repo_name stringlengths 9 75 | topic stringclasses 30
values | issue_number int64 1 203k | title stringlengths 1 976 | body stringlengths 0 254k | state stringclasses 2
values | created_at stringlengths 20 20 | updated_at stringlengths 20 20 | url stringlengths 38 105 | labels listlengths 0 9 | user_login stringlengths 1 39 | comments_count int64 0 452 |
|---|---|---|---|---|---|---|---|---|---|---|---|
kaliiiiiiiiii/Selenium-Driverless | web-scraping | 310 | await driver.close() closes the entire window instead of current Ta | I expected await driver.close() to close only the current tab, but it closes the entire browser window instead.
Steps to Reproduce
Open multiple tabs using execute_script("window.open(...);").
Switch to the new tab.
Call await driver.close().
Instead of closing only the new tab, the entire browser session terminates.
... | open | 2025-02-02T15:07:32Z | 2025-02-02T22:10:13Z | https://github.com/kaliiiiiiiiii/Selenium-Driverless/issues/310 | [
"invalid"
] | bizjaya | 1 |
apachecn/ailearning | nlp | 564 | 几乎所有的图都挂了。。。 | 几乎所有的图都挂了。。。 | closed | 2019-12-23T03:32:12Z | 2019-12-30T02:28:29Z | https://github.com/apachecn/ailearning/issues/564 | [] | llwinner | 2 |
sqlalchemy/alembic | sqlalchemy | 1,098 | Postgres Index with opperators dont autogenerate | **Describe the bug**
I'm using a postgresql index with a inet_ops operator. Autogenerated code does not include the index.
**Expected behavior**
Autogenerated code should include the index.
**To Reproduce**
****env.py****
```py
from logging.config import fileConfig
from sqlalchemy import Column, Index,... | open | 2022-10-12T11:25:37Z | 2022-12-01T16:19:52Z | https://github.com/sqlalchemy/alembic/issues/1098 | [
"autogenerate - detection",
"postgresql",
"use case"
] | garyvdm | 4 |
django-import-export/django-import-export | django | 1,812 | Can't import file without an `id` column in V4 | **Describe the bug**
In V4 I get an error attempting to import data without an `id` column.
**To Reproduce**
When updating to V4 it seems I'm no longer able to import data without an `id` field present in the import data. My expectation (and I think this was the V3 behavior?) is that if `id` is present it is u... | closed | 2024-05-03T05:04:53Z | 2024-05-08T08:10:49Z | https://github.com/django-import-export/django-import-export/issues/1812 | [
"bug"
] | cdubz | 4 |
desec-io/desec-stack | rest-api | 752 | Webapp: refactor form inputs | ## Problem
Some form inputs are C&P and code is redundant.
Affected inputs:
- [x] E-Mail
- [x] Password
- [ ] Captcha
## Solution
Refactor to generic component.
Implementation already done, but must be evaluated. PR in some days... | open | 2023-07-06T07:08:56Z | 2023-09-16T11:43:37Z | https://github.com/desec-io/desec-stack/issues/752 | [] | Rotzbua | 1 |
open-mmlab/mmdetection | pytorch | 12,020 | ImportError: cannot import name 'make_res_layer' from 'mmdet.models.backbones' | >>> import mmdet
>>> mmdet.__version__
'1.0.rc0+34e9957' | open | 2024-10-29T12:08:51Z | 2024-10-29T12:09:07Z | https://github.com/open-mmlab/mmdetection/issues/12020 | [] | jiyuwangbupt | 0 |
CorentinJ/Real-Time-Voice-Cloning | python | 1,189 | Support for Arabic language. | Hello, I'm working on an Arabic model for this project. I'm planning on adding Arabic characters support as well as having the software synthesize audio that speaks Arabic. So far I'm trying to do the following :
1. Downloading the corpus Arabic dataset linked here : http://en.arabicspeechcorpus.com/arabic-speech-co... | open | 2023-04-12T11:48:25Z | 2023-04-12T11:48:25Z | https://github.com/CorentinJ/Real-Time-Voice-Cloning/issues/1189 | [] | ghost | 0 |
ultralytics/yolov5 | pytorch | 13,330 | about save-txt in yolov5-seg | ### Search before asking
- [X] I have searched the YOLOv5 [issues](https://github.com/ultralytics/yolov5/issues) and [discussions](https://github.com/ultralytics/yolov5/discussions) and found no similar questions.
### Question
when activating "save-txt" in yolov5-seg.py, a txt with the coordinates of the pre... | open | 2024-09-23T09:40:49Z | 2024-10-27T13:30:36Z | https://github.com/ultralytics/yolov5/issues/13330 | [
"question"
] | Powerfulidot | 5 |
Yorko/mlcourse.ai | data-science | 757 | Proofread topic 6 | - Fix issues
- Fix typos
- Correct the translation where needed
- Add images where necessary | closed | 2023-10-24T07:41:40Z | 2024-08-25T08:08:32Z | https://github.com/Yorko/mlcourse.ai/issues/757 | [
"enhancement",
"articles"
] | Yorko | 0 |
Anjok07/ultimatevocalremovergui | pytorch | 1,496 | Low VRAM error | I'm using the openCL version with a 1GB VRAM Radeon RX580, so definitely on the lowers of the low VRAM range. I was trying to use the Ensemble Mode, but it didn't run. I have 16gb of RAM though, so it would be lovely if it could use my RAM instead, but that's a PyTorch problem, isn't it? Could you guys create a low_vr... | open | 2024-08-06T11:09:11Z | 2024-08-06T11:31:05Z | https://github.com/Anjok07/ultimatevocalremovergui/issues/1496 | [] | RutraNickers | 1 |
alpacahq/alpaca-trade-api-python | rest-api | 276 | paper trading api keys don't authenticate on StreamConn | I'm using my paper trading keys to instantiate a StreamConn because I want to receive trade updates on my paper account. But it looks like StreamConn only authenticates on live trading keys. | closed | 2020-07-24T03:00:14Z | 2020-07-24T03:34:28Z | https://github.com/alpacahq/alpaca-trade-api-python/issues/276 | [] | connor-roche | 0 |
ultralytics/yolov5 | deep-learning | 12,815 | load images in batch size | ### Search before asking
- [X] I have searched the YOLOv5 [issues](https://github.com/ultralytics/yolov5/issues) and [discussions](https://github.com/ultralytics/yolov5/discussions) and found no similar questions.
### Question
Hi, I would like to ask when using torch.hub.load to load a model, is there a method to p... | closed | 2024-03-12T01:59:59Z | 2024-05-09T00:21:09Z | https://github.com/ultralytics/yolov5/issues/12815 | [
"question",
"Stale"
] | KnightInsight | 6 |
Miserlou/Zappa | flask | 2,233 | ImportError: cannot import name 'Flask' from 'flask' (unknown location) | I am trying to deploy a simple application using zappa which uses flask and moviepy.
## Context
Neither the name of the folder nor the any file is named as flask.
Locally `python ./app.py` is just working fine.
I am running on python 3.9
## Expected Behavior
To run normally on lambda.
## Actual Behavior
Whe... | open | 2021-12-12T13:24:19Z | 2022-08-22T13:27:45Z | https://github.com/Miserlou/Zappa/issues/2233 | [] | Shubham-Kumar-2000 | 2 |
allenai/allennlp | nlp | 5,018 | Visual Genome Question Answering | Visual Genome has a set of questions for every image, 1.7M in total. This task is about `Step` that produces a `DatasetDict` for this task, and then some more steps to connect it to the model for [VQA](https://github.com/allenai/allennlp-models/blob/main/allennlp_models/vision/dataset_readers/vqav2.py) and [GQA](https:... | open | 2021-02-24T22:40:24Z | 2021-08-28T00:25:23Z | https://github.com/allenai/allennlp/issues/5018 | [
"Contributions welcome",
"Models",
"medium"
] | dirkgr | 0 |
flairNLP/flair | nlp | 2,953 | How to use the trained model for named entity recognition | I've trained a named entity-related model, now how can I use the model to recognize named entities in my own sentences? Can you provide relevant examples? I looked at this case
`model = SequenceTagger.load('resources/taggers/example-upos/final-model.pt')
sentence = Sentence('I love Berlin')
model.predict(sentence)
... | closed | 2022-10-01T03:10:17Z | 2023-04-02T16:54:25Z | https://github.com/flairNLP/flair/issues/2953 | [
"question",
"wontfix"
] | yaoysyao | 1 |
jina-ai/serve | deep-learning | 6,168 | Engine consumes a large amount of Memory. What methods can be used to optimize memory usage? | jina ver: 3.25.1 latest
When using an empty example to observe the idle memory usage without adding any executor, the situation is as follows:
flow.py
```python
from jina import Flow
f = Flow()
with f:
f.block()
```
In the x86 environment:
```scss
python(16985)───python(17051)─┬─{default-execut... | closed | 2024-05-16T07:00:52Z | 2024-08-29T00:21:17Z | https://github.com/jina-ai/serve/issues/6168 | [
"Stale"
] | Janus-Xu | 8 |
ansible/ansible | python | 83,915 | local_action does not delegate to controller when remote is localhost | ### Summary
When port forwarding a remote SSH server to a local port and using that local port in Ansible, delegation to the controller node does not seem to be possible with either the `local_action` or `delegate_to` as, in both cases, Ansible runs delegated tasks on the remote node.
While it does make sense to me t... | closed | 2024-09-07T00:30:35Z | 2024-09-21T13:00:02Z | https://github.com/ansible/ansible/issues/83915 | [
"bug",
"affects_2.17"
] | Jackenmen | 5 |
vitalik/django-ninja | pydantic | 977 | [BUG] pagination does not support async view | **Describe the bug**
I created async APIs with ninja pagination. error occurs as below.
The qs passed into pagination class is coroutine.

**Versions (please complete the following information):**
- Python vers... | closed | 2023-12-04T07:01:13Z | 2023-12-04T08:39:03Z | https://github.com/vitalik/django-ninja/issues/977 | [] | samuelchen | 1 |
numpy/numpy | numpy | 27,679 | BUG: Inconsistent casting error for masked-arrays in-place operations | ### Describe the issue:
The changes in scalar casting of 2.0 seem to affect NumPy arrays and masked-arrays differently for in-place operations, resulting in unexpected errors for masked-arrays. See example below (which only occurs in NumPy 2.X, but not on 1.26.4).
Maybe linked to https://github.com/numpy/numpy/issu... | open | 2024-10-30T23:00:40Z | 2024-11-01T18:31:09Z | https://github.com/numpy/numpy/issues/27679 | [
"00 - Bug",
"component: numpy.ma"
] | rhugonnet | 3 |
gradio-app/gradio | deep-learning | 9,992 | Update the hugging face space header | - [x] I have searched to see if a similar issue already exists.
We need to update the Huggingface space header version to latest to pull in some bug fixes. | open | 2024-11-19T03:49:34Z | 2024-11-25T23:32:04Z | https://github.com/gradio-app/gradio/issues/9992 | [
"good first issue",
"svelte"
] | pngwn | 1 |
tortoise/tortoise-orm | asyncio | 1,109 | HOW to use 'using_db' in 'in_transaction' when using 'bulk_create' | tortoise-orm==0.18.1
when I use 'in_transaction', I want to use 'MyModel.bulk_create', but has no attribute 'using_db'
```
async with in_transaction('default') as tconn:
objects = [
MyModel(...) for _ in range(10)
]
await MyModel.bulk_create(objects) ## this has no attribute 'using_db', ho... | closed | 2022-04-21T08:16:32Z | 2022-04-21T08:36:31Z | https://github.com/tortoise/tortoise-orm/issues/1109 | [] | wchpeng | 3 |
ymcui/Chinese-LLaMA-Alpaca | nlp | 440 | 预训练后效果不佳,请教过程问题 | 我训练的过程如下:
1、下载chinese_llama_plus_lora_7b、chinese_alpaca_plus_lora_7b,并进行合并,脚本如下:
python scripts/merge_llama_with_chinese_lora.py \
--base_model /home/dell/project/Chinese-LLaMA-Alpaca/models/llama_hf_models \
--lora_model /home/dell/project/Chinese-LLaMA-Alpaca/models/chinese_llama_plus_lora_7b,/home/dell/p... | closed | 2023-05-27T05:34:13Z | 2023-06-19T22:02:44Z | https://github.com/ymcui/Chinese-LLaMA-Alpaca/issues/440 | [
"stale"
] | chyyf006 | 30 |
benbusby/whoogle-search | flask | 416 | [FEATURE] <Autocomplete URL for browsers> | Since whoogle has autocomplete functionality, what about adding an autocomplete URL for the browsers that support it?
`https://<instance>/auto?q=` something like this?
This is vivaldi and you can add a "suggest URL thing"
.
Below is the full sc... | open | 2024-12-11T10:08:14Z | 2024-12-19T17:17:56Z | https://github.com/deepspeedai/DeepSpeed/issues/6853 | [
"bug",
"compression"
] | cyx96 | 3 |
NVIDIA/pix2pixHD | computer-vision | 64 | Is it possible to freeze the model in pytorch ? | Thanks to this code that I succeed to train with my custom data. As I set `--ngf=64`, at last I obtained a model .pth very heavy: about **700MB.**
I would like to ask if there exist some methods to freeze the model (before I worked with tensorflow so you can freeze your ckpt to .pb so at last you get a much lighter m... | open | 2018-09-26T13:17:06Z | 2018-09-26T13:17:06Z | https://github.com/NVIDIA/pix2pixHD/issues/64 | [] | chenyuZha | 0 |
ets-labs/python-dependency-injector | flask | 821 | a simplest fastapi app, di does not work | I've been playing with the lib for a while. Cannot make it work :(
Python 3.12.3
dependency-injector = "^4.42.0"
fastapi = "^0.110.1"
uvicorn = "^0.29.0"
```
import uvicorn
from dependency_injector import containers, providers
from dependency_injector.wiring import Provide, inject
from fastapi import FastA... | closed | 2024-10-02T20:50:24Z | 2024-10-03T08:30:55Z | https://github.com/ets-labs/python-dependency-injector/issues/821 | [] | antonio-antuan | 1 |
neuml/txtai | nlp | 626 | QUESTION. Debugging, tracing, observability? | Hey, i like how minimalistic txtai is, a way better to get started with then competitors.
Thinking about debbuging, i don't find txtai's solution such as [langchain debugging](https://python.langchain.com/docs/guides/debugging) or [llamaindex tracing](https://docs.llamaindex.ai/en/stable/understanding/tracing_and_de... | closed | 2023-12-31T23:46:09Z | 2024-11-19T12:49:55Z | https://github.com/neuml/txtai/issues/626 | [] | 4l1fe | 5 |
onnx/onnx | machine-learning | 6,583 | ImportError when importing onnx in python script packed in snap on arm machine | # Bug Report
### Is the issue related to model conversion?
No
### Describe the bug
There is an ImportError when trying to import onnx into a python program when the python file is packed into a snap package and run on an arm device. The error is: `ImportError: cannot import name 'ONNX_ML' from 'onnx.onnx_cpp2p... | open | 2024-12-13T17:21:28Z | 2025-02-19T17:32:44Z | https://github.com/onnx/onnx/issues/6583 | [
"bug"
] | BjGoCraft | 0 |
chatanywhere/GPT_API_free | api | 286 | zotero翻译插件和iTerm均出现405报错 | **Describe the bug 描述bug**
确定配置没有问题,zotero- translator和iTerm出现405报错,沉浸式翻译可以正常使用,.tech和.cn都出现此问题
**Screenshots 截图**
<img width="467" alt="image" src="https://github.com/user-attachments/assets/49d4df98-e63b-4fad-9b10-209dfe1875a0">
<img width="248" alt="image" src="https://github.com/user-attachments/assets/fae9... | closed | 2024-08-24T02:56:21Z | 2024-08-26T05:50:40Z | https://github.com/chatanywhere/GPT_API_free/issues/286 | [] | JYDAG | 5 |
python-gino/gino | asyncio | 26 | Table inheritance? | open | 2017-08-04T05:48:11Z | 2018-05-21T07:15:24Z | https://github.com/python-gino/gino/issues/26 | [
"help wanted",
"question"
] | fantix | 2 | |
pallets-eco/flask-sqlalchemy | flask | 820 | README example throws error | If I try to run the example from the repository README, it throws this error:
> python3 app.py/home/username/.local/lib/python3.8/site-packages/flask_sqlalchemy/__init__.py:834: FSADeprecationWarning: SQLALCHEMY_TRACK_MODIFICATIONS adds significant overhead and will be disabled by default in the future. Set it to T... | closed | 2020-05-15T20:37:49Z | 2020-12-05T19:58:35Z | https://github.com/pallets-eco/flask-sqlalchemy/issues/820 | [] | rafaellehmkuhl | 2 |
CorentinJ/Real-Time-Voice-Cloning | pytorch | 1,062 | synthesizer params varries for same input audio and text?? | If run a the demo.cli.py for same audio and text multiple time i can see variation in synthesizer
EX-Synthesizing the waveform:
{| ████████████████ 57000/57600 | Batch Size: 6 | Gen Rate: 5.1kHz | }float64
Synthesizing the waveform:
{| ████████████████ 47500/48000 | Batch Size: 5 | Gen Rate: 4.3kHz | }float64
... | open | 2022-05-05T12:15:12Z | 2022-05-25T20:19:15Z | https://github.com/CorentinJ/Real-Time-Voice-Cloning/issues/1062 | [] | ayush431 | 8 |
huggingface/datasets | pytorch | 6,801 | got fileNotFound | ### Describe the bug
When I use load_dataset to load the nyanko7/danbooru2023 data set, the cache is read in the form of a symlink. There may be a problem with the arrow_dataset initialization process and I get FileNotFoundError: [Errno 2] No such file or directory: '2945000.jpg'
### Steps to reproduce the bug
#code... | closed | 2024-04-11T04:57:41Z | 2024-04-12T16:47:43Z | https://github.com/huggingface/datasets/issues/6801 | [] | laoniandisko | 2 |
recommenders-team/recommenders | data-science | 1,929 | [BUG] News recommendation method: npa_MIND.ipynb cannot run properly! | ### Description
I'm running `recommenders/examples/00_quick_start/npa_MIND.ipynb`, and I encountered the following error message when I ran `print(model.run_eval(valid_news_file, valid_behaviors_file))`:
> File ~/anaconda3/envs/tf2/lib/python3.8/site-packages/tensorflow/python/client/session.py:1480, in BaseSessio... | closed | 2023-05-08T08:06:59Z | 2023-05-13T09:02:39Z | https://github.com/recommenders-team/recommenders/issues/1929 | [
"bug"
] | SnowyMeteor | 1 |
okken/pytest-check | pytest | 125 | Using pytest_check asserts that there will be errors | My error is shown below
```shell
../venv/lib/python3.11/site-packages/_pytest/config/__init__.py:1173
/Users/shanyingqing/code/weeeTest/venv/lib/python3.11/site-packages/_pytest/config/__init__.py:1173: PytestAssertRewriteWarning: Module already imported so cannot be rewritten: pytest_check
self._mark_plu... | closed | 2023-04-04T08:35:40Z | 2023-06-06T21:03:58Z | https://github.com/okken/pytest-check/issues/125 | [
"need more info"
] | Yingqingshan | 2 |
Gozargah/Marzban | api | 1,141 | raw client config template! | as client configs are so complicated, some users needs a lot of things like External Config (https://github.com/Gozargah/Marzban/pull/1118) & fake config (to show user expire time and else) without creating additional inbound & many options like tcpFastOpen that is not usable with Marzban currently
I have a idea to so... | closed | 2024-07-19T10:07:16Z | 2024-07-20T21:07:53Z | https://github.com/Gozargah/Marzban/issues/1141 | [
"Feature"
] | fodhelper | 1 |
NullArray/AutoSploit | automation | 1,229 | Unhandled Exception (09a4fcccc) | Autosploit version: `2.2.3`
OS information: `Linux-3.18.130-dieD-Rebase-armv8l-with-libc`
Running context: `/data/data/com.thecrackertechnology.andrax/ANDRAX/AutoSploit/autosploit.py`
Error meesage: `[Errno 2] No such file or directory: 'host'`
Error traceback:
```
Traceback (most recent call):
File "/data/data/com.th... | closed | 2019-12-31T10:53:45Z | 2020-02-02T01:20:00Z | https://github.com/NullArray/AutoSploit/issues/1229 | [] | AutosploitReporter | 0 |
deezer/spleeter | tensorflow | 667 | [Bug] Windows CLI: 2.3.0 crashes when working in GPU mode with RTX3060 laptop 6GB VRAM | - [Y] I didn't find a similar issue already open.
- [Y] I read the documentation (README AND Wiki)
- [Y] I have installed FFMpeg
- [Y] My problem is related to Spleeter only, not a derivative product (such as Webapplication, or GUI provided by others)
## Description
I use spleeter 2.3.0 in
![微信图片_202109261858... | open | 2021-09-26T10:58:56Z | 2021-12-21T10:29:52Z | https://github.com/deezer/spleeter/issues/667 | [
"bug",
"invalid"
] | ths0013 | 3 |
ipyflow/ipyflow | jupyter | 116 | add observable-style minimap to display cell dependencies | open | 2022-12-21T19:25:23Z | 2023-08-13T04:32:18Z | https://github.com/ipyflow/ipyflow/issues/116 | [
"enhancement"
] | smacke | 1 | |
CorentinJ/Real-Time-Voice-Cloning | pytorch | 1,218 | demo_cli melspectrogram error | I get this error when running demo_cli.py:
`
File "/home/carlos/Downloads/Real-Time-Voice-Cloning-master/encoder/audio.py", line 58, in wav_to_mel_spectrogram
frames = librosa.feature.melspectrogram(
TypeError: melspectrogram() takes 0 positional arguments but 2 positional arguments (and 2 keyword-only ar... | open | 2023-05-14T19:40:25Z | 2023-05-17T07:20:26Z | https://github.com/CorentinJ/Real-Time-Voice-Cloning/issues/1218 | [] | Noobos100 | 1 |
pydata/pandas-datareader | pandas | 931 | RemoteDataError | Yesteday I found this error.
I suppose yahoo finace has changed html structure.
```
[/usr/local/lib/python3.7/dist-packages/pandas_datareader/base.py](https://localhost:8080/#) in _get_response(self, url, params, headers)
179 msg += "\nResponse Text:\n{0}".format(last_response_text)
180
-->... | open | 2022-04-24T23:08:59Z | 2022-04-24T23:08:59Z | https://github.com/pydata/pandas-datareader/issues/931 | [] | higebobo | 0 |
roboflow/supervision | pytorch | 748 | [JSONSink] - allowing to serialise Detections to a JSON file | ### Description
The `JSONSink` class should be designed to efficiently convert and store detection data into a JSON file format. This class should be capable of handling bounding box coordinates in the `xyxy` format and converting them into a JSON structure with fields: `x_min`, `y_min`, `x_max`, `y_max`. It should ... | closed | 2024-01-18T20:25:20Z | 2024-03-01T00:23:47Z | https://github.com/roboflow/supervision/issues/748 | [
"enhancement",
"Q1.2024"
] | SkalskiP | 0 |
adbar/trafilatura | web-scraping | 42 | Replace requests with bare urllib3 | The current use of `requests` sessions in `cli_utils.py` doesn't appear to be thread-safe (https://github.com/psf/requests/issues/2766).
The full functionality of the module isn't really needed here and a change would help reducing the total number of dependencies as mentioned in https://github.com/adbar/trafilatura... | closed | 2020-12-15T16:19:35Z | 2020-12-16T15:45:29Z | https://github.com/adbar/trafilatura/issues/42 | [
"enhancement"
] | adbar | 1 |
lucidrains/vit-pytorch | computer-vision | 130 | Transformer in Convolutional Neural Networks [Paper Implementation] | Hi @lucidrains
Are you considering implementing the titled "Transformer in Convolutional Neural Networks" paper in your repo?
https://arxiv.org/pdf/2106.03180.pdf
Regards,
Khawar | open | 2021-07-04T07:18:25Z | 2021-07-04T07:18:25Z | https://github.com/lucidrains/vit-pytorch/issues/130 | [] | khawar-islam | 0 |
huggingface/datasets | pytorch | 6,796 | CI is broken due to hf-internal-testing/dataset_with_script | CI is broken for test_load_dataset_distributed_with_script. See: https://github.com/huggingface/datasets/actions/runs/8614926216/job/23609378127
```
FAILED tests/test_load.py::test_load_dataset_distributed_with_script[None] - assert False
+ where False = all(<generator object test_load_dataset_distributed_with_scr... | closed | 2024-04-10T06:56:02Z | 2024-04-12T09:02:13Z | https://github.com/huggingface/datasets/issues/6796 | [
"bug"
] | albertvillanova | 4 |
JaidedAI/EasyOCR | pytorch | 579 | Failed to detect and determine text with confidence. | 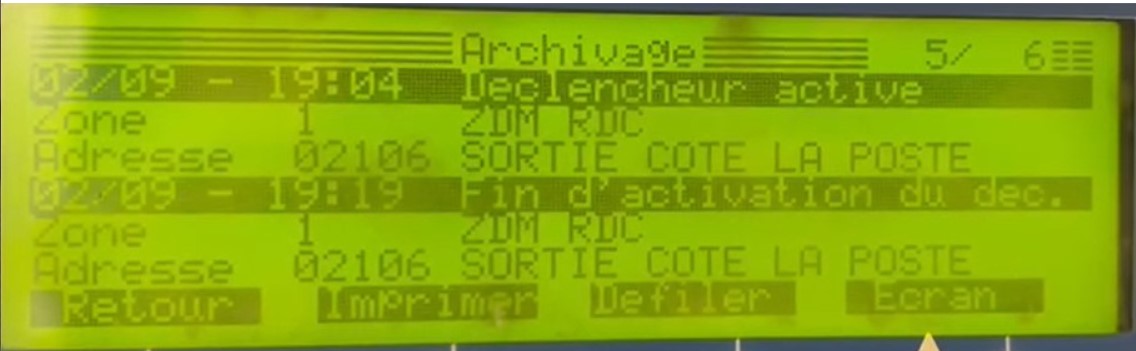
I have a well uniform writting on a led screen.
2 problematics:
- evolving light
- black and green background changing row by row
With an adaptive treshold and by analyzing the row one by one, w... | closed | 2021-10-28T15:01:51Z | 2021-12-15T16:30:05Z | https://github.com/JaidedAI/EasyOCR/issues/579 | [] | AvekIA | 2 |
oegedijk/explainerdashboard | plotly | 212 | get_feature_names_out() not working with FunctionTransformer | Sheers,
if you use a FunctionTransformer in a columntransformer explainerdashboard does not recognize the column names.
I think this is because you use get_features_names_out() in your code rather than the attribute. | closed | 2022-04-27T12:41:46Z | 2022-04-27T22:03:37Z | https://github.com/oegedijk/explainerdashboard/issues/212 | [] | nilslacroix | 1 |
Lightning-AI/pytorch-lightning | machine-learning | 20,348 | Add support S3 as a storage option for profiling results | ### Description & Motivation
Description: Currently, when using the <code>default_root_dir</code> parameter to specify an S3 bucket as the storage location for profiling results, the profiler fails to work due to lack of support for S3 I/O. This limitation prevents users from storing profiling results in a scalable an... | open | 2024-10-18T15:23:04Z | 2024-10-18T15:23:27Z | https://github.com/Lightning-AI/pytorch-lightning/issues/20348 | [
"feature",
"needs triage"
] | kimminw00 | 0 |
custom-components/pyscript | jupyter | 439 | Strange behaviour of @event_trigger | Hi,
I just found some strange behaviour when using the event_trigger.
Having somehting like this:
`
@event_trigger("some_event", "action == 'some_action' and state=='finished'")
def dummy(**kwargs):
log.debug(f"{kwargs}")
`
a and triggering the event with `event.fire("some_event")` leads to the cor... | closed | 2023-02-23T10:57:29Z | 2023-04-08T21:17:12Z | https://github.com/custom-components/pyscript/issues/439 | [] | Michael-CGN | 3 |
deepfakes/faceswap | deep-learning | 1,085 | Being informed on manual preview refresh | On slower hardware and with demanding model configurations it can take several minutes until a manual preview refresh actually completes.
For that reason I suggest that another message "Refresh preview done" will be added, so that the user can focus on other things in the meantime and still reliably tell whether the... | closed | 2020-11-10T10:37:25Z | 2021-05-30T10:48:41Z | https://github.com/deepfakes/faceswap/issues/1085 | [
"feature"
] | OreSeq | 1 |
SYSTRAN/faster-whisper | deep-learning | 941 | can't get it to use GPU | so I've been trying to use whisperX but can't get it to work, so decided to test with faster whisper since it is built on that.
but I still can't get it working, it only uses CPU.
I've also downloaded the 11 and 12 files from here and pasted them in the bin https://github.com/Purfview/whisper-standalone-win/releases/... | open | 2024-07-29T12:32:01Z | 2024-08-13T08:46:00Z | https://github.com/SYSTRAN/faster-whisper/issues/941 | [] | M2ATrail | 4 |
yunjey/pytorch-tutorial | deep-learning | 72 | RuntimeError: invalid argument 2: out of range | HI, @jtoy @hunkim @Kongsea @DingKe @JayParks
I met this error in image caption run:
kraken@devBox1:~/pytorch-tutorial/tutorials/03-advanced/image_captioning$ sudo python3 sample.py --image='./png/example.png'
[sudo] password for kraken:
Traceback (most recent call last):
File "sample.py", line 97, in <mod... | closed | 2017-10-13T07:34:32Z | 2018-05-10T08:58:41Z | https://github.com/yunjey/pytorch-tutorial/issues/72 | [] | bemoregt | 2 |
huggingface/datasets | nlp | 6,585 | losing DatasetInfo in Dataset.map when num_proc > 1 | ### Describe the bug
Hello and thanks for developing this package!
When I process a Dataset with the map function using multiple processors some set attributes of the DatasetInfo get lost and are None in the resulting Dataset.
### Steps to reproduce the bug
```python
from datasets import Dataset, DatasetInfo... | open | 2024-01-12T13:39:19Z | 2024-01-12T14:08:24Z | https://github.com/huggingface/datasets/issues/6585 | [] | JochenSiegWork | 2 |
encode/httpx | asyncio | 3,203 | Authentification docs reference non-existing class | The [auth docs](https://www.python-httpx.org/advanced/authentication/) reference `httpx.BasicAuthentication()` 3 times, but this class does not exist, it should be replaced with `httpx.BasicAuth()`. | closed | 2024-05-17T07:53:03Z | 2024-08-27T12:10:34Z | https://github.com/encode/httpx/issues/3203 | [] | alexprengere | 2 |
vastsa/FileCodeBox | fastapi | 54 | docker起来之后怎么开启HTTPS,直接反向代理? 使用CF的CDN提示525错误 | 不知道从哪里开始排查错误/ | closed | 2023-02-28T06:27:37Z | 2024-04-29T15:08:36Z | https://github.com/vastsa/FileCodeBox/issues/54 | [] | asseywang | 2 |
quantmind/pulsar | asyncio | 126 | WsgiHandler doesn't chain response middlewares | According to the documentation the `response_middleware` parameter in the `WsgiHandler` supports a list of functions:
``` python
middleware = [...]
response_middleware = [
CustomMiddleware(),
GZipMiddleware()
]
return wsgi.WsgiHandler(middleware=middleware, response_middleware=response_middleware)
```
However... | closed | 2014-08-12T13:55:24Z | 2014-10-14T13:51:09Z | https://github.com/quantmind/pulsar/issues/126 | [
"wsgi"
] | sekrause | 11 |
piskvorky/gensim | machine-learning | 3,051 | Release 3.8.3: vector_size in docs doesn't match release constructor argument for word2vec and fasttext | A fix seems to be currently in the develop branch, but the pypi release branch (release-3.8.3) [Word2Vec](https://github.com/RaRe-Technologies/gensim/blob/release-3.8.3/gensim/models/word2vec.py#L477) and [FastText](https://github.com/RaRe-Technologies/gensim/blob/release-3.8.3/gensim/models/fasttext.py#L355) files use... | closed | 2021-02-25T22:25:57Z | 2021-02-26T07:47:47Z | https://github.com/piskvorky/gensim/issues/3051 | [] | hakunanatasha | 1 |
ultralytics/yolov5 | deep-learning | 13,246 | divide the objects into small and large categories based on the size of the bonding boxes | ### Search before asking
- [X] I have searched the YOLOv5 [issues](https://github.com/ultralytics/yolov5/issues) and [discussions](https://github.com/ultralytics/yolov5/discussions) and found no similar questions.
### Question
Hello. I had a question that was not similar. I want to divide the objects into small and... | open | 2024-08-06T10:53:11Z | 2024-10-20T19:51:27Z | https://github.com/ultralytics/yolov5/issues/13246 | [
"question"
] | EmmaLevine94 | 8 |
PaddlePaddle/PaddleHub | nlp | 2,144 | PaddleHub一键OCR中文识别-五、部署服务器 | Sir, thanks for your guidance. I have completed online image recognition according the article
https://aistudio.baidu.com/aistudio/projectdetail/5121515
While, I am unable to know how to deploy the server through the notebook.I will appreciate it ,if you give me detail steps about deploying the server -"五、部署服务器" by t... | open | 2022-11-28T05:34:47Z | 2022-12-01T02:36:20Z | https://github.com/PaddlePaddle/PaddleHub/issues/2144 | [] | MaxokDavid | 9 |
prkumar/uplink | rest-api | 61 | Code suggestion: use abc instead of raising NotImplementedError in interfaces | ``abc`` module provides the infrastructure for defining abstract base classes (ABCs) in Python, as outlined in PEP 3119. The point is that not fully defined successors of abstract base class cannot be instantiated unlike objects with methods raising ``NotImplementedError``. | open | 2018-02-01T13:32:23Z | 2018-02-01T19:08:52Z | https://github.com/prkumar/uplink/issues/61 | [
"Needs Maintainer Input"
] | daa | 0 |
aleju/imgaug | deep-learning | 90 | Make the bounding box more tighten for the object after image rotation | I use this code for image and bounding box rotation:
```
ia.seed(1)
image = # read image
bbs = ia.BoundingBoxesOnImage(
[ia.BoundingBox(x1=95, y1=52, x2=250, y2=245)],
shape=image.shape)
seq = iaa.Sequential([
iaa.Multiply((1.2, 1.5)), # change brightness, doesn't af... | closed | 2018-01-10T13:42:46Z | 2020-11-05T17:42:34Z | https://github.com/aleju/imgaug/issues/90 | [] | panovr | 5 |
noirbizarre/flask-restplus | api | 438 | Refactor for accepting Marshmallow and other marshaling libraries | This is less of an issue and more of an **intent to implement** :shipit:
## Result
The result of this work should be these two main things:
- It should be possible to build alternative request parsing implementations.
- It should be possible to use Marshmallow as easily as the current request parsing impleme... | closed | 2018-05-15T17:40:18Z | 2019-12-30T10:26:45Z | https://github.com/noirbizarre/flask-restplus/issues/438 | [] | martijnarts | 24 |
Gerapy/Gerapy | django | 233 | 通过克隆的方法部署项目,可以运行起来,但是登录不了 | **描述错误**
通过克隆的方法部署项目,可以运行,但是登录不了
**重现**
重现行为的步骤:
1. 通过 git clone 克隆仓库到本地
2. 安装好 requirements.txt 所需库,在 Pycharm 里面配置好 `*\Gerapy\gerapy\cmd\__init__.py `运行配置
3. 启动 scrapyd,运行 `*\Gerapy\gerapy\cmd\__init__.py `文件,在 gerapy/client 文件夹下 通过 `npm run serve` 命令启动前端
4. 控制台报错
**Traceback**
将控制台中显示的 traceback 复制到此处:
... | closed | 2022-04-02T02:47:05Z | 2022-04-29T19:40:37Z | https://github.com/Gerapy/Gerapy/issues/233 | [
"bug"
] | kesyupeng | 2 |
deepspeedai/DeepSpeed | deep-learning | 5,793 | [BUG] Excessive CPU and GPU Memory Usage with Multi-GPU Inference Using DeepSpeed | I am experiencing excessive CPU and GPU memory usage when running multi-GPU inference with DeepSpeed. Specifically, the memory usage does not scale as expected when increasing the number of GPUs. Below is the code I am using for inference:
```python
import os
import torch
import deepspeed
import time
from transfo... | open | 2024-07-23T07:55:48Z | 2024-10-10T03:02:31Z | https://github.com/deepspeedai/DeepSpeed/issues/5793 | [
"bug",
"inference"
] | gawain000000 | 3 |
JaidedAI/EasyOCR | machine-learning | 775 | Tips and trick | Hi,
could you share some tip and tirck about creating data and config file
for train arabic model
thanks | closed | 2022-07-04T16:08:56Z | 2022-11-18T11:00:19Z | https://github.com/JaidedAI/EasyOCR/issues/775 | [] | uniquefan | 4 |
ndleah/python-mini-project | data-visualization | 270 | Add Docstring in the Caesar Cipher file | # Description
In this issue, I want to add docstrings to the function in the Caesar Cipher code. The purpose is to have a clear and well documented code that will help us to understand it better.
<!-- Please include a summary of the issue.-->
## Type of issue
- [ ] Feature (New Script)
- [ ] Bug
- [x] Documen... | open | 2024-06-07T08:33:33Z | 2024-06-09T08:20:29Z | https://github.com/ndleah/python-mini-project/issues/270 | [] | Gabriela20103967 | 0 |
eamigo86/graphene-django-extras | graphql | 152 | DurationField is incorrectly converted as float | Hi!
Doing [this](https://github.com/eamigo86/graphene-django-extras/blob/master/graphene_django_extras/converter.py#L264):
```python
@convert_django_field.register(models.DurationField)
def convert_field_to_float(field, registry=None, input_flag=None, nested_field=False):
return Float(
description... | open | 2020-09-18T12:42:00Z | 2020-09-18T12:42:32Z | https://github.com/eamigo86/graphene-django-extras/issues/152 | [] | karlosss | 0 |
tflearn/tflearn | tensorflow | 1,075 | Calling regression() with parameter loss='weighted_crossentropy' | Hello, I am currently also trying to implement a weighted CE loss function. I'd really appreciate some guidance on how to call this function from the `loss=` parameter of the `tflearn.regression()` function.
The following attempt to use the above method in my code yields:
```
net_2 = net = tflearn.input_data(shape... | closed | 2018-07-19T22:15:44Z | 2018-07-20T00:26:44Z | https://github.com/tflearn/tflearn/issues/1075 | [] | tnightengale | 0 |
hbldh/bleak | asyncio | 754 | Collect notify data from two devices (different MACs) with same UUIDs + Identify devices | * bleak version: 0.14.2
* Python version: 3.9.7
* Operating System: Ubuntu 21.10
* BlueZ version (`bluetoothctl -v`) in case of Linux: 5.60
### Description
*I apologize in advance, I am sure the library is working fine here, this is just my limited BLE knowledge causing issues*
- I will hopefully repay any he... | closed | 2022-01-30T17:22:33Z | 2022-01-30T21:56:58Z | https://github.com/hbldh/bleak/issues/754 | [] | cooperlees | 4 |
neuml/txtai | nlp | 827 | How to index() or upsert() only on specific index? | Heyo,
Love txtai, amazing work.
I have two subindexes and want to perform an `upsert()` on only one of the subindexes:
Basically I'd want something like this:
```python
embeddings = Embeddings({
"indexes": {
"raw" : CONFIG,
"llm": CONFIG
}
})
embedding... | closed | 2024-12-02T18:18:48Z | 2025-02-23T14:54:34Z | https://github.com/neuml/txtai/issues/827 | [] | byt3bl33d3r | 6 |
plotly/dash | data-visualization | 2,925 | _validate.py "RuntimeError: dictionary changed size during iteration" | **Environment**
```
dash 2.17.1
dash-bootstrap-components 1.5.0
dash-bootstrap-templates 1.0.8
dash-core-components 2.0.0
dash-html-components 2.0.0
dash-loading-spinners 1.0.2
dash-table 5.0.0
```
**Describe the bug**
When starting a large Dash app (c... | open | 2024-07-20T11:34:29Z | 2024-08-13T19:55:53Z | https://github.com/plotly/dash/issues/2925 | [
"bug",
"P3"
] | mbworth | 1 |
Guovin/iptv-api | api | 226 | 关于docker运行的问题 | 建议生成的文件和配置的文件在一个单独的文件夹,比如
/tv-driver/output
/tv-driver/config
这样映射只需要映射子文件夹,就不会提示脚本不存在了
```
实现宿主机文件与容器文件同步,修改模板、配置、获取更新结果文件可直接在宿主机文件夹下操作
注:使用此命令运行容器,请务必先clone本项目至宿主机
``` | closed | 2024-08-04T05:27:09Z | 2024-08-14T10:01:37Z | https://github.com/Guovin/iptv-api/issues/226 | [
"enhancement"
] | QAQQL | 3 |
numba/numba | numpy | 9,994 | Numba spams output on startup | <!--
Thanks for opening an issue! To help the Numba team handle your information
efficiently, please first ensure that there is no other issue present that
already describes the issue you have
(search at https://github.com/numba/numba/issues?&q=is%3Aissue).
-->
## Reporting a bug
<!--
Before submitting a bug repor... | open | 2025-03-17T19:54:27Z | 2025-03-19T14:19:02Z | https://github.com/numba/numba/issues/9994 | [
"needtriage"
] | JC3 | 7 |
pytorch/vision | machine-learning | 8,819 | torchvision' object has no attribute '_cuda_version' | ### 🐛 Describe the bug
I am using a MacBook 15.1.1 (24B91) running Python 3.10.
I installed torch and torchvision through pip with `pip install -U torch torchvision` and it gave the following output:
```
Installing collected packages: torch, torchvision
Attempting uninstall: torch
Found existing insta... | closed | 2024-12-19T21:53:57Z | 2025-01-22T20:17:24Z | https://github.com/pytorch/vision/issues/8819 | [] | Flippchen | 2 |
recommenders-team/recommenders | data-science | 1,861 | [ASK] Error in NCFDataset creation | ### Description
Hello all,
i'm trying to use the NCF_deep_dive notebook with my own data.
With the following structure
<html>
<body>
<!--StartFragment-->
| usr_id | code_id | amt_trx | bestelldatum
-- | -- | -- | -- | --
0 | 0 | 35 | 1 | 2022-03-01
1 | 0 | 2 | 1 | 2022-03-01
2 | 0 | 18 | 1 | 2022-0... | open | 2022-11-28T09:13:17Z | 2022-11-28T14:20:49Z | https://github.com/recommenders-team/recommenders/issues/1861 | [
"help wanted"
] | mrcmoresi | 0 |
adbar/trafilatura | web-scraping | 245 | Use a class to gather all extraction settings | Implementing an extraction class instead of passing series of arguments to extraction functions would simplify the code. | closed | 2022-09-08T10:32:21Z | 2022-09-12T15:12:58Z | https://github.com/adbar/trafilatura/issues/245 | [
"enhancement"
] | adbar | 1 |
vimalloc/flask-jwt-extended | flask | 525 | Signature verification failed with just generated tokens | I have flask-jwt-extended configured with the following settings, with an app running on a docker container behind nginx:
```
JWT_SECRET_KEY = secrets.token_urlsafe(24)
JWT_ACCESS_TOKEN_EXPIRES = timedelta(hours=1)
JWT_TOKEN_LOCATION = ['cookies']
JWT_COOKIE_CSRF_PROTECT = False
JWT_CSRF_CHECK_FORM = False
JWT_C... | closed | 2023-09-23T19:30:19Z | 2023-09-24T19:23:16Z | https://github.com/vimalloc/flask-jwt-extended/issues/525 | [] | flixman | 1 |
benbusby/whoogle-search | flask | 303 | [BUG] Slow search on Raspberry Pi and incorrect env variables | **Describe the bug**
* Whoogle takes a long time to return search results (between 5-15s)
* Autocomplete doesn't work.
* whoogle.env seems to be ignored.
**Deployment Method**
- [ ] Heroku (one-click deploy)
- [x] Docker -- buildx/experimental
- [ ] `run` executable
- [ ] pip/pipx
- [ ] Other: [describe setu... | closed | 2021-05-02T14:16:05Z | 2021-05-20T13:59:00Z | https://github.com/benbusby/whoogle-search/issues/303 | [
"bug"
] | accountForIssues | 10 |
huggingface/datasets | deep-learning | 6,842 | Datasets with files with colon : in filenames cannot be used on Windows | ### Describe the bug
Datasets (such as https://huggingface.co/datasets/MLCommons/peoples_speech) cannot be used on Windows due to the fact that windows does not allow colons ":" in filenames. These should be converted into alternative strings.
### Steps to reproduce the bug
1. Attempt to run load_dataset on MLCo... | open | 2024-04-26T00:14:16Z | 2024-04-26T00:14:16Z | https://github.com/huggingface/datasets/issues/6842 | [] | jacobjennings | 0 |
dmlc/gluon-nlp | numpy | 1,034 | Error when using fp16 trainer | ## Description
I use fp16 trainer in fp16_utils.py to train model. I got the following error.
@eric-haibin-lin
### Error Message
File "/home/ec2-user/project/src/deep/utils/fp16_utils.py", line 179, in step
overflow = self._scaler.has_overflow(self.fp32_trainer._params)
File "/home/ec2-user/project/... | closed | 2019-12-03T01:04:51Z | 2019-12-05T02:48:29Z | https://github.com/dmlc/gluon-nlp/issues/1034 | [
"bug"
] | rich-junwang | 0 |
raphaelvallat/pingouin | pandas | 204 | Homoscedasticity is Incorrectly Calculated for Wide-Format DataFrames | Hello,
First, thanks for this great package! It's been great to use in place of scipy.stats and statsmodels.
It seems the following part of the homoscedasticity function (lines 342 - 347 in distribution.py) performs incorrectly:
```python
if dv is None and group is None:
# Wide-format
... | closed | 2021-10-20T22:41:18Z | 2021-10-28T22:11:51Z | https://github.com/raphaelvallat/pingouin/issues/204 | [
"bug :boom:",
"URGENT :warning:"
] | StephenB1289 | 4 |
pytorch/vision | machine-learning | 8,902 | Different Behaviors of tranforms.ToTensor and transforms.v2.ToTensor | ## 🐛 Describe the bug
In the [docs](https://pytorch.org/vision/stable/transforms.html#conversion) it says
> Deprecated
>
> | Func | Desc |
> |---|---|
> | v2.ToTensor() | [DEPRECATED] Use v2.Compose([v2.ToImage(), v2.ToDtype(torch.float32, scale=True)]) instead. |
But when using the suggested code, the values are ... | closed | 2025-02-07T21:36:36Z | 2025-02-19T16:34:40Z | https://github.com/pytorch/vision/issues/8902 | [] | jneuendorf | 1 |
gto76/python-cheatsheet | python | 126 | Python | thank you | closed | 2022-03-10T18:52:28Z | 2022-12-14T05:30:29Z | https://github.com/gto76/python-cheatsheet/issues/126 | [] | SouthKoreanLee | 1 |
Zeyi-Lin/HivisionIDPhotos | machine-learning | 1 | 换装的话是否替换背景? | closed | 2023-07-03T03:18:08Z | 2023-07-04T05:14:23Z | https://github.com/Zeyi-Lin/HivisionIDPhotos/issues/1 | [] | hai411741962 | 1 | |
python-visualization/folium | data-visualization | 1,142 | Working on a Colab Notebook (link included) to help teachers to use in the classroom | As a teacher, folium is a great tool. Sometimes students and teachers do not have enough coding knowledge to work with packages. Therefore, by writing it in a Colab, teachers can easily edit specific areas to use in the classroom. I just started this and I am working through the documentation in getting started.
... | closed | 2019-05-04T21:17:20Z | 2022-11-26T16:56:49Z | https://github.com/python-visualization/folium/issues/1142 | [
"MentoredSprintsPyCon2019"
] | KellyPared | 0 |
JaidedAI/EasyOCR | deep-learning | 404 | Tajik Language | Here's needed data. Can you tell me when your ocr can learn it, so i can use it? Thank you!
[easyocr.zip](https://github.com/JaidedAI/EasyOCR/files/6220472/easyocr.zip)
| closed | 2021-03-29T08:59:30Z | 2021-03-31T01:36:32Z | https://github.com/JaidedAI/EasyOCR/issues/404 | [] | KhayrulloevDD | 1 |
jupyterhub/zero-to-jupyterhub-k8s | jupyter | 3,545 | hook and continuous image pullers' DaemonSets: configuring k8s ServiceAccount - yes or no? | ## Update
My take is that we should help people pass some compliance tests etc, even though it doesn't improve security in this case. There could be some edge case where use of service accounts can be relevant still, such as when PSPs were around or similar.
## Background
There are two sets of machinery to pul... | closed | 2024-10-15T09:20:01Z | 2025-01-12T15:26:20Z | https://github.com/jupyterhub/zero-to-jupyterhub-k8s/issues/3545 | [
"new"
] | consideRatio | 11 |
plotly/dash | plotly | 2,710 | [Feature Request] support multiple URL path levels in path template | I'd like to suggest the following behavior for interpreting path templates as part of the pages feature.
The following example can illustrate the requested behavior:
```
dash.register_page("reports", path_template="/reports/<product>/<feature>/<report_type>/<data_version>")
def layout(product: str | None = No... | closed | 2023-12-10T07:45:07Z | 2024-05-31T20:14:05Z | https://github.com/plotly/dash/issues/2710 | [] | yreiss | 1 |
alteryx/featuretools | data-science | 2,611 | Update for compatibility with pyarrow 13.0.0 in Featuretools | Pyarrow v13.0.0 appears to have introduced changes that are causing some unit tests to fail. These failures should be investigated and, once resolved, the upper version restriction on pyarrow in pyproject.toml should be removed (both in test requirements and spark requirements). | closed | 2023-09-06T14:13:10Z | 2024-02-15T21:58:20Z | https://github.com/alteryx/featuretools/issues/2611 | [] | thehomebrewnerd | 1 |
sqlalchemy/alembic | sqlalchemy | 575 | Insert values to created table | How i can insert variables into created tables?
When i create them and commit it will raise error, that table is not created | closed | 2019-06-05T07:41:47Z | 2019-06-05T07:45:13Z | https://github.com/sqlalchemy/alembic/issues/575 | [] | mrquokka | 0 |
2noise/ChatTTS | python | 155 | 这个报错是怎么回事 | To create a public link, set `share=True` in `launch()`.
INFO:ChatTTS.core:All initialized.
WARNING:ChatTTS.core:Package WeTextProcessing not found! Run: conda install -c conda-forge pynini=2.1.5 && pip install WeTextProcessing
Traceback (most recent call last):
File "C:\Users\20514\AppDat... | closed | 2024-05-31T22:57:23Z | 2024-06-24T08:29:28Z | https://github.com/2noise/ChatTTS/issues/155 | [
"bug"
] | iscc-top | 4 |
microsoft/nni | data-science | 5,350 | . | closed | 2023-02-14T11:44:10Z | 2023-02-15T10:20:42Z | https://github.com/microsoft/nni/issues/5350 | [] | Nafees-060 | 6 | |
onnx/onnx | tensorflow | 6,577 | ONNX produces different results compared to torch.geometric | # Bug Report
### Is the issue related to model conversion?
I have a torch model that i need to convert to onnx to run in C++. However, i cannot produce the same output using the torch model and the one with onnx.
### Describe the bug
I think SAGEConv Layer causes some problems when converted to onnx. It corre... | open | 2024-12-06T16:04:24Z | 2024-12-06T16:05:12Z | https://github.com/onnx/onnx/issues/6577 | [
"bug"
] | PliniLeonardo | 0 |
tortoise/tortoise-orm | asyncio | 1,744 | Robyn registe tortoise, but unable to close the robyn program properl | I tried using tortoise_orm with the Robyn. I referenced the registration method from Sanic. When I didn't remove the '@app.shutdown_handler' function, I couldn't properly close the Robyn program using Ctrl+C. After I removed the '@app.shutdown_handler' function, it closed normally
```from robyn import Robyn, jsonify... | closed | 2024-10-22T08:57:00Z | 2024-11-26T12:23:53Z | https://github.com/tortoise/tortoise-orm/issues/1744 | [] | JiaLiangChen99 | 1 |
jina-ai/clip-as-service | pytorch | 359 | server does not start successfully | **Prerequisites**
> Please fill in by replacing `[ ]` with `[x]`.
* [x ] Are you running the latest `bert-as-service`?
* [x] Did you follow [the installation](https://github.com/hanxiao/bert-as-service#install) and [the usage](https://github.com/hanxiao/bert-as-service#usage) instructions in `README.md`?
* [x ]... | open | 2019-05-22T11:25:45Z | 2020-08-19T17:54:34Z | https://github.com/jina-ai/clip-as-service/issues/359 | [] | mainulquraishi | 2 |
OpenInterpreter/open-interpreter | python | 1,049 | UnicodeDecodeError: 'utf-8' codec can't decode byte 0xff in position 0: invalid start byte | ### Describe the bug
(oi) C:\Users\Matas>interpreter
Traceback (most recent call last):
File "<frozen runpy>", line 198, in _run_module_as_main
File "<frozen runpy>", line 88, in _run_code
File "C:\Users\Matas\anaconda3\envs\oi\Scripts\interpreter.exe\__main__.py", line 4, in <module>
File "C:\Users\Matas... | open | 2024-03-01T19:38:53Z | 2024-03-01T19:38:53Z | https://github.com/OpenInterpreter/open-interpreter/issues/1049 | [
"Bug"
] | Politas380 | 0 |
ludwig-ai/ludwig | computer-vision | 3,759 | test issue notification | closed | 2023-10-30T23:36:37Z | 2023-10-30T23:37:12Z | https://github.com/ludwig-ai/ludwig/issues/3759 | [] | geoffreyangus | 1 | |
marshmallow-code/flask-marshmallow | sqlalchemy | 194 | URLFor and many relationships | I haven't found much luck with the following.
I'd like to use URLFor to describe a URL for a relationship. I am rendering the child model `Lease`, which back_populates `resources`.
When I am deserializing the `Lease` object and `links` is generated, the parent `Resource` object is desereialized as well, but sinc... | open | 2020-06-19T22:36:35Z | 2020-06-19T22:36:57Z | https://github.com/marshmallow-code/flask-marshmallow/issues/194 | [] | retr0h | 0 |
StackStorm/st2 | automation | 5,787 | Concurrency issues with workflow or action | Scene:
Receive and consume messages in bulk using kafka
issue:


The workflow i... | open | 2022-10-26T03:13:54Z | 2022-10-26T03:13:54Z | https://github.com/StackStorm/st2/issues/5787 | [] | simonli866 | 0 |
jupyter/nbviewer | jupyter | 820 | nbviewer.jupyter.org: External images are not updated | If you have links to externalized images in a notebook (but located in the same git repo), when those images change, they are not updated by https://nbviewer.jupyter.org/, even when adding `?flush_cache=true`.
**To Reproduce**
Compare these 2 URLs:
* https://nbviewer.jupyter.org/github/jhermann/jupyter-by-exampl... | closed | 2019-03-11T13:52:05Z | 2019-03-18T11:12:52Z | https://github.com/jupyter/nbviewer/issues/820 | [] | jhermann | 2 |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.