The full dataset viewer is not available (click to read why). Only showing a preview of the rows.
Error code: DatasetGenerationCastError
Exception: DatasetGenerationCastError
Message: An error occurred while generating the dataset
All the data files must have the same columns, but at some point there are 1074 new columns ({'h2a', 'znf2', 'tbp', 'ssrp1', 'ercc2', 'macroh2a2', 'smc3', 'irf1', 'hsf1', 'nfrkb', 'znf486', 'znf784', 'h2afx', 'nfatc3', 'nfic', 'znf341', 'atrx', 'hmg20a', 'znf222', 'cdca2', 'nipbl', 'prpf4', 'zfp69b', 'pdx1', 'znf217', 'znf280d', 'nkx2-5', 'rfx5', 'lmo1', 'rbck1', 'znf567', 'hoxa4', 'znf697', 'ybx1', 'grhl3', 'atf5', 'nfkb2', 'lmx1b', 'ttf1', 'creb3', 'fosl2', 'h3k36ac', 'hnf1b', 'stat5a', 'cebpb', 'znf549', 'csde1', 'chd7', 'glis3', 'zbtb24', 'znf343', 'dlx1', 'zbtb7a', 'znf467', 'six5', 'aebp2', 'elf3', 'xrcc4', 'hdac3', 'znf362', 'znf282', 'zbtb42', 'h3r26cit', 'stat5b', 'atf4', 'spdef', 'znf565', 'runx2', 'six2', 'stat5', 'erg', 'znf320', 'fxr1', 'alkbh3', 'creb3l4', 'znf33a', 'atf3', 'auts2', 'tfcp2', 'znf561', 'e2f4', 'ddx20', 'greb1', 'znf175', 'spib', 'smad2/3', 'egln2', 'ezh1', 'lrwd1', 'sap30', 'rxrg', 'gli4', 'zbtb8a', 'h3.3,h2a.z', 'scrt1', 'zbtb40', 'znf860', 'cnot3', 'glyr1', 'grhl2', 'rag2', 'zc3h8', 'h2bk120ub', 'tle3', 'smc1a', 'nfe2l2', 'camta2', 'nr1h2', 'znf165', 'hey1', 'tcf7l1', 'egr1', 'egr3', 'tead2', 'znf317', 'hoxa9', 'myb', 'brd3', 'smarca4', 'ehmt2', 'hdac8', 'znf440', 'dnase', 'mafb', 'mbd1', 'irf2', 'sox15', 'sirt6', 'znf141', 'ahrr', 'rexo1', 'klf16', 'tshz1', 'nfia', 'tfap2c', 'apobec3b', 'grip1', 'nr5a2', 'supt5h', 'bcor', 'stag1', 'suz12', 'zim3', 'hotair', 'hic1', 'lana', 'hnf4a', 'clock', 'tfam', 'hes2', 'id3', 'wrnip1', 'cdk9', 'znf281', 'znf250', 'hltf', 'foxq1', 'sox4', 'nfil3', 'dnmt1', 'rloop', 'h3k9k14ac', 'gucy1b3', 'tbx3', '
...
'h2bk5ac', 'znf808', 'foxa1/2', 'znf543', 'brd9', 'pax6', 'erf', 'ncapg2', 'znf664', 'epcam', 'ccdc101', 'h4ac', 'zbtb16', 'chd4', 'prdm6', 'phf2', 'gata6', 'znf468', 'zc3h11a', 'jmjd6', 'tbx21', 'kdm4b', 'med1', 'znf519', 'zfp69', 'nfya', 'znf512', 'arid3a', 'zbed1', 'drap1', 'smarce1', 'vdr', 'lmo2', 'rxra/b/g', 'eed', 'sp7', 'dpf1', 'znf547', 'ctbp2', 'rel', 'znf257', 'bahd1', 'h4', 'zxdb', 'taf7', 'ptpn11', 'h3f3a', 'nr1h3', 'h3-t45ph', 'psip1', 'znf121', 'mbd3', 'arntl2', 'usf2', 'pml', 'h2ax', 'h2a.zac', 'h2ak119ub', 'h3k122ac', 'tcf12', 'cebpg', 'rfx1', 'h2bk15ac', 'znf800', 'znf586', 'wnt3a', 'polr3g', 'znf266', 'krab', 'patz1', 'crem', 'cbx3', 'e2f3', 'h2bk12ac', 'h3k23ac', 'trrap', 'elk4', 'chd8', 'polr2b', 'atf1', 'foxp1', 'znf273', 'prmt1', 'faire', 'fos', 'srebf2', 'gata2', 'znf680', 'kdm3b', 'klf3', 'brd4', 'hnf1a', 'histone lysine acetylation', 'nr2f6', 'pygo2', 'relb', 'bmi1', 'tbx5', 'znf304', 'sumo2/3', 'klf12', 'mafg', 'nanog', 'pcgf1', 'fli1', 'h2az', 'h3k4me1', 'nfe2', 'wwtr1', 'ovol3', 'irf4', 'prdm2', 'ikzf1', 'id2', 'mnt', 'znf488', 'ercc6', 'mybl2', 'zfp42', 'znf589', 'hdac4', 'junb', 'znf93', 'zeb1', 'znf169', 'zbed4', 'znf391', 'h3k36me2', 'znf30', 'creb3l2', 'ar', 'zbtb48', 'h3k36me1', 'znf518a', 'hes4', 'klf8', 'rad51', 'hbp1', 'h3k79me2', 'phip', 'zscan16', 'ing5', 'larp7', 'macroh2a1', 'rpa2', 'carm1', 'rnf2', 'ccnt2', 'prdm12', 'znf17', 'fev', 'znf667', 'sp4', 'tsc22d4', 'hcfc1r1', 'h4k20me1', 'nfyc', 'cbx8', 'drosha', 'epc1', 'ing2', 'znf623'}) and 11 missing columns ({'end', 'gene_name', 'start', 'gene_biotype', 'build_region_index', 'strand', 'tss', 'loc2', 'gene_id', 'loc1', 'chrom'}).
This happened while the csv dataset builder was generating data using
hf://datasets/TongjiZhanglab/chrombert/hg38_6k_mask_matrix.tsv (at revision 721c67aecb0789c4eb204bc304f21779de7cd201), ['hf://datasets/TongjiZhanglab/chrombert@721c67aecb0789c4eb204bc304f21779de7cd201/hg38_1kb_gene_meta.tsv', 'hf://datasets/TongjiZhanglab/chrombert@721c67aecb0789c4eb204bc304f21779de7cd201/hg38_200bp_gene_meta.tsv', 'hf://datasets/TongjiZhanglab/chrombert@721c67aecb0789c4eb204bc304f21779de7cd201/hg38_2kb_gene_meta.tsv', 'hf://datasets/TongjiZhanglab/chrombert@721c67aecb0789c4eb204bc304f21779de7cd201/hg38_4kb_gene_meta.tsv', 'hf://datasets/TongjiZhanglab/chrombert@721c67aecb0789c4eb204bc304f21779de7cd201/hg38_6k_mask_matrix.tsv', 'hf://datasets/TongjiZhanglab/chrombert@721c67aecb0789c4eb204bc304f21779de7cd201/hg38_6k_meta.tsv', 'hf://datasets/TongjiZhanglab/chrombert@721c67aecb0789c4eb204bc304f21779de7cd201/mm10_1kb_gene_meta.tsv', 'hf://datasets/TongjiZhanglab/chrombert@721c67aecb0789c4eb204bc304f21779de7cd201/mm10_5k_mask_matrix.tsv', 'hf://datasets/TongjiZhanglab/chrombert@721c67aecb0789c4eb204bc304f21779de7cd201/mm10_5k_meta.tsv']
Please either edit the data files to have matching columns, or separate them into different configurations (see docs at https://hf.co/docs/hub/datasets-manual-configuration#multiple-configurations)
Traceback: Traceback (most recent call last):
File "/usr/local/lib/python3.12/site-packages/datasets/builder.py", line 1887, in _prepare_split_single
writer.write_table(table)
File "/usr/local/lib/python3.12/site-packages/datasets/arrow_writer.py", line 675, in write_table
pa_table = table_cast(pa_table, self._schema)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/usr/local/lib/python3.12/site-packages/datasets/table.py", line 2272, in table_cast
return cast_table_to_schema(table, schema)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/usr/local/lib/python3.12/site-packages/datasets/table.py", line 2218, in cast_table_to_schema
raise CastError(
datasets.table.CastError: Couldn't cast
Unnamed: 0: string
5hmc: int64
adnp: int64
aebp2: int64
aff1: int64
aff4: int64
ago1: int64
ago2: int64
ahr: int64
ahrr: int64
alkbh3: int64
anp32e: int64
apobec3b: int64
ar: int64
arid1a: int64
arid2: int64
arid3a: int64
arid5b: int64
arnt: int64
arnt2: int64
arntl: int64
arntl2: int64
arrb1: int64
ascl1: int64
ascl2: int64
ash1l: int64
ash2l: int64
asxl1: int64
atac-seq: int64
atf1: int64
atf2: int64
atf3: int64
atf4: int64
atf5: int64
atoh1: int64
atrx: int64
auts2: int64
bach1: int64
bach2: int64
bahd1: int64
banf1: int64
banp: int64
barhl1: int64
barx1: int64
barx2: int64
batf: int64
bcat1: int64
bcl11a: int64
bcl11b: int64
bcl3: int64
bcl6: int64
bclaf1: int64
bcor: int64
bdp1: int64
bhlhe40: int64
bmi1: int64
bptf: int64
brca1: int64
brd1: int64
brd1-s: int64
brd2: int64
brd3: int64
brd4: int64
brd4-nut: int64
brd7: int64
brd9: int64
brdu: int64
brf1: int64
brf2: int64
brpf3: int64
c17orf49: int64
c17orf96: int64
camta2: int64
carm1: int64
casp8ap2: int64
cbfa2t2: int64
cbfb: int64
cbx1: int64
cbx2: int64
cbx3: int64
cbx4: int64
cbx5: int64
cbx6: int64
cbx7: int64
cbx8: int64
ccdc101: int64
ccnd2: int64
ccnt2: int64
cd74: int64
cdc5l: int64
cdca2: int64
cdk12: int64
cdk6: int64
cdk7: int64
cdk8: int64
cdk9: int64
cdx1: int64
cdx2: int64
cebpa: int64
cebpb: int64
cebpd: int64
cebpg: int64
cebpz: int64
cenpa: int64
cenpc: int64
cfp1: int64
champ1: int64
chd1: int64
chd2: int64
chd4: int64
chd5: int64
chd7: int64
chd8: int64
ciita: int64
clock: int64
cnot3: int64
cops2: i
...
4
znf501: int64
znf506: int64
znf512: int64
znf518a: int64
znf519: int64
znf528: int64
znf534: int64
znf543: int64
znf547: int64
znf549: int64
znf554: int64
znf558: int64
znf561: int64
znf563: int64
znf565: int64
znf567: int64
znf571: int64
znf573: int64
znf574: int64
znf580: int64
znf581: int64
znf584: int64
znf585a: int64
znf585b: int64
znf586: int64
znf589: int64
znf592: int64
znf596: int64
znf600: int64
znf605: int64
znf610: int64
znf611: int64
znf616: int64
znf623: int64
znf627: int64
znf639: int64
znf641: int64
znf644: int64
znf649: int64
znf654: int64
znf664: int64
znf667: int64
znf671: int64
znf674: int64
znf675: int64
znf680: int64
znf682: int64
znf687: int64
znf692: int64
znf695: int64
znf697: int64
znf701: int64
znf706: int64
znf707: int64
znf71: int64
znf711: int64
znf714: int64
znf730: int64
znf740: int64
znf750: int64
znf75a: int64
znf75d: int64
znf76: int64
znf765: int64
znf766: int64
znf770: int64
znf778: int64
znf780a: int64
znf781: int64
znf783: int64
znf784: int64
znf786: int64
znf788: int64
znf792: int64
znf793: int64
znf8: int64
znf800: int64
znf808: int64
znf816: int64
znf823: int64
znf83: int64
znf837: int64
znf84: int64
znf846: int64
znf85: int64
znf860: int64
znf92: int64
znf93: int64
zscan16: int64
zscan2: int64
zscan20: int64
zscan22: int64
zscan23: int64
zscan29: int64
zscan31: int64
zscan5a: int64
zta: int64
zxdb: int64
zxdc: int64
zzz3: int64
-- schema metadata --
pandas: '{"index_columns": [{"kind": "range", "name": null, "start": 0, "' + 115721
to
{'chrom': Value('string'), 'loc1': Value('int64'), 'loc2': Value('int64'), 'strand': Value('string'), 'tss': Value('int64'), 'gene_id': Value('string'), 'gene_name': Value('string'), 'gene_biotype': Value('string'), 'start': Value('int64'), 'end': Value('int64'), 'build_region_index': Value('int64')}
because column names don't match
During handling of the above exception, another exception occurred:
Traceback (most recent call last):
File "/src/services/worker/src/worker/job_runners/config/parquet_and_info.py", line 1342, in compute_config_parquet_and_info_response
parquet_operations, partial, estimated_dataset_info = stream_convert_to_parquet(
^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/src/services/worker/src/worker/job_runners/config/parquet_and_info.py", line 907, in stream_convert_to_parquet
builder._prepare_split(split_generator=splits_generators[split], file_format="parquet")
File "/usr/local/lib/python3.12/site-packages/datasets/builder.py", line 1736, in _prepare_split
for job_id, done, content in self._prepare_split_single(
^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/usr/local/lib/python3.12/site-packages/datasets/builder.py", line 1889, in _prepare_split_single
raise DatasetGenerationCastError.from_cast_error(
datasets.exceptions.DatasetGenerationCastError: An error occurred while generating the dataset
All the data files must have the same columns, but at some point there are 1074 new columns ({'h2a', 'znf2', 'tbp', 'ssrp1', 'ercc2', 'macroh2a2', 'smc3', 'irf1', 'hsf1', 'nfrkb', 'znf486', 'znf784', 'h2afx', 'nfatc3', 'nfic', 'znf341', 'atrx', 'hmg20a', 'znf222', 'cdca2', 'nipbl', 'prpf4', 'zfp69b', 'pdx1', 'znf217', 'znf280d', 'nkx2-5', 'rfx5', 'lmo1', 'rbck1', 'znf567', 'hoxa4', 'znf697', 'ybx1', 'grhl3', 'atf5', 'nfkb2', 'lmx1b', 'ttf1', 'creb3', 'fosl2', 'h3k36ac', 'hnf1b', 'stat5a', 'cebpb', 'znf549', 'csde1', 'chd7', 'glis3', 'zbtb24', 'znf343', 'dlx1', 'zbtb7a', 'znf467', 'six5', 'aebp2', 'elf3', 'xrcc4', 'hdac3', 'znf362', 'znf282', 'zbtb42', 'h3r26cit', 'stat5b', 'atf4', 'spdef', 'znf565', 'runx2', 'six2', 'stat5', 'erg', 'znf320', 'fxr1', 'alkbh3', 'creb3l4', 'znf33a', 'atf3', 'auts2', 'tfcp2', 'znf561', 'e2f4', 'ddx20', 'greb1', 'znf175', 'spib', 'smad2/3', 'egln2', 'ezh1', 'lrwd1', 'sap30', 'rxrg', 'gli4', 'zbtb8a', 'h3.3,h2a.z', 'scrt1', 'zbtb40', 'znf860', 'cnot3', 'glyr1', 'grhl2', 'rag2', 'zc3h8', 'h2bk120ub', 'tle3', 'smc1a', 'nfe2l2', 'camta2', 'nr1h2', 'znf165', 'hey1', 'tcf7l1', 'egr1', 'egr3', 'tead2', 'znf317', 'hoxa9', 'myb', 'brd3', 'smarca4', 'ehmt2', 'hdac8', 'znf440', 'dnase', 'mafb', 'mbd1', 'irf2', 'sox15', 'sirt6', 'znf141', 'ahrr', 'rexo1', 'klf16', 'tshz1', 'nfia', 'tfap2c', 'apobec3b', 'grip1', 'nr5a2', 'supt5h', 'bcor', 'stag1', 'suz12', 'zim3', 'hotair', 'hic1', 'lana', 'hnf4a', 'clock', 'tfam', 'hes2', 'id3', 'wrnip1', 'cdk9', 'znf281', 'znf250', 'hltf', 'foxq1', 'sox4', 'nfil3', 'dnmt1', 'rloop', 'h3k9k14ac', 'gucy1b3', 'tbx3', '
...
'h2bk5ac', 'znf808', 'foxa1/2', 'znf543', 'brd9', 'pax6', 'erf', 'ncapg2', 'znf664', 'epcam', 'ccdc101', 'h4ac', 'zbtb16', 'chd4', 'prdm6', 'phf2', 'gata6', 'znf468', 'zc3h11a', 'jmjd6', 'tbx21', 'kdm4b', 'med1', 'znf519', 'zfp69', 'nfya', 'znf512', 'arid3a', 'zbed1', 'drap1', 'smarce1', 'vdr', 'lmo2', 'rxra/b/g', 'eed', 'sp7', 'dpf1', 'znf547', 'ctbp2', 'rel', 'znf257', 'bahd1', 'h4', 'zxdb', 'taf7', 'ptpn11', 'h3f3a', 'nr1h3', 'h3-t45ph', 'psip1', 'znf121', 'mbd3', 'arntl2', 'usf2', 'pml', 'h2ax', 'h2a.zac', 'h2ak119ub', 'h3k122ac', 'tcf12', 'cebpg', 'rfx1', 'h2bk15ac', 'znf800', 'znf586', 'wnt3a', 'polr3g', 'znf266', 'krab', 'patz1', 'crem', 'cbx3', 'e2f3', 'h2bk12ac', 'h3k23ac', 'trrap', 'elk4', 'chd8', 'polr2b', 'atf1', 'foxp1', 'znf273', 'prmt1', 'faire', 'fos', 'srebf2', 'gata2', 'znf680', 'kdm3b', 'klf3', 'brd4', 'hnf1a', 'histone lysine acetylation', 'nr2f6', 'pygo2', 'relb', 'bmi1', 'tbx5', 'znf304', 'sumo2/3', 'klf12', 'mafg', 'nanog', 'pcgf1', 'fli1', 'h2az', 'h3k4me1', 'nfe2', 'wwtr1', 'ovol3', 'irf4', 'prdm2', 'ikzf1', 'id2', 'mnt', 'znf488', 'ercc6', 'mybl2', 'zfp42', 'znf589', 'hdac4', 'junb', 'znf93', 'zeb1', 'znf169', 'zbed4', 'znf391', 'h3k36me2', 'znf30', 'creb3l2', 'ar', 'zbtb48', 'h3k36me1', 'znf518a', 'hes4', 'klf8', 'rad51', 'hbp1', 'h3k79me2', 'phip', 'zscan16', 'ing5', 'larp7', 'macroh2a1', 'rpa2', 'carm1', 'rnf2', 'ccnt2', 'prdm12', 'znf17', 'fev', 'znf667', 'sp4', 'tsc22d4', 'hcfc1r1', 'h4k20me1', 'nfyc', 'cbx8', 'drosha', 'epc1', 'ing2', 'znf623'}) and 11 missing columns ({'end', 'gene_name', 'start', 'gene_biotype', 'build_region_index', 'strand', 'tss', 'loc2', 'gene_id', 'loc1', 'chrom'}).
This happened while the csv dataset builder was generating data using
hf://datasets/TongjiZhanglab/chrombert/hg38_6k_mask_matrix.tsv (at revision 721c67aecb0789c4eb204bc304f21779de7cd201), ['hf://datasets/TongjiZhanglab/chrombert@721c67aecb0789c4eb204bc304f21779de7cd201/hg38_1kb_gene_meta.tsv', 'hf://datasets/TongjiZhanglab/chrombert@721c67aecb0789c4eb204bc304f21779de7cd201/hg38_200bp_gene_meta.tsv', 'hf://datasets/TongjiZhanglab/chrombert@721c67aecb0789c4eb204bc304f21779de7cd201/hg38_2kb_gene_meta.tsv', 'hf://datasets/TongjiZhanglab/chrombert@721c67aecb0789c4eb204bc304f21779de7cd201/hg38_4kb_gene_meta.tsv', 'hf://datasets/TongjiZhanglab/chrombert@721c67aecb0789c4eb204bc304f21779de7cd201/hg38_6k_mask_matrix.tsv', 'hf://datasets/TongjiZhanglab/chrombert@721c67aecb0789c4eb204bc304f21779de7cd201/hg38_6k_meta.tsv', 'hf://datasets/TongjiZhanglab/chrombert@721c67aecb0789c4eb204bc304f21779de7cd201/mm10_1kb_gene_meta.tsv', 'hf://datasets/TongjiZhanglab/chrombert@721c67aecb0789c4eb204bc304f21779de7cd201/mm10_5k_mask_matrix.tsv', 'hf://datasets/TongjiZhanglab/chrombert@721c67aecb0789c4eb204bc304f21779de7cd201/mm10_5k_meta.tsv']
Please either edit the data files to have matching columns, or separate them into different configurations (see docs at https://hf.co/docs/hub/datasets-manual-configuration#multiple-configurations)Need help to make the dataset viewer work? Make sure to review how to configure the dataset viewer, and open a discussion for direct support.
chrom string | loc1 int64 | loc2 int64 | strand string | tss int64 | gene_id string | gene_name string | gene_biotype string | start int64 | end int64 | build_region_index int64 |
|---|---|---|---|---|---|---|---|---|---|---|
chr1 | 182,696 | 184,174 | + | 182,696 | ENSG00000279928 | DDX11L17 | unprocessed_pseudogene | 182,000 | 183,000 | 40 |
chr1 | 2,581,560 | 2,584,533 | + | 2,581,560 | ENSG00000228037 | null | lncRNA | 2,581,000 | 2,582,000 | 1,762 |
chr1 | 3,069,168 | 3,438,621 | + | 3,069,168 | ENSG00000142611 | PRDM16 | protein_coding | 3,069,000 | 3,070,000 | 2,125 |
chr1 | 5,301,928 | 5,307,394 | - | 5,307,394 | ENSG00000284616 | null | lncRNA | 5,307,000 | 5,308,000 | 3,910 |
chr1 | 2,403,964 | 2,413,797 | - | 2,413,797 | ENSG00000157911 | PEX10 | protein_coding | 2,413,000 | 2,414,000 | 1,600 |
chr1 | 2,350,414 | 2,352,820 | - | 2,352,820 | ENSG00000269896 | null | transcribed_processed_pseudogene | 2,352,000 | 2,353,000 | 1,545 |
chr1 | 3,944,547 | 3,949,024 | - | 3,949,024 | ENSG00000226374 | LINC01345 | lncRNA | 3,949,000 | 3,950,000 | 2,933 |
chr1 | 4,175,528 | 4,175,899 | - | 4,175,899 | ENSG00000229280 | EEF1DP6 | processed_pseudogene | 4,175,000 | 4,176,000 | 3,112 |
chr1 | 10,472,288 | 10,630,758 | + | 10,472,288 | ENSG00000142655 | PEX14 | protein_coding | 10,472,000 | 10,473,000 | 8,554 |
chr1 | 4,571,481 | 4,594,016 | + | 4,571,481 | ENSG00000232596 | LINC01646 | lncRNA | 4,571,000 | 4,572,000 | 3,400 |
chr1 | 4,412,027 | 4,424,689 | + | 4,412,027 | ENSG00000235054 | LINC01777 | lncRNA | 4,412,000 | 4,413,000 | 3,284 |
chr1 | 5,086,459 | 5,090,899 | - | 5,090,899 | ENSG00000231510 | LINC02782 | lncRNA | 5,090,000 | 5,091,000 | 3,757 |
chr1 | 2,425,980 | 2,505,532 | + | 2,425,980 | ENSG00000149527 | PLCH2 | protein_coding | 2,425,000 | 2,426,000 | 1,611 |
chr1 | 4,963,954 | 4,973,298 | - | 4,973,298 | ENSG00000284739 | null | lncRNA | 4,973,000 | 4,974,000 | 3,685 |
chr1 | 9,292,894 | 9,369,532 | + | 9,292,894 | ENSG00000171621 | SPSB1 | protein_coding | 9,292,000 | 9,293,000 | 7,491 |
chr1 | 3,306,636 | 3,310,096 | - | 3,310,096 | ENSG00000272235 | null | lncRNA | 3,310,000 | 3,311,000 | 2,347 |
chr1 | 4,479,131 | 4,484,375 | - | 4,484,375 | ENSG00000284694 | null | lncRNA | 4,484,000 | 4,485,000 | 3,340 |
chr1 | 2,492,300 | 2,493,258 | - | 2,493,258 | ENSG00000224387 | null | lncRNA | 2,493,000 | 2,494,000 | 1,676 |
chr1 | 9,035,106 | 9,088,478 | - | 9,088,478 | ENSG00000142583 | SLC2A5 | protein_coding | 9,088,000 | 9,089,000 | 7,306 |
chr1 | 4,973,381 | 4,981,568 | + | 4,973,381 | ENSG00000284674 | LINC02781 | lncRNA | 4,973,000 | 4,974,000 | 3,685 |
chr1 | 10,612,222 | 10,616,452 | - | 10,616,452 | ENSG00000287727 | null | lncRNA | 10,616,000 | 10,617,000 | 8,693 |
chr1 | 266,855 | 268,655 | + | 266,855 | ENSG00000286448 | null | lncRNA | 266,000 | 267,000 | 52 |
chr1 | 4,012,921 | 4,019,508 | - | 4,019,508 | ENSG00000284703 | null | lncRNA | 4,019,000 | 4,020,000 | 2,992 |
chr1 | 9,943,428 | 9,985,501 | + | 9,943,428 | ENSG00000173614 | NMNAT1 | protein_coding | 9,943,000 | 9,944,000 | 8,093 |
chr1 | 15,191,828 | 15,192,845 | - | 15,192,845 | ENSG00000215720 | MFFP1 | processed_pseudogene | 15,192,000 | 15,193,000 | 12,298 |
chr1 | 9,983,141 | 9,984,568 | + | 9,983,141 | ENSG00000241326 | null | lncRNA | 9,983,000 | 9,984,000 | 8,120 |
chr1 | 10,058,671 | 10,059,648 | + | 10,058,671 | ENSG00000233623 | PGAM1P11 | processed_pseudogene | 10,058,000 | 10,059,000 | 8,182 |
chr1 | 3,746,460 | 3,771,645 | + | 3,746,460 | ENSG00000162592 | CCDC27 | protein_coding | 3,746,000 | 3,747,000 | 2,757 |
chr1 | 11,479,155 | 11,537,551 | + | 11,479,155 | ENSG00000204624 | DISP3 | protein_coding | 11,479,000 | 11,480,000 | 9,466 |
chr1 | 2,590,639 | 2,633,016 | - | 2,633,016 | ENSG00000142606 | MMEL1 | protein_coding | 2,633,000 | 2,634,000 | 1,810 |
chr1 | 2,566,410 | 2,569,888 | + | 2,566,410 | ENSG00000225931 | null | TEC | 2,566,000 | 2,567,000 | 1,747 |
chr1 | 15,152,532 | 15,220,478 | + | 15,152,532 | ENSG00000171729 | TMEM51 | protein_coding | 15,152,000 | 15,153,000 | 12,259 |
chr1 | 2,493,437 | 2,494,479 | - | 2,494,479 | ENSG00000229393 | null | lncRNA | 2,494,000 | 2,495,000 | 1,677 |
chr1 | 16,246,840 | 16,352,480 | - | 16,352,480 | ENSG00000037637 | FBXO42 | protein_coding | 16,352,000 | 16,353,000 | 13,372 |
chr1 | 4,730,211 | 4,734,992 | - | 4,734,992 | ENSG00000287586 | null | lncRNA | 4,734,000 | 4,735,000 | 3,530 |
chr1 | 18,871,430 | 18,902,724 | - | 18,902,724 | ENSG00000159423 | ALDH4A1 | protein_coding | 18,902,000 | 18,903,000 | 15,543 |
chr1 | 4,551,735 | 4,552,145 | - | 4,552,145 | ENSG00000227169 | null | lncRNA | 4,552,000 | 4,553,000 | 3,386 |
chr1 | 5,554,747 | 5,554,881 | + | 5,554,747 | ENSG00000283356 | null | unprocessed_pseudogene | 5,554,000 | 5,555,000 | 4,098 |
chr1 | 2,391,775 | 2,405,442 | + | 2,391,775 | ENSG00000157916 | RER1 | protein_coding | 2,391,000 | 2,392,000 | 1,579 |
chr1 | 2,508,537 | 2,526,597 | - | 2,526,597 | ENSG00000157881 | PANK4 | protein_coding | 2,526,000 | 2,527,000 | 1,708 |
chr1 | 8,026,738 | 8,122,702 | + | 8,026,738 | ENSG00000238290 | ERRFI1-DT | lncRNA | 8,026,000 | 8,027,000 | 6,307 |
chr1 | 18,385,829 | 18,388,514 | + | 18,385,829 | ENSG00000225387 | null | lncRNA | 18,385,000 | 18,386,000 | 15,080 |
chr1 | 12,230,030 | 12,512,047 | + | 12,230,030 | ENSG00000048707 | VPS13D | protein_coding | 12,230,000 | 12,231,000 | 10,157 |
chr1 | 12,220,794 | 12,221,109 | - | 12,221,109 | ENSG00000225196 | RPL10P17 | processed_pseudogene | 12,221,000 | 12,222,000 | 10,149 |
chr1 | 9,997,206 | 10,016,021 | + | 9,997,206 | ENSG00000162444 | RBP7 | protein_coding | 9,997,000 | 9,998,000 | 8,129 |
chr1 | 22,364,630 | 22,366,482 | - | 22,366,482 | ENSG00000279625 | null | TEC | 22,366,000 | 22,367,000 | 18,700 |
chr1 | 9,100,305 | 9,129,102 | - | 9,129,102 | ENSG00000180758 | GPR157 | protein_coding | 9,129,000 | 9,130,000 | 7,344 |
chr1 | 22,428,838 | 22,531,157 | + | 22,428,838 | ENSG00000184677 | ZBTB40 | protein_coding | 22,428,000 | 22,429,000 | 18,757 |
chr1 | 16,228,674 | 16,231,335 | - | 16,231,335 | ENSG00000288398 | null | lncRNA | 16,231,000 | 16,232,000 | 13,262 |
chr1 | 16,159,266 | 16,161,883 | + | 16,159,266 | ENSG00000224621 | null | lncRNA | 16,159,000 | 16,160,000 | 13,194 |
chr1 | 11,500,803 | 11,502,016 | - | 11,502,016 | ENSG00000285833 | null | lncRNA | 11,502,000 | 11,503,000 | 9,486 |
chr1 | 5,862,811 | 5,992,473 | - | 5,992,473 | ENSG00000131697 | NPHP4 | protein_coding | 5,992,000 | 5,993,000 | 4,469 |
chr1 | 5,478,736 | 5,493,057 | - | 5,493,057 | ENSG00000284692 | null | lncRNA | 5,493,000 | 5,494,000 | 4,055 |
chr1 | 2,960,658 | 2,968,707 | - | 2,968,707 | ENSG00000284745 | null | lncRNA | 2,968,000 | 2,969,000 | 2,036 |
chr1 | 18,109,389 | 18,115,861 | + | 18,109,389 | ENSG00000280222 | null | TEC | 18,109,000 | 18,110,000 | 14,836 |
chr1 | 2,768,091 | 2,784,733 | + | 2,768,091 | ENSG00000285945 | null | lncRNA | 2,768,000 | 2,769,000 | 1,862 |
chr1 | 2,528,745 | 2,530,263 | - | 2,530,263 | ENSG00000197921 | HES5 | protein_coding | 2,530,000 | 2,531,000 | 1,712 |
chr1 | 25,043,707 | 25,113,120 | - | 25,113,120 | ENSG00000233755 | null | lncRNA | 25,113,000 | 25,114,000 | 21,248 |
chr1 | 16,241,213 | 16,241,398 | - | 16,241,398 | ENSG00000233929 | MT1XP1 | processed_pseudogene | 16,241,000 | 16,242,000 | 13,270 |
chr1 | 16,197,854 | 16,198,357 | + | 16,197,854 | ENSG00000234166 | ARHGEF19-AS1 | lncRNA | 16,197,000 | 16,198,000 | 13,232 |
chr1 | 18,513,118 | 18,513,383 | + | 18,513,118 | ENSG00000235282 | DYNLL1P3 | processed_pseudogene | 18,513,000 | 18,514,000 | 15,197 |
chr1 | 16,352,575 | 16,398,145 | + | 16,352,575 | ENSG00000055070 | SZRD1 | protein_coding | 16,352,000 | 16,353,000 | 13,372 |
chr1 | 26,111,165 | 26,125,555 | + | 26,111,165 | ENSG00000175087 | PDIK1L | protein_coding | 26,111,000 | 26,112,000 | 22,166 |
chr1 | 18,166,929 | 18,179,346 | - | 18,179,346 | ENSG00000230035 | IGSF21-AS1 | lncRNA | 18,179,000 | 18,180,000 | 14,896 |
chr1 | 25,581,478 | 25,590,356 | + | 25,581,478 | ENSG00000225643 | null | lncRNA | 25,581,000 | 25,582,000 | 21,664 |
chr1 | 5,480,787 | 5,482,028 | - | 5,482,028 | ENSG00000284666 | null | lncRNA | 5,482,000 | 5,483,000 | 4,047 |
chr1 | 28,109,739 | 28,110,298 | + | 28,109,739 | ENSG00000228943 | null | processed_pseudogene | 28,109,000 | 28,110,000 | 23,982 |
chr1 | 22,322,840 | 22,323,331 | - | 22,323,331 | ENSG00000234397 | PPIAP34 | processed_pseudogene | 22,323,000 | 22,324,000 | 18,662 |
chr1 | 26,046,665 | 26,049,099 | + | 26,046,665 | ENSG00000284309 | null | lncRNA | 26,046,000 | 26,047,000 | 22,105 |
chr1 | 30,732,469 | 30,757,774 | - | 30,757,774 | ENSG00000162511 | LAPTM5 | protein_coding | 30,757,000 | 30,758,000 | 26,333 |
chr1 | 26,037,252 | 26,046,118 | - | 26,046,118 | ENSG00000158014 | SLC30A2 | protein_coding | 26,046,000 | 26,047,000 | 22,105 |
chr1 | 28,369,582 | 28,500,364 | + | 28,369,582 | ENSG00000204138 | PHACTR4 | protein_coding | 28,369,000 | 28,370,000 | 24,222 |
chr1 | 25,125,053 | 25,125,454 | - | 25,125,454 | ENSG00000233419 | IFITM3P7 | processed_pseudogene | 25,125,000 | 25,126,000 | 21,259 |
chr1 | 9,501,092 | 9,503,471 | - | 9,503,471 | ENSG00000235263 | null | lncRNA | 9,503,000 | 9,504,000 | 7,688 |
chr1 | 29,708,851 | 29,709,547 | - | 29,709,547 | ENSG00000228176 | null | lncRNA | 29,709,000 | 29,710,000 | 25,424 |
chr1 | 30,226,523 | 30,226,615 | + | 30,226,523 | ENSG00000231251 | null | processed_pseudogene | 30,226,000 | 30,227,000 | 25,864 |
chr1 | 26,177,484 | 26,189,884 | + | 26,177,484 | ENSG00000142675 | CNKSR1 | protein_coding | 26,177,000 | 26,178,000 | 22,228 |
chr1 | 26,159,079 | 26,163,962 | + | 26,159,079 | ENSG00000197245 | FAM110D | protein_coding | 26,159,000 | 26,160,000 | 22,211 |
chr1 | 28,329,002 | 28,335,965 | + | 28,329,002 | ENSG00000130772 | MED18 | protein_coding | 28,329,000 | 28,330,000 | 24,189 |
chr1 | 31,487,589 | 31,488,930 | - | 31,488,930 | ENSG00000215899 | EEF1A1P46 | processed_pseudogene | 31,488,000 | 31,489,000 | 27,018 |
chr1 | 28,247,144 | 28,247,568 | + | 28,247,144 | ENSG00000271398 | null | lncRNA | 28,247,000 | 28,248,000 | 24,111 |
chr1 | 29,488,193 | 29,496,893 | - | 29,496,893 | ENSG00000225011 | null | unprocessed_pseudogene | 29,496,000 | 29,497,000 | 25,242 |
chr1 | 26,647,447 | 26,647,681 | + | 26,647,447 | ENSG00000235069 | null | processed_pseudogene | 26,647,000 | 26,648,000 | 22,647 |
chr1 | 29,755,175 | 29,790,597 | + | 29,755,175 | ENSG00000284676 | null | lncRNA | 29,755,000 | 29,756,000 | 25,459 |
chr1 | 2,632,568 | 2,636,620 | + | 2,632,568 | ENSG00000237058 | MMEL1-AS1 | lncRNA | 2,632,000 | 2,633,000 | 1,809 |
chr1 | 30,013,952 | 30,037,612 | - | 30,037,612 | ENSG00000233399 | LINC01648 | lncRNA | 30,037,000 | 30,038,000 | 25,698 |
chr1 | 27,098,809 | 27,166,981 | - | 27,166,981 | ENSG00000090020 | SLC9A1 | protein_coding | 27,166,000 | 27,167,000 | 23,122 |
chr1 | 25,543,606 | 25,568,886 | + | 25,543,606 | ENSG00000157978 | LDLRAP1 | protein_coding | 25,543,000 | 25,544,000 | 21,628 |
chr1 | 28,236,109 | 28,246,906 | + | 28,236,109 | ENSG00000130770 | ATP5IF1 | protein_coding | 28,236,000 | 28,237,000 | 24,100 |
chr1 | 31,333,067 | 31,346,799 | - | 31,346,799 | ENSG00000229044 | null | lncRNA | 31,346,000 | 31,347,000 | 26,886 |
chr1 | 30,869,466 | 30,908,758 | - | 30,908,758 | ENSG00000162512 | SDC3 | protein_coding | 30,908,000 | 30,909,000 | 26,480 |
chr1 | 25,959,767 | 25,998,117 | - | 25,998,117 | ENSG00000158006 | PAFAH2 | protein_coding | 25,998,000 | 25,999,000 | 22,061 |
chr1 | 28,120,449 | 28,121,321 | - | 28,121,321 | ENSG00000214812 | ARL8BP2 | processed_pseudogene | 28,121,000 | 28,122,000 | 23,992 |
chr1 | 28,199,456 | 28,233,029 | - | 28,233,029 | ENSG00000126698 | DNAJC8 | protein_coding | 28,233,000 | 28,234,000 | 24,097 |
chr1 | 31,263,245 | 31,263,681 | - | 31,263,681 | ENSG00000229447 | null | processed_pseudogene | 31,263,000 | 31,264,000 | 26,809 |
chr1 | 30,810,378 | 30,815,553 | - | 30,815,553 | ENSG00000229607 | null | lncRNA | 30,815,000 | 30,816,000 | 26,391 |
chr1 | 28,259,518 | 28,282,491 | + | 28,259,518 | ENSG00000130766 | SESN2 | protein_coding | 28,259,000 | 28,260,000 | 24,121 |
chr1 | 31,409,565 | 31,434,680 | + | 31,409,565 | ENSG00000168528 | SERINC2 | protein_coding | 31,409,000 | 31,410,000 | 26,944 |
chr1 | 4,654,609 | 4,792,534 | + | 4,654,609 | ENSG00000196581 | AJAP1 | protein_coding | 4,654,000 | 4,655,000 | 3,463 |
chr1 | 28,147,166 | 28,193,936 | - | 28,193,936 | ENSG00000169403 | PTAFR | protein_coding | 28,193,000 | 28,194,000 | 24,062 |
ChromBERT: A foundation model for learning interpretable representations for context-specific transcriptional regulatory networks
ChromBERT is a pre-trained deep learning model designed to capture the genome-wide co-association patterns of approximately one thousand transcription regulators, thereby enabling accurate representations of context-specific transcriptional regulatory networks (TRNs). As a foundational model, ChromBERT can be fine-tuned to adapt to various biological contexts through transfer learning. This significantly enhances our understanding of transcription regulation and offers a powerful tool for a broad range of research and clinical applications in different biological settings.
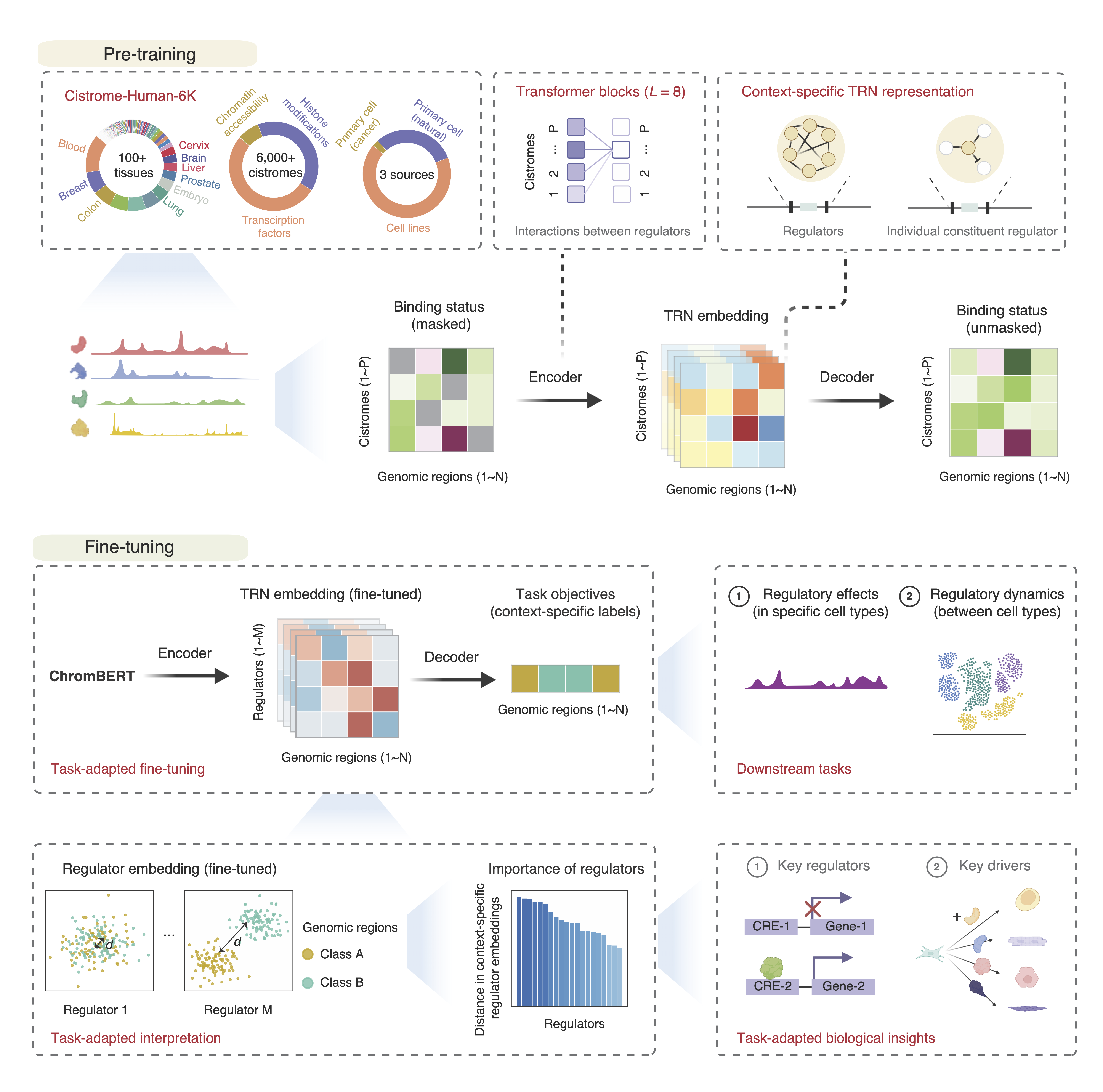
This repository provides the checkpoints and required dependencies for ChromBERT.
For usage, see ChromBERT for detail.
- Downloads last month
- 164

