dset:
with dset.prepare_for_task(task=task_with_labels) as dset:
self.assertSetEqual({"labels", "text"}, set(dset.column_names))
self.assertDictEqual(features_after_cast, dset.features)
def test_task_question_answering(self):
features_before_cast = Features(
{
"input_context": Value("string"),
"input_question": Value("string"),
"input_answers": Sequence(
{
"text": Value("string"),
"answer_start": Value("int32"),
}
),
}
)
features_after_cast = Features(
{
"context": Value("string"),
"question": Value("string"),
"answers": Sequence(
{
"text": Value("string"),
"answer_start": Value("int32"),
}
),
}
)
task = QuestionAnsweringExtractive(
context_column="input_context", question_column="input_question", answers_column="input_answers"
)
info = DatasetInfo(features=features_before_cast, task_templates=task)
data = {
"input_context": ["huggingface is going to the moon!"],
"input_question": ["where is huggingface going?"],
"input_answers": [{"text": ["to the moon!"], "answer_start": [2]}],
}
# Test we can load from task name
with Dataset.from_dict(data, info=info) as dset:
self.assertSetEqual(
{"input_context", "input_question", "input_answers.text", "input_answers.answer_start"},
set(dset.flatten().column_names),
)
self.assertDictEqual(features_before_cast, dset.features)
with dset.prepare_for_task(task="question-answering-extractive") as dset:
self.assertSetEqual(
{"context", "question", "answers.text", "answers.answer_start"},
set(dset.flatten().column_names),
)
self.assertDictEqual(features_after_cast, dset.features)
# Test we can load from QuestionAnsweringExtractive template
info.task_templates = None
with Dataset.from_dict(data, info=info) as dset:
with dset.prepare_for_task(task=task) as dset:
self.assertSetEqual(
{"context", "question", "answers.text", "answers.answer_start"},
set(dset.flatten().column_names),
)
self.assertDictEqual(features_after_cast, dset.features)
def test_task_summarization(self):
# Include a dummy extra column `dummy` to test we drop it correctly
features_before_cast = Features(
{"input_text": Value("string"), "input_summary": Value("string"), "dummy": Value("string")}
)
features_after_cast = Features({"text": Value("string"), "summary": Value("string")})
task = Summarization(text_column="input_text", summary_column="input_summary")
info = DatasetInfo(features=features_before_cast, task_templates=task)
data = {
"input_text": ["jack and jill took a taxi to attend a super duper party in the city."],
"input_summary": ["jack and jill attend party"],
"dummy": ["123456"],
}
# Test we can load from task name
with Dataset.from_dict(data, info=info) as dset:
with dset.prepare_for_task(task="summarization") as dset:
self.assertSetEqual(
{"text", "summary"},
set(dset.column_names),
)
self.assertDictEqual(features_after_cast, dset.features)
# Test we can load from Summarization template
info.task_templates = None
with Dataset.from_dict(data, info=info) as dset:
with dset.prepare_for_task(task=task) as dset:
self.assertSetEqual(
{"text", "summary"},
set(dset.column_names),
)
self.assertDictEqual(features_after_cast, dset.features)
def test_task_automatic_speech_recognition(self):
# Include a dummy extra column `dummy` to test we drop it correctly
features_before_cast = Features(
{
"input_audio": Audio(sampling_rate=16_000),
"input_transcription": Value("string"),
"dummy": Value("string"),
}
)
features_after_cast = Features({"audio": Audio(sampling_rate=16_000), "transcription": Value("string")})
task = AutomaticSpeechRecognition(audio_column="input_audio", transcription_column="input_transcription")
info = DatasetInfo(features=features_before_cast, task_templates=task)
data = {
"input_audio": [{"bytes": None, "path": "path/to/some/audio/file.wav"}],
"input_transcription": ["hello, my name is bob!"],
"dummy": ["123456"],
}
# Test we can load from task name
with Dataset.from_dict(data, info=info) as dset:
with dset.prepare_for_task(task="automatic-speech-recognition") as dset:
self.assertSetEqual(
{"audio", "transcription"},
set(dset.column_names),
)
self.assertDictEqual(features_after_cast, dset.features)
# Test we can load from Summarization template
info.task_templates = None
with Dataset.from_dict(data, info=info) as dset:
with dset.prepare_for_task(task=task) as dset:
self.assertSetEqual(
{"audio", "transcription"},
set(dset.column_names),
)
self.assertDictEqual(features_after_cast, dset.features)
def test_task_with_no_template(self):
data = {"input_text": ["i love transformers!"], "input_labels": [1]}
with Dataset.from_dict(data) as dset:
with self.assertRaises(ValueError):
dset.prepare_for_task("text-classification")
def test_task_with_incompatible_templates(self):
labels = sorted(["pos", "neg"])
features = Features(
{
"input_text": Value("string"),
"input_labels": ClassLabel(names=labels),
}
)
task = TextClassification(text_column="input_text", label_column="input_labels")
info = DatasetInfo(
features=features,
task_templates=task,
)
data = {"input_text": ["i love transformers!"], "input_labels": [1]}
with Dataset.from_dict(data, info=info) as dset:
# Invalid task name
self.assertRaises(ValueError, dset.prepare_for_task, "this-task-does-not-exist")
# Invalid task type
self.assertRaises(ValueError, dset.prepare_for_task, 1)
def test_task_with_multiple_compatible_task_templates(self):
features = Features(
{
"text1": Value("string"),
"text2": Value("string"),
}
)
task1 = LanguageModeling(text_column="text1")
task2 = LanguageModeling(text_column="text2")
info = DatasetInfo(
features=features,
task_templates=[task1, task2],
)
data = {"text1": ["i love transformers!"], "text2": ["i love datasets!"]}
with Dataset.from_dict(data, info=info) as dset:
self.assertRaises(ValueError, dset.prepare_for_task, "language-modeling", id=3)
with dset.prepare_for_task("language-modeling") as dset1:
self.assertEqual(dset1[0]["text"], "i love transformers!")
with dset.prepare_for_task("language-modeling", id=1) as dset2:
self.assertEqual(dset2[0]["text"], "i love datasets!")
def test_task_templates_empty_after_preparation(self):
features = Features(
{
"input_text": Value("string"),
"input_labels": ClassLabel(names=["pos", "neg"]),
}
)
task = TextClassification(text_column="input_text", label_column="input_labels")
info = DatasetInfo(
features=features,
task_templates=task,
)
data = {"input_text": ["i love transformers!"], "input_labels": [1]}
with Dataset.from_dict(data, info=info) as dset:
with dset.prepare_for_task(task="text-classification") as dset:
self.assertIsNone(dset.info.task_templates)
def test_align_labels_with_mapping_classification(self):
features = Features(
{
"input_text": Value("string"),
"input_labels": ClassLabel(num_classes=3, names=["entailment", "neutral", "contradiction"]),
}
)
data = {"input_text": ["a", "a", "b", "b", "c", "c"], "input_labels": [0, 0, 1, 1, 2, 2]}
label2id = {"CONTRADICTION": 0, "ENTAILMENT": 2, "NEUTRAL": 1}
id2label = {v: k for k, v in label2id.items()}
expected_labels = [2, 2, 1, 1, 0, 0]
expected_label_names = [id2label[idx] for idx in expected_labels]
with Dataset.from_dict(data, features=features) as dset:
with dset.align_labels_with_mapping(label2id, "input_labels") as dset:
self.assertListEqual(expected_labels, dset["input_labels"])
aligned_label_names = [dset.features["input_labels"].int2str(idx) for idx in dset["input_labels"]]
self.assertListEqual(expected_label_names, aligned_label_names)
def test_align_labels_with_mapping_ner(self):
features = Features(
{
"input_text": Value("string"),
"input_labels": Sequence(
ClassLabel(
names=[
"b-per",
"i-per",
"o",
]
)
),
}
)
data = {"input_text": [["Optimus", "Prime", "is", "a", "Transformer"]], "input_labels": [[0, 1, 2, 2, 2]]}
label2id = {"B-PER": 2, "I-PER": 1, "O": 0}
id2label = {v: k for k, v in label2id.items()}
expected_labels = [[2, 1, 0, 0, 0]]
expected_label_names = [[id2label[idx] for idx in seq] for seq in expected_labels]
with Dataset.from_dict(data, features=features) as dset:
with dset.align_labels_with_mapping(label2id, "input_labels") as dset:
self.assertListEqual(expected_labels, dset["input_labels"])
aligned_label_names = [
dset.features["input_labels"].feature.int2str(idx) for idx in dset["input_labels"]
]
self.assertListEqual(expected_label_names, aligned_label_names)
def test_concatenate_with_no_task_templates(self):
info = DatasetInfo(task_templates=None)
data = {"text": ["i love transformers!"], "labels": [1]}
with Dataset.from_dict(data, info=info) as dset1, Dataset.from_dict(
data, info=info
) as dset2, Dataset.from_dict(data, info=info) as dset3:
with concatenate_datasets([dset1, dset2, dset3]) as dset_concat:
self.assertEqual(dset_concat.info.task_templates, None)
def test_concatenate_with_equal_task_templates(self):
labels = ["neg", "pos"]
task_template = TextClassification(text_column="text", label_column="labels")
info = DatasetInfo(
features=Features({"text": Value("string"), "labels": ClassLabel(names=labels)}),
# Label names are added in `DatasetInfo.__post_init__` so not included here
task_templates=TextClassification(text_column="text", label_column="labels"),
)
data = {"text": ["i love transformers!"], "labels": [1]}
with Dataset.from_dict(data, info=info) as dset1, Dataset.from_dict(
data, info=info
) as dset2, Dataset.from_dict(data, info=info) as dset3:
with concatenate_datasets([dset1, dset2, dset3]) as dset_concat:
self.assertListEqual(dset_concat.info.task_templates, [task_template])
def test_concatenate_with_mixed_task_templates_in_common(self):
tc_template = TextClassification(text_column="text", label_column="labels")
qa_template = QuestionAnsweringExtractive(
question_column="question", context_column="context", answers_column="answers"
)
info1 = DatasetInfo(
task_templates=[qa_template],
features=Features(
{
"text": Value("string"),
"labels": ClassLabel(names=["pos", "neg"]),
"context": Value("string"),
"question": Value("string"),
"answers": Sequence(
{
"text": Value("string"),
"answer_start": Value("int32"),
}
),
}
),
)
info2 = DatasetInfo(
task_templates=[qa_template, tc_template],
features=Features(
{
"text": Value("string"),
"labels": ClassLabel(names=["pos", "neg"]),
"context": Value("string"),
"question": Value("string"),
"answers": Sequence(
{
"text": Value("string"),
"answer_start": Value("int32"),
}
),
}
),
)
data = {
"text": ["i love transformers!"],
"labels": [1],
"context": ["huggingface is going to the moon!"],
"question": ["where is huggingface going?"],
"answers": [{"text": ["to the moon!"], "answer_start": [2]}],
}
with Dataset.from_dict(data, info=info1) as dset1, Dataset.from_dict(
data, info=info2
) as dset2, Dataset.from_dict(data, info=info2) as dset3:
with concatenate_datasets([dset1, dset2, dset3]) as dset_concat:
self.assertListEqual(dset_concat.info.task_templates, [qa_template])
def test_concatenate_with_no_mixed_task_templates_in_common(self):
tc_template1 = TextClassification(text_column="text", label_column="labels")
tc_template2 = TextClassification(text_column="text", label_column="sentiment")
qa_template = QuestionAnsweringExtractive(
question_column="question", context_column="context", answers_column="answers"
)
info1 = DatasetInfo(
features=Features(
{
"text": Value("string"),
"labels": ClassLabel(names=["pos", "neg"]),
"sentiment": ClassLabel(names=["pos", "neg", "neutral"]),
"context": Value("string"),
"question": Value("string"),
"answers": Sequence(
{
"text": Value("string"),
"answer_start": Value("int32"),
}
),
}
),
task_templates=[tc_template1],
)
info2 = DatasetInfo(
features=Features(
{
"text": Value("string"),
"labels": ClassLabel(names=["pos", "neg"]),
"sentiment": ClassLabel(names=["pos", "neg", "neutral"]),
"context": Value("string"),
"question": Value("string"),
"answers": Sequence(
{
"text": Value("string"),
"answer_start": Value("int32"),
}
),
}
),
task_templates=[tc_template2],
)
info3 = DatasetInfo(
features=Features(
{
"text": Value("string"),
"labels": ClassLabel(names=["pos", "neg"]),
"sentiment": ClassLabel(names=["pos", "neg", "neutral"]),
"context": Value("string"),
"question": Value("string"),
"answers": Sequence(
{
"text": Value("string"),
"answer_start": Value("int32"),
}
),
}
),
task_templates=[qa_template],
)
data = {
"text": ["i love transformers!"],
"labels": [1],
"sentiment": [0],
"context": ["huggingface is going to the moon!"],
"question": ["where is huggingface going?"],
"answers": [{"text": ["to the moon!"], "answer_start": [2]}],
}
with Dataset.from_dict(data, info=info1) as dset1, Dataset.from_dict(
data, info=info2
) as dset2, Dataset.from_dict(data, info=info3) as dset3:
with concatenate_datasets([dset1, dset2, dset3]) as dset_concat:
self.assertEqual(dset_concat.info.task_templates, None)
def test_task_text_classification_when_columns_removed(self):
labels = sorted(["pos", "neg"])
features_before_map = Features(
{
"input_text": Value("string"),
"input_labels": ClassLabel(names=labels),
}
)
features_after_map = Features({"new_column": Value("int64")})
# Label names are added in `DatasetInfo.__post_init__` so not needed here
task = TextClassification(text_column="input_text", label_column="input_labels")
info = DatasetInfo(
features=features_before_map,
task_templates=task,
)
data = {"input_text": ["i love transformers!"], "input_labels": [1]}
with Dataset.from_dict(data, info=info) as dset:
with dset.map(lambda x: {"new_column": 0}, remove_columns=dset.column_names) as dset:
self.assertDictEqual(dset.features, features_after_map)
class StratifiedTest(TestCase):
def test_errors_train_test_split_stratify(self):
ys = [
np.array([0, 0, 0, 0, 1, 1, 1, 2, 2, 2, 2, 2]),
np.array([0, 1, 1, 1, 2, 2, 2, 3, 3, 3]),
np.array([0, 1, 2, 3, 0, 1, 2, 3, 0, 1, 2, 3, 0, 1, 2] * 2),
np.array([0, 0, 1, 1, 2, 2, 3, 3, 4, 4, 5, 5]),
np.array([0, 0, 1, 1, 2, 2, 3, 3, 4, 4, 5, 5]),
]
for i in range(len(ys)):
features = Features({"text": Value("int64"), "label": ClassLabel(len(np.unique(ys[i])))})
data = {"text": np.ones(len(ys[i])), "label": ys[i]}
d1 = Dataset.from_dict(data, features=features)
# For checking stratify_by_column exist as key in self.features.keys()
if i == 0:
self.assertRaises(ValueError, d1.train_test_split, 0.33, stratify_by_column="labl")
# For checking minimum class count error
elif i == 1:
self.assertRaises(ValueError, d1.train_test_split, 0.33, stratify_by_column="label")
# For check typeof label as ClassLabel type
elif i == 2:
d1 = Dataset.from_dict(data)
self.assertRaises(ValueError, d1.train_test_split, 0.33, stratify_by_column="label")
# For checking test_size should be greater than or equal to number of classes
elif i == 3:
self.assertRaises(ValueError, d1.train_test_split, 0.30, stratify_by_column="label")
# For checking train_size should be greater than or equal to number of classes
elif i == 4:
self.assertRaises(ValueError, d1.train_test_split, 0.60, stratify_by_column="label")
def test_train_test_split_startify(self):
ys = [
np.array([0, 0, 0, 0, 1, 1, 1, 2, 2, 2, 2, 2]),
np.array([0, 0, 0, 1, 1, 1, 2, 2, 2, 3, 3, 3]),
np.array([0, 1, 2, 3, 0, 1, 2, 3, 0, 1, 2, 3, 0, 1, 2] * 2),
np.array([0, 0, 1, 1, 1, 2, 2, 2, 3, 3, 3, 3, 3, 3, 3, 3]),
np.array([0] * 800 + [1] * 50),
]
for y in ys:
features = Features({"text": Value("int64"), "label": ClassLabel(len(np.unique(y)))})
data = {"text": np.ones(len(y)), "label": y}
d1 = Dataset.from_dict(data, features=features)
d1 = d1.train_test_split(test_size=0.33, stratify_by_column="label")
y = np.asanyarray(y) # To make it indexable for y[train]
test_size = np.ceil(0.33 * len(y))
train_size = len(y) - test_size
npt.assert_array_equal(np.unique(d1["train"]["label"]), np.unique(d1["test"]["label"]))
# checking classes proportion
p_train = np.bincount(np.unique(d1["train"]["label"], return_inverse=True)[1]) / float(
len(d1["train"]["label"])
)
p_test = np.bincount(np.unique(d1["test"]["label"], return_inverse=True)[1]) / float(
len(d1["test"]["label"])
)
npt.assert_array_almost_equal(p_train, p_test, 1)
assert len(d1["train"]["text"]) + len(d1["test"]["text"]) == y.size
assert len(d1["train"]["text"]) == train_size
assert len(d1["test"]["text"]) == test_size
def test_dataset_estimate_nbytes():
ds = Dataset.from_dict({"a": ["0" * 100] * 100})
assert 0.9 * ds._estimate_nbytes() < 100 * 100, "must be smaller than full dataset size"
ds = Dataset.from_dict({"a": ["0" * 100] * 100}).select([0])
assert 0.9 * ds._estimate_nbytes() < 100 * 100, "must be smaller than one chunk"
ds = Dataset.from_dict({"a": ["0" * 100] * 100})
ds = concatenate_datasets([ds] * 100)
assert 0.9 * ds._estimate_nbytes() < 100 * 100 * 100, "must be smaller than full dataset size"
assert 1.1 * ds._estimate_nbytes() > 100 * 100 * 100, "must be bigger than full dataset size"
ds = Dataset.from_dict({"a": ["0" * 100] * 100})
ds = concatenate_datasets([ds] * 100).select([0])
assert 0.9 * ds._estimate_nbytes() < 100 * 100, "must be smaller than one chunk"
def test_dataset_to_iterable_dataset(dataset: Dataset):
iterable_dataset = dataset.to_iterable_dataset()
assert isinstance(iterable_dataset, IterableDataset)
assert list(iterable_dataset) == list(dataset)
assert iterable_dataset.features == dataset.features
iterable_dataset = dataset.to_iterable_dataset(num_shards=3)
assert isinstance(iterable_dataset, IterableDataset)
assert list(iterable_dataset) == list(dataset)
assert iterable_dataset.features == dataset.features
assert iterable_dataset.n_shards == 3
with pytest.raises(ValueError):
dataset.to_iterable_dataset(num_shards=len(dataset) + 1)
with pytest.raises(NotImplementedError):
dataset.with_format("torch").to_iterable_dataset()
@require_pil
def test_dataset_format_with_unformatted_image():
import PIL
ds = Dataset.from_dict(
{"a": [np.arange(4 * 4 * 3).reshape(4, 4, 3)] * 10, "b": [[0, 1]] * 10},
Features({"a": Image(), "b": Sequence(Value("int64"))}),
)
ds.set_format("np", columns=["b"], output_all_columns=True)
assert isinstance(ds[0]["a"], PIL.Image.Image)
assert isinstance(ds[0]["b"], np.ndarray)
@pytest.mark.parametrize("batch_size", [1, 4])
@require_torch
def test_dataset_with_torch_dataloader(dataset, batch_size):
from torch.utils.data import DataLoader
from datasets import config
dataloader = DataLoader(dataset, batch_size=batch_size)
with patch.object(dataset, "_getitem", wraps=dataset._getitem) as mock_getitem:
out = list(dataloader)
getitem_call_count = mock_getitem.call_count
assert len(out) == len(dataset) // batch_size + int(len(dataset) % batch_size > 0)
# calling dataset[list_of_indices] is much more efficient than [dataset[idx] for idx in list of indices]
if config.TORCH_VERSION >= version.parse("1.13.0"):
assert getitem_call_count == len(dataset) // batch_size + int(len(dataset) % batch_size > 0)
@pytest.mark.parametrize("return_lazy_dict", [True, False, "mix"])
def test_map_cases(return_lazy_dict):
def f(x):
"""May return a mix of LazyDict and regular Dict"""
if x["a"] < 2:
x["a"] = -1
return dict(x) if return_lazy_dict is False else x
else:
return x if return_lazy_dict is True else {}
ds = Dataset.from_dict({"a": [0, 1, 2, 3]})
ds = ds.map(f)
outputs = ds[:]
assert outputs == {"a": [-1, -1, 2, 3]}
def f(x):
"""May return a mix of LazyDict and regular Dict, but sometimes with None values"""
if x["a"] < 2:
x["a"] = None
return dict(x) if return_lazy_dict is False else x
else:
return x if return_lazy_dict is True else {}
ds = Dataset.from_dict({"a": [0, 1, 2, 3]})
ds = ds.map(f)
outputs = ds[:]
assert outputs == {"a": [None, None, 2, 3]}
def f(x):
"""Return a LazyDict, but we remove a lazy column and add a new one"""
if x["a"] < 2:
x["b"] = -1
return x
else:
x["b"] = x["a"]
return x
ds = Dataset.from_dict({"a": [0, 1, 2, 3]})
ds = ds.map(f, remove_columns=["a"])
outputs = ds[:]
assert outputs == {"b": [-1, -1, 2, 3]}
# The formatted dataset version removes the lazy column from a different dictionary, hence it should be preserved in the output
ds = Dataset.from_dict({"a": [0, 1, 2, 3]})
ds = ds.with_format("numpy")
ds = ds.map(f, remove_columns=["a"])
ds = ds.with_format(None)
outputs = ds[:]
assert outputs == {"a": [0, 1, 2, 3], "b": [-1, -1, 2, 3]}
def f(x):
"""May return a mix of LazyDict and regular Dict, but we replace a lazy column"""
if x["a"] < 2:
x["a"] = -1
return dict(x) if return_lazy_dict is False else x
else:
x["a"] = x["a"]
return x if return_lazy_dict is True else {"a": x["a"]}
ds = Dataset.from_dict({"a": [0, 1, 2, 3]})
ds = ds.map(f, remove_columns=["a"])
outputs = ds[:]
assert outputs == ({"a": [-1, -1, 2, 3]} if return_lazy_dict is False else {})
def f(x):
"""May return a mix of LazyDict and regular Dict, but we modify a nested lazy column in-place"""
if x["a"]["b"] < 2:
x["a"]["c"] = -1
return dict(x) if return_lazy_dict is False else x
else:
x["a"]["c"] = x["a"]["b"]
return x if return_lazy_dict is True else {}
ds = Dataset.from_dict({"a": [{"b": 0}, {"b": 1}, {"b": 2}, {"b": 3}]})
ds = ds.map(f)
outputs = ds[:]
assert outputs == {"a": [{"b": 0, "c": -1}, {"b": 1, "c": -1}, {"b": 2, "c": 2}, {"b": 3, "c": 3}]}
def f(x):
"""May return a mix of LazyDict and regular Dict, but using an extension type"""
if x["a"][0][0] < 2:
x["a"] = [[-1]]
return dict(x) if return_lazy_dict is False else x
else:
return x if return_lazy_dict is True else {}
features = Features({"a": Array2D(shape=(1, 1), dtype="int32")})
ds = Dataset.from_dict({"a": [[[i]] for i in [0, 1, 2, 3]]}, features=features)
ds = ds.map(f)
outputs = ds[:]
assert outputs == {"a": [[[i]] for i in [-1, -1, 2, 3]]}
def f(x):
"""May return a mix of LazyDict and regular Dict, but using a nested extension type"""
if x["a"]["nested"][0][0] < 2:
x["a"] = {"nested": [[-1]]}
return dict(x) if return_lazy_dict is False else x
else:
return x if return_lazy_dict is True else {}
features = Features({"a": {"nested": Array2D(shape=(1, 1), dtype="int64")}})
ds = Dataset.from_dict({"a": [{"nested": [[i]]} for i in [0, 1, 2, 3]]}, features=features)
ds = ds.map(f)
outputs = ds[:]
assert outputs == {"a": [{"nested": [[i]]} for i in [-1, -1, 2, 3]]}
def test_dataset_getitem_raises():
ds = Dataset.from_dict({"a": [0, 1, 2, 3]})
with pytest.raises(TypeError):
ds[False]
with pytest.raises(TypeError):
ds._getitem(True)
| 0
|
hf_public_repos/datasets
|
hf_public_repos/datasets/tests/test_formatting.py
|
import datetime
from pathlib import Path
from unittest import TestCase
import numpy as np
import pandas as pd
import pyarrow as pa
import pytest
from datasets import Audio, Features, Image, IterableDataset
from datasets.formatting import NumpyFormatter, PandasFormatter, PythonFormatter, query_table
from datasets.formatting.formatting import (
LazyBatch,
LazyRow,
NumpyArrowExtractor,
PandasArrowExtractor,
PythonArrowExtractor,
)
from datasets.table import InMemoryTable
from .utils import require_jax, require_pil, require_sndfile, require_tf, require_torch
class AnyArray:
def __init__(self, data) -> None:
self.data = data
def __array__(self) -> np.ndarray:
return np.asarray(self.data)
def _gen_any_arrays():
for _ in range(10):
yield {"array": AnyArray(list(range(10)))}
@pytest.fixture
def any_arrays_dataset():
return IterableDataset.from_generator(_gen_any_arrays)
_COL_A = [0, 1, 2]
_COL_B = ["foo", "bar", "foobar"]
_COL_C = [[[1.0, 0.0, 0.0]] * 2, [[0.0, 1.0, 0.0]] * 2, [[0.0, 0.0, 1.0]] * 2]
_COL_D = [datetime.datetime(2023, 1, 1, 0, 0, tzinfo=datetime.timezone.utc)] * 3
_INDICES = [1, 0]
IMAGE_PATH_1 = Path(__file__).parent / "features" / "data" / "test_image_rgb.jpg"
IMAGE_PATH_2 = Path(__file__).parent / "features" / "data" / "test_image_rgba.png"
AUDIO_PATH_1 = Path(__file__).parent / "features" / "data" / "test_audio_44100.wav"
class ArrowExtractorTest(TestCase):
def _create_dummy_table(self):
return pa.Table.from_pydict({"a": _COL_A, "b": _COL_B, "c": _COL_C, "d": _COL_D})
def test_python_extractor(self):
pa_table = self._create_dummy_table()
extractor = PythonArrowExtractor()
row = extractor.extract_row(pa_table)
self.assertEqual(row, {"a": _COL_A[0], "b": _COL_B[0], "c": _COL_C[0], "d": _COL_D[0]})
col = extractor.extract_column(pa_table)
self.assertEqual(col, _COL_A)
batch = extractor.extract_batch(pa_table)
self.assertEqual(batch, {"a": _COL_A, "b": _COL_B, "c": _COL_C, "d": _COL_D})
def test_numpy_extractor(self):
pa_table = self._create_dummy_table().drop(["c", "d"])
extractor = NumpyArrowExtractor()
row = extractor.extract_row(pa_table)
np.testing.assert_equal(row, {"a": _COL_A[0], "b": _COL_B[0]})
col = extractor.extract_column(pa_table)
np.testing.assert_equal(col, np.array(_COL_A))
batch = extractor.extract_batch(pa_table)
np.testing.assert_equal(batch, {"a": np.array(_COL_A), "b": np.array(_COL_B)})
def test_numpy_extractor_nested(self):
pa_table = self._create_dummy_table().drop(["a", "b", "d"])
extractor = NumpyArrowExtractor()
row = extractor.extract_row(pa_table)
self.assertEqual(row["c"][0].dtype, np.float64)
self.assertEqual(row["c"].dtype, object)
col = extractor.extract_column(pa_table)
self.assertEqual(col[0][0].dtype, np.float64)
self.assertEqual(col[0].dtype, object)
self.assertEqual(col.dtype, object)
batch = extractor.extract_batch(pa_table)
self.assertEqual(batch["c"][0][0].dtype, np.float64)
self.assertEqual(batch["c"][0].dtype, object)
self.assertEqual(batch["c"].dtype, object)
def test_numpy_extractor_temporal(self):
pa_table = self._create_dummy_table().drop(["a", "b", "c"])
extractor = NumpyArrowExtractor()
row = extractor.extract_row(pa_table)
self.assertTrue(np.issubdtype(row["d"].dtype, np.datetime64))
col = extractor.extract_column(pa_table)
self.assertTrue(np.issubdtype(col[0].dtype, np.datetime64))
self.assertTrue(np.issubdtype(col.dtype, np.datetime64))
batch = extractor.extract_batch(pa_table)
self.assertTrue(np.issubdtype(batch["d"][0].dtype, np.datetime64))
self.assertTrue(np.issubdtype(batch["d"].dtype, np.datetime64))
def test_pandas_extractor(self):
pa_table = self._create_dummy_table()
extractor = PandasArrowExtractor()
row = extractor.extract_row(pa_table)
self.assertIsInstance(row, pd.DataFrame)
pd.testing.assert_series_equal(row["a"], pd.Series(_COL_A, name="a")[:1])
pd.testing.assert_series_equal(row["b"], pd.Series(_COL_B, name="b")[:1])
col = extractor.extract_column(pa_table)
pd.testing.assert_series_equal(col, pd.Series(_COL_A, name="a"))
batch = extractor.extract_batch(pa_table)
self.assertIsInstance(batch, pd.DataFrame)
pd.testing.assert_series_equal(batch["a"], pd.Series(_COL_A, name="a"))
pd.testing.assert_series_equal(batch["b"], pd.Series(_COL_B, name="b"))
def test_pandas_extractor_nested(self):
pa_table = self._create_dummy_table().drop(["a", "b", "d"])
extractor = PandasArrowExtractor()
row = extractor.extract_row(pa_table)
self.assertEqual(row["c"][0][0].dtype, np.float64)
self.assertEqual(row["c"].dtype, object)
col = extractor.extract_column(pa_table)
self.assertEqual(col[0][0].dtype, np.float64)
self.assertEqual(col[0].dtype, object)
self.assertEqual(col.dtype, object)
batch = extractor.extract_batch(pa_table)
self.assertEqual(batch["c"][0][0].dtype, np.float64)
self.assertEqual(batch["c"][0].dtype, object)
self.assertEqual(batch["c"].dtype, object)
def test_pandas_extractor_temporal(self):
pa_table = self._create_dummy_table().drop(["a", "b", "c"])
extractor = PandasArrowExtractor()
row = extractor.extract_row(pa_table)
self.assertTrue(pd.api.types.is_datetime64_any_dtype(row["d"].dtype))
col = extractor.extract_column(pa_table)
self.assertTrue(isinstance(col[0], datetime.datetime))
self.assertTrue(pd.api.types.is_datetime64_any_dtype(col.dtype))
batch = extractor.extract_batch(pa_table)
self.assertTrue(isinstance(batch["d"][0], datetime.datetime))
self.assertTrue(pd.api.types.is_datetime64_any_dtype(batch["d"].dtype))
class LazyDictTest(TestCase):
def _create_dummy_table(self):
return pa.Table.from_pydict({"a": _COL_A, "b": _COL_B, "c": _COL_C})
def _create_dummy_formatter(self):
return PythonFormatter(lazy=True)
def test_lazy_dict_copy(self):
pa_table = self._create_dummy_table()
formatter = self._create_dummy_formatter()
lazy_batch = formatter.format_batch(pa_table)
lazy_batch_copy = lazy_batch.copy()
self.assertEqual(type(lazy_batch), type(lazy_batch_copy))
self.assertEqual(lazy_batch.items(), lazy_batch_copy.items())
lazy_batch["d"] = [1, 2, 3]
self.assertNotEqual(lazy_batch.items(), lazy_batch_copy.items())
class FormatterTest(TestCase):
def _create_dummy_table(self):
return pa.Table.from_pydict({"a": _COL_A, "b": _COL_B, "c": _COL_C})
def test_python_formatter(self):
pa_table = self._create_dummy_table()
formatter = PythonFormatter()
row = formatter.format_row(pa_table)
self.assertEqual(row, {"a": _COL_A[0], "b": _COL_B[0], "c": _COL_C[0]})
col = formatter.format_column(pa_table)
self.assertEqual(col, _COL_A)
batch = formatter.format_batch(pa_table)
self.assertEqual(batch, {"a": _COL_A, "b": _COL_B, "c": _COL_C})
def test_python_formatter_lazy(self):
pa_table = self._create_dummy_table()
formatter = PythonFormatter(lazy=True)
row = formatter.format_row(pa_table)
self.assertIsInstance(row, LazyRow)
self.assertEqual(row["a"], _COL_A[0])
self.assertEqual(row["b"], _COL_B[0])
self.assertEqual(row["c"], _COL_C[0])
batch = formatter.format_batch(pa_table)
self.assertIsInstance(batch, LazyBatch)
self.assertEqual(batch["a"], _COL_A)
self.assertEqual(batch["b"], _COL_B)
self.assertEqual(batch["c"], _COL_C)
def test_numpy_formatter(self):
pa_table = self._create_dummy_table()
formatter = NumpyFormatter()
row = formatter.format_row(pa_table)
np.testing.assert_equal(row, {"a": _COL_A[0], "b": _COL_B[0], "c": np.array(_COL_C[0])})
col = formatter.format_column(pa_table)
np.testing.assert_equal(col, np.array(_COL_A))
batch = formatter.format_batch(pa_table)
np.testing.assert_equal(batch, {"a": np.array(_COL_A), "b": np.array(_COL_B), "c": np.array(_COL_C)})
assert batch["c"].shape == np.array(_COL_C).shape
def test_numpy_formatter_np_array_kwargs(self):
pa_table = self._create_dummy_table().drop(["b"])
formatter = NumpyFormatter(dtype=np.float16)
row = formatter.format_row(pa_table)
self.assertEqual(row["c"].dtype, np.dtype(np.float16))
col = formatter.format_column(pa_table)
self.assertEqual(col.dtype, np.float16)
batch = formatter.format_batch(pa_table)
self.assertEqual(batch["a"].dtype, np.dtype(np.float16))
self.assertEqual(batch["c"].dtype, np.dtype(np.float16))
@require_pil
def test_numpy_formatter_image(self):
# same dimensions
pa_table = pa.table({"image": [{"bytes": None, "path": str(IMAGE_PATH_1)}] * 2})
formatter = NumpyFormatter(features=Features({"image": Image()}))
row = formatter.format_row(pa_table)
self.assertEqual(row["image"].dtype, np.uint8)
self.assertEqual(row["image"].shape, (480, 640, 3))
col = formatter.format_column(pa_table)
self.assertEqual(col.dtype, np.uint8)
self.assertEqual(col.shape, (2, 480, 640, 3))
batch = formatter.format_batch(pa_table)
self.assertEqual(batch["image"].dtype, np.uint8)
self.assertEqual(batch["image"].shape, (2, 480, 640, 3))
# different dimensions
pa_table = pa.table(
{"image": [{"bytes": None, "path": str(IMAGE_PATH_1)}, {"bytes": None, "path": str(IMAGE_PATH_2)}]}
)
formatter = NumpyFormatter(features=Features({"image": Image()}))
row = formatter.format_row(pa_table)
self.assertEqual(row["image"].dtype, np.uint8)
self.assertEqual(row["image"].shape, (480, 640, 3))
col = formatter.format_column(pa_table)
self.assertIsInstance(col, np.ndarray)
self.assertEqual(col.dtype, object)
self.assertEqual(col[0].dtype, np.uint8)
self.assertEqual(col[0].shape, (480, 640, 3))
batch = formatter.format_batch(pa_table)
self.assertIsInstance(batch["image"], np.ndarray)
self.assertEqual(batch["image"].dtype, object)
self.assertEqual(batch["image"][0].dtype, np.uint8)
self.assertEqual(batch["image"][0].shape, (480, 640, 3))
@require_sndfile
def test_numpy_formatter_audio(self):
pa_table = pa.table({"audio": [{"bytes": None, "path": str(AUDIO_PATH_1)}]})
formatter = NumpyFormatter(features=Features({"audio": Audio()}))
row = formatter.format_row(pa_table)
self.assertEqual(row["audio"]["array"].dtype, np.dtype(np.float32))
col = formatter.format_column(pa_table)
self.assertEqual(col[0]["array"].dtype, np.float32)
batch = formatter.format_batch(pa_table)
self.assertEqual(batch["audio"][0]["array"].dtype, np.dtype(np.float32))
def test_pandas_formatter(self):
pa_table = self._create_dummy_table()
formatter = PandasFormatter()
row = formatter.format_row(pa_table)
self.assertIsInstance(row, pd.DataFrame)
pd.testing.assert_series_equal(row["a"], pd.Series(_COL_A, name="a")[:1])
pd.testing.assert_series_equal(row["b"], pd.Series(_COL_B, name="b")[:1])
col = formatter.format_column(pa_table)
pd.testing.assert_series_equal(col, pd.Series(_COL_A, name="a"))
batch = formatter.format_batch(pa_table)
self.assertIsInstance(batch, pd.DataFrame)
pd.testing.assert_series_equal(batch["a"], pd.Series(_COL_A, name="a"))
pd.testing.assert_series_equal(batch["b"], pd.Series(_COL_B, name="b"))
@require_torch
def test_torch_formatter(self):
import torch
from datasets.formatting import TorchFormatter
pa_table = self._create_dummy_table()
formatter = TorchFormatter()
row = formatter.format_row(pa_table)
torch.testing.assert_close(row["a"], torch.tensor(_COL_A, dtype=torch.int64)[0])
assert row["b"] == _COL_B[0]
torch.testing.assert_close(row["c"], torch.tensor(_COL_C, dtype=torch.float32)[0])
col = formatter.format_column(pa_table)
torch.testing.assert_close(col, torch.tensor(_COL_A, dtype=torch.int64))
batch = formatter.format_batch(pa_table)
torch.testing.assert_close(batch["a"], torch.tensor(_COL_A, dtype=torch.int64))
assert batch["b"] == _COL_B
torch.testing.assert_close(batch["c"], torch.tensor(_COL_C, dtype=torch.float32))
assert batch["c"].shape == np.array(_COL_C).shape
@require_torch
def test_torch_formatter_torch_tensor_kwargs(self):
import torch
from datasets.formatting import TorchFormatter
pa_table = self._create_dummy_table().drop(["b"])
formatter = TorchFormatter(dtype=torch.float16)
row = formatter.format_row(pa_table)
self.assertEqual(row["c"].dtype, torch.float16)
col = formatter.format_column(pa_table)
self.assertEqual(col.dtype, torch.float16)
batch = formatter.format_batch(pa_table)
self.assertEqual(batch["a"].dtype, torch.float16)
self.assertEqual(batch["c"].dtype, torch.float16)
@require_torch
@require_pil
def test_torch_formatter_image(self):
import torch
from datasets.formatting import TorchFormatter
# same dimensions
pa_table = pa.table({"image": [{"bytes": None, "path": str(IMAGE_PATH_1)}] * 2})
formatter = TorchFormatter(features=Features({"image": Image()}))
row = formatter.format_row(pa_table)
self.assertEqual(row["image"].dtype, torch.uint8)
self.assertEqual(row["image"].shape, (480, 640, 3))
col = formatter.format_column(pa_table)
self.assertEqual(col.dtype, torch.uint8)
self.assertEqual(col.shape, (2, 480, 640, 3))
batch = formatter.format_batch(pa_table)
self.assertEqual(batch["image"].dtype, torch.uint8)
self.assertEqual(batch["image"].shape, (2, 480, 640, 3))
# different dimensions
pa_table = pa.table(
{"image": [{"bytes": None, "path": str(IMAGE_PATH_1)}, {"bytes": None, "path": str(IMAGE_PATH_2)}]}
)
formatter = TorchFormatter(features=Features({"image": Image()}))
row = formatter.format_row(pa_table)
self.assertEqual(row["image"].dtype, torch.uint8)
self.assertEqual(row["image"].shape, (480, 640, 3))
col = formatter.format_column(pa_table)
self.assertIsInstance(col, list)
self.assertEqual(col[0].dtype, torch.uint8)
self.assertEqual(col[0].shape, (480, 640, 3))
batch = formatter.format_batch(pa_table)
self.assertIsInstance(batch["image"], list)
self.assertEqual(batch["image"][0].dtype, torch.uint8)
self.assertEqual(batch["image"][0].shape, (480, 640, 3))
@require_torch
@require_sndfile
def test_torch_formatter_audio(self):
import torch
from datasets.formatting import TorchFormatter
pa_table = pa.table({"audio": [{"bytes": None, "path": str(AUDIO_PATH_1)}]})
formatter = TorchFormatter(features=Features({"audio": Audio()}))
row = formatter.format_row(pa_table)
self.assertEqual(row["audio"]["array"].dtype, torch.float32)
col = formatter.format_column(pa_table)
self.assertEqual(col[0]["array"].dtype, torch.float32)
batch = formatter.format_batch(pa_table)
self.assertEqual(batch["audio"][0]["array"].dtype, torch.float32)
@require_tf
def test_tf_formatter(self):
import tensorflow as tf
from datasets.formatting import TFFormatter
pa_table = self._create_dummy_table()
formatter = TFFormatter()
row = formatter.format_row(pa_table)
tf.debugging.assert_equal(row["a"], tf.convert_to_tensor(_COL_A, dtype=tf.int64)[0])
tf.debugging.assert_equal(row["b"], tf.convert_to_tensor(_COL_B, dtype=tf.string)[0])
tf.debugging.assert_equal(row["c"], tf.convert_to_tensor(_COL_C, dtype=tf.float32)[0])
col = formatter.format_column(pa_table)
tf.debugging.assert_equal(col, tf.ragged.constant(_COL_A, dtype=tf.int64))
batch = formatter.format_batch(pa_table)
tf.debugging.assert_equal(batch["a"], tf.convert_to_tensor(_COL_A, dtype=tf.int64))
tf.debugging.assert_equal(batch["b"], tf.convert_to_tensor(_COL_B, dtype=tf.string))
self.assertIsInstance(batch["c"], tf.Tensor)
self.assertEqual(batch["c"].dtype, tf.float32)
tf.debugging.assert_equal(
batch["c"].shape.as_list(), tf.convert_to_tensor(_COL_C, dtype=tf.float32).shape.as_list()
)
tf.debugging.assert_equal(tf.convert_to_tensor(batch["c"]), tf.convert_to_tensor(_COL_C, dtype=tf.float32))
@require_tf
def test_tf_formatter_tf_tensor_kwargs(self):
import tensorflow as tf
from datasets.formatting import TFFormatter
pa_table = self._create_dummy_table().drop(["b"])
formatter = TFFormatter(dtype=tf.float16)
row = formatter.format_row(pa_table)
self.assertEqual(row["c"].dtype, tf.float16)
col = formatter.format_column(pa_table)
self.assertEqual(col.dtype, tf.float16)
batch = formatter.format_batch(pa_table)
self.assertEqual(batch["a"].dtype, tf.float16)
self.assertEqual(batch["c"].dtype, tf.float16)
@require_tf
@require_pil
def test_tf_formatter_image(self):
import tensorflow as tf
from datasets.formatting import TFFormatter
# same dimensions
pa_table = pa.table({"image": [{"bytes": None, "path": str(IMAGE_PATH_1)}] * 2})
formatter = TFFormatter(features=Features({"image": Image()}))
row = formatter.format_row(pa_table)
self.assertEqual(row["image"].dtype, tf.uint8)
self.assertEqual(row["image"].shape, (480, 640, 3))
col = formatter.format_column(pa_table)
self.assertEqual(col.dtype, tf.uint8)
self.assertEqual(col.shape, (2, 480, 640, 3))
batch = formatter.format_batch(pa_table)
self.assertEqual(batch["image"][0].dtype, tf.uint8)
self.assertEqual(batch["image"].shape, (2, 480, 640, 3))
# different dimensions
pa_table = pa.table(
{"image": [{"bytes": None, "path": str(IMAGE_PATH_1)}, {"bytes": None, "path": str(IMAGE_PATH_2)}]}
)
formatter = TFFormatter(features=Features({"image": Image()}))
row = formatter.format_row(pa_table)
self.assertEqual(row["image"].dtype, tf.uint8)
self.assertEqual(row["image"].shape, (480, 640, 3))
col = formatter.format_column(pa_table)
self.assertIsInstance(col, list)
self.assertEqual(col[0].dtype, tf.uint8)
self.assertEqual(col[0].shape, (480, 640, 3))
batch = formatter.format_batch(pa_table)
self.assertIsInstance(batch["image"], list)
self.assertEqual(batch["image"][0].dtype, tf.uint8)
self.assertEqual(batch["image"][0].shape, (480, 640, 3))
@require_tf
@require_sndfile
def test_tf_formatter_audio(self):
import tensorflow as tf
from datasets.formatting import TFFormatter
pa_table = pa.table({"audio": [{"bytes": None, "path": str(AUDIO_PATH_1)}]})
formatter = TFFormatter(features=Features({"audio": Audio()}))
row = formatter.format_row(pa_table)
self.assertEqual(row["audio"]["array"].dtype, tf.float32)
col = formatter.format_column(pa_table)
self.assertEqual(col[0]["array"].dtype, tf.float32)
batch = formatter.format_batch(pa_table)
self.assertEqual(batch["audio"][0]["array"].dtype, tf.float32)
@require_jax
def test_jax_formatter(self):
import jax
import jax.numpy as jnp
from datasets.formatting import JaxFormatter
pa_table = self._create_dummy_table()
formatter = JaxFormatter()
row = formatter.format_row(pa_table)
jnp.allclose(row["a"], jnp.array(_COL_A, dtype=jnp.int64 if jax.config.jax_enable_x64 else jnp.int32)[0])
assert row["b"] == _COL_B[0]
jnp.allclose(row["c"], jnp.array(_COL_C, dtype=jnp.float32)[0])
col = formatter.format_column(pa_table)
jnp.allclose(col, jnp.array(_COL_A, dtype=jnp.int64 if jax.config.jax_enable_x64 else jnp.int32))
batch = formatter.format_batch(pa_table)
jnp.allclose(batch["a"], jnp.array(_COL_A, dtype=jnp.int64 if jax.config.jax_enable_x64 else jnp.int32))
assert batch["b"] == _COL_B
jnp.allclose(batch["c"], jnp.array(_COL_C, dtype=jnp.float32))
assert batch["c"].shape == np.array(_COL_C).shape
@require_jax
def test_jax_formatter_jnp_array_kwargs(self):
import jax.numpy as jnp
from datasets.formatting import JaxFormatter
pa_table = self._create_dummy_table().drop(["b"])
formatter = JaxFormatter(dtype=jnp.float16)
row = formatter.format_row(pa_table)
self.assertEqual(row["c"].dtype, jnp.float16)
col = formatter.format_column(pa_table)
self.assertEqual(col.dtype, jnp.float16)
batch = formatter.format_batch(pa_table)
self.assertEqual(batch["a"].dtype, jnp.float16)
self.assertEqual(batch["c"].dtype, jnp.float16)
@require_jax
@require_pil
def test_jax_formatter_image(self):
import jax.numpy as jnp
from datasets.formatting import JaxFormatter
# same dimensions
pa_table = pa.table({"image": [{"bytes": None, "path": str(IMAGE_PATH_1)}] * 2})
formatter = JaxFormatter(features=Features({"image": Image()}))
row = formatter.format_row(pa_table)
self.assertEqual(row["image"].dtype, jnp.uint8)
self.assertEqual(row["image"].shape, (480, 640, 3))
col = formatter.format_column(pa_table)
self.assertEqual(col.dtype, jnp.uint8)
self.assertEqual(col.shape, (2, 480, 640, 3))
batch = formatter.format_batch(pa_table)
self.assertEqual(batch["image"].dtype, jnp.uint8)
self.assertEqual(batch["image"].shape, (2, 480, 640, 3))
# different dimensions
pa_table = pa.table(
{"image": [{"bytes": None, "path": str(IMAGE_PATH_1)}, {"bytes": None, "path": str(IMAGE_PATH_2)}]}
)
formatter = JaxFormatter(features=Features({"image": Image()}))
row = formatter.format_row(pa_table)
self.assertEqual(row["image"].dtype, jnp.uint8)
self.assertEqual(row["image"].shape, (480, 640, 3))
col = formatter.format_column(pa_table)
self.assertIsInstance(col, list)
self.assertEqual(col[0].dtype, jnp.uint8)
self.assertEqual(col[0].shape, (480, 640, 3))
batch = formatter.format_batch(pa_table)
self.assertIsInstance(batch["image"], list)
self.assertEqual(batch["image"][0].dtype, jnp.uint8)
self.assertEqual(batch["image"][0].shape, (480, 640, 3))
@require_jax
@require_sndfile
def test_jax_formatter_audio(self):
import jax.numpy as jnp
from datasets.formatting import JaxFormatter
pa_table = pa.table({"audio": [{"bytes": None, "path": str(AUDIO_PATH_1)}]})
formatter = JaxFormatter(features=Features({"audio": Audio()}))
row = formatter.format_row(pa_table)
self.assertEqual(row["audio"]["array"].dtype, jnp.float32)
col = formatter.format_column(pa_table)
self.assertEqual(col[0]["array"].dtype, jnp.float32)
batch = formatter.format_batch(pa_table)
self.assertEqual(batch["audio"][0]["array"].dtype, jnp.float32)
@require_jax
def test_jax_formatter_device(self):
import jax
from datasets.formatting import JaxFormatter
pa_table = self._create_dummy_table()
device = jax.devices()[0]
formatter = JaxFormatter(device=str(device))
row = formatter.format_row(pa_table)
assert row["a"].device() == device
assert row["c"].device() == device
col = formatter.format_column(pa_table)
assert col.device() == device
batch = formatter.format_batch(pa_table)
assert batch["a"].device() == device
assert batch["c"].device() == device
class QueryTest(TestCase):
def _create_dummy_table(self):
return pa.Table.from_pydict({"a": _COL_A, "b": _COL_B, "c": _COL_C})
def _create_dummy_arrow_indices(self):
return pa.Table.from_arrays([pa.array(_INDICES, type=pa.uint64())], names=["indices"])
def assertTableEqual(self, first: pa.Table, second: pa.Table):
self.assertEqual(first.schema, second.schema)
for first_array, second_array in zip(first, second):
self.assertEqual(first_array, second_array)
self.assertEqual(first, second)
def test_query_table_int(self):
pa_table = self._create_dummy_table()
table = InMemoryTable(pa_table)
n = pa_table.num_rows
# classical usage
subtable = query_table(table, 0)
self.assertTableEqual(subtable, pa.Table.from_pydict({"a": _COL_A[:1], "b": _COL_B[:1], "c": _COL_C[:1]}))
subtable = query_table(table, 1)
self.assertTableEqual(subtable, pa.Table.from_pydict({"a": _COL_A[1:2], "b": _COL_B[1:2], "c": _COL_C[1:2]}))
subtable = query_table(table, -1)
self.assertTableEqual(subtable, pa.Table.from_pydict({"a": _COL_A[-1:], "b": _COL_B[-1:], "c": _COL_C[-1:]}))
# raise an IndexError
with self.assertRaises(IndexError):
query_table(table, n)
with self.assertRaises(IndexError):
query_table(table, -(n + 1))
# with indices
indices = InMemoryTable(self._create_dummy_arrow_indices())
subtable = query_table(table, 0, indices=indices)
self.assertTableEqual(
subtable,
pa.Table.from_pydict({"a": [_COL_A[_INDICES[0]]], "b": [_COL_B[_INDICES[0]]], "c": [_COL_C[_INDICES[0]]]}),
)
with self.assertRaises(IndexError):
assert len(indices) < n
query_table(table, len(indices), indices=indices)
def test_query_table_slice(self):
pa_table = self._create_dummy_table()
table = InMemoryTable(pa_table)
n = pa_table.num_rows
# classical usage
subtable = query_table(table, slice(0, 1))
self.assertTableEqual(subtable, pa.Table.from_pydict({"a": _COL_A[:1], "b": _COL_B[:1], "c": _COL_C[:1]}))
subtable = query_table(table, slice(1, 2))
self.assertTableEqual(subtable, pa.Table.from_pydict({"a": _COL_A[1:2], "b": _COL_B[1:2], "c": _COL_C[1:2]}))
subtable = query_table(table, slice(-2, -1))
self.assertTableEqual(
subtable, pa.Table.from_pydict({"a": _COL_A[-2:-1], "b": _COL_B[-2:-1], "c": _COL_C[-2:-1]})
)
# usage with None
subtable = query_table(table, slice(-1, None))
self.assertTableEqual(subtable, pa.Table.from_pydict({"a": _COL_A[-1:], "b": _COL_B[-1:], "c": _COL_C[-1:]}))
subtable = query_table(table, slice(None, n + 1))
self.assertTableEqual(
subtable, pa.Table.from_pydict({"a": _COL_A[: n + 1], "b": _COL_B[: n + 1], "c": _COL_C[: n + 1]})
)
self.assertTableEqual(subtable, pa.Table.from_pydict({"a": _COL_A, "b": _COL_B, "c": _COL_C}))
subtable = query_table(table, slice(-(n + 1), None))
self.assertTableEqual(
subtable, pa.Table.from_pydict({"a": _COL_A[-(n + 1) :], "b": _COL_B[-(n + 1) :], "c": _COL_C[-(n + 1) :]})
)
self.assertTableEqual(subtable, pa.Table.from_pydict({"a": _COL_A, "b": _COL_B, "c": _COL_C}))
# usage with step
subtable = query_table(table, slice(None, None, 2))
self.assertTableEqual(subtable, pa.Table.from_pydict({"a": _COL_A[::2], "b": _COL_B[::2], "c": _COL_C[::2]}))
# empty ouput but no errors
subtable = query_table(table, slice(-1, 0)) # usage with both negative and positive idx
assert len(_COL_A[-1:0]) == 0
self.assertTableEqual(subtable, pa_table.slice(0, 0))
subtable = query_table(table, slice(2, 1))
assert len(_COL_A[2:1]) == 0
self.assertTableEqual(subtable, pa_table.slice(0, 0))
subtable = query_table(table, slice(n, n))
assert len(_COL_A[n:n]) == 0
self.assertTableEqual(subtable, pa_table.slice(0, 0))
subtable = query_table(table, slice(n, n + 1))
assert len(_COL_A[n : n + 1]) == 0
self.assertTableEqual(subtable, pa_table.slice(0, 0))
# it's not possible to get an error with a slice
# with indices
indices = InMemoryTable(self._create_dummy_arrow_indices())
subtable = query_table(table, slice(0, 1), indices=indices)
self.assertTableEqual(
subtable,
pa.Table.from_pydict({"a": [_COL_A[_INDICES[0]]], "b": [_COL_B[_INDICES[0]]], "c": [_COL_C[_INDICES[0]]]}),
)
subtable = query_table(table, slice(n - 1, n), indices=indices)
assert len(indices.column(0).to_pylist()[n - 1 : n]) == 0
self.assertTableEqual(subtable, pa_table.slice(0, 0))
def test_query_table_range(self):
pa_table = self._create_dummy_table()
table = InMemoryTable(pa_table)
n = pa_table.num_rows
np_A, np_B, np_C = np.array(_COL_A, dtype=np.int64), np.array(_COL_B), np.array(_COL_C)
# classical usage
subtable = query_table(table, range(0, 1))
self.assertTableEqual(
subtable,
pa.Table.from_pydict({"a": np_A[range(0, 1)], "b": np_B[range(0, 1)], "c": np_C[range(0, 1)].tolist()}),
)
subtable = query_table(table, range(1, 2))
self.assertTableEqual(
subtable,
pa.Table.from_pydict({"a": np_A[range(1, 2)], "b": np_B[range(1, 2)], "c": np_C[range(1, 2)].tolist()}),
)
subtable = query_table(table, range(-2, -1))
self.assertTableEqual(
subtable,
pa.Table.from_pydict(
{"a": np_A[range(-2, -1)], "b": np_B[range(-2, -1)], "c": np_C[range(-2, -1)].tolist()}
),
)
# usage with both negative and positive idx
subtable = query_table(table, range(-1, 0))
self.assertTableEqual(
subtable,
pa.Table.from_pydict({"a": np_A[range(-1, 0)], "b": np_B[range(-1, 0)], "c": np_C[range(-1, 0)].tolist()}),
)
subtable = query_table(table, range(-1, n))
self.assertTableEqual(
subtable,
pa.Table.from_pydict({"a": np_A[range(-1, n)], "b": np_B[range(-1, n)], "c": np_C[range(-1, n)].tolist()}),
)
# usage with step
subtable = query_table(table, range(0, n, 2))
self.assertTableEqual(
subtable,
pa.Table.from_pydict(
{"a": np_A[range(0, n, 2)], "b": np_B[range(0, n, 2)], "c": np_C[range(0, n, 2)].tolist()}
),
)
subtable = query_table(table, range(0, n + 1, 2 * n))
self.assertTableEqual(
subtable,
pa.Table.from_pydict(
{
"a": np_A[range(0, n + 1, 2 * n)],
"b": np_B[range(0, n + 1, 2 * n)],
"c": np_C[range(0, n + 1, 2 * n)].tolist(),
}
),
)
# empty ouput but no errors
subtable = query_table(table, range(2, 1))
assert len(np_A[range(2, 1)]) == 0
self.assertTableEqual(subtable, pa.Table.from_batches([], schema=pa_table.schema))
subtable = query_table(table, range(n, n))
assert len(np_A[range(n, n)]) == 0
self.assertTableEqual(subtable, pa.Table.from_batches([], schema=pa_table.schema))
# raise an IndexError
with self.assertRaises(IndexError):
with self.assertRaises(IndexError):
np_A[range(0, n + 1)]
query_table(table, range(0, n + 1))
with self.assertRaises(IndexError):
with self.assertRaises(IndexError):
np_A[range(-(n + 1), -1)]
query_table(table, range(-(n + 1), -1))
with self.assertRaises(IndexError):
with self.assertRaises(IndexError):
np_A[range(n, n + 1)]
query_table(table, range(n, n + 1))
# with indices
indices = InMemoryTable(self._create_dummy_arrow_indices())
subtable = query_table(table, range(0, 1), indices=indices)
self.assertTableEqual(
subtable,
pa.Table.from_pydict({"a": [_COL_A[_INDICES[0]]], "b": [_COL_B[_INDICES[0]]], "c": [_COL_C[_INDICES[0]]]}),
)
with self.assertRaises(IndexError):
assert len(indices) < n
query_table(table, range(len(indices), len(indices) + 1), indices=indices)
def test_query_table_str(self):
pa_table = self._create_dummy_table()
table = InMemoryTable(pa_table)
subtable = query_table(table, "a")
self.assertTableEqual(subtable, pa.Table.from_pydict({"a": _COL_A}))
with self.assertRaises(KeyError):
query_table(table, "z")
indices = InMemoryTable(self._create_dummy_arrow_indices())
subtable = query_table(table, "a", indices=indices)
self.assertTableEqual(subtable, pa.Table.from_pydict({"a": [_COL_A[i] for i in _INDICES]}))
def test_query_table_iterable(self):
pa_table = self._create_dummy_table()
table = InMemoryTable(pa_table)
n = pa_table.num_rows
np_A, np_B, np_C = np.array(_COL_A, dtype=np.int64), np.array(_COL_B), np.array(_COL_C)
# classical usage
subtable = query_table(table, [0])
self.assertTableEqual(
subtable, pa.Table.from_pydict({"a": np_A[[0]], "b": np_B[[0]], "c": np_C[[0]].tolist()})
)
subtable = query_table(table, [1])
self.assertTableEqual(
subtable, pa.Table.from_pydict({"a": np_A[[1]], "b": np_B[[1]], "c": np_C[[1]].tolist()})
)
subtable = query_table(table, [-1])
self.assertTableEqual(
subtable, pa.Table.from_pydict({"a": np_A[[-1]], "b": np_B[[-1]], "c": np_C[[-1]].tolist()})
)
subtable = query_table(table, [0, -1, 1])
self.assertTableEqual(
subtable,
pa.Table.from_pydict({"a": np_A[[0, -1, 1]], "b": np_B[[0, -1, 1]], "c": np_C[[0, -1, 1]].tolist()}),
)
# numpy iterable
subtable = query_table(table, np.array([0, -1, 1]))
self.assertTableEqual(
subtable,
pa.Table.from_pydict({"a": np_A[[0, -1, 1]], "b": np_B[[0, -1, 1]], "c": np_C[[0, -1, 1]].tolist()}),
)
# empty ouput but no errors
subtable = query_table(table, [])
assert len(np_A[[]]) == 0
self.assertTableEqual(subtable, pa.Table.from_batches([], schema=pa_table.schema))
# raise an IndexError
with self.assertRaises(IndexError):
with self.assertRaises(IndexError):
np_A[[n]]
query_table(table, [n])
with self.assertRaises(IndexError):
with self.assertRaises(IndexError):
np_A[[-(n + 1)]]
query_table(table, [-(n + 1)])
# with indices
indices = InMemoryTable(self._create_dummy_arrow_indices())
subtable = query_table(table, [0], indices=indices)
self.assertTableEqual(
subtable,
pa.Table.from_pydict({"a": [_COL_A[_INDICES[0]]], "b": [_COL_B[_INDICES[0]]], "c": [_COL_C[_INDICES[0]]]}),
)
with self.assertRaises(IndexError):
assert len(indices) < n
query_table(table, [len(indices)], indices=indices)
def test_query_table_invalid_key_type(self):
pa_table = self._create_dummy_table()
table = InMemoryTable(pa_table)
with self.assertRaises(TypeError):
query_table(table, 0.0)
with self.assertRaises(TypeError):
query_table(table, [0, "a"])
with self.assertRaises(TypeError):
query_table(table, int)
with self.assertRaises(TypeError):
def iter_to_inf(start=0):
while True:
yield start
start += 1
query_table(table, iter_to_inf())
@pytest.fixture(scope="session")
def arrow_table():
return pa.Table.from_pydict({"col_int": [0, 1, 2], "col_float": [0.0, 1.0, 2.0]})
@require_tf
@pytest.mark.parametrize(
"cast_schema",
[
None,
[("col_int", pa.int64()), ("col_float", pa.float64())],
[("col_int", pa.int32()), ("col_float", pa.float64())],
[("col_int", pa.int64()), ("col_float", pa.float32())],
],
)
def test_tf_formatter_sets_default_dtypes(cast_schema, arrow_table):
import tensorflow as tf
from datasets.formatting import TFFormatter
if cast_schema:
arrow_table = arrow_table.cast(pa.schema(cast_schema))
arrow_table_dict = arrow_table.to_pydict()
list_int = arrow_table_dict["col_int"]
list_float = arrow_table_dict["col_float"]
formatter = TFFormatter()
row = formatter.format_row(arrow_table)
tf.debugging.assert_equal(row["col_int"], tf.ragged.constant(list_int, dtype=tf.int64)[0])
tf.debugging.assert_equal(row["col_float"], tf.ragged.constant(list_float, dtype=tf.float32)[0])
col = formatter.format_column(arrow_table)
tf.debugging.assert_equal(col, tf.ragged.constant(list_int, dtype=tf.int64))
batch = formatter.format_batch(arrow_table)
tf.debugging.assert_equal(batch["col_int"], tf.ragged.constant(list_int, dtype=tf.int64))
tf.debugging.assert_equal(batch["col_float"], tf.ragged.constant(list_float, dtype=tf.float32))
@require_torch
@pytest.mark.parametrize(
"cast_schema",
[
None,
[("col_int", pa.int64()), ("col_float", pa.float64())],
[("col_int", pa.int32()), ("col_float", pa.float64())],
[("col_int", pa.int64()), ("col_float", pa.float32())],
],
)
def test_torch_formatter_sets_default_dtypes(cast_schema, arrow_table):
import torch
from datasets.formatting import TorchFormatter
if cast_schema:
arrow_table = arrow_table.cast(pa.schema(cast_schema))
arrow_table_dict = arrow_table.to_pydict()
list_int = arrow_table_dict["col_int"]
list_float = arrow_table_dict["col_float"]
formatter = TorchFormatter()
row = formatter.format_row(arrow_table)
torch.testing.assert_close(row["col_int"], torch.tensor(list_int, dtype=torch.int64)[0])
torch.testing.assert_close(row["col_float"], torch.tensor(list_float, dtype=torch.float32)[0])
col = formatter.format_column(arrow_table)
torch.testing.assert_close(col, torch.tensor(list_int, dtype=torch.int64))
batch = formatter.format_batch(arrow_table)
torch.testing.assert_close(batch["col_int"], torch.tensor(list_int, dtype=torch.int64))
torch.testing.assert_close(batch["col_float"], torch.tensor(list_float, dtype=torch.float32))
def test_iterable_dataset_of_arrays_format_to_arrow(any_arrays_dataset: IterableDataset):
formatted = any_arrays_dataset.with_format("arrow")
assert all(isinstance(example, pa.Table) for example in formatted)
def test_iterable_dataset_of_arrays_format_to_numpy(any_arrays_dataset: IterableDataset):
formatted = any_arrays_dataset.with_format("np")
assert all(isinstance(example["array"], np.ndarray) for example in formatted)
@require_torch
def test_iterable_dataset_of_arrays_format_to_torch(any_arrays_dataset: IterableDataset):
import torch
formatted = any_arrays_dataset.with_format("torch")
assert all(isinstance(example["array"], torch.Tensor) for example in formatted)
@require_tf
def test_iterable_dataset_of_arrays_format_to_tf(any_arrays_dataset: IterableDataset):
import tensorflow as tf
formatted = any_arrays_dataset.with_format("tf")
assert all(isinstance(example["array"], tf.Tensor) for example in formatted)
@require_jax
def test_iterable_dataset_of_arrays_format_to_jax(any_arrays_dataset: IterableDataset):
import jax.numpy as jnp
formatted = any_arrays_dataset.with_format("jax")
assert all(isinstance(example["array"], jnp.ndarray) for example in formatted)
| 0
|
hf_public_repos/datasets
|
hf_public_repos/datasets/tests/test_hub.py
|
from urllib.parse import quote
import pytest
from datasets.utils.hub import hf_hub_url
@pytest.mark.parametrize("repo_id", ["canonical_dataset_name", "org-name/dataset-name"])
@pytest.mark.parametrize("path", ["filename.csv", "filename with blanks.csv"])
@pytest.mark.parametrize("revision", [None, "v2"])
def test_hf_hub_url(repo_id, path, revision):
url = hf_hub_url(repo_id=repo_id, path=path, revision=revision)
assert url == f"https://huggingface.co/datasets/{repo_id}/resolve/{revision or 'main'}/{quote(path)}"
| 0
|
hf_public_repos/datasets
|
hf_public_repos/datasets/tests/test_parallel.py
|
import pytest
from datasets.parallel import ParallelBackendConfig, parallel_backend
from datasets.utils.py_utils import map_nested
from .utils import require_dill_gt_0_3_2, require_joblibspark, require_not_windows
def add_one(i): # picklable for multiprocessing
return i + 1
@require_dill_gt_0_3_2
@require_joblibspark
@require_not_windows
def test_parallel_backend_input():
with parallel_backend("spark"):
assert ParallelBackendConfig.backend_name == "spark"
lst = [1, 2, 3]
with pytest.raises(ValueError):
with parallel_backend("unsupported backend"):
map_nested(add_one, lst, num_proc=2)
with pytest.raises(ValueError):
with parallel_backend("unsupported backend"):
map_nested(add_one, lst, num_proc=-1)
@require_dill_gt_0_3_2
@require_joblibspark
@require_not_windows
@pytest.mark.parametrize("num_proc", [2, -1])
def test_parallel_backend_map_nested(num_proc):
s1 = [1, 2]
s2 = {"a": 1, "b": 2}
s3 = {"a": [1, 2], "b": [3, 4]}
s4 = {"a": {"1": 1}, "b": 2}
s5 = {"a": 1, "b": 2, "c": 3, "d": 4}
expected_map_nested_s1 = [2, 3]
expected_map_nested_s2 = {"a": 2, "b": 3}
expected_map_nested_s3 = {"a": [2, 3], "b": [4, 5]}
expected_map_nested_s4 = {"a": {"1": 2}, "b": 3}
expected_map_nested_s5 = {"a": 2, "b": 3, "c": 4, "d": 5}
with parallel_backend("spark"):
assert map_nested(add_one, s1, num_proc=num_proc) == expected_map_nested_s1
assert map_nested(add_one, s2, num_proc=num_proc) == expected_map_nested_s2
assert map_nested(add_one, s3, num_proc=num_proc) == expected_map_nested_s3
assert map_nested(add_one, s4, num_proc=num_proc) == expected_map_nested_s4
assert map_nested(add_one, s5, num_proc=num_proc) == expected_map_nested_s5
| 0
|
hf_public_repos/datasets
|
hf_public_repos/datasets/tests/test_metric_common.py
|
# Copyright 2020 HuggingFace Inc.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
import doctest
import glob
import importlib
import inspect
import os
import re
from contextlib import contextmanager
from functools import wraps
from unittest.mock import patch
import numpy as np
import pytest
from absl.testing import parameterized
import datasets
from datasets import load_metric
from .utils import for_all_test_methods, local, slow
# mark all tests as integration
pytestmark = pytest.mark.integration
REQUIRE_FAIRSEQ = {"comet"}
_has_fairseq = importlib.util.find_spec("fairseq") is not None
UNSUPPORTED_ON_WINDOWS = {"code_eval"}
_on_windows = os.name == "nt"
REQUIRE_TRANSFORMERS = {"bertscore", "frugalscore", "perplexity"}
_has_transformers = importlib.util.find_spec("transformers") is not None
def skip_if_metric_requires_fairseq(test_case):
@wraps(test_case)
def wrapper(self, metric_name):
if not _has_fairseq and metric_name in REQUIRE_FAIRSEQ:
self.skipTest('"test requires Fairseq"')
else:
test_case(self, metric_name)
return wrapper
def skip_if_metric_requires_transformers(test_case):
@wraps(test_case)
def wrapper(self, metric_name):
if not _has_transformers and metric_name in REQUIRE_TRANSFORMERS:
self.skipTest('"test requires transformers"')
else:
test_case(self, metric_name)
return wrapper
def skip_on_windows_if_not_windows_compatible(test_case):
@wraps(test_case)
def wrapper(self, metric_name):
if _on_windows and metric_name in UNSUPPORTED_ON_WINDOWS:
self.skipTest('"test not supported on Windows"')
else:
test_case(self, metric_name)
return wrapper
def get_local_metric_names():
metrics = [metric_dir.split(os.sep)[-2] for metric_dir in glob.glob("./metrics/*/")]
return [{"testcase_name": x, "metric_name": x} for x in metrics if x != "gleu"] # gleu is unfinished
@parameterized.named_parameters(get_local_metric_names())
@for_all_test_methods(
skip_if_metric_requires_fairseq, skip_if_metric_requires_transformers, skip_on_windows_if_not_windows_compatible
)
@local
class LocalMetricTest(parameterized.TestCase):
INTENSIVE_CALLS_PATCHER = {}
metric_name = None
@pytest.mark.filterwarnings("ignore:metric_module_factory is deprecated:FutureWarning")
@pytest.mark.filterwarnings("ignore:load_metric is deprecated:FutureWarning")
def test_load_metric(self, metric_name):
doctest.ELLIPSIS_MARKER = "[...]"
metric_module = importlib.import_module(
datasets.load.metric_module_factory(os.path.join("metrics", metric_name)).module_path
)
metric = datasets.load.import_main_class(metric_module.__name__, dataset=False)
# check parameters
parameters = inspect.signature(metric._compute).parameters
self.assertTrue(all(p.kind != p.VAR_KEYWORD for p in parameters.values())) # no **kwargs
# run doctest
with self.patch_intensive_calls(metric_name, metric_module.__name__):
with self.use_local_metrics():
try:
results = doctest.testmod(metric_module, verbose=True, raise_on_error=True)
except doctest.UnexpectedException as e:
raise e.exc_info[1] # raise the exception that doctest caught
self.assertEqual(results.failed, 0)
self.assertGreater(results.attempted, 1)
@slow
def test_load_real_metric(self, metric_name):
doctest.ELLIPSIS_MARKER = "[...]"
metric_module = importlib.import_module(
datasets.load.metric_module_factory(os.path.join("metrics", metric_name)).module_path
)
# run doctest
with self.use_local_metrics():
results = doctest.testmod(metric_module, verbose=True, raise_on_error=True)
self.assertEqual(results.failed, 0)
self.assertGreater(results.attempted, 1)
@contextmanager
def patch_intensive_calls(self, metric_name, module_name):
if metric_name in self.INTENSIVE_CALLS_PATCHER:
with self.INTENSIVE_CALLS_PATCHER[metric_name](module_name):
yield
else:
yield
@contextmanager
def use_local_metrics(self):
def load_local_metric(metric_name, *args, **kwargs):
return load_metric(os.path.join("metrics", metric_name), *args, **kwargs)
with patch("datasets.load_metric") as mock_load_metric:
mock_load_metric.side_effect = load_local_metric
yield
@classmethod
def register_intensive_calls_patcher(cls, metric_name):
def wrapper(patcher):
patcher = contextmanager(patcher)
cls.INTENSIVE_CALLS_PATCHER[metric_name] = patcher
return patcher
return wrapper
# Metrics intensive calls patchers
# --------------------------------
@LocalMetricTest.register_intensive_calls_patcher("bleurt")
def patch_bleurt(module_name):
import tensorflow.compat.v1 as tf
from bleurt.score import Predictor
tf.flags.DEFINE_string("sv", "", "") # handle pytest cli flags
class MockedPredictor(Predictor):
def predict(self, input_dict):
assert len(input_dict["input_ids"]) == 2
return np.array([1.03, 1.04])
# mock predict_fn which is supposed to do a forward pass with a bleurt model
with patch("bleurt.score._create_predictor") as mock_create_predictor:
mock_create_predictor.return_value = MockedPredictor()
yield
@LocalMetricTest.register_intensive_calls_patcher("bertscore")
def patch_bertscore(module_name):
import torch
def bert_cos_score_idf(model, refs, *args, **kwargs):
return torch.tensor([[1.0, 1.0, 1.0]] * len(refs))
# mock get_model which is supposed to do download a bert model
# mock bert_cos_score_idf which is supposed to do a forward pass with a bert model
with patch("bert_score.scorer.get_model"), patch(
"bert_score.scorer.bert_cos_score_idf"
) as mock_bert_cos_score_idf:
mock_bert_cos_score_idf.side_effect = bert_cos_score_idf
yield
@LocalMetricTest.register_intensive_calls_patcher("comet")
def patch_comet(module_name):
def load_from_checkpoint(model_path):
class Model:
def predict(self, data, *args, **kwargs):
assert len(data) == 2
scores = [0.19, 0.92]
return scores, sum(scores) / len(scores)
return Model()
# mock load_from_checkpoint which is supposed to do download a bert model
# mock load_from_checkpoint which is supposed to do download a bert model
with patch("comet.download_model") as mock_download_model:
mock_download_model.return_value = None
with patch("comet.load_from_checkpoint") as mock_load_from_checkpoint:
mock_load_from_checkpoint.side_effect = load_from_checkpoint
yield
def test_seqeval_raises_when_incorrect_scheme():
metric = load_metric(os.path.join("metrics", "seqeval"))
wrong_scheme = "ERROR"
error_message = f"Scheme should be one of [IOB1, IOB2, IOE1, IOE2, IOBES, BILOU], got {wrong_scheme}"
with pytest.raises(ValueError, match=re.escape(error_message)):
metric.compute(predictions=[], references=[], scheme=wrong_scheme)
| 0
|
hf_public_repos/datasets
|
hf_public_repos/datasets/tests/test_offline_util.py
|
import pytest
import requests
from datasets.utils.file_utils import http_head
from .utils import OfflineSimulationMode, RequestWouldHangIndefinitelyError, offline
@pytest.mark.integration
def test_offline_with_timeout():
with offline(OfflineSimulationMode.CONNECTION_TIMES_OUT):
with pytest.raises(RequestWouldHangIndefinitelyError):
requests.request("GET", "https://huggingface.co")
with pytest.raises(requests.exceptions.ConnectTimeout):
requests.request("GET", "https://huggingface.co", timeout=1.0)
@pytest.mark.integration
def test_offline_with_connection_error():
with offline(OfflineSimulationMode.CONNECTION_FAILS):
with pytest.raises(requests.exceptions.ConnectionError):
requests.request("GET", "https://huggingface.co")
def test_offline_with_datasets_offline_mode_enabled():
with offline(OfflineSimulationMode.HF_DATASETS_OFFLINE_SET_TO_1):
with pytest.raises(ConnectionError):
http_head("https://huggingface.co")
| 0
|
hf_public_repos/datasets
|
hf_public_repos/datasets/tests/_test_patching.py
|
# isort: skip_file
# This is the module that test_patching.py uses to test patch_submodule()
import os # noqa: F401 - this is just for tests
import os as renamed_os # noqa: F401 - this is just for tests
from os import path # noqa: F401 - this is just for tests
from os import path as renamed_path # noqa: F401 - this is just for tests
from os.path import join # noqa: F401 - this is just for tests
from os.path import join as renamed_join # noqa: F401 - this is just for tests
open = open # noqa we just need to have a builtin inside this module to test it properly
| 0
|
hf_public_repos/datasets
|
hf_public_repos/datasets/tests/test_arrow_reader.py
|
import os
import tempfile
from pathlib import Path
from unittest import TestCase
import pyarrow as pa
import pytest
from datasets.arrow_dataset import Dataset
from datasets.arrow_reader import ArrowReader, BaseReader, FileInstructions, ReadInstruction, make_file_instructions
from datasets.info import DatasetInfo
from datasets.splits import NamedSplit, Split, SplitDict, SplitInfo
from .utils import assert_arrow_memory_doesnt_increase, assert_arrow_memory_increases
class ReaderTest(BaseReader):
"""
Build a Dataset object out of Instruction instance(s).
This reader is made for testing. It mocks file reads.
"""
def _get_table_from_filename(self, filename_skip_take, in_memory=False):
"""Returns a Dataset instance from given (filename, skip, take)."""
filename, skip, take = (
filename_skip_take["filename"],
filename_skip_take["skip"] if "skip" in filename_skip_take else None,
filename_skip_take["take"] if "take" in filename_skip_take else None,
)
open(os.path.join(filename), "wb").close()
pa_table = pa.Table.from_pydict({"filename": [Path(filename).name] * 100})
if take == -1:
take = len(pa_table) - skip
if skip is not None and take is not None:
pa_table = pa_table.slice(skip, take)
return pa_table
class BaseReaderTest(TestCase):
def test_read(self):
name = "my_name"
train_info = SplitInfo(name="train", num_examples=100)
test_info = SplitInfo(name="test", num_examples=100)
split_infos = [train_info, test_info]
split_dict = SplitDict()
split_dict.add(train_info)
split_dict.add(test_info)
info = DatasetInfo(splits=split_dict)
with tempfile.TemporaryDirectory() as tmp_dir:
reader = ReaderTest(tmp_dir, info)
instructions = "test[:33%]"
dset = Dataset(**reader.read(name, instructions, split_infos))
self.assertEqual(dset["filename"][0], f"{name}-test")
self.assertEqual(dset.num_rows, 33)
self.assertEqual(dset.num_columns, 1)
instructions1 = ["train", "test[:33%]"]
instructions2 = [Split.TRAIN, ReadInstruction.from_spec("test[:33%]")]
for instructions in [instructions1, instructions2]:
datasets_kwargs = [reader.read(name, instr, split_infos) for instr in instructions]
train_dset, test_dset = (Dataset(**dataset_kwargs) for dataset_kwargs in datasets_kwargs)
self.assertEqual(train_dset["filename"][0], f"{name}-train")
self.assertEqual(train_dset.num_rows, 100)
self.assertEqual(train_dset.num_columns, 1)
self.assertIsInstance(train_dset.split, NamedSplit)
self.assertEqual(str(train_dset.split), "train")
self.assertEqual(test_dset["filename"][0], f"{name}-test")
self.assertEqual(test_dset.num_rows, 33)
self.assertEqual(test_dset.num_columns, 1)
self.assertIsInstance(test_dset.split, NamedSplit)
self.assertEqual(str(test_dset.split), "test[:33%]")
del train_dset, test_dset
def test_read_sharded(self):
name = "my_name"
train_info = SplitInfo(name="train", num_examples=1000, shard_lengths=[100] * 10)
split_infos = [train_info]
split_dict = SplitDict()
split_dict.add(train_info)
info = DatasetInfo(splits=split_dict)
with tempfile.TemporaryDirectory() as tmp_dir:
reader = ReaderTest(tmp_dir, info)
instructions = "train[:33%]"
dset = Dataset(**reader.read(name, instructions, split_infos))
self.assertEqual(dset["filename"][0], f"{name}-train-00000-of-00010")
self.assertEqual(dset["filename"][-1], f"{name}-train-00003-of-00010")
self.assertEqual(dset.num_rows, 330)
self.assertEqual(dset.num_columns, 1)
def test_read_files(self):
train_info = SplitInfo(name="train", num_examples=100)
test_info = SplitInfo(name="test", num_examples=100)
split_dict = SplitDict()
split_dict.add(train_info)
split_dict.add(test_info)
info = DatasetInfo(splits=split_dict)
with tempfile.TemporaryDirectory() as tmp_dir:
reader = ReaderTest(tmp_dir, info)
files = [
{"filename": os.path.join(tmp_dir, "train")},
{"filename": os.path.join(tmp_dir, "test"), "skip": 10, "take": 10},
]
dset = Dataset(**reader.read_files(files, original_instructions="train+test[10:20]"))
self.assertEqual(dset.num_rows, 110)
self.assertEqual(dset.num_columns, 1)
del dset
@pytest.mark.parametrize("in_memory", [False, True])
def test_read_table(in_memory, dataset, arrow_file):
filename = arrow_file
with assert_arrow_memory_increases() if in_memory else assert_arrow_memory_doesnt_increase():
table = ArrowReader.read_table(filename, in_memory=in_memory)
assert table.shape == dataset.data.shape
assert set(table.column_names) == set(dataset.data.column_names)
assert dict(table.to_pydict()) == dict(dataset.data.to_pydict()) # to_pydict returns OrderedDict
@pytest.mark.parametrize("in_memory", [False, True])
def test_read_files(in_memory, dataset, arrow_file):
filename = arrow_file
reader = ArrowReader("", None)
with assert_arrow_memory_increases() if in_memory else assert_arrow_memory_doesnt_increase():
dataset_kwargs = reader.read_files([{"filename": filename}], in_memory=in_memory)
assert dataset_kwargs.keys() == {"arrow_table", "info", "split"}
table = dataset_kwargs["arrow_table"]
assert table.shape == dataset.data.shape
assert set(table.column_names) == set(dataset.data.column_names)
assert dict(table.to_pydict()) == dict(dataset.data.to_pydict()) # to_pydict returns OrderedDict
def test_read_instruction_spec():
assert ReadInstruction("train", to=10, unit="abs").to_spec() == "train[:10]"
assert ReadInstruction("train", from_=-80, to=10, unit="%").to_spec() == "train[-80%:10%]"
spec_train_test = "train+test"
assert ReadInstruction.from_spec(spec_train_test).to_spec() == spec_train_test
spec_train_abs = "train[2:10]"
assert ReadInstruction.from_spec(spec_train_abs).to_spec() == spec_train_abs
spec_train_pct = "train[15%:-20%]"
assert ReadInstruction.from_spec(spec_train_pct).to_spec() == spec_train_pct
spec_train_pct_rounding = "train[:10%](closest)"
assert ReadInstruction.from_spec(spec_train_pct_rounding).to_spec() == "train[:10%]"
spec_train_pct_rounding = "train[:10%](pct1_dropremainder)"
assert ReadInstruction.from_spec(spec_train_pct_rounding).to_spec() == spec_train_pct_rounding
spec_train_test_pct_rounding = "train[:10%](pct1_dropremainder)+test[-10%:](pct1_dropremainder)"
assert ReadInstruction.from_spec(spec_train_test_pct_rounding).to_spec() == spec_train_test_pct_rounding
def test_make_file_instructions():
name = "dummy"
split_infos = [SplitInfo(name="train", num_examples=100)]
instruction = "train[:33%]"
filetype_suffix = "arrow"
prefix_path = "prefix"
file_instructions = make_file_instructions(name, split_infos, instruction, filetype_suffix, prefix_path)
assert isinstance(file_instructions, FileInstructions)
assert file_instructions.num_examples == 33
assert file_instructions.file_instructions == [
{"filename": os.path.join(prefix_path, f"{name}-train.arrow"), "skip": 0, "take": 33}
]
split_infos = [SplitInfo(name="train", num_examples=100, shard_lengths=[10] * 10)]
file_instructions = make_file_instructions(name, split_infos, instruction, filetype_suffix, prefix_path)
assert isinstance(file_instructions, FileInstructions)
assert file_instructions.num_examples == 33
assert file_instructions.file_instructions == [
{"filename": os.path.join(prefix_path, f"{name}-train-00000-of-00010.arrow"), "skip": 0, "take": -1},
{"filename": os.path.join(prefix_path, f"{name}-train-00001-of-00010.arrow"), "skip": 0, "take": -1},
{"filename": os.path.join(prefix_path, f"{name}-train-00002-of-00010.arrow"), "skip": 0, "take": -1},
{"filename": os.path.join(prefix_path, f"{name}-train-00003-of-00010.arrow"), "skip": 0, "take": 3},
]
@pytest.mark.parametrize("name, expected_exception", [(None, TypeError), ("", ValueError)])
def test_make_file_instructions_raises(name, expected_exception):
split_infos = [SplitInfo(name="train", num_examples=100)]
instruction = "train"
filetype_suffix = "arrow"
prefix_path = "prefix_path"
with pytest.raises(expected_exception):
_ = make_file_instructions(name, split_infos, instruction, filetype_suffix, prefix_path)
| 0
|
hf_public_repos/datasets
|
hf_public_repos/datasets/tests/test_tasks.py
|
from copy import deepcopy
from unittest.case import TestCase
import pytest
from datasets.arrow_dataset import Dataset
from datasets.features import Audio, ClassLabel, Features, Image, Sequence, Value
from datasets.info import DatasetInfo
from datasets.tasks import (
AudioClassification,
AutomaticSpeechRecognition,
ImageClassification,
LanguageModeling,
QuestionAnsweringExtractive,
Summarization,
TextClassification,
task_template_from_dict,
)
from datasets.utils.py_utils import asdict
SAMPLE_QUESTION_ANSWERING_EXTRACTIVE = {
"id": "5733be284776f41900661182",
"title": "University_of_Notre_Dame",
"context": 'Architecturally, the school has a Catholic character. Atop the Main Building\'s gold dome is a golden statue of the Virgin Mary. Immediately in front of the Main Building and facing it, is a copper statue of Christ with arms upraised with the legend "Venite Ad Me Omnes". Next to the Main Building is the Basilica of the Sacred Heart. Immediately behind the basilica is the Grotto, a Marian place of prayer and reflection. It is a replica of the grotto at Lourdes, France where the Virgin Mary reputedly appeared to Saint Bernadette Soubirous in 1858. At the end of the main drive (and in a direct line that connects through 3 statues and the Gold Dome), is a simple, modern stone statue of Mary.',
"question": "To whom did the Virgin Mary allegedly appear in 1858 in Lourdes France?",
"answers": {"text": ["Saint Bernadette Soubirous"], "answer_start": [515]},
}
@pytest.mark.parametrize(
"task_cls",
[
AudioClassification,
AutomaticSpeechRecognition,
ImageClassification,
LanguageModeling,
QuestionAnsweringExtractive,
Summarization,
TextClassification,
],
)
def test_reload_task_from_dict(task_cls):
task = task_cls()
task_dict = asdict(task)
reloaded = task_template_from_dict(task_dict)
assert task == reloaded
class TestLanguageModeling:
def test_column_mapping(self):
task = LanguageModeling(text_column="input_text")
assert {"input_text": "text"} == task.column_mapping
def test_from_dict(self):
input_schema = Features({"text": Value("string")})
template_dict = {"text_column": "input_text"}
task = LanguageModeling.from_dict(template_dict)
assert "language-modeling" == task.task
assert input_schema == task.input_schema
class TextClassificationTest(TestCase):
def setUp(self):
self.labels = sorted(["pos", "neg"])
def test_column_mapping(self):
task = TextClassification(text_column="input_text", label_column="input_label")
self.assertDictEqual({"input_text": "text", "input_label": "labels"}, task.column_mapping)
def test_from_dict(self):
input_schema = Features({"text": Value("string")})
# Labels are cast to tuple during `TextClassification.__post_init__`, so we do the same here
label_schema = Features({"labels": ClassLabel})
template_dict = {"text_column": "input_text", "label_column": "input_labels"}
task = TextClassification.from_dict(template_dict)
self.assertEqual("text-classification", task.task)
self.assertEqual(input_schema, task.input_schema)
self.assertEqual(label_schema, task.label_schema)
def test_align_with_features(self):
task = TextClassification(text_column="input_text", label_column="input_label")
self.assertEqual(task.label_schema["labels"], ClassLabel)
task = task.align_with_features(Features({"input_label": ClassLabel(names=self.labels)}))
self.assertEqual(task.label_schema["labels"], ClassLabel(names=self.labels))
class QuestionAnsweringTest(TestCase):
def test_column_mapping(self):
task = QuestionAnsweringExtractive(
context_column="input_context", question_column="input_question", answers_column="input_answers"
)
self.assertDictEqual(
{"input_context": "context", "input_question": "question", "input_answers": "answers"}, task.column_mapping
)
def test_from_dict(self):
input_schema = Features({"question": Value("string"), "context": Value("string")})
label_schema = Features(
{
"answers": Sequence(
{
"text": Value("string"),
"answer_start": Value("int32"),
}
)
}
)
template_dict = {
"context_column": "input_input_context",
"question_column": "input_question",
"answers_column": "input_answers",
}
task = QuestionAnsweringExtractive.from_dict(template_dict)
self.assertEqual("question-answering-extractive", task.task)
self.assertEqual(input_schema, task.input_schema)
self.assertEqual(label_schema, task.label_schema)
class SummarizationTest(TestCase):
def test_column_mapping(self):
task = Summarization(text_column="input_text", summary_column="input_summary")
self.assertDictEqual({"input_text": "text", "input_summary": "summary"}, task.column_mapping)
def test_from_dict(self):
input_schema = Features({"text": Value("string")})
label_schema = Features({"summary": Value("string")})
template_dict = {"text_column": "input_text", "summary_column": "input_summary"}
task = Summarization.from_dict(template_dict)
self.assertEqual("summarization", task.task)
self.assertEqual(input_schema, task.input_schema)
self.assertEqual(label_schema, task.label_schema)
class AutomaticSpeechRecognitionTest(TestCase):
def test_column_mapping(self):
task = AutomaticSpeechRecognition(audio_column="input_audio", transcription_column="input_transcription")
self.assertDictEqual({"input_audio": "audio", "input_transcription": "transcription"}, task.column_mapping)
def test_from_dict(self):
input_schema = Features({"audio": Audio()})
label_schema = Features({"transcription": Value("string")})
template_dict = {
"audio_column": "input_audio",
"transcription_column": "input_transcription",
}
task = AutomaticSpeechRecognition.from_dict(template_dict)
self.assertEqual("automatic-speech-recognition", task.task)
self.assertEqual(input_schema, task.input_schema)
self.assertEqual(label_schema, task.label_schema)
class AudioClassificationTest(TestCase):
def setUp(self):
self.labels = sorted(["pos", "neg"])
def test_column_mapping(self):
task = AudioClassification(audio_column="input_audio", label_column="input_label")
self.assertDictEqual({"input_audio": "audio", "input_label": "labels"}, task.column_mapping)
def test_from_dict(self):
input_schema = Features({"audio": Audio()})
label_schema = Features({"labels": ClassLabel})
template_dict = {
"audio_column": "input_image",
"label_column": "input_label",
}
task = AudioClassification.from_dict(template_dict)
self.assertEqual("audio-classification", task.task)
self.assertEqual(input_schema, task.input_schema)
self.assertEqual(label_schema, task.label_schema)
def test_align_with_features(self):
task = AudioClassification(audio_column="input_audio", label_column="input_label")
self.assertEqual(task.label_schema["labels"], ClassLabel)
task = task.align_with_features(Features({"input_label": ClassLabel(names=self.labels)}))
self.assertEqual(task.label_schema["labels"], ClassLabel(names=self.labels))
class ImageClassificationTest(TestCase):
def setUp(self):
self.labels = sorted(["pos", "neg"])
def test_column_mapping(self):
task = ImageClassification(image_column="input_image", label_column="input_label")
self.assertDictEqual({"input_image": "image", "input_label": "labels"}, task.column_mapping)
def test_from_dict(self):
input_schema = Features({"image": Image()})
label_schema = Features({"labels": ClassLabel})
template_dict = {
"image_column": "input_image",
"label_column": "input_label",
}
task = ImageClassification.from_dict(template_dict)
self.assertEqual("image-classification", task.task)
self.assertEqual(input_schema, task.input_schema)
self.assertEqual(label_schema, task.label_schema)
def test_align_with_features(self):
task = ImageClassification(image_column="input_image", label_column="input_label")
self.assertEqual(task.label_schema["labels"], ClassLabel)
task = task.align_with_features(Features({"input_label": ClassLabel(names=self.labels)}))
self.assertEqual(task.label_schema["labels"], ClassLabel(names=self.labels))
class DatasetWithTaskProcessingTest(TestCase):
def test_map_on_task_template(self):
info = DatasetInfo(task_templates=QuestionAnsweringExtractive())
dataset = Dataset.from_dict({k: [v] for k, v in SAMPLE_QUESTION_ANSWERING_EXTRACTIVE.items()}, info=info)
assert isinstance(dataset.info.task_templates, list)
assert len(dataset.info.task_templates) == 1
def keep_task(x):
return x
def dont_keep_task(x):
out = deepcopy(SAMPLE_QUESTION_ANSWERING_EXTRACTIVE)
out["answers"]["foobar"] = 0
return out
mapped_dataset = dataset.map(keep_task)
assert mapped_dataset.info.task_templates == dataset.info.task_templates
# reload from cache
mapped_dataset = dataset.map(keep_task)
assert mapped_dataset.info.task_templates == dataset.info.task_templates
mapped_dataset = dataset.map(dont_keep_task)
assert mapped_dataset.info.task_templates == []
# reload from cache
mapped_dataset = dataset.map(dont_keep_task)
assert mapped_dataset.info.task_templates == []
def test_remove_and_map_on_task_template(self):
features = Features({"text": Value("string"), "label": ClassLabel(names=("pos", "neg"))})
task_templates = TextClassification(text_column="text", label_column="label")
info = DatasetInfo(features=features, task_templates=task_templates)
dataset = Dataset.from_dict({"text": ["A sentence."], "label": ["pos"]}, info=info)
def process(example):
return example
modified_dataset = dataset.remove_columns("label")
mapped_dataset = modified_dataset.map(process)
assert mapped_dataset.info.task_templates == []
| 0
|
hf_public_repos/datasets
|
hf_public_repos/datasets/tests/test_patching.py
|
from datasets.utils.patching import _PatchedModuleObj, patch_submodule
from . import _test_patching
def test_patch_submodule():
import os as original_os
from os import path as original_path
from os import rename as original_rename
from os.path import dirname as original_dirname
from os.path import join as original_join
assert _test_patching.os is original_os
assert _test_patching.path is original_path
assert _test_patching.join is original_join
assert _test_patching.renamed_os is original_os
assert _test_patching.renamed_path is original_path
assert _test_patching.renamed_join is original_join
mock = "__test_patch_submodule_mock__"
with patch_submodule(_test_patching, "os.path.join", mock):
# Every way to access os.path.join must be patched, and the rest must stay untouched
# check os.path.join
assert isinstance(_test_patching.os, _PatchedModuleObj)
assert isinstance(_test_patching.os.path, _PatchedModuleObj)
assert _test_patching.os.path.join is mock
# check path.join
assert isinstance(_test_patching.path, _PatchedModuleObj)
assert _test_patching.path.join is mock
# check join
assert _test_patching.join is mock
# check that the other attributes are untouched
assert _test_patching.os.rename is original_rename
assert _test_patching.path.dirname is original_dirname
assert _test_patching.os.path.dirname is original_dirname
# Even renamed modules or objects must be patched
# check renamed_os.path.join
assert isinstance(_test_patching.renamed_os, _PatchedModuleObj)
assert isinstance(_test_patching.renamed_os.path, _PatchedModuleObj)
assert _test_patching.renamed_os.path.join is mock
# check renamed_path.join
assert isinstance(_test_patching.renamed_path, _PatchedModuleObj)
assert _test_patching.renamed_path.join is mock
# check renamed_join
assert _test_patching.renamed_join is mock
# check that the other attributes are untouched
assert _test_patching.renamed_os.rename is original_rename
assert _test_patching.renamed_path.dirname is original_dirname
assert _test_patching.renamed_os.path.dirname is original_dirname
# check that everthing is back to normal when the patch is over
assert _test_patching.os is original_os
assert _test_patching.path is original_path
assert _test_patching.join is original_join
assert _test_patching.renamed_os is original_os
assert _test_patching.renamed_path is original_path
assert _test_patching.renamed_join is original_join
def test_patch_submodule_builtin():
assert _test_patching.open is open
mock = "__test_patch_submodule_builtin_mock__"
# _test_patching has "open" in its globals
assert _test_patching.open is open
with patch_submodule(_test_patching, "open", mock):
assert _test_patching.open is mock
# check that everthing is back to normal when the patch is over
assert _test_patching.open is open
def test_patch_submodule_missing():
# pandas.read_csv is not present in _test_patching
mock = "__test_patch_submodule_missing_mock__"
with patch_submodule(_test_patching, "pandas.read_csv", mock):
pass
def test_patch_submodule_missing_builtin():
# builtin should always be mocked even if they're not in the globals
# in case they're loaded at one point
mock = "__test_patch_submodule_missing_builtin_mock__"
# _test_patching doesn't have "len" in its globals
assert getattr(_test_patching, "len", None) is None
with patch_submodule(_test_patching, "len", mock):
assert _test_patching.len is mock
assert _test_patching.len is len
def test_patch_submodule_start_and_stop():
mock = "__test_patch_submodule_start_and_stop_mock__"
patch = patch_submodule(_test_patching, "open", mock)
assert _test_patching.open is open
patch.start()
assert _test_patching.open is mock
patch.stop()
assert _test_patching.open is open
def test_patch_submodule_successive():
from os import rename as original_rename
from os.path import dirname as original_dirname
from os.path import join as original_join
mock_join = "__test_patch_submodule_successive_join__"
mock_dirname = "__test_patch_submodule_successive_dirname__"
mock_rename = "__test_patch_submodule_successive_rename__"
assert _test_patching.os.path.join is original_join
assert _test_patching.os.path.dirname is original_dirname
assert _test_patching.os.rename is original_rename
with patch_submodule(_test_patching, "os.path.join", mock_join):
with patch_submodule(_test_patching, "os.rename", mock_rename):
with patch_submodule(_test_patching, "os.path.dirname", mock_dirname):
assert _test_patching.os.path.join is mock_join
assert _test_patching.os.path.dirname is mock_dirname
assert _test_patching.os.rename is mock_rename
# try another order
with patch_submodule(_test_patching, "os.rename", mock_rename):
with patch_submodule(_test_patching, "os.path.join", mock_join):
with patch_submodule(_test_patching, "os.path.dirname", mock_dirname):
assert _test_patching.os.path.join is mock_join
assert _test_patching.os.path.dirname is mock_dirname
assert _test_patching.os.rename is mock_rename
assert _test_patching.os.path.join is original_join
assert _test_patching.os.path.dirname is original_dirname
assert _test_patching.os.rename is original_rename
def test_patch_submodule_doesnt_exist():
mock = "__test_patch_submodule_doesnt_exist_mock__"
with patch_submodule(_test_patching, "__module_that_doesn_exist__.__attribute_that_doesn_exist__", mock):
pass
with patch_submodule(_test_patching, "os.__attribute_that_doesn_exist__", mock):
pass
| 0
|
hf_public_repos/datasets
|
hf_public_repos/datasets/tests/test_readme_util.py
|
import re
import tempfile
from pathlib import Path
import pytest
import yaml
from datasets.utils.readme import ReadMe
# @pytest.fixture
# def example_yaml_structure():
example_yaml_structure = yaml.safe_load(
"""\
name: ""
allow_empty: false
allow_empty_text: true
subsections:
- name: "Dataset Card for X" # First-level markdown heading
allow_empty: false
allow_empty_text: true
subsections:
- name: "Table of Contents"
allow_empty: false
allow_empty_text: false
subsections: null
- name: "Dataset Description"
allow_empty: false
allow_empty_text: false
subsections:
- name: "Dataset Summary"
allow_empty: false
allow_empty_text: false
subsections: null
- name: "Supported Tasks and Leaderboards"
allow_empty: true
allow_empty_text: true
subsections: null
- name: Languages
allow_empty: false
allow_empty_text: true
subsections: null
"""
)
CORRECT_DICT = {
"name": "root",
"text": "",
"is_empty_text": True,
"subsections": [
{
"name": "Dataset Card for My Dataset",
"text": "",
"is_empty_text": True,
"subsections": [
{"name": "Table of Contents", "text": "Some text here.", "is_empty_text": False, "subsections": []},
{
"name": "Dataset Description",
"text": "Some text here.",
"is_empty_text": False,
"subsections": [
{
"name": "Dataset Summary",
"text": "Some text here.",
"is_empty_text": False,
"subsections": [],
},
{
"name": "Supported Tasks and Leaderboards",
"text": "",
"is_empty_text": True,
"subsections": [],
},
{"name": "Languages", "text": "Language Text", "is_empty_text": False, "subsections": []},
],
},
],
}
],
}
README_CORRECT = """\
---
language:
- zh
- en
---
# Dataset Card for My Dataset
## Table of Contents
Some text here.
## Dataset Description
Some text here.
### Dataset Summary
Some text here.
### Supported Tasks and Leaderboards
### Languages
Language Text
"""
README_CORRECT_FOUR_LEVEL = """\
---
language:
- zh
- en
---
# Dataset Card for My Dataset
## Table of Contents
Some text here.
## Dataset Description
Some text here.
### Dataset Summary
Some text here.
#### Extra Ignored Subsection
### Supported Tasks and Leaderboards
### Languages
Language Text
"""
CORRECT_DICT_FOUR_LEVEL = {
"name": "root",
"text": "",
"is_empty_text": True,
"subsections": [
{
"name": "Dataset Card for My Dataset",
"text": "",
"is_empty_text": True,
"subsections": [
{"name": "Table of Contents", "text": "Some text here.", "is_empty_text": False, "subsections": []},
{
"name": "Dataset Description",
"text": "Some text here.",
"is_empty_text": False,
"subsections": [
{
"name": "Dataset Summary",
"text": "Some text here.",
"is_empty_text": False,
"subsections": [
{
"name": "Extra Ignored Subsection",
"text": "",
"is_empty_text": True,
"subsections": [],
}
],
},
{
"name": "Supported Tasks and Leaderboards",
"text": "",
"is_empty_text": True,
"subsections": [],
},
{"name": "Languages", "text": "Language Text", "is_empty_text": False, "subsections": []},
],
},
],
}
],
}
README_EMPTY_YAML = """\
---
---
# Dataset Card for My Dataset
## Table of Contents
Some text here.
## Dataset Description
Some text here.
### Dataset Summary
Some text here.
### Supported Tasks and Leaderboards
### Languages
Language Text
"""
EXPECTED_ERROR_README_EMPTY_YAML = (
"The following issues were found for the README at `{path}`:\n-\tEmpty YAML markers are present in the README."
)
README_NO_YAML = """\
# Dataset Card for My Dataset
## Table of Contents
Some text here.
## Dataset Description
Some text here.
### Dataset Summary
Some text here.
### Supported Tasks and Leaderboards
### Languages
Language Text
"""
EXPECTED_ERROR_README_NO_YAML = (
"The following issues were found for the README at `{path}`:\n-\tNo YAML markers are present in the README."
)
README_INCORRECT_YAML = """\
---
# Dataset Card for My Dataset
## Table of Contents
Some text here.
## Dataset Description
Some text here.
### Dataset Summary
Some text here.
### Supported Tasks and Leaderboards
### Languages
Language Text
"""
EXPECTED_ERROR_README_INCORRECT_YAML = "The following issues were found for the README at `{path}`:\n-\tOnly the start of YAML tags present in the README."
README_MISSING_TEXT = """\
---
language:
- zh
- en
---
# Dataset Card for My Dataset
## Table of Contents
Some text here.
## Dataset Description
Some text here.
### Dataset Summary
### Supported Tasks and Leaderboards
### Languages
Language Text
"""
EXPECTED_ERROR_README_MISSING_TEXT = "The following issues were found for the README at `{path}`:\n-\tExpected some content in section `Dataset Summary` but it is empty.\n-\tExpected some text in section `Dataset Summary` but it is empty (text in subsections are ignored)."
README_NONE_SUBSECTION = """\
---
language:
- zh
- en
---
# Dataset Card for My Dataset
"""
EXPECTED_ERROR_README_NONE_SUBSECTION = "The following issues were found for the README at `{path}`:\n-\tExpected some content in section `Dataset Card for My Dataset` but it is empty.\n-\tSection `Dataset Card for My Dataset` expected the following subsections: `Table of Contents`, `Dataset Description`. Found 'None'."
README_MISSING_SUBSECTION = """\
---
language:
- zh
- en
---
# Dataset Card for My Dataset
## Table of Contents
Some text here.
## Dataset Description
Some text here.
### Dataset Summary
Some text here.
### Languages
Language Text
"""
EXPECTED_ERROR_README_MISSING_SUBSECTION = "The following issues were found for the README at `{path}`:\n-\tSection `Dataset Description` is missing subsection: `Supported Tasks and Leaderboards`."
README_MISSING_CONTENT = """\
---
language:
- zh
- en
---
# Dataset Card for My Dataset
## Table of Contents
Some text here.
## Dataset Description
Some text here.
### Dataset Summary
Some text here.
### Supported Tasks and Leaderboards
### Languages
"""
EXPECTED_ERROR_README_MISSING_CONTENT = "The following issues were found for the README at `{path}`:\n-\tExpected some content in section `Languages` but it is empty."
README_MISSING_FIRST_LEVEL = """\
---
language:
- zh
- en
---
## Table of Contents
Some text here.
## Dataset Description
Some text here.
### Dataset Summary
Some text here.
### Supported Tasks and Leaderboards
### Languages
Language Text
"""
EXPECTED_ERROR_README_MISSING_FIRST_LEVEL = "The following issues were found for the README at `{path}`:\n-\tThe README has no first-level headings. One heading is expected. Skipping further validation for this README."
README_MULTIPLE_WRONG_FIRST_LEVEL = """\
---
language:
- zh
- en
---
# Dataset Card for My Dataset
## Table of Contents
Some text here.
## Dataset Description
Some text here.
### Dataset Summary
Some text here.
### Supported Tasks and Leaderboards
### Languages
Language Text
# Dataset Card My Dataset
"""
EXPECTED_ERROR_README_MULTIPLE_WRONG_FIRST_LEVEL = "The following issues were found for the README at `{path}`:\n-\tThe README has several first-level headings: `Dataset Card for My Dataset`, `Dataset Card My Dataset`. Only one heading is expected. Skipping further validation for this README."
README_WRONG_FIRST_LEVEL = """\
---
language:
- zh
- en
---
# Dataset Card My Dataset
## Table of Contents
Some text here.
## Dataset Description
Some text here.
### Dataset Summary
Some text here.
### Supported Tasks and Leaderboards
### Languages
Language Text
"""
EXPECTED_ERROR_README_WRONG_FIRST_LEVEL = "The following issues were found for the README at `{path}`:\n-\tNo first-level heading starting with `Dataset Card for` found in README. Skipping further validation for this README."
README_EMPTY = ""
EXPECTED_ERROR_README_EMPTY = "The following issues were found for the README at `{path}`:\n-\tThe README has no first-level headings. One heading is expected. Skipping further validation for this README.\n-\tNo YAML markers are present in the README."
README_MULTIPLE_SAME_HEADING_1 = """\
---
language:
- zh
- en
---
# Dataset Card for My Dataset
# Dataset Card for My Dataset
## Table of Contents
Some text here.
## Dataset Description
Some text here.
### Dataset Summary
Some text here.
### Supported Tasks and Leaderboards
### Languages
Language Text
"""
EXPECTED_ERROR_README_MULTIPLE_SAME_HEADING_1 = "The following issues were found while parsing the README at `{path}`:\n-\tMultiple sections with the same heading `Dataset Card for My Dataset` have been found. Please keep only one of these sections."
@pytest.mark.parametrize(
"readme_md, expected_dict",
[
(README_CORRECT, CORRECT_DICT),
(README_CORRECT_FOUR_LEVEL, CORRECT_DICT_FOUR_LEVEL),
],
)
def test_readme_from_string_correct(readme_md, expected_dict):
assert ReadMe.from_string(readme_md, example_yaml_structure).to_dict() == expected_dict
@pytest.mark.parametrize(
"readme_md, expected_error",
[
(README_NO_YAML, EXPECTED_ERROR_README_NO_YAML),
(README_EMPTY_YAML, EXPECTED_ERROR_README_EMPTY_YAML),
(README_INCORRECT_YAML, EXPECTED_ERROR_README_INCORRECT_YAML),
(README_EMPTY, EXPECTED_ERROR_README_EMPTY),
(README_NONE_SUBSECTION, EXPECTED_ERROR_README_NONE_SUBSECTION),
(README_MISSING_FIRST_LEVEL, EXPECTED_ERROR_README_MISSING_FIRST_LEVEL),
(README_MISSING_SUBSECTION, EXPECTED_ERROR_README_MISSING_SUBSECTION),
(README_MISSING_TEXT, EXPECTED_ERROR_README_MISSING_TEXT),
(README_WRONG_FIRST_LEVEL, EXPECTED_ERROR_README_WRONG_FIRST_LEVEL),
(README_MULTIPLE_WRONG_FIRST_LEVEL, EXPECTED_ERROR_README_MULTIPLE_WRONG_FIRST_LEVEL),
(README_MISSING_CONTENT, EXPECTED_ERROR_README_MISSING_CONTENT),
],
)
def test_readme_from_string_validation_errors(readme_md, expected_error):
with pytest.raises(ValueError, match=re.escape(expected_error.format(path="root"))):
readme = ReadMe.from_string(readme_md, example_yaml_structure)
readme.validate()
@pytest.mark.parametrize(
"readme_md, expected_error",
[
(README_MULTIPLE_SAME_HEADING_1, EXPECTED_ERROR_README_MULTIPLE_SAME_HEADING_1),
],
)
def test_readme_from_string_parsing_errors(readme_md, expected_error):
with pytest.raises(ValueError, match=re.escape(expected_error.format(path="root"))):
ReadMe.from_string(readme_md, example_yaml_structure)
@pytest.mark.parametrize(
"readme_md,",
[
(README_MULTIPLE_SAME_HEADING_1),
],
)
def test_readme_from_string_suppress_parsing_errors(readme_md):
ReadMe.from_string(readme_md, example_yaml_structure, suppress_parsing_errors=True)
@pytest.mark.parametrize(
"readme_md, expected_dict",
[
(README_CORRECT, CORRECT_DICT),
(README_CORRECT_FOUR_LEVEL, CORRECT_DICT_FOUR_LEVEL),
],
)
def test_readme_from_readme_correct(readme_md, expected_dict):
with tempfile.TemporaryDirectory() as tmp_dir:
path = Path(tmp_dir) / "README.md"
with open(path, "w+") as readme_file:
readme_file.write(readme_md)
out = ReadMe.from_readme(path, example_yaml_structure).to_dict()
assert out["name"] == path
assert out["text"] == ""
assert out["is_empty_text"]
assert out["subsections"] == expected_dict["subsections"]
@pytest.mark.parametrize(
"readme_md, expected_error",
[
(README_NO_YAML, EXPECTED_ERROR_README_NO_YAML),
(README_EMPTY_YAML, EXPECTED_ERROR_README_EMPTY_YAML),
(README_INCORRECT_YAML, EXPECTED_ERROR_README_INCORRECT_YAML),
(README_EMPTY, EXPECTED_ERROR_README_EMPTY),
(README_NONE_SUBSECTION, EXPECTED_ERROR_README_NONE_SUBSECTION),
(README_MISSING_FIRST_LEVEL, EXPECTED_ERROR_README_MISSING_FIRST_LEVEL),
(README_MISSING_SUBSECTION, EXPECTED_ERROR_README_MISSING_SUBSECTION),
(README_MISSING_TEXT, EXPECTED_ERROR_README_MISSING_TEXT),
(README_WRONG_FIRST_LEVEL, EXPECTED_ERROR_README_WRONG_FIRST_LEVEL),
(README_MULTIPLE_WRONG_FIRST_LEVEL, EXPECTED_ERROR_README_MULTIPLE_WRONG_FIRST_LEVEL),
(README_MISSING_CONTENT, EXPECTED_ERROR_README_MISSING_CONTENT),
],
)
def test_readme_from_readme_error(readme_md, expected_error):
with tempfile.TemporaryDirectory() as tmp_dir:
path = Path(tmp_dir) / "README.md"
with open(path, "w+") as readme_file:
readme_file.write(readme_md)
expected_error = expected_error.format(path=path)
with pytest.raises(ValueError, match=re.escape(expected_error)):
readme = ReadMe.from_readme(path, example_yaml_structure)
readme.validate()
@pytest.mark.parametrize(
"readme_md, expected_error",
[
(README_MULTIPLE_SAME_HEADING_1, EXPECTED_ERROR_README_MULTIPLE_SAME_HEADING_1),
],
)
def test_readme_from_readme_parsing_errors(readme_md, expected_error):
with tempfile.TemporaryDirectory() as tmp_dir:
path = Path(tmp_dir) / "README.md"
with open(path, "w+") as readme_file:
readme_file.write(readme_md)
expected_error = expected_error.format(path=path)
with pytest.raises(ValueError, match=re.escape(expected_error)):
ReadMe.from_readme(path, example_yaml_structure)
@pytest.mark.parametrize(
"readme_md,",
[
(README_MULTIPLE_SAME_HEADING_1),
],
)
def test_readme_from_readme_suppress_parsing_errors(readme_md):
with tempfile.TemporaryDirectory() as tmp_dir:
path = Path(tmp_dir) / "README.md"
with open(path, "w+") as readme_file:
readme_file.write(readme_md)
ReadMe.from_readme(path, example_yaml_structure, suppress_parsing_errors=True)
| 0
|
hf_public_repos/datasets
|
hf_public_repos/datasets/tests/test_py_utils.py
|
import time
from dataclasses import dataclass
from multiprocessing import Pool
from unittest import TestCase
from unittest.mock import patch
import multiprocess
import numpy as np
import pytest
from datasets.utils.py_utils import (
NestedDataStructure,
asdict,
iflatmap_unordered,
map_nested,
temp_seed,
temporary_assignment,
zip_dict,
)
from .utils import require_tf, require_torch
def np_sum(x): # picklable for multiprocessing
return x.sum()
def add_one(i): # picklable for multiprocessing
return i + 1
@dataclass
class A:
x: int
y: str
class PyUtilsTest(TestCase):
def test_map_nested(self):
s1 = {}
s2 = []
s3 = 1
s4 = [1, 2]
s5 = {"a": 1, "b": 2}
s6 = {"a": [1, 2], "b": [3, 4]}
s7 = {"a": {"1": 1}, "b": 2}
s8 = {"a": 1, "b": 2, "c": 3, "d": 4}
expected_map_nested_s1 = {}
expected_map_nested_s2 = []
expected_map_nested_s3 = 2
expected_map_nested_s4 = [2, 3]
expected_map_nested_s5 = {"a": 2, "b": 3}
expected_map_nested_s6 = {"a": [2, 3], "b": [4, 5]}
expected_map_nested_s7 = {"a": {"1": 2}, "b": 3}
expected_map_nested_s8 = {"a": 2, "b": 3, "c": 4, "d": 5}
self.assertEqual(map_nested(add_one, s1), expected_map_nested_s1)
self.assertEqual(map_nested(add_one, s2), expected_map_nested_s2)
self.assertEqual(map_nested(add_one, s3), expected_map_nested_s3)
self.assertEqual(map_nested(add_one, s4), expected_map_nested_s4)
self.assertEqual(map_nested(add_one, s5), expected_map_nested_s5)
self.assertEqual(map_nested(add_one, s6), expected_map_nested_s6)
self.assertEqual(map_nested(add_one, s7), expected_map_nested_s7)
self.assertEqual(map_nested(add_one, s8), expected_map_nested_s8)
num_proc = 2
self.assertEqual(map_nested(add_one, s1, num_proc=num_proc), expected_map_nested_s1)
self.assertEqual(map_nested(add_one, s2, num_proc=num_proc), expected_map_nested_s2)
self.assertEqual(map_nested(add_one, s3, num_proc=num_proc), expected_map_nested_s3)
self.assertEqual(map_nested(add_one, s4, num_proc=num_proc), expected_map_nested_s4)
self.assertEqual(map_nested(add_one, s5, num_proc=num_proc), expected_map_nested_s5)
self.assertEqual(map_nested(add_one, s6, num_proc=num_proc), expected_map_nested_s6)
self.assertEqual(map_nested(add_one, s7, num_proc=num_proc), expected_map_nested_s7)
self.assertEqual(map_nested(add_one, s8, num_proc=num_proc), expected_map_nested_s8)
sn1 = {"a": np.eye(2), "b": np.zeros(3), "c": np.ones(2)}
expected_map_nested_sn1_sum = {"a": 2, "b": 0, "c": 2}
expected_map_nested_sn1_int = {
"a": np.eye(2).astype(int),
"b": np.zeros(3).astype(int),
"c": np.ones(2).astype(int),
}
self.assertEqual(map_nested(np_sum, sn1, map_numpy=False), expected_map_nested_sn1_sum)
self.assertEqual(
{k: v.tolist() for k, v in map_nested(int, sn1, map_numpy=True).items()},
{k: v.tolist() for k, v in expected_map_nested_sn1_int.items()},
)
self.assertEqual(map_nested(np_sum, sn1, map_numpy=False, num_proc=num_proc), expected_map_nested_sn1_sum)
self.assertEqual(
{k: v.tolist() for k, v in map_nested(int, sn1, map_numpy=True, num_proc=num_proc).items()},
{k: v.tolist() for k, v in expected_map_nested_sn1_int.items()},
)
with self.assertRaises(AttributeError): # can't pickle a local lambda
map_nested(lambda x: x + 1, sn1, num_proc=num_proc)
def test_zip_dict(self):
d1 = {"a": 1, "b": 2}
d2 = {"a": 3, "b": 4}
d3 = {"a": 5, "b": 6}
expected_zip_dict_result = sorted([("a", (1, 3, 5)), ("b", (2, 4, 6))])
self.assertEqual(sorted(zip_dict(d1, d2, d3)), expected_zip_dict_result)
def test_temporary_assignment(self):
class Foo:
my_attr = "bar"
foo = Foo()
self.assertEqual(foo.my_attr, "bar")
with temporary_assignment(foo, "my_attr", "BAR"):
self.assertEqual(foo.my_attr, "BAR")
self.assertEqual(foo.my_attr, "bar")
@pytest.mark.parametrize(
"iterable_length, num_proc, expected_num_proc",
[
(1, None, 1),
(1, 1, 1),
(2, None, 1),
(2, 1, 1),
(2, 2, 1),
(2, 3, 1),
(3, 2, 1),
(16, 16, 16),
(16, 17, 16),
(17, 16, 16),
],
)
def test_map_nested_num_proc(iterable_length, num_proc, expected_num_proc):
with patch("datasets.utils.py_utils._single_map_nested") as mock_single_map_nested, patch(
"datasets.parallel.parallel.Pool"
) as mock_multiprocessing_pool:
data_struct = {f"{i}": i for i in range(iterable_length)}
_ = map_nested(lambda x: x + 10, data_struct, num_proc=num_proc, parallel_min_length=16)
if expected_num_proc == 1:
assert mock_single_map_nested.called
assert not mock_multiprocessing_pool.called
else:
assert not mock_single_map_nested.called
assert mock_multiprocessing_pool.called
assert mock_multiprocessing_pool.call_args[0][0] == expected_num_proc
class TempSeedTest(TestCase):
@require_tf
def test_tensorflow(self):
import tensorflow as tf
from tensorflow.keras import layers
model = layers.Dense(2)
def gen_random_output():
x = tf.random.uniform((1, 3))
return model(x).numpy()
with temp_seed(42, set_tensorflow=True):
out1 = gen_random_output()
with temp_seed(42, set_tensorflow=True):
out2 = gen_random_output()
out3 = gen_random_output()
np.testing.assert_equal(out1, out2)
self.assertGreater(np.abs(out1 - out3).sum(), 0)
@require_torch
def test_torch(self):
import torch
def gen_random_output():
model = torch.nn.Linear(3, 2)
x = torch.rand(1, 3)
return model(x).detach().numpy()
with temp_seed(42, set_pytorch=True):
out1 = gen_random_output()
with temp_seed(42, set_pytorch=True):
out2 = gen_random_output()
out3 = gen_random_output()
np.testing.assert_equal(out1, out2)
self.assertGreater(np.abs(out1 - out3).sum(), 0)
def test_numpy(self):
def gen_random_output():
return np.random.rand(1, 3)
with temp_seed(42):
out1 = gen_random_output()
with temp_seed(42):
out2 = gen_random_output()
out3 = gen_random_output()
np.testing.assert_equal(out1, out2)
self.assertGreater(np.abs(out1 - out3).sum(), 0)
@pytest.mark.parametrize("input_data", [{}])
def test_nested_data_structure_data(input_data):
output_data = NestedDataStructure(input_data).data
assert output_data == input_data
@pytest.mark.parametrize(
"data, expected_output",
[
({}, []),
([], []),
("foo", ["foo"]),
(["foo", "bar"], ["foo", "bar"]),
([["foo", "bar"]], ["foo", "bar"]),
([[["foo"], ["bar"]]], ["foo", "bar"]),
([[["foo"], "bar"]], ["foo", "bar"]),
({"a": 1, "b": 2}, [1, 2]),
({"a": [1, 2], "b": [3, 4]}, [1, 2, 3, 4]),
({"a": [[1, 2]], "b": [[3, 4]]}, [1, 2, 3, 4]),
({"a": [[1, 2]], "b": [3, 4]}, [1, 2, 3, 4]),
({"a": [[[1], [2]]], "b": [[[3], [4]]]}, [1, 2, 3, 4]),
({"a": [[[1], [2]]], "b": [[3, 4]]}, [1, 2, 3, 4]),
({"a": [[[1], [2]]], "b": [3, 4]}, [1, 2, 3, 4]),
({"a": [[[1], [2]]], "b": [3, [4]]}, [1, 2, 3, 4]),
({"a": {"1": 1}, "b": 2}, [1, 2]),
({"a": {"1": [1]}, "b": 2}, [1, 2]),
({"a": {"1": [1]}, "b": [2]}, [1, 2]),
],
)
def test_flatten(data, expected_output):
output = NestedDataStructure(data).flatten()
assert output == expected_output
def test_asdict():
input = A(x=1, y="foobar")
expected_output = {"x": 1, "y": "foobar"}
assert asdict(input) == expected_output
input = {"a": {"b": A(x=10, y="foo")}, "c": [A(x=20, y="bar")]}
expected_output = {"a": {"b": {"x": 10, "y": "foo"}}, "c": [{"x": 20, "y": "bar"}]}
assert asdict(input) == expected_output
with pytest.raises(TypeError):
asdict([1, A(x=10, y="foo")])
def _split_text(text: str):
return text.split()
def _2seconds_generator_of_2items_with_timing(content):
yield (time.time(), content)
time.sleep(2)
yield (time.time(), content)
def test_iflatmap_unordered():
with Pool(2) as pool:
out = list(iflatmap_unordered(pool, _split_text, kwargs_iterable=[{"text": "hello there"}] * 10))
assert out.count("hello") == 10
assert out.count("there") == 10
assert len(out) == 20
# check multiprocess from pathos (uses dill for pickling)
with multiprocess.Pool(2) as pool:
out = list(iflatmap_unordered(pool, _split_text, kwargs_iterable=[{"text": "hello there"}] * 10))
assert out.count("hello") == 10
assert out.count("there") == 10
assert len(out) == 20
# check that we get items as fast as possible
with Pool(2) as pool:
out = []
for yield_time, content in iflatmap_unordered(
pool, _2seconds_generator_of_2items_with_timing, kwargs_iterable=[{"content": "a"}, {"content": "b"}]
):
assert yield_time < time.time() + 0.1, "we should each item directly after it was yielded"
out.append(content)
assert out.count("a") == 2
assert out.count("b") == 2
assert len(out) == 4
| 0
|
hf_public_repos/datasets
|
hf_public_repos/datasets/tests/test_info_utils.py
|
import pytest
import datasets.config
from datasets.utils.info_utils import is_small_dataset
@pytest.mark.parametrize("dataset_size", [None, 400 * 2**20, 600 * 2**20])
@pytest.mark.parametrize("input_in_memory_max_size", ["default", 0, 100 * 2**20, 900 * 2**20])
def test_is_small_dataset(dataset_size, input_in_memory_max_size, monkeypatch):
if input_in_memory_max_size != "default":
monkeypatch.setattr(datasets.config, "IN_MEMORY_MAX_SIZE", input_in_memory_max_size)
in_memory_max_size = datasets.config.IN_MEMORY_MAX_SIZE
if input_in_memory_max_size == "default":
assert in_memory_max_size == 0
else:
assert in_memory_max_size == input_in_memory_max_size
if dataset_size and in_memory_max_size:
expected = dataset_size < in_memory_max_size
else:
expected = False
result = is_small_dataset(dataset_size)
assert result == expected
| 0
|
hf_public_repos/datasets
|
hf_public_repos/datasets/tests/test_data_files.py
|
import copy
import os
from pathlib import Path
from typing import List
from unittest.mock import patch
import fsspec
import pytest
from fsspec.registry import _registry as _fsspec_registry
from fsspec.spec import AbstractFileSystem
from datasets.data_files import (
DataFilesDict,
DataFilesList,
_get_data_files_patterns,
_get_metadata_files_patterns,
_is_inside_unrequested_special_dir,
_is_unrequested_hidden_file_or_is_inside_unrequested_hidden_dir,
get_data_patterns,
resolve_pattern,
)
from datasets.fingerprint import Hasher
_TEST_PATTERNS = ["*", "**", "**/*", "*.txt", "data/*", "**/*.txt", "**/train.txt"]
_FILES_TO_IGNORE = {".dummy", "README.md", "dummy_data.zip", "dataset_infos.json"}
_DIRS_TO_IGNORE = {"data/.dummy_subdir", "__pycache__"}
_TEST_PATTERNS_SIZES = {
"*": 0,
"**": 4,
"**/*": 4,
"*.txt": 0,
"data/*": 2,
"data/**": 4,
"**/*.txt": 4,
"**/train.txt": 2,
}
_TEST_URL = "https://raw.githubusercontent.com/huggingface/datasets/9675a5a1e7b99a86f9c250f6ea5fa5d1e6d5cc7d/setup.py"
@pytest.fixture
def complex_data_dir(tmp_path):
data_dir = tmp_path / "complex_data_dir"
data_dir.mkdir()
(data_dir / "data").mkdir()
with open(data_dir / "data" / "train.txt", "w") as f:
f.write("foo\n" * 10)
with open(data_dir / "data" / "test.txt", "w") as f:
f.write("bar\n" * 10)
with open(data_dir / "README.md", "w") as f:
f.write("This is a readme")
with open(data_dir / ".dummy", "w") as f:
f.write("this is a dummy file that is not a data file")
(data_dir / "data" / "subdir").mkdir()
with open(data_dir / "data" / "subdir" / "train.txt", "w") as f:
f.write("foo\n" * 10)
with open(data_dir / "data" / "subdir" / "test.txt", "w") as f:
f.write("bar\n" * 10)
(data_dir / "data" / ".dummy_subdir").mkdir()
with open(data_dir / "data" / ".dummy_subdir" / "train.txt", "w") as f:
f.write("foo\n" * 10)
with open(data_dir / "data" / ".dummy_subdir" / "test.txt", "w") as f:
f.write("bar\n" * 10)
(data_dir / "__pycache__").mkdir()
with open(data_dir / "__pycache__" / "script.py", "w") as f:
f.write("foo\n" * 10)
return str(data_dir)
def is_relative_to(path, *other):
# A built-in method in Python 3.9+
try:
path.relative_to(*other)
return True
except ValueError:
return False
@pytest.fixture
def pattern_results(complex_data_dir):
# We use fsspec glob as a reference for data files resolution from patterns.
# This is the same as dask for example.
#
# /!\ Here are some behaviors specific to fsspec glob that are different from glob.glob, Path.glob, Path.match or fnmatch:
# - '*' matches only first level items
# - '**' matches all items
# - '**/*' matches all at least second level items
#
# More generally:
# - '*' matches any character except a forward-slash (to match just the file or directory name)
# - '**' matches any character including a forward-slash /
return {
pattern: sorted(
Path(os.path.abspath(path)).as_posix()
for path in fsspec.filesystem("file").glob(os.path.join(complex_data_dir, pattern))
if Path(path).name not in _FILES_TO_IGNORE
and not any(
is_relative_to(Path(path), os.path.join(complex_data_dir, dir_path)) for dir_path in _DIRS_TO_IGNORE
)
and Path(path).is_file()
)
for pattern in _TEST_PATTERNS
}
@pytest.fixture
def hub_dataset_repo_path(tmpfs, complex_data_dir):
for path in Path(complex_data_dir).rglob("*"):
if path.is_file():
with tmpfs.open(path.relative_to(complex_data_dir).as_posix(), "wb") as f:
f.write(path.read_bytes())
yield "tmp://"
@pytest.fixture
def hub_dataset_repo_patterns_results(hub_dataset_repo_path, complex_data_dir, pattern_results):
return {
pattern: [
hub_dataset_repo_path + Path(path).relative_to(complex_data_dir).as_posix()
for path in pattern_results[pattern]
]
for pattern in pattern_results
}
def test_is_inside_unrequested_special_dir(complex_data_dir, pattern_results):
# usual patterns outside special dir work fine
for pattern, result in pattern_results.items():
if result:
matched_rel_path = str(Path(result[0]).relative_to(complex_data_dir))
assert _is_inside_unrequested_special_dir(matched_rel_path, pattern) is False
# check behavior for special dir
f = _is_inside_unrequested_special_dir
assert f("__pycache__/b.txt", "**") is True
assert f("__pycache__/b.txt", "*/b.txt") is True
assert f("__pycache__/b.txt", "__pycache__/*") is False
assert f("__pycache__/__b.txt", "__pycache__/*") is False
assert f("__pycache__/__b.txt", "__*/*") is False
assert f("__b.txt", "*") is False
def test_is_unrequested_hidden_file_or_is_inside_unrequested_hidden_dir(complex_data_dir, pattern_results):
# usual patterns outside hidden dir work fine
for pattern, result in pattern_results.items():
if result:
matched_rel_path = str(Path(result[0]).relative_to(complex_data_dir))
assert _is_inside_unrequested_special_dir(matched_rel_path, pattern) is False
# check behavior for hidden dir and file
f = _is_unrequested_hidden_file_or_is_inside_unrequested_hidden_dir
assert f(".hidden_file.txt", "**") is True
assert f(".hidden_file.txt", ".*") is False
assert f(".hidden_dir/a.txt", "**") is True
assert f(".hidden_dir/a.txt", ".*/*") is False
assert f(".hidden_dir/a.txt", ".hidden_dir/*") is False
assert f(".hidden_dir/.hidden_file.txt", "**") is True
assert f(".hidden_dir/.hidden_file.txt", ".*/*") is True
assert f(".hidden_dir/.hidden_file.txt", ".*/.*") is False
assert f(".hidden_dir/.hidden_file.txt", ".hidden_dir/*") is True
assert f(".hidden_dir/.hidden_file.txt", ".hidden_dir/.*") is False
@pytest.mark.parametrize("pattern", _TEST_PATTERNS)
def test_pattern_results_fixture(pattern_results, pattern):
assert len(pattern_results[pattern]) == _TEST_PATTERNS_SIZES[pattern]
assert all(Path(path).is_file() for path in pattern_results[pattern])
@pytest.mark.parametrize("pattern", _TEST_PATTERNS)
def test_resolve_pattern_locally(complex_data_dir, pattern, pattern_results):
try:
resolved_data_files = resolve_pattern(pattern, complex_data_dir)
assert sorted(str(f) for f in resolved_data_files) == pattern_results[pattern]
except FileNotFoundError:
assert len(pattern_results[pattern]) == 0
def test_resolve_pattern_locally_with_dot_in_base_path(complex_data_dir):
base_path_with_dot = os.path.join(complex_data_dir, "data", ".dummy_subdir")
resolved_data_files = resolve_pattern(os.path.join(base_path_with_dot, "train.txt"), base_path_with_dot)
assert len(resolved_data_files) == 1
def test_resolve_pattern_locally_with_absolute_path(tmp_path, complex_data_dir):
abs_path = os.path.join(complex_data_dir, "data", "train.txt")
resolved_data_files = resolve_pattern(abs_path, str(tmp_path / "blabla"))
assert len(resolved_data_files) == 1
def test_resolve_pattern_locally_with_double_dots(tmp_path, complex_data_dir):
path_with_double_dots = os.path.join(complex_data_dir, "data", "subdir", "..", "train.txt")
resolved_data_files = resolve_pattern(path_with_double_dots, str(tmp_path / "blabla"))
assert len(resolved_data_files) == 1
def test_resolve_pattern_locally_returns_hidden_file_only_if_requested(complex_data_dir):
with pytest.raises(FileNotFoundError):
resolve_pattern("*dummy", complex_data_dir)
resolved_data_files = resolve_pattern(".dummy", complex_data_dir)
assert len(resolved_data_files) == 1
def test_resolve_pattern_locally_hidden_base_path(tmp_path):
hidden = tmp_path / ".test_hidden_base_path"
hidden.mkdir()
(tmp_path / ".test_hidden_base_path" / "a.txt").touch()
resolved_data_files = resolve_pattern("*", str(hidden))
assert len(resolved_data_files) == 1
def test_resolve_pattern_locallyreturns_hidden_dir_only_if_requested(complex_data_dir):
with pytest.raises(FileNotFoundError):
resolve_pattern("data/*dummy_subdir/train.txt", complex_data_dir)
resolved_data_files = resolve_pattern("data/.dummy_subdir/train.txt", complex_data_dir)
assert len(resolved_data_files) == 1
resolved_data_files = resolve_pattern("*/.dummy_subdir/train.txt", complex_data_dir)
assert len(resolved_data_files) == 1
def test_resolve_pattern_locally_returns_special_dir_only_if_requested(complex_data_dir):
with pytest.raises(FileNotFoundError):
resolve_pattern("data/*dummy_subdir/train.txt", complex_data_dir)
resolved_data_files = resolve_pattern("data/.dummy_subdir/train.txt", complex_data_dir)
assert len(resolved_data_files) == 1
resolved_data_files = resolve_pattern("*/.dummy_subdir/train.txt", complex_data_dir)
assert len(resolved_data_files) == 1
def test_resolve_pattern_locally_special_base_path(tmp_path):
special = tmp_path / "__test_special_base_path__"
special.mkdir()
(tmp_path / "__test_special_base_path__" / "a.txt").touch()
resolved_data_files = resolve_pattern("*", str(special))
assert len(resolved_data_files) == 1
@pytest.mark.parametrize("pattern,size,extensions", [("**", 4, [".txt"]), ("**", 4, None), ("**", 0, [".blablabla"])])
def test_resolve_pattern_locally_with_extensions(complex_data_dir, pattern, size, extensions):
if size > 0:
resolved_data_files = resolve_pattern(pattern, complex_data_dir, allowed_extensions=extensions)
assert len(resolved_data_files) == size
else:
with pytest.raises(FileNotFoundError):
resolve_pattern(pattern, complex_data_dir, allowed_extensions=extensions)
def test_fail_resolve_pattern_locally(complex_data_dir):
with pytest.raises(FileNotFoundError):
resolve_pattern(complex_data_dir, ["blablabla"])
@pytest.mark.skipif(os.name == "nt", reason="Windows does not support symlinks in the default mode")
def test_resolve_pattern_locally_does_not_resolve_symbolic_links(tmp_path, complex_data_dir):
(tmp_path / "train_data_symlink.txt").symlink_to(os.path.join(complex_data_dir, "data", "train.txt"))
resolved_data_files = resolve_pattern("train_data_symlink.txt", str(tmp_path))
assert len(resolved_data_files) == 1
assert Path(resolved_data_files[0]) == tmp_path / "train_data_symlink.txt"
def test_resolve_pattern_locally_sorted_files(tmp_path_factory):
path = str(tmp_path_factory.mktemp("unsorted_text_files"))
unsorted_names = ["0.txt", "2.txt", "3.txt"]
for name in unsorted_names:
with open(os.path.join(path, name), "w"):
pass
resolved_data_files = resolve_pattern("*", path)
resolved_names = [os.path.basename(data_file) for data_file in resolved_data_files]
assert resolved_names == sorted(unsorted_names)
@pytest.mark.parametrize("pattern", _TEST_PATTERNS)
def test_resolve_pattern_in_dataset_repository(hub_dataset_repo_path, pattern, hub_dataset_repo_patterns_results):
try:
resolved_data_files = resolve_pattern(pattern, hub_dataset_repo_path)
assert sorted(str(f) for f in resolved_data_files) == hub_dataset_repo_patterns_results[pattern]
except FileNotFoundError:
assert len(hub_dataset_repo_patterns_results[pattern]) == 0
@pytest.mark.parametrize(
"pattern,size,base_path", [("**", 4, None), ("**", 4, "data"), ("**", 2, "data/subdir"), ("**", 0, "data/subdir2")]
)
def test_resolve_pattern_in_dataset_repository_with_base_path(hub_dataset_repo_path, pattern, size, base_path):
base_path = hub_dataset_repo_path + (base_path or "")
if size > 0:
resolved_data_files = resolve_pattern(pattern, base_path)
assert len(resolved_data_files) == size
else:
with pytest.raises(FileNotFoundError):
resolve_pattern(pattern, base_path)
@pytest.mark.parametrize("pattern,size,extensions", [("**", 4, [".txt"]), ("**", 4, None), ("**", 0, [".blablabla"])])
def test_resolve_pattern_in_dataset_repository_with_extensions(hub_dataset_repo_path, pattern, size, extensions):
if size > 0:
resolved_data_files = resolve_pattern(pattern, hub_dataset_repo_path, allowed_extensions=extensions)
assert len(resolved_data_files) == size
else:
with pytest.raises(FileNotFoundError):
resolved_data_files = resolve_pattern(pattern, hub_dataset_repo_path, allowed_extensions=extensions)
def test_fail_resolve_pattern_in_dataset_repository(hub_dataset_repo_path):
with pytest.raises(FileNotFoundError):
resolve_pattern("blablabla", hub_dataset_repo_path)
def test_resolve_pattern_in_dataset_repository_returns_hidden_file_only_if_requested(hub_dataset_repo_path):
with pytest.raises(FileNotFoundError):
resolve_pattern("*dummy", hub_dataset_repo_path)
resolved_data_files = resolve_pattern(".dummy", hub_dataset_repo_path)
assert len(resolved_data_files) == 1
def test_resolve_pattern_in_dataset_repository_hidden_base_path(tmpfs):
tmpfs.touch(".hidden/a.txt")
resolved_data_files = resolve_pattern("*", base_path="tmp://.hidden")
assert len(resolved_data_files) == 1
def test_resolve_pattern_in_dataset_repository_returns_hidden_dir_only_if_requested(hub_dataset_repo_path):
with pytest.raises(FileNotFoundError):
resolve_pattern("data/*dummy_subdir/train.txt", hub_dataset_repo_path)
resolved_data_files = resolve_pattern("data/.dummy_subdir/train.txt", hub_dataset_repo_path)
assert len(resolved_data_files) == 1
resolved_data_files = resolve_pattern("*/.dummy_subdir/train.txt", hub_dataset_repo_path)
assert len(resolved_data_files) == 1
def test_resolve_pattern_in_dataset_repository_returns_special_dir_only_if_requested(hub_dataset_repo_path):
with pytest.raises(FileNotFoundError):
resolve_pattern("data/*dummy_subdir/train.txt", hub_dataset_repo_path)
resolved_data_files = resolve_pattern("data/.dummy_subdir/train.txt", hub_dataset_repo_path)
assert len(resolved_data_files) == 1
resolved_data_files = resolve_pattern("*/.dummy_subdir/train.txt", hub_dataset_repo_path)
assert len(resolved_data_files) == 1
def test_resolve_pattern_in_dataset_repository_special_base_path(tmpfs):
tmpfs.touch("__special__/a.txt")
resolved_data_files = resolve_pattern("*", base_path="tmp://__special__")
assert len(resolved_data_files) == 1
@pytest.fixture
def dummy_fs():
DummyTestFS = mock_fs(["train.txt", "test.txt"])
_fsspec_registry["mock"] = DummyTestFS
_fsspec_registry["dummy"] = DummyTestFS
yield
del _fsspec_registry["mock"]
del _fsspec_registry["dummy"]
def test_resolve_pattern_fs(dummy_fs):
resolved_data_files = resolve_pattern("mock://train.txt", base_path="")
assert resolved_data_files == ["mock://train.txt"]
@pytest.mark.parametrize("pattern", _TEST_PATTERNS)
def test_DataFilesList_from_patterns_in_dataset_repository_(
hub_dataset_repo_path, hub_dataset_repo_patterns_results, pattern
):
try:
data_files_list = DataFilesList.from_patterns([pattern], hub_dataset_repo_path)
assert sorted(data_files_list) == hub_dataset_repo_patterns_results[pattern]
assert len(data_files_list.origin_metadata) == len(data_files_list)
except FileNotFoundError:
assert len(hub_dataset_repo_patterns_results[pattern]) == 0
def test_DataFilesList_from_patterns_locally_with_extra_files(complex_data_dir, text_file):
data_files_list = DataFilesList.from_patterns([_TEST_URL, text_file.as_posix()], complex_data_dir)
assert list(data_files_list) == [_TEST_URL, text_file.as_posix()]
assert len(data_files_list.origin_metadata) == 2
def test_DataFilesList_from_patterns_raises_FileNotFoundError(complex_data_dir):
with pytest.raises(FileNotFoundError):
DataFilesList.from_patterns(["file_that_doesnt_exist.txt"], complex_data_dir)
class TestDataFilesDict:
def test_key_order_after_copy(self):
data_files = DataFilesDict({"train": "train.csv", "test": "test.csv"})
copied_data_files = copy.deepcopy(data_files)
assert list(copied_data_files.keys()) == list(data_files.keys()) # test split order with list()
@pytest.mark.parametrize("pattern", _TEST_PATTERNS)
def test_DataFilesDict_from_patterns_in_dataset_repository(
hub_dataset_repo_path, hub_dataset_repo_patterns_results, pattern
):
split_name = "train"
try:
data_files = DataFilesDict.from_patterns({split_name: [pattern]}, hub_dataset_repo_path)
assert all(isinstance(data_files_list, DataFilesList) for data_files_list in data_files.values())
assert sorted(data_files[split_name]) == hub_dataset_repo_patterns_results[pattern]
except FileNotFoundError:
assert len(hub_dataset_repo_patterns_results[pattern]) == 0
@pytest.mark.parametrize(
"pattern,size,base_path,split_name",
[
("**", 4, None, "train"),
("**", 4, "data", "train"),
("**", 2, "data/subdir", "train"),
("**train*", 1, "data/subdir", "train"),
("**test*", 1, "data/subdir", "test"),
("**", 0, "data/subdir2", "train"),
],
)
def test_DataFilesDict_from_patterns_in_dataset_repository_with_base_path(
hub_dataset_repo_path, pattern, size, base_path, split_name
):
base_path = hub_dataset_repo_path + (base_path or "")
if size > 0:
data_files = DataFilesDict.from_patterns({split_name: [pattern]}, base_path=base_path)
assert len(data_files[split_name]) == size
else:
with pytest.raises(FileNotFoundError):
resolve_pattern(pattern, base_path)
@pytest.mark.parametrize("pattern", _TEST_PATTERNS)
def test_DataFilesDict_from_patterns_locally(complex_data_dir, pattern_results, pattern):
split_name = "train"
try:
data_files = DataFilesDict.from_patterns({split_name: [pattern]}, complex_data_dir)
assert all(isinstance(data_files_list, DataFilesList) for data_files_list in data_files.values())
assert sorted(data_files[split_name]) == pattern_results[pattern]
except FileNotFoundError:
assert len(pattern_results[pattern]) == 0
def test_DataFilesDict_from_patterns_in_dataset_repository_hashing(hub_dataset_repo_path):
patterns = {"train": ["**/train.txt"], "test": ["**/test.txt"]}
data_files1 = DataFilesDict.from_patterns(patterns, hub_dataset_repo_path)
data_files2 = DataFilesDict.from_patterns(patterns, hub_dataset_repo_path)
assert Hasher.hash(data_files1) == Hasher.hash(data_files2)
data_files2 = DataFilesDict(sorted(data_files1.items(), reverse=True))
assert Hasher.hash(data_files1) == Hasher.hash(data_files2)
patterns2 = {"train": ["data/**train.txt"], "test": ["data/**test.txt"]}
data_files2 = DataFilesDict.from_patterns(patterns2, hub_dataset_repo_path)
assert Hasher.hash(data_files1) == Hasher.hash(data_files2)
patterns2 = {"train": ["data/**train.txt"], "test": ["data/**train.txt"]}
data_files2 = DataFilesDict.from_patterns(patterns2, hub_dataset_repo_path)
assert Hasher.hash(data_files1) != Hasher.hash(data_files2)
# the tmpfs used to mock the hub repo is based on a local directory
# therefore os.stat is used to get the mtime of the data files
with patch("os.stat", return_value=os.stat(__file__)):
data_files2 = DataFilesDict.from_patterns(patterns, hub_dataset_repo_path)
assert Hasher.hash(data_files1) != Hasher.hash(data_files2)
def test_DataFilesDict_from_patterns_locally_or_remote_hashing(text_file):
patterns = {"train": [_TEST_URL], "test": [str(text_file)]}
data_files1 = DataFilesDict.from_patterns(patterns)
data_files2 = DataFilesDict.from_patterns(patterns)
assert Hasher.hash(data_files1) == Hasher.hash(data_files2)
data_files2 = DataFilesDict(sorted(data_files1.items(), reverse=True))
assert Hasher.hash(data_files1) == Hasher.hash(data_files2)
patterns2 = {"train": [_TEST_URL], "test": [_TEST_URL]}
data_files2 = DataFilesDict.from_patterns(patterns2)
assert Hasher.hash(data_files1) != Hasher.hash(data_files2)
with patch("fsspec.implementations.http._file_info", return_value={}):
data_files2 = DataFilesDict.from_patterns(patterns)
assert Hasher.hash(data_files1) != Hasher.hash(data_files2)
with patch("os.stat", return_value=os.stat(__file__)):
data_files2 = DataFilesDict.from_patterns(patterns)
assert Hasher.hash(data_files1) != Hasher.hash(data_files2)
def mock_fs(file_paths: List[str]):
"""
Set up a mock filesystem for fsspec containing the provided files
Example:
```py
>>> DummyTestFS = mock_fs(["data/train.txt", "data.test.txt"])
>>> fs = DummyTestFS()
>>> assert fsspec.get_filesystem_class("mock").__name__ == "DummyTestFS"
>>> assert type(fs).__name__ == "DummyTestFS"
>>> print(fs.glob("**"))
["data", "data/train.txt", "data.test.txt"]
```
"""
file_paths = [file_path.split("://")[-1] for file_path in file_paths]
dir_paths = {
"/".join(file_path.split("/")[: i + 1]) for file_path in file_paths for i in range(file_path.count("/"))
}
fs_contents = [{"name": dir_path, "type": "directory"} for dir_path in dir_paths] + [
{"name": file_path, "type": "file", "size": 10} for file_path in file_paths
]
class DummyTestFS(AbstractFileSystem):
protocol = ("mock", "dummy")
_fs_contents = fs_contents
def ls(self, path, detail=True, refresh=True, **kwargs):
if kwargs.pop("strip_proto", True):
path = self._strip_protocol(path)
files = not refresh and self._ls_from_cache(path)
if not files:
files = [file for file in self._fs_contents if path == self._parent(file["name"])]
files.sort(key=lambda file: file["name"])
self.dircache[path.rstrip("/")] = files
if detail:
return files
return [file["name"] for file in files]
return DummyTestFS
@pytest.mark.parametrize("base_path", ["", "mock://", "my_dir"])
@pytest.mark.parametrize(
"data_file_per_split",
[
# === Main cases ===
# file named after split at the root
{"train": "train.txt", "validation": "valid.txt", "test": "test.txt"},
# file named after split in a directory
{
"train": "data/train.txt",
"validation": "data/valid.txt",
"test": "data/test.txt",
},
# directory named after split
{
"train": "train/split.txt",
"validation": "valid/split.txt",
"test": "test/split.txt",
},
# sharded splits
{
"train": [f"data/train_{i}.txt" for i in range(3)],
"validation": [f"data/validation_{i}.txt" for i in range(3)],
"test": [f"data/test_{i}.txt" for i in range(3)],
},
# sharded splits with standard format (+ custom split name)
{
"train": [f"data/train-0000{i}-of-00003.txt" for i in range(3)],
"validation": [f"data/validation-0000{i}-of-00003.txt" for i in range(3)],
"test": [f"data/test-0000{i}-of-00003.txt" for i in range(3)],
"random": [f"data/random-0000{i}-of-00003.txt" for i in range(3)],
},
# === Secondary cases ===
# Default to train split
{"train": "dataset.txt"},
{"train": "data/dataset.txt"},
{"train": ["data/image.jpg", "metadata.jsonl"]},
{"train": ["data/image.jpg", "metadata.csv"]},
# With prefix or suffix in directory or file names
{"train": "my_train_dir/dataset.txt"},
{"train": "data/my_train_file.txt"},
{"test": "my_test_dir/dataset.txt"},
{"test": "data/my_test_file.txt"},
{"validation": "my_validation_dir/dataset.txt"},
{"validation": "data/my_validation_file.txt"},
# With test<>eval aliases
{"test": "eval.txt"},
{"test": "data/eval.txt"},
{"test": "eval/dataset.txt"},
# With valid<>dev aliases
{"validation": "dev.txt"},
{"validation": "data/dev.txt"},
{"validation": "dev/dataset.txt"},
# With valid<>val aliases
{"validation": "val.txt"},
{"validation": "data/val.txt"},
# With other extensions
{"train": "train.parquet", "validation": "valid.parquet", "test": "test.parquet"},
# With "dev" or "eval" without separators
{"train": "developers_list.txt"},
{"train": "data/seqeval_results.txt"},
{"train": "contest.txt"},
# With supported separators
{"test": "my.test.file.txt"},
{"test": "my-test-file.txt"},
{"test": "my_test_file.txt"},
{"test": "my test file.txt"},
{"test": "test00001.txt"},
],
)
def test_get_data_files_patterns(base_path, data_file_per_split):
data_file_per_split = {k: v if isinstance(v, list) else [v] for k, v in data_file_per_split.items()}
data_file_per_split = {
split: [
base_path + ("/" if base_path and base_path[-1] != "/" else "") + file_path
for file_path in data_file_per_split[split]
]
for split in data_file_per_split
}
file_paths = sum(data_file_per_split.values(), [])
DummyTestFS = mock_fs(file_paths)
fs = DummyTestFS()
def resolver(pattern):
pattern = base_path + ("/" if base_path and base_path[-1] != "/" else "") + pattern
return [
file_path[len(fs._strip_protocol(base_path)) :].lstrip("/")
for file_path in fs.glob(pattern)
if fs.isfile(file_path)
]
patterns_per_split = _get_data_files_patterns(resolver)
assert list(patterns_per_split.keys()) == list(data_file_per_split.keys()) # Test split order with list()
for split, patterns in patterns_per_split.items():
matched = [file_path for pattern in patterns for file_path in resolver(pattern)]
expected = [
fs._strip_protocol(file_path)[len(fs._strip_protocol(base_path)) :].lstrip("/")
for file_path in data_file_per_split[split]
]
assert matched == expected
@pytest.mark.parametrize(
"metadata_files",
[
# metadata files at the root
["metadata.jsonl"],
["metadata.csv"],
# nested metadata files
["metadata.jsonl", "data/metadata.jsonl"],
["metadata.csv", "data/metadata.csv"],
],
)
def test_get_metadata_files_patterns(metadata_files):
DummyTestFS = mock_fs(metadata_files)
fs = DummyTestFS()
def resolver(pattern):
return [file_path for file_path in fs.glob(pattern) if fs.isfile(file_path)]
patterns = _get_metadata_files_patterns(resolver)
matched = [file_path for pattern in patterns for file_path in resolver(pattern)]
assert sorted(matched) == sorted(metadata_files)
def test_get_data_patterns_from_directory_with_the_word_data_twice(tmp_path):
repo_dir = tmp_path / "directory-name-ending-with-the-word-data" # parent directory contains the word "data/"
data_dir = repo_dir / "data"
data_dir.mkdir(parents=True)
data_file = data_dir / "train-00001-of-00009.parquet"
data_file.touch()
data_file_patterns = get_data_patterns(repo_dir.as_posix())
assert data_file_patterns == {"train": ["data/train-[0-9][0-9][0-9][0-9][0-9]-of-[0-9][0-9][0-9][0-9][0-9]*.*"]}
| 0
|
hf_public_repos/datasets
|
hf_public_repos/datasets/tests/test_warnings.py
|
import pytest
from datasets import inspect_metric, list_metrics, load_metric
@pytest.fixture
def mock_emitted_deprecation_warnings(monkeypatch):
monkeypatch.setattr("datasets.utils.deprecation_utils._emitted_deprecation_warnings", set())
# Used by list_metrics
@pytest.fixture
def mock_hfh(monkeypatch):
class MetricMock:
def __init__(self, metric_id):
self.id = metric_id
class HfhMock:
_metrics = [MetricMock(metric_id) for metric_id in ["accuracy", "mse", "precision", "codeparrot/apps_metric"]]
def list_metrics(self):
return self._metrics
monkeypatch.setattr("datasets.inspect.huggingface_hub", HfhMock())
@pytest.mark.parametrize(
"func, args", [(load_metric, ("metrics/mse",)), (list_metrics, ()), (inspect_metric, ("metrics/mse", "tmp_path"))]
)
def test_metric_deprecation_warning(func, args, mock_emitted_deprecation_warnings, mock_hfh, tmp_path):
if "tmp_path" in args:
args = tuple(arg if arg != "tmp_path" else tmp_path for arg in args)
with pytest.warns(FutureWarning, match="https://huggingface.co/docs/evaluate"):
func(*args)
| 0
|
hf_public_repos/datasets
|
hf_public_repos/datasets/tests/test_hf_gcp.py
|
import os
from tempfile import TemporaryDirectory
from unittest import TestCase
import pytest
from absl.testing import parameterized
from datasets import config
from datasets.arrow_reader import HF_GCP_BASE_URL
from datasets.builder import DatasetBuilder
from datasets.dataset_dict import IterableDatasetDict
from datasets.iterable_dataset import IterableDataset
from datasets.load import dataset_module_factory, import_main_class
from datasets.utils.file_utils import cached_path
DATASETS_ON_HF_GCP = [
{"dataset": "wikipedia", "config_name": "20220301.de"},
{"dataset": "wikipedia", "config_name": "20220301.en"},
{"dataset": "wikipedia", "config_name": "20220301.fr"},
{"dataset": "wikipedia", "config_name": "20220301.frr"},
{"dataset": "wikipedia", "config_name": "20220301.it"},
{"dataset": "wikipedia", "config_name": "20220301.simple"},
{"dataset": "snli", "config_name": "plain_text"},
{"dataset": "eli5", "config_name": "LFQA_reddit"},
{"dataset": "wiki40b", "config_name": "en"},
{"dataset": "wiki_dpr", "config_name": "psgs_w100.nq.compressed"},
{"dataset": "wiki_dpr", "config_name": "psgs_w100.nq.no_index"},
{"dataset": "wiki_dpr", "config_name": "psgs_w100.multiset.no_index"},
{"dataset": "natural_questions", "config_name": "default"},
]
def list_datasets_on_hf_gcp_parameters(with_config=True):
if with_config:
return [
{
"testcase_name": d["dataset"] + "/" + d["config_name"],
"dataset": d["dataset"],
"config_name": d["config_name"],
}
for d in DATASETS_ON_HF_GCP
]
else:
return [
{"testcase_name": dataset, "dataset": dataset} for dataset in {d["dataset"] for d in DATASETS_ON_HF_GCP}
]
@parameterized.named_parameters(list_datasets_on_hf_gcp_parameters(with_config=True))
class TestDatasetOnHfGcp(TestCase):
dataset = None
config_name = None
def test_dataset_info_available(self, dataset, config_name):
with TemporaryDirectory() as tmp_dir:
dataset_module = dataset_module_factory(dataset, cache_dir=tmp_dir)
builder_cls = import_main_class(dataset_module.module_path, dataset=True)
builder_instance: DatasetBuilder = builder_cls(
cache_dir=tmp_dir,
config_name=config_name,
hash=dataset_module.hash,
)
dataset_info_url = "/".join(
[
HF_GCP_BASE_URL,
builder_instance._relative_data_dir(with_hash=False).replace(os.sep, "/"),
config.DATASET_INFO_FILENAME,
]
)
datset_info_path = cached_path(dataset_info_url, cache_dir=tmp_dir)
self.assertTrue(os.path.exists(datset_info_path))
@pytest.mark.integration
def test_as_dataset_from_hf_gcs(tmp_path_factory):
tmp_dir = tmp_path_factory.mktemp("test_hf_gcp") / "test_wikipedia_simple"
dataset_module = dataset_module_factory("wikipedia", cache_dir=tmp_dir)
builder_cls = import_main_class(dataset_module.module_path)
builder_instance: DatasetBuilder = builder_cls(
cache_dir=tmp_dir,
config_name="20220301.frr",
hash=dataset_module.hash,
)
# use the HF cloud storage, not the original download_and_prepare that uses apache-beam
builder_instance._download_and_prepare = None
builder_instance.download_and_prepare()
ds = builder_instance.as_dataset()
assert ds
@pytest.mark.integration
def test_as_streaming_dataset_from_hf_gcs(tmp_path):
dataset_module = dataset_module_factory("wikipedia", cache_dir=tmp_path)
builder_cls = import_main_class(dataset_module.module_path, dataset=True)
builder_instance: DatasetBuilder = builder_cls(
cache_dir=tmp_path,
config_name="20220301.frr",
hash=dataset_module.hash,
)
ds = builder_instance.as_streaming_dataset()
assert ds
assert isinstance(ds, IterableDatasetDict)
assert "train" in ds
assert isinstance(ds["train"], IterableDataset)
assert next(iter(ds["train"]))
| 0
|
hf_public_repos/datasets/tests
|
hf_public_repos/datasets/tests/distributed_scripts/run_torch_distributed.py
|
import os
from argparse import ArgumentParser
from typing import List
import torch.utils.data
from datasets import Dataset, IterableDataset
from datasets.distributed import split_dataset_by_node
NUM_SHARDS = 4
NUM_ITEMS_PER_SHARD = 3
class FailedTestError(RuntimeError):
pass
def gen(shards: List[str]):
for shard in shards:
for i in range(NUM_ITEMS_PER_SHARD):
yield {"i": i, "shard": shard}
def main():
rank = int(os.environ["RANK"])
world_size = int(os.environ["WORLD_SIZE"])
parser = ArgumentParser()
parser.add_argument("--streaming", type=bool)
parser.add_argument("--local_rank", type=int)
parser.add_argument("--num_workers", type=int, default=0)
args = parser.parse_args()
streaming = args.streaming
num_workers = args.num_workers
gen_kwargs = {"shards": [f"shard_{shard_idx}" for shard_idx in range(NUM_SHARDS)]}
ds = IterableDataset.from_generator(gen, gen_kwargs=gen_kwargs)
if not streaming:
ds = Dataset.from_list(list(ds))
ds = split_dataset_by_node(ds, rank=rank, world_size=world_size)
dataloader = torch.utils.data.DataLoader(ds, num_workers=num_workers)
full_size = NUM_SHARDS * NUM_ITEMS_PER_SHARD
expected_local_size = full_size // world_size
expected_local_size += int(rank < (full_size % world_size))
local_size = sum(1 for _ in dataloader)
if local_size != expected_local_size:
raise FailedTestError(f"local_size {local_size} != expected_local_size {expected_local_size}")
if __name__ == "__main__":
main()
| 0
|
hf_public_repos/datasets/tests
|
hf_public_repos/datasets/tests/packaged_modules/test_json.py
|
import textwrap
import pyarrow as pa
import pytest
from datasets import Features, Value
from datasets.packaged_modules.json.json import Json
@pytest.fixture
def jsonl_file(tmp_path):
filename = tmp_path / "file.jsonl"
data = textwrap.dedent(
"""\
{"col_1": -1}
{"col_1": 1, "col_2": 2}
{"col_1": 10, "col_2": 20}
"""
)
with open(filename, "w") as f:
f.write(data)
return str(filename)
@pytest.fixture
def jsonl_file_utf16_encoded(tmp_path):
filename = tmp_path / "file_utf16_encoded.jsonl"
data = textwrap.dedent(
"""\
{"col_1": -1}
{"col_1": 1, "col_2": 2}
{"col_1": 10, "col_2": 20}
"""
)
with open(filename, "w", encoding="utf-16") as f:
f.write(data)
return str(filename)
@pytest.fixture
def json_file_with_list_of_dicts(tmp_path):
filename = tmp_path / "file_with_list_of_dicts.json"
data = textwrap.dedent(
"""\
[
{"col_1": -1},
{"col_1": 1, "col_2": 2},
{"col_1": 10, "col_2": 20}
]
"""
)
with open(filename, "w") as f:
f.write(data)
return str(filename)
@pytest.fixture
def json_file_with_list_of_dicts_field(tmp_path):
filename = tmp_path / "file_with_list_of_dicts_field.json"
data = textwrap.dedent(
"""\
{
"field1": 1,
"field2": "aabb",
"field3": [
{"col_1": -1},
{"col_1": 1, "col_2": 2},
{"col_1": 10, "col_2": 20}
]
}
"""
)
with open(filename, "w") as f:
f.write(data)
return str(filename)
@pytest.mark.parametrize(
"file_fixture, config_kwargs",
[
("jsonl_file", {}),
("jsonl_file_utf16_encoded", {"encoding": "utf-16"}),
("json_file_with_list_of_dicts", {}),
("json_file_with_list_of_dicts_field", {"field": "field3"}),
],
)
def test_json_generate_tables(file_fixture, config_kwargs, request):
json = Json(**config_kwargs)
generator = json._generate_tables([[request.getfixturevalue(file_fixture)]])
pa_table = pa.concat_tables([table for _, table in generator])
assert pa_table.to_pydict() == {"col_1": [-1, 1, 10], "col_2": [None, 2, 20]}
@pytest.mark.parametrize(
"file_fixture, config_kwargs",
[
(
"jsonl_file",
{"features": Features({"col_1": Value("int64"), "col_2": Value("int64"), "missing_col": Value("string")})},
),
(
"json_file_with_list_of_dicts",
{"features": Features({"col_1": Value("int64"), "col_2": Value("int64"), "missing_col": Value("string")})},
),
(
"json_file_with_list_of_dicts_field",
{
"field": "field3",
"features": Features(
{"col_1": Value("int64"), "col_2": Value("int64"), "missing_col": Value("string")}
),
},
),
],
)
def test_json_generate_tables_with_missing_features(file_fixture, config_kwargs, request):
json = Json(**config_kwargs)
generator = json._generate_tables([[request.getfixturevalue(file_fixture)]])
pa_table = pa.concat_tables([table for _, table in generator])
assert pa_table.to_pydict() == {"col_1": [-1, 1, 10], "col_2": [None, 2, 20], "missing_col": [None, None, None]}
| 0
|
hf_public_repos/datasets/tests
|
hf_public_repos/datasets/tests/packaged_modules/test_webdataset.py
|
import tarfile
import pytest
from datasets import DownloadManager, Features, Image, Value
from datasets.packaged_modules.webdataset.webdataset import WebDataset
from ..utils import require_pil
@pytest.fixture
def tar_file(tmp_path, image_file, text_file):
filename = tmp_path / "file.tar"
num_examples = 3
with tarfile.open(str(filename), "w") as f:
for example_idx in range(num_examples):
f.add(text_file, f"{example_idx:05d}.txt")
f.add(image_file, f"{example_idx:05d}.jpg")
return str(filename)
@pytest.fixture
def bad_tar_file(tmp_path, image_file, text_file):
filename = tmp_path / "bad_file.tar"
with tarfile.open(str(filename), "w") as f:
f.add(image_file)
f.add(text_file)
return str(filename)
@require_pil
def test_webdataset(tar_file):
import PIL.Image
data_files = {"train": [tar_file]}
webdataset = WebDataset(data_files=data_files)
split_generators = webdataset._split_generators(DownloadManager())
assert webdataset.info.features == Features(
{
"__key__": Value("string"),
"__url__": Value("string"),
"txt": Value("string"),
"jpg": Image(),
}
)
assert len(split_generators) == 1
split_generator = split_generators[0]
assert split_generator.name == "train"
generator = webdataset._generate_examples(**split_generator.gen_kwargs)
_, examples = zip(*generator)
assert len(examples) == 3
assert isinstance(examples[0]["txt"], str)
assert isinstance(examples[0]["jpg"], dict) # keep encoded to avoid unecessary copies
decoded = webdataset.info.features.decode_example(examples[0])
assert isinstance(decoded["txt"], str)
assert isinstance(decoded["jpg"], PIL.Image.Image)
def test_webdataset_errors_on_bad_file(bad_tar_file):
data_files = {"train": [bad_tar_file]}
webdataset = WebDataset(data_files=data_files)
with pytest.raises(ValueError):
webdataset._split_generators(DownloadManager())
| 0
|
hf_public_repos/datasets/tests
|
hf_public_repos/datasets/tests/packaged_modules/test_imagefolder.py
|
import shutil
import textwrap
import numpy as np
import pytest
from datasets import ClassLabel, Features, Image, Value
from datasets.data_files import DataFilesDict, get_data_patterns
from datasets.download.streaming_download_manager import StreamingDownloadManager
from datasets.packaged_modules.imagefolder.imagefolder import ImageFolder
from ..utils import require_pil
@pytest.fixture
def cache_dir(tmp_path):
return str(tmp_path / "imagefolder_cache_dir")
@pytest.fixture
def data_files_with_labels_no_metadata(tmp_path, image_file):
data_dir = tmp_path / "data_files_with_labels_no_metadata"
data_dir.mkdir(parents=True, exist_ok=True)
subdir_class_0 = data_dir / "cat"
subdir_class_0.mkdir(parents=True, exist_ok=True)
subdir_class_1 = data_dir / "dog"
subdir_class_1.mkdir(parents=True, exist_ok=True)
image_filename = subdir_class_0 / "image_cat.jpg"
shutil.copyfile(image_file, image_filename)
image_filename2 = subdir_class_1 / "image_dog.jpg"
shutil.copyfile(image_file, image_filename2)
data_files_with_labels_no_metadata = DataFilesDict.from_patterns(
get_data_patterns(str(data_dir)), data_dir.as_posix()
)
return data_files_with_labels_no_metadata
@pytest.fixture
def image_files_with_labels_and_duplicated_label_key_in_metadata(tmp_path, image_file):
data_dir = tmp_path / "image_files_with_labels_and_label_key_in_metadata"
data_dir.mkdir(parents=True, exist_ok=True)
subdir_class_0 = data_dir / "cat"
subdir_class_0.mkdir(parents=True, exist_ok=True)
subdir_class_1 = data_dir / "dog"
subdir_class_1.mkdir(parents=True, exist_ok=True)
image_filename = subdir_class_0 / "image_cat.jpg"
shutil.copyfile(image_file, image_filename)
image_filename2 = subdir_class_1 / "image_dog.jpg"
shutil.copyfile(image_file, image_filename2)
image_metadata_filename = tmp_path / data_dir / "metadata.jsonl"
image_metadata = textwrap.dedent(
"""\
{"file_name": "cat/image_cat.jpg", "caption": "Nice image of a cat", "label": "Cat"}
{"file_name": "dog/image_dog.jpg", "caption": "Nice image of a dog", "label": "Dog"}
"""
)
with open(image_metadata_filename, "w", encoding="utf-8") as f:
f.write(image_metadata)
return str(image_filename), str(image_filename2), str(image_metadata_filename)
@pytest.fixture
def image_file_with_metadata(tmp_path, image_file):
image_filename = tmp_path / "image_rgb.jpg"
shutil.copyfile(image_file, image_filename)
image_metadata_filename = tmp_path / "metadata.jsonl"
image_metadata = textwrap.dedent(
"""\
{"file_name": "image_rgb.jpg", "caption": "Nice image"}
"""
)
with open(image_metadata_filename, "w", encoding="utf-8") as f:
f.write(image_metadata)
return str(image_filename), str(image_metadata_filename)
@pytest.fixture
def image_files_with_metadata_that_misses_one_image(tmp_path, image_file):
image_filename = tmp_path / "image_rgb.jpg"
shutil.copyfile(image_file, image_filename)
image_filename2 = tmp_path / "image_rgb2.jpg"
shutil.copyfile(image_file, image_filename2)
image_metadata_filename = tmp_path / "metadata.jsonl"
image_metadata = textwrap.dedent(
"""\
{"file_name": "image_rgb.jpg", "caption": "Nice image"}
"""
)
with open(image_metadata_filename, "w", encoding="utf-8") as f:
f.write(image_metadata)
return str(image_filename), str(image_filename2), str(image_metadata_filename)
@pytest.fixture(params=["jsonl", "csv"])
def data_files_with_one_split_and_metadata(request, tmp_path, image_file):
data_dir = tmp_path / "imagefolder_data_dir_with_metadata_one_split"
data_dir.mkdir(parents=True, exist_ok=True)
subdir = data_dir / "subdir"
subdir.mkdir(parents=True, exist_ok=True)
image_filename = data_dir / "image_rgb.jpg"
shutil.copyfile(image_file, image_filename)
image_filename2 = data_dir / "image_rgb2.jpg"
shutil.copyfile(image_file, image_filename2)
image_filename3 = subdir / "image_rgb3.jpg" # in subdir
shutil.copyfile(image_file, image_filename3)
image_metadata_filename = data_dir / f"metadata.{request.param}"
image_metadata = (
textwrap.dedent(
"""\
{"file_name": "image_rgb.jpg", "caption": "Nice image"}
{"file_name": "image_rgb2.jpg", "caption": "Nice second image"}
{"file_name": "subdir/image_rgb3.jpg", "caption": "Nice third image"}
"""
)
if request.param == "jsonl"
else textwrap.dedent(
"""\
file_name,caption
image_rgb.jpg,Nice image
image_rgb2.jpg,Nice second image
subdir/image_rgb3.jpg,Nice third image
"""
)
)
with open(image_metadata_filename, "w", encoding="utf-8") as f:
f.write(image_metadata)
data_files_with_one_split_and_metadata = DataFilesDict.from_patterns(
get_data_patterns(str(data_dir)), data_dir.as_posix()
)
assert len(data_files_with_one_split_and_metadata) == 1
assert len(data_files_with_one_split_and_metadata["train"]) == 4
return data_files_with_one_split_and_metadata
@pytest.fixture(params=["jsonl", "csv"])
def data_files_with_two_splits_and_metadata(request, tmp_path, image_file):
data_dir = tmp_path / "imagefolder_data_dir_with_metadata_two_splits"
data_dir.mkdir(parents=True, exist_ok=True)
train_dir = data_dir / "train"
train_dir.mkdir(parents=True, exist_ok=True)
test_dir = data_dir / "test"
test_dir.mkdir(parents=True, exist_ok=True)
image_filename = train_dir / "image_rgb.jpg" # train image
shutil.copyfile(image_file, image_filename)
image_filename2 = train_dir / "image_rgb2.jpg" # train image
shutil.copyfile(image_file, image_filename2)
image_filename3 = test_dir / "image_rgb3.jpg" # test image
shutil.copyfile(image_file, image_filename3)
train_image_metadata_filename = train_dir / f"metadata.{request.param}"
image_metadata = (
textwrap.dedent(
"""\
{"file_name": "image_rgb.jpg", "caption": "Nice train image"}
{"file_name": "image_rgb2.jpg", "caption": "Nice second train image"}
"""
)
if request.param == "jsonl"
else textwrap.dedent(
"""\
file_name,caption
image_rgb.jpg,Nice train image
image_rgb2.jpg,Nice second train image
"""
)
)
with open(train_image_metadata_filename, "w", encoding="utf-8") as f:
f.write(image_metadata)
test_image_metadata_filename = test_dir / f"metadata.{request.param}"
image_metadata = (
textwrap.dedent(
"""\
{"file_name": "image_rgb3.jpg", "caption": "Nice test image"}
"""
)
if request.param == "jsonl"
else textwrap.dedent(
"""\
file_name,caption
image_rgb3.jpg,Nice test image
"""
)
)
with open(test_image_metadata_filename, "w", encoding="utf-8") as f:
f.write(image_metadata)
data_files_with_two_splits_and_metadata = DataFilesDict.from_patterns(
get_data_patterns(str(data_dir)), data_dir.as_posix()
)
assert len(data_files_with_two_splits_and_metadata) == 2
assert len(data_files_with_two_splits_and_metadata["train"]) == 3
assert len(data_files_with_two_splits_and_metadata["test"]) == 2
return data_files_with_two_splits_and_metadata
@pytest.fixture
def data_files_with_zip_archives(tmp_path, image_file):
from PIL import Image, ImageOps
data_dir = tmp_path / "imagefolder_data_dir_with_zip_archives"
data_dir.mkdir(parents=True, exist_ok=True)
archive_dir = data_dir / "archive"
archive_dir.mkdir(parents=True, exist_ok=True)
subdir = archive_dir / "subdir"
subdir.mkdir(parents=True, exist_ok=True)
image_filename = archive_dir / "image_rgb.jpg"
shutil.copyfile(image_file, image_filename)
image_filename2 = subdir / "image_rgb2.jpg" # in subdir
# make sure they're two different images
# Indeed we won't be able to compare the image.filename, since the archive is not extracted in streaming mode
ImageOps.flip(Image.open(image_file)).save(image_filename2)
image_metadata_filename = archive_dir / "metadata.jsonl"
image_metadata = textwrap.dedent(
"""\
{"file_name": "image_rgb.jpg", "caption": "Nice image"}
{"file_name": "subdir/image_rgb2.jpg", "caption": "Nice second image"}
"""
)
with open(image_metadata_filename, "w", encoding="utf-8") as f:
f.write(image_metadata)
shutil.make_archive(archive_dir, "zip", archive_dir)
shutil.rmtree(str(archive_dir))
data_files_with_zip_archives = DataFilesDict.from_patterns(get_data_patterns(str(data_dir)), data_dir.as_posix())
assert len(data_files_with_zip_archives) == 1
assert len(data_files_with_zip_archives["train"]) == 1
return data_files_with_zip_archives
@require_pil
# check that labels are inferred correctly from dir names
def test_generate_examples_with_labels(data_files_with_labels_no_metadata, cache_dir):
# there are no metadata.jsonl files in this test case
imagefolder = ImageFolder(data_files=data_files_with_labels_no_metadata, cache_dir=cache_dir, drop_labels=False)
imagefolder.download_and_prepare()
assert imagefolder.info.features == Features({"image": Image(), "label": ClassLabel(names=["cat", "dog"])})
dataset = list(imagefolder.as_dataset()["train"])
label_feature = imagefolder.info.features["label"]
assert dataset[0]["label"] == label_feature._str2int["cat"]
assert dataset[1]["label"] == label_feature._str2int["dog"]
@require_pil
@pytest.mark.parametrize("drop_metadata", [None, True, False])
@pytest.mark.parametrize("drop_labels", [None, True, False])
def test_generate_examples_duplicated_label_key(
image_files_with_labels_and_duplicated_label_key_in_metadata, drop_metadata, drop_labels, cache_dir, caplog
):
cat_image_file, dog_image_file, image_metadata_file = image_files_with_labels_and_duplicated_label_key_in_metadata
imagefolder = ImageFolder(
drop_metadata=drop_metadata,
drop_labels=drop_labels,
data_files=[cat_image_file, dog_image_file, image_metadata_file],
cache_dir=cache_dir,
)
if drop_labels is False:
# infer labels from directories even if metadata files are found
imagefolder.download_and_prepare()
warning_in_logs = any("ignoring metadata columns" in record.msg.lower() for record in caplog.records)
assert warning_in_logs if drop_metadata is not True else not warning_in_logs
dataset = imagefolder.as_dataset()["train"]
assert imagefolder.info.features["label"] == ClassLabel(names=["cat", "dog"])
assert all(example["label"] in imagefolder.info.features["label"]._str2int.values() for example in dataset)
else:
imagefolder.download_and_prepare()
dataset = imagefolder.as_dataset()["train"]
if drop_metadata is not True:
# labels are from metadata
assert imagefolder.info.features["label"] == Value("string")
assert all(example["label"] in ["Cat", "Dog"] for example in dataset)
else:
# drop both labels and metadata
assert imagefolder.info.features == Features({"image": Image()})
assert all(example.keys() == {"image"} for example in dataset)
@require_pil
@pytest.mark.parametrize("drop_metadata", [None, True, False])
@pytest.mark.parametrize("drop_labels", [None, True, False])
def test_generate_examples_drop_labels(data_files_with_labels_no_metadata, drop_metadata, drop_labels):
imagefolder = ImageFolder(
drop_metadata=drop_metadata, drop_labels=drop_labels, data_files=data_files_with_labels_no_metadata
)
gen_kwargs = imagefolder._split_generators(StreamingDownloadManager())[0].gen_kwargs
# removing the labels explicitly requires drop_labels=True
assert gen_kwargs["add_labels"] is not bool(drop_labels)
assert gen_kwargs["add_metadata"] is False
generator = imagefolder._generate_examples(**gen_kwargs)
if not drop_labels:
assert all(
example.keys() == {"image", "label"} and all(val is not None for val in example.values())
for _, example in generator
)
else:
assert all(
example.keys() == {"image"} and all(val is not None for val in example.values())
for _, example in generator
)
@require_pil
@pytest.mark.parametrize("drop_metadata", [None, True, False])
@pytest.mark.parametrize("drop_labels", [None, True, False])
def test_generate_examples_drop_metadata(image_file_with_metadata, drop_metadata, drop_labels):
image_file, image_metadata_file = image_file_with_metadata
imagefolder = ImageFolder(
drop_metadata=drop_metadata, drop_labels=drop_labels, data_files={"train": [image_file, image_metadata_file]}
)
gen_kwargs = imagefolder._split_generators(StreamingDownloadManager())[0].gen_kwargs
# since the dataset has metadata, removing the metadata explicitly requires drop_metadata=True
assert gen_kwargs["add_metadata"] is not bool(drop_metadata)
# since the dataset has metadata, adding the labels explicitly requires drop_labels=False
assert gen_kwargs["add_labels"] is (drop_labels is False)
generator = imagefolder._generate_examples(**gen_kwargs)
expected_columns = {"image"}
if gen_kwargs["add_metadata"]:
expected_columns.add("caption")
if gen_kwargs["add_labels"]:
expected_columns.add("label")
result = [example for _, example in generator]
assert len(result) == 1
example = result[0]
assert example.keys() == expected_columns
for column in expected_columns:
assert example[column] is not None
@require_pil
@pytest.mark.parametrize("drop_metadata", [None, True, False])
def test_generate_examples_with_metadata_in_wrong_location(image_file, image_file_with_metadata, drop_metadata):
_, image_metadata_file = image_file_with_metadata
imagefolder = ImageFolder(drop_metadata=drop_metadata, data_files={"train": [image_file, image_metadata_file]})
gen_kwargs = imagefolder._split_generators(StreamingDownloadManager())[0].gen_kwargs
generator = imagefolder._generate_examples(**gen_kwargs)
if not drop_metadata:
with pytest.raises(ValueError):
list(generator)
else:
assert all(
example.keys() == {"image"} and all(val is not None for val in example.values())
for _, example in generator
)
@require_pil
@pytest.mark.parametrize("drop_metadata", [None, True, False])
def test_generate_examples_with_metadata_that_misses_one_image(
image_files_with_metadata_that_misses_one_image, drop_metadata
):
image_file, image_file2, image_metadata_file = image_files_with_metadata_that_misses_one_image
if not drop_metadata:
features = Features({"image": Image(), "caption": Value("string")})
else:
features = Features({"image": Image()})
imagefolder = ImageFolder(
drop_metadata=drop_metadata,
features=features,
data_files={"train": [image_file, image_file2, image_metadata_file]},
)
gen_kwargs = imagefolder._split_generators(StreamingDownloadManager())[0].gen_kwargs
generator = imagefolder._generate_examples(**gen_kwargs)
if not drop_metadata:
with pytest.raises(ValueError):
list(generator)
else:
assert all(
example.keys() == {"image"} and all(val is not None for val in example.values())
for _, example in generator
)
@require_pil
@pytest.mark.parametrize("streaming", [False, True])
def test_data_files_with_metadata_and_single_split(streaming, cache_dir, data_files_with_one_split_and_metadata):
data_files = data_files_with_one_split_and_metadata
imagefolder = ImageFolder(data_files=data_files, cache_dir=cache_dir)
imagefolder.download_and_prepare()
datasets = imagefolder.as_streaming_dataset() if streaming else imagefolder.as_dataset()
for split, data_files in data_files.items():
expected_num_of_images = len(data_files) - 1 # don't count the metadata file
assert split in datasets
dataset = list(datasets[split])
assert len(dataset) == expected_num_of_images
# make sure each sample has its own image and metadata
assert len({example["image"].filename for example in dataset}) == expected_num_of_images
assert len({example["caption"] for example in dataset}) == expected_num_of_images
assert all(example["caption"] is not None for example in dataset)
@require_pil
@pytest.mark.parametrize("streaming", [False, True])
def test_data_files_with_metadata_and_multiple_splits(streaming, cache_dir, data_files_with_two_splits_and_metadata):
data_files = data_files_with_two_splits_and_metadata
imagefolder = ImageFolder(data_files=data_files, cache_dir=cache_dir)
imagefolder.download_and_prepare()
datasets = imagefolder.as_streaming_dataset() if streaming else imagefolder.as_dataset()
for split, data_files in data_files.items():
expected_num_of_images = len(data_files) - 1 # don't count the metadata file
assert split in datasets
dataset = list(datasets[split])
assert len(dataset) == expected_num_of_images
# make sure each sample has its own image and metadata
assert len({example["image"].filename for example in dataset}) == expected_num_of_images
assert len({example["caption"] for example in dataset}) == expected_num_of_images
assert all(example["caption"] is not None for example in dataset)
@require_pil
@pytest.mark.parametrize("streaming", [False, True])
def test_data_files_with_metadata_and_archives(streaming, cache_dir, data_files_with_zip_archives):
imagefolder = ImageFolder(data_files=data_files_with_zip_archives, cache_dir=cache_dir)
imagefolder.download_and_prepare()
datasets = imagefolder.as_streaming_dataset() if streaming else imagefolder.as_dataset()
for split, data_files in data_files_with_zip_archives.items():
num_of_archives = len(data_files) # the metadata file is inside the archive
expected_num_of_images = 2 * num_of_archives
assert split in datasets
dataset = list(datasets[split])
assert len(dataset) == expected_num_of_images
# make sure each sample has its own image and metadata
assert len({np.array(example["image"])[0, 0, 0] for example in dataset}) == expected_num_of_images
assert len({example["caption"] for example in dataset}) == expected_num_of_images
assert all(example["caption"] is not None for example in dataset)
@require_pil
def test_data_files_with_wrong_metadata_file_name(cache_dir, tmp_path, image_file):
data_dir = tmp_path / "data_dir_with_bad_metadata"
data_dir.mkdir(parents=True, exist_ok=True)
shutil.copyfile(image_file, data_dir / "image_rgb.jpg")
image_metadata_filename = data_dir / "bad_metadata.jsonl" # bad file
image_metadata = textwrap.dedent(
"""\
{"file_name": "image_rgb.jpg", "caption": "Nice image"}
"""
)
with open(image_metadata_filename, "w", encoding="utf-8") as f:
f.write(image_metadata)
data_files_with_bad_metadata = DataFilesDict.from_patterns(get_data_patterns(str(data_dir)), data_dir.as_posix())
imagefolder = ImageFolder(data_files=data_files_with_bad_metadata, cache_dir=cache_dir)
imagefolder.download_and_prepare()
dataset = imagefolder.as_dataset(split="train")
# check that there are no metadata, since the metadata file name doesn't have the right name
assert "caption" not in dataset.column_names
@require_pil
def test_data_files_with_wrong_image_file_name_column_in_metadata_file(cache_dir, tmp_path, image_file):
data_dir = tmp_path / "data_dir_with_bad_metadata"
data_dir.mkdir(parents=True, exist_ok=True)
shutil.copyfile(image_file, data_dir / "image_rgb.jpg")
image_metadata_filename = data_dir / "metadata.jsonl"
image_metadata = textwrap.dedent( # with bad column "bad_file_name" instead of "file_name"
"""\
{"bad_file_name": "image_rgb.jpg", "caption": "Nice image"}
"""
)
with open(image_metadata_filename, "w", encoding="utf-8") as f:
f.write(image_metadata)
data_files_with_bad_metadata = DataFilesDict.from_patterns(get_data_patterns(str(data_dir)), data_dir.as_posix())
imagefolder = ImageFolder(data_files=data_files_with_bad_metadata, cache_dir=cache_dir)
with pytest.raises(ValueError) as exc_info:
imagefolder.download_and_prepare()
assert "`file_name` must be present" in str(exc_info.value)
@require_pil
def test_data_files_with_with_metadata_in_different_formats(cache_dir, tmp_path, image_file):
data_dir = tmp_path / "data_dir_with_metadata_in_different_format"
data_dir.mkdir(parents=True, exist_ok=True)
shutil.copyfile(image_file, data_dir / "image_rgb.jpg")
image_metadata_filename_jsonl = data_dir / "metadata.jsonl"
image_metadata_jsonl = textwrap.dedent(
"""\
{"file_name": "image_rgb.jpg", "caption": "Nice image"}
"""
)
with open(image_metadata_filename_jsonl, "w", encoding="utf-8") as f:
f.write(image_metadata_jsonl)
image_metadata_filename_csv = data_dir / "metadata.csv"
image_metadata_csv = textwrap.dedent(
"""\
file_name,caption
image_rgb.jpg,Nice image
"""
)
with open(image_metadata_filename_csv, "w", encoding="utf-8") as f:
f.write(image_metadata_csv)
data_files_with_bad_metadata = DataFilesDict.from_patterns(get_data_patterns(str(data_dir)), data_dir.as_posix())
imagefolder = ImageFolder(data_files=data_files_with_bad_metadata, cache_dir=cache_dir)
with pytest.raises(ValueError) as exc_info:
imagefolder.download_and_prepare()
assert "metadata files with different extensions" in str(exc_info.value)
| 0
|
hf_public_repos/datasets/tests
|
hf_public_repos/datasets/tests/packaged_modules/test_folder_based_builder.py
|
import importlib
import shutil
import textwrap
import pytest
from datasets import ClassLabel, DownloadManager, Features, Value
from datasets.data_files import DataFilesDict, get_data_patterns
from datasets.download.streaming_download_manager import StreamingDownloadManager
from datasets.packaged_modules.folder_based_builder.folder_based_builder import (
FolderBasedBuilder,
FolderBasedBuilderConfig,
)
from datasets.tasks import TextClassification
remote_files = [
"https://huggingface.co/datasets/hf-internal-testing/textfolder/resolve/main/hallo.txt",
"https://huggingface.co/datasets/hf-internal-testing/textfolder/resolve/main/hello.txt",
"https://huggingface.co/datasets/hf-internal-testing/textfolder/resolve/main/class1/bonjour.txt",
"https://huggingface.co/datasets/hf-internal-testing/textfolder/resolve/main/class1/bonjour2.txt",
]
class DummyFolderBasedBuilder(FolderBasedBuilder):
BASE_FEATURE = dict
BASE_COLUMN_NAME = "base"
BUILDER_CONFIG_CLASS = FolderBasedBuilderConfig
EXTENSIONS = [".txt"]
CLASSIFICATION_TASK = TextClassification(text_column="base", label_column="label")
@pytest.fixture
def cache_dir(tmp_path):
return str(tmp_path / "autofolder_cache_dir")
@pytest.fixture
def auto_text_file(text_file):
return str(text_file)
@pytest.fixture
def data_files_with_labels_no_metadata(tmp_path, auto_text_file):
data_dir = tmp_path / "data_files_with_labels_no_metadata"
data_dir.mkdir(parents=True, exist_ok=True)
subdir_class_0 = data_dir / "class0"
subdir_class_0.mkdir(parents=True, exist_ok=True)
subdir_class_1 = data_dir / "class1"
subdir_class_1.mkdir(parents=True, exist_ok=True)
filename = subdir_class_0 / "file0.txt"
shutil.copyfile(auto_text_file, filename)
filename2 = subdir_class_1 / "file1.txt"
shutil.copyfile(auto_text_file, filename2)
data_files_with_labels_no_metadata = DataFilesDict.from_patterns(
get_data_patterns(str(data_dir)), data_dir.as_posix()
)
return data_files_with_labels_no_metadata
@pytest.fixture
def data_files_with_different_levels_no_metadata(tmp_path, auto_text_file):
data_dir = tmp_path / "data_files_with_different_levels"
data_dir.mkdir(parents=True, exist_ok=True)
subdir_class_0 = data_dir / "class0"
subdir_class_0.mkdir(parents=True, exist_ok=True)
subdir_class_1 = data_dir / "subdir" / "class1"
subdir_class_1.mkdir(parents=True, exist_ok=True)
filename = subdir_class_0 / "file0.txt"
shutil.copyfile(auto_text_file, filename)
filename2 = subdir_class_1 / "file1.txt"
shutil.copyfile(auto_text_file, filename2)
data_files_with_different_levels = DataFilesDict.from_patterns(
get_data_patterns(str(data_dir)), data_dir.as_posix()
)
return data_files_with_different_levels
@pytest.fixture
def data_files_with_one_label_no_metadata(tmp_path, auto_text_file):
# only one label found = all files in a single dir/in a root dir
data_dir = tmp_path / "data_files_with_one_label"
data_dir.mkdir(parents=True, exist_ok=True)
filename = data_dir / "file0.txt"
shutil.copyfile(auto_text_file, filename)
filename2 = data_dir / "file1.txt"
shutil.copyfile(auto_text_file, filename2)
data_files_with_one_label = DataFilesDict.from_patterns(get_data_patterns(str(data_dir)), data_dir.as_posix())
return data_files_with_one_label
@pytest.fixture
def files_with_labels_and_duplicated_label_key_in_metadata(tmp_path, auto_text_file):
data_dir = tmp_path / "files_with_labels_and_label_key_in_metadata"
data_dir.mkdir(parents=True, exist_ok=True)
subdir_class_0 = data_dir / "class0"
subdir_class_0.mkdir(parents=True, exist_ok=True)
subdir_class_1 = data_dir / "class1"
subdir_class_1.mkdir(parents=True, exist_ok=True)
filename = subdir_class_0 / "file_class0.txt"
shutil.copyfile(auto_text_file, filename)
filename2 = subdir_class_1 / "file_class1.txt"
shutil.copyfile(auto_text_file, filename2)
metadata_filename = tmp_path / data_dir / "metadata.jsonl"
metadata = textwrap.dedent(
"""\
{"file_name": "class0/file_class0.txt", "additional_feature": "First dummy file", "label": "CLASS_0"}
{"file_name": "class1/file_class1.txt", "additional_feature": "Second dummy file", "label": "CLASS_1"}
"""
)
with open(metadata_filename, "w", encoding="utf-8") as f:
f.write(metadata)
return str(filename), str(filename2), str(metadata_filename)
@pytest.fixture
def file_with_metadata(tmp_path, text_file):
filename = tmp_path / "file.txt"
shutil.copyfile(text_file, filename)
metadata_filename = tmp_path / "metadata.jsonl"
metadata = textwrap.dedent(
"""\
{"file_name": "file.txt", "additional_feature": "Dummy file"}
"""
)
with open(metadata_filename, "w", encoding="utf-8") as f:
f.write(metadata)
return str(filename), str(metadata_filename)
@pytest.fixture()
def files_with_metadata_that_misses_one_sample(tmp_path, auto_text_file):
filename = tmp_path / "file.txt"
shutil.copyfile(auto_text_file, filename)
filename2 = tmp_path / "file2.txt"
shutil.copyfile(auto_text_file, filename2)
metadata_filename = tmp_path / "metadata.jsonl"
metadata = textwrap.dedent(
"""\
{"file_name": "file.txt", "additional_feature": "Dummy file"}
"""
)
with open(metadata_filename, "w", encoding="utf-8") as f:
f.write(metadata)
return str(filename), str(filename2), str(metadata_filename)
@pytest.fixture
def data_files_with_one_split_and_metadata(tmp_path, auto_text_file):
data_dir = tmp_path / "autofolder_data_dir_with_metadata_one_split"
data_dir.mkdir(parents=True, exist_ok=True)
subdir = data_dir / "subdir"
subdir.mkdir(parents=True, exist_ok=True)
filename = data_dir / "file.txt"
shutil.copyfile(auto_text_file, filename)
filename2 = data_dir / "file2.txt"
shutil.copyfile(auto_text_file, filename2)
filename3 = subdir / "file3.txt" # in subdir
shutil.copyfile(auto_text_file, filename3)
metadata_filename = data_dir / "metadata.jsonl"
metadata = textwrap.dedent(
"""\
{"file_name": "file.txt", "additional_feature": "Dummy file"}
{"file_name": "file2.txt", "additional_feature": "Second dummy file"}
{"file_name": "./subdir/file3.txt", "additional_feature": "Third dummy file"}
"""
)
with open(metadata_filename, "w", encoding="utf-8") as f:
f.write(metadata)
data_files_with_one_split_and_metadata = DataFilesDict.from_patterns(
get_data_patterns(str(data_dir)), data_dir.as_posix()
)
assert len(data_files_with_one_split_and_metadata) == 1
assert len(data_files_with_one_split_and_metadata["train"]) == 4
return data_files_with_one_split_and_metadata
@pytest.fixture
def data_files_with_two_splits_and_metadata(tmp_path, auto_text_file):
data_dir = tmp_path / "autofolder_data_dir_with_metadata_two_splits"
data_dir.mkdir(parents=True, exist_ok=True)
train_dir = data_dir / "train"
train_dir.mkdir(parents=True, exist_ok=True)
test_dir = data_dir / "test"
test_dir.mkdir(parents=True, exist_ok=True)
filename = train_dir / "file.txt" # train
shutil.copyfile(auto_text_file, filename)
filename2 = train_dir / "file2.txt" # train
shutil.copyfile(auto_text_file, filename2)
filename3 = test_dir / "file3.txt" # test
shutil.copyfile(auto_text_file, filename3)
train_metadata_filename = train_dir / "metadata.jsonl"
train_metadata = textwrap.dedent(
"""\
{"file_name": "file.txt", "additional_feature": "Train dummy file"}
{"file_name": "file2.txt", "additional_feature": "Second train dummy file"}
"""
)
with open(train_metadata_filename, "w", encoding="utf-8") as f:
f.write(train_metadata)
test_metadata_filename = test_dir / "metadata.jsonl"
test_metadata = textwrap.dedent(
"""\
{"file_name": "file3.txt", "additional_feature": "Test dummy file"}
"""
)
with open(test_metadata_filename, "w", encoding="utf-8") as f:
f.write(test_metadata)
data_files_with_two_splits_and_metadata = DataFilesDict.from_patterns(
get_data_patterns(str(data_dir)), data_dir.as_posix()
)
assert len(data_files_with_two_splits_and_metadata) == 2
assert len(data_files_with_two_splits_and_metadata["train"]) == 3
assert len(data_files_with_two_splits_and_metadata["test"]) == 2
return data_files_with_two_splits_and_metadata
@pytest.fixture
def data_files_with_zip_archives(tmp_path, auto_text_file):
data_dir = tmp_path / "autofolder_data_dir_with_zip_archives"
data_dir.mkdir(parents=True, exist_ok=True)
archive_dir = data_dir / "archive"
archive_dir.mkdir(parents=True, exist_ok=True)
subdir = archive_dir / "subdir"
subdir.mkdir(parents=True, exist_ok=True)
filename = archive_dir / "file.txt"
shutil.copyfile(auto_text_file, filename)
filename2 = subdir / "file2.txt" # in subdir
shutil.copyfile(auto_text_file, filename2)
metadata_filename = archive_dir / "metadata.jsonl"
metadata = textwrap.dedent(
"""\
{"file_name": "file.txt", "additional_feature": "Dummy file"}
{"file_name": "subdir/file2.txt", "additional_feature": "Second dummy file"}
"""
)
with open(metadata_filename, "w", encoding="utf-8") as f:
f.write(metadata)
shutil.make_archive(archive_dir, "zip", archive_dir)
shutil.rmtree(str(archive_dir))
data_files_with_zip_archives = DataFilesDict.from_patterns(get_data_patterns(str(data_dir)), data_dir.as_posix())
assert len(data_files_with_zip_archives) == 1
assert len(data_files_with_zip_archives["train"]) == 1
return data_files_with_zip_archives
def test_inferring_labels_from_data_dirs(data_files_with_labels_no_metadata, cache_dir):
autofolder = DummyFolderBasedBuilder(
data_files=data_files_with_labels_no_metadata, cache_dir=cache_dir, drop_labels=False
)
gen_kwargs = autofolder._split_generators(StreamingDownloadManager())[0].gen_kwargs
assert autofolder.info.features == Features({"base": {}, "label": ClassLabel(names=["class0", "class1"])})
generator = autofolder._generate_examples(**gen_kwargs)
assert all(example["label"] in {"class0", "class1"} for _, example in generator)
def test_default_folder_builder_not_usable(data_files_with_labels_no_metadata, cache_dir):
# builder would try to access non-existing attributes of a default `BuilderConfig` class
# as a custom one is not provided
with pytest.raises(AttributeError):
_ = FolderBasedBuilder(
data_files=data_files_with_labels_no_metadata,
cache_dir=cache_dir,
)
# test that AutoFolder is extended for streaming when it's child class is instantiated:
# see line 115 in src/datasets/streaming.py
def test_streaming_patched():
_ = DummyFolderBasedBuilder()
module = importlib.import_module(FolderBasedBuilder.__module__)
assert hasattr(module, "_patched_for_streaming")
assert module._patched_for_streaming
@pytest.mark.parametrize("drop_metadata", [None, True, False])
@pytest.mark.parametrize("drop_labels", [None, True, False])
def test_generate_examples_duplicated_label_key(
files_with_labels_and_duplicated_label_key_in_metadata, drop_metadata, drop_labels, cache_dir, caplog
):
class0_file, class1_file, metadata_file = files_with_labels_and_duplicated_label_key_in_metadata
autofolder = DummyFolderBasedBuilder(
data_files=[class0_file, class1_file, metadata_file],
cache_dir=cache_dir,
drop_metadata=drop_metadata,
drop_labels=drop_labels,
)
gen_kwargs = autofolder._split_generators(StreamingDownloadManager())[0].gen_kwargs
generator = autofolder._generate_examples(**gen_kwargs)
if drop_labels is False:
# infer labels from directories even if metadata files are found
warning_in_logs = any("ignoring metadata columns" in record.msg.lower() for record in caplog.records)
assert warning_in_logs if drop_metadata is not True else not warning_in_logs
assert autofolder.info.features["label"] == ClassLabel(names=["class0", "class1"])
assert all(example["label"] in ["class0", "class1"] for _, example in generator)
else:
if drop_metadata is not True:
# labels are from metadata
assert autofolder.info.features["label"] == Value("string")
assert all(example["label"] in ["CLASS_0", "CLASS_1"] for _, example in generator)
else:
# drop both labels and metadata
assert autofolder.info.features == Features({"base": {}})
assert all(example.keys() == {"base"} for _, example in generator)
@pytest.mark.parametrize("drop_metadata", [None, True, False])
@pytest.mark.parametrize("drop_labels", [None, True, False])
def test_generate_examples_drop_labels(
data_files_with_labels_no_metadata, auto_text_file, drop_metadata, drop_labels, cache_dir
):
autofolder = DummyFolderBasedBuilder(
data_files=data_files_with_labels_no_metadata,
drop_metadata=drop_metadata,
drop_labels=drop_labels,
cache_dir=cache_dir,
)
gen_kwargs = autofolder._split_generators(StreamingDownloadManager())[0].gen_kwargs
# removing labels explicitly requires drop_labels=True
assert gen_kwargs["add_labels"] is not bool(drop_labels)
assert gen_kwargs["add_metadata"] is False
generator = autofolder._generate_examples(**gen_kwargs)
if not drop_labels:
assert all(
example.keys() == {"base", "label"} and all(val is not None for val in example.values())
for _, example in generator
)
else:
assert all(
example.keys() == {"base"} and all(val is not None for val in example.values()) for _, example in generator
)
@pytest.mark.parametrize("drop_metadata", [None, True, False])
@pytest.mark.parametrize("drop_labels", [None, True, False])
def test_generate_examples_drop_metadata(file_with_metadata, drop_metadata, drop_labels, cache_dir):
file, metadata_file = file_with_metadata
autofolder = DummyFolderBasedBuilder(
data_files=[file, metadata_file],
drop_metadata=drop_metadata,
drop_labels=drop_labels,
cache_dir=cache_dir,
)
gen_kwargs = autofolder._split_generators(StreamingDownloadManager())[0].gen_kwargs
# since the dataset has metadata, removing the metadata explicitly requires drop_metadata=True
assert gen_kwargs["add_metadata"] is not bool(drop_metadata)
# since the dataset has metadata, adding the labels explicitly requires drop_labels=False
assert gen_kwargs["add_labels"] is (drop_labels is False)
generator = autofolder._generate_examples(**gen_kwargs)
expected_columns = {"base"}
if gen_kwargs["add_metadata"]:
expected_columns.add("additional_feature")
if gen_kwargs["add_labels"]:
expected_columns.add("label")
result = [example for _, example in generator]
assert len(result) == 1
example = result[0]
assert example.keys() == expected_columns
for column in expected_columns:
assert example[column] is not None
@pytest.mark.parametrize("remote", [True, False])
@pytest.mark.parametrize("drop_labels", [None, True, False])
def test_data_files_with_different_levels_no_metadata(
data_files_with_different_levels_no_metadata, drop_labels, remote, cache_dir
):
data_files = remote_files if remote else data_files_with_different_levels_no_metadata
autofolder = DummyFolderBasedBuilder(
data_files=data_files,
cache_dir=cache_dir,
drop_labels=drop_labels,
)
gen_kwargs = autofolder._split_generators(StreamingDownloadManager())[0].gen_kwargs
generator = autofolder._generate_examples(**gen_kwargs)
if drop_labels is not False:
# with None (default) we should drop labels if files are on different levels in dir structure
assert "label" not in autofolder.info.features
assert all(example.keys() == {"base"} for _, example in generator)
else:
assert "label" in autofolder.info.features
assert isinstance(autofolder.info.features["label"], ClassLabel)
assert all(example.keys() == {"base", "label"} for _, example in generator)
@pytest.mark.parametrize("remote", [False, True])
@pytest.mark.parametrize("drop_labels", [None, True, False])
def test_data_files_with_one_label_no_metadata(data_files_with_one_label_no_metadata, drop_labels, remote, cache_dir):
data_files = remote_files[:2] if remote else data_files_with_one_label_no_metadata
autofolder = DummyFolderBasedBuilder(
data_files=data_files,
cache_dir=cache_dir,
drop_labels=drop_labels,
)
gen_kwargs = autofolder._split_generators(StreamingDownloadManager())[0].gen_kwargs
generator = autofolder._generate_examples(**gen_kwargs)
if drop_labels is not False:
# with None (default) we should drop labels if only one label is found (=if there is a single dir)
assert "label" not in autofolder.info.features
assert all(example.keys() == {"base"} for _, example in generator)
else:
assert "label" in autofolder.info.features
assert isinstance(autofolder.info.features["label"], ClassLabel)
assert all(example.keys() == {"base", "label"} for _, example in generator)
@pytest.mark.parametrize("drop_metadata", [None, True, False])
def test_data_files_with_metadata_that_misses_one_sample(
files_with_metadata_that_misses_one_sample, drop_metadata, cache_dir
):
file, file2, metadata_file = files_with_metadata_that_misses_one_sample
if not drop_metadata:
features = Features({"base": None, "additional_feature": Value("string")})
else:
features = Features({"base": None})
autofolder = DummyFolderBasedBuilder(
data_files=[file, file2, metadata_file],
drop_metadata=drop_metadata,
features=features,
cache_dir=cache_dir,
)
gen_kwargs = autofolder._split_generators(StreamingDownloadManager())[0].gen_kwargs
generator = autofolder._generate_examples(**gen_kwargs)
if not drop_metadata:
with pytest.raises(ValueError):
list(generator)
else:
assert all(
example.keys() == {"base"} and all(val is not None for val in example.values()) for _, example in generator
)
@pytest.mark.parametrize("streaming", [False, True])
@pytest.mark.parametrize("n_splits", [1, 2])
def test_data_files_with_metadata_and_splits(
streaming, cache_dir, n_splits, data_files_with_one_split_and_metadata, data_files_with_two_splits_and_metadata
):
data_files = data_files_with_one_split_and_metadata if n_splits == 1 else data_files_with_two_splits_and_metadata
autofolder = DummyFolderBasedBuilder(
data_files=data_files,
cache_dir=cache_dir,
)
download_manager = StreamingDownloadManager() if streaming else DownloadManager()
generated_splits = autofolder._split_generators(download_manager)
for (split, files), generated_split in zip(data_files.items(), generated_splits):
assert split == generated_split.name
expected_num_of_examples = len(files) - 1
generated_examples = list(autofolder._generate_examples(**generated_split.gen_kwargs))
assert len(generated_examples) == expected_num_of_examples
assert len({example["base"] for _, example in generated_examples}) == expected_num_of_examples
assert len({example["additional_feature"] for _, example in generated_examples}) == expected_num_of_examples
assert all(example["additional_feature"] is not None for _, example in generated_examples)
@pytest.mark.parametrize("streaming", [False, True])
def test_data_files_with_metadata_and_archives(streaming, cache_dir, data_files_with_zip_archives):
autofolder = DummyFolderBasedBuilder(data_files=data_files_with_zip_archives, cache_dir=cache_dir)
download_manager = StreamingDownloadManager() if streaming else DownloadManager()
generated_splits = autofolder._split_generators(download_manager)
for (split, files), generated_split in zip(data_files_with_zip_archives.items(), generated_splits):
assert split == generated_split.name
num_of_archives = len(files)
expected_num_of_examples = 2 * num_of_archives
generated_examples = list(autofolder._generate_examples(**generated_split.gen_kwargs))
assert len(generated_examples) == expected_num_of_examples
assert len({example["base"] for _, example in generated_examples}) == expected_num_of_examples
assert len({example["additional_feature"] for _, example in generated_examples}) == expected_num_of_examples
assert all(example["additional_feature"] is not None for _, example in generated_examples)
def test_data_files_with_wrong_metadata_file_name(cache_dir, tmp_path, auto_text_file):
data_dir = tmp_path / "data_dir_with_bad_metadata"
data_dir.mkdir(parents=True, exist_ok=True)
shutil.copyfile(auto_text_file, data_dir / "file.txt")
metadata_filename = data_dir / "bad_metadata.jsonl" # bad file
metadata = textwrap.dedent(
"""\
{"file_name": "file.txt", "additional_feature": "Dummy file"}
"""
)
with open(metadata_filename, "w", encoding="utf-8") as f:
f.write(metadata)
data_files_with_bad_metadata = DataFilesDict.from_patterns(get_data_patterns(str(data_dir)), data_dir.as_posix())
autofolder = DummyFolderBasedBuilder(data_files=data_files_with_bad_metadata, cache_dir=cache_dir)
gen_kwargs = autofolder._split_generators(StreamingDownloadManager())[0].gen_kwargs
generator = autofolder._generate_examples(**gen_kwargs)
assert all("additional_feature" not in example for _, example in generator)
def test_data_files_with_wrong_file_name_column_in_metadata_file(cache_dir, tmp_path, auto_text_file):
data_dir = tmp_path / "data_dir_with_bad_metadata"
data_dir.mkdir(parents=True, exist_ok=True)
shutil.copyfile(auto_text_file, data_dir / "file.txt")
metadata_filename = data_dir / "metadata.jsonl"
metadata = textwrap.dedent( # with bad column "bad_file_name" instead of "file_name"
"""\
{"bad_file_name": "file.txt", "additional_feature": "Dummy file"}
"""
)
with open(metadata_filename, "w", encoding="utf-8") as f:
f.write(metadata)
data_files_with_bad_metadata = DataFilesDict.from_patterns(get_data_patterns(str(data_dir)), data_dir.as_posix())
autofolder = DummyFolderBasedBuilder(data_files=data_files_with_bad_metadata, cache_dir=cache_dir)
with pytest.raises(ValueError) as exc_info:
_ = autofolder._split_generators(StreamingDownloadManager())[0].gen_kwargs
assert "`file_name` must be present" in str(exc_info.value)
| 0
|
hf_public_repos/datasets/tests
|
hf_public_repos/datasets/tests/packaged_modules/test_spark.py
|
from unittest.mock import patch
import pyspark
from datasets.packaged_modules.spark.spark import (
Spark,
SparkExamplesIterable,
_generate_iterable_examples,
)
from ..utils import (
require_dill_gt_0_3_2,
require_not_windows,
)
def _get_expected_row_ids_and_row_dicts_for_partition_order(df, partition_order):
expected_row_ids_and_row_dicts = []
for part_id in partition_order:
partition = df.where(f"SPARK_PARTITION_ID() = {part_id}").collect()
for row_idx, row in enumerate(partition):
expected_row_ids_and_row_dicts.append((f"{part_id}_{row_idx}", row.asDict()))
return expected_row_ids_and_row_dicts
@require_not_windows
@require_dill_gt_0_3_2
def test_repartition_df_if_needed():
spark = pyspark.sql.SparkSession.builder.master("local[*]").appName("pyspark").getOrCreate()
df = spark.range(100).repartition(1)
spark_builder = Spark(df)
# The id ints will be converted to Pyarrow int64s, so each row will be 8 bytes. Setting a max_shard_size of 16 means
# that each partition can hold 2 rows.
spark_builder._repartition_df_if_needed(max_shard_size=16)
# Given that the dataframe has 100 rows and each partition has 2 rows, we expect 50 partitions.
assert spark_builder.df.rdd.getNumPartitions() == 50
@require_not_windows
@require_dill_gt_0_3_2
def test_generate_iterable_examples():
spark = pyspark.sql.SparkSession.builder.master("local[*]").appName("pyspark").getOrCreate()
df = spark.range(10).repartition(2)
partition_order = [1, 0]
generate_fn = _generate_iterable_examples(df, partition_order) # Reverse the partitions.
expected_row_ids_and_row_dicts = _get_expected_row_ids_and_row_dicts_for_partition_order(df, partition_order)
for i, (row_id, row_dict) in enumerate(generate_fn()):
expected_row_id, expected_row_dict = expected_row_ids_and_row_dicts[i]
assert row_id == expected_row_id
assert row_dict == expected_row_dict
@require_not_windows
@require_dill_gt_0_3_2
def test_spark_examples_iterable():
spark = pyspark.sql.SparkSession.builder.master("local[*]").appName("pyspark").getOrCreate()
df = spark.range(10).repartition(1)
it = SparkExamplesIterable(df)
assert it.n_shards == 1
for i, (row_id, row_dict) in enumerate(it):
assert row_id == f"0_{i}"
assert row_dict == {"id": i}
@require_not_windows
@require_dill_gt_0_3_2
def test_spark_examples_iterable_shuffle():
spark = pyspark.sql.SparkSession.builder.master("local[*]").appName("pyspark").getOrCreate()
df = spark.range(30).repartition(3)
# Mock the generator so that shuffle reverses the partition indices.
with patch("numpy.random.Generator") as generator_mock:
generator_mock.shuffle.side_effect = lambda x: x.reverse()
expected_row_ids_and_row_dicts = _get_expected_row_ids_and_row_dicts_for_partition_order(df, [2, 1, 0])
shuffled_it = SparkExamplesIterable(df).shuffle_data_sources(generator_mock)
assert shuffled_it.n_shards == 3
for i, (row_id, row_dict) in enumerate(shuffled_it):
expected_row_id, expected_row_dict = expected_row_ids_and_row_dicts[i]
assert row_id == expected_row_id
assert row_dict == expected_row_dict
@require_not_windows
@require_dill_gt_0_3_2
def test_spark_examples_iterable_shard():
spark = pyspark.sql.SparkSession.builder.master("local[*]").appName("pyspark").getOrCreate()
df = spark.range(20).repartition(4)
# Partitions 0 and 2
shard_it_1 = SparkExamplesIterable(df).shard_data_sources(worker_id=0, num_workers=2)
assert shard_it_1.n_shards == 2
expected_row_ids_and_row_dicts_1 = _get_expected_row_ids_and_row_dicts_for_partition_order(df, [0, 2])
for i, (row_id, row_dict) in enumerate(shard_it_1):
expected_row_id, expected_row_dict = expected_row_ids_and_row_dicts_1[i]
assert row_id == expected_row_id
assert row_dict == expected_row_dict
# Partitions 1 and 3
shard_it_2 = SparkExamplesIterable(df).shard_data_sources(worker_id=1, num_workers=2)
assert shard_it_2.n_shards == 2
expected_row_ids_and_row_dicts_2 = _get_expected_row_ids_and_row_dicts_for_partition_order(df, [1, 3])
for i, (row_id, row_dict) in enumerate(shard_it_2):
expected_row_id, expected_row_dict = expected_row_ids_and_row_dicts_2[i]
assert row_id == expected_row_id
assert row_dict == expected_row_dict
@require_not_windows
@require_dill_gt_0_3_2
def test_repartition_df_if_needed_max_num_df_rows():
spark = pyspark.sql.SparkSession.builder.master("local[*]").appName("pyspark").getOrCreate()
df = spark.range(100).repartition(1)
spark_builder = Spark(df)
# Choose a small max_shard_size for maximum partitioning.
spark_builder._repartition_df_if_needed(max_shard_size=1)
# The new number of partitions should not be greater than the number of rows.
assert spark_builder.df.rdd.getNumPartitions() == 100
| 0
|
hf_public_repos/datasets/tests
|
hf_public_repos/datasets/tests/packaged_modules/test_csv.py
|
import os
import textwrap
import pyarrow as pa
import pytest
from datasets import ClassLabel, Features, Image
from datasets.packaged_modules.csv.csv import Csv
from ..utils import require_pil
@pytest.fixture
def csv_file(tmp_path):
filename = tmp_path / "file.csv"
data = textwrap.dedent(
"""\
header1,header2
1,2
10,20
"""
)
with open(filename, "w") as f:
f.write(data)
return str(filename)
@pytest.fixture
def malformed_csv_file(tmp_path):
filename = tmp_path / "malformed_file.csv"
data = textwrap.dedent(
"""\
header1,header2
1,2
10,20,
"""
)
with open(filename, "w") as f:
f.write(data)
return str(filename)
@pytest.fixture
def csv_file_with_image(tmp_path, image_file):
filename = tmp_path / "csv_with_image.csv"
data = textwrap.dedent(
f"""\
image
{image_file}
"""
)
with open(filename, "w") as f:
f.write(data)
return str(filename)
@pytest.fixture
def csv_file_with_label(tmp_path):
filename = tmp_path / "csv_with_label.csv"
data = textwrap.dedent(
"""\
label
good
bad
good
"""
)
with open(filename, "w") as f:
f.write(data)
return str(filename)
@pytest.fixture
def csv_file_with_int_list(tmp_path):
filename = tmp_path / "csv_with_int_list.csv"
data = textwrap.dedent(
"""\
int_list
1 2 3
4 5 6
7 8 9
"""
)
with open(filename, "w") as f:
f.write(data)
return str(filename)
def test_csv_generate_tables_raises_error_with_malformed_csv(csv_file, malformed_csv_file, caplog):
csv = Csv()
generator = csv._generate_tables([[csv_file, malformed_csv_file]])
with pytest.raises(ValueError, match="Error tokenizing data"):
for _ in generator:
pass
assert any(
record.levelname == "ERROR"
and "Failed to read file" in record.message
and os.path.basename(malformed_csv_file) in record.message
for record in caplog.records
)
@require_pil
def test_csv_cast_image(csv_file_with_image):
with open(csv_file_with_image, encoding="utf-8") as f:
image_file = f.read().splitlines()[1]
csv = Csv(encoding="utf-8", features=Features({"image": Image()}))
generator = csv._generate_tables([[csv_file_with_image]])
pa_table = pa.concat_tables([table for _, table in generator])
assert pa_table.schema.field("image").type == Image()()
generated_content = pa_table.to_pydict()["image"]
assert generated_content == [{"path": image_file, "bytes": None}]
def test_csv_cast_label(csv_file_with_label):
with open(csv_file_with_label, encoding="utf-8") as f:
labels = f.read().splitlines()[1:]
csv = Csv(encoding="utf-8", features=Features({"label": ClassLabel(names=["good", "bad"])}))
generator = csv._generate_tables([[csv_file_with_label]])
pa_table = pa.concat_tables([table for _, table in generator])
assert pa_table.schema.field("label").type == ClassLabel(names=["good", "bad"])()
generated_content = pa_table.to_pydict()["label"]
assert generated_content == [ClassLabel(names=["good", "bad"]).str2int(label) for label in labels]
def test_csv_convert_int_list(csv_file_with_int_list):
csv = Csv(encoding="utf-8", sep=",", converters={"int_list": lambda x: [int(i) for i in x.split()]})
generator = csv._generate_tables([[csv_file_with_int_list]])
pa_table = pa.concat_tables([table for _, table in generator])
assert pa.types.is_list(pa_table.schema.field("int_list").type)
generated_content = pa_table.to_pydict()["int_list"]
assert generated_content == [[1, 2, 3], [4, 5, 6], [7, 8, 9]]
| 0
|
hf_public_repos/datasets/tests
|
hf_public_repos/datasets/tests/packaged_modules/test_text.py
|
import textwrap
import pyarrow as pa
import pytest
from datasets import Features, Image
from datasets.packaged_modules.text.text import Text
from ..utils import require_pil
@pytest.fixture
def text_file(tmp_path):
filename = tmp_path / "text.txt"
data = textwrap.dedent(
"""\
Lorem ipsum dolor sit amet, consectetur adipiscing elit, sed do eiusmod tempor incididunt ut labore et dolore magna aliqua.
Ut enim ad minim veniam, quis nostrud exercitation ullamco laboris nisi ut aliquip ex ea commodo consequat.
Duis aute irure dolor in reprehenderit in voluptate velit esse cillum dolore eu fugiat nulla pariatur.
Excepteur sint occaecat cupidatat non proident, sunt in culpa qui officia deserunt mollit anim id est laborum.
Second paragraph:
Lorem ipsum dolor sit amet, consectetur adipiscing elit, sed do eiusmod tempor incididunt ut labore et dolore magna aliqua.
Ut enim ad minim veniam, quis nostrud exercitation ullamco laboris nisi ut aliquip ex ea commodo consequat.
Duis aute irure dolor in reprehenderit in voluptate velit esse cillum dolore eu fugiat nulla pariatur.
Excepteur sint occaecat cupidatat non proident, sunt in culpa qui officia deserunt mollit anim id est laborum.
"""
)
with open(filename, "w", encoding="utf-8") as f:
f.write(data)
return str(filename)
@pytest.fixture
def text_file_with_image(tmp_path, image_file):
filename = tmp_path / "text_with_image.txt"
with open(filename, "w", encoding="utf-8") as f:
f.write(image_file)
return str(filename)
@pytest.mark.parametrize("keep_linebreaks", [True, False])
def test_text_linebreaks(text_file, keep_linebreaks):
with open(text_file, encoding="utf-8") as f:
expected_content = f.read().splitlines(keepends=keep_linebreaks)
text = Text(keep_linebreaks=keep_linebreaks, encoding="utf-8")
generator = text._generate_tables([[text_file]])
generated_content = pa.concat_tables([table for _, table in generator]).to_pydict()["text"]
assert generated_content == expected_content
@require_pil
def test_text_cast_image(text_file_with_image):
with open(text_file_with_image, encoding="utf-8") as f:
image_file = f.read().splitlines()[0]
text = Text(encoding="utf-8", features=Features({"image": Image()}))
generator = text._generate_tables([[text_file_with_image]])
pa_table = pa.concat_tables([table for _, table in generator])
assert pa_table.schema.field("image").type == Image()()
generated_content = pa_table.to_pydict()["image"]
assert generated_content == [{"path": image_file, "bytes": None}]
@pytest.mark.parametrize("sample_by", ["line", "paragraph", "document"])
def test_text_sample_by(sample_by, text_file):
with open(text_file, encoding="utf-8") as f:
expected_content = f.read()
if sample_by == "line":
expected_content = expected_content.splitlines()
elif sample_by == "paragraph":
expected_content = expected_content.split("\n\n")
elif sample_by == "document":
expected_content = [expected_content]
text = Text(sample_by=sample_by, encoding="utf-8", chunksize=100)
generator = text._generate_tables([[text_file]])
generated_content = pa.concat_tables([table for _, table in generator]).to_pydict()["text"]
assert generated_content == expected_content
| 0
|
hf_public_repos/datasets/tests
|
hf_public_repos/datasets/tests/packaged_modules/test_audiofolder.py
|
import shutil
import textwrap
import librosa
import numpy as np
import pytest
import soundfile as sf
from datasets import Audio, ClassLabel, Features, Value
from datasets.data_files import DataFilesDict, get_data_patterns
from datasets.download.streaming_download_manager import StreamingDownloadManager
from datasets.packaged_modules.audiofolder.audiofolder import AudioFolder
from ..utils import require_sndfile
@pytest.fixture
def cache_dir(tmp_path):
return str(tmp_path / "audiofolder_cache_dir")
@pytest.fixture
def data_files_with_labels_no_metadata(tmp_path, audio_file):
data_dir = tmp_path / "data_files_with_labels_no_metadata"
data_dir.mkdir(parents=True, exist_ok=True)
subdir_class_0 = data_dir / "fr"
subdir_class_0.mkdir(parents=True, exist_ok=True)
subdir_class_1 = data_dir / "uk"
subdir_class_1.mkdir(parents=True, exist_ok=True)
audio_filename = subdir_class_0 / "audio_fr.wav"
shutil.copyfile(audio_file, audio_filename)
audio_filename2 = subdir_class_1 / "audio_uk.wav"
shutil.copyfile(audio_file, audio_filename2)
data_files_with_labels_no_metadata = DataFilesDict.from_patterns(
get_data_patterns(str(data_dir)), data_dir.as_posix()
)
return data_files_with_labels_no_metadata
@pytest.fixture
def audio_files_with_labels_and_duplicated_label_key_in_metadata(tmp_path, audio_file):
data_dir = tmp_path / "audio_files_with_labels_and_label_key_in_metadata"
data_dir.mkdir(parents=True, exist_ok=True)
subdir_class_0 = data_dir / "fr"
subdir_class_0.mkdir(parents=True, exist_ok=True)
subdir_class_1 = data_dir / "uk"
subdir_class_1.mkdir(parents=True, exist_ok=True)
audio_filename = subdir_class_0 / "audio_fr.wav"
shutil.copyfile(audio_file, audio_filename)
audio_filename2 = subdir_class_1 / "audio_uk.wav"
shutil.copyfile(audio_file, audio_filename2)
audio_metadata_filename = tmp_path / data_dir / "metadata.jsonl"
audio_metadata = textwrap.dedent(
"""\
{"file_name": "fr/audio_fr.wav", "text": "Audio in French", "label": "Fr"}
{"file_name": "uk/audio_uk.wav", "text": "Audio in Ukrainian", "label": "Uk"}
"""
)
with open(audio_metadata_filename, "w", encoding="utf-8") as f:
f.write(audio_metadata)
return str(audio_filename), str(audio_filename2), str(audio_metadata_filename)
@pytest.fixture
def audio_file_with_metadata(tmp_path, audio_file):
audio_filename = tmp_path / "audio_file.wav"
shutil.copyfile(audio_file, audio_filename)
audio_metadata_filename = tmp_path / "metadata.jsonl"
audio_metadata = textwrap.dedent(
"""\
{"file_name": "audio_file.wav", "text": "Audio transcription"}
"""
)
with open(audio_metadata_filename, "w", encoding="utf-8") as f:
f.write(audio_metadata)
return str(audio_filename), str(audio_metadata_filename)
@pytest.fixture
def audio_files_with_metadata_that_misses_one_audio(tmp_path, audio_file):
audio_filename = tmp_path / "audio_file.wav"
shutil.copyfile(audio_file, audio_filename)
audio_filename2 = tmp_path / "audio_file2.wav"
shutil.copyfile(audio_file, audio_filename2)
audio_metadata_filename = tmp_path / "metadata.jsonl"
audio_metadata = textwrap.dedent(
"""\
{"file_name": "audio_file.wav", "text": "Audio transcription"}
"""
)
with open(audio_metadata_filename, "w", encoding="utf-8") as f:
f.write(audio_metadata)
return str(audio_filename), str(audio_filename2), str(audio_metadata_filename)
@pytest.fixture
def data_files_with_one_split_and_metadata(tmp_path, audio_file):
data_dir = tmp_path / "audiofolder_data_dir_with_metadata"
data_dir.mkdir(parents=True, exist_ok=True)
subdir = data_dir / "subdir"
subdir.mkdir(parents=True, exist_ok=True)
audio_filename = data_dir / "audio_file.wav"
shutil.copyfile(audio_file, audio_filename)
audio_filename2 = data_dir / "audio_file2.wav"
shutil.copyfile(audio_file, audio_filename2)
audio_filename3 = subdir / "audio_file3.wav" # in subdir
shutil.copyfile(audio_file, audio_filename3)
audio_metadata_filename = data_dir / "metadata.jsonl"
audio_metadata = textwrap.dedent(
"""\
{"file_name": "audio_file.wav", "text": "First audio transcription"}
{"file_name": "audio_file2.wav", "text": "Second audio transcription"}
{"file_name": "subdir/audio_file3.wav", "text": "Third audio transcription (in subdir)"}
"""
)
with open(audio_metadata_filename, "w", encoding="utf-8") as f:
f.write(audio_metadata)
data_files_with_one_split_and_metadata = DataFilesDict.from_patterns(
get_data_patterns(str(data_dir)), data_dir.as_posix()
)
assert len(data_files_with_one_split_and_metadata) == 1
assert len(data_files_with_one_split_and_metadata["train"]) == 4
return data_files_with_one_split_and_metadata
@pytest.fixture(params=["jsonl", "csv"])
def data_files_with_two_splits_and_metadata(request, tmp_path, audio_file):
data_dir = tmp_path / "audiofolder_data_dir_with_metadata"
data_dir.mkdir(parents=True, exist_ok=True)
train_dir = data_dir / "train"
train_dir.mkdir(parents=True, exist_ok=True)
test_dir = data_dir / "test"
test_dir.mkdir(parents=True, exist_ok=True)
audio_filename = train_dir / "audio_file.wav" # train audio
shutil.copyfile(audio_file, audio_filename)
audio_filename2 = train_dir / "audio_file2.wav" # train audio
shutil.copyfile(audio_file, audio_filename2)
audio_filename3 = test_dir / "audio_file3.wav" # test audio
shutil.copyfile(audio_file, audio_filename3)
train_audio_metadata_filename = train_dir / f"metadata.{request.param}"
audio_metadata = (
textwrap.dedent(
"""\
{"file_name": "audio_file.wav", "text": "First train audio transcription"}
{"file_name": "audio_file2.wav", "text": "Second train audio transcription"}
"""
)
if request.param == "jsonl"
else textwrap.dedent(
"""\
file_name,text
audio_file.wav,First train audio transcription
audio_file2.wav,Second train audio transcription
"""
)
)
with open(train_audio_metadata_filename, "w", encoding="utf-8") as f:
f.write(audio_metadata)
test_audio_metadata_filename = test_dir / f"metadata.{request.param}"
audio_metadata = (
textwrap.dedent(
"""\
{"file_name": "audio_file3.wav", "text": "Test audio transcription"}
"""
)
if request.param == "jsonl"
else textwrap.dedent(
"""\
file_name,text
audio_file3.wav,Test audio transcription
"""
)
)
with open(test_audio_metadata_filename, "w", encoding="utf-8") as f:
f.write(audio_metadata)
data_files_with_two_splits_and_metadata = DataFilesDict.from_patterns(
get_data_patterns(str(data_dir)), data_dir.as_posix()
)
assert len(data_files_with_two_splits_and_metadata) == 2
assert len(data_files_with_two_splits_and_metadata["train"]) == 3
assert len(data_files_with_two_splits_and_metadata["test"]) == 2
return data_files_with_two_splits_and_metadata
@pytest.fixture
def data_files_with_zip_archives(tmp_path, audio_file):
data_dir = tmp_path / "audiofolder_data_dir_with_zip_archives"
data_dir.mkdir(parents=True, exist_ok=True)
archive_dir = data_dir / "archive"
archive_dir.mkdir(parents=True, exist_ok=True)
subdir = archive_dir / "subdir"
subdir.mkdir(parents=True, exist_ok=True)
audio_filename = archive_dir / "audio_file.wav"
shutil.copyfile(audio_file, audio_filename)
audio_filename2 = subdir / "audio_file2.wav" # in subdir
# make sure they're two different audios
# Indeed we won't be able to compare the audio filenames, since the archive is not extracted in streaming mode
array, sampling_rate = librosa.load(str(audio_filename), sr=16000) # original sampling rate is 44100
sf.write(str(audio_filename2), array, samplerate=16000)
audio_metadata_filename = archive_dir / "metadata.jsonl"
audio_metadata = textwrap.dedent(
"""\
{"file_name": "audio_file.wav", "text": "First audio transcription"}
{"file_name": "subdir/audio_file2.wav", "text": "Second audio transcription (in subdir)"}
"""
)
with open(audio_metadata_filename, "w", encoding="utf-8") as f:
f.write(audio_metadata)
shutil.make_archive(str(archive_dir), "zip", archive_dir)
shutil.rmtree(str(archive_dir))
data_files_with_zip_archives = DataFilesDict.from_patterns(get_data_patterns(str(data_dir)), data_dir.as_posix())
assert len(data_files_with_zip_archives) == 1
assert len(data_files_with_zip_archives["train"]) == 1
return data_files_with_zip_archives
@require_sndfile
# check that labels are inferred correctly from dir names
def test_generate_examples_with_labels(data_files_with_labels_no_metadata, cache_dir):
# there are no metadata.jsonl files in this test case
audiofolder = AudioFolder(data_files=data_files_with_labels_no_metadata, cache_dir=cache_dir, drop_labels=False)
audiofolder.download_and_prepare()
assert audiofolder.info.features == Features({"audio": Audio(), "label": ClassLabel(names=["fr", "uk"])})
dataset = list(audiofolder.as_dataset()["train"])
label_feature = audiofolder.info.features["label"]
assert dataset[0]["label"] == label_feature._str2int["fr"]
assert dataset[1]["label"] == label_feature._str2int["uk"]
@require_sndfile
@pytest.mark.parametrize("drop_metadata", [None, True, False])
@pytest.mark.parametrize("drop_labels", [None, True, False])
def test_generate_examples_duplicated_label_key(
audio_files_with_labels_and_duplicated_label_key_in_metadata, drop_metadata, drop_labels, cache_dir, caplog
):
fr_audio_file, uk_audio_file, audio_metadata_file = audio_files_with_labels_and_duplicated_label_key_in_metadata
audiofolder = AudioFolder(
drop_metadata=drop_metadata,
drop_labels=drop_labels,
data_files=[fr_audio_file, uk_audio_file, audio_metadata_file],
cache_dir=cache_dir,
)
if drop_labels is False:
# infer labels from directories even if metadata files are found
audiofolder.download_and_prepare()
warning_in_logs = any("ignoring metadata columns" in record.msg.lower() for record in caplog.records)
assert warning_in_logs if drop_metadata is not True else not warning_in_logs
dataset = audiofolder.as_dataset()["train"]
assert audiofolder.info.features["label"] == ClassLabel(names=["fr", "uk"])
assert all(example["label"] in audiofolder.info.features["label"]._str2int.values() for example in dataset)
else:
audiofolder.download_and_prepare()
dataset = audiofolder.as_dataset()["train"]
if drop_metadata is not True:
# labels are from metadata
assert audiofolder.info.features["label"] == Value("string")
assert all(example["label"] in ["Fr", "Uk"] for example in dataset)
else:
# drop both labels and metadata
assert audiofolder.info.features == Features({"audio": Audio()})
assert all(example.keys() == {"audio"} for example in dataset)
@require_sndfile
@pytest.mark.parametrize("drop_metadata", [None, True, False])
@pytest.mark.parametrize("drop_labels", [None, True, False])
def test_generate_examples_drop_labels(data_files_with_labels_no_metadata, drop_metadata, drop_labels):
audiofolder = AudioFolder(
drop_metadata=drop_metadata, drop_labels=drop_labels, data_files=data_files_with_labels_no_metadata
)
gen_kwargs = audiofolder._split_generators(StreamingDownloadManager())[0].gen_kwargs
# removing the labels explicitly requires drop_labels=True
assert gen_kwargs["add_labels"] is not bool(drop_labels)
assert gen_kwargs["add_metadata"] is False # metadata files is not present in this case
generator = audiofolder._generate_examples(**gen_kwargs)
if not drop_labels:
assert all(
example.keys() == {"audio", "label"} and all(val is not None for val in example.values())
for _, example in generator
)
else:
assert all(
example.keys() == {"audio"} and all(val is not None for val in example.values())
for _, example in generator
)
@require_sndfile
@pytest.mark.parametrize("drop_metadata", [None, True, False])
@pytest.mark.parametrize("drop_labels", [None, True, False])
def test_generate_examples_drop_metadata(audio_file_with_metadata, drop_metadata, drop_labels):
audio_file, audio_metadata_file = audio_file_with_metadata
audiofolder = AudioFolder(
drop_metadata=drop_metadata, drop_labels=drop_labels, data_files={"train": [audio_file, audio_metadata_file]}
)
gen_kwargs = audiofolder._split_generators(StreamingDownloadManager())[0].gen_kwargs
# since the dataset has metadata, removing the metadata explicitly requires drop_metadata=True
assert gen_kwargs["add_metadata"] is not bool(drop_metadata)
# since the dataset has metadata, adding the labels explicitly requires drop_labels=False
assert gen_kwargs["add_labels"] is (drop_labels is False)
generator = audiofolder._generate_examples(**gen_kwargs)
expected_columns = {"audio"}
if gen_kwargs["add_metadata"]:
expected_columns.add("text")
if gen_kwargs["add_labels"]:
expected_columns.add("label")
result = [example for _, example in generator]
assert len(result) == 1
example = result[0]
assert example.keys() == expected_columns
for column in expected_columns:
assert example[column] is not None
@require_sndfile
@pytest.mark.parametrize("drop_metadata", [None, True, False])
def test_generate_examples_with_metadata_in_wrong_location(audio_file, audio_file_with_metadata, drop_metadata):
_, audio_metadata_file = audio_file_with_metadata
audiofolder = AudioFolder(drop_metadata=drop_metadata, data_files={"train": [audio_file, audio_metadata_file]})
gen_kwargs = audiofolder._split_generators(StreamingDownloadManager())[0].gen_kwargs
generator = audiofolder._generate_examples(**gen_kwargs)
if not drop_metadata:
with pytest.raises(ValueError):
list(generator)
else:
assert all(
example.keys() == {"audio"} and all(val is not None for val in example.values())
for _, example in generator
)
@require_sndfile
@pytest.mark.parametrize("drop_metadata", [None, True, False])
def test_generate_examples_with_metadata_that_misses_one_audio(
audio_files_with_metadata_that_misses_one_audio, drop_metadata
):
audio_file, audio_file2, audio_metadata_file = audio_files_with_metadata_that_misses_one_audio
if not drop_metadata:
features = Features({"audio": Audio(), "text": Value("string")})
else:
features = Features({"audio": Audio()})
audiofolder = AudioFolder(
drop_metadata=drop_metadata,
features=features,
data_files={"train": [audio_file, audio_file2, audio_metadata_file]},
)
gen_kwargs = audiofolder._split_generators(StreamingDownloadManager())[0].gen_kwargs
generator = audiofolder._generate_examples(**gen_kwargs)
if not drop_metadata:
with pytest.raises(ValueError):
_ = list(generator)
else:
assert all(
example.keys() == {"audio"} and all(val is not None for val in example.values())
for _, example in generator
)
@require_sndfile
@pytest.mark.parametrize("streaming", [False, True])
def test_data_files_with_metadata_and_single_split(streaming, cache_dir, data_files_with_one_split_and_metadata):
data_files = data_files_with_one_split_and_metadata
audiofolder = AudioFolder(data_files=data_files, cache_dir=cache_dir)
audiofolder.download_and_prepare()
datasets = audiofolder.as_streaming_dataset() if streaming else audiofolder.as_dataset()
for split, data_files in data_files.items():
expected_num_of_audios = len(data_files) - 1 # don't count the metadata file
assert split in datasets
dataset = list(datasets[split])
assert len(dataset) == expected_num_of_audios
# make sure each sample has its own audio and metadata
assert len({example["audio"]["path"] for example in dataset}) == expected_num_of_audios
assert len({example["text"] for example in dataset}) == expected_num_of_audios
assert all(example["text"] is not None for example in dataset)
@require_sndfile
@pytest.mark.parametrize("streaming", [False, True])
def test_data_files_with_metadata_and_multiple_splits(streaming, cache_dir, data_files_with_two_splits_and_metadata):
data_files = data_files_with_two_splits_and_metadata
audiofolder = AudioFolder(data_files=data_files, cache_dir=cache_dir)
audiofolder.download_and_prepare()
datasets = audiofolder.as_streaming_dataset() if streaming else audiofolder.as_dataset()
for split, data_files in data_files.items():
expected_num_of_audios = len(data_files) - 1 # don't count the metadata file
assert split in datasets
dataset = list(datasets[split])
assert len(dataset) == expected_num_of_audios
# make sure each sample has its own audio and metadata
assert len({example["audio"]["path"] for example in dataset}) == expected_num_of_audios
assert len({example["text"] for example in dataset}) == expected_num_of_audios
assert all(example["text"] is not None for example in dataset)
@require_sndfile
@pytest.mark.parametrize("streaming", [False, True])
def test_data_files_with_metadata_and_archives(streaming, cache_dir, data_files_with_zip_archives):
audiofolder = AudioFolder(data_files=data_files_with_zip_archives, cache_dir=cache_dir)
audiofolder.download_and_prepare()
datasets = audiofolder.as_streaming_dataset() if streaming else audiofolder.as_dataset()
for split, data_files in data_files_with_zip_archives.items():
num_of_archives = len(data_files) # the metadata file is inside the archive
expected_num_of_audios = 2 * num_of_archives
assert split in datasets
dataset = list(datasets[split])
assert len(dataset) == expected_num_of_audios
# make sure each sample has its own audio (all arrays are different) and metadata
assert (
sum(np.array_equal(dataset[0]["audio"]["array"], example["audio"]["array"]) for example in dataset[1:])
== 0
)
assert len({example["text"] for example in dataset}) == expected_num_of_audios
assert all(example["text"] is not None for example in dataset)
@require_sndfile
def test_data_files_with_wrong_metadata_file_name(cache_dir, tmp_path, audio_file):
data_dir = tmp_path / "data_dir_with_bad_metadata"
data_dir.mkdir(parents=True, exist_ok=True)
shutil.copyfile(audio_file, data_dir / "audio_file.wav")
audio_metadata_filename = data_dir / "bad_metadata.jsonl" # bad file
audio_metadata = textwrap.dedent(
"""\
{"file_name": "audio_file.wav", "text": "Audio transcription"}
"""
)
with open(audio_metadata_filename, "w", encoding="utf-8") as f:
f.write(audio_metadata)
data_files_with_bad_metadata = DataFilesDict.from_patterns(get_data_patterns(str(data_dir)), data_dir.as_posix())
audiofolder = AudioFolder(data_files=data_files_with_bad_metadata, cache_dir=cache_dir)
audiofolder.download_and_prepare()
dataset = audiofolder.as_dataset(split="train")
# check that there are no metadata, since the metadata file name doesn't have the right name
assert "text" not in dataset.column_names
@require_sndfile
def test_data_files_with_wrong_audio_file_name_column_in_metadata_file(cache_dir, tmp_path, audio_file):
data_dir = tmp_path / "data_dir_with_bad_metadata"
data_dir.mkdir(parents=True, exist_ok=True)
shutil.copyfile(audio_file, data_dir / "audio_file.wav")
audio_metadata_filename = data_dir / "metadata.jsonl"
audio_metadata = textwrap.dedent( # with bad column "bad_file_name" instead of "file_name"
"""\
{"bad_file_name_column": "audio_file.wav", "text": "Audio transcription"}
"""
)
with open(audio_metadata_filename, "w", encoding="utf-8") as f:
f.write(audio_metadata)
data_files_with_bad_metadata = DataFilesDict.from_patterns(get_data_patterns(str(data_dir)), data_dir.as_posix())
audiofolder = AudioFolder(data_files=data_files_with_bad_metadata, cache_dir=cache_dir)
with pytest.raises(ValueError) as exc_info:
audiofolder.download_and_prepare()
assert "`file_name` must be present" in str(exc_info.value)
@require_sndfile
def test_data_files_with_with_metadata_in_different_formats(cache_dir, tmp_path, audio_file):
data_dir = tmp_path / "data_dir_with_metadata_in_different_format"
data_dir.mkdir(parents=True, exist_ok=True)
shutil.copyfile(audio_file, data_dir / "audio_file.wav")
audio_metadata_filename_jsonl = data_dir / "metadata.jsonl"
audio_metadata_jsonl = textwrap.dedent(
"""\
{"file_name": "audio_file.wav", "text": "Audio transcription"}
"""
)
with open(audio_metadata_filename_jsonl, "w", encoding="utf-8") as f:
f.write(audio_metadata_jsonl)
audio_metadata_filename_csv = data_dir / "metadata.csv"
audio_metadata_csv = textwrap.dedent(
"""\
file_name,text
audio_file.wav,Audio transcription
"""
)
with open(audio_metadata_filename_csv, "w", encoding="utf-8") as f:
f.write(audio_metadata_csv)
data_files_with_bad_metadata = DataFilesDict.from_patterns(get_data_patterns(str(data_dir)), data_dir.as_posix())
audiofolder = AudioFolder(data_files=data_files_with_bad_metadata, cache_dir=cache_dir)
with pytest.raises(ValueError) as exc_info:
audiofolder.download_and_prepare()
assert "metadata files with different extensions" in str(exc_info.value)
| 0
|
hf_public_repos/datasets/tests
|
hf_public_repos/datasets/tests/io/test_json.py
|
import io
import json
import fsspec
import pytest
from datasets import Dataset, DatasetDict, Features, NamedSplit, Value
from datasets.io.json import JsonDatasetReader, JsonDatasetWriter
from ..utils import assert_arrow_memory_doesnt_increase, assert_arrow_memory_increases
def _check_json_dataset(dataset, expected_features):
assert isinstance(dataset, Dataset)
assert dataset.num_rows == 4
assert dataset.num_columns == 3
assert dataset.column_names == ["col_1", "col_2", "col_3"]
for feature, expected_dtype in expected_features.items():
assert dataset.features[feature].dtype == expected_dtype
@pytest.mark.parametrize("keep_in_memory", [False, True])
def test_dataset_from_json_keep_in_memory(keep_in_memory, jsonl_path, tmp_path):
cache_dir = tmp_path / "cache"
expected_features = {"col_1": "string", "col_2": "int64", "col_3": "float64"}
with assert_arrow_memory_increases() if keep_in_memory else assert_arrow_memory_doesnt_increase():
dataset = JsonDatasetReader(jsonl_path, cache_dir=cache_dir, keep_in_memory=keep_in_memory).read()
_check_json_dataset(dataset, expected_features)
@pytest.mark.parametrize(
"features",
[
None,
{"col_1": "string", "col_2": "int64", "col_3": "float64"},
{"col_1": "string", "col_2": "string", "col_3": "string"},
{"col_1": "int32", "col_2": "int32", "col_3": "int32"},
{"col_1": "float32", "col_2": "float32", "col_3": "float32"},
],
)
def test_dataset_from_json_features(features, jsonl_path, tmp_path):
cache_dir = tmp_path / "cache"
default_expected_features = {"col_1": "string", "col_2": "int64", "col_3": "float64"}
expected_features = features.copy() if features else default_expected_features
features = (
Features({feature: Value(dtype) for feature, dtype in features.items()}) if features is not None else None
)
dataset = JsonDatasetReader(jsonl_path, features=features, cache_dir=cache_dir).read()
_check_json_dataset(dataset, expected_features)
@pytest.mark.parametrize(
"features",
[
None,
{"col_3": "float64", "col_1": "string", "col_2": "int64"},
],
)
def test_dataset_from_json_with_unsorted_column_names(features, jsonl_312_path, tmp_path):
cache_dir = tmp_path / "cache"
default_expected_features = {"col_3": "float64", "col_1": "string", "col_2": "int64"}
expected_features = features.copy() if features else default_expected_features
features = (
Features({feature: Value(dtype) for feature, dtype in features.items()}) if features is not None else None
)
dataset = JsonDatasetReader(jsonl_312_path, features=features, cache_dir=cache_dir).read()
assert isinstance(dataset, Dataset)
assert dataset.num_rows == 2
assert dataset.num_columns == 3
assert dataset.column_names == ["col_3", "col_1", "col_2"]
for feature, expected_dtype in expected_features.items():
assert dataset.features[feature].dtype == expected_dtype
def test_dataset_from_json_with_mismatched_features(jsonl_312_path, tmp_path):
# jsonl_312_path features are {"col_3": "float64", "col_1": "string", "col_2": "int64"}
features = {"col_2": "int64", "col_3": "float64", "col_1": "string"}
expected_features = features.copy()
features = (
Features({feature: Value(dtype) for feature, dtype in features.items()}) if features is not None else None
)
cache_dir = tmp_path / "cache"
dataset = JsonDatasetReader(jsonl_312_path, features=features, cache_dir=cache_dir).read()
assert isinstance(dataset, Dataset)
assert dataset.num_rows == 2
assert dataset.num_columns == 3
assert dataset.column_names == ["col_2", "col_3", "col_1"]
for feature, expected_dtype in expected_features.items():
assert dataset.features[feature].dtype == expected_dtype
@pytest.mark.parametrize("split", [None, NamedSplit("train"), "train", "test"])
def test_dataset_from_json_split(split, jsonl_path, tmp_path):
cache_dir = tmp_path / "cache"
expected_features = {"col_1": "string", "col_2": "int64", "col_3": "float64"}
dataset = JsonDatasetReader(jsonl_path, cache_dir=cache_dir, split=split).read()
_check_json_dataset(dataset, expected_features)
assert dataset.split == split if split else "train"
@pytest.mark.parametrize("path_type", [str, list])
def test_dataset_from_json_path_type(path_type, jsonl_path, tmp_path):
if issubclass(path_type, str):
path = jsonl_path
elif issubclass(path_type, list):
path = [jsonl_path]
cache_dir = tmp_path / "cache"
expected_features = {"col_1": "string", "col_2": "int64", "col_3": "float64"}
dataset = JsonDatasetReader(path, cache_dir=cache_dir).read()
_check_json_dataset(dataset, expected_features)
def _check_json_datasetdict(dataset_dict, expected_features, splits=("train",)):
assert isinstance(dataset_dict, DatasetDict)
for split in splits:
dataset = dataset_dict[split]
assert dataset.num_rows == 4
assert dataset.num_columns == 3
assert dataset.column_names == ["col_1", "col_2", "col_3"]
for feature, expected_dtype in expected_features.items():
assert dataset.features[feature].dtype == expected_dtype
@pytest.mark.parametrize("keep_in_memory", [False, True])
def test_datasetdict_from_json_keep_in_memory(keep_in_memory, jsonl_path, tmp_path):
cache_dir = tmp_path / "cache"
expected_features = {"col_1": "string", "col_2": "int64", "col_3": "float64"}
with assert_arrow_memory_increases() if keep_in_memory else assert_arrow_memory_doesnt_increase():
dataset = JsonDatasetReader({"train": jsonl_path}, cache_dir=cache_dir, keep_in_memory=keep_in_memory).read()
_check_json_datasetdict(dataset, expected_features)
@pytest.mark.parametrize(
"features",
[
None,
{"col_1": "string", "col_2": "int64", "col_3": "float64"},
{"col_1": "string", "col_2": "string", "col_3": "string"},
{"col_1": "int32", "col_2": "int32", "col_3": "int32"},
{"col_1": "float32", "col_2": "float32", "col_3": "float32"},
],
)
def test_datasetdict_from_json_features(features, jsonl_path, tmp_path):
cache_dir = tmp_path / "cache"
default_expected_features = {"col_1": "string", "col_2": "int64", "col_3": "float64"}
expected_features = features.copy() if features else default_expected_features
features = (
Features({feature: Value(dtype) for feature, dtype in features.items()}) if features is not None else None
)
dataset = JsonDatasetReader({"train": jsonl_path}, features=features, cache_dir=cache_dir).read()
_check_json_datasetdict(dataset, expected_features)
@pytest.mark.parametrize("split", [None, NamedSplit("train"), "train", "test"])
def test_datasetdict_from_json_splits(split, jsonl_path, tmp_path):
if split:
path = {split: jsonl_path}
else:
split = "train"
path = {"train": jsonl_path, "test": jsonl_path}
cache_dir = tmp_path / "cache"
expected_features = {"col_1": "string", "col_2": "int64", "col_3": "float64"}
dataset = JsonDatasetReader(path, cache_dir=cache_dir).read()
_check_json_datasetdict(dataset, expected_features, splits=list(path.keys()))
assert all(dataset[split].split == split for split in path.keys())
def load_json(buffer):
return json.load(buffer)
def load_json_lines(buffer):
return [json.loads(line) for line in buffer]
class TestJsonDatasetWriter:
@pytest.mark.parametrize("lines, load_json_function", [(True, load_json_lines), (False, load_json)])
def test_dataset_to_json_lines(self, lines, load_json_function, dataset):
with io.BytesIO() as buffer:
JsonDatasetWriter(dataset, buffer, lines=lines).write()
buffer.seek(0)
exported_content = load_json_function(buffer)
assert isinstance(exported_content, list)
assert isinstance(exported_content[0], dict)
assert len(exported_content) == 10
@pytest.mark.parametrize(
"orient, container, keys, len_at",
[
("records", list, {"tokens", "labels", "answers", "id"}, None),
("split", dict, {"columns", "data"}, "data"),
("index", dict, set("0123456789"), None),
("columns", dict, {"tokens", "labels", "answers", "id"}, "tokens"),
("values", list, None, None),
("table", dict, {"schema", "data"}, "data"),
],
)
def test_dataset_to_json_orient(self, orient, container, keys, len_at, dataset):
with io.BytesIO() as buffer:
JsonDatasetWriter(dataset, buffer, lines=False, orient=orient).write()
buffer.seek(0)
exported_content = load_json(buffer)
assert isinstance(exported_content, container)
if keys:
if container is dict:
assert exported_content.keys() == keys
else:
assert exported_content[0].keys() == keys
else:
assert not hasattr(exported_content, "keys") and not hasattr(exported_content[0], "keys")
if len_at:
assert len(exported_content[len_at]) == 10
else:
assert len(exported_content) == 10
@pytest.mark.parametrize("lines, load_json_function", [(True, load_json_lines), (False, load_json)])
def test_dataset_to_json_lines_multiproc(self, lines, load_json_function, dataset):
with io.BytesIO() as buffer:
JsonDatasetWriter(dataset, buffer, lines=lines, num_proc=2).write()
buffer.seek(0)
exported_content = load_json_function(buffer)
assert isinstance(exported_content, list)
assert isinstance(exported_content[0], dict)
assert len(exported_content) == 10
@pytest.mark.parametrize(
"orient, container, keys, len_at",
[
("records", list, {"tokens", "labels", "answers", "id"}, None),
("split", dict, {"columns", "data"}, "data"),
("index", dict, set("0123456789"), None),
("columns", dict, {"tokens", "labels", "answers", "id"}, "tokens"),
("values", list, None, None),
("table", dict, {"schema", "data"}, "data"),
],
)
def test_dataset_to_json_orient_multiproc(self, orient, container, keys, len_at, dataset):
with io.BytesIO() as buffer:
JsonDatasetWriter(dataset, buffer, lines=False, orient=orient, num_proc=2).write()
buffer.seek(0)
exported_content = load_json(buffer)
assert isinstance(exported_content, container)
if keys:
if container is dict:
assert exported_content.keys() == keys
else:
assert exported_content[0].keys() == keys
else:
assert not hasattr(exported_content, "keys") and not hasattr(exported_content[0], "keys")
if len_at:
assert len(exported_content[len_at]) == 10
else:
assert len(exported_content) == 10
def test_dataset_to_json_orient_invalidproc(self, dataset):
with pytest.raises(ValueError):
with io.BytesIO() as buffer:
JsonDatasetWriter(dataset, buffer, num_proc=0)
@pytest.mark.parametrize("compression, extension", [("gzip", "gz"), ("bz2", "bz2"), ("xz", "xz")])
def test_dataset_to_json_compression(self, shared_datadir, tmp_path_factory, extension, compression, dataset):
path = tmp_path_factory.mktemp("data") / f"test.json.{extension}"
original_path = str(shared_datadir / f"test_file.json.{extension}")
JsonDatasetWriter(dataset, path, compression=compression).write()
with fsspec.open(path, "rb", compression="infer") as f:
exported_content = f.read()
with fsspec.open(original_path, "rb", compression="infer") as f:
original_content = f.read()
assert exported_content == original_content
| 0
|
hf_public_repos/datasets/tests
|
hf_public_repos/datasets/tests/io/test_parquet.py
|
import pyarrow.parquet as pq
import pytest
from datasets import Audio, Dataset, DatasetDict, Features, IterableDatasetDict, NamedSplit, Sequence, Value, config
from datasets.features.image import Image
from datasets.info import DatasetInfo
from datasets.io.parquet import ParquetDatasetReader, ParquetDatasetWriter, get_writer_batch_size
from ..utils import assert_arrow_memory_doesnt_increase, assert_arrow_memory_increases
def _check_parquet_dataset(dataset, expected_features):
assert isinstance(dataset, Dataset)
assert dataset.num_rows == 4
assert dataset.num_columns == 3
assert dataset.column_names == ["col_1", "col_2", "col_3"]
for feature, expected_dtype in expected_features.items():
assert dataset.features[feature].dtype == expected_dtype
@pytest.mark.parametrize("keep_in_memory", [False, True])
def test_dataset_from_parquet_keep_in_memory(keep_in_memory, parquet_path, tmp_path):
cache_dir = tmp_path / "cache"
expected_features = {"col_1": "string", "col_2": "int64", "col_3": "float64"}
with assert_arrow_memory_increases() if keep_in_memory else assert_arrow_memory_doesnt_increase():
dataset = ParquetDatasetReader(parquet_path, cache_dir=cache_dir, keep_in_memory=keep_in_memory).read()
_check_parquet_dataset(dataset, expected_features)
@pytest.mark.parametrize(
"features",
[
None,
{"col_1": "string", "col_2": "int64", "col_3": "float64"},
{"col_1": "string", "col_2": "string", "col_3": "string"},
{"col_1": "int32", "col_2": "int32", "col_3": "int32"},
{"col_1": "float32", "col_2": "float32", "col_3": "float32"},
],
)
def test_dataset_from_parquet_features(features, parquet_path, tmp_path):
cache_dir = tmp_path / "cache"
default_expected_features = {"col_1": "string", "col_2": "int64", "col_3": "float64"}
expected_features = features.copy() if features else default_expected_features
features = (
Features({feature: Value(dtype) for feature, dtype in features.items()}) if features is not None else None
)
dataset = ParquetDatasetReader(parquet_path, features=features, cache_dir=cache_dir).read()
_check_parquet_dataset(dataset, expected_features)
@pytest.mark.parametrize("split", [None, NamedSplit("train"), "train", "test"])
def test_dataset_from_parquet_split(split, parquet_path, tmp_path):
cache_dir = tmp_path / "cache"
expected_features = {"col_1": "string", "col_2": "int64", "col_3": "float64"}
dataset = ParquetDatasetReader(parquet_path, cache_dir=cache_dir, split=split).read()
_check_parquet_dataset(dataset, expected_features)
assert dataset.split == split if split else "train"
@pytest.mark.parametrize("path_type", [str, list])
def test_dataset_from_parquet_path_type(path_type, parquet_path, tmp_path):
if issubclass(path_type, str):
path = parquet_path
elif issubclass(path_type, list):
path = [parquet_path]
cache_dir = tmp_path / "cache"
expected_features = {"col_1": "string", "col_2": "int64", "col_3": "float64"}
dataset = ParquetDatasetReader(path, cache_dir=cache_dir).read()
_check_parquet_dataset(dataset, expected_features)
def _check_parquet_datasetdict(dataset_dict, expected_features, splits=("train",)):
assert isinstance(dataset_dict, (DatasetDict, IterableDatasetDict))
for split in splits:
dataset = dataset_dict[split]
assert len(list(dataset)) == 4
assert dataset.features is not None
assert set(dataset.features) == set(expected_features)
for feature, expected_dtype in expected_features.items():
assert dataset.features[feature].dtype == expected_dtype
@pytest.mark.parametrize("keep_in_memory", [False, True])
def test_parquet_datasetdict_reader_keep_in_memory(keep_in_memory, parquet_path, tmp_path):
cache_dir = tmp_path / "cache"
expected_features = {"col_1": "string", "col_2": "int64", "col_3": "float64"}
with assert_arrow_memory_increases() if keep_in_memory else assert_arrow_memory_doesnt_increase():
dataset = ParquetDatasetReader(
{"train": parquet_path}, cache_dir=cache_dir, keep_in_memory=keep_in_memory
).read()
_check_parquet_datasetdict(dataset, expected_features)
@pytest.mark.parametrize("streaming", [False, True])
@pytest.mark.parametrize(
"features",
[
None,
{"col_1": "string", "col_2": "int64", "col_3": "float64"},
{"col_1": "string", "col_2": "string", "col_3": "string"},
{"col_1": "int32", "col_2": "int32", "col_3": "int32"},
{"col_1": "float32", "col_2": "float32", "col_3": "float32"},
],
)
def test_parquet_datasetdict_reader_features(streaming, features, parquet_path, tmp_path):
cache_dir = tmp_path / "cache"
default_expected_features = {"col_1": "string", "col_2": "int64", "col_3": "float64"}
expected_features = features.copy() if features else default_expected_features
features = (
Features({feature: Value(dtype) for feature, dtype in features.items()}) if features is not None else None
)
dataset = ParquetDatasetReader(
{"train": parquet_path}, features=features, cache_dir=cache_dir, streaming=streaming
).read()
_check_parquet_datasetdict(dataset, expected_features)
@pytest.mark.parametrize("streaming", [False, True])
@pytest.mark.parametrize("columns", [None, ["col_1"]])
@pytest.mark.parametrize("pass_features", [False, True])
@pytest.mark.parametrize("pass_info", [False, True])
def test_parquet_datasetdict_reader_columns(streaming, columns, pass_features, pass_info, parquet_path, tmp_path):
cache_dir = tmp_path / "cache"
default_expected_features = {"col_1": "string", "col_2": "int64", "col_3": "float64"}
info = (
DatasetInfo(features=Features({feature: Value(dtype) for feature, dtype in default_expected_features.items()}))
if pass_info
else None
)
expected_features = (
{col: default_expected_features[col] for col in columns} if columns else default_expected_features
)
features = (
Features({feature: Value(dtype) for feature, dtype in expected_features.items()}) if pass_features else None
)
dataset = ParquetDatasetReader(
{"train": parquet_path},
columns=columns,
features=features,
info=info,
cache_dir=cache_dir,
streaming=streaming,
).read()
_check_parquet_datasetdict(dataset, expected_features)
@pytest.mark.parametrize("split", [None, NamedSplit("train"), "train", "test"])
def test_parquet_datasetdict_reader_split(split, parquet_path, tmp_path):
if split:
path = {split: parquet_path}
else:
split = "train"
path = {"train": parquet_path, "test": parquet_path}
cache_dir = tmp_path / "cache"
expected_features = {"col_1": "string", "col_2": "int64", "col_3": "float64"}
dataset = ParquetDatasetReader(path, cache_dir=cache_dir).read()
_check_parquet_datasetdict(dataset, expected_features, splits=list(path.keys()))
assert all(dataset[split].split == split for split in path.keys())
def test_parquet_write(dataset, tmp_path):
writer = ParquetDatasetWriter(dataset, tmp_path / "foo.parquet")
assert writer.write() > 0
pf = pq.ParquetFile(tmp_path / "foo.parquet")
output_table = pf.read()
assert dataset.data.table == output_table
def test_dataset_to_parquet_keeps_features(shared_datadir, tmp_path):
image_path = str(shared_datadir / "test_image_rgb.jpg")
data = {"image": [image_path]}
features = Features({"image": Image()})
dataset = Dataset.from_dict(data, features=features)
writer = ParquetDatasetWriter(dataset, tmp_path / "foo.parquet")
assert writer.write() > 0
reloaded_dataset = Dataset.from_parquet(str(tmp_path / "foo.parquet"))
assert dataset.features == reloaded_dataset.features
reloaded_iterable_dataset = ParquetDatasetReader(str(tmp_path / "foo.parquet"), streaming=True).read()
assert dataset.features == reloaded_iterable_dataset.features
@pytest.mark.parametrize(
"feature, expected",
[
(Features({"foo": Value("int32")}), None),
(Features({"image": Image(), "foo": Value("int32")}), config.PARQUET_ROW_GROUP_SIZE_FOR_IMAGE_DATASETS),
(Features({"nested": Sequence(Audio())}), config.PARQUET_ROW_GROUP_SIZE_FOR_AUDIO_DATASETS),
],
)
def test_get_writer_batch_size(feature, expected):
assert get_writer_batch_size(feature) == expected
| 0
|
hf_public_repos/datasets/tests
|
hf_public_repos/datasets/tests/io/test_sql.py
|
import contextlib
import os
import sqlite3
import pytest
from datasets import Dataset, Features, Value
from datasets.io.sql import SqlDatasetReader, SqlDatasetWriter
from ..utils import assert_arrow_memory_doesnt_increase, assert_arrow_memory_increases, require_sqlalchemy
def _check_sql_dataset(dataset, expected_features):
assert isinstance(dataset, Dataset)
assert dataset.num_rows == 4
assert dataset.num_columns == 3
assert dataset.column_names == ["col_1", "col_2", "col_3"]
for feature, expected_dtype in expected_features.items():
assert dataset.features[feature].dtype == expected_dtype
@require_sqlalchemy
@pytest.mark.parametrize("keep_in_memory", [False, True])
def test_dataset_from_sql_keep_in_memory(keep_in_memory, sqlite_path, tmp_path, set_sqlalchemy_silence_uber_warning):
cache_dir = tmp_path / "cache"
expected_features = {"col_1": "string", "col_2": "int64", "col_3": "float64"}
with assert_arrow_memory_increases() if keep_in_memory else assert_arrow_memory_doesnt_increase():
dataset = SqlDatasetReader(
"dataset", "sqlite:///" + sqlite_path, cache_dir=cache_dir, keep_in_memory=keep_in_memory
).read()
_check_sql_dataset(dataset, expected_features)
@require_sqlalchemy
@pytest.mark.parametrize(
"features",
[
None,
{"col_1": "string", "col_2": "int64", "col_3": "float64"},
{"col_1": "string", "col_2": "string", "col_3": "string"},
{"col_1": "int32", "col_2": "int32", "col_3": "int32"},
{"col_1": "float32", "col_2": "float32", "col_3": "float32"},
],
)
def test_dataset_from_sql_features(features, sqlite_path, tmp_path, set_sqlalchemy_silence_uber_warning):
cache_dir = tmp_path / "cache"
default_expected_features = {"col_1": "string", "col_2": "int64", "col_3": "float64"}
expected_features = features.copy() if features else default_expected_features
features = (
Features({feature: Value(dtype) for feature, dtype in features.items()}) if features is not None else None
)
dataset = SqlDatasetReader("dataset", "sqlite:///" + sqlite_path, features=features, cache_dir=cache_dir).read()
_check_sql_dataset(dataset, expected_features)
def iter_sql_file(sqlite_path):
with contextlib.closing(sqlite3.connect(sqlite_path)) as con:
cur = con.cursor()
cur.execute("SELECT * FROM dataset")
for row in cur:
yield row
@require_sqlalchemy
def test_dataset_to_sql(sqlite_path, tmp_path, set_sqlalchemy_silence_uber_warning):
cache_dir = tmp_path / "cache"
output_sqlite_path = os.path.join(cache_dir, "tmp.sql")
dataset = SqlDatasetReader("dataset", "sqlite:///" + sqlite_path, cache_dir=cache_dir).read()
SqlDatasetWriter(dataset, "dataset", "sqlite:///" + output_sqlite_path, num_proc=1).write()
original_sql = iter_sql_file(sqlite_path)
expected_sql = iter_sql_file(output_sqlite_path)
for row1, row2 in zip(original_sql, expected_sql):
assert row1 == row2
@require_sqlalchemy
def test_dataset_to_sql_multiproc(sqlite_path, tmp_path, set_sqlalchemy_silence_uber_warning):
cache_dir = tmp_path / "cache"
output_sqlite_path = os.path.join(cache_dir, "tmp.sql")
dataset = SqlDatasetReader("dataset", "sqlite:///" + sqlite_path, cache_dir=cache_dir).read()
SqlDatasetWriter(dataset, "dataset", "sqlite:///" + output_sqlite_path, num_proc=2).write()
original_sql = iter_sql_file(sqlite_path)
expected_sql = iter_sql_file(output_sqlite_path)
for row1, row2 in zip(original_sql, expected_sql):
assert row1 == row2
@require_sqlalchemy
def test_dataset_to_sql_invalidproc(sqlite_path, tmp_path, set_sqlalchemy_silence_uber_warning):
cache_dir = tmp_path / "cache"
output_sqlite_path = os.path.join(cache_dir, "tmp.sql")
dataset = SqlDatasetReader("dataset", "sqlite:///" + sqlite_path, cache_dir=cache_dir).read()
with pytest.raises(ValueError):
SqlDatasetWriter(dataset, "dataset", "sqlite:///" + output_sqlite_path, num_proc=0).write()
| 0
|
hf_public_repos/datasets/tests
|
hf_public_repos/datasets/tests/io/test_csv.py
|
import csv
import os
import pytest
from datasets import Dataset, DatasetDict, Features, NamedSplit, Value
from datasets.io.csv import CsvDatasetReader, CsvDatasetWriter
from ..utils import assert_arrow_memory_doesnt_increase, assert_arrow_memory_increases
def _check_csv_dataset(dataset, expected_features):
assert isinstance(dataset, Dataset)
assert dataset.num_rows == 4
assert dataset.num_columns == 3
assert dataset.column_names == ["col_1", "col_2", "col_3"]
for feature, expected_dtype in expected_features.items():
assert dataset.features[feature].dtype == expected_dtype
@pytest.mark.parametrize("keep_in_memory", [False, True])
def test_dataset_from_csv_keep_in_memory(keep_in_memory, csv_path, tmp_path):
cache_dir = tmp_path / "cache"
expected_features = {"col_1": "int64", "col_2": "int64", "col_3": "float64"}
with assert_arrow_memory_increases() if keep_in_memory else assert_arrow_memory_doesnt_increase():
dataset = CsvDatasetReader(csv_path, cache_dir=cache_dir, keep_in_memory=keep_in_memory).read()
_check_csv_dataset(dataset, expected_features)
@pytest.mark.parametrize(
"features",
[
None,
{"col_1": "string", "col_2": "int64", "col_3": "float64"},
{"col_1": "string", "col_2": "string", "col_3": "string"},
{"col_1": "int32", "col_2": "int32", "col_3": "int32"},
{"col_1": "float32", "col_2": "float32", "col_3": "float32"},
],
)
def test_dataset_from_csv_features(features, csv_path, tmp_path):
cache_dir = tmp_path / "cache"
# CSV file loses col_1 string dtype information: default now is "int64" instead of "string"
default_expected_features = {"col_1": "int64", "col_2": "int64", "col_3": "float64"}
expected_features = features.copy() if features else default_expected_features
features = (
Features({feature: Value(dtype) for feature, dtype in features.items()}) if features is not None else None
)
dataset = CsvDatasetReader(csv_path, features=features, cache_dir=cache_dir).read()
_check_csv_dataset(dataset, expected_features)
@pytest.mark.parametrize("split", [None, NamedSplit("train"), "train", "test"])
def test_dataset_from_csv_split(split, csv_path, tmp_path):
cache_dir = tmp_path / "cache"
expected_features = {"col_1": "int64", "col_2": "int64", "col_3": "float64"}
dataset = CsvDatasetReader(csv_path, cache_dir=cache_dir, split=split).read()
_check_csv_dataset(dataset, expected_features)
assert dataset.split == split if split else "train"
@pytest.mark.parametrize("path_type", [str, list])
def test_dataset_from_csv_path_type(path_type, csv_path, tmp_path):
if issubclass(path_type, str):
path = csv_path
elif issubclass(path_type, list):
path = [csv_path]
cache_dir = tmp_path / "cache"
expected_features = {"col_1": "int64", "col_2": "int64", "col_3": "float64"}
dataset = CsvDatasetReader(path, cache_dir=cache_dir).read()
_check_csv_dataset(dataset, expected_features)
def _check_csv_datasetdict(dataset_dict, expected_features, splits=("train",)):
assert isinstance(dataset_dict, DatasetDict)
for split in splits:
dataset = dataset_dict[split]
assert dataset.num_rows == 4
assert dataset.num_columns == 3
assert dataset.column_names == ["col_1", "col_2", "col_3"]
for feature, expected_dtype in expected_features.items():
assert dataset.features[feature].dtype == expected_dtype
@pytest.mark.parametrize("keep_in_memory", [False, True])
def test_csv_datasetdict_reader_keep_in_memory(keep_in_memory, csv_path, tmp_path):
cache_dir = tmp_path / "cache"
expected_features = {"col_1": "int64", "col_2": "int64", "col_3": "float64"}
with assert_arrow_memory_increases() if keep_in_memory else assert_arrow_memory_doesnt_increase():
dataset = CsvDatasetReader({"train": csv_path}, cache_dir=cache_dir, keep_in_memory=keep_in_memory).read()
_check_csv_datasetdict(dataset, expected_features)
@pytest.mark.parametrize(
"features",
[
None,
{"col_1": "string", "col_2": "int64", "col_3": "float64"},
{"col_1": "string", "col_2": "string", "col_3": "string"},
{"col_1": "int32", "col_2": "int32", "col_3": "int32"},
{"col_1": "float32", "col_2": "float32", "col_3": "float32"},
],
)
def test_csv_datasetdict_reader_features(features, csv_path, tmp_path):
cache_dir = tmp_path / "cache"
# CSV file loses col_1 string dtype information: default now is "int64" instead of "string"
default_expected_features = {"col_1": "int64", "col_2": "int64", "col_3": "float64"}
expected_features = features.copy() if features else default_expected_features
features = (
Features({feature: Value(dtype) for feature, dtype in features.items()}) if features is not None else None
)
dataset = CsvDatasetReader({"train": csv_path}, features=features, cache_dir=cache_dir).read()
_check_csv_datasetdict(dataset, expected_features)
@pytest.mark.parametrize("split", [None, NamedSplit("train"), "train", "test"])
def test_csv_datasetdict_reader_split(split, csv_path, tmp_path):
if split:
path = {split: csv_path}
else:
path = {"train": csv_path, "test": csv_path}
cache_dir = tmp_path / "cache"
expected_features = {"col_1": "int64", "col_2": "int64", "col_3": "float64"}
dataset = CsvDatasetReader(path, cache_dir=cache_dir).read()
_check_csv_datasetdict(dataset, expected_features, splits=list(path.keys()))
assert all(dataset[split].split == split for split in path.keys())
def iter_csv_file(csv_path):
with open(csv_path, encoding="utf-8") as csvfile:
yield from csv.reader(csvfile)
def test_dataset_to_csv(csv_path, tmp_path):
cache_dir = tmp_path / "cache"
output_csv = os.path.join(cache_dir, "tmp.csv")
dataset = CsvDatasetReader({"train": csv_path}, cache_dir=cache_dir).read()
CsvDatasetWriter(dataset["train"], output_csv, num_proc=1).write()
original_csv = iter_csv_file(csv_path)
expected_csv = iter_csv_file(output_csv)
for row1, row2 in zip(original_csv, expected_csv):
assert row1 == row2
def test_dataset_to_csv_multiproc(csv_path, tmp_path):
cache_dir = tmp_path / "cache"
output_csv = os.path.join(cache_dir, "tmp.csv")
dataset = CsvDatasetReader({"train": csv_path}, cache_dir=cache_dir).read()
CsvDatasetWriter(dataset["train"], output_csv, num_proc=2).write()
original_csv = iter_csv_file(csv_path)
expected_csv = iter_csv_file(output_csv)
for row1, row2 in zip(original_csv, expected_csv):
assert row1 == row2
def test_dataset_to_csv_invalidproc(csv_path, tmp_path):
cache_dir = tmp_path / "cache"
output_csv = os.path.join(cache_dir, "tmp.csv")
dataset = CsvDatasetReader({"train": csv_path}, cache_dir=cache_dir).read()
with pytest.raises(ValueError):
CsvDatasetWriter(dataset["train"], output_csv, num_proc=0)
| 0
|
hf_public_repos/datasets/tests
|
hf_public_repos/datasets/tests/io/test_text.py
|
import pytest
from datasets import Dataset, DatasetDict, Features, NamedSplit, Value
from datasets.io.text import TextDatasetReader
from ..utils import assert_arrow_memory_doesnt_increase, assert_arrow_memory_increases
def _check_text_dataset(dataset, expected_features):
assert isinstance(dataset, Dataset)
assert dataset.num_rows == 4
assert dataset.num_columns == 1
assert dataset.column_names == ["text"]
for feature, expected_dtype in expected_features.items():
assert dataset.features[feature].dtype == expected_dtype
@pytest.mark.parametrize("keep_in_memory", [False, True])
def test_dataset_from_text_keep_in_memory(keep_in_memory, text_path, tmp_path):
cache_dir = tmp_path / "cache"
expected_features = {"text": "string"}
with assert_arrow_memory_increases() if keep_in_memory else assert_arrow_memory_doesnt_increase():
dataset = TextDatasetReader(text_path, cache_dir=cache_dir, keep_in_memory=keep_in_memory).read()
_check_text_dataset(dataset, expected_features)
@pytest.mark.parametrize(
"features",
[
None,
{"text": "string"},
{"text": "int32"},
{"text": "float32"},
],
)
def test_dataset_from_text_features(features, text_path, tmp_path):
cache_dir = tmp_path / "cache"
default_expected_features = {"text": "string"}
expected_features = features.copy() if features else default_expected_features
features = (
Features({feature: Value(dtype) for feature, dtype in features.items()}) if features is not None else None
)
dataset = TextDatasetReader(text_path, features=features, cache_dir=cache_dir).read()
_check_text_dataset(dataset, expected_features)
@pytest.mark.parametrize("split", [None, NamedSplit("train"), "train", "test"])
def test_dataset_from_text_split(split, text_path, tmp_path):
cache_dir = tmp_path / "cache"
expected_features = {"text": "string"}
dataset = TextDatasetReader(text_path, cache_dir=cache_dir, split=split).read()
_check_text_dataset(dataset, expected_features)
assert dataset.split == split if split else "train"
@pytest.mark.parametrize("path_type", [str, list])
def test_dataset_from_text_path_type(path_type, text_path, tmp_path):
if issubclass(path_type, str):
path = text_path
elif issubclass(path_type, list):
path = [text_path]
cache_dir = tmp_path / "cache"
expected_features = {"text": "string"}
dataset = TextDatasetReader(path, cache_dir=cache_dir).read()
_check_text_dataset(dataset, expected_features)
def _check_text_datasetdict(dataset_dict, expected_features, splits=("train",)):
assert isinstance(dataset_dict, DatasetDict)
for split in splits:
dataset = dataset_dict[split]
assert dataset.num_rows == 4
assert dataset.num_columns == 1
assert dataset.column_names == ["text"]
for feature, expected_dtype in expected_features.items():
assert dataset.features[feature].dtype == expected_dtype
@pytest.mark.parametrize("keep_in_memory", [False, True])
def test_datasetdict_from_text_keep_in_memory(keep_in_memory, text_path, tmp_path):
cache_dir = tmp_path / "cache"
expected_features = {"text": "string"}
with assert_arrow_memory_increases() if keep_in_memory else assert_arrow_memory_doesnt_increase():
dataset = TextDatasetReader({"train": text_path}, cache_dir=cache_dir, keep_in_memory=keep_in_memory).read()
_check_text_datasetdict(dataset, expected_features)
@pytest.mark.parametrize(
"features",
[
None,
{"text": "string"},
{"text": "int32"},
{"text": "float32"},
],
)
def test_datasetdict_from_text_features(features, text_path, tmp_path):
cache_dir = tmp_path / "cache"
# CSV file loses col_1 string dtype information: default now is "int64" instead of "string"
default_expected_features = {"text": "string"}
expected_features = features.copy() if features else default_expected_features
features = (
Features({feature: Value(dtype) for feature, dtype in features.items()}) if features is not None else None
)
dataset = TextDatasetReader({"train": text_path}, features=features, cache_dir=cache_dir).read()
_check_text_datasetdict(dataset, expected_features)
@pytest.mark.parametrize("split", [None, NamedSplit("train"), "train", "test"])
def test_datasetdict_from_text_split(split, text_path, tmp_path):
if split:
path = {split: text_path}
else:
split = "train"
path = {"train": text_path, "test": text_path}
cache_dir = tmp_path / "cache"
expected_features = {"text": "string"}
dataset = TextDatasetReader(path, cache_dir=cache_dir).read()
_check_text_datasetdict(dataset, expected_features, splits=list(path.keys()))
assert all(dataset[split].split == split for split in path.keys())
| 0
|
hf_public_repos/datasets/tests
|
hf_public_repos/datasets/tests/features/test_audio.py
|
import os
import tarfile
import pyarrow as pa
import pytest
from datasets import Dataset, concatenate_datasets, load_dataset
from datasets.features import Audio, Features, Sequence, Value
from ..utils import (
require_sndfile,
)
@pytest.fixture()
def tar_wav_path(shared_datadir, tmp_path_factory):
audio_path = str(shared_datadir / "test_audio_44100.wav")
path = tmp_path_factory.mktemp("data") / "audio_data.wav.tar"
with tarfile.TarFile(path, "w") as f:
f.add(audio_path, arcname=os.path.basename(audio_path))
return path
@pytest.fixture()
def tar_mp3_path(shared_datadir, tmp_path_factory):
audio_path = str(shared_datadir / "test_audio_44100.mp3")
path = tmp_path_factory.mktemp("data") / "audio_data.mp3.tar"
with tarfile.TarFile(path, "w") as f:
f.add(audio_path, arcname=os.path.basename(audio_path))
return path
def iter_archive(archive_path):
with tarfile.open(archive_path) as tar:
for tarinfo in tar:
file_path = tarinfo.name
file_obj = tar.extractfile(tarinfo)
yield file_path, file_obj
def test_audio_instantiation():
audio = Audio()
assert audio.sampling_rate is None
assert audio.mono is True
assert audio.id is None
assert audio.dtype == "dict"
assert audio.pa_type == pa.struct({"bytes": pa.binary(), "path": pa.string()})
assert audio._type == "Audio"
def test_audio_feature_type_to_arrow():
features = Features({"audio": Audio()})
assert features.arrow_schema == pa.schema({"audio": Audio().pa_type})
features = Features({"struct_containing_an_audio": {"audio": Audio()}})
assert features.arrow_schema == pa.schema({"struct_containing_an_audio": pa.struct({"audio": Audio().pa_type})})
features = Features({"sequence_of_audios": Sequence(Audio())})
assert features.arrow_schema == pa.schema({"sequence_of_audios": pa.list_(Audio().pa_type)})
@pytest.mark.parametrize(
"build_example",
[
lambda audio_path: audio_path,
lambda audio_path: open(audio_path, "rb").read(),
lambda audio_path: {"path": audio_path},
lambda audio_path: {"path": audio_path, "bytes": None},
lambda audio_path: {"path": audio_path, "bytes": open(audio_path, "rb").read()},
lambda audio_path: {"path": None, "bytes": open(audio_path, "rb").read()},
lambda audio_path: {"bytes": open(audio_path, "rb").read()},
lambda audio_path: {"array": [0.1, 0.2, 0.3], "sampling_rate": 16_000},
],
)
def test_audio_feature_encode_example(shared_datadir, build_example):
audio_path = str(shared_datadir / "test_audio_44100.wav")
audio = Audio()
encoded_example = audio.encode_example(build_example(audio_path))
assert isinstance(encoded_example, dict)
assert encoded_example.keys() == {"bytes", "path"}
assert encoded_example["bytes"] is not None or encoded_example["path"] is not None
decoded_example = audio.decode_example(encoded_example)
assert decoded_example.keys() == {"path", "array", "sampling_rate"}
@pytest.mark.parametrize(
"build_example",
[
lambda audio_path: {"path": audio_path, "sampling_rate": 16_000},
lambda audio_path: {"path": audio_path, "bytes": None, "sampling_rate": 16_000},
lambda audio_path: {"path": audio_path, "bytes": open(audio_path, "rb").read(), "sampling_rate": 16_000},
lambda audio_path: {"array": [0.1, 0.2, 0.3], "sampling_rate": 16_000},
],
)
def test_audio_feature_encode_example_pcm(shared_datadir, build_example):
audio_path = str(shared_datadir / "test_audio_16000.pcm")
audio = Audio(sampling_rate=16_000)
encoded_example = audio.encode_example(build_example(audio_path))
assert isinstance(encoded_example, dict)
assert encoded_example.keys() == {"bytes", "path"}
assert encoded_example["bytes"] is not None or encoded_example["path"] is not None
decoded_example = audio.decode_example(encoded_example)
assert decoded_example.keys() == {"path", "array", "sampling_rate"}
@require_sndfile
def test_audio_decode_example(shared_datadir):
audio_path = str(shared_datadir / "test_audio_44100.wav")
audio = Audio()
decoded_example = audio.decode_example(audio.encode_example(audio_path))
assert decoded_example.keys() == {"path", "array", "sampling_rate"}
assert decoded_example["path"] == audio_path
assert decoded_example["array"].shape == (202311,)
assert decoded_example["sampling_rate"] == 44100
with pytest.raises(RuntimeError):
Audio(decode=False).decode_example(audio_path)
@require_sndfile
def test_audio_resampling(shared_datadir):
audio_path = str(shared_datadir / "test_audio_44100.wav")
audio = Audio(sampling_rate=16000)
decoded_example = audio.decode_example(audio.encode_example(audio_path))
assert decoded_example.keys() == {"path", "array", "sampling_rate"}
assert decoded_example["path"] == audio_path
assert decoded_example["array"].shape == (73401,)
assert decoded_example["sampling_rate"] == 16000
@require_sndfile
def test_audio_decode_example_mp3(shared_datadir):
audio_path = str(shared_datadir / "test_audio_44100.mp3")
audio = Audio()
decoded_example = audio.decode_example(audio.encode_example(audio_path))
assert decoded_example.keys() == {"path", "array", "sampling_rate"}
assert decoded_example["path"] == audio_path
assert decoded_example["array"].shape == (110592,)
assert decoded_example["sampling_rate"] == 44100
@require_sndfile
def test_audio_decode_example_opus(shared_datadir):
audio_path = str(shared_datadir / "test_audio_48000.opus")
audio = Audio()
decoded_example = audio.decode_example(audio.encode_example(audio_path))
assert decoded_example.keys() == {"path", "array", "sampling_rate"}
assert decoded_example["path"] == audio_path
assert decoded_example["array"].shape == (48000,)
assert decoded_example["sampling_rate"] == 48000
@pytest.mark.parametrize("sampling_rate", [16_000, 48_000])
def test_audio_decode_example_pcm(shared_datadir, sampling_rate):
audio_path = str(shared_datadir / "test_audio_16000.pcm")
audio_input = {"path": audio_path, "sampling_rate": 16_000}
audio = Audio(sampling_rate=sampling_rate)
decoded_example = audio.decode_example(audio.encode_example(audio_input))
assert decoded_example.keys() == {"path", "array", "sampling_rate"}
assert decoded_example["path"] is None
assert decoded_example["array"].shape == (16208 * sampling_rate // 16_000,)
assert decoded_example["sampling_rate"] == sampling_rate
@require_sndfile
def test_audio_resampling_mp3_different_sampling_rates(shared_datadir):
audio_path = str(shared_datadir / "test_audio_44100.mp3")
audio_path2 = str(shared_datadir / "test_audio_16000.mp3")
audio = Audio(sampling_rate=48000)
decoded_example = audio.decode_example(audio.encode_example(audio_path))
assert decoded_example.keys() == {"path", "array", "sampling_rate"}
assert decoded_example["path"] == audio_path
assert decoded_example["array"].shape == (120373,)
assert decoded_example["sampling_rate"] == 48000
decoded_example = audio.decode_example(audio.encode_example(audio_path2))
assert decoded_example.keys() == {"path", "array", "sampling_rate"}
assert decoded_example["path"] == audio_path2
assert decoded_example["array"].shape == (122688,)
assert decoded_example["sampling_rate"] == 48000
@require_sndfile
def test_dataset_with_audio_feature(shared_datadir):
audio_path = str(shared_datadir / "test_audio_44100.wav")
data = {"audio": [audio_path]}
features = Features({"audio": Audio()})
dset = Dataset.from_dict(data, features=features)
item = dset[0]
assert item.keys() == {"audio"}
assert item["audio"].keys() == {"path", "array", "sampling_rate"}
assert item["audio"]["path"] == audio_path
assert item["audio"]["array"].shape == (202311,)
assert item["audio"]["sampling_rate"] == 44100
batch = dset[:1]
assert batch.keys() == {"audio"}
assert len(batch["audio"]) == 1
assert batch["audio"][0].keys() == {"path", "array", "sampling_rate"}
assert batch["audio"][0]["path"] == audio_path
assert batch["audio"][0]["array"].shape == (202311,)
assert batch["audio"][0]["sampling_rate"] == 44100
column = dset["audio"]
assert len(column) == 1
assert column[0].keys() == {"path", "array", "sampling_rate"}
assert column[0]["path"] == audio_path
assert column[0]["array"].shape == (202311,)
assert column[0]["sampling_rate"] == 44100
@require_sndfile
def test_dataset_with_audio_feature_tar_wav(tar_wav_path):
audio_filename = "test_audio_44100.wav"
data = {"audio": []}
for file_path, file_obj in iter_archive(tar_wav_path):
data["audio"].append({"path": file_path, "bytes": file_obj.read()})
break
features = Features({"audio": Audio()})
dset = Dataset.from_dict(data, features=features)
item = dset[0]
assert item.keys() == {"audio"}
assert item["audio"].keys() == {"path", "array", "sampling_rate"}
assert item["audio"]["path"] == audio_filename
assert item["audio"]["array"].shape == (202311,)
assert item["audio"]["sampling_rate"] == 44100
batch = dset[:1]
assert batch.keys() == {"audio"}
assert len(batch["audio"]) == 1
assert batch["audio"][0].keys() == {"path", "array", "sampling_rate"}
assert batch["audio"][0]["path"] == audio_filename
assert batch["audio"][0]["array"].shape == (202311,)
assert batch["audio"][0]["sampling_rate"] == 44100
column = dset["audio"]
assert len(column) == 1
assert column[0].keys() == {"path", "array", "sampling_rate"}
assert column[0]["path"] == audio_filename
assert column[0]["array"].shape == (202311,)
assert column[0]["sampling_rate"] == 44100
@require_sndfile
def test_dataset_with_audio_feature_tar_mp3(tar_mp3_path):
audio_filename = "test_audio_44100.mp3"
data = {"audio": []}
for file_path, file_obj in iter_archive(tar_mp3_path):
data["audio"].append({"path": file_path, "bytes": file_obj.read()})
break
features = Features({"audio": Audio()})
dset = Dataset.from_dict(data, features=features)
item = dset[0]
assert item.keys() == {"audio"}
assert item["audio"].keys() == {"path", "array", "sampling_rate"}
assert item["audio"]["path"] == audio_filename
assert item["audio"]["array"].shape == (110592,)
assert item["audio"]["sampling_rate"] == 44100
batch = dset[:1]
assert batch.keys() == {"audio"}
assert len(batch["audio"]) == 1
assert batch["audio"][0].keys() == {"path", "array", "sampling_rate"}
assert batch["audio"][0]["path"] == audio_filename
assert batch["audio"][0]["array"].shape == (110592,)
assert batch["audio"][0]["sampling_rate"] == 44100
column = dset["audio"]
assert len(column) == 1
assert column[0].keys() == {"path", "array", "sampling_rate"}
assert column[0]["path"] == audio_filename
assert column[0]["array"].shape == (110592,)
assert column[0]["sampling_rate"] == 44100
@require_sndfile
def test_dataset_with_audio_feature_with_none():
data = {"audio": [None]}
features = Features({"audio": Audio()})
dset = Dataset.from_dict(data, features=features)
item = dset[0]
assert item.keys() == {"audio"}
assert item["audio"] is None
batch = dset[:1]
assert len(batch) == 1
assert batch.keys() == {"audio"}
assert isinstance(batch["audio"], list) and all(item is None for item in batch["audio"])
column = dset["audio"]
assert len(column) == 1
assert isinstance(column, list) and all(item is None for item in column)
# nested tests
data = {"audio": [[None]]}
features = Features({"audio": Sequence(Audio())})
dset = Dataset.from_dict(data, features=features)
item = dset[0]
assert item.keys() == {"audio"}
assert all(i is None for i in item["audio"])
data = {"nested": [{"audio": None}]}
features = Features({"nested": {"audio": Audio()}})
dset = Dataset.from_dict(data, features=features)
item = dset[0]
assert item.keys() == {"nested"}
assert item["nested"].keys() == {"audio"}
assert item["nested"]["audio"] is None
@require_sndfile
def test_resampling_at_loading_dataset_with_audio_feature(shared_datadir):
audio_path = str(shared_datadir / "test_audio_44100.wav")
data = {"audio": [audio_path]}
features = Features({"audio": Audio(sampling_rate=16000)})
dset = Dataset.from_dict(data, features=features)
item = dset[0]
assert item.keys() == {"audio"}
assert item["audio"].keys() == {"path", "array", "sampling_rate"}
assert item["audio"]["path"] == audio_path
assert item["audio"]["array"].shape == (73401,)
assert item["audio"]["sampling_rate"] == 16000
batch = dset[:1]
assert batch.keys() == {"audio"}
assert len(batch["audio"]) == 1
assert batch["audio"][0].keys() == {"path", "array", "sampling_rate"}
assert batch["audio"][0]["path"] == audio_path
assert batch["audio"][0]["array"].shape == (73401,)
assert batch["audio"][0]["sampling_rate"] == 16000
column = dset["audio"]
assert len(column) == 1
assert column[0].keys() == {"path", "array", "sampling_rate"}
assert column[0]["path"] == audio_path
assert column[0]["array"].shape == (73401,)
assert column[0]["sampling_rate"] == 16000
@require_sndfile
def test_resampling_at_loading_dataset_with_audio_feature_mp3(shared_datadir):
audio_path = str(shared_datadir / "test_audio_44100.mp3")
data = {"audio": [audio_path]}
features = Features({"audio": Audio(sampling_rate=16000)})
dset = Dataset.from_dict(data, features=features)
item = dset[0]
assert item.keys() == {"audio"}
assert item["audio"].keys() == {"path", "array", "sampling_rate"}
assert item["audio"]["path"] == audio_path
assert item["audio"]["array"].shape == (40125,)
assert item["audio"]["sampling_rate"] == 16000
batch = dset[:1]
assert batch.keys() == {"audio"}
assert len(batch["audio"]) == 1
assert batch["audio"][0].keys() == {"path", "array", "sampling_rate"}
assert batch["audio"][0]["path"] == audio_path
assert batch["audio"][0]["array"].shape == (40125,)
assert batch["audio"][0]["sampling_rate"] == 16000
column = dset["audio"]
assert len(column) == 1
assert column[0].keys() == {"path", "array", "sampling_rate"}
assert column[0]["path"] == audio_path
assert column[0]["array"].shape == (40125,)
assert column[0]["sampling_rate"] == 16000
@require_sndfile
def test_resampling_after_loading_dataset_with_audio_feature(shared_datadir):
audio_path = str(shared_datadir / "test_audio_44100.wav")
data = {"audio": [audio_path]}
features = Features({"audio": Audio()})
dset = Dataset.from_dict(data, features=features)
item = dset[0]
assert item["audio"]["sampling_rate"] == 44100
dset = dset.cast_column("audio", Audio(sampling_rate=16000))
item = dset[0]
assert item.keys() == {"audio"}
assert item["audio"].keys() == {"path", "array", "sampling_rate"}
assert item["audio"]["path"] == audio_path
assert item["audio"]["array"].shape == (73401,)
assert item["audio"]["sampling_rate"] == 16000
batch = dset[:1]
assert batch.keys() == {"audio"}
assert len(batch["audio"]) == 1
assert batch["audio"][0].keys() == {"path", "array", "sampling_rate"}
assert batch["audio"][0]["path"] == audio_path
assert batch["audio"][0]["array"].shape == (73401,)
assert batch["audio"][0]["sampling_rate"] == 16000
column = dset["audio"]
assert len(column) == 1
assert column[0].keys() == {"path", "array", "sampling_rate"}
assert column[0]["path"] == audio_path
assert column[0]["array"].shape == (73401,)
assert column[0]["sampling_rate"] == 16000
@require_sndfile
def test_resampling_after_loading_dataset_with_audio_feature_mp3(shared_datadir):
audio_path = str(shared_datadir / "test_audio_44100.mp3")
data = {"audio": [audio_path]}
features = Features({"audio": Audio()})
dset = Dataset.from_dict(data, features=features)
item = dset[0]
assert item["audio"]["sampling_rate"] == 44100
dset = dset.cast_column("audio", Audio(sampling_rate=16000))
item = dset[0]
assert item.keys() == {"audio"}
assert item["audio"].keys() == {"path", "array", "sampling_rate"}
assert item["audio"]["path"] == audio_path
assert item["audio"]["array"].shape == (40125,)
assert item["audio"]["sampling_rate"] == 16000
batch = dset[:1]
assert batch.keys() == {"audio"}
assert len(batch["audio"]) == 1
assert batch["audio"][0].keys() == {"path", "array", "sampling_rate"}
assert batch["audio"][0]["path"] == audio_path
assert batch["audio"][0]["array"].shape == (40125,)
assert batch["audio"][0]["sampling_rate"] == 16000
column = dset["audio"]
assert len(column) == 1
assert column[0].keys() == {"path", "array", "sampling_rate"}
assert column[0]["path"] == audio_path
assert column[0]["array"].shape == (40125,)
assert column[0]["sampling_rate"] == 16000
@pytest.mark.parametrize(
"build_data",
[
lambda audio_path: {"audio": [audio_path]},
lambda audio_path: {"audio": [open(audio_path, "rb").read()]},
lambda audio_path: {"audio": [{"path": audio_path}]},
lambda audio_path: {"audio": [{"path": audio_path, "bytes": None}]},
lambda audio_path: {"audio": [{"path": audio_path, "bytes": open(audio_path, "rb").read()}]},
lambda audio_path: {"audio": [{"path": None, "bytes": open(audio_path, "rb").read()}]},
lambda audio_path: {"audio": [{"bytes": open(audio_path, "rb").read()}]},
],
)
def test_dataset_cast_to_audio_features(shared_datadir, build_data):
audio_path = str(shared_datadir / "test_audio_44100.wav")
data = build_data(audio_path)
dset = Dataset.from_dict(data)
item = dset.cast(Features({"audio": Audio()}))[0]
assert item.keys() == {"audio"}
assert item["audio"].keys() == {"path", "array", "sampling_rate"}
item = dset.cast_column("audio", Audio())[0]
assert item.keys() == {"audio"}
assert item["audio"].keys() == {"path", "array", "sampling_rate"}
def test_dataset_concatenate_audio_features(shared_datadir):
# we use a different data structure between 1 and 2 to make sure they are compatible with each other
audio_path = str(shared_datadir / "test_audio_44100.wav")
data1 = {"audio": [audio_path]}
dset1 = Dataset.from_dict(data1, features=Features({"audio": Audio()}))
data2 = {"audio": [{"bytes": open(audio_path, "rb").read()}]}
dset2 = Dataset.from_dict(data2, features=Features({"audio": Audio()}))
concatenated_dataset = concatenate_datasets([dset1, dset2])
assert len(concatenated_dataset) == len(dset1) + len(dset2)
assert concatenated_dataset[0]["audio"]["array"].shape == dset1[0]["audio"]["array"].shape
assert concatenated_dataset[1]["audio"]["array"].shape == dset2[0]["audio"]["array"].shape
def test_dataset_concatenate_nested_audio_features(shared_datadir):
# we use a different data structure between 1 and 2 to make sure they are compatible with each other
audio_path = str(shared_datadir / "test_audio_44100.wav")
features = Features({"list_of_structs_of_audios": [{"audio": Audio()}]})
data1 = {"list_of_structs_of_audios": [[{"audio": audio_path}]]}
dset1 = Dataset.from_dict(data1, features=features)
data2 = {"list_of_structs_of_audios": [[{"audio": {"bytes": open(audio_path, "rb").read()}}]]}
dset2 = Dataset.from_dict(data2, features=features)
concatenated_dataset = concatenate_datasets([dset1, dset2])
assert len(concatenated_dataset) == len(dset1) + len(dset2)
assert (
concatenated_dataset[0]["list_of_structs_of_audios"][0]["audio"]["array"].shape
== dset1[0]["list_of_structs_of_audios"][0]["audio"]["array"].shape
)
assert (
concatenated_dataset[1]["list_of_structs_of_audios"][0]["audio"]["array"].shape
== dset2[0]["list_of_structs_of_audios"][0]["audio"]["array"].shape
)
@require_sndfile
def test_dataset_with_audio_feature_map_is_not_decoded(shared_datadir):
audio_path = str(shared_datadir / "test_audio_44100.wav")
data = {"audio": [audio_path], "text": ["Hello"]}
features = Features({"audio": Audio(), "text": Value("string")})
dset = Dataset.from_dict(data, features=features)
expected_audio = features.encode_batch(data)["audio"][0]
for item in dset.cast_column("audio", Audio(decode=False)):
assert item.keys() == {"audio", "text"}
assert item == {"audio": expected_audio, "text": "Hello"}
def process_text(example):
example["text"] = example["text"] + " World!"
return example
processed_dset = dset.map(process_text)
for item in processed_dset.cast_column("audio", Audio(decode=False)):
assert item.keys() == {"audio", "text"}
assert item == {"audio": expected_audio, "text": "Hello World!"}
@require_sndfile
def test_dataset_with_audio_feature_map_is_decoded(shared_datadir):
audio_path = str(shared_datadir / "test_audio_44100.wav")
data = {"audio": [audio_path], "text": ["Hello"]}
features = Features({"audio": Audio(), "text": Value("string")})
dset = Dataset.from_dict(data, features=features)
def process_audio_sampling_rate_by_example(example):
example["double_sampling_rate"] = 2 * example["audio"]["sampling_rate"]
return example
decoded_dset = dset.map(process_audio_sampling_rate_by_example)
for item in decoded_dset.cast_column("audio", Audio(decode=False)):
assert item.keys() == {"audio", "text", "double_sampling_rate"}
assert item["double_sampling_rate"] == 88200
def process_audio_sampling_rate_by_batch(batch):
double_sampling_rates = []
for audio in batch["audio"]:
double_sampling_rates.append(2 * audio["sampling_rate"])
batch["double_sampling_rate"] = double_sampling_rates
return batch
decoded_dset = dset.map(process_audio_sampling_rate_by_batch, batched=True)
for item in decoded_dset.cast_column("audio", Audio(decode=False)):
assert item.keys() == {"audio", "text", "double_sampling_rate"}
assert item["double_sampling_rate"] == 88200
@require_sndfile
def test_formatted_dataset_with_audio_feature(shared_datadir):
audio_path = str(shared_datadir / "test_audio_44100.wav")
data = {"audio": [audio_path, audio_path]}
features = Features({"audio": Audio()})
dset = Dataset.from_dict(data, features=features)
with dset.formatted_as("numpy"):
item = dset[0]
assert item.keys() == {"audio"}
assert item["audio"].keys() == {"path", "array", "sampling_rate"}
assert item["audio"]["path"] == audio_path
assert item["audio"]["array"].shape == (202311,)
assert item["audio"]["sampling_rate"] == 44100
batch = dset[:1]
assert batch.keys() == {"audio"}
assert len(batch["audio"]) == 1
assert batch["audio"][0].keys() == {"path", "array", "sampling_rate"}
assert batch["audio"][0]["path"] == audio_path
assert batch["audio"][0]["array"].shape == (202311,)
assert batch["audio"][0]["sampling_rate"] == 44100
column = dset["audio"]
assert len(column) == 2
assert column[0].keys() == {"path", "array", "sampling_rate"}
assert column[0]["path"] == audio_path
assert column[0]["array"].shape == (202311,)
assert column[0]["sampling_rate"] == 44100
with dset.formatted_as("pandas"):
item = dset[0]
assert item.shape == (1, 1)
assert item.columns == ["audio"]
assert item["audio"][0].keys() == {"path", "array", "sampling_rate"}
assert item["audio"][0]["path"] == audio_path
assert item["audio"][0]["array"].shape == (202311,)
assert item["audio"][0]["sampling_rate"] == 44100
batch = dset[:1]
assert batch.shape == (1, 1)
assert batch.columns == ["audio"]
assert batch["audio"][0].keys() == {"path", "array", "sampling_rate"}
assert batch["audio"][0]["path"] == audio_path
assert batch["audio"][0]["array"].shape == (202311,)
assert batch["audio"][0]["sampling_rate"] == 44100
column = dset["audio"]
assert len(column) == 2
assert column[0].keys() == {"path", "array", "sampling_rate"}
assert column[0]["path"] == audio_path
assert column[0]["array"].shape == (202311,)
assert column[0]["sampling_rate"] == 44100
@pytest.fixture
def jsonl_audio_dataset_path(shared_datadir, tmp_path_factory):
import json
audio_path = str(shared_datadir / "test_audio_44100.wav")
data = [{"audio": audio_path, "text": "Hello world!"}]
path = str(tmp_path_factory.mktemp("data") / "audio_dataset.jsonl")
with open(path, "w") as f:
for item in data:
f.write(json.dumps(item) + "\n")
return path
@require_sndfile
@pytest.mark.parametrize("streaming", [False, True])
def test_load_dataset_with_audio_feature(streaming, jsonl_audio_dataset_path, shared_datadir):
audio_path = str(shared_datadir / "test_audio_44100.wav")
data_files = jsonl_audio_dataset_path
features = Features({"audio": Audio(), "text": Value("string")})
dset = load_dataset("json", split="train", data_files=data_files, features=features, streaming=streaming)
item = dset[0] if not streaming else next(iter(dset))
assert item.keys() == {"audio", "text"}
assert item["audio"].keys() == {"path", "array", "sampling_rate"}
assert item["audio"]["path"] == audio_path
assert item["audio"]["array"].shape == (202311,)
assert item["audio"]["sampling_rate"] == 44100
@require_sndfile
@pytest.mark.integration
def test_dataset_with_audio_feature_loaded_from_cache():
# load first time
ds = load_dataset("patrickvonplaten/librispeech_asr_dummy", "clean")
# load from cache
ds = load_dataset("patrickvonplaten/librispeech_asr_dummy", "clean", split="validation")
assert isinstance(ds, Dataset)
def test_dataset_with_audio_feature_undecoded(shared_datadir):
audio_path = str(shared_datadir / "test_audio_44100.wav")
data = {"audio": [audio_path]}
features = Features({"audio": Audio(decode=False)})
dset = Dataset.from_dict(data, features=features)
item = dset[0]
assert item.keys() == {"audio"}
assert item["audio"] == {"path": audio_path, "bytes": None}
batch = dset[:1]
assert batch.keys() == {"audio"}
assert len(batch["audio"]) == 1
assert batch["audio"][0] == {"path": audio_path, "bytes": None}
column = dset["audio"]
assert len(column) == 1
assert column[0] == {"path": audio_path, "bytes": None}
def test_formatted_dataset_with_audio_feature_undecoded(shared_datadir):
audio_path = str(shared_datadir / "test_audio_44100.wav")
data = {"audio": [audio_path]}
features = Features({"audio": Audio(decode=False)})
dset = Dataset.from_dict(data, features=features)
with dset.formatted_as("numpy"):
item = dset[0]
assert item.keys() == {"audio"}
assert item["audio"] == {"path": audio_path, "bytes": None}
batch = dset[:1]
assert batch.keys() == {"audio"}
assert len(batch["audio"]) == 1
assert batch["audio"][0] == {"path": audio_path, "bytes": None}
column = dset["audio"]
assert len(column) == 1
assert column[0] == {"path": audio_path, "bytes": None}
with dset.formatted_as("pandas"):
item = dset[0]
assert item.shape == (1, 1)
assert item.columns == ["audio"]
assert item["audio"][0] == {"path": audio_path, "bytes": None}
batch = dset[:1]
assert batch.shape == (1, 1)
assert batch.columns == ["audio"]
assert batch["audio"][0] == {"path": audio_path, "bytes": None}
column = dset["audio"]
assert len(column) == 1
assert column[0] == {"path": audio_path, "bytes": None}
def test_dataset_with_audio_feature_map_undecoded(shared_datadir):
audio_path = str(shared_datadir / "test_audio_44100.wav")
data = {"audio": [audio_path]}
features = Features({"audio": Audio(decode=False)})
dset = Dataset.from_dict(data, features=features)
def assert_audio_example_undecoded(example):
assert example["audio"] == {"path": audio_path, "bytes": None}
dset.map(assert_audio_example_undecoded)
def assert_audio_batch_undecoded(batch):
for audio in batch["audio"]:
assert audio == {"path": audio_path, "bytes": None}
dset.map(assert_audio_batch_undecoded, batched=True)
def test_audio_embed_storage(shared_datadir):
audio_path = str(shared_datadir / "test_audio_44100.wav")
example = {"bytes": None, "path": audio_path}
storage = pa.array([example], type=pa.struct({"bytes": pa.binary(), "path": pa.string()}))
embedded_storage = Audio().embed_storage(storage)
embedded_example = embedded_storage.to_pylist()[0]
assert embedded_example == {"bytes": open(audio_path, "rb").read(), "path": "test_audio_44100.wav"}
| 0
|
hf_public_repos/datasets/tests
|
hf_public_repos/datasets/tests/features/test_array_xd.py
|
import os
import random
import tempfile
import unittest
import numpy as np
import pandas as pd
import pyarrow as pa
import pytest
from absl.testing import parameterized
import datasets
from datasets.arrow_writer import ArrowWriter
from datasets.features import Array2D, Array3D, Array4D, Array5D, Value
from datasets.features.features import Array3DExtensionType, PandasArrayExtensionDtype, _ArrayXD
from datasets.formatting.formatting import NumpyArrowExtractor, SimpleArrowExtractor
SHAPE_TEST_1 = (30, 487)
SHAPE_TEST_2 = (36, 1024)
SHAPE_TEST_3 = (None, 100)
SPEED_TEST_SHAPE = (100, 100)
SPEED_TEST_N_EXAMPLES = 100
DEFAULT_FEATURES = datasets.Features(
{
"text": Array2D(SHAPE_TEST_1, dtype="float32"),
"image": Array2D(SHAPE_TEST_2, dtype="float32"),
"dynamic": Array2D(SHAPE_TEST_3, dtype="float32"),
}
)
def generate_examples(features: dict, num_examples=100, seq_shapes=None):
dummy_data = []
seq_shapes = seq_shapes or {}
for i in range(num_examples):
example = {}
for col_id, (k, v) in enumerate(features.items()):
if isinstance(v, _ArrayXD):
if k == "dynamic":
first_dim = random.randint(1, 3)
data = np.random.rand(first_dim, *v.shape[1:]).astype(v.dtype)
else:
data = np.random.rand(*v.shape).astype(v.dtype)
elif isinstance(v, datasets.Value):
data = "foo"
elif isinstance(v, datasets.Sequence):
while isinstance(v, datasets.Sequence):
v = v.feature
shape = seq_shapes[k]
data = np.random.rand(*shape).astype(v.dtype)
example[k] = data
dummy_data.append((i, example))
return dummy_data
class ExtensionTypeCompatibilityTest(unittest.TestCase):
def test_array2d_nonspecific_shape(self):
with tempfile.TemporaryDirectory() as tmp_dir:
my_features = DEFAULT_FEATURES.copy()
with ArrowWriter(features=my_features, path=os.path.join(tmp_dir, "beta.arrow")) as writer:
for key, record in generate_examples(
features=my_features,
num_examples=1,
):
example = my_features.encode_example(record)
writer.write(example)
num_examples, num_bytes = writer.finalize()
dataset = datasets.Dataset.from_file(os.path.join(tmp_dir, "beta.arrow"))
dataset.set_format("numpy")
row = dataset[0]
first_shape = row["image"].shape
second_shape = row["text"].shape
self.assertTrue(first_shape is not None and second_shape is not None, "need atleast 2 different shapes")
self.assertEqual(len(first_shape), len(second_shape), "both shapes are supposed to be equal length")
self.assertNotEqual(first_shape, second_shape, "shapes must not be the same")
del dataset
def test_multiple_extensions_same_row(self):
with tempfile.TemporaryDirectory() as tmp_dir:
my_features = DEFAULT_FEATURES.copy()
with ArrowWriter(features=my_features, path=os.path.join(tmp_dir, "beta.arrow")) as writer:
for key, record in generate_examples(features=my_features, num_examples=1):
example = my_features.encode_example(record)
writer.write(example)
num_examples, num_bytes = writer.finalize()
dataset = datasets.Dataset.from_file(os.path.join(tmp_dir, "beta.arrow"))
dataset.set_format("numpy")
row = dataset[0]
first_len = len(row["image"].shape)
second_len = len(row["text"].shape)
third_len = len(row["dynamic"].shape)
self.assertEqual(first_len, 2, "use a sequence type if dim is < 2")
self.assertEqual(second_len, 2, "use a sequence type if dim is < 2")
self.assertEqual(third_len, 2, "use a sequence type if dim is < 2")
del dataset
def test_compatability_with_string_values(self):
with tempfile.TemporaryDirectory() as tmp_dir:
my_features = DEFAULT_FEATURES.copy()
my_features["image_id"] = datasets.Value("string")
with ArrowWriter(features=my_features, path=os.path.join(tmp_dir, "beta.arrow")) as writer:
for key, record in generate_examples(features=my_features, num_examples=1):
example = my_features.encode_example(record)
writer.write(example)
num_examples, num_bytes = writer.finalize()
dataset = datasets.Dataset.from_file(os.path.join(tmp_dir, "beta.arrow"))
self.assertIsInstance(dataset[0]["image_id"], str, "image id must be of type string")
del dataset
def test_extension_indexing(self):
with tempfile.TemporaryDirectory() as tmp_dir:
my_features = DEFAULT_FEATURES.copy()
my_features["explicit_ext"] = Array2D((3, 3), dtype="float32")
with ArrowWriter(features=my_features, path=os.path.join(tmp_dir, "beta.arrow")) as writer:
for key, record in generate_examples(features=my_features, num_examples=1):
example = my_features.encode_example(record)
writer.write(example)
num_examples, num_bytes = writer.finalize()
dataset = datasets.Dataset.from_file(os.path.join(tmp_dir, "beta.arrow"))
dataset.set_format("numpy")
data = dataset[0]["explicit_ext"]
self.assertIsInstance(data, np.ndarray, "indexed extension must return numpy.ndarray")
del dataset
def get_array_feature_types():
shape_1 = [3] * 5
shape_2 = [3, 4, 5, 6, 7]
return [
{
"testcase_name": f"{d}d",
"array_feature": array_feature,
"shape_1": tuple(shape_1[:d]),
"shape_2": tuple(shape_2[:d]),
}
for d, array_feature in zip(range(2, 6), [Array2D, Array3D, Array4D, Array5D])
]
@parameterized.named_parameters(get_array_feature_types())
class ArrayXDTest(unittest.TestCase):
def get_features(self, array_feature, shape_1, shape_2):
return datasets.Features(
{
"image": array_feature(shape_1, dtype="float32"),
"source": Value("string"),
"matrix": array_feature(shape_2, dtype="float32"),
}
)
def get_dict_example_0(self, shape_1, shape_2):
return {
"image": np.random.rand(*shape_1).astype("float32"),
"source": "foo",
"matrix": np.random.rand(*shape_2).astype("float32"),
}
def get_dict_example_1(self, shape_1, shape_2):
return {
"image": np.random.rand(*shape_1).astype("float32"),
"matrix": np.random.rand(*shape_2).astype("float32"),
"source": "bar",
}
def get_dict_examples(self, shape_1, shape_2):
return {
"image": np.random.rand(2, *shape_1).astype("float32").tolist(),
"source": ["foo", "bar"],
"matrix": np.random.rand(2, *shape_2).astype("float32").tolist(),
}
def _check_getitem_output_type(self, dataset, shape_1, shape_2, first_matrix):
matrix_column = dataset["matrix"]
self.assertIsInstance(matrix_column, list)
self.assertIsInstance(matrix_column[0], list)
self.assertIsInstance(matrix_column[0][0], list)
self.assertTupleEqual(np.array(matrix_column).shape, (2, *shape_2))
matrix_field_of_first_example = dataset[0]["matrix"]
self.assertIsInstance(matrix_field_of_first_example, list)
self.assertIsInstance(matrix_field_of_first_example, list)
self.assertEqual(np.array(matrix_field_of_first_example).shape, shape_2)
np.testing.assert_array_equal(np.array(matrix_field_of_first_example), np.array(first_matrix))
matrix_field_of_first_two_examples = dataset[:2]["matrix"]
self.assertIsInstance(matrix_field_of_first_two_examples, list)
self.assertIsInstance(matrix_field_of_first_two_examples[0], list)
self.assertIsInstance(matrix_field_of_first_two_examples[0][0], list)
self.assertTupleEqual(np.array(matrix_field_of_first_two_examples).shape, (2, *shape_2))
with dataset.formatted_as("numpy"):
self.assertTupleEqual(dataset["matrix"].shape, (2, *shape_2))
self.assertEqual(dataset[0]["matrix"].shape, shape_2)
self.assertTupleEqual(dataset[:2]["matrix"].shape, (2, *shape_2))
with dataset.formatted_as("pandas"):
self.assertIsInstance(dataset["matrix"], pd.Series)
self.assertIsInstance(dataset[0]["matrix"], pd.Series)
self.assertIsInstance(dataset[:2]["matrix"], pd.Series)
self.assertTupleEqual(dataset["matrix"].to_numpy().shape, (2, *shape_2))
self.assertTupleEqual(dataset[0]["matrix"].to_numpy().shape, (1, *shape_2))
self.assertTupleEqual(dataset[:2]["matrix"].to_numpy().shape, (2, *shape_2))
def test_write(self, array_feature, shape_1, shape_2):
with tempfile.TemporaryDirectory() as tmp_dir:
my_features = self.get_features(array_feature, shape_1, shape_2)
my_examples = [
(0, self.get_dict_example_0(shape_1, shape_2)),
(1, self.get_dict_example_1(shape_1, shape_2)),
]
with ArrowWriter(features=my_features, path=os.path.join(tmp_dir, "beta.arrow")) as writer:
for key, record in my_examples:
example = my_features.encode_example(record)
writer.write(example)
num_examples, num_bytes = writer.finalize()
dataset = datasets.Dataset.from_file(os.path.join(tmp_dir, "beta.arrow"))
self._check_getitem_output_type(dataset, shape_1, shape_2, my_examples[0][1]["matrix"])
del dataset
def test_write_batch(self, array_feature, shape_1, shape_2):
with tempfile.TemporaryDirectory() as tmp_dir:
my_features = self.get_features(array_feature, shape_1, shape_2)
dict_examples = self.get_dict_examples(shape_1, shape_2)
dict_examples = my_features.encode_batch(dict_examples)
with ArrowWriter(features=my_features, path=os.path.join(tmp_dir, "beta.arrow")) as writer:
writer.write_batch(dict_examples)
num_examples, num_bytes = writer.finalize()
dataset = datasets.Dataset.from_file(os.path.join(tmp_dir, "beta.arrow"))
self._check_getitem_output_type(dataset, shape_1, shape_2, dict_examples["matrix"][0])
del dataset
def test_from_dict(self, array_feature, shape_1, shape_2):
dict_examples = self.get_dict_examples(shape_1, shape_2)
dataset = datasets.Dataset.from_dict(
dict_examples, features=self.get_features(array_feature, shape_1, shape_2)
)
self._check_getitem_output_type(dataset, shape_1, shape_2, dict_examples["matrix"][0])
del dataset
class ArrayXDDynamicTest(unittest.TestCase):
def get_one_col_dataset(self, first_dim_list, fixed_shape):
features = datasets.Features({"image": Array3D(shape=(None, *fixed_shape), dtype="float32")})
dict_values = {"image": [np.random.rand(fdim, *fixed_shape).astype("float32") for fdim in first_dim_list]}
dataset = datasets.Dataset.from_dict(dict_values, features=features)
return dataset
def get_two_col_datasset(self, first_dim_list, fixed_shape):
features = datasets.Features(
{"image": Array3D(shape=(None, *fixed_shape), dtype="float32"), "text": Value("string")}
)
dict_values = {
"image": [np.random.rand(fdim, *fixed_shape).astype("float32") for fdim in first_dim_list],
"text": ["text" for _ in first_dim_list],
}
dataset = datasets.Dataset.from_dict(dict_values, features=features)
return dataset
def test_to_pylist(self):
fixed_shape = (2, 2)
first_dim_list = [1, 3, 10]
dataset = self.get_one_col_dataset(first_dim_list, fixed_shape)
arr_xd = SimpleArrowExtractor().extract_column(dataset._data)
self.assertIsInstance(arr_xd.type, Array3DExtensionType)
pylist = arr_xd.to_pylist()
for first_dim, single_arr in zip(first_dim_list, pylist):
self.assertIsInstance(single_arr, list)
self.assertTupleEqual(np.array(single_arr).shape, (first_dim, *fixed_shape))
def test_to_numpy(self):
fixed_shape = (2, 2)
# ragged
first_dim_list = [1, 3, 10]
dataset = self.get_one_col_dataset(first_dim_list, fixed_shape)
arr_xd = SimpleArrowExtractor().extract_column(dataset._data)
self.assertIsInstance(arr_xd.type, Array3DExtensionType)
# replace with arr_xd = arr_xd.combine_chunks() when 12.0.0 will be the minimal required PyArrow version
arr_xd = arr_xd.type.wrap_array(pa.concat_arrays([chunk.storage for chunk in arr_xd.chunks]))
numpy_arr = arr_xd.to_numpy()
self.assertIsInstance(numpy_arr, np.ndarray)
self.assertEqual(numpy_arr.dtype, object)
for first_dim, single_arr in zip(first_dim_list, numpy_arr):
self.assertIsInstance(single_arr, np.ndarray)
self.assertTupleEqual(single_arr.shape, (first_dim, *fixed_shape))
# non-ragged
first_dim_list = [4, 4, 4]
dataset = self.get_one_col_dataset(first_dim_list, fixed_shape)
arr_xd = SimpleArrowExtractor().extract_column(dataset._data)
self.assertIsInstance(arr_xd.type, Array3DExtensionType)
# replace with arr_xd = arr_xd.combine_chunks() when 12.0.0 will be the minimal required PyArrow version
arr_xd = arr_xd.type.wrap_array(pa.concat_arrays([chunk.storage for chunk in arr_xd.chunks]))
numpy_arr = arr_xd.to_numpy()
self.assertIsInstance(numpy_arr, np.ndarray)
self.assertNotEqual(numpy_arr.dtype, object)
for first_dim, single_arr in zip(first_dim_list, numpy_arr):
self.assertIsInstance(single_arr, np.ndarray)
self.assertTupleEqual(single_arr.shape, (first_dim, *fixed_shape))
def test_iter_dataset(self):
fixed_shape = (2, 2)
first_dim_list = [1, 3, 10]
dataset = self.get_one_col_dataset(first_dim_list, fixed_shape)
for first_dim, ds_row in zip(first_dim_list, dataset):
single_arr = ds_row["image"]
self.assertIsInstance(single_arr, list)
self.assertTupleEqual(np.array(single_arr).shape, (first_dim, *fixed_shape))
def test_to_pandas(self):
fixed_shape = (2, 2)
# ragged
first_dim_list = [1, 3, 10]
dataset = self.get_one_col_dataset(first_dim_list, fixed_shape)
df = dataset.to_pandas()
self.assertEqual(type(df.image.dtype), PandasArrayExtensionDtype)
numpy_arr = df.image.to_numpy()
self.assertIsInstance(numpy_arr, np.ndarray)
self.assertEqual(numpy_arr.dtype, object)
for first_dim, single_arr in zip(first_dim_list, numpy_arr):
self.assertIsInstance(single_arr, np.ndarray)
self.assertTupleEqual(single_arr.shape, (first_dim, *fixed_shape))
# non-ragged
first_dim_list = [4, 4, 4]
dataset = self.get_one_col_dataset(first_dim_list, fixed_shape)
df = dataset.to_pandas()
self.assertEqual(type(df.image.dtype), PandasArrayExtensionDtype)
numpy_arr = df.image.to_numpy()
self.assertIsInstance(numpy_arr, np.ndarray)
self.assertNotEqual(numpy_arr.dtype, object)
for first_dim, single_arr in zip(first_dim_list, numpy_arr):
self.assertIsInstance(single_arr, np.ndarray)
self.assertTupleEqual(single_arr.shape, (first_dim, *fixed_shape))
def test_map_dataset(self):
fixed_shape = (2, 2)
first_dim_list = [1, 3, 10]
dataset = self.get_one_col_dataset(first_dim_list, fixed_shape)
dataset = dataset.map(lambda a: {"image": np.concatenate([a] * 2)}, input_columns="image")
# check also if above function resulted with 2x bigger first dim
for first_dim, ds_row in zip(first_dim_list, dataset):
single_arr = ds_row["image"]
self.assertIsInstance(single_arr, list)
self.assertTupleEqual(np.array(single_arr).shape, (first_dim * 2, *fixed_shape))
@pytest.mark.parametrize("dtype, dummy_value", [("int32", 1), ("bool", True), ("float64", 1)])
def test_table_to_pandas(dtype, dummy_value):
features = datasets.Features({"foo": datasets.Array2D(dtype=dtype, shape=(2, 2))})
dataset = datasets.Dataset.from_dict({"foo": [[[dummy_value] * 2] * 2]}, features=features)
df = dataset._data.to_pandas()
assert type(df.foo.dtype) == PandasArrayExtensionDtype
arr = df.foo.to_numpy()
np.testing.assert_equal(arr, np.array([[[dummy_value] * 2] * 2], dtype=np.dtype(dtype)))
@pytest.mark.parametrize("dtype, dummy_value", [("int32", 1), ("bool", True), ("float64", 1)])
def test_array_xd_numpy_arrow_extractor(dtype, dummy_value):
features = datasets.Features({"foo": datasets.Array2D(dtype=dtype, shape=(2, 2))})
dataset = datasets.Dataset.from_dict({"foo": [[[dummy_value] * 2] * 2]}, features=features)
arr = NumpyArrowExtractor().extract_column(dataset._data)
assert isinstance(arr, np.ndarray)
np.testing.assert_equal(arr, np.array([[[dummy_value] * 2] * 2], dtype=np.dtype(dtype)))
def test_array_xd_with_none():
# Fixed shape
features = datasets.Features({"foo": datasets.Array2D(dtype="int32", shape=(2, 2))})
dummy_array = np.array([[1, 2], [3, 4]], dtype="int32")
dataset = datasets.Dataset.from_dict({"foo": [dummy_array, None, dummy_array, None]}, features=features)
arr = NumpyArrowExtractor().extract_column(dataset._data)
assert isinstance(arr, np.ndarray) and arr.dtype == np.float64 and arr.shape == (4, 2, 2)
assert np.allclose(arr[0], dummy_array) and np.allclose(arr[2], dummy_array)
assert np.all(np.isnan(arr[1])) and np.all(np.isnan(arr[3])) # broadcasted np.nan - use np.all
# Dynamic shape
features = datasets.Features({"foo": datasets.Array2D(dtype="int32", shape=(None, 2))})
dummy_array = np.array([[1, 2], [3, 4]], dtype="int32")
dataset = datasets.Dataset.from_dict({"foo": [dummy_array, None, dummy_array, None]}, features=features)
arr = NumpyArrowExtractor().extract_column(dataset._data)
assert isinstance(arr, np.ndarray) and arr.dtype == object and arr.shape == (4,)
np.testing.assert_equal(arr[0], dummy_array)
np.testing.assert_equal(arr[2], dummy_array)
assert np.isnan(arr[1]) and np.isnan(arr[3]) # a single np.nan value - np.all not needed
@pytest.mark.parametrize("seq_type", ["no_sequence", "sequence", "sequence_of_sequence"])
@pytest.mark.parametrize(
"dtype",
[
"bool",
"int8",
"int16",
"int32",
"int64",
"uint8",
"uint16",
"uint32",
"uint64",
"float16",
"float32",
"float64",
],
)
@pytest.mark.parametrize("shape, feature_class", [((2, 3), datasets.Array2D), ((2, 3, 4), datasets.Array3D)])
def test_array_xd_with_np(seq_type, dtype, shape, feature_class):
feature = feature_class(dtype=dtype, shape=shape)
data = np.zeros(shape, dtype=dtype)
expected = data.tolist()
if seq_type == "sequence":
feature = datasets.Sequence(feature)
data = [data]
expected = [expected]
elif seq_type == "sequence_of_sequence":
feature = datasets.Sequence(datasets.Sequence(feature))
data = [[data]]
expected = [[expected]]
ds = datasets.Dataset.from_dict({"col": [data]}, features=datasets.Features({"col": feature}))
assert ds[0]["col"] == expected
@pytest.mark.parametrize("with_none", [False, True])
def test_dataset_map(with_none):
ds = datasets.Dataset.from_dict({"path": ["path1", "path2"]})
def process_data(batch):
batch = {
"image": [
np.array(
[
[[1, 2, 3], [4, 5, 6], [7, 8, 9]],
[[10, 20, 30], [40, 50, 60], [70, 80, 90]],
[[100, 200, 300], [400, 500, 600], [700, 800, 900]],
]
)
for _ in batch["path"]
]
}
if with_none:
batch["image"][0] = None
return batch
features = datasets.Features({"image": Array3D(dtype="int32", shape=(3, 3, 3))})
processed_ds = ds.map(process_data, batched=True, remove_columns=ds.column_names, features=features)
assert processed_ds.shape == (2, 1)
with processed_ds.with_format("numpy") as pds:
for i, example in enumerate(pds):
assert "image" in example
assert isinstance(example["image"], np.ndarray)
assert example["image"].shape == (3, 3, 3)
if with_none and i == 0:
assert np.all(np.isnan(example["image"]))
| 0
|
hf_public_repos/datasets/tests
|
hf_public_repos/datasets/tests/features/test_features.py
|
import datetime
from unittest import TestCase
from unittest.mock import patch
import numpy as np
import pandas as pd
import pyarrow as pa
import pytest
from datasets import Array2D
from datasets.arrow_dataset import Dataset
from datasets.features import Audio, ClassLabel, Features, Image, Sequence, Value
from datasets.features.features import (
_arrow_to_datasets_dtype,
_cast_to_python_objects,
cast_to_python_objects,
encode_nested_example,
generate_from_dict,
string_to_arrow,
)
from datasets.features.translation import Translation, TranslationVariableLanguages
from datasets.info import DatasetInfo
from datasets.utils.py_utils import asdict
from ..utils import require_jax, require_tf, require_torch
class FeaturesTest(TestCase):
def test_from_arrow_schema_simple(self):
data = {"a": [{"b": {"c": "text"}}] * 10, "foo": [1] * 10}
original_features = Features({"a": {"b": {"c": Value("string")}}, "foo": Value("int64")})
dset = Dataset.from_dict(data, features=original_features)
new_features = dset.features
new_dset = Dataset.from_dict(data, features=new_features)
self.assertEqual(original_features.type, new_features.type)
self.assertDictEqual(dset[0], new_dset[0])
self.assertDictEqual(dset[:], new_dset[:])
def test_from_arrow_schema_with_sequence(self):
data = {"a": [{"b": {"c": ["text"]}}] * 10, "foo": [1] * 10}
original_features = Features({"a": {"b": Sequence({"c": Value("string")})}, "foo": Value("int64")})
dset = Dataset.from_dict(data, features=original_features)
new_features = dset.features
new_dset = Dataset.from_dict(data, features=new_features)
self.assertEqual(original_features.type, new_features.type)
self.assertDictEqual(dset[0], new_dset[0])
self.assertDictEqual(dset[:], new_dset[:])
def test_string_to_arrow_bijection_for_primitive_types(self):
supported_pyarrow_datatypes = [
pa.time32("s"),
pa.time64("us"),
pa.timestamp("s"),
pa.timestamp("ns", tz="America/New_York"),
pa.date32(),
pa.date64(),
pa.duration("s"),
pa.decimal128(10, 2),
pa.decimal256(40, -3),
pa.string(),
pa.int32(),
pa.float64(),
pa.array([datetime.time(1, 1, 1)]).type, # arrow type: DataType(time64[us])
]
for dt in supported_pyarrow_datatypes:
self.assertEqual(dt, string_to_arrow(_arrow_to_datasets_dtype(dt)))
unsupported_pyarrow_datatypes = [pa.list_(pa.float64())]
for dt in unsupported_pyarrow_datatypes:
with self.assertRaises(ValueError):
string_to_arrow(_arrow_to_datasets_dtype(dt))
supported_datasets_dtypes = [
"time32[s]",
"timestamp[ns]",
"timestamp[ns, tz=+07:30]",
"duration[us]",
"decimal128(30, -4)",
"int32",
"float64",
]
for sdt in supported_datasets_dtypes:
self.assertEqual(sdt, _arrow_to_datasets_dtype(string_to_arrow(sdt)))
unsupported_datasets_dtypes = [
"time32[ns]",
"timestamp[blob]",
"timestamp[[ns]]",
"timestamp[ns, tz=[ns]]",
"duration[[us]]",
"decimal20(30, -4)",
"int",
]
for sdt in unsupported_datasets_dtypes:
with self.assertRaises(ValueError):
string_to_arrow(sdt)
def test_feature_named_type(self):
"""reference: issue #1110"""
features = Features({"_type": Value("string")})
ds_info = DatasetInfo(features=features)
reloaded_features = Features.from_dict(asdict(ds_info)["features"])
assert features == reloaded_features
def test_feature_named_self_as_kwarg(self):
"""reference: issue #5641"""
features = Features(self=Value("string"))
ds_info = DatasetInfo(features=features)
reloaded_features = Features.from_dict(asdict(ds_info)["features"])
assert features == reloaded_features
def test_class_label_feature_with_no_labels(self):
"""reference: issue #4681"""
features = Features({"label": ClassLabel(names=[])})
ds_info = DatasetInfo(features=features)
reloaded_features = Features.from_dict(asdict(ds_info)["features"])
assert features == reloaded_features
def test_reorder_fields_as(self):
features = Features(
{
"id": Value("string"),
"document": {
"title": Value("string"),
"url": Value("string"),
"html": Value("string"),
"tokens": Sequence({"token": Value("string"), "is_html": Value("bool")}),
},
"question": {
"text": Value("string"),
"tokens": Sequence(Value("string")),
},
"annotations": Sequence(
{
"id": Value("string"),
"long_answer": {
"start_token": Value("int64"),
"end_token": Value("int64"),
"start_byte": Value("int64"),
"end_byte": Value("int64"),
},
"short_answers": Sequence(
{
"start_token": Value("int64"),
"end_token": Value("int64"),
"start_byte": Value("int64"),
"end_byte": Value("int64"),
"text": Value("string"),
}
),
"yes_no_answer": ClassLabel(names=["NO", "YES"]),
}
),
}
)
other = Features( # same but with [] instead of sequences, and with a shuffled fields order
{
"id": Value("string"),
"document": {
"tokens": Sequence({"token": Value("string"), "is_html": Value("bool")}),
"title": Value("string"),
"url": Value("string"),
"html": Value("string"),
},
"question": {
"text": Value("string"),
"tokens": [Value("string")],
},
"annotations": {
"yes_no_answer": [ClassLabel(names=["NO", "YES"])],
"id": [Value("string")],
"long_answer": [
{
"end_byte": Value("int64"),
"start_token": Value("int64"),
"end_token": Value("int64"),
"start_byte": Value("int64"),
}
],
"short_answers": [
Sequence(
{
"text": Value("string"),
"start_token": Value("int64"),
"end_token": Value("int64"),
"start_byte": Value("int64"),
"end_byte": Value("int64"),
}
)
],
},
}
)
expected = Features(
{
"id": Value("string"),
"document": {
"tokens": Sequence({"token": Value("string"), "is_html": Value("bool")}),
"title": Value("string"),
"url": Value("string"),
"html": Value("string"),
},
"question": {
"text": Value("string"),
"tokens": Sequence(Value("string")),
},
"annotations": Sequence(
{
"yes_no_answer": ClassLabel(names=["NO", "YES"]),
"id": Value("string"),
"long_answer": {
"end_byte": Value("int64"),
"start_token": Value("int64"),
"end_token": Value("int64"),
"start_byte": Value("int64"),
},
"short_answers": Sequence(
{
"text": Value("string"),
"start_token": Value("int64"),
"end_token": Value("int64"),
"start_byte": Value("int64"),
"end_byte": Value("int64"),
}
),
}
),
}
)
reordered_features = features.reorder_fields_as(other)
self.assertDictEqual(reordered_features, expected)
self.assertEqual(reordered_features.type, other.type)
self.assertEqual(reordered_features.type, expected.type)
self.assertNotEqual(reordered_features.type, features.type)
def test_flatten(self):
features = Features({"foo": {"bar1": Value("int32"), "bar2": {"foobar": Value("string")}}})
_features = features.copy()
flattened_features = features.flatten()
assert flattened_features == {"foo.bar1": Value("int32"), "foo.bar2.foobar": Value("string")}
assert features == _features, "calling flatten shouldn't alter the current features"
def test_flatten_with_sequence(self):
features = Features({"foo": Sequence({"bar": {"my_value": Value("int32")}})})
_features = features.copy()
flattened_features = features.flatten()
assert flattened_features == {"foo.bar": [{"my_value": Value("int32")}]}
assert features == _features, "calling flatten shouldn't alter the current features"
def test_features_dicts_are_synced(self):
def assert_features_dicts_are_synced(features: Features):
assert (
hasattr(features, "_column_requires_decoding")
and features.keys() == features._column_requires_decoding.keys()
)
features = Features({"foo": Sequence({"bar": {"my_value": Value("int32")}})})
assert_features_dicts_are_synced(features)
features["barfoo"] = Image()
assert_features_dicts_are_synced(features)
del features["barfoo"]
assert_features_dicts_are_synced(features)
features.update({"foobar": Value("string")})
assert_features_dicts_are_synced(features)
features.pop("foobar")
assert_features_dicts_are_synced(features)
features.popitem()
assert_features_dicts_are_synced(features)
features.setdefault("xyz", Value("bool"))
assert_features_dicts_are_synced(features)
features.clear()
assert_features_dicts_are_synced(features)
def test_classlabel_init(tmp_path_factory):
names = ["negative", "positive"]
names_file = str(tmp_path_factory.mktemp("features") / "labels.txt")
with open(names_file, "w", encoding="utf-8") as f:
f.write("\n".join(names))
classlabel = ClassLabel(names=names)
assert classlabel.names == names and classlabel.num_classes == len(names)
classlabel = ClassLabel(names_file=names_file)
assert classlabel.names == names and classlabel.num_classes == len(names)
classlabel = ClassLabel(num_classes=len(names), names=names)
assert classlabel.names == names and classlabel.num_classes == len(names)
classlabel = ClassLabel(num_classes=len(names))
assert classlabel.names == [str(i) for i in range(len(names))] and classlabel.num_classes == len(names)
with pytest.raises(ValueError):
classlabel = ClassLabel(num_classes=len(names) + 1, names=names)
with pytest.raises(ValueError):
classlabel = ClassLabel(names=names, names_file=names_file)
with pytest.raises(ValueError):
classlabel = ClassLabel()
with pytest.raises(TypeError):
classlabel = ClassLabel(names=np.array(names))
def test_classlabel_str2int():
names = ["negative", "positive"]
classlabel = ClassLabel(names=names)
for label in names:
assert classlabel.str2int(label) == names.index(label)
with pytest.raises(ValueError):
classlabel.str2int("__bad_label_name__")
with pytest.raises(ValueError):
classlabel.str2int(1)
with pytest.raises(ValueError):
classlabel.str2int(None)
def test_classlabel_int2str():
names = ["negative", "positive"]
classlabel = ClassLabel(names=names)
for i in range(len(names)):
assert classlabel.int2str(i) == names[i]
with pytest.raises(ValueError):
classlabel.int2str(len(names))
with pytest.raises(ValueError):
classlabel.int2str(-1)
with pytest.raises(ValueError):
classlabel.int2str(None)
def test_classlabel_cast_storage():
names = ["negative", "positive"]
classlabel = ClassLabel(names=names)
# from integers
arr = pa.array([0, 1, -1, -100], type=pa.int64())
result = classlabel.cast_storage(arr)
assert result.type == pa.int64()
assert result.to_pylist() == [0, 1, -1, -100]
arr = pa.array([0, 1, -1, -100], type=pa.int32())
result = classlabel.cast_storage(arr)
assert result.type == pa.int64()
assert result.to_pylist() == [0, 1, -1, -100]
arr = pa.array([3])
with pytest.raises(ValueError):
classlabel.cast_storage(arr)
# from strings
arr = pa.array(["negative", "positive"])
result = classlabel.cast_storage(arr)
assert result.type == pa.int64()
assert result.to_pylist() == [0, 1]
arr = pa.array(["__label_that_doesnt_exist__"])
with pytest.raises(ValueError):
classlabel.cast_storage(arr)
# from nulls
arr = pa.array([None])
result = classlabel.cast_storage(arr)
assert result.type == pa.int64()
assert result.to_pylist() == [None]
# from empty
arr = pa.array([], pa.int64())
result = classlabel.cast_storage(arr)
assert result.type == pa.int64()
assert result.to_pylist() == []
arr = pa.array([], pa.string())
result = classlabel.cast_storage(arr)
assert result.type == pa.int64()
assert result.to_pylist() == []
@pytest.mark.parametrize("class_label_arg", ["names", "names_file"])
def test_class_label_to_and_from_dict(class_label_arg, tmp_path_factory):
names = ["negative", "positive"]
names_file = str(tmp_path_factory.mktemp("features") / "labels.txt")
with open(names_file, "w", encoding="utf-8") as f:
f.write("\n".join(names))
if class_label_arg == "names":
class_label = ClassLabel(names=names)
elif class_label_arg == "names_file":
class_label = ClassLabel(names_file=names_file)
generated_class_label = generate_from_dict(asdict(class_label))
assert generated_class_label == class_label
@pytest.mark.parametrize("inner_type", [Value("int32"), {"subcolumn": Value("int32")}])
def test_encode_nested_example_sequence_with_none(inner_type):
schema = Sequence(inner_type)
obj = None
result = encode_nested_example(schema, obj)
assert result is None
def test_encode_batch_with_example_with_empty_first_elem():
features = Features(
{
"x": Sequence(Sequence(ClassLabel(names=["a", "b"]))),
}
)
encoded_batch = features.encode_batch(
{
"x": [
[["a"], ["b"]],
[[], ["b"]],
]
}
)
assert encoded_batch == {"x": [[[0], [1]], [[], [1]]]}
@pytest.mark.parametrize(
"feature",
[
Value("int32"),
ClassLabel(num_classes=2),
Translation(languages=["en", "fr"]),
TranslationVariableLanguages(languages=["en", "fr"]),
],
)
def test_dataset_feature_with_none(feature):
data = {"col": [None]}
features = Features({"col": feature})
dset = Dataset.from_dict(data, features=features)
item = dset[0]
assert item.keys() == {"col"}
assert item["col"] is None
batch = dset[:1]
assert len(batch) == 1
assert batch.keys() == {"col"}
assert isinstance(batch["col"], list) and all(item is None for item in batch["col"])
column = dset["col"]
assert len(column) == 1
assert isinstance(column, list) and all(item is None for item in column)
# nested tests
data = {"col": [[None]]}
features = Features({"col": Sequence(feature)})
dset = Dataset.from_dict(data, features=features)
item = dset[0]
assert item.keys() == {"col"}
assert all(i is None for i in item["col"])
data = {"nested": [{"col": None}]}
features = Features({"nested": {"col": feature}})
dset = Dataset.from_dict(data, features=features)
item = dset[0]
assert item.keys() == {"nested"}
assert item["nested"].keys() == {"col"}
assert item["nested"]["col"] is None
def iternumpy(key1, value1, value2):
if value1.dtype != value2.dtype: # check only for dtype
raise AssertionError(
f"dtype of '{key1}' key for casted object: {value1.dtype} and expected object: {value2.dtype} not matching"
)
def dict_diff(d1: dict, d2: dict): # check if 2 dictionaries are equal
np.testing.assert_equal(d1, d2) # sanity check if dict values are equal or not
for (k1, v1), (k2, v2) in zip(d1.items(), d2.items()): # check if their values have same dtype or not
if isinstance(v1, dict): # nested dictionary case
dict_diff(v1, v2)
elif isinstance(v1, np.ndarray): # checks if dtype and value of np.ndarray is equal
iternumpy(k1, v1, v2)
elif isinstance(v1, list):
for element1, element2 in zip(v1, v2): # iterates over all elements of list
if isinstance(element1, dict):
dict_diff(element1, element2)
elif isinstance(element1, np.ndarray):
iternumpy(k1, element1, element2)
class CastToPythonObjectsTest(TestCase):
def test_cast_to_python_objects_list(self):
obj = {"col_1": [{"vec": [1, 2, 3], "txt": "foo"}] * 3, "col_2": [[1, 2], [3, 4], [5, 6]]}
expected_obj = {"col_1": [{"vec": [1, 2, 3], "txt": "foo"}] * 3, "col_2": [[1, 2], [3, 4], [5, 6]]}
casted_obj = cast_to_python_objects(obj)
self.assertDictEqual(casted_obj, expected_obj)
def test_cast_to_python_objects_tuple(self):
obj = {"col_1": [{"vec": (1, 2, 3), "txt": "foo"}] * 3, "col_2": [(1, 2), (3, 4), (5, 6)]}
expected_obj = {"col_1": [{"vec": (1, 2, 3), "txt": "foo"}] * 3, "col_2": [(1, 2), (3, 4), (5, 6)]}
casted_obj = cast_to_python_objects(obj)
self.assertDictEqual(casted_obj, expected_obj)
def test_cast_to_python_or_numpy(self):
obj = {"col_1": [{"vec": np.arange(1, 4), "txt": "foo"}] * 3, "col_2": np.arange(1, 7).reshape(3, 2)}
expected_obj = {
"col_1": [{"vec": np.array([1, 2, 3]), "txt": "foo"}] * 3,
"col_2": np.array([[1, 2], [3, 4], [5, 6]]),
}
casted_obj = cast_to_python_objects(obj)
dict_diff(casted_obj, expected_obj)
def test_cast_to_python_objects_series(self):
obj = {
"col_1": pd.Series([{"vec": [1, 2, 3], "txt": "foo"}] * 3),
"col_2": pd.Series([[1, 2], [3, 4], [5, 6]]),
}
expected_obj = {"col_1": [{"vec": [1, 2, 3], "txt": "foo"}] * 3, "col_2": [[1, 2], [3, 4], [5, 6]]}
casted_obj = cast_to_python_objects(obj)
self.assertDictEqual(casted_obj, expected_obj)
def test_cast_to_python_objects_dataframe(self):
obj = pd.DataFrame({"col_1": [{"vec": [1, 2, 3], "txt": "foo"}] * 3, "col_2": [[1, 2], [3, 4], [5, 6]]})
expected_obj = {"col_1": [{"vec": [1, 2, 3], "txt": "foo"}] * 3, "col_2": [[1, 2], [3, 4], [5, 6]]}
casted_obj = cast_to_python_objects(obj)
self.assertDictEqual(casted_obj, expected_obj)
def test_cast_to_python_objects_pandas_timestamp(self):
obj = pd.Timestamp(2020, 1, 1)
expected_obj = obj.to_pydatetime()
casted_obj = cast_to_python_objects(obj)
self.assertEqual(casted_obj, expected_obj)
casted_obj = cast_to_python_objects(pd.Series([obj]))
self.assertListEqual(casted_obj, [expected_obj])
casted_obj = cast_to_python_objects(pd.DataFrame({"a": [obj]}))
self.assertDictEqual(casted_obj, {"a": [expected_obj]})
def test_cast_to_python_objects_pandas_timedelta(self):
obj = pd.Timedelta(seconds=1)
expected_obj = obj.to_pytimedelta()
casted_obj = cast_to_python_objects(obj)
self.assertEqual(casted_obj, expected_obj)
casted_obj = cast_to_python_objects(pd.Series([obj]))
self.assertListEqual(casted_obj, [expected_obj])
casted_obj = cast_to_python_objects(pd.DataFrame({"a": [obj]}))
self.assertDictEqual(casted_obj, {"a": [expected_obj]})
@require_torch
def test_cast_to_python_objects_torch(self):
import torch
obj = {
"col_1": [{"vec": torch.tensor(np.arange(1, 4)), "txt": "foo"}] * 3,
"col_2": torch.tensor(np.arange(1, 7).reshape(3, 2)),
}
expected_obj = {
"col_1": [{"vec": np.array([1, 2, 3]), "txt": "foo"}] * 3,
"col_2": np.array([[1, 2], [3, 4], [5, 6]]),
}
casted_obj = cast_to_python_objects(obj)
dict_diff(casted_obj, expected_obj)
@require_tf
def test_cast_to_python_objects_tf(self):
import tensorflow as tf
obj = {
"col_1": [{"vec": tf.constant(np.arange(1, 4)), "txt": "foo"}] * 3,
"col_2": tf.constant(np.arange(1, 7).reshape(3, 2)),
}
expected_obj = {
"col_1": [{"vec": np.array([1, 2, 3]), "txt": "foo"}] * 3,
"col_2": np.array([[1, 2], [3, 4], [5, 6]]),
}
casted_obj = cast_to_python_objects(obj)
dict_diff(casted_obj, expected_obj)
@require_jax
def test_cast_to_python_objects_jax(self):
import jax.numpy as jnp
obj = {
"col_1": [{"vec": jnp.array(np.arange(1, 4)), "txt": "foo"}] * 3,
"col_2": jnp.array(np.arange(1, 7).reshape(3, 2)),
}
assert obj["col_2"].dtype == jnp.int32
expected_obj = {
"col_1": [{"vec": np.array([1, 2, 3], dtype=np.int32), "txt": "foo"}] * 3,
"col_2": np.array([[1, 2], [3, 4], [5, 6]], dtype=np.int32),
}
casted_obj = cast_to_python_objects(obj)
dict_diff(casted_obj, expected_obj)
@patch("datasets.features.features._cast_to_python_objects", side_effect=_cast_to_python_objects)
def test_dont_iterate_over_each_element_in_a_list(self, mocked_cast):
obj = {"col_1": [[1, 2], [3, 4], [5, 6]]}
cast_to_python_objects(obj)
self.assertEqual(mocked_cast.call_count, 4) # 4 = depth of obj
SIMPLE_FEATURES = [
Features(),
Features({"a": Value("int32")}),
Features({"a": Value("int32", id="my feature")}),
Features({"a": Value("int32"), "b": Value("float64"), "c": Value("string")}),
]
CUSTOM_FEATURES = [
Features({"label": ClassLabel(names=["negative", "positive"])}),
Features({"array": Array2D(dtype="float32", shape=(4, 4))}),
Features({"image": Image()}),
Features({"audio": Audio()}),
Features({"image": Image(decode=False)}),
Features({"audio": Audio(decode=False)}),
Features({"translation": Translation(["en", "fr"])}),
Features({"translation": TranslationVariableLanguages(["en", "fr"])}),
]
NESTED_FEATURES = [
Features({"foo": {}}),
Features({"foo": {"bar": Value("int32")}}),
Features({"foo": {"bar1": Value("int32"), "bar2": Value("float64")}}),
Features({"foo": Sequence(Value("int32"))}),
Features({"foo": Sequence({})}),
Features({"foo": Sequence({"bar": Value("int32")})}),
Features({"foo": [Value("int32")]}),
Features({"foo": [{"bar": Value("int32")}]}),
]
NESTED_CUSTOM_FEATURES = [
Features({"foo": {"bar": ClassLabel(names=["negative", "positive"])}}),
Features({"foo": Sequence(ClassLabel(names=["negative", "positive"]))}),
Features({"foo": Sequence({"bar": ClassLabel(names=["negative", "positive"])})}),
Features({"foo": [ClassLabel(names=["negative", "positive"])]}),
Features({"foo": [{"bar": ClassLabel(names=["negative", "positive"])}]}),
]
@pytest.mark.parametrize("features", SIMPLE_FEATURES + CUSTOM_FEATURES + NESTED_FEATURES + NESTED_CUSTOM_FEATURES)
def test_features_to_dict(features: Features):
features_dict = features.to_dict()
assert isinstance(features_dict, dict)
reloaded = Features.from_dict(features_dict)
assert features == reloaded
@pytest.mark.parametrize("features", SIMPLE_FEATURES + CUSTOM_FEATURES + NESTED_FEATURES + NESTED_CUSTOM_FEATURES)
def test_features_to_yaml_list(features: Features):
features_yaml_list = features._to_yaml_list()
assert isinstance(features_yaml_list, list)
reloaded = Features._from_yaml_list(features_yaml_list)
assert features == reloaded
@pytest.mark.parametrize("features", SIMPLE_FEATURES + CUSTOM_FEATURES + NESTED_FEATURES + NESTED_CUSTOM_FEATURES)
def test_features_to_arrow_schema(features: Features):
arrow_schema = features.arrow_schema
assert isinstance(arrow_schema, pa.Schema)
reloaded = Features.from_arrow_schema(arrow_schema)
assert features == reloaded
| 0
|
hf_public_repos/datasets/tests
|
hf_public_repos/datasets/tests/features/test_image.py
|
import os
import tarfile
import warnings
import numpy as np
import pandas as pd
import pyarrow as pa
import pytest
from datasets import Dataset, Features, Image, Sequence, Value, concatenate_datasets, load_dataset
from datasets.features.image import encode_np_array, image_to_bytes
from ..utils import require_pil
@pytest.fixture
def tar_jpg_path(shared_datadir, tmp_path_factory):
image_path = str(shared_datadir / "test_image_rgb.jpg")
path = tmp_path_factory.mktemp("data") / "image_data.jpg.tar"
with tarfile.TarFile(path, "w") as f:
f.add(image_path, arcname=os.path.basename(image_path))
return path
def iter_archive(archive_path):
with tarfile.open(archive_path) as tar:
for tarinfo in tar:
file_path = tarinfo.name
file_obj = tar.extractfile(tarinfo)
yield file_path, file_obj
def test_image_instantiation():
image = Image()
assert image.id is None
assert image.dtype == "PIL.Image.Image"
assert image.pa_type == pa.struct({"bytes": pa.binary(), "path": pa.string()})
assert image._type == "Image"
def test_image_feature_type_to_arrow():
features = Features({"image": Image()})
assert features.arrow_schema == pa.schema({"image": Image().pa_type})
features = Features({"struct_containing_an_image": {"image": Image()}})
assert features.arrow_schema == pa.schema({"struct_containing_an_image": pa.struct({"image": Image().pa_type})})
features = Features({"sequence_of_images": Sequence(Image())})
assert features.arrow_schema == pa.schema({"sequence_of_images": pa.list_(Image().pa_type)})
@require_pil
@pytest.mark.parametrize(
"build_example",
[
lambda image_path: image_path,
lambda image_path: open(image_path, "rb").read(),
lambda image_path: {"path": image_path},
lambda image_path: {"path": image_path, "bytes": None},
lambda image_path: {"path": image_path, "bytes": open(image_path, "rb").read()},
lambda image_path: {"path": None, "bytes": open(image_path, "rb").read()},
lambda image_path: {"bytes": open(image_path, "rb").read()},
],
)
def test_image_feature_encode_example(shared_datadir, build_example):
import PIL.Image
image_path = str(shared_datadir / "test_image_rgb.jpg")
image = Image()
encoded_example = image.encode_example(build_example(image_path))
assert isinstance(encoded_example, dict)
assert encoded_example.keys() == {"bytes", "path"}
assert encoded_example["bytes"] is not None or encoded_example["path"] is not None
decoded_example = image.decode_example(encoded_example)
assert isinstance(decoded_example, PIL.Image.Image)
@require_pil
def test_image_decode_example(shared_datadir):
import PIL.Image
image_path = str(shared_datadir / "test_image_rgb.jpg")
image = Image()
decoded_example = image.decode_example({"path": image_path, "bytes": None})
assert isinstance(decoded_example, PIL.Image.Image)
assert os.path.samefile(decoded_example.filename, image_path)
assert decoded_example.size == (640, 480)
assert decoded_example.mode == "RGB"
with pytest.raises(RuntimeError):
Image(decode=False).decode_example(image_path)
@require_pil
def test_dataset_with_image_feature(shared_datadir):
import PIL.Image
image_path = str(shared_datadir / "test_image_rgb.jpg")
data = {"image": [image_path]}
features = Features({"image": Image()})
dset = Dataset.from_dict(data, features=features)
item = dset[0]
assert item.keys() == {"image"}
assert isinstance(item["image"], PIL.Image.Image)
assert os.path.samefile(item["image"].filename, image_path)
assert item["image"].format == "JPEG"
assert item["image"].size == (640, 480)
assert item["image"].mode == "RGB"
batch = dset[:1]
assert len(batch) == 1
assert batch.keys() == {"image"}
assert isinstance(batch["image"], list) and all(isinstance(item, PIL.Image.Image) for item in batch["image"])
assert os.path.samefile(batch["image"][0].filename, image_path)
assert batch["image"][0].format == "JPEG"
assert batch["image"][0].size == (640, 480)
assert batch["image"][0].mode == "RGB"
column = dset["image"]
assert len(column) == 1
assert isinstance(column, list) and all(isinstance(item, PIL.Image.Image) for item in column)
assert os.path.samefile(column[0].filename, image_path)
assert column[0].format == "JPEG"
assert column[0].size == (640, 480)
assert column[0].mode == "RGB"
@require_pil
@pytest.mark.parametrize("infer_feature", [False, True])
def test_dataset_with_image_feature_from_pil_image(infer_feature, shared_datadir):
import PIL.Image
image_path = str(shared_datadir / "test_image_rgb.jpg")
data = {"image": [PIL.Image.open(image_path)]}
features = Features({"image": Image()}) if not infer_feature else None
dset = Dataset.from_dict(data, features=features)
item = dset[0]
assert item.keys() == {"image"}
assert isinstance(item["image"], PIL.Image.Image)
assert os.path.samefile(item["image"].filename, image_path)
assert item["image"].format == "JPEG"
assert item["image"].size == (640, 480)
assert item["image"].mode == "RGB"
batch = dset[:1]
assert len(batch) == 1
assert batch.keys() == {"image"}
assert isinstance(batch["image"], list) and all(isinstance(item, PIL.Image.Image) for item in batch["image"])
assert os.path.samefile(batch["image"][0].filename, image_path)
assert batch["image"][0].format == "JPEG"
assert batch["image"][0].size == (640, 480)
assert batch["image"][0].mode == "RGB"
column = dset["image"]
assert len(column) == 1
assert isinstance(column, list) and all(isinstance(item, PIL.Image.Image) for item in column)
assert os.path.samefile(column[0].filename, image_path)
assert column[0].format == "JPEG"
assert column[0].size == (640, 480)
assert column[0].mode == "RGB"
@require_pil
def test_dataset_with_image_feature_from_np_array():
import PIL.Image
image_array = np.arange(640 * 480, dtype=np.int32).reshape(480, 640)
data = {"image": [image_array]}
features = Features({"image": Image()})
dset = Dataset.from_dict(data, features=features)
item = dset[0]
assert item.keys() == {"image"}
assert isinstance(item["image"], PIL.Image.Image)
np.testing.assert_array_equal(np.array(item["image"]), image_array)
assert item["image"].filename == ""
assert item["image"].format in ["PNG", "TIFF"]
assert item["image"].size == (640, 480)
batch = dset[:1]
assert len(batch) == 1
assert batch.keys() == {"image"}
assert isinstance(batch["image"], list) and all(isinstance(item, PIL.Image.Image) for item in batch["image"])
np.testing.assert_array_equal(np.array(batch["image"][0]), image_array)
assert batch["image"][0].filename == ""
assert batch["image"][0].format in ["PNG", "TIFF"]
assert batch["image"][0].size == (640, 480)
column = dset["image"]
assert len(column) == 1
assert isinstance(column, list) and all(isinstance(item, PIL.Image.Image) for item in column)
np.testing.assert_array_equal(np.array(column[0]), image_array)
assert column[0].filename == ""
assert column[0].format in ["PNG", "TIFF"]
assert column[0].size == (640, 480)
@require_pil
def test_dataset_with_image_feature_tar_jpg(tar_jpg_path):
import PIL.Image
data = {"image": []}
for file_path, file_obj in iter_archive(tar_jpg_path):
data["image"].append({"path": file_path, "bytes": file_obj.read()})
break
features = Features({"image": Image()})
dset = Dataset.from_dict(data, features=features)
item = dset[0]
assert item.keys() == {"image"}
assert isinstance(item["image"], PIL.Image.Image)
assert item["image"].filename == ""
assert item["image"].format == "JPEG"
assert item["image"].size == (640, 480)
assert item["image"].mode == "RGB"
batch = dset[:1]
assert len(batch) == 1
assert batch.keys() == {"image"}
assert isinstance(batch["image"], list) and all(isinstance(item, PIL.Image.Image) for item in batch["image"])
assert batch["image"][0].filename == ""
assert batch["image"][0].format == "JPEG"
assert batch["image"][0].size == (640, 480)
assert batch["image"][0].mode == "RGB"
column = dset["image"]
assert len(column) == 1
assert isinstance(column, list) and all(isinstance(item, PIL.Image.Image) for item in column)
assert column[0].filename == ""
assert column[0].format == "JPEG"
assert column[0].size == (640, 480)
assert column[0].mode == "RGB"
@require_pil
def test_dataset_with_image_feature_with_none():
data = {"image": [None]}
features = Features({"image": Image()})
dset = Dataset.from_dict(data, features=features)
item = dset[0]
assert item.keys() == {"image"}
assert item["image"] is None
batch = dset[:1]
assert len(batch) == 1
assert batch.keys() == {"image"}
assert isinstance(batch["image"], list) and all(item is None for item in batch["image"])
column = dset["image"]
assert len(column) == 1
assert isinstance(column, list) and all(item is None for item in column)
# nested tests
data = {"images": [[None]]}
features = Features({"images": Sequence(Image())})
dset = Dataset.from_dict(data, features=features)
item = dset[0]
assert item.keys() == {"images"}
assert all(i is None for i in item["images"])
data = {"nested": [{"image": None}]}
features = Features({"nested": {"image": Image()}})
dset = Dataset.from_dict(data, features=features)
item = dset[0]
assert item.keys() == {"nested"}
assert item["nested"].keys() == {"image"}
assert item["nested"]["image"] is None
@require_pil
@pytest.mark.parametrize(
"build_data",
[
lambda image_path: {"image": [image_path]},
lambda image_path: {"image": [open(image_path, "rb").read()]},
lambda image_path: {"image": [{"path": image_path}]},
lambda image_path: {"image": [{"path": image_path, "bytes": None}]},
lambda image_path: {"image": [{"path": image_path, "bytes": open(image_path, "rb").read()}]},
lambda image_path: {"image": [{"path": None, "bytes": open(image_path, "rb").read()}]},
lambda image_path: {"image": [{"bytes": open(image_path, "rb").read()}]},
],
)
def test_dataset_cast_to_image_features(shared_datadir, build_data):
import PIL.Image
image_path = str(shared_datadir / "test_image_rgb.jpg")
data = build_data(image_path)
dset = Dataset.from_dict(data)
item = dset.cast(Features({"image": Image()}))[0]
assert item.keys() == {"image"}
assert isinstance(item["image"], PIL.Image.Image)
item = dset.cast_column("image", Image())[0]
assert item.keys() == {"image"}
assert isinstance(item["image"], PIL.Image.Image)
@require_pil
def test_dataset_concatenate_image_features(shared_datadir):
# we use a different data structure between 1 and 2 to make sure they are compatible with each other
image_path = str(shared_datadir / "test_image_rgb.jpg")
data1 = {"image": [image_path]}
dset1 = Dataset.from_dict(data1, features=Features({"image": Image()}))
data2 = {"image": [{"bytes": open(image_path, "rb").read()}]}
dset2 = Dataset.from_dict(data2, features=Features({"image": Image()}))
concatenated_dataset = concatenate_datasets([dset1, dset2])
assert len(concatenated_dataset) == len(dset1) + len(dset2)
assert concatenated_dataset[0]["image"] == dset1[0]["image"]
assert concatenated_dataset[1]["image"] == dset2[0]["image"]
@require_pil
def test_dataset_concatenate_nested_image_features(shared_datadir):
# we use a different data structure between 1 and 2 to make sure they are compatible with each other
image_path = str(shared_datadir / "test_image_rgb.jpg")
features = Features({"list_of_structs_of_images": [{"image": Image()}]})
data1 = {"list_of_structs_of_images": [[{"image": image_path}]]}
dset1 = Dataset.from_dict(data1, features=features)
data2 = {"list_of_structs_of_images": [[{"image": {"bytes": open(image_path, "rb").read()}}]]}
dset2 = Dataset.from_dict(data2, features=features)
concatenated_dataset = concatenate_datasets([dset1, dset2])
assert len(concatenated_dataset) == len(dset1) + len(dset2)
assert (
concatenated_dataset[0]["list_of_structs_of_images"][0]["image"]
== dset1[0]["list_of_structs_of_images"][0]["image"]
)
assert (
concatenated_dataset[1]["list_of_structs_of_images"][0]["image"]
== dset2[0]["list_of_structs_of_images"][0]["image"]
)
@require_pil
def test_dataset_with_image_feature_map(shared_datadir):
image_path = str(shared_datadir / "test_image_rgb.jpg")
data = {"image": [image_path], "caption": ["cats sleeping"]}
features = Features({"image": Image(), "caption": Value("string")})
dset = Dataset.from_dict(data, features=features)
for item in dset.cast_column("image", Image(decode=False)):
assert item.keys() == {"image", "caption"}
assert item == {"image": {"path": image_path, "bytes": None}, "caption": "cats sleeping"}
# no decoding
def process_caption(example):
example["caption"] = "Two " + example["caption"]
return example
processed_dset = dset.map(process_caption)
for item in processed_dset.cast_column("image", Image(decode=False)):
assert item.keys() == {"image", "caption"}
assert item == {"image": {"path": image_path, "bytes": None}, "caption": "Two cats sleeping"}
# decoding example
def process_image_by_example(example):
example["mode"] = example["image"].mode
return example
decoded_dset = dset.map(process_image_by_example)
for item in decoded_dset.cast_column("image", Image(decode=False)):
assert item.keys() == {"image", "caption", "mode"}
assert os.path.samefile(item["image"]["path"], image_path)
assert item["caption"] == "cats sleeping"
assert item["mode"] == "RGB"
# decoding batch
def process_image_by_batch(batch):
batch["mode"] = [image.mode for image in batch["image"]]
return batch
decoded_dset = dset.map(process_image_by_batch, batched=True)
for item in decoded_dset.cast_column("image", Image(decode=False)):
assert item.keys() == {"image", "caption", "mode"}
assert os.path.samefile(item["image"]["path"], image_path)
assert item["caption"] == "cats sleeping"
assert item["mode"] == "RGB"
@require_pil
def test_formatted_dataset_with_image_feature_map(shared_datadir):
image_path = str(shared_datadir / "test_image_rgb.jpg")
pil_image = Image().decode_example({"path": image_path, "bytes": None})
data = {"image": [image_path], "caption": ["cats sleeping"]}
features = Features({"image": Image(), "caption": Value("string")})
dset = Dataset.from_dict(data, features=features)
for item in dset.cast_column("image", Image(decode=False)):
assert item.keys() == {"image", "caption"}
assert item == {"image": {"path": image_path, "bytes": None}, "caption": "cats sleeping"}
def process_image_by_example(example):
example["num_channels"] = example["image"].shape[-1]
return example
decoded_dset = dset.with_format("numpy").map(process_image_by_example)
for item in decoded_dset.cast_column("image", Image(decode=False)):
assert item.keys() == {"image", "caption", "num_channels"}
assert item["image"] == encode_np_array(np.array(pil_image))
assert item["caption"] == "cats sleeping"
assert item["num_channels"] == 3
def process_image_by_batch(batch):
batch["num_channels"] = [image.shape[-1] for image in batch["image"]]
return batch
decoded_dset = dset.with_format("numpy").map(process_image_by_batch, batched=True)
for item in decoded_dset.cast_column("image", Image(decode=False)):
assert item.keys() == {"image", "caption", "num_channels"}
assert item["image"] == encode_np_array(np.array(pil_image))
assert item["caption"] == "cats sleeping"
assert item["num_channels"] == 3
@require_pil
def test_dataset_with_image_feature_map_change_image(shared_datadir):
import PIL.Image
image_path = str(shared_datadir / "test_image_rgb.jpg")
pil_image = Image().decode_example({"path": image_path, "bytes": None})
data = {"image": [image_path]}
features = Features({"image": Image()})
dset = Dataset.from_dict(data, features=features)
for item in dset.cast_column("image", Image(decode=False)):
assert item.keys() == {"image"}
assert item == {
"image": {
"bytes": None,
"path": image_path,
}
}
# return pil image
def process_image_resize_by_example(example):
example["image"] = example["image"].resize((100, 100))
return example
decoded_dset = dset.map(process_image_resize_by_example)
for item in decoded_dset.cast_column("image", Image(decode=False)):
assert item.keys() == {"image"}
assert item == {"image": {"bytes": image_to_bytes(pil_image.resize((100, 100))), "path": None}}
def process_image_resize_by_batch(batch):
batch["image"] = [image.resize((100, 100)) for image in batch["image"]]
return batch
decoded_dset = dset.map(process_image_resize_by_batch, batched=True)
for item in decoded_dset.cast_column("image", Image(decode=False)):
assert item.keys() == {"image"}
assert item == {"image": {"bytes": image_to_bytes(pil_image.resize((100, 100))), "path": None}}
# return np.ndarray (e.g. when using albumentations)
def process_image_resize_by_example_return_np_array(example):
example["image"] = np.array(example["image"].resize((100, 100)))
return example
decoded_dset = dset.map(process_image_resize_by_example_return_np_array)
for item in decoded_dset.cast_column("image", Image(decode=False)):
assert item.keys() == {"image"}
assert item == {
"image": {
"bytes": image_to_bytes(PIL.Image.fromarray(np.array(pil_image.resize((100, 100))))),
"path": None,
}
}
def process_image_resize_by_batch_return_np_array(batch):
batch["image"] = [np.array(image.resize((100, 100))) for image in batch["image"]]
return batch
decoded_dset = dset.map(process_image_resize_by_batch_return_np_array, batched=True)
for item in decoded_dset.cast_column("image", Image(decode=False)):
assert item.keys() == {"image"}
assert item == {
"image": {
"bytes": image_to_bytes(PIL.Image.fromarray(np.array(pil_image.resize((100, 100))))),
"path": None,
}
}
@require_pil
def test_formatted_dataset_with_image_feature(shared_datadir):
import PIL.Image
image_path = str(shared_datadir / "test_image_rgb.jpg")
data = {"image": [image_path, image_path]}
features = Features({"image": Image()})
dset = Dataset.from_dict(data, features=features)
with dset.formatted_as("numpy"):
item = dset[0]
assert item.keys() == {"image"}
assert isinstance(item["image"], np.ndarray)
assert item["image"].shape == (480, 640, 3)
batch = dset[:1]
assert batch.keys() == {"image"}
assert len(batch) == 1
assert isinstance(batch["image"], np.ndarray)
assert batch["image"].shape == (1, 480, 640, 3)
column = dset["image"]
assert len(column) == 2
assert isinstance(column, np.ndarray)
assert column.shape == (2, 480, 640, 3)
with dset.formatted_as("pandas"):
item = dset[0]
assert item.shape == (1, 1)
assert item.columns == ["image"]
assert isinstance(item["image"][0], PIL.Image.Image)
assert os.path.samefile(item["image"][0].filename, image_path)
assert item["image"][0].format == "JPEG"
assert item["image"][0].size == (640, 480)
assert item["image"][0].mode == "RGB"
batch = dset[:1]
assert batch.shape == (1, 1)
assert batch.columns == ["image"]
assert isinstance(batch["image"], pd.Series) and all(
isinstance(item, PIL.Image.Image) for item in batch["image"]
)
assert os.path.samefile(batch["image"][0].filename, image_path)
assert batch["image"][0].format == "JPEG"
assert batch["image"][0].size == (640, 480)
assert batch["image"][0].mode == "RGB"
column = dset["image"]
assert len(column) == 2
assert isinstance(column, pd.Series) and all(isinstance(item, PIL.Image.Image) for item in column)
assert os.path.samefile(column[0].filename, image_path)
assert column[0].format == "JPEG"
assert column[0].size == (640, 480)
assert column[0].mode == "RGB"
# Currently, the JSONL reader doesn't support complex feature types so we create a temporary dataset script
# to test streaming (without uploading the test dataset to the hub).
DATASET_LOADING_SCRIPT_NAME = "__dummy_dataset__"
DATASET_LOADING_SCRIPT_CODE = """
import os
import datasets
from datasets import DatasetInfo, Features, Image, Split, SplitGenerator, Value
class __DummyDataset__(datasets.GeneratorBasedBuilder):
def _info(self) -> DatasetInfo:
return DatasetInfo(features=Features({"image": Image(), "caption": Value("string")}))
def _split_generators(self, dl_manager):
return [
SplitGenerator(Split.TRAIN, gen_kwargs={"filepath": os.path.join(dl_manager.manual_dir, "train.txt")}),
]
def _generate_examples(self, filepath, **kwargs):
with open(filepath, encoding="utf-8") as f:
for i, line in enumerate(f):
image_path, caption = line.split(",")
yield i, {"image": image_path.strip(), "caption": caption.strip()}
"""
@pytest.fixture
def data_dir(shared_datadir, tmp_path):
data_dir = tmp_path / "dummy_dataset_data"
data_dir.mkdir()
image_path = str(shared_datadir / "test_image_rgb.jpg")
with open(data_dir / "train.txt", "w") as f:
f.write(f"{image_path},Two cats sleeping\n")
return str(data_dir)
@pytest.fixture
def dataset_loading_script_dir(tmp_path):
script_name = DATASET_LOADING_SCRIPT_NAME
script_dir = tmp_path / script_name
script_dir.mkdir()
script_path = script_dir / f"{script_name}.py"
with open(script_path, "w") as f:
f.write(DATASET_LOADING_SCRIPT_CODE)
return str(script_dir)
@require_pil
@pytest.mark.parametrize("streaming", [False, True])
def test_load_dataset_with_image_feature(shared_datadir, data_dir, dataset_loading_script_dir, streaming):
import PIL.Image
image_path = str(shared_datadir / "test_image_rgb.jpg")
dset = load_dataset(dataset_loading_script_dir, split="train", data_dir=data_dir, streaming=streaming)
item = dset[0] if not streaming else next(iter(dset))
assert item.keys() == {"image", "caption"}
assert isinstance(item["image"], PIL.Image.Image)
assert os.path.samefile(item["image"].filename, image_path)
assert item["image"].format == "JPEG"
assert item["image"].size == (640, 480)
assert item["image"].mode == "RGB"
@require_pil
def test_dataset_with_image_feature_undecoded(shared_datadir):
image_path = str(shared_datadir / "test_image_rgb.jpg")
data = {"image": [image_path]}
features = Features({"image": Image(decode=False)})
dset = Dataset.from_dict(data, features=features)
item = dset[0]
assert item.keys() == {"image"}
assert item["image"] == {"path": image_path, "bytes": None}
batch = dset[:1]
assert batch.keys() == {"image"}
assert len(batch["image"]) == 1
assert batch["image"][0] == {"path": image_path, "bytes": None}
column = dset["image"]
assert len(column) == 1
assert column[0] == {"path": image_path, "bytes": None}
@require_pil
def test_formatted_dataset_with_image_feature_undecoded(shared_datadir):
image_path = str(shared_datadir / "test_image_rgb.jpg")
data = {"image": [image_path]}
features = Features({"image": Image(decode=False)})
dset = Dataset.from_dict(data, features=features)
with dset.formatted_as("numpy"):
item = dset[0]
assert item.keys() == {"image"}
assert item["image"] == {"path": image_path, "bytes": None}
batch = dset[:1]
assert batch.keys() == {"image"}
assert len(batch["image"]) == 1
assert batch["image"][0] == {"path": image_path, "bytes": None}
column = dset["image"]
assert len(column) == 1
assert column[0] == {"path": image_path, "bytes": None}
with dset.formatted_as("pandas"):
item = dset[0]
assert item.shape == (1, 1)
assert item.columns == ["image"]
assert item["image"][0] == {"path": image_path, "bytes": None}
batch = dset[:1]
assert batch.shape == (1, 1)
assert batch.columns == ["image"]
assert batch["image"][0] == {"path": image_path, "bytes": None}
column = dset["image"]
assert len(column) == 1
assert column[0] == {"path": image_path, "bytes": None}
@require_pil
def test_dataset_with_image_feature_map_undecoded(shared_datadir):
image_path = str(shared_datadir / "test_image_rgb.jpg")
data = {"image": [image_path]}
features = Features({"image": Image(decode=False)})
dset = Dataset.from_dict(data, features=features)
def assert_image_example_undecoded(example):
assert example["image"] == {"path": image_path, "bytes": None}
dset.map(assert_image_example_undecoded)
def assert_image_batch_undecoded(batch):
for image in batch["image"]:
assert image == {"path": image_path, "bytes": None}
dset.map(assert_image_batch_undecoded, batched=True)
@require_pil
def test_image_embed_storage(shared_datadir):
image_path = str(shared_datadir / "test_image_rgb.jpg")
example = {"bytes": None, "path": image_path}
storage = pa.array([example], type=pa.struct({"bytes": pa.binary(), "path": pa.string()}))
embedded_storage = Image().embed_storage(storage)
embedded_example = embedded_storage.to_pylist()[0]
assert embedded_example == {"bytes": open(image_path, "rb").read(), "path": "test_image_rgb.jpg"}
@require_pil
@pytest.mark.parametrize(
"array, dtype_cast, expected_image_format",
[
(np.arange(16).reshape(4, 4).astype(np.uint8), "exact_match", "PNG"),
(np.arange(16).reshape(4, 4).astype(np.uint16), "exact_match", "TIFF"),
(np.arange(16).reshape(4, 4).astype(np.int64), "downcast->|i4", "TIFF"),
(np.arange(16).reshape(4, 4).astype(np.complex128), "error", None),
(np.arange(16).reshape(2, 2, 4).astype(np.uint8), "exact_match", "PNG"),
(np.arange(16).reshape(2, 2, 4), "downcast->|u1", "PNG"),
(np.arange(16).reshape(2, 2, 4).astype(np.float64), "error", None),
],
)
def test_encode_np_array(array, dtype_cast, expected_image_format):
if dtype_cast.startswith("downcast"):
_, dest_dtype = dtype_cast.split("->")
dest_dtype = np.dtype(dest_dtype)
with pytest.warns(UserWarning, match=f"Downcasting array dtype.+{dest_dtype}.+"):
encoded_image = Image().encode_example(array)
elif dtype_cast == "error":
with pytest.raises(TypeError):
Image().encode_example(array)
return
else: # exact_match (no warnings are raised)
with warnings.catch_warnings():
warnings.simplefilter("error")
encoded_image = Image().encode_example(array)
assert isinstance(encoded_image, dict)
assert encoded_image.keys() == {"path", "bytes"}
assert encoded_image["path"] is None
assert encoded_image["bytes"] is not None and isinstance(encoded_image["bytes"], bytes)
decoded_image = Image().decode_example(encoded_image)
assert decoded_image.format == expected_image_format
np.testing.assert_array_equal(np.array(decoded_image), array)
| 0
|
hf_public_repos/datasets/tests
|
hf_public_repos/datasets/tests/fixtures/hub.py
|
import time
import uuid
from contextlib import contextmanager
from pathlib import Path
from typing import Optional
import pytest
import requests
from huggingface_hub.hf_api import HfApi, HfFolder, RepositoryNotFoundError
CI_HUB_USER = "__DUMMY_TRANSFORMERS_USER__"
CI_HUB_USER_FULL_NAME = "Dummy User"
CI_HUB_USER_TOKEN = "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"
CI_HUB_ENDPOINT = "https://hub-ci.huggingface.co"
CI_HUB_DATASETS_URL = CI_HUB_ENDPOINT + "/datasets/{repo_id}/resolve/{revision}/{path}"
CI_HFH_HUGGINGFACE_CO_URL_TEMPLATE = CI_HUB_ENDPOINT + "/{repo_id}/resolve/{revision}/{filename}"
CI_HUB_TOKEN_PATH = Path("~/.huggingface/hub_ci_token").expanduser()
@pytest.fixture
def ci_hfh_hf_hub_url(monkeypatch):
monkeypatch.setattr(
"huggingface_hub.file_download.HUGGINGFACE_CO_URL_TEMPLATE", CI_HFH_HUGGINGFACE_CO_URL_TEMPLATE
)
@pytest.fixture
def ci_hub_config(monkeypatch):
monkeypatch.setattr("datasets.config.HF_ENDPOINT", CI_HUB_ENDPOINT)
monkeypatch.setattr("datasets.config.HUB_DATASETS_URL", CI_HUB_DATASETS_URL)
@pytest.fixture
def ci_hub_token_path(monkeypatch):
monkeypatch.setattr("huggingface_hub.hf_api.HfFolder.path_token", CI_HUB_TOKEN_PATH)
@pytest.fixture
def set_ci_hub_access_token(ci_hub_config, ci_hub_token_path):
HfFolder.save_token(CI_HUB_USER_TOKEN)
yield
HfFolder.delete_token()
@pytest.fixture(scope="session")
def hf_api():
return HfApi(endpoint=CI_HUB_ENDPOINT)
@pytest.fixture(scope="session")
def hf_token():
yield CI_HUB_USER_TOKEN
@pytest.fixture
def cleanup_repo(hf_api):
def _cleanup_repo(repo_id):
hf_api.delete_repo(repo_id, token=CI_HUB_USER_TOKEN, repo_type="dataset")
return _cleanup_repo
@pytest.fixture
def temporary_repo(cleanup_repo):
@contextmanager
def _temporary_repo(repo_id: Optional[str] = None):
repo_id = repo_id or f"{CI_HUB_USER}/test-dataset-{uuid.uuid4().hex[:6]}-{int(time.time() * 10e3)}"
try:
yield repo_id
finally:
try:
cleanup_repo(repo_id)
except RepositoryNotFoundError:
pass
return _temporary_repo
@pytest.fixture(scope="session")
def hf_private_dataset_repo_txt_data_(hf_api: HfApi, hf_token, text_file):
repo_name = f"repo_txt_data-{int(time.time() * 10e6)}"
repo_id = f"{CI_HUB_USER}/{repo_name}"
hf_api.create_repo(repo_id, token=hf_token, repo_type="dataset", private=True)
hf_api.upload_file(
token=hf_token,
path_or_fileobj=str(text_file),
path_in_repo="data/text_data.txt",
repo_id=repo_id,
repo_type="dataset",
)
yield repo_id
try:
hf_api.delete_repo(repo_id, token=hf_token, repo_type="dataset")
except (requests.exceptions.HTTPError, ValueError): # catch http error and token invalid error
pass
@pytest.fixture()
def hf_private_dataset_repo_txt_data(hf_private_dataset_repo_txt_data_, ci_hub_config, ci_hfh_hf_hub_url):
return hf_private_dataset_repo_txt_data_
@pytest.fixture(scope="session")
def hf_private_dataset_repo_zipped_txt_data_(hf_api: HfApi, hf_token, zip_csv_with_dir_path):
repo_name = f"repo_zipped_txt_data-{int(time.time() * 10e6)}"
repo_id = f"{CI_HUB_USER}/{repo_name}"
hf_api.create_repo(repo_id, token=hf_token, repo_type="dataset", private=True)
hf_api.upload_file(
token=hf_token,
path_or_fileobj=str(zip_csv_with_dir_path),
path_in_repo="data.zip",
repo_id=repo_id,
repo_type="dataset",
)
yield repo_id
try:
hf_api.delete_repo(repo_id, token=hf_token, repo_type="dataset")
except (requests.exceptions.HTTPError, ValueError): # catch http error and token invalid error
pass
@pytest.fixture()
def hf_private_dataset_repo_zipped_txt_data(
hf_private_dataset_repo_zipped_txt_data_, ci_hub_config, ci_hfh_hf_hub_url
):
return hf_private_dataset_repo_zipped_txt_data_
@pytest.fixture(scope="session")
def hf_private_dataset_repo_zipped_img_data_(hf_api: HfApi, hf_token, zip_image_path):
repo_name = f"repo_zipped_img_data-{int(time.time() * 10e6)}"
repo_id = f"{CI_HUB_USER}/{repo_name}"
hf_api.create_repo(repo_id, token=hf_token, repo_type="dataset", private=True)
hf_api.upload_file(
token=hf_token,
path_or_fileobj=str(zip_image_path),
path_in_repo="data.zip",
repo_id=repo_id,
repo_type="dataset",
)
yield repo_id
try:
hf_api.delete_repo(repo_id, token=hf_token, repo_type="dataset")
except (requests.exceptions.HTTPError, ValueError): # catch http error and token invalid error
pass
@pytest.fixture()
def hf_private_dataset_repo_zipped_img_data(
hf_private_dataset_repo_zipped_img_data_, ci_hub_config, ci_hfh_hf_hub_url
):
return hf_private_dataset_repo_zipped_img_data_
| 0
|
hf_public_repos/datasets/tests
|
hf_public_repos/datasets/tests/fixtures/fsspec.py
|
import posixpath
from pathlib import Path
from unittest.mock import patch
import pytest
from fsspec.implementations.local import AbstractFileSystem, LocalFileSystem, stringify_path
from fsspec.registry import _registry as _fsspec_registry
class MockFileSystem(AbstractFileSystem):
protocol = "mock"
def __init__(self, *args, local_root_dir, **kwargs):
super().__init__()
self._fs = LocalFileSystem(*args, **kwargs)
self.local_root_dir = Path(local_root_dir).resolve().as_posix() + "/"
def mkdir(self, path, *args, **kwargs):
path = posixpath.join(self.local_root_dir, self._strip_protocol(path))
return self._fs.mkdir(path, *args, **kwargs)
def makedirs(self, path, *args, **kwargs):
path = posixpath.join(self.local_root_dir, self._strip_protocol(path))
return self._fs.makedirs(path, *args, **kwargs)
def rmdir(self, path):
path = posixpath.join(self.local_root_dir, self._strip_protocol(path))
return self._fs.rmdir(path)
def ls(self, path, detail=True, *args, **kwargs):
path = posixpath.join(self.local_root_dir, self._strip_protocol(path))
out = self._fs.ls(path, detail=detail, *args, **kwargs)
if detail:
return [{**info, "name": info["name"][len(self.local_root_dir) :]} for info in out]
else:
return [name[len(self.local_root_dir) :] for name in out]
def info(self, path, *args, **kwargs):
path = posixpath.join(self.local_root_dir, self._strip_protocol(path))
out = dict(self._fs.info(path, *args, **kwargs))
out["name"] = out["name"][len(self.local_root_dir) :]
return out
def cp_file(self, path1, path2, *args, **kwargs):
path1 = posixpath.join(self.local_root_dir, self._strip_protocol(path1))
path2 = posixpath.join(self.local_root_dir, self._strip_protocol(path2))
return self._fs.cp_file(path1, path2, *args, **kwargs)
def rm_file(self, path, *args, **kwargs):
path = posixpath.join(self.local_root_dir, self._strip_protocol(path))
return self._fs.rm_file(path, *args, **kwargs)
def rm(self, path, *args, **kwargs):
path = posixpath.join(self.local_root_dir, self._strip_protocol(path))
return self._fs.rm(path, *args, **kwargs)
def _open(self, path, *args, **kwargs):
path = posixpath.join(self.local_root_dir, self._strip_protocol(path))
return self._fs._open(path, *args, **kwargs)
def created(self, path):
path = posixpath.join(self.local_root_dir, self._strip_protocol(path))
return self._fs.created(path)
def modified(self, path):
path = posixpath.join(self.local_root_dir, self._strip_protocol(path))
return self._fs.modified(path)
@classmethod
def _strip_protocol(cls, path):
path = stringify_path(path)
if path.startswith("mock://"):
path = path[7:]
return path
class TmpDirFileSystem(MockFileSystem):
protocol = "tmp"
tmp_dir = None
def __init__(self, *args, **kwargs):
assert self.tmp_dir is not None, "TmpDirFileSystem.tmp_dir is not set"
super().__init__(*args, **kwargs, local_root_dir=self.tmp_dir, auto_mkdir=True)
@classmethod
def _strip_protocol(cls, path):
path = stringify_path(path)
if path.startswith("tmp://"):
path = path[6:]
return path
@pytest.fixture
def mock_fsspec():
_fsspec_registry["mock"] = MockFileSystem
_fsspec_registry["tmp"] = TmpDirFileSystem
yield
del _fsspec_registry["mock"]
del _fsspec_registry["tmp"]
@pytest.fixture
def mockfs(tmp_path_factory, mock_fsspec):
local_fs_dir = tmp_path_factory.mktemp("mockfs")
return MockFileSystem(local_root_dir=local_fs_dir, auto_mkdir=True)
@pytest.fixture
def tmpfs(tmp_path_factory, mock_fsspec):
tmp_fs_dir = tmp_path_factory.mktemp("tmpfs")
with patch.object(TmpDirFileSystem, "tmp_dir", tmp_fs_dir):
yield TmpDirFileSystem()
TmpDirFileSystem.clear_instance_cache()
| 0
|
hf_public_repos/datasets/tests
|
hf_public_repos/datasets/tests/fixtures/files.py
|
import contextlib
import csv
import json
import os
import sqlite3
import tarfile
import textwrap
import zipfile
import pyarrow as pa
import pyarrow.parquet as pq
import pytest
import datasets
import datasets.config
# dataset + arrow_file
@pytest.fixture(scope="session")
def dataset():
n = 10
features = datasets.Features(
{
"tokens": datasets.Sequence(datasets.Value("string")),
"labels": datasets.Sequence(datasets.ClassLabel(names=["negative", "positive"])),
"answers": datasets.Sequence(
{
"text": datasets.Value("string"),
"answer_start": datasets.Value("int32"),
}
),
"id": datasets.Value("int64"),
}
)
dataset = datasets.Dataset.from_dict(
{
"tokens": [["foo"] * 5] * n,
"labels": [[1] * 5] * n,
"answers": [{"answer_start": [97], "text": ["1976"]}] * 10,
"id": list(range(n)),
},
features=features,
)
return dataset
@pytest.fixture(scope="session")
def arrow_file(tmp_path_factory, dataset):
filename = str(tmp_path_factory.mktemp("data") / "file.arrow")
dataset.map(cache_file_name=filename)
return filename
# FILE_CONTENT + files
FILE_CONTENT = """\
Text data.
Second line of data."""
@pytest.fixture(scope="session")
def text_file(tmp_path_factory):
filename = tmp_path_factory.mktemp("data") / "file.txt"
data = FILE_CONTENT
with open(filename, "w") as f:
f.write(data)
return filename
@pytest.fixture(scope="session")
def bz2_file(tmp_path_factory):
import bz2
path = tmp_path_factory.mktemp("data") / "file.txt.bz2"
data = bytes(FILE_CONTENT, "utf-8")
with bz2.open(path, "wb") as f:
f.write(data)
return path
@pytest.fixture(scope="session")
def gz_file(tmp_path_factory):
import gzip
path = str(tmp_path_factory.mktemp("data") / "file.txt.gz")
data = bytes(FILE_CONTENT, "utf-8")
with gzip.open(path, "wb") as f:
f.write(data)
return path
@pytest.fixture(scope="session")
def lz4_file(tmp_path_factory):
if datasets.config.LZ4_AVAILABLE:
import lz4.frame
path = tmp_path_factory.mktemp("data") / "file.txt.lz4"
data = bytes(FILE_CONTENT, "utf-8")
with lz4.frame.open(path, "wb") as f:
f.write(data)
return path
@pytest.fixture(scope="session")
def seven_zip_file(tmp_path_factory, text_file):
if datasets.config.PY7ZR_AVAILABLE:
import py7zr
path = tmp_path_factory.mktemp("data") / "file.txt.7z"
with py7zr.SevenZipFile(path, "w") as archive:
archive.write(text_file, arcname=os.path.basename(text_file))
return path
@pytest.fixture(scope="session")
def tar_file(tmp_path_factory, text_file):
import tarfile
path = tmp_path_factory.mktemp("data") / "file.txt.tar"
with tarfile.TarFile(path, "w") as f:
f.add(text_file, arcname=os.path.basename(text_file))
return path
@pytest.fixture(scope="session")
def xz_file(tmp_path_factory):
import lzma
path = tmp_path_factory.mktemp("data") / "file.txt.xz"
data = bytes(FILE_CONTENT, "utf-8")
with lzma.open(path, "wb") as f:
f.write(data)
return path
@pytest.fixture(scope="session")
def zip_file(tmp_path_factory, text_file):
import zipfile
path = tmp_path_factory.mktemp("data") / "file.txt.zip"
with zipfile.ZipFile(path, "w") as f:
f.write(text_file, arcname=os.path.basename(text_file))
return path
@pytest.fixture(scope="session")
def zstd_file(tmp_path_factory):
if datasets.config.ZSTANDARD_AVAILABLE:
import zstandard as zstd
path = tmp_path_factory.mktemp("data") / "file.txt.zst"
data = bytes(FILE_CONTENT, "utf-8")
with zstd.open(path, "wb") as f:
f.write(data)
return path
# xml_file
@pytest.fixture(scope="session")
def xml_file(tmp_path_factory):
filename = tmp_path_factory.mktemp("data") / "file.xml"
data = textwrap.dedent(
"""\
<?xml version="1.0" encoding="UTF-8" ?>
<tmx version="1.4">
<header segtype="sentence" srclang="ca" />
<body>
<tu>
<tuv xml:lang="ca"><seg>Contingut 1</seg></tuv>
<tuv xml:lang="en"><seg>Content 1</seg></tuv>
</tu>
<tu>
<tuv xml:lang="ca"><seg>Contingut 2</seg></tuv>
<tuv xml:lang="en"><seg>Content 2</seg></tuv>
</tu>
<tu>
<tuv xml:lang="ca"><seg>Contingut 3</seg></tuv>
<tuv xml:lang="en"><seg>Content 3</seg></tuv>
</tu>
<tu>
<tuv xml:lang="ca"><seg>Contingut 4</seg></tuv>
<tuv xml:lang="en"><seg>Content 4</seg></tuv>
</tu>
<tu>
<tuv xml:lang="ca"><seg>Contingut 5</seg></tuv>
<tuv xml:lang="en"><seg>Content 5</seg></tuv>
</tu>
</body>
</tmx>"""
)
with open(filename, "w") as f:
f.write(data)
return filename
DATA = [
{"col_1": "0", "col_2": 0, "col_3": 0.0},
{"col_1": "1", "col_2": 1, "col_3": 1.0},
{"col_1": "2", "col_2": 2, "col_3": 2.0},
{"col_1": "3", "col_2": 3, "col_3": 3.0},
]
DATA2 = [
{"col_1": "4", "col_2": 4, "col_3": 4.0},
{"col_1": "5", "col_2": 5, "col_3": 5.0},
]
DATA_DICT_OF_LISTS = {
"col_1": ["0", "1", "2", "3"],
"col_2": [0, 1, 2, 3],
"col_3": [0.0, 1.0, 2.0, 3.0],
}
DATA_312 = [
{"col_3": 0.0, "col_1": "0", "col_2": 0},
{"col_3": 1.0, "col_1": "1", "col_2": 1},
]
DATA_STR = [
{"col_1": "s0", "col_2": 0, "col_3": 0.0},
{"col_1": "s1", "col_2": 1, "col_3": 1.0},
{"col_1": "s2", "col_2": 2, "col_3": 2.0},
{"col_1": "s3", "col_2": 3, "col_3": 3.0},
]
@pytest.fixture(scope="session")
def dataset_dict():
return DATA_DICT_OF_LISTS
@pytest.fixture(scope="session")
def arrow_path(tmp_path_factory):
dataset = datasets.Dataset.from_dict(DATA_DICT_OF_LISTS)
path = str(tmp_path_factory.mktemp("data") / "dataset.arrow")
dataset.map(cache_file_name=path)
return path
@pytest.fixture(scope="session")
def sqlite_path(tmp_path_factory):
path = str(tmp_path_factory.mktemp("data") / "dataset.sqlite")
with contextlib.closing(sqlite3.connect(path)) as con:
cur = con.cursor()
cur.execute("CREATE TABLE dataset(col_1 text, col_2 int, col_3 real)")
for item in DATA:
cur.execute("INSERT INTO dataset(col_1, col_2, col_3) VALUES (?, ?, ?)", tuple(item.values()))
con.commit()
return path
@pytest.fixture(scope="session")
def csv_path(tmp_path_factory):
path = str(tmp_path_factory.mktemp("data") / "dataset.csv")
with open(path, "w", newline="") as f:
writer = csv.DictWriter(f, fieldnames=["col_1", "col_2", "col_3"])
writer.writeheader()
for item in DATA:
writer.writerow(item)
return path
@pytest.fixture(scope="session")
def csv2_path(tmp_path_factory):
path = str(tmp_path_factory.mktemp("data") / "dataset2.csv")
with open(path, "w", newline="") as f:
writer = csv.DictWriter(f, fieldnames=["col_1", "col_2", "col_3"])
writer.writeheader()
for item in DATA:
writer.writerow(item)
return path
@pytest.fixture(scope="session")
def bz2_csv_path(csv_path, tmp_path_factory):
import bz2
path = tmp_path_factory.mktemp("data") / "dataset.csv.bz2"
with open(csv_path, "rb") as f:
data = f.read()
# data = bytes(FILE_CONTENT, "utf-8")
with bz2.open(path, "wb") as f:
f.write(data)
return path
@pytest.fixture(scope="session")
def zip_csv_path(csv_path, csv2_path, tmp_path_factory):
path = tmp_path_factory.mktemp("zip_csv_path") / "csv-dataset.zip"
with zipfile.ZipFile(path, "w") as f:
f.write(csv_path, arcname=os.path.basename(csv_path))
f.write(csv2_path, arcname=os.path.basename(csv2_path))
return path
@pytest.fixture(scope="session")
def zip_uppercase_csv_path(csv_path, csv2_path, tmp_path_factory):
path = tmp_path_factory.mktemp("data") / "dataset.csv.zip"
with zipfile.ZipFile(path, "w") as f:
f.write(csv_path, arcname=os.path.basename(csv_path.replace(".csv", ".CSV")))
f.write(csv2_path, arcname=os.path.basename(csv2_path.replace(".csv", ".CSV")))
return path
@pytest.fixture(scope="session")
def zip_csv_with_dir_path(csv_path, csv2_path, tmp_path_factory):
path = tmp_path_factory.mktemp("data") / "dataset_with_dir.csv.zip"
with zipfile.ZipFile(path, "w") as f:
f.write(csv_path, arcname=os.path.join("main_dir", os.path.basename(csv_path)))
f.write(csv2_path, arcname=os.path.join("main_dir", os.path.basename(csv2_path)))
return path
@pytest.fixture(scope="session")
def parquet_path(tmp_path_factory):
path = str(tmp_path_factory.mktemp("data") / "dataset.parquet")
schema = pa.schema(
{
"col_1": pa.string(),
"col_2": pa.int64(),
"col_3": pa.float64(),
}
)
with open(path, "wb") as f:
writer = pq.ParquetWriter(f, schema=schema)
pa_table = pa.Table.from_pydict({k: [DATA[i][k] for i in range(len(DATA))] for k in DATA[0]}, schema=schema)
writer.write_table(pa_table)
writer.close()
return path
@pytest.fixture(scope="session")
def json_list_of_dicts_path(tmp_path_factory):
path = str(tmp_path_factory.mktemp("data") / "dataset.json")
data = {"data": DATA}
with open(path, "w") as f:
json.dump(data, f)
return path
@pytest.fixture(scope="session")
def json_dict_of_lists_path(tmp_path_factory):
path = str(tmp_path_factory.mktemp("data") / "dataset.json")
data = {"data": DATA_DICT_OF_LISTS}
with open(path, "w") as f:
json.dump(data, f)
return path
@pytest.fixture(scope="session")
def jsonl_path(tmp_path_factory):
path = str(tmp_path_factory.mktemp("data") / "dataset.jsonl")
with open(path, "w") as f:
for item in DATA:
f.write(json.dumps(item) + "\n")
return path
@pytest.fixture(scope="session")
def jsonl2_path(tmp_path_factory):
path = str(tmp_path_factory.mktemp("data") / "dataset2.jsonl")
with open(path, "w") as f:
for item in DATA:
f.write(json.dumps(item) + "\n")
return path
@pytest.fixture(scope="session")
def jsonl_312_path(tmp_path_factory):
path = str(tmp_path_factory.mktemp("data") / "dataset_312.jsonl")
with open(path, "w") as f:
for item in DATA_312:
f.write(json.dumps(item) + "\n")
return path
@pytest.fixture(scope="session")
def jsonl_str_path(tmp_path_factory):
path = str(tmp_path_factory.mktemp("data") / "dataset-str.jsonl")
with open(path, "w") as f:
for item in DATA_STR:
f.write(json.dumps(item) + "\n")
return path
@pytest.fixture(scope="session")
def text_gz_path(tmp_path_factory, text_path):
import gzip
path = str(tmp_path_factory.mktemp("data") / "dataset.txt.gz")
with open(text_path, "rb") as orig_file:
with gzip.open(path, "wb") as zipped_file:
zipped_file.writelines(orig_file)
return path
@pytest.fixture(scope="session")
def jsonl_gz_path(tmp_path_factory, jsonl_path):
import gzip
path = str(tmp_path_factory.mktemp("data") / "dataset.jsonl.gz")
with open(jsonl_path, "rb") as orig_file:
with gzip.open(path, "wb") as zipped_file:
zipped_file.writelines(orig_file)
return path
@pytest.fixture(scope="session")
def zip_jsonl_path(jsonl_path, jsonl2_path, tmp_path_factory):
path = tmp_path_factory.mktemp("data") / "dataset.jsonl.zip"
with zipfile.ZipFile(path, "w") as f:
f.write(jsonl_path, arcname=os.path.basename(jsonl_path))
f.write(jsonl2_path, arcname=os.path.basename(jsonl2_path))
return path
@pytest.fixture(scope="session")
def zip_nested_jsonl_path(zip_jsonl_path, jsonl_path, jsonl2_path, tmp_path_factory):
path = tmp_path_factory.mktemp("data") / "dataset_nested.jsonl.zip"
with zipfile.ZipFile(path, "w") as f:
f.write(zip_jsonl_path, arcname=os.path.join("nested", os.path.basename(zip_jsonl_path)))
return path
@pytest.fixture(scope="session")
def zip_jsonl_with_dir_path(jsonl_path, jsonl2_path, tmp_path_factory):
path = tmp_path_factory.mktemp("data") / "dataset_with_dir.jsonl.zip"
with zipfile.ZipFile(path, "w") as f:
f.write(jsonl_path, arcname=os.path.join("main_dir", os.path.basename(jsonl_path)))
f.write(jsonl2_path, arcname=os.path.join("main_dir", os.path.basename(jsonl2_path)))
return path
@pytest.fixture(scope="session")
def tar_jsonl_path(jsonl_path, jsonl2_path, tmp_path_factory):
path = tmp_path_factory.mktemp("data") / "dataset.jsonl.tar"
with tarfile.TarFile(path, "w") as f:
f.add(jsonl_path, arcname=os.path.basename(jsonl_path))
f.add(jsonl2_path, arcname=os.path.basename(jsonl2_path))
return path
@pytest.fixture(scope="session")
def tar_nested_jsonl_path(tar_jsonl_path, jsonl_path, jsonl2_path, tmp_path_factory):
path = tmp_path_factory.mktemp("data") / "dataset_nested.jsonl.tar"
with tarfile.TarFile(path, "w") as f:
f.add(tar_jsonl_path, arcname=os.path.join("nested", os.path.basename(tar_jsonl_path)))
return path
@pytest.fixture(scope="session")
def text_path(tmp_path_factory):
data = ["0", "1", "2", "3"]
path = str(tmp_path_factory.mktemp("data") / "dataset.txt")
with open(path, "w") as f:
for item in data:
f.write(item + "\n")
return path
@pytest.fixture(scope="session")
def text2_path(tmp_path_factory):
data = ["0", "1", "2", "3"]
path = str(tmp_path_factory.mktemp("data") / "dataset2.txt")
with open(path, "w") as f:
for item in data:
f.write(item + "\n")
return path
@pytest.fixture(scope="session")
def text_dir_with_unsupported_extension(tmp_path_factory):
data = ["0", "1", "2", "3"]
path = tmp_path_factory.mktemp("data") / "dataset.abc"
with open(path, "w") as f:
for item in data:
f.write(item + "\n")
return path
@pytest.fixture(scope="session")
def zip_text_path(text_path, text2_path, tmp_path_factory):
path = tmp_path_factory.mktemp("data") / "dataset.text.zip"
with zipfile.ZipFile(path, "w") as f:
f.write(text_path, arcname=os.path.basename(text_path))
f.write(text2_path, arcname=os.path.basename(text2_path))
return path
@pytest.fixture(scope="session")
def zip_text_with_dir_path(text_path, text2_path, tmp_path_factory):
path = tmp_path_factory.mktemp("data") / "dataset_with_dir.text.zip"
with zipfile.ZipFile(path, "w") as f:
f.write(text_path, arcname=os.path.join("main_dir", os.path.basename(text_path)))
f.write(text2_path, arcname=os.path.join("main_dir", os.path.basename(text2_path)))
return path
@pytest.fixture(scope="session")
def zip_unsupported_ext_path(text_path, text2_path, tmp_path_factory):
path = tmp_path_factory.mktemp("data") / "dataset.ext.zip"
with zipfile.ZipFile(path, "w") as f:
f.write(text_path, arcname=os.path.basename("unsupported.ext"))
f.write(text2_path, arcname=os.path.basename("unsupported_2.ext"))
return path
@pytest.fixture(scope="session")
def text_path_with_unicode_new_lines(tmp_path_factory):
text = "\n".join(["First", "Second\u2029with Unicode new line", "Third"])
path = str(tmp_path_factory.mktemp("data") / "dataset_with_unicode_new_lines.txt")
with open(path, "w", encoding="utf-8") as f:
f.write(text)
return path
@pytest.fixture(scope="session")
def image_file():
return os.path.join("tests", "features", "data", "test_image_rgb.jpg")
@pytest.fixture(scope="session")
def audio_file():
return os.path.join("tests", "features", "data", "test_audio_44100.wav")
@pytest.fixture(scope="session")
def zip_image_path(image_file, tmp_path_factory):
path = tmp_path_factory.mktemp("data") / "dataset.img.zip"
with zipfile.ZipFile(path, "w") as f:
f.write(image_file, arcname=os.path.basename(image_file))
f.write(image_file, arcname=os.path.basename(image_file).replace(".jpg", "2.jpg"))
return path
@pytest.fixture(scope="session")
def data_dir_with_hidden_files(tmp_path_factory):
data_dir = tmp_path_factory.mktemp("data_dir")
(data_dir / "subdir").mkdir()
with open(data_dir / "subdir" / "train.txt", "w") as f:
f.write("foo\n" * 10)
with open(data_dir / "subdir" / "test.txt", "w") as f:
f.write("bar\n" * 10)
# hidden file
with open(data_dir / "subdir" / ".test.txt", "w") as f:
f.write("bar\n" * 10)
# hidden directory
(data_dir / ".subdir").mkdir()
with open(data_dir / ".subdir" / "train.txt", "w") as f:
f.write("foo\n" * 10)
with open(data_dir / ".subdir" / "test.txt", "w") as f:
f.write("bar\n" * 10)
return data_dir
| 0
|
hf_public_repos/datasets/tests
|
hf_public_repos/datasets/tests/commands/conftest.py
|
import pytest
DATASET_LOADING_SCRIPT_NAME = "__dummy_dataset1__"
DATASET_LOADING_SCRIPT_CODE = """
import json
import os
import datasets
REPO_URL = "https://huggingface.co/datasets/hf-internal-testing/raw_jsonl/resolve/main/"
URLS = {"train": REPO_URL + "wikiann-bn-train.jsonl", "validation": REPO_URL + "wikiann-bn-validation.jsonl"}
class __DummyDataset1__(datasets.GeneratorBasedBuilder):
def _info(self):
features = datasets.Features(
{
"tokens": datasets.Sequence(datasets.Value("string")),
"ner_tags": datasets.Sequence(
datasets.features.ClassLabel(
names=[
"O",
"B-PER",
"I-PER",
"B-ORG",
"I-ORG",
"B-LOC",
"I-LOC",
]
)
),
"langs": datasets.Sequence(datasets.Value("string")),
"spans": datasets.Sequence(datasets.Value("string")),
}
)
return datasets.DatasetInfo(features=features)
def _split_generators(self, dl_manager):
dl_path = dl_manager.download(URLS)
return [
datasets.SplitGenerator(datasets.Split.TRAIN, gen_kwargs={"filepath": dl_path["train"]}),
datasets.SplitGenerator(datasets.Split.VALIDATION, gen_kwargs={"filepath": dl_path["validation"]}),
]
def _generate_examples(self, filepath):
with open(filepath, "r", encoding="utf-8") as f:
for i, line in enumerate(f):
yield i, json.loads(line)
"""
@pytest.fixture
def dataset_loading_script_name():
return DATASET_LOADING_SCRIPT_NAME
@pytest.fixture
def dataset_loading_script_code():
return DATASET_LOADING_SCRIPT_CODE
@pytest.fixture
def dataset_loading_script_dir(dataset_loading_script_name, dataset_loading_script_code, tmp_path):
script_name = dataset_loading_script_name
script_dir = tmp_path / "datasets" / script_name
script_dir.mkdir(parents=True)
script_path = script_dir / f"{script_name}.py"
with open(script_path, "w") as f:
f.write(dataset_loading_script_code)
return str(script_dir)
| 0
|
hf_public_repos/datasets/tests
|
hf_public_repos/datasets/tests/commands/test_test.py
|
import os
from collections import namedtuple
import pytest
from datasets import ClassLabel, Features, Sequence, Value
from datasets.commands.test import TestCommand
from datasets.info import DatasetInfo, DatasetInfosDict
_TestCommandArgs = namedtuple(
"_TestCommandArgs",
[
"dataset",
"name",
"cache_dir",
"data_dir",
"all_configs",
"save_infos",
"ignore_verifications",
"force_redownload",
"clear_cache",
],
defaults=[None, None, None, False, False, False, False, False],
)
def is_1percent_close(source, target):
return (abs(source - target) / target) < 0.01
@pytest.mark.integration
def test_test_command(dataset_loading_script_dir):
args = _TestCommandArgs(dataset=dataset_loading_script_dir, all_configs=True, save_infos=True)
test_command = TestCommand(*args)
test_command.run()
dataset_readme_path = os.path.join(dataset_loading_script_dir, "README.md")
assert os.path.exists(dataset_readme_path)
dataset_infos = DatasetInfosDict.from_directory(dataset_loading_script_dir)
expected_dataset_infos = DatasetInfosDict(
{
"default": DatasetInfo(
features=Features(
{
"tokens": Sequence(Value("string")),
"ner_tags": Sequence(
ClassLabel(names=["O", "B-PER", "I-PER", "B-ORG", "I-ORG", "B-LOC", "I-LOC"])
),
"langs": Sequence(Value("string")),
"spans": Sequence(Value("string")),
}
),
splits=[
{
"name": "train",
"num_bytes": 2351563,
"num_examples": 10000,
},
{
"name": "validation",
"num_bytes": 238418,
"num_examples": 1000,
},
],
download_size=3940680,
dataset_size=2589981,
)
}
)
assert dataset_infos.keys() == expected_dataset_infos.keys()
for key in DatasetInfo._INCLUDED_INFO_IN_YAML:
result, expected = getattr(dataset_infos["default"], key), getattr(expected_dataset_infos["default"], key)
if key == "num_bytes":
assert is_1percent_close(result, expected)
elif key == "splits":
assert list(result) == list(expected)
for split in result:
assert result[split].name == expected[split].name
assert result[split].num_examples == expected[split].num_examples
assert is_1percent_close(result[split].num_bytes, expected[split].num_bytes)
else:
result == expected
| 0
|
hf_public_repos/datasets
|
hf_public_repos/datasets/notebooks/Overview.ipynb
|
# install datasets
!pip install datasets# Let's import the library. We typically only need at most two methods:
from datasets import list_datasets, load_dataset
from pprint import pprint# Currently available datasets
datasets = list_datasets()
print(f"🤩 Currently {len(datasets)} datasets are available on the hub:")
pprint(datasets[:100] + [f"{len(datasets) - 100} more..."], compact=True)# You can access various attributes of the datasets before downloading them
squad_dataset = list_datasets(with_details=True)[datasets.index('squad')]
pprint(squad_dataset.__dict__) # It's a simple python dataclass# Downloading and loading a dataset
dataset = load_dataset('squad', split='validation[:10%]')# Informations on the dataset (description, citation, size, splits, format...)
# are provided in `dataset.info` (a simple python dataclass) and also as direct attributes in the dataset object
pprint(dataset.info.__dict__)print(dataset)print(f"👉 Dataset len(dataset): {len(dataset)}")
print("\n👉 First item 'dataset[0]':")
pprint(dataset[0])# Or get slices with several examples:
print("\n👉Slice of the two items 'dataset[10:12]':")
pprint(dataset[10:12])# You can get a full column of the dataset by indexing with its name as a string:
print(dataset['question'][:10])print(dataset[0]['question'] == dataset['question'][0])
print(dataset[10:20]['context'] == dataset['context'][10:20])# You can inspect the dataset column names and types
print("Column names:")
pprint(dataset.column_names)
print("Features:")
pprint(dataset.features)# Datasets also have shapes informations
print("The number of rows", dataset.num_rows, "also available as len(dataset)", len(dataset))
print("The number of columns", dataset.num_columns)
print("The shape (rows, columns)", dataset.shape)# Let's print the length of each `context` string in our subset of the dataset
# (10% of the validation i.e. 1057 examples)
dataset.map(lambda example: print(len(example['context']), end=','))from datasets import logging
logging.set_verbosity_warning()
dataset.map(lambda example: print(len(example['context']), end=','))# Let's keep it verbose for our tutorial though
from datasets import logging
logging.set_verbosity_info()# Let's add a prefix 'My cute title: ' to each of our titles
def add_prefix_to_title(example):
example['title'] = 'My cute title: ' + example['title']
return example
prefixed_dataset = dataset.map(add_prefix_to_title)
print(prefixed_dataset.unique('title')) # `.unique()` is a super fast way to print the unique elemnts in a column (see the doc for all the methods)# Since the input example dict is updated with our function output dict,
# we can actually just return the updated 'title' field
titled_dataset = dataset.map(lambda example: {'title': 'My cutest title: ' + example['title']})
print(titled_dataset.unique('title'))# This will remove the 'title' column while doing the update (after having send it the the mapped function so you can use it in your function!)
less_columns_dataset = dataset.map(lambda example: {'new_title': 'Wouhahh: ' + example['title']}, remove_columns=['title'])
print(less_columns_dataset.column_names)
print(less_columns_dataset.unique('new_title'))# This will add the index in the dataset to the 'question' field
with_indices_dataset = dataset.map(lambda example, idx: {'question': f'{idx}: ' + example['question']},
with_indices=True)
pprint(with_indices_dataset['question'][:5])# Let's import a fast tokenizer that can work on batched inputs
# (the 'Fast' tokenizers in HuggingFace)
from transformers import BertTokenizerFast, logging as transformers_logging
transformers_logging.set_verbosity_warning()
tokenizer = BertTokenizerFast.from_pretrained('bert-base-cased')# Now let's batch tokenize our dataset 'context'
encoded_dataset = dataset.map(lambda example: tokenizer(example['context']), batched=True)
print("encoded_dataset[0]")
pprint(encoded_dataset[0], compact=True)# we have added additional columns
pprint(encoded_dataset.column_names)# Let show a more complex processing with the full preparation of the SQuAD dataset
# for training a model from Transformers
def convert_to_features(batch):
# Tokenize contexts and questions (as pairs of inputs)
encodings = tokenizer(batch['context'], batch['question'], truncation=True)
# Compute start and end tokens for labels
start_positions, end_positions = [], []
for i, answer in enumerate(batch['answers']):
first_char = answer['answer_start'][0]
last_char = first_char + len(answer['text'][0]) - 1
start_positions.append(encodings.char_to_token(i, first_char))
end_positions.append(encodings.char_to_token(i, last_char))
encodings.update({'start_positions': start_positions, 'end_positions': end_positions})
return encodings
encoded_dataset = dataset.map(convert_to_features, batched=True)# Now our dataset comprise the labels for the start and end position
# as well as the offsets for converting back tokens
# in span of the original string for evaluation
print("column_names", encoded_dataset.column_names)
print("start_positions", encoded_dataset[:5]['start_positions'])image_dataset = load_dataset("cats_vs_dogs", split="train")
image_dataset[0]image_dataset[0]["image"]from datasets import load_dataset
audio_dataset = load_dataset("common_voice", "fi", split="train")
audio_dataset[0]audio_dataset[0]["audio"]["array"], audio_dataset[0]["audio"]["sampling_rate"]from datasets import Audio
audio_dataset = audio_dataset.cast_column("audio", Audio(sampling_rate=16_000))
audio_dataset[0]["audio"]["array"], audio_dataset[0]["audio"]["sampling_rate"]columns_to_return = ['input_ids', 'token_type_ids', 'attention_mask', 'start_positions', 'end_positions']
# Uncomment whichever one is appropriate for you
# encoded_dataset.set_format(type='torch', columns=columns_to_return)
encoded_dataset.set_format(type='tensorflow', columns=columns_to_return)
# Our dataset indexing output is now ready for being used in a pytorch dataloader
pprint(encoded_dataset[1], compact=True)# Note that the columns are not removed from the dataset, just not returned when calling __getitem__
# Similarly the inner type of the dataset is not changed to torch.Tensor, the conversion and filtering is done on-the-fly when querying the dataset
print(encoded_dataset.column_names)# We can remove the formatting with `.reset_format()`
# or, identically, a call to `.set_format()` with no arguments
encoded_dataset.reset_format()
pprint(encoded_dataset[1], compact=True)# The current format can be checked with `.format`,
# which is a dict of the type and formatting
pprint(encoded_dataset.format)import torch
from datasets import load_dataset
from transformers import BertTokenizerFast
# Load our training dataset and tokenizer
dataset = load_dataset('squad')
tokenizer = BertTokenizerFast.from_pretrained('bert-base-cased')
def get_correct_alignement(context, answer):
""" Some original examples in SQuAD have indices wrong by 1 or 2 character. We test and fix this here. """
gold_text = answer['text'][0]
start_idx = answer['answer_start'][0]
end_idx = start_idx + len(gold_text)
if context[start_idx:end_idx] == gold_text:
return start_idx, end_idx # When the gold label position is good
elif context[start_idx-1:end_idx-1] == gold_text:
return start_idx-1, end_idx-1 # When the gold label is off by one character
elif context[start_idx-2:end_idx-2] == gold_text:
return start_idx-2, end_idx-2 # When the gold label is off by two character
else:
raise ValueError()
# Tokenize our training dataset
def convert_to_features(example_batch):
# Tokenize contexts and questions (as pairs of inputs)
encodings = tokenizer(example_batch['context'], example_batch['question'], truncation=True)
# Compute start and end tokens for labels using Transformers's fast tokenizers alignement methods.
start_positions, end_positions = [], []
for i, (context, answer) in enumerate(zip(example_batch['context'], example_batch['answers'])):
start_idx, end_idx = get_correct_alignement(context, answer)
start_positions.append(encodings.char_to_token(i, start_idx))
end_positions.append(encodings.char_to_token(i, end_idx-1))
encodings.update({'start_positions': start_positions, 'end_positions': end_positions})
return encodings
encoded_dataset = dataset.map(convert_to_features, batched=True)
# Format our dataset to outputs torch.Tensor to train a pytorch model
columns = ['input_ids', 'token_type_ids', 'attention_mask', 'start_positions', 'end_positions']
encoded_dataset.set_format(type='torch', columns=columns)
# Instantiate a PyTorch Dataloader around our dataset
# Let's do dynamic batching (pad on the fly with our own collate_fn)
def collate_fn(examples):
return tokenizer.pad(examples, return_tensors='pt')
dataloader = torch.utils.data.DataLoader(encoded_dataset['train'], collate_fn=collate_fn, batch_size=8)columns = ['input_ids', 'token_type_ids', 'attention_mask', 'start_positions', 'end_positions']
# Let's do dynamic batching (pad on the fly with our own collate_fn)
def collate_fn(examples):
return tokenizer.pad(examples, return_tensors='np')
# to_tf_dataset() returns a tf.data.Dataset that we can pass straight to model.fit().
encoded_tf_dataset = encoded_dataset['train'].to_tf_dataset(
columns=columns,
collate_fn=collate_fn,
batch_size=8,
shuffle=True,
)# Let's load a pretrained Bert model and a simple optimizer
from transformers import AutoModelForQuestionAnswering
model = AutoModelForQuestionAnswering.from_pretrained('bert-base-cased', return_dict=True)
optimizer = torch.optim.Adam(model.parameters(), lr=1e-5)# Now let's train our model
device = 'cuda' if torch.cuda.is_available() else 'cpu'
model.train().to(device)
for i, batch in enumerate(dataloader):
batch.to(device)
outputs = model(**batch)
loss = outputs.loss
loss.backward()
optimizer.step()
model.zero_grad()
print(f'Step {i} - loss: {loss:.3}')
if i > 5:
break# Let's load a pretrained Bert model and a simple optimizer
from transformers import TFAutoModelForQuestionAnswering
import tensorflow as tf
model = TFAutoModelForQuestionAnswering.from_pretrained('bert-base-cased')
# No loss argument!
model.compile(optimizer=tf.keras.optimizers.Adam(1e-5))model.fit(encoded_tf_dataset, epochs=1, steps_per_epoch=3)
| 0
|
hf_public_repos/datasets
|
hf_public_repos/datasets/notebooks/README.md
|
<!---
Copyright 2023 The HuggingFace Team. All rights reserved.
Licensed under the Apache License, Version 2.0 (the "License");
you may not use this file except in compliance with the License.
You may obtain a copy of the License at
http://www.apache.org/licenses/LICENSE-2.0
Unless required by applicable law or agreed to in writing, software
distributed under the License is distributed on an "AS IS" BASIS,
WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
See the License for the specific language governing permissions and
limitations under the License.
-->
# 🤗 Datasets Notebooks
You can find here a list of the official notebooks provided by Hugging Face.
Also, we would like to list here interesting content created by the community.
If you wrote some notebook(s) leveraging 🤗 Datasets and would like it to be listed here, please open a
Pull Request so it can be included under the Community notebooks.
## Hugging Face's notebooks 🤗
### Documentation notebooks
You can open any page of the documentation as a notebook in Colab (there is a button directly on said pages) but they are also listed here if you need them:
| Notebook | Description | | |
|:----------|:-------------|:-------------|------:|
| [Quickstart](https://github.com/huggingface/notebooks/blob/main/datasets_doc/en/quickstart.ipynb) | A quick presentation on integrating Datasets into a model training workflow |[](https://colab.research.google.com/github/huggingface/notebooks/blob/main/datasets_doc/en/quickstart.ipynb)| [](https://studiolab.sagemaker.aws/import/github/huggingface/notebooks/blob/main/datasets_doc/en/quickstart.ipynb)|
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/mahalanobis/README.md
|
# Metric Card for Mahalanobis Distance
## Metric Description
Mahalonobis distance is the distance between a point and a distribution (as opposed to the distance between two points), making it the multivariate equivalent of the Euclidean distance.
It is often used in multivariate anomaly detection, classification on highly imbalanced datasets and one-class classification.
## How to Use
At minimum, this metric requires two `list`s of datapoints:
```python
>>> mahalanobis_metric = datasets.load_metric("mahalanobis")
>>> results = mahalanobis_metric.compute(reference_distribution=[[0, 1], [1, 0]], X=[[0, 1]])
```
### Inputs
- `X` (`list`): data points to be compared with the `reference_distribution`.
- `reference_distribution` (`list`): data points from the reference distribution that we want to compare to.
### Output Values
`mahalanobis` (`array`): the Mahalonobis distance for each data point in `X`.
```python
>>> print(results)
{'mahalanobis': array([0.5])}
```
#### Values from Popular Papers
*N/A*
### Example
```python
>>> mahalanobis_metric = datasets.load_metric("mahalanobis")
>>> results = mahalanobis_metric.compute(reference_distribution=[[0, 1], [1, 0]], X=[[0, 1]])
>>> print(results)
{'mahalanobis': array([0.5])}
```
## Limitations and Bias
The Mahalanobis distance is only able to capture linear relationships between the variables, which means it cannot capture all types of outliers. Mahalanobis distance also fails to faithfully represent data that is highly skewed or multimodal.
## Citation
```bibtex
@inproceedings{mahalanobis1936generalized,
title={On the generalized distance in statistics},
author={Mahalanobis, Prasanta Chandra},
year={1936},
organization={National Institute of Science of India}
}
```
```bibtex
@article{de2000mahalanobis,
title={The Mahalanobis distance},
author={De Maesschalck, Roy and Jouan-Rimbaud, Delphine and Massart, D{\'e}sir{\'e} L},
journal={Chemometrics and intelligent laboratory systems},
volume={50},
number={1},
pages={1--18},
year={2000},
publisher={Elsevier}
}
```
## Further References
-[Wikipedia -- Mahalanobis Distance](https://en.wikipedia.org/wiki/Mahalanobis_distance)
-[Machine Learning Plus -- Mahalanobis Distance](https://www.machinelearningplus.com/statistics/mahalanobis-distance/)
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/mahalanobis/mahalanobis.py
|
# Copyright 2021 The HuggingFace Datasets Authors and the current dataset script contributor.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""Mahalanobis metric."""
import numpy as np
import datasets
_DESCRIPTION = """
Compute the Mahalanobis Distance
Mahalonobis distance is the distance between a point and a distribution.
And not between two distinct points. It is effectively a multivariate equivalent of the Euclidean distance.
It was introduced by Prof. P. C. Mahalanobis in 1936
and has been used in various statistical applications ever since
[source: https://www.machinelearningplus.com/statistics/mahalanobis-distance/]
"""
_CITATION = """\
@article{de2000mahalanobis,
title={The mahalanobis distance},
author={De Maesschalck, Roy and Jouan-Rimbaud, Delphine and Massart, D{\'e}sir{\'e} L},
journal={Chemometrics and intelligent laboratory systems},
volume={50},
number={1},
pages={1--18},
year={2000},
publisher={Elsevier}
}
"""
_KWARGS_DESCRIPTION = """
Args:
X: List of datapoints to be compared with the `reference_distribution`.
reference_distribution: List of datapoints from the reference distribution we want to compare to.
Returns:
mahalanobis: The Mahalonobis distance for each datapoint in `X`.
Examples:
>>> mahalanobis_metric = datasets.load_metric("mahalanobis")
>>> results = mahalanobis_metric.compute(reference_distribution=[[0, 1], [1, 0]], X=[[0, 1]])
>>> print(results)
{'mahalanobis': array([0.5])}
"""
@datasets.utils.file_utils.add_start_docstrings(_DESCRIPTION, _KWARGS_DESCRIPTION)
class Mahalanobis(datasets.Metric):
def _info(self):
return datasets.MetricInfo(
description=_DESCRIPTION,
citation=_CITATION,
inputs_description=_KWARGS_DESCRIPTION,
features=datasets.Features(
{
"X": datasets.Sequence(datasets.Value("float", id="sequence"), id="X"),
}
),
)
def _compute(self, X, reference_distribution):
# convert to numpy arrays
X = np.array(X)
reference_distribution = np.array(reference_distribution)
# Assert that arrays are 2D
if len(X.shape) != 2:
raise ValueError("Expected `X` to be a 2D vector")
if len(reference_distribution.shape) != 2:
raise ValueError("Expected `reference_distribution` to be a 2D vector")
if reference_distribution.shape[0] < 2:
raise ValueError(
"Expected `reference_distribution` to be a 2D vector with more than one element in the first dimension"
)
# Get mahalanobis distance for each prediction
X_minus_mu = X - np.mean(reference_distribution)
cov = np.cov(reference_distribution.T)
try:
inv_covmat = np.linalg.inv(cov)
except np.linalg.LinAlgError:
inv_covmat = np.linalg.pinv(cov)
left_term = np.dot(X_minus_mu, inv_covmat)
mahal_dist = np.dot(left_term, X_minus_mu.T).diagonal()
return {"mahalanobis": mahal_dist}
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/frugalscore/README.md
|
# Metric Card for FrugalScore
## Metric Description
FrugalScore is a reference-based metric for Natural Language Generation (NLG) model evaluation. It is based on a distillation approach that allows to learn a fixed, low cost version of any expensive NLG metric, while retaining most of its original performance.
The FrugalScore models are obtained by continuing the pretraining of small models on a synthetic dataset constructed using summarization, backtranslation and denoising models. During the training, the small models learn the internal mapping of the expensive metric, including any similarity function.
## How to use
When loading FrugalScore, you can indicate the model you wish to use to compute the score. The default model is `moussaKam/frugalscore_tiny_bert-base_bert-score`, and a full list of models can be found in the [Limitations and bias](#Limitations-and-bias) section.
```python
>>> from datasets import load_metric
>>> frugalscore = load_metric("frugalscore", "moussaKam/frugalscore_medium_bert-base_mover-score")
```
FrugalScore calculates how good are the predictions given some references, based on a set of scores.
The inputs it takes are:
`predictions`: a list of strings representing the predictions to score.
`references`: a list of string representing the references for each prediction.
Its optional arguments are:
`batch_size`: the batch size for predictions (default value is `32`).
`max_length`: the maximum sequence length (default value is `128`).
`device`: either "gpu" or "cpu" (default value is `None`).
```python
>>> results = frugalscore.compute(predictions=['hello there', 'huggingface'], references=['hello world', 'hugging face'], batch_size=16, max_length=64, device="gpu")
```
## Output values
The output of FrugalScore is a dictionary with the list of scores for each prediction-reference pair:
```python
{'scores': [0.6307541, 0.6449357]}
```
### Values from popular papers
The [original FrugalScore paper](https://arxiv.org/abs/2110.08559) reported that FrugalScore-Tiny retains 97.7/94.7% of the original performance compared to [BertScore](https://huggingface.co/metrics/bertscore) while running 54 times faster and having 84 times less parameters.
## Examples
Maximal values (exact match between `references` and `predictions`):
```python
>>> from datasets import load_metric
>>> frugalscore = load_metric("frugalscore")
>>> results = frugalscore.compute(predictions=['hello world'], references=['hello world'])
>>> print(results)
{'scores': [0.9891098]}
```
Partial values:
```python
>>> from datasets import load_metric
>>> frugalscore = load_metric("frugalscore")
>>> results = frugalscore.compute(predictions=['hello world'], references=['hugging face'])
>>> print(results)
{'scores': [0.42482382]}
```
## Limitations and bias
FrugalScore is based on [BertScore](https://huggingface.co/metrics/bertscore) and [MoverScore](https://arxiv.org/abs/1909.02622), and the models used are based on the original models used for these scores.
The full list of available models for FrugalScore is:
| FrugalScore | Student | Teacher | Method |
|----------------------------------------------------|-------------|----------------|------------|
| [moussaKam/frugalscore_tiny_bert-base_bert-score](https://huggingface.co/moussaKam/frugalscore_tiny_bert-base_bert-score) | BERT-tiny | BERT-Base | BERTScore |
| [moussaKam/frugalscore_small_bert-base_bert-score](https://huggingface.co/moussaKam/frugalscore_small_bert-base_bert-score) | BERT-small | BERT-Base | BERTScore |
| [moussaKam/frugalscore_medium_bert-base_bert-score](https://huggingface.co/moussaKam/frugalscore_medium_bert-base_bert-score) | BERT-medium | BERT-Base | BERTScore |
| [moussaKam/frugalscore_tiny_roberta_bert-score](https://huggingface.co/moussaKam/frugalscore_tiny_roberta_bert-score) | BERT-tiny | RoBERTa-Large | BERTScore |
| [moussaKam/frugalscore_small_roberta_bert-score](https://huggingface.co/moussaKam/frugalscore_small_roberta_bert-score) | BERT-small | RoBERTa-Large | BERTScore |
| [moussaKam/frugalscore_medium_roberta_bert-score](https://huggingface.co/moussaKam/frugalscore_medium_roberta_bert-score) | BERT-medium | RoBERTa-Large | BERTScore |
| [moussaKam/frugalscore_tiny_deberta_bert-score](https://huggingface.co/moussaKam/frugalscore_tiny_deberta_bert-score) | BERT-tiny | DeBERTa-XLarge | BERTScore |
| [moussaKam/frugalscore_small_deberta_bert-score](https://huggingface.co/moussaKam/frugalscore_small_deberta_bert-score) | BERT-small | DeBERTa-XLarge | BERTScore |
| [moussaKam/frugalscore_medium_deberta_bert-score](https://huggingface.co/moussaKam/frugalscore_medium_deberta_bert-score) | BERT-medium | DeBERTa-XLarge | BERTScore |
| [moussaKam/frugalscore_tiny_bert-base_mover-score](https://huggingface.co/moussaKam/frugalscore_tiny_bert-base_mover-score) | BERT-tiny | BERT-Base | MoverScore |
| [moussaKam/frugalscore_small_bert-base_mover-score](https://huggingface.co/moussaKam/frugalscore_small_bert-base_mover-score) | BERT-small | BERT-Base | MoverScore |
| [moussaKam/frugalscore_medium_bert-base_mover-score](https://huggingface.co/moussaKam/frugalscore_medium_bert-base_mover-score) | BERT-medium | BERT-Base | MoverScore |
Depending on the size of the model picked, the loading time will vary: the `tiny` models will load very quickly, whereas the `medium` ones can take several minutes, depending on your Internet connection.
## Citation
```bibtex
@article{eddine2021frugalscore,
title={FrugalScore: Learning Cheaper, Lighter and Faster Evaluation Metrics for Automatic Text Generation},
author={Eddine, Moussa Kamal and Shang, Guokan and Tixier, Antoine J-P and Vazirgiannis, Michalis},
journal={arXiv preprint arXiv:2110.08559},
year={2021}
}
```
## Further References
- [Original FrugalScore code](https://github.com/moussaKam/FrugalScore)
- [FrugalScore paper](https://arxiv.org/abs/2110.08559)
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/frugalscore/frugalscore.py
|
# Copyright 2022 The HuggingFace Datasets Authors and the current metric script contributor.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""FrugalScore metric."""
import torch
from transformers import AutoModelForSequenceClassification, AutoTokenizer, Trainer, TrainingArguments
import datasets
_CITATION = """\
@article{eddine2021frugalscore,
title={FrugalScore: Learning Cheaper, Lighter and Faster Evaluation Metrics for Automatic Text Generation},
author={Eddine, Moussa Kamal and Shang, Guokan and Tixier, Antoine J-P and Vazirgiannis, Michalis},
journal={arXiv preprint arXiv:2110.08559},
year={2021}
}
"""
_DESCRIPTION = """\
FrugalScore is a reference-based metric for NLG models evaluation. It is based on a distillation approach that allows to learn a fixed, low cost version of any expensive NLG metric, while retaining most of its original performance.
"""
_KWARGS_DESCRIPTION = """
Calculates how good are predictions given some references, using certain scores.
Args:
predictions (list of str): list of predictions to score. Each predictions
should be a string.
references (list of str): list of reference for each prediction. Each
reference should be a string.
batch_size (int): the batch size for predictions.
max_length (int): maximum sequence length.
device (str): either gpu or cpu
Returns:
scores (list of int): list of scores.
Examples:
>>> frugalscore = datasets.load_metric("frugalscore")
>>> results = frugalscore.compute(predictions=['hello there', 'huggingface'], references=['hello world', 'hugging face'])
>>> print([round(s, 3) for s in results["scores"]])
[0.631, 0.645]
"""
@datasets.utils.file_utils.add_start_docstrings(_DESCRIPTION, _KWARGS_DESCRIPTION)
class FRUGALSCORE(datasets.Metric):
def _info(self):
return datasets.MetricInfo(
description=_DESCRIPTION,
citation=_CITATION,
inputs_description=_KWARGS_DESCRIPTION,
features=datasets.Features(
{
"predictions": datasets.Value("string"),
"references": datasets.Value("string"),
}
),
homepage="https://github.com/moussaKam/FrugalScore",
)
def _download_and_prepare(self, dl_manager):
if self.config_name == "default":
checkpoint = "moussaKam/frugalscore_tiny_bert-base_bert-score"
else:
checkpoint = self.config_name
self.model = AutoModelForSequenceClassification.from_pretrained(checkpoint)
self.tokenizer = AutoTokenizer.from_pretrained(checkpoint)
def _compute(
self,
predictions,
references,
batch_size=32,
max_length=128,
device=None,
):
"""Returns the scores"""
assert len(predictions) == len(
references
), "predictions and references should have the same number of sentences."
if device is not None:
assert device in ["gpu", "cpu"], "device should be either gpu or cpu."
else:
device = "gpu" if torch.cuda.is_available() else "cpu"
training_args = TrainingArguments(
"trainer",
fp16=(device == "gpu"),
per_device_eval_batch_size=batch_size,
report_to="all",
no_cuda=(device == "cpu"),
log_level="warning",
)
dataset = {"sentence1": predictions, "sentence2": references}
raw_datasets = datasets.Dataset.from_dict(dataset)
def tokenize_function(data):
return self.tokenizer(
data["sentence1"], data["sentence2"], max_length=max_length, truncation=True, padding=True
)
tokenized_datasets = raw_datasets.map(tokenize_function, batched=True)
tokenized_datasets.remove_columns(["sentence1", "sentence2"])
trainer = Trainer(self.model, training_args, tokenizer=self.tokenizer)
predictions = trainer.predict(tokenized_datasets)
return {"scores": list(predictions.predictions.squeeze(-1))}
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/google_bleu/README.md
|
# Metric Card for Google BLEU (GLEU)
## Metric Description
The BLEU score has some undesirable properties when used for single
sentences, as it was designed to be a corpus measure. The Google BLEU score, also known as GLEU score, is designed to limit these undesirable properties when used for single sentences.
To calculate this score, all sub-sequences of 1, 2, 3 or 4 tokens in output and target sequence (n-grams) are recorded. The precision and recall, described below, are then computed.
- **precision:** the ratio of the number of matching n-grams to the number of total n-grams in the generated output sequence
- **recall:** the ratio of the number of matching n-grams to the number of total n-grams in the target (ground truth) sequence
The minimum value of precision and recall is then returned as the score.
## Intended Uses
This metric is generally used to evaluate machine translation models. It is especially used when scores of individual (prediction, reference) sentence pairs are needed, as opposed to when averaging over the (prediction, reference) scores for a whole corpus. That being said, it can also be used when averaging over the scores for a whole corpus.
Because it performs better on individual sentence pairs as compared to BLEU, Google BLEU has also been used in RL experiments.
## How to Use
At minimum, this metric takes predictions and references:
```python
>>> sentence1 = "the cat sat on the mat".split()
>>> sentence2 = "the cat ate the mat".split()
>>> google_bleu = datasets.load_metric("google_bleu")
>>> result = google_bleu.compute(predictions=[sentence1], references=[[sentence2]])
>>> print(result)
{'google_bleu': 0.3333333333333333}
```
### Inputs
- **predictions** (list of list of str): list of translations to score. Each translation should be tokenized into a list of tokens.
- **references** (list of list of list of str): list of lists of references for each translation. Each reference should be tokenized into a list of tokens.
- **min_len** (int): The minimum order of n-gram this function should extract. Defaults to 1.
- **max_len** (int): The maximum order of n-gram this function should extract. Defaults to 4.
### Output Values
This metric returns the following in a dict:
- **google_bleu** (float): google_bleu score
The output format is as follows:
```python
{'google_bleu': google_bleu score}
```
This metric can take on values from 0 to 1, inclusive. Higher scores are better, with 0 indicating no matches, and 1 indicating a perfect match.
Note that this score is symmetrical when switching output and target. This means that, given two sentences, `sentence1` and `sentence2`, whatever score is output when `sentence1` is the predicted sentence and `sencence2` is the reference sentence will be the same as when the sentences are swapped and `sentence2` is the predicted sentence while `sentence1` is the reference sentence. In code, this looks like:
```python
sentence1 = "the cat sat on the mat".split()
sentence2 = "the cat ate the mat".split()
google_bleu = datasets.load_metric("google_bleu")
result_a = google_bleu.compute(predictions=[sentence1], references=[[sentence2]])
result_b = google_bleu.compute(predictions=[sentence2], references=[[sentence1]])
print(result_a == result_b)
>>> True
```
#### Values from Popular Papers
### Examples
Example with one reference per sample:
```python
>>> hyp1 = ['It', 'is', 'a', 'guide', 'to', 'action', 'which',
... 'ensures', 'that', 'the', 'rubber', 'duck', 'always',
... 'disobeys', 'the', 'commands', 'of', 'the', 'cat']
>>> ref1a = ['It', 'is', 'the', 'guiding', 'principle', 'which',
... 'guarantees', 'the', 'rubber', 'duck', 'forces', 'never',
... 'being', 'under', 'the', 'command', 'of', 'the', 'cat']
>>> hyp2 = ['he', 'read', 'the', 'book', 'because', 'he', 'was',
... 'interested', 'in', 'world', 'history']
>>> ref2a = ['he', 'was', 'interested', 'in', 'world', 'history',
... 'because', 'he', 'read', 'the', 'book']
>>> list_of_references = [[ref1a], [ref2a]]
>>> hypotheses = [hyp1, hyp2]
>>> google_bleu = datasets.load_metric("google_bleu")
>>> results = google_bleu.compute(predictions=hypotheses, references=list_of_references)
>>> print(round(results["google_bleu"], 2))
0.44
```
Example with multiple references for the first sample:
```python
>>> hyp1 = ['It', 'is', 'a', 'guide', 'to', 'action', 'which',
... 'ensures', 'that', 'the', 'rubber', 'duck', 'always',
... 'disobeys', 'the', 'commands', 'of', 'the', 'cat']
>>> ref1a = ['It', 'is', 'the', 'guiding', 'principle', 'which',
... 'guarantees', 'the', 'rubber', 'duck', 'forces', 'never',
... 'being', 'under', 'the', 'command', 'of', 'the', 'cat']
>>> ref1b = ['It', 'is', 'a', 'guide', 'to', 'action', 'that',
... 'ensures', 'that', 'the', 'rubber', 'duck', 'will', 'never',
... 'heed', 'the', 'cat', 'commands']
>>> ref1c = ['It', 'is', 'the', 'practical', 'guide', 'for', 'the',
... 'rubber', 'duck', 'army', 'never', 'to', 'heed', 'the', 'directions',
... 'of', 'the', 'cat']
>>> hyp2 = ['he', 'read', 'the', 'book', 'because', 'he', 'was',
... 'interested', 'in', 'world', 'history']
>>> ref2a = ['he', 'was', 'interested', 'in', 'world', 'history',
... 'because', 'he', 'read', 'the', 'book']
>>> list_of_references = [[ref1a, ref1b, ref1c], [ref2a]]
>>> hypotheses = [hyp1, hyp2]
>>> google_bleu = datasets.load_metric("google_bleu")
>>> results = google_bleu.compute(predictions=hypotheses, references=list_of_references)
>>> print(round(results["google_bleu"], 2))
0.61
```
Example with multiple references for the first sample, and with `min_len` adjusted to `2`, instead of the default `1`:
```python
>>> hyp1 = ['It', 'is', 'a', 'guide', 'to', 'action', 'which',
... 'ensures', 'that', 'the', 'rubber', 'duck', 'always',
... 'disobeys', 'the', 'commands', 'of', 'the', 'cat']
>>> ref1a = ['It', 'is', 'the', 'guiding', 'principle', 'which',
... 'guarantees', 'the', 'rubber', 'duck', 'forces', 'never',
... 'being', 'under', 'the', 'command', 'of', 'the', 'cat']
>>> ref1b = ['It', 'is', 'a', 'guide', 'to', 'action', 'that',
... 'ensures', 'that', 'the', 'rubber', 'duck', 'will', 'never',
... 'heed', 'the', 'cat', 'commands']
>>> ref1c = ['It', 'is', 'the', 'practical', 'guide', 'for', 'the',
... 'rubber', 'duck', 'army', 'never', 'to', 'heed', 'the', 'directions',
... 'of', 'the', 'cat']
>>> hyp2 = ['he', 'read', 'the', 'book', 'because', 'he', 'was',
... 'interested', 'in', 'world', 'history']
>>> ref2a = ['he', 'was', 'interested', 'in', 'world', 'history',
... 'because', 'he', 'read', 'the', 'book']
>>> list_of_references = [[ref1a, ref1b, ref1c], [ref2a]]
>>> hypotheses = [hyp1, hyp2]
>>> google_bleu = datasets.load_metric("google_bleu")
>>> results = google_bleu.compute(predictions=hypotheses, references=list_of_references, min_len=2)
>>> print(round(results["google_bleu"], 2))
0.53
```
Example with multiple references for the first sample, with `min_len` adjusted to `2`, instead of the default `1`, and `max_len` adjusted to `6` instead of the default `4`:
```python
>>> hyp1 = ['It', 'is', 'a', 'guide', 'to', 'action', 'which',
... 'ensures', 'that', 'the', 'rubber', 'duck', 'always',
... 'disobeys', 'the', 'commands', 'of', 'the', 'cat']
>>> ref1a = ['It', 'is', 'the', 'guiding', 'principle', 'which',
... 'guarantees', 'the', 'rubber', 'duck', 'forces', 'never',
... 'being', 'under', 'the', 'command', 'of', 'the', 'cat']
>>> ref1b = ['It', 'is', 'a', 'guide', 'to', 'action', 'that',
... 'ensures', 'that', 'the', 'rubber', 'duck', 'will', 'never',
... 'heed', 'the', 'cat', 'commands']
>>> ref1c = ['It', 'is', 'the', 'practical', 'guide', 'for', 'the',
... 'rubber', 'duck', 'army', 'never', 'to', 'heed', 'the', 'directions',
... 'of', 'the', 'cat']
>>> hyp2 = ['he', 'read', 'the', 'book', 'because', 'he', 'was',
... 'interested', 'in', 'world', 'history']
>>> ref2a = ['he', 'was', 'interested', 'in', 'world', 'history',
... 'because', 'he', 'read', 'the', 'book']
>>> list_of_references = [[ref1a, ref1b, ref1c], [ref2a]]
>>> hypotheses = [hyp1, hyp2]
>>> google_bleu = datasets.load_metric("google_bleu")
>>> results = google_bleu.compute(predictions=hypotheses,references=list_of_references, min_len=2, max_len=6)
>>> print(round(results["google_bleu"], 2))
0.4
```
## Limitations and Bias
## Citation
```bibtex
@misc{wu2016googles,
title={Google's Neural Machine Translation System: Bridging the Gap between Human and Machine Translation},
author={Yonghui Wu and Mike Schuster and Zhifeng Chen and Quoc V. Le and Mohammad Norouzi and Wolfgang Macherey and Maxim Krikun and Yuan Cao and Qin Gao and Klaus Macherey and Jeff Klingner and Apurva Shah and Melvin Johnson and Xiaobing Liu and Łukasz Kaiser and Stephan Gouws and Yoshikiyo Kato and Taku Kudo and Hideto Kazawa and Keith Stevens and George Kurian and Nishant Patil and Wei Wang and Cliff Young and Jason Smith and Jason Riesa and Alex Rudnick and Oriol Vinyals and Greg Corrado and Macduff Hughes and Jeffrey Dean},
year={2016},
eprint={1609.08144},
archivePrefix={arXiv},
primaryClass={cs.CL}
}
```
## Further References
- This Hugging Face implementation uses the [nltk.translate.gleu_score implementation](https://www.nltk.org/_modules/nltk/translate/gleu_score.html)
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/google_bleu/google_bleu.py
|
# Copyright 2020 The HuggingFace Datasets Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
""" Google BLEU (aka GLEU) metric. """
from typing import Dict, List
from nltk.translate import gleu_score
import datasets
from datasets import MetricInfo
_CITATION = """\
@misc{wu2016googles,
title={Google's Neural Machine Translation System: Bridging the Gap between Human and Machine Translation},
author={Yonghui Wu and Mike Schuster and Zhifeng Chen and Quoc V. Le and Mohammad Norouzi and Wolfgang Macherey
and Maxim Krikun and Yuan Cao and Qin Gao and Klaus Macherey and Jeff Klingner and Apurva Shah and Melvin
Johnson and Xiaobing Liu and Łukasz Kaiser and Stephan Gouws and Yoshikiyo Kato and Taku Kudo and Hideto
Kazawa and Keith Stevens and George Kurian and Nishant Patil and Wei Wang and Cliff Young and
Jason Smith and Jason Riesa and Alex Rudnick and Oriol Vinyals and Greg Corrado and Macduff Hughes
and Jeffrey Dean},
year={2016},
eprint={1609.08144},
archivePrefix={arXiv},
primaryClass={cs.CL}
}
"""
_DESCRIPTION = """\
The BLEU score has some undesirable properties when used for single
sentences, as it was designed to be a corpus measure. We therefore
use a slightly different score for our RL experiments which we call
the 'GLEU score'. For the GLEU score, we record all sub-sequences of
1, 2, 3 or 4 tokens in output and target sequence (n-grams). We then
compute a recall, which is the ratio of the number of matching n-grams
to the number of total n-grams in the target (ground truth) sequence,
and a precision, which is the ratio of the number of matching n-grams
to the number of total n-grams in the generated output sequence. Then
GLEU score is simply the minimum of recall and precision. This GLEU
score's range is always between 0 (no matches) and 1 (all match) and
it is symmetrical when switching output and target. According to
our experiments, GLEU score correlates quite well with the BLEU
metric on a corpus level but does not have its drawbacks for our per
sentence reward objective.
"""
_KWARGS_DESCRIPTION = """\
Computes corpus-level Google BLEU (GLEU) score of translated segments against one or more references.
Instead of averaging the sentence level GLEU scores (i.e. macro-average precision), Wu et al. (2016) sum up the matching
tokens and the max of hypothesis and reference tokens for each sentence, then compute using the aggregate values.
Args:
predictions (list of str): list of translations to score.
Each translation should be tokenized into a list of tokens.
references (list of list of str): list of lists of references for each translation.
Each reference should be tokenized into a list of tokens.
min_len (int): The minimum order of n-gram this function should extract. Defaults to 1.
max_len (int): The maximum order of n-gram this function should extract. Defaults to 4.
Returns:
'google_bleu': google_bleu score
Examples:
Example 1:
>>> hyp1 = ['It', 'is', 'a', 'guide', 'to', 'action', 'which',
... 'ensures', 'that', 'the', 'rubber', 'duck', 'always',
... 'disobeys', 'the', 'commands', 'of', 'the', 'cat']
>>> ref1a = ['It', 'is', 'the', 'guiding', 'principle', 'which',
... 'guarantees', 'the', 'rubber', 'duck', 'forces', 'never',
... 'being', 'under', 'the', 'command', 'of', 'the', 'cat']
>>> hyp2 = ['he', 'read', 'the', 'book', 'because', 'he', 'was',
... 'interested', 'in', 'world', 'history']
>>> ref2a = ['he', 'was', 'interested', 'in', 'world', 'history',
... 'because', 'he', 'read', 'the', 'book']
>>> list_of_references = [[ref1a], [ref2a]]
>>> hypotheses = [hyp1, hyp2]
>>> google_bleu = datasets.load_metric("google_bleu")
>>> results = google_bleu.compute(predictions=hypotheses, references=list_of_references)
>>> print(round(results["google_bleu"], 2))
0.44
Example 2:
>>> hyp1 = ['It', 'is', 'a', 'guide', 'to', 'action', 'which',
... 'ensures', 'that', 'the', 'rubber', 'duck', 'always',
... 'disobeys', 'the', 'commands', 'of', 'the', 'cat']
>>> ref1a = ['It', 'is', 'the', 'guiding', 'principle', 'which',
... 'guarantees', 'the', 'rubber', 'duck', 'forces', 'never',
... 'being', 'under', 'the', 'command', 'of', 'the', 'cat']
>>> ref1b = ['It', 'is', 'a', 'guide', 'to', 'action', 'that',
... 'ensures', 'that', 'the', 'rubber', 'duck', 'will', 'never',
... 'heed', 'the', 'cat', 'commands']
>>> ref1c = ['It', 'is', 'the', 'practical', 'guide', 'for', 'the',
... 'rubber', 'duck', 'army', 'never', 'to', 'heed', 'the', 'directions',
... 'of', 'the', 'cat']
>>> hyp2 = ['he', 'read', 'the', 'book', 'because', 'he', 'was',
... 'interested', 'in', 'world', 'history']
>>> ref2a = ['he', 'was', 'interested', 'in', 'world', 'history',
... 'because', 'he', 'read', 'the', 'book']
>>> list_of_references = [[ref1a, ref1b, ref1c], [ref2a]]
>>> hypotheses = [hyp1, hyp2]
>>> google_bleu = datasets.load_metric("google_bleu")
>>> results = google_bleu.compute(predictions=hypotheses, references=list_of_references)
>>> print(round(results["google_bleu"], 2))
0.61
Example 3:
>>> hyp1 = ['It', 'is', 'a', 'guide', 'to', 'action', 'which',
... 'ensures', 'that', 'the', 'rubber', 'duck', 'always',
... 'disobeys', 'the', 'commands', 'of', 'the', 'cat']
>>> ref1a = ['It', 'is', 'the', 'guiding', 'principle', 'which',
... 'guarantees', 'the', 'rubber', 'duck', 'forces', 'never',
... 'being', 'under', 'the', 'command', 'of', 'the', 'cat']
>>> ref1b = ['It', 'is', 'a', 'guide', 'to', 'action', 'that',
... 'ensures', 'that', 'the', 'rubber', 'duck', 'will', 'never',
... 'heed', 'the', 'cat', 'commands']
>>> ref1c = ['It', 'is', 'the', 'practical', 'guide', 'for', 'the',
... 'rubber', 'duck', 'army', 'never', 'to', 'heed', 'the', 'directions',
... 'of', 'the', 'cat']
>>> hyp2 = ['he', 'read', 'the', 'book', 'because', 'he', 'was',
... 'interested', 'in', 'world', 'history']
>>> ref2a = ['he', 'was', 'interested', 'in', 'world', 'history',
... 'because', 'he', 'read', 'the', 'book']
>>> list_of_references = [[ref1a, ref1b, ref1c], [ref2a]]
>>> hypotheses = [hyp1, hyp2]
>>> google_bleu = datasets.load_metric("google_bleu")
>>> results = google_bleu.compute(predictions=hypotheses, references=list_of_references, min_len=2)
>>> print(round(results["google_bleu"], 2))
0.53
Example 4:
>>> hyp1 = ['It', 'is', 'a', 'guide', 'to', 'action', 'which',
... 'ensures', 'that', 'the', 'rubber', 'duck', 'always',
... 'disobeys', 'the', 'commands', 'of', 'the', 'cat']
>>> ref1a = ['It', 'is', 'the', 'guiding', 'principle', 'which',
... 'guarantees', 'the', 'rubber', 'duck', 'forces', 'never',
... 'being', 'under', 'the', 'command', 'of', 'the', 'cat']
>>> ref1b = ['It', 'is', 'a', 'guide', 'to', 'action', 'that',
... 'ensures', 'that', 'the', 'rubber', 'duck', 'will', 'never',
... 'heed', 'the', 'cat', 'commands']
>>> ref1c = ['It', 'is', 'the', 'practical', 'guide', 'for', 'the',
... 'rubber', 'duck', 'army', 'never', 'to', 'heed', 'the', 'directions',
... 'of', 'the', 'cat']
>>> hyp2 = ['he', 'read', 'the', 'book', 'because', 'he', 'was',
... 'interested', 'in', 'world', 'history']
>>> ref2a = ['he', 'was', 'interested', 'in', 'world', 'history',
... 'because', 'he', 'read', 'the', 'book']
>>> list_of_references = [[ref1a, ref1b, ref1c], [ref2a]]
>>> hypotheses = [hyp1, hyp2]
>>> google_bleu = datasets.load_metric("google_bleu")
>>> results = google_bleu.compute(predictions=hypotheses,references=list_of_references, min_len=2, max_len=6)
>>> print(round(results["google_bleu"], 2))
0.4
"""
@datasets.utils.file_utils.add_start_docstrings(_DESCRIPTION, _KWARGS_DESCRIPTION)
class GoogleBleu(datasets.Metric):
def _info(self) -> MetricInfo:
return datasets.MetricInfo(
description=_DESCRIPTION,
citation=_CITATION,
inputs_description=_KWARGS_DESCRIPTION,
features=datasets.Features(
{
"predictions": datasets.Sequence(datasets.Value("string", id="token"), id="sequence"),
"references": datasets.Sequence(
datasets.Sequence(datasets.Value("string", id="token"), id="sequence"), id="references"
),
}
),
)
def _compute(
self,
predictions: List[List[List[str]]],
references: List[List[str]],
min_len: int = 1,
max_len: int = 4,
) -> Dict[str, float]:
return {
"google_bleu": gleu_score.corpus_gleu(
list_of_references=references, hypotheses=predictions, min_len=min_len, max_len=max_len
)
}
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/xtreme_s/README.md
|
# Metric Card for XTREME-S
## Metric Description
The XTREME-S metric aims to evaluate model performance on the Cross-lingual TRansfer Evaluation of Multilingual Encoders for Speech (XTREME-S) benchmark.
This benchmark was designed to evaluate speech representations across languages, tasks, domains and data regimes. It covers 102 languages from 10+ language families, 3 different domains and 4 task families: speech recognition, translation, classification and retrieval.
## How to Use
There are two steps: (1) loading the XTREME-S metric relevant to the subset of the benchmark being used for evaluation; and (2) calculating the metric.
1. **Loading the relevant XTREME-S metric** : the subsets of XTREME-S are the following: `mls`, `voxpopuli`, `covost2`, `fleurs-asr`, `fleurs-lang_id`, `minds14` and `babel`. More information about the different subsets can be found on the [XTREME-S benchmark page](https://huggingface.co/datasets/google/xtreme_s).
```python
>>> from datasets import load_metric
>>> xtreme_s_metric = datasets.load_metric('xtreme_s', 'mls')
```
2. **Calculating the metric**: the metric takes two inputs :
- `predictions`: a list of predictions to score, with each prediction a `str`.
- `references`: a list of lists of references for each translation, with each reference a `str`.
```python
>>> references = ["it is sunny here", "paper and pen are essentials"]
>>> predictions = ["it's sunny", "paper pen are essential"]
>>> results = xtreme_s_metric.compute(predictions=predictions, references=references)
```
It also has two optional arguments:
- `bleu_kwargs`: a `dict` of keywords to be passed when computing the `bleu` metric for the `covost2` subset. Keywords can be one of `smooth_method`, `smooth_value`, `force`, `lowercase`, `tokenize`, `use_effective_order`.
- `wer_kwargs`: optional dict of keywords to be passed when computing `wer` and `cer`, which are computed for the `mls`, `fleurs-asr`, `voxpopuli`, and `babel` subsets. Keywords are `concatenate_texts`.
## Output values
The output of the metric depends on the XTREME-S subset chosen, consisting of a dictionary that contains one or several of the following metrics:
- `accuracy`: the proportion of correct predictions among the total number of cases processed, with a range between 0 and 1 (see [accuracy](https://huggingface.co/metrics/accuracy) for more information). This is returned for the `fleurs-lang_id` and `minds14` subsets.
- `f1`: the harmonic mean of the precision and recall (see [F1 score](https://huggingface.co/metrics/f1) for more information). Its range is 0-1 -- its lowest possible value is 0, if either the precision or the recall is 0, and its highest possible value is 1.0, which means perfect precision and recall. It is returned for the `minds14` subset.
- `wer`: Word error rate (WER) is a common metric of the performance of an automatic speech recognition system. The lower the value, the better the performance of the ASR system, with a WER of 0 being a perfect score (see [WER score](https://huggingface.co/metrics/wer) for more information). It is returned for the `mls`, `fleurs-asr`, `voxpopuli` and `babel` subsets of the benchmark.
- `cer`: Character error rate (CER) is similar to WER, but operates on character instead of word. The lower the CER value, the better the performance of the ASR system, with a CER of 0 being a perfect score (see [CER score](https://huggingface.co/metrics/cer) for more information). It is returned for the `mls`, `fleurs-asr`, `voxpopuli` and `babel` subsets of the benchmark.
- `bleu`: the BLEU score, calculated according to the SacreBLEU metric approach. It can take any value between 0.0 and 100.0, inclusive, with higher values being better (see [SacreBLEU](https://huggingface.co/metrics/sacrebleu) for more details). This is returned for the `covost2` subset.
### Values from popular papers
The [original XTREME-S paper](https://arxiv.org/pdf/2203.10752.pdf) reported average WERs ranging from 9.2 to 14.6, a BLEU score of 20.6, an accuracy of 73.3 and F1 score of 86.9, depending on the subsets of the dataset tested on.
## Examples
For the `mls` subset (which outputs `wer` and `cer`):
```python
>>> from datasets import load_metric
>>> xtreme_s_metric = datasets.load_metric('xtreme_s', 'mls')
>>> references = ["it is sunny here", "paper and pen are essentials"]
>>> predictions = ["it's sunny", "paper pen are essential"]
>>> results = xtreme_s_metric.compute(predictions=predictions, references=references)
>>> print({k: round(v, 2) for k, v in results.items()})
{'wer': 0.56, 'cer': 0.27}
```
For the `covost2` subset (which outputs `bleu`):
```python
>>> from datasets import load_metric
>>> xtreme_s_metric = datasets.load_metric('xtreme_s', 'covost2')
>>> references = ["bonjour paris", "il est necessaire de faire du sport de temps en temp"]
>>> predictions = ["bonjour paris", "il est important de faire du sport souvent"]
>>> results = xtreme_s_metric.compute(predictions=predictions, references=references)
>>> print({k: round(v, 2) for k, v in results.items()})
{'bleu': 31.65}
```
For the `fleurs-lang_id` subset (which outputs `accuracy`):
```python
>>> from datasets import load_metric
>>> xtreme_s_metric = datasets.load_metric('xtreme_s', 'fleurs-lang_id')
>>> references = [0, 1, 0, 0, 1]
>>> predictions = [0, 1, 1, 0, 0]
>>> results = xtreme_s_metric.compute(predictions=predictions, references=references)
>>> print({k: round(v, 2) for k, v in results.items()})
{'accuracy': 0.6}
```
For the `minds14` subset (which outputs `f1` and `accuracy`):
```python
>>> from datasets import load_metric
>>> xtreme_s_metric = datasets.load_metric('xtreme_s', 'minds14')
>>> references = [0, 1, 0, 0, 1]
>>> predictions = [0, 1, 1, 0, 0]
>>> results = xtreme_s_metric.compute(predictions=predictions, references=references)
>>> print({k: round(v, 2) for k, v in results.items()})
{'f1': 0.58, 'accuracy': 0.6}
```
## Limitations and bias
This metric works only with datasets that have the same format as the [XTREME-S dataset](https://huggingface.co/datasets/google/xtreme_s).
While the XTREME-S dataset is meant to represent a variety of languages and tasks, it has inherent biases: it is missing many languages that are important and under-represented in NLP datasets.
It also has a particular focus on read-speech because common evaluation benchmarks like CoVoST-2 or LibriSpeech evaluate on this type of speech, which results in a mismatch between performance obtained in a read-speech setting and a more noisy setting (in production or live deployment, for instance).
## Citation
```bibtex
@article{conneau2022xtreme,
title={XTREME-S: Evaluating Cross-lingual Speech Representations},
author={Conneau, Alexis and Bapna, Ankur and Zhang, Yu and Ma, Min and von Platen, Patrick and Lozhkov, Anton and Cherry, Colin and Jia, Ye and Rivera, Clara and Kale, Mihir and others},
journal={arXiv preprint arXiv:2203.10752},
year={2022}
}
```
## Further References
- [XTREME-S dataset](https://huggingface.co/datasets/google/xtreme_s)
- [XTREME-S github repository](https://github.com/google-research/xtreme)
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/xtreme_s/xtreme_s.py
|
# Copyright 2022 The HuggingFace Datasets Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
""" XTREME-S benchmark metric. """
from typing import List
from packaging import version
from sklearn.metrics import f1_score
import datasets
from datasets.config import PY_VERSION
if PY_VERSION < version.parse("3.8"):
import importlib_metadata
else:
import importlib.metadata as importlib_metadata
# TODO(Patrick/Anton)
_CITATION = """\
"""
_DESCRIPTION = """\
XTREME-S is a benchmark to evaluate universal cross-lingual speech representations in many languages.
XTREME-S covers four task families: speech recognition, classification, speech-to-text translation and retrieval.
"""
_KWARGS_DESCRIPTION = """
Compute XTREME-S evaluation metric associated to each XTREME-S dataset.
Args:
predictions: list of predictions to score.
Each translation should be tokenized into a list of tokens.
references: list of lists of references for each translation.
Each reference should be tokenized into a list of tokens.
bleu_kwargs: optional dict of keywords to be passed when computing 'bleu'.
Keywords include Dict can be one of 'smooth_method', 'smooth_value', 'force', 'lowercase',
'tokenize', 'use_effective_order'.
wer_kwargs: optional dict of keywords to be passed when computing 'wer' and 'cer'.
Keywords include 'concatenate_texts'.
Returns: depending on the XTREME-S task, one or several of:
"accuracy": Accuracy - for 'fleurs-lang_id', 'minds14'
"f1": F1 score - for 'minds14'
"wer": Word error rate - for 'mls', 'fleurs-asr', 'voxpopuli', 'babel'
"cer": Character error rate - for 'mls', 'fleurs-asr', 'voxpopuli', 'babel'
"bleu": BLEU score according to the `sacrebleu` metric - for 'covost2'
Examples:
>>> xtreme_s_metric = datasets.load_metric('xtreme_s', 'mls') # 'mls', 'voxpopuli', 'fleurs-asr' or 'babel'
>>> references = ["it is sunny here", "paper and pen are essentials"]
>>> predictions = ["it's sunny", "paper pen are essential"]
>>> results = xtreme_s_metric.compute(predictions=predictions, references=references)
>>> print({k: round(v, 2) for k, v in results.items()})
{'wer': 0.56, 'cer': 0.27}
>>> xtreme_s_metric = datasets.load_metric('xtreme_s', 'covost2')
>>> references = ["bonjour paris", "il est necessaire de faire du sport de temps en temp"]
>>> predictions = ["bonjour paris", "il est important de faire du sport souvent"]
>>> results = xtreme_s_metric.compute(predictions=predictions, references=references)
>>> print({k: round(v, 2) for k, v in results.items()})
{'bleu': 31.65}
>>> xtreme_s_metric = datasets.load_metric('xtreme_s', 'fleurs-lang_id')
>>> references = [0, 1, 0, 0, 1]
>>> predictions = [0, 1, 1, 0, 0]
>>> results = xtreme_s_metric.compute(predictions=predictions, references=references)
>>> print({k: round(v, 2) for k, v in results.items()})
{'accuracy': 0.6}
>>> xtreme_s_metric = datasets.load_metric('xtreme_s', 'minds14')
>>> references = [0, 1, 0, 0, 1]
>>> predictions = [0, 1, 1, 0, 0]
>>> results = xtreme_s_metric.compute(predictions=predictions, references=references)
>>> print({k: round(v, 2) for k, v in results.items()})
{'f1': 0.58, 'accuracy': 0.6}
"""
_CONFIG_NAMES = ["fleurs-asr", "mls", "voxpopuli", "babel", "covost2", "fleurs-lang_id", "minds14"]
SENTENCE_DELIMITER = ""
try:
from jiwer import transforms as tr
_jiwer_available = True
except ImportError:
_jiwer_available = False
if _jiwer_available and version.parse(importlib_metadata.version("jiwer")) < version.parse("2.3.0"):
class SentencesToListOfCharacters(tr.AbstractTransform):
def __init__(self, sentence_delimiter: str = " "):
self.sentence_delimiter = sentence_delimiter
def process_string(self, s: str):
return list(s)
def process_list(self, inp: List[str]):
chars = []
for sent_idx, sentence in enumerate(inp):
chars.extend(self.process_string(sentence))
if self.sentence_delimiter is not None and self.sentence_delimiter != "" and sent_idx < len(inp) - 1:
chars.append(self.sentence_delimiter)
return chars
cer_transform = tr.Compose(
[tr.RemoveMultipleSpaces(), tr.Strip(), SentencesToListOfCharacters(SENTENCE_DELIMITER)]
)
elif _jiwer_available:
cer_transform = tr.Compose(
[
tr.RemoveMultipleSpaces(),
tr.Strip(),
tr.ReduceToSingleSentence(SENTENCE_DELIMITER),
tr.ReduceToListOfListOfChars(),
]
)
else:
cer_transform = None
def simple_accuracy(preds, labels):
return float((preds == labels).mean())
def f1_and_simple_accuracy(preds, labels):
return {
"f1": float(f1_score(y_true=labels, y_pred=preds, average="macro")),
"accuracy": simple_accuracy(preds, labels),
}
def bleu(
preds,
labels,
smooth_method="exp",
smooth_value=None,
force=False,
lowercase=False,
tokenize=None,
use_effective_order=False,
):
# xtreme-s can only have one label
labels = [[label] for label in labels]
preds = list(preds)
try:
import sacrebleu as scb
except ImportError:
raise ValueError(
"sacrebleu has to be installed in order to apply the bleu metric for covost2."
"You can install it via `pip install sacrebleu`."
)
if version.parse(scb.__version__) < version.parse("1.4.12"):
raise ImportWarning(
"To use `sacrebleu`, the module `sacrebleu>=1.4.12` is required, and the current version of `sacrebleu` doesn't match this condition.\n"
'You can install it with `pip install "sacrebleu>=1.4.12"`.'
)
references_per_prediction = len(labels[0])
if any(len(refs) != references_per_prediction for refs in labels):
raise ValueError("Sacrebleu requires the same number of references for each prediction")
transformed_references = [[refs[i] for refs in labels] for i in range(references_per_prediction)]
output = scb.corpus_bleu(
preds,
transformed_references,
smooth_method=smooth_method,
smooth_value=smooth_value,
force=force,
lowercase=lowercase,
use_effective_order=use_effective_order,
**({"tokenize": tokenize} if tokenize else {}),
)
return {"bleu": output.score}
def wer_and_cer(preds, labels, concatenate_texts, config_name):
try:
from jiwer import compute_measures
except ImportError:
raise ValueError(
f"jiwer has to be installed in order to apply the wer metric for {config_name}."
"You can install it via `pip install jiwer`."
)
if concatenate_texts:
wer = compute_measures(labels, preds)["wer"]
cer = compute_measures(labels, preds, truth_transform=cer_transform, hypothesis_transform=cer_transform)["wer"]
return {"wer": wer, "cer": cer}
else:
def compute_score(preds, labels, score_type="wer"):
incorrect = 0
total = 0
for prediction, reference in zip(preds, labels):
if score_type == "wer":
measures = compute_measures(reference, prediction)
elif score_type == "cer":
measures = compute_measures(
reference, prediction, truth_transform=cer_transform, hypothesis_transform=cer_transform
)
incorrect += measures["substitutions"] + measures["deletions"] + measures["insertions"]
total += measures["substitutions"] + measures["deletions"] + measures["hits"]
return incorrect / total
return {"wer": compute_score(preds, labels, "wer"), "cer": compute_score(preds, labels, "cer")}
@datasets.utils.file_utils.add_start_docstrings(_DESCRIPTION, _KWARGS_DESCRIPTION)
class XtremeS(datasets.Metric):
def _info(self):
if self.config_name not in _CONFIG_NAMES:
raise KeyError(f"You should supply a configuration name selected in {_CONFIG_NAMES}")
pred_type = "int64" if self.config_name in ["fleurs-lang_id", "minds14"] else "string"
return datasets.MetricInfo(
description=_DESCRIPTION,
citation=_CITATION,
inputs_description=_KWARGS_DESCRIPTION,
features=datasets.Features(
{"predictions": datasets.Value(pred_type), "references": datasets.Value(pred_type)}
),
codebase_urls=[],
reference_urls=[],
format="numpy",
)
def _compute(self, predictions, references, bleu_kwargs=None, wer_kwargs=None):
bleu_kwargs = bleu_kwargs if bleu_kwargs is not None else {}
wer_kwargs = wer_kwargs if wer_kwargs is not None else {}
if self.config_name == "fleurs-lang_id":
return {"accuracy": simple_accuracy(predictions, references)}
elif self.config_name == "minds14":
return f1_and_simple_accuracy(predictions, references)
elif self.config_name == "covost2":
smooth_method = bleu_kwargs.pop("smooth_method", "exp")
smooth_value = bleu_kwargs.pop("smooth_value", None)
force = bleu_kwargs.pop("force", False)
lowercase = bleu_kwargs.pop("lowercase", False)
tokenize = bleu_kwargs.pop("tokenize", None)
use_effective_order = bleu_kwargs.pop("use_effective_order", False)
return bleu(
preds=predictions,
labels=references,
smooth_method=smooth_method,
smooth_value=smooth_value,
force=force,
lowercase=lowercase,
tokenize=tokenize,
use_effective_order=use_effective_order,
)
elif self.config_name in ["fleurs-asr", "mls", "voxpopuli", "babel"]:
concatenate_texts = wer_kwargs.pop("concatenate_texts", False)
return wer_and_cer(predictions, references, concatenate_texts, self.config_name)
else:
raise KeyError(f"You should supply a configuration name selected in {_CONFIG_NAMES}")
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/mse/README.md
|
# Metric Card for MSE
## Metric Description
Mean Squared Error(MSE) represents the average of the squares of errors -- i.e. the average squared difference between the estimated values and the actual values.
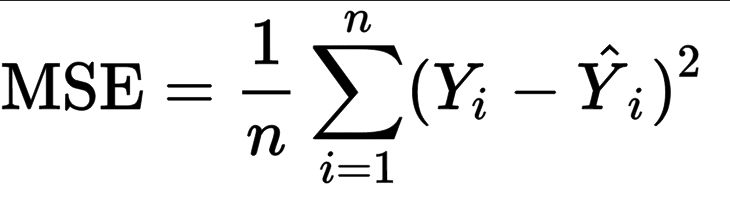
## How to Use
At minimum, this metric requires predictions and references as inputs.
```python
>>> mse_metric = datasets.load_metric("mse")
>>> predictions = [2.5, 0.0, 2, 8]
>>> references = [3, -0.5, 2, 7]
>>> results = mse_metric.compute(predictions=predictions, references=references)
```
### Inputs
Mandatory inputs:
- `predictions`: numeric array-like of shape (`n_samples,`) or (`n_samples`, `n_outputs`), representing the estimated target values.
- `references`: numeric array-like of shape (`n_samples,`) or (`n_samples`, `n_outputs`), representing the ground truth (correct) target values.
Optional arguments:
- `sample_weight`: numeric array-like of shape (`n_samples,`) representing sample weights. The default is `None`.
- `multioutput`: `raw_values`, `uniform_average` or numeric array-like of shape (`n_outputs,`), which defines the aggregation of multiple output values. The default value is `uniform_average`.
- `raw_values` returns a full set of errors in case of multioutput input.
- `uniform_average` means that the errors of all outputs are averaged with uniform weight.
- the array-like value defines weights used to average errors.
- `squared` (`bool`): If `True` returns MSE value, if `False` returns RMSE (Root Mean Squared Error). The default value is `True`.
### Output Values
This metric outputs a dictionary, containing the mean squared error score, which is of type:
- `float`: if multioutput is `uniform_average` or an ndarray of weights, then the weighted average of all output errors is returned.
- numeric array-like of shape (`n_outputs,`): if multioutput is `raw_values`, then the score is returned for each output separately.
Each MSE `float` value ranges from `0.0` to `1.0`, with the best value being `0.0`.
Output Example(s):
```python
{'mse': 0.5}
```
If `multioutput="raw_values"`:
```python
{'mse': array([0.41666667, 1. ])}
```
#### Values from Popular Papers
### Examples
Example with the `uniform_average` config:
```python
>>> from datasets import load_metric
>>> mse_metric = load_metric("mse")
>>> predictions = [2.5, 0.0, 2, 8]
>>> references = [3, -0.5, 2, 7]
>>> results = mse_metric.compute(predictions=predictions, references=references)
>>> print(results)
{'mse': 0.375}
```
Example with `squared = True`, which returns the RMSE:
```python
>>> from datasets import load_metric
>>> mse_metric = load_metric("mse")
>>> predictions = [2.5, 0.0, 2, 8]
>>> references = [3, -0.5, 2, 7]
>>> rmse_result = mse_metric.compute(predictions=predictions, references=references, squared=False)
>>> print(rmse_result)
{'mse': 0.6123724356957945}
```
Example with multi-dimensional lists, and the `raw_values` config:
```python
>>> from datasets import load_metric
>>> mse_metric = load_metric("mse", "multilist")
>>> predictions = [[0.5, 1], [-1, 1], [7, -6]]
>>> references = [[0, 2], [-1, 2], [8, -5]]
>>> results = mse_metric.compute(predictions=predictions, references=references, multioutput='raw_values')
>>> print(results)
{'mse': array([0.41666667, 1. ])}
"""
```
## Limitations and Bias
MSE has the disadvantage of heavily weighting outliers -- given that it squares them, this results in large errors weighing more heavily than small ones. It can be used alongside [MAE](https://huggingface.co/metrics/mae), which is complementary given that it does not square the errors.
## Citation(s)
```bibtex
@article{scikit-learn,
title={Scikit-learn: Machine Learning in {P}ython},
author={Pedregosa, F. and Varoquaux, G. and Gramfort, A. and Michel, V.
and Thirion, B. and Grisel, O. and Blondel, M. and Prettenhofer, P.
and Weiss, R. and Dubourg, V. and Vanderplas, J. and Passos, A. and
Cournapeau, D. and Brucher, M. and Perrot, M. and Duchesnay, E.},
journal={Journal of Machine Learning Research},
volume={12},
pages={2825--2830},
year={2011}
}
```
```bibtex
@article{willmott2005advantages,
title={Advantages of the mean absolute error (MAE) over the root mean square error (RMSE) in assessing average model performance},
author={Willmott, Cort J and Matsuura, Kenji},
journal={Climate research},
volume={30},
number={1},
pages={79--82},
year={2005}
}
```
## Further References
- [Mean Squared Error - Wikipedia](https://en.wikipedia.org/wiki/Mean_squared_error)
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/mse/mse.py
|
# Copyright 2022 The HuggingFace Datasets Authors and the current dataset script contributor.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""MSE - Mean Squared Error Metric"""
from sklearn.metrics import mean_squared_error
import datasets
_CITATION = """\
@article{scikit-learn,
title={Scikit-learn: Machine Learning in {P}ython},
author={Pedregosa, F. and Varoquaux, G. and Gramfort, A. and Michel, V.
and Thirion, B. and Grisel, O. and Blondel, M. and Prettenhofer, P.
and Weiss, R. and Dubourg, V. and Vanderplas, J. and Passos, A. and
Cournapeau, D. and Brucher, M. and Perrot, M. and Duchesnay, E.},
journal={Journal of Machine Learning Research},
volume={12},
pages={2825--2830},
year={2011}
}
"""
_DESCRIPTION = """\
Mean Squared Error(MSE) is the average of the square of difference between the predicted
and actual values.
"""
_KWARGS_DESCRIPTION = """
Args:
predictions: array-like of shape (n_samples,) or (n_samples, n_outputs)
Estimated target values.
references: array-like of shape (n_samples,) or (n_samples, n_outputs)
Ground truth (correct) target values.
sample_weight: array-like of shape (n_samples,), default=None
Sample weights.
multioutput: {"raw_values", "uniform_average"} or array-like of shape (n_outputs,), default="uniform_average"
Defines aggregating of multiple output values. Array-like value defines weights used to average errors.
"raw_values" : Returns a full set of errors in case of multioutput input.
"uniform_average" : Errors of all outputs are averaged with uniform weight.
squared : bool, default=True
If True returns MSE value, if False returns RMSE (Root Mean Squared Error) value.
Returns:
mse : mean squared error.
Examples:
>>> mse_metric = datasets.load_metric("mse")
>>> predictions = [2.5, 0.0, 2, 8]
>>> references = [3, -0.5, 2, 7]
>>> results = mse_metric.compute(predictions=predictions, references=references)
>>> print(results)
{'mse': 0.375}
>>> rmse_result = mse_metric.compute(predictions=predictions, references=references, squared=False)
>>> print(rmse_result)
{'mse': 0.6123724356957945}
If you're using multi-dimensional lists, then set the config as follows :
>>> mse_metric = datasets.load_metric("mse", "multilist")
>>> predictions = [[0.5, 1], [-1, 1], [7, -6]]
>>> references = [[0, 2], [-1, 2], [8, -5]]
>>> results = mse_metric.compute(predictions=predictions, references=references)
>>> print(results)
{'mse': 0.7083333333333334}
>>> results = mse_metric.compute(predictions=predictions, references=references, multioutput='raw_values')
>>> print(results) # doctest: +NORMALIZE_WHITESPACE
{'mse': array([0.41666667, 1. ])}
"""
@datasets.utils.file_utils.add_start_docstrings(_DESCRIPTION, _KWARGS_DESCRIPTION)
class Mse(datasets.Metric):
def _info(self):
return datasets.MetricInfo(
description=_DESCRIPTION,
citation=_CITATION,
inputs_description=_KWARGS_DESCRIPTION,
features=datasets.Features(self._get_feature_types()),
reference_urls=[
"https://scikit-learn.org/stable/modules/generated/sklearn.metrics.mean_squared_error.html"
],
)
def _get_feature_types(self):
if self.config_name == "multilist":
return {
"predictions": datasets.Sequence(datasets.Value("float")),
"references": datasets.Sequence(datasets.Value("float")),
}
else:
return {
"predictions": datasets.Value("float"),
"references": datasets.Value("float"),
}
def _compute(self, predictions, references, sample_weight=None, multioutput="uniform_average", squared=True):
mse = mean_squared_error(
references, predictions, sample_weight=sample_weight, multioutput=multioutput, squared=squared
)
return {"mse": mse}
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/code_eval/README.md
|
# Metric Card for Code Eval
## Metric description
The CodeEval metric estimates the pass@k metric for code synthesis.
It implements the evaluation harness for the HumanEval problem solving dataset described in the paper ["Evaluating Large Language Models Trained on Code"](https://arxiv.org/abs/2107.03374).
## How to use
The Code Eval metric calculates how good are predictions given a set of references. Its arguments are:
`predictions`: a list of candidates to evaluate. Each candidate should be a list of strings with several code candidates to solve the problem.
`references`: a list with a test for each prediction. Each test should evaluate the correctness of a code candidate.
`k`: number of code candidates to consider in the evaluation. The default value is `[1, 10, 100]`.
`num_workers`: the number of workers used to evaluate the candidate programs (The default value is `4`).
`timeout`: The maximum time taken to produce a prediction before it is considered a "timeout". The default value is `3.0` (i.e. 3 seconds).
```python
from datasets import load_metric
code_eval = load_metric("code_eval")
test_cases = ["assert add(2,3)==5"]
candidates = [["def add(a,b): return a*b", "def add(a, b): return a+b"]]
pass_at_k, results = code_eval.compute(references=test_cases, predictions=candidates, k=[1, 2])
```
N.B.
This metric exists to run untrusted model-generated code. Users are strongly encouraged not to do so outside of a robust security sandbox. Before running this metric and once you've taken the necessary precautions, you will need to set the `HF_ALLOW_CODE_EVAL` environment variable. Use it at your own risk:
```python
import os
os.environ["HF_ALLOW_CODE_EVAL"] = "1"`
```
## Output values
The Code Eval metric outputs two things:
`pass_at_k`: a dictionary with the pass rates for each k value defined in the arguments.
`results`: a dictionary with granular results of each unit test.
### Values from popular papers
The [original CODEX paper](https://arxiv.org/pdf/2107.03374.pdf) reported that the CODEX-12B model had a pass@k score of 28.8% at `k=1`, 46.8% at `k=10` and 72.3% at `k=100`. However, since the CODEX model is not open source, it is hard to verify these numbers.
## Examples
Full match at `k=1`:
```python
from datasets import load_metric
code_eval = load_metric("code_eval")
test_cases = ["assert add(2,3)==5"]
candidates = [["def add(a, b): return a+b"]]
pass_at_k, results = code_eval.compute(references=test_cases, predictions=candidates, k=[1])
print(pass_at_k)
{'pass@1': 1.0}
```
No match for k = 1:
```python
from datasets import load_metric
code_eval = load_metric("code_eval")
test_cases = ["assert add(2,3)==5"]
candidates = [["def add(a,b): return a*b"]]
pass_at_k, results = code_eval.compute(references=test_cases, predictions=candidates, k=[1])
print(pass_at_k)
{'pass@1': 0.0}
```
Partial match at k=1, full match at k=2:
```python
from datasets import load_metric
code_eval = load_metric("code_eval")
test_cases = ["assert add(2,3)==5"]
candidates = [["def add(a, b): return a+b", "def add(a,b): return a*b"]]
pass_at_k, results = code_eval.compute(references=test_cases, predictions=candidates, k=[1, 2])
print(pass_at_k)
{'pass@1': 0.5, 'pass@2': 1.0}
```
## Limitations and bias
As per the warning included in the metric code itself:
> This program exists to execute untrusted model-generated code. Although it is highly unlikely that model-generated code will do something overtly malicious in response to this test suite, model-generated code may act destructively due to a lack of model capability or alignment. Users are strongly encouraged to sandbox this evaluation suite so that it does not perform destructive actions on their host or network. For more information on how OpenAI sandboxes its code, see the accompanying paper. Once you have read this disclaimer and taken appropriate precautions, uncomment the following line and proceed at your own risk:
More information about the limitations of the code can be found on the [Human Eval Github repository](https://github.com/openai/human-eval).
## Citation
```bibtex
@misc{chen2021evaluating,
title={Evaluating Large Language Models Trained on Code},
author={Mark Chen and Jerry Tworek and Heewoo Jun and Qiming Yuan \
and Henrique Ponde de Oliveira Pinto and Jared Kaplan and Harri Edwards \
and Yuri Burda and Nicholas Joseph and Greg Brockman and Alex Ray \
and Raul Puri and Gretchen Krueger and Michael Petrov and Heidy Khlaaf \
and Girish Sastry and Pamela Mishkin and Brooke Chan and Scott Gray \
and Nick Ryder and Mikhail Pavlov and Alethea Power and Lukasz Kaiser \
and Mohammad Bavarian and Clemens Winter and Philippe Tillet \
and Felipe Petroski Such and Dave Cummings and Matthias Plappert \
and Fotios Chantzis and Elizabeth Barnes and Ariel Herbert-Voss \
and William Hebgen Guss and Alex Nichol and Alex Paino and Nikolas Tezak \
and Jie Tang and Igor Babuschkin and Suchir Balaji and Shantanu Jain \
and William Saunders and Christopher Hesse and Andrew N. Carr \
and Jan Leike and Josh Achiam and Vedant Misra and Evan Morikawa \
and Alec Radford and Matthew Knight and Miles Brundage and Mira Murati \
and Katie Mayer and Peter Welinder and Bob McGrew and Dario Amodei \
and Sam McCandlish and Ilya Sutskever and Wojciech Zaremba},
year={2021},
eprint={2107.03374},
archivePrefix={arXiv},
primaryClass={cs.LG}
}
```
## Further References
- [Human Eval Github repository](https://github.com/openai/human-eval)
- [OpenAI Codex website](https://openai.com/blog/openai-codex/)
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/code_eval/code_eval.py
|
# Copyright 2020 The HuggingFace Datasets Authors and the current dataset script contributor.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""The CodeEval metric estimates the pass@k metric for code synthesis.
This is an evaluation harness for the HumanEval problem solving dataset
described in the paper "Evaluating Large Language Models Trained on Code"
(https://arxiv.org/abs/2107.03374)."""
import itertools
import os
from collections import Counter, defaultdict
from concurrent.futures import ThreadPoolExecutor, as_completed
import numpy as np
import datasets
from .execute import check_correctness
_CITATION = """\
@misc{chen2021evaluating,
title={Evaluating Large Language Models Trained on Code},
author={Mark Chen and Jerry Tworek and Heewoo Jun and Qiming Yuan \
and Henrique Ponde de Oliveira Pinto and Jared Kaplan and Harri Edwards \
and Yuri Burda and Nicholas Joseph and Greg Brockman and Alex Ray \
and Raul Puri and Gretchen Krueger and Michael Petrov and Heidy Khlaaf \
and Girish Sastry and Pamela Mishkin and Brooke Chan and Scott Gray \
and Nick Ryder and Mikhail Pavlov and Alethea Power and Lukasz Kaiser \
and Mohammad Bavarian and Clemens Winter and Philippe Tillet \
and Felipe Petroski Such and Dave Cummings and Matthias Plappert \
and Fotios Chantzis and Elizabeth Barnes and Ariel Herbert-Voss \
and William Hebgen Guss and Alex Nichol and Alex Paino and Nikolas Tezak \
and Jie Tang and Igor Babuschkin and Suchir Balaji and Shantanu Jain \
and William Saunders and Christopher Hesse and Andrew N. Carr \
and Jan Leike and Josh Achiam and Vedant Misra and Evan Morikawa \
and Alec Radford and Matthew Knight and Miles Brundage and Mira Murati \
and Katie Mayer and Peter Welinder and Bob McGrew and Dario Amodei \
and Sam McCandlish and Ilya Sutskever and Wojciech Zaremba},
year={2021},
eprint={2107.03374},
archivePrefix={arXiv},
primaryClass={cs.LG}
}
"""
_DESCRIPTION = """\
This metric implements the evaluation harness for the HumanEval problem solving dataset
described in the paper "Evaluating Large Language Models Trained on Code"
(https://arxiv.org/abs/2107.03374).
"""
_KWARGS_DESCRIPTION = """
Calculates how good are predictions given some references, using certain scores
Args:
predictions: list of candidates to evaluate. Each candidates should be a list
of strings with several code candidates to solve the problem.
references: a list with a test for each prediction. Each test should evaluate the
correctness of a code candidate.
k: number of code candidates to consider in the evaluation (Default: [1, 10, 100])
num_workers: number of workers used to evaluate the canidate programs (Default: 4).
timeout:
Returns:
pass_at_k: dict with pass rates for each k
results: dict with granular results of each unittest
Examples:
>>> code_eval = datasets.load_metric("code_eval")
>>> test_cases = ["assert add(2,3)==5"]
>>> candidates = [["def add(a,b): return a*b", "def add(a, b): return a+b"]]
>>> pass_at_k, results = code_eval.compute(references=test_cases, predictions=candidates, k=[1, 2])
>>> print(pass_at_k)
{'pass@1': 0.5, 'pass@2': 1.0}
"""
_WARNING = """
################################################################################
!!!WARNING!!!
################################################################################
The "code_eval" metric executes untrusted model-generated code in Python.
Although it is highly unlikely that model-generated code will do something
overtly malicious in response to this test suite, model-generated code may act
destructively due to a lack of model capability or alignment.
Users are strongly encouraged to sandbox this evaluation suite so that it
does not perform destructive actions on their host or network. For more
information on how OpenAI sandboxes its code, see the paper "Evaluating Large
Language Models Trained on Code" (https://arxiv.org/abs/2107.03374).
Once you have read this disclaimer and taken appropriate precautions,
set the environment variable HF_ALLOW_CODE_EVAL="1". Within Python you can to this
with:
>>> import os
>>> os.environ["HF_ALLOW_CODE_EVAL"] = "1"
################################################################################\
"""
_LICENSE = """The MIT License
Copyright (c) OpenAI (https://openai.com)
Permission is hereby granted, free of charge, to any person obtaining a copy
of this software and associated documentation files (the "Software"), to deal
in the Software without restriction, including without limitation the rights
to use, copy, modify, merge, publish, distribute, sublicense, and/or sell
copies of the Software, and to permit persons to whom the Software is
furnished to do so, subject to the following conditions:
The above copyright notice and this permission notice shall be included in
all copies or substantial portions of the Software.
THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE
AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM,
OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN
THE SOFTWARE."""
@datasets.utils.file_utils.add_start_docstrings(_DESCRIPTION, _KWARGS_DESCRIPTION)
class CodeEval(datasets.Metric):
def _info(self):
return datasets.MetricInfo(
# This is the description that will appear on the metrics page.
description=_DESCRIPTION,
citation=_CITATION,
inputs_description=_KWARGS_DESCRIPTION,
# This defines the format of each prediction and reference
features=datasets.Features(
{
"predictions": datasets.Sequence(datasets.Value("string")),
"references": datasets.Value("string"),
}
),
homepage="https://github.com/openai/human-eval",
codebase_urls=["https://github.com/openai/human-eval"],
reference_urls=["https://github.com/openai/human-eval"],
license=_LICENSE,
)
def _compute(self, predictions, references, k=[1, 10, 100], num_workers=4, timeout=3.0):
"""Returns the scores"""
if os.getenv("HF_ALLOW_CODE_EVAL", 0) != "1":
raise ValueError(_WARNING)
if os.name == "nt":
raise NotImplementedError("This metric is currently not supported on Windows.")
with ThreadPoolExecutor(max_workers=num_workers) as executor:
futures = []
completion_id = Counter()
n_samples = 0
results = defaultdict(list)
for task_id, (candidates, test_case) in enumerate(zip(predictions, references)):
for candidate in candidates:
test_program = candidate + "\n" + test_case
args = (test_program, timeout, task_id, completion_id[task_id])
future = executor.submit(check_correctness, *args)
futures.append(future)
completion_id[task_id] += 1
n_samples += 1
for future in as_completed(futures):
result = future.result()
results[result["task_id"]].append((result["completion_id"], result))
total, correct = [], []
for result in results.values():
result.sort()
passed = [r[1]["passed"] for r in result]
total.append(len(passed))
correct.append(sum(passed))
total = np.array(total)
correct = np.array(correct)
ks = k
pass_at_k = {f"pass@{k}": estimate_pass_at_k(total, correct, k).mean() for k in ks if (total >= k).all()}
return pass_at_k, results
def estimate_pass_at_k(num_samples, num_correct, k):
"""Estimates pass@k of each problem and returns them in an array."""
def estimator(n: int, c: int, k: int) -> float:
"""Calculates 1 - comb(n - c, k) / comb(n, k)."""
if n - c < k:
return 1.0
return 1.0 - np.prod(1.0 - k / np.arange(n - c + 1, n + 1))
if isinstance(num_samples, int):
num_samples_it = itertools.repeat(num_samples, len(num_correct))
else:
assert len(num_samples) == len(num_correct)
num_samples_it = iter(num_samples)
return np.array([estimator(int(n), int(c), k) for n, c in zip(num_samples_it, num_correct)])
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/code_eval/execute.py
|
# Copyright 2020 The HuggingFace Datasets Authors and the current dataset script contributor.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# This code is adapted from OpenAI's release
# https://github.com/openai/human-eval/blob/master/human_eval/execution.py
import contextlib
import faulthandler
import io
import multiprocessing
import os
import platform
import signal
import tempfile
def check_correctness(check_program, timeout, task_id, completion_id):
"""
Evaluates the functional correctness of a completion by running the test
suite provided in the problem.
:param completion_id: an optional completion ID so we can match
the results later even if execution finishes asynchronously.
"""
manager = multiprocessing.Manager()
result = manager.list()
p = multiprocessing.Process(target=unsafe_execute, args=(check_program, result, timeout))
p.start()
p.join(timeout=timeout + 1)
if p.is_alive():
p.kill()
if not result:
result.append("timed out")
return {
"task_id": task_id,
"passed": result[0] == "passed",
"result": result[0],
"completion_id": completion_id,
}
def unsafe_execute(check_program, result, timeout):
with create_tempdir():
# These system calls are needed when cleaning up tempdir.
import os
import shutil
rmtree = shutil.rmtree
rmdir = os.rmdir
chdir = os.chdir
# Disable functionalities that can make destructive changes to the test.
reliability_guard()
# Run program.
try:
exec_globals = {}
with swallow_io():
with time_limit(timeout):
exec(check_program, exec_globals)
result.append("passed")
except TimeoutException:
result.append("timed out")
except BaseException as e:
result.append(f"failed: {e}")
# Needed for cleaning up.
shutil.rmtree = rmtree
os.rmdir = rmdir
os.chdir = chdir
@contextlib.contextmanager
def time_limit(seconds):
def signal_handler(signum, frame):
raise TimeoutException("Timed out!")
signal.setitimer(signal.ITIMER_REAL, seconds)
signal.signal(signal.SIGALRM, signal_handler)
try:
yield
finally:
signal.setitimer(signal.ITIMER_REAL, 0)
@contextlib.contextmanager
def swallow_io():
stream = WriteOnlyStringIO()
with contextlib.redirect_stdout(stream):
with contextlib.redirect_stderr(stream):
with redirect_stdin(stream):
yield
@contextlib.contextmanager
def create_tempdir():
with tempfile.TemporaryDirectory() as dirname:
with chdir(dirname):
yield dirname
class TimeoutException(Exception):
pass
class WriteOnlyStringIO(io.StringIO):
"""StringIO that throws an exception when it's read from"""
def read(self, *args, **kwargs):
raise OSError
def readline(self, *args, **kwargs):
raise OSError
def readlines(self, *args, **kwargs):
raise OSError
def readable(self, *args, **kwargs):
"""Returns True if the IO object can be read."""
return False
class redirect_stdin(contextlib._RedirectStream): # type: ignore
_stream = "stdin"
@contextlib.contextmanager
def chdir(root):
if root == ".":
yield
return
cwd = os.getcwd()
os.chdir(root)
try:
yield
except BaseException as exc:
raise exc
finally:
os.chdir(cwd)
def reliability_guard(maximum_memory_bytes=None):
"""
This disables various destructive functions and prevents the generated code
from interfering with the test (e.g. fork bomb, killing other processes,
removing filesystem files, etc.)
WARNING
This function is NOT a security sandbox. Untrusted code, including, model-
generated code, should not be blindly executed outside of one. See the
Codex paper for more information about OpenAI's code sandbox, and proceed
with caution.
"""
if maximum_memory_bytes is not None:
import resource
resource.setrlimit(resource.RLIMIT_AS, (maximum_memory_bytes, maximum_memory_bytes))
resource.setrlimit(resource.RLIMIT_DATA, (maximum_memory_bytes, maximum_memory_bytes))
if not platform.uname().system == "Darwin":
resource.setrlimit(resource.RLIMIT_STACK, (maximum_memory_bytes, maximum_memory_bytes))
faulthandler.disable()
import builtins
builtins.exit = None
builtins.quit = None
import os
os.environ["OMP_NUM_THREADS"] = "1"
os.kill = None
os.system = None
os.putenv = None
os.remove = None
os.removedirs = None
os.rmdir = None
os.fchdir = None
os.setuid = None
os.fork = None
os.forkpty = None
os.killpg = None
os.rename = None
os.renames = None
os.truncate = None
os.replace = None
os.unlink = None
os.fchmod = None
os.fchown = None
os.chmod = None
os.chown = None
os.chroot = None
os.fchdir = None
os.lchflags = None
os.lchmod = None
os.lchown = None
os.getcwd = None
os.chdir = None
import shutil
shutil.rmtree = None
shutil.move = None
shutil.chown = None
import subprocess
subprocess.Popen = None # type: ignore
__builtins__["help"] = None
import sys
sys.modules["ipdb"] = None
sys.modules["joblib"] = None
sys.modules["resource"] = None
sys.modules["psutil"] = None
sys.modules["tkinter"] = None
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/precision/README.md
|
# Metric Card for Precision
## Metric Description
Precision is the fraction of correctly labeled positive examples out of all of the examples that were labeled as positive. It is computed via the equation:
Precision = TP / (TP + FP)
where TP is the True positives (i.e. the examples correctly labeled as positive) and FP is the False positive examples (i.e. the examples incorrectly labeled as positive).
## How to Use
At minimum, precision takes as input a list of predicted labels, `predictions`, and a list of output labels, `references`.
```python
>>> precision_metric = datasets.load_metric("precision")
>>> results = precision_metric.compute(references=[0, 1], predictions=[0, 1])
>>> print(results)
{'precision': 1.0}
```
### Inputs
- **predictions** (`list` of `int`): Predicted class labels.
- **references** (`list` of `int`): Actual class labels.
- **labels** (`list` of `int`): The set of labels to include when `average` is not set to `'binary'`. If `average` is `None`, it should be the label order. Labels present in the data can be excluded, for example to calculate a multiclass average ignoring a majority negative class. Labels not present in the data will result in 0 components in a macro average. For multilabel targets, labels are column indices. By default, all labels in `predictions` and `references` are used in sorted order. Defaults to None.
- **pos_label** (`int`): The class to be considered the positive class, in the case where `average` is set to `binary`. Defaults to 1.
- **average** (`string`): This parameter is required for multiclass/multilabel targets. If set to `None`, the scores for each class are returned. Otherwise, this determines the type of averaging performed on the data. Defaults to `'binary'`.
- 'binary': Only report results for the class specified by `pos_label`. This is applicable only if the classes found in `predictions` and `references` are binary.
- 'micro': Calculate metrics globally by counting the total true positives, false negatives and false positives.
- 'macro': Calculate metrics for each label, and find their unweighted mean. This does not take label imbalance into account.
- 'weighted': Calculate metrics for each label, and find their average weighted by support (the number of true instances for each label). This alters `'macro'` to account for label imbalance. This option can result in an F-score that is not between precision and recall.
- 'samples': Calculate metrics for each instance, and find their average (only meaningful for multilabel classification).
- **sample_weight** (`list` of `float`): Sample weights Defaults to None.
- **zero_division** (): Sets the value to return when there is a zero division. Defaults to .
- 0: Returns 0 when there is a zero division.
- 1: Returns 1 when there is a zero division.
- 'warn': Raises warnings and then returns 0 when there is a zero division.
### Output Values
- **precision**(`float` or `array` of `float`): Precision score or list of precision scores, depending on the value passed to `average`. Minimum possible value is 0. Maximum possible value is 1. Higher values indicate that fewer negative examples were incorrectly labeled as positive, which means that, generally, higher scores are better.
Output Example(s):
```python
{'precision': 0.2222222222222222}
```
```python
{'precision': array([0.66666667, 0.0, 0.0])}
```
#### Values from Popular Papers
### Examples
Example 1-A simple binary example
```python
>>> precision_metric = datasets.load_metric("precision")
>>> results = precision_metric.compute(references=[0, 1, 0, 1, 0], predictions=[0, 0, 1, 1, 0])
>>> print(results)
{'precision': 0.5}
```
Example 2-The same simple binary example as in Example 1, but with `pos_label` set to `0`.
```python
>>> precision_metric = datasets.load_metric("precision")
>>> results = precision_metric.compute(references=[0, 1, 0, 1, 0], predictions=[0, 0, 1, 1, 0], pos_label=0)
>>> print(round(results['precision'], 2))
0.67
```
Example 3-The same simple binary example as in Example 1, but with `sample_weight` included.
```python
>>> precision_metric = datasets.load_metric("precision")
>>> results = precision_metric.compute(references=[0, 1, 0, 1, 0], predictions=[0, 0, 1, 1, 0], sample_weight=[0.9, 0.5, 3.9, 1.2, 0.3])
>>> print(results)
{'precision': 0.23529411764705882}
```
Example 4-A multiclass example, with different values for the `average` input.
```python
>>> predictions = [0, 2, 1, 0, 0, 1]
>>> references = [0, 1, 2, 0, 1, 2]
>>> results = precision_metric.compute(predictions=predictions, references=references, average='macro')
>>> print(results)
{'precision': 0.2222222222222222}
>>> results = precision_metric.compute(predictions=predictions, references=references, average='micro')
>>> print(results)
{'precision': 0.3333333333333333}
>>> results = precision_metric.compute(predictions=predictions, references=references, average='weighted')
>>> print(results)
{'precision': 0.2222222222222222}
>>> results = precision_metric.compute(predictions=predictions, references=references, average=None)
>>> print([round(res, 2) for res in results['precision']])
[0.67, 0.0, 0.0]
```
## Limitations and Bias
[Precision](https://huggingface.co/metrics/precision) and [recall](https://huggingface.co/metrics/recall) are complementary and can be used to measure different aspects of model performance -- using both of them (or an averaged measure like [F1 score](https://huggingface.co/metrics/F1) to better represent different aspects of performance. See [Wikipedia](https://en.wikipedia.org/wiki/Precision_and_recall) for more information.
## Citation(s)
```bibtex
@article{scikit-learn,
title={Scikit-learn: Machine Learning in {P}ython},
author={Pedregosa, F. and Varoquaux, G. and Gramfort, A. and Michel, V.
and Thirion, B. and Grisel, O. and Blondel, M. and Prettenhofer, P.
and Weiss, R. and Dubourg, V. and Vanderplas, J. and Passos, A. and
Cournapeau, D. and Brucher, M. and Perrot, M. and Duchesnay, E.},
journal={Journal of Machine Learning Research},
volume={12},
pages={2825--2830},
year={2011}
}
```
## Further References
- [Wikipedia -- Precision and recall](https://en.wikipedia.org/wiki/Precision_and_recall)
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/precision/precision.py
|
# Copyright 2020 The HuggingFace Datasets Authors and the current dataset script contributor.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""Precision metric."""
from sklearn.metrics import precision_score
import datasets
_DESCRIPTION = """
Precision is the fraction of correctly labeled positive examples out of all of the examples that were labeled as positive. It is computed via the equation:
Precision = TP / (TP + FP)
where TP is the True positives (i.e. the examples correctly labeled as positive) and FP is the False positive examples (i.e. the examples incorrectly labeled as positive).
"""
_KWARGS_DESCRIPTION = """
Args:
predictions (`list` of `int`): Predicted class labels.
references (`list` of `int`): Actual class labels.
labels (`list` of `int`): The set of labels to include when `average` is not set to `'binary'`. If `average` is `None`, it should be the label order. Labels present in the data can be excluded, for example to calculate a multiclass average ignoring a majority negative class. Labels not present in the data will result in 0 components in a macro average. For multilabel targets, labels are column indices. By default, all labels in `predictions` and `references` are used in sorted order. Defaults to None.
pos_label (`int`): The class to be considered the positive class, in the case where `average` is set to `binary`. Defaults to 1.
average (`string`): This parameter is required for multiclass/multilabel targets. If set to `None`, the scores for each class are returned. Otherwise, this determines the type of averaging performed on the data. Defaults to `'binary'`.
- 'binary': Only report results for the class specified by `pos_label`. This is applicable only if the classes found in `predictions` and `references` are binary.
- 'micro': Calculate metrics globally by counting the total true positives, false negatives and false positives.
- 'macro': Calculate metrics for each label, and find their unweighted mean. This does not take label imbalance into account.
- 'weighted': Calculate metrics for each label, and find their average weighted by support (the number of true instances for each label). This alters `'macro'` to account for label imbalance. This option can result in an F-score that is not between precision and recall.
- 'samples': Calculate metrics for each instance, and find their average (only meaningful for multilabel classification).
sample_weight (`list` of `float`): Sample weights Defaults to None.
zero_division (`int` or `string`): Sets the value to return when there is a zero division. Defaults to 'warn'.
- 0: Returns 0 when there is a zero division.
- 1: Returns 1 when there is a zero division.
- 'warn': Raises warnings and then returns 0 when there is a zero division.
Returns:
precision (`float` or `array` of `float`): Precision score or list of precision scores, depending on the value passed to `average`. Minimum possible value is 0. Maximum possible value is 1. Higher values indicate that fewer negative examples were incorrectly labeled as positive, which means that, generally, higher scores are better.
Examples:
Example 1-A simple binary example
>>> precision_metric = datasets.load_metric("precision")
>>> results = precision_metric.compute(references=[0, 1, 0, 1, 0], predictions=[0, 0, 1, 1, 0])
>>> print(results)
{'precision': 0.5}
Example 2-The same simple binary example as in Example 1, but with `pos_label` set to `0`.
>>> precision_metric = datasets.load_metric("precision")
>>> results = precision_metric.compute(references=[0, 1, 0, 1, 0], predictions=[0, 0, 1, 1, 0], pos_label=0)
>>> print(round(results['precision'], 2))
0.67
Example 3-The same simple binary example as in Example 1, but with `sample_weight` included.
>>> precision_metric = datasets.load_metric("precision")
>>> results = precision_metric.compute(references=[0, 1, 0, 1, 0], predictions=[0, 0, 1, 1, 0], sample_weight=[0.9, 0.5, 3.9, 1.2, 0.3])
>>> print(results)
{'precision': 0.23529411764705882}
Example 4-A multiclass example, with different values for the `average` input.
>>> predictions = [0, 2, 1, 0, 0, 1]
>>> references = [0, 1, 2, 0, 1, 2]
>>> results = precision_metric.compute(predictions=predictions, references=references, average='macro')
>>> print(results)
{'precision': 0.2222222222222222}
>>> results = precision_metric.compute(predictions=predictions, references=references, average='micro')
>>> print(results)
{'precision': 0.3333333333333333}
>>> results = precision_metric.compute(predictions=predictions, references=references, average='weighted')
>>> print(results)
{'precision': 0.2222222222222222}
>>> results = precision_metric.compute(predictions=predictions, references=references, average=None)
>>> print([round(res, 2) for res in results['precision']])
[0.67, 0.0, 0.0]
"""
_CITATION = """
@article{scikit-learn,
title={Scikit-learn: Machine Learning in {P}ython},
author={Pedregosa, F. and Varoquaux, G. and Gramfort, A. and Michel, V.
and Thirion, B. and Grisel, O. and Blondel, M. and Prettenhofer, P.
and Weiss, R. and Dubourg, V. and Vanderplas, J. and Passos, A. and
Cournapeau, D. and Brucher, M. and Perrot, M. and Duchesnay, E.},
journal={Journal of Machine Learning Research},
volume={12},
pages={2825--2830},
year={2011}
}
"""
@datasets.utils.file_utils.add_start_docstrings(_DESCRIPTION, _KWARGS_DESCRIPTION)
class Precision(datasets.Metric):
def _info(self):
return datasets.MetricInfo(
description=_DESCRIPTION,
citation=_CITATION,
inputs_description=_KWARGS_DESCRIPTION,
features=datasets.Features(
{
"predictions": datasets.Sequence(datasets.Value("int32")),
"references": datasets.Sequence(datasets.Value("int32")),
}
if self.config_name == "multilabel"
else {
"predictions": datasets.Value("int32"),
"references": datasets.Value("int32"),
}
),
reference_urls=["https://scikit-learn.org/stable/modules/generated/sklearn.metrics.precision_score.html"],
)
def _compute(
self,
predictions,
references,
labels=None,
pos_label=1,
average="binary",
sample_weight=None,
zero_division="warn",
):
score = precision_score(
references,
predictions,
labels=labels,
pos_label=pos_label,
average=average,
sample_weight=sample_weight,
zero_division=zero_division,
)
return {"precision": float(score) if score.size == 1 else score}
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/cer/cer.py
|
# Copyright 2021 The HuggingFace Datasets Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
""" Character Error Ratio (CER) metric. """
from typing import List
import jiwer
import jiwer.transforms as tr
from packaging import version
import datasets
from datasets.config import PY_VERSION
if PY_VERSION < version.parse("3.8"):
import importlib_metadata
else:
import importlib.metadata as importlib_metadata
SENTENCE_DELIMITER = ""
if version.parse(importlib_metadata.version("jiwer")) < version.parse("2.3.0"):
class SentencesToListOfCharacters(tr.AbstractTransform):
def __init__(self, sentence_delimiter: str = " "):
self.sentence_delimiter = sentence_delimiter
def process_string(self, s: str):
return list(s)
def process_list(self, inp: List[str]):
chars = []
for sent_idx, sentence in enumerate(inp):
chars.extend(self.process_string(sentence))
if self.sentence_delimiter is not None and self.sentence_delimiter != "" and sent_idx < len(inp) - 1:
chars.append(self.sentence_delimiter)
return chars
cer_transform = tr.Compose(
[tr.RemoveMultipleSpaces(), tr.Strip(), SentencesToListOfCharacters(SENTENCE_DELIMITER)]
)
else:
cer_transform = tr.Compose(
[
tr.RemoveMultipleSpaces(),
tr.Strip(),
tr.ReduceToSingleSentence(SENTENCE_DELIMITER),
tr.ReduceToListOfListOfChars(),
]
)
_CITATION = """\
@inproceedings{inproceedings,
author = {Morris, Andrew and Maier, Viktoria and Green, Phil},
year = {2004},
month = {01},
pages = {},
title = {From WER and RIL to MER and WIL: improved evaluation measures for connected speech recognition.}
}
"""
_DESCRIPTION = """\
Character error rate (CER) is a common metric of the performance of an automatic speech recognition system.
CER is similar to Word Error Rate (WER), but operates on character instead of word. Please refer to docs of WER for further information.
Character error rate can be computed as:
CER = (S + D + I) / N = (S + D + I) / (S + D + C)
where
S is the number of substitutions,
D is the number of deletions,
I is the number of insertions,
C is the number of correct characters,
N is the number of characters in the reference (N=S+D+C).
CER's output is not always a number between 0 and 1, in particular when there is a high number of insertions. This value is often associated to the percentage of characters that were incorrectly predicted. The lower the value, the better the
performance of the ASR system with a CER of 0 being a perfect score.
"""
_KWARGS_DESCRIPTION = """
Computes CER score of transcribed segments against references.
Args:
references: list of references for each speech input.
predictions: list of transcribtions to score.
concatenate_texts: Whether or not to concatenate sentences before evaluation, set to True for more accurate result.
Returns:
(float): the character error rate
Examples:
>>> predictions = ["this is the prediction", "there is an other sample"]
>>> references = ["this is the reference", "there is another one"]
>>> cer = datasets.load_metric("cer")
>>> cer_score = cer.compute(predictions=predictions, references=references)
>>> print(cer_score)
0.34146341463414637
"""
@datasets.utils.file_utils.add_start_docstrings(_DESCRIPTION, _KWARGS_DESCRIPTION)
class CER(datasets.Metric):
def _info(self):
return datasets.MetricInfo(
description=_DESCRIPTION,
citation=_CITATION,
inputs_description=_KWARGS_DESCRIPTION,
features=datasets.Features(
{
"predictions": datasets.Value("string", id="sequence"),
"references": datasets.Value("string", id="sequence"),
}
),
codebase_urls=["https://github.com/jitsi/jiwer/"],
reference_urls=[
"https://en.wikipedia.org/wiki/Word_error_rate",
"https://sites.google.com/site/textdigitisation/qualitymeasures/computingerrorrates",
],
)
def _compute(self, predictions, references, concatenate_texts=False):
if concatenate_texts:
return jiwer.compute_measures(
references,
predictions,
truth_transform=cer_transform,
hypothesis_transform=cer_transform,
)["wer"]
incorrect = 0
total = 0
for prediction, reference in zip(predictions, references):
measures = jiwer.compute_measures(
reference,
prediction,
truth_transform=cer_transform,
hypothesis_transform=cer_transform,
)
incorrect += measures["substitutions"] + measures["deletions"] + measures["insertions"]
total += measures["substitutions"] + measures["deletions"] + measures["hits"]
return incorrect / total
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/cer/README.md
|
# Metric Card for CER
## Metric description
Character error rate (CER) is a common metric of the performance of an automatic speech recognition (ASR) system. CER is similar to Word Error Rate (WER), but operates on character instead of word.
Character error rate can be computed as:
`CER = (S + D + I) / N = (S + D + I) / (S + D + C)`
where
`S` is the number of substitutions,
`D` is the number of deletions,
`I` is the number of insertions,
`C` is the number of correct characters,
`N` is the number of characters in the reference (`N=S+D+C`).
## How to use
The metric takes two inputs: references (a list of references for each speech input) and predictions (a list of transcriptions to score).
```python
from datasets import load_metric
cer = load_metric("cer")
cer_score = cer.compute(predictions=predictions, references=references)
```
## Output values
This metric outputs a float representing the character error rate.
```
print(cer_score)
0.34146341463414637
```
The **lower** the CER value, the **better** the performance of the ASR system, with a CER of 0 being a perfect score.
However, CER's output is not always a number between 0 and 1, in particular when there is a high number of insertions (see [Examples](#Examples) below).
### Values from popular papers
This metric is highly dependent on the content and quality of the dataset, and therefore users can expect very different values for the same model but on different datasets.
Multilingual datasets such as [Common Voice](https://huggingface.co/datasets/common_voice) report different CERs depending on the language, ranging from 0.02-0.03 for languages such as French and Italian, to 0.05-0.07 for English (see [here](https://github.com/speechbrain/speechbrain/tree/develop/recipes/CommonVoice/ASR/CTC) for more values).
## Examples
Perfect match between prediction and reference:
```python
from datasets import load_metric
cer = load_metric("cer")
predictions = ["hello world", "good night moon"]
references = ["hello world", "good night moon"]
cer_score = cer.compute(predictions=predictions, references=references)
print(cer_score)
0.0
```
Partial match between prediction and reference:
```python
from datasets import load_metric
cer = load_metric("cer")
predictions = ["this is the prediction", "there is an other sample"]
references = ["this is the reference", "there is another one"]
cer_score = cer.compute(predictions=predictions, references=references)
print(cer_score)
0.34146341463414637
```
No match between prediction and reference:
```python
from datasets import load_metric
cer = load_metric("cer")
predictions = ["hello"]
references = ["gracias"]
cer_score = cer.compute(predictions=predictions, references=references)
print(cer_score)
1.0
```
CER above 1 due to insertion errors:
```python
from datasets import load_metric
cer = load_metric("cer")
predictions = ["hello world"]
references = ["hello"]
cer_score = cer.compute(predictions=predictions, references=references)
print(cer_score)
1.2
```
## Limitations and bias
CER is useful for comparing different models for tasks such as automatic speech recognition (ASR) and optic character recognition (OCR), especially for multilingual datasets where WER is not suitable given the diversity of languages. However, CER provides no details on the nature of translation errors and further work is therefore required to identify the main source(s) of error and to focus any research effort.
Also, in some cases, instead of reporting the raw CER, a normalized CER is reported where the number of mistakes is divided by the sum of the number of edit operations (`I` + `S` + `D`) and `C` (the number of correct characters), which results in CER values that fall within the range of 0–100%.
## Citation
```bibtex
@inproceedings{morris2004,
author = {Morris, Andrew and Maier, Viktoria and Green, Phil},
year = {2004},
month = {01},
pages = {},
title = {From WER and RIL to MER and WIL: improved evaluation measures for connected speech recognition.}
}
```
## Further References
- [Hugging Face Tasks -- Automatic Speech Recognition](https://huggingface.co/tasks/automatic-speech-recognition)
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/cer/test_cer.py
|
# Copyright 2021 The HuggingFace Datasets Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
import unittest
from cer import CER
cer = CER()
class TestCER(unittest.TestCase):
def test_cer_case_senstive(self):
refs = ["White House"]
preds = ["white house"]
# S = 2, D = 0, I = 0, N = 11, CER = 2 / 11
char_error_rate = cer.compute(predictions=preds, references=refs)
self.assertTrue(abs(char_error_rate - 0.1818181818) < 1e-6)
def test_cer_whitespace(self):
refs = ["were wolf"]
preds = ["werewolf"]
# S = 0, D = 0, I = 1, N = 9, CER = 1 / 9
char_error_rate = cer.compute(predictions=preds, references=refs)
self.assertTrue(abs(char_error_rate - 0.1111111) < 1e-6)
refs = ["werewolf"]
preds = ["weae wolf"]
# S = 1, D = 1, I = 0, N = 8, CER = 0.25
char_error_rate = cer.compute(predictions=preds, references=refs)
self.assertTrue(abs(char_error_rate - 0.25) < 1e-6)
# consecutive whitespaces case 1
refs = ["were wolf"]
preds = ["were wolf"]
# S = 0, D = 0, I = 0, N = 9, CER = 0
char_error_rate = cer.compute(predictions=preds, references=refs)
self.assertTrue(abs(char_error_rate - 0.0) < 1e-6)
# consecutive whitespaces case 2
refs = ["were wolf"]
preds = ["were wolf"]
# S = 0, D = 0, I = 0, N = 9, CER = 0
char_error_rate = cer.compute(predictions=preds, references=refs)
self.assertTrue(abs(char_error_rate - 0.0) < 1e-6)
def test_cer_sub(self):
refs = ["werewolf"]
preds = ["weaewolf"]
# S = 1, D = 0, I = 0, N = 8, CER = 0.125
char_error_rate = cer.compute(predictions=preds, references=refs)
self.assertTrue(abs(char_error_rate - 0.125) < 1e-6)
def test_cer_del(self):
refs = ["werewolf"]
preds = ["wereawolf"]
# S = 0, D = 1, I = 0, N = 8, CER = 0.125
char_error_rate = cer.compute(predictions=preds, references=refs)
self.assertTrue(abs(char_error_rate - 0.125) < 1e-6)
def test_cer_insert(self):
refs = ["werewolf"]
preds = ["wereolf"]
# S = 0, D = 0, I = 1, N = 8, CER = 0.125
char_error_rate = cer.compute(predictions=preds, references=refs)
self.assertTrue(abs(char_error_rate - 0.125) < 1e-6)
def test_cer_equal(self):
refs = ["werewolf"]
char_error_rate = cer.compute(predictions=refs, references=refs)
self.assertEqual(char_error_rate, 0.0)
def test_cer_list_of_seqs(self):
refs = ["werewolf", "I am your father"]
char_error_rate = cer.compute(predictions=refs, references=refs)
self.assertEqual(char_error_rate, 0.0)
refs = ["werewolf", "I am your father", "doge"]
preds = ["werxwolf", "I am your father", "doge"]
# S = 1, D = 0, I = 0, N = 28, CER = 1 / 28
char_error_rate = cer.compute(predictions=preds, references=refs)
self.assertTrue(abs(char_error_rate - 0.03571428) < 1e-6)
def test_correlated_sentences(self):
refs = ["My hovercraft", "is full of eels"]
preds = ["My hovercraft is full", " of eels"]
# S = 0, D = 0, I = 2, N = 28, CER = 2 / 28
# whitespace at the front of " of eels" will be strip during preporcessing
# so need to insert 2 whitespaces
char_error_rate = cer.compute(predictions=preds, references=refs, concatenate_texts=True)
self.assertTrue(abs(char_error_rate - 0.071428) < 1e-6)
def test_cer_unicode(self):
refs = ["我能吞下玻璃而不伤身体"]
preds = [" 能吞虾玻璃而 不霜身体啦"]
# S = 3, D = 2, I = 0, N = 11, CER = 5 / 11
char_error_rate = cer.compute(predictions=preds, references=refs)
self.assertTrue(abs(char_error_rate - 0.4545454545) < 1e-6)
refs = ["我能吞下玻璃", "而不伤身体"]
preds = ["我 能 吞 下 玻 璃", "而不伤身体"]
# S = 0, D = 5, I = 0, N = 11, CER = 5 / 11
char_error_rate = cer.compute(predictions=preds, references=refs)
self.assertTrue(abs(char_error_rate - 0.454545454545) < 1e-6)
refs = ["我能吞下玻璃而不伤身体"]
char_error_rate = cer.compute(predictions=refs, references=refs)
self.assertFalse(char_error_rate, 0.0)
def test_cer_empty(self):
refs = [""]
preds = ["Hypothesis"]
with self.assertRaises(ValueError):
cer.compute(predictions=preds, references=refs)
if __name__ == "__main__":
unittest.main()
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/glue/README.md
|
# Metric Card for GLUE
## Metric description
This metric is used to compute the GLUE evaluation metric associated to each [GLUE dataset](https://huggingface.co/datasets/glue).
GLUE, the General Language Understanding Evaluation benchmark is a collection of resources for training, evaluating, and analyzing natural language understanding systems.
## How to use
There are two steps: (1) loading the GLUE metric relevant to the subset of the GLUE dataset being used for evaluation; and (2) calculating the metric.
1. **Loading the relevant GLUE metric** : the subsets of GLUE are the following: `sst2`, `mnli`, `mnli_mismatched`, `mnli_matched`, `qnli`, `rte`, `wnli`, `cola`,`stsb`, `mrpc`, `qqp`, and `hans`.
More information about the different subsets of the GLUE dataset can be found on the [GLUE dataset page](https://huggingface.co/datasets/glue).
2. **Calculating the metric**: the metric takes two inputs : one list with the predictions of the model to score and one lists of references for each translation.
```python
from datasets import load_metric
glue_metric = load_metric('glue', 'sst2')
references = [0, 1]
predictions = [0, 1]
results = glue_metric.compute(predictions=predictions, references=references)
```
## Output values
The output of the metric depends on the GLUE subset chosen, consisting of a dictionary that contains one or several of the following metrics:
`accuracy`: the proportion of correct predictions among the total number of cases processed, with a range between 0 and 1 (see [accuracy](https://huggingface.co/metrics/accuracy) for more information).
`f1`: the harmonic mean of the precision and recall (see [F1 score](https://huggingface.co/metrics/f1) for more information). Its range is 0-1 -- its lowest possible value is 0, if either the precision or the recall is 0, and its highest possible value is 1.0, which means perfect precision and recall.
`pearson`: a measure of the linear relationship between two datasets (see [Pearson correlation](https://huggingface.co/metrics/pearsonr) for more information). Its range is between -1 and +1, with 0 implying no correlation, and -1/+1 implying an exact linear relationship. Positive correlations imply that as x increases, so does y, whereas negative correlations imply that as x increases, y decreases.
`spearmanr`: a nonparametric measure of the monotonicity of the relationship between two datasets(see [Spearman Correlation](https://huggingface.co/metrics/spearmanr) for more information). `spearmanr` has the same range as `pearson`.
`matthews_correlation`: a measure of the quality of binary and multiclass classifications (see [Matthews Correlation](https://huggingface.co/metrics/matthews_correlation) for more information). Its range of values is between -1 and +1, where a coefficient of +1 represents a perfect prediction, 0 an average random prediction and -1 an inverse prediction.
The `cola` subset returns `matthews_correlation`, the `stsb` subset returns `pearson` and `spearmanr`, the `mrpc` and `qqp` subsets return both `accuracy` and `f1`, and all other subsets of GLUE return only accuracy.
### Values from popular papers
The [original GLUE paper](https://huggingface.co/datasets/glue) reported average scores ranging from 58 to 64%, depending on the model used (with all evaluation values scaled by 100 to make computing the average possible).
For more recent model performance, see the [dataset leaderboard](https://paperswithcode.com/dataset/glue).
## Examples
Maximal values for the MRPC subset (which outputs `accuracy` and `f1`):
```python
from datasets import load_metric
glue_metric = load_metric('glue', 'mrpc') # 'mrpc' or 'qqp'
references = [0, 1]
predictions = [0, 1]
results = glue_metric.compute(predictions=predictions, references=references)
print(results)
{'accuracy': 1.0, 'f1': 1.0}
```
Minimal values for the STSB subset (which outputs `pearson` and `spearmanr`):
```python
from datasets import load_metric
glue_metric = load_metric('glue', 'stsb')
references = [0., 1., 2., 3., 4., 5.]
predictions = [-10., -11., -12., -13., -14., -15.]
results = glue_metric.compute(predictions=predictions, references=references)
print(results)
{'pearson': -1.0, 'spearmanr': -1.0}
```
Partial match for the COLA subset (which outputs `matthews_correlation`)
```python
from datasets import load_metric
glue_metric = load_metric('glue', 'cola')
references = [0, 1]
predictions = [1, 1]
results = glue_metric.compute(predictions=predictions, references=references)
results
{'matthews_correlation': 0.0}
```
## Limitations and bias
This metric works only with datasets that have the same format as the [GLUE dataset](https://huggingface.co/datasets/glue).
While the GLUE dataset is meant to represent "General Language Understanding", the tasks represented in it are not necessarily representative of language understanding, and should not be interpreted as such.
Also, while the GLUE subtasks were considered challenging during its creation in 2019, they are no longer considered as such given the impressive progress made since then. A more complex (or "stickier") version of it, called [SuperGLUE](https://huggingface.co/datasets/super_glue), was subsequently created.
## Citation
```bibtex
@inproceedings{wang2019glue,
title={{GLUE}: A Multi-Task Benchmark and Analysis Platform for Natural Language Understanding},
author={Wang, Alex and Singh, Amanpreet and Michael, Julian and Hill, Felix and Levy, Omer and Bowman, Samuel R.},
note={In the Proceedings of ICLR.},
year={2019}
}
```
## Further References
- [GLUE benchmark homepage](https://gluebenchmark.com/)
- [Fine-tuning a model with the Trainer API](https://huggingface.co/course/chapter3/3?)
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/glue/glue.py
|
# Copyright 2020 The HuggingFace Datasets Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
""" GLUE benchmark metric. """
from scipy.stats import pearsonr, spearmanr
from sklearn.metrics import f1_score, matthews_corrcoef
import datasets
_CITATION = """\
@inproceedings{wang2019glue,
title={{GLUE}: A Multi-Task Benchmark and Analysis Platform for Natural Language Understanding},
author={Wang, Alex and Singh, Amanpreet and Michael, Julian and Hill, Felix and Levy, Omer and Bowman, Samuel R.},
note={In the Proceedings of ICLR.},
year={2019}
}
"""
_DESCRIPTION = """\
GLUE, the General Language Understanding Evaluation benchmark
(https://gluebenchmark.com/) is a collection of resources for training,
evaluating, and analyzing natural language understanding systems.
"""
_KWARGS_DESCRIPTION = """
Compute GLUE evaluation metric associated to each GLUE dataset.
Args:
predictions: list of predictions to score.
Each translation should be tokenized into a list of tokens.
references: list of lists of references for each translation.
Each reference should be tokenized into a list of tokens.
Returns: depending on the GLUE subset, one or several of:
"accuracy": Accuracy
"f1": F1 score
"pearson": Pearson Correlation
"spearmanr": Spearman Correlation
"matthews_correlation": Matthew Correlation
Examples:
>>> glue_metric = datasets.load_metric('glue', 'sst2') # 'sst2' or any of ["mnli", "mnli_mismatched", "mnli_matched", "qnli", "rte", "wnli", "hans"]
>>> references = [0, 1]
>>> predictions = [0, 1]
>>> results = glue_metric.compute(predictions=predictions, references=references)
>>> print(results)
{'accuracy': 1.0}
>>> glue_metric = datasets.load_metric('glue', 'mrpc') # 'mrpc' or 'qqp'
>>> references = [0, 1]
>>> predictions = [0, 1]
>>> results = glue_metric.compute(predictions=predictions, references=references)
>>> print(results)
{'accuracy': 1.0, 'f1': 1.0}
>>> glue_metric = datasets.load_metric('glue', 'stsb')
>>> references = [0., 1., 2., 3., 4., 5.]
>>> predictions = [0., 1., 2., 3., 4., 5.]
>>> results = glue_metric.compute(predictions=predictions, references=references)
>>> print({"pearson": round(results["pearson"], 2), "spearmanr": round(results["spearmanr"], 2)})
{'pearson': 1.0, 'spearmanr': 1.0}
>>> glue_metric = datasets.load_metric('glue', 'cola')
>>> references = [0, 1]
>>> predictions = [0, 1]
>>> results = glue_metric.compute(predictions=predictions, references=references)
>>> print(results)
{'matthews_correlation': 1.0}
"""
def simple_accuracy(preds, labels):
return float((preds == labels).mean())
def acc_and_f1(preds, labels):
acc = simple_accuracy(preds, labels)
f1 = float(f1_score(y_true=labels, y_pred=preds))
return {
"accuracy": acc,
"f1": f1,
}
def pearson_and_spearman(preds, labels):
pearson_corr = float(pearsonr(preds, labels)[0])
spearman_corr = float(spearmanr(preds, labels)[0])
return {
"pearson": pearson_corr,
"spearmanr": spearman_corr,
}
@datasets.utils.file_utils.add_start_docstrings(_DESCRIPTION, _KWARGS_DESCRIPTION)
class Glue(datasets.Metric):
def _info(self):
if self.config_name not in [
"sst2",
"mnli",
"mnli_mismatched",
"mnli_matched",
"cola",
"stsb",
"mrpc",
"qqp",
"qnli",
"rte",
"wnli",
"hans",
]:
raise KeyError(
"You should supply a configuration name selected in "
'["sst2", "mnli", "mnli_mismatched", "mnli_matched", '
'"cola", "stsb", "mrpc", "qqp", "qnli", "rte", "wnli", "hans"]'
)
return datasets.MetricInfo(
description=_DESCRIPTION,
citation=_CITATION,
inputs_description=_KWARGS_DESCRIPTION,
features=datasets.Features(
{
"predictions": datasets.Value("int64" if self.config_name != "stsb" else "float32"),
"references": datasets.Value("int64" if self.config_name != "stsb" else "float32"),
}
),
codebase_urls=[],
reference_urls=[],
format="numpy",
)
def _compute(self, predictions, references):
if self.config_name == "cola":
return {"matthews_correlation": matthews_corrcoef(references, predictions)}
elif self.config_name == "stsb":
return pearson_and_spearman(predictions, references)
elif self.config_name in ["mrpc", "qqp"]:
return acc_and_f1(predictions, references)
elif self.config_name in ["sst2", "mnli", "mnli_mismatched", "mnli_matched", "qnli", "rte", "wnli", "hans"]:
return {"accuracy": simple_accuracy(predictions, references)}
else:
raise KeyError(
"You should supply a configuration name selected in "
'["sst2", "mnli", "mnli_mismatched", "mnli_matched", '
'"cola", "stsb", "mrpc", "qqp", "qnli", "rte", "wnli", "hans"]'
)
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/accuracy/README.md
|
# Metric Card for Accuracy
## Metric Description
Accuracy is the proportion of correct predictions among the total number of cases processed. It can be computed with:
Accuracy = (TP + TN) / (TP + TN + FP + FN)
Where:
TP: True positive
TN: True negative
FP: False positive
FN: False negative
## How to Use
At minimum, this metric requires predictions and references as inputs.
```python
>>> accuracy_metric = datasets.load_metric("accuracy")
>>> results = accuracy_metric.compute(references=[0, 1], predictions=[0, 1])
>>> print(results)
{'accuracy': 1.0}
```
### Inputs
- **predictions** (`list` of `int`): Predicted labels.
- **references** (`list` of `int`): Ground truth labels.
- **normalize** (`boolean`): If set to False, returns the number of correctly classified samples. Otherwise, returns the fraction of correctly classified samples. Defaults to True.
- **sample_weight** (`list` of `float`): Sample weights Defaults to None.
### Output Values
- **accuracy**(`float` or `int`): Accuracy score. Minimum possible value is 0. Maximum possible value is 1.0, or the number of examples input, if `normalize` is set to `True`.. A higher score means higher accuracy.
Output Example(s):
```python
{'accuracy': 1.0}
```
This metric outputs a dictionary, containing the accuracy score.
#### Values from Popular Papers
Top-1 or top-5 accuracy is often used to report performance on supervised classification tasks such as image classification (e.g. on [ImageNet](https://paperswithcode.com/sota/image-classification-on-imagenet)) or sentiment analysis (e.g. on [IMDB](https://paperswithcode.com/sota/text-classification-on-imdb)).
### Examples
Example 1-A simple example
```python
>>> accuracy_metric = datasets.load_metric("accuracy")
>>> results = accuracy_metric.compute(references=[0, 1, 2, 0, 1, 2], predictions=[0, 1, 1, 2, 1, 0])
>>> print(results)
{'accuracy': 0.5}
```
Example 2-The same as Example 1, except with `normalize` set to `False`.
```python
>>> accuracy_metric = datasets.load_metric("accuracy")
>>> results = accuracy_metric.compute(references=[0, 1, 2, 0, 1, 2], predictions=[0, 1, 1, 2, 1, 0], normalize=False)
>>> print(results)
{'accuracy': 3.0}
```
Example 3-The same as Example 1, except with `sample_weight` set.
```python
>>> accuracy_metric = datasets.load_metric("accuracy")
>>> results = accuracy_metric.compute(references=[0, 1, 2, 0, 1, 2], predictions=[0, 1, 1, 2, 1, 0], sample_weight=[0.5, 2, 0.7, 0.5, 9, 0.4])
>>> print(results)
{'accuracy': 0.8778625954198473}
```
## Limitations and Bias
This metric can be easily misleading, especially in the case of unbalanced classes. For example, a high accuracy might be because a model is doing well, but if the data is unbalanced, it might also be because the model is only accurately labeling the high-frequency class. In such cases, a more detailed analysis of the model's behavior, or the use of a different metric entirely, is necessary to determine how well the model is actually performing.
## Citation(s)
```bibtex
@article{scikit-learn,
title={Scikit-learn: Machine Learning in {P}ython},
author={Pedregosa, F. and Varoquaux, G. and Gramfort, A. and Michel, V.
and Thirion, B. and Grisel, O. and Blondel, M. and Prettenhofer, P.
and Weiss, R. and Dubourg, V. and Vanderplas, J. and Passos, A. and
Cournapeau, D. and Brucher, M. and Perrot, M. and Duchesnay, E.},
journal={Journal of Machine Learning Research},
volume={12},
pages={2825--2830},
year={2011}
}
```
## Further References
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/accuracy/accuracy.py
|
# Copyright 2020 The HuggingFace Datasets Authors and the current dataset script contributor.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""Accuracy metric."""
from sklearn.metrics import accuracy_score
import datasets
_DESCRIPTION = """
Accuracy is the proportion of correct predictions among the total number of cases processed. It can be computed with:
Accuracy = (TP + TN) / (TP + TN + FP + FN)
Where:
TP: True positive
TN: True negative
FP: False positive
FN: False negative
"""
_KWARGS_DESCRIPTION = """
Args:
predictions (`list` of `int`): Predicted labels.
references (`list` of `int`): Ground truth labels.
normalize (`boolean`): If set to False, returns the number of correctly classified samples. Otherwise, returns the fraction of correctly classified samples. Defaults to True.
sample_weight (`list` of `float`): Sample weights Defaults to None.
Returns:
accuracy (`float` or `int`): Accuracy score. Minimum possible value is 0. Maximum possible value is 1.0, or the number of examples input, if `normalize` is set to `True`.. A higher score means higher accuracy.
Examples:
Example 1-A simple example
>>> accuracy_metric = datasets.load_metric("accuracy")
>>> results = accuracy_metric.compute(references=[0, 1, 2, 0, 1, 2], predictions=[0, 1, 1, 2, 1, 0])
>>> print(results)
{'accuracy': 0.5}
Example 2-The same as Example 1, except with `normalize` set to `False`.
>>> accuracy_metric = datasets.load_metric("accuracy")
>>> results = accuracy_metric.compute(references=[0, 1, 2, 0, 1, 2], predictions=[0, 1, 1, 2, 1, 0], normalize=False)
>>> print(results)
{'accuracy': 3.0}
Example 3-The same as Example 1, except with `sample_weight` set.
>>> accuracy_metric = datasets.load_metric("accuracy")
>>> results = accuracy_metric.compute(references=[0, 1, 2, 0, 1, 2], predictions=[0, 1, 1, 2, 1, 0], sample_weight=[0.5, 2, 0.7, 0.5, 9, 0.4])
>>> print(results)
{'accuracy': 0.8778625954198473}
"""
_CITATION = """
@article{scikit-learn,
title={Scikit-learn: Machine Learning in {P}ython},
author={Pedregosa, F. and Varoquaux, G. and Gramfort, A. and Michel, V.
and Thirion, B. and Grisel, O. and Blondel, M. and Prettenhofer, P.
and Weiss, R. and Dubourg, V. and Vanderplas, J. and Passos, A. and
Cournapeau, D. and Brucher, M. and Perrot, M. and Duchesnay, E.},
journal={Journal of Machine Learning Research},
volume={12},
pages={2825--2830},
year={2011}
}
"""
@datasets.utils.file_utils.add_start_docstrings(_DESCRIPTION, _KWARGS_DESCRIPTION)
class Accuracy(datasets.Metric):
def _info(self):
return datasets.MetricInfo(
description=_DESCRIPTION,
citation=_CITATION,
inputs_description=_KWARGS_DESCRIPTION,
features=datasets.Features(
{
"predictions": datasets.Sequence(datasets.Value("int32")),
"references": datasets.Sequence(datasets.Value("int32")),
}
if self.config_name == "multilabel"
else {
"predictions": datasets.Value("int32"),
"references": datasets.Value("int32"),
}
),
reference_urls=["https://scikit-learn.org/stable/modules/generated/sklearn.metrics.accuracy_score.html"],
)
def _compute(self, predictions, references, normalize=True, sample_weight=None):
return {
"accuracy": float(
accuracy_score(references, predictions, normalize=normalize, sample_weight=sample_weight)
)
}
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/ter/README.md
|
# Metric Card for TER
## Metric Description
TER (Translation Edit Rate, also called Translation Error Rate) is a metric to quantify the edit operations that a hypothesis requires to match a reference translation. We use the implementation that is already present in [sacrebleu](https://github.com/mjpost/sacreBLEU#ter), which in turn is inspired by the [TERCOM implementation](https://github.com/jhclark/tercom).
The implementation here is slightly different from sacrebleu in terms of the required input format. The length of the references and hypotheses lists need to be the same, so you may need to transpose your references compared to sacrebleu's required input format. See [this github issue](https://github.com/huggingface/datasets/issues/3154#issuecomment-950746534).
See the README.md file at https://github.com/mjpost/sacreBLEU#ter for more information.
## How to Use
This metric takes, at minimum, predicted sentences and reference sentences:
```python
>>> predictions = ["does this sentence match??",
... "what about this sentence?",
... "What did the TER metric user say to the developer?"]
>>> references = [["does this sentence match", "does this sentence match!?!"],
... ["wHaT aBoUt ThIs SeNtEnCe?", "wHaT aBoUt ThIs SeNtEnCe?"],
... ["Your jokes are...", "...TERrible"]]
>>> ter = datasets.load_metric("ter")
>>> results = ter.compute(predictions=predictions,
... references=references,
... case_sensitive=True)
>>> print(results)
{'score': 150.0, 'num_edits': 15, 'ref_length': 10.0}
```
### Inputs
This metric takes the following as input:
- **`predictions`** (`list` of `str`): The system stream (a sequence of segments).
- **`references`** (`list` of `list` of `str`): A list of one or more reference streams (each a sequence of segments).
- **`normalized`** (`boolean`): If `True`, applies basic tokenization and normalization to sentences. Defaults to `False`.
- **`ignore_punct`** (`boolean`): If `True`, applies basic tokenization and normalization to sentences. Defaults to `False`.
- **`support_zh_ja_chars`** (`boolean`): If `True`, tokenization/normalization supports processing of Chinese characters, as well as Japanese Kanji, Hiragana, Katakana, and Phonetic Extensions of Katakana. Only applies if `normalized = True`. Defaults to `False`.
- **`case_sensitive`** (`boolean`): If `False`, makes all predictions and references lowercase to ignore differences in case. Defaults to `False`.
### Output Values
This metric returns the following:
- **`score`** (`float`): TER score (num_edits / sum_ref_lengths * 100)
- **`num_edits`** (`int`): The cumulative number of edits
- **`ref_length`** (`float`): The cumulative average reference length
The output takes the following form:
```python
{'score': ter_score, 'num_edits': num_edits, 'ref_length': ref_length}
```
The metric can take on any value `0` and above. `0` is a perfect score, meaning the predictions exactly match the references and no edits were necessary. Higher scores are worse. Scores above 100 mean that the cumulative number of edits, `num_edits`, is higher than the cumulative length of the references, `ref_length`.
#### Values from Popular Papers
### Examples
Basic example with only predictions and references as inputs:
```python
>>> predictions = ["does this sentence match??",
... "what about this sentence?"]
>>> references = [["does this sentence match", "does this sentence match!?!"],
... ["wHaT aBoUt ThIs SeNtEnCe?", "wHaT aBoUt ThIs SeNtEnCe?"]]
>>> ter = datasets.load_metric("ter")
>>> results = ter.compute(predictions=predictions,
... references=references,
... case_sensitive=True)
>>> print(results)
{'score': 62.5, 'num_edits': 5, 'ref_length': 8.0}
```
Example with `normalization = True`:
```python
>>> predictions = ["does this sentence match??",
... "what about this sentence?"]
>>> references = [["does this sentence match", "does this sentence match!?!"],
... ["wHaT aBoUt ThIs SeNtEnCe?", "wHaT aBoUt ThIs SeNtEnCe?"]]
>>> ter = datasets.load_metric("ter")
>>> results = ter.compute(predictions=predictions,
... references=references,
... normalized=True,
... case_sensitive=True)
>>> print(results)
{'score': 57.14285714285714, 'num_edits': 6, 'ref_length': 10.5}
```
Example ignoring punctuation and capitalization, and everything matches:
```python
>>> predictions = ["does this sentence match??",
... "what about this sentence?"]
>>> references = [["does this sentence match", "does this sentence match!?!"],
... ["wHaT aBoUt ThIs SeNtEnCe?", "wHaT aBoUt ThIs SeNtEnCe?"]]
>>> ter = datasets.load_metric("ter")
>>> results = ter.compute(predictions=predictions,
... references=references,
... ignore_punct=True,
... case_sensitive=False)
>>> print(results)
{'score': 0.0, 'num_edits': 0, 'ref_length': 8.0}
```
Example ignoring punctuation and capitalization, but with an extra (incorrect) sample:
```python
>>> predictions = ["does this sentence match??",
... "what about this sentence?",
... "What did the TER metric user say to the developer?"]
>>> references = [["does this sentence match", "does this sentence match!?!"],
... ["wHaT aBoUt ThIs SeNtEnCe?", "wHaT aBoUt ThIs SeNtEnCe?"],
... ["Your jokes are...", "...TERrible"]]
>>> ter = datasets.load_metric("ter")
>>> results = ter.compute(predictions=predictions,
... references=references,
... ignore_punct=True,
... case_sensitive=False)
>>> print(results)
{'score': 100.0, 'num_edits': 10, 'ref_length': 10.0}
```
## Limitations and Bias
## Citation
```bibtex
@inproceedings{snover-etal-2006-study,
title = "A Study of Translation Edit Rate with Targeted Human Annotation",
author = "Snover, Matthew and
Dorr, Bonnie and
Schwartz, Rich and
Micciulla, Linnea and
Makhoul, John",
booktitle = "Proceedings of the 7th Conference of the Association for Machine Translation in the Americas: Technical Papers",
month = aug # " 8-12",
year = "2006",
address = "Cambridge, Massachusetts, USA",
publisher = "Association for Machine Translation in the Americas",
url = "https://aclanthology.org/2006.amta-papers.25",
pages = "223--231",
}
@inproceedings{post-2018-call,
title = "A Call for Clarity in Reporting {BLEU} Scores",
author = "Post, Matt",
booktitle = "Proceedings of the Third Conference on Machine Translation: Research Papers",
month = oct,
year = "2018",
address = "Belgium, Brussels",
publisher = "Association for Computational Linguistics",
url = "https://www.aclweb.org/anthology/W18-6319",
pages = "186--191",
}
```
## Further References
- See [the sacreBLEU github repo](https://github.com/mjpost/sacreBLEU#ter) for more information.
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/ter/ter.py
|
# Copyright 2021 The HuggingFace Datasets Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
""" TER metric as available in sacrebleu. """
import sacrebleu as scb
from packaging import version
from sacrebleu import TER
import datasets
_CITATION = """\
@inproceedings{snover-etal-2006-study,
title = "A Study of Translation Edit Rate with Targeted Human Annotation",
author = "Snover, Matthew and
Dorr, Bonnie and
Schwartz, Rich and
Micciulla, Linnea and
Makhoul, John",
booktitle = "Proceedings of the 7th Conference of the Association for Machine Translation in the Americas: Technical Papers",
month = aug # " 8-12",
year = "2006",
address = "Cambridge, Massachusetts, USA",
publisher = "Association for Machine Translation in the Americas",
url = "https://aclanthology.org/2006.amta-papers.25",
pages = "223--231",
}
@inproceedings{post-2018-call,
title = "A Call for Clarity in Reporting {BLEU} Scores",
author = "Post, Matt",
booktitle = "Proceedings of the Third Conference on Machine Translation: Research Papers",
month = oct,
year = "2018",
address = "Belgium, Brussels",
publisher = "Association for Computational Linguistics",
url = "https://www.aclweb.org/anthology/W18-6319",
pages = "186--191",
}
"""
_DESCRIPTION = """\
TER (Translation Edit Rate, also called Translation Error Rate) is a metric to quantify the edit operations that a
hypothesis requires to match a reference translation. We use the implementation that is already present in sacrebleu
(https://github.com/mjpost/sacreBLEU#ter), which in turn is inspired by the TERCOM implementation, which can be found
here: https://github.com/jhclark/tercom.
The implementation here is slightly different from sacrebleu in terms of the required input format. The length of
the references and hypotheses lists need to be the same, so you may need to transpose your references compared to
sacrebleu's required input format. See https://github.com/huggingface/datasets/issues/3154#issuecomment-950746534
See the README.md file at https://github.com/mjpost/sacreBLEU#ter for more information.
"""
_KWARGS_DESCRIPTION = """
Produces TER scores alongside the number of edits and reference length.
Args:
predictions (list of str): The system stream (a sequence of segments).
references (list of list of str): A list of one or more reference streams (each a sequence of segments).
normalized (boolean): If `True`, applies basic tokenization and normalization to sentences. Defaults to `False`.
ignore_punct (boolean): If `True`, applies basic tokenization and normalization to sentences. Defaults to `False`.
support_zh_ja_chars (boolean): If `True`, tokenization/normalization supports processing of Chinese characters,
as well as Japanese Kanji, Hiragana, Katakana, and Phonetic Extensions of Katakana.
Only applies if `normalized = True`. Defaults to `False`.
case_sensitive (boolean): If `False`, makes all predictions and references lowercase to ignore differences in case. Defaults to `False`.
Returns:
'score' (float): TER score (num_edits / sum_ref_lengths * 100)
'num_edits' (int): The cumulative number of edits
'ref_length' (float): The cumulative average reference length
Examples:
Example 1:
>>> predictions = ["does this sentence match??",
... "what about this sentence?",
... "What did the TER metric user say to the developer?"]
>>> references = [["does this sentence match", "does this sentence match!?!"],
... ["wHaT aBoUt ThIs SeNtEnCe?", "wHaT aBoUt ThIs SeNtEnCe?"],
... ["Your jokes are...", "...TERrible"]]
>>> ter = datasets.load_metric("ter")
>>> results = ter.compute(predictions=predictions,
... references=references,
... case_sensitive=True)
>>> print(results)
{'score': 150.0, 'num_edits': 15, 'ref_length': 10.0}
Example 2:
>>> predictions = ["does this sentence match??",
... "what about this sentence?"]
>>> references = [["does this sentence match", "does this sentence match!?!"],
... ["wHaT aBoUt ThIs SeNtEnCe?", "wHaT aBoUt ThIs SeNtEnCe?"]]
>>> ter = datasets.load_metric("ter")
>>> results = ter.compute(predictions=predictions,
... references=references,
... case_sensitive=True)
>>> print(results)
{'score': 62.5, 'num_edits': 5, 'ref_length': 8.0}
Example 3:
>>> predictions = ["does this sentence match??",
... "what about this sentence?"]
>>> references = [["does this sentence match", "does this sentence match!?!"],
... ["wHaT aBoUt ThIs SeNtEnCe?", "wHaT aBoUt ThIs SeNtEnCe?"]]
>>> ter = datasets.load_metric("ter")
>>> results = ter.compute(predictions=predictions,
... references=references,
... normalized=True,
... case_sensitive=True)
>>> print(results)
{'score': 57.14285714285714, 'num_edits': 6, 'ref_length': 10.5}
Example 4:
>>> predictions = ["does this sentence match??",
... "what about this sentence?"]
>>> references = [["does this sentence match", "does this sentence match!?!"],
... ["wHaT aBoUt ThIs SeNtEnCe?", "wHaT aBoUt ThIs SeNtEnCe?"]]
>>> ter = datasets.load_metric("ter")
>>> results = ter.compute(predictions=predictions,
... references=references,
... ignore_punct=True,
... case_sensitive=False)
>>> print(results)
{'score': 0.0, 'num_edits': 0, 'ref_length': 8.0}
Example 5:
>>> predictions = ["does this sentence match??",
... "what about this sentence?",
... "What did the TER metric user say to the developer?"]
>>> references = [["does this sentence match", "does this sentence match!?!"],
... ["wHaT aBoUt ThIs SeNtEnCe?", "wHaT aBoUt ThIs SeNtEnCe?"],
... ["Your jokes are...", "...TERrible"]]
>>> ter = datasets.load_metric("ter")
>>> results = ter.compute(predictions=predictions,
... references=references,
... ignore_punct=True,
... case_sensitive=False)
>>> print(results)
{'score': 100.0, 'num_edits': 10, 'ref_length': 10.0}
"""
@datasets.utils.file_utils.add_start_docstrings(_DESCRIPTION, _KWARGS_DESCRIPTION)
class Ter(datasets.Metric):
def _info(self):
if version.parse(scb.__version__) < version.parse("1.4.12"):
raise ImportWarning(
"To use `sacrebleu`, the module `sacrebleu>=1.4.12` is required, and the current version of `sacrebleu` doesn't match this condition.\n"
'You can install it with `pip install "sacrebleu>=1.4.12"`.'
)
return datasets.MetricInfo(
description=_DESCRIPTION,
citation=_CITATION,
homepage="http://www.cs.umd.edu/~snover/tercom/",
inputs_description=_KWARGS_DESCRIPTION,
features=datasets.Features(
{
"predictions": datasets.Value("string", id="sequence"),
"references": datasets.Sequence(datasets.Value("string", id="sequence"), id="references"),
}
),
codebase_urls=["https://github.com/mjpost/sacreBLEU#ter"],
reference_urls=[
"https://github.com/jhclark/tercom",
],
)
def _compute(
self,
predictions,
references,
normalized: bool = False,
ignore_punct: bool = False,
support_zh_ja_chars: bool = False,
case_sensitive: bool = False,
):
references_per_prediction = len(references[0])
if any(len(refs) != references_per_prediction for refs in references):
raise ValueError("Sacrebleu requires the same number of references for each prediction")
transformed_references = [[refs[i] for refs in references] for i in range(references_per_prediction)]
sb_ter = TER(
normalized=normalized,
no_punct=ignore_punct,
asian_support=support_zh_ja_chars,
case_sensitive=case_sensitive,
)
output = sb_ter.corpus_score(predictions, transformed_references)
return {"score": output.score, "num_edits": output.num_edits, "ref_length": output.ref_length}
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/indic_glue/README.md
|
# Metric Card for IndicGLUE
## Metric description
This metric is used to compute the evaluation metric for the [IndicGLUE dataset](https://huggingface.co/datasets/indic_glue).
IndicGLUE is a natural language understanding benchmark for Indian languages. It contains a wide variety of tasks and covers 11 major Indian languages - Assamese (`as`), Bengali (`bn`), Gujarati (`gu`), Hindi (`hi`), Kannada (`kn`), Malayalam (`ml`), Marathi(`mr`), Oriya(`or`), Panjabi (`pa`), Tamil(`ta`) and Telugu (`te`).
## How to use
There are two steps: (1) loading the IndicGLUE metric relevant to the subset of the dataset being used for evaluation; and (2) calculating the metric.
1. **Loading the relevant IndicGLUE metric** : the subsets of IndicGLUE are the following: `wnli`, `copa`, `sna`, `csqa`, `wstp`, `inltkh`, `bbca`, `cvit-mkb-clsr`, `iitp-mr`, `iitp-pr`, `actsa-sc`, `md`, and`wiki-ner`.
More information about the different subsets of the Indic GLUE dataset can be found on the [IndicGLUE dataset page](https://indicnlp.ai4bharat.org/indic-glue/).
2. **Calculating the metric**: the metric takes two inputs : one list with the predictions of the model to score and one lists of references for each translation for all subsets of the dataset except for `cvit-mkb-clsr`, where each prediction and reference is a vector of floats.
```python
from datasets import load_metric
indic_glue_metric = load_metric('indic_glue', 'wnli')
references = [0, 1]
predictions = [0, 1]
results = indic_glue_metric.compute(predictions=predictions, references=references)
```
## Output values
The output of the metric depends on the IndicGLUE subset chosen, consisting of a dictionary that contains one or several of the following metrics:
`accuracy`: the proportion of correct predictions among the total number of cases processed, with a range between 0 and 1 (see [accuracy](https://huggingface.co/metrics/accuracy) for more information).
`f1`: the harmonic mean of the precision and recall (see [F1 score](https://huggingface.co/metrics/f1) for more information). Its range is 0-1 -- its lowest possible value is 0, if either the precision or the recall is 0, and its highest possible value is 1.0, which means perfect precision and recall.
`precision@10`: the fraction of the true examples among the top 10 predicted examples, with a range between 0 and 1 (see [precision](https://huggingface.co/metrics/precision) for more information).
The `cvit-mkb-clsr` subset returns `precision@10`, the `wiki-ner` subset returns `accuracy` and `f1`, and all other subsets of Indic GLUE return only accuracy.
### Values from popular papers
The [original IndicGlue paper](https://aclanthology.org/2020.findings-emnlp.445.pdf) reported an average accuracy of 0.766 on the dataset, which varies depending on the subset selected.
## Examples
Maximal values for the WNLI subset (which outputs `accuracy`):
```python
from datasets import load_metric
indic_glue_metric = load_metric('indic_glue', 'wnli')
references = [0, 1]
predictions = [0, 1]
results = indic_glue_metric.compute(predictions=predictions, references=references)
print(results)
{'accuracy': 1.0}
```
Minimal values for the Wiki-NER subset (which outputs `accuracy` and `f1`):
```python
>>> from datasets import load_metric
>>> indic_glue_metric = load_metric('indic_glue', 'wiki-ner')
>>> references = [0, 1]
>>> predictions = [1,0]
>>> results = indic_glue_metric.compute(predictions=predictions, references=references)
>>> print(results)
{'accuracy': 1.0, 'f1': 1.0}
```
Partial match for the CVIT-Mann Ki Baat subset (which outputs `precision@10`)
```python
>>> from datasets import load_metric
>>> indic_glue_metric = load_metric('indic_glue', 'cvit-mkb-clsr')
>>> references = [[0.5, 0.5, 0.5], [0.1, 0.2, 0.3]]
>>> predictions = [[0.5, 0.5, 0.5], [0.1, 0.2, 0.3]]
>>> results = indic_glue_metric.compute(predictions=predictions, references=references)
>>> print(results)
{'precision@10': 1.0}
```
## Limitations and bias
This metric works only with datasets that have the same format as the [IndicGLUE dataset](https://huggingface.co/datasets/glue).
## Citation
```bibtex
@inproceedings{kakwani2020indicnlpsuite,
title={{IndicNLPSuite: Monolingual Corpora, Evaluation Benchmarks and Pre-trained Multilingual Language Models for Indian Languages}},
author={Divyanshu Kakwani and Anoop Kunchukuttan and Satish Golla and Gokul N.C. and Avik Bhattacharyya and Mitesh M. Khapra and Pratyush Kumar},
year={2020},
booktitle={Findings of EMNLP},
}
```
## Further References
- [IndicNLP website](https://indicnlp.ai4bharat.org/home/)
-
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/indic_glue/indic_glue.py
|
# Copyright 2020 The HuggingFace Datasets Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
""" IndicGLUE benchmark metric. """
import numpy as np
from scipy.spatial.distance import cdist
from sklearn.metrics import f1_score
import datasets
_CITATION = """\
@inproceedings{kakwani2020indicnlpsuite,
title={{IndicNLPSuite: Monolingual Corpora, Evaluation Benchmarks and Pre-trained Multilingual Language Models for Indian Languages}},
author={Divyanshu Kakwani and Anoop Kunchukuttan and Satish Golla and Gokul N.C. and Avik Bhattacharyya and Mitesh M. Khapra and Pratyush Kumar},
year={2020},
booktitle={Findings of EMNLP},
}
"""
_DESCRIPTION = """\
IndicGLUE is a natural language understanding benchmark for Indian languages. It contains a wide
variety of tasks and covers 11 major Indian languages - as, bn, gu, hi, kn, ml, mr, or, pa, ta, te.
"""
_KWARGS_DESCRIPTION = """
Compute IndicGLUE evaluation metric associated to each IndicGLUE dataset.
Args:
predictions: list of predictions to score (as int64),
except for 'cvit-mkb-clsr' where each prediction is a vector (of float32).
references: list of ground truth labels corresponding to the predictions (as int64),
except for 'cvit-mkb-clsr' where each reference is a vector (of float32).
Returns: depending on the IndicGLUE subset, one or several of:
"accuracy": Accuracy
"f1": F1 score
"precision": Precision@10
Examples:
>>> indic_glue_metric = datasets.load_metric('indic_glue', 'wnli') # 'wnli' or any of ["copa", "sna", "csqa", "wstp", "inltkh", "bbca", "iitp-mr", "iitp-pr", "actsa-sc", "md"]
>>> references = [0, 1]
>>> predictions = [0, 1]
>>> results = indic_glue_metric.compute(predictions=predictions, references=references)
>>> print(results)
{'accuracy': 1.0}
>>> indic_glue_metric = datasets.load_metric('indic_glue', 'wiki-ner')
>>> references = [0, 1]
>>> predictions = [0, 1]
>>> results = indic_glue_metric.compute(predictions=predictions, references=references)
>>> print(results)
{'accuracy': 1.0, 'f1': 1.0}
>>> indic_glue_metric = datasets.load_metric('indic_glue', 'cvit-mkb-clsr')
>>> references = [[0.5, 0.5, 0.5], [0.1, 0.2, 0.3]]
>>> predictions = [[0.5, 0.5, 0.5], [0.1, 0.2, 0.3]]
>>> results = indic_glue_metric.compute(predictions=predictions, references=references)
>>> print(results)
{'precision@10': 1.0}
"""
def simple_accuracy(preds, labels):
return float((preds == labels).mean())
def acc_and_f1(preds, labels):
acc = simple_accuracy(preds, labels)
f1 = float(f1_score(y_true=labels, y_pred=preds))
return {
"accuracy": acc,
"f1": f1,
}
def precision_at_10(en_sentvecs, in_sentvecs):
en_sentvecs = np.array(en_sentvecs)
in_sentvecs = np.array(in_sentvecs)
n = en_sentvecs.shape[0]
# mean centering
en_sentvecs = en_sentvecs - np.mean(en_sentvecs, axis=0)
in_sentvecs = in_sentvecs - np.mean(in_sentvecs, axis=0)
sim = cdist(en_sentvecs, in_sentvecs, "cosine")
actual = np.array(range(n))
preds = sim.argsort(axis=1)[:, :10]
matches = np.any(preds == actual[:, None], axis=1)
return float(matches.mean())
@datasets.utils.file_utils.add_start_docstrings(_DESCRIPTION, _KWARGS_DESCRIPTION)
class IndicGlue(datasets.Metric):
def _info(self):
if self.config_name not in [
"wnli",
"copa",
"sna",
"csqa",
"wstp",
"inltkh",
"bbca",
"cvit-mkb-clsr",
"iitp-mr",
"iitp-pr",
"actsa-sc",
"md",
"wiki-ner",
]:
raise KeyError(
"You should supply a configuration name selected in "
'["wnli", "copa", "sna", "csqa", "wstp", "inltkh", "bbca", '
'"cvit-mkb-clsr", "iitp-mr", "iitp-pr", "actsa-sc", "md", '
'"wiki-ner"]'
)
return datasets.MetricInfo(
description=_DESCRIPTION,
citation=_CITATION,
inputs_description=_KWARGS_DESCRIPTION,
features=datasets.Features(
{
"predictions": datasets.Value("int64")
if self.config_name != "cvit-mkb-clsr"
else datasets.Sequence(datasets.Value("float32")),
"references": datasets.Value("int64")
if self.config_name != "cvit-mkb-clsr"
else datasets.Sequence(datasets.Value("float32")),
}
),
codebase_urls=[],
reference_urls=[],
format="numpy" if self.config_name != "cvit-mkb-clsr" else None,
)
def _compute(self, predictions, references):
if self.config_name == "cvit-mkb-clsr":
return {"precision@10": precision_at_10(predictions, references)}
elif self.config_name in ["wiki-ner"]:
return acc_and_f1(predictions, references)
elif self.config_name in [
"wnli",
"copa",
"sna",
"csqa",
"wstp",
"inltkh",
"bbca",
"iitp-mr",
"iitp-pr",
"actsa-sc",
"md",
]:
return {"accuracy": simple_accuracy(predictions, references)}
else:
raise KeyError(
"You should supply a configuration name selected in "
'["wnli", "copa", "sna", "csqa", "wstp", "inltkh", "bbca", '
'"cvit-mkb-clsr", "iitp-mr", "iitp-pr", "actsa-sc", "md", '
'"wiki-ner"]'
)
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/coval/coval.py
|
# Copyright 2020 The HuggingFace Datasets Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
""" CoVal metric. """
import coval # From: git+https://github.com/ns-moosavi/coval.git # noqa: F401
from coval.conll import reader, util
from coval.eval import evaluator
import datasets
logger = datasets.logging.get_logger(__name__)
_CITATION = """\
@InProceedings{moosavi2019minimum,
author = { Nafise Sadat Moosavi, Leo Born, Massimo Poesio and Michael Strube},
title = {Using Automatically Extracted Minimum Spans to Disentangle Coreference Evaluation from Boundary Detection},
year = {2019},
booktitle = {Proceedings of the 57th Annual Meeting of
the Association for Computational Linguistics (Volume 1: Long Papers)},
publisher = {Association for Computational Linguistics},
address = {Florence, Italy},
}
@inproceedings{10.3115/1072399.1072405,
author = {Vilain, Marc and Burger, John and Aberdeen, John and Connolly, Dennis and Hirschman, Lynette},
title = {A Model-Theoretic Coreference Scoring Scheme},
year = {1995},
isbn = {1558604022},
publisher = {Association for Computational Linguistics},
address = {USA},
url = {https://doi.org/10.3115/1072399.1072405},
doi = {10.3115/1072399.1072405},
booktitle = {Proceedings of the 6th Conference on Message Understanding},
pages = {45–52},
numpages = {8},
location = {Columbia, Maryland},
series = {MUC6 ’95}
}
@INPROCEEDINGS{Bagga98algorithmsfor,
author = {Amit Bagga and Breck Baldwin},
title = {Algorithms for Scoring Coreference Chains},
booktitle = {In The First International Conference on Language Resources and Evaluation Workshop on Linguistics Coreference},
year = {1998},
pages = {563--566}
}
@INPROCEEDINGS{Luo05oncoreference,
author = {Xiaoqiang Luo},
title = {On coreference resolution performance metrics},
booktitle = {In Proc. of HLT/EMNLP},
year = {2005},
pages = {25--32},
publisher = {URL}
}
@inproceedings{moosavi-strube-2016-coreference,
title = "Which Coreference Evaluation Metric Do You Trust? A Proposal for a Link-based Entity Aware Metric",
author = "Moosavi, Nafise Sadat and
Strube, Michael",
booktitle = "Proceedings of the 54th Annual Meeting of the Association for Computational Linguistics (Volume 1: Long Papers)",
month = aug,
year = "2016",
address = "Berlin, Germany",
publisher = "Association for Computational Linguistics",
url = "https://www.aclweb.org/anthology/P16-1060",
doi = "10.18653/v1/P16-1060",
pages = "632--642",
}
"""
_DESCRIPTION = """\
CoVal is a coreference evaluation tool for the CoNLL and ARRAU datasets which
implements of the common evaluation metrics including MUC [Vilain et al, 1995],
B-cubed [Bagga and Baldwin, 1998], CEAFe [Luo et al., 2005],
LEA [Moosavi and Strube, 2016] and the averaged CoNLL score
(the average of the F1 values of MUC, B-cubed and CEAFe)
[Denis and Baldridge, 2009a; Pradhan et al., 2011].
This wrapper of CoVal currently only work with CoNLL line format:
The CoNLL format has one word per line with all the annotation for this word in column separated by spaces:
Column Type Description
1 Document ID This is a variation on the document filename
2 Part number Some files are divided into multiple parts numbered as 000, 001, 002, ... etc.
3 Word number
4 Word itself This is the token as segmented/tokenized in the Treebank. Initially the *_skel file contain the placeholder [WORD] which gets replaced by the actual token from the Treebank which is part of the OntoNotes release.
5 Part-of-Speech
6 Parse bit This is the bracketed structure broken before the first open parenthesis in the parse, and the word/part-of-speech leaf replaced with a *. The full parse can be created by substituting the asterix with the "([pos] [word])" string (or leaf) and concatenating the items in the rows of that column.
7 Predicate lemma The predicate lemma is mentioned for the rows for which we have semantic role information. All other rows are marked with a "-"
8 Predicate Frameset ID This is the PropBank frameset ID of the predicate in Column 7.
9 Word sense This is the word sense of the word in Column 3.
10 Speaker/Author This is the speaker or author name where available. Mostly in Broadcast Conversation and Web Log data.
11 Named Entities These columns identifies the spans representing various named entities.
12:N Predicate Arguments There is one column each of predicate argument structure information for the predicate mentioned in Column 7.
N Coreference Coreference chain information encoded in a parenthesis structure.
More informations on the format can be found here (section "*_conll File Format"): http://www.conll.cemantix.org/2012/data.html
Details on the evaluation on CoNLL can be found here: https://github.com/ns-moosavi/coval/blob/master/conll/README.md
CoVal code was written by @ns-moosavi.
Some parts are borrowed from https://github.com/clarkkev/deep-coref/blob/master/evaluation.py
The test suite is taken from https://github.com/conll/reference-coreference-scorers/
Mention evaluation and the test suite are added by @andreasvc.
Parsing CoNLL files is developed by Leo Born.
"""
_KWARGS_DESCRIPTION = """
Calculates coreference evaluation metrics.
Args:
predictions: list of sentences. Each sentence is a list of word predictions to score in the CoNLL format.
Each prediction is a word with its annotations as a string made of columns joined with spaces.
Only columns 4, 5, 6 and the last column are used (word, POS, Pars and coreference annotation)
See the details on the format in the description of the metric.
references: list of sentences. Each sentence is a list of word reference to score in the CoNLL format.
Each reference is a word with its annotations as a string made of columns joined with spaces.
Only columns 4, 5, 6 and the last column are used (word, POS, Pars and coreference annotation)
See the details on the format in the description of the metric.
keep_singletons: After extracting all mentions of key or system files,
mentions whose corresponding coreference chain is of size one,
are considered as singletons. The default evaluation mode will include
singletons in evaluations if they are included in the key or the system files.
By setting 'keep_singletons=False', all singletons in the key and system files
will be excluded from the evaluation.
NP_only: Most of the recent coreference resolvers only resolve NP mentions and
leave out the resolution of VPs. By setting the 'NP_only' option, the scorer will only evaluate the resolution of NPs.
min_span: By setting 'min_span', the scorer reports the results based on automatically detected minimum spans.
Minimum spans are determined using the MINA algorithm.
Returns:
'mentions': mentions
'muc': MUC metric [Vilain et al, 1995]
'bcub': B-cubed [Bagga and Baldwin, 1998]
'ceafe': CEAFe [Luo et al., 2005]
'lea': LEA [Moosavi and Strube, 2016]
'conll_score': averaged CoNLL score (the average of the F1 values of MUC, B-cubed and CEAFe)
Examples:
>>> coval = datasets.load_metric('coval')
>>> words = ['bc/cctv/00/cctv_0005 0 0 Thank VBP (TOP(S(VP* thank 01 1 Xu_li * (V*) * -',
... 'bc/cctv/00/cctv_0005 0 1 you PRP (NP*) - - - Xu_li * (ARG1*) (ARG0*) (116)',
... 'bc/cctv/00/cctv_0005 0 2 everyone NN (NP*) - - - Xu_li * (ARGM-DIS*) * (116)',
... 'bc/cctv/00/cctv_0005 0 3 for IN (PP* - - - Xu_li * (ARG2* * -',
... 'bc/cctv/00/cctv_0005 0 4 watching VBG (S(VP*)))) watch 01 1 Xu_li * *) (V*) -',
... 'bc/cctv/00/cctv_0005 0 5 . . *)) - - - Xu_li * * * -']
>>> references = [words]
>>> predictions = [words]
>>> results = coval.compute(predictions=predictions, references=references)
>>> print(results) # doctest:+ELLIPSIS
{'mentions/recall': 1.0,[...] 'conll_score': 100.0}
"""
def get_coref_infos(
key_lines, sys_lines, NP_only=False, remove_nested=False, keep_singletons=True, min_span=False, doc="dummy_doc"
):
key_doc_lines = {doc: key_lines}
sys_doc_lines = {doc: sys_lines}
doc_coref_infos = {}
key_nested_coref_num = 0
sys_nested_coref_num = 0
key_removed_nested_clusters = 0
sys_removed_nested_clusters = 0
key_singletons_num = 0
sys_singletons_num = 0
key_clusters, singletons_num = reader.get_doc_mentions(doc, key_doc_lines[doc], keep_singletons)
key_singletons_num += singletons_num
if NP_only or min_span:
key_clusters = reader.set_annotated_parse_trees(key_clusters, key_doc_lines[doc], NP_only, min_span)
sys_clusters, singletons_num = reader.get_doc_mentions(doc, sys_doc_lines[doc], keep_singletons)
sys_singletons_num += singletons_num
if NP_only or min_span:
sys_clusters = reader.set_annotated_parse_trees(sys_clusters, key_doc_lines[doc], NP_only, min_span)
if remove_nested:
nested_mentions, removed_clusters = reader.remove_nested_coref_mentions(key_clusters, keep_singletons)
key_nested_coref_num += nested_mentions
key_removed_nested_clusters += removed_clusters
nested_mentions, removed_clusters = reader.remove_nested_coref_mentions(sys_clusters, keep_singletons)
sys_nested_coref_num += nested_mentions
sys_removed_nested_clusters += removed_clusters
sys_mention_key_cluster = reader.get_mention_assignments(sys_clusters, key_clusters)
key_mention_sys_cluster = reader.get_mention_assignments(key_clusters, sys_clusters)
doc_coref_infos[doc] = (key_clusters, sys_clusters, key_mention_sys_cluster, sys_mention_key_cluster)
if remove_nested:
logger.info(
"Number of removed nested coreferring mentions in the key "
f"annotation: {key_nested_coref_num}; and system annotation: {sys_nested_coref_num}"
)
logger.info(
"Number of resulting singleton clusters in the key "
f"annotation: {key_removed_nested_clusters}; and system annotation: {sys_removed_nested_clusters}"
)
if not keep_singletons:
logger.info(
f"{key_singletons_num:d} and {sys_singletons_num:d} singletons are removed from the key and system "
"files, respectively"
)
return doc_coref_infos
def evaluate(key_lines, sys_lines, metrics, NP_only, remove_nested, keep_singletons, min_span):
doc_coref_infos = get_coref_infos(key_lines, sys_lines, NP_only, remove_nested, keep_singletons, min_span)
output_scores = {}
conll = 0
conll_subparts_num = 0
for name, metric in metrics:
recall, precision, f1 = evaluator.evaluate_documents(doc_coref_infos, metric, beta=1)
if name in ["muc", "bcub", "ceafe"]:
conll += f1
conll_subparts_num += 1
output_scores.update({f"{name}/recall": recall, f"{name}/precision": precision, f"{name}/f1": f1})
logger.info(
name.ljust(10),
f"Recall: {recall * 100:.2f}",
f" Precision: {precision * 100:.2f}",
f" F1: {f1 * 100:.2f}",
)
if conll_subparts_num == 3:
conll = (conll / 3) * 100
logger.info(f"CoNLL score: {conll:.2f}")
output_scores.update({"conll_score": conll})
return output_scores
def check_gold_parse_annotation(key_lines):
has_gold_parse = False
for line in key_lines:
if not line.startswith("#"):
if len(line.split()) > 6:
parse_col = line.split()[5]
if not parse_col == "-":
has_gold_parse = True
break
else:
break
return has_gold_parse
@datasets.utils.file_utils.add_start_docstrings(_DESCRIPTION, _KWARGS_DESCRIPTION)
class Coval(datasets.Metric):
def _info(self):
return datasets.MetricInfo(
description=_DESCRIPTION,
citation=_CITATION,
inputs_description=_KWARGS_DESCRIPTION,
features=datasets.Features(
{
"predictions": datasets.Sequence(datasets.Value("string")),
"references": datasets.Sequence(datasets.Value("string")),
}
),
codebase_urls=["https://github.com/ns-moosavi/coval"],
reference_urls=[
"https://github.com/ns-moosavi/coval",
"https://www.aclweb.org/anthology/P16-1060",
"http://www.conll.cemantix.org/2012/data.html",
],
)
def _compute(
self, predictions, references, keep_singletons=True, NP_only=False, min_span=False, remove_nested=False
):
allmetrics = [
("mentions", evaluator.mentions),
("muc", evaluator.muc),
("bcub", evaluator.b_cubed),
("ceafe", evaluator.ceafe),
("lea", evaluator.lea),
]
if min_span:
has_gold_parse = util.check_gold_parse_annotation(references)
if not has_gold_parse:
raise NotImplementedError("References should have gold parse annotation to use 'min_span'.")
# util.parse_key_file(key_file)
# key_file = key_file + ".parsed"
score = evaluate(
key_lines=references,
sys_lines=predictions,
metrics=allmetrics,
NP_only=NP_only,
remove_nested=remove_nested,
keep_singletons=keep_singletons,
min_span=min_span,
)
return score
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/coval/README.md
|
# Metric Card for COVAL
## Metric description
CoVal is a coreference evaluation tool for the [CoNLL](https://huggingface.co/datasets/conll2003) and [ARRAU](https://catalog.ldc.upenn.edu/LDC2013T22) datasets which implements of the common evaluation metrics including MUC [Vilain et al, 1995](https://aclanthology.org/M95-1005.pdf), B-cubed [Bagga and Baldwin, 1998](https://citeseerx.ist.psu.edu/viewdoc/download?doi=10.1.1.34.2578&rep=rep1&type=pdf), CEAFe [Luo et al., 2005](https://aclanthology.org/H05-1004.pdf), LEA [Moosavi and Strube, 2016](https://aclanthology.org/P16-1060.pdf) and the averaged CoNLL score (the average of the F1 values of MUC, B-cubed and CEAFe).
CoVal code was written by [`@ns-moosavi`](https://github.com/ns-moosavi), with some parts borrowed from [Deep Coref](https://github.com/clarkkev/deep-coref/blob/master/evaluation.py). The test suite is taken from the [official CoNLL code](https://github.com/conll/reference-coreference-scorers/), with additions by [`@andreasvc`](https://github.com/andreasvc) and file parsing developed by Leo Born.
## How to use
The metric takes two lists of sentences as input: one representing `predictions` and `references`, with the sentences consisting of words in the CoNLL format (see the [Limitations and bias](#Limitations-and-bias) section below for more details on the CoNLL format).
```python
from datasets import load_metric
coval = load_metric('coval')
words = ['bc/cctv/00/cctv_0005 0 0 Thank VBP (TOP(S(VP* thank 01 1 Xu_li * (V*) * -',
... 'bc/cctv/00/cctv_0005 0 1 you PRP (NP*) - - - Xu_li * (ARG1*) (ARG0*) (116)',
... 'bc/cctv/00/cctv_0005 0 2 everyone NN (NP*) - - - Xu_li * (ARGM-DIS*) * (116)',
... 'bc/cctv/00/cctv_0005 0 3 for IN (PP* - - - Xu_li * (ARG2* * -',
... 'bc/cctv/00/cctv_0005 0 4 watching VBG (S(VP*)))) watch 01 1 Xu_li * *) (V*) -',
... 'bc/cctv/00/cctv_0005 0 5 . . *)) - - - Xu_li * * * -']
references = [words]
predictions = [words]
results = coval.compute(predictions=predictions, references=references)
```
It also has several optional arguments:
`keep_singletons`: After extracting all mentions of key or system file mentions whose corresponding coreference chain is of size one are considered as singletons. The default evaluation mode will include singletons in evaluations if they are included in the key or the system files. By setting `keep_singletons=False`, all singletons in the key and system files will be excluded from the evaluation.
`NP_only`: Most of the recent coreference resolvers only resolve NP mentions and leave out the resolution of VPs. By setting the `NP_only` option, the scorer will only evaluate the resolution of NPs.
`min_span`: By setting `min_span`, the scorer reports the results based on automatically detected minimum spans. Minimum spans are determined using the [MINA algorithm](https://arxiv.org/pdf/1906.06703.pdf).
## Output values
The metric outputs a dictionary with the following key-value pairs:
`mentions`: number of mentions, ranges from 0-1
`muc`: MUC metric, which expresses performance in terms of recall and precision, ranging from 0-1.
`bcub`: B-cubed metric, which is the averaged precision of all items in the distribution, ranging from 0-1.
`ceafe`: CEAFe (Constrained Entity Alignment F-Measure) is computed by aligning reference and system entities with the constraint that a reference entity is aligned with at most one reference entity. It ranges from 0-1
`lea`: LEA is a Link-Based Entity-Aware metric which, for each entity, considers how important the entity is and how well it is resolved. It ranges from 0-1.
`conll_score`: averaged CoNLL score (the average of the F1 values of `muc`, `bcub` and `ceafe`), ranging from 0 to 100.
### Values from popular papers
Given that many of the metrics returned by COVAL come from different sources, is it hard to cite reference values for all of them.
The CoNLL score is used to track progress on different datasets such as the [ARRAU corpus](https://paperswithcode.com/sota/coreference-resolution-on-the-arrau-corpus) and [CoNLL 2012](https://paperswithcode.com/sota/coreference-resolution-on-conll-2012).
## Examples
Maximal values
```python
from datasets import load_metric
coval = load_metric('coval')
words = ['bc/cctv/00/cctv_0005 0 0 Thank VBP (TOP(S(VP* thank 01 1 Xu_li * (V*) * -',
... 'bc/cctv/00/cctv_0005 0 1 you PRP (NP*) - - - Xu_li * (ARG1*) (ARG0*) (116)',
... 'bc/cctv/00/cctv_0005 0 2 everyone NN (NP*) - - - Xu_li * (ARGM-DIS*) * (116)',
... 'bc/cctv/00/cctv_0005 0 3 for IN (PP* - - - Xu_li * (ARG2* * -',
... 'bc/cctv/00/cctv_0005 0 4 watching VBG (S(VP*)))) watch 01 1 Xu_li * *) (V*) -',
... 'bc/cctv/00/cctv_0005 0 5 . . *)) - - - Xu_li * * * -']
references = [words]
predictions = [words]
results = coval.compute(predictions=predictions, references=references)
print(results)
{'mentions/recall': 1.0, 'mentions/precision': 1.0, 'mentions/f1': 1.0, 'muc/recall': 1.0, 'muc/precision': 1.0, 'muc/f1': 1.0, 'bcub/recall': 1.0, 'bcub/precision': 1.0, 'bcub/f1': 1.0, 'ceafe/recall': 1.0, 'ceafe/precision': 1.0, 'ceafe/f1': 1.0, 'lea/recall': 1.0, 'lea/precision': 1.0, 'lea/f1': 1.0, 'conll_score': 100.0}
```
## Limitations and bias
This wrapper of CoVal currently only works with [CoNLL line format](https://huggingface.co/datasets/conll2003), which has one word per line with all the annotation for this word in column separated by spaces:
| Column | Type | Description |
|:-------|:----------------------|:--------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------|
| 1 | Document ID | This is a variation on the document filename |
| 2 | Part number | Some files are divided into multiple parts numbered as 000, 001, 002, ... etc. |
| 3 | Word number | |
| 4 | Word | This is the token as segmented/tokenized in the Treebank. Initially the *_skel file contain the placeholder [WORD] which gets replaced by the actual token from the Treebank which is part of the OntoNotes release. |
| 5 | Part-of-Speech | |
| 6 | Parse bit | This is the bracketed structure broken before the first open parenthesis in the parse, and the word/part-of-speech leaf replaced with a *. The full parse can be created by substituting the asterix with the "([pos] [word])" string (or leaf) and concatenating the items in the rows of that column. |
| 7 | Predicate lemma | The predicate lemma is mentioned for the rows for which we have semantic role information. All other rows are marked with a "-". |
| 8 | Predicate Frameset ID | This is the PropBank frameset ID of the predicate in Column 7. |
| 9 | Word sense | This is the word sense of the word in Column 3. |
| 10 | Speaker/Author | This is the speaker or author name where available. Mostly in Broadcast Conversation and Web Log data. |
| 11 | Named Entities | These columns identifies the spans representing various named entities. |
| 12:N | Predicate Arguments | There is one column each of predicate argument structure information for the predicate mentioned in Column 7. |
| N | Coreference | Coreference chain information encoded in a parenthesis structure. |
## Citations
```bibtex
@InProceedings{moosavi2019minimum,
author = { Nafise Sadat Moosavi, Leo Born, Massimo Poesio and Michael Strube},
title = {Using Automatically Extracted Minimum Spans to Disentangle Coreference Evaluation from Boundary Detection},
year = {2019},
booktitle = {Proceedings of the 57th Annual Meeting of
the Association for Computational Linguistics (Volume 1: Long Papers)},
publisher = {Association for Computational Linguistics},
address = {Florence, Italy},
}
```
```bibtex
@inproceedings{10.3115/1072399.1072405,
author = {Vilain, Marc and Burger, John and Aberdeen, John and Connolly, Dennis and Hirschman, Lynette},
title = {A Model-Theoretic Coreference Scoring Scheme},
year = {1995},
isbn = {1558604022},
publisher = {Association for Computational Linguistics},
address = {USA},
url = {https://doi.org/10.3115/1072399.1072405},
doi = {10.3115/1072399.1072405},
booktitle = {Proceedings of the 6th Conference on Message Understanding},
pages = {45–52},
numpages = {8},
location = {Columbia, Maryland},
series = {MUC6 ’95}
}
```
```bibtex
@INPROCEEDINGS{Bagga98algorithmsfor,
author = {Amit Bagga and Breck Baldwin},
title = {Algorithms for Scoring Coreference Chains},
booktitle = {In The First International Conference on Language Resources and Evaluation Workshop on Linguistics Coreference},
year = {1998},
pages = {563--566}
}
```
```bibtex
@INPROCEEDINGS{Luo05oncoreference,
author = {Xiaoqiang Luo},
title = {On coreference resolution performance metrics},
booktitle = {In Proc. of HLT/EMNLP},
year = {2005},
pages = {25--32},
publisher = {URL}
}
```
```bibtex
@inproceedings{moosavi-strube-2016-coreference,
title = "Which Coreference Evaluation Metric Do You Trust? A Proposal for a Link-based Entity Aware Metric",
author = "Moosavi, Nafise Sadat and
Strube, Michael",
booktitle = "Proceedings of the 54th Annual Meeting of the Association for Computational Linguistics (Volume 1: Long Papers)",
month = aug,
year = "2016",
address = "Berlin, Germany",
publisher = "Association for Computational Linguistics",
url = "https://www.aclweb.org/anthology/P16-1060",
doi = "10.18653/v1/P16-1060",
pages = "632--642",
}
```
## Further References
- [CoNLL 2012 Task Description](http://www.conll.cemantix.org/2012/data.html): for information on the format (section "*_conll File Format")
- [CoNLL Evaluation details](https://github.com/ns-moosavi/coval/blob/master/conll/README.md)
- [Hugging Face - Neural Coreference Resolution (Neuralcoref)](https://huggingface.co/coref/)
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/meteor/README.md
|
# Metric Card for METEOR
## Metric description
METEOR (Metric for Evaluation of Translation with Explicit ORdering) is a machine translation evaluation metric, which is calculated based on the harmonic mean of precision and recall, with recall weighted more than precision.
METEOR is based on a generalized concept of unigram matching between the machine-produced translation and human-produced reference translations. Unigrams can be matched based on their surface forms, stemmed forms, and meanings. Once all generalized unigram matches between the two strings have been found, METEOR computes a score for this matching using a combination of unigram-precision, unigram-recall, and a measure of fragmentation that is designed to directly capture how well-ordered the matched words in the machine translation are in relation to the reference.
## How to use
METEOR has two mandatory arguments:
`predictions`: a list of predictions to score. Each prediction should be a string with tokens separated by spaces.
`references`: a list of references for each prediction. Each reference should be a string with tokens separated by spaces.
It also has several optional parameters:
`alpha`: Parameter for controlling relative weights of precision and recall. The default value is `0.9`.
`beta`: Parameter for controlling shape of penalty as a function of fragmentation. The default value is `3`.
`gamma`: The relative weight assigned to fragmentation penalty. The default is `0.5`.
Refer to the [METEOR paper](https://aclanthology.org/W05-0909.pdf) for more information about parameter values and ranges.
```python
>>> from datasets import load_metric
>>> meteor = load_metric('meteor')
>>> predictions = ["It is a guide to action which ensures that the military always obeys the commands of the party"]
>>> references = ["It is a guide to action that ensures that the military will forever heed Party commands"]
>>> results = meteor.compute(predictions=predictions, references=references)
```
## Output values
The metric outputs a dictionary containing the METEOR score. Its values range from 0 to 1.
### Values from popular papers
The [METEOR paper](https://aclanthology.org/W05-0909.pdf) does not report METEOR score values for different models, but it does report that METEOR gets an R correlation value of 0.347 with human evaluation on the Arabic data and 0.331 on the Chinese data.
## Examples
Maximal values :
```python
>>> from datasets import load_metric
>>> meteor = load_metric('meteor')
>>> predictions = ["It is a guide to action which ensures that the military always obeys the commands of the party"]
>>> references = ["It is a guide to action which ensures that the military always obeys the commands of the party"]
>>> results = meteor.compute(predictions=predictions, references=references)
>>> print(round(results['meteor'], 2))
1.0
```
Minimal values:
```python
>>> from datasets import load_metric
>>> meteor = load_metric('meteor')
>>> predictions = ["It is a guide to action which ensures that the military always obeys the commands of the party"]
>>> references = ["Hello world"]
>>> results = meteor.compute(predictions=predictions, references=references)
>>> print(round(results['meteor'], 2))
0.0
```
Partial match:
```python
>>> from datasets import load_metric
>>> meteor = load_metric('meteor')
>>> predictions = ["It is a guide to action which ensures that the military always obeys the commands of the party"]
>>> references = ["It is a guide to action that ensures that the military will forever heed Party commands"]
>>> results = meteor.compute(predictions=predictions, references=references)
>>> print(round(results['meteor'], 2))
0.69
```
## Limitations and bias
While the correlation between METEOR and human judgments was measured for Chinese and Arabic and found to be significant, further experimentation is needed to check its correlation for other languages.
Furthermore, while the alignment and matching done in METEOR is based on unigrams, using multiple word entities (e.g. bigrams) could contribute to improving its accuracy -- this has been proposed in [more recent publications](https://www.cs.cmu.edu/~alavie/METEOR/pdf/meteor-naacl-2010.pdf) on the subject.
## Citation
```bibtex
@inproceedings{banarjee2005,
title = {{METEOR}: An Automatic Metric for {MT} Evaluation with Improved Correlation with Human Judgments},
author = {Banerjee, Satanjeev and Lavie, Alon},
booktitle = {Proceedings of the {ACL} Workshop on Intrinsic and Extrinsic Evaluation Measures for Machine Translation and/or Summarization},
month = jun,
year = {2005},
address = {Ann Arbor, Michigan},
publisher = {Association for Computational Linguistics},
url = {https://www.aclweb.org/anthology/W05-0909},
pages = {65--72},
}
```
## Further References
- [METEOR -- Wikipedia](https://en.wikipedia.org/wiki/METEOR)
- [METEOR score -- NLTK](https://www.nltk.org/_modules/nltk/translate/meteor_score.html)
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/meteor/meteor.py
|
# Copyright 2020 The HuggingFace Datasets Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
""" METEOR metric. """
import importlib.metadata
import numpy as np
from nltk.translate import meteor_score
from packaging import version
import datasets
NLTK_VERSION = version.parse(importlib.metadata.version("nltk"))
if NLTK_VERSION >= version.Version("3.6.4"):
from nltk import word_tokenize
_CITATION = """\
@inproceedings{banarjee2005,
title = {{METEOR}: An Automatic Metric for {MT} Evaluation with Improved Correlation with Human Judgments},
author = {Banerjee, Satanjeev and Lavie, Alon},
booktitle = {Proceedings of the {ACL} Workshop on Intrinsic and Extrinsic Evaluation Measures for Machine Translation and/or Summarization},
month = jun,
year = {2005},
address = {Ann Arbor, Michigan},
publisher = {Association for Computational Linguistics},
url = {https://www.aclweb.org/anthology/W05-0909},
pages = {65--72},
}
"""
_DESCRIPTION = """\
METEOR, an automatic metric for machine translation evaluation
that is based on a generalized concept of unigram matching between the
machine-produced translation and human-produced reference translations.
Unigrams can be matched based on their surface forms, stemmed forms,
and meanings; furthermore, METEOR can be easily extended to include more
advanced matching strategies. Once all generalized unigram matches
between the two strings have been found, METEOR computes a score for
this matching using a combination of unigram-precision, unigram-recall, and
a measure of fragmentation that is designed to directly capture how
well-ordered the matched words in the machine translation are in relation
to the reference.
METEOR gets an R correlation value of 0.347 with human evaluation on the Arabic
data and 0.331 on the Chinese data. This is shown to be an improvement on
using simply unigram-precision, unigram-recall and their harmonic F1
combination.
"""
_KWARGS_DESCRIPTION = """
Computes METEOR score of translated segments against one or more references.
Args:
predictions: list of predictions to score. Each prediction
should be a string with tokens separated by spaces.
references: list of reference for each prediction. Each
reference should be a string with tokens separated by spaces.
alpha: Parameter for controlling relative weights of precision and recall. default: 0.9
beta: Parameter for controlling shape of penalty as a function of fragmentation. default: 3
gamma: Relative weight assigned to fragmentation penalty. default: 0.5
Returns:
'meteor': meteor score.
Examples:
>>> meteor = datasets.load_metric('meteor')
>>> predictions = ["It is a guide to action which ensures that the military always obeys the commands of the party"]
>>> references = ["It is a guide to action that ensures that the military will forever heed Party commands"]
>>> results = meteor.compute(predictions=predictions, references=references)
>>> print(round(results["meteor"], 4))
0.6944
"""
@datasets.utils.file_utils.add_start_docstrings(_DESCRIPTION, _KWARGS_DESCRIPTION)
class Meteor(datasets.Metric):
def _info(self):
return datasets.MetricInfo(
description=_DESCRIPTION,
citation=_CITATION,
inputs_description=_KWARGS_DESCRIPTION,
features=datasets.Features(
{
"predictions": datasets.Value("string", id="sequence"),
"references": datasets.Value("string", id="sequence"),
}
),
codebase_urls=["https://github.com/nltk/nltk/blob/develop/nltk/translate/meteor_score.py"],
reference_urls=[
"https://www.nltk.org/api/nltk.translate.html#module-nltk.translate.meteor_score",
"https://en.wikipedia.org/wiki/METEOR",
],
)
def _download_and_prepare(self, dl_manager):
import nltk
nltk.download("wordnet")
if NLTK_VERSION >= version.Version("3.6.5"):
nltk.download("punkt")
if NLTK_VERSION >= version.Version("3.6.6"):
nltk.download("omw-1.4")
def _compute(self, predictions, references, alpha=0.9, beta=3, gamma=0.5):
if NLTK_VERSION >= version.Version("3.6.5"):
scores = [
meteor_score.single_meteor_score(
word_tokenize(ref), word_tokenize(pred), alpha=alpha, beta=beta, gamma=gamma
)
for ref, pred in zip(references, predictions)
]
else:
scores = [
meteor_score.single_meteor_score(ref, pred, alpha=alpha, beta=beta, gamma=gamma)
for ref, pred in zip(references, predictions)
]
return {"meteor": np.mean(scores)}
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/bertscore/README.md
|
# Metric Card for BERT Score
## Metric description
BERTScore is an automatic evaluation metric for text generation that computes a similarity score for each token in the candidate sentence with each token in the reference sentence. It leverages the pre-trained contextual embeddings from [BERT](https://huggingface.co/bert-base-uncased) models and matches words in candidate and reference sentences by cosine similarity.
Moreover, BERTScore computes precision, recall, and F1 measure, which can be useful for evaluating different language generation tasks.
## How to use
BERTScore takes 3 mandatory arguments : `predictions` (a list of string of candidate sentences), `references` (a list of strings or list of list of strings of reference sentences) and either `lang` (a string of two letters indicating the language of the sentences, in [ISO 639-1 format](https://en.wikipedia.org/wiki/List_of_ISO_639-1_codes)) or `model_type` (a string specififying which model to use, according to the BERT specification). The default behavior of the metric is to use the suggested model for the target language when one is specified, otherwise to use the `model_type` indicated.
```python
from datasets import load_metric
bertscore = load_metric("bertscore")
predictions = ["hello there", "general kenobi"]
references = ["hello there", "general kenobi"]
results = bertscore.compute(predictions=predictions, references=references, lang="en")
```
BERTScore also accepts multiple optional arguments:
`num_layers` (int): The layer of representation to use. The default is the number of layers tuned on WMT16 correlation data, which depends on the `model_type` used.
`verbose` (bool): Turn on intermediate status update. The default value is `False`.
`idf` (bool or dict): Use idf weighting; can also be a precomputed idf_dict.
`device` (str): On which the contextual embedding model will be allocated on. If this argument is `None`, the model lives on `cuda:0` if cuda is available.
`nthreads` (int): Number of threads used for computation. The default value is `4`.
`rescale_with_baseline` (bool): Rescale BERTScore with the pre-computed baseline. The default value is `False`.
`batch_size` (int): BERTScore processing batch size, at least one of `model_type` or `lang`. `lang` needs to be specified when `rescale_with_baseline` is `True`.
`baseline_path` (str): Customized baseline file.
`use_fast_tokenizer` (bool): `use_fast` parameter passed to HF tokenizer. The default value is `False`.
## Output values
BERTScore outputs a dictionary with the following values:
`precision`: The [precision](https://huggingface.co/metrics/precision) for each sentence from the `predictions` + `references` lists, which ranges from 0.0 to 1.0.
`recall`: The [recall](https://huggingface.co/metrics/recall) for each sentence from the `predictions` + `references` lists, which ranges from 0.0 to 1.0.
`f1`: The [F1 score](https://huggingface.co/metrics/f1) for each sentence from the `predictions` + `references` lists, which ranges from 0.0 to 1.0.
`hashcode:` The hashcode of the library.
### Values from popular papers
The [original BERTScore paper](https://openreview.net/pdf?id=SkeHuCVFDr) reported average model selection accuracies (Hits@1) on WMT18 hybrid systems for different language pairs, which ranged from 0.004 for `en<->tr` to 0.824 for `en<->de`.
For more recent model performance, see the [metric leaderboard](https://paperswithcode.com/paper/bertscore-evaluating-text-generation-with).
## Examples
Maximal values with the `distilbert-base-uncased` model:
```python
from datasets import load_metric
bertscore = load_metric("bertscore")
predictions = ["hello world", "general kenobi"]
references = ["hello world", "general kenobi"]
results = bertscore.compute(predictions=predictions, references=references, model_type="distilbert-base-uncased")
print(results)
{'precision': [1.0, 1.0], 'recall': [1.0, 1.0], 'f1': [1.0, 1.0], 'hashcode': 'distilbert-base-uncased_L5_no-idf_version=0.3.10(hug_trans=4.10.3)'}
```
Partial match with the `bert-base-uncased` model:
```python
from datasets import load_metric
bertscore = load_metric("bertscore")
predictions = ["hello world", "general kenobi"]
references = ["goodnight moon", "the sun is shining"]
results = bertscore.compute(predictions=predictions, references=references, model_type="distilbert-base-uncased")
print(results)
{'precision': [0.7380737066268921, 0.5584042072296143], 'recall': [0.7380737066268921, 0.5889028906822205], 'f1': [0.7380737066268921, 0.5732481479644775], 'hashcode': 'bert-base-uncased_L5_no-idf_version=0.3.10(hug_trans=4.10.3)'}
```
## Limitations and bias
The [original BERTScore paper](https://openreview.net/pdf?id=SkeHuCVFDr) showed that BERTScore correlates well with human judgment on sentence-level and system-level evaluation, but this depends on the model and language pair selected.
Furthermore, not all languages are supported by the metric -- see the [BERTScore supported language list](https://github.com/google-research/bert/blob/master/multilingual.md#list-of-languages) for more information.
Finally, calculating the BERTScore metric involves downloading the BERT model that is used to compute the score-- the default model for `en`, `roberta-large`, takes over 1.4GB of storage space and downloading it can take a significant amount of time depending on the speed of your internet connection. If this is an issue, choose a smaller model; for instance `distilbert-base-uncased` is 268MB. A full list of compatible models can be found [here](https://docs.google.com/spreadsheets/d/1RKOVpselB98Nnh_EOC4A2BYn8_201tmPODpNWu4w7xI/edit#gid=0).
## Citation
```bibtex
@inproceedings{bert-score,
title={BERTScore: Evaluating Text Generation with BERT},
author={Tianyi Zhang* and Varsha Kishore* and Felix Wu* and Kilian Q. Weinberger and Yoav Artzi},
booktitle={International Conference on Learning Representations},
year={2020},
url={https://openreview.net/forum?id=SkeHuCVFDr}
}
```
## Further References
- [BERTScore Project README](https://github.com/Tiiiger/bert_score#readme)
- [BERTScore ICLR 2020 Poster Presentation](https://iclr.cc/virtual_2020/poster_SkeHuCVFDr.html)
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/bertscore/bertscore.py
|
# Copyright 2020 The HuggingFace Datasets Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
""" BERTScore metric. """
import functools
from contextlib import contextmanager
import bert_score
from packaging import version
import datasets
@contextmanager
def filter_logging_context():
def filter_log(record):
return False if "This IS expected if you are initializing" in record.msg else True
logger = datasets.utils.logging.get_logger("transformers.modeling_utils")
logger.addFilter(filter_log)
try:
yield
finally:
logger.removeFilter(filter_log)
_CITATION = """\
@inproceedings{bert-score,
title={BERTScore: Evaluating Text Generation with BERT},
author={Tianyi Zhang* and Varsha Kishore* and Felix Wu* and Kilian Q. Weinberger and Yoav Artzi},
booktitle={International Conference on Learning Representations},
year={2020},
url={https://openreview.net/forum?id=SkeHuCVFDr}
}
"""
_DESCRIPTION = """\
BERTScore leverages the pre-trained contextual embeddings from BERT and matches words in candidate and reference
sentences by cosine similarity.
It has been shown to correlate with human judgment on sentence-level and system-level evaluation.
Moreover, BERTScore computes precision, recall, and F1 measure, which can be useful for evaluating different language
generation tasks.
See the project's README at https://github.com/Tiiiger/bert_score#readme for more information.
"""
_KWARGS_DESCRIPTION = """
BERTScore Metrics with the hashcode from a source against one or more references.
Args:
predictions (list of str): Prediction/candidate sentences.
references (list of str or list of list of str): Reference sentences.
lang (str): Language of the sentences; required (e.g. 'en').
model_type (str): Bert specification, default using the suggested
model for the target language; has to specify at least one of
`model_type` or `lang`.
num_layers (int): The layer of representation to use,
default using the number of layers tuned on WMT16 correlation data.
verbose (bool): Turn on intermediate status update.
idf (bool or dict): Use idf weighting; can also be a precomputed idf_dict.
device (str): On which the contextual embedding model will be allocated on.
If this argument is None, the model lives on cuda:0 if cuda is available.
nthreads (int): Number of threads.
batch_size (int): Bert score processing batch size,
at least one of `model_type` or `lang`. `lang` needs to be
specified when `rescale_with_baseline` is True.
rescale_with_baseline (bool): Rescale bertscore with pre-computed baseline.
baseline_path (str): Customized baseline file.
use_fast_tokenizer (bool): `use_fast` parameter passed to HF tokenizer. New in version 0.3.10.
Returns:
precision: Precision.
recall: Recall.
f1: F1 score.
hashcode: Hashcode of the library.
Examples:
>>> predictions = ["hello there", "general kenobi"]
>>> references = ["hello there", "general kenobi"]
>>> bertscore = datasets.load_metric("bertscore")
>>> results = bertscore.compute(predictions=predictions, references=references, lang="en")
>>> print([round(v, 2) for v in results["f1"]])
[1.0, 1.0]
"""
@datasets.utils.file_utils.add_start_docstrings(_DESCRIPTION, _KWARGS_DESCRIPTION)
class BERTScore(datasets.Metric):
def _info(self):
return datasets.MetricInfo(
description=_DESCRIPTION,
citation=_CITATION,
homepage="https://github.com/Tiiiger/bert_score",
inputs_description=_KWARGS_DESCRIPTION,
features=datasets.Features(
{
"predictions": datasets.Value("string", id="sequence"),
"references": datasets.Sequence(datasets.Value("string", id="sequence"), id="references"),
}
),
codebase_urls=["https://github.com/Tiiiger/bert_score"],
reference_urls=[
"https://github.com/Tiiiger/bert_score",
"https://arxiv.org/abs/1904.09675",
],
)
def _compute(
self,
predictions,
references,
lang=None,
model_type=None,
num_layers=None,
verbose=False,
idf=False,
device=None,
batch_size=64,
nthreads=4,
all_layers=False,
rescale_with_baseline=False,
baseline_path=None,
use_fast_tokenizer=False,
):
get_hash = bert_score.utils.get_hash
scorer = bert_score.BERTScorer
if version.parse(bert_score.__version__) >= version.parse("0.3.10"):
get_hash = functools.partial(get_hash, use_fast_tokenizer=use_fast_tokenizer)
scorer = functools.partial(scorer, use_fast_tokenizer=use_fast_tokenizer)
elif use_fast_tokenizer:
raise ImportWarning(
"To use a fast tokenizer, the module `bert-score>=0.3.10` is required, and the current version of `bert-score` doesn't match this condition.\n"
'You can install it with `pip install "bert-score>=0.3.10"`.'
)
if model_type is None:
assert lang is not None, "either lang or model_type should be specified"
model_type = bert_score.utils.lang2model[lang.lower()]
if num_layers is None:
num_layers = bert_score.utils.model2layers[model_type]
hashcode = get_hash(
model=model_type,
num_layers=num_layers,
idf=idf,
rescale_with_baseline=rescale_with_baseline,
use_custom_baseline=baseline_path is not None,
)
with filter_logging_context():
if not hasattr(self, "cached_bertscorer") or self.cached_bertscorer.hash != hashcode:
self.cached_bertscorer = scorer(
model_type=model_type,
num_layers=num_layers,
batch_size=batch_size,
nthreads=nthreads,
all_layers=all_layers,
idf=idf,
device=device,
lang=lang,
rescale_with_baseline=rescale_with_baseline,
baseline_path=baseline_path,
)
(P, R, F) = self.cached_bertscorer.score(
cands=predictions,
refs=references,
verbose=verbose,
batch_size=batch_size,
)
output_dict = {
"precision": P.tolist(),
"recall": R.tolist(),
"f1": F.tolist(),
"hashcode": hashcode,
}
return output_dict
def add_batch(self, predictions=None, references=None, **kwargs):
"""Add a batch of predictions and references for the metric's stack."""
# References can be strings or lists of strings
# Let's change strings to lists of strings with one element
if references is not None:
references = [[ref] if isinstance(ref, str) else ref for ref in references]
super().add_batch(predictions=predictions, references=references, **kwargs)
def add(self, prediction=None, reference=None, **kwargs):
"""Add one prediction and reference for the metric's stack."""
# References can be strings or lists of strings
# Let's change strings to lists of strings with one element
if isinstance(reference, str):
reference = [reference]
super().add(prediction=prediction, reference=reference, **kwargs)
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/sacrebleu/README.md
|
# Metric Card for SacreBLEU
## Metric Description
SacreBLEU provides hassle-free computation of shareable, comparable, and reproducible BLEU scores. Inspired by Rico Sennrich's `multi-bleu-detok.perl`, it produces the official Workshop on Machine Translation (WMT) scores but works with plain text. It also knows all the standard test sets and handles downloading, processing, and tokenization.
See the [README.md] file at https://github.com/mjpost/sacreBLEU for more information.
## How to Use
This metric takes a set of predictions and a set of references as input, along with various optional parameters.
```python
>>> predictions = ["hello there general kenobi", "foo bar foobar"]
>>> references = [["hello there general kenobi", "hello there !"],
... ["foo bar foobar", "foo bar foobar"]]
>>> sacrebleu = datasets.load_metric("sacrebleu")
>>> results = sacrebleu.compute(predictions=predictions,
... references=references)
>>> print(list(results.keys()))
['score', 'counts', 'totals', 'precisions', 'bp', 'sys_len', 'ref_len']
>>> print(round(results["score"], 1))
100.0
```
### Inputs
- **`predictions`** (`list` of `str`): list of translations to score. Each translation should be tokenized into a list of tokens.
- **`references`** (`list` of `list` of `str`): A list of lists of references. The contents of the first sub-list are the references for the first prediction, the contents of the second sub-list are for the second prediction, etc. Note that there must be the same number of references for each prediction (i.e. all sub-lists must be of the same length).
- **`smooth_method`** (`str`): The smoothing method to use, defaults to `'exp'`. Possible values are:
- `'none'`: no smoothing
- `'floor'`: increment zero counts
- `'add-k'`: increment num/denom by k for n>1
- `'exp'`: exponential decay
- **`smooth_value`** (`float`): The smoothing value. Only valid when `smooth_method='floor'` (in which case `smooth_value` defaults to `0.1`) or `smooth_method='add-k'` (in which case `smooth_value` defaults to `1`).
- **`tokenize`** (`str`): Tokenization method to use for BLEU. If not provided, defaults to `'zh'` for Chinese, `'ja-mecab'` for Japanese and `'13a'` (mteval) otherwise. Possible values are:
- `'none'`: No tokenization.
- `'zh'`: Chinese tokenization.
- `'13a'`: mimics the `mteval-v13a` script from Moses.
- `'intl'`: International tokenization, mimics the `mteval-v14` script from Moses
- `'char'`: Language-agnostic character-level tokenization.
- `'ja-mecab'`: Japanese tokenization. Uses the [MeCab tokenizer](https://pypi.org/project/mecab-python3).
- **`lowercase`** (`bool`): If `True`, lowercases the input, enabling case-insensitivity. Defaults to `False`.
- **`force`** (`bool`): If `True`, insists that your tokenized input is actually detokenized. Defaults to `False`.
- **`use_effective_order`** (`bool`): If `True`, stops including n-gram orders for which precision is 0. This should be `True`, if sentence-level BLEU will be computed. Defaults to `False`.
### Output Values
- `score`: BLEU score
- `counts`: Counts
- `totals`: Totals
- `precisions`: Precisions
- `bp`: Brevity penalty
- `sys_len`: predictions length
- `ref_len`: reference length
The output is in the following format:
```python
{'score': 39.76353643835252, 'counts': [6, 4, 2, 1], 'totals': [10, 8, 6, 4], 'precisions': [60.0, 50.0, 33.333333333333336, 25.0], 'bp': 1.0, 'sys_len': 10, 'ref_len': 7}
```
The score can take any value between `0.0` and `100.0`, inclusive.
#### Values from Popular Papers
### Examples
```python
>>> predictions = ["hello there general kenobi",
... "on our way to ankh morpork"]
>>> references = [["hello there general kenobi", "hello there !"],
... ["goodbye ankh morpork", "ankh morpork"]]
>>> sacrebleu = datasets.load_metric("sacrebleu")
>>> results = sacrebleu.compute(predictions=predictions,
... references=references)
>>> print(list(results.keys()))
['score', 'counts', 'totals', 'precisions', 'bp', 'sys_len', 'ref_len']
>>> print(round(results["score"], 1))
39.8
```
## Limitations and Bias
Because what this metric calculates is BLEU scores, it has the same limitations as that metric, except that sacreBLEU is more easily reproducible.
## Citation
```bibtex
@inproceedings{post-2018-call,
title = "A Call for Clarity in Reporting {BLEU} Scores",
author = "Post, Matt",
booktitle = "Proceedings of the Third Conference on Machine Translation: Research Papers",
month = oct,
year = "2018",
address = "Belgium, Brussels",
publisher = "Association for Computational Linguistics",
url = "https://www.aclweb.org/anthology/W18-6319",
pages = "186--191",
}
```
## Further References
- See the [sacreBLEU README.md file](https://github.com/mjpost/sacreBLEU) for more information.
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/sacrebleu/sacrebleu.py
|
# Copyright 2020 The HuggingFace Datasets Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
""" SACREBLEU metric. """
import sacrebleu as scb
from packaging import version
import datasets
_CITATION = """\
@inproceedings{post-2018-call,
title = "A Call for Clarity in Reporting {BLEU} Scores",
author = "Post, Matt",
booktitle = "Proceedings of the Third Conference on Machine Translation: Research Papers",
month = oct,
year = "2018",
address = "Belgium, Brussels",
publisher = "Association for Computational Linguistics",
url = "https://www.aclweb.org/anthology/W18-6319",
pages = "186--191",
}
"""
_DESCRIPTION = """\
SacreBLEU provides hassle-free computation of shareable, comparable, and reproducible BLEU scores.
Inspired by Rico Sennrich's `multi-bleu-detok.perl`, it produces the official WMT scores but works with plain text.
It also knows all the standard test sets and handles downloading, processing, and tokenization for you.
See the [README.md] file at https://github.com/mjpost/sacreBLEU for more information.
"""
_KWARGS_DESCRIPTION = """
Produces BLEU scores along with its sufficient statistics
from a source against one or more references.
Args:
predictions (`list` of `str`): list of translations to score. Each translation should be tokenized into a list of tokens.
references (`list` of `list` of `str`): A list of lists of references. The contents of the first sub-list are the references for the first prediction, the contents of the second sub-list are for the second prediction, etc. Note that there must be the same number of references for each prediction (i.e. all sub-lists must be of the same length).
smooth_method (`str`): The smoothing method to use, defaults to `'exp'`. Possible values are:
- `'none'`: no smoothing
- `'floor'`: increment zero counts
- `'add-k'`: increment num/denom by k for n>1
- `'exp'`: exponential decay
smooth_value (`float`): The smoothing value. Only valid when `smooth_method='floor'` (in which case `smooth_value` defaults to `0.1`) or `smooth_method='add-k'` (in which case `smooth_value` defaults to `1`).
tokenize (`str`): Tokenization method to use for BLEU. If not provided, defaults to `'zh'` for Chinese, `'ja-mecab'` for Japanese and `'13a'` (mteval) otherwise. Possible values are:
- `'none'`: No tokenization.
- `'zh'`: Chinese tokenization.
- `'13a'`: mimics the `mteval-v13a` script from Moses.
- `'intl'`: International tokenization, mimics the `mteval-v14` script from Moses
- `'char'`: Language-agnostic character-level tokenization.
- `'ja-mecab'`: Japanese tokenization. Uses the [MeCab tokenizer](https://pypi.org/project/mecab-python3).
lowercase (`bool`): If `True`, lowercases the input, enabling case-insensitivity. Defaults to `False`.
force (`bool`): If `True`, insists that your tokenized input is actually detokenized. Defaults to `False`.
use_effective_order (`bool`): If `True`, stops including n-gram orders for which precision is 0. This should be `True`, if sentence-level BLEU will be computed. Defaults to `False`.
Returns:
'score': BLEU score,
'counts': Counts,
'totals': Totals,
'precisions': Precisions,
'bp': Brevity penalty,
'sys_len': predictions length,
'ref_len': reference length,
Examples:
Example 1:
>>> predictions = ["hello there general kenobi", "foo bar foobar"]
>>> references = [["hello there general kenobi", "hello there !"], ["foo bar foobar", "foo bar foobar"]]
>>> sacrebleu = datasets.load_metric("sacrebleu")
>>> results = sacrebleu.compute(predictions=predictions, references=references)
>>> print(list(results.keys()))
['score', 'counts', 'totals', 'precisions', 'bp', 'sys_len', 'ref_len']
>>> print(round(results["score"], 1))
100.0
Example 2:
>>> predictions = ["hello there general kenobi",
... "on our way to ankh morpork"]
>>> references = [["hello there general kenobi", "hello there !"],
... ["goodbye ankh morpork", "ankh morpork"]]
>>> sacrebleu = datasets.load_metric("sacrebleu")
>>> results = sacrebleu.compute(predictions=predictions,
... references=references)
>>> print(list(results.keys()))
['score', 'counts', 'totals', 'precisions', 'bp', 'sys_len', 'ref_len']
>>> print(round(results["score"], 1))
39.8
"""
@datasets.utils.file_utils.add_start_docstrings(_DESCRIPTION, _KWARGS_DESCRIPTION)
class Sacrebleu(datasets.Metric):
def _info(self):
if version.parse(scb.__version__) < version.parse("1.4.12"):
raise ImportWarning(
"To use `sacrebleu`, the module `sacrebleu>=1.4.12` is required, and the current version of `sacrebleu` doesn't match this condition.\n"
'You can install it with `pip install "sacrebleu>=1.4.12"`.'
)
return datasets.MetricInfo(
description=_DESCRIPTION,
citation=_CITATION,
homepage="https://github.com/mjpost/sacreBLEU",
inputs_description=_KWARGS_DESCRIPTION,
features=datasets.Features(
{
"predictions": datasets.Value("string", id="sequence"),
"references": datasets.Sequence(datasets.Value("string", id="sequence"), id="references"),
}
),
codebase_urls=["https://github.com/mjpost/sacreBLEU"],
reference_urls=[
"https://github.com/mjpost/sacreBLEU",
"https://en.wikipedia.org/wiki/BLEU",
"https://towardsdatascience.com/evaluating-text-output-in-nlp-bleu-at-your-own-risk-e8609665a213",
],
)
def _compute(
self,
predictions,
references,
smooth_method="exp",
smooth_value=None,
force=False,
lowercase=False,
tokenize=None,
use_effective_order=False,
):
references_per_prediction = len(references[0])
if any(len(refs) != references_per_prediction for refs in references):
raise ValueError("Sacrebleu requires the same number of references for each prediction")
transformed_references = [[refs[i] for refs in references] for i in range(references_per_prediction)]
output = scb.corpus_bleu(
predictions,
transformed_references,
smooth_method=smooth_method,
smooth_value=smooth_value,
force=force,
lowercase=lowercase,
use_effective_order=use_effective_order,
**({"tokenize": tokenize} if tokenize else {}),
)
output_dict = {
"score": output.score,
"counts": output.counts,
"totals": output.totals,
"precisions": output.precisions,
"bp": output.bp,
"sys_len": output.sys_len,
"ref_len": output.ref_len,
}
return output_dict
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/spearmanr/spearmanr.py
|
# Copyright 2021 The HuggingFace Datasets Authors and the current dataset script contributor.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""Spearman correlation coefficient metric."""
from scipy.stats import spearmanr
import datasets
_DESCRIPTION = """
The Spearman rank-order correlation coefficient is a measure of the
relationship between two datasets. Like other correlation coefficients,
this one varies between -1 and +1 with 0 implying no correlation.
Positive correlations imply that as data in dataset x increases, so
does data in dataset y. Negative correlations imply that as x increases,
y decreases. Correlations of -1 or +1 imply an exact monotonic relationship.
Unlike the Pearson correlation, the Spearman correlation does not
assume that both datasets are normally distributed.
The p-value roughly indicates the probability of an uncorrelated system
producing datasets that have a Spearman correlation at least as extreme
as the one computed from these datasets. The p-values are not entirely
reliable but are probably reasonable for datasets larger than 500 or so.
"""
_KWARGS_DESCRIPTION = """
Args:
predictions (`List[float]`): Predicted labels, as returned by a model.
references (`List[float]`): Ground truth labels.
return_pvalue (`bool`): If `True`, returns the p-value. If `False`, returns
only the spearmanr score. Defaults to `False`.
Returns:
spearmanr (`float`): Spearman correlation coefficient.
p-value (`float`): p-value. **Note**: is only returned if `return_pvalue=True` is input.
Examples:
Example 1:
>>> spearmanr_metric = datasets.load_metric("spearmanr")
>>> results = spearmanr_metric.compute(references=[1, 2, 3, 4, 5], predictions=[10, 9, 2.5, 6, 4])
>>> print(results)
{'spearmanr': -0.7}
Example 2:
>>> spearmanr_metric = datasets.load_metric("spearmanr")
>>> results = spearmanr_metric.compute(references=[1, 2, 3, 4, 5],
... predictions=[10, 9, 2.5, 6, 4],
... return_pvalue=True)
>>> print(results['spearmanr'])
-0.7
>>> print(round(results['spearmanr_pvalue'], 2))
0.19
"""
_CITATION = r"""\
@book{kokoska2000crc,
title={CRC standard probability and statistics tables and formulae},
author={Kokoska, Stephen and Zwillinger, Daniel},
year={2000},
publisher={Crc Press}
}
@article{2020SciPy-NMeth,
author = {Virtanen, Pauli and Gommers, Ralf and Oliphant, Travis E. and
Haberland, Matt and Reddy, Tyler and Cournapeau, David and
Burovski, Evgeni and Peterson, Pearu and Weckesser, Warren and
Bright, Jonathan and {van der Walt}, St{\'e}fan J. and
Brett, Matthew and Wilson, Joshua and Millman, K. Jarrod and
Mayorov, Nikolay and Nelson, Andrew R. J. and Jones, Eric and
Kern, Robert and Larson, Eric and Carey, C J and
Polat, {\.I}lhan and Feng, Yu and Moore, Eric W. and
{VanderPlas}, Jake and Laxalde, Denis and Perktold, Josef and
Cimrman, Robert and Henriksen, Ian and Quintero, E. A. and
Harris, Charles R. and Archibald, Anne M. and
Ribeiro, Ant{\^o}nio H. and Pedregosa, Fabian and
{van Mulbregt}, Paul and {SciPy 1.0 Contributors}},
title = {{{SciPy} 1.0: Fundamental Algorithms for Scientific
Computing in Python}},
journal = {Nature Methods},
year = {2020},
volume = {17},
pages = {261--272},
adsurl = {https://rdcu.be/b08Wh},
doi = {10.1038/s41592-019-0686-2},
}
"""
@datasets.utils.file_utils.add_start_docstrings(_DESCRIPTION, _KWARGS_DESCRIPTION)
class Spearmanr(datasets.Metric):
def _info(self):
return datasets.MetricInfo(
description=_DESCRIPTION,
citation=_CITATION,
inputs_description=_KWARGS_DESCRIPTION,
features=datasets.Features(
{
"predictions": datasets.Value("float"),
"references": datasets.Value("float"),
}
),
reference_urls=["https://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.spearmanr.html"],
)
def _compute(self, predictions, references, return_pvalue=False):
results = spearmanr(references, predictions)
if return_pvalue:
return {"spearmanr": results[0], "spearmanr_pvalue": results[1]}
else:
return {"spearmanr": results[0]}
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/spearmanr/README.md
|
# Metric Card for Spearman Correlation Coefficient Metric (spearmanr)
## Metric Description
The Spearman rank-order correlation coefficient is a measure of the
relationship between two datasets. Like other correlation coefficients,
this one varies between -1 and +1 with 0 implying no correlation.
Positive correlations imply that as data in dataset x increases, so
does data in dataset y. Negative correlations imply that as x increases,
y decreases. Correlations of -1 or +1 imply an exact monotonic relationship.
Unlike the Pearson correlation, the Spearman correlation does not
assume that both datasets are normally distributed.
The p-value roughly indicates the probability of an uncorrelated system
producing datasets that have a Spearman correlation at least as extreme
as the one computed from these datasets. The p-values are not entirely
reliable but are probably reasonable for datasets larger than 500 or so.
## How to Use
At minimum, this metric only requires a `list` of predictions and a `list` of references:
```python
>>> spearmanr_metric = datasets.load_metric("spearmanr")
>>> results = spearmanr_metric.compute(references=[1, 2, 3, 4, 5], predictions=[10, 9, 2.5, 6, 4])
>>> print(results)
{'spearmanr': -0.7}
```
### Inputs
- **`predictions`** (`list` of `float`): Predicted labels, as returned by a model.
- **`references`** (`list` of `float`): Ground truth labels.
- **`return_pvalue`** (`bool`): If `True`, returns the p-value. If `False`, returns
only the spearmanr score. Defaults to `False`.
### Output Values
- **`spearmanr`** (`float`): Spearman correlation coefficient.
- **`p-value`** (`float`): p-value. **Note**: is only returned
if `return_pvalue=True` is input.
If `return_pvalue=False`, the output is a `dict` with one value, as below:
```python
{'spearmanr': -0.7}
```
Otherwise, if `return_pvalue=True`, the output is a `dict` containing a the `spearmanr` value as well as the corresponding `pvalue`:
```python
{'spearmanr': -0.7, 'spearmanr_pvalue': 0.1881204043741873}
```
Spearman rank-order correlations can take on any value from `-1` to `1`, inclusive.
The p-values can take on any value from `0` to `1`, inclusive.
#### Values from Popular Papers
### Examples
A basic example:
```python
>>> spearmanr_metric = datasets.load_metric("spearmanr")
>>> results = spearmanr_metric.compute(references=[1, 2, 3, 4, 5], predictions=[10, 9, 2.5, 6, 4])
>>> print(results)
{'spearmanr': -0.7}
```
The same example, but that also returns the pvalue:
```python
>>> spearmanr_metric = datasets.load_metric("spearmanr")
>>> results = spearmanr_metric.compute(references=[1, 2, 3, 4, 5], predictions=[10, 9, 2.5, 6, 4], return_pvalue=True)
>>> print(results)
{'spearmanr': -0.7, 'spearmanr_pvalue': 0.1881204043741873
>>> print(results['spearmanr'])
-0.7
>>> print(results['spearmanr_pvalue'])
0.1881204043741873
```
## Limitations and Bias
## Citation
```bibtex
@book{kokoska2000crc,
title={CRC standard probability and statistics tables and formulae},
author={Kokoska, Stephen and Zwillinger, Daniel},
year={2000},
publisher={Crc Press}
}
@article{2020SciPy-NMeth,
author = {Virtanen, Pauli and Gommers, Ralf and Oliphant, Travis E. and
Haberland, Matt and Reddy, Tyler and Cournapeau, David and
Burovski, Evgeni and Peterson, Pearu and Weckesser, Warren and
Bright, Jonathan and {van der Walt}, St{\'e}fan J. and
Brett, Matthew and Wilson, Joshua and Millman, K. Jarrod and
Mayorov, Nikolay and Nelson, Andrew R. J. and Jones, Eric and
Kern, Robert and Larson, Eric and Carey, C J and
Polat, {\.I}lhan and Feng, Yu and Moore, Eric W. and
{VanderPlas}, Jake and Laxalde, Denis and Perktold, Josef and
Cimrman, Robert and Henriksen, Ian and Quintero, E. A. and
Harris, Charles R. and Archibald, Anne M. and
Ribeiro, Ant{\^o}nio H. and Pedregosa, Fabian and
{van Mulbregt}, Paul and {SciPy 1.0 Contributors}},
title = {{{SciPy} 1.0: Fundamental Algorithms for Scientific
Computing in Python}},
journal = {Nature Methods},
year = {2020},
volume = {17},
pages = {261--272},
adsurl = {https://rdcu.be/b08Wh},
doi = {10.1038/s41592-019-0686-2},
}
```
## Further References
*Add any useful further references.*
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/squad/README.md
|
# Metric Card for SQuAD
## Metric description
This metric wraps the official scoring script for version 1 of the [Stanford Question Answering Dataset (SQuAD)](https://huggingface.co/datasets/squad).
SQuAD is a reading comprehension dataset, consisting of questions posed by crowdworkers on a set of Wikipedia articles, where the answer to every question is a segment of text, or span, from the corresponding reading passage, or the question might be unanswerable.
## How to use
The metric takes two files or two lists of question-answers dictionaries as inputs : one with the predictions of the model and the other with the references to be compared to:
```python
from datasets import load_metric
squad_metric = load_metric("squad")
results = squad_metric.compute(predictions=predictions, references=references)
```
## Output values
This metric outputs a dictionary with two values: the average exact match score and the average [F1 score](https://huggingface.co/metrics/f1).
```
{'exact_match': 100.0, 'f1': 100.0}
```
The range of `exact_match` is 0-100, where 0.0 means no answers were matched and 100.0 means all answers were matched.
The range of `f1` is 0-1 -- its lowest possible value is 0, if either the precision or the recall is 0, and its highest possible value is 1.0, which means perfect precision and recall.
### Values from popular papers
The [original SQuAD paper](https://nlp.stanford.edu/pubs/rajpurkar2016squad.pdf) reported an F1 score of 51.0% and an Exact Match score of 40.0%. They also report that human performance on the dataset represents an F1 score of 90.5% and an Exact Match score of 80.3%.
For more recent model performance, see the [dataset leaderboard](https://paperswithcode.com/dataset/squad).
## Examples
Maximal values for both exact match and F1 (perfect match):
```python
from datasets import load_metric
squad_metric = load_metric("squad")
predictions = [{'prediction_text': '1976', 'id': '56e10a3be3433e1400422b22'}]
references = [{'answers': {'answer_start': [97], 'text': ['1976']}, 'id': '56e10a3be3433e1400422b22'}]
results = squad_metric.compute(predictions=predictions, references=references)
results
{'exact_match': 100.0, 'f1': 100.0}
```
Minimal values for both exact match and F1 (no match):
```python
from datasets import load_metric
squad_metric = load_metric("squad")
predictions = [{'prediction_text': '1999', 'id': '56e10a3be3433e1400422b22'}]
references = [{'answers': {'answer_start': [97], 'text': ['1976']}, 'id': '56e10a3be3433e1400422b22'}]
results = squad_metric.compute(predictions=predictions, references=references)
results
{'exact_match': 0.0, 'f1': 0.0}
```
Partial match (2 out of 3 answers correct) :
```python
from datasets import load_metric
squad_metric = load_metric("squad")
predictions = [{'prediction_text': '1976', 'id': '56e10a3be3433e1400422b22'}, {'prediction_text': 'Beyonce', 'id': '56d2051ce7d4791d0090260b'}, {'prediction_text': 'climate change', 'id': '5733b5344776f419006610e1'}]
references = [{'answers': {'answer_start': [97], 'text': ['1976']}, 'id': '56e10a3be3433e1400422b22'}, {'answers': {'answer_start': [233], 'text': ['Beyoncé and Bruno Mars']}, 'id': '56d2051ce7d4791d0090260b'}, {'answers': {'answer_start': [891], 'text': ['climate change']}, 'id': '5733b5344776f419006610e1'}]
results = squad_metric.compute(predictions=predictions, references=references)
results
{'exact_match': 66.66666666666667, 'f1': 66.66666666666667}
```
## Limitations and bias
This metric works only with datasets that have the same format as [SQuAD v.1 dataset](https://huggingface.co/datasets/squad).
The SQuAD dataset does contain a certain amount of noise, such as duplicate questions as well as missing answers, but these represent a minority of the 100,000 question-answer pairs. Also, neither exact match nor F1 score reflect whether models do better on certain types of questions (e.g. who questions) or those that cover a certain gender or geographical area -- carrying out more in-depth error analysis can complement these numbers.
## Citation
@inproceedings{Rajpurkar2016SQuAD10,
title={SQuAD: 100, 000+ Questions for Machine Comprehension of Text},
author={Pranav Rajpurkar and Jian Zhang and Konstantin Lopyrev and Percy Liang},
booktitle={EMNLP},
year={2016}
}
## Further References
- [The Stanford Question Answering Dataset: Background, Challenges, Progress (blog post)](https://rajpurkar.github.io/mlx/qa-and-squad/)
- [Hugging Face Course -- Question Answering](https://huggingface.co/course/chapter7/7)
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/squad/squad.py
|
# Copyright 2020 The HuggingFace Datasets Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
""" SQuAD metric. """
import datasets
from .evaluate import evaluate
_CITATION = """\
@inproceedings{Rajpurkar2016SQuAD10,
title={SQuAD: 100, 000+ Questions for Machine Comprehension of Text},
author={Pranav Rajpurkar and Jian Zhang and Konstantin Lopyrev and Percy Liang},
booktitle={EMNLP},
year={2016}
}
"""
_DESCRIPTION = """
This metric wrap the official scoring script for version 1 of the Stanford Question Answering Dataset (SQuAD).
Stanford Question Answering Dataset (SQuAD) is a reading comprehension dataset, consisting of questions posed by
crowdworkers on a set of Wikipedia articles, where the answer to every question is a segment of text, or span,
from the corresponding reading passage, or the question might be unanswerable.
"""
_KWARGS_DESCRIPTION = """
Computes SQuAD scores (F1 and EM).
Args:
predictions: List of question-answers dictionaries with the following key-values:
- 'id': id of the question-answer pair as given in the references (see below)
- 'prediction_text': the text of the answer
references: List of question-answers dictionaries with the following key-values:
- 'id': id of the question-answer pair (see above),
- 'answers': a Dict in the SQuAD dataset format
{
'text': list of possible texts for the answer, as a list of strings
'answer_start': list of start positions for the answer, as a list of ints
}
Note that answer_start values are not taken into account to compute the metric.
Returns:
'exact_match': Exact match (the normalized answer exactly match the gold answer)
'f1': The F-score of predicted tokens versus the gold answer
Examples:
>>> predictions = [{'prediction_text': '1976', 'id': '56e10a3be3433e1400422b22'}]
>>> references = [{'answers': {'answer_start': [97], 'text': ['1976']}, 'id': '56e10a3be3433e1400422b22'}]
>>> squad_metric = datasets.load_metric("squad")
>>> results = squad_metric.compute(predictions=predictions, references=references)
>>> print(results)
{'exact_match': 100.0, 'f1': 100.0}
"""
@datasets.utils.file_utils.add_start_docstrings(_DESCRIPTION, _KWARGS_DESCRIPTION)
class Squad(datasets.Metric):
def _info(self):
return datasets.MetricInfo(
description=_DESCRIPTION,
citation=_CITATION,
inputs_description=_KWARGS_DESCRIPTION,
features=datasets.Features(
{
"predictions": {"id": datasets.Value("string"), "prediction_text": datasets.Value("string")},
"references": {
"id": datasets.Value("string"),
"answers": datasets.features.Sequence(
{
"text": datasets.Value("string"),
"answer_start": datasets.Value("int32"),
}
),
},
}
),
codebase_urls=["https://rajpurkar.github.io/SQuAD-explorer/"],
reference_urls=["https://rajpurkar.github.io/SQuAD-explorer/"],
)
def _compute(self, predictions, references):
pred_dict = {prediction["id"]: prediction["prediction_text"] for prediction in predictions}
dataset = [
{
"paragraphs": [
{
"qas": [
{
"answers": [{"text": answer_text} for answer_text in ref["answers"]["text"]],
"id": ref["id"],
}
for ref in references
]
}
]
}
]
score = evaluate(dataset=dataset, predictions=pred_dict)
return score
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/squad/evaluate.py
|
""" Official evaluation script for v1.1 of the SQuAD dataset. """
import argparse
import json
import re
import string
import sys
from collections import Counter
def normalize_answer(s):
"""Lower text and remove punctuation, articles and extra whitespace."""
def remove_articles(text):
return re.sub(r"\b(a|an|the)\b", " ", text)
def white_space_fix(text):
return " ".join(text.split())
def remove_punc(text):
exclude = set(string.punctuation)
return "".join(ch for ch in text if ch not in exclude)
def lower(text):
return text.lower()
return white_space_fix(remove_articles(remove_punc(lower(s))))
def f1_score(prediction, ground_truth):
prediction_tokens = normalize_answer(prediction).split()
ground_truth_tokens = normalize_answer(ground_truth).split()
common = Counter(prediction_tokens) & Counter(ground_truth_tokens)
num_same = sum(common.values())
if num_same == 0:
return 0
precision = 1.0 * num_same / len(prediction_tokens)
recall = 1.0 * num_same / len(ground_truth_tokens)
f1 = (2 * precision * recall) / (precision + recall)
return f1
def exact_match_score(prediction, ground_truth):
return normalize_answer(prediction) == normalize_answer(ground_truth)
def metric_max_over_ground_truths(metric_fn, prediction, ground_truths):
scores_for_ground_truths = []
for ground_truth in ground_truths:
score = metric_fn(prediction, ground_truth)
scores_for_ground_truths.append(score)
return max(scores_for_ground_truths)
def evaluate(dataset, predictions):
f1 = exact_match = total = 0
for article in dataset:
for paragraph in article["paragraphs"]:
for qa in paragraph["qas"]:
total += 1
if qa["id"] not in predictions:
message = "Unanswered question " + qa["id"] + " will receive score 0."
print(message, file=sys.stderr)
continue
ground_truths = [x["text"] for x in qa["answers"]]
prediction = predictions[qa["id"]]
exact_match += metric_max_over_ground_truths(exact_match_score, prediction, ground_truths)
f1 += metric_max_over_ground_truths(f1_score, prediction, ground_truths)
exact_match = 100.0 * exact_match / total
f1 = 100.0 * f1 / total
return {"exact_match": exact_match, "f1": f1}
if __name__ == "__main__":
expected_version = "1.1"
parser = argparse.ArgumentParser(description="Evaluation for SQuAD " + expected_version)
parser.add_argument("dataset_file", help="Dataset file")
parser.add_argument("prediction_file", help="Prediction File")
args = parser.parse_args()
with open(args.dataset_file) as dataset_file:
dataset_json = json.load(dataset_file)
if dataset_json["version"] != expected_version:
print(
"Evaluation expects v-" + expected_version + ", but got dataset with v-" + dataset_json["version"],
file=sys.stderr,
)
dataset = dataset_json["data"]
with open(args.prediction_file) as prediction_file:
predictions = json.load(prediction_file)
print(json.dumps(evaluate(dataset, predictions)))
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/squad_v2/README.md
|
# Metric Card for SQuAD v2
## Metric description
This metric wraps the official scoring script for version 2 of the [Stanford Question Answering Dataset (SQuAD)](https://huggingface.co/datasets/squad_v2).
SQuAD is a reading comprehension dataset, consisting of questions posed by crowdworkers on a set of Wikipedia articles, where the answer to every question is a segment of text, or span, from the corresponding reading passage, or the question might be unanswerable.
SQuAD 2.0 combines the 100,000 questions in SQuAD 1.1 with over 50,000 unanswerable questions written adversarially by crowdworkers to look similar to answerable ones. To do well on SQuAD2.0, systems must not only answer questions when possible, but also determine when no answer is supported by the paragraph and abstain from answering.
## How to use
The metric takes two files or two lists - one representing model predictions and the other the references to compare them to.
*Predictions* : List of triple for question-answers to score with the following key-value pairs:
* `'id'`: the question-answer identification field of the question and answer pair
* `'prediction_text'` : the text of the answer
* `'no_answer_probability'` : the probability that the question has no answer
*References*: List of question-answers dictionaries with the following key-value pairs:
* `'id'`: id of the question-answer pair (see above),
* `'answers'`: a list of Dict {'text': text of the answer as a string}
* `'no_answer_threshold'`: the probability threshold to decide that a question has no answer.
```python
from datasets import load_metric
squad_metric = load_metric("squad_v2")
results = squad_metric.compute(predictions=predictions, references=references)
```
## Output values
This metric outputs a dictionary with 13 values:
* `'exact'`: Exact match (the normalized answer exactly match the gold answer) (see the `exact_match` metric (forthcoming))
* `'f1'`: The average F1-score of predicted tokens versus the gold answer (see the [F1 score](https://huggingface.co/metrics/f1) metric)
* `'total'`: Number of scores considered
* `'HasAns_exact'`: Exact match (the normalized answer exactly match the gold answer)
* `'HasAns_f1'`: The F-score of predicted tokens versus the gold answer
* `'HasAns_total'`: How many of the questions have answers
* `'NoAns_exact'`: Exact match (the normalized answer exactly match the gold answer)
* `'NoAns_f1'`: The F-score of predicted tokens versus the gold answer
* `'NoAns_total'`: How many of the questions have no answers
* `'best_exact'` : Best exact match (with varying threshold)
* `'best_exact_thresh'`: No-answer probability threshold associated to the best exact match
* `'best_f1'`: Best F1 score (with varying threshold)
* `'best_f1_thresh'`: No-answer probability threshold associated to the best F1
The range of `exact_match` is 0-100, where 0.0 means no answers were matched and 100.0 means all answers were matched.
The range of `f1` is 0-1 -- its lowest possible value is 0, if either the precision or the recall is 0, and its highest possible value is 1.0, which means perfect precision and recall.
The range of `total` depends on the length of predictions/references: its minimal value is 0, and maximal value is the total number of questions in the predictions and references.
### Values from popular papers
The [SQuAD v2 paper](https://arxiv.org/pdf/1806.03822.pdf) reported an F1 score of 66.3% and an Exact Match score of 63.4%.
They also report that human performance on the dataset represents an F1 score of 89.5% and an Exact Match score of 86.9%.
For more recent model performance, see the [dataset leaderboard](https://paperswithcode.com/dataset/squad).
## Examples
Maximal values for both exact match and F1 (perfect match):
```python
from datasets import load_metric
squad_v2_ metric = load_metric("squad_v2")
predictions = [{'prediction_text': '1976', 'id': '56e10a3be3433e1400422b22', 'no_answer_probability': 0.}]
references = [{'answers': {'answer_start': [97], 'text': ['1976']}, 'id': '56e10a3be3433e1400422b22'}]
results = squad_v2_metric.compute(predictions=predictions, references=references)
results
{'exact': 100.0, 'f1': 100.0, 'total': 1, 'HasAns_exact': 100.0, 'HasAns_f1': 100.0, 'HasAns_total': 1, 'best_exact': 100.0, 'best_exact_thresh': 0.0, 'best_f1': 100.0, 'best_f1_thresh': 0.0}
```
Minimal values for both exact match and F1 (no match):
```python
from datasets import load_metric
squad_metric = load_metric("squad_v2")
predictions = [{'prediction_text': '1999', 'id': '56e10a3be3433e1400422b22', 'no_answer_probability': 0.}]
references = [{'answers': {'answer_start': [97], 'text': ['1976']}, 'id': '56e10a3be3433e1400422b22'}]
results = squad_v2_metric.compute(predictions=predictions, references=references)
results
{'exact': 0.0, 'f1': 0.0, 'total': 1, 'HasAns_exact': 0.0, 'HasAns_f1': 0.0, 'HasAns_total': 1, 'best_exact': 0.0, 'best_exact_thresh': 0.0, 'best_f1': 0.0, 'best_f1_thresh': 0.0}
```
Partial match (2 out of 3 answers correct) :
```python
from datasets import load_metric
squad_metric = load_metric("squad_v2")
predictions = [{'prediction_text': '1976', 'id': '56e10a3be3433e1400422b22', 'no_answer_probability': 0.}, {'prediction_text': 'Beyonce', 'id': '56d2051ce7d4791d0090260b', 'no_answer_probability': 0.}, {'prediction_text': 'climate change', 'id': '5733b5344776f419006610e1', 'no_answer_probability': 0.}]
references = [{'answers': {'answer_start': [97], 'text': ['1976']}, 'id': '56e10a3be3433e1400422b22'}, {'answers': {'answer_start': [233], 'text': ['Beyoncé and Bruno Mars']}, 'id': '56d2051ce7d4791d0090260b'}, {'answers': {'answer_start': [891], 'text': ['climate change']}, 'id': '5733b5344776f419006610e1'}]
results = squad_v2_metric.compute(predictions=predictions, references=references)
results
{'exact': 66.66666666666667, 'f1': 66.66666666666667, 'total': 3, 'HasAns_exact': 66.66666666666667, 'HasAns_f1': 66.66666666666667, 'HasAns_total': 3, 'best_exact': 66.66666666666667, 'best_exact_thresh': 0.0, 'best_f1': 66.66666666666667, 'best_f1_thresh': 0.0}
```
## Limitations and bias
This metric works only with the datasets in the same format as the [SQuAD v.2 dataset](https://huggingface.co/datasets/squad_v2).
The SQuAD datasets do contain a certain amount of noise, such as duplicate questions as well as missing answers, but these represent a minority of the 100,000 question-answer pairs. Also, neither exact match nor F1 score reflect whether models do better on certain types of questions (e.g. who questions) or those that cover a certain gender or geographical area -- carrying out more in-depth error analysis can complement these numbers.
## Citation
```bibtex
@inproceedings{Rajpurkar2018SQuAD2,
title={Know What You Don't Know: Unanswerable Questions for SQuAD},
author={Pranav Rajpurkar and Jian Zhang and Percy Liang},
booktitle={ACL 2018},
year={2018}
}
```
## Further References
- [The Stanford Question Answering Dataset: Background, Challenges, Progress (blog post)](https://rajpurkar.github.io/mlx/qa-and-squad/)
- [Hugging Face Course -- Question Answering](https://huggingface.co/course/chapter7/7)
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/squad_v2/evaluate.py
|
"""Official evaluation script for SQuAD version 2.0.
In addition to basic functionality, we also compute additional statistics and
plot precision-recall curves if an additional na_prob.json file is provided.
This file is expected to map question ID's to the model's predicted probability
that a question is unanswerable.
"""
import argparse
import collections
import json
import os
import re
import string
import sys
import numpy as np
ARTICLES_REGEX = re.compile(r"\b(a|an|the)\b", re.UNICODE)
OPTS = None
def parse_args():
parser = argparse.ArgumentParser("Official evaluation script for SQuAD version 2.0.")
parser.add_argument("data_file", metavar="data.json", help="Input data JSON file.")
parser.add_argument("pred_file", metavar="pred.json", help="Model predictions.")
parser.add_argument(
"--out-file", "-o", metavar="eval.json", help="Write accuracy metrics to file (default is stdout)."
)
parser.add_argument(
"--na-prob-file", "-n", metavar="na_prob.json", help="Model estimates of probability of no answer."
)
parser.add_argument(
"--na-prob-thresh",
"-t",
type=float,
default=1.0,
help='Predict "" if no-answer probability exceeds this (default = 1.0).',
)
parser.add_argument(
"--out-image-dir", "-p", metavar="out_images", default=None, help="Save precision-recall curves to directory."
)
parser.add_argument("--verbose", "-v", action="store_true")
if len(sys.argv) == 1:
parser.print_help()
sys.exit(1)
return parser.parse_args()
def make_qid_to_has_ans(dataset):
qid_to_has_ans = {}
for article in dataset:
for p in article["paragraphs"]:
for qa in p["qas"]:
qid_to_has_ans[qa["id"]] = bool(qa["answers"]["text"])
return qid_to_has_ans
def normalize_answer(s):
"""Lower text and remove punctuation, articles and extra whitespace."""
def remove_articles(text):
return ARTICLES_REGEX.sub(" ", text)
def white_space_fix(text):
return " ".join(text.split())
def remove_punc(text):
exclude = set(string.punctuation)
return "".join(ch for ch in text if ch not in exclude)
def lower(text):
return text.lower()
return white_space_fix(remove_articles(remove_punc(lower(s))))
def get_tokens(s):
if not s:
return []
return normalize_answer(s).split()
def compute_exact(a_gold, a_pred):
return int(normalize_answer(a_gold) == normalize_answer(a_pred))
def compute_f1(a_gold, a_pred):
gold_toks = get_tokens(a_gold)
pred_toks = get_tokens(a_pred)
common = collections.Counter(gold_toks) & collections.Counter(pred_toks)
num_same = sum(common.values())
if len(gold_toks) == 0 or len(pred_toks) == 0:
# If either is no-answer, then F1 is 1 if they agree, 0 otherwise
return int(gold_toks == pred_toks)
if num_same == 0:
return 0
precision = 1.0 * num_same / len(pred_toks)
recall = 1.0 * num_same / len(gold_toks)
f1 = (2 * precision * recall) / (precision + recall)
return f1
def get_raw_scores(dataset, preds):
exact_scores = {}
f1_scores = {}
for article in dataset:
for p in article["paragraphs"]:
for qa in p["qas"]:
qid = qa["id"]
gold_answers = [t for t in qa["answers"]["text"] if normalize_answer(t)]
if not gold_answers:
# For unanswerable questions, only correct answer is empty string
gold_answers = [""]
if qid not in preds:
print(f"Missing prediction for {qid}")
continue
a_pred = preds[qid]
# Take max over all gold answers
exact_scores[qid] = max(compute_exact(a, a_pred) for a in gold_answers)
f1_scores[qid] = max(compute_f1(a, a_pred) for a in gold_answers)
return exact_scores, f1_scores
def apply_no_ans_threshold(scores, na_probs, qid_to_has_ans, na_prob_thresh):
new_scores = {}
for qid, s in scores.items():
pred_na = na_probs[qid] > na_prob_thresh
if pred_na:
new_scores[qid] = float(not qid_to_has_ans[qid])
else:
new_scores[qid] = s
return new_scores
def make_eval_dict(exact_scores, f1_scores, qid_list=None):
if not qid_list:
total = len(exact_scores)
return collections.OrderedDict(
[
("exact", 100.0 * sum(exact_scores.values()) / total),
("f1", 100.0 * sum(f1_scores.values()) / total),
("total", total),
]
)
else:
total = len(qid_list)
return collections.OrderedDict(
[
("exact", 100.0 * sum(exact_scores[k] for k in qid_list) / total),
("f1", 100.0 * sum(f1_scores[k] for k in qid_list) / total),
("total", total),
]
)
def merge_eval(main_eval, new_eval, prefix):
for k in new_eval:
main_eval[f"{prefix}_{k}"] = new_eval[k]
def plot_pr_curve(precisions, recalls, out_image, title):
plt.step(recalls, precisions, color="b", alpha=0.2, where="post")
plt.fill_between(recalls, precisions, step="post", alpha=0.2, color="b")
plt.xlabel("Recall")
plt.ylabel("Precision")
plt.xlim([0.0, 1.05])
plt.ylim([0.0, 1.05])
plt.title(title)
plt.savefig(out_image)
plt.clf()
def make_precision_recall_eval(scores, na_probs, num_true_pos, qid_to_has_ans, out_image=None, title=None):
qid_list = sorted(na_probs, key=lambda k: na_probs[k])
true_pos = 0.0
cur_p = 1.0
cur_r = 0.0
precisions = [1.0]
recalls = [0.0]
avg_prec = 0.0
for i, qid in enumerate(qid_list):
if qid_to_has_ans[qid]:
true_pos += scores[qid]
cur_p = true_pos / float(i + 1)
cur_r = true_pos / float(num_true_pos)
if i == len(qid_list) - 1 or na_probs[qid] != na_probs[qid_list[i + 1]]:
# i.e., if we can put a threshold after this point
avg_prec += cur_p * (cur_r - recalls[-1])
precisions.append(cur_p)
recalls.append(cur_r)
if out_image:
plot_pr_curve(precisions, recalls, out_image, title)
return {"ap": 100.0 * avg_prec}
def run_precision_recall_analysis(main_eval, exact_raw, f1_raw, na_probs, qid_to_has_ans, out_image_dir):
if out_image_dir and not os.path.exists(out_image_dir):
os.makedirs(out_image_dir)
num_true_pos = sum(1 for v in qid_to_has_ans.values() if v)
if num_true_pos == 0:
return
pr_exact = make_precision_recall_eval(
exact_raw,
na_probs,
num_true_pos,
qid_to_has_ans,
out_image=os.path.join(out_image_dir, "pr_exact.png"),
title="Precision-Recall curve for Exact Match score",
)
pr_f1 = make_precision_recall_eval(
f1_raw,
na_probs,
num_true_pos,
qid_to_has_ans,
out_image=os.path.join(out_image_dir, "pr_f1.png"),
title="Precision-Recall curve for F1 score",
)
oracle_scores = {k: float(v) for k, v in qid_to_has_ans.items()}
pr_oracle = make_precision_recall_eval(
oracle_scores,
na_probs,
num_true_pos,
qid_to_has_ans,
out_image=os.path.join(out_image_dir, "pr_oracle.png"),
title="Oracle Precision-Recall curve (binary task of HasAns vs. NoAns)",
)
merge_eval(main_eval, pr_exact, "pr_exact")
merge_eval(main_eval, pr_f1, "pr_f1")
merge_eval(main_eval, pr_oracle, "pr_oracle")
def histogram_na_prob(na_probs, qid_list, image_dir, name):
if not qid_list:
return
x = [na_probs[k] for k in qid_list]
weights = np.ones_like(x) / float(len(x))
plt.hist(x, weights=weights, bins=20, range=(0.0, 1.0))
plt.xlabel("Model probability of no-answer")
plt.ylabel("Proportion of dataset")
plt.title(f"Histogram of no-answer probability: {name}")
plt.savefig(os.path.join(image_dir, f"na_prob_hist_{name}.png"))
plt.clf()
def find_best_thresh(preds, scores, na_probs, qid_to_has_ans):
num_no_ans = sum(1 for k in qid_to_has_ans if not qid_to_has_ans[k])
cur_score = num_no_ans
best_score = cur_score
best_thresh = 0.0
qid_list = sorted(na_probs, key=lambda k: na_probs[k])
for i, qid in enumerate(qid_list):
if qid not in scores:
continue
if qid_to_has_ans[qid]:
diff = scores[qid]
else:
if preds[qid]:
diff = -1
else:
diff = 0
cur_score += diff
if cur_score > best_score:
best_score = cur_score
best_thresh = na_probs[qid]
return 100.0 * best_score / len(scores), best_thresh
def find_all_best_thresh(main_eval, preds, exact_raw, f1_raw, na_probs, qid_to_has_ans):
best_exact, exact_thresh = find_best_thresh(preds, exact_raw, na_probs, qid_to_has_ans)
best_f1, f1_thresh = find_best_thresh(preds, f1_raw, na_probs, qid_to_has_ans)
main_eval["best_exact"] = best_exact
main_eval["best_exact_thresh"] = exact_thresh
main_eval["best_f1"] = best_f1
main_eval["best_f1_thresh"] = f1_thresh
def main():
with open(OPTS.data_file) as f:
dataset_json = json.load(f)
dataset = dataset_json["data"]
with open(OPTS.pred_file) as f:
preds = json.load(f)
if OPTS.na_prob_file:
with open(OPTS.na_prob_file) as f:
na_probs = json.load(f)
else:
na_probs = {k: 0.0 for k in preds}
qid_to_has_ans = make_qid_to_has_ans(dataset) # maps qid to True/False
has_ans_qids = [k for k, v in qid_to_has_ans.items() if v]
no_ans_qids = [k for k, v in qid_to_has_ans.items() if not v]
exact_raw, f1_raw = get_raw_scores(dataset, preds)
exact_thresh = apply_no_ans_threshold(exact_raw, na_probs, qid_to_has_ans, OPTS.na_prob_thresh)
f1_thresh = apply_no_ans_threshold(f1_raw, na_probs, qid_to_has_ans, OPTS.na_prob_thresh)
out_eval = make_eval_dict(exact_thresh, f1_thresh)
if has_ans_qids:
has_ans_eval = make_eval_dict(exact_thresh, f1_thresh, qid_list=has_ans_qids)
merge_eval(out_eval, has_ans_eval, "HasAns")
if no_ans_qids:
no_ans_eval = make_eval_dict(exact_thresh, f1_thresh, qid_list=no_ans_qids)
merge_eval(out_eval, no_ans_eval, "NoAns")
if OPTS.na_prob_file:
find_all_best_thresh(out_eval, preds, exact_raw, f1_raw, na_probs, qid_to_has_ans)
if OPTS.na_prob_file and OPTS.out_image_dir:
run_precision_recall_analysis(out_eval, exact_raw, f1_raw, na_probs, qid_to_has_ans, OPTS.out_image_dir)
histogram_na_prob(na_probs, has_ans_qids, OPTS.out_image_dir, "hasAns")
histogram_na_prob(na_probs, no_ans_qids, OPTS.out_image_dir, "noAns")
if OPTS.out_file:
with open(OPTS.out_file, "w") as f:
json.dump(out_eval, f)
else:
print(json.dumps(out_eval, indent=2))
if __name__ == "__main__":
OPTS = parse_args()
if OPTS.out_image_dir:
import matplotlib
matplotlib.use("Agg")
import matplotlib.pyplot as plt
main()
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/squad_v2/squad_v2.py
|
# Copyright 2020 The HuggingFace Datasets Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
""" SQuAD v2 metric. """
import datasets
from .evaluate import (
apply_no_ans_threshold,
find_all_best_thresh,
get_raw_scores,
make_eval_dict,
make_qid_to_has_ans,
merge_eval,
)
_CITATION = """\
@inproceedings{Rajpurkar2016SQuAD10,
title={SQuAD: 100, 000+ Questions for Machine Comprehension of Text},
author={Pranav Rajpurkar and Jian Zhang and Konstantin Lopyrev and Percy Liang},
booktitle={EMNLP},
year={2016}
}
"""
_DESCRIPTION = """
This metric wrap the official scoring script for version 2 of the Stanford Question
Answering Dataset (SQuAD).
Stanford Question Answering Dataset (SQuAD) is a reading comprehension dataset, consisting of questions posed by
crowdworkers on a set of Wikipedia articles, where the answer to every question is a segment of text, or span,
from the corresponding reading passage, or the question might be unanswerable.
SQuAD2.0 combines the 100,000 questions in SQuAD1.1 with over 50,000 unanswerable questions
written adversarially by crowdworkers to look similar to answerable ones.
To do well on SQuAD2.0, systems must not only answer questions when possible, but also
determine when no answer is supported by the paragraph and abstain from answering.
"""
_KWARGS_DESCRIPTION = """
Computes SQuAD v2 scores (F1 and EM).
Args:
predictions: List of triple for question-answers to score with the following elements:
- the question-answer 'id' field as given in the references (see below)
- the text of the answer
- the probability that the question has no answer
references: List of question-answers dictionaries with the following key-values:
- 'id': id of the question-answer pair (see above),
- 'answers': a list of Dict {'text': text of the answer as a string}
no_answer_threshold: float
Probability threshold to decide that a question has no answer.
Returns:
'exact': Exact match (the normalized answer exactly match the gold answer)
'f1': The F-score of predicted tokens versus the gold answer
'total': Number of score considered
'HasAns_exact': Exact match (the normalized answer exactly match the gold answer)
'HasAns_f1': The F-score of predicted tokens versus the gold answer
'HasAns_total': Number of score considered
'NoAns_exact': Exact match (the normalized answer exactly match the gold answer)
'NoAns_f1': The F-score of predicted tokens versus the gold answer
'NoAns_total': Number of score considered
'best_exact': Best exact match (with varying threshold)
'best_exact_thresh': No-answer probability threshold associated to the best exact match
'best_f1': Best F1 (with varying threshold)
'best_f1_thresh': No-answer probability threshold associated to the best F1
Examples:
>>> predictions = [{'prediction_text': '1976', 'id': '56e10a3be3433e1400422b22', 'no_answer_probability': 0.}]
>>> references = [{'answers': {'answer_start': [97], 'text': ['1976']}, 'id': '56e10a3be3433e1400422b22'}]
>>> squad_v2_metric = datasets.load_metric("squad_v2")
>>> results = squad_v2_metric.compute(predictions=predictions, references=references)
>>> print(results)
{'exact': 100.0, 'f1': 100.0, 'total': 1, 'HasAns_exact': 100.0, 'HasAns_f1': 100.0, 'HasAns_total': 1, 'best_exact': 100.0, 'best_exact_thresh': 0.0, 'best_f1': 100.0, 'best_f1_thresh': 0.0}
"""
@datasets.utils.file_utils.add_start_docstrings(_DESCRIPTION, _KWARGS_DESCRIPTION)
class SquadV2(datasets.Metric):
def _info(self):
return datasets.MetricInfo(
description=_DESCRIPTION,
citation=_CITATION,
inputs_description=_KWARGS_DESCRIPTION,
features=datasets.Features(
{
"predictions": {
"id": datasets.Value("string"),
"prediction_text": datasets.Value("string"),
"no_answer_probability": datasets.Value("float32"),
},
"references": {
"id": datasets.Value("string"),
"answers": datasets.features.Sequence(
{"text": datasets.Value("string"), "answer_start": datasets.Value("int32")}
),
},
}
),
codebase_urls=["https://rajpurkar.github.io/SQuAD-explorer/"],
reference_urls=["https://rajpurkar.github.io/SQuAD-explorer/"],
)
def _compute(self, predictions, references, no_answer_threshold=1.0):
no_answer_probabilities = {p["id"]: p["no_answer_probability"] for p in predictions}
dataset = [{"paragraphs": [{"qas": references}]}]
predictions = {p["id"]: p["prediction_text"] for p in predictions}
qid_to_has_ans = make_qid_to_has_ans(dataset) # maps qid to True/False
has_ans_qids = [k for k, v in qid_to_has_ans.items() if v]
no_ans_qids = [k for k, v in qid_to_has_ans.items() if not v]
exact_raw, f1_raw = get_raw_scores(dataset, predictions)
exact_thresh = apply_no_ans_threshold(exact_raw, no_answer_probabilities, qid_to_has_ans, no_answer_threshold)
f1_thresh = apply_no_ans_threshold(f1_raw, no_answer_probabilities, qid_to_has_ans, no_answer_threshold)
out_eval = make_eval_dict(exact_thresh, f1_thresh)
if has_ans_qids:
has_ans_eval = make_eval_dict(exact_thresh, f1_thresh, qid_list=has_ans_qids)
merge_eval(out_eval, has_ans_eval, "HasAns")
if no_ans_qids:
no_ans_eval = make_eval_dict(exact_thresh, f1_thresh, qid_list=no_ans_qids)
merge_eval(out_eval, no_ans_eval, "NoAns")
find_all_best_thresh(out_eval, predictions, exact_raw, f1_raw, no_answer_probabilities, qid_to_has_ans)
return dict(out_eval)
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/perplexity/README.md
|
# Metric Card for Perplexity
## Metric Description
Given a model and an input text sequence, perplexity measures how likely the model is to generate the input text sequence. This can be used in two main ways:
1. to evaluate how well the model has learned the distribution of the text it was trained on
- In this case, the model input should be the trained model to be evaluated, and the input texts should be the text that the model was trained on.
2. to evaluate how well a selection of text matches the distribution of text that the input model was trained on
- In this case, the model input should be a trained model, and the input texts should be the text to be evaluated.
## Intended Uses
Any language generation task.
## How to Use
The metric takes a list of text as input, as well as the name of the model used to compute the metric:
```python
from datasets import load_metric
perplexity = load_metric("perplexity")
results = perplexity.compute(input_texts=input_texts, model_id='gpt2')
```
### Inputs
- **model_id** (str): model used for calculating Perplexity. NOTE: Perplexity can only be calculated for causal language models.
- This includes models such as gpt2, causal variations of bert, causal versions of t5, and more (the full list can be found in the AutoModelForCausalLM documentation here: https://huggingface.co/docs/transformers/master/en/model_doc/auto#transformers.AutoModelForCausalLM )
- **input_texts** (list of str): input text, each separate text snippet is one list entry.
- **batch_size** (int): the batch size to run texts through the model. Defaults to 16.
- **add_start_token** (bool): whether to add the start token to the texts, so the perplexity can include the probability of the first word. Defaults to True.
- **device** (str): device to run on, defaults to 'cuda' when available
### Output Values
This metric outputs a dictionary with the perplexity scores for the text input in the list, and the average perplexity.
If one of the input texts is longer than the max input length of the model, then it is truncated to the max length for the perplexity computation.
```
{'perplexities': [8.182524681091309, 33.42122268676758, 27.012239456176758], 'mean_perplexity': 22.871995608011883}
```
This metric's range is 0 and up. A lower score is better.
#### Values from Popular Papers
### Examples
Calculating perplexity on input_texts defined here:
```python
perplexity = datasets.load_metric("perplexity")
input_texts = ["lorem ipsum", "Happy Birthday!", "Bienvenue"]
results = perplexity.compute(model_id='gpt2',
add_start_token=False,
input_texts=input_texts)
print(list(results.keys()))
>>>['perplexities', 'mean_perplexity']
print(round(results["mean_perplexity"], 2))
>>>78.22
print(round(results["perplexities"][0], 2))
>>>11.11
```
Calculating perplexity on input_texts loaded in from a dataset:
```python
perplexity = datasets.load_metric("perplexity")
input_texts = datasets.load_dataset("wikitext",
"wikitext-2-raw-v1",
split="test")["text"][:50]
input_texts = [s for s in input_texts if s!='']
results = perplexity.compute(model_id='gpt2',
input_texts=input_texts)
print(list(results.keys()))
>>>['perplexities', 'mean_perplexity']
print(round(results["mean_perplexity"], 2))
>>>60.35
print(round(results["perplexities"][0], 2))
>>>81.12
```
## Limitations and Bias
Note that the output value is based heavily on what text the model was trained on. This means that perplexity scores are not comparable between models or datasets.
## Citation
```bibtex
@article{jelinek1977perplexity,
title={Perplexity—a measure of the difficulty of speech recognition tasks},
author={Jelinek, Fred and Mercer, Robert L and Bahl, Lalit R and Baker, James K},
journal={The Journal of the Acoustical Society of America},
volume={62},
number={S1},
pages={S63--S63},
year={1977},
publisher={Acoustical Society of America}
}
```
## Further References
- [Hugging Face Perplexity Blog Post](https://huggingface.co/docs/transformers/perplexity)
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/perplexity/perplexity.py
|
# Copyright 2022 The HuggingFace Datasets Authors and the current dataset script contributor.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""Perplexity Metric."""
import numpy as np
import torch
from torch.nn import CrossEntropyLoss
from transformers import AutoModelForCausalLM, AutoTokenizer
import datasets
from datasets import logging
_CITATION = """\
"""
_DESCRIPTION = """
Perplexity (PPL) is one of the most common metrics for evaluating language models.
It is defined as the exponentiated average negative log-likelihood of a sequence.
For more information, see https://huggingface.co/docs/transformers/perplexity
"""
_KWARGS_DESCRIPTION = """
Args:
model_id (str): model used for calculating Perplexity
NOTE: Perplexity can only be calculated for causal language models.
This includes models such as gpt2, causal variations of bert,
causal versions of t5, and more (the full list can be found
in the AutoModelForCausalLM documentation here:
https://huggingface.co/docs/transformers/master/en/model_doc/auto#transformers.AutoModelForCausalLM )
input_texts (list of str): input text, each separate text snippet
is one list entry.
batch_size (int): the batch size to run texts through the model. Defaults to 16.
add_start_token (bool): whether to add the start token to the texts,
so the perplexity can include the probability of the first word. Defaults to True.
device (str): device to run on, defaults to 'cuda' when available
Returns:
perplexity: dictionary containing the perplexity scores for the texts
in the input list, as well as the mean perplexity. If one of the input texts is
longer than the max input length of the model, then it is truncated to the
max length for the perplexity computation.
Examples:
Example 1:
>>> perplexity = datasets.load_metric("perplexity")
>>> input_texts = ["lorem ipsum", "Happy Birthday!", "Bienvenue"]
>>> results = perplexity.compute(model_id='gpt2',
... add_start_token=False,
... input_texts=input_texts) # doctest:+ELLIPSIS
>>> print(list(results.keys()))
['perplexities', 'mean_perplexity']
>>> print(round(results["mean_perplexity"], 2))
78.22
>>> print(round(results["perplexities"][0], 2))
11.11
Example 2:
>>> perplexity = datasets.load_metric("perplexity")
>>> input_texts = datasets.load_dataset("wikitext",
... "wikitext-2-raw-v1",
... split="test")["text"][:50]
>>> input_texts = [s for s in input_texts if s!='']
>>> results = perplexity.compute(model_id='gpt2',
... input_texts=input_texts) # doctest:+ELLIPSIS
>>> print(list(results.keys()))
['perplexities', 'mean_perplexity']
>>> print(round(results["mean_perplexity"], 2))
60.35
>>> print(round(results["perplexities"][0], 2))
81.12
"""
@datasets.utils.file_utils.add_start_docstrings(_DESCRIPTION, _KWARGS_DESCRIPTION)
class Perplexity(datasets.Metric):
def _info(self):
return datasets.MetricInfo(
description=_DESCRIPTION,
citation=_CITATION,
inputs_description=_KWARGS_DESCRIPTION,
features=datasets.Features(
{
"input_texts": datasets.Value("string"),
}
),
reference_urls=["https://huggingface.co/docs/transformers/perplexity"],
)
def _compute(self, input_texts, model_id, batch_size: int = 16, add_start_token: bool = True, device=None):
if device is not None:
assert device in ["gpu", "cpu", "cuda"], "device should be either gpu or cpu."
if device == "gpu":
device = "cuda"
else:
device = "cuda" if torch.cuda.is_available() else "cpu"
model = AutoModelForCausalLM.from_pretrained(model_id)
model = model.to(device)
tokenizer = AutoTokenizer.from_pretrained(model_id)
# if batch_size > 1 (which generally leads to padding being required), and
# if there is not an already assigned pad_token, assign an existing
# special token to also be the padding token
if tokenizer.pad_token is None and batch_size > 1:
existing_special_tokens = list(tokenizer.special_tokens_map_extended.values())
# check that the model already has at least one special token defined
assert (
len(existing_special_tokens) > 0
), "If batch_size > 1, model must have at least one special token to use for padding. Please use a different model or set batch_size=1."
# assign one of the special tokens to also be the pad token
tokenizer.add_special_tokens({"pad_token": existing_special_tokens[0]})
if add_start_token:
# leave room for <BOS> token to be added:
assert (
tokenizer.bos_token is not None
), "Input model must already have a BOS token if using add_start_token=True. Please use a different model, or set add_start_token=False"
max_tokenized_len = model.config.max_length - 1
else:
max_tokenized_len = model.config.max_length
encodings = tokenizer(
input_texts,
add_special_tokens=False,
padding=True,
truncation=True,
max_length=max_tokenized_len,
return_tensors="pt",
return_attention_mask=True,
).to(device)
encoded_texts = encodings["input_ids"]
attn_masks = encodings["attention_mask"]
# check that each input is long enough:
if add_start_token:
assert torch.all(torch.ge(attn_masks.sum(1), 1)), "Each input text must be at least one token long."
else:
assert torch.all(
torch.ge(attn_masks.sum(1), 2)
), "When add_start_token=False, each input text must be at least two tokens long. Run with add_start_token=True if inputting strings of only one token, and remove all empty input strings."
ppls = []
loss_fct = CrossEntropyLoss(reduction="none")
for start_index in logging.tqdm(range(0, len(encoded_texts), batch_size)):
end_index = min(start_index + batch_size, len(encoded_texts))
encoded_batch = encoded_texts[start_index:end_index]
attn_mask = attn_masks[start_index:end_index]
if add_start_token:
bos_tokens_tensor = torch.tensor([[tokenizer.bos_token_id]] * encoded_batch.size(dim=0)).to(device)
encoded_batch = torch.cat([bos_tokens_tensor, encoded_batch], dim=1)
attn_mask = torch.cat(
[torch.ones(bos_tokens_tensor.size(), dtype=torch.int64).to(device), attn_mask], dim=1
)
labels = encoded_batch
with torch.no_grad():
out_logits = model(encoded_batch, attention_mask=attn_mask).logits
shift_logits = out_logits[..., :-1, :].contiguous()
shift_labels = labels[..., 1:].contiguous()
shift_attention_mask_batch = attn_mask[..., 1:].contiguous()
perplexity_batch = torch.exp2(
(loss_fct(shift_logits.transpose(1, 2), shift_labels) * shift_attention_mask_batch).sum(1)
/ shift_attention_mask_batch.sum(1)
)
ppls += perplexity_batch.tolist()
return {"perplexities": ppls, "mean_perplexity": np.mean(ppls)}
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/recall/README.md
|
# Metric Card for Recall
## Metric Description
Recall is the fraction of the positive examples that were correctly labeled by the model as positive. It can be computed with the equation:
Recall = TP / (TP + FN)
Where TP is the number of true positives and FN is the number of false negatives.
## How to Use
At minimum, this metric takes as input two `list`s, each containing `int`s: predictions and references.
```python
>>> recall_metric = datasets.load_metric('recall')
>>> results = recall_metric.compute(references=[0, 1], predictions=[0, 1])
>>> print(results)
["{'recall': 1.0}"]
```
### Inputs
- **predictions** (`list` of `int`): The predicted labels.
- **references** (`list` of `int`): The ground truth labels.
- **labels** (`list` of `int`): The set of labels to include when `average` is not set to `binary`, and their order when average is `None`. Labels present in the data can be excluded in this input, for example to calculate a multiclass average ignoring a majority negative class, while labels not present in the data will result in 0 components in a macro average. For multilabel targets, labels are column indices. By default, all labels in y_true and y_pred are used in sorted order. Defaults to None.
- **pos_label** (`int`): The class label to use as the 'positive class' when calculating the recall. Defaults to `1`.
- **average** (`string`): This parameter is required for multiclass/multilabel targets. If None, the scores for each class are returned. Otherwise, this determines the type of averaging performed on the data. Defaults to `'binary'`.
- `'binary'`: Only report results for the class specified by `pos_label`. This is applicable only if the target labels and predictions are binary.
- `'micro'`: Calculate metrics globally by counting the total true positives, false negatives, and false positives.
- `'macro'`: Calculate metrics for each label, and find their unweighted mean. This does not take label imbalance into account.
- `'weighted'`: Calculate metrics for each label, and find their average weighted by support (the number of true instances for each label). This alters `'macro'` to account for label imbalance. Note that it can result in an F-score that is not between precision and recall.
- `'samples'`: Calculate metrics for each instance, and find their average (only meaningful for multilabel classification).
- **sample_weight** (`list` of `float`): Sample weights Defaults to `None`.
- **zero_division** (): Sets the value to return when there is a zero division. Defaults to .
- `'warn'`: If there is a zero division, the return value is `0`, but warnings are also raised.
- `0`: If there is a zero division, the return value is `0`.
- `1`: If there is a zero division, the return value is `1`.
### Output Values
- **recall**(`float`, or `array` of `float`, for multiclass targets): Either the general recall score, or the recall scores for individual classes, depending on the values input to `labels` and `average`. Minimum possible value is 0. Maximum possible value is 1. A higher recall means that more of the positive examples have been labeled correctly. Therefore, a higher recall is generally considered better.
Output Example(s):
```python
{'recall': 1.0}
```
```python
{'recall': array([1., 0., 0.])}
```
This metric outputs a dictionary with one entry, `'recall'`.
#### Values from Popular Papers
### Examples
Example 1-A simple example with some errors
```python
>>> recall_metric = datasets.load_metric('recall')
>>> results = recall_metric.compute(references=[0, 0, 1, 1, 1], predictions=[0, 1, 0, 1, 1])
>>> print(results)
{'recall': 0.6666666666666666}
```
Example 2-The same example as Example 1, but with `pos_label=0` instead of the default `pos_label=1`.
```python
>>> recall_metric = datasets.load_metric('recall')
>>> results = recall_metric.compute(references=[0, 0, 1, 1, 1], predictions=[0, 1, 0, 1, 1], pos_label=0)
>>> print(results)
{'recall': 0.5}
```
Example 3-The same example as Example 1, but with `sample_weight` included.
```python
>>> recall_metric = datasets.load_metric('recall')
>>> sample_weight = [0.9, 0.2, 0.9, 0.3, 0.8]
>>> results = recall_metric.compute(references=[0, 0, 1, 1, 1], predictions=[0, 1, 0, 1, 1], sample_weight=sample_weight)
>>> print(results)
{'recall': 0.55}
```
Example 4-A multiclass example, using different averages.
```python
>>> recall_metric = datasets.load_metric('recall')
>>> predictions = [0, 2, 1, 0, 0, 1]
>>> references = [0, 1, 2, 0, 1, 2]
>>> results = recall_metric.compute(predictions=predictions, references=references, average='macro')
>>> print(results)
{'recall': 0.3333333333333333}
>>> results = recall_metric.compute(predictions=predictions, references=references, average='micro')
>>> print(results)
{'recall': 0.3333333333333333}
>>> results = recall_metric.compute(predictions=predictions, references=references, average='weighted')
>>> print(results)
{'recall': 0.3333333333333333}
>>> results = recall_metric.compute(predictions=predictions, references=references, average=None)
>>> print(results)
{'recall': array([1., 0., 0.])}
```
## Limitations and Bias
## Citation(s)
```bibtex
@article{scikit-learn, title={Scikit-learn: Machine Learning in {P}ython}, author={Pedregosa, F. and Varoquaux, G. and Gramfort, A. and Michel, V. and Thirion, B. and Grisel, O. and Blondel, M. and Prettenhofer, P. and Weiss, R. and Dubourg, V. and Vanderplas, J. and Passos, A. and Cournapeau, D. and Brucher, M. and Perrot, M. and Duchesnay, E.}, journal={Journal of Machine Learning Research}, volume={12}, pages={2825--2830}, year={2011}
```
## Further References
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/recall/recall.py
|
# Copyright 2020 The HuggingFace Datasets Authors and the current dataset script contributor.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""Recall metric."""
from sklearn.metrics import recall_score
import datasets
_DESCRIPTION = """
Recall is the fraction of the positive examples that were correctly labeled by the model as positive. It can be computed with the equation:
Recall = TP / (TP + FN)
Where TP is the true positives and FN is the false negatives.
"""
_KWARGS_DESCRIPTION = """
Args:
- **predictions** (`list` of `int`): The predicted labels.
- **references** (`list` of `int`): The ground truth labels.
- **labels** (`list` of `int`): The set of labels to include when `average` is not set to `binary`, and their order when average is `None`. Labels present in the data can be excluded in this input, for example to calculate a multiclass average ignoring a majority negative class, while labels not present in the data will result in 0 components in a macro average. For multilabel targets, labels are column indices. By default, all labels in y_true and y_pred are used in sorted order. Defaults to None.
- **pos_label** (`int`): The class label to use as the 'positive class' when calculating the recall. Defaults to `1`.
- **average** (`string`): This parameter is required for multiclass/multilabel targets. If None, the scores for each class are returned. Otherwise, this determines the type of averaging performed on the data. Defaults to `'binary'`.
- `'binary'`: Only report results for the class specified by `pos_label`. This is applicable only if the target labels and predictions are binary.
- `'micro'`: Calculate metrics globally by counting the total true positives, false negatives, and false positives.
- `'macro'`: Calculate metrics for each label, and find their unweighted mean. This does not take label imbalance into account.
- `'weighted'`: Calculate metrics for each label, and find their average weighted by support (the number of true instances for each label). This alters `'macro'` to account for label imbalance. Note that it can result in an F-score that is not between precision and recall.
- `'samples'`: Calculate metrics for each instance, and find their average (only meaningful for multilabel classification).
- **sample_weight** (`list` of `float`): Sample weights Defaults to `None`.
- **zero_division** (): Sets the value to return when there is a zero division. Defaults to .
- `'warn'`: If there is a zero division, the return value is `0`, but warnings are also raised.
- `0`: If there is a zero division, the return value is `0`.
- `1`: If there is a zero division, the return value is `1`.
Returns:
- **recall** (`float`, or `array` of `float`): Either the general recall score, or the recall scores for individual classes, depending on the values input to `labels` and `average`. Minimum possible value is 0. Maximum possible value is 1. A higher recall means that more of the positive examples have been labeled correctly. Therefore, a higher recall is generally considered better.
Examples:
Example 1-A simple example with some errors
>>> recall_metric = datasets.load_metric('recall')
>>> results = recall_metric.compute(references=[0, 0, 1, 1, 1], predictions=[0, 1, 0, 1, 1])
>>> print(results)
{'recall': 0.6666666666666666}
Example 2-The same example as Example 1, but with `pos_label=0` instead of the default `pos_label=1`.
>>> recall_metric = datasets.load_metric('recall')
>>> results = recall_metric.compute(references=[0, 0, 1, 1, 1], predictions=[0, 1, 0, 1, 1], pos_label=0)
>>> print(results)
{'recall': 0.5}
Example 3-The same example as Example 1, but with `sample_weight` included.
>>> recall_metric = datasets.load_metric('recall')
>>> sample_weight = [0.9, 0.2, 0.9, 0.3, 0.8]
>>> results = recall_metric.compute(references=[0, 0, 1, 1, 1], predictions=[0, 1, 0, 1, 1], sample_weight=sample_weight)
>>> print(results)
{'recall': 0.55}
Example 4-A multiclass example, using different averages.
>>> recall_metric = datasets.load_metric('recall')
>>> predictions = [0, 2, 1, 0, 0, 1]
>>> references = [0, 1, 2, 0, 1, 2]
>>> results = recall_metric.compute(predictions=predictions, references=references, average='macro')
>>> print(results)
{'recall': 0.3333333333333333}
>>> results = recall_metric.compute(predictions=predictions, references=references, average='micro')
>>> print(results)
{'recall': 0.3333333333333333}
>>> results = recall_metric.compute(predictions=predictions, references=references, average='weighted')
>>> print(results)
{'recall': 0.3333333333333333}
>>> results = recall_metric.compute(predictions=predictions, references=references, average=None)
>>> print(results)
{'recall': array([1., 0., 0.])}
"""
_CITATION = """
@article{scikit-learn, title={Scikit-learn: Machine Learning in {P}ython}, author={Pedregosa, F. and Varoquaux, G. and Gramfort, A. and Michel, V. and Thirion, B. and Grisel, O. and Blondel, M. and Prettenhofer, P. and Weiss, R. and Dubourg, V. and Vanderplas, J. and Passos, A. and Cournapeau, D. and Brucher, M. and Perrot, M. and Duchesnay, E.}, journal={Journal of Machine Learning Research}, volume={12}, pages={2825--2830}, year={2011}
"""
@datasets.utils.file_utils.add_start_docstrings(_DESCRIPTION, _KWARGS_DESCRIPTION)
class Recall(datasets.Metric):
def _info(self):
return datasets.MetricInfo(
description=_DESCRIPTION,
citation=_CITATION,
inputs_description=_KWARGS_DESCRIPTION,
features=datasets.Features(
{
"predictions": datasets.Sequence(datasets.Value("int32")),
"references": datasets.Sequence(datasets.Value("int32")),
}
if self.config_name == "multilabel"
else {
"predictions": datasets.Value("int32"),
"references": datasets.Value("int32"),
}
),
reference_urls=["https://scikit-learn.org/stable/modules/generated/sklearn.metrics.recall_score.html"],
)
def _compute(
self,
predictions,
references,
labels=None,
pos_label=1,
average="binary",
sample_weight=None,
zero_division="warn",
):
score = recall_score(
references,
predictions,
labels=labels,
pos_label=pos_label,
average=average,
sample_weight=sample_weight,
zero_division=zero_division,
)
return {"recall": float(score) if score.size == 1 else score}
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/sari/README.md
|
# Metric Card for SARI
## Metric description
SARI (***s**ystem output **a**gainst **r**eferences and against the **i**nput sentence*) is a metric used for evaluating automatic text simplification systems.
The metric compares the predicted simplified sentences against the reference and the source sentences. It explicitly measures the goodness of words that are added, deleted and kept by the system.
SARI can be computed as:
`sari = ( F1_add + F1_keep + P_del) / 3`
where
`F1_add` is the n-gram F1 score for add operations
`F1_keep` is the n-gram F1 score for keep operations
`P_del` is the n-gram precision score for delete operations
The number of n grams, `n`, is equal to 4, as in the original paper.
This implementation is adapted from [Tensorflow's tensor2tensor implementation](https://github.com/tensorflow/tensor2tensor/blob/master/tensor2tensor/utils/sari_hook.py).
It has two differences with the [original GitHub implementation](https://github.com/cocoxu/simplification/blob/master/SARI.py):
1) It defines 0/0=1 instead of 0 to give higher scores for predictions that match a target exactly.
2) It fixes an [alleged bug](https://github.com/cocoxu/simplification/issues/6) in the keep score computation.
## How to use
The metric takes 3 inputs: sources (a list of source sentence strings), predictions (a list of predicted sentence strings) and references (a list of lists of reference sentence strings)
```python
from datasets import load_metric
sari = load_metric("sari")
sources=["About 95 species are currently accepted."]
predictions=["About 95 you now get in."]
references=[["About 95 species are currently known.","About 95 species are now accepted.","95 species are now accepted."]]
sari_score = sari.compute(sources=sources, predictions=predictions, references=references)
```
## Output values
This metric outputs a dictionary with the SARI score:
```
print(sari_score)
{'sari': 26.953601953601954}
```
The range of values for the SARI score is between 0 and 100 -- the higher the value, the better the performance of the model being evaluated, with a SARI of 100 being a perfect score.
### Values from popular papers
The [original paper that proposes the SARI metric](https://aclanthology.org/Q16-1029.pdf) reports scores ranging from 26 to 43 for different simplification systems and different datasets. They also find that the metric ranks all of the simplification systems and human references in the same order as the human assessment used as a comparison, and that it correlates reasonably with human judgments.
More recent SARI scores for text simplification can be found on leaderboards for datasets such as [TurkCorpus](https://paperswithcode.com/sota/text-simplification-on-turkcorpus) and [Newsela](https://paperswithcode.com/sota/text-simplification-on-newsela).
## Examples
Perfect match between prediction and reference:
```python
from datasets import load_metric
sari = load_metric("sari")
sources=["About 95 species are currently accepted ."]
predictions=["About 95 species are currently accepted ."]
references=[["About 95 species are currently accepted ."]]
sari_score = sari.compute(sources=sources, predictions=predictions, references=references)
print(sari_score)
{'sari': 100.0}
```
Partial match between prediction and reference:
```python
from datasets import load_metric
sari = load_metric("sari")
sources=["About 95 species are currently accepted ."]
predictions=["About 95 you now get in ."]
references=[["About 95 species are currently known .","About 95 species are now accepted .","95 species are now accepted ."]]
sari_score = sari.compute(sources=sources, predictions=predictions, references=references)
print(sari_score)
{'sari': 26.953601953601954}
```
## Limitations and bias
SARI is a valuable measure for comparing different text simplification systems as well as one that can assist the iterative development of a system.
However, while the [original paper presenting SARI](https://aclanthology.org/Q16-1029.pdf) states that it captures "the notion of grammaticality and meaning preservation", this is a difficult claim to empirically validate.
## Citation
```bibtex
@inproceedings{xu-etal-2016-optimizing,
title = {Optimizing Statistical Machine Translation for Text Simplification},
authors={Xu, Wei and Napoles, Courtney and Pavlick, Ellie and Chen, Quanze and Callison-Burch, Chris},
journal = {Transactions of the Association for Computational Linguistics},
volume = {4},
year={2016},
url = {https://www.aclweb.org/anthology/Q16-1029},
pages = {401--415},
}
```
## Further References
- [NLP Progress -- Text Simplification](http://nlpprogress.com/english/simplification.html)
- [Hugging Face Hub -- Text Simplification Models](https://huggingface.co/datasets?filter=task_ids:text-simplification)
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/sari/sari.py
|
# Copyright 2020 The HuggingFace Datasets Authors and the current dataset script contributor.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
""" SARI metric."""
from collections import Counter
import sacrebleu
import sacremoses
from packaging import version
import datasets
_CITATION = """\
@inproceedings{xu-etal-2016-optimizing,
title = {Optimizing Statistical Machine Translation for Text Simplification},
authors={Xu, Wei and Napoles, Courtney and Pavlick, Ellie and Chen, Quanze and Callison-Burch, Chris},
journal = {Transactions of the Association for Computational Linguistics},
volume = {4},
year={2016},
url = {https://www.aclweb.org/anthology/Q16-1029},
pages = {401--415},
}
"""
_DESCRIPTION = """\
SARI is a metric used for evaluating automatic text simplification systems.
The metric compares the predicted simplified sentences against the reference
and the source sentences. It explicitly measures the goodness of words that are
added, deleted and kept by the system.
Sari = (F1_add + F1_keep + P_del) / 3
where
F1_add: n-gram F1 score for add operation
F1_keep: n-gram F1 score for keep operation
P_del: n-gram precision score for delete operation
n = 4, as in the original paper.
This implementation is adapted from Tensorflow's tensor2tensor implementation [3].
It has two differences with the original GitHub [1] implementation:
(1) Defines 0/0=1 instead of 0 to give higher scores for predictions that match
a target exactly.
(2) Fixes an alleged bug [2] in the keep score computation.
[1] https://github.com/cocoxu/simplification/blob/master/SARI.py
(commit 0210f15)
[2] https://github.com/cocoxu/simplification/issues/6
[3] https://github.com/tensorflow/tensor2tensor/blob/master/tensor2tensor/utils/sari_hook.py
"""
_KWARGS_DESCRIPTION = """
Calculates sari score (between 0 and 100) given a list of source and predicted
sentences, and a list of lists of reference sentences.
Args:
sources: list of source sentences where each sentence should be a string.
predictions: list of predicted sentences where each sentence should be a string.
references: list of lists of reference sentences where each sentence should be a string.
Returns:
sari: sari score
Examples:
>>> sources=["About 95 species are currently accepted ."]
>>> predictions=["About 95 you now get in ."]
>>> references=[["About 95 species are currently known .","About 95 species are now accepted .","95 species are now accepted ."]]
>>> sari = datasets.load_metric("sari")
>>> results = sari.compute(sources=sources, predictions=predictions, references=references)
>>> print(results)
{'sari': 26.953601953601954}
"""
def SARIngram(sgrams, cgrams, rgramslist, numref):
rgramsall = [rgram for rgrams in rgramslist for rgram in rgrams]
rgramcounter = Counter(rgramsall)
sgramcounter = Counter(sgrams)
sgramcounter_rep = Counter()
for sgram, scount in sgramcounter.items():
sgramcounter_rep[sgram] = scount * numref
cgramcounter = Counter(cgrams)
cgramcounter_rep = Counter()
for cgram, ccount in cgramcounter.items():
cgramcounter_rep[cgram] = ccount * numref
# KEEP
keepgramcounter_rep = sgramcounter_rep & cgramcounter_rep
keepgramcountergood_rep = keepgramcounter_rep & rgramcounter
keepgramcounterall_rep = sgramcounter_rep & rgramcounter
keeptmpscore1 = 0
keeptmpscore2 = 0
for keepgram in keepgramcountergood_rep:
keeptmpscore1 += keepgramcountergood_rep[keepgram] / keepgramcounter_rep[keepgram]
# Fix an alleged bug [2] in the keep score computation.
# keeptmpscore2 += keepgramcountergood_rep[keepgram] / keepgramcounterall_rep[keepgram]
keeptmpscore2 += keepgramcountergood_rep[keepgram]
# Define 0/0=1 instead of 0 to give higher scores for predictions that match
# a target exactly.
keepscore_precision = 1
keepscore_recall = 1
if len(keepgramcounter_rep) > 0:
keepscore_precision = keeptmpscore1 / len(keepgramcounter_rep)
if len(keepgramcounterall_rep) > 0:
# Fix an alleged bug [2] in the keep score computation.
# keepscore_recall = keeptmpscore2 / len(keepgramcounterall_rep)
keepscore_recall = keeptmpscore2 / sum(keepgramcounterall_rep.values())
keepscore = 0
if keepscore_precision > 0 or keepscore_recall > 0:
keepscore = 2 * keepscore_precision * keepscore_recall / (keepscore_precision + keepscore_recall)
# DELETION
delgramcounter_rep = sgramcounter_rep - cgramcounter_rep
delgramcountergood_rep = delgramcounter_rep - rgramcounter
delgramcounterall_rep = sgramcounter_rep - rgramcounter
deltmpscore1 = 0
deltmpscore2 = 0
for delgram in delgramcountergood_rep:
deltmpscore1 += delgramcountergood_rep[delgram] / delgramcounter_rep[delgram]
deltmpscore2 += delgramcountergood_rep[delgram] / delgramcounterall_rep[delgram]
# Define 0/0=1 instead of 0 to give higher scores for predictions that match
# a target exactly.
delscore_precision = 1
if len(delgramcounter_rep) > 0:
delscore_precision = deltmpscore1 / len(delgramcounter_rep)
# ADDITION
addgramcounter = set(cgramcounter) - set(sgramcounter)
addgramcountergood = set(addgramcounter) & set(rgramcounter)
addgramcounterall = set(rgramcounter) - set(sgramcounter)
addtmpscore = 0
for addgram in addgramcountergood:
addtmpscore += 1
# Define 0/0=1 instead of 0 to give higher scores for predictions that match
# a target exactly.
addscore_precision = 1
addscore_recall = 1
if len(addgramcounter) > 0:
addscore_precision = addtmpscore / len(addgramcounter)
if len(addgramcounterall) > 0:
addscore_recall = addtmpscore / len(addgramcounterall)
addscore = 0
if addscore_precision > 0 or addscore_recall > 0:
addscore = 2 * addscore_precision * addscore_recall / (addscore_precision + addscore_recall)
return (keepscore, delscore_precision, addscore)
def SARIsent(ssent, csent, rsents):
numref = len(rsents)
s1grams = ssent.split(" ")
c1grams = csent.split(" ")
s2grams = []
c2grams = []
s3grams = []
c3grams = []
s4grams = []
c4grams = []
r1gramslist = []
r2gramslist = []
r3gramslist = []
r4gramslist = []
for rsent in rsents:
r1grams = rsent.split(" ")
r2grams = []
r3grams = []
r4grams = []
r1gramslist.append(r1grams)
for i in range(0, len(r1grams) - 1):
if i < len(r1grams) - 1:
r2gram = r1grams[i] + " " + r1grams[i + 1]
r2grams.append(r2gram)
if i < len(r1grams) - 2:
r3gram = r1grams[i] + " " + r1grams[i + 1] + " " + r1grams[i + 2]
r3grams.append(r3gram)
if i < len(r1grams) - 3:
r4gram = r1grams[i] + " " + r1grams[i + 1] + " " + r1grams[i + 2] + " " + r1grams[i + 3]
r4grams.append(r4gram)
r2gramslist.append(r2grams)
r3gramslist.append(r3grams)
r4gramslist.append(r4grams)
for i in range(0, len(s1grams) - 1):
if i < len(s1grams) - 1:
s2gram = s1grams[i] + " " + s1grams[i + 1]
s2grams.append(s2gram)
if i < len(s1grams) - 2:
s3gram = s1grams[i] + " " + s1grams[i + 1] + " " + s1grams[i + 2]
s3grams.append(s3gram)
if i < len(s1grams) - 3:
s4gram = s1grams[i] + " " + s1grams[i + 1] + " " + s1grams[i + 2] + " " + s1grams[i + 3]
s4grams.append(s4gram)
for i in range(0, len(c1grams) - 1):
if i < len(c1grams) - 1:
c2gram = c1grams[i] + " " + c1grams[i + 1]
c2grams.append(c2gram)
if i < len(c1grams) - 2:
c3gram = c1grams[i] + " " + c1grams[i + 1] + " " + c1grams[i + 2]
c3grams.append(c3gram)
if i < len(c1grams) - 3:
c4gram = c1grams[i] + " " + c1grams[i + 1] + " " + c1grams[i + 2] + " " + c1grams[i + 3]
c4grams.append(c4gram)
(keep1score, del1score, add1score) = SARIngram(s1grams, c1grams, r1gramslist, numref)
(keep2score, del2score, add2score) = SARIngram(s2grams, c2grams, r2gramslist, numref)
(keep3score, del3score, add3score) = SARIngram(s3grams, c3grams, r3gramslist, numref)
(keep4score, del4score, add4score) = SARIngram(s4grams, c4grams, r4gramslist, numref)
avgkeepscore = sum([keep1score, keep2score, keep3score, keep4score]) / 4
avgdelscore = sum([del1score, del2score, del3score, del4score]) / 4
avgaddscore = sum([add1score, add2score, add3score, add4score]) / 4
finalscore = (avgkeepscore + avgdelscore + avgaddscore) / 3
return finalscore
def normalize(sentence, lowercase: bool = True, tokenizer: str = "13a", return_str: bool = True):
# Normalization is requried for the ASSET dataset (one of the primary
# datasets in sentence simplification) to allow using space
# to split the sentence. Even though Wiki-Auto and TURK datasets,
# do not require normalization, we do it for consistency.
# Code adapted from the EASSE library [1] written by the authors of the ASSET dataset.
# [1] https://github.com/feralvam/easse/blob/580bba7e1378fc8289c663f864e0487188fe8067/easse/utils/preprocessing.py#L7
if lowercase:
sentence = sentence.lower()
if tokenizer in ["13a", "intl"]:
if version.parse(sacrebleu.__version__).major >= 2:
normalized_sent = sacrebleu.metrics.bleu._get_tokenizer(tokenizer)()(sentence)
else:
normalized_sent = sacrebleu.TOKENIZERS[tokenizer]()(sentence)
elif tokenizer == "moses":
normalized_sent = sacremoses.MosesTokenizer().tokenize(sentence, return_str=True, escape=False)
elif tokenizer == "penn":
normalized_sent = sacremoses.MosesTokenizer().penn_tokenize(sentence, return_str=True)
else:
normalized_sent = sentence
if not return_str:
normalized_sent = normalized_sent.split()
return normalized_sent
@datasets.utils.file_utils.add_start_docstrings(_DESCRIPTION, _KWARGS_DESCRIPTION)
class Sari(datasets.Metric):
def _info(self):
return datasets.MetricInfo(
description=_DESCRIPTION,
citation=_CITATION,
inputs_description=_KWARGS_DESCRIPTION,
features=datasets.Features(
{
"sources": datasets.Value("string", id="sequence"),
"predictions": datasets.Value("string", id="sequence"),
"references": datasets.Sequence(datasets.Value("string", id="sequence"), id="references"),
}
),
codebase_urls=[
"https://github.com/cocoxu/simplification/blob/master/SARI.py",
"https://github.com/tensorflow/tensor2tensor/blob/master/tensor2tensor/utils/sari_hook.py",
],
reference_urls=["https://www.aclweb.org/anthology/Q16-1029.pdf"],
)
def _compute(self, sources, predictions, references):
if not (len(sources) == len(predictions) == len(references)):
raise ValueError("Sources length must match predictions and references lengths.")
sari_score = 0
for src, pred, refs in zip(sources, predictions, references):
sari_score += SARIsent(normalize(src), normalize(pred), [normalize(sent) for sent in refs])
sari_score = sari_score / len(predictions)
return {"sari": 100 * sari_score}
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/mean_iou/README.md
|
# Metric Card for Mean IoU
## Metric Description
IoU (Intersection over Union) is the area of overlap between the predicted segmentation and the ground truth divided by the area of union between the predicted segmentation and the ground truth.
For binary (two classes) or multi-class segmentation, the *mean IoU* of the image is calculated by taking the IoU of each class and averaging them.
## How to Use
The Mean IoU metric takes two numeric arrays as input corresponding to the predicted and ground truth segmentations:
```python
>>> import numpy as np
>>> mean_iou = datasets.load_metric("mean_iou")
>>> predicted = np.array([[2, 2, 3], [8, 2, 4], [3, 255, 2]])
>>> ground_truth = np.array([[1, 2, 2], [8, 2, 1], [3, 255, 1]])
>>> results = mean_iou.compute(predictions=predicted, references=ground_truth, num_labels=10, ignore_index=255)
```
### Inputs
**Mandatory inputs**
- `predictions` (`List[ndarray]`): List of predicted segmentation maps, each of shape (height, width). Each segmentation map can be of a different size.
- `references` (`List[ndarray]`): List of ground truth segmentation maps, each of shape (height, width). Each segmentation map can be of a different size.
- `num_labels` (`int`): Number of classes (categories).
- `ignore_index` (`int`): Index that will be ignored during evaluation.
**Optional inputs**
- `nan_to_num` (`int`): If specified, NaN values will be replaced by the number defined by the user.
- `label_map` (`dict`): If specified, dictionary mapping old label indices to new label indices.
- `reduce_labels` (`bool`): Whether or not to reduce all label values of segmentation maps by 1. Usually used for datasets where 0 is used for background, and background itself is not included in all classes of a dataset (e.g. ADE20k). The background label will be replaced by 255. The default value is `False`.
### Output Values
The metric returns a dictionary with the following elements:
- `mean_iou` (`float`): Mean Intersection-over-Union (IoU averaged over all categories).
- `mean_accuracy` (`float`): Mean accuracy (averaged over all categories).
- `overall_accuracy` (`float`): Overall accuracy on all images.
- `per_category_accuracy` (`ndarray` of shape `(num_labels,)`): Per category accuracy.
- `per_category_iou` (`ndarray` of shape `(num_labels,)`): Per category IoU.
The values of all of the scores reported range from from `0.0` (minimum) and `1.0` (maximum).
Output Example:
```python
{'mean_iou': 0.47750000000000004, 'mean_accuracy': 0.5916666666666666, 'overall_accuracy': 0.5263157894736842, 'per_category_iou': array([0. , 0. , 0.375, 0.4 , 0.5 , 0. , 0.5 , 1. , 1. , 1. ]), 'per_category_accuracy': array([0. , 0. , 0.75 , 0.66666667, 1. , 0. , 0.5 , 1. , 1. , 1. ])}
```
#### Values from Popular Papers
The [leaderboard for the CityScapes dataset](https://paperswithcode.com/sota/semantic-segmentation-on-cityscapes) reports a Mean IOU ranging from 64 to 84; that of [ADE20k](https://paperswithcode.com/sota/semantic-segmentation-on-ade20k) ranges from 30 to a peak of 59.9, indicating that the dataset is more difficult for current approaches (as of 2022).
### Examples
```python
>>> from datasets import load_metric
>>> import numpy as np
>>> mean_iou = load_metric("mean_iou")
>>> # suppose one has 3 different segmentation maps predicted
>>> predicted_1 = np.array([[1, 2], [3, 4], [5, 255]])
>>> actual_1 = np.array([[0, 3], [5, 4], [6, 255]])
>>> predicted_2 = np.array([[2, 7], [9, 2], [3, 6]])
>>> actual_2 = np.array([[1, 7], [9, 2], [3, 6]])
>>> predicted_3 = np.array([[2, 2, 3], [8, 2, 4], [3, 255, 2]])
>>> actual_3 = np.array([[1, 2, 2], [8, 2, 1], [3, 255, 1]])
>>> predictions = [predicted_1, predicted_2, predicted_3]
>>> references = [actual_1, actual_2, actual_3]
>>> results = mean_iou.compute(predictions=predictions, references=references, num_labels=10, ignore_index=255, reduce_labels=False)
>>> print(results) # doctest: +NORMALIZE_WHITESPACE
{'mean_iou': 0.47750000000000004, 'mean_accuracy': 0.5916666666666666, 'overall_accuracy': 0.5263157894736842, 'per_category_iou': array([0. , 0. , 0.375, 0.4 , 0.5 , 0. , 0.5 , 1. , 1. , 1. ]), 'per_category_accuracy': array([0. , 0. , 0.75 , 0.66666667, 1. , 0. , 0.5 , 1. , 1. , 1. ])}
```
## Limitations and Bias
Mean IOU is an average metric, so it will not show you where model predictions differ from the ground truth (i.e. if there are particular regions or classes that the model does poorly on). Further error analysis is needed to gather actional insights that can be used to inform model improvements.
## Citation(s)
```bibtex
@software{MMSegmentation_Contributors_OpenMMLab_Semantic_Segmentation_2020,
author = {{MMSegmentation Contributors}},
license = {Apache-2.0},
month = {7},
title = {{OpenMMLab Semantic Segmentation Toolbox and Benchmark}},
url = {https://github.com/open-mmlab/mmsegmentation},
year = {2020}
}"
```
## Further References
- [Wikipedia article - Jaccard Index](https://en.wikipedia.org/wiki/Jaccard_index)
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/mean_iou/mean_iou.py
|
# Copyright 2022 The HuggingFace Datasets Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""Mean IoU (Intersection-over-Union) metric."""
from typing import Dict, Optional
import numpy as np
import datasets
_DESCRIPTION = """
IoU is the area of overlap between the predicted segmentation and the ground truth divided by the area of union
between the predicted segmentation and the ground truth. For binary (two classes) or multi-class segmentation,
the mean IoU of the image is calculated by taking the IoU of each class and averaging them.
"""
_KWARGS_DESCRIPTION = """
Args:
predictions (`List[ndarray]`):
List of predicted segmentation maps, each of shape (height, width). Each segmentation map can be of a different size.
references (`List[ndarray]`):
List of ground truth segmentation maps, each of shape (height, width). Each segmentation map can be of a different size.
num_labels (`int`):
Number of classes (categories).
ignore_index (`int`):
Index that will be ignored during evaluation.
nan_to_num (`int`, *optional*):
If specified, NaN values will be replaced by the number defined by the user.
label_map (`dict`, *optional*):
If specified, dictionary mapping old label indices to new label indices.
reduce_labels (`bool`, *optional*, defaults to `False`):
Whether or not to reduce all label values of segmentation maps by 1. Usually used for datasets where 0 is used for background,
and background itself is not included in all classes of a dataset (e.g. ADE20k). The background label will be replaced by 255.
Returns:
`Dict[str, float | ndarray]` comprising various elements:
- *mean_iou* (`float`):
Mean Intersection-over-Union (IoU averaged over all categories).
- *mean_accuracy* (`float`):
Mean accuracy (averaged over all categories).
- *overall_accuracy* (`float`):
Overall accuracy on all images.
- *per_category_accuracy* (`ndarray` of shape `(num_labels,)`):
Per category accuracy.
- *per_category_iou* (`ndarray` of shape `(num_labels,)`):
Per category IoU.
Examples:
>>> import numpy as np
>>> mean_iou = datasets.load_metric("mean_iou")
>>> # suppose one has 3 different segmentation maps predicted
>>> predicted_1 = np.array([[1, 2], [3, 4], [5, 255]])
>>> actual_1 = np.array([[0, 3], [5, 4], [6, 255]])
>>> predicted_2 = np.array([[2, 7], [9, 2], [3, 6]])
>>> actual_2 = np.array([[1, 7], [9, 2], [3, 6]])
>>> predicted_3 = np.array([[2, 2, 3], [8, 2, 4], [3, 255, 2]])
>>> actual_3 = np.array([[1, 2, 2], [8, 2, 1], [3, 255, 1]])
>>> predicted = [predicted_1, predicted_2, predicted_3]
>>> ground_truth = [actual_1, actual_2, actual_3]
>>> results = mean_iou.compute(predictions=predicted, references=ground_truth, num_labels=10, ignore_index=255, reduce_labels=False)
>>> print(results) # doctest: +NORMALIZE_WHITESPACE
{'mean_iou': 0.47750000000000004, 'mean_accuracy': 0.5916666666666666, 'overall_accuracy': 0.5263157894736842, 'per_category_iou': array([0. , 0. , 0.375, 0.4 , 0.5 , 0. , 0.5 , 1. , 1. , 1. ]), 'per_category_accuracy': array([0. , 0. , 0.75 , 0.66666667, 1. , 0. , 0.5 , 1. , 1. , 1. ])}
"""
_CITATION = """\
@software{MMSegmentation_Contributors_OpenMMLab_Semantic_Segmentation_2020,
author = {{MMSegmentation Contributors}},
license = {Apache-2.0},
month = {7},
title = {{OpenMMLab Semantic Segmentation Toolbox and Benchmark}},
url = {https://github.com/open-mmlab/mmsegmentation},
year = {2020}
}"""
def intersect_and_union(
pred_label,
label,
num_labels,
ignore_index: bool,
label_map: Optional[Dict[int, int]] = None,
reduce_labels: bool = False,
):
"""Calculate intersection and Union.
Args:
pred_label (`ndarray`):
Prediction segmentation map of shape (height, width).
label (`ndarray`):
Ground truth segmentation map of shape (height, width).
num_labels (`int`):
Number of categories.
ignore_index (`int`):
Index that will be ignored during evaluation.
label_map (`dict`, *optional*):
Mapping old labels to new labels. The parameter will work only when label is str.
reduce_labels (`bool`, *optional*, defaults to `False`):
Whether or not to reduce all label values of segmentation maps by 1. Usually used for datasets where 0 is used for background,
and background itself is not included in all classes of a dataset (e.g. ADE20k). The background label will be replaced by 255.
Returns:
area_intersect (`ndarray`):
The intersection of prediction and ground truth histogram on all classes.
area_union (`ndarray`):
The union of prediction and ground truth histogram on all classes.
area_pred_label (`ndarray`):
The prediction histogram on all classes.
area_label (`ndarray`):
The ground truth histogram on all classes.
"""
if label_map is not None:
for old_id, new_id in label_map.items():
label[label == old_id] = new_id
# turn into Numpy arrays
pred_label = np.array(pred_label)
label = np.array(label)
if reduce_labels:
label[label == 0] = 255
label = label - 1
label[label == 254] = 255
mask = label != ignore_index
mask = np.not_equal(label, ignore_index)
pred_label = pred_label[mask]
label = np.array(label)[mask]
intersect = pred_label[pred_label == label]
area_intersect = np.histogram(intersect, bins=num_labels, range=(0, num_labels - 1))[0]
area_pred_label = np.histogram(pred_label, bins=num_labels, range=(0, num_labels - 1))[0]
area_label = np.histogram(label, bins=num_labels, range=(0, num_labels - 1))[0]
area_union = area_pred_label + area_label - area_intersect
return area_intersect, area_union, area_pred_label, area_label
def total_intersect_and_union(
results,
gt_seg_maps,
num_labels,
ignore_index: bool,
label_map: Optional[Dict[int, int]] = None,
reduce_labels: bool = False,
):
"""Calculate Total Intersection and Union, by calculating `intersect_and_union` for each (predicted, ground truth) pair.
Args:
results (`ndarray`):
List of prediction segmentation maps, each of shape (height, width).
gt_seg_maps (`ndarray`):
List of ground truth segmentation maps, each of shape (height, width).
num_labels (`int`):
Number of categories.
ignore_index (`int`):
Index that will be ignored during evaluation.
label_map (`dict`, *optional*):
Mapping old labels to new labels. The parameter will work only when label is str.
reduce_labels (`bool`, *optional*, defaults to `False`):
Whether or not to reduce all label values of segmentation maps by 1. Usually used for datasets where 0 is used for background,
and background itself is not included in all classes of a dataset (e.g. ADE20k). The background label will be replaced by 255.
Returns:
total_area_intersect (`ndarray`):
The intersection of prediction and ground truth histogram on all classes.
total_area_union (`ndarray`):
The union of prediction and ground truth histogram on all classes.
total_area_pred_label (`ndarray`):
The prediction histogram on all classes.
total_area_label (`ndarray`):
The ground truth histogram on all classes.
"""
total_area_intersect = np.zeros((num_labels,), dtype=np.float64)
total_area_union = np.zeros((num_labels,), dtype=np.float64)
total_area_pred_label = np.zeros((num_labels,), dtype=np.float64)
total_area_label = np.zeros((num_labels,), dtype=np.float64)
for result, gt_seg_map in zip(results, gt_seg_maps):
area_intersect, area_union, area_pred_label, area_label = intersect_and_union(
result, gt_seg_map, num_labels, ignore_index, label_map, reduce_labels
)
total_area_intersect += area_intersect
total_area_union += area_union
total_area_pred_label += area_pred_label
total_area_label += area_label
return total_area_intersect, total_area_union, total_area_pred_label, total_area_label
def mean_iou(
results,
gt_seg_maps,
num_labels,
ignore_index: bool,
nan_to_num: Optional[int] = None,
label_map: Optional[Dict[int, int]] = None,
reduce_labels: bool = False,
):
"""Calculate Mean Intersection and Union (mIoU).
Args:
results (`ndarray`):
List of prediction segmentation maps, each of shape (height, width).
gt_seg_maps (`ndarray`):
List of ground truth segmentation maps, each of shape (height, width).
num_labels (`int`):
Number of categories.
ignore_index (`int`):
Index that will be ignored during evaluation.
nan_to_num (`int`, *optional*):
If specified, NaN values will be replaced by the number defined by the user.
label_map (`dict`, *optional*):
Mapping old labels to new labels. The parameter will work only when label is str.
reduce_labels (`bool`, *optional*, defaults to `False`):
Whether or not to reduce all label values of segmentation maps by 1. Usually used for datasets where 0 is used for background,
and background itself is not included in all classes of a dataset (e.g. ADE20k). The background label will be replaced by 255.
Returns:
`Dict[str, float | ndarray]` comprising various elements:
- *mean_iou* (`float`):
Mean Intersection-over-Union (IoU averaged over all categories).
- *mean_accuracy* (`float`):
Mean accuracy (averaged over all categories).
- *overall_accuracy* (`float`):
Overall accuracy on all images.
- *per_category_accuracy* (`ndarray` of shape `(num_labels,)`):
Per category accuracy.
- *per_category_iou* (`ndarray` of shape `(num_labels,)`):
Per category IoU.
"""
total_area_intersect, total_area_union, total_area_pred_label, total_area_label = total_intersect_and_union(
results, gt_seg_maps, num_labels, ignore_index, label_map, reduce_labels
)
# compute metrics
metrics = {}
all_acc = total_area_intersect.sum() / total_area_label.sum()
iou = total_area_intersect / total_area_union
acc = total_area_intersect / total_area_label
metrics["mean_iou"] = np.nanmean(iou)
metrics["mean_accuracy"] = np.nanmean(acc)
metrics["overall_accuracy"] = all_acc
metrics["per_category_iou"] = iou
metrics["per_category_accuracy"] = acc
if nan_to_num is not None:
metrics = {metric: np.nan_to_num(metric_value, nan=nan_to_num) for metric, metric_value in metrics.items()}
return metrics
@datasets.utils.file_utils.add_start_docstrings(_DESCRIPTION, _KWARGS_DESCRIPTION)
class MeanIoU(datasets.Metric):
def _info(self):
return datasets.MetricInfo(
description=_DESCRIPTION,
citation=_CITATION,
inputs_description=_KWARGS_DESCRIPTION,
features=datasets.Features(
# 1st Seq - height dim, 2nd - width dim
{
"predictions": datasets.Sequence(datasets.Sequence(datasets.Value("uint16"))),
"references": datasets.Sequence(datasets.Sequence(datasets.Value("uint16"))),
}
),
reference_urls=[
"https://github.com/open-mmlab/mmsegmentation/blob/71c201b1813267d78764f306a297ca717827c4bf/mmseg/core/evaluation/metrics.py"
],
)
def _compute(
self,
predictions,
references,
num_labels: int,
ignore_index: bool,
nan_to_num: Optional[int] = None,
label_map: Optional[Dict[int, int]] = None,
reduce_labels: bool = False,
):
iou_result = mean_iou(
results=predictions,
gt_seg_maps=references,
num_labels=num_labels,
ignore_index=ignore_index,
nan_to_num=nan_to_num,
label_map=label_map,
reduce_labels=reduce_labels,
)
return iou_result
| 0
|
hf_public_repos/datasets/metrics
|
hf_public_repos/datasets/metrics/comet/README.md
|
# Metric Card for COMET
## Metric description
Crosslingual Optimized Metric for Evaluation of Translation (COMET) is an open-source framework used to train Machine Translation metrics that achieve high levels of correlation with different types of human judgments.
## How to use
COMET takes 3 lists of strings as input: `sources` (a list of source sentences), `predictions` (a list of candidate translations) and `references` (a list of reference translations).
```python
from datasets import load_metric
comet_metric = load_metric('comet')
source = ["Dem Feuer konnte Einhalt geboten werden", "Schulen und Kindergärten wurden eröffnet."]
hypothesis = ["The fire could be stopped", "Schools and kindergartens were open"]
reference = ["They were able to control the fire.", "Schools and kindergartens opened"]
comet_score = comet_metric.compute(predictions=hypothesis, references=reference, sources=source)
```
It has several configurations, named after the COMET model to be used. It will default to `wmt20-comet-da` (previously known as `wmt-large-da-estimator-1719`). Alternate models that can be chosen include `wmt20-comet-qe-da`, `wmt21-comet-mqm`, `wmt21-cometinho-da`, `wmt21-comet-qe-mqm` and `emnlp20-comet-rank`.
It also has several optional arguments:
`gpus`: optional, an integer (number of GPUs to train on) or a list of integers (which GPUs to train on). Set to 0 to use CPU. The default value is `None` (uses one GPU if possible, else use CPU).
`progress_bar`a boolean -- if set to `True`, progress updates will be printed out. The default value is `False`.
More information about model characteristics can be found on the [COMET website](https://unbabel.github.io/COMET/html/models.html).
## Output values
The COMET metric outputs two lists:
`scores`: a list of COMET scores for each of the input sentences, ranging from 0-1.
`mean_score`: the mean value of COMET scores `scores` over all the input sentences, ranging from 0-1.
### Values from popular papers
The [original COMET paper](https://arxiv.org/pdf/2009.09025.pdf) reported average COMET scores ranging from 0.4 to 0.6, depending on the language pairs used for evaluating translation models. They also illustrate that COMET correlates well with human judgements compared to other metrics such as [BLEU](https://huggingface.co/metrics/bleu) and [CHRF](https://huggingface.co/metrics/chrf).
## Examples
Full match:
```python
from datasets import load_metric
comet_metric = load_metric('comet')
source = ["Dem Feuer konnte Einhalt geboten werden", "Schulen und Kindergärten wurden eröffnet."]
hypothesis = ["They were able to control the fire.", "Schools and kindergartens opened"]
reference = ["They were able to control the fire.", "Schools and kindergartens opened"]
results = comet_metric.compute(predictions=hypothesis, references=reference, sources=source)
print([round(v, 1) for v in results["scores"]])
[1.0, 1.0]
```
Partial match:
```python
from datasets import load_metric
comet_metric = load_metric('comet')
source = ["Dem Feuer konnte Einhalt geboten werden", "Schulen und Kindergärten wurden eröffnet."]
hypothesis = ["The fire could be stopped", "Schools and kindergartens were open"]
reference = ["They were able to control the fire", "Schools and kindergartens opened"]
results = comet_metric.compute(predictions=hypothesis, references=reference, sources=source)
print([round(v, 2) for v in results["scores"]])
[0.19, 0.92]
```
No match:
```python
from datasets import load_metric
comet_metric = load_metric('comet')
source = ["Dem Feuer konnte Einhalt geboten werden", "Schulen und Kindergärten wurden eröffnet."]
hypothesis = ["The girl went for a walk", "The boy was sleeping"]
reference = ["They were able to control the fire", "Schools and kindergartens opened"]
results = comet_metric.compute(predictions=hypothesis, references=reference, sources=source)
print([round(v, 2) for v in results["scores"]])
[0.00, 0.00]
```
## Limitations and bias
The models provided for calculating the COMET metric are built on top of XLM-R and cover the following languages:
Afrikaans, Albanian, Amharic, Arabic, Armenian, Assamese, Azerbaijani, Basque, Belarusian, Bengali, Bengali Romanized, Bosnian, Breton, Bulgarian, Burmese, Burmese, Catalan, Chinese (Simplified), Chinese (Traditional), Croatian, Czech, Danish, Dutch, English, Esperanto, Estonian, Filipino, Finnish, French, Galician, Georgian, German, Greek, Gujarati, Hausa, Hebrew, Hindi, Hindi Romanized, Hungarian, Icelandic, Indonesian, Irish, Italian, Japanese, Javanese, Kannada, Kazakh, Khmer, Korean, Kurdish (Kurmanji), Kyrgyz, Lao, Latin, Latvian, Lithuanian, Macedonian, Malagasy, Malay, Malayalam, Marathi, Mongolian, Nepali, Norwegian, Oriya, Oromo, Pashto, Persian, Polish, Portuguese, Punjabi, Romanian, Russian, Sanskri, Scottish, Gaelic, Serbian, Sindhi, Sinhala, Slovak, Slovenian, Somali, Spanish, Sundanese, Swahili, Swedish, Tamil, Tamil Romanized, Telugu, Telugu Romanized, Thai, Turkish, Ukrainian, Urdu, Urdu Romanized, Uyghur, Uzbek, Vietnamese, Welsh, Western, Frisian, Xhosa, Yiddish.
Thus, results for language pairs containing uncovered languages are unreliable, as per the [COMET website](https://github.com/Unbabel/COMET)
Also, calculating the COMET metric involves downloading the model from which features are obtained -- the default model, `wmt20-comet-da`, takes over 1.79GB of storage space and downloading it can take a significant amount of time depending on the speed of your internet connection. If this is an issue, choose a smaller model; for instance `wmt21-cometinho-da` is 344MB.
## Citation
```bibtex
@inproceedings{rei-EtAl:2020:WMT,
author = {Rei, Ricardo and Stewart, Craig and Farinha, Ana C and Lavie, Alon},
title = {Unbabel's Participation in the WMT20 Metrics Shared Task},
booktitle = {Proceedings of the Fifth Conference on Machine Translation},
month = {November},
year = {2020},
address = {Online},
publisher = {Association for Computational Linguistics},
pages = {909--918},
}
```
```bibtex
@inproceedings{rei-etal-2020-comet,
title = "{COMET}: A Neural Framework for {MT} Evaluation",
author = "Rei, Ricardo and
Stewart, Craig and
Farinha, Ana C and
Lavie, Alon",
booktitle = "Proceedings of the 2020 Conference on Empirical Methods in Natural Language Processing (EMNLP)",
month = nov,
year = "2020",
address = "Online",
publisher = "Association for Computational Linguistics",
url = "https://www.aclweb.org/anthology/2020.emnlp-main.213",
pages = "2685--2702",
```
## Further References
- [COMET website](https://unbabel.github.io/COMET/html/index.html)
- [Hugging Face Tasks - Machine Translation](https://huggingface.co/tasks/translation)
| 0
|
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.