markdown stringlengths 0 1.02M | code stringlengths 0 832k | output stringlengths 0 1.02M | license stringlengths 3 36 | path stringlengths 6 265 | repo_name stringlengths 6 127 |
|---|---|---|---|---|---|
Observations:* The expected error is always smaller under the Projection DPP distribution compared to the Volume Sampling distribution.* The expected error for the Volume Sampling distribution is constant for a contant D* However the expected error for the Projection DPP distribution depends on the k-sum* For X_0 and X_1, the profile of the k-leverage scores are highly concentrated (k-sum > k-1) thus epsilon is smaller than 1, in this regime the determinstic algorithm have the lower approximation error and it performs better than expected (the theoretical bound is 1/(1-epsilon).* However, for the other datasets, the (k-sum 1 and the deterministic algorithm have no guarantee in this regime: we observe that the approximation error for the deterministic algorithm can be very high in this regime. Recall:We recall here some geometrical properties of the matrices $$X_i$$$$X_i = UD_{j}V_{i}$$Where for every i, the first k columns of $$V_{i}$$ are the $$Q_{i}$$ while the other columns are gernerated randomly | previous_Q = NAL_1.multi_Q
lv_0 = estimate_leverage_scores_from_orthogonal_matrix(previous_Q[0,:,:])
lv_1 = estimate_leverage_scores_from_orthogonal_matrix(previous_Q[1,:,:])
lv_2 = estimate_leverage_scores_from_orthogonal_matrix(previous_Q[2,:,:])
lv_3 = estimate_leverage_scores_from_orthogonal_matrix(previous_Q[3,:,:])
lv_4 = estimate_leverage_scores_from_orthogonal_matrix(previous_Q[4,:,:])
lv_5 = estimate_leverage_scores_from_orthogonal_matrix(previous_Q[5,:,:])
index_list = list(range(real_dim)) | _____no_output_____ | MIT | .ipynb_checkpoints/Numerical_Analysis_Volume_Sampling_vs_Projection_DPP-ollld-checkpoint.ipynb | AyoubBelhadji/FrameBuilder |
In this example the objective is Q and the initialization is Q_0 (the rectangular identity)We have with respect to the Schur-order (or the majorization):$$Q = Q_5 \prec_{S} Q_4 \prec_{S} Q_3 \prec_{S} Q_2 \prec_{S} Q_1 \prec_{S} Q_0 $$ | plt.plot(index_list[0:10], lv_0[0:10], 'c--',index_list[0:10], lv_1[0:10], 'k--', index_list[0:10], lv_2[0:10], 'r--', index_list[0:10], lv_3[0:10], 'b--',index_list[0:10], lv_4[0:10], 'g--',index_list[0:10], lv_5[0:10], 'y--')
plt.xlabel('index')
plt.ylabel('leverage score')
cyan_patch = mpatches.Patch(color='cyan', label='Q_0')
black_patch = mpatches.Patch(color='black', label='Q_1')
red_patch = mpatches.Patch(color='red', label='Q_2')
blue_patch = mpatches.Patch(color='blue', label='Q_3')
green_patch = mpatches.Patch(color='green', label='Q_4')
yellow_patch = mpatches.Patch(color='yellow', label='Q = Q_5')
plt.legend(handles=[cyan_patch,black_patch,red_patch,blue_patch,green_patch,yellow_patch])
plt.show() | _____no_output_____ | MIT | .ipynb_checkpoints/Numerical_Analysis_Volume_Sampling_vs_Projection_DPP-ollld-checkpoint.ipynb | AyoubBelhadji/FrameBuilder |
III-1-2 The case of a projection spectrumWe mean by a projection spectrum matrix, a matrix with equal the first k singular values.We observe that the two distributions are very similar.... \todo{reword} | cov_2 = np.diag(np.concatenate(([1000,1000,1000,1,0.1],cov_test)))
NAL_2 = Numrerical_Analysis_DPP(N,real_dim,r,k,versions_number,mean,cov_2,lv_scores_vector)
projection_DPP_res_2 = NAL_2.get_expected_error_for_projection_DPP()
volume_sampling_res_2 = NAL_2.get_expected_error_for_volume_sampling()
deterministic_selection_res_2 = NAL_1.get_error_for_deterministic_selection()
sum_U_res_2 = NAL_2.get_sum_k_leverage_scores()
deterministic_upper_bound_res_2 = NAL_2.get_deterministic_upper_bound()
results = [["Dataset","Using Volume Sampling","Using Projection DPP","k-sum","1/(1-epsilon)","Using Deterministic Algorithm"],["X_0",volume_sampling_res_2[0],projection_DPP_res_2[0],sum_U_res_2[0],deterministic_upper_bound_res_2[0],deterministic_selection_res_2[0]],["X_1",volume_sampling_res_2[1],projection_DPP_res_2[1],sum_U_res_2[1],deterministic_upper_bound_res_2[1],deterministic_selection_res_2[1]],
["X_2",volume_sampling_res_2[2],projection_DPP_res_2[2],sum_U_res_2[2],deterministic_upper_bound_res_2[2],deterministic_selection_res_2[2]],["X_3",volume_sampling_res_2[3],projection_DPP_res_2[3],sum_U_res_2[3],deterministic_upper_bound_res_2[3],deterministic_selection_res_2[3]],["X_4",volume_sampling_res_2[4],projection_DPP_res_2[4],sum_U_res_2[4],deterministic_upper_bound_res_2[4],deterministic_selection_res_2[4]],["X_5",volume_sampling_res_2[5],projection_DPP_res_2[5],sum_U_res_2[5],deterministic_upper_bound_res_2[5],deterministic_selection_res_2[5]]]
display(HTML(
'<center><b>The expected approximation error (divided by the optimal error) according to a sampling scheme for different distribution</b><br><table><tr>{}</tr></table>'.format(
'</tr><tr>'.join(
'<td>{}</td>'.format('</td><td>'.join(str(_) for _ in row)) for row in results)
)
)) | [ 0.3 0.3 0.3 0.3 0.3 0.3 0.3 0.3 0.3 0.3 0.3 0.3 0.3 0.3 0.3
0.3 0.3 0.3 0.3 0.3]
[ 1. 1. 1. 1. 1. 1. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0.
0. 0.]
[-0.7 -0.7 -0.7 -0.7 -0.7 -0.7 0.3 0.3 0.3 0.3 0.3 0.3 0.3 0.3 0.3
0.3 0.3 0.3 0.3 0.3]
5
6
| MIT | .ipynb_checkpoints/Numerical_Analysis_Volume_Sampling_vs_Projection_DPP-ollld-checkpoint.ipynb | AyoubBelhadji/FrameBuilder |
III-2 The influence of the "spread" of VIn this section we investigate the influence of the "spread" (to be defined formally) of the cloud of points. We can change this "spread" by changing the initialization of the generator of orthogonal matrices: we replace the rectangular identity by "other" orthogonal matrices. Technically, this boils down to change the generator mode in the constructor call from "nonspread" to "spread". | np.power(np.linspace(1, k, num=k),2)
matrices_array = [ np.zeros((4,4)) for comb in combinations(range(5),4)]
matrix_sum = np.sum(matrices_array)
matrix_sum
matrices_array | _____no_output_____ | MIT | .ipynb_checkpoints/Numerical_Analysis_Volume_Sampling_vs_Projection_DPP-ollld-checkpoint.ipynb | AyoubBelhadji/FrameBuilder |
Convolutional Neural Network ExampleBuild a convolutional neural network with TensorFlow.- Author: Aymeric Damien- Project: https://github.com/aymericdamien/TensorFlow-Examples/These lessons are adapted from [aymericdamien TensorFlow tutorials](https://github.com/aymericdamien/TensorFlow-Examples) / [GitHub](https://github.com/aymericdamien/TensorFlow-Examples) which are published under the [MIT License](https://github.com/Hvass-Labs/TensorFlow-Tutorials/blob/master/LICENSE) which allows very broad use for both academic and commercial purposes. CNN Overview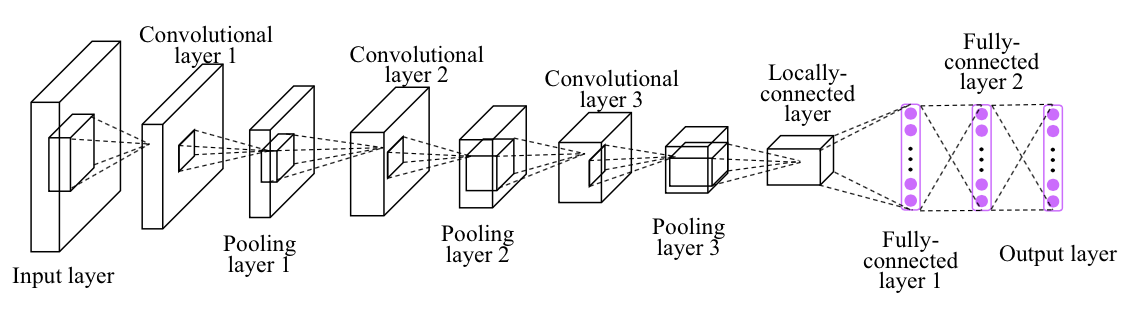 MNIST Dataset OverviewThis example is using MNIST handwritten digits. The dataset contains 60,000 examples for training and 10,000 examples for testing. The digits have been size-normalized and centered in a fixed-size image (28x28 pixels) with values from 0 to 1. For simplicity, each image has been flattened and converted to a 1-D numpy array of 784 features (28*28).More info: http://yann.lecun.com/exdb/mnist/ | from __future__ import division, print_function, absolute_import
import tensorflow as tf
# Import MNIST data
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets("/tmp/data/", one_hot=True)
# Training Parameters
learning_rate = 0.001
num_steps = 500
batch_size = 128
display_step = 10
# Network Parameters
num_input = 784 # MNIST data input (img shape: 28*28)
num_classes = 10 # MNIST total classes (0-9 digits)
dropout = 0.75 # Dropout, probability to keep units
# tf Graph input
X = tf.placeholder(tf.float32, [None, num_input])
Y = tf.placeholder(tf.float32, [None, num_classes])
keep_prob = tf.placeholder(tf.float32) # dropout (keep probability)
# Create some wrappers for simplicity
def conv2d(x, W, b, strides=1):
# Conv2D wrapper, with bias and relu activation
x = tf.nn.conv2d(x, W, strides=[1, strides, strides, 1], padding='SAME')
x = tf.nn.bias_add(x, b)
return tf.nn.relu(x)
def maxpool2d(x, k=2):
# MaxPool2D wrapper
return tf.nn.max_pool(x, ksize=[1, k, k, 1], strides=[1, k, k, 1],
padding='SAME')
# Create model
def conv_net(x, weights, biases, dropout):
# MNIST data input is a 1-D vector of 784 features (28*28 pixels)
# Reshape to match picture format [Height x Width x Channel]
# Tensor input become 4-D: [Batch Size, Height, Width, Channel]
x = tf.reshape(x, shape=[-1, 28, 28, 1])
# Convolution Layer
conv1 = conv2d(x, weights['wc1'], biases['bc1'])
# Max Pooling (down-sampling)
conv1 = maxpool2d(conv1, k=2)
# Convolution Layer
conv2 = conv2d(conv1, weights['wc2'], biases['bc2'])
# Max Pooling (down-sampling)
conv2 = maxpool2d(conv2, k=2)
# Fully connected layer
# Reshape conv2 output to fit fully connected layer input
fc1 = tf.reshape(conv2, [-1, weights['wd1'].get_shape().as_list()[0]])
fc1 = tf.add(tf.matmul(fc1, weights['wd1']), biases['bd1'])
fc1 = tf.nn.relu(fc1)
# Apply Dropout
fc1 = tf.nn.dropout(fc1, dropout)
# Output, class prediction
out = tf.add(tf.matmul(fc1, weights['out']), biases['out'])
return out
# Store layers weight & bias
weights = {
# 5x5 conv, 1 input, 32 outputs
'wc1': tf.Variable(tf.random_normal([5, 5, 1, 32])),
# 5x5 conv, 32 inputs, 64 outputs
'wc2': tf.Variable(tf.random_normal([5, 5, 32, 64])),
# fully connected, 7*7*64 inputs, 1024 outputs
'wd1': tf.Variable(tf.random_normal([7*7*64, 1024])),
# 1024 inputs, 10 outputs (class prediction)
'out': tf.Variable(tf.random_normal([1024, num_classes]))
}
biases = {
'bc1': tf.Variable(tf.random_normal([32])),
'bc2': tf.Variable(tf.random_normal([64])),
'bd1': tf.Variable(tf.random_normal([1024])),
'out': tf.Variable(tf.random_normal([num_classes]))
}
# Construct model
logits = conv_net(X, weights, biases, keep_prob)
prediction = tf.nn.softmax(logits)
# Define loss and optimizer
loss_op = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(
logits=logits, labels=Y))
optimizer = tf.train.AdamOptimizer(learning_rate=learning_rate)
train_op = optimizer.minimize(loss_op)
# Evaluate model
correct_pred = tf.equal(tf.argmax(prediction, 1), tf.argmax(Y, 1))
accuracy = tf.reduce_mean(tf.cast(correct_pred, tf.float32))
# Initialize the variables (i.e. assign their default value)
init = tf.global_variables_initializer()
# Start training
with tf.Session() as sess:
# Run the initializer
sess.run(init)
for step in range(1, num_steps+1):
batch_x, batch_y = mnist.train.next_batch(batch_size)
# Run optimization op (backprop)
sess.run(train_op, feed_dict={X: batch_x, Y: batch_y, keep_prob: dropout})
if step % display_step == 0 or step == 1:
# Calculate batch loss and accuracy
loss, acc = sess.run([loss_op, accuracy], feed_dict={X: batch_x,
Y: batch_y,
keep_prob: 1.0})
print("Step " + str(step) + ", Minibatch Loss= " + \
"{:.4f}".format(loss) + ", Training Accuracy= " + \
"{:.3f}".format(acc))
print("Optimization Finished!")
# Calculate accuracy for 256 MNIST test images
print("Testing Accuracy:", \
sess.run(accuracy, feed_dict={X: mnist.test.images[:256],
Y: mnist.test.labels[:256],
keep_prob: 1.0}))
| Step 1, Minibatch Loss= 63763.3047, Training Accuracy= 0.141
Step 10, Minibatch Loss= 26429.6680, Training Accuracy= 0.242
Step 20, Minibatch Loss= 12171.8584, Training Accuracy= 0.586
Step 30, Minibatch Loss= 6306.6318, Training Accuracy= 0.734
Step 40, Minibatch Loss= 5113.7583, Training Accuracy= 0.711
Step 50, Minibatch Loss= 4022.2131, Training Accuracy= 0.805
Step 60, Minibatch Loss= 3125.4949, Training Accuracy= 0.867
Step 70, Minibatch Loss= 2225.4875, Training Accuracy= 0.875
Step 80, Minibatch Loss= 1843.3540, Training Accuracy= 0.867
Step 90, Minibatch Loss= 1715.7744, Training Accuracy= 0.875
Step 100, Minibatch Loss= 2611.2708, Training Accuracy= 0.906
Step 110, Minibatch Loss= 4804.0913, Training Accuracy= 0.875
Step 120, Minibatch Loss= 1067.5258, Training Accuracy= 0.938
Step 130, Minibatch Loss= 2519.1514, Training Accuracy= 0.898
Step 140, Minibatch Loss= 2687.9292, Training Accuracy= 0.906
Step 150, Minibatch Loss= 1983.4077, Training Accuracy= 0.938
Step 160, Minibatch Loss= 2844.6553, Training Accuracy= 0.930
Step 170, Minibatch Loss= 3602.2524, Training Accuracy= 0.914
Step 180, Minibatch Loss= 175.3922, Training Accuracy= 0.961
Step 190, Minibatch Loss= 645.1918, Training Accuracy= 0.945
Step 200, Minibatch Loss= 1147.6567, Training Accuracy= 0.938
Step 210, Minibatch Loss= 1140.4148, Training Accuracy= 0.914
Step 220, Minibatch Loss= 1572.8756, Training Accuracy= 0.906
Step 230, Minibatch Loss= 1292.9274, Training Accuracy= 0.898
Step 240, Minibatch Loss= 1501.4623, Training Accuracy= 0.953
Step 250, Minibatch Loss= 1908.2997, Training Accuracy= 0.898
Step 260, Minibatch Loss= 2182.2380, Training Accuracy= 0.898
Step 270, Minibatch Loss= 487.5807, Training Accuracy= 0.961
Step 280, Minibatch Loss= 1284.1130, Training Accuracy= 0.945
Step 290, Minibatch Loss= 1232.4919, Training Accuracy= 0.891
Step 300, Minibatch Loss= 1198.8336, Training Accuracy= 0.945
Step 310, Minibatch Loss= 2010.5345, Training Accuracy= 0.906
Step 320, Minibatch Loss= 786.3917, Training Accuracy= 0.945
Step 330, Minibatch Loss= 1408.3556, Training Accuracy= 0.898
Step 340, Minibatch Loss= 1453.7538, Training Accuracy= 0.953
Step 350, Minibatch Loss= 999.8901, Training Accuracy= 0.906
Step 360, Minibatch Loss= 914.3958, Training Accuracy= 0.961
Step 370, Minibatch Loss= 488.0052, Training Accuracy= 0.938
Step 380, Minibatch Loss= 1070.8710, Training Accuracy= 0.922
Step 390, Minibatch Loss= 151.4658, Training Accuracy= 0.961
Step 400, Minibatch Loss= 555.3539, Training Accuracy= 0.953
Step 410, Minibatch Loss= 765.5746, Training Accuracy= 0.945
Step 420, Minibatch Loss= 326.9393, Training Accuracy= 0.969
Step 430, Minibatch Loss= 530.8968, Training Accuracy= 0.977
Step 440, Minibatch Loss= 463.3909, Training Accuracy= 0.977
Step 450, Minibatch Loss= 362.2226, Training Accuracy= 0.977
Step 460, Minibatch Loss= 414.0034, Training Accuracy= 0.953
Step 470, Minibatch Loss= 583.4587, Training Accuracy= 0.945
Step 480, Minibatch Loss= 566.1262, Training Accuracy= 0.969
Step 490, Minibatch Loss= 691.1143, Training Accuracy= 0.961
Step 500, Minibatch Loss= 282.8893, Training Accuracy= 0.984
Optimization Finished!
Testing Accuracy: 0.976562
| MIT | Deep_Learning/TensorFlow-aymericdamien/notebooks/3_NeuralNetworks/convolutional_network_raw.ipynb | Chau-Xochitl/INFO_7390 |
Google Cloud CMIP6 Public Data: Basic Python ExampleThis notebooks shows how to query the catalog and load the data using python | from matplotlib import pyplot as plt
import numpy as np
import pandas as pd
import xarray as xr
import zarr
import fsspec
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
plt.rcParams['figure.figsize'] = 12, 6 | _____no_output_____ | MIT | basic_search_and_load.ipynb | pangeo-gallery/cmip6 |
Browse CatalogThe data catatalog is stored as a CSV file. Here we read it with Pandas. | df = pd.read_csv('https://storage.googleapis.com/cmip6/cmip6-zarr-consolidated-stores.csv')
df.head() | _____no_output_____ | MIT | basic_search_and_load.ipynb | pangeo-gallery/cmip6 |
The columns of the dataframe correspond to the CMI6 controlled vocabulary. A beginners' guide to these terms is available in [this document](https://docs.google.com/document/d/1yUx6jr9EdedCOLd--CPdTfGDwEwzPpCF6p1jRmqx-0Q). Here we filter the data to find monthly surface air temperature for historical experiments. | df_ta = df.query("activity_id=='CMIP' & table_id == 'Amon' & variable_id == 'tas' & experiment_id == 'historical'")
df_ta | _____no_output_____ | MIT | basic_search_and_load.ipynb | pangeo-gallery/cmip6 |
Now we do further filtering to find just the models from NCAR. | df_ta_ncar = df_ta.query('institution_id == "NCAR"')
df_ta_ncar | _____no_output_____ | MIT | basic_search_and_load.ipynb | pangeo-gallery/cmip6 |
Load DataNow we will load a single store using fsspec, zarr, and xarray. | # get the path to a specific zarr store (the first one from the dataframe above)
zstore = df_ta_ncar.zstore.values[-1]
print(zstore)
# create a mutable-mapping-style interface to the store
mapper = fsspec.get_mapper(zstore)
# open it using xarray and zarr
ds = xr.open_zarr(mapper, consolidated=True)
ds | _____no_output_____ | MIT | basic_search_and_load.ipynb | pangeo-gallery/cmip6 |
Plot a map from a specific date. | ds.tas.sel(time='1950-01').squeeze().plot() | _____no_output_____ | MIT | basic_search_and_load.ipynb | pangeo-gallery/cmip6 |
Create a timeseries of global-average surface air temperature. For this we need the area weighting factor for each gridpoint. | df_area = df.query("variable_id == 'areacella' & source_id == 'CESM2'")
ds_area = xr.open_zarr(fsspec.get_mapper(df_area.zstore.values[0]), consolidated=True)
ds_area
total_area = ds_area.areacella.sum(dim=['lon', 'lat'])
ta_timeseries = (ds.tas * ds_area.areacella).sum(dim=['lon', 'lat']) / total_area
ta_timeseries | _____no_output_____ | MIT | basic_search_and_load.ipynb | pangeo-gallery/cmip6 |
By default the data are loaded lazily, as Dask arrays. Here we trigger computation explicitly. | %time ta_timeseries.load()
ta_timeseries.plot(label='monthly')
ta_timeseries.rolling(time=12).mean().plot(label='12 month rolling mean')
plt.legend()
plt.title('Global Mean Surface Air Temperature') | _____no_output_____ | MIT | basic_search_and_load.ipynb | pangeo-gallery/cmip6 |
Figures for the section on approximately computing the stream structure | # Load the smooth and peppered stream
sdf_smooth= gd1_util.setup_gd1model()
pepperfilename= 'gd1pepper.pkl'
if os.path.exists(pepperfilename):
with open(pepperfilename,'rb') as savefile:
sdf_pepper= pickle.load(savefile)
else:
timpacts= simulate_streampepper.parse_times('256sampling',9.)
sdf_pepper= gd1_util.setup_gd1model(timpact=timpacts,
hernquist=True)
save_pickles(pepperfilename,sdf_pepper) | galpyWarning: WARNING: Rnorm keyword input to streamdf is deprecated in favor of the standard ro keyword
galpyWarning: WARNING: Vnorm keyword input to streamdf is deprecated in favor of the standard vo keyword
galpyWarning: Using C implementation to integrate orbits
| BSD-3-Clause | meanOperpAndApproxImpacts.ipynb | jobovy/streamgap-pepper |
Is the mean perpendicular frequency close to zero? | # Sampling functions
massrange=[5.,9.]
plummer= False
Xrs= 5.
nsubhalo= simulate_streampepper.nsubhalo
rs= simulate_streampepper.rs
dNencdm= simulate_streampepper.dNencdm
sample_GM= lambda: (10.**((-0.5)*massrange[0])\
+(10.**((-0.5)*massrange[1])\
-10.**((-0.5)*massrange[0]))\
*numpy.random.uniform())**(1./(-0.5))\
/bovy_conversion.mass_in_msol(V0,R0)
rate_range= numpy.arange(massrange[0]+0.5,massrange[1]+0.5,1)
rate= numpy.sum([dNencdm(sdf_pepper,10.**r,Xrs=Xrs,
plummer=plummer)
for r in rate_range])
sample_rs= lambda x: rs(x*bovy_conversion.mass_in_1010msol(V0,R0)*10.**10.,
plummer=plummer)
numpy.random.seed(2)
sdf_pepper.simulate(rate=rate,sample_GM=sample_GM,sample_rs=sample_rs,Xrs=Xrs)
n= 100000
aa_mock_per= sdf_pepper.sample(n=n,returnaAdt=True)
dO= numpy.dot(aa_mock_per[0].T-sdf_pepper._progenitor_Omega,
sdf_pepper._sigomatrixEig[1][:,sdf_pepper._sigomatrixEigsortIndx])
dO[:,2]*= sdf_pepper._sigMeanSign
da= numpy.dot(aa_mock_per[1].T-sdf_pepper._progenitor_angle,
sdf_pepper._sigomatrixEig[1][:,sdf_pepper._sigomatrixEigsortIndx])
da[:,2]*= sdf_pepper._sigMeanSign
apar= da[:,2]
xs= numpy.linspace(0.,1.5,1001)
mO_unp= numpy.array([sdf_smooth.meanOmega(x,oned=True,use_physical=False) for x in xs])
mOint= interpolate.InterpolatedUnivariateSpline(xs,mO_unp,k=3)
mOs= mOint(apar)
frac= 0.02
alpha=0.01
linecolor='0.65'
bovy_plot.bovy_print(axes_labelsize=18.,xtick_labelsize=14.,ytick_labelsize=14.)
figsize(12,4)
subplot(1,3,1)
bovy_plot.bovy_plot(apar[::3],dO[::3,2]/mOs[::3]-1,'k.',alpha=alpha*2,gcf=True,
rasterized=True,xrange=[0.,1.5],yrange=[-1.2,1.2])
z= lowess(dO[:,2]/mOs-1,apar,frac=frac)
plot(z[::100,0],z[::100,1],color=linecolor,lw=2.5)
#xlim(0.,1.5)
#ylim(-1.2,1.2)
xlabel(r'$\Delta\theta_\parallel$')
bovy_plot.bovy_text(r'$\Delta\Omega_\parallel/\langle\Delta\Omega^0_\parallel\rangle-1$',top_left=True,
size=18.)
subplot(1,3,2)
bovy_plot.bovy_plot(apar[::3],dO[::3,1]/mOs[::3],'k.',alpha=alpha*2,gcf=True,
rasterized=True,xrange=[0.,1.5],yrange=[-0.05,0.05])
z= lowess(dO[:,1]/mOs,apar,frac=frac)
plot(z[::100,0],z[::100,1],color=linecolor,lw=2.5)
#xlim(0.,1.5)
#ylim(-0.05,0.05)
xlabel(r'$\Delta\theta_\parallel$')
bovy_plot.bovy_text(r'$\Delta\Omega_{\perp,1}/\langle\Delta\Omega^0_\parallel\rangle$',top_left=True,
size=18.)
subplot(1,3,3)
bovy_plot.bovy_plot(apar[::3],dO[::3,0]/mOs[::3],'k.',alpha=alpha,gcf=True,
rasterized=True,xrange=[0.,1.5],yrange=[-0.05,0.05])
z= lowess(dO[:,0]/mOs,apar,frac=frac)
plot(z[::100,0],z[::100,1],color=linecolor,lw=2.5)
#xlim(0.,1.5)
#ylim(-0.05,0.05)
xlabel(r'$\Delta\theta_\parallel$')
bovy_plot.bovy_text(r'$\Delta\Omega_{\perp,2}/\langle\Delta\Omega^0_\parallel\rangle$',top_left=True,
size=18.)
if save_figures:
tight_layout()
bovy_plot.bovy_end_print(os.path.join(os.getenv('PAPERSDIR'),'2016-stream-stats','gd1like_meanOparOperp.pdf'))
print "This stream had %i impacts" % len(sdf_pepper._GM) | This stream had 61 impacts
| BSD-3-Clause | meanOperpAndApproxImpacts.ipynb | jobovy/streamgap-pepper |
Test the single-impact approximations | # Setup a single, large impact
m= 10.**8.
GM= 10**8./bovy_conversion.mass_in_msol(V0,R0)
timpactIndx= numpy.argmin(numpy.fabs(numpy.array(sdf_pepper._uniq_timpact)-1.3/bovy_conversion.time_in_Gyr(V0,R0)))
# Load the single-impact stream
gapfilename= 'gd1single.pkl'
if os.path.exists(gapfilename):
with open(gapfilename,'rb') as savefile:
sdf_gap= pickle.load(savefile)
else:
sdf_gap= gd1_util.setup_gd1model(hernquist=True,
singleImpact=True,
impactb=0.5*rs(m),
subhalovel=numpy.array([-25.,155.,30.])/V0,
impact_angle=0.6,
timpact=sdf_pepper._uniq_timpact[timpactIndx],
GM=GM,rs=rs(m))
save_pickles(gapfilename,sdf_gap)
n= 100000
aa_mock_per= sdf_gap.sample(n=n,returnaAdt=True)
dO= numpy.dot(aa_mock_per[0].T-sdf_gap._progenitor_Omega,
sdf_gap._sigomatrixEig[1][:,sdf_gap._sigomatrixEigsortIndx])
dO[:,2]*= sdf_gap._sigMeanSign
da= numpy.dot(aa_mock_per[1].T-sdf_gap._progenitor_angle,
sdf_gap._sigomatrixEig[1][:,sdf_gap._sigomatrixEigsortIndx])
da[:,2]*= sdf_gap._sigMeanSign
num= True
apar= numpy.arange(0.,sdf_smooth.length()+0.003,0.003)
dens_unp= numpy.array([sdf_smooth._density_par(x) for x in apar])
dens_approx= numpy.array([sdf_gap.density_par(x,approx=True) for x in apar])
dens_approx_higherorder= numpy.array([sdf_gap._density_par(x,approx=True,higherorder=True) for x in apar])
# normalize
dens_unp= dens_unp/numpy.sum(dens_unp)/(apar[1]-apar[0])
dens_approx= dens_approx/numpy.sum(dens_approx)/(apar[1]-apar[0])
dens_approx_higherorder= dens_approx_higherorder/numpy.sum(dens_approx_higherorder)/(apar[1]-apar[0])
if num:
dens_num= numpy.array([sdf_gap.density_par(x,approx=False) for x in apar])
dens_num= dens_num/numpy.sum(dens_num)/(apar[1]-apar[0])
bovy_plot.bovy_print(axes_labelsize=18.,xtick_labelsize=14.,ytick_labelsize=14.)
figsize(6,7)
axTop= pyplot.axes([0.15,0.3,0.825,0.65])
fig= pyplot.gcf()
fig.sca(axTop)
bovy_plot.bovy_plot(apar,dens_approx,lw=2.5,gcf=True,
color='k',
xrange=[0.,1.],
yrange=[0.,2.24],
ylabel=r'$\mathrm{density}$')
plot(apar,dens_unp,lw=3.5,color='k',ls='--',zorder=0)
nullfmt = NullFormatter() # no labels
axTop.xaxis.set_major_formatter(nullfmt)
dum= hist(da[:,2],bins=101,normed=True,range=[apar[0],apar[-1]],
histtype='step',color='0.55',zorder=0,lw=3.)
axBottom= pyplot.axes([0.15,0.1,0.825,0.2])
fig= pyplot.gcf()
fig.sca(axBottom)
bovy_plot.bovy_plot(apar,100.*(dens_approx_higherorder-dens_approx)/dens_approx_higherorder,
lw=2.5,gcf=True,color='k',
xrange=[0.,1.],
yrange=[-0.145,0.145],
zorder=2,
xlabel=r'$\Delta \theta_\parallel$',
ylabel=r'$\mathrm{relative\ difference\ in}\ \%$')
if num:
plot(apar,100.*(dens_num-dens_approx_higherorder)/dens_approx_higherorder,
lw=2.5,zorder=1,color='0.55')
# label
aparIndx= numpy.argmin(numpy.fabs(apar-0.64))
plot([0.45,apar[aparIndx]],[0.06,(100.*(dens_approx_higherorder-dens_approx)/dens_approx_higherorder)[aparIndx]],
'k',lw=1.)
bovy_plot.bovy_text(0.1,0.07,r'$\mathrm{higher\!\!-\!\!order\ minus\ linear}$',size=17.)
if num:
aparIndx= numpy.argmin(numpy.fabs(apar-0.62))
plot([0.45,apar[aparIndx]],[-0.07,(100.*(dens_num-dens_approx_higherorder)/dens_approx_higherorder)[aparIndx]],
'k',lw=1.)
bovy_plot.bovy_text(0.05,-0.12,r'$\mathrm{numerical\ minus\ higher\!\!-\!\!order}$',size=17.)
if save_figures:
bovy_plot.bovy_end_print(os.path.join(os.getenv('PAPERSDIR'),'2016-stream-stats','gd1likeSingle_densapprox.pdf'))
mO_unp= numpy.array([sdf_smooth.meanOmega(x,oned=True) for x in apar])\
*bovy_conversion.freq_in_Gyr(V0,R0)
mO_approx= numpy.array([sdf_gap.meanOmega(x,approx=True,oned=True) for x in apar])\
*bovy_conversion.freq_in_Gyr(V0,R0)
mO_approx_higherorder= numpy.array([sdf_gap.meanOmega(x,oned=True,approx=True,higherorder=True) for x in apar])\
*bovy_conversion.freq_in_Gyr(V0,R0)
if num:
mO_num= numpy.array([sdf_gap.meanOmega(x,approx=False,oned=True) for x in apar])\
*bovy_conversion.freq_in_Gyr(V0,R0)
frac= 0.005
alpha=0.01
linecolor='0.65'
bovy_plot.bovy_print(axes_labelsize=18.,xtick_labelsize=14.,ytick_labelsize=14.)
figsize(6,7)
axTop= pyplot.axes([0.15,0.3,0.825,0.65])
fig= pyplot.gcf()
fig.sca(axTop)
bovy_plot.bovy_plot(apar,mO_approx,lw=2.5,gcf=True,
color='k',
xrange=[0.,1.],
yrange=[0.,0.2],
ylabel=r'$\Delta \Omega_\parallel\,(\mathrm{Gyr}^{-1})$')
plot(apar,mO_unp,lw=2.5,color='k',ls='--')
plot(da[::3,2],dO[::3,2]*bovy_conversion.freq_in_Gyr(V0,R0),
'k.',alpha=alpha*2,rasterized=True)
nullfmt = NullFormatter() # no labels
axTop.xaxis.set_major_formatter(nullfmt)
axBottom= pyplot.axes([0.15,0.1,0.825,0.2])
fig= pyplot.gcf()
fig.sca(axBottom)
bovy_plot.bovy_plot(apar,100.*(mO_approx_higherorder-mO_approx)/mO_approx_higherorder,
lw=2.5,gcf=True,color='k',
xrange=[0.,1.],zorder=1,
yrange=[-0.039,0.039],
xlabel=r'$\Delta \theta_\parallel$',
ylabel=r'$\mathrm{relative\ difference\ in\ \%}$')
if num:
plot(apar,100.*(mO_num-mO_approx_higherorder)/mO_approx_higherorder,
lw=2.5,color='0.55',zorder=0)
# label
aparIndx= numpy.argmin(numpy.fabs(apar-0.64))
plot([0.45,apar[aparIndx]],[0.024,(100.*(mO_approx_higherorder-mO_approx)/mO_approx_higherorder)[aparIndx]],
'k',lw=1.)
bovy_plot.bovy_text(0.1,0.026,r'$\mathrm{higher\!\!-\!\!order\ minus\ linear}$',size=17.)
aparIndx= numpy.argmin(numpy.fabs(apar-0.6))
if num:
plot([0.45,apar[aparIndx]],[-0.02,(100.*(mO_num-mO_approx_higherorder)/mO_approx_higherorder)[aparIndx]],
'k',lw=1.)
bovy_plot.bovy_text(0.05,-0.03,r'$\mathrm{numerical\ minus\ higher\!\!-\!\!order}$',size=17.)
if save_figures:
bovy_plot.bovy_end_print(os.path.join(os.getenv('PAPERSDIR'),'2016-stream-stats','gd1likeSingle_mOparapprox.pdf'))
start= time.time()
numpy.array([sdf_gap.density_par(x,approx=False) for x in apar[::10]])
end= time.time()
print (end-start)*1000.*10./len(apar)
start= time.time()
numpy.array([sdf_gap.density_par(x,approx=True) for x in apar[::10]])
end= time.time()
print (end-start)*1000.*10./len(apar)
start= time.time()
numpy.array([sdf_gap.density_par(x,approx=True,higherorder=True) for x in apar[::10]])
end= time.time()
print (end-start)*1000.*10./len(apar)
start= time.time()
numpy.array([sdf_gap.meanOmega(x,approx=False,oned=True) for x in apar[::10]])
end= time.time()
print (end-start)*1000.*10./len(apar)
start= time.time()
numpy.array([sdf_gap.meanOmega(x,approx=True,oned=True) for x in apar[::10]])
end= time.time()
print (end-start)*1000.*10./len(apar)
start= time.time()
numpy.array([sdf_gap.meanOmega(x,approx=True,oned=True,higherorder=True) for x in apar[::10]])
end= time.time()
print (end-start)*1000.*10./len(apar) | 186.439969323
0.519882548939
5.57736194495
| BSD-3-Clause | meanOperpAndApproxImpacts.ipynb | jobovy/streamgap-pepper |
Test the multiple-impact approximations | # Setup a four, intermediate impacts
m= [10.**7.,10.**7.25,10.**6.75,10.**7.5]
GM= [mm/bovy_conversion.mass_in_msol(V0,R0) for mm in m]
timpactIndx= [numpy.argmin(numpy.fabs(numpy.array(sdf_pepper._uniq_timpact)-1.3/bovy_conversion.time_in_Gyr(V0,R0))),
numpy.argmin(numpy.fabs(numpy.array(sdf_pepper._uniq_timpact)-2.3/bovy_conversion.time_in_Gyr(V0,R0))),
numpy.argmin(numpy.fabs(numpy.array(sdf_pepper._uniq_timpact)-3.3/bovy_conversion.time_in_Gyr(V0,R0))),
numpy.argmin(numpy.fabs(numpy.array(sdf_pepper._uniq_timpact)-4.3/bovy_conversion.time_in_Gyr(V0,R0)))]
sdf_pepper.set_impacts(impactb=[0.5*rs(m[0]),2.*rs(m[1]),1.*rs(m[2]),2.5*rs(m[3])],
subhalovel=numpy.array([[-25.,155.,30.],
[125.,35.,80.],
[-225.,5.,-40.],
[25.,-155.,37.]])/V0,
impact_angle=[0.6,0.4,0.3,0.3],
timpact=[sdf_pepper._uniq_timpact[ti] for ti in timpactIndx],
GM=GM,rs=[rs(mm) for mm in m])
sdf_gap= sdf_pepper
n= 100000
aa_mock_per= sdf_pepper.sample(n=n,returnaAdt=True)
dO= numpy.dot(aa_mock_per[0].T-sdf_gap._progenitor_Omega,
sdf_gap._sigomatrixEig[1][:,sdf_gap._sigomatrixEigsortIndx])
dO[:,2]*= sdf_gap._sigMeanSign
da= numpy.dot(aa_mock_per[1].T-sdf_gap._progenitor_angle,
sdf_gap._sigomatrixEig[1][:,sdf_gap._sigomatrixEigsortIndx])
da[:,2]*= sdf_gap._sigMeanSign
num= True
apar= numpy.arange(0.,sdf_smooth.length()+0.003,0.003)
dens_unp= numpy.array([sdf_smooth._density_par(x) for x in apar])
dens_approx= numpy.array([sdf_gap.density_par(x,approx=True) for x in apar])
# normalize
dens_unp= dens_unp/numpy.sum(dens_unp)/(apar[1]-apar[0])
dens_approx= dens_approx/numpy.sum(dens_approx)/(apar[1]-apar[0])
if num:
dens_num= numpy.array([sdf_gap.density_par(x,approx=False) for x in apar])
dens_num= dens_num/numpy.sum(dens_num)/(apar[1]-apar[0])
bovy_plot.bovy_print(axes_labelsize=18.,xtick_labelsize=14.,ytick_labelsize=14.)
figsize(6,7)
axTop= pyplot.axes([0.15,0.3,0.825,0.65])
fig= pyplot.gcf()
fig.sca(axTop)
bovy_plot.bovy_plot(apar,dens_approx,lw=2.5,gcf=True,
color='k',
xrange=[0.,1.],
yrange=[0.,2.24],
ylabel=r'$\mathrm{density}$')
plot(apar,dens_unp,lw=3.5,color='k',ls='--',zorder=0)
nullfmt = NullFormatter() # no labels
axTop.xaxis.set_major_formatter(nullfmt)
dum= hist(da[:,2],bins=101,normed=True,range=[apar[0],apar[-1]],
histtype='step',color='0.55',zorder=0,lw=3.)
axBottom= pyplot.axes([0.15,0.1,0.825,0.2])
fig= pyplot.gcf()
fig.sca(axBottom)
if num:
bovy_plot.bovy_plot(apar,100.*(dens_num-dens_approx)/dens_approx,
lw=2.5,gcf=True,color='k',
xrange=[0.,1.],
yrange=[-1.45,1.45],
zorder=2,
xlabel=r'$\Delta \theta_\parallel$',
ylabel=r'$\mathrm{relative\ difference\ in}\ \%$')
# label
if num:
aparIndx= numpy.argmin(numpy.fabs(apar-0.6))
plot([0.45,apar[aparIndx]],[0.7,(100.*(dens_num-dens_approx)/dens_approx)[aparIndx]],
'k',lw=1.)
bovy_plot.bovy_text(0.15,0.4,r'$\mathrm{numerical\ minus}$'+'\n'+r'$\mathrm{approximation}$',size=17.)
if save_figures:
bovy_plot.bovy_end_print(os.path.join(os.getenv('PAPERSDIR'),'2016-stream-stats','gd1likeMulti_densapprox.pdf'))
mO_unp= numpy.array([sdf_smooth.meanOmega(x,oned=True) for x in apar])\
*bovy_conversion.freq_in_Gyr(V0,R0)
mO_approx= numpy.array([sdf_gap.meanOmega(x,approx=True,oned=True) for x in apar])\
*bovy_conversion.freq_in_Gyr(V0,R0)
if num:
mO_num= numpy.array([sdf_gap.meanOmega(x,approx=False,oned=True) for x in apar])\
*bovy_conversion.freq_in_Gyr(V0,R0)
frac= 0.005
alpha=0.01
linecolor='0.65'
bovy_plot.bovy_print(axes_labelsize=18.,xtick_labelsize=14.,ytick_labelsize=14.)
figsize(6,7)
axTop= pyplot.axes([0.15,0.3,0.825,0.65])
fig= pyplot.gcf()
fig.sca(axTop)
bovy_plot.bovy_plot(apar,mO_approx,lw=2.5,gcf=True,
color='k',
xrange=[0.,1.],
yrange=[0.,0.2],
ylabel=r'$\Delta \Omega_\parallel\,(\mathrm{Gyr}^{-1})$')
plot(apar,mO_unp,lw=2.5,color='k',ls='--')
plot(da[::3,2],dO[::3,2]*bovy_conversion.freq_in_Gyr(V0,R0),
'k.',alpha=alpha*2,rasterized=True)
nullfmt = NullFormatter() # no labels
axTop.xaxis.set_major_formatter(nullfmt)
axBottom= pyplot.axes([0.15,0.1,0.825,0.2])
fig= pyplot.gcf()
fig.sca(axBottom)
if num:
bovy_plot.bovy_plot(apar,100.*(mO_num-mO_approx)/mO_approx,
lw=2.5,gcf=True,color='k',
xrange=[0.,1.],zorder=1,
yrange=[-0.39,0.39],
xlabel=r'$\Delta \theta_\parallel$',
ylabel=r'$\mathrm{relative\ difference\ in\ \%}$')
# label
if num:
aparIndx= numpy.argmin(numpy.fabs(apar-0.6))
plot([0.35,apar[aparIndx]],[0.2,(100.*(mO_num-mO_approx)/mO_approx)[aparIndx]],
'k',lw=1.)
bovy_plot.bovy_text(0.05,0.1,r'$\mathrm{numerical\ minus}$'+'\n'+r'$\mathrm{approximation}$',size=17.)
if save_figures:
bovy_plot.bovy_end_print(os.path.join(os.getenv('PAPERSDIR'),'2016-stream-stats','gd1likeMulti_mOparapprox.pdf'))
start= time.time()
numpy.array([sdf_gap.density_par(x,approx=False) for x in apar[::10]])
end= time.time()
print (end-start)*1000.*10./len(apar)
start= time.time()
numpy.array([sdf_gap.density_par(x,approx=True) for x in apar[::10]])
end= time.time()
print (end-start)*1000.*10./len(apar)
start= time.time()
numpy.array([sdf_gap.meanOmega(x,approx=False,oned=True) for x in apar[::10]])
end= time.time()
print (end-start)*1000.*10./len(apar)
start= time.time()
numpy.array([sdf_gap.meanOmega(x,approx=True,oned=True) for x in apar[::10]])
end= time.time()
print (end-start)*1000.*10./len(apar) | 2007.35482303
8.63206025326
| BSD-3-Clause | meanOperpAndApproxImpacts.ipynb | jobovy/streamgap-pepper |
Computational speed | nimp= 2**numpy.arange(1,9)
ntrials= 3
nsample= [10,10,10,10,10,10,33,33,33]
compt= numpy.zeros(len(nimp))
for ii,ni in enumerate(nimp):
tcompt= 0.
for t in range(ntrials):
nimpact=ni
timpacts= numpy.random.permutation(numpy.array(sdf_pepper._uniq_timpact))[:ni]
print len(timpacts)
impact_angles= numpy.array([\
sdf_pepper._icdf_stream_len[ti](numpy.random.uniform())
for ti in timpacts])
GMs= numpy.array([sample_GM() for a in impact_angles])
rss= numpy.array([sample_rs(gm) for gm in GMs])
impactbs= numpy.random.uniform(size=len(impact_angles))*Xrs*rss
subhalovels= numpy.empty((len(impact_angles),3))
for jj in range(len(timpacts)):
subhalovels[jj]=\
sdf_pepper._draw_impact_velocities(timpacts[jj],120./V0,
impact_angles[jj],n=1)[0]
# Flip angle sign if necessary
if not sdf_pepper._gap_leading: impact_angles*= -1.
# Setup
sdf_pepper.set_impacts(impact_angle=impact_angles,
impactb=impactbs,
subhalovel=subhalovels,
timpact=timpacts,
GM=GMs,rs=rss)
start= time.time()
numpy.array([sdf_pepper.density_par(x,approx=True) for x in apar[::nsample[ii]]])
end= time.time()
tcompt+= (end-start)*1000.*nsample[ii]/len(apar)
compt[ii]= tcompt/ntrials
bovy_plot.bovy_print(axes_labelsize=18.,xtick_labelsize=14.,ytick_labelsize=14.)
figsize(6,4)
bovy_plot.bovy_plot(numpy.log2(nimp),compt,'ko',
semilogy=True,
xrange=[0.,9.],
yrange=[.5,100000.],
ylabel=r'$\mathrm{time}\,(\mathrm{ms})$',
xlabel=r'$\mathrm{number\ of\ impacts}$')
p= numpy.polyfit(numpy.log10(nimp),numpy.log10(compt),deg=1)
bovy_plot.bovy_plot(numpy.log2(nimp),10.**(p[0]*numpy.log10(nimp)+p[1]),
'-',lw=2.,
color=(0.0, 0.4470588235294118, 0.6980392156862745),
overplot=True)
pyplot.text(0.3,0.075,
r'$\log_{10}\ \mathrm{time/ms} = %.2f \,\log_{10} N %.2f$' % (p[0],p[1]),
transform=pyplot.gca().transAxes,size=14.)
# Use 100, 1000 instead of 10^2, 10^3
gca().yaxis.set_major_formatter(ScalarFormatter())
def twoto(x,pos):
return r'$%i$' % (2**x)
formatter = FuncFormatter(twoto)
gca().xaxis.set_major_formatter(formatter)
gcf().subplots_adjust(left=0.175,bottom=0.15,right=0.95,top=0.95)
if save_figures:
bovy_plot.bovy_end_print(os.path.join(os.getenv('PAPERSDIR'),'2016-stream-stats','gd1likeMulti_compTime.pdf')) | _____no_output_____ | BSD-3-Clause | meanOperpAndApproxImpacts.ipynb | jobovy/streamgap-pepper |
Example densities and tracks Single masses | # Load our fiducial simulation's output, for apars and smooth stream
data= numpy.genfromtxt(os.path.join(os.getenv('DATADIR'),'streamgap-pepper','gd1_multtime',
'gd1_t64sampling_X5_5-9_dens.dat'),
delimiter=',',max_rows=2)
apars= data[0]
dens_unp= data[1]
data= numpy.genfromtxt(os.path.join(os.getenv('DATADIR'),'streamgap-pepper','gd1_multtime',
'gd1_t64sampling_X5_5-9_omega.dat'),
delimiter=',',max_rows=2)
omega_unp= data[1]
dens_example= []
omega_example= []
# Perform some simulations, for different mass ranges
numpy.random.seed(3)
nexample= 4
masses= [5.5,6.5,7.5,8.5]
for ii in range(nexample):
# Sampling functions
sample_GM= lambda: 10.**(masses[ii]-10.)\
/bovy_conversion.mass_in_1010msol(V0,R0)
rate= dNencdm(sdf_pepper,10.**masses[ii],Xrs=Xrs,
plummer=plummer)
sdf_pepper.simulate(rate=rate,sample_GM=sample_GM,sample_rs=sample_rs,Xrs=Xrs)
densOmega= numpy.array([sdf_pepper._densityAndOmega_par_approx(a) for a in apars]).T
dens_example.append(densOmega[0])
omega_example.append(densOmega[1])
bovy_plot.bovy_print(axes_labelsize=18.,xtick_labelsize=14.,ytick_labelsize=18.)
figsize(6,7)
overplot= False
for ii in range(nexample):
bovy_plot.bovy_plot(apars,dens_example[ii]/dens_unp+2.*ii+0.5*(ii>2),lw=2.5,
color='k',
xrange=[0.,1.3],
yrange=[0.,2.*nexample+1],
xlabel=r'$\Delta \theta_\parallel$',
ylabel=r'$\mathrm{density}/\mathrm{smooth\ density}+\mathrm{constant}$',
overplot=overplot)
plot(apars,apars*0.+1.+2.*ii+0.5*(ii>2),lw=1.5,color='k',ls='--',zorder=0)
bovy_plot.bovy_text(1.025,1.+2.*ii+0.5*(ii>2),r'$10^{%.1f}\,M_\odot$' % masses[ii],verticalalignment='center',size=18.)
overplot=True
if save_figures:
bovy_plot.bovy_end_print(os.path.join(os.getenv('PAPERSDIR'),'2016-stream-stats',
'gd1like_densexample_singlemasses.pdf'))
bovy_plot.bovy_print(axes_labelsize=18.,xtick_labelsize=14.,ytick_labelsize=18.)
figsize(6,7)
overplot= False
mult= [3.,3.,1.,1.]
for ii in range(nexample):
bovy_plot.bovy_plot(apars,mult[ii]*(omega_example[ii]/omega_unp-1.)+1.+2.*ii+0.5*(ii>2),
lw=2.5,
color='k',
xrange=[0.,1.3],
yrange=[0.,2.*nexample+1.],
xlabel=r'$\Delta \theta_\parallel$',
ylabel=r'$\langle\Delta \Omega_\parallel\rangle\big/\langle\Delta \Omega_\parallel^0\rangle+\mathrm{constant}$',
overplot=overplot)
plot(apars,apars*0.+1.+2.*ii+0.5*(ii>2),lw=1.5,color='k',ls='--',zorder=0)
bovy_plot.bovy_text(1.025,1.+2.*ii+0.5*(ii>2),r'$10^{%.1f}\,M_\odot$' % masses[ii],verticalalignment='center',size=18.)
bovy_plot.bovy_text(0.025,1.+2.*ii+0.1+0.5*(ii>2),r'$\times%i$' % mult[ii],size=18.)
overplot= True
if save_figures:
bovy_plot.bovy_end_print(os.path.join(os.getenv('PAPERSDIR'),'2016-stream-stats',
'gd1like_omegaexample_singlemasses.pdf')) | _____no_output_____ | BSD-3-Clause | meanOperpAndApproxImpacts.ipynb | jobovy/streamgap-pepper |
Full mass range First look at low apar resolution: | apars= apars[::30]
dens_unp= dens_unp[::30]
omega_unp= omega_unp[::30]
# Sampling functions
massrange=[5.,9.]
plummer= False
Xrs= 5.
nsubhalo= simulate_streampepper.nsubhalo
rs= simulate_streampepper.rs
dNencdm= simulate_streampepper.dNencdm
sample_GM= lambda: (10.**((-0.5)*massrange[0])\
+(10.**((-0.5)*massrange[1])\
-10.**((-0.5)*massrange[0]))\
*numpy.random.uniform())**(1./(-0.5))\
/bovy_conversion.mass_in_msol(V0,R0)
rate_range= numpy.arange(massrange[0]+0.5,massrange[1]+0.5,1)
rate= numpy.sum([dNencdm(sdf_pepper,10.**r,Xrs=Xrs,
plummer=plummer)
for r in rate_range])
sample_rs= lambda x: rs(x*bovy_conversion.mass_in_1010msol(V0,R0)*10.**10.,
plummer=plummer)
dens_example2= []
omega_example2= []
# Perform some simulations
numpy.random.seed(3)
nexample= 4
for ii in range(nexample):
sdf_pepper.simulate(rate=rate,sample_GM=sample_GM,sample_rs=sample_rs,Xrs=Xrs)
densOmega= numpy.array([sdf_pepper._densityAndOmega_par_approx(a) for a in apars]).T
dens_example2.append(densOmega[0])
omega_example2.append(densOmega[1])
bovy_plot.bovy_print(axes_labelsize=18.,xtick_labelsize=14.,ytick_labelsize=18.)
figsize(6,7)
overplot= False
for ii in range(nexample):
bovy_plot.bovy_plot(apars,dens_example2[ii]/dens_unp+2.*ii,lw=2.5,
color='k',
xrange=[0.,1.],
yrange=[0.,2.*nexample+1.],
xlabel=r'$\Delta \theta_\parallel$',
ylabel=r'$\mathrm{density}/\mathrm{smooth\ density}+\mathrm{constant}$',
overplot=overplot)
plot(apars,apars*0.+1.+2.*ii,lw=1.5,color='k',ls='--',zorder=0)
overplot=True
bovy_plot.bovy_print(axes_labelsize=18.,xtick_labelsize=14.,ytick_labelsize=18.)
figsize(6,7)
overplot= False
for ii in range(nexample):
bovy_plot.bovy_plot(apars,omega_example2[ii]/omega_unp+2.*ii,lw=2.5,
color='k',
xrange=[0.,1.],
yrange=[0.,2.*nexample],
xlabel=r'$\Delta \theta_\parallel$',
ylabel=r'$\langle\Delta \Omega_\parallel\rangle\big/\langle\Delta \Omega_\parallel^0\rangle+\mathrm{constant}$',
overplot=overplot)
plot(apars,apars*0.+1.+2.*ii,lw=1.5,color='k',ls='--',zorder=0)
overplot= True | _____no_output_____ | BSD-3-Clause | meanOperpAndApproxImpacts.ipynb | jobovy/streamgap-pepper |
At full apar resolution: | # Load our fiducial simulation's output, for apars and smooth stream
data= numpy.genfromtxt(os.path.join(os.getenv('DATADIR'),'streamgap-pepper','gd1_multtime',
'gd1_t64sampling_X5_5-9_dens.dat'),
delimiter=',',max_rows=2)
apars= data[0]
dens_unp= data[1]
data= numpy.genfromtxt(os.path.join(os.getenv('DATADIR'),'streamgap-pepper','gd1_multtime',
'gd1_t64sampling_X5_5-9_omega.dat'),
delimiter=',',max_rows=2)
omega_unp= data[1]
dens_example2= []
omega_example2= []
# Perform some simulations
numpy.random.seed(3)
nexample= 4
for ii in range(nexample):
sdf_pepper.simulate(rate=rate,sample_GM=sample_GM,sample_rs=sample_rs,Xrs=Xrs)
densOmega= numpy.array([sdf_pepper._densityAndOmega_par_approx(a) for a in apars]).T
dens_example2.append(densOmega[0])
omega_example2.append(densOmega[1])
bovy_plot.bovy_print(axes_labelsize=18.,xtick_labelsize=14.,ytick_labelsize=18.)
figsize(6,7)
overplot= False
for ii in range(nexample):
bovy_plot.bovy_plot(apars,dens_example2[ii]/dens_unp+2.*ii,lw=2.5,
color='k',
xrange=[0.,1.],
yrange=[0.,2.*nexample+1.],
xlabel=r'$\Delta \theta_\parallel$',
ylabel=r'$\mathrm{density}/\mathrm{smooth\ density}+\mathrm{constant}$',
overplot=overplot)
plot(apars,apars*0.+1.+2.*ii,lw=1.5,color='k',ls='--',zorder=0)
overplot=True
if save_figures:
bovy_plot.bovy_end_print(os.path.join(os.getenv('PAPERSDIR'),'2016-stream-stats','gd1like_densexample.pdf'))
bovy_plot.bovy_print(axes_labelsize=18.,xtick_labelsize=14.,ytick_labelsize=18.)
figsize(6,7)
overplot= False
for ii in range(nexample):
bovy_plot.bovy_plot(apars,omega_example2[ii]/omega_unp+2.*ii,lw=2.5,
color='k',
xrange=[0.,1.],
yrange=[0.,2.*nexample],
xlabel=r'$\Delta \theta_\parallel$',
ylabel=r'$\langle\Delta \Omega_\parallel\rangle\big/\langle\Delta \Omega_\parallel^0\rangle+\mathrm{constant}$',
overplot=overplot)
plot(apars,apars*0.+1.+2.*ii,lw=1.5,color='k',ls='--',zorder=0)
overplot= True
if save_figures:
bovy_plot.bovy_end_print(os.path.join(os.getenv('PAPERSDIR'),'2016-stream-stats','gd1like_omegaexample.pdf')) | _____no_output_____ | BSD-3-Clause | meanOperpAndApproxImpacts.ipynb | jobovy/streamgap-pepper |
Examples Example 1: Move A FileAKA cut and paste. Moving files is a fast operation because we are just changing a pointer to the ones and zeros to point to something else. We are not actually pushing bits around disk to make the move happen.The module shutil is filled with all kinds of file handling goodies.In the root of the example directory is a small file. While it is in the root folder, it is simulating being outside the boundaries of the data warehouse environment. Let’s bring it inside by moving it to the In folder. | import shutil as sh
import os
if not 'script_dir' in globals():
script_dir = os.getcwd()
data_directory = 'data\\'
example_directory = 'BasicFileOpsExample\\'
target_directory = 'In\\'
file_name = 'forestfires.csv'
source_path = os.path.join(script_dir,data_directory,example_directory,file_name)
target_path = os.path.join(script_dir,data_directory,example_directory,target_directory,file_name)
sh.move(source_path, target_path) | _____no_output_____ | MIT | 03 Advanced/20-basic-file-io.ipynb | MassStreetUniversity/tutorial-python |
Example 2: Archiving A FileWe are now done processing the file and we need to archive it in case we need to drag it out and reload the system. The process of archiving is multi step. Zip up the file. Move the file to the Archive folder. Blow away the original.Once you run the example, check the Archive folder and the In folder. You should see a zip file in Archive and nothing in the In folder. | import zipfile as zf
import os
if not 'script_dir' in globals():
script_dir = os.getcwd()
data_directory = 'data\\'
example_directory = 'BasicFileOpsExample\\'
source_directory = 'In\\'
target_directory = 'Archive\\'
file_name = 'forestfires.csv'
archive_name = 'forestfires.zip'
target_path = os.path.join(script_dir,data_directory,example_directory,target_directory,archive_name)
source_path = os.path.join(data_directory,example_directory,source_directory)
archive = zf.ZipFile(target_path, "w")
os.chdir(source_path)
archive.write(file_name)
archive.close() | _____no_output_____ | MIT | 03 Advanced/20-basic-file-io.ipynb | MassStreetUniversity/tutorial-python |
Exercises Q 1The energy required to get from point $\vec{r}_1$ to point $\vec{r}_2$ for a plane is given by$$ E = \alpha \int_{C} \left| \frac{d\vec{r}}{dt} \right| dt - \int_C \vec{F} \cdot \frac{d\vec{r}}{dt}dt $$Suppose that $\alpha=5$ and our start and ends points are $\vec{r}_1 = (0,0)$ and $\vec{r}_2 = (0, 10)$. On this particular day, the wind produces a force field $\vec{F} = (0, -2/(x+1)^2)$. Find the optimal value of $A$ in $\vec{r}(t) = A\sin(\pi t/10)\hat{x} + t\hat{y}$ that minimizes the work. Then $x=A\sin(\pi t/10)$, $y=t$, and$$\left| \frac{d\vec{r}}{dt} \right| = \sqrt{1+(\pi A /10)^2 \cos^2(\pi t/10)}$$$$\vec{F} \cdot d\vec{r} = \begin{bmatrix} 0 \\ -2/(A\sin(\pi t/10) +1)^2\\ \end{bmatrix} \cdot \begin{bmatrix} \pi A/10 \cos(\pi t/10)\\ 1\\ \end{bmatrix} = -2/(A\sin(\pi t/10) +1)^2 $$so that$$ E = \int_{0}^{10} \left(5\sqrt{1+(\pi A /10)^2 \cos^2(\pi t/10)} + \frac{2}{(A\sin(\pi t/10) +100)^2} \right) dt$$ Q2Newton's law of cooling is$$\frac{dT}{dt} = -k(T-T_s(t)) $$where $T$ is the temperature of an object in the surroundings with temperature $T_s(t)$ (which may depend on time). Suppose $T$ represents the temperature of a shallow pool of water and $T_s(t)$ represents the temperature of outside. Find $T(t)$ given that you collected measurements of the outside: | t_m = np.array([ 0., 1.04347826, 2.08695652, 3.13043478, 4.17391304,
5.2173913 , 6.26086957, 7.30434783, 8.34782609, 9.39130435,
10.43478261, 11.47826087, 12.52173913, 13.56521739, 14.60869565,
15.65217391, 16.69565217, 17.73913043, 18.7826087 , 19.82608696,
20.86956522, 21.91304348, 22.95652174, 24. ])
temp_m = np.array([283.2322975, 284.6945461, 286.2259041, 287.8603625, 289.6440635,
291.6187583, 293.7939994, 296.1148895, 298.4395788, 300.5430675,
302.1566609, 303.0363609, 303.0363609, 302.1566609, 300.5430675,
298.4395788, 296.1148895, 293.7939994, 291.6187583, 289.6440635,
287.8603625, 286.2259041, 284.6945461, 283.2322975])
times = np.linspace(1, 23, 1000)
T0 = 284.6945461 | _____no_output_____ | Unlicense | day4/04. Scipy - Exercises.ipynb | ubutnux/bosscha-python-workshop-2022 |
Look At It | # Load image - individual 100307.
img = ci.load("fmri_data/rfMRI_REST1_LR_Atlas_hp2000_clean_filt_sm6.HCPMMP.ptseries.nii")
fmri_data = np.array(img.get_fdata())
# Visualize data, i.e. inspect the first 60 timesteps of each parcel.
# Generate heatmap.
timesteps = 60
displayed_data = np.transpose(fmri_data[range(timesteps),:])
plt.subplots(figsize=(15,10))
ax = sns.heatmap(displayed_data, yticklabels=False)
# Plot parameters.
plt.title('Resting fMRI Signal by Parcel - Individual 100307 LR', fontsize = 20)
plt.ylabel('Parcel', fontsize = 15)
plt.xlabel('Timestep', fontsize = 15)
plt.show() | _____no_output_____ | Apache-2.0 | docs/tutorials/time_series/fmri_analysis.ipynb | RebeccaYin7/hyppo |
Set Up Hyperparameters | # From Ting: Regions-of-Interest (ROIs)
roi_keys = np.array([1, 23, 18, 53, 24, 96, 117, 50, 143, 109, 148, 60, 38, 135, 93, 83, 149, 150, 65, 161, 132, 71]) - 1
roi_labels = np.array([
"Visual",
"Visual",
"Visual",
"SM",
"SM",
"dAtt",
"dAtt",
"dAtt",
"dAtt",
"vAtt",
"vAtt",
"vAtt",
"vAtt",
"Limbic",
"Limbic",
"FP",
"FP",
"DMN",
"DMN",
"DMN",
"DMN",
"DMN",
])
roi_data = fmri_data[0:300, roi_keys]
num_roi = len(roi_keys)
# Hyperparameters
max_lag = 1 # number of lags to check in the past
reps = 1000 # number of bootstrap replicates
workers = 1 # number of workers in internal MGC parallelization
# Subsample to test experiment.
# pairs = list(product(range(num_roi), repeat = 2)) # Fake param for testing.
pairs = list(product(range(num_roi), repeat = 2)) | _____no_output_____ | Apache-2.0 | docs/tutorials/time_series/fmri_analysis.ipynb | RebeccaYin7/hyppo |
Run Experiment | mgcx = MGCX(max_lag = max_lag)
def worker(i, j):
X = roi_data[:, i]
Y = roi_data[:, j]
stat, pval, mgcx_dict = mgcx.test(X, Y, reps = reps, workers = workers)
opt_lag = mgcx_dict['opt_lag']
opt_scale_x, opt_scale_y = mgcx_dict['opt_scale']
return stat, pval, opt_lag, opt_scale_x, opt_scale_y
output = np.array(Parallel(n_jobs=-2)(delayed(worker)(i, j) for i, j in pairs))
pickle.dump(output, open('fmri_data/mgcx_fmri_output.pkl', 'wb'))
# Load results into num_roi-by-num_roi matrices.
results = pickle.load(open('fmri_data/mgcx_fmri_output.pkl', 'rb'))
test_outputs = ['stat', 'pval', 'opt_lag', 'opt_scale_x', 'opt_scale_y']
matrices = np.zeros((len(test_outputs), num_roi, num_roi))
for p, pair in enumerate(pairs):
i, j = pair
for t in range(len(test_outputs)):
matrices[t, i, j] = results[p, t]
for t, test_output in enumerate(test_outputs):
pickle.dump(matrices[t], open('fmri_data/%s_matrix.pkl' % test_output, 'wb')) | _____no_output_____ | Apache-2.0 | docs/tutorials/time_series/fmri_analysis.ipynb | RebeccaYin7/hyppo |
Visualize Matrices | def plot_heatmap(matrix, labels, title, filename):
# sns.set()
cmap = mpl.cm.get_cmap('Purples')
cc = np.linspace(0, 1, 256)
cmap = mpl.colors.ListedColormap(cmap(cc))
heatmap_kws = dict(
cbar=False,
font_scale=1.4,
inner_hier_labels=labels,
hier_label_fontsize=20,
cmap=cmap,
center=None,
)
side_label_kws = dict(labelpad=45, fontsize=24)
fig, ax = plt.subplots(1, 1, figsize=(20, 16))
# Plot heatmap via graspy.
heatmap(matrix, ax=ax, **heatmap_kws)
ax.set_title(title, pad = 100, fontdict = {'fontsize' : 23})
# Create ticks.
num_ticks = 8
top_val = np.max(matrix)
ticks = [i * np.max(matrix) / num_ticks for i in range(num_ticks+1)]
yticks = [('%.2f' % np.round(10 ** -p, 2)) for p in ticks]
# Add colorbar.
sm = plt.cm.ScalarMappable(cmap=cmap)
sm.set_array(matrix)
cbar = fig.colorbar(sm, ax=ax, fraction=0.0475, pad=-0.1, ticks=ticks)
cbar.ax.set_yticklabels(yticks)
cbar.ax.tick_params(labelsize=25)
plt.savefig(
"%s.pdf" % filename,
facecolor="w",
format="pdf",
bbox_inches="tight",
)
plt.tight_layout()
plt.show() | _____no_output_____ | Apache-2.0 | docs/tutorials/time_series/fmri_analysis.ipynb | RebeccaYin7/hyppo |
p-value Matrix | # Apply negative log10 transform.
# matrix = pickle.load(open('fmri_data/pval_matrix.pkl', 'rb'))
# matrix = -np.log10(matrix)
# pickle.dump(matrix, open('fmri_data/nl10_pval_matrix.pkl', 'wb'))
matrix = pickle.load(open('fmri_data/nl10_pval_matrix.pkl', 'rb'))
plot_heatmap(matrix, roi_labels, 'p-Value', 'pval') | _____no_output_____ | Apache-2.0 | docs/tutorials/time_series/fmri_analysis.ipynb | RebeccaYin7/hyppo |
Buffered Text-to-SpeechIn this tutorial, we are going to build a state machine that controls a text-to-speech synthesis. The problem we solve is the following:- Speaking the text takes time, depending on how long the text is that the computer should speak.- Commands for speaking can arrive at any time, and we would like our state machine to process one of them at a time. So, even if we send three messages to it shortly after each other, it processes them one after the other.While solving this problem, we can learn more about the following concepts in STMPY state machines:- **Do-Activities**, which allow us to encapsulate the long-running text-to-speech function in a state machine.- **Deferred Events**, which allow us to ignore incoming messages until a later state, when we are ready again. Text-to-Speech MacOn a Mac, this is a function to make your computer speak: | from os import system
def text_to_speech(text):
system('say {}'.format(text)) | _____no_output_____ | MIT | notebooks/Buffered Text-to-Speech.ipynb | Hallvardd/ttm4115-project |
Run the above cell so the function is available in the following, and then execute the following cell to test it: | text_to_speech("Hello. I am a computer.") | _____no_output_____ | MIT | notebooks/Buffered Text-to-Speech.ipynb | Hallvardd/ttm4115-project |
WindowsTODO: We should have some code to run text to speech on Windows, too! State Machine 1With this function, we can create our first state machine that accepts a message and then speaks out some text. (Let's for now ignore how we get the text into the method, we will do that later.)Unfortunately, this state machine has a problem. This is because the method `text_to_speech(text)` is taking a long time to complete. This means, for the entire time that it takes to speak the text, nothing else can happen in all the state machines that are part of the same driver! State Machine 2 Long-Running ActionsThe way this function is implented makes that it **blocks**. This means, the Python program is busy executing this function as long as the speech takes to pronouce the message. Longer message, longer blocking.You can test this by putting some debugging aroud the function, to see when the functions returns: | print('Before speaking.')
text_to_speech("Hello. I am a computer.")
print('After speaking.') | _____no_output_____ | MIT | notebooks/Buffered Text-to-Speech.ipynb | Hallvardd/ttm4115-project |
You see that the string _"After speaking"_ is printed after the speaking is finished. During the execution, the program is blocked and does not do anything else. When our program should also do other stuff at the same time, either completely unrelated to speech or even just accepting new speech commands, this is not working! The driver is now completely blocked with executing the speech method, not being able to do anything else. Do-ActivitiesInstead of executing the method as part of a transition, we execute it as part of a state. This is called a **Do-Activity**, and it is declared as part of a state. The do-activity is started when the state is entered. Once the activity is finished, the state machine receives the event `done`, which triggers it to switch into another state.You may think now that the do-activity is similar to an entry action, as it is started when entering a state. However, a do-activity is started as part of its own thread, so that it does not block any other behavior from happening. Our state machine stays responsive, and so does any of the other state machines that may be assigned to the same driver. This happens in the background, STMPY is creating a new thread for a do-activity, starts it, and dispatches the `done` event once the do-activity finishes.When the do-activity finishes (in the case of the text-to-speech function, this means when the computer is finished talking), the state machine dispatches _automatically_ the event `done`, which brings the state machine into the next state. - A state with a do activity can therefore only declare one single outgoing transition that is triggered by the event `done`. - A state can have at most one do-activity. - A do-activity cannot be aborted. Instead, it should be programmed so that the function itself terminates, indicated for instance by the change of a variable.The following things are still possible in a state with a do-activity:- A state with a do-activity can have entry and exit actions. They are simply executed before or after the do activities.- A state with a do-activity can have internal transitions, since they don't leave the state. | from stmpy import Machine, Driver
from os import system
import logging
debug_level = logging.DEBUG
logger = logging.getLogger('stmpy')
logger.setLevel(debug_level)
ch = logging.StreamHandler()
ch.setLevel(debug_level)
formatter = logging.Formatter('%(asctime)s - %(name)-12s - %(levelname)-8s - %(message)s')
ch.setFormatter(formatter)
logger.addHandler(ch)
class Speaker:
def speak(self, string):
system('say {}'.format(string))
speaker = Speaker()
t0 = {'source': 'initial', 'target': 'ready'}
t1 = {'trigger': 'speak', 'source': 'ready', 'target': 'speaking'}
t2 = {'trigger': 'done', 'source': 'speaking', 'target': 'ready'}
s1 = {'name': 'speaking', 'do': 'speak(*)'}
stm = Machine(name='stm', transitions=[t0, t1, t2], states=[s1], obj=speaker)
speaker.stm = stm
driver = Driver()
driver.add_machine(stm)
driver.start()
driver.send('speak', 'stm', args=['My first sentence.'])
driver.send('speak', 'stm', args=['My second sentence.'])
driver.send('speak', 'stm', args=['My third sentence.'])
driver.send('speak', 'stm', args=['My fourth sentence.'])
driver.wait_until_finished() | _____no_output_____ | MIT | notebooks/Buffered Text-to-Speech.ipynb | Hallvardd/ttm4115-project |
The state machine 2 still has a problem, but this time another one: If we receive a new message with more text to speak _while_ we are in state `speaking`, this message is discarded. Our next state machine will fix this. State Machine 3As you know, events arriving in a state that do not declare outgoing triggers with that event, are discarded (that means, thrown away). For our state machine 2 above this means that when we are in state `speaking` and a new message arrives, this message is discarded. However, what we ideally want is that this message is handled once the currently spoken text is finished. There are two ways of achieving this:1. We could build a queue variable into our logic, and declare a transition that puts any arriving `speak` message into that queue. Whenever the currently spoken text finishes, we take another one from the queue until the queue is empty again. This has the drawback that we need to code the queue ourselves.2. We use a mechanism called **deferred event**, which is part of the state machine mechanics. This is the one we are going to use below. Deferred EventsA state can declare that it wants to **defer** an event, which simply means to not handle it. For our speech state machine it means that state `speaking` can declare that it defers event `speak`. Any event that arrives in a state that defers it, is ignored by that state. It is as if it never arrived, or as if it is invisible in the incoming event queue. Only once we switch into a next state that does not defer it, it gets visible again, and then either consumed by a transition, or discarded if the state does not declare any transition triggered by it. | s1 = {'name': 'speaking', 'do': 'speak(*)', 'speak': 'defer'}
stm = Machine(name='stm', transitions=[t0, t1, t2], states=[s1], obj=speaker)
speaker.stm = stm
driver = Driver()
driver.add_machine(stm)
driver.start()
driver.send('speak', 'stm', args=['My first sentence.'])
driver.send('speak', 'stm', args=['My second sentence.'])
driver.send('speak', 'stm', args=['My third sentence.'])
driver.send('speak', 'stm', args=['My fourth sentence.'])
driver.wait_until_finished() | _____no_output_____ | MIT | notebooks/Buffered Text-to-Speech.ipynb | Hallvardd/ttm4115-project |
概要- 101クラス分類- 対象:料理画像- VGG16による転移学習 1. 全結合層 1. 全層- RXなし | RUN = 100 | _____no_output_____ | MIT | VGG16.ipynb | KoshinoK/RandomExchange |
使用するGPUメモリの制限 | import tensorflow as tf
tf_ver = tf.__version__
if tf_ver.startswith('1.'):
from tensorflow.keras.backend import set_session
config = tf.ConfigProto()
config.gpu_options.allow_growth = True
config.log_device_placement = True
sess = tf.Session(config=config)
set_session(sess) | _____no_output_____ | MIT | VGG16.ipynb | KoshinoK/RandomExchange |
使用するGPUを指定 | import os
os.environ["CUDA_VISIBLE_DEVICES"]="0" | _____no_output_____ | MIT | VGG16.ipynb | KoshinoK/RandomExchange |
matplotlibでプロットしたグラフをファイルへ保存 | import os
def save_fig(plt, file_prefix):
if file_prefix == '':
return
parent = os.path.dirname(os.path.abspath(file_prefix))
os.makedirs(parent, exist_ok=True)
plt.savefig(f'{file_prefix}.pdf', transparent=True, bbox_inches='tight', pad_inches = 0)
plt.savefig(f'{file_prefix}.png', transparent=True, dpi=300, bbox_inches='tight', pad_inches = 0) | _____no_output_____ | MIT | VGG16.ipynb | KoshinoK/RandomExchange |
指定フォルダ以下にある画像リストを作成- サブフォルダはラベルに対応する数字であること- TOP_DIR - 0 - 00001.jpg - 00002.jpg - 1 - 00003.jpg - 00004.jpg | import pathlib
import random
import os
TOP_DIR = '/data1/Datasets/Food-101/03_all'
sub_dirs = pathlib.Path(TOP_DIR).glob('*/**')
label2files = dict()
for s in sub_dirs:
files = pathlib.Path(s).glob('**/*.jpg')
label = int(os.path.basename(s))
label2files[label] = list(files) | _____no_output_____ | MIT | VGG16.ipynb | KoshinoK/RandomExchange |
画像とラベルを訓練データと検証データに分割する | ratio = 0.8
train_list = []
train_labels = []
val_list = []
val_labels = []
for k, v in label2files.items():
random.shuffle(v)
N = len(v)
N_train = int(N * ratio)
train_list.extend(v[:N_train])
train_labels.extend([k] * N_train)
val_list.extend(v[N_train:])
val_labels.extend([k] * (N - N_train))
NUM_CLASSES = len(label2files.keys()) | _____no_output_____ | MIT | VGG16.ipynb | KoshinoK/RandomExchange |
画像ファイルリストとラベルから教師データを生成するクラス | import math
import numpy as np
from PIL import Image
import matplotlib.pyplot as plt
%matplotlib inline
import keras
from tensorflow.keras.utils import to_categorical
from tensorflow.keras.preprocessing.image import img_to_array
from tensorflow.keras.applications.vgg16 import preprocess_input
import tensorflow as tf
class ImageSequence(tf.keras.utils.Sequence):
def __init__(self, file_list, labels, batch_size, image_shape=(224, 224), shuffle=True, horizontal_flip=True):
self.file_list = np.array(file_list)
self.labels = to_categorical(labels)
self.batch_size = batch_size
self.image_shape = image_shape
self.shuffle = shuffle
self.horizontal_flip = horizontal_flip
self.indexes = np.arange(len(self.file_list))
if self.shuffle:
random.shuffle(self.indexes)
def __getitem__(self, index):
idx = self.indexes[index * self.batch_size : (index + 1) * self.batch_size]
y = self.labels[idx]
files = self.file_list[idx]
x = []
for f in files:
try:
img = Image.open(f)
# 正しいデータはRGB画像
# データセットの中には、グレースケール画像が入っている可能性がある。
# RGBに変換して、正しいデータと次元を揃える
img = img.convert('RGB')
img = img.resize(self.image_shape, Image.BILINEAR)
img = img_to_array(img)
img = preprocess_input(img) / 255.0
if self.horizontal_flip and np.random.random() > 0.5:
img = img[:,::-1, :]
x.append(np.expand_dims(img, axis=0))
except:
print(f)
return np.concatenate(x, axis=0), y
def __len__(self):
return len(self.file_list) // self.batch_size
def on_epoch_end(self):
if self.shuffle:
random.shuffle(self.indexes) | Using TensorFlow backend.
| MIT | VGG16.ipynb | KoshinoK/RandomExchange |
モデル保存用のディレクトリを作成★ | import os
from datetime import datetime
# モデル保存用ディレクトリの準備
model_dir = os.path.join(
f'../run/VGG16_run{RUN}'
)
os.makedirs(model_dir, exist_ok=True)
print('model_dir:', model_dir) # 保存先のディレクトリ名を表示
dir_weights = model_dir
os.makedirs(dir_weights, exist_ok=True) | model_dir: ../run/VGG16_run100
| MIT | VGG16.ipynb | KoshinoK/RandomExchange |
VGGモデルのロード | from tensorflow.keras.applications.vgg16 import VGG16
# 既存の1000クラスの出力を使わないため、
# `incliude_top=False`として出力層を含まない状態でロード
vgg16 = VGG16(include_top=False, input_shape=(224, 224, 3))
# モデルのサマリを確認。出力層が含まれてないことがわかる
vgg16.summary() | _________________________________________________________________
Layer (type) Output Shape Param #
=================================================================
input_1 (InputLayer) (None, 224, 224, 3) 0
_________________________________________________________________
block1_conv1 (Conv2D) (None, 224, 224, 64) 1792
_________________________________________________________________
block1_conv2 (Conv2D) (None, 224, 224, 64) 36928
_________________________________________________________________
block1_pool (MaxPooling2D) (None, 112, 112, 64) 0
_________________________________________________________________
block2_conv1 (Conv2D) (None, 112, 112, 128) 73856
_________________________________________________________________
block2_conv2 (Conv2D) (None, 112, 112, 128) 147584
_________________________________________________________________
block2_pool (MaxPooling2D) (None, 56, 56, 128) 0
_________________________________________________________________
block3_conv1 (Conv2D) (None, 56, 56, 256) 295168
_________________________________________________________________
block3_conv2 (Conv2D) (None, 56, 56, 256) 590080
_________________________________________________________________
block3_conv3 (Conv2D) (None, 56, 56, 256) 590080
_________________________________________________________________
block3_pool (MaxPooling2D) (None, 28, 28, 256) 0
_________________________________________________________________
block4_conv1 (Conv2D) (None, 28, 28, 512) 1180160
_________________________________________________________________
block4_conv2 (Conv2D) (None, 28, 28, 512) 2359808
_________________________________________________________________
block4_conv3 (Conv2D) (None, 28, 28, 512) 2359808
_________________________________________________________________
block4_pool (MaxPooling2D) (None, 14, 14, 512) 0
_________________________________________________________________
block5_conv1 (Conv2D) (None, 14, 14, 512) 2359808
_________________________________________________________________
block5_conv2 (Conv2D) (None, 14, 14, 512) 2359808
_________________________________________________________________
block5_conv3 (Conv2D) (None, 14, 14, 512) 2359808
_________________________________________________________________
block5_pool (MaxPooling2D) (None, 7, 7, 512) 0
=================================================================
Total params: 14,714,688
Trainable params: 14,714,688
Non-trainable params: 0
_________________________________________________________________
| MIT | VGG16.ipynb | KoshinoK/RandomExchange |
VGG16を利用したモデルの作成と学習方法の設定★ | from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Dropout, Flatten
# モデルを編集し、ネットワークを生成する関数の定義
def build_transfer_model(vgg16):
# 読み出したモデルを使って、新しいモデルを作成
model = Sequential(vgg16.layers)
# 読み出した重みの一部は再学習しないように設定。
# ここでは、追加する層と出力層に近い層の重みのみを再学習
for layer in model.layers[:15]:
layer.trainable = False
# 追加する出力部分の層を構築
model.add(Flatten())
model.add(Dense(1024, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(1024, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(NUM_CLASSES, activation='softmax'))
return model | _____no_output_____ | MIT | VGG16.ipynb | KoshinoK/RandomExchange |
全結合層とそれに近い畳み込み層の学習★ モデル作成 | # 定義した関数を呼び出して、ネットワークを生成
model = build_transfer_model(vgg16) | _____no_output_____ | MIT | VGG16.ipynb | KoshinoK/RandomExchange |
ネットワーク構造の保存★ | import json
import pickle
# ネットワークの保存
model_json = os.path.join(model_dir, 'model.json')
with open(model_json, 'w') as f:
json.dump(model.to_json(), f) | _____no_output_____ | MIT | VGG16.ipynb | KoshinoK/RandomExchange |
最適化アルゴリズムなどを指定してモデルをコンパイルする | from tensorflow.keras.optimizers import SGD
model.compile(
loss='categorical_crossentropy',
optimizer=SGD(lr=1e-4, momentum=0.9),
metrics=['accuracy']
)
# モデルのサマリを確認
model.summary() | _________________________________________________________________
Layer (type) Output Shape Param #
=================================================================
block1_conv1 (Conv2D) (None, 224, 224, 64) 1792
_________________________________________________________________
block1_conv2 (Conv2D) (None, 224, 224, 64) 36928
_________________________________________________________________
block1_pool (MaxPooling2D) (None, 112, 112, 64) 0
_________________________________________________________________
block2_conv1 (Conv2D) (None, 112, 112, 128) 73856
_________________________________________________________________
block2_conv2 (Conv2D) (None, 112, 112, 128) 147584
_________________________________________________________________
block2_pool (MaxPooling2D) (None, 56, 56, 128) 0
_________________________________________________________________
block3_conv1 (Conv2D) (None, 56, 56, 256) 295168
_________________________________________________________________
block3_conv2 (Conv2D) (None, 56, 56, 256) 590080
_________________________________________________________________
block3_conv3 (Conv2D) (None, 56, 56, 256) 590080
_________________________________________________________________
block3_pool (MaxPooling2D) (None, 28, 28, 256) 0
_________________________________________________________________
block4_conv1 (Conv2D) (None, 28, 28, 512) 1180160
_________________________________________________________________
block4_conv2 (Conv2D) (None, 28, 28, 512) 2359808
_________________________________________________________________
block4_conv3 (Conv2D) (None, 28, 28, 512) 2359808
_________________________________________________________________
block4_pool (MaxPooling2D) (None, 14, 14, 512) 0
_________________________________________________________________
block5_conv1 (Conv2D) (None, 14, 14, 512) 2359808
_________________________________________________________________
block5_conv2 (Conv2D) (None, 14, 14, 512) 2359808
_________________________________________________________________
block5_conv3 (Conv2D) (None, 14, 14, 512) 2359808
_________________________________________________________________
block5_pool (MaxPooling2D) (None, 7, 7, 512) 0
_________________________________________________________________
flatten (Flatten) (None, 25088) 0
_________________________________________________________________
dense (Dense) (None, 1024) 25691136
_________________________________________________________________
dropout (Dropout) (None, 1024) 0
_________________________________________________________________
dense_1 (Dense) (None, 1024) 1049600
_________________________________________________________________
dropout_1 (Dropout) (None, 1024) 0
_________________________________________________________________
dense_2 (Dense) (None, 101) 103525
=================================================================
Total params: 41,558,949
Trainable params: 31,563,877
Non-trainable params: 9,995,072
_________________________________________________________________
| MIT | VGG16.ipynb | KoshinoK/RandomExchange |
シーケンス生成 | batch_size = 25
img_seq_train = ImageSequence(train_list, train_labels, batch_size=batch_size)
img_seq_validation = ImageSequence(val_list, val_labels, batch_size=batch_size)
print('Train images =', len(img_seq_train) * batch_size)
print('Validation images =', len(img_seq_validation) * batch_size) | Train images = 80800
Validation images = 20200
| MIT | VGG16.ipynb | KoshinoK/RandomExchange |
Callbackの生成★ | from tensorflow.keras.callbacks import ModelCheckpoint, CSVLogger, EarlyStopping, ReduceLROnPlateau
# Callbacksの設定
cp_filepath = os.path.join(dir_weights, 'ep_{epoch:04d}_ls_{loss:.1f}.h5')
cp = ModelCheckpoint(
cp_filepath,
monitor='val_acc',
verbose=0,
save_best_only=True,
save_weights_only=True,
mode='auto'
)
csv_filepath = os.path.join(model_dir, 'loss.csv')
csv = CSVLogger(csv_filepath, append=True)
es = EarlyStopping(monitor='val_acc', patience=20, verbose=1, mode='auto')
rl = ReduceLROnPlateau(monitor='val_loss', factor=0.5, patience=5, verbose=1, mode='auto', epsilon=0.0001, cooldown=0, min_lr=0) | WARNING:tensorflow:`epsilon` argument is deprecated and will be removed, use `min_delta` instead.
| MIT | VGG16.ipynb | KoshinoK/RandomExchange |
学習 | n_epoch = 200
# モデルの学習
history = model.fit_generator(
img_seq_train,
epochs=n_epoch, # 学習するエポック数
steps_per_epoch=len(img_seq_train),
validation_data=img_seq_validation,
validation_steps=len(img_seq_validation),
verbose=1,
callbacks=[cp, csv, es, rl]
) | Epoch 1/200
3232/3232 [==============================] - 1217s 377ms/step - loss: 4.6143 - acc: 0.0148 - val_loss: 4.5334 - val_acc: 0.0410
Epoch 2/200
3232/3232 [==============================] - 499s 155ms/step - loss: 4.4336 - acc: 0.0381 - val_loss: 4.0951 - val_acc: 0.1068
Epoch 3/200
3232/3232 [==============================] - 500s 155ms/step - loss: 4.0705 - acc: 0.0861 - val_loss: 3.5528 - val_acc: 0.2125
Epoch 4/200
3232/3232 [==============================] - 499s 154ms/step - loss: 3.6836 - acc: 0.1451 - val_loss: 3.1393 - val_acc: 0.2758
Epoch 5/200
3232/3232 [==============================] - 502s 155ms/step - loss: 3.3705 - acc: 0.2016 - val_loss: 2.8485 - val_acc: 0.3291
Epoch 6/200
3232/3232 [==============================] - 508s 157ms/step - loss: 3.1329 - acc: 0.2452 - val_loss: 2.6545 - val_acc: 0.3693
Epoch 7/200
3232/3232 [==============================] - 500s 155ms/step - loss: 2.9475 - acc: 0.2812 - val_loss: 2.5029 - val_acc: 0.3941
Epoch 8/200
3232/3232 [==============================] - 499s 154ms/step - loss: 2.7907 - acc: 0.3143 - val_loss: 2.3997 - val_acc: 0.4174
Epoch 9/200
3232/3232 [==============================] - 498s 154ms/step - loss: 2.6628 - acc: 0.3397 - val_loss: 2.2895 - val_acc: 0.4424
Epoch 10/200
3232/3232 [==============================] - 500s 155ms/step - loss: 2.5539 - acc: 0.3637 - val_loss: 2.2035 - val_acc: 0.4557
Epoch 11/200
3232/3232 [==============================] - 502s 155ms/step - loss: 2.4609 - acc: 0.3858 - val_loss: 2.1512 - val_acc: 0.4664
Epoch 12/200
3232/3232 [==============================] - 501s 155ms/step - loss: 2.3779 - acc: 0.4038 - val_loss: 2.0793 - val_acc: 0.4807
Epoch 13/200
3232/3232 [==============================] - 498s 154ms/step - loss: 2.3044 - acc: 0.4187 - val_loss: 2.0313 - val_acc: 0.4899
Epoch 14/200
3232/3232 [==============================] - 500s 155ms/step - loss: 2.2252 - acc: 0.4368 - val_loss: 2.0015 - val_acc: 0.4949
Epoch 15/200
3232/3232 [==============================] - 502s 155ms/step - loss: 2.1651 - acc: 0.4520 - val_loss: 1.9603 - val_acc: 0.5070
Epoch 16/200
3232/3232 [==============================] - 502s 155ms/step - loss: 2.1022 - acc: 0.4646 - val_loss: 1.9267 - val_acc: 0.5141
Epoch 17/200
3232/3232 [==============================] - 499s 154ms/step - loss: 2.0475 - acc: 0.4769 - val_loss: 1.8840 - val_acc: 0.5234
Epoch 18/200
3232/3232 [==============================] - 503s 156ms/step - loss: 1.9960 - acc: 0.4895 - val_loss: 1.8575 - val_acc: 0.5267
Epoch 19/200
3232/3232 [==============================] - 502s 155ms/step - loss: 1.9521 - acc: 0.4995 - val_loss: 1.8374 - val_acc: 0.5341
Epoch 20/200
3232/3232 [==============================] - 498s 154ms/step - loss: 1.9016 - acc: 0.5120 - val_loss: 1.8023 - val_acc: 0.5405
Epoch 21/200
3232/3232 [==============================] - 498s 154ms/step - loss: 1.8537 - acc: 0.5233 - val_loss: 1.7787 - val_acc: 0.5486
Epoch 22/200
3232/3232 [==============================] - 484s 150ms/step - loss: 1.8121 - acc: 0.5312 - val_loss: 1.7634 - val_acc: 0.5505
Epoch 23/200
3232/3232 [==============================] - 485s 150ms/step - loss: 1.7748 - acc: 0.5378 - val_loss: 1.7524 - val_acc: 0.5514
Epoch 24/200
3232/3232 [==============================] - 486s 150ms/step - loss: 1.7294 - acc: 0.5470 - val_loss: 1.7392 - val_acc: 0.5547
Epoch 25/200
3232/3232 [==============================] - 487s 151ms/step - loss: 1.6912 - acc: 0.5574 - val_loss: 1.7472 - val_acc: 0.5532
Epoch 26/200
3232/3232 [==============================] - 485s 150ms/step - loss: 1.6500 - acc: 0.5659 - val_loss: 1.7059 - val_acc: 0.5616
Epoch 27/200
3232/3232 [==============================] - 492s 152ms/step - loss: 1.6187 - acc: 0.5736 - val_loss: 1.6946 - val_acc: 0.5647
Epoch 28/200
3232/3232 [==============================] - 492s 152ms/step - loss: 1.5838 - acc: 0.5820 - val_loss: 1.6890 - val_acc: 0.5670
Epoch 29/200
3232/3232 [==============================] - 488s 151ms/step - loss: 1.5471 - acc: 0.5917 - val_loss: 1.6749 - val_acc: 0.5668
Epoch 30/200
3232/3232 [==============================] - 489s 151ms/step - loss: 1.5096 - acc: 0.5964 - val_loss: 1.6665 - val_acc: 0.5695
Epoch 31/200
3232/3232 [==============================] - 487s 151ms/step - loss: 1.4784 - acc: 0.6067 - val_loss: 1.6452 - val_acc: 0.5782
Epoch 32/200
3232/3232 [==============================] - 492s 152ms/step - loss: 1.4428 - acc: 0.6163 - val_loss: 1.6375 - val_acc: 0.5794
Epoch 33/200
3232/3232 [==============================] - 486s 150ms/step - loss: 1.4086 - acc: 0.6217 - val_loss: 1.6253 - val_acc: 0.5825
Epoch 34/200
3232/3232 [==============================] - 491s 152ms/step - loss: 1.3763 - acc: 0.6286 - val_loss: 1.6203 - val_acc: 0.5832
Epoch 35/200
3232/3232 [==============================] - 490s 151ms/step - loss: 1.3446 - acc: 0.6359 - val_loss: 1.6395 - val_acc: 0.5834
Epoch 36/200
3232/3232 [==============================] - 489s 151ms/step - loss: 1.3171 - acc: 0.6429 - val_loss: 1.6277 - val_acc: 0.5800
Epoch 37/200
3232/3232 [==============================] - 488s 151ms/step - loss: 1.2863 - acc: 0.6489 - val_loss: 1.6106 - val_acc: 0.5874
Epoch 38/200
3232/3232 [==============================] - 484s 150ms/step - loss: 1.2545 - acc: 0.6583 - val_loss: 1.6117 - val_acc: 0.5884
Epoch 39/200
3232/3232 [==============================] - 488s 151ms/step - loss: 1.2276 - acc: 0.6656 - val_loss: 1.6133 - val_acc: 0.5885
Epoch 40/200
3232/3232 [==============================] - 487s 151ms/step - loss: 1.1959 - acc: 0.6734 - val_loss: 1.6090 - val_acc: 0.5882
Epoch 41/200
3232/3232 [==============================] - 486s 150ms/step - loss: 1.1637 - acc: 0.6792 - val_loss: 1.6263 - val_acc: 0.5867
Epoch 42/200
3232/3232 [==============================] - 484s 150ms/step - loss: 1.1359 - acc: 0.6874 - val_loss: 1.6013 - val_acc: 0.5909
Epoch 43/200
3232/3232 [==============================] - 486s 151ms/step - loss: 1.1095 - acc: 0.6927 - val_loss: 1.6142 - val_acc: 0.5935
Epoch 44/200
3232/3232 [==============================] - 490s 152ms/step - loss: 1.0810 - acc: 0.7000 - val_loss: 1.6058 - val_acc: 0.5960
Epoch 45/200
3232/3232 [==============================] - 492s 152ms/step - loss: 1.0511 - acc: 0.7065 - val_loss: 1.6033 - val_acc: 0.5926
Epoch 46/200
3232/3232 [==============================] - 487s 151ms/step - loss: 1.0232 - acc: 0.7130 - val_loss: 1.6130 - val_acc: 0.5941
Epoch 47/200
3232/3232 [==============================] - 487s 151ms/step - loss: 0.9957 - acc: 0.7211 - val_loss: 1.5959 - val_acc: 0.5958
Epoch 48/200
3232/3232 [==============================] - 488s 151ms/step - loss: 0.9706 - acc: 0.7258 - val_loss: 1.6057 - val_acc: 0.5959
Epoch 49/200
3232/3232 [==============================] - 485s 150ms/step - loss: 0.9440 - acc: 0.7347 - val_loss: 1.6165 - val_acc: 0.5961
Epoch 50/200
3232/3232 [==============================] - 490s 152ms/step - loss: 0.9121 - acc: 0.7419 - val_loss: 1.6494 - val_acc: 0.5919
Epoch 51/200
3232/3232 [==============================] - 486s 151ms/step - loss: 0.8922 - acc: 0.7459 - val_loss: 1.6133 - val_acc: 0.6008
Epoch 52/200
3231/3232 [============================>.] - ETA: 0s - loss: 0.8630 - acc: 0.7528
Epoch 00052: ReduceLROnPlateau reducing learning rate to 4.999999873689376e-05.
3232/3232 [==============================] - 485s 150ms/step - loss: 0.8631 - acc: 0.7528 - val_loss: 1.6334 - val_acc: 0.5990
Epoch 53/200
3232/3232 [==============================] - 489s 151ms/step - loss: 0.7975 - acc: 0.7695 - val_loss: 1.6231 - val_acc: 0.6012
Epoch 54/200
3232/3232 [==============================] - 487s 151ms/step - loss: 0.7781 - acc: 0.7774 - val_loss: 1.6193 - val_acc: 0.6026
Epoch 55/200
3232/3232 [==============================] - 487s 151ms/step - loss: 0.7586 - acc: 0.7803 - val_loss: 1.6249 - val_acc: 0.6021
Epoch 56/200
3232/3232 [==============================] - 484s 150ms/step - loss: 0.7481 - acc: 0.7845 - val_loss: 1.6249 - val_acc: 0.6005
Epoch 57/200
3231/3232 [============================>.] - ETA: 0s - loss: 0.7362 - acc: 0.7866
Epoch 00057: ReduceLROnPlateau reducing learning rate to 2.499999936844688e-05.
3232/3232 [==============================] - 486s 150ms/step - loss: 0.7361 - acc: 0.7866 - val_loss: 1.6208 - val_acc: 0.6024
Epoch 58/200
3232/3232 [==============================] - 481s 149ms/step - loss: 0.6953 - acc: 0.8007 - val_loss: 1.6324 - val_acc: 0.6047
Epoch 59/200
3232/3232 [==============================] - 485s 150ms/step - loss: 0.6884 - acc: 0.8003 - val_loss: 1.6305 - val_acc: 0.6027
Epoch 60/200
3232/3232 [==============================] - 488s 151ms/step - loss: 0.6805 - acc: 0.8024 - val_loss: 1.6360 - val_acc: 0.6024
Epoch 61/200
3232/3232 [==============================] - 485s 150ms/step - loss: 0.6705 - acc: 0.8058 - val_loss: 1.6367 - val_acc: 0.6048
Epoch 62/200
3231/3232 [============================>.] - ETA: 0s - loss: 0.6591 - acc: 0.8081
Epoch 00062: ReduceLROnPlateau reducing learning rate to 1.249999968422344e-05.
3232/3232 [==============================] - 492s 152ms/step - loss: 0.6592 - acc: 0.8081 - val_loss: 1.6445 - val_acc: 0.6037
Epoch 63/200
3232/3232 [==============================] - 487s 151ms/step - loss: 0.6491 - acc: 0.8123 - val_loss: 1.6431 - val_acc: 0.6056
Epoch 64/200
3232/3232 [==============================] - 488s 151ms/step - loss: 0.6441 - acc: 0.8133 - val_loss: 1.6480 - val_acc: 0.6049
Epoch 65/200
3232/3232 [==============================] - 487s 151ms/step - loss: 0.6403 - acc: 0.8150 - val_loss: 1.6382 - val_acc: 0.6069
Epoch 66/200
3232/3232 [==============================] - 489s 151ms/step - loss: 0.6359 - acc: 0.8164 - val_loss: 1.6350 - val_acc: 0.6071
Epoch 67/200
3231/3232 [============================>.] - ETA: 0s - loss: 0.6319 - acc: 0.8168
Epoch 00067: ReduceLROnPlateau reducing learning rate to 6.24999984211172e-06.
3232/3232 [==============================] - 485s 150ms/step - loss: 0.6319 - acc: 0.8168 - val_loss: 1.6379 - val_acc: 0.6045
Epoch 68/200
3232/3232 [==============================] - 490s 151ms/step - loss: 0.6226 - acc: 0.8195 - val_loss: 1.6434 - val_acc: 0.6070
Epoch 69/200
3232/3232 [==============================] - 490s 152ms/step - loss: 0.6222 - acc: 0.8189 - val_loss: 1.6373 - val_acc: 0.6062
Epoch 70/200
3232/3232 [==============================] - 484s 150ms/step - loss: 0.6175 - acc: 0.8204 - val_loss: 1.6485 - val_acc: 0.6068
Epoch 71/200
3232/3232 [==============================] - 482s 149ms/step - loss: 0.6155 - acc: 0.8208 - val_loss: 1.6381 - val_acc: 0.6078
Epoch 72/200
3231/3232 [============================>.] - ETA: 0s - loss: 0.6161 - acc: 0.8215
Epoch 00072: ReduceLROnPlateau reducing learning rate to 3.12499992105586e-06.
3232/3232 [==============================] - 490s 152ms/step - loss: 0.6161 - acc: 0.8215 - val_loss: 1.6511 - val_acc: 0.6053
Epoch 73/200
3232/3232 [==============================] - 491s 152ms/step - loss: 0.6129 - acc: 0.8235 - val_loss: 1.6444 - val_acc: 0.6074
Epoch 74/200
3232/3232 [==============================] - 486s 150ms/step - loss: 0.6060 - acc: 0.8232 - val_loss: 1.6510 - val_acc: 0.6060
Epoch 75/200
3232/3232 [==============================] - 491s 152ms/step - loss: 0.6067 - acc: 0.8230 - val_loss: 1.6488 - val_acc: 0.6053
Epoch 76/200
3232/3232 [==============================] - 486s 150ms/step - loss: 0.6073 - acc: 0.8237 - val_loss: 1.6531 - val_acc: 0.6060
Epoch 77/200
3231/3232 [============================>.] - ETA: 0s - loss: 0.6054 - acc: 0.8248
Epoch 00077: ReduceLROnPlateau reducing learning rate to 1.56249996052793e-06.
3232/3232 [==============================] - 489s 151ms/step - loss: 0.6053 - acc: 0.8248 - val_loss: 1.6554 - val_acc: 0.6069
Epoch 78/200
3232/3232 [==============================] - 482s 149ms/step - loss: 0.6018 - acc: 0.8254 - val_loss: 1.6492 - val_acc: 0.6067
Epoch 79/200
3232/3232 [==============================] - 479s 148ms/step - loss: 0.6064 - acc: 0.8244 - val_loss: 1.6446 - val_acc: 0.6083
Epoch 80/200
3232/3232 [==============================] - 485s 150ms/step - loss: 0.6015 - acc: 0.8242 - val_loss: 1.6471 - val_acc: 0.6060
Epoch 81/200
3232/3232 [==============================] - 487s 151ms/step - loss: 0.6046 - acc: 0.8235 - val_loss: 1.6474 - val_acc: 0.6076
Epoch 82/200
3231/3232 [============================>.] - ETA: 0s - loss: 0.6039 - acc: 0.8248
Epoch 00082: ReduceLROnPlateau reducing learning rate to 7.81249980263965e-07.
3232/3232 [==============================] - 485s 150ms/step - loss: 0.6040 - acc: 0.8248 - val_loss: 1.6481 - val_acc: 0.6039
Epoch 83/200
3232/3232 [==============================] - 487s 151ms/step - loss: 0.6030 - acc: 0.8251 - val_loss: 1.6449 - val_acc: 0.6076
Epoch 84/200
3232/3232 [==============================] - 480s 149ms/step - loss: 0.6040 - acc: 0.8235 - val_loss: 1.6508 - val_acc: 0.6060
Epoch 85/200
3232/3232 [==============================] - 487s 151ms/step - loss: 0.6027 - acc: 0.8234 - val_loss: 1.6452 - val_acc: 0.6099
Epoch 86/200
3232/3232 [==============================] - 487s 151ms/step - loss: 0.6000 - acc: 0.8257 - val_loss: 1.6515 - val_acc: 0.6083
Epoch 87/200
3231/3232 [============================>.] - ETA: 0s - loss: 0.5978 - acc: 0.8245
Epoch 00087: ReduceLROnPlateau reducing learning rate to 3.906249901319825e-07.
3232/3232 [==============================] - 489s 151ms/step - loss: 0.5978 - acc: 0.8245 - val_loss: 1.6468 - val_acc: 0.6073
Epoch 88/200
3232/3232 [==============================] - 487s 151ms/step - loss: 0.6009 - acc: 0.8248 - val_loss: 1.6499 - val_acc: 0.6068
Epoch 89/200
3232/3232 [==============================] - 480s 149ms/step - loss: 0.6071 - acc: 0.8235 - val_loss: 1.6483 - val_acc: 0.6074
Epoch 90/200
3232/3232 [==============================] - 487s 151ms/step - loss: 0.6026 - acc: 0.8246 - val_loss: 1.6489 - val_acc: 0.6072
Epoch 91/200
3232/3232 [==============================] - 482s 149ms/step - loss: 0.5966 - acc: 0.8254 - val_loss: 1.6461 - val_acc: 0.6067
Epoch 92/200
3231/3232 [============================>.] - ETA: 0s - loss: 0.6014 - acc: 0.8238
Epoch 00092: ReduceLROnPlateau reducing learning rate to 1.9531249506599124e-07.
3232/3232 [==============================] - 482s 149ms/step - loss: 0.6014 - acc: 0.8238 - val_loss: 1.6436 - val_acc: 0.6103
Epoch 93/200
3232/3232 [==============================] - 483s 150ms/step - loss: 0.6033 - acc: 0.8245 - val_loss: 1.6448 - val_acc: 0.6070
Epoch 94/200
3232/3232 [==============================] - 484s 150ms/step - loss: 0.6004 - acc: 0.8246 - val_loss: 1.6473 - val_acc: 0.6079
Epoch 95/200
3232/3232 [==============================] - 485s 150ms/step - loss: 0.5992 - acc: 0.8260 - val_loss: 1.6506 - val_acc: 0.6059
Epoch 96/200
3232/3232 [==============================] - 486s 150ms/step - loss: 0.6007 - acc: 0.8251 - val_loss: 1.6441 - val_acc: 0.6085
Epoch 97/200
3231/3232 [============================>.] - ETA: 0s - loss: 0.6011 - acc: 0.8257
Epoch 00097: ReduceLROnPlateau reducing learning rate to 9.765624753299562e-08.
3232/3232 [==============================] - 491s 152ms/step - loss: 0.6011 - acc: 0.8256 - val_loss: 1.6462 - val_acc: 0.6092
Epoch 98/200
3232/3232 [==============================] - 490s 152ms/step - loss: 0.6027 - acc: 0.8254 - val_loss: 1.6427 - val_acc: 0.6078
Epoch 99/200
3232/3232 [==============================] - 487s 151ms/step - loss: 0.5980 - acc: 0.8254 - val_loss: 1.6453 - val_acc: 0.6057
Epoch 100/200
3232/3232 [==============================] - 480s 148ms/step - loss: 0.6035 - acc: 0.8231 - val_loss: 1.6516 - val_acc: 0.6065
Epoch 101/200
3232/3232 [==============================] - 489s 151ms/step - loss: 0.5982 - acc: 0.8244 - val_loss: 1.6424 - val_acc: 0.6075
Epoch 102/200
3231/3232 [============================>.] - ETA: 0s - loss: 0.6012 - acc: 0.8250
Epoch 00102: ReduceLROnPlateau reducing learning rate to 4.882812376649781e-08.
3232/3232 [==============================] - 487s 151ms/step - loss: 0.6012 - acc: 0.8250 - val_loss: 1.6413 - val_acc: 0.6091
Epoch 103/200
3232/3232 [==============================] - 491s 152ms/step - loss: 0.5998 - acc: 0.8236 - val_loss: 1.6475 - val_acc: 0.6065
Epoch 104/200
3232/3232 [==============================] - 488s 151ms/step - loss: 0.5979 - acc: 0.8264 - val_loss: 1.6496 - val_acc: 0.6053
Epoch 105/200
3232/3232 [==============================] - 484s 150ms/step - loss: 0.6012 - acc: 0.8242 - val_loss: 1.6449 - val_acc: 0.6067
Epoch 106/200
3232/3232 [==============================] - 489s 151ms/step - loss: 0.5984 - acc: 0.8270 - val_loss: 1.6476 - val_acc: 0.6081
Epoch 107/200
3231/3232 [============================>.] - ETA: 0s - loss: 0.5999 - acc: 0.8235
Epoch 00107: ReduceLROnPlateau reducing learning rate to 2.4414061883248905e-08.
3232/3232 [==============================] - 486s 151ms/step - loss: 0.5999 - acc: 0.8235 - val_loss: 1.6499 - val_acc: 0.6068
Epoch 108/200
3232/3232 [==============================] - 486s 150ms/step - loss: 0.5964 - acc: 0.8263 - val_loss: 1.6412 - val_acc: 0.6097
Epoch 109/200
3232/3232 [==============================] - 482s 149ms/step - loss: 0.6018 - acc: 0.8237 - val_loss: 1.6465 - val_acc: 0.6089
Epoch 110/200
3232/3232 [==============================] - 485s 150ms/step - loss: 0.5970 - acc: 0.8252 - val_loss: 1.6416 - val_acc: 0.6082
Epoch 111/200
3232/3232 [==============================] - 480s 148ms/step - loss: 0.5968 - acc: 0.8265 - val_loss: 1.6495 - val_acc: 0.6059
Epoch 112/200
3231/3232 [============================>.] - ETA: 0s - loss: 0.5972 - acc: 0.8255
Epoch 00112: ReduceLROnPlateau reducing learning rate to 1.2207030941624453e-08.
3232/3232 [==============================] - 493s 152ms/step - loss: 0.5973 - acc: 0.8255 - val_loss: 1.6476 - val_acc: 0.6081
Epoch 00112: early stopping
| MIT | VGG16.ipynb | KoshinoK/RandomExchange |
Stage1の損失と正解率の保存 | h = history.history
stage1_loss = h['loss']
stage1_val_loss = h['val_loss']
stage1_acc = h['acc']
stage1_val_acc = h['val_acc'] | _____no_output_____ | MIT | VGG16.ipynb | KoshinoK/RandomExchange |
全層の学習 Stage1の最良モデルパラメータをロード | import pathlib
checkpoints = pathlib.Path(model_dir).glob('*.h5')
checkpoints = sorted(checkpoints, key=lambda cp:cp.stat().st_mtime)
latest = str(checkpoints[-1])
model.load_weights(latest) | _____no_output_____ | MIT | VGG16.ipynb | KoshinoK/RandomExchange |
全層を学習可能にする | for l in model.layers:
l.trainable = True
model.summary() | _________________________________________________________________
Layer (type) Output Shape Param #
=================================================================
block1_conv1 (Conv2D) (None, 224, 224, 64) 1792
_________________________________________________________________
block1_conv2 (Conv2D) (None, 224, 224, 64) 36928
_________________________________________________________________
block1_pool (MaxPooling2D) (None, 112, 112, 64) 0
_________________________________________________________________
block2_conv1 (Conv2D) (None, 112, 112, 128) 73856
_________________________________________________________________
block2_conv2 (Conv2D) (None, 112, 112, 128) 147584
_________________________________________________________________
block2_pool (MaxPooling2D) (None, 56, 56, 128) 0
_________________________________________________________________
block3_conv1 (Conv2D) (None, 56, 56, 256) 295168
_________________________________________________________________
block3_conv2 (Conv2D) (None, 56, 56, 256) 590080
_________________________________________________________________
block3_conv3 (Conv2D) (None, 56, 56, 256) 590080
_________________________________________________________________
block3_pool (MaxPooling2D) (None, 28, 28, 256) 0
_________________________________________________________________
block4_conv1 (Conv2D) (None, 28, 28, 512) 1180160
_________________________________________________________________
block4_conv2 (Conv2D) (None, 28, 28, 512) 2359808
_________________________________________________________________
block4_conv3 (Conv2D) (None, 28, 28, 512) 2359808
_________________________________________________________________
block4_pool (MaxPooling2D) (None, 14, 14, 512) 0
_________________________________________________________________
block5_conv1 (Conv2D) (None, 14, 14, 512) 2359808
_________________________________________________________________
block5_conv2 (Conv2D) (None, 14, 14, 512) 2359808
_________________________________________________________________
block5_conv3 (Conv2D) (None, 14, 14, 512) 2359808
_________________________________________________________________
block5_pool (MaxPooling2D) (None, 7, 7, 512) 0
_________________________________________________________________
flatten (Flatten) (None, 25088) 0
_________________________________________________________________
dense (Dense) (None, 1024) 25691136
_________________________________________________________________
dropout (Dropout) (None, 1024) 0
_________________________________________________________________
dense_1 (Dense) (None, 1024) 1049600
_________________________________________________________________
dropout_1 (Dropout) (None, 1024) 0
_________________________________________________________________
dense_2 (Dense) (None, 101) 103525
=================================================================
WARNING:tensorflow:Discrepancy between trainable weights and collected trainable weights, did you set `model.trainable` without calling `model.compile` after ?
Total params: 31,563,877
Trainable params: 31,563,877
Non-trainable params: 0
_________________________________________________________________
| MIT | VGG16.ipynb | KoshinoK/RandomExchange |
最適化アルゴリズムなどを指定してモデルをコンパイルする | from tensorflow.keras.optimizers import SGD
model.compile(
loss='categorical_crossentropy',
optimizer=SGD(lr=1e-4, momentum=0.9),
metrics=['accuracy']
) | _____no_output_____ | MIT | VGG16.ipynb | KoshinoK/RandomExchange |
シーケンス生成 | batch_size = 25
img_seq_train = ImageSequence(train_list, train_labels, batch_size=batch_size)
img_seq_validation = ImageSequence(val_list, val_labels, batch_size=batch_size)
print('Train images =', len(img_seq_train) * batch_size)
print('Validation images =', len(img_seq_validation) * batch_size) | Train images = 80800
Validation images = 20200
| MIT | VGG16.ipynb | KoshinoK/RandomExchange |
Callbackの生成★ | from tensorflow.keras.callbacks import ModelCheckpoint, CSVLogger, EarlyStopping, ReduceLROnPlateau
# Callbacksの設定
cp_filepath = os.path.join(dir_weights, 'ep_{epoch:04d}_ls_{loss:.1f}.h5')
cp = ModelCheckpoint(
cp_filepath,
monitor='val_acc',
verbose=0,
save_best_only=True,
save_weights_only=True,
mode='auto'
)
csv_filepath = os.path.join(model_dir, 'stage2_loss.csv')
csv = CSVLogger(csv_filepath, append=True)
es = EarlyStopping(monitor='val_acc', patience=20, verbose=1, mode='auto')
rl = ReduceLROnPlateau(monitor='val_loss', factor=0.5, patience=5, verbose=1, mode='auto', epsilon=0.0001, cooldown=0, min_lr=0) | WARNING:tensorflow:`epsilon` argument is deprecated and will be removed, use `min_delta` instead.
| MIT | VGG16.ipynb | KoshinoK/RandomExchange |
学習 | n_epoch = 500
initial_epoch = len(stage1_loss)
# モデルの学習
history = model.fit_generator(
img_seq_train,
epochs=n_epoch, # 学習するエポック数
steps_per_epoch=len(img_seq_train),
validation_data=img_seq_validation,
validation_steps=len(img_seq_validation),
verbose=1,
callbacks=[cp, csv, es, rl],
initial_epoch=initial_epoch
) | Epoch 113/500
3232/3232 [==============================] - 800s 247ms/step - loss: 1.0005 - acc: 0.7156 - val_loss: 1.5914 - val_acc: 0.6010
Epoch 114/500
3232/3232 [==============================] - 797s 247ms/step - loss: 0.8781 - acc: 0.7448 - val_loss: 1.5427 - val_acc: 0.6180
Epoch 115/500
3232/3232 [==============================] - 797s 247ms/step - loss: 0.8167 - acc: 0.7629 - val_loss: 1.5274 - val_acc: 0.6247
Epoch 116/500
3232/3232 [==============================] - 798s 247ms/step - loss: 0.7525 - acc: 0.7781 - val_loss: 1.5708 - val_acc: 0.6297
Epoch 117/500
3232/3232 [==============================] - 797s 247ms/step - loss: 0.7048 - acc: 0.7932 - val_loss: 1.5540 - val_acc: 0.6318
Epoch 118/500
3232/3232 [==============================] - 797s 247ms/step - loss: 0.6641 - acc: 0.8020 - val_loss: 1.5949 - val_acc: 0.6291
Epoch 119/500
3232/3232 [==============================] - 797s 247ms/step - loss: 0.6321 - acc: 0.8123 - val_loss: 1.6130 - val_acc: 0.6234
Epoch 120/500
3231/3232 [============================>.] - ETA: 0s - loss: 0.5969 - acc: 0.8211
Epoch 00120: ReduceLROnPlateau reducing learning rate to 4.999999873689376e-05.
3232/3232 [==============================] - 798s 247ms/step - loss: 0.5970 - acc: 0.8211 - val_loss: 1.6113 - val_acc: 0.6308
Epoch 121/500
3232/3232 [==============================] - 795s 246ms/step - loss: 0.4443 - acc: 0.8633 - val_loss: 1.6167 - val_acc: 0.6447
Epoch 122/500
3232/3232 [==============================] - 795s 246ms/step - loss: 0.4167 - acc: 0.8718 - val_loss: 1.5966 - val_acc: 0.6505
Epoch 123/500
3232/3232 [==============================] - 797s 247ms/step - loss: 0.4008 - acc: 0.8754 - val_loss: 1.6456 - val_acc: 0.6497
Epoch 124/500
3232/3232 [==============================] - 796s 246ms/step - loss: 0.3804 - acc: 0.8810 - val_loss: 1.6828 - val_acc: 0.6416
Epoch 125/500
3231/3232 [============================>.] - ETA: 0s - loss: 0.3632 - acc: 0.8867
Epoch 00125: ReduceLROnPlateau reducing learning rate to 2.499999936844688e-05.
3232/3232 [==============================] - 797s 247ms/step - loss: 0.3631 - acc: 0.8867 - val_loss: 1.7106 - val_acc: 0.6474
Epoch 126/500
3232/3232 [==============================] - 798s 247ms/step - loss: 0.3005 - acc: 0.9048 - val_loss: 1.6939 - val_acc: 0.6538
Epoch 127/500
3232/3232 [==============================] - 806s 249ms/step - loss: 0.2825 - acc: 0.9110 - val_loss: 1.7165 - val_acc: 0.6579
Epoch 128/500
3232/3232 [==============================] - 817s 253ms/step - loss: 0.2749 - acc: 0.9121 - val_loss: 1.7501 - val_acc: 0.6525
Epoch 129/500
3232/3232 [==============================] - 820s 254ms/step - loss: 0.2679 - acc: 0.9148 - val_loss: 1.7399 - val_acc: 0.6551
Epoch 130/500
3231/3232 [============================>.] - ETA: 0s - loss: 0.2595 - acc: 0.9176
Epoch 00130: ReduceLROnPlateau reducing learning rate to 1.249999968422344e-05.
3232/3232 [==============================] - 820s 254ms/step - loss: 0.2596 - acc: 0.9175 - val_loss: 1.7587 - val_acc: 0.6537
Epoch 131/500
3232/3232 [==============================] - 820s 254ms/step - loss: 0.2359 - acc: 0.9251 - val_loss: 1.7814 - val_acc: 0.6569
Epoch 132/500
3232/3232 [==============================] - 821s 254ms/step - loss: 0.2293 - acc: 0.9266 - val_loss: 1.7864 - val_acc: 0.6538
Epoch 133/500
3232/3232 [==============================] - 820s 254ms/step - loss: 0.2224 - acc: 0.9280 - val_loss: 1.8053 - val_acc: 0.6551
Epoch 134/500
3232/3232 [==============================] - 821s 254ms/step - loss: 0.2197 - acc: 0.9296 - val_loss: 1.8303 - val_acc: 0.6556
Epoch 135/500
3231/3232 [============================>.] - ETA: 0s - loss: 0.2140 - acc: 0.9309
Epoch 00135: ReduceLROnPlateau reducing learning rate to 6.24999984211172e-06.
3232/3232 [==============================] - 822s 254ms/step - loss: 0.2140 - acc: 0.9309 - val_loss: 1.8030 - val_acc: 0.6560
Epoch 136/500
3232/3232 [==============================] - 821s 254ms/step - loss: 0.2012 - acc: 0.9344 - val_loss: 1.8238 - val_acc: 0.6561
Epoch 137/500
3232/3232 [==============================] - 819s 253ms/step - loss: 0.1930 - acc: 0.9365 - val_loss: 1.8182 - val_acc: 0.6564
Epoch 138/500
3232/3232 [==============================] - 820s 254ms/step - loss: 0.1957 - acc: 0.9361 - val_loss: 1.8219 - val_acc: 0.6602
Epoch 139/500
3232/3232 [==============================] - 820s 254ms/step - loss: 0.1906 - acc: 0.9378 - val_loss: 1.8461 - val_acc: 0.6584
Epoch 140/500
3231/3232 [============================>.] - ETA: 0s - loss: 0.1932 - acc: 0.9371
Epoch 00140: ReduceLROnPlateau reducing learning rate to 3.12499992105586e-06.
3232/3232 [==============================] - 820s 254ms/step - loss: 0.1932 - acc: 0.9370 - val_loss: 1.8270 - val_acc: 0.6564
Epoch 141/500
3232/3232 [==============================] - 820s 254ms/step - loss: 0.1843 - acc: 0.9399 - val_loss: 1.8421 - val_acc: 0.6561
Epoch 142/500
3232/3232 [==============================] - 819s 253ms/step - loss: 0.1839 - acc: 0.9395 - val_loss: 1.8519 - val_acc: 0.6555
Epoch 143/500
3232/3232 [==============================] - 819s 253ms/step - loss: 0.1874 - acc: 0.9393 - val_loss: 1.8423 - val_acc: 0.6567
Epoch 144/500
3232/3232 [==============================] - 819s 253ms/step - loss: 0.1796 - acc: 0.9407 - val_loss: 1.8570 - val_acc: 0.6567
Epoch 145/500
3231/3232 [============================>.] - ETA: 0s - loss: 0.1810 - acc: 0.9410
Epoch 00145: ReduceLROnPlateau reducing learning rate to 1.56249996052793e-06.
3232/3232 [==============================] - 820s 254ms/step - loss: 0.1811 - acc: 0.9410 - val_loss: 1.8457 - val_acc: 0.6565
Epoch 146/500
3232/3232 [==============================] - 820s 254ms/step - loss: 0.1767 - acc: 0.9418 - val_loss: 1.8540 - val_acc: 0.6580
Epoch 147/500
3232/3232 [==============================] - 819s 253ms/step - loss: 0.1770 - acc: 0.9426 - val_loss: 1.8753 - val_acc: 0.6570
Epoch 148/500
3232/3232 [==============================] - 819s 253ms/step - loss: 0.1750 - acc: 0.9437 - val_loss: 1.8755 - val_acc: 0.6546
Epoch 149/500
3232/3232 [==============================] - 820s 254ms/step - loss: 0.1793 - acc: 0.9416 - val_loss: 1.8487 - val_acc: 0.6605
Epoch 150/500
3231/3232 [============================>.] - ETA: 0s - loss: 0.1760 - acc: 0.9432
Epoch 00150: ReduceLROnPlateau reducing learning rate to 7.81249980263965e-07.
3232/3232 [==============================] - 820s 254ms/step - loss: 0.1760 - acc: 0.9431 - val_loss: 1.8431 - val_acc: 0.6596
Epoch 151/500
3232/3232 [==============================] - 819s 253ms/step - loss: 0.1734 - acc: 0.9431 - val_loss: 1.8619 - val_acc: 0.6577
Epoch 152/500
3232/3232 [==============================] - 820s 254ms/step - loss: 0.1739 - acc: 0.9435 - val_loss: 1.8627 - val_acc: 0.6601
Epoch 153/500
3232/3232 [==============================] - 821s 254ms/step - loss: 0.1764 - acc: 0.9424 - val_loss: 1.8549 - val_acc: 0.6571
Epoch 154/500
3232/3232 [==============================] - 820s 254ms/step - loss: 0.1703 - acc: 0.9441 - val_loss: 1.8689 - val_acc: 0.6584
Epoch 155/500
3231/3232 [============================>.] - ETA: 0s - loss: 0.1709 - acc: 0.9447
Epoch 00155: ReduceLROnPlateau reducing learning rate to 3.906249901319825e-07.
3232/3232 [==============================] - 820s 254ms/step - loss: 0.1710 - acc: 0.9446 - val_loss: 1.8620 - val_acc: 0.6582
Epoch 156/500
3232/3232 [==============================] - 820s 254ms/step - loss: 0.1721 - acc: 0.9444 - val_loss: 1.8677 - val_acc: 0.6552
Epoch 157/500
3232/3232 [==============================] - 818s 253ms/step - loss: 0.1732 - acc: 0.9437 - val_loss: 1.8700 - val_acc: 0.6587
Epoch 158/500
3232/3232 [==============================] - 820s 254ms/step - loss: 0.1721 - acc: 0.9439 - val_loss: 1.8744 - val_acc: 0.6567
Epoch 159/500
3232/3232 [==============================] - 820s 254ms/step - loss: 0.1722 - acc: 0.9440 - val_loss: 1.8635 - val_acc: 0.6587
Epoch 160/500
3231/3232 [============================>.] - ETA: 0s - loss: 0.1714 - acc: 0.9446
Epoch 00160: ReduceLROnPlateau reducing learning rate to 1.9531249506599124e-07.
3232/3232 [==============================] - 819s 253ms/step - loss: 0.1713 - acc: 0.9446 - val_loss: 1.8742 - val_acc: 0.6561
Epoch 161/500
3232/3232 [==============================] - 821s 254ms/step - loss: 0.1694 - acc: 0.9455 - val_loss: 1.8670 - val_acc: 0.6567
Epoch 162/500
3232/3232 [==============================] - 819s 253ms/step - loss: 0.1721 - acc: 0.9440 - val_loss: 1.8659 - val_acc: 0.6552
Epoch 163/500
3232/3232 [==============================] - 819s 253ms/step - loss: 0.1702 - acc: 0.9442 - val_loss: 1.8738 - val_acc: 0.6578
Epoch 164/500
3232/3232 [==============================] - 819s 254ms/step - loss: 0.1724 - acc: 0.9434 - val_loss: 1.8574 - val_acc: 0.6582
Epoch 165/500
3231/3232 [============================>.] - ETA: 0s - loss: 0.1730 - acc: 0.9443
Epoch 00165: ReduceLROnPlateau reducing learning rate to 9.765624753299562e-08.
3232/3232 [==============================] - 819s 253ms/step - loss: 0.1730 - acc: 0.9443 - val_loss: 1.8597 - val_acc: 0.6591
Epoch 166/500
3232/3232 [==============================] - 819s 253ms/step - loss: 0.1693 - acc: 0.9452 - val_loss: 1.8612 - val_acc: 0.6589
Epoch 167/500
3232/3232 [==============================] - 819s 253ms/step - loss: 0.1726 - acc: 0.9441 - val_loss: 1.8629 - val_acc: 0.6604
Epoch 168/500
3232/3232 [==============================] - 820s 254ms/step - loss: 0.1700 - acc: 0.9450 - val_loss: 1.8632 - val_acc: 0.6578
Epoch 169/500
3232/3232 [==============================] - 818s 253ms/step - loss: 0.1698 - acc: 0.9444 - val_loss: 1.8565 - val_acc: 0.6595
Epoch 00169: early stopping
| MIT | VGG16.ipynb | KoshinoK/RandomExchange |
結果 損失関数のプロット | plt.figure(figsize=(8, 6))
h = history.history
loss = stage1_loss + h['loss']
val_loss = stage1_val_loss + h['val_loss']
ep = np.arange(1, len(loss) + 1)
plt.title('Loss', fontsize=16)
plt.plot(ep, loss, label='Training')
plt.plot(ep, val_loss, label='Validation')
plt.legend(fontsize=16)
plt.xticks(fontsize=14)
plt.yticks(fontsize=14)
plt.xlabel('Epoch', fontsize=16)
plt.ylabel('Loss', fontsize=16)
plt.tight_layout()
save_fig(plt, file_prefix=os.path.join(model_dir, 'Loss'))
plt.show() | _____no_output_____ | MIT | VGG16.ipynb | KoshinoK/RandomExchange |
正解率のプロット | plt.figure(figsize=(8, 6))
h = history.history
acc = stage1_acc + h['acc']
val_acc = stage1_val_acc + h['val_acc']
ep = np.arange(1, len(loss) + 1)
plt.title('Accuracy', fontsize=16)
plt.plot(ep, acc, label='Training')
plt.plot(ep, val_acc, label='Validation')
plt.legend(fontsize=16)
plt.xticks(fontsize=14)
plt.yticks(fontsize=14)
plt.xlabel('Epoch', fontsize=16)
plt.ylabel('Accuracy', fontsize=16)
plt.tight_layout()
save_fig(plt, file_prefix=os.path.join(model_dir, 'Loss'))
plt.show() | _____no_output_____ | MIT | VGG16.ipynb | KoshinoK/RandomExchange |
汎化能力の推定 | from sklearn.metrics import confusion_matrix, accuracy_score, precision_score, recall_score
import seaborn as sns
def evalulate(y_true, y_pred, file_prefix=''):
cm = confusion_matrix(y_true, y_pred)
# print(cm)
accuracy = accuracy_score(y_true, y_pred)
precision = precision_score(y_true, y_pred, average=None)
recall = recall_score(y_true, y_pred, average=None)
print('正解率')
print(f' {accuracy}')
class_labels = []
for i in range(y_true.max() + 1):
class_labels.append(f'{i:4d}')
precision_str = []
recall_str = []
for i in range(y_true.max() + 1):
precision_str.append(f'{precision[i]}')
recall_str.append(f'{recall[i]}')
print('精度')
print(' ' + ' '.join(class_labels))
print(' ' + ' '.join(precision_str))
print('再現率')
print(' ' + ' '.join(class_labels))
print(' ' + ' '.join(recall_str))
plt.figure(figsize = (10,7))
sns.heatmap(cm, annot=True, fmt='3d', square=True, cmap='hot')
plt.tight_layout()
save_fig(plt, file_prefix=file_prefix)
plt.show()
import pathlib
checkpoints = pathlib.Path(model_dir).glob('*.h5')
checkpoints = sorted(checkpoints, key=lambda cp:cp.stat().st_mtime)
latest = str(checkpoints[-1])
model.load_weights(latest)
batch_size = 25
img_seq_validation = ImageSequence(val_list, val_labels, shuffle=False, batch_size=batch_size)
y_pred = model.predict_generator(img_seq_validation)
y_pred_classes = y_pred.argmax(axis=1)
y_true = np.array(val_labels)
evalulate(y_true, y_pred_classes, file_prefix=os.path.join(model_dir, 'cm')) | 正解率
0.6573267326732674
精度
0 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100
0.38235294117647056 0.6367713004484304 0.6298076923076923 0.7309644670050761 0.5566037735849056 0.5891089108910891 0.797979797979798 0.803030303030303 0.45251396648044695 0.505 0.543010752688172 0.7149532710280374 0.7486631016042781 0.6633663366336634 0.5778894472361809 0.44559585492227977 0.5807860262008734 0.5285714285714286 0.5314285714285715 0.6101694915254238 0.6730769230769231 0.5431034482758621 0.4878048780487805 0.7323232323232324 0.6651376146788991 0.7336683417085427 0.46 0.7777777777777778 0.7336448598130841 0.672645739910314 0.8009950248756219 0.6326530612244898 0.8118811881188119 0.9402985074626866 0.7142857142857143 0.7932960893854749 0.6573033707865169 0.4752475247524752 0.6359447004608295 0.39655172413793105 0.7035398230088495 0.7512437810945274 0.5652173913043478 0.7391304347826086 0.6965174129353234 0.7853658536585366 0.6594594594594595 0.5977653631284916 0.6551724137931034 0.5441176470588235 0.5396825396825397 0.7247706422018348 0.7371134020618557 0.648936170212766 0.8571428571428571 0.6666666666666666 0.4647058823529412 0.6136363636363636 0.5625 0.5707070707070707 0.7288888888888889 0.6543778801843319 0.6445497630331753 0.7533039647577092 0.8373205741626795 0.8038277511961722 0.7027027027027027 0.5112359550561798 0.8029556650246306 0.845 0.7453703703703703 0.6966824644549763 0.676923076923077 0.509009009009009 0.7037037037037037 0.8043478260869565 0.7488789237668162 0.3952380952380952 0.7122641509433962 0.7653631284916201 0.5603864734299517 0.7225130890052356 0.5345911949685535 0.7547169811320755 0.5756097560975609 0.65625 0.7395348837209302 0.5110132158590308 0.8238095238095238 0.551219512195122 0.8306878306878307 0.8608247422680413 0.7336956521739131 0.36904761904761907 0.6179245283018868 0.6118721461187214 0.5435897435897435 0.7512953367875648 0.6413043478260869 0.5029940119760479 0.7580645161290323
再現率
0 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100
0.325 0.71 0.655 0.72 0.59 0.595 0.79 0.795 0.405 0.505 0.505 0.765 0.7 0.67 0.575 0.43 0.665 0.555 0.465 0.72 0.7 0.63 0.5 0.725 0.725 0.73 0.46 0.77 0.785 0.75 0.805 0.62 0.82 0.945 0.775 0.71 0.585 0.48 0.69 0.345 0.795 0.755 0.52 0.68 0.7 0.805 0.61 0.535 0.76 0.555 0.51 0.79 0.715 0.61 0.87 0.62 0.395 0.405 0.495 0.565 0.82 0.71 0.68 0.855 0.875 0.84 0.65 0.455 0.815 0.845 0.805 0.735 0.66 0.565 0.665 0.925 0.835 0.415 0.755 0.685 0.58 0.69 0.425 0.8 0.59 0.63 0.795 0.58 0.865 0.565 0.785 0.835 0.675 0.31 0.655 0.67 0.53 0.725 0.59 0.42 0.705
| MIT | VGG16.ipynb | KoshinoK/RandomExchange |
Query Classifier Tutorial[](https://colab.research.google.com/github/deepset-ai/haystack/blob/master/tutorials/Tutorial14_Query_Classifier.ipynb)In this tutorial we introduce the query classifier the goal of introducing this feature was to optimize the overall flow of Haystack pipeline by detecting the nature of user queries. Now, the Haystack can detect primarily three types of queries using both light-weight SKLearn Gradient Boosted classifier or Transformer based more robust classifier. The three categories of queries are as follows: 1. Keyword Queries: Such queries don't have semantic meaning and merely consist of keywords. For instance these three are the examples of keyword queries.* arya stark father* jon snow country* arya stark younger brothers 2. Interrogative Queries: In such queries users usually ask a question, regardless of presence of "?" in the query the goal here is to detect the intent of the user whether any question is asked or not in the query. For example:* who is the father of arya stark ?* which country was jon snow filmed ?* who are the younger brothers of arya stark ? 3. Declarative Queries: Such queries are variation of keyword queries, however, there is semantic relationship between words. Fo example:* Arya stark was a daughter of a lord.* Jon snow was filmed in a country in UK.* Bran was brother of a princess.In this tutorial, you will learn how the `TransformersQueryClassifier` and `SklearnQueryClassifier` classes can be used to intelligently route your queries, based on the nature of the user query. Also, you can choose between a lightweight Gradients boosted classifier or a transformer based classifier.Furthermore, there are two types of classifiers you can use out of the box from Haystack.1. Keyword vs Statement/Question Query Classifier2. Statement vs Question Query ClassifierAs evident from the name the first classifier detects the keywords search queries and semantic statements like sentences/questions. The second classifier differentiates between question based queries and declarative sentences. Prepare environment Colab: Enable the GPU runtimeMake sure you enable the GPU runtime to experience decent speed in this tutorial. **Runtime -> Change Runtime type -> Hardware accelerator -> GPU** These lines are to install Haystack through pip | # Install the latest release of Haystack in your own environment
#! pip install farm-haystack
# Install the latest master of Haystack
!pip install --upgrade pip
!pip install git+https://github.com/deepset-ai/haystack.git#egg=farm-haystack[colab]
# Install pygraphviz
!apt install libgraphviz-dev
!pip install pygraphviz
# In Colab / No Docker environments: Start Elasticsearch from source
! wget https://artifacts.elastic.co/downloads/elasticsearch/elasticsearch-7.9.2-linux-x86_64.tar.gz -q
! tar -xzf elasticsearch-7.9.2-linux-x86_64.tar.gz
! chown -R daemon:daemon elasticsearch-7.9.2
import os
from subprocess import Popen, PIPE, STDOUT
es_server = Popen(
["elasticsearch-7.9.2/bin/elasticsearch"], stdout=PIPE, stderr=STDOUT, preexec_fn=lambda: os.setuid(1) # as daemon
)
# wait until ES has started
! sleep 30 | _____no_output_____ | Apache-2.0 | tutorials/Tutorial14_Query_Classifier.ipynb | askainet/haystack |
If running from Colab or a no Docker environment, you will want to start Elasticsearch from source InitializationHere are some core imports Then let's fetch some data (in this case, pages from the Game of Thrones wiki) and prepare it so that it canbe used indexed into our `DocumentStore` | from haystack.utils import (
print_answers,
print_documents,
fetch_archive_from_http,
convert_files_to_docs,
clean_wiki_text,
launch_es,
)
from haystack.pipelines import Pipeline
from haystack.document_stores import ElasticsearchDocumentStore
from haystack.nodes import (
BM25Retriever,
EmbeddingRetriever,
FARMReader,
TransformersQueryClassifier,
SklearnQueryClassifier,
)
# Download and prepare data - 517 Wikipedia articles for Game of Thrones
doc_dir = "data/tutorial14"
s3_url = "https://s3.eu-central-1.amazonaws.com/deepset.ai-farm-qa/datasets/documents/wiki_gameofthrones_txt14.zip"
fetch_archive_from_http(url=s3_url, output_dir=doc_dir)
# convert files to dicts containing documents that can be indexed to our datastore
got_docs = convert_files_to_docs(dir_path=doc_dir, clean_func=clean_wiki_text, split_paragraphs=True)
# Initialize DocumentStore and index documents
launch_es()
document_store = ElasticsearchDocumentStore()
document_store.delete_documents()
document_store.write_documents(got_docs)
# Initialize Sparse retriever
bm25_retriever = BM25Retriever(document_store=document_store)
# Initialize dense retriever
embedding_retriever = EmbeddingRetriever(
document_store=document_store,
model_format="sentence_transformers",
embedding_model="sentence-transformers/multi-qa-mpnet-base-dot-v1",
)
document_store.update_embeddings(embedding_retriever, update_existing_embeddings=False)
reader = FARMReader(model_name_or_path="deepset/roberta-base-squad2") | _____no_output_____ | Apache-2.0 | tutorials/Tutorial14_Query_Classifier.ipynb | askainet/haystack |
Keyword vs Question/Statement ClassifierThe keyword vs question/statement query classifier essentially distinguishes between the keyword queries and statements/questions. So you can intelligently route to different retrieval nodes based on the nature of the query. Using this classifier can potentially yield the following benefits:* Getting better search results (e.g. by routing only proper questions to DPR / QA branches and not keyword queries)* Less GPU costs (e.g. if 50% of your traffic is only keyword queries you could just use elastic here and save the GPU resources for the other 50% of traffic with semantic queries)![image]() Below, we define a `SklearnQueryClassifier` and show how to use it:Read more about the trained model and dataset used [here](https://ext-models-haystack.s3.eu-central-1.amazonaws.com/gradboost_query_classifier/readme.txt) | # Here we build the pipeline
sklearn_keyword_classifier = Pipeline()
sklearn_keyword_classifier.add_node(component=SklearnQueryClassifier(), name="QueryClassifier", inputs=["Query"])
sklearn_keyword_classifier.add_node(
component=embedding_retriever, name="EmbeddingRetriever", inputs=["QueryClassifier.output_1"]
)
sklearn_keyword_classifier.add_node(component=bm25_retriever, name="ESRetriever", inputs=["QueryClassifier.output_2"])
sklearn_keyword_classifier.add_node(component=reader, name="QAReader", inputs=["ESRetriever", "EmbeddingRetriever"])
sklearn_keyword_classifier.draw("pipeline_classifier.png")
# Run only the dense retriever on the full sentence query
res_1 = sklearn_keyword_classifier.run(query="Who is the father of Arya Stark?")
print("Embedding Retriever Results" + "\n" + "=" * 15)
print_answers(res_1, details="minimum")
# Run only the sparse retriever on a keyword based query
res_2 = sklearn_keyword_classifier.run(query="arya stark father")
print("ES Results" + "\n" + "=" * 15)
print_answers(res_2, details="minimum")
# Run only the dense retriever on the full sentence query
res_3 = sklearn_keyword_classifier.run(query="which country was jon snow filmed ?")
print("Embedding Retriever Results" + "\n" + "=" * 15)
print_answers(res_3, details="minimum")
# Run only the sparse retriever on a keyword based query
res_4 = sklearn_keyword_classifier.run(query="jon snow country")
print("ES Results" + "\n" + "=" * 15)
print_answers(res_4, details="minimum")
# Run only the dense retriever on the full sentence query
res_5 = sklearn_keyword_classifier.run(query="who are the younger brothers of arya stark ?")
print("Embedding Retriever Results" + "\n" + "=" * 15)
print_answers(res_5, details="minimum")
# Run only the sparse retriever on a keyword based query
res_6 = sklearn_keyword_classifier.run(query="arya stark younger brothers")
print("ES Results" + "\n" + "=" * 15)
print_answers(res_6, details="minimum") | _____no_output_____ | Apache-2.0 | tutorials/Tutorial14_Query_Classifier.ipynb | askainet/haystack |
Transformer Keyword vs Question/Statement ClassifierFirstly, it's essential to understand the trade-offs between SkLearn and Transformer query classifiers. The transformer classifier is more accurate than SkLearn classifier however, it requires more memory and most probably GPU for faster inference however the transformer size is roughly `50 MBs`. Whereas, SkLearn is less accurate however is much more faster and doesn't require GPU for inference.Below, we define a `TransformersQueryClassifier` and show how to use it:Read more about the trained model and dataset used [here](https://huggingface.co/shahrukhx01/bert-mini-finetune-question-detection) | # Here we build the pipeline
transformer_keyword_classifier = Pipeline()
transformer_keyword_classifier.add_node(
component=TransformersQueryClassifier(), name="QueryClassifier", inputs=["Query"]
)
transformer_keyword_classifier.add_node(
component=embedding_retriever, name="EmbeddingRetriever", inputs=["QueryClassifier.output_1"]
)
transformer_keyword_classifier.add_node(
component=bm25_retriever, name="ESRetriever", inputs=["QueryClassifier.output_2"]
)
transformer_keyword_classifier.add_node(component=reader, name="QAReader", inputs=["ESRetriever", "EmbeddingRetriever"])
transformer_keyword_classifier.draw("pipeline_classifier.png")
# Run only the dense retriever on the full sentence query
res_1 = transformer_keyword_classifier.run(query="Who is the father of Arya Stark?")
print("Embedding Retriever Results" + "\n" + "=" * 15)
print_answers(res_1, details="minimum")
# Run only the sparse retriever on a keyword based query
res_2 = transformer_keyword_classifier.run(query="arya stark father")
print("ES Results" + "\n" + "=" * 15)
print_answers(res_2, details="minimum")
# Run only the dense retriever on the full sentence query
res_3 = transformer_keyword_classifier.run(query="which country was jon snow filmed ?")
print("Embedding Retriever Results" + "\n" + "=" * 15)
print_answers(res_3, details="minimum")
# Run only the sparse retriever on a keyword based query
res_4 = transformer_keyword_classifier.run(query="jon snow country")
print("ES Results" + "\n" + "=" * 15)
print_answers(res_4, details="minimum")
# Run only the dense retriever on the full sentence query
res_5 = transformer_keyword_classifier.run(query="who are the younger brothers of arya stark ?")
print("Embedding Retriever Results" + "\n" + "=" * 15)
print_answers(res_5, details="minimum")
# Run only the sparse retriever on a keyword based query
res_6 = transformer_keyword_classifier.run(query="arya stark younger brothers")
print("ES Results" + "\n" + "=" * 15)
print_answers(res_6, details="minimum") | _____no_output_____ | Apache-2.0 | tutorials/Tutorial14_Query_Classifier.ipynb | askainet/haystack |
Question vs Statement ClassifierOne possible use case of this classifier could be to route queries after the document retrieval to only send questions to QA reader and in case of declarative sentence, just return the DPR/ES results back to user to enhance user experience and only show answers when user explicitly asks it.![image]() Below, we define a `TransformersQueryClassifier` and show how to use it:Read more about the trained model and dataset used [here](https://huggingface.co/shahrukhx01/question-vs-statement-classifier) | # Here we build the pipeline
transformer_question_classifier = Pipeline()
transformer_question_classifier.add_node(component=embedding_retriever, name="EmbeddingRetriever", inputs=["Query"])
transformer_question_classifier.add_node(
component=TransformersQueryClassifier(model_name_or_path="shahrukhx01/question-vs-statement-classifier"),
name="QueryClassifier",
inputs=["EmbeddingRetriever"],
)
transformer_question_classifier.add_node(component=reader, name="QAReader", inputs=["QueryClassifier.output_1"])
transformer_question_classifier.draw("question_classifier.png")
# Run only the QA reader on the question query
res_1 = transformer_question_classifier.run(query="Who is the father of Arya Stark?")
print("Embedding Retriever Results" + "\n" + "=" * 15)
print_answers(res_1, details="minimum")
res_2 = transformer_question_classifier.run(query="Arya Stark was the daughter of a Lord.")
print("ES Results" + "\n" + "=" * 15)
print_documents(res_2) | _____no_output_____ | Apache-2.0 | tutorials/Tutorial14_Query_Classifier.ipynb | askainet/haystack |
Standalone Query ClassifierBelow we run queries classifiers standalone to better understand their outputs on each of the three types of queries | # Here we create the keyword vs question/statement query classifier
from haystack.nodes import TransformersQueryClassifier
queries = [
"arya stark father",
"jon snow country",
"who is the father of arya stark",
"which country was jon snow filmed?",
]
keyword_classifier = TransformersQueryClassifier()
for query in queries:
result = keyword_classifier.run(query=query)
if result[1] == "output_1":
category = "question/statement"
else:
category = "keyword"
print(f"Query: {query}, raw_output: {result}, class: {category}")
# Here we create the question vs statement query classifier
from haystack.nodes import TransformersQueryClassifier
queries = [
"Lord Eddard was the father of Arya Stark.",
"Jon Snow was filmed in United Kingdom.",
"who is the father of arya stark?",
"Which country was jon snow filmed in?",
]
question_classifier = TransformersQueryClassifier(model_name_or_path="shahrukhx01/question-vs-statement-classifier")
for query in queries:
result = question_classifier.run(query=query)
if result[1] == "output_1":
category = "question"
else:
category = "statement"
print(f"Query: {query}, raw_output: {result}, class: {category}") | _____no_output_____ | Apache-2.0 | tutorials/Tutorial14_Query_Classifier.ipynb | askainet/haystack |
Chapter 3: Inferential statistics[Link to outline](https://docs.google.com/document/d/1fwep23-95U-w1QMPU31nOvUnUXE2X3s_Dbk5JuLlKAY/editheading=h.uutryzqeo2av)Concept map: Notebook setup | # loading Python modules
import math
import random
import numpy as np
import pandas as pd
import seaborn as sns
from scipy.stats.distributions import norm
# set random seed for repeatability
np.random.seed(42)
# notebooks figs setup
%matplotlib inline
import matplotlib.pyplot as plt
sns.set(rc={'figure.figsize':(8,5)})
blue, orange = sns.color_palette()[0], sns.color_palette()[1]
# silence annoying warnings
import warnings; warnings.filterwarnings('ignore') | _____no_output_____ | MIT | stats_overview/03_STATS.ipynb | minireference/noBSstatsnotebooks |
Overview- Main idea = learn about a population based on a sample- Recall Amy's two research questions about the employee lifetime value (ELV) data: - Question 1 = Is there a difference between ELV of the two groups? → **hypothesis testing** - Question 2 = How much difference in ELV does stats training provide? → **estimation**- Inferential statistics provides us with tools to answer both of these questions EstimatorsWe'll begin our study of inferential statistics by introducing **estimators**,which are used for both **hypothesis testing** and **estimation**. $\def\stderr1{\mathbf{se}_{1}}$$\def\stderrhat1{\hat{\mathbf{se}}_{1}}$ Definitions- We use the term "estimator" to describe a function $f$ that takes samples as inputs, which is written mathematically as: $$ f \ \colon \underbrace{\mathcal{X}\times \mathcal{X}\times \cdots \times \mathcal{X}}_{n \textrm{ copies}} \quad \to \quad \mathbb{R}, $$ where $n$ is the samples size and $\mathcal{X}$ denotes the possible values of the random variable $X$.- We give different names to estimators, depending on the use case: - **statistic** = a function computed from samples (descriptive statistics) - **parameter estimators** = statistics that estimates population parameters - **test statistic** = an estimator used as part of hypothesis testing procedure- The **value** of the estimator $f(\mathbf{x})$ is computer from a particular sample $\mathbf{x}$.- The **sampling distribution** of an estimator is when $f$ is the distribution of $f(\mathbf{X})$, where $\mathbf{X}$ is a random sample.- Example of estimators we discussed in descriptive statistics: - Sample mean - estimator: $\overline{x} = g(\mathbf{x}) = \frac{1}{n}\sum_{i=1}^n x_i$ - gives an estimate for the population mean $\mu$ - sampling distribution: $\overline{X} = g(\mathbf{X}) = \frac{1}{n}\sum_{i=1}^n X_i$ - Sample variance - estimator: $s^2 = h(\mathbf{x}) = \frac{1}{n-1}\sum_{i=1}^n (x_i-\overline{x})^2$ - gives an estimate for the population variance $\sigma^2$ - sampling distribution: $S^2 = h(\mathbf{X}) = \frac{1}{n-1}\sum_{i=1}^n (X_i-\overline{X})^2$ - In this notebook we focus on one estimator: **difference between group means** - estimator: $d = \texttt{mean}(\mathbf{x}_A) - \texttt{mean}(\mathbf{x}_{B}) = \overline{x}_{A} - \overline{x}_{B}$ - gives an estimate for the difference between population means: $\Delta = \mu_A - \mu_{B}$ - sampling distribution: $D = \overline{X}_A - \overline{X}_{B}$, which is a random variable Difference between group meansConsider two random variables $X_A$ and $X_B$:$$ \largeX_A \sim \mathcal{N}\!\left(\mu_A, \sigma^2_A \right)\qquad\textrm{and}\qquadX_B \sim \mathcal{N}\!\left(\mu_B, \sigma^2_B \right)$$that describe the probability distribution for groups A and B, respectively.- A sample of size $n_A$ from $X_A$ is denoted $\mathbf{x}_A = x_1x_2\cdots x_{n_A}$=`xA`, and let $\mathbf{x}_B = x_1x_2\cdots x_{n_B}$=`xB` be a random sample of size $n_B$ from $X_B$.- We compute the mean in each group: $\overline{x}_{A} = \texttt{mean}(\mathbf{x}_A)$ and $\overline{x}_{B} = \texttt{mean}(\mathbf{x}_B)$- The value of the estimator is $d = \overline{x}_{A} - \overline{x}_{B}$ | def dmeans(xA, xB):
"""
Estimator for the difference between group means.
"""
d = np.mean(xA) - np.mean(xB)
return d | _____no_output_____ | MIT | stats_overview/03_STATS.ipynb | minireference/noBSstatsnotebooks |
Note the difference between group means is precisely the estimator Amy need for her analysis (**Group S** and **Group NS**). We intentionally use the labels **A** and **B** to illustrate the general case. | # example parameters for each group
muA, sigmaA = 300, 10
muB, sigmaB = 200, 20
# size of samples for each group
nA = 5
nB = 4 | _____no_output_____ | MIT | stats_overview/03_STATS.ipynb | minireference/noBSstatsnotebooks |
Particular value of the estimator `dmeans` | xA = norm(muA, sigmaA).rvs(nA) # random sample from Group A
xB = norm(muB, sigmaB).rvs(nB) # random sample from Group B
d = dmeans(xA, xB)
d | _____no_output_____ | MIT | stats_overview/03_STATS.ipynb | minireference/noBSstatsnotebooks |
The value of $d$ computed from the samples is an estimate for the difference between means of two groups: $\Delta = \mu_A - \mu_{B}$ (which we know is $100$ in this example). Sampling distribution of the estimator `dmeans`How well does the estimate $d$ approximate the true value $\Delta$?**What is the accuracy and variability of the estimates we can expect?**To answer these questions, consider the random samples$\mathbf{X}_A = X_1X_2\cdots X_{n_A}$and $\mathbf{X}_B = X_1X_2\cdots X_{n_B}$,then compute the **sampling distribution**: $D = \overline{X}_A - \overline{X}_{B}$.By definition, the sampling distribution of the estimator is obtained by repeatedly generating samples `xA` and `xB` from the two distributions and computing `dmeans` on the random samples. For example, we can obtain the sampling distribution by generating $N=1000$ samples. | def get_sampling_dist(statfunc, meanA, stdA, nA, meanB, stdB, nB, N=1000):
"""
Obtain the sampling distribution of the statistic `statfunc`
from `N` random samples drawn from groups A and B with parmeters:
- Group A: `nA` values taken from `norm(meanA, stdA)`
- Group B: `nB` values taken from `norm(meanB, stdB)`
Returns a list of samples from the sampling distribution of `statfunc`.
"""
sampling_dist = []
for i in range(0, N):
xA = norm(meanA, stdA).rvs(nA) # random sample from Group A
xB = norm(meanB, stdB).rvs(nB) # random sample from Group B
stat = statfunc(xA, xB) # evaluate `statfunc`
sampling_dist.append(stat) # record the value of statfunc
return sampling_dist
# Generate the sampling distirbution for dmeans
dmeans_sdist = get_sampling_dist(statfunc=dmeans,
meanA=muA, stdA=sigmaA, nA=nA,
meanB=muB, stdB=sigmaB, nB=nB)
print("Generated", len(dmeans_sdist), "values from `dmeans(XA, XB)`")
# first 3 values
dmeans_sdist[0:3] | _____no_output_____ | MIT | stats_overview/03_STATS.ipynb | minireference/noBSstatsnotebooks |
Plot the sampling distribution of `dmeans` | fig3, ax3 = plt.subplots()
title3 = "Samping distribution of D = mean($\mathbf{X}_A$) - mean($\mathbf{X}_B$) " + \
"for samples of size $n_A$ = " + str(nA) + \
" from $\mathcal{N}$(" + str(muA) + "," + str(sigmaA) + ")" + \
" and $n_B$ = " + str(nB) + \
" from $\mathcal{N}$(" + str(muB) + "," + str(sigmaB) + ")"
sns.distplot(dmeans_sdist, kde=False, norm_hist=True, ax=ax3)
_ = ax3.set_title(title3) | _____no_output_____ | MIT | stats_overview/03_STATS.ipynb | minireference/noBSstatsnotebooks |
Theoretical model for the sampling distribution of `dmeans` Let's use probability theory to build a theoretical model for the sampling distribution of the difference-between-means estimator `dmeans`.- The central limit theorem the rules of to obtain a model for the random variable $D = \overline{X}_A - \overline{X}_{B}$,which describes the sampling distribution of `dmeans`.- The central limit theorem tells us the sample mean within the two group are $$ \large \overline{X}_A \sim \mathcal{N}\!\left(\mu_A, \tfrac{\sigma^2_A}{n_A} \right) \qquad \textrm{and} \qquad \overline{X}_B \sim \mathcal{N}\!\left(\mu_B, \tfrac{\sigma^2_B}{n_B} \right) $$- The rules of probability theory tells us that the [difference of two normal random variables](https://en.wikipedia.org/wiki/Sum_of_normally_distributed_random_variablesIndependent_random_variables) requires subtracting their means and adding their variance, so we get: $$ \large D \sim \mathcal{N}\!\left(\mu_A - \mu_B, \ \tfrac{\sigma^2_A}{n_A} + \tfrac{\sigma^2_B}{n_B} \right) $$In other words, the sampling distribution for the difference of means estimator has mean and standard deviation given by:$$ \large \mu_D = \mu_A - \mu_B \qquad \textrm{and} \qquad \sigma_D = \sqrt{ \tfrac{\sigma^2_A}{n_A} + \tfrac{\sigma^2_B}{n_B} }$$Let's plot the theoretical prediction on top of the simulated data to see if they are a good fit. | Dmean = muA - muB
Dstd = np.sqrt(sigmaA**2/nA + sigmaB**2/nB)
print("Probability theory predicts the sampling distribution had"
"mean", round(Dmean, 3),
"and standard deviation", round(Dstd, 3))
x = np.linspace(min(dmeans_sdist), max(dmeans_sdist), 10000)
D = norm(Dmean, Dstd).pdf(x)
label = 'Theory prediction'
ax3 = sns.lineplot(x, D, ax=ax3, label=label, color=blue)
fig3 | Probability theory predicts the sampling distribution hadmean 100 and standard deviation 10.954
| MIT | stats_overview/03_STATS.ipynb | minireference/noBSstatsnotebooks |
Regroup and reality checkHow are you doing, dear readers?I know this was a lot of math and a lot of code, but the good news is we're done now!The key things to remember is that we have two ways to compute sampling distribution for any estimator: - Repeatedly generate random samples from model and compute the estimator values (histogram) - Use probability theory to obtain a analytical formula Why are we doing all this modelling?The estimator `dmeans` we defined above measures the quantity we're interested in: the difference between the means of two groups (**Group S** and **Group NS** in Amy's statistical analysis of ELV data).Using the functions we developed above, we now have the ability to simulate the data from any two groups by simply choosing the appropriate parameters. In particular if we choose `stdS=266`, `nS=30`; and `stdNS=233`, `nNS=31`,we can generate random data that has similar variability to Amy ELV measurements. Okay, dear reader, we're about to jump into the deep end of the statistics pool: **hypothesis testing**,which is one of the two major ideas in the STATS 101 curriculum.Heads up this will get complicated, but we have to go into it because it is an essential procedurethat is used widely in science, engineering, business, and other types of research.You need to trust me this one: it's worth knowing this stuff, even if it is boring.Don't worry about it though, since you have all the prerequisites needed to get through this! ____Recall Amy's research Question 1: Is there a difference between ELV of the employees in **Group S** and the employees in **Group NS**? Hypothesis testing- An approach to formulating research questions as **yes-no decisions** and a **procedure for making these decisions**- Hypothesis testing is a standardized procedure for doing statistical analysis (also, using stats jargon makes everything look more convincing ;)- We formulate research question as two **competing hypotheses**: - **Null hypothesis $H_0$** = no effect in our example: "no difference between means," which is written as $\color{red}{\mu_S = \mu_{NS} = \mu_0}$. In other words, the probability models for the two groups are: $$ \large H_0: \qquad X_S = \mathcal{N}(\color{red}{\mu_0}, \sigma_S) \quad \textrm{and} \quad X_{NS} = \mathcal{N}(\color{red}{\mu_0}, \sigma_{NS}) \quad $$ - **Alternative hypothesis $H_A$** = an effect exists in our example: "means for Group S different from mean for Group NS", $\color{blue}{\mu_S} \neq \color{orange}{\mu_{NS}}$. The probability models for the two groups are: $$ H_A: \qquad X_S = \mathcal{N}(\color{blue}{\mu_S}, \sigma_S) \quad \textrm{and} \quad X_{NS} = \mathcal{N}(\color{orange}{\mu_{NS}}, \sigma_{NS}) $$- The purpose of hypothesis testing is to perform a basic sanity-check to show the difference between the group means we observed ($d = \overline{x}_{S} - \overline{x}_{NS} = 130$) is **unlikely to have occurred by chance**- NEW CONCEPT: $p$-value is the probability of observing $d$ or more extreme under the null hypothesis. Overview of the hypothesis testing procedureHere is the high-level overview of the hypothesis testing procedure:- **inputs**: sample statistics computed from the observed data (in our case the signal $\overline{x}_S$, $\overline{x}_{NS}$, and our estimates of the noise $s^2_S$, and $s^2_{NS}$)- **outputs**: a decision that is one of: "reject the null hypothesis" or "fail to reject the null hypothesis"We'll now look at two different approaches for computing the sampling distribution ofthe difference between group means statistic, $D = \overline{X}_S - \overline{X}_{NS}$:permutation tests and analytical approximations. Interpreting the results of hypothesis testing (optional) - The implication of rejecting the null hypothesis (no difference) is that there must is a difference between the group means. In other words, the ELV data for employees who took the statistics training (**Group S**) is different form the average ELV for employees who didn't take the statistics training (**Group NS**), which is what Amy is trying to show. - Note that rejecting null hypothesis (H0) is not the same as "proving" the alternative hypothesis (HA), we have just shown that the data is unlikely under the null hypothesis and we must be *some* difference between the groups, so is worth looking for *some* alternative hypothesis. - The alternative hypothesis we picked above, $\mu_S \neq \mu_{NS}$, is just a placeholder, that includes desirable effect: $\mu_S > \mu_{NS}$ (stats training improves ELV), but also includes the opposite effect: $\mu_S < \mu_{NS}$ (stats training decreases ELV). - Using statistics jargon, when we reject the hypothesis H0 we say we've observed a "statistically significant" result, which sounds a lot more impressive statement than it actually is. Recall hypothesis test is just used to rule out "occurred by chance," which is a very basic sanity check. - The implication of failing to reject the null hypothesis is that the observed difference between means is "not significant," meaning it could have occurred by chance, so there is no need to search for an alternative hypothesis. - Note that "failing to reject" is not the same as "proving" the null hypothesis - Note also "failing to reject H0" doesn't mean we reject HA. In fact, the alternative hypothesis didn't play any role in the calculations whatsoever.I know all this sounds super complicated and roundabout (an it is!),but you will get a hang of it in no time with some practice.Trust me, you need to know this shit. Start by load data again...First things first, let's reload the data which we prepared back in the DATA where we left off back in the [01_DATA.ipynb](./01_DATA.ipynb) notebook. | df = pd.read_csv('data/employee_lifetime_values.csv')
df
# remember the descriptive statistics
df.groupby("group").describe()
def dmeans(sample):
"""
Compute the difference between groups means.
"""
xS = sample[sample["group"]=="S"]["ELV"]
xNS = sample[sample["group"]=="NS"]["ELV"]
d = np.mean(xS) - np.mean(xNS)
return d
# the observed value in Amy's data
dmeans(df) | _____no_output_____ | MIT | stats_overview/03_STATS.ipynb | minireference/noBSstatsnotebooks |
Our goal is to determine how likely or unlikely this observed value is under the null hypothesis $H_0$.In the next two sections, we'll look at two different approaches for obtaining the sampling distribution of $D$ under $H_0$. Approach 1: Permutation test for hypothesis testing- The permutation test allow us to reject $H_0$ using existing sample $\mathbf{x}$ that we have, treating the sample as if it were a population.- Relevant probability distributions: - Sampling distribution = obtained from repeated samples from a hypothetical population under $H_0$. - Approximate sampling distribution: obtained by **resampling data from the single sample we have**.- Recall Goal 1: make sure data cannot be explained by $H_0$ (observed difference due to natural variability) - We want to obtain an approximation of the sampling distribution under $H_0$ - The $H_0$ probability model describes a hypothetical scenario with **no difference between groups**, which means data from **Group S** and **Group NS** comes the same distribution. - To generate a new random sample $\mathbf{x}^p$ from $H_0$ model we can reuse the sample we have obtained $\mathbf{x}$, but randomly mix-up the group labels. Since under the $H_0$ model, the **S** and **NS** populations are identical, mixing up the labels should have no effect. - The math term for "mixing up" is **permutation**, meaning each value is input is randomly reassigned to a new random place in the output. | def resample_under_H0(sample, groupcol="group"):
"""
Return a copy of the dataframe `sample` with the labels in the column `groupcol`
modified based on a random permutation of the values in the original sample.
"""
resample = sample.copy()
labels = sample[groupcol].values
newlabels = np.random.permutation(labels)
resample[groupcol] = newlabels
return resample
resample_under_H0(df)
# resample
resample = resample_under_H0(df)
# compute the difference in means for the new labels
dmeans(resample) | _____no_output_____ | MIT | stats_overview/03_STATS.ipynb | minireference/noBSstatsnotebooks |
The steps in the above code cell give us a simple way to generate samples from the null hypothesis and compute the value of `dmeans` statistic for these samples. We used the assumption of "no difference" under the null hypothesis, and translated this to the "forget the labels" interpretation. Running a permutation testWe can repeat the resampling procedure `10000` times to get the sampling distribution of $D$ under $H_0$,as illustrated in the code procedure below. | def permutation_test(sample, statfunc, groupcol="group", permutations=10000):
"""
Compute the p-value of the observed `statfunc(sample)` under the null hypothesis
where the labels in the `groupcol` are randomized.
"""
# 1. compute the observed value of the statistic for the sample
obsstat = statfunc(sample)
# 2. generate the sampling distr. using random permutations of the group labels
resampled_stats = []
for i in range(0, permutations):
resample = resample_under_H0(sample, groupcol=groupcol)
restat = statfunc(resample)
resampled_stats.append(restat)
# 3. compute p-value: how many `restat`s are equal-or-more-extreme than `obsstat`
tailstats = [restat for restat in resampled_stats \
if restat <= -abs(obsstat) or restat >= abs(obsstat)]
pvalue = len(tailstats) / len(resampled_stats)
return resampled_stats, pvalue
sampling_dist, pvalue = permutation_test(df, statfunc=dmeans)
# plot the sampling distribution in blue
sns.displot(sampling_dist, bins=200)
# plot red line for the observed statistic
obsstat = dmeans(df)
plt.axvline(obsstat, color='r')
# plot the values that are equal or more extreme in red
tailstats = [rs for rs in sampling_dist if rs <= -obsstat or rs >= obsstat]
_ = sns.histplot(tailstats, bins=200, color="red") | _____no_output_____ | MIT | stats_overview/03_STATS.ipynb | minireference/noBSstatsnotebooks |
- Once we have the sampling distribution of `D` under $H_0$, we can see where the observed value $d=130$ falls within this distribution.- p-value: the probability of observing value $d$ or more extreme under the null hypothesis | pvalue | _____no_output_____ | MIT | stats_overview/03_STATS.ipynb | minireference/noBSstatsnotebooks |
We can now make the decision based on the $p$-value and a pre-determined threshold:- If the observed value $d$ is unlikely under $H_0$ ($p$-value less than 5% chance of occurring), then our decision will be to "reject the null hypothesis."- Otherwise, if the observed value $d$ is not that unusual ($p$-value greater than 5%), we conclude that we have "failed to reject the null hypothesis." | if pvalue < 0.05:
print("DECISION: Reject H0", "( p-value =", pvalue, ")")
print(" There is a statistically significant difference between xS and xNS means")
else:
print("DECISION: Fail to reject H0")
print(" The difference between groups means could have occurred by chance") | DECISION: Reject H0 ( p-value = 0.046 )
There is a statistically significant difference between xS and xNS means
| MIT | stats_overview/03_STATS.ipynb | minireference/noBSstatsnotebooks |
Permutations test using SciPy The above code was given only for illustrative purposes.In practice, you can use the SciPy implementation of permutation test,by calling `ttest_ind(..., permutations=10000)` to perform a permutation test, then obtain the $p$-value. | from scipy.stats import ttest_ind
xS = df[df["group"]=="S"]["ELV"]
xNS = df[df["group"]=="NS"]["ELV"]
ttest_ind(xS, xNS, permutations=10000).pvalue | _____no_output_____ | MIT | stats_overview/03_STATS.ipynb | minireference/noBSstatsnotebooks |
Discussion - The procedure we used is called a **permutations test** for comparison of group means. - The permutation test takes it's name from the action of mixing up the group-membership labels and computing a statistic which is a way to generate samples from the null hypothesis in situations where we're comparing two groups. - Permutation tests are very versatile since we can use them for any estimator $h(\mathbf{x})$. For example, we could have used difference in medians by specifying the `median` as the input `statfunc`. Approach 2: Analytical approximations for hypothesis testingWe'll now look at another approach for answering Question 1:using and analytical approximation,which is the way normally taught in STATS 101 courses.How likely or unlikely is the observed difference $d=130$ under the null hypothesis?- Analytical approximations are math models for describing the sampling distribution under $H_0$ - Sampling distributions = obtained by repeated sampling from $H_0$ - Analytical approximation = probability distribution model based on estimated parameters- Assumption: population is normally distributed- Based on this assumption we can use the theoretical model we developed above for difference between group means to obtain a **closed form expression** for the sampling distribution of $D$- In particular, the probability model for the two groups under $H_0$ are: $$ \large H_0: \qquad X_S = \mathcal{N}(\color{red}{\mu_0}, \sigma_S) \quad \textrm{and} \quad X_{NS} = \mathcal{N}(\color{red}{\mu_0}, \sigma_{NS}), \quad $$ from which we can derive the model for $D = \overline{X}_S - \overline{X}_{NS}$: $$ \large D \sim \mathcal{N}\!\left( \color{red}{0}, \ \tfrac{\sigma^2_S}{n_S} + \tfrac{\sigma^2_{NS}}{n_{NS}} \right) $$ In words, the sampling distribution of the difference between group means is normally distributed with mean $\mu_D = 0$ and variance $\sigma^2_D$ dependent on the variance of the two groups $\sigma^2_S$ and $\sigma^2_{NS}$. Recall we obtained this expression earlier when we discussed difference of means between groups A and B.- However, the population variances are unknown $\sigma^2_S$ and $\sigma^2_{NS}$, and we only have the estimated variances $s_S^2$ and $s_{NS}^2$ calculated from the sample.- That's OK though, since sample variances are good approximation to the population variances. There are two common ways to obtain an approximation for $\sigma^2_D$: - Pooled variance: $\sigma^2_D \approx s^2_p = \frac{(n_S-1)s_S^2 \; + \; (n_{NS}-1)s_{NS}^2}{n_S + n_{NS} - 2}$ (takes advantage of assumption that both samples come from the same population under $H_0$) - Unpooled variance: $\sigma^2_D \approx s^2_u = \tfrac{s^2_S}{n_S} + \tfrac{s^2_{NS}}{n_{NS}}$ (follows from general rule of prob theory)- NEW CONCEPT: **Student's $t$-distribution** is a model for $D$ which takes into account we are using $s_S^2$ and $s_{NS}^2$ instead of $\sigma_S^2$ and $\sigma_{NS}^2$.- NEW CONCEPT: **degrees of freedom**, denoted `dof` in code or $\nu$ (Greek letter *nu*) in equations, is the parameter Student's $t$ distribution related to the sample size used to estimate quantities. Student's t-test (pooled variance)[Student's t-test for comparison of difference between groups means](https://statkat.com/stattest.php?&t=9),is a procedure that makes use of the pooled variance $s^2_p$. Black-box approachThe `scipy.stats` function `ttest_ind` will perform all the steps of the $t$-test procedure,without the need for us to understand the details. | from scipy.stats import ttest_ind
# extract data for two groups
xS = df[df["group"]=="S"]['ELV']
xNS = df[df["group"]=="NS"]['ELV']
# run the complete t-test procedure for ind-ependent samples:
result = ttest_ind(xS, xNS)
result.pvalue | _____no_output_____ | MIT | stats_overview/03_STATS.ipynb | minireference/noBSstatsnotebooks |
The $p$-value is less than 0.05 so our decision is to **reject the null hypothesis**. Student's t-test under the hoodThe computations hidden behind the function `ttest_ind` involve a six step procedure that makes use of the pooled variance $s^2_p$. | from statistics import stdev
from scipy.stats.distributions import t
# 1. calculate the mean in each group
meanS, meanNS = np.mean(xS), np.mean(xNS)
# 2. calculate d, the observed difference between means
d = meanS - meanNS
# 3. calculate the standard deviations in each group
stdS, stdNS = stdev(xS), stdev(xNS)
nS, nNS = len(xS), len(xNS)
# 4. compute the pooled variance and standard error
var_pooled = ((nS-1)*stdS**2 + (nNS-1)*stdNS**2)/(nS + nNS - 2)
std_pooled = np.sqrt(var_pooled)
std_err = np.sqrt(std_pooled**2/nS + std_pooled**2/nNS)
# 5. compute the value of the t-statistic
tstat = d / std_err
# 6. obtain the p-value for the t-statistic from a
# t-distribution with 31+30-2 = 59 degrees of freedom
dof = nS + nNS - 2
pvalue = 2 * t(dof).cdf(-abs(tstat)) # 2* because two-sided
pvalue | _____no_output_____ | MIT | stats_overview/03_STATS.ipynb | minireference/noBSstatsnotebooks |
Welch's t-test (unpooled variances)An [alternative t-test procedure](https://statkat.com/stattest.php?&t=9) that doesn't assume the variances in groups are equal. | result2 = ttest_ind(xS, xNS, equal_var=False)
result2.pvalue | _____no_output_____ | MIT | stats_overview/03_STATS.ipynb | minireference/noBSstatsnotebooks |
Welch's $t$-test differs only in steps 4 through 6 as shown below: | # 4'. compute the unpooled standard deviation of D
stdD = np.sqrt(stdS**2/nS + stdNS**2/nNS)
# 5'. compute the value of the t-statistic
tstat = d / stdD
# 6'. obtain the p-value from a t-distribution with
# (insert crazy formula here) degrees of freedom
dof = (stdS**2/nS + stdNS**2/nNS)**2 / \
((stdS**2/nS)**2/(nS-1) + (stdNS**2/nNS)**2/(nNS-1) )
pvalue = 2 * t(dof).cdf(-abs(tstat)) # 2* because two-sided
pvalue | _____no_output_____ | MIT | stats_overview/03_STATS.ipynb | minireference/noBSstatsnotebooks |
Summary of Question 1We saw two ways to answer Question 1 (is there a difference between group means) and obtain the p-value.We interpreted the small p-values as evidence that the observed difference, $d=130$, is unlikely to be due to chance,i.e. we rejected the null hypothesis.Note this whole procedure is just a sanity check—we haven't touched the alternative hypothesis at all yet,and for all we know the stats training could have the effect of decreasing ELV! ____ It's time to study Question 2, which is to estimate the magnitude of the change in ELV obtained from completing the stats training, which is called *effect size* in statistics. Estimating the effect size- Question 2 of statistical analysis is to estimate the difference in ELV gained by stats training.- NEW CONCEPT: **effect size** is a measure of difference between intervention and control groups.- We assume the data of **Group S** and **Group NS** come from different populations with means $\mu_S$ and $\mu_{NS}$- We're interested in the difference between population means, denoted $\Delta = \mu_S - \mu_{NS}$.- By analyzing the sample, we have obtained an estimate $d=130$ for the unknown $\Delta$, but we know our data contains lots of variability, so we know our estimate might be off.- We want an answer to Question 2 (What is the estimated difference between group means?) that takes into account the variability of the data.- NEW CONCEPT: **confidence interval** is a way to describe a range of values for an estimate- We want to provide an answer to Question 2 in the form of a confidence interval that tells us a range of values where we believe the true value of $\Delta$ falls.- Similar to how we showed to approaches for hypothesis testing, we'll work on effect size estimation using two approaches: resampling methods and analytical approximations. Approach 1: estimate the effect size using bootstrap method- We want to estimate the distribution of ELV values for the two groups, and compute the difference between the means of these distributions.- Distributions: - Sampling distributions = obtained by repeated sampling from the populations - Bootstrap sampling distributions = resampling data from the samples we have (with replacement)- Intuition: treat the samples as if they were the population- We'll compute $B=5000$ bootstrap samples from the two groups and compute the difference, then look at the distribution of the bootstrap sample difference to obtain $CI_{\Delta}$, the confidence interval for the difference between population means. | from statistics import mean
def bootstrap_stat(sample, statfunc=mean, B=5000):
"""
Compute the bootstrap estimate of the function `statfunc` from the sample.
Returns a list of statistic values from bootstrap samples.
"""
n = len(sample)
bstats = []
for i in range(0, B):
bsample = np.random.choice(sample, n, replace=True)
bstat = statfunc(bsample)
bstats.append(bstat)
return bstats
# load data for two groups
df = pd.read_csv('data/employee_lifetime_values.csv')
xS = df[df["group"]=="S"]['ELV']
xNS = df[df["group"]=="NS"]['ELV']
# compute bootstrap estimates for mean in each group
meanS_bstats = bootstrap_stat(xS, statfunc=mean)
meanNS_bstats = bootstrap_stat(xNS, statfunc=mean)
# compute the difference between means from bootstrap samples
dmeans_bstats = []
for bmeanS, bmeanNS in zip(meanS_bstats, meanNS_bstats):
d = bmeanS - bmeanNS
dmeans_bstats.append(d)
sns.displot(dmeans_bstats)
# 90% confidence interval for the difference in means
CI_boot = [np.percentile(dmeans_bstats, 5), np.percentile(dmeans_bstats, 95)]
CI_boot | _____no_output_____ | MIT | stats_overview/03_STATS.ipynb | minireference/noBSstatsnotebooks |
SciPy bootstrap method | from scipy.stats import bootstrap
def dmeans2(sample1, sample2):
return np.mean(sample1) - np.mean(sample2)
res = bootstrap((xS, xNS), statistic=dmeans2, vectorized=False,
confidence_level=0.9, n_resamples=5000, method='percentile')
CI_boot2 = [res.confidence_interval.low, res.confidence_interval.high]
CI_boot2 | _____no_output_____ | MIT | stats_overview/03_STATS.ipynb | minireference/noBSstatsnotebooks |
Approach 2: Estimates using analytical approximation method - Assumption 1: populations for **Group S** and **Group NS** are normally distributed - Assumption 2: the variance of the two populations is the same (or approximately equal) - Using the theoretical model for the populations, we can obtain a formula for CI of effect size $\Delta$: $$ \textrm{CI}_{(1-\alpha)} = \left[ d - t^*\!\cdot\!\sigma_D, \, d + t^*\!\cdot\!\sigma_D \right]. $$ The confidence interval is centred at $d$, with width proportional to the standard deviation $\sigma_D$. The constant $t^*$ denotes the value of the inverse CDF of Student's $t$-distribution with appropriate number of degrees of freedom `dof` evaluated at $1-\frac{\alpha}{2}$. For a 90% confidence interval, we choose $\alpha=0.10$, which gives $(1-\frac{\alpha}{2}) = 0.95$, $t^* = F_{T_{\textrm{dof}}}^{-1}\left(0.95\right)$. - We can use the two different analytical approximations to obtain a formula for $\sigma_D$ just as we did in the hypothesis testing: - Pooled variance: $\sigma^2_p = \frac{(n_S-1)s_S^2 + (n_{NS}-1)s_{NS}^2}{n_S + n_{NS} - 2}$, and `dof` = $n_S + n_{NS} -2$ - Unpooled variance: $\sigma^2_u = \tfrac{s^2_A}{n_A} + \tfrac{s^2_B}{n_B}$, and `dof` = [...](https://en.wikipedia.org/wiki/Student%27s_t-testEqual_or_unequal_sample_sizes,_unequal_variances_(sX1_%3E_2sX2_or_sX2_%3E_2sX1)) Using pooled varianceThe calculations are similar to Student's t-test for hypothesis testing. | from scipy.stats.distributions import t
d = np.mean(xS) - np.mean(xNS)
nS, nNS = len(xS), len(xNS)
stdS, stdNS = stdev(xS), stdev(xNS)
var_pooled = ((nS-1)*stdS**2 + (nNS-1)*stdNS**2)/(nS + nNS - 2)
std_pooled = np.sqrt(var_pooled)
std_err = std_pooled * np.sqrt(1/nS + 1/nNS)
dof = nS + nNS - 2
# for 90% confidence interval, need 10% in tails
alpha = 0.10
# now use inverse-CDF of Students t-distribution
tstar = abs(t(dof).ppf(alpha/2))
CI_tpooled = [d - tstar*std_err, d + tstar*std_err]
CI_tpooled | _____no_output_____ | MIT | stats_overview/03_STATS.ipynb | minireference/noBSstatsnotebooks |
Using unpooled varianceThe calculations are similar to the Welch's t-test for hypothesis testing. | d = np.mean(xS) - np.mean(xNS)
nS, nNS = len(xS), len(xNS)
stdS, stdNS = stdev(xS), stdev(xNS)
stdD = np.sqrt(stdS**2/nS + stdNS**2/nNS)
dof = (stdS**2/nS + stdNS**2/nNS)**2 / \
((stdS**2/nS)**2/(nS-1) + (stdNS**2/nNS)**2/(nNS-1) )
# for 90% confidence interval, need 10% in tails
alpha = 0.10
# now use inverse-CDF of Students t-distribution
tstar = abs(t(dof).ppf(alpha/2))
CI_tunpooled = [d - tstar*stdD, d + tstar*stdD]
CI_tunpooled | _____no_output_____ | MIT | stats_overview/03_STATS.ipynb | minireference/noBSstatsnotebooks |
Summary of Question 2 resultsWe now have all the information we need to give a precise and nuanced answer to Question 2: "How big is the increase in ELV produced by stats training?".The basic estimate of the difference is $130$ can be reported, and additionally can can report the 90% confidence interval for the difference between group means, that takes into account the variability in the data we have observed.Note the CIs obtained using different approaches are all similar (+/- 5 ELV points), so it doesn't matter much which approach we use: | CI_boot, CI_boot2, CI_tpooled, CI_tunpooled | _____no_output_____ | MIT | stats_overview/03_STATS.ipynb | minireference/noBSstatsnotebooks |
Standardized effect size (optional)It is sometimes useful to report the effect size using a "standardized" measure for effect sizes.*Cohen's $d$* one such measure, and it is defined as the difference between two means divided by the pooled standard deviation. | def cohend(sample1, sample2):
"""
Compute Cohen's d measure of effect size for two independent samples.
"""
n1, n2 = len(sample1), len(sample2)
mean1, mean2 = np.mean(sample1), np.mean(sample2)
var1, var2 = np.var(sample1, ddof=1), np.var(sample2, ddof=1)
# calculate the pooled variance and standard deviaiton
var_pooled = ((n1-1)*var1 + (n2-1)*var2) / (n1 + n2 - 2)
std_pooled = np.sqrt(var_pooled)
# compute Cohen's d
cohend = (mean1 - mean2) / std_pooled
return cohend
cohend(xS, xNS) | _____no_output_____ | MIT | stats_overview/03_STATS.ipynb | minireference/noBSstatsnotebooks |
We can interpret the value of Cohen's d obtained using the [reference table](https://en.wikipedia.org/wiki/Effect_sizeCohen's_d) of values:| Cohen's d | Effect size || ----------- | ----------- || 0.01 | very small || 0.20 | small || 0.50 | medium || 0.80 | large |We can therefore say the effect size of offering statistics training for employees has an **medium** effect size. Conclusion of Amy's statistical analysisRecall the two research questions that Amy set out to answer in the beginning of this video series:- Question 1: Is there a difference between the means in the two groups?- Question 2: How much does statistics training improve the ELV of employees?The statistical analysis we did allows us to answer these two questions as follows:- Answer 1: There is a statistically significant difference between Group S and Group NS, p = 0.048.- Answer 2: The estimated improvement in ELV is 130 points, which is corresponds to Cohen's d value of 0.52 (medium effect size). A 90% confidence interval for the true effect size is [25.9, 234.2].Note: we used the numerical results obtained from resampling methods (Approach 1), but conclusions would be qualitatively the same if we reported results obtained from analytical approximations (Approach 2). Using statistics for convincing othersYou may be wondering if all this probabilistic modelling and complicated statistical analysis was worth it to reach a conclusion that seems obvious in retrospect. Was all this work worth it? The purpose of all this work is to obtains something close to an objective conclusion. Without statistics it is very easy to fool ourselves and interpret patterns in data the way we want to, or alternatively, not see patterns that are present. By following the standard statistical procedures, we're less likely to fool ourselves, and more likely to be able to convince others.It can very useful to imagine Amy explaining the results to a skeptical colleague. Suppose the colleague is very much against the idea of statistical training, and sees it as a distraction, saying things like "We hire employees to do a job, not to play with Python." and "I don't know any statistics and I'm doing my job just fine!" You get the picture.Imagine Amy presenting her findings about how 100 hours of statistical training improves employee lifetime value (ELV) results after one year, and suggesting the statistical training be implemented for all new hires from now on. The skeptical colleague immediately rejects the idea and questions Amy's recommendation using emotional arguments like about necessity, time wasting, and how statistics is a specialty topic that is not required for all employees. Instead of arguing based on opinions and emotions with her colleague, Amy explains her recommendation is based on a statistical experiment she conducted, and shows the results.- When the colleague asks if the observed difference could be due to chance, Amy says that this is unlikely, and quotes the p-value of 0.048 (less than 0.05), and interprets the result as saying the probability of observed difference between **Group S** and **Group NS** to be due to chance is less than 5%.- The skeptical colleague is forced to concede that statistical training does improve ELV, but then asks about the effect size of the improvement: "How much more ELV can we expect if we provide statistics training?" Amy is ready to answer quoting the observed difference of $130$ ELV points, and further specifies the 90% confidence interval of [25.9, 234.2] for the improvement, meaning in the worst case there is 25 ELV points improvement.The skeptic is forced to back down from their objections, and the "stats training for all" program is adopted in the company. Not only was Amy able to win the argument using statistics, but she was also able to set appropriate expectations for the results. In other words, she hasn't promised a guaranteed +130 ELV improvement, but a realistic range of values that can be expected. Comparison of resampling methods and analytical approximationsIn this notebook we saw two different approaches for doing statistical analysis: resampling methods and analytical approximations. This is a general pattern in statistics where there is not only one correct answer: multiple approaches to data analysis are valid, and you need to think about the specifics of each data analysis situation. You'll learn about both approaches in the book.Analytical approximations currently taught in most stats courses (STAT 101). Historically, analytical approximations have been used more widely because they require only simple arithmetic calculations: statistic practitioners (scientists, engineers, etc.) simply need to compute sample statistics, plug them into a formula, and obtain a $p$-value. This convenience is at the cost of numerous assumptions about the data distribution, which often don't hold in practice (e.g. assuming population is normal, when it is isn't).In recent years, resampling methods like the permutation test and bootstrap estimation are becoming more popular and widely in industry, and increasingly also taught at to university students (*modern statistics*). **The main advantage so resampling methods is that they require less modelling assumptions.** Procedures like the permutation test can be applied broadly to any scenarios where two groups are compared, and don't require developing specific formulas for different cases. Resampling methods are easier to understand since the statistical procedure they require are directly related to the sampling distribution, and there are no formulas to memorize.Understanding resampling methods requires some basic familiarity with programming, but the skills required are not advanced: knowledge of variables, expressions, and basic `for` loop is sufficient. If you were able to follow the code examples described above (see `resample_under_H0`, `permutation_test`, and `bootstrap_stat`), then you've already **seen all the code you will need for the entire book!** Other statistics topics in the bookThe goal of this notebook was to focus on the two main ideas of inferential statistics ([Chapter 3](https://docs.google.com/document/d/1fwep23-95U-w1QMPU31nOvUnUXE2X3s_Dbk5JuLlKAY/editheading=h.uutryzqeo2av)): hypothesis testing and estimation. We didn't have time to cover many of the other important topics in statistics, which will be covered in the book (and in future notebooks). Here is a list of some of these topics:- Null Hypothesis Significance Testing (NHST) procedure in full details (Type I and Type II error, power, sample size calculations) - Statistical assumptions behind analytical approximations- Cookbook of statistical analysis recipes (analytical approximations for different scenarios)- Experimental design (how to plan and conduct statistical experiments)- Misuses of statistics (caveats to watch out for and mistakes to avoid)- Bayesian statistics (very deep topic; we'll cover only main ideas)- Practice problems and exercises (real knowledge is when you can do the calculations yourself) ___ So far our statistical analysis was limited to comparing two groups, which is referred to as **categorical predictor variable** using stats jargon. In the next notebook we'll learn about statistical analysis with **continuous predictor variables**: instead of comparing stats vs. no-stats, we analyze what happens when variable amount of stats training is provided (a continuous predictor variable).Open the notebook [04_LINEAR_MODELS.ipynb](./04_LINEAR_MODELS.ipynb) when you're ready to continue. | code = list(["um"]) | _____no_output_____ | MIT | stats_overview/03_STATS.ipynb | minireference/noBSstatsnotebooks |
Compare built-in Sagemaker classification algorithms for a binary classification problem using Iris dataset In the notebook tutorial, we build 3 classification models using HPO and then compare the AUC on test dataset on 3 deployed modelsIRIS is perhaps the best known database to be found in the pattern recognition literature. Fisher's paper is a classic in the field and is referenced frequently to this day. (See Duda & Hart, for example.) The data set contains 3 classes of 50 instances each, where each class refers to a type of iris plant. The dataset is built-in by default into R or can also be downloaded from https://archive.ics.uci.edu/ml/datasets/irisThe iris dataset, besides its historical importance, is also a fun dataset to play with since it can educate us about various ML techniques such as clustering, classification and regression, all in one dataset.The dataset is built into any base R installation, so no download is required.Attribute Information:1. sepal length in cm2. sepal width in cm3. petal length in cm4. petal width in cm5. Species of flowers: Iris setosa, Iris versicolor, Iris virginicaThe prediction we will perform is `Species ~ f(sepal.length,sepal.width,petal.width,petal.length)`Predicted attribute: Species of iris plant. Load required libraries and initialize variables. | rm(list=ls())
library(reticulate) # be careful not to install reticulate again. since it can cause problems.
library(tidyverse)
library(pROC)
set.seed(1324) | _____no_output_____ | Apache-2.0 | r_examples/r_sagemaker_binary_classification_algorithms/R_binary_classification_algorithms_comparison.ipynb | vllyakho/amazon-sagemaker-examples |
SageMaker needs to be imported using the reticulate library. If this was performed in a local computer, we would have to make sure that Python and appropriate SageMaker libraries are installed, but inside a SageMaker notebook R kernels, these are all pre-loaded and the R user does not have to worry about installing reticulate or Python. Session is the unique session ID associated with each SageMaker call. It remains the same throughout the execution of the program and can be recalled later to close a session or open a new session.The bucket is the Amazon S3 bucket where we will be storing our data output. The Amazon S3 bucket and prefix that you want to use for training and model data. This should be within the same region as the Notebook Instance, training, and hosting.The role is the role of the SageMaker notebook as when it was initially deployed. The IAM role arn used to give training and hosting access to your data. See the documentation for how to create these. Note, if more than one role is required for notebook instances, training, and/or hosting, please replace the boto regexp with appropriate full IAM role arn string(s). | sagemaker <- import('sagemaker')
session <- sagemaker$Session()
bucket <- session$default_bucket() # you may replace with name of your personal S3 bucket
role_arn <- sagemaker$get_execution_role() | _____no_output_____ | Apache-2.0 | r_examples/r_sagemaker_binary_classification_algorithms/R_binary_classification_algorithms_comparison.ipynb | vllyakho/amazon-sagemaker-examples |
Input the data and basic pre-processing | head(iris)
summary(iris) | _____no_output_____ | Apache-2.0 | r_examples/r_sagemaker_binary_classification_algorithms/R_binary_classification_algorithms_comparison.ipynb | vllyakho/amazon-sagemaker-examples |
In above, we see that there are 50 flowers of the setosa species, 50 flowers of the versicolor species, and 50 flowers of the virginica species. In this case, the target variable is the Species prediction. We are trying to predict the species of the flower given its numerical measurements of Sepal length, sepal width, petal length, and petal width. Since we are trying to do binary classification, we will only take the flower species setosa and versicolor for simplicity. Also we will perform one-hot encoding on the categorical variable Species. | iris1 <- iris %>%
dplyr::select(Species,Sepal.Length,Sepal.Width,Petal.Length,Petal.Width) %>% # change order of columns such that the label column is the first column.
dplyr::filter(Species %in% c("setosa","versicolor")) %>% #only select two flower for binary classification.
dplyr::mutate(Species = as.numeric(Species) -1) # one-hot encoding,starting with 0 as setosa and 1 as versicolor.
head(iris1) | _____no_output_____ | Apache-2.0 | r_examples/r_sagemaker_binary_classification_algorithms/R_binary_classification_algorithms_comparison.ipynb | vllyakho/amazon-sagemaker-examples |
We now obtain some basic descriptive statistics of the features. | iris1 %>% group_by(Species) %>% summarize(mean_sepal_length = mean(Sepal.Length),
mean_petal_length = mean(Petal.Length),
mean_sepal_width = mean(Sepal.Width),
mean_petal_width = mean(Petal.Width),
) | _____no_output_____ | Apache-2.0 | r_examples/r_sagemaker_binary_classification_algorithms/R_binary_classification_algorithms_comparison.ipynb | vllyakho/amazon-sagemaker-examples |
In the summary statistics, we observe that mean sepal length is longer than mean petal length for both flowers. Prepare for modelling We split the train and test and validate into 70%, 15%, and 15%, using random sampling. | iris_train <- iris1 %>%
sample_frac(size = 0.7)
iris_test <- anti_join(iris1, iris_train) %>%
sample_frac(size = 0.5)
iris_validate <- anti_join(iris1, iris_train) %>%
anti_join(., iris_test) | _____no_output_____ | Apache-2.0 | r_examples/r_sagemaker_binary_classification_algorithms/R_binary_classification_algorithms_comparison.ipynb | vllyakho/amazon-sagemaker-examples |
We do a check of the summary statistics to make sure train, test, validate datasets are appropriately split and have proper class balance. | table(iris_train$Species)
nrow(iris_train) | _____no_output_____ | Apache-2.0 | r_examples/r_sagemaker_binary_classification_algorithms/R_binary_classification_algorithms_comparison.ipynb | vllyakho/amazon-sagemaker-examples |
We see that the class balance between 0 and 1 is almost 50% each for the binary classification. We also see that there are 70 rows in the train dataset. | table(iris_validate$Species)
nrow(iris_validate) | _____no_output_____ | Apache-2.0 | r_examples/r_sagemaker_binary_classification_algorithms/R_binary_classification_algorithms_comparison.ipynb | vllyakho/amazon-sagemaker-examples |
We see that the class balance in validation dataset between 0 and 1 is almost 50% each for the binary classification. We also see that there are 15 rows in the validation dataset. | table(iris_test$Species)
nrow(iris_test) | _____no_output_____ | Apache-2.0 | r_examples/r_sagemaker_binary_classification_algorithms/R_binary_classification_algorithms_comparison.ipynb | vllyakho/amazon-sagemaker-examples |
We see that the class balance in test dataset between 0 and 1 is almost 50% each for the binary classification. We also see that there are 15 rows in the test dataset. Write the data to Amazon S3 Different algorithms in SageMaker will have different data formats required for training and for testing. These formats are created to make model production easier. csv is the most well known of these formats and has been used here as input in all algorithms to make it consistent.SageMaker algorithms take in data from an Amazon S3 object and output data to an Amazon S3 object, so data has to be stored in Amazon S3 as csv,json, proto-buf or any format that is supported by the algorithm that you are going to use. | write_csv(iris_train, 'iris_train.csv', col_names = FALSE)
write_csv(iris_validate, 'iris_valid.csv', col_names = FALSE)
write_csv(iris_test, 'iris_test.csv', col_names = FALSE)
s3_train <- session$upload_data(path = 'iris_train.csv',
bucket = bucket,
key_prefix = 'data')
s3_valid <- session$upload_data(path = 'iris_valid.csv',
bucket = bucket,
key_prefix = 'data')
s3_test <- session$upload_data(path = 'iris_test.csv',
bucket = bucket,
key_prefix = 'data')
s3_train_input <- sagemaker$inputs$TrainingInput(s3_data = s3_train,
content_type = 'text/csv')
s3_valid_input <- sagemaker$inputs$TrainingInput(s3_data = s3_valid,
content_type = 'text/csv')
s3_test_input <- sagemaker$inputs$TrainingInput(s3_data = s3_test,
content_type = 'text/csv')
| _____no_output_____ | Apache-2.0 | r_examples/r_sagemaker_binary_classification_algorithms/R_binary_classification_algorithms_comparison.ipynb | vllyakho/amazon-sagemaker-examples |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.