genes listlengths 393 5.45k | expressions listlengths 393 5.45k | drug stringclasses 95 values | sample stringclasses 96 values | BARCODE_SUB_LIB_ID stringlengths 19 19 | cell_line_id stringclasses 50 values | moa-fine stringclasses 15 values | canonical_smiles stringclasses 95 values | pubchem_cid stringclasses 95 values | plate stringclasses 1 value |
|---|---|---|---|---|---|---|---|---|---|
[
1,
5,
19,
21,
31,
56,
68,
77,
78,
85,
99,
100,
106,
107,
108,
112,
117,
127,
128,
130,
138,
151,
155,
156,
171,
174,
212,
214,
232,
233,
234,
235,
244,
246,
252,
257,
273,
275,
292,
298,
300,
302,
321,
326,
328,
334,
344,
... | [
-2,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
2,
1,
1,
1,
1,
2,
2,
2,
1,
1,
1,
1,
2,
2,
1,
1,
1,
2,
3,
2,
1,
1,
2,
1,
1,
2,
1,
1,
1,
3,
1,
1,
1,
1,
1,
1,
1,
1,
3,
1,
1,
1,
3,
1,
1,
1,
2,
3,
... | 8-Hydroxyquinoline | smp_1783 | 01_001_052-lib_1105 | CVCL_0480 | unclear | C1=CC2=C(C(=C1)O)N=CC=C2 | 1923.0 | plate4 |
[
1,
15,
19,
26,
32,
35,
38,
59,
70,
76,
78,
109,
117,
121,
136,
137,
144,
163,
164,
205,
206,
211,
221,
236,
246,
255,
257,
267,
270,
274,
290,
293,
300,
307,
312,
335,
342,
354,
383,
402,
412,
417,
427,
436,
441,
471,
474,
... | [
-2,
1,
1,
1,
1,
1,
1,
2,
1,
1,
2,
1,
1,
1,
1,
1,
1,
3,
1,
1,
1,
1,
1,
1,
2,
1,
3,
1,
1,
1,
1,
1,
3,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
2,
1,
1,
1,
1,
1,
2,
1,
1,
2,
1,
1,
1,
2,
... | 8-Hydroxyquinoline | smp_1783 | 01_001_105-lib_1105 | CVCL_0546 | unclear | C1=CC2=C(C(=C1)O)N=CC=C2 | 1923.0 | plate4 |
[
1,
10,
11,
19,
23,
26,
31,
35,
43,
45,
56,
58,
68,
70,
75,
76,
78,
82,
84,
89,
95,
103,
113,
114,
120,
137,
138,
139,
140,
146,
148,
151,
153,
154,
156,
163,
169,
174,
177,
184,
187,
202,
206,
210,
214,
217,
231,
234,
24... | [
-2,
1,
1,
2,
3,
1,
1,
1,
1,
2,
5,
1,
3,
1,
1,
1,
2,
1,
1,
1,
1,
3,
1,
1,
1,
1,
2,
1,
1,
1,
2,
3,
1,
1,
2,
1,
1,
2,
1,
1,
1,
1,
1,
2,
3,
1,
1,
4,
1,
1,
1,
2,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
... | 8-Hydroxyquinoline | smp_1783 | 01_001_165-lib_1105 | CVCL_1717 | unclear | C1=CC2=C(C(=C1)O)N=CC=C2 | 1923.0 | plate4 |
[
1,
20,
21,
31,
45,
56,
68,
69,
77,
88,
103,
104,
124,
138,
139,
140,
141,
142,
149,
174,
202,
210,
214,
226,
235,
259,
266,
267,
287,
290,
293,
307,
313,
332,
344,
354,
359,
363,
365,
368,
375,
378,
387,
390,
398,
404,
447,
... | [
-2,
1,
1,
1,
2,
1,
1,
2,
1,
1,
1,
1,
1,
3,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
2,
1,
1,
1,
1,
1,
1,
1,
1,
1,
2,
1,
1,
3,
1,
2,
1,
1,
1,
1,
1,
2,
4,
2,
2,
2,
1,
2,
1,
1,
1,
1,
1,
1,
2,
1,
1,
1,
1,
... | 8-Hydroxyquinoline | smp_1783 | 01_003_094-lib_1105 | CVCL_1717 | unclear | C1=CC2=C(C(=C1)O)N=CC=C2 | 1923.0 | plate4 |
[
1,
10,
19,
43,
56,
77,
86,
95,
103,
109,
112,
124,
128,
137,
151,
210,
214,
216,
220,
226,
233,
236,
290,
293,
298,
302,
307,
310,
362,
414,
429,
431,
436,
455,
459,
470,
475,
478,
483,
494,
521,
527,
528,
541,
545,
548,
549... | [
-2,
1,
1,
1,
1,
1,
2,
1,
1,
1,
1,
1,
2,
1,
2,
1,
2,
2,
1,
1,
2,
1,
1,
2,
2,
2,
2,
1,
1,
2,
1,
1,
4,
2,
1,
2,
1,
1,
2,
2,
1,
1,
1,
1,
1,
1,
1,
1,
3,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
3,
... | 8-Hydroxyquinoline | smp_1783 | 01_003_164-lib_1105 | CVCL_1056 | unclear | C1=CC2=C(C(=C1)O)N=CC=C2 | 1923.0 | plate4 |
[
1,
5,
19,
32,
43,
57,
77,
128,
139,
146,
154,
214,
227,
236,
252,
293,
299,
390,
413,
435,
439,
447,
455,
459,
472,
507,
519,
521,
529,
538,
548,
553,
563,
605,
609,
610,
611,
623,
638,
658,
726,
728,
740,
751,
770,
785,
841... | [
-2,
1,
1,
1,
2,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
5,
1,
1,
1,
1,
1,
1,
1,
1,
8,
1,
1,
1,
1,
2,
1,
1,
1,
1,
1,
1,
1,
6,
1,
1,
1,
1,
1,
1,
3,
2,
1,
1,
1,
1,
1,
1,
1,
1,
1,
2,
... | 8-Hydroxyquinoline | smp_1783 | 01_005_047-lib_1105 | CVCL_0131 | unclear | C1=CC2=C(C(=C1)O)N=CC=C2 | 1923.0 | plate4 |
[
1,
21,
22,
26,
32,
49,
112,
127,
131,
138,
139,
154,
212,
224,
234,
246,
252,
283,
293,
302,
324,
344,
353,
404,
418,
497,
515,
534,
553,
569,
605,
609,
610,
618,
670,
690,
714,
717,
731,
733,
736,
751,
783,
785,
793,
815,
8... | [
-2,
1,
1,
1,
1,
1,
3,
1,
1,
2,
2,
3,
1,
1,
1,
1,
1,
1,
1,
1,
1,
3,
1,
1,
1,
1,
1,
1,
3,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
3,
1,
1,
1,
2,
1,
2,
1,
5,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
2,
1,
1,
2,
... | 8-Hydroxyquinoline | smp_1783 | 01_005_055-lib_1105 | CVCL_0179 | unclear | C1=CC2=C(C(=C1)O)N=CC=C2 | 1923.0 | plate4 |
[
1,
11,
19,
22,
32,
42,
43,
56,
77,
103,
107,
112,
149,
164,
167,
224,
231,
233,
235,
245,
273,
280,
307,
321,
363,
382,
390,
403,
429,
434,
455,
459,
477,
495,
500,
501,
505,
509,
511,
521,
524,
549,
553,
554,
563,
576,
577,... | [
-2,
1,
1,
1,
3,
1,
2,
1,
1,
1,
1,
1,
1,
1,
1,
2,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
2,
1,
1,
3,
1,
3,
1,
2,
2,
1,
2,
1,
1,
1,
1,
2,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
... | 8-Hydroxyquinoline | smp_1783 | 01_005_087-lib_1105 | CVCL_1056 | unclear | C1=CC2=C(C(=C1)O)N=CC=C2 | 1923.0 | plate4 |
[
1,
21,
26,
59,
69,
70,
90,
95,
106,
108,
140,
190,
200,
203,
214,
220,
221,
231,
250,
266,
275,
284,
292,
299,
301,
303,
337,
344,
356,
378,
392,
404,
408,
412,
428,
431,
440,
450,
454,
455,
463,
477,
485,
490,
504,
505,
515... | [
-2,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
3,
1,
1,
1,
1,
1,
2,
1,
1,
1,
1,
1,
2,
7,
1,
1,
5,
2,
2,
1,
1,
2,
1,
1,
1,
1,
1,
2,
1,
1,
1,
1,
1,
1,
2,
1,
1,
4,
2,
1,
1,
4,
1,
1,
1,
4,
1,
1,
1,
... | 8-Hydroxyquinoline | smp_1783 | 01_006_070-lib_1105 | CVCL_1478 | unclear | C1=CC2=C(C(=C1)O)N=CC=C2 | 1923.0 | plate4 |
[
1,
5,
9,
11,
42,
43,
58,
68,
90,
95,
99,
103,
114,
139,
143,
157,
171,
187,
200,
203,
210,
214,
220,
233,
246,
257,
273,
278,
284,
287,
290,
311,
344,
351,
363,
375,
378,
380,
382,
392,
404,
415,
440,
459,
472,
505,
511,
5... | [
-2,
1,
1,
3,
1,
2,
1,
1,
1,
1,
1,
2,
1,
1,
1,
1,
1,
1,
1,
1,
2,
1,
1,
1,
1,
1,
1,
1,
1,
2,
1,
1,
5,
1,
1,
1,
1,
1,
1,
6,
1,
1,
1,
1,
1,
3,
1,
1,
1,
1,
1,
2,
1,
1,
2,
1,
1,
2,
1,
1,
2,
1,
2,
... | 8-Hydroxyquinoline | smp_1783 | 01_006_120-lib_1105 | CVCL_1478 | unclear | C1=CC2=C(C(=C1)O)N=CC=C2 | 1923.0 | plate4 |
[
1,
9,
19,
25,
26,
41,
42,
53,
103,
106,
108,
112,
114,
128,
144,
147,
160,
275,
299,
307,
344,
379,
382,
394,
396,
403,
431,
434,
477,
478,
505,
513,
515,
521,
529,
542,
549,
569,
577,
584,
595,
598,
605,
610,
612,
624,
638,... | [
-2,
2,
2,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
2,
4,
2,
2,
1,
5,
1,
1,
2,
2,
2,
1,
1,
2,
1,
2,
1,
1,
1,
1,
1,
2,
1,
1,
1,
2,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
3,
1,
3,
1,
1,
2,
1,
1,
1,
1,
... | 8-Hydroxyquinoline | smp_1783 | 01_006_154-lib_1105 | CVCL_1119 | unclear | C1=CC2=C(C(=C1)O)N=CC=C2 | 1923.0 | plate4 |
[
1,
15,
19,
21,
31,
32,
41,
54,
56,
69,
78,
103,
108,
112,
114,
136,
137,
139,
151,
167,
171,
182,
187,
203,
205,
206,
210,
212,
220,
231,
233,
234,
235,
236,
246,
252,
253,
255,
266,
270,
274,
275,
277,
290,
292,
293,
298,
... | [
-2,
1,
2,
2,
1,
3,
1,
1,
2,
3,
2,
1,
6,
2,
2,
1,
2,
1,
2,
1,
1,
1,
1,
1,
1,
1,
1,
2,
1,
2,
1,
4,
3,
2,
3,
1,
1,
2,
1,
1,
1,
1,
1,
1,
1,
1,
1,
2,
1,
1,
1,
2,
1,
1,
3,
1,
1,
1,
8,
1,
3,
1,
2,
... | 8-Hydroxyquinoline | smp_1783 | 01_007_068-lib_1105 | CVCL_0546 | unclear | C1=CC2=C(C(=C1)O)N=CC=C2 | 1923.0 | plate4 |
[
1,
5,
12,
19,
30,
31,
33,
58,
62,
70,
77,
95,
103,
108,
112,
124,
149,
151,
163,
182,
187,
195,
200,
202,
210,
220,
229,
234,
237,
246,
247,
249,
252,
275,
284,
292,
297,
299,
304,
311,
312,
319,
326,
334,
349,
353,
357,
3... | [
-2,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
2,
1,
1,
1,
1,
1,
3,
1,
2,
1,
1,
1,
1,
2,
1,
1,
1,
2,
1,
1,
1,
1,
2,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
2,
1,
1,
1,
1,
1,
1,
1,
1,
1,
1,
3,
1,
... | 8-Hydroxyquinoline | smp_1783 | 01_008_027-lib_1105 | CVCL_1666 | unclear | C1=CC2=C(C(=C1)O)N=CC=C2 | 1923.0 | plate4 |
Tahoe-100M
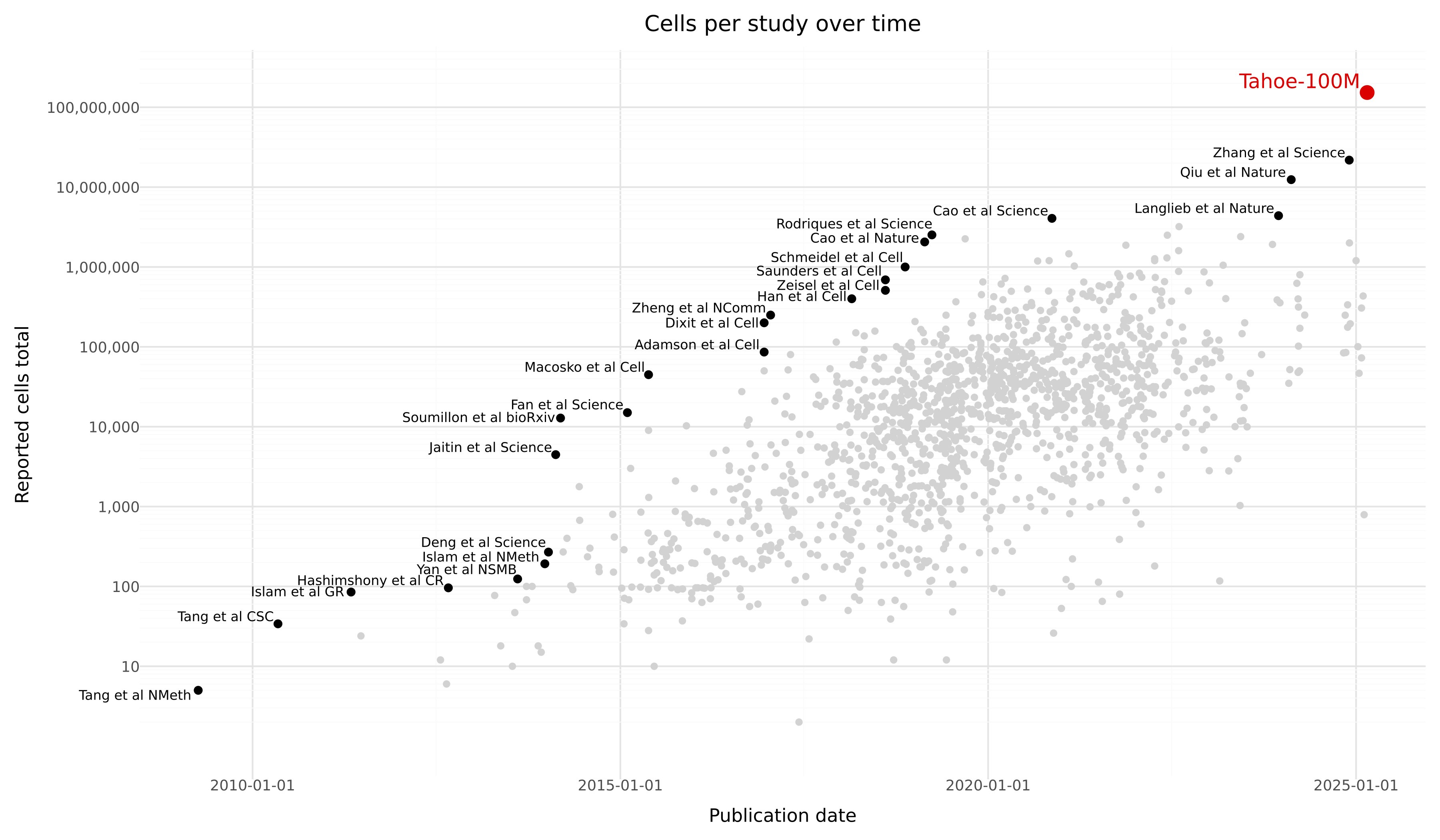
Tahoe-100M is a giga-scale single-cell perturbation atlas consisting of over 100 million transcriptomic profiles from 50 cancer cell lines exposed to 1,100 small-molecule perturbations. Generated using Vevo Therapeutics' Mosaic high-throughput platform, Tahoe-100M enables deep, context-aware exploration of gene function, cellular states, and drug responses at unprecedented scale and resolution. This dataset is designed to power the development of next-generation AI models of cell biology, offering broad applications across systems biology, drug discovery, and precision medicine.
Quickstart
from datasets import load_dataset
# Load dataset in streaming mode
ds = load_dataset("tahoebio/Tahoe-100m", streaming=True, split="train")
# View the first record
next(ds.iter(1))
Tutorials
Please refer to our tutorials for examples on using the data, accessing metadata tables and converting to/from the anndata format.
Please see the Data Loading Tutorial for a walkthrough on using the data.
| Notebook | URL | Colab |
|---|---|---|
| Loading the dataset from huggingface, accessing metadata, mapping to anndata | Link |
|
Community Resources
Here are a links to few resources created by the community. We would love to feature additional tutorials from the community, if you have built something on top of Tahoe-100M, please let us know and we would love to feature your work.
| Resource | Contributor | URL |
|---|---|---|
| Analysis guide for Tahoe-100M using rapids-single-cell, scanpy and dask | SCVERSE | Link |
| Tutorial for accessing Tahoe-100M h5ad files hosted by the Arc Institute | Arc Institute | Link |
Dataset Features
We provide multiple tables with the dataset including the main data (raw counts) in the expression_data table as well as
various metadata in the gene_metadata,sample_metadata,drug_metadata,cell_line_metadata,obs_metadata tables.
The main data can be downloaded as follows:
from datasets import load_dataset
tahoe_100m_ds = load_dataset("tahoebio/Tahoe-100M", streaming=True, split="train")
Setting stream=True instantiates an IterableDataset and prevents needing to
download the full dataset first. See tutorial for an end-to-end example.
The expression_data table has the following fields:
| Field Name | Type | Description |
|---|---|---|
genes |
sequence<int64> |
Gene identifiers (integer token IDs) corresponding to each gene with non-zero expression in the cell. This sequence aligns with the expressions field. The gene_metadata table can be used to map the token_IDs to gene_symbols or ensembl_IDs. The first entry for each row is just a marker token and should be ignored (See data-loading tutorial) |
expressions |
sequence<float32> |
Raw count values for each gene, aligned with the genes field. The first entry just marks a CLS token and should be ignored when parsing. |
drug |
string |
Name of the treatment. DMSO_TF marks vehicle controls, use DMSO_TF along with plate to get plate matched controls. |
sample |
string |
Unique identifier for the sample from which the cell was derived. Can be used to merge information from the sample_metadata table. Distinguishes replicate treatments. |
BARCODE_SUB_LIB_ID |
string |
Combination of barcode and sublibary identifiers. Unique for each cell in the dataset. Can be used as an index key when referencing to the obs_metadata table. |
cell_line_id |
string |
Unique identifier for the cancer cell line from which the cell originated. We use Cellosaurus IDs were, but additional identifiers such as DepMap IDs are provided in the cell_line_metadata table. |
moa-fine |
string |
Fine-grained mechanism of action (MOA) annotation for the drug, specifying the biological process or molecular target affected. Derived from MedChemExpress and curated with GPT-based annotations. |
canonical_smiles |
string |
Canonical SMILES (Simplified Molecular Input Line Entry System) string representing the molecular structure of the perturbing compound. |
pubchem_cid |
string |
PubChem Compound Identifier for the drug, allowing cross-referencing with public chemical databases. An empty string is used for DMSO controls. Please cast to int before querrying pubchem. |
plate |
string |
Identifier for the 96-well plate (1–14) in which the mixed-cell spheroid was seeded and treated. |
Additional metadata
Gene Metadata
gene_metadata = load_dataset("taheobio/Tahoe-100M","gene_metadata", split="train")
| Column Name | Description |
|---|---|
gene_symbol |
The HGNC-approved gene symbol corresponding to each gene (e.g., TP53, BRCA1). |
ensembl_id |
The Ensembl gene identifier (e.g., ENSG00000000003) based on Ensembl release 109 and genome build 38. |
token_id |
An integer token ID used to represent each gene. This is the ID used in the genes field in the main data. |
Sample Metadata
sample_metadata = load_dataset("tahoebio/Tahoe-100M","sample_metadata", split="train")
The sample_metadata has additional information for aggregate quality metrics for the sample as well as the concentration.
| Column Name | Description |
|---|---|
sample |
Unique identifier for the sample from which the cell was derived. Unique key for this table. |
plate |
Identifier (1–14) for the 96-well plate for the sample |
mean_gene_count |
Average number of unique genes detected per cell for the given sample. |
mean_tscp_count |
Average number of transcripts (UMIs) detected per cell in the sample. |
mean_mread_count |
Average number of reads per cell. |
mean_pcnt_mito |
Mean percentage of total reads that map to mitochondrial genes, across cells in the sample. |
drug |
Name of the treatment used to perturb the cells in the sample. |
drugname_drugconc |
String combining the compound name, concentration and concentration unit (e.g., [('8-Hydroxyquinoline',0.05,'uM')]), used to uniquely label each treatment condition. |
Drug Metadata
drug_metadata = load_dataset("tahoebio/Tahoe-100M","drug_metadata", split="train")
The drug_metadata has additional information about each treatment.
| Column Name | Description |
|---|---|
drug |
Name of the treatment used to perturb the cells in the sample. Unique key for this table |
targets |
List of gene symbols representing the known molecular targets of the compound. Targets were proposed by GPT-4o based on compound names and then validated against MedChemExpress information. |
moa-broad |
Broad classification of the compound’s mechanism of action (MOA), typically categorized as "inhibitor/antagonist," "activator/agonist," or "unclear." GPT-4o inferred this using compound target data and curated descriptions from MedChemExpress. |
moa-fine |
Specific functional annotation of the compound's MOA (e.g., "Proteasome inhibitor" or "MEK inhibitor"). These fine-grained labels were selected from a curated list of 25 MOA categories and assigned by GPT-4o with validation against compound descriptions. |
human-approved |
Indicates whether the compound is approved for human use ("yes" or "no"). GPT-4o provided these labels using prior knowledge and validation from public sources such as clinicaltrials.gov. |
clinical-trials |
Indicates whether the compound has been evaluated in any registered clinical trials ("yes" or "no"). Determined using GPT-4o and corroborated using clinicaltrials.gov searches. |
gpt-notes-approval |
Contextual notes generated by GPT-4o summarizing the compound’s approval status, common clinical usage, or nuances such as formulation-specific approvals. |
canonical_smiles |
The compound's SMILES (Simplified Molecular Input Line Entry System) representation, capturing its molecular structure as a text string. |
pubchem_cid |
The PubChem Compound Identifier (CID), a unique numerical ID linking the compound to its entry in the PubChem database. |
Cell Line Metadata
cell_line_metadata = load_dataset("tahoebio/Tahoe-100M","cell_line_metadata", split="train")
The cell-line metadata table has additional information about the key driver mutations for each cell line.
| Column Name | Description |
|---|---|
cell_name |
Standard name of the cancer cell line (e.g., A549). |
Cell_ID_DepMap |
Unique identifier for the cell line in the DepMap project (e.g., ACH-000681) |
Cell_ID_Cellosaur |
Cellosaurus accession ID (e.g., CVCL_0023). This is the ID used in the main dataset. |
Organ |
Tissue or organ of origin for the cell line (e.g., Lung), used to interpret lineage-specific responses and biological context. |
Driver_Gene_Symbol |
HGNC-approved symbol of a known or putative driver gene with functional alterations in this cell line (e.g., KRAS, CDKN2A). We report a curated list of driver mutations per cell-line. |
Driver_VarZyg |
Zygosity of the driver variant (e.g., Hom for homozygous, Het for heterozygous) |
Driver_VarType |
Type of genetic alteration (e.g., Missense, Frameshift, Stopgain, Deletion) |
Driver_ProtEffect_or_CdnaEffect |
Specific protein or cDNA-level annotation of the mutation (e.g., p.G12S, p.Q37), providing precise information on the variant’s consequence. |
Driver_Mech_InferDM |
Inferred functional mechanism of the mutation (e.g., LoF for loss-of-function, GoF for gain-of-function) |
Driver_GeneType_DM |
Classification of the driver gene as an Oncogene or Suppressor |
Citation
Please cite:
@article{zhang2025tahoe,
title={Tahoe-100M: A Giga-Scale Single-Cell Perturbation Atlas for Context-Dependent Gene Function and Cellular Modeling},
author={Zhang, Jesse and Ubas, Airol A and de Borja, Richard and Svensson, Valentine and Thomas, Nicole and Thakar, Neha and Lai, Ian and Winters, Aidan and Khan, Umair and Jones, Matthew G and others},
journal={bioRxiv},
pages={2025--02},
year={2025},
publisher={Cold Spring Harbor Laboratory}
}
- Downloads last month
- 2,822