repo_name stringlengths 9 75 | topic stringclasses 30
values | issue_number int64 1 203k | title stringlengths 1 976 | body stringlengths 0 254k | state stringclasses 2
values | created_at stringlengths 20 20 | updated_at stringlengths 20 20 | url stringlengths 38 105 | labels listlengths 0 9 | user_login stringlengths 1 39 | comments_count int64 0 452 |
|---|---|---|---|---|---|---|---|---|---|---|---|
kynan/nbstripout | jupyter | 88 | pip install fails | Here's what happened when I tried the pip install on my py3.6 on my macOS 10.12.6:
```bash
pip install --upgrade nbstripout (py36)
Requirement already up-to-date: nbstripout in /Users/klay6683/miniconda3/envs/py36/lib/python3.6/site-packages (0.3.3)
Requirement ... | closed | 2018-10-05T20:16:41Z | 2018-10-16T18:31:36Z | https://github.com/kynan/nbstripout/issues/88 | [
"resolution:invalid"
] | michaelaye | 2 |
thp/urlwatch | automation | 3 | BeautifulSoup usage? | Hello,
First, thank you for your great product. I created a set of RPMs on [AXIVO repository](https://www.axivo.com/packages/setup), available for CentOS 6 (and soon for CentOS 7 with BeautifulSoup4):
```
$ yum --enablerepo=axivo list | egrep 'urlwatch|futures|beautifulsoup'
python-beautifulsoup.noarch 3... | closed | 2014-08-21T23:32:42Z | 2014-10-27T08:06:56Z | https://github.com/thp/urlwatch/issues/3 | [] | ghost | 3 |
davidteather/TikTok-Api | api | 1,031 | Empty response | Someone experiencing the same problem with video api? Can't get info of video becuase api is giving empty page
Traceback (most recent call last):
File "E:\tiktok\s.py", line 94, in <module>
data = video.info()
File "E:\tiktok\venv\lib\site-packages\TikTokApi\api\video.py", line 74, in info
return sel... | closed | 2023-07-18T15:51:59Z | 2023-08-08T21:39:53Z | https://github.com/davidteather/TikTok-Api/issues/1031 | [] | dortesy | 6 |
microsoft/unilm | nlp | 1,096 | error when fine-tuning VLMO | **Describe**
Model I am using (UniLM, MiniLM, LayoutLM ...): VLMO

error happens when I fine-tuned pre-trained VLMO model on COCO retrieval task... | closed | 2023-05-17T13:27:26Z | 2023-06-14T11:51:02Z | https://github.com/microsoft/unilm/issues/1096 | [] | det-tu | 5 |
CorentinJ/Real-Time-Voice-Cloning | python | 1,060 | Quality of generated audio | When using a recording from the LibriSpeech downloaded dataset, a good ratio of the generated audio pieces sound good and accurate. However, whenever I record some audio and use that, no matter who the speaker is, all the generated audio pieces sound the same. Is there any way I can fix this, or am I not understanding ... | open | 2022-05-01T20:45:45Z | 2022-10-21T17:38:13Z | https://github.com/CorentinJ/Real-Time-Voice-Cloning/issues/1060 | [] | aryanpanpalia | 3 |
Kanaries/pygwalker | plotly | 589 | How to use pygwalker in fastapi? | I used the following code to read the file and send it to pygwalker, but the returned result rendering has no effect and cannot produce dynamic rendering results like flask.
```html index.html
<!DOCTYPE html>
<html lang="en">
<head>
<meta charset="UTF-8">
<meta name="viewport" content="width=device-width... | closed | 2024-07-05T02:34:44Z | 2024-07-15T12:50:05Z | https://github.com/Kanaries/pygwalker/issues/589 | [] | soundmemories | 1 |
quokkaproject/quokka | flask | 399 | Example of making forms using Custom Values | Is there an example to make a new form that contain fields based on existing Post custom values?
For example:
A post have Price custom value, I want to make a form that has Price as one of the fields.
| closed | 2016-10-09T17:04:19Z | 2018-02-06T13:45:53Z | https://github.com/quokkaproject/quokka/issues/399 | [] | ncmonger | 0 |
koxudaxi/datamodel-code-generator | fastapi | 2,346 | When I using as module calling generate method and pass in disable_timestamp=True, the timestamp will still be displayed | ```py
from datamodel_code_generator import (
DataModelType, # 注意:DataModelType 不在 __all__ 中定义,可能在未来的版本中发生变化
InputFileType,
PythonVersion,
generate,
)
generate(
json_str,
input_file_type=InputFileType.Json, # JSON类型
output=model_file, # 输出文件路径
output_mo... | open | 2025-03-15T05:22:21Z | 2025-03-15T05:22:21Z | https://github.com/koxudaxi/datamodel-code-generator/issues/2346 | [] | Airpy | 0 |
recommenders-team/recommenders | machine-learning | 1,807 | [ASK] Running evaluation on test set for DKN | ### Description
I've have been using [DKN Deep Dive](https://github.com/microsoft/recommenders/blob/aeb6b0b12e177b3eaf55bb7ab2b747549a541394/examples/02_model_content_based_filtering/dkn_deep_dive.ipynb), but I wanted to add evaluation on the test data. When I run `model.run_eval(test_file)` I am getting an error. I b... | open | 2022-08-04T17:16:03Z | 2022-08-04T17:16:03Z | https://github.com/recommenders-team/recommenders/issues/1807 | [
"help wanted"
] | Bhammin | 0 |
deepspeedai/DeepSpeed | pytorch | 7,117 | safe_get_full_grad & safe_set_full_grad | deepspeed 0.15.3
zero 3 is used
For "safe_get_full_grad", does it return the same gradient values on each process/rank?
As for "safe_set_full_grad", should it be called on all the processes/ranks? or just one of them is enough?
If it's the former one, users will need to ensure gradient values to be set on each proces... | open | 2025-03-09T10:10:19Z | 2025-03-21T22:12:20Z | https://github.com/deepspeedai/DeepSpeed/issues/7117 | [] | ProjectDisR | 3 |
apache/airflow | machine-learning | 47,718 | Support AssetRef when binding to AssetAlias | ### Body
A followup to #47677. This will need some changes to the `outlet_events` exchange format.
### Committer
- [x] I acknowledge that I am a maintainer/committer of the Apache Airflow project. | open | 2025-03-13T09:51:07Z | 2025-03-13T09:53:27Z | https://github.com/apache/airflow/issues/47718 | [
"kind:meta",
"area:datasets"
] | uranusjr | 0 |
sqlalchemy/sqlalchemy | sqlalchemy | 10,497 | Make ORM private api more explicitly so | Related to the discussion in https://github.com/sqlalchemy/sqlalchemy/discussions/10474
Rename function to have underscore in them / rename private modules to have an udnerstore.
This in an attempt to discourage usage of private ORM api by 3-rd party library.
If such api is required then a clear feature request ... | closed | 2023-10-18T15:31:27Z | 2024-11-19T12:50:47Z | https://github.com/sqlalchemy/sqlalchemy/issues/10497 | [
"orm"
] | CaselIT | 9 |
huggingface/transformers | machine-learning | 36,410 | Conflicting Keras 3 mitigations | ### System Info
- `transformers` version: 4.49.0
- Platform: Linux-6.13.4-zen1-1-zen-x86_64-with-glibc2.41
- Python version: 3.13.2
- Huggingface_hub version: 0.29.1
- Safetensors version: 0.5.2
- Accelerate version: 1.4.0
- Accelerate config: not found
- DeepSpeed version: not installed
- PyTorch version (GPU?): 2.6... | closed | 2025-02-26T05:40:32Z | 2025-02-26T14:38:42Z | https://github.com/huggingface/transformers/issues/36410 | [
"bug"
] | mistersmee | 2 |
huggingface/datasets | numpy | 7,299 | Efficient Image Augmentation in Hugging Face Datasets | ### Describe the bug
I'm using the Hugging Face datasets library to load images in batch and would like to apply a torchvision transform to solve the inconsistent image sizes in the dataset and apply some on the fly image augmentation. I can just think about using the collate_fn, but seems quite inefficient.
... | open | 2024-11-26T16:50:32Z | 2024-11-26T16:53:53Z | https://github.com/huggingface/datasets/issues/7299 | [] | fabiozappo | 0 |
thtrieu/darkflow | tensorflow | 659 | How boxes in annotation are mapped into output vector | Say I have only 1 class A to detect and data set with a few jpg images.
Each image contains a bounding box(s) with the class A.
Should I somehow provide negative samples where there is no class A?
If so, how the annotation should look like in this case?
Or darkflow gets those negative samples outside the boxes... | open | 2018-03-23T09:03:03Z | 2018-03-23T09:03:03Z | https://github.com/thtrieu/darkflow/issues/659 | [] | vfateev | 0 |
ploomber/ploomber | jupyter | 175 | Make it easy to add new tasks | We should have a way to quickly create new tasks by providing templates.
Proposed API:
```sh
ploomber add python/sql
```
Asks for name and then creates the file.
Python:
```python
# ---
# jupyter:
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + ... | closed | 2020-07-07T04:34:04Z | 2020-07-08T06:18:25Z | https://github.com/ploomber/ploomber/issues/175 | [] | edublancas | 0 |
davidsandberg/facenet | computer-vision | 829 | Large loss with pretrained_model trained on CASIA-WebFace_align | I run:
python src/train_softmax.py --data_dir /home/han/facenet/CASIA-WebFace_align --image_size 160 __embedding_size 512 /home/han/facenet/src/pretrained_models/20180408-102900/model-2018408-102900.ckpt-90
Theoretically, the accuracy of pretrained model trained 90 epoch should be nearly 0, but when I trained on the ... | open | 2018-07-31T04:44:08Z | 2018-07-31T04:50:43Z | https://github.com/davidsandberg/facenet/issues/829 | [] | Victoria2333 | 0 |
Anjok07/ultimatevocalremovergui | pytorch | 644 | Is it possible to seperate the backing vocal to a channel? | How to remove backing vocal? | open | 2023-07-04T05:59:26Z | 2023-07-04T05:59:26Z | https://github.com/Anjok07/ultimatevocalremovergui/issues/644 | [] | zhouhao27 | 0 |
dadadel/pyment | numpy | 35 | Problem to install on RHEL-7 (with python 2.7) | ```
matej@mitmanek: pyment (master)$ python setup.py develop --user
running develop
running egg_info
writing Pyment.egg-info/PKG-INFO
writing top-level names to Pyment.egg-info/top_level.txt
writing dependency_links to Pyment.egg-info/dependency_links.txt
writing entry points to Pyment.egg-info/entry_points.txt
... | closed | 2017-04-20T08:55:59Z | 2021-03-08T14:51:11Z | https://github.com/dadadel/pyment/issues/35 | [] | mcepl | 2 |
fastapi/fastapi | python | 12,246 | OpenAPI servers not being returned according how the docs say they should be | ### Discussed in https://github.com/fastapi/fastapi/discussions/12226
<div type='discussions-op-text'>
<sup>Originally posted by **mzealey** September 19, 2024</sup>
### First Check
- [X] I added a very descriptive title here.
- [X] I used the GitHub search to find a similar question and didn't find it.
- [... | open | 2024-09-22T10:29:30Z | 2024-09-22T16:10:30Z | https://github.com/fastapi/fastapi/issues/12246 | [
"question"
] | Kludex | 3 |
521xueweihan/HelloGitHub | python | 2,818 | 【开源自荐】shares: vscode股票看盘插件 | ## 推荐项目
<!-- 这里是 HelloGitHub 月刊推荐项目的入口,欢迎自荐和推荐开源项目,唯一要求:请按照下面的提示介绍项目。-->
<!-- 点击上方 “Preview” 立刻查看提交的内容 -->
<!--仅收录 GitHub 上的开源项目,请填写 GitHub 的项目地址-->
- 项目地址:https://github.com/xxjwxc/shares
<!--请从中选择(C、C#、C++、CSS、Go、Java、JS、Kotlin、Objective-C、PHP、Python、Ruby、Rust、Swift、其它、书籍、机器学习)-->
- 类别:(Go,vue)
<... | open | 2024-09-29T02:49:33Z | 2024-10-23T01:53:54Z | https://github.com/521xueweihan/HelloGitHub/issues/2818 | [] | xxjwxc | 1 |
paperless-ngx/paperless-ngx | django | 7,470 | [BUG] User cannot login after upgrade | ### Description
User cannot login after Upgrade from 2.11.0 to 2.11.4
However, Admin can login.
### Steps to reproduce
Load Login page.
login as regular user.
### Webserver logs
```bash
{"headers":{"normalizedNames":{},"lazyUpdate":null},"status":403,"statusText":"OK","url":"https://docs.my.domain/api/ui_setting... | closed | 2024-08-14T19:27:30Z | 2024-09-14T03:04:25Z | https://github.com/paperless-ngx/paperless-ngx/issues/7470 | [
"not a bug"
] | manuelkamp | 3 |
littlecodersh/ItChat | api | 65 | 如何设置屏蔽手机端发送的消息 | ``` Python
@itchat.msg_register('Text', isGroupChat=False)
def auto_reply(msg):
try:
print itchat.get_friends(userName=msg['FromUserName'])['NickName']
print itchat.get_friends(userName=msg['ToUserName'])['NickName']
print msg['Content']
except:
pass
```
结果,只输出自己的NickName。。。。
| closed | 2016-08-15T16:14:06Z | 2016-08-16T11:46:48Z | https://github.com/littlecodersh/ItChat/issues/65 | [
"bug"
] | hadwinfu | 2 |
pyg-team/pytorch_geometric | deep-learning | 8,936 | Cannot install both torch-scatter and torch-sparse on PyTorch 2.2 | ### 🐛 Describe the bug
When attempting to import the `torch_geometric` package in my jupyter notebook, I encounter the warnings below:
```
[/Library/Frameworks/Python.framework/Versions/3.11/lib/python3.11/site-packages/torch_geometric/typing.py:72](https://file+.vscode-resource.vscode-cdn.net/Library/Frameworks/Py... | closed | 2024-02-19T06:45:46Z | 2024-02-22T15:34:08Z | https://github.com/pyg-team/pytorch_geometric/issues/8936 | [
"bug"
] | Panichito | 6 |
CorentinJ/Real-Time-Voice-Cloning | pytorch | 888 | The voice is way too different can anyone explain how to improve? | I was trying an audio from this clip https://youtu.be/ZZuWSkuqjPA downloaded and converted to wav.
There is a lot of krrrr krrr noise in the output.
I tried passing a 30sec, 5 min, 10min clips, and also cleaned the WAV files with noise reduction in audacity and gave them as input; still no difference | closed | 2021-11-08T20:35:40Z | 2021-11-08T22:59:17Z | https://github.com/CorentinJ/Real-Time-Voice-Cloning/issues/888 | [] | 0xrushi | 1 |
pytest-dev/pytest-qt | pytest | 147 | Missing bits from pytest-qt 2.0 docs | @The-Compiler
> I'm a bit late, but I just realised check_params_cb and order aren't mentioned in the signal docs at all, and order isn't even in the changelog.
| closed | 2016-07-29T11:06:45Z | 2016-10-19T00:08:52Z | https://github.com/pytest-dev/pytest-qt/issues/147 | [] | nicoddemus | 3 |
fastapi/sqlmodel | sqlalchemy | 50 | Is dynamic schema supported like in SQLAlchemy? | ### First Check
- [X] I added a very descriptive title to this issue.
- [X] I used the GitHub search to find a similar issue and didn't find it.
- [X] I searched the SQLModel documentation, with the integrated search.
- [X] I already searched in Google "How to X in SQLModel" and didn't find any information.
- [X] I al... | open | 2021-08-28T06:14:59Z | 2021-11-19T06:00:42Z | https://github.com/fastapi/sqlmodel/issues/50 | [
"question"
] | aghanti7 | 2 |
ploomber/ploomber | jupyter | 365 | HTML - Display youtube videos on a responsive table | The videos section in the docs embeds youtube videos one after another: https://ploomber.readthedocs.io/en/latest/videos.html
it'd be better to create a responsive table that displays 2-3 columns and collapses them on mobile, we're using the bootstrap framework so it should be simple to do it
source code: https:/... | closed | 2021-10-15T16:05:54Z | 2021-10-18T23:19:45Z | https://github.com/ploomber/ploomber/issues/365 | [
"documentation",
"good first issue"
] | edublancas | 4 |
graphql-python/graphene-django | graphql | 570 | [question] How to use generated inner types? | For instance:
I have some django model:
```python
class Foo(models.Model):
field = models.CharField(max_length=255, choices=(('foo', 'Foo'), ('bar', 'Bar')))
```
Then I generate object type using `DjangoObjectType`:
```python
class FooNode(DjangoObjectType):
class Meta:
model = Foo
```
G... | closed | 2019-01-13T22:44:02Z | 2021-06-22T18:32:34Z | https://github.com/graphql-python/graphene-django/issues/570 | [] | vanyakosmos | 2 |
ray-project/ray | data-science | 51,506 | CI test windows://python/ray/tests:test_multi_tenancy is consistently_failing | CI test **windows://python/ray/tests:test_multi_tenancy** is consistently_failing. Recent failures:
- https://buildkite.com/ray-project/postmerge/builds/8965#0195aaf1-9737-4a02-a7f8-1d7087c16fb1
- https://buildkite.com/ray-project/postmerge/builds/8965#0195aa03-5c4f-4156-97c5-9793049512c1
DataCaseName-windows://pyt... | closed | 2025-03-19T00:07:58Z | 2025-03-19T21:53:33Z | https://github.com/ray-project/ray/issues/51506 | [
"bug",
"triage",
"core",
"flaky-tracker",
"ray-test-bot",
"ci-test",
"weekly-release-blocker",
"stability"
] | can-anyscale | 2 |
ivy-llc/ivy | pytorch | 28,219 | Fix Frontend Failing Test: torch - tensor.torch.Tensor.__gt__ | closed | 2024-02-07T22:37:13Z | 2024-02-25T10:38:42Z | https://github.com/ivy-llc/ivy/issues/28219 | [
"Sub Task"
] | jacksondm33 | 0 | |
deepset-ai/haystack | pytorch | 8,190 | 🧪 Tools: support for tools in OllamaChatGenerator | closed | 2024-08-09T14:40:08Z | 2024-10-02T13:35:41Z | https://github.com/deepset-ai/haystack/issues/8190 | [
"P2"
] | anakin87 | 0 | |
tatsu-lab/stanford_alpaca | deep-learning | 127 | epoch | closed | 2023-03-23T01:31:56Z | 2023-03-23T01:32:02Z | https://github.com/tatsu-lab/stanford_alpaca/issues/127 | [] | zhengzangw | 0 | |
google-research/bert | tensorflow | 1,149 | Question about fine-tuning BERT on domain-specific dataset | ### Hi.
### I'm asking <ins>do we need to do a random search for hyperparameters - learning rate, batch size, weight decay - when we fine-tune pretrained BERT on a domain-specific dataset?</ins> A common practice introduced in the paper is that we can set the learning rate to 2e-5, batch size to 32, 64. Does this me... | open | 2020-09-16T07:15:15Z | 2020-09-23T01:59:19Z | https://github.com/google-research/bert/issues/1149 | [] | guoxuxu | 0 |
vimalloc/flask-jwt-extended | flask | 152 | Token becomes invalid after reload of service | Hi, I notice next issue:
If you serve flask app with uwsgi, and run your application with "service" command (create service file, put it to /etc/systemd/system), after service reload all tokens become invalid (e.g. for all requests with previously generated tokens returned response with 422 error).
If you need an... | closed | 2018-05-09T11:47:00Z | 2023-12-16T10:33:00Z | https://github.com/vimalloc/flask-jwt-extended/issues/152 | [] | mudrila | 5 |
FlareSolverr/FlareSolverr | api | 1,424 | [TorrentLeech] Error solving the challenge. Timeout after 60.0 seconds. | ### Have you checked our README?
- [X] I have checked the README
### Have you followed our Troubleshooting?
- [X] I have followed your Troubleshooting
### Is there already an issue for your problem?
- [X] I have checked older issues, open and closed
### Have you checked the discussions?
- [X] I hav... | closed | 2024-12-20T01:49:40Z | 2024-12-22T00:57:18Z | https://github.com/FlareSolverr/FlareSolverr/issues/1424 | [
"duplicate"
] | fansollo22 | 11 |
open-mmlab/mmdetection | pytorch | 11,930 | get_flops in Co-DETR ERROR!!!!!!!!!!! | I just run the get_flops.py with config is co_dino_5scale_r50_lsj_8xb2_1x_coco.py
the work done, but the resule is error like this:

I print the state table like this:

Hello Adam sir! First of all, let me tell you that face_recognition is an extremely Robust and Highly Accurate API till date for Facial Detection/ Recognition. KUDOS to your work !!!
I was wondering... | open | 2019-11-23T07:13:54Z | 2022-05-20T01:42:07Z | https://github.com/ageitgey/face_recognition/issues/985 | [] | DixitIshan | 2 |
microsoft/unilm | nlp | 1,066 | The project of E5: How to fine-tune on the E5 model? | Dear author:
The following informations is about the paper and project of E5: [Text Embeddings by Weakly-Supervised Contrastive Pre-training](https://arxiv.org/pdf/2212.03533.pdf). Liang Wang, Nan Yang, Xiaolong Huang, Binxing Jiao, Linjun Yang, Daxin Jiang, Rangan Majumder, Furu Wei, arXiv 2022
I want to fine-tune o... | closed | 2023-04-14T07:26:54Z | 2024-04-25T07:51:26Z | https://github.com/microsoft/unilm/issues/1066 | [] | EasyLuck | 14 |
BMW-InnovationLab/BMW-YOLOv4-Training-Automation | rest-api | 35 | No such file or directory | Hi I have the same issue as others in this issue history.
I have tried solution to set DOWNLOAD_ALL=1 in dockerfile but not works for me.
I have yolov4.weights in the right folder under config/darknet/yolov4_default_weights/
Any help? Thank you. Robert
<img width="1203" alt="image" src="https://user-images.githubus... | closed | 2022-02-24T23:19:27Z | 2022-12-14T09:40:23Z | https://github.com/BMW-InnovationLab/BMW-YOLOv4-Training-Automation/issues/35 | [
"enhancement",
"help wanted"
] | rsicak | 7 |
voila-dashboards/voila | jupyter | 692 | start Jupyter Notebook as Voila app from the OS Windows manager | Similarly to nbopen I would like to start a Jupyter Notebook from e.g. my Windows explorer with a double-click in voila. Has somebody seen something like that? Is it possible? | open | 2020-09-03T19:13:01Z | 2020-12-31T13:18:19Z | https://github.com/voila-dashboards/voila/issues/692 | [] | 1kastner | 4 |
hzwer/ECCV2022-RIFE | computer-vision | 309 | 如何对视频流中特定的某一段进行插帧 | open | 2023-04-21T01:39:13Z | 2023-04-21T02:30:01Z | https://github.com/hzwer/ECCV2022-RIFE/issues/309 | [] | HAL900000 | 0 | |
ansible/awx | django | 15,773 | job_wait times out when job ends with status error | ### Please confirm the following
- [x] I agree to follow this project's [code of conduct](https://docs.ansible.com/ansible/latest/community/code_of_conduct.html).
- [x] I have checked the [current issues](https://github.com/ansible/awx/issues) for duplicates.
- [x] I understand that AWX is open source software provide... | closed | 2025-01-24T07:59:17Z | 2025-02-05T18:16:36Z | https://github.com/ansible/awx/issues/15773 | [
"type:bug",
"needs_triage",
"community"
] | mhallin2 | 1 |
home-assistant/core | python | 141,160 | Zone triggers should be based on state transitions rather than GPS location updates | ### The problem
The device_tracker may not always determine the state based on the GPS location, because GPS location is not always accurate.
I use a third-party APP for positioning. It determines whether I am at home by scanning nearby WiFi. For me, GPS only tells my family my approximate location.
However, today I... | open | 2025-03-23T01:56:27Z | 2025-03-23T07:45:41Z | https://github.com/home-assistant/core/issues/141160 | [] | NXY666 | 0 |
roboflow/supervision | tensorflow | 1,182 | HaloAnnotator does not work | ### Search before asking
- [X] I have searched the Supervision [issues](https://github.com/roboflow/supervision/issues) and found no similar feature requests.
### Question
can you give me a complete code? when i use HaloAnnotator, nothing changes in the image, the detections.mask is None
```python
import cv2
i... | closed | 2024-05-09T07:33:38Z | 2024-05-09T08:11:32Z | https://github.com/roboflow/supervision/issues/1182 | [
"question"
] | wilsonlv | 1 |
httpie/cli | api | 1,461 | Failing tests with responses ≥ 0.22.0 | ## Checklist
- [x] I've searched for similar issues.
- [x] I'm using the latest version of HTTPie.
---
## Minimal reproduction code and steps
1. `git clone https://github.com/httpie/httpie; cd httpie`
2. `pip install 'responses>=0.22.0' .[test]`
3. `pytest`
## Current result
A multitude of failures... | closed | 2022-12-25T18:36:06Z | 2023-01-15T16:58:59Z | https://github.com/httpie/cli/issues/1461 | [
"bug",
"new"
] | alexshpilkin | 1 |
howie6879/owllook | asyncio | 56 | 我又用debian9试着装,出现这个问题 | 我又用debian9试着装,
debian9上安装了宝塔面板,搭建了lnmp
在安装你的教程,安装,都蛮顺利的,到这里
(python36) root@ip-172-26-13-169:~/owllook# pipenv run pip install pip==18.0
Creating a virtualenv for this project…
Pipfile: /root/owllook/Pipfile
Using /root/anaconda3/envs/python36/bin/python3.6m (3.6.8) to create virtualenv…
⠦ Creating virtual envir... | closed | 2019-01-23T02:33:11Z | 2019-01-23T03:36:17Z | https://github.com/howie6879/owllook/issues/56 | [] | aff2018 | 7 |
thtrieu/darkflow | tensorflow | 410 | Anyone knows Conv2d-kernel's format is 'HWCN' or 'WHCN' ? | I am trying to simplify tiny-yolo net.
Conv2d-kernel weights were collected, using [CheckpointReader.get_tensor(key)](https://github.com/tensorflow/tensorflow/blob/master/tensorflow/python/tools/inspect_checkpoint.py)
from official sample
and I got ndarray shaped like (3, 3, 128, 256), for example.
I want ... | open | 2017-09-28T14:54:08Z | 2017-09-28T15:00:59Z | https://github.com/thtrieu/darkflow/issues/410 | [] | Serbipunk | 0 |
ydataai/ydata-profiling | jupyter | 1,197 | Does pandas-profiling work in Jupyter Notebooks on AWS? | Does pandas-profiling work in Jupyter Notebooks on AWS? I understand there are a lot of configuration differences that can lead to issues but whenever I try to produce a profiling report, I get the following errors when I run:
```
profile = ProfileReport(df, 'myreport')
profile.to_file('s3://myfolder/myreport.html... | open | 2022-12-05T18:01:37Z | 2022-12-20T12:12:24Z | https://github.com/ydataai/ydata-profiling/issues/1197 | [
"question/discussion ❓",
"information requested ❔"
] | JohnTravolski | 3 |
aio-libs/aiohttp | asyncio | 9,878 | Interacting with aiohttp.request results in error (version 3.11.0) | ### Describe the bug
We're using the aiohttp.request coroutine, however since updating to v3.11.0 we see the following error:
```python
TypeError: RequestInfo.__new__() missing 1 required positional argument: 'real_url'
```
### To Reproduce
1. Make a call using aiohttp.request, e.g.,
```python
aiohttp.reque... | closed | 2024-11-14T16:48:41Z | 2024-11-14T18:18:24Z | https://github.com/aio-libs/aiohttp/issues/9878 | [
"bug"
] | brent-cybrid | 4 |
roboflow/supervision | computer-vision | 809 | No module name 'supervision', installed supervision==0.18.0 and imported supervision as sv | ### Search before asking
- [X] I have searched the Supervision [issues](https://github.com/roboflow/supervision/issues) and found no similar bug report.
### Bug
Traceback (most recent call last):
File "/home/shivakrishnakarnati/Documents/Programming/ROS/ros_supervision_obj/install/object_det/lib/object_det/obj... | closed | 2024-01-29T14:29:43Z | 2024-01-29T15:21:54Z | https://github.com/roboflow/supervision/issues/809 | [
"bug"
] | shivakarnati | 7 |
schemathesis/schemathesis | pytest | 2,136 | [BUG] Parametrization incompatibility with falcon ASGI apps | ### Checklist
- [x] I checked the [FAQ section](https://schemathesis.readthedocs.io/en/stable/faq.html#frequently-asked-questions) of the documentation
- [x] I looked for similar issues in the [issue tracker](https://github.com/schemathesis/schemathesis/issues)
- [x] I am using the latest version of Schemathesis
... | closed | 2024-04-16T20:33:12Z | 2024-04-29T19:48:47Z | https://github.com/schemathesis/schemathesis/issues/2136 | [
"Priority: High",
"Type: Bug",
"Python: ASGI"
] | BrandonWiebe | 5 |
huggingface/datasets | tensorflow | 7,061 | Custom Dataset | Still Raise Error while handling errors in _generate_examples | ### Describe the bug
I follow this [example](https://discuss.huggingface.co/t/error-handling-in-iterabledataset/72827/3) to handle errors in custom dataset. I am writing a dataset script which read jsonl files and i need to handle errors and continue reading files without raising exception and exit the execution.
`... | open | 2024-07-22T21:18:12Z | 2024-09-09T14:48:07Z | https://github.com/huggingface/datasets/issues/7061 | [] | hahmad2008 | 0 |
matplotlib/matplotlib | data-visualization | 29,732 | [Bug]: Unit tests: MacOS 14 failures: gi-invoke-error-quark | ### Bug summary
During a recent GitHub Actions workflow for unrelated pull request #29721, the following error appeared:
```
E gi._error.GError: gi-invoke-error-quark: Could not locate g_option_error_quark: dlopen(libglib-2.0.0.dylib, 0x0009): tried: 'libglib-2.0.0.dylib' (no such file), '/System/Volumes/Pr... | open | 2025-03-11T14:10:28Z | 2025-03-19T11:37:08Z | https://github.com/matplotlib/matplotlib/issues/29732 | [] | jayaddison | 10 |
ray-project/ray | data-science | 51,188 | [<Ray component: Data] Async map_batches return empty result when execution_options.preserve_order = True | ### What happened + What you expected to happen
### The bug
With async generator, `ray.data.map_batches` will return empty result if we set `ray.data.DataContext.execution_options.preserve_order = True`.
### Expected behavior
The result should not be empty, and is expected to have the same amount of data as the input ... | open | 2025-03-09T03:21:02Z | 2025-03-10T17:27:31Z | https://github.com/ray-project/ray/issues/51188 | [
"bug",
"triage",
"data"
] | Drice1999 | 0 |
apachecn/ailearning | python | 347 | 图像压缩 | 在使用SVD对图像压缩的时候,为什么最后32*32显示的全是数字“0”, | closed | 2018-04-11T14:19:55Z | 2018-04-15T06:25:51Z | https://github.com/apachecn/ailearning/issues/347 | [] | cjr0106 | 4 |
QuivrHQ/quivr | api | 3,055 | Display all knowledge in a knowledge management system tab | For Integrations, each integration tool (Gdrive, Notion, etc) is a folder, and each time, each account is a subfolder. | closed | 2024-08-22T14:37:29Z | 2024-10-23T08:04:01Z | https://github.com/QuivrHQ/quivr/issues/3055 | [
"Feature"
] | linear[bot] | 1 |
microsoft/MMdnn | tensorflow | 491 | Failed to convert tf1.4 model to caffe mode,TypeError | Platform (like ubuntu 16.04/win10):
Ubuntu14.04
Python version:
2.7
Source framework with version (like Tensorflow 1.4.1 with GPU):
TensorFlow1.4 gpu
Destination framework with version (like CNTK 2.3 with GPU):
caffe gpu
Pre-trained model path (webpath or webdisk path):
https://github.com/Jackiq/TF-1.4-MODEL
... | open | 2018-11-08T10:16:57Z | 2018-11-08T11:56:23Z | https://github.com/microsoft/MMdnn/issues/491 | [] | Jackiq | 1 |
ray-project/ray | pytorch | 50,961 | [Feedback] Feedback for ray + uv | Hello everyone! As of [Ray 2.43.0](https://github.com/ray-project/ray/releases/tag/ray-2.43.0), we have launched a new integration with `uv run` that we are super excited to share with you all. This will serve as the main Github issue to track any issues or feedback that ya'll might have while using this.
Please shar... | open | 2025-02-27T21:33:22Z | 2025-03-21T17:54:50Z | https://github.com/ray-project/ray/issues/50961 | [] | cszhu | 17 |
jupyter/nbgrader | jupyter | 1,357 | [ERROR] No notebooks were matched by autograded/* | <!--
Thanks for helping to improve nbgrader!
If you are submitting a bug report or looking for support, please use the below
template so we can efficiently solve the problem.
If you are requesting a new feature, feel free to remove irrelevant pieces of
the issue template.
-->
### macOS catalina 10.15.6
... | open | 2020-08-14T08:48:32Z | 2020-08-14T16:48:50Z | https://github.com/jupyter/nbgrader/issues/1357 | [] | ghost | 0 |
robotframework/robotframework | automation | 5,248 | [CLI error] No command line arguments are accepted after the Argumentfile | Robot framework versions: 6.1.1 and 7.1.1
Python version: 3.10.x
OS: Windows (not tested on any other OS)
When the "robot" command is called from command line interface and an argument file is provided through "-A" or "--argumentfile", any arguments passed **after** the argumentfile cause an error (including anoth... | closed | 2024-10-28T00:32:08Z | 2024-10-29T11:10:02Z | https://github.com/robotframework/robotframework/issues/5248 | [] | Mushahar | 1 |
opengeos/streamlit-geospatial | streamlit | 58 | Link is broken | The streamlit app doesn't load | closed | 2022-07-18T20:23:09Z | 2022-07-19T14:30:46Z | https://github.com/opengeos/streamlit-geospatial/issues/58 | [] | osbm | 2 |
keras-team/keras | deep-learning | 20,763 | Use Multi GPU(Mirrored Strategy) training with XLA and AMP(Mixed Precision) | Hi I've encountered some issues while trying to perform **multi-GPU training with XLA** (Accelerated Linear Algebra), **and AMP** (Automatic Mixed Precision).
I'm reaching out to understand if it's possible to use **multi-GPU training with XLA and AMP** together.
If so, I'd like guidance on which versions of tensorfl... | open | 2025-01-15T09:09:15Z | 2025-01-24T04:46:42Z | https://github.com/keras-team/keras/issues/20763 | [
"type:Bug"
] | keshusharmamrt | 4 |
HumanSignal/labelImg | deep-learning | 558 | Find unlabeled images | <!--
I've labeled over 5000 image in a folder but the were around 11 images that i messed. Is there a way to quickly find unlabeled images amongst all the labeled images?
-->
- **OS: Mac**
- **PyQt version:**
| open | 2020-02-25T22:25:16Z | 2020-04-06T19:12:38Z | https://github.com/HumanSignal/labelImg/issues/558 | [] | rojackson13 | 2 |
ageitgey/face_recognition | machine-learning | 1,122 | Getting error while installing sparse module | * face_recognition version:
* Python version:2.7
* Operating System: macOS Catalina version 10.15.3
### Description
I am trying to run face_recognition on my system. Installing sparse Module getting error.
DEPRECATION: Python 2.7 reached the end of its life on January 1st, 2020. Please upgrade your Python as Py... | open | 2020-04-23T16:42:40Z | 2020-05-02T20:01:49Z | https://github.com/ageitgey/face_recognition/issues/1122 | [] | harshitKyal | 0 |
huggingface/datasets | pandas | 6,887 | FAISS load to None | ### Describe the bug
I've use FAISS with Datasets and save to FAISS.
Then load to save FAISS then no error, then ds to None
```python
ds.load_faiss_index('embeddings', 'my_index.faiss')
```
### Steps to reproduce the bug
# 1.
```python
ds_with_embeddings = ds.map(lambda example: {'embeddings': model(transf... | open | 2024-05-09T02:43:50Z | 2024-05-16T20:44:23Z | https://github.com/huggingface/datasets/issues/6887 | [] | brainer3220 | 1 |
marcomusy/vedo | numpy | 825 | Create a mesh in the y-axis direction with partial mesh. | Hi, I looked at the mesh generation method I wanted.
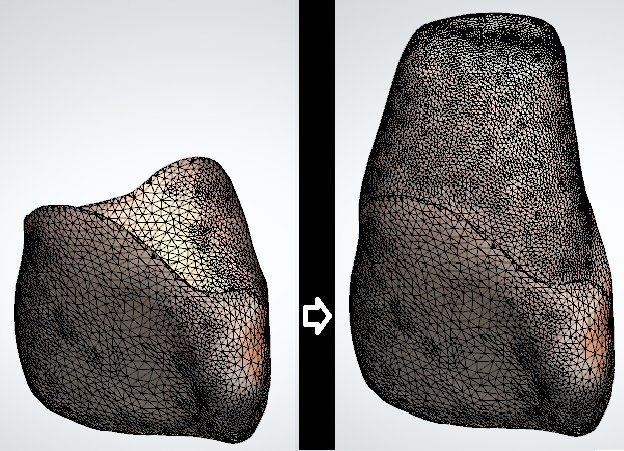

Can I use this library to make a part of the mesh full? Like the image above.
Or is it using AI?
Or Is there a way to create the mesh above?
I at... | closed | 2023-03-06T01:19:45Z | 2023-03-10T00:59:18Z | https://github.com/marcomusy/vedo/issues/825 | [] | ack9437 | 3 |
JoeanAmier/TikTokDownloader | api | 412 | 下载视频后没有声音 | **问题描述**
下载视频后没有声音
**重现步骤**
重现该问题的步骤:
Steps to reproduce the behavior:
1. 选择终端交互
2. 批量下载链接作品(抖音)
3. 下载地址:https://v.douyin.com/i5rhkCP8
**预期结果**
正常播放视频应该有声音 | closed | 2025-02-28T13:02:00Z | 2025-03-01T00:27:59Z | https://github.com/JoeanAmier/TikTokDownloader/issues/412 | [] | shiyigit312 | 3 |
yt-dlp/yt-dlp | python | 12,484 | Faster Way to Check for Manually Created Subtitles | ### Checklist
- [x] I'm asking a question and **not** reporting a bug or requesting a feature
- [x] I've looked through the [README](https://github.com/yt-dlp/yt-dlp#readme)
- [x] I've verified that I have **updated yt-dlp to nightly or master** ([update instructions](https://github.com/yt-dlp/yt-dlp#update-channels))... | closed | 2025-02-26T07:40:16Z | 2025-02-26T16:23:52Z | https://github.com/yt-dlp/yt-dlp/issues/12484 | [
"question"
] | srezasm | 3 |
widgetti/solara | fastapi | 550 | How to disable dynamic rendering/event driven rendering for some components | Hello me again,
I'm continuing to build my application with Solara 😍😍😍, and some of the elements I want to chart takes quite some time to display due a lot of data points, around 30-60 seconds. (The charting library library also needs to be optimised more ). Having dynamic rendering is awesome feature 99% of the t... | closed | 2024-03-11T13:03:40Z | 2024-03-24T10:30:34Z | https://github.com/widgetti/solara/issues/550 | [] | BFAGIT | 7 |
deezer/spleeter | tensorflow | 407 | Error message | I just installed Spleeter v2.5 and got this message.
Starting processing of all songs
Processing D:\~ STUDIO ONE\Backing Tracks\songs\Senza Fine\Spleeter\Monica Mancini - Senza Fine - Reference track.mp3
Traceback (most recent call last):
File "C:\Users\richa\AppData\Roaming\SpleeterGUI\python\Lib\site-package... | closed | 2020-05-31T23:44:15Z | 2020-05-31T23:49:06Z | https://github.com/deezer/spleeter/issues/407 | [] | wsaxmm | 1 |
mars-project/mars | numpy | 2,769 | [BUG] Failed to create Mars DataFrame when mars object exists in a list | <!--
Thank you for your contribution!
Please review https://github.com/mars-project/mars/blob/master/CONTRIBUTING.rst before opening an issue.
-->
**Describe the bug**
Failed to create Mars DataFrame when mars object exists in a list.
**To Reproduce**
To help us reproducing this bug, please provide infor... | closed | 2022-03-01T08:09:12Z | 2022-03-02T10:32:47Z | https://github.com/mars-project/mars/issues/2769 | [
"type: bug",
"mod: dataframe",
"task: medium"
] | qinxuye | 0 |
quokkaproject/quokka | flask | 214 | start error ,cannot load library 'libcairo.so.2 ,Error importing flask-weasyprint! | # python manage.py run0
```
Error importing flask-weasyprint!
PDF support is temporarily disabled.
Manual dependencies may need to be installed.
See,
`http://weasyprint.org/docs/install/#by-platform`_
`https://github.com/Kozea/WeasyPrint/issues/79`_
cannot load library 'libcairo.so.2': libcairo.so.2: cannot o... | closed | 2015-06-16T06:21:36Z | 2015-07-16T02:56:10Z | https://github.com/quokkaproject/quokka/issues/214 | [] | netqyq | 1 |
miguelgrinberg/microblog | flask | 171 | I'm getting an error on Chapter 16 | Hello, I spent few hours but couldn't find any solution. Here is my problem:
When I used reindex method with Post model, I got this error:
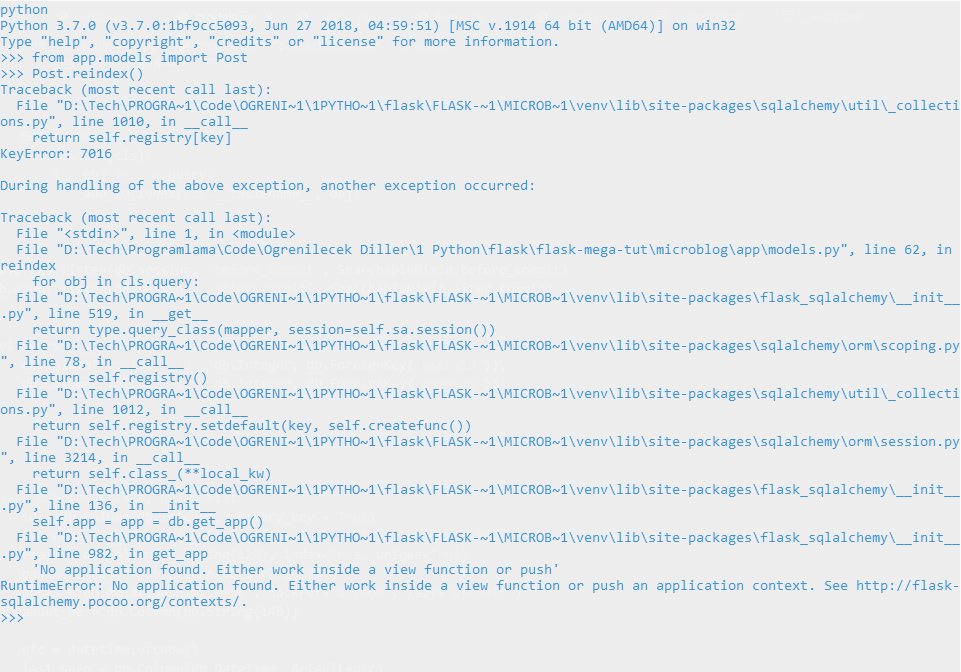
Also I get this when I call query from Post or User model, I don'... | closed | 2019-07-05T23:18:10Z | 2019-07-06T07:18:53Z | https://github.com/miguelgrinberg/microblog/issues/171 | [
"question"
] | alperiox | 2 |
CTFd/CTFd | flask | 2,099 | Theme template project | Vite supports a project template system. We need to create a theme template to create CTFd themes so people can create themes more easily.
https://vitejs.dev/guide/#community-templates | open | 2022-04-22T19:46:07Z | 2024-01-24T06:41:12Z | https://github.com/CTFd/CTFd/issues/2099 | [] | ColdHeat | 1 |
deeppavlov/DeepPavlov | nlp | 1,312 | TorchBertClassifier does not use token_type_ids | Want to contribute to DeepPavlov? Please read the [contributing guideline](http://docs.deeppavlov.ai/en/master/devguides/contribution_guide.html) first.
Please enter all the information below, otherwise your issue may be closed without a warning.
**Issue**:
TorchBertClassifier does not use token_type_ids in [ca... | closed | 2020-09-07T12:45:25Z | 2020-11-13T11:16:23Z | https://github.com/deeppavlov/DeepPavlov/issues/1312 | [
"bug"
] | yurakuratov | 0 |
junyanz/pytorch-CycleGAN-and-pix2pix | pytorch | 757 | Issue with Colab | Hi! I'm using CycleGAN in Google Colab. In some reason when training it doesn't show anything in Checkpoints folder - no folders and no files. I tried with different browsers and different accounts. The files themselves are there and model continues to train after stop with no problem. Anybody knows why that is?
And a... | closed | 2019-09-05T17:38:02Z | 2019-09-30T21:23:15Z | https://github.com/junyanz/pytorch-CycleGAN-and-pix2pix/issues/757 | [] | muxgt | 8 |
Miserlou/Zappa | flask | 1,847 | Zappa deployment throwing 404 error | <!--- Provide a general summary of the issue in the Title above -->
## Context
I am using connexion for my Flask application. I am getting a 404 error \ while accessing teh endpoint. I have tried to dummy down the issue I am facing at https://github.com/ctippur/zappa-connexion-sample
<!--- Provide a more detailed ... | open | 2019-04-03T16:30:40Z | 2022-05-25T04:12:34Z | https://github.com/Miserlou/Zappa/issues/1847 | [] | ctippur | 6 |
nalepae/pandarallel | pandas | 49 | groupby and apply does not work | ```python
def cumulate_asset_scores(dataset):
dataset['counts'] = list(range(len(dataset)))
dataset['corrects'] = dataset['score'].cumsum()
return dataset
from pandarallel import pandarallel
pandarallel.initialize(progress_bar=True, shm_size_mb=30000)
df = dataframe.groupby('user_id').par... | closed | 2019-10-08T12:21:01Z | 2020-01-29T12:15:04Z | https://github.com/nalepae/pandarallel/issues/49 | [] | JonasRSV | 2 |
python-restx/flask-restx | api | 506 | support for OpenAPI Version 3.0.3 - (Swagger) | **Question**
Are you willing to add support for open api spec version 3 guys ?
flask-restx seems to support only swagger spec 2.0 (seen in code here: https://github.com/python-restx/flask-restx/blob/4c748d00eccef675afbde457d43bca5062715a5c/flask_restx/swagger.py#L295)
| open | 2023-01-09T12:08:56Z | 2024-10-11T00:32:08Z | https://github.com/python-restx/flask-restx/issues/506 | [
"question"
] | nskley | 2 |
streamlit/streamlit | python | 10,160 | Expose OAuth errors during `st.login` | ### Checklist
- [X] I have searched the [existing issues](https://github.com/streamlit/streamlit/issues) for similar feature requests.
- [X] I added a descriptive title and summary to this issue.
### Summary
We're soon launching native authentication in Streamlit (see #8518). One thing we left out for now is ... | open | 2025-01-10T23:32:58Z | 2025-01-10T23:33:46Z | https://github.com/streamlit/streamlit/issues/10160 | [
"type:enhancement",
"feature:st.user",
"feature:st.login"
] | jrieke | 1 |
allenai/allennlp | pytorch | 5,703 | Is AllenNLP biased towards BERT model type? | Hi there! I asked a [question](https://stackoverflow.com/questions/73310991/is-allennlp-biased-towards-bert) on stack overflow, as suggested, about a week ago. Just wanted to ping in here in case no one saw it yet. | closed | 2022-08-17T17:32:00Z | 2022-09-01T16:10:13Z | https://github.com/allenai/allennlp/issues/5703 | [
"question",
"stale"
] | pvcastro | 1 |
modelscope/modelscope | nlp | 960 | download_mode = "force_redownload" 未清理干净.arrow缓存 | MsDataset.load(...,download_mode="force_redownload",...
只能保证重新下载最新版附件,
但是解压后generate split的时候若以往有遗留的 .arrow 缓存它还是会去自动用的
若这个 .arrow 是旧附件解压生成的映射,新附件配旧映射会报错(报split内数据量不匹配啥的)
需要让 download_mode="force_redownload" 自动清理.arrow 缓存
调试数据集的时候会遇到这类问题,若更新数据集的zip压缩包或者jsonl附件后,测试调用时即使加了download_mode="force_redownload"也会出现这个问题... | closed | 2024-08-28T01:48:39Z | 2024-10-01T05:32:20Z | https://github.com/modelscope/modelscope/issues/960 | [] | monetjoe | 1 |
roboflow/supervision | tensorflow | 1,769 | Include additional file format in load_yolo_annotations | ### Search before asking
- [X] I have searched the Supervision [issues](https://github.com/roboflow/supervision/issues) and found no similar feature requests.
### Question
Hi,
I wanted to load my yolo dataset which uses `.bmp` images via `sv.DetectionDataset.from_yolo()`. Currently only `["jpg", "jpeg", "png"]`... | closed | 2025-01-07T14:31:50Z | 2025-01-08T09:29:23Z | https://github.com/roboflow/supervision/issues/1769 | [
"question"
] | pirnerjonas | 5 |
deeppavlov/DeepPavlov | nlp | 1,358 | 👩💻📞 DeepPavlov Community Call #4 | > Update: [DeepPavlov Community Call #4 Recording](http://bit.ly/DPCommunityCall4_Video)
> Subscribe for future calls here (last Thursday of the month, 8am Pacific/7pm MSK):
[http://bit.ly/MonthlyDPCommunityCall2021](http://bit.ly/MonthlyDPCommunityCall2021)
Dear DeepPavlov community,
The online DeepPavlov Com... | closed | 2020-12-14T15:17:54Z | 2021-01-21T12:08:38Z | https://github.com/deeppavlov/DeepPavlov/issues/1358 | [
"discussion"
] | moryshka | 0 |
nonebot/nonebot2 | fastapi | 2,672 | Plugin: nonebot-plugin-sanyao | ### PyPI 项目名
nonebot-plugin-sanyao
### 插件 import 包名
nonebot_plugin_sanyao
### 标签
[{"label":"占卜","color":"#52e5ea"}]
### 插件配置项
_No response_ | closed | 2024-04-21T07:46:24Z | 2024-04-22T04:34:36Z | https://github.com/nonebot/nonebot2/issues/2672 | [
"Plugin"
] | afterow | 2 |
supabase/supabase-py | flask | 1,073 | Installing via Conda forgets h2 | # Bug report
<!--
⚠️ We receive a lot of bug reports which have already been solved or discussed. If you are looking for help, please try these first:
- Docs: https://docs.supabase.com
- Discussions: https://github.com/supabase/supabase/discussions
- Discord: https://discord.supabase.com
Before opening a bug repor... | open | 2025-03-10T16:52:42Z | 2025-03-10T16:52:50Z | https://github.com/supabase/supabase-py/issues/1073 | [
"bug"
] | PierreMesure | 1 |
zappa/Zappa | django | 960 | Retained versions and alias | ## Context
When a `num_retained_versions` is set in the zappa_settings.py and an alias has been created manually, a boto3 error trigger.
## Expected Behavior
Versions referenced by alias should be skipped from the deletion process.
## Actual Behavior
The following error trigger and the `zappa update` fails :
... | closed | 2021-04-07T09:25:57Z | 2024-04-13T19:37:02Z | https://github.com/zappa/Zappa/issues/960 | [
"no-activity",
"auto-closed"
] | Yaronn44 | 2 |
ipython/ipython | data-science | 13,970 | Auto suggestions cannot be completed when in the middle of multi-line input | <!-- This is the repository for IPython command line, if you can try to make sure this question/bug/feature belong here and not on one of the Jupyter repositories.
If it's a generic Python/Jupyter question, try other forums or discourse.jupyter.org.
If you are unsure, it's ok to post here, though, there are few ... | closed | 2023-03-12T20:15:57Z | 2023-03-30T08:29:51Z | https://github.com/ipython/ipython/issues/13970 | [
"bug",
"autosuggestions"
] | lukelbd | 3 |
nvbn/thefuck | python | 1,449 | Having a hard time getting set up on Ubuntu 24.04 | The output of `thefuck --version` (something like `The Fuck 3.1 using Python
3.5.0 and Bash 4.4.12(1)-release`):
```
pblanton@ThreadRipper:~$ thefuck --version
Traceback (most recent call last):
File "/home/pblanton/.local/bin/thefuck", line 5, in <module>
from thefuck.entrypoints.main import main
File... | open | 2024-05-22T19:03:56Z | 2024-09-20T14:07:08Z | https://github.com/nvbn/thefuck/issues/1449 | [] | pblanton | 18 |
dynaconf/dynaconf | fastapi | 207 | [RFC] Allow python.module.path on INCLUDES_FOR_DYNACONF | Dynaconf right now can load includes, the includes should be a list of paths like `['/path/to/file.yaml', '/path/to/glob/*.toml']`
**Problem**
It should also allow includes by `['python.module.paths'..]` instead of only `glob-able` paths.
**Describe the solution you'd like**
I want to be able to use:
```py... | closed | 2019-08-16T16:43:05Z | 2019-08-22T12:33:22Z | https://github.com/dynaconf/dynaconf/issues/207 | [
"in progress",
"Not a Bug",
"RFC"
] | rochacbruno | 0 |
allenai/allennlp | nlp | 5,024 | A google colab for the guide | **Is your feature request related to a problem? Please describe.**
(Continuing the discussion from here https://github.com/allenai/allennlp/issues/5017)
- While going through the guide, I felt that it would be a great addition if a google colab notebook can be created.
- As one begins to go through a new ML librar... | open | 2021-02-26T16:51:47Z | 2021-02-26T18:38:55Z | https://github.com/allenai/allennlp/issues/5024 | [
"Contributions welcome",
"Feature request"
] | ekdnam | 4 |
dropbox/sqlalchemy-stubs | sqlalchemy | 238 | Column comment type | Running the SQLAlchemy [`inspect`](https://docs.sqlalchemy.org/en/14/core/inspection.html#sqlalchemy.inspect) method on an existing MySQL server on a table with a column with an empty comment field is returning `comment=None` so I guess `comment` to be `Optional[str]` but mypy output complains that it needs to be `str`... | open | 2022-02-10T11:39:33Z | 2022-02-10T12:49:03Z | https://github.com/dropbox/sqlalchemy-stubs/issues/238 | [] | sdfordham | 0 |
sqlalchemy/alembic | sqlalchemy | 706 | Newly created Oracle Indexes are trying to be added as new. | How do I debug Alembic to find out why it is trying to add existing newly-created indexes?
I have a table defined as such:
```
class EditEOB(BASE):
'''This is the Edit EOB cross reference table.'''
__tablename__ = 'edit_eob'
__table_args__ = (Index('eob_edit_idx', 'eob_rid', 'edit_rid', 'relations... | closed | 2020-06-25T02:50:55Z | 2020-06-25T15:33:38Z | https://github.com/sqlalchemy/alembic/issues/706 | [
"bug",
"external SQLAlchemy issues",
"oracle"
] | davebyrnew | 14 |
jupyter-book/jupyter-book | jupyter | 1,554 | Add --builder singlehtml to docs? | Would a contribution to the docs that `--builder singlehtml` now exists and what it does be welcome? I spent a while looking for something like this and only discovered that it exists through @TomasBeuzen kindly pointing it out in the CHANGELOG. I am happy to put together a small PR if so. | closed | 2021-12-02T22:14:33Z | 2021-12-04T00:12:16Z | https://github.com/jupyter-book/jupyter-book/issues/1554 | [] | ttimbers | 1 |
marcomusy/vedo | numpy | 545 | did actor.scalar_colors disappear? | I have some old code that used actor.scalar_colors to specify the color for the vertices.
Workes fine in vedo 2021.0.3 but breaks in later versions. Has this been replaced by something else? | closed | 2021-11-23T14:54:32Z | 2021-11-23T15:53:19Z | https://github.com/marcomusy/vedo/issues/545 | [] | RubendeBruin | 4 |
pandas-dev/pandas | data-science | 61,123 | DOC: `read_excel` `nrows` parameter reads extra rows when tables are adjacent (no blank row) | ### Pandas version checks
- [x] I have checked that this issue has not already been reported.
- [x] I have confirmed this bug exists on the [latest version](https://pandas.pydata.org/docs/whatsnew/index.html) of pandas.
- [x] I have confirmed this bug exists on the [main branch](https://pandas.pydata.org/docs/dev/ge... | closed | 2025-03-14T21:30:43Z | 2025-03-19T20:37:24Z | https://github.com/pandas-dev/pandas/issues/61123 | [
"Docs",
"IO Excel",
"good first issue"
] | robertutterback | 1 |
litestar-org/litestar | pydantic | 3,737 | Bug: NameError: name 'BigInteger' is not defined | ### Description
subclassing
```python
class BigIntAuditBase(CommonTableAttributes, BigIntPrimaryKey, AuditColumns, DeclarativeBase):
"""Base for declarative models with BigInt primary keys and audit columns."""
registry = orm_registry
```
cause the mentioned error
### URL to code causing the ... | closed | 2024-09-14T12:41:10Z | 2025-03-20T15:54:55Z | https://github.com/litestar-org/litestar/issues/3737 | [
"Bug :bug:"
] | cbdiesse | 3 |
slackapi/bolt-python | fastapi | 583 | Is it possible to handle OAuth flow entirely with websockets? | Our team has an app that currently uses Flask and the OAuth flow to allow our users to set up incoming webhooks in a self-service fashion. We are trying to rewrite the app to use Socket Mode. Using OAuth flow seems to require a redirect URL, but the redirect URL requires HTTP/S. We verified that the SocketModeHandle... | closed | 2022-01-31T21:26:46Z | 2024-04-27T00:21:13Z | https://github.com/slackapi/bolt-python/issues/583 | [
"enhancement"
] | nimjor | 11 |
ageitgey/face_recognition | python | 1,364 | return np.linalg.norm(face_encodings - face_to_compare, axis=1) | * face_recognition version:
* Python version: 3.9
* Operating System: Windows 10
### Description
i was trying this code by sentdex in my compiler but i was getting a error, sentdex in his code have assigned [0] in this line
encoding = face_recognition.face_encodings(image)[0]
while my system is giving error l... | open | 2021-08-30T11:30:02Z | 2022-10-12T12:01:24Z | https://github.com/ageitgey/face_recognition/issues/1364 | [] | TanishqRajawat12 | 2 |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.