markdown stringlengths 0 1.02M | code stringlengths 0 832k | output stringlengths 0 1.02M | license stringlengths 3 36 | path stringlengths 6 265 | repo_name stringlengths 6 127 |
|---|---|---|---|---|---|
数学和统计方法数组的数学函数对数组或数组某个轴向的数据进行统计计算;既可以通过数据实例方法调用;也可以当做nump顶级函数使用. | arr = np.arange(10).reshape(2,5)
print arr
print arr.mean()#计算数组平均值
print arr.mean(axis = 0)#可指定轴,用以统计该轴上的值
print arr.mean(axis = 1)
print np.mean(arr)
print np.mean(arr,axis = 0)
print np.mean(arr,axis = 1)
print arr.sum()#计算数组和
print arr.sum(axis = 0)
print arr.sum(axis = 1)
print np.sum(arr)
print np.sum(arr,axis = 0)
print np.sum(arr,axis = 1)
print arr.var()#计算方差差
print arr.var(axis = 0)#
print arr.var(axis = 1)#
print np.var(arr)#计算方差差
print np.var(arr,axis = 0)#
print np.var(arr,axis = 1)#
print arr.std()#计算标准差
print arr.std(axis = 0)#
print arr.std(axis = 1)#
print np.std(arr)#计算标准差
print np.std(arr,axis = 0)#
print np.std(arr,axis = 1)#
print arr
print arr.min()#计算最小值
print arr.min(axis = 0)#
print arr.min(axis = 1)#
print np.min(arr)#
print np.min(arr,axis = 0)#
print np.min(arr,axis = 1)#
print arr
print arr.max()#计算最大值
print arr.max(axis = 0)#
print arr.max(axis = 1)#
print np.max(arr)
print np.max(arr,axis = 0)#
print np.max(arr,axis = 1)#
print arr
print arr[0].argmin()#最小值的索引
print arr[1].argmin()#最小值的索引
print arr[0].argmax()#最大值的索引
print arr[1].argmax()#最大值的索引
print arr
print arr.cumsum()#不聚合,所有元素的累积和,而是返回中间结果构成的数组
arr = arr + 1
print arr
print arr.cumprod()#不聚合,所有元素的累积积,而是返回中间结果构成的数组 | [[ 1 2 3 4 5]
[ 6 7 8 9 10]]
[ 1 2 6 24 120 720 5040 40320 362880
3628800]
| Apache-2.0 | numpy/2-numpy-middle.ipynb | GmZhang3/data-science-ipython-notebooks |
用于布尔型数组的方法any()测试布尔型数组是否存在一个或多个Trueall()检查数组中所有值是否都为True | arr = np.random.randn(10)
# print
bools = arr > 0
print bools
print bools.any()
print np.any(bools)
print bools.all()
print np.all(bools)
arr = np.array([0,1,2,3,4])
print arr.any()#非布尔型数组,所有非0元素会被当做True | [ True False True True True False True True True False]
True
True
False
False
True
| Apache-2.0 | numpy/2-numpy-middle.ipynb | GmZhang3/data-science-ipython-notebooks |
排序 | arr = np.random.randn(10)
print arr
arr.sort()#与python内置列表排序一样;就地排序,会修改数组本身
print arr
arr = np.random.randn(10)
print arr
print np.sort(arr)#返回数组已排序副本 | [ 1.00826129 -1.33200021 -2.09423152 1.14618429 -1.94373366 -1.19964262
-0.97484352 -0.25508501 0.95594379 -0.92684744]
[-2.09423152 -1.94373366 -1.33200021 -1.19964262 -0.97484352 -0.92684744
-0.25508501 0.95594379 1.00826129 1.14618429]
[-0.13764091 -0.44657322 -0.4725095 -2.25229905 0.92481373 -0.4272677
-2.3459137 0.82869414 -0.2215158 -0.28151528]
[-2.3459137 -2.25229905 -0.4725095 -0.44657322 -0.4272677 -0.28151528
-0.2215158 -0.13764091 0.82869414 0.92481373]
| Apache-2.0 | numpy/2-numpy-middle.ipynb | GmZhang3/data-science-ipython-notebooks |
唯一化及其他的集合逻辑numpy针对一维数组的基本集合运算np.unique()找出数组中的唯一值,并返回已排序的结果 | names = np.array(['bob', 'joe', 'will', 'bob', 'will', 'joe', 'joe'])
print names
print np.unique(names) | ['bob' 'joe' 'will' 'bob' 'will' 'joe' 'joe']
['bob' 'joe' 'will']
| Apache-2.0 | numpy/2-numpy-middle.ipynb | GmZhang3/data-science-ipython-notebooks |
线性代数 | #矩阵乘法
x = np.array([[1,2,3],[4,5,6]])
y = np.array([[6,23],[-1,7],[8,9]])
print x
print y
print x.dot(y)
print np.dot(x,y) | [[1 2 3]
[4 5 6]]
[[ 6 23]
[-1 7]
[ 8 9]]
[[ 28 64]
[ 67 181]]
[[ 28 64]
[ 67 181]]
| Apache-2.0 | numpy/2-numpy-middle.ipynb | GmZhang3/data-science-ipython-notebooks |
随机数生成numpy.random模块对python内置的random进行补充,增接了一些用于高效生成多生概率分布的样本值的函数 | #从给定的上下限范围内随机选取整数
print np.random.randint(10)
#产生正态分布的样本值
print np.random.randn(3,2) | [[-0.05199971 -1.95031704]
[-1.42560357 0.78544126]
[ 0.47068984 -0.51157053]]
| Apache-2.0 | numpy/2-numpy-middle.ipynb | GmZhang3/data-science-ipython-notebooks |
数组组合 | a = np.array([1,2,3])
b = np.array([4,5,6])
c = np.arange(6).reshape(2,3)
d = np.arange(2,8).reshape(2,3)
print(a)
print(b)
print(c)
print(d)
np.concatenate([c,d])
# In machine learning, useful to enrich or
# add new/concatenate features with hstack
np.hstack([c, d])
# Use broadcasting when needed to do this automatically
np.vstack([a,b, d]) | _____no_output_____ | Apache-2.0 | numpy/2-numpy-middle.ipynb | GmZhang3/data-science-ipython-notebooks |
A Scientific Deep Dive Into SageMaker LDA1. [Introduction](Introduction)1. [Setup](Setup)1. [Data Exploration](DataExploration)1. [Training](Training)1. [Inference](Inference)1. [Epilogue](Epilogue) Introduction***Amazon SageMaker LDA is an unsupervised learning algorithm that attempts to describe a set of observations as a mixture of distinct categories. Latent Dirichlet Allocation (LDA) is most commonly used to discover a user-specified number of topics shared by documents within a text corpus. Here each observation is a document, the features are the presence (or occurrence count) of each word, and the categories are the topics. Since the method is unsupervised, the topics are not specified up front, and are not guaranteed to align with how a human may naturally categorize documents. The topics are learned as a probability distribution over the words that occur in each document. Each document, in turn, is described as a mixture of topics.This notebook is similar to **LDA-Introduction.ipynb** but its objective and scope are a different. We will be taking a deeper dive into the theory. The primary goals of this notebook are,* to understand the LDA model and the example dataset,* understand how the Amazon SageMaker LDA algorithm works,* interpret the meaning of the inference output.Former knowledge of LDA is not required. However, we will run through concepts rather quickly and at least a foundational knowledge of mathematics or machine learning is recommended. Suggested references are provided, as appropriate. | !conda install -y scipy
%matplotlib inline
import os, re, tarfile
import boto3
import matplotlib.pyplot as plt
import mxnet as mx
import numpy as np
np.set_printoptions(precision=3, suppress=True)
# some helpful utility functions are defined in the Python module
# "generate_example_data" located in the same directory as this
# notebook
from generate_example_data import (
generate_griffiths_data, match_estimated_topics,
plot_lda, plot_lda_topics)
# accessing the SageMaker Python SDK
import sagemaker
from sagemaker.amazon.common import numpy_to_record_serializer
from sagemaker.predictor import csv_serializer, json_deserializer | _____no_output_____ | Apache-2.0 | scientific_details_of_algorithms/lda_topic_modeling/LDA-Science.ipynb | karim7262/amazon-sagemaker-examples |
Setup****This notebook was created and tested on an ml.m4.xlarge notebook instance.*We first need to specify some AWS credentials; specifically data locations and access roles. This is the only cell of this notebook that you will need to edit. In particular, we need the following data:* `bucket` - An S3 bucket accessible by this account. * Used to store input training data and model data output. * Should be withing the same region as this notebook instance, training, and hosting.* `prefix` - The location in the bucket where this notebook's input and and output data will be stored. (The default value is sufficient.)* `role` - The IAM Role ARN used to give training and hosting access to your data. * See documentation on how to create these. * The script below will try to determine an appropriate Role ARN. | from sagemaker import get_execution_role
role = get_execution_role()
bucket = '<your_s3_bucket_name_here>'
prefix = 'sagemaker/lda_science'
print('Training input/output will be stored in {}/{}'.format(bucket, prefix))
print('\nIAM Role: {}'.format(role)) | _____no_output_____ | Apache-2.0 | scientific_details_of_algorithms/lda_topic_modeling/LDA-Science.ipynb | karim7262/amazon-sagemaker-examples |
The LDA ModelAs mentioned above, LDA is a model for discovering latent topics describing a collection of documents. In this section we will give a brief introduction to the model. Let,* $M$ = the number of *documents* in a corpus* $N$ = the average *length* of a document.* $V$ = the size of the *vocabulary* (the total number of unique words)We denote a *document* by a vector $w \in \mathbb{R}^V$ where $w_i$ equals the number of times the $i$th word in the vocabulary occurs within the document. This is called the "bag-of-words" format of representing a document.$$\underbrace{w}_{\text{document}} = \overbrace{\big[ w_1, w_2, \ldots, w_V \big] }^{\text{word counts}},\quadV = \text{vocabulary size}$$The *length* of a document is equal to the total number of words in the document: $N_w = \sum_{i=1}^V w_i$.An LDA model is defined by two parameters: a topic-word distribution matrix $\beta \in \mathbb{R}^{K \times V}$ and a Dirichlet topic prior $\alpha \in \mathbb{R}^K$. In particular, let,$$\beta = \left[ \beta_1, \ldots, \beta_K \right]$$be a collection of $K$ *topics* where each topic $\beta_k \in \mathbb{R}^V$ is represented as probability distribution over the vocabulary. One of the utilities of the LDA model is that a given word is allowed to appear in multiple topics with positive probability. The Dirichlet topic prior is a vector $\alpha \in \mathbb{R}^K$ such that $\alpha_k > 0$ for all $k$. Data Exploration--- An Example DatasetBefore explaining further let's get our hands dirty with an example dataset. The following synthetic data comes from [1] and comes with a very useful visual interpretation.> [1] Thomas Griffiths and Mark Steyvers. *Finding Scientific Topics.* Proceedings of the National Academy of Science, 101(suppl 1):5228-5235, 2004. | print('Generating example data...')
num_documents = 6000
known_alpha, known_beta, documents, topic_mixtures = generate_griffiths_data(
num_documents=num_documents, num_topics=10)
num_topics, vocabulary_size = known_beta.shape
# separate the generated data into training and tests subsets
num_documents_training = int(0.9*num_documents)
num_documents_test = num_documents - num_documents_training
documents_training = documents[:num_documents_training]
documents_test = documents[num_documents_training:]
topic_mixtures_training = topic_mixtures[:num_documents_training]
topic_mixtures_test = topic_mixtures[num_documents_training:]
print('documents_training.shape = {}'.format(documents_training.shape))
print('documents_test.shape = {}'.format(documents_test.shape)) | _____no_output_____ | Apache-2.0 | scientific_details_of_algorithms/lda_topic_modeling/LDA-Science.ipynb | karim7262/amazon-sagemaker-examples |
Let's start by taking a closer look at the documents. Note that the vocabulary size of these data is $V = 25$. The average length of each document in this data set is 150. (See `generate_griffiths_data.py`.) | print('First training document =\n{}'.format(documents_training[0]))
print('\nVocabulary size = {}'.format(vocabulary_size))
print('Length of first document = {}'.format(documents_training[0].sum()))
average_document_length = documents.sum(axis=1).mean()
print('Observed average document length = {}'.format(average_document_length)) | _____no_output_____ | Apache-2.0 | scientific_details_of_algorithms/lda_topic_modeling/LDA-Science.ipynb | karim7262/amazon-sagemaker-examples |
The example data set above also returns the LDA parameters,$$(\alpha, \beta)$$used to generate the documents. Let's examine the first topic and verify that it is a probability distribution on the vocabulary. | print('First topic =\n{}'.format(known_beta[0]))
print('\nTopic-word probability matrix (beta) shape: (num_topics, vocabulary_size) = {}'.format(known_beta.shape))
print('\nSum of elements of first topic = {}'.format(known_beta[0].sum())) | _____no_output_____ | Apache-2.0 | scientific_details_of_algorithms/lda_topic_modeling/LDA-Science.ipynb | karim7262/amazon-sagemaker-examples |
Unlike some clustering algorithms, one of the versatilities of the LDA model is that a given word can belong to multiple topics. The probability of that word occurring in each topic may differ, as well. This is reflective of real-world data where, for example, the word *"rover"* appears in a *"dogs"* topic as well as in a *"space exploration"* topic.In our synthetic example dataset, the first word in the vocabulary belongs to both Topic 1 and Topic 6 with non-zero probability. | print('Topic #1:\n{}'.format(known_beta[0]))
print('Topic #6:\n{}'.format(known_beta[5])) | _____no_output_____ | Apache-2.0 | scientific_details_of_algorithms/lda_topic_modeling/LDA-Science.ipynb | karim7262/amazon-sagemaker-examples |
Human beings are visual creatures, so it might be helpful to come up with a visual representation of these documents.In the below plots, each pixel of a document represents a word. The greyscale intensity is a measure of how frequently that word occurs within the document. Below we plot the first few documents of the training set reshaped into 5x5 pixel grids. | %matplotlib inline
fig = plot_lda(documents_training, nrows=3, ncols=4, cmap='gray_r', with_colorbar=True)
fig.suptitle('$w$ - Document Word Counts')
fig.set_dpi(160) | _____no_output_____ | Apache-2.0 | scientific_details_of_algorithms/lda_topic_modeling/LDA-Science.ipynb | karim7262/amazon-sagemaker-examples |
When taking a close look at these documents we can see some patterns in the word distributions suggesting that, perhaps, each topic represents a "column" or "row" of words with non-zero probability and that each document is composed primarily of a handful of topics.Below we plots the *known* topic-word probability distributions, $\beta$. Similar to the documents we reshape each probability distribution to a $5 \times 5$ pixel image where the color represents the probability of that each word occurring in the topic. | %matplotlib inline
fig = plot_lda(known_beta, nrows=1, ncols=10)
fig.suptitle(r'Known $\beta$ - Topic-Word Probability Distributions')
fig.set_dpi(160)
fig.set_figheight(2) | _____no_output_____ | Apache-2.0 | scientific_details_of_algorithms/lda_topic_modeling/LDA-Science.ipynb | karim7262/amazon-sagemaker-examples |
These 10 topics were used to generate the document corpus. Next, we will learn about how this is done. Generating DocumentsLDA is a generative model, meaning that the LDA parameters $(\alpha, \beta)$ are used to construct documents word-by-word by drawing from the topic-word distributions. In fact, looking closely at the example documents above you can see that some documents sample more words from some topics than from others.LDA works as follows: given * $M$ documents $w^{(1)}, w^{(2)}, \ldots, w^{(M)}$,* an average document length of $N$,* and an LDA model $(\alpha, \beta)$.**For** each document, $w^{(m)}$:* sample a topic mixture: $\theta^{(m)} \sim \text{Dirichlet}(\alpha)$* **For** each word $n$ in the document: * Sample a topic $z_n^{(m)} \sim \text{Multinomial}\big( \theta^{(m)} \big)$ * Sample a word from this topic, $w_n^{(m)} \sim \text{Multinomial}\big( \beta_{z_n^{(m)}} \; \big)$ * Add to documentThe [plate notation](https://en.wikipedia.org/wiki/Plate_notation) for the LDA model, introduced in [2], encapsulates this process pictorially.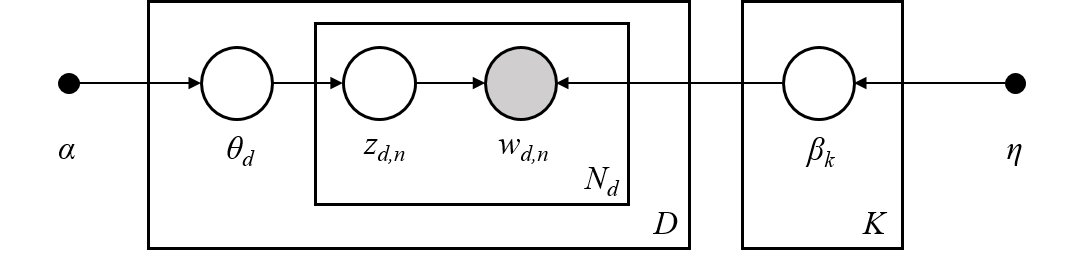> [2] David M Blei, Andrew Y Ng, and Michael I Jordan. Latent Dirichlet Allocation. Journal of Machine Learning Research, 3(Jan):993–1022, 2003. Topic MixturesFor the documents we generated above lets look at their corresponding topic mixtures, $\theta \in \mathbb{R}^K$. The topic mixtures represent the probablility that a given word of the document is sampled from a particular topic. For example, if the topic mixture of an input document $w$ is,$$\theta = \left[ 0.3, 0.2, 0, 0.5, 0, \ldots, 0 \right]$$then $w$ is 30% generated from the first topic, 20% from the second topic, and 50% from the fourth topic. In particular, the words contained in the document are sampled from the first topic-word probability distribution 30% of the time, from the second distribution 20% of the time, and the fourth disribution 50% of the time.The objective of inference, also known as scoring, is to determine the most likely topic mixture of a given input document. Colloquially, this means figuring out which topics appear within a given document and at what ratios. We will perform infernece later in the [Inference](Inference) section.Since we generated these example documents using the LDA model we know the topic mixture generating them. Let's examine these topic mixtures. | print('First training document =\n{}'.format(documents_training[0]))
print('\nVocabulary size = {}'.format(vocabulary_size))
print('Length of first document = {}'.format(documents_training[0].sum()))
print('First training document topic mixture =\n{}'.format(topic_mixtures_training[0]))
print('\nNumber of topics = {}'.format(num_topics))
print('sum(theta) = {}'.format(topic_mixtures_training[0].sum())) | _____no_output_____ | Apache-2.0 | scientific_details_of_algorithms/lda_topic_modeling/LDA-Science.ipynb | karim7262/amazon-sagemaker-examples |
We plot the first document along with its topic mixture. We also plot the topic-word probability distributions again for reference. | %matplotlib inline
fig, (ax1,ax2) = plt.subplots(2, 1)
ax1.matshow(documents[0].reshape(5,5), cmap='gray_r')
ax1.set_title(r'$w$ - Document', fontsize=20)
ax1.set_xticks([])
ax1.set_yticks([])
cax2 = ax2.matshow(topic_mixtures[0].reshape(1,-1), cmap='Reds', vmin=0, vmax=1)
cbar = fig.colorbar(cax2, orientation='horizontal')
ax2.set_title(r'$\theta$ - Topic Mixture', fontsize=20)
ax2.set_xticks([])
ax2.set_yticks([])
fig.set_dpi(100)
%matplotlib inline
# pot
fig = plot_lda(known_beta, nrows=1, ncols=10)
fig.suptitle(r'Known $\beta$ - Topic-Word Probability Distributions')
fig.set_dpi(160)
fig.set_figheight(1.5) | _____no_output_____ | Apache-2.0 | scientific_details_of_algorithms/lda_topic_modeling/LDA-Science.ipynb | karim7262/amazon-sagemaker-examples |
Finally, let's plot several documents with their corresponding topic mixtures. We can see how topics with large weight in the document lead to more words in the document within the corresponding "row" or "column". | %matplotlib inline
fig = plot_lda_topics(documents_training, 3, 4, topic_mixtures=topic_mixtures)
fig.suptitle(r'$(w,\theta)$ - Documents with Known Topic Mixtures')
fig.set_dpi(160) | _____no_output_____ | Apache-2.0 | scientific_details_of_algorithms/lda_topic_modeling/LDA-Science.ipynb | karim7262/amazon-sagemaker-examples |
Training***In this section we will give some insight into how AWS SageMaker LDA fits an LDA model to a corpus, create an run a SageMaker LDA training job, and examine the output trained model. Topic Estimation using Tensor DecompositionsGiven a document corpus, Amazon SageMaker LDA uses a spectral tensor decomposition technique to determine the LDA model $(\alpha, \beta)$ which most likely describes the corpus. See [1] for a primary reference of the theory behind the algorithm. The spectral decomposition, itself, is computed using the CPDecomp algorithm described in [2].The overall idea is the following: given a corpus of documents $\mathcal{W} = \{w^{(1)}, \ldots, w^{(M)}\}, \; w^{(m)} \in \mathbb{R}^V,$ we construct a statistic tensor,$$T \in \bigotimes^3 \mathbb{R}^V$$such that the spectral decomposition of the tensor is approximately the LDA parameters $\alpha \in \mathbb{R}^K$ and $\beta \in \mathbb{R}^{K \times V}$ which maximize the likelihood of observing the corpus for a given number of topics, $K$,$$T \approx \sum_{k=1}^K \alpha_k \; (\beta_k \otimes \beta_k \otimes \beta_k)$$This statistic tensor encapsulates information from the corpus such as the document mean, cross correlation, and higher order statistics. For details, see [1].> [1] Animashree Anandkumar, Rong Ge, Daniel Hsu, Sham Kakade, and Matus Telgarsky. *"Tensor Decompositions for Learning Latent Variable Models"*, Journal of Machine Learning Research, 15:2773–2832, 2014.>> [2] Tamara Kolda and Brett Bader. *"Tensor Decompositions and Applications"*. SIAM Review, 51(3):455–500, 2009. Store Data on S3Before we run training we need to prepare the data.A SageMaker training job needs access to training data stored in an S3 bucket. Although training can accept data of various formats we convert the documents MXNet RecordIO Protobuf format before uploading to the S3 bucket defined at the beginning of this notebook. | # convert documents_training to Protobuf RecordIO format
recordio_protobuf_serializer = numpy_to_record_serializer()
fbuffer = recordio_protobuf_serializer(documents_training)
# upload to S3 in bucket/prefix/train
fname = 'lda.data'
s3_object = os.path.join(prefix, 'train', fname)
boto3.Session().resource('s3').Bucket(bucket).Object(s3_object).upload_fileobj(fbuffer)
s3_train_data = 's3://{}/{}'.format(bucket, s3_object)
print('Uploaded data to S3: {}'.format(s3_train_data)) | _____no_output_____ | Apache-2.0 | scientific_details_of_algorithms/lda_topic_modeling/LDA-Science.ipynb | karim7262/amazon-sagemaker-examples |
Next, we specify a Docker container containing the SageMaker LDA algorithm. For your convenience, a region-specific container is automatically chosen for you to minimize cross-region data communication | containers = {
'us-west-2': '266724342769.dkr.ecr.us-west-2.amazonaws.com/lda:latest',
'us-east-1': '766337827248.dkr.ecr.us-east-1.amazonaws.com/lda:latest',
'us-east-2': '999911452149.dkr.ecr.us-east-2.amazonaws.com/lda:latest',
'eu-west-1': '999678624901.dkr.ecr.eu-west-1.amazonaws.com/lda:latest'
}
region_name = boto3.Session().region_name
container = containers[region_name]
print('Using SageMaker LDA container: {} ({})'.format(container, region_name)) | _____no_output_____ | Apache-2.0 | scientific_details_of_algorithms/lda_topic_modeling/LDA-Science.ipynb | karim7262/amazon-sagemaker-examples |
Training ParametersParticular to a SageMaker LDA training job are the following hyperparameters:* **`num_topics`** - The number of topics or categories in the LDA model. * Usually, this is not known a priori. * In this example, howevever, we know that the data is generated by five topics.* **`feature_dim`** - The size of the *"vocabulary"*, in LDA parlance. * In this example, this is equal 25.* **`mini_batch_size`** - The number of input training documents.* **`alpha0`** - *(optional)* a measurement of how "mixed" are the topic-mixtures. * When `alpha0` is small the data tends to be represented by one or few topics. * When `alpha0` is large the data tends to be an even combination of several or many topics. * The default value is `alpha0 = 1.0`.In addition to these LDA model hyperparameters, we provide additional parameters defining things like the EC2 instance type on which training will run, the S3 bucket containing the data, and the AWS access role. Note that,* Recommended instance type: `ml.c4`* Current limitations: * SageMaker LDA *training* can only run on a single instance. * SageMaker LDA does not take advantage of GPU hardware. * (The Amazon AI Algorithms team is working hard to provide these capabilities in a future release!) Using the above configuration create a SageMaker client and use the client to create a training job. | session = sagemaker.Session()
# specify general training job information
lda = sagemaker.estimator.Estimator(
container,
role,
output_path='s3://{}/{}/output'.format(bucket, prefix),
train_instance_count=1,
train_instance_type='ml.c4.2xlarge',
sagemaker_session=session,
)
# set algorithm-specific hyperparameters
lda.set_hyperparameters(
num_topics=num_topics,
feature_dim=vocabulary_size,
mini_batch_size=num_documents_training,
alpha0=1.0,
)
# run the training job on input data stored in S3
lda.fit({'train': s3_train_data}) | _____no_output_____ | Apache-2.0 | scientific_details_of_algorithms/lda_topic_modeling/LDA-Science.ipynb | karim7262/amazon-sagemaker-examples |
If you see the message> `===== Job Complete =====`at the bottom of the output logs then that means training sucessfully completed and the output LDA model was stored in the specified output path. You can also view information about and the status of a training job using the AWS SageMaker console. Just click on the "Jobs" tab and select training job matching the training job name, below: | print('Training job name: {}'.format(lda.latest_training_job.job_name)) | _____no_output_____ | Apache-2.0 | scientific_details_of_algorithms/lda_topic_modeling/LDA-Science.ipynb | karim7262/amazon-sagemaker-examples |
Inspecting the Trained ModelWe know the LDA parameters $(\alpha, \beta)$ used to generate the example data. How does the learned model compare the known one? In this section we will download the model data and measure how well SageMaker LDA did in learning the model.First, we download the model data. SageMaker will output the model in > `s3:////output//output/model.tar.gz`.SageMaker LDA stores the model as a two-tuple $(\alpha, \beta)$ where each LDA parameter is an MXNet NDArray. | # download and extract the model file from S3
job_name = lda.latest_training_job.job_name
model_fname = 'model.tar.gz'
model_object = os.path.join(prefix, 'output', job_name, 'output', model_fname)
boto3.Session().resource('s3').Bucket(bucket).Object(model_object).download_file(fname)
with tarfile.open(fname) as tar:
tar.extractall()
print('Downloaded and extracted model tarball: {}'.format(model_object))
# obtain the model file
model_list = [fname for fname in os.listdir('.') if fname.startswith('model_')]
model_fname = model_list[0]
print('Found model file: {}'.format(model_fname))
# get the model from the model file and store in Numpy arrays
alpha, beta = mx.ndarray.load(model_fname)
learned_alpha_permuted = alpha.asnumpy()
learned_beta_permuted = beta.asnumpy()
print('\nLearned alpha.shape = {}'.format(learned_alpha_permuted.shape))
print('Learned beta.shape = {}'.format(learned_beta_permuted.shape)) | _____no_output_____ | Apache-2.0 | scientific_details_of_algorithms/lda_topic_modeling/LDA-Science.ipynb | karim7262/amazon-sagemaker-examples |
Presumably, SageMaker LDA has found the topics most likely used to generate the training corpus. However, even if this is case the topics would not be returned in any particular order. Therefore, we match the found topics to the known topics closest in L1-norm in order to find the topic permutation.Note that we will use the `permutation` later during inference to match known topic mixtures to found topic mixtures.Below plot the known topic-word probability distribution, $\beta \in \mathbb{R}^{K \times V}$ next to the distributions found by SageMaker LDA as well as the L1-norm errors between the two. | permutation, learned_beta = match_estimated_topics(known_beta, learned_beta_permuted)
learned_alpha = learned_alpha_permuted[permutation]
fig = plot_lda(np.vstack([known_beta, learned_beta]), 2, 10)
fig.set_dpi(160)
fig.suptitle('Known vs. Found Topic-Word Probability Distributions')
fig.set_figheight(3)
beta_error = np.linalg.norm(known_beta - learned_beta, 1)
alpha_error = np.linalg.norm(known_alpha - learned_alpha, 1)
print('L1-error (beta) = {}'.format(beta_error))
print('L1-error (alpha) = {}'.format(alpha_error)) | _____no_output_____ | Apache-2.0 | scientific_details_of_algorithms/lda_topic_modeling/LDA-Science.ipynb | karim7262/amazon-sagemaker-examples |
Not bad!In the eyeball-norm the topics match quite well. In fact, the topic-word distribution error is approximately 2%. Inference***A trained model does nothing on its own. We now want to use the model we computed to perform inference on data. For this example, that means predicting the topic mixture representing a given document.We create an inference endpoint using the SageMaker Python SDK `deploy()` function from the job we defined above. We specify the instance type where inference is computed as well as an initial number of instances to spin up. | lda_inference = lda.deploy(
initial_instance_count=1,
instance_type='ml.m4.xlarge', # LDA inference may work better at scale on ml.c4 instances
) | _____no_output_____ | Apache-2.0 | scientific_details_of_algorithms/lda_topic_modeling/LDA-Science.ipynb | karim7262/amazon-sagemaker-examples |
Congratulations! You now have a functioning SageMaker LDA inference endpoint. You can confirm the endpoint configuration and status by navigating to the "Endpoints" tab in the AWS SageMaker console and selecting the endpoint matching the endpoint name, below: | print('Endpoint name: {}'.format(lda_inference.endpoint)) | _____no_output_____ | Apache-2.0 | scientific_details_of_algorithms/lda_topic_modeling/LDA-Science.ipynb | karim7262/amazon-sagemaker-examples |
With this realtime endpoint at our fingertips we can finally perform inference on our training and test data.We can pass a variety of data formats to our inference endpoint. In this example we will demonstrate passing CSV-formatted data. Other available formats are JSON-formatted, JSON-sparse-formatter, and RecordIO Protobuf. We make use of the SageMaker Python SDK utilities `csv_serializer` and `json_deserializer` when configuring the inference endpoint. | lda_inference.content_type = 'text/csv'
lda_inference.serializer = csv_serializer
lda_inference.deserializer = json_deserializer | _____no_output_____ | Apache-2.0 | scientific_details_of_algorithms/lda_topic_modeling/LDA-Science.ipynb | karim7262/amazon-sagemaker-examples |
We pass some test documents to the inference endpoint. Note that the serializer and deserializer will atuomatically take care of the datatype conversion. | results = lda_inference.predict(documents_test[:12])
print(results) | _____no_output_____ | Apache-2.0 | scientific_details_of_algorithms/lda_topic_modeling/LDA-Science.ipynb | karim7262/amazon-sagemaker-examples |
It may be hard to see but the output format of SageMaker LDA inference endpoint is a Python dictionary with the following format.```{ 'predictions': [ {'topic_mixture': [ ... ] }, {'topic_mixture': [ ... ] }, {'topic_mixture': [ ... ] }, ... ]}```We extract the topic mixtures, themselves, corresponding to each of the input documents. | inferred_topic_mixtures_permuted = np.array([prediction['topic_mixture'] for prediction in results['predictions']])
print('Inferred topic mixtures (permuted):\n\n{}'.format(inferred_topic_mixtures_permuted)) | _____no_output_____ | Apache-2.0 | scientific_details_of_algorithms/lda_topic_modeling/LDA-Science.ipynb | karim7262/amazon-sagemaker-examples |
Inference AnalysisRecall that although SageMaker LDA successfully learned the underlying topics which generated the sample data the topics were in a different order. Before we compare to known topic mixtures $\theta \in \mathbb{R}^K$ we should also permute the inferred topic mixtures | inferred_topic_mixtures = inferred_topic_mixtures_permuted[:,permutation]
print('Inferred topic mixtures:\n\n{}'.format(inferred_topic_mixtures)) | _____no_output_____ | Apache-2.0 | scientific_details_of_algorithms/lda_topic_modeling/LDA-Science.ipynb | karim7262/amazon-sagemaker-examples |
Let's plot these topic mixture probability distributions alongside the known ones. | %matplotlib inline
# create array of bar plots
width = 0.4
x = np.arange(10)
nrows, ncols = 3, 4
fig, ax = plt.subplots(nrows, ncols, sharey=True)
for i in range(nrows):
for j in range(ncols):
index = i*ncols + j
ax[i,j].bar(x, topic_mixtures_test[index], width, color='C0')
ax[i,j].bar(x+width, inferred_topic_mixtures[index], width, color='C1')
ax[i,j].set_xticks(range(num_topics))
ax[i,j].set_yticks(np.linspace(0,1,5))
ax[i,j].grid(which='major', axis='y')
ax[i,j].set_ylim([0,1])
ax[i,j].set_xticklabels([])
if (i==(nrows-1)):
ax[i,j].set_xticklabels(range(num_topics), fontsize=7)
if (j==0):
ax[i,j].set_yticklabels([0,'',0.5,'',1.0], fontsize=7)
fig.suptitle('Known vs. Inferred Topic Mixtures')
ax_super = fig.add_subplot(111, frameon=False)
ax_super.tick_params(labelcolor='none', top='off', bottom='off', left='off', right='off')
ax_super.grid(False)
ax_super.set_xlabel('Topic Index')
ax_super.set_ylabel('Topic Probability')
fig.set_dpi(160) | _____no_output_____ | Apache-2.0 | scientific_details_of_algorithms/lda_topic_modeling/LDA-Science.ipynb | karim7262/amazon-sagemaker-examples |
In the eyeball-norm these look quite comparable.Let's be more scientific about this. Below we compute and plot the distribution of L1-errors from **all** of the test documents. Note that we send a new payload of test documents to the inference endpoint and apply the appropriate permutation to the output. | %%time
# create a payload containing all of the test documents and run inference again
#
# TRY THIS:
# try switching between the test data set and a subset of the training
# data set. It is likely that LDA inference will perform better against
# the training set than the holdout test set.
#
payload_documents = documents_test # Example 1
known_topic_mixtures = topic_mixtures_test # Example 1
#payload_documents = documents_training[:600]; # Example 2
#known_topic_mixtures = topic_mixtures_training[:600] # Example 2
print('Invoking endpoint...\n')
results = lda_inference.predict(payload_documents)
inferred_topic_mixtures_permuted = np.array([prediction['topic_mixture'] for prediction in results['predictions']])
inferred_topic_mixtures = inferred_topic_mixtures_permuted[:,permutation]
print('known_topics_mixtures.shape = {}'.format(known_topic_mixtures.shape))
print('inferred_topics_mixtures_test.shape = {}\n'.format(inferred_topic_mixtures.shape))
%matplotlib inline
l1_errors = np.linalg.norm((inferred_topic_mixtures - known_topic_mixtures), 1, axis=1)
# plot the error freqency
fig, ax_frequency = plt.subplots()
bins = np.linspace(0,1,40)
weights = np.ones_like(l1_errors)/len(l1_errors)
freq, bins, _ = ax_frequency.hist(l1_errors, bins=50, weights=weights, color='C0')
ax_frequency.set_xlabel('L1-Error')
ax_frequency.set_ylabel('Frequency', color='C0')
# plot the cumulative error
shift = (bins[1]-bins[0])/2
x = bins[1:] - shift
ax_cumulative = ax_frequency.twinx()
cumulative = np.cumsum(freq)/sum(freq)
ax_cumulative.plot(x, cumulative, marker='o', color='C1')
ax_cumulative.set_ylabel('Cumulative Frequency', color='C1')
# align grids and show
freq_ticks = np.linspace(0, 1.5*freq.max(), 5)
freq_ticklabels = np.round(100*freq_ticks)/100
ax_frequency.set_yticks(freq_ticks)
ax_frequency.set_yticklabels(freq_ticklabels)
ax_cumulative.set_yticks(np.linspace(0, 1, 5))
ax_cumulative.grid(which='major', axis='y')
ax_cumulative.set_ylim((0,1))
fig.suptitle('Topic Mixutre L1-Errors')
fig.set_dpi(110) | _____no_output_____ | Apache-2.0 | scientific_details_of_algorithms/lda_topic_modeling/LDA-Science.ipynb | karim7262/amazon-sagemaker-examples |
Machine learning algorithms are not perfect and the data above suggests this is true of SageMaker LDA. With more documents and some hyperparameter tuning we can obtain more accurate results against the known topic-mixtures.For now, let's just investigate the documents-topic mixture pairs that seem to do well as well as those that do not. Below we retreive a document and topic mixture corresponding to a small L1-error as well as one with a large L1-error. | N = 6
good_idx = (l1_errors < 0.05)
good_documents = payload_documents[good_idx][:N]
good_topic_mixtures = inferred_topic_mixtures[good_idx][:N]
poor_idx = (l1_errors > 0.3)
poor_documents = payload_documents[poor_idx][:N]
poor_topic_mixtures = inferred_topic_mixtures[poor_idx][:N]
%matplotlib inline
fig = plot_lda_topics(good_documents, 2, 3, topic_mixtures=good_topic_mixtures)
fig.suptitle('Documents With Accurate Inferred Topic-Mixtures')
fig.set_dpi(120)
%matplotlib inline
fig = plot_lda_topics(poor_documents, 2, 3, topic_mixtures=poor_topic_mixtures)
fig.suptitle('Documents With Inaccurate Inferred Topic-Mixtures')
fig.set_dpi(120) | _____no_output_____ | Apache-2.0 | scientific_details_of_algorithms/lda_topic_modeling/LDA-Science.ipynb | karim7262/amazon-sagemaker-examples |
In this example set the documents on which inference was not as accurate tend to have a denser topic-mixture. This makes sense when extrapolated to real-world datasets: it can be difficult to nail down which topics are represented in a document when the document uses words from a large subset of the vocabulary. Stop / Close the EndpointFinally, we should delete the endpoint before we close the notebook.To do so execute the cell below. Alternately, you can navigate to the "Endpoints" tab in the SageMaker console, select the endpoint with the name stored in the variable `endpoint_name`, and select "Delete" from the "Actions" dropdown menu. | sagemaker.Session().delete_endpoint(lda_inference.endpoint) | _____no_output_____ | Apache-2.0 | scientific_details_of_algorithms/lda_topic_modeling/LDA-Science.ipynb | karim7262/amazon-sagemaker-examples |
Load Library | !pip install wget
!pip install keras-tcn
import wget
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from tensorflow.keras import Input, Model
from tensorflow.keras.layers import Dense
from tqdm.notebook import tqdm
from tcn import TCN
wget.download("https://github.com/philipperemy/keras-tcn/raw/master/tasks/monthly-milk-production-pounds-p.csv") | _____no_output_____ | MIT | TCN_TimeSeries_Approach.ipynb | ashishpatel26/tcn-keras-Examples |
Read the dataset | milk = pd.read_csv('monthly-milk-production-pounds-p.csv', index_col=0, parse_dates=True) | _____no_output_____ | MIT | TCN_TimeSeries_Approach.ipynb | ashishpatel26/tcn-keras-Examples |
Display top5 Record | print(milk.shape)
milk.head() | (168, 1)
| MIT | TCN_TimeSeries_Approach.ipynb | ashishpatel26/tcn-keras-Examples |
Lookback 12 month windows | lookback_window = 12 | _____no_output_____ | MIT | TCN_TimeSeries_Approach.ipynb | ashishpatel26/tcn-keras-Examples |
Convert Milk Data into Numpy Array | milk = milk.values | _____no_output_____ | MIT | TCN_TimeSeries_Approach.ipynb | ashishpatel26/tcn-keras-Examples |
Convert in to X, y format | x = []
y = []
for i in tqdm(range(lookback_window, len(milk))):
x.append(milk[i - lookback_window:i])
y.append(milk[i]) | _____no_output_____ | MIT | TCN_TimeSeries_Approach.ipynb | ashishpatel26/tcn-keras-Examples |
Generate Array of list x and y | x = np.array(x)
y = np.array(y)
print(x.shape)
print(y.shape) | (156, 12, 1)
(156, 1)
| MIT | TCN_TimeSeries_Approach.ipynb | ashishpatel26/tcn-keras-Examples |
Model Design | i = Input(shape=(lookback_window, 1))
m = TCN()(i)
m = Dense(1, activation='linear')(m)
model = Model(inputs=[i], outputs=[m])
model.summary()
model.compile('adam','mae') | _____no_output_____ | MIT | TCN_TimeSeries_Approach.ipynb | ashishpatel26/tcn-keras-Examples |
Time for Model training... | print('Train...')
model.fit(x, y, epochs=100) | Train...
Epoch 1/100
5/5 [==============================] - 0s 25ms/step - loss: 271.5721
Epoch 2/100
5/5 [==============================] - 0s 26ms/step - loss: 202.3601
Epoch 3/100
5/5 [==============================] - 0s 23ms/step - loss: 129.1283
Epoch 4/100
5/5 [==============================] - 0s 25ms/step - loss: 119.5586
Epoch 5/100
5/5 [==============================] - 0s 25ms/step - loss: 82.7962
Epoch 6/100
5/5 [==============================] - 0s 23ms/step - loss: 29.5778
Epoch 7/100
5/5 [==============================] - 0s 23ms/step - loss: 26.9950
Epoch 8/100
5/5 [==============================] - 0s 25ms/step - loss: 23.8581
Epoch 9/100
5/5 [==============================] - 0s 23ms/step - loss: 27.1797
Epoch 10/100
5/5 [==============================] - 0s 24ms/step - loss: 37.1005
Epoch 11/100
5/5 [==============================] - 0s 27ms/step - loss: 38.2245
Epoch 12/100
5/5 [==============================] - 0s 23ms/step - loss: 27.9166
Epoch 13/100
5/5 [==============================] - 0s 27ms/step - loss: 49.9146
Epoch 14/100
5/5 [==============================] - 0s 24ms/step - loss: 31.0408
Epoch 15/100
5/5 [==============================] - 0s 25ms/step - loss: 23.8059
Epoch 16/100
5/5 [==============================] - 0s 24ms/step - loss: 18.7888
Epoch 17/100
5/5 [==============================] - 0s 24ms/step - loss: 25.1425
Epoch 18/100
5/5 [==============================] - 0s 25ms/step - loss: 32.6869
Epoch 19/100
5/5 [==============================] - 0s 23ms/step - loss: 31.6958
Epoch 20/100
5/5 [==============================] - 0s 23ms/step - loss: 31.2011
Epoch 21/100
5/5 [==============================] - 0s 23ms/step - loss: 22.9983
Epoch 22/100
5/5 [==============================] - 0s 25ms/step - loss: 27.7421
Epoch 23/100
5/5 [==============================] - 0s 24ms/step - loss: 29.9864
Epoch 24/100
5/5 [==============================] - 0s 23ms/step - loss: 68.7743
Epoch 25/100
5/5 [==============================] - 0s 23ms/step - loss: 41.1511
Epoch 26/100
5/5 [==============================] - 0s 24ms/step - loss: 35.2367
Epoch 27/100
5/5 [==============================] - 0s 23ms/step - loss: 66.6526
Epoch 28/100
5/5 [==============================] - 0s 24ms/step - loss: 40.3432
Epoch 29/100
5/5 [==============================] - 0s 23ms/step - loss: 50.4716
Epoch 30/100
5/5 [==============================] - 0s 25ms/step - loss: 48.0320
Epoch 31/100
5/5 [==============================] - 0s 24ms/step - loss: 34.7563
Epoch 32/100
5/5 [==============================] - 0s 24ms/step - loss: 27.2386
Epoch 33/100
5/5 [==============================] - 0s 24ms/step - loss: 32.2853
Epoch 34/100
5/5 [==============================] - 0s 25ms/step - loss: 26.1964
Epoch 35/100
5/5 [==============================] - 0s 25ms/step - loss: 22.9360
Epoch 36/100
5/5 [==============================] - 0s 23ms/step - loss: 25.1416
Epoch 37/100
5/5 [==============================] - 0s 24ms/step - loss: 30.5617
Epoch 38/100
5/5 [==============================] - 0s 25ms/step - loss: 24.7003
Epoch 39/100
5/5 [==============================] - 0s 23ms/step - loss: 14.7676
Epoch 40/100
5/5 [==============================] - 0s 24ms/step - loss: 14.6580
Epoch 41/100
5/5 [==============================] - 0s 24ms/step - loss: 12.4486
Epoch 42/100
5/5 [==============================] - 0s 24ms/step - loss: 15.5033
Epoch 43/100
5/5 [==============================] - 0s 23ms/step - loss: 20.4509
Epoch 44/100
5/5 [==============================] - 0s 23ms/step - loss: 27.3433
Epoch 45/100
5/5 [==============================] - 0s 23ms/step - loss: 25.6425
Epoch 46/100
5/5 [==============================] - 0s 24ms/step - loss: 23.2722
Epoch 47/100
5/5 [==============================] - 0s 24ms/step - loss: 19.9241
Epoch 48/100
5/5 [==============================] - 0s 23ms/step - loss: 26.6964
Epoch 49/100
5/5 [==============================] - 0s 23ms/step - loss: 51.5725
Epoch 50/100
5/5 [==============================] - 0s 24ms/step - loss: 46.1796
Epoch 51/100
5/5 [==============================] - 0s 24ms/step - loss: 43.5478
Epoch 52/100
5/5 [==============================] - 0s 23ms/step - loss: 48.7085
Epoch 53/100
5/5 [==============================] - 0s 23ms/step - loss: 46.6810
Epoch 54/100
5/5 [==============================] - 0s 23ms/step - loss: 37.4476
Epoch 55/100
5/5 [==============================] - 0s 24ms/step - loss: 40.0287
Epoch 56/100
5/5 [==============================] - 0s 25ms/step - loss: 31.3757
Epoch 57/100
5/5 [==============================] - 0s 23ms/step - loss: 23.3440
Epoch 58/100
5/5 [==============================] - 0s 24ms/step - loss: 25.6717
Epoch 59/100
5/5 [==============================] - 0s 23ms/step - loss: 22.8951
Epoch 60/100
5/5 [==============================] - 0s 24ms/step - loss: 16.8260
Epoch 61/100
5/5 [==============================] - 0s 23ms/step - loss: 19.1047
Epoch 62/100
5/5 [==============================] - 0s 23ms/step - loss: 21.6529
Epoch 63/100
5/5 [==============================] - 0s 23ms/step - loss: 16.5754
Epoch 64/100
5/5 [==============================] - 0s 24ms/step - loss: 12.2943
Epoch 65/100
5/5 [==============================] - 0s 25ms/step - loss: 14.4700
Epoch 66/100
5/5 [==============================] - 0s 24ms/step - loss: 12.4727
Epoch 67/100
5/5 [==============================] - 0s 24ms/step - loss: 12.2146
Epoch 68/100
5/5 [==============================] - 0s 23ms/step - loss: 24.3314
Epoch 69/100
5/5 [==============================] - 0s 23ms/step - loss: 23.2591
Epoch 70/100
5/5 [==============================] - 0s 23ms/step - loss: 21.5991
Epoch 71/100
5/5 [==============================] - 0s 24ms/step - loss: 34.6669
Epoch 72/100
5/5 [==============================] - 0s 24ms/step - loss: 58.7299
Epoch 73/100
5/5 [==============================] - 0s 24ms/step - loss: 30.8628
Epoch 74/100
5/5 [==============================] - 0s 23ms/step - loss: 36.4043
Epoch 75/100
5/5 [==============================] - 0s 23ms/step - loss: 45.6252
Epoch 76/100
5/5 [==============================] - 0s 24ms/step - loss: 29.5786
Epoch 77/100
5/5 [==============================] - 0s 23ms/step - loss: 16.1575
Epoch 78/100
5/5 [==============================] - 0s 24ms/step - loss: 24.9460
Epoch 79/100
5/5 [==============================] - 0s 23ms/step - loss: 17.0462
Epoch 80/100
5/5 [==============================] - 0s 23ms/step - loss: 13.5069
Epoch 81/100
5/5 [==============================] - 0s 24ms/step - loss: 14.0239
Epoch 82/100
5/5 [==============================] - 0s 23ms/step - loss: 12.9607
Epoch 83/100
5/5 [==============================] - 0s 23ms/step - loss: 13.6181
Epoch 84/100
5/5 [==============================] - 0s 23ms/step - loss: 12.3904
Epoch 85/100
5/5 [==============================] - 0s 26ms/step - loss: 13.5270
Epoch 86/100
5/5 [==============================] - 0s 23ms/step - loss: 18.3469
Epoch 87/100
5/5 [==============================] - 0s 24ms/step - loss: 20.0561
Epoch 88/100
5/5 [==============================] - 0s 22ms/step - loss: 19.5092
Epoch 89/100
5/5 [==============================] - 0s 23ms/step - loss: 22.7791
Epoch 90/100
5/5 [==============================] - 0s 22ms/step - loss: 20.7960
Epoch 91/100
5/5 [==============================] - 0s 23ms/step - loss: 26.3365
Epoch 92/100
5/5 [==============================] - 0s 24ms/step - loss: 21.1719
Epoch 93/100
5/5 [==============================] - 0s 22ms/step - loss: 19.5793
Epoch 94/100
5/5 [==============================] - 0s 23ms/step - loss: 22.3828
Epoch 95/100
5/5 [==============================] - 0s 23ms/step - loss: 20.0138
Epoch 96/100
5/5 [==============================] - 0s 23ms/step - loss: 26.0298
Epoch 97/100
5/5 [==============================] - 0s 23ms/step - loss: 15.8576
Epoch 98/100
5/5 [==============================] - 0s 23ms/step - loss: 26.3697
Epoch 99/100
5/5 [==============================] - 0s 24ms/step - loss: 14.8465
Epoch 100/100
5/5 [==============================] - 0s 22ms/step - loss: 13.5129
| MIT | TCN_TimeSeries_Approach.ipynb | ashishpatel26/tcn-keras-Examples |
Prediction with TCN Model | predict = model.predict(x) | _____no_output_____ | MIT | TCN_TimeSeries_Approach.ipynb | ashishpatel26/tcn-keras-Examples |
Plot the Result | plt.style.use("fivethirtyeight")
plt.figure(figsize = (15,7))
plt.plot(predict)
plt.plot(y)
plt.title('Monthly Milk Production (in pounds)')
plt.legend(['predicted', 'actual'])
plt.xlabel("Moths Counts")
plt.ylabel("Milk Production in Pounds")
plt.show() | _____no_output_____ | MIT | TCN_TimeSeries_Approach.ipynb | ashishpatel26/tcn-keras-Examples |
synchro.extracting> Function to extract data of an experiment from 3rd party programs To align timeseries of an experiment, we need to read logs and import data produced by 3rd party softwares used during the experiment. It includes:* QDSpy logging* Numpy arrays of the stimuli* SpykingCircus spike sorting refined with Phy* Eye tracking results from MaskRCNN | #export
import numpy as np
import datetime
import os, glob
import csv
import re
from theonerig.synchro.io import *
from theonerig.utils import *
def get_QDSpy_logs(log_dir):
"""Factory function to generate QDSpy_log objects from all the QDSpy logs of the folder `log_dir`"""
log_names = glob.glob(os.path.join(log_dir,'[0-9]*.log'))
qdspy_logs = [QDSpy_log(log_name) for log_name in log_names]
for qdspy_log in qdspy_logs:
qdspy_log.find_stimuli()
return qdspy_logs
class QDSpy_log:
"""Class defining a QDSpy log.
It reads the log it represent and extract the stimuli information from it:
- Start and end time
- Parameters like the md5 key
- Frame delays
"""
def __init__(self, log_path):
self.log_path = log_path
self.stimuli = []
self.comments = []
def _extract_data(self, data_line):
data = data_line[data_line.find('{')+1:data_line.find('}')]
data_splitted = data.split(',')
data_dict = {}
for data in data_splitted:
ind = data.find("'")
if type(data[data.find(":")+2:]) is str:
data_dict[data[ind+1:data.find("'",ind+1)]] = data[data.find(":")+2:][1:-1]
else:
data_dict[data[ind+1:data.find("'",ind+1)]] = data[data.find(":")+2:]
return data_dict
def _extract_time(self,data_line):
return datetime.datetime.strptime(data_line.split()[0], '%Y%m%d_%H%M%S')
def _extract_delay(self,data_line):
ind = data_line.find('#')
index_frame = int(data_line[ind+1:data_line.find(' ',ind)])
ind = data_line.find('was')
delay = float(data_line[ind:].split(" ")[1])
return (index_frame, delay)
def __repr__(self):
return "\n".join([str(stim) for stim in self.stimuli])
@property
def n_stim(self):
return len(self.stimuli)
@property
def stim_names(self):
return [stim.name for stim in self.stimuli]
def find_stimuli(self):
"""Find the stimuli in the log file and return the list of the stimuli
found by this object."""
with open(self.log_path, 'r', encoding="ISO-8859-1") as log_file:
for line in log_file:
if "DATA" in line:
data_juice = self._extract_data(line)
if 'stimState' in data_juice.keys():
if data_juice['stimState'] == "STARTED" :
curr_stim = Stimulus(self._extract_time(line))
curr_stim.set_parameters(data_juice)
self.stimuli.append(curr_stim)
stimulus_ON = True
elif data_juice['stimState'] == "FINISHED" or data_juice['stimState'] == "ABORTED":
curr_stim.is_aborted = data_juice['stimState'] == "ABORTED"
curr_stim.stop_time = self._extract_time(line)
stimulus_ON = False
elif 'userComment' in data_juice.keys():
pass
#print("userComment, use it to bind logs to records")
elif stimulus_ON: #Information on stimulus parameters
curr_stim.set_parameters(data_juice)
# elif 'probeX' in data_juice.keys():
# print("Probe center not implemented yet")
if "WARNING" in line and "dt of frame" and stimulus_ON:
curr_stim.frame_delay.append(self._extract_delay(line))
if curr_stim.frame_delay[-1][1] > 2000/60: #if longer than 2 frames could be bad
print(curr_stim.name, " ".join(line.split()[1:])[:-1])
return self.stimuli
class Stimulus:
"""Stimulus object containing information about it's presentation.
- start_time : a datetime object)
- stop_time : a datetime object)
- parameters : Parameters extracted from the QDSpy
- md5 : The md5 hash of that compiled version of the stimulus
- name : The name of the stimulus
"""
def __init__(self,start):
self.start_time = start
self.stop_time = None
self.parameters = {}
self.md5 = None
self.name = "NoName"
self.frame_delay = []
self.is_aborted = False
def set_parameters(self, parameters):
self.parameters.update(parameters)
if "_sName" in parameters.keys():
self.name = parameters["_sName"]
if "stimMD5" in parameters.keys():
self.md5 = parameters["stimMD5"]
def __str__(self):
return "%s %s at %s" %(self.name+" "*(24-len(self.name)),self.md5,self.start_time)
def __repr__(self):
return self.__str__() | _____no_output_____ | Apache-2.0 | 11_synchro.extracting.ipynb | Ines-Filipa/theonerig |
To read QDSpy logs of your experiment, simply provide the folder containing the log you want to read to `get_QDSpy_logs` | #logs = get_QDSpy_logs("./files/basic_synchro") | flickering_bars_pr WARNING dt of frame #15864 was 50.315 m
flickering_bars_pr WARNING dt of frame #19477 was 137.235 m
| Apache-2.0 | 11_synchro.extracting.ipynb | Ines-Filipa/theonerig |
It returns a list fo the QDSpy logs. Stimuli are contained in a list inside each log: | #logs[0].stimuli | _____no_output_____ | Apache-2.0 | 11_synchro.extracting.ipynb | Ines-Filipa/theonerig |
The stimuli objects contains informations on how their display went: | # stim = logs[0].stimuli[5]
# print(stim.name, stim.start_time, stim.frame_delay, stim.md5)
#export
def unpack_stim_npy(npy_dir, md5_hash):
"""Find the stimuli of a given hash key in the npy stimulus folder. The stimuli are in a compressed version
comprising three files. inten for the stimulus values on the screen, marker for the values of the marker
read by a photodiode to get the stimulus timing during a record, and an optional shader that is used to
specify informations about a shader when used, like for the moving gratings."""
#Stimuli can be either npy or npz (useful when working remotely)
def find_file(ftype):
flist = glob.glob(os.path.join(npy_dir, "*_"+ftype+"_"+md5_hash+".npy"))
if len(flist)==0:
flist = glob.glob(os.path.join(npy_dir, "*_"+ftype+"_"+md5_hash+".npz"))
res = np.load(flist[0])["arr_0"]
else:
res = np.load(flist[0])
return res
inten = find_file("intensities")
marker = find_file("marker")
shader, unpack_shader = None, None
if len(glob.glob(os.path.join(npy_dir, "*_shader_"+md5_hash+".np*")))>0:
shader = find_file("shader")
unpack_shader = np.empty((np.sum(marker[:,0]), *shader.shape[1:]))
#The latter unpacks the arrays
unpack_inten = np.empty((np.sum(marker[:,0]), *inten.shape[1:]))
unpack_marker = np.empty(np.sum(marker[:,0]))
cursor = 0
for i, n_frame in enumerate(marker[:,0]):
unpack_inten[cursor:cursor+n_frame] = inten[i]
unpack_marker[cursor:cursor+n_frame] = marker[i, 1]
if shader is not None:
unpack_shader[cursor:cursor+n_frame] = shader[i]
cursor += n_frame
return unpack_inten, unpack_marker, unpack_shader
# logs = get_QDSpy_logs("./files/basic_synchro") | flickering_bars_pr WARNING dt of frame #15864 was 50.315 m
flickering_bars_pr WARNING dt of frame #19477 was 137.235 m
| Apache-2.0 | 11_synchro.extracting.ipynb | Ines-Filipa/theonerig |
To unpack the stimulus values, provide the folder of the numpy arrays and the hash of the stimulus: | # unpacked = unpack_stim_npy("./files/basic_synchro/stimulus_data", "eed21bda540934a428e93897908d049e") | _____no_output_____ | Apache-2.0 | 11_synchro.extracting.ipynb | Ines-Filipa/theonerig |
Unpacked is a tuple, where the first element is the intensity of shape (n_frames, n_colors, y, x) | # unpacked[0].shape | _____no_output_____ | Apache-2.0 | 11_synchro.extracting.ipynb | Ines-Filipa/theonerig |
The second element of the tuple repesents the marker values for the timing. QDSpy defaults are zero and ones, but I used custom red squares taking intensities [50,100,150,200,250] to time with five different signals | # unpacked[1][:50] | _____no_output_____ | Apache-2.0 | 11_synchro.extracting.ipynb | Ines-Filipa/theonerig |
Each stimulus is also starting with a barcode, of the form:0 0 0 0 0 0 4 0 4\*[1-4] 0 4\*[1-4] 0 4\*[1-4] 0 4\*[1-4] 0 4 0 0 0 0 0 0 and ends with 0 0 0 0 0 0 | #export
def extract_spyking_circus_results(dir_, record_basename):
"""Extract the good cells of a record. Overlap with phy_results_dict."""
phy_dir = os.path.join(dir_,record_basename+"/"+record_basename+".GUI")
phy_dict = phy_results_dict(phy_dir)
good_clusters = []
with open(os.path.join(phy_dir,'cluster_group.tsv'), 'r') as tsvfile:
spamreader = csv.reader(tsvfile, delimiter='\t', quotechar='|')
for i,row in enumerate(spamreader):
if row[1] == "good":
good_clusters.append(int(row[0]))
good_clusters = np.array(good_clusters)
phy_dict["good_clusters"] = good_clusters
return phy_dict
#export
def extract_best_pupil(fn):
"""From results of MaskRCNN, go over all or None pupil detected and select the best pupil.
Each pupil returned is (x,y,width,height,angle,probability)"""
pupil = np.load(fn, allow_pickle=True)
filtered_pupil = np.empty((len(pupil), 6))
for i, detected in enumerate(pupil):
if len(detected)>0:
best = detected[0]
for detect in detected[1:]:
if detect[5]>best[5]:
best = detect
filtered_pupil[i] = np.array(best)
else:
filtered_pupil[i] = np.array([0,0,0,0,0,0])
return filtered_pupil
#export
def stack_len_extraction(stack_info_dir):
"""Extract from ImageJ macro directives the size of the stacks acquired."""
ptrn_nFrame = r".*number=(\d*) .*"
l_epochs = []
for fn in glob.glob(os.path.join(stack_info_dir, "*.txt")):
with open(fn) as f:
line = f.readline()
l_epochs.append(int(re.findall(ptrn_nFrame, line)[0]))
return l_epochs
#hide
from nbdev.export import *
notebook2script() | Converted 00_core.ipynb.
Converted 01_utils.ipynb.
Converted 02_processing.ipynb.
Converted 03_modelling.ipynb.
Converted 04_plotting.ipynb.
Converted 05_database.ipynb.
Converted 10_synchro.io.ipynb.
Converted 11_synchro.extracting.ipynb.
Converted 12_synchro.processing.ipynb.
Converted 99_testdata.ipynb.
Converted index.ipynb.
| Apache-2.0 | 11_synchro.extracting.ipynb | Ines-Filipa/theonerig |
Training a Plant Disease Diagnosis Model with PlantVillage Dataset | import numpy as np
import os
import matplotlib.pyplot as plt
from skimage.io import imread
from sklearn.metrics import classification_report, confusion_matrix
from sklearn .model_selection import train_test_split
import keras
import keras.backend as K
from keras.preprocessing.image import load_img, img_to_array, ImageDataGenerator
from keras.utils.np_utils import to_categorical
from keras import layers
from keras.models import Sequential, Model
from keras.callbacks import EarlyStopping, ModelCheckpoint | _____no_output_____ | Net-SNMP | notebooks/PlantDisease_tutorial.ipynb | Julia2505/deep_learning_for_biologists |
Preparation Data Preparation | !apt-get install subversion > /dev/null
#Retreive specifc diseases of tomato for training
!svn export https://github.com/spMohanty/PlantVillage-Dataset/trunk/raw/color/Tomato___Bacterial_spot image/Tomato___Bacterial_spot > /dev/null
!svn export https://github.com/spMohanty/PlantVillage-Dataset/trunk/raw/color/Tomato___Early_blight image/Tomato___Early_blight > /dev/null
!svn export https://github.com/spMohanty/PlantVillage-Dataset/trunk/raw/color/Tomato___Late_blight image/Tomato___Late_blight > /dev/null
!svn export https://github.com/spMohanty/PlantVillage-Dataset/trunk/raw/color/Tomato___Septoria_leaf_spot image/Tomato___Septoria_leaf_spot > /dev/null
!svn export https://github.com/spMohanty/PlantVillage-Dataset/trunk/raw/color/Tomato___Target_Spot image/Tomato___Target_Spot > /dev/null
!svn export https://github.com/spMohanty/PlantVillage-Dataset/trunk/raw/color/Tomato___healthy image/Tomato___healthy > /dev/null
#folder structure
!ls image
plt.figure(figsize=(15,10))
#visualize several images
parent_directory = "image"
for i, folder in enumerate(os.listdir(parent_directory)):
print(folder)
folder_directory = os.path.join(parent_directory,folder)
files = os.listdir(folder_directory)
#will inspect only 1 image per folder
file = files[0]
file_path = os.path.join(folder_directory,file)
image = imread(file_path)
plt.subplot(1,6,i+1)
plt.imshow(image)
plt.axis("off")
name = folder.split("___")[1][:-1]
plt.title(name)
#plt.show()
#load everything into memory
x = []
y = []
class_names = []
parent_directory = "image"
for i,folder in enumerate(os.listdir(parent_directory)):
print(i,folder)
class_names.append(folder)
folder_directory = os.path.join(parent_directory,folder)
files = os.listdir(folder_directory)
#will inspect only 1 image per folder
for file in files:
file_path = os.path.join(folder_directory,file)
image = load_img(file_path,target_size=(64,64))
image = img_to_array(image)/255.
x.append(image)
y.append(i)
x = np.array(x)
y = to_categorical(y)
#check the data shape
print(x.shape)
print(y.shape)
print(y[0])
x_train, _x, y_train, _y = train_test_split(x,y,test_size=0.2, stratify = y, random_state = 1)
x_valid,x_test, y_valid, y_test = train_test_split(_x,_y,test_size=0.4, stratify = _y, random_state = 1)
print("train data:",x_train.shape,y_train.shape)
print("validation data:",x_valid.shape,y_valid.shape)
print("test data:",x_test.shape,y_test.shape)
| _____no_output_____ | Net-SNMP | notebooks/PlantDisease_tutorial.ipynb | Julia2505/deep_learning_for_biologists |
Model Preparation | K.clear_session()
nfilter = 32
#VGG16 like model
model = Sequential([
#block1
layers.Conv2D(nfilter,(3,3),padding="same",name="block1_conv1",input_shape=(64,64,3)),
layers.Activation("relu"),
layers.BatchNormalization(),
#layers.Dropout(rate=0.2),
layers.Conv2D(nfilter,(3,3),padding="same",name="block1_conv2"),
layers.BatchNormalization(),
layers.Activation("relu"),
#layers.Dropout(rate=0.2),
layers.MaxPooling2D((2,2),strides=(2,2),name="block1_pool"),
#block2
layers.Conv2D(nfilter*2,(3,3),padding="same",name="block2_conv1"),
layers.BatchNormalization(),
layers.Activation("relu"),
#layers.Dropout(rate=0.2),
layers.Conv2D(nfilter*2,(3,3),padding="same",name="block2_conv2"),
layers.BatchNormalization(),
layers.Activation("relu"),
#layers.Dropout(rate=0.2),
layers.MaxPooling2D((2,2),strides=(2,2),name="block2_pool"),
#block3
layers.Conv2D(nfilter*2,(3,3),padding="same",name="block3_conv1"),
layers.BatchNormalization(),
layers.Activation("relu"),
#layers.Dropout(rate=0.2),
layers.Conv2D(nfilter*4,(3,3),padding="same",name="block3_conv2"),
layers.BatchNormalization(),
layers.Activation("relu"),
#layers.Dropout(rate=0.2),
layers.Conv2D(nfilter*4,(3,3),padding="same",name="block3_conv3"),
layers.BatchNormalization(),
layers.Activation("relu"),
#layers.Dropout(rate=0.2),
layers.MaxPooling2D((2,2),strides=(2,2),name="block3_pool"),
#layers.Flatten(),
layers.GlobalAveragePooling2D(),
#inference layer
layers.Dense(128,name="fc1"),
layers.BatchNormalization(),
layers.Activation("relu"),
#layers.Dropout(rate=0.2),
layers.Dense(128,name="fc2"),
layers.BatchNormalization(),
layers.Activation("relu"),
#layers.Dropout(rate=0.2),
layers.Dense(6,name="prepredictions"),
layers.Activation("softmax",name="predictions")
])
model.compile(optimizer = "adam", loss="categorical_crossentropy", metrics=["accuracy"])
model.summary() | _____no_output_____ | Net-SNMP | notebooks/PlantDisease_tutorial.ipynb | Julia2505/deep_learning_for_biologists |
Training | #utilize early stopping function to stop at the lowest validation loss
es = EarlyStopping(monitor='val_loss', patience=10, verbose=1, mode='auto')
#utilize save best weight model during training
ckpt = ModelCheckpoint("PlantDiseaseCNNmodel.hdf5", monitor='val_loss', verbose=1, save_best_only=True, save_weights_only=False, mode='auto', period=1)
#we will define a generator class for training data and validation data seperately, as no augmentation is not required for validation data
t_gen = ImageDataGenerator(rotation_range=90,horizontal_flip=True)
v_gen = ImageDataGenerator()
train_gen = t_gen.flow(x_train,y_train,batch_size=98)
valid_gen = v_gen.flow(x_valid,y_valid,batch_size=98)
history = model.fit_generator(
train_gen,
steps_per_epoch = train_gen.n // 98,
callbacks = [es,ckpt],
validation_data = valid_gen,
validation_steps = valid_gen.n // 98,
epochs=50) | _____no_output_____ | Net-SNMP | notebooks/PlantDisease_tutorial.ipynb | Julia2505/deep_learning_for_biologists |
Evaluation | #load the model weight file with lowest validation loss
model.load_weights("PlantDiseaseCNNmodel.hdf5")
#or can obtain the pretrained model from the github repo.
#check the model metrics
print(model.metrics_names)
#evaluate training data
print(model.evaluate(x= x_train, y = y_train))
#evaluate validation data
print(model.evaluate(x= x_valid, y = y_valid))
#evaluate test data
print(model.evaluate(x= x_test, y = y_test))
#draw a confusion matrix
#true label
y_true = np.argmax(y_test,axis=1)
#prediction label
Y_pred = model.predict(x_test)
y_pred = np.argmax(Y_pred, axis=1)
print(y_true)
print(y_pred)
#https://scikit-learn.org/stable/auto_examples/model_selection/plot_confusion_matrix.html#sphx-glr-auto-examples-model-selection-plot-confusion-matrix-py
from sklearn.metrics import confusion_matrix
from sklearn.utils.multiclass import unique_labels
def plot_confusion_matrix(y_true, y_pred, classes,
normalize=False,
title=None,
cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
if not title:
if normalize:
title = 'Normalized confusion matrix'
else:
title = 'Confusion matrix, without normalization'
# Compute confusion matrix
cm = confusion_matrix(y_true, y_pred)
# Only use the labels that appear in the data
#classes = classes[unique_labels(y_true, y_pred)]
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Normalized confusion matrix")
else:
print('Confusion matrix, without normalization')
print(cm)
fig, ax = plt.subplots(figsize=(5,5))
im = ax.imshow(cm, interpolation='nearest', cmap=cmap)
#ax.figure.colorbar(im, ax=ax)
# We want to show all ticks...
ax.set(xticks=np.arange(cm.shape[1]),
yticks=np.arange(cm.shape[0]),
# ... and label them with the respective list entries
xticklabels=classes, yticklabels=classes,
title=title,
ylabel='True label',
xlabel='Predicted label')
# Rotate the tick labels and set their alignment.
plt.setp(ax.get_xticklabels(), rotation=45, ha="right",
rotation_mode="anchor")
# Loop over data dimensions and create text annotations.
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i in range(cm.shape[0]):
for j in range(cm.shape[1]):
ax.text(j, i, format(cm[i, j], fmt),
ha="center", va="center",
color="white" if cm[i, j] > thresh else "black")
fig.tight_layout()
return ax
np.set_printoptions(precision=2)
plot_confusion_matrix(y_true, y_pred, classes=class_names, normalize=True,
title='Normalized confusion matrix')
| _____no_output_____ | Net-SNMP | notebooks/PlantDisease_tutorial.ipynb | Julia2505/deep_learning_for_biologists |
Predicting Indivisual Images | n = 15 #do not exceed (number of test image - 1)
plt.imshow(x_test[n])
plt.show()
true_label = np.argmax(y_test,axis=1)[n]
print("true_label is:",true_label,":",class_names[true_label])
prediction = model.predict(x_test[n][np.newaxis,...])[0]
print("predicted_value is:",prediction)
predicted_label = np.argmax(prediction)
print("predicted_label is:",predicted_label,":",class_names[predicted_label])
if true_label == predicted_label:
print("correct prediction")
else:
print("wrong prediction") | _____no_output_____ | Net-SNMP | notebooks/PlantDisease_tutorial.ipynb | Julia2505/deep_learning_for_biologists |
LeNet Lab SolutionSource: Yan LeCun Load DataLoad the MNIST data, which comes pre-loaded with TensorFlow.You do not need to modify this section. | from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets("MNIST_data/", reshape=False)
X_train, y_train = mnist.train.images, mnist.train.labels
X_validation, y_validation = mnist.validation.images, mnist.validation.labels
X_test, y_test = mnist.test.images, mnist.test.labels
assert(len(X_train) == len(y_train))
assert(len(X_validation) == len(y_validation))
assert(len(X_test) == len(y_test))
print()
print("Image Shape: {}".format(X_train[0].shape))
print()
print("Training Set: {} samples".format(len(X_train)))
print("Validation Set: {} samples".format(len(X_validation)))
print("Test Set: {} samples".format(len(X_test))) | _____no_output_____ | MIT | LeNet-Lab-Solution.ipynb | LiYan1988/LeNet-2 |
The MNIST data that TensorFlow pre-loads comes as 28x28x1 images.However, the LeNet architecture only accepts 32x32xC images, where C is the number of color channels.In order to reformat the MNIST data into a shape that LeNet will accept, we pad the data with two rows of zeros on the top and bottom, and two columns of zeros on the left and right (28+2+2 = 32).You do not need to modify this section. | import numpy as np
# Pad images with 0s
X_train = np.pad(X_train, ((0,0),(2,2),(2,2),(0,0)), 'constant')
X_validation = np.pad(X_validation, ((0,0),(2,2),(2,2),(0,0)), 'constant')
X_test = np.pad(X_test, ((0,0),(2,2),(2,2),(0,0)), 'constant')
print("Updated Image Shape: {}".format(X_train[0].shape)) | _____no_output_____ | MIT | LeNet-Lab-Solution.ipynb | LiYan1988/LeNet-2 |
Visualize DataView a sample from the dataset.You do not need to modify this section. | import random
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
index = random.randint(0, len(X_train))
image = X_train[index].squeeze()
plt.figure(figsize=(1,1))
plt.imshow(image, cmap="gray")
print(y_train[index]) | _____no_output_____ | MIT | LeNet-Lab-Solution.ipynb | LiYan1988/LeNet-2 |
Preprocess DataShuffle the training data.You do not need to modify this section. | from sklearn.utils import shuffle
X_train, y_train = shuffle(X_train, y_train) | _____no_output_____ | MIT | LeNet-Lab-Solution.ipynb | LiYan1988/LeNet-2 |
Setup TensorFlowThe `EPOCH` and `BATCH_SIZE` values affect the training speed and model accuracy.You do not need to modify this section. | import tensorflow as tf
EPOCHS = 10
BATCH_SIZE = 128 | _____no_output_____ | MIT | LeNet-Lab-Solution.ipynb | LiYan1988/LeNet-2 |
SOLUTION: Implement LeNet-5Implement the [LeNet-5](http://yann.lecun.com/exdb/lenet/) neural network architecture.This is the only cell you need to edit. InputThe LeNet architecture accepts a 32x32xC image as input, where C is the number of color channels. Since MNIST images are grayscale, C is 1 in this case. Architecture**Layer 1: Convolutional.** The output shape should be 28x28x6.**Activation.** Your choice of activation function.**Pooling.** The output shape should be 14x14x6.**Layer 2: Convolutional.** The output shape should be 10x10x16.**Activation.** Your choice of activation function.**Pooling.** The output shape should be 5x5x16.**Flatten.** Flatten the output shape of the final pooling layer such that it's 1D instead of 3D. The easiest way to do is by using `tf.contrib.layers.flatten`, which is already imported for you.**Layer 3: Fully Connected.** This should have 120 outputs.**Activation.** Your choice of activation function.**Layer 4: Fully Connected.** This should have 84 outputs.**Activation.** Your choice of activation function.**Layer 5: Fully Connected (Logits).** This should have 10 outputs. OutputReturn the result of the 2nd fully connected layer. | from tensorflow.contrib.layers import flatten
def LeNet(x):
# Arguments used for tf.truncated_normal, randomly defines variables for the weights and biases for each layer
mu = 0
sigma = 0.1
# SOLUTION: Layer 1: Convolutional. Input = 32x32x1. Output = 28x28x6.
conv1_W = tf.Variable(tf.truncated_normal(shape=(5, 5, 1, 6), mean = mu, stddev = sigma))
conv1_b = tf.Variable(tf.zeros(6))
conv1 = tf.nn.conv2d(x, conv1_W, strides=[1, 1, 1, 1], padding='VALID') + conv1_b
# SOLUTION: Activation.
conv1 = tf.nn.relu(conv1)
# SOLUTION: Pooling. Input = 28x28x6. Output = 14x14x6.
conv1 = tf.nn.max_pool(conv1, ksize=[1, 2, 2, 1], strides=[1, 2, 2, 1], padding='VALID')
# SOLUTION: Layer 2: Convolutional. Output = 10x10x16.
conv2_W = tf.Variable(tf.truncated_normal(shape=(5, 5, 6, 16), mean = mu, stddev = sigma))
conv2_b = tf.Variable(tf.zeros(16))
conv2 = tf.nn.conv2d(conv1, conv2_W, strides=[1, 1, 1, 1], padding='VALID') + conv2_b
# SOLUTION: Activation.
conv2 = tf.nn.relu(conv2)
# SOLUTION: Pooling. Input = 10x10x16. Output = 5x5x16.
conv2 = tf.nn.max_pool(conv2, ksize=[1, 2, 2, 1], strides=[1, 2, 2, 1], padding='VALID')
# SOLUTION: Flatten. Input = 5x5x16. Output = 400.
fc0 = flatten(conv2)
# SOLUTION: Layer 3: Fully Connected. Input = 400. Output = 120.
fc1_W = tf.Variable(tf.truncated_normal(shape=(400, 120), mean = mu, stddev = sigma))
fc1_b = tf.Variable(tf.zeros(120))
fc1 = tf.matmul(fc0, fc1_W) + fc1_b
# SOLUTION: Activation.
fc1 = tf.nn.relu(fc1)
# SOLUTION: Layer 4: Fully Connected. Input = 120. Output = 84.
fc2_W = tf.Variable(tf.truncated_normal(shape=(120, 84), mean = mu, stddev = sigma))
fc2_b = tf.Variable(tf.zeros(84))
fc2 = tf.matmul(fc1, fc2_W) + fc2_b
# SOLUTION: Activation.
fc2 = tf.nn.relu(fc2)
# SOLUTION: Layer 5: Fully Connected. Input = 84. Output = 10.
fc3_W = tf.Variable(tf.truncated_normal(shape=(84, 10), mean = mu, stddev = sigma))
fc3_b = tf.Variable(tf.zeros(10))
logits = tf.matmul(fc2, fc3_W) + fc3_b
return logits | _____no_output_____ | MIT | LeNet-Lab-Solution.ipynb | LiYan1988/LeNet-2 |
Features and LabelsTrain LeNet to classify [MNIST](http://yann.lecun.com/exdb/mnist/) data.`x` is a placeholder for a batch of input images.`y` is a placeholder for a batch of output labels.You do not need to modify this section. | x = tf.placeholder(tf.float32, (None, 32, 32, 1))
y = tf.placeholder(tf.int32, (None))
one_hot_y = tf.one_hot(y, 10) | _____no_output_____ | MIT | LeNet-Lab-Solution.ipynb | LiYan1988/LeNet-2 |
Training PipelineCreate a training pipeline that uses the model to classify MNIST data.You do not need to modify this section. | rate = 0.001
logits = LeNet(x)
cross_entropy = tf.nn.softmax_cross_entropy_with_logits(labels=one_hot_y, logits=logits)
loss_operation = tf.reduce_mean(cross_entropy)
optimizer = tf.train.AdamOptimizer(learning_rate = rate)
training_operation = optimizer.minimize(loss_operation) | _____no_output_____ | MIT | LeNet-Lab-Solution.ipynb | LiYan1988/LeNet-2 |
Model EvaluationEvaluate how well the loss and accuracy of the model for a given dataset.You do not need to modify this section. | correct_prediction = tf.equal(tf.argmax(logits, 1), tf.argmax(one_hot_y, 1))
accuracy_operation = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
saver = tf.train.Saver()
def evaluate(X_data, y_data):
num_examples = len(X_data)
total_accuracy = 0
sess = tf.get_default_session()
for offset in range(0, num_examples, BATCH_SIZE):
batch_x, batch_y = X_data[offset:offset+BATCH_SIZE], y_data[offset:offset+BATCH_SIZE]
accuracy = sess.run(accuracy_operation, feed_dict={x: batch_x, y: batch_y})
total_accuracy += (accuracy * len(batch_x))
return total_accuracy / num_examples | _____no_output_____ | MIT | LeNet-Lab-Solution.ipynb | LiYan1988/LeNet-2 |
Train the ModelRun the training data through the training pipeline to train the model.Before each epoch, shuffle the training set.After each epoch, measure the loss and accuracy of the validation set.Save the model after training.You do not need to modify this section. | with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
num_examples = len(X_train)
print("Training...")
print()
for i in range(EPOCHS):
X_train, y_train = shuffle(X_train, y_train)
for offset in range(0, num_examples, BATCH_SIZE):
end = offset + BATCH_SIZE
batch_x, batch_y = X_train[offset:end], y_train[offset:end]
sess.run(training_operation, feed_dict={x: batch_x, y: batch_y})
validation_accuracy = evaluate(X_validation, y_validation)
print("EPOCH {} ...".format(i+1))
print("Validation Accuracy = {:.3f}".format(validation_accuracy))
print()
saver.save(sess, './lenet')
print("Model saved") | _____no_output_____ | MIT | LeNet-Lab-Solution.ipynb | LiYan1988/LeNet-2 |
Evaluate the ModelOnce you are completely satisfied with your model, evaluate the performance of the model on the test set.Be sure to only do this once!If you were to measure the performance of your trained model on the test set, then improve your model, and then measure the performance of your model on the test set again, that would invalidate your test results. You wouldn't get a true measure of how well your model would perform against real data.You do not need to modify this section. | with tf.Session() as sess:
saver.restore(sess, tf.train.latest_checkpoint('.'))
test_accuracy = evaluate(X_test, y_test)
print("Test Accuracy = {:.3f}".format(test_accuracy)) | _____no_output_____ | MIT | LeNet-Lab-Solution.ipynb | LiYan1988/LeNet-2 |
Polynomial Regression What if your data doesn't look linear at all? Let's look at some more realistic-looking page speed / purchase data: | %matplotlib inline
from pylab import *
import numpy as np
np.random.seed(2)
pageSpeeds = np.random.normal(3.0, 1.0, 1000)
purchaseAmount = np.random.normal(50.0, 10.0, 1000) / pageSpeeds
scatter(pageSpeeds, purchaseAmount) | _____no_output_____ | MIT | PolynomialRegression.ipynb | Lucian-N/DataScience |
numpy has a handy polyfit function we can use, to let us construct an nth-degree polynomial model of our data that minimizes squared error. Let's try it with a 4th degree polynomial: | x = np.array(pageSpeeds)
y = np.array(purchaseAmount)
p4 = np.poly1d(np.polyfit(x, y, 4))
| _____no_output_____ | MIT | PolynomialRegression.ipynb | Lucian-N/DataScience |
We'll visualize our original scatter plot, together with a plot of our predicted values using the polynomial for page speed times ranging from 0-7 seconds: | import matplotlib.pyplot as plt
xp = np.linspace(0, 7, 100)
plt.scatter(x, y)
plt.plot(xp, p4(xp), c='r')
plt.show() | _____no_output_____ | MIT | PolynomialRegression.ipynb | Lucian-N/DataScience |
Looks pretty good! Let's measure the r-squared error: | from sklearn.metrics import r2_score
r2 = r2_score(y, p4(x))
print(r2)
| 0.82937663963
| MIT | PolynomialRegression.ipynb | Lucian-N/DataScience |
Single-stepping the `logictools` Pattern Generator * This notebook will show how to use single-stepping mode with the pattern generator* Note that all generators in the _logictools_ library may be **single-stepped** Visually ... The _logictools_ library on the Zynq device on the PYNQ board  Demonstrator notes* For this demo, the pattern generator implements a simple, 4-bit binary, up-counter* We will single-step the clock and verify the counter operation* The output is verified using the waveforms captured by the trace analyzer Points to note* __Everything__ runs on the Zynq chip on the PYNQ board, even this slide show! * We will specify and implement circuits __using only Python code__ * __No__ Xilinx CAD tools are used * We can create live, real-time circuits __instantaneously__ | # Specify a stimulus waveform and display it
from pynq.overlays.logictools import LogicToolsOverlay
from pynq.lib.logictools import Waveform
logictools_olay = LogicToolsOverlay('logictools.bit')
up_counter_stimulus = {'signal': [
{'name': 'bit0', 'pin': 'D0', 'wave': 'lh' * 8},
{'name': 'bit1', 'pin': 'D1', 'wave': 'l.h.' * 4},
{'name': 'bit2', 'pin': 'D2', 'wave': 'l...h...' * 2},
{'name': 'bit3', 'pin': 'D3', 'wave': 'l.......h.......'}]}
# Check visually that the stimulus pattern is correct
waveform = Waveform(up_counter_stimulus)
waveform.display()
# Add the signals we want to analyze
up_counter = {'signal': [
['stimulus',
{'name': 'bit0', 'pin': 'D0', 'wave': 'lh' * 8},
{'name': 'bit1', 'pin': 'D1', 'wave': 'l.h.' * 4},
{'name': 'bit2', 'pin': 'D2', 'wave': 'l...h...' * 2},
{'name': 'bit3', 'pin': 'D3', 'wave': 'l.......h.......'}],
{},
['analysis',
{'name': 'bit0_output', 'pin': 'D0'},
{'name': 'bit1_output', 'pin': 'D1'},
{'name': 'bit2_output', 'pin': 'D2'},
{'name': 'bit3_output', 'pin': 'D3'}]]}
# Display the stimulus and analysis signal groups
waveform = Waveform(up_counter)
waveform.display()
# Configure the pattern generator and analyzer
pattern_generator = logictools_olay.pattern_generator
pattern_generator.trace(num_analyzer_samples=16)
pattern_generator.setup(up_counter,
stimulus_group_name='stimulus',
analysis_group_name='analysis')
# Press `cntrl-enter` to advance the pattern generator by one clock cycle
pattern_generator.step()
pattern_generator.show_waveform()
# Advance an arbitrary number of cycles
no_of_cycles = 7
for _ in range(no_of_cycles):
pattern_generator.step()
pattern_generator.show_waveform()
# Finally, reset the pattern generator after use
pattern_generator.reset() | _____no_output_____ | BSD-3-Clause | Session_3/3_pattern_generator.ipynb | xupsh/PYNQ_Workshop |
Practice: approximate q-learning_Reference: based on Practical RL_ [week04](https://github.com/yandexdataschool/Practical_RL/tree/master/week04_approx_rl)In this notebook you will teach a __pytorch__ neural network to do Q-learning. | # # in google colab uncomment this
# import os
# os.system('apt-get install -y xvfb')
# os.system('wget https://raw.githubusercontent.com/yandexdataschool/Practical_DL/fall18/xvfb -O ../xvfb')
# os.system('apt-get install -y python-opengl ffmpeg')
# XVFB will be launched if you run on a server
import os
if type(os.environ.get("DISPLAY")) is not str or len(os.environ.get("DISPLAY")) == 0:
!bash ../xvfb start
%env DISPLAY = : 1
import gym
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
%matplotlib inline
env = gym.make("CartPole-v0").env
env.reset()
n_actions = env.action_space.n
state_dim = env.observation_space.shape
plt.imshow(env.render("rgb_array"))
env.close() | _____no_output_____ | MIT | week10_Model_free_learning_peek/practice_approximate_Q_learning.ipynb | GimmeDanger/made_nlp_course |
Approximate Q-learning: building the networkTo train a neural network policy one must have a neural network policy. Let's build it.Since we're working with a pre-extracted features (cart positions, angles and velocities), we don't need a complicated network yet. In fact, let's build something like this for starters: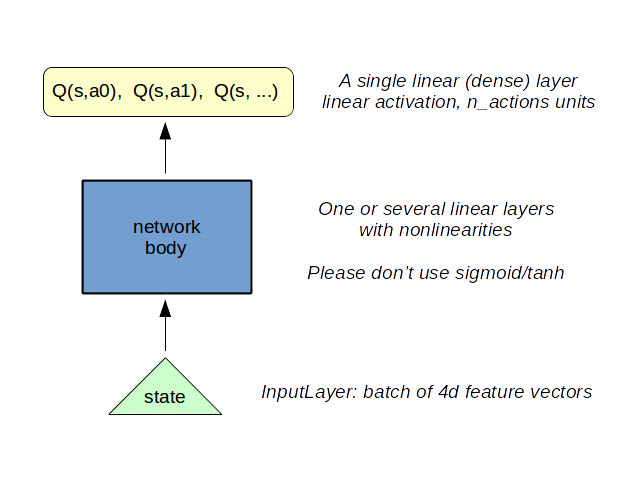For your first run, please only use linear layers (nn.Linear) and activations. Stuff like batch normalization or dropout may ruin everything if used haphazardly. Also please avoid using nonlinearities like sigmoid & tanh: agent's observations are not normalized so sigmoids may become saturated from init.Ideally you should start small with maybe 1-2 hidden layers with < 200 neurons and then increase network size if agent doesn't beat the target score. | import torch
import torch.nn as nn
import torch.nn.functional as F
network = nn.Sequential()
network.add_module('layer1', < ... >)
<YOUR CODE: stack layers!!!1 >
# hint: use state_dim[0] as input size
def get_action(state, epsilon=0):
"""
sample actions with epsilon-greedy policy
recap: with p = epsilon pick random action, else pick action with highest Q(s,a)
"""
state = torch.tensor(state[None], dtype=torch.float32)
q_values = network(state).detach().numpy()
# YOUR CODE
return int( < epsilon-greedily selected action > )
s = env.reset()
assert tuple(network(torch.tensor([s]*3, dtype=torch.float32)).size()) == (
3, n_actions), "please make sure your model maps state s -> [Q(s,a0), ..., Q(s, a_last)]"
assert isinstance(list(network.modules(
))[-1], nn.Linear), "please make sure you predict q-values without nonlinearity (ignore if you know what you're doing)"
assert isinstance(get_action(
s), int), "get_action(s) must return int, not %s. try int(action)" % (type(get_action(s)))
# test epsilon-greedy exploration
for eps in [0., 0.1, 0.5, 1.0]:
state_frequencies = np.bincount(
[get_action(s, epsilon=eps) for i in range(10000)], minlength=n_actions)
best_action = state_frequencies.argmax()
assert abs(state_frequencies[best_action] -
10000 * (1 - eps + eps / n_actions)) < 200
for other_action in range(n_actions):
if other_action != best_action:
assert abs(state_frequencies[other_action] -
10000 * (eps / n_actions)) < 200
print('e=%.1f tests passed' % eps) | _____no_output_____ | MIT | week10_Model_free_learning_peek/practice_approximate_Q_learning.ipynb | GimmeDanger/made_nlp_course |
Q-learning via gradient descentWe shall now train our agent's Q-function by minimizing the TD loss:$$ L = { 1 \over N} \sum_i (Q_{\theta}(s,a) - [r(s,a) + \gamma \cdot max_{a'} Q_{-}(s', a')]) ^2 $$Where* $s, a, r, s'$ are current state, action, reward and next state respectively* $\gamma$ is a discount factor defined two cells above.The tricky part is with $Q_{-}(s',a')$. From an engineering standpoint, it's the same as $Q_{\theta}$ - the output of your neural network policy. However, when doing gradient descent, __we won't propagate gradients through it__ to make training more stable (see lectures).To do so, we shall use `x.detach()` function which basically says "consider this thing constant when doingbackprop". | def to_one_hot(y_tensor, n_dims=None):
""" helper: take an integer vector and convert it to 1-hot matrix. """
y_tensor = y_tensor.type(torch.LongTensor).view(-1, 1)
n_dims = n_dims if n_dims is not None else int(torch.max(y_tensor)) + 1
y_one_hot = torch.zeros(
y_tensor.size()[0], n_dims).scatter_(1, y_tensor, 1)
return y_one_hot
def where(cond, x_1, x_2):
""" helper: like np.where but in pytorch. """
return (cond * x_1) + ((1-cond) * x_2)
def compute_td_loss(states, actions, rewards, next_states, is_done, gamma=0.99, check_shapes=False):
""" Compute td loss using torch operations only. Use the formula above. """
states = torch.tensor(
states, dtype=torch.float32) # shape: [batch_size, state_size]
actions = torch.tensor(actions, dtype=torch.int32) # shape: [batch_size]
rewards = torch.tensor(rewards, dtype=torch.float32) # shape: [batch_size]
# shape: [batch_size, state_size]
next_states = torch.tensor(next_states, dtype=torch.float32)
is_done = torch.tensor(is_done, dtype=torch.float32) # shape: [batch_size]
# get q-values for all actions in current states
predicted_qvalues = network(states)
# select q-values for chosen actions
predicted_qvalues_for_actions = torch.sum(
predicted_qvalues * to_one_hot(actions, n_actions), dim=1)
# compute q-values for all actions in next states
predicted_next_qvalues = # YOUR CODE
# compute V*(next_states) using predicted next q-values
next_state_values = # YOUR CODE
assert next_state_values.dtype == torch.float32
# compute "target q-values" for loss - it's what's inside square parentheses in the above formula.
target_qvalues_for_actions = # YOUR CODE
# at the last state we shall use simplified formula: Q(s,a) = r(s,a) since s' doesn't exist
target_qvalues_for_actions = where(
is_done, rewards, target_qvalues_for_actions)
# mean squared error loss to minimize
loss = torch.mean((predicted_qvalues_for_actions -
target_qvalues_for_actions.detach()) ** 2)
if check_shapes:
assert predicted_next_qvalues.data.dim(
) == 2, "make sure you predicted q-values for all actions in next state"
assert next_state_values.data.dim(
) == 1, "make sure you computed V(s') as maximum over just the actions axis and not all axes"
assert target_qvalues_for_actions.data.dim(
) == 1, "there's something wrong with target q-values, they must be a vector"
return loss
# sanity checks
s = env.reset()
a = env.action_space.sample()
next_s, r, done, _ = env.step(a)
loss = compute_td_loss([s], [a], [r], [next_s], [done], check_shapes=True)
loss.backward()
assert len(loss.size()) == 0, "you must return scalar loss - mean over batch"
assert np.any(next(network.parameters()).grad.detach().numpy() !=
0), "loss must be differentiable w.r.t. network weights" | _____no_output_____ | MIT | week10_Model_free_learning_peek/practice_approximate_Q_learning.ipynb | GimmeDanger/made_nlp_course |
Playing the game | opt = torch.optim.Adam(network.parameters(), lr=1e-4)
epsilon = 0.5
def generate_session(t_max=1000, epsilon=0, train=False):
"""play env with approximate q-learning agent and train it at the same time"""
total_reward = 0
s = env.reset()
for t in range(t_max):
a = get_action(s, epsilon=epsilon)
next_s, r, done, _ = env.step(a)
if train:
opt.zero_grad()
compute_td_loss([s], [a], [r], [next_s], [done]).backward()
opt.step()
total_reward += r
s = next_s
if done:
break
return total_reward
for i in range(1000):
session_rewards = [generate_session(
epsilon=epsilon, train=True) for _ in range(100)]
print("epoch #{}\tmean reward = {:.3f}\tepsilon = {:.3f}".format(
i, np.mean(session_rewards), epsilon))
epsilon *= 0.99
assert epsilon >= 1e-4, "Make sure epsilon is always nonzero during training"
if np.mean(session_rewards) > 300:
print("You Win!")
break | _____no_output_____ | MIT | week10_Model_free_learning_peek/practice_approximate_Q_learning.ipynb | GimmeDanger/made_nlp_course |
How to interpret resultsWelcome to the f.. world of deep f...n reinforcement learning. Don't expect agent's reward to smoothly go up. Hope for it to go increase eventually. If it deems you worthy.Seriously though,* __ mean reward__ is the average reward per game. For a correct implementation it may stay low for some 10 epochs, then start growing while oscilating insanely and converges by ~50-100 steps depending on the network architecture. * If it never reaches target score by the end of for loop, try increasing the number of hidden neurons or look at the epsilon.* __ epsilon__ - agent's willingness to explore. If you see that agent's already at < 0.01 epsilon before it's is at least 200, just reset it back to 0.1 - 0.5. Record videosAs usual, we now use `gym.wrappers.Monitor` to record a video of our agent playing the game. Unlike our previous attempts with state binarization, this time we expect our agent to act ~~(or fail)~~ more smoothly since there's no more binarization error at play.As you already did with tabular q-learning, we set epsilon=0 for final evaluation to prevent agent from exploring himself to death. | # record sessions
import gym.wrappers
env = gym.wrappers.Monitor(gym.make("CartPole-v0"),
directory="videos", force=True)
sessions = [generate_session(epsilon=0, train=False) for _ in range(100)]
env.close()
# Show video. This may not work in some setups. If it doesn't
# work for you, you can download the videos and view them locally.
import sys
from pathlib import Path
from base64 import b64encode
from IPython.display import HTML
video_paths = sorted([s for s in Path('videos').iterdir() if s.suffix == '.mp4'])
video_path = video_paths[-3] # You can also try other indices
if 'google.colab' in sys.modules:
# https://stackoverflow.com/a/57378660/1214547
with video_path.open('rb') as fp:
mp4 = fp.read()
data_url = 'data:video/mp4;base64,' + b64encode(mp4).decode()
else:
data_url = str(video_path)
HTML("""
<video width="640" height="480" controls>
<source src="{}" type="video/mp4">
</video>
""".format(data_url)) | _____no_output_____ | MIT | week10_Model_free_learning_peek/practice_approximate_Q_learning.ipynb | GimmeDanger/made_nlp_course |
Frame of reference> Marcos Duarte, Renato Naville Watanabe > [Laboratory of Biomechanics and Motor Control](http://pesquisa.ufabc.edu.br/bmclab) > Federal University of ABC, Brazil Contents1 Frame of reference for human motion analysis2 Cartesian coordinate system2.1 Standardizations in movement analysis 3 Determination of a coordinate system3.1 Definition of a basis3.2 Using the cross product3.3 Gram–Schmidt process4 Polar and spherical coordinate systems4.1 Polar coordinate system4.2 Spherical coordinate system 5 Generalized coordinates6 Further reading7 Video lectures on the Internet8 Problems9 References Motion (a change of position in space with respect to time) is not an absolute concept; a reference is needed to describe the motion of the object in relation to this reference. Likewise, the state of such reference cannot be absolute in space and so motion is relative. A [frame of reference](http://en.wikipedia.org/wiki/Frame_of_reference) is the place with respect to we choose to describe the motion of an object. In this reference frame, we define a [coordinate system](http://en.wikipedia.org/wiki/Coordinate_system) (a set of axes) within which we measure the motion of an object (but frame of reference and coordinate system are often used interchangeably). Often, the choice of reference frame and coordinate system is made by convenience. However, there is an important distinction between reference frames when we deal with the dynamics of motion, where we are interested to understand the forces related to the motion of the object. In dynamics, we refer to [inertial frame of reference](http://en.wikipedia.org/wiki/Inertial_frame_of_reference) (a.k.a., Galilean reference frame) when the Newton's laws of motion in their simple form are valid in this frame and to non-inertial frame of reference when the Newton's laws in their simple form are not valid (in such reference frame, fictitious accelerations/forces appear). An inertial reference frame is at rest or moves at constant speed (because there is no absolute rest!), whereas a non-inertial reference frame is under acceleration (with respect to an inertial reference frame).The concept of frame of reference has changed drastically since Aristotle, Galileo, Newton, and Einstein. To read more about that and its philosophical implications, see [Space and Time: Inertial Frames](http://plato.stanford.edu/entries/spacetime-iframes/). Frame of reference for human motion analysisIn anatomy, we use a simplified reference frame composed by perpendicular planes to provide a standard reference for qualitatively describing the structures and movements of the human body, as shown in the next figure.Figure. Anatomical body position and body planes (image from Wikipedia). Cartesian coordinate systemAs we perceive the surrounding space as three-dimensional, a convenient coordinate system is the [Cartesian coordinate system](http://en.wikipedia.org/wiki/Cartesian_coordinate_system) in the [Euclidean space](http://en.wikipedia.org/wiki/Euclidean_space) with three orthogonal axes as shown below. The axes directions are commonly defined by the [right-hand rule](http://en.wikipedia.org/wiki/Right-hand_rule) and attributed the letters X, Y, Z. The orthogonality of the Cartesian coordinate system is convenient for its use in classical mechanics, most of the times the structure of space is assumed having the [Euclidean geometry](http://en.wikipedia.org/wiki/Euclidean_geometry) and as consequence, the motion in different directions are independent of each other. Figure. A point in three-dimensional Euclidean space described in a Cartesian coordinate system. Standardizations in movement analysisThe concept of reference frame in Biomechanics and motor control is very important and central to the understanding of human motion. For example, do we see, plan and control the movement of our hand with respect to reference frames within our body or in the environment we move? Or a combination of both? The figure below, although derived for a robotic system, illustrates well the concept that we might have to deal with multiple coordinate systems. Figure. Multiple coordinate systems for use in robots (figure from Corke (2017)).For three-dimensional motion analysis in Biomechanics, we may use several different references frames for convenience and refer to them as global, laboratory, local, anatomical, or technical reference frames or coordinate systems (we will study this later). There has been proposed different standardizations on how to define frame of references for the main segments and joints of the human body. For instance, the International Society of Biomechanics has a [page listing standardization proposals](https://isbweb.org/activities/standards) by its standardization committee and subcommittees: | from IPython.display import IFrame
IFrame('https://isbweb.org/activities/standards', width='100%', height=400) | _____no_output_____ | CC-BY-4.0 | notebooks/ReferenceFrame.ipynb | rnwatanabe/BMC |
Another initiative for the standardization of references frames is from the [Virtual Animation of the Kinematics of the Human for Industrial, Educational and Research Purposes (VAKHUM)](https://raw.githubusercontent.com/demotu/BMC/master/refs/VAKHUM.pdf) project. Determination of a coordinate systemIn Biomechanics, we may use different coordinate systems for convenience and refer to them as global, laboratory, local, anatomical, or technical reference frames or coordinate systems. For example, in a standard gait analysis, we define a global or laboratory coordinate system and a different coordinate system for each segment of the body to be able to describe the motion of a segment in relation to anatomical axes of another segment. To define this anatomical coordinate system, we need to place markers on anatomical landmarks on each segment. We also may use other markers (technical markers) on the segment to improve the motion analysis and then we will also have to define a technical coordinate system for each segment.As we perceive the surrounding space as three-dimensional, a convenient coordinate system to use is the [Cartesian coordinate system](http://en.wikipedia.org/wiki/Cartesian_coordinate_system) with three orthogonal axes in the [Euclidean space](http://en.wikipedia.org/wiki/Euclidean_space). From [linear algebra](http://en.wikipedia.org/wiki/Linear_algebra), a set of unit linearly independent vectors (orthogonal in the Euclidean space and each with norm (length) equals to one) that can represent any vector via [linear combination](http://en.wikipedia.org/wiki/Linear_combination) is called a basis (or orthonormal basis). The figure below shows a point and its position vector in the Cartesian coordinate system and the corresponding versors (unit vectors) of the basis for this coordinate system. See the notebook [Scalar and vector](http://nbviewer.ipython.org/github/demotu/BMC/blob/master/notebooks/ScalarVector.ipynb) for a description on vectors. Figure. Representation of a point **P** and its position vector $\overrightarrow{\mathbf{r}}$ in a Cartesian coordinate system. The versors $\hat{\mathbf{i}}, \hat{\mathbf{j}}, \hat{\mathbf{k}}$ form a basis for this coordinate system and are usually represented in the color sequence RGB (red, green, blue) for easier visualization.One can see that the versors of the basis shown in the figure above have the following coordinates in the Cartesian coordinate system:\begin{equation}\hat{\mathbf{i}} = \begin{bmatrix}1\\0\\0 \end{bmatrix}, \quad \hat{\mathbf{j}} = \begin{bmatrix}0\\1\\0 \end{bmatrix}, \quad \hat{\mathbf{k}} = \begin{bmatrix} 0 \\ 0 \\ 1 \end{bmatrix}\end{equation}Using the notation described in the figure above, the position vector $\overrightarrow{\mathbf{r}}$ (or the point $\overrightarrow{\mathbf{P}}$) can be expressed as:\begin{equation}\overrightarrow{\mathbf{r}} = x\hat{\mathbf{i}} + y\hat{\mathbf{j}} + z\hat{\mathbf{k}}\end{equation} Definition of a basisThe mathematical problem of determination of a coordinate system is to find a basis and an origin for it (a basis is only a set of vectors, with no origin). There are different methods to calculate a basis given a set of points (coordinates), for example, one can use the scalar product or the cross product for this problem. Using the cross productLet's now define a basis using a common method in motion analysis (employing the cross product): Given the coordinates of three noncollinear points in 3D space (points that do not all lie on the same line), $\overrightarrow{\mathbf{m}}_1, \overrightarrow{\mathbf{m}}_2, \overrightarrow{\mathbf{m}}_3$, which would represent the positions of markers captured from a motion analysis session, a basis can be found following these steps: 1. First axis, $\overrightarrow{\mathbf{v}}_1$, the vector $\overrightarrow{\mathbf{m}}_2-\overrightarrow{\mathbf{m}}_1$ (or any other vector difference); 2. Second axis, $\overrightarrow{\mathbf{v}}_2$, the cross or vector product between the vectors $\overrightarrow{\mathbf{v}}_1$ and $\overrightarrow{\mathbf{m}}_3-\overrightarrow{\mathbf{m}}_1$ (or $\overrightarrow{\mathbf{m}}_3-\overrightarrow{\mathbf{m}}_2$); 3. Third axis, $\overrightarrow{\mathbf{v}}_3$, the cross product between the vectors $\overrightarrow{\mathbf{v}}_1$ and $\overrightarrow{\mathbf{v}}_2$. 4. Make all vectors to have norm 1 dividing each vector by its norm. The positions of the points used to construct a coordinate system have, by definition, to be specified in relation to an already existing coordinate system. In motion analysis, this coordinate system is the coordinate system from the motion capture system and it is established in the calibration phase. In this phase, the positions of markers placed on an object with perpendicular axes and known distances between the markers are captured and used as the reference (laboratory) coordinate system. For example, given the positions $\overrightarrow{\mathbf{m}}_1 = [1,2,5], \overrightarrow{\mathbf{m}}_2 = [2,3,3], \overrightarrow{\mathbf{m}}_3 = [4,0,2]$, a basis can be found with: | import numpy as np
m1 = np.array([1, 2, 5])
m2 = np.array([2, 3, 3])
m3 = np.array([4, 0, 2])
v1 = m2 - m1 # first axis
v2 = np.cross(v1, m3 - m1) # second axis
v3 = np.cross(v1, v2) # third axis
# Vector normalization
e1 = v1/np.linalg.norm(v1)
e2 = v2/np.linalg.norm(v2)
e3 = v3/np.linalg.norm(v3)
print('Versors:', '\ne1 =', e1, '\ne2 =', e2, '\ne3 =', e3)
print('\nTest of orthogonality (cross product between versors):',
'\ne1 x e2:', np.linalg.norm(np.cross(e1, e2)),
'\ne1 x e3:', np.linalg.norm(np.cross(e1, e3)),
'\ne2 x e3:', np.linalg.norm(np.cross(e2, e3)))
print('\nNorm of each versor:',
'\n||e1|| =', np.linalg.norm(e1),
'\n||e2|| =', np.linalg.norm(e2),
'\n||e3|| =', np.linalg.norm(e3)) | Versors:
e1 = [ 0.40824829 0.40824829 -0.81649658]
e2 = [-0.76834982 -0.32929278 -0.5488213 ]
e3 = [-0.49292179 0.85141036 0.17924429]
Test of orthogonality (cross product between versors):
e1 x e2: 1.0
e1 x e3: 1.0000000000000002
e2 x e3: 0.9999999999999999
Norm of each versor:
||e1|| = 1.0
||e2|| = 1.0
||e3|| = 1.0
| CC-BY-4.0 | notebooks/ReferenceFrame.ipynb | rnwatanabe/BMC |
To define a coordinate system using the calculated basis, we also need to define an origin. In principle, we could use any point as origin, but if the calculated coordinate system should follow anatomical conventions, e.g., the coordinate system origin should be at a joint center, we will have to calculate the basis and origin according to standards used in motion analysis as discussed before. If the coordinate system is a technical basis and not anatomic-based, a common procedure in motion analysis is to define the origin for the coordinate system as the centroid (average) position among the markers at the reference frame. Using the average position across markers potentially reduces the effect of noise (for example, from soft tissue artifact) on the calculation. For the markers in the example above, the origin of the coordinate system will be: | origin = np.mean((m1, m2, m3), axis=0)
print('Origin: ', origin) | Origin: [2.33333333 1.66666667 3.33333333]
| CC-BY-4.0 | notebooks/ReferenceFrame.ipynb | rnwatanabe/BMC |
Let's plot the coordinate system and the basis using the custom Python function `CCS.py`: | import sys
sys.path.insert(1, r'./../functions') # add to pythonpath
from CCS import CCS
markers = np.vstack((m1, m2, m3))
basis = np.vstack((e1, e2, e3)) | _____no_output_____ | CC-BY-4.0 | notebooks/ReferenceFrame.ipynb | rnwatanabe/BMC |
Create figure in this page (inline): | %matplotlib notebook
markers = np.vstack((m1, m2, m3))
basis = np.vstack((e1, e2, e3))
CCS(xyz=[], Oijk=origin, ijk=basis, point=markers, vector=True); | _____no_output_____ | CC-BY-4.0 | notebooks/ReferenceFrame.ipynb | rnwatanabe/BMC |
Gram–Schmidt processAnother classical procedure in mathematics, employing the scalar product, is known as the [Gram–Schmidt process](http://en.wikipedia.org/wiki/Gram%E2%80%93Schmidt_process). See the notebook [Scalar and Vector](http://nbviewer.jupyter.org/github/bmclab/BMC/blob/master/notebooks/ScalarVector.ipynb) for a demonstration of the Gram–Schmidt process and how to implement it in Python.The [Gram–Schmidt process](http://en.wikipedia.org/wiki/Gram%E2%80%93Schmidt_process) is a method for orthonormalizing (orthogonal unit versors) a set of vectors using the scalar product. The Gram–Schmidt process works for any number of vectors. For example, given three vectors, $\overrightarrow{\mathbf{a}}, \overrightarrow{\mathbf{b}}, \overrightarrow{\mathbf{c}}$, in the 3D space, a basis $\{\hat{e}_a, \hat{e}_b, \hat{e}_c\}$ can be found using the Gram–Schmidt process by: The first versor is in the $\overrightarrow{\mathbf{a}}$ direction (or in the direction of any of the other vectors): \begin{equation}\hat{e}_a = \frac{\overrightarrow{\mathbf{a}}}{||\overrightarrow{\mathbf{a}}||}\end{equation} The second versor, orthogonal to $\hat{e}_a$, can be found considering we can express vector $\overrightarrow{\mathbf{b}}$ in terms of the $\hat{e}_a$ direction as: $$ \overrightarrow{\mathbf{b}} = \overrightarrow{\mathbf{b}}^\| + \overrightarrow{\mathbf{b}}^\bot $$Then:$$ \overrightarrow{\mathbf{b}}^\bot = \overrightarrow{\mathbf{b}} - \overrightarrow{\mathbf{b}}^\| = \overrightarrow{\mathbf{b}} - (\overrightarrow{\mathbf{b}} \cdot \hat{e}_a ) \hat{e}_a $$Finally:$$ \hat{e}_b = \frac{\overrightarrow{\mathbf{b}}^\bot}{||\overrightarrow{\mathbf{b}}^\bot||} $$ The third versor, orthogonal to $\{\hat{e}_a, \hat{e}_b\}$, can be found expressing the vector $\overrightarrow{\mathbf{C}}$ in terms of $\hat{e}_a$ and $\hat{e}_b$ directions as:$$ \overrightarrow{\mathbf{c}} = \overrightarrow{\mathbf{c}}^\| + \overrightarrow{\mathbf{c}}^\bot $$Then:$$ \overrightarrow{\mathbf{c}}^\bot = \overrightarrow{\mathbf{c}} - \overrightarrow{\mathbf{c}}^\| $$Where:$$ \overrightarrow{\mathbf{c}}^\| = (\overrightarrow{\mathbf{c}} \cdot \hat{e}_a ) \hat{e}_a + (\overrightarrow{\mathbf{c}} \cdot \hat{e}_b ) \hat{e}_b $$Finally:$$ \hat{e}_c = \frac{\overrightarrow{\mathbf{c}}^\bot}{||\overrightarrow{\mathbf{c}}^\bot||} $$ Let's implement the Gram–Schmidt process in Python.For example, consider the positions (vectors) $\overrightarrow{\mathbf{a}} = [1,2,0], \overrightarrow{\mathbf{b}} = [0,1,3], \overrightarrow{\mathbf{c}} = [1,0,1]$: | import numpy as np
a = np.array([1, 2, 0])
b = np.array([0, 1, 3])
c = np.array([1, 0, 1]) | _____no_output_____ | CC-BY-4.0 | notebooks/ReferenceFrame.ipynb | rnwatanabe/BMC |
The first versor is: | ea = a/np.linalg.norm(a)
print(ea) | [ 0.4472136 0.89442719 0. ]
| CC-BY-4.0 | notebooks/ReferenceFrame.ipynb | rnwatanabe/BMC |
The second versor is: | eb = b - np.dot(b, ea)*ea
eb = eb/np.linalg.norm(eb)
print(eb) | [-0.13187609 0.06593805 0.98907071]
| CC-BY-4.0 | notebooks/ReferenceFrame.ipynb | rnwatanabe/BMC |
And the third version is: | ec = c - np.dot(c, ea)*ea - np.dot(c, eb)*eb
ec = ec/np.linalg.norm(ec)
print(ec) | [ 0.88465174 -0.44232587 0.14744196]
| CC-BY-4.0 | notebooks/ReferenceFrame.ipynb | rnwatanabe/BMC |
Let's check the orthonormality between these versors: | print(' Versors:', '\nea =', ea, '\neb =', eb, '\nec =', ec)
print('\n Test of orthogonality (scalar product between versors):',
'\n ea x eb:', np.dot(ea, eb),
'\n eb x ec:', np.dot(eb, ec),
'\n ec x ea:', np.dot(ec, ea))
print('\n Norm of each versor:',
'\n ||ea|| =', np.linalg.norm(ea),
'\n ||eb|| =', np.linalg.norm(eb),
'\n ||ec|| =', np.linalg.norm(ec)) | Versors:
ea = [ 0.4472136 0.89442719 0. ]
eb = [-0.13187609 0.06593805 0.98907071]
ec = [ 0.88465174 -0.44232587 0.14744196]
Test of orthogonality (scalar product between versors):
ea x eb: 2.08166817117e-17
eb x ec: -2.77555756156e-17
ec x ea: 5.55111512313e-17
Norm of each versor:
||ea|| = 1.0
||eb|| = 1.0
||ec|| = 1.0
| CC-BY-4.0 | notebooks/ReferenceFrame.ipynb | rnwatanabe/BMC |
Volume Sampling vs projection DPP for low rank approximation Introduction In this notebook we compare the volume sampling and projection DPP for low rank approximation.We recall the result proved in the article [DRVW]:\\Let S be a random subset of k columns of X chosen with probability: $$P(S) = \frac{1}{Z_{k}} det(X_{.,S}^{T}X_{.,S})$$ with $$Z_{k} = \sum\limits_{S \subset [N], |S| = k} det(X_{.,S}^{T}X_{.,S})$$Then$$\begin{equation}E(\| X - \pi_{X_{.,S}}(X) \|_{Fr}^{2}) \leq (k+1)\| X - \pi_{k}(X) \|_{Fr}^{2}\end{equation}$$We can prove that the volume sampling distribution is a mixture of projection DPPs distributions..., in particular one projection DPP distribution stands out for the problem of low rank approximation: ....\\For the moment, there is no analytical expression for $$\begin{equation}E(\| X - \pi_{X_{.,S}}(X) \|_{Fr}^{2}) \end{equation}$$ under the distribution of projection DPP.\\However, we can calculate this quantity using simulation on some matrices representing cloud points with some specific geometric constraints.Let $$X \in R^{n \times m}$$ a matrix representing a cloud of points.We can write the SVD of $$X = UDV^{T}$$ In this notebook we investigate the influence of some structures enforced to V and D on the expected error expressed above for different algorithms: Volume Sampling, Projection DPP and the deterministic algorithm. As for the Volume Sampling distribution, we can express the expected approximation error using only the elements of D. We can test this theoretical property in the next Numerical Study below. However, there is no closed formula (for the moment) for the expected approximation error under Projection DPP distribution. We will see in the Numerical Study section, that this value cannot depends only on the elements of D. References[DRVW] Deshpande, Amit and Rademacher, Luis and Vempala, Santosh and Wang, Grant - Matrix Approximation and Projective Clustering via Volume Sampling 2006[BoDr] Boutsidis, Christos and Drineas, Petros - Deterministic and randomized column selection algorithms for matrices 2014[] INDERJIT S. DHILLON , ROBERT W. HEATH JR., MA ́TYA ́S A. SUSTIK, ANDJOEL A. TROPP - GENERALIZED FINITE ALGORITHMS FOR CONSTRUCTING HERMITIAN MATRICESWITH PRESCRIBED DIAGONAL AND SPECTRUM 2005 I- Generating a cloud of points with geometric constraintsIn this simulation we will enforce some structure on the matrix V for two values of the matrix D. While the matrix U will be choosen randomly. We want to investigate the influence of the profile of the norms of the V_k rows: the k-leverage scores. For this purpose we use an algorithm proposed in the article []: this algorithm outputs a ( dxk) matrix Q with orthonormal columns and a prescribed profile of the norms of the rows. If we consider the Gram matrix H= QQ^{T}, this boils down to enforce the diagonal of H while keeping its spectrum containing k ones and d-k zeros. The algorithm proceed as following:* Initialization of the matrix Q by the rectangular identity* Apply a Givens Rotation (of dimension d) to the matrix Q: this step will enforce the norm of a row every iteration* Outputs the resulting matrix when all the rows norms are enforced. | import numpy as np
import pandas as pd
from itertools import combinations
from scipy.stats import binom
import scipy.special
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
from IPython.display import display, HTML
from FrameBuilder.eigenstepsbuilder import *
from decimal import *
u = np.random.uniform(0,1)
u | _____no_output_____ | MIT | .ipynb_checkpoints/Numerical_Analysis_Volume_Sampling_vs_Projection_DPP-ollld-checkpoint.ipynb | AyoubBelhadji/FrameBuilder |
I-1- Givens Rotations generatorsThese functions generate a Givens rotation | def t_func(q_i,q_j,q_ij,l_i,l_j):
# t in section 3.1 Dhillon (2005)
delta = np.power(q_ij,2)-(q_i-l_i)*(q_j-l_i)
if delta<0:
print(delta)
print("error sqrt")
t = q_ij - np.sqrt(delta)
t = t/(q_j-l_i)
return t
def G_func(i,j,q_i,q_j,q_ij,l_i,l_j,N):
# Gitens Rotation
G=np.eye(N)
t = t_func(q_i,q_j,q_ij,l_i,l_j)
c = 1/(np.sqrt(np.power(t,2)+1))
s = t*c
G[i,i]=c
G[i,j]=s
G[j,i]= -s
G[j,j]= c
return G | _____no_output_____ | MIT | .ipynb_checkpoints/Numerical_Analysis_Volume_Sampling_vs_Projection_DPP-ollld-checkpoint.ipynb | AyoubBelhadji/FrameBuilder |
The following function is an implementation of the algorithm [] figuring in the article [] to generate an orthogonal matrix with a prescribed profile of leverage scores.In fact this is a simplification of the algorithm ..... | class Data_Set_Generator:
def __init__(self, N, d, nu, Sigma):
self.N = N
self.d = d
self.nu = nu
self.Sigma = Sigma
self.mean = np.zeros(d)
def multivariate_t_rvs(self):
x = np.random.chisquare(self.nu, self.N)/self.nu
z = np.random.multivariate_normal(self.mean,self.Sigma,(self.N,))
return self.mean + z/np.sqrt(x)[:,None]
def generate_orthonormal_matrix_with_leverage_scores_ES(N,d,lv_scores_vector,versions_number,nn_cardinal_list):
lambda_vector = np.zeros((N))
lambda_vector[0:d] = np.ones((d))
#mu_vector = np.linspace(1, 0.1, num=N)
#sum_mu_vector = np.sum(mu_vector)
#mu_vector = d/sum_mu_vector*mu_vector
Q = np.zeros((N,d))
previous_Q = np.zeros((versions_number+1,N,d))
#mu_vector = d/N*np.ones((N,1))
E = np.zeros((N,N)) #(d,N)
counter = 0
for j in nn_cardinal_list:
print("counter")
print(counter)
mu_vector = generate_leverage_scores_vector_with_dirichlet(N,d,j)
print(np.sum(mu_vector))
print(mu_vector)
E_test = get_eigensteps_random(mu_vector,lambda_vector,N,d)
E_ = np.zeros((d,N+1))
for i in range(d):
E_[i,1:N+1] = E_test[i,:]
print(E_test)
#F_test = get_F(d,N,np.asmatrix(E_),mu_vector)
#previous_Q[counter,:,:] = np.transpose(F_test)
#Q = np.transpose(F_test)
counter = counter +1
return Q,previous_Q
Q,previous_Q = generate_orthonormal_matrix_with_leverage_scores_ES(20,2,[],3,[18,15,10])
def generate_leverage_scores_vector_with_dirichlet(d,k,nn_cardinal):
getcontext().prec = 3
mu_vector = np.float16(np.zeros((d,)))
mu_vector_2 = np.float16(np.zeros((d,)))
not_bounded = 1
while(not_bounded == 1):
mu_vector[0:nn_cardinal] = (k*np.random.dirichlet([1]*nn_cardinal, 1))[0]
mu_vector = np.flip(np.sort(mu_vector),axis = 0)
if max(mu_vector)<=1:
not_bounded = 0
for i in range(nn_cardinal):
mu_vector_2[i] = round(mu_vector[i],4)
mu_vector_2 = k*mu_vector_2/np.sum(mu_vector_2)
return list(mu_vector_2)
l = generate_leverage_scores_vector_with_dirichlet(10,2,6)
print(l)
print(np.sum(l))
def generate_orthonormal_matrix_with_leverage_scores(N,d,lv_scores_vector,versions_number,mode):
#Transforming an idendity matrix to an orthogonal matrix with prescribed lengths
Q = np.zeros((N,d))
previous_Q = np.zeros((versions_number+1,N,d))
versionning_period = (int)(N/versions_number)
if mode == 'identity':
for _ in range(0,d):
Q[_,_] = 1
if mode == 'spread':
nu = 1
Sigma = np.diag(np.ones(d))
mean = np.zeros(d)
x = np.random.chisquare(nu, N)/nu
z = np.random.multivariate_normal(mean,Sigma,(N,))
dataset = mean + z/np.sqrt(x)[:,None]
[Q,_,_] = np.linalg.svd(dataset,full_matrices=False)
print(np.shape(Q))
I_sorting = list(reversed(np.argsort(lv_scores_vector)))
lv_scores_vector = np.asarray(list(reversed(np.sort(lv_scores_vector))))
initial_lv_scores_vector = np.diag(np.dot(Q,Q.T))
I_initial_sorting = list(reversed(np.argsort(initial_lv_scores_vector)))
initial_lv_scores_vector = np.asarray(list(reversed(np.sort(np.diag(np.dot(Q,Q.T))))))
#initial_lv_scores_vector =
Q[I_initial_sorting,:] = Q
print(lv_scores_vector)
print(initial_lv_scores_vector)
delta_lv_scores_vector = lv_scores_vector - initial_lv_scores_vector
print(delta_lv_scores_vector)
min_index = next((i for i, x in enumerate(delta_lv_scores_vector) if x>0), None)
i = min_index-1
j = min_index
print(i)
print(j)
#if mode == 'identity':
# i = d-1
# j = d
#if mode == 'spread':
# i = d-2
# j = d-1
v_counter =0
for t in range(N-1):
#print(i)
#print(j)
delta_i = np.abs(lv_scores_vector[i] - np.power(np.linalg.norm(Q[i,:]),2))
delta_j = np.abs(lv_scores_vector[j] - np.power(np.linalg.norm(Q[j,:]),2))
q_i = np.power(np.linalg.norm(Q[i,:]),2)
q_j = np.power(np.linalg.norm(Q[j,:]),2)
q_ij = np.dot(Q[i,:],Q[j,:].T)
l_i = lv_scores_vector[i]
l_j = lv_scores_vector[j]
G = np.eye(N)
if t%versionning_period ==0:
previous_Q[v_counter,:,:] = Q
v_counter = v_counter +1
if delta_i <= delta_j:
l_k = q_i + q_j -l_i
G = G_func(i,j,q_i,q_j,q_ij,l_i,l_k,N)
Q = np.dot(G,Q)
i = i-1
else:
l_k = q_i + q_j -l_j
G = G_func(i,j,q_j,q_i,q_ij,l_j,l_k,N)
Q = np.dot(G,Q)
j = j+1
previous_Q[versions_number,:,:] = Q
return Q,previous_Q | _____no_output_____ | MIT | .ipynb_checkpoints/Numerical_Analysis_Volume_Sampling_vs_Projection_DPP-ollld-checkpoint.ipynb | AyoubBelhadji/FrameBuilder |
The following function allows to estimate the leverage scores for an orthogonal matrix Q: the function calculates the diagonoal of the matrix $$Q Q^{T}$$ | def estimate_leverage_scores_from_orthogonal_matrix(Q):
[N,_] = np.shape(Q)
lv_scores_vector = np.zeros((N,1))
lv_scores_vector = np.diag(np.dot(Q,np.transpose(Q)))
lv_scores_vector = np.asarray(list(reversed(np.sort(lv_scores_vector))))
return lv_scores_vector
def estimate_sum_first_k_leverage_scores(Q,k):
lv_scores_vector = estimate_leverage_scores_from_orthogonal_matrix(Q)
res = np.sum(lv_scores_vector[0:k])
return res | _____no_output_____ | MIT | .ipynb_checkpoints/Numerical_Analysis_Volume_Sampling_vs_Projection_DPP-ollld-checkpoint.ipynb | AyoubBelhadji/FrameBuilder |
I-2- Extending the orthogonal matrices For the task of low rank approximation, we have seen that only the information contained in the first right k eigenvectors of the matrix X are relevant. In the previous step we build only the first right k eigenvectors but we still need to complete these orthogonal matrices with d-k columns. We proceed as following:Generate a random vector (Nx1) using independent standard gaussian variables,Project this vector in the orthogonal of the span of QNormalize the obtained vector after the projectionExtend the matrix QNote that this procedure is not the unique way to extend the matrix Q to an orthogonal (Nxd) matrix. | def extend_orthogonal_matrix(Q,d_target):
[N,d] = np.shape(Q)
Q_target = np.zeros((N,d))
Q_target = Q
delta = d_target - d
for t in range(delta):
Q_test = np.random.normal(0, 1, N)
for _ in range(d):
Q_test = Q_test - np.dot(Q_test,Q[:,_])*Q[:,_]
Q_test = Q_test/np.linalg.norm(Q_test)
Q_test = Q_test.reshape(N,1)
Q_target = np.append(Q_target,Q_test,1)
return Q_target
#extended_Q = extend_orthogonal_matrix(Q,r) | _____no_output_____ | MIT | .ipynb_checkpoints/Numerical_Analysis_Volume_Sampling_vs_Projection_DPP-ollld-checkpoint.ipynb | AyoubBelhadji/FrameBuilder |
I-3 - Constructing a dataset for every extended orthogonal matrix The previous step allow us to build (N x d) orthogonal matrices such that the extracted (N x k) matrix have a prescribed profile of leverage scores.Now we construct a cloud of point by assigning a covariance matrix D and a matrix V | def contruct_dataset_from_orthogonal_matrix(multi_Q,N,target_d,cov,mean,versions_number):
multi_X = np.zeros((versions_number+1,N,real_dim))
for t in range(versions_number+1):
test_X = np.random.multivariate_normal(mean, cov, N)
[U,_,_] = np.linalg.svd(test_X, full_matrices=False)
Q_test = extend_orthogonal_matrix(multi_Q[t,:,:],target_d)
multi_X[t,:,:] = np.dot(np.dot(Q_test,cov),U.T).T
return multi_X
| _____no_output_____ | MIT | .ipynb_checkpoints/Numerical_Analysis_Volume_Sampling_vs_Projection_DPP-ollld-checkpoint.ipynb | AyoubBelhadji/FrameBuilder |
II- Volume sampling vs Projection DPP for low rank approximationThese functions allow to quantify the approximation error:* approximation_error_function_fro calculate the ratio of the approximation error of a subset of columns to the optimal approximatione error given by the first k left eigenvectors of the matrix X* expected_approximation_error_for_sampling_scheme calculate the expected value of the ratio of the approximatione error under some sampling distribution | def approximation_error_function_fro(Sigma,k,X,X_S):
## Sigma is the spectrum of the matrix X: we need to calculate the optimal approximation error given by the PCA
## k is the rank of the approximation
## X is the initial matrix
## X_S is the subset of columns of the matrix X for witch we calculate the approximation error ratio
d = list(Sigma.shape)[0] # the dimension of the matrix X
Sigma = np.multiply(Sigma,Sigma) # Sigma power 2 -> we are intersted in the approximation error square
sigma_S_temp = np.linalg.inv(np.dot(X_S.T,X_S)) # just a temporary matrix to construct the projection matrix
projection_S = np.dot(np.dot(X_S,sigma_S_temp),X_S.T) # the projection matrix P_S
res_X = X - np.dot(projection_S,X) # The projection of the matrix X in the orthogonal of S
approximation_error_ratio = np.power(np.linalg.norm(res_X,'fro'),2)/np.sum(Sigma[k:d])
# Calculate the apparoximation error ratio
return approximation_error_ratio
def approximation_error_function_spectral(Sigma,k,X,X_S):
## Sigma is the spectrum of the matrix X: we need to calculate the optimal approximation error given by the PCA
## k is the rank of the approximation
## X is the initial matrix
## X_S is the subset of columns of the matrix X for witch we calculate the approximation error ratio
d = list(Sigma.shape)[0] # the dimension of the matrix X
Sigma = np.multiply(Sigma,Sigma) # Sigma power 2 -> we are intersted in the approximation error square
sigma_S_temp = np.linalg.inv(np.dot(X_S.T,X_S)) # just a temporary matrix to construct the projection matrix
projection_S = np.dot(np.dot(X_S,sigma_S_temp),X_S.T) # the projection matrix P_S
res_X = X - np.dot(projection_S,X) # The projection of the matrix X in the orthogonal of S
approximation_error_ratio = np.power(np.linalg.norm(res_X,ord = 2),2)/np.sum(Sigma[k:k+1])
# Calculate the apparoximation error ratio
return approximation_error_ratio
def upper_bound_error_function_for_projection_DPP(k,X,X_S):
## Sigma is the spectrum of the matrix X: we need to calculate the optimal approximation error given by the PCA
## k is the rank of the approximation
## X is the initial matrix
## X_S is the subset of columns of the matrix X for witch we calculate the approximation error ratio
_,sigma_S_temp,_ = np.linalg.svd(X_S, full_matrices=False) # just a temporary matrix to construct the projection matrix
trunc_product = np.power(np.prod(sigma_S_temp[0:k-1]),2)
if np.power(np.prod(sigma_S_temp[0:k]),2) == 0:
trunc_product = 0
# Calculate the apparoximation error ratio
return trunc_product
def tight_upper_bound_error_function_fro(k,X,X_S,V_k,V_k_S):
## Sigma is the spectrum of the matrix X: we need to calculate the optimal approximation error given by the PCA
## k is the rank of the approximation
## X is the initial matrix
## X_S is the subset of columns of the matrix X for witch we calculate the approximation error ratio
_,Sigma,_ = np.linalg.svd(X, full_matrices=False)
d = list(Sigma.shape)[0]
Sigma = np.multiply(Sigma,Sigma)
if np.linalg.matrix_rank(V_k_S,0.000001) == k:
temp_T = np.dot(np.linalg.inv(V_k_S),V_k)
temp_matrix = X - np.dot(X_S,temp_T)
return np.power(np.linalg.norm(temp_matrix,'fro'),2)/np.sum(Sigma[k:d])
else:
return 0
def get_the_matrix_sum_T_S(k,d,V_k,V_d_k):
## Sigma is the spectrum of the matrix X: we need to calculate the optimal approximation error given by the PCA
## k is the rank of the approximation
## X is the initial matrix
## X_S is the subset of columns of the matrix X for witch we calculate the approximation error ratio
#Sigma = np.multiply(Sigma,Sigma)
#matrices_array = [ np.dot(V_d_k[:,list(comb)],np.dot(np.dot(np.linalg.inv(V_k[:,list(comb)]),np.linalg.inv(V_k[:,list(comb)]))),np.transpose(V_d_k[:,list(comb)])) for comb in combinations(range(d),k) if np.linalg.matrix_rank(V_k[:,list(comb)],0.000001) == k]
T = np.zeros((d-k,d-k))
for comb in combinations(range(d),k):
if np.linalg.matrix_rank(V_k[:,list(comb)],0.0000000001) == k:
V_k_S_inv = np.linalg.inv(V_k[:,list(comb)])
V_d_k_S = V_d_k[:,list(comb)]
V_k_S_inv_2 = np.transpose(np.dot(V_k_S_inv,np.transpose(V_k_S_inv)))
#T = np.dot(np.dot(np.dot(V_d_k_S,np.dot(V_k_S_inv,np.transpose(V_k_S_inv)))),np.transpose(V_d_k_S)) + T
T = np.power(np.linalg.det(V_k[:,list(comb)]),2)*np.dot(V_d_k_S,np.dot(V_k_S_inv_2,np.transpose(V_d_k_S))) +T
return T
def tight_approximation_error_fro_for_sampling_scheme(X,U,k,N):
## X is the matrix X :)
## U is the matrix used in the sampling: we sample propotional to the volume of UU^{T}_{S,S}:
## we are not sampling but we need the weigth to estimate the expected error
## k is the rank of the approximation
## N is the number of columns (to be changed to avoid confusion with the number of points)
_,Sigma,V = np.linalg.svd(X, full_matrices=False)
V_k = V[0:k,:]
## Estimating the spectrum of X -> needed in approximation_error_function_fro
volumes_array = [np.abs(np.linalg.det(np.dot(U[:,list(comb)].T,U[:,list(comb)]))) for comb in combinations(range(N),k)]
## Construct the array of weights: the volumes of UU^{T}_{S,S}
volumes_array_sum = np.sum(volumes_array)
## The normalization constant
volumes_array = volumes_array/volumes_array_sum
## The weigths normalized
approximation_error_array = [tight_upper_bound_error_function_fro(k,X,X[:,list(comb)],V_k,V_k[:,list(comb)]) for comb in combinations(range(N),k)]
## Calculating the approximation error for every k-tuple
expected_value = np.dot(approximation_error_array,volumes_array)
## The expected value of the approximatione error is just the dot product of the two arrays above
return expected_value
def expected_approximation_error_fro_for_sampling_scheme(X,U,k,N):
## X is the matrix X :)
## U is the matrix used in the sampling: we sample propotional to the volume of UU^{T}_{S,S}:
## we are not sampling but we need the weigth to estimate the expected error
## k is the rank of the approximation
## N is the number of columns (to be changed to avoid confusion with the number of points)
_,Sigma,_ = np.linalg.svd(X, full_matrices=False)
## Estimating the spectrum of X -> needed in approximation_error_function_fro
volumes_array = [np.abs(np.linalg.det(np.dot(U[:,list(comb)].T,U[:,list(comb)]))) for comb in combinations(range(N),k)]
## Construct the array of weights: the volumes of UU^{T}_{S,S}
volumes_array_sum = np.sum(volumes_array)
## The normalization constant
volumes_array = volumes_array/volumes_array_sum
## The weigths normalized
approximation_error_array = [approximation_error_function_fro(Sigma,k,X,X[:,list(comb)]) for comb in combinations(range(N),k)]
## Calculating the approximation error for every k-tuple
expected_value = np.dot(approximation_error_array,volumes_array)
## The expected value of the approximatione error is just the dot product of the two arrays above
return expected_value
def expected_approximation_error_spectral_for_sampling_scheme(X,U,k,N):
## X is the matrix X :)
## U is the matrix used in the sampling: we sample propotional to the volume of UU^{T}_{S,S}:
## we are not sampling but we need the weigth to estimate the expected error
## k is the rank of the approximation
## N is the number of columns (to be changed to avoid confusion with the number of points)
_,Sigma,_ = np.linalg.svd(X, full_matrices=False)
## Estimating the spectrum of X -> needed in approximation_error_function_fro
volumes_array = [np.abs(np.linalg.det(np.dot(U[:,list(comb)].T,U[:,list(comb)]))) for comb in combinations(range(N),k)]
## Construct the array of weights: the volumes of UU^{T}_{S,S}
volumes_array_sum = np.sum(volumes_array)
## The normalization constant
volumes_array = volumes_array/volumes_array_sum
## The weigths normalized
approximation_error_array = [approximation_error_function_spectral(Sigma,k,X,X[:,list(comb)]) for comb in combinations(range(N),k)]
## Calculating the approximation error for every k-tuple
expected_value = np.dot(approximation_error_array,volumes_array)
## The expected value of the approximatione error is just the dot product of the two arrays above
return expected_value
def expected_upper_bound_for_projection_DPP(X,U,k,N):
## X is the matrix X :)
## U is the matrix used in the sampling: we sample propotional to the volume of UU^{T}_{S,S}:
## we are not sampling but we need the weigth to estimate the expected error
## k is the rank of the approximation
## N is the number of columns (to be changed to avoid confusion with the number of points)
approximation_error_array = [upper_bound_error_function_for_projection_DPP(k,X,U[:,list(comb)]) for comb in combinations(range(N),k)]
## Calculating the approximation error for every k-tuple
## The expected value of the approximatione error is just the dot product of the two arrays above
#return expected_value
return np.sum(approximation_error_array) | _____no_output_____ | MIT | .ipynb_checkpoints/Numerical_Analysis_Volume_Sampling_vs_Projection_DPP-ollld-checkpoint.ipynb | AyoubBelhadji/FrameBuilder |
III - Numerical analysis In this section we use the functions developed previously to investigate the influence of two parameters: the spectrum of X and the k-leverage scores. For this purpose, we assemble these functionalities in a class allowing fast numerical experiments. | class Numrerical_Analysis_DPP:
def __init__(self,N,real_dim,r,k,versions_number,mean,cov,lv_scores,versions_list):
self.N = N
self.real_dim = real_dim
self.r = r
self.k = k
self.versions_number = versions_number
self.mean = mean
self.cov = cov
self.lv_scores = lv_scores
self.Q = np.zeros((real_dim,k))
self.multi_Q = np.zeros((self.versions_number+1,real_dim,k))
self.X = np.zeros((N,real_dim))
self.multi_X = np.zeros((self.versions_number+1,N,real_dim))
#[self.Q,self.multi_Q] = generate_orthonormal_matrix_with_leverage_scores(real_dim,k,lv_scores,versions_number,'identity')
[self.Q,self.multi_Q] = generate_orthonormal_matrix_with_leverage_scores_ES(self.real_dim,self.k,[],self.versions_number+1,versions_list)
self.multi_X = contruct_dataset_from_orthogonal_matrix(self.multi_Q,self.N,self.real_dim,self.cov,self.mean,self.versions_number)
def contruct_dataset_from_orthogonal_matrix_4(self,multi_Q,N,target_d,cov,mean,versions_number):
test_multi_X = np.zeros((self.versions_number+1,N,real_dim))
for t in range(self.versions_number+1):
test_X = np.random.multivariate_normal(mean, cov, N)
[U,_,_] = np.linalg.svd(test_X, full_matrices=False)
Q_test = extend_orthogonal_matrix(self.multi_Q[t,:,:],target_d)
test_multi_X[t,:,:] = np.dot(np.dot(Q_test,cov),U.T).T
return test_multi_X
def get_effective_kernel_from_orthogonal_matrix(self):
test_eff_V = np.zeros((self.versions_number+1,self.real_dim,self.k))
p_eff_list = self.get_p_eff()
for t in range(self.versions_number+1):
test_V = self.multi_Q[t,:,:]
p_eff = p_eff_list[t]
diag_Q_t = np.diag(np.dot(test_V,test_V.T))
#diag_Q_t = list(diag_Q_t[::-1].sort())
print(diag_Q_t)
permutation_t = list(reversed(np.argsort(diag_Q_t)))
print(permutation_t)
for i in range(self.real_dim):
if i >p_eff-1:
test_V[permutation_t[i],:] = 0
#Q_test = extend_orthogonal_matrix(self.multi_Q[t,:,:],target_d)
test_eff_V[t,:,:] = test_V
return test_eff_V
def get_expected_error_fro_for_volume_sampling(self):
## Calculate the expected error ratio for the Volume Sampling distribution for every dataset
res_list = np.zeros(self.versions_number+1)
for t in range(self.versions_number+1):
test_X = self.multi_X[t,:,:]
res_list[t] = expected_approximation_error_fro_for_sampling_scheme(test_X,test_X,self.k,self.real_dim)
return res_list
def get_expected_error_fro_for_effective_kernel_sampling(self):
## Calculate the expected error ratio for the Volume Sampling distribution for every dataset
res_list = np.zeros(self.versions_number+1)
test_eff_V = self.get_effective_kernel_from_orthogonal_matrix()
for t in range(self.versions_number+1):
test_X = self.multi_X[t,:,:]
test_U = test_eff_V[t,:,:].T
res_list[t] = expected_approximation_error_fro_for_sampling_scheme(test_X,test_U,self.k,self.real_dim)
return res_list
def get_tight_upper_bound_error_fro_for_projection_DPP(self):
res_list = np.zeros(self.versions_number+1)
for t in range(self.versions_number+1):
test_X = self.multi_X[t,:,:]
test_U = self.multi_Q[t,:,:].T
res_list[t] = tight_approximation_error_fro_for_sampling_scheme(test_X,test_U,self.k,self.real_dim)
return res_list
def get_max_diag_sum_T_matrices(self):
res_list = np.zeros((self.versions_number+1))
for t in range(self.versions_number+1):
test_X = self.multi_X[t,:,:]
_,_,test_V = np.linalg.svd(test_X, full_matrices=False)
test_V_k = test_V[0:self.k,:]
test_V_d_k = test_V[self.k:self.real_dim,:]
res_list[t] = 1+np.max(np.diag(get_the_matrix_sum_T_S(self.k,self.real_dim,test_V_k,test_V_d_k)))
return res_list
def get_max_spectrum_sum_T_matrices(self):
res_list = np.zeros((self.versions_number+1))
for t in range(self.versions_number+1):
test_X = self.multi_X[t,:,:]
_,_,test_V = np.linalg.svd(test_X, full_matrices=False)
test_V_k = test_V[0:self.k,:]
test_V_d_k = test_V[self.k:self.real_dim,:]
res_list[t] = 1+np.max(np.diag(get_the_matrix_sum_T_S(self.k,self.real_dim,test_V_k,test_V_d_k)))
return res_list
def get_expected_error_fro_for_projection_DPP(self):
## Calculate the expected error ratio for the Projection DPP distribution for every dataset
res_list = np.zeros(self.versions_number+1)
for t in range(self.versions_number+1):
test_X = self.multi_X[t,:,:]
test_U = self.multi_Q[t,:,:].T
res_list[t] = expected_approximation_error_fro_for_sampling_scheme(test_X,test_U,self.k,self.real_dim)
return res_list
def get_expected_error_spectral_for_volume_sampling(self):
## Calculate the expected error ratio for the Volume Sampling distribution for every dataset
res_list = np.zeros(self.versions_number+1)
for t in range(self.versions_number+1):
test_X = self.multi_X[t,:,:]
res_list[t] = expected_approximation_error_spectral_for_sampling_scheme(test_X,test_X,self.k,self.real_dim)
return res_list
def get_expected_error_spectral_for_projection_DPP(self):
## Calculate the expected error ratio for the Projection DPP distribution for every dataset
res_list = np.zeros(self.versions_number+1)
for t in range(self.versions_number+1):
test_X = self.multi_X[t,:,:]
test_U = self.multi_Q[t,:,:].T
res_list[t] = expected_approximation_error_spectral_for_sampling_scheme(test_X,test_U,self.k,self.real_dim)
return res_list
def get_upper_bound_error_for_projection_DPP(self):
## Calculate the expected error ratio for the Projection DPP distribution for every dataset
#res_list = np.zeros(self.versions_number+1)
res_list = []
for t in range(self.versions_number+1):
test_X = self.multi_X[t,:,:]
test_U = self.multi_Q[t,:,:].T
#res_list[t] = expected_upper_bound_for_projection_DPP(test_X,test_U,self.k,self.real_dim)
res_list.append( expected_upper_bound_for_projection_DPP(test_X,test_U,self.k,self.real_dim))
return res_list
def get_error_fro_for_deterministic_selection(self):
## Calculate the error ratio for the k-tuple selected by the deterministic algorithm for every dataset
res_list = np.zeros(self.versions_number+1)
for t in range(self.versions_number+1):
test_X = self.multi_X[t,:,:]
test_U = self.multi_Q[t,:,:].T
lv_scores_vector = np.diag(np.dot(np.transpose(test_U),test_U))
test_I_k = list(np.argsort(lv_scores_vector)[self.real_dim-self.k:self.real_dim])
_,test_Sigma,_ = np.linalg.svd(test_X, full_matrices=False)
res_list[t] = approximation_error_function_fro(test_Sigma,self.k,test_X,test_X[:,test_I_k])
#res_list.append(test_I_k)
return res_list
def get_error_spectral_for_deterministic_selection(self):
## Calculate the error ratio for the k-tuple selected by the deterministic algorithm for every dataset
res_list = np.zeros(self.versions_number+1)
for t in range(self.versions_number+1):
test_X = self.multi_X[t,:,:]
test_U = self.multi_Q[t,:,:].T
lv_scores_vector = np.diag(np.dot(np.transpose(test_U),test_U))
test_I_k = list(np.argsort(lv_scores_vector)[self.real_dim-self.k:self.real_dim])
_,test_Sigma,_ = np.linalg.svd(test_X, full_matrices=False)
res_list[t] = approximation_error_function_spectral(test_Sigma,self.k,test_X,test_X[:,test_I_k])
#res_list.append(test_I_k)
return res_list
def get_p_eff(self):
## A function that calculate the p_eff.
## It is a measure of the concentration of V_k. This is done for every dataset
res_list = np.zeros(self.versions_number+1)
for t in range(self.versions_number+1):
diag_Q_t = np.diag(np.dot(self.multi_Q[t,:,:],self.multi_Q[t,:,:].T))
#diag_Q_t = list(diag_Q_t[::-1].sort())
diag_Q_t = list(np.sort(diag_Q_t)[::-1])
p = self.real_dim
print(diag_Q_t)
while np.sum(diag_Q_t[0:p-1]) > float(self.k-1.0/2):
p = p-1
res_list[t] = p
return res_list
def get_sum_k_leverage_scores(self):
## A function that calculate the k-sum: the sum of the first k k-leverage scores. It is a measure of the concentration of V_k
## This is done for every dataset
res_list = np.zeros(self.versions_number+1)
for t in range(self.versions_number+1):
res_list[t] = estimate_sum_first_k_leverage_scores(self.multi_Q[t,:,:],self.k)
return res_list
def get_deterministic_upper_bound(self):
## A function that calculate the theoretical upper bound for the deterministic algorithm for every dataset
res_list = np.zeros(self.versions_number+1)
for t in range(self.versions_number+1):
res_list[t] = 1/(1+estimate_sum_first_k_leverage_scores(self.multi_Q[t,:,:],self.k)-self.k)
return res_list
def get_alpha_sum_k_leverage_scores(self,alpha):
## A function that calculate the theoretical upper bound for the deterministic algorithm for every dataset
res_list = np.zeros(self.versions_number+1)
#k_l = self.get_sum_k_leverage_scores()
for t in range(self.versions_number+1):
k_l = estimate_leverage_scores_from_orthogonal_matrix(self.multi_Q[t,:,:])[0:k]
func_k = np.power(np.linspace(1, k, num=k),alpha)
res_list[t] = np.dot(func_k,k_l)
return res_list | _____no_output_____ | MIT | .ipynb_checkpoints/Numerical_Analysis_Volume_Sampling_vs_Projection_DPP-ollld-checkpoint.ipynb | AyoubBelhadji/FrameBuilder |
III- 0 Parameters of the simultations | ## The dimensions of the design matrix X
N = 100 # The number of observations in the dataset
real_dim = 20 # The dimension of the dataset
## The low rank paramters
k = 2 # The rank of the low rank approximation
## The covariance matrix parameters
r = 6 # Just a parameter to control the number of non trivial singular values in the covariance matrix
mean = np.zeros((real_dim)) # The mean vector useful to generate U (X = UDV^T)
cov_test = 0.01*np.ones((real_dim-r)) # The "trivial" singular values in the covariance matrix (there are real_dim-r)
## The paramters of the matrix V
versions_number = 3 # The number of orthogonal matrices (and therefor datasets) (-1) generated by the algorithm above
lv_scores_vector = k/real_dim*np.ones(real_dim) # The vector of leverage scores (the last one)
l = [1,5,2,10]
ll = list(reversed(np.argsort(l)))
ll | _____no_output_____ | MIT | .ipynb_checkpoints/Numerical_Analysis_Volume_Sampling_vs_Projection_DPP-ollld-checkpoint.ipynb | AyoubBelhadji/FrameBuilder |
III-1 The influence of the spectrum In this subsection we compare the Volume Sampling distribution to the projection DPP distribution and the deterministic algorithm of [] for different profiles of the spectrum with k-leverage scores profile fixed. In other words, if we note $$X = UDV^{T}$$ We keep V_{k} constant and we investigate the effect of D. III-1-1 The case of a non-projection spectrumWe mean by a projection spectrum matrix, a matrix with equal the first k singular values.We observe that the two distributions are very similar.... \todo{reword} | cov_1 = np.diag(np.concatenate(([100,100,1,1,1,1],cov_test)))
versions_list = [20,19,18,17]
NAL_1 = Numrerical_Analysis_DPP(N,real_dim,r,k,versions_number,mean,cov_1,lv_scores_vector,versions_list)
projection_DPP_res_fro_1 = NAL_1.get_expected_error_fro_for_projection_DPP()
volume_sampling_res_fro_1 = NAL_1.get_expected_error_fro_for_volume_sampling()
deterministic_selection_res_fro_1 = NAL_1.get_error_fro_for_deterministic_selection()
projection_DPP_res_spectral_1 = NAL_1.get_expected_error_spectral_for_projection_DPP()
volume_sampling_res_spectral_1 = NAL_1.get_expected_error_spectral_for_volume_sampling()
deterministic_selection_res_spectral_1 = NAL_1.get_error_spectral_for_deterministic_selection()
effective_kernel_sampling_res_fro_1 = NAL_1.get_expected_error_fro_for_effective_kernel_sampling()
#sss = NAL_1.get_effective_kernel_from_orthogonal_matrix()
#p_eff_res_1 = NAL_1.get_p_eff()
upper_tight_bound_projection_DPP_res_fro_1 = NAL_1.get_tight_upper_bound_error_fro_for_projection_DPP()
alpha_sum_res_1 = NAL_1.get_alpha_sum_k_leverage_scores(1)
sum_U_res_1 = NAL_1.get_sum_k_leverage_scores()
deterministic_upper_bound_res_1 = NAL_1.get_deterministic_upper_bound()
expected_upper_bound_res_1 = NAL_1.get_upper_bound_error_for_projection_DPP()
multi_Q_1 = NAL_1.multi_Q[1,:,:].T
p_eff_res_1 = NAL_1.get_p_eff()
eff_kernel_upper_bound_1 = 1+ (p_eff_res_1-k)/(real_dim-k)*(k+1)
eff_kernel_upper_bound
print(k*(real_dim-k+1))
sum_T_matrices = NAL_1.get_sum_T_matrices()
pd_1 = pd.DataFrame(
{'k-sum (ratio)': sum_U_res_1/k,
'p_eff':p_eff_res_1,
'alpha k-sum': alpha_sum_res_1,
'Expected Upper Bound for Projection DPP': expected_upper_bound_res_1,
'Volume Sampling(Fro)': volume_sampling_res_fro_1,
'Projection DPP(Fro)': projection_DPP_res_fro_1,
'Effective kernel(Fro)' : effective_kernel_sampling_res_fro_1,
'Effective kernel upper bound (Fro)':eff_kernel_upper_bound_1,
'Very sharp approximation of Projection DPP(Fro)': upper_tight_bound_projection_DPP_res_fro_1,
'Deterministic Algorithm(Fro)': deterministic_selection_res_fro_1,
'Volume Sampling(Spectral)': volume_sampling_res_spectral_1,
'Projection DPP(Spectral)': projection_DPP_res_spectral_1,
'Deterministic Algorithm(Spectral)': deterministic_selection_res_spectral_1,
'Deterministic Upper Bound': deterministic_upper_bound_res_1
})
pd_1 = pd_1[['k-sum (ratio)','p_eff', 'alpha k-sum','Expected Upper Bound for Projection DPP','Volume Sampling(Fro)','Projection DPP(Fro)','Effective kernel(Fro)','Effective kernel upper bound (Fro)','Very sharp approximation of Projection DPP(Fro)','Deterministic Algorithm(Fro)','Volume Sampling(Spectral)','Projection DPP(Spectral)','Deterministic Algorithm(Spectral)','Deterministic Upper Bound']]
p_eff_res_1[3]
#'1+Largest eigenvalue of sum_T': sum_T_matrices,
pd_1 | _____no_output_____ | MIT | .ipynb_checkpoints/Numerical_Analysis_Volume_Sampling_vs_Projection_DPP-ollld-checkpoint.ipynb | AyoubBelhadji/FrameBuilder |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.