code stringlengths 2.5k 150k | kind stringclasses 1 value |
|---|---|
# Launch Delay Model
Brian M. Gardner <bgardne1@stevens.edu>
Paul T. Grogan <pgrogan@stevens.edu>
## Introduction
This script performs statistical analysis and probabilistic modeling for launch delays using a [pre-processed data set](LaunchDelayDaya.ipynb).
Most of the required imports below are standard on typical Python 3 distributions; however, you may have to install tabulate separately (`!pip install tabulate`).
```
from IPython.display import HTML, display
from tabulate import tabulate
import numpy as np
import matplotlib.pyplot as plt
from scipy import integrate, optimize, stats
from datetime import datetime, timedelta
```
## Data Set
The data set includes lead times (time before planned launch) and delay times (time between planned and actual launches). Please see the [corresponding data notebook](LaunchDelayDaya.ipynb) for the source.
```
lead = np.array([108, 156, 71, 244, 131, 17, 273, 170, 144, 61, 23, 321, 230, 145, 119, 89, 38, 390, 257, 295, 205, 157, 84, 48, 8, 515, 382, 323, 233, 204, 108, 94, 22, 426, 376, 263, 196, 193, 169, 107, 71, 42, 508, 376, 443, 353, 312, 239, 163, 110, 67, 26, 509, 500, 387, 332, 291, 294, 163, 151, 115, 67, 8, 633, 285, 506, 612, 375, 302, 226, 216, 180, 132, 121, 47, 783, 650, 624, 534, 493, 420, 344, 345, 309, 261, 187, 113, 595, 462, 349, 259, 218, 145, 284, 213, 177, 326, 252, 178, 689, 921, 540, 450, 409, 285, 260, 272, 236, 188, 117, 237, 1032, 899, 901, 811, 770, 697, 621, 638, 602, 554, 480, 496, 873, 740, 702, 612, 206, 133, 422, 514, 370, 342, 268, 194, 998, 865, 1213, 757, 716, 643, 567, 586, 484, 436, 362, 288, 817, 684, 737, 647, 606, 533, 457, 471, 413, 382, 308, 404, 19, 960, 827, 936, 846, 805, 732, 656, 585, 549, 768, 694, 620, 80, 1148, 1025, 947, 857, 816, 743, 667, 704, 550, 502, 428, 358, 52, 16])
delay = np.array([0, 47, -1, 19, -1, 0, 149, 119, 32, 25, 22, 200, 158, 130, 66, 55, 33, 217, 217, 66, 66, 73, 73, 33, 2, 108, 108, 54, 54, 42, 65, 3, 4, 251, 168, 168, 145, 107, 58, 44, 9, 2, 206, 205, 25, 25, 25, 25, 25, 7, 14, 7, 279, 155, 155, 120, 120, 44, 99, 40, 40, 40, 25, 249, 464, 130, -66, 130, 130, 130, 69, 69, 69, 6, 6, 174, 174, 87, 87, 87, 87, 87, 15, 15, 15, 15, 15, 414, 414, 414, 414, 414, 414, 199, 199, 199, 2, 2, 2, 429, 64, 332, 332, 332, 383, 332, 249, 249, 249, 246, 52, 194, 194, 79, 79, 79, 79, 79, -9, -9, -9, -9, -99, 369, 369, 294, 294, 659, 659, 294, 131, 239, 219, 219, 219, 345, 345, -116, 250, 250, 250, 250, 160, 226, 226, 226, 226, 617, 617, 451, 451, 451, 451, 451, 366, 388, 371, 371, 201, 8, 527, 527, 305, 305, 305, 305, 305, 305, 305, 38, 38, 38, 0, 411, 401, 366, 366, 366, 366, 366, 258, 376, 376, 376, 372, 100, 14])
```
The analysis splits the data into three planning horizons:
* Near-term: lead time between 0 and 90 days.
* Intermediate: lead time between 90 and 180 days.
* Long-term: lead time between 180 and 360 days.
Descriptive statistics characterize the lead and delay times.
```
lead_90 = lead[lead <= 90]
delay_90 = delay[lead <= 90]
lead_180 = lead[np.logical_and(lead > 90, lead <= 180)]
delay_180 = delay[np.logical_and(lead > 90, lead <= 180)]
lead_360 = lead[np.logical_and(lead > 180, lead <= 360)]
delay_360 = delay[np.logical_and(lead > 180, lead <= 360)]
display(HTML(tabulate([
["Lead Time ($t$, days)", "Near-term", "Intermediate", "Long-term"],
["Count", f"{np.size(lead_90):.0f}", f"{np.size(lead_180):.0f}", f"{np.size(lead_360):.0f}"],
["Minimum", f"{np.min(lead_90):.0f}", f"{np.min(lead_180):.0f}", f"{np.min(lead_360):.0f}"],
["Maximum", f"{np.max(lead_90):.0f}", f"{np.max(lead_180):.0f}", f"{np.max(lead_360):.0f}"],
["Mean", f"{np.mean(lead_90):.1f}", f"{np.mean(lead_180):.1f}", f"{np.mean(lead_360):.1f}"],
["Std. Dev.", f"{np.std(lead_90, ddof=1):.1f}", f"{np.std(lead_180, ddof=1):.1f}", f"{np.std(lead_360, ddof=1):.1f}"]
], tablefmt='html')))
display(HTML(tabulate([
["Delay Time ($t$, days)", "Near-term", "Intermediate", "Long-term"],
["Count (Zero, %)",
f"{np.size(delay_90):.0f} ({np.sum(delay_90<=0):.0f}, {np.sum(delay_90<=0)/np.size(delay_90):.1%})",
f"{np.size(delay_180):.0f} ({np.sum(delay_180<=0):.0f}, {np.sum(delay_180<=0)/np.size(delay_180):.1%})",
f"{np.size(delay_360):.0f} ({np.sum(delay_360<=0):.0f}, {np.sum(delay_360<=0)/np.size(delay_360):.1%})"],
["Minimum", f"{np.min(delay_90):.0f}", f"{np.min(delay_180):.0f}", f"{np.min(delay_360):.0f}"],
["Maximum", f"{np.max(delay_90):.0f}", f"{np.max(delay_180):.0f}", f"{np.max(delay_360):.0f}"],
["Mean", f"{np.mean(delay_90):.1f}", f"{np.mean(delay_180):.1f}", f"{np.mean(delay_360):.1f}"],
["Std. Dev.", f"{np.std(delay_90, ddof=1):.1f}", f"{np.std(delay_180, ddof=1):.1f}", f"{np.std(delay_360, ddof=1):.1f}"]
], tablefmt='html')))
```
A scatter plot shows generally increasing delay time with increasing planning lead time.
```
plt.figure()
plt.semilogy(lead[lead<=360], delay[lead<=360], '.k')
plt.annotate('Short-term\n $x\leq 90$', (35,150), ha='center', rotation=0)
plt.axvline(x=90, ls='--', c='k')
plt.annotate('Intermediate\n$90 <x \leq 180$', (138,2.5), ha='center', rotation=0)
plt.axvline(x=180, ls='--', c='k')
plt.annotate('Long-term\n$180 <x\leq 360$', (250,4), ha='center', rotation=0)
plt.xlabel('Lead Time ($x$, days)')
plt.ylabel('Delay Time ($t$, days)')
plt.show()
```
And a cumulative distribution function (CDF) compares the delay distribution for the three planning horizons.
```
plt.figure()
t = np.linspace(0, np.round(np.max(delay), -2))
plt.step(t, [np.sum(delay_90 <= i)/np.size(delay_90) for i in t], '-k', where='post', label='Short-term ($x\leq 90$)')
plt.step(t, [np.sum(delay_180 <= i)/np.size(delay_180) for i in t], '--k', where='post', label='Intermediate ($90 < x \leq 180$)')
plt.step(t, [np.sum(delay_360 <= i)/np.size(delay_360) for i in t], ':k', where='post', label='Long-term ($180 < x \leq 360$)')
plt.xlabel('Delay, $t$ (days)')
plt.ylabel('Cumulative Relative Frequency, $F(t)$')
plt.legend()
plt.show()
```
## Maximum Likelihood Estimation
This model uses a hybrid Weibull distribution with a probability mass $p_0$ for zero delay and a probability density for positive delay based on the scale factor $\alpha$ and shape factor $\gamma$. The probability density function (PDF) $f(t)$ and cumulative distribution function (CDF) $F(t)$ resemble a standard Weibull distribution.
$$
f(t) = \left(1-p_0\right)\frac{\gamma}{\alpha} \left(\frac{t}{\alpha}\right)^{\gamma-1}e^{-\left(t/\alpha\right)^\gamma}, \quad t>0
$$
$$
F(t) = p_0 + \left(1-p_0\right) \left( 1-e^{-\left(t/\alpha\right)^\gamma}\right), \quad t \geq 0
$$
Maximum likelihood estimation (MLE) is a statistical inference procedure to fit model parameters using observed data. This method seeks to maximize the likelihood function (transformed using a logarithm for numerical efficiency) or, equivalently, minimize the minus log likelihood function.
The log likelihood function for the hybrid Weibull distribution is given as
$$
\log L\left(\lbrace t_i \rbrace, \gamma, \alpha\right) = \begin{cases}
\sum_{t_i} \log\left(f(t_i)\right) & N_0 = 0 \\
N_0\log\left(p_0\right) + \sum_{t_i>0} \log\left(f(t_i)\right) & N_0 > 0
\end{cases}
$$
where $N_0$ is the number of zero-delay observations, $N$ is the number of positive delay observations and $p_0=N_0/(N_0+N)$.
```
def hybrid_weibull_pdf(t, p_0, shape, scale):
"""
Computes the probability density function (PDF) for a hybrid Weibull distribution.
Args:
t (float): delay (days)
p_0 (float): probability mass for zero delay
shape: shape parameter (γ)
scale: scale parameter (α)
Returns:
float: the probability density
"""
return (1-p_0)*stats.weibull_min.pdf(t, shape, loc=0, scale=scale)
def hybrid_weibull_cdf(t, p_0, shape, scale):
"""
Computes the cumulative distribution function (CDF) for a hybrid Weibull distribution.
Args:
t (float): delay (days)
p_0 (float): probability mass for zero delay
shape: shape parameter (γ)
scale: scale parameter (α)
Returns:
float: the probability density
"""
return p_0*(t>=0) + (1-p_0)*stats.weibull_min.cdf(t, shape, loc=0, scale=scale)
def minus_log_likelihood_hybrid_weibull(params, obs):
"""
Compute the minus log likelihood function for a hybrid Weibull distribution.
Args:
params (:obj:`tuple`): The Weibull distribution parameters (shape, scale).
obs (:obj:`list`): The list of flight delay observations.
Returns:
float: the minus log likelihood value.
"""
# determine the number of zero-delay launches
N_0 = np.sum(obs<=0)
# compute the probability mass for zero-delay
p_0 = N_0/np.size(obs)
# compute the probability density for each positive delay
p_i = hybrid_weibull_pdf(obs[obs>0], p_0, params[0], params[1])
if np.any(p_i == 0):
# if any probability densities are zero, return an infinite value
return np.inf
elif N_0 > 0:
# if there are any zero-delay launches, return the hybrid minus log likelihood
return -N_0*np.log(p_0) - np.sum(np.log(p_i))
else:
# otherwise, return the simple minus log likelihood
return -np.sum(np.log(p_i))
```
The maximum likelihood method uses numerical optimization methods to minimize the minus log likelihood function.
```
def mle_hybrid_weibull(obs, params_0, bounds=None):
"""
Maximize the log likehood value for a hybrid Weibull distribution.
Args:
obs (:obj:`list`): The list of flight delay observations.
params_0 (:obj:`tuple`): The initial value of Weibull distribution parameters (shape, scale).
bounds (:obj:`tuple`, optional): The bounds for Weibull distribution parameters (shape, scale).
Returns:
(float, :obj:`tuple`): the probability mass (p_0) and distribution parameters (shape, scale).
"""
# determine the number of zero-delay launches
N_0 = np.sum(obs<=0)
# compute the probability mass for zero-delay
p_0 = N_0/np.size(obs)
# minimize the minus log likelihood function
res = optimize.minimize(minus_log_likelihood_hybrid_weibull, params_0, obs, method='SLSQP', bounds=bounds)
# return the probability mass and distribution parameters
return (p_0, res)
```
Finally, MLE is performed for each of the three planning horizons.
```
LL = np.zeros(3)
p_0_mle = np.zeros(3)
shape_mle = np.zeros(3)
scale_mle = np.zeros(3)
labels = ['Lead $\leq 90$', '$90 <$ Lead$ \leq 180$', '$180 <$ Lead$ \leq 360$']
for ii, obs in enumerate([delay_90, delay_180, delay_360]):
p_0_mle[ii], res = mle_hybrid_weibull(obs, (1, 30))
shape_mle[ii] = res.x[0]
scale_mle[ii] = res.x[1]
LL[ii] = -minus_log_likelihood_hybrid_weibull(res.x, obs)
plt.figure()
t = np.arange(0,np.max(obs)*1.5)
plt.title(labels[ii])
plt.step(t, [np.sum(obs <= i)/np.size(obs) for i in t], '-k', where='post', label='Observed')
plt.plot(t, hybrid_weibull_cdf(t, p_0_mle[ii], shape_mle[ii], scale_mle[ii]), ':k', label='MLE')
plt.xlabel('Delay Duration, $t$ (days)')
plt.ylabel('Cumulative Probability, $F(t)$')
plt.ylim([0,1])
plt.legend(loc='best')
plt.show()
display(HTML(tabulate([
["Horizon", "Log Likelihood", "Zero-delay p_0", "Shape (γ)", "Scale (α)"],
["Near-term", f"{LL[0]:.2f}", f"{p_0_mle[0]:.1%}", f"{shape_mle[0]:.2f}", f"{scale_mle[0]:.1f}"],
["Intermediate", f"{LL[1]:.2f}", f"{p_0_mle[1]:.1%}", f"{shape_mle[1]:.2f}", f"{scale_mle[1]:.1f}"],
["Long-term", f"{LL[2]:.2f}", f"{p_0_mle[2]:.1%}", f"{shape_mle[2]:.2f}", f"{scale_mle[2]:.1f}"]
], tablefmt='html')))
```
## Bootstrapping Analysis
Bootstrapping analysis performs repeated MLE samples by resampling (with replacement) from the source data set. It can help understand how model parameters are affected by small sample sizes.
```
N_SAMPLES = 100
p_0_mean = np.zeros(3)
shape_mean = np.zeros(3)
scale_mean = np.zeros(3)
table_labels = ['Short-term', 'Intermediate', 'Long-term']
labels = ['Short-term ($x\leq 90$)', 'Intermediate ($90 < x \leq 180$)', 'Long-term ($180 < x\leq 360$)']
for ii, obs in enumerate([delay_90, delay_180, delay_360]):
p_0 = np.zeros(N_SAMPLES)
shape = np.zeros(N_SAMPLES)
scale = np.zeros(N_SAMPLES)
np.random.seed(0)
for i in range(N_SAMPLES):
obs_bootstrap = np.random.choice(obs, size=len(obs), replace=True)
p_0[i], res = mle_hybrid_weibull(obs_bootstrap, (1, 30))
shape[i] = res.x[0]
scale[i] = res.x[1]
t = np.arange(0,np.max(obs)*1.5)
p, res = mle_hybrid_weibull(obs, (1, 30))
p_0_mean[ii] = p
shape_mean[ii] = res.x[0]
scale_mean[ii] = res.x[1]
plt.figure()
for i in range(N_SAMPLES):
F_t = p_0[i] + (1-p_0[i])*stats.weibull_min.cdf(t, shape[i], scale=scale[i])
plt.plot(t, F_t, '-', color='grey', alpha=0.01, label='Bootstrap' if i == 0 else None)
plt.step(t, [np.sum(obs <= i)/np.size(obs) for i in t], '-k', where='post', label='Empirical')
plt.plot(t, p + (1-p)*stats.weibull_min.cdf(t, res.x[0], scale=res.x[1]), ':k', label='MLE')
plt.xlabel('Delay Duration, $t$ (days)')
plt.ylabel('Cumulative Probability, $F(t)$')
plt.title(labels[ii])
leg = plt.legend()
for lh in leg.legendHandles:
lh.set_alpha(1)
plt.show()
display(HTML(tabulate([
[table_labels[ii], "Mean", "95% CI (Lower)", "95% CI (Upper)"],
["Zero Delay (p_0)", f"{p_0_mean[ii]:.1%}", f"{np.percentile(p_0, 5):.1%}", f"{np.percentile(p_0, 95):.1%}"],
["Shape (γ)", f"{shape_mean[ii]:.2f}", f"{np.percentile(shape, 5):.2f}", f"{np.percentile(shape, 95):.2f}"],
["Scale (α)", f"{scale_mean[ii]:.1f}", f"{np.percentile(scale, 5):.1f}", f"{np.percentile(scale, 95):.1f}"]
], tablefmt='html')))
fig, axes = plt.subplots(1,3,figsize=(12,3))
axes[0].hist(p_0, weights=np.ones_like(p_0)/np.size(p_0), color='k')
axes[0].set_xlabel('Zero Delay $P_0$')
axes[0].set_ylabel('Frequency Observed')
axes[1].hist(shape, weights=np.ones_like(shape)/np.size(shape), color='k')
axes[1].set_xlabel('Shape Parameter')
axes[1].set_ylabel('Frequency Observed')
axes[2].hist(scale, weights=np.ones_like(scale)/np.size(scale), color='k')
axes[2].set_xlabel('Scale Parameter')
axes[2].set_ylabel('Frequency Observed')
plt.suptitle(labels[ii])
plt.tight_layout()
plt.show()
```
## Quantization (Lloyd-Max Algorithm)
The Lloyd-Max algorithm can be used to convert the continuous distribution to a discrete distribution suitable for decision trees or other probabilistic scenario analysis. The method assumes $n$ quantization regions for discrete delays between $T_{min}$ and $T_{max}$ values. For example, one may start with evenly-spaced representation points $a_i$:
$$
a_i = T_{min} + \frac{ (T_{max}-T_{min})}{n} \cdot\frac{(2i-1)}{2}, \;\; 1\leq i \leq n
$$
Quantization region endpoints $b_i$ are defined by:
$$
b_i = \begin{cases} T_{min} & \mbox{if }i=0 \\ \frac{1}{2}\left(a_i+a_{i+1}\right) & \mbox{if } 0 < i < n \\ T_{max} & \mbox{if }i=n \end{cases}
$$
For a set of quantization regions ($a_i$) and region endpoints ($b_i$), the mean square error (MSE) is given by:
$$
MSE = \sum_{i=1}^{n} \int_{b_{i-1}}^{b_i} f(t)\cdot \left(t-a_i\right)^2 \, dt
$$
and updated representation points are computed by:
$$
a_i = \frac{\int_{b_{i-1}}^{b_i} t \cdot f(t) \, dt}{\int_{b_{i-1}}^{b_i} f(t) \, dt}
$$
The Lloyd-Max algorithm improves MSE by iteratively updating representation points until reaching a specfied convergence threshold.
The associated probability mass of each quantization region is found by integrating the continuous function:
$$
p(a_i) = \int_{b_{i-1}}^{b_i} f(t) \, dt
$$
```
def lloyd_max_hybrid_weibull(a, t_min, t_max, p_0, shape, scale):
# compute the interval bounds as the midpoints between each a, bounded by t_min and t_max
b = np.array(
[t_min] + [
(a[i]+a[i+1])/2
for i in range(len(a)-1)
] + [t_max]
)
# compute the equivalent probability mass function by numerically integrating the PDF over each interval
p_a = np.array([
integrate.quad(hybrid_weibull_pdf, b[i], b[i+1], args=(p_0, shape, scale))[0]
for i in range(len(a))
])
# define a helper function to compute the mean square error
def mse_fun(t, a, p_0, shape, scale):
return (t-a)**2*hybrid_weibull_pdf(t, p_0, shape, scale)
# compute the mean square error between the PDF and derived PMF
mse = np.sum([
integrate.quad(mse_fun, b[i], b[i+1], args=(a[i],p_0, shape, scale))[0]
for i in range(len(a))
])
# define a helper function to compute the numerator of the expected value
def num_fun(t, p_0, shape, scale):
return t*hybrid_weibull_pdf(t, p_0, shape, scale)
# compute the next values of a using the expected value for each region
next_a = np.array([
integrate.quad(num_fun, b[i], b[i+1], args=(p_0, shape, scale))[0]
/ integrate.quad(hybrid_weibull_pdf, b[i], b[i+1], args=(p_0, shape, scale))[0]
for i in range(len(a))
])
return b, p_a, mse, next_a
# define minimum and maximum delay values
t_min = 0
t_max = stats.weibull_min.ppf((0.999-p_0_mle[0])/(1-p_0_mle[0]), shape_mle[0], loc=0, scale=scale_mle[0])
# define number of quantization regions and compute initial representation points
n = 6
a_0 = [t_min + (t_max-t_min)/n*(2*i+1)/2 for i in range(n)]
# perform three iterations
b_0, p_a_0, mse_0, a_1 = lloyd_max_hybrid_weibull(a_0, t_min, t_max, p_0_mle[0], shape_mle[0], scale_mle[0])
b_1, p_a_1, mse_1, a_2 = lloyd_max_hybrid_weibull(a_1, t_min, t_max, p_0_mle[0], shape_mle[0], scale_mle[0])
b_2, p_a_2, mse_2, a_3 = lloyd_max_hybrid_weibull(a_2, t_min, t_max, p_0_mle[0], shape_mle[0], scale_mle[0])
display(HTML(tabulate([
["Iteration", "MSE", "$a_1$ ($p(a_1)$)", "$a_2$ ($p(a_2)$)", "$a_3$ ($p(a_3)$)", "$a_4$ ($p(a_4)$)", "$a_5$ ($p(a_5)$)", "$a_6$ ($p(a_6)$)"],
[0, f"{mse_0:.1f}", f"{a_0[0]:.1f} ({p_a_0[0]:.3f})", f"{a_0[1]:.1f} ({p_a_0[1]:.3f})", f"{a_0[2]:.1f} ({p_a_0[2]:.3f})", f"{a_0[3]:.1f} ({p_a_0[3]:.3f})", f"{a_0[4]:.1f} ({p_a_0[4]:.3f})", f"{a_0[5]:.1f} ({p_a_0[5]:.3f})"],
[1, f"{mse_1:.1f}", f"{a_1[0]:.1f} ({p_a_1[0]:.3f})", f"{a_1[1]:.1f} ({p_a_1[1]:.3f})", f"{a_1[2]:.1f} ({p_a_1[2]:.3f})", f"{a_1[3]:.1f} ({p_a_1[3]:.3f})", f"{a_1[4]:.1f} ({p_a_1[4]:.3f})", f"{a_1[5]:.1f} ({p_a_1[5]:.3f})"],
[2, f"{mse_2:.1f}", f"{a_2[0]:.1f} ({p_a_2[0]:.3f})", f"{a_2[1]:.1f} ({p_a_2[1]:.3f})", f"{a_2[2]:.1f} ({p_a_2[2]:.3f})", f"{a_2[3]:.1f} ({p_a_2[3]:.3f})", f"{a_2[4]:.1f} ({p_a_2[4]:.3f})", f"{a_2[5]:.1f} ({p_a_2[5]:.3f})"]
], tablefmt='html')))
t = np.linspace(t_min, t_max, 1000)
plt.figure()
plt.plot(t, hybrid_weibull_cdf(t, p_0_mle[0], shape_mle[0], scale_mle[0]), 'k', label='MLE')
plt.step(np.append([0], a_0), np.cumsum(np.append(p_0_mle[0], p_a_0)), where='post', label='Lloyd-Max Iter 0')
plt.step(np.append([0], a_1), np.cumsum(np.append(p_0_mle[0], p_a_1)), where='post', label='Lloyd-Max Iter 1')
plt.step(np.append([0], a_2), np.cumsum(np.append(p_0_mle[0], p_a_2)), where='post', label='Lloyd-Max Iter 2')
plt.xlabel('Delay Duration, $t$ (days)')
plt.ylabel('Cumulative Probability, $F(t)$')
plt.ylim([0,1])
plt.legend(loc='best')
plt.title('Short-term ($x\leq 90$)')
plt.show()
# array to record number of iterations, representation points, and probability masses
num_iter = np.zeros(3)
a = np.zeros((3, n))
p_a = np.zeros((3, n))
t_max = np.zeros(3)
# convergence criteria
delta_mse = 1e-2
for ii, horizon in enumerate(['Near-term', 'Intermediate', 'Long-term']):
t_min = 0
t_max[ii] = stats.weibull_min.ppf((0.999-p_0_mle[ii])/(1-p_0_mle[ii]), shape_mle[ii], loc=0, scale=scale_mle[ii])
a[ii,:] = [t_min + (t_max[ii]-t_min)/n*(2*i+1)/2 for i in range(n)]
prev_mse = mse = None
while prev_mse is None or mse is None or prev_mse-mse > delta_mse:
prev_mse = mse
num_iter[ii]+=1
b, p_a[ii,:], mse, a[ii,:] = lloyd_max_hybrid_weibull(a[ii,:], t_min, t_max[ii], p_0_mle[ii], shape_mle[ii], scale_mle[ii])
display(HTML(tabulate(
[
["Quantity", "Short-term", "Intermediate", "Long-term"],
["$T_{max}$", f"{t_max[0]:.0f}", f"{t_max[1]:.0f}", f"{t_max[2]:.0f}"],
["Iterations", f"{num_iter[0]:.0f}", f"{num_iter[1]:.0f}", f"{num_iter[2]:.0f}"],
["$i$", "$a_i$ ($p(a_i)$)", "$a_i$ ($p(a_i)$)", "$a_i$ ($p(a_i)$)"]
] + [
[i+1, f"{a[0,i]:.1f} ({p_a[0,i]:.3f})", f"{a[1,i]:.1f} ({p_a[1,i]:.3f})", f"{a[2,i]:.1f} ({p_a[2,i]:.3f})"]
for i in range(n)
], tablefmt='html'
)))
```
| github_jupyter |
# NuSVR with MinMaxScaler & Power Transformer
This Code template is for regression analysis using a Nu-Support Vector Regressor(NuSVR) based on the Support Vector Machine algorithm with PowerTransformer as Feature Transformation Technique and MinMaxScaler for Feature Scaling in a pipeline.
### Required Packages
```
import warnings
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as se
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import MinMaxScaler, PowerTransformer
from sklearn.model_selection import train_test_split
from sklearn.svm import NuSVR
from sklearn.metrics import r2_score, mean_absolute_error, mean_squared_error
warnings.filterwarnings('ignore')
```
### Initialization
Filepath of CSV file
```
file_path= ""
```
List of features which are required for model training .
```
features =[]
```
Target feature for prediction.
```
target=''
```
### Data Fetching
Pandas is an open-source, BSD-licensed library providing high-performance, easy-to-use data manipulation and data analysis tools.
We will use panda's library to read the CSV file using its storage path.And we use the head function to display the initial row or entry.
```
df=pd.read_csv(file_path)
df.head()
```
### Feature Selections
It is the process of reducing the number of input variables when developing a predictive model. Used to reduce the number of input variables to both reduce the computational cost of modelling and, in some cases, to improve the performance of the model.
We will assign all the required input features to X and target/outcome to Y.
```
X=df[features]
Y=df[target]
```
### Data Preprocessing
Since the majority of the machine learning models in the Sklearn library doesn't handle string category data and Null value, we have to explicitly remove or replace null values. The below snippet have functions, which removes the null value if any exists. And convert the string classes data in the datasets by encoding them to integer classes.
```
def NullClearner(df):
if(isinstance(df, pd.Series) and (df.dtype in ["float64","int64"])):
df.fillna(df.mean(),inplace=True)
return df
elif(isinstance(df, pd.Series)):
df.fillna(df.mode()[0],inplace=True)
return df
else:return df
def EncodeX(df):
return pd.get_dummies(df)
```
Calling preprocessing functions on the feature and target set.
```
x=X.columns.to_list()
for i in x:
X[i]=NullClearner(X[i])
Y=NullClearner(Y)
X=EncodeX(X)
X.head()
```
#### Correlation Map
In order to check the correlation between the features, we will plot a correlation matrix. It is effective in summarizing a large amount of data where the goal is to see patterns.
```
f,ax = plt.subplots(figsize=(18, 18))
matrix = np.triu(X.corr())
se.heatmap(X.corr(), annot=True, linewidths=.5, fmt= '.1f',ax=ax, mask=matrix)
plt.show()
```
### Data Splitting
The train-test split is a procedure for evaluating the performance of an algorithm. The procedure involves taking a dataset and dividing it into two subsets. The first subset is utilized to fit/train the model. The second subset is used for prediction. The main motive is to estimate the performance of the model on new data.
```
X_train,X_test,y_train,y_test=train_test_split(X,Y,test_size=0.2,random_state=123)
```
### Model
Support vector machines (SVMs) are a set of supervised learning methods used for classification, regression and outliers detection.
A Support Vector Machine is a discriminative classifier formally defined by a separating hyperplane. In other terms, for a given known/labelled data points, the SVM outputs an appropriate hyperplane that classifies the inputted new cases based on the hyperplane. In 2-Dimensional space, this hyperplane is a line separating a plane into two segments where each class or group occupied on either side.
Here we will use NuSVR, the NuSVR implementation is based on libsvm. Similar to NuSVC, for regression, uses a parameter nu to control the number of support vectors. However, unlike NuSVC, where nu replaces C, here nu replaces the parameter epsilon of epsilon-SVR.
#### Model Tuning Parameters
1. nu : float, default=0.5
> An upper bound on the fraction of training errors and a lower bound of the fraction of support vectors. Should be in the interval (0, 1]. By default 0.5 will be taken.
2. C : float, default=1.0
> Regularization parameter. The strength of the regularization is inversely proportional to C. Must be strictly positive. The penalty is a squared l2 penalty.
3. kernel : {‘linear’, ‘poly’, ‘rbf’, ‘sigmoid’, ‘precomputed’}, default=’rbf’
> Specifies the kernel type to be used in the algorithm. It must be one of ‘linear’, ‘poly’, ‘rbf’, ‘sigmoid’, ‘precomputed’ or a callable. If none is given, ‘rbf’ will be used. If a callable is given it is used to pre-compute the kernel matrix from data matrices; that matrix should be an array of shape (n_samples, n_samples).
4. gamma : {‘scale’, ‘auto’} or float, default=’scale’
> Gamma is a hyperparameter that we have to set before the training model. Gamma decides how much curvature we want in a decision boundary.
5. degree : int, default=3
> Degree of the polynomial kernel function (‘poly’). Ignored by all other kernels.Using degree 1 is similar to using a linear kernel. Also, increasing degree parameter leads to higher training times.
#### Rescaling technique
MinMax Scaler scales the data between 0 and 1.Minimum is subtracted from all values – thereby marking a scale from Min to Max. Then it is divided by the difference between Min and Max.
Refer [API](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.MinMaxScaler.html) for the parameters
#### Feature Transformation
Power transforms are a family of parametric, monotonic transformations that are applied to make data more Gaussian-like. This is useful for modeling issues related to heteroscedasticity (non-constant variance), or other situations where normality is desired.
```
model=make_pipeline(MinMaxScaler(),PowerTransformer(),NuSVR())
model.fit(X_train,y_train)
```
#### Model Accuracy
We will use the trained model to make a prediction on the test set.Then use the predicted value for measuring the accuracy of our model.
> **score**: The **score** function returns the coefficient of determination <code>R<sup>2</sup></code> of the prediction.
```
print("Accuracy score {:.2f} %\n".format(model.score(X_test,y_test)*100))
```
> **r2_score**: The **r2_score** function computes the percentage variablility explained by our model, either the fraction or the count of correct predictions.
> **mae**: The **mean abosolute error** function calculates the amount of total error(absolute average distance between the real data and the predicted data) by our model.
> **mse**: The **mean squared error** function squares the error(penalizes the model for large errors) by our model.
```
y_pred=model.predict(X_test)
print("R2 Score: {:.2f} %".format(r2_score(y_test,y_pred)*100))
print("Mean Absolute Error {:.2f}".format(mean_absolute_error(y_test,y_pred)))
print("Mean Squared Error {:.2f}".format(mean_squared_error(y_test,y_pred)))
```
#### Prediction Plot
First, we make use of a scatter plot to plot the actual observations, with x_train on the x-axis and y_train on the y-axis.
For the regression line, we will use x_train on the x-axis and then the predictions of the x_train observations on the y-axis.
```
n=len(X_test) if len(X_test)<20 else 20
plt.figure(figsize=(14,10))
plt.plot(range(n),y_test[0:n], color = "green")
plt.plot(range(n),model.predict(X_test[0:n]), color = "red")
plt.legend(["Actual","prediction"])
plt.title("Predicted vs True Value")
plt.xlabel("Record number")
plt.ylabel(target)
plt.show()
```
#### Creator: Aishwarya Guntoju , Github: [Profile](https://github.com/DSAishwaryaG)
| github_jupyter |
```
import numpy as np
import librosa
import os
from matplotlib import pyplot as plt
!pip install pretty_midi
import pretty_midi
pretty_midi.pretty_midi.MAX_TICK = 1e10
#preparing the data from the original audio files
entries = os.scandir('/content/drive/MyDrive/train_source')#loading the data to process which we are going to use for training
for file in entries:
file_name,file_extensions=os.path.splitext(file)
if file_extensions=='.wav':
y, sr = librosa.load(file)
n_fft=2048
hop_length=512
stft=librosa.stft(y, n_fft=n_fft, hop_length=hop_length)#using the librosa library to calculate the short time fourier transform of the audio file which is returned as a 2D vector
c=len(y) # it is being stored because it will be used for the frequency while making the piano roll file
stft=np.abs(stft)# the returned 2d vector is a complex quantity
stft=librosa.amplitude_to_db(stft)# taking the amplitude on logarithmic scale gives a better visualization of data
b=np.shape(stft)[1]
out = '/content/drive/MyDrive/trash/wav_'+str(file_name[40:])+'.npy'
entries1 = os.scandir('/content/drive/MyDrive/train_source')#it is being loaded again so that we can get the corresponding midi file to the wav file which we have just used
for file1 in entries1:
file_name1,file_extensions1=os.path.splitext(file1)
if file_name1==file_name and file_extensions1==".mid":
file2=file_name + file_extensions1
pm = pretty_midi.PrettyMIDI(file2)
a=pm.instruments[0].get_piano_roll(float(sr*float(b)/float(c)))# The frequency has been taken such that its shape and the stft files shape is same
a=np.append(np.transpose(a), np.zeros((b-np.shape(a)[1],128)), axis=0)# there is a slight difference between the length of the stft data and midi data , to make the number of features same for training a padding of few row is added
a=np.transpose(a)
out = '/content/drive/MyDrive/trash/mid_'+str(file_name1[40:])+'mid'+'.npy'
np.save(out,a)
path='/content/drive/MyDrive/trainfiles/trainwav'
def concatenate_wav(path):#we are dividing the number of features of the stft file into batches of 100 and stacking them up to create a 3D array
entries = os.scandir(path)
f=0
d=np.zeros((1,252,100))#initialising the array with the correct shape
for file in entries:
a=np.load(file)
a=a[np.newaxis,:,:]# adding a new axis to the original files
for b in range(int(np.shape(a)[2]/100)):
c=a[:,:,b*100:(b*100)+100]# dividing them into batches of 100
d=np.concatenate((d,c), axis=0)# stacking them up on the first axis
out = '/content/drive/MyDrive/trainfiles/trainwav/concatenate'+'wav'+'.npy'
d=d[1:,:,:]# array was initialise with zeros so excluding that part
d=np.transpose(d,(0,2,1))# taking transpose so that the the second and third dimensions are interchanged
np.save(out,d)
concatenate_wav(path)
path='/content/drive/MyDrive/trainfiles/trainmid'
def concatenate_mid(path):#we are dividing the number of features of the pianoroll file into batches of 100 and stacking them up to create a 3D array
entries = os.scandir(path)
f=0
d=np.zeros((1,88,100))
for file in entries:
a=np.load(file)
f=f+np.shape(a)[1]
a=a[np.newaxis,20:108,:]
for b in range(int(np.shape(a)[2]/100)):
c=a[:,:,b*100:(b*100)+100]
d=np.concatenate((d,c), axis=0)
out = '/content/drive/MyDrive/trainlabels/concatenate'+'mid'+'.npy'
print(f)
print(np.shape(d)[0]*100)
d=d[1:,:,:]
d=np.transpose(d,(0,2,1))
np.save(out,d)
concatenate_mid(path)
#for the test files
entries = os.scandir('/content/drive/MyDrive/test_source')
os.mkdir('/content/drive/MyDrive/testfiles')
os.mkdir('/content/drive/MyDrive/testfiles/testwav')
os.mkdir('/content/drive/MyDrive/testfiles/testmid')
for file in entries:
file_name,file_extensions=os.path.splitext(file)
if file_extensions=='.wav':
y, sr = librosa.load(file)
n_fft=2048
hop_length=512
sr=16000
stft=librosa.stft(y, n_fft=n_fft, hop_length=hop_length)
c=len(y)
stft=np.abs(stft)
stft=librosa.amplitude_to_db(stft)
b=np.shape(stft)[1]
out = '/content/drive/MyDrive/trainfiles/trainwav/wav_'+str(file_name[40:])+'.npy'
np.save(out,stft)
print('wrote file', out)
entries1 = os.scandir('/content/drive/MyDrive/train_source')
for file1 in entries1:
file_name1,file_extensions1=os.path.splitext(file1)
if file_name1==file_name and file_extensions1==".mid":
file2=file_name + file_extensions1
pm = pretty_midi.PrettyMIDI(file2)
a=pm.instruments[0].get_piano_roll(float(sr*float(b)/float(c)))
a=np.append(np.transpose(a), np.zeros((b-np.shape(a)[1],128)), axis=0)
a=np.transpose(a)
out = '/content/drive/MyDrive/trainfiles/trainmid/mid_'+str(file_name1[40:])+'mid'+'.npy'
np.save(out,a)
print("saving", out)
path='/content/drive/MyDrive/testfiles/testwav'
def concatenate_wav(path):
entries = os.scandir(path)
f=0
d=np.zeros((1,252,100))
for file in entries:
a=np.load(file)
f=f+np.shape(a)[1]
a=a[np.newaxis,:,:]
for b in range(int(np.shape(a)[2]/100)):
c=a[:,:,b*100:(b*100)+100]
d=np.concatenate((d,c), axis=0)
out = '/content/drive/MyDrive/testfiles/testwav/concatenate'+'wav'+'.npy'
print(f)
print(np.shape(d)[0]*100)
d=d[1:,:,:]
d=np.transpose(d,(0,2,1))
np.save(out,d)
concatenate_wav(path)
path='/content/drive/MyDrive/testfiles/testmid'
def concatenate_mid(path):
entries = os.scandir(path)
f=0
d=np.zeros((1,88,100))
for file in entries:
a=np.load(file)
f=f+np.shape(a)[1]
a=a[np.newaxis,20:108,:]
for b in range(int(np.shape(a)[2]/100)):
c=a[:,:,b*100:(b*100)+100]
d=np.concatenate((d,c), axis=0)
out = '/content/drive/MyDrive/testfiles/testmid/concatenate'+'mid'+'.npy'
print(f)
print(np.shape(d)[0]*100)
d=d[1:,:,:]
d=np.transpose(d,(0,2,1))
np.save(out,d)
concatenate_mid(path)
```
| github_jupyter |
# Neural style transfer
**Modified by:** [Sergio Nieto](https://twitter.com/SergioSNieto)<br>
**Original:** [fchollet](https://twitter.com/fchollet)<br>
**Date created:** 2016/01/11<br>
**Last modified:** 2021/06/25<br>
**Description:** Transfering the style of a reference image to target image using gradient descent.
## Introduction
Style transfer consists in generating an image
with the same "content" as a base image, but with the
"style" of a different picture (typically artistic).
This is achieved through the optimization of a loss function
that has 3 components: "style loss", "content loss",
and "total variation loss":
- The total variation loss imposes local spatial continuity between
the pixels of the combination image, giving it visual coherence.
- The style loss is where the deep learning keeps in --that one is defined
using a deep convolutional neural network. Precisely, it consists in a sum of
L2 distances between the Gram matrices of the representations of
the base image and the style reference image, extracted from
different layers of a convnet (trained on ImageNet). The general idea
is to capture color/texture information at different spatial
scales (fairly large scales --defined by the depth of the layer considered).
- The content loss is a L2 distance between the features of the base
image (extracted from a deep layer) and the features of the combination image,
keeping the generated image close enough to the original one.
**Reference:** [A Neural Algorithm of Artistic Style](
http://arxiv.org/abs/1508.06576)
## Setup
```
import numpy as np
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras.applications import vgg19
base_image_path = keras.preprocessing.image.load_img("./base_image.jpg")
#keras.utils.get_file("base_image.jpg")
style_reference_image_path = keras.utils.get_file(
"EaG5URr.jpg", "https://i.imgur.com/EaG5URr.jpg"
)
result_prefix = "paris_generated"
# Weights of the different loss components
total_variation_weight = 1e-6
style_weight = 1e-6
content_weight = 2.5e-8
# Dimensions of the generated picture.
width, height = base_image_path.size#keras.preprocessing.image.load_img(base_image_path).size
img_nrows = 400
img_ncols = int(width * img_nrows / height)
```
## Let's take a look at our base (content) image and our style reference image
```
from IPython.display import Image, display
display(Image(base_image_path))
display(Image(style_reference_image_path))
display(Image("base_image.jpg"))
```
## Image preprocessing / deprocessing utilities
```
def preprocess_image(image_path):
# Util function to open, resize and format pictures into appropriate tensors
img = keras.preprocessing.image.load_img(
image_path, target_size=(img_nrows, img_ncols)
)
img = keras.preprocessing.image.img_to_array(img)
img = np.expand_dims(img, axis=0)
img = vgg19.preprocess_input(img)
return tf.convert_to_tensor(img)
def preprocess_image_nop():
# Util function to open, resize and format pictures into appropriate tensors
# Change to use base image in local folder
#base_image_path = keras.preprocessing.image.load_img("./base_image.jpg")
img = keras.preprocessing.image.load_img(
"./base_image.jpg", target_size=(img_nrows, img_ncols)
)
img = keras.preprocessing.image.img_to_array(img)
img = np.expand_dims(img, axis=0)
img = vgg19.preprocess_input(img)
return tf.convert_to_tensor(img)
def deprocess_image(x):
# Util function to convert a tensor into a valid image
x = x.reshape((img_nrows, img_ncols, 3))
# Remove zero-center by mean pixel
x[:, :, 0] += 103.939
x[:, :, 1] += 116.779
x[:, :, 2] += 123.68
# 'BGR'->'RGB'
x = x[:, :, ::-1]
x = np.clip(x, 0, 255).astype("uint8")
return x
```
## Compute the style transfer loss
First, we need to define 4 utility functions:
- `gram_matrix` (used to compute the style loss)
- The `style_loss` function, which keeps the generated image close to the local textures
of the style reference image
- The `content_loss` function, which keeps the high-level representation of the
generated image close to that of the base image
- The `total_variation_loss` function, a regularization loss which keeps the generated
image locally-coherent
```
# The gram matrix of an image tensor (feature-wise outer product)
def gram_matrix(x):
x = tf.transpose(x, (2, 0, 1))
features = tf.reshape(x, (tf.shape(x)[0], -1))
gram = tf.matmul(features, tf.transpose(features))
return gram
# The "style loss" is designed to maintain
# the style of the reference image in the generated image.
# It is based on the gram matrices (which capture style) of
# feature maps from the style reference image
# and from the generated image
def style_loss(style, combination):
S = gram_matrix(style)
C = gram_matrix(combination)
channels = 3
size = img_nrows * img_ncols
return tf.reduce_sum(tf.square(S - C)) / (4.0 * (channels ** 2) * (size ** 2))
# An auxiliary loss function
# designed to maintain the "content" of the
# base image in the generated image
def content_loss(base, combination):
return tf.reduce_sum(tf.square(combination - base))
# The 3rd loss function, total variation loss,
# designed to keep the generated image locally coherent
def total_variation_loss(x):
a = tf.square(
x[:, : img_nrows - 1, : img_ncols - 1, :] - x[:, 1:, : img_ncols - 1, :]
)
b = tf.square(
x[:, : img_nrows - 1, : img_ncols - 1, :] - x[:, : img_nrows - 1, 1:, :]
)
return tf.reduce_sum(tf.pow(a + b, 1.25))
```
Next, let's create a feature extraction model that retrieves the intermediate activations
of VGG19 (as a dict, by name).
```
# Build a VGG19 model loaded with pre-trained ImageNet weights
model = vgg19.VGG19(weights="imagenet", include_top=False)
# Get the symbolic outputs of each "key" layer (we gave them unique names).
outputs_dict = dict([(layer.name, layer.output) for layer in model.layers])
# Set up a model that returns the activation values for every layer in
# VGG19 (as a dict).
feature_extractor = keras.Model(inputs=model.inputs, outputs=outputs_dict)
```
Finally, here's the code that computes the style transfer loss.
```
# List of layers to use for the style loss.
style_layer_names = [
"block1_conv1",
"block2_conv1",
"block3_conv1",
"block4_conv1",
"block5_conv1",
]
# The layer to use for the content loss.
content_layer_name = "block5_conv2"
def compute_loss(combination_image, base_image, style_reference_image):
input_tensor = tf.concat(
[base_image, style_reference_image, combination_image], axis=0
)
features = feature_extractor(input_tensor)
# Initialize the loss
loss = tf.zeros(shape=())
# Add content loss
layer_features = features[content_layer_name]
base_image_features = layer_features[0, :, :, :]
combination_features = layer_features[2, :, :, :]
loss = loss + content_weight * content_loss(
base_image_features, combination_features
)
# Add style loss
for layer_name in style_layer_names:
layer_features = features[layer_name]
style_reference_features = layer_features[1, :, :, :]
combination_features = layer_features[2, :, :, :]
sl = style_loss(style_reference_features, combination_features)
loss += (style_weight / len(style_layer_names)) * sl
# Add total variation loss
loss += total_variation_weight * total_variation_loss(combination_image)
return loss
```
## Add a tf.function decorator to loss & gradient computation
To compile it, and thus make it fast.
```
@tf.function
def compute_loss_and_grads(combination_image, base_image, style_reference_image):
with tf.GradientTape() as tape:
loss = compute_loss(combination_image, base_image, style_reference_image)
grads = tape.gradient(loss, combination_image)
return loss, grads
```
## The training loop
Repeatedly run vanilla gradient descent steps to minimize the loss, and save the
resulting image every 100 iterations.
We decay the learning rate by 0.96 every 100 steps.
```
optimizer = keras.optimizers.SGD(
keras.optimizers.schedules.ExponentialDecay(
initial_learning_rate=100.0, decay_steps=100, decay_rate=0.96
)
)
#base_image_path
base_image = preprocess_image_nop()
style_reference_image = preprocess_image(style_reference_image_path)
combination_image = tf.Variable(preprocess_image_nop())
iterations = 4000
for i in range(1, iterations + 1):
loss, grads = compute_loss_and_grads(
combination_image, base_image, style_reference_image
)
optimizer.apply_gradients([(grads, combination_image)])
if i % 100 == 0:
print("Iteration %d: loss=%.2f" % (i, loss))
img = deprocess_image(combination_image.numpy())
fname = result_prefix + "_at_iteration_%d.png" % i
keras.preprocessing.image.save_img(fname, img)
```
After 4000 iterations, you get the following result:
```
display(Image(result_prefix + "_at_iteration_1000.png"))
```
| github_jupyter |
What is the 10-digit integer whose digits are all distinct and the number formed by the first i of them is divisible
by i for each i from 1 to 10?
```
# Student 1 using first principles
import random
# import sys
even = [2,4,6,8]
odd = [1,3,5,7,9]
for i in range(999999):
a = random.choice(odd)
b = random.choice(even)
c = random.choice(odd)
d = random.choice(even)
e = 5
f = random.choice(even)
g = random.choice(odd)
h = random.choice(even)
i = random.choice(odd)
j = 0
if (a != b != c != d != e != f != g != h != i != j and
a != c != e != g != i and a != d != g != j and
a != e != i and a != f and a != g and a != h and
a != i and a != j and b != d != f != h != j and
b != e != h and b != f != j and b != g and b != h and
b != i and b != j and c != f != i and c != g and
c != h and c != i and c != j and d != h and d != i and
d != j and e != j):
num = (a * 10 ** 9 + b * 10 ** 8 + c * 10 ** 7 +
d * 10 ** 6 + e * 10 ** 5 + f * 10 ** 4 +
g * 10 ** 3 + h * 10 ** 2 + i * 10 ** 1 + j)
if ((a * 10 + b) % 2 == 0 and (a * 100 + b * 10 + c) % 3 == 0 and
(a * 1000 + b * 100 + c * 10 + d) % 4 == 0 and
(a * 10000 + b * 1000 + c * 100 + d * 10 + e) % 5 == 0
and (a * 100000 + b * 10000 + c * 1000 + d * 100 + e * 10 + f) % 6 == 0
and (a * 10 ** 6 + b * 100000 + c * 10000 + d * 1000 + e * 100 + f * 10 + g) % 7 == 0
and (a * 10 ** 7 + b * 10 ** 6 + c * 100000 + d * 10000 + e * 1000 + f * 100 + g * 10 + h) % 8 == 0
and (a * 10 ** 8 + b * 10 ** 7 + c * 10 ** 6 + d * 100000 + e * 10000 + f * 1000 + g * 100 + h * 10 + i) % 9 == 0
and (a * 10 ** 9 + b * 10 ** 8 + c * 10 ** 7 + d * 10 ** 6 + e * 100000 + f * 10000 + g * 1000 + h * 100 + i * 10 + j) % 10 == 0):
print ("The 10-digit number is %s." %num)
break # sys.exit()
else:
None
else:
None
```
**Comments**
* The 2 else are redundant. If can stand on its own (i.e. else part is optional).
* Please include comments. :)
* Removed sys as this does not work well in a web browser environment. replaced with break.
```
# Student 2 using itertools
import time
start = time.time()
import itertools
digits = ['1','2','3','4','5','6','7','8','9','0']
permutation = itertools.permutations(digits)
all_nums = [''.join(item[0:10]) for item in permutation]
for i in range (10,1,-1):
number = [x for x in all_nums if int(x[0:i])%i==0]
all_nums = number
elapsed=time.time()-start
print("result", int(''.join(number)), "returned in", elapsed, "seconds")
```
** Comments **
* Consider grouping import statements at the start for logical separation of different aspects of code.
* Good to have time profiling to 'measure' efficiency.
* Comments will be helpful to explain what you are trying to achieve in list comprehension.
```
# Alternative approach 1 - brute force through all possibilities
from itertools import permutations
for digits in permutations("1234567890"):
number = "".join(digits)
for i in range(1, 11):
if int(number[:i]) % i:
break
else:
print(number)
break
# Alternative approach 2 - since number will be multiple of 10, treat as separate case to improve efficiency
from itertools import permutations
for digits in permutations("123456789"):
number = "".join(digits)
for i in range(1, 10):
if int(number[:i]) % i:
break
else:
print(number + "0")
break
# Alternative approach 3 - separate even and odd numbers to further improve efficiency
from itertools import permutations
for evens in permutations("2468"):
for odds in permutations("1379"):
number = odds[0]+evens[0]+odds[1]+evens[1]+'5'+evens[2]+odds[2]+evens[3]+odds[3]+'0'
for i in range(1, 10):
if int(number[:i]) % i:
break
else:
print(number)
break
```
Comments? Email buildingblocs@computing.sg :)
| github_jupyter |
# Misc tests used for evaluating how well RSSI translates to distance
Note - this notebook still needs to be cleaned. We include it here so this work won't be lost
```
%matplotlib inline
import pandas as pd
import matplotlib
import matplotlib.pyplot as plt
onemeter_file_path = '../data/rssi_distance/expt4/expt_07_11_'
data = pd.read_csv(onemeter_file_path+'1m.dat', skiprows=1, skipfooter=1, parse_dates=True, sep=' ', header=None, names=['MAC', 'RSSI', 'TIME'])
macs = ['D9:CB:6E:1F:48:82,', 'F2:50:9D:7E:C8:0C,']
signal_strength_1m = []
for mac in macs:
signal_strength_1m.append(pd.rolling_mean(data[data['MAC'] == mac]['RSSI'], 5, min_periods=4))
# Get a threshold value of signal strength for inide and outside the case based on an accuracy level
def getThresholds(accuracy, arr_vals):
result = []
for i in xrange(len(arr_vals)):
sample_num = int((1-accuracy)*len(arr_vals[i]))
result.append(sorted(arr_vals[i])[sample_num])
return result
for badge in signal_strength_1m:
badge.plot(kind='hist', alpha=0.5)
plt.xlabel('Signal Strength')
plt.ylabel('Count')
plt.title('Distribution of signal values for 1 meter distance')
plt.legend(macs, loc='upper right')
[out_90, in_90] = getThresholds(0.9, [signal_strength_1m[0], signal_strength_1m[1]])
plt.axvline(x=out_90, linewidth=2.0, color='r')
plt.axvline(x=in_90, linewidth=2.0, color='r')
# D9 is outside the case and F2 is inside ... menttion on the graph
def getDistance(rssi, n, a):
return 10**((a-rssi)/(10*n))
```
### Compare the obtained constants against experiment conducted with multiple badges
### Combine signal distribution from multiple badge experiment as follows:
### Focus only on the 2 badges facing front in the top two levels (4 badges in total)
### Combine the data from 3 badges outside the case and treat the one outside the case separately
### Plot the distribution for the two groups of badges from the experiment and the function for distance using both these constants
```
multibadge_file_path = '../data/rssi_distance/expt6/expt_07_11_'
raw_data = []
for i in xrange(2,9,2):
raw_data.append(pd.read_csv(multibadge_file_path+str(i)+'f.dat', header=None, sep=' ', skiprows=1, skipfooter=1, parse_dates=True, names=['MAC', 'RSSI', 'TIME']))
macs = [['F2:50:9D:7E:C8:0C,'], ['D9:CB:6E:1F:48:82,', 'CD:A3:F0:C5:68:73,', 'D2:67:85:48:D5:EF,']]
for i,distance in enumerate(raw_data):
vals = []
for mac in macs:
temp = distance[distance['MAC'].isin(mac)]['RSSI']
vals.append(pd.rolling_mean(temp, 5, min_periods=4))
[inside, outside] = getThresholds(0.9, [vals[0], vals[1]])
plt.figure(figsize=(10,7))
for j,val in enumerate(vals):
val.plot(kind='hist', alpha=0.5)
plt.xlabel('Signal Strength')
plt.ylabel('Count')
plt.title('Signal Strength Distribution For ' + str(2*(i+1)) + ' ft')
plt.axvline(x=outside, linewidth=2.0, color='r', label='outside')
plt.axvline(x=inside, linewidth=2.0, color='purple', label='inside')
plt.legend(['outside', 'inside'], loc='upper left')
signal_level = range(-70,-50)
[outside, inside] = getThresholds(0.9, [signal_strength_1m[0], signal_strength_1m[1]])
distances = [[getDistance(level, 2.4, A)*3.33 for level in signal_level] for A in [outside, inside]]
for i in xrange(len(distances)):
plt.plot(signal_level, distances[i], linewidth=2.0)
plt.xlabel('Signal strength (dB)')
plt.ylabel('Distance (feet)')
plt.title("Variation of distance with RSSI value for different 'n'")
plt.legend(['outside', 'inside'], loc='upper right')
labels = ['inside', 'outside']
for i,mac in enumerate(macs):
vals = []
for distance in raw_data:
temp = distance[distance['MAC'].isin(mac)]['RSSI']
vals.append(pd.rolling_mean(temp, 5, min_periods=4))
thresholds = getThresholds(0.9, vals)
plt.figure(figsize=(10,7))
for j,val in enumerate(vals):
val.plot(kind='hist', alpha=0.5)
plt.axvline(x=thresholds[j], linewidth=2.0, color='red')
plt.xlabel('Signal Strength')
plt.ylabel('Count')
plt.title('Signal Strength Distribution For Groups of badges across 2ft-8ft : ' + labels[i])
```
## Analysis for experiment with all badges inside the case and the receiver outside the case
```
cased_badges_file_path = '../data/rssi_distance/expt7/expt_07_29_'
raw_data = []
for i in xrange(2,11,2):
raw_data.append(pd.read_csv(cased_badges_file_path+str(i)+'f.dat', header=None, sep=' ', skiprows=1, skipfooter=1, parse_dates=True, names=['MAC', 'RSSI', 'TIME']))
vals = []
for i,distance in enumerate(raw_data):
temp = distance['RSSI']
vals.append(pd.rolling_mean(temp, 5, min_periods=4))
thresholds = getThresholds(0.9, vals)
plt.figure(figsize=(10,7))
for j,val in enumerate(vals):
val.plot(kind='hist', alpha=0.5)
plt.axvline(x=thresholds[j], linewidth=2.0, color='red', label=str(j))
plt.xlabel('Signal Strength')
plt.ylabel('Count')
plt.title('Signal Strength Distribution For 4 Badges In a Case (Receiver Outside) across 2ft-10ft : ')
plt.legend(['t1', 't2', 't3', 't4', 't5','2ft', '4ft', '6ft', '8ft', '10ft'], loc='upper left')
```
## Analysis For Badges and Receiver Inside the Case
```
receiver_inside_file_path = '../data/rssi_distance/expt8/expt_07_29_'
raw_data = []
for i in xrange(2,11,2):
raw_data.append(pd.read_csv(receiver_inside_file_path+str(i)+'f.dat', header=None, sep=' ', skiprows=1, skipfooter=1, parse_dates=True, names=['MAC', 'RSSI', 'TIME']))
vals = []
for i,distance in enumerate(raw_data):
temp = distance['RSSI']
vals.append(pd.rolling_mean(temp, 5, min_periods=4))
thresholds = getThresholds(0.9, vals)
plt.figure(figsize=(10,7))
for j,val in enumerate(vals):
val.plot(kind='hist', alpha=0.5)
plt.axvline(x=thresholds[j], linewidth=2.0, color='red', label=str(j))
plt.xlabel('Signal Strength')
plt.ylabel('Count')
plt.title('Signal Strength Distribution For 4 Badges In a Case (Receiver Inside) across 2ft-10ft : ')
plt.legend(['t1', 't2', 't3', 't4', 't5','2ft', '4ft', '6ft', '8ft', '10ft'], loc='upper left')
```
| github_jupyter |
```
import requests
from bs4 import BeautifulSoup
import csv
import re
import json
import sqlite3
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from time import sleep
import os
from collections import Counter
import pickle
import warnings
import time
warnings.filterwarnings("ignore")
from wordcloud import WordCloud
import matplotlib.pyplot as plt
import PIL
from PIL import Image, ImageFilter
from selenium import webdriver
from selenium.webdriver.common.action_chains import ActionChains
from selenium.webdriver.chrome.options import Options
import boto3
import botocore
%matplotlib inline
# Use proxy and headers for safe web scraping
# os.environ['HTTPS_PROXY'] = 'http://3.112.188.39:8080'
# pd.options.mode.chained_assignment = None
headers = {
'user-agent': 'Mozilla/5.0 (Windows NT 10.0; Win64; x64) AppleWebKit/'
'537.36 (KHTML, like Gecko) Chrome/75.0.3770.100 Safari/537.36'}
countries_link = {'USA':'https://www.amazon.com',
'Australia':'https://www.amazon.com.au',
'UK':'https://www.amazon.co.uk',
'India':'https://www.amazon.in',
'Japan':'https://www.amazon.co.jp/',
'UAE':'https://amazon.ae'}
```
##### List of Products
```
amazon_usa = {'health_and_beauty':{'hair_products':{'shampoo':'https://www.amazon.com/s?i=beauty-intl-ship&bbn=16225006011&rh=n%3A%2116225006011%2Cn%3A11057241%2Cn%3A17911764011%2Cn%3A11057651&dc&',
'conditioner':'https://www.amazon.com/s?i=beauty-intl-ship&bbn=16225006011&rh=n%3A%2116225006011%2Cn%3A11057241%2Cn%3A17911764011%2Cn%3A11057251&dc&',
'hair_scalp_treatment':'https://www.amazon.com/s?i=beauty-intl-ship&bbn=16225006011&rh=n%3A%2116225006011%2Cn%3A11057241%2Cn%3A11057431&dc&',
'treatment_oil':'https://www.amazon.com/s?i=beauty-intl-ship&bbn=16225006011&rh=n%3A%2116225006011%2Cn%3A11057241%2Cn%3A10666439011&dc&',
'hair_loss':'https://www.amazon.com/s?i=beauty-intl-ship&bbn=16225006011&rh=n%3A%2116225006011%2Cn%3A11057241%2Cn%3A10898755011&dc&'},
'skin_care':{'body':{'cleansers':'https://www.amazon.com/s?i=beauty-intl-ship&bbn=16225006011&rh=n%3A%2116225006011%2Cn%3A11060451%2Cn%3A11060521%2Cn%3A11056281&dc&',
'moisturizers':'https://www.amazon.com/s?i=beauty-intl-ship&bbn=16225006011&rh=n%3A%2116225006011%2Cn%3A11060451%2Cn%3A11060521%2Cn%3A11060661&dc&',
'treatments':'https://www.amazon.com/s?i=beauty-intl-ship&bbn=16225006011&rh=n%3A%2116225006011%2Cn%3A11060451%2Cn%3A11060521%2Cn%3A11056421&dc&'},
'eyes':{'creams':'https://www.amazon.com/s?i=beauty-intl-ship&bbn=16225006011&rh=n%3A%2116225006011%2Cn%3A11060451%2Cn%3A11061941%2Cn%3A7730090011&dc&',
'gels':'https://www.amazon.com/s?i=beauty-intl-ship&bbn=16225006011&rh=n%3A%2116225006011%2Cn%3A11060451%2Cn%3A11061941%2Cn%3A7730092011&dc&',
'serums':'https://www.amazon.com/s?i=beauty-intl-ship&bbn=16225006011&rh=n%3A%2116225006011%2Cn%3A11060451%2Cn%3A11061941%2Cn%3A7730098011&dc&'},
'face':{'f_cleansers':'https://www.amazon.com/s?i=beauty-intl-ship&bbn=16225006011&rh=n%3A%2116225006011%2Cn%3A11060451%2Cn%3A11060711%2Cn%3A11060901&dc&',
'f_moisturizers':'https://www.amazon.com/s?i=beauty-intl-ship&bbn=16225006011&rh=n%3A%2116225006011%2Cn%3A11060451%2Cn%3A11060711%2Cn%3A11060901&dc&',
'scrubs':'https://www.amazon.com/s?i=beauty-intl-ship&bbn=16225006011&rh=n%3A%2116225006011%2Cn%3A11060451%2Cn%3A11060711%2Cn%3A11061091&dc&',
'toners':'https://www.amazon.com/s?i=beauty-intl-ship&bbn=16225006011&rh=n%3A%2116225006011%2Cn%3A11060451%2Cn%3A11060711%2Cn%3A11061931&dc&',
'f_treatments':'https://www.amazon.com/s?i=beauty-intl-ship&bbn=16225006011&rh=n%3A%2116225006011%2Cn%3A11060451%2Cn%3A11060711%2Cn%3A11061931&dc&'},
'lipcare':'https://www.amazon.com/s?i=beauty-intl-ship&bbn=16225006011&rh=n%3A%2116225006011%2Cn%3A11060451%2Cn%3A3761351&dc&'}},
'food':{'tea':{'herbal':'https://www.amazon.com/s?k=tea&i=grocery&rh=n%3A16310101%2Cn%3A16310231%2Cn%3A16521305011%2Cn%3A16318401%2Cn%3A16318511&dc&',
'green':'https://www.amazon.com/s?k=tea&i=grocery&rh=n%3A16310101%2Cn%3A16310231%2Cn%3A16521305011%2Cn%3A16318401%2Cn%3A16318471&dc&',
'black':'https://www.amazon.com/s?k=tea&i=grocery&rh=n%3A16310101%2Cn%3A16310231%2Cn%3A16521305011%2Cn%3A16318401%2Cn%3A16318411&dc&',
'chai':'https://www.amazon.com/s?k=tea&i=grocery&rh=n%3A16310101%2Cn%3A16310231%2Cn%3A16521305011%2Cn%3A16318401%2Cn%3A348022011&dc&'},
'coffee':'https://www.amazon.com/s?k=tea&i=grocery&rh=n%3A16310101%2Cn%3A16310231%2Cn%3A16521305011%2Cn%3A16318031%2Cn%3A2251593011&dc&',
'dried_fruits':{'mixed':'https://www.amazon.com/s?k=dried+fruits&i=grocery&rh=n%3A16310101%2Cn%3A6506977011%2Cn%3A9865332011%2Cn%3A9865334011%2Cn%3A9865348011&dc&',
'mangoes':'https://www.amazon.com/s?k=dried+fruits&rh=n%3A16310101%2Cn%3A9865346011&dc&'},
'nuts':{'mixed':'https://www.amazon.com/s?k=nuts&rh=n%3A16310101%2Cn%3A16322931&dc&',
'peanuts':'https://www.amazon.com/s?k=nuts&i=grocery&rh=n%3A16310101%2Cn%3A18787303011%2Cn%3A16310221%2Cn%3A16322881%2Cn%3A16322941&dc&',
'cashews':'https://www.amazon.com/s?k=nuts&i=grocery&rh=n%3A16310101%2Cn%3A18787303011%2Cn%3A16310221%2Cn%3A16322881%2Cn%3A16322901&dc&'}},
'supplements':{'sports':{'pre_workout':'https://www.amazon.com/s?k=supplements&i=hpc&rh=n%3A3760901%2Cn%3A6973663011%2Cn%3A6973697011&dc&',
'protein':'https://www.amazon.com/s?k=supplements&i=hpc&rh=n%3A3760901%2Cn%3A6973663011%2Cn%3A6973704011&dc&',
'fat_burner':'https://www.amazon.com/s?k=supplements&i=hpc&rh=n%3A3760901%2Cn%3A6973663011%2Cn%3A6973679011&dc&',
'weight_gainer':'https://www.amazon.com/s?k=supplements&i=hpc&rh=n%3A3760901%2Cn%3A6973663011%2Cn%3A6973725011&dc&'},
'vitamins_dietary':{'supplements':'https://www.amazon.com/s?k=supplements&i=hpc&rh=n%3A3760901%2Cn%3A3764441%2Cn%3A6939426011&dc&',
'multivitamins':'https://www.amazon.com/s?k=supplements&i=hpc&rh=n%3A3760901%2Cn%3A3774861&dc&'}},
'wellness':{'ayurveda':'https://www.amazon.com/s?k=supplements&i=hpc&rh=n%3A3760901%2Cn%3A10079996011%2Cn%3A13052911%2Cn%3A13052941&dc&',
'essential_oil_set':'https://www.amazon.com/s?k=supplements&i=hpc&rh=n%3A3760901%2Cn%3A10079996011%2Cn%3A13052911%2Cn%3A18502613011&dc&',
'massage_oil':'https://www.amazon.com/s?k=supplements&i=hpc&rh=n%3A3760901%2Cn%3A10079996011%2Cn%3A14442631&dc&'},
'personal_accessories':{'bags':{'women':{'clutches':'https://www.amazon.com/s?k=bags&i=fashion-womens-handbags&bbn=15743631&rh=n%3A7141123011%2Cn%3A%217141124011%2Cn%3A7147440011%2Cn%3A15743631%2Cn%3A17037745011&dc&',
'crossbody':'https://www.amazon.com/s?k=bags&i=fashion-womens-handbags&bbn=15743631&rh=n%3A7141123011%2Cn%3A%217141124011%2Cn%3A7147440011%2Cn%3A15743631%2Cn%3A2475899011&dc&',
'fashion':'https://www.amazon.com/s?k=bags&i=fashion-womens-handbags&bbn=15743631&rh=n%3A7141123011%2Cn%3A%217141124011%2Cn%3A7147440011%2Cn%3A15743631%2Cn%3A16977745011&dc&',
'hobo':'https://www.amazon.com/s?k=bags&i=fashion-womens-handbags&bbn=15743631&rh=n%3A7141123011%2Cn%3A%217141124011%2Cn%3A7147440011%2Cn%3A15743631%2Cn%3A16977747011&dc&'}},
'jewelry':{'anklets':'https://www.amazon.com/s?i=fashion-womens-intl-ship&bbn=16225018011&rh=n%3A16225018011%2Cn%3A7192394011%2Cn%3A7454897011&dc&',
'bracelets':'https://www.amazon.com/s?i=fashion-womens-intl-ship&bbn=16225018011&rh=n%3A16225018011%2Cn%3A7192394011%2Cn%3A7454898011&dc&',
'earrings':'https://www.amazon.com/s?i=fashion-womens-intl-ship&bbn=16225018011&rh=n%3A16225018011%2Cn%3A7192394011%2Cn%3A7454917011&dc&',
'necklaces':'https://www.amazon.com/s?i=fashion-womens-intl-ship&bbn=16225018011&rh=n%3A16225018011%2Cn%3A7192394011%2Cn%3A7454917011&dc&',
'rings':'https://www.amazon.com/s?i=fashion-womens-intl-ship&bbn=16225018011&rh=n%3A16225018011%2Cn%3A7192394011%2Cn%3A7454939011&dc&'},
'artisan_fabrics':'https://www.amazon.com/s?k=fabrics&rh=n%3A2617941011%2Cn%3A12899121&dc&'}}
amazon_uk = {'health_and_beauty':{'hair_products':{'shampoo':'https://www.amazon.co.uk/b/ref=amb_link_5?ie=UTF8&node=74094031&pf_rd_m=A3P5ROKL5A1OLE&pf_rd_s=merchandised-search-leftnav&pf_rd_r=KF9SM53J2HXHP4EJD3AH&pf_rd_r=KF9SM53J2HXHP4EJD3AH&pf_rd_t=101&pf_rd_p=aaaa7182-fdd6-4b35-8f0b-993e78880b69&pf_rd_p=aaaa7182-fdd6-4b35-8f0b-993e78880b69&pf_rd_i=66469031',
'conditioner':'https://www.amazon.co.uk/b/ref=amb_link_6?ie=UTF8&node=2867976031&pf_rd_m=A3P5ROKL5A1OLE&pf_rd_s=merchandised-search-leftnav&pf_rd_r=KF9SM53J2HXHP4EJD3AH&pf_rd_r=KF9SM53J2HXHP4EJD3AH&pf_rd_t=101&pf_rd_p=aaaa7182-fdd6-4b35-8f0b-993e78880b69&pf_rd_p=aaaa7182-fdd6-4b35-8f0b-993e78880b69&pf_rd_i=66469031',
'hair_loss':'https://www.amazon.co.uk/b/ref=amb_link_11?ie=UTF8&node=2867979031&pf_rd_m=A3P5ROKL5A1OLE&pf_rd_s=merchandised-search-leftnav&pf_rd_r=KF9SM53J2HXHP4EJD3AH&pf_rd_r=KF9SM53J2HXHP4EJD3AH&pf_rd_t=101&pf_rd_p=aaaa7182-fdd6-4b35-8f0b-993e78880b69&pf_rd_p=aaaa7182-fdd6-4b35-8f0b-993e78880b69&pf_rd_i=66469031',
'hair_scalp_treatment':'https://www.amazon.co.uk/b/ref=amb_link_7?ie=UTF8&node=2867977031&pf_rd_m=A3P5ROKL5A1OLE&pf_rd_s=merchandised-search-leftnav&pf_rd_r=KF9SM53J2HXHP4EJD3AH&pf_rd_r=KF9SM53J2HXHP4EJD3AH&pf_rd_t=101&pf_rd_p=aaaa7182-fdd6-4b35-8f0b-993e78880b69&pf_rd_p=aaaa7182-fdd6-4b35-8f0b-993e78880b69&pf_rd_i=66469031',
'treatment_oil':'https://www.amazon.co.uk/hair-oil-argan/b/ref=amb_link_8?ie=UTF8&node=2867981031&pf_rd_m=A3P5ROKL5A1OLE&pf_rd_s=merchandised-search-leftnav&pf_rd_r=KF9SM53J2HXHP4EJD3AH&pf_rd_r=KF9SM53J2HXHP4EJD3AH&pf_rd_t=101&pf_rd_p=aaaa7182-fdd6-4b35-8f0b-993e78880b69&pf_rd_p=aaaa7182-fdd6-4b35-8f0b-993e78880b69&pf_rd_i=66469031'},
'skin_care':{'body':{'cleanser':'https://www.amazon.co.uk/s/ref=lp_344269031_nr_n_3?fst=as%3Aoff&rh=n%3A117332031%2Cn%3A%21117333031%2Cn%3A118464031%2Cn%3A344269031%2Cn%3A344282031&bbn=344269031&ie=UTF8&qid=1581612722&rnid=344269031',
'moisturizers':'https://www.amazon.co.uk/s/ref=lp_344269031_nr_n_1?fst=as%3Aoff&rh=n%3A117332031%2Cn%3A%21117333031%2Cn%3A118464031%2Cn%3A344269031%2Cn%3A2805272031&bbn=344269031&ie=UTF8&qid=1581612722&rnid=344269031'},
'eyes':{'creams':'https://www.amazon.co.uk/s/ref=lp_118465031_nr_n_0?fst=as%3Aoff&rh=n%3A117332031%2Cn%3A%21117333031%2Cn%3A118464031%2Cn%3A118465031%2Cn%3A344259031&bbn=118465031&ie=UTF8&qid=1581612984&rnid=118465031',
'gels':'https://www.amazon.co.uk/s/ref=lp_118465031_nr_n_1?fst=as%3Aoff&rh=n%3A117332031%2Cn%3A%21117333031%2Cn%3A118464031%2Cn%3A118465031%2Cn%3A344258031&bbn=118465031&ie=UTF8&qid=1581613044&rnid=118465031',
'serums':'https://www.amazon.co.uk/s/ref=lp_118465031_nr_n_3?fst=as%3Aoff&rh=n%3A117332031%2Cn%3A%21117333031%2Cn%3A118464031%2Cn%3A118465031%2Cn%3A344257031&bbn=118465031&ie=UTF8&qid=1581613044&rnid=118465031'},
'face':{'cleansers':'https://www.amazon.co.uk/s/ref=lp_118466031_nr_n_1?fst=as%3Aoff&rh=n%3A117332031%2Cn%3A%21117333031%2Cn%3A118464031%2Cn%3A118466031%2Cn%3A344265031&bbn=118466031&ie=UTF8&qid=1581613120&rnid=118466031',
'moisturizers':'https://www.amazon.co.uk/s/ref=lp_118466031_nr_n_3?fst=as%3Aoff&rh=n%3A117332031%2Cn%3A%21117333031%2Cn%3A118464031%2Cn%3A118466031%2Cn%3A2805291031&bbn=118466031&ie=UTF8&qid=1581613120&rnid=118466031',
'toners':'https://www.amazon.co.uk/s/ref=lp_118466031_nr_n_0?fst=as%3Aoff&rh=n%3A117332031%2Cn%3A%21117333031%2Cn%3A118464031%2Cn%3A118466031%2Cn%3A344267031&bbn=118466031&ie=UTF8&qid=1581613120&rnid=118466031',
'treatments':'https://www.amazon.co.uk/s?bbn=118466031&rh=n%3A117332031%2Cn%3A%21117333031%2Cn%3A118464031%2Cn%3A118466031%2Cn%3A18918424031&dc&fst=as%3Aoff&qid=1581613120&rnid=118466031&ref=lp_118466031_nr_n_7'},
'lipcare':'https://www.amazon.co.uk/s/ref=lp_118464031_nr_n_4?fst=as%3Aoff&rh=n%3A117332031%2Cn%3A%21117333031%2Cn%3A118464031%2Cn%3A118467031&bbn=118464031&ie=UTF8&qid=1581613357&rnid=118464031'}},
'food':{'tea':{'herbal':'https://www.amazon.co.uk/s?k=tea&i=grocery&rh=n%3A340834031%2Cn%3A358584031%2Cn%3A11711401%2Cn%3A406567031&dc&qid=1581613483&rnid=344155031&ref=sr_nr_n_1',
'green':'https://www.amazon.co.uk/s?k=tea&i=grocery&rh=n%3A340834031%2Cn%3A358584031%2Cn%3A11711401%2Cn%3A406566031&dc&qid=1581613483&rnid=344155031&ref=sr_nr_n_3',
'black':'https://www.amazon.co.uk/s?k=tea&i=grocery&rh=n%3A340834031%2Cn%3A358584031%2Cn%3A11711401%2Cn%3A406564031&dc&qid=1581613483&rnid=344155031&ref=sr_nr_n_2'},
'coffee':'https://www.amazon.co.uk/s?k=coffee&rh=n%3A340834031%2Cn%3A11711391&dc&qid=1581613715&rnid=1642204031&ref=sr_nr_n_2',
'dried_fruits':{'mixed':'https://www.amazon.co.uk/s?k=dried+fruits&rh=n%3A340834031%2Cn%3A9733163031&dc&qid=1581613770&rnid=1642204031&ref=sr_nr_n_2'},
'nuts':{'mixed':'https://www.amazon.co.uk/s?k=mixed&rh=n%3A359964031&ref=nb_sb_noss',
'peanuts':'https://www.amazon.co.uk/s?k=peanuts&rh=n%3A359964031&ref=nb_sb_noss',
'cashews':'https://www.amazon.co.uk/s?k=cashew&rh=n%3A359964031&ref=nb_sb_noss'}},
'supplements':{'sports':{'pre_workout':'https://www.amazon.co.uk/b/?node=5977685031&ref_=Oct_s9_apbd_odnav_hd_bw_b35Hc3L_1&pf_rd_r=C5MZHH5TH5F868B6FQWD&pf_rd_p=8086b6c9-ae16-5c3c-a879-030afa4ee08f&pf_rd_s=merchandised-search-11&pf_rd_t=BROWSE&pf_rd_i=2826478031',
'protein':'https://www.amazon.co.uk/b/?node=2826510031&ref_=Oct_s9_apbd_odnav_hd_bw_b35Hc3L_0&pf_rd_r=C5MZHH5TH5F868B6FQWD&pf_rd_p=8086b6c9-ae16-5c3c-a879-030afa4ee08f&pf_rd_s=merchandised-search-11&pf_rd_t=BROWSE&pf_rd_i=2826478031',
'fat_burner':'https://www.amazon.co.uk/b/?node=5977737031&ref_=Oct_s9_apbd_odnav_hd_bw_b35Hc3L_2&pf_rd_r=C5MZHH5TH5F868B6FQWD&pf_rd_p=8086b6c9-ae16-5c3c-a879-030afa4ee08f&pf_rd_s=merchandised-search-11&pf_rd_t=BROWSE&pf_rd_i=2826478031'},
'vitamins_dietary':{'supplements':'https://www.amazon.co.uk/b/?_encoding=UTF8&node=2826534031&bbn=65801031&ref_=Oct_s9_apbd_odnav_hd_bw_b35Hdc7_2&pf_rd_r=AY01DQVCB4SE7VVE7MTK&pf_rd_p=1ecdbf02-af23-502a-b7ab-9916ddd6690c&pf_rd_s=merchandised-search-11&pf_rd_t=BROWSE&pf_rd_i=2826484031',
'multivitamins':'https://www.amazon.co.uk/b/?_encoding=UTF8&node=2826506031&bbn=65801031&ref_=Oct_s9_apbd_odnav_hd_bw_b35Hdc7_1&pf_rd_r=AY01DQVCB4SE7VVE7MTK&pf_rd_p=1ecdbf02-af23-502a-b7ab-9916ddd6690c&pf_rd_s=merchandised-search-11&pf_rd_t=BROWSE&pf_rd_i=2826484031'}},
'wellness':{'massage_oil':'https://www.amazon.co.uk/b/?node=3360479031&ref_=Oct_s9_apbd_odnav_hd_bw_b50nmJ_4&pf_rd_r=GYVYF52HT2004EDTY67W&pf_rd_p=3f8e4361-c00b-588b-a07d-ff259bf98bbc&pf_rd_s=merchandised-search-11&pf_rd_t=BROWSE&pf_rd_i=74073031',
'ayurveda':'https://www.amazon.co.uk/s?k=ayurveda&rh=n%3A65801031%2Cn%3A2826449031&dc&qid=1581686978&rnid=1642204031&ref=sr_nr_n_22'},
'personal_accessories':{'bags':{'women':{'clutches':'https://www.amazon.co.uk/b/?node=1769563031&ref_=Oct_s9_apbd_odnav_hd_bw_b1vkt8h_3&pf_rd_r=VC8RX89R4V4JJ5TEBANF&pf_rd_p=cefca17f-8dac-5c80-848f-812aff1bfdd7&pf_rd_s=merchandised-search-11&pf_rd_t=BROWSE&pf_rd_i=1769559031',
'crossbody':'https://www.amazon.co.uk/b/?node=1769564031&ref_=Oct_s9_apbd_odnav_hd_bw_b1vkt8h_1&pf_rd_r=VC8RX89R4V4JJ5TEBANF&pf_rd_p=cefca17f-8dac-5c80-848f-812aff1bfdd7&pf_rd_s=merchandised-search-11&pf_rd_t=BROWSE&pf_rd_i=1769559031',
'fashion':'https://www.amazon.co.uk/b/?node=1769560031&ref_=Oct_s9_apbd_odnav_hd_bw_b1vkt8h_5&pf_rd_r=VC8RX89R4V4JJ5TEBANF&pf_rd_p=cefca17f-8dac-5c80-848f-812aff1bfdd7&pf_rd_s=merchandised-search-11&pf_rd_t=BROWSE&pf_rd_i=1769559031',
'hobo':'https://www.amazon.co.uk/b/?node=1769565031&ref_=Oct_s9_apbd_odnav_hd_bw_b1vkt8h_4&pf_rd_r=VC8RX89R4V4JJ5TEBANF&pf_rd_p=cefca17f-8dac-5c80-848f-812aff1bfdd7&pf_rd_s=merchandised-search-11&pf_rd_t=BROWSE&pf_rd_i=1769559031'}},
'jewelry':{'anklets':'https://www.amazon.co.uk/s/ref=lp_10382835031_nr_n_0?fst=as%3Aoff&rh=n%3A193716031%2Cn%3A%21193717031%2Cn%3A10382835031%2Cn%3A10382860031&bbn=10382835031&ie=UTF8&qid=1581687575&rnid=10382835031',
'bracelets':'https://www.amazon.co.uk/s/ref=lp_10382835031_nr_n_1?fst=as%3Aoff&rh=n%3A193716031%2Cn%3A%21193717031%2Cn%3A10382835031%2Cn%3A10382861031&bbn=10382835031&ie=UTF8&qid=1581687575&rnid=10382835031',
'earrings':'https://www.amazon.co.uk/s/ref=lp_10382835031_nr_n_4?fst=as%3Aoff&rh=n%3A193716031%2Cn%3A%21193717031%2Cn%3A10382835031%2Cn%3A10382865031&bbn=10382835031&ie=UTF8&qid=1581687575&rnid=10382835031',
'necklaces':'https://www.amazon.co.uk/s/ref=lp_10382835031_nr_n_7?fst=as%3Aoff&rh=n%3A193716031%2Cn%3A%21193717031%2Cn%3A10382835031%2Cn%3A10382868031&bbn=10382835031&ie=UTF8&qid=1581687575&rnid=10382835031',
'rings':'https://www.amazon.co.uk/s/ref=lp_10382835031_nr_n_10?fst=as%3Aoff&rh=n%3A193716031%2Cn%3A%21193717031%2Cn%3A10382835031%2Cn%3A10382871031&bbn=10382835031&ie=UTF8&qid=1581687575&rnid=10382835031'},
'artisan_fabrics':'https://www.amazon.co.uk/s?k=fabric&rh=n%3A11052681%2Cn%3A3063518031&dc&qid=1581687726&rnid=1642204031&ref=a9_sc_1'}}
amazon_india = {'health_and_beauty':{'hair_products':{'shampoo':'https://www.amazon.in/b/ref=s9_acss_bw_cg_btyH1_2a1_w?ie=UTF8&node=1374334031&pf_rd_m=A1K21FY43GMZF8&pf_rd_s=merchandised-search-5&pf_rd_r=JHDJ4QHM0APVS05NGF4G&pf_rd_t=101&pf_rd_p=41b9c06b-1514-47de-a1c6-f4f13fb55ffe&pf_rd_i=1374305031',
'conditioner':'https://www.amazon.in/b/ref=s9_acss_bw_cg_btyH1_2b1_w?ie=UTF8&node=1374306031&pf_rd_m=A1K21FY43GMZF8&pf_rd_s=merchandised-search-5&pf_rd_r=CBABMCW6C69JRBGZNWWP&pf_rd_t=101&pf_rd_p=41b9c06b-1514-47de-a1c6-f4f13fb55ffe&pf_rd_i=1374305031',
'treatment_oil':''},
'skin_care':[],
'wellness_product':[]},
'food':{'tea':[],
'coffee':[],
'dried_fruits':[],
'nuts':[],
'supplements':[]},
'personal_accessories':{'bags':[],
'jewelry':[],
'artisan_fabrics':[]}}
amazon_aus = {'health_and_beauty':{'hair_products':{'shampoo':'https://www.amazon.com.au/b/?_encoding=UTF8&node=5150253051&bbn=4851917051&ref_=Oct_s9_apbd_odnav_hd_bw_b5cXATz&pf_rd_r=6SEM7GFDN7CQ2W4KXM9M&pf_rd_p=9dd4b462-1094-5e36-890d-bb1b694c8b53&pf_rd_s=merchandised-search-12&pf_rd_t=BROWSE&pf_rd_i=5150070051',
'conditioner':'https://www.amazon.com.au/b/?_encoding=UTF8&node=5150226051&bbn=4851917051&ref_=Oct_s9_apbd_odnav_hd_bw_b5cXATz&pf_rd_r=6SEM7GFDN7CQ2W4KXM9M&pf_rd_p=9dd4b462-1094-5e36-890d-bb1b694c8b53&pf_rd_s=merchandised-search-12&pf_rd_t=BROWSE&pf_rd_i=5150070051'},
'skin_care':[],
'wellness_product':[]},
'food':{'tea':[],
'coffee':[],
'dried_fruits':[],
'nuts':[],
'supplements':[]},
'personal_accessories':{'bags':[],
'jewelry':[],
'artisan_fabrics':[]}}
amazon = {'USA':amazon_usa,
'UK':amazon_uk,
'India':amazon_india,
'Australia':amazon_aus}
def hover(browser, xpath):
'''
This function makes an automated mouse hovering in the selenium webdriver
element based on its xpath.
PARAMETER
---------
browser: Selenium based webbrowser
xpath: str
xpath of the element in the webpage where hover operation has to be
performed.
'''
element_to_hover_over = browser.find_element_by_xpath(xpath)
hover = ActionChains(browser).move_to_element(element_to_hover_over)
hover.perform()
element_to_hover_over.click()
def browser(link):
'''This funtion opens a selenium based chromebrowser specifically tuned
to work for amazon product(singular item) webpages. Few functionality
includes translation of webpage, clicking the initial popups, and hovering
over product imagesso that the images can be scrape
PARAMETER
---------
link: str
Amazon Product item link
RETURN
------
driver: Selenium web browser with operated functions
'''
options = Options()
prefs = {
"translate_whitelists": {"ja":"en","de":'en'},
"translate":{"enabled":"true"}
}
# helium = r'C:\Users\Dell-pc\AppData\Local\Google\Chrome\User Data\Default\Extensions\njmehopjdpcckochcggncklnlmikcbnb\4.2.12_0'
# options.add_argument(helium)
options.add_experimental_option("prefs", prefs)
options.headless = True
driver = webdriver.Chrome(chrome_options=options)
driver.get(link)
try:
driver.find_element_by_xpath('//*[@id="nav-main"]/div[1]/div[2]/div/div[3]/span[1]/span/input').click()
except:
pass
try:
hover(driver,'//*[@id="altImages"]/ul/li[3]')
except:
pass
try:
driver.find_element_by_xpath('//*[@id="a-popover-6"]/div/header/button/i').click()
except:
pass
try:
hover(driver,'//*[@id="altImages"]/ul/li[4]')
except:
pass
try:
driver.find_element_by_xpath('//*[@id="a-popover-6"]/div/header/button/i').click()
except:
pass
try:
hover(driver,'//*[@id="altImages"]/ul/li[5]')
except:
pass
try:
driver.find_element_by_xpath('//*[@id="a-popover-6"]/div/header/button/i').click()
except:
pass
try:
hover(driver,'//*[@id="altImages"]/ul/li[6]')
except:
pass
try:
driver.find_element_by_xpath('//*[@id="a-popover-6"]/div/header/button/i').click()
except:
pass
try:
hover(driver,'//*[@id="altImages"]/ul/li[7]')
except:
pass
try:
driver.find_element_by_xpath('//*[@id="a-popover-6"]/div/header/button/i').click()
except:
pass
try:
hover(driver,'//*[@id="altImages"]/ul/li[8]')
except:
pass
try:
driver.find_element_by_xpath('//*[@id="a-popover-6"]/div/header/button/i').click()
except:
pass
try:
hover(driver,'//*[@id="altImages"]/ul/li[9]')
except:
pass
try:
driver.find_element_by_xpath('//*[@id="a-popover-6"]/div/header/button/i').click()
except:
pass
return driver
def scroll_temp(driver):
'''
Automated Scroller in Selenium Webbrowser
PARAMETER
---------
driver: Selenium Webbrowser
'''
pre_scroll_height = driver.execute_script('return document.body.scrollHeight;')
run_time, max_run_time = 0, 2
while True:
iteration_start = time.time()
# Scroll webpage, the 100 allows for a more 'aggressive' scroll
driver.execute_script('window.scrollTo(0,0.6*document.body.scrollHeight);')
post_scroll_height = driver.execute_script('return document.body.scrollHeight;')
scrolled = post_scroll_height != pre_scroll_height
timed_out = run_time >= max_run_time
if scrolled:
run_time = 0
pre_scroll_height = post_scroll_height
elif not scrolled and not timed_out:
run_time += time.time() - iteration_start
elif not scrolled and timed_out:
break
def scroll(driver):
scroll_temp(driver)
from selenium.common.exceptions import NoSuchElementException
try:
element = driver.find_element_by_xpath('//*[@id="reviewsMedley"]/div/div[1]')
except NoSuchElementException:
try:
element = driver.find_element_by_xpath('//*[@id="reviewsMedley"]')
except NoSuchElementException:
element = driver.find_element_by_xpath('//*[@id="detail-bullets_feature_div"]')
actions = ActionChains(driver)
actions.move_to_element(element).perform()
def browser_link(product_link,country):
'''Returns all the web link of the products based on the first
page of the product category. It captures product link of all the pages for
that specific product.
PARAMETER
---------
link: str
The initial web link of the product page. This is generally the
first page of the all the items for that specfic product
RETURN
------
links: list
It is a list of strings which contains all the links of the items
for the specific product
'''
driver = browser(product_link)
soup = BeautifulSoup(driver.page_source, 'lxml')
try:
pages_soup = soup.findAll("ul",{"class":"a-pagination"})
pages = int(pages_soup[0].findAll("li",{'class':'a-disabled'})[1].text)
except:
pass
try:
pages_soup = soup.findAll("div",{"id":"pagn"})
pages = int(pages_soup[0].findAll("span",{'class':'pagnDisabled'})[0].text)
except:
try:
pages_soup = soup.findAll("div",{"id":"pagn"})
pages = int(pages_soup[0].findAll("span",{'class':'pagnDisabled'})[1].text)
except:
pass
print(pages)
links = []
for page in range(1,pages+1):
print(page)
link_page = product_link + '&page=' + str(page)
driver_temp = browser(link_page)
time.sleep(2)
soup_temp = BeautifulSoup(driver_temp.page_source, 'lxml')
try:
search = soup_temp.findAll("div",{"id":"mainResults"})
temp_search = search[1].findAll("a",{'class':'a-link-normal s-access-detail-page s-color-twister-title-link a-text-normal'})
for i in range(len(temp_search)):
if country == 'Australia':
link = temp_search[i].get('href')
else:
link = countries_link[country] + temp_search[i].get('href')
links.append(link)
print(len(links))
except:
try:
search = soup_temp.findAll("div",{"class":"s-result-list s-search-results sg-row"})
temp_search = search[1].findAll("h2")
if len(temp_search) < 2:
for i in range(len(search[0].findAll("h2"))):
temp = search[0].findAll("h2")[i]
for j in range(len(temp.findAll('a'))):
link = countries_link[country]+temp.findAll('a')[j].get('href')
links.append(link)
print(len(links))
else:
for i in range(len(search[1].findAll("h2"))):
temp = search[1].findAll("h2")[i]
for j in range(len(temp.findAll('a'))):
link = countries_link[country]+temp.findAll('a')[j].get('href')
links.append(link)
print(len(links))
except:
pass
try:
search = soup_temp.findAll("div",{"id":"mainResults"})
temp_search = search[0].findAll("a",{'class':'a-link-normal s-access-detail-page s-color-twister-title-link a-text-normal'})
for i in range(len(temp_search)):
if country == 'Australia':
link = temp_search[i].get('href')
else:
link = countries_link[country] + temp_search[i].get('href')
links.append(link)
print(len(links))
except:
try:
search = soup_temp.findAll("div",{"class":"s-result-list s-search-results sg-row"})
temp_search = search[1].findAll("h2")
if len(temp_search) < 2:
for i in range(len(search[0].findAll("h2"))):
temp = search[0].findAll("h2")[i]
for j in range(len(temp.findAll('a'))):
link = countries_link[country]+temp.findAll('a')[j].get('href')
links.append(link)
print(len(links))
else:
for i in range(len(search[1].findAll("h2"))):
temp = search[1].findAll("h2")[i]
for j in range(len(temp.findAll('a'))):
link = countries_link[country]+temp.findAll('a')[j].get('href')
links.append(link)
print(len(links))
except:
print('Not Scrapable')
return links
def indexes(amazon_links,link_list):
amazon_dict = amazon_links
if len(link_list) == 5:
return amazon_dict[link_list[0]][link_list[1]][link_list[2]][link_list[3]][link_list[4]]
elif len(link_list) == 4:
return amazon_dict[link_list[0]][link_list[1]][link_list[2]][link_list[3]]
elif len(link_list) == 3:
return amazon_dict[link_list[0]][link_list[1]][link_list[2]]
elif len(link_list) == 2:
return amazon_dict[link_list[0]][link_list[1]]
elif len(link_list) == 1:
return amazon_dict[link_list[0]]
else:
return print("Invalid Product")
def products_links(country, **kwargs):
amazon_links = amazon[country]
directory_temp = []
for key, value in kwargs.items():
directory_temp.append(value)
directory = '/'.join(directory_temp)
print(directory)
product_link = indexes(amazon_links,directory_temp)
main_links = browser_link(product_link,country=country)
return main_links,directory
```
### Product Scraper Function
```
def delete_images(filename):
import os
file_path = '/home/jishu/'
os.remove(file_path + filename)
def upload_s3(filename,key):
key_id = 'AKIAWR6YW7N5ZKW35OJI'
access_key = 'h/xrcI9A2SRU0ds+zts4EClKAqbzU+/iXdiDcgzm'
bucket_name = 'amazon-data-ecfullfill'
s3 = boto3.client('s3',aws_access_key_id=key_id,
aws_secret_access_key=access_key)
try:
s3.upload_file(filename,bucket_name,key)
except FileNotFoundError:
pass
def product_info(link,directory,country):
'''Get all the product information of an Amazon Product'''
#Opening Selenium Webdrive with Amazon product
driver = browser(link)
time.sleep(4)
scroll(driver)
time.sleep(2)
#Initializing BeautifulSoup operation in selenium browser
selenium_soup = BeautifulSoup(driver.page_source, 'lxml')
time.sleep(2)
#Product Title
try:
product_title = driver.find_element_by_xpath('//*[@id="productTitle"]').text
except:
product_title = 'Not Scrapable'
print(product_title)
#Ratings - Star
try:
rating_star = float(selenium_soup.findAll('span',{'class':'a-icon-alt'})[0].text.split()[0])
except:
rating_star = 'Not Scrapable'
print(rating_star)
#Rating - Overall
try:
overall_rating = int(selenium_soup.findAll('span',{'id':'acrCustomerReviewText'})[0].text.split()[0].replace(',',''))
except:
overall_rating = 'Not Scrapable'
print(overall_rating)
#Company
try:
company = selenium_soup.findAll('a',{'id':'bylineInfo'})[0].text
except:
company = 'Not Scrapable'
print(country)
#Price
try:
if country=='UAE':
denomination = selenium_soup.findAll('span',{'id':'priceblock_ourprice'})[0].text[:3]
price = float(selenium_soup.findAll('span',{'id':'priceblock_ourprice'})[0].text[3:])
else:
denomination = selenium_soup.findAll('span',{'id':'priceblock_ourprice'})[0].text[0]
price = float(selenium_soup.findAll('span',{'id':'priceblock_ourprice'})[0].text[1:])
except:
try:
if country=='UAE':
try:
price = float(selenium_soup.findAll('span',{'id':'priceblock_ourprice'})[0].text[3:].replace(',',''))
except:
price = float(selenium_soup.findAll('span',{'id':'priceblock_dealprice'})[0].text[3:].replace(',',''))
else:
try:
price = float(selenium_soup.findAll('span',{'id':'priceblock_ourprice'})[0].text[3:].replace(',',''))
except:
price = float(selenium_soup.findAll('span',{'id':'priceblock_dealprice'})[0].text[3:].replace(',',''))
except:
denomination = 'Not Scrapable'
price = 'Not Scrapable'
print(denomination,price)
#Product Highlights
try:
temp_ph = selenium_soup.findAll('ul',{'class':'a-unordered-list a-vertical a-spacing-none'})[0].findAll('li')
counter_ph = len(temp_ph)
product_highlights = []
for i in range(counter_ph):
raw = temp_ph[i].text
clean = raw.strip()
product_highlights.append(clean)
product_highlights = '<CPT14>'.join(product_highlights)
except:
try:
temp_ph = selenium_soup.findAll('div',{'id':'rich-product-description'})[0].findAll('p')
counter_ph = len(temp_ph)
product_highlights = []
for i in range(counter_ph):
raw = temp_ph[i].text
clean = raw.strip()
product_highlights.append(clean)
product_highlights = '<CPT14>'.join(product_highlights)
except:
product_highlights = 'Not Available'
print(product_highlights)
#Product Details/Dimensions:
#USA
try:
temp_pd = selenium_soup.findAll('div',{'class':'content'})[0].findAll('ul')[0].findAll('li')
counter_pd = len(temp_pd)
for i in range(counter_pd):
try:
if re.findall('ASIN',temp_pd[i].text)[0]:
try:
asin = temp_pd[i].text.split(' ')[1]
except:
pass
except IndexError:
pass
try:
if re.findall('Product Dimensions|Product Dimension|Product dimensions',temp_pd[i].text)[0]:
pd_temp = temp_pd[i].text.strip().split('\n')[2].strip().split(';')
try:
product_length = float(pd_temp[0].split('x')[0])
except IndexError:
pass
try:
product_width = float(pd_temp[0].split('x')[1])
except IndexError:
pass
try:
product_height = float(pd_temp[0].split('x')[2].split(' ')[1])
except IndexError:
pass
try:
pd_unit = pd_temp[0].split('x')[2].split(' ')[2]
except IndexError:
pass
try:
product_weight = float(pd_temp[1].split(' ')[1])
except IndexError:
pass
try:
weight_unit = pd_temp[1].split(' ')[2]
except IndexError:
pass
except:
pass
try:
if re.findall('Shipping Weight|Shipping weight|shipping weight',temp_pd[i].text)[0]:
sweight_temp = temp_pd[i].text.split(':')[1].strip().split(' ')
shipping_weight = float(sweight_temp[0])
shipping_weight_unit = sweight_temp[1]
except IndexError:
pass
try:
if re.findall('Amazon Best Sellers Rank|Amazon Bestsellers Rank',temp_pd[i].text)[0]:
x = temp_pd[i].text.replace('\n','').split(' ')
indexes = []
for j,k in enumerate(x):
if re.findall('#',k):
indexes.append(j)
try:
best_seller_cat = int(temp_pd[i].text.strip().replace('\n','').split(' ')[3].replace(',',''))
best_seller_prod = int(x[indexes[0]].split('#')[1].split('in')[0])
except:
try:
best_seller_cat = x[indexes[0]].split('#')[1]
except:
pass
try:
best_seller_prod = x[indexes[1]].split('#')[1].split('in')[0]
except:
pass
except IndexError:
pass
print(asin)
except:
pass
try:
temp_pd = selenium_soup.findAll('div',{'class':'content'})[1].findAll('ul')[0].findAll('li')
counter_pd = len(temp_pd)
for i in range(counter_pd):
try:
if re.findall('ASIN',temp_pd[i].text)[0]:
try:
asin = temp_pd[i].text.split(' ')[1]
except:
pass
except IndexError:
pass
try:
if re.findall('Product Dimensions|Product Dimension|Product dimensions',temp_pd[i].text)[0]:
pd_temp = temp_pd[i].text.strip().split('\n')[2].strip().split(';')
try:
product_length = float(pd_temp[0].split('x')[0])
except IndexError:
pass
try:
product_width = float(pd_temp[0].split('x')[1])
except IndexError:
pass
try:
product_height = float(pd_temp[0].split('x')[2].split(' ')[1])
except IndexError:
pass
try:
pd_unit = pd_temp[0].split('x')[2].split(' ')[2]
except IndexError:
pass
try:
product_weight = float(pd_temp[1].split(' ')[1])
except IndexError:
pass
try:
weight_unit = pd_temp[1].split(' ')[2]
except IndexError:
pass
except:
pass
try:
if re.findall('Shipping Weight|Shipping weight|shipping weight',temp_pd[i].text)[0]:
sweight_temp = temp_pd[i].text.split(':')[1].strip().split(' ')
shipping_weight = float(sweight_temp[0])
shipping_weight_unit = sweight_temp[1]
except IndexError:
pass
try:
if re.findall('Amazon Best Sellers Rank|Amazon Bestsellers Rank',temp_pd[i].text)[0]:
x = temp_pd[i].text.replace('\n','').split(' ')
indexes = []
for j,k in enumerate(x):
if re.findall('#',k):
indexes.append(j)
try:
best_seller_cat = int(temp_pd[i].text.strip().replace('\n','').split(' ')[3].replace(',',''))
best_seller_prod = int(x[indexes[0]].split('#')[1].split('in')[0])
except:
try:
best_seller_cat = x[indexes[0]].split('#')[1]
except:
pass
try:
best_seller_prod = x[indexes[1]].split('#')[1].split('in')[0]
except:
pass
except IndexError:
pass
print(asin)
except:
pass
#India
try:
temp_pd = selenium_soup.findAll('div',{'class':'content'})[0].findAll('ul')[0].findAll('li')
counter_pd = len(temp_pd)
for i in range(counter_pd):
try:
if re.findall('ASIN',temp_pd[i].text)[0]:
asin = temp_pd[i].text.split(' ')[1]
except:
pass
try:
if re.findall('Product Dimensions|Product Dimension|Product dimensions',temp_pd[i].text)[0]:
pd_temp = temp_pd[i].text.strip().split('\n')[2].strip().split(' ')
try:
product_length = float(pd_temp[0])
except:
pass
try:
product_width = float(pd_temp[2])
except:
pass
try:
product_height = float(pd_temp[4])
except:
pass
try:
pd_unit = pd_temp[5]
except:
pass
try:
product_weight = float(pd_temp[1].split(' ')[1])
except:
pass
try:
weight_unit = pd_temp[1].split(' ')[2]
except:
pass
print(asin)
except IndexError:
pass
try:
if re.findall('Shipping Weight|Shipping weight|shipping weight',temp_pd[i].text)[0]:
sweight_temp = temp_pd[i].text.split(':')[1].strip().split(' ')
shipping_weight = float(sweight_temp[0])
shipping_weight_unit = sweight_temp[1]
except IndexError:
pass
try:
if re.findall('Item Weight|Product Weight|Item weight|Product weight|Boxed-product Weight',temp_pd[i].text)[0]:
pd_weight_temp = temp_pd[i].text.replace('\n','').strip().split(' ')[1].strip()
product_weight = float(pd_weight_temp.split(' ')[0])
weight_unit = pd_weight_temp.split(' ')[1]
except IndexError:
pass
try:
if re.findall('Amazon Best Sellers Rank|Amazon Bestsellers Rank',temp_pd[i].text)[0]:
x = temp_pd[i].text.strip().replace('\n','').split(' ')
indexes = []
for j,k in enumerate(x):
if re.findall('#',k):
indexes.append(j)
try:
best_seller_cat = int(temp_pd[i].text.strip().replace('\n','').split(' ')[3].replace(',',''))
best_seller_prod = int(x[indexes[0]].split('#')[1].split('in')[0])
except:
try:
best_seller_cat = x[indexes[0]].split('#')[1]
except:
pass
try:
best_seller_prod = x[indexes[1]].split('#')[1].split('in')[0]
except:
pass
except IndexError:
pass
print(asin)
except:
pass
try:
try:
asin = list(selenium_soup.findAll('div',{'class':'pdTab'})[1].findAll('tr')[0].findAll('td')[1])[0]
except:
pass
try:
dimensions = list(selenium_soup.findAll('div',{'class':'pdTab'})[0].findAll('tr')[0].findAll('td')[1])[0]
except:
pass
try:
weight_temp = list(selenium_soup.findAll('div',{'class':'pdTab'})[1].findAll('tr')[1].findAll('td')[1])[0]
except:
pass
try:
best_seller_cat = float(list(selenium_soup.findAll('div',{'class':'pdTab'})[1].findAll('tr')[5].findAll('td')[1])[0].split('\n')[-1].split(' ')[0].replace(',',''))
except:
pass
try:
best_seller_prod = int(list(list(list(list(selenium_soup.findAll('div',{'class':'pdTab'})[1].findAll('tr')[5].findAll('td')[1])[5])[1])[1])[0].replace('#',''))
except:
pass
try:
product_length = float(dimensions.split('x')[0])
except:
pass
try:
product_width = float(dimensions.split('x')[1])
except:
pass
try:
product_height = float(dimensions.split('x')[2].split(' ')[1])
except:
pass
try:
product_weight = weight_temp.split(' ')[0]
except:
pass
try:
weight_unit = weight_temp.split(' ')[1]
except:
pass
try:
pd_unit = dimensions.split(' ')[-1]
except:
pass
print(asin)
except:
try:
for j in [0,1]:
temp_pd = selenium_soup.findAll('table',{'class':'a-keyvalue prodDetTable'})[j].findAll('tr')
for i in range(len(temp_pd)):
if re.findall('ASIN',temp_pd[i].text):
asin = temp_pd[i].text.strip().split('\n')[3].strip()
if re.findall('Item Model Number|Item model number',temp_pd[i].text):
bait = temp_pd[i].text.strip().split('\n')[3].strip()
if re.findall('Best Sellers Rank|Amazon Best Sellers Rank|Amazon Bestsellers Rank',temp_pd[i].text):
x = temp_pd[i].text.strip().replace('\n','').split(' ')
indexes = []
for j,k in enumerate(x):
if re.findall('#',k):
indexes.append(j)
best_seller_cat = int(x[indexes[0]].split('#')[1])
best_seller_prod = int(x[indexes[1]].split('#')[1].split('in')[0])
if re.findall('Product Dimensions|Product dimension|Product Dimension',temp_pd[i].text):
dimensions = temp_pd[i].text.strip().split('\n')[3].strip().split('x')
product_length = float(dimensions[0].strip())
product_width = float(dimensions[1].strip())
product_height = float(dimensions[2].strip().split(' ')[0])
pd_unit = dimensions[2].strip().split(' ')[1]
if re.findall('Item Weight|Product Weight|Item weight|Boxed-product Weight',temp_pd[i].text):
weight_temp = temp_pd[i].text.strip().split('\n')[3].strip()
product_weight = float(weight_temp.split(' ')[0])
weight_unit = weight_temp.split(' ')[1]
if re.findall('Shipping Weight|Shipping weight|shipping weight',temp_pd[i].text):
sweight_temp = temp_pd[i].text.replace('\n','').strip().split(' ')[1].lstrip().split(' ')
shipping_weight = float(sweight_temp[0])
shipping_weight_unit = sweight_temp[1]
print(asin,bait)
except:
try:
temp_pd = selenium_soup.findAll('div',{'id':'prodDetails'})[0].findAll('tr')
for i in range(len(temp_pd)):
if re.findall('ASIN',temp_pd[i].text):
asin = temp_pd[i].text.strip().split('\n')[3].strip()
if re.findall('Best Sellers Rank|Amazon Best Sellers Rank|Amazon Bestsellers Rank',temp_pd[i].text):
x = temp_pd[i].text.strip().replace('\n','').split(' ')
indexes = []
for j,k in enumerate(x):
if re.findall('#',k):
indexes.append(j)
best_seller_cat = int(x[indexes[0]].split('#')[1])
best_seller_prod = int(x[indexes[1]].split('#')[1].split('in')[0])
if re.findall('Product Dimensions|Product dimension|Product Dimension',temp_pd[i].text):
dimensions = temp_pd[i].text.strip().split('\n')[3].strip().split('x')
product_length = float(dimensions[0].strip())
product_width = float(dimensions[1].strip())
product_height = float(dimensions[2].strip().split(' ')[0])
pd_unit = dimensions[2].strip().split(' ')[1]
if re.findall('Item Weight|Product Weight|Item weight|Boxed-product Weight',temp_pd[i].text):
weight_temp = temp_pd[i].text.strip().split('\n')[3].strip()
product_weight = float(weight_temp.split(' ')[0])
weight_unit = weight_temp.split(' ')[1]
if re.findall('Shipping Weight|Shipping weight|shipping weight',temp_pd[i].text):
sweight_temp = temp_pd[i].text.replace('\n','').strip().split(' ')[1].lstrip().split(' ')
shipping_weight = float(sweight_temp[0])
shipping_weight_unit = sweight_temp[1]
except:
try:
temp_pd = selenium_soup.findAll('div',{'id':'detail_bullets_id'})[0].findAll('tr')[0].findAll('li')
for i in range(len(temp_pd)):
if re.findall('ASIN',temp_pd[i].text):
asin = temp_pd[i].text.strip().split(':')[1].strip()
if re.findall('Best Sellers Rank|Amazon Best Sellers Rank|Amazon Bestsellers Rank',temp_pd[i].text):
x = temp_pd[i].text.strip().replace('\n','').split(' ')
indexes = []
for j,k in enumerate(x):
if re.findall('#',k):
indexes.append(j)
best_seller_cat = int(x[indexes[0]].split('#')[1])
best_seller_prod = int(x[indexes[1]].split('#')[1].split('in')[0])
if re.findall('Product Dimensions|Product dimension|Product Dimension',temp_pd[i].text):
dimensions = temp_pd[i].text.strip().split('\n')[2].strip().split('x')
product_length = float(dimensions[0].strip())
product_width = float(dimensions[1].strip())
product_height = float(dimensions[2].strip().split(' ')[0])
pd_unit = dimensions[2].strip().split(' ')[1]
if re.findall('Item Weight|Product Weight|Item weight|Boxed-product Weight',temp_pd[i].text):
weight_temp = temp_pd[i].text.strip().split('\n')[2].strip()
product_weight = float(weight_temp.split(' ')[0])
weight_unit = weight_temp.split(' ')[1]
if re.findall('Shipping Weight|Shipping weight|shipping weight',temp_pd[i].text):
sweight_temp = temp_pd[i].text.replace('\n','').strip().split(' ')[1].lstrip().split(' ')
shipping_weight = float(sweight_temp[0])
shipping_weight_unit = sweight_temp[1]
except:
pass
try:
print(asin)
except NameError:
asin = 'Not Scrapable'
try:
print(best_seller_cat)
except NameError:
best_seller_cat = 'Not Scrapable'
try:
print(best_seller_prod)
except NameError:
best_seller_prod = 'Not Scrapable'
try:
print(product_length)
except NameError:
product_length = 'Not Scrapable'
try:
print(product_width)
except NameError:
product_width = 'Not Scrapable'
try:
print(product_height)
except NameError:
product_height = 'Not Scrapable'
try:
print(product_weight)
except NameError:
product_weight = 'Not Scrapable'
try:
print(weight_unit)
except NameError:
weight_unit = 'Not Scrapable'
try:
print(pd_unit)
except NameError:
pd_unit = 'Not Scrapable'
try:
print(shipping_weight_unit)
except NameError:
shipping_weight_unit = 'Not Scrapable'
try:
print(shipping_weight)
except NameError:
shipping_weight = 'Not Scrapable'
print(product_length,product_width,product_height,product_weight,asin,pd_unit,
best_seller_cat,best_seller_prod,weight_unit,shipping_weight,shipping_weight_unit)
#Customer Review Ratings - Overall
time.sleep(0.5)
try:
temp_crr = selenium_soup.findAll('table',{'id':'histogramTable'})[1].findAll('a')
crr_main = {}
crr_temp = []
counter_crr = len(temp_crr)
for i in range(counter_crr):
crr_temp.append(temp_crr[i]['title'])
crr_temp = list(set(crr_temp))
for j in range(len(crr_temp)):
crr_temp[j] = crr_temp[j].split(' ')
stopwords = ['stars','represent','of','rating','reviews','have']
for word in list(crr_temp[j]):
if word in stopwords:
crr_temp[j].remove(word)
print(crr_temp[j])
try:
if re.findall(r'%',crr_temp[j][1])[0]:
crr_main.update({int(crr_temp[j][0]): int(crr_temp[j][1].replace('%',''))})
except:
crr_main.update({int(crr_temp[j][1]): int(crr_temp[j][0].replace('%',''))})
except:
try:
temp_crr = selenium_soup.findAll('table',{'id':'histogramTable'})[1].findAll('span',{'class':'a-offscreen'})
crr_main = {}
counter_crr = len(temp_crr)
star = counter_crr
for i in range(counter_crr):
crr_main.update({star:int(temp_crr[i].text.strip().split('/n')[0].split(' ')[0].replace('%',''))})
star -= 1
except:
pass
try:
crr_5 = crr_main[5]
except:
crr_5 = 0
try:
crr_4 = crr_main[4]
except:
crr_4 = 0
try:
crr_3 = crr_main[3]
except:
crr_3 = 0
try:
crr_2 = crr_main[2]
except:
crr_2 = 0
try:
crr_1 = crr_main[1]
except:
crr_1 = 0
#Customer Review Ratings - By Feature
time.sleep(1)
try:
driver.find_element_by_xpath('//*[@id="cr-summarization-attributes-list"]/div[4]/a/span').click()
temp_fr = driver.find_element_by_xpath('//*[@id="cr-summarization-attributes-list"]').text
temp_fr = temp_fr.split('\n')
crr_feature_title = []
crr_feature_rating = []
for i in [0,2,4]:
crr_feature_title.append(temp_fr[i])
for j in [1,3,5]:
crr_feature_rating.append(temp_fr[j])
crr_feature = dict(zip(crr_feature_title,crr_feature_rating))
except:
try:
temp_fr = driver.find_element_by_xpath('//*[@id="cr-summarization-attributes-list"]').text
temp_fr = temp_fr.split('\n')
crr_feature_title = []
crr_feature_rating = []
for i in [0,2,4]:
crr_feature_title.append(temp_fr[i])
for j in [1,3,5]:
crr_feature_rating.append(temp_fr[j])
crr_feature = dict(zip(crr_feature_title,crr_feature_rating))
except:
crr_feature = 'Not Defined'
try:
crr_feature_key = list(crr_feature.keys())
except:
pass
try:
crr_fr_1 = crr_feature[crr_feature_key[0]]
except:
crr_fr_1 = 0
try:
crr_fr_2 = crr_feature[crr_feature_key[1]]
except:
crr_fr_2 = 0
try:
crr_fr_3 = crr_feature[crr_feature_key[2]]
except:
crr_fr_3 = 0
#Tags:
time.sleep(1)
try:
temp_tags = selenium_soup.findAll('div',{'class':'cr-lighthouse-terms'})[0]
counter_tags = len(temp_tags)
print('Counter Tags:',counter_tags)
tags = []
for i in range(counter_tags):
tags.append(temp_tags.findAll('span')[i].text.strip())
print(tags[i])
except:
tags = ['None']
try:
for feature in crr_feature_key:
tags.append(feature)
except:
pass
tags = list(set(tags))
tags = '<CPT14>'.join(tags)
print(tags)
#Images
images = []
for i in [0,3,4,5,6,7,8,9]:
try:
images.append(selenium_soup.findAll('div',{'class':'imgTagWrapper'})[i].find('img')['src'])
except:
pass
import urllib.request
for i in range(len(images)):
if asin =='Not Scrapable':
product_image = "{}_{}.jpg".format(product_title,i)
product_image = product_image.replace('/','')
urllib.request.urlretrieve(images[i],product_image)
upload_s3("{}_{}.jpg".format(product_title,i),
directory+"/images/" + product_image)
delete_images(product_image)
else:
product_image = "{}_{}.jpg".format(asin,i)
product_image = product_image.replace('/','')
urllib.request.urlretrieve(images[i],product_image)
upload_s3("{}_{}.jpg".format(asin,i),
directory+"/images/" + product_image)
delete_images(product_image)
return [product_title,rating_star,overall_rating,company,price,
product_highlights,product_length,product_width,product_height,
product_weight,asin,pd_unit,best_seller_cat,best_seller_prod,
weight_unit,shipping_weight,shipping_weight_unit,crr_5,crr_4,
crr_3,crr_2,crr_1,crr_fr_1,crr_fr_2,crr_fr_3,tags,directory]
```
### Data Wrangling
```
def database(product_data,**kwargs):
try:
try:
link = kwargs['link']
except KeyError:
print('Error in Link')
try:
country = kwargs['country']
except KeyError:
print("Enter Country Name")
try:
cat1 = kwargs['cat1']
except KeyError:
pass
try:
cat2 = kwargs['cat2']
except KeyError:
pass
try:
cat3 = kwargs['cat3']
except KeyError:
pass
try:
cat4 = kwargs['cat4']
except KeyError:
pass
try:
product = kwargs['product']
except KeyError:
print("Enter Product Name")
metadata = [link,country,cat1,cat2,cat3,cat4,product]
except NameError:
try:
cat4 = None
metadata = [link,country,cat1,cat2,cat3,cat4,product]
except NameError:
try:
cat4 = None
cat3 = None
metadata = [link,country,cat1,cat2,cat3,cat4,product]
except NameError:
cat4 = None
cat3 = None
cat2 = None
metadata = [link,country,cat1,cat2,cat3,cat4,product]
conn = sqlite3.connect('{}.db'.format(product))
headers = ['link','country','cat1','cat2','cat3','cat4','product','product_title',
'rating_star','overall_rating','company','price',
'product_highlights','product_length','product_width','product_height',
'product_weight','asin','pd_unit','best_seller_cat','best_seller_prod',
'weight_unit','shipping_weight','shipping_weight_unit','crr_5','crr_4',
'crr_3','crr_2','crr_1','crr_fr_1','crr_fr_2','crr_fr_3','tags','images_link']
product_data.append(metadata)
product_data = product_data[-1] + product_data[:len(product_data)-1]
temp = pd.DataFrame(data= [product_data],columns=headers)
temp.to_sql('Product',conn,if_exists='append')
upload_s3(product+'.db',directory+'/'+product+'.db')
conn.close()
def checkpoint(link_list,directory,product):
BUCKET_NAME = 'amazon-data-ecfullfill'
key_id = 'AKIAWR6YW7N5ZKW35OJI'
access_key = 'h/xrcI9A2SRU0ds+zts4EClKAqbzU+/iXdiDcgzm'
KEY = '{}/{}.db'.format(directory,product)
s3 = boto3.resource('s3',aws_access_key_id=key_id,
aws_secret_access_key=access_key)
try:
s3.Bucket(BUCKET_NAME).download_file(KEY, 'test.db')
except botocore.exceptions.ClientError as e:
if e.response['Error']['Code'] == "404":
print("The object does not exist.")
else:
raise
conn = sqlite3.connect('test.db')
try:
df = pd.read_sql('''SELECT * FROM Product''', conn)
product_link = df['link'].unique()
new_list = []
for i in link_list:
if i in product_link:
pass
else:
new_list.append(i)
except:
new_list = link_list
return new_list
```
### Execution
```
#Initializing the product per Jupyter Notebook
country = 'USA'
cat1 = 'food'
cat2='tea'
# cat3='None'
# cat4 = 'None'
product='herbal'
links,directory = products_links(country=country,category=cat1,cat2=cat2,product=product)
# test_1 = {'links':links,'directory':directory}
# import pickle
# with open('food_tea_herbal.pkl', 'wb') as f:
# pickle.dump(test_1, f)
with open('food_tea_herbal.pkl', 'rb') as f:
file = pickle.load(f)
links = file['links']
directory = 'Amazon_USA/food/tea/herbal'
#replace links with new_links if interruption
for link in new_links:
data = product_info(link=link,directory=directory,country='USA')
conn = sqlite3.connect('{}.db'.format(product))
database(product_data=data,link=link,country=country,
cat1=cat1,cat2=cat2,product=product)
# Run if there is an interruption
new_links = checkpoint(links,directory,product)
len(new_links)
len(links)
```
#### Testing the datasets in S3
```
BUCKET_NAME = 'amazon-data-ecfullfill' # replace with your bucket name
key_id = 'AKIAWR6YW7N5ZKW35OJI'
access_key = 'h/xrcI9A2SRU0ds+zts4EClKAqbzU+/iXdiDcgzm'
KEY = 'Amazon_USA/health_and_beauty/hair_products/shampoo/shampoo.db' # replace with your object key
s3 = boto3.resource('s3',aws_access_key_id=key_id,
aws_secret_access_key=access_key)
try:
s3.Bucket(BUCKET_NAME).download_file(KEY, 'test.db')
except botocore.exceptions.ClientError as e:
if e.response['Error']['Code'] == "404":
print("The object does not exist.")
else:
raise
conn = sqlite3.connect('shampoo.db')
df_USA = pd.read_sql("SELECT * FROM Product",conn)
df_USA.iloc[:,:15]
df_USA.iloc[:,15:]
len(link_db)
# def upload_s3(filename,key):
# key_id = 'AKIAWR6YW7N5ZKW35OJI'
# access_key = 'h/xrcI9A2SRU0ds+zts4EClKAqbzU+/iXdiDcgzm'
# bucket_name = 'amazon-data-ecfullfill'
# s3 = boto3.client('s3',aws_access_key_id=key_id,
# aws_secret_access_key=access_key)
# # s3.put_object(Bucket=bucket_name, Key='Amazon/health_and_beauty/hair_product/shampoo')
# s3.upload_file(filename,bucket_name,key)
```
| github_jupyter |
# ML Pipeline Preparation
Follow the instructions below to help you create your ML pipeline.
### 1. Import libraries and load data from database.
- Import Python libraries
- Load dataset from database with [`read_sql_table`](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.read_sql_table.html)
- Define feature and target variables X and Y
```
# import libraries
import nltk
nltk.download(['punkt', 'wordnet', 'averaged_perceptron_tagger','omw-1.4'])
from sqlalchemy import create_engine
import pandas as pd
from nltk.tokenize import word_tokenize
from nltk.stem import WordNetLemmatizer
%matplotlib inline
import re
import numpy as np
import pandas as pd
from nltk.tokenize import word_tokenize
from nltk.stem import WordNetLemmatizer
# !pip install -U scikit-learn
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import confusion_matrix
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
from sklearn.pipeline import Pipeline, FeatureUnion
from sklearn.base import BaseEstimator, TransformerMixin
from sklearn.feature_extraction.text import CountVectorizer, TfidfTransformer
from sklearn.multioutput import MultiOutputClassifier
from sklearn.linear_model import SGDClassifier
from sklearn.base import BaseEstimator
from sklearn.svm import SVC
# from sentence_transformers import SentenceTransformer
# load data from database
engine = create_engine('sqlite:///disaster_reponse.db')
df = pd.read_sql('combined_df',con=engine)
X = df['message']
Y = df.iloc[:,4:]
df.genre.value_counts().sort_values().plot(kind = 'bar')
df
dic={}
for i,column in enumerate(Y.columns):
print(Y[column].value_counts())
dic['column']=column
dic['']
yy=Y.apply(pd.Series.value_counts)
yy= yy.reset_index().rename(columns={'index': 'category'})
yy =yy.melt(id_vars=["category"],
var_name="response type",
value_name="Value")
yy
dic={}
graphs=[]
for cat in yy.category:
yy_temp=yy[yy.category==cat]
dic_2={}
dic_2['x']=yy_temp['response type'].tolist()
dic_2['y']=yy_temp['Value'].tolist()
dic_2['name']=cat
dic[cat]=dic_2
graphs.append(dic_2)
yy=Y.apply(pd.Series.value_counts)
yy.index.astype(str, copy = False)
```
### 2. Write a tokenization function to process your text data
```
def tokenize(text):
url_regex = 'http[s]?://(?:[a-zA-Z]|[0-9]|[$-_@.&+]|[!*\(\),]|(?:%[0-9a-fA-F][0-9a-fA-F]))+'
detected_urls = re.findall(url_regex, text)
for url in detected_urls:
text = text.replace(url, "urlplaceholder")
tokens = word_tokenize(text)
lemmatizer = WordNetLemmatizer()
clean_tokens = []
for tok in tokens:
clean_tok = lemmatizer.lemmatize(tok).lower().strip()
clean_tokens.append(clean_tok)
return clean_tokens
class StartingVerbExtractor(BaseEstimator, TransformerMixin):
def starting_verb(self, text):
sentence_list = nltk.sent_tokenize(text)
for sentence in sentence_list:
pos_tags = nltk.pos_tag(tokenize(sentence))
first_word, first_tag = pos_tags[0]
if first_tag in ['VB', 'VBP'] or first_word == 'RT':
return 1
return 0
def fit(self, x, y=None):
return self
def transform(self, X):
X_tagged = pd.Series(X).apply(self.starting_verb)
return pd.DataFrame(X_tagged)
class ClfSwitcher(BaseEstimator):
def __init__(
self,
estimator = SGDClassifier(),
):
"""
A Custom BaseEstimator that can switch between classifiers.
:param estimator: sklearn object - The classifier
"""
self.estimator = estimator
def fit(self, X, y=None, **kwargs):
self.estimator.fit(X, y)
return self
def predict(self, X, y=None):
return self.estimator.predict(X)
def predict_proba(self, X):
return self.estimator.predict_proba(X)
def score(self, X, y):
return self.estimator.score(X, y)
```
### 3. Build a machine learning pipeline
This machine pipeline should take in the `message` column as input and output classification results on the other 36 categories in the dataset. You may find the [MultiOutputClassifier](http://scikit-learn.org/stable/modules/generated/sklearn.multioutput.MultiOutputClassifier.html) helpful for predicting multiple target variables.
```
pipeline = Pipeline([
('features', FeatureUnion([
('text_pipeline', Pipeline([
('vect', CountVectorizer(tokenizer=tokenize)),
('tfidf', TfidfTransformer())
])),
('starting_verb', StartingVerbExtractor())
])),
('clf', MultiOutputClassifier(RandomForestClassifier()))
])
```
### 4. Train pipeline
- Split data into train and test sets
- Train pipeline
```
X_train, X_test, y_train, y_test = train_test_split(X, Y)
model = pipeline.fit(X_train,y_train)
y_pred = model.predict(X_test)
```
### 5. Test your model
Report the f1 score, precision and recall for each output category of the dataset. You can do this by iterating through the columns and calling sklearn's `classification_report` on each.
```
from sklearn.metrics import classification_report
y_pred_df = pd.DataFrame(y_pred, columns=Y.columns)
for i,column in enumerate(Y.columns):
print(column,classification_report(y_test[column], y_pred_df[column]))
```
### 6. Improve your model
Use grid search to find better parameters.
```
def build_model():
pipeline = Pipeline([
('features', FeatureUnion([
('text_pipeline', Pipeline([
('vect', CountVectorizer(tokenizer=tokenize)),
('tfidf', TfidfTransformer())
])),
('starting_verb', StartingVerbExtractor())
])),
('clf', MultiOutputClassifier(ClfSwitcher()))
])
# parameters = {
# 'features__text_pipeline__vect__ngram_range': ((1, 1), (1, 2)),
# 'features__text_pipeline__vect__max_df': (0.5, 0.75, 1.0),
# 'features__text_pipeline__vect__max_features': (None, 5000, 10000),
# 'features__text_pipeline__tfidf__use_idf': (True, False),
# 'clf__estimator__n_estimators': [50, 100, 200],
# 'clf__estimator__min_samples_split': [2, 3, 4],
# 'features__transformer_weights': (
# {'text_pipeline': 1, 'starting_verb': 0.5},
# {'text_pipeline': 0.5, 'starting_verb': 1},
# {'text_pipeline': 0.8, 'starting_verb': 1},
# )
# }
parameters = [
{
'clf__estimator': [SVC()], # SVM if hinge loss / logreg if log loss
'features__text_pipeline__vect__max_df': (0.5, 0.75, 1.0),
'clf__estimator__gamma': ('scale', 'auto'),
'clf__estimator__C': [0.1, 1,100],
},
{
'clf__estimator': [RandomForestClassifier()],
'features__text_pipeline__vect__max_df': (0.5, 0.75, 1.0),
'clf__estimator__n_estimators': [50, 100, 200],
'clf__estimator__min_samples_split': [2, 3, 4],
},]
cv = GridSearchCV(pipeline, param_grid=parameters, verbose=3)
return cv
```
### 7. Test your model
Show the accuracy, precision, and recall of the tuned model.
Since this project focuses on code quality, process, and pipelines, there is no minimum performance metric needed to pass. However, make sure to fine tune your models for accuracy, precision and recall to make your project stand out - especially for your portfolio!
```
def load_data():
engine = create_engine('sqlite:///disaster_reponse.db')
df = pd.read_sql('combined_df',con=engine)
X = df['message']
Y = df.iloc[:,4:]
return X, Y
def main():
X, y = load_data()
X_train, X_test, y_train, y_test = train_test_split(X, y)
model = build_model()
model.fit(X_train, y_train)
y_pred = model.predict(X_test)
return model
model = main()
```
### 8. Try improving your model further. Here are a few ideas:
* try other machine learning algorithms
* add other features besides the TF-IDF
### 9. Export your model as a pickle file
```
import pickle
with open('model_pkl', 'wb') as files:
pickle.dump(model, files)
```
### 10. Use this notebook to complete `train.py`
Use the template file attached in the Resources folder to write a script that runs the steps above to create a database and export a model based on a new dataset specified by the user.
| github_jupyter |
<a href="https://colab.research.google.com/github/akrambchiri123/deep-learning-coursera/blob/master/Python_%7C_NLP_analysis_of_Restaurant_reviews.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
# Importing Libraries
import numpy as np
import pandas as pd
# Import dataset
dataset = pd.read_csv('/content/Restaurant_Reviews.tsv', delimiter = '\t')
dataset.head(10)
```
Cleaning processing
```
# library to clean data
import re
# Natural Language Tool Kit
import nltk
nltk.download('stopwords')
# to remove stopword
from nltk.corpus import stopwords
# for Stemming propose
from nltk.stem.porter import PorterStemmer
# Initialize empty array
# to append clean text
corpus = []
# 1000 (reviews) rows to clean
for i in range(0, 1000):
# column : "Review", row ith
review = re.sub('[^a-zA-Z]', ' ', dataset['Review'][i])
# convert all cases to lower cases
review = review.lower()
# split to array(default delimiter is " ")
review = review.split()
# creating PorterStemmer object to
# take main stem of each word
ps = PorterStemmer()
# loop for stemming each word
# in string array at ith row
review = [ps.stem(word) for word in review
if not word in set(stopwords.words('english'))]
# rejoin all string array elements
# to create back into a string
review = ' '.join(review)
# append each string to create
# array of clean text
corpus.append(review)
len(corpus)
print(corpus)
```
jusqua maintenant nadhfna each row by removing white space ,lower,removing stops words
**Making the bag of words via sparse matrix**
```
# Creating the Bag of Words model
from sklearn.feature_extraction.text import CountVectorizer
# To extract max 1500 feature.
# "max_features" is attribute to
# experiment with to get better results
cv = CountVectorizer(max_features = 1500)
# X contains corpus (dependent variable)
X = cv.fit_transform(corpus).toarray()
# y contains answers if review
# is positive or negative
y = dataset.iloc[:, 1].values
X
```
Splitting Corpus into Training and Test set. For this, we need class train_test_split from sklearn.cross_validation. Split can be made 70/30 or 80/20 or 85/15 or 75/25, here I choose 75/25 via “test_size”.
X is the bag of words, y is 0 or 1 (positive or negative). **texte en gras**
```
# Splitting the dataset into
# the Training set and Test set
from sklearn.model_selection import train_test_split
# experiment with "test_size"
# to get better results
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.25)
```
**Fitting a Predictive Model (here random forest)**
```
# Fitting Random Forest Classification
# to the Training set
from sklearn.ensemble import RandomForestClassifier
# n_estimators can be said as number of
# trees, experiment with n_estimators
# to get better results
model = RandomForestClassifier(n_estimators = 501,
criterion = 'entropy')
model.fit(X_train, y_train)
```
** Pridicting Final Results via using .predict() method with attribute X_test**
```
# Predicting the Test set results
y_pred = model.predict(X_test)
y_pred
# Making the Confusion Matrix
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(y_test, y_pred)
cm
X_test
myreview='I m not bad'
myreview = myreview.lower()
# split to array(default delimiter is " ")
myreview = myreview.split()
ps = PorterStemmer()
myreview = [ps.stem(word) for word in myreview
if not word in set(stopwords.words('english'))]
myreview = ' '.join(myreview)
l=[]
l.append(myreview)
myrv = cv.transform(l).toarray()
y=model.predict(myrv)
if (y==1):
print('Positive review')
else:
print('negative review')
```
| github_jupyter |
# Displacement controlled normal contact
***
In this notebook we will make a contact model which solves a normal contact problem with a specified displacement.
For normal contact problems with specified loads see the 'recreating the hertz solition numerically' example.
Here again we will use the hertz solution as an easy way to verify that we are getting sensible results.
First lets import everything we will need (no one actually writes these first it's just convention to put them at the top of the file)
```
%matplotlib inline
import slippy.surface as s # surface generation and manipulation
import slippy.contact as c # contact modelling
import numpy as np # numerical functions
import matplotlib.pyplot as plt # plotting
```
## Making the surfaces
In order to solve the problem the geometry must be set. As we are solving the hertz problem we will use an analytically defined round surface and an analytically defined flat surface. For your own analyses the geometry data can come from experimental data or be generated randomly. For more information on this see examples about the surface class and the analytical surfaces.
Importantly at least one of the surfaces must be discrete to be used in a contact model.
```
flat_surface = s.FlatSurface(shift=(0, 0))
round_surface = s.RoundSurface((1, 1, 1), extent=(0.006, 0.006),
shape=(255, 255), generate=True)
```
## Setting the materials
The material for each surface must also be set. This material controls the deformation behaviour of the surface, the fricition and wear behaviour must be set separately. Please see tutorials on adding sub models to the step for more information.
```
# set materials
steel = c.Elastic('Steel', {'E': 200e9, 'v': 0.3})
aluminum = c.Elastic('Aluminum', {'E': 70e9, 'v': 0.33})
flat_surface.material = aluminum
round_surface.material = steel
```
## Making a contact model
Now we have surfaces and materials, but we need a contact model to hold this information and control the solving of model steps, this will become more important when we have multiple steps but for now think of the contact model as a container that all of the information is put in.
```
# create model
my_model = c.ContactModel('model-1', round_surface, flat_surface)
```
## Making a model step and adding it to the model
A step defines what happens to the surfaces during a set period. Here we will add a step that sets the interferance between the surfaces from the point of first touching. The resulting loads on the surfaces and deflections at each point on the surface will be found.
This can then be combined with ther sub models to specify friction or wear behaviour etc. however in this example we will simply compare the results back to the hertz solution. In order to do this we will use the analytical hertz solver to generate a sensible interferance.
```
# Find the analytical result
analy_result = c.hertz_full([1, 1], [np.inf, np.inf], [200e9, 70e9], [0.3, 0.33], 100)
# Make the model step
my_step = c.StaticStep(step_name='This is the step name',
interference=analy_result['total_deflection'])
# Add the step to the model
my_model.add_step(my_step)
```
## Model steps
The steps of the model are stored in the steps property of the model, this is an ordered dictionary, with the keys being the same as the step names. To retrieve a step you can index thie dictionary with the step name. However, if you try to add two steps with the same name the first will be overwritten.
```
my_model.steps
```
## Solving the model
The entire model can then be solved using the solve method of the contact model. This will run through all the steps in order and return the model state at the end of the last step. Other information can be saved using output requests, but as we only have one step there is no need for this.
Before running, by default the model will data check it's self, this action checks that each step and sub model can be solved with the information from the current state. It dosen't check for numerical stabiltiy or accuracy. This can be skipped if necessary.
```
final_state = my_model.solve()
```
## Checking the model against the analytical result
Now lets check the model results against the analytical result from the hertz solution. Althoug this particular step directly sets the interferance, most steps in slippy are solved iteratively, so it is a good idea to check that the set parameter converged to the desired value:
```
print('Solution converged at: ', final_state['interference'], ' interferance')
print('Set interferance was:', analy_result['total_deflection'])
```
Lets check the maximum pressure, contact area and total load all match up with the analytical values:
```
print('Analytical total load: ', 100)
print('Numerical total load: ',
round_surface.grid_spacing**2*sum(final_state['loads_z'].flatten()))
print('Analytical max pressure: ', analy_result['max_pressure'])
print('Numerical max pressure: ', max(final_state['loads_z'].flatten()))
print('Analytical contact area: ', analy_result['contact_area'])
print('Numerical contact area: ',
round_surface.grid_spacing**2* sum(final_state['contact_nodes'].flatten()))
```
## Checking the form of the result
We can also check that the individual surface loads line up with the analytical solution:
```
fig, axes = plt.subplots(1, 3, figsize=(20, 4))
X,Y = round_surface.get_points_from_extent()
X,Y = X-X[-1,-1]/2 , Y-Y[-1,-1]/2
Z_n = final_state['loads_z']
axes[0].imshow(Z_n)
axes[0].set_title('Numerical Result')
R = np.sqrt(X**2+Y**2)
Z_a = analy_result['pressure_f'](R)
axes[1].imshow(Z_a)
axes[1].set_title('Analytical Result')
axes[2].imshow(np.abs(Z_a-Z_n))
for im in axes[2].get_images():
im.set_clim(0, analy_result['max_pressure'])
_ = axes[2].set_title('Absolute Error')
```
## Other items in the state dict
Other items can be listed from the state dict by the following code:
```
print(list(final_state.keys()))
```
* 'just_touching_gap' The gap between the surface at the point when they first touch
* 'surface_1_points' The points of the first surface which are in the solution domain
* 'surface_2_points' The points of the second surface which are in the solution domain
* 'time' The current modelled time which is 0 at the start of the model. This is used for submodels which introdce time dependent behaviour
* 'time_step' The current time step, again used for submodels
* 'new_step' Ture if this is the first substep in a model step
* 'off_set' The tangential displacement between the surfaces at the end of the step
* 'loads_z' The loads acting on each point of the surface
* 'total_displacement_z' The total displacement of each point of the surface pair
* 'surface_1_displacement_z' The displacement of each point on surface 1
* 'surface_2_displacement_z' The displacement of each point on surface 2
* 'contact_nodes' A boolean array showing which nodes are in contact, to find the percentage in contact simply take the mean of this array
* 'total_normal_load' The total load pressing the surfaces together
* 'interference' The distance the surfaces have pushed into eachother from the point of first touching
* 'converged' True if the step converged to a soltuion
* 'gap' The gap between the surfaces when loaded, including deformation
```
# getting the deformed gap between the surfaces
def_gap = (final_state['gap'])
plt.imshow(def_gap)
```
# Saving outputs
You can also save outputs from the model by using an output request, but this is not needed for single step models
| github_jupyter |
# Cell Editing
DataGrid cells can be edited using in-place editors built into DataGrid. Editing can be initiated by double clicking on a cell or by starting typing the new value for the cell.
DataGrids are not editable by default. Editing can be enabled by setting `editable` property to `True`. Selection enablement is required for editing to work and it is set automatically to `cell` mode if it is `none` when editing is enabled.
### Cursor Movement
Editing is initiated for the `cursor` cell. Cursor cell is the same as the selected cell if there is a single cell selected. If there are multiple cells / rectangles selected then cursor cell is the cell where the last selection rectangle was started.
Cursor can be moved in four directions by using the following keyboard keys.
- **Down**: Enter
- **Up**: Shift + Enter
- **Right**: Tab
- **Left**: Shift + Tab
Once done with editing a cell, cursor can be moved to next cell based on the keyboard hit following the rules above.
```
from ipydatagrid import DataGrid
from json import load
import pandas as pd
with open("./cars.json") as fobj:
data = load(fobj)
df = pd.DataFrame(data["data"]).drop("index", axis=1)
datagrid = DataGrid(df, editable=True, layout={"height": "200px"})
datagrid
```
All grid views are updated simultaneously to reflect cell edit changes.
```
datagrid
# keep track of changed cells
changed_cells = {}
def create_cell_key(cell):
return "{row}:{column}".format(row=cell["row"], column=cell["column_index"])
def track_changed_cell(cell):
key = create_cell_key(cell)
changed_cells[key] = cell
```
Changes to cell values can be tracked by subscribing to `on_cell_change` event as below.
```
def on_cell_changed(cell):
track_changed_cell(cell)
print(
"Cell at primary key {row} and column '{column}'({column_index}) changed to {value}".format(
row=cell["row"],
column=cell["column"],
column_index=cell["column_index"],
value=cell["value"],
)
)
datagrid.on_cell_change(on_cell_changed)
```
A cell's value can also be changed programmatically by using the DataGrid methods `set_cell_value` and `set_cell_value_by_index`
```
datagrid.set_cell_value("Cylinders", 2, 12)
```
Whether new cell values are entered using UI or programmatically, both the DataGrid cell rendering and the underlying python data are updated.
```
datagrid.data.iloc[2]["Cylinders"]
datagrid.set_cell_value_by_index("Horsepower", 3, 169)
datagrid.data.iloc[3]["Origin"]
def select_all_changed_cells():
datagrid.clear_selection()
for cell in changed_cells.values():
datagrid.select(cell["row"], cell["column_index"])
return datagrid.selected_cells
```
Show all cells changed using UI or programmatically by selecting them.
```
select_all_changed_cells()
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import os
import tensorflow as tf
from keras import backend as K
# os.environ["CUDA_VISIBLE_DEVICES"] = '1'
config = tf.ConfigProto()
config.gpu_options.allow_growth = True
sess = tf.Session(config=config)
K.set_session(sess)
from keras.regularizers import l2
from keras.layers import Input, Dense, Flatten, GlobalAveragePooling2D, Activation, Conv2D, MaxPooling2D, BatchNormalization, Lambda, Dropout
from keras.layers import SeparableConv2D, Add, Convolution2D, concatenate, Layer, ReLU, DepthwiseConv2D, Reshape, Multiply, InputSpec
from keras.models import Model, load_model, model_from_json
from keras.preprocessing.image import ImageDataGenerator
from keras.optimizers import Adam, SGD
from keras.callbacks import EarlyStopping, ReduceLROnPlateau
from keras.utils import to_categorical
from sklearn import metrics
from sklearn.metrics import roc_curve, roc_auc_score, confusion_matrix, classification_report
import matplotlib.pyplot as plt
from scipy.optimize import brentq
from scipy.interpolate import interp1d
import glob
from PIL import Image
# from tqdm import trange
import random
from keras.applications import Xception, ResNet152
from PIL import ImageFile
ImageFile.LOAD_TRUNCATED_IMAGES = True
import cv2
nb_classes = 2 # number of classes
img_width, img_height = 64, 64 # change based on the shape/structure of your images
batch_size = 64 # try 4, 8, 16, 32, 64, 128, 256 dependent on CPU/GPU memory capacity (powers of 2 values).
nb_epoch = 300 # number of iteration the algorithm gets trained.
def bgr(img):
return cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
train_dir = '/mnt/a/fakedata/face2face/train'
validation_dir = '/mnt/a/fakedata/face2face/val'
test50_dir = '/mnt/a/fakedata/face2face/test'
def res_block(x, in_planes, out_planes, bottleneck_ratio=4, strides=1):
bottleneck_planes = in_planes // bottleneck_ratio
out = BatchNormalization()(x)
out = Activation('relu')(out)
if strides == 2 or in_planes != out_planes:
x_res = Conv2D(out_planes, kernel_size=1, strides=strides, use_bias=False, kernel_initializer='he_normal')(x)
else:
x_res = x
out = Conv2D(bottleneck_planes, kernel_size=1, strides=1, use_bias=False, kernel_initializer='he_normal')(out)
out = BatchNormalization()(out)
out = Activation('relu')(out)
out = Conv2D(bottleneck_planes, kernel_size=3, padding='same', strides=strides, use_bias=False, kernel_initializer='he_normal')(out)
out = BatchNormalization()(out)
out = Activation('relu')(out)
out = Conv2D(out_planes, kernel_size=1, strides=1, use_bias=False, kernel_initializer='he_normal')(out)
out = Add()([out, x_res])
return out
img_input = Input(shape=[img_width, img_height, 3])
x = Conv2D(16, kernel_size=7, strides=2, padding='same', use_bias=False, kernel_initializer='he_normal')(img_input)
x = BatchNormalization()(x)
x = Activation('relu')(x)
x = res_block(x, 16, 64, strides=1)
for i in range(49):
x = res_block(x, 64, 64, strides=1)
x = res_block(x, 64, 256, strides=2)
for i in range(49):
x = res_block(x, 256, 256)
x = res_block(x, 256, 512, strides=2)
for i in range(49):
x = res_block(x, 512, 512)
x_gap = GlobalAveragePooling2D()(x)
x_dense = Dense(nb_classes)(x_gap)
x_sm = Activation('softmax')(x_dense)
model = Model(img_input, x_sm)
model.summary()
model.compile(optimizer=Adam(),
loss='categorical_crossentropy',
metrics=['accuracy'])
print(len(model.trainable_weights))
train_datagen = ImageDataGenerator(rescale=1./255, preprocessing_function=bgr)
test_datagen = ImageDataGenerator(rescale=1./255, preprocessing_function=bgr)
train_generator = train_datagen.flow_from_directory(train_dir,
target_size=(img_height, img_width),
batch_size=batch_size,
shuffle=True,
class_mode='categorical')
validation_generator = train_datagen.flow_from_directory(validation_dir,
target_size=(img_height, img_width),
batch_size=batch_size,
shuffle=False,
class_mode='categorical')
test50_generator = test_datagen.flow_from_directory(test50_dir,
target_size=(img_height, img_width),
batch_size=batch_size,
shuffle=False,
class_mode='categorical')
callback_list = [EarlyStopping(monitor='val_accuracy', patience=10),
ReduceLROnPlateau(monitor='val_loss', factor=0.1, patience=3)]
history = model.fit_generator(train_generator,
steps_per_epoch=200,
epochs=100,
validation_data=validation_generator,
validation_steps=len(validation_generator),
callbacks=callback_list)
model.save('/home/www/fake_detection/model/face2face_resnet.h5')
model = load_model('/home/www/fake_detection/model/face2face_resnet.h5', compile=False)
output = model.predict_generator(test50_generator, steps=len(test50_generator), verbose=1)
np.set_printoptions(formatter={'float': lambda x: "{0:0.3f}".format(x)})
print(test50_generator.class_indices)
print(output)
output_score50 = []
output_class50 = []
answer_class50 = []
answer_class50_1 =[]
for i in range(len(test50_generator)):
output50 = model.predict_on_batch(test50_generator[i][0])
output_score50.append(output50)
answer_class50.append(test50_generator[i][1])
output_score50 = np.concatenate(output_score50)
answer_class50 = np.concatenate(answer_class50)
output_class50 = np.argmax(output_score50, axis=1)
answer_class50_1 = np.argmax(answer_class50, axis=1)
print(output_class50)
print(answer_class50_1)
cm50 = confusion_matrix(answer_class50_1, output_class50)
report50 = classification_report(answer_class50_1, output_class50)
recall50 = cm50[0][0] / (cm50[0][0] + cm50[0][1])
fallout50 = cm50[1][0] / (cm50[1][0] + cm50[1][1])
fpr50, tpr50, thresholds50 = roc_curve(answer_class50_1, output_score50[:, 1], pos_label=1.)
eer50 = brentq(lambda x : 1. - x - interp1d(fpr50, tpr50)(x), 0., 1.)
thresh50 = interp1d(fpr50, thresholds50)(eer50)
print(report50)
print(cm50)
print("AUROC: %f" %(roc_auc_score(answer_class50_1, output_score50[:, 1])))
print(thresh50)
print('test_acc: ', len(output_class50[np.equal(output_class50, answer_class50_1)]) / len(output_class50))
def cutout(img):
"""
# Function: RandomCrop (ZeroPadded (4, 4)) + random occulusion image
# Arguments:
img: image
# Returns:
img
"""
img = bgr(img)
height = img.shape[0]
width = img.shape[1]
channels = img.shape[2]
MAX_CUTS = 3 # chance to get more cuts
MAX_LENGTH_MUTIPLIER = 5 # chance to get larger cuts
# 16 for cifar10, 8 for cifar100
# Zero-padded (4, 4)
# img = np.pad(img, ((4,4),(4,4),(0,0)), mode='constant', constant_values=(0))
# # random-crop 64x64
# dy, dx = height, width
# x = np.random.randint(0, width - dx + 1)
# y = np.random.randint(0, height - dy + 1)
# img = img[y:(y+dy), x:(x+dx)]
# mean norm
# mean = img.mean(keepdims=True)
# img -= mean
img *= 1./255
mask = np.ones((height, width, channels), dtype=np.float32)
nb_cuts = np.random.randint(0, MAX_CUTS + 1)
# cutout
for i in range(nb_cuts):
y = np.random.randint(height)
x = np.random.randint(width)
length = 4 * np.random.randint(1, MAX_LENGTH_MUTIPLIER+1)
y1 = np.clip(y-length//2, 0, height)
y2 = np.clip(y+length//2, 0, height)
x1 = np.clip(x-length//2, 0, width)
x2 = np.clip(x+length//2, 0, width)
mask[y1:y2, x1:x2, :] = 0.
img = img * mask
return img
class ReLU6(Layer):
def __init__(self):
super().__init__(name="ReLU6")
self.relu6 = ReLU(max_value=6, name="ReLU6")
def call(self, input):
return self.relu6(input)
class HardSigmoid(Layer):
def __init__(self):
super().__init__()
self.relu6 = ReLU6()
def call(self, input):
return self.relu6(input + 3.0) / 6.0
class HardSwish(Layer):
def __init__(self):
super().__init__()
self.hard_sigmoid = HardSigmoid()
def call(self, input):
return input * self.hard_sigmoid(input)
class Attention(Layer):
def __init__(self, ch, **kwargs):
super(Attention, self).__init__(**kwargs)
self.channels = ch
self.filters_f_g = self.channels // 8
self.filters_h = self.channels
def build(self, input_shape):
kernel_shape_f_g = (1, 1) + (self.channels, self.filters_f_g)
print(kernel_shape_f_g)
kernel_shape_h = (1, 1) + (self.channels, self.filters_h)
# Create a trainable weight variable for this layer:
self.gamma = self.add_weight(name='gamma', shape=[1], initializer='zeros', trainable=True)
self.kernel_f = self.add_weight(shape=kernel_shape_f_g,
initializer='glorot_uniform',
name='kernel_f')
self.kernel_g = self.add_weight(shape=kernel_shape_f_g,
initializer='glorot_uniform',
name='kernel_g')
self.kernel_h = self.add_weight(shape=kernel_shape_h,
initializer='glorot_uniform',
name='kernel_h')
self.bias_f = self.add_weight(shape=(self.filters_f_g,),
initializer='zeros',
name='bias_F')
self.bias_g = self.add_weight(shape=(self.filters_f_g,),
initializer='zeros',
name='bias_g')
self.bias_h = self.add_weight(shape=(self.filters_h,),
initializer='zeros',
name='bias_h')
super(Attention, self).build(input_shape)
# Set input spec.
self.input_spec = InputSpec(ndim=4,
axes={3: input_shape[-1]})
self.built = True
def call(self, x):
def hw_flatten(x):
return K.reshape(x, shape=[K.shape(x)[0], K.shape(x)[1]*K.shape(x)[2], K.shape(x)[-1]])
f = K.conv2d(x,
kernel=self.kernel_f,
strides=(1, 1), padding='same') # [bs, h, w, c']
f = K.bias_add(f, self.bias_f)
g = K.conv2d(x,
kernel=self.kernel_g,
strides=(1, 1), padding='same') # [bs, h, w, c']
g = K.bias_add(g, self.bias_g)
h = K.conv2d(x,
kernel=self.kernel_h,
strides=(1, 1), padding='same') # [bs, h, w, c]
h = K.bias_add(h, self.bias_h)
s = tf.matmul(hw_flatten(g), hw_flatten(f), transpose_b=True) # # [bs, N, N]
beta = K.softmax(s, axis=-1) # attention map
o = K.batch_dot(beta, hw_flatten(h)) # [bs, N, C]
o = K.reshape(o, shape=K.shape(x)) # [bs, h, w, C]
x = self.gamma * o + x
return x
def compute_output_shape(self, input_shape):
return input_shape
ft_dir = '/mnt/a/fakedata/face2face/finetune'
train_gen_aug = ImageDataGenerator(shear_range=0,
zoom_range=0,
rotation_range=0.2,
width_shift_range=2.,
height_shift_range=2.,
horizontal_flip=True,
zca_whitening=False,
fill_mode='nearest',
preprocessing_function=cutout)
test_datagen = ImageDataGenerator(rescale=1./255, preprocessing_function=bgr)
ft_gen = train_gen_aug.flow_from_directory(ft_dir,
target_size=(img_height, img_width),
batch_size=batch_size,
shuffle=True,
class_mode='categorical')
validation_generator = test_datagen.flow_from_directory(validation_dir,
target_size=(img_height, img_width),
batch_size=batch_size,
shuffle=False,
class_mode='categorical')
test50_generator = test_datagen.flow_from_directory(test50_dir,
target_size=(img_height, img_width),
batch_size=batch_size,
shuffle=False,
class_mode='categorical')
model_ft = load_model('/home/www/fake_detection/model/face2face_resnet.h5', compile=False)
for i in range(3):
model_ft.layers.pop()
im_in = Input(shape=(img_width, img_height, 3))
base_model = Model(img_input, x)
base_model.set_weights(model_ft.get_weights())
# for i in range(len(base_model.layers) - 0):
# base_model.layers[i].trainable = False
x1 = base_model(im_in) # (12, 12, 32)
########### Mobilenet block bneck 3x3 (32 --> 128) #################
expand1 = Conv2D(576, kernel_size=1, strides=1, kernel_regularizer=l2(1e-5), use_bias=False)(x1)
expand1 = BatchNormalization()(expand1)
expand1 = HardSwish()(expand1)
dw1 = DepthwiseConv2D(kernel_size=(3,3), strides=(2,2), padding='same', depthwise_regularizer=l2(1e-5), use_bias=False)(expand1)
dw1 = BatchNormalization()(dw1)
se_gap1 = GlobalAveragePooling2D()(dw1)
se_gap1 = Reshape([1, 1, -1])(se_gap1)
se1 = Conv2D(144, kernel_size=1, strides=1, padding='valid', kernel_regularizer=l2(1e-5), use_bias=False)(se_gap1)
se1 = Activation('relu')(se1)
se1 = Conv2D(576, kernel_size=1, strides=1, padding='valid', kernel_regularizer=l2(1e-5), use_bias=False)(se1)
se1 = HardSigmoid()(se1)
se1 = Multiply()([expand1, se1])
project1 = HardSwish()(se1)
project1 = Conv2D(128, kernel_size=(1, 1), padding='valid', kernel_regularizer=l2(1e-5), use_bias=False)(project1)
project1 = BatchNormalization()(project1)
########### Mobilenet block bneck 5x5 (128 --> 128) #################
expand2 = Conv2D(576, kernel_size=1, strides=1, kernel_regularizer=l2(1e-5), use_bias=False)(project1)
expand2 = BatchNormalization()(expand2)
expand2 = HardSwish()(expand2)
dw2 = DepthwiseConv2D(kernel_size=(5,5), strides=(1,1), padding='same', depthwise_regularizer=l2(1e-5), use_bias=False)(expand2)
dw2 = BatchNormalization()(dw2)
se_gap2 = GlobalAveragePooling2D()(dw2)
se_gap2 = Reshape([1, 1, -1])(se_gap2)
se2 = Conv2D(144, kernel_size=1, strides=1, padding='valid', kernel_regularizer=l2(1e-5), use_bias=False)(se_gap2)
se2 = Activation('relu')(se2)
se2 = Conv2D(576, kernel_size=1, strides=1, padding='valid', kernel_regularizer=l2(1e-5), use_bias=False)(se2)
se2 = HardSigmoid()(se2)
se2 = Multiply()([expand2, se2])
project2 = HardSwish()(se2)
project2 = Conv2D(128, kernel_size=(1, 1), padding='valid', kernel_regularizer=l2(1e-5), use_bias=False)(project2)
project2 = BatchNormalization()(project2)
project2 = Add()([project1, project2])
########### Mobilenet block bneck 5x5 (128 --> 128) #################
expand3 = Conv2D(576, kernel_size=1, strides=1, kernel_regularizer=l2(1e-5), use_bias=False)(project2)
expand3 = BatchNormalization()(expand3)
expand3 = HardSwish()(expand3)
dw3 = DepthwiseConv2D(kernel_size=(5,5), strides=(1,1), padding='same', depthwise_regularizer=l2(1e-5), use_bias=False)(expand3)
dw3 = BatchNormalization()(dw3)
se_gap3 = GlobalAveragePooling2D()(dw3)
se_gap3 = Reshape([1, 1, -1])(se_gap3)
se3 = Conv2D(144, kernel_size=1, strides=1, padding='valid', kernel_regularizer=l2(1e-5), use_bias=False)(se_gap3)
se3 = Activation('relu')(se3)
se3 = Conv2D(576, kernel_size=1, strides=1, padding='valid', kernel_regularizer=l2(1e-5), use_bias=False)(se3)
se3 = HardSigmoid()(se3)
se3 = Multiply()([expand3, se3])
project3 = HardSwish()(se3)
project3 = Conv2D(128, kernel_size=(1, 1), padding='valid', kernel_regularizer=l2(1e-5), use_bias=False)(project3)
project3 = BatchNormalization()(project3)
project3 = Add()([project2, project3])
expand4 = Conv2D(576, kernel_size=1, strides=1, kernel_regularizer=l2(1e-5), use_bias=False)(project3)
expand4 = BatchNormalization()(expand4)
expand4 = HardSwish()(expand4)
dw4 = DepthwiseConv2D(kernel_size=(5,5), strides=(1,1), padding='same', depthwise_regularizer=l2(1e-5), use_bias=False)(expand4)
dw4 = BatchNormalization()(dw4)
se_gap4 = GlobalAveragePooling2D()(dw4)
se_gap4 = Reshape([1, 1, -1])(se_gap4)
se4 = Conv2D(144, kernel_size=1, strides=1, padding='valid', kernel_regularizer=l2(1e-5), use_bias=False)(se_gap4)
se4 = Activation('relu')(se4)
se4 = Conv2D(576, kernel_size=1, strides=1, padding='valid', kernel_regularizer=l2(1e-5), use_bias=False)(se4)
se4 = HardSigmoid()(se4)
se4 = Multiply()([expand4, se4])
project4 = HardSwish()(se4)
project4 = Conv2D(128, kernel_size=(1, 1), padding='valid', kernel_regularizer=l2(1e-5), use_bias=False)(project4)
project4 = BatchNormalization()(project4)
project4 = Add()([project3, project4])
########## Classification ##########
x2 = Conv2D(576, kernel_size=1, strides=1, padding='valid', kernel_regularizer=l2(1e-5), use_bias=False)(project4)
x2 = BatchNormalization()(x2)
x2 = HardSwish()(x2)
x2 = GlobalAveragePooling2D()(x2)
######### Image Attention Model #########
### Block 1 ###
x3 = SeparableConv2D(32, kernel_size=(3, 3), strides=(2,2), padding='same', depthwise_regularizer=l2(1e-5), pointwise_regularizer=l2(1e-5), use_bias=False)(im_in)
x3 = BatchNormalization()(x3)
x3 = Activation('relu')(x3)
x3 = Attention(32)(x3)
### Block 2 ###
x4 = SeparableConv2D(64, kernel_size=(3, 3), strides=(2,2), padding='same', depthwise_regularizer=l2(1e-5), pointwise_regularizer=l2(1e-5), use_bias=False)(x3)
x4 = BatchNormalization()(x4)
x4 = Activation('relu')(x4)
x4 = Attention(64)(x4)
### Block 3 ###
x5 = SeparableConv2D(128, kernel_size=(3, 3), strides=(2,2), padding='same', depthwise_regularizer=l2(1e-5), pointwise_regularizer=l2(1e-5), use_bias=False)(x4)
x5 = BatchNormalization()(x5)
x5 = Activation('relu')(x5)
x5 = Attention(128)(x5)
### final stage ###
x6 = Conv2D(576, kernel_size=1, strides=1, padding='valid', kernel_regularizer=l2(1e-5), use_bias=False)(x5)
x6 = BatchNormalization()(x6)
x6 = Activation('relu')(x6)
x6 = GlobalAveragePooling2D()(x6)
######## final addition #########
x2 = Add()([x2, x6])
x2 = Dense(2)(x2)
x2 = Activation('softmax')(x2)
model_top = Model(inputs=im_in, outputs=x2)
model_top.summary()
# optimizer = SGD(lr=1e-3, momentum=0.9, nesterov=True)
optimizer = Adam()
model_top.compile(optimizer=optimizer, loss='categorical_crossentropy', metrics=['acc'])
callback_list = [EarlyStopping(monitor='val_acc', patience=30),
ReduceLROnPlateau(monitor='loss', factor=np.sqrt(0.5), cooldown=0, patience=5, min_lr=0.5e-5)]
output = model_top.fit_generator(ft_gen, steps_per_epoch=200, epochs=300,
validation_data=validation_generator, validation_steps=len(validation_generator), callbacks=callback_list)
from tqdm import trange
output_score50 = []
output_class50 = []
answer_class50 = []
answer_class50_1 =[]
for i in trange(len(test50_generator)):
output50 = model_top.predict_on_batch(test50_generator[i][0])
output_score50.append(output50)
answer_class50.append(test50_generator[i][1])
output_score50 = np.concatenate(output_score50)
answer_class50 = np.concatenate(answer_class50)
output_class50 = np.argmax(output_score50, axis=1)
answer_class50_1 = np.argmax(answer_class50, axis=1)
print(output_class50)
print(answer_class50_1)
cm50 = confusion_matrix(answer_class50_1, output_class50)
report50 = classification_report(answer_class50_1, output_class50)
recall50 = cm50[0][0] / (cm50[0][0] + cm50[0][1])
fallout50 = cm50[1][0] / (cm50[1][0] + cm50[1][1])
fpr50, tpr50, thresholds50 = roc_curve(answer_class50_1, output_score50[:, 1], pos_label=1.)
eer50 = brentq(lambda x : 1. - x - interp1d(fpr50, tpr50)(x), 0., 1.)
thresh50 = interp1d(fpr50, thresholds50)(eer50)
print(report50)
print(cm50)
print("AUROC: %f" %(roc_auc_score(answer_class50_1, output_score50[:, 1])))
print(thresh50)
print('test_acc: ', len(output_class50[np.equal(output_class50, answer_class50_1)]) / len(output_class50))
model_top.save('/home/www/fake_detection/model/face2face_resnetv2_ft.h5')
```
| github_jupyter |
```
#convert
```
# babilim.training.losses
> A package containing all losses.
```
#export
from collections import defaultdict
from typing import Any
import json
import numpy as np
import babilim
from babilim.core.itensor import ITensor
from babilim.core.logging import info
from babilim.core.tensor import Tensor
from babilim.core.module import Module
#export
class Loss(Module):
def __init__(self, reduction: str = "mean"):
"""
A loss is a statefull object which computes the difference between the prediction and the target.
:param log_std: When true the loss will log its standard deviation. (default: False)
:param log_min: When true the loss will log its minimum values. (default: False)
:param log_max: When true the loss will log its maximal values. (default: False)
:param reduction: Specifies the reduction to apply to the output: `'none'` | `'mean'` | `'sum'`. `'none'`: no reduction will be applied, `'mean'`: the sum of the output will be divided by the number of elements in the output, `'sum'`: the output will be summed. Default: 'mean'.
"""
super().__init__()
self._accumulators = defaultdict(list)
self.reduction = reduction
if reduction not in ["none", "mean", "sum"]:
raise NotImplementedError()
def call(self, y_pred: Any, y_true: Any) -> ITensor:
"""
Implement a loss function between preds and true outputs.
**DO NOT**:
* Overwrite this function (overwrite `self.loss(...)` instead)
* Call this function (call the module instead `self(y_pred, y_true)`)
Arguments:
:param y_pred: The predictions of the network. Either a NamedTuple pointing at ITensors or a Dict or Tuple of ITensors.
:param y_true: The desired outputs of the network (labels). Either a NamedTuple pointing at ITensors or a Dict or Tuple of ITensors.
"""
loss = self.loss(y_pred, y_true)
if loss.is_nan().any():
raise ValueError("Loss is nan. Loss value: {}".format(loss))
if self.reduction == "mean":
loss = loss.mean()
elif self.reduction == "sum":
loss = loss.sum()
return loss
def loss(self, y_pred: Any, y_true: Any) -> ITensor:
"""
Implement a loss function between preds and true outputs.
**`loss` must be overwritten by subclasses.**
**DO NOT**:
* Call this function (call the module instead `self(y_pred, y_true)`)
Arguments:
:param y_pred: The predictions of the network. Either a NamedTuple pointing at ITensors or a Dict or Tuple of ITensors.
:param y_true: The desired outputs of the network (labels). Either a NamedTuple pointing at ITensors or a Dict or Tuple of ITensors.
"""
raise NotImplementedError("Every loss must implement the call method.")
def log(self, name: str, value: ITensor) -> None:
"""
Log a tensor under a name.
These logged values then can be used for example by tensorboard loggers.
:param name: The name under which to log the tensor.
:param value: The tensor that should be logged.
"""
if isinstance(value, ITensor):
val = value.numpy()
if len(val.shape) > 0:
self._accumulators[name].append(val)
else:
self._accumulators[name].append(np.array([val]))
else:
self._accumulators[name].append(np.array([value]))
def reset_avg(self) -> None:
"""
Reset the accumulation of tensors in the logging.
Should only be called by a tensorboard logger.
"""
self._accumulators = defaultdict(list)
def summary(self, samples_seen, summary_writer=None, summary_txt=None, log_std=False, log_min=False, log_max=False) -> None:
"""
Write a summary of the accumulated logs into tensorboard.
:param samples_seen: The number of samples the training algorithm has seen so far (not iterations!).
This is used for the x axis in the plot. If you use the samples seen it is independant of the batch size.
If the network was trained for 4 batches with 32 batch size or for 32 batches with 4 batchsize does not matter.
:param summary_writer: The summary writer to use for writing the summary. If none is provided it will use the tensorflow default.
:param summary_txt: The file where to write the summary in csv format.
"""
results = {}
if summary_writer is not None:
for k in self._accumulators:
if not self._accumulators[k]:
continue
combined = np.concatenate(self._accumulators[k], axis=0)
summary_writer.add_scalar("{}".format(k), combined.mean(), global_step=samples_seen)
results[f"{k}"] = combined.mean()
if log_std:
results[f"{k}_std"] = combined.std()
summary_writer.add_scalar("{}_std".format(k), results[f"{k}_std"], global_step=samples_seen)
if log_min:
results[f"{k}_min"] = combined.min()
summary_writer.add_scalar("{}_min".format(k), results[f"{k}_min"], global_step=samples_seen)
if log_max:
results[f"{k}_max"] = combined.max()
summary_writer.add_scalar("{}_max".format(k), results[f"{k}_max"], global_step=samples_seen)
else:
import tensorflow as tf
for k in self._accumulators:
if not self._accumulators[k]:
continue
combined = np.concatenate(self._accumulators[k], axis=0)
tf.summary.scalar("{}".format(k), combined.mean(), step=samples_seen)
results[f"{k}"] = combined.mean()
if log_std:
results[f"{k}_std"] = combined.std()
tf.summary.scalar("{}_std".format(k), results[f"{k}_std"], step=samples_seen)
if log_min:
results[f"{k}_min"] = combined.min()
tf.summary.scalar("{}_min".format(k), results[f"{k}_min"], step=samples_seen)
if log_max:
results[f"{k}_max"] = combined.max()
tf.summary.scalar("{}_max".format(k), results[f"{k}_max"], step=samples_seen)
if summary_txt is not None:
results["samples_seen"] = samples_seen
for k in results:
results[k] = f"{results[k]:.5f}"
with open(summary_txt, "a") as f:
f.write(json.dumps(results)+"\n")
@property
def avg(self):
"""
Get the average of the loged values.
This is helpfull to print values that are more stable than values from a single iteration.
"""
avgs = {}
for k in self._accumulators:
if not self._accumulators[k]:
continue
combined = np.concatenate(self._accumulators[k], axis=0)
avgs[k] = combined.mean()
return avgs
#export
class NativeLossWrapper(Loss):
def __init__(self, loss, reduction: str = "mean"):
"""
Wrap a native loss as a babilim loss.
The wrapped object must have the following signature:
```python
Callable(y_pred, y_true, log_val) -> Tensor
```
where log_val will be a function which can be used for logging scalar tensors/values.
:param loss: The loss that should be wrapped.
:param reduction: Specifies the reduction to apply to the output: `'none'` | `'mean'` | `'sum'`. `'none'`: no reduction will be applied, `'mean'`: the sum of the output will be divided by the number of elements in the output, `'sum'`: the output will be summed. Default: 'mean'.
"""
super().__init__(reduction=reduction)
self.native_loss = loss
self._auto_device()
def _auto_device(self):
if babilim.is_backend(babilim.PYTORCH_BACKEND):
import torch
self.native_loss = self.native_loss.to(torch.device(self.device))
return self
def loss(self, y_pred: Any, y_true: Any) -> ITensor:
"""
Compute the loss using the native loss function provided in the constructor.
:param y_pred: The predictions of the network. Either a NamedTuple pointing at ITensors or a Dict or Tuple of ITensors.
:param y_true: The desired outputs of the network (labels). Either a NamedTuple pointing at ITensors or a Dict or Tuple of ITensors.
"""
# Unwrap arguments
tmp = y_true._asdict()
y_true_tmp = {k: tmp[k].native for k in tmp}
y_true = type(y_true)(**y_true_tmp)
tmp = y_pred._asdict()
y_pred_tmp = {k: tmp[k].native for k in tmp}
y_pred = type(y_pred)(**y_pred_tmp)
# call function
result = self.native_loss(y_pred=y_pred, y_true=y_true,
log_val=lambda name, tensor: self.log(name, Tensor(data=tensor, trainable=True)))
return Tensor(data=result, trainable=True)
#export
class SparseCrossEntropyLossFromLogits(Loss):
def __init__(self, reduction: str = "mean"):
"""
Compute a sparse cross entropy.
This means that the preds are logits and the targets are not one hot encoded.
:param reduction: Specifies the reduction to apply to the output: `'none'` | `'mean'` | `'sum'`. `'none'`: no reduction will be applied, `'mean'`: the sum of the output will be divided by the number of elements in the output, `'sum'`: the output will be summed. Default: 'mean'.
"""
super().__init__(reduction=reduction)
if babilim.is_backend(babilim.PYTORCH_BACKEND):
from torch.nn import CrossEntropyLoss
self.loss_fun = CrossEntropyLoss(reduction="none")
else:
from tensorflow.nn import sparse_softmax_cross_entropy_with_logits
self.loss_fun = sparse_softmax_cross_entropy_with_logits
def loss(self, y_pred: ITensor, y_true: ITensor) -> ITensor:
"""
Compute the sparse cross entropy assuming y_pred to be logits.
:param y_pred: The predictions of the network. Either a NamedTuple pointing at ITensors or a Dict or Tuple of ITensors.
:param y_true: The desired outputs of the network (labels). Either a NamedTuple pointing at ITensors or a Dict or Tuple of ITensors.
"""
y_true = y_true.cast("int64")
if babilim.is_backend(babilim.PYTORCH_BACKEND):
return Tensor(data=self.loss_fun(y_pred.native, y_true.native), trainable=True)
else:
return Tensor(data=self.loss_fun(labels=y_true.native, logits=y_pred.native), trainable=True)
#export
class BinaryCrossEntropyLossFromLogits(Loss):
def __init__(self, reduction: str = "mean"):
"""
Compute a binary cross entropy.
This means that the preds are logits and the targets are a binary (1 or 0) tensor of same shape as logits.
:param reduction: Specifies the reduction to apply to the output: `'none'` | `'mean'` | `'sum'`. `'none'`: no reduction will be applied, `'mean'`: the sum of the output will be divided by the number of elements in the output, `'sum'`: the output will be summed. Default: 'mean'.
"""
super().__init__(reduction=reduction)
if babilim.is_backend(babilim.PYTORCH_BACKEND):
from torch.nn import BCEWithLogitsLoss
self.loss_fun = BCEWithLogitsLoss(reduction="none")
else:
from tensorflow.nn import sigmoid_cross_entropy_with_logits
self.loss_fun = sigmoid_cross_entropy_with_logits
def loss(self, y_pred: ITensor, y_true: ITensor) -> ITensor:
"""
Compute the sparse cross entropy assuming y_pred to be logits.
:param y_pred: The predictions of the network. Either a NamedTuple pointing at ITensors or a Dict or Tuple of ITensors.
:param y_true: The desired outputs of the network (labels). Either a NamedTuple pointing at ITensors or a Dict or Tuple of ITensors.
"""
if babilim.is_backend(babilim.PYTORCH_BACKEND):
return Tensor(data=self.loss_fun(y_pred.native, y_true.native), trainable=True)
else:
return Tensor(data=self.loss_fun(labels=y_true.native, logits=y_pred.native), trainable=True)
#export
class SmoothL1Loss(Loss):
def __init__(self, reduction: str = "mean"):
"""
Compute a binary cross entropy.
This means that the preds are logits and the targets are a binary (1 or 0) tensor of same shape as logits.
:param reduction: Specifies the reduction to apply to the output: `'none'` | `'mean'` | `'sum'`. `'none'`: no reduction will be applied, `'mean'`: the sum of the output will be divided by the number of elements in the output, `'sum'`: the output will be summed. Default: 'mean'.
"""
super().__init__(reduction=reduction)
if babilim.is_backend(babilim.PYTORCH_BACKEND):
from torch.nn import SmoothL1Loss
self.loss_fun = SmoothL1Loss(reduction="none")
else:
from tensorflow.keras.losses import huber
self.loss_fun = huber
self.delta = 1.0
def loss(self, y_pred: ITensor, y_true: ITensor) -> ITensor:
"""
Compute the sparse cross entropy assuming y_pred to be logits.
:param y_pred: The predictions of the network. Either a NamedTuple pointing at ITensors or a Dict or Tuple of ITensors.
:param y_true: The desired outputs of the network (labels). Either a NamedTuple pointing at ITensors or a Dict or Tuple of ITensors.
"""
if babilim.is_backend(babilim.PYTORCH_BACKEND):
return Tensor(data=self.loss_fun(y_pred.native, y_true.native), trainable=True)
else:
return Tensor(data=self.loss_fun(labels=y_true.native, logits=y_pred.native, delta=self.delta), trainable=True)
#export
class MeanSquaredError(Loss):
def __init__(self, reduction: str = "mean"):
"""
Compute the mean squared error.
:param reduction: Specifies the reduction to apply to the output: `'none'` | `'mean'` | `'sum'`. `'none'`: no reduction will be applied, `'mean'`: the sum of the output will be divided by the number of elements in the output, `'sum'`: the output will be summed. Default: 'mean'.
"""
super().__init__(reduction=reduction)
def loss(self, y_pred: ITensor, y_true: ITensor, axis: int=-1) -> ITensor:
"""
Compute the mean squared error.
:param y_pred: The predictions of the network. Either a NamedTuple pointing at ITensors or a Dict or Tuple of ITensors.
:param y_true: The desired outputs of the network (labels). Either a NamedTuple pointing at ITensors or a Dict or Tuple of ITensors.
:param axis: (Optional) The axis along which to compute the mean squared error.
"""
return ((y_pred - y_true) ** 2).mean(axis=axis)
#export
class SparseCategoricalAccuracy(Loss):
def __init__(self, reduction: str = "mean"):
"""
Compute the sparse mean squared error.
Sparse means that the targets are not one hot encoded.
:param reduction: Specifies the reduction to apply to the output: `'none'` | `'mean'` | `'sum'`. `'none'`: no reduction will be applied, `'mean'`: the sum of the output will be divided by the number of elements in the output, `'sum'`: the output will be summed. Default: 'mean'.
"""
super().__init__(reduction=reduction)
def loss(self, y_pred: ITensor, y_true: ITensor, axis: int=-1) -> ITensor:
"""
Compute the sparse categorical accuracy.
:param y_pred: The predictions of the network. Either a NamedTuple pointing at ITensors or a Dict or Tuple of ITensors.
:param y_true: The desired outputs of the network (labels). Either a NamedTuple pointing at ITensors or a Dict or Tuple of ITensors.
:param axis: (Optional) The axis along which to compute the sparse categorical accuracy.
"""
pred_class = y_pred.argmax(axis=axis)
true_class = y_true.cast("int64")
correct_predictions = pred_class == true_class
return correct_predictions.cast("float32").mean(axis=axis)
#export
class NaNMaskedLoss(Loss):
def __init__(self, loss):
"""
Compute a sparse cross entropy.
This means that the preds are logits and the targets are not one hot encoded.
:param loss: The loss that should be wrapped and only applied on non nan values.
"""
super().__init__(reduction="none")
self.wrapped_loss = loss
self.zero = Tensor(data=np.array(0), trainable=False)
def loss(self, y_pred: ITensor, y_true: ITensor) -> ITensor:
"""
Compute the loss given in the constructor only on values where the GT is not NaN.
:param y_pred: The predictions of the network. Either a NamedTuple pointing at ITensors or a Dict or Tuple of ITensors.
:param y_true: The desired outputs of the network (labels). Either a NamedTuple pointing at ITensors or a Dict or Tuple of ITensors.
"""
binary_mask = (~y_true.is_nan())
mask = binary_mask.cast("float32")
masked_y_true = (y_true * mask)[binary_mask]
if y_pred.shape[-1] != binary_mask.shape[-1] and binary_mask.shape[-1] == 1:
new_shape = binary_mask.shape[:-1]
binary_mask = binary_mask.reshape(new_shape)
masked_y_pred = (y_pred * mask)[binary_mask]
if masked_y_pred.shape[0] > 0:
loss = self.wrapped_loss(masked_y_pred, masked_y_true)
else:
loss = self.zero
return loss
```
| github_jupyter |
<a href="https://colab.research.google.com/github/kzafeiroudi/QuestRecommend/blob/master/Preprocessing_SQuAD.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Preprocessing the SQuAD dataset
## The dataset
After looking into the Quora Question Pairs dataset, it is now time to apply the deveoped process on the SQuAD dataset. The part of the SQuAD dataset that we are going to use here consists of:
* 3111 unique questions
* 30 unique article topics from Wikipedia
Each instance of this part of the dataset has the following attributes:
* sentID : A unique identifier for each question
* Question: The full text of the question
* Class: The article title of the Wikipedia page the question was derived from, used here as a nominal class
## Upload the dataset
Upload the file `squad.csv` that will be used in this Python 3 notebook.
```
from google.colab import files
# Choose from your own machine the file to upload - name should be "squad.csv"
uploaded = files.upload()
```
## Download a pre-trained English language model
We will be using the large pre-trained statistical model for English, available by the **spaCy** free open-source library for NLP in Python. Find more [here](https://spacy.io/models/en#en_core_web_lg).
```
!python -m spacy download en_core_web_lg
# Load the model
import spacy
nlp = spacy.load('en_core_web_lg')
```
## Importing Python libraries
```
import csv
import numpy as np
import random
from prettytable import PrettyTable
```
## Load the dataset
```
# Loading the SQuAD dataset
data_file = 'squad.csv'
new_data = []
with open(data_file) as csvfile:
reader = csv.DictReader(csvfile)
for row in reader:
new_data.append(row)
# Extracting the unique questions from the dataset, and all unique topics
uniqueQ = []
topics = {}
for dd in new_data:
uniqueQ.append(dd['Question'])
if (dd['Class'] in topics):
topics[dd['Class']].append(dd['Question'])
else:
topics[dd['Class']] = [dd['Question']]
# Print stats for the dataset
print('Unique Questions: ', len(uniqueQ))
print('Unique Topics: ', len(topics.keys()))
print()
# Print stats for each topic
t = PrettyTable(['Topic', '# of Questions'])
for tt in topics:
t.add_row([tt, len(topics[tt])])
print(t)
```
## Implementing a cosine similarity function
`cosine(vA, vB)` calculates the cosine similarity between two vectors `vA` and `vB`.
```
def cosine(vA, vB):
cos = np.dot(vA, vB) / (np.sqrt(np.dot(vA, vA)) * np.sqrt(np.dot(vB, vB)))
return cos.astype('float64')
```
## Calculating the question vector representation
We will use only the main verb arguments of the question to calculate the question embeddings, since the process we followed for the Quora Question Pairs dataset showed that question are less likely to be considered duplicates when they are not.
The final result is saved in the file `squad_vectors.csv`, which will then be used to perform classification and clustering.
```
squad = []
for i in range(len(new_data)):
foo = nlp(new_data[i]['Question'])
calc = [np.zeros(300, dtype='float64')]
for nn in foo.noun_chunks:
calc.append(nn.vector)
vA = np.mean(calc, axis=0).astype('float64')
squad.append(vA)
toCSV = []
for i in range(len(new_data)):
boo = {}
boo['Class'] = new_data[i]['Class']
for j in range(300):
boo['vector_' + str(j+1)] = squad[i][j]
toCSV.append(boo)
with open('squad_vectors.csv', 'w') as csvfile:
fieldnames = list(boo.keys())
writer = csv.DictWriter(csvfile, fieldnames=fieldnames)
writer.writeheader()
for i in range(len(toCSV)):
writer.writerow(toCSV[i])
```
## Testing whether the dataset is large enough for the task
In this section, we calculate the mean vector representation across all questions that belong in the same class, showcasing that these vectors do not coincide, and thus we have a heterogenous dataset good enough for the task.
As we can see, by running this code, there is no occurence of *Same means* in the standard output.
```
# cl: all different classes
classes = list(topics.keys())
# calc: to calculate the mean vector representation per class
calc = {}
for cc in classes:
calc[cc] = []
for i in range(len(new_data)):
calc[new_data[i]['Class']].append(squad[i])
# means: to store the mean vector representation of each class
means = {}
for cc in classes:
means[cc] = np.mean(calc[cc], axis=0)
for i in range(len(classes)):
for j in range(i+1, len(classes)):
if (any(means[keys[i]] == means[keys[j]])):
print('Same means')
```
## Evaluating the results if we were to rely completely on the cosine similarity
For this task, we will pick randomly 100 questions, we will calculate the vector representation on the full body of the question, and by calculating the cosine similarity between split1 (34 questions) and split2 (66 questions), we will show the questions that is suggestive to group together due to the process we have developed so far.
```
# Shuffling the sequence of questions
new = list(range(len(new_data)))
random.shuffle(new)
# Picking randomly 340 for split1
# and 660 for split2
split1 = {}
split2 = {}
for i in new[:340]:
q = new_data[i]['Question']
split1[q] = {'vec': nlp(q), 'class' : new_data[i]['Class']}
for i in new[340:1000]:
q = new_data[i]['Question']
split2[q] = {'vec': nlp(q), 'class' : new_data[i]['Class']}
# Calculate the question vector on the full body of the question
# and save the questions from split2 that are closely related to
# questions from split1 (cosine similarity > 0.93 as per our first
# classification task)
rank = {}
for tt in split1:
rank[tt] = []
vA = split1[tt]['vec'].vector
for tr in split2:
vB = split2[tr]['vec'].vector
cos = cosine(vA, vB)
if (cos > 0.93):
rank[tt].append(tr)
for q in rank:
if(rank[q] != []):
print('Input Question:')
print('\t-',q)
print('Related Questions:')
for rr in rank[q]:
print('\t-', rr)
print()
```
## Upload the context data
```
# Choose from your own machine the file to upload - name should be "squad_contexts.csv"
uploaded = files.upload()
```
## Load the context data
```
# Loading the SQuAD context dataset
data_file = 'squad_contexts.csv'
new_data = []
with open(data_file) as csvfile:
reader = csv.DictReader(csvfile)
for row in reader:
new_data.append(row)
# Extracting the unique classes/topics from the dataset, and all contexts
contexts = {}
for dd in new_data:
if (dd['Superclass'] in contexts):
contexts[dd['Superclass']].append(dd['Context'])
else:
contexts[dd['Superclass']] = [dd['Context']]
# Print stats for the dataset
print('Unique Topics: ', len(contexts))
print()
# Print stats for each topic
t = PrettyTable(['Topic', '# of Contexts'])
for tt in contexts:
t.add_row([tt, len(contexts[tt])])
print(t)
```
## Calculating the context vector representation
We will use only the main verb arguments of the context to calculate the context embeddings, as before.
The final result is saved in the file `squad_contextsVectors.csv`, which will then be used to perform classification and clustering.
```
squad_cont = []
for i in range(len(new_data)):
foo = nlp(new_data[i]['Context'])
calc = [np.zeros(300, dtype='float64')]
for nn in foo.noun_chunks:
calc.append(nn.vector)
vA = np.mean(calc, axis=0).astype('float64')
squad_cont.append(vA)
toCSV = []
for i in range(len(new_data)):
boo = {}
boo['Class'] = new_data[i]['Superclass']
for j in range(300):
boo['vector_' + str(j+1)] = squad_cont[i][j]
toCSV.append(boo)
with open('squad_contextsVectors.csv', 'w') as csvfile:
fieldnames = list(boo.keys())
writer = csv.DictWriter(csvfile, fieldnames=fieldnames)
writer.writeheader()
for i in range(len(toCSV)):
writer.writerow(toCSV[i])
```
| github_jupyter |
Copyright (c) Microsoft Corporation. All rights reserved.
Licensed under the MIT License.
# Inference Bert Model for High Performance with ONNX Runtime on AzureML #
This tutorial includes how to pretrain and finetune Bert models using AzureML, convert it to ONNX, and then deploy the ONNX model with ONNX Runtime through Azure ML. In the following sections, we are going to use the Bert model trained with Stanford Question Answering Dataset (SQuAD) dataset as an example. Bert SQuAD model is used in question answering scenarios, where the answer to every question is a segment of text, or span, from the corresponding reading passage, or the question might be unanswerable.
## Roadmap
0. **Prerequisites** to set up your Azure ML work environments.
1. **Pre-train, finetune and export Bert model** from other framework using Azure ML.
2. **Deploy Bert model using ONNX Runtime and AzureML**
## Step 0 - Prerequisites
If you are using an [Azure Machine Learning Notebook VM](https://docs.microsoft.com/en-us/azure/machine-learning/service/quickstart-run-cloud-notebook), you are all set. Otherwise, refer to the [configuration Notebook](https://github.com/Azure/MachineLearningNotebooks/blob/56e0ebc5acb9614fac51d8b98ede5acee8003820/configuration.ipynb) first if you haven't already to establish your connection to the AzureML Workspace. Prerequisites are:
* Azure subscription
* Azure Machine Learning Workspace
* Azure Machine Learning SDK
Also to make the best use of your time, make sure you have done the following:
* Understand the [architecture and terms](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture) introduced by Azure Machine Learning
* [Azure Portal](https://portal.azure.com) allows you to track the status of your deployments.
## Step 1 - Pretrain, Finetune and Export Bert Model (PyTorch)
If you'd like to pre-train and finetune a Bert model from scratch, follow the instructions in [
Pretraining of the BERT model](https://github.com/microsoft/AzureML-BERT/blob/master/pretrain/PyTorch/notebooks/BERT_Pretrain.ipynb) to pretrain a Bert model in PyTorch using AzureML. Once you have the pretrained model, refer to [AzureML Bert Eval Squad](https://github.com/microsoft/AzureML-BERT/blob/master/finetune/PyTorch/notebooks/BERT_Eval_SQUAD.ipynb) or [AzureML Bert Eval GLUE](https://github.com/microsoft/AzureML-BERT/blob/master/finetune/PyTorch/notebooks/BERT_Eval_GLUE.ipynb) to finetune your model with your desired dataset. Follow the tutorials all the way through **Create a PyTorch estimator for fine-tuning**. Before creating a Pytorch estimator, we need to prepare an entry file that trains and exports the PyTorch model together. Make sure the entry file has the following code to create an ONNX file:
```
output_model_path = "bert_azureml_large_uncased.onnx"
# set the model to inference mode
# It is important to call torch_model.eval() or torch_model.train(False) before exporting the model,
# to turn the model to inference mode. This is required since operators like dropout or batchnorm
# behave differently in inference and training mode.
model.eval()
# Generate dummy inputs to the model. Adjust if neccessary
inputs = {
'input_ids': torch.randint(32, [2, 32], dtype=torch.long).to(device), # list of numerical ids for the tokenised text
'attention_mask': torch.ones([2, 32], dtype=torch.long).to(device), # dummy list of ones
'token_type_ids': torch.ones([2, 32], dtype=torch.long).to(device), # dummy list of ones
}
symbolic_names = {0: 'batch_size', 1: 'max_seq_len'}
torch.onnx.export(model, # model being run
(inputs['input_ids'],
inputs['attention_mask'],
inputs['token_type_ids']), # model input (or a tuple for multiple inputs)
output_model_path, # where to save the model (can be a file or file-like object)
opset_version=11, # the ONNX version to export the model to
do_constant_folding=True, # whether to execute constant folding for optimization
input_names=['input_ids',
'input_mask',
'segment_ids'], # the model's input names
output_names=['start', "end"], # the model's output names
dynamic_axes={'input_ids': symbolic_names,
'input_mask' : symbolic_names,
'segment_ids' : symbolic_names,
'start' : symbolic_names,
'end': symbolic_names}) # variable length axes
```
In this directory, a `run_squad_azureml.py` containing the above code is available for use. Copy the training script `run_squad_azureml.py` to your `project_root` (defined at an earlier step in [AzureML Bert Eval Squad](https://github.com/microsoft/AzureML-BERT/blob/master/finetune/PyTorch/notebooks/BERT_Eval_SQUAD.ipynb))
```
shutil.copy('run_squad_azureml.py', project_root)
```
Now you may continue to follow the **Create a PyTorch estimator for fine-tuning** section in [AzureML Bert Eval Squad](https://github.com/microsoft/AzureML-BERT/blob/master/finetune/PyTorch/notebooks/BERT_Eval_SQUAD.ipynb). In creating the estimator, change `entry_script` parameter to point to the `run_squad_azureml.py` we just copied as noted in the following code.
```
estimator = PyTorch(source_directory=project_roots,
script_params={'--output-dir': './outputs'},
compute_target=gpu_compute_target,
use_docker=True
custom_docker_image=image_name
script_params = {...}
entry_script='run_squad_azureml.py', # change here
node_count=1,
process_count_per_node=4,
distributed_backend='mpi',
use_gpu=True)
```
Follow the rest of the [AzureML Bert Eval Squad](https://github.com/microsoft/AzureML-BERT/blob/master/finetune/PyTorch/notebooks/BERT_Eval_SQUAD.ipynb) to run and export your model.
## Step 2 - Deploy Bert model with ONNX Runtime through AzureML
In Step 1 and 2, we have prepared an optimized ONNX Bert model and now we can deploy this model as a web service using Azure Machine Learning services and the ONNX Runtime.
We're now going to deploy our ONNX model on Azure ML using the following steps.
1. **Register our model** in our Azure Machine Learning workspace
2. **Write a scoring file** to evaluate our model with ONNX Runtime
3. **Write environment file** for our Docker container image.
4. **Deploy to the cloud** using an Azure Container Instances VM and use it to make predictions using ONNX Runtime Python APIs
5. **Classify sample text input** so we can explore inference with our deployed service.

## Step 2.0 - Check your AzureML environment
```
# Check core SDK version number
import azureml.core
from PIL import Image, ImageDraw, ImageFont
import json
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
print("SDK version:", azureml.core.VERSION)
```
### Load your Azure ML workspace
We begin by instantiating a workspace object from the existing workspace created earlier in the configuration notebook.
```
from azureml.core import Workspace
ws = Workspace.from_config()
print(ws.name, ws.location, ws.resource_group, sep = '\n')
```
## Step 2.1 - Register your model with Azure ML
Now we upload the model and register it in the workspace. In the following tutorial. we use the bert SQuAD model outputted from Step 1 as an example.
You can also register the model from your run to your workspace. The model_path parameter takes in the relative path on the remote VM to the model file in your outputs directory. You can then deploy this registered model as a web service through the AML SDK.
```
model = run.register_model(model_path = "./bert_azureml_large_uncased.onnx", # Name of the registered model in your workspace.
model_name = "bert-squad-large-uncased", # Local ONNX model to upload and register as a model
model_framework=Model.Framework.ONNX , # Framework used to create the model.
model_framework_version='1.6', # Version of ONNX used to create the model.
tags = {"onnx": "demo"},
description = "Bert-large-uncased squad model exported from PyTorch",
workspace = ws)
```
Alternatively, if you're working on a local model and want to deploy it to AzureML, upload your model to the same directory as this notebook and register it with `Model.register()`
```
from azureml.core.model import Model
model = Model.register(model_path = "./bert_azureml_large_uncased.onnx", # Name of the registered model in your workspace.
model_name = "bert-squad-large-uncased", # Local ONNX model to upload and register as a model
model_framework=Model.Framework.ONNX , # Framework used to create the model.
model_framework_version='1.6', # Version of ONNX used to create the model.
tags = {"onnx": "demo"},
description = "Bert-large-uncased squad model exported from PyTorch",
workspace = ws)
```
#### Displaying your registered models
You can optionally list out all the models that you have registered in this workspace.
```
models = ws.models
for name, m in models.items():
print("Name:", name,"\tVersion:", m.version, "\tDescription:", m.description, m.tags)
# # If you'd like to delete the models from workspace
# model_to_delete = Model(ws, name)
# model_to_delete.delete()
```
## Step 2.2 - Write scoring file
We are now going to deploy our ONNX model on Azure ML using the ONNX Runtime. We begin by writing a score.py file that will be invoked by the web service call. The `init()` function is called once when the container is started so we load the model using the ONNX Runtime into a global session object. Then the `run()` function is called when we run the model using the Azure ML web service. Add neccessary `preprocess()` and `postprocess()` steps. The following score.py file uses `bert-squad` as an example and assumes the inputs will be in the following format.
```
inputs_json = {
"version": "1.4",
"data": [
{
"paragraphs": [
{
"context": "In its early years, the new convention center failed to meet attendance and revenue expectations.[12] By 2002, many Silicon Valley businesses were choosing the much larger Moscone Center in San Francisco over the San Jose Convention Center due to the latter's limited space. A ballot measure to finance an expansion via a hotel tax failed to reach the required two-thirds majority to pass. In June 2005, Team San Jose built the South Hall, a $6.77 million, blue and white tent, adding 80,000 square feet (7,400 m2) of exhibit space",
"qas": [
{
"question": "where is the businesses choosing to go?",
"id": "1"
},
{
"question": "how may votes did the ballot measure need?",
"id": "2"
},
{
"question": "When did businesses choose Moscone Center?",
"id": "3"
}
]
}
],
"title": "Conference Center"
}
]
}
%%writefile score.py
import os
import collections
import json
import time
from azureml.core.model import Model
import numpy as np # we're going to use numpy to process input and output data
import onnxruntime # to inference ONNX models, we use the ONNX Runtime
import wget
from pytorch_pretrained_bert.tokenization import whitespace_tokenize, BasicTokenizer, BertTokenizer
def init():
global session, tokenizer
# use AZUREML_MODEL_DIR to get your deployed model(s). If multiple models are deployed,
# model_path = os.path.join(os.getenv('AZUREML_MODEL_DIR'), '$MODEL_NAME/$VERSION/$MODEL_FILE_NAME')
model_path = os.path.join(os.getenv('AZUREML_MODEL_DIR'), 'bert_azureml_large_uncased.onnx')
sess_options = onnxruntime.SessionOptions()
# Set graph optimization level to ORT_ENABLE_EXTENDED to enable bert optimization
session = onnxruntime.InferenceSession(model_path, sess_options)
tokenizer = BertTokenizer.from_pretrained("bert-large-uncased", do_lower_case=True)
# download run_squad.py and tokenization.py from
# https://github.com/onnx/models/tree/master/text/machine_comprehension/bert-squad to
# help with preprocessing and post-processing.
if not os.path.exists('./run_onnx_squad.py'):
url = "https://raw.githubusercontent.com/onnx/models/master/text/machine_comprehension/bert-squad/dependencies/run_onnx_squad.py"
wget.download(url, './run_onnx_squad.py')
if not os.path.exists('./tokenization.py'):
url = "https://raw.githubusercontent.com/onnx/models/master/text/machine_comprehension/bert-squad/dependencies/tokenization.py"
wget.download(url, './tokenization.py')
def preprocess(input_data_json):
global all_examples, extra_data
# Model configs. Adjust as needed.
max_seq_length = 128
doc_stride = 128
max_query_length = 64
# Write the input json to file to be used by read_squad_examples()
input_data_file = "input.json"
with open(input_data_file, 'w') as outfile:
json.dump(json.loads(input_data_json), outfile)
from run_onnx_squad import read_squad_examples, convert_examples_to_features
# Use read_squad_examples method from run_onnx_squad to read the input file
all_examples = read_squad_examples(input_file=input_data_file)
# Use convert_examples_to_features method from run_onnx_squad to get parameters from the input
input_ids, input_mask, segment_ids, extra_data = convert_examples_to_features(all_examples, tokenizer,
max_seq_length, doc_stride, max_query_length)
return input_ids, input_mask, segment_ids
def postprocess(all_results):
# postprocess results
from run_onnx_squad import write_predictions
n_best_size = 20
max_answer_length = 30
output_dir = 'predictions'
os.makedirs(output_dir, exist_ok=True)
output_prediction_file = os.path.join(output_dir, "predictions.json")
output_nbest_file = os.path.join(output_dir, "nbest_predictions.json")
# Write the predictions (answers to the questions) in a file.
write_predictions(all_examples, extra_data, all_results,
n_best_size, max_answer_length,
True, output_prediction_file, output_nbest_file)
# Retrieve best results from file.
result = {}
with open(output_prediction_file, "r") as f:
result = json.load(f)
return result
def run(input_data_json):
try:
# load in our data
input_ids, input_mask, segment_ids = preprocess(input_data_json)
RawResult = collections.namedtuple("RawResult", ["unique_id", "start_logits", "end_logits"])
n = len(input_ids)
bs = 1
all_results = []
start = time.time()
for idx in range(0, n):
item = all_examples[idx]
# this is using batch_size=1
# feed the input data as int64
data = {
"segment_ids": segment_ids[idx:idx+bs],
"input_ids": input_ids[idx:idx+bs],
"input_mask": input_mask[idx:idx+bs]
}
result = session.run(["start", "end"], data)
in_batch = result[0].shape[0]
start_logits = [float(x) for x in result[1][0].flat]
end_logits = [float(x) for x in result[0][0].flat]
for i in range(0, in_batch):
unique_id = len(all_results)
all_results.append(RawResult(unique_id=unique_id, start_logits=start_logits, end_logits=end_logits))
end = time.time()
print("total time: {}sec, {}sec per item".format(end - start, (end - start) / len(all_results)))
return {"result": postprocess(all_results),
"total_time": end - start,
"time_per_item": (end - start) / len(all_results)}
except Exception as e:
result = str(e)
return {"error": result}
```
## Step 2.3 - Write Environment File
We create a YAML file that specifies which dependencies we would like to see in our container.
```
from azureml.core.conda_dependencies import CondaDependencies
myenv = CondaDependencies.create(pip_packages=["numpy","onnxruntime","azureml-core", "azureml-defaults", "tensorflow", "wget", "pytorch_pretrained_bert"])
with open("myenv.yml","w") as f:
f.write(myenv.serialize_to_string())
```
We're all set! Let's get our model chugging.
## Step 2.4 - Deploy Model as Webservice on Azure Container Instance
```
from azureml.core.webservice import AciWebservice
from azureml.core.model import InferenceConfig
from azureml.core.environment import Environment
myenv = Environment.from_conda_specification(name="myenv", file_path="myenv.yml")
inference_config = InferenceConfig(entry_script="score.py", environment=myenv)
aciconfig = AciWebservice.deploy_configuration(cpu_cores = 1,
memory_gb = 4,
tags = {'demo': 'onnx'},
description = 'web service for Bert-squad-large-uncased ONNX model')
```
The following cell will likely take a few minutes to run as well.
```
from azureml.core.webservice import Webservice
from random import randint
aci_service_name = 'onnx-bert-squad-large-uncased-'+str(randint(0,100))
print("Service", aci_service_name)
aci_service = Model.deploy(ws,
aci_service_name,
[model],
inference_config,
aciconfig)
aci_service.wait_for_deployment(True)
print(aci_service.state)
```
In case the deployment fails, you can check the logs. Make sure to delete your aci_service before trying again.
```
if aci_service.state != 'Healthy':
# run this command for debugging.
print(aci_service.get_logs())
aci_service.delete()
```
## Success!
If you've made it this far, you've deployed a working web service that does image classification using an ONNX model. You can get the URL for the webservice with the code below.
```
print(aci_service.scoring_uri)
```
## Step 2.5 - Inference Bert Model using our WebService
**Input**: Context paragraph and questions as formatted in `inputs.json`
**Task**: For each question about the context paragraph, the model predicts a start and an end token from the paragraph that most likely answers the questions.
**Output**: The best answer for each question.
```
# Use the inputs from step 2.2
print("========= INPUT DATA =========")
print(json.dumps(inputs_json, indent=2))
azure_result = aci_service.run(json.dumps(inputs_json))
print("\n")
print("========= RESULT =========")
print(json.dumps(azure_result, indent=2))
res = azure_result['result']
inference_time = np.round(azure_result['total_time'] * 1000, 2)
time_per_item = np.round(azure_result['time_per_item'] * 1000, 2)
print('========================================')
print('Final predictions are: ')
for key in res:
print("Question: ", inputs_json['data'][0]['paragraphs'][0]['qas'][int(key) - 1]['question'])
print("Best Answer: ", res[key])
print()
print('========================================')
print('Inference time: ' + str(inference_time) + " ms")
print('Average inference time for each question: ' + str(time_per_item) + " ms")
print('========================================')
```
When you are eventually done using the web service, remember to delete it.
```
aci_service.delete()
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import re, nltk, spacy, gensim
import en_core_web_sm
from tqdm import tqdm
# Sklearn
from sklearn.decomposition import LatentDirichletAllocation, TruncatedSVD, NMF
from sklearn.feature_extraction.text import CountVectorizer, TfidfVectorizer
from sklearn.model_selection import GridSearchCV
from pprint import pprint
from sklearn.datasets import fetch_20newsgroups
from sklearn.metrics import silhouette_score
# Plotting tools
import pyLDAvis
import pyLDAvis.sklearn
import matplotlib.pyplot as plt
%matplotlib inline
nlp = spacy.load('en_core_web_sm', disable=['parser', 'ner'])
newsgroups_data = fetch_20newsgroups()
data = pd.DataFrame()
data['content'] = newsgroups_data.data[:1000] # take only first 1000 texts
data['label'] = newsgroups_data.target[:1000]
newsgroups_data.target_names
data.head(3)
# Convert to list
texts = data.content.values.tolist()
# Remove Emails
texts = [re.sub('\S*@\S*\s?', '', sent) for sent in texts]
# Remove new line characters
texts = [re.sub('\s+', ' ', sent) for sent in texts]
# Remove distracting single quotes
texts = [re.sub("\'", "", sent) for sent in texts]
pprint(texts[:1])
def sent_to_words(sentences):
for sentence in sentences:
yield(gensim.utils.simple_preprocess(str(sentence), deacc=True)) # deacc=True removes punctuations
data_words = list(sent_to_words(texts))
print(data_words[:1])
def lemmatization(texts, allowed_postags=['NOUN', 'ADJ', 'VERB', 'ADV']):
"""https://spacy.io/api/annotation"""
texts_out = []
for sent in texts:
doc = nlp(" ".join(sent))
texts_out.append(" ".join([token.lemma_ if token.lemma_ not in ['-PRON-'] else '' for token in doc if token.pos_ in allowed_postags]))
return texts_out
# Initialize spacy 'en' model, keeping only tagger component (for efficiency)
# Run in terminal: python3 -m spacy download en
# Do lemmatization keeping only Noun, Adj, Verb, Adverb
data_lemmatized = lemmatization(data_words, allowed_postags=['NOUN', 'ADJ', 'VERB', 'ADV'])
print(data_lemmatized[:2])
def print_top_words(model, feature_names, n_top_words, print_topics=True):
all_top_words = []
for topic_idx, topic in enumerate(model.components_):
message = "Topic #%d: " % topic_idx
message += " ".join([feature_names[i]
for i in topic.argsort()[:-n_top_words - 1:-1]])
if print_topics:
print(message)
all_top_words.append([feature_names[i]
for i in topic.argsort()[:-n_top_words - 1:-1]])
return all_top_words
def coherence_score(tf,vocabulary,all_top_words):
totalcnt = len(all_top_words)
tf = tf > 0
total = 0
count = 0
for i, allwords in enumerate(all_top_words):
for word1 in allwords:
for word2 in allwords:
if word1 != word2:
ind1 = vocabulary[word1]
ind2 = vocabulary[word2]
#print (ind1,ind2)
total += np.log((np.matmul(np.array(tf[:,ind1]).ravel(),np.array(tf[:,ind2]).ravel()) + 1)/np.sum(tf[:,ind2]))
count += 1
return total/count
vectorizer = CountVectorizer(max_df=0.7, min_df=2,
max_features=50000,
stop_words='english')
countvector = vectorizer.fit_transform(data_lemmatized)
countvector = countvector.todense()
vocabulary = dict(zip(vectorizer.get_feature_names(),np.arange(countvector.shape[1])))
print (countvector.shape)
n_topics = 10
nmf = NMF(n_components=n_topics, random_state=123,
alpha=.1, l1_ratio=.5).fit(countvector)
topic_words = print_top_words(nmf, tfidf_vectorizer.get_feature_names(), n_top_words=5)
print (coherence_score(countvector,vocabulary,topic_words))
top_topics = nmf.transform(countvector).argmax(axis=1)
print (silhouette_score(countvector,top_topics))
lda = LatentDirichletAllocation(n_components=n_topics, max_iter=50,
learning_method='online',
learning_offset=50.,
random_state=0)
lda.fit(countvector)
topic_words = print_top_words(lda, tfidf_vectorizer.get_feature_names(), n_top_words=5)
print (coherence_score(countvector,vocabulary,topic_words))
lda.perplexity(countvector)
top_topics = lda.transform(countvector).argmax(axis=1)
print (silhouette_score(countvector,top_topics))
nmf_coh_scores, nmf_sil_scores, lda_coh_scores, lda_sil_scores, lda_perplexity_scores = [], [], [], [], []
for numtopic in tqdm([10,20,50,75,100]):
nmf = NMF(n_components=numtopic, random_state=123,
alpha=.1, l1_ratio=.5).fit(countvector)
topic_words = print_top_words(nmf, tfidf_vectorizer.get_feature_names(), n_top_words=5, print_topics=False)
coh_nmf = coherence_score(countvector,vocabulary,topic_words)
top_topics = nmf.transform(countvector).argmax(axis=1)
sil_nmf = silhouette_score(countvector,top_topics)
lda = LatentDirichletAllocation(n_components=numtopic, max_iter=50,
learning_method='online',
learning_offset=50.,
random_state=0)
lda.fit(countvector)
topic_words = print_top_words(lda, tfidf_vectorizer.get_feature_names(), n_top_words=5, print_topics=False)
coh_lda = coherence_score(countvector,vocabulary,topic_words)
top_topics = lda.transform(countvector).argmax(axis=1)
sil_lda = silhouette_score(countvector,top_topics)
nmf_coh_scores.append(coh_nmf)
nmf_sil_scores.append(sil_nmf)
lda_coh_scores.append(coh_lda)
lda_sil_scores.append(sil_lda)
lda_perplexity_scores.append(lda.perplexity(countvector))
plt.figure(figsize=(12, 8))
plt.plot([10,20,50,75,100], nmf_coh_scores, label='NMF')
plt.plot([10,20,50,75,100], lda_coh_scores, label='LDA')
plt.title("Coherence Scores")
plt.xlabel("Num Topics")
plt.ylabel("UMass Coh Scores")
plt.legend(title='Model', loc='best')
plt.show()
plt.figure(figsize=(12, 8))
plt.plot([10,20,50,75,100], nmf_sil_scores, label='NMF')
plt.plot([10,20,50,75,100], lda_sil_scores, label='LDA')
plt.title("Silhouette Scores")
plt.xlabel("Num Topics")
plt.ylabel("Silhouette Score")
plt.legend(title='Model', loc='best')
plt.show()
plt.figure(figsize=(12, 8))
plt.plot([10,20,50,75,100], lda_perplexity_scores)
plt.title("Perplexity Scores")
plt.xlabel("Num Topics")
plt.ylabel("Perplexity Score")
plt.show()
nmf_sil_scores
lda_sil_scores
lda_perplexity_scores
```
| github_jupyter |
# Comparison of full discretizations using approximate Riemann solvers
In [Euler_approximate_solvers.ipynb](Euler_approximate_solvers.ipynb) we introduced several approximate Riemann solvers for the Euler equations of compressible gas dynamics. How do these solvers impact the solution accuracy when used within a finite volume discretization? To investigate, we will use them within [PyClaw](http://www.clawpack.org/pyclaw/) to solve two standard test problems for one-dimensional compressible flow:
1. The Sod shocktube problem. This is a Riemann problem, which we have considered previously.
2. The Woodward-Colella blast wave problem. The initial data consists of two Riemann problems, with resulting shock waves of differing strengths. These shock waves later interact with each other.
In this chapter, unlike previous chapters, we include extensive sections of code in the notebook. This is meant to more easily allow the reader to use these as templates for setting up other problems. For more information about the software and algorithms used here, see [this paper](https://peerj.com/articles/cs-68/) and references therein.
```
%matplotlib inline
%config InlineBackend.figure_format = 'svg'
import numpy as np
from exact_solvers import euler
from clawpack import riemann
import matplotlib.pyplot as plt
from ipywidgets import interact
from ipywidgets import widgets
from clawpack.riemann.euler_with_efix_1D_constants \
import density, momentum, energy, num_eqn
def shocktube(q_l, q_r, N=50, riemann_solver='HLL',
solver_type='classic'):
from clawpack import pyclaw
from clawpack import riemann
if riemann_solver == 'Roe':
rs = riemann.euler_1D_py.euler_roe_1D
elif riemann_solver == 'HLL':
rs = riemann.euler_1D_py.euler_hll_1D
if solver_type == 'classic':
solver = pyclaw.ClawSolver1D(rs)
solver.limiters = pyclaw.limiters.tvd.MC
else:
solver = pyclaw.SharpClawSolver1D(rs)
solver.kernel_language = 'Python'
solver.bc_lower[0]=pyclaw.BC.extrap
solver.bc_upper[0]=pyclaw.BC.extrap
x = pyclaw.Dimension(-1.0,1.0,N,name='x')
domain = pyclaw.Domain([x])
state = pyclaw.State(domain,num_eqn)
gamma = 1.4
state.problem_data['gamma']= gamma
state.problem_data['gamma1']= gamma-1.
state.problem_data['efix'] = False
xc = state.grid.p_centers[0]
velocity = (xc<=0)*q_l[1] + (xc>0)*q_r[1]
pressure = (xc<=0)*q_l[2] + (xc>0)*q_r[2]
state.q[density ,:] = (xc<=0)*q_l[0] + (xc>0)*q_r[0]
state.q[momentum,:] = velocity * state.q[density,:]
state.q[energy ,:] = pressure/(gamma - 1.) + \
0.5 * state.q[density,:] * velocity**2
claw = pyclaw.Controller()
claw.tfinal = 0.5
claw.solution = pyclaw.Solution(state,domain)
claw.solver = solver
claw.num_output_times = 10
claw.keep_copy = True
claw.verbosity=0
return claw
from clawpack.riemann.euler_with_efix_1D_constants \
import density, momentum, energy, num_eqn
def blastwave(N=400, riemann_solver='HLL', solver_type='classic'):
from clawpack import pyclaw
from clawpack import riemann
if riemann_solver == 'Roe':
kernel_language = 'Fortran'
rs = riemann.euler_with_efix_1D
elif riemann_solver == 'HLL':
kernel_language = 'Python'
rs = riemann.euler_1D_py.euler_hll_1D
if solver_type == 'classic':
solver = pyclaw.ClawSolver1D(rs)
solver.limiters = pyclaw.limiters.tvd.MC
else:
solver = pyclaw.SharpClawSolver1D(rs)
solver.kernel_language = kernel_language
solver.bc_lower[0]=pyclaw.BC.wall
solver.bc_upper[0]=pyclaw.BC.wall
x = pyclaw.Dimension(0.0,1.0,N,name='x')
domain = pyclaw.Domain([x])
state = pyclaw.State(domain,num_eqn)
gamma = 1.4
state.problem_data['gamma']= gamma
state.problem_data['gamma1']= gamma-1.
state.problem_data['efix'] = False
xc = state.grid.p_centers[0]
pressure = (xc<0.1)*1.e3 + (0.1<=xc)*(xc<0.9)*1.e-2 + (0.9<=xc)*1.e2
state.q[density ,:] = 1.
state.q[momentum,:] = 0.
state.q[energy ,:] = pressure / (gamma - 1.)
claw = pyclaw.Controller()
claw.tfinal = 0.038
claw.solution = pyclaw.Solution(state,domain)
claw.solver = solver
claw.num_output_times = 30
claw.keep_copy = True
claw.verbosity=0
return claw
```
## Classic Clawpack algorithm with HLL versus Roe
We compare results obtained with these two solvers within a second-order Lax-Wendroff based scheme (with limiters to avoid nonphysical oscillations). This method is the basis of Clawpack.
### Sod shock tube problem
We first consider the classic shocktube problem proposed by Sod and already discussed in [Euler.ipynb](Euler.ipynb). This is a particular Riemann problem in which the initial velocity is zero on both sides of a discontinuity in pressure and/or density of a gas, and so the exact Riemann solver for the Euler equations would provide the exact solution for all time, consisting of a right-going shock, a left-going rarefaction, and an intermediate contact discontinuity.
In the numerical experiments done in this notebook, we use this initial data for a more general finite volume method that could be used to approximate the solution for any initial data. In the first time step there is a single cell interface with nontrivial Riemann data, but as the solution evolves on the grid the Riemann problems that arise in subsequent time steps are very different from the single problem we started with. Depending on the accuracy of the numerical method, the resolution of the grid, and the choice of approximate Riemann solver to use at each grid cell every time step, the numerical solution may deviate significantly from the exact solution to the original shocktube problem. This makes a good initial test problem for numerical methods because the exact solution can be computed for comparison purposes, and because it clearly shows whether the method introduces oscillations around discontinuities and/or smears them out.
```
prim_l = [1.,0.,1.]
prim_r = [1./8,0.,1./10]
q_l = euler.conservative_to_primitive(*prim_l)
q_r = euler.conservative_to_primitive(*prim_r)
# Roe-based solution
roe_st = shocktube(q_l,q_r,N=50,riemann_solver='Roe')
roe_st.run()
xc_st = roe_st.solution.state.grid.p_centers[0]
# HLL-based solution
hll_st = shocktube(q_l,q_r,N=50,riemann_solver='HLL')
hll_st.run()
# Exact solution
xc_exact_st = np.linspace(-1,1,2000)
states, speeds, reval, wave_types = euler.exact_riemann_solution(prim_l, prim_r)
def plot_frame(i):
t = roe_st.frames[i].t
fig, ax = plt.subplots(figsize=(12,6))
ax.set_xlim((-1,1)); ax.set_ylim((0,1.1))
ax.plot(xc_exact_st,reval(xc_exact_st/(t+1.e-16))[0],'-k',lw=1)
ax.plot(xc_st,hll_st.frames[i].q[density,:],'-ob',lw=2)
ax.plot(xc_st,roe_st.frames[i].q[density,:],'-or',lw=2)
plt.legend(['Exact','HLL','Roe'],loc='best')
plt.title('Density at t={:.2f}'.format(t))
plt.show()
interact(plot_frame, i=widgets.IntSlider(min=0, max=10, description='Frame'));
```
As you might expect, the HLL solver smears the middle wave (contact discontinuity) significantly more than the Roe solver does. Perhaps surprisingly, it captures the shock just as accurately as the Roe solver does.
### Woodward-Colella blast wave
Next we consider the Woodward-Colella blast wave problem, which is discussed for example in <cite data-cite="fvmhp"><a href="riemann.html#fvmhp">(LeVeque 2002)</a></cite>. Here the initial velocity is zero and the density is one everywhere. The pressure is
\begin{align}
p_0(x) = \begin{cases} 1000 & 0 \le x \le 0.1 \\
0.01 & 0.1 \le x \le 0.9 \\
100 & 0.9 \le x \le 1
\end{cases}
\end{align}
The boundaries at $x=0$ and $x=1$ are solid walls. The solution involves a Riemann problem at $x=0.1$ and another at $x=0.9$. Later, the waves resulting from these Riemann problems interact with each other.
```
roe_bw = blastwave(riemann_solver='Roe')
roe_bw.run()
hll_bw = blastwave(riemann_solver='HLL')
hll_bw.run()
fine_bw = blastwave(N=4000,riemann_solver='Roe')
fine_bw.run();
xc_bw = roe_bw.solution.state.grid.p_centers[0]
xc_fine_bw = fine_bw.solution.state.grid.p_centers[0]
def plot_frame(i):
t = roe_bw.frames[i].t
fig, ax = plt.subplots(figsize=(12,6))
ax.set_xlim((0.,1)); ax.set_ylim((0,10))
ax.plot(xc_fine_bw,fine_bw.frames[i].q[density,:],'-k',lw=1)
ax.plot(xc_bw,hll_bw.frames[i].q[density,:],'-ob',lw=2)
ax.plot(xc_bw,roe_bw.frames[i].q[density,:],'-or',lw=2)
plt.legend(['Fine','HLL','Roe'],loc='best')
plt.title('Density at t={:.3f}'.format(t))
plt.show()
interact(plot_frame, i=widgets.IntSlider(min=0, max=30, description='Frame'));
```
Here no exact solution is available, so we compare with a solution computed on a finer grid.
Again the solutions are fairly similar, though the HLL solution is a bit more smeared.
One should not conclude from these tests that, for instance, the Roe solver is *better* than the HLL solver. Many factors besides accuracy should be considered, including cost and robustness. As we have seen the HLL solver is more robust in the presence of near-vacuum states.
## High-order WENO + Runge-Kutta
Next we look at the difference between the HLL and Roe solution when these solvers are employed within a higher-order method of lines discretization using fifth-order WENO and a 4th-order Runge-Kutta scheme.
```
prim_l = [1.,0.,1.]
prim_r = [1./8,0.,1./10]
q_l = euler.conservative_to_primitive(*prim_l)
q_r = euler.conservative_to_primitive(*prim_r)
roe_weno = shocktube(q_l,q_r,N=50,riemann_solver='Roe',solver_type='sharpclaw')
roe_weno.run()
hll_weno = shocktube(q_l,q_r,N=50,riemann_solver='HLL',solver_type='sharpclaw')
hll_weno.run()
xc = roe_weno.solution.state.grid.p_centers[0]
# Exact solution
xc_exact = np.linspace(-1,1,2000)
states, speeds, reval, wave_types = euler.exact_riemann_solution(prim_l, prim_r)
def plot_frame(i):
t = roe_weno.frames[i].t
fig, ax = plt.subplots(figsize=(12,6))
ax.set_xlim((-1,1)); ax.set_ylim((0,1.1))
ax.plot(xc_exact,reval(xc_exact/(t+1.e-16))[0],'-k',lw=1)
ax.plot(xc,hll_weno.frames[i].q[density,:],'-ob',lw=2)
ax.plot(xc,roe_weno.frames[i].q[density,:],'-or',lw=2)
plt.legend(['Exact','HLL','Roe'],loc='best')
plt.title('Density at t={:.2f}'.format(t))
plt.show()
interact(plot_frame, i=widgets.IntSlider(min=0, max=10, description='Frame'));
```
With higher-order discretizations, the difference in solutions due to using different Riemann solvers is less significant. This is partly because these high-order schemes use more accurate values as inputs to the Riemann problem, so that in smooth regions the jump between most cells is very small.
| github_jupyter |
```
import numpy as np
import cv2
# read image
img = cv2.imread('sample.jpg', 0)# IMREAD_GRAYSCALE, IMREAD_COLOR
%matplotlib inline
from matplotlib import pyplot as plt
plt.imshow(img, cmap = 'gray', interpolation = 'bicubic')
plt.xticks([]), plt.yticks([]) # to hide tick values on X and Y axis
plt.show()
```
# 1. “亮点”缺陷检测
Detect part of defects by pre-process
```
equ = cv2.equalizeHist(img)# Histograms Equalization
ret2,img_th2 = cv2.threshold(img,150,255,cv2.THRESH_BINARY)
_, contours, _ = cv2.findContours(img_th2.copy(),cv2.RETR_TREE,cv2.CHAIN_APPROX_SIMPLE)
im = cv2.imread('sample.jpg', cv2.IMREAD_COLOR)# IMREAD_GRAYSCALE, IMREAD_COLOR
cv2.drawContours(im,contours,-1,(255,0,0),5)
%matplotlib inline
from matplotlib import pyplot as plt
width = 15
height = 10
plt.figure(figsize=(width, height))
plt.subplot(2, 2, 1)
plt.title('original')
plt.imshow(img, cmap = 'gray', interpolation = 'bicubic')
plt.xticks([]), plt.yticks([]) # to hide tick values on X and Y axis
plt.subplot(2, 2, 2)
plt.title('hist equlize')
plt.imshow(equ, cmap = 'gray', interpolation = 'bicubic')
plt.xticks([]), plt.yticks([]) # to hide tick values on X and Y axis
plt.subplot(2, 2, 3)
plt.title('img th2')
plt.imshow(img_th2, cmap = 'gray', interpolation = 'bicubic')
plt.xticks([]), plt.yticks([]) # to hide tick values on X and Y axis
plt.subplot(2, 2, 4)
plt.title('defect-1')
plt.imshow(im, cmap = 'gray', interpolation = 'bicubic')
plt.xticks([]), plt.yticks([]) # to hide tick values on X and Y axis
plt.show()
```
# 2. “阴影点”缺陷检测
Detect others by removing part of shadowimage
```
# read image
img2 = cv2.imread('sample_2.jpg', 0)# IMREAD_GRAYSCALE, IMREAD_COLOR
# average
blur2 = cv2.blur(img2,(5,5))
%matplotlib inline
from matplotlib import pyplot as plt
width = 15
height = 10
plt.figure(figsize=(width, height))
plt.subplot(1, 2, 1)
plt.title('original')
plt.imshow(img2, cmap = 'gray', interpolation = 'bicubic')
plt.xticks([]), plt.yticks([]) # to hide tick values on X and Y axis
plt.subplot(1, 2, 2)
plt.title('average')
plt.imshow(blur2, cmap = 'gray', interpolation = 'bicubic')
plt.xticks([]), plt.yticks([]) # to hide tick values on X and Y axis
plt.show()
equ2 = cv2.equalizeHist(blur2)# Histograms Equalization
ret2,img2_th2 = cv2.threshold(equ2,100,255,cv2.THRESH_BINARY)
plt.imshow(img2_th2, cmap = 'gray', interpolation = 'bicubic')
plt.xticks([]), plt.yticks([]) # to hide tick values on X and Y axis
plt.show()
# revert color
img2_th2_rvt = cv2.bitwise_not(img2_th2)
# erode
kernel = cv2.getStructuringElement(cv2.MORPH_RECT,(6, 6))
dilate2 = cv2.dilate(img2_th2_rvt, kernel)
eroded2 = cv2.erode(dilate2,kernel)
%matplotlib inline
from matplotlib import pyplot as plt
width = 15
height = 10
plt.figure(figsize=(width, height))
plt.subplot(1, 2, 1)
plt.title('rvt')
plt.imshow(img2_th2_rvt, cmap = 'gray', interpolation = 'bicubic')
plt.xticks([]), plt.yticks([]) # to hide tick values on X and Y axis
plt.subplot(1, 2, 2)
plt.title('erode')
plt.imshow(eroded2, cmap = 'gray', interpolation = 'bicubic')
plt.xticks([]), plt.yticks([]) # to hide tick values on X and Y axis
plt.show()
_, contours, _ = cv2.findContours(eroded2.copy(),cv2.RETR_TREE,cv2.CHAIN_APPROX_SIMPLE)
selected_contours = []
for c in contours:
area = cv2.contourArea(c)
perimeter = cv2.arcLength(c, True)# 周长
if(area>620 and perimeter<1000 and area/perimeter>5):
selected_contours.append(c)
print('area={0}, perimeter={1}'.format(area, perimeter))
len(selected_contours)
im2 = cv2.imread('sample_2.jpg', cv2.IMREAD_COLOR)# IMREAD_GRAYSCALE, IMREAD_COLOR
cv2.drawContours(im2,selected_contours,-1,(255,0,0),5)
width = 15
height = 10
plt.figure(figsize=(width, height))
plt.imshow(im2, cmap = 'gray', interpolation = 'bicubic')
plt.xticks([]), plt.yticks([]) # to hide tick values on X and Y axis
plt.show()
%matplotlib inline
from matplotlib import pyplot as plt
width = 15
height = 10
plt.figure(figsize=(width, height))
plt.subplot(2, 4, 1)
plt.title('original')
plt.imshow(img2, cmap = 'gray', interpolation = 'bicubic')
plt.xticks([]), plt.yticks([]) # to hide tick values on X and Y axis
plt.subplot(2, 4, 2)
plt.title('average')
plt.imshow(blur2, cmap = 'gray', interpolation = 'bicubic')
plt.xticks([]), plt.yticks([]) # to hide tick values on X and Y axis
plt.subplot(2,4,3)
plt.title('equ2')
plt.imshow(equ2, cmap = 'gray', interpolation = 'bicubic')
plt.xticks([]), plt.yticks([]) # to hide tick values on X and Y axis
plt.subplot(2,4,4)
plt.title('th2')
plt.imshow(img2_th2, cmap = 'gray', interpolation = 'bicubic')
plt.xticks([]), plt.yticks([]) # to hide tick values on X and Y axis
plt.subplot(2,4,5)
plt.title('revert')
plt.imshow(img2_th2_rvt, cmap = 'gray', interpolation = 'bicubic')
plt.xticks([]), plt.yticks([]) # to hide tick values on X and Y axis
plt.subplot(2,4,6)
plt.title('dilate')
plt.imshow(dilate2, cmap = 'gray', interpolation = 'bicubic')
plt.xticks([]), plt.yticks([]) # to hide tick values on X and Y axis
plt.subplot(2,4,7)
plt.title('eroded')
plt.imshow(eroded2, cmap = 'gray', interpolation = 'bicubic')
plt.xticks([]), plt.yticks([]) # to hide tick values on X and Y axis
plt.subplot(2,4,8)
plt.title('result')
plt.imshow(im2, cmap = 'gray', interpolation = 'bicubic')
plt.xticks([]), plt.yticks([]) # to hide tick values on X and Y axis
plt.show()
```
| github_jupyter |
# In-Class Coding Lab: Data Visualization
The goals of this lab are to help you understand:
- The value of visualization: A picture is worth 1,000 words!
- The various ways to visualize information
- The basic requirements for any visualization
- How to plot complex visualizations such as multi-series charts and maps
- Visualization Tools:
- Matplolib
- Plot.ly
- Folium Maps
```
!pip install cufflinks
!pip install folium
%matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
import plotly
import plotly.plotly as py
import plotly.graph_objs as go
import cufflinks as cf
import pandas as pd
import folium
import warnings
#matplotlib.rcParams['figure.figsize'] = (20.0, 10.0) # larger figure size
warnings.filterwarnings('ignore')
```
## Back to the movie goers data set
For this lab, we will once again use the movie goers dataset. As you may recall this data set is a survey demographic survey of people who go to the movies. Let's reload the data and setup our `age_group` feature again.
```
goers = pd.read_csv('CCL-moviegoers.csv')
goers['age_group'] = ''
goers['age_group'][goers['age'] <=18] = 'Youth'
goers['age_group'][(goers['age'] >=19) & (goers['age'] <=55)] = 'Adult'
goers['age_group'][goers['age'] >=56] = 'Senior'
goers.sample(5)
```
## Visualizing Data
There are many ways your can visualize information. Which one is the most appropriate? It depends on the data, of course.
- **Counting Categorial data** belongs in *charts like pie charts and bar charts*.
- **Counting Numerical data** is best suited for *histograms*.
- **Timeseries data and continuous data** belongs in *line charts*.
- **A comparision of two continuous values** is best suited for a *scatter plot*.
- **Geographical data** is best displauyed on *maps*.
Let's use this knowledge to plot some data in the `goers` `DataFrame`!
## Males or Females?
The first think we might want to visualize is a count gender in the dataset. A **pie** chart is well suited for this task as it displays data as a portion of a whole. To create a pie chart we need the data to count and the labels for the counts.
Let's try it.
First we get the value counts as a series `gender`:
```
gender = goers['gender'].value_counts()
gender
```
Then we make it into a dataframe:
```
gender_df = pd.DataFrame( { 'Gender' : gender.index, "Counts" : gender })
gender_df
```
Then we plot! The index has the labels, and the value at the index is what we want to plot:
```
gender_df.plot.pie( y = 'Counts') # y are the values we are plotting
```
### Now You Try it!
Create a pie chart based on `age_group` first create a series of the `value_counts()` second, create the `DataFrame` with two columns `AgeGroup` and `Counts` then plot with `.plot.pie()`.
Follow the steps we did in the previous three cells, but comvine into one cell!
```
#todo write code here
age_group = goers['age'].value_counts()
age_group_df = pd.DataFrame( { 'AgeGroup' : age_group.index, 'counts': age_group})
age_group_df
age_group_df.plot.pie( y = 'counts')
```
## Too many pieces of the pie?
Pie charts are nice, but they are only useful when you have a small number of labels. More that 5-7 labels and the pie becomes messy. For example take a look at this pie chart of `occupation`:
```
occ = goers['occupation'].value_counts()
occ_df = pd.DataFrame( { 'occupation' : occ.index, "counts" : occ })
occ_df.plot.pie(y = 'counts')
```
That's crazy... and difficult to comprehend. Also pie charts visualize data as part of the whole. We have no idea how many students there are. Sometimes we want to know actual counts. This is where the **bar chart** comes in handy!
## Raising the bar!
Let's reproduce the same plot as a bar:
```
occ_df.plot.bar()
```
Ahh. that's much better. So much easier to understand!
### Now you try it!
Write a one-liner to plot `groups_df` as a Bar!
```
# todo write code here
groups_df.plot.bar()
#doesn't tell us what groups_df is
```
## When bar charts fail...
Bar charts have the same problem as pie charts. Too many categories overcomplicate the chart, or show the data in a meaningless way. For example, let's create a bart chart for ages:
```
ages = goers['age'].value_counts()
ages_df = pd.DataFrame( { 'age' : ages.index, "counts" : ages })
ages_df.plot.bar(y = 'counts')
```
Meaningless. For two key reasons:
1. too many categories
2. age is a continuous variable not a categorical variable. In plain English, this means there's a relationship between one age and the next. 20 < 21 < 22. This is not represented in a bar chart.
## ...Call in the Histogram!
What we want is a **historgram**, which takes a continuous variable and loads counts into "buckets". Notice how we didn't have to lump data with `value_counts()`. Histograms can do that automatically because the `age` variable is continuous. Let's try it:
```
goers.hist(column ='age')
```
## Plot.ly
[Plot.ly](https://plot.ly) is data visualization as a service. You give it data, it gives you back a web-based plot. Plot.ly is free and works with a variety of environments and programming languages, including Python.
For Python is has bindings so that you can use it just like `matplotlib`! No need to manually invoke the web service call.
To get started with plot.ly you must sign up for an account and get a set of credentials:
- Visit [https://plot.ly/settings/api](https://plot.ly/settings/api)
- Create an account or sign-in with Google or GitHub
- Generate your API key and paste your username and key in the code below:
```
# todo: setup the credentials replace ??? and ??? with your Plot.ly username and api_key
plotly.tools.set_credentials_file(username='???', api_key='???')
```
Using plot.ly is as easy as, or sometimes easier than `matplotlib`. In most cases all you need to do is call `iplot()` on the data frame. For example, here's out first pie chart, plotly style:
```
gender_df.iplot(kind="pie", labels = 'Gender', values='Counts')
```
Notice that plot.ly is a bit more interactive. You can hover over the part of the pie chart and see counts!
### Now You Try it!
Use plotly's `iplot()` method to create a bar chart on the `occ_df` Data Frame:
```
# todo: write code here
occ_df.iplot(kind='bar', labels= 'Occupation', values = 'Counts')
```
## Folium with Leaflet.js
Folium is a Python module wrapper for [Leaflet.js](http://leafletjs.com/), which uses [Open Street Maps](https://www.openstreetmap.us/). These are two, popular open source mapping libraries. Unlike Google maps API, its 100% free!
You can use Folium to render maps in Python and put data on the maps. Here's how easy it is to bring up a map:
```
CENTER_US = (39.8333333,-98.585522)
london = (51.5074, -0.1278)
map = folium.Map(location=CENTER_US, zoom_start=4)
map
```
You can zoom right down to the street level and get a amazing detail. There's also different maps you can use, as was covered in this week's reading.
## Mapping the students.
Let's take the largest category of movie goers and map their whereabouts. We will first need to import a data set to give us a lat/lng for the `zip_code` we have in the dataframe. We could look this up with Google's geolookup API, but that's too slow as we will be making 100's of requests. It's better to have them stored already and merge them with `goers`!
Let's import the zipcode database into a Pandas DataFrame, then merge it with the `goers` DataFrame:
```
zipcodes = pd.read_csv('https://raw.githubusercontent.com/mafudge/datasets/master/zipcodes/free-zipcode-database-Primary.csv', dtype = {'Zipcode' :object})
data = goers.merge(zipcodes, how ='inner', left_on='zip_code', right_on='Zipcode')
students = data[ data['occupation'] == 'student']
students.sample()
```
Let's explain the code, as a Pandas refresher course:
1. in the first line I added `dtype = {'Zipcode' :object}` to force the `Zipcode` column to be of type `object` without that, it imports as type `int` and cannot match with the `goers` DataFrame.
1. the next line merges the two dataframes together where the `zip_code` in `doers` (on_left) matches `Zipcode` in `zipcodes` (on_right)
1. the result `data` is a combined DataFrame, which we then filter to only `student` occupations, sorting that in the `students` DataFrame
## Slapping those students on a map!
We're ready to place the students on a map. It's easy:
1. For each row in the students dataframe:
1. get the coordinates (lat /lng )
1. make a `marker` with the coordinates
1. add the marker to the map with `add_children()`
Here we go!
```
for row in students.to_records():
pos = (row['Lat'],row['Long'])
message = "%d year old %s from %s, %s" % (row['age'],row['gender'], row['City'], row['State'])
marker = folium.Marker(location=pos,
popup=message
)
map.add_children(marker)
map
```
### Now you try it!
1. use the `data` DataFrame to retrieve only the occupation `programmer`
1. create another map `map2` plot the programmers on that map!
```
## todo write code here!
zipcodes = pd.read_csv('https://raw.githubusercontent.com/mafudge/datasets/master/zipcodes/free-zipcode-database-Primary.csv', dtype = {'Zipcode' :object})
data = goers.merge(zipcodes, how ='inner', left_on='zip_code', right_on='Zipcode')
programmers = data[ data['occupation'] == 'programmers']
programmers.sample()
for row in programmers.to_records():
pos = (row['Lat'],row['Long'])
message = "%d year old %s from %s, %s" % (row['age'],row['gender'], row['City'], row['State'])
marker = folium.Marker(location=pos,
popup=message
)
map.add_children(marker)
map
```
| github_jupyter |
<a href="https://colab.research.google.com/github/harvardnlp/pytorch-struct/blob/master/notebooks/Unsupervised_CFG.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
!pip install -qqq torchtext -qqq pytorch-transformers dgl
!pip install -qqqU git+https://github.com/harvardnlp/pytorch-struct
import torchtext
import torch
from torch_struct import SentCFG
from examples.networks import NeuralCFG
import examples.data
# Download and the load default data.
WORD = torchtext.data.Field(include_lengths=True)
UD_TAG = torchtext.data.Field(init_token="<bos>", eos_token="<eos>", include_lengths=True)
# Download and the load default data.
train, val, test = torchtext.datasets.UDPOS.splits(
fields=(('word', WORD), ('udtag', UD_TAG), (None, None)),
filter_pred=lambda ex: 5 < len(ex.word) < 30
)
WORD.build_vocab(train.word, min_freq=3)
UD_TAG.build_vocab(train.udtag)
train_iter = examples.data.TokenBucket(train,
batch_size=200,
device="cuda:0")
H = 256
T = 30
NT = 30
model = NeuralCFG(len(WORD.vocab), T, NT, H)
model.cuda()
opt = torch.optim.Adam(model.parameters(), lr=0.001, betas=[0.75, 0.999])
def train():
#model.train()
losses = []
for epoch in range(10):
for i, ex in enumerate(train_iter):
opt.zero_grad()
words, lengths = ex.word
N, batch = words.shape
words = words.long()
params = model(words.cuda().transpose(0, 1))
dist = SentCFG(params, lengths=lengths)
loss = dist.partition.mean()
(-loss).backward()
losses.append(loss.detach())
torch.nn.utils.clip_grad_norm_(model.parameters(), 3.0)
opt.step()
if i % 100 == 1:
print(-torch.tensor(losses).mean(), words.shape)
losses = []
train()
for i, ex in enumerate(train_iter):
opt.zero_grad()
words, lengths = ex.word
N, batch = words.shape
words = words.long()
params = terms(words.transpose(0, 1)), rules(batch), roots(batch)
tree = CKY(MaxSemiring).marginals(params, lengths=lengths, _autograd=True)
print(tree)
break
def split(spans):
batch, N = spans.shape[:2]
splits = []
for b in range(batch):
cover = spans[b].nonzero()
left = {i: [] for i in range(N)}
right = {i: [] for i in range(N)}
batch_split = {}
for i in range(cover.shape[0]):
i, j, A = cover[i].tolist()
left[i].append((A, j, j - i + 1))
right[j].append((A, i, j - i + 1))
for i in range(cover.shape[0]):
i, j, A = cover[i].tolist()
B = None
for B_p, k, a_span in left[i]:
for C_p, k_2, b_span in right[j]:
if k_2 == k + 1 and a_span + b_span == j - i + 1:
B, C = B_p, C_p
k_final = k
break
if j > i:
batch_split[(i, j)] =k
splits.append(batch_split)
return splits
splits = split(spans)
```
| github_jupyter |
<a href="https://qworld.net" target="_blank" align="left"><img src="../qworld/images/header.jpg" align="left"></a>
$ \newcommand{\bra}[1]{\langle #1|} $
$ \newcommand{\ket}[1]{|#1\rangle} $
$ \newcommand{\braket}[2]{\langle #1|#2\rangle} $
$ \newcommand{\dot}[2]{ #1 \cdot #2} $
$ \newcommand{\biginner}[2]{\left\langle #1,#2\right\rangle} $
$ \newcommand{\mymatrix}[2]{\left( \begin{array}{#1} #2\end{array} \right)} $
$ \newcommand{\myvector}[1]{\mymatrix{c}{#1}} $
$ \newcommand{\myrvector}[1]{\mymatrix{r}{#1}} $
$ \newcommand{\mypar}[1]{\left( #1 \right)} $
$ \newcommand{\mybigpar}[1]{ \Big( #1 \Big)} $
$ \newcommand{\sqrttwo}{\frac{1}{\sqrt{2}}} $
$ \newcommand{\dsqrttwo}{\dfrac{1}{\sqrt{2}}} $
$ \newcommand{\onehalf}{\frac{1}{2}} $
$ \newcommand{\donehalf}{\dfrac{1}{2}} $
$ \newcommand{\hadamard}{ \mymatrix{rr}{ \sqrttwo & \sqrttwo \\ \sqrttwo & -\sqrttwo }} $
$ \newcommand{\vzero}{\myvector{1\\0}} $
$ \newcommand{\vone}{\myvector{0\\1}} $
$ \newcommand{\stateplus}{\myvector{ \sqrttwo \\ \sqrttwo } } $
$ \newcommand{\stateminus}{ \myrvector{ \sqrttwo \\ -\sqrttwo } } $
$ \newcommand{\myarray}[2]{ \begin{array}{#1}#2\end{array}} $
$ \newcommand{\X}{ \mymatrix{cc}{0 & 1 \\ 1 & 0} } $
$ \newcommand{\I}{ \mymatrix{rr}{1 & 0 \\ 0 & 1} } $
$ \newcommand{\Z}{ \mymatrix{rr}{1 & 0 \\ 0 & -1} } $
$ \newcommand{\Htwo}{ \mymatrix{rrrr}{ \frac{1}{2} & \frac{1}{2} & \frac{1}{2} & \frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & \frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} & \frac{1}{2} } } $
$ \newcommand{\CNOT}{ \mymatrix{cccc}{1 & 0 & 0 & 0 \\ 0 & 1 & 0 & 0 \\ 0 & 0 & 0 & 1 \\ 0 & 0 & 1 & 0} } $
$ \newcommand{\norm}[1]{ \left\lVert #1 \right\rVert } $
$ \newcommand{\pstate}[1]{ \lceil \mspace{-1mu} #1 \mspace{-1.5mu} \rfloor } $
$ \newcommand{\greenbit}[1] {\mathbf{{\color{green}#1}}} $
$ \newcommand{\bluebit}[1] {\mathbf{{\color{blue}#1}}} $
$ \newcommand{\redbit}[1] {\mathbf{{\color{red}#1}}} $
$ \newcommand{\brownbit}[1] {\mathbf{{\color{brown}#1}}} $
$ \newcommand{\blackbit}[1] {\mathbf{{\color{black}#1}}} $
<font style="font-size:28px;" align="left"><b> Superposition </b></font>
<br>
_prepared by Abuzer Yakaryilmaz_
<br><br>
[<img src="../qworld/images/watch_lecture.jpg" align="left">](https://youtu.be/uJZtxWHAlPI)
<br><br><br>
There is no classical counterpart of the concept "superposition".
But, we can still use a classical analogy that might help us to give some intuitions.
<h3> Probability distribution </h3>
Suppose that Asja starts in $ \myvector{1\\0} $ and secretly applies the probabilistic operator $ \mymatrix{cc}{ 0.3 & 0.6 \\ 0.7 & 0.4 } $.
Because she applies her operator secretly, our information about her state is probabilistic, which is calculated as
$$
\myvector{0.3 \\ 0.7} = \mymatrix{cc}{ 0.3 & 0.6 \\ 0.7 & 0.4 } \myvector{1\\0}.
$$
Asja is either in state 0 or in state 1.
However, from our point of view, Asja is in state 0 with probability $ 0.3 $ and in state 1 with probability $ 0.7 $.
We can say that Asja is in a probability distribution of states 0 and 1, being in both states at the same time but with different weights.
On the other hand, if we observe Asja's state, then our information about Asja becomes deterministic: either $ \myvector{1 \\ 0} $ or $ \myvector{0 \\ 1} $.
We can say that, after observing Asja's state, the probabilistic state $ \myvector{0.3 \\ 0.7} $ collapses to either $ \myvector{1 \\ 0} $ or $ \myvector{0 \\ 1} $.
<h3> The third experiment </h3>
Remember the following experiment. We trace it step by step by matrix-vector multiplication.
<img src="../photon/images/photon7.jpg" width="65%">
<b> The initial Step </b>
The photon is in state $ \ket{v_0} = \vzero $.
<b> The first step </b>
Hadamard is applied:
$ \ket{v_1} = \hadamard \vzero = \stateplus $.
At this point, the photon is in a <b>superposition</b> of state $ \ket{0} $ and state $ \ket{1} $, <u>being in both states with the amplitudes</u> $ \frac{1}{\sqrt{2}} $ and $ \frac{1}{\sqrt{2}} $, respectively.
The state of photon is $ \ket{v_1} = \stateplus $, and we can also represent it as follows:
$ \ket{v_1} = \frac{1}{\sqrt{2}} \ket{0} + \frac{1}{\sqrt{2}} \ket{1} $.
<b> The second step </b>
Hadamard is applied again:
We write the effect of Hadamard on states $ \ket{0} $ and $ \ket{1} $ as follows:
$ H \ket{0} = \frac{1}{\sqrt{2}} \ket{0} + \frac{1}{\sqrt{2}} \ket{1} $ <i>(These are the transition amplitudes of the first column.)</i>
$ H \ket{1} = \frac{1}{\sqrt{2}} \ket{0} - \frac{1}{\sqrt{2}} \ket{1} $ <i>(These are the transition amplitudes of the second column.)</i>
This representation helps us to see clearly why the state $ \ket{1} $ disappears.
Now, let's see the effect of Hadamard on the quantum state $ \ket{v_1} = \frac{1}{\sqrt{2}} \ket{0} + \frac{1}{\sqrt{2}} \ket{1} $:
$ \ket{v_2} = H \ket{v_1} = H \mybigpar{ \frac{1}{\sqrt{2}} \ket{0} + \frac{1}{\sqrt{2}} \ket{1} } = \frac{1}{\sqrt{2}} H \ket{0} + \frac{1}{\sqrt{2}} H \ket{1} $
We can replace $ H\ket{0} $ and $ H\ket{1} $ as described above. $ \ket{v_2} $ is formed by the summation of the following terms:
$~~~~~~~~ \dsqrttwo H \ket{0} = $ <font color="green">$\donehalf \ket{0} $</font> <font color="red">$ + \donehalf \ket{1} $</font>
$~~~~~~~~ \dsqrttwo H \ket{1} = $ <font color="green">$\donehalf \ket{0} $</font> <font color="red">$ - \donehalf \ket{1} $</font>
<br>
<font size="+1">$ \mathbf{+}\mbox{___________________} $</font>
$ ~~ $ <font color="green"> $\mypar{ \donehalf+\donehalf } \ket{0} $</font> +
<font color="red"> $\mypar{ \donehalf-\donehalf } \ket{1} $ </font> $ = \mathbf{\ket{0}} $.
<font color="green">The amplitude of $ \ket{0} $ becomes 1,</font> <font color="red"> but the amplitude of $ \ket{1} $ becomes 0 because of cancellation.</font>
The photon was in both states at the same time with <u>certain amplitudes</u>.
After the second Hadamard, the "outcomes" are <u>interfered with each other</u>.
The interference can be constructive or destructive.
In our examples, <font color="green"><b>the outcome $ \ket{0} $s are interfered constructively</b></font>, but <font color="red"><b>the outcome $ \ket{1} $s are interfered destructively</b></font>.
<h3> Observations </h3>
<u>Probabilistic systems</u>: If there is a nonzero transition to a state, then it contributes to the probability of this state positively.
<u>Quantum systems</u>: If there is a nonzero transition to a state, then we cannot know its contribution without knowing the other transitions to this state.
If it is the only transition, then it contributes to the amplitude (and probability) of the state, and it does not matter whether the sign of the transition is positive or negative.
If there is more than one transition, then depending on the summation of all transitions, we can determine whether a specific transition contributes or not.
As a simple rule, if the final amplitude of the state and nonzero transition have the same sign, then it is a positive contribution; and, if they have the opposite signs, then it is a negative contribution.
<h3> Task 1 </h3>
[on paper]
Start in state $ \ket{u_0} = \ket{1} $.
Apply Hadamard operator to $ \ket{u_0} $, i.e, find $ \ket{u_1} = H \ket{u_0} $.
Apply Hadamard operator to $\ket{u_1}$, i.e, find $ \ket{u_2} = H \ket{u_1} $.
Observe the constructive and destructive interferences, when calculating $ \ket{u_2} $.
<h3> Being in a superposition </h3>
A quantum system can be in more than one state with nonzero amplitudes.
Then, we say that our system is in a superposition of these states.
When evolving from a superposition, the resulting transitions may affect each other constructively and destructively.
This happens because of having opposite sign transition amplitudes.
Otherwise, all nonzero transitions are added up to each other as in probabilistic systems.
<hr>
<h2> Measurement </h2>
We can measure a quantum system, and then the system is observed in one of its states. This is the most basic type of measurement in quantum computing. (There are more generic measurement operators, but we will not mention about them.)
The probability of the system to be observed in a specified state is the square value of its amplitude.
<ul>
<li> If the amplitude of a state is zero, then this state cannot be observed. </li>
<li> If the amplitude of a state is nonzero, then this state can be observed. </li>
</ul>
For example, if the system is in quantum state
$$
\myrvector{ -\frac{\sqrt{2}}{\sqrt{3}} \\ \frac{1}{\sqrt{3}} },
$$
then, after a measurement, we can observe the system in state $\ket{0} $ with probability $ \frac{2}{3} $ and in state $\ket{1}$ with probability $ \frac{1}{3} $.
<h4> Collapsing </h4>
After the measurement, the system collapses to the observed state, and so the system is no longer in a superposition. Thus, the information kept in a superposition is lost.
- In the above example, when the system is observed in state $\ket{0}$, then the new state becomes $ \myvector{1 \\ 0} $.
- If it is observed in state $\ket{1}$, then the new state becomes $ \myvector{0 \\ 1} $.
<h3> The second experiment of the quantum coin flipping </h3>
Remember the experiment set-up.
<img src="../photon/images/photon5.jpg" width="65%">
In this experiment, after the first quantum coin-flipping, we make a measurement.
If the measurement outcome is state $ \ket{0} $, then we apply a second Hadamard.
First, we trace the experiment analytically.
<table width="100%"><tr>
<td width="400px" style="background-color:white;text-align:center;vertical-align:middle;" cellpadding=0>
<img src="images/tracing-2nd-exp.png">
<br><br>
<font size="-2"><a href="../images/tracing-2nd-exp.txt" target="_blank">the tex code of the image</a></font>
</td>
<td width="*" style="background-color:white;text-align:left;vertical-align:top;">
<b> The first Hadamard </b>
<br><br>
We start in state $ \ket{0} = \vzero $. Then, we apply Hadamard operator:
<br><br>
$ \stateplus = \hadamard \vzero $ <hr>
<!---------->
<b> The first measurement </b>
<br><br>
Due to the photon detector A, the photon cannot be in superposition, and so it forces the photon to be observed in state $\ket{0}$ or state $ \ket{1} $. This is a measurement.
<br><br>
Since the amplitudes are $ \sqrttwo $, we observe each state with equal probability.
<br><br>
Thus, with probability $ \frac{1}{2} $, the new quantum state is $ \ket{0} = \vzero $.
<br><br>
And, with probability $ \frac{1}{2} $, the new quantum state is $ \ket{1} = \vone $. <hr>
<!---------->
<b> The second Hadamard </b>
<br><br>
If the photon is in state $ \ket{0} $, then another Hadamard operator is applied.
<br><br>
In other words, with probability $ \frac{1}{2} $, the computation continues and another Hadamard is applied:
<br><br>
$ \stateplus = \hadamard \vzero $ <hr>
<b> The second measurement </b>
<br><br>
Due to photon detectors B1 and B2, we make another measurement.
<br><br>
Thus, we observe state $ \ket{0} $ with probability $ \frac{1}{4} $ and state $ \ket{1} $ with probability $ \frac{1}{4} $.
<br><br>
At the end, the state $ \ket{0} $ can be observed with probability $ \frac{1}{4} $, and the state $ \ket{1} $ can be observed with probability $ \frac{3}{4} $.
</td>
</tr></table>
<h3> Implementing the second experiment </h3>
By using the simulator, we can implement the second experiment.
For this purpose, qiskit provides a conditional operator based on the value of a classical register.
In the following example, the last operator (x-gate) on the quantum register will be executed if the value of the classical register is 1.
q = QuantumRegister(1)
c = ClassicalRegister(1)
qc = QuantumCircuit(q,c)
...
qc.measure(q,c)
qc.x(q[0]).c_if(c,1)
In our experiment, we use such classical control after the first measurement.
```
# import all necessary objects and methods for quantum circuits
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
# define a quantum register with a single qubit
q = QuantumRegister(1,"q")
# define a classical register with a single bit
c = ClassicalRegister(1,"c")
# define a quantum circuit
qc = QuantumCircuit(q,c)
# apply the first Hadamard
qc.h(q[0])
# the first measurement
qc.measure(q,c)
# apply the second Hadamard if the measurement outcome is 0
qc.h(q[0]).c_if(c,0)
# the second measurement
qc.measure(q[0],c)
# draw the circuit
display(qc.draw(output="mpl"))
```
<h3> Task 2 </h3>
If we execute this circuit 1000 times, what are the expected numbers of observing the outcomes '0' and '1'?
Test your result by executing the following code.
```
job = execute(qc,Aer.get_backend('qasm_simulator'),shots=1000)
counts = job.result().get_counts(qc)
print(counts)
```
<h3> Task 3 </h3>
Repeat the second experiment with the following modifications.
Start in state $ \ket{1} $.
Apply a Hadamard gate.
Make a measurement.
If the measurement outcome is 0, stop.
Otherwise, apply a second Hadamard, and then make a measurement.
Execute your circuit 1000 times.
Calculate the expected values of observing '0' and '1', and then compare your result with the simulator result.
```
# import all necessary objects and methods for quantum circuits
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
#
# your code is here
#
```
<a href="Q36_Superposition_and_Measurement_Solutions.ipynb#task3">click for our solution</a>
<h3> Task 4 </h3>
Design the following quantum circuit.
Start in state $ \ket{0} $.
Repeat 3 times:
if the classical bit is 0:
apply a Hadamard operator
make a measurement
Execute your circuit 1000 times.
Calculate the expected values of observing '0' and '1', and then compare your result with the simulator result.
```
# import all necessary objects and methods for quantum circuits
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
#
# your code is here
#
```
<a href="Q36_Superposition_and_Measurement_Solutions.ipynb#task4">click for our solution</a>
---
<h3> Extra: Task 5 </h3>
Design the following randomly created quantum circuit.
Start in state $ \ket{0} $.
apply a Hadamard operator
make a measurement
REPEAT 4 times:
randomly pick x in {0,1}
if the classical bit is x:
apply a Hadamard operator
make a measurement
Draw your circuit, and guess the expected frequency of observing '0' and '1' if the circuit is executed 10000 times.
Then, execute your circuit 10000 times, and compare your result with the simulator result.
Repeat execution a few more times.
```
# import all necessary objects and methods for quantum circuits
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
# import randrange for random choices
from random import randrange
#
# your code is here
#
```
<a href="Q36_Superposition_and_Measurement_Solutions.ipynb#task5">click for our solution</a>
| github_jupyter |
This notebook will be sketching out my implementation of the Resnet-18 network used in this [assignment](http://cs231n.stanford.edu/reports/2017/pdfs/406.pdf) mentioned in initial_work.ipynb. I want to try building it out with the base components built into PyTorch and compare that to PyTorch's built-in Resnet constructors.
```
import torch
import torch.nn as nn
import torch.nn.functional as F
import common_utils
class ResBlock(nn.Module):
"""
Class representing one two-layer residual learning block, as outlined in the paper
"Deep Residual Learning for Image Recognition"
"""
def __init__(self, in_dimension, dimension, stride=1):
super(ResBlock, self).__init__()
self.dimension = dimension
self.in_dimension = in_dimension
self.in_conv = nn.Conv2d(in_dimension, dimension, 3, stride=stride, padding=1)
self.bnorm = nn.BatchNorm2d(dimension)
self.conv = nn.Conv2d(dimension, dimension, 3, stride=1, padding=1)
def forward(self, x):
"""
Connectivity follows the diagram from the paper 'Deep Residual Learning for Image Recognition'
"""
res = x
out = F.relu(self.bnorm(self.in_conv(x)), inplace=True)
out = self.bnorm(self.conv(out))
# adding the residual weights to the layer output
# need to downsample the residual in first layer of blocks with nonzero stride
if (res.size() != out.size()):
conv = nn.Conv2d(self.in_dimension, self.dimension, 1, stride=2)
bnorm = nn.BatchNorm2d(self.dimension)
res = bnorm(conv(res))
out += res
out = F.relu(out)
return out
class HomegrownResnet18(nn.Module):
def __init__(self, num_classes):
super(HomegrownResnet18, self).__init__()
self.in_conv = nn.Conv2d(3, 64, 7, stride=2, padding=3)
self.bnorm = nn.BatchNorm2d(64)
self.max_pool = nn.MaxPool2d(3,stride=2, padding = 1)
self.chunk64 = nn.Sequential(*[ResBlock(64,64),ResBlock(64,64)])
self.chunk128 = nn.Sequential(*[ResBlock(64,128,stride=2),ResBlock(128,128)])
self.chunk256 = nn.Sequential(*[ResBlock(128,256,stride=2),ResBlock(256,256)])
self.chunk512 = nn.Sequential(*[ResBlock(256,512,stride=2),ResBlock(512,512)])
self.avgpool = nn.AvgPool2d(7,stride=2)
self.fc = nn.Linear(512,num_classes)
# initialize weights in a way that makes sense for ReLU layers
for m in self.modules():
if isinstance(m, nn.Conv2d):
nn.init.kaiming_normal_(m.weight,nonlinearity='relu')
elif isinstance(m, nn.BatchNorm2d):
nn.init.constant_(m.weight, 1)
nn.init.constant_(m.bias, 0)
def forward(self, x):
x = F.relu(self.bnorm(self.in_conv(x)),inplace=True)
x = self.max_pool(x)
x = self.chunk64(x)
x = self.chunk128(x)
x = self.chunk256(x)
x = self.chunk512(x)
x = self.avgpool(x)
x = x.view(x.size()[0], -1)
x = self.fc(x)
return x
net = HomegrownResnet18(38)
print(net)
train, test, val = common_utils.get_dataloaders()
import torch.optim as optim
import matplotlib.pyplot as plt
criterion = nn.CrossEntropyLoss()
optimizer = optim.Adam(net.parameters(), lr = 0.001)
val_acc = []
iterations = []
train_len = len(train)
for epoch in range(30):
running_loss = 0
for i, sample in enumerate(train):
images, labels = sample['images'], sample['labels']
optimizer.zero_grad()
outputs = net(images)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
running_loss += loss.item()
if i % 80 == 79:
print('[%d, %5d] loss: %.3f' %
(epoch + 1, i + 1, running_loss / 80))
running_loss = 0.0
with torch.no_grad():
total = 0
correct = 0
for sample in val:
images,labels = sample['images'], sample['labels']
outputs = net(images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
val_acc.append(100*correct/total)
iterations.append((i+1) + (train_len * epoch))
print("[%d, %5d] test accuracy: %.3f" % (epoch + 1, i + 1, 100*correct/total))
print('done')
plt.plot(iterations, val_acc)
plt.xlabel('number of iterations')
plt.ylabel('percent accuracy')
plt.show()
```
| github_jupyter |
# Object-based filtering of pixel classifications <img align="right" src="../figs/DE_Africa_Logo_Stacked_RGB_small.jpg">
## Background
Geographic Object-Based Image Analysis (GEOBIA), which aims to group pixels together into meaningful image-objects. There are two advantages to a GEOBIA worklow; one, we can reduce the 'salt and pepper' effect typical of classifying pixels; and two, we can increase the computational efficiency of our workflow by grouping pixels into fewer, larger, but meaningful objects. A review of the emerging trends in GEOBIA can be found in [Chen et al. (2017)](https://www.tandfonline.com/doi/abs/10.1080/15481603.2018.1426092).
## Description
In this notebook, we take the pixel-based classifications generated in the `4_Predict.ipynb` notebook, and filter the classifications by image-objects. To do this, we first need to conduct image segmentation using the function `rsgislib.segmentation.runShepherdSegmentation`. This image sgementation algorithm is fast and scalable. The image segmentation is conducted on the `NDVI_S1` and `NDVI_S2` layers (NDVI season 1, NDVI season 2).
To filter the pixel observations, we assign to each segment the majority (mode) pixel classification.
1. Load the NDVI_S1 and NDVI_S2 layers
2. Convert the NDVI layers to a .kea file format (a requirement for RSGSISLIB)
3. Run the image segmentation
4. Calculate the mode statistic for each segment
5. Write the new object-based classification to disk as COG
***
## Getting started
To run this analysis, run all the cells in the notebook, starting with the "Load packages" cell.
### Load Packages
```
import os
import sys
import gdal
import shutil
import xarray as xr
import numpy as np
import geopandas as gpd
import subprocess as sp
from datacube.utils.cog import write_cog
from rsgislib.segmentation import segutils
from scipy.ndimage.measurements import _stats
sys.path.append('../../Scripts')
from deafrica_classificationtools import HiddenPrints
```
# Analysis Parameters
```
test_shapefile = 'data/imagesegtiles.shp'
results = 'results/classifications/predicted/20210401/'
model_type='gm_mads_two_seasons_20210401'
min_seg_size=50 #in number of pixels
```
### Open testing tile shapefile
```
gdf = gpd.read_file(test_shapefile)
```
## Image segmentation
```
%%time
for g_id in gdf['title'].values:
print('working on grid: ' + g_id)
#store temp files somewhere
directory=results+'tmp_'+g_id
if not os.path.exists(directory):
os.mkdir(directory)
tmp='tmp_'+g_id+'/'
#inputs to image seg
tiff_to_segment = results+'ndvi/Eastern_tile_'+g_id+'_NDVI.tif'
kea_file = results+'ndvi/Eastern_tile_'+g_id+'_NDVI.kea'
segmented_kea_file = results+'ndvi/Eastern_tile_'+g_id+'_segmented.kea'
#convert tiff to kea
gdal.Translate(destName=kea_file,
srcDS=tiff_to_segment,
format='KEA',
outputSRS='EPSG:6933')
#run image seg
print(' image segmentation...')
with HiddenPrints():
segutils.runShepherdSegmentation(inputImg=kea_file,
outputClumps=segmented_kea_file,
tmpath=results+tmp,
numClusters=60,
minPxls=min_seg_size)
#open segments, and predictions
segments=xr.open_rasterio(segmented_kea_file).squeeze().values
t = results+ 'tiles/Eastern_tile_'+g_id+'_prediction_pixel_'+model_type+'.tif'
pred = xr.open_rasterio(t).squeeze().drop_vars('band')
#calculate mode
print(' calculating mode...')
count, _sum =_stats(pred, labels=segments, index=segments)
mode = _sum > (count/2)
mode = xr.DataArray(mode, coords=pred.coords, dims=pred.dims, attrs=pred.attrs).astype(np.int16)
#write to disk
print(' writing to disk...')
write_cog(mode, results+ 'segmented/Eastern_tile_'+g_id+'_prediction_filtered_'+model_type+'.tif', overwrite=True)
#remove the tmp folder
shutil.rmtree(results+tmp)
os.remove(kea_file)
os.remove(segmented_kea_file)
os.remove(tiff_to_segment)
mode.plot(size=12);
pred.plot(size=12)
```
## Next steps
To continue working through the notebooks in this `Eastern Africa Cropland Mask` workflow, go to the next notebook `6_Accuracy_assessment.ipynb`.
1. [Extracting_training_data](1_Extracting_training_data.ipynb)
2. [Inspect_training_data](2_Inspect_training_data.ipynb)
3. [Train_fit_evaluate_classifier](3_Train_fit_evaluate_classifier.ipynb)
4. [Predict](4_Predict.ipynb)
5. **Object-based_filtering (this notebook)**
6. [Accuracy_assessment](6_Accuracy_assessment.ipynb)
***
## Additional information
**License:** The code in this notebook is licensed under the [Apache License, Version 2.0](https://www.apache.org/licenses/LICENSE-2.0).
Digital Earth Africa data is licensed under the [Creative Commons by Attribution 4.0](https://creativecommons.org/licenses/by/4.0/) license.
**Contact:** If you need assistance, please post a question on the [Open Data Cube Slack channel](http://slack.opendatacube.org/) or on the [GIS Stack Exchange](https://gis.stackexchange.com/questions/ask?tags=open-data-cube) using the `open-data-cube` tag (you can view previously asked questions [here](https://gis.stackexchange.com/questions/tagged/open-data-cube)).
If you would like to report an issue with this notebook, you can file one on [Github](https://github.com/digitalearthafrica/deafrica-sandbox-notebooks).
**Last modified:** Dec 2020
| github_jupyter |
<a href="https://colab.research.google.com/github/AI4Finance-LLC/FinRL/blob/master/FinRL_ensemble_stock_trading_ICAIF_2020.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Deep Reinforcement Learning for Stock Trading from Scratch: Multiple Stock Trading Using Ensemble Strategy
Tutorials to use OpenAI DRL to trade multiple stocks using ensemble strategy in one Jupyter Notebook | Presented at ICAIF 2020
* This notebook is the reimplementation of our paper: Deep Reinforcement Learning for Automated Stock Trading: An Ensemble Strategy, using FinRL.
* Check out medium blog for detailed explanations: https://medium.com/@ai4finance/deep-reinforcement-learning-for-automated-stock-trading-f1dad0126a02
* Please report any issues to our Github: https://github.com/AI4Finance-LLC/FinRL-Library/issues
* **Pytorch Version**
# Content
* [1. Problem Definition](#0)
* [2. Getting Started - Load Python packages](#1)
* [2.1. Install Packages](#1.1)
* [2.2. Check Additional Packages](#1.2)
* [2.3. Import Packages](#1.3)
* [2.4. Create Folders](#1.4)
* [3. Download Data](#2)
* [4. Preprocess Data](#3)
* [4.1. Technical Indicators](#3.1)
* [4.2. Perform Feature Engineering](#3.2)
* [5.Build Environment](#4)
* [5.1. Training & Trade Data Split](#4.1)
* [5.2. User-defined Environment](#4.2)
* [5.3. Initialize Environment](#4.3)
* [6.Implement DRL Algorithms](#5)
* [7.Backtesting Performance](#6)
* [7.1. BackTestStats](#6.1)
* [7.2. BackTestPlot](#6.2)
* [7.3. Baseline Stats](#6.3)
* [7.3. Compare to Stock Market Index](#6.4)
<a id='0'></a>
# Part 1. Problem Definition
This problem is to design an automated trading solution for single stock trading. We model the stock trading process as a Markov Decision Process (MDP). We then formulate our trading goal as a maximization problem.
The algorithm is trained using Deep Reinforcement Learning (DRL) algorithms and the components of the reinforcement learning environment are:
* Action: The action space describes the allowed actions that the agent interacts with the
environment. Normally, a ∈ A includes three actions: a ∈ {−1, 0, 1}, where −1, 0, 1 represent
selling, holding, and buying one stock. Also, an action can be carried upon multiple shares. We use
an action space {−k, ..., −1, 0, 1, ..., k}, where k denotes the number of shares. For example, "Buy
10 shares of AAPL" or "Sell 10 shares of AAPL" are 10 or −10, respectively
* Reward function: r(s, a, s′) is the incentive mechanism for an agent to learn a better action. The change of the portfolio value when action a is taken at state s and arriving at new state s', i.e., r(s, a, s′) = v′ − v, where v′ and v represent the portfolio
values at state s′ and s, respectively
* State: The state space describes the observations that the agent receives from the environment. Just as a human trader needs to analyze various information before executing a trade, so
our trading agent observes many different features to better learn in an interactive environment.
* Environment: Dow 30 consituents
The data of the single stock that we will be using for this case study is obtained from Yahoo Finance API. The data contains Open-High-Low-Close price and volume.
<a id='1'></a>
# Part 2. Getting Started- Load Python Packages
<a id='1.1'></a>
## 2.1. Install all the packages through FinRL library
```
# ## install finrl library
#!pip install git+https://github.com/AI4Finance-LLC/FinRL-Library.git
```
<a id='1.2'></a>
## 2.2. Check if the additional packages needed are present, if not install them.
* Yahoo Finance API
* pandas
* numpy
* matplotlib
* stockstats
* OpenAI gym
* stable-baselines
* tensorflow
* pyfolio
<a id='1.3'></a>
## 2.3. Import Packages
```
import warnings
warnings.filterwarnings("ignore")
import pandas as pd
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
# matplotlib.use('Agg')
from datetime import datetime, timedelta
%matplotlib inline
from finrl.apps import config
from finrl.neo_finrl.preprocessor.yahoodownloader import YahooDownloader
from finrl.neo_finrl.preprocessor.preprocessors import FeatureEngineer, data_split
from finrl.neo_finrl.env_stock_trading.env_stocktrading import StockTradingEnv
from finrl.drl_agents.stablebaselines3.models import DRLAgent,DRLEnsembleAgent
from finrl.plot import backtest_stats, backtest_plot, get_daily_return, get_baseline
from pprint import pprint
import sys
sys.path.append("../FinRL-Library")
import itertools
```
<a id='1.4'></a>
## 2.4. Create Folders
```
import os
if not os.path.exists("./" + config.DATA_SAVE_DIR):
os.makedirs("./" + config.DATA_SAVE_DIR)
if not os.path.exists("./" + config.TRAINED_MODEL_DIR):
os.makedirs("./" + config.TRAINED_MODEL_DIR)
if not os.path.exists("./" + config.TENSORBOARD_LOG_DIR):
os.makedirs("./" + config.TENSORBOARD_LOG_DIR)
if not os.path.exists("./" + config.RESULTS_DIR):
os.makedirs("./" + config.RESULTS_DIR)
import yfinance as yf
data_prefix = [f"{config.DOW_30_TICKER}", f"{config.nas_choosen}", f"{config.sp_choosen}"]
path_mark = ["dow", "nas", "sp"]
choose_number = 0
#注意更改ticker_list
config.START_DATE
config.END_DATE
print(data_prefix[choose_number])
time_current = datetime.now() + timedelta(hours=8)
time_current = time_current.strftime("%Y-%m-%d_%H:%M")
print(f"time_current ====={time_current}")
```
<a id='2'></a>
# Part 3. Download Data
Yahoo Finance is a website that provides stock data, financial news, financial reports, etc. All the data provided by Yahoo Finance is free.
* FinRL uses a class **YahooDownloader** to fetch data from Yahoo Finance API
* Call Limit: Using the Public API (without authentication), you are limited to 2,000 requests per hour per IP (or up to a total of 48,000 requests a day).
-----
class YahooDownloader:
Provides methods for retrieving daily stock data from
Yahoo Finance API
Attributes
----------
start_date : str
start date of the data (modified from config.py)
end_date : str
end date of the data (modified from config.py)
ticker_list : list
a list of stock tickers (modified from config.py)
Methods
-------
fetch_data()
Fetches data from yahoo API
```
#ticker_list = []
#ticker_list = [ticks[i] for i in index]
#getattr(ticker_list, data_prefix[choose_number])
# 缓存数据,如果日期或者股票列表发生变化,需要删除该缓存文件重新下载
SAVE_PATH = f"./datasets/{path_mark[choose_number]}.csv"
if os.path.exists(SAVE_PATH):
df = pd.read_csv(SAVE_PATH)
else:
#注意更改ticker_list
df = YahooDownloader(
config.START_DATE, #'2000-01-01',
config.END_DATE, # 2021-01-01,预计将改日期改为'2021-07-03'(今日日期)
ticker_list=config.DOW_30_TICKER#config.DOW_30_TICKER, config.nas_choosen, config.sp_choosen
).fetch_data()
df.to_csv(SAVE_PATH)
df.head()
df.tail()
df.shape
df.sort_values(['date','tic']).head()
```
# Part 4: Preprocess Data
Data preprocessing is a crucial step for training a high quality machine learning model. We need to check for missing data and do feature engineering in order to convert the data into a model-ready state.
* Add technical indicators. In practical trading, various information needs to be taken into account, for example the historical stock prices, current holding shares, technical indicators, etc. In this article, we demonstrate two trend-following technical indicators: MACD and RSI.
* Add turbulence index. Risk-aversion reflects whether an investor will choose to preserve the capital. It also influences one's trading strategy when facing different market volatility level. To control the risk in a worst-case scenario, such as financial crisis of 2007–2008, FinRL employs the financial turbulence index that measures extreme asset price fluctuation.
```
tech_indicators = ['macd',
'rsi_30',
'cci_30',
'dx_30']
fe = FeatureEngineer(
use_technical_indicator=True,
tech_indicator_list = tech_indicators,
use_turbulence=True,
user_defined_feature = False)
csv_name_func_processed = lambda time: f"./datasets/ensemble_{path_mark[choose_number]}_{time}_processed.csv"
SAVE_PATH = csv_name_func_processed(time_current)
if os.path.exists(SAVE_PATH):
processed = pd.read_csv(SAVE_PATH)
else:
processed = fe.preprocess_data(df)
processed.to_csv(SAVE_PATH)
list_ticker = processed["tic"].unique().tolist()
list_date = list(pd.date_range(processed['date'].min(),processed['date'].max()).astype(str))
combination = list(itertools.product(list_date,list_ticker))
processed_full = pd.DataFrame(combination,columns=["date","tic"]).merge(processed,on=["date","tic"],how="left")
processed_full = processed_full[processed_full['date'].isin(processed['date'])]
processed_full = processed_full.sort_values(['date','tic'])
processed_full = processed_full.fillna(0)
processed_full.sample(5)
```
<a id='4'></a>
# Part 5. Design Environment
Considering the stochastic and interactive nature of the automated stock trading tasks, a financial task is modeled as a **Markov Decision Process (MDP)** problem. The training process involves observing stock price change, taking an action and reward's calculation to have the agent adjusting its strategy accordingly. By interacting with the environment, the trading agent will derive a trading strategy with the maximized rewards as time proceeds.
Our trading environments, based on OpenAI Gym framework, simulate live stock markets with real market data according to the principle of time-driven simulation.
The action space describes the allowed actions that the agent interacts with the environment. Normally, action a includes three actions: {-1, 0, 1}, where -1, 0, 1 represent selling, holding, and buying one share. Also, an action can be carried upon multiple shares. We use an action space {-k,…,-1, 0, 1, …, k}, where k denotes the number of shares to buy and -k denotes the number of shares to sell. For example, "Buy 10 shares of AAPL" or "Sell 10 shares of AAPL" are 10 or -10, respectively. The continuous action space needs to be normalized to [-1, 1], since the policy is defined on a Gaussian distribution, which needs to be normalized and symmetric.
```
#train = data_split(processed_full, '2009-01-01','2019-01-01')
#trade = data_split(processed_full, '2019-01-01','2021-01-01')
#print(len(train))
#print(len(trade))
stock_dimension = len(processed_full.tic.unique())
state_space = 1 + 2*stock_dimension + len(tech_indicators)*stock_dimension
print(f"Stock Dimension: {stock_dimension}, State Space: {state_space}")
env_kwargs = {
"hmax": 100,
"initial_amount": 1000000,
"buy_cost_pct": 0.001,
"sell_cost_pct": 0.001,
"state_space": state_space,
"stock_dim": stock_dimension,
"tech_indicator_list": tech_indicators,
"action_space": stock_dimension,
"reward_scaling": 1e-4,
"print_verbosity":5
}
```
<a id='5'></a>
# Part 6: Implement DRL Algorithms
* The implementation of the DRL algorithms are based on **OpenAI Baselines** and **Stable Baselines**. Stable Baselines is a fork of OpenAI Baselines, with a major structural refactoring, and code cleanups.
* FinRL library includes fine-tuned standard DRL algorithms, such as DQN, DDPG,
Multi-Agent DDPG, PPO, SAC, A2C and TD3. We also allow users to
design their own DRL algorithms by adapting these DRL algorithms.
* In this notebook, we are training and validating 3 agents (A2C, PPO, DDPG) using Rolling-window Ensemble Method ([reference code](https://github.com/AI4Finance-LLC/Deep-Reinforcement-Learning-for-Automated-Stock-Trading-Ensemble-Strategy-ICAIF-2020/blob/80415db8fa7b2179df6bd7e81ce4fe8dbf913806/model/models.py#L92))
```
rebalance_window = 63 # rebalance_window is the number of days to retrain the model
validation_window = 63 # validation_window is the number of days to do validation and trading (e.g. if validation_window=63, then both validation and trading period will be 63 days)
"""
train_start = '2009-01-01'
train_end = "2016-10-03"
val_test_start = "2016-10-03"
val_test_end = "2021-09-18"
"""
train_start = "2009-01-01"
train_end = "2015-12-31"
val_test_start = "2015-12-31"
val_test_end = "2021-05-03"
ensemble_agent = DRLEnsembleAgent(df=processed_full,
train_period=(train_start,train_end),
val_test_period=(val_test_start,val_test_end),
rebalance_window=rebalance_window,
validation_window=validation_window,
**env_kwargs)
A2C_model_kwargs = {
'n_steps': 5,
'ent_coef': 0.01,
'learning_rate': 0.0005
}
PPO_model_kwargs = {
"ent_coef":0.01,
"n_steps": 2048,
"learning_rate": 0.00025,
"batch_size": 128
}
DDPG_model_kwargs = {
"action_noise":"ornstein_uhlenbeck",
"buffer_size": 50_000,
"learning_rate": 0.000005,
"batch_size": 128
}
timesteps_dict = {'a2c' : 1_000,
'ppo' : 1_000,
'ddpg' : 1_000
}
df_summary = ensemble_agent.run_ensemble_strategy(A2C_model_kwargs,
PPO_model_kwargs,
DDPG_model_kwargs,
timesteps_dict)
df_summary
```
<a id='6'></a>
# Part 7: Backtest Our Strategy
Backtesting plays a key role in evaluating the performance of a trading strategy. Automated backtesting tool is preferred because it reduces the human error. We usually use the Quantopian pyfolio package to backtest our trading strategies. It is easy to use and consists of various individual plots that provide a comprehensive image of the performance of a trading strategy.
```
unique_trade_date = processed_full[(processed_full.date > val_test_start)&(processed_full.date <= val_test_end)].date.unique()
import functools
def compare(A, B): # 名字可以随便取,不一定得是“compare"
return -1 if int(A.split("_")[-1][:-4])<int(B.split("_")[-1][:-4]) else 1
from typing import List
with open("paths.txt") as f:
paths: List[str] = f.readlines()
paths = [path.strip().split(" ")[-1] for path in paths]
paths.sort(key=functools.cmp_to_key(compare))
#paths.sort()
#print(paths)
df_account_value=pd.DataFrame()
for i in range(len(df_summary)):
iter = df_summary.iloc[i]["Iter"]
al = df_summary.iloc[i]["Model Used"]
path = f"results/account_value_validation_{al}_{iter}.csv"
#print(path, os.path.exists(path))
df_tmp = pd.read_csv(path)
df_account_value = df_account_value.append(df_tmp,ignore_index=True)
df_account_value
df_account_value.tail()
df_account_value.to_csv("results/account_value_all.csv", index=False)
df_account_value.head()
%matplotlib inline
df_account_value.account_value.plot()
```
<a id='6.1'></a>
## 7.1 BackTestStats
pass in df_account_value, this information is stored in env class
```
print("==============Get Backtest Results===========")
now = datetime.now().strftime('%Y%m%d-%Hh%M')
perf_stats_all = backtest_stats(account_value=df_account_value)
perf_stats_all = pd.DataFrame(perf_stats_all)
#baseline stats
print("==============Get Baseline Stats===========")
baseline_df = get_baseline(
ticker="^GSPC",
start = df_account_value.loc[0,'date'],
end = df_account_value.loc[len(df_account_value)-1,'date'])
stats = backtest_stats(baseline_df, value_col_name = 'close')
```
<a id='6.2'></a>
## 7.2 BackTestPlot
```
print("==============Compare to DJIA===========")
import pandas as pd
%matplotlib inline
# S&P 500: ^GSPC
# Dow Jones Index: ^DJI
# NASDAQ 100: ^NDX
#df_account_value = pd.read_csv("/mnt/wanyao/guiyi/hhf/RL-FIN/datasets/ensemble_2021-09-06_19:36_account_value.csv")
backtest_plot(df_account_value,
baseline_ticker = '^DJI',
baseline_start = df_account_value.loc[0,'date'],
baseline_end = df_account_value.loc[len(df_account_value)-1,'date'])
```
| github_jupyter |
# Text Classification using LSTM
This Code Template is for Text Classification using Long short-term memory in python
<img src="https://cdn.blobcity.com/assets/gpu_required.png" height="25" style="margin-bottom:-15px" />
### Required Packages
```
!pip install tensorflow
!pip install nltk
import pandas as pd
import tensorflow as tf
from tensorflow.keras.layers import Embedding
from tensorflow.keras.preprocessing.sequence import pad_sequences
from tensorflow.keras.models import Sequential
from tensorflow.keras.preprocessing.text import one_hot
from tensorflow.keras.layers import LSTM
from tensorflow.keras.layers import Dense
import nltk
import re
from nltk.corpus import stopwords
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
from sklearn.metrics import confusion_matrix
pip install tensorflow==2.1.0
```
### Initialization
Filepath of CSV file
```
df=pd.read_csv('/content/train.csv')
```
### Data Fetching
Pandas is an open-source, BSD-licensed library providing high-performance, easy-to-use data manipulation and data analysis tools.
We will use panda's library to read the CSV file using its storage path.And we use the head function to display the initial row or entry.
```
df.head()
###Drop Nan Values
df=df.dropna()
```
Target variable for prediction.
```
target=''
```
Text column containing all the text data
```
text=""
tf.__version__
### Vocabulary size
voc_size=5000
nltk.download('stopwords')
```
### Data Preprocessing
Since the majority of the machine learning models in the Sklearn library doesn't handle string category data and Null value, we have to explicitly remove or replace null values. The below snippet have functions, which removes the null value if any exists. And convert the string classes data in the datasets by encoding them to integer classes.
```
### Dataset Preprocessing
from nltk.stem.porter import PorterStemmer
ps = PorterStemmer()
corpus = []
for i in range(0, len(df)):
review = re.sub('[^a-zA-Z]', ' ', df[text][i])
review = review.lower()
review = review.split()
review = [ps.stem(word) for word in review if not word in stopwords.words('english')]
review = ' '.join(review)
corpus.append(review)
corpus[1:10]
onehot_repr=[one_hot(words,voc_size)for words in corpus]
onehot_repr
```
### Embedding Representation
```
sent_length=20
embedded_docs=pad_sequences(onehot_repr,padding='pre',maxlen=sent_length)
print(embedded_docs)
embedded_docs[0]
```
### Model
#LSTM
The LSTM model will learn a function that maps a sequence of past observations as input to an output observation. As such, the sequence of observations must be transformed into multiple examples from which the LSTM can learn.
Refer [API](https://towardsdatascience.com/illustrated-guide-to-lstms-and-gru-s-a-step-by-step-explanation-44e9eb85bf21) for the parameters
```
## Creating model
embedding_vector_features=40
model=Sequential()
model.add(Embedding(voc_size,embedding_vector_features,input_length=sent_length))
model.add(LSTM(100))
model.add(Dense(1,activation='sigmoid'))
model.compile(loss='binary_crossentropy',optimizer='adam',metrics=['accuracy'])
print(model.summary())
len(embedded_docs),y.shape
X=np.array(embedded_docs)
Y=df[target]
X_train, X_test, y_train, y_test = train_test_split(X, Y, test_size=0.33, random_state=42)
```
### Model Training
```
### Finally Training
model.fit(X_train,y_train,validation_data=(X_test,y_test),epochs=10,batch_size=64)
```
### Performance Metrics And Accuracy
```
y_pred=model.predict_classes(X_test)
confusion_matrix(y_test,y_pred)
```
## Model Accuracy
```
accuracy_score(y_test,y_pred)
```
#### Creator: Ageer Harikrishna , Github: [Profile](https://github.com/ageerHarikrishna)
| github_jupyter |
```
import bokeh.charts as bc
import matplotlib.pyplot as plt
import pandas as pd
from scipy import stats
import requests
import qgrid
import seaborn as sns
from IPython.display import Image
from scipy.stats import pearsonr
%matplotlib inline
qgrid.nbinstall()
```
### The purpose of this notebook is to document data analysis for a [Reproducibility Study](https://osf.io/ezcuj/wiki/home/?_ga=1.257932747.1792380294.1420926336) conducted in collaboration with the [Center for Open Science](https://cos.io) and fulfillment of my [undergraduate thesis](https://osf.io/3k4uy/) at Reed College.
###The original article was ["Errors are aversive: defensive motivation and the error-related negativity."](http://www.ncbi.nlm.nih.gov/pubmed/18271855)
```
data = pd.read_csv("rp.csv")
qgrid.show_grid(data, remote_js=True)
# subset trials depending on whether participant made an error,
# made an error in the previous trial ('predict'), or
# was correct in current and previous trial ('unpred')
error_trials = data[data['startle_type'] == 'error']
pred_trials = data[data['startle_type'] == 'predict']
unpred_trials = data[data['startle_type'] == 'unpred']
# restructure dataframe for python analysis
pred_error = pd.merge(error_trials, pred_trials, how="outer", on=["participant","gender","ERN","EPS","errors"], suffixes=('_error', '_pred'))
# further restructuring
final = pd.merge(pred_error, unpred_trials, how="outer", on=["participant","gender","ERN","EPS","errors"], suffixes=('_', '_unpred'))
final.rename(columns={'startle':'startle_unpred','startle_peak':'startle_peak_unpred'},inplace=True)
# drop superfluous columns naming startle type
final.drop(['startle_type_error','startle_type_pred','startle_type'], axis=1, inplace=True)
# na in participant 21's unpred startle trial excludes it from some analyes
error_trial = error_trials[error_trials.participant != 21]
pred_trials = pred_trials[pred_trials.participant != 21]
unpred_trials = unpred_trials[unpred_trials.participant != 21]
final = final[final.participant != 21]
# mean error-potentiated startle (EPS) amplitude
round(final['EPS'].mean(),2)
#standard error of the mean
round(stats.sem(final['EPS']),2)
# mean difference between error and correct trials
(final['startle_error'] - final['startle_unpred']).mean()
round(stats.sem(final['startle_error'] - final['startle_unpred']),2)
#main finding using one trial type for appropriate DF
corr_data = data[['ERN','EPS']]
corr_data.corr(method='pearson', min_periods=1)
```
# calculation of ERN from correct vs error ERPs
```
correct = pd.read_csv('data/rep_Cor_all.txt',delim_whitespace=True)
incorrect = pd.read_csv('data/rep_Inc_All.txt',delim_whitespace=True)
correct['File'] = correct['File'].apply(lambda x: x.translate(None, '_repCor'))
incorrect['File'] = incorrect['File'].apply(lambda x: x.translate(None, '_repInc'))
incorrect = incorrect[['File','2-rep_Inc']]
correct = correct[['File','2-rep_Cor']]
erp = pd.merge(correct, incorrect, on='File')
erp.rename(columns={'File':'participant','2-rep_Cor':'correct','2-rep_Inc':'error'},inplace=True)
erp['participant'] = erp['participant'].apply(lambda x: int(x))
erp['ERN'] = erp['error']-erp['correct']
erp.sort('participant',inplace=True)
# difference between ERPs on correct vs error trials
stats.ttest_rel(erp['correct'], erp['error'])
#mean ERN amplitude
round((erp['error'] - erp['correct']).mean(),2)
# ERN amplitude SEM
round(stats.sem(erp['error'] - erp['correct']),2)
```
### The main finding of the article replicated is A in the following figure.
```
Image(url="http://www.frontiersin.org/files/Articles/82577/fnhum-08-00064-HTML/image_m/fnhum-08-00064-g001.jpg")
```
### I failed to replicate this finding:
```
sns.jointplot(error_trials['ERN'],error_trials['EPS'],kind="reg",stat_func=pearsonr, color = "slategray")
```
However, the original author also found a more robust subsequent finding by dividing participants along median ERN and assessing correlation among those whose amplitudes were relatively high. We performed the same
```
high_amplitude = final[final['ERN'] < -6.91]
sns.jointplot(high_amplitude['ERN'],high_amplitude['EPS'],kind="reg",stat_func=pearsonr, color = "slategray")
```
and also found greater significance (greater correlation, *p* < .05) However, to investigate whether the outlier participant 1 had any bearing, we removed that data and reran the analysis:
```
high_amp_san_outlier = high_amplitude[high_amplitude['participant'] != 1]
sns.jointplot(high_amp_san_outlier['ERN'],high_amp_san_outlier['EPS'],kind="reg",stat_func=pearsonr, color = "slategray")
```
and found the correlation diminished again.
The original author was communicative and eager to compare findings, and so offered his original ERN and EPS data to pool with our own:
```
collaboration = pd.read_csv('data/collaboration.csv')
collaboration.rename(columns={'ERN (window difference wave)':'ERN','Error-potentiated startle':'EPS'},inplace=True)
sns.jointplot(collaboration['ERN'],collaboration['EPS'],kind="reg",stat_func=pearsonr, color = "slategray")
```
The correlation is present here, *p* < .05, but by coincidence we each had one participant with unusually high ERN amplitudes. We analyzed the correlation with these removed:
```
collab_sans_outlier = collaboration[collaboration['ERN'] > -20]
plot = sns.jointplot(collab_sans_outlier['ERN'],collab_sans_outlier['EPS'],kind="reg",stat_func=pearsonr, color = "slategray")
```
and the correlation is no longer present. It could be the case that the correlation is weak but real (and strong in 1 out of every 40 subjects), or it could be the case that in any sample of 40 subjects, random noise will cause at least one subject to show abnormally large ERN and EPS magnitudes, thereby strongly influencing the statistical relationship between these two variables.
We thank the original author Dr. Hajcak for providing input throughout the project. Quality replications are typically only possible with close correspondence with the original authors and we appreciate the willingness of Dr. Hajcak to contribute to this replication attempt.
| github_jupyter |
# Deep learning models for age prediction on EEG data
This notebook uses deep learning methods to predict the age of infants using EEG data. The EEG data is preprocessed as shown in the notebook 'Deep learning EEG_dataset preprocessing raw'.
```
import sys, os, fnmatch, csv
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
sys.path.insert(0, os.path.dirname(os.getcwd()))
from config import PATH_DATA_PROCESSED_DL_REDUCED, PATH_MODELS
```
## Load preprocessed data (reduced)
The data can be found in 'PATH_DATA_PROCESSED_DL_REDUCED'. This is a single folder with all the data and metadata. The EEG data is in the .zarr files and the metadata is in .csv files. The .zarr files are divided in chunks of 1-second epochs (average of 10 original EEG epochs) from the same subject and the metadata contains the information like the subject's identification code and age. The notebook "Deep learning EEG_DL dataset_reduction.ipynb" takes care of reducing the original processed data set to the reduced size.
Generator loads all the data into memory. The generators generate averaged epochs on the fly. The data is split in train/validation/test and no subject can be found in more than one of these splits.
Originally, we used the original EEG epochs and averaged 30 of them into a new EEG epoch in the generator object. This had two disadvantages: (1) many files had to be openened and closed during the training process, and (2) the data set was too large to fit into memory. Therefore, we decided to randomly create chunks of 10 EEG epochs and average those for each subject/age group. This reduced the data set from ±145GB to ±14.5 GB. We now use these already averaged EEG epochs as "original" epcohs, and average those another 3-5 times to reduce noise. We have also experimented with averaging all EEG epochs of a subject at a specific age into a single EEG epoch, but performance was lower, most likely because this reduced the size of the data set a lot.
```
%%time
# Load all the metadata
from sklearn.model_selection import train_test_split
# Step 1: Get all the files in the output folder
file_names = os.listdir(PATH_DATA_PROCESSED_DL_REDUCED)
# Step 2: Get the full paths of the files (without extensions)
files = [os.path.splitext(os.path.join(PATH_DATA_PROCESSED_DL_REDUCED, file_name))[0] for file_name in fnmatch.filter(file_names, "*.zarr")]
# Step 3: Load all the metadata
frames = []
for idx, feature_file in enumerate(files):
df_metadata = pd.read_csv(feature_file + ".csv")
frames.append(df_metadata)
df_metadata = pd.concat(frames)
# Step 4: Add missing age information based on the age group the subject is in
df_metadata['age_months'].fillna(df_metadata['age_group'], inplace=True)
df_metadata['age_days'].fillna(df_metadata['age_group']*30, inplace=True)
df_metadata['age_years'].fillna(df_metadata['age_group']/12, inplace=True)
# Step 5: List all the unique subject IDs
subject_ids = sorted(list(set(df_metadata["code"].tolist())))
from sklearn.model_selection import train_test_split
IDs_train, IDs_temp = train_test_split(subject_ids, test_size=0.3, random_state=42)
IDs_test, IDs_val = train_test_split(IDs_temp, test_size=0.5, random_state=42)
df_metadata
from dataset_generator_reduced import DataGeneratorReduced
# # Import libraries
# from tensorflow.keras.utils import Sequence
# import numpy as np
# import zarr
# import os
# class DataGeneratorReduced(Sequence):
# """Generates data for loading (preprocessed) EEG timeseries data.
# Create batches for training or prediction from given folders and filenames.
# """
# def __init__(self,
# list_IDs,
# BASE_PATH,
# metadata,
# gaussian_noise=0.0,
# n_average = 3,
# batch_size=10,
# iter_per_epoch = 1,
# n_timepoints = 501,
# n_channels=30,
# shuffle=True,
# warnings=False):
# """Initialization
# Args:
# --------
# list_IDs:
# list of all filename/label ids to use in the generator
# metadata:
# DataFrame containing all the metadata.
# n_average: int
# Number of EEG/time series epochs to average.
# batch_size:
# batch size at each iteration
# iter_per_epoch: int
# Number of iterations over all data points within one epoch.
# n_timepoints: int
# Timepoint dimension of data.
# n_channels:
# number of input channels
# shuffle:
# True to shuffle label indexes after every epoch
# """
# self.list_IDs = list_IDs
# self.BASE_PATH = BASE_PATH
# self.metadata = metadata
# self.metadata_temp = None
# self.gaussian_noise = gaussian_noise
# self.n_average = n_average
# self.batch_size = batch_size
# self.iter_per_epoch = iter_per_epoch
# self.n_timepoints = n_timepoints
# self.n_channels = n_channels
# self.shuffle = shuffle
# self.warnings = warnings
# self.on_epoch_end()
# # Store all data in here
# self.X_data_all = []
# self.y_data_all = []
# self.load_all_data()
# def __len__(self):
# """Denotes the number of batches per epoch
# return: number of batches per epoch
# """
# return int(np.floor(len(self.metadata_temp) / self.batch_size))
# def __getitem__(self, index):
# """Generate one batch of data
# Args:
# --------
# index: int
# index of the batch
# return: X and y when fitting. X only when predicting
# """
# # Generate indexes of the batch
# indexes = self.indexes[index * self.batch_size:((index + 1) * self.batch_size)]
# # Generate data
# X, y = self.generate_data(indexes)
# return X, y
# def load_all_data(self):
# """ Loads all data into memory. """
# for i, metadata_file in self.metadata_temp.iterrows():
# filename = os.path.join(self.BASE_PATH, metadata_file['cnt_file'] + '.zarr')
# X_data = np.zeros((0, self.n_channels, self.n_timepoints))
# data_signal = self.load_signal(filename)
# if (len(data_signal) == 0) and self.warnings:
# print(f"EMPTY SIGNAL, filename: {filename}")
# X_data = np.concatenate((X_data, data_signal), axis=0)
# self.X_data_all.append(X_data)
# self.y_data_all.append(metadata_file['age_months'])
# def on_epoch_end(self):
# """Updates indexes after each epoch."""
# # Create new metadata DataFrame with only the current subject IDs
# if self.metadata_temp is None:
# self.metadata_temp = self.metadata[self.metadata['code'].isin(self.list_IDs)].reset_index(drop=True)
# idx_base = np.arange(len(self.metadata_temp))
# idx_epoch = np.tile(idx_base, self.iter_per_epoch)
# self.indexes = idx_epoch
# if self.shuffle == True:
# np.random.shuffle(self.indexes)
# def generate_data(self, indexes):
# """Generates data containing batch_size averaged time series.
# Args:
# -------
# list_IDs_temp: list
# list of label ids to load
# return: batch of averaged time series
# """
# X_data = np.zeros((0, self.n_channels, self.n_timepoints))
# y_data = []
# for index in indexes:
# X = self.create_averaged_epoch(self.X_data_all[index])
# X_data = np.concatenate((X_data, X), axis=0)
# y_data.append(self.y_data_all[index])
# return np.swapaxes(X_data,1,2), np.array(y_data).reshape((-1,1))
# def create_averaged_epoch(self,
# data_signal):
# """
# Function to create averages of self.n_average epochs.
# Will create one averaged epoch per found unique label from self.n_average random epochs.
# Args:
# --------
# data_signal: numpy array
# Data from one person as numpy array
# """
# # Create new data collection:
# X_data = np.zeros((0, self.n_channels, self.n_timepoints))
# num_epochs = len(data_signal)
# if num_epochs >= self.n_average:
# select = np.random.choice(num_epochs, self.n_average, replace=False)
# signal_averaged = np.mean(data_signal[select,:,:], axis=0)
# else:
# if self.warnings:
# print("Found only", num_epochs, " epochs and will take those!")
# signal_averaged = np.mean(data_signal[:,:,:], axis=0)
# X_data = np.concatenate([X_data, np.expand_dims(signal_averaged, axis=0)], axis=0)
# if self.gaussian_noise != 0.0:
# X_data += np.random.normal(0, self.gaussian_noise, X_data.shape)
# return X_data
# def load_signal(self,
# filename):
# """Load EEG signal from one person.
# Args:
# -------
# filename: str
# filename...
# return: loaded array
# """
# return zarr.open(os.path.join(filename), mode='r')
%%time
# IDs_train = [152, 18, 11, 435, 617]
# IDs_val = [180, 105, 19, 332]
train_generator_noise = DataGeneratorReduced(list_IDs = IDs_train,
BASE_PATH = PATH_DATA_PROCESSED_DL_REDUCED,
metadata = df_metadata,
n_average = 3,
batch_size = 10,
gaussian_noise=0.01,
iter_per_epoch = 3,
n_timepoints = 501,
n_channels=30,
shuffle=True)
val_generator = DataGeneratorReduced(list_IDs = IDs_val,
BASE_PATH = PATH_DATA_PROCESSED_DL_REDUCED,
metadata = df_metadata,
n_average = 3,
batch_size = 10,
n_timepoints = 501,
iter_per_epoch = 3,
n_channels=30,
shuffle=True)
```
# Helper functions
```
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers, Input, Sequential
from tensorflow.keras.layers import Bidirectional, LSTM, Dropout, BatchNormalization, Dense, Conv1D, LeakyReLU, AveragePooling1D, Flatten, Reshape, MaxPooling1D
from tensorflow.keras.optimizers import Adam, Adadelta, SGD
from tensorflow.keras.callbacks import ModelCheckpoint, EarlyStopping, ReduceLROnPlateau
from tensorflow.keras.metrics import RootMeanSquaredError, MeanAbsoluteError
import time
n_timesteps = 501
n_features = 30
n_outputs = 1
input_shape = (n_timesteps, n_features)
```
# Testing of different architectures
Below we will test multiple different architectures, most of them as discussed in "Deep learning for time series classification: a review", by Ismail Fawaz et al (2019). Most of them are inspired again on other papers. Refer to the Ismail Fawaz paper for the original papers.
1. Fully-connected NN
2. CNN
3. ResNet
4. Encoder
5. Time-CNN
Other architectures to test:
6. BLSTM-LSTM
7. InceptionTime
# 1. Fully connected NN
```
def fully_connected_model():
""" Returns the fully connected model from Ismail Fawaz et al. (2019). """
input_layer = keras.layers.Input(input_shape)
input_layer_flattened = keras.layers.Flatten()(input_layer)
layer_1 = keras.layers.Dropout(0.1)(input_layer_flattened)
layer_1 = keras.layers.Dense(500, activation='relu')(layer_1)
layer_2 = keras.layers.Dropout(0.2)(layer_1)
layer_2 = keras.layers.Dense(500, activation='relu')(layer_2)
layer_3 = keras.layers.Dropout(0.2)(layer_2)
layer_3 = keras.layers.Dense(500, activation='relu')(layer_3)
output_layer = keras.layers.Dropout(0.3)(layer_3)
output_layer = keras.layers.Dense(1)(output_layer)
model = keras.models.Model(inputs=input_layer, outputs=output_layer)
return model
model = fully_connected_model()
optimizer = Adadelta(learning_rate=0.01)
model.compile(loss='mean_squared_error',
optimizer=optimizer,
metrics=[RootMeanSquaredError(), MeanAbsoluteError()])
# 01 seems to be incorrect (makes too many predictions, changed model)
# Fully_connected_regressor_01: MSE, Adadelta, N_average=30, 5000 epochs, ES=1000, RLR=200, gaussian=0.01
# Fully_connected_regressor_02: MSE, Adadelta, N_average=30, 5000 epochs, ES=1000, RLR=200, gaussian=0.01
output_filename = 'Fully_connected_regressor_03'
output_file = os.path.join(PATH_MODELS, output_filename)
checkpointer = ModelCheckpoint(filepath = output_file + ".hdf5", monitor='val_loss', verbose=1, save_best_only=True)
earlystopper = EarlyStopping(monitor='val_loss', patience=1000, verbose=1)
reduce_lr = ReduceLROnPlateau(monitor='val_loss', factor=0.5, patience=200, min_lr=0.0001, verbose=1)
%%time
epochs = 5000
# fit network
history = model.fit(x=train_generator_noise,
validation_data=val_generator,
epochs=epochs,
callbacks=[checkpointer, earlystopper, reduce_lr])
```
# 2. CNN
```
def cnn_model():
""" Returns the CNN (FCN) model from Ismail Fawaz et al. (2019). """
input_layer = keras.layers.Input(input_shape)
conv1 = keras.layers.Conv1D(filters=128, kernel_size=8, padding='same')(input_layer)
conv1 = keras.layers.BatchNormalization()(conv1)
conv1 = keras.layers.Activation(activation='relu')(conv1)
conv2 = keras.layers.Conv1D(filters=256, kernel_size=5, padding='same')(conv1)
conv2 = keras.layers.BatchNormalization()(conv2)
conv2 = keras.layers.Activation('relu')(conv2)
conv3 = keras.layers.Conv1D(128, kernel_size=3, padding='same')(conv2)
conv3 = keras.layers.BatchNormalization()(conv3)
conv3 = keras.layers.Activation('relu')(conv3)
gap_layer = keras.layers.GlobalAveragePooling1D()(conv3)
output_layer = keras.layers.Dense(1)(gap_layer)
model = keras.models.Model(inputs=input_layer, outputs=output_layer)
return model
model = cnn_model()
optimizer = Adam(learning_rate=0.01)
model.compile(loss='mean_squared_error',
optimizer=optimizer,
metrics=[RootMeanSquaredError(), MeanAbsoluteError()])
# CNN_regressor_01: MSE, Adam, N_average=30, 2000 epochs, ES=250, RLR=50, gaussian=0.01
output_filename = 'CNN_regressor_03'
output_file = os.path.join(PATH_MODELS, output_filename)
checkpointer = ModelCheckpoint(filepath = output_file + ".hdf5", monitor='val_loss', verbose=1, save_best_only=True)
earlystopper = EarlyStopping(monitor='val_loss', patience=250, verbose=1)
reduce_lr = ReduceLROnPlateau(monitor='val_loss', factor=0.5, patience=50, min_lr=0.0001, verbose=1)
%%time
epochs = 2000
# fit network
history = model.fit(x=train_generator_noise,
validation_data=val_generator,
epochs=epochs,
callbacks=[checkpointer, earlystopper, reduce_lr])
```
# 3. ResNet
```
def resnet_model():
""" Returns the ResNet model from Ismail Fawaz et al. (2019). """
n_feature_maps = 64
input_layer = keras.layers.Input(input_shape)
# BLOCK 1
conv_x = keras.layers.Conv1D(filters=n_feature_maps, kernel_size=8, padding='same')(input_layer)
conv_x = keras.layers.BatchNormalization()(conv_x)
conv_x = keras.layers.Activation('relu')(conv_x)
conv_y = keras.layers.Conv1D(filters=n_feature_maps, kernel_size=5, padding='same')(conv_x)
conv_y = keras.layers.BatchNormalization()(conv_y)
conv_y = keras.layers.Activation('relu')(conv_y)
conv_z = keras.layers.Conv1D(filters=n_feature_maps, kernel_size=3, padding='same')(conv_y)
conv_z = keras.layers.BatchNormalization()(conv_z)
# expand channels for the sum
shortcut_y = keras.layers.Conv1D(filters=n_feature_maps, kernel_size=1, padding='same')(input_layer)
shortcut_y = keras.layers.BatchNormalization()(shortcut_y)
output_block_1 = keras.layers.add([shortcut_y, conv_z])
output_block_1 = keras.layers.Activation('relu')(output_block_1)
# BLOCK 2
conv_x = keras.layers.Conv1D(filters=n_feature_maps * 2, kernel_size=8, padding='same')(output_block_1)
conv_x = keras.layers.BatchNormalization()(conv_x)
conv_x = keras.layers.Activation('relu')(conv_x)
conv_y = keras.layers.Conv1D(filters=n_feature_maps * 2, kernel_size=5, padding='same')(conv_x)
conv_y = keras.layers.BatchNormalization()(conv_y)
conv_y = keras.layers.Activation('relu')(conv_y)
conv_z = keras.layers.Conv1D(filters=n_feature_maps * 2, kernel_size=3, padding='same')(conv_y)
conv_z = keras.layers.BatchNormalization()(conv_z)
# expand channels for the sum
shortcut_y = keras.layers.Conv1D(filters=n_feature_maps * 2, kernel_size=1, padding='same')(output_block_1)
shortcut_y = keras.layers.BatchNormalization()(shortcut_y)
output_block_2 = keras.layers.add([shortcut_y, conv_z])
output_block_2 = keras.layers.Activation('relu')(output_block_2)
# BLOCK 3
conv_x = keras.layers.Conv1D(filters=n_feature_maps * 2, kernel_size=8, padding='same')(output_block_2)
conv_x = keras.layers.BatchNormalization()(conv_x)
conv_x = keras.layers.Activation('relu')(conv_x)
conv_y = keras.layers.Conv1D(filters=n_feature_maps * 2, kernel_size=5, padding='same')(conv_x)
conv_y = keras.layers.BatchNormalization()(conv_y)
conv_y = keras.layers.Activation('relu')(conv_y)
conv_z = keras.layers.Conv1D(filters=n_feature_maps * 2, kernel_size=3, padding='same')(conv_y)
conv_z = keras.layers.BatchNormalization()(conv_z)
# no need to expand channels because they are equal
shortcut_y = keras.layers.BatchNormalization()(output_block_2)
output_block_3 = keras.layers.add([shortcut_y, conv_z])
output_block_3 = keras.layers.Activation('relu')(output_block_3)
# FINAL
gap_layer = keras.layers.GlobalAveragePooling1D()(output_block_3)
output_layer = keras.layers.Dense(1)(gap_layer)
model = keras.models.Model(inputs=input_layer, outputs=output_layer)
return model
model = resnet_model()
optimizer = Adam(learning_rate=0.01)
model.compile(loss='mean_squared_error',
optimizer=optimizer,
metrics=[RootMeanSquaredError(), MeanAbsoluteError()])
# ResNet_regressor_01: MSE, Adam, N_average=30, 1500 epochs, ES=250, RLR=50, gaussian=0.01
output_filename = 'ResNet_regressor_02'
output_file = os.path.join(PATH_MODELS, output_filename)
checkpointer = ModelCheckpoint(filepath = output_file + ".hdf5", monitor='val_loss', verbose=1, save_best_only=True)
earlystopper = EarlyStopping(monitor='val_loss', patience=250, verbose=1)
reduce_lr = ReduceLROnPlateau(monitor='val_loss', factor=0.5, patience=50, min_lr=0.0001, verbose=1)
%%time
epochs = 1500
# fit network
history = model.fit(x=train_generator_noise,
validation_data=val_generator,
epochs=epochs,
callbacks=[checkpointer, earlystopper, reduce_lr])
```
# 4. Encoder
```
import tensorflow_addons as tfa
def encoder_model():
""" Returns the Encoder model from Ismail Fawaz et al. (2019). """
input_layer = keras.layers.Input(input_shape)
# conv block -1
conv1 = keras.layers.Conv1D(filters=128,kernel_size=5,strides=1,padding='same')(input_layer)
conv1 = tfa.layers.InstanceNormalization()(conv1)
conv1 = keras.layers.PReLU(shared_axes=[1])(conv1)
conv1 = keras.layers.Dropout(rate=0.2)(conv1)
conv1 = keras.layers.MaxPooling1D(pool_size=2)(conv1)
# conv block -2
conv2 = keras.layers.Conv1D(filters=256,kernel_size=11,strides=1,padding='same')(conv1)
conv2 = tfa.layers.InstanceNormalization()(conv2)
conv2 = keras.layers.PReLU(shared_axes=[1])(conv2)
conv2 = keras.layers.Dropout(rate=0.2)(conv2)
conv2 = keras.layers.MaxPooling1D(pool_size=2)(conv2)
# conv block -3
conv3 = keras.layers.Conv1D(filters=512,kernel_size=21,strides=1,padding='same')(conv2)
conv3 = tfa.layers.InstanceNormalization()(conv3)
conv3 = keras.layers.PReLU(shared_axes=[1])(conv3)
conv3 = keras.layers.Dropout(rate=0.2)(conv3)
# split for attention
attention_data = keras.layers.Lambda(lambda x: x[:,:,:256])(conv3)
attention_softmax = keras.layers.Lambda(lambda x: x[:,:,256:])(conv3)
# attention mechanism
attention_softmax = keras.layers.Softmax()(attention_softmax)
multiply_layer = keras.layers.Multiply()([attention_softmax,attention_data])
# last layer
dense_layer = keras.layers.Dense(units=256,activation='sigmoid')(multiply_layer)
dense_layer = tfa.layers.InstanceNormalization()(dense_layer)
# output layer
flatten_layer = keras.layers.Flatten()(dense_layer)
output_layer = keras.layers.Dense(1)(flatten_layer)
model = keras.models.Model(inputs=input_layer, outputs=output_layer)
return model
model = encoder_model()
optimizer = Adam(learning_rate=0.00001)
model.compile(loss='mean_squared_error',
optimizer=optimizer,
metrics=[RootMeanSquaredError(), MeanAbsoluteError()])
# Encoder_regressor_01: MSE, Adam, N_average=30, 1500 epochs, ES=250, RLR=50, gaussian=0.01 (LR = 0.0001, no reduction)
output_filename = 'Encoder_regressor_02'
output_file = os.path.join(PATH_MODELS, output_filename)
checkpointer = ModelCheckpoint(filepath = output_file + ".hdf5", monitor='val_loss', verbose=1, save_best_only=True)
%%time
epochs = 100
# fit network
history = model.fit(x=train_generator_noise,
validation_data=val_generator,
epochs=epochs,
callbacks=[checkpointer])
```
# 5. Time-CNN
```
# https://github.com/hfawaz/dl-4-tsc/blob/master/classifiers/cnn.py
def timecnn_model():
""" Returns the Time-CNN model from Ismail Fawaz et al. (2019). """
padding = 'valid'
input_layer = keras.layers.Input(input_shape)
conv1 = keras.layers.Conv1D(filters=6,kernel_size=7,padding=padding,activation='sigmoid')(input_layer)
conv1 = keras.layers.AveragePooling1D(pool_size=3)(conv1)
conv2 = keras.layers.Conv1D(filters=12,kernel_size=7,padding=padding,activation='sigmoid')(conv1)
conv2 = keras.layers.AveragePooling1D(pool_size=3)(conv2)
flatten_layer = keras.layers.Flatten()(conv2)
output_layer = keras.layers.Dense(units=1)(flatten_layer)
model = keras.models.Model(inputs=input_layer, outputs=output_layer)
return model
model = cnn_model()
optimizer = Adam(learning_rate=0.01)
model.compile(loss='mean_squared_error',
optimizer=optimizer,
metrics=[RootMeanSquaredError(), MeanAbsoluteError()])
# TimeCNN_regressor_01: MSE, Adam, N_average=30, 2000 epochs, ES=250, RLR=50, gaussian=0.01
output_filename = 'TimeCNN_regressor_02'
output_file = os.path.join(PATH_MODELS, output_filename)
checkpointer = ModelCheckpoint(filepath = output_file + ".hdf5", monitor='val_loss', verbose=1, save_best_only=True)
earlystopper = EarlyStopping(monitor='val_loss', patience=250, verbose=1)
reduce_lr = ReduceLROnPlateau(monitor='val_loss', factor=0.5, patience=50, min_lr=0.0001, verbose=1)
%%time
epochs = 2000
# fit network
history = model.fit(x=train_generator_noise,
validation_data=val_generator,
epochs=epochs,
callbacks=[checkpointer, earlystopper, reduce_lr])
```
# 6. BLSTM-LSTM
```
def blstm_lstm_model():
""" Returns the BLSTM-LSTM model from Kaushik et al. (2019). """
# MARK: This model compresses too much in the last phase, check if possible to improve.
model = keras.Sequential()
# BLSTM layer
model.add(Bidirectional(LSTM(256, return_sequences=True), input_shape=input_shape))
model.add(Dropout(.2))
model.add(BatchNormalization())
# LSTM layer
model.add(LSTM(128, return_sequences=True))
model.add(BatchNormalization())
# LSTM layer
model.add(LSTM(64, return_sequences=False))
model.add(BatchNormalization())
# Fully connected layer
model.add(Dense(32))
model.add(Dense(n_outputs))
return model
model = blstm_lstm_model()
optimizer = Adam(learning_rate=0.01)
model.compile(loss='mean_squared_error',
optimizer=optimizer,
metrics=[RootMeanSquaredError(), MeanAbsoluteError()])
# BLSTM_regressor_01: MSE, Adam, N_average=30, 1500 epochs, ES=250, RLR=50, gaussian=0.01
output_filename = 'BLSTM_regressor_01'
output_file = os.path.join(PATH_MODELS, output_filename)
checkpointer = ModelCheckpoint(filepath = output_file + ".hdf5", monitor='val_loss', verbose=1, save_best_only=True)
earlystopper = EarlyStopping(monitor='val_loss', patience=250, verbose=1)
reduce_lr = ReduceLROnPlateau(monitor='val_loss', factor=0.5, patience=50, min_lr=0.0001, verbose=1)
model.summary()
%%time
epochs = 1500
# fit network
history = model.fit(x=train_generator_noise,
validation_data=val_generator,
epochs=epochs,
callbacks=[checkpointer, earlystopper, reduce_lr])
```
# 7. InceptionTime
```
class Regressor_Inception:
def __init__(self, output_directory, input_shape, verbose=False, build=True, batch_size=64,
nb_filters=32, use_residual=True, use_bottleneck=True, depth=6, kernel_size=41, nb_epochs=1500):
self.output_directory = output_directory
self.nb_filters = nb_filters
self.use_residual = use_residual
self.use_bottleneck = use_bottleneck
self.depth = depth
self.kernel_size = kernel_size - 1
self.callbacks = None
self.batch_size = batch_size
self.bottleneck_size = 32
self.nb_epochs = nb_epochs
if build == True:
self.model = self.build_model(input_shape)
if (verbose == True):
self.model.summary()
self.verbose = verbose
self.model.save_weights(self.output_directory + 'inception_model_init.hdf5')
def _inception_module(self, input_tensor, stride=1, activation='linear'):
if self.use_bottleneck and int(input_tensor.shape[-1]) > 1:
input_inception = tf.keras.layers.Conv1D(filters=self.bottleneck_size, kernel_size=1,
padding='same', activation=activation, use_bias=False)(input_tensor)
else:
input_inception = input_tensor
# kernel_size_s = [3, 5, 8, 11, 17]
kernel_size_s = [self.kernel_size // (2 ** i) for i in range(3)]
conv_list = []
for i in range(len(kernel_size_s)):
conv_list.append(tf.keras.layers.Conv1D(filters=self.nb_filters, kernel_size=kernel_size_s[i],
strides=stride, padding='same', activation=activation, use_bias=False)(
input_inception))
max_pool_1 = tf.keras.layers.MaxPool1D(pool_size=3, strides=stride, padding='same')(input_tensor)
conv_6 = tf.keras.layers.Conv1D(filters=self.nb_filters, kernel_size=1,
padding='same', activation=activation, use_bias=False)(max_pool_1)
conv_list.append(conv_6)
x = tf.keras.layers.Concatenate(axis=2)(conv_list)
x = tf.keras.layers.BatchNormalization()(x)
x = tf.keras.layers.Activation(activation='relu')(x)
return x
def _shortcut_layer(self, input_tensor, out_tensor):
shortcut_y = tf.keras.layers.Conv1D(filters=int(out_tensor.shape[-1]), kernel_size=1,
padding='same', use_bias=False)(input_tensor)
shortcut_y = tf.keras.layers.BatchNormalization()(shortcut_y)
x = tf.keras.layers.Add()([shortcut_y, out_tensor])
x = tf.keras.layers.Activation('relu')(x)
return x
def build_model(self, input_shape):
input_layer = tf.keras.layers.Input(input_shape)
x = input_layer
input_res = input_layer
for d in range(self.depth):
x = self._inception_module(x)
if self.use_residual and d % 3 == 2:
x = self._shortcut_layer(input_res, x)
input_res = x
pooling_layer = tf.keras.layers.AveragePooling1D(pool_size=50)(x)
flat_layer = tf.keras.layers.Flatten()(pooling_layer)
dense_layer = tf.keras.layers.Dense(128, activation='relu')(flat_layer)
output_layer = tf.keras.layers.Dense(1)(dense_layer)
model = tf.keras.models.Model(inputs=input_layer, outputs=output_layer)
return model
model = Regressor_Inception(PATH_MODELS, input_shape, verbose=True).model
optimizer = Adam(learning_rate=0.01)
model.compile(loss='mean_squared_error',
optimizer=optimizer,
metrics=[RootMeanSquaredError(), MeanAbsoluteError()])
# 'Inception_regressor_01' (n_average = 40, gaussian_noise = 0.01, MAE)
# 'Inception_regressor_02' (n_average = 1, gaussian_noise = 0.01, MAE)
# 'Inception_regressor_03' (n_average = 40, gaussian_noise = 0.01, MSE)
# 'Inception_regressor_04' (n_average = 1, gaussian_noise = 0.01, MSE)
# 'Inception_regressor_05' (n_average = 100, gaussian_noise = 0.01, MAE)
output_filename = 'Inception_regressor_05'
output_file = os.path.join(PATH_MODELS, output_filename)
checkpointer = ModelCheckpoint(filepath = output_file + ".hdf5", monitor='val_loss', verbose=1, save_best_only=True)
earlystopper = EarlyStopping(monitor='val_loss', patience=100, verbose=1)
reduce_lr = ReduceLROnPlateau(monitor='val_loss', factor=0.5, patience=20, min_lr=0.0001, verbose=1)
%%time
epochs = 1500
# fit network
history = model.fit(x=train_generator_noise,
validation_data=val_generator,
epochs=epochs,
callbacks = [checkpointer, earlystopper, reduce_lr])
```
# Compare models
### Helper functions for comparison
```
# 'Inception_regressor_01' (n_average = 40, gaussian_noise = 0.01, MAE)
model_path = os.path.join(PATH_MODELS, 'Inception_regressor_01.hdf5')
loaded_model = tf.keras.models.load_model(model_path)
evaluate_model(loaded_model)
# 'Inception_regressor_02' (n_average = 1, gaussian_noise = 0.01, MAE)
model_path = os.path.join(PATH_MODELS, 'Inception_regressor_02.hdf5')
loaded_model = tf.keras.models.load_model(model_path)
evaluate_model(loaded_model)
# 'Inception_regressor_03' (n_average = 40, gaussian_noise = 0.01, MSE)
model_path = os.path.join(PATH_MODELS, 'Inception_regressor_03.hdf5')
loaded_model = tf.keras.models.load_model(model_path)
evaluate_model(loaded_model)
# 'Inception_regressor_04' (n_average = 1, gaussian_noise = 0.01, MSE)
model_path = os.path.join(PATH_MODELS, 'Inception_regressor_04.hdf5')
loaded_model = tf.keras.models.load_model(model_path)
evaluate_model(loaded_model)
# 'Inception_regressor_05' (n_average = 100, gaussian_noise = 0.01, MAE)
model_path = os.path.join(PATH_MODELS, 'Inception_regressor_05.hdf5')
loaded_model = tf.keras.models.load_model(model_path)
evaluate_model(loaded_model)
# Fully_connected_regressor_02: MSE, Adadelta, N_average=30, 5000 epochs, ES=1000, RLR=200, gaussian=0.01
model_path = os.path.join(PATH_MODELS, 'Fully_connected_regressor_02.hdf5')
loaded_model = tf.keras.models.load_model(model_path)
evaluate_model(loaded_model)
# CNN_regressor_01: MSE, Adam, N_average=30, 2000 epochs, ES=250, RLR=50, gaussian=0.01
model_path = os.path.join(PATH_MODELS, 'CNN_regressor_01.hdf5')
loaded_model = tf.keras.models.load_model(model_path)
evaluate_model(loaded_model)
# ResNet_regressor_01: MSE, Adam, N_average=30, 1500 epochs, ES=250, RLR=50, gaussian=0.01
model_path = os.path.join(PATH_MODELS, 'ResNet_regressor_01.hdf5')
loaded_model = tf.keras.models.load_model(model_path)
evaluate_model(loaded_model)
# Encoder_regressor_01: MSE, Adam, N_average=30, 1500 epochs, ES=250, RLR=50, gaussian=0.01 (LR = 0.0001, no reduction)
model_path = os.path.join(PATH_MODELS, 'Encoder_regressor_01.hdf5')
loaded_model = tf.keras.models.load_model(model_path)
evaluate_model(loaded_model)
# TimeCNN_regressor_01: MSE, Adam, N_average=30, 2000 epochs, ES=250, RLR=50, gaussian=0.01
model_path = os.path.join(PATH_MODELS, 'TimeCNN_regressor_01.hdf5')
loaded_model = tf.keras.models.load_model(model_path)
evaluate_model(loaded_model)
```
| github_jupyter |
# Descriptive statistics
```
import numpy as np
import seaborn as sns
import scipy.stats as st
import matplotlib.pyplot as plt
import matplotlib.mlab as mlab
import pandas as pd
import statsmodels.api as sm
import statistics
import os
from scipy.stats import norm
```
## Probability data, binomial distribution
We already got to know data that follow a binomial distribution, but we actually had not looked at the distribution. We will do this now. 10% of the 100 cells we count have deformed nuclei. To illustrate the distribution we will count repeatedly....
```
n = 100 # number of trials
p = 0.1 # probability of each trial
s = np.random.binomial(n, p, 1000) #simulation repeating the experiment 1000 times
print(s)
```
As you can see, the result of the distribution is in absolute counts, not proportions - they can easyly be converted by deviding with n, but they dont have to...
```
props = s/n
print(props)
```
Now we plot the distribution. The easiest first look is a histogram.
```
plt.hist(props, bins = 5)
plt.xlabel("proportion")
plt.ylabel("frequency")
plt.show()
```
The resolution is a bit inappropriate, given that we deal with integers. To increase the bin number would be a good idea. Maybe we should also plot a confidence interval.
```
CI= sm.stats.proportion_confint(n*p, n, alpha=0.05)
print(CI)
plt.axvspan(CI[0],CI[1], alpha=0.2, color='yellow')
plt.hist(props, bins = 50)
plt.xlabel("proportion")
plt.ylabel("frequency")
plt.axvline(p, color="black")
```
In a binomial distribution, the distribution is given by the proportion and the sample size. Therefore we could calculate a confidence interval from one measurement.
#### How can we now describe the distribution?
Summary statistics:
```
print("the minimum is:", min(props))
print("the maximum is:", max(props))
print(statistics.mean(props))
```
Is the mean a good way to look at our distribution?
```
n = 50 # number of trials
p = 0.02 # probability of each trial
s = np.random.binomial(n, p, 1000) #simulation repeating the experiment 1000 times
props = s/n
CI= sm.stats.proportion_confint(n*p, n, alpha=0.05)
print(CI)
plt.axvspan(CI[0],CI[1], alpha=0.2, color='yellow')
plt.hist(props, bins = 20)
plt.xlabel("proportion")
plt.ylabel("frequency")
plt.axvline(p, color="black")
plt.axvline(statistics.mean(props), color="red")
print(statistics.mean(props))
```
## Count data/ the Poisson distribution
The Poisson distribution is built on count data, e.g. the numbers of raisins in a Dresdner Christstollen, the number of geese at any given day between Blaues Wunder and Waldschlösschenbrücke, or radioactive decay. So lets use a Geiger counter and count the numbers of decay per min.
```
freq =1.6
s = np.random.poisson(freq, 1000)
plt.hist(s, bins = 20)
plt.xlabel("counts per minute")
plt.ylabel("frequency")
plt.axvline(freq, color="black")
```
### Confidence intervals for a Poisson distribution
Similar to the binomial distribution, the distribution is defined by sample size and the mean.
Also for Poisson, one can calculate an also asymmetrical confidence interval:
```
freq =1.6
s = np.random.poisson(freq, 1000)
CI = st.poisson.interval(0.95,freq)
plt.axvspan(CI[0],CI[1], alpha=0.2, color='yellow')
plt.hist(s, bins = 20)
plt.xlabel("counts per minute")
plt.ylabel("frequency")
plt.axvline(freq, color="black")
```
For a poisson distribution, poisson error can be reduced, when increasing the counting population, in our case lets count for 10 min instead of 1 min, and see what happens.
```
CI = np.true_divide(st.poisson.interval(0.95,freq*10),10)
print(CI)
s = np.true_divide(np.random.poisson(freq*10, 1000),10)
plt.axvspan(CI[0],CI[1], alpha=0.2, color='yellow')
plt.hist(s, bins = 70)
plt.xlabel("counts per minute")
plt.ylabel("frequency")
plt.axvline(freq, color="black")
```
What is the difference between Poisson and Binomial? Aren't they both kind of looking at count data?
Yes, BUT:
Binomial counts events versus another event, e.g. for the cells there are two options, normal versus deformed. A binomial distribution is about comparing the two options.
Poisson counts with an open end, e.g. number of mutations.
## Continuous data
Let's import the count data you have generated with Robert. When you download it from Google sheets (https://docs.google.com/spreadsheets/d/1Ek-23Soro5XZ3y1kJHpvaTaa1f4n2C7G3WX0qddD-78/edit#gid=0), it comes with spaces. Try to avoid spaces and special characters in file names, they tend to make trouble.
I renamed it to 'BBBC001.csv'.
```
dat = pd.read_csv(os.path.join('data','BBBC001.csv'), header=1, sep=';')
print(dat)
```
For now we will focus on the manual counts, visualise it and perform summary statistics.
```
man_count = dat.iloc[:,1].values
plt.hist(man_count,bins=100)
```
There are different alternatives of displaying such data, some of which independent of distribution. You will find documentation in the graph galery: https://www.python-graph-gallery.com/
```
sns.kdeplot(man_count)
```
A density plot is sometimes helpful to see the distribution, but be aware of the smoothing and that you loose the information on sample size.
```
sns.stripplot(y=man_count)
sns.swarmplot(y=man_count)
sns.violinplot(y=man_count)
```
this plot is useful, but the density function can sometimes be misleading and lead to artefacts dependent on the sample size. Unless explicitely stated, sample sizes are usually normalised and therefore hidden!
```
sns.boxplot(y=man_count)
```
Be aware that boxplots hide the underlying distribution and the sample size.
So the "safest" plot, when in doubt, is to combine boxplot and jitter:
```
ax = sns.swarmplot(y=man_count, color="black")
ax = sns.boxplot(y=man_count,color="lightgrey")
```
The boxplot is very useful, because it directly provides non-parametric summary statistics:
Min, Max, Median, Quartiles and therefore the inter-quartile range (IQR). The whiskers are usually the highest point that is within 1.5x the quartile plus the IQR. Everything beyond that is considered an outlier. Whiskers are however not always used in this way!
The mean and standard diviation are not visible in a boxplot, because it is only meaningful in distributions that center around the mean. It is however a part of summary statistics:
```
dat["BBBC001 manual count"].describe()
```
## Normal distribution
We assume that our distribution is "normal".
First we fit a normal distribution to our data.
```
(mu, sigma) = norm.fit(man_count)
n, bins, patches = plt.hist(man_count, 100,density=1)
# add a 'best fit' line
y = norm.pdf(bins, mu, sigma)
l = plt.plot(bins, y, 'r--', linewidth=2)
#plot
plt.xlabel('manual counts counts')
plt.ylabel('binned counts')
plt.title(r'$\mathrm{Histogram\ of\ manual\ counts:}\ \mu=%.3f,\ \sigma=%.3f$' %(mu, sigma))
plt.show()
```
Is it really normally distributed? What we see here is already one of the most problematic properties of a normal distribution: The susceptibility to outliers.
In normal distributions the confidence interval is determined by the standard diviation. A confidence level of 95% equals 1.96 x sigma.
```
#plot
(mu, sigma) = norm.fit(man_count)
n, bins, patches = plt.hist(man_count, 100,density=1)
# add a 'best fit' line
y = norm.pdf(bins, mu, sigma)
l = plt.plot(bins, y, 'r--', linewidth=2)
plt.xlabel('manual counts counts')
plt.ylabel('binned counts')
plt.title(r'$\mathrm{Histogram\ of\ manual\ counts:}\ \mu=%.3f,\ \sigma=%.3f$' %(mu, sigma))
plt.axvspan((mu-1.96*sigma),(mu+1.96*sigma), alpha=0.2, color='yellow')
plt.axvline(mu, color="black")
plt.show()
```
This shows even nicer that our outlier messes up the distribution :-)
How can we solve this in practise?
1. Ignore the problem and continue with the knowledge that we are overestimating the width of the distribution and underestimating the mean.
2. Censor the outlier.
3. Decide that we cannot assume normality and move to either a different distribution or non-parametric statistics.
## Other distributions
Of course there are many more distributions, e.g.
Lognormal is a distribution that becomes normal, when log transformed. It is important for the "geometric mean".
Bimodal distributions may arise from imaging data with background signal, or DNA methylation data.
Negative binomial distributions are very important in genomics, especially RNA-Seq analysis.
## Exercise
1. We had imported the total table with also the automated counts. Visualise the distribution of the data next to each other
2. Generate the summary statistics and compare the different distributions
3. Two weeks ago you learned how to analyze a folder of images and measured the average size of beads:
https://nbviewer.jupyter.org/github/BiAPoL/Bio-image_Analysis_with_Python/blob/main/image_processing/12_process_folders.ipynb
Go back to the bead-analysis two weeks ago and measure the intensity of the individual beads (do not average over the image). Plot the beads' intensities as different plots. Which one do you find most approproiate for these data?
| github_jupyter |
```
from google.colab import drive
drive.mount('/content/drive')
import torch
# If there's a GPU available...
if torch.cuda.is_available():
# Tell PyTorch to use the GPU.
device = torch.device("cuda")
print('There are %d GPU(s) available.' % torch.cuda.device_count())
print('We will use the GPU:', torch.cuda.get_device_name(0))
# If not...
else:
print('No GPU available, using the CPU instead.')
device = torch.device("cpu")
!ls drive/My\ Drive/Colab\ Notebooks/bert
BERT_PATH = "drive/My Drive/Colab Notebooks/bert"
COLUMN_NAME = "raw_tweet_features_text_tokens"
PATH = BERT_PATH+"/tweet_features_text_tokens_padded.csv.gz"
N_ROWS = 1000
CHUNKSIZE = 10
PAD = int(0)
MAX_LENGTH = 179
VOCAB = 'bert-base-multilingual-cased'
!pip install transformers
import pandas as pd
import numpy as np
from transformers import BertTokenizer
tokenizer = BertTokenizer.from_pretrained(VOCAB)
vocab_size = len(tokenizer.vocab.keys())
vocab_size
tokenizer.vocab['[PAD]']
```
## Load tweets from csv file as lists of int tokens
```
# Declaring two lambdas in order to cast a string to a numpy array of integers
f_to_int = lambda x: int(x)
f_int = lambda x: list(map(f_to_int, x.replace('[', '').replace(']', '').replace(' ', '').split(',')))
def read_tweets_list(path):
list_of_tweets = []
for chunk in pd.read_csv(path,
chunksize=CHUNKSIZE,
names=[COLUMN_NAME],
#dtype={COLUMN_NAME: pd.Int32Dtype()},
nrows=N_ROWS,
header=None,
index_col=0,
compression='gzip'):
#print(chunk)
tweets = chunk[COLUMN_NAME]
for t in tweets:
t_list = f_int(t)
list_of_tweets.append(t_list)
return list_of_tweets
tokenized_tweets = read_tweets_list(PATH)
n_tweets = len(tokenized_tweets)
n_tweets
```
## Check that at least one tweet having the last token different from PAD exists
```
for tweet in tokenized_tweets:
if tweet[-1] != PAD:
print('Ok')
```
## Decode all the tweets in the list
```
decoded_tweets = []
for t in tokenized_tweets:
decoded_tweets.append(tokenizer.decode(t))
len(decoded_tweets)
```
## Find two tweets containing the words "obama" and "president"
```
i = 0
for t in decoded_tweets:
if "president" in t or "obama" in t:
print(i)
i += 1
```
## Create the model from a pre-trained one
```
from transformers import BertModel, AdamW, BertConfig
# Load BERT model, the pretrained BERT model with a single
# linear classification layer on top.
model = BertModel.from_pretrained(
VOCAB, # Use the 12-layer BERT model, with an uncased vocab.
)
# Tell pytorch to run this model on the GPU.
model.cuda()
# Get all of the model's parameters as a list of tuples.
params = list(model.named_parameters())
print('The BERT model has {:} different named parameters.\n'.format(len(params)))
print('==== Embedding Layer ====\n')
for p in params[0:5]:
print("{:<55} {:>12}".format(p[0], str(tuple(p[1].size()))))
print('\n==== First Transformer ====\n')
for p in params[5:21]:
print("{:<55} {:>12}".format(p[0], str(tuple(p[1].size()))))
print('\n==== Output Layer ====\n')
for p in params[-2:]:
print("{:<55} {:>12}".format(p[0], str(tuple(p[1].size()))))
```
## Create inputs tensor
```
decoded_tweets[754] # it contains the word "president"
decoded_tweets[813] # it contains the word "obama"
model.eval()
tweet1 = tokenized_tweets[754]
tweet2 = tokenized_tweets[813]
inputs
```
## Create attention mask
```
masks = []
masks.append([int(token_id > 0) for token_id in tweet1]) # mask PAD tokens (PAD == 0)
masks.append([int(token_id > 0) for token_id in tweet2])
masks = torch.tensor(masks)
masks.shape
masks
```
## Move tensors to GPU
```
inputs = inputs.to(device)
masks = masks.to(device)
```
## Get model outputs
```
outputs = model(input_ids=inputs, attention_mask=masks)
outputs[0][0].size()
outputs[0][1].size()
outputs[1][0].size()
outputs[1][1].size()
```
#### The output is a tuple with 2 elements:
* a list containing lists of lists: for each input token we obtain a vector with 768 elements ---> for each input we have a list of MAX_LENGTH lists (one for each token), each containing 768 values (refer to this link http://jalammar.github.io/a-visual-guide-to-using-bert-for-the-first-time/, in the "Flowing through DistillBERT" section)
* the second element is a list containing an embedding vector for each input
```
outputs
```
## Compute similarity between the two embedding vectors
```
emb1 = outputs[1][0]
emb2 = outputs[1][1]
torch.dot(emb1, emb2) # dot product
cosine = torch.nn.CosineSimilarity(dim=0)
cosine(emb1, emb2) # cosine similarity
```
## Similarity between other two tweets
```
decoded_tweets[2]
decoded_tweets[200]
tweet1 = tokenized_tweets[2]
tweet2 = tokenized_tweets[200]
inputs = torch.tensor([tweet1, tweet2])
masks = []
masks.append([int(token_id > 0) for token_id in tweet1]) # mask PAD tokens (PAD == 0)
masks.append([int(token_id > 0) for token_id in tweet2])
masks = torch.tensor(masks)
inputs = inputs.to(device)
masks = masks.to(device)
outputs = model(input_ids=inputs, attention_mask=masks)
emb1 = outputs[1][0]
emb2 = outputs[1][1]
torch.dot(emb1, emb2) # dot product
cosine = torch.nn.CosineSimilarity(dim=0)
cosine(emb1, emb2) # cosine similarity
```
| github_jupyter |
```
import os
os.environ['CUDA_VISIBLE_DEVICES'] = ''
from malaya_speech.train.model import best_rq, ctc
from malaya_speech.train.model.conformer.model import Model as ConformerModel
import malaya_speech
import tensorflow as tf
import numpy as np
import json
from glob import glob
import string
unique_vocab = [''] + list(
string.ascii_lowercase + string.digits
) + [' ']
len(unique_vocab)
X = tf.compat.v1.placeholder(tf.float32, [None, None], name = 'X_placeholder')
X_len = tf.compat.v1.placeholder(tf.int32, [None], name = 'X_len_placeholder')
training = True
class Encoder:
def __init__(self, config):
self.config = config
self.encoder = ConformerModel(**self.config)
def __call__(self, x, input_mask, training = True):
return self.encoder(x, training = training)
config_conformer = malaya_speech.config.conformer_large_encoder_config
config_conformer['subsampling']['type'] = 'none'
config_conformer['dropout'] = 0.0
encoder = Encoder(config_conformer)
cfg = best_rq.Best_RQConfig(dropout=0.0,
attention_dropout=0.0,
encoder_layerdrop=0.0,
dropout_input=0.0,
dropout_features=0.0)
model = best_rq.Model(cfg, encoder)
r = model(X, padding_mask = X_len, features_only = True, mask = False)
logits = tf.layers.dense(r['x'], len(unique_vocab) + 1)
seq_lens = tf.reduce_sum(
tf.cast(tf.logical_not(r['padding_mask']), tf.int32), axis = 1
)
logits = tf.transpose(logits, [1, 0, 2])
logits = tf.identity(logits, name = 'logits')
seq_lens = tf.identity(seq_lens, name = 'seq_lens')
sess = tf.Session()
sess.run(tf.global_variables_initializer())
var_list = tf.get_collection(tf.GraphKeys.GLOBAL_VARIABLES)
saver = tf.train.Saver(var_list = var_list)
saver.restore(sess, 'best-rq-conformer-large-ctc-char/model.ckpt-2000000')
saver = tf.train.Saver()
saver.save(sess, 'output-best-rq-conformer-large-ctc/model.ckpt')
strings = ','.join(
[
n.name
for n in tf.get_default_graph().as_graph_def().node
if ('Variable' in n.op
or 'gather' in n.op.lower()
or 'placeholder' in n.name
or 'logits' in n.name
or 'seq_lens' in n.name)
and 'adam' not in n.name
and 'global_step' not in n.name
and 'Assign' not in n.name
and 'ReadVariableOp' not in n.name
and 'Gather' not in n.name
]
)
strings.split(',')
def freeze_graph(model_dir, output_node_names):
if not tf.gfile.Exists(model_dir):
raise AssertionError(
"Export directory doesn't exists. Please specify an export "
'directory: %s' % model_dir
)
checkpoint = tf.train.get_checkpoint_state(model_dir)
input_checkpoint = checkpoint.model_checkpoint_path
absolute_model_dir = '/'.join(input_checkpoint.split('/')[:-1])
output_graph = absolute_model_dir + '/frozen_model.pb'
clear_devices = True
with tf.Session(graph = tf.Graph()) as sess:
saver = tf.train.import_meta_graph(
input_checkpoint + '.meta', clear_devices = clear_devices
)
saver.restore(sess, input_checkpoint)
output_graph_def = tf.graph_util.convert_variables_to_constants(
sess,
tf.get_default_graph().as_graph_def(),
output_node_names.split(','),
)
with tf.gfile.GFile(output_graph, 'wb') as f:
f.write(output_graph_def.SerializeToString())
print('%d ops in the final graph.' % len(output_graph_def.node))
freeze_graph('output-best-rq-conformer-large-ctc', strings)
def load_graph(frozen_graph_filename):
with tf.gfile.GFile(frozen_graph_filename, 'rb') as f:
graph_def = tf.GraphDef()
graph_def.ParseFromString(f.read())
with tf.Graph().as_default() as graph:
tf.import_graph_def(graph_def)
return graph
files = [
'speech/record/savewav_2020-11-26_22-36-06_294832.wav',
'speech/record/savewav_2020-11-26_22-40-56_929661.wav',
'speech/record/675.wav',
'speech/record/664.wav',
'speech/example-speaker/husein-zolkepli.wav',
'speech/example-speaker/mas-aisyah.wav',
'speech/example-speaker/khalil-nooh.wav',
'speech/example-speaker/shafiqah-idayu.wav',
'speech/khutbah/wadi-annuar.wav',
]
ys = [malaya_speech.load(f)[0] for f in files]
padded, lens = malaya_speech.padding.sequence_1d(ys, return_len = True)
g = load_graph('output-best-rq-conformer-large-ctc/frozen_model.pb')
input_nodes = [
'X_placeholder',
'X_len_placeholder',
]
output_nodes = [
'logits',
'seq_lens',
]
inputs = {n: g.get_tensor_by_name(f'import/{n}:0') for n in input_nodes}
outputs = {n: g.get_tensor_by_name(f'import/{n}:0') for n in output_nodes}
test_sess = tf.Session(graph = g)
r = test_sess.run(outputs['logits'], feed_dict = {inputs['X_placeholder']: padded,
inputs['X_len_placeholder']: lens})
from tensorflow.tools.graph_transforms import TransformGraph
transforms = ['add_default_attributes',
'remove_nodes(op=Identity, op=CheckNumerics, op=Dropout)',
'fold_batch_norms',
'fold_old_batch_norms',
'quantize_weights(fallback_min=-10, fallback_max=10)',
'strip_unused_nodes',
'sort_by_execution_order']
pb = 'output-best-rq-conformer-large-ctc/frozen_model.pb'
input_graph_def = tf.GraphDef()
with tf.gfile.FastGFile(pb, 'rb') as f:
input_graph_def.ParseFromString(f.read())
transformed_graph_def = TransformGraph(input_graph_def,
input_nodes,
output_nodes, transforms)
with tf.gfile.GFile(f'{pb}.quantized', 'wb') as f:
f.write(transformed_graph_def.SerializeToString())
g = load_graph('output-best-rq-conformer-large-ctc/frozen_model.pb.quantized')
!tar -czvf output-best-rq-conformer-large-ctc.tar.gz output-best-rq-conformer-large-ctc
b2_application_key_id = os.environ['b2_application_key_id']
b2_application_key = os.environ['b2_application_key']
from b2sdk.v1 import *
info = InMemoryAccountInfo()
b2_api = B2Api(info)
application_key_id = b2_application_key_id
application_key = b2_application_key
b2_api.authorize_account("production", application_key_id, application_key)
file_info = {'how': 'good-file'}
b2_bucket = b2_api.get_bucket_by_name('malaya-speech-model')
key = 'output-best-rq-conformer-large-ctc.tar.gz'
outPutname = "pretrained/output-best-rq-conformer-large-ctc.tar.gz"
b2_bucket.upload_local_file(
local_file=key,
file_name=outPutname,
file_infos=file_info,
)
file = 'output-best-rq-conformer-large-ctc/frozen_model.pb'
outPutname = 'speech-to-text-ctc-v2/best-rq-conformer-large/model.pb'
b2_bucket.upload_local_file(
local_file=file,
file_name=outPutname,
file_infos=file_info,
)
file = 'output-best-rq-conformer-large-ctc/frozen_model.pb.quantized'
outPutname = 'speech-to-text-ctc-v2/best-rq-conformer-large-quantized/model.pb'
b2_bucket.upload_local_file(
local_file=file,
file_name=outPutname,
file_infos=file_info,
)
!rm -rf output-best-rq-conformer-large-ctc*
```
| github_jupyter |
# Qiskit: Open-Source Quantum Development, an introduction
---
### Workshop contents
1. Intro IBM Quantum Lab and Qiskit modules
2. Circuits, backends, visualization
3. Quantum info, circuit lib, algorithms
4. Circuit compilation, pulse, opflow
## 1. Intro IBM Quantum Lab and Qiskit modules
### https://quantum-computing.ibm.com/lab
### https://qiskit.org/documentation/
### https://github.com/qiskit
## 2. Circuits, backends and visualization
```
from qiskit import IBMQ
# Loading your IBM Quantum account(s)
#provider = IBMQ.load_account()
#provider = IBMQ.enable_account(<token>)
IBMQ.providers()
```
### Your first quantum circuit
Let's begin exploring the different tools in Qiskit Terra. For that, we will now create a Quantum Circuit.
```
from qiskit import QuantumCircuit
# Create circuit
# <INSERT CODE>
# print circuit
# <INSERT CODE>
```
Now let's run the circuit in the Aer simulator and plot the results in a histogram.
```
from qiskit import Aer
# run circuit on Aer simulator
# <INSERT CODE>
from qiskit.visualization import plot_histogram
# and display it on a histogram
# <INSERT CODE>
```
### Qiskit Visualization tools
```
from qiskit.visualization import plot_state_city
from qiskit.visualization import plot_state_paulivec, plot_state_hinton
circuit = QuantumCircuit(2, 2)
circuit.h(0)
circuit.cx(0, 1)
backend = Aer.get_backend('statevector_simulator') # the device to run on
result = backend.run(circuit).result()
psi = result.get_statevector(circuit)
# plot state city
# <INSERT CODE>
# plot state hinton
# <INSERT CODE>
# plot state paulivec
# <INSERT CODE>
```
#### Circuit Visualization
```
from qiskit import QuantumCircuit, ClassicalRegister, QuantumRegister
# Build a quantum circuit
circuit = QuantumCircuit(3, 3)
circuit.x(1)
circuit.h(range(3))
circuit.cx(0, 1)
circuit.measure(range(3), range(3));
# print circuit
# <INSERT CODE>
# print circuit using draw method
# <INSERT CODE>
```
There are different drawing formats. The parameter output (str) selects the output method to use for drawing the circuit. Valid choices are ``text, mpl, latex, latex_source``. See [qiskit.circuit.QuantumCircuit.draw](https://qiskit.org/documentation/stubs/qiskit.circuit.QuantumCircuit.draw.html?highlight=draw)
```
# print circuit using different drawer (mlp for example)
# <INSERT CODE>
```
##### Disable Plot Barriers and Reversing Bit Order
```
# Draw a new circuit with barriers and more registers
q_a = QuantumRegister(3, name='qa')
q_b = QuantumRegister(5, name='qb')
c_a = ClassicalRegister(3)
c_b = ClassicalRegister(5)
circuit = QuantumCircuit(q_a, q_b, c_a, c_b)
circuit.x(q_a[1])
circuit.x(q_b[1])
circuit.x(q_b[2])
circuit.x(q_b[4])
circuit.barrier()
circuit.h(q_a)
circuit.barrier(q_a)
circuit.h(q_b)
circuit.cswap(q_b[0], q_b[1], q_b[2])
circuit.cswap(q_b[2], q_b[3], q_b[4])
circuit.cswap(q_b[3], q_b[4], q_b[0])
circuit.barrier(q_b)
circuit.measure(q_a, c_a)
circuit.measure(q_b, c_b);
# Draw the circuit
# <INSERT CODE>
# Draw the circuit with reversed bit order
# <INSERT CODE>
# Draw the circuit without barriers
# <INSERT CODE>
```
##### MPL specific costumizations
```
# Change the background color in mpl
# <INSERT CODE>
# Scale the mpl output to 1/2 the normal size
# <INSERT CODE>
```
### Simulators
```
import numpy as np
# Import Qiskit
from qiskit import QuantumCircuit
from qiskit import Aer, transpile
from qiskit.tools.visualization import plot_histogram, plot_state_city
import qiskit.quantum_info as qi
Aer.backends()
simulator = Aer.get_backend('aer_simulator')
# Create circuit
circ = QuantumCircuit(2)
circ.h(0)
circ.cx(0, 1)
circ.measure_all()
# Transpile for simulator
simulator = Aer.get_backend('aer_simulator')
circ = transpile(circ, simulator)
# Run and get counts
result = simulator.run(circ).result()
counts = result.get_counts(circ)
plot_histogram(counts, title='Bell-State counts')
# Run and get memory (measurement outcomes for each individual shot)
result = simulator.run(circ, shots=10, memory=True).result()
memory = result.get_memory(circ)
print(memory)
```
##### Simulation methods
```
# Increase shots to reduce sampling variance
shots = 10000
# Stabilizer simulation method
sim_stabilizer = Aer.get_backend('aer_simulator_stabilizer')
job_stabilizer = sim_stabilizer.run(circ, shots=shots)
counts_stabilizer = job_stabilizer.result().get_counts(0)
# Statevector simulation method
sim_statevector = Aer.get_backend('aer_simulator_statevector')
job_statevector = sim_statevector.run(circ, shots=shots)
counts_statevector = job_statevector.result().get_counts(0)
# Density Matrix simulation method
sim_density = Aer.get_backend('aer_simulator_density_matrix')
job_density = sim_density.run(circ, shots=shots)
counts_density = job_density.result().get_counts(0)
# Matrix Product State simulation method
sim_mps = Aer.get_backend('aer_simulator_matrix_product_state')
job_mps = sim_mps.run(circ, shots=shots)
counts_mps = job_mps.result().get_counts(0)
plot_histogram([counts_stabilizer, counts_statevector, counts_density, counts_mps],
title='Counts for different simulation methods',
legend=['stabilizer', 'statevector',
'density_matrix', 'matrix_product_state'])
```
##### Simulation precision
```
# Configure a single-precision statevector simulator backend
simulator = Aer.get_backend('aer_simulator_statevector')
simulator.set_options(precision='single')
# Run and get counts
result = simulator.run(circ).result()
counts = result.get_counts(circ)
print(counts)
```
##### Device backend noise model simulations
```
from qiskit import IBMQ, transpile
from qiskit import QuantumCircuit
from qiskit.providers.aer import AerSimulator
from qiskit.tools.visualization import plot_histogram
from qiskit.test.mock import FakeVigo
device_backend = FakeVigo()
# Construct quantum circuit
circ = QuantumCircuit(3, 3)
circ.h(0)
circ.cx(0, 1)
circ.cx(1, 2)
circ.measure([0, 1, 2], [0, 1, 2])
# Create ideal simulator and run
# <INSERT CODE>
# Create simulator from backend
# <INSERT CODE>
# Transpile the circuit for the noisy basis gates and get results
# <INSERT CODE>
```
#### Usefull operations with circuits
```
qc = QuantumCircuit(12)
for idx in range(5):
qc.h(idx)
qc.cx(idx, idx+5)
qc.cx(1, 7)
qc.x(8)
qc.cx(1, 9)
qc.x(7)
qc.cx(1, 11)
qc.swap(6, 11)
qc.swap(6, 9)
qc.swap(6, 10)
qc.x(6)
qc.draw()
# width of circuit
# <INSERT CODE>
# number of qubits
# <INSERT CODE>
# count of operations
# <INSERT CODE>
# size of circuit
# <INSERT CODE>
# depth of circuit
# <INSERT CODE>
```
#### Final statevector
```
# Saving the final statevector
# Construct quantum circuit without measure
from qiskit.visualization import array_to_latex
circuit = QuantumCircuit(2)
circuit.h(0)
circuit.cx(0, 1)
# save statevector, run circuit and get results
# <INSERT CODE>
# Saving the circuit unitary
# Construct quantum circuit without measure
circuit = QuantumCircuit(2)
circuit.h(0)
circuit.cx(0, 1)
# save unitary, run circuit and get results
# <INSERT CODE>
```
Saving multiple statevectors
```
# Saving multiple states
# Construct quantum circuit without measure
steps = 5
circ = QuantumCircuit(1)
for i in range(steps):
circ.save_statevector(label=f'psi_{i}')
circ.rx(i * np.pi / steps, 0)
circ.save_statevector(label=f'psi_{steps}')
# Transpile for simulator
simulator = Aer.get_backend('aer_simulator')
circ = transpile(circ, simulator)
# Run and get saved data
result = simulator.run(circ).result()
data = result.data(0)
data
```
Saving custom statevector
```
# Generate a random statevector
num_qubits = 2
psi = qi.random_statevector(2 ** num_qubits, seed=100)
# Set initial state to generated statevector
circ = QuantumCircuit(num_qubits)
circ.set_statevector(psi)
circ.save_state()
# Transpile for simulator
simulator = Aer.get_backend('aer_simulator')
circ = transpile(circ, simulator)
# Run and get saved data
result = simulator.run(circ).result()
result.data(0)
```
### Parametric circuits
```
# Parameterized Quantum Circuits
from qiskit.circuit import Parameter
# create parameter and use it in circuit
# <INSERT CODE>
res = sim.run(circuit, parameter_binds=[{theta: [np.pi/2, np.pi, 0]}]).result() # Different bindings
res.get_counts()
from qiskit.circuit import Parameter
theta = Parameter('θ')
n = 5
qc = QuantumCircuit(5, 1)
qc.h(0)
for i in range(n-1):
qc.cx(i, i+1)
qc.barrier()
qc.rz(theta, range(5))
qc.barrier()
for i in reversed(range(n-1)):
qc.cx(i, i+1)
qc.h(0)
qc.measure(0, 0)
qc.draw('mpl')
#We can inspect the circuit’s parameters
# <INSERT CODE>
import numpy as np
theta_range = np.linspace(0, 2 * np.pi, 128)
circuits = [qc.bind_parameters({theta: theta_val})
for theta_val in theta_range]
circuits[-1].draw()
backend = Aer.get_backend('aer_simulator')
job = backend.run(transpile(circuits, backend))
counts = job.result().get_counts()
import matplotlib.pyplot as plt
fig = plt.figure(figsize=(8,6))
ax = fig.add_subplot(111)
ax.plot(theta_range, list(map(lambda c: c.get('0', 0), counts)), '.-', label='0')
ax.plot(theta_range, list(map(lambda c: c.get('1', 0), counts)), '.-', label='1')
ax.set_xticks([i * np.pi / 2 for i in range(5)])
ax.set_xticklabels(['0', r'$\frac{\pi}{2}$', r'$\pi$', r'$\frac{3\pi}{2}$', r'$2\pi$'], fontsize=14)
ax.set_xlabel('θ', fontsize=14)
ax.set_ylabel('Counts', fontsize=14)
ax.legend(fontsize=14)
# Random Circuit
from qiskit.circuit.random import random_circuit
# create random circuit
# <INSERT CODE>
# add unitary matrix to circuit
matrix = [[0, 0, 0, 1],
[0, 0, 1, 0],
[1, 0, 0, 0],
[0, 1, 0, 0]]
# <INSERT CODE>
# Classical logic
from qiskit.circuit import classical_function, Int1
@classical_function
def oracle(x: Int1, y: Int1, z: Int1) -> Int1:
return not x and (y or z)
circuit = QuantumCircuit(4)
circuit.append(oracle, [0, 1, 2, 3])
circuit.draw()
# circuit.decompose().draw() #synthesis
# Classical logic
from qiskit.circuit import classical_function, Int1
@classical_function
def oracle(x: Int1) -> Int1:
return not x
circuit = QuantumCircuit(2)
circuit.append(oracle, [0, 1])
circuit.draw()
circuit.decompose().draw()
```
https://qiskit.org/documentation/tutorials/circuits_advanced/02_operators_overview.html
## 3. Quantum info, circuit lib and algorithms
### Circuit lib
```
from qiskit.circuit.library import InnerProduct, QuantumVolume, clifford_6_2, C3XGate
# inner product circuit
# <INSERT CODE>
# clifford
# <INSERT CODE>
```
### Quantum info
```
from qiskit.quantum_info.operators import Operator
# Create an operator
# <INSERT CODE>
# add operator to circuit
# <INSERT CODE>
# Pauli
from qiskit.quantum_info.operators import Pauli
# use Pauli operator
# <INSERT CODE>
# Pauli with phase
from qiskit.quantum_info.operators import Pauli
circuit = QuantumCircuit(4)
iIXYZ = Pauli('iIXYZ') # ['', '-i', '-', 'i']
circuit.append(iIXYZ, [0, 1, 2, 3])
circuit.draw()
# create clifford
from qiskit.quantum_info import random_clifford
# random clifford
# <INSERT CODE>
# stabilizer and destabilizer
# <INSERT CODE>
```
### Algorithms
#### VQE
```
from qiskit.algorithms import VQE
from qiskit.algorithms.optimizers import SLSQP
from qiskit.circuit.library import TwoLocal
num_qubits = 2
ansatz = TwoLocal(num_qubits, 'ry', 'cz')
opt = SLSQP(maxiter=1000)
vqe = VQE(ansatz, optimizer=opt)
from qiskit.opflow import X, Z, I
H2_op = (-1.052373245772859 * I ^ I) + \
(0.39793742484318045 * I ^ Z) + \
(-0.39793742484318045 * Z ^ I) + \
(-0.01128010425623538 * Z ^ Z) + \
(0.18093119978423156 * X ^ X)
from qiskit.utils import algorithm_globals
seed = 50
algorithm_globals.random_seed = seed
qi = QuantumInstance(Aer.get_backend('statevector_simulator'), seed_transpiler=seed, seed_simulator=seed)
ansatz = TwoLocal(rotation_blocks='ry', entanglement_blocks='cz')
slsqp = SLSQP(maxiter=1000)
vqe = VQE(ansatz, optimizer=slsqp, quantum_instance=qi)
result = vqe.compute_minimum_eigenvalue(H2_op)
print(result)
```
#### Grover's algorithm
```
from qiskit.algorithms import Grover
from qiskit.algorithms import AmplificationProblem
# the state we desire to find is '11'
good_state = ['11']
# specify the oracle that marks the state '11' as a good solution
oracle = QuantumCircuit(2)
oracle.cz(0, 1)
# define Grover's algorithm
problem = AmplificationProblem(oracle, is_good_state=good_state)
# now we can have a look at the Grover operator that is used in running the algorithm
problem.grover_operator.draw(output='mpl')
from qiskit import Aer
from qiskit.utils import QuantumInstance
from qiskit.algorithms import Grover
aer_simulator = Aer.get_backend('aer_simulator')
grover = Grover(quantum_instance=aer_simulator)
result = grover.amplify(problem)
print('Result type:', type(result))
print('Success!' if result.oracle_evaluation else 'Failure!')
print('Top measurement:', result.top_measurement)
```
## 4. Transpiling, pulse and opflow
### Compiling circuits
```
[(b.name(), b.configuration().n_qubits) for b in provider.backends()]
from qiskit.tools.jupyter import *
%qiskit_backend_overview
from qiskit.providers.ibmq import least_busy
# get least busy backend
# <INSERT CODE>
# bell state
circuit = QuantumCircuit(2)
circuit.h(0)
circuit.cx(0, 1)
circuit.measure_all()
circuit.draw()
# run it in simulator
sim = Aer.get_backend('aer_simulator')
result = sim.run(circuit).result()
counts = result.get_counts()
plot_histogram(counts)
# run on least busy backend
# <INSERT CODE>
# get results
# <INSERT CODE>
circuit.draw('mpl')
from qiskit import transpile
# transpile with specified backend
# <INSERT CODE>
# rerun job
# <INSERT CODE>
# job status
# <INSERT CODE>
# get results
# <INSERT CODE>
from qiskit.visualization import plot_circuit_layout, plot_gate_map
display(transpiled_circuit.draw(idle_wires=False))
display(plot_gate_map(backend))
plot_circuit_layout(transpiled_circuit, backend)
# a slightly more interesting example:
circuit = QuantumCircuit(3)
circuit.h([0,1,2])
circuit.ccx(0, 1, 2)
circuit.h([0,1,2])
circuit.ccx(2, 0, 1)
circuit.h([0,1,2])
circuit.measure_all()
circuit.draw()
transpiled = transpile(circuit, backend)
transpiled.draw(idle_wires=False, fold=-1)
# Initial layout
# transpiling with initial layout
# <INSERT CODE>
transpiled.draw(idle_wires=False, fold=-1)
level0 = transpile(circuit, backend, optimization_level=0)
level1 = transpile(circuit, backend, optimization_level=1)
level2 = transpile(circuit, backend, optimization_level=2)
level3 = transpile(circuit, backend, optimization_level=3)
for level in [level0, level1, level2, level3]:
print(level.count_ops()['cx'], level.depth())
# transpiling is a stochastic process
transpiled = transpile(circuit, backend, optimization_level=2, seed_transpiler=42)
transpiled.depth()
transpiled = transpile(circuit, backend, optimization_level=2, seed_transpiler=1)
transpiled.depth()
# Playing with other transpiler options (without a backend)
transpiled = transpile(circuit)
transpiled.draw(fold=-1)
# Set a basis gates
backend.configuration().basis_gates
# specify basis gates
# <INSERT CODE>
# Set a coupling map
backend.configuration().coupling_map
from qiskit.transpiler import CouplingMap
# specify coupling map
# <INSERT CODE>
# Set an initial layout in a coupling map
transpiled = transpile(circuit,
coupling_map=CouplingMap([(0,1),(1,2)]),
initial_layout=[1, 0, 2])
transpiled.draw(fold=-1)
# Set an initial_layout in the coupling map with basis gates
transpiled = transpile(circuit,
coupling_map=CouplingMap([(0,1),(1,2)]),
initial_layout=[1, 0, 2],
basis_gates=['x', 'cx', 'h', 'p']
)
transpiled.draw(fold=-1)
transpiled.count_ops()['cx']
# Plus optimization level
transpiled = transpile(circuit,
coupling_map=CouplingMap([(0,1),(1,2)]),
initial_layout=[1, 0, 2],
basis_gates=['x', 'cx', 'h', 'p'],
optimization_level=3
)
transpiled.draw(fold=-1)
transpiled.count_ops()['cx']
# Last parameter, approximation degree
transpiled = transpile(circuit,
coupling_map=CouplingMap([(0,1),(1,2)]),
initial_layout=[1, 0, 2],
basis_gates=['x', 'cx', 'h', 'p'],
approximation_degree=0.99,
optimization_level=3
)
transpiled.draw(fold=-1)
transpiled.depth()
transpiled = transpile(circuit,
coupling_map=CouplingMap([(0,1),(1,2)]),
initial_layout=[1, 0, 2],
basis_gates=['x', 'cx', 'h', 'p'],
approximation_degree=0.01,
optimization_level=3
)
transpiled.draw(fold=-1)
transpiled.depth()
```
#### Qiskit is hardware agnostic!
```
# !pip install qiskit-ionq
# from qiskit_ionq import IonQProvider
# provider = IonQProvider(<your token>)
# circuit = QuantumCircuit(2)
# circuit.h(0)
# circuit.cx(0, 1)
# circuit.measure_all()
# circuit.draw()
# backend = provider.get_backend("ionq_qpu")
# job = backend.run(circuit)
# plot_histogram(job.get_counts())
```
### Pulse
```
from qiskit import pulse
# create dummy pusle program
# <INSERT CODE>
from qiskit.pulse import DriveChannel
channel = DriveChannel(0)
from qiskit.test.mock import FakeValencia
# build backend aware pulse schedule
# <INSERT CODE>
```
#### Delay instruction
```
# delay instruction
# <INSERT CODE>
```
#### Play instruction
#### Parametric pulses
```
from qiskit.pulse import library
# build parametric pulse
# <INSERT CODE>
# play parametric pulse
# <INSERT CODE>
# set frequency
# <INSERT CODE>
# shift phase
# <INSERT CODE>
from qiskit.pulse import Acquire, AcquireChannel, MemorySlot
# aqure instruction
# <INSERT CODE>
# example with left align schedule
# <INSERT CODE>
from qiskit import pulse
dc = pulse.DriveChannel
d0, d1, d2, d3, d4 = dc(0), dc(1), dc(2), dc(3), dc(4)
with pulse.build(name='pulse_programming_in') as pulse_prog:
pulse.play([1, 1, 1, 0, 0, 1, 1, 1, 0, 1, 1, 1, 0, 1, 0, 1, 0, 1, 1, 1, 0, 1, 1, 1], d0)
pulse.play([1, 0, 1, 0, 0, 0, 1, 0, 0, 1, 0, 0, 0, 1, 1, 0, 0, 0, 1, 0, 0, 0, 1, 0], d1)
pulse.play([1, 0, 1, 0, 0, 0, 1, 0, 0, 1, 1, 1, 0, 1, 0, 0, 0, 0, 1, 0, 0, 0, 1, 0], d2)
pulse.play([1, 0, 1, 0, 0, 0, 1, 0, 0, 0, 0, 1, 0, 1, 1, 0, 0, 0, 1, 0, 0, 0, 1, 0], d3)
pulse.play([1, 1, 1, 1, 0, 1, 1, 1, 0, 1, 1, 1, 0, 1, 0, 1, 0, 1, 1, 1, 0, 0, 1, 0], d4)
pulse_prog.draw()
```
### Opflow
```
from qiskit.opflow import I, X, Y, Z
print(I, X, Y, Z)
# These operators may also carry a coefficient.
# <INSERT CODE>
# These coefficients allow the operators to be used as terms in a sum.
# <INSERT CODE>
# Tensor products are denoted with a caret, like this.
# <INSERT CODE>
# Composition is denoted by the @ symbol.
# <INSERT CODE>
```
### State functions and measurements
```
from qiskit.opflow import (StateFn, Zero, One, Plus, Minus, H,
DictStateFn, VectorStateFn, CircuitStateFn, OperatorStateFn)
# zero, one
# <INSERT CODE>
# plus, minus
# <INSERT CODE>
# evaulation
# <INSERT CODE>
# adjoint
# <INSERT CODE>
# other way of writing adjoint
# <INSERT CODE>
```
#### Algebraic operations
```
import math
v_zero_one = (Zero + One) / math.sqrt(2)
print(v_zero_one)
print(StateFn({'0':1}))
print(StateFn({'0':1}) == Zero)
print(StateFn([0,1,1,0]))
from qiskit.circuit.library import RealAmplitudes
print(StateFn(RealAmplitudes(2)))
from qiskit.opflow import Zero, One, H, CX, I
# bell state
# <INSERT CODE>
# implement arbitrary circuits
# <INSERT CODE>
```
| github_jupyter |
# Pivotal method vs Percentile Method
In this notebook we will explore the difference between the **pivotal** and **percentile** bootstrapping methods.
tldr -
* The **percentile method** generates a bunch of re-samples and esimates confidence intervals based on the percentile values of those re-samples.
* The **pivotal method** is similar to percentile but does a correction for the fact that your input sample may not be a good representation of your population. Bootstrapped uses this as the default.
We will show that the pviotal method has generally better power. This does come at a cost - the pivotal method can warp the confidence interval to give non-sencical interval values.
See [Link1](https://ocw.mit.edu/courses/mathematics/18-05-introduction-to-probability-and-statistics-spring-2014/readings/MIT18_05S14_Reading24.pdf), [Link2](http://www.stat.cmu.edu/~cshalizi/402/lectures/08-bootstrap/lecture-08.pdf) for explanations of both methods.
```
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import numpy.random as npr
import bootstrapped.bootstrap as bs
import bootstrapped.stats_functions as bs_stats
import bootstrapped.power as bs_power
```
### Setup
The bootstrap is based on a sample of your larger population. A sample is only as good as how representitave it is. If you happen to be unlucky in your sample then you are going to make some very bad inferences!
We pick the exponential distribution because it should differentiate the difference between the two methods somewhat. We will also look at an extreme case.
```
population = np.random.exponential(scale=1000, size=500000)
# Plot the population
count, bins, ignored = plt.hist(population, 30, normed=True)
plt.title('Distribution of the population')
plt.show()
population = pd.Series(population)
# Do a bunch of simpulations and track the percent of the time the error bars overlap the true mean
def bootstrap_vs_pop_mean(population, num_samples, is_pivotal, num_loops=3000):
population_mean = population.mean()
pop_results = []
for _ in range(num_loops):
samples = population.sample(num_samples)
result = bs.bootstrap(samples.values, stat_func=bs_stats.mean, is_pivotal=is_pivotal)
# we want to 0 center this for our power plotting below
# we want our error bars to overlap zero
result = result - population_mean
pop_results.append(result)
return pop_results
def bounds_squared_distance(results):
'''The squared distance from zero for both the lower and the upper bound
This is a rough measure of how 'good' the confidence intervals are in terms of near misses vs extreme misses.
It is minimized when (1) the confidence interval is symmetric over zero and (2) when it is narrow.
'''
return np.sum([r.upper_bound**2 for r in results]) + np.sum([r.lower_bound**2 for r in results])
def squared_dist_ratio(x, y):
'Compare bounds_squared_distance for two sets of bootstrap results'
return bounds_squared_distance(x) / bounds_squared_distance(y)
```
### Pivotal vs Percentile for very small input sample size - 10 elements
```
pivotal_tiny_sample_count = bootstrap_vs_pop_mean(population, num_samples=10, is_pivotal=True)
percentile_tiny_sample_count = bootstrap_vs_pop_mean(population, num_samples=10, is_pivotal=False)
squared_dist_ratio(pivotal_tiny_sample_count, percentile_tiny_sample_count)
# more insignificant results is better
print(bs_power.power_stats(pivotal_tiny_sample_count))
bs_power.plot_power(pivotal_tiny_sample_count[::20])
# more insignificant results is better
print(bs_power.power_stats(percentile_tiny_sample_count))
bs_power.plot_power(percentile_tiny_sample_count[::20])
```
### Pivotal vs Percentile for small input sample size - 100 elements
```
pivotal_small_sample_count = bootstrap_vs_pop_mean(population, num_samples=100, is_pivotal=True)
percentile_small_sample_count = bootstrap_vs_pop_mean(population, num_samples=100, is_pivotal=False)
squared_dist_ratio(pivotal_small_sample_count, percentile_small_sample_count)
print(bs_power.power_stats(pivotal_small_sample_count))
bs_power.plot_power(pivotal_small_sample_count[::20])
print(bs_power.power_stats(percentile_small_sample_count))
bs_power.plot_power(percentile_small_sample_count[::20])
```
### Pivotal vs Percentile for medium input sample size - 1000 elements
```
pivotal_med_sample_count = bootstrap_vs_pop_mean(population, num_samples=1000, is_pivotal=True)
percentile_med_sample_count = bootstrap_vs_pop_mean(population, num_samples=1000, is_pivotal=False)
squared_dist_ratio(pivotal_med_sample_count, percentile_med_sample_count)
print(bs_power.power_stats(pivotal_med_sample_count))
bs_power.plot_power(pivotal_med_sample_count[::20])
print(bs_power.power_stats(percentile_med_sample_count))
bs_power.plot_power(percentile_med_sample_count[::20])
```
### Pivotal vs Percentile for somewhat large input sample size - 10k elements
### Bad Populaiton
```
bad_population = pd.Series([1]*10000 + [100000])
# Plot the population
count, bins, ignored = plt.hist(bad_population, 30, normed=True)
plt.title('Distribution of the population')
plt.show()
samples = bad_population.sample(10000)
print('Mean:\t\t{}'.format(bad_population.mean()))
print('Pivotal CI:\t{}'.format(bs.bootstrap(samples.values, stat_func=bs_stats.mean, is_pivotal=True)))
print('Percentile CI:\t{}'.format(bs.bootstrap(samples.values, stat_func=bs_stats.mean, is_pivotal=False)))
```
**Analysis:** The correction from the pivotal method causes a negative lower bound for the estimate in this instance. Generalization: the pivotal correction may not always produce realisitc intervals. In this situation we know that the mean cant be less than one. Still, we have seen from above that the piovtal method seems to be more reliable - this is why it is the default.
| github_jupyter |
Cesar Andrés Galindo Villalobos /
Juan Sebastian Correa Paez
**MOVIMIENTO DE UN CUERPO HACÍA UN PLANETA**
Un cuerpo de masa m2, parte desde una posición (a,b), con una velocidad horizontal Vx y una velocidad vertical Vy hacía un planeta de masa 1, radio R y con un centro de gravedad ubicado en el punto (h,k) del sistema de coordenadas rectangulares y a una distancia r. Establecer un modelo fisico matemático para definir si el cuerpo choca o no con el planeta.
**Elementos trigonométricos**
- Ángulo de desviación (alpha)
sen(alpha)=R/r entonces alpha=arcsen(R/r) **(1)**
donde R es constante, r es el vector posición inicial. Si r -> alpha, entonces alpha = 0
- Ángulo de inclinación de vector de la fuerza de gravitación universal
tan(tetha)=(b-k)/(a-h) -> tetha=arctan((b-k)/(a-h)) **(2)**
- Ángulo direccional del vector de la velocidad
tan(betha)=Vy/Vx -> betha=arctan(Vy/Vx) **(3)**
- Relación entre ángulos
tetha=alpha + betha -> alpha = tetha - alpha
alpha = arcsen(R/r) =arctan((b-k)/(a-h))=arctan(Vy/Vx) **(4)**
**Situaciones físicas**
Si alpha<=arcsen(R/r), el cuerpo hará contacto con el planeta
Si alpha>arcsen(R/r), el cuerpo no hará contacto con el planeta
**Modelo fisico matemático que relaciona el ánguo de desviación con las variables físicas: fuerza gravitacional, posición.**
- Vector posición en función de masas y fuerza gravitacional
F=(Gm1m2)/r^2 -> r^2=(G*m1*m2)/F -> r=((Gm1m2)/F)^(1/2) **(5)**
- Reemplazar r en ecuación (1)
alpha=arcsen(R/((Gm1m2)/F)^(1/2)) = arcsen=(R(F/(Gm1m2))^(1/2))
Si F -> 0, entonces alpha -> 0
- Modelo Físico matemático
alpha=arcsen(R*(F/(G*m1*m2))^(1/2)) = arctan((b-k)/(a-h))- arctan(Vy/Vx)
**FORMA DE SOLUCIÓN NO.1 SEGÚN ÁNGULO DE DESVIACIÓN**
```
import math
f = open ('tarea4-01.txt','r')
mensaje = f.read()
print("Los datos son:", mensaje)
G=float(6.67*(10**-11))
dato=str(mensaje).split(",")
h=float(dato[0])
a=float(dato[1])
k=float(dato[2])
b=float(dato[3])
m1=float(dato[4])
m2=float(dato[5])
Vx=float(dato[6])
r=float(dato[7])
R=float(dato[8])
Vy=float(dato[9])
F=(G*m1*m2)/(r**2)
alpharad=math.asin((R*math.sqrt(F))/(math.sqrt(G*m1*m2)))
alpha=math.degrees(alpharad)
print("El ángulo de desviación es", alpha)
condicion=math.asin(R/r)
print("La condición es:",condicion)
if alpha <= condicion:
print("Como el ángulo de desviación no es mayor a la condición, el cuerpo hará contacto con el planeta")
else:
print("Como el ángulo de desviación es mayor a la condición, el cuerpo no hará contacto con el planeta")
f.close()
```
**FORMA DE SOLUCIÓN NO.2 SEGÚN ÁNGULO DE DESVIACIÓN**
```
import math
f = open ('tarea4-01.txt','r')
mensaje = f.read()
print("Los datos son:", mensaje)
G=float(6.67*(10**-11))
dato=str(mensaje).split(",")
h=float(dato[0])
a=float(dato[1])
k=float(dato[2])
b=float(dato[3])
m1=float(dato[4])
m2=float(dato[5])
Vx=float(dato[6])
r=float(dato[7])
R=float(dato[8])
Vy=float(dato[9])
alpha=math.atan((b-k)/(a-h))-math.atan(Vy/Vx)
alpha=math.degrees(alpharad)
print("El ángulo de desviación es", alpha)
condicion=math.asin(R/r)
print("La condición es:",condicion)
if alpha <= condicion:
print("Como el ángulo de desviación no es mayor a la condición, el cuerpo hará contacto con el planeta")
else:
print("Como el ángulo de desviación es mayor a la condición, el cuerpo no hará contacto con el planeta")
f.close()
```
| github_jupyter |
```
import numpy as np # linear algebra
import pandas as pd # data processing
import warnings # ignore the warnings
warnings.filterwarnings('ignore')
```
**Reading the Datasets**
```
#ADA
ada=pd.read_csv("../input/crypto-data/ada_data.csv",index_col=0)
ada.rename(columns={"0":"Date", "1":"Price"},inplace = True)
ada.head()
#BTC
btc=pd.read_csv("../input/crypto-data/btc_data.csv",index_col=0)
btc.rename(columns={"0" : "Date","1":"Price"},inplace = True)
btc.head()
#ETH
eth=pd.read_csv("../input/crypto-data/btc_data.csv",index_col=0)
eth.rename(columns={"0" : "Date","1":"Price"},inplace = True)
eth.head()
```
## Data Analysis and Visualisation
```
import matplotlib.pyplot as plt
import seaborn as sns
```
**Adding a column crypto_name for all the 3 datasets separately**
```
for i in range(0,len(ada)):
ada['crypto_name']='ada'
for i in range(0,len(btc)):
btc['crypto_name']='btc'
for i in range(0,len(eth)):
eth['crypto_name']='eth'
```
**Viewing the updated datasets**
```
ada.head()
btc.head()
eth.head()
```
**Concatenating the dataframes to form a new dataframe**
```
new_df = pd.concat([ada,btc,eth])
```
**Viewing the new dataset**
```
new_df
```
**Save the file and then remove the previous index**
```
## Save file
new_df.to_csv('new_df.csv')
## Read the saved file
new_df = pd.read_csv('new_df.csv')
## Drop the column "Unnamed: 0" (previous index)
new_df.drop('Unnamed: 0',axis=1,inplace=True)
## View the dataset with proper index
new_df
```
**Crypto with highest value**
```
new_df[(new_df['Price']) == (new_df['Price'].max())]
```
**Crypto with lowest value**
```
new_df[(new_df['Price']) == (new_df['Price'].min())]
```
**Number of crypto prices present in each year**
```
pd.DatetimeIndex(new_df['Date']).year.value_counts()
```
**Different crypto entries in each year**
```
plt.figure(figsize=(15,4))
sns.countplot(x=pd.DatetimeIndex(new_df['Date']).year,hue='crypto_name',data=new_df)
plt.show()
```
**Creating a new dataframe for comparison on crypto_currencies**
```
## Merging the dataframes
df12 = ada.merge(btc, on='Date')
df13 = df12.merge(eth, on='Date')
## Removing the crypto names
df13.drop(['crypto_name_y','crypto_name','crypto_name_x'],axis=1,inplace=True)
## Renamming the columns
df13.rename(columns = {'Price_x':'ada','Price_y':'btc','Price':'eth'},inplace=True)
## Viewing the updated dataset
df13.head()
```
**Cumulative returns of the largest crypto-currency**
```
df13pct = df13[['ada','btc','eth']].pct_change()
df13_cum_returns = (df13pct + 1).cumprod() - 1
df13_cum_returns.head()
df13_cum_returns.plot(figsize=(15,5))
plt.title('Cumulative Returns of the largest crypto-currencies')
plt.show()
```
**Plotting date vs price for all the crypto-currencies**
```
plt.figure(figsize=(15,10))
plt.subplot(3, 1, 1 )
plt.title('ada')
sns.lineplot(x='Date', y='Price', data = ada, color= 'r')
plt.xticks(ticks=[0,27.5,55,82.5,110,137.5,165,192.5,220],rotation=45)
plt.subplot(3,1,2)
plt.title('btc')
sns.lineplot(x='Date', y='Price', data= btc, color= 'b')
plt.xticks(ticks=[0,48.75,97.5,146.5,195,247.5,300,345,390],rotation=45)
plt.subplot(3,1,3)
plt.title('eth')
sns.lineplot(x='Date', y='Price', data= eth, color= 'g')
plt.xticks(ticks=[0,48.75,97.5,146.5,195,247.5,300,345,390],rotation=45)
plt.tight_layout()
plt.savefig('individual_plots_data_vs_price')
```
### Predictions
The following set of code predicts the price of bitcoin, ada and etherium for the coming set of days
**Creating new dataframes with only Price columns**
```
ada1 = pd.read_csv('../input/cryptocurrencypricehistory/coin_Aave.csv')
btc1 = pd.read_csv('../input/cryptocurrencypricehistory/coin_Bitcoin.csv')
eth1 = pd.read_csv('../input/cryptocurrencypricehistory/coin_Ethereum.csv')
ada1 = ada1[['Date','Close']]
eth1 = eth1[['Date','Close']]
btc1 = btc1[['Date','Close']]
ada1['Date'] = pd.DatetimeIndex(ada1['Date']).date
eth1['Date'] = pd.DatetimeIndex(eth1['Date']).date
btc1['Date'] = pd.DatetimeIndex(btc1['Date']).date
ada1.rename(columns = {'Close':'Price'}, inplace = True)
eth1.rename(columns = {'Close':'Price'}, inplace = True)
btc1.rename(columns = {'Close':'Price'}, inplace = True)
new_btc = pd.DataFrame(btc1['Price'])
new_eth = pd.DataFrame(eth1['Price'])
new_ada = pd.DataFrame(ada1['Price'])
#A variable for predicting 'n' days into the future
prediction_days = 30
# another column for shifted n units up
new_ada['Prediction'] = new_ada['Price'].shift(-prediction_days)
new_btc['Prediction'] = new_btc['Price'].shift(-prediction_days)
new_eth['Prediction'] = new_eth['Price'].shift(-prediction_days)
# Create independent data set
# Convert the dataframe to a numpy array and drop prediction column
X1 = np.array(new_ada.drop(['Prediction'], 1))
X2 = np.array(new_eth.drop(['Prediction'], 1))
X3 = np.array(new_btc.drop(['Prediction'], 1))
# Remove the last n rows where n is the prediction_days
X1 = X1[:len(new_ada)-prediction_days]
X2 = X2[:len(new_eth)-prediction_days]
X3 = X3[:len(new_btc)-prediction_days]
# Create a depedent data set
# Convert the dataframe to a numpy array and drop prediction column
y1 = np.array(new_ada['Prediction'])
y1 = y1[: -prediction_days]
y2 = np.array(new_eth['Prediction'])
y2 = y2[: -prediction_days]
y3 = np.array(new_btc['Prediction'])
y3 = y3[: -prediction_days]
# Split data into training and testing data
from sklearn.model_selection import train_test_split
# for ada
X1_train, X1_test, y1_train, y1_test =train_test_split(X1,y1,test_size=0.2)
# for eth
X2_train, X2_test, y2_train, y2_test =train_test_split(X2,y2,test_size=0.2)
# for btc
X3_train, X3_test, y3_train, y3_test =train_test_split(X3,y3,test_size=0.2)
# Set the prediction days array equal to last 30 days from the original data
# for ada
prediction_days_array1 = (new_ada.drop(['Prediction'],1))[-prediction_days:]
# for eth
prediction_days_array2 = (new_eth.drop(['Prediction'],1))[-prediction_days:]
# btc
prediction_days_array3 = (new_btc.drop(['Prediction'],1))[-prediction_days:]
from sklearn.svm import SVR
# Create and train the Support Vector Machine (Regression) using radial basis function
svr_rbf = SVR(kernel='rbf', C=1e3, gamma=0.00001)
# for ada
svr_rbf.fit(X1_train, y1_train)
# for eth
svr_rbf.fit(X2_train, y2_train)
# btc
svr_rbf.fit(X3_train, y3_train)
# Test the model
# for ada
svr_rbf_confidence1 = svr_rbf.score(X1_test, y1_test)
print('svr_rbf accuracy for ada',svr_rbf_confidence1 )
# for eth
svr_rbf_confidence2 = svr_rbf.score(X2_test, y2_test)
print('svr_rbf accuracy for eth ',svr_rbf_confidence2 )
# for btc
svr_rbf_confidence3 = svr_rbf.score(X3_test, y3_test)
print('svr_rbf accuracy for ada',svr_rbf_confidence3 )
# Print the predicted values
# for ada
svm_prediction1 = svr_rbf.predict(X1_test)
print(svm_prediction1)
# Print the actual values
print(y1_test)
# for eth
svm_prediction2 = svr_rbf.predict(X2_test)
print(svm_prediction2)
# Print the actual values
print(y2_test)
# for ada
svm_prediction3 = svr_rbf.predict(X3_test)
print(svm_prediction3)
# Print the actual values
print(y3_test)
# print the model predictions for next 30 days
#for ada
svm_prediction1 = svr_rbf.predict(prediction_days_array1)
print('FOR ADA ',svm_prediction1)
#for eth
svm_prediction2 = svr_rbf.predict(prediction_days_array2)
print('FOR ETH ',svm_prediction2)
#for btc
svm_prediction3 = svr_rbf.predict(prediction_days_array3)
print('FOR BTC ',svm_prediction3[:30])
# Predictions for each crypto_currency in a tabular format for 30 days
new_predictions = pd.DataFrame([svm_prediction1,y1_test, svm_prediction2, y2_test, svm_prediction3[:30],y3_test], index=['ADA','ADA_ACTUAL','ETH','ETH_ACTUAL','BTC','BTC_ACTUAL'])
new_predictions = new_predictions.transpose()
new_predictions.head()
# Plotting the attained dataset
lisst = []
for i in range(1,31):
lisst.append(i)
plt.figure(figsize=(15,10))
plt.subplot(3,2,1)
sns.lineplot(x = lisst, y = new_predictions['ADA'][:30], marker = 'o',color='orange' )
plt.title("ADA Prediction Prices vs Days")
plt.xlabel("Days")
plt.ylabel("Predicted Price")
plt.subplot(3,2,2)
new_predictions['ADA_ACTUAL'][:30].plot(marker='o',color='blue')
plt.title("ADA Actual Prices vs Days")
plt.xlabel("Days")
plt.ylabel("Actual Price")
plt.subplot(3,2,3)
sns.lineplot(x = lisst, y = new_predictions['ETH'][:30], marker = 'o' )
plt.title("ETH Prediction Prices vs Days")
plt.xlabel("Days")
plt.ylabel("Predicted Price")
plt.subplot(3,2,4)
new_predictions['ETH_ACTUAL'][:30].plot(marker='o',color='red')
plt.title("ETH Actual Prices vs Days")
plt.xlabel("Days")
plt.ylabel("Actual Price")
plt.subplot(3,2,5)
sns.lineplot(x = lisst, y = new_predictions['BTC'][:30], marker = 'o',color='green' )
plt.title("BTC Prediction Prices vs Days")
plt.xlabel("Days")
plt.ylabel("Predicted Price")
plt.subplot(3,2,6)
new_predictions['BTC_ACTUAL'][:30].plot(marker='o',color='magenta')
plt.title("BTC Actual Prices vs Days")
plt.xlabel("Days")
plt.ylabel("Actual Price")
plt.tight_layout()
plt.show()
```
| github_jupyter |
```
import os
import pandas as pd
import numpy as np
import matplotlib as mpl
from matplotlib
import pyplot as plt
import datetime
import json
import seaborn as sns
#Setting Working Directory
os.getcwd()
os.chdir('/Users/Bhaktii/Downloads')
asia = pd.read_csv('ASIA_trending.csv')
eu = pd.read_csv('EU_trending.csv')
na = pd.read_csv('NA_trending.csv')
vd = pd.read_csv('videos.csv')
asia
eu
na
vd
#EDA
print(vd.shape)
print(vd.head())
vd.tail()
vd.info()
#Splitting publish time column within the videos dataset to time and date columns
vd['publish_time'].str.split( ' ', n = 1, expand = True)
#Frequency table for different categories in the videos dataset
CategoryCount = pd.crosstab(index = vd["category_id"], columns = "count")
CategoryCount
#Bar Graph for the above frquency table for categories and their respective counts
vd["category_id"].value_counts().plot(kind="bar", title= "Category Counts", xlabel="Categories", ylabel="No. of Videos", color="#8B3A62")
print ("Conclusion: From the table and the chart, it is evident that category ID 24 has the highest number of trending videos. As per our assumption, it is the Entertainment Category.")
#Merging the three datasets for different regions, that is, Asia, EU, NA into one consolidated dataset for ease of analysis
youtube = pd.concat(map(pd.read_csv, ['Asia_trending.csv','EU_trending.csv','NA_trending.csv']), ignore_index = True)
youtube
#EDA
print(youtube.shape)
#EDA
youtube.head()
#EDA
youtube.tail()
#EDA
youtube.info()
#Changing dataset column of likes/dislikes with split value columns for likes and dislikes
youtube [["likes","dislikes"]] = youtube ["likes/dislikes"].str.split("/", n = 1, expand = True)
youtube ['likes'] = pd.to_numeric(youtube['likes'])
youtube['dislikes'] = pd.to_numeric(youtube['dislikes'])
youtube
#Drop unnecessary columns w.r.t to the analysis
youtube = youtube.drop (columns = ['thumbnail_link','likes/dislikes'])
youtube.info()
#Change Datatypes as required
youtube['trending_date']= youtube['trending_date'].astype('category')
youtube['comments_disabled']= youtube['comments_disabled'].astype('category')
youtube['country']= youtube['country'].astype('category')
#EDA
youtube.describe()
#Formatting the given trending date in the dataset
youtube.trending_date = pd.to_datetime(youtube.trending_date, format='%y.%d.%m')
dates = youtube['trending_date'].dt
youtube["Year"] = dates.year
youtube["Month"] = dates.month
youtube["Day"] = dates.day
youtube['Weekday'] = dates.day_name()
#EDA
youtube.head()
#Table for Countries and Videos with Error or Videos removed
table1 = pd.crosstab(index=youtube["country"],columns="video_error_or_removed")
print(table1)
print("Conclusion: Germany and Canada noticably have the highest number of videos with an error or videos removed.")
#How many trending videos have their ratings disabled?
youtube["ratings_disabled"].value_counts().plot.pie(y='How many trending videos have their ratings disabled?', figsize=(5, 5), title='How many trending videos have their ratings disabled?', ylabel=' ', colors=['#003f5c', '#ffa600'])
print('Conclusion:Very few trending videos had their ratings disabled from the total cohort in the dataset. Since ratings play a significant role in the matter of whether a video goes viral or not and hence the lower fraction.')
youtube["ratings_disabled"].value_counts()
#How many trending videos have their comments disabled?
youtube["comments_disabled"].value_counts().plot.pie(y='How many trending videos have their comments disabled?', figsize=(5, 5), title='How many trending videos have their comments disabled?', ylabel=' ', colors=['#003f5c', '#ffa600'])
print("Conclusion: Very few trending videos have their comments disabled because, as in the case of ratings, comments too are instrumental in impacting the reach if a video.")
youtube["comments_disabled"].value_counts(normalize=True)
#How many trending videos have an error ir were removed?
youtube["video_error_or_removed"].value_counts().plot.pie(y='How many trending videos have an error or were removed?', figsize=(5, 5), title='How many trending videos have an error or were removed?', ylabel=' ', colors=['#003f5c', '#ffa600'])
print("Conclusion: A negligible fraction of the trending videos had an error or were removed.")
youtube["video_error_or_removed"].value_counts()
#Correlation calculations for the merged dataset of the given three regions
correlation=youtube.corr()
correlation
# Heatmap for representation of the given correlation
sns.heatmap(correlation)
plt.show()
print("Conclusion: It can be noticed that comment counts, views, and likes and dislikes show a moderate to high positive correlation.")
#How many trending video titles contain capitalized word?
def contains_capitalized_word(s):
for w in s.split():
if w.isupper():
return True
return False
vd["contains_cap"] = vd["title"].apply(contains_capitalized_word)
vd["contains_cap"].value_counts().plot.pie(y='Does the Title Contain a Capital Word', figsize=(5, 5), title='Does the Title Contain a Capital Word?', ylabel=' ', explode=[0,0.1])
print("Conclusion: Approximately 45% of titles contain a capitalized word which seems to confirm the human tendency to click on a video title with a capital word - the capitalization creates an impression of curiosity and excitement.")
#Bar Graph for No. of trending videos by country
youtube["country"].value_counts().plot(kind="bar", color= "#FF6A6A", title= "Trending Videos by Country")
print("Conclusion: With Japan having the lowest number of trending videos, followed by India. We observe that Germany, Canada, France and the US tie together with the highestnumer of trending videos.")
# Publishing year for the trending videos count
youtube['Year'].value_counts()
# Graph for Publishing year for the trending videos
youtube["Year"].value_counts().plot(kind="bar", color="#CAFF70", title= "Publishing Year of the Trending Videos", ylabel="No. of Videos")
# Histrogram for the Title Length of trending videos
vd["title_length"] = vd["title"].apply(lambda x: len(x))
plt.hist(vd['title_length'],bins=10,align='right', color='#EED2EE', edgecolor='black')
plt.xlabel('Title Length')
plt.ylabel('No. of Videos')
plt.title('Title Length of Trending Videos')
# Scatter Plot for views and title length within the merged 'youtube' datasset
views_sample=youtube['views'].sample(n=100000, random_state=1)
titlelength_sample=vd['title_length'].sample(n=100000, random_state=1)
sequence=np.arange(100000)
plt.scatter(views_sample,titlelength_sample,c=sequence)
plt.show()
print("Conclusion: It can be noticed that on a majority of the trending videos have an average length of 50-60 characters. Analyzing the title length in conjunction with the views shows that there is no apparent association between the two variables. However, we also notice that videos that have 100,000,000 views and more have title length between 20 and 80 characters approximately")
# Scatter Plot for views and likes within the merged datasset
plt.scatter(youtube['views'],youtube['likes'],c=np.arange(248252))
plt.show()
print("Conclusion: We see that views and likes are truly positively correlated: as one increases, the other increases too.")
#Line Graph for number of trending videos published on particular day of the week
youtube['Weekday'].value_counts()
publish_day=pd.DataFrame({'No. of Videos':[35151,36298,34908,34721,35314,36691,35169]},index=['Monday','Tuesday','Wednesday','Thursday','Friday','Saturday','Sunday'])
publish_day.plot.line(xlabel='Day of the Week', ylabel='No. of Videos')
print("On observation, Tuesdays and Saturdays were largely the best days chosen by content creators to upload videos while Wednesday adn Thursday were seen as the worst.")
#Which channels have the largest number of trending videos?
channels=vd['channel_title'].value_counts(ascending=False).head(20)
print(channels)
channels.plot(kind="barh",color='#00868B',edgecolor='blue', title= 'Highest number of trending videos')
print("Conclusion: Indian Channels - SET India, VikatanTV, and SAB TV - have the highest number of trending videos. Even in general, the top 20 channels are dominated by Indian channels.")
# Finding the most controversial videos
#Controversial videos could be defined here as the ikes and dislikes that have a 10% difference.
# Sorting based on highest number of views
youtube = youtube.sort_values(by=["views"],
ascending=False)
#Making a new column with the percentage difference between likes and dislikes
youtube["Controversial score"] = np.abs(youtube["likes"] - youtube["dislikes"])/(youtube["likes"]+youtube["dislikes"])
youtube1= youtube[(youtube["Controversial score"] < 0.1 ) ]
youtube1
print(youtube["Controversial score"].describe())
print(" ")
print("Conclusion: It is generally assumed that controversial videos tend to get viral and thus, I attempted to construct a controversial score for the trending videos. It can be seen here that this score ranges from 0 to 0.5.")
```
| github_jupyter |
```
import pandas as pd
import numpy as np
from funcoes_microdados import ler_microdados, ler_escolas_censo
pd.set_option('display.max_columns', 500)
#pd.set_option('display.max_rows', 500)
# Leitura dos dados: MICRODADOS_ENEM são os microdados do ENEM, ESCOLAS.csv é uma tabela do Censo Escolar do respectivo ano
ano = 2019
escolas = ler_escolas_censo(ano, f'./dados/{ano}/ESCOLAS_{ano}.csv')
dados = ler_microdados(ano, f'./dados/{ano}/MICRODADOS_ENEM_{ano}.csv')
# Função que retorna ranking dataframe com filtro de número mínimo de alunos da escola (padrão 10 alunos mínimos)
def medias_escolas(df, min_alunos = 10):
geral = df[df.PARTICIPANTES >= min_alunos] # Seleciona somente os alunos cuja escola tem número de participantes >= min_alunos
# Calcula as médias por escola mantendo outras variáveis de escola
geral = geral.groupby('CO_ESCOLA').agg(CH = ('NU_NOTA_CH', 'mean'),CN = ('NU_NOTA_CN', 'mean'), LC = ('NU_NOTA_LC', 'mean'), MT = ('NU_NOTA_MT', 'mean'),
MEDIAS_OBJETIVA = ('OBJETIVA', 'mean'), RED = ('NU_NOTA_REDACAO', 'mean'), OBJETIVAS_COM_RED = ('MEDIA_RD', 'mean'),
MUNICIPIO = ('NO_MUNICIPIO_ESC', 'first'),PARTICIPANTES = ('PARTICIPANTES', 'first'),
ESTADO = ('SG_UF_ESC', 'first')).reset_index().sort_values('MEDIAS_OBJETIVA', ascending = False)
# Insere o nome das escolas usando o Censo Escolar, se o codInep não constar no Censo, deixa o nome vazio
geral = geral.merge(escolas, on='CO_ESCOLA', how='left')
geral.fillna(value="", inplace=True)
# Arredonda as notas pra duas casas decimais, muda a ordem e renomeia as colunas
geral[['CN', 'CH', 'LC', 'MT', 'MEDIAS_OBJETIVA', 'RED', 'OBJETIVAS_COM_RED']] = geral[['CN', 'CH', 'LC', 'MT', 'MEDIAS_OBJETIVA', 'RED', 'OBJETIVAS_COM_RED']].round(2)
geral = geral[['CO_ESCOLA', 'NOME', 'MUNICIPIO','ESTADO', 'PARTICIPANTES', 'CH', 'CN', 'LC', 'MT', 'MEDIAS_OBJETIVA', 'RED', 'OBJETIVAS_COM_RED' ]]
geral.rename(columns= {"CO_ESCOLA": "codInep",
"NOME": "nome",
"MUNICIPIO": "municipio",
"ESTADO": "estado",
"PARTICIPANTES": "participantes",
"CH": "mediaCH","CN": "mediaCN","LC": "mediaLC","MT": "mediaMT","MEDIAS_OBJETIVA": "mediaObj","RED": "mediaRed","OBJETIVAS_COM_RED": "mediaGeral"}, inplace=True)
return geral
# Filtro: Notas objetivas não nulas, possui Cod Escola, concluindo o ensino médio. Adiciona colunas com médias objetiva e com redação
dados_filtrado = dados[(dados.NU_NOTA_CH > 0) &(dados.NU_NOTA_LC > 0) &(dados.NU_NOTA_MT > 0) &(dados.NU_NOTA_CN > 0) &(dados.CO_ESCOLA > 0) & (dados.TP_ST_CONCLUSAO == 2) & (dados.TP_ENSINO == 1)]
dados_filtrado = dados_filtrado.assign(OBJETIVA = (dados_filtrado.NU_NOTA_CH + dados_filtrado.NU_NOTA_CN + dados_filtrado.NU_NOTA_LC + dados_filtrado.NU_NOTA_MT)/4)
dados_filtrado.fillna({'NU_NOTA_REDACAO' : 0}, inplace=True)
dados_filtrado = dados_filtrado.assign(MEDIA_RD = (dados_filtrado.NU_NOTA_CH + dados_filtrado.NU_NOTA_CN + dados_filtrado.NU_NOTA_LC + dados_filtrado.NU_NOTA_MT + dados_filtrado.NU_NOTA_REDACAO)/5)
# Adiciona número de participantes da escola do aluno
dados_filtrado = dados_filtrado.merge(dados_filtrado.groupby('CO_ESCOLA').agg(PARTICIPANTES = ('NU_INSCRICAO', 'count')).reset_index())
df = medias_escolas(dados_filtrado, 0)
nomeJson = f'mediasEnem_{ano}.json'
df.to_json(nomeJson, orient='records')
# Salva num banco MongoDB local
#!mongoimport --db=apiNotasEnem --collection=2019 --type=json --file={nomeJson} --jsonArray --drop
# Criar index na variável `codInep` posteriormente para melhor performace
```
| github_jupyter |
```
import sys
sys.path.append('/Users/mic.fell/Documents/venvs/jupyter/lib/python3.6/site-packages')
import pandas as pd
import html
from functools import reduce
import re
import numpy as np
from nltk import word_tokenize
import spacy
import pronouncing
from textblob import TextBlob
from sklearn.preprocessing import StandardScaler
from keras.models import load_model
import _pickle
from sklearn import preprocessing
nlp = spacy.load('en_core_web_lg')
mpd_with_lyrics = pd.read_csv('/Users/mic.fell/Documents/Jupyter Orbit/resources/recsys_challenge_2018/mpd_wasabi_aligned_langdetect.csv', sep='\t', encoding='utf8')
mpd_with_lyrics = mpd_with_lyrics.drop(['Unnamed: 0'], axis=1)
mpd_with_lyrics.head()
features = read_file('/Users/mic.fell/Documents/Jupyter Orbit/resources/recsys_challenge_2018/mpd_ids_english_features.pickle')
features.head()
# use only english lyrics
mpd_with_lyrics = mpd_with_lyrics[mpd_with_lyrics['langdetect'] == 'en']
print('aligned indices :', len(mpd_with_lyrics))
print('unique Spotify URIs:', len(set(mpd_with_lyrics['spotify_track_uri'])))
print('unique lyrics :', len(set(mpd_with_lyrics['lyrics'])))
def read_file(name):
with open(name, 'rb') as f:
content = _pickle.load(f)
return content
def write_file(content, name):
with open(name, 'wb') as f:
_pickle.dump(content, f)
def robust_min_max_scaler(matrix):
matrix_robust = preprocessing.RobustScaler().fit_transform(matrix)
matrix_mmax = preprocessing.MinMaxScaler().fit_transform(matrix_robust)
return matrix_mmax
# parse lyrics to segment-line-structure, assuming lines are separated by line_border_indicator and
# segments are separated by multiple consecutive line_border_indicator occurences
# assuming line_border_indicator is <br> (standard in lyrics.wikia.com)
def tree_structure(text):
#normalize segment border encoding
segment_border_encoder = '<segmentborder>'
line_border_encoder = '<lineborder>'
tree_string = re.sub('(( )*<br>( )*){2,}', segment_border_encoder, text)
tree_string = re.sub('( )*<br>( )*', line_border_encoder, tree_string)
#parse tree_string
segment_structure = tree_string.split(segment_border_encoder)
tree_structure = list(map(lambda segment: segment.split(line_border_encoder), segment_structure))
return tree_structure
#flattened tree structure, does not differentiate between segment and line border
def line_structure(lyric_tree):
return reduce(lambda x, segment: x + segment, lyric_tree, [])
# flattened line_structure
def token_structure(lyric_tree, tokenizer=word_tokenize):
return reduce(lambda x, line: extend_with_return(x, tokenizer(line)), line_structure(lyric_tree), [])
def extend_with_return(some_list, other_list):
some_list.extend(other_list)
return some_list
# normalizations we want to apply to all lyrics go here
def normalize_lyric(lyric):
lyric = html.unescape(lyric)
lyric = lyric.lower()
return lyric
# Reduce a list of numbers to single number / feature (cf. np.average, np.std, ...)
def list_span(some_list):
min_list = min(some_list)
return max(some_list) / min_list if min_list > 0 else 1e-10
######################################
########Stylometric features##########
######################################
def type_token_ratio(lyric_tokens):
return len(set(lyric_tokens)) / len(lyric_tokens)
def line_lengths_in_chars(lyric_lines):
return list(map(len, lyric_lines))
def line_lengths_in_tokens(lyric_lines):
return list(map(lambda line: len(word_tokenize(line)), lyric_lines))
def pos_tag_distribution(lyric_lines):
# Look at https://spacy.io/api/annotation for a better description of each tag
tags = ['ADJ', 'ADP', 'ADV', 'AUX', 'CONJ', 'CCONJ', 'DET', 'INTJ', 'NOUN',
'NUM', 'PART', 'PRON', 'PROPN', 'PUNCT', 'SCONJ', 'SYM', 'VERB', 'X', 'SPACE']
freq = dict()
for tag in tags:
freq[tag] = 0
for line in lyric_lines:
doc = nlp(line)
for word in doc:
if word.pos_ in tags:
freq[word.pos_] += 1
wc = sum(line_lengths_in_tokens(lyric_lines))
for key in freq:
freq[key] /= wc
return freq
def get_rhymes(lyric_lines):
count = 0
lyric_length = len(lyric_lines)
for i in range(lyric_length-1):
words = lyric_lines[i].split()
if len(words) < 1:
continue
rhymes = pronouncing.rhymes(words[-1])
next_line_words = lyric_lines[i+1].split()
if next_line_words is not None and len(next_line_words) > 0 and next_line_words[-1] in rhymes:
count += 1
return count / lyric_length if lyric_length > 0 else 0
def get_echoisms(lyric_lines):
vowels = ['a', 'e', 'i', 'o', 'u']
# Do echoism count on a word level
echoism_count = 0
for line in lyric_lines:
doc = nlp(line)
for i in range(len(doc) - 1):
echoism_count += doc[i].text.lower() == doc[i+1].text.lower()
# Count echoisms inside words e.g. yeeeeeeah
for tk in doc:
for i in range(len(tk.text) - 1):
if tk.text[i] == tk.text[i+1] and tk.text in vowels:
echoism_count += 1
break
return echoism_count / sum(line_lengths_in_tokens(lyric_lines))
def is_title_in_lyrics(title, lyric_lines):
for line in lyric_lines:
if title in line:
return True
return False
def count_duplicate_lines(lyric_lines):
wc = sum(line_lengths_in_tokens(lyric_lines))
wc = wc if wc > 0 else 1
return sum([lyric_lines.count(x) for x in list(set(lyric_lines)) if lyric_lines.count(x) > 1]) / wc
##################################
########Segment features##########
##################################
#The indices of lines that end a segment
def segment_borders(lyric_tree):
segment_lengths = reduce(lambda x, block: x + [len(block)], lyric_tree, [])
segment_indices = []
running_sum = -1
for i in range(len(segment_lengths)):
running_sum += segment_lengths[i]
segment_indices.append(running_sum)
return segment_indices[:-1]
# lengths of the segments
def segment_lengths(lyric_tree):
return reduce(lambda x, block: x + [len(block)], lyric_tree, [])
##################################
########Orientation feature#######
##################################
def get_verb_tense_frequencies(lyric_lines):
freq = dict()
freq['present'] = 0
freq['future'] = 0
freq['past'] = 0
verbs_no = 0
for line in lyric_lines:
doc = nlp(line)
for i in range(len(doc)):
token = doc[i]
if token.pos_ == 'VERB' and token.tag_ != 'MD':
verbs_no += 1
if 'present' in spacy.explain(token.tag_):
freq['present'] += 1
elif 'past' in spacy.explain(token.tag_):
freq['past'] += 1
elif token.pos_ == 'VERB' and token.tag_ == 'MD' and token.text.lower() == 'will':
if i < len(doc) - 1:
i += 1
next_token = doc[i]
if next_token is not None and next_token.text == 'VB':
verbs_no += 1
freq['future'] += 1
if verbs_no > 0:
for key, value in freq.items():
freq[key] = value/verbs_no
return freq
def get_polarity_and_subjectivity(lyric_lines):
text = '\n'.join(lyric_lines)
opinion = TextBlob(text)
sentiment = opinion.sentiment
return (sentiment.polarity, sentiment.subjectivity)
##################################
########Emotion features##########
##################################
#import emoclassify as clf
#
#def get_emotion_vector(lyric, title, modelpath='emodetect.h5'):
# return clf.classify(0, '', title, lyric_content = html.unescape(lyric).replace('<br>', '\n'))
def representations_from(lyric):
"""Compute different representations of lyric: tree (with paragraphs), lines, tokens"""
lyric_tree = tree_structure(lyric)
lyric_lines = line_structure(lyric_tree)
lyric_tokens = token_structure(lyric_tree)
return lyric_tree, lyric_lines, lyric_tokens
def feat_vect_from(feature_list):
"""Assuming a list of features of the lyric"""
feat_vect = []
feat_vect.append(np.median(feature_list))
feat_vect.append(np.std(feature_list))
feat_vect.append(list_span(feature_list))
return feat_vect
def extend_feat_vect(feat_vect, feature_list):
feat_vect.extend(feat_vect_from(feature_list))
return feat_vect
def feature_vector_from(lyric, title):
lyric_tree, lyric_lines, lyric_tokens = representations_from(lyric)
# lump everything in a single feature vector
feat_vect = []
# segmentation features
feat_vect = extend_feat_vect(feat_vect, segment_lengths(lyric_tree))
# stylometric features
feat_vect.append(type_token_ratio(lyric_tokens))
ln_lengths_chars = line_lengths_in_chars(lyric_lines)
# line count
feat_vect.append(len(ln_lengths_chars))
feat_vect = extend_feat_vect(feat_vect, ln_lengths_chars)
feat_vect = extend_feat_vect(feat_vect, line_lengths_in_tokens(lyric_lines))
feat_vect.append(get_rhymes(lyric_lines))
# orientation features
feat_vect.extend(get_polarity_and_subjectivity(lyric_lines))
# emotion features
#emo = get_emotion_vector(lyric, title)
#feat_vect.extend(get_emotion_vector(lyric, title))
#features using expensive nlp(.) calls
#feat_vect.extend(pos_tag_distribution(lyric_lines).values())
#feat_vect.extend(get_verb_tense_frequencies(lyric_lines).values())
#feat_vect.append(get_echoisms(lyric_lines))
return feat_vect
def feature_vectors_from(many_lyrics: list, many_titles: list) -> np.ndarray:
many_count = len(many_lyrics)
first_feat_vect = feature_vector_from(many_lyrics.iloc[0], many_titles.iloc[0])
feat_vects = np.empty((many_count, len(first_feat_vect)), dtype=object)
feat_vects[0] = first_feat_vect
for i in range(1, many_count):
feat_vects[i] = feature_vector_from(many_lyrics.iloc[i], many_titles.iloc[i])
if i % 100 == 0:
print('Progress:', i, 'of', many_count, '(' + str(round(i/many_count*100, 1)) + '%)')
return feat_vects
# Define subset to compute features on
mpd_with_lyrics_subset = mpd_with_lyrics.iloc[:10]
all_uris = mpd_with_lyrics_subset['spotify_track_uri']
all_lyrics = mpd_with_lyrics_subset['lyrics']
all_titles = mpd_with_lyrics_subset['title']
assert len(all_uris) == len(all_lyrics) == len(all_titles)
matrix = feature_vectors_from(all_lyrics, all_titles)
print(matrix.shape)
#Put back into dataframe
fvec = {}
for row in range(matrix.shape[0]):
fvec[all_uris.iloc[row]] = matrix[row, :]
xf = pd.DataFrame(list(fvec.items()), columns=['spotify_track_uri', 'feature_vector'])
xf.head()
write_file(content=xf, fl='/Users/mic.fell/Documents/Jupyter Orbit/resources/recsys_challenge_2018/mpd_ids_english_features.csv')
xx = read_file('/Users/mic.fell/Documents/Jupyter Orbit/resources/recsys_challenge_2018/mpd_ids_english_features.pickle')
xx
xx.iloc[567].feature_vector.shape
from time import time
mpd_limited = mpd_with_lyrics_subset
all_lyrics, all_titles = (mpd_limited['lyrics'], mpd_limited['title'])
t = time()
matrix = feature_vectors_from(all_lyrics, all_titles)
print('Lyrics count :', len(all_lyrics))
time_taken = time()-t
print('Time taken [s] :', round(time_taken, 2))
print('Time / lyric [s] :', round(time_taken / len(all_lyrics), 3))
print('Time / 416k [h/m] :', round(time_taken / len(all_lyrics) * 416000 / 60 / 60, 1), '/', round(time_taken / len(all_lyrics) * 416000 / 60, 1))
t = time()
scaled_matrix = apply_to_columns(min_max_scaler, matrix)
print('Scaling', matrix.shape, ':', round(time() - t, 3))
```
| github_jupyter |
# An Informal Introduction to Python
[The [source material](https://docs.python.org/3.5/tutorial/introduction.html) is from Python 3.5.1, but the contents of this tutorial should apply to almost any version of Python 3]
Many of the examples in this manual, even those entered at the interactive prompt, include comments. Comments in Python start with the hash character, `#`, and extend to the end of the physical line. A comment may appear at the start of a line or following whitespace or code, but not within a string literal. A hash character within a string literal is just a hash character. Since comments are to clarify code and are not interpreted by Python, they may be omitted when typing in examples.
Some examples:
```
# This is the first comment
spam = 1 # and this is the second comment
# ... and now a third!
text = "# This is not a comment because it's inside quotes."
```
## Using Python as a Calculator
Letâs try some simple Python commands.
### Numbers
The interpreter acts as a simple calculator: you can type an expression at it and it will write the value. Expression syntax is straightforward: the operators `+`, `-`, `*` and `/` work just like in most other languages (for example, Pascal or C); parentheses (`()`) can be used for grouping. For example:
```
2 + 2
50 - 5*6
(50 - 5*6) / 4
8 / 5 # Division always returns a floating point number.
```
The integer numbers (e.g. `2`, `4`, `20`) have type [`int`](https://docs.python.org/3.5/library/functions.html#int), the ones with a fractional part (e.g. `5.0`, `1.6`) have type [`float`](https://docs.python.org/3.5/library/functions.html#float). We will see more about numeric types later in the tutorial.
Division (`/`) always returns a float. To do [floor division](https://docs.python.org/3.5/glossary.html#term-floor-division) and get an integer result (discarding any fractional result) you can use the `//` operator; to calculate the remainder you can use `%`:
```
17 / 3 # Classic division returns a float.
17 // 3 # Floor division discards the fractional part.
17 % 3 # The % operator returns the remainder of the division.
5 * 3 + 2 # result * divisor + remainder
```
With Python, it is possible to use the `**` operator to calculate powers:
```
5 ** 2 # 5 squared
2 ** 7 # 2 to the power of 7
```
Do note that `**` has higher precedence than `-`, so if you want a negative base you will need parentheses:
```
-3**2 # Same as -(3**2)
(-3)**2
```
The equal sign (`=`) is used to assign a value to a variable. Afterwards, no result is displayed before the next interactive prompt:
```
width = 20
height = 5 * 9
width * height
```
If a variable is not âdefinedâ (assigned a value), trying to use it will give you an error:
```
n # Try to access an undefined variable.
```
There is full support for floating point; operators with mixed type operands convert the integer operand to floating point:
```
3 * 3.75 / 1.5
7.0 / 2
```
In interactive mode, the last printed expression is assigned to the variable `_`. This means that when you are using Python as a desk calculator, it is somewhat easier to continue calculations, for example:
```
tax = 12.5 / 100
price = 100.50
price * tax
price + _
round(_, 2)
```
This variable should be treated as read-only by the user. Donât explicitly assign a value to it â you would create an independent local variable with the same name masking the built-in variable with its magic behavior.
In addition to `int` and `float`, Python supports other types of numbers, such as [`Decimal`](https://docs.python.org/3.5/library/decimal.html#decimal.Decimal) and [`Fraction`](https://docs.python.org/3.5/library/fractions.html#fractions.Fraction). Python also has built-in support for [complex numbers](https://docs.python.org/3.5/library/stdtypes.html#typesnumeric), and uses the `j` or `J` suffix to indicate the imaginary part (e.g. `3+5j`).
### Strings
Besides numbers, Python can also manipulate strings, which can be expressed in several ways. They can be enclosed in single quotes (`'...'`) or double quotes (`"..."`) with the same result. `\` can be used to escape quotes:
```
'spam eggs' # Single quotes.
'doesn\'t' # Use \' to escape the single quote...
"doesn't" # ...or use double quotes instead.
'"Yes," he said.'
"\"Yes,\" he said."
'"Isn\'t," she said.'
```
In the interactive interpreter, the output string is enclosed in quotes and special characters are escaped with backslashes. While this might sometimes look different from the input (the enclosing quotes could change), the two strings are equivalent. The string is enclosed in double quotes if the string contains a single quote and no double quotes, otherwise it is enclosed in single quotes. The [`print()`](https://docs.python.org/3.5/library/functions.html#print) function produces a more readable output, by omitting the enclosing quotes and by printing escaped and special characters:
```
'"Isn\'t," she said.'
print('"Isn\'t," she said.')
s = 'First line.\nSecond line.' # \n means newline.
s # Without print(), \n is included in the output.
print(s) # With print(), \n produces a new line.
```
If you donât want characters prefaced by `\` to be interpreted as special characters, you can use _raw strings_ by adding an `r` before the first quote:
```
print('C:\some\name') # Here \n means newline!
print(r'C:\some\name') # Note the r before the quote.
```
String literals can span multiple lines. One way is using triple-quotes: `"""..."""` or `'''...'''`. End of lines are automatically included in the string, but itâs possible to prevent this by adding a `\` at the end of the line. The following example:
```
print("""\
Usage: thingy [OPTIONS]
-h Display this usage message
-H hostname Hostname to connect to
""")
```
Strings can be concatenated (glued together) with the `+` operator, and repeated with `*`:
```
# 3 times 'un', followed by 'ium'
3 * 'un' + 'ium'
```
Two or more _string literals_ (i.e. the ones enclosed between quotes) next to each other are automatically concatenated.
```
'Py' 'thon'
```
This only works with two literals though, not with variables or expressions:
```
prefix = 'Py'
prefix 'thon' # Can't concatenate a variable and a string literal.
('un' * 3) 'ium'
```
If you want to concatenate variables or a variable and a literal, use `+`:
```
prefix = 'Py'
prefix + 'thon'
```
This feature is particularly useful when you want to break long strings:
```
text = ('Put several strings within parentheses '
'to have them joined together.')
text
```
Strings can be _indexed_ (subscripted), with the first character having index 0. There is no separate character type; a character is simply a string of size one:
```
word = 'Python'
word[0] # Character in position 0.
word[5] # Character in position 5.
```
Indices may also be negative numbers, to start counting from the right:
```
word[-1] # Last character.
word[-2] # Second-last character.
word[-6]
```
Note that since -0 is the same as 0, negative indices start from -1.
In addition to indexing, _slicing_ is also supported. While indexing is used to obtain individual characters, slicing allows you to obtain substring:
```
word[0:2] # Characters from position 0 (included) to 2 (excluded).
word[2:5] # Characters from position 2 (included) to 5 (excluded).
```
Note how the start is always included, and the end always excluded. This makes sure that `s[:i] + s[i:]` is always equal to `s`:
```
word[:2] + word[2:]
word[:4] + word[4:]
```
Slice indices have useful defaults; an omitted first index defaults to zero, an omitted second index defaults to the size of the string being sliced.
```
word[:2] # Character from the beginning to position 2 (excluded).
word[4:] # Characters from position 4 (included) to the end.
word[-2:] # Characters from the second-last (included) to the end.
```
One way to remember how slices work is to think of the indices as pointing between characters, with the left edge of the first character numbered 0. Then the right edge of the last character of a string of _n_ characters has index _n_, for example:
The first row of numbers gives the position of the indices 0...6 in the string; the second row gives the corresponding negative indices. The slice from _i_ to _j_ consists of all characters between the edges labeled _i_ and _j_, respectively.
For non-negative indices, the length of a slice is the difference of the indices, if both are within bounds. For example, the length of `word[1:3]` is 2.
Attempting to use an index that is too large will result in an error:
```
word[42] # The word only has 6 characters.
word[4:42]
word[42:]
```
Python strings cannot be changed â they are [immutable](https://docs.python.org/3.5/glossary.html#term-immutable). Therefore, assigning to an indexed position in the string results in an error:
```
word[0] = 'J'
word[2:] = 'py'
'J' + word[1:]
word[:2] + 'Py'
```
The built-in function [`len()`](https://docs.python.org/3.5/library/functions.html#len) returns the length of a string:
```
s = 'supercalifragilisticexpialidocious'
len(s)
```
See also:
- [Text Sequence Type â str](https://docs.python.org/3.5/library/stdtypes.html#textseq): Strings are examples of _sequence types_, and support the common operations supported by such types.
- [String Methods](https://docs.python.org/3.5/library/stdtypes.html#string-methods): Strings support a large number of methods for basic transformations and searching.
- [Format String Syntax](https://docs.python.org/3.5/library/string.html#formatstrings): Information about string formatting with [`str.format()`](https://docs.python.org/3.5/library/string.html#formatstrings).
- [`printf`-style String Formatting](https://docs.python.org/3.5/library/stdtypes.html#old-string-formatting): The old formatting operations invoked when strings and Unicode strings are the left operand of the `%` operator.
### Lists
Python knows a number of _compound_ data types, used to group together other values. The most versatile is the [_list_](https://docs.python.org/3.5/library/stdtypes.html#typesseq-list), which can be written as a list of comma-separated values (items) between square brackets. Lists might contain items of different types, but usually the items all have the same type.
```
squares = [1, 4, 9, 16, 25]
squares
```
Like strings (and all other built-in [sequence](https://docs.python.org/3.5/glossary.html#term-sequence) type), lists can be indexed and sliced:
```
squares[0] # Indexing returns the item.
squares[-1]
squares[-3:] # Slicing returns a new list.
```
All slice operations return a new list containing the requested elements. This means that the following slice returns a new (shallow) copy of the list:
```
squares[:]
```
Lists also support operations like concatenation:
```
squares + [36, 49, 64, 81, 100]
```
Unlike strings, which are [immutable](https://docs.python.org/3.5/glossary.html#term-immutable), lists are a [mutable](https://docs.python.org/3.5/glossary.html#term-mutable) type, i.e. it is possible to change their content:
```
cubes = [1, 8, 27, 65, 125] # Something's wrong here ...
4 ** 3 # the cube of 4 is 64, not 65!
cubes[3] = 64 # Replace the wrong value.
cubes
```
You can also add new items at the end of the list, by using the `append()` method (we will see more about methods later):
```
cubes.append(216) # Add the cube of 6 ...
cubes.append(7 ** 3) # and the cube of 7.
cubes
```
Assignment to slices is also possible, and this can even change the size of the list or clear it entirely:
```
letters = ['a', 'b', 'c', 'd', 'e', 'f', 'g']
letters
# Replace some values.
letters[2:5] = ['C', 'D', 'E']
letters
# Now remove them.
letters[2:5] = []
letters
# Clear the list by replacing all the elements with an empty list.
letters[:] = []
letters
```
The built-in function [`len()`](https://docs.python.org/3.5/library/functions.html#len) also applies to lists:
```
letters = ['a', 'b', 'c', 'd']
len(letters)
```
It is possible to nest lists (create lists containing other lists), for example:
```
a = ['a', 'b', 'c']
n = [1, 2, 3]
x = [a, n]
x
x[0]
x[0][1]
```
## First Steps Towards Programming
Of course, we can use Python for more complicated tasks than adding two and two together. For instance, we can write an initial sub-sequence of the Fibonacci series as follows:
```
# Fibonacci series:
# the sum of two elements defines the next.
a, b = 0, 1
while b < 10:
print(b)
a, b = b, a+b
```
This example introduces several new features.
- The first line contains a _multiple assignment_: the variables `a` and `b` simultaneously get the new values 0 and 1. On the last line this is used again, demonstrating that the expressions on the right-hand side are all evaluated first before any of the assignments take place. The right-hand side expressions are evaluated from the left to the right.
- The [`while`](https://docs.python.org/3.5/reference/compound_stmts.html#while) loop executes as long as the condition (here: `b < 10`) remains true. In Python, like in C, any non-zero integer value is true; zero is false. The condition may also be a string or list value, in fact any sequence; anything with a non-zero length is true, empty sequences are false. The test used in the example is a simple comparison. The standard comparison operators are written the same as in C: `<` (less than), `>` (greater than), `==` (equal to), `<=` (less than or equal to), `>=` (greater than or equal to) and `!=` (not equal to).
- The _body_ of the loop is _indented_: indentation is Pythonâs way of grouping statements. At the interactive prompt, you have to type a tab or space(s) for each indented line. In practice you will prepare more complicated input for Python with a text editor; all decent text editors have an auto-indent facility. When a compound statement is entered interactively, it must be followed by a blank line to indicate completion (since the parser cannot guess when you have typed the last line). Note that each line within a basic block must be indented by the same amount.
- The [`print()`](https://docs.python.org/3.5/library/functions.html#print) function writes the value of the argument(s) it is given. It differs from just writing the expression you want to write (as we did earlier in the calculator examples) in the way it handles multiple arguments, floating point quantities, and strings. Strings are printed without quotes, and a space is inserted between items, so you can format things nicely, like this:
```
i = 256*256
print('The value of i is', i)
```
The keyword argument `end` can be used to avoid the newline after the output, or end the output with a different string:
```
a, b = 0, 1
while b < 1000:
print(b, end=',')
a, b = b, a+b
```
| github_jupyter |
# Plagiarism Detection, Feature Engineering
In this project, you will be tasked with building a plagiarism detector that examines an answer text file and performs binary classification; labeling that file as either plagiarized or not, depending on how similar that text file is to a provided, source text.
Your first task will be to create some features that can then be used to train a classification model. This task will be broken down into a few discrete steps:
* Clean and pre-process the data.
* Define features for comparing the similarity of an answer text and a source text, and extract similarity features.
* Select "good" features, by analyzing the correlations between different features.
* Create train/test `.csv` files that hold the relevant features and class labels for train/test data points.
In the _next_ notebook, Notebook 3, you'll use the features and `.csv` files you create in _this_ notebook to train a binary classification model in a SageMaker notebook instance.
You'll be defining a few different similarity features, as outlined in [this paper](https://s3.amazonaws.com/video.udacity-data.com/topher/2019/January/5c412841_developing-a-corpus-of-plagiarised-short-answers/developing-a-corpus-of-plagiarised-short-answers.pdf), which should help you build a robust plagiarism detector!
To complete this notebook, you'll have to complete all given exercises and answer all the questions in this notebook.
> All your tasks will be clearly labeled **EXERCISE** and questions as **QUESTION**.
It will be up to you to decide on the features to include in your final training and test data.
---
## Read in the Data
The cell below will download the necessary, project data and extract the files into the folder `data/`.
This data is a slightly modified version of a dataset created by Paul Clough (Information Studies) and Mark Stevenson (Computer Science), at the University of Sheffield. You can read all about the data collection and corpus, at [their university webpage](https://ir.shef.ac.uk/cloughie/resources/plagiarism_corpus.html).
> **Citation for data**: Clough, P. and Stevenson, M. Developing A Corpus of Plagiarised Short Answers, Language Resources and Evaluation: Special Issue on Plagiarism and Authorship Analysis, In Press. [Download]
```
# NOTE:
# you only need to run this cell if you have not yet downloaded the data
# otherwise you may skip this cell or comment it out
!wget https://s3.amazonaws.com/video.udacity-data.com/topher/2019/January/5c4147f9_data/data.zip
!unzip data
# import libraries
import pandas as pd
import numpy as np
import os
```
This plagiarism dataset is made of multiple text files; each of these files has characteristics that are is summarized in a `.csv` file named `file_information.csv`, which we can read in using `pandas`.
```
csv_file = 'data/file_information.csv'
plagiarism_df = pd.read_csv(csv_file)
# print out the first few rows of data info
plagiarism_df.head()
```
## Types of Plagiarism
Each text file is associated with one **Task** (task A-E) and one **Category** of plagiarism, which you can see in the above DataFrame.
### Tasks, A-E
Each text file contains an answer to one short question; these questions are labeled as tasks A-E. For example, Task A asks the question: "What is inheritance in object oriented programming?"
### Categories of plagiarism
Each text file has an associated plagiarism label/category:
**1. Plagiarized categories: `cut`, `light`, and `heavy`.**
* These categories represent different levels of plagiarized answer texts. `cut` answers copy directly from a source text, `light` answers are based on the source text but include some light rephrasing, and `heavy` answers are based on the source text, but *heavily* rephrased (and will likely be the most challenging kind of plagiarism to detect).
**2. Non-plagiarized category: `non`.**
* `non` indicates that an answer is not plagiarized; the Wikipedia source text is not used to create this answer.
**3. Special, source text category: `orig`.**
* This is a specific category for the original, Wikipedia source text. We will use these files only for comparison purposes.
---
## Pre-Process the Data
In the next few cells, you'll be tasked with creating a new DataFrame of desired information about all of the files in the `data/` directory. This will prepare the data for feature extraction and for training a binary, plagiarism classifier.
### EXERCISE: Convert categorical to numerical data
You'll notice that the `Category` column in the data, contains string or categorical values, and to prepare these for feature extraction, we'll want to convert these into numerical values. Additionally, our goal is to create a binary classifier and so we'll need a binary class label that indicates whether an answer text is plagiarized (1) or not (0). Complete the below function `numerical_dataframe` that reads in a `file_information.csv` file by name, and returns a *new* DataFrame with a numerical `Category` column and a new `Class` column that labels each answer as plagiarized or not.
Your function should return a new DataFrame with the following properties:
* 4 columns: `File`, `Task`, `Category`, `Class`. The `File` and `Task` columns can remain unchanged from the original `.csv` file.
* Convert all `Category` labels to numerical labels according to the following rules (a higher value indicates a higher degree of plagiarism):
* 0 = `non`
* 1 = `heavy`
* 2 = `light`
* 3 = `cut`
* -1 = `orig`, this is a special value that indicates an original file.
* For the new `Class` column
* Any answer text that is not plagiarized (`non`) should have the class label `0`.
* Any plagiarized answer texts should have the class label `1`.
* And any `orig` texts will have a special label `-1`.
### Expected output
After running your function, you should get a DataFrame with rows that looks like the following:
```
File Task Category Class
0 g0pA_taska.txt a 0 0
1 g0pA_taskb.txt b 3 1
2 g0pA_taskc.txt c 2 1
3 g0pA_taskd.txt d 1 1
4 g0pA_taske.txt e 0 0
...
...
99 orig_taske.txt e -1 -1
```
```
# Read in a csv file and return a transformed dataframe
def numerical_dataframe(csv_file='data/file_information.csv'):
'''Reads in a csv file which is assumed to have `File`, `Category` and `Task` columns.
This function does two things:
1) converts `Category` column values to numerical values
2) Adds a new, numerical `Class` label column.
The `Class` column will label plagiarized answers as 1 and non-plagiarized as 0.
Source texts have a special label, -1.
:param csv_file: The directory for the file_information.csv file
:return: A dataframe with numerical categories and a new `Class` label column'''
# your code here
df = pd.read_csv(csv_file)
df['Category'] = df['Category'].map({'non':0, 'heavy':1, 'light':2, 'cut':3, 'orig':-1})
df['Class'] = df['Category'].clip(-1,1)
return df
```
### Test cells
Below are a couple of test cells. The first is an informal test where you can check that your code is working as expected by calling your function and printing out the returned result.
The **second** cell below is a more rigorous test cell. The goal of a cell like this is to ensure that your code is working as expected, and to form any variables that might be used in _later_ tests/code, in this case, the data frame, `transformed_df`.
> The cells in this notebook should be run in chronological order (the order they appear in the notebook). This is especially important for test cells.
Often, later cells rely on the functions, imports, or variables defined in earlier cells. For example, some tests rely on previous tests to work.
These tests do not test all cases, but they are a great way to check that you are on the right track!
```
# informal testing, print out the results of a called function
# create new `transformed_df`
transformed_df = numerical_dataframe(csv_file ='data/file_information.csv')
# check work
# check that all categories of plagiarism have a class label = 1
transformed_df.head(10)
# test cell that creates `transformed_df`, if tests are passed
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
# importing tests
import problem_unittests as tests
# test numerical_dataframe function
tests.test_numerical_df(numerical_dataframe)
# if above test is passed, create NEW `transformed_df`
transformed_df = numerical_dataframe(csv_file ='data/file_information.csv')
# check work
print('\nExample data: ')
transformed_df.head()
```
## Text Processing & Splitting Data
Recall that the goal of this project is to build a plagiarism classifier. At it's heart, this task is a comparison text; one that looks at a given answer and a source text, compares them and predicts whether an answer has plagiarized from the source. To effectively do this comparison, and train a classifier we'll need to do a few more things: pre-process all of our text data and prepare the text files (in this case, the 95 answer files and 5 original source files) to be easily compared, and split our data into a `train` and `test` set that can be used to train a classifier and evaluate it, respectively.
To this end, you've been provided code that adds additional information to your `transformed_df` from above. The next two cells need not be changed; they add two additional columns to the `transformed_df`:
1. A `Text` column; this holds all the lowercase text for a `File`, with extraneous punctuation removed.
2. A `Datatype` column; this is a string value `train`, `test`, or `orig` that labels a data point as part of our train or test set
The details of how these additional columns are created can be found in the `helpers.py` file in the project directory. You're encouraged to read through that file to see exactly how text is processed and how data is split.
Run the cells below to get a `complete_df` that has all the information you need to proceed with plagiarism detection and feature engineering.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
import helpers
# create a text column
text_df = helpers.create_text_column(transformed_df)
text_df.head()
# after running the cell above
# check out the processed text for a single file, by row index
row_idx = 0 # feel free to change this index
sample_text = text_df.iloc[0]['Text']
print('Sample processed text:\n\n', sample_text)
```
## Split data into training and test sets
The next cell will add a `Datatype` column to a given DataFrame to indicate if the record is:
* `train` - Training data, for model training.
* `test` - Testing data, for model evaluation.
* `orig` - The task's original answer from wikipedia.
### Stratified sampling
The given code uses a helper function which you can view in the `helpers.py` file in the main project directory. This implements [stratified random sampling](https://en.wikipedia.org/wiki/Stratified_sampling) to randomly split data by task & plagiarism amount. Stratified sampling ensures that we get training and test data that is fairly evenly distributed across task & plagiarism combinations. Approximately 26% of the data is held out for testing and 74% of the data is used for training.
The function **train_test_dataframe** takes in a DataFrame that it assumes has `Task` and `Category` columns, and, returns a modified frame that indicates which `Datatype` (train, test, or orig) a file falls into. This sampling will change slightly based on a passed in *random_seed*. Due to a small sample size, this stratified random sampling will provide more stable results for a binary plagiarism classifier. Stability here is smaller *variance* in the accuracy of classifier, given a random seed.
```
random_seed = 1 # can change; set for reproducibility
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
import helpers
# create new df with Datatype (train, test, orig) column
# pass in `text_df` from above to create a complete dataframe, with all the information you need
complete_df = helpers.train_test_dataframe(text_df, random_seed=random_seed)
# check results
complete_df.head(10)
```
# Determining Plagiarism
Now that you've prepared this data and created a `complete_df` of information, including the text and class associated with each file, you can move on to the task of extracting similarity features that will be useful for plagiarism classification.
> Note: The following code exercises, assume that the `complete_df` as it exists now, will **not** have its existing columns modified.
The `complete_df` should always include the columns: `['File', 'Task', 'Category', 'Class', 'Text', 'Datatype']`. You can add additional columns, and you can create any new DataFrames you need by copying the parts of the `complete_df` as long as you do not modify the existing values, directly.
---
# Similarity Features
One of the ways we might go about detecting plagiarism, is by computing **similarity features** that measure how similar a given answer text is as compared to the original wikipedia source text (for a specific task, a-e). The similarity features you will use are informed by [this paper on plagiarism detection](https://s3.amazonaws.com/video.udacity-data.com/topher/2019/January/5c412841_developing-a-corpus-of-plagiarised-short-answers/developing-a-corpus-of-plagiarised-short-answers.pdf).
> In this paper, researchers created features called **containment** and **longest common subsequence**.
Using these features as input, you will train a model to distinguish between plagiarized and not-plagiarized text files.
## Feature Engineering
Let's talk a bit more about the features we want to include in a plagiarism detection model and how to calculate such features. In the following explanations, I'll refer to a submitted text file as a **Student Answer Text (A)** and the original, wikipedia source file (that we want to compare that answer to) as the **Wikipedia Source Text (S)**.
### Containment
Your first task will be to create **containment features**. To understand containment, let's first revisit a definition of [n-grams](https://en.wikipedia.org/wiki/N-gram). An *n-gram* is a sequential word grouping. For example, in a line like "bayes rule gives us a way to combine prior knowledge with new information," a 1-gram is just one word, like "bayes." A 2-gram might be "bayes rule" and a 3-gram might be "combine prior knowledge."
> Containment is defined as the **intersection** of the n-gram word count of the Wikipedia Source Text (S) with the n-gram word count of the Student Answer Text (S) *divided* by the n-gram word count of the Student Answer Text.
$$ \frac{\sum{count(\text{ngram}_{A}) \cap count(\text{ngram}_{S})}}{\sum{count(\text{ngram}_{A})}} $$
If the two texts have no n-grams in common, the containment will be 0, but if _all_ their n-grams intersect then the containment will be 1. Intuitively, you can see how having longer n-gram's in common, might be an indication of cut-and-paste plagiarism. In this project, it will be up to you to decide on the appropriate `n` or several `n`'s to use in your final model.
### EXERCISE: Create containment features
Given the `complete_df` that you've created, you should have all the information you need to compare any Student Answer Text (A) with its appropriate Wikipedia Source Text (S). An answer for task A should be compared to the source text for task A, just as answers to tasks B, C, D, and E should be compared to the corresponding original source text.
In this exercise, you'll complete the function, `calculate_containment` which calculates containment based upon the following parameters:
* A given DataFrame, `df` (which is assumed to be the `complete_df` from above)
* An `answer_filename`, such as 'g0pB_taskd.txt'
* An n-gram length, `n`
### Containment calculation
The general steps to complete this function are as follows:
1. From *all* of the text files in a given `df`, create an array of n-gram counts; it is suggested that you use a [CountVectorizer](https://scikit-learn.org/stable/modules/generated/sklearn.feature_extraction.text.CountVectorizer.html) for this purpose.
2. Get the processed answer and source texts for the given `answer_filename`.
3. Calculate the containment between an answer and source text according to the following equation.
>$$ \frac{\sum{count(\text{ngram}_{A}) \cap count(\text{ngram}_{S})}}{\sum{count(\text{ngram}_{A})}} $$
4. Return that containment value.
You are encouraged to write any helper functions that you need to complete the function below.
```
# Calculate the ngram containment for one answer file/source file pair in a df
from sklearn.feature_extraction.text import CountVectorizer
def calculate_containment(df, n, answer_filename):
'''Calculates the containment between a given answer text and its associated source text.
This function creates a count of ngrams (of a size, n) for each text file in our data.
Then calculates the containment by finding the ngram count for a given answer text,
and its associated source text, and calculating the normalized intersection of those counts.
:param df: A dataframe with columns,
'File', 'Task', 'Category', 'Class', 'Text', and 'Datatype'
:param n: An integer that defines the ngram size
:param answer_filename: A filename for an answer text in the df, ex. 'g0pB_taskd.txt'
:return: A single containment value that represents the similarity
between an answer text and its source text.
'''
# your code here
source_filename = 'orig_' + answer_filename.split('_')[1]
answer = df['Text'].loc[df['File']==answer_filename].values[0]
source = df['Text'].loc[df['File']==source_filename].values[0]
counts = CountVectorizer(analyzer='word', ngram_range=(n,n))
ngrams = counts.fit_transform([answer, source])
ngram_array = ngrams.toarray()
return np.sum(np.min(ngram_array, axis=0))/sum(ngram_array[0])
```
### Test cells
After you've implemented the containment function, you can test out its behavior.
The cell below iterates through the first few files, and calculates the original category _and_ containment values for a specified n and file.
>If you've implemented this correctly, you should see that the non-plagiarized have low or close to 0 containment values and that plagiarized examples have higher containment values, closer to 1.
Note what happens when you change the value of n. I recommend applying your code to multiple files and comparing the resultant containment values. You should see that the highest containment values correspond to files with the highest category (`cut`) of plagiarism level.
```
# select a value for n
n = 1
# indices for first few files
test_indices = range(5)
# iterate through files and calculate containment
category_vals = []
containment_vals = []
for i in test_indices:
# get level of plagiarism for a given file index
category_vals.append(complete_df.loc[i, 'Category'])
# calculate containment for given file and n
filename = complete_df.loc[i, 'File']
c = calculate_containment(complete_df, n, filename)
containment_vals.append(c)
# print out result, does it make sense?
print('Original category values: \n', category_vals)
print()
print(str(n)+'-gram containment values: \n', containment_vals)
# run this test cell
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
# test containment calculation
# params: complete_df from before, and containment function
tests.test_containment(complete_df, calculate_containment)
```
### QUESTION 1: Why can we calculate containment features across *all* data (training & test), prior to splitting the DataFrame for modeling? That is, what about the containment calculation means that the test and training data do not influence each other?
**Answer:**
Each sample's containment calculation is independent of each other. No information of test set is used in training set feature.
---
## Longest Common Subsequence
Containment a good way to find overlap in word usage between two documents; it may help identify cases of cut-and-paste as well as paraphrased levels of plagiarism. Since plagiarism is a fairly complex task with varying levels, it's often useful to include other measures of similarity. The paper also discusses a feature called **longest common subsequence**.
> The longest common subsequence is the longest string of words (or letters) that are *the same* between the Wikipedia Source Text (S) and the Student Answer Text (A). This value is also normalized by dividing by the total number of words (or letters) in the Student Answer Text.
In this exercise, we'll ask you to calculate the longest common subsequence of words between two texts.
### EXERCISE: Calculate the longest common subsequence
Complete the function `lcs_norm_word`; this should calculate the *longest common subsequence* of words between a Student Answer Text and corresponding Wikipedia Source Text.
It may be helpful to think of this in a concrete example. A Longest Common Subsequence (LCS) problem may look as follows:
* Given two texts: text A (answer text) of length n, and string S (original source text) of length m. Our goal is to produce their longest common subsequence of words: the longest sequence of words that appear left-to-right in both texts (though the words don't have to be in continuous order).
* Consider:
* A = "i think pagerank is a link analysis algorithm used by google that uses a system of weights attached to each element of a hyperlinked set of documents"
* S = "pagerank is a link analysis algorithm used by the google internet search engine that assigns a numerical weighting to each element of a hyperlinked set of documents"
* In this case, we can see that the start of each sentence of fairly similar, having overlap in the sequence of words, "pagerank is a link analysis algorithm used by" before diverging slightly. Then we **continue moving left -to-right along both texts** until we see the next common sequence; in this case it is only one word, "google". Next we find "that" and "a" and finally the same ending "to each element of a hyperlinked set of documents".
* Below, is a clear visual of how these sequences were found, sequentially, in each text.
<img src='notebook_ims/common_subseq_words.png' width=40% />
* Now, those words appear in left-to-right order in each document, sequentially, and even though there are some words in between, we count this as the longest common subsequence between the two texts.
* If I count up each word that I found in common I get the value 20. **So, LCS has length 20**.
* Next, to normalize this value, divide by the total length of the student answer; in this example that length is only 27. **So, the function `lcs_norm_word` should return the value `20/27` or about `0.7408`.**
In this way, LCS is a great indicator of cut-and-paste plagiarism or if someone has referenced the same source text multiple times in an answer.
### LCS, dynamic programming
If you read through the scenario above, you can see that this algorithm depends on looking at two texts and comparing them word by word. You can solve this problem in multiple ways. First, it may be useful to `.split()` each text into lists of comma separated words to compare. Then, you can iterate through each word in the texts and compare them, adding to your value for LCS as you go.
The method I recommend for implementing an efficient LCS algorithm is: using a matrix and dynamic programming. **Dynamic programming** is all about breaking a larger problem into a smaller set of subproblems, and building up a complete result without having to repeat any subproblems.
This approach assumes that you can split up a large LCS task into a combination of smaller LCS tasks. Let's look at a simple example that compares letters:
* A = "ABCD"
* S = "BD"
We can see right away that the longest subsequence of _letters_ here is 2 (B and D are in sequence in both strings). And we can calculate this by looking at relationships between each letter in the two strings, A and S.
Here, I have a matrix with the letters of A on top and the letters of S on the left side:
<img src='notebook_ims/matrix_1.png' width=40% />
This starts out as a matrix that has as many columns and rows as letters in the strings S and O **+1** additional row and column, filled with zeros on the top and left sides. So, in this case, instead of a 2x4 matrix it is a 3x5.
Now, we can fill this matrix up by breaking it into smaller LCS problems. For example, let's first look at the shortest substrings: the starting letter of A and S. We'll first ask, what is the Longest Common Subsequence between these two letters "A" and "B"?
**Here, the answer is zero and we fill in the corresponding grid cell with that value.**
<img src='notebook_ims/matrix_2.png' width=30% />
Then, we ask the next question, what is the LCS between "AB" and "B"?
**Here, we have a match, and can fill in the appropriate value 1**.
<img src='notebook_ims/matrix_3_match.png' width=25% />
If we continue, we get to a final matrix that looks as follows, with a **2** in the bottom right corner.
<img src='notebook_ims/matrix_6_complete.png' width=25% />
The final LCS will be that value **2** *normalized* by the number of n-grams in A. So, our normalized value is 2/4 = **0.5**.
### The matrix rules
One thing to notice here is that, you can efficiently fill up this matrix one cell at a time. Each grid cell only depends on the values in the grid cells that are directly on top and to the left of it, or on the diagonal/top-left. The rules are as follows:
* Start with a matrix that has one extra row and column of zeros.
* As you traverse your string:
* If there is a match, fill that grid cell with the value to the top-left of that cell *plus* one. So, in our case, when we found a matching B-B, we added +1 to the value in the top-left of the matching cell, 0.
* If there is not a match, take the *maximum* value from either directly to the left or the top cell, and carry that value over to the non-match cell.
<img src='notebook_ims/matrix_rules.png' width=50% />
After completely filling the matrix, **the bottom-right cell will hold the non-normalized LCS value**.
This matrix treatment can be applied to a set of words instead of letters. Your function should apply this to the words in two texts and return the normalized LCS value.
```
# Compute the normalized LCS given an answer text and a source text
def lcs_norm_word(answer_text, source_text):
'''Computes the longest common subsequence of words in two texts; returns a normalized value.
:param answer_text: The pre-processed text for an answer text
:param source_text: The pre-processed text for an answer's associated source text
:return: A normalized LCS value'''
# your code here
A = answer_text.split()
B = source_text.split()
dp = np.zeros([len(A)+1, len(B)+1])
for i, a in enumerate(A):
for j, b in enumerate(B):
if a!=b:
dp[i+1,j+1] = max(dp[i,j+1], dp[i+1,j])
else:
dp[i+1,j+1] = dp[i,j]+1
return dp[-1,-1]/len(A)
```
### Test cells
Let's start by testing out your code on the example given in the initial description.
In the below cell, we have specified strings A (answer text) and S (original source text). We know that these texts have 20 words in common and the submitted answer is 27 words long, so the normalized, longest common subsequence should be 20/27.
```
# Run the test scenario from above
# does your function return the expected value?
A = "i think pagerank is a link analysis algorithm used by google that uses a system of weights attached to each element of a hyperlinked set of documents"
S = "pagerank is a link analysis algorithm used by the google internet search engine that assigns a numerical weighting to each element of a hyperlinked set of documents"
# calculate LCS
lcs = lcs_norm_word(A, S)
print('LCS = ', lcs)
# expected value test
assert lcs==20/27., "Incorrect LCS value, expected about 0.7408, got "+str(lcs)
print('Test passed!')
```
This next cell runs a more rigorous test.
```
# run test cell
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
# test lcs implementation
# params: complete_df from before, and lcs_norm_word function
tests.test_lcs(complete_df, lcs_norm_word)
```
Finally, take a look at a few resultant values for `lcs_norm_word`. Just like before, you should see that higher values correspond to higher levels of plagiarism.
```
# test on your own
test_indices = range(5) # look at first few files
category_vals = []
lcs_norm_vals = []
# iterate through first few docs and calculate LCS
for i in test_indices:
category_vals.append(complete_df.loc[i, 'Category'])
# get texts to compare
answer_text = complete_df.loc[i, 'Text']
task = complete_df.loc[i, 'Task']
# we know that source texts have Class = -1
orig_rows = complete_df[(complete_df['Class'] == -1)]
orig_row = orig_rows[(orig_rows['Task'] == task)]
source_text = orig_row['Text'].values[0]
# calculate lcs
lcs_val = lcs_norm_word(answer_text, source_text)
lcs_norm_vals.append(lcs_val)
# print out result, does it make sense?
print('Original category values: \n', category_vals)
print()
print('Normalized LCS values: \n', lcs_norm_vals)
```
---
# Create All Features
Now that you've completed the feature calculation functions, it's time to actually create multiple features and decide on which ones to use in your final model! In the below cells, you're provided two helper functions to help you create multiple features and store those in a DataFrame, `features_df`.
### Creating multiple containment features
Your completed `calculate_containment` function will be called in the next cell, which defines the helper function `create_containment_features`.
> This function returns a list of containment features, calculated for a given `n` and for *all* files in a df (assumed to the the `complete_df`).
For our original files, the containment value is set to a special value, -1.
This function gives you the ability to easily create several containment features, of different n-gram lengths, for each of our text files.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
# Function returns a list of containment features, calculated for a given n
# Should return a list of length 100 for all files in a complete_df
def create_containment_features(df, n, column_name=None):
containment_values = []
if(column_name==None):
column_name = 'c_'+str(n) # c_1, c_2, .. c_n
# iterates through dataframe rows
for i in df.index:
file = df.loc[i, 'File']
# Computes features using calculate_containment function
if df.loc[i,'Category'] > -1:
c = calculate_containment(df, n, file)
containment_values.append(c)
# Sets value to -1 for original tasks
else:
containment_values.append(-1)
print(str(n)+'-gram containment features created!')
return containment_values
```
### Creating LCS features
Below, your complete `lcs_norm_word` function is used to create a list of LCS features for all the answer files in a given DataFrame (again, this assumes you are passing in the `complete_df`. It assigns a special value for our original, source files, -1.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
# Function creates lcs feature and add it to the dataframe
def create_lcs_features(df, column_name='lcs_word'):
lcs_values = []
# iterate through files in dataframe
for i in df.index:
# Computes LCS_norm words feature using function above for answer tasks
if df.loc[i,'Category'] > -1:
# get texts to compare
answer_text = df.loc[i, 'Text']
task = df.loc[i, 'Task']
# we know that source texts have Class = -1
orig_rows = df[(df['Class'] == -1)]
orig_row = orig_rows[(orig_rows['Task'] == task)]
source_text = orig_row['Text'].values[0]
# calculate lcs
lcs = lcs_norm_word(answer_text, source_text)
lcs_values.append(lcs)
# Sets to -1 for original tasks
else:
lcs_values.append(-1)
print('LCS features created!')
return lcs_values
```
## EXERCISE: Create a features DataFrame by selecting an `ngram_range`
The paper suggests calculating the following features: containment *1-gram to 5-gram* and *longest common subsequence*.
> In this exercise, you can choose to create even more features, for example from *1-gram to 7-gram* containment features and *longest common subsequence*.
You'll want to create at least 6 features to choose from as you think about which to give to your final, classification model. Defining and comparing at least 6 different features allows you to discard any features that seem redundant, and choose to use the best features for your final model!
In the below cell **define an n-gram range**; these will be the n's you use to create n-gram containment features. The rest of the feature creation code is provided.
```
# Define an ngram range
ngram_range = range(1,7)
# The following code may take a minute to run, depending on your ngram_range
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
features_list = []
# Create features in a features_df
all_features = np.zeros((len(ngram_range)+1, len(complete_df)))
# Calculate features for containment for ngrams in range
i=0
for n in ngram_range:
column_name = 'c_'+str(n)
features_list.append(column_name)
# create containment features
all_features[i]=np.squeeze(create_containment_features(complete_df, n))
i+=1
# Calculate features for LCS_Norm Words
features_list.append('lcs_word')
all_features[i]= np.squeeze(create_lcs_features(complete_df))
# create a features dataframe
features_df = pd.DataFrame(np.transpose(all_features), columns=features_list)
# Print all features/columns
print()
print('Features: ', features_list)
print()
# print some results
features_df.head(10)
```
## Correlated Features
You should use feature correlation across the *entire* dataset to determine which features are ***too*** **highly-correlated** with each other to include both features in a single model. For this analysis, you can use the *entire* dataset due to the small sample size we have.
All of our features try to measure the similarity between two texts. Since our features are designed to measure similarity, it is expected that these features will be highly-correlated. Many classification models, for example a Naive Bayes classifier, rely on the assumption that features are *not* highly correlated; highly-correlated features may over-inflate the importance of a single feature.
So, you'll want to choose your features based on which pairings have the lowest correlation. These correlation values range between 0 and 1; from low to high correlation, and are displayed in a [correlation matrix](https://www.displayr.com/what-is-a-correlation-matrix/), below.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
# Create correlation matrix for just Features to determine different models to test
corr_matrix = features_df.corr().abs().round(2)
# display shows all of a dataframe
import seaborn as sns
sns.heatmap(corr_matrix, annot=True, cmap='coolwarm')
```
## EXERCISE: Create selected train/test data
Complete the `train_test_data` function below. This function should take in the following parameters:
* `complete_df`: A DataFrame that contains all of our processed text data, file info, datatypes, and class labels
* `features_df`: A DataFrame of all calculated features, such as containment for ngrams, n= 1-5, and lcs values for each text file listed in the `complete_df` (this was created in the above cells)
* `selected_features`: A list of feature column names, ex. `['c_1', 'lcs_word']`, which will be used to select the final features in creating train/test sets of data.
It should return two tuples:
* `(train_x, train_y)`, selected training features and their corresponding class labels (0/1)
* `(test_x, test_y)`, selected training features and their corresponding class labels (0/1)
** Note: x and y should be arrays of feature values and numerical class labels, respectively; not DataFrames.**
Looking at the above correlation matrix, you should decide on a **cutoff** correlation value, less than 1.0, to determine which sets of features are *too* highly-correlated to be included in the final training and test data. If you cannot find features that are less correlated than some cutoff value, it is suggested that you increase the number of features (longer n-grams) to choose from or use *only one or two* features in your final model to avoid introducing highly-correlated features.
Recall that the `complete_df` has a `Datatype` column that indicates whether data should be `train` or `test` data; this should help you split the data appropriately.
```
# Takes in dataframes and a list of selected features (column names)
# and returns (train_x, train_y), (test_x, test_y)
def train_test_data(complete_df, features_df, selected_features):
'''Gets selected training and test features from given dataframes, and
returns tuples for training and test features and their corresponding class labels.
:param complete_df: A dataframe with all of our processed text data, datatypes, and labels
:param features_df: A dataframe of all computed, similarity features
:param selected_features: An array of selected features that correspond to certain columns in `features_df`
:return: training and test features and labels: (train_x, train_y), (test_x, test_y)'''
# get the training features
train_x = features_df[selected_features].loc[complete_df['Datatype']=='train'].values
# And training class labels (0 or 1)
train_y = complete_df['Class'].loc[complete_df['Datatype']=='train'].values
# get the test features and labels
test_x = features_df[selected_features].loc[complete_df['Datatype']=='test'].values
test_y = complete_df['Class'].loc[complete_df['Datatype']=='test'].values
return (train_x, train_y), (test_x, test_y)
```
### Test cells
Below, test out your implementation and create the final train/test data.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
test_selection = list(features_df)[:2] # first couple columns as a test
# test that the correct train/test data is created
(train_x, train_y), (test_x, test_y) = train_test_data(complete_df, features_df, test_selection)
# params: generated train/test data
tests.test_data_split(train_x, train_y, test_x, test_y)
```
## EXERCISE: Select "good" features
If you passed the test above, you can create your own train/test data, below.
Define a list of features you'd like to include in your final mode, `selected_features`; this is a list of the features names you want to include.
```
# Select your list of features, this should be column names from features_df
# ex. ['c_1', 'lcs_word']
selected_features = ['c_1', 'c_6', 'lcs_word']
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
(train_x, train_y), (test_x, test_y) = train_test_data(complete_df, features_df, selected_features)
# check that division of samples seems correct
# these should add up to 95 (100 - 5 original files)
print('Training size: ', len(train_x))
print('Test size: ', len(test_x))
print()
print('Training df sample: \n', train_x[:10])
```
### Question 2: How did you decide on which features to include in your final model?
**Answer:**
---
## Creating Final Data Files
Now, you are almost ready to move on to training a model in SageMaker!
You'll want to access your train and test data in SageMaker and upload it to S3. In this project, SageMaker will expect the following format for your train/test data:
* Training and test data should be saved in one `.csv` file each, ex `train.csv` and `test.csv`
* These files should have class labels in the first column and features in the rest of the columns
This format follows the practice, outlined in the [SageMaker documentation](https://docs.aws.amazon.com/sagemaker/latest/dg/cdf-training.html), which reads: "Amazon SageMaker requires that a CSV file doesn't have a header record and that the target variable [class label] is in the first column."
## EXERCISE: Create csv files
Define a function that takes in x (features) and y (labels) and saves them to one `.csv` file at the path `data_dir/filename`.
It may be useful to use pandas to merge your features and labels into one DataFrame and then convert that into a csv file. You can make sure to get rid of any incomplete rows, in a DataFrame, by using `dropna`.
```
def make_csv(x, y, filename, data_dir):
'''Merges features and labels and converts them into one csv file with labels in the first column.
:param x: Data features
:param y: Data labels
:param file_name: Name of csv file, ex. 'train.csv'
:param data_dir: The directory where files will be saved
'''
# make data dir, if it does not exist
if not os.path.exists(data_dir):
os.makedirs(data_dir)
# your code here
pd.concat([pd.DataFrame(y), pd.DataFrame(x)], axis=1).to_csv(os.path.join(data_dir, filename), header=False, index=False)
# nothing is returned, but a print statement indicates that the function has run
print('Path created: '+str(data_dir)+'/'+str(filename))
```
### Test cells
Test that your code produces the correct format for a `.csv` file, given some text features and labels.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
fake_x = [ [0.39814815, 0.0001, 0.19178082],
[0.86936937, 0.44954128, 0.84649123],
[0.44086022, 0., 0.22395833] ]
fake_y = [0, 1, 1]
make_csv(fake_x, fake_y, filename='to_delete.csv', data_dir='test_csv')
# read in and test dimensions
fake_df = pd.read_csv('test_csv/to_delete.csv', header=None)
# check shape
assert fake_df.shape==(3, 4), \
'The file should have as many rows as data_points and as many columns as features+1 (for indices).'
# check that first column = labels
assert np.all(fake_df.iloc[:,0].values==fake_y), 'First column is not equal to the labels, fake_y.'
print('Tests passed!')
# delete the test csv file, generated above
! rm -rf test_csv
```
If you've passed the tests above, run the following cell to create `train.csv` and `test.csv` files in a directory that you specify! This will save the data in a local directory. Remember the name of this directory because you will reference it again when uploading this data to S3.
```
# can change directory, if you want
data_dir = 'plagiarism_data'
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
make_csv(train_x, train_y, filename='train.csv', data_dir=data_dir)
make_csv(test_x, test_y, filename='test.csv', data_dir=data_dir)
```
## Up Next
Now that you've done some feature engineering and created some training and test data, you are ready to train and deploy a plagiarism classification model. The next notebook will utilize SageMaker resources to train and test a model that you design.
| github_jupyter |
<a href="https://colab.research.google.com/github/sushantburnawal/awaaziitkgp/blob/master/C3_W2_Lab_3_Quantization_and_Pruning.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Ungraded Lab: Quantization and Pruning
In this lab, you will get some hands-on practice with the mobile optimization techniques discussed in the lectures. These enable reduced model size and latency which makes it ideal for edge and IOT devices. You will start by training a Keras model then compare its model size and accuracy after going through these techniques:
* post-training quantization
* quantization aware training
* weight pruning
Let's begin!
## Imports
Let's first import a few common libraries that you'll be using throughout the notebook.
```
import tensorflow as tf
import numpy as np
import os
import tempfile
import zipfile
```
<a name='utilities'>
## Utilities and constants
Let's first define a few string constants and utility functions to make our code easier to maintain.
```
# GLOBAL VARIABLES
# String constants for model filenames
FILE_WEIGHTS = 'baseline_weights.h5'
FILE_NON_QUANTIZED_H5 = 'non_quantized.h5'
FILE_NON_QUANTIZED_TFLITE = 'non_quantized.tflite'
FILE_PT_QUANTIZED = 'post_training_quantized.tflite'
FILE_QAT_QUANTIZED = 'quant_aware_quantized.tflite'
FILE_PRUNED_MODEL_H5 = 'pruned_model.h5'
FILE_PRUNED_QUANTIZED_TFLITE = 'pruned_quantized.tflite'
FILE_PRUNED_NON_QUANTIZED_TFLITE = 'pruned_non_quantized.tflite'
# Dictionaries to hold measurements
MODEL_SIZE = {}
ACCURACY = {}
# UTILITY FUNCTIONS
def print_metric(metric_dict, metric_name):
'''Prints key and values stored in a dictionary'''
for metric, value in metric_dict.items():
print(f'{metric_name} for {metric}: {value}')
def model_builder():
'''Returns a shallow CNN for training on the MNIST dataset'''
keras = tf.keras
# Define the model architecture.
model = keras.Sequential([
keras.layers.InputLayer(input_shape=(28, 28)),
keras.layers.Reshape(target_shape=(28, 28, 1)),
keras.layers.Conv2D(filters=12, kernel_size=(3, 3), activation='relu'),
keras.layers.MaxPooling2D(pool_size=(2, 2)),
keras.layers.Flatten(),
keras.layers.Dense(10, activation='softmax')
])
return model
def evaluate_tflite_model(filename, x_test, y_test):
'''
Measures the accuracy of a given TF Lite model and test set
Args:
filename (string) - filename of the model to load
x_test (numpy array) - test images
y_test (numpy array) - test labels
Returns
float showing the accuracy against the test set
'''
# Initialize the TF Lite Interpreter and allocate tensors
interpreter = tf.lite.Interpreter(model_path=filename)
interpreter.allocate_tensors()
# Get input and output index
input_index = interpreter.get_input_details()[0]["index"]
output_index = interpreter.get_output_details()[0]["index"]
# Initialize empty predictions list
prediction_digits = []
# Run predictions on every image in the "test" dataset.
for i, test_image in enumerate(x_test):
# Pre-processing: add batch dimension and convert to float32 to match with
# the model's input data format.
test_image = np.expand_dims(test_image, axis=0).astype(np.float32)
interpreter.set_tensor(input_index, test_image)
# Run inference.
interpreter.invoke()
# Post-processing: remove batch dimension and find the digit with highest
# probability.
output = interpreter.tensor(output_index)
digit = np.argmax(output()[0])
prediction_digits.append(digit)
# Compare prediction results with ground truth labels to calculate accuracy.
prediction_digits = np.array(prediction_digits)
accuracy = (prediction_digits == y_test).mean()
return accuracy
def get_gzipped_model_size(file):
'''Returns size of gzipped model, in bytes.'''
_, zipped_file = tempfile.mkstemp('.zip')
with zipfile.ZipFile(zipped_file, 'w', compression=zipfile.ZIP_DEFLATED) as f:
f.write(file)
return os.path.getsize(zipped_file)
```
## Download and Prepare the Dataset
You will be using the [MNIST](https://keras.io/api/datasets/mnist/) dataset which is hosted in [Keras Datasets](https://keras.io/api/datasets/). Some of the helper files in this notebook are made to work with this dataset so if you decide to switch to a different dataset, make sure to check if those helper functions need to be modified (e.g. shape of the Flatten layer in your model).
```
# Load MNIST dataset
mnist = tf.keras.datasets.mnist
(train_images, train_labels), (test_images, test_labels) = mnist.load_data()
# Normalize the input image so that each pixel value is between 0 to 1.
train_images = train_images / 255.0
test_images = test_images / 255.0
```
## Baseline Model
You will first build and train a Keras model. This will be the baseline where you will be comparing the mobile optimized versions later on. This will just be a shallow CNN with a softmax output to classify a given MNIST digit. You can review the `model_builder()` function in the utilities at the top of this notebook but we also printed the model summary below to show the architecture.
You will also save the weights so you can reinitialize the other models later the same way. This is not needed in real projects but for this demo notebook, it would be good to have the same initial state later so you can compare the effects of the optimizations.
```
# Create the baseline model
baseline_model = model_builder()
# Save the initial weights for use later
baseline_model.save_weights(FILE_WEIGHTS)
# Print the model summary
baseline_model.summary()
```
You can then compile and train the model. In practice, it's best to shuffle the train set but for this demo, it is set to `False` for reproducibility of the results. One epoch below will reach around 91% accuracy.
```
# Setup the model for training
baseline_model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
# Train the model
baseline_model.fit(train_images, train_labels, epochs=1, shuffle=False)
```
Let's save the accuracy of the model against the test set so you can compare later.
```
# Get the baseline accuracy
_, ACCURACY['baseline Keras model'] = baseline_model.evaluate(test_images, test_labels)
```
Next, you will save the Keras model as a file and record its size as well.
```
# Save the Keras model
baseline_model.save(FILE_NON_QUANTIZED_H5, include_optimizer=False)
# Save and get the model size
MODEL_SIZE['baseline h5'] = os.path.getsize(FILE_NON_QUANTIZED_H5)
# Print records so far
print_metric(ACCURACY, "test accuracy")
print_metric(MODEL_SIZE, "model size in bytes")
```
### Convert the model to TF Lite format
Next, you will convert the model to [Tensorflow Lite (TF Lite)](https://www.tensorflow.org/lite/guide) format. This is designed to make Tensorflow models more efficient and lightweight when running on mobile, embedded, and IOT devices.
You can convert a Keras model with TF Lite's [Converter](https://www.tensorflow.org/lite/convert/index) class and we've incorporated it in the short helper function below. Notice that there is a `quantize` flag which you can use to quantize the model.
```
def convert_tflite(model, filename, quantize=False):
'''
Converts the model to TF Lite format and writes to a file
Args:
model (Keras model) - model to convert to TF Lite
filename (string) - string to use when saving the file
quantize (bool) - flag to indicate quantization
Returns:
None
'''
# Initialize the converter
converter = tf.lite.TFLiteConverter.from_keras_model(model)
# Set for quantization if flag is set to True
if quantize:
converter.optimizations = [tf.lite.Optimize.DEFAULT]
# Convert the model
tflite_model = converter.convert()
# Save the model.
with open(filename, 'wb') as f:
f.write(tflite_model)
```
You will use the helper function to convert the Keras model then get its size and accuracy. Take note that this is *not yet* quantized.
```
# Convert baseline model
convert_tflite(baseline_model, FILE_NON_QUANTIZED_TFLITE)
```
You will notice that there is already a slight decrease in model size when converting to `.tflite` format.
```
MODEL_SIZE['non quantized tflite'] = os.path.getsize(FILE_NON_QUANTIZED_TFLITE)
print_metric(MODEL_SIZE, 'model size in bytes')
```
The accuracy will also be nearly identical when converting between formats. You can setup a TF Lite model for input-output using its [Interpreter](https://www.tensorflow.org/api_docs/python/tf/lite/Interpreter) class. This is shown in the `evaluate_tflite_model()` helper function provided in the `Utilities` section earlier.
*Note: If you see a `Runtime Error: There is at least 1 reference to internal data in the interpreter in the form of a numpy array or slice.` , please try re-running the cell.*
```
ACCURACY['non quantized tflite'] = evaluate_tflite_model(FILE_NON_QUANTIZED_TFLITE, test_images, test_labels)
print_metric(ACCURACY, 'test accuracy')
```
### Post-Training Quantization
Now that you have the baseline metrics, you can now observe the effects of quantization. As mentioned in the lectures, this process involves converting floating point representations into integer to reduce model size and achieve faster computation.
As shown in the `convert_tflite()` helper function earlier, you can easily do [post-training quantization](https://www.tensorflow.org/lite/performance/post_training_quantization) with the TF Lite API. You just need to set the converter optimization and assign an [Optimize](https://www.tensorflow.org/api_docs/python/tf/lite/Optimize) Enum.
You will set the `quantize` flag to do that and get the metrics again.
```
# Convert and quantize the baseline model
convert_tflite(baseline_model, FILE_PT_QUANTIZED, quantize=True)
# Get the model size
MODEL_SIZE['post training quantized tflite'] = os.path.getsize(FILE_PT_QUANTIZED)
print_metric(MODEL_SIZE, 'model size')
```
You should see around a 4X reduction in model size in the quantized version. This comes from converting the 32 bit representations (float) into 8 bits (integer).
```
ACCURACY['post training quantized tflite'] = evaluate_tflite_model(FILE_PT_QUANTIZED, test_images, test_labels)
print_metric(ACCURACY, 'test accuracy')
```
As mentioned in the lecture, you can expect the accuracy to not be the same when quantizing the model. Most of the time it will decrease but in some cases, it can even increase. Again, this can be attributed to the loss of precision when you remove the extra bits from the float data.
## Quantization Aware Training
When post-training quantization results in loss of accuracy that is unacceptable for your application, you can consider doing [quantization aware training](https://www.tensorflow.org/model_optimization/guide/quantization/training) before quantizing the model. This simulates the loss of precision by inserting fake quant nodes in the model during training. That way, your model will learn to adapt with the loss of precision to get more accurate predictions.
The [Tensorflow Model Optimization Toolkit](https://www.tensorflow.org/model_optimization) provides a [quantize_model()](https://www.tensorflow.org/model_optimization/api_docs/python/tfmot/quantization/keras/quantize_model) method to do this quickly and you will see that below. But first, let's install the toolkit into the notebook environment.
```
# Install the toolkit
!pip install tensorflow_model_optimization
```
You will build the baseline model again but this time, you will pass it into the `quantize_model()` method to indicate quantization aware training.
Take note that in case you decide to pass in a model that is already trained, then make sure to recompile before you continue training.
```
import tensorflow_model_optimization as tfmot
# method to quantize a Keras model
quantize_model = tfmot.quantization.keras.quantize_model
# Define the model architecture.
model_to_quantize = model_builder()
# Reinitialize weights with saved file
model_to_quantize.load_weights(FILE_WEIGHTS)
# Quantize the model
q_aware_model = quantize_model(model_to_quantize)
# `quantize_model` requires a recompile.
q_aware_model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
q_aware_model.summary()
```
You may have noticed a slight difference in the model summary above compared to the baseline model summary in the earlier sections. The total params count increased as expected because of the nodes added by the `quantize_model()` method.
With that, you can now train the model. You will notice that the accuracy is a bit lower because the model is simulating the loss of precision. The training will take a bit longer if you want to achieve the same training accuracy as the earlier run. For this exercise though, we will keep to 1 epoch.
```
# Train the model
q_aware_model.fit(train_images, train_labels, epochs=1, shuffle=False)
```
You can then get the accuracy of the Keras model before and after quantizing the model. The accuracy is expected to be nearly identical because the model is trained to counter the effects of quantization.
```
# Reinitialize the dictionary
ACCURACY = {}
# Get the accuracy of the quantization aware trained model (not yet quantized)
_, ACCURACY['quantization aware non-quantized'] = q_aware_model.evaluate(test_images, test_labels, verbose=0)
print_metric(ACCURACY, 'test accuracy')
# Convert and quantize the model.
convert_tflite(q_aware_model, FILE_QAT_QUANTIZED, quantize=True)
# Get the accuracy of the quantized model
ACCURACY['quantization aware quantized'] = evaluate_tflite_model(FILE_QAT_QUANTIZED, test_images, test_labels)
print_metric(ACCURACY, 'test accuracy')
```
## Pruning
Let's now move on to another technique for reducing model size: [Pruning](https://www.tensorflow.org/model_optimization/guide/pruning/pruning_with_keras). This process involves zeroing out insignificant (i.e. low magnitude) weights. The intuition is these weights do not contribute as much to making predictions so you can remove them and get the same result. Making the weights sparse helps in compressing the model more efficiently and you will see that in this section.
The Tensorflow Model Optimization Toolkit again has a convenience method for this. The [prune_low_magnitude()](https://www.tensorflow.org/model_optimization/api_docs/python/tfmot/sparsity/keras/prune_low_magnitude) method puts wrappers in a Keras model so it can be pruned during training. You will pass in the baseline model that you already trained earlier. You will notice that the model summary show increased params because of the wrapper layers added by the pruning method.
You can set how the pruning is done during training. Below, you will use [PolynomialDecay](https://www.tensorflow.org/model_optimization/api_docs/python/tfmot/sparsity/keras/PolynomialDecay) to indicate how the sparsity ramps up with each step. Another option available in the library is [Constant Sparsity](https://www.tensorflow.org/model_optimization/api_docs/python/tfmot/sparsity/keras/ConstantSparsity).
```
# Get the pruning method
prune_low_magnitude = tfmot.sparsity.keras.prune_low_magnitude
# Compute end step to finish pruning after 2 epochs.
batch_size = 128
epochs = 2
validation_split = 0.1 # 10% of training set will be used for validation set.
num_images = train_images.shape[0] * (1 - validation_split)
end_step = np.ceil(num_images / batch_size).astype(np.int32) * epochs
# Define pruning schedule.
pruning_params = {
'pruning_schedule': tfmot.sparsity.keras.PolynomialDecay(initial_sparsity=0.50,
final_sparsity=0.80,
begin_step=0,
end_step=end_step)
}
# Pass in the trained baseline model
model_for_pruning = prune_low_magnitude(baseline_model, **pruning_params)
# `prune_low_magnitude` requires a recompile.
model_for_pruning.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
model_for_pruning.summary()
```
You can also peek at the weights of one of the layers in your model. After pruning, you will notice that many of these will be zeroed out.
```
# Preview model weights
model_for_pruning.weights[1]
```
With that, you can now start re-training the model. Take note that the [UpdatePruningStep()](https://www.tensorflow.org/model_optimization/api_docs/python/tfmot/sparsity/keras/UpdatePruningStep) callback is required.
```
# Callback to update pruning wrappers at each step
callbacks = [
tfmot.sparsity.keras.UpdatePruningStep(),
]
# Train and prune the model
model_for_pruning.fit(train_images, train_labels,
epochs=epochs, validation_split=validation_split,
callbacks=callbacks)
```
Now see how the weights in the same layer looks like after pruning.
```
# Preview model weights
model_for_pruning.weights[1]
```
After pruning, you can remove the wrapper layers to have the same layers and params as the baseline model. You can do that with the [strip_pruning()](https://www.tensorflow.org/model_optimization/api_docs/python/tfmot/sparsity/keras/strip_pruning) method as shown below. You will do this so you can save the model and also export to TF Lite format just like in the previous sections.
```
# Remove pruning wrappers
model_for_export = tfmot.sparsity.keras.strip_pruning(model_for_pruning)
model_for_export.summary()
```
You will see the same model weights but the index is different because the wrappers were removed.
```
# Preview model weights (index 1 earlier is now 0 because pruning wrappers were removed)
model_for_export.weights[0]
```
You will notice below that the pruned model will have the same file size as the baseline_model when saved as H5. This is to be expected. The improvement will be noticeable when you compress the model as will be shown in the cell after this.
```
# Save Keras model
model_for_export.save(FILE_PRUNED_MODEL_H5, include_optimizer=False)
# Get uncompressed model size of baseline and pruned models
MODEL_SIZE = {}
MODEL_SIZE['baseline h5'] = os.path.getsize(FILE_NON_QUANTIZED_H5)
MODEL_SIZE['pruned non quantized h5'] = os.path.getsize(FILE_PRUNED_MODEL_H5)
print_metric(MODEL_SIZE, 'model_size in bytes')
```
You will use the `get_gzipped_model_size()` helper function in the `Utilities` to compress the models and get its resulting file size. You will notice that the pruned model is about 3 times smaller. This is because of the sparse weights generated by the pruning process. The zeros can be compressed much more efficiently than the low magnitude weights before pruning.
```
# Get compressed size of baseline and pruned models
MODEL_SIZE = {}
MODEL_SIZE['baseline h5'] = get_gzipped_model_size(FILE_NON_QUANTIZED_H5)
MODEL_SIZE['pruned non quantized h5'] = get_gzipped_model_size(FILE_PRUNED_MODEL_H5)
print_metric(MODEL_SIZE, "gzipped model size in bytes")
```
You can make the model even more lightweight by quantizing the pruned model. This achieves around 10X reduction in compressed model size as compared to the baseline.
```
# Convert and quantize the pruned model.
pruned_quantized_tflite = convert_tflite(model_for_export, FILE_PRUNED_QUANTIZED_TFLITE, quantize=True)
# Compress and get the model size
MODEL_SIZE['pruned quantized tflite'] = get_gzipped_model_size(FILE_PRUNED_QUANTIZED_TFLITE)
print_metric(MODEL_SIZE, "gzipped model size in bytes")
```
As expected, the TF Lite model's accuracy will also be close to the Keras model.
```
# Get accuracy of pruned Keras and TF Lite models
ACCURACY = {}
_, ACCURACY['pruned model h5'] = model_for_pruning.evaluate(test_images, test_labels)
ACCURACY['pruned and quantized tflite'] = evaluate_tflite_model(FILE_PRUNED_QUANTIZED_TFLITE, test_images, test_labels)
print_metric(ACCURACY, 'accuracy')
```
## Wrap Up
In this notebook, you practiced several techniques in optimizing your models for mobile and embedded applications. You used quantization to reduce floating point representations into integer, then used pruning to make the weights sparse for efficient model compression. These make your models lightweight for efficient transport and storage without sacrificing model accuracy. Try this in your own models and see what performance you get. For more information, here are a few other resources:
* [Post Training Quantization Guide](https://www.tensorflow.org/lite/performance/post_training_quantization)
* [Quantization Aware Training Comprehensive Guide](https://www.tensorflow.org/model_optimization/guide/quantization/training_comprehensive_guide)
* [Pruning Comprehensive Guide](https://www.tensorflow.org/model_optimization/guide/pruning/comprehensive_guide)
**Congratulations and enjoy the rest of the course!**
| github_jupyter |
```
import os
import sys
from os.path import dirname
proj_path = dirname(os.getcwd())
sys.path.append(proj_path)
from typing import Mapping
import torch
import torch.distributions as D
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
import tqdm
from absl import app
from absl import flags
from absl import logging
from oatomobile.baselines.torch.dim.model import ImitativeModel
from oatomobile.datasets.carla import CARLADataset
from oatomobile.torch import types
from oatomobile.torch.loggers import TensorBoardLogger
from oatomobile.torch.savers import Checkpointer
import numpy as np
import ray.train as train
from ray.train import Trainer, TorchConfig
from ray.train.callbacks import JsonLoggerCallback, TBXLoggerCallback
from torch.nn.parallel import DistributedDataParallel
from torch.utils.data import DistributedSampler
import ray
ray_address = "127.0.0.1"
num_workers = 4
smoke_test = True
dataset_dir = "nonlocalexamples"
output_dir = "models"
batch_size = 512
num_epochs = 3
save_model_frequency = 4
learning_rate = 1e-3
num_timesteps_to_keep = 4
weight_decay = 0.0
clip_gradients = False
def train_func(confg):
batch_size = config.get("batch_size", 32)
hidden_size = config.get("hidden_size", 1)
lr = config.get("lr", 1e-2)
epochs = config.get("epochs", 3)
weight_decay = config.get("weight_decay", 0)
# # Creates the necessary output directory.
# os.makedirs(output_dir, exist_ok=True)
# log_dir = os.path.join(output_dir, "logs")
# os.makedirs(log_dir, exist_ok=True)
# ckpt_dir = os.path.join(output_dir, "ckpts")
# os.makedirs(ckpt_dir, exist_ok=True)
# Initializes the model and its optimizer.
output_shape = [num_timesteps_to_keep, 2]
model = ImitativeModel(output_shape=output_shape).to(device)
model = DistributedDataParallel(model)
optimizer = optim.Adam(
model.parameters(),
lr=lr,
weight_decay=weight_decay,
)
# writer = TensorBoardLogger(log_dir=log_dir)
# checkpointer = Checkpointer(model=model, ckpt_dir=ckpt_dir)
def transform(batch: Mapping[str, types.Array]) -> Mapping[str, torch.Tensor]:
"""Preprocesses a batch for the model.
Args:
batch: (keyword arguments) The raw batch variables.
Returns:
The processed batch.
"""
# Sends tensors to `device`.
batch = {key: tensor.to(device) for (key, tensor) in batch.items()}
# Preprocesses batch for the model.
batch = model.transform(batch)
return batch
# Setups the dataset and the dataloader.
modalities = (
"lidar",
"is_at_traffic_light",
"traffic_light_state",
"player_future",
"velocity",
)
dataset_train = CARLADataset.as_torch(
dataset_dir=os.path.join(dataset_dir, "train"),
modalities=modalities,
)
dataloader_train = torch.utils.data.DataLoader(
dataset_train,
batch_size=batch_size,
shuffle=True,
num_workers=5,
)
dataset_val = CARLADataset.as_torch(
dataset_dir=os.path.join(dataset_dir, "val"),
modalities=modalities,
)
dataloader_val = torch.utils.data.DataLoader(
dataset_val,
batch_size=batch_size * 5,
shuffle=True,
num_workers=5,
)
# Theoretical limit of NLL.
nll_limit = -torch.sum( # pylint: disable=no-member
D.MultivariateNormal(
loc=torch.zeros(output_shape[-2] * output_shape[-1]), # pylint: disable=no-member
scale_tril=torch.eye(output_shape[-2] * output_shape[-1]) * # pylint: disable=no-member
noise_level, # pylint: disable=no-member
).log_prob(torch.zeros(output_shape[-2] * output_shape[-1]))) # pylint: disable=no-member
def train_step(
model: ImitativeModel,
optimizer: optim.Optimizer,
batch: Mapping[str, torch.Tensor],
clip: bool = False,
) -> torch.Tensor:
"""Performs a single gradient-descent optimisation step."""
# Resets optimizer's gradients.
optimizer.zero_grad()
# Perturb target.
y = torch.normal( # pylint: disable=no-member
mean=batch["player_future"][..., :2],
std=torch.ones_like(batch["player_future"][..., :2]) * noise_level, # pylint: disable=no-member
)
# Forward pass from the model.
z, _ = model._params(
velocity=batch["velocity"],
visual_features=batch["visual_features"],
is_at_traffic_light=batch["is_at_traffic_light"],
traffic_light_state=batch["traffic_light_state"],
)
_, log_prob, logabsdet = model._decoder._inverse(y=y, z=z)
# Calculates loss (NLL).
loss = -torch.mean(log_prob - logabsdet, dim=0) # pylint: disable=no-member
# Backward pass.
loss.backward()
# Clips gradients norm.
if clip:
torch.nn.utils.clip_grad_norm(model.parameters(), 1.0)
# Performs a gradient descent step.
optimizer.step()
return loss
def train_epoch(
model: ImitativeModel,
optimizer: optim.Optimizer,
dataloader: torch.utils.data.DataLoader,
) -> torch.Tensor:
"""Performs an epoch of gradient descent optimization on `dataloader`."""
model.train()
loss = 0.0
with tqdm.tqdm(dataloader) as pbar:
for batch in pbar:
# Prepares the batch.
batch = transform(batch)
# Performs a gradien-descent step.
loss += train_step(model, optimizer, batch, clip=clip_gradients)
return loss / len(dataloader)
def evaluate_step(
model: ImitativeModel,
batch: Mapping[str, torch.Tensor],
) -> torch.Tensor:
"""Evaluates `model` on a `batch`."""
# Forward pass from the model.
z, _ = model._params(
velocity=batch["velocity"],
visual_features=batch["visual_features"],
is_at_traffic_light=batch["is_at_traffic_light"],
traffic_light_state=batch["traffic_light_state"],
)
_, log_prob, logabsdet = model._decoder._inverse(
y=batch["player_future"][..., :2],
z=z,
)
# Calculates loss (NLL).
loss = -torch.mean(log_prob - logabsdet, dim=0) # pylint: disable=no-member
return loss
def evaluate_epoch(
model: ImitativeModel,
dataloader: torch.utils.data.DataLoader,
) -> torch.Tensor:
"""Performs an evaluation of the `model` on the `dataloader."""
model.eval()
loss = 0.0
with tqdm.tqdm(dataloader) as pbar:
for batch in pbar:
# Prepares the batch.
batch = transform(batch)
# Accumulates loss in dataset.
with torch.no_grad():
loss += evaluate_step(model, batch)
return loss / len(dataloader)
results = []
for _ in range(epochs):
# Trains model on whole training dataset, and writes on `TensorBoard`.
loss_train = train_epoch(model, optimizer, dataloader_train)
# write(model, dataloader_train, writer, "train", loss_train, epoch)
# Evaluates model on whole validation dataset, and writes on `TensorBoard`.
loss_val = evaluate_epoch(model, dataloader_val)
# write(model, dataloader_val, writer, "val", loss_val, epoch)
# # Checkpoints model weights.
# if epoch % save_model_frequency == 0:
# checkpointer.save(epoch)
# train_epoch(train_loader, model, loss_fn, optimizer)
# result = validate_epoch(validation_loader, model, loss_fn)
train.report(**result)
results.append(result)
return results
def train_carla():
trainer = Trainer(TorchConfig(backend="gloo"), num_workers=num_workers)
config = {"lr": learning_rate, "hidden_size": 1, "batch_size": batch_size, "epochs": num_epochs}
trainer.start()
results = trainer.run(
train_func,
config,
callbacks=[JsonLoggerCallback(),
TBXLoggerCallback()])
trainer.shutdown()
print(results)
return results
if __name__ == "__main__":
# Start Ray Cluster
if smoke_test:
ray.init(num_cpus=2)
else:
ray.init(address=address)
# Train carla
train_carla()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/peeyushsinghal/EVA/blob/main/S5-Assignment%20Solution/EVA_S5_Exp2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
### Importing Libraries
```
import torch
from torchvision import datasets,transforms
%matplotlib inline
import matplotlib.pyplot as plt # for visualizing images
import random # for random image index
import torch.nn as nn # for network
import torch.nn.functional as F # for forward method
import torch.optim as optim # for optimizer
!pip install torchsummary
from torchsummary import summary # for model summary and params
from tqdm import tqdm # for beautiful model training updates
```
Seed and Cuda
```
# check for cuda
cuda = torch.cuda.is_available()
print (f' Cuda Status : {cuda}')
# setting seed
SEED = 42 # arbit seed, why 42 - because in hitch hikers guide to galaxy it is answer to everything
# torch.cuda.seed(SEED)
torch.cuda.manual_seed_all(SEED) if cuda else torch.manual_seed(SEED)
```
### Downloading dataset, splitting datasets
loading dataset
```
train = datasets.MNIST(
root = './',# directory where data needs to be stored
train = True, # get the training portion of the dataset
download = True, # downloads
transform = transforms.Compose([
transforms.ToTensor(),# converts to tesnor
transforms.Normalize((0.1307,), (0.3081,))# Normalize
])
)
test = datasets.MNIST(
root = './',# directory where data needs to be stored
train = False, # get the test portion of the dataset
download = True, # downloads
transform = transforms.Compose([
transforms.ToTensor(),# converts to tesnor
transforms.Normalize((0.1307,), (0.3081,))# Normalize
])
)
```
Train and Test Dataloader
```
dataloader_args = dict(shuffle=True, batch_size=128, num_workers=4, pin_memory = True) if cuda else dict(shuffle=True, batch_size=64)
train_loader = torch.utils.data.DataLoader(
dataset=train,# train dataset
**dataloader_args # the dataloader arguments change dependent on cuda is available or not
)
test_loader = torch.utils.data.DataLoader(
dataset = test,# test dataset
**dataloader_args # the dataloader arguments change dependent on cuda is available or not
)
```
Checking Dataloaders
- sample data
```
images, labels = next(iter(train_loader))
print(images.shape)
print(labels.shape)
# printing random image and seeing
plt.imshow(images[random.randint(0,len(images))].numpy().squeeze(), cmap='gray_r')
# Looking at more images
figure = plt.figure()
for index in range(1, len(images) + 1): # assumption: batch size would be atleast 8
plt.subplot(8, int(len(images)/8), index)
plt.axis('off')
plt.imshow(images[index-1].numpy().squeeze(), cmap='gray_r')
```
### Network
```
class Network(nn.Module):
def __init__(self):
super(Network,self).__init__() # extending super class method
drop_out_value = 0.1
# Input Block
self.convblock1 = nn.Sequential(
nn.Conv2d(1,8,3), # In- 1x28x28, Out- 8x26x26, RF- 3x3, Jump_in -1, Jump_out -1
nn.ReLU(),
nn.BatchNorm2d(8),
nn.Dropout(drop_out_value)
)
# Conv Block 2
self.convblock2 = nn.Sequential(
nn.Conv2d(8,8,3), # In- 8x26x26, Out- 8x24x24, RF- 5x5, Jump_in -1, Jump_out -1
nn.ReLU(),
nn.BatchNorm2d(8),
nn.Dropout(drop_out_value)
)
# Conv Block 3
self.convblock3 = nn.Sequential(
nn.Conv2d(8,16,3), # In- 16x24x24, Out- 16x22x22, RF- 7x7, Jump_in -1, Jump_out -1
nn.ReLU(),
nn.BatchNorm2d(16),
nn.Dropout(drop_out_value)
)
# Transition Block 1 (this also includes a conv block)
self.pool1 = nn.MaxPool2d(kernel_size=2, stride=2) # In- 16x22x22, Out- 16x11x11, RF- 8x8, Jump_in -1, Jump_out -2
self.convblock4 = nn.Sequential(
nn.Conv2d(16,8,1), # In- 16x11x11, Out- 8x11x11, RF- 8x8, Jump_in -2, Jump_out -2
nn.ReLU(),
nn.BatchNorm2d(8),
nn.Dropout(drop_out_value)
)
# Conv Block 5
self.convblock5 = nn.Sequential(
nn.Conv2d(8,8,3), # In- 8x11x11, Out- 8x9x9, RF- 12x12, Jump_in -2, Jump_out -2
nn.ReLU(),
nn.BatchNorm2d(8),
nn.Dropout(drop_out_value)
)
# Conv Block 6
self.convblock6 = nn.Sequential(
nn.Conv2d(8,16,3), # In- 8x9x9, Out- 16x7x7, RF- 16x16, Jump_in -2, Jump_out -2
nn.ReLU(),
nn.BatchNorm2d(16),
nn.Dropout(drop_out_value)
)
# Output Block
self.convblock7 = nn.Sequential(
nn.Conv2d(16,10,1), # In- 16x7x7, Out- 10x7x7, RF- 16x16, Jump_in -2, Jump_out -2
nn.ReLU(),
nn.BatchNorm2d(10),
nn.Dropout(drop_out_value)
)
self.gap = nn.AvgPool2d(7) # In- 10x7x7, Out- 10x1x1, RF- 28x28, Jump_in -2, Jump_out -2
def forward(self,x):
x = self.convblock1(x)
x = self.convblock2(x)
x = self.convblock3(x)
x = self.pool1(x)
x = self.convblock4(x)
x = self.convblock5(x)
x = self.convblock6(x)
x = self.convblock7(x)
x = self.gap(x)
# Flattening
x = x.view(-1,10)
return F.log_softmax(x,dim=-1)
# model = Network()
# print(model)
```
### Model Params
- Checking the model summary and number of parameters
```
device = torch.device("cuda" if cuda else "cpu")
print(device)
model = Network().to(device)
# print(model)
summary(model, input_size=(1, 28, 28))
```
### Training and Testing
- includes test and train functions
- includes loop function, where test can happen after each epoch is trained
```
# Training Function
train_losses = [] # to capture train losses over training epochs
train_accuracy = [] # to capture train accuracy over training epochs
def train(model,device, train_loader,optimizer,epoch):
model.train() # setting the model in training mode
pbar = tqdm(train_loader) # putting the iterator in pbar
correct = 0 # for accuracy numerator
processed =0 # for accuracy denominator
for batch_idx, (images,labels) in enumerate(pbar):
images, labels = images.to(device),labels.to(device)#sending data to CPU or GPU as per device
optimizer.zero_grad() # setting gradients to zero to avoid accumulation
y_preds = model(images) # forward pass, result captured in y_preds (plural as there are many images in a batch)
# the predictions are in one hot vector
loss = F.nll_loss(y_preds,labels) # capturing loss
train_losses.append(loss) # to capture loss over many epochs
loss.backward() # backpropagation
optimizer.step() # updating the params
preds = y_preds.argmax(dim=1, keepdim=True) # get the index of the max log-probability
correct += preds.eq(labels.view_as(preds)).sum().item()
processed += len(images)
pbar.set_description(desc= f'Loss={loss.item()} Batch_id={batch_idx} Accuracy={100*correct/processed:0.2f}')
train_accuracy.append(100*correct/processed)
# Test Function
test_losses = [] # to capture test losses
test_accuracy = [] # to capture test accuracy
def test(model,device, test_loader):
model.eval() # setting the model in evaluation mode
test_loss = 0
correct = 0 # for accuracy numerator
with torch.no_grad():
for (images,labels) in test_loader:
images, labels = images.to(device),labels.to(device)#sending data to CPU or GPU as per device
outputs = model(images) # forward pass, result captured in outputs (plural as there are many images in a batch)
# the outputs are in batch size x one hot vector
test_loss = F.nll_loss(outputs,labels, reduction='sum').item() # sum up batch loss
preds = outputs.argmax(dim=1, keepdim=True) # get the index of the max log-probability
correct += preds.eq(labels.view_as(preds)).sum().item()
test_loss /= len(test_loader.dataset) # average test loss
test_losses.append(test_loss) # to capture loss over many batches
print('\nTest set: Average loss: {:.4f}, Accuracy: {}/{} ({:.2f}%)\n'.format(
test_loss, correct, len(test_loader.dataset),
100. * correct / len(test_loader.dataset)))
test_accuracy.append(100*correct/len(test_loader.dataset))
model = Network().to(device)
optimizer = optim.SGD(model.parameters(), lr=0.01, momentum=0.9)
# EPOCHS = 1
EPOCHS = 15
for epoch in range(EPOCHS):
print("EPOCH:", epoch)
train(model, device, train_loader, optimizer, epoch)
test(model, device, test_loader)
# Graphs
fig, axs = plt.subplots(2,2,figsize=(15,10))
axs[0, 0].plot(train_losses)
axs[0, 0].set_title("Training Loss")
axs[1, 0].plot(train_accuracy)
axs[1, 0].set_title("Training Accuracy")
axs[0, 1].plot(test_losses)
axs[0, 1].set_title("Test Loss")
axs[1, 1].plot(test_accuracy)
axs[1, 1].set_title("Test Accuracy")
```
### Conclusion
Experiment Number : 2
Objective / Target
1. Reduce params (number of channels are reduced, use of GAP)
2. Reduce Overfitting (used dropout, and BN)
Results
- Parameters: 4,038
- Best Train Accuracy: 97.84 %
- Best Test Accuracy: 98.46 %
Analysis
1. Small Model, but not hitting the accuracy mark
2. Overfitting (train - test accuracy) < 0 is largely containted
Next Steps
- Look to increase accuracy by increasing number of params
| github_jupyter |
<table align="left" width="100%"> <tr>
<td style="background-color:#ffffff;"><a href="https://qsoftware.lu.lv/index.php/qworld/" target="_blank"><img src="../images/qworld.jpg" width="35%" align="left"></a></td>
<td align="right" style="background-color:#ffffff;vertical-align:bottom;horizontal-align:right">
prepared by Özlem Salehi (<a href="http://qworld.lu.lv/index.php/qturkey/" target="_blank">QTurkey</a>)
</td>
</tr></table>
<table width="100%"><tr><td style="color:#bbbbbb;background-color:#ffffff;font-size:11px;font-style:italic;text-align:right;">This cell contains some macros. If there is a problem with displaying mathematical formulas, please run this cell to load these macros. </td></tr></table>
$ \newcommand{\bra}[1]{\langle #1|} $
$ \newcommand{\ket}[1]{|#1\rangle} $
$ \newcommand{\braket}[2]{\langle #1|#2\rangle} $
$ \newcommand{\dot}[2]{ #1 \cdot #2} $
$ \newcommand{\biginner}[2]{\left\langle #1,#2\right\rangle} $
$ \newcommand{\mymatrix}[2]{\left( \begin{array}{#1} #2\end{array} \right)} $
$ \newcommand{\myvector}[1]{\mymatrix{c}{#1}} $
$ \newcommand{\myrvector}[1]{\mymatrix{r}{#1}} $
$ \newcommand{\mypar}[1]{\left( #1 \right)} $
$ \newcommand{\mybigpar}[1]{ \Big( #1 \Big)} $
$ \newcommand{\sqrttwo}{\frac{1}{\sqrt{2}}} $
$ \newcommand{\dsqrttwo}{\dfrac{1}{\sqrt{2}}} $
$ \newcommand{\onehalf}{\frac{1}{2}} $
$ \newcommand{\donehalf}{\dfrac{1}{2}} $
$ \newcommand{\hadamard}{ \mymatrix{rr}{ \sqrttwo & \sqrttwo \\ \sqrttwo & -\sqrttwo }} $
$ \newcommand{\vzero}{\myvector{1\\0}} $
$ \newcommand{\vone}{\myvector{0\\1}} $
$ \newcommand{\stateplus}{\myvector{ \sqrttwo \\ \sqrttwo } } $
$ \newcommand{\stateminus}{ \myrvector{ \sqrttwo \\ -\sqrttwo } } $
$ \newcommand{\myarray}[2]{ \begin{array}{#1}#2\end{array}} $
$ \newcommand{\X}{ \mymatrix{cc}{0 & 1 \\ 1 & 0} } $
$ \newcommand{\Z}{ \mymatrix{rr}{1 & 0 \\ 0 & -1} } $
$ \newcommand{\Htwo}{ \mymatrix{rrrr}{ \frac{1}{2} & \frac{1}{2} & \frac{1}{2} & \frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & \frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} & \frac{1}{2} } } $
$ \newcommand{\CNOT}{ \mymatrix{cccc}{1 & 0 & 0 & 0 \\ 0 & 1 & 0 & 0 \\ 0 & 0 & 0 & 1 \\ 0 & 0 & 1 & 0} } $
$ \newcommand{\norm}[1]{ \left\lVert #1 \right\rVert } $
$ \newcommand{\pstate}[1]{ \lceil \mspace{-1mu} #1 \mspace{-1.5mu} \rfloor } $
<h1> <font color="blue"> Solutions for </font> Phase Estimation</h1>
<h3>Task 1 (on paper)</h3>
<a id="task1"></a>
Show that $\ket{-}$ and $\ket{+}$ are eigenvectors for the $X$ operator with eigenvalues -1 and 1 respectively.
<h3>Solution </h3>
\begin{align*}
X \ket{-} = X \frac {\ket{0} - \ket{1}} {\sqrt{2}}
& = \frac {\ket{1} - \ket{0}} {\sqrt{2}}\\
& = -\frac {\ket{0} - \ket{1}} {\sqrt{2}}\\
& = -\ket{-}
\end{align*}
\begin{align*}
X \ket{+} = X \frac {\ket{0} + \ket{1}} {\sqrt{2}}
& = \frac {\ket{1} + \ket{0}} {\sqrt{2}}\\
& = \frac {\ket{0} + \ket{1}} {\sqrt{2}}\\
& = \ket{+}
\end{align*}
Hence $ \ket{-}$ and $\ket{+}$ are eigenvectors of $ X $ operator with the eigenvalues -1 and 1 respectively.
<h3>Task 2 (on paper)</h3>
<a id="task2"></a>
Consider the following quantum state where $ x=0 $ or $ 1 $. How can you find out the value of $x$ by applying a single operator?
$$
\frac {\ket{0} + (-1)^x \ket{1}} {\sqrt{2}}
$$
<h3>Solution </h3>
If our aim is to find out $ x $, then applying a Hadamard gate we can determine whether $ x=0 $ or $ 1 $ depending on the outcome. If the outcome is $ \ket{0} $, then $ x=0 $ and if the outcome is $ \ket{1} $, then $ x=1 $.
<hr>
```
from qiskit import QuantumCircuit, QuantumRegister, ClassicalRegister, Aer, execute
from qiskit.circuit.library import PhaseGate
from math import pi
qasm_simulator = Aer.get_backend('qasm_simulator')
```
### Task 3
Consider the unitary operator $U$ with eigenvector $\ket{1}$ and eigenvalue $e^{2\pi i \phi}$ where $\phi = \frac{5}{16}$. $CU$ operator can be realized via a `PhaseGate` gate. Pick $t=4$ and implement the phase estimation circuit to estimate $\phi$. Write a function named <i>qpe</i> which takes as parameters <i>t</i>, <i>control</i> qubits, <i>target</i> qubits, <i>circuit</i> and the operator <i>CU</i>.
Note that you will get an exact result since $t=4$ is precise enough.
_Remarks:_
- You can apply arithmetical operations to gates, such as taking powers or inverses.
- To apply $QFT^{\dagger}$, you can use the function `iQFT` from our `silver` module. Create a `iQFT` operator by providing the number of qubits, and apply it like any other gate provided by `qiskit.circuit.library`.
- To create a controlled $U^{2^k}$ gate from any $U$ in `qiskit.circuit.library`, you can use the `CUPowFactory` function in our module.
<center><pre>
──────■──────
┌───┐ ┌────┴────┐
─┤ U ├─ to ─┤ U^(2^k) ├─
└───┘ └─────────┘
</pre></center>
```
from silver import CUPowFactory
CPhasePowGate = CUPowFactory(PhaseGate(2*pi*5/16)) # Do not forget the 2π factor
```
<h3>Solution </h3>
```
from silver import CUPowFactory
CPhasePowGate = CUPowFactory(PhaseGate(2*pi*5/16)) # Do not forget the 2π factor
from silver import iQFT
def qpe(circuit, CU, control, target):
t = control.size
# Go over each control qubit in the reversed
# order
for i, c in enumerate(reversed(control)):
# Apply Hadamard
circuit.h(c)
# Apply
circuit.append(CU(i) ,[c, *target])
#Apply inverse QFT to the control qubits
iqft_gate = iQFT(len(control))
circuit.append(iqft_gate, control)
CONTROL_QUBIT_NUM = 4
# Create a cirucit
circuit = QuantumCircuit(
control_register := QuantumRegister(CONTROL_QUBIT_NUM),
target_register := QuantumRegister(1),
classical_register := ClassicalRegister(CONTROL_QUBIT_NUM),
)
# Set target qubit to store eigenvector
for q in target_register:
circuit.x(q)
circuit.barrier()
# Apply QPE
qpe(circuit, CPhasePowGate, control_register, target_register)
# Mesaure and print the results
circuit.measure(control_register, classical_register)
counts = execute(circuit.reverse_bits(), qasm_simulator) \
.result() \
.get_counts()
for i, j in counts.items():
print(f'φ = {int(i,2)}/{2**CONTROL_QUBIT_NUM} observerd {j} times')
# Select the maximum outcome as the current estimation
max_key = max(counts.keys(), key=lambda key: counts[key])
print(f'Estimation is {int(max_key, 2) / 2**CONTROL_QUBIT_NUM}')
```
<a id="task4"></a>
<h3>Task 4</h3>
Try Task 3 this time by setting $t=3$. What do you expect to see?
<h3>Solution </h3>
```
CONTROL_QUBIT_NUM = 3
# Create a cirucit
circuit = QuantumCircuit(
control_register := QuantumRegister(CONTROL_QUBIT_NUM),
target_register := QuantumRegister(1),
classical_register := ClassicalRegister(CONTROL_QUBIT_NUM),
)
# Set target qubit to store eigenvector
for q in target_register:
circuit.x(q)
circuit.barrier()
# Apply QPE
qpe(circuit, CPhasePowGate, control_register, target_register)
# Mesaure and print the results
circuit.measure(control_register, classical_register)
counts = execute(circuit.reverse_bits(), qasm_simulator) \
.result() \
.get_counts()
for i, j in counts.items():
print(f'{int(i,2)} observerd {j} times')
# Select the maximum outcome as the current estimation
max_key = max(counts.keys(), key=lambda key: counts[key])
print(f'\nEstimation is {int(max_key, 2) / 2**CONTROL_QUBIT_NUM}')
```
<a id="task5"></a>
<h3>Task 5</h3>
Try Task 3 this time by setting $\phi=0.685$. Calculate and print the estimates for different $t$ values between 1 and 10.
<h3>Solution </h3>
```
from silver import CUPowFactory
from qiskit.circuit.library import PhaseGate
# Create CUPow operator with phase 0.685
CPhasePowGate = CUPowFactory(PhaseGate(2*pi*0.685)) # Do not forget the 2π factor
for t in range (1,10):
# Create a cirucit
circuit = QuantumCircuit(
control_register := QuantumRegister(t),
target_register := QuantumRegister(1),
classical_register := ClassicalRegister(t),
)
# Set target qubit to store eigenvector
for q in target_register:
circuit.x(q)
circuit.barrier()
# Apply QPE
qpe(circuit, CPhasePowGate, control_register, target_register)
# Mesaure and print the results
circuit.measure(control_register, classical_register)
counts = execute(circuit.reverse_bits(), qasm_simulator) \
.result() \
.get_counts()
# Select the maximum outcome as the current estimation
max_key = max(counts.keys(), key=lambda key: counts[key])
print(f'Estimation for t={t} is {int(max_key, 2)/2**t}')
```
<a id="task6"></a>
<h3>Task 6</h3>
You are given an operator $U$ with eigenvector $\ket{11}$ and eigenvalue $\phi$, which is this time unknown to you. Use phase estimation to estimate $\phi$ by trying different $t$ values.
Run the following cell to load operator $U$
```
from silver import CUPowFactory
from silver.phaseEstimation import U
misteryGate = CUPowFactory(U)
```
<h3>Solution </h3>
```
for t in range (1,10):
# Create a cirucit
circuit = QuantumCircuit(
control_register := QuantumRegister(t),
target_register := QuantumRegister(2),
classical_register := ClassicalRegister(t),
)
# Set target qubit to store eigenvector
for q in target_register:
circuit.x(q)
circuit.barrier()
# Apply QPE
qpe(circuit, misteryGate, control_register, target_register)
# Mesaure and print the results
circuit.measure(control_register, classical_register)
counts = execute(circuit.reverse_bits(), qasm_simulator) \
.result() \
.get_counts()
max_key = max(counts.keys(), key=lambda key: counts[key])
print(f'Estimation for t={t} is {int(max_key, 2)/2**t}')
```
| github_jupyter |
# Programming with Python
## Episode 1a - Introduction - Analysing Patient Data
Teaching: 60 min,
Exercises: 30 min
Objectives
- Assign values to variables.
- Explain what a library is and what libraries are used for.
- Import a Python library and use the functions it contains.
- Read tabular data from a file into a program.
- Select individual values and subsections from data.
- Perform operations on arrays of data.
## Our Dataset
In this episode we will learn how to work with CSV files in Python. Our dataset contains patient inflammation data - where each row represents a different patient and the column represent inflammation data over a series of days.

However, before we discuss how to deal with many data points, let's learn how to work with single data values.
## Variables
Any Python interpreter can be used as a calculator:
```
3 + 5 * 4
```
```
3 + 5 * 4
```
This is great but not very interesting. To do anything useful with data, we need to assign its value to a variable. In Python, we can assign a value to a variable, using the equals sign ``=``. For example, to assign value 60 to a variable ``weight_kg``, we would execute:
```
weight_kg = 60
```
```
weight_kg = 60
```
From now on, whenever we use ``weight_kg``, Python will substitute the value we assigned to it. In essence, a variable is just a name for a value.
```
weight_kg + 5
```
```
weight_kg + 5
```
In Python, variable names:
- can include letters, digits, and underscores - `A-z, a-z, _`
- cannot start with a digit
- are case sensitive.
This means that, for example:
`weight0` is a valid variable name, whereas `0weight` is not
`weight` and `Weight` are different variables
#### Types of data
Python knows various types of data. Three common ones are:
- integer numbers (whole numbers)
- floating point numbers (numbers with a decimal point)
- and strings (of characters).
In the example above, variable `weight_kg` has an integer value of `60`. To create a variable with a floating point value, we can execute:
```
weight_kg = 60.0
```
```
weight_kg = 60.0
```
And to create a string we simply have to add single or double quotes around some text, for example:
```
weight_kg_text = 'weight in kilograms:'
```
To display the value of a variable to the screen in Python, we can use the print function:
```
print(weight_kg)
```
```
print(weight_kg)
```
We can display multiple things at once using only one print command:
```
print(weight_kg_text, weight_kg)
```
```
weight_kg_text = "weight in kilograms:"
print (weight_kg_text, weight_kg)
```
Moreover, we can do arithmetic with variables right inside the print function:
```
print('weight in pounds:', 2.2 * weight_kg)
```
```
print ("weight in ponds:", 2.2* weight_kg)
```
The above command, however, did not change the value of ``weight_kg``:
```
print(weight_kg)
```
```
print(weight_kg)
```
To change the value of the ``weight_kg`` variable, we have to assign `weight_kg` a new value using the equals `=` sign:
```
weight_kg = 65.0
print('weight in kilograms is now:', weight_kg)
```
```
weight_kg = 65.0
print('weight in kilo is now', weight_kg)
print(weight_kg)
```
#### Variables as Sticky Notes
A variable is analogous to a sticky note with a name written on it: assigning a value to a variable is like writing a value on the sticky note with a particular name.
This means that assigning a value to one variable does not change values of other variables (or sticky notes). For example, let's store the subject's weight in pounds in its own variable:
```
# There are 2.2 pounds per kilogram
weight_lb = 2.2 * weight_kg
print(weight_kg_text, weight_kg, 'and in pounds:', weight_lb)
```
```
# There are 2.2 pounds per kilogram
weight_lb = 2.2 * weight_kg
print(weight_kg_text, weight_kg, 'and in pounds:', weight_lb)
```
#### Updating a Variable
Variables calculated from other variables do not change their value just because the original variable changed its value (unlike cells in Excel):
```
weight_kg = 100.0
print('weight in kilograms is now:', weight_kg, 'and weight in pounds is still:', weight_lb)
```
```
weight_kg = 100.0
print('weight in kilograms is now:', weight_kg, 'and weight in pounds is:', weight_lb)
```
Since `weight_lb` doesn't *remember* where its value comes from, so it is not updated when we change `weight_kg`.
## Libraries
Words are useful, but what's more useful are the sentences and stories we build with them (or indeed entire books or whole libraries). Similarly, while a lot of powerful, general tools are built into Python, specialised tools built up from these basic units live in *libraries* that can be called upon when needed.
### Loading data into Python
In order to load our inflammation dataset into Python, we need to access (import in Python terminology) a library called `NumPy` (which stands for Numerical Python).
In general you should use this library if you want to do fancy things with numbers, especially if you have matrices or arrays. We can import `NumPy` using:
```
import numpy
```
```
import numpy
```
Importing a library is like getting a piece of lab equipment out of a storage locker and setting it up on the bench. Libraries provide additional functionality to the basic Python package, much like a new piece of equipment adds functionality to a lab space. Just like in the lab, importing too many libraries can sometimes complicate and slow down your programs - so we only import what we need for each program. Once we've imported the library, we can ask the library to read our data file for us:
```
numpy.loadtxt(fname='data/inflammation-01.csv', delimiter=',')
```
The expression `numpy.loadtxt(...)` is a function call that asks Python to run the function `loadtxt` which belongs to the `numpy` library. This dot `.` notation is used everywhere in Python: the thing that appears before the dot contains the thing that appears after.
As an example, John Smith is the John that belongs to the Smith family. We could use the dot notation to write his name smith.john, just as `loadtxt` is a function that belongs to the `numpy` library.
`numpy.loadtxt` has two parameters: the name of the file we want to read and the delimiter that separates values on a line. These both need to be character strings (or strings for short), so we put them in quotes.
Since we haven't told it to do anything else with the function's output, the notebook displays it. In this case, that output is the data we just loaded. By default, only a few rows and columns are shown (with ... to omit elements when displaying big arrays). To save space, Python displays numbers as 1. instead of 1.0 when there's nothing interesting after the decimal point.
Our call to `numpy.loadtxt` read our file but didn't save the data in memory. To do that, we need to assign the array to a variable. Just as we can assign a single value to a variable, we can also assign an array of values to a variable using the same syntax. Let's re-run `numpy.loadtxt` and save the returned data:
```
data = numpy.loadtxt(fname='data/inflammation-01.csv', delimiter=',')
```
```
data = numpy.loadtxt(fname='data/inflammation-01.csv', delimiter=',')
```
This statement doesn't produce any output because we've assigned the output to the variable `data`. If we want to check that the data has been loaded, we can print the variable's value:
```
print(data)
```
```
print(data)
```
Now that the data is in memory, we can manipulate it. First, let's ask Python what type of thing `data` refers to:
```
print(type(data))
```
```
print(type(data))
```
The output tells us that data currently refers to an N-dimensional array, the functionality for which is provided by the `NumPy` library. This data correspond to arthritis patients' inflammation. The rows are the individual patients, and the columns are their daily inflammation measurements.
#### Data Type
A NumPy array contains one or more elements of the same type. The type function will only tell you that a variable is a NumPy array but won't tell you the type of thing inside the array. We can find out the type of the data contained in the NumPy array.
```
print(data.dtype)
```
```
print(data.dtype)
data.dtype, data.shape, data.size
```
This tells us that the NumPy array's elements are floating-point numbers.
With the following command, we can see the array's shape:
```
print(data.shape)
```
```
print(data.shape)
```
The output tells us that the data array variable contains 60 rows and 40 columns. When we created the variable data to store our arthritis data, we didn't just create the array; we also created information about the array, called members or attributes. This extra information describes data in the same way an adjective describes a noun. data.shape is an attribute of data which describes the dimensions of data. We use the same dotted notation for the attributes of variables that we use for the functions in libraries because they have the same part-and-whole relationship.
If we want to get a single number from the array, we must provide an index in square brackets after the variable name, just as we do in math when referring to an element of a matrix. Our inflammation data has two dimensions, so we will need to use two indices to refer to one specific value:
```
print('first value in data:', data[0, 0])
print('middle value in data:', data[30, 20])
```
The expression `data[30, 20]` accesses the element at row 30, column 20. While this expression may not surprise you, `data[0, 0]` might.
#### Zero Indexing
Programming languages like Fortran, MATLAB and R start counting at 1 because that's what human beings have done for thousands of years. Languages in the C family (including C++, Java, Perl, and Python) count from 0 because it represents an offset from the first value in the array (the second value is offset by one index from the first value). This is closer to the way that computers represent arrays (if you are interested in the historical reasons behind counting indices from zero, you can read Mike Hoye's blog post).
As a result, if we have an M×N array in Python, its indices go from 0 to M-1 on the first axis and 0 to N-1 on the second. It takes a bit of getting used to, but one way to remember the rule is that the index is how many steps we have to take from the start to get the item we want.
#### In the Corner
What may also surprise you is that when Python displays an array, it shows the element with index `[0, 0]` in the upper left corner rather than the lower left. This is consistent with the way mathematicians draw matrices but different from the Cartesian coordinates. The indices are (row, column) instead of (column, row) for the same reason, which can be confusing when plotting data.
#### Slicing data
An index like `[30, 20]` selects a single element of an array, but we can select whole sections as well. For example, we can select the first ten days (columns) of values for the first four patients (rows) like this:
```
print(data[0:4, 0:10])
```
```
print(data[0:4, 0:10])
```
The slice `[0:4]` means, *Start at index 0 and go up to, but not including, index 4*.
Again, the up-to-but-not-including takes a bit of getting used to, but the rule is that the difference between the upper and lower bounds is the number of values in the slice.
Also, we don't have to start slices at `0`:
```
print(data[5:10, 0:10])
```
and we don't have to include the upper or lower bound on the slice.
If we don't include the lower bound, Python uses 0 by default; if we don't include the upper, the slice runs to the end of the axis, and if we don't include either (i.e., if we just use `:` on its own), the slice includes everything:
```
small = data[:3, 36:]
print('small is:')
print(small)
```
The above example selects rows 0 through 2 and columns 36 through to the end of the array.
thus small is:
```
[[ 2. 3. 0. 0.]
[ 1. 1. 0. 1.]
[ 2. 2. 1. 1.]]
```
Arrays also know how to perform common mathematical operations on their values. The simplest operations with data are arithmetic: addition, subtraction, multiplication, and division. When you do such operations on arrays, the operation is done element-by-element. Thus:
```
doubledata = data * 2.0
```
will create a new array doubledata each element of which is twice the value of the corresponding element in data:
```
print('original:')
print(data[:3, 36:])
print('doubledata:')
print(doubledata[:3, 36:])
```
If, instead of taking an array and doing arithmetic with a single value (as above), you did the arithmetic operation with another array of the same shape, the operation will be done on corresponding elements of the two arrays. Thus:
```
tripledata = doubledata + data
```
will give you an array where `tripledata[0,0]` will equal `doubledata[0,0]` plus `data[0,0]`, and so on for all other elements of the arrays.
```
print('tripledata:')
print(tripledata[:3, 36:])
```
## Exercises
### Variables
What values do the variables mass and age have after each statement in the following program?
```
mass = 47.5
age = 122
mass = mass * 2.0
age = age - 20
print(mass, age)
```
Test your answers by executing the commands.
```
mass = 47.5
age = 122.0
mass = mass * 2.0
age = age - 20
print(mass, age)
```
Solution:
### Sorting Out References
What does the following program print out?
```
first, second = 'Grace', 'Hopper'
third, fourth = second, first
print(third, fourth)
```
```
first, second = 'Grace', 'Hopper'
third, fourth = second, first
print(third, fourth)
```
Solution:
### Slicing Strings
A section of an array is called a slice. We can take slices of character strings as well:
```
element = 'oxygen'
print('first three characters:', element[0:3])
print('last three characters:', element[3:6])
```
What is the value of `element[:4]` ? What about `element[4:]`? Or `element[:]` ?
What about `element[-1]` and `element[-2]` ?
```
element = 'oxygen'
print('first three characters:', element[0:3])
print('last three characters:', element[3:6])
print('up to :4:', element[:4])
print('4: and after', element[4:])
print('everything', element[:])
print('index -1:', element[-1])
print('index -2:', element[-2])
```
Solution:
Given those answers, explain what `element[1:-1]` does.
```
print('iddd2:', element[1:-1])
```
Solution:
### Thin Slices
The expression `element[3:3]` produces an empty string, i.e., a string that contains no characters. If data holds our array of patient data, what does `data[3:3, 4:4]` produce? What about `data[3:3, :]` ?
```
print(data[3:3], )
```
Solution:
## Key Points
Import a library into a program using import library_name.
Use the numpy library to work with arrays in Python.
Use `variable` `=` `value` to assign a value to a variable in order to record it in memory.
Variables are created on demand whenever a value is assigned to them.
Use `print(something)` to display the value of something.
The expression `array.shape` gives the shape of an array.
Use `array[x, y]` to select a single element from a 2D array.
Array indices start at 0, not 1.
Use `low:high` to specify a slice that includes the indices from low to high-1.
All the indexing and slicing that works on arrays also works on strings.
Use `#` and some kind of explanation to add comments to programs.
# Save, and version control your changes
- save your work: `File -> Save`
- add all your changes to your local repository: `Terminal -> git add .`
- commit your updates a new Git version: `Terminal -> git commit -m "End of Episode 1"`
- push your latest commits to GitHub: `Terminal -> git push`
| github_jupyter |
# Analyzer for individual Participant results
```
import pandas as pd
import math
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
sns.set() # Setting seaborn as default style
# declare global helpers
condition_values = ['0ms','10ms','400ms','10ms echo','400ms echo']
groupnames = [0,1,2,3,4]
#read in source csv
df = pd.read_csv (r'C:\Users\aron\Google Drive\BA\thesis\data_analysis\Database_AllParticipants_av_experiment_122522_2021-04-12_13h09.36_56b82ada-9b90-11eb-a6de-ac1f6b405aea.csv')
df.head()
#cut down df to individual participant
df = df[df['participant'] == 22]
df.shape
# Here we can see all tracked variables
df.info()
```
## General Info
```
df.describe()
list(df.columns)
exp = df['expName'].iloc[0]
print('Exp: ',exp)
os = df['OS'].iloc[0]
print('OS: ',os)
browser = df['browser'].iloc[0]
print('Browser: ',browser)
res = (df['xResolution'].iloc[0], df['yResolution'].iloc[0])
print('Resolution: ',res)
fr = df['frameRate'].iloc[0]
print('Framerate: ',browser)
eid = df['id'].iloc[0]
print('Experiment ID: ',eid)
pid = df['participant'].iloc[0]
print('Participant ID: ',pid)
date = df['date'].iloc[0]
print('Date: ',date)
group = df['group'].iloc[0]
print('Group: ',group)
switch = df['target_loc'].median() == 1
print('Images reversed: ',switch)
trials_ = df['trials.ran'].count()
```
# Analysis of choice trial
```
rt = df['choice_response.rt']
fig = plt.gcf()
# Change seaborn plot size
fig.set_size_inches(10, 8)
# Increase font size
sns.set(font_scale=1.5)
sns.histplot(rt, kde=True)
# Change Axis labels:
plt.xlabel('Response Time')
plt.ylabel('Occurrence')
plt.title('Distribution of Choice RT overall')
trials = df['trials.ran'].count()
trials
means_by_cond = df.groupby('cond')['choice_response.rt'].mean()
print(' Means over conditions: ', means_by_cond)
fig = plt.gcf()
# Change seaborn plot size
fig.set_size_inches(10, 8)
# Increase font size
sns.set(font_scale=1.5)
sns.violinplot(y='choice_response.rt',
x="cond",
data=df,
showmeans=True).set_xticklabels(condition_values)
# Change Axis labels:
plt.xlabel('Condition')
plt.ylabel('Response Time (MSec)')
plt.title('Response times over conditions in choice trial')
fig = plt.gcf()
# Change seaborn plot size
fig.set_size_inches(10, 8)
# Increase font size
sns.set(font_scale=1.5)
sns.boxplot(y='choice_response.rt', x="cond",
data=df,
showmeans=True).set_xticklabels(condition_values)
# Change Axis labels:
plt.xlabel('Condition')
plt.ylabel('Response Time (MSec)')
plt.title('Response times over different conditions')
fig = plt.gcf()
# Change seaborn plot size
fig.set_size_inches(10, 8)
# Increase font size
sns.set(font_scale=1.5)
sns.countplot(hue='choice_response.corr', x="cond",
data=df).set_xticklabels(condition_values)
# Change Axis labels:
plt.legend(loc = 'upper left')
plt.xlabel('Condition')
plt.ylabel('Count')
plt.title('Correct Responses over conditions in choice trial')
```
## Response Time over experiment Progress
```
fig = plt.gcf()
# Change seaborn plot size
fig.set_size_inches(10, 8)
# Increase font size
sns.set(font_scale=1.5)
sns.regplot(y='choice_response.rt',
x='trials.thisTrialN',
#hue = 'choice_response.corr',
data=df)
# Change Axis labels:
plt.legend(loc = 'upper left')
plt.xlabel('Choice Trials')
plt.ylabel('RT')
plt.title('RTs over all trials')
# Filter out correct responses
df_incorr = df.drop(df[df['choice_response.corr'] == 1].index)
df_incorr = df_incorr.drop(df_incorr[df_incorr['target_loc'] == 2].index)
df_incorr.shape
fig = plt.gcf()
# Change seaborn plot size
fig.set_size_inches(10, 8)
# Increase font size
sns.set(font_scale=1.5)
sns.regplot(y='choice_response.rt',
x='trials.thisTrialN',
#hue = 'choice_response.corr',
data=df)
# Change Axis labels:
plt.legend(loc = 'upper left')
plt.xlabel('Choice Trials')
plt.ylabel('RT')
plt.title('RTs over all incorrectly answered trials')
# Filter out incorrect responses
df_corr = df.drop(df[df['choice_response.corr'] == 0].index)
df_corr = df_corr.drop(df_corr[df_corr['target_loc'] == 2].index)
df_corr.shape
fig = plt.gcf()
# Change seaborn plot size
fig.set_size_inches(10, 8)
# Increase font size
sns.set(font_scale=1.5)
sns.regplot(y='choice_response.rt',
x='trials.thisTrialN',
#hue = 'choice_response.corr',
data=df_corr)
# Change Axis labels:
plt.legend(loc = 'upper left')
plt.xlabel('Choice Trials')
plt.ylabel('RT')
plt.title('RTs over correctly answered non-filler trials')
# Filter out incorrect responses
df_filler = df.drop(df[df['target_loc'] < 2].index)
df_filler['target_loc'].unique()
fig = plt.gcf()
# Change seaborn plot size
fig.set_size_inches(10, 8)
# Increase font size
sns.set(font_scale=1.5)
sns.regplot(y='choice_response.rt',
x='trials.thisTrialN',
#hue = 'choice_response.corr',
data=df_filler)
# Change Axis labels:
plt.legend(loc = 'upper left')
plt.xlabel('Choice Trials')
plt.ylabel('RT')
plt.title('RTs over filler trials')
fig = plt.gcf()
# Change seaborn plot size
fig.set_size_inches(10, 8)
# Increase font size
sns.set(font_scale=1.5)
sns.countplot(hue='choice_response.corr', x="cond",
data=df_filler).set_xticklabels(condition_values)
# Change Axis labels:
plt.legend(loc = 'upper left')
plt.xlabel('Condition')
plt.ylabel('Count')
plt.title('Correct Responses over conditions in filler trials')
fig = plt.gcf()
# Change seaborn plot size
fig.set_size_inches(10, 8)
# Increase font size
sns.set(font_scale=1.5)
sns.relplot(y='choice_response.rt', x='trials.thisTrialN', hue="choice_response.corr", data=df);
# Change Axis labels:
plt.xlabel('Choice Trials')
plt.ylabel('RT')
plt.title('Correct / Incorrect RTs over all trials')
```
# Analysis of SJ task
```
fig, axes = plt.subplots(1, 2)
# Change seaborn plot size
fig.set_size_inches(16, 8)
# Increase font size
sns.set(font_scale=1.5)
fig.suptitle('Correct Responses In SJ Task')
axes[0].set_title('Synchrony Question')
axes[1].set_title('Distortion Question')
sns.countplot(ax=axes[0],hue='response_sync.corr', x="cond",data=df).set_xticklabels(condition_values)
sns.countplot(ax=axes[1],hue='response_distortion.corr', x="cond",data=df).set_xticklabels(condition_values)
# Change Axis labels:
axes[0].legend(loc = 'upper left')
axes[1].legend(loc = 'upper right')
fig, axes = plt.subplots(3, 1)
# Change seaborn plot size
fig.set_size_inches(10, 8)
# Increase font size
sns.set(font_scale=1.5)
sns.regplot(y='response_sync.corr',
x = 'SJ_trials_before.thisTrialN',
data=df,
truncate = True,
#logistic = True,
ax = axes[0])
sns.regplot(y='response_sync.corr',
x = 'SJ_trials_between.thisTrialN',
data=df,
truncate = True,
#logistic = True,
ax = axes[1])
sns.regplot(y='response_sync.corr',
x = 'SJ_trials_after.thisTrialN',
data=df,
truncate = True,
#logistic = True,
ax = axes[2])
# Change Axis labels:
axes[0].set_title('Block 1')
axes[1].set_title('Block 2')
axes[2].set_title('Block 3')
plt.suptitle('Correctness of SJ synchrony answers over trials')
```
| github_jupyter |
# Image Classification Neo Compilation Example - Local Mode
This notebook shows an intermediate step in the process of developing an Edge image classification algorithm.
## Notebook Setup
```
%matplotlib inline
import time
import os
#os.environ["CUDA_VISIBLE_DEVICES"]="0"
import tensorflow as tf
import numpy as np
import cv2
from tensorflow.python.client import device_lib
tf.__version__
```
## Helper Functions and scripts
```
def get_img(img_path):
img = cv2.imread(img_path)
img = cv2.resize(img.astype(float), (224, 224)) #resize
imgMean = np.array([104, 117, 124], np.float)
img -= imgMean #subtract image mean
return img
def get_label_encode(class_id):
enc = np.zeros((257),dtype=int)
enc[class_id] = 1
return enc
def freeze_graph(model_dir, output_node_names):
"""Extract the sub graph defined by the output nodes and convert
all its variables into constant
Args:
model_dir: the root folder containing the checkpoint state file
output_node_names: a string, containing all the output node's names,
comma separated
"""
if not tf.gfile.Exists(model_dir):
raise AssertionError(
"Export directory doesn't exists. Please specify an export "
"directory: %s" % model_dir)
if not output_node_names:
print("You need to supply the name of a node to --output_node_names.")
return -1
# We retrieve our checkpoint fullpath
checkpoint = tf.train.get_checkpoint_state(model_dir)
input_checkpoint = checkpoint.model_checkpoint_path
# We precise the file fullname of our freezed graph
absolute_model_dir = "/".join(input_checkpoint.split('/')[:-1])
output_graph = absolute_model_dir + "/frozen_model.pb"
# We clear devices to allow TensorFlow to control on which device it will load operations
clear_devices = True
# We start a session using a temporary fresh Graph
with tf.Session(graph=tf.Graph()) as sess:
# We import the meta graph in the current default Graph
saver = tf.train.import_meta_graph(input_checkpoint + '.meta', clear_devices=clear_devices)
# We restore the weights
saver.restore(sess, input_checkpoint)
# We use a built-in TF helper to export variables to constants
output_graph_def = tf.graph_util.convert_variables_to_constants(
sess, # The session is used to retrieve the weights
tf.get_default_graph().as_graph_def(), # The graph_def is used to retrieve the nodes
output_node_names.split(",") # The output node names are used to select the usefull nodes
)
# Finally we serialize and dump the output graph to the filesystem
with tf.gfile.GFile(output_graph, "wb") as f:
f.write(output_graph_def.SerializeToString())
print("%d ops in the final graph." % len(output_graph_def.node))
return output_graph_def
class VGG(object):
"""alexNet model"""
def __init__(self, n_classes, batch_size=None):
self.NUM_CLASSES = n_classes
self.BATCH_SIZE = batch_size
self.x = tf.placeholder(tf.float32, [None, 224, 224, 3])
self.y = tf.placeholder(tf.float32, [None, self.NUM_CLASSES])
self.buildCNN()
def buildCNN(self):
"""build model"""
input_layer = self.x
conv1_1 = tf.layers.conv2d(
input_layer, filters=64, kernel_size=(3, 3), strides=(1, 1), activation=tf.nn.relu, padding='SAME', name='conv1_1'
)
conv1_2 = tf.layers.conv2d(
conv1_1, filters=64, kernel_size=(3, 3), strides=(1, 1), activation=tf.nn.relu, padding='SAME', name='conv1_2'
)
pool1 = tf.layers.max_pooling2d(conv1_2, pool_size=(2, 2), strides=(2, 2), padding='SAME', name='pool1')
# conv2
conv2_1 = tf.layers.conv2d(
pool1, filters=128, kernel_size=(3, 3), strides=(1, 1), activation=tf.nn.relu, padding='SAME', name='conv2_1'
)
conv2_2 = tf.layers.conv2d(
conv2_1, filters=128, kernel_size=(3, 3), strides=(1, 1), activation=tf.nn.relu, padding='SAME', name='conv2_2'
)
pool2 = tf.layers.max_pooling2d(conv2_2, pool_size=(2, 2), strides=(2, 2), padding='SAME', name='pool2')
# conv3
conv3_1 = tf.layers.conv2d(
pool2, filters=256, kernel_size=(3, 3), strides=(1, 1), activation=tf.nn.relu, padding='SAME', name='conv3_1'
)
conv3_2 = tf.layers.conv2d(
conv3_1, filters=256, kernel_size=(3, 3), strides=(1, 1), activation=tf.nn.relu, padding='SAME', name='conv3_2'
)
conv3_3 = tf.layers.conv2d(
conv3_2, filters=256, kernel_size=(3, 3), strides=(1, 1), activation=tf.nn.relu, padding='SAME', name='conv3_3'
)
pool3 = tf.layers.max_pooling2d(conv3_3, pool_size=(2, 2), strides=(2, 2), padding='SAME', name='pool3')
# conv4
conv4_1 = tf.layers.conv2d(
pool3, filters=512, kernel_size=(3, 3), strides=(1, 1), activation=tf.nn.relu, padding='SAME', name='conv4_1'
)
conv4_2 = tf.layers.conv2d(
conv4_1, filters=512, kernel_size=(3, 3), strides=(1, 1), activation=tf.nn.relu, padding='SAME', name='conv4_2'
)
conv4_3 = tf.layers.conv2d(
conv4_2, filters=512, kernel_size=(3, 3), strides=(1, 1), activation=tf.nn.relu, padding='SAME', name='conv4_3'
)
pool4 = tf.layers.max_pooling2d(conv4_3, pool_size=(2, 2), strides=(2, 2), padding='SAME', name='pool4')
# conv5
conv5_1 = tf.layers.conv2d(
pool4, filters=512, kernel_size=(3, 3), strides=(1, 1), activation=tf.nn.relu, padding='SAME', name='conv5_1'
)
conv5_2 = tf.layers.conv2d(
conv5_1, filters=512, kernel_size=(3, 3), strides=(1, 1), activation=tf.nn.relu, padding='SAME', name='conv5_2'
)
conv5_3 = tf.layers.conv2d(
conv5_2, filters=512, kernel_size=(3, 3), strides=(1, 1), activation=tf.nn.relu, padding='SAME', name='conv5_3'
)
pool5 = tf.layers.max_pooling2d(conv5_3, pool_size=(2, 2), strides=(2, 2), padding='SAME', name='pool5')
#print('POOL5', pool5.shape)
#CULPADO flatten = tf.layers.flatten(pool5, name='flatten')
flatten = tf.reshape(pool5, [-1, 7*7*512])
fc1_relu = tf.layers.dense(flatten, units=4096, activation=tf.nn.relu, name='fc1_relu')
fc2_relu = tf.layers.dense(fc1_relu, units=4096, activation=tf.nn.relu, name='fc2_relu')
self.logits = tf.layers.dense(fc2_relu, units=self.NUM_CLASSES, name='fc3_relu')
#fc3 = tf.nn.softmax(logits)
print(self.logits.shape)
# Return the complete AlexNet model
return self.logits
net = VGG(257)
net.x, net.y, net.logits
```
## Dataset
The example is no longer on this URL. Now it has to be downloaded from Google Drive: https://drive.google.com/uc?id=1r6o0pSROcV1_VwT4oSjA2FBUSCWGuxLK
```
!conda install -c conda-forge -y gdown
!mkdir -p ../data/Caltech256
import gdown
url = "https://drive.google.com/uc?id=1r6o0pSROcV1_VwT4oSjA2FBUSCWGuxLK"
destination = "../data/Caltech256/256_ObjectCategories.tar"
gdown.download(url, destination, quiet=False)
#!curl http://www.vision.caltech.edu/Image_Datasets/Caltech256/256_ObjectCategories.tar -o ../data/Caltech256/256_ObjectCategories.tar
%%sh
pushd ../data/Caltech256/
tar -xvf 256_ObjectCategories.tar
popd
import glob
import random
from sklearn.model_selection import train_test_split
counter = 0
classes = {}
all_image_paths = list(glob.glob('../data/Caltech256/256_ObjectCategories/*/*.jpg'))
x = []
y = []
for i in all_image_paths:
_,cat,fname = i.split('/')[3:]
if classes.get(cat) is None:
classes[cat] = counter
counter += 1
x.append(i)
y.append(classes.get(cat))
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size=0.33, random_state=42)
```
## Training
```
# (6) Define model's cost and optimizer
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits_v2(logits=net.logits, labels=net.y))
optimizer = tf.train.AdamOptimizer().minimize(cost)
# (7) Defining evaluation metrics
correct_prediction = tf.equal(tf.argmax(net.logits, 1), tf.argmax(net.y, 1))
accuracy_pct = tf.reduce_mean(tf.cast(correct_prediction, tf.float32)) * 100
# (8) initialize
initializer_op = tf.global_variables_initializer()
epochs = 1
batch_size = 128
test_batch_size = 256
# (9) Run
with tf.Session() as session:
session.run(initializer_op)
print("Training for", epochs, "epochs.")
# looping over epochs:
for epoch in range(epochs):
# To monitor performance during training
avg_cost = 0.0
avg_acc_pct = 0.0
# loop over all batches of the epoch- 1088 records
# batch_size = 128 is already defined
n_batches = int(len(x_train) / batch_size)
counter = 1
for i in range(n_batches):
# Get the random int for batch
#random_indices = np.random.randint(len(x_train), size=batch_size) # 1088 is the no of training set records
pivot = i * batch_size
feed = {
net.x: [get_img(i) for i in x_train[pivot:pivot+batch_size]],
net.y: [get_label_encode(i) for i in y_train[pivot:pivot+batch_size]]
}
# feed batch data to run optimization and fetching cost and accuracy:
_, batch_cost, batch_acc = session.run([optimizer, cost, accuracy_pct],
feed_dict=feed)
# Print batch cost to see the code is working (optional)
# print('Batch no. {}: batch_cost: {}, batch_acc: {}'.format(counter, batch_cost, batch_acc))
# Get the average cost and accuracy for all batches:
avg_cost += batch_cost / n_batches
avg_acc_pct += batch_acc / n_batches
counter += 1
if counter % 50 == 0:
print("Batch {}/{}: batch_cost={:.3f}, batch_acc={:.3f},avg_cost={:.3f},avg_acc={:.3f}".format(
i,n_batches, batch_cost, batch_acc, avg_cost, avg_acc_pct
))
# Get cost and accuracy after one iteration
test_cost = cost.eval({net.x: [get_img(i) for i in x_test[:test_batch_size]], net.y: [get_label_encode(i) for i in y_test[:test_batch_size]]})
test_acc_pct = accuracy_pct.eval({net.x: [get_img(i) for i in x_test[:test_batch_size]], net.y: [get_label_encode(i) for i in y_test[:test_batch_size]]})
# output logs at end of each epoch of training:
print("Epoch {}: Training Cost = {:.3f}, Training Acc = {:.2f} -- Test Cost = {:.3f}, Test Acc = {:.2f}"\
.format(epoch + 1, avg_cost, avg_acc_pct, test_cost, test_acc_pct))
# Getting Final Test Evaluation
print('\n')
print("Training Completed. Final Evaluation on Test Data Set.\n")
test_cost = cost.eval({net.x: [get_img(i) for i in x_test[:test_batch_size]], net.y: [get_label_encode(i) for i in y_test[:test_batch_size]]})
test_accy_pct = accuracy_pct.eval({net.x: [get_img(i) for i in x_test[:test_batch_size]], net.y: [get_label_encode(i) for i in y_test[:test_batch_size]]})
print("Test Cost:", '{:.3f}'.format(test_cost))
print("Test Accuracy: ", '{:.2f}'.format(test_accy_pct), "%", sep='')
print('\n')
# Getting accuracy on Validation set
val_cost = cost.eval({net.x: [get_img(i) for i in x_test[test_batch_size:test_batch_size*2]], net.y: [get_label_encode(i) for i in y_test[test_batch_size:test_batch_size*2]]})
val_acc_pct = accuracy_pct.eval({net.x: [get_img(i) for i in x_test[test_batch_size:test_batch_size*2]], net.y: [get_label_encode(i) for i in y_test[test_batch_size:test_batch_size*2]]})
print("Evaluation on Validation Data Set.\n")
print("Evaluation Cost:", '{:.3f}'.format(val_cost))
print("Evaluation Accuracy: ", '{:.2f}'.format(val_acc_pct), "%", sep='')
```
## Saving/exporting the model
```
!rm -rf vgg16/
exporter = tf.saved_model.builder.SavedModelBuilder('vgg16/model')
saver = tf.train.Saver()
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
img = get_img('TestImage.jpg')
batch = np.array([img for i in range(batch_size)]).reshape((batch_size,224,224,3))
print(batch.shape)
x = net.x
y = net.logits
start_time = time.time()
out = sess.run(y, feed_dict = {x: batch})
print("Elapsed time: {}".format(time.time() - start_time))
exporter.add_meta_graph_and_variables(
sess,
tags=[tf.saved_model.tag_constants.SERVING],
signature_def_map={
tf.saved_model.signature_constants.DEFAULT_SERVING_SIGNATURE_DEF_KEY:
tf.saved_model.signature_def_utils.predict_signature_def(
inputs={"inputs": net.x},
outputs={"outputs": net.logits }
)
},
strip_default_attrs=True)
#exporter.save()
saver.save(sess, 'vgg16/model/model')
_ = freeze_graph('vgg16/model', 'fc3_relu/BiasAdd')
!cd vgg16 && rm -f model.tar.gz && cd model && tar -czvf ../model.tar.gz frozen_model.pb
import time
import sagemaker
import os
import json
import boto3
# Retrieve the default bucket
sagemaker_session = sagemaker.Session()
default_bucket = sagemaker_session.default_bucket()
type(sagemaker_session)
role=sagemaker.session.get_execution_role()
job_prefix='VGG16'
path='neo/%s' % job_prefix
sm = boto3.client('sagemaker')
!aws s3 cp vgg16/model.tar.gz s3://$default_bucket/$path/model.tar.gz
target_device='ml_m5'
# 'lambda'|'ml_m4'|'ml_m5'|'ml_c4'|'ml_c5'|'ml_p2'|'ml_p3'|'ml_g4dn'|'ml_inf1'|'jetson_tx1'|'jetson_tx2'|'jetson_nano'|'jetson_xavier'|
# 'rasp3b'|'imx8qm'|'deeplens'|'rk3399'|'rk3288'|'aisage'|'sbe_c'|'qcs605'|'qcs603'|'sitara_am57x'|'amba_cv22'|'x86_win32'|'x86_win64'
job_name="%s-%d" % (job_prefix, int(time.time()))
sm.create_compilation_job(
CompilationJobName=job_name,
RoleArn=role,
InputConfig={
'S3Uri': "s3://%s/%s/model.tar.gz" % (default_bucket, path),
'DataInputConfig': '{"data":[1,224,224,3]}',
'Framework': 'TENSORFLOW'
},
OutputConfig={
'S3OutputLocation': "s3://%s/%s/" % (default_bucket, path),
'TargetDevice': target_device
},
StoppingCondition={
'MaxRuntimeInSeconds': 300
}
)
print(job_name)
default_bucket
```
## NEO
```
!echo s3://$default_bucket/neo/VGG16/model-$target_device\.tar.gz
!aws s3 cp s3://$default_bucket/neo/VGG16/model-$target_device\.tar.gz .
!tar -tzvf model-$target_device\.tar.gz
!rm -rf neo_test && mkdir neo_test
!tar -xzvf model-$target_device\.tar.gz -C neo_test
!ldd neo_test/compiled.so
%%time
import os
import numpy as np
import cv2
import time
from dlr import DLRModel
model_path='neo_test'
imgMean = np.array([104, 117, 124], np.float)
img = cv2.imread("TestImage.jpg")
img = cv2.resize(img.astype(float), (224, 224)) # resize
img -= imgMean #subtract image mean
img = img.reshape((1, 224, 224, 3))
device = 'cpu' # Go, Raspberry Pi, go!
model = DLRModel(model_path, dev_type=device)
print(model.get_input_names())
def predict(img):
start = time.time()
input_data = {'Placeholder': img}
out = model.run(input_data)
return (out, time.time()-start)
startTime = time.time()
out = [predict(img)[1] for i in (range(1))]
print("Elapsed time: {}".format((time.time() - start_time)))
#top1 = np.argmax(out[0])
#prob = np.max(out)
#print("Class: %d, probability: %f" % (top1, prob))
%matplotlib inline
import matplotlib.pyplot as plt
fig = plt.figure(figsize=(20,10))
ax = plt.axes()
ax.plot(out)
```
| github_jupyter |
# Basic Distributions
### A. Taylan Cemgil
### Boğaziçi University, Dept. of Computer Engineering
### Notebook Summary
* We review the notation and parametrization of densities of some basic distributions that are often encountered
* We show how random numbers are generated using python libraries
* We show some basic visualization methods such as displaying histograms
# Sampling From Basic Distributions
Sampling from basic distribution is easy using the numpy library.
Formally we will write
$x \sim p(X|\theta)$
where $\theta$ is the _parameter vector_, $p(X| \theta)$ denotes the _density_ of the random variable $X$ and $x$ is a _realization_, a particular draw from the density $p$.
The following distributions are building blocks from which more complicated processes may be constructed. It is important to have a basic understanding of these distributions.
### Continuous Univariate
* Uniform $\mathcal{U}$
* Univariate Gaussian $\mathcal{N}$
* Gamma $\mathcal{G}$
* Inverse Gamma $\mathcal{IG}$
* Beta $\mathcal{B}$
### Discrete
* Poisson $\mathcal{P}$
* Bernoulli $\mathcal{BE}$
* Binomial $\mathcal{BI}$
* Categorical $\mathcal{M}$
* Multinomial $\mathcal{M}$
### Continuous Multivariate (todo)
* Multivariate Gaussian $\mathcal{N}$
* Dirichlet $\mathcal{D}$
### Continuous Matrix-variate (todo)
* Wishart $\mathcal{W}$
* Inverse Wishart $\mathcal{IW}$
* Matrix Gaussian $\mathcal{N}$
## Sampling from the standard uniform $\mathcal{U}(0,1)$
For generating a single random number in the interval $[0, 1)$ we use the notation
$$
x_1 \sim \mathcal{U}(x; 0,1)
$$
In python, this is implemented as
```
import numpy as np
x_1 = np.random.rand()
print(x_1)
```
We can also generate an array of realizations $x_i$ for $i=1 \dots N$,
$$
x_i \sim \mathcal{U}(x; 0,1)
$$
```
import numpy as np
N = 5
x = np.random.rand(N)
print(x)
```
For large $N$, it is more informative to display an histogram of generated data:
```
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
# Number of realizations
N = 50000
x = np.random.rand(N)
plt.hist(x, bins=20)
plt.xlabel('x')
plt.ylabel('Count')
plt.show()
```
$\newcommand{\indi}[1]{\left[{#1}\right]}$
$\newcommand{\E}[1]{\left\langle{#1}\right\rangle}$
We know that the density of the uniform distribution $\mathcal{U}(0,1)$ is
$$
\mathcal{U}(x; 0,1) = \left\{ \begin{array}{cc} 1 & 0 \leq x < 1 \\ 0 & \text{otherwise} \end{array} \right.
$$
or using the indicator notation
$$
\mathcal{U}(x; 0,1) = \left[ x \in [0,1) \right]
$$
#### Indicator function
To write and manipulate discrete probability distributions in algebraic expression, the *indicator* function is useful:
$$ \left[x\right] = \left\{ \begin{array}{cc}
1 & x\;\;\text{is true} \\
0 & x\;\;\text{is false}
\end{array}
\right.$$
This notation is also known as the Iverson's convention.
#### Aside: How to plot the density and the histogram onto the same plot?
In one dimension, the histogram is simply the count of the data points that fall to a given interval. Mathematically, we have
$j = 1\dots J$ intervals where $B_j = [b_{j-1}, b_j]$ and $b_j$ are bin boundries such that $b_0 < b_1 < \dots < b_J$.
$$
h(x) = \sum_{j=1}^J \sum_{i=1}^N \indi{x \in B_j} \indi{x_i \in B_j}
$$
This expression, at the first sight looks somewhat more complicated than it really is. The indicator product just encodes the logical condition $x \in B_j$ __and__ $x_i \in B_j$. The sum over $j$ is just a convenient way of writing the result instead of specifying the histogram as a case by case basis for each bin. It is important to get used to such nested sums.
When the density $p(x)$ is given, the probability that a single realization is in bin $B_j$ is given by
$$
\Pr\left\{x \in B_j\right\} = \int_{B_j} dx p(x) = \int_{-\infty}^{\infty} dx \indi{x\in B_j} p(x) = \E{\indi{x\in B_j}}
$$
In other words, the probability is just the expectation of the indicator.
The histogram can be written as follows
$$
h(x) = \sum_{j=1}^J \indi{x \in B_j} \sum_{i=1}^N \indi{x_i \in B_j}
$$
We define the counts at each bin as
$$
c_j \equiv \sum_{i=1}^N \indi{x_i \in B_j}
$$
If all bins have the same width, i.e., $b_j - b_{j-1} = \Delta$ for $\forall j$, and if $\Delta$ is sufficiently small we have
$$
\E{\indi{x\in B_j}} \approx p(b_{j-1}+\Delta/2) \Delta
$$
i.e., the probability is roughly the interval width times the density evaluated at the middle point of the bin. The expected value of the counts is
$$
\E{c_j} = \sum_{i=1}^N \E{\indi{x_i \in B_j}} \approx N \Delta p(b_{j-1}+\Delta/2)
$$
Hence, the density should be roughly
$$
p(b_{j-1}+\Delta/2) \approx \frac{\E{c_j} }{N \Delta}
$$
The $N$ term is intuitive but the $\Delta$ term is easily forgotten. When plotting the histograms on top of the corresponding densities, we should scale the normalized histogram ${ c_j }/{N}$ by dividing by $\Delta$.
```
N = 1000
# Bin width
Delta = 0.02
# Bin edges
b = np.arange(0 ,1+Delta, Delta)
# Evaluate the density
g = np.ones(b.size)
# Draw the samples
u = np.random.rand(N)
counts,edges = np.histogram(u, bins=b)
plt.bar(b[:-1], counts/N/Delta, width=Delta)
#plt.hold(True)
plt.plot(b, g, linewidth=3, color='y')
#plt.hold(False)
plt.show()
```
The __plt.hist__ function (calling __np.histogram__) can do this calculation automatically if the option normed=True. However, when the grid is not uniform, it is better to write your own code to be sure what is going on.
```
N = 1000
Delta = 0.05
b = np.arange(0 ,1+Delta, Delta)
g = np.ones(b.size)
u = np.random.rand(N)
#plt.hold(True)
plt.plot(b, g, linewidth=3, color='y')
plt.hist(u, bins=b, normed=True)
#plt.hold(False)
plt.show()
```
# Continuous Univariate Distributions
* Uniform $\mathcal{U}$
* Univariate Gaussian $\mathcal{N}$
$${\cal N}(x;\mu, v) = \frac{1}{\sqrt{2\pi v}} \exp\left(-\frac12 \frac{(x - \mu)^2}{v}\right) $$
* Gamma $\mathcal{G}$
$${\cal G}(\lambda; a, b) = \frac{b^a \lambda^{a-1}}{\Gamma(a)} \exp( - b \lambda)$$
* Inverse Gamma $\mathcal{IG}$
$${\cal IG}(v; \alpha, \beta) = \frac{\beta^\alpha}{\Gamma(\alpha) v^{\alpha+1}} \exp(- \frac{\beta}{v}) $$
* Beta $\mathcal{B}$
$${\cal B}(r; \alpha, \beta) = \frac{\Gamma(\alpha + \beta)}{\Gamma(\alpha) \Gamma(\beta) } r^{\alpha-1} (1-r)^{\beta-1}$$
In derivations, the distributions are often needed as building blocks. The following code segment prints the latex strings to be copied and pasted.
$\DeclareMathOperator{\trace}{Tr}$
```
from IPython.display import display, Math, Latex, HTML
import notes_utilities as nut
print('Gaussian')
L = nut.pdf2latex_gauss(x=r'Z_{i,j}', m=r'\mu_{i,j}',v=r'l_{i,j}')
display(HTML(nut.eqs2html_table(L)))
print('Gamma')
L = nut.pdf2latex_gamma(x=r'u', a=r'a',b=r'b')
display(HTML(nut.eqs2html_table(L)))
print('Inverse Gamma')
L = nut.pdf2latex_invgamma(x=r'z', a=r'a',b=r'b')
display(HTML(nut.eqs2html_table(L)))
print('Beta')
L = nut.pdf2latex_beta(x=r'\pi', a=r'\alpha',b=r'\beta')
display(HTML(nut.eqs2html_table(L)))
```
We will illustrate two alternative ways for sampling from continuous distributions.
- The first method has minimal dependence on the numpy and scipy libraries. This is initially the preferred method. Only random variable generators and the $\log \Gamma(x)$ (__gammaln__) function is used and nothing more.
- The second method uses scipy. This is a lot more practical but requires knowing more about the internals of the library.
### Aside: The Gamma function $\Gamma(x)$
The gamma function $\Gamma(x)$ is the (generalized) factorial.
- Defined by
$$\Gamma(x) = \int_0^{\infty} t^{x-1} e^{-t}\, dt$$
- For integer $x$, $\Gamma(x) = (x-1)!$. Remember that for positive integers $x$, the factorial function can be defined recursively $x! = (x-1)! x $ for $x\geq 1$.
- For real $x>1$, the gamma function satisfies
$$
\Gamma(x+1) = \Gamma(x) x
$$
- Interestingly, we have
$$\Gamma(1/2) = \sqrt{\pi}$$
- Hence
$$\Gamma(3/2) = \Gamma(1/2 + 1) = \Gamma(1/2) (1/2) = \sqrt{\pi}/2$$
- It is available in many numerical computation packages, in python it is available as __scipy.special.gamma__.
- To compute $\log \Gamma(x)$, you should always use the implementation as __scipy.special.gammaln__. The gamma function blows up super-exponentially so numerically you should never evaluate $\log \Gamma(x)$ as
```python
import numpy as np
import scipy.special as sps
np.log(sps.gamma(x)) # Don't
sps.gammaln(x) # Do
```
- A related function is the Beta function
$$B(x,y) = \int_0^{1} t^{x-1} (1-t)^{y-1}\, dt$$
- We have
$$B(x,y) = \frac{\Gamma(x)\Gamma(y)}{\Gamma(x+y)}$$
- Both $\Gamma(x)$ and $B(x)$ pop up as normalizing constant of the gamma and beta distributions.
#### Derivatives of $\Gamma(x)$
- <span style="color:red"> </span> The derivatives of $\log \Gamma(x)$ pop up quite often when fitting densities. The first derivative has a specific name, often called the digamma function or the psi function.
$$
\Psi(x) \equiv \frac{d}{d x} \log \Gamma(x)
$$
- It is available as __scipy.special.digamma__ or __scipy.special.psi__
- Higher order derivatives of the $\log \Gamma(x)$ function (including digamma itself) are available as __scipy.special.polygamma__
```
import numpy as np
import scipy.special as sps
import matplotlib.pyplot as plt
x = np.arange(0.1,5,0.01)
f = sps.gammaln(x)
df = sps.psi(x)
# First derivative of the digamma function
ddf = sps.polygamma(1,x)
# sps.psi(x) == sps.polygamma(0,x)
plt.figure(figsize=(8,10))
plt.subplot(3,1,1)
plt.plot(x, f, 'r')
plt.grid(True)
plt.xlabel('x')
plt.ylabel('log Gamma(x)')
plt.subplot(3,1,2)
plt.grid(True)
plt.plot(x, df, 'b')
plt.xlabel('x')
plt.ylabel('Psi(x)')
plt.subplot(3,1,3)
plt.plot(x, ddf, 'k')
plt.grid(True)
plt.xlabel('x')
plt.ylabel('Psi\'(x)')
plt.show()
```
#### Stirling's approximation
An important approximation to the factorial is the famous Stirling's approximation
\begin{align}
n! \sim \sqrt{2 \pi n}\left(\frac{n}{e}\right)^n
\end{align}
\begin{align}
\log \Gamma(x+1) \approx \frac{1}{2}\log(2 \pi) + x \log(x) - \frac{1}{2} \log(x)
\end{align}
```
import matplotlib.pylab as plt
import numpy as np
from scipy.special import polygamma
from scipy.special import gammaln as loggamma
from scipy.special import psi
x = np.arange(0.001,6,0.001)
ylim = [-1,8]
xlim = [-1,6]
plt.plot(x, loggamma(x), 'b')
stir = x*np.log(x)-x +0.5*np.log(2*np.pi*x)
plt.plot(x+1, stir,'r')
plt.hlines(0,0,8)
plt.vlines([0,1,2],ylim[0],ylim[1],linestyles=':')
plt.hlines(range(ylim[0],ylim[1]),xlim[0],xlim[1],linestyles=':',colors='g')
plt.ylim(ylim)
plt.xlim(xlim)
plt.legend([r'\log\Gamma(x)',r'\log(x-1)'],loc=1)
plt.xlabel('x')
plt.show()
```
# Sampling from Continuous Univariate Distributions
## Sampling using numpy.random
```
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
from scipy.special import gammaln
def plot_histogram_and_density(N, c, edges, dx, g, title='Put a title'):
'''
N : Number of Datapoints
c : Counts, as obtained from np.histogram function
edges : bin edges, as obtained from np.histogram
dx : The bin width
g : Density evaluated at the points given in edges
title : for the plot
'''
plt.bar(edges[:-1], c/N/dx, width=dx)
# plt.hold(True)
plt.plot(edges, g, linewidth=3, color='y')
# plt.hold(False)
plt.title(title)
def log_gaussian_pdf(x, mu, V):
return -0.5*np.log(2*np.pi*V) -0.5*(x-mu)**2/V
def log_gamma_pdf(x, a, b):
return (a-1)*np.log(x) - b*x - gammaln(a) + a*np.log(b)
def log_invgamma_pdf(x, a, b):
return -(a+1)*np.log(x) - b/x - gammaln(a) + a*np.log(b)
def log_beta_pdf(x, a, b):
return - gammaln(a) - gammaln(b) + gammaln(a+b) + np.log(x)*(a-1) + np.log(1-x)*(b-1)
N = 1000
# Univariate Gaussian
mu = 2 # mean
V = 1.2 # Variance
x_normal = np.random.normal(mu, np.sqrt(V), N)
dx = 10*np.sqrt(V)/50
x = np.arange(mu-5*np.sqrt(V) ,mu+5*np.sqrt(V),dx)
g = np.exp(log_gaussian_pdf(x, mu, V))
#g = scs.norm.pdf(x, loc=mu, scale=np.sqrt(V))
c,edges = np.histogram(x_normal, bins=x)
plt.figure(num=None, figsize=(16, 5), dpi=80, facecolor='w', edgecolor='k')
plt.subplot(2,2,1)
plot_histogram_and_density(N, c, x, dx, g, 'Gaussian')
## Gamma
# Shape
a = 1.2
# inverse scale
b = 30
# Generate unit scale first than scale with inverse scale parameter b
x_gamma = np.random.gamma(a, 1, N)/b
dx = np.max(x_gamma)/500
x = np.arange(dx, 250*dx, dx)
g = np.exp(log_gamma_pdf(x, a, b))
c,edges = np.histogram(x_gamma, bins=x)
plt.subplot(2,2,2)
plot_histogram_and_density(N, c, x, dx, g, 'Gamma')
## Inverse Gamma
a = 3.5
b = 0.2
x_invgamma = b/np.random.gamma(a, 1, N)
dx = np.max(x_invgamma)/500
x = np.arange(dx, 150*dx, dx)
g = np.exp(log_invgamma_pdf(x,a,b))
c,edges = np.histogram(x_invgamma, bins=x)
plt.subplot(2,2,3)
plot_histogram_and_density(N, c, x, dx, g, 'Inverse Gamma')
## Beta
a = 0.5
b = 1
x_beta = np.random.beta(a, b, N)
dx = 0.01
x = np.arange(dx, 1, dx)
g = np.exp(log_beta_pdf(x, a, b))
c,edges = np.histogram(x_beta, bins=x)
plt.subplot(2,2,4)
plot_histogram_and_density(N, c, x, dx, g, 'Beta')
plt.show()
```
## Sampling using scipy.stats
```
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import scipy.stats as scs
N = 2000
# Univariate Gaussian
mu = 2 # mean
V = 1.2 # Variance
rv_normal = scs.norm(loc=mu, scale=np.sqrt(V))
x_normal = rv_normal.rvs(size=N)
dx = 10*np.sqrt(V)/50
x = np.arange(mu-5*np.sqrt(V) ,mu+5*np.sqrt(V),dx)
g = rv_normal.pdf(x)
c,edges = np.histogram(x_normal, bins=x)
plt.figure(num=None, figsize=(16, 5), dpi=80, facecolor='w', edgecolor='k')
plt.subplot(2,2,1)
plot_histogram_and_density(N, c, x, dx, g, 'Gaussian')
## Gamma
a = 3.2
b = 30
# The following is equivalent to our parametrization of gamma, note the 1/b term
rv_gamma = scs.gamma(a, scale=1/b)
x_gamma = rv_gamma.rvs(N)
dx = np.max(x_gamma)/500
x = np.arange(0, 250*dx, dx)
g = rv_gamma.pdf(x)
c,edges = np.histogram(x_gamma, bins=x)
plt.subplot(2,2,2)
plot_histogram_and_density(N, c, x, dx, g, 'Gamma')
## Inverse Gamma
a = 3.5
b = 0.2
# Note the b term
rv_invgamma = scs.invgamma(a, scale=b)
x_invgamma = rv_invgamma.rvs(N)
dx = np.max(x_invgamma)/500
x = np.arange(dx, 150*dx, dx)
g = rv_invgamma.pdf(x)
c,edges = np.histogram(x_invgamma, bins=x)
plt.subplot(2,2,3)
plot_histogram_and_density(N, c, x, dx, g, 'Inverse Gamma')
## Beta
a = 0.7
b = 0.8
rv_beta = scs.beta(a, b)
x_beta = rv_beta.rvs(N)
dx = 0.02
x = np.arange(0, 1+dx, dx)
g = rv_beta.pdf(x)
c,edges = np.histogram(x_beta, bins=x)
plt.subplot(2,2,4)
plot_histogram_and_density(N, c, x, dx, g, 'Beta')
plt.show()
```
# Sampling from Discrete Densities
* Bernoulli $\mathcal{BE}$
$$
{\cal BE}(r; w) = w^r (1-w)^{1-r} \;\; \text{if} \; r \in \{0, 1\}
$$
* Binomial $\mathcal{BI}$
$${\cal BI}(r; L, w) = \binom{L}{r, (L-r)} w^r (1-w)^{L-r} \;\; \text{if} \; r \in \{0, 1, \dots, L\} $$
Here, the binomial coefficient is defined as
$$
\binom{L}{r, (L-r)} = \frac{N!}{r!(L-r)!}
$$
Note that
$$
{\cal BE}(r; w) = {\cal BI}(r; L=1, w)
$$
* Poisson $\mathcal{PO}$, with intensity $\lambda$
$${\cal PO}(x;\lambda) = \frac{e^{-\lambda} \lambda^x}{x!} = \exp(x \log \lambda - \lambda - \log\Gamma(x+1)) $$
Given samples on nonnegative integers, we can obtain histograms easily using __np.bincount__.
```python
c = np.bincount(samples)
```
The functionality is equivalent to the following sniplet, while implementation is possibly different and more efficient.
```python
upper_bound = np.max()
c = np.zeros(upper_bound+1)
for i in samples:
c[i] += 1
```
```
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
def plot_histogram_and_pmf(N, c, domain, dx, g, title='Put a title'):
'''
N : Number of Datapoints
c : Counts, as obtained from np.bincount function
domain : integers for each c, same size as c
dx : The bin width
g : Density evaluated at the points given in edges
title : for the plot
'''
plt.bar(domain-dx/2, c/N, width=dx)
# plt.hold(True)
plt.plot(domain, g, 'ro:', linewidth=3, color='y')
# plt.hold(False)
plt.title(title)
def log_poisson_pdf(x, lam):
return -lam + x*np.log(lam) - gammaln(x+1)
def log_bernoulli_pdf(r, pr):
return r*np.log(pr) + (1-r)*np.log(1 - pr)
def log_binomial_pdf(r, pr, L):
return gammaln(L+1) - gammaln(r+1) - gammaln(L-r+1) + r*np.log(pr) + (L-r)*np.log(1 - pr)
N = 100
pr = 0.8
# For plots
bin_width = 0.3
# Bernoulli
L = 1
x_bern = np.random.binomial(n=L, p=pr, size=N)
c = np.bincount(x_bern, minlength=L+1)
g = np.exp(log_bernoulli_pdf(np.arange(L+1), pr))
plt.figure(figsize=(20,4))
plt.subplot(1,3,1)
plot_histogram_and_pmf(N, c, np.arange(L+1), bin_width, g, 'Bernoulli')
plt.xticks([0,1])
# Binomial
L = 10
pr = 0.7
x_binom = np.random.binomial(n=L, p=pr, size=N)
c = np.bincount(x_binom, minlength=L+1)
g = np.exp(log_binomial_pdf(np.arange(L+1), pr, L))
plt.subplot(1,3,2)
plot_histogram_and_pmf(N, c, np.arange(L+1), bin_width, g, 'Binomial')
plt.xticks(np.arange(L+1))
# Poisson
intensity = 10.5
x_poiss = np.random.poisson(intensity, size =N )
c = np.bincount(x_poiss)
x = np.arange(len(c))
g = np.exp(log_poisson_pdf(x, intensity))
plt.subplot(1,3,3)
plot_histogram_and_pmf(N, c, x, bin_width, g, 'Poisson')
```
## Bernoulli, Binomial, Categorical and Multinomial Distributions
The Bernoulli and Binomial distributions are quite simple and well known distributions on small integers, so it may come as a surprise that they have another, less obvious but arguably more useful representation as discrete multivariate densities. This representation makes the link to categorical distributions where there are more than two possible outcomes. Finally, all Bernoulli, Binomial or Categorical distributions are special cases of Multinomial distribution.
### Bernoulli
Recall the Bernoulli distribution $r \in \{0, 1\}$
$$
{\cal BE}(r; w) = w^r (1-w)^{1-r}
$$
We will define $\pi_0 = 1-w$ and $\pi_1 = w$, such that $\pi_0 + \pi_1 = 1$. The parameter vector is $\pi = (\pi_0, \pi_1)$
We will also introduce a positional encoding such that
\begin{eqnarray}
r = 0 & \Rightarrow & s = (1, 0) \\
r = 1 & \Rightarrow & s = (0, 1)
\end{eqnarray}
In other words $s = (s_0, s_1)$ is a 2-dimensional vector where
$$s_0, s_1 \in \{0,1\}\;\text{and}\; s_0 + s_1 = 1$$
We can now write the Bernoulli density
$$
p(s | \pi) = \pi_0^{s_0} \pi_1^{s_1}
$$
### Binomial
Similarly, recall the Binomial density where $r \in \{0, 1, \dots, L\}$
$${\cal BI}(r; L, w) = \binom{L}{r, (L-r)} w^r (1-w)^{L-r} $$
We will again define $\pi_0 = 1-w$ and $\pi_1 = w$, such that $\pi_0 + \pi_1 = 1$. The parameter vector is $\pi = (\pi_0, \pi_1)$
\begin{eqnarray}
r = 0 & \Rightarrow & s = (L, 0) \\
r = 1 & \Rightarrow & s = (L-1, 1)\\
r = 2 & \Rightarrow & s = (L-2, 2)\\
\dots \\
r = L & \Rightarrow & s = (0, L)
\end{eqnarray}
where $s = (s_0, s_1)$ is a 2-dimensional vector where $$s_0, s_1 \in \{0,\dots,L\} \;\text{and}\; s_0 + s_1 = L$$
We can now write the Binomial density as
$$
p(s | \pi) = \binom{L}{s_0, s_1} \pi_0^{s_0} \pi_1^{s_1}
$$
### Categorical (Multinouilli)
One of the advantages of this new notation is that we can write the density even if the outcomes are not numerical. For example, the result of a single coin flip experiment when $r \in \{$ 'Tail', 'Head' $\}$ where the probability of 'Tail' is $w$ can be written as
$$
p(r | w) = w^{\indi{r=\text{'Tail'}}} (1-w)^{\indi{r=\text{'Head'}}}
$$
We define $s_0 = \indi{r=\text{'Head'}}$ and $s_1 = \indi{r=\text{'Tail'}}$, then the density can be written in the same form as
$$
p(s | \pi) = \pi_0^{s_0} \pi_1^{s_1}
$$
where $\pi_0 = 1-w$ and $\pi_1 = w$.
More generally, when $r$ is from a set with $K$ elements, i.e., $r \in R = \{ v_0, v_1, \dots, v_{K-1} \}$ with probability of the event $r = v_k$ given as $\pi_k$, we define $s = (s_0, s_1, \dots, s_{K-1})$ for $k=0,\dots, K-1$
$$
s_k = \indi{r=v_k}
$$
Note that by construction, we have $\sum_k s_k = 1$.
The resulting density, known as the Categorical density, can be writen as
$$
p(s|\pi) = \pi_0^{s_0} \pi_1^{s_1} \dots \pi_{K-1}^{s_{K-1}}
$$
### Multinomial
When drawing from a categorical distribution, one chooses a single category from $K$ options with given probabilities. A standard model for this is placing a single ball into $K$ different bins. The vector $s = (s_0, s_1, \dots,s_k, \dots, s_{K-1})$ represents how many balls eack bin $k$ contains.
Now, place $L$ balls instead of one into $K$ bins with placing each ball idependently into bin $k$ where $k \in\{0,\dots,K-1\}$ with the probability $\pi_k$. The multinomial is the joint distribution of $s$ where $s_k$ is the number of balls placed into bin $k$.
The density will be denoted as
$${\cal M}(s; L, \pi) = \binom{L}{s_0, s_1, \dots, s_{K-1}}\prod_{k=0}^{K-1} \pi_k^{s_k} $$
Here $\pi \equiv [\pi_0, \pi_2, \dots, \pi_{K-1} ]$ is the probability vector and $L$ is referred as the _index parameter_.
Clearly, we have the normalization constraint $ \sum_k \pi_k = 1$ and realization of the counts $s$ satisfy
$ \sum_k s_k = L $.
Here, the _multinomial_ coefficient is defined as
$$\binom{L}{s_0, s_1, \dots, s_{K-1}} = \frac{L!}{s_0! s_1! \dots s_{K-1}!}$$
Binomial, Bernoulli and Categorical distributions are all special cases of the Multinomial distribution, with a suitable representation.
The picture is as follows:
~~~
|Balls/Bins | $2$ Bins | $K$ Bins |
|-------- | -------- | ---------|
| $1$ Ball | Bernoulli ${\cal BE}$ | Categorical ${\cal C}$ |
|-------- | -------- | ---------|
| $L$ Balls | Binomial ${\cal BI}$ | Multinomial ${\cal M}$ |
~~~
Murphy calls the categorical distribution ($1$ Ball, $K$ Bins) as the Multinoulli. This is non-standard but logical (and somewhat cute).
It is common to consider Bernoulli and Binomial as scalar random variables. However, when we think of them as special case of a Multinomial it is better to think of them as bivariate, albeit degenerate, random variables, as
illustrated in the following cell along with an alternative visualization.
```
# The probability parameter
pr = 0.3
fig = plt.figure(figsize=(16,50), edgecolor=None)
maxL = 12
plt.subplot(maxL-1,2,1)
plt.grid(False)
# Set up the scalar binomial density as a bivariate density
for L in range(1,maxL):
r = np.arange(L+1)
p = np.exp(log_binomial_pdf(r, pr=pr, L=L))
A = np.zeros(shape=(13,13))
for s in range(L):
s0 = s
s1 = L-s
A[s0, s1] = p[s]
#plt.subplot(maxL-1,2,2*L-1)
# plt.bar(r-0.25, p, width=0.5)
# ax.set_xlim(-1,maxL)
# ax.set_xticks(range(0,maxL))
if True:
plt.subplot(maxL-1,2,2*L-1)
plt.barh(bottom=r-0.25, width=p, height=0.5)
ax2 = fig.gca()
pos = ax2.get_position()
pos2 = [pos.x0, pos.y0, 0.04, pos.height]
ax2.set_position(pos2)
ax2.set_ylim(-1,maxL)
ax2.set_yticks(range(0,maxL))
ax2.set_xlim([0,1])
ax2.set_xticks([0,1])
plt.ylabel('s1')
ax2.invert_xaxis()
plt.subplot(maxL-1,2,2*L)
plt.imshow(A, interpolation='nearest', origin='lower',cmap='gray_r',vmin=0,vmax=0.7)
plt.xlabel('s0')
ax1 = fig.gca()
pos = ax1.get_position()
pos2 = [pos.x0-0.45, pos.y0, pos.width, pos.height]
ax1.set_position(pos2)
ax1.set_ylim(-1,maxL)
ax1.set_yticks(range(0,maxL))
ax1.set_xlim(-1,maxL)
ax1.set_xticks(range(0,maxL))
plt.show()
```
The following cell illustrates sampling from the Multinomial density.
```
# Number of samples
sz = 3
# Multinomial
p = np.array([0.3, 0.1, 0.1, 0.5])
K = len(p) # number of Bins
L = 20 # number of Balls
print('Multinomial with number of bins K = {K} and Number of balls L = {L}'.format(K=K,L=L))
print(np.random.multinomial(L, p, size=sz))
# Categorical
L = 1 # number of Balls
print('Categorical with number of bins K = {K} and a single ball L=1'.format(K=K))
print(np.random.multinomial(L, p, size=sz))
# Binomial
p = np.array([0.3, 0.7])
K = len(p) # number of Bins = 2
L = 20 # number of Balls
print('Binomial with two bins K=2 and L={L} balls'.format(L=L))
print(np.random.multinomial(L, p, size=sz))
# Bernoulli
L = 1 # number of Balls
p = np.array([0.3, 0.7])
K = len(p) # number of Bins = 2
print('Bernoulli, two bins and a single ball')
print(np.random.multinomial(L, p, size=sz))
```
| github_jupyter |
# 4. Bayesian Networks
This notebook will show you how to query the main class of this package: `thomas.core.BayesianNetwork`.
```
%run '_preamble.ipynb'
from thomas.core.models.bn import BayesianNetwork
from thomas.core import examples
from thomas.core.reader import oobn
from IPython.display import display, HTML
```
## Student example
```
# Load the Student Network from the examples
Gs = examples.get_student_network()
Gs
# Show the network and its relations
Gs.draw()
# Nodes can be accessed by using the BN as a `dict`:
Gs['S']
# The Node-property 'states' will return a list of allowed states.
Gs['S'].states
# Accessing individual Nodes/CPTs is easy
Gs['S'].cpt
# Querying the network is also simple. To compute the marginals over I, S and D:
Gs.compute_marginals(['I', 'S', 'D'])
# To compute the marginals over I, S and D given evidence I=i0:
Gs.compute_marginals(['I', 'S', 'D'], {'I':'i0'})
# Complex queries can be run simply by providing the query as string:
Gs.P('I=i0|S')
# This should be equal to Gs['S']
Gs.P('S|I')
Gs.S.cpt
# Alternatively, parameters can be specified separately through
# `compute_posterior()`:
Gs.compute_posterior(qd=['I'], qv={}, ed=['S'], ev={})
# A CPT can be converted to a pandas.DataFrame by calling `unstack()` without
# parameters. Note that this only works if there are conditioning variables.
unstacked = Gs.P('I|S').as_series().unstack()
print(f'type(unstacked): {type(unstacked)}')
print()
display(unstacked)
# The junction tree (jointree) can be inspected by calling `jt.draw()`:
Gs.jt.draw()
```
## Importing Bayesian Networks
`thomas` supports loading Bayesian Networks from different file formats:
* `.net`: the Netica file format
* `.oobn`: the Hugin Object Oriented Bayesian Network file format
* `.json`: thomas' custom, JSON based format.
As an example, we'll open an `oobn` file.
```
from thomas.core.reader import oobn, net
# Read a file in OOBN format
filename = thomas.core.get_pkg_filename('lungcancer.oobn')
bn = oobn.read(filename)
bn.P('death').as_factor()
# Read a file in NET format
filename = thomas.core.get_pkg_filename('lungcancer.net')
bn = net.read(filename)
bn.P('death').as_factor()
# Download and read another file in NET format
import requests
import gzip
filename = 'asia.net'
if not os.path.exists(filename):
url = 'https://www.bnlearn.com/bnrepository/asia/asia.net.gz'
target_path = 'asia.net.gz'
print(f'Current directory: "{os.getcwd()}"')
print(f'Downloading "{target_path}" from "{url}"')
response = requests.get(url, stream=True)
if response.status_code == 200:
with open(target_path, 'wb') as f:
f.write(response.raw.read())
print(f'Unzipping "{target_path}"')
data = gzip.open(target_path)
print(f'Saving "{filename}"')
with open(filename, 'wb') as fp:
fp.write(data.read())
else:
print('ERROR: Could not download file!?')
bn = net.read(filename)
display(bn.P('asia'))
```
| github_jupyter |
<h2 style='color:blue' align="center" font='Verdana'>KNN (K Nearest Neighbors) machine learning algorithm by Maryam Nisha </h2>
**installing dependencies**
```
!pip install pandas
!pip install matplotlib
!pip install sklearn
!pip install seaborn
```
**loading iris builtin dataset of flowers in sklearn**
```
import pandas as pd
from sklearn.datasets import load_iris
iris = load_iris()
iris
```
<img height=300 width=300 src="iris_petal_sepal.png" />
**Features in Iris to implement KNN**
```
iris.feature_names
```
**Names of flowers in Iris**
```
iris.target_names
```
**Making data_frame by using Pandas**
```
df = pd.DataFrame(iris.data,columns=iris.feature_names)
df.head(10)
```
**Adding target index in dataframe**
```
df['target'] = iris.target
df.head()
df[df.target==1].head()
df[df.target==2].head()
```
**Adding flower name in dataframe with respect to index**
```
df['flower_name'] =df.target.apply(lambda x: iris.target_names[x])
df.head()
df[45:55]
```
**Slicing dataframe in three equal parts**
```
df0 = df[:50]
df1 = df[50:100]
df2 = df[100:]
```
**Using matplotlib to visualize dataset**
```
import matplotlib.pyplot as plt
%matplotlib inline
```
**Sepal length vs Sepal Width (Setosa vs Versicolor)**
```
plt.xlabel('Sepal Length')
plt.ylabel('Sepal Width')
plt.scatter(df0['sepal length (cm)'], df0['sepal width (cm)'],color="green",marker='+')
plt.scatter(df1['sepal length (cm)'], df1['sepal width (cm)'],color="blue",marker='.')
```
**Petal length vs Pepal Width (Setosa vs Versicolor)**
```
plt.xlabel('Petal Length')
plt.ylabel('Petal Width')
plt.scatter(df0['petal length (cm)'], df0['petal width (cm)'],color="green",marker='+')
plt.scatter(df1['petal length (cm)'], df1['petal width (cm)'],color="blue",marker='.')
```
**Train test split**
```
from sklearn.model_selection import train_test_split
X = df.drop(['target','flower_name'], axis='columns')
y = df.target
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=1)
len(X_train)
len(X_test)
```
**Training KNN (K Neighrest Neighbour Classifier) on Iris**
```
from sklearn.neighbors import KNeighborsClassifier
knn = KNeighborsClassifier(n_neighbors=10)
knn.fit(X_train, y_train)
```
**Accuracy of our model**
```
knn.score(X_test, y_test)
```
**prediction of new flower which family of flower it belongs in Iris by train KNN**
**[[sepal length (cm),sepal width (cm),petal length (cm),petal width (cm)]]**
```
knn.predict([[4.8,3.0,1.5,0.3]])
```
**Ploting Confusion Matrix by predicting all test data of flowers(split from iris) by KNN VS true_value of Iris**
```
from sklearn.metrics import confusion_matrix
y_pred = knn.predict(X_test)
cm = confusion_matrix(y_test, y_pred)
cm
%matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sn
plt.figure(figsize=(7,5))
sn.heatmap(cm, annot=True)
plt.xlabel('Predicted')
plt.ylabel('Truth')
```
**Print classification report for precesion, recall and f1-score for each classes**
```
from sklearn.metrics import classification_report
print(classification_report(y_test, y_pred))
```
**Thankyou Maryam Nisha**
| github_jupyter |
# Convolutional Neural Networks
## Project: Write an Algorithm for a Dog Identification App
---
In this notebook, some template code has already been provided for you, and you will need to implement additional functionality to successfully complete this project. You will not need to modify the included code beyond what is requested. Sections that begin with **'(IMPLEMENTATION)'** in the header indicate that the following block of code will require additional functionality which you must provide. Instructions will be provided for each section, and the specifics of the implementation are marked in the code block with a 'TODO' statement. Please be sure to read the instructions carefully!
> **Note**: Once you have completed all of the code implementations, you need to finalize your work by exporting the Jupyter Notebook as an HTML document. Before exporting the notebook to html, all of the code cells need to have been run so that reviewers can see the final implementation and output. You can then export the notebook by using the menu above and navigating to **File -> Download as -> HTML (.html)**. Include the finished document along with this notebook as your submission.
In addition to implementing code, there will be questions that you must answer which relate to the project and your implementation. Each section where you will answer a question is preceded by a **'Question X'** header. Carefully read each question and provide thorough answers in the following text boxes that begin with **'Answer:'**. Your project submission will be evaluated based on your answers to each of the questions and the implementation you provide.
>**Note:** Code and Markdown cells can be executed using the **Shift + Enter** keyboard shortcut. Markdown cells can be edited by double-clicking the cell to enter edit mode.
The rubric contains _optional_ "Stand Out Suggestions" for enhancing the project beyond the minimum requirements. If you decide to pursue the "Stand Out Suggestions", you should include the code in this Jupyter notebook.
---
### Why We're Here
In this notebook, you will make the first steps towards developing an algorithm that could be used as part of a mobile or web app. At the end of this project, your code will accept any user-supplied image as input. If a dog is detected in the image, it will provide an estimate of the dog's breed. If a human is detected, it will provide an estimate of the dog breed that is most resembling. The image below displays potential sample output of your finished project (... but we expect that each student's algorithm will behave differently!).

In this real-world setting, you will need to piece together a series of models to perform different tasks; for instance, the algorithm that detects humans in an image will be different from the CNN that infers dog breed. There are many points of possible failure, and no perfect algorithm exists. Your imperfect solution will nonetheless create a fun user experience!
### The Road Ahead
We break the notebook into separate steps. Feel free to use the links below to navigate the notebook.
* [Step 0](#step0): Import Datasets
* [Step 1](#step1): Detect Humans
* [Step 2](#step2): Detect Dogs
* [Step 3](#step3): Create a CNN to Classify Dog Breeds (from Scratch)
* [Step 4](#step4): Create a CNN to Classify Dog Breeds (using Transfer Learning)
* [Step 5](#step5): Write your Algorithm
* [Step 6](#step6): Test Your Algorithm
---
<a id='step0'></a>
## Step 0: Import Datasets
Make sure that you've downloaded the required human and dog datasets:
* Download the [dog dataset](https://s3-us-west-1.amazonaws.com/udacity-aind/dog-project/dogImages.zip). Unzip the folder and place it in this project's home directory, at the location `/dogImages`.
* Download the [human dataset](https://s3-us-west-1.amazonaws.com/udacity-aind/dog-project/lfw.zip). Unzip the folder and place it in the home directory, at location `/lfw`.
*Note: If you are using a Windows machine, you are encouraged to use [7zip](http://www.7-zip.org/) to extract the folder.*
In the code cell below, we save the file paths for both the human (LFW) dataset and dog dataset in the numpy arrays `human_files` and `dog_files`.
```
import numpy as np
from glob import glob
# load filenames for human and dog images
human_files = np.array(glob("lfw/*/*"))
dog_files = np.array(glob("dogImages/*/*/*"))
# print number of images in each dataset
print('There are %d total human images.' % len(human_files))
print('There are %d total dog images.' % len(dog_files))
```
<a id='step1'></a>
## Step 1: Detect Humans
In this section, we use OpenCV's implementation of [Haar feature-based cascade classifiers](http://docs.opencv.org/trunk/d7/d8b/tutorial_py_face_detection.html) to detect human faces in images.
OpenCV provides many pre-trained face detectors, stored as XML files on [github](https://github.com/opencv/opencv/tree/master/data/haarcascades). We have downloaded one of these detectors and stored it in the `haarcascades` directory. In the next code cell, we demonstrate how to use this detector to find human faces in a sample image.
```
import cv2
import matplotlib.pyplot as plt
%matplotlib inline
# extract pre-trained face detector
face_cascade = cv2.CascadeClassifier('haarcascades/haarcascade_frontalface_alt.xml')
# load color (BGR) image
img = cv2.imread(human_files[0])
# convert BGR image to grayscale
gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
# find faces in image
faces = face_cascade.detectMultiScale(gray)
# print number of faces detected in the image
print('Number of faces detected:', len(faces))
# get bounding box for each detected face
for (x,y,w,h) in faces:
# add bounding box to color image
cv2.rectangle(img,(x,y),(x+w,y+h),(255,0,0),2)
# convert BGR image to RGB for plotting
cv_rgb = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
# display the image, along with bounding box
plt.imshow(cv_rgb)
plt.show()
```
Before using any of the face detectors, it is standard procedure to convert the images to grayscale. The `detectMultiScale` function executes the classifier stored in `face_cascade` and takes the grayscale image as a parameter.
In the above code, `faces` is a numpy array of detected faces, where each row corresponds to a detected face. Each detected face is a 1D array with four entries that specifies the bounding box of the detected face. The first two entries in the array (extracted in the above code as `x` and `y`) specify the horizontal and vertical positions of the top left corner of the bounding box. The last two entries in the array (extracted here as `w` and `h`) specify the width and height of the box.
### Write a Human Face Detector
We can use this procedure to write a function that returns `True` if a human face is detected in an image and `False` otherwise. This function, aptly named `face_detector`, takes a string-valued file path to an image as input and appears in the code block below.
```
# returns "True" if face is detected in image stored at img_path
def face_detector(img_path):
img = cv2.imread(img_path)
gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
faces = face_cascade.detectMultiScale(gray)
return len(faces) > 0
```
### (IMPLEMENTATION) Assess the Human Face Detector
__Question 1:__ Use the code cell below to test the performance of the `face_detector` function.
- What percentage of the first 100 images in `human_files` have a detected human face?
- What percentage of the first 100 images in `dog_files` have a detected human face?
Ideally, we would like 100% of human images with a detected face and 0% of dog images with a detected face. You will see that our algorithm falls short of this goal, but still gives acceptable performance. We extract the file paths for the first 100 images from each of the datasets and store them in the numpy arrays `human_files_short` and `dog_files_short`.
__Answer:__
(You can print out your results and/or write your percentages in this cell)
```
from tqdm import tqdm
human_files_short = human_files[:100]
dog_files_short = dog_files[:100]
#-#-# Do NOT modify the code above this line. #-#-#
## TODO: Test the performance of the face_detector algorithm
## on the images in human_files_short and dog_files_short.
```
We suggest the face detector from OpenCV as a potential way to detect human images in your algorithm, but you are free to explore other approaches, especially approaches that make use of deep learning :). Please use the code cell below to design and test your own face detection algorithm. If you decide to pursue this _optional_ task, report performance on `human_files_short` and `dog_files_short`.
```
### (Optional)
### TODO: Test performance of another face detection algorithm.
### Feel free to use as many code cells as needed.
```
---
<a id='step2'></a>
## Step 2: Detect Dogs
In this section, we use a [pre-trained model](http://pytorch.org/docs/master/torchvision/models.html) to detect dogs in images.
### Obtain Pre-trained VGG-16 Model
The code cell below downloads the VGG-16 model, along with weights that have been trained on [ImageNet](http://www.image-net.org/), a very large, very popular dataset used for image classification and other vision tasks. ImageNet contains over 10 million URLs, each linking to an image containing an object from one of [1000 categories](https://gist.github.com/yrevar/942d3a0ac09ec9e5eb3a).
```
import torch
import torchvision.models as models
# define VGG16 model
VGG16 = models.vgg16(pretrained=True)
# check if CUDA is available
use_cuda = torch.cuda.is_available()
# move model to GPU if CUDA is available
if use_cuda:
VGG16 = VGG16.cuda()
```
Given an image, this pre-trained VGG-16 model returns a prediction (derived from the 1000 possible categories in ImageNet) for the object that is contained in the image.
### (IMPLEMENTATION) Making Predictions with a Pre-trained Model
In the next code cell, you will write a function that accepts a path to an image (such as `'dogImages/train/001.Affenpinscher/Affenpinscher_00001.jpg'`) as input and returns the index corresponding to the ImageNet class that is predicted by the pre-trained VGG-16 model. The output should always be an integer between 0 and 999, inclusive.
Before writing the function, make sure that you take the time to learn how to appropriately pre-process tensors for pre-trained models in the [PyTorch documentation](http://pytorch.org/docs/stable/torchvision/models.html).
```
from PIL import Image
import torchvision.transforms as transforms
# Set PIL to be tolerant of image files that are truncated.
from PIL import ImageFile
ImageFile.LOAD_TRUNCATED_IMAGES = True
def VGG16_predict(img_path):
'''
Use pre-trained VGG-16 model to obtain index corresponding to
predicted ImageNet class for image at specified path
Args:
img_path: path to an image
Returns:
Index corresponding to VGG-16 model's prediction
'''
## TODO: Complete the function.
## Load and pre-process an image from the given img_path
## Return the *index* of the predicted class for that image
return None # predicted class index
```
### (IMPLEMENTATION) Write a Dog Detector
While looking at the [dictionary](https://gist.github.com/yrevar/942d3a0ac09ec9e5eb3a), you will notice that the categories corresponding to dogs appear in an uninterrupted sequence and correspond to dictionary keys 151-268, inclusive, to include all categories from `'Chihuahua'` to `'Mexican hairless'`. Thus, in order to check to see if an image is predicted to contain a dog by the pre-trained VGG-16 model, we need only check if the pre-trained model predicts an index between 151 and 268 (inclusive).
Use these ideas to complete the `dog_detector` function below, which returns `True` if a dog is detected in an image (and `False` if not).
```
### returns "True" if a dog is detected in the image stored at img_path
def dog_detector(img_path):
## TODO: Complete the function.
return None # true/false
```
### (IMPLEMENTATION) Assess the Dog Detector
__Question 2:__ Use the code cell below to test the performance of your `dog_detector` function.
- What percentage of the images in `human_files_short` have a detected dog?
- What percentage of the images in `dog_files_short` have a detected dog?
__Answer:__
```
### TODO: Test the performance of the dog_detector function
### on the images in human_files_short and dog_files_short.
```
We suggest VGG-16 as a potential network to detect dog images in your algorithm, but you are free to explore other pre-trained networks (such as [Inception-v3](http://pytorch.org/docs/master/torchvision/models.html#inception-v3), [ResNet-50](http://pytorch.org/docs/master/torchvision/models.html#id3), etc). Please use the code cell below to test other pre-trained PyTorch models. If you decide to pursue this _optional_ task, report performance on `human_files_short` and `dog_files_short`.
```
### (Optional)
### TODO: Report the performance of another pre-trained network.
### Feel free to use as many code cells as needed.
```
---
<a id='step3'></a>
## Step 3: Create a CNN to Classify Dog Breeds (from Scratch)
Now that we have functions for detecting humans and dogs in images, we need a way to predict breed from images. In this step, you will create a CNN that classifies dog breeds. You must create your CNN _from scratch_ (so, you can't use transfer learning _yet_!), and you must attain a test accuracy of at least 10%. In Step 4 of this notebook, you will have the opportunity to use transfer learning to create a CNN that attains greatly improved accuracy.
We mention that the task of assigning breed to dogs from images is considered exceptionally challenging. To see why, consider that *even a human* would have trouble distinguishing between a Brittany and a Welsh Springer Spaniel.
Brittany | Welsh Springer Spaniel
- | -
<img src="images/Brittany_02625.jpg" width="100"> | <img src="images/Welsh_springer_spaniel_08203.jpg" width="200">
It is not difficult to find other dog breed pairs with minimal inter-class variation (for instance, Curly-Coated Retrievers and American Water Spaniels).
Curly-Coated Retriever | American Water Spaniel
- | -
<img src="images/Curly-coated_retriever_03896.jpg" width="200"> | <img src="images/American_water_spaniel_00648.jpg" width="200">
Likewise, recall that labradors come in yellow, chocolate, and black. Your vision-based algorithm will have to conquer this high intra-class variation to determine how to classify all of these different shades as the same breed.
Yellow Labrador | Chocolate Labrador | Black Labrador
- | -
<img src="images/Labrador_retriever_06457.jpg" width="150"> | <img src="images/Labrador_retriever_06455.jpg" width="240"> | <img src="images/Labrador_retriever_06449.jpg" width="220">
We also mention that random chance presents an exceptionally low bar: setting aside the fact that the classes are slightly imabalanced, a random guess will provide a correct answer roughly 1 in 133 times, which corresponds to an accuracy of less than 1%.
Remember that the practice is far ahead of the theory in deep learning. Experiment with many different architectures, and trust your intuition. And, of course, have fun!
### (IMPLEMENTATION) Specify Data Loaders for the Dog Dataset
Use the code cell below to write three separate [data loaders](http://pytorch.org/docs/stable/data.html#torch.utils.data.DataLoader) for the training, validation, and test datasets of dog images (located at `dogImages/train`, `dogImages/valid`, and `dogImages/test`, respectively). You may find [this documentation on custom datasets](http://pytorch.org/docs/stable/torchvision/datasets.html) to be a useful resource. If you are interested in augmenting your training and/or validation data, check out the wide variety of [transforms](http://pytorch.org/docs/stable/torchvision/transforms.html?highlight=transform)!
```
import os
from torchvision import datasets
### TODO: Write data loaders for training, validation, and test sets
## Specify appropriate transforms, and batch_sizes
```
**Question 3:** Describe your chosen procedure for preprocessing the data.
- How does your code resize the images (by cropping, stretching, etc)? What size did you pick for the input tensor, and why?
- Did you decide to augment the dataset? If so, how (through translations, flips, rotations, etc)? If not, why not?
**Answer**:
### (IMPLEMENTATION) Model Architecture
Create a CNN to classify dog breed. Use the template in the code cell below.
```
import torch.nn as nn
import torch.nn.functional as F
# define the CNN architecture
class Net(nn.Module):
### TODO: choose an architecture, and complete the class
def __init__(self):
super(Net, self).__init__()
## Define layers of a CNN
def forward(self, x):
## Define forward behavior
return x
#-#-# You do NOT have to modify the code below this line. #-#-#
# instantiate the CNN
model_scratch = Net()
# move tensors to GPU if CUDA is available
if use_cuda:
model_scratch.cuda()
```
__Question 4:__ Outline the steps you took to get to your final CNN architecture and your reasoning at each step.
__Answer:__
### (IMPLEMENTATION) Specify Loss Function and Optimizer
Use the next code cell to specify a [loss function](http://pytorch.org/docs/stable/nn.html#loss-functions) and [optimizer](http://pytorch.org/docs/stable/optim.html). Save the chosen loss function as `criterion_scratch`, and the optimizer as `optimizer_scratch` below.
```
import torch.optim as optim
### TODO: select loss function
criterion_scratch = None
### TODO: select optimizer
optimizer_scratch = None
```
### (IMPLEMENTATION) Train and Validate the Model
Train and validate your model in the code cell below. [Save the final model parameters](http://pytorch.org/docs/master/notes/serialization.html) at filepath `'model_scratch.pt'`.
```
# the following import is required for training to be robust to truncated images
from PIL import ImageFile
ImageFile.LOAD_TRUNCATED_IMAGES = True
def train(n_epochs, loaders, model, optimizer, criterion, use_cuda, save_path):
"""returns trained model"""
# initialize tracker for minimum validation loss
valid_loss_min = np.Inf
for epoch in range(1, n_epochs+1):
# initialize variables to monitor training and validation loss
train_loss = 0.0
valid_loss = 0.0
###################
# train the model #
###################
model.train()
for batch_idx, (data, target) in enumerate(loaders['train']):
# move to GPU
if use_cuda:
data, target = data.cuda(), target.cuda()
## find the loss and update the model parameters accordingly
## record the average training loss, using something like
## train_loss = train_loss + ((1 / (batch_idx + 1)) * (loss.data - train_loss))
######################
# validate the model #
######################
model.eval()
for batch_idx, (data, target) in enumerate(loaders['valid']):
# move to GPU
if use_cuda:
data, target = data.cuda(), target.cuda()
## update the average validation loss
# print training/validation statistics
print('Epoch: {} \tTraining Loss: {:.6f} \tValidation Loss: {:.6f}'.format(
epoch,
train_loss,
valid_loss
))
## TODO: save the model if validation loss has decreased
# return trained model
return model
# train the model
model_scratch = train(100, loaders_scratch, model_scratch, optimizer_scratch,
criterion_scratch, use_cuda, 'model_scratch.pt')
# load the model that got the best validation accuracy
model_scratch.load_state_dict(torch.load('model_scratch.pt'))
```
### (IMPLEMENTATION) Test the Model
Try out your model on the test dataset of dog images. Use the code cell below to calculate and print the test loss and accuracy. Ensure that your test accuracy is greater than 10%.
```
def test(loaders, model, criterion, use_cuda):
# monitor test loss and accuracy
test_loss = 0.
correct = 0.
total = 0.
model.eval()
for batch_idx, (data, target) in enumerate(loaders['test']):
# move to GPU
if use_cuda:
data, target = data.cuda(), target.cuda()
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the loss
loss = criterion(output, target)
# update average test loss
test_loss = test_loss + ((1 / (batch_idx + 1)) * (loss.data - test_loss))
# convert output probabilities to predicted class
pred = output.data.max(1, keepdim=True)[1]
# compare predictions to true label
correct += np.sum(np.squeeze(pred.eq(target.data.view_as(pred))).cpu().numpy())
total += data.size(0)
print('Test Loss: {:.6f}\n'.format(test_loss))
print('\nTest Accuracy: %2d%% (%2d/%2d)' % (
100. * correct / total, correct, total))
# call test function
test(loaders_scratch, model_scratch, criterion_scratch, use_cuda)
```
---
<a id='step4'></a>
## Step 4: Create a CNN to Classify Dog Breeds (using Transfer Learning)
You will now use transfer learning to create a CNN that can identify dog breed from images. Your CNN must attain at least 60% accuracy on the test set.
### (IMPLEMENTATION) Specify Data Loaders for the Dog Dataset
Use the code cell below to write three separate [data loaders](http://pytorch.org/docs/master/data.html#torch.utils.data.DataLoader) for the training, validation, and test datasets of dog images (located at `dogImages/train`, `dogImages/valid`, and `dogImages/test`, respectively).
If you like, **you are welcome to use the same data loaders from the previous step**, when you created a CNN from scratch.
```
## TODO: Specify data loaders
```
### (IMPLEMENTATION) Model Architecture
Use transfer learning to create a CNN to classify dog breed. Use the code cell below, and save your initialized model as the variable `model_transfer`.
```
import torchvision.models as models
import torch.nn as nn
## TODO: Specify model architecture
if use_cuda:
model_transfer = model_transfer.cuda()
```
__Question 5:__ Outline the steps you took to get to your final CNN architecture and your reasoning at each step. Describe why you think the architecture is suitable for the current problem.
__Answer:__
### (IMPLEMENTATION) Specify Loss Function and Optimizer
Use the next code cell to specify a [loss function](http://pytorch.org/docs/master/nn.html#loss-functions) and [optimizer](http://pytorch.org/docs/master/optim.html). Save the chosen loss function as `criterion_transfer`, and the optimizer as `optimizer_transfer` below.
```
criterion_transfer = None
optimizer_transfer = None
```
### (IMPLEMENTATION) Train and Validate the Model
Train and validate your model in the code cell below. [Save the final model parameters](http://pytorch.org/docs/master/notes/serialization.html) at filepath `'model_transfer.pt'`.
```
# train the model
model_transfer = # train(n_epochs, loaders_transfer, model_transfer, optimizer_transfer, criterion_transfer, use_cuda, 'model_transfer.pt')
# load the model that got the best validation accuracy (uncomment the line below)
#model_transfer.load_state_dict(torch.load('model_transfer.pt'))
```
### (IMPLEMENTATION) Test the Model
Try out your model on the test dataset of dog images. Use the code cell below to calculate and print the test loss and accuracy. Ensure that your test accuracy is greater than 60%.
```
test(loaders_transfer, model_transfer, criterion_transfer, use_cuda)
```
### (IMPLEMENTATION) Predict Dog Breed with the Model
Write a function that takes an image path as input and returns the dog breed (`Affenpinscher`, `Afghan hound`, etc) that is predicted by your model.
```
### TODO: Write a function that takes a path to an image as input
### and returns the dog breed that is predicted by the model.
# list of class names by index, i.e. a name can be accessed like class_names[0]
class_names = [item[4:].replace("_", " ") for item in data_transfer['train'].classes]
def predict_breed_transfer(img_path):
# load the image and return the predicted breed
return None
```
---
<a id='step5'></a>
## Step 5: Write your Algorithm
Write an algorithm that accepts a file path to an image and first determines whether the image contains a human, dog, or neither. Then,
- if a __dog__ is detected in the image, return the predicted breed.
- if a __human__ is detected in the image, return the resembling dog breed.
- if __neither__ is detected in the image, provide output that indicates an error.
You are welcome to write your own functions for detecting humans and dogs in images, but feel free to use the `face_detector` and `dog_detector` functions developed above. You are __required__ to use your CNN from Step 4 to predict dog breed.
Some sample output for our algorithm is provided below, but feel free to design your own user experience!

### (IMPLEMENTATION) Write your Algorithm
```
### TODO: Write your algorithm.
### Feel free to use as many code cells as needed.
def run_app(img_path):
## handle cases for a human face, dog, and neither
```
---
<a id='step6'></a>
## Step 6: Test Your Algorithm
In this section, you will take your new algorithm for a spin! What kind of dog does the algorithm think that _you_ look like? If you have a dog, does it predict your dog's breed accurately? If you have a cat, does it mistakenly think that your cat is a dog?
### (IMPLEMENTATION) Test Your Algorithm on Sample Images!
Test your algorithm at least six images on your computer. Feel free to use any images you like. Use at least two human and two dog images.
__Question 6:__ Is the output better than you expected :) ? Or worse :( ? Provide at least three possible points of improvement for your algorithm.
__Answer:__ (Three possible points for improvement)
```
## TODO: Execute your algorithm from Step 6 on
## at least 6 images on your computer.
## Feel free to use as many code cells as needed.
## suggested code, below
for file in np.hstack((human_files[:3], dog_files[:3])):
run_app(file)
```
| github_jupyter |
<h1>Analyzing US Economic Data and Building a Dashboard </h1>
<h2>Description</h2>
Extracting essential data from a dataset and displaying it is a necessary part of data science; therefore individuals can make correct decisions based on the data. In this assignment, you will extract some essential economic indicators from some data, you will then display these economic indicators in a Dashboard. You can then share the dashboard via an URL.
<p>
<a href="https://en.wikipedia.org/wiki/Gross_domestic_product"> Gross domestic product (GDP)</a> is a measure of the market value of all the final goods and services produced in a period. GDP is an indicator of how well the economy is doing. A drop in GDP indicates the economy is producing less; similarly an increase in GDP suggests the economy is performing better. In this lab, you will examine how changes in GDP impact the unemployment rate. You will take screen shots of every step, you will share the notebook and the URL pointing to the dashboard.</p>
<h2>Table of Contents</h2>
<div class="alert alert-block alert-info" style="margin-top: 20px">
<ul>
<li><a href="#Section_1"> Define a Function that Makes a Dashboard </a></li>
<li><a href="#Section_2">Question 1: Create a dataframe that contains the GDP data and display it</a> </li>
<li><a href="#Section_3">Question 2: Create a dataframe that contains the unemployment data and display it</a></li>
<li><a href="#Section_4">Question 3: Display a dataframe where unemployment was greater than 8.5%</a></li>
<li><a href="#Section_5">Question 4: Use the function make_dashboard to make a dashboard</a></li>
<li><a href="#Section_6"><b>(Optional not marked)</b> Save the dashboard on IBM cloud and display it</a></li>
</ul>
<p>
Estimated Time Needed: <strong>180 min</strong></p>
</div>
<hr>
<h2 id="Section_1"> Define Function that Makes a Dashboard </h2>
We will import the following libraries.
```
import pandas as pd
from bokeh.plotting import figure, output_file, show,output_notebook
output_notebook()
```
In this section, we define the function <code>make_dashboard</code>.
You don't have to know how the function works, you should only care about the inputs. The function will produce a dashboard as well as an html file. You can then use this html file to share your dashboard. If you do not know what an html file is don't worry everything you need to know will be provided in the lab.
```
def make_dashboard(x, gdp_change, unemployment, title, file_name):
output_file(file_name)
p = figure(title=title, x_axis_label='year', y_axis_label='%')
p.line(x.squeeze(), gdp_change.squeeze(), color="firebrick", line_width=4, legend="% GDP change")
p.line(x.squeeze(), unemployment.squeeze(), line_width=4, legend="% unemployed")
show(p)
```
The dictionary <code>links</code> contain the CSV files with all the data. The value for the key <code>GDP</code> is the file that contains the GDP data. The value for the key <code>unemployment</code> contains the unemployment data.
```
links={'GDP':'https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/projects/coursera_project/clean_gdp.csv',\
'unemployment':'https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/projects/coursera_project/clean_unemployment.csv'}
#links is dictionary - key value pair
```
<h3 id="Section_2"> Question 1: Create a dataframe that contains the GDP data and display the first five rows of the dataframe.</h3>
Use the dictionary <code>links</code> and the function <code>pd.read_csv</code> to create a Pandas dataframes that contains the GDP data.
<b>Hint: <code>links["GDP"]</code> contains the path or name of the file.</b>
```
# Type your code here
df=pd.read_csv(links["GDP"])
```
Use the method <code>head()</code> to display the first five rows of the GDP data, then take a screen-shot.
```
df.head()
# Type your code here
df.head(1000)
```
<h3 id="Section_2"> Question 2: Create a dataframe that contains the unemployment data. Display the first five rows of the dataframe. </h3>
Use the dictionary <code>links</code> and the function <code>pd.read_csv</code> to create a Pandas dataframes that contains the unemployment data.
```
# Type your code here
df2=pd.read_csv(links['unemployment'])
```
Use the method <code>head()</code> to display the first five rows of the GDP data, then take a screen-shot.
```
# Type your code here
df2.head()
df2.head(1000)
```
<h3 id="Section_3">Question 3: Display a dataframe where unemployment was greater than 8.5%. Take a screen-shot.</h3>
```
# Type your code here
df2.loc[df2['unemployment']>8.5]
```
<h3 id="Section_4">Question 4: Use the function make_dashboard to make a dashboard</h3>
In this section, you will call the function <code>make_dashboard</code> , to produce a dashboard. We will use the convention of giving each variable the same name as the function parameter.
Create a new dataframe with the column <code>'date'</code> called <code>x</code> from the dataframe that contains the GDP data.
```
x = df['date']
# Create your dataframe with column date
x
```
Create a new dataframe with the column <code>'change-current' </code> called <code>gdp_change</code> from the dataframe that contains the GDP data.
```
gdp_change = df['change-current']
# Create your dataframe with column change-current
gdp_change
```
Create a new dataframe with the column <code>'unemployment' </code> called <code>unemployment</code> from the dataframe that contains the unemployment data.
```
unemployment = df2['unemployment']
# Create your dataframe with column unemployment
unemployment
```
Give your dashboard a string title, and assign it to the variable <code>title</code>
```
title = 'Analysing US economic data'
# Give your dashboard a string title
```
Finally, the function <code>make_dashboard</code> will output an <code>.html</code> in your direictory, just like a <code>csv</code> file. The name of the file is <code>"index.html"</code> and it will be stored in the varable <code>file_name</code>.
```
file_name = "index.html"
```
Call the function <code>make_dashboard</code> , to produce a dashboard. Assign the parameter values accordingly take a the <b>, take a screen shot of the dashboard and submit it</b>.
```
# Fill up the parameters in the following function:
make_dashboard(x, gdp_change, unemployment, title, file_name)
```
<h3 id="Section_5"> <b>(Optional not marked)</b>Save the dashboard on IBM cloud and display it </h3>
From the tutorial <i>PROVISIONING AN OBJECT STORAGE INSTANCE ON IBM CLOUD</i> copy the JSON object containing the credentials you created. You’ll want to store everything you see in a credentials variable like the one below (obviously, replace the placeholder values with your own). Take special note of your <code>access_key_id</code> and <code>secret_access_key</code>. <b>Do not delete <code># @hidden_cell </code> as this will not allow people to see your credentials when you share your notebook. </b>
<code>
credentials = {<br>
"apikey": "your-api-key",<br>
"cos_hmac_keys": {<br>
"access_key_id": "your-access-key-here", <br>
"secret_access_key": "your-secret-access-key-here"<br>
},<br>
</code>
<code>
"endpoints": "your-endpoints",<br>
"iam_apikey_description": "your-iam_apikey_description",<br>
"iam_apikey_name": "your-iam_apikey_name",<br>
"iam_role_crn": "your-iam_apikey_name",<br>
"iam_serviceid_crn": "your-iam_serviceid_crn",<br>
"resource_instance_id": "your-resource_instance_id"<br>
}
</code>
```
# @hidden_cell
#
```
You will need the endpoint make sure the setting are the same as <i> PROVISIONING AN OBJECT STORAGE INSTANCE ON IBM CLOUD </i> assign the name of your bucket to the variable <code>bucket_name </code>
```
endpoint = 'https://s3-api.us-geo.objectstorage.softlayer.net'
```
From the tutorial <i> PROVISIONING AN OBJECT STORAGE INSTANCE ON IBM CLOUD </i> assign the name of your bucket to the variable <code>bucket_name </code>
```
bucket_name = # Type your bucket name on IBM Cloud
```
We can access IBM Cloud Object Storage with Python useing the <code>boto3</code> library, which we’ll import below:
```
import boto3
```
We can interact with IBM Cloud Object Storage through a <code>boto3</code> resource object.
```
resource = boto3.resource(
's3',
aws_access_key_id = credentials["cos_hmac_keys"]['access_key_id'],
aws_secret_access_key = credentials["cos_hmac_keys"]["secret_access_key"],
endpoint_url = endpoint,
)
```
We are going to use <code>open</code> to create a file object. To get the path of the file, you are going to concatenate the name of the file stored in the variable <code>file_name</code>. The directory stored in the variable directory using the <code>+</code> operator and assign it to the variable
<code>html_path</code>. We will use the function <code>getcwd()</code> to find current the working directory.
```
import os
directory = os.getcwd()
html_path = directory + "/" + file_name
```
Now you must read the html file, use the function <code>f = open(html_path, mode)</code> to create a file object and assign it to the variable <code>f</code>. The parameter <code>file</code> should be the variable <code>html_path</code>, the mode should be <code>"r"</code> for read.
```
# Type your code here
```
To load your dataset into the bucket we will use the method <code>put_object</code>, you must set the parameter name to the name of the bucket, the parameter <code>Key</code> should be the name of the HTML file and the value for the parameter Body should be set to <code>f.read()</code>.
```
# Fill up the parameters in the following function:
# resource.Bucket(name=).put_object(Key=, Body=)
```
In the dictionary <code>Params</code> provide the bucket name as the value for the key <i>'Bucket'</i>. Also for the value of the key <i>'Key'</i> add the name of the <code>html</code> file, both values should be strings.
```
# Fill in the value for each key
# Params = {'Bucket': ,'Key': }
```
The following lines of code will generate a URL to share your dashboard. The URL only last seven days, but don't worry you will get full marks if the URL is visible in your notebook.
```
import sys
time = 7*24*60**2
client = boto3.client(
's3',
aws_access_key_id = credentials["cos_hmac_keys"]['access_key_id'],
aws_secret_access_key = credentials["cos_hmac_keys"]["secret_access_key"],
endpoint_url=endpoint,
)
url = client.generate_presigned_url('get_object',Params=Params,ExpiresIn=time)
print(url)
```
<h2 id="Section_5"> How to submit </h2>
<p>Once you complete your notebook you will have to share it to be marked. Select the icon on the top right a marked in red in the image below, a dialogue box should open, select the option all content excluding sensitive code cells.</p>
<p><img height="440" width="700" src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/projects/EdX/ReadMe%20files/share_noteook1.png" alt="share notebook" /></p>
<p></p>
<p>You can then share the notebook via a URL by scrolling down as shown in the following image:</p>
<p style="text-align: center;"> <img height="308" width="350" src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/projects/EdX/ReadMe%20files/link2.png" alt="share notebook" /> </p>
<hr>
<p>Copyright © 2019 IBM Developer Skills Network. This notebook and its source code are released under the terms of the <a href="https://cognitiveclass.ai/mit-license/">MIT License</a>.</p>
<h2>About the Authors:</h2>
<a href="https://www.linkedin.com/in/joseph-s-50398b136/">Joseph Santarcangelo</a> has a PhD in Electrical Engineering, his research focused on using machine learning, signal processing, and computer vision to determine how videos impact human cognition. Joseph has been working for IBM since he completed his PhD.
<p>
Other contributors: <a href="https://www.linkedin.com/in/yi-leng-yao-84451275/">Yi leng Yao</a>, <a href="www.linkedin.com/in/jiahui-mavis-zhou-a4537814a">Mavis Zhou</a>
</p>
<h2>References :</h2>
<ul>
<il>
1) <a href="https://research.stlouisfed.org/">Economic Research at the St. Louis Fed </a>:<a href="https://fred.stlouisfed.org/series/UNRATE/"> Civilian Unemployment Rate</a>
</il>
<p>
<il>
2) <a href="https://github.com/datasets">Data Packaged Core Datasets
</a>
</il>
</p>
</ul>
</div>
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
import sys
sys.path.append('../')
import scripts.visualization as viz
from scripts.data_utils import load_results
viz.load_matplotlib()
plt = viz.plt
```
### Parameters:
- $\beta$ = 0.7
- $\gamma$ = 0.5
- $\mu$ = 0.85
- $\kappa$ = 0.1
- max_infected_time = 10
- NSTEPS = 50 000
- NAGENTS = 1 000
- NFRACLINKS = 0.1
- n_runs = 10
And we consider $q = \{3, 4, 5\}$
```
dr_q3_params, dr_q3_df = load_results(
"../data/new_experiments/p_q_random_opinion/p_3_q/dead_ratio_p_3_q_L1-beta=0.7_gamma=0.5_mu=0.85_kappa=0.1_max_infected_time=10_L2-q=4_p=0.5_xi=0.1_n=100_NRUNS=10_NSTEPS=50000_NAGENTS=1000_NFRACLINKS=0.1.csv")
ir_q3_params, ir_q3_df = load_results(
"../data/new_experiments/p_q_random_opinion/p_3_q/infected_ratio_p_3_q_L1-beta=0.7_gamma=0.5_mu=0.85_kappa=0.1_max_infected_time=10_L2-q=4_p=0.5_xi=0.1_n=100_NRUNS=10_NSTEPS=50000_NAGENTS=1000_NFRACLINKS=0.1.csv")
opinion_q3_params, opinion_q3_df = load_results(
"../data/new_experiments/p_q_random_opinion/p_3_q/mean_opinion_p_3_q_L1-beta=0.7_gamma=0.5_mu=0.85_kappa=0.1_max_infected_time=10_L2-q=4_p=0.5_xi=0.1_n=100_NRUNS=10_NSTEPS=50000_NAGENTS=1000_NFRACLINKS=0.1.csv")
dr_q4_params, dr_q4_df = load_results(
"../data/new_experiments/p_q_random_opinion/p_4_q/dead_ratio_p_4_q_L1-beta=0.7_gamma=0.5_mu=0.85_kappa=0.1_max_infected_time=10_L2-q=4_p=0.5_xi=0.1_n=100_NRUNS=10_NSTEPS=50000_NAGENTS=1000_NFRACLINKS=0.1.csv")
ir_q4_params, ir_q4_df = load_results(
"../data/new_experiments/p_q_random_opinion/p_4_q/infected_ratio_p_4_q_L1-beta=0.7_gamma=0.5_mu=0.85_kappa=0.1_max_infected_time=10_L2-q=4_p=0.5_xi=0.1_n=100_NRUNS=10_NSTEPS=50000_NAGENTS=1000_NFRACLINKS=0.1.csv")
opinion_q4_params, opinion_q4_df = load_results(
"../data/new_experiments/p_q_random_opinion/p_4_q/mean_opinion_p_4_q_L1-beta=0.7_gamma=0.5_mu=0.85_kappa=0.1_max_infected_time=10_L2-q=4_p=0.5_xi=0.1_n=100_NRUNS=10_NSTEPS=50000_NAGENTS=1000_NFRACLINKS=0.1.csv")
dr_q5_params, dr_q5_df = load_results(
"../data/new_experiments/p_q_random_opinion/p_5_q/dead_ratio_p_5_q_L1-beta=0.7_gamma=0.5_mu=0.85_kappa=0.1_max_infected_time=10_L2-q=4_p=0.5_xi=0.1_n=100_NRUNS=10_NSTEPS=50000_NAGENTS=1000_NFRACLINKS=0.1.csv")
ir_q5_params, ir_q5_df = load_results(
"../data/new_experiments/p_q_random_opinion/p_5_q/infected_ratio_p_5_q_L1-beta=0.7_gamma=0.5_mu=0.85_kappa=0.1_max_infected_time=10_L2-q=4_p=0.5_xi=0.1_n=100_NRUNS=10_NSTEPS=50000_NAGENTS=1000_NFRACLINKS=0.1.csv")
opinion_q5_params, opinion_q5_df = load_results(
"../data/new_experiments/p_q_random_opinion/p_5_q/mean_opinion_p_5_q_L1-beta=0.7_gamma=0.5_mu=0.85_kappa=0.1_max_infected_time=10_L2-q=4_p=0.5_xi=0.1_n=100_NRUNS=10_NSTEPS=50000_NAGENTS=1000_NFRACLINKS=0.1.csv")
colors = ['red', 'green', 'blue']
colors = ['xkcd:' + c for c in colors]
markers = ['o', 's', 'v']
markersize = 6
labels = [f'q={x}' for x in [3, 4, 5]]
ps = [float(x) for x in opinion_q3_df.columns]
ys = [opinion_q3_df, opinion_q4_df, opinion_q5_df]
for i, y in enumerate(ys):
plt.plot(ps, y.values[0], color=colors[i], label=labels[i], marker=markers[i],
markersize=markersize)
plt.ylabel('opinion', fontsize=20)
plt.xlabel('p', fontsize=20)
plt.legend()
ys = [dr_q3_df, dr_q4_df, dr_q5_df]
for i, y in enumerate(ys):
plt.plot(ps, y.values[0], color=colors[i], label=labels[i], marker=markers[i],
markersize=markersize)
plt.ylabel('dead rate', fontsize=20)
plt.xlabel('p', fontsize=20)
plt.legend()
ys = [ir_q3_df, ir_q4_df, ir_q5_df]
for i, y in enumerate(ys):
plt.plot(ps, y.values[0], color=colors[i], label=labels[i], marker=markers[i],
markersize=markersize)
plt.ylabel('max infected rate', fontsize=20)
plt.xlabel('p', fontsize=20)
plt.legend()
```
# Large network
## Parameters
- $\beta$ = 0.7
- $\gamma$ = 0.5
- $\mu$ = 0.85
- $\kappa$ = 0.1
- max_infected_time = 10
- NSTEPS = 200 000
- NAGENTS = 10 000
- NFRACLINKS = 0.1
- n_runs = 5
And we consider $q = \{3, 4, 5\}$
```
dr_q3_params, dr_q3_df_large = load_results(
"../data/new_experiments/p_q_random_opinion/p_3_q/dead_ratio_p_3_q_L1-beta=0.7_gamma=0.5_mu=0.85_kappa=0.1_max_infected_time=10_L2-q=4_p=0.5_xi=0.1_n=100_NRUNS=5_NSTEPS=200000_NAGENTS=10000_NFRACLINKS=0.1.csv")
ir_q3_params, ir_q3_df_large = load_results(
"../data/new_experiments/p_q_random_opinion/p_3_q/infected_ratio_p_3_q_L1-beta=0.7_gamma=0.5_mu=0.85_kappa=0.1_max_infected_time=10_L2-q=4_p=0.5_xi=0.1_n=100_NRUNS=5_NSTEPS=200000_NAGENTS=10000_NFRACLINKS=0.1.csv")
opinion_q3_params, opinion_q3_df_large = load_results(
"../data/new_experiments/p_q_random_opinion/p_3_q/mean_opinion_p_3_q_L1-beta=0.7_gamma=0.5_mu=0.85_kappa=0.1_max_infected_time=10_L2-q=4_p=0.5_xi=0.1_n=100_NRUNS=5_NSTEPS=200000_NAGENTS=10000_NFRACLINKS=0.1.csv")
dr_q4_params, dr_q4_df_large = load_results(
"../data/new_experiments/p_q_random_opinion/p_4_q/dead_ratio_p_4_q_L1-beta=0.7_gamma=0.5_mu=0.85_kappa=0.1_max_infected_time=10_L2-q=4_p=0.5_xi=0.1_n=100_NRUNS=5_NSTEPS=200000_NAGENTS=10000_NFRACLINKS=0.1.csv")
ir_q4_params, ir_q4_df_large = load_results(
"../data/new_experiments/p_q_random_opinion/p_4_q/infected_ratio_p_4_q_L1-beta=0.7_gamma=0.5_mu=0.85_kappa=0.1_max_infected_time=10_L2-q=4_p=0.5_xi=0.1_n=100_NRUNS=5_NSTEPS=200000_NAGENTS=10000_NFRACLINKS=0.1.csv")
opinion_q4_params, opinion_q4_df_large = load_results(
"../data/new_experiments/p_q_random_opinion/p_4_q/mean_opinion_p_4_q_L1-beta=0.7_gamma=0.5_mu=0.85_kappa=0.1_max_infected_time=10_L2-q=4_p=0.5_xi=0.1_n=100_NRUNS=5_NSTEPS=200000_NAGENTS=10000_NFRACLINKS=0.1.csv")
dr_q5_params, dr_q5_df_large = load_results(
"../data/new_experiments/p_q_random_opinion/p_5_q/dead_ratio_p_5_q_L1-beta=0.7_gamma=0.5_mu=0.85_kappa=0.1_max_infected_time=10_L2-q=4_p=0.5_xi=0.1_n=100_NRUNS=5_NSTEPS=200000_NAGENTS=10000_NFRACLINKS=0.1.csv")
ir_q5_params, ir_q5_df_large = load_results(
"../data/new_experiments/p_q_random_opinion/p_5_q/infected_ratio_p_5_q_L1-beta=0.7_gamma=0.5_mu=0.85_kappa=0.1_max_infected_time=10_L2-q=4_p=0.5_xi=0.1_n=100_NRUNS=5_NSTEPS=200000_NAGENTS=10000_NFRACLINKS=0.1.csv")
opinion_q5_params, opinion_q5_df_large = load_results(
"../data/new_experiments/p_q_random_opinion/p_5_q/mean_opinion_p_5_q_L1-beta=0.7_gamma=0.5_mu=0.85_kappa=0.1_max_infected_time=10_L2-q=4_p=0.5_xi=0.1_n=100_NRUNS=5_NSTEPS=200000_NAGENTS=10000_NFRACLINKS=0.1.csv")
labels = [f'q={x}' for x in [3, 4, 5]]
ps = [float(x) for x in opinion_q3_df_large.columns]
ys = [opinion_q3_df_large, opinion_q4_df_large, opinion_q5_df_large]
for i, y in enumerate(ys):
plt.plot(ps, y.values[0], color=colors[i], label=labels[i], marker=markers[i],
markersize=markersize)
plt.ylabel('opinion', fontsize=20)
plt.xlabel('p', fontsize=20)
plt.legend()
ys = [dr_q3_df_large, dr_q4_df_large, dr_q5_df_large]
for i, y in enumerate(ys):
plt.plot(ps, y.values[0], color=colors[i], label=labels[i], marker=markers[i],
markersize=markersize)
plt.ylabel('dead rate', fontsize=20)
plt.xlabel('p', fontsize=20)
plt.legend()
ys = [ir_q3_df_large, ir_q4_df_large, ir_q5_df_large]
for i, y in enumerate(ys):
plt.plot(ps, y.values[0], color=colors[i], label=labels[i], marker=markers[i],
markersize=markersize)
plt.ylabel('max infected rate', fontsize=20)
plt.xlabel('p', fontsize=20)
plt.legend()
```
| github_jupyter |
```
import sys
if not './' in sys.path:
sys.path.append('./')
import pandas as pd
import numpy as np
import io
import os
from datetime import datetime
from sklearn import preprocessing
from sklearn.model_selection import train_test_split
from envs.stocks_env_multiaction import Stocks_env
from datasets import nyse
import tensorflow as tf
from IPython.display import clear_output
import matplotlib.pyplot as plt
%matplotlib inline
#data = nyse.load_data('../data/')
data, _, _ = nyse.load_data_with_industry('../data/')
fit_data = data.drop(["symbol"], axis=1)
scaler = preprocessing.StandardScaler().fit(fit_data)
# Hyper params:
input_shape = np.shape(data)[1]-1
lr = 1e-3
seed = 42
epochs = 8
batch_size = 256
# log
save_directory = 'results/logreg/'
date = datetime.now().strftime("%Y_%m_%d-%H:%M:%S")
identifier = "logreg-" + date
train_summary_writer = tf.summary.create_file_writer('results/summaries/logreg/train/' + identifier)
test_summary_writer = tf.summary.create_file_writer('results/summaries/logreg/test/' + identifier)
train_loss = tf.keras.metrics.Mean('train_loss', dtype=tf.float32)
train_accuracy = tf.keras.metrics.BinaryAccuracy('train_accuracy')
test_loss = tf.keras.metrics.Mean('test_loss', dtype=tf.float32)
test_accuracy = tf.keras.metrics.BinaryAccuracy('test_accuracy')
# format data
symbols = data['symbol'].unique().tolist()
X = []
Y = []
for sym in symbols:
sym_data = scaler.transform(data.loc[data.symbol==sym].drop(["symbol"], axis=1))
for i in range(len(sym_data)-1):
X.append(sym_data[i])
Y.append(1 if (sym_data[i][1]<sym_data[i+1][1]) else 0)
X_train, X_test, y_train, y_test = train_test_split(X, Y, test_size=0.33, random_state=seed)
train_dataset = tf.data.Dataset.from_tensor_slices((X_train, y_train))
test_dataset = tf.data.Dataset.from_tensor_slices((X_test, y_test))
train_dataset = train_dataset.shuffle(60000).batch(batch_size)
test_dataset = test_dataset.batch(batch_size)
# initialize the model
model = tf.keras.Sequential(tf.keras.layers.Dense(1, input_shape=(input_shape,), activation='sigmoid'))
optimizer = tf.keras.optimizers.Adam(lr)
loss_object = tf.keras.losses.BinaryCrossentropy(from_logits=True)
def train_step(model, optimizer, x_train, y_train):
with tf.GradientTape() as tape:
predictions = model(x_train, training=True)
loss = loss_object(y_train, predictions)
grads = tape.gradient(loss, model.trainable_variables)
optimizer.apply_gradients(zip(grads, model.trainable_variables))
train_loss(loss)
train_accuracy(y_train, predictions)
def test_step(model, x_test, y_test):
predictions = model(x_test)
loss = loss_object(y_test, predictions)
test_loss(loss)
test_accuracy(y_test, predictions)
test_total_profits = []
for epoch in range(epochs):
for (x_train, y_train) in train_dataset:
train_step(model, optimizer, x_train, y_train)
with train_summary_writer.as_default():
tf.summary.scalar('loss', train_loss.result(), step=epoch)
tf.summary.scalar('accuracy', train_accuracy.result(), step=epoch)
for (x_test, y_test) in test_dataset:
test_step(model, x_test, y_test)
with test_summary_writer.as_default():
tf.summary.scalar('loss', test_loss.result(), step=epoch)
tf.summary.scalar('accuracy', test_accuracy.result(), step=epoch)
template = 'Epoch {}, Loss: {}, Accuracy: {}, Test Loss: {}, Test Accuracy: {}'
print (template.format(epoch+1,
train_loss.result(),
train_accuracy.result()*100,
test_loss.result(),
test_accuracy.result()*100))
# Reset metrics every epoch
train_loss.reset_states()
test_loss.reset_states()
train_accuracy.reset_states()
test_accuracy.reset_states()
columns = data.columns[1:].tolist()
weights = model.layers[0].get_weights()
a = [(weights[0][i][0], columns[i]) for i in range(len(columns))]
a
window_size = 1
run_lenght = 10
initial_money = 100
train_test_ratio = 0.2
env = Stocks_env(data, window_size, run_lenght, batch_size=batch_size, train_test_ratio = train_test_ratio,
test_seed=seed, initial_money=initial_money)
batch_size = len(env.get_test_symbols())
def test_env(record_days=False):
state = env.reset(training=False, batch_size=batch_size, run_lenght=run_lenght, initial_money=initial_money)
state = np.reshape(state, (batch_size, len(columns)))
done = False
operation_array = []
days_array = []
rewards_array = []
total_profit = np.zeros(batch_size)
while not done:
actions = []
for result in model(state):
buy = 1 if result>0 else 0
sell = 1 if result<0 else 0
actions += [[buy, sell]]
next_state, reward, done, operations, day, profit = env.step(actions)
state = next_state
if record_days:
operation_array.append(np.array(operations))
days_array.append(np.array(day))
rewards_array.append(np.array(reward))
mean_test_reward(np.array(reward))
total_profit += profit
total_profit = total_profit/initial_money
return operation_array, days_array, rewards_array, total_profit
save_directory = 'results/test-all/'
date = datetime.now().strftime("%Y_%m_%d-%H:%M:%S")
identifier = "logreg-" + date
test_summary_writer = tf.summary.create_file_writer('results/summaries/test/' + identifier)
mean_test_reward = tf.keras.metrics.Mean(name='mean_test_reward')
repeat = 100
test_total_profits = []
for i in range(repeat):
print(i)
operation_array, days_array, rewards_array, test_total_profit = test_env(record_days=True)
test_total_profits.append(test_total_profit)
with test_summary_writer.as_default():
tf.summary.scalar('mean_test_reward', mean_test_reward.result(), step=i)
# serialize weights to HDF5
if not os.path.exists(save_directory):
os.makedirs(save_directory)
if not os.path.exists(save_directory+'operations/'):
os.makedirs(save_directory+'operations/')
if not os.path.exists(save_directory+'endingdays/'):
os.makedirs(save_directory+'endingdays/')
if not os.path.exists(save_directory+'rewards/'):
os.makedirs(save_directory+'rewards/')
if not os.path.exists(save_directory+'profits/'):
os.makedirs(save_directory+'profits/')
pd.DataFrame(operation_array).to_csv(save_directory+"operations/{}-iteration{}.csv".format(identifier, i),
header=env.get_current_symbols(), index=None)
pd.DataFrame(days_array).to_csv(save_directory+"endingdays/{}-iteration{}.csv".format(identifier, i),
header=env.get_current_symbols(), index=None)
pd.DataFrame(rewards_array).to_csv(save_directory+"rewards/{}-iteration{}.csv".format(identifier, i),
header=env.get_current_symbols(), index=None)
pd.DataFrame(test_total_profits).to_csv(save_directory+"profits/{}.csv".format(identifier),
index=None)
mean_test_reward.reset_states()
a = [(-0.15989657, 'open'),
(-0.19772957, 'close'),
(0.11944062, 'low'),
(-0.07273539, 'high'),
(-0.05442702, 'volume'),
(0.08966919, 'Accounts Payable'),
(0.021010576, 'Accounts Receivable'),
(-0.027171114, "Add'l income/expense items"),
(0.01937542, 'After Tax ROE'),
(0.22196895, 'Capital Expenditures'),
(0.02117831, 'Capital Surplus'),
(-0.062394377, 'Cash Ratio'),
(-0.1270617, 'Cash and Cash Equivalents'),
(0.047790807, 'Changes in Inventories'),
(0.07777014, 'Common Stocks'),
(0.106798984, 'Cost of Revenue'),
(-0.021594241, 'Current Ratio'),
(-0.015595697, 'Deferred Asset Charges'),
(-0.031983275, 'Deferred Liability Charges'),
(0.1295437, 'Depreciation'),
(-0.19305989, 'Earnings Before Interest and Tax'),
(-0.10321899, 'Earnings Before Tax'),
(0.0041140895, 'Effect of Exchange Rate'),
(0.23839177, 'Equity Earnings/Loss Unconsolidated Subsidiary'),
(-0.20699006, 'Fixed Assets'),
(0.018536765, 'Goodwill'),
(0.03508943, 'Gross Margin'),
(0.10541249, 'Gross Profit'),
(0.19734253, 'Income Tax'),
(0.13355829, 'Intangible Assets'),
(-0.17602208, 'Interest Expense'),
(-0.0025003892, 'Inventory'),
(-0.081173904, 'Investments'),
(0.028858133, 'Liabilities'),
(-0.015410017, 'Long-Term Debt'),
(-0.059088428, 'Long-Term Investments'),
(-0.060415223, 'Minority Interest'),
(0.052735303, 'Misc. Stocks'),
(-0.16270235, 'Net Borrowings'),
(0.30459827, 'Net Cash Flow'),
(-0.051255636, 'Net Cash Flow-Operating'),
(-0.17853276, 'Net Cash Flows-Financing'),
(-0.012475527, 'Net Cash Flows-Investing'),
(-0.007237089, 'Net Income'),
(0.030531632, 'Net Income Adjustments'),
(-0.044338573, 'Net Income Applicable to Common Shareholders'),
(-0.019390859, 'Net Income-Cont. Operations'),
(-0.042641692, 'Net Receivables'),
(-0.19151591, 'Non-Recurring Items'),
(0.2126374, 'Operating Income'),
(-0.115024894, 'Operating Margin'),
(-0.12695658, 'Other Assets'),
(-0.11095659, 'Other Current Assets'),
(-0.21254101, 'Other Current Liabilities'),
(0.08447041, 'Other Equity'),
(0.017648121, 'Other Financing Activities'),
(-0.08386561, 'Other Investing Activities'),
(0.10737695, 'Other Liabilities'),
(-0.09191378, 'Other Operating Activities'),
(-0.057637196, 'Other Operating Items'),
(-0.023080353, 'Pre-Tax Margin'),
(-0.048542995, 'Pre-Tax ROE'),
(0.13798867, 'Profit Margin'),
(0.02022573, 'Quick Ratio'),
(-0.05218117, 'Research and Development'),
(-0.32454595, 'Retained Earnings'),
(-0.01603744, 'Sale and Purchase of Stock'),
(-0.07250982, 'Sales, General and Admin.'),
(0.02872207, 'Short-Term Debt / Current Portion of Long-Term Debt'),
(0.29647112, 'Short-Term Investments'),
(-0.0073608444, 'Total Assets'),
(-0.07006701, 'Total Current Assets'),
(-0.014121261, 'Total Current Liabilities'),
(0.011505632, 'Total Equity'),
(0.16001107, 'Total Liabilities'),
(0.010682229, 'Total Liabilities & Equity'),
(0.0726818, 'Total Revenue'),
(-0.07457304, 'Treasury Stock'),
(0.21740438, 'Earnings Per Share'),
(-0.2218255, 'Estimated Shares Outstanding')]
def myFunc(x):
return abs(x[0])
a.sort(key=myFunc)
def myFunc(x):
return abs(x[0])
a.sort(key=myFunc)
positive_list = [x>0 for x,y in a[-15:]]
x = [abs(x) for x,y in a[-15:]]
y = [y for x,y in a[-15:]]
plt.xlabel('Absolute coefficient')
plt.title('Logistic reggression coefficients')
plot = plt.barh(y,x, alpha=0.5)
for i in range(len(positive_list)):
if positive_list[i]:
plot[i].set_color('g')
else:
plot[i].set_color('r')
import matplotlib.patches as mpatches
red_patch = mpatches.Patch(color='r', label='Negative')
green_patch = mpatches.Patch(color='g', label='Positive')
plt.legend(handles=[green_patch,red_patch], loc="lower right")
plt.tight_layout()
plt.savefig('lr_coefficients.png', dpi=1200)
positive_list
```
| github_jupyter |
## 知识点梳理
1. 相关概念(生成模型、判别模型)
2. 先验概率、条件概率
3. 贝叶斯决策理论
4. 贝叶斯定理公式
5. 极值问题情况下的每个类的分类概率
6. 下溢问题如何解决
7. 零概率问题如何解决?
8. 优缺点
9. sklearn参数详解,Python绘制决策树
### sklearn接口
```
from sklearn.naive_bayes import GaussianNB
from sklearn.datasets import load_iris
import pandas as pd
from sklearn.model_selection import train_test_split
iris = load_iris()
X_train, X_test, y_train, y_test = train_test_split(iris.data, iris.target, test_size=0.2)
clf = GaussianNB().fit(X_train, y_train)
print ("Classifier Score:", clf.score(X_test, y_test))
```
<table>
<thead>
<tr>
<th align="center">编号</th>
<th align="center">色泽</th>
<th align="center">根蒂</th>
<th align="center">敲声</th>
<th align="center">纹理</th>
<th align="center">脐部</th>
<th align="center">触感</th>
<th align="center">好瓜</th>
</tr>
</thead>
<tbody><tr>
<td align="center">1</td>
<td align="center">青绿</td>
<td align="center">蜷缩</td>
<td align="center">浊响</td>
<td align="center">清晰</td>
<td align="center">凹陷</td>
<td align="center">硬滑</td>
<td align="center">是</td>
</tr>
<tr>
<td align="center">2</td>
<td align="center">乌黑</td>
<td align="center">蜷缩</td>
<td align="center">沉闷</td>
<td align="center">清晰</td>
<td align="center">凹陷</td>
<td align="center">硬滑</td>
<td align="center">是</td>
</tr>
<tr>
<td align="center">3</td>
<td align="center">乌黑</td>
<td align="center">蜷缩</td>
<td align="center">浊响</td>
<td align="center">清晰</td>
<td align="center">凹陷</td>
<td align="center">硬滑</td>
<td align="center">是</td>
</tr>
<tr>
<td align="center">4</td>
<td align="center">青绿</td>
<td align="center">蜷缩</td>
<td align="center">沉闷</td>
<td align="center">清晰</td>
<td align="center">凹陷</td>
<td align="center">硬滑</td>
<td align="center">是</td>
</tr>
<tr>
<td align="center">5</td>
<td align="center">浅白</td>
<td align="center">蜷缩</td>
<td align="center">浊响</td>
<td align="center">清晰</td>
<td align="center">凹陷</td>
<td align="center">硬滑</td>
<td align="center">是</td>
</tr>
<tr>
<td align="center">6</td>
<td align="center">青绿</td>
<td align="center">稍蜷</td>
<td align="center">浊响</td>
<td align="center">清晰</td>
<td align="center">稍凹</td>
<td align="center">软粘</td>
<td align="center">是</td>
</tr>
<tr>
<td align="center">7</td>
<td align="center">乌黑</td>
<td align="center">稍蜷</td>
<td align="center">浊响</td>
<td align="center">稍糊</td>
<td align="center">稍凹</td>
<td align="center">软粘</td>
<td align="center">是</td>
</tr>
<tr>
<td align="center">8</td>
<td align="center">乌黑</td>
<td align="center">稍蜷</td>
<td align="center">浊响</td>
<td align="center">清晰</td>
<td align="center">稍凹</td>
<td align="center">硬滑</td>
<td align="center">是</td>
</tr>
<tr>
<td align="center">9</td>
<td align="center">乌黑</td>
<td align="center">稍蜷</td>
<td align="center">沉闷</td>
<td align="center">稍糊</td>
<td align="center">稍凹</td>
<td align="center">硬滑</td>
<td align="center">否</td>
</tr>
<tr>
<td align="center">10</td>
<td align="center">青绿</td>
<td align="center">硬挺</td>
<td align="center">清脆</td>
<td align="center">清晰</td>
<td align="center">平坦</td>
<td align="center">软粘</td>
<td align="center">否</td>
</tr>
<tr>
<td align="center">11</td>
<td align="center">浅白</td>
<td align="center">硬挺</td>
<td align="center">清脆</td>
<td align="center">模糊</td>
<td align="center">平坦</td>
<td align="center">硬滑</td>
<td align="center">否</td>
</tr>
<tr>
<td align="center">12</td>
<td align="center">浅白</td>
<td align="center">蜷缩</td>
<td align="center">浊响</td>
<td align="center">模糊</td>
<td align="center">平坦</td>
<td align="center">软粘</td>
<td align="center">否</td>
</tr>
<tr>
<td align="center">13</td>
<td align="center">青绿</td>
<td align="center">稍蜷</td>
<td align="center">浊响</td>
<td align="center">稍糊</td>
<td align="center">凹陷</td>
<td align="center">硬滑</td>
<td align="center">否</td>
</tr>
<tr>
<td align="center">14</td>
<td align="center">浅白</td>
<td align="center">稍蜷</td>
<td align="center">沉闷</td>
<td align="center">稍糊</td>
<td align="center">凹陷</td>
<td align="center">硬滑</td>
<td align="center">否</td>
</tr>
<tr>
<td align="center">15</td>
<td align="center">乌黑</td>
<td align="center">稍蜷</td>
<td align="center">浊响</td>
<td align="center">清晰</td>
<td align="center">稍凹</td>
<td align="center">软粘</td>
<td align="center">否</td>
</tr>
<tr>
<td align="center">16</td>
<td align="center">浅白</td>
<td align="center">蜷缩</td>
<td align="center">浊响</td>
<td align="center">模糊</td>
<td align="center">平坦</td>
<td align="center">硬滑</td>
<td align="center">否</td>
</tr>
<tr>
<td align="center">17</td>
<td align="center">青绿</td>
<td align="center">蜷缩</td>
<td align="center">沉闷</td>
<td align="center">稍糊</td>
<td align="center">稍凹</td>
<td align="center">硬滑</td>
<td align="center">否</td>
</tr>
</tbody></table>
### 1. 相关概念
生成模型:在概率统计理论中, 生成模型是指能够随机生成观测数据的模型,尤其是在给定某些隐含参数的条件下。它给观测值和标注数据序列指定一个联合概率分布。在机器学习中,生成模型可以用来直接对数据建模(例如根据某个变量的概率密度函数进行数据采样),也可以用来建立变量间的条件概率分布。条件概率分布可以由生成模型根据贝叶斯定理形成。常见的基于生成模型算法有高斯混合模型和其他混合模型、隐马尔可夫模型、随机上下文无关文法、朴素贝叶斯分类器、AODE分类器、潜在狄利克雷分配模型、受限玻尔兹曼机
<br>
举例:要确定一个瓜是好瓜还是坏瓜,用判别模型的方法是从历史数据中学习到模型,然后通过提取这个瓜的特征来预测出这只瓜是好瓜的概率,是坏瓜的概率。
<br>
判别模型: 在机器学习领域判别模型是一种对未知数据 y 与已知数据 x 之间关系进行建模的方法。判别模型是一种基于概率理论的方法。已知输入变量 x ,判别模型通过构建条件概率分布 P(y|x) 预测 y 。常见的基于判别模型算法有逻辑回归、线性回归、支持向量机、提升方法、条件随机场、人工神经网络、随机森林、感知器
举例:利用生成模型是根据好瓜的特征首先学习出一个好瓜的模型,然后根据坏瓜的特征学习得到一个坏瓜的模型,然后从需要预测的瓜中提取特征,放到生成好的好瓜的模型中看概率是多少,在放到生产的坏瓜模型中看概率是多少,哪个概率大就预测其为哪个。
<br>
生成模型是所有变量的全概率模型,而判别模型是在给定观测变量值前提下目标变量条件概率模型。因此生成模型能够用于模拟(即生成)模型中任意变量的分布情况,而判别模型只能根据观测变量得到目标变量的采样。判别模型不对观测变量的分布建模,因此它不能够表达观测变量与目标变量之间更复杂的关系。因此,生成模型更适用于无监督的任务,如分类和聚类。
### 2. 先验概率、条件概率
条件概率: 就是事件A在事件B发生的条件下发生的概率。条件概率表示为P(A|B),读作“A在B发生的条件下发生的概率”。
<br>
先验概率: 在贝叶斯统计中,某一不确定量 p 的先验概率分布是在考虑"观测数据"前,能表达 p 不确定性的概率分布。它旨在描述这个不确定量的不确定程度,而不是这个不确定量的随机性。这个不确定量可以是一个参数,或者是一个隐含变量。
<br>
后验概率: 在贝叶斯统计中,一个随机事件或者一个不确定事件的后验概率是在考虑和给出相关证据或数据后所得到的条件概率。同样,后验概率分布是一个未知量(视为随机变量)基于试验和调查后得到的概率分布。“后验”在本文中代表考虑了被测试事件的相关证据。
<br>
通过上述西瓜的数据集来看
条件概率,就是在条件为瓜的颜色是青绿的情况下,瓜是好瓜的概率
先验概率,就是常识、经验、统计学所透露出的“因”的概率,即瓜的颜色是青绿的概率。
后验概率,就是在知道“果”之后,去推测“因”的概率,也就是说,如果已经知道瓜是好瓜,那么瓜的颜色是青绿的概率是多少。后验和先验的关系就需要运用贝叶斯决策理论来求解。
### 3. 贝叶斯决策理论
贝叶斯决策论是概率框架下实施决策的基本方法,对分类任务来说,在所有相关概率都已知的理想情形下,贝叶斯决策论考虑如何基于这些概率和误判损失来选择最优的类别标记。
假设有N种可能标记, $λ_{ij}$是将类$c_j$误分类为$c_i$所产生的损失,基于后验概率$ P(c_i | x)$ 可以获得样本x分类为$c_i$所产生的期望损失 ,即在样本x上的条件风险:
$$R(c_i|\mathbf{x}) = \sum_{j=1}^N \lambda_{ij} P(c_j|\mathbf{x})$$
我们的任务是寻找一个判定准则 $h:X→Y$以最小化总体风险
$$R(h)= \mathbb{E}_x [R(h(\mathbf(x)) | \mathbf(x))]$$
显然,对每个样本x,若h能最小化条件风险 $R(h((x))|(x))$,则总体风险R(h)也将被最小化。这就产生了贝叶斯判定准则:为最小化总体风险,只需要在每个样本上选择那个能使条件风险R(c|x)最小的类别标记,即:
$$h^* (x) = argmin_{c\in y} R(c|\mathbf{x})$$
此时,h 称作贝叶斯最优分类器,与之对应的总体风险R(h )称为贝叶斯风险,1-R(h*)反映了分类器能达到的最好性能,即机器学习所产生的模型精度的上限。
具体来说,若目标是最小化分类错误率(对应0/1损失),则$λ_{ij}$可以用0/1损失改写,得到条件风险和最小化分类错误率的最优分类器分别为:
$$R(c|\mathbf{x}) = 1- P(c|\mathbf{x})$$
$$h^*(x) = argmax_{c\in \mathcal{Y}} P(c|\mathbf{x})$$
即对每个样本x,选择能使后验概率P(c|x)最大的类别标识。
获得后验概率的两种方法:
1. 判别式模型:给定x,可以通过直接建模P(c|x)来预测c。
2. 生成模型:先对联合分布p(x, c)建模,然后再有此获得P(c|x)。
### 4. 贝叶斯公式
对生成模型来说,必然考虑:
$$P(c|x) = \frac{P(x,c)}{P(x)} = \frac{P(c) P(x|c)}{P(x)}$$
其中P(c)是“先验概率”;
P(x|c)是样本x对于类标记c的类条件概率,或称为“似然”;
P(x)是用于归一化的“证据”因子。
上式即为贝叶斯公式。
可以将其看做$$P(类别|特征) = \frac{P(特征,类别)}{P(特征)} = \frac{P(类别) P(特征|类别)}{P(特征)}$$
对类条件概率P(x|c)来说,直接根据样本出现的频率来估计将会遇到严重的困难,所以引入了极大似然估计。
#### 极大似然估计
估计类条件概率有一种常用的策略就是先假定其具有某种确定的概率分布形式,再基于训练样本对概率分布的参数进行估计。假设P(x|c)具有某种确定的形式并且被参数$θ_c$ 唯一确定,则我们的任务就是利用训练结D估计参数 $θ_c$。为了明确期间,我们将P(x|c)记为$p(x|θc)$.
举个通俗的例子:假设一个袋子装有白球与红球,比例未知,现在抽取10次(每次抽完都放回,保证事件独立性),假设抽到了7次白球和3次红球,在此数据样本条件下,可以采用最大似然估计法求解袋子中白球的比例(最大似然估计是一种“模型已定,参数未知”的方法)。当然,这种数据情况下很明显,白球的比例是70%,但如何通过理论的方法得到这个答案呢?一些复杂的条件下,是很难通过直观的方式获得答案的,这时候理论分析就尤为重要了,这也是学者们为何要提出最大似然估计的原因。我们可以定义从袋子中抽取白球和红球的概率如下:

x1为第一次采样,x2为第二次采样,f为模型, theta为模型参数
其中$\theta$是未知的,因此,我们定义似然L为:

L为似然的符号
两边取ln,取ln是为了将右边的乘号变为加号,方便求导。

两边取ln的结果,左边的通常称之为对数似然。
这是平均对数似然
最大似然估计的过程,就是找一个合适的theta,使得平均对数似然的值为最大。因此,可以得到以下公式:
最大似然估计的公式
这里讨论的是2次采样的情况,当然也可以拓展到多次采样的情况:

最大似然估计的公式(n次采样)
我们定义M为模型(也就是之前公式中的f),表示抽到白球的概率为theta,而抽到红球的概率为(1-theta),因此10次抽取抽到白球7次的概率可以表示为:

10次抽取抽到白球7次的概率
将其描述为平均似然可得:
10次抽取抽到白球7次的平均对数似然,抽球的情况比较简单,可以直接用平均似然来求解
那么最大似然就是找到一个合适的theta,获得最大的平均似然。因此我们可以对平均似然的公式对theta求导,并另导数为0。

求导过程
由此可得,当抽取白球的概率为0.7时,最可能产生10次抽取抽到白球7次的事件。
以上就用到了最大似然估计的思想
令Dc表示训练集D中第c类样本组成的集合,假设这些集合是独立同分布的,则对参数θcθc对于数据集Dc的似然是:
$$P(D_c|\theta_c) = \prod P(\mathbf{x}|\theta_c)$$
对$θ_c$进行激发似然估计买就是去寻找能最大化似然函数的参数值$θ_c$.直观上,极大似然估计是在试图在$θ_c$的所有可能的去职中,找到一个能使数据出现最大“可能性”的最大值。
上面的式子中的连乘操作容易造成下溢,通常使用对数似然:
$$L(\theta_c) = \log P(D_c| \theta_c) = \sum_{x\in D_c} \log P(x|\theta_c)$$
此时,参数$θ_c$的极大似然估计$\hat{\theta_c}$为
$\hat{\theta_c} = argmax_{\theta_c} LL(\theta_c)$
例如,在连续属性的情形下,假设概率密度函数$p(x|c) \sim \mathcal{N}(\mu_c , \sigma^2)$,则参数$μ_c$和$σ_2$的极大似然估计为:
$\hat{\mu_c} = \frac{1}{|D_c|} \sum_{x\in D_c} x$
$\hat{\sigma_c}^2 = \frac{1}{|D_c|} \sum_{x\in D_c} (x-\hat{\mu_c} )(x-\hat{\mu_c} ^T)$
也就是说通过极大似然发得到的额正态分布均值就是样本均值,方差就是$(x-\hat{\mu_c} )(x-\hat{\mu_c} ^T)$的均值。这显然是一个符合只觉得结果,在离散属性情形下,也可以通过类似的方法来估计类条件概率。
需要注意的是这种方法虽然能够使类条件概率估计变得简单,但是估计结果准确性严重依赖于所假设的概率分布形式是否符合潜在的真实数据分布。在显示生活中往往需要应用任务本身的经验知识,“猜测”则会导致误导性的结果。
贝叶斯分类器的训练过程就是参数估计。总结最大似然法估计参数的过程,一般分为以下四个步骤:
```
1.写出似然函数;
2.对似然函数取对数,并整理;
3.求导数,令偏导数为0,得到似然方程组;
4.解似然方程组,得到所有参数即为所求。
```
#### 朴素贝叶斯分类器
基于贝叶斯公式来估计后验概率P(c|x)主要困难在于类条件概率P(x|c)是所有属性上的联合概率,难以从有限的训练样本直接估计而得。
基于有限训练样本直接计算联合概率,在计算上将会遭遇组合爆炸问题;在数据上将会遭遇样本稀疏问题;属性越多,问题越严重。
为了避开这个障碍,朴素贝叶斯分类器采用了$“属性条件独立性假设”$:对已知类别,假设所有属性相互独立。换言之,假设每个属性独立的对分类结果发生影响相互独立。
回答西瓜的例子就可以认为{色泽 根蒂 敲声 纹理 脐部 触感}这些属性对西瓜是好还是坏的结果所产生的影响相互独立。
基于条件独立性假设,对于多个属性的后验概率可以写成:
$$P(c|\mathbf{x}) = \frac{P(C)P(\mathbf{x}|c)}{P(\mathbf{x})} = \frac{P(c)}{P(\mathbf{x})}\prod_{i=1}^d P(x_i|c)$$
d为属性数目,$x_i$是x在第i个属性上取值。
对于所有的类别来说P(x)相同,基于极大似然的贝叶斯判定准则有朴素贝叶斯的表达式:
$$h_{nb}(\mathbf{x}) = \arg max_{c\in \mathcal{Y}}P(c)\prod_{i=1}^d P(x_i|c) \quad (1)$$
### 5. 极值问题情况下的每个类的分类概率
极值问题
很多时候遇到求出各种目标函数(object function)的最值问题(最大值或者最小值)。关于函数最值问题,其实在高中的时候我们就已经了解不少,最经典的方法就是:直接求出极值点。这些极值点的梯度为0。若极值点唯一,则这个点就是代入函数得出的就是最值;若极值点不唯一,那么这些点中,必定存在最小值或者最大值(去除函数的左右的最端点),所以把极值代入函数,经对比后可得到结果。
请注意:并不一定所有函数的极值都可以通过设置导数为0的方式求 出。也就是说,有些问题中当我们设定导数为0时,未必能直接计算出满足导数为0的点(比如逻辑回归模型),这时候就需要利用数值计算相关的技术(最典型为梯度下降法,牛顿法……)。
### 6. 下溢问题如何解决
数值下溢问题:是指计算机浮点数计算的结果小于可以表示的最小数,因为计算机的能力有限,当数值小于一定数时,其无法精确保存,会造成数值的精度丢失,由上述公式可以看到,求概率时多个概率值相乘,得到的结果往往非常小;因此通常采用取对数的方式,将连乘转化为连加,以避免数值下溢。
### 7. 零概率问题如何解决?
零概率问题,就是在计算实例的概率时,如果某个量x,在观察样本库(训练集)中没有出现过,会导致整个实例的概率结果是0.
在实际的模型训练过程中,可能会出现零概率问题(因为先验概率和反条件概率是根据训练样本算的,但训练样本数量不是无限的,所以可能出现有的情况在实际中存在,但在训练样本中没有,导致为0的概率值,影响后面后验概率的计算),即便可以继续增加训练数据量,但对于有些问题来说,数据怎么增多也是不够的。这时我们说模型是不平滑的,我们要使之平滑,一种方法就是将训练(学习)的方法换成贝叶斯估计。
现在看一个示例,及$P(敲声=清脆|好瓜=是)=\frac{8}{0}=0$
不论样本的其他属性如何,分类结果都会为“好瓜=否”,这样显然不太合理。
朴素贝叶斯算法的先天缺陷:其他属性携带的信息被训练集中某个分类下未出现的属性值“抹去”,造成预测出来的概率绝对为0。为了拟补这一缺陷,前辈们引入了拉普拉斯平滑的方法:对先验概率的分子(划分的计数)加1,分母加上类别数;对条件概率分子加1,分母加上对应特征的可能取值数量。这样在解决零概率问题的同时,也保证了概率和依然为1:
$$P(c) = \frac{{|{D_c}|}}{{|D|}} \to P(c) = \frac{{|{D_c}| + 1}}{{|D| + N}}$$
$$P({x_i}|c) = \frac{{|{D_{{x_i}|c}}|}}{{|{D_c}|}} \to P({x_i}|c) = \frac{{|{D_{{x_i}|c}}| + 1}}{{|{D_c}| + {N_i}}}$$
其中,N表示数据集中分类标签,$N_i$表示第$i$个属性的取值类别数,|D|样本容量,$|D_c|$表示类别c的记录数量,${|{D_{{x_i}|c}}|}$表示类别c中第i个属性取值为$x_i$的记录数量。
将这两个式子应用到上面的计算过程中,就可以弥补朴素贝叶斯算法的这一缺陷问题。
用西瓜的数据来看,当我们计算
P(好瓜=是)时,样本有17个,所以|D| = 17,N,好瓜标签可以分为{是,否}两类,所以N=2,(好瓜=是)的样本个数有8个,所以这里$|D_c|$=8。
综上,根据拉普拉斯平滑后有 $$P(好瓜=是) = \frac{{|{D_c}| + 1}}{{|D| + N}} = \frac{{|{8}| + 1}}{{|17| + 2}}$$
P(色泽=青绿|好瓜=是)时,色泽青绿的样本有8个,所以|D_c| = 8,N,色泽标签可以分为{青绿,浅白,乌黑}三类,所以N=3,(好瓜=是)的样本个数有3个,所以这里$|D_{c,x_i}|$=3。
综上,根据拉普拉斯平滑后有$$P(色泽=青绿|好瓜=是)= \frac{{|{D_{{x_i}|c}}| + 1}}{{|{D_c}| + {N_i}}}=\frac{{|{3}}| + 1}{{|{8}| + {3}}}$$
同理,分析可知,之前不合理的$P(敲声=清脆|好瓜=是)=\frac{8}{0}=0$在进行拉普拉斯平滑后为$$ P(敲声=清脆|好瓜=是)= \frac{{|{D_{{x_i}|c}}| + 1}}{{|{D_c}| + {N_i}}}=\frac{{|{0}}| + 1}{{|{8}| + {3}}}$$显然结果不是0,使结果变得合理。
### 8. sklearn参数详解
1. 高斯朴素贝叶斯算法是假设特征的可能性(即概率)为高斯分布。
```
class sklearn.naive_bayes.GaussianNB(priors=None)
参数:
priors:先验概率大小,如果没有给定,模型则根据样本数据自己计算(利用极大似然法)。
var_smoothing:可选参数,所有特征的最大方差
属性:
class_prior_:每个样本的概率
class_count:每个类别的样本数量
classes_:分类器已知的标签类型
theta_:每个类别中每个特征的均值
sigma_:每个类别中每个特征的方差
epsilon_:方差的绝对加值方法
# 贝叶斯的方法和其他模型的方法一致。
fit(X,Y):在数据集(X,Y)上拟合模型。
get_params():获取模型参数。
predict(X):对数据集X进行预测。
predict_log_proba(X):对数据集X预测,得到每个类别的概率对数值。predict_proba(X):对数据集X预测,得到每个类别的概率。
score(X,Y):得到模型在数据集(X,Y)的得分情况。
```
根据李航老师的代码构建自己的朴素贝叶斯模型
这里采用GaussianNB 高斯朴素贝叶斯,概率密度函数为
$$P(x_{i}|y_{k}) = \frac{1}{\sqrt{2\pi\sigma_{y_{k}}^{2}}}exp( -\frac{(x_{i}-\mu_{y_{k}})^2} {2\sigma_{y_{k}}^{2}} )$$
数学期望:$\mu$
方差:$\sigma ^2=\frac{1}{n}\sum_i^n(x_i-\overline x)^2$
https://github.com/fengdu78/lihang-code/blob/master/%E7%AC%AC04%E7%AB%A0%20%E6%9C%B4%E7%B4%A0%E8%B4%9D%E5%8F%B6%E6%96%AF/4.NaiveBayes.ipynb
```
import math
class NaiveBayes:
def __init__(self):
self.model = None
# 数学期望
@staticmethod
def mean(X):
# 根据期望的公式直接返回(≈1 line)
return None
# 标准差(方差)
def stdev(self, X):
# 根据期望的公式计算(≈2 line)
avg = None
stdev = None
return stdev
# 概率密度函数
def gaussian_probability(self, x, mean, stdev):
# 根据概率密度公式计算(≈2 line)
exponent = None
probability = None
return probability
# 处理X_train
def summarize(self, train_data):
summaries = [(self.mean(i), self.stdev(i)) for i in zip(*train_data)]
return summaries
# 分类别求出数学期望和标准差
def fit(self, X, y):
labels = list(set(y))
data = {label: [] for label in labels}
for f, label in zip(X, y):
data[label].append(f)
self.model = {
label: self.summarize(value)
for label, value in data.items()
}
return 'gaussianNB train done!'
# 计算概率
def calculate_probabilities(self, input_data):
# summaries:{0.0: [(5.0, 0.37),(3.42, 0.40)], 1.0: [(5.8, 0.449),(2.7, 0.27)]}
# input_data:[1.1, 2.2]
probabilities = {}
for label, value in self.model.items():
probabilities[label] = 1
for i in range(len(value)):
# 获取计算好的均值和方差(≈1 line)
mean, stdev = None
# 根据上面的描述计算概率(≈1 line)
probabilities[label] *= None
return probabilities
# 类别
def predict(self, X_test):
# {0.0: 2.9680340789325763e-27, 1.0: 3.5749783019849535e-26}
label = sorted(
self.calculate_probabilities(X_test).items(),
key=lambda x: x[-1])[-1][0]
return label
# 计算得分
def score(self, X_test, y_test):
right = 0
for X, y in zip(X_test, y_test):
label = self.predict(X)
if label == y:
right += 1
return right / float(len(X_test))
model = NaiveBayes()
model.fit(X_train, y_train)
print(model.predict([4.4, 3.2, 1.3, 0.2]))
model.score(X_test, y_test)
```
### 9. 优缺点
优点
1. 朴素贝叶斯模型有稳定的分类效率。
2. 对小规模的数据表现很好,能处理多分类任务,适合增量式训练,尤其是数据量超出内存时,可以一批批的去增量训练。
3. 对缺失数据不太敏感,算法也比较简单,常用于文本分类。
缺点:
1. 理论上,朴素贝叶斯模型与其他分类方法相比具有最小的误差率。但是实际上并非总是如此,这是因为朴素贝叶斯模型给定输出类别的情况下,假设属性之间相互独立,这个假设在实际应用中往往是不成立的,在属性个数比较多或者属性之间相关性较大时,分类效果不好。而在属性相关性较小时,朴素贝叶斯性能最为良好。对于这一点,有半朴素贝叶斯之类的算法通过考虑部分关联性适度改进。
2. 需要知道先验概率,且先验概率很多时候取决于假设,假设的模型可以有很多种,因此在某些时候会由于假设的先验模型的原因导致预测效果不佳。
3. 由于我们是通过先验和数据来决定后验的概率从而决定分类,所以分类决策存在一定的错误率。
4. 对输入数据的表达形式很敏感。
### 10. 参考文献
西瓜书
https://samanthachen.github.io/2016/08/05/%E6%9C%BA%E5%99%A8%E5%AD%A6%E4%B9%A0_%E5%91%A8%E5%BF%97%E5%8D%8E_%E7%AC%94%E8%AE%B07/
https://www.jianshu.com/p/f1d3906e4a3e
https://zhuanlan.zhihu.com/p/66117273
https://zhuanlan.zhihu.com/p/39780650
https://blog.csdn.net/zrh_CSDN/article/details/81007851
| github_jupyter |
### Package Imports
```
#Package imports
import pandas as pd
import numpy as np
import calendar
import matplotlib.pyplot as plt
import seaborn as sns
import plotly
import utils
sns.set_style('darkgrid')
from pandas_datareader import data #Package for pulling data from the web
from datetime import date
from fbprophet import Prophet
```
### Pulling in Data
```
#Pulling in the tickers and economic predicators we're interested in
#ONLN = Online Retail ETF, FTXD = Nasdaq Retail ETF, XLY = S&P Consumer Discretionary ETF,
#IYW= iShares US Tech ETF, AMZN = Amazon, EBAY = Ebay, FB = Facebook
#JETS = Global Airline ETF (Recovery trade), XOP = S&P O&G Exploration and Production ETF (Recovery Trade)
#GOVT = US T-Bond ETF, CL=F = Crude, GC=F = Gold, SI=F = Silver, HG=F = Copper, ^VIX = CBOE Vix
tickers = ['ETSY','SPY','ONLN','FTXD','XLY','IHE','AMZN','EBAY','FB','JETS','XOP','GOVT','CL=F','GC=F','SI=F','HG=F','^VIX']
#Getting the most up to date data we can
today = date.today() #This function automatically updates with today's data
#Pulling the timeseries data directly from yahoo finance into a dataframe
etf_df= data.DataReader(tickers,
start='2020-06-01', #start date
end = today, #charting up to today's date
data_source='yahoo')['Adj Close'] #obtaining price at close
#Dropping missing vals
etf_df = etf_df.dropna()
#Checking the 5 most recent values
etf_df.head(5)
#Creating correlation matrix
corr_mat = etf_df.corr()
fig, ax = plt.subplots(figsize = (10,10))
sns.heatmap(corr_mat)
#Quantifying the Correlation Matrix
i = 12
columns = corr_mat.nlargest(i,'ETSY')['ETSY'].index
corrmat = np.corrcoef(etf_df[columns].values.T)
heatmap = sns.heatmap(corrmat,
cbar=True,
annot=True,
xticklabels = columns.values,
yticklabels = columns.values)
ax.set_title('Correlation Matrix for ETSY')
plt.show()
```
### Forecasting Predicator Variables Individually
```
#Creating dfs to model with
#SPY, XOP, ONLN
#Regressor 1 (SPY)
reg1_prophet = pd.DataFrame()
reg1_prophet['ds'] = etf_df.index
reg1_prophet['y'] = etf_df['SPY'].values
#Regressor 2 (ONLN)
reg2_prophet = pd.DataFrame()
reg2_prophet['ds'] = etf_df.index
reg2_prophet['y'] = etf_df['ONLN'].values
#Regressor 3 (XOP)
reg3_prophet = pd.DataFrame()
reg3_prophet['ds'] = etf_df.index
reg3_prophet['y'] = etf_df['XOP'].values
```
### Fitting Models
```
#Pred 1
reg1_m = Prophet()
reg1_m.fit(reg1_prophet)
reg1_future = reg1_m.make_future_dataframe(freq='m', periods=3);
#Pred 2
reg2_m = Prophet()
reg2_m.fit(reg2_prophet)
reg2_future = reg2_m.make_future_dataframe(freq='m', periods=3);
#Pred 3
reg3_m = Prophet()
reg3_m.fit(reg3_prophet)
reg3_future = reg3_m.make_future_dataframe(freq='m', periods=3);
```
### Creating Forecast for Next 6 Months of Predicator Variables
```
#Defining length of forecast
future_pred_length = reg1_m.make_future_dataframe(freq='m',periods = 3)
#Pred 1
reg1_forecast = reg1_m.predict(future_pred_length)
reg1_pred = pd.DataFrame(reg1_forecast['trend'].values)
#Pred 2
reg2_forecast = reg2_m.predict(future_pred_length)
reg2_pred = pd.DataFrame(reg2_forecast['trend'].values)
#Pred 3
reg3_forecast = reg3_m.predict(future_pred_length)
reg3_pred = pd.DataFrame(reg3_forecast['trend'].values)
#Combining predicators into one df
frames = [future_pred_length,reg1_pred,reg2_pred,reg3_pred]
predicator_forecast = pd.concat(frames,axis=1)
predicator_forecast.columns = ['ds','Reg_1','Reg_2','Reg_3']
predicator_forecast.head()
```
### Forecasting ETF Performance with Predicator Variables
```
#Creating dfs to train model with (FB Prophet format)
etf_prophet = pd.DataFrame()
etf_prophet['ds'] = etf_df.index
etf_prophet['y'] = etf_df['ETSY'].values
etf_prophet['Reg_1'] = etf_df['SPY'].values
etf_prophet['Reg_2'] = etf_df['ONLN'].values
etf_prophet['Reg_3'] = etf_df['XOP'].values
#Checking Prophet df
etf_prophet.head()
#Defining model, adding additional regressors and fitting model
m = Prophet()
#m.add_regressor('Reg_1')
m.add_regressor('Reg_2')
m.add_regressor('Reg_3')
m.fit(etf_prophet)
#Forecasting the next 90 days
etf_forecast = m.predict(predicator_forecast)
etf_forecast.tail()
#Visualizing forecast
plt.figure(figsize = (20,10))
m.plot(etf_forecast, xlabel='Year',ylabel ='Price [USD]');
plt.title('3 Month ETSY Forecast [ETSY]')
#Plotting forecast components
plt.figure(figsize = (20,10))
m.plot_components(etf_forecast);
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import seaborn as sb
%matplotlib inline
import matplotlib.pyplot as plt
df = pd.read_csv('C:\\\\Users\kimte\\git\\data-analytics-and-science\\exercises\\exercise 1 - loan prediction problem\\data\\train.csv')
df.shape
type(df)
df.info()
df.head()
```
# Missing values identification
```
df.isnull().any()
df[df.isnull().any(axis=1)]
```
# Target value identification and recoding
### target Loan_Status
```
df["Loan_Status"].isnull().value_counts()
df["Loan_Status"].value_counts()
df["Loan_Status_int"] = df["Loan_Status"].astype('category').cat.codes
pd.crosstab(df["Loan_Status"],df["Loan_Status_int"])
```
# Predictive variable identification
Opties om data in dummys om te zetten:
cat_vars=['job','marital','education','default','housing','loan','contact','month','day_of_week','poutcome']
for var in cat_vars:
cat_list='var'+'_'+var
cat_list = pd.get_dummies(data[var], prefix=var)
data1=data.join(cat_list)
data=data1
df["Gender"].astype('category').cat.codes
pd.get_dummies(df["Gender"])
## make list for predictive vars
```
x_selected = []
x_selected
```
### Var Gender
```
df["Gender"].isnull().value_counts()
pd.crosstab(df["Loan_Status"], df["Gender"].isnull())\
,pd.crosstab(df["Loan_Status"], df["Gender"].isnull(),normalize="columns")
import scipy.stats as scs
scs.chi2_contingency(pd.crosstab(df["Loan_Status"], df["Gender"].isnull()))
df["Gender"].value_counts()
pd.crosstab(df["Loan_Status"], df["Gender"], normalize="columns").plot(kind='bar')
df.groupby("Loan_Status")["Gender"].value_counts(normalize=True)
import scipy.stats as scs
scs.chi2_contingency(pd.crosstab(df["Loan_Status"],df["Gender"]))
```
### Var Married
```
df["Married"].isnull().value_counts()
df["Married"].value_counts()
pd.crosstab(df["Loan_Status"], df["Married"], normalize="columns").plot(kind='bar')
df.groupby("Loan_Status")["Married"].value_counts(normalize=True)
import scipy.stats as scs
scs.chi2_contingency(pd.crosstab(df["Loan_Status"], df["Married"]))
df = pd.concat([df, pd.get_dummies(df["Married"], prefix="Married", prefix_sep="_")], axis=1)
df.info()
x_selected += ["Married_No"]
x_selected += ["Married_Yes"]
x_selected
```
### var Dependents
```
df["Dependents"].isnull().value_counts()
df["Dependents"].value_counts()
pd.crosstab(df["Loan_Status"], df["Dependents"], normalize="columns").plot(kind='bar')
df.groupby("Loan_Status")["Dependents"].value_counts(normalize=True)
import scipy.stats as scs
scs.chi2_contingency(pd.crosstab(df["Loan_Status"],df["Dependents"]))
df = pd.concat([df, pd.get_dummies(df["Dependents"], prefix="Dependents", prefix_sep="_")], axis=1)
df.info()
x_selected += ["Dependents_0"]
x_selected += ["Dependents_1"]
x_selected += ["Dependents_2"]
x_selected += ["Dependents_3+"]
x_selected
```
### var Education
```
df["Education"].isnull().value_counts()
df["Education"].value_counts()
pd.crosstab(df["Loan_Status"], df["Education"], normalize="columns").plot(kind='bar')
df.groupby("Loan_Status")["Education"].value_counts(normalize=True)
import scipy.stats as scs
scs.chi2_contingency(pd.crosstab(df["Loan_Status"],df["Education"]))
df = pd.concat([df, pd.get_dummies(df["Education"], prefix="Education", prefix_sep="_")], axis=1)
df.info()
x_selected += ["Education_Graduate"]
x_selected += ["Education_Not Graduate"]
x_selected
```
### var Self_Employed
```
df["Self_Employed"].isnull().value_counts()
pd.crosstab(df["Loan_Status"], df["Self_Employed"].isnull())\
,pd.crosstab(df["Loan_Status"], df["Self_Employed"].isnull(),normalize="columns")
import scipy.stats as scs
scs.chi2_contingency(pd.crosstab(df["Loan_Status"], df["Self_Employed"].isnull()))
df["Self_Employed"].value_counts()
pd.crosstab(df["Loan_Status"], df["Self_Employed"], normalize="columns").plot(kind='bar')
df.groupby("Loan_Status")["Self_Employed"].value_counts(normalize=True)
```
### var ApplicantIncome
```
df["ApplicantIncome"].isnull().value_counts()
df["ApplicantIncome"].min(), df["ApplicantIncome"].max(), df["ApplicantIncome"].mean(), df["ApplicantIncome"].median()
ax = df["ApplicantIncome"].plot.box()
df[["ApplicantIncome","Loan_Status"]].boxplot(by=["Loan_Status"])
df.groupby("Loan_Status")["ApplicantIncome"].median()
df['ApplicantIncome'][(df["Loan_Status"]=='N')].hist(alpha=0.6)
df['ApplicantIncome'][(df["Loan_Status"]=='Y')].hist(alpha=0.6)
import scipy.stats as scs
scs.ttest_rel(df["Loan_Status_int"],df["ApplicantIncome"])
x_selected += ["ApplicantIncome"]
x_selected
```
### var CoapplicantIncome
```
df["CoapplicantIncome"].isnull().value_counts()
df["CoapplicantIncome"].min(), df["CoapplicantIncome"].max(), df["CoapplicantIncome"].mean(), df["CoapplicantIncome"].median()
ax = df["CoapplicantIncome"].plot.box()
df["CoapplicantIncome"][df["CoapplicantIncome"]>0].shape
df["CoapplicantIncome"][df["CoapplicantIncome"]>0].min(), df["CoapplicantIncome"][df["CoapplicantIncome"]>0].max(), df["CoapplicantIncome"][df["CoapplicantIncome"]>0].mean(), df["CoapplicantIncome"][df["CoapplicantIncome"]>0].median()
ax = df["CoapplicantIncome"][df["CoapplicantIncome"]>0].plot.box()
df[["CoapplicantIncome","Loan_Status"]].boxplot(by=["Loan_Status"])
df.groupby("Loan_Status")["CoapplicantIncome"].median()
df['CoapplicantIncome'][(df["Loan_Status"]=='N')].hist(alpha=0.6)
df['CoapplicantIncome'][(df["Loan_Status"]=='Y')].hist(alpha=0.6)
import scipy.stats as scs
scs.ttest_rel(df["Loan_Status_int"],df["CoapplicantIncome"])
x_selected += ["CoapplicantIncome"]
x_selected
```
### var LoanAmount
```
df["LoanAmount"].isnull().value_counts()
df["LoanAmount"].fillna(df["LoanAmount"].mean(), inplace=True)
df["LoanAmount"].isnull().value_counts()
df["LoanAmount"].min(), df["LoanAmount"].max(), df["LoanAmount"].mean(), df["LoanAmount"].median()
ax = df["LoanAmount"].plot.box()
df[["LoanAmount","Loan_Status"]].boxplot(by=["Loan_Status"])
df.groupby("Loan_Status")["LoanAmount"].median()
df['LoanAmount'][(df["Loan_Status"]=='N')].hist(alpha=0.6)
df['LoanAmount'][(df["Loan_Status"]=='Y')].hist(alpha=0.6)
import scipy.stats as scs
scs.ttest_rel(df["Loan_Status_int"],df["LoanAmount"])
x_selected += ["LoanAmount"]
x_selected
```
### Var Loan_Amount_Term
```
df["Loan_Amount_Term"].isnull().value_counts()
df["Loan_Amount_Term"].fillna(df["Loan_Amount_Term"].mean(), inplace=True)
df["Loan_Amount_Term"].isnull().value_counts()
df["Loan_Amount_Term"].min(), df["Loan_Amount_Term"].max(), df["Loan_Amount_Term"].mean(), df["Loan_Amount_Term"].median()
df[["Loan_Amount_Term","Loan_Status"]].boxplot(by=["Loan_Status"])
df.groupby("Loan_Status")["Loan_Amount_Term"].median()
df['Loan_Amount_Term'][(df["Loan_Status"]=='N')].hist(alpha=0.6)
df['Loan_Amount_Term'][(df["Loan_Status"]=='Y')].hist(alpha=0.6)
import scipy.stats as scs
scs.ttest_rel(df["Loan_Amount_Term"],df["CoapplicantIncome"])
```
### var Credit_History
```
df["Credit_History"].isnull().value_counts()
df["Credit_History"].value_counts()
pd.crosstab(df["Loan_Status"], df["Credit_History"], normalize="columns").plot(kind='bar')
df.groupby("Loan_Status")["Credit_History"].value_counts(normalize=True)
import scipy.stats as scs
scs.chi2_contingency(pd.crosstab(df["Loan_Status"],df["Credit_History"]))
df = pd.concat([df, pd.get_dummies(df["Credit_History"], prefix="Credit_History", prefix_sep="_")], axis=1)
df.info()
x_selected += ["Credit_History_0.0"]
x_selected += ["Credit_History_1.0"]
x_selected
```
### var Property_Area
```
df["Property_Area"].isnull().value_counts()
df["Property_Area"].value_counts()
pd.crosstab(df["Loan_Status"], df["Property_Area"], normalize="columns").plot(kind='bar')
df.groupby("Loan_Status")["Property_Area"].value_counts(normalize=True)
import scipy.stats as scs
scs.chi2_contingency(pd.crosstab(df["Loan_Status"],df["Property_Area"]))
df = pd.concat([df, pd.get_dummies(df["Property_Area"], prefix="Property_Area", prefix_sep="_")], axis=1)
df.info()
x_selected += ["Property_Area_Rural"]
x_selected += ["Property_Area_Semiurban"]
x_selected += ["Property_Area_Urban"]
x_selected
```
# Model bouwen
```
x_selected
x=df[x_selected]
y=df["Loan_Status_int"]
x.info()
```
### Full model (own feature selection)
```
import statsmodels.api as sm
logit_model=sm.Logit(y,x)
result=logit_model.fit()
print(result.summary())
```
### Automated feature selection
```
from sklearn import datasets
from sklearn.feature_selection import RFE
from sklearn.linear_model import LogisticRegression
logreg = LogisticRegression()
rfe = RFE(logreg, 6)
rfe = rfe.fit(x, y)
print(rfe.support_)
print(rfe.ranking_)
list_features = np.atleast_1d(rfe.support_)
x_features = x.loc[:,list_features]
x_features.info()
```
### Final model (6 features selected)
```
import statsmodels.api as sm
logit_model=sm.Logit(y,x_features)
result=logit_model.fit()
print(result.summary())
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(x_features, y, test_size=0.3, random_state=0)
from sklearn.linear_model import LogisticRegression
from sklearn import metrics
model = LogisticRegression()
model.fit(x_train, y_train)
```
### Accuracy of final model
```
y_pred = model.predict(x_test)
print('Accuracy of logistic regression classifier on test set: {:.2f}'.format(model.score(x_test, y_test)))
```
### Accuracy in cross validation of final model
```
from sklearn import model_selection
from sklearn.model_selection import cross_val_score
kfold = model_selection.KFold(n_splits=10, random_state=7)
scoring = 'accuracy'
results = model_selection.cross_val_score(model, x_train, y_train, cv=kfold, scoring=scoring)
print("10-fold cross validation average accuracy: %.3f" % (results.mean()))
```
### Confusion matrix
```
from sklearn.metrics import confusion_matrix
confusion_matrix = confusion_matrix(y_test, y_pred)
print(confusion_matrix)
```
*The result is telling us that we have 21+132 correct predictions and 2+30 incorrect predictions.
### Precision, recall, F-measure and support
```
from sklearn.metrics import classification_report
print(classification_report(y_test, y_pred))
```
### ROC curve
```
from sklearn.metrics import roc_auc_score
from sklearn.metrics import roc_curve
logit_roc_auc = roc_auc_score(y_test, model.predict(x_test))
fpr, tpr, thresholds = roc_curve(y_test, model.predict_proba(x_test)[:,1])
plt.figure()
plt.plot(fpr, tpr, label='Logistic Regression (area = %0.2f)' % logit_roc_auc)
plt.plot([0, 1], [0, 1],'r--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic')
plt.legend(loc="lower right")
plt.savefig('Log_ROC')
plt.show()
```
# Prediction using the model
```
x_features.info()
```
### test whether prediction using the model works
```
myvals = np.array([0, 0, 0, 0, 0, 0]).reshape(1, -1)
myvals
model.predict(myvals) # Closer to 0 is higher likelyhood of getting the loan
```
## Test data where loan must be predicted
```
df_test = pd.read_csv('C:\\\\Users\kimte\\git\\data-analytics-and-science\\exercises\\exercise 1 - loan prediction problem\\data\\test.csv')
df_test.shape
type(df_test)
df_test.info()
df_test = pd.concat([df_test, pd.get_dummies(df_test["Married"], prefix="Married", prefix_sep="_")], axis=1)
df_test = pd.concat([df_test, pd.get_dummies(df_test["Dependents"], prefix="Dependents", prefix_sep="_")], axis=1)
df_test = pd.concat([df_test, pd.get_dummies(df_test["Education"], prefix="Education", prefix_sep="_")], axis=1)
df_test = pd.concat([df_test, pd.get_dummies(df_test["Credit_History"], prefix="Credit_History", prefix_sep="_")], axis=1)
df_test = pd.concat([df_test, pd.get_dummies(df_test["Property_Area"], prefix="Property_Area", prefix_sep="_")], axis=1)
list_features = x_features.columns.values.tolist()
x_features_test = df_test.loc[:,list_features]
x_features_test.info()
df_test["loan_predicted"] = model.predict(x_features_test)
df_test["loan_predicted"].value_counts()
```
| github_jupyter |
# Using multi-armed bandits to choose the best model for predicting credit card default
## Dependencies
- [helm](https://github.com/helm/helm)
- [minikube](https://github.com/kubernetes/minikube) --> install 0.25.2
- [s2i](https://github.com/openshift/source-to-image)
- Kaggle account to download data.
- Python packages:
```
!pip install -r requirements.txt
```
## Getting data
Either head to https://www.kaggle.com/uciml/default-of-credit-card-clients-dataset or use the Kaggle API (instructions at https://github.com/Kaggle/kaggle-api) to download the dataset:
```
!kaggle datasets download -d uciml/default-of-credit-card-clients-dataset
!unzip default-of-credit-card-clients-dataset.zip
```
## Load and inspect data
```
import pandas as pd
data = pd.read_csv('UCI_Credit_Card.csv')
data.shape
data.columns
target = 'default.payment.next.month'
data[target].value_counts()
```
Note that we have a class imbalance, so if we use accuracy as the performance measure of a classifier, we need to be able to beat the "dummy" model that classifies every instance as 0 (no default):
```
data[target].value_counts().max()/data.shape[0]
```
## Case study for using multi-armed bandits
In deploying a new ML model, it is rarely the case that the existing (if any) model is decommissioned immediately in favour of the new one. More commonly the new model is deployed alongside the existing one(s) and the incoming traffic is shared between the models.
Typically A/B testing is performed in which traffic is routed between existing models randomly, this is called the experiment stage. After a set period of time performance statistics are calculated and the best-performing model is chosen to serve 100% of the requests while the other model(s) are decommissioned.
An alternative method is to route traffic dynamically to the best performing model using multi-armed bandits. This avoids the opportunity cost of consistently routing a lot of traffic to the worst performing model(s) during an experiment as in A/B testing.
This notebook is a case study in deploying two models in parallel and routing traffic between them dynamically using multi-armed bandits (Epsilon-greedy and Thompson sampling in particular).
We will use the dataset to simulate a real-world scenario consisting of several steps:
1. Split the data set in half (15K samples in each set) and treat the first half as the only data observes so far
2. Split the first half of the data in proportion 10K:5K samples to use as train:test sets for a first simple model (Random Forest)
3. After training the first model, simulate a "live" environment on the first 5K of data in the second half of the dataset
4. Use the so far observed 20K samples to train a second model (XGBoost)
5. Deploy the second model alongside the first together with a multi-armed bandit and simulate a "live" environment on the last 10K of the unobserved data, routing requests between the two models
The following diagram illustrates the proposed simulation design:

## Data preparation
```
import numpy as np
from sklearn.model_selection import train_test_split
OBSERVED_DATA = 15000
TRAIN_1 = 10000
TEST_1 = 5000
REST_DATA = 15000
RUN_DATA = 5000
ROUTE_DATA = 10000
# get features and target
X = data.loc[:, data.columns!=target].values
y = data[target].values
# observed/unobserved split
X_obs, X_rest, y_obs, y_rest = train_test_split(X, y, random_state=1, test_size=REST_DATA)
# observed split into train1/test1
X_train1, X_test1, y_train1, y_test1 = train_test_split(X_obs, y_obs, random_state=1, test_size=TEST_1)
# unobserved split into run/route
X_run, X_route, y_run, y_route = train_test_split(X_rest, y_rest, random_state=1, test_size=ROUTE_DATA)
# observed+run split into train2/test2
X_rest = np.vstack((X_run, X_route))
y_rest = np.hstack((y_run, y_route))
X_train2 = np.vstack((X_train1, X_test1))
X_test2 = X_run
y_train2 = np.hstack((y_train1, y_test1))
y_test2 = y_run
```
## Model training
We will train both models at once, but defer evaluation of the second model until simulating the live environment.
```
from sklearn.ensemble import RandomForestClassifier
rf = RandomForestClassifier(random_state=1)
rf.fit(X_train1, y_train1)
```
Now let's see how good our first model is on the test1 set:
```
from sklearn.metrics import accuracy_score, precision_score, recall_score, \
f1_score, confusion_matrix, classification_report
y_preds1 = rf.predict(X_test1)
print(classification_report(y_test1, y_preds1,
target_names=['No default','Default']))
for score in [accuracy_score, precision_score, recall_score, f1_score, confusion_matrix]:
print(score.__name__ + ':\n', score(y_test1, y_preds1))
%matplotlib inline
from utils import plot_confusion_matrix
cm = confusion_matrix(y_test1, y_preds1)
plot_confusion_matrix(cm, classes=['No default','Default'], normalize=True)
```
So a simple random forest model without any optimizations is able to outperform random guessing on accuracy and achieves a baseline F1 score of ~0.44. However, it is a poor predictor of default as it only achieves a recall of ~0.34.
Train the second model in advance, but defer evaluation:
```
from xgboost import XGBClassifier
xgb = XGBClassifier(random_state=1)
xgb.fit(X_train2, y_train2)
```
Save trained models to disk:
```
from sklearn.externals import joblib
joblib.dump(rf, 'models/rf_model/RFModel.sav')
joblib.dump(xgb, 'models/xgb_model/XGBModel.sav')
```
## Set up Kubernetes for live simulation
Pick Kubernetes cluster on GCP or Minikube.
```
minikube = False
if minikube:
!minikube start --vm-driver kvm2 --memory 4096 --cpus 6
else:
!gcloud container clusters get-credentials standard-cluster-1 --zone europe-west1-b --project seldon-demos
```
Create a cluster-wide cluster-admin role assigned to a service account named “default” in the namespace “kube-system”.
```
!kubectl create clusterrolebinding kube-system-cluster-admin --clusterrole=cluster-admin \
--serviceaccount=kube-system:default
!kubectl create namespace seldon
```
Add current context details to the configuration file in the seldon namespace.
```
!kubectl config set-context $(kubectl config current-context) --namespace=seldon
```
Create tiller service account and give it a cluster-wide cluster-admin role.
```
!kubectl -n kube-system create sa tiller
!kubectl create clusterrolebinding tiller --clusterrole cluster-admin --serviceaccount=kube-system:tiller
!helm init --service-account tiller
```
Check deployment rollout status and deploy seldon/spartakus helm charts.
```
!kubectl rollout status deploy/tiller-deploy -n kube-system
!helm install ../../../helm-charts/seldon-core-operator --name seldon-core --set usageMetrics.enabled=true --namespace seldon-system
!kubectl rollout status deploy/seldon-controller-manager -n seldon-system
%%bash
helm repo add datawire https://www.getambassador.io
helm repo update
helm install ambassador datawire/ambassador \
--set image.repository=quay.io/datawire/ambassador \
--set enableAES=false \
--set crds.keep=false
!kubectl rollout status deployment.apps/ambassador
```
Install analytics (Prometheus for metrics and Grafana for visualisation):
```
!helm install ../../../helm-charts/seldon-core-analytics --name seldon-core-analytics \
--set grafana_prom_admin_password=password \
--set persistence.enabled=false \
--namespace seldon
```
Port forward Ambassador (run command in terminal):
```sh
kubectl port-forward $(kubectl get pods -n seldon -l app.kubernetes.io/name=ambassador -o jsonpath='{.items[0].metadata.name}') -n seldon 8003:8080
```
Port forward Grafana (run command in terminal):
```sh
kubectl port-forward $(kubectl get pods -n seldon -l app=grafana-prom-server -o jsonpath='{.items[0].metadata.name}') -n seldon 3000:3000
```
You can then view an analytics dashboard inside the cluster at http://localhost:3000/d/rs_zGKYiz/mab?refresh=1s&orgId=1&from=now-2m&to=now. Login with:
Username : admin
password : password (as set when starting seldon-core-analytics above)
**Import the mab dashboard from ```assets/mab.json```.**
### Wrap model and router images with s2i
We have prepared the model classes under ```models/rf_model/RFModel.py``` and ```models/xgb_model/XGBModel.py``` for wrapping the trained models as docker images using s2i. The structure of the files is as follows:
```
!pygmentize models/rf_model/RFModel.py
```
Note that we define our own custom metrics which are the entries of the confusion matrix that will be exposed to Prometheus and visualized in Grafana as the model runs in the simulated live environment.
If Minikube used: create docker image for the trained models and routers inside Minikube using s2i.
```
if minikube:
!eval $(minikube docker-env) && \
make -C models/rf_model build && \
make -C models/xgb_model build && \
make -C ../epsilon-greedy build && \
make -C ../thompson-sampling build
```
## Deploy the first model
```
!kubectl apply -f assets/rf_deployment.json -n seldon
!kubectl rollout status deploy/rf-model-rf-model-7c4643e
```
## Simulate the first model in production for 5000 samples
```
from utils import rest_request_ambassador, send_feedback_rest
for i in range(X_run.shape[0]):
if i%1000 == 0:
print(f'Processed {i}/{X_run.shape[0]} samples', flush=True)
# fetch sample and make a request payload
x = X_run[i].reshape(1,-1).tolist()
request = {'data':{'ndarray':x}}
# send request to model
response = rest_request_ambassador('rf-deployment', 'seldon', request)
# extract prediction
probs = response.get('data').get('ndarray')[0]
pred = np.argmax(probs)
# send feedback to the model informing it if it made the right decision
truth_val = int(y_run[i])
reward = int(pred==truth_val)
truth = [truth_val]
_ = send_feedback_rest('rf-deployment', 'seldon', request, response, reward, truth)
```
We can see the model performance on the Grafana dashboard:
http://localhost:3000/d/rs_zGKYiz/mab?refresh=1s&orgId=1&from=now-2m&to=now (refresh to update)
## Deploy the original model and the new model with a router in front
Suppose now we have come up with a new model and want to deploy it alongside the first model with a multi-armed bandit router to make decisions which model should make predictions. We will delete the original deployment and make a new one that has both models in parallel and a router/multi-armed bandit in front.
To make things interesting, we will actually deploy 2 parallel deployments with the same 2 models but a different router in front (Epsilon-greedy and Thompson sampling) to compare the performance of two very different multi-armed bandit algorithms. One can think of the first deployment as a production deployment and the second parallel one as a shadow deployment whose responses are used for testing only.
But first, let's see what the performance of the new XGBoost model is on its test2 data:
```
y_preds2 = xgb.predict(X_test2)
print(classification_report(y_test2, y_preds2,
target_names=['No default','Default']))
for score in [accuracy_score, precision_score, recall_score, f1_score, confusion_matrix]:
print(score.__name__ + ':\n', score(y_test2, y_preds2))
cm = confusion_matrix(y_test2, y_preds2)
plot_confusion_matrix(cm, classes=['No default','Default'], normalize=True)
```
So the XGBoost model is slightly better than the old RFModel, so we expect any decent multi-armed bandit router to pick this up on live data, let's try this out.
First, delete the existing deployment of the old RFModel:
```
!kubectl delete sdep rf-deployment
```
Deploy the following two deployments:
```
from utils import get_graph
get_graph('assets/eg_deployment.json')
get_graph('assets/ts_deployment.json')
!kubectl apply -f assets/eg_deployment.json -n seldon
!kubectl apply -f assets/ts_deployment.json -n seldon
!kubectl rollout status deploy/poc-eg-eg-2-47fb8da
!kubectl rollout status deploy/poc-ts-ts-2-75f9d39
```
## Simulate both deployments in parellel with the remaining 10000 data samples
Here we send request and feedback to both parallel deployments, thus assessing the performance of the Epsilon-greedy router versus Thompson sampling as a method of routing to the best performing model.
```
for i in range(X_route.shape[0]):
if i%1000 == 0:
print(f'Processed {i}/{X_route.shape[0]} samples', flush=True)
# fetch sample and make a request payload
x = X_route[i].reshape(1,-1).tolist()
request = {'data':{'ndarray':x}}
# send request to both deployments
eg_response = rest_request_ambassador('eg-deployment', 'seldon', request)
ts_response = rest_request_ambassador('ts-deployment', 'seldon', request)
# extract predictions
eg_probs = eg_response.get('data').get('ndarray')[0]
ts_probs = ts_response.get('data').get('ndarray')[0]
eg_pred = np.argmax(eg_probs)
ts_pred = np.argmax(ts_probs)
# send feedback to the model informing it if it made the right decision
truth_val = int(y_route[i])
eg_reward = int(eg_pred==truth_val)
ts_reward = int(ts_pred==truth_val)
truth = [truth_val]
_ = send_feedback_rest('eg-deployment', 'seldon', request, eg_response, eg_reward, truth)
_ = send_feedback_rest('ts-deployment', 'seldon', request, ts_response, ts_reward, truth)
```
We can see the model performance on the Grafana dashboard:
http://localhost:3000/dashboard/db/mab?refresh=5s&orgId=1 (refresh to update)
We note that both the Epsilon greedy and Thompson sampling allocate more traffic to the better performing model (XGBoost) over time, but Thompson Sampling does so at a quicker rate as evidenced by the superior metrics (F1 score in particular).
## Clean-up
```
# delete data
!rm default-of-credit-card-clients-dataset.zip
!rm UCI_Credit_Card.csv
# delete trained models
!rm models/rf_model/RFModel.sav
!rm models/xgb_model/XGBModel.sav
# delete Seldon deployment from the cluster
!kubectl delete sdep --all
```
| github_jupyter |
<img src="NotebookAddons/blackboard-banner.png" width="100%" />
<font face="Calibri">
<br>
<font size="7"> <b> GEOS 657: Microwave Remote Sensing<b> </font>
<font size="5"> <b>Lab 9: InSAR Time Series Analysis using GIAnT within Jupyter Notebooks<br>Part 2: GIAnT <font color='rgba(200,0,0,0.2)'> -- [## Points] </font> </b> </font>
<br>
<font size="4"> <b> Franz J Meyer & Joshua J C Knicely; University of Alaska Fairbanks</b> <br>
<img src="NotebookAddons/UAFLogo_A_647.png" width="170" align="right" /><font color='rgba(200,0,0,0.2)'> <b>Due Date: </b>NONE</font>
</font>
<font size="3"> This Lab is part of the UAF course <a href="https://radar.community.uaf.edu/" target="_blank">GEOS 657: Microwave Remote Sensing</a>. This lab is divided into 3 parts: 1) data download and preprocessing, 2) GIAnT time series, and 3) a simple Mogi source inversion. The primary goal of this lab is to demonstrate how to process InSAR data, specifically interferograms, using the Generic InSAR Analysis Toolbox (<a href="http://earthdef.caltech.edu/projects/giant/wiki" target="_blank">GIAnT</a>) in the framework of *Jupyter Notebooks*.<br>
<b>Our specific objectives for this lab are to:</b>
- Prepare data for GIAnT:
- Create necessary ancillary files and functions.
- Reformat the interferograms.
- Run GIAnT
- Use GIAnT to create maps of surface deformation:
- Understand its capabilities.
- Understand its limitations.
</font>
<br>
<font face="Calibri">
<font size="5"> <b> Target Description </b> </font>
<font size="4"> <font color='rgba(200,0,0,0.2)'> <b>THIS NOTEBOOK INCLUDES NO HOMEWORK ASSIGNMENTS.</b></font> <br>
Contact me at fjmeyer@alaska.edu should you run into any problems.
</font>
```
%%javascript
var kernel = Jupyter.notebook.kernel;
var command = ["notebookUrl = ",
"'", window.location, "'" ].join('')
kernel.execute(command)
from IPython.display import Markdown
from IPython.display import display
user = !echo $JUPYTERHUB_USER
env = !echo $CONDA_PREFIX
if env[0] == '':
env[0] = 'Python 3 (base)'
if env[0] != '/home/jovyan/.local/envs/insar_analysis':
display(Markdown(f'<text style=color:red><strong>WARNING:</strong></text>'))
display(Markdown(f'<text style=color:red>This notebook should be run using the "insar_analysis" conda environment.</text>'))
display(Markdown(f'<text style=color:red>It is currently using the "{env[0].split("/")[-1]}" environment.</text>'))
display(Markdown(f'<text style=color:red>Select "insar_analysis" from the "Change Kernel" submenu of the "Kernel" menu.</text>'))
display(Markdown(f'<text style=color:red>If the "insar_analysis" environment is not present, use <a href="{notebookUrl.split("/user")[0]}/user/{user[0]}/notebooks/conda_environments/Create_OSL_Conda_Environments.ipynb"> Create_OSL_Conda_Environments.ipynb </a> to create it.</text>'))
display(Markdown(f'<text style=color:red>Note that you must restart your server after creating a new environment before it is usable by notebooks.</text>'))
```
<font face='Calibri'><font size='5'><b>Overview</b></font>
<br>
<font size='3'><b>About GIAnT</b>
<br>
GIAnT is a Python framework that allows rapid time series analysis of low amplitude deformation signals. It allows users to use multiple time series analysis technqiues: Small Baseline Subset (SBAS), New Small Baseline Subset (N-SBAS), and Multiscale InSAR Time-Series (MInTS). As a part of this, it includes the ability to correct for atmospheric delays by assuming a spatially uniform stratified atmosphere.
<br><br>
<b>Limitations</b>
<br>
GIAnT has a number of limitations that are important to keep in mind as these can affect its effectiveness for certain applications. It implements the simplest time-series inversion methods. Its single coherence threshold is very conservative in terms of pixel selection. It does not include any consistency checks for unwrapping errors. It has a limited dictionary of temporal model functions. It cannot correct for atmospheric effects due to differing surface elevations.
<br><br>
<b>Steps to use GIAnT</b><br>
Although GIAnT is an incredibly powerful tool, it requires very specific input. Because of the input requirements, the bulk of the lab and code below is dedicated to getting our data into a form that GIAnT can manipulate and to creating files that tell GIAnT what to do. The general steps to use GIAnT are below.
<br><br>
More information about GIAnT can be found here, <a href="http://earthdef.caltech.edu/projects/giant/wiki" target="_blank">http://earthdef.caltech.edu/projects/giant/wiki</a>.
<br><br>
- Download Data
- Identify Area of Interest
- Subset (Crop) Data to Area of Interest
- Prepare Data for GIAnT (Today's lab starts here)
- Adjust file names
- Remove potentially disruptive default values (optional)
- Convert data from '.tiff' to '.flt' format
- Create Input Files for GIAnT
- Create 'ifg.list'
- Create 'date.mli.par'
- Make prepxml_SBAS.py
- Run prepxml_SBAS.py
- Make userfn.py
- Run GIAnT
- PrepIgramStack.py
- ProcessStack.py
- SBASInvert.py
- SBASxval.py
- Data Visualization
<br><br>
When you use GIAnT, cite the creator's work using:<br>
Agram et al., (2013). "New Radar Interferometric Time Series Analysis Toolbox Released." Eos Trans. AGU, 94, 69.
<br>AND<br>
Agram et al., (2012). "Generic InSAR Analysis Toolbox (GIAnT) - User Guide." <http://earthdef.caltech.edu>
<br><br><b><i>DO WE NEED TO ALSO GIVE ASF A CITATION???</i></b>
<font face='Calibri'><font size='5'><b>Required Files</b></font>
<br>
<font size='3'>Before we begin, each student should have a specific subset of files with a specific folder structure as described below. If your folder structure differs, the code will need to be modified.
<br>
- Folder: Corrected Subsets
- Files: TRAIN corrected unwrapped phase subsets
- Files: Uncorrected unwrapped phase subsets
- Folder: Ingram Subsets
- Files: Amplitude subsets
- Files: Coherence subsets
- Files: Unwrapped phase (ingram) subsets
- Theoretically, these are identical to the uncorrected unwrapped phase subsets in the Corrected Subsets folder.
</font></font>
<font face='Calibri'><font size='5'><b>0 System Setup</b></font><br>
<font size='3'>We will first do some system setup. This involves importing requiesite Python libraries and defining all of our user inputs. </font></font>
<font face='Calibri'><font size='4'><b>0.0 Import Python Packages</b></font><br>
<font size='3'>Let's import the Python libraries we will need to run this lab. </font></font>
```
%%capture
import os
import h5py
import shutil
import re
from datetime import date
import glob
import pickle
import pathlib
import subprocess
from math import ceil
from osgeo import gdal
from osgeo import osr
import pandas as pd
%matplotlib notebook
import matplotlib.pyplot as plt
import matplotlib.animation
import matplotlib.patches as patches # for Rectangle
plt.rcParams.update({'font.size': 10})
import numpy as np
from numpy import isneginf
from IPython.display import HTML
from matplotlib import animation, rc
import asf_notebook as asfn
asfn.jupytertheme_matplotlib_format()
```
<font face='Calibri' size='4'><b>0.1 Enter an analysis directory prepared in the Part 1 notebook</b></font>
```
pickle_pth = "part1_pickle"
while True:
analysis_directory = input("Enter the absolute path to the directory holding your data from Part 1:\n")
if os.path.exists(f"{analysis_directory}/{pickle_pth}"):
break
else:
print(f"\n{analysis_directory} does not contain {pickle_pth}")
continue
os.chdir(analysis_directory)
print(f"\nanalysis_directory: {analysis_directory}")
```
<font face='Calibri' size='4'><b>0.2 Open our pickle from the Part 1 notebook</b></font>
```
pickle_pth = "part1_pickle"
if os.path.exists(pickle_pth):
with open(pickle_pth,'rb') as infile:
part1_pickle = pickle.load(infile)
print(f"part1_pickle = {part1_pickle}")
else:
print('Invalid Path. Did you follow part 1 and generate your pickle?')
```
<font face='Calibri' size='4'><b>0.3 Define some important paths</b></font>
```
ingram_folder = part1_pickle['ingram_folder'] # Location of the original ingram folders
corrected_folder = part1_pickle['corrected_folder'] # Location of the TRAIN corrected subsets
subset_folder = part1_pickle['subset_folder'] # Location of the original (uncorrected) subsets
giant_dir = f'{analysis_directory}/GIAnT' # directory where we will perform GIAnT analysis
giant_data = f'{analysis_directory}/GIAnT_Data' # directory where we will store our data for use by GIAnT
output_dir = f'{analysis_directory}/geotiffs_and_animations'
pathlib.Path(output_dir).mkdir(parents=True, exist_ok=True)
```
<font face='Calibri'>
<font size='4'><b>0.4 Decide whether or not to delete old data</b></font>
<br>
<font size='3'>True = delete
<br>
False = save</font>
```
replace_giant_dir = True # True or False; if True, any folder with the same name will be deleted and recreated.
replace_giant_data = True
replace_output_dir = True
# set things to find files
unw = '_unw_phase.'
unw_corr = '_unw_phase_corrected.'
corr = '_corr.'
amp = '_amp.'
```
<font face='Calibri' size='4'><b>0.5 Delete the GIAnT data and working directories if 'replace_* = True'</b></font>
```
if replace_giant_dir:
try:
shutil.rmtree(giant_dir)
except:
pass
if replace_giant_data:
try:
shutil.rmtree(giant_data)
except:
pass
if replace_output_dir:
try:
shutil.rmtree(output_dir)
except:
pass
# If the GIAnT data and working directories don't exist, create them.
pathlib.Path(giant_dir).mkdir(parents=True, exist_ok=True)
pathlib.Path(giant_data).mkdir(parents=True, exist_ok=True)
pathlib.Path(output_dir).mkdir(parents=True, exist_ok=True)
```
<font face='Calibri' size='4'><b>0.7 Load the pickled heading_avg</b></font>
```
heading_avg = part1_pickle['heading_avg']
print(f"heading_avg = {heading_avg}")
```
<font face='Calibri' size='4'><b>0.8 Copy a single clipped geotiff into our working GIAnT directory for later data visualization</b></font>
```
amps = [f for f in os.listdir(subset_folder) if '_amp' in f]
amps.sort()
shutil.copy(os.path.join(subset_folder, amps[0]), giant_dir)
radar_tiff = os.path.join(giant_dir, amps[0])
```
<font face='Calibri'><font size='5'><b>1. Create Input Files And Code for GIAnT</b></font>
<br>
<font size ='3'>Let's create the input files and specialty code that GIAnT requires. These are listed below.
<br>
- ifg.list
- List of the interferogram properties including primary and secondary dates, perpendicular baseline, and sensor name.
- date.mli.par
- File from which GIAnT pulls requisite information about the sensor.
- This is specifically for GAMMA files. When using other interferogram processing techniques, an alternate file is required.
- prepxml_SBAS.py
- Python function to create an xml file that specifies the processing options to GIAnT.
- This must be modified by the user for their particular application.
- userfn.py
- Python function to map the interferogram dates to a phyiscal file on disk.
- This must be modified by the user for their particular application.
</font>
</font>
<font face='Calibri' size='4'><b>1.1 Create 'ifg.list' File </b></font>
<br><br>
<font face='Calibri' size='3'><b>1.1.0 Create a list of unwrapped, phase-corrected data</b></font>
```
files = [f for f in os.listdir(corrected_folder) if f.endswith(f'{unw_corr}tif')] # Just get one of each file name.
files.sort()
print(len(files))
print(*files, sep="\n")
```
<font face='Calibri' size='3'><b>1.1.1 Create a 4 column text file to communicate network information to GIAnT within the GIAnT folder</b>
<br>
Contains primary date, secondary date, perpendicular baseline, and sensor designation
</font>
```
reference_dates = []
secondary_dates = []
for file in files:
reference_dates.append(file[0:8])
secondary_dates.append(file[9:17])
# Sort the dates according to the primary dates.
p_dates, s_dates = (list(t) for t in zip(*sorted(zip(reference_dates, secondary_dates))))
with open(os.path.join(giant_dir, 'ifg.list'), 'w') as fid:
for i, dt in enumerate(p_dates):
primary_date = dt # pull out Primary Date (first set of numbers)
secondary_date = s_dates[i] # pull out Secondary Date (second set of numbers)
bperp = '0.0' # perpendicular baseline.
sensor = 'S1' # Sensor designation.
# write values to the 'ifg.list' file.
fid.write(f'{primary_date} {secondary_date} {bperp} {sensor}\n')
```
<font face='Calibri' size='3'><b>1.1.2 Print ifg.list</b></font>
```
with open(os.path.join(giant_dir, 'ifg.list'), 'r') as fid:
print(fid.read())
```
<font face='Calibri'><font size='3'>You may notice that the code above sets the perpendicular baseline to a value of 0.0 m. This is not the true perpendicular baseline. That value can be found in metadata file (titled <font face='Courier New'>$<$primary timestamp$>$_$<$secondary timestamp$>$.txt</font>) that comes with the original interferogram. Generally, we would want the true baseline for each interferogram. However, since Sentinel-1 has such a short baseline, a value of 0.0 m is sufficient for our purposes. </font></font>
<font face='Calibri' size='4'> <b>1.2 Create 'date.mli.par' File </b></font>
<br>
<font face='Calibri' font size='3'> As we are using GAMMA products, we must create a 'date.mli.par' file from which GIAnT will pull necessary information. If another processing technique is used to create the interferograms, an alternate file name and file inputs are required.
<br><br>
<b>1.2.0 Make a list of paths to all of the unwrapped interferograms</b>
</font>
```
files = [f for f in os.listdir(corrected_folder) if f.endswith(f'{unw_corr}tif')]
print(len(files))
print(*files, sep="\n")
print(f'\n\nCurrent Working Directory: {os.getcwd()}')
print(f'radar_tiff path: {radar_tiff}')
```
<font face='Calibri' font size='3'><b>1.2.1 Gather metadata required by GIAnT and save it to a date.mli.par file</b></font>
```
def get_primary_insar_aquisition_date(ingram_name):
regex = '[0-9]{8}T[0-9]{6}'
match = re.search(regex, ingram_name)
if match:
return match[0]
ds = gdal.Open(radar_tiff, gdal.GA_ReadOnly)
# Get WIDTH (xsize; AKA 'Pixels') and FILE_LENGTH (ysize; AKA 'Lines') information
n_lines = ds.RasterYSize
n_pixels = ds.RasterXSize
ds = None
# Get the center line UTC time stamp; can also be found inside <date>_<date>.txt file
ingram_name = os.listdir(ingram_folder)[0]
tstamp = get_primary_insar_aquisition_date(ingram_name)
c_l_utc = int(tstamp[0:2])*3600 + int(tstamp[2:4])*60 + int(tstamp[4:6])
# radar frequency; speed of light divided by radar wavelength of Sentinel-1 in meters
rfreq = 299792548.0 / 0.055465763
# write the 'date.mli.par' file
with open(os.path.join(giant_dir, 'date.mli.par'), 'w') as fid:
# when using GAMMA products, GIAnT requires the radar frequency. Everything else is in wavelength (m)
fid.write(f'radar_frequency: {rfreq} \n')
# Method from Tom Logan's prepGIAnT code; can also be found inside <date>_<date>.txt file
fid.write(f'center_time: {c_l_utc} \n')
# inside <date>_<date>.txt file; can be hardcoded or set up so code finds it.
fid.write(f'heading: {heading_avg} \n')
# number of lines in direction of the satellite's flight path
fid.write(f'azimuth_lines: {n_lines} \n')
# number of pixels in direction perpendicular to satellite's flight path
fid.write(f'range_samples: {n_pixels} \n')
```
<font face='Calibri' font size='3'><b>1.2.2 Print the metadata stored in 'date.mli.par'</b></font>
```
with open(os.path.join(giant_dir, 'date.mli.par'), 'r') as fid:
print(fid.read())
```
<font face='Calibri'><font size='4'><b>1.3 Create prepxml_SBAS.py Function</b> </font>
<br>
<font size='3'>We will create a prepxml_SBAS.py function and put it into our GIAnT working directory.</font>
</font>
<font face='Calibri'> <font size='3'><b>Necessary prepxml_SBAS.py edits</b></font>
<br>
<font size='3'> GIAnT comes with an example prepxml_SBAS.py, but requries significant edits for our purposes. These alterations have already been made, so we don't have to do anything now, but it is good to know the kinds of things that have to be altered. The details of some of these options can be found in the GIAnT documentation. The rest must be found in the GIAnT processing files themselves, most notably the tsxml.py and tsio.py functions. <br>The following alterations were made:
<br>
- Changed 'example' ► 'date.mli.par'
- Removed 'xlim', 'ylim', 'rxlim', and 'rylim'
- These are used for clipping the files in GIAnT. As we have already done this, it is not necessary.
- Removed latfile='lat.map' and lonfile='lon.map'
- These are optional inputs for the latitude and longitude maps.
- Removed hgtfile='hgt.map'
- This is an optional altitude file for the sensor.
- Removed inc=21.
- This is the optional incidence angle information.
- It can be a constant float value or incidence angle file.
- For Sentinel1, it varies from 29.1-46.0°.
- Removed masktype='f4'
- This is the mask designation.
- We are not using any masks for this.
- Changed unwfmt='RMG' ► unwfmt='GRD'
- Read data using GDAL.
- Removed demfmt='RMG'
- Changed corfmt='RMG' ► corfmt='GRD'
- Read data using GDAL.
- Changed nvalid=30 -> nvalid=1
- This is the minimum number of interferograms in which a pixel must be coherent. A particular pixel will be included only if its coherence is above the coherence threshold, cohth, in more than nvalid number of interferograms.
- Removed atmos='ECMWF'
- This is an amtospheric correction command. It depends on a library called 'pyaps' developed for GIAnT. This library has not been installed yet.
- Changed masterdate='19920604' ► masterdate='20161119'
- Use our actual masterdate.
- I simply selected the earliest date as the masterdate.</font>
<br><br>
<font size='3'><b>1.3.0 Define some prepxml inputs</b>
<br>
The reference region and related variables</font></font></font>
```
use_ref_region = False
filt = 1.0 / 12 # Start with a temporal filter in length of years
ref_region_size = [5, 5] # Reference region size in Lines and Pixels
ref_region_center = [-0.9, -91.3] # Center of reference region in lat, lon coordinates
rxlim = [0, 10]
rylim = [95, 105]
primary_date = min([amps[i][0:8] for i in range(len(amps))], key=int) # set the primary date.
print(f"Primary Date: {primary_date}")
```
<font face='Calibri'><font size='3'><b>1.3.1 Create the prepxml Python script</b></font>
```
if use_ref_region == True:
use_ref = ''
else:
use_ref = '#'
prepxml_sbas_template = '''
#!/usr/bin/env python
"""Example script for creating XML files for use with the SBAS processing chain. This script is supposed to be copied to the working directory and modified as needed."""
import sys
tsinsar_pth = '/home/jovyan/.local/GIAnT'
if tsinsar_pth not in sys.path:
sys.path.append(tsinsar_pth)
import tsinsar as ts
import argparse
import numpy as np
def parse():
parser= argparse.ArgumentParser(description='Preparation of XML files for setting up the processing chain. Check tsinsar/tsxml.py for details on the parameters.')
parser.parse_args()
parse()
g = ts.TSXML('data')
g.prepare_data_xml(
'{0}/date.mli.par', proc='GAMMA',
{7}rxlim = [{1},{2}], rylim=[{3},{4}],
inc = 21., cohth=0.10,
unwfmt='GRD', corfmt='GRD', chgendian='True', endianlist=['UNW','COR'])
g.writexml('data.xml')
g = ts.TSXML('params')
g.prepare_sbas_xml(nvalid=1, netramp=True,
#atmos='NARR',
demerr=False, uwcheck=False, regu=True, masterdate='{5}', filt={6})
g.writexml('sbas.xml')
############################################################
# Program is part of GIAnT v1.0 #
# Copyright 2012, by the California Institute of Technology#
# Contact: earthdef@gps.caltech.edu #
############################################################
'''
with open(os.path.join(giant_dir, 'prepxml_SBAS.py'), 'w') as fid:
fid.write(prepxml_sbas_template.format(giant_dir,
rxlim[0], rxlim[1],
rylim[0], rylim[1],
primary_date, filt,use_ref))
```
<font face='Calibri'><font size='3'><b>1.3.2 Print the prepxml script</b></font>
```
with open(os.path.join(giant_dir, 'prepxml_SBAS.py'), 'r') as fid:
print(fid.read())
```
<font face='Calibri'><font size='4'><b>1.4 Create userfn.py script</b></font>
<br>
<font size='3'>This file maps the interferogram dates to a physical file on disk. This python file must be in our working directory, <b>/GIAnT</b>.
<br><br>
<b>1.4.0 Write userfn.py</b>
</font>
```
userfn_template = """
#!/usr/bin/env python
import os
def makefnames(dates1, dates2, sensor):
dirname = '{0}'
root = os.path.join(dirname, dates1+'_'+dates2)
unwname = root+'{1}'+'flt' # for potentially disruptive default values removed.
corname = root+'_corr.flt'
return unwname, corname
"""
with open(os.path.join(giant_dir, 'userfn.py'), 'w') as fid:
fid.write(userfn_template.format(giant_data, unw_corr))
```
<font face='Calibri'><font size='3'><b>1.4.1 Print userfn.py</b></font>
```
with open(os.path.join(giant_dir, 'userfn.py'), 'r') as fid:
print(fid.read())
```
<font face='Calibri'>
<font size='5'> <b> 2. Prepare Data for GIAnT </b> </font>
<br>
<font size='3'> Our resultant files need to be adjusted for input into GIAnT. This involves:
<ul>
<li>Adjusting filenames to start with the date</li>
<li>(Optional) Removing potentially disruptive default values</li>
<li>Converting data format from '.tif' to '.flt'</li>
</ul>
</font>
</font>
<font face='Calibri'>
<font size='4'><b>2.1 Adjust File Names</b></font>
<font size='3'><br>GIAnT requires the files to have a simple format of <font face='Courier New'><primary_date>-<secondary_date>_<unwrapped or coherence designation>.tiff</font>.
<br><br>
<font color='green'><b>We already did this step in Part 1, so section 2.1 is just a reminder that it is also a requirement for GIAnT.</b></font>
</font> </font>
<font face='Calibri'> <font size='4'><b>2.2 Remove disruptive default values</b></font>
<br>
<font size='3'>Now we can remove the potentially disruptive default values from the unwrapped interferograms. For this lab, this step is optional. This is because the Sentinel-1 data does not have any of the disruptive default values. However, for other datasources, this may be required.<br>This works by creating an entirely new geotiff with the text <font face='Courier New'>'\_no\_default'</font> added to the file name.
<br><br>
<b>2.2.0 Write functions to gather and print paths via a wildcard path.</b>
</font>
```
def get_tiff_paths(paths):
tiff_paths = glob.glob(paths)
tiff_paths.sort()
return tiff_paths
def print_tiff_paths(tiff_paths):
for p in tiff_paths:
print(f"{p}\n")
```
<font face='Calibri'><font size='3'><b>2.2.1 Gather the paths to the uncorrected and corrected unw_phase tifs</b></font>
```
paths_unc = f"{corrected_folder}/*{unw}tif"
paths_corr = f"{corrected_folder}/*{unw_corr}tif"
files_unc = get_tiff_paths(paths_unc)
files_corr = get_tiff_paths(paths_corr)
print_tiff_paths(files_unc)
print_tiff_paths(files_corr)
```
<font face='Calibri'><font size='3'><b>2.2.2 Decide if you wish to replace dispruptive NaN values with zeros</b>
<br>
True = replace NaNs
<br>
False = keep NaNs
</font>
```
replace_nans = False
```
<font face='Calibri'><font size='3'><b>2.2.3 Write a function to remove disruptive (NaN) values</b></font>
```
def replace_nan_values(paths):
for file in paths:
outfile = os.path.join(analysis_directory, file.replace('.tif','_no_default.tif'))
cmd = f"gdal_calc.py --calc=\"(A>-1000)*A\" --outfile={outfile} -A {analysis_directory}/{file} --NoDataValue=0 --format=GTIFF"
subprocess.call(cmd, shell=True)
```
<font face='Calibri'> <font size='3'><b>2.2.4 Remove files that have potentially disruptive default values.</b> This step will run only if there is a file with '\_no\_default' in the <font face='Courier New' size='2'>corrected_folder</font> directory.</font> </font>
```
if replace_nans:
replace_nan_values(files_unc)
replace_nan_values(files_corr)
# change 'unw' and 'unw_corr' so the appropriate files will still be found in later code.
unw = unw.replace('_unw_phase', '_unw_phase_no_default')
unw_corr = unw_corr.replace('_corrected', '_corrected_no_default')
else:
print("Skipping the removal of disruptive values.")
```
<font face='Calibri'>
<font size='4'><b>2.3 Convert data format from '.tif' to '.flt'</b><br></font>
<font size='3'>This is a requirement of GIAnT
<br><br>
<b>2.3.0 Gather the paths to the files to convert</b></font></font>
```
# Find the unwrapped phase files to convert.
paths_unc = f"{corrected_folder}/*{unw}tif"
paths_corr = f"{corrected_folder}/*{unw_corr}tif"
paths_cohr = f"{subset_folder}/*{corr}tif"
files_unc = get_tiff_paths(paths_unc)
files_corr = get_tiff_paths(paths_corr)
files_cohr = get_tiff_paths(paths_cohr)
# file format should appear as below
# YYYYMMDD-YYYYMMDD_unw_phase.flt
# OR, if default values have been removed...
# YYYYMMDD-YYYYMMDD_unw_phase_no_default.flt
print_tiff_paths(files_unc)
print_tiff_paths(files_corr)
print_tiff_paths(files_cohr)
```
<font face='Calibri'><font size='3'><b>2.3.1 Convert the .tif files to .flt files</b>
<br>
tiff_to_flt creates 3 files: '*.flt', '*.flt.aux.xml', and '*.hdr', where the asterisk, '*', is replaced with the file name</font>
```
def tiff_to_flt(paths):
for file in paths:
newname, old_ext = os.path.splitext(file)
# Create system command that will convert the file from a .tiff to a .flt
cmd = f"gdal_translate -of ENVI {file} {file.replace('.tif','.flt')}"
try:
#print(f"cmd = {cmd}") # display what the command looks like.
# pass command to the system
subprocess.call(cmd, shell=True)
except:
print(f"Problem with command: {cmd}")
print("Conversion from '.tif' to '.flt' complete.")
tiff_to_flt(files_unc)
tiff_to_flt(files_corr)
tiff_to_flt(files_cohr)
```
<font face = 'Calibri' size='3'><b>2.3.1 Gather the paths to the '\*.flt', '\*.hdr', and '\*.aux.xml' files</b></font>
```
paths_flt = f"*/*.flt"
paths_hdr = f"*/*.hdr"
paths_aux = f"*/*.flt.aux.xml"
files_flt = get_tiff_paths(paths_flt)
files_hdr = get_tiff_paths(paths_hdr)
files_aux = get_tiff_paths(paths_aux)
print_tiff_paths(files_flt)
print_tiff_paths(files_hdr)
print_tiff_paths(files_aux)
```
<font face='Calibri'><font size='3'><b>2.3.2 Move the files and delete the extraneous '.flt.aux.xml' files</b></font>
```
def move_list_of_files(paths, new_analysis_directory):
for file in paths:
path, name = os.path.split(file)
try:
shutil.move(file,os.path.join(new_analysis_directory, name))
except FileNotFoundError as e:
raise type(e)(f"Failed to find {file}")
move_list_of_files(files_flt, giant_data)
move_list_of_files(files_hdr, giant_data)
for file in files_aux:
os.remove(file)
```
<font face='Calibri'> <font size='4'> <b>2.4 Run prepxml_SBAS.py </b> </font>
<br>
<font size='3'> Here we run <b>prepxml_SBAS.py</b> to create the 2 needed files
- data.xml
- sbas.xml
To use MinTS, we would create and run <b>prepxml_MinTS.py</b> to create
- data.xml
- mints.xml
These files are needed by <b>PrepIgramStack.py</b>.
</font>
<br><br>
<font size='3'><b>Define the path to the GIAnT module, change into giantDir, and run prepxml_SBAS.py</b>
<br>
Note: We have to change into giant_dir due to GIAnT's use of relative paths to find expected files.
<br><br>
<b>2.4.0 Define the path to the GIAnT installation and move into giant_dir (in your analysis directory)</b></font>
```
giant_path = "/home/jovyan/.local/GIAnT/SCR"
#install cython (replaces weave. Needed for running GIAnT in Python 3)
# !pip install --user cython
try:
os.chdir(giant_dir)
except FileNotFoundError as e:
raise type(e)(f"Failed to find {file}")
print(f"current directory: {os.getcwd()}\n")
for file in glob.glob('*.*'):
print(file)
```
<font face='Calibri' size='3'><b>2.4.1 Download GIAnT from the asf-jupyter-data S3 bucket</b>
<br>
GIAnT is no longer supported (Python 2). This unofficial version of GIAnT has been partially ported to Python 3 to run this notebook. Only the portions of GIAnT used in this notebook have been tested.
</font>
```
if not os.path.exists("/home/jovyan/.local/GIAnT"):
download_path = 's3://asf-jupyter-data/GIAnT_5_21.zip'
output_path = f"/home/jovyan/.local/{os.path.basename(download_path)}"
!aws --region=us-east-1 --no-sign-request s3 cp $download_path $output_path
if os.path.isfile(output_path):
!unzip $output_path -d /home/jovyan/.local/
os.remove(output_path)
```
<font face='Calibri'><font size='3'><b>2.4.2 Run prepxml_SBAS.py</b></font>
```
!python prepxml_SBAS.py
```
<font face='Calibri' size='3'><b>2.4.3 View the newly created data.xml</b></font>
```
with open('data.xml', 'r') as f:
print(f.read())
```
<font face='Calibri' size='3'><b>2.4.4 View the newly created 'sbas.xml'</b></font>
```
with open('sbas.xml', 'r') as f:
print(f.read())
```
<font face='Calibri'><font size='5'><b>3. Run GIAnT</b></font>
<br>
<font size='3'>We have now created all of the necessary files to run GIAnT. The full GIAnT process requires 3 function calls.
<ul>
<li>PrepIgramStack.py</li>
<li>ProcessStack.py</li>
<li>SBASInvert.py</li>
</ul>
We will make a 4th function call that is not necessary, but provides some error estimation that can be useful.
<ul>
<li>SBASxval.py</li>
</ul>
</font>
<font face='Calibri'> <font size='4'> <b>3.0 Run PrepIgramStack.py </b> </font>
<br>
<font size='3'> Here we run <b>PrepIgramStack.py</b> to create the files for GIAnT. This will read in the input data and the files we previously created. This will output HDF5 files. As we did not have to modify <b>PrepIgramStack.py</b>, we will call from the installed GIAnT library. <br>
Inputs:
- ifg.list
- data.xml
- sbas.xml
Outputs:
- RAW-STACK.h5
- PNG previews under 'Igrams' folder
</font> </font>
<br>
<font face='Calibri' size='3'><b>3.0.0 Display the help information for PrepIgramStack.py</b></font>
```
!python $giant_path/PrepIgramStack.py -h
```
<font face='Calibri' size='3'><b>3.0.1 Run PrepIgramStack.py</b>
<br>
Note: The processing status bar may not fully complete when the script is finished running. This is okay.</font>
```
!python $giant_path/PrepIgramStack.py -i ifg.list
# The '-i ifg.list' is technically unnecessary as that is the default.
```
<font face='Calibri' size='3'><b>3.0.2 Confirm the correct HDF5 file format</b></font>
```
file = os.path.join('Stack','RAW-STACK.h5')
if not h5py.is_hdf5(file):
print(f'Not an HDF5 file:{file}')
else:
print("It's an HDF5 file")
```
<font face='Calibri'> <font size='4'> <b>3.1 Run ProcessStack.py </b> </font>
<br>
<font size='3'> This seems to be an optional step. Does atmospheric corrections and estimation of orbit residuals. <br>
Inputs:
- HDF5 files from PrepIgramStack.py, RAW-STACK.h5
- data.xml
- sbas.xml
- GPS Data (optional; we don't have this)
- Weather models (downloaded automatically)
Outputs:
- HDF5 files, PROC-STACK.h5
These files are then fed into SBAS.
</font> </font>
<br><br>
<font face='Calibri' size='3'><b>3.1.0 Display the help information for ProcessStack.py</b></font>
```
!python $giant_path/ProcessStack.py -h
```
<font face='Calibri' size='3'><b>3.1.1 Run ProcessStack.py</b>
<br>
Note: The processing status bars may never fully complete. This is okay.</font>
```
!python $giant_path/ProcessStack.py
```
<font face='Calibri' size='3'><b>3.1.2 Confirm the correct HDF5 file format</b></font>
```
file = os.path.join('Stack','PROC-STACK.h5')
#print(files)
if not h5py.is_hdf5(file):
print(f'Not an HDF5 file:{file}')
else:
print("It's an HDF5 file!")
```
<font face='Calibri'> <font size='4'> <b>3.2 Run SBASInvert.py </b> </font>
<br>
<font size='3'> Run the time series.
Inputs
<ul>
<li>HDF5 file, PROC-STACK.h5</li>
<li>data.xml</li>
<li>sbas.xml</li>
</ul>
Outputs
<ul>
<li>HDF5 file: LS-PARAMS.h5</li>
</ul>
</font> </font>
<br>
<font face='Calibri' size='3'><b>3.2.0 Display the help information for SBASInvert.py</b></font>
```
!python $giant_path/SBASInvert.py -h
```
<font face='Calibri' size='3'><b>3.2.1 Run SBASInvert.py</b>
<br><br>
Note: The processing status bar may not fully complete when the script is finished running. This is okay.</font>
```
!python $giant_path/SBASInvert.py
```
<font face='Calibri'> <font size='4'> <b>3.3 Run SBASxval.py </b> </font>
<br>
<font size='3'> Get an uncertainty estimate for each pixel and epoch using a Jacknife test. This is often the most time consuming portion of the code.
Inputs:
<ul>
<li>HDF5 files, PROC-STACK.h5</li>
<li>data.xml</li>
<li>sbas.xml</li>
</ul>
Outputs:
<ul>
<li>HDF5 file, LS-xval.h5</li>
</ul>
</font> </font>
<br>
<font face='Calibri' size='3'><b>3.3.0 Display the help information for SBASxval.py</b></font>
```
!python $giant_path/SBASxval.py -h
```
<font face='Calibri' size='3'><b>3.3.1 Run SBASInvert.py</b>
<br><br>
Note: The processing status bar may not fully complete when the script is finished running. This is okay.</font>
```
!python $giant_path/SBASxval.py
```
<font face='Calibri' size='4'><b>3.4 Having completed our GIAnT processing, return to the analysis' analysis_directory directory</b></font>
```
os.chdir(analysis_directory)
print(f'Current directory: {os.getcwd()}')
```
<font face='Calibri'><font size='5'><b>4. Data Visualization</b></font>
<br><br>
<font size='4'><b>4.0 Prepare data for visualization</b></font>
<br>
<font size='3'><b>4.0.0 Load the LS-PARAMS stack produced by GIAnT and read it into an array so we can manipulate and display it.</b></font></font>
```
filename = os.path.join(giant_dir, 'Stack/LS-PARAMS.h5')
f = h5py.File(filename, 'r')
# List all groups
group_key = list(f.keys()) # get list of all keys.
print(f"group_key: {group_key}") # print list of keys.
```
<font face='Calibri' size='3'><b>4.0.1 Display LS-PARAMS info with gdalinfo</b></font>
```
!gdalinfo {filename}
```
<font face='Calibri'><font size='3'><b>4.0.2 Load the LS-xval stack produced by GIAnT and read it into an array so we can manipulate and display it.</b></font></font>
```
filename = os.path.join(giant_dir, 'Stack/LS-xval.h5')
error = h5py.File(filename, 'r')
# List all groups
error_key = list(error.keys()) # get list of all keys.
print(f"error_key: {error_key}") # print list of keys.
```
<font face='Calibri' size='3'><b>4.0.3 Display LS-xval info with gdalinfo</b></font>
```
print(gdal.Info(filename))
```
<font face='Calibri'><font size='3'>Details on what each of these keys in group_key and error_key means can be found in the GIAnT documentation. For now, the only keys with which we are concerned are 'recons' (the filtered time series of each pixel) and 'dates' (the dates of acquisition). It is important to note that the dates are given in a type of Julian Day number called Rata Die number which we convert to Gregorian dates below. </font></font>
<br><br>
<font face='Calibri'><font size='3'><b>4.0.4 Get the relevant data and error values from our stacks and confirm that the number of rasters in our data stack, the number of error points, and the number of dates are all the same.</b></font></font>
```
# Get our data from the stack
data_cube = f['recons'][()]
# Get our error from the stack
error_cube = error['error'][()]
# Get the dates for each raster from the stack
dates = list(f['dates']) # these dates appear to be given in Rata Die style: floor(Julian Day Number - 1721424.5).
# Check that the number of rasters in the data cube, error cube, and the number of dates are the same.
if data_cube.shape[0] is not len(dates) or error_cube.shape[0] is not len(dates):
print('Problem')
print('Number of rasters in data_cube: ', data_cube.shape[0])
print('Number of rasters in error_cube: ', error_cube.shape[0])
print('Number of dates: ', len(dates))
else:
print('The number of dates and rasters match.')
```
<font face='Calibri'><font size='3'><b>4.0.5 Convert from Rata Die number (similar to Julian Day number) contained in 'dates' to Gregorian date. </b></font></font>
```
tindex = []
for d in dates:
tindex.append(date.fromordinal(int(d)))
```
<font face='Calibri'><font size='3'><b>4.0.6 Create an amplitude image of the volcano.</b></font></font>
```
radar = gdal.Open(radar_tiff) # open my radar tiff using GDAL
im_radar = radar.GetRasterBand(1).ReadAsArray() # access the band information
dbplot = np.ma.log10(im_radar) # plot as a log10
dbplot[isneginf(dbplot)] = 0
```
<font face='Calibri'>
<font size='4'><b>4.1 Select a Point-of-Interest</b></font>
<br><br>
<font size='3'><b>4.1.0 Write a class to create an interactive matplotlib plot that displays an overlay of the clipped deformation map and amplitude image, and allows the user to select a point of interest.</b></font></font>
```
class pixelPicker:
def __init__(self, image1, image2, width, height):
self.x = None
self.y = None
self.fig = plt.figure(figsize=(width, height), tight_layout=True)
self.fig.tight_layout()
self.ax = self.fig.add_subplot(111, visible=False)
self.rect = patches.Rectangle(
(0.0, 0.0), width, height,
fill=False, clip_on=False, visible=False,
)
self.rect_patch = self.ax.add_patch(self.rect)
self.cid = self.rect_patch.figure.canvas.mpl_connect(
'button_press_event',
self
)
self.image1 = image1
self.image2 = image2
self.plot = self.defNradar_plot(self.image1, self.image2, self.fig, return_ax=True)
self.plot.set_title('Select a Point of Interest from db-Scaled Amplitude Image')
def defNradar_plot(self, deformation, radar, fig, return_ax=False):
ax = fig.add_subplot(111)
vmin = np.percentile(radar, 3)
vmax = np.percentile(radar, 97)
im = ax.imshow(radar, cmap='gray', vmin=vmin, vmax=vmax)
vmin = np.percentile(deformation, 3)
vmax = np.percentile(deformation, 97)
fin_plot = ax.imshow(deformation, cmap='RdBu', vmin=vmin, vmax=vmax, alpha=0.75)
cbar = fig.colorbar(fin_plot, fraction=0.24, pad=0.02)
cbar.ax.set_xlabel('mm')
cbar.ax.set_ylabel('Deformation', rotation=270, labelpad=20)
#ax.set(title="Integrated Defo [mm] Overlaid on Clipped db-Scaled Amplitude Image")
plt.grid()
if return_ax:
return(ax)
def __call__(self, event):
print('click', event)
self.x = event.xdata
self.y = event.ydata
for pnt in self.plot.get_lines():
pnt.remove()
plt.plot(self.x, self.y, 'ro')
```
<font face='Calibri'><font size='3'><b>4.1.1 Create the interactive point of interest plot, and select a point</b></font></font>
```
fig_xsize = 9
fig_ysize = 5
deformation = data_cube[data_cube.shape[0]-1]
my_plot = pixelPicker(deformation, dbplot, fig_xsize, fig_ysize)
```
<font face='Calibri' size='3'><b>4.1.2 Grab the line and pixel for the selected POI</b></font>
```
my_line = ceil(my_plot.y)
my_pixel = ceil(my_plot.x)
print(f'my_line: {my_line}')
print(f'my_pixel: {my_pixel}')
```
<font face='Calibri'>
<font size='4'><b>4.2 Visualize the data for the selected POI</b></font>
<br><br>
<font size='3'><b>4.2.0 Write a function to visualize the time series for a single pixel (POI) as well as the associated error for that pixel.</b></font></font>
```
def time_series_with_error(deformation, radar, series_d,
series_e, my_line, my_pixel,
tindex, fig_xsize, fig_ysize):
# Create base figure; 2 rows, 1 column
fig, axes = plt.subplots(nrows=2, ncols=1,
figsize=(fig_xsize,fig_ysize*2),
gridspec_kw={'width_ratios':[1],'height_ratios':[1,1]})
# Plot radar image with deformation in left column with crosshair over pixel of interest
vmin = np.percentile(radar, 3)
vmax = np.percentile(radar, 97)
axes[0].imshow(radar, cmap='gray', vmin=vmin, vmax=vmax, aspect='equal')
vmin = np.percentile(deformation, 3)
vmax = np.percentile(deformation, 97)
fin_plot = axes[0].imshow(deformation, cmap='RdBu', vmin=vmin, vmax=vmax, alpha=0.75, aspect='equal')
cbar = fig.colorbar(fin_plot, fraction=0.24, pad=0.02, ax=axes[0])
cbar.ax.set_xlabel('mm')
cbar.ax.set_ylabel('Deformation', rotation=270, labelpad=20)
axes[0].set(title="Integrated Defo [mm] Overlain on\nClipped db-Scaled Amplitude Image")
axes[0].grid() # overlay a grid on the image.
# Put crosshair over POI in first figure.
axes[0].plot(my_pixel, my_line, 'cx', markersize=12, markeredgewidth=4) # create a cyan x ('cx') at location [myPixel,myLine]
axes[0].legend(labels=['POI']) # Put in a legend with the label 'POI'
# Plot time series of pixel in right column with error bars
# Make pandas time series object
ts_d = pd.Series(series_d, index=tindex)
# Plot the time series object with the associated error.
ts_d.plot(marker='.', markersize=6, yerr=series_e)
axes[1].legend(labels=[f'Time Series for POI at ({my_line}, {my_pixel})'])
axes[1].set(title="Integrated Defo [mm] with Error Estimates for a Single Pixel")
plt.xlabel('Date')
plt.ylabel('Deformation [mm]')
for tick in axes[1].get_xticklabels():
tick.set_rotation(90)
plt.grid() # overlay a grid on the time series plot.
```
<font face='Calibri'><font size='3'><b>4.2.1 Plot the time series for the selected POI</b></font></font>
```
series_d = data_cube[:, my_line, my_pixel]
series_e = error_cube[:, my_line, my_pixel]
time_series_with_error(deformation, dbplot, series_d,
series_e, my_line, my_pixel,
tindex, fig_xsize, fig_ysize)
```
<font face='Calibri'><font size='3'><b>4.2.2 Create an animation of the deformation</b></font></font>
```
%%capture
fig = plt.figure(figsize=(14,8))
ax = fig.add_subplot(111)
#ax.axis('off')
vmin=np.percentile(data_cube.flatten(), 5)
vmax=np.percentile(data_cube.flatten(), 95)
vmin = np.percentile(deformation, 3)
vmax = np.percentile(deformation, 97)
im = ax.imshow(data_cube[0], cmap='RdBu', vmin=vmin, vmax=vmax)
ax.set_title("Animation of Deformation Time Series - Sierra Negra, Galapagos")
cbar = fig.colorbar(im)
cbar.ax.set_xlabel('mm')
cbar.ax.set_ylabel('Deformation', rotation=270, labelpad=20)
plt.grid()
def animate(i):
ax.set_title("Date: {}".format(tindex[i]))
im.set_data(data_cube[i])
ani = matplotlib.animation.FuncAnimation(fig, animate, frames=data_cube.shape[0], interval=400)
rc('animation', embed_limit=10.0**9) # set the maximum animation size allowed.
```
<font face='Calibri'><font size='3'><b>4.2.3 Display the animation</b></font></font>
```
HTML(ani.to_jshtml())
```
<font face='Calibri'><font size='3'><b>Define a prefix to prepend to all output files for identification purposes</b></font></font>
```
while True:
prefix = input("Enter an identifiable prefix to prepend to output files: ")
if re.search('\W', prefix):
print(f"\n\"{prefix}\" contains invalid characters. Please try again.")
else:
break
```
<font face='Calibri'><font size='3'><b>4.2.5 Save the animation as a gif</b></font></font>
```
ani_path = os.path.join(output_dir, f"{prefix}_filt={filt}_{date.today()}.gif")
ani.save(ani_path, writer='pillow', fps=2)
```
<font face='Calibri'><font size='3'>Note: 'date.today()' returns the current date from the system on which it is called. As OpenSAR Lab runs in machines on the East coast of the United States, date.today() will return the date based on Eastern Standard Time (EST).</font>
</font>
<font face='Calibri'><font size='5'><b>5. HDF5 TO GEOTIFF</b></font>
<br>
<font size='3'>Convert the HDF5 file to a series of geolocated geotiffs. This method comes from an answer to this <a href="https://gis.stackexchange.com/questions/37238/writing-numpy-array-to-raster-file" target="_blank">StackExchange question</a> which is based on a recipe from the <a href="https://pcjericks.github.io/py-gdalogr-cookbook/raster_layers.html#create-raster-from-array">Python GDAL/OGR Cookbook</a>.</font>
<br>
<font size='4'>
<b>5.0 Display the size of the data-stack</b>
</font>
</font>
```
print(f"Number of rasters: {data_cube.shape[0]}")
print(f"Height & Width in Pixels: {data_cube.shape[1]}, {data_cube.shape[2]}")
```
<font face='Calibri'><font size='4'><b>5.1 Get raster information from one of the subsets.</b></font></font>
```
print(files_corr[0])
src = gdal.Open(files_corr[0])
geo_transform = src.GetGeoTransform()
del src
```
<font face='Calibri'><font size='4'><b>5.2 Write a function to convert a numpy array into a geotiff</b></font></font>
```
def array_to_geotiff(in_raster, geo_transform, array, epsg):
cols = array.shape[1]
rows = array.shape[0]
driver = gdal.GetDriverByName('GTiff')
out_raster = driver.Create(in_raster, cols, rows, 1, gdal.GDT_Float32)
out_raster.SetGeoTransform(geo_transform)
outband = out_raster.GetRasterBand(1)
outband.WriteArray(array)
out_raster_srs = osr.SpatialReference()
out_raster_srs.ImportFromEPSG(epsg)
out_raster.SetProjection(out_raster_srs.ExportToWkt())
outband.FlushCache()
```
<font face='Calibri'><font size='4'><b>5.3 Create the geotiffs</b></font></font>
```
for i in range(0, data_cube.shape[0]):
try:
geotiff_path = os.path.join(output_dir, f"{prefix}_{tindex[i]}.tif")
print(f"{i}: {geotiff_path}")
array = data_cube[i, :, :].copy()
array_to_geotiff(geotiff_path, geo_transform, array, int(part1_pickle['utm']))
print(f"Geotiff {i+1} of {data_cube.shape[0]} created\nFile path: {geotiff_path}")
except:
print(f"Geotiff {i+1} of {data_cube.shape[0]} FAILED\nFile path: {geotiff_path}")
```
<font face='Calibri'><font size='5'><b>6. Alter the time filter parameter</b></font><br>
<font size='3'>Looking at the video above, you may notice that the deformation has a very smoothed appearance. This may be because of our time filter which is currently set to 1 year ('filt=1.0' in the prepxml_SBAS.py code). Let's repeat the lab from there with 2 different time filters. <br>First, using no time filter ('filt=0.0') and then using a 1 month time filter ('filt=0.082'). Change the output filename prefix for anything you want saved (e.g., 'SierraNegraDeformationTS.gif' to 'YourDesiredFileName.gif'). Otherwise, it will be overwritten. <br>
<ul>
<li>How did these changes affect the output time series?</li>
<li>How might we figure out the right filter length?</li>
<li>What does this say about the parameters we select?</li>
</ul>
</font></font>
<font face='Calibri'><font size='5'><b>7. Clear data from the Notebook (optional)</b></font>
<br>
<font size='3'>This lab has produced a large quantity of data. If you look at this notebook in your analysis_directory directory, it should now be ~80 MB. This can take a long time to load in a Jupyter Notebook. It may be useful to clear the cell outputs, which will restore the Notebook to its original size. <br>To clear the cell outputs, go Cell->All Output->Clear. This will clear the outputs of the Jupyter Notebook and restore it to its original size of ~60 kB. This will not delete any of the files we have created. </font>
</font>
<br>
<div class="alert alert-success">
<font face="Calibri" size="5"> <b> <font color='rgba(200,0,0,0.2)'> <u>ASSIGNMENT ##</u>: </font> Explore the effect of Atmospheric Correction </b> <font color='rgba(200,0,0,0.2)'> -- [## Points] </font> </font>
<font face="Calibri" size="3"> <u>Compare corrected and uncorrected time series for selected pixels</u>:
<ol type="1">
<li>Repeat the time series using the original subsets (i.e., those that TRAIN did not correct). MAKE SURE TO SAVE IT AS A DIFFERENT FILE. Hint: userfn.py will need to be modified. <font color='rgba(200,0,0,0.2)'> -- [## Point] </font></li>
<br>
<li>Show a plot of deformation through time for 3 selected pixels. Each plot should have the corrected and uncorrected time series for the selected pixels. Makes sure to select at least 1 point near the summit of the volcano (where deformation should be the greatest) and 1 point at low elevation (where deformation should be relatively small). The location of the third point is at your discretion. Hint: the variables 'myLine' and 'myPixel' can be made into a pair of lists over which a loop is run. <font color='rgba(200,0,0,0.2)'> -- [## Points] </font></li>
</ol>
</font>
</div>
<hr>
<font face="Calibri" size="2"> <i>GEOS 657-Lab9-InSARTimeSeriesAnalysis-Part2-GIAnT.ipynb - Version 3.0.1 - May 2021
<br>
<b>Version Changes:</b>
<ul>
<li>Prepare for insar_analysis conda env</li>
</ul>
</i>
</font>
</font>
| github_jupyter |
```
from collections import defaultdict
from nltk.tokenize import WordPunctTokenizer # splits all punctuations into separate tokens
from nltk.stem import WordNetLemmatizer
from nltk import pos_tag
wnl = WordNetLemmatizer()
word_punct_tokenizer = WordPunctTokenizer()
def bow_movie_nltk(root,start,end):
# list of dict where each element of bow_per_movie is bow for that movie
bow_per_movie = []
for i in range(start,end):
bow = defaultdict(float)
string = ""
for j in range(1,len(root[i])):
string += root[i][j].text
tokens = word_punct_tokenizer.tokenize(string)
l_tokens = map(lambda t: t.lower(), tokens)
### Lemmatizing using wordnetlemmatizer
l_tokens = [wnl.lemmatize(i,j[0].lower()) if j[0].lower() in ['a','n','v'] else wnl.lemmatize(i) for i,j in pos_tag(l_tokens)]
for token in l_tokens:
bow[token] += 1.0
bow_per_movie.append(bow)
return bow_per_movie
## lemmatizing positive-negative words
def pos_tagging_for_list(l):
new=[]
for i,j in pos_tag(l):
if j[0].lower() in ['a','n','v']:
new.append(wnl.lemmatize(i,j[0].lower()))
else:
new.append(wnl.lemmatize(i))
return set(new)
from sklearn.linear_model import Ridge
from sklearn import linear_model
from sklearn.tree import DecisionTreeRegressor
from sklearn.model_selection import GridSearchCV
from sklearn.neural_network import MLPRegressor
import scipy
import numpy as np
import numpy as np
def train(train_x,train_y,test_x):
clf=DecisionTreeRegressor(max_depth=10)
clf.fit(train_x, train_y)
return clf.predict(test_x)
def cal_mae(y_hat,y):
return np.mean(abs(y_hat-y))
# Function that returns a list of target variables i.e. revenue for all movies in the given set(train/dev/test) of given file root
def true_rev(start,end,root):
rev = []
for i in range(start,end):
rev.append(root[i][0].attrib['yvalue'])
rev=np.array(rev).astype(np.float)
return rev
def mpl(train_x,train_y,test_x):
clf = MLPRegressor(hidden_layer_sizes=(300,300,300,300,300),alpha=0.01)
clf.fit(train_x, train_y)
return clf.predict(test_x)
import xml.etree.ElementTree as ET
tree = ET.parse('dataset\\movies-data-v1.0\\movies-data-v1.0\\7domains-train-dev.tl.xml')
root_traindev_to = tree.getroot()
bow_per_movie_train = bow_movie_nltk(root_traindev_to,0,1147)
bow_per_movie_dev=bow_movie_nltk(root_traindev_to,1147,1464)
## uploading the positive/ negative words
pos_list=open('positive-words.txt','r').readlines()
neg_list=open('negative-words.txt','r').readlines()
## refining it
for i in range(len(pos_list)):
pos_list[i]=pos_list[i].replace('\n','')
for i in range(len(neg_list)):
neg_list[i]=neg_list[i].replace('\n','')
positive=[]
negative=[]
## lemmatizing it
positive=pos_tagging_for_list(pos_list)
negative=pos_tagging_for_list(neg_list)
## create the feature vector based on the absense and presence of pos-neg words
feature=[]
for bow in bow_per_movie_train:
feat=[]
for pos in positive:
if pos in bow.keys():
feat.append(1)
else:
feat.append(0)
for neg in negative:
if neg in bow.keys():
feat.append(-1)
else:
feat.append(0)
feature.append(feat)
feature_dev=[]
for bow in bow_per_movie_dev:
feat=[]
for pos in positive:
if pos in bow.keys():
feat.append(1)
else:
feat.append(0)
for neg in negative:
if neg in bow.keys():
feat.append(-1)
else:
feat.append(0)
feature_dev.append(feat)
f=open('train_y_to.txt', 'r')
train_y= pickle.load(f)
f=open('dev_y_to.txt', 'r')
test_y= pickle.load(f)
y_hat=mpl(feature,train_y,feature_dev)
print "MAE is ", cal_mae(test_y, y_hat)
import pickle
f=open('train_pos_feat.txt', 'w')
pickle.dump(feature,f)
f=open('dev_pos_feat.txt', 'w')
pickle.dump(feature_dev,f)
```
| github_jupyter |
# GEOL 7720 - Exercise 8
### November 27$^{th}$
### Ian Deniset
## Problem
Calculate the Z-Transform of a simple five element wavelet and design a ten element Wiener inverse filter.
## Theory
Deconvolution can be posed as a simple inverse filtering problem:
$D(\omega)=S(\omega)G(\omega)$
$G(\omega)=\frac{D(\omega)}{S(\omega)}$
$g(t)=s^{-1}(t)*d(t)$
where $g(t)$ is the Green's function, and $s(t)$ is the source function or wavelet.
This means that deconvolution can be accomplished by designing a filter as close to the inverse of the source time function as possible and subsequently convolving it with the data. For sampled data, this can be achieved by calculating and convolving the z-transform dipole inverses of the wavelet. However, this method does not work well in the presence of non minimum phase data and noise since some dipoles may not an inverse.
## Solution
### Z-Transform the Wavelet and Find Dipoles:
```
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
```
#### The initial wavelet:
The initial five element wavelet is:
$w=[-1.5,-1.25,7.5,5,-6]$
```
wavelet = np.array([-1.5,-1.25,7.5,5,-6])
print('The initial wavelet is: ', wavelet)
```
#### Z-Transform the Wavelet:
The z-transform of the wavelet is:
$W(Z)=-1.5 -1.25z+7.5z^2+5z^3-6z^4$
#### Determine Zeros:
The zeros of the z-transformed wavelet can be found numerically:
```
zCoeff = np.asarray([-6,5,7.5,-1.25,-1.5])
zeros = np.roots(zCoeff)
print('The zeros of the wavelet are: ', zeros)
```
Writing the zeros out in terms of the z-domain gives:
$W(z) = A(z-\frac{3}{2})(z-\frac{1}{2})(z+\frac{2}{3})(z+\frac{1}{2})$
where $A$ is the amplitude obtained through expansion:
$W(z)= A(z-\frac{3}{2})(z-\frac{1}{2})(z+\frac{2}{3})(z+\frac{1}{2})$
$=A\big[z^4-\frac{5}{6}z^3-\frac{5}{4}z^2+\frac{5}{24}z+\frac{1}{4}\big]$
$=-\frac{3}{2} -\frac{5}{4}z+\frac{15}{2}z^2+5z^3-6z^4$
giving an amplitude value, $A$, of $-6$:
$W(z) = -6(z-\frac{3}{2})(z-\frac{1}{2})(z+\frac{2}{3})(z+\frac{1}{2})$
This has the normalized form of:
$W(z)=-\frac{3}{2}(1-\frac{2}{3}z)(1-2z)(1+\frac{3}{2}z)(1+2z)$
Since some coefficients are less than or equal to 1, while others are greater than 1, the wavelet is mixed phase. This can be better visualized by plotting since minimum phase dipoles plot outside the unit circle in the complex plane while maximum phase dipoles plot inside.
```
t = np.arange(0,2*np.pi,0.001)
fig, ax = plt.subplots(figsize=(10,10))
ax.plot(np.cos(t), np.sin(t),ls='--')
ax.axhline(y=0,c='k')
ax.axvline(x=0,c='k')
ax.scatter(zeros,[0,0,0,0],c='g',s=90, label='Poles')
ax.legend()
ax.set_xlim(-1.75,1.75)
ax.set_ylim(-1.75,1.75)
ax.grid()
plt.show()
```
With the dipoles and associated phase found, the inverse of each can be written out as a series expansion to investigate convergence:
##### Zero of 3/2:
$(z-\frac{3}{2})^{-1}=-\frac{1}{a}-\frac{z}{a^2}-\frac{z^2}{a^3}...$
$=-\frac{1}{3/2}-\frac{1}{3/2^2}-\frac{1}{3/2^3}-\frac{1}{3/2^4}-\frac{1}{3/2^5}-\frac{1}{3/2^6}...=-0.66-0.44-0.29-0.19-0.13-0.087...$
##### Zero of 1/2:
$(z-\frac{1}{2})^{-1}=z^{-1}+az^{-2}+a^2z^{-3}+a^3z^{-4}...$
$=1^{-1}+\frac{1}{2}1^{-2}+\frac{1}{2}^{2}1^{-3}...=1+0.5+0.25+0.125+0.0625+0.0312...$
##### Zero of -2/3:
$(z-\frac{2}{3})^{-1}=z^{-1}+az^{-2}+a^2z^{-3}+a^3z^{-4}...$
$=1^{-1}+-\frac{2}{3}1^{-2}+-\frac{2}{3}^{2}1^{-3}...=1-0.66+0.44-0.29+0.19-0.13+0.087...$
##### Zero of -1/2:
$(z-\frac{1}{2})^{-1}=z^{-1}+az^{-2}+a^2z^{-3}+a^3z^{-4}...$
$=1^{-1}+-\frac{1}{2}1^{-2}+-\frac{1}{2}^{2}1^{-3}...=1-0.5+0.25-0.125+0.0625-0.0312...$
As can be seen above, since the zeros for this wavelet lie close to the unit circle, convergence for each dipole is somewhat slow.
## Wiener Filtering
To avoid the issue of minimum and maximum phase dipoles as well as problems with slow convergence, a Wiener filtering approach can be used to find the inverse of a wavelet. The inverse filter is obtained through a damped least squares inversion of the form:
$\begin{bmatrix} w_0 & 0 & \cdots & 0 & 0 \\
w_1 & w_0 & \cdots & 0 & 0 \\
w_2 & w_1 & \cdots & 0 & 0 \\
\vdots & \vdots & \ddots & \vdots & \vdots \\
0 & 0 & \cdots & w_m & w_{m-1} \\
0 & 0 & \cdots & 0 & w_m \end{bmatrix}
\begin{bmatrix} f_0 \\ f_1 \\ \vdots \\ f_n \end{bmatrix}
=
\begin{bmatrix} 1 \\ 0\\ 0 \\ \vdots \\ 0 \end{bmatrix}$
where $w$ is the wavelet, $f$ is the desired inverse filter, and the right hand side is an impulse response containing a single spike. The location of the spike can be moved in order to handle acausality.
Using the above formulation, a ten point inverse filter for the initial wavelet can be found:
###### Create Wavelet Matrix:
```
#parameters
filterLen = 10
waveletLen = len(wavelet)
numRows = filterLen + (waveletLen-1)
#create the wavelet matrix and ensure it is the correct size
waveletMatrix = np.zeros((numRows,filterLen))
col = 0
for row in range(filterLen):
waveletMatrix[row:(row+5),col]=wavelet
col += 1
print('The shape of the wavelet matrix is :', waveletMatrix.shape)
print('\nThe wavelet matrix is of the form: \n', waveletMatrix)
```
###### Set up inverse problem with damping and solve
An initial first past inversion can be completed to investigate the impact of different damping weights and spike locations.
Each calculated inverse filter will be convolved with the original wavelet to investigate how well the original impulse response series is recovered.
```
#damping values to try
dampVal = np.asarray([0.001,0.01,0.1,1.0,10,100])
filterDamp = []
for dampingV in range(len(dampVal)):
for spikeLoc in range(8):
eyeDamp = dampVal[dampingV]*np.identity(len(waveletMatrix[0,:]))
augA = np.concatenate((waveletMatrix,eyeDamp))
#print(augA.shape)
#set spike location
response = np.zeros((numRows,1))
response[spikeLoc] = 1
#print(spikeLoc)
zeros = np.zeros((len(waveletMatrix[0,:]),1))
augD = np.concatenate((response,zeros))
#print(augD.shape)
ans = np.linalg.lstsq(augA,augD)
filterDamp.append(ans[0])
#plot the resulting models
fig, ax = plt.subplots(6,8,figsize=(15,10))
i=0
for row in range(6):
for col in range(8):
ax[row,col].plot(np.convolve(wavelet,filterDamp[i][:,0]), label='Recovered Response')
ax[row,col].plot(wavelet/7.5, label='Initial Wavelet')
if col > 0:
ax[row,col].set_yticklabels([])
if row < 5:
ax[row,col].set_xticklabels([])
if col == 0:
ax[row,col].set_ylabel('Damp: %g' %dampVal[row])
if row == 0:
ax[row,col].set_xlabel('Spike Loc: %g' %col)
ax[row,col].xaxis.set_label_position('top')
i+=1
fig.legend()
plt.show()
```
Looking at the above plots it can be seen that only a small amount of damping is required, and that due to the mostly maximum phase nature of the wavelet, the spike location is best placed at some point after the first sample to allow for acausality.
With a general idea of the best parameters to use, a second inversion can be run to further refine the result.
```
#damping values to try
dampVal = np.asarray([0.001,0.01,0.1,1.0])
filterDamp = []
for dampingV in range(len(dampVal)):
for spikeLoc in range(6):
eyeDamp = dampVal[dampingV]*np.identity(len(waveletMatrix[0,:]))
augA = np.concatenate((waveletMatrix,eyeDamp))
#print(augA.shape)
#set spike location
response = np.zeros((numRows,1))
response[spikeLoc+3] = 1
#print(spikeLoc)
zeros = np.zeros((len(waveletMatrix[0,:]),1))
augD = np.concatenate((response,zeros))
#print(augD.shape)
ans = np.linalg.lstsq(augA,augD)
filterDamp.append(ans[0])
#plot the resulting models
fig, ax = plt.subplots(4,6,figsize=(15,10))
i=0
for row in range(4):
for col in range(6):
ax[row,col].plot(np.convolve(wavelet,filterDamp[i][:,0]), label='Recovered Response')
ax[row,col].plot(wavelet/7.5, label='Initial Wavelet')
if col > 0:
ax[row,col].set_yticklabels([])
if row < 5:
ax[row,col].set_xticklabels([])
if col == 0:
ax[row,col].set_ylabel('Damp: %g' %dampVal[row])
if row == 0:
ax[row,col].set_xlabel('Spike Loc: %g' %(col+3))
ax[row,col].xaxis.set_label_position('top')
i+=1
fig.legend()
plt.show()
```
Visually, there appears to be almost no difference between the recovered response based on damping value alone. On the other hand, the location of the spike significantly influences the response, with a spike placed at the sixth sample point giving the highest recovered amplitude and reasonably flat side lobes. Based on this, a damping value of 1 and spike location of 6 will be selected as the final parameters for the inverse filter. The results for these values along with the final inverse filter are plotted in more detail below.
```
response[6]=1
response[8]=0
fig, ax = plt.subplots(figsize=(15,10))
ax.plot(np.convolve(wavelet,filterDamp[21][:,0]), label='Recovered Response', alpha=0.75)
ax.plot(wavelet/7.5, label='Initial Wavelet', alpha=0.75, ls='--')
ax.plot(response, label='Original Response', alpha=0.75, ls='--')
ax.plot(filterDamp[21][:,0]*7.5, label='Final Inverse Filter', alpha=0.75)
ax.grid(which='both')
plt.legend()
plt.show()
```
| github_jupyter |
*Python Machine Learning 2nd Edition* by [Sebastian Raschka](https://sebastianraschka.com), Packt Publishing Ltd. 2017
Code Repository: https://github.com/rasbt/python-machine-learning-book-2nd-edition
Code License: [MIT License](https://github.com/rasbt/python-machine-learning-book-2nd-edition/blob/master/LICENSE.txt)
```
from google.colab import drive
drive.mount('/content/drive')
```
# Implementing a Multi-Layer Perceptron learning algorithm in Python
```
from IPython.display import Image
%matplotlib inline
from keras import layers
from keras import models
from keras.utils import to_categorical
```
# Classifying handwritten digits
## An object-oriented multi-layer perceptron API
```
import numpy as np
import sys
class NeuralNetMLP(object):
""" Feedforward neural network / Multi-layer perceptron classifier.
Parameters
------------
n_hidden : int (default: 30)
Number of hidden units.
l2 : float (default: 0.)
Lambda value for L2-regularization.
No regularization if l2=0. (default)
epochs : int (default: 100)
Number of passes over the training set.
eta : float (default: 0.001)
Learning rate.
shuffle : bool (default: True)
Shuffles training data every epoch if True to prevent circles.
minibatch_size : int (default: 1)
Number of training samples per minibatch.
seed : int (default: None)
Random seed for initalizing weights and shuffling.
Attributes
-----------
eval_ : dict
Dictionary collecting the cost, training accuracy,
and validation accuracy for each epoch during training.
"""
def __init__(self, n_hidden=30,
l2=0., epochs=100, eta=0.001,
shuffle=True, minibatch_size=1, seed=None):
self.random = np.random.RandomState(seed)
self.n_hidden = n_hidden
self.l2 = l2
self.epochs = epochs
self.eta = eta
self.shuffle = shuffle
self.minibatch_size = minibatch_size
def _onehot(self, y, n_classes):
"""Encode labels into one-hot representation
Parameters
------------
y : array, shape = [n_samples]
Target values.
Returns
-----------
onehot : array, shape = (n_samples, n_labels)
"""
onehot = np.zeros((n_classes, y.shape[0]))
for idx, val in enumerate(y.astype(int)):
onehot[val, idx] = 1.
return onehot.T
def _sigmoid(self, z):
"""Compute logistic function (sigmoid)"""
return 1. / (1. + np.exp(-np.clip(z, -250, 250)))
def _forward(self, X):
"""Compute forward propagation step"""
# step 1: net input of hidden layer
# [n_samples, n_features] dot [n_features, n_hidden]
# -> [n_samples, n_hidden]
z_h = np.dot(X, self.w_h) + self.b_h
# step 2: activation of hidden layer
a_h = self._sigmoid(z_h)
# step 3: net input of output layer
# [n_samples, n_hidden] dot [n_hidden, n_classlabels]
# -> [n_samples, n_classlabels]
z_out = np.dot(a_h, self.w_out) + self.b_out
# step 4: activation output layer
a_out = self._sigmoid(z_out)
return z_h, a_h, z_out, a_out
def _compute_cost(self, y_enc, output):
"""Compute cost function.
Parameters
----------
y_enc : array, shape = (n_samples, n_labels)
one-hot encoded class labels.
output : array, shape = [n_samples, n_output_units]
Activation of the output layer (forward propagation)
Returns
---------
cost : float
Regularized cost
"""
L2_term = (self.l2 *
(np.sum(self.w_h ** 2.) +
np.sum(self.w_out ** 2.)))
term1 = -y_enc * (np.log(output))
term2 = (1. - y_enc) * np.log(1. - output)
cost = np.sum(term1 - term2) + L2_term
# If you are applying this cost function to other
# datasets where activation
# values maybe become more extreme (closer to zero or 1)
# you may encounter "ZeroDivisionError"s due to numerical
# instabilities in Python & NumPy for the current implementation.
# I.e., the code tries to evaluate log(0), which is undefined.
# To address this issue, you could add a small constant to the
# activation values that are passed to the log function.
#
# For example:
#
# term1 = -y_enc * (np.log(output + 1e-5))
# term2 = (1. - y_enc) * np.log(1. - output + 1e-5)
return cost
def predict(self, X):
"""Predict class labels
Parameters
-----------
X : array, shape = [n_samples, n_features]
Input layer with original features.
Returns:
----------
y_pred : array, shape = [n_samples]
Predicted class labels.
"""
z_h, a_h, z_out, a_out = self._forward(X)
y_pred = np.argmax(z_out, axis=1)
return y_pred
def fit(self, X_train, y_train, X_valid, y_valid):
""" Learn weights from training data.
Parameters
-----------
X_train : array, shape = [n_samples, n_features]
Input layer with original features.
y_train : array, shape = [n_samples]
Target class labels.
X_valid : array, shape = [n_samples, n_features]
Sample features for validation during training
y_valid : array, shape = [n_samples]
Sample labels for validation during training
Returns:
----------
self
"""
n_output = np.unique(y_train).shape[0] # number of class labels
n_features = X_train.shape[1]
########################
# Weight initialization
########################
# weights for input -> hidden
self.b_h = np.zeros(self.n_hidden)
self.w_h = self.random.normal(loc=0.0, scale=0.1,
size=(n_features, self.n_hidden))
# weights for hidden -> output
self.b_out = np.zeros(n_output)
self.w_out = self.random.normal(loc=0.0, scale=0.1,
size=(self.n_hidden, n_output))
epoch_strlen = len(str(self.epochs)) # for progress formatting
self.eval_ = {'cost': [], 'train_acc': [], 'valid_acc': []}
y_train_enc = self._onehot(y_train, n_output)
# iterate over training epochs
for i in range(self.epochs):
# iterate over minibatches
indices = np.arange(X_train.shape[0])
if self.shuffle:
self.random.shuffle(indices)
for start_idx in range(0, indices.shape[0] - self.minibatch_size +
1, self.minibatch_size):
batch_idx = indices[start_idx:start_idx + self.minibatch_size]
# forward propagation
z_h, a_h, z_out, a_out = self._forward(X_train[batch_idx])
##################
# Backpropagation
##################
# [n_samples, n_classlabels]
sigma_out = a_out - y_train_enc[batch_idx]
# [n_samples, n_hidden]
sigmoid_derivative_h = a_h * (1. - a_h)
# [n_samples, n_classlabels] dot [n_classlabels, n_hidden]
# -> [n_samples, n_hidden]
sigma_h = (np.dot(sigma_out, self.w_out.T) *
sigmoid_derivative_h)
# [n_features, n_samples] dot [n_samples, n_hidden]
# -> [n_features, n_hidden]
grad_w_h = np.dot(X_train[batch_idx].T, sigma_h)
grad_b_h = np.sum(sigma_h, axis=0)
# [n_hidden, n_samples] dot [n_samples, n_classlabels]
# -> [n_hidden, n_classlabels]
grad_w_out = np.dot(a_h.T, sigma_out)
grad_b_out = np.sum(sigma_out, axis=0)
# Regularization and weight updates
delta_w_h = (grad_w_h + self.l2*self.w_h)
delta_b_h = grad_b_h # bias is not regularized
self.w_h -= self.eta * delta_w_h
self.b_h -= self.eta * delta_b_h
delta_w_out = (grad_w_out + self.l2*self.w_out)
delta_b_out = grad_b_out # bias is not regularized
self.w_out -= self.eta * delta_w_out
self.b_out -= self.eta * delta_b_out
#############
# Evaluation
#############
# Evaluation after each epoch during training
z_h, a_h, z_out, a_out = self._forward(X_train)
cost = self._compute_cost(y_enc=y_train_enc,
output=a_out)
y_train_pred = self.predict(X_train)
y_valid_pred = self.predict(X_valid)
train_acc = ((np.sum(y_train == y_train_pred)).astype(np.float) /
X_train.shape[0])
valid_acc = ((np.sum(y_valid == y_valid_pred)).astype(np.float) /
X_valid.shape[0])
sys.stderr.write('\r%0*d/%d | Cost: %.2f '
'| Train/Valid Acc.: %.2f%%/%.2f%% ' %
(epoch_strlen, i+1, self.epochs, cost,
train_acc*100, valid_acc*100))
sys.stderr.flush()
self.eval_['cost'].append(cost)
self.eval_['train_acc'].append(train_acc)
self.eval_['valid_acc'].append(valid_acc)
return self
```
## Obtaining the MNIST dataset
The MNIST dataset is publicly available at http://yann.lecun.com/exdb/mnist/ and consists of the following four parts:
- Training set images: train-images-idx3-ubyte.gz (9.9 MB, 47 MB unzipped, 60,000 samples)
- Training set labels: train-labels-idx1-ubyte.gz (29 KB, 60 KB unzipped, 60,000 labels)
- Test set images: t10k-images-idx3-ubyte.gz (1.6 MB, 7.8 MB, 10,000 samples)
- Test set labels: t10k-labels-idx1-ubyte.gz (5 KB, 10 KB unzipped, 10,000 labels)
After downloading the files, simply run the next code cell to unzip the files.
```
# this code cell unzips mnist
import sys
import gzip
import shutil
import os
if (sys.version_info > (3, 0)):
writemode = 'wb'
else:
writemode = 'w'
zipped_mnist = [f for f in os.listdir('./') if f.endswith('ubyte.gz')]
for z in zipped_mnist:
with gzip.GzipFile(z, mode='rb') as decompressed, open(z[:-3], writemode) as outfile:
outfile.write(decompressed.read())
import os
import struct
import numpy as np
def load_mnist(path, kind='train'):
os.chdir('/content/drive/My Drive/deeplearning_cursoCienciaDados_Insight/notebooks/data')
"""Load MNIST data from `path`"""
labels_path = os.path.join(path,
'%s-labels-idx1-ubyte' % kind)
images_path = os.path.join(path,
'%s-images-idx3-ubyte' % kind)
with open(labels_path, 'rb') as lbpath:
magic, n = struct.unpack('>II',
lbpath.read(8))
labels = np.fromfile(lbpath,
dtype=np.uint8)
with open(images_path, 'rb') as imgpath:
magic, num, rows, cols = struct.unpack(">IIII",
imgpath.read(16))
images = np.fromfile(imgpath,
dtype=np.uint8).reshape(len(labels), 784)
images = ((images / 255.) - .5) * 2
return images, labels
!ls
path = ''
X_train, y_train = load_mnist(path, kind='train')
print('Rows: %d, columns: %d' % (X_train.shape[0], X_train.shape[1]))
X_test, y_test = load_mnist(path, kind='t10k')
print('Rows: %d, columns: %d' % (X_test.shape[0], X_test.shape[1]))
```
## Visualize the first digit of each class:
```
import matplotlib.pyplot as plt
fig, ax = plt.subplots(nrows=2, ncols=5, sharex=True, sharey=True,)
ax = ax.flatten()
for i in range(10):
img = X_train[y_train == i][0].reshape(28, 28)
ax[i].imshow(img, cmap='Greys')
ax[0].set_xticks([])
ax[0].set_yticks([])
plt.tight_layout()
# plt.savefig('images/12_5.png', dpi=300)
plt.show()
```
## Visualize 25 different versions of "7":
```
fig, ax = plt.subplots(nrows=5, ncols=5, sharex=True, sharey=True,)
ax = ax.flatten()
for i in range(25):
img = X_train[y_train == 7][i].reshape(28, 28)
ax[i].imshow(img, cmap='Greys')
ax[0].set_xticks([])
ax[0].set_yticks([])
plt.tight_layout()
# plt.savefig('images/12_6.png', dpi=300)
plt.show()
X_train.shape, X_test.shape, y_train.shape, y_test.shape
n_epochs = 200
nn = NeuralNetMLP(n_hidden=100,
l2=0.01,
epochs=n_epochs,
eta=0.0005,
minibatch_size=100,
shuffle=True,
seed=1)
nn.fit(X_train=X_train[:55000],
y_train=y_train[:55000],
X_valid=X_train[55000:],
y_valid=y_train[55000:])
import matplotlib.pyplot as plt
plt.plot(range(nn.epochs), nn.eval_['cost'])
plt.ylabel('Cost')
plt.xlabel('Epochs')
#plt.savefig('images/12_07.png', dpi=300)
plt.show()
plt.plot(range(nn.epochs), nn.eval_['train_acc'],
label='training')
plt.plot(range(nn.epochs), nn.eval_['valid_acc'],
label='validation', linestyle='--')
plt.ylabel('Accuracy')
plt.xlabel('Epochs')
plt.legend()
#plt.savefig('images/12_08.png', dpi=300)
plt.show()
y_test_pred = nn.predict(X_test)
acc = (np.sum(y_test == y_test_pred)
.astype(np.float) / X_test.shape[0])
print('Test accuracy: %.2f%%' % (acc * 100))
miscl_img = X_test[y_test != y_test_pred][:25]
correct_lab = y_test[y_test != y_test_pred][:25]
miscl_lab = y_test_pred[y_test != y_test_pred][:25]
fig, ax = plt.subplots(nrows=5, ncols=5, sharex=True, sharey=True,)
ax = ax.flatten()
for i in range(25):
img = miscl_img[i].reshape(28, 28)
ax[i].imshow(img, cmap='Greys', interpolation='nearest')
ax[i].set_title('%d) t: %d p: %d' % (i+1, correct_lab[i], miscl_lab[i]))
ax[0].set_xticks([])
ax[0].set_yticks([])
plt.tight_layout()
#plt.savefig('images/12_09.png', dpi=300)
plt.show()
```
| github_jupyter |
```
%matplotlib inline
```
(beta) Static Quantization with Eager Mode in PyTorch
=========================================================
**Author**: `Raghuraman Krishnamoorthi <https://github.com/raghuramank100>`_
**Edited by**: `Parth Patel <https://github.com/ParthPatel-ES>`_
This tutorial shows how to do post-training static quantization, as well as illustrating
two more advanced techniques - per-channel quantization and quantization-aware training -
to further improve the model's accuracy. Note that quantization is currently only supported
for CPUs, so we will not be utilizing GPUs / CUDA in this tutorial.
By the end of this tutorial, you will see how quantization in PyTorch can result in
significant decreases in model size while increasing speed. Furthermore, you'll see how
to easily apply some advanced quantization techniques shown
`here <https://arxiv.org/abs/1806.08342>`_ so that your quantized models take much less
of an accuracy hit than they would otherwise.
Warning: we use a lot of boilerplate code from other PyTorch repos to, for example,
define the ``MobileNetV2`` model archtecture, define data loaders, and so on. We of course
encourage you to read it; but if you want to get to the quantization features, feel free
to skip to the "4. Post-training static quantization" section.
We'll start by doing the necessary imports:
```
import numpy as np
import torch
import torch.nn as nn
import torchvision
from torch.utils.data import DataLoader
from torchvision import datasets
import torchvision.transforms as transforms
import os
import time
import sys
import torch.quantization
# # Setup warnings
import warnings
warnings.filterwarnings(
action='ignore',
category=DeprecationWarning,
module=r'.*'
)
warnings.filterwarnings(
action='default',
module=r'torch.quantization'
)
# Specify random seed for repeatable results
torch.manual_seed(191009)
```
1. Model architecture
---------------------
We first define the MobileNetV2 model architecture, with several notable modifications
to enable quantization:
- Replacing addition with ``nn.quantized.FloatFunctional``
- Insert ``QuantStub`` and ``DeQuantStub`` at the beginning and end of the network.
- Replace ReLU6 with ReLU
Note: this code is taken from
`here <https://github.com/pytorch/vision/blob/master/torchvision/models/mobilenet.py>`_.
```
from torch.quantization import QuantStub, DeQuantStub
def _make_divisible(v, divisor, min_value=None):
"""
This function is taken from the original tf repo.
It ensures that all layers have a channel number that is divisible by 8
It can be seen here:
https://github.com/tensorflow/models/blob/master/research/slim/nets/mobilenet/mobilenet.py
:param v:
:param divisor:
:param min_value:
:return:
"""
if min_value is None:
min_value = divisor
new_v = max(min_value, int(v + divisor / 2) // divisor * divisor)
# Make sure that round down does not go down by more than 10%.
if new_v < 0.9 * v:
new_v += divisor
return new_v
class ConvBNReLU(nn.Sequential):
def __init__(self, in_planes, out_planes, kernel_size=3, stride=1, groups=1):
padding = (kernel_size - 1) // 2
super(ConvBNReLU, self).__init__(
nn.Conv2d(in_planes, out_planes, kernel_size, stride, padding, groups=groups, bias=False),
nn.BatchNorm2d(out_planes, momentum=0.1),
# Replace with ReLU
nn.ReLU(inplace=False)
)
class InvertedResidual(nn.Module):
def __init__(self, inp, oup, stride, expand_ratio):
super(InvertedResidual, self).__init__()
self.stride = stride
assert stride in [1, 2]
hidden_dim = int(round(inp * expand_ratio))
self.use_res_connect = self.stride == 1 and inp == oup
layers = []
if expand_ratio != 1:
# pw
layers.append(ConvBNReLU(inp, hidden_dim, kernel_size=1))
layers.extend([
# dw
ConvBNReLU(hidden_dim, hidden_dim, stride=stride, groups=hidden_dim),
# pw-linear
nn.Conv2d(hidden_dim, oup, 1, 1, 0, bias=False),
nn.BatchNorm2d(oup, momentum=0.1),
])
self.conv = nn.Sequential(*layers)
# Replace torch.add with floatfunctional
self.skip_add = nn.quantized.FloatFunctional()
def forward(self, x):
if self.use_res_connect:
return self.skip_add.add(x, self.conv(x))
else:
return self.conv(x)
class MobileNetV2(nn.Module):
def __init__(self, num_classes=1000, width_mult=1.0, inverted_residual_setting=None, round_nearest=8):
"""
MobileNet V2 main class
Args:
num_classes (int): Number of classes
width_mult (float): Width multiplier - adjusts number of channels in each layer by this amount
inverted_residual_setting: Network structure
round_nearest (int): Round the number of channels in each layer to be a multiple of this number
Set to 1 to turn off rounding
"""
super(MobileNetV2, self).__init__()
block = InvertedResidual
input_channel = 32
last_channel = 1280
if inverted_residual_setting is None:
inverted_residual_setting = [
# t, c, n, s
[1, 16, 1, 1],
[6, 24, 2, 2],
[6, 32, 3, 2],
[6, 64, 4, 2],
[6, 96, 3, 1],
[6, 160, 3, 2],
[6, 320, 1, 1],
]
# only check the first element, assuming user knows t,c,n,s are required
if len(inverted_residual_setting) == 0 or len(inverted_residual_setting[0]) != 4:
raise ValueError("inverted_residual_setting should be non-empty "
"or a 4-element list, got {}".format(inverted_residual_setting))
# building first layer
input_channel = _make_divisible(input_channel * width_mult, round_nearest)
self.last_channel = _make_divisible(last_channel * max(1.0, width_mult), round_nearest)
features = [ConvBNReLU(3, input_channel, stride=2)]
# building inverted residual blocks
for t, c, n, s in inverted_residual_setting:
output_channel = _make_divisible(c * width_mult, round_nearest)
for i in range(n):
stride = s if i == 0 else 1
features.append(block(input_channel, output_channel, stride, expand_ratio=t))
input_channel = output_channel
# building last several layers
features.append(ConvBNReLU(input_channel, self.last_channel, kernel_size=1))
# make it nn.Sequential
self.features = nn.Sequential(*features)
self.quant = QuantStub()
self.dequant = DeQuantStub()
# building classifier
self.classifier = nn.Sequential(
nn.Dropout(0.2),
nn.Linear(self.last_channel, num_classes),
)
# weight initialization
for m in self.modules():
if isinstance(m, nn.Conv2d):
nn.init.kaiming_normal_(m.weight, mode='fan_out')
if m.bias is not None:
nn.init.zeros_(m.bias)
elif isinstance(m, nn.BatchNorm2d):
nn.init.ones_(m.weight)
nn.init.zeros_(m.bias)
elif isinstance(m, nn.Linear):
nn.init.normal_(m.weight, 0, 0.01)
nn.init.zeros_(m.bias)
def forward(self, x):
x = self.quant(x)
x = self.features(x)
x = x.mean([2, 3])
x = self.classifier(x)
x = self.dequant(x)
return x
# Fuse Conv+BN and Conv+BN+Relu modules prior to quantization
# This operation does not change the numerics
def fuse_model(self):
for m in self.modules():
if type(m) == ConvBNReLU:
torch.quantization.fuse_modules(m, ['0', '1', '2'], inplace=True)
if type(m) == InvertedResidual:
for idx in range(len(m.conv)):
if type(m.conv[idx]) == nn.Conv2d:
torch.quantization.fuse_modules(m.conv, [str(idx), str(idx + 1)], inplace=True)
```
2. Helper functions
-------------------
We next define several helper functions to help with model evaluation. These mostly come from
`here <https://github.com/pytorch/examples/blob/master/imagenet/main.py>`_.
```
class AverageMeter(object):
"""Computes and stores the average and current value"""
def __init__(self, name, fmt=':f'):
self.name = name
self.fmt = fmt
self.reset()
def reset(self):
self.val = 0
self.avg = 0
self.sum = 0
self.count = 0
def update(self, val, n=1):
self.val = val
self.sum += val * n
self.count += n
self.avg = self.sum / self.count
def __str__(self):
fmtstr = '{name} {val' + self.fmt + '} ({avg' + self.fmt + '})'
return fmtstr.format(**self.__dict__)
def accuracy(output, target, topk=(1,)):
"""Computes the accuracy over the k top predictions for the specified values of k"""
with torch.no_grad():
maxk = max(topk)
batch_size = target.size(0)
_, pred = output.topk(maxk, 1, True, True)
pred = pred.t()
correct = pred.eq(target.view(1, -1).expand_as(pred))
res = []
for k in topk:
correct_k = correct[:k].view(-1).float().sum(0, keepdim=True)
res.append(correct_k.mul_(100.0 / batch_size))
return res
def evaluate(model, criterion, data_loader, neval_batches):
model.eval()
top1 = AverageMeter('Acc@1', ':6.2f')
top5 = AverageMeter('Acc@5', ':6.2f')
cnt = 0
with torch.no_grad():
for image, target in data_loader:
output = model(image)
loss = criterion(output, target)
cnt += 1
acc1, acc5 = accuracy(output, target, topk=(1, 5))
print('.', end = '')
top1.update(acc1[0], image.size(0))
top5.update(acc5[0], image.size(0))
if cnt >= neval_batches:
return top1, top5
return top1, top5
def load_model(model_file):
model = MobileNetV2()
state_dict = torch.load(model_file)
model.load_state_dict(state_dict)
model.to('cpu')
return model
def print_size_of_model(model):
torch.save(model.state_dict(), "temp.p")
print('Size (MB):', os.path.getsize("temp.p")/1e6)
os.remove('temp.p')
```
3. Define dataset and data loaders
----------------------------------
As our last major setup step, we define our dataloaders for our training and testing set.
The specific dataset we've created for this tutorial contains just 1000 images from the ImageNet data, one from
each class (this dataset, at just over 250 MB, is small enough that it can be downloaded
relatively easily). The URL for this custom dataset is:
https://s3.amazonaws.com/pytorch-tutorial-assets/imagenet_1k.zip
For the tutorial to run, you can also download this data and move it to the right place using
`these lines <https://github.com/pytorch/tutorials/blob/master/Makefile#L97-L98>`_
from the `Makefile <https://github.com/pytorch/tutorials/blob/master/Makefile>` if not on Google Colab.
To run the code in this tutorial using the entire ImageNet dataset, on the other hand, you could download
the data using ``torchvision`` following
`here <https://pytorch.org/docs/stable/torchvision/datasets.html#imagenet>`_. Which might not be publicly available.
```
import requests
import zipfile
url = 'https://s3.amazonaws.com/pytorch-tutorial-assets/imagenet_1k.zip'
r = requests.get(url, allow_redirects=True)
open('imagenet_1k.zip', 'wb').write(r.content)
with zipfile.ZipFile('/content/imagenet_1k.zip', 'r') as zip_ref:
zip_ref.extractall('./data')
def prepare_data_loaders(data_path):
traindir = os.path.join(data_path, 'train')
valdir = os.path.join(data_path, 'val')
normalize = transforms.Normalize(mean=[0.485, 0.456, 0.406],
std=[0.229, 0.224, 0.225])
dataset = torchvision.datasets.ImageFolder(
traindir,
transforms.Compose([
transforms.RandomResizedCrop(224),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
normalize,
]))
dataset_test = torchvision.datasets.ImageFolder(
valdir,
transforms.Compose([
transforms.Resize(256),
transforms.CenterCrop(224),
transforms.ToTensor(),
normalize,
]))
train_sampler = torch.utils.data.RandomSampler(dataset)
test_sampler = torch.utils.data.SequentialSampler(dataset_test)
data_loader = torch.utils.data.DataLoader(
dataset, batch_size=train_batch_size,
sampler=train_sampler)
data_loader_test = torch.utils.data.DataLoader(
dataset_test, batch_size=eval_batch_size,
sampler=test_sampler)
return data_loader, data_loader_test
```
Next, we'll load in the pre-trained MobileNetV2 model. We provide the URL to download the data from in ``torchvision``
`here <https://github.com/pytorch/vision/blob/master/torchvision/models/mobilenet.py#L9>`_.
```
url = 'https://download.pytorch.org/models/mobilenet_v2-b0353104.pth'
r = requests.get(url, allow_redirects=True)
open('./data/mobilenet_pretrained_float.pth', 'wb').write(r.content)
data_path = 'data/imagenet_1k'
saved_model_dir = 'data/'
float_model_file = 'mobilenet_pretrained_float.pth'
scripted_float_model_file = 'mobilenet_quantization_scripted.pth'
scripted_quantized_model_file = 'mobilenet_quantization_scripted_quantized.pth'
train_batch_size = 30
eval_batch_size = 30
data_loader, data_loader_test = prepare_data_loaders(data_path)
criterion = nn.CrossEntropyLoss()
float_model = load_model(saved_model_dir + float_model_file).to('cpu')
```
Next, we'll "fuse modules"; this can both make the model faster by saving on memory access
while also improving numerical accuracy. While this can be used with any model, this is
especially common with quantized models.
```
print('\n Inverted Residual Block: Before fusion \n\n', float_model.features[1].conv)
float_model.eval()
# Fuses modules
float_model.fuse_model()
# Note fusion of Conv+BN+Relu and Conv+Relu
print('\n Inverted Residual Block: After fusion\n\n',float_model.features[1].conv)
```
Finally to get a "baseline" accuracy, let's see the accuracy of our un-quantized model
with fused modules
```
num_eval_batches = 10
print("Size of baseline model")
print_size_of_model(float_model)
top1, top5 = evaluate(float_model, criterion, data_loader_test, neval_batches=num_eval_batches)
print('Evaluation accuracy on %d images, %2.2f'%(num_eval_batches * eval_batch_size, top1.avg))
torch.jit.save(torch.jit.script(float_model), saved_model_dir + scripted_float_model_file)
```
We see 78% accuracy on 300 images, a solid baseline for ImageNet,
especially considering our model is just 14.0 MB.
This will be our baseline to compare to. Next, let's try different quantization methods
4. Post-training static quantization
------------------------------------
Post-training static quantization involves not just converting the weights from float to int,
as in dynamic quantization, but also performing the additional step of first feeding batches
of data through the network and computing the resulting distributions of the different activations
(specifically, this is done by inserting `observer` modules at different points that record this
data). These distributions are then used to determine how the specifically the different activations
should be quantized at inference time (a simple technique would be to simply divide the entire range
of activations into 256 levels, but we support more sophisticated methods as well). Importantly,
this additional step allows us to pass quantized values between operations instead of converting these
values to floats - and then back to ints - between every operation, resulting in a significant speed-up.
```
num_calibration_batches = 10
myModel = load_model(saved_model_dir + float_model_file).to('cpu')
myModel.eval()
# Fuse Conv, bn and relu
myModel.fuse_model()
# Specify quantization configuration
# Start with simple min/max range estimation and per-tensor quantization of weights
myModel.qconfig = torch.quantization.default_qconfig
print(myModel.qconfig)
torch.quantization.prepare(myModel, inplace=True)
# Calibrate first
print('Post Training Quantization Prepare: Inserting Observers')
print('\n Inverted Residual Block:After observer insertion \n\n', myModel.features[1].conv)
# Calibrate with the training set
evaluate(myModel, criterion, data_loader, neval_batches=num_calibration_batches)
print('Post Training Quantization: Calibration done')
# Convert to quantized model
torch.quantization.convert(myModel, inplace=True)
print('Post Training Quantization: Convert done')
print('\n Inverted Residual Block: After fusion and quantization, note fused modules: \n\n',myModel.features[1].conv)
print("Size of model after quantization")
print_size_of_model(myModel)
top1, top5 = evaluate(myModel, criterion, data_loader_test, neval_batches=num_eval_batches)
print('Evaluation accuracy on %d images, %2.2f'%(num_eval_batches * eval_batch_size, top1.avg))
```
For this quantized model, we see a significantly lower accuracy of just ~62% on these same 300
images. Nevertheless, we did reduce the size of our model down to just under 3.6 MB, almost a 4x
decrease.
In addition, we can significantly improve on the accuracy simply by using a different
quantization configuration. We repeat the same exercise with the recommended configuration for
quantizing for x86 architectures. This configuration does the following:
- Quantizes weights on a per-channel basis
- Uses a histogram observer that collects a histogram of activations and then picks
quantization parameters in an optimal manner.
```
per_channel_quantized_model = load_model(saved_model_dir + float_model_file)
per_channel_quantized_model.eval()
per_channel_quantized_model.fuse_model()
per_channel_quantized_model.qconfig = torch.quantization.get_default_qconfig('fbgemm')
print(per_channel_quantized_model.qconfig)
torch.quantization.prepare(per_channel_quantized_model, inplace=True)
evaluate(per_channel_quantized_model,criterion, data_loader, num_calibration_batches)
torch.quantization.convert(per_channel_quantized_model, inplace=True)
top1, top5 = evaluate(per_channel_quantized_model, criterion, data_loader_test, neval_batches=num_eval_batches)
print('Evaluation accuracy on %d images, %2.2f'%(num_eval_batches * eval_batch_size, top1.avg))
torch.jit.save(torch.jit.script(per_channel_quantized_model), saved_model_dir + scripted_quantized_model_file)
```
Changing just this quantization configuration method resulted in an increase
of the accuracy to over 76%! Still, this is 1-2% worse than the baseline of 78% achieved above.
So lets try quantization aware training.
5. Quantization-aware training
------------------------------
Quantization-aware training (QAT) is the quantization method that typically results in the highest accuracy.
With QAT, all weights and activations are “fake quantized” during both the forward and backward passes of
training: that is, float values are rounded to mimic int8 values, but all computations are still done with
floating point numbers. Thus, all the weight adjustments during training are made while “aware” of the fact
that the model will ultimately be quantized; after quantizing, therefore, this method will usually yield
higher accuracy than either dynamic quantization or post-training static quantization.
The overall workflow for actually performing QAT is very similar to before:
- We can use the same model as before: there is no additional preparation needed for quantization-aware
training.
- We need to use a ``qconfig`` specifying what kind of fake-quantization is to be inserted after weights
and activations, instead of specifying observers
We first define a training function:
```
def train_one_epoch(model, criterion, optimizer, data_loader, device, ntrain_batches):
model.train()
top1 = AverageMeter('Acc@1', ':6.2f')
top5 = AverageMeter('Acc@5', ':6.2f')
avgloss = AverageMeter('Loss', '1.5f')
cnt = 0
for image, target in data_loader:
start_time = time.time()
print('.', end = '')
cnt += 1
image, target = image.to(device), target.to(device)
output = model(image)
loss = criterion(output, target)
optimizer.zero_grad()
loss.backward()
optimizer.step()
acc1, acc5 = accuracy(output, target, topk=(1, 5))
top1.update(acc1[0], image.size(0))
top5.update(acc5[0], image.size(0))
avgloss.update(loss, image.size(0))
if cnt >= ntrain_batches:
print('Loss', avgloss.avg)
print('Training: * Acc@1 {top1.avg:.3f} Acc@5 {top5.avg:.3f}'
.format(top1=top1, top5=top5))
return
print('Full imagenet train set: * Acc@1 {top1.global_avg:.3f} Acc@5 {top5.global_avg:.3f}'
.format(top1=top1, top5=top5))
return
```
We fuse modules as before
```
qat_model = load_model(saved_model_dir + float_model_file)
qat_model.fuse_model()
optimizer = torch.optim.SGD(qat_model.parameters(), lr = 0.0001)
qat_model.qconfig = torch.quantization.get_default_qat_qconfig('fbgemm')
```
Finally, ``prepare_qat`` performs the "fake quantization", preparing the model for quantization-aware
training
```
torch.quantization.prepare_qat(qat_model, inplace=True)
print('Inverted Residual Block: After preparation for QAT, note fake-quantization modules \n',qat_model.features[1].conv)
```
Training a quantized model with high accuracy requires accurate modeling of numerics at
inference. For quantization aware training, therefore, we modify the training loop by:
- Switch batch norm to use running mean and variance towards the end of training to better
match inference numerics.
- We also freeze the quantizer parameters (scale and zero-point) and fine tune the weights.
```
num_train_batches = 20
# Train and check accuracy after each epoch
for nepoch in range(8):
train_one_epoch(qat_model, criterion, optimizer, data_loader, torch.device('cpu'), num_train_batches)
if nepoch > 3:
# Freeze quantizer parameters
qat_model.apply(torch.quantization.disable_observer)
if nepoch > 2:
# Freeze batch norm mean and variance estimates
qat_model.apply(torch.nn.intrinsic.qat.freeze_bn_stats)
# Check the accuracy after each epoch
quantized_model = torch.quantization.convert(qat_model.eval(), inplace=False)
quantized_model.eval()
top1, top5 = evaluate(quantized_model,criterion, data_loader_test, neval_batches=num_eval_batches)
print('Epoch %d :Evaluation accuracy on %d images, %2.2f'%(nepoch, num_eval_batches * eval_batch_size, top1.avg))
```
Here, we just perform quantization-aware training for a small number of epochs. Nevertheless,
quantization-aware training yields an accuracy of over 71% on the entire imagenet dataset,
which is close to the floating point accuracy of 71.9%.
More on quantization-aware training:
- QAT is a super-set of post training quant techniques that allows for more debugging.
For example, we can analyze if the accuracy of the model is limited by weight or activation
quantization.
- We can also simulate the accuracy of a quantized model in floating point since
we are using fake-quantization to model the numerics of actual quantized arithmetic.
- We can mimic post training quantization easily too.
Speedup from quantization
^^^^^^^^^^^^^^^^^^^^^^^^^
Finally, let's confirm something we alluded to above: do our quantized models actually perform inference
faster? Let's test:
```
def run_benchmark(model_file, img_loader):
elapsed = 0
model = torch.jit.load(model_file)
model.eval()
num_batches = 5
# Run the scripted model on a few batches of images
for i, (images, target) in enumerate(img_loader):
if i < num_batches:
start = time.time()
output = model(images)
end = time.time()
elapsed = elapsed + (end-start)
else:
break
num_images = images.size()[0] * num_batches
print('Elapsed time: %3.0f ms' % (elapsed/num_images*1000))
return elapsed
run_benchmark(saved_model_dir + scripted_float_model_file, data_loader_test)
run_benchmark(saved_model_dir + scripted_quantized_model_file, data_loader_test)
```
Running this locally on a MacBook pro yielded 61 ms for the regular model, and
just 20 ms for the quantized model, illustrating the typical 2-4x speedup
we see for quantized models compared to floating point ones.
Conclusion
----------
In this tutorial, we showed two quantization methods - post-training static quantization,
and quantization-aware training - describing what they do "under the hood" and how to use
them in PyTorch.
Thanks for reading! As always, we welcome any feedback, so please create an issue
`here <https://github.com/pytorch/pytorch/issues>`_ if you have any.
| github_jupyter |
##### Copyright 2018 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Ragged tensors
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/guide/ragged_tensor"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/guide/ragged_tensor.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/guide/ragged_tensor.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/guide/ragged_tensor.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table>
## Setup
```
!pip install -q tf_nightly
import math
import tensorflow as tf
```
## Overview
Your data comes in many shapes; your tensors should too.
*Ragged tensors* are the TensorFlow equivalent of nested variable-length
lists. They make it easy to store and process data with non-uniform shapes,
including:
* Variable-length features, such as the set of actors in a movie.
* Batches of variable-length sequential inputs, such as sentences or video
clips.
* Hierarchical inputs, such as text documents that are subdivided into
sections, paragraphs, sentences, and words.
* Individual fields in structured inputs, such as protocol buffers.
### What you can do with a ragged tensor
Ragged tensors are supported by more than a hundred TensorFlow operations,
including math operations (such as `tf.add` and `tf.reduce_mean`), array operations
(such as `tf.concat` and `tf.tile`), string manipulation ops (such as
`tf.substr`), control flow operations (such as `tf.while_loop` and `tf.map_fn`), and many others:
```
digits = tf.ragged.constant([[3, 1, 4, 1], [], [5, 9, 2], [6], []])
words = tf.ragged.constant([["So", "long"], ["thanks", "for", "all", "the", "fish"]])
print(tf.add(digits, 3))
print(tf.reduce_mean(digits, axis=1))
print(tf.concat([digits, [[5, 3]]], axis=0))
print(tf.tile(digits, [1, 2]))
print(tf.strings.substr(words, 0, 2))
print(tf.map_fn(tf.math.square, digits))
```
There are also a number of methods and operations that are
specific to ragged tensors, including factory methods, conversion methods,
and value-mapping operations.
For a list of supported ops, see the **`tf.ragged` package
documentation**.
Ragged tensors are supported by many TensorFlow APIs, including [Keras](https://www.tensorflow.org/guide/keras), [Datasets](https://www.tensorflow.org/guide/data), [tf.function](https://www.tensorflow.org/guide/function), [SavedModels](https://www.tensorflow.org/guide/saved_model), and [tf.Example](https://www.tensorflow.org/tutorials/load_data/tfrecord). For more information, see the section on **TensorFlow APIs** below.
As with normal tensors, you can use Python-style indexing to access specific
slices of a ragged tensor. For more information, see the section on
**Indexing** below.
```
print(digits[0]) # First row
print(digits[:, :2]) # First two values in each row.
print(digits[:, -2:]) # Last two values in each row.
```
And just like normal tensors, you can use Python arithmetic and comparison
operators to perform elementwise operations. For more information, see the section on
**Overloaded Operators** below.
```
print(digits + 3)
print(digits + tf.ragged.constant([[1, 2, 3, 4], [], [5, 6, 7], [8], []]))
```
If you need to perform an elementwise transformation to the values of a `RaggedTensor`, you can use `tf.ragged.map_flat_values`, which takes a function plus one or more arguments, and applies the function to transform the `RaggedTensor`'s values.
```
times_two_plus_one = lambda x: x * 2 + 1
print(tf.ragged.map_flat_values(times_two_plus_one, digits))
```
Ragged tensors can be converted to nested Python `list`s and numpy `array`s:
```
digits.to_list()
digits.numpy()
```
### Constructing a ragged tensor
The simplest way to construct a ragged tensor is using
`tf.ragged.constant`, which builds the
`RaggedTensor` corresponding to a given nested Python `list` or numpy `array`:
```
sentences = tf.ragged.constant([
["Let's", "build", "some", "ragged", "tensors", "!"],
["We", "can", "use", "tf.ragged.constant", "."]])
print(sentences)
paragraphs = tf.ragged.constant([
[['I', 'have', 'a', 'cat'], ['His', 'name', 'is', 'Mat']],
[['Do', 'you', 'want', 'to', 'come', 'visit'], ["I'm", 'free', 'tomorrow']],
])
print(paragraphs)
```
Ragged tensors can also be constructed by pairing flat *values* tensors with
*row-partitioning* tensors indicating how those values should be divided into
rows, using factory classmethods such as `tf.RaggedTensor.from_value_rowids`,
`tf.RaggedTensor.from_row_lengths`, and
`tf.RaggedTensor.from_row_splits`.
#### `tf.RaggedTensor.from_value_rowids`
If you know which row each value belongs in, then you can build a `RaggedTensor` using a `value_rowids` row-partitioning tensor:
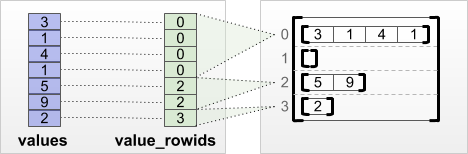
```
print(tf.RaggedTensor.from_value_rowids(
values=[3, 1, 4, 1, 5, 9, 2],
value_rowids=[0, 0, 0, 0, 2, 2, 3]))
```
#### `tf.RaggedTensor.from_row_lengths`
If you know how long each row is, then you can use a `row_lengths` row-partitioning tensor:
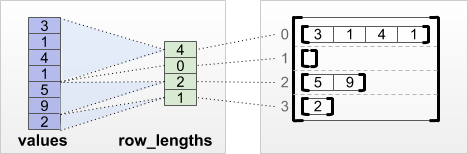
```
print(tf.RaggedTensor.from_row_lengths(
values=[3, 1, 4, 1, 5, 9, 2],
row_lengths=[4, 0, 2, 1]))
```
#### `tf.RaggedTensor.from_row_splits`
If you know the index where each row starts and ends, then you can use a `row_splits` row-partitioning tensor:
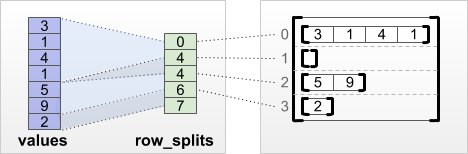
```
print(tf.RaggedTensor.from_row_splits(
values=[3, 1, 4, 1, 5, 9, 2],
row_splits=[0, 4, 4, 6, 7]))
```
See the `tf.RaggedTensor` class documentation for a full list of factory methods.
Note: By default, these factory methods add assertions that the row partition tensor is well-formed and consistent with the number of values. The `validate=False` parameter can be used to skip these checks if you can guarantee that the inputs are well-formed and consistent.
### What you can store in a ragged tensor
As with normal `Tensor`s, the values in a `RaggedTensor` must all have the same
type; and the values must all be at the same nesting depth (the *rank* of the
tensor):
```
print(tf.ragged.constant([["Hi"], ["How", "are", "you"]])) # ok: type=string, rank=2
print(tf.ragged.constant([[[1, 2], [3]], [[4, 5]]])) # ok: type=int32, rank=3
try:
tf.ragged.constant([["one", "two"], [3, 4]]) # bad: multiple types
except ValueError as exception:
print(exception)
try:
tf.ragged.constant(["A", ["B", "C"]]) # bad: multiple nesting depths
except ValueError as exception:
print(exception)
```
## Example use case
The following example demonstrates how `RaggedTensor`s can be used to construct
and combine unigram and bigram embeddings for a batch of variable-length
queries, using special markers for the beginning and end of each sentence.
For more details on the ops used in this example, see the `tf.ragged` package documentation.
```
queries = tf.ragged.constant([['Who', 'is', 'Dan', 'Smith'],
['Pause'],
['Will', 'it', 'rain', 'later', 'today']])
# Create an embedding table.
num_buckets = 1024
embedding_size = 4
embedding_table = tf.Variable(
tf.random.truncated_normal([num_buckets, embedding_size],
stddev=1.0 / math.sqrt(embedding_size)))
# Look up the embedding for each word.
word_buckets = tf.strings.to_hash_bucket_fast(queries, num_buckets)
word_embeddings = tf.nn.embedding_lookup(embedding_table, word_buckets) # ①
# Add markers to the beginning and end of each sentence.
marker = tf.fill([queries.nrows(), 1], '#')
padded = tf.concat([marker, queries, marker], axis=1) # ②
# Build word bigrams & look up embeddings.
bigrams = tf.strings.join([padded[:, :-1], padded[:, 1:]], separator='+') # ③
bigram_buckets = tf.strings.to_hash_bucket_fast(bigrams, num_buckets)
bigram_embeddings = tf.nn.embedding_lookup(embedding_table, bigram_buckets) # ④
# Find the average embedding for each sentence
all_embeddings = tf.concat([word_embeddings, bigram_embeddings], axis=1) # ⑤
avg_embedding = tf.reduce_mean(all_embeddings, axis=1) # ⑥
print(avg_embedding)
```
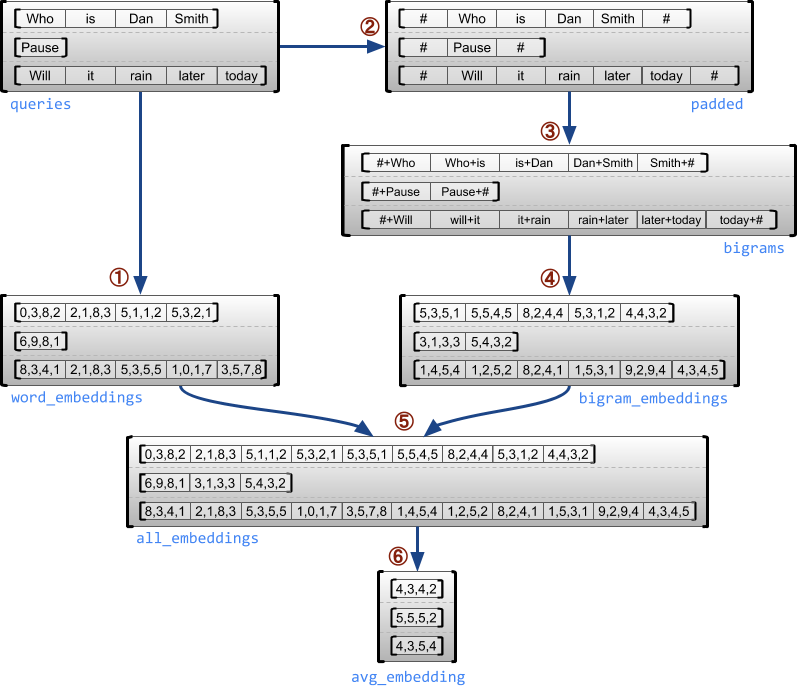
## Ragged and uniform dimensions
A ***ragged dimension*** is a dimension whose slices may have different lengths. For example, the
inner (column) dimension of `rt=[[3, 1, 4, 1], [], [5, 9, 2], [6], []]` is
ragged, since the column slices (`rt[0, :]`, ..., `rt[4, :]`) have different
lengths. Dimensions whose slices all have the same length are called *uniform
dimensions*.
The outermost dimension of a ragged tensor is always uniform, since it consists
of a single slice (and so there is no possibility for differing slice
lengths). The remaining dimensions may be either ragged or uniform. For
example, we might store the word embeddings for
each word in a batch of sentences using a ragged tensor with shape
`[num_sentences, (num_words), embedding_size]`, where the parentheses around
`(num_words)` indicate that the dimension is ragged.
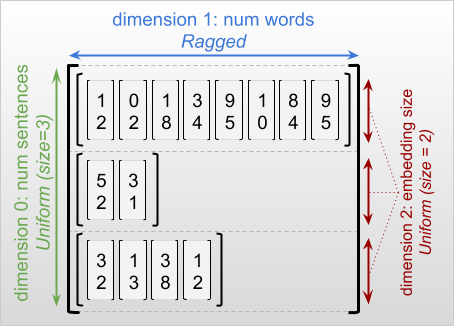
Ragged tensors may have multiple ragged dimensions. For example, we could store
a batch of structured text documents using a tensor with shape `[num_documents,
(num_paragraphs), (num_sentences), (num_words)]` (where again parentheses are
used to indicate ragged dimensions).
As with `tf.Tensor`, the ***rank*** of a ragged tensor is its total number of dimensions (including both ragged and uniform dimensions).
A ***potentially ragged tensor*** is a value that might be
either a `tf.Tensor` or a `tf.RaggedTensor`.
When describing the shape of a RaggedTensor, ragged dimensions are conventionally indicated by
enclosing them in parentheses. For example, as we saw above, the shape of a 3-D
RaggedTensor that stores word embeddings for each word in a batch of sentences
can be written as `[num_sentences, (num_words), embedding_size]`.
The `RaggedTensor.shape` attribute returns a `tf.TensorShape` for a
ragged tensor, where ragged dimensions have size `None`:
```
tf.ragged.constant([["Hi"], ["How", "are", "you"]]).shape
```
The method `tf.RaggedTensor.bounding_shape` can be used to find a tight
bounding shape for a given `RaggedTensor`:
```
print(tf.ragged.constant([["Hi"], ["How", "are", "you"]]).bounding_shape())
```
## Ragged vs. sparse
A ragged tensor should *not* be thought of as a type of sparse tensor. In particular, sparse tensors are *efficient encodings for tf.Tensor*, that model the same data in a compact format; but ragged tensor is an *extension to tf.Tensor*, that models an expanded class of data. This difference is crucial when defining operations:
* Applying an op to a sparse or dense tensor should always give the same result.
* Applying an op to a ragged or sparse tensor may give different results.
As an illustrative example, consider how array operations such as `concat`,
`stack`, and `tile` are defined for ragged vs. sparse tensors. Concatenating
ragged tensors joins each row to form a single row with the combined length:

```
ragged_x = tf.ragged.constant([["John"], ["a", "big", "dog"], ["my", "cat"]])
ragged_y = tf.ragged.constant([["fell", "asleep"], ["barked"], ["is", "fuzzy"]])
print(tf.concat([ragged_x, ragged_y], axis=1))
```
But concatenating sparse tensors is equivalent to concatenating the corresponding dense tensors,
as illustrated by the following example (where Ø indicates missing values):
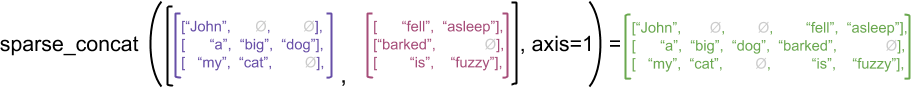
```
sparse_x = ragged_x.to_sparse()
sparse_y = ragged_y.to_sparse()
sparse_result = tf.sparse.concat(sp_inputs=[sparse_x, sparse_y], axis=1)
print(tf.sparse.to_dense(sparse_result, ''))
```
For another example of why this distinction is important, consider the
definition of “the mean value of each row” for an op such as `tf.reduce_mean`.
For a ragged tensor, the mean value for a row is the sum of the
row’s values divided by the row’s width.
But for a sparse tensor, the mean value for a row is the sum of the
row’s values divided by the sparse tensor’s overall width (which is
greater than or equal to the width of the longest row).
## TensorFlow APIs
### Keras
[tf.keras](https://www.tensorflow.org/guide/keras) is TensorFlow's high-level API for building and training deep learning models. Ragged tensors may be passed as inputs to a Keras model by setting `ragged=True` on `tf.keras.Input` or `tf.keras.layers.InputLayer`. Ragged tensors may also be passed between Keras layers, and returned by Keras models. The following example shows a toy LSTM model that is trained using ragged tensors.
```
# Task: predict whether each sentence is a question or not.
sentences = tf.constant(
['What makes you think she is a witch?',
'She turned me into a newt.',
'A newt?',
'Well, I got better.'])
is_question = tf.constant([True, False, True, False])
# Preprocess the input strings.
hash_buckets = 1000
words = tf.strings.split(sentences, ' ')
hashed_words = tf.strings.to_hash_bucket_fast(words, hash_buckets)
# Build the Keras model.
keras_model = tf.keras.Sequential([
tf.keras.layers.Input(shape=[None], dtype=tf.int64, ragged=True),
tf.keras.layers.Embedding(hash_buckets, 16),
tf.keras.layers.LSTM(32, use_bias=False),
tf.keras.layers.Dense(32),
tf.keras.layers.Activation(tf.nn.relu),
tf.keras.layers.Dense(1)
])
keras_model.compile(loss='binary_crossentropy', optimizer='rmsprop')
keras_model.fit(hashed_words, is_question, epochs=5)
print(keras_model.predict(hashed_words))
```
### tf.Example
[tf.Example](https://www.tensorflow.org/tutorials/load_data/tfrecord) is a standard [protobuf](https://developers.google.com/protocol-buffers/) encoding for TensorFlow data. Data encoded with `tf.Example`s often includes variable-length features. For example, the following code defines a batch of four `tf.Example` messages with different feature lengths:
```
import google.protobuf.text_format as pbtext
def build_tf_example(s):
return pbtext.Merge(s, tf.train.Example()).SerializeToString()
example_batch = [
build_tf_example(r'''
features {
feature {key: "colors" value {bytes_list {value: ["red", "blue"]} } }
feature {key: "lengths" value {int64_list {value: [7]} } } }'''),
build_tf_example(r'''
features {
feature {key: "colors" value {bytes_list {value: ["orange"]} } }
feature {key: "lengths" value {int64_list {value: []} } } }'''),
build_tf_example(r'''
features {
feature {key: "colors" value {bytes_list {value: ["black", "yellow"]} } }
feature {key: "lengths" value {int64_list {value: [1, 3]} } } }'''),
build_tf_example(r'''
features {
feature {key: "colors" value {bytes_list {value: ["green"]} } }
feature {key: "lengths" value {int64_list {value: [3, 5, 2]} } } }''')]
```
We can parse this encoded data using `tf.io.parse_example`, which takes a tensor of serialized strings and a feature specification dictionary, and returns a dictionary mapping feature names to tensors. To read the variable-length features into ragged tensors, we simply use `tf.io.RaggedFeature` in the feature specification dictionary:
```
feature_specification = {
'colors': tf.io.RaggedFeature(tf.string),
'lengths': tf.io.RaggedFeature(tf.int64),
}
feature_tensors = tf.io.parse_example(example_batch, feature_specification)
for name, value in feature_tensors.items():
print("{}={}".format(name, value))
```
`tf.io.RaggedFeature` can also be used to read features with multiple ragged dimensions. For details, see the [API documentation](https://www.tensorflow.org/api_docs/python/tf/io/RaggedFeature).
### Datasets
[tf.data](https://www.tensorflow.org/guide/data) is an API that enables you to build complex input pipelines from simple, reusable pieces. Its core data structure is `tf.data.Dataset`, which represents a sequence of elements, in which each element consists of one or more components.
```
# Helper function used to print datasets in the examples below.
def print_dictionary_dataset(dataset):
for i, element in enumerate(dataset):
print("Element {}:".format(i))
for (feature_name, feature_value) in element.items():
print('{:>14} = {}'.format(feature_name, feature_value))
```
#### Building Datasets with ragged tensors
Datasets can be built from ragged tensors using the same methods that are used to build them from `tf.Tensor`s or numpy `array`s, such as `Dataset.from_tensor_slices`:
```
dataset = tf.data.Dataset.from_tensor_slices(feature_tensors)
print_dictionary_dataset(dataset)
```
Note: `Dataset.from_generator` does not support ragged tensors yet, but support will be added soon.
#### Batching and unbatching Datasets with ragged tensors
Datasets with ragged tensors can be batched (which combines *n* consecutive elements into a single elements) using the `Dataset.batch` method.
```
batched_dataset = dataset.batch(2)
print_dictionary_dataset(batched_dataset)
```
Conversely, a batched dataset can be transformed into a flat dataset using `Dataset.unbatch`.
```
unbatched_dataset = batched_dataset.unbatch()
print_dictionary_dataset(unbatched_dataset)
```
#### Batching Datasets with variable-length non-ragged tensors
If you have a Dataset that contains non-ragged tensors, and tensor lengths vary across elements, then you can batch those non-ragged tensors into ragged tensors by applying the `dense_to_ragged_batch` transformation:
```
non_ragged_dataset = tf.data.Dataset.from_tensor_slices([1, 5, 3, 2, 8])
non_ragged_dataset = non_ragged_dataset.map(tf.range)
batched_non_ragged_dataset = non_ragged_dataset.apply(
tf.data.experimental.dense_to_ragged_batch(2))
for element in batched_non_ragged_dataset:
print(element)
```
#### Transforming Datasets with ragged tensors
Ragged tensors in Datasets can also be created or transformed using `Dataset.map`.
```
def transform_lengths(features):
return {
'mean_length': tf.math.reduce_mean(features['lengths']),
'length_ranges': tf.ragged.range(features['lengths'])}
transformed_dataset = dataset.map(transform_lengths)
print_dictionary_dataset(transformed_dataset)
```
### tf.function
[tf.function](https://www.tensorflow.org/guide/function) is a decorator that precomputes TensorFlow graphs for Python functions, which can substantially improve the performance of your TensorFlow code. Ragged tensors can be used transparently with `@tf.function`-decorated functions. For example, the following function works with both ragged and non-ragged tensors:
```
@tf.function
def make_palindrome(x, axis):
return tf.concat([x, tf.reverse(x, [axis])], axis)
make_palindrome(tf.constant([[1, 2], [3, 4], [5, 6]]), axis=1)
make_palindrome(tf.ragged.constant([[1, 2], [3], [4, 5, 6]]), axis=1)
```
If you wish to explicitly specify the `input_signature` for the `tf.function`, then you can do so using `tf.RaggedTensorSpec`.
```
@tf.function(
input_signature=[tf.RaggedTensorSpec(shape=[None, None], dtype=tf.int32)])
def max_and_min(rt):
return (tf.math.reduce_max(rt, axis=-1), tf.math.reduce_min(rt, axis=-1))
max_and_min(tf.ragged.constant([[1, 2], [3], [4, 5, 6]]))
```
#### Concrete functions
[Concrete functions](https://www.tensorflow.org/guide/concrete_function) encapsulate individual traced graphs that are built by `tf.function`. Starting with TensorFlow 2.3 (and in `tf-nightly`), ragged tensors can be used transparently with concrete functions.
```
# Preferred way to use ragged tensors with concrete functions (TF 2.3+):
try:
@tf.function
def increment(x):
return x + 1
rt = tf.ragged.constant([[1, 2], [3], [4, 5, 6]])
cf = increment.get_concrete_function(rt)
print(cf(rt))
except Exception as e:
print(f"Not supported before TF 2.3: {type(e)}: {e}")
```
If you need to use ragged tensors with concrete functions prior to TensorFlow 2.3, then we recommend decomposing ragged tensors into their components (`values` and `row_splits`), and passing them in as separate arguments.
```
# Backwards-compatible way to use ragged tensors with concrete functions:
@tf.function
def decomposed_ragged_increment(x_values, x_splits):
x = tf.RaggedTensor.from_row_splits(x_values, x_splits)
return x + 1
rt = tf.ragged.constant([[1, 2], [3], [4, 5, 6]])
cf = decomposed_ragged_increment.get_concrete_function(rt.values, rt.row_splits)
print(cf(rt.values, rt.row_splits))
```
### SavedModels
A [SavedModel](https://www.tensorflow.org/guide/saved_model) is a serialized TensorFlow program, including both weights and computation. It can be built from a Keras model or from a custom model. In either case, ragged tensors can be used transparently with the functions and methods defined by a SavedModel.
#### Example: saving a Keras model
```
import tempfile
keras_module_path = tempfile.mkdtemp()
tf.saved_model.save(keras_model, keras_module_path)
imported_model = tf.saved_model.load(keras_module_path)
imported_model(hashed_words)
```
#### Example: saving a custom model
```
class CustomModule(tf.Module):
def __init__(self, variable_value):
super(CustomModule, self).__init__()
self.v = tf.Variable(variable_value)
@tf.function
def grow(self, x):
return x * self.v
module = CustomModule(100.0)
# Before saving a custom model, we must ensure that concrete functions are
# built for each input signature that we will need.
module.grow.get_concrete_function(tf.RaggedTensorSpec(shape=[None, None],
dtype=tf.float32))
custom_module_path = tempfile.mkdtemp()
tf.saved_model.save(module, custom_module_path)
imported_model = tf.saved_model.load(custom_module_path)
imported_model.grow(tf.ragged.constant([[1.0, 4.0, 3.0], [2.0]]))
```
Note: SavedModel [signatures](https://www.tensorflow.org/guide/saved_model#specifying_signatures_during_export) are concrete functions. As discussed in the section on Concrete Functions above, ragged tensors are only handled correctly by concrete functions starting with TensorFlow 2.3 (and in `tf_nightly`). If you need to use SavedModel signatures in a previous version of TensorFlow, then we recommend decomposing the ragged tensor into its component tensors.
## Overloaded operators
The `RaggedTensor` class overloads the standard Python arithmetic and comparison
operators, making it easy to perform basic elementwise math:
```
x = tf.ragged.constant([[1, 2], [3], [4, 5, 6]])
y = tf.ragged.constant([[1, 1], [2], [3, 3, 3]])
print(x + y)
```
Since the overloaded operators perform elementwise computations, the inputs to
all binary operations must have the same shape, or be broadcastable to the same
shape. In the simplest broadcasting case, a single scalar is combined
elementwise with each value in a ragged tensor:
```
x = tf.ragged.constant([[1, 2], [3], [4, 5, 6]])
print(x + 3)
```
For a discussion of more advanced cases, see the section on
**Broadcasting**.
Ragged tensors overload the same set of operators as normal `Tensor`s: the unary
operators `-`, `~`, and `abs()`; and the binary operators `+`, `-`, `*`, `/`,
`//`, `%`, `**`, `&`, `|`, `^`, `==`, `<`, `<=`, `>`, and `>=`.
## Indexing
Ragged tensors support Python-style indexing, including multidimensional
indexing and slicing. The following examples demonstrate ragged tensor indexing
with a 2-D and a 3-D ragged tensor.
### Indexing examples: 2D ragged tensor
```
queries = tf.ragged.constant(
[['Who', 'is', 'George', 'Washington'],
['What', 'is', 'the', 'weather', 'tomorrow'],
['Goodnight']])
print(queries[1]) # A single query
print(queries[1, 2]) # A single word
print(queries[1:]) # Everything but the first row
print(queries[:, :3]) # The first 3 words of each query
print(queries[:, -2:]) # The last 2 words of each query
```
### Indexing examples 3D ragged tensor
```
rt = tf.ragged.constant([[[1, 2, 3], [4]],
[[5], [], [6]],
[[7]],
[[8, 9], [10]]])
print(rt[1]) # Second row (2-D RaggedTensor)
print(rt[3, 0]) # First element of fourth row (1-D Tensor)
print(rt[:, 1:3]) # Items 1-3 of each row (3-D RaggedTensor)
print(rt[:, -1:]) # Last item of each row (3-D RaggedTensor)
```
`RaggedTensor`s supports multidimensional indexing and slicing, with one
restriction: indexing into a ragged dimension is not allowed. This case is
problematic because the indicated value may exist in some rows but not others.
In such cases, it's not obvious whether we should (1) raise an `IndexError`; (2)
use a default value; or (3) skip that value and return a tensor with fewer rows
than we started with. Following the
[guiding principles of Python](https://www.python.org/dev/peps/pep-0020/)
("In the face
of ambiguity, refuse the temptation to guess" ), we currently disallow this
operation.
## Tensor type conversion
The `RaggedTensor` class defines methods that can be used to convert
between `RaggedTensor`s and `tf.Tensor`s or `tf.SparseTensors`:
```
ragged_sentences = tf.ragged.constant([
['Hi'], ['Welcome', 'to', 'the', 'fair'], ['Have', 'fun']])
# RaggedTensor -> Tensor
print(ragged_sentences.to_tensor(default_value='', shape=[None, 10]))
# Tensor -> RaggedTensor
x = [[1, 3, -1, -1], [2, -1, -1, -1], [4, 5, 8, 9]]
print(tf.RaggedTensor.from_tensor(x, padding=-1))
#RaggedTensor -> SparseTensor
print(ragged_sentences.to_sparse())
# SparseTensor -> RaggedTensor
st = tf.SparseTensor(indices=[[0, 0], [2, 0], [2, 1]],
values=['a', 'b', 'c'],
dense_shape=[3, 3])
print(tf.RaggedTensor.from_sparse(st))
```
## Evaluating ragged tensors
To access the values in a ragged tensor, you can:
1. Use `tf.RaggedTensor.to_list()` to convert the ragged tensor to a
nested python list.
1. Use `tf.RaggedTensor.numpy()` to convert the ragged tensor to a numpy array
whose values are nested numpy arrays.
1. Decompose the ragged tensor into its components, using the
`tf.RaggedTensor.values` and `tf.RaggedTensor.row_splits`
properties, or row-paritioning methods such as
`tf.RaggedTensor.row_lengths()` and `tf.RaggedTensor.value_rowids()`.
1. Use Python indexing to select values from the ragged tensor.
```
rt = tf.ragged.constant([[1, 2], [3, 4, 5], [6], [], [7]])
print("python list:", rt.to_list())
print("numpy array:", rt.numpy())
print("values:", rt.values.numpy())
print("splits:", rt.row_splits.numpy())
print("indexed value:", rt[1].numpy())
```
## Broadcasting
Broadcasting is the process of making tensors with different shapes have
compatible shapes for elementwise operations. For more background on
broadcasting, see:
* [Numpy: Broadcasting](https://docs.scipy.org/doc/numpy/user/basics.broadcasting.html)
* `tf.broadcast_dynamic_shape`
* `tf.broadcast_to`
The basic steps for broadcasting two inputs `x` and `y` to have compatible
shapes are:
1. If `x` and `y` do not have the same number of dimensions, then add outer
dimensions (with size 1) until they do.
2. For each dimension where `x` and `y` have different sizes:
* If `x` or `y` have size `1` in dimension `d`, then repeat its values
across dimension `d` to match the other input's size.
* Otherwise, raise an exception (`x` and `y` are not broadcast
compatible).
Where the size of a tensor in a uniform dimension is a single number (the size
of slices across that dimension); and the size of a tensor in a ragged dimension
is a list of slice lengths (for all slices across that dimension).
### Broadcasting examples
```
# x (2D ragged): 2 x (num_rows)
# y (scalar)
# result (2D ragged): 2 x (num_rows)
x = tf.ragged.constant([[1, 2], [3]])
y = 3
print(x + y)
# x (2d ragged): 3 x (num_rows)
# y (2d tensor): 3 x 1
# Result (2d ragged): 3 x (num_rows)
x = tf.ragged.constant(
[[10, 87, 12],
[19, 53],
[12, 32]])
y = [[1000], [2000], [3000]]
print(x + y)
# x (3d ragged): 2 x (r1) x 2
# y (2d ragged): 1 x 1
# Result (3d ragged): 2 x (r1) x 2
x = tf.ragged.constant(
[[[1, 2], [3, 4], [5, 6]],
[[7, 8]]],
ragged_rank=1)
y = tf.constant([[10]])
print(x + y)
# x (3d ragged): 2 x (r1) x (r2) x 1
# y (1d tensor): 3
# Result (3d ragged): 2 x (r1) x (r2) x 3
x = tf.ragged.constant(
[
[
[[1], [2]],
[],
[[3]],
[[4]],
],
[
[[5], [6]],
[[7]]
]
],
ragged_rank=2)
y = tf.constant([10, 20, 30])
print(x + y)
```
Here are some examples of shapes that do not broadcast:
```
# x (2d ragged): 3 x (r1)
# y (2d tensor): 3 x 4 # trailing dimensions do not match
x = tf.ragged.constant([[1, 2], [3, 4, 5, 6], [7]])
y = tf.constant([[1, 2, 3, 4], [5, 6, 7, 8], [9, 10, 11, 12]])
try:
x + y
except tf.errors.InvalidArgumentError as exception:
print(exception)
# x (2d ragged): 3 x (r1)
# y (2d ragged): 3 x (r2) # ragged dimensions do not match.
x = tf.ragged.constant([[1, 2, 3], [4], [5, 6]])
y = tf.ragged.constant([[10, 20], [30, 40], [50]])
try:
x + y
except tf.errors.InvalidArgumentError as exception:
print(exception)
# x (3d ragged): 3 x (r1) x 2
# y (3d ragged): 3 x (r1) x 3 # trailing dimensions do not match
x = tf.ragged.constant([[[1, 2], [3, 4], [5, 6]],
[[7, 8], [9, 10]]])
y = tf.ragged.constant([[[1, 2, 0], [3, 4, 0], [5, 6, 0]],
[[7, 8, 0], [9, 10, 0]]])
try:
x + y
except tf.errors.InvalidArgumentError as exception:
print(exception)
```
## RaggedTensor encoding
Ragged tensors are encoded using the `RaggedTensor` class. Internally, each
`RaggedTensor` consists of:
* A `values` tensor, which concatenates the variable-length rows into a
flattened list.
* A `row_partition`, which indicates how those flattened values are divided
into rows.
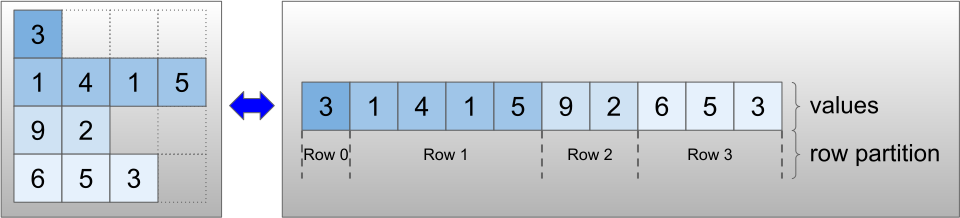
The `row_partition` can be stored using four different encodings:
* `row_splits` is an integer vector specifying the split points between rows.
* `value_rowids` is an integer vector specifying the row index for each value.
* `row_lengths` is an integer vector specifying the length of each row.
* `uniform_row_length` is an integer scalar specifying a single length for
all rows.
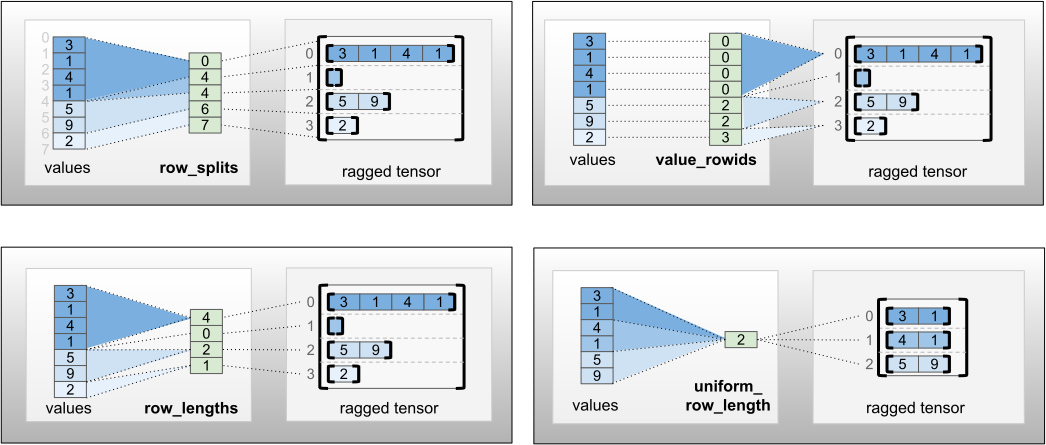
An integer scalar `nrows` can also be included in the `row_partition` encoding, to account for empty trailing rows with `value_rowids`, or empty rows with `uniform_row_length`.
```
rt = tf.RaggedTensor.from_row_splits(
values=[3, 1, 4, 1, 5, 9, 2],
row_splits=[0, 4, 4, 6, 7])
print(rt)
```
The choice of which encoding to use for row partitions is managed internally by ragged tensors, to improve efficiency in some contexts. In particular, some of the advantages and disadvantages of the different row-partitioning
schemes are:
+ **Efficient indexing**:
The `row_splits` encoding enables
constant-time indexing and slicing into ragged tensors.
+ **Efficient concatenation**:
The `row_lengths` encoding is more efficient when concatenating ragged
tensors, since row lengths do not change when two tensors are concatenated
together.
+ **Small encoding size**:
The `value_rowids` encoding is more efficient when storing ragged tensors
that have a large number of empty rows, since the size of the tensor
depends only on the total number of values. On the other hand, the
`row_splits` and `row_lengths` encodings
are more efficient when storing ragged tensors with longer rows, since they
require only one scalar value for each row.
+ **Compatibility**:
The `value_rowids` scheme matches the
[segmentation](https://www.tensorflow.org/api_docs/python/tf/math#about_segmentation)
format used by operations such as `tf.segment_sum`. The `row_limits` scheme
matches the format used by ops such as `tf.sequence_mask`.
+ **Uniform dimensions**:
As discussed below, the `uniform_row_length` encoding is used to encode
ragged tensors with uniform dimensions.
### Multiple ragged dimensions
A ragged tensor with multiple ragged dimensions is encoded by using a nested
`RaggedTensor` for the `values` tensor. Each nested `RaggedTensor` adds a single
ragged dimension.
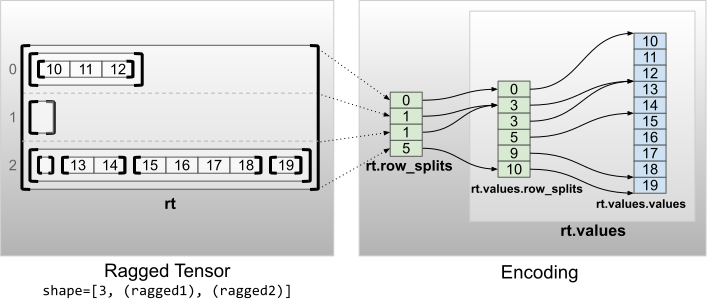
```
rt = tf.RaggedTensor.from_row_splits(
values=tf.RaggedTensor.from_row_splits(
values=[10, 11, 12, 13, 14, 15, 16, 17, 18, 19],
row_splits=[0, 3, 3, 5, 9, 10]),
row_splits=[0, 1, 1, 5])
print(rt)
print("Shape: {}".format(rt.shape))
print("Number of partitioned dimensions: {}".format(rt.ragged_rank))
```
The factory function `tf.RaggedTensor.from_nested_row_splits` may be used to construct a
RaggedTensor with multiple ragged dimensions directly, by providing a list of
`row_splits` tensors:
```
rt = tf.RaggedTensor.from_nested_row_splits(
flat_values=[10, 11, 12, 13, 14, 15, 16, 17, 18, 19],
nested_row_splits=([0, 1, 1, 5], [0, 3, 3, 5, 9, 10]))
print(rt)
```
### Ragged rank and flat values
A ragged tensor's ***ragged rank*** is the number of times that the underlying
`values` Tensor has been partitioned (i.e., the nesting depth of `RaggedTensor` objects). The innermost `values` tensor is known as its ***flat_values***. In the following example, `conversations` has ragged_rank=3, and its `flat_values` is a 1D `Tensor` with 24 strings:
```
# shape = [batch, (paragraph), (sentence), (word)]
conversations = tf.ragged.constant(
[[[["I", "like", "ragged", "tensors."]],
[["Oh", "yeah?"], ["What", "can", "you", "use", "them", "for?"]],
[["Processing", "variable", "length", "data!"]]],
[[["I", "like", "cheese."], ["Do", "you?"]],
[["Yes."], ["I", "do."]]]])
conversations.shape
assert conversations.ragged_rank == len(conversations.nested_row_splits)
conversations.ragged_rank # Number of partitioned dimensions.
conversations.flat_values.numpy()
```
### Uniform inner dimensions
Ragged tensors with uniform inner dimensions are encoded by using a
multidimensional `tf.Tensor` for the flat_values (i.e., the innermost `values`).
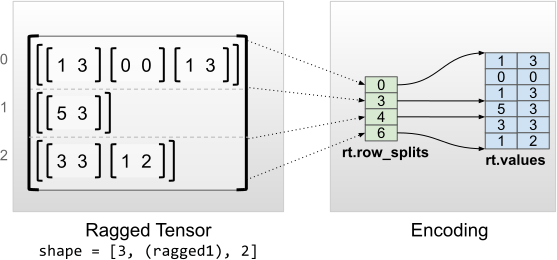
```
rt = tf.RaggedTensor.from_row_splits(
values=[[1, 3], [0, 0], [1, 3], [5, 3], [3, 3], [1, 2]],
row_splits=[0, 3, 4, 6])
print(rt)
print("Shape: {}".format(rt.shape))
print("Number of partitioned dimensions: {}".format(rt.ragged_rank))
print("Flat values shape: {}".format(rt.flat_values.shape))
print("Flat values:\n{}".format(rt.flat_values))
```
### Uniform non-inner dimensions
Ragged tensors with uniform non-inner dimensions are encoded by partitioning rows with `uniform_row_length`.
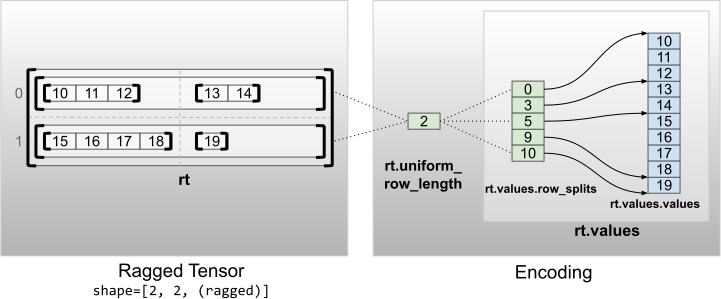
```
rt = tf.RaggedTensor.from_uniform_row_length(
values=tf.RaggedTensor.from_row_splits(
values=[10, 11, 12, 13, 14, 15, 16, 17, 18, 19],
row_splits=[0, 3, 5, 9, 10]),
uniform_row_length=2)
print(rt)
print("Shape: {}".format(rt.shape))
print("Number of partitioned dimensions: {}".format(rt.ragged_rank))
```
| github_jupyter |
```
#Imported relevant and necessary libraries and data cleaning tools
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import hypertools as hyp
from glob import glob as lsdir
import os
import re
import datetime as dt
from sklearn import linear_model
from sklearn.neural_network import MLPRegressor
from sklearn.model_selection import train_test_split
%matplotlib inline
#Code from Professor Manning to set up and read in the relevant UVLT data
data_readers = {'xlsx': pd.read_excel, 'xls': pd.read_excel, 'dta': pd.read_stata}
get_extension = lambda x: x.split('.')[-1]
def read_data(datadir, readers):
files = lsdir(os.path.join(datadir, '*'))
readable_files = []
data = []
for f in files:
ext = get_extension(f)
if ext in readers.keys():
readable_files.append(f)
data.append(data_readers[ext](f))
return readable_files, data
fnames, data = read_data('data', data_readers)
#A summary of the data files that are now read into the notebook
fnames
#This is the individual data
#Most of the initial cleaning that I have done has been on this data set, though obviously it can be applied if necessary to any of the data sets
data[0].head()
#Renaming relevant columns in UVLT individual data to be more easily readable
names={'DeceasedDateYN' : 'Is the donor Deceased?',
'U_Tot_Amt': 'Total Unrestricted Donations',
'U_Tot_Cnt': 'Total Number of Unrestricted Donations Given',
'ConservedOwner' : 'Owns Conserved Land?',
'RTotAmt' : 'Total Restricted Donations',
'RTotCnt': 'Total Number of Restricted Donations Given',
'VTotCnt' : 'Total Volunteer Occurances',
'ETotCnt' : 'Total Event Attendances'}
data[0].rename(names, inplace=True, axis=1)
#Summary of the column values in data set 1
data[0].columns.values
#copying each set of data into more memorably named versions
#I figured different analyses require different aspects of each dataframe, so starting here and using copies of the data for different analyses may be helpful for organizationa and fidelity
Individual_data=data[0].copy()
Final_data=data[1].copy()
Mailing_data=data[2].copy()
Town_Data=data[5].copy()
#Similarly to the individual data, the final data could benefit from some cleaner column names
Final_data.rename(names, inplace=True, axis=1)
to_drop={'U200001',
'U200102',
'U200203',
'U200304',
'U200405',
'U200506',
'U200607',
'U200708',
'U200809',
'U200910',
'U201011',
'U201112',
'U201213',
'U201314',
'U201415',
'U201516',
'U201617',
'U201718',
'U201819',
'R200001',
'R200102',
'R200203',
'R200304',
'R200405',
'R200506',
'R200607',
'R200708',
'R200809',
'R200910',
'R201011',
'R201112',
'R201213',
'R201314',
'R201415',
'R201516',
'R201617',
'R201718',
'R201819',
'V200001',
'V200102',
'V200203',
'V200304',
'V200405',
'V200506',
'V200607',
'V200708',
'V200809',
'V200910',
'V201011',
'V201112',
'V201213',
'V201314',
'V201415',
'V201516',
'V201617',
'V201718',
'V201819',
'E200001',
'E200102',
'E200203',
'E200304',
'E200405',
'E200506',
'E200607',
'E200708',
'E200809',
'E200910',
'E201011',
'E201112',
'E201213',
'E201314',
'E201415',
'E201516',
'E201617',
'E201718',
'E201819'}
Individual_data_SUMMARY=data[0].copy()
Individual_data_SUMMARY.drop(to_drop, inplace=True, axis=1)
#The summary cleaning gets rid of the year-by-year data and leaves only the summary of the donations for analyses
#obviously there are pros and cons to this, but I figured it was a tedious process to undertake so now it's a simple dictionary that can be used on the various data sets
Individual_data_SUMMARY.head(15)
Final_data_SUMMARY=Final_data.copy()
Final_data_SUMMARY.drop(to_drop, inplace=True, axis=1)
Final_data_SUMMARY.head(15)
```
| github_jupyter |
# Ex 12
```
import tensorflow as tf
from tensorflow import keras
import os
import numpy as np
import matplotlib.pyplot as plt
from tensorflow.keras.datasets import mnist
from tensorflow.keras import models
from tensorflow.keras import layers
from tensorflow.keras.utils import to_categorical
from matplotlib.pyplot import figure
from time import time
seed=0
np.random.seed(seed) # fix random seed
start_time = time()
OTTIMIZZATORE = 'rmsprop'
LOSS = 'categorical_crossentropy'
BATCH = 120
EPOCHS = 10
BATCH_CONV = 120
EPOCHS_CONV = 10
```
### Exercise 12.1
By keeping fixed all the other parameters, try to use at least two other optimizers, different from SGD. <span style="color:red">Watch to accuracy and loss for training and validation data and comment on the performances</span>.
```
(train_images, train_labels), (test_images, test_labels) = mnist.load_data()
pixel_rows = train_images.shape[1]
pixel_col = train_images.shape[2]
def prepare_data(array):
array = array.reshape(array.shape[0], array.shape[1]*array.shape[2])
return array.astype('float32')/255.
x_train = prepare_data(train_images)
x_test = prepare_data(test_images)
y_train = to_categorical(train_labels)
y_test = to_categorical(test_labels)
def dense_model():
model = models.Sequential()
model.add(layers.Dense(512, activation='relu', input_shape=(28 * 28,)))
model.add(layers.Dropout(0.5))
model.add(layers.Dense(10, activation='softmax'))
return model
network = dense_model()
network.compile(optimizer=OTTIMIZZATORE,
loss=LOSS,
metrics=['accuracy'])
history = network.fit(x_train, y_train,
batch_size = BATCH,
epochs = EPOCHS,
validation_split = 0.1,
verbose=True,
shuffle=True)
figure(figsize=((10,5)), dpi=200)
plt.subplot(1,2,1)
plt.plot(history.history['loss'], label='Loss')
plt.plot(history.history['val_loss'], label='Validation Loss')
plt.title("Model Loss")
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend(loc='best')
plt.subplot(1,2,2)
plt.plot(history.history['accuracy'], label='Accuracy')
plt.plot(history.history['val_accuracy'], label='Validation Accuracy')
plt.title("Model Accuracy")
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.legend(loc='best')
plt.show()
network.evaluate(x_test, y_test)
```
### Exercise 12.2
Change the architecture of your DNN using convolutional layers. Use `Conv2D`, `MaxPooling2D`, `Dropout`, but also do not forget `Flatten`, a standard `Dense` layer and `soft-max` in the end. I have merged step 2 and 3 in the following definition of `create_CNN()` that **<span style="color:red">you should complete</span>**:
```
def prepare_data_conv(array):
if keras.backend.image_data_format() == 'channels_first':
array = array.reshape(array.shape[0], 1, array.shape[1], array.shape[2])
shape = (1, array.shape[1], array.shape[2])
else:
array = array.reshape(array.shape[0], array.shape[1], array.shape[2], 1)
shape = (array.shape[1], array.shape[2], 1)
return array.astype('float32')/255., shape
x_train, INPUT_SHAPE = prepare_data_conv(train_images)
x_test, test_shape = prepare_data_conv(test_images)
def conv_model():
model = models.Sequential()
model.add(layers.Conv2D(32,
(3,3),
activation='relu',
input_shape=INPUT_SHAPE))
model.add(layers.MaxPooling2D((2,2)))
model.add(layers.Conv2D(64, (3,3), activation='relu',))
model.add(layers.MaxPooling2D((2,2)))
model.add(layers.Conv2D(64, (3,3), activation='relu',))
model.add(layers.Flatten())
model.add(layers.Dense(64, activation='relu'))
model.add(layers.Dense(10, activation='softmax'))
return model
network_conv = conv_model()
network_conv.compile(optimizer=OTTIMIZZATORE, loss=LOSS, metrics=['accuracy'])
history = network_conv.fit(x_train,
y_train,
validation_split=0.1,
verbose=True,
batch_size=BATCH,
epochs=5,
shuffle=True)
figure(figsize=((10,5)), dpi=200)
plt.subplot(1,2,1)
plt.plot(history.history['loss'], label='Loss')
plt.plot(history.history['val_loss'], label='Validation Loss')
plt.title("Model Loss")
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.grid(True)
plt.legend(loc='best')
plt.subplot(1,2,2)
plt.plot(history.history['accuracy'], label='Accuracy')
plt.plot(history.history['val_accuracy'], label='Validation Accuracy')
plt.title("Model Accuracy")
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.legend(loc='best')
plt.grid(True)
plt.show()
network_conv.evaluate(x_test, y_test)
```
### Exercise 12.3
Use the `gimp` application to create 10 pictures of your "handwritten" digits, import them in your jupyter-notebook and try to see if your CNN is able to recognize your handwritten digits.
For example, you can use the following code to import a picture of an handwritten digit:
```
from PIL import Image
import os
plt.figure(figsize=(20,40), dpi=50)
full_data = np.zeros((10,28,28))
for k in range(0,10):
digit_filename = str(k)+".png"
digit_in = Image.open(digit_filename).convert('L')
pix=digit_in.load();
data = np.zeros((28, 28))
for j in range(28):
for i in range(28):
data[i,j]=pix[j,i]
plt.subplot(1,10,k+1)
plt.imshow(data, cmap='gray')
full_data[k,:,:] = data[:,:]
#data /= 255
plt.show()
print(data.shape)
```
Test di accuratezza con i numeri scritti a mano da me.
```
plt.figure(figsize=(20,40), dpi=50)
for k in range(0,10):
data = full_data[k,:,:]
data_conv = np.zeros((1,data.shape[0], data.shape[1]))
data_conv[0,:,:] = data[:,:]
data_conv, aa = prepare_data_conv(data_conv)
data = data.reshape(1, 28*28)
pred_0 = network.predict(data)
pred_1 = network_conv.predict(data_conv)
data = data.reshape(28,28)
plt.subplot(1,10,k+1)
plt.imshow(data, cmap='gray')
plt.title("Dense {}".format(np.argmax(pred_0))+"\n \nConv {}".format(np.argmax(pred_1)))
plt.axis('off')
plt.show()
```
Test di accuratezza con alcune cifre sempre dal ```MNIST```.
```
#X_test = X_test.reshape(X_test.shape[0], img_rows*img_cols)
predictions = network_conv.predict(x_test)
x_test = x_test.reshape(x_test.shape[0], 28, 28,1)
plt.figure(figsize=(15, 15))
for i in range(10):
ax = plt.subplot(2, 10, i + 1)
plt.imshow(x_test[i, :, :, 0], cmap='gray')
plt.title("Digit: {}\nPredicted: {}".format(np.argmax(y_test[i]), np.argmax(predictions[i])))
plt.axis('off')
plt.show()
end_time = time()
minutes = int((end_time - start_time)/60.)
seconds = int((end_time - start_time) - minutes*60)
print("Computation time: ", minutes, "min ", seconds, "sec.")
```
| github_jupyter |
# Prophecy of ATM Withdrawals
Agus Gunawan, Holy Lovenia
## Importing dataset
```
from datetime import datetime
from pandas import read_csv
import pandas as pd
from os import listdir, mkdir
from os.path import exists, isfile, join
```
### Read train data
#### Functions
```
def get_files_from_dir_path(dir_path):
files = []
for file in listdir(dir_path):
if file.endswith('.csv'):
files.append(file)
return files
def import_all_datasets(base_src_dir_path):
src_files = get_files_from_dir_path(base_src_dir_path)
date_parser = lambda dates: datetime.strptime(dates, '%Y-%m-%d')
datasets = {}
for i in range(0, len(src_files)):
current_src_file = src_files[i]
current_id = current_src_file.split('.')[0]
current_id = current_id.split('_')[0]
datasets[int(current_id)] = read_csv(base_src_dir_path + current_src_file, parse_dates=['date'], date_parser=date_parser)
datasets[int(current_id)] = datasets[int(current_id)].rename(columns={'date': 'ds', 'Withdrawals': 'y'})
return datasets
```
### Import all split training datasets
Due to different natures and patterns generated by each ATM machine, the training dataset was split based on the ATM machines, e.g. K1, K2, ... ATM machine has its own dataset respectively.
```
train_datasets = import_all_datasets('dataset/train/')
```
## Prophet model building
```
from fbprophet import Prophet
from fbprophet.diagnostics import cross_validation, performance_metrics
from fbprophet.plot import plot_cross_validation_metric, plot_yearly, plot_weekly
import calendar
```
### Define Payday (holiday seasonality)
During the end of the month, usually the `Withdrawals` value gets higher
```
gajian = pd.DataFrame({
'holiday' : 'gajian',
'ds' : pd.to_datetime(['2018-03-30', '2018-02-28', '2018-01-31']),
'lower_window' : -2,
'upper_window' : 2}
)
holidays = gajian
```
### Define weekly seasonality for Sunday
On Sundays, `Withdrawals` is almost half of the other days
```
def take_money(ds):
date = pd.to_datetime(ds)
switcher = {
6: 0.5
}
return switcher.get(date.weekday(), 1)
```
### Adding regressor column for Sunday's `take_money` in dataset
```
for i in range(1, len(train_datasets) + 1):
train_datasets[i]['take_money'] = train_datasets[i]['ds'].apply(take_money)
```
### Training Prophet models for each dataset
In this step, each model is trained using its own dataset. An additional regressor for Sunday's `take_money` (weekly seasonality) is added for every model.
```
prophets = {}
for i in range(1, len(train_datasets) + 1):
prophet = Prophet(yearly_seasonality=False,
weekly_seasonality=False,
daily_seasonality=False,
holidays=holidays)
prophet.add_regressor(name='take_money', mode='multiplicative')
prophet.fit(train_datasets[i])
prophets[i] = prophet
```
### Forecasting `Withdrawals`
For the sake of demonstration, let's just predict the next seven days using the first 10 ATM machines.
```
forecast_data = {}
for i in range(1, 10 + 1):
if i % 10 == 0:
print(str(i) + ' from ' + str(len(prophets)))
future_data = prophets[i].make_future_dataframe(periods=7, freq='d')
future_data['take_money'] = future_data['ds'].apply(take_money)
forecast_data[i] = prophets[i].predict(future_data)
```
## Performance measure
The performance measure for 10 first ATM machines is computed using cross-validation.
### Get performance metrics with cross-validation
```
pm = {}
for i in range(1, 10 + 1):
cv = cross_validation(prophets[i], horizon='7 days')
pm[i] = performance_metrics(cv)
```
### Show averaged MSE and MAPE from each data point
```
for i in range(1, len(pm) + 1):
print(i, pm[i][['mse']].mean(), pm[i][['mape']].mean())
```
## Result
### Preparing the answers
```
temp_forecast_data = forecast_data.copy()
for i in range(1, len(temp_forecast_data) + 1):
temp_forecast_data[i]['no. ATM'] = "K" + str(i)
for i in range(1, len(temp_forecast_data) + 1):
temp_forecast_data[i] = temp_forecast_data[i].rename(columns={'ds': 'date'})
answer = {}
for i in range(1, len(temp_forecast_data) + 1):
answer[i] = temp_forecast_data[i].loc[temp_forecast_data[i]['date'] > '2018-03-24']
answer[i] = answer[i][['no. ATM', 'date', 'yhat']]
answer[i] = answer[i].rename(columns={'yhat': 'prediction'})
if i % 10 == 0:
print(str(i) + ' from ' + str(len(temp_forecast_data)))
```
### Concat all of the answers into a single `DataFrame`
```
final_answer = pd.DataFrame()
final_answer_list = []
for i in range(1, len(answer) + 1):
final_answer_list.append(answer[i])
final_answer = pd.concat(final_answer_list)
```
### Save it as CSV
```
final_answer.to_csv('result/prediction.csv', index=False)
```
| github_jupyter |
```
import os
os.chdir('/Users/sheldon/git/podly_app/new_files')
import glob
files = [f for f in os.listdir('.') if os.path.isfile(f)]
files = glob.glob('*.txt')
import pandas as pd
series = []
for i in files:
series.append(i.split('.mp3')[0])
x = pd.DataFrame(series)
def read_podcast(file):
tempFile = open(file).read()
return tempFile
transcriptions = []
for i in files:
transcription = open(i).read()
transcriptions.append(transcription)
x['transcription'] = transcriptions
x.head()
x.columns = ['file','transcription']
fileList = range(1, len(files)+1)
x['index'] = fileList
x.head()
from nltk.corpus import stopwords
stop = set(stopwords.words('english'))
def tokenize_and_lower(textfile):
tokens = word_tokenize(textfile)
lower = [w.lower() for w in tokens]
filtered_words = [word for word in lower if word not in stop]
return filtered_words
from nltk.corpus import stopwords
from collections import Counter
import pandas as pd
import numpy as np
import nltk.data
from __future__ import division # Python 2 users only
import nltk, re, pprint
from nltk import word_tokenize
tokenized_all_docs = [tokenize_and_lower(doc) for doc in transcriptions]
tokenized_all_docs
from gensim import corpora
dictionary = corpora.Dictionary(tokenized_all_docs)
dictionary.save('sample.dict')
print dictionary
corpus = [dictionary.doc2bow(text) for text in tokenized_all_docs]
corpora.MmCorpus.serialize('corpus.mm', corpus)
from gensim import models, corpora, similarities
tfidf = models.TfidfModel(corpus)
corpus_tfidf = tfidf[corpus]
lsi = models.LsiModel(corpus_tfidf, id2word=dictionary, num_topics=10)
corpus_lsi = lsi[corpus_tfidf]
index = similarities.MatrixSimilarity(lsi[corpus])
lsi.print_topics(10)
def get_related_podcasts_query(query):
query = query.lower()
vec_box = dictionary.doc2bow(query.split())
vec_lsi = lsi[vec_box]
sims = index[vec_lsi]
sims = sorted(enumerate(sims), key=lambda item: -item[1])[0:10]
related_df = pd.DataFrame(sims,columns=['index','score'])
final_df = pd.merge(related_df, x, on='index')[['index','file','score']]
return final_df
def get_related_podcasts(index):
def getKey(item):
return item[1]
corpus = corpus_lsi[index]
corpus = sorted(corpus, key=getKey, reverse=True)[0:10]
related_df = pd.DataFrame(corpus,columns=['index','score'])
final_df = pd.merge(related_df, x, on='index')[['index','file','score']]
return final_df
get_related_podcasts_query('math')
related_podcasts = list(get_related_podcasts(1)['index'])
def get_topics_per_podcast(podcast_index):
def getKey(item):
return item[1]
topic_ids = [i for i in sorted(corpus_lsi[podcast_index], key=getKey, reverse=True) if i[1] > 0.10]
def get_topic_arrays(topic_ids):
x = []
for id in topic_ids:
list_of_words = sorted(lsi.show_topic(id[0], topn=5),key=getKey, reverse=True)
z = []
for word in list_of_words:
if word[1] > .05:
z.append(word)
x.append(z)
return x
topic_arrays = get_topic_arrays(topic_ids)
return topic_arrays
related_podcasts_topics_words = [[related_podcasts[i],get_topics_per_podcast(related_podcasts[i])] for i in range(0, len(related_podcasts))]
episode_podcasts = list(get_related_podcasts(1)['file'])
for i,k in enumerate(related_podcasts_topics_words):
print "Podcast: {}, ID: {}".format(i+1, k[0])
print "Episode Title: {}".format(episode_podcasts[i])
for num, topic in enumerate(k[1]):
print "topic: {}".format(num)
for word in topic:
print "word: {}, score:{}".format(word[0], word[1])
```
| github_jupyter |
# Clustering FOX Data #
In this notebook I try Kmeans clustering on FOX data to see if automatically assigned clusters can help determine which sentences are ads.
```
import re
import pandas as pd
import matplotlib.pyplot as plt
import pickle
import nltk
from nltk.corpus import stopwords
from nltk import word_tokenize
from nltk.stem import WordNetLemmatizer
from sklearn.model_selection import train_test_split
from sklearn.feature_extraction.text import TfidfVectorizer, CountVectorizer
from sklearn.cluster import KMeans
from sklearn.decomposition import TruncatedSVD
nltk.download('wordnet')
fox_df = pd.read_csv('../data/interim/fox-last-year-sent-comb.csv')
fox_df.head()
len(fox_df)
```
This is a massive dataset, nearly 3 million sentences, so I'm going to get a random 10% of it to work with.
```
fox_df, _ = train_test_split(fox_df.drop(columns=['Unnamed: 0',
'Unnamed: 0.1']).dropna(), test_size=0.9, random_state=18)
len(fox_df)
stop_words = stopwords.words('english')
stop_words.extend(['from', 'subject', 're', 'edu', 'use', 'not', 'would', 'say', 'could',
'_', 'be', 'know', 'good', 'go', 'get', 'do', 'done', 'try', 'many',
'some', 'nice', 'thank', 'think', 'see', 'rather', 'easy', 'easily',
'lot', 'lack', 'make', 'want', 'seem', 'run', 'need', 'even', 'right',
'line', 'even', 'also', 'may', 'take', 'come', 'hi', 'ha', 'le', 'u', 'wa', 'thi',
'to', 'one'])
def clean_sent(sentences):
for sent in sentences:
sent = re.sub('\s+', ' ', sent) # remove newline chars
sent = re.sub("\'", "", sent) # remove single quotes
sent = re.sub("([\d,\,\./!#$%&\'\":;>\?@\[\]`)(\+])+", "", sent) # remove digits and remove punctuation
sent = re.sub("([-])+", " ", sent)
yield(sent)
corpus = list(clean_sent(fox_df.sentence.values.tolist()))
corpus[:5]
```
From some previous experimentation with Fox data, I found that TfidF vectorization with bigrams gave the easiest-to-interpret kmeans clusters and that 75 clusters separated out a lot of ad copy
```
#lemmatize before vectorizing
class LemmaTokenizer:
def __init__(self):
self.wnl = WordNetLemmatizer()
def __call__(self, doc):
return [self.wnl.lemmatize(t) for t in word_tokenize(doc)]
vect = TfidfVectorizer(tokenizer=LemmaTokenizer(), strip_accents='unicode', stop_words='english',
min_df=2, max_df=0.3, ngram_range=(2,2))
pickle.dump(vect, open('../models/fox_vect.p', 'wb'))
fox_bow = vect.fit_transform(corpus)
vect.get_feature_names()
kmeans = KMeans(n_clusters=75, random_state=18)
results = kmeans.fit_predict(fox_bow)
pickle.dump(kmeans, open('../models/fox_kmeans.p', 'wb'))
#print out most indicative words
terms = vect.get_feature_names()
order_centroids = kmeans.cluster_centers_.argsort()[:, ::-1]
for i in range(75):
print ("Cluster %d:" % i)
for ind in order_centroids[i, :10]:
print (' %s' % terms[ind])
print()
```
I did attempt other types of clustering algorithms, but they all killed my computer with even 10,000 sentences, so I've stuck with KMeans and LDA (to follow).
```
# add results to fox data frame
fox_df['cluster'] = results
# view sentences for each cluster
file_contents = ''
for i in range(75):
file_contents += 'Cluster {}\n'.format(i)
counter = 0
for index, row in fox_df[fox_df.cluster == i].iterrows():
file_contents += row['sentence'] + '\n'
counter += 1
if counter > 20:
break
file_contents += '\n'
with open('../data/interim/fox-sentence-check-20200611.txt', 'w') as f:
f.write(file_contents)
# count number of sentences in each cluster
fox_df[['cluster', 'sentence']].groupby('cluster').count().sort_values(by='sentence', ascending=False)
```
Clearly Cluster 1 has the vast majority of the sentences in it. Kmeans clustering, at least with this number of clusters, is not effective on its own but does help pick out some ads with repetitive copy, and so clusters may be an effective feature for future supervised learning.
I can see that clusters with a smaller vocabulary are more likely to be ads. This might be something to look at in the future.
I manually assigned all the clusters to be ad clusters, news clusters, or mixed, and then added that information to the dataframe.
```
ad_clusters = [0, 2, 3, 6, 9, 11, 15, 16, 17, 18, 20, 21, 22, 23, 29, 30, 33, 34, 37,
39, 41, 42, 45, 47, 49, 51, 55, 56, 58, 61, 62, 65, 66, 67, 69, 73]
news_clusters = [4, 5, 7, 8, 10, 12, 13, 14, 19, 24, 25, 26, 27, 28, 31, 32, 35, 36,
38, 40, 43, 44, 46, 48, 50, 52, 53, 54, 57, 59, 60, 63, 64, 68, 70, 71,
72, 74]
mixed = [1]
fox_df['ad_cluster'] = 0
fox_df['news_cluster'] = 0
fox_df.head()
fox_df['ad_cluster'] = fox_df['cluster'].isin(ad_clusters)
fox_df['news_cluster'] = fox_df['cluster'].isin(news_clusters)
fox_df = fox_df.mask(fox_df == True, 1)
fox_df = fox_df.mask(fox_df == False, 0)
fox_df[['news_cluster', 'sentence']].groupby('news_cluster').count()
fox_df.to_csv('../data/interim/mini-fox-clustered.csv')
```
Now to visualize the effectiveness of the clustering, I'm using PCA dimension reduction to graph the cluster separation into news, ads, and otherwise.
```
svd = TruncatedSVD(n_components=2, n_iter=7, random_state=18)
X_new = svd.fit_transform(fox_bow)
fox_df = fox_df.reset_index()
fox_df.head()
X_new_df = pd.DataFrame(data=X_new, columns=['Component 1', 'Component 2'])
X_new_df['Cluster'] = fox_df['cluster']
X_new_df['Is News'] = fox_df['news_cluster']
X_new_df['Is Ad'] = fox_df['ad_cluster']
X_new_df.head()
X_new_df['Type'] = X_new_df.apply(lambda x: 'Ad' if x['Is Ad'] == 1 else 'Mixed', axis=1)
X_new_df['Type'] = X_new_df.apply(lambda x: 'News' if x['Is News'] == 1 else x['Type'], axis=1)
X_new_df.head()
fig = plt.figure(figsize = (8,8))
ax = fig.add_subplot(1,1,1)
ax.set_xlabel('Component 1', fontsize = 15)
ax.set_ylabel('Component 2', fontsize = 15)
ax.set_title('FOX 2 Component SVD', fontsize = 20)
targets = ['Mixed', 'News', 'Ad']
colors = ['r', 'g', 'b']
for target, color in zip(targets,colors):
indicesToKeep = X_new_df['Type'] == target
ax.scatter(X_new_df.loc[indicesToKeep, 'Component 1']
, X_new_df.loc[indicesToKeep, 'Component 2']
, c = color
, s = 50)
ax.legend(targets)
ax.grid()
```
Well, this isn't a very interesting visualization, but also it was mostly to see if there were visual ways to separate these.
| github_jupyter |
# Imports
```
import pandas as pd
import numpy as np
import seaborn as sns
import pycountry_convert as pc
import statsmodels.formula.api as sm
from statsmodels.tsa.seasonal import STL
from scipy.stats import pearsonr
from scipy.misc import derivative
from scipy.optimize import fsolve
import numpy as np
sns.set_style('darkgrid')
```
# Data view
```
df = pd.read_csv('final_data.csv')
df.head()
df.info()
df.dt = pd.to_datetime(df.dt)
df.describe()
df.isna().sum()
```
# Aggregate to continents
```
def get_continent(country):
try:
return pc.country_alpha2_to_continent_code(pc.country_name_to_country_alpha2(country))
except:
pass
df['Continent'] = df.apply(lambda row: get_continent(row.Country), axis=1)
df[df.Continent.isna()].loc[:, 'Country'].unique() # to be mapped/removed
#TODO instead of this there should be mapping of not detected countries to continent
df.dropna(inplace=True)
def avg_continent(continent):
time_series = pd.DataFrame(df[df.Continent==continent].groupby('dt').AverageTemperature.mean())
time_series['Continent'] = continent
time_series['x'] = list(range(len(time_series)))
return time_series
agg_continents = pd.concat([avg_continent(continent) for continent in df.Continent.unique()])
```
# Growth significancy
```
g = sns.lmplot(data=agg_continents, x='x', y='AverageTemperature', hue='Continent', scatter=False, lowess=True)
g.axes.flat[0].set_xticks(list(range(0, len(agg_continents.index.unique()), 120)))
g.axes.flat[0].set_xticklabels(list(df.year.unique())[::10], rotation=45)
g
def linear_regression(continental):
data = pd.DataFrame({'x': list(range(len(continental))), 'y': continental.values})
return sm.ols(data=data, formula='y ~ x').fit().params.x
agg_continents.groupby('Continent').AverageTemperature.agg(linear_regression).reset_index()
# consider using another test
def correlation_test(continental):
# optional but gives different results - to consider
res = STL(continental.values, period=12, seasonal=3).fit() # value of `seasonal` can be changed
continental -= res.seasonal
data = pd.DataFrame({'x': list(range(len(continental))), 'y': continental.values})
return pearsonr(data.x, data.y)
agg_continents.groupby('Continent').AverageTemperature.agg(correlation_test).reset_index()
```
# Start of global warming
```
def inflection_point(y, deg=3):
x = np.arange(len(y))
x0 = np.mean([x[0], x[-1]])
coef = np.polyfit(x, y, deg)
construct_polynomial = lambda coef: np.vectorize(
lambda x: np.dot(coef, np.array([x**i for i in range(len(coef)-1, -1, -1)])))
return y.index[int(round(fsolve(lambda x_prime: derivative(construct_polynomial(coef), x_prime, n=2), x0)[0]))], coef[0]
agg_continents.groupby('Continent').AverageTemperature.agg(inflection_point).reset_index()
```
| github_jupyter |
```
import numpy as np
import pandas as pd
np.random.seed(42)
import tensorflow as tf
tf.set_random_seed(42)
from keras.models import Model
from keras.layers import Dense, Conv2D, BatchNormalization, MaxPooling2D, Flatten, Dropout, Input
from keras.preprocessing.image import ImageDataGenerator
from keras.optimizers import SGD
import os
print(os.listdir("../input/"))
from PIL import Image
# Create a class to store global variables. Easier for adjustments.
class Configuration:
def __init__(self):
self.epochs = 50
self.batch_size = 16
self.maxwidth =0
self.maxheight=0
self.minwidth = 35000
self.minheight = 35000
self.imgcount=0
self.img_width_adjust = 224
self.img_height_adjust= 224
#Kaggle
self.data_dir = "../input/train/"
config = Configuration()
```
## Data Exploration
```
#Load an example photo
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
img=mpimg.imread('../input/train/c0/img_4013.jpg')
imgplot = plt.imshow(img)
img.shape
plt.show()
#Find the largest and smallest dimensions of all the pictures
def findPictureDims(path):
for subdir, dirs, files in os.walk(path):
for file in files:
if file.endswith(".jpg"):
config.imgcount+=1
filename = os.path.join(subdir, file)
image = Image.open(filename)
width, height = image.size
if width < config.minwidth:
config.minwidth = width
if height < config.minheight:
config.minheight = height
if width > config.maxwidth:
config.maxwidth = width
if height > config.maxheight:
config.maxheight = height
return
#Count the number of files in each subdirectory
def listDirectoryCounts(path):
d = []
for subdir, dirs, files in os.walk(path,topdown=False):
filecount = len(files)
dirname = subdir
d.append((dirname,filecount))
return d
def SplitCat(df):
for index, row in df.iterrows():
directory=row['Category'].split('/')
if directory[3]!='':
directory=directory[3]
df.at[index,'Category']=directory
else:
df.drop(index, inplace=True)
return
#Get image count per category
dirCount=listDirectoryCounts(config.data_dir)
categoryInfo = pd.DataFrame(dirCount, columns=['Category','Count'])
SplitCat(categoryInfo)
categoryInfo=categoryInfo.sort_values(by=['Category'])
print(categoryInfo.to_string(index=False))
#Print out mins and maxes and the image counts
findPictureDims(config.data_dir)
print("Minimum Width:\t",config.minwidth, "\tMinimum Height:",config.minheight)
print("Maximum Width:\t",config.maxwidth, "\tMaximum Height:",config.maxheight, "\tImage Count:\t",config.imgcount)
```
## Analysis
-All of the data in the training directory is of the same height and width.
-The aspect ratio of the pictures is 4:3, so any adjustments are made should be close to that ratio (see configuration)
## Building the Model
```
import numpy as np
from keras import layers
from keras.layers import Input, Add, Dense, Activation, ZeroPadding2D, BatchNormalization, Flatten, Conv2D, AveragePooling2D, MaxPooling2D, GlobalMaxPooling2D
from keras.models import Model, load_model
from keras.preprocessing import image
from keras.utils import layer_utils
from keras.utils.data_utils import get_file
from keras.applications.imagenet_utils import preprocess_input
import pydot
from IPython.display import SVG
from keras.utils.vis_utils import model_to_dot
from keras.utils import plot_model
#from resnets_utils import *
from keras.initializers import glorot_uniform
import scipy.misc
from matplotlib.pyplot import imshow
import keras.backend as K
K.set_image_data_format('channels_last')
K.set_learning_phase(1)
%matplotlib inline
# In[2]:
def identity_block(X, f, filters, stage, block):
"""
Arguments:
X -- input tensor of shape (m, n_H_prev, n_W_prev, n_C_prev)
f -- integer, specifying the shape of the middle CONV's window for the main path
filters -- python list of integers, defining the number of filters in the CONV layers of the main path
stage -- integer, used to name the layers, depending on their position in the network
block -- string/character, used to name the layers, depending on their position in the network
Returns:
X -- output of the identity block, tensor of shape (n_H, n_W, n_C)
"""
# defining name basis
conv_name_base = 'res' + str(stage) + block + '_branch'
bn_name_base = 'bn' + str(stage) + block + '_branch'
# Retrieve Filters
F1, F2, F3 = filters
# Save the input value. You'll need this later to add back to the main path.
X_shortcut = X
# First component of main path
X = Conv2D(filters = F1, kernel_size = (1, 1), strides = (1,1), padding = 'valid', name = conv_name_base + '2a', kernel_initializer = glorot_uniform(seed=0))(X)
X = BatchNormalization(axis = 3, name = bn_name_base + '2a')(X)
X = Activation('relu')(X)
# Second component of main path
X = Conv2D(filters = F2, kernel_size = (f, f), strides = (1,1), padding = 'same', name = conv_name_base + '2b', kernel_initializer = glorot_uniform(seed=0))(X)
X = BatchNormalization(axis = 3, name = bn_name_base + '2b')(X)
X = Activation('relu')(X)
# Third component of main path
X = Conv2D(filters = F3, kernel_size = (1, 1), strides = (1,1), padding = 'valid', name = conv_name_base + '2c', kernel_initializer = glorot_uniform(seed=0))(X)
X = BatchNormalization(axis = 3, name = bn_name_base + '2c')(X)
# Final step: Add shortcut value to main path, and pass it through a RELU activation
X = Add()([X, X_shortcut])
X = Activation('relu')(X)
return X
# In[3]:
def convolutional_block(X, f, filters, stage, block, s = 2):
"""
Arguments:
X -- input tensor of shape (m, n_H_prev, n_W_prev, n_C_prev)
f -- integer, specifying the shape of the middle CONV's window for the main path
filters -- python list of integers, defining the number of filters in the CONV layers of the main path
stage -- integer, used to name the layers, depending on their position in the network
block -- string/character, used to name the layers, depending on their position in the network
s -- Integer, specifying the stride to be used
Returns:
X -- output of the convolutional block, tensor of shape (n_H, n_W, n_C)
"""
# defining name basis
conv_name_base = 'res' + str(stage) + block + '_branch'
bn_name_base = 'bn' + str(stage) + block + '_branch'
# Retrieve Filters
F1, F2, F3 = filters
# Save the input value
X_shortcut = X
##### MAIN PATH #####
# First component of main path
X = Conv2D(F1, (1, 1), strides = (s,s), name = conv_name_base + '2a', kernel_initializer = glorot_uniform(seed=0))(X)
X = BatchNormalization(axis = 3, name = bn_name_base + '2a')(X)
X = Activation('relu')(X)
# Second component of main path
X = Conv2D(filters = F2, kernel_size = (f, f), strides = (1,1), padding = 'same', name = conv_name_base + '2b', kernel_initializer = glorot_uniform(seed=0))(X)
X = BatchNormalization(axis = 3, name = bn_name_base + '2b')(X)
X = Activation('relu')(X)
# Third component of main path
X = Conv2D(filters = F3, kernel_size = (1, 1), strides = (1,1), padding = 'valid', name = conv_name_base + '2c', kernel_initializer = glorot_uniform(seed=0))(X)
X = BatchNormalization(axis = 3, name = bn_name_base + '2c')(X)
##### SHORTCUT PATH ####
X_shortcut = Conv2D(filters = F3, kernel_size = (1, 1), strides = (s,s), padding = 'valid', name = conv_name_base + '1', kernel_initializer = glorot_uniform(seed=0))(X_shortcut)
X_shortcut = BatchNormalization(axis = 3, name = bn_name_base + '1')(X_shortcut)
# Final step: Add shortcut value to main path, and pass it through a RELU activation
X = Add()([X, X_shortcut])
X = Activation('relu')(X)
return X
# In[49]:
def ResNet50(input_shape = (224,224,3), classes = 10):
"""
Implementation of the popular ResNet50 the following architecture:
CONV2D -> BATCHNORM -> RELU -> MAXPOOL -> CONVBLOCK -> IDBLOCK*2 -> CONVBLOCK -> IDBLOCK*3
-> CONVBLOCK -> IDBLOCK*5 -> CONVBLOCK -> IDBLOCK*2 -> AVGPOOL -> TOPLAYER
Arguments:
input_shape -- shape of the images of the dataset
classes -- integer, number of classes
Returns:
model -- a Model() instance in Keras
"""
# Define the input as a tensor with shape input_shape
X_input = Input(input_shape)
# Zero-Padding
X = ZeroPadding2D((3, 3))(X_input)
# Stage 1
X = Conv2D(32, (7, 7), strides = (2, 2), name = 'conv1', kernel_initializer = glorot_uniform(seed=0))(X)
X = BatchNormalization(axis = 3, name = 'bn_conv1')(X)
X = Activation('relu')(X)
X = MaxPooling2D((3, 3), strides=(2, 2))(X)
# Stage 2
X = convolutional_block(X, f = 5, filters = [32, 32, 128], stage = 2, block='a', s = 1)
X = identity_block(X, 5, [32, 32, 128], stage=2, block='b')
X = identity_block(X, 5, [32, 32, 128], stage=2, block='c')
# Stage 3
X = convolutional_block(X, f = 3, filters = [64,64,256], stage = 3, block='a', s = 2)
X = identity_block(X, 3, [64,64,256], stage=3, block='b')
X = identity_block(X, 3, [64,64,256], stage=3, block='c')
X = identity_block(X, 3, [64,64,256], stage=3, block='d')
# Stage 4
X = convolutional_block(X, f = 3, filters = [128,128,512], stage = 4, block='a', s = 2)
X = identity_block(X, 3, [128,128,512], stage=4, block='b')
X = identity_block(X, 3, [128,128,512], stage=4, block='c')
X = identity_block(X, 3, [128,128,512], stage=4, block='d')
X = identity_block(X, 3, [128,128,512], stage=4, block='e')
X = identity_block(X, 3, [128,128,512], stage=4, block='f')
# Stage 5
X = convolutional_block(X, f = 3, filters = [256,256,1024], stage = 5, block='a', s = 2)
X = identity_block(X, 3, [256,256,1024], stage=5, block='b')
X = identity_block(X, 3, [256,256,1024], stage=5, block='c')
# AVGPOOL
X = AveragePooling2D(pool_size=(5, 5), padding='valid')(X)
# output layer
X = Flatten()(X)
X = Dense(classes, activation='softmax', name='fc' + str(classes), kernel_initializer = glorot_uniform(seed=0))(X)
# Create model
model = Model(inputs = X_input, outputs = X, name='ResNet50')
return model
# In[50]:
model = ResNet50(input_shape = (224, 224, 3), classes = 10)
model.summary()
#Model Definition
def build_model():
inputs = Input(shape=(config.img_width_adjust,config.img_height_adjust,3), name="input")
#Convolution 1
conv1 = Conv2D(128, kernel_size=(3,3), activation="relu", name="conv_1")(inputs)
pool1 = MaxPooling2D(pool_size=(2, 2), name="pool_1")(conv1)
#Convolution 2
conv2 = Conv2D(64, kernel_size=(3,3), activation="relu", name="conv_2")(pool1)
pool2 = MaxPooling2D(pool_size=(2, 2), name="pool_2")(conv2)
#Convolution 3
conv3 = Conv2D(32, kernel_size=(3,3), activation="relu", name="conv_3")(pool2)
pool3 = MaxPooling2D(pool_size=(2, 2), name="pool_3")(conv3)
#Convolution 4
conv4 = Conv2D(16, kernel_size=(3,3), activation="relu", name="conv_4")(pool3)
pool4 = MaxPooling2D(pool_size=(2, 2), name="pool_4")(conv4)
#Fully Connected Layer
flatten = Flatten()(pool4)
fc1 = Dense(1024, activation="relu", name="fc_1")(flatten)
#output
output=Dense(10, activation="softmax", name ="softmax")(fc1)
# finalize and compile
model = Model(inputs=inputs, outputs=output)
model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=["accuracy"])
return model
model1 = build_model()
model1.summary()
#Setup data, and create split for training, testing 80/20
def setup_data(train_data_dir, val_data_dir, img_width=config.img_width_adjust, img_height=config.img_height_adjust, batch_size=config.batch_size):
train_datagen = ImageDataGenerator(rescale=1./255,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True,
validation_split=0.2) # set validation split
train_generator = train_datagen.flow_from_directory(
train_data_dir,
target_size=(img_width, img_height),
batch_size=batch_size,
class_mode='categorical',
subset='training')
validation_generator = train_datagen.flow_from_directory(
train_data_dir,
target_size=(img_width, img_height),
batch_size=batch_size,
class_mode='categorical',
subset='validation')
#Note uses training dataflow generator
return train_generator, validation_generator
def fit_model(model, train_generator, val_generator, batch_size, epochs):
model.fit_generator(
train_generator,
steps_per_epoch=train_generator.samples // batch_size,
epochs=epochs,
validation_data=val_generator,
validation_steps=val_generator.samples // batch_size,
verbose=1)
return model
#Verbose: 0: no output, 1: output with status bar, 2: Epochs Only
# Model Evaluation
def eval_model(model, val_generator, batch_size):
scores = model.evaluate_generator(val_generator, steps=val_generator.samples // batch_size)
print("Loss: " + str(scores[0]) + " Accuracy: " + str(scores[1]))
# Create Data 80/20
train_generator, val_generator = setup_data(config.data_dir, config.data_dir, batch_size=config.batch_size)
import time
#let's also import the abstract base class for our callback
from keras.callbacks import Callback
#defining the callback
class TimerCallback(Callback):
def __init__(self, maxExecutionTime, byBatch = False, on_interrupt=None):
# Arguments:
# maxExecutionTime (number): Time in minutes. The model will keep training
# until shortly before this limit
# (If you need safety, provide a time with a certain tolerance)
# byBatch (boolean) : If True, will try to interrupt training at the end of each batch
# If False, will try to interrupt the model at the end of each epoch
# (use `byBatch = True` only if each epoch is going to take hours)
# on_interrupt (method) : called when training is interrupted
# signature: func(model,elapsedTime), where...
# model: the model being trained
# elapsedTime: the time passed since the beginning until interruption
self.maxExecutionTime = maxExecutionTime * 60
self.on_interrupt = on_interrupt
#the same handler is used for checking each batch or each epoch
if byBatch == True:
#on_batch_end is called by keras every time a batch finishes
self.on_batch_end = self.on_end_handler
else:
#on_epoch_end is called by keras every time an epoch finishes
self.on_epoch_end = self.on_end_handler
#Keras will call this when training begins
def on_train_begin(self, logs):
self.startTime = time.time()
self.longestTime = 0 #time taken by the longest epoch or batch
self.lastTime = self.startTime #time when the last trained epoch or batch was finished
#this is our custom handler that will be used in place of the keras methods:
#`on_batch_end(batch,logs)` or `on_epoch_end(epoch,logs)`
def on_end_handler(self, index, logs):
currentTime = time.time()
self.elapsedTime = currentTime - self.startTime #total time taken until now
thisTime = currentTime - self.lastTime #time taken for the current epoch
#or batch to finish
self.lastTime = currentTime
#verifications will be made based on the longest epoch or batch
if thisTime > self.longestTime:
self.longestTime = thisTime
#if the (assumed) time taken by the next epoch or batch is greater than the
#remaining time, stop training
remainingTime = self.maxExecutionTime - self.elapsedTime
if remainingTime < self.longestTime:
self.model.stop_training = True #this tells Keras to not continue training
print("\n\nTimerCallback: Finishing model training before it takes too much time. (Elapsed time: " + str(self.elapsedTime/60.) + " minutes )\n\n")
#if we have passed the `on_interrupt` callback, call it here
if self.on_interrupt is not None:
self.on_interrupt(self.model, self.elapsedTime)
timerCallback = TimerCallback(350)
from keras.callbacks import ModelCheckpoint
model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
batch_size = 16
model.fit_generator(
train_generator,
steps_per_epoch=train_generator.samples // 16,
epochs=config.epochs,
validation_data=val_generator,
validation_steps=val_generator.samples // 16,
verbose=1,callbacks = [timerCallback, ModelCheckpoint('my_weights.h5')])
```
## Evaluation
```
# Evaluate your model.
eval_model(model, val_generator, batch_size=config.batch_size)
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import glob
%matplotlib inline
### Reading all file from glob module
### The glob module is used to retrieve files/pathnames matching a specified pattern.
path = r'E:\project\Transport_Vehicle_Online_Sales\Data_Vehicle_Sales'
all_files = glob.glob(path + "/*.csv")
all_file_list= []
for filename in all_files:
df = pd.read_csv(filename, index_col=None, header=0, encoding = "latin")
all_file_list.append(df)
### Concating all the dataframes of differnt csv files
data1 = pd.concat(all_file_list[1:], axis=0, ignore_index=True)
### Droped 'fromdata' and 'todate' columns from "data1" dataframe
data1 = data1.drop(['fromdate', 'todate'], axis=1)
### Droped 'Approved_Dt' columns
### data2 in this dataframe InsuranceCompany name is given
data2 = all_file_list[0].drop(['Apprved_Dt'], axis=1)
### Droped 'InsuranceCompany' columns from data2 inorder to make the frame collumns and data2Temp columns equal
data2Temp = data2.drop(['InsuranceCompany'], axis=1)
data2Temp.columns
### change columns names
data2Temp.columns = ['slno', 'modelDesc', 'fuel', 'colour', 'vehicleClass', 'makeYear',
'seatCapacity', 'insuranceValidity', 'secondVehicle',
'tempRegistrationNumber', 'category', 'makerName', 'OfficeCd']
### Again concating two data frames i.e frame and data2Temp
li=[data1, data2Temp]
data1 = pd.concat(li, axis=0, ignore_index=True)
### Dropped slno columns from data1
data1=data1.drop(['slno'], axis=1)
data1.head(2)
### drop insuranceValidity
data1 = data1.drop(['insuranceValidity'], axis=1)
data1['fuel'].unique()
data1.loc[data1['fuel']=='D', 'fuel'] = 'DIESEL'
data1.loc[data1['fuel']=='P', 'fuel'] = 'PETROL'
data1.loc[data1['fuel']=='L', 'fuel'] = 'LPG'
data1.loc[data1['fuel']=='B', 'fuel'] = 'BATTERY'
data1.loc[data1['fuel']=='C', 'fuel'] = 'CNG'
data1['fuel'].unique()
```
## Handling missing values
```
data1.isnull().sum()
data1[data1['fuel'].isnull()]['vehicleClass'].value_counts()
sop=data1[data1['vehicleClass']=='Trailer For Commercial Use']['fuel'].notnull()
sop=list(sop[sop].index)
data1.loc[sop, 'fuel'].value_counts()
### Gathering all notnull value of the different catogery of model name i.e 'modelDesc' depending on its fuel type
### Gathering of fuel type value from 'vehicleClass' of 'Trailer For Commercial Use'
TC = data1[data1['vehicleClass'] == 'Trailer For Commercial Use']['fuel'].notnull()
TC = list(TC[TC].index)
dieselName = data1.loc[TC, 'fuel'] == 'DIESEL'
dieselName = list(dieselName[dieselName].index)
dieselName = list(data1.iloc[dieselName,1].unique())
oneName = data1.loc[TC, 'fuel'] == '-1'
oneName = list(oneName[oneName].index)
oneName = list(data1.loc[oneName, 'modelDesc'].unique())
TC = data1[data1['vehicleClass'] == 'Trailer For Commercial Use']['fuel'].isnull()
TC = list(TC[TC].index)
data1.loc[TC, 'modelDesc'].isin(dieselName).value_counts()
TC = data1[data1['vehicleClass'] == 'Trailer For Commercial Use']['fuel'].isnull()
TC = list(TC[TC].index)
dcc=0
occ=0
for i in TC:
if data1.loc[i, 'modelDesc'] in dieselName:
dcc=dcc+1
elif data1.loc[i, 'modelDesc'] in oneName:
occ=occ+1
print("diesel name of notnull fuel value match with nan fuel value's model name",dcc)
print("-1 name of notnull fuel value match with nan fuel value's model name",occ)
### filling missing value based on same model name
### also noticed that some name are common among diesel and non fuel varient name
### so as per data without fuel varient is large in number and diesel varient is less in number
### so we try to fill diesel value first
dieN = data1.loc[TC,'modelDesc'].isin(dieselName)
oneN = data1.loc[TC,'modelDesc'].isin(oneName)
data1.loc[dieN[dieN].index, 'fuel'] = 'DIESEL'
data1.loc[oneN[oneN].index, 'fuel'] = '-1'
data1[data1['fuel'].isnull()]['vehicleClass'].value_counts()
### fuel type in "Trailer For Commercial Use"
dddd=data1[data1['fuel'].notnull()]['vehicleClass']=='Trailer For Commercial Use'
data1.loc[dddd[dddd].index,'fuel'].value_counts().plot.bar()
### filling rest missing value with most occured value
catt=data1[data1['fuel'].isnull()]['vehicleClass'] == 'Trailer For Commercial Use'
data1.loc[catt[catt].index, 'fuel'] = '-1'
data1[data1['fuel'].isnull()]['vehicleClass'].value_counts()
### Handling missing 'fuel' value of 'vehicleClass' sub category "Trailer for Agriculture Purpose"
dddd=data1[data1['fuel'].notnull()]['vehicleClass'] == 'Trailer for Agriculture Purpose'
data1.loc[dddd[dddd].index,'fuel'].value_counts().plot.bar()
### Gathering all notnull value of the different catogery of model name i.e 'modelDesc' depending on its fuel type
### Gathering of fuel type value from 'vehicleClass' of 'Trailer for Agriculture Purpose'
TC = data1[data1['vehicleClass'] == 'Trailer for Agriculture Purpose']['fuel'].notnull()
TC = list(TC[TC].index)
dieselName = data1.loc[TC, 'fuel'] == 'DIESEL'
dieselName = list(dieselName[dieselName].index)
dieselName = list(data1.loc[dieselName, 'modelDesc'].unique())
petrolName = data1.loc[TC, 'fuel'] == 'PETROL'
petrolName = list(petrolName[petrolName].index)
petrolName = list(data1.loc[petrolName,'modelDesc'].unique())
oneName = data1.loc[TC, 'fuel'] == '-1'
oneName = list(oneName[oneName].index)
oneName = list(data1.loc[oneName,'modelDesc'].unique())
TC = data1[data1['vehicleClass'] == 'Trailer for Agriculture Purpose']['fuel'].isnull()
TC = list(TC[TC].index)
data1.loc[TC, 'modelDesc'].isin(oneName).value_counts()
### notnull fuel type model name match with nan fuel value model name
TC = data1[data1['vehicleClass'] == 'Trailer for Agriculture Purpose']['fuel'].isnull()
TC = list(TC[TC].index)
dcc=0
occ=0
pcc=0
for i in TC:
if data1.loc[i, 'modelDesc'] in dieselName:
dcc=dcc+1
elif data1.loc[i, 'modelDesc'] in oneName:
occ=occ+1
elif data1.loc[i, 'modelDesc'] in petrolName:
pcc=pcc+1
print("Diesel name of notnull fuel value match with nan fuel value's model name = ",dcc)
print("")
print("Petrol name of notnull fuel value match with nan fuel value's model name = ",pcc)
print("")
print("-1 name of notnull fuel value match with nan fuel value's model name = ",occ)
dieN = data1.loc[TC, 'modelDesc'].isin(dieselName)
oneN = data1.loc[TC, 'modelDesc'].isin(oneName)
data1.loc[dieN[dieN].index, 'fuel'] = 'DIESEL'
data1.loc[oneN[oneN].index, 'fuel'] = '-1'
data1[data1['fuel'].isnull()]['vehicleClass'].value_counts()
### fuel type in "Trailer for Agriculture Purpose"
dddd=data1[data1['fuel'].notnull()]['vehicleClass']=='Trailer for Agriculture Purpose'
data1.loc[dddd[dddd].index,'fuel'].value_counts().plot.bar()
### filling rest missing value with most occured value
catt=data1[data1['fuel'].isnull()]['vehicleClass'] == 'Trailer for Agriculture Purpose'
data1.loc[catt[catt].index, 'fuel'] = '-1'
#### handling missing values of 'MOTOR CYCLE' of vehicleClass
### fuel type in "MOTOR CYCLE"
dddd=data1[data1['fuel'].notnull()]['vehicleClass']=='MOTOR CYCLE'
data1.loc[dddd[dddd].index,'fuel'].value_counts()
### filling rest missing value with most occured value of 'MOTOR CYCLE'
catt=data1[data1['fuel'].isnull()]['vehicleClass'] == 'MOTOR CYCLE'
data1.loc[catt[catt].index, 'fuel'] = 'PETROL'
#### handling missing values of 'MOTOR CAR' of vehicleClass
### fuel type in "MOTOR CAR"
dddd=data1[data1['fuel'].notnull()]['vehicleClass']=='MOTOR CAR'
data1.loc[dddd[dddd].index,'fuel'].value_counts().plot.bar()
### Motor car missing value filled by random value i.e (Petrol,Diesel) because occurrence of both is maximum
car=data1[data1['vehicleClass'] == 'MOTOR CAR']['fuel'].isnull()
count_car = len(car[car])
np.random.seed(0)
car_varient = np.random.choice(['DIESEL','PETROL'], size=count_car)
data1.loc[car[car].index, 'fuel'] = car_varient
data1[data1['fuel'].isnull()]['vehicleClass'].value_counts().sort_values( ascending=False)
### maximum missing value of top 3 vehicleClass fuel varient
val=data1[data1['fuel'].isnull()]['vehicleClass'].unique()[:3]
fig, axes = plt.subplots(nrows=1,ncols=3, figsize=(15,4),constrained_layout=True)
data1.loc[data1['vehicleClass']=='Tractor for Agricultural Purpose','fuel'].value_counts().plot(ax=axes[0], kind='bar', title='Tractor for Agricultural Purpose')
data1.loc[data1['vehicleClass']=='Luxory Tourist Cab','fuel'].value_counts().plot(ax=axes[1], kind='bar', title='Luxory Tourist Cab')
data1.loc[data1['vehicleClass']=='Motor Cab','fuel'].value_counts().plot(ax=axes[2], kind='bar', title='Motor Cab')
data1.loc[data1['fuel'].isnull(),'fuel']='DIESEL'
```
### Handling OfficeCd missing value
```
d=data1.isnull().sum().sort_values(ascending=False)
d[d.values>0]
### vehicle first 4 digit number
data1["rto"]=data1['tempRegistrationNumber'].astype(str).str[0:4]
### vehicle state
data1['state'] = data1['rto'].astype(str).str[:2]
data1.apply(lambda x : pd.factorize(x)[0]).corr(method='pearson', min_periods=1)
### for OfficeCd missing values
### 1.) group the rto and category
### 2.) take the top 3 most frequent OfficeCd of peticular group
### 3.) randomly fill that value in perticular group
c_test=data1.groupby(['rto', 'category']).groups
for cate_rto in c_test.keys():
missing_office=data1.loc[c_test[cate_rto], 'OfficeCd'].isnull().sum()
if missing_office:
dat = data1.loc[c_test[cate_rto], 'OfficeCd'].value_counts().sort_values(ascending=False).keys()[:3]
count_office = data1.loc[c_test[cate_rto],'OfficeCd'].isnull().sum()
np.random.seed(0)
choice_cate = np.random.choice(dat, size=count_office)
d=data1.loc[c_test[cate_rto], 'OfficeCd'].isnull()
data1.loc[d[d].index, 'OfficeCd'] = choice_cate
d=data1.isnull().sum()
d[d>0]
### Replacing Colour NIL value with most frequent colour of perticular vehicle class
d=data1['colour']=='NIL'
vehi_class = data1.loc[d[d].index, 'vehicleClass'].unique()
for clas in vehi_class:
val = data1[data1['vehicleClass'] == clas]['colour'].value_counts().sort_values(ascending=False).keys()
if val[0].upper() == 'NIL':
val = val[1]
else:
val=val[0]
d = data1[(data1['vehicleClass'] == clas)&(data1['colour']=='NIL')]
data1.loc[d.index, 'colour'] = val
### handling colour missing value
### filling nan value with most frequent colour of perticular vehicle class
d=data1['colour'].isnull()
vehi_class = data1.loc[d[d].index, 'vehicleClass'].unique()
data1['colour'] = data1['colour'].str.upper()
for clas in vehi_class:
val = data1[data1['vehicleClass'] == clas]['colour'].value_counts().sort_values(ascending=False).keys()
if val[0].upper() == 'NIL':
print(val)
else:
val=val[0]
d = data1[data1['vehicleClass'] == clas]['colour'].isnull()
data1.loc[d[d].index, 'colour'] = val
data1.isnull().sum()
```
### Features play
```
### manufacturing month and manufacturing year
d=data1['makeYear'].notnull()
data1['manufacturing_year']=data1.loc[d[d].index,'makeYear'].apply(lambda x: str('20') + x.split('/')[2] if len(x.split('/')[2])==2 else x.split('/')[2])
data1['manufacturing_year'] = data1['manufacturing_year'].astype(int)
dict_month={'02':'February', '01':'January', '04':'April ', '03':'March', '05':'May', '11':'November',
'07':'July', '12':'December', '09':'September', '10':'October', '08':'August',
'06':'Jun'}
data1['manufacturing_month']=data1.loc[d[d].index,'makeYear'].apply(lambda x: x.split('/')[1]).map(dict_month)
c=data1['manufacturing_year'].astype(int)
c[c>2020].index
data1.loc[3577031,'manufacturing_year']=1985
data1['howOld']=2020-data1['manufacturing_year'].astype(int)
data1.columns
### maximum missing value of top 3 vehicleClass fuel varient
#fig, axes = plt.subplots(nrows=2,ncols=2, figsize=(15,6),constrained_layout=True)
fig = plt.figure(figsize=(15,6))
ax1 = fig.add_subplot(221)
ax2 = fig.add_subplot(222)
ax3 = fig.add_subplot(212)
data1.boxplot(column = 'manufacturing_year', ax=ax1, vert=False)
data1.boxplot(column='howOld', ax=ax2, vert=False)
data1.boxplot(column='seatCapacity', ax=ax3, vert=False)
import matplotlib
fig, ax1 = plt.subplots(figsize=(18, 15))
c=[c for c in data1.groupby(['howOld']).groups.keys()]
cy = [len(i) for i in list(data1.groupby(['howOld']).groups.values())]
ax1.plot(c,cy, '*g--')
ax1.get_yaxis().set_major_formatter(
matplotlib.ticker.FuncFormatter(lambda x, p: format(int(x), ',')))
ax1.grid()
plt.xlabel("How old vehicle", axes=ax1, size=15)
plt.ylabel("Total registerd vehicle", axes=ax1, size=15)
plt.title("Total vehicle distribution as per its age", axes=ax1, size=20)
plt.show()
fig, ax1 = plt.subplots(figsize=(18, 15))
c=[c for c in data1.groupby(['seatCapacity']).groups.keys()]
cy = [len(i) for i in list(data1.groupby(['seatCapacity']).groups.values())]
ax1.get_yaxis().set_major_formatter(
matplotlib.ticker.FuncFormatter(lambda x, p: format(int(x), ',')))
ax1.plot(c,cy, '*r--')
#plt.xticks(keys, rotation='vertical', size=15, axes=ax1)
ax1.grid()
plt.xlabel("Vehicle count", axes=ax1, size=15)
plt.ylabel("Vehicle Seat capacity", axes=ax1, size=15)
plt.title("Total vehicle distribution as per its Seat Capacity", axes=ax1, size=20)
plt.show()
### year sold vehicle class
keys = [pair for pair in data1.groupby(['manufacturing_year']).groups]
fig, ax = plt.subplots(figsize=(25, 20))
ax.get_yaxis().set_major_formatter(
matplotlib.ticker.FuncFormatter(lambda x, p: format(int(x), ',')))
c=[c for c in data1.groupby(['manufacturing_year']).groups.keys()]
cy = [len(i) for i in list(data1.groupby(['manufacturing_year']).groups.values())]
ax.plot(c,cy, '*r--')
plt.xticks(keys, rotation='vertical', size=15, axes=ax)
plt.yticks(np.arange(0, max(cy), 100000), axes=ax, size=20)
ax.grid(color='lightgreen')
plt.xlabel("Vehicle count", axes=ax1, size=20)
plt.ylabel("Vehicle Manufacturing Year", axes=ax1, size=20)
plt.title("Total vehicle distribution as per its Manufacturing Year", size=25)
plt.show()
data1['vehicleClass'].unique()
from random import randrange
c=[i for i in data1.groupby(['vehicleClass']).groups.keys()]
cy=[len(i) for i in data1.groupby('vehicleClass').groups.values()]
fig, ax =plt.subplots(figsize=(25, 20))
c_data = [randrange(50,250) for i in range(len(c))]
my_cmap = matplotlib.cm.get_cmap('jet')
plt.bar(c, cy, axes=ax, color=my_cmap(c_data))
plt.xticks(c, rotation='vertical', axes=ax, size=20)
plt.yticks(np.arange(0, max(cy), 100000), axes=ax, size=20)
ax.get_yaxis().set_major_formatter(
matplotlib.ticker.FuncFormatter(lambda x, p: format(int(x), ',')))
plt.xlabel("Vehicle count", axes=ax, size=30)
plt.ylabel("Vehicle class", axes=ax, size=30)
plt.title("Total vehicle distribution as per Vehicle class", axes=ax, size=30)
plt.grid(color='lightblue')
plt.show()
```
| github_jupyter |
# EEP/IAS 118 - Coding Bootcamp
## Introduction to R and Jupyter Notebooks
### August 2021
## Jupyter Notebook Basics
We're going to start off this term by using _Jupyter Notebooks_ to run __R__, our preferred programming language for statistical computing. Jupyter notebooks are an interactive computing environment that lets us combine both text elements and active **R** code. Importantly for us, notebooks let us
* Write and run __R__ code through our web browsers, and
* Add narrative text that describes our code and the output from said code
This means that we can use __R__ without having to install any software on our personal computers, and can ignore errors that might pop up from local conflicts or issues with packages on personal machines. Further, it lets us answer written exercises and coding problems all in one place.
When you clicked the link on bCourses, you were taken to a folder in your web browser. This folder is hosted on a remote web server through Berkeley's __Datahub__ and should look something like this:
<p style="text-align: center;"> Datahub Folder </p>
<img src="images/Section_Screen.png" width="800" />
Today we are doing Coding Resources, so you will want to click on that folder.
You should see the following files.
1. The first file _Coding Bootcamp Session.ipynb_ is our notebook for today
2. _autos.csv_ is a data file in comma separated value (csv) format
3. _autos.dta_ is the same data file, but formatted for Stata in .dta format
We'll be using both csv and dta data this term, so we'll practice reading in both formats.
(There is also a folder of images...don't worry about those!)
### Running Jupyter Notebooks
Double clicking on the _Coding Bootcamp.ipynb_ notebook will open up the notebook in another browser window. The notebook that opens should look... well, like this!
At the top of the page you'll find the menubar and toolbar:
<img src="images/menubar_toolbar.png" width="800" />
From this menu we can run our code, display our text, save our notebook, and download a pdf or other format of our notebook.
Clicking on **File** in the menu bar lets us save the notebook and build a checkpoint. You can use **Revert to Checkpoint** to return to previous checkpoints if you want to go back to a previous version of the notebook.
**Print Preview** lets you view the notebook as it would look when printed.
We will be using the **Download As > PDF via HTML** command to export a pdf copy of the notebook after running all text/code for submission. This is the first option on the menu.
### Editing Cells
This and all notebooks are comprised of a linear collection of boxes, called _cells_. For the sake of this class, we'll be working with two types of cells: *Markdown* cells for text, and *Code* for writing and executing **R** code.
Markdown cells support plain text as well as markdown code, html, and LaTeX math mode. For this class, plain text answers are totally fine. If you want to dive into Markdown formatting, [this cheat sheet](https://github.com/adam-p/markdown-here/wiki/Markdown-Cheatsheet) has information on formatting text, building tables, and adding html. If you want to add pretty math equations, see [this LaTeX Math Mode Guide](https://www.overleaf.com/learn/latex/Mathematical_expressions).
Select a cell by right clicking on it. A grey box with a blue bar to the left will appear around the cell, like so:
<img src="images/command_mode.png" width="800" />
This means you are in __command mode__. In command mode you can see the cell type (whether it is markdown for text or code for __R__ commands) in the toolbar but can't edit the content of the cell.
To edit the content of cells, double click on the cell to enter __edit mode__. When in edit mode, the box surrounding the cell will turn green - as will the left margin.
A text (markdown) cell in edit mode should look like this:
<img src="images/edit_mode_text.png" width="800" />
and a code (R) cell like this:
<img src="images/edit_mode_code.png" width="800" />
Use edit mode to type in all your text and code. When you are done with your paragraph or want to try out your code, it's time to run the cell.
Try selecting this cell by right clicking on it, then double click to enter edit mode.
### Running Cells
When you are ready to run a cell (done typing your text in a cell and want to display it in formatted mode, or want to run **R** code), hit `shift` + `enter`, `control` + `enter`, or hit `Run` in the toolbar.
Running a text cell will exit edit mode, format the cell's text, and select the next cell down.
You can also run a cell from edit or command mode - try this by right clicking on the above cell that you were typing in, and hitting `shift` + `enter`.
We'll see later that running code cells will oftentimes add output to our notebook. This will be how we follow what we're doing in **R** and how we'll get our output.
## R
**R** is our programming language for statistical programming. It is open source and free, handles lots of different types of data, and has tons of different packages that allow it to a ton of different things (regression analysis, plotting, working with spatial data, web scraping, creating applications, machine learning to name a few).
### R in Code Cells
Thanks to *Datahub*, **R** is running in the background of our Jupyter notebooks. As a result, we can type **R** code into code cells, run the cell, and get results all in one place.
Let's try it: in the code cell below, type '2 + 2' and run the cell.
**R** took our code as input, and spit out the result - in this case the value of our summation, 4.
### R Syntax
*Note of caution:* **R** is a stickler for typos. Precise syntax is essential. Capital letters, commas, or parentheses must be in the correct place. Any deviation from required syntax will lead your code to either fail or produce unintended (and incorrect) results. You will spare yourself a lot of aggravation if you take the time to go slowly and carefully as you’re getting started until you have gotten more familiar with the commands and their required syntax.
### Packages, Libraries, and Paths
One of **R**’s best features is that it’s based on packages. Base **R** does have some stats functions, but **R** plus packages is incredibly powerful. Nearly any time you do anything in **R**, you’ll be using functions contained within a package.
Every time we want to use a function contained in a package, we must first call that package using the `library()` function. For example, we can load the **haven** package by typing and running the command `library(haven)` in the below cell:
```
library(haven)
```
The package was loaded correctly in the background. Since there was no output to display nor an error to show, we received no input. We can confirm that the package was loaded correctly by running the `sessionInfo()` function below.
```
sessionInfo()
```
This shows us information about our **R** version, local settings, active packages, ones we've manually loaded, and others that are loaded through our system.
If we had written the package name wrong, and instead tried to load a nonexistent package "hevon", we would receive an error in the notebook:
```
library(hevon)
```
### Paths
When we start loading in data files we will need to account for the location of files in our file paths. **R** handles paths by basing itself in the *working directory*, and all file paths are defined relative to that working directory.
We can view the working directory by using the `getwd()` function.
```
getwd()
```
The above path is the same as the folder our linked opened to. When calling files, we will start our paths in this folder - if the file is located in the same folder as our notebook, we do not need to include all the "/home/rstudio..." path, and instead can reference it by name. If the file we want to reference (let's call it *enviro.csv*) is in a subfolder (say called *Data*), we will need to include that subfolder in our path: "Data/enviro.csv"
We'll deal more with working directories once we move into **RStudio** - for now all the files we need will be in the same folder as our template, so we can call the files directly by name.
### Loading Files in R
We're going to read in our data, but before we can read in our dataset we should refresh ourselves on what the file is called. To do this, we can use the `list.files()` command to see all the files in our current working directory.
```
list.files()
```
The file we're interested in first is the **autos.dta** file. Since this is a stata-formatted file (.dta), we'll need to use the `read_dta()` function in the **haven** package (which we already loaded). To use the function we need to include the filename with quotes in the parentheses: `read_dta("autos.dta")`.
Here we run into an important feature about **R**: it is an object-oriented language. When we load a dataset or want to store an object to memory (such that we can do things with it later), we have to *assign it a name*. To do this, we use the syntax
` name <- function(arguments)`
The arrow tells us we are assigning the output of the function (given the function's arguments/inputs) to *name*. The arrow is read as "gets," so if we wanted to store a vector of integers between 1 through 10 as `integers` using the command
` integers <- 1:10`
We read it as "integers 'gets' 1 through 10."
Okay, let's load in the **autos** data and save it to the object `carsdata`:
```
carsdata<-read_dta("autos.dta")
```
The dataset has now been stored to memory under the name `carsdata`. To view it in our notebook, we can just type the name and hit run... or if we want to view only the first few lines of the dataset we can use the `head()` command to see only a few observations.
```
head(carsdata)
```
The very first bit of information tells us the format of the data (data frame). The first row tells us the variable names, and the second row tells us the variable type (character string or type of numeric variable). Each row in the table is a different observation - here giving us info on a car make and model, its price, mpg, and other characteristics.
If we forget the info in our dataframe, we can check the names of the variables:
```
names(carsdata)
```
If we want to interact with just one column of our data frame, we can refer to it using the `$` command. This is useful if we're interested in, say, obtaining the mean of only miles per gallon for the cars in our sample.
```
avg_mpg <- mean(carsdata$mpg)
avg_mpg
```
#### Other File Formats
If we wanted to load a .csv file, we'd want to use the `read.csv()` command to load it, once again using quotation marks around the file name and assigning it a name.
### Manipulating Data Frames
We have a lot of flexibility to select certain observations, certain variables, or certain values within our data frame. We can also perform a lot of operations to variables - change their values or create new variables. One of the packages we'll be using for this is the **tidyverse** package. It's actually a collection of packages designed for data science, and includes a number that we'll use throughout this term.
To start, let's load **tidyverse** and use it to perform transformations on our dataset.
```
library(tidyverse)
carsdata_low_price <- filter(carsdata, price <= 10000)
head(carsdata_low_price)
carsdata_low_price <- arrange(carsdata_low_price, desc(price))
head(carsdata_low_price)
carsdata_low_price <- select(carsdata_low_price, make, price, mpg, weight)
head(carsdata_low_price)
```
Phew, that was a lot of stuff! Let's back up and work through it.
The first thing we did after loading the **tidyverse** package was to `filter()` our data frame. Here we are selecting only the observations with price less than or equal to $10,000. We save this to the new `carsdata_low_price` object.
Next, we arrange the new `carsdata_low_price` data frame in descending order of price using the `arrange()` function and overwrite our `carsdata_low_price` with this new arrangement.
Finally, we `select()` only a few variables - here we choose to keep just vehicle make, price, and miles per gallon - and once again overwrite our `carsdata_low_price` object.
Now we'll create new versions of our variables using the `mutate()` command. If I wanted to rescale price to be in units of $1,000, we can do this by typing
```
carsdata_low_price <- mutate(carsdata_low_price, price_thousand = price/1000)
head(carsdata_low_price)
```
Here we are replacing `carsdata_low_price` object with itself after adding in a new variable names `price_thousand` that is `price` divided by 1000.
### Summarizing Variables
There are a number of different ways for us to obtain information about our variables in **R**. While we saw earlier one way to directly get the mean of a variable, we could instead have used the `summarise` command, which lets us obtain a number of different statistics - either one at a time, or multiple together
```
avg_mpg <- summarise(carsdata_low_price, mean(mpg))
avg_mpg
multi_stats <- summarise(carsdata_low_price, max(mpg), min(price), n())
multi_stats
```
`summarise()` is useful because we can use it to select specific summary statistics we're insterested in or create truly custom summary stats. If instead we wanted to get a bunch of basic stats all at once, we could use the `summary()` command for a given variable - say to get information on the variable `price`. We can refer to specific variables in a data frame with a `$` . So for example, `carsdata_low_price$price`.
```
summary(carsdata_low_price$price)
```
### Defining New Variables
Suppose we want to add new columns to our data frame. This can be done easily, by simply defining them with our `$` notation. Suppose we want to make a new column called price_sq which is the squared price of each car. We can do so by defining `carsdata_low_price$price_sq<-` (with an appropriately defining expression on the right side). To do this, note that you can call other columns and values you already have defined or loaded in to R. R also uses most basic math symbols in the ways you would expect: `+`, `-`, `*`,`\`, and `^`. Try it below!. Note: Be careful with order of operations and use parentheses when necessary.
### Plotting in R
Today we're going to learn how to use the basic plot function in **R**. In the coming weeks, we're going to learn **ggplot2**, a fantastic graphics package that is one of the great reasons for using **R** to produce graphics. Its syntax is a little bit more involved, so we're going to start off with some simple functions.
First, we can generate a histogram of vehicle prices using the `hist()` command.
```
hist(carsdata_low_price$price,
main = "Price Distribution",
xlab = "Price")
```
Now let's look at fuel economy, and add a blue line of width three at the average mpg:
```
hist(carsdata_low_price$mpg,
main = "Fuel Economy Distribution",
xlab = "Miles per Gallon")
abline(v = avg_mpg, col = "blue", lwd = 3)
```
We can also make a scatterplot of vehicle price and fuel efficiency using the `plot()` function.
```
plot(carsdata_low_price$mpg, carsdata_low_price$price, # x-axis first, y-axis second
main = "Price vs. Fuel Efficiency",
xlab = "Miles per Gallon",
ylab = "Price ($)") # don't forget the last parenthesis!
```
There seems to be a negative correlation here: the higher MPG, the lower the price. Is this potentially an SUV effect?
### Regression
Running regressions with **R** is quite easy. Later in the course we'll get into some more complex regression commands, but for now we'll stick with simple linear regression using the `lm()` command.
First, let's take a step back and see how mpg and price are correlated in the data.
```
mpg_price_cor <- cor(carsdata_low_price$mpg, carsdata_low_price$price)
mpg_price_cor
```
What if we used a simple OLS regression instead?
```
mpg_price_regression <- lm(mpg ~ price, data = carsdata_low_price)
# basic syntax: lm(depvar ~ indvar1 + indvar2 + ..., data=data)
mpg_price_regression
```
Our `mpg_price_regression` object contains our regression coefficients as well as a bunch of other objects, most notably residuals and fitted values.
```
head(mpg_price_regression$fitted.values)
mean(mpg_price_regression$residuals)
```
## Help with R
The internet is your best friend. While we will introduce you to some R commands during section, you may find that there are times where you are having trouble with a function's syntax or need a new function to perform a certain task. Asking Google, using the [R Documentation site](https://www.rdocumentation.org/) as well as [R Project Package Reference Manuals/Vignettes](https://cran.r-project.org/) for help with functions and syntax will get you quite far. If something is unclear, searching for the R task and perusing answers on [StackExchange](https://stackexchange.com/) can be a helpful resource. I will try my best to make sure homework assignments mention the functions/packages needed for a new task, and that we've at least seen them in section or lecture.
| github_jupyter |
```
from collections import Counter
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import ast
import json
df = pd.read_csv('/home/amir/projects/light-sa-type-inf/ManyTypes4Py_processed_fix_feb2/all_fns.csv',
low_memory=False)
jsons_merged = json.load(open('/home/amir/ManyTypes4Py_processed_Mar1/merged_projects.json', 'r'))
df.head()
print("Number of projects: ", len(set(df.apply(lambda x: x['author'] + "/" + x['repo'], axis=1))))
print(f"Number of source code files: {len(df['file'].unique()):,}")
print(f"Number of files with type annotations: {len(df[df['has_type'] == True].drop_duplicates('file', keep='last')):,}")
print(f"Number of functions: {len(df.fn_name):,}")
print(f"Number of classes: {df.cls_name.count():,}")
print(f"Number of functions with a type: {len(df[df['has_type'] == True]):,}")
print(f"Number of arguemts: {sum(df['arg_names'].apply(lambda x: len(ast.literal_eval(x)))):,}")
print(f"Number of arguemts with types: {sum(df['arg_types'].apply(lambda x: len([t for t in ast.literal_eval(x) if t != '']))):,}")
print(f"Number of functions with a return type: {df['return_type'].count():,}")
print(f"Number of functions with docstring: {df['docstring'].count():,}")
print(f"Number of functions with description: {df['func_descr'].count():,}")
print(f"Number of arguemts with comments: {sum(df['arg_descrs'].apply(lambda x: len([t for t in ast.literal_eval(x) if t != '']))):,}")
print(f"Number of functions with a comment: {df['docstring'].count() + df['fn_descr'].count() + sum(df['arg_descrs'].apply(lambda x: len([t for t in ast.literal_eval(x) if t != '']))) + df['return_descr'].count():,}")
print(f"Number functions with a return type comment: {df['return_descr'].count():,}")
args_t = list(df['arg_types'].apply(lambda x: [t for t in ast.literal_eval(x) if t != '']))
ret_t = list(df[df['return_type'].notnull()]['return_type'].values)
fn_vars_t = list(df['fn_variables_types'].apply(lambda x: [t for t in ast.literal_eval(x) if t != '']))
cls_vars_t = [t for cls_t in df.drop_duplicates('cls_name', keep='last')[df['cls_variables_types'].notnull()]['cls_variables_types'] for t in ast.literal_eval(cls_t) if t != '']
mod_vars_t = list(df.drop_duplicates('file', keep='last')['mod_variables_types'].apply(lambda x: [t for t in ast.literal_eval(x) if t != '']))
print(f"Total Number of type annotations: {sum([len(t)for t in args_t]) + len(ret_t) + sum([len(t) for t in fn_vars_t]) + len(cls_vars_t) + sum([len(t) for t in mod_vars_t]):,}")
print(f"Number of unique types: {len(set([t for l in args_t for t in l] + ret_t + [t for l in fn_vars_t for t in l] + cls_vars_t + [t for l in mod_vars_t for t in l])):,}")
def set_stats(d, s):
print(f"Number of projects: ", len(set(d[d['set'] == s].apply(lambda x: x['author'] + "/" + x['repo'], axis=1))))
print(f"Number of source code files: {len(d[d['set'] == s]['file'].unique()):,}")
print(f"Number of files with type annotations: {len(d[d['set'] == s][d['has_type'] == True].drop_duplicates('file', keep='last')):,}")
print(f"Number of functions: {len(d[d['set'] == s].fn_name):,}")
print(f"Number of functions with a type: {len(d[d['set'] == s][d['has_type'] == True]):,}")
print(f"Number of functions with a return type: {d[d['set'] == s]['return_type'].count():,}")
print(f"Number of functions with docstring: {d[d['set'] == s]['docstring'].count():,}")
print(f"Number of functions with description: {d[d['set'] == s]['fn_descr'].count():,}")
print(f"Number functions with a return type comment: {d[d['set'] == s]['return_descr'].count():,}")
print(f"Number of functions with a comment: {d[d['set'] == s]['docstring'].count() + d[d['set'] == s]['fn_descr'].count() + sum(d[d['set'] == s]['arg_descrs'].apply(lambda x: len([t for t in ast.literal_eval(x) if t != '']))) + d[d['set'] == s]['return_descr'].count():,}")
print(f"Number of arguemts: {sum(d[d['set'] == s]['arg_names'].apply(lambda x: len(ast.literal_eval(x)))):,}")
print(f"Number of arguemts with types: {sum(d[d['set'] == s]['arg_types'].apply(lambda x: len([t for t in ast.literal_eval(x) if t != '']))):,}")
print(f"Number of arguemts with comments: {sum(d[d['set'] == s]['arg_descrs'].apply(lambda x: len([t for t in ast.literal_eval(x) if t != '']))):,}")
set_stats(df, 'test')
set_stats(df, 'train')
def find_all_types_set(d, s):
args_t = list(d[d['set'] == s]['arg_types'].apply(lambda x: [t for t in ast.literal_eval(x) if t != '']))
ret_t = list(d[d['set'] == s][d['return_type'].notnull()]['return_type'].values)
fn_vars_t = list(d[d['set'] == s]['fn_variables_types'].apply(lambda x: [t for t in ast.literal_eval(x) if t != '']))
cls_vars_t = [t for cls_t in d.drop_duplicates('cls_name', keep='last')[d['cls_variables_types'].notnull()]['cls_variables_types'] for t in ast.literal_eval(cls_t) if t != '']
mod_vars_t = list(d.drop_duplicates('file', keep='last')['mod_variables_types'].apply(lambda x: [t for t in ast.literal_eval(x) if t != '']))
print(f"Total Number of type annotations: {sum([len(t)for t in args_t]) + len(ret_t) + sum([len(t) for t in fn_vars_t]) + len(cls_vars_t) + sum([len(t) for t in mod_vars_t]):,}")
print(f"Number of unique types: {len(set([t for l in args_t for t in l] + ret_t + [t for l in fn_vars_t for t in l] + cls_vars_t + [t for l in mod_vars_t for t in l])):,}")
find_all_types_set(df, 'train')
```
## Distribution of Top 10 frequent types in the dataset
```
all_types = [l for t in args_t for l in t] + ret_t + \
[l for t in fn_vars_t for l in t] + cls_vars_t + [l for t in mod_vars_t for l in t]
top_n=10
# Already sorted
df_plot = pd.DataFrame.from_records(Counter(all_types).most_common(top_n), columns=['type', 'count'])
print(f"Sum of Top {top_n} frequent types: {sum(counts)}")
types = df_plot['type'].values[:top_n]
counts = df_plot['count'].values[:top_n]
y_pos = np.arange(len(counts))
plt.barh(y_pos, counts, align='center', alpha=0.5)
plt.yticks(y_pos, types)
plt.xlabel('Count')
plt.ylabel('Types')
#plt.title('Top %d most frequent types in the dataset' % top_n)
plt.tight_layout() # Fixes the Y axis lables to be shown completely.
# plt.savefig('top-%d-most-frequent-types.pdf' % top_n, format='pdf', bbox_inches='tight',
# dpi=256)
plt.show()
all_types
```
Finding type annotation coverage
```
p_type_annote_cove = [(p, round(jsons_merged['projects'][p]['type_annot_cove'] * 100, 1)) for p in jsons_merged['projects'].keys()]
len(list(filter(lambda x: x[1] > 0, p_type_annote_cove)))
f_type_annote_cove = [(f, round(jsons_merged['projects'][p]['src_files'][f]['type_annot_cove'] * 100, 1)) for p in jsons_merged['projects'].keys() for f in jsons_merged['projects'][p]['src_files']]
f_type_annote_cove
type_cove_interval = {"[%d%%-%d%%%s" % (i, i+20, ']' if i + 20 == 100 else ')' ):[(i, i+20), 0] for i in range(0, 100, 20)}
for p, t_c in p_type_annote_cove:
for i, v in type_cove_interval.items():
if t_c >= v[0][0] and t_c < v[0][1]:
type_cove_interval[i][1] += 1
print(type_cove_interval)
f_type_cove_interval = {"[%d%%-%d%%%s" % (i, i+20, ']' if i + 20 == 100 else ')' ):[(i, i+20), 0] for i in range(0, 100, 20)}
for p, t_c in f_type_annote_cove:
for i, v in f_type_cove_interval.items():
if t_c >= v[0][0] and t_c < v[0][1]:
f_type_cove_interval[i][1] += 1
f_type_cove_interval
```
## Type annotations coverage for projects
```
f = plt.bar([i for i, v in type_cove_interval.items()], [v[1] for i, v in type_cove_interval.items()])
plt.xlabel("Percentage of type annotation coverage")
plt.ylabel("No. of projects")
for rect, v in zip(f.patches, [v[1] for i, v in type_cove_interval.items()]):
plt.text(rect.get_x() + rect.get_width() / 2, rect.get_height()+5, str(v),
fontweight='bold', ha='center', va='bottom')
plt.tight_layout()
plt.show()
```
## Type annotations coverage for source code files
```
f = plt.bar([i for i, v in f_type_cove_interval.items()], [v[1] for i, v in f_type_cove_interval.items()])
plt.xlabel("Percentage of type annotation coverage")
plt.ylabel("No. of source code files")
for rect, v in zip(f.patches, [v[1] for i, v in f_type_cove_interval.items()]):
plt.text(rect.get_x() + rect.get_width() / 2, rect.get_height()+5, str(v),
fontweight='bold', ha='center', va='bottom')
plt.tight_layout()
plt.show()
```
| github_jupyter |
```
d
```
## Import Required Libraries
```
import glob
import numpy as np
import pandas as pd
EXPERIMENT_ID = '1005'
experiment_dir = 'outputs/experiment_{}/'.format(EXPERIMENT_ID)
results_path = experiment_dir + 'results.npy'
schedules_path = experiment_dir + 'schedulesWithMakespan.npy'
log_paths = glob.glob(experiment_dir + '*.log')
```
## Read log files, and results
```
# logs = pd.concat([pd.read_csv(log_file) for log_file in log_paths], axis=0)
result_columns = ['batch_id', 'makespan_mean', 'cum_makespan_mean', 'time', 'cum_time']
results = pd.DataFrame(np.load(results_path), columns=result_columns)
with open(schedules_path, 'rb') as f:
batch_schedules = np.load(f)
batch_makespans = np.load(f)
def convert_schedules_np_to_df(batch_schedules, batch_makespans):
"""Convert batch schedules from numpy array to dataframe"""
n_batch_instance, batch_size, n_nodes, _ = batch_schedules.shape
schedules = batch_schedules.reshape((n_batch_instance*batch_size, n_nodes, 2))
# remove machine-to-machine pair
schedules = schedules[
np.not_equal(
schedules[:,:,:1], schedules[:,:,1:]
)[:,:,-1]
].reshape(schedules.shape[0], -1, schedules.shape[-1]).astype(str)
n_jobs = schedules.shape[1]
makespans = batch_makespans.reshape((n_batch_instance*batch_size,1))
df_schedules = pd.DataFrame(
np.concatenate([
np.apply_along_axis(
' -> '.join, 0, [schedules[:,:,0], schedules[:,:,1]]),
makespans
], axis=1),
columns= list(map(lambda v: "J_"+str(v), range(n_jobs)))+['makespan']
)
return df_schedules
df_schedules = convert_schedules_np_to_df(batch_schedules, batch_makespans)
df_schedules.head(10)
makespans.shape, schedules.shape
np.apply_along_axis(
' -> '.join, 0, [schedules[:,:,0], schedules[:,:,1]]).shape
np.concatenate([
np.apply_along_axis(
' -> '.join, 0, [schedules[:,:,0], schedules[:,:,1]]),
makespans
], axis=1)
#"V_".join(list(range(n_nodes)))
list(map(lambda v: "V_"+str(v), range(1,n_nodes+1)))+['makespan']
"V_".join(map(str, range(1, n_nodes+1)))
makespans.shape
batch_makespans
# r = best_schedules.reshape((64, 7, 2))
# #np.transpose(r, (0,2,1))
# r = r.astype(str)
# r[:,:,0]# + "," + r[:,:,1]
# #np.core.defchararray.add(r[:,:,0], r[:,:,1])
# # list(zip(r[:,:,0], r[:,:,1]))
# # numpy.char.join(r[:,:,0], r[:,:,1])
#best_makespans.shape,
a = np.arange(6).reshape((3, 2))
np.chararray((3, 3))
path = 'outputs/experiment_{}/schedulesWithMakespan.npy'.format(experiment_id)
with open(path, 'rb') as f:
best_schedules = np.load(f)
best_makespans = np.load(f)
path = 'outputs/experiment_{}/results.npy'.format(experiment_id)
with open(path, 'rb') as f:
results = np.load(f)
log_path = 'outputs/experiment_{}/results.npy'.format(experiment_id)
_columns = ['batch_id', 'makespan_mean', 'cum_makespan_mean', 'time', 'cum_time']
df = pd.DataFrame(results, columns=_columns)
df
```
# Modules: Datasets and Graphs
```
import tensorflow as tf
import tensorflow as tf
from tensorflow.keras.layers import Layer
from msp.datasets import make_data, load_sample_data, make_sparse_data
from msp.graphs import MSPGraph, MSPSparseGraph
from msp.models.encoders import GGCNEncoder
from msp.models.decoders import AttentionDecoder
%load_ext autoreload
%autoreload 2
dataset = make_sparse_data(40, msp_size=(2,5), random_state=2021)
type_ = tf.constant([[[4,3,6,1]],[[9,3,6,1]]])
# tf.slice(type_, [0,0,2], [-1,-1,-1])
temp = tf.zeros(type_.shape)
cur_node = tf.constant([[2],[1]])
tf.tensor_scatter_nd_update(temp, indices, [1,1])
self._visited = tf.tensor_scatter_nd_update(
tf.squeeze(temp, axis=-2),
tf.concat([tf.reshape(tf.range(batch_size, dtype=tf.int64),cur_node.shape), cur_node], axis=1),
tf.ones((batch_size,), dtype=self._visited.dtype)
)[:,tf.newaxis,:]
indices = tf.constant([[2],[1]])
tf.gather(tf.squeeze(type_), indices, batch_dims=1)
tf.squeeze(type_)
batch_size = 2
batch_dataset = dataset.batch(batch_size)
inputs = list(batch_dataset.take(1))[0]
inputs
```
# Modules: Layers and Encoders
```
units = 6
layers = 2
ecoder_model = GGCNEncoder(units, layers)
embedded_inputs = ecoder_model(inputs)
embedded_inputs.node_embed
embedded_inputs.edge_embed
```
# Modules: Decoder
```
model = AttentionDecoder(units, aggregation_graph='mean', n_heads=3)
makespan, ll, pi = model(embedded_inputs)
pi
```
# All together
```
n_instances = 640
instance_size = (2, 8)
batch_size = 64
units = 64 # embedding dims
layers = 2
dataset = make_sparse_data(n_instances, msp_size=instance_size, random_state=2021)
batch_dataset = dataset.batch(batch_size)
inputs = list(batch_dataset.take(1))[0]
ecoder_model = GGCNEncoder(units, layers)
embedded_inputs = ecoder_model(inputs)
n_heads = 8
model = AttentionDecoder(units, aggregation_graph='mean', n_heads=n_heads)
makespan, ll, pi = model(embedded_inputs)
pi
inputs.adj_matrix[0]
tf.math.reduce_all(tf.math.equal(inputs.adj_matrix[1], inputs.adj_matrix[2]))
import sys
import numpy
numpy.set_printoptions(threshold=sys.maxsize)
print(inputs.adj_matrix[0].numpy())
print(inputs.adj_matrix[7].numpy())
from typing import NamedTuple
from tensorflow import Tensor
class MSPState:
def __init__(self, inputs):
""" """
self.adj_matrix = inputs.adj_matrix
self.node_embed = inputs.node_embed
batch_size, num_nodes, node_embed_dims = self.node_embed.shape
self._first_node = tf.zeros((batch_size,1), dtype=tf.int64)
self._last_node = self._first_node
self._visited = tf.zeros((batch_size,1,num_nodes), dtype=tf.int64)
self._makespan = tf.zeros((batch_size,1))
self.i = tf.zeros(1, dtype=tf.int64) # # Vector with length num_steps
self.ids = tf.range(5, delta=1, dtype=tf.int64)[:, None] # # Add steps dimension
#self._step_num = tf.zeros(1, dtype=tf.int64)
@property
def first_node(self):
return self._first_node
# self._node_embed = None
# self._edge_embed = None
# self._adj_matrix = None
# self._first_job = None
# self._last_job = None
# @property
# def node_embed(self):
# return self._node_embed
# @node_embed.setter
# def node_embed(self, node_embed):
# self._node_embed = node_embed
# @property
# def adj_matrix(self):
# return self._adj_matrix
# @adj_matrix.setter
# def adj_matrix(self, adj_matrix):
# self._adj_matrix = adj_matrix
# def initialize(self, inputs):
# self.adj_matrix = inputs.adj_matrix
# return self
class MSPSolution:
def __init__(self):
pass
# sequence: Tensor
# makespan: Tensor
# a: int
import torch
torch.arange(5, dtype=torch.int64)[:, None], tf.range(5, delta=1, dtype=tf.int64)[:, None]
class MSPSparseGraph2(NamedTuple):
""" """
adj_matrix: Tensor
node_features: Tensor
edge_features: Tensor
alpha: Tensor
@property
def num_node(self):
return self.adj_matrix.shape[-2]
from collections import namedtuple
class A(NamedTuple):
a: int
@property
def num(self):
return self.a
# jane._replace(a=26)
class C(
NamedTuple(
'SSSSS',
A._field_types.items()
),
A
):
pass
class B(
NamedTuple(
'SSSB',
[
*A._field_types.items(),
('weight', int)
]
),
A
):
@property
def h(self):
return self.weight
# class B(namedtuple('SSSB', [*A._fields, "weight"]), A):
# #[*A._fields, "weight"]
# @property
# def h(self):
# return self.weight
# def __getitem__(self, key):
# return self[key]
#B(A(a=90)._asdict(), weight=90)
#B(**A(a=9090)._asdict(), weight=90).num
#Ba = 2343, weight=23).num
#dir(B)
#C(a=90)
B(*(34,), weight=34)
list(A._field_types.items())
A._field_types
def fun(a):
print(a)
fun(**inputs._asdict())
[*A._field_types, ('v','d')]
# for attr, val in inputs._asdict().items():
# print(attr, val)
from typing import NamedTuple
class AttentionModelFixed(NamedTuple):
"""
Context for AttentionModel decoder that is fixed during decoding so can be precomputed/cached
This class allows for efficient indexing of multiple Tensors at once
"""
node_embeddings: torch.Tensor
context_node_projected: torch.Tensor
glimpse_key: torch.Tensor
glimpse_val: torch.Tensor
logit_key: torch.Tensor
def __getitem__(self, key):
if tf.is_tensor(key) or isinstance(key, slice):
return AttentionModelFixed(
node_embeddings=self.node_embeddings[key],
context_node_projected=self.context_node_projected[key],
glimpse_key=self.glimpse_key[:, key], # dim 0 are the heads
glimpse_val=self.glimpse_val[:, key], # dim 0 are the heads
logit_key=self.logit_key[key]
)
return super(AttentionModelFixed, self).__getitem__(key)
"""
"""
import math
import tensorflow as tf
import tensorflow_probability as tfp
from tensorflow.keras import Model
from msp.layers import GGCNLayer
from msp.graphs import MSPSparseGraph
from msp.solutions import MSPState
class AttentionDecoder(Model):
def __init__(self,
units,
*args,
activation='relu',
aggregation_graph='mean',
n_heads=2, # make it 8
mask_inner=True,
tanh_clipping=10,
decode_type='sampling',
extra_logging=False,
**kwargs):
""" """
super(AttentionDecoder, self).__init__(*args, **kwargs)
self.aggregation_graph = aggregation_graph
self.n_heads = n_heads
self.mask_inner = mask_inner
self.tanh_clipping = tanh_clipping
self.decode_type = decode_type
self.extra_logging = extra_logging
embedding_dim = units
self.W_placeholder = self.add_weight(shape=(2*embedding_dim,),
initializer='random_uniform', #Placeholder should be in range of activations (think)
name='W_placeholder',
trainable=True)
graph_embed_shape = tf.TensorShape((None, units))
self.fixed_context_layer = tf.keras.layers.Dense(units, use_bias=False)
self.fixed_context_layer.build(graph_embed_shape)
# For each node we compute (glimpse key, glimpse value, logit key) so 3 * embedding_dim
project_node_embeddings_shape = tf.TensorShape((None, None, None, units))
self.project_node_embeddings = tf.keras.layers.Dense(3*units, use_bias=False)
self.project_node_embeddings.build(project_node_embeddings_shape)
#
# Embedding of first and last node
step_context_dim = 2*units
project_step_context_shape = tf.TensorShape((None, None, step_context_dim))
self.project_step_context = tf.keras.layers.Dense(embedding_dim, use_bias=False)
self.project_step_context.build(project_step_context_shape)
assert embedding_dim % n_heads == 0
# Note n_heads * val_dim == embedding_dim so input to project_out is embedding_dim
project_out_shape = tf.TensorShape((None, None, 1, embedding_dim))
self.project_out = tf.keras.layers.Dense(embedding_dim, use_bias=False)
self.project_out.build(project_out_shape)
# self.context_layer = tf.keras.layers.Dense(units, use_bias=False)
# self.mha_layer = None
# dynamic router
def call(self, inputs, training=False, return_pi=False):
""" """
state = MSPState(inputs)
node_embedding = inputs.node_embed
# AttentionModelFixed(node_embedding, fixed_context, *fixed_attention_node_data)
fixed = self._precompute(node_embedding)
# for i in range(num_steps):
# i == 0 should be machine
# AttentionCell(inputs, states)
outputs = []
sequences = []
# i = 0
while not state.all_finished():
# B x 1 x V
# Get log probabilities of next action
log_p, mask = self._get_log_p(fixed, state)
selected = self._select_node(
tf.squeeze(tf.exp(log_p), axis=-2), tf.squeeze(mask, axis=-2)) # Squeeze out steps dimension
state.update(selected)
outputs.append(log_p[:, 0, :])
sequences.append(selected)
# if i == 1:
# break
# i+=1
_log_p, pi = tf.stack(outputs, axis=1), tf.stack(sequences, axis=1)
if self.extra_logging:
self.log_p_batch = _log_p
self.log_p_sel_batch = tf.gather(tf.squeeze(_log_p,axis=-2), pi, batch_dims=1)
# # Get predicted costs
# cost, mask = self.problem.get_costs(nodes, pi)
mask = None
###################################################
# Need Clarity #############################################################
# loglikelihood
ll = self._calc_log_likelihood(_log_p, pi, mask)
## Just for checking
return_pi = True
if return_pi:
return state.makespan, ll, pi
return state.makespan, ll
def _precompute(self, node_embedding, num_steps=1):
graph_embed = self._get_graph_embed(node_embedding)
fixed_context = self.fixed_context_layer(graph_embed)
# fixed context = (batch_size, 1, embed_dim) to make broadcastable with parallel timesteps
fixed_context = tf.expand_dims(fixed_context, axis=-2)
glimpse_key_fixed, glimpse_val_fixed, logit_key_fixed = tf.split(
self.project_node_embeddings(tf.expand_dims(node_embedding, axis=-3)),
num_or_size_splits=3,
axis=-1
)
# No need to rearrange key for logit as there is a single head
fixed_attention_node_data = (
self._make_heads(glimpse_key_fixed, num_steps),
self._make_heads(glimpse_val_fixed, num_steps),
logit_key_fixed
)
return AttentionModelFixed(node_embedding, fixed_context, *fixed_attention_node_data)
def _get_graph_embed(self, node_embedding):
""" """
if self.aggregation_graph == "sum":
graph_embed = tf.reduce_sum(node_embedding, axis=-2)
elif self.aggregation_graph == "max":
graph_embed = tf.reduce_max(node_embedding, axis=-2)
elif self.aggregation_graph == "mean":
graph_embed = tf.reduce_mean(node_embedding, axis=-2)
else: # Default: dissable graph embedding
graph_embed = tf.reduce_sum(node_embedding, axis=-2) * 0.0
return graph_embed
def _make_heads(self, v, num_steps=None):
assert num_steps is None or v.shape[1] == 1 or v.shape[1] == num_steps
batch_size, _, num_nodes, embed_dims = v.shape
num_steps = num_steps if num_steps else 1
head_dims = embed_dims//self.n_heads
# M x B x N x V x (H/M)
return tf.transpose(
tf.broadcast_to(
tf.reshape(v, shape=[batch_size, v.shape[1], num_nodes, self.n_heads, head_dims]),
shape=[batch_size, num_steps, num_nodes, self.n_heads, head_dims]
),
perm=[3, 0, 1, 2, 4]
)
def _get_log_p(self, fixed, state, normalize=True):
# Compute query = context node embedding
# B x 1 x H
query = fixed.context_node_projected + \
self.project_step_context(self._get_parallel_step_context(fixed.node_embeddings, state))
# Compute keys and values for the nodes
glimpse_K, glimpse_V, logit_K = self._get_attention_node_data(fixed, state)
# Compute the mask, for masking next action based on previous actions
mask = state.get_mask()
graph_mask = state.get_graph_mask()
# Compute logits (unnormalized log_p)
log_p, glimpse = self._one_to_many_logits(query, glimpse_K, glimpse_V, logit_K, mask, graph_mask)
# [B x N x V]
# log-softmax activation function so that we get log probabilities over actions
if normalize:
log_p = tf.nn.log_softmax(log_p/1.0, axis=-1)
assert not tf.reduce_any(tf.math.is_nan(log_p)), "Log probabilities over the nodes should be defined"
return log_p, mask
def _get_parallel_step_context(self, node_embedding, state, from_depot=False):
"""
Returns the context per step, optionally for multiple steps at once
(for efficient evaluation of the model)
"""
# last_node at time t
last_node = state.get_current_node()
batch_size, num_steps = last_node.shape
if num_steps == 1: # We need to special case if we have only 1 step, may be the first or not
if state.i.numpy()[0] == 0:
# First and only step, ignore prev_a (this is a placeholder)
# B x 1 x 2H
return tf.broadcast_to(self.W_placeholder[None, None, :],
shape=[batch_size, 1, self.W_placeholder.shape[-1]])
else:
return tf.concat(
[
tf.gather(node_embedding,state.first_node,batch_dims=1),
tf.gather(node_embedding,last_node,batch_dims=1)
],
axis=-1
)
# print('$'*20)
# node_embedding = torch.from_numpy(node_embedding.numpy())
# f = torch.from_numpy(state.first_node.numpy())
# l = torch.from_numpy(state.last_node.numpy())
# GG = node_embedding\
# .gather(
# 1,
# torch.cat((f, l), 1)[:, :, None].expand(batch_size, 2, node_embedding.size(-1))
# ).view(batch_size, 1, -1)
# print(GG)
# print('$'*20)
# ##############################################
# # PENDING
# ##############################################
# pass
def _get_attention_node_data(self, fixed, state):
return fixed.glimpse_key, fixed.glimpse_val, fixed.logit_key
def _one_to_many_logits(self, query, glimpse_K, glimpse_V, logit_K, mask, graph_mask=None):
batch_size, num_steps, embed_dim = query.shape
query_size = key_size = val_size = embed_dim // self.n_heads
# M x B x N x 1 x (H/M)
# Compute the glimpse, rearrange dimensions to (n_heads, batch_size, num_steps, 1, key_size)
glimpse_Q = tf.transpose(
tf.reshape(
query, # B x 1 x H
shape=[batch_size, num_steps, self.n_heads, 1, query_size]
),
perm=[2, 0, 1, 3, 4]
)
# [M x B x N x 1 x (H/M)] X [M x B x N x (H/M) x V] = [M x B x N x 1 x V]
# Batch matrix multiplication to compute compatibilities (n_heads, batch_size, num_steps, graph_size)
compatibility = tf.matmul(glimpse_Q, tf.transpose(glimpse_K, [0,1,2,4,3])) / math.sqrt(query_size)
mask_temp = tf.cast(tf.broadcast_to(mask[None, :, :, None, :], shape=compatibility.shape), dtype=tf.double)
compatibility = tf.cast(compatibility, dtype=tf.double) + (mask_temp * -1e9)
graph_mask_temp = tf.cast(tf.broadcast_to(graph_mask[None, :, :, None, :], shape=compatibility.shape), dtype=tf.double)
compatibility = tf.cast(compatibility, dtype=tf.double) + (graph_mask_temp * -1e9)
compatibility = tf.cast(compatibility, dtype=tf.float32)
# compatibility[tf.broadcast_to(mask[None, :, :, None, :], shape=compatibility.shape)] = -1e10
# compatibility[tf.broadcast_to(graph_mask[None, :, :, None, :], shape=compatibility.shape)] = -1e10
# attention weights a(c,j):
attention_weights = tf.nn.softmax(compatibility, axis=-1)
# [M x B x N x 1 x V] x [M x B x N x V x (H/M)] = [M x B x N x 1 x (H/M)]
heads = tf.matmul(attention_weights, glimpse_V)
# B x N x 1 x H
# Project to get glimpse/updated context node embedding (batch_size, num_steps, embedding_dim)
glimpse = self.project_out(
tf.reshape(
tf.transpose(heads, perm=[1, 2, 3, 0, 4]),
shape=[batch_size, num_steps, 1, self.n_heads*val_size]
)
)
# B x N x 1 x H
# Now projecting the glimpse is not needed since this can be absorbed into project_out
# final_Q = self.project_glimpse(glimpse)
final_Q = glimpse
# [B x N x 1 x H] x [B x 1 x H x V] = [B x N x 1 x V] --> [B x N x V] (Squeeze)
# Batch matrix multiplication to compute logits (batch_size, num_steps, graph_size)
# logits = 'compatibility'
logits = tf.squeeze(tf.matmul(final_Q, tf.transpose(logit_K, perm=[0,1,3,2])),
axis=-2) / math.sqrt(final_Q.shape[-1])
logits = logits + ( tf.cast(graph_mask, dtype=tf.float32) * -1e9)
logits = tf.math.tanh(logits) * self.tanh_clipping
logits = logits + ( tf.cast(mask, dtype=tf.float32) * -1e9)
# logits[graph_mask] = -1e10
# logits = torch.tanh(logits) * self.tanh_clipping
# logits[mask] = -1e10
return logits, tf.squeeze(glimpse, axis=-2)
def _select_node(self, probs, mask):
assert tf.reduce_all(probs == probs) == True, "Probs should not contain any nans"
if self.decode_type == "greedy":
selected = tf.math.argmax(probs, axis=1)
assert not tf.reduce_any(tf.cast(tf.gather_nd(mask, tf.expand_dims(selected, axis=-1), batch_dims=1), dtype=tf.bool)), "Decode greedy: infeasible action has maximum probability"
elif self.decode_type == "sampling":
dist = tfp.distributions.Multinomial(total_count=1, probs=probs)
selected = tf.argmax(dist.sample(), axis=1)
# Check if sampling went OK
while tf.reduce_any(tf.cast(tf.gather_nd(mask, tf.expand_dims(selected, axis=-1), batch_dims=1), dtype=tf.bool)):
print('Sampled bad values, resampling!')
selected = tf.argmax(dist.sample(), axis=1)
else:
assert False, "Unknown decode type"
return selected
def _calc_log_likelihood(self, _log_p, a, mask):
# Get log_p corresponding to selected actions
batch_size, steps_count = a.shape
indices = tf.concat([
tf.expand_dims(tf.broadcast_to(tf.range(steps_count, dtype=tf.int64), shape=a.shape), axis=-1),
tf.expand_dims(a, axis=-1)],
axis=-1
)
log_p = tf.gather_nd(_log_p, indices, batch_dims=1)
# _log_p = torch.from_numpy(_log_p.numpy())
# a = torch.from_numpy(a.numpy())
# AA = _log_p.gather(2, a.unsqueeze(-1)).squeeze(-1)
# print(AA)
# print(log_p)
# print('DONE')
# Get log_p corresponding to selected actions
# log_p = tf.gather(tf.squeeze(_log_p,axis=-2), a, batch_dims=1) #_log_p.gather(2, a.unsqueeze(-1)).squeeze(-1)
# Optional: mask out actions irrelevant to objective so they do not get reinforced
if mask is not None:
log_p[mask] = 0
# Why??????
# assert (log_p > -1000).data.all(), "Logprobs should not be -inf, check sampling procedure!"
# Calculate log_likelihood
return tf.reduce_sum(log_p, axis=1) # log_p.sum(1)
# import numpy as np
# # x = tf.constant([[True, True], [False, False]])
# # assert not tf.reduce_any(x), "asdf"
# tf.keras.losses.SparseCategoricalCrossentropy(
# from_logits=True, reduction='none')([]).dtype
# t = torch.tensor([[[2,3],[5,2]], [[6,1],[0,4]]])
# indices = torch.tensor([[[0, 0]],[[0, 0]]])
# torch.gather(t, 1, indices)
# # t = tf.constant([[[2,3],[5,2]], [[6,1],[0,4]]])
# # indices = tf.constant([[0,0,0]])
# # tf.gather_nd(t, indices=indices).numpy()
# # #tf.gather_nd()
# # t = tf.constant([[[1,0],[1,0]], [[0,1],[1,1]]])
# # t = embedded_inputs.adj_matrix
# # indices = tf.constant([[0], [0]])
# # indices = tf.expand_dims(indices, axis=-1)
# # ~tf.cast(tf.gather_nd(t, indices=indices, batch_dims=1), tf.bool)
# a = tf.constant([-3.0,-1.0, 0.0,1.0,3.0], dtype = tf.float32)
# b = tf.keras.activations.tanh(a)
# c = tf.math.tanh(a)
# b.numpy(), c.numpy()
model = AttentionDecoder(6, aggregation_graph='mean', n_heads=3)
makespan, ll, pi = model(embedded_inputs)
pi
tf.range(4, dtype=tf.int64)
indices = [[[1, 0]], [[0, 1]]]
params = [[['a0', 'b0'], ['c0', 'd0']],
[['a1', 'b1'], ['c1', 'd1']]]
#output = [['c0'], ['b1']]
tf.gather
a = tf.Tensor(
[[[0,1]
[1,2]],
[[0,0]
# [1 1]]], shape=(2, 2, 2), dtype=int64)
# tf.Tensor(
# [[[-1.0886807e+00 -1.2036482e+00 -1.0126851e+00]
# [-9.8199100e+00 -1.0000000e+09 -5.4357959e-05]]
# [[-5.4339290e-01 -1.6555539e+00 -1.4773605e+00]
# [-1.0000000e+09 -7.7492166e-01 -6.1755723e-01]]], shape=(2, 2, 3), dtype=float32)
embeddings.gather(
1,
torch.cat((state.first_a, current_node), 1)[:, :, None].expand(batch_size, 2, embeddings.size(-1))
).view(batch_size, 1, -1)
_log_p, pi#tf.expand_dims(pi,axis=-1)
_log_p
tf.gather(tf.squeeze(_log_p,axis=-2), pi, batch_dims=1)
_log_p = torch.Tensor(_log_p.numpy())
pi = torch.from_numpy(tf.cast(pi, dtype=tf.int64).numpy())
log_p = _log_p.gather(2, pi.unsqueeze(-1)).squeeze(-1)
log_p.sum(1)
log_p
# pi.unsqueeze(-1)
# tf.cast(pi, dtype=tf.int64)
_log_p
# seq = [torch.Tensor(i.numpy()) for i in seq]
# torch.stack(seq, 1)
pi
last_node = tf.constant([[0], [1]], dtype=tf.int64)
cur_node = selected[:, tf.newaxis]
cur_node #cur_node
tf.gather_nd(
tf.gather_nd(embedded_inputs.edge_features, tf.concat([last_node, cur_node], axis=1), batch_dims=1),
tf.reshape(tf.range(2), shape=(2,1)), # feature_indexfor processing time
batch_dims=0
)
A = tf.gather_nd(embedded_inputs.edge_features, tf.concat([last_node, cur_node], axis=1), batch_dims=1)[:,0:1]
A
tf.reshape(tf.range(2), shape=(2,1))
embedded_inputs.node_features[:,]
cur_node[:,tf.newaxis,:] #embedded_inputs.node_features[:, ]
tf.gather_nd(embedded_inputs.edge_features, tf.concat([last_node, cur_node], axis=1), batch_dims=1)[:,0:1] + \
tf.gather_nd(embedded_inputs.node_features, cur_node, batch_dims=1)[:,0:1]
cur_node
tf.rank(tf.ones(30,10))
t = tf.constant([[[1, 1, 1, 4], [2, 2, 2, 4]], [[3, 3, 3, 4], [4, 4, 4, 4]]])
tf.rank(t)
_visited = tf.zeros((batch_size,1,3), dtype=tf.uint8)
_visited
cur_node #[:,:,None] #, tf.ones([2,1,1])
tf.concat([tf.reshape(tf.range(2, dtype=tf.int64),cur_node.shape), cur_node], axis=1)
_visited.dtype
batch_size, _, _ = _visited.shape
tf.tensor_scatter_nd_update(
tf.squeeze(_visited, axis=-2),
tf.concat([tf.reshape(tf.range(batch_size, dtype=tf.int64),cur_node.shape), cur_node], axis=1),
tf.ones((batch_size,), dtype=_visited.dtype)
)[:,tf.newaxis,:]
#_visited, cur_node[:,:,None], tf.ones([2,1,1], dtype=tf.uint8))
cur_node = selected[:, tf.newaxis]
cur_node #cur_node
tf.concat([cur_node, cur_node], axis=1)
tf.gather_nd(
tf.gather_nd(embedded_inputs.edge_features, tf.concat([cur_node, cur_node], axis=1), batch_dims=1),
tf.zeros((2,1), dtype=tf.int32)
)
tf.gather_nd(embedded_inputs.edge_features, tf.concat([cur_node, cur_node], axis=1), batch_dims=1)
tf.zeros((2,), dtype=tf.int32)
tf.range(2)
embedded_inputs.edge_features
probs_ = tf.squeeze(tf.exp(log_p), axis=-2)
tf.squeeze(mask, axis=-2)
embedded_inputs.node_features
probs = torch.Tensor(log_p.numpy()).exp()[:, 0, :]
mask_ = torch.Tensor(mask.numpy())[:, 0, :]
probs, probs.multinomial(1) #.squeeze(1)
# _, selected = probs.max(1)
# assert not mask_.gather(1, selected.unsqueeze(
# -1)).data.any(), "Decode greedy: infeasible action has maximum probability"
# mask_.gather(1, selected.unsqueeze(-1)) #.data.any()
probs_
probs.multinomial(1)
# p = [.2, .3, .5]
# dist = tfp.distributions.Multinomial(total_count=1, probs=probs_)
p = [[.1, .2, .7], [.3, .3, .4]] # Shape [2, 3]
dist = tfp.distributions.Multinomial(total_count=1, probs=probs_)
selected = tf.argmax(dist.sample(), axis=1)
selected = tf.math.argmax(tf.squeeze(tf.exp(log_p), axis=-2), axis=1)
assert not tf.reduce_any(tf.cast(tf.gather_nd(mask_, tf.expand_dims(selected, axis=-1), batch_dims=1), dtype=tf.bool)), "Decode greedy: infeasible action has maximum probability"
tf.expand_dims(selected, axis=-1), mask_,
probs_
tf.broadcast_to(mask[None, :, None, :, :], shape=compatibility.shape)
compatibility
compatibility[graph_mask[None, :, :, None, :].expand_as(compatibility)] = -1e10
def _make_heads(v, num_steps=None):
assert num_steps is None or v.size(1) == 1 or v.size(1) == num_steps
n_heads = 2
return (
v.contiguous().view(v.size(0), v.size(1), v.size(2), n_heads, -1)
.expand(v.size(0), v.size(1) if num_steps is None else num_steps, v.size(2), n_heads, -1)
.permute(3, 0, 1, 2, 4) # (n_heads, batch_size, num_steps, graph_size, head_dim)
)
from torch import nn
W_placeholder = nn.Parameter(torch.Tensor(2 * 6))
W_placeholder[None, None, :].expand(2, 1, W_placeholder.size(-1))
tf.ones((3,4)).dtype
_make_heads2(A, num_steps=4)
_make_heads2(A, num_steps=2)
A.shape
tf.broadcast_to(A, tf.TensorShape([-1, 4, None, None]))
tf.split(
tf.matmul(tf.ones((5,3))[None,:,:], tf.ones((3,3*3))),
3,
axis=-1
)
tf.matmul(tf.ones((5,3))[None,:,:], tf.ones((3,3*3)))
import torch
embeddings = torch.Tensor(embedded_inputs.node_features.numpy())
torch.cat((state.first_a, current_node), 1)[:, :, None]
embedded_inputs.node_features
import torch
from torch import nn
W_placeholder = nn.Parameter(torch.Tensor(2 * 5))
W_placeholder[None, None, :].expand(batch_size, 1, W_placeholder.size(-1)).shape
tf.constant(W_placeholder[None, None, :].detach().numpy()).shape[-1]
tf.broadcast_to?
W_placeholder[None, None, :].shape
self.w = self.add_weight(shape=(input_shape[-1], self.units),
initializer='random_normal',
trainable=True)
tf.reduce_mean(embedded_inputs.node_features, axis=-2)
import torch
torch.Tensor(embedded_inputs.node_features.numpy()).max(1)[0]
embedded_inputs.node_features.numpy()
"""
"""
import tensorflow as tf
from tensorflow.keras import Model
from msp.layers import GGCNLayer
from msp.graphs import MSPSparseGraph
from msp.solutions import MSPState
class AttentionDecoder(Model):
def __init__(self,
units,
*args,
activation='relu',
aggregation_graph='mean',
n_heads=8,
**kwargs):
""" """
super(AttentionDecoder, self).__init__(*args, **kwargs)
self.aggregation_graph = aggregation_graph
self.n_heads = n_heads
embedding_dim = units
self.W_placeholder = self.add_weight(shape=(2*embedding_dim,),
initializer='random_uniform', #Placeholder should be in range of activations (think)
trainable=True)
graph_embed_shape = tf.TensorShape((None, units))
self.fixed_context_layer = tf.keras.layers.Dense(units, use_bias=False)
self.fixed_context_layer.build(graph_embed_shape)
# For each node we compute (glimpse key, glimpse value, logit key) so 3 * embedding_dim
project_node_embeddings_shape = tf.TensorShape((None, None, None, units))
self.project_node_embeddings = tf.keras.layers.Dense(3*units, use_bias=False)
self.project_node_embeddings.build(project_node_embeddings_shape)
#
# Embedding of first and last node
step_context_dim = 2*units
project_step_context_shape = tf.TensorShape((None, None, step_context_dim))
self.project_step_context = tf.keras.layers.Dense(embedding_dim, use_bias=False)
self.project_step_context.build(project_step_context_shape)
#nn.Linear(step_context_dim, embedding_dim, bias=False)
# self.context_layer = tf.keras.layers.Dense(units, use_bias=False)
# self.mha_layer = None
# dynamic router
# self.initial_layer_1 = tf.keras.layers.Dense(units)
# self.initial_layer_2 = tf.keras.layers.Dense(units)
# self.ggcn_layers = [GGCNLayer(units, activation=activation)
# for _ in range(layers)]
def call(self, inputs, training=False):
""" """
state = MSPState(inputs)
node_embedding = inputs.node_embed
# AttentionModelFixed(node_embedding, fixed_context, *fixed_attention_node_data)
fixed = self._precompute(node_embedding)
return fixed
while not state.all_finished():
log_p, mask = self._get_log_p(fixed, state)
node_embed = inputs.node_features
return self._get_parallel_step_context(node_embed)
def _precompute(self, node_embedding, num_steps=1):
graph_embed = self._get_graph_embed(node_embedding)
fixed_context = self.fixed_context_layer(graph_embed)
# fixed context = (batch_size, 1, embed_dim) to make broadcastable with parallel timesteps
fixed_context = tf.expand_dims(fixed_context, axis=-2)
glimpse_key_fixed, glimpse_val_fixed, logit_key_fixed = tf.split(
self.project_node_embeddings(tf.expand_dims(node_embedding, axis=-3)),
num_or_size_splits=3,
axis=-1
)
return glimpse_key_fixed
# No need to rearrange key for logit as there is a single head
fixed_attention_node_data = (
self._make_heads(glimpse_key_fixed, num_steps),
self._make_heads(glimpse_val_fixed, num_steps),
logit_key_fixed
)
return AttentionModelFixed(node_embedding, fixed_context, *fixed_attention_node_data)
def _get_graph_embed(self, node_embedding):
""" """
if self.aggregation_graph == "sum":
graph_embed = tf.reduce_sum(node_embedding, axis=-2)
elif self.aggregation_graph == "max":
graph_embed = tf.reduce_max(node_embedding, axis=-2)
elif self.aggregation_graph == "mean":
graph_embed = tf.reduce_mean(node_embedding, axis=-2)
else: # Default: dissable graph embedding
graph_embed = tf.reduce_sum(node_embedding, axis=-2) * 0.0
return graph_embed
def _make_heads(self, v, num_steps=None):
assert num_steps is None or v.shape[1] == 1 or v.shape[1] == num_steps
batch_size, _, num_nodes, embed_dims = v.shape
num_steps = num_steps if num_steps else 1
# M x B x N x V x H
return tf.broadcast_to(
tf.broadcast_to(v, shape=[batch_size, num_steps, num_nodes, embed_dims])[None,:,:,:,:],
shape=[self.n_heads, batch_size, num_steps, num_nodes, embed_dims]
)
def _get_log_p(self, fixed, state, normalize=True):
# Compute query = context node embedding
# B x 1 x H
query = fixed.context_node_projected + \
self.project_step_context(self._get_parallel_step_context(fixed.node_embeddings, state))
# Compute keys and values for the nodes
glimpse_K, glimpse_V, logit_K = self._get_attention_node_data(fixed, state)
graph_mask = None
if self.mask_graph:
# Compute the graph mask, for masking next action based on graph structure
graph_mask = state.get_graph_mask() # Pending...........................................
# Compute logits (unnormalized log_p)
log_p, glimpse = self._one_to_many_logits(query, glimpse_K, glimpse_V, logit_K, mask, graph_mask)
def _get_parallel_step_context(self, node_embedding, state, from_depot=False):
"""
Returns the context per step, optionally for multiple steps at once
(for efficient evaluation of the model)
"""
current_node = state.get_current_node()
batch_size, num_steps = current_node.shape
if num_steps == 1: # We need to special case if we have only 1 step, may be the first or not
if self.i.numpy()[0] == 0:
# First and only step, ignore prev_a (this is a placeholder)
# B x 1 x 2H
return tf.broadcast_to(self.W_placeholder[None, None, :],
shape=[batch_size, 1, self.W_placeholder.shape[-1]])
# else:
# return embeddings.gather(
# 1,
# torch.cat((state.first_a, current_node), 1)[:, :, None].expand(batch_size, 2, embeddings.size(-1))
# ).view(batch_size, 1, -1)
def _get_attention_node_data(self, fixed, state):
return fixed.glimpse_key, fixed.glimpse_val, fixed.logit_key
def _one_to_many_logits(self, query, glimpse_K, glimpse_V, logit_K, mask, graph_mask=None):
pass
tf.boolean_mask
MSPSparseGraph._fields
dataset = make_sparse_data(4, msp_size=(1,2), random_state=2021)
graph = list(dataset.take(1))[0]
graph
indices= tf.cast(tf.transpose(
tf.concat([
edge_index,
tf.scatter_nd(
tf.constant([[1],[0]]),
edge_index,
edge_index.shape
)
], axis=-1)
), tf.int64)
indices
adj_matrix = tf.sparse.SparseTensor(
indices= indices,
values = tf.ones([2*edge_index.shape[-1]]),
dense_shape = [3,3]
)
adj_matrix
tf.sparse.to_dense(tf.sparse.reorder(adj_matrix))
reorder
tf.ones([edge_index.shape[-1]])
E = tf.sparse.SparseTensor(indices=[[0, 0], [1, 2], [2, 0]], values=[1, 2], dense_shape=[3, 4])
tf.sparse.to_dense()
tf.ones((2,3))
class GGCNLayer(Layer):
# @validate_hyperparams
def __init__(self,
units,
*args,
activation='relu',
use_bias=True,
normalization='batch',
aggregation='mean',
**kwargs):
""" """
super(GGCNLayer, self).__init__(*args, **kwargs)
self.units = units
self.activation = tf.keras.activations.get(activation)
self.use_bias = use_bias
self.normalization= normalization
self.aggregation = aggregation
def build(self, input_shape):
"""Create the state of the layer (weights)"""
print('Build')
node_features_shape = input_shape.node_features
edge_featues_shape = input_shape.edge_features
embedded_shape = tf.TensorShape((None, self.units))
# _initial_projection_layer (think on it)
with tf.name_scope('node'):
with tf.name_scope('U'):
self.U = tf.keras.layers.Dense(self.units, use_bias=self.use_bias)
self.U.build(node_features_shape)
with tf.name_scope('V'):
self.V = tf.keras.layers.Dense(self.units, use_bias=self.use_bias)
self.V.build(node_features_shape)
with tf.name_scope('norm'):
self.norm_h = {
"batch": tf.keras.layers.BatchNormalization(),
"layer": tf.keras.layers.LayerNormalization()
}.get(self.normalization, None)
if self.norm_h:
self.norm_h.build(embedded_shape)
with tf.name_scope('edge'):
with tf.name_scope('A'):
self.A = tf.keras.layers.Dense(self.units, use_bias=self.use_bias)
self.A.build(tf.TensorShape((None, node_features_shape[-1])))
with tf.name_scope('B'):
self.B = tf.keras.layers.Dense(self.units, use_bias=self.use_bias)
self.B.build(node_features_shape)
with tf.name_scope('C'):
self.C = tf.keras.layers.Dense(self.units, use_bias=self.use_bias)
self.C.build(edge_featues_shape)
with tf.name_scope('norm'):
self.norm_e = {
'batch': tf.keras.layers.BatchNormalization(),
'layer': tf.keras.layers.LayerNormalization(axis=-2)
}.get(self.normalization, None)
if self.norm_e:
self.norm_e.build(embedded_shape)
super().build(input_shape)
def call(self, inputs):
""" """
print('call')
adj_matrix = inputs.adj_matrix
h = inputs.node_features
e = inputs.edge_features
# Edges Featuers
Ah = self.A(h)
Bh = self.B(h)
Ce = self.C(e)
e = self._update_edges(e, [Ah, Bh, Ce])
edge_gates = tf.sigmoid(e)
# Nodes Features
Uh = self.U(h)
Vh = self.V(h)
h = self._update_nodes(
h,
[Uh, self._aggregate(Vh, edge_gates, adj_matrix)]
)
outputs = MSPSparseGraph(adj_matrix, h, e, inputs.alpha)
return inputs
def _update_edges(self, e, transformations:list):
"""Update edges features"""
Ah, Bh, Ce = transformations
e_new = tf.expand_dims(Ah, axis=1) + tf.expand_dims(Bh, axis=2) + Ce
# Normalization
batch_size, num_nodes, num_nodes, hidden_dim = e_new.shape
if self.norm_e:
e_new = tf.reshape(
self.norm_e(
tf.reshape(e_new, [batch_size*num_nodes*num_nodes, hidden_dim])
), e_new.shape
)
# Activation
e_new = self.activation(e_new)
# Skip/residual Connection
e_new = e + e_new
return e_new
def _update_nodes(self, h, transformations:list):
""" """
Uh, aggregated_messages = transformations
h_new = tf.math.add_n([Uh, aggregated_messages])
# Normalization
batch_size, num_nodes, hidden_dim = h_new.shape
if self.norm_h:
h_new = tf.reshape(
self.norm_h(
tf.reshape(h_new, [batch_size*num_nodes, hidden_dim])
), h_new.shape
)
# Activation
h_new = self.activation(h_new)
# Skip/residual Connection
h_new = h + h_new
return h_new
def _aggregate(self, Vh, edge_gates, adj_matrix):
""" """
# Reshape as edge_gates
Vh = tf.broadcast_to(
tf.expand_dims(Vh, axis=1),
edge_gates.shape
)
# Gating mechanism
Vh = edge_gates * Vh
# Enforce graph structure
# mask = tf.broadcast_to(tf.expand_dims(adj_matrix,axis=-1), Vh.shape)
# Vh[~mask] = 0
# message aggregation
if self.aggregation == 'mean':
total_messages = tf.cast(
tf.expand_dims(
tf.math.reduce_sum(adj_matrix, axis=-1),
axis=-1
),
Vh.dtype
)
return tf.math.reduce_sum(Vh, axis=2) / total_messages
elif self.aggregation == 'sum':
return tf.math.reduce_sum(Vh, axis=2)
#
B, V, H = 1, 3, 2
h = tf.ones((B, V, H))
dataset = make_sparse_data(4, msp_size=(1,2), random_state=2020)
graphs = list(dataset.batch(2))
VVh = list(graphs)[0].edge_features
A = list(graphs)[0].adj_matrix
VVh.shape, A.shape
# mask = tf.cast(tf.broadcast_to(
# tf.expand_dims(A, axis=-1),
# VVh.shape
# ), tf.bool)
# VVh[~mask]
ggcn.variables
units = graphs[0].edge_features.shape[-1]
ggcn = GGCNLayer(units=units, use_bias=True, name='GGCN_Layer')
output = ggcn(graphs[0])
output
tf.cast(
tf.expand_dims(
tf.math.reduce_sum(list(graphs)[0].adj_matrix, axis=-1),
axis=-1
),
Vh.dtype
)
tf.expand_dims(
tf.math.reduce_sum(list(graphs)[0].adj_matrix, axis=-1),
axis=-1).dtype
tf.math.reduce_max(graphs[0].edge_features, axis=-2 )[0]
list(graphs)[0].adj_matrix
# ggcn = GGCNLayer(units=units, use_bias=True, name='GGCN_Layer')
# output = ggcn(graphs[0])
# output
graphs = list(make_data(1, msp_size=(1,2), random_state=2021).take(1))[0]
num_edges = 3
A = tf.sparse.SparseTensor(
indices= tf.cast(tf.transpose(graphs['edge_index']), tf.int64),
values = tf.ones([num_edges]),
dense_shape = [3,3]
)
tf.sparse.to_dense(A)
tf.cast(tf.transpose(graphs['edge_index']), tf.int64)
# ggcn.variables
tf.cast(tf.transpose(graphs['edge_index']), tf.int64)
edge_index = tf.cast(tf.transpose(
tf.concat([
graphs['edge_index'],
tf.scatter_nd(
tf.constant([[1],[0]]),
graphs['edge_index'],
graphs['edge_index'].shape
)
], axis=-1)
), tf.int32)
edge_index
model = tf.keras.Sequential([
GGCNLayer(units=units, use_bias=True, name='GGCN_Layer')
])
model.compile(optimizer='sgd', loss='mse')
model.fit(dataset)
model.weights
```
| github_jupyter |
# 序列到序列学习(seq2seq)
:label:`sec_seq2seq`
正如我们在 :numref:`sec_machine_translation`中看到的,
机器翻译中的输入序列和输出序列都是长度可变的。
为了解决这类问题,我们在 :numref:`sec_encoder-decoder`中
设计了一个通用的”编码器-解码器“架构。
本节,我们将使用两个循环神经网络的编码器和解码器,
并将其应用于*序列到序列*(sequence to sequence,seq2seq)类的学习任务
:cite:`Sutskever.Vinyals.Le.2014,Cho.Van-Merrienboer.Gulcehre.ea.2014`。
遵循编码器-解码器架构的设计原则,
循环神经网络编码器使用长度可变的序列作为输入,
将其转换为固定形状的隐状态。
换言之,输入序列的信息被*编码*到循环神经网络编码器的隐状态中。
为了连续生成输出序列的词元,
独立的循环神经网络解码器是基于输入序列的编码信息
和输出序列已经看见的或者生成的词元来预测下一个词元。
:numref:`fig_seq2seq`演示了
如何在机器翻译中使用两个循环神经网络进行序列到序列学习。

:label:`fig_seq2seq`
在 :numref:`fig_seq2seq`中,
特定的“<eos>”表示序列结束词元。
一旦输出序列生成此词元,模型就会停止预测。
在循环神经网络解码器的初始化时间步,有两个特定的设计决定:
首先,特定的“<bos>”表示序列开始词元,它是解码器的输入序列的第一个词元。
其次,使用循环神经网络编码器最终的隐状态来初始化解码器的隐状态。
例如,在 :cite:`Sutskever.Vinyals.Le.2014`的设计中,
正是基于这种设计将输入序列的编码信息送入到解码器中来生成输出序列的。
在其他一些设计中 :cite:`Cho.Van-Merrienboer.Gulcehre.ea.2014`,
如 :numref:`fig_seq2seq`所示,
编码器最终的隐状态在每一个时间步都作为解码器的输入序列的一部分。
类似于 :numref:`sec_language_model`中语言模型的训练,
可以允许标签成为原始的输出序列,
从源序列词元“<bos>”、“Ils”、“regardent”、“.”
到新序列词元
“Ils”、“regardent”、“.”、“<eos>”来移动预测的位置。
下面,我们动手构建 :numref:`fig_seq2seq`的设计,
并将基于 :numref:`sec_machine_translation`中
介绍的“英-法”数据集来训练这个机器翻译模型。
```
import collections
import math
import torch
from torch import nn
from d2l import torch as d2l
```
## 编码器
从技术上讲,编码器将长度可变的输入序列转换成
形状固定的上下文变量$\mathbf{c}$,
并且将输入序列的信息在该上下文变量中进行编码。
如 :numref:`fig_seq2seq`所示,可以使用循环神经网络来设计编码器。
考虑由一个序列组成的样本(批量大小是$1$)。
假设输入序列是$x_1, \ldots, x_T$,
其中$x_t$是输入文本序列中的第$t$个词元。
在时间步$t$,循环神经网络将词元$x_t$的输入特征向量
$\mathbf{x}_t$和$\mathbf{h} _{t-1}$(即上一时间步的隐状态)
转换为$\mathbf{h}_t$(即当前步的隐状态)。
使用一个函数$f$来描述循环神经网络的循环层所做的变换:
$$\mathbf{h}_t = f(\mathbf{x}_t, \mathbf{h}_{t-1}). $$
总之,编码器通过选定的函数$q$,
将所有时间步的隐状态转换为上下文变量:
$$\mathbf{c} = q(\mathbf{h}_1, \ldots, \mathbf{h}_T).$$
比如,当选择$q(\mathbf{h}_1, \ldots, \mathbf{h}_T) = \mathbf{h}_T$时
(就像 :numref:`fig_seq2seq`中一样),
上下文变量仅仅是输入序列在最后时间步的隐状态$\mathbf{h}_T$。
到目前为止,我们使用的是一个单向循环神经网络来设计编码器,
其中隐状态只依赖于输入子序列,
这个子序列是由输入序列的开始位置到隐状态所在的时间步的位置
(包括隐状态所在的时间步)组成。
我们也可以使用双向循环神经网络构造编码器,
其中隐状态依赖于两个输入子序列,
两个子序列是由隐状态所在的时间步的位置之前的序列和之后的序列
(包括隐状态所在的时间步),
因此隐状态对整个序列的信息都进行了编码。
现在,让我们[**实现循环神经网络编码器**]。
注意,我们使用了*嵌入层*(embedding layer)
来获得输入序列中每个词元的特征向量。
嵌入层的权重是一个矩阵,
其行数等于输入词表的大小(`vocab_size`),
其列数等于特征向量的维度(`embed_size`)。
对于任意输入词元的索引$i$,
嵌入层获取权重矩阵的第$i$行(从$0$开始)以返回其特征向量。
另外,本文选择了一个多层门控循环单元来实现编码器。
```
#@save
class Seq2SeqEncoder(d2l.Encoder):
"""用于序列到序列学习的循环神经网络编码器"""
def __init__(self, vocab_size, embed_size, num_hiddens, num_layers,
dropout=0, **kwargs):
super(Seq2SeqEncoder, self).__init__(**kwargs)
# 嵌入层
self.embedding = nn.Embedding(vocab_size, embed_size)
self.rnn = nn.GRU(embed_size, num_hiddens, num_layers,
dropout=dropout)
def forward(self, X, *args):
# 输出'X'的形状:(batch_size,num_steps,embed_size)
X = self.embedding(X)
# 在循环神经网络模型中,第一个轴对应于时间步
X = X.permute(1, 0, 2)
# 如果未提及状态,则默认为0
output, state = self.rnn(X)
# output的形状:(num_steps,batch_size,num_hiddens)
# state[0]的形状:(num_layers,batch_size,num_hiddens)
return output, state
```
循环层返回变量的说明可以参考 :numref:`sec_rnn-concise`。
下面,我们实例化[**上述编码器的实现**]:
我们使用一个两层门控循环单元编码器,其隐藏单元数为$16$。
给定一小批量的输入序列`X`(批量大小为$4$,时间步为$7$)。
在完成所有时间步后,
最后一层的隐状态的输出是一个张量(`output`由编码器的循环层返回),
其形状为(时间步数,批量大小,隐藏单元数)。
```
encoder = Seq2SeqEncoder(vocab_size=10, embed_size=8, num_hiddens=16,
num_layers=2)
encoder.eval()
X = torch.zeros((4, 7), dtype=torch.long)
output, state = encoder(X)
output.shape
```
由于这里使用的是门控循环单元,
所以在最后一个时间步的多层隐状态的形状是
(隐藏层的数量,批量大小,隐藏单元的数量)。
如果使用长短期记忆网络,`state`中还将包含记忆单元信息。
```
state.shape
```
## [**解码器**]
:label:`sec_seq2seq_decoder`
正如上文提到的,编码器输出的上下文变量$\mathbf{c}$
对整个输入序列$x_1, \ldots, x_T$进行编码。
来自训练数据集的输出序列$y_1, y_2, \ldots, y_{T'}$,
对于每个时间步$t'$(与输入序列或编码器的时间步$t$不同),
解码器输出$y_{t'}$的概率取决于先前的输出子序列
$y_1, \ldots, y_{t'-1}$和上下文变量$\mathbf{c}$,
即$P(y_{t'} \mid y_1, \ldots, y_{t'-1}, \mathbf{c})$。
为了在序列上模型化这种条件概率,
我们可以使用另一个循环神经网络作为解码器。
在输出序列上的任意时间步$t^\prime$,
循环神经网络将来自上一时间步的输出$y_{t^\prime-1}$
和上下文变量$\mathbf{c}$作为其输入,
然后在当前时间步将它们和上一隐状态
$\mathbf{s}_{t^\prime-1}$转换为
隐状态$\mathbf{s}_{t^\prime}$。
因此,可以使用函数$g$来表示解码器的隐藏层的变换:
$$\mathbf{s}_{t^\prime} = g(y_{t^\prime-1}, \mathbf{c}, \mathbf{s}_{t^\prime-1}).$$
:eqlabel:`eq_seq2seq_s_t`
在获得解码器的隐状态之后,
我们可以使用输出层和softmax操作
来计算在时间步$t^\prime$时输出$y_{t^\prime}$的条件概率分布
$P(y_{t^\prime} \mid y_1, \ldots, y_{t^\prime-1}, \mathbf{c})$。
根据 :numref:`fig_seq2seq`,当实现解码器时,
我们直接使用编码器最后一个时间步的隐状态来初始化解码器的隐状态。
这就要求使用循环神经网络实现的编码器和解码器具有相同数量的层和隐藏单元。
为了进一步包含经过编码的输入序列的信息,
上下文变量在所有的时间步与解码器的输入进行拼接(concatenate)。
为了预测输出词元的概率分布,
在循环神经网络解码器的最后一层使用全连接层来变换隐状态。
```
class Seq2SeqDecoder(d2l.Decoder):
"""用于序列到序列学习的循环神经网络解码器"""
def __init__(self, vocab_size, embed_size, num_hiddens, num_layers,
dropout=0, **kwargs):
super(Seq2SeqDecoder, self).__init__(**kwargs)
self.embedding = nn.Embedding(vocab_size, embed_size)
self.rnn = nn.GRU(embed_size + num_hiddens, num_hiddens, num_layers,
dropout=dropout)
self.dense = nn.Linear(num_hiddens, vocab_size)
def init_state(self, enc_outputs, *args):
return enc_outputs[1]
def forward(self, X, state):
# 输出'X'的形状:(batch_size,num_steps,embed_size)
X = self.embedding(X).permute(1, 0, 2)
# 广播context,使其具有与X相同的num_steps
context = state[-1].repeat(X.shape[0], 1, 1)
X_and_context = torch.cat((X, context), 2)
output, state = self.rnn(X_and_context, state)
output = self.dense(output).permute(1, 0, 2)
# output的形状:(batch_size,num_steps,vocab_size)
# state[0]的形状:(num_layers,batch_size,num_hiddens)
return output, state
```
下面,我们用与前面提到的编码器中相同的超参数来[**实例化解码器**]。
如我们所见,解码器的输出形状变为(批量大小,时间步数,词表大小),
其中张量的最后一个维度存储预测的词元分布。
```
decoder = Seq2SeqDecoder(vocab_size=10, embed_size=8, num_hiddens=16,
num_layers=2)
decoder.eval()
state = decoder.init_state(encoder(X))
output, state = decoder(X, state)
output.shape, state.shape
```
总之,上述循环神经网络“编码器-解码器”模型中的各层如
:numref:`fig_seq2seq_details`所示。

:label:`fig_seq2seq_details`
## 损失函数
在每个时间步,解码器预测了输出词元的概率分布。
类似于语言模型,可以使用softmax来获得分布,
并通过计算交叉熵损失函数来进行优化。
回想一下 :numref:`sec_machine_translation`中,
特定的填充词元被添加到序列的末尾,
因此不同长度的序列可以以相同形状的小批量加载。
但是,我们应该将填充词元的预测排除在损失函数的计算之外。
为此,我们可以使用下面的`sequence_mask`函数
[**通过零值化屏蔽不相关的项**],
以便后面任何不相关预测的计算都是与零的乘积,结果都等于零。
例如,如果两个序列的有效长度(不包括填充词元)分别为$1$和$2$,
则第一个序列的第一项和第二个序列的前两项之后的剩余项将被清除为零。
```
#@save
def sequence_mask(X, valid_len, value=0):
"""在序列中屏蔽不相关的项"""
maxlen = X.size(1)
mask = torch.arange((maxlen), dtype=torch.float32,
device=X.device)[None, :] < valid_len[:, None]
X[~mask] = value
return X
X = torch.tensor([[1, 2, 3], [4, 5, 6]])
sequence_mask(X, torch.tensor([1, 2]))
```
(**我们还可以使用此函数屏蔽最后几个轴上的所有项。**)如果愿意,也可以使用指定的非零值来替换这些项。
```
X = torch.ones(2, 3, 4)
sequence_mask(X, torch.tensor([1, 2]), value=-1)
```
现在,我们可以[**通过扩展softmax交叉熵损失函数来遮蔽不相关的预测**]。
最初,所有预测词元的掩码都设置为1。
一旦给定了有效长度,与填充词元对应的掩码将被设置为0。
最后,将所有词元的损失乘以掩码,以过滤掉损失中填充词元产生的不相关预测。
```
#@save
class MaskedSoftmaxCELoss(nn.CrossEntropyLoss):
"""带遮蔽的softmax交叉熵损失函数"""
# pred的形状:(batch_size,num_steps,vocab_size)
# label的形状:(batch_size,num_steps)
# valid_len的形状:(batch_size,)
def forward(self, pred, label, valid_len):
weights = torch.ones_like(label)
weights = sequence_mask(weights, valid_len)
self.reduction='none'
unweighted_loss = super(MaskedSoftmaxCELoss, self).forward(
pred.permute(0, 2, 1), label)
weighted_loss = (unweighted_loss * weights).mean(dim=1)
return weighted_loss
```
我们可以创建三个相同的序列来进行[**代码健全性检查**],
然后分别指定这些序列的有效长度为$4$、$2$和$0$。
结果就是,第一个序列的损失应为第二个序列的两倍,而第三个序列的损失应为零。
```
loss = MaskedSoftmaxCELoss()
loss(torch.ones(3, 4, 10), torch.ones((3, 4), dtype=torch.long),
torch.tensor([4, 2, 0]))
```
## [**训练**]
:label:`sec_seq2seq_training`
在下面的循环训练过程中,如 :numref:`fig_seq2seq`所示,
特定的序列开始词元(“<bos>”)和
原始的输出序列(不包括序列结束词元“<eos>”)
拼接在一起作为解码器的输入。
这被称为*强制教学*(teacher forcing),
因为原始的输出序列(词元的标签)被送入解码器。
或者,将来自上一个时间步的*预测*得到的词元作为解码器的当前输入。
```
#@save
def train_seq2seq(net, data_iter, lr, num_epochs, tgt_vocab, device):
"""训练序列到序列模型"""
def xavier_init_weights(m):
if type(m) == nn.Linear:
nn.init.xavier_uniform_(m.weight)
if type(m) == nn.GRU:
for param in m._flat_weights_names:
if "weight" in param:
nn.init.xavier_uniform_(m._parameters[param])
net.apply(xavier_init_weights)
net.to(device)
optimizer = torch.optim.Adam(net.parameters(), lr=lr)
loss = MaskedSoftmaxCELoss()
net.train()
animator = d2l.Animator(xlabel='epoch', ylabel='loss',
xlim=[10, num_epochs])
for epoch in range(num_epochs):
timer = d2l.Timer()
metric = d2l.Accumulator(2) # 训练损失总和,词元数量
for batch in data_iter:
optimizer.zero_grad()
X, X_valid_len, Y, Y_valid_len = [x.to(device) for x in batch]
bos = torch.tensor([tgt_vocab['<bos>']] * Y.shape[0],
device=device).reshape(-1, 1)
dec_input = torch.cat([bos, Y[:, :-1]], 1) # 强制教学
Y_hat, _ = net(X, dec_input, X_valid_len)
l = loss(Y_hat, Y, Y_valid_len)
l.sum().backward() # 损失函数的标量进行“反向传播”
d2l.grad_clipping(net, 1)
num_tokens = Y_valid_len.sum()
optimizer.step()
with torch.no_grad():
metric.add(l.sum(), num_tokens)
if (epoch + 1) % 10 == 0:
animator.add(epoch + 1, (metric[0] / metric[1],))
print(f'loss {metric[0] / metric[1]:.3f}, {metric[1] / timer.stop():.1f} '
f'tokens/sec on {str(device)}')
```
现在,在机器翻译数据集上,我们可以
[**创建和训练一个循环神经网络“编码器-解码器”模型**]用于序列到序列的学习。
```
embed_size, num_hiddens, num_layers, dropout = 32, 32, 2, 0.1
batch_size, num_steps = 64, 10
lr, num_epochs, device = 0.005, 300, d2l.try_gpu()
train_iter, src_vocab, tgt_vocab = d2l.load_data_nmt(batch_size, num_steps)
encoder = Seq2SeqEncoder(len(src_vocab), embed_size, num_hiddens, num_layers,
dropout)
decoder = Seq2SeqDecoder(len(tgt_vocab), embed_size, num_hiddens, num_layers,
dropout)
net = d2l.EncoderDecoder(encoder, decoder)
train_seq2seq(net, train_iter, lr, num_epochs, tgt_vocab, device)
```
## [**预测**]
为了采用一个接着一个词元的方式预测输出序列,
每个解码器当前时间步的输入都将来自于前一时间步的预测词元。
与训练类似,序列开始词元(“<bos>”)
在初始时间步被输入到解码器中。
该预测过程如 :numref:`fig_seq2seq_predict`所示,
当输出序列的预测遇到序列结束词元(“<eos>”)时,预测就结束了。

:label:`fig_seq2seq_predict`
我们将在 :numref:`sec_beam-search`中介绍不同的序列生成策略。
```
#@save
def predict_seq2seq(net, src_sentence, src_vocab, tgt_vocab, num_steps,
device, save_attention_weights=False):
"""序列到序列模型的预测"""
# 在预测时将net设置为评估模式
net.eval()
src_tokens = src_vocab[src_sentence.lower().split(' ')] + [
src_vocab['<eos>']]
enc_valid_len = torch.tensor([len(src_tokens)], device=device)
src_tokens = d2l.truncate_pad(src_tokens, num_steps, src_vocab['<pad>'])
# 添加批量轴
enc_X = torch.unsqueeze(
torch.tensor(src_tokens, dtype=torch.long, device=device), dim=0)
enc_outputs = net.encoder(enc_X, enc_valid_len)
dec_state = net.decoder.init_state(enc_outputs, enc_valid_len)
# 添加批量轴
dec_X = torch.unsqueeze(torch.tensor(
[tgt_vocab['<bos>']], dtype=torch.long, device=device), dim=0)
output_seq, attention_weight_seq = [], []
for _ in range(num_steps):
Y, dec_state = net.decoder(dec_X, dec_state)
# 我们使用具有预测最高可能性的词元,作为解码器在下一时间步的输入
dec_X = Y.argmax(dim=2)
pred = dec_X.squeeze(dim=0).type(torch.int32).item()
# 保存注意力权重(稍后讨论)
if save_attention_weights:
attention_weight_seq.append(net.decoder.attention_weights)
# 一旦序列结束词元被预测,输出序列的生成就完成了
if pred == tgt_vocab['<eos>']:
break
output_seq.append(pred)
return ' '.join(tgt_vocab.to_tokens(output_seq)), attention_weight_seq
```
## 预测序列的评估
我们可以通过与真实的标签序列进行比较来评估预测序列。
虽然 :cite:`Papineni.Roukos.Ward.ea.2002`
提出的BLEU(bilingual evaluation understudy)
最先是用于评估机器翻译的结果,
但现在它已经被广泛用于测量许多应用的输出序列的质量。
原则上说,对于预测序列中的任意$n$元语法(n-grams),
BLEU的评估都是这个$n$元语法是否出现在标签序列中。
我们将BLEU定义为:
$$ \exp\left(\min\left(0, 1 - \frac{\mathrm{len}_{\text{label}}}{\mathrm{len}_{\text{pred}}}\right)\right) \prod_{n=1}^k p_n^{1/2^n},$$
:eqlabel:`eq_bleu`
其中$\mathrm{len}_{\text{label}}$表示标签序列中的词元数和
$\mathrm{len}_{\text{pred}}$表示预测序列中的词元数,
$k$是用于匹配的最长的$n$元语法。
另外,用$p_n$表示$n$元语法的精确度,它是两个数量的比值:
第一个是预测序列与标签序列中匹配的$n$元语法的数量,
第二个是预测序列中$n$元语法的数量的比率。
具体地说,给定标签序列$A$、$B$、$C$、$D$、$E$、$F$
和预测序列$A$、$B$、$B$、$C$、$D$,
我们有$p_1 = 4/5$、$p_2 = 3/4$、$p_3 = 1/3$和$p_4 = 0$。
根据 :eqref:`eq_bleu`中BLEU的定义,
当预测序列与标签序列完全相同时,BLEU为$1$。
此外,由于$n$元语法越长则匹配难度越大,
所以BLEU为更长的$n$元语法的精确度分配更大的权重。
具体来说,当$p_n$固定时,$p_n^{1/2^n}$
会随着$n$的增长而增加(原始论文使用$p_n^{1/n}$)。
而且,由于预测的序列越短获得的$p_n$值越高,
所以 :eqref:`eq_bleu`中乘法项之前的系数用于惩罚较短的预测序列。
例如,当$k=2$时,给定标签序列$A$、$B$、$C$、$D$、$E$、$F$
和预测序列$A$、$B$,尽管$p_1 = p_2 = 1$,
惩罚因子$\exp(1-6/2) \approx 0.14$会降低BLEU。
[**BLEU的代码实现**]如下。
```
def bleu(pred_seq, label_seq, k): #@save
"""计算BLEU"""
pred_tokens, label_tokens = pred_seq.split(' '), label_seq.split(' ')
len_pred, len_label = len(pred_tokens), len(label_tokens)
score = math.exp(min(0, 1 - len_label / len_pred))
for n in range(1, k + 1):
num_matches, label_subs = 0, collections.defaultdict(int)
for i in range(len_label - n + 1):
label_subs[' '.join(label_tokens[i: i + n])] += 1
for i in range(len_pred - n + 1):
if label_subs[' '.join(pred_tokens[i: i + n])] > 0:
num_matches += 1
label_subs[' '.join(pred_tokens[i: i + n])] -= 1
score *= math.pow(num_matches / (len_pred - n + 1), math.pow(0.5, n))
return score
```
最后,利用训练好的循环神经网络“编码器-解码器”模型,
[**将几个英语句子翻译成法语**],并计算BLEU的最终结果。
```
engs = ['go .', "i lost .", 'he\'s calm .', 'i\'m home .']
fras = ['va !', 'j\'ai perdu .', 'il est calme .', 'je suis chez moi .']
for eng, fra in zip(engs, fras):
translation, attention_weight_seq = predict_seq2seq(
net, eng, src_vocab, tgt_vocab, num_steps, device)
print(f'{eng} => {translation}, bleu {bleu(translation, fra, k=2):.3f}')
```
## 小结
* 根据“编码器-解码器”架构的设计,
我们可以使用两个循环神经网络来设计一个序列到序列学习的模型。
* 在实现编码器和解码器时,我们可以使用多层循环神经网络。
* 我们可以使用遮蔽来过滤不相关的计算,例如在计算损失时。
* 在“编码器-解码器”训练中,强制教学方法将原始输出序列(而非预测结果)输入解码器。
* BLEU是一种常用的评估方法,它通过测量预测序列和标签序列之间的$n$元语法的匹配度来评估预测。
## 练习
1. 你能通过调整超参数来改善翻译效果吗?
1. 重新运行实验并在计算损失时不使用遮蔽。你观察到什么结果?为什么?
1. 如果编码器和解码器的层数或者隐藏单元数不同,那么如何初始化解码器的隐状态?
1. 在训练中,如果用前一时间步的预测输入到解码器来代替强制教学,对性能有何影响?
1. 用长短期记忆网络替换门控循环单元重新运行实验。
1. 有没有其他方法来设计解码器的输出层?
[Discussions](https://discuss.d2l.ai/t/2782)
| github_jupyter |
```
# code for loading the format for the notebook
import os
# path : store the current path to convert back to it later
path = os.getcwd()
os.chdir( os.path.join('..', 'notebook_format') )
from formats import load_style
load_style()
os.chdir(path)
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# 1. magic for inline plot
# 2. magic to print version
# 3. magic so that the notebook will reload external python modules
%matplotlib inline
%load_ext watermark
%load_ext autoreload
%autoreload 2
from sklearn import datasets
from sklearn.metrics import accuracy_score
from sklearn.preprocessing import StandardScaler
from sklearn.linear_model import LogisticRegression
%watermark -a 'Ethen' -d -t -v -p numpy,pandas,matplotlib,sklearn
```
# Softmax Regression
**Softmax Regression** is a generalization of logistic regression that we can use for multi-class classification. If we want to assign probabilities to an object being one of several different things, softmax is the thing to do. Even later on, when we start training neural network models, the final step will be a layer of softmax.
A softmax regression has two steps: first we add up the evidence of our input being in certain classes, and then we convert that evidence into probabilities.
In **Softmax Regression**, we replace the sigmoid logistic function by the so-called *softmax* function $\phi(\cdot)$.
$$P(y=j \mid z^{(i)}) = \phi(z^{(i)}) = \frac{e^{z^{(i)}}}{\sum_{j=1}^{k} e^{z_{j}^{(i)}}}$$
where we define the net input *z* as
$$z = w_1x_1 + ... + w_mx_m + b= \sum_{l=1}^{m} w_l x_l + b= \mathbf{w}^T\mathbf{x} + b$$
(**w** is the weight vector, $\mathbf{x}$ is the feature vector of 1 training sample. Each $w$ corresponds to a feature $x$ and there're $m$ of them in total. $b$ is the bias unit. $k$ denotes the total number of classes.)
Now, this softmax function computes the probability that the $i_{th}$ training sample $\mathbf{x}^{(i)}$ belongs to class $l$ given the weight and net input $z^{(i)}$. So given the obtained weight $w$, we're basically compute the probability, $p(y = j \mid \mathbf{x^{(i)}; w}_j)$, the probability of the training sample belonging to class $j$ for each class label in $j = 1, \ldots, k$. Note the normalization term in the denominator which causes these class probabilities to sum up to one.
We can picture our softmax regression as looking something like the following, although with a lot more $x_s$. For each output, we compute a weighted sum of the $x_s$, add a bias, and then apply softmax.
<img src='images/softmax1.png' width="60%">
If we write that out as equations, we get:
<img src='images/softmax2.png' width="60%">
We can "vectorize" this procedure, turning it into a matrix multiplication and vector addition. This is helpful for computational efficiency. (It's also a useful way to think.)
<img src='images/softmax3.png' width="60%">
To illustrate the concept of softmax, let us walk through a concrete example. Suppose we have a training set consisting of 4 samples from 3 different classes (0, 1, and 2)
- $x_0 \rightarrow \text{class }0$
- $x_1 \rightarrow \text{class }1$
- $x_2 \rightarrow \text{class }2$
- $x_3 \rightarrow \text{class }2$
First, we apply one-hot encoding to encode the class labels into a format that we can more easily work with.
```
y = np.array([0, 1, 2, 2])
def one_hot_encode(y):
n_class = np.unique(y).shape[0]
y_encode = np.zeros((y.shape[0], n_class))
for idx, val in enumerate(y):
y_encode[idx, val] = 1.0
return y_encode
y_encode = one_hot_encode(y)
y_encode
```
A sample that belongs to class 0 (the first row) has a 1 in the first cell, a sample that belongs to class 1 has a 1 in the second cell of its row, and so forth.
Next, let us define the feature matrix of our 4 training samples. Here, we assume that our dataset consists of 2 features; thus, we create a 4x2 dimensional matrix of our samples and features.
Similarly, we create a 2x3 dimensional weight matrix (one row per feature and one column for each class).
```
X = np.array([[0.1, 0.5],
[1.1, 2.3],
[-1.1, -2.3],
[-1.5, -2.5]])
W = np.array([[0.1, 0.2, 0.3],
[0.1, 0.2, 0.3]])
bias = np.array([0.01, 0.1, 0.1])
print('Inputs X:\n', X)
print('\nWeights W:\n', W)
print('\nbias:\n', bias)
```
To compute the net input, we multiply the 4x2 matrix feature matrix `X` with the 2x3 (n_features x n_classes) weight matrix `W`, which yields a 4x3 output matrix (n_samples x n_classes) to which we then add the bias unit:
$$\mathbf{Z} = \mathbf{X}\mathbf{W} + \mathbf{b}$$
```
def net_input(X, W, b):
return X.dot(W) + b
net_in = net_input(X, W, bias)
print('net input:\n', net_in)
```
Now, it's time to compute the softmax activation that we discussed earlier:
$$P(y=j \mid z^{(i)}) = \phi_{softmax}(z^{(i)}) = \frac{e^{z^{(i)}}}{\sum_{j=1}^{k} e^{z_{j}^{(i)}}}$$
```
def softmax(z):
return np.exp(z) / np.sum(np.exp(z), axis = 1, keepdims = True)
smax = softmax(net_in)
print('softmax:\n', smax)
```
As we can see, the values for each sample (row) nicely sum up to 1 now. E.g., we can say that the first sample `[ 0.29450637 0.34216758 0.36332605]` has a 29.45% probability to belong to class 0. Now, in order to turn these probabilities back into class labels, we could simply take the argmax-index position of each row:
[[ 0.29450637 0.34216758 **0.36332605**] -> 2
[ 0.21290077 0.32728332 **0.45981591**] -> 2
[ **0.42860913** 0.33380113 0.23758974] -> 0
[ **0.44941979** 0.32962558 0.22095463]] -> 0
```
def to_classlabel(z):
return z.argmax(axis = 1)
print('predicted class labels: ', to_classlabel(smax))
```
As we can see, our predictions are terribly wrong, since the correct class labels are `[0, 1, 2, 2]`. Now, in order to train our model we need to measuring how inefficient our predictions are for describing the truth and then optimize on it. To do so we first need to define a loss/cost function $J(\cdot)$ that we want to minimize. One very common function is "cross-entropy":
$$J(\mathbf{W}; \mathbf{b}) = \frac{1}{n} \sum_{i=1}^{n} H( T^{(i)}, O^{(i)} )$$
which is the average of all cross-entropies $H$ over our $n$ training samples. The cross-entropy function is defined as:
$$H( T^{(i)}, O^{(i)} ) = -\sum_k T^{(i)} \cdot log(O^{(i)})$$
Where:
- $T$ stands for "target" (i.e., the *true* class labels)
- $O$ stands for output -- the computed *probability* via softmax; **not** the predicted class label.
- $\sum_k$ denotes adding up the difference between the target and the output for all classes.
```
def cross_entropy_cost(y_target, output):
return np.mean(-np.sum(y_target * np.log(output), axis = 1))
cost = cross_entropy_cost(y_target = y_encode, output = smax)
print('Cross Entropy Cost:', cost)
```
## Gradient Descent
Our objective in training a neural network is to find a set of weights that gives us the lowest error when we run it against our training data. There're many ways to find these weights and simplest one is so called **gradient descent**. It does this by giving us directions (using derivatives) on how to "shift" our weights to an optimum. It tells us whether we should increase or decrease the value of a specific weight in order to lower the error function.
Let's imagine we have a function $f(x) = x^4 - 3x^3 + 2$ and we want to find the minimum of this function using gradient descent. Here's a graph of that function:
```
from sympy.plotting import plot
from sympy import symbols, init_printing
# change default figure and font size
plt.rcParams['figure.figsize'] = 6, 4
plt.rcParams['font.size'] = 12
# plotting f(x) = x^4 - 3x^3 + 2, showing -2 < x <4
init_printing()
x = symbols('x')
fx = x ** 4 - 3 * x ** 3 + 2
p1 = plot(fx, (x, -2, 4), ylim = (-10, 50))
```
As you can see, there appears to be a minimum around ~2.3 or so. Gradient descent answers this question: If we were to start with a random value of x, which direction should we go if we want to get to the lowest point on this function? Let's imagine we pick a random x value, say <b>x = 4</b>, which would be somewhere way up on the right side of the graph. We obviously need to start going to the left if we want to get to the bottom. This is obvious when the function is an easily visualizable 2d plot, but when dealing with functions of multiple variables, we need to rely on the raw mathematics.
Calculus tells us that the derivative of a function at a particular point is the rate of change/slope of the tangent to that part of the function. So let's use derivatives to help us get to the bottom of this function. The derivative of $f(x) = x^4 - 3x^3 + 2$ is $f'(x) = 4x^3 - 9x^2$. So if we plug in our random point from above (x=4) into the first derivative of $f(x)$ we get $f'(4) = 4(4)^3 - 9(4)^2 = 112$. So how does 112 tell us where to go? Well, first of all, it's positive. If we were to compute $f'(-1)$ we get a negative number (-13). So it looks like we can say that whenever the $f'(x)$ for a particular $x$ is positive, we should move to the left (decrease x) and whenever it's negative, we should move to the right (increase x).
We'll now formalize this: When we start with a random x and compute it's deriative $f'(x)$, our <b>new x</b> should then be proportional to $x - f'(x)$. The word proportional is there because we wish to control <em>to what degree</em> we move at each step, for example when we compute $f'(4)=112$, do we really want our new $x$ to be $x - 112 = -108$? No, if we jump all the way to -108, we're even farther from the minimum than we were before. Instead, we want to take relatively <em>small</em> steps toward the minimum.
Let's say that for any random $x$, we want to take a step (change $x$ a little bit) such that our <b>new $x$</b> $ = x - \alpha*f'(x)$. We'll call $\alpha$ (alpha) our <em>learning rate or step size</em> because it determines how big of a step we take. $\alpha$ is something we will just have to play around with to find a good value. Some functions might require bigger steps, others smaller steps.
Suppose we've set our $\alpha$ to be 0.001. This means, if we randomly started at $f'(4)=112$ then our new $x$ will be $ = 4 - (0.001 * 112) = 3.888$. So we moved to the left a little bit, toward the optimum. Let's do it again. $x_{new} = x - \alpha*f'(3.888) = 3.888 - (0.001 * 99.0436) = 3.79$. Nice, we're indeed moving to the left, closer to the minimum of $f(x)$, little by little. And we'll keep on doing this until we've reached convergence. By convergence, we mean that if the absolute value of the difference between the updated $x$ and the old $x$ is smaller than some randomly small number that we set, denoted as $\epsilon$ (epsilon).
```
x_old = 0
x_new = 4 # The algorithm starts at x = 4
alpha = 0.01 # step size
epsilon = 0.00001
def f_derivative(x):
return 4 * x ** 3 - 9 * x ** 2
while abs(x_new - x_old) > epsilon:
x_old = x_new
x_new = x_old - alpha * f_derivative(x_old)
print("Local minimum occurs at", x_new)
```
The script above says that if the absolute difference of $x$ between the two iterations is not changing by more than 0.00001, then we're probably at the bottom of the "bowl" because our slope is approaching 0, and therefore we should stop and call it a day. Now, if you remember some calculus and algebra, you could have solved for this minimum analytically, and you should get 2.25. Very close to what our gradient descent algorithm above found.
## More Gradient Descent...
As you might imagine, when we use gradient descent for a neural network, things get a lot more complicated. Not because gradient descent gets more complicated, it still ends up just being a matter of taking small steps downhill, it's that we need that pesky derivative in order to use gradient descent, and the derivative of a neural network cost function (with respect to its weights) is pretty intense. It's not a matter of just analytically solving $f(x)=x^2, f'(x)=2x$ , because the output of a neural net has many nested or "inner" functions.
Also unlike our toy math problem above, a neural network may have many weights. We need to find the optimal value for each individual weight to lower the cost for our entire neural net output. This requires taking the partial derivative of the cost/error function with respect to a single weight, and then running gradient descent for each individual weight. Thus, for any individual weight $W_j$, we'll compute the following:
$$ W_j^{(t + 1)} = W_j^{(t)} - \alpha * \frac{\partial L}{\partial W_j}$$
Where:
- $L$ denotes the loss function that we've defined.
- $W_j^{(t)}$ denotes the weight of the $j_{th}$ feature at iteration $t$.
And as before, we do this iteratively for each weight, many times, until the whole network's cost function is minimized.
In order to learn the weight for our softmax model via gradient descent, we then need to compute the gradient of our cost function for each class $j \in \{0, 1, ..., k\}$.
$$\nabla \mathbf{w}_j \, J(\mathbf{W}; \mathbf{b})$$
We won't be going through the tedious details here, but this cost's gradient turns out to be simply:
$$\nabla \mathbf{w}_j \, J(\mathbf{W}; \mathbf{b}) = \frac{1}{n} \sum^{n}_{i=0} \big[\mathbf{x}^{(i)}_j\ \big( O^{(i)} - T^{(i)} \big) \big]$$
We can then use the cost derivate to update the weights in opposite direction of the cost gradient with learning rate $\eta$:
$$\mathbf{w}_j := \mathbf{w}_j - \eta \nabla \mathbf{w}_j \, J(\mathbf{W}; \mathbf{b})$$
(note that $\mathbf{w}_j$ is the weight vector for the class $y=j$), and we update the bias units using:
$$
\mathbf{b}_j := \mathbf{b}_j - \eta \bigg[ \frac{1}{n} \sum^{n}_{i=0} \big( O^{(i)} - T^{(i)} \big) \bigg]
$$
As a penalty against complexity, an approach to reduce the variance of our model and decrease the degree of overfitting by adding additional bias, we can further add a regularization term such as the L2 term with the regularization parameter $\lambda$:
$$\frac{\lambda}{2} ||\mathbf{w}||_{2}^{2}$$
where $||\mathbf{w}||_{2}^{2}$ simply means adding up the squared weights across all the features and classes.
$$||\mathbf{w}||_{2}^{2} = \sum^{m}_{l=0} \sum^{k}_{j=0} w_{l, j}^2$$
so that our cost function becomes
$$
J(\mathbf{W}; \mathbf{b}) = \frac{1}{n} \sum_{i=1}^{n} H( T^{(i)}, O^{(i)} ) + \frac{\lambda}{2} ||\mathbf{w}||_{2}^{2}
$$
and we define the "regularized" weight update as
$$
\mathbf{w}_j := \mathbf{w}_j - \eta \big[\nabla \mathbf{w}_j \, J(\mathbf{W}) + \lambda \mathbf{w}_j \big]
$$
Note that we don't regularize the bias term, thus the update function for it stays the same.
## Softmax Regression Code
Bringing the concepts together, we could come up with an implementation as follows: Note that for the weight and bias parameter, we'll have initialize a value for it. Here we'll simply draw the weights from a normal distribution and set the bias as zero. The code can be obtained [here](https://github.com/ethen8181/machine-learning/blob/master/deep_learning/softmax.py).
```
# import some data to play with
iris = datasets.load_iris()
X = iris.data
y = iris.target
# standardize the input features
scaler = StandardScaler()
X_std = scaler.fit_transform(X)
from softmax import SoftmaxRegression
# train the softmax using batch gradient descent,
# eta: learning rate, epochs : number of iterations, minibatches, number of
# training data to use for training at each iteration
softmax_reg = SoftmaxRegression(eta = 0.1, epochs = 10, minibatches = y.shape[0])
softmax_reg.fit(X_std, y)
# print the training accuracy
y_pred = softmax_reg.predict(X_std)
accuracy = np.sum(y_pred == y) / y.shape[0]
print('accuracy: ', accuracy)
# use a library to ensure comparable results
log_reg = LogisticRegression()
log_reg.fit(X_std, y)
y_pred = log_reg.predict(X_std)
print('accuracy library: ', accuracy_score(y_true = y, y_pred = y_pred))
```
# Reference
- [Blog: Softmax Regression](http://nbviewer.jupyter.org/github/rasbt/python-machine-learning-book/blob/master/code/bonus/softmax-regression.ipynb)
- [Blog: Gradient Descent with Backpropagation](http://outlace.com/Beginner-Tutorial-Backpropagation/)
- [TensorFlow Documentation: MNIST For ML Beginners](https://www.tensorflow.org/get_started/mnist/beginners)
| github_jupyter |
```
import torch
import torch.optim as optim
import torch.nn.functional as F
import torchvision
import torchvision.datasets as datasets
import torchvision.models as models
import torchvision.transforms as transforms
import time
import matplotlib.pyplot as plt
import numpy as np
dataset = datasets.ImageFolder(
'dataset',
transforms.Compose([
transforms.ColorJitter(0.1, 0.1, 0.1, 0.1),
transforms.Resize((224, 224)),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225])
])
)
train_dataset, test_dataset = torch.utils.data.random_split(dataset, [len(dataset) - 65, 65])
train_loader = torch.utils.data.DataLoader(
train_dataset,
batch_size=8,
shuffle=True,
num_workers=0,
)
test_loader = torch.utils.data.DataLoader(
test_dataset,
batch_size=8,
shuffle=True,
num_workers=0,
)
model = models.alexnet(pretrained=True)
model.classifier[6] = torch.nn.Linear(model.classifier[6].in_features, 2)
device = torch.device('cuda')
model = model.to(device)
import datetime
today = datetime.date.today()
strToday = str(today.year) + '_' + str(today.month) + '_' + str(today.day)
RESNET_MODEL = 'Botline_CA_model_alexnet_' + strToday + '.pth'
TRT_MODEL = 'Botline_CA_model_alexnet_trt_' + strToday + '.pth'
NUM_EPOCHS = 50
best_accuracy = 0.0
optimizer = optim.SGD(model.parameters(), lr=0.001, momentum=0.9)
accuracyList = list()
timeList = list()
startTime = time.time()
for epoch in range(NUM_EPOCHS):
t0 = time.time()
for images, labels in iter(train_loader):
images = images.to(device)
labels = labels.to(device)
optimizer.zero_grad()
outputs = model(images)
loss = F.cross_entropy(outputs, labels)
loss.backward()
optimizer.step()
test_error_count = 0.0
for images, labels in iter(test_loader):
images = images.to(device)
labels = labels.to(device)
outputs = model(images)
test_error_count += float(torch.sum(torch.abs(labels - outputs.argmax(1))))
elapsedTime = time.time() - t0
test_accuracy = 1.0 - float(test_error_count) / float(len(test_dataset))
accuracyList.append(test_accuracy)
timeList.append(elapsedTime)
print('[%2d] Accuracy: %f\tTime : %.2f sec' % (epoch, test_accuracy, elapsedTime))
if test_accuracy > best_accuracy:
torch.save(model.state_dict(), RESNET_MODEL)
best_accuracy = test_accuracy
endTime = time.time()
print('Time: {0:.2f}'.format(endTime - startTime))
plt.figure(figsize=(12, 3))
accX = range(len(accuracyList))
plt.subplot(121)
plt.plot(accX, accuracyList)
plt.ylabel('Accuracy')
timeX = range(len(timeList))
plt.subplot(122)
plt.plot(timeX, timeList)
plt.ylabel('Time')
plt.show()
```
| github_jupyter |
## Imports
```
%matplotlib inline
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split, cross_val_score
from sklearn.metrics import mean_squared_error
from sklearn.preprocessing import StandardScaler
from sklearn.linear_model import LinearRegression, Lasso, Ridge
```
## Generate Data
```
samples = 500
np.random.seed(10) ## seed for reproducibility
f1 = np.random.uniform(low=0, high=10, size=samples) ## garbage feature
f2 = np.random.rand(samples)
f3 = np.random.binomial(n=1, p=0.5, size=samples)
f4 = None
f5 = np.random.normal(1, 2.5, samples)
d = {'f1':f1, 'f2':f2, 'f3':f3, 'f4':f4, 'f5':f5}
df = pd.DataFrame(d)
df['target'] = None
df.head()
# Set target values w/noise
for i, _ in df.iterrows():
df.loc[i, 'target'] = df.loc[i, 'f2'] * df.loc[i, 'f3'] + df.loc[i, 'f5'] * df.loc[i, 'f3'] + np.random.rand() * 3
df.loc[i, 'f4'] = df.loc[i, 'f2'] ** 2.8 + np.random.normal(loc=0, scale=1.25, size=1)
df['f4'] = df.f4.astype('float')
df['target'] = df.target.astype('float')
df.head()
df.describe()
df.corr()
sns.pairplot(df);
```
## Train/Test Split
```
X_train, X_test, y_train, y_test = train_test_split(df[['f1', 'f2', 'f3', 'f4', 'f5']],
df['target'],
test_size=0.2,
random_state=42)
```
## StandardScale
```
ss = StandardScaler()
ss.fit(X_train)
X_train_std = ss.transform(X_train)
X_test_std = ss.transform(X_test)
```
## Modeling
```
lr = LinearRegression()
lasso = Lasso(alpha=0.01, random_state=42)
ridge = Ridge(alpha=0.01, random_state=42)
lr.fit(X_train_std, y_train)
lasso.fit(X_train_std, y_train)
ridge.fit(X_train_std, y_train)
```
## Results
```
def pretty_print_coef(obj):
print('intercept: {0:.4}'.format(obj.intercept_))
print('coef: {0:.3} {1:.4} {2:.4} {3:.3} {4:.3}'.format(obj.coef_[0],
obj.coef_[1],
obj.coef_[2],
obj.coef_[3],
obj.coef_[4]))
models = (lr, lasso, ridge)
for model in models:
print(str(model))
pretty_print_coef(model)
print('R^2:', model.score(X_train_std, y_train))
print('MSE:', mean_squared_error(y_test, model.predict(X_test_std)))
print()
np.mean(cross_val_score(lr, X_train_std, y_train, scoring='neg_mean_squared_error', cv=5, n_jobs=-1) * -1)
np.mean(cross_val_score(lasso, X_train_std, y_train, scoring='neg_mean_squared_error', cv=5, n_jobs=-1) * -1)
np.mean(cross_val_score(ridge, X_train_std, y_train, scoring='neg_mean_squared_error', cv=5, n_jobs=-1) * -1)
```
## Residuals
```
fig, axes = plt.subplots(1, 3, sharex=False, sharey=False)
fig.suptitle('[Residual Plots]')
fig.set_size_inches(18,5)
axes[0].plot(lr.predict(X_test_std), y_test-lr.predict(X_test_std), 'bo')
axes[0].axhline(y=0, color='k')
axes[0].grid()
axes[0].set_title('Linear')
axes[0].set_xlabel('predicted values')
axes[0].set_ylabel('residuals')
axes[1].plot(lasso.predict(X_test_std), y_test-lasso.predict(X_test_std), 'go')
axes[1].axhline(y=0, color='k')
axes[1].grid()
axes[1].set_title('Lasso')
axes[1].set_xlabel('predicted values')
axes[1].set_ylabel('residuals')
axes[2].plot(ridge.predict(X_test_std), y_test-ridge.predict(X_test_std), 'ro')
axes[2].axhline(y=0, color='k')
axes[2].grid()
axes[2].set_title('Ridge')
axes[2].set_xlabel('predicted values')
axes[2].set_ylabel('residuals');
```
## Notes
We can tell from the residuals that there's signal that we're not catching with our current model. The reason is obvious in this case, because we know how the data was generated. The **target** is built on interaction terms.
## Random Forest (for fun)
```
from sklearn.ensemble import RandomForestRegressor
rf = RandomForestRegressor(n_estimators=10, max_depth=5, n_jobs=-1, random_state=42)
rf.fit(X_train, y_train)
mean_squared_error(y_test, rf.predict(X_test))
rf.feature_importances_
fig, axes = plt.subplots(1, 4, sharex=False, sharey=False)
fig.suptitle('[Residual Plots]')
fig.set_size_inches(18,5)
axes[0].plot(lr.predict(X_test_std), y_test-lr.predict(X_test_std), 'bo')
axes[0].axhline(y=0, color='k')
axes[0].grid()
axes[0].set_title('Linear')
axes[0].set_xlabel('predicted values')
axes[0].set_ylabel('residuals')
axes[1].plot(lasso.predict(X_test_std), y_test-lasso.predict(X_test_std), 'go')
axes[1].axhline(y=0, color='k')
axes[1].grid()
axes[1].set_title('Lasso')
axes[1].set_xlabel('predicted values')
axes[1].set_ylabel('residuals')
axes[2].plot(ridge.predict(X_test_std), y_test-ridge.predict(X_test_std), 'ro')
axes[2].axhline(y=0, color='k')
axes[2].grid()
axes[2].set_title('Ridge')
axes[2].set_xlabel('predicted values')
axes[2].set_ylabel('residuals');
axes[3].plot(rf.predict(X_test), y_test-rf.predict(X_test), 'ko')
axes[3].axhline(y=0, color='k')
axes[3].grid()
axes[3].set_title('RF')
axes[3].set_xlabel('predicted values')
axes[3].set_ylabel('residuals');
```
While random forest does a better job catching the interaction between variables, we still see some pattern in the residuals meaning we haven't captured all the signal. Nonetheless, random forest is signficantly better than linear regression, lasso, and ridge on the raw features. We can combat this with feature engineering, however.
| github_jupyter |
##### Copyright 2019 The TensorFlow Hub Authors.
Licensed under the Apache License, Version 2.0 (the "License");
```
# Copyright 2019 The TensorFlow Hub Authors. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
```
## TF-Hub Demo for Fast Style Transfer for Arbitrary Styles
Based on the model code in [magenta](https://github.com/tensorflow/magenta/tree/master/magenta/models/arbitrary_image_stylization) and the publication:
[Exploring the structure of a real-time, arbitrary neural artistic stylization
network](https://arxiv.org/abs/1705.06830).
*Golnaz Ghiasi, Honglak Lee,
Manjunath Kudlur, Vincent Dumoulin, Jonathon Shlens*,
Proceedings of the British Machine Vision Conference (BMVC), 2017.
Let's start with importing TF-2 and all relevant dependencies.
```
# We want to use TensorFlow 2.0 in the Eager mode for this demonstration. But this module works as well with the Graph mode.
!pip install tensorflow --quiet
from __future__ import absolute_import, division, print_function
import functools
import os
from matplotlib import gridspec
import matplotlib.pylab as plt
import numpy as np
import tensorflow as tf
import tensorflow_hub as hub
print("TF Version: ", tf.__version__)
print("TF-Hub version: ", hub.__version__)
print("Eager mode enabled: ", tf.executing_eagerly())
print("GPU available: ", tf.test.is_gpu_available())
# @title Define image loading and visualization functions { display-mode: "form" }
def crop_center(image):
"""Returns a cropped square image."""
shape = image.shape
new_shape = min(shape[1], shape[2])
offset_y = max(shape[1] - shape[2], 0) // 2
offset_x = max(shape[2] - shape[1], 0) // 2
image = tf.image.crop_to_bounding_box(
image, offset_y, offset_x, new_shape, new_shape)
return image
@functools.lru_cache(maxsize=None)
def load_image(image_url, image_size=(256, 256), preserve_aspect_ratio=True):
"""Loads and preprocesses images."""
# Cache image file locally.
image_path = tf.keras.utils.get_file(os.path.basename(image_url)[-128:], image_url)
# Load and convert to float32 numpy array, add batch dimension, and normalize to range [0, 1].
img = plt.imread(image_path).astype(np.float32)[np.newaxis, ...]
if img.max() > 1.0:
img = img / 255.
if len(img.shape) == 3:
img = tf.stack([img, img, img], axis=-1)
img = crop_center(img)
img = tf.image.resize(img, image_size, preserve_aspect_ratio=True)
return img
def show_n(images, titles=('',)):
n = len(images)
image_sizes = [image.shape[1] for image in images]
w = (image_sizes[0] * 6) // 320
plt.figure(figsize=(w * n, w))
gs = gridspec.GridSpec(1, n, width_ratios=image_sizes)
for i in range(n):
plt.subplot(gs[i])
plt.imshow(images[i][0], aspect='equal')
plt.axis('off')
plt.title(titles[i] if len(titles) > i else '')
plt.show()
```
Let's get as well some images to play with.
```
# @title Load example images { display-mode: "form" }
content_image_url = 'https://upload.wikimedia.org/wikipedia/commons/thumb/f/fd/Golden_Gate_Bridge_from_Battery_Spencer.jpg/640px-Golden_Gate_Bridge_from_Battery_Spencer.jpg' # @param {type:"string"}
style_image_url = 'https://upload.wikimedia.org/wikipedia/commons/0/0a/The_Great_Wave_off_Kanagawa.jpg' # @param {type:"string"}
output_image_size = 384 # @param {type:"integer"}
# The content image size can be arbitrary.
content_img_size = (output_image_size, output_image_size)
# The style prediction model was trained with image size 256 and it's the
# recommended image size for the style image (though, other sizes work as
# well but will lead to different results).
style_img_size = (256, 256) # Recommended to keep it at 256.
content_image = load_image(content_image_url, content_img_size)
style_image = load_image(style_image_url, style_img_size)
style_image = tf.nn.avg_pool(style_image, ksize=[3,3], strides=[1,1], padding='SAME')
show_n([content_image, style_image], ['Content image', 'Style image'])
```
## Import TF-Hub module
```
# Load TF-Hub module.
hub_handle = 'https://tfhub.dev/google/magenta/arbitrary-image-stylization-v1-256/2'
hub_module = hub.load(hub_handle)
```
The signature of this hub module for image stylization is:
```
outputs = hub_module(content_image, style_image)
stylized_image = outputs[0]
```
Where `content_image`, `style_image`, and `stylized_image` are expected to be 4-D Tensors with shapes `[batch_size, image_height, image_width, 3]`.
In the current example we provide only single images and therefore the batch dimension is 1, but one can use the same module to process more images at the same time.
The input and output values of the images should be in the range [0, 1].
The shapes of content and style image don't have to match. Output image shape
is the same as the content image shape.
## Demonstrate image stylization
```
# Stylize content image with given style image.
# This is pretty fast within a few milliseconds on a GPU.
outputs = hub_module(tf.constant(content_image), tf.constant(style_image))
stylized_image = outputs[0]
# Visualize input images and the generated stylized image.
show_n([content_image, style_image, stylized_image], titles=['Original content image', 'Style image', 'Stylized image'])
```
## Let's try it on more images
```
# @title To Run: Load more images { display-mode: "form" }
content_urls = dict(
sea_turtle='https://upload.wikimedia.org/wikipedia/commons/d/d7/Green_Sea_Turtle_grazing_seagrass.jpg',
tuebingen='https://upload.wikimedia.org/wikipedia/commons/0/00/Tuebingen_Neckarfront.jpg',
grace_hopper='https://storage.googleapis.com/download.tensorflow.org/example_images/grace_hopper.jpg',
)
style_urls = dict(
kanagawa_great_wave='https://upload.wikimedia.org/wikipedia/commons/0/0a/The_Great_Wave_off_Kanagawa.jpg',
kandinsky_composition_7='https://upload.wikimedia.org/wikipedia/commons/b/b4/Vassily_Kandinsky%2C_1913_-_Composition_7.jpg',
hubble_pillars_of_creation='https://upload.wikimedia.org/wikipedia/commons/6/68/Pillars_of_creation_2014_HST_WFC3-UVIS_full-res_denoised.jpg',
van_gogh_starry_night='https://upload.wikimedia.org/wikipedia/commons/thumb/e/ea/Van_Gogh_-_Starry_Night_-_Google_Art_Project.jpg/1024px-Van_Gogh_-_Starry_Night_-_Google_Art_Project.jpg',
turner_nantes='https://upload.wikimedia.org/wikipedia/commons/b/b7/JMW_Turner_-_Nantes_from_the_Ile_Feydeau.jpg',
munch_scream='https://upload.wikimedia.org/wikipedia/commons/c/c5/Edvard_Munch%2C_1893%2C_The_Scream%2C_oil%2C_tempera_and_pastel_on_cardboard%2C_91_x_73_cm%2C_National_Gallery_of_Norway.jpg',
picasso_demoiselles_avignon='https://upload.wikimedia.org/wikipedia/en/4/4c/Les_Demoiselles_d%27Avignon.jpg',
picasso_violin='https://upload.wikimedia.org/wikipedia/en/3/3c/Pablo_Picasso%2C_1911-12%2C_Violon_%28Violin%29%2C_oil_on_canvas%2C_Kr%C3%B6ller-M%C3%BCller_Museum%2C_Otterlo%2C_Netherlands.jpg',
picasso_bottle_of_rum='https://upload.wikimedia.org/wikipedia/en/7/7f/Pablo_Picasso%2C_1911%2C_Still_Life_with_a_Bottle_of_Rum%2C_oil_on_canvas%2C_61.3_x_50.5_cm%2C_Metropolitan_Museum_of_Art%2C_New_York.jpg',
fire='https://upload.wikimedia.org/wikipedia/commons/3/36/Large_bonfire.jpg',
derkovits_woman_head='https://upload.wikimedia.org/wikipedia/commons/0/0d/Derkovits_Gyula_Woman_head_1922.jpg',
amadeo_style_life='https://upload.wikimedia.org/wikipedia/commons/8/8e/Untitled_%28Still_life%29_%281913%29_-_Amadeo_Souza-Cardoso_%281887-1918%29_%2817385824283%29.jpg',
derkovtis_talig='https://upload.wikimedia.org/wikipedia/commons/3/37/Derkovits_Gyula_Talig%C3%A1s_1920.jpg',
amadeo_cardoso='https://upload.wikimedia.org/wikipedia/commons/7/7d/Amadeo_de_Souza-Cardoso%2C_1915_-_Landscape_with_black_figure.jpg'
)
content_image_size = 384
style_image_size = 256
content_images = {k: load_image(v, (content_image_size, content_image_size)) for k, v in content_urls.items()}
style_images = {k: load_image(v, (style_image_size, style_image_size)) for k, v in style_urls.items()}
style_images = {k: tf.nn.avg_pool(style_image, ksize=[3,3], strides=[1,1], padding='SAME') for k, style_image in style_images.items()}
#@title Specify the main content image and the style you want to use. { display-mode: "form" }
content_name = 'sea_turtle' # @param ['sea_turtle', 'tuebingen', 'grace_hopper']
style_name = 'munch_scream' # @param ['kanagawa_great_wave', 'kandinsky_composition_7', 'hubble_pillars_of_creation', 'van_gogh_starry_night', 'turner_nantes', 'munch_scream', 'picasso_demoiselles_avignon', 'picasso_violin', 'picasso_bottle_of_rum', 'fire', 'derkovits_woman_head', 'amadeo_style_life', 'derkovtis_talig', 'amadeo_cardoso']
stylized_image = hub_module(tf.constant(content_images[content_name]),
tf.constant(style_images[style_name]))[0]
show_n([content_images[content_name], style_images[style_name], stylized_image],
titles=['Original content image', 'Style image', 'Stylized image'])
```
# Try it out with your own images.
Paste URLs to your own image files in the boxes below.
```
# @title Load example images { display-mode: "form" }
content_image_url = '' # @param {type:"string"}
style_image_url = '' # @param {type:"string"}
output_image_size = 384 # @param {type:"integer"}
# The content image size can be arbitrary.
content_img_size = (output_image_size, output_image_size)
# The style prediction model was trained with image size 256 and it's the
# recommended image size for the style image (though, other sizes work as
# well but will lead to different results).
style_img_size = (256, 256) # Recommended to keep it at 256.
content_image = load_image(content_image_url, content_img_size)
style_image = load_image(style_image_url, style_img_size)
style_image = tf.nn.avg_pool(style_image, ksize=[3,3], strides=[1,1], padding='SAME')
show_n([content_image, style_image], ['Content image', 'Style image'])
outputs = hub_module(tf.constant(content_image), tf.constant(style_image))
show_n([outputs[0]], titles=['Stylized image'])
```
| github_jupyter |
## Dependencies
```
import json, warnings, shutil, glob
from jigsaw_utility_scripts import *
from scripts_step_lr_schedulers import *
from transformers import TFXLMRobertaModel, XLMRobertaConfig
from tensorflow.keras.models import Model
from tensorflow.keras import optimizers, metrics, losses, layers
SEED = 0
seed_everything(SEED)
warnings.filterwarnings("ignore")
```
## TPU configuration
```
strategy, tpu = set_up_strategy()
print("REPLICAS: ", strategy.num_replicas_in_sync)
AUTO = tf.data.experimental.AUTOTUNE
```
# Load data
```
database_base_path = '/kaggle/input/jigsaw-data-split-roberta-192-ratio-2-clean-tail4/'
k_fold = pd.read_csv(database_base_path + '5-fold.csv')
valid_df = pd.read_csv("/kaggle/input/jigsaw-multilingual-toxic-comment-classification/validation.csv",
usecols=['comment_text', 'toxic', 'lang'])
print('Train samples: %d' % len(k_fold))
display(k_fold.head())
print('Validation samples: %d' % len(valid_df))
display(valid_df.head())
base_data_path = 'fold_1/'
fold_n = 1
# Unzip files
!tar -xvf /kaggle/input/jigsaw-data-split-roberta-192-ratio-2-clean-tail4/fold_1.tar.gz
```
# Model parameters
```
base_path = '/kaggle/input/jigsaw-transformers/XLM-RoBERTa/'
config = {
"MAX_LEN": 192,
"BATCH_SIZE": 128,
"EPOCHS": 4,
"LEARNING_RATE": 1e-5,
"ES_PATIENCE": None,
"base_model_path": base_path + 'tf-xlm-roberta-large-tf_model.h5',
"config_path": base_path + 'xlm-roberta-large-config.json'
}
with open('config.json', 'w') as json_file:
json.dump(json.loads(json.dumps(config)), json_file)
```
## Learning rate schedule
```
lr_min = 1e-7
lr_start = 1e-7
lr_max = config['LEARNING_RATE']
step_size = len(k_fold[k_fold[f'fold_{fold_n}'] == 'train']) // config['BATCH_SIZE']
total_steps = config['EPOCHS'] * step_size
hold_max_steps = 0
warmup_steps = step_size * 1
decay = .9997
rng = [i for i in range(0, total_steps, config['BATCH_SIZE'])]
y = [exponential_schedule_with_warmup(tf.cast(x, tf.float32), warmup_steps, hold_max_steps,
lr_start, lr_max, lr_min, decay) for x in rng]
sns.set(style="whitegrid")
fig, ax = plt.subplots(figsize=(20, 6))
plt.plot(rng, y)
print("Learning rate schedule: {:.3g} to {:.3g} to {:.3g}".format(y[0], max(y), y[-1]))
```
# Model
```
module_config = XLMRobertaConfig.from_pretrained(config['config_path'], output_hidden_states=False)
def model_fn(MAX_LEN):
input_ids = layers.Input(shape=(MAX_LEN,), dtype=tf.int32, name='input_ids')
attention_mask = layers.Input(shape=(MAX_LEN,), dtype=tf.int32, name='attention_mask')
base_model = TFXLMRobertaModel.from_pretrained(config['base_model_path'], config=module_config)
last_hidden_state, _ = base_model({'input_ids': input_ids, 'attention_mask': attention_mask})
cls_token = last_hidden_state[:, 0, :]
output = layers.Dense(1, activation='sigmoid', name='output')(cls_token)
model = Model(inputs=[input_ids, attention_mask], outputs=output)
return model
```
# Train
```
# Load data
x_train = np.load(base_data_path + 'x_train.npy')
y_train = np.load(base_data_path + 'y_train_int.npy').reshape(x_train.shape[1], 1).astype(np.float32)
x_valid_ml = np.load(database_base_path + 'x_valid.npy')
y_valid_ml = np.load(database_base_path + 'y_valid.npy').reshape(x_valid_ml.shape[1], 1).astype(np.float32)
#################### ADD TAIL ####################
x_train_tail = np.load(base_data_path + 'x_train_tail.npy')
y_train_tail = np.load(base_data_path + 'y_train_int_tail.npy').reshape(x_train_tail.shape[1], 1).astype(np.float32)
x_train = np.hstack([x_train, x_train_tail])
y_train = np.vstack([y_train, y_train_tail])
step_size = x_train.shape[1] // config['BATCH_SIZE']
valid_step_size = x_valid_ml.shape[1] // config['BATCH_SIZE']
# Build TF datasets
train_dist_ds = strategy.experimental_distribute_dataset(get_training_dataset(x_train, y_train, config['BATCH_SIZE'], AUTO, seed=SEED))
valid_dist_ds = strategy.experimental_distribute_dataset(get_validation_dataset(x_valid_ml, y_valid_ml, config['BATCH_SIZE'], AUTO, repeated=True, seed=SEED))
train_data_iter = iter(train_dist_ds)
valid_data_iter = iter(valid_dist_ds)
# Step functions
@tf.function
def train_step(data_iter):
def train_step_fn(x, y):
with tf.GradientTape() as tape:
probabilities = model(x, training=True)
loss = loss_fn(y, probabilities)
grads = tape.gradient(loss, model.trainable_variables)
optimizer.apply_gradients(zip(grads, model.trainable_variables))
train_auc.update_state(y, probabilities)
train_loss.update_state(loss)
for _ in tf.range(step_size):
strategy.experimental_run_v2(train_step_fn, next(data_iter))
@tf.function
def valid_step(data_iter):
def valid_step_fn(x, y):
probabilities = model(x, training=False)
loss = loss_fn(y, probabilities)
valid_auc.update_state(y, probabilities)
valid_loss.update_state(loss)
for _ in tf.range(valid_step_size):
strategy.experimental_run_v2(valid_step_fn, next(data_iter))
# Train model
with strategy.scope():
model = model_fn(config['MAX_LEN'])
optimizer = optimizers.Adam(learning_rate=lambda:
exponential_schedule_with_warmup(tf.cast(optimizer.iterations, tf.float32),
warmup_steps, hold_max_steps, lr_start,
lr_max, lr_min, decay))
loss_fn = losses.binary_crossentropy
train_auc = metrics.AUC()
valid_auc = metrics.AUC()
train_loss = metrics.Sum()
valid_loss = metrics.Sum()
metrics_dict = {'loss': train_loss, 'auc': train_auc,
'val_loss': valid_loss, 'val_auc': valid_auc}
history = custom_fit(model, metrics_dict, train_step, valid_step, train_data_iter, valid_data_iter,
step_size, valid_step_size, config['BATCH_SIZE'], config['EPOCHS'],
config['ES_PATIENCE'], save_last=False)
# model.save_weights('model.h5')
# Make predictions
x_train = np.load(base_data_path + 'x_train.npy')
x_valid = np.load(base_data_path + 'x_valid.npy')
x_valid_ml_eval = np.load(database_base_path + 'x_valid.npy')
train_preds = model.predict(get_test_dataset(x_train, config['BATCH_SIZE'], AUTO))
valid_preds = model.predict(get_test_dataset(x_valid, config['BATCH_SIZE'], AUTO))
valid_ml_preds = model.predict(get_test_dataset(x_valid_ml_eval, config['BATCH_SIZE'], AUTO))
k_fold.loc[k_fold[f'fold_{fold_n}'] == 'train', f'pred_{fold_n}'] = np.round(train_preds)
k_fold.loc[k_fold[f'fold_{fold_n}'] == 'validation', f'pred_{fold_n}'] = np.round(valid_preds)
valid_df[f'pred_{fold_n}'] = valid_ml_preds
# Fine-tune on validation set
#################### ADD TAIL ####################
x_valid_ml_tail = np.hstack([x_valid_ml, np.load(database_base_path + 'x_valid_tail.npy')])
y_valid_ml_tail = np.vstack([y_valid_ml, y_valid_ml])
valid_step_size_tail = x_valid_ml_tail.shape[1] // config['BATCH_SIZE']
# Build TF datasets
train_ml_dist_ds = strategy.experimental_distribute_dataset(get_training_dataset(x_valid_ml_tail, y_valid_ml_tail,
config['BATCH_SIZE'], AUTO, seed=SEED))
train_ml_data_iter = iter(train_ml_dist_ds)
history_ml = custom_fit(model, metrics_dict, train_step, valid_step, train_ml_data_iter, valid_data_iter,
valid_step_size_tail, valid_step_size, config['BATCH_SIZE'], 1,
config['ES_PATIENCE'], save_last=False)
# Join history
for key in history_ml.keys():
history[key] += history_ml[key]
model.save_weights('model_ml.h5')
# Make predictions
valid_ml_preds = model.predict(get_test_dataset(x_valid_ml_eval, config['BATCH_SIZE'], AUTO))
valid_df[f'pred_ml_{fold_n}'] = valid_ml_preds
### Delete data dir
shutil.rmtree(base_data_path)
```
## Model loss graph
```
plot_metrics(history)
```
# Model evaluation
```
display(evaluate_model_single_fold(k_fold, fold_n, label_col='toxic_int').style.applymap(color_map))
```
# Confusion matrix
```
train_set = k_fold[k_fold[f'fold_{fold_n}'] == 'train']
validation_set = k_fold[k_fold[f'fold_{fold_n}'] == 'validation']
plot_confusion_matrix(train_set['toxic_int'], train_set[f'pred_{fold_n}'],
validation_set['toxic_int'], validation_set[f'pred_{fold_n}'])
```
# Model evaluation by language
```
display(evaluate_model_single_fold_lang(valid_df, fold_n).style.applymap(color_map))
# ML fine-tunned preds
display(evaluate_model_single_fold_lang(valid_df, fold_n, pred_col='pred_ml').style.applymap(color_map))
```
# Visualize predictions
```
pd.set_option('max_colwidth', 120)
print('English validation set')
display(k_fold[['comment_text', 'toxic'] + [c for c in k_fold.columns if c.startswith('pred')]].head(10))
print('Multilingual validation set')
display(valid_df[['comment_text', 'toxic'] + [c for c in valid_df.columns if c.startswith('pred')]].head(10))
```
# Test set predictions
```
x_test = np.load(database_base_path + 'x_test.npy')
test_preds = model.predict(get_test_dataset(x_test, config['BATCH_SIZE'], AUTO))
submission = pd.read_csv('/kaggle/input/jigsaw-multilingual-toxic-comment-classification/sample_submission.csv')
submission['toxic'] = test_preds
submission.to_csv('submission.csv', index=False)
display(submission.describe())
display(submission.head(10))
```
| github_jupyter |
# Introduction to Deep Learning with PyTorch
In this notebook, you'll get introduced to [PyTorch](http://pytorch.org/), a framework for building and training neural networks. PyTorch in a lot of ways behaves like the arrays you love from Numpy. These Numpy arrays, after all, are just tensors. PyTorch takes these tensors and makes it simple to move them to GPUs for the faster processing needed when training neural networks. It also provides a module that automatically calculates gradients (for backpropagation!) and another module specifically for building neural networks. All together, PyTorch ends up being more coherent with Python and the Numpy/Scipy stack compared to TensorFlow and other frameworks.
## Neural Networks
Deep Learning is based on artificial neural networks which have been around in some form since the late 1950s. The networks are built from individual parts approximating neurons, typically called units or simply "neurons." Each unit has some number of weighted inputs. These weighted inputs are summed together (a linear combination) then passed through an activation function to get the unit's output.
<img src="assets/simple_neuron.png" width=400px>
Mathematically this looks like:
$$
\begin{align}
y &= f(w_1 x_1 + w_2 x_2 + b) \\
y &= f\left(\sum_i w_i x_i +b \right)
\end{align}
$$
With vectors this is the dot/inner product of two vectors:
$$
h = \begin{bmatrix}
x_1 \, x_2 \cdots x_n
\end{bmatrix}
\cdot
\begin{bmatrix}
w_1 \\
w_2 \\
\vdots \\
w_n
\end{bmatrix}
$$
## Tensors
It turns out neural network computations are just a bunch of linear algebra operations on *tensors*, a generalization of matrices. A vector is a 1-dimensional tensor, a matrix is a 2-dimensional tensor, an array with three indices is a 3-dimensional tensor (RGB color images for example). The fundamental data structure for neural networks are tensors and PyTorch (as well as pretty much every other deep learning framework) is built around tensors.
<img src="assets/tensor_examples.svg" width=600px>
With the basics covered, it's time to explore how we can use PyTorch to build a simple neural network.
```
# First, import PyTorch
import torch
def activation(x):
""" Sigmoid activation function
Arguments
---------
x: torch.Tensor
"""
return 1/(1+torch.exp(-x))
### Generate some data
torch.manual_seed(7) # Set the random seed so things are predictable
# Features are 5 random normal variables
features = torch.randn((1, 5))
# True weights for our data, random normal variables again
weights = torch.randn_like(features)
# and a true bias term
bias = torch.randn((1, 1))
```
Above I generated data we can use to get the output of our simple network. This is all just random for now, going forward we'll start using normal data. Going through each relevant line:
`features = torch.randn((1, 5))` creates a tensor with shape `(1, 5)`, one row and five columns, that contains values randomly distributed according to the normal distribution with a mean of zero and standard deviation of one.
`weights = torch.randn_like(features)` creates another tensor with the same shape as `features`, again containing values from a normal distribution.
Finally, `bias = torch.randn((1, 1))` creates a single value from a normal distribution.
PyTorch tensors can be added, multiplied, subtracted, etc, just like Numpy arrays. In general, you'll use PyTorch tensors pretty much the same way you'd use Numpy arrays. They come with some nice benefits though such as GPU acceleration which we'll get to later. For now, use the generated data to calculate the output of this simple single layer network.
> **Exercise**: Calculate the output of the network with input features `features`, weights `weights`, and bias `bias`. Similar to Numpy, PyTorch has a [`torch.sum()`](https://pytorch.org/docs/stable/torch.html#torch.sum) function, as well as a `.sum()` method on tensors, for taking sums. Use the function `activation` defined above as the activation function.
```
## Calculate the output of this network using the weights and bias tensors
y = activation(torch.sum(features * weights) + bias)
y = activation((features * weights).sum() + bias)
y
```
You can do the multiplication and sum in the same operation using a matrix multiplication. In general, you'll want to use matrix multiplications since they are more efficient and accelerated using modern libraries and high-performance computing on GPUs.
Here, we want to do a matrix multiplication of the features and the weights. For this we can use [`torch.mm()`](https://pytorch.org/docs/stable/torch.html#torch.mm) or [`torch.matmul()`](https://pytorch.org/docs/stable/torch.html#torch.matmul) which is somewhat more complicated and supports broadcasting. If we try to do it with `features` and `weights` as they are, we'll get an error
```python
>> torch.mm(features, weights)
---------------------------------------------------------------------------
RuntimeError Traceback (most recent call last)
<ipython-input-13-15d592eb5279> in <module>()
----> 1 torch.mm(features, weights)
RuntimeError: size mismatch, m1: [1 x 5], m2: [1 x 5] at /Users/soumith/minicondabuild3/conda-bld/pytorch_1524590658547/work/aten/src/TH/generic/THTensorMath.c:2033
```
As you're building neural networks in any framework, you'll see this often. Really often. What's happening here is our tensors aren't the correct shapes to perform a matrix multiplication. Remember that for matrix multiplications, the number of columns in the first tensor must equal to the number of rows in the second column. Both `features` and `weights` have the same shape, `(1, 5)`. This means we need to change the shape of `weights` to get the matrix multiplication to work.
**Note:** To see the shape of a tensor called `tensor`, use `tensor.shape`. If you're building neural networks, you'll be using this method often.
There are a few options here: [`weights.reshape()`](https://pytorch.org/docs/stable/tensors.html#torch.Tensor.reshape), [`weights.resize_()`](https://pytorch.org/docs/stable/tensors.html#torch.Tensor.resize_), and [`weights.view()`](https://pytorch.org/docs/stable/tensors.html#torch.Tensor.view).
* `weights.reshape(a, b)` will return a new tensor with the same data as `weights` with size `(a, b)` sometimes, and sometimes a clone, as in it copies the data to another part of memory.
* `weights.resize_(a, b)` returns the same tensor with a different shape. However, if the new shape results in fewer elements than the original tensor, some elements will be removed from the tensor (but not from memory). If the new shape results in more elements than the original tensor, new elements will be uninitialized in memory. Here I should note that the underscore at the end of the method denotes that this method is performed **in-place**. Here is a great forum thread to [read more about in-place operations](https://discuss.pytorch.org/t/what-is-in-place-operation/16244) in PyTorch.
* `weights.view(a, b)` will return a new tensor with the same data as `weights` with size `(a, b)`.
I usually use `.view()`, but any of the three methods will work for this. So, now we can reshape `weights` to have five rows and one column with something like `weights.view(5, 1)`.
> **Exercise**: Calculate the output of our little network using matrix multiplication.
```
## Calculate the output of this network using matrix multiplication
y = activation(torch.mm(weights,features.view(5,1))+bias)
y
```
### Stack them up!
That's how you can calculate the output for a single neuron. The real power of this algorithm happens when you start stacking these individual units into layers and stacks of layers, into a network of neurons. The output of one layer of neurons becomes the input for the next layer. With multiple input units and output units, we now need to express the weights as a matrix.
<img src='assets/multilayer_diagram_weights.png' width=450px>
The first layer shown on the bottom here are the inputs, understandably called the **input layer**. The middle layer is called the **hidden layer**, and the final layer (on the right) is the **output layer**. We can express this network mathematically with matrices again and use matrix multiplication to get linear combinations for each unit in one operation. For example, the hidden layer ($h_1$ and $h_2$ here) can be calculated
$$
\vec{h} = [h_1 \, h_2] =
\begin{bmatrix}
x_1 \, x_2 \cdots \, x_n
\end{bmatrix}
\cdot
\begin{bmatrix}
w_{11} & w_{12} \\
w_{21} &w_{22} \\
\vdots &\vdots \\
w_{n1} &w_{n2}
\end{bmatrix}
$$
The output for this small network is found by treating the hidden layer as inputs for the output unit. The network output is expressed simply
$$
y = f_2 \! \left(\, f_1 \! \left(\vec{x} \, \mathbf{W_1}\right) \mathbf{W_2} \right)
$$
```
### Generate some data
torch.manual_seed(7) # Set the random seed so things are predictable
# Features are 3 random normal variables
features = torch.randn((1, 3))
# Define the size of each layer in our network
n_input = features.shape[1] # Number of input units, must match number of input features
n_hidden = 2 # Number of hidden units
n_output = 1 # Number of output units
# Weights for inputs to hidden layer
W1 = torch.randn(n_input, n_hidden)
# Weights for hidden layer to output layer
W2 = torch.randn(n_hidden, n_output)
# and bias terms for hidden and output layers
B1 = torch.randn((1, n_hidden))
B2 = torch.randn((1, n_output))
```
> **Exercise:** Calculate the output for this multi-layer network using the weights `W1` & `W2`, and the biases, `B1` & `B2`.
```
## Your solution here
h = activation(torch.mm(features, W1) + B1)
output = activation(torch.mm(h, W2) + B2)
print(output)
```
If you did this correctly, you should see the output `tensor([[ 0.3171]])`.
The number of hidden units a parameter of the network, often called a **hyperparameter** to differentiate it from the weights and biases parameters. As you'll see later when we discuss training a neural network, the more hidden units a network has, and the more layers, the better able it is to learn from data and make accurate predictions.
## Numpy to Torch and back
Special bonus section! PyTorch has a great feature for converting between Numpy arrays and Torch tensors. To create a tensor from a Numpy array, use `torch.from_numpy()`. To convert a tensor to a Numpy array, use the `.numpy()` method.
```
import numpy as np
a = np.random.rand(4,3)
a
b = torch.from_numpy(a)
b
b.numpy()
```
The memory is shared between the Numpy array and Torch tensor, so if you change the values in-place of one object, the other will change as well.
```
# Multiply PyTorch Tensor by 2, in place
b.mul_(2)
# Numpy array matches new values from Tensor
a
```
| github_jupyter |
<h1>Table of Contents<span class="tocSkip"></span></h1>
<div class="toc"><ul class="toc-item"><li><span><a href="#3.8-Classification-연습문제" data-toc-modified-id="3.8-Classification-연습문제-1">3.8 Classification 연습문제</a></span><ul class="toc-item"><li><span><a href="#(Practice3)-타이타닉-데이터셋에-도전(Kaggle)" data-toc-modified-id="(Practice3)-타이타닉-데이터셋에-도전(Kaggle)-1.1">(Practice3) 타이타닉 데이터셋에 도전(Kaggle)</a></span><ul class="toc-item"><li><span><a href="#데이터를-적재합니다." data-toc-modified-id="데이터를-적재합니다.-1.1.1">데이터를 적재합니다.</a></span></li><li><span><a href="#데이터-살펴보기" data-toc-modified-id="데이터-살펴보기-1.1.2">데이터 살펴보기</a></span><ul class="toc-item"><li><span><a href="#속성은-다음과-같은-의미를-가집니다:" data-toc-modified-id="속성은-다음과-같은-의미를-가집니다:-1.1.2.1">속성은 다음과 같은 의미를 가집니다:</a></span></li><li><span><a href="#누락된-데이터가-얼마나-되는지-알아보겠습니다." data-toc-modified-id="누락된-데이터가-얼마나-되는지-알아보겠습니다.-1.1.2.2">누락된 데이터가 얼마나 되는지 알아보겠습니다.</a></span></li><li><span><a href="#훈련-데이터의-숫자형-특성들의-통계치를-살펴-보겠습니다." data-toc-modified-id="훈련-데이터의-숫자형-특성들의-통계치를-살펴-보겠습니다.-1.1.2.3">훈련 데이터의 숫자형 특성들의 통계치를 살펴 보겠습니다.</a></span></li><li><span><a href="#특성별-생존자-구성-비율을-분석해봅니다." data-toc-modified-id="특성별-생존자-구성-비율을-분석해봅니다.-1.1.2.4">특성별 생존자 구성 비율을 분석해봅니다.</a></span></li></ul></li><li><span><a href="#Feature-engineering" data-toc-modified-id="Feature-engineering-1.1.3">Feature engineering</a></span><ul class="toc-item"><li><span><a href="#Name의-호칭-부분을-추출하여-Initial-특성을-추가합니다." data-toc-modified-id="Name의-호칭-부분을-추출하여-Initial-특성을-추가합니다.-1.1.3.1">Name의 호칭 부분을 추출하여 Initial 특성을 추가합니다.</a></span></li><li><span><a href="#누락된-Age-값을-Initial(호칭)별-중간값으로-채웁니다." data-toc-modified-id="누락된-Age-값을-Initial(호칭)별-중간값으로-채웁니다.-1.1.3.2">누락된 Age 값을 Initial(호칭)별 중간값으로 채웁니다.</a></span></li><li><span><a href="#누락된-Fare값을-Pclass의-평균값(mean)으로으로-채웁니다." data-toc-modified-id="누락된-Fare값을-Pclass의-평균값(mean)으로으로-채웁니다.-1.1.3.3">누락된 Fare값을 Pclass의 평균값(mean)으로으로 채웁니다.</a></span></li><li><span><a href="#누락된-Embarked-값은-가장-많은-범주-값(most_frequent)으로-채웁니다." data-toc-modified-id="누락된-Embarked-값은-가장-많은-범주-값(most_frequent)으로-채웁니다.-1.1.3.4">누락된 Embarked 값은 가장 많은 범주 값(most_frequent)으로 채웁니다.</a></span></li><li><span><a href="#Cabin-특성의-누락된-데이터를-채우는-방법을-찾아봅니다." data-toc-modified-id="Cabin-특성의-누락된-데이터를-채우는-방법을-찾아봅니다.-1.1.3.5">Cabin 특성의 누락된 데이터를 채우는 방법을 찾아봅니다.</a></span></li><li><span><a href="#Pclass와-Cabin을-이용해-갑판(Deck)-특성을-만듭니다." data-toc-modified-id="Pclass와-Cabin을-이용해-갑판(Deck)-특성을-만듭니다.-1.1.3.6">Pclass와 Cabin을 이용해 갑판(Deck) 특성을 만듭니다.</a></span></li><li><span><a href="#숫자형-Age-특성으로-부터-범주형-AgeBucket을-만듭니다." data-toc-modified-id="숫자형-Age-특성으로-부터-범주형-AgeBucket을-만듭니다.-1.1.3.7">숫자형 Age 특성으로 부터 범주형 AgeBucket을 만듭니다.</a></span></li><li><span><a href="#형제배우자와-부모자녀-특성을-더해서-FamilySize-특성을-만듭니다." data-toc-modified-id="형제배우자와-부모자녀-특성을-더해서-FamilySize-특성을-만듭니다.-1.1.3.8">형제배우자와 부모자녀 특성을 더해서 FamilySize 특성을 만듭니다.</a></span></li><li><span><a href="#Ticket을-분석해서-TicketNumber와-TicketCode를-만듭니다." data-toc-modified-id="Ticket을-분석해서-TicketNumber와-TicketCode를-만듭니다.-1.1.3.9">Ticket을 분석해서 TicketNumber와 TicketCode를 만듭니다.</a></span></li><li><span><a href="#TicketNumber를-티켓번호-구간을-기준으로-범주형으로-변환-합니다." data-toc-modified-id="TicketNumber를-티켓번호-구간을-기준으로-범주형으로-변환-합니다.-1.1.3.10">TicketNumber를 티켓번호 구간을 기준으로 범주형으로 변환 합니다.</a></span></li><li><span><a href="#숫자형-특성-Standardization(표준화)와-범주형-특성-OneHotEncoding을-수행-합니다." data-toc-modified-id="숫자형-특성-Standardization(표준화)와-범주형-특성-OneHotEncoding을-수행-합니다.-1.1.3.11">숫자형 특성 Standardization(표준화)와 범주형 특성 OneHotEncoding을 수행 합니다.</a></span></li><li><span><a href="#전처리가-끝난-titanic_data에서-훈련-데이터와-테스트-데이터를-분리-합니다." data-toc-modified-id="전처리가-끝난-titanic_data에서-훈련-데이터와-테스트-데이터를-분리-합니다.-1.1.3.12">전처리가 끝난 titanic_data에서 훈련 데이터와 테스트 데이터를 분리 합니다.</a></span></li></ul></li><li><span><a href="#모델을-훈련-합니다." data-toc-modified-id="모델을-훈련-합니다.-1.1.4">모델을 훈련 합니다.</a></span><ul class="toc-item"><li><span><a href="#SVM" data-toc-modified-id="SVM-1.1.4.1">SVM</a></span></li><li><span><a href="#kNN" data-toc-modified-id="kNN-1.1.4.2">kNN</a></span></li><li><span><a href="#Decision-Tree" data-toc-modified-id="Decision-Tree-1.1.4.3">Decision Tree</a></span></li><li><span><a href="#Ramdom-Forest" data-toc-modified-id="Ramdom-Forest-1.1.4.4">Ramdom Forest</a></span></li><li><span><a href="#그리드-탐색으로-최적의-하이퍼파라미터를-튜닝-해봅니다." data-toc-modified-id="그리드-탐색으로-최적의-하이퍼파라미터를-튜닝-해봅니다.-1.1.4.5">그리드 탐색으로 최적의 하이퍼파라미터를 튜닝 해봅니다.</a></span></li></ul></li><li><span><a href="#테스트-데이터에-대한-예측을-만들어-Kaggle에-업로드-합니다." data-toc-modified-id="테스트-데이터에-대한-예측을-만들어-Kaggle에-업로드-합니다.-1.1.5">테스트 데이터에 대한 예측을 만들어 Kaggle에 업로드 합니다.</a></span></li></ul></li></ul></li></ul></div>
## 3.8 Classification 연습문제
### (Practice3) 타이타닉 데이터셋에 도전(Kaggle)
승객의 나이, 성별, 승객 등급, 승선 위치 같은 속성을 기반으로 하여 승객의 생존 여부를 예측하는 것이 목표입니다.
먼저 [캐글](https://www.kaggle.com)에 로그인 하고, [타이타닉 챌린지](https://www.kaggle.com/c/titanic)에서 `train.csv`와 `test.csv`를 다운로드합니다. 두 파일을 `datasets/titanic` 디렉토리에 저장하세요.
#### 데이터를 적재합니다.
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import warnings
%matplotlib inline
sns.set() # setting seaborn default for plots
warnings.filterwarnings("ignore", category=FutureWarning)
warnings.filterwarnings("ignore", category=DeprecationWarning)
train_data = pd.read_csv("datasets/titanic/train.csv")
test_data = pd.read_csv("datasets/titanic/test.csv")
titanic_data = pd.concat([train_data, test_data])
```
- 데이터 분석에는 train_data를 사용하고, <br>
데이터 전처리는 titanic_data로 한꺼번에 처리 후 모델 훈련 전에 훈련 데이터와 테스트 데이터를 분리해줍니다.
#### 데이터 살펴보기
```
train_data.shape
test_data.shape
titanic_data.head()
```
##### 속성은 다음과 같은 의미를 가집니다:
* **Survived**: 타깃입니다. 0은 생존하지 못한 것이고 1은 생존을 의미합니다.
* **Pclass**: 승객 등급. 1, 2, 3등석.
* **Name**, **Sex**, **Age**: 이름 그대로 의미입니다.
* **SibSp**: 함께 탑승한 형제, 배우자의 수.
* **Parch**: 함께 탑승한 자녀, 부모의 수.
* **Ticket**: 티켓 아이디
* **Fare**: 티켓 요금 (파운드)
* **Cabin**: 객실 번호
* **Embarked**: 승객이 탑승한 곳. C(Cherbourg), Q(Queenstown), S(Southampton)
##### 누락된 데이터가 얼마나 되는지 알아보겠습니다.
```
titanic_data.info()
```
- Survived는 출력 레이블 입니다.
- Age, Cabin, Embarked, Fare 특성은 일부 데이터가 Null 입니다.
- Object형 특성 중에 Name, Ticket, Cabin은 Text형 특성이고, Sex, Embarked는 범주형 특성 입니다.
- int64형 특성 중에 Pclass는 범주형 특성 입니다.
- 1,309개 데이터 중에 891개는 label값(Survived)이 있는 훈련 데이터 이고, 418개는 테스트 데이터 입니다.
##### 훈련 데이터의 숫자형 특성들의 통계치를 살펴 보겠습니다.
```
train_data.describe()
```
- 생존자는 38% 정도 입니다.
- 부모자식 수는 최대 6명, 형제/배우자 수는 최대 8명, 나이는 0.42~80.0세 까지 분포해있습니다.
##### 특성별 생존자 구성 비율을 분석해봅니다.
```
# 특성별 생존자 구성을 분석하기 위한 바-차트 함수를 만듭니다.
def bar_chart(feature):
survived = train_data[train_data['Survived']==1][feature].value_counts()
dead = train_data[train_data['Survived']==0][feature].value_counts()
df = pd.DataFrame([survived,dead])
df.index = ['Survived', 'Dead']
df = df.T
df.plot(kind='bar', stacked=True, figsize=(10,5))
bar_chart("Sex")
```
- 남성의 사망자 비율이 높게 보입니다.
```
bar_chart("Pclass")
```
- 3등급의 사망자 비율이 높게 보입니다.
```
bar_chart("Embarked")
```
- S(Southampton) 탑승자들의 사망 비율이 상대적으로 높습니다.
#### Feature engineering
Age, Cabin, Embarked, Fare 특성의 누락된 데이터를 적당한 값으로 채우고, 더 가치가 있는 새로운 특성들을 만듭니다.
##### Name의 호칭 부분을 추출하여 Initial 특성을 추가합니다.
Age는 중간값(media) 값으로 채울 수도 있지만, <br>
**`Name`** 컬럼의 Initial(호칭)을 분석해서 **`Age`** 컬럼의 누락된 데이트를 Initial의 평균값으로 대체해보겠습니다.
Name의 Initial 추출은 Inintial이 알파벳 문자 뒤에 "."으로 끝나는 점을 이용해 정규표현식을 사용 합니다.
```
titanic_data["Name"].head()
titanic_data["Initial"] = titanic_data.Name.str.extract('([A-Za-z]+)\.')
titanic_data["Initial"].value_counts()
```
Inintial의 대부분은 Mr, Miss, Mrs, Master, Dr 입니다. <br>
Mr, Miss, Mrs, Master, Dr로 구분 가능한 것들은 replace 하고, 나머지는 Others로 통합합니다.
```
titanic_data['Initial'].replace(['Mlle','Mme','Ms','Dr','Major','Lady','Countess','Jonkheer','Col','Rev','Capt','Sir','Don', 'Dona'],
['Miss','Mrs','Miss','Dr','Mr','Mrs','Mrs','Others','Others','Others','Mr','Mr','Mr', 'Mrs'], inplace=True)
# "Mr", "Miss", "Mrs", "Master", "Dr"를 제외한 모두를 "Others"로 간단하게 변환도 가능
# titanic_data["Initial"] = titanic_data["Initial"].apply(lambda x: x if x in ["Mr", "Miss", "Mrs", "Master", "Dr"] else "Others")
titanic_data["Initial"].value_counts()
```
##### 누락된 Age 값을 Initial(호칭)별 중간값으로 채웁니다.
```
titanic_data[["Initial", "Age"]].groupby(["Initial"]).median()
titanic_data["Age"] = titanic_data.groupby("Initial").transform(lambda x: x.fillna(x.median()))["Age"]
```
##### 누락된 Fare값을 Pclass의 평균값(mean)으로으로 채웁니다.
```
titanic_data[["Pclass", "Fare"]].groupby(["Pclass"]).mean()
titanic_data["Fare"] = titanic_data.groupby("Pclass").transform(lambda x: x.fillna(x.mean()))["Fare"]
```
##### 누락된 Embarked 값은 가장 많은 범주 값(most_frequent)으로 채웁니다.
```
titanic_data["Embarked"].value_counts()
from sklearn.impute import SimpleImputer
imputer = SimpleImputer(strategy="most_frequent")
titanic_data["Embarked"] = imputer.fit_transform(titanic_data["Embarked"].reshape(-1,1))
titanic_data["Embarked"].value_counts()
```
##### Cabin 특성의 누락된 데이터를 채우는 방법을 찾아봅니다.
```
titanic_data["Cabin"].value_counts()[:20]
```
`Cabin에는 객실의 갑판(Deck) 정보가 맨 앞 영문자에 포함되어 있습니다.`
타이타닉의 각 층(갑판)은 아래와 같이 이루어져 있었다. (https://ko.wikipedia.org/wiki/RMS_타이타닉)
- **보트 갑판** - 최상층으로, 구명보트들은 모두 여기에 나열되어 있었다. <br>
맨 앞쪽에는 선교와 조타실이 있었으며 선장 & 간부(항해사) 숙소와 바로 연결되어 있었다. <br>
그리고 그 뒤에는 중앙계단의 최상층과 체육관이 연결되어 있었다. <br>
넓은 산책로가 있었으며, 1등실, 2등실, 항해사, 기관사마다 산책로의 영역이 정해져 있었다. <br>
1등실 산책로의 경우 외관을 잘 볼 수 있게 하기 위해 구명보트가 설치되어 있지 않았는데, 이것이 참사의 원인 중 하나가 되었다.
- **A 갑판** - `산책 갑판`이라고도 불렸다. 거의 모든 영역이 1등실 전용이었고 1등실 객실과 라운지, 흡연실, 독서실, 야자수 코트가 있었다. <br>
산책 갑판이라는 이름답게 기나긴 산책로가 있었다.
- **B 갑판** - 선교루 갑판이라고도 불렸다. 객실은 `1등실이 상당수를 차지`하고 있었으며, 어떤 객실에는 개인 산책로가 있었다. <br>
레스토랑과 카페도 있었다. 이 갑판에는 선두와 선미 윗쪽도 포함하고 있는데, 선미는 3등실 산책로로 쓰였다
- **C 갑판** - 배에서 선수, 선미와 분리되는 지점이자 하얗게 칠한 곳이 시작하는 지점으로, <br>
3등실의 산책로로 쓰이고 선수와 선미로 각각 이어지는 요갑판과 이어져 있었다. <br>
`선수는 주로 선원들의 숙소`로 쓰였고, 선미는 3등실 전용 휴게실과 흡연실이 있었다. 그 사이에는 `1등실 객실과 2등실 도서관`이 있었다.
- **D 갑판** - 3개의 넓은 공공시설이 상당부분을 차지하고 있었는데, 1등실 대합실, 1등실 식당과 2등실 식당이 있었다. <br>
3등실을 위한 열린 공간도 있어서 밤만 되면 신나는 연회가 열렸다. `1등실, 2등실, 3등실 객실`과 화부 숙소가 있던 곳이기도 했다. <br>
`방수격벽이 위치해 있던 가장 높은 갑판` 이기도 했다.
- **E 갑판** - 주로 `1등실, 2등실, 3등실 모두의 객실과 선원 숙소`가 있었다. <br>
"스코틀랜드 로드"라고 불리던 아주 긴 복도가 있어 선원과 승객이 오고갈수 있었다.<br>
(스코틀랜드 로드는 후에 타이타닉 선체 내부에 물이 들어차기 시작했을때 E갑판의 선수쪽 전체를 침수시키고 방수격실을 무력화시키는 원인 중 하나였다.)
- **F 갑판** - 객실은 `3등실이 대부분`을 차지했지만 `2등실 객실과 선원 숙소`도 있었다. <br>
3등실 식당은 여기에 위치했다. 또 1등실 승객을 위한 수영장과 터키탕도 있었다.
- **G 갑판** - 수면 위에서 가장 낮은 층으로, `선원과 승객 객실이 있는 가장 낮은 갑판`이었다. <br>
스쿼시 코트도 있었으며 우편 저장소도 여기에 있었다. 대다수의 영역이 보일러실이 있는 최하 갑판과 겹쳐 있어 선미와 후미가 각각 따로 떨어져 막혀 있었다.
- **최하 갑판** : 주로 창고가 대부분이었다.
- **탱크 톱** : 보일러실과 기관실이 있었다.
##### Pclass와 Cabin을 이용해 갑판(Deck) 특성을 만듭니다.
Cabin으로 부터 Deck 특성을 추출하고,<br>
Pclass를 숫자형 범주 값으로 변경한 후 Deck의 누락된 데이터를 Pclass 중간값으로 채웁니다.
```
titanic_data["Deck"] = titanic_data.Cabin.str[0]
titanic_data["Deck"].value_counts()
titanic_data["Deck"].replace(['A', 'B', 'C', 'D', 'E', 'F', 'G', 'T'], [1, 2, 3, 4, 5, 6, 7, 8], inplace=True)
titanic_data["Deck"].fillna(titanic_data.groupby("Pclass")["Deck"].transform("median"), inplace=True)
titanic_data["Deck"].value_counts()
```
##### 숫자형 Age 특성으로 부터 범주형 AgeBucket을 만듭니다.
```
# 간단하게 일정 구간을 나눠서 범주형으로 변환할 수도 있습니다.
titanic_data["AgeBucket"] = titanic_data["Age"] // 15 * 15
titanic_data[["AgeBucket", "Survived"]].groupby(['AgeBucket']).mean()
```
AgeBucket을 좀더 정확한 범주형 특성으로 만들기 위해 Age의 구간별 특성을 분석해봅니다.
```
facet = sns.FacetGrid(titanic_data, hue="Survived", aspect=4)
facet.map(sns.kdeplot, 'Age', shade= True)
facet.set(xlim=(0, train_data['Age'].max()))
facet.add_legend()
facet = sns.FacetGrid(titanic_data, hue="Survived", aspect=4)
facet.map(sns.kdeplot, 'Age', shade= True)
facet.set(xlim=(0, train_data['Age'].max()))
facet.add_legend()
plt.xlim(0, 20)
facet = sns.FacetGrid(titanic_data, hue="Survived", aspect=4)
facet.map(sns.kdeplot, 'Age', shade= True)
facet.set(xlim=(0, train_data['Age'].max()))
facet.add_legend()
plt.xlim(20, 40)
facet = sns.FacetGrid(titanic_data, hue="Survived", aspect=4)
facet.map(sns.kdeplot, 'Age', shade= True)
facet.set(xlim=(0, train_data['Age'].max()))
facet.add_legend()
plt.xlim(40, 60)
facet = sns.FacetGrid(titanic_data, hue="Survived", aspect=4)
facet.map(sns.kdeplot, 'Age', shade= True)
facet.set(xlim=(0, train_data['Age'].max()))
facet.add_legend()
plt.xlim(60, 80)
```
생존자와 사망자의 비율이 변동되는 나이를 기준으로 Age의 그룹을 나누어 AgeBucket을 만듭니다.
```
titanic_data.loc[titanic_data['Age'] <= 17, 'AgeBucket'] = 0
titanic_data.loc[(titanic_data['Age'] > 17) & (titanic_data['Age'] <= 34), 'AgeBucket'] = 1
titanic_data.loc[(titanic_data['Age'] > 34) & (titanic_data['Age'] <= 26), 'AgeBucket'] = 2
titanic_data.loc[(titanic_data['Age'] > 46) & (titanic_data['Age'] <= 60), 'AgeBucket'] = 3
titanic_data.loc[titanic_data['Age'] > 60, 'AgeBucket'] = 4
titanic_data["AgeBucket"].value_counts()
```
##### 형제배우자와 부모자녀 특성을 더해서 FamilySize 특성을 만듭니다.
```
titanic_data["FamilySize"] = titanic_data["SibSp"] + titanic_data["Parch"]
titanic_data[["FamilySize", "Survived"]].groupby(['FamilySize']).mean()
```
##### Ticket을 분석해서 TicketNumber와 TicketCode를 만듭니다.
```
titanic_data["Ticket"].value_counts()[:30]
```
Ticket 컬럼을 보면 알파벳 코드가 있는 티켓과 없는 티켓이 있고, 일정한 범위의 숫자로 구성된 것으로 보입니다.<br>
"LINE" 처럼 숫자가 없는 문자로 만 구성된 Ticket도 보입니다.
알파벳 코드와 티켓 일련번호를 공백을 기준으로 구분해봅니다. 코드가 없는 데이터는 "NoCode"로 대체 합니다.
```
titanic_data["TicketNumber"] = titanic_data["Ticket"].apply(lambda x: x.split()[-1])
titanic_data["TicketNumber"] = titanic_data["TicketNumber"].replace("LINE", "0")
titanic_data["TicketCode"] = titanic_data["Ticket"].apply(lambda x: x.split()[0] if len(x.split())!= 1 else "NoCode")
titanic_data["TicketCode"].value_counts()[:20]
```
TicketCode는 의미를 해석할 수 없어서 일단 보류합니다.
##### TicketNumber를 티켓번호 구간을 기준으로 범주형으로 변환 합니다.
TicketNumber 데이터를 히스토그램으로 확인해봅니다.
```
titanic_data["TicketNumber"].astype('int64').hist(bins=100)
plt.show()
```
0부터 500,000 이하에 값이 집중되어 있고, 3,000,000 이상의 값도 일부 있습니다.<br>
십만단위의 범위로 범주형 데이터를 만들어 봅니다.
```
titanic_data["TicketNumber"] = pd.to_numeric(titanic_data["TicketNumber"]) // 100000 * 100000
titanic_data["TicketNumber"].value_counts()
```
##### 숫자형 특성 Standardization(표준화)와 범주형 특성 OneHotEncoding을 수행 합니다.
```
# 이제 불필요한 특성을 지웁니다. (Age 특성을 AgeBucket과 함께 유지합니다.)
titanic_data_imputed = titanic_data.drop(["Cabin", "Name", "Parch", "SibSp", "PassengerId", "Survived", "Ticket", "TicketCode"], axis=1)
titanic_data_imputed.info()
titanic_data_imputed.columns
```
숫자 특성을 표준화 하는 파이프라인과 범주형 특성을 OneHotEncoding 하는 파이프라인 각각 만들고,<br>
사이킷런의 **`ColumnTransformer`** 클래스를 사용하여 두 개의 파이프라인을 연결합니다.
`StandardScaler`는 숫자형 데이터를 표준편차가 1이 되도록 조정 합니다.<br>
`OneHotEncoder`는 정수형 또는 문자열 범주에 대해 각 범주형 값을 원-핫 벡터로 변경합니다.
```
from sklearn.pipeline import Pipeline
from sklearn.compose import ColumnTransformer
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import OneHotEncoder
num_pipeline = Pipeline([
("std_scaler", StandardScaler()),
])
cat_pipeline = Pipeline([
("cat_encoder", OneHotEncoder(sparse=False)),
])
preprocess_pipeline = ColumnTransformer([
("num_pipeline", num_pipeline, ["Age", "Fare"]),
("cat_pipeline", cat_pipeline, ['Embarked', 'Pclass', 'Sex', 'Initial', 'Deck', 'AgeBucket', 'FamilySize', 'TicketNumber']),
])
titanic_data_preprocessed = preprocess_pipeline.fit_transform(titanic_data_imputed)
titanic_data_preprocessed.shape
```
##### 전처리가 끝난 titanic_data에서 훈련 데이터와 테스트 데이터를 분리 합니다.
```
# 훈련 데이터와 테스트 데이터로 구분합니다.
X_train = titanic_data_preprocessed[:891]
X_test = titanic_data_preprocessed[891:]
# 훈련 데이터에 대한 레이블을 추출합니다.
y_train = train_data["Survived"]
```
이제 원본 데이터를 받아 머신러닝 모델에 주입할 숫자 입력 특성으로 구성된 훈련 세트와 테스트 세트가 만들어졌습니다.
#### 모델을 훈련 합니다.
이제 분류기를 훈련시킬 차례입니다.
```
from sklearn.svm import SVC
from sklearn.neighbors import KNeighborsClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.model_selection import KFold
from sklearn.model_selection import cross_val_score
k_fold = KFold(n_splits=10, shuffle=True, random_state=42)
```
##### SVM
먼저 `SVC`를 사용해 보겠습니다.
```
svm_clf = SVC(gamma='auto')
svm_clf.fit(X_train, y_train)
```
훈련된 모델을 사용해서 테스트 세트에 대한 예측을 만들어 Kaggle에 바로 업로드 할 수 있지만, <br>
교차 검증으로 모델이 얼마나 좋은지 평가해봅니다.<br>
정밀도, 재현율, F1 점수도 확인해봅니다.
```
svm_scores = cross_val_score(svm_clf, X_train, y_train, cv=k_fold, n_jobs=1, scoring='accuracy')
svm_scores.mean()
from sklearn.model_selection import cross_val_predict
y_train_svc = cross_val_predict(svm_clf, X_train, y_train, cv=10)
from sklearn.metrics import confusion_matrix
confusion_matrix(y_train, y_train_svc)
from sklearn.metrics import precision_score, recall_score
precision_score(y_train, y_train_svc)
recall_score(y_train, y_train_svc)
from sklearn.metrics import f1_score
f1_score(y_train, y_train_svc)
```
정확도 79%, 정밀도 75% 수준 입니다. 무작위로 선택한 것보다는 좋지만 아주 높은 점수는 아닙니다.<br>
캐글에서 타이타닉 경연 대회의 [리더보드](https://www.kaggle.com/c/titanic/leaderboard)를 보면 상위 10%내에 들려면 80% 이상의 정확도를 내야합니다.
다른 알고리즘들을 적용해 좀 더 정확도를 개선한 모델을 만들어 보겠습니다.
##### kNN
```
knn_clf = KNeighborsClassifier(n_neighbors = 10)
knn_scores = cross_val_score(knn_clf, X_train, y_train, cv=k_fold, n_jobs=1, scoring='accuracy')
knn_scores.mean()
y_train_knn = cross_val_predict(knn_clf, X_train, y_train, cv=10)
precision_score(y_train, y_train_knn)
f1_score(y_train, y_train_knn)
```
##### Decision Tree
```
decision_clf = DecisionTreeClassifier()
decision_scores = cross_val_score(decision_clf, X_train, y_train, cv=k_fold, n_jobs=1, scoring='accuracy')
decision_scores.mean()
y_train_decision = cross_val_predict(decision_clf, X_train, y_train, cv=10)
precision_score(y_train, y_train_decision)
f1_score(y_train, y_train_decision)
```
##### Ramdom Forest
```
forest_clf = RandomForestClassifier(n_estimators=10, random_state=42)
forest_scores = cross_val_score(forest_clf, X_train, y_train, cv=k_fold, n_jobs=1, scoring='accuracy')
forest_scores.mean()
y_train_forest = cross_val_predict(forest_clf, X_train, y_train, cv=10)
precision_score(y_train, y_train_forest)
f1_score(y_train, y_train_forest)
```
##### 그리드 탐색으로 최적의 하이퍼파라미터를 튜닝 해봅니다.
```
from sklearn.model_selection import GridSearchCV
```
**그리드 탐색으로 `RandomForestClassifier` 하이퍼파라미터를 튜닝 해봅니다.**
```
param_grid = [
{'n_estimators': [3, 10, 20, 30, 40, 50], 'max_features': [10, 15, 20, 25, 30, 40], 'max_leaf_nodes': [5, 10, 20, 30, 40]},
{'n_estimators': [3, 10, 20, 30, 40], 'max_features': [10, 20, 25, 30, 35, 40, 42], 'min_samples_leaf': [1, 5, 10, 15]},
]
forest_clf = RandomForestClassifier(random_state=42)
grid_search_forest = GridSearchCV(forest_clf, param_grid, cv=10, iid=False,
scoring='neg_mean_squared_error',
return_train_score=True)
grid_search_forest.fit(X_train, y_train)
grid_search_forest.best_params_
best_estimator_forest = grid_search_forest.best_estimator_
best_estimator_forest
best_scores_forest = cross_val_score(best_estimator_forest, X_train, y_train, cv=k_fold, n_jobs=1, scoring='accuracy')
best_scores_forest.mean()
```
**그리드 탐색으로 `KNeighborsClassifier` 하이퍼파라미터를 튜닝 해봅니다.**
```
param_grid = [
{'n_neighbors': [3, 4, 5, 6, 7, 8, 9, 10, 15, 20]},
]
knn_clf = KNeighborsClassifier()
grid_search_knn = GridSearchCV(knn_clf, param_grid, cv=10, iid=False,
scoring='neg_mean_squared_error',
return_train_score=True)
grid_search_knn.fit(X_train, y_train)
grid_search_knn.best_params_
best_estimator_knn = grid_search_knn.best_estimator_
best_scores_knn = cross_val_score(best_estimator_knn, X_train, y_train, cv=k_fold, n_jobs=1, scoring='accuracy')
best_scores_knn.mean()
```
**그리드 탐색으로 `SVC` 하이퍼파라미터를 튜닝 해봅니다.**
```
param_grid = [
{'gamma': [0.001, 0.01, 0.1, 1, 10, 100], 'kernel': ['linear', 'poly', 'rbf', 'sigmoid']},
]
svm_clf = SVC(random_state=42)
grid_search_svm = GridSearchCV(svm_clf, param_grid, cv=10, iid=False,
scoring='neg_mean_squared_error',
return_train_score=True)
grid_search_svm.fit(X_train, y_train)
grid_search_svm.best_params_
best_estimator_svm = grid_search_svm.best_estimator_
best_scores_svm = cross_val_score(best_estimator_svm, X_train, y_train, cv=k_fold, n_jobs=1, scoring='accuracy')
best_scores_svm.mean()
```
**그리드 탐색으로 `DecisionTreeClassifier` 하이퍼파라미터를 튜닝 해봅니다.**
```
param_grid = [
{'criterion': ['gini', 'entropy'], 'max_features': [10, 15, 20, 25, 30, 40], 'max_leaf_nodes': [5, 10, 20, 30, 40, 50]},
]
decision_clf = DecisionTreeClassifier(random_state=42)
grid_search_decision = GridSearchCV(decision_clf, param_grid, cv=10, iid=False,
scoring='neg_mean_squared_error',
return_train_score=True)
grid_search_decision.fit(X_train, y_train)
grid_search_decision.best_params_
grid_search_decision = grid_search_decision.best_estimator_
best_scores_decision = cross_val_score(grid_search_decision, X_train, y_train, cv=k_fold, n_jobs=1, scoring='accuracy')
best_scores_decision.mean()
```
#### 테스트 데이터에 대한 예측을 만들어 Kaggle에 업로드 합니다.
훈련 결과가 가장 좋은 best_estimator_svm 모델로 테스트 데이터에 대한 예측을 만듭니다.
```
best_estimator_svm.fit(X_train, y_train)
y_test_pred = best_estimator_svm.predict(X_test)
```
예측 값에 PassengerId를 추가하고, Kaggle Competition에서 지정한 제목(컬럼 이름)을 추가한 후, <br>
csv 파일로 저정하여 Kaggle에 업로드 합니다.
```
submission = np.r_[np.array([["PassengerId", "Survived"]]), np.c_[test_data["PassengerId"], y_test_pred]]
np.savetxt("titanic_survive_submission.csv", submission, fmt="%s", delimiter=",")
```
best_estimator_forest 모델로도 테스트에 대한 예측을 만들어 봅니다.
```
best_estimator_forest.fit(X_train, y_train)
y_test_pred = best_estimator_forest.predict(X_test)
submission = np.r_[np.array([["PassengerId", "Survived"]]), np.c_[test_data["PassengerId"], y_test_pred]]
np.savetxt("titanic_survive_submission.csv", submission, fmt="%s", delimiter=",")
```
| github_jupyter |
# CS446/546 - Class Session 19 - Correlation networks
In this class session we are going to analyze gene expression data from a human bladder cancer cohort. We will load a data matrix of expression measurements of 4,473 genes in 414 different bladder cancer samples. These genes have been selected because they are differentially expressed between normal bladder and bladder cancer (thus more likely to have a function in bladder cancer specifically), but the columns in the data matrix are restricted to bladder cancer samples (not normal bladder) because we want to obtain a network representing variation across cancers. The measurements in the matrix have already been normalized to account for inter-sample heterogeneity and then log2 transformed. Our job is to compute Pearson correlation coefficients between all pairs of genes, obtain Fisher-transformed *z*-scores for all pairs of genes, test each pair of genes for significance of the z score, adjust for multiple hypothesis testing, filter to eliminate any pair for which *R* < 0.75 or *P*adj > 0.01, load the graph into an `igraph::Graph`, and plot the degree distribution on log-log scale. We will then answer two questions: (1) does the network look to be scale-free? and (2) what is it's best-fit scaling exponent?
Let's start by loading the packages that we will need for this notebook. Note the difference in language-design philosophy between R (which requires one package for this analysis) and python (where we have to load eight modules). Python keeps its core minimal, whereas R has a lot of statistical and plotting functions in the base language (or in packages that are loaded by default).
```
suppressPackageStartupMessages(
library(igraph)
)
```
Read the tab-deliminted text file of gene expression measurements (rows correspond to genes, columns correspond to bladder tumor samples). (use `read.table` with `row.names=1`). As always, sanity check that the file that you loaded has the expected dimensions (4,473 x 414)
```
gene_matrix_for_network <- read.table("shared/bladder_cancer_genes_tcga.txt",
sep="\t",
header=TRUE,
row.names=1,
stringsAsFactors=FALSE)
dim(gene_matrix_for_network)
```
Look up the online help for the `cor(x)` function, using the command `?cor`. When the `x` argument is a matrix, does `cor` compute the correlation coefficient of pairs of columns or pairs of rows?
```
?cor
```
Since `cor(x)` computes the correlation coefficients of pairs of *columns* of its matrix argument but we want to compute correlation coefficients of pairs of *genes* (and recall that genes are *rows* in our matrix `gene_matrix_for_network`), use the `t` function to transpose the matrix. Then use the `cor` function to compute the 4,473 x 4,473 matrix of gene-gene Pearson correlation coefficients, and call this matrix `gene_matrix_for_network_cor`. It is in your interest to do this as a one-line command (by composing the function calls), so we store only one big matrix in the environment:
```
gene_matrix_for_network_cor <- cor(t(gene_matrix_for_network))
```
Look up the online help for the `upper.tri` function. Does it return the matrix's upper triangle entries or the *index values* of the upper triangle entries of the matrix?
```
?upper.tri
```
Set the upper-triangle of the matrix `gene_matrix_for_network_cor` to zero, using (from left to right) three things: array indexing `[`, the function `upper.tri`, and the assignment operator `<-`. In the call to `upper.tri`, specify `diag=TRUE` to also zero out the diagonal.
```
gene_matrix_for_network_cor[upper.tri(gene_matrix_for_network_cor, diag=TRUE)] <- 0
```
Use the function `which` in order to obtain a vector `inds_correl_above_thresh` containing the indices of the entries of the matrix `gene_matrix_for_network_cor` for which *R* >= 0.75. Use array indexing to obtain the *R* values for these matrix entries, as a numpy array `cor_coeff_values_above_thresh`.
```
inds_correl_above_thresh <- which(gene_matrix_for_network_cor >= 0.75)
cor_coeff_values_above_thresh <- gene_matrix_for_network_cor[inds_correl_above_thresh]
```
Refer to Eq. (13.5) in the assigned readding for today's class (p9 of the PDF). Obtain a vector of the correlation coefficients that exceeded 0.75, and Fisher-transform the correlation coefficient values to get a vector `z_scores` of *z* scores. Each of these *z* scores will correspond to an **edge** in the network, unless the absolute *z* score is too small such that we can't exclude the null hypothesis that the corresponding two genes' expression values are indepdenent (we will perform that check in the next step).
```
z_scores <- 0.5*log((1+gene_matrix_for_network_cor[inds_correl_above_thresh])/
(1-gene_matrix_for_network_cor[inds_correl_above_thresh]))
```
Delete the correlation matrix object in order to save memory (we won't need it from here on out).
```
rm(gene_matrix_for_network_cor)
```
Now we are going to perform a statistical test on *each* correlation coefficient that was >= 0.75. Assume that under the null hypothesis that two genes are independent, then sqrt(M-3)z for the pair of genes is independent sample from the normal distribution with zero mean and unit variance, where M is the number of samples used to compute the Pearson correlation coefficient (i.e., M = 414). For each entry in `z_scores` compute a P value as the area under two tails of the normal distribution N(x), where the two tails are x < -sqrt(M-3)z and x > sqrt(M-3)z. (You'll know you are doing it right if z=0 means you get a P value of 1). You will want to use the functions `sqrt` and `pnorm`.
```
M <- ncol(gene_matrix_for_network)
P_values <- 2*(pnorm(-z_scores*sqrt(M-3)))
```
Look up the function `p.adjust` in the online help
```
?p.adjust
```
Adjust the P values for multiple hypothesis testing, using the `p.adjust` function with `method="hochberg"`
```
P_values_adj <- p.adjust(P_values, method="hochberg")
```
Verify that we don't need to drop any entries due to the adjusted P value not being small enough (use `which` and `length`); this should produce zero since we have M=414 samples per gene.
```
length(which(P_values_adj > 0.01))
```
Make an undirected graph from the row/column indices of the (upper-triangle) gene pairs whose correlations were above our threshold (you'll want to use the `arrayInd`, `cbind`, and `graph_from_data_frame` functions for this). Print a summary of the network, as a sanity check. Make sure to specify `directed=FALSE` when you construct the graph.
```
N <- nrow(gene_matrix_for_network)
final_network <- graph_from_data_frame(cbind(arrayInd(inds_correl_above_thresh,
.dim=c(N,N))), directed=FALSE)
summary(final_network)
```
Plot the degree distribution on log-log scale; does it appear to be scale-free?
```
suppressWarnings(
plot(degree.distribution(final_network), log="xy", xlab="k", ylab="Pk")
)
```
Use the `igraph::power.law.fit` function to estimate the scaling exponent *alpha* of the degree distribution:
```
power.law.fit(degree(final_network))$alpha
```
## extra challenge:
If you got this far, see if you can scatter plot the relationship between R (as the independent variable) and -log10(P) value (as the dependent variable). When the effect size variable (e.g., *R*) can range from negative to positive, this plot is sometimes called a "volcano plot".
```
inds_use = which(P_values_adj > 0)
plot(cor_coeff_values_above_thresh[inds_use], -log10(P_values_adj[inds_use]), xlab="R", ylab="-log10(P)")
```
## extra-extra challenge
For each of the gene pairs for which R>0.75, see if you can compute the t-test P value for each correlation coefficient (don't bother adjusting for false discovery rate control). Compare to the (un-adjusted) P values that you got using the Fisher transformation, using a scatter plot. How do they compare? Which test has better statistical power, for this case where M = 414? (If you are wondering, *general* advice is to use Fisher if M>=10; for very small numbers of samples, use the Student t test).
```
ts = cor_coeff_values_above_thresh * sqrt(M-2) / sqrt((1 - cor_coeff_values_above_thresh^2))
P_values_studentT = 2*pt(-ts, M-2)
inds_use = which(P_values > 0 & P_values_studentT > 0)
plot(-log10(P_values[inds_use]), -log10(P_values_studentT[inds_use]), xlab="Fisher transformation", ylab="Student t")
```
| github_jupyter |
# JECRC LABS
```
import pandas as pd
# import numpy as np
from numpy import * # Import all functions directly from name!
import matplotlib.pyplot as plt
import scipy as sp
```
## Lab 2: Numpy
```
# Creating a numpy array
arr = array([[1, 2, 3, 4, 5]])
print(arr)
arr.shape
type(arr)
# Creating a multi dimensional array
arr_mul_dim = array([ [1, 2, 3, 4, 5], [1, 2, 3, 4, 5] ])
arr_mul_dim.shape
arr_mul_dim
arr
arr.min()
arr.max()
arr.sum()
arr_mul_dim1 = array([[1, 2, 3, 4, 5], [0, -1, 2, 3, 4]])
arr_mul_dim1[1][1]
arr_mul_dim1.size
x = linspace(-1, 3, 5000)
# function = lambda x: x ** 3 - 3 * x ** 2 + x
function = lambda x: 100 * x ** 2 - 50 * x + 10
plt.plot(x, function(x))
plt.show()
l = [[1, 2, 3], [4, 5, 6], [7, 8, 9]]
l
l[0][1], l[1][1], l[2][1]
for i in range(3):
print(l[i][1])
import numpy as np
a = np.array([1, "Harshit Dawar", 5.7])
a
l_a = np.array(l)
l_a
# Slicing
l_a[:, 2]
l_a.sum(axis = 0)
# import numpy as np
import random
# build up 2 sets of random numbers
# setup empty array 2 rows, 500 columns
numbers = np.empty([2,500], int)
# set seed so we can repeat results
random.seed(10)
# step to populate the array
for num in range(0, 500):
numbers[0,num] = random.randint(0, 500)
numbers[1,num] = random.randint(0, 500)
# produce a histogram of the data
(hist, bins) = np.histogram(numbers, bins = 10, range = (0,500))
print ("Histogram is ",hist)
# calculate correlation between the 2 columns
corrs = np.correlate(numbers[:,1], numbers[:,2], mode='valid')
print ("Correlation of the two rows is ", corrs)
numbers.shape
l
l + [10]
l_a + 10
l_a.shape
np.random.random()
```
## Lab 3: Pandas
```
import pandas as pd
titanic_data = pd.read_csv("titanic_train.csv")
titanic_data
titanic_data.head()
titanic_data.tail()
titanic_data.head(10)
titanic_data_2 = pd.read_csv("https://vincentarelbundock.github.io/Rdatasets/csv/datasets/Titanic.csv")
titanic_data_2
amazon_data = pd.read_excel("amazonLabelled.xlsx")
amazon_data
titanic_data.isna().sum()
titanic_data.head()
titanic_data.dropna()["Age"].sort_values()
titanic_data["Age"].dropna().sort_values()
891 - 177
titanic_data.columns
titanic_data[["Age"]].groupby("Age")["Age"].count()
processed_ages = titanic_data["Age"].dropna()
processed_ages
mean_of_age = processed_ages.mean()
mean_of_age
titanic_data["Age - Mean of Age"] = abs(processed_ages - mean_of_age)
titanic_data[["Age", "Age - Mean of Age"]].dropna().head(15)
std_of_age = processed_ages.std()
outlier_threshold = std_of_age * 1.96
outlier_threshold
titanic_data["Outlier_Threshold"] = outlier_threshold
titanic_data
titanic_data[["Age", "Age - Mean of Age", "Outlier_Threshold"]].dropna().head(15)
titanic_data["Is Age an Outlier?"] = titanic_data["Age - Mean of Age"] > titanic_data["Outlier_Threshold"]
titanic_data
titanic_data[["Age", "Age - Mean of Age", "Outlier_Threshold", "Is Age an Outlier?"]].dropna().head(15)
titanic_data["Is Age an Outlier?"].value_counts()
titanic_data[titanic_data["Is Age an Outlier?"] == True][["Is Age an Outlier?", "Name"]]
import matplotlib.pyplot as plt
plt.hist(titanic_data["Age"]);
plt.figure(figsize = (15, 9))
plt.scatter(range(titanic_data.shape[0]), titanic_data["Age"])
plt.show()
```
## Lab 4: Scipy Library
```
import scipy as sc
# First Practical: Integration
from scipy.integrate import quad
cubic_calculator = lambda x: x ** 3 # => (x ** 4 / 4)
cubic_calculator(5)
quad(cubic_calculator, 1, 5)
(5 ** 4 - 1 ** 4) / 4
from scipy.optimize import minimize
demo_fun = lambda x: (x + 10) ** 2
demo_fun(-10)
minimize(demo_fun, x0 = 100)
import matplotlib.pyplot as plt
plt.scatter([1, 5, 7, 9, 11], [10, 20, 70, 90, 150])
plt.scatter(9.5, 105, c = "g")
plt.plot([1, 5, 7, 9, 11], [10, 20, 70, 90, 150], c = "r")
plt.show()
from scipy.interpolate import interp1d
interpolate = interp1d([1, 5, 7, 9, 11], [10, 20, 70, 90, 150])
interpolate(9.5)
import numpy as np
x = np.linspace(0.0, 100, 100)
y = np.sin(5 * x) + 7
plt.plot(x, y);
from scipy.fftpack import fft
fty = fft(y)
plt.plot(x, fty);
a = np.array([[1, 2], [3, 5]])
a
b = np.array([1, 2])
b
np.linalg.solve(a, b)
```
## Lab 5: Implementing Linear Regression from Scratch
```
import numpy as np
import matplotlib.pyplot as plt
# equation = 3*x + 9
x = np.arange(10, 150)
y = 3 * x + 9
plt.figure(figsize = (15, 7))
plt.scatter(x, y)
plt.plot(x, y, c = 'r')
plt.xlabel("x", size = 50)
plt.ylabel("y", size = 50)
plt.title("Custom Data for LR!", size = 50)
plt.show()
x = x.reshape(-1, 1)
y = y.reshape(-1, 1)
x.shape, y.shape
def compute_cost(weights, bias, independent_variable, dependent_variable):
N = float(independent_variable.shape[0])
total_cost = np.sum((dependent_variable - (weights * independent_variable + bias)) ** 2) / N
return total_cost
def stochastic_gradient_descent(independednt_variable, dependent_variable, learning_rate, I_bias, I_weights):
total_cost = []
N = float(independednt_variable.shape[0])
weights, bias = 0,0
change_in_weight, change_in_bias = 0,0
for j in range(5000):
for i in range(int(N)):
change_in_weight = (-2 / N) * independednt_variable[i] * (dependent_variable[i] - (weights * independednt_variable[i] + bias))
change_in_bias = (-2 / N) * (dependent_variable[i] - (weights * independednt_variable[i] + bias))
weights -= learning_rate * change_in_weight
bias -= learning_rate * change_in_bias
total_cost.append(compute_cost(weights, bias, independednt_variable, dependent_variable))
return total_cost, weights[0], bias[0]
errors, w, b = stochastic_gradient_descent(x, y, 0.01, 0, 0)
w, b
len(errors)
errors[-5:]
plt.figure(figsize = (15, 7))
plt.plot(range(len(errors)), errors)
plt.grid()
plt.xlabel("Epochs/Iterations")
plt.ylabel("Error")
plt.show()
plt.figure(figsize = (15, 7))
plt.scatter(x, y)
plt.plot(x, w*x + b, c = 'r')
plt.xlabel("x", size = 50)
plt.ylabel("y", size = 50)
plt.title("Custom Data for LR!", size = 50)
plt.show()
y_pred = w * x + b
from sklearn.metrics import r2_score, mean_squared_error
r2_score(y, y_pred), mean_squared_error(y, y_pred)
```
## Lab 6: Linear, LASSO & Ridge Regression on the House Prediction Dataset!
```
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
# Loading the dataset
train_data = pd.read_csv("train.csv")
test_data = pd.read_csv("test.csv")
train_data
test_data
data = pd.concat([train_data, test_data], keys = ["train", "test"])
data.drop(["Id", "SalePrice"], axis = 1, inplace = True)
data
data.columns
```
### Performing Sanity Checks
1. Check for Features that represent years should not go take values larger than 2018
2. Make sure, Areas, distances and prices should not take negative values
3. Months should be between 1 and 12
```
# Sanity Check 1
data[['YearBuilt', 'YearRemodAdd', 'GarageYrBlt','YrSold']].describe()
mask = (data[['YearBuilt', 'YearRemodAdd', 'GarageYrBlt','YrSold']] > 2018).any(axis=1) # take any index
data[mask][['YearBuilt', 'YearRemodAdd', 'GarageYrBlt','YrSold']]
# Repliacing the value of 'GarageYrBlt' by the value as 'YearBuilt' as the value of the former is > 2018
data.loc[mask, 'GarageYrBlt'] = data[mask]['YearBuilt']
data[mask][['YearBuilt', 'YearRemodAdd', 'GarageYrBlt','YrSold']]
# Sanity Check 2
# Make sure that the Areas, distances and prices should not take negative values
metrics = ['LotFrontage', 'LotArea', 'MasVnrArea','BsmtFinSF1', 'BsmtFinSF2','BsmtUnfSF','TotalBsmtSF',
'1stFlrSF', '2ndFlrSF', 'LowQualFinSF', 'GrLivArea', 'GarageArea', 'WoodDeckSF', 'OpenPorchSF',
'EnclosedPorch', '3SsnPorch', 'ScreenPorch', 'PoolArea', 'MiscVal']
mask2 = data[data[metrics] < 0].any(axis = 1) # Checking any row in the dataset for the condition
data[mask2]
# Sanity Check 3
mask3 = ((data["MoSold"] > 12) | (data["MoSold"] < 1))
data[mask3]
# No column with Name Grades, therefore, leave this processing step that is present in lab
# data["Grades"]
for i in data.columns:
if i.startswith("G"):
print(i)
# Fetching all the categorical Features
data.columns[data.dtypes == 'O']
```
### Dealing with Skewness
```
skewed_features = data[[col for col in data.columns if data[col].dtype != 'object']].skew(skipna=True)
skewed_features[abs(skewed_features) > 0.75]
data[skewed_features.index] = np.log1p(data[skewed_features.index])
data[[col for col in data.columns if data[col].dtype != 'object']].skew(skipna=True)
```
### Dealing wiith Missing Values
```
# Fetching all the variables with the Missing Values
data.isnull().sum()[data.isnull().sum() > 0]
# Fetching all the variables with the minimal Missing Values
data.isnull().sum()[(data.isnull().sum() > 0) & (data.isnull().sum() < 5)]
features_with_few_missing_values = data.isnull().sum()[(data.isnull().sum() > 0) & (data.isnull().sum() < 5)].index
# Checking out the results of the groupby
data.loc["train"].groupby("Neighborhood")[features_with_few_missing_values].first()
# Filling few variables with the minimum
features_filled = data.loc["train"].groupby("Neighborhood")[features_with_few_missing_values].apply(lambda x: x.mode().iloc[0])
features_filled
# Filling the missing values of the features having few missing values
for feature in features_with_few_missing_values:
data[feature] = data[feature].fillna(data["Neighborhood"].map(features_filled[feature]))
# Dealing with the missing values of LotFrontage Variable
plt.subplots(figsize=(15,5))
boxdata = data.loc['train'].groupby('LotConfig')['LotFrontage'].median().sort_values(ascending=False)
order = boxdata.index
sns.boxplot(x='LotConfig', y='LotFrontage', order=order,data=data.loc['train'])
plt.show()
# data['LotFrontage'] = data['LotFrontage'].fillna(data.loc['train','LotFrontage'].median())
# # bsmt=['BsmtQual', 'BsmtCond', 'BsmtExposure','BsmtFinSF1','BsmtFinType1', 'BsmtFinType2', 'BsmtFinSF2', 'BsmtUnfSF', 'BsmtFullBath', 'BsmtHalfBath', 'TotalBsmtSF']
# # fire = [' Fireplaces', 'FirePlaceQu']
# # garage = ['GarageQual', 'GarageCond', 'GarageType', 'GarageFinish','GarageArea', 'GarageCars','GarageYrBlt']
# # masn = ['MasVnrType','MasVnrArea']
# # others = ['Alley', 'Fence', 'PoolQC', 'MiscFeature']
categorical_features = data.columns[data.dtypes == 'object']
numerical_features = list(set(data.columns) - set(categorical_features))
categorical_features
numerical_features
data['MasVnrType'].replace({'None': np.nan}, inplace=True)
data.isnull().sum().sum()
# Filling all the missing values
data[categorical_features] = data[categorical_features].fillna('0')
data[numerical_features] = data[numerical_features].fillna(0)
data.isnull().sum().sum()
print(data.dtypes)
data['BsmtFullBath'] = data['BsmtFullBath'].astype('int64', copy=False)
data['BsmtHalfBath'] = data['BsmtHalfBath'].astype('int64', copy=False)
data['GarageCars'] = data['GarageCars'].astype('int64', copy=False)
data[['YearBuilt', 'YearRemodAdd', 'GarageYrBlt','YrSold']] = data[['YearBuilt', 'YearRemodAdd', 'GarageYrBlt','YrSold']].astype('int64', copy=False)
plt.hist(train_data.SalePrice)
plt.show()
# Removing the skewness of the price variable by applying the log transformation
price = np.log1p(train_data.SalePrice)
plt.hist(price)
plt.show()
# Concatenating column wise
categorical_data_with_labels = pd.concat((data.loc["train"][categorical_features], price), axis = 1)
categorical_data_with_labels
# Dropping the first column to get rid of the dummy variable trap
final_training_data = pd.concat((pd.get_dummies(categorical_data_with_labels.iloc[:, :-1], drop_first = True),
data.loc["train"][numerical_features],
categorical_data_with_labels.SalePrice,
), axis = 1)
final_training_data
final_testing_data = pd.concat((pd.get_dummies((data.loc["test"][categorical_features]), drop_first = True),
data.loc["test"][numerical_features],
categorical_data_with_labels.SalePrice,
), axis = 1)
final_testing_data
data.loc["test"][categorical_features].columns.size
data.loc["train"][categorical_features].columns.size
data.loc["test"][numerical_features].columns.size
data.loc["train"][numerical_features].columns.size
```
**Since, the number of columns in training & testing doesn't match, we can perform dimensionality reduction, but let's not go off topic from the subject, let's just fetch the columns from training data that are present in testing data.**
```
final_training_data = final_training_data[final_testing_data.columns]
final_training_data.shape, final_testing_data.shape
final_testing_data.dropna(inplace = True)
final_testing_data.shape
```
### Scaling the Data
```
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
training_data_scaled = scaler.fit_transform(final_training_data.iloc[:, :-1])
# testing_data_scaled = scaler.transform(final_testing_data)
```
### Splitting the Model
```
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(training_data_scaled, final_training_data.iloc[:, -1], test_size = 0.2)
# Not using this, as the explicit testing data doesn't had all the transformations applied.
# x_train, x_test, y_train, y_test = training_data_scaled[:, :-1], testing_data_scaled[:, :-1], training_data_scaled[:, -1], testing_data_scaled[:, -1]
x_train.shape, x_test.shape, y_train.shape, y_test.shape
```
### Applying Linear, LASSO & Ridge Regression
```
from sklearn.linear_model import LinearRegression, Lasso, Ridge, LassoCV, RidgeCV
from sklearn.metrics import r2_score
lin_reg = LinearRegression()
lin_reg.fit(x_train, y_train)
r2_score(lin_reg.predict(x_test), y_test)
# Lasso Regression with the default Penalty that is 1
las_reg = Lasso()
las_reg.fit(x_train, y_train)
r2_score(las_reg.predict(x_test), y_test)
# Lasso Regression having penalty as 0.01
las_reg = Lasso(alpha = 0.01)
las_reg.fit(x_train, y_train)
r2_score(las_reg.predict(x_test), y_test)
# Running Lasso with Cross Validation
las_reg = LassoCV()
las_reg.fit(x_train, y_train)
r2_score(las_reg.predict(x_test), y_test)
# Printing all the tried alphas
las_reg.alphas_
# Getting the best Alpha
las_reg.alpha_
# Number of features eliminated by Lasso Regression
las_reg.coef_[las_reg.coef_ == 0].shape
ridge_reg = Ridge()
ridge_reg.fit(x_train, y_train)
r2_score(ridge_reg.predict(x_test), y_test)
ridge_reg = RidgeCV()
ridge_reg.fit(x_train, y_train)
r2_score(ridge_reg.predict(x_test), y_test)
# Getting the best Alpha
ridge_reg.alpha_
```
### Getting the mean of all the Models
```
from sklearn.model_selection import cross_val_score
models = [LinearRegression(), LassoCV(), Ridge()]
CV = 5
results = np.empty((len(models), CV))
for i in range(len(models)):
results[i, :] = cross_val_score(models[i], x_train, y_train, cv = CV)
# Computing the Mean Accuracy of the Model (Row Wise Mean Calculation)
for i in results.mean(axis = 1):
print(round(i, 5))
```
# Lab 7: Decision Tree
```
import numpy as np
from sklearn.datasets import load_iris
X, y = load_iris(return_X_y = True)
classes_names = load_iris().target_names
X.shape, y.shape
X[-5:, :], y[-5:]
np.unique(y)
classes_names
```
| github_jupyter |
```
import numpy as np
import pylab as plt
# from tensorflow.keras.datasets.mnist import load_data
# # load the images into memory
# (trainX, trainy), (testX, testy) = load_data()
# # summarize the shape of the dataset
# print('Train', trainX.shape, trainy.shape)
# print('Test', testX.shape, testy.shape)
def generate_samples(n_sample,nf = 500,noise=0.03,real=True):
ys = []
for _ in range(n_sample):
x = np.linspace(0,10,nf)
a = np.random.uniform(0.9,1.1)
b = np.random.uniform(-2,2)
y = a*np.sin(x+b)
if real:
a = np.random.uniform(0.05,0.1)
b = np.random.uniform(-2,2)
y = y+a*np.sin(20*x+b)
y = y+np.random.normal(0,noise,nf)
ys.append(y)
ys = np.array(ys).reshape(n_sample,nf,1)
if real:
label = np.ones((n_sample, 1))
else:
label = np.zeros((n_sample, 1))
ys = ys-ys.min()
ys = ys/ys.max()
return ys,label
y,_ = generate_samples(n_sample=1,real=True)
plt.plot(y[0,:,0],'b')
y,_ = generate_samples(n_sample=1,real=False)
plt.plot(y[0,:,0],'r')
X_real, y_real = generate_samples(n_sample=20//2,real=True)
X_fake, y_fake = generate_samples(n_sample=20//2,real=False)
# example of training the discriminator model on real and random mnist images
from numpy import expand_dims
from numpy import ones
from numpy import zeros
from numpy.random import rand
from numpy.random import randint
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense
from tensorflow.keras.layers import Conv1D
from tensorflow.keras.layers import Flatten
from tensorflow.keras.layers import Dropout
from tensorflow.keras.layers import LeakyReLU
# define the standalone discriminator model
def define_discriminator(in_shape=(500,1)):
model = Sequential()
model.add(Conv1D(64, 7, strides=2, padding='same', input_shape=in_shape))
model.add(LeakyReLU(alpha=0.2))
model.add(Conv1D(16, 7, strides=2, padding='same'))
model.add(LeakyReLU(alpha=0.2))
model.add(Conv1D(4, 7, strides=2, padding='same'))
model.add(LeakyReLU(alpha=0.2))
model.add(Flatten())
model.add(Dropout(0.4))
model.add(Dense(1, activation='sigmoid'))
# compile model
opt = Adam(lr=0.0002, beta_1=0.5)
model.compile(loss='binary_crossentropy', optimizer=opt, metrics=['accuracy'])
return model
# train the discriminator model
def train_discriminator(model, n_iter=350, n_batch=501):
half_batch = int(n_batch / 2)
# manually enumerate epochs
for i in range(n_iter):
X_real, y_real = generate_samples(n_sample=half_batch,real=True)
X_fake, y_fake = generate_samples(n_sample=half_batch,real=False)
X = np.concatenate([X_real,X_fake],axis=0)
y = np.concatenate([y_real,y_fake],axis=0)
_, acc = model.train_on_batch(X, y)
if i%50==0:
print('{} acc={:2.2f}'.format(i+1, acc*100))
# define the discriminator model
model = define_discriminator()
model.summary()
# fit the model
train_discriminator(model)
from numpy import expand_dims
from numpy import zeros
from numpy import ones
from numpy import vstack
from numpy.random import randn
from numpy.random import randint
import tensorflow as tf
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense
from tensorflow.keras.layers import Reshape
from tensorflow.keras.layers import Flatten
from tensorflow.keras.layers import Conv2D
from tensorflow.keras.layers import Conv2DTranspose
from tensorflow.keras.layers import LeakyReLU
from tensorflow.keras.layers import Dropout
from matplotlib import pyplot
import logging
tf.get_logger().setLevel(logging.ERROR)
# define the standalone discriminator model
def define_discriminator(in_shape=(500,1)):
model = Sequential()
model.add(Conv1D(64, 7, strides=2, padding='same', input_shape=in_shape))
model.add(LeakyReLU(alpha=0.2))
model.add(Conv1D(16, 7, strides=2, padding='same'))
model.add(LeakyReLU(alpha=0.2))
model.add(Conv1D(4, 7, strides=2, padding='same'))
model.add(LeakyReLU(alpha=0.2))
model.add(Flatten())
model.add(Dropout(0.4))
model.add(Dense(1, activation='sigmoid'))
# compile model
opt = Adam(lr=0.0002, beta_1=0.5)
model.compile(loss='binary_crossentropy', optimizer=opt, metrics=['accuracy'])
return model
# define the standalone generator model
def define_convertor(in_shape=(500,1)):
model = Sequential()
model.add(Conv1D(12, 7, padding='same', input_shape=in_shape))
model.add(LeakyReLU(alpha=0.2))
model.add(Conv1D(12, 7, padding='same'))
model.add(LeakyReLU(alpha=0.2))
model.add(Conv1D(1, 7, padding='same'))
model.add(LeakyReLU(alpha=0.2))
return model
# define the combined generator and discriminator model, for updating the generator
def define_gan(c_model, d_model):
# make weights in the discriminator not trainable
d_model.trainable = False
# connect them
model = Sequential()
# add generator
model.add(c_model)
# add the discriminator
model.add(d_model)
# compile model
opt = Adam(lr=0.0002, beta_1=0.5)
model.compile(loss='binary_crossentropy', optimizer=opt, metrics=['accuracy'])
return model
# train the generator and discriminator
def train(d_model, gan_model, n_iter=2, n_batch=256):
half_batch = int(n_batch / 2)
# manually enumerate epochs
for i in range(n_iter):
X_real, y_real = generate_samples(n_sample=half_batch,real=True)
X_fake, y_fake = generate_samples(n_sample=half_batch,real=False)
X = np.concatenate([X_real,X_fake],axis=0)
y = np.concatenate([y_real,y_fake],axis=0)
_, acc = d_model.train_on_batch(X, y)
if i%10==0:
print('d: {} acc={:2.2f}'.format(i+1, acc*100))
X_fake, y_fake = generate_samples(n_sample=n_batch,real=False)
y = np.concatenate([y_real,y_real],axis=0)
_, acc = gan_model.train_on_batch(X_fake, y)
# summarize loss on this batch
if i%10==0:
print('g: {} acc={:2.2f}'.format(i+1, acc*100))
d_model = define_discriminator()
c_model = define_convertor()
gan_model = define_gan(c_model, d_model)
# n_batch=400
# half_batch = int(n_batch / 2)
# X_real, y_real = generate_samples(n_sample=half_batch,real=True)
# X_fake, y_fake = generate_samples(n_sample=n_batch,real=False)
# y = np.concatenate([y_real,y_real],axis=0)
# print(X_fake.shape, y.shape)
# _, acc = gan_model.train_on_batch(X_fake, y)
# gan_model.summary()
train(d_model, gan_model, n_iter=350, n_batch=500)
y,_ = generate_samples(n_sample=1,real=True)
plt.plot(y[0,:,0],'b')
y,_ = generate_samples(n_sample=1,real=False)
plt.plot(y[0,:,0],'r')
y = c_model.predict(y)
plt.plot(y[0,:,0],'b--')
plt.ylim(0,1)
```
| github_jupyter |
<h1> Time series prediction using RNNs, with TensorFlow and Cloud ML Engine </h1>
This notebook illustrates:
<ol>
<li> Creating a Recurrent Neural Network in TensorFlow
<li> Creating a Custom Estimator in tf.contrib.learn
<li> Training on Cloud ML Engine
</ol>
<p>
<h3> Simulate some time-series data </h3>
Essentially a set of sinusoids with random amplitudes and frequencies.
```
!pip install --upgrade tensorflow
import tensorflow as tf
print tf.__version__
import numpy as np
import tensorflow as tf
import seaborn as sns
import pandas as pd
SEQ_LEN = 10
def create_time_series():
freq = (np.random.random()*0.5) + 0.1 # 0.1 to 0.6
ampl = np.random.random() + 0.5 # 0.5 to 1.5
x = np.sin(np.arange(0,SEQ_LEN) * freq) * ampl
return x
for i in xrange(0, 5):
sns.tsplot( create_time_series() ); # 5 series
def to_csv(filename, N):
with open(filename, 'w') as ofp:
for lineno in xrange(0, N):
seq = create_time_series()
line = ",".join(map(str, seq))
ofp.write(line + '\n')
to_csv('train.csv', 1000) # 1000 sequences
to_csv('valid.csv', 50)
!head -5 train.csv valid.csv
```
<h2> RNN </h2>
For more info, see:
<ol>
<li> http://colah.github.io/posts/2015-08-Understanding-LSTMs/ for the theory
<li> https://www.tensorflow.org/tutorials/recurrent for explanations
<li> https://github.com/tensorflow/models/tree/master/tutorials/rnn/ptb for sample code
</ol>
Here, we are trying to predict from 8 values of a timeseries, the next two values.
<p>
<h3> Imports </h3>
Several tensorflow packages and shutil
```
import tensorflow as tf
import shutil
import tensorflow.contrib.learn as tflearn
import tensorflow.contrib.layers as tflayers
from tensorflow.contrib.learn.python.learn import learn_runner
import tensorflow.contrib.metrics as metrics
import tensorflow.contrib.rnn as rnn
```
<h3> Input Fn to read CSV </h3>
Our CSV file structure is quite simple -- a bunch of floating point numbers (note the type of DEFAULTS). We ask for the data to be read BATCH_SIZE sequences at a time. The Estimator API in tf.contrib.learn wants the features returned as a dict. We'll just call this timeseries column 'rawdata'.
<p>
Our CSV file sequences consist of 10 numbers. We'll assume that 8 of them are inputs and we need to predict the next two.
```
DEFAULTS = [[0.0] for x in xrange(0, SEQ_LEN)]
BATCH_SIZE = 20
TIMESERIES_COL = 'rawdata'
N_OUTPUTS = 2 # in each sequence, 1-8 are features, and 9-10 is label
N_INPUTS = SEQ_LEN - N_OUTPUTS
```
Reading data using the Estimator API in tf.learn requires an input_fn. This input_fn needs to return a dict of features and the corresponding labels.
<p>
So, we read the CSV file. The Tensor format here will be batchsize x 1 -- entire line. We then decode the CSV. At this point, all_data will contain a list of Tensors. Each tensor has a shape batchsize x 1. There will be 10 of these tensors, since SEQ_LEN is 10.
<p>
We split these 10 into 8 and 2 (N_OUTPUTS is 2). Put the 8 into a dict, call it features. The other 2 are the ground truth, so labels.
```
# read data and convert to needed format
def read_dataset(filename, mode=tf.contrib.learn.ModeKeys.TRAIN):
def _input_fn():
num_epochs = 100 if mode == tf.contrib.learn.ModeKeys.TRAIN else 1
# could be a path to one file or a file pattern.
input_file_names = tf.train.match_filenames_once(filename)
filename_queue = tf.train.string_input_producer(
input_file_names, num_epochs=num_epochs, shuffle=True)
reader = tf.TextLineReader()
_, value = reader.read_up_to(filename_queue, num_records=BATCH_SIZE)
value_column = tf.expand_dims(value, -1)
print 'readcsv={}'.format(value_column)
# all_data is a list of tensors
all_data = tf.decode_csv(value_column, record_defaults=DEFAULTS)
inputs = all_data[:len(all_data)-N_OUTPUTS] # first few values
label = all_data[len(all_data)-N_OUTPUTS : ] # last few values
# from list of tensors to tensor with one more dimension
inputs = tf.concat(inputs, axis=1)
label = tf.concat(label, axis=1)
print 'inputs={}'.format(inputs)
return {TIMESERIES_COL: inputs}, label # dict of features, label
return _input_fn
```
<h3> Define RNN </h3>
A recursive neural network consists of possibly stacked LSTM cells.
<p>
The RNN has one output per input, so it will have 8 output cells. We use only the last output cell, but rather use it directly, we do a matrix multiplication of that cell by a set of weights to get the actual predictions. This allows for a degree of scaling between inputs and predictions if necessary (we don't really need it in this problem).
<p>
Finally, to supply a model function to the Estimator API, you need to return a ModelFnOps. The rest of the function creates the necessary objects.
```
LSTM_SIZE = 3 # number of hidden layers in each of the LSTM cells
# create the inference model
def simple_rnn(features, targets, mode):
# 0. Reformat input shape to become a sequence
x = tf.split(features[TIMESERIES_COL], N_INPUTS, 1)
#print 'x={}'.format(x)
# 1. configure the RNN
lstm_cell = rnn.BasicLSTMCell(LSTM_SIZE, forget_bias=1.0)
outputs, _ = rnn.static_rnn(lstm_cell, x, dtype=tf.float32)
# slice to keep only the last cell of the RNN
outputs = outputs[-1]
#print 'last outputs={}'.format(outputs)
# output is result of linear activation of last layer of RNN
weight = tf.Variable(tf.random_normal([LSTM_SIZE, N_OUTPUTS]))
bias = tf.Variable(tf.random_normal([N_OUTPUTS]))
predictions = tf.matmul(outputs, weight) + bias
# 2. loss function, training/eval ops
if mode == tf.contrib.learn.ModeKeys.TRAIN or mode == tf.contrib.learn.ModeKeys.EVAL:
loss = tf.losses.mean_squared_error(targets, predictions)
train_op = tf.contrib.layers.optimize_loss(
loss=loss,
global_step=tf.contrib.framework.get_global_step(),
learning_rate=0.01,
optimizer="SGD")
eval_metric_ops = {
"rmse": tf.metrics.root_mean_squared_error(targets, predictions)
}
else:
loss = None
train_op = None
eval_metric_ops = None
# 3. Create predictions
predictions_dict = {"predicted": predictions}
# 4. return ModelFnOps
return tflearn.ModelFnOps(
mode=mode,
predictions=predictions_dict,
loss=loss,
train_op=train_op,
eval_metric_ops=eval_metric_ops)
```
<h3> Experiment </h3>
Distributed training is launched off using an Experiment. The key line here is that we use tflearn.Estimator rather than, say tflearn.DNNRegressor. This allows us to provide a model_fn, which will be our RNN defined above. Note also that we specify a serving_input_fn -- this is how we parse the input data provided to us at prediction time.
```
def get_train():
return read_dataset('train.csv', mode=tf.contrib.learn.ModeKeys.TRAIN)
def get_valid():
return read_dataset('valid.csv', mode=tf.contrib.learn.ModeKeys.EVAL)
def serving_input_fn():
feature_placeholders = {
TIMESERIES_COL: tf.placeholder(tf.float32, [None, N_INPUTS])
}
features = {
key: tf.expand_dims(tensor, -1)
for key, tensor in feature_placeholders.items()
}
features[TIMESERIES_COL] = tf.squeeze(features[TIMESERIES_COL], axis=[2])
print 'serving: features={}'.format(features[TIMESERIES_COL])
return tflearn.utils.input_fn_utils.InputFnOps(
features,
None,
feature_placeholders
)
from tensorflow.contrib.learn.python.learn.utils import saved_model_export_utils
def experiment_fn(output_dir):
# run experiment
return tflearn.Experiment(
tflearn.Estimator(model_fn=simple_rnn, model_dir=output_dir),
train_input_fn=get_train(),
eval_input_fn=get_valid(),
eval_metrics={
'rmse': tflearn.MetricSpec(
metric_fn=metrics.streaming_root_mean_squared_error
)
},
export_strategies=[saved_model_export_utils.make_export_strategy(
serving_input_fn,
default_output_alternative_key=None,
exports_to_keep=1
)]
)
shutil.rmtree('outputdir', ignore_errors=True) # start fresh each time
learn_runner.run(experiment_fn, 'outputdir')
```
<h3> Standalone Python module </h3>
To train this on Cloud ML Engine, we take the code in this notebook, make an standalone Python module.
```
%bash
# run module as-is
REPO=$(pwd)
echo $REPO
rm -rf outputdir
export PYTHONPATH=${PYTHONPATH}:${REPO}/simplernn
python -m trainer.task \
--train_data_paths="${REPO}/train.csv*" \
--eval_data_paths="${REPO}/valid.csv*" \
--output_dir=${REPO}/outputdir \
--job-dir=./tmp
```
Try out online prediction. This is how the REST API will work after you train on Cloud ML Engine
```
%writefile test.json
{"rawdata": [0.0,0.0527,0.10498,0.1561,0.2056,0.253,0.2978,0.3395]}
%bash
MODEL_DIR=$(ls ./outputdir/export/Servo/)
gcloud ml-engine local predict --model-dir=./outputdir/export/Servo/$MODEL_DIR --json-instances=test.json
```
<h3> Cloud ML Engine </h3>
Now to train on Cloud ML Engine.
```
%bash
# run module on Cloud ML Engine
REPO=$(pwd)
BUCKET=cloud-training-demos-ml # CHANGE AS NEEDED
OUTDIR=gs://${BUCKET}/simplernn/model_trained
JOBNAME=simplernn_$(date -u +%y%m%d_%H%M%S)
REGION=us-central1
gsutil -m rm -rf $OUTDIR
gcloud ml-engine jobs submit training $JOBNAME \
--region=$REGION \
--module-name=trainer.task \
--package-path=${REPO}/simplernn/trainer \
--job-dir=$OUTDIR \
--staging-bucket=gs://$BUCKET \
--scale-tier=BASIC \
--runtime-version=1.2 \
-- \
--train_data_paths="gs://${BUCKET}/train.csv*" \
--eval_data_paths="gs://${BUCKET}/valid.csv*" \
--output_dir=$OUTDIR \
--num_epochs=100
```
<h2> Variant: long sequence </h2>
To create short sequences from a very long sequence.
```
import tensorflow as tf
import numpy as np
def breakup(sess, x, lookback_len):
N = sess.run(tf.size(x))
windows = [tf.slice(x, [b], [lookback_len]) for b in xrange(0, N-lookback_len)]
windows = tf.stack(windows)
return windows
x = tf.constant(np.arange(1,11, dtype=np.float32))
with tf.Session() as sess:
print 'input=', x.eval()
seqx = breakup(sess, x, 5)
print 'output=', seqx.eval()
```
Copyright 2017 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License
| github_jupyter |
# Matrix Factorization with Expander Graphs
## Supporting Functions
### Functions for generating and checking data
```
# IMPORTS
import numpy as np
import copy
import matplotlib.pyplot as plt
from timeit import default_timer as timer
from numpy import linalg as LA
# DATA GENERATION AND PERFORMANCE CHECKING FUNCTIONS
def generate_latent(n,k,c1,c2):
b = c2-c1
a = c1/b
x = np.zeros(n)
locs = np.random.permutation(n)[:k]
x[locs] = 1
signs = np.sign(np.random.randn(k))
vals = b*(np.random.rand(k)+a)
x[locs] = x[locs]*vals*signs
return x
def generate_exp_dictionary1(m,n,d):
A = np.zeros((m,n))
for i in range(n):
locs = np.random.permutation(m)[:d]
A[locs,i] = 1
return A
def generate_exp_dictionary2(m,n,d):
A = np.zeros((m,n))
alpha = int(np.floor(m/d))
c = 0
while c<n:
perm = np.random.permutation(m)
zeta = min(n-c, alpha)
for j in range(zeta):
locs = perm[j*d:(j+1)*d]
A[locs,c] = 1
c = c+1
return A
def check_dictionary(eps,n,d,A):
t = (1-4*eps)*d
mu_A = np.max(np.transpose(A)@A - d*np.identity(n))
if mu_A >= t:
print("Generation error, A is not expander for this value of epsilon, mu_A = ", str.format('{0:.2f}', mu_A),", tau = ", str.format('{0:.2f}', t), ".")
passed = False
else:
print("A passes coherence test, mu_A = ", str.format('{0:.2f}', mu_A),", tau = ", str.format('{0:.2f}', t), ".")
passed = True
return passed
def check_recon_A(A,A_rec,d,eps,n):
print("Checking reconstruction of A.")
col_sum = sum(A_rec)
print(str.format('{0:.1f}',100*len(col_sum[col_sum==d])/n), "% of columns fully formed.")
print(str.format('{0:.1f}',100*(n - len(col_sum[col_sum==d]) - len(col_sum[col_sum==0]))/n) , "% of columns partially formed.")
print(str.format('{0:.1f}', 100*len(col_sum[col_sum==0])/n), "% of columns have no entries.")
overlaps = np.transpose(A_rec)@A
A_rec_col, A_col = np.where(overlaps>(1-4*eps)*d)
if len(col_sum[col_sum==0])<n:
print(str.format('{0:.1f}', 100*len(A_rec_col)/(n - len(col_sum[col_sum==0]))), "% of reconstructed (fully or partially) columns match with a column in target matrix.")
error = 0
missing = 0
P = np.zeros((n,n))
for i in range(len(A_rec_col)):
diff = A_rec[:, A_rec_col[i]] - A[:,A_col[i]]
error = error + sum(abs(diff[diff==1]))
missing = missing + sum(abs(diff[diff==-1]))
P[A_col[i], A_rec_col[i]] = 1
print(str.format('{0:.0f}', 100*(missing+d*n-len(A_rec_col)*d)/(d*n)), "% of entries missing in total.")
print(str.format('{0:.0f}', missing), "entries missing in matched columns.")
print(str.format('{0:.0f}', error), "entries wrong in matched columns.")
return P
def check_recon_X(X,X_rec,P,N,k,n):
X_perm = P@X_rec
print("Checking the values of X recovered.")
tol = 10**-6
total_entries = N*k
total_entries_recovered = 0
correct_loc_correct_val = 0
correct_loc_wrong_val = 0
wrong_loc = 0
missing = 0
passed = True
for i in range(N):
for j in range(n):
if abs(X[j,i])>tol and abs(X_perm[j,i])>tol:
error = abs(X[j,i] - X_perm[j,i])
if error < tol:
correct_loc_correct_val +=1
else:
correct_loc_wrong_val += 1
print("Column number = ", i, ", row number = ", j, ", true value = ", X[j,i], "value found = ", X_perm[j,i])
elif abs(X[j,i])>tol and abs(X_perm[j,i])<tol:
missing += 1
elif abs(X[j,i])<tol and abs(X_perm[j,i])>tol:
wrong_loc += 1
print(str.format('{0:.1f}', 100*(correct_loc_correct_val/total_entries)), "% of entries correctly recovered.")
print(str.format('{0:.1f}', 100*(correct_loc_wrong_val/total_entries)), "% of entries in correct location but have wrong value for location.")
print(str.format('{0:.1f}', 100*(missing/total_entries)), "% of entries missing.")
return X_perm
```
### Auxillary functions
```
# USED IN BOTH EXPERIMENTS 1 and 2
def xps2(y,x_vals,x_locs,A_rec,m,eps,d,k):
tol1 = 10**-8
tol2 = 10**-8
y_copy = copy.deepcopy(y)
t = (1-2*eps)*d
z1 = len(y_copy);
W = np.zeros((m,k))
vals = np.zeros(k)
q = 0
for i in range(z1):
if np.abs(y_copy[i])> tol1:
locs = list()
locs.append(i)
for j in range(i+1, m):
if np.abs(y_copy[j] - y_copy[i]) < tol2:
locs.append(j)
y_copy[j] = 0
match = np.where(y_copy[i]==x_vals)[0]
if not match:
if len(locs)>t:
W[locs,q] = 1
vals[q] = y_copy[i]
q = q+1
elif len(match)==1:
A_rec[locs, x_locs[match[0]]] = 1
else:
print('ERROR - column value matches with more than 1 singleton value already extracted')
y_copy[i] = 0
W = W[:,:q]
vals = vals[:q]
return [W, A_rec, vals, q]
def process_column2(y, A_rec, x, m, eps, d, k, c, dpid):
exit_flag = True
tau = (1-4*eps)*d
x_locs = np.nonzero(x)[0]
x_vals = np.zeros(len(x_locs))
for i in range(len(x_locs)):
x_vals[i] = x[x_locs[i]]
[W, A_rec, vals, q] = xps2(y,x_vals,x_locs,A_rec,m,eps,d,k)
if q > 0:
match_check = np.zeros(q)
match_col_id = -0.5*np.ones(q)
data_id = dpid*np.ones(q)
if c > 0:
matches = np.transpose(W)@A_rec
matches[matches<=tau] = 0
matches[matches>tau] = 1
num_matches = matches.sum(axis=1)
col_ind = matches@np.arange(c)
for i in range(q):
if num_matches[i] == 0:
if c>=n:
print("Error: trying to add new column which exceeds total number of columns")
exit_flag==False
elif num_matches[i] == 1:
match_check[i] = 1
z = int(col_ind[i])
match_col_id[i] = z
elif num_matches[i] > 1:
print("Error: partial support matches with ", num_matches[i]," partially reconstructed columns of A.")
exit_flag = False
else:
exit_flag = False
match_check = []
match_col_id = []
data_id = []
return [W, A_rec, match_check, match_col_id, data_id, vals, exit_flag]
def sort_unmatched_columns(W, eps, d, c):
q = W.shape[1]
col_id_new = -0.5*np.zeros(q)
tau = (1-4*eps)*d
matches = np.transpose(W)@W
matches[matches<=tau] = 0
matches[matches>tau] = 1
i =0
while i<q:
if sum(matches[:,i]) > 0:
temp = np.where(matches[:,i]==1)[0]
col_id_new[temp] = c
c +=1
for j in range(len(temp)):
matches[temp[j],:] = np.zeros(q)
matches[:,temp[j]] = np.zeros(q)
i +=1
return [col_id_new, c]
def update_reconstruction(W, col_id, data_id, vals, A_rec, X_rec):
for i in range(len(col_id)):
A_rec[:, int(col_id[i])] += W[:,i]
X_rec[int(col_id[i]), int(data_id[i])] = vals[i]
A_rec[A_rec>0] = 1
return [A_rec, X_rec]
def l0_decode(y, A, num_comp, alpha):
tol = 10**-3
tol2 = 10**-9
converged = False
r = copy.deepcopy(y)
r_old = copy.deepcopy(y)
x = np.zeros(num_comp)
iters=1
while converged == False and iters<10:
vals = np.unique(r)
vals = vals[vals!=0]
for j in range(num_comp):
r_0 = np.count_nonzero(r)
for v in np.nditer(vals):
u = r - v*A[:,j]
u_0 = np.count_nonzero(u)
if r_0 - u_0 >= alpha:
x[j] = x[j] + v
r = y - A@x
r[np.absolute(r)<tol2]=0
if LA.norm(r - r_old) <= LA.norm(r_old)*tol or LA.norm(r)<=LA.norm(y)*tol:
converged = True
else:
r_old = copy.deepcopy(r)
iters +=1
if iters == 10:
print("Warning: max number of iterations of while loop in l0 decode step reached.")
print('Percentage improvement in residual from penultimate iteration: ', str.format('{0:.5f}', LA.norm(r - r_old)))
print('Size of final residual: ' , str.format('{0:.5f}', LA.norm(r)))
return x
```
### EBR Algorithm
```
def EBR(Y,N,m,n,d,k,eps,max_epochs,A_sparsity,X_sparsity,Y_frob):
epoch = 0
c = 0
tol = 10**-8 # make sure is smaller than 1
exit_program=False
A_rec = np.zeros((m,n))
X_rec = np.zeros((n,N))
frob_error_series = np.ones(max_epoch)
A_entries = np.zeros(max_epoch)
X_entries = np.zeros(max_epoch)
total_entries = np.zeros(max_epoch)
R = copy.deepcopy(Y)
start_time = timer()
while epoch < max_epoch and frob_error_series[epoch] > tol and exit_program==False:
num_matched = 0
num_new = 0
for i in range(N):
[W, A_rec[:,:c], match_check, match_col_id, data_id, vals, supports_found]=process_column2(R[:,i], A_rec[:,:c], X_rec[:,i], m, eps, d, k, c, i)
if supports_found == True:
matches = np.where(match_check==1)[0]
new = np.where(match_check==0)[0]
if len(matches)>0:
if num_matched > 0:
W_match = np.concatenate((W_match, W[:,matches]), axis=1)
col_id_match = np.concatenate((col_id_match, match_col_id[matches]), axis=0)
data_id_match = np.concatenate((data_id_match, data_id[matches]), axis=0)
val_match = np.concatenate((val_match, vals[matches]), axis=0)
else:
W_match = W[:,matches]
col_id_match = match_col_id[matches]
data_id_match = data_id[matches]
val_match = vals[matches]
num_matched += W_match.shape[1]
if len(new)>0:
if num_new > 0:
W_new = np.concatenate((W_new, W[:,new]), axis=1)
data_id_new = np.concatenate((data_id_new, data_id[new]), axis=0)
val_new = np.concatenate((val_new, vals[new]), axis=0)
else:
W_new = W[:,new]
data_id_new = data_id[new]
val_new = vals[new]
num_new += W_new.shape[1]
if num_matched > 0 and num_new > 0:
[col_id_new, c] = sort_unmatched_columns(W_new, eps, d, c)
W_all = np.concatenate((W_match, W_new), axis=1)
col_id_all = np.concatenate((col_id_match, col_id_new), axis=0)
data_id_all = np.concatenate((data_id_match, data_id_new), axis=0)
val_all = np.concatenate((val_match, val_new), axis=0)
elif num_matched > 0 and num_new == 0:
W_all = W_match
col_id_all = col_id_match
data_id_all = data_id_match
val_all = val_match
elif num_matched == 0 and num_new > 0:
[col_id_all, c] = sort_unmatched_columns(W_new, eps, d, c)
W_all = W_new
data_id_all = data_id_new
val_all = val_new
elif num_matched == 0 and num_new == 0:
print("No partial supports recovered, terminating algorithm")
frob_error_series[epoch:] = frob_error_series[epoch-1]
A_entries[epoch:] = A_entries[epoch-1]
X_entries[epoch:] = X_entries[epoch-1]
total_entries[epoch:] = total_entries[epoch-1]
exit_program = True
if exit_program == False:
[A_rec, X_rec] = update_reconstruction(W_all, col_id_all, data_id_all, val_all, A_rec, X_rec)
R = Y - A_rec@X_rec
R[np.absolute(R)<tol] = 0
frob_error_series[epoch] = np.sqrt(sum(sum(R**2)))
A_entries[epoch] = np.count_nonzero(A_rec)
X_entries[epoch] = np.count_nonzero(X_rec)
total_entries[epoch] = A_entries[epoch] + X_entries[epoch]
print('Epoch ', str.format('{0:.0f}', epoch), " processed, l_2 error = ", str.format('{0:.2f}', 100*frob_error_series[epoch]/Y_frob), '%, A l_0 error =', str.format('{0:.2f}', 100*(A_sparsity - A_entries[epoch])/A_sparsity), '%, X l_0 error =', str.format('{0:.2f}', 100*(X_sparsity - X_entries[epoch])/X_sparsity), '%.')
if epoch == max_epoch:
print("Maximum number of epochs reached.")
epoch +=1
end_time = timer()
comp_time = end_time - start_time
return [A_rec, X_rec, frob_error_series, total_entries, A_entries, X_entries, comp_time]
def EBR_l0(Y,N,m,n,d,k,eps,max_epochs,A_sparsity,X_sparsity,Y_frob, alpha):
epoch = 0
c = 0
tol = 10**-8 # make sure is smaller than 1
exit_program=False
A_rec = np.zeros((m,n))
X_rec = np.zeros((n,N))
frob_error_series = np.ones(max_epoch)
A_entries = np.zeros(max_epoch)
X_entries = np.zeros(max_epoch)
total_entries = np.zeros(max_epoch)
R = copy.deepcopy(Y)
start_time = timer()
while epoch < max_epoch and frob_error_series[epoch] > tol and exit_program==False:
num_matched = 0
num_new = 0
for i in range(N):
[W, A_rec[:,:c], match_check, match_col_id, data_id, vals, supports_found]=process_column2(R[:,i], A_rec[:,:c], X_rec[:,i], m, eps, d, k, c, i)
if supports_found == True:
matches = np.where(match_check==1)[0]
new = np.where(match_check==0)[0]
if len(matches)>0:
if num_matched > 0:
W_match = np.concatenate((W_match, W[:,matches]), axis=1)
col_id_match = np.concatenate((col_id_match, match_col_id[matches]), axis=0)
data_id_match = np.concatenate((data_id_match, data_id[matches]), axis=0)
val_match = np.concatenate((val_match, vals[matches]), axis=0)
else:
W_match = W[:,matches]
col_id_match = match_col_id[matches]
data_id_match = data_id[matches]
val_match = vals[matches]
num_matched += W_match.shape[1]
if len(new)>0:
if num_new > 0:
W_new = np.concatenate((W_new, W[:,new]), axis=1)
data_id_new = np.concatenate((data_id_new, data_id[new]), axis=0)
val_new = np.concatenate((val_new, vals[new]), axis=0)
else:
W_new = W[:,new]
data_id_new = data_id[new]
val_new = vals[new]
num_new += W_new.shape[1]
if num_matched > 0 and num_new > 0:
[col_id_new, c] = sort_unmatched_columns(W_new, eps, d, c)
W_all = np.concatenate((W_match, W_new), axis=1)
col_id_all = np.concatenate((col_id_match, col_id_new), axis=0)
data_id_all = np.concatenate((data_id_match, data_id_new), axis=0)
val_all = np.concatenate((val_match, val_new), axis=0)
elif num_matched > 0 and num_new == 0:
W_all = W_match
col_id_all = col_id_match
data_id_all = data_id_match
val_all = val_match
elif num_matched == 0 and num_new > 0:
[col_id_all, c] = sort_unmatched_columns(W_new, eps, d, c)
W_all = W_new
data_id_all = data_id_new
val_all = val_new
elif num_matched == 0 and num_new == 0:
print("No partial supports recovered, terminating algorithm")
frob_error_series[epoch:] = frob_error_series[epoch-1]
A_entries[epoch:] = A_entries[epoch-1]
X_entries[epoch:] = X_entries[epoch-1]
total_entries[epoch:] = total_entries[epoch-1]
exit_program = True
if exit_program == False:
[A_rec, X_rec] = update_reconstruction(W_all, col_id_all, data_id_all, val_all, A_rec, X_rec)
### RUN l-0 HERE using completed columns ####
# Identify the completed columns of A at this iteration
col_sumA = np.sum(A_rec, axis = 0)
col_sumA[col_sumA != d] = 0
comp = np.nonzero(col_sumA)[0]
num_comp = len(comp)
# Compute the residual based on the completed columns only
R_l0 = Y - A_rec[:,comp]@X_rec[comp, :]
R_l0[np.absolute(R_l0)<tol] = 0
# Identify which columns of the residual need to still try and decode
col_sumR = np.sum(np.absolute(R_l0), axis = 0)
col_sumR[np.absolute(col_sumR) < tol] = 0
not_dec = np.nonzero(col_sumR)[0]
num_2_process = len(not_dec)
print("Running l0 decode on epoch ", str.format('{0:.0f}', epoch+1), ': n. of data points of Y still to decode = ', str.format('{0:.0f}', num_2_process), ", n. columns of A completed = ", str.format('{0:.0f}', num_comp),)
# Iterate through each column in turn and run l0 decode
for j in range(num_2_process):
X_l0_update = l0_decode(R_l0[:,not_dec[j]], A_rec[:,comp], num_comp, alpha)
X_rec[comp, not_dec[j]] = X_rec[comp, not_dec[j]] + X_l0_update
if np.mod(int(not_dec[j]),np.ceil(num_2_process/5)) == 0:
print("Processed ", str.format('{0:.0f}', j), " data points out of ", str.format('{0:.0f}', num_2_process))
############################################
R = Y - A_rec@X_rec
R[np.absolute(R)<tol] = 0
frob_error_series[epoch] = np.sqrt(sum(sum(R**2)))
A_entries[epoch] = np.count_nonzero(A_rec)
X_entries[epoch] = np.count_nonzero(X_rec)
total_entries[epoch] = A_entries[epoch] + X_entries[epoch]
print('Epoch ', str.format('{0:.0f}', epoch), " processed, l_2 error = ", str.format('{0:.2f}', 100*frob_error_series[epoch]/Y_frob), '%, A l_0 error =', str.format('{0:.2f}', 100*(A_sparsity - A_entries[epoch])/A_sparsity), '%, X l_0 error =', str.format('{0:.2f}', 100*(X_sparsity - X_entries[epoch])/X_sparsity), '%.')
if epoch == max_epoch:
print("Maximum number of epochs reached.")
epoch +=1
end_time = timer()
comp_time = end_time - start_time
return [A_rec, X_rec, frob_error_series, total_entries, A_entries, X_entries, comp_time]
```
# Experiments analyzing performance of EBR
## Generate synthetic data to run algorithms on
#### Generating a valid dictionary and data
```
# Dictionary parameters parameters
n = 1200
# beta = 900
m = 900
d = 10;
eps = 1/6;
N = 200
# Generate a dictionary and check that passes necessary test.
num_generations = 0
passed = False
while passed==False and num_generations < 100:
A = generate_exp_dictionary2(m,n,d);
passed = check_dictionary(eps,n,d,A)
num_generations += 1
if passed == True:
print("Generated dictionary that passes the coherency test")
else:
print("Failed to generate dictionary that passes the coherency test, consider different parameters")
A_sparsity = np.count_nonzero(A)
```
#### Generate data using dictionary
```
##### GENERATE LATENT REPRESENTATION
b1=1;
b2=5;
# k = [int(np.ceil(0.01*n)), int(np.ceil(0.04*n)), int(np.ceil(0.07*n)), int(np.ceil(0.1*n))]
k = [int(np.ceil(0.01*n)), int(np.ceil(0.03*n)), int(np.ceil(0.05*n)), int(np.ceil(0.07*n))]
X = np.zeros((len(k), n, N))
Y = np.zeros((len(k), m, N))
Y_frob = np.zeros(len(k))
X_sparsity = np.zeros(len(k))
number_k_to_process = len(k)
for j in range(len(k)):
for i in range(N):
X[j,:,i] = generate_latent(n,k[j],b1,b2)
Y[j] = A@X[j]
Y_frob[j] = np.sqrt(sum(sum(Y[j]**2)))
X_sparsity[j] = k[j]*N
```
## Check that l0 function is working as it should be
```
ind = 3
alpha=3
N_check = 1
Z = np.zeros((n, N_check))
offset = 28
for j in range(N_check):
Z[:,j] = l0_decode(Y[ind,:,offset+j], A, n, alpha)
if np.mod(j,np.ceil(N_check/10)) == 0:
print(j)
check_recon_X(X[ind,:,offset:offset+N_check],Z,np.eye(n),N_check,k[ind],n)
```
## Running EBR to recover A and X varying $k/n$
```
if passed == True:
eps = 1/6;
max_epoch = 8
tol = 10**-8
comp_time = np.zeros(len(k))
A_rec = np.zeros((len(k),m,n))
X_rec = np.zeros((len(k),n,N))
frob_error_series = np.ones((len(k), max_epoch))
A_entries = np.zeros((len(k), max_epoch))
X_entries = np.zeros((len(k), max_epoch))
total_entries = np.zeros((len(k), max_epoch))
for j in range(number_k_to_process):
print("Solving problem with k/n = ", str.format('{0:.1f}', 100*k[j]/n), '%, N = ', str.format('{0:.0f}', N))
[A_rec[j], X_rec[j], frob_error_series[j], total_entries[j], A_entries[j], X_entries[j], comp_time[j]] = EBR(Y[j],N,m,n,d,k[j],eps,max_epoch,A_sparsity,X_sparsity[j],Y_frob[j])
print("")
P = check_recon_A(A, A_rec[j],d,eps,n)
print("")
check_recon_X(X[j],X_rec[j],P,N,k[j],n)
print("")
plt.figure(1).clear()
plot_A_entries = 100*A_entries/A_sparsity
plot_X_entries = np.zeros((len(k), max_epoch))
plot_frob = np.zeros((len(k), max_epoch))
for j in range(number_k_to_process):
plot_X_entries[j,:] = 100*X_entries[j,:]/X_sparsity[j]
plot_epoch = max_epoch
print("Running times:")
print("k/n=%s %%, %s s"%(100*k[0]/n, comp_time[0]))
print("k/n=%s %%, %s s"%(100*k[1]/n, comp_time[1]))
print("k/n=%s %%, %s s"%(100*k[2]/n, comp_time[2]))
plt.figure(1)
plt.figure(figsize=(15,20))
plt.subplot(2,2,1)
plt.plot(np.arange(max_epoch)+1, plot_A_entries[0], 'g-', label="A, k/n=%s %%"%(int(round(100*k[0]/n, 0))))
plt.plot(np.arange(max_epoch)+1, plot_A_entries[1], 'b-', label="A, k/n=%s %%"%(int(round(100*k[1]/n, 0))))
plt.plot(np.arange(max_epoch)+1, plot_A_entries[2], 'r-', label="A, k/n=%s %%"%(int(round(100*k[2]/n, 0))))
plt.plot(np.arange(max_epoch)+1, plot_A_entries[3], 'k-', label="A, k/n=%s %%"%(int(round(100*k[3]/n, 0))))
plt.grid(True)
plt.title('Percentage of entries of A recovered by EBR varying k/n')
plt.xlabel('Epochs')
plt.ylabel('Entries of A recovered (%)')
plt.xlim(1,plot_epoch)
plt.legend()
plt.subplot(2,2,2)
plt.plot(np.arange(max_epoch)+1, plot_X_entries[0], 'g-',label="X, k/n=%s %%"%(int(round(100*k[0]/n, 0))))
plt.plot(np.arange(max_epoch)+1, plot_X_entries[1], 'b-',label="X, k/n=%s %%"%(int(round(100*k[1]/n, 0))))
plt.plot(np.arange(max_epoch)+1, plot_X_entries[2], 'r-',label="X, k/n=%s %%"%(int(round(100*k[2]/n, 0))))
plt.plot(np.arange(max_epoch)+1, plot_X_entries[3], 'k-',label="X, k/n=%s %%"%(int(round(100*k[3]/n, 0))))
plt.grid(True)
plt.title('Percentage of entries of A recovered by EBR varying k/n')
plt.xlabel('Epochs')
plt.ylabel('Entries of X recovered (%)')
plt.xlim(1,plot_epoch)
plt.legend()
# plt.savefig('./figures_ACHA/EBR_performance.eps', format='eps', bbox_inches='tight')
```
## Running EBR-l0 to recover A and X varying $k/n$
```
# Extra parameters required for EBR-l0
alpha=3
# Run EBR interweaved with l-0 decode
if passed == True:
eps = 1/6;
tol = 10**-8
comp_time_l0 = np.zeros(len(k))
A_rec_l0 = np.zeros((len(k),m,n))
X_rec_l0 = np.zeros((len(k),n,N))
frob_error_series_l0 = np.ones((len(k), max_epoch))
A_entries_l0 = np.zeros((len(k), max_epoch))
X_entries_l0 = np.zeros((len(k), max_epoch))
total_entries_l0 = np.zeros((len(k), max_epoch))
for j in range(number_k_to_process):
print("Solving problem with k/n = ", str.format('{0:.1f}', 100*k[j]/n), '%, N = ', str.format('{0:.0f}', N))
[A_rec_l0[j], X_rec_l0[j], frob_error_series_l0[j], total_entries_l0[j], A_entries_l0[j], X_entries_l0[j], comp_time_l0[j]] = EBR_l0(Y[j],N,m,n,d,k[j],eps,max_epoch,A_sparsity,X_sparsity[j],Y_frob[j],alpha)
print("")
P = check_recon_A(A, A_rec_l0[j],d,eps,n)
print("")
check_recon_X(X[j],X_rec_l0[j],P,N,k[j],n)
print("")
plt.figure(2).clear()
plot_X_entries_l0 = np.zeros((len(k), max_epoch))
plot_A_entries_l0 = 100*A_entries_l0/A_sparsity
for j in range(number_k_to_process):
plot_X_entries_l0[j,:] = 100*X_entries_l0[j,:]/X_sparsity[j]
plot_epoch = max_epoch
print("Running times:")
print("k/n=%s %%, %s s"%(100*k[0]/n, comp_time[0]))
print("k/n=%s %%, %s s"%(100*k[1]/n, comp_time[1]))
print("k/n=%s %%, %s s"%(100*k[2]/n, comp_time[2]))
plt.figure(2)
plt.figure(figsize=(15,20))
plt.subplot(2,2,1)
plt.plot(np.arange(max_epoch)+1, plot_A_entries_l0[0], 'g--', label="A, k/n=%s %%"%(int(round(100*k[0]/n, 0))))
plt.plot(np.arange(max_epoch)+1, plot_A_entries_l0[1], 'b--', label="A, k/n=%s %%"%(int(round(100*k[1]/n, 0))))
plt.plot(np.arange(max_epoch)+1, plot_A_entries_l0[2], 'r--', label="A, k/n=%s %%"%(int(round(100*k[2]/n, 0))))
plt.plot(np.arange(max_epoch)+1, plot_A_entries_l0[3], 'k--', label="A, k/n=%s %%"%(int(round(100*k[3]/n, 0))))
plt.grid(True)
plt.title('Percentage of entries of A recovered by EBR-l0 varying k/n')
plt.xlabel('Epochs')
plt.ylabel('Entries of A recovered (%)')
plt.xlim(1,plot_epoch)
plt.legend()
plt.subplot(2,2,2)
plt.plot(np.arange(max_epoch)+1, plot_X_entries_l0[0], 'g--',label="X, k/n=%s %%"%(int(round(100*k[0]/n, 0))))
plt.plot(np.arange(max_epoch)+1, plot_X_entries_l0[1], 'b--',label="X, k/n=%s %%"%(int(round(100*k[1]/n, 0))))
plt.plot(np.arange(max_epoch)+1, plot_X_entries_l0[2], 'r--',label="X, k/n=%s %%"%(int(round(100*k[2]/n, 0))))
plt.plot(np.arange(max_epoch)+1, plot_X_entries_l0[3], 'k--',label="X, k/n=%s %%"%(int(round(100*k[3]/n, 0))))
plt.grid(True)
plt.title('Percentage of entries of A recovered by EBR-l0 varying k/n')
plt.xlabel('Epochs')
plt.ylabel('Entries of X recovered (%)')
plt.xlim(1,plot_epoch)
plt.legend()
plt.savefig('./figures_ACHA/EBR_l0_performance.eps', format='eps', bbox_inches='tight')
```
## Side by side comparison of EBR vs EBR-l0
```
plot_epoch = 6
plt.figure(3)
plt.figure(figsize=(15,20))
plt.subplot(2,2,1)
plt.plot(np.arange(max_epoch)+1, plot_A_entries[0], 'g-', label="EBR, k/n=%s %%"%(int(round(100*k[0]/n, 0))))
plt.plot(np.arange(max_epoch)+1, plot_A_entries[1], 'b-', label="EBR, k/n=%s %%"%(int(round(100*k[1]/n, 0))))
plt.plot(np.arange(max_epoch)+1, plot_A_entries[2], 'r-', label="EBR, k/n=%s %%"%(int(round(100*k[2]/n, 0))))
plt.plot(np.arange(max_epoch)+1, plot_A_entries[3], 'k-', label="EBR, k/n=%s %%"%(int(round(100*k[3]/n, 0))))
plt.plot(np.arange(max_epoch)+1, plot_A_entries_l0[0], 'g--', label="EBR-l0, k/n=%s %%"%(int(round(100*k[0]/n, 0))))
plt.plot(np.arange(max_epoch)+1, plot_A_entries_l0[1], 'b--', label="EBR-l0, k/n=%s %%"%(int(round(100*k[1]/n, 0))))
plt.plot(np.arange(max_epoch)+1, plot_A_entries_l0[2], 'r--', label="EBR-l0, k/n=%s %%"%(int(round(100*k[2]/n, 0))))
plt.plot(np.arange(max_epoch)+1, plot_A_entries_l0[3], 'k--', label="EBR-l0, k/n=%s %%"%(int(round(100*k[3]/n, 0))))
plt.grid(True)
plt.title('Percentage of entries of A recovered by EBR and EBR-l0 varying k/n')
plt.xlabel('Epochs')
plt.ylabel('Entries of A recovered (%)')
plt.xlim(1,plot_epoch)
plt.legend()
plt.subplot(2,2,2)
plt.plot(np.arange(max_epoch)+1, plot_X_entries[0], 'g-',label="EBR, k/n=%s %%"%(int(round(100*k[0]/n, 0))))
plt.plot(np.arange(max_epoch)+1, plot_X_entries[1], 'b-',label="EBR, k/n=%s %%"%(int(round(100*k[1]/n, 0))))
plt.plot(np.arange(max_epoch)+1, plot_X_entries[2], 'r-',label="EBR, k/n=%s %%"%(int(round(100*k[2]/n, 0))))
plt.plot(np.arange(max_epoch)+1, plot_X_entries[3], 'k-',label="EBR, k/n=%s %%"%(int(round(100*k[3]/n, 0))))
plt.plot(np.arange(max_epoch)+1, plot_X_entries_l0[0], 'g--',label="EBR-l0, k/n=%s %%"%(int(round(100*k[0]/n, 0))))
plt.plot(np.arange(max_epoch)+1, plot_X_entries_l0[1], 'b--',label="EBR-l0, k/n=%s %%"%(int(round(100*k[1]/n, 0))))
plt.plot(np.arange(max_epoch)+1, plot_X_entries_l0[2], 'r--',label="EBR-l0, k/n=%s %%"%(int(round(100*k[2]/n, 0))))
plt.plot(np.arange(max_epoch)+1, plot_X_entries_l0[3], 'k--',label="EBR-l0, k/n=%s %%"%(int(round(100*k[3]/n, 0))))
plt.grid(True)
plt.title('Percentage of entries of X recovered by EBR and EBR-l0 varying k/n')
plt.xlabel('Epochs')
plt.ylabel('Entries of X recovered (%)')
plt.xlim(1,plot_epoch)
# plt.legend()
plt.savefig('./figures_ACHA/EBR_vs_EBR_l0_comparison.eps', format='eps', bbox_inches='tight')
print(np.count_nonzero(np.sum(X[0],axis=1)))
```
| github_jupyter |
# The Monty Hall problem, with lists
For inspiration, see this simulation of [the Monty Hall
Problem](../more-simulation/monty_hall) using arrays.
We use arrays often in data science, but sometimes, it is more efficient to use
Python [lists](../data-types/lists).
To follow along in this section, you will also need [more on
lists](more_on_lists).
## Simulating one trial
To operate on lists we use the Python standard `random` module, instead of the Numpy `random` module. The Numpy module always returns arrays, but in our case, we want to return lists.
```
import random
```
In particular, we are going to use the `shuffle` function in the Python
standard `random` module.
```
doors = ['car', 'goat', 'goat']
random.shuffle(doors)
doors
```
Here we chose a door at random. We use the standard `random.choice` instead of
`np.random.choice`.
```
my_door_index = random.choice([0, 1, 2])
my_door_index
```
We get the result of staying with our original choice, and remove that option from the list of available doors.
```
stay_result = doors.pop(my_door_index)
stay_result
```
We are left with the two doors that Monty has to choose from.
```
doors
```
Behind one of these doors, Monty knows there is a goat. He opens the door. We simulate that by removing the first door with a goat behind it.
[Remember](more_on_lists), `remove` removes only the first instance of "goat", leaving the second, if there is one.
```
doors.remove('goat')
doors
```
Now we have only one remaining door. The item behind that door is the result from switching from our original door.
```
switch_result = doors[0]
switch_result
```
## Many trials.
That's one trial. Now let's do that 100000 times.
Here we are using `range` instead of `np.arange`. `range` is the standard
Python equivalent of `np.arange`; it has the same effect, in this case, when we
use it in a loop.
```
# Make 10000 trials.
n_tries = 100000
# Lists to store results from stay and switch strategy
stay_results = []
switch_results = []
for i in range(n_tries):
# Same code as above, for one trial
doors = ['car', 'goat', 'goat']
random.shuffle(doors)
my_door_index = random.choice([0, 1, 2])
stay_result = doors.pop(my_door_index)
doors.remove('goat')
switch_result = doors[0]
# Put results into result lists
stay_results.append(stay_result)
switch_results.append(switch_result)
```
We use the `count` method of the list to count the number of "car" element in
each list, and divide by the length of the list, to get the proportion of
successes.
```
stay_results.count('car') / n_tries
switch_results.count('car') / n_tries
```
Compare this solution to the [solution using arrays](../more-simulation/monty_hall). Which solution is easier to read and understand?
| github_jupyter |
# Predicting Student Admissions with Neural Networks in Keras
In this notebook, we predict student admissions to graduate school at UCLA based on three pieces of data:
- GRE Scores (Test)
- GPA Scores (Grades)
- Class rank (1-4)
The dataset originally came from here: http://www.ats.ucla.edu/
## Loading the data
To load the data and format it nicely, we will use two very useful packages called Pandas and Numpy. You can read on the documentation here:
- https://pandas.pydata.org/pandas-docs/stable/
- https://docs.scipy.org/
```
# Importing pandas and numpy
import pandas as pd
import numpy as np
# Reading the csv file into a pandas DataFrame
data = pd.read_csv('student_data.csv')
# Printing out the first 10 rows of our data
data[:10]
```
## Plotting the data
First let's make a plot of our data to see how it looks. In order to have a 2D plot, let's ingore the rank.
```
# Importing matplotlib
import matplotlib.pyplot as plt
# Function to help us plot
def plot_points(data):
X = np.array(data[["gre","gpa"]])
y = np.array(data["admit"])
admitted = X[np.argwhere(y==1)]
rejected = X[np.argwhere(y==0)]
plt.scatter([s[0][0] for s in rejected], [s[0][1] for s in rejected], s = 25, color = 'red', edgecolor = 'k')
plt.scatter([s[0][0] for s in admitted], [s[0][1] for s in admitted], s = 25, color = 'cyan', edgecolor = 'k')
plt.xlabel('Test (GRE)')
plt.ylabel('Grades (GPA)')
# Plotting the points
plot_points(data)
plt.show()
```
Roughly, it looks like the students with high scores in the grades and test passed, while the ones with low scores didn't, but the data is not as nicely separable as we hoped it would. Maybe it would help to take the rank into account? Let's make 4 plots, each one for each rank.
```
# Separating the ranks
data_rank1 = data[data["rank"]==1]
data_rank2 = data[data["rank"]==2]
data_rank3 = data[data["rank"]==3]
data_rank4 = data[data["rank"]==4]
# Plotting the graphs
plot_points(data_rank1)
plt.title("Rank 1")
plt.show()
plot_points(data_rank2)
plt.title("Rank 2")
plt.show()
plot_points(data_rank3)
plt.title("Rank 3")
plt.show()
plot_points(data_rank4)
plt.title("Rank 4")
plt.show()
```
This looks more promising, as it seems that the lower the rank, the higher the acceptance rate. Let's use the rank as one of our inputs. In order to do this, we should one-hot encode it.
## One-hot encoding the rank
For this, we'll use the `get_dummies` function in pandas.
```
# Make dummy variables for rank
one_hot_data = pd.concat([data, pd.get_dummies(data['rank'], prefix='rank')], axis=1)
# Drop the previous rank column
one_hot_data = one_hot_data.drop('rank', axis=1)
# Print the first 10 rows of our data
one_hot_data[:10]
```
## Scaling the data
The next step is to scale the data. We notice that the range for grades is 1.0-4.0, whereas the range for test scores is roughly 200-800, which is much larger. This means our data is skewed, and that makes it hard for a neural network to handle. Let's fit our two features into a range of 0-1, by dividing the grades by 4.0, and the test score by 800.
```
# Copying our data
processed_data = one_hot_data[:]
# Scaling the columns
processed_data['gre'] = processed_data['gre']/800
processed_data['gpa'] = processed_data['gpa']/4.0
processed_data[:10]
```
## Splitting the data into Training and Testing
In order to test our algorithm, we'll split the data into a Training and a Testing set. The size of the testing set will be 10% of the total data.
```
sample = np.random.choice(processed_data.index, size=int(len(processed_data)*0.9), replace=False)
train_data, test_data = processed_data.iloc[sample], processed_data.drop(sample)
print("Number of training samples is", len(train_data))
print("Number of testing samples is", len(test_data))
print(train_data[:10])
print(test_data[:10])
```
## Splitting the data into features and targets (labels)
Now, as a final step before the training, we'll split the data into features (X) and targets (y).
Also, in Keras, we need to one-hot encode the output. We'll do this with the `to_categorical function`.
```
import keras
# Separate data and one-hot encode the output
# Note: We're also turning the data into numpy arrays, in order to train the model in Keras
features = np.array(train_data.drop('admit', axis=1))
targets = np.array(keras.utils.to_categorical(train_data['admit'], 2))
features_test = np.array(test_data.drop('admit', axis=1))
targets_test = np.array(keras.utils.to_categorical(test_data['admit'], 2))
print(features[:10])
print(targets[:10])
```
## Defining the model architecture
Here's where we use Keras to build our neural network.
```
# Imports
import numpy as np
from keras.models import Sequential
from keras.layers.core import Dense, Dropout, Activation
from keras.optimizers import SGD
from keras.utils import np_utils
# Building the model
model = Sequential()
model.add(Dense(128, activation='relu', input_shape=(6,)))
model.add(Dropout(.2))
model.add(Dense(64, activation='relu'))
model.add(Dropout(.1))
model.add(Dense(2, activation='softmax'))
# Compiling the model
model.compile(loss = 'categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
model.summary()
```
## Training the model
```
# Training the model
model.fit(features, targets, epochs=200, batch_size=100, verbose=0)
```
## Scoring the model
```
# Evaluating the model on the training and testing set
score = model.evaluate(features, targets)
print("\n Training Accuracy:", score[1])
score = model.evaluate(features_test, targets_test)
print("\n Testing Accuracy:", score[1])
```
## Challenge: Play with the parameters!
You can see that we made several decisions in our training. For instance, the number of layers, the sizes of the layers, the number of epochs, etc.
It's your turn to play with parameters! Can you improve the accuracy? The following are other suggestions for these parameters. We'll learn the definitions later in the class:
- Activation function: relu and sigmoid
- Loss function: categorical_crossentropy, mean_squared_error
- Optimizer: rmsprop, adam, ada
| github_jupyter |
```
import numpy as np
%matplotlib inline
import matplotlib.pyplot as plt
import scipy.integrate as integrate
import scipy.optimize as optimize
import scipy.stats as stats
```
# Regression without knowing the underlying model
In the first part of the lesson, we studied linear regression and made a crucial assumption: to estimate the parameter, we knew that the underling model was linear. What happens if we remove this assumption?
To understand it, we will still look at a regression task but we relax this assumption as follows: we know that the underlying model is a polynomial with additive noise:
$$
y(i) = p^*(x(i)) + \epsilon(i),
$$
where $p^*$ is an unknown polynomial that we need to estimate.
```
t = 0.5
sigma = 0.2
def model(x):
return t*x**3 + sigma*np.random.randn()
D = np.random.uniform(-2,2,25)
D = np.sort(D)
print(D)
Y = [model(d) for d in D]
plt.scatter(D,Y)
D_plot = np.arange(-2, 2.1, 0.015)
plt.plot(D_plot, t*D_plot**3)
```
## What goes wrong with our previous approach?
We can define a quadratic loss:
$$
J(p) = \frac{1}{m}\sum_{i=1}^m \left(y(i)-p(x(i)) \right)^2,
$$
and minimizes it among all polynomials.
```
z = np.polyfit(D, Y, len(D)-1)
p = np.poly1d(z)
plt.scatter(D,Y)
plt.plot(D_plot, t*D_plot**3)
plt.ylim(-4,4)
# plt.xlim(-2,2)
plt.plot(D_plot, p(D_plot))
```
Great, we achieve a loss of 0 but clearly our solution does not seem right!
Let cheat a bit a see what happens if we are looking at a solution with the right degree:
```
z = np.polyfit(D, Y, 5)
p = np.poly1d(z)
plt.scatter(D, Y)
plt.plot(D_plot, t*D_plot**3)
plt.ylim(-4, 4)
plt.plot(D_plot, p(D_plot))
```
OK, this looks much better, but we cheated! Indeed, you can modify the degree for your polynomial fit and see that it is not easy to decide between degrees 3,4,5...
We can now formalize our problem as follows. Given the points $(x(i),y(i))$, we need to do two things:
- decide on the degree of the polynomial $p^*$;
- once the degree is fixed, estimate the parameters of the polynomial.
One natural way to deal with this new formulation of the problem is to check all possible degrees and make an estimation of the parameters for each possible choice. But then, we need to decide which degree to select. In order to do that, we will split the dataset in a training set and a validation set. We will use the training set to estimate the parameters of the polynomial for all possible degrees. To decide which degree we should select, we will compute the loss of the obtained polynomial on the validation set and pick the one with minimal validation loss.
Let see if this works?
```
D_train = D[1::2]
Y_train = Y[1::2]
D_val = D[0::2]
Y_val = Y[0::2]
plt.scatter(D_train, Y_train)
plt.scatter(D_val, Y_val)
def get_error(deg):
val_error = np.zeros(deg)
train_error = np.zeros(deg)
for i in range(deg):
z = np.polyfit(D_train, Y_train, i)
p = np.poly1d(z)
train_error[i] = (np.mean((p(D_train)-Y_train)**2))
val_error[i] = (np.mean((p(D_val)-Y_val)**2))
return train_error, val_error
train_error, val_error = get_error(len(D_train)-1)
plt.figure(figsize=(14,7))
plt.plot(train_error, label='Training error')
plt.ylim(0, 3)
plt.plot(val_error, label='Validation error')
plt.xlabel("Degree")
plt.legend()
```
We see that the error on the training set is decreasing until it reaches 0 when the polynomial is able to inpterpolate all the points of the training set.
The error on the validation set is first decreasing as the training error but then starts to increase again. This is because the polynomial interpolating through the points of the training set is now missing a lot of points of the validation set, as shown below:
```
z = np.polyfit(D_train, Y_train, len(D_train)-1)
p = np.poly1d(z)
plt.scatter(D_val, Y_val)
plt.scatter(D_train, Y_train)
plt.plot(D_plot, t*D_plot**3)
plt.ylim(-4, 4)
plt.plot(D_plot, p(D_plot))
```
To summarize:
- for low degrees, our parametric model is not expressive enough to capture the true model resulting in a high error both on the training and validation set.
- for high degrees, our model becomes very expressive and start to actually fit the noise in the dataset, resulting in a low error on the training set and a high error on the validation set.
To formalize a bit what happens, we need to introduce the notion of risk. For an estimator $f:\mathbb{R}\to\mathbb{R}$, we define the risk as:
$$
\mathcal{R}(f) = \mathbb{E}\left[(f(X)-Y)^2\right],
$$
where the average is taken over randomness of $(X,Y)$. In our case, we simulate the true model with $X\sim Unif[-2,2]$ and $Y = 0.5*X^3+\sigma \epsilon$, where $\epsilon\sim \mathcal{N}(0,1)$.
We also define $\mathcal{H}_k$ as the set of polynomial of maximum degree $k$. $\mathcal{H}_k$ is the hypothesis space. We denote by $H_\infty$ the set of all polynomials.
Our goal is to find:
$$
p^* = \arg\min_{p\in \mathcal{H}_\infty}\mathcal{R}(p).
$$
In our case, we can compute the risk:
\begin{eqnarray*}
\min_{p\in \mathcal{H}_\infty}\mathcal{R}(p) &=& \mathbb{E}\left[(Y-\mathbb{E}[Y|X])^2\right]\\
&=& \mathbb{E}\left[ (Y-0.5 X^3)^2\right]\\
&=& \sigma^2
\end{eqnarray*}
But in practice, we do not have access to the true underlying model defining the distribution of $(X,Y)$, hence we are not able to evaluate the average defining the risk.
Hence, we define the empirical risk:
$$
\hat{\mathcal{R}}(p) = \frac{1}{m}\sum_{i=1}^m \left(y(i)-p((x(i))\right)^2,
$$
which is an approximation of the true risk.
To be more precise, we defined two differetn empirical risks: for $\hat{\mathcal{R}}_{train}(p)$, the average is taken over the training set and for $\hat{\mathcal{R}}_{val}(p)$, the average is taken over the validation set.
Now we define the polynomial of degree at most $k$ minimizing the empirical risk on the training set:
$$
\hat{p}_k = \arg\min_{p\in \mathcal{H}_k}\hat{\mathcal{R}}_{train}(p).
$$
The training error above is given by $\hat{\mathcal{R}}_{train}(\hat{p}_k)$ and the validation error by $\hat{\mathcal{R}}_{val}(\hat{p}_k)$.
Since the data points in the validation set are not used for the polynomial fit, we have $\hat{\mathcal{R}}_{val}(\hat{p}_k)\approx \mathcal{R}(\hat{p}_k)$.
Unfortunately, what we would like to compute is
$$
p^*_k = \arg\min_{p\in \mathcal{H}_k}\mathcal{R}(p).
$$
Note that $\mathcal{R}(p^*_k) \downarrow \mathcal{R}(p^*)$ as $k\to \infty$.
Unfortunately, our experiment above shows us that $p^*_k\neq \hat{p}_k$, especially for large values of $k$.
In all cases, we can decompose the risk of our estimator in the following non-negative terms:
$$
\mathcal{R}(\hat{p}_k) = \underbrace{\mathcal{R}(\hat{p}_k)-\mathcal{R}(p^*_k)}_{(1)} + \underbrace{\mathcal{R}(p^*_k)-\mathcal{R}(p^*)}_{(2)}+\mathcal{R}(p^*).
$$
The first term is called the **estimation error**, the second term is called the **approximation error** and the last term $\mathcal{R}(p^*)$ is the true risk.
Clearly, as $k\to \infty$, the approximation error (2) vanishes as our model becomes more and more expressive. In our case, the approximation error is 0 for $k\geq 3$. But the estimation error (1) will unfortunately grows with $k$. Going back to our empirical findings above, we see that for low values of $k$, $\mathcal{R}(\hat{p}_k)$ is high because the approximation error (2) is high and the for high values of $k$, $\mathcal{R}(\hat{p}_k)$ is high because the estimation error (1) is high. In practice, we take the minimum of this curve as an estimate of $\mathcal{R}(p^*)$.
```
def get_risk(deg):
risk = np.zeros(deg)
for i in range(deg):
z = np.polyfit(D_train, Y_train, i)
p = np.poly1d(z)
fun_int= lambda e,x : (p(x)-t*x**3 - sigma*e)**2*stats.norm.pdf(e)/4
risk[i] = integrate.dblquad(fun_int,-2,2,lambda x: -10, lambda x: 10)[0]
return risk
risk = get_risk(len(D_train)-1)
def opti_risk(x):
if int(x) == 0:
return t**2*2**6/7+sigma**2
elif int(x) == 1:
return t**2*2**6/7+sigma**2-t**2*2**6/5**2*3
elif int(x) == 2:
pass# left as an exercise!
else:
return sigma**2
deg = [0,1,3,4,5,6,7,8,9,10]
plt.figure(figsize=(14,7))
plt.ylim(0, 3)
plt.plot(deg, [risk[int(d)]-opti_risk(d) for d in deg], color='red',label='estimation error')
plt.plot(deg, [opti_risk(d)-sigma**2 for d in deg],color='green',label='approximation error')
plt.plot(val_error, label='empirical validation error')
plt.xlabel("Degree")
plt.legend()
plt.figure(figsize=(14,7))
plt.ylim(0, 3)
plt.plot(risk, color='red',label='risk of esimator $\mathcal{R}(\hat{p}_k)$')
plt.plot(val_error, label='empirical validation error $\hat{\mathcal{R}}_{val}(\hat{p}_k)$')
plt.xlabel("Degree")
plt.legend()
```
| github_jupyter |
#### Download and install TensorFlow2 package. Import TensorFlow into your program
```
from __future__ import absolute_import, division, print_function, unicode_literals
import tensorflow as tf
from tensorflow.keras.layers import Dense, Flatten, Conv2D
from tensorflow.keras import Model
```
#### Load and prepare the MNIST dataset
```
mnist = tf.keras.datasets.mnist
(x_train, y_train), (x_test, y_test) = mnist.load_data()
x_train, x_test = x_train/255.0, x_test/255.0
# Add a channels dimension
x_train = x_train[..., tf.newaxis]
x_test = x_test[..., tf.newaxis]
```
#### Use `tf.data` to batch and shuffle the dataset
```
train_ds = tf.data.Dataset.from_tensor_slices((x_train, y_train)).shuffle(10000).batch(32)
test_ds = tf.data.Dataset.from_tensor_slices((x_test, y_test)).batch(32)
```
#### Build the `tf.keras` model using the Keras model subclassing API
```
class MyModel(Model):
def __init__(self):
super(MyModel, self).__init__()
self.conv1 = Conv2D(32, 3, activation='relu')
self.flatten = Flatten()
self.d1 = Dense(128, activation='relu')
self.d2 = Dense(10, activation='softmax')
def call(self, x):
x = self.conv1(x)
x = self.flatten(x)
x = self.d1(x)
return self.d2(x)
# Create an instance of the model
model = MyModel()
```
#### Choose an optimizer and loss function for training
```
loss_object = tf.keras.losses.SparseCategoricalCrossentropy()
optimizer = tf.keras.optimizers.Adam()
```
#### Select metrics to measure the loss and the accuracy of the model. These metrics accumulate the values over epochs and then print the overall result
```
train_loss = tf.keras.metrics.Mean(name='train_loss')
train_accuracy = tf.keras.metrics.SparseCategoricalAccuracy(name='train_accuracy')
test_loss = tf.keras.metrics.Mean(name='test_loss')
test_accuracy = tf.keras.metrics.SparseCategoricalAccuracy(name='test_accuracy')
```
#### Use `tf.GradientTape` to train the model
```
@tf.function
def train_step(images, labels):
with tf.GradientTape() as tape:
predictions = model(images)
loss = loss_object(labels, predictions)
gradients = tape.gradient(loss, model.trainable_variables)
optimizer.apply_gradients(zip(gradients, model.trainable_variables))
train_loss(loss)
train_accuracy(labels, predictions)
```
#### Test the model
```
@tf.function # what is the role of @ in front of tf? (@tf)
def test_step(images, labels):
predictions = model(images)
t_loss = loss_object(labels, predictions)
test_loss(t_loss)
test_accuracy(labels, predictions)
EPOCHS = 5
for epoch in range(EPOCHS):
#Reset the metrics at the start of the next epoch
train_loss.reset_states()
train_accuracy.reset_states()
test_loss.reset_states()
test_accuracy.reset_states()
for images, labels in train_ds:
train_step(images, labels)
for test_images, test_labels in test_ds:
test_step(test_images, test_labels)
template = 'Epoch: {}, Loss: {}, Accuracy: {}, Test Loss: {}, Test Accuracy: {}'
print(template.format(epoch+1,
train_loss.result(),
train_accuracy.result()*100,
test_loss.result(),
test_accuracy.result()*100))
```
end -
##### reference
https://www.tensorflow.org/tutorials/quickstart/advanced
| github_jupyter |
```
# coding: utf-8
# python2.7
from __future__ import division, print_function
from parsers import CitationWindowParser # use parse and empty_parse methods
from context_parsing_functions import *
from indexing_functions import *
from os import listdir, makedirs
from os.path import isfile, exists
from collections import defaultdict
import re
import subprocess
from multiprocessing import Pool
INDEXES_DIR = "../Retrieval/Indexes/" # ../Retrieval/Indexes/
COLLECTIONS_DIR = "../Documents/PF+PN_Collection/XML/"
DOC_DIR = COLLECTIONS_DIR + "test/" # "PF+PN/"
CONTEXT_DIR = COLLECTIONS_DIR + "C.l080b080a.manual_clean/" # "C.l080b080a.raw", "C.l080b080a.detex"
WINDOW_SPECS = {'Sentence': [(3,3)], 'Word': [(50,50)]}
# {'Sentence': [(0,0),(1,0),(1,1),(2,0),(2,1),(2,2),(3,0),(3,1),(3,2),(3,3),(4,0),(4,1),(4,2),(4,3),(4,4),(5,0),(5,1),(5,2),(5,3),(5,4),(5,5)], 'Word': [(25,0),(25,25),(50,0),(50,25),(50,50),(75,0),(75,25),(75,50),(75,75),(100,0),(100,25),(100,50),(100,75),(100,100)]}
def append_citation_context_windows(isearch_file, doc_dir=DOC_DIR):
"""
Appends sentence and word-based citation context windows
to isearch document content in a new file.
Args:
isearch_file (str)
doc_dir=DOC_DIR (str)
"""
# prepare document text
print("...Reading: {}".format(isearch_file))
doc_text = prepare_document_text(doc_dir + isearch_file)
# prepare citation windows
print("...Preparing citation windows")
citation_windows = prepare_citation_windows(isearch_file)
# write files with document text and citation context windows appended
print("...Writing text files to collections\n")
write_files_with_cit_windows(isearch_file, doc_text, citation_windows)
def prepare_citation_windows(isearch_file, context_dir=CONTEXT_DIR):
"""
Prepares XML-formatted citation context windows
of different sizes for each citation context.
Args:
isearch_file (str)
context_dir=CONTEXT_DIR (str)
Returns:
citation_windows (dict of list of str)
"""
citation_windows = defaultdict(list)
# initiate parsers
parsers = create_context_parsers(WINDOW_SPECS)
# identify if file with citation contexts exists
contexts_file = "{}-citation_contexts.xml".format(find_isearch_id(isearch_file))
if contexts_file in listdir(context_dir):
# prepare citation context texts
citation_contexts = prepare_context_texts(context_dir + contexts_file)
# parse citation contexts into windows
for context in citation_contexts:
for parser in parsers:
stringy_name, window = parser.parse(context)
citation_windows[stringy_name].append(window)
# otherwise...
else:
# make empty windows
for parser in parsers:
stringy_name, window = parser.empty_parse()
citation_windows[stringy_name] = window
return citation_windows
def prepare_document_text(filepath):
"""
Extracts content from an iSearch XML file,
returns content within an <ISEARCHDOC> XML field.
Args:
filepath (str)
Returns:
formatted_text (str)
"""
with open(filepath) as doc:
text = doc.readlines()
begin_index = text.index('<DOC>\n')
text.insert(begin_index + 1, '<ISEARCHDOC>\n')
text[text.index('</DOC>\n')] = '</ISEARCHDOC>\n'
formatted_text = "".join(text)
return formatted_text
def find_isearch_id(filename):
"""
Returns an isearch id when found as an XML filename.
Args:
filename (str)
Returns:
isearch_id (str)
"""
isearch_id = re.search('(P[FN][0-9]{6})\.xml', filename).group(1)
return isearch_id
def write_files_with_cit_windows(isearch_file, doc_text, citation_windows, collection_dir=COLLECTIONS_DIR):
"""
Writes isearch document text and citation context windows
to a new file.
Args:
isearch_file (str)
doc_text (str)
citation_windows (dict of list of str)
collection_dir=COLLECTIONS_DIR (str)
"""
for size, windows in citation_windows.items():
windows = "".join(windows)
write_dir = collection_dir + "PF+PN+C." + size + "/"
if not exists(write_dir):
makedirs(write_dir)
with open(write_dir + isearch_file, 'a') as combo:
combo.write("{}<CITATIONS>\n{}</CITATIONS>\n</DOC>"
.format(doc_text, windows))
return
def main():
p = Pool(1)
p.map(append_citation_context_windows, listdir(DOC_DIR))
index_collections(WINDOW_SPECS, INDEXES_DIR, COLLECTIONS_DIR)
if __name__ == '__main__':
main()
def main():
p = Pool(1)
p.map(append_citation_context_windows, listdir(DOC_DIR))
# create_context_parsers()
# print(prepare_context_texts("../Append_Citation_Contexts/Cits/PF309700-citation_contexts.xml"))
if __name__ == '__main__':
main()
import subprocess
text =""" la la ala \begin{abstract}
I present a toy model for the Berkovits pure spinor superparticle.
It is a $D=1$, $\N=2$ superparticle with no physical degrees of
freedom. We study the cohomology in various ways, in particular
finding an explicit expression for the `b'-field. Finally, we
construct the topological string B-model from a straightforward
generalization of the system.
\end{abstract}
%\pacs{}% PACS, the Physics and Astronomy
% Classification Scheme.
\keywords{BRST, Superstring, Pure spinors}
\maketitle
"""
# "/usr/texbin/detex"
def detex_text(text):
detex = subprocess.Popen(["detex", "-t"], stdout=subprocess.PIPE, stdin=subprocess.PIPE, stderr=subprocess.STDOUT)
detex_stdout = detex.communicate(input=text)[0]
return detex_stdout.decode()
detex_text(text)
```
| github_jupyter |
# *CoNNear*: A convolutional neural-network model of human cochlear mechanics and filter tuning for real-time applications
Python notebook for reproducing the evaluation results of the proposed CoNNear model.
## Prerequisites
- First, let us compile the cochlea_utils.c file that is used for solving the transmission line (TL) model of the cochlea. This requires some C++ compiler which should be installed beforehand. Then go the connear folder from the terminal and run:
```
gcc -shared -fpic -O3 -ffast-math -o tridiag.so cochlea_utils.c
```
- Install numpy, scipy, keras and tensorflow
## Import required python packages and functions
Import required python packages and load the connear model.
```
import numpy as np
from scipy import signal
import matplotlib.pyplot as plt
import keras
from keras.models import model_from_json
from keras.utils import CustomObjectScope
from keras.initializers import glorot_uniform
from tlmodel.get_tl_vbm_and_oae import tl_vbm_and_oae
json_file = open("connear/Gmodel.json", "r")
loaded_model_json = json_file.read()
json_file.close()
connear = model_from_json(loaded_model_json)
connear.load_weights("connear/Gmodel.h5")
connear.summary()
```
Define some functions here
```
def rms (x):
# compute rms of a matrix
sq = np.mean(np.square(x), axis = 0)
return np.sqrt(sq)
# Define model specific variables
down_rate = 2
fs = 20e3
fs_tl = 100e3
p0 = 2e-5
factor_fs = int(fs_tl / fs)
right_context = 256
left_context = 256
# load CFs
CF = np.loadtxt('tlmodel/cf.txt')
```
## Click response
Compare the responses of the models to a click stimulus.
**Notice that for all the simulations, TL model operates at 100kHz and the CoNNear model operates at 20kHz.**
```
#Define the click stimulus
dur = 128.0e-3 # for 2560 samples #CONTEXT
click_duration = 2 # 100 us click
stim = np.zeros((1, int(dur * fs)))
L = 70.0
samples = dur * fs
click_duration = 2 # 100 us click
click_duration_tl = factor_fs * click_duration
silence = 60 #samples in silence
samples = int(samples - right_context - left_context)
'''
# GET TL model response
stim = np.zeros((1, (samples + right_context + left_context)*factor_fs))
stim[0, (factor_fs * (right_context+silence)) : (factor_fs * (right_context+silence)) + click_duration_tl] = 2 * np.sqrt(2) * p0 * 10**(L/20)
output = tl_vbm_and_oae(stim , L)
CF = output[0]['cf'][::down_rate]
# basilar membrane motion for click response
# the context samples (first and last 256 samples)
# are removed. Also downsample it to 20kHz
bmm_click_out_full = np.array(output[0]['v'])
stimrange = range(right_context*factor_fs, (right_context*factor_fs) + (factor_fs*samples))
bmm_click_tl = sp_sig.resample_poly(output[0]['v'][stimrange,::down_rate], fs, fs_tl)
bmm_click_tl = bmm_click_tl.T
'''
# Prepare the same for CoNNear model
stim = np.zeros((1, int(dur * fs)))
stim[0, right_context + silence : right_context + silence + click_duration] = 2 * np.sqrt(2) * p0 * 10**(L/20)
# Get the CoNNear response
stim = np.expand_dims(stim, axis=2)
connear_pred_click = connear.predict(stim.T, verbose=1)
bmm_click_connear = connear_pred_click[0,:,:].T * 1e-6
```
Plotting the results.
```
plt.plot(stim[0,256:-256]), plt.xlim(0,2000)
plt.show()
'''
plt.imshow(bmm_click_tl, aspect='auto', cmap='jet')
plt.xlim(0,2000), plt.clim(-4e-7,5e-7)
plt.colorbar()
plt.show()
'''
plt.imshow(bmm_click_connear, aspect='auto', cmap='jet')
plt.xlim(0,2000), plt.clim(-4e-7,5e-7)
plt.colorbar()
plt.show()
```
## Cochlear Excitation Patterns
Here, we plot the simulated RMS levels of basilar memberane (BM) displacement across CF for tone stimuli presented at SPLs between 0 and 90 dB SPL.
```
f_tone = 1e3 # You can change this tone frequency to see how the excitation pattern changes
# with stimulus frequency
fs = 20e3
p0 = 2e-5
dur = 102.4e-3 # for 2048 samples
window_len = int(fs * dur)
L = np.arange(0., 91.0, 10.) # SPLs from 0 to 90dB
#CoNNear
t = np.arange(0., dur, 1./fs)
hanlength = int(10e-3 * fs) # 10ms length hanning window
stim_sin = np.sin(2 * np.pi * f_tone * t)
han = signal.windows.hann(hanlength)
stim_sin[:int(hanlength/2)] = stim_sin[:int(hanlength/2)] * han[:int(hanlength/2)]
stim_sin[-int(hanlength/2):] = stim_sin[-int(hanlength/2):] * han[int(hanlength/2):]
stim = np.zeros((len(L), int(len(stim_sin))))
#total_length = 2560 #CONTEXT
total_length = window_len + right_context + left_context # CONTEXT
stim = np.zeros((len(L), total_length)) #CONTEXT
for j in range(len(L)):
stim[j,right_context:window_len+right_context] = p0 * np.sqrt(2) * 10**(L[j]/20) * stim_sin
# prepare for feeding to the DNN
stim = np.expand_dims(stim, axis=2)
connear_pred_tone = connear.predict(stim, verbose=1)
bmm_tone_connear = connear_pred_tone# * 1e-6
bmm_tone_connear.shape
# Compute rms for each level
cochlear_pred_tone_rms = np.vstack([rms(bmm_tone_connear[i]) for i in range(len(L))])
# Plot the RMS
cftile=np.tile(CF, (len(L),1))
plt.semilogx(cftile.T, 20.*np.log10(cochlear_pred_tone_rms.T))
plt.xlim(0.25,8.), plt.grid(which='both'),
plt.xticks(ticks=(0.25, 0.5, 1., 2., 4., 8.) , labels=(0.25, 0.5, 1., 2., 4., 8.))
plt.ylim(-80, 20)
plt.xlabel('CF (kHz)')
plt.ylabel('RMS of y_bm (dB)')
plt.title('CoNNear Predicted')
plt.show()
```
| github_jupyter |
##### Copyright 2020 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# MLMD Model Card Toolkit Demo
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/responsible_ai/model_card_toolkit/examples/MLMD_Model_Card_Toolkit_Demo"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/model-card-toolkit/blob/master/model_card_toolkit/documentation/examples/MLMD_Model_Card_Toolkit_Demo.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/model-card-toolkit/blob/master/model_card_toolkit/documentation/examples/MLMD_Model_Card_Toolkit_Demo.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/model-card-toolkit/model_card_toolkit/documentation/examples/MLMD_Model_Card_Toolkit_Demo.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table>
## Background
This notebook demonstrates how to generate a model card using the Model Card Toolkit with MLMD and TFX pipeline in a Jupyter/Colab environment. You can learn more about model cards at https://modelcards.withgoogle.com/about.
## Setup
We first need to a) install and import the necessary packages, and b) download the data.
### Upgrade to Pip 20.2 and Install TFX
```
!pip install --upgrade pip==20.2
!pip install "tfx==0.26.0"
!pip install model-card-toolkit
```
### Did you restart the runtime?
If you are using Google Colab, the first time that you run the cell above, you must restart the runtime (Runtime > Restart runtime ...). This is because of the way that Colab loads packages.
### Import packages
We import necessary packages, including standard TFX component classes and check the library versions.
```
import os
import pprint
import tempfile
import urllib
import absl
import tensorflow as tf
import tensorflow_model_analysis as tfma
tf.get_logger().propagate = False
pp = pprint.PrettyPrinter()
import tfx
from tfx.components import CsvExampleGen
from tfx.components import Evaluator
from tfx.components import Pusher
from tfx.components import ResolverNode
from tfx.components import SchemaGen
from tfx.components import StatisticsGen
from tfx.components import Trainer
from tfx.components import Transform
from tfx.components.base import executor_spec
from tfx.components.trainer.executor import GenericExecutor
from tfx.dsl.experimental import latest_blessed_model_resolver
from tfx.orchestration import metadata
from tfx.orchestration import pipeline
from tfx.orchestration.experimental.interactive.interactive_context import InteractiveContext
from tfx.proto import pusher_pb2
from tfx.proto import trainer_pb2
from tfx.types import Channel
from tfx.types.standard_artifacts import Model
from tfx.types.standard_artifacts import ModelBlessing
from tfx.utils.dsl_utils import external_input
import ml_metadata as mlmd
print('TensorFlow version: {}'.format(tf.__version__))
print('TFX version: {}'.format(tfx.version.__version__))
print('MLMD version: {}'.format(mlmd.__version__))
```
### Set up pipeline paths
```
# This is the root directory for your TFX pip package installation.
_tfx_root = tfx.__path__
# Set up logging.
absl.logging.set_verbosity(absl.logging.INFO)
```
### Download example data
We download the example dataset for use in our TFX pipeline.
```
DATA_PATH = 'https://archive.ics.uci.edu/ml/machine-learning-databases/adult/' \
'adult.data'
_data_root = tempfile.mkdtemp(prefix='tfx-data')
_data_filepath = os.path.join(_data_root, "data.csv")
urllib.request.urlretrieve(DATA_PATH, _data_filepath)
columns = [
"Age", "Workclass", "fnlwgt", "Education", "Education-Num", "Marital-Status",
"Occupation", "Relationship", "Race", "Sex", "Capital-Gain", "Capital-Loss",
"Hours-per-week", "Country", "Over-50K"]
with open(_data_filepath, 'r') as f:
content = f.read()
content = content.replace(", <=50K", ', 0').replace(", >50K", ', 1')
with open(_data_filepath, 'w') as f:
f.write(','.join(columns) + '\n' + content)
```
Take a quick look at the CSV file.
```
!head {_data_filepath}
```
### Create the InteractiveContext
Last, we create an InteractiveContext, which will allow us to run TFX components interactively in this notebook.
```
# Here, we create an InteractiveContext using default parameters. This will
# use a temporary directory with an ephemeral ML Metadata database instance.
# To use your own pipeline root or database, the optional properties
# `pipeline_root` and `metadata_connection_config` may be passed to
# InteractiveContext. Calls to InteractiveContext are no-ops outside of the
# notebook.
context = InteractiveContext()
```
## Run TFX components interactively
In the cells that follow, we create TFX components one-by-one, run each of them, and visualize their output artifacts. In this notebook, we won’t provide detailed explanations of each TFX component, but you can see what each does at [TFX Colab workshop](https://github.com/tensorflow/workshops/blob/master/tfx_labs/Lab_1_Pipeline_in_Colab.ipynb).
### ExampleGen
Create the `ExampleGen` component to split data into training and evaluation sets, convert the data into `tf.Example` format, and copy data into the `_tfx_root` directory for other components to access.
```
example_gen = CsvExampleGen(input=external_input(_data_root))
context.run(example_gen)
artifact = example_gen.outputs['examples'].get()[0]
print(artifact.split_names, artifact.uri)
```
Let’s take a look at the first three training examples:
```
# Get the URI of the output artifact representing the training examples, which is a directory
train_uri = os.path.join(example_gen.outputs['examples'].get()[0].uri, 'train')
# Get the list of files in this directory (all compressed TFRecord files)
tfrecord_filenames = [os.path.join(train_uri, name)
for name in os.listdir(train_uri)]
# Create a `TFRecordDataset` to read these files
dataset = tf.data.TFRecordDataset(tfrecord_filenames, compression_type="GZIP")
# Iterate over the first 3 records and decode them.
for tfrecord in dataset.take(3):
serialized_example = tfrecord.numpy()
example = tf.train.Example()
example.ParseFromString(serialized_example)
pp.pprint(example)
```
### StatisticsGen
`StatisticsGen` takes as input the dataset we just ingested using `ExampleGen` and allows you to perform some analysis of your dataset using TensorFlow Data Validation (TFDV).
```
statistics_gen = StatisticsGen(
examples=example_gen.outputs['examples'])
context.run(statistics_gen)
```
After `StatisticsGen` finishes running, we can visualize the outputted statistics. Try playing with the different plots!
```
context.show(statistics_gen.outputs['statistics'])
```
### SchemaGen
`SchemaGen` will take as input the statistics that we generated with `StatisticsGen`, looking at the training split by default.
```
schema_gen = SchemaGen(
statistics=statistics_gen.outputs['statistics'],
infer_feature_shape=False)
context.run(schema_gen)
context.show(schema_gen.outputs['schema'])
```
To learn more about schemas, see [the SchemaGen documentation](https://www.tensorflow.org/tfx/guide/schemagen).
### Transform
`Transform` will take as input the data from `ExampleGen`, the schema from `SchemaGen`, as well as a module that contains user-defined Transform code.
Let's see an example of user-defined Transform code below (for an introduction to the TensorFlow Transform APIs, [see the tutorial](https://www.tensorflow.org/tfx/tutorials/transform/simple)).
```
_census_income_constants_module_file = 'census_income_constants.py'
%%writefile {_census_income_constants_module_file}
# Categorical features are assumed to each have a maximum value in the dataset.
MAX_CATEGORICAL_FEATURE_VALUES = [20]
CATEGORICAL_FEATURE_KEYS = ["Education-Num"]
DENSE_FLOAT_FEATURE_KEYS = ["Capital-Gain", "Hours-per-week", "Capital-Loss"]
# Number of buckets used by tf.transform for encoding each feature.
FEATURE_BUCKET_COUNT = 10
BUCKET_FEATURE_KEYS = ["Age"]
# Number of vocabulary terms used for encoding VOCAB_FEATURES by tf.transform
VOCAB_SIZE = 200
# Count of out-of-vocab buckets in which unrecognized VOCAB_FEATURES are hashed.
OOV_SIZE = 10
VOCAB_FEATURE_KEYS = ["Workclass", "Education", "Marital-Status", "Occupation",
"Relationship", "Race", "Sex", "Country"]
# Keys
LABEL_KEY = "Over-50K"
def transformed_name(key):
return key + '_xf'
_census_income_transform_module_file = 'census_income_transform.py'
%%writefile {_census_income_transform_module_file}
import tensorflow as tf
import tensorflow_transform as tft
import census_income_constants
_DENSE_FLOAT_FEATURE_KEYS = census_income_constants.DENSE_FLOAT_FEATURE_KEYS
_VOCAB_FEATURE_KEYS = census_income_constants.VOCAB_FEATURE_KEYS
_VOCAB_SIZE = census_income_constants.VOCAB_SIZE
_OOV_SIZE = census_income_constants.OOV_SIZE
_FEATURE_BUCKET_COUNT = census_income_constants.FEATURE_BUCKET_COUNT
_BUCKET_FEATURE_KEYS = census_income_constants.BUCKET_FEATURE_KEYS
_CATEGORICAL_FEATURE_KEYS = census_income_constants.CATEGORICAL_FEATURE_KEYS
_LABEL_KEY = census_income_constants.LABEL_KEY
_transformed_name = census_income_constants.transformed_name
def preprocessing_fn(inputs):
"""tf.transform's callback function for preprocessing inputs.
Args:
inputs: map from feature keys to raw not-yet-transformed features.
Returns:
Map from string feature key to transformed feature operations.
"""
outputs = {}
for key in _DENSE_FLOAT_FEATURE_KEYS:
# Preserve this feature as a dense float, setting nan's to the mean.
outputs[_transformed_name(key)] = tft.scale_to_z_score(
_fill_in_missing(inputs[key]))
for key in _VOCAB_FEATURE_KEYS:
# Build a vocabulary for this feature.
outputs[_transformed_name(key)] = tft.compute_and_apply_vocabulary(
_fill_in_missing(inputs[key]),
top_k=_VOCAB_SIZE,
num_oov_buckets=_OOV_SIZE)
for key in _BUCKET_FEATURE_KEYS:
outputs[_transformed_name(key)] = tft.bucketize(
_fill_in_missing(inputs[key]), _FEATURE_BUCKET_COUNT)
for key in _CATEGORICAL_FEATURE_KEYS:
outputs[_transformed_name(key)] = _fill_in_missing(inputs[key])
label = _fill_in_missing(inputs[_LABEL_KEY])
outputs[_transformed_name(_LABEL_KEY)] = label
return outputs
def _fill_in_missing(x):
"""Replace missing values in a SparseTensor.
Fills in missing values of `x` with '' or 0, and converts to a dense tensor.
Args:
x: A `SparseTensor` of rank 2. Its dense shape should have size at most 1
in the second dimension.
Returns:
A rank 1 tensor where missing values of `x` have been filled in.
"""
default_value = '' if x.dtype == tf.string else 0
return tf.squeeze(
tf.sparse.to_dense(
tf.SparseTensor(x.indices, x.values, [x.dense_shape[0], 1]),
default_value),
axis=1)
transform = Transform(
examples=example_gen.outputs['examples'],
schema=schema_gen.outputs['schema'],
module_file=os.path.abspath(_census_income_transform_module_file))
context.run(transform)
transform.outputs
```
### Trainer
Let's see an example of user-defined model code below (for an introduction to the TensorFlow Keras APIs, [see the tutorial](https://www.tensorflow.org/guide/keras)):
```
_census_income_trainer_module_file = 'census_income_trainer.py'
%%writefile {_census_income_trainer_module_file}
from typing import List, Text
import os
import absl
import datetime
import tensorflow as tf
import tensorflow_transform as tft
from tfx.components.trainer.executor import TrainerFnArgs
import census_income_constants
_DENSE_FLOAT_FEATURE_KEYS = census_income_constants.DENSE_FLOAT_FEATURE_KEYS
_VOCAB_FEATURE_KEYS = census_income_constants.VOCAB_FEATURE_KEYS
_VOCAB_SIZE = census_income_constants.VOCAB_SIZE
_OOV_SIZE = census_income_constants.OOV_SIZE
_FEATURE_BUCKET_COUNT = census_income_constants.FEATURE_BUCKET_COUNT
_BUCKET_FEATURE_KEYS = census_income_constants.BUCKET_FEATURE_KEYS
_CATEGORICAL_FEATURE_KEYS = census_income_constants.CATEGORICAL_FEATURE_KEYS
_MAX_CATEGORICAL_FEATURE_VALUES = census_income_constants.MAX_CATEGORICAL_FEATURE_VALUES
_LABEL_KEY = census_income_constants.LABEL_KEY
_transformed_name = census_income_constants.transformed_name
def _transformed_names(keys):
return [_transformed_name(key) for key in keys]
def _gzip_reader_fn(filenames):
"""Small utility returning a record reader that can read gzip'ed files."""
return tf.data.TFRecordDataset(
filenames,
compression_type='GZIP')
def _get_serve_tf_examples_fn(model, tf_transform_output):
"""Returns a function that parses a serialized tf.Example and applies TFT."""
model.tft_layer = tf_transform_output.transform_features_layer()
@tf.function
def serve_tf_examples_fn(serialized_tf_examples):
"""Returns the output to be used in the serving signature."""
feature_spec = tf_transform_output.raw_feature_spec()
feature_spec.pop(_LABEL_KEY)
parsed_features = tf.io.parse_example(serialized_tf_examples, feature_spec)
transformed_features = model.tft_layer(parsed_features)
if _transformed_name(_LABEL_KEY) in transformed_features:
transformed_features.pop(_transformed_name(_LABEL_KEY))
return model(transformed_features)
return serve_tf_examples_fn
def _input_fn(file_pattern: List[Text],
tf_transform_output: tft.TFTransformOutput,
batch_size: int = 200) -> tf.data.Dataset:
"""Generates features and label for tuning/training.
Args:
file_pattern: List of paths or patterns of input tfrecord files.
tf_transform_output: A TFTransformOutput.
batch_size: representing the number of consecutive elements of returned
dataset to combine in a single batch
Returns:
A dataset that contains (features, indices) tuple where features is a
dictionary of Tensors, and indices is a single Tensor of label indices.
"""
transformed_feature_spec = (
tf_transform_output.transformed_feature_spec().copy())
dataset = tf.data.experimental.make_batched_features_dataset(
file_pattern=file_pattern,
batch_size=batch_size,
features=transformed_feature_spec,
reader=_gzip_reader_fn,
label_key=_transformed_name(_LABEL_KEY))
return dataset
def _build_keras_model(hidden_units: List[int] = None) -> tf.keras.Model:
"""Creates a DNN Keras model.
Args:
hidden_units: [int], the layer sizes of the DNN (input layer first).
Returns:
A keras Model.
"""
real_valued_columns = [
tf.feature_column.numeric_column(key, shape=())
for key in _transformed_names(_DENSE_FLOAT_FEATURE_KEYS)
]
categorical_columns = [
tf.feature_column.categorical_column_with_identity(
key, num_buckets=_VOCAB_SIZE + _OOV_SIZE, default_value=0)
for key in _transformed_names(_VOCAB_FEATURE_KEYS)
]
categorical_columns += [
tf.feature_column.categorical_column_with_identity(
key, num_buckets=_FEATURE_BUCKET_COUNT, default_value=0)
for key in _transformed_names(_BUCKET_FEATURE_KEYS)
]
categorical_columns += [
tf.feature_column.categorical_column_with_identity( # pylint: disable=g-complex-comprehension
key,
num_buckets=num_buckets,
default_value=0) for key, num_buckets in zip(
_transformed_names(_CATEGORICAL_FEATURE_KEYS),
_MAX_CATEGORICAL_FEATURE_VALUES)
]
indicator_column = [
tf.feature_column.indicator_column(categorical_column)
for categorical_column in categorical_columns
]
model = _wide_and_deep_classifier(
# TODO(b/139668410) replace with premade wide_and_deep keras model
wide_columns=indicator_column,
deep_columns=real_valued_columns,
dnn_hidden_units=hidden_units or [100, 70, 50, 25])
return model
def _wide_and_deep_classifier(wide_columns, deep_columns, dnn_hidden_units):
"""Build a simple keras wide and deep model.
Args:
wide_columns: Feature columns wrapped in indicator_column for wide (linear)
part of the model.
deep_columns: Feature columns for deep part of the model.
dnn_hidden_units: [int], the layer sizes of the hidden DNN.
Returns:
A Wide and Deep Keras model
"""
# Following values are hard coded for simplicity in this example,
# However prefarably they should be passsed in as hparams.
# Keras needs the feature definitions at compile time.
# TODO(b/139081439): Automate generation of input layers from FeatureColumn.
input_layers = {
colname: tf.keras.layers.Input(name=colname, shape=(), dtype=tf.float32)
for colname in _transformed_names(_DENSE_FLOAT_FEATURE_KEYS)
}
input_layers.update({
colname: tf.keras.layers.Input(name=colname, shape=(), dtype='int32')
for colname in _transformed_names(_VOCAB_FEATURE_KEYS)
})
input_layers.update({
colname: tf.keras.layers.Input(name=colname, shape=(), dtype='int32')
for colname in _transformed_names(_BUCKET_FEATURE_KEYS)
})
input_layers.update({
colname: tf.keras.layers.Input(name=colname, shape=(), dtype='int32')
for colname in _transformed_names(_CATEGORICAL_FEATURE_KEYS)
})
# TODO(b/161816639): SparseFeatures for feature columns + Keras.
deep = tf.keras.layers.DenseFeatures(deep_columns)(input_layers)
for numnodes in dnn_hidden_units:
deep = tf.keras.layers.Dense(numnodes)(deep)
wide = tf.keras.layers.DenseFeatures(wide_columns)(input_layers)
output = tf.keras.layers.Dense(
1, activation='sigmoid')(
tf.keras.layers.concatenate([deep, wide]))
model = tf.keras.Model(input_layers, output)
model.compile(
loss='binary_crossentropy',
optimizer=tf.keras.optimizers.Adam(lr=0.001),
metrics=[tf.keras.metrics.BinaryAccuracy()])
model.summary(print_fn=absl.logging.info)
return model
# TFX Trainer will call this function.
def run_fn(fn_args: TrainerFnArgs):
"""Train the model based on given args.
Args:
fn_args: Holds args used to train the model as name/value pairs.
"""
# Number of nodes in the first layer of the DNN
first_dnn_layer_size = 100
num_dnn_layers = 4
dnn_decay_factor = 0.7
tf_transform_output = tft.TFTransformOutput(fn_args.transform_output)
train_dataset = _input_fn(fn_args.train_files, tf_transform_output, 40)
eval_dataset = _input_fn(fn_args.eval_files, tf_transform_output, 40)
model = _build_keras_model(
# Construct layers sizes with exponetial decay
hidden_units=[
max(2, int(first_dnn_layer_size * dnn_decay_factor**i))
for i in range(num_dnn_layers)
])
# This log path might change in the future.
log_dir = os.path.join(os.path.dirname(fn_args.serving_model_dir), 'logs')
tensorboard_callback = tf.keras.callbacks.TensorBoard(
log_dir=log_dir, update_freq='batch')
model.fit(
train_dataset,
steps_per_epoch=fn_args.train_steps,
validation_data=eval_dataset,
validation_steps=fn_args.eval_steps,
callbacks=[tensorboard_callback])
signatures = {
'serving_default':
_get_serve_tf_examples_fn(model,
tf_transform_output).get_concrete_function(
tf.TensorSpec(
shape=[None],
dtype=tf.string,
name='examples')),
}
model.save(fn_args.serving_model_dir, save_format='tf', signatures=signatures)
trainer = Trainer(
module_file=os.path.abspath(_census_income_trainer_module_file),
custom_executor_spec=executor_spec.ExecutorClassSpec(GenericExecutor),
examples=transform.outputs['transformed_examples'],
transform_graph=transform.outputs['transform_graph'],
schema=schema_gen.outputs['schema'],
train_args=trainer_pb2.TrainArgs(num_steps=100),
eval_args=trainer_pb2.EvalArgs(num_steps=50))
context.run(trainer)
```
### Evaluator
The `Evaluator` component computes model performance metrics over the evaluation set. It uses the [TensorFlow Model Analysis](https://www.tensorflow.org/tfx/model_analysis/get_started) library.
`Evaluator` will take as input the data from `ExampleGen`, the trained model from `Trainer`, and slicing configuration. The slicing configuration allows you to slice your metrics on feature values. See an example of this configuration below:
```
from google.protobuf.wrappers_pb2 import BoolValue
eval_config = tfma.EvalConfig(
model_specs=[
# This assumes a serving model with signature 'serving_default'. If
# using estimator based EvalSavedModel, add signature_name: 'eval' and
# remove the label_key.
tfma.ModelSpec(label_key="Over-50K")
],
metrics_specs=[
tfma.MetricsSpec(
# The metrics added here are in addition to those saved with the
# model (assuming either a keras model or EvalSavedModel is used).
# Any metrics added into the saved model (for example using
# model.compile(..., metrics=[...]), etc) will be computed
# automatically.
# To add validation thresholds for metrics saved with the model,
# add them keyed by metric name to the thresholds map.
metrics=[
tfma.MetricConfig(class_name='ExampleCount'),
tfma.MetricConfig(class_name='BinaryAccuracy'),
tfma.MetricConfig(class_name='FairnessIndicators',
config='{ "thresholds": [0.5] }'),
]
)
],
slicing_specs=[
# An empty slice spec means the overall slice, i.e. the whole dataset.
tfma.SlicingSpec(),
# Data can be sliced along a feature column. In this case, data is
# sliced by feature column Race and Sex.
tfma.SlicingSpec(feature_keys=['Race']),
tfma.SlicingSpec(feature_keys=['Sex']),
tfma.SlicingSpec(feature_keys=['Race', 'Sex']),
],
options = tfma.Options(compute_confidence_intervals=BoolValue(value=True))
)
# Use TFMA to compute a evaluation statistics over features of a model and
# validate them against a baseline.
evaluator = Evaluator(
examples=example_gen.outputs['examples'],
model=trainer.outputs['model'],
eval_config=eval_config)
context.run(evaluator)
evaluator.outputs
```
Using the `evaluation` output we can show the default visualization of global metrics on the entire evaluation set.
```
context.show(evaluator.outputs['evaluation'])
```
## Populate Properties from ModelCard with Model Card Toolkit
Now that we’ve set up our TFX pipeline, we will use the Model Card Toolkit to extract key artifacts from the run and populate a Model Card.
### Connect to the MLMD store used by the InteractiveContext
```
from ml_metadata.metadata_store import metadata_store
from IPython import display
mlmd_store = metadata_store.MetadataStore(context.metadata_connection_config)
model_uri = trainer.outputs["model"].get()[0].uri
```
### Use Model Card Toolkit
#### Initialize the Model Card Toolkit.
```
from model_card_toolkit import ModelCardToolkit
mct = ModelCardToolkit(mlmd_store=mlmd_store, model_uri=model_uri)
```
#### Create Model Card workspace.
```
model_card = mct.scaffold_assets()
```
#### Annotate more information into Model Card.
It is also important to document model information that might be important to downstream users, such as its limitations, intended use cases, trade offs, and ethical considerations. For each of these sections, we can directly add new JSON objects to represent this information.
```
model_card.model_details.name = 'Census Income Classifier'
model_card.model_details.overview = (
'This is a wide and deep Keras model which aims to classify whether or not '
'an individual has an income of over $50,000 based on various demographic '
'features. The model is trained on the UCI Census Income Dataset. This is '
'not a production model, and this dataset has traditionally only been used '
'for research purposes. In this Model Card, you can review quantitative '
'components of the model’s performance and data, as well as information '
'about the model’s intended uses, limitations, and ethical considerations.'
)
model_card.model_details.owners = [
{'name': 'Model Cards Team', 'contact': 'model-cards@google.com'}
]
model_card.considerations.use_cases = [
'This dataset that this model was trained on was originally created to '
'support the machine learning community in conducting empirical analysis '
'of ML algorithms. The Adult Data Set can be used in fairness-related '
'studies that compare inequalities across sex and race, based on '
'people’s annual incomes.'
]
model_card.considerations.limitations = [
'This is a class-imbalanced dataset across a variety of sensitive classes.'
' The ratio of male-to-female examples is about 2:1 and there are far more'
' examples with the “white” attribute than every other race combined. '
'Furthermore, the ratio of $50,000 or less earners to $50,000 or more '
'earners is just over 3:1. Due to the imbalance across income levels, we '
'can see that our true negative rate seems quite high, while our true '
'positive rate seems quite low. This is true to an even greater degree '
'when we only look at the “female” sub-group, because there are even '
'fewer female examples in the $50,000+ earner group, causing our model to '
'overfit these examples. To avoid this, we can try various remediation '
'strategies in future iterations (e.g. undersampling, hyperparameter '
'tuning, etc), but we may not be able to fix all of the fairness issues.'
]
model_card.considerations.ethical_considerations = [{
'name':
'We risk expressing the viewpoint that the attributes in this dataset '
'are the only ones that are predictive of someone’s income, even '
'though we know this is not the case.',
'mitigation_strategy':
'As mentioned, some interventions may need to be performed to address '
'the class imbalances in the dataset.'
}]
```
#### Filter and Add Graphs.
We can filter the graphs generated by the TFX components to include those most relevant for the Model Card using the function defined below. In this example, we filter for `race` and `sex`, two potentially sensitive attributes.
Each Model Card will have up to three sections for graphs -- training dataset statistics, evaluation dataset statistics, and quantitative analysis of our model’s performance.
```
# These are the graphs that will appear in the Quantiative Analysis portion of
# the Model Card. Feel free to add or remove from this list.
TARGET_EVAL_GRAPH_NAMES = [
'fairness_indicators_metrics/false_positive_rate@0.5',
'fairness_indicators_metrics/false_negative_rate@0.5',
'binary_accuracy',
'example_count | Race_X_Sex',
]
# These are the graphs that will appear in both the Train Set and Eval Set
# portions of the Model Card. Feel free to add or remove from this list.
TARGET_DATASET_GRAPH_NAMES = [
'counts | Race',
'counts | Sex',
]
def filter_graphs(graphics, target_graph_names):
result = []
for graph in graphics:
for target_graph_name in target_graph_names:
if graph.name.startswith(target_graph_name):
result.append(graph)
result.sort(key=lambda g: g.name)
return result
# Populating the three different sections using the filter defined above. To
# see all the graphs available in a section, we can iterate through each of the
# different collections.
model_card.quantitative_analysis.graphics.collection = filter_graphs(
model_card.quantitative_analysis.graphics.collection, TARGET_EVAL_GRAPH_NAMES)
model_card.model_parameters.data.eval.graphics.collection = filter_graphs(
model_card.model_parameters.data.eval.graphics.collection, TARGET_DATASET_GRAPH_NAMES)
model_card.model_parameters.data.train.graphics.collection = filter_graphs(
model_card.model_parameters.data.train.graphics.collection, TARGET_DATASET_GRAPH_NAMES)
```
We then add (optional) descriptions for each of the each of the graph sections.
```
model_card.model_parameters.data.train.graphics.description = (
'This section includes graphs displaying the class distribution for the '
'“Race” and “Sex” attributes in our training dataset. We chose to '
'show these graphs in particular because we felt it was important that '
'users see the class imbalance.'
)
model_card.model_parameters.data.eval.graphics.description = (
'Like the training set, we provide graphs showing the class distribution '
'of the data we used to evaluate our model’s performance. '
)
model_card.quantitative_analysis.graphics.description = (
'These graphs show how the model performs for data sliced by “Race”, '
'“Sex” and the intersection of these attributes. The metrics we chose '
'to display are “Accuracy”, “False Positive Rate”, and “False '
'Negative Rate”, because we anticipated that the class imbalances might '
'cause our model to underperform for certain groups.'
)
mct.update_model_card_json(model_card)
```
#### Generate the Model Card.
We can now display the Model Card in HTML format.
```
html = mct.export_format()
display.display(display.HTML(html))
```
| github_jupyter |
# Navigation
---
In this notebook, you will learn how to use the Unity ML-Agents environment for the first project of the [Deep Reinforcement Learning Nanodegree](https://www.udacity.com/course/deep-reinforcement-learning-nanodegree--nd893).
### 1. Start the Environment
We begin by importing some necessary packages. If the code cell below returns an error, please revisit the project instructions to double-check that you have installed [Unity ML-Agents](https://github.com/Unity-Technologies/ml-agents/blob/master/docs/Installation.md) and [NumPy](http://www.numpy.org/).
```
from unityagents import UnityEnvironment
import numpy as np
```
Next, we will start the environment! **_Before running the code cell below_**, change the `file_name` parameter to match the location of the Unity environment that you downloaded.
- **Mac**: `"path/to/Banana.app"`
- **Windows** (x86): `"path/to/Banana_Windows_x86/Banana.exe"`
- **Windows** (x86_64): `"path/to/Banana_Windows_x86_64/Banana.exe"`
- **Linux** (x86): `"path/to/Banana_Linux/Banana.x86"`
- **Linux** (x86_64): `"path/to/Banana_Linux/Banana.x86_64"`
- **Linux** (x86, headless): `"path/to/Banana_Linux_NoVis/Banana.x86"`
- **Linux** (x86_64, headless): `"path/to/Banana_Linux_NoVis/Banana.x86_64"`
For instance, if you are using a Mac, then you downloaded `Banana.app`. If this file is in the same folder as the notebook, then the line below should appear as follows:
```
env = UnityEnvironment(file_name="Banana.app")
```
```
env = UnityEnvironment(file_name="./Banana.app")
#env = UnityEnvironment(file_name="./Banana_Linux_NoVis/Banana.x86_64") # non visual
```
Environments contain **_brains_** which are responsible for deciding the actions of their associated agents. Here we check for the first brain available, and set it as the default brain we will be controlling from Python.
```
# get the default brain
brain_name = env.brain_names[0]
brain = env.brains[brain_name]
```
### 2. Examine the State and Action Spaces
The simulation contains a single agent that navigates a large environment. At each time step, it has four actions at its disposal:
- `0` - walk forward
- `1` - walk backward
- `2` - turn left
- `3` - turn right
The state space has `37` dimensions and contains the agent's velocity, along with ray-based perception of objects around agent's forward direction. A reward of `+1` is provided for collecting a yellow banana, and a reward of `-1` is provided for collecting a blue banana.
Run the code cell below to print some information about the environment.
```
# reset the environment
env_info = env.reset(train_mode=True)[brain_name]
# number of agents in the environment
print('Number of agents:', len(env_info.agents))
# number of actions
action_size = brain.vector_action_space_size
print('Number of actions:', action_size)
# examine the state space
state = env_info.vector_observations[0]
print('States look like:', state)
state_size = len(state)
print('States have length:', state_size)
```
### 3. Take Random Actions in the Environment
In the next code cell, you will learn how to use the Python API to control the agent and receive feedback from the environment.
Once this cell is executed, you will watch the agent's performance, if it selects an action (uniformly) at random with each time step. A window should pop up that allows you to observe the agent, as it moves through the environment.
Of course, as part of the project, you'll have to change the code so that the agent is able to use its experience to gradually choose better actions when interacting with the environment!
env_info = env.reset(train_mode=False)[brain_name] # reset the environment
state = env_info.vector_observations[0] # get the current state
score = 0 # initialize the score
while True:
action = np.random.randint(action_size) # select an action
env_info = env.step(action)[brain_name] # send the action to the environment
next_state = env_info.vector_observations[0] # get the next state
reward = env_info.rewards[0] # get the reward
done = env_info.local_done[0] # see if episode has finished
score += reward # update the score
state = next_state # roll over the state to next time step
if done: # exit loop if episode finished
break
print("Score: {}".format(score))
### 4. It's Your Turn!
Now it's your turn to train your own agent to solve the environment! When training the environment, set `train_mode=True`, so that the line for resetting the environment looks like the following:
```python
env_info = env.reset(train_mode=True)[brain_name]
```
```
from dqn_agent import Agent
import torch
device = torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')
from collections import deque, namedtuple
import matplotlib.pyplot as plt
%matplotlib inline
# Some default settings
EPS_START = 1.0
EPS_END = 0.01
EPS_DECAY = 0.995
MAX_T = 1000
N_EPISODES = 2000
# how often to update the network
def dqn(n_episodes = N_EPISODES, max_t = MAX_T, eps_start = EPS_START, eps_end = EPS_END, eps_decay = EPS_DECAY):
scores = []
scores_recent = []
eps = eps_start
for i_episode in range(1, n_episodes+1):
env_info = env.reset(train_mode=True)[brain_name]
state = env_info.vector_observations[0] # get the initial state
score = 0
for i_t in range(max_t): # considered as episodic with 100 steps
action = agent.act(state, eps) # select an action
env_info = env.step(action)[brain_name] # send the action to the environment
next_state = env_info.vector_observations[0] # get the next state
reward = env_info.rewards[0] # get the reward
done = env_info.local_done[0] # see if episode has finished
agent.step(state, action, reward, next_state, done) # Sample & Learn step of Q learning
score += reward # update the score
state = next_state # roll over the state to next time step
if done: # exit loop if episode finished
break
scores.append(score)
scores_recent.append(score)
eps = max(eps_end, eps*eps_decay) # decrease epsilon
print('\rEpisode {}\tAverage Score: {:.2f}'.format(i_episode, np.mean(scores_recent)), end="")
if i_episode % 100 == 0:
print('\rEpisode {}\tAverage Score: {:.2f}'.format(i_episode, np.mean(scores_recent)))
if np.mean(scores_recent)>=13.0:
print('\nEnvironment solved in {:d} episodes!\tAverage Score: {:.2f}'.format(i_episode-100, np.mean(scores_recent)))
torch.save(agent.qnetwork_local.state_dict(), 'checkpoint.pth')
break
return scores
BUFFER_SIZE = int(1e5)
BATCH_SIZE = 64
GAMMA = 0.99
TAU = 1e-3
LR = 5e-4
UPDATE_EVERY = 4
EPS_START = 1.0
EPS_END = 0.01
EPS_DECAY = 0.995
MAX_T = 1000
N_EPISODES = 100
agent = Agent(state_size, action_size, seed=0, buffer_size = BUFFER_SIZE, batch_size = BATCH_SIZE,
gamma = GAMMA, tau = TAU, lr = LR, update_every = UPDATE_EVERY)
scores = dqn(n_episodes = N_EPISODES, max_t = MAX_T, eps_start = EPS_START, eps_end = EPS_END, eps_decay = EPS_DECAY)
# plot the scores
fig = plt.figure()
ax = fig.add_subplot(111)
plt.plot(np.arange(len(scores)), scores)
plt.ylabel('Score')
plt.xlabel('Episode #')
plt.show()
```
When finished, you can close the environment.
```
env.close()
```
| github_jupyter |
# XAI
## Links
### [SOTA Resources Listing](https://github.com/rehmanzafar/xai-iml-sota)
### Application Frameworks:
- [ExplainerDashboard](https://explainerdashboard.readthedocs.io/en/latest/index.html)
- [ExplainX](https://github.com/explainX/explainx)
- [Explainer](https://explainer.ai/)
### Toolkits (many are integrated into Application Frameworks):
- [SHAP](https://shap.readthedocs.io/en/latest/index.html)
- [LIME](https://github.com/marcotcr/lime)
- [shapash](https://shapash.readthedocs.io/en/latest/)
- [Shapley-Flow](https://github.com/nathanwang000/Shapley-Flow)
- [DALEX](https://github.com/ModelOriented/DALEX)
- [tf-explain](https://tf-explain.readthedocs.io/en/latest/)
### Articles:
- [x] [Explainble AI (XAI) -- A guide to 7 python packages to explain your models](https://towardsdatascience.com/explainable-ai-xai-a-guide-to-7-packages-in-python-to-explain-your-models-932967f0634b)
- [x] [5 Explainable Machine Learning Models You Should Understand](https://towardsdatascience.com/explainable-ai-9a9af94931ff)
- [SHAPling around a custom network](https://towardsdatascience.com/shapling-around-a-custom-network-636b97b40628)
- [Are ML models interpretable](https://gowtamsingulur.medium.com/are-ml-models-interpretable-ba4765f4be40)
- [x] [AI Explainability Requires Robustness](https://towardsdatascience.com/ai-explainability-requires-robustness-2028ac200e9a)
- [x] [New frontiers in Explainable AI](https://towardsdatascience.com/new-frontiers-in-explainable-ai-af43bba18348)
- [Can SHAP trigger a paradigm shift in Risk Analytics?](https://towardsdatascience.com/can-shap-trigger-a-paradigm-shift-in-risk-analytics-c01278e4dd77)
- [Explainable AI at OLX: understanding models by understanding data](https://tech.olx.com/explainable-ai-at-olx-understanding-models-by-understanding-data-c8905ee688ae)
- [Are we thinking about eXplAInability backwards?](https://towardsdatascience.com/are-we-thinking-about-explainability-backwards-c9a719cb1250)
- [x] [The most important types of XAI](https://towardsdatascience.com/the-most-important-types-of-xai-72f5beb9e77e)
- [x] [Understanding SHAP for Interpretable Machine Learning](https://ai.plainenglish.io/understanding-shap-for-interpretable-machine-learning-35e8639d03db)
- [x] [Feature Attribution in Explainable AI](https://medium.com/geekculture/feature-attribution-in-explainable-ai-626f0a1d95e2)
- [x] [Behold: The Confusion Matrix](https://vijayasriiyer.medium.com/behold-the-confusion-matrix-10afd3feb603)
- [Counterfactuals and their Evaluation](https://medium.com/@urjapawar/counterfactuals-and-their-evaluation-574ef58d34ac)
- [x] [How not to stumble while evaluating interpretability?](https://medium.com/geekculture/how-not-to-stumble-while-evaluating-interpretability-3af6c122621)
- [Explainable AI (XAI) with Class Maps](https://towardsdatascience.com/explainable-ai-xai-with-class-maps-d0e137a91d2c)
- [Class Activation Maps in Deep Learning](https://valentinaalto.medium.com/class-activation-maps-in-deep-learning-14101e2ec7e1)
- [x] [Building Confidence on Explainability Methods](https://towardsdatascience.com/building-confidence-on-explainability-methods-66b9ee575514)
- [Boruta SHAP: A Tool for Feature Selection Every Data Scientist Should Know](https://towardsdatascience.com/boruta-shap-an-amazing-tool-for-feature-selection-every-data-scientist-should-know-33a5f01285c0)
- [x] [Shapley Value aided Recipe for Materials Design](https://towardsdatascience.com/shapley-value-aided-recipe-for-materials-design-d122fb851cef)
- [x] [Deep Dive into Neural Network Explanations with Integrated Gradients](https://towardsdatascience.com/deep-dive-into-neural-network-explanations-with-integrated-gradients-7f4b7be855a2)
- [x] [Should you explain your predictions with SHAP or IG?](https://towardsdatascience.com/should-you-explain-your-predictions-with-shap-or-ig-9cabe218b5cc)
- [Explainable AI (XAI) Methods Part 3 — Accumulated Local Effects (ALE)](https://towardsdatascience.com/explainable-ai-xai-methods-part-3-accumulated-local-effects-ale-cf6ba3387fde)
- [Alternative Feature Selection Methods in Machine Learning](https://www.kdnuggets.com/2021/12/alternative-feature-selection-methods-machine-learning.html)
- [The Power of Feature Engineering](https://towardsdatascience.com/the-power-of-feature-engineering-b6f3bb7de39c)
- [Hypothesis Tests Explained](https://towardsdatascience.com/hypothesis-tests-explained-8a070636bd28)
- [How to do Bayesian hyper-parameter tuning on a blackbox model](https://towardsdatascience.com/how-to-do-bayesian-hyper-parameter-tuning-on-a-blackbox-model-882009552c6d)
- [How many samples do you need?](https://towardsdatascience.com/how-many-samples-do-you-need-simulation-in-r-a60891a6e549)
- [Monte Carlo Value at Risk (VaR) in R](https://towardsdev.com/monte-carlo-value-at-risk-var-in-r-960e62f0f53f)
- [Feature Engineering Techniques](https://singhharsh246.medium.com/feature-engineering-techniques-c1a5a8d202df)
- [Feature Engineering Libraries in Python …](https://paulaleksis.medium.com/feature-engineering-libraries-in-python-6ea5dd0aec79)
- [x] [Explainable Defect Detection Using Convolutional Neural Networks: Case Study](https://medium.com/geekculture/explainable-defect-detection-using-convolutional-neural-networks-case-study-2b58bc17c8b1)
- [x] [A Hands-on Introduction to Explaining Neural Networks with TruLens](https://medium.com/trulens/a-hands-on-introduction[-to-explaining-neural-networks-with-trulens-504bfab1a578)
- [Gain Trust in Your Model and Generate Explanations With LIME and SHAP](https://towardsdatascience.com/gain-trust-in-your-model-and-generate-explanations-with-lime-and-shap-94288694c154)
- [Explainable Boosting Machines](https://pub.towardsai.net/explainable-boosting-machines-c71b207231b5)
- [Which Explanation Method Should You Use for Your Machine Learning Model?](https://medium.datadriveninvestor.com/which-explanation-method-should-you-use-for-your-machine-learning-model-1e38e1011225)
- [How to Train Deep Neural Networks Over Data Streams](https://towardsdatascience.com/how-to-train-deep-neural-networks-over-data-streams-fdab15704e66)
- [x] [Explainable AI](https://link.medium.com/QN7ugaSjGnb)
- [x] [Interpretable Machine Learning using SHAP — theory and applications](https://medium.com/@khalilz/interpretable-machine-learning-using-shap-theory-and-applications-26c12f7a7f1a)
- [x] [A Novel Approach to Feature Importance — Shapley Additive Explanations](https://towardsdatascience.com/a-novel-approach-to-feature-importance-shapley-additive-explanations-d18af30fc21b)
- [x] [Explain your model using SHAP values](https://towardsdatascience.com/explain-your-model-with-the-shap-values-bc36aac4de3d)
### Notebooks:
- [SHAP Materials Design](https://github.com/Bjoyita/SHAP_MaterialsDesign)
- [The SHAP Values with More Elegant Charts](https://github.com/dataman-git/codes_for_articles/blob/master/The%20SHAP%20values%20with%20More%20Charts%20for%20article.ipynb)
- [Explain Your Model with Microsoft's InterpretML](https://github.com/dataman-git/codes_for_articles/blob/master/Explain%20Your%20Model%20with%20Microsoft's%20InterpretML-Github.ipynb)
### Talks:
- [Explainable AI for Science and Medicine by Scott Lundberg](https://www.youtube.com/watch?v=B-c8tIgchu0)
### Relevant books:
- [Interpretable Machine Learning](https://christophm.github.io/interpretable-ml-book)
- [Explanatory Model Analysis](https://ema.drwhy.ai/)
- [Interpretable AI](https://www.manning.com/books/interpretable-ai)
- [Limits of Interpretable Machine Learning Methods](https://compstat-lmu.github.io/iml_methods_limitations/)
### Relevant courses:
- [Trustworthy Deep Learning](https://berkeley-deep-learning.github.io/cs294-131-s19/)
# Knowledge Graph (static)
```
import graphviz
with open('assets/xai-mind-map.gv', 'r') as f:
diagram_src = f.read()
graphviz.Source(diagram_src)
```
## Problems
### 1. Too much information to look at
### 2. Too difficult to Configure
### 3. Requires constant manual updates
### 4. Doesn't update using a map generator
---
# Knowledge Graph Refactoring
## 1. Provide a more stylistic view

## 2. Add a left side that is related to pre-model processing

## 3. Add Links that will open Notebooks

## 4. Highlight Prior Presentations

## 5. Knowledge Graph generation (underlying development structure)

| github_jupyter |
##### Copyright 2018 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#@title MIT License
#
# Copyright (c) 2017 François Chollet
#
# Permission is hereby granted, free of charge, to any person obtaining a
# copy of this software and associated documentation files (the "Software"),
# to deal in the Software without restriction, including without limitation
# the rights to use, copy, modify, merge, publish, distribute, sublicense,
# and/or sell copies of the Software, and to permit persons to whom the
# Software is furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in
# all copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
# IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
# FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL
# THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
# LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING
# FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER
# DEALINGS IN THE SOFTWARE.
```
# 過学習と学習不足について知る
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/ja/r1/tutorials/keras/overfit_and_underfit.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/ja/r1/tutorials/keras/overfit_and_underfit.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
</table>
Note: これらのドキュメントは私たちTensorFlowコミュニティが翻訳したものです。コミュニティによる 翻訳は**ベストエフォート**であるため、この翻訳が正確であることや[英語の公式ドキュメント](https://www.tensorflow.org/?hl=en)の 最新の状態を反映したものであることを保証することはできません。 この翻訳の品質を向上させるためのご意見をお持ちの方は、GitHubリポジトリ[tensorflow/docs](https://github.com/tensorflow/docs)にプルリクエストをお送りください。 コミュニティによる翻訳やレビューに参加していただける方は、 [docs-ja@tensorflow.org メーリングリスト](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs-ja)にご連絡ください。
いつものように、この例のプログラムは`tf.keras` APIを使用します。詳しくはTensorFlowの[Keras guide](https://www.tensorflow.org/r1/guide/keras)を参照してください。
これまでの例、つまり、映画レビューの分類と燃費の推定では、検証用データでのモデルの正解率が、数エポックでピークを迎え、その後低下するという現象が見られました。
言い換えると、モデルが訓練用データを**過学習**したと考えられます。過学習への対処の仕方を学ぶことは重要です。**訓練用データセット**で高い正解率を達成することは難しくありませんが、我々は、(これまで見たこともない)**テスト用データ**に汎化したモデルを開発したいのです。
過学習の反対語は**学習不足**(underfitting)です。学習不足は、モデルがテストデータに対してまだ改善の余地がある場合に発生します。学習不足の原因は様々です。モデルが十分強力でないとか、正則化のしすぎだとか、単に訓練時間が短すぎるといった理由があります。学習不足は、訓練用データの中の関連したパターンを学習しきっていないということを意味します。
モデルの訓練をやりすぎると、モデルは過学習を始め、訓練用データの中のパターンで、テストデータには一般的ではないパターンを学習します。我々は、過学習と学習不足の中間を目指す必要があります。これから見ていくように、ちょうどよいエポック数だけ訓練を行うというのは必要なスキルなのです。
過学習を防止するための、最良の解決策は、より多くの訓練用データを使うことです。多くのデータで訓練を行えば行うほど、モデルは自然により汎化していく様になります。これが不可能な場合、次善の策は正則化のようなテクニックを使うことです。正則化は、モデルに保存される情報の量とタイプに制約を課すものです。ネットワークが少数のパターンしか記憶できなければ、最適化プロセスにより、最も主要なパターンのみを学習することになり、より汎化される可能性が高くなります。
このノートブックでは、重みの正則化とドロップアウトという、よく使われる2つの正則化テクニックをご紹介します。これらを使って、IMDBの映画レビューを分類するノートブックの改善を図ります。
```
import tensorflow.compat.v1 as tf
from tensorflow import keras
import numpy as np
import matplotlib.pyplot as plt
print(tf.__version__)
```
## IMDBデータセットのダウンロード
以前のノートブックで使用したエンベディングの代わりに、ここでは文をマルチホットエンコードします。このモデルは、訓練用データセットをすぐに過学習します。このモデルを使って、過学習がいつ起きるかということと、どうやって過学習と戦うかをデモします。
リストをマルチホットエンコードすると言うのは、0と1のベクトルにするということです。具体的にいうと、例えば`[3, 5]`というシーケンスを、インデックス3と5の値が1で、それ以外がすべて0の、10,000次元のベクトルに変換するということを意味します。
```
NUM_WORDS = 10000
(train_data, train_labels), (test_data, test_labels) = keras.datasets.imdb.load_data(num_words=NUM_WORDS)
def multi_hot_sequences(sequences, dimension):
# 形状が (len(sequences), dimension)ですべて0の行列を作る
results = np.zeros((len(sequences), dimension))
for i, word_indices in enumerate(sequences):
results[i, word_indices] = 1.0 # 特定のインデックスに対してresults[i] を1に設定する
return results
train_data = multi_hot_sequences(train_data, dimension=NUM_WORDS)
test_data = multi_hot_sequences(test_data, dimension=NUM_WORDS)
```
結果として得られるマルチホットベクトルの1つを見てみましょう。単語のインデックスは頻度順にソートされています。このため、インデックスが0に近いほど1が多く出現するはずです。分布を見てみましょう。
```
plt.plot(train_data[0])
```
## 過学習のデモ
過学習を防止するための最も単純な方法は、モデルのサイズ、すなわち、モデル内の学習可能なパラメータの数を小さくすることです(学習パラメータの数は、層の数と層ごとのユニット数で決まります)。ディープラーニングでは、モデルの学習可能なパラメータ数を、しばしばモデルの「キャパシティ」と呼びます。直感的に考えれば、パラメータ数の多いモデルほど「記憶容量」が大きくなり、訓練用のサンプルとその目的変数の間の辞書のようなマッピングをたやすく学習することができます。このマッピングには汎化能力がまったくなく、これまで見たことが無いデータを使って予測をする際には役に立ちません。
ディープラーニングのモデルは訓練用データに適応しやすいけれど、本当のチャレレンジは汎化であって適応ではないということを、肝に銘じておく必要があります。
一方、ネットワークの記憶容量が限られている場合、前述のようなマッピングを簡単に学習することはできません。損失を減らすためには、より予測能力が高い圧縮された表現を学習しなければなりません。同時に、モデルを小さくしすぎると、訓練用データに適応するのが難しくなります。「多すぎる容量」と「容量不足」の間にちょうどよい容量があるのです。
残念ながら、(層の数や、層ごとの大きさといった)モデルの適切なサイズやアーキテクチャを決める魔法の方程式はありません。一連の異なるアーキテクチャを使って実験を行う必要があります。
適切なモデルのサイズを見つけるには、比較的少ない層の数とパラメータから始めるのがベストです。それから、検証用データでの損失値の改善が見られなくなるまで、徐々に層の大きさを増やしたり、新たな層を加えたりします。映画レビューの分類ネットワークでこれを試してみましょう。
比較基準として、```Dense```層だけを使ったシンプルなモデルを構築し、その後、それより小さいバージョンと大きいバージョンを作って比較します。
### 比較基準を作る
```
baseline_model = keras.Sequential([
# `.summary` を見るために`input_shape`が必要
keras.layers.Dense(16, activation=tf.nn.relu, input_shape=(NUM_WORDS,)),
keras.layers.Dense(16, activation=tf.nn.relu),
keras.layers.Dense(1, activation=tf.nn.sigmoid)
])
baseline_model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy', 'binary_crossentropy'])
baseline_model.summary()
baseline_history = baseline_model.fit(train_data,
train_labels,
epochs=20,
batch_size=512,
validation_data=(test_data, test_labels),
verbose=2)
```
### より小さいモデルの構築
今作成したばかりの比較基準となるモデルに比べて隠れユニット数が少ないモデルを作りましょう。
```
smaller_model = keras.Sequential([
keras.layers.Dense(4, activation=tf.nn.relu, input_shape=(NUM_WORDS,)),
keras.layers.Dense(4, activation=tf.nn.relu),
keras.layers.Dense(1, activation=tf.nn.sigmoid)
])
smaller_model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy', 'binary_crossentropy'])
smaller_model.summary()
```
同じデータを使って訓練します。
```
smaller_history = smaller_model.fit(train_data,
train_labels,
epochs=20,
batch_size=512,
validation_data=(test_data, test_labels),
verbose=2)
```
### より大きなモデルの構築
練習として、より大きなモデルを作成し、どれほど急速に過学習が起きるかを見ることもできます。次はこのベンチマークに、この問題が必要とするよりはるかに容量の大きなネットワークを追加しましょう。
```
bigger_model = keras.models.Sequential([
keras.layers.Dense(512, activation=tf.nn.relu, input_shape=(NUM_WORDS,)),
keras.layers.Dense(512, activation=tf.nn.relu),
keras.layers.Dense(1, activation=tf.nn.sigmoid)
])
bigger_model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy','binary_crossentropy'])
bigger_model.summary()
```
このモデルもまた同じデータを使って訓練します。
```
bigger_history = bigger_model.fit(train_data, train_labels,
epochs=20,
batch_size=512,
validation_data=(test_data, test_labels),
verbose=2)
```
### 訓練時と検証時の損失をグラフにする
<!--TODO(markdaoust): This should be a one-liner with tensorboard -->
実線は訓練用データセットの損失、破線は検証用データセットでの損失です(検証用データでの損失が小さい方が良いモデルです)。これをみると、小さいネットワークのほうが比較基準のモデルよりも過学習が始まるのが遅いことがわかります(4エポックではなく6エポック後)。また、過学習が始まっても性能の低下がよりゆっくりしています。
```
def plot_history(histories, key='binary_crossentropy'):
plt.figure(figsize=(16,10))
for name, history in histories:
val = plt.plot(history.epoch, history.history['val_'+key],
'--', label=name.title()+' Val')
plt.plot(history.epoch, history.history[key], color=val[0].get_color(),
label=name.title()+' Train')
plt.xlabel('Epochs')
plt.ylabel(key.replace('_',' ').title())
plt.legend()
plt.xlim([0,max(history.epoch)])
plot_history([('baseline', baseline_history),
('smaller', smaller_history),
('bigger', bigger_history)])
```
より大きなネットワークでは、すぐに、1エポックで過学習が始まり、その度合も強いことに注目してください。ネットワークの容量が大きいほど訓練用データをモデル化するスピードが早くなり(結果として訓練時の損失値が小さくなり)ますが、より過学習しやすく(結果として訓練時の損失値と検証時の損失値が大きく乖離しやすく)なります。
## 戦略
### 重みの正則化を加える
「オッカムの剃刀」の原則をご存知でしょうか。何かの説明が2つあるとすると、最も正しいと考えられる説明は、仮定の数が最も少ない「一番単純な」説明だというものです。この原則は、ニューラルネットワークを使って学習されたモデルにも当てはまります。ある訓練用データとネットワーク構造があって、そのデータを説明できる重みの集合が複数ある時(つまり、複数のモデルがある時)、単純なモデルのほうが複雑なものよりも過学習しにくいのです。
ここで言う「単純なモデル」とは、パラメータ値の分布のエントロピーが小さいもの(あるいは、上記で見たように、そもそもパラメータの数が少ないもの)です。したがって、過学習を緩和するための一般的な手法は、重みが小さい値のみをとることで、重み値の分布がより整然となる(正則)様に制約を与えるものです。これを「重みの正則化」と呼ばれ、ネットワークの損失関数に、重みの大きさに関連するコストを加えることで行われます。このコストには2つの種類があります。
* [L1正則化](https://developers.google.com/machine-learning/glossary/#L1_regularization) 重み係数の絶対値に比例するコストを加える(重みの「L1ノルム」と呼ばれる)。
* [L2正則化](https://developers.google.com/machine-learning/glossary/#L2_regularization) 重み係数の二乗に比例するコストを加える(重み係数の二乗「L2ノルム」と呼ばれる)。L2正則化はニューラルネットワーク用語では重み減衰(Weight Decay)と呼ばれる。呼び方が違うので混乱しないように。重み減衰は数学的にはL2正則化と同義である。
`tf.keras`では、重みの正則化をするために、重み正則化のインスタンスをキーワード引数として層に加えます。ここでは、L2正則化を追加してみましょう。
```
l2_model = keras.models.Sequential([
keras.layers.Dense(16, kernel_regularizer=keras.regularizers.l2(0.001),
activation=tf.nn.relu, input_shape=(NUM_WORDS,)),
keras.layers.Dense(16, kernel_regularizer=keras.regularizers.l2(0.001),
activation=tf.nn.relu),
keras.layers.Dense(1, activation=tf.nn.sigmoid)
])
l2_model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy', 'binary_crossentropy'])
l2_model_history = l2_model.fit(train_data, train_labels,
epochs=20,
batch_size=512,
validation_data=(test_data, test_labels),
verbose=2)
```
```l2(0.001)```というのは、層の重み行列の係数全てに対して```0.001 * 重み係数の値 **2```をネットワークの損失値合計に加えることを意味します。このペナルティは訓練時のみに加えられるため、このネットワークの損失値は、訓練時にはテスト時に比べて大きくなることに注意してください。
L2正則化の影響を見てみましょう。
```
plot_history([('baseline', baseline_history),
('l2', l2_model_history)])
```
ご覧のように、L2正則化ありのモデルは比較基準のモデルに比べて過学習しにくくなっています。両方のモデルのパラメータ数は同じであるにもかかわらずです。
### ドロップアウトを追加する
ドロップアウトは、ニューラルネットワークの正則化テクニックとして最もよく使われる手法の一つです。この手法は、トロント大学のヒントンと彼の学生が開発したものです。ドロップアウトは層に適用するもので、訓練時に層から出力された特徴量に対してランダムに「ドロップアウト(つまりゼロ化)」を行うものです。例えば、ある層が訓練時にある入力サンプルに対して、普通は`[0.2, 0.5, 1.3, 0.8, 1.1]` というベクトルを出力するとします。ドロップアウトを適用すると、このベクトルは例えば`[0, 0.5, 1.3, 0, 1.1]`のようにランダムに散らばったいくつかのゼロを含むようになります。「ドロップアウト率」はゼロ化される特徴の割合で、通常は0.2から0.5の間に設定します。テスト時は、どのユニットもドロップアウトされず、代わりに出力値がドロップアウト率と同じ比率でスケールダウンされます。これは、訓練時に比べてたくさんのユニットがアクティブであることに対してバランスをとるためです。
`tf.keras`では、Dropout層を使ってドロップアウトをネットワークに導入できます。ドロップアウト層は、その直前の層の出力に対してドロップアウトを適用します。
それでは、IMDBネットワークに2つのドロップアウト層を追加しましょう。
```
dpt_model = keras.models.Sequential([
keras.layers.Dense(16, activation=tf.nn.relu, input_shape=(NUM_WORDS,)),
keras.layers.Dropout(rate=0.5),
keras.layers.Dense(16, activation=tf.nn.relu),
keras.layers.Dropout(rate=0.5),
keras.layers.Dense(1, activation=tf.nn.sigmoid)
])
dpt_model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy','binary_crossentropy'])
dpt_model_history = dpt_model.fit(train_data, train_labels,
epochs=20,
batch_size=512,
validation_data=(test_data, test_labels),
verbose=2)
plot_history([('baseline', baseline_history),
('dropout', dpt_model_history)])
```
ドロップアウトを追加することで、比較対象モデルより明らかに改善が見られます。
まとめ:ニューラルネットワークにおいて過学習を防ぐ最も一般的な方法は次のとおりです。
* 訓練データを増やす
* ネットワークの容量をへらす
* 重みの正則化を行う
* ドロップアウトを追加する
このガイドで触れていない2つの重要なアプローチがあります。データ拡張とバッチ正規化です。
| github_jupyter |
```
from deeplesion.ENVIRON import data_root
anchor_scales = [4, 6, 8, 12, 24, 48]#, 64
# fp16 = dict(loss_scale=96.)
dataset_transform = dict(
IMG_DO_CLIP = False,
# PIXEL_MEAN = 0.5,
WINDOWING = [-1024, 2050],
DATA_AUG_POSITION = False,
NORM_SPACING = 0.,
SLICE_INTV = 2.0,
NUM_SLICES = 7,
GROUNP_ZSAPACING = False,
)
input_channel = dataset_transform['NUM_SLICES']
feature_channel = 512
# model settings
weights2d_path = None,
model = dict(
type='MaskRCNN',
pretrained= False,
backbone=dict(
type='DenseNetCustomTrunc3dACS',
n_cts=input_channel,
out_dim=feature_channel,
fpn_finest_layer=2,
n_fold=8,
memory_efficient=True),
rpn_head=dict(
type='RPNHead',
in_channels=feature_channel,
feat_channels=feature_channel,
anchor_scales=anchor_scales,
anchor_ratios=[0.5, 1., 2.0],
anchor_base_sizes=[4],
anchor_strides=[4],
target_means=[.0, .0, .0, .0],
target_stds=[1., 1., 1., 1.],
loss_cls=dict(
type='CrossEntropyLoss', use_sigmoid=True, loss_weight=1.0),
loss_bbox=dict(type='SmoothL1Loss', beta=1.0 / 9.0, loss_weight=1.0)),
bbox_roi_extractor=dict(
type='SingleRoIExtractor',
roi_layer=dict(type='RoIAlign', out_size=7, sample_num=2),
out_channels=feature_channel,
featmap_strides=[4]),
bbox_head=dict(
type='SharedFCBBoxHead',
num_fcs=2,
in_channels=feature_channel,
fc_out_channels=feature_channel * 4,
roi_feat_size=7,
num_classes=2,
num_shared_convs=0,
target_means=[0., 0., 0., 0.],
target_stds=[0.1, 0.1, 0.2, 0.2],
# target_stds=[1., 1., 1., 1.],
reg_class_agnostic=False,
loss_cls=dict(
type='CrossEntropyLoss', use_sigmoid=False, loss_weight=1.0),
loss_bbox=dict(type='SmoothL1Loss', beta=1.0, loss_weight=1.0)),
mask_roi_extractor=dict(
type='SingleRoIExtractor',
roi_layer=dict(type='RoIAlign', out_size=14, sample_num=2),
finest_scale=196,
out_channels=feature_channel,
featmap_strides=[4]),
mask_head=dict(
type='FCNMaskHead',
num_convs=4,
in_channels=feature_channel,
conv_out_channels=feature_channel // 2,
num_classes=2,
loss_mask=dict(
type='DiceLoss', use_mask=True, loss_weight=1.0)))##DiceLoss, CrossEntropyLoss
# model training and testing settings
train_cfg = dict(
rpn=dict(
assigner=dict(
type='MaxIoUAssigner',
pos_iou_thr=0.7,
neg_iou_thr=0.3,
min_pos_iou=0.3,
ignore_iof_thr=-1),
sampler=dict(
type='RandomSampler',
num=32,
pos_fraction=0.5,
neg_pos_ub=-1,
add_gt_as_proposals=False),
allowed_border=0,
pos_weight=-1,
debug=False),
rpn_proposal=dict(
nms_across_levels=False,
nms_pre=12000,
nms_post=2000,
max_num=2000,
nms_thr=0.7,# 越大 抑制越少
min_bbox_size=0),
rcnn=dict(
assigner=dict(
type='MaxIoUAssigner',
pos_iou_thr=0.5,
neg_iou_thr=0.5,
min_pos_iou=0.5,
ignore_iof_thr=-1),
sampler=dict(
type='RandomSampler',
num=64,
pos_fraction=0.25,
neg_pos_ub=-1,
add_gt_as_proposals=True),
mask_size=28,
pos_weight=-1,
debug=False))
test_cfg = dict(
avgFP=[0.5, 1, 2, 4, 8, 16],
iou_th_astrue=0.5,
rpn=dict(
nms_across_levels=False,
nms_pre=6000,
nms_post=300,
max_num=300,
nms_thr=0.7,
min_bbox_size=0),
rcnn=dict(
score_thr=0.00005,
nms=dict(type='nms', iou_thr=0.5),
max_per_img=50,
mask_thr_binary=0.5))
# dataset settings
dataset_type = 'DeepLesionDatasetACS'
img_norm_cfg = dict(
mean=[0.] * input_channel, std=[1.] * input_channel, to_rgb=False)
pre_pipeline=[
dict(
type='Albu',
transforms=[dict(
type='ShiftScaleRotate',
shift_limit=0.04,
scale_limit=0.2,
rotate_limit=15,
interpolation=2,## cv::BORDER_REFLECT
p=0.8),],#dict(type='Rotate', limit=15, p=0.5)
bbox_params=dict(
type='BboxParams',
format='pascal_voc',
label_fields=['gt_labels'],
min_visibility=0.0),
keymap={
'img': 'image',
'gt_masks': 'masks',
'gt_bboxes': 'bboxes'
},
update_pad_shape=False,
skip_img_without_anno=False),
dict(type='Resize', img_scale=(512, 512), keep_ratio=True),
dict(type='RandomFlip', flip_ratio=0.5),
dict(type='Normalize', **img_norm_cfg),
dict(type='Pad', size_divisor=32),
]
pre_pipeline_test=[
dict(type='Resize', img_scale=(512, 512), keep_ratio=True),
dict(type='RandomFlip', flip_ratio=0),
dict(type='Normalize', **img_norm_cfg),
dict(type='Pad', size_divisor=32),
]
train_pipeline = [
dict(type='DefaultFormatBundle_3d'),
dict(type='Collect', keys=['img', 'gt_bboxes', 'gt_labels', 'gt_masks', 'thickness']),#, 'flage'#, 'img_info'#, 'z_spacing', , 'thickness'
]
test_pipeline = [
dict(type='ImageToTensor_3d', keys=['img']),
dict(type='Collect', keys=['img', 'gt_bboxes']),
]
data = dict(
imgs_per_gpu=2,
workers_per_gpu=0,
train=dict(
type=dataset_type,
ann_file=data_root + 'train_ann.pkl',
image_path=data_root + 'Images_png/',
pipeline=train_pipeline,
pre_pipeline = pre_pipeline,
dicm2png_cfg=dataset_transform),
with_mask=True,
with_label=True,
val=dict(
type=dataset_type,
ann_file=data_root + 'val_ann.pkl',
image_path=data_root + 'Images_png/',
pipeline=train_pipeline,
pre_pipeline = pre_pipeline_test,
dicm2png_cfg=dataset_transform),
test=dict(
type=dataset_type,
ann_file=data_root + 'test_ann.pkl',
image_path=data_root + 'Images_png/',
pipeline=train_pipeline,
pre_pipeline = pre_pipeline_test,
dicm2png_cfg=dataset_transform))
# optimizer
optimizer = dict(type='SGD', lr=0.02, momentum=0.9, weight_decay=0.0001)#momentum=0.9,
optimizer_config = dict(grad_clip=dict(max_norm=35, norm_type=2))#, step_inteval=2
# learning policy
lr_config = dict(
policy='step',
warmup='linear',
warmup_iters=500,
warmup_ratio=1/100,
step=[9, 16])#10,13
checkpoint_config = dict(interval=1)
# yapf:disable
log_config = dict(
interval=100,
hooks=[
dict(type='TextLoggerHook'),
dict(type='TensorboardLoggerHook')
])
# yapf:enable
#evaluation = dict(interval=1)
# runtime settings
total_epochs = 20#16
dist_params = dict(backend='nccl')
log_level = 'INFO'
work_dir = f'./work_dirs/densenet_3d_acs_r1'
load_from = None
resume_from = None
workflow = [('train', 1)]#, ('val',1)
GPU = '0,1,2,3'
description='acs'
```
| github_jupyter |
# Extractive QA to build structured data
_This notebook is part of a tutorial series on [txtai](https://github.com/neuml/txtai), an AI-powered semantic search platform._
Traditional ETL/data parsing systems establish rules to extract information of interest. Regular expressions, string parsing and similar methods define fixed rules. This works in many cases but what if you are working with unstructured data containing numerous variations? The rules can be cumbersome and hard to maintain over time.
This notebook uses machine learning and extractive question-answering (QA) to utilize the vast knowledge built into large language models. These models have been trained on extremely large datasets, learning the many variations of natural language.
# Install dependencies
Install `txtai` and all dependencies.
```
%%capture
!pip install git+https://github.com/neuml/txtai
```
# Train a QA model with few-shot learning
The code below trains a new QA model using a few examples. These examples gives the model hints on the type of questions that will be asked and the type of answers to look for. It doesn't take a lot of examples to do this as shown below.
```
import pandas as pd
from txtai.pipeline import HFTrainer, Questions, Labels
# Training data for few-shot learning
data = [
{"question": "What is the url?",
"context": "Faiss (https://github.com/facebookresearch/faiss) is a library for efficient similarity search.",
"answers": "https://github.com/facebookresearch/faiss"},
{"question": "What is the url", "context": "The last release was Wed Sept 25 2021", "answers": None},
{"question": "What is the date?", "context": "The last release was Wed Sept 25 2021", "answers": "Wed Sept 25 2021"},
{"question": "What is the date?", "context": "The order total comes to $44.33", "answers": None},
{"question": "What is the amount?", "context": "The order total comes to $44.33", "answers": "$44.33"},
{"question": "What is the amount?", "context": "The last release was Wed Sept 25 2021", "answers": None},
]
# Fine-tune QA model
trainer = HFTrainer()
model, tokenizer = trainer("distilbert-base-cased-distilled-squad", data, task="question-answering")
```
# Parse data into a structured table
The next section takes a series of rows of text and runs a set of questions against each row. The answers are then used to build a pandas DataFrame.
```
# Input data
context = ["Released on 6/03/2021",
"Release delayed until the 11th of August",
"Documentation can be found here: neuml.github.io/txtai",
"The stock price fell to three dollars",
"Great day: closing price for March 23rd is $33.11, for details - https://finance.google.com"]
# Define column queries
queries = ["What is the url?", "What is the date?", "What is the amount?"]
# Extract fields
questions = Questions(path=(model, tokenizer), gpu=True)
results = [questions([question] * len(context), context) for question in queries]
results.append(context)
# Load into DataFrame
pd.DataFrame(list(zip(*results)), columns=["URL", "Date", "Amount", "Text"])
```
# Add additional columns
This method can be combined with other models to categorize, group or otherwise derive additional columns. The code below derives an additional sentiment column.
```
# Add sentiment
labels = Labels(path="distilbert-base-uncased-finetuned-sst-2-english", dynamic=False)
labels = ["POSITIVE" if x[0][0] == 1 else "NEGATIVE" for x in labels(context)]
results.insert(len(results) - 1, labels)
# Load into DataFrame
pd.DataFrame(list(zip(*results)), columns=["URL", "Date", "Amount", "Sentiment", "Text"])
```
| github_jupyter |
```
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from tqdm.notebook import tqdm
sns.set_style("ticks")
sns.set_context("paper", 1.5, {"lines.linewidth": 2})
seed=666
epoch=20
env_name='hopper'
reward_state_action = 'state'
dataset_name = 'medium'
epoch_list = [0, 1, 2, 3, 10, 20, 30, 40]
grad_norms_epochs = []
for epoch in tqdm(epoch_list):
grad_norms = np.load(f'results/gradnorms_{epoch}_gpt2_igpt_dt_{env_name}_{dataset_name}_{seed}_{reward_state_action}.npy').swapaxes(0, 1)
model_name_label = ['GPT2', 'iGPT', 'Random Init']
colors = [(0.372, 0.537, 0.537), (0.627, 0.352, 0.470), (0.733, 0.737, 0.870)]
my_palette = sns.color_palette(colors)
sns.boxplot(data=grad_norms, palette=my_palette) # "PuBuGn_r"
plt.xticks(np.arange(3), model_name_label)
plt.ylabel('Clipped Gradient Norm')
plt.ylim(top=0.25, bottom=0)
plt.title(f'Epoch {epoch}')
plt.savefig(f'figs/gradnorms_{epoch}_gpt2_igpt_dt_{env_name}_{dataset_name}_{seed}_{reward_state_action}.pdf')
plt.show()
grad_norms_epochs.append(grad_norms)
grad_norms_mean_gpt2 = []
grad_norms_std_gpt2 = []
grad_norms_mean_igpt = []
grad_norms_std_igpt = []
grad_norms_mean_dt = []
grad_norms_std_dt = []
for epoch, grad_norms in enumerate(grad_norms_epochs[:4]):
grad_norms_mean_gpt2.append(grad_norms[0].mean())
grad_norms_std_gpt2.append(grad_norms[0].std())
grad_norms_mean_igpt.append(grad_norms[1].mean())
grad_norms_std_igpt.append(grad_norms[1].std())
grad_norms_mean_dt.append(grad_norms[2].mean())
grad_norms_std_dt.append(grad_norms[2].std())
grad_norms_mean_gpt2 = np.array(grad_norms_mean_gpt2)
grad_norms_std_gpt2 = np.array(grad_norms_std_gpt2)
grad_norms_mean_igpt = np.array(grad_norms_mean_igpt)
grad_norms_std_igpt = np.array(grad_norms_std_igpt)
grad_norms_mean_dt = np.array(grad_norms_mean_dt)
grad_norms_std_dt = np.array(grad_norms_std_dt)
plt.fill_between(np.arange(len(grad_norms_mean_gpt2)), grad_norms_mean_gpt2 - grad_norms_std_gpt2, grad_norms_mean_gpt2 + grad_norms_std_gpt2, alpha=0.2, color=(0.372, 0.537, 0.537))
plt.plot(grad_norms_mean_gpt2, color=(0.372, 0.537, 0.537), label='GPT2')
plt.scatter(np.arange(len(grad_norms_mean_gpt2)), grad_norms_mean_gpt2, color=(0.372, 0.537, 0.537))
plt.fill_between(np.arange(len(grad_norms_mean_igpt)), grad_norms_mean_igpt - grad_norms_std_igpt, grad_norms_mean_igpt + grad_norms_std_igpt, alpha=0.2, color=(0.627, 0.352, 0.470))
plt.plot(grad_norms_mean_igpt, color=(0.627, 0.352, 0.470), label='iGPT')
plt.scatter(np.arange(len(grad_norms_mean_igpt)), grad_norms_mean_igpt, color=(0.627, 0.352, 0.470))
plt.fill_between(np.arange(len(grad_norms_mean_dt)), grad_norms_mean_dt - grad_norms_std_dt, grad_norms_mean_dt + grad_norms_std_dt, alpha=0.2, color=(0.733, 0.737, 0.870))
plt.plot(grad_norms_mean_dt, color=(0.733, 0.737, 0.870), label='Random Init')
plt.scatter(np.arange(len(grad_norms_mean_dt)), grad_norms_mean_dt, color=(0.733, 0.737, 0.870))
plt.legend()
plt.tight_layout()
plt.show()
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.