code stringlengths 2.5k 150k | kind stringclasses 1 value |
|---|---|
# Data union
4 archives are presented with data from year 2017 to 2019 for each hive (names: Wurzburg and Schwartau)
- flow(nameofthehive).csv : For a date it contains the number of departures and arrivals from/to the beehive. A positive number indicates the number of arrivals and a negative number of departures. Note that this 2 values are in the data set with the same timestamp.
- humidty(nameofthehive).csv : Level of humidity through time of the beehive expressed in %
- temperature(nameofthehive).csv : Temperature of the beehive through time of the beehive in Cº
- weight(nameofthehive).csv : Weight of the beehive through time in Kg.
Resample daily information and join everything into a single dataframe.
```
# Pandas for data loading and processing
import pandas as pd
#visualization libraries
import matplotlib.pyplot as plt
import seaborn as sns
# Reading data from schwartau
df_flow_schwartau = pd.read_csv('data/flow_schwartau.csv')
df_humidity_schwartau = pd.read_csv('data/humidity_schwartau.csv')
df_temperature_schwartau = pd.read_csv('data/temperature_schwartau.csv')
df_weight_schwartau = pd.read_csv('data/weight_schwartau.csv')
# Reading data from wurzburg
df_flow_wurzburg = pd.read_csv('data/flow_wurzburg.csv')
df_humidity_wurzburg = pd.read_csv('data/humidity_wurzburg.csv')
df_temperature_wurzburg = pd.read_csv('data/temperature_wurzburg.csv')
df_weight_wurzburg = pd.read_csv('data/weight_wurzburg.csv')
# Changing data type to timestamp from schwartau
df_flow_schwartau['timestamp'] = pd.to_datetime(df_flow_schwartau['timestamp'], format='%Y-%m-%d %H:%M:%S')
df_humidity_schwartau['timestamp'] = pd.to_datetime(df_humidity_schwartau['timestamp'], format='%Y-%m-%d %H:%M:%S')
df_temperature_schwartau['timestamp'] = pd.to_datetime(df_temperature_schwartau['timestamp'], format='%Y-%m-%d %H:%M:%S')
df_weight_schwartau['timestamp'] = pd.to_datetime(df_weight_schwartau['timestamp'], format='%Y-%m-%d %H:%M:%S')
# Changing data type to timestamp from wurzburg
df_flow_wurzburg['timestamp'] = pd.to_datetime(df_flow_wurzburg['timestamp'], format='%Y-%m-%d %H:%M:%S')
df_humidity_wurzburg['timestamp'] = pd.to_datetime(df_humidity_wurzburg['timestamp'], format='%Y-%m-%d %H:%M:%S')
df_temperature_wurzburg['timestamp'] = pd.to_datetime(df_temperature_wurzburg['timestamp'], format='%Y-%m-%d %H:%M:%S')
df_weight_wurzburg['timestamp'] = pd.to_datetime(df_weight_wurzburg['timestamp'], format='%Y-%m-%d %H:%M:%S')
print(df_flow_schwartau.head(5))
print(df_humidity_schwartau.head(5))
print(df_temperature_schwartau.head(5))
print(df_weight_schwartau.head(5))
print(df_flow_wurzburg.head(5))
print(df_humidity_wurzburg.head(5))
print(df_temperature_wurzburg.head(5))
print(df_weight_wurzburg.head(5))
# Resampling data daily
flow_schwartau_daily = df_flow_schwartau.groupby(pd.Grouper(key = 'timestamp',freq='D')).sum()
humidity_schwartau_daily = df_humidity_schwartau.groupby(pd.Grouper(key = 'timestamp',freq='D')).mean()
temperature_schwartau_daily = df_temperature_schwartau.groupby(pd.Grouper(key = 'timestamp',freq='D')).mean()
weight_schwartau_daily = df_weight_schwartau.groupby(pd.Grouper(key = 'timestamp',freq='D')).mean()
# Resampling data daily
flow_wurzburg_daily = df_flow_wurzburg.groupby(pd.Grouper(key = 'timestamp',freq='D')).sum()
humidity_wurzburg_daily = df_humidity_wurzburg.groupby(pd.Grouper(key = 'timestamp',freq='D')).mean()
temperature_wurzburg_daily = df_temperature_wurzburg.groupby(pd.Grouper(key = 'timestamp',freq='D')).mean()
weight_wurzburg_daily = df_weight_wurzburg.groupby(pd.Grouper(key = 'timestamp',freq='D')).mean()
print(flow_schwartau_daily.head(5))
print(humidity_schwartau_daily.head(5))
print(temperature_schwartau_daily.head(5))
print(weight_schwartau_daily.head(5))
print(flow_wurzburg_daily.head(5))
print(humidity_wurzburg_daily.head(5))
print(temperature_wurzburg_daily.head(5))
print(weight_wurzburg_daily.head(5))
schwartau_daily=flow_schwartau_daily.merge(humidity_schwartau_daily,on='timestamp').merge(temperature_schwartau_daily,on='timestamp').merge(weight_schwartau_daily,on='timestamp')
wurzburg_daily=flow_wurzburg_daily.merge(humidity_wurzburg_daily,on='timestamp').merge(temperature_wurzburg_daily,on='timestamp').merge(weight_wurzburg_daily,on='timestamp')
schwartau_daily.head(10)
wurzburg_daily.head(10)
schwartau_daily.to_csv('data/summary/schwartau_daily.csv', index = True, header=True)
wurzburg_daily.to_csv('data/summary/wurzburg_daily.csv', index = True, header=True)
#describe our data
schwartau_daily[schwartau_daily.select_dtypes(exclude='object').columns].describe().\
style.background_gradient(axis=1,cmap=sns.light_palette('green', as_cmap=True))
#describe our data
wurzburg_daily[wurzburg_daily.select_dtypes(exclude='object').columns].describe().\
style.background_gradient(axis=1,cmap=sns.light_palette('green', as_cmap=True))
# Resampling data hourly
flow_schwartau_hourly = df_flow_schwartau.groupby(pd.Grouper(key = 'timestamp',freq='H')).sum()
humidity_schwartau_hourly = df_humidity_schwartau.groupby(pd.Grouper(key = 'timestamp',freq='H')).mean()
temperature_schwartau_hourly = df_temperature_schwartau.groupby(pd.Grouper(key = 'timestamp',freq='H')).mean()
weight_schwartau_hourly = df_weight_schwartau.groupby(pd.Grouper(key = 'timestamp',freq='H')).mean()
# Resampling data hourly
flow_wurzburg_hourly = df_flow_wurzburg.groupby(pd.Grouper(key = 'timestamp',freq='H')).sum()
humidity_wurzburg_hourly = df_humidity_wurzburg.groupby(pd.Grouper(key = 'timestamp',freq='H')).mean()
temperature_wurzburg_hourly = df_temperature_wurzburg.groupby(pd.Grouper(key = 'timestamp',freq='H')).mean()
weight_wurzburg_hourly = df_weight_wurzburg.groupby(pd.Grouper(key = 'timestamp',freq='H')).mean()
schwartau_hourly=flow_schwartau_hourly.merge(humidity_schwartau_hourly,on='timestamp').merge(temperature_schwartau_hourly,on='timestamp').merge(weight_schwartau_hourly,on='timestamp')
wurzburg_hourly=flow_wurzburg_hourly.merge(humidity_wurzburg_hourly,on='timestamp').merge(temperature_wurzburg_hourly,on='timestamp').merge(weight_wurzburg_hourly,on='timestamp')
schwartau_hourly.to_csv('data/summary/schwartau_hourly.csv', index = True, header=True)
wurzburg_hourly.to_csv('data/summary/wurzburg_hourly.csv', index = True, header=True)
#describe our data
schwartau_hourly[schwartau_hourly.select_dtypes(exclude='object').columns].describe().\
style.background_gradient(axis=1,cmap=sns.light_palette('green', as_cmap=True))
#describe our data
wurzburg_hourly[wurzburg_hourly.select_dtypes(exclude='object').columns].describe().\
style.background_gradient(axis=1,cmap=sns.light_palette('green', as_cmap=True))
```
| github_jupyter |
# Matrix Factorization for Recommender Systems - Part 2
As seen in [Part 1](/examples/matrix-factorization-for-recommender-systems-part-1), strength of [Matrix Factorization (MF)](https://en.wikipedia.org/wiki/Matrix_factorization_(recommender_systems)) lies in its ability to deal with sparse and high cardinality categorical variables. In this second tutorial we will have a look at Factorization Machines (FM) algorithm and study how it generalizes the power of MF.
**Table of contents of this tutorial series on matrix factorization for recommender systems:**
- [Part 1 - Traditional Matrix Factorization methods for Recommender Systems](https://online-ml.github.io/examples/matrix-factorization-for-recommender-systems-part-1)
- [Part 2 - Factorization Machines and Field-aware Factorization Machines](https://online-ml.github.io/examples/matrix-factorization-for-recommender-systems-part-2)
- [Part 3 - Large scale learning and better predictive power with multiple pass learning](https://online-ml.github.io/examples/matrix-factorization-for-recommender-systems-part-3)
## Factorization Machines
Steffen Rendel came up in 2010 with [Factorization Machines](https://www.csie.ntu.edu.tw/~b97053/paper/Rendle2010FM.pdf), an algorithm able to handle any real valued feature vector, combining the advantages of general predictors with factorization models. It became quite popular in the field of online advertising, notably after winning several Kaggle competitions. The modeling technique starts with a linear regression to capture the effects of each variable individually:
$$
\normalsize
\hat{y}(x) = w_{0} + \sum_{j=1}^{p} w_{j} x_{j}
$$
Then are added interaction terms to learn features relations. Instead of learning a single and specific weight per interaction (as in [polynomial regression](https://en.wikipedia.org/wiki/Polynomial_regression)), a set of latent factors is learnt per feature (as in MF). An interaction is calculated by multiplying involved features product with their latent vectors dot product. The degree of factorization — or model order — represents the maximum number of features per interaction considered. The model equation for a factorization machine of degree $d$ = 2 is defined as:
$$
\normalsize
\hat{y}(x) = w_{0} + \sum_{j=1}^{p} w_{j} x_{j} + \sum_{j=1}^{p} \sum_{j'=j+1}^{p} \langle \mathbf{v}_j, \mathbf{v}_{j'} \rangle x_{j} x_{j'}
$$
Where $\normalsize \langle \mathbf{v}_j, \mathbf{v}_{j'} \rangle$ is the dot product of $j$ and $j'$ latent vectors:
$$
\normalsize
\langle \mathbf{v}_j, \mathbf{v}_{j'} \rangle = \sum_{f=1}^{k} \mathbf{v}_{j, f} \cdot \mathbf{v}_{j', f}
$$
Higher-order FM will be covered in a following section, just note that factorization models express their power in sparse settings, which is also where higher-order interactions are hard to estimate.
Strong emphasis must be placed on feature engineering as it allows FM to mimic most factorization models and significantly impact its performance. High cardinality categorical variables one hot encoding is the most frequent step before feeding the model with data. For more efficiency, `river` FM implementation considers string values as categorical variables and automatically one hot encode them. FM models have their own module [river.facto](/api/overview/#facto).
## Mimic Biased Matrix Factorization (BiasedMF)
Let's start with a simple example where we want to reproduce the Biased Matrix Factorization model we trained in the previous tutorial. For a fair comparison with [Part 1 example](/examples/matrix-factorization-for-recommender-systems-part-1/#biased-matrix-factorization-biasedmf), let's set the same evaluation framework:
```
from river import datasets
from river import metrics
from river.evaluate import progressive_val_score
def evaluate(model):
X_y = datasets.MovieLens100K()
metric = metrics.MAE() + metrics.RMSE()
_ = progressive_val_score(X_y, model, metric, print_every=25_000, show_time=True, show_memory=True)
```
In order to build an equivalent model we need to use the same hyper-parameters. As we can't replace FM intercept by the global running mean we won't be able to build the exact same model:
```
from river import compose
from river import facto
from river import meta
from river import optim
from river import stats
fm_params = {
'n_factors': 10,
'weight_optimizer': optim.SGD(0.025),
'latent_optimizer': optim.SGD(0.05),
'sample_normalization': False,
'l1_weight': 0.,
'l2_weight': 0.,
'l1_latent': 0.,
'l2_latent': 0.,
'intercept': 3,
'intercept_lr': .01,
'weight_initializer': optim.initializers.Zeros(),
'latent_initializer': optim.initializers.Normal(mu=0., sigma=0.1, seed=73),
}
regressor = compose.Select('user', 'item')
regressor |= facto.FMRegressor(**fm_params)
model = meta.PredClipper(
regressor=regressor,
y_min=1,
y_max=5
)
evaluate(model)
```
Both MAE are very close to each other (0.7486 vs 0.7485) showing that we almost reproduced [reco.BiasedMF](/api/reco/BiasedMF/) algorithm. The cost is a naturally slower running time as FM implementation offers more flexibility.
## Feature engineering for FM models
Let's study the basics of how to properly encode data for FM models. We are going to keep using MovieLens 100K as it provides various feature types:
```
import json
for x, y in datasets.MovieLens100K():
print(f'x = {json.dumps(x, indent=4)}\ny = {y}')
break
```
The features we are going to add to our model don't improve its predictive power. Nevertheless, they are useful to illustrate different methods of data encoding:
1. Set-categorical variables
We have seen that categorical variables are one hot encoded automatically if set to strings, in the other hand, set-categorical variables must be encoded explicitly by the user. A good way of doing so is to assign them a value of $1/m$, where $m$ is the number of elements of the sample set. It gives the feature a constant "weight" across all samples preserving model's stability. Let's create a routine to encode movies genres this way:
```
def split_genres(x):
genres = x['genres'].split(', ')
return {f'genre_{genre}': 1 / len(genres) for genre in genres}
```
2. Numerical variables
In practice, transforming numerical features into categorical ones works better in most cases. Feature binning is the natural way, but finding good bins is sometimes more an art than a science. Let's encode users age with something simple:
```
def bin_age(x):
if x['age'] <= 18:
return {'age_0-18': 1}
elif x['age'] <= 32:
return {'age_19-32': 1}
elif x['age'] < 55:
return {'age_33-54': 1}
else:
return {'age_55-100': 1}
```
Let's put everything together:
```
fm_params = {
'n_factors': 14,
'weight_optimizer': optim.SGD(0.01),
'latent_optimizer': optim.SGD(0.025),
'intercept': 3,
'latent_initializer': optim.initializers.Normal(mu=0., sigma=0.05, seed=73),
}
regressor = compose.Select('user', 'item')
regressor += (
compose.Select('genres') |
compose.FuncTransformer(split_genres)
)
regressor += (
compose.Select('age') |
compose.FuncTransformer(bin_age)
)
regressor |= facto.FMRegressor(**fm_params)
model = meta.PredClipper(
regressor=regressor,
y_min=1,
y_max=5
)
evaluate(model)
```
Note that using more variables involves factorizing a larger latent space, then increasing the number of latent factors $k$ often helps capturing more information.
Some other feature engineering tips from [3 idiots' winning solution](https://www.kaggle.com/c/criteo-display-ad-challenge/discussion/10555) for Kaggle [Criteo display ads](https://www.kaggle.com/c/criteo-display-ad-challenge) competition in 2014:
- Infrequent modalities often bring noise and little information, transforming them into a special tag can help
- In some cases, sample-wise normalization seems to make the optimization problem easier to be solved
## Higher-Order Factorization Machines (HOFM)
The model equation generalized to any order $d \geq 2$ is defined as:
$$
\normalsize
\hat{y}(x) = w_{0} + \sum_{j=1}^{p} w_{j} x_{j} + \sum_{l=2}^{d} \sum_{j_1=1}^{p} \cdots \sum_{j_l=j_{l-1}+1}^{p} \left(\prod_{j'=1}^{l} x_{j_{j'}} \right) \left(\sum_{f=1}^{k_l} \prod_{j'=1}^{l} v_{j_{j'}, f}^{(l)} \right)
$$
```
hofm_params = {
'degree': 3,
'n_factors': 12,
'weight_optimizer': optim.SGD(0.01),
'latent_optimizer': optim.SGD(0.025),
'intercept': 3,
'latent_initializer': optim.initializers.Normal(mu=0., sigma=0.05, seed=73),
}
regressor = compose.Select('user', 'item')
regressor += (
compose.Select('genres') |
compose.FuncTransformer(split_genres)
)
regressor += (
compose.Select('age') |
compose.FuncTransformer(bin_age)
)
regressor |= facto.HOFMRegressor(**hofm_params)
model = meta.PredClipper(
regressor=regressor,
y_min=1,
y_max=5
)
evaluate(model)
```
As said previously, high-order interactions are often hard to estimate due to too much sparsity, that's why we won't spend too much time here.
## Field-aware Factorization Machines (FFM)
[Field-aware variant of FM (FFM)](https://www.csie.ntu.edu.tw/~cjlin/papers/ffm.pdf) improved the original method by adding the notion of "*fields*". A "*field*" is a group of features that belong to a specific domain (e.g. the "*users*" field, the "*items*" field, or the "*movie genres*" field).
FFM restricts itself to pairwise interactions and factorizes separated latent spaces — one per combination of fields (e.g. users/items, users/movie genres, or items/movie genres) — instead of a common one shared by all fields. Therefore, each feature has one latent vector per field it can interact with — so that it can learn the specific effect with each different field.
The model equation is defined by:
$$
\normalsize
\hat{y}(x) = w_{0} + \sum_{j=1}^{p} w_{j} x_{j} + \sum_{j=1}^{p} \sum_{j'=j+1}^{p} \langle \mathbf{v}_{j, f_{j'}}, \mathbf{v}_{j', f_{j}} \rangle x_{j} x_{j'}
$$
Where $f_j$ and $f_{j'}$ are the fields corresponding to $j$ and $j'$ features, respectively.
```
ffm_params = {
'n_factors': 8,
'weight_optimizer': optim.SGD(0.01),
'latent_optimizer': optim.SGD(0.025),
'intercept': 3,
'latent_initializer': optim.initializers.Normal(mu=0., sigma=0.05, seed=73),
}
regressor = compose.Select('user', 'item')
regressor += (
compose.Select('genres') |
compose.FuncTransformer(split_genres)
)
regressor += (
compose.Select('age') |
compose.FuncTransformer(bin_age)
)
regressor |= facto.FFMRegressor(**ffm_params)
model = meta.PredClipper(
regressor=regressor,
y_min=1,
y_max=5
)
evaluate(model)
```
Note that FFM usually needs to learn smaller number of latent factors $k$ than FM as each latent vector only deals with one field.
## Field-weighted Factorization Machines (FwFM)
[Field-weighted Factorization Machines (FwFM)](https://arxiv.org/abs/1806.03514) address FFM memory issues caused by its large number of parameters, which is in the order of *feature number* times *field number*. As FFM, FwFM is an extension of FM restricted to pairwise interactions, but instead of factorizing separated latent spaces, it learns a specific weight $r_{f_j, f_{j'}}$ for each field combination modelling the interaction strength.
The model equation is defined as:
$$
\normalsize
\hat{y}(x) = w_{0} + \sum_{j=1}^{p} w_{j} x_{j} + \sum_{j=1}^{p} \sum_{j'=j+1}^{p} r_{f_j, f_{j'}} \langle \mathbf{v}_j, \mathbf{v}_{j'} \rangle x_{j} x_{j'}
$$
```
fwfm_params = {
'n_factors': 10,
'weight_optimizer': optim.SGD(0.01),
'latent_optimizer': optim.SGD(0.025),
'intercept': 3,
'seed': 73,
}
regressor = compose.Select('user', 'item')
regressor += (
compose.Select('genres') |
compose.FuncTransformer(split_genres)
)
regressor += (
compose.Select('age') |
compose.FuncTransformer(bin_age)
)
regressor |= facto.FwFMRegressor(**fwfm_params)
model = meta.PredClipper(
regressor=regressor,
y_min=1,
y_max=5
)
evaluate(model)
```
| github_jupyter |
# Multi-Head Attention
:label:`sec_multihead-attention`
In practice,
given the same set of queries, keys, and values
we may want our model to
combine knowledge from
different behaviors of the same attention mechanism,
such as capturing dependencies of various ranges (e.g., shorter-range vs. longer-range)
within a sequence.
Thus,
it may be beneficial
to allow our attention mechanism
to jointly use different representation subspaces
of queries, keys, and values.
To this end,
instead of performing a single attention pooling,
queries, keys, and values
can be transformed
with $h$ independently learned linear projections.
Then these $h$ projected queries, keys, and values
are fed into attention pooling in parallel.
In the end,
$h$ attention pooling outputs
are concatenated and
transformed with another learned linear projection
to produce the final output.
This design
is called *multi-head attention*,
where each of the $h$ attention pooling outputs
is a *head* :cite:`Vaswani.Shazeer.Parmar.ea.2017`.
Using fully-connected layers
to perform learnable linear transformations,
:numref:`fig_multi-head-attention`
describes multi-head attention.

:label:`fig_multi-head-attention`
## Model
Before providing the implementation of multi-head attention,
let us formalize this model mathematically.
Given a query $\mathbf{q} \in \mathbb{R}^{d_q}$,
a key $\mathbf{k} \in \mathbb{R}^{d_k}$,
and a value $\mathbf{v} \in \mathbb{R}^{d_v}$,
each attention head $\mathbf{h}_i$ ($i = 1, \ldots, h$)
is computed as
$$\mathbf{h}_i = f(\mathbf W_i^{(q)}\mathbf q, \mathbf W_i^{(k)}\mathbf k,\mathbf W_i^{(v)}\mathbf v) \in \mathbb R^{p_v},$$
where learnable parameters
$\mathbf W_i^{(q)}\in\mathbb R^{p_q\times d_q}$,
$\mathbf W_i^{(k)}\in\mathbb R^{p_k\times d_k}$
and $\mathbf W_i^{(v)}\in\mathbb R^{p_v\times d_v}$,
and
$f$ is attention pooling,
such as
additive attention and scaled dot-product attention
in :numref:`sec_attention-scoring-functions`.
The multi-head attention output
is another linear transformation via
learnable parameters
$\mathbf W_o\in\mathbb R^{p_o\times h p_v}$
of the concatenation of $h$ heads:
$$\mathbf W_o \begin{bmatrix}\mathbf h_1\\\vdots\\\mathbf h_h\end{bmatrix} \in \mathbb{R}^{p_o}.$$
Based on this design,
each head may attend to different parts of the input.
More sophisticated functions than the simple weighted average
can be expressed.
```
import math
from mxnet import autograd, np, npx
from mxnet.gluon import nn
from d2l import mxnet as d2l
npx.set_np()
```
## Implementation
In our implementation,
we choose the scaled dot-product attention
for each head of the multi-head attention.
To avoid significant growth
of computational cost and parameterization cost,
we set
$p_q = p_k = p_v = p_o / h$.
Note that $h$ heads
can be computed in parallel
if we set
the number of outputs of linear transformations
for the query, key, and value
to $p_q h = p_k h = p_v h = p_o$.
In the following implementation,
$p_o$ is specified via the argument `num_hiddens`.
```
#@save
class MultiHeadAttention(nn.Block):
def __init__(self, num_hiddens, num_heads, dropout, use_bias=False,
**kwargs):
super(MultiHeadAttention, self).__init__(**kwargs)
self.num_heads = num_heads
self.attention = d2l.DotProductAttention(dropout)
self.W_q = nn.Dense(num_hiddens, use_bias=use_bias, flatten=False)
self.W_k = nn.Dense(num_hiddens, use_bias=use_bias, flatten=False)
self.W_v = nn.Dense(num_hiddens, use_bias=use_bias, flatten=False)
self.W_o = nn.Dense(num_hiddens, use_bias=use_bias, flatten=False)
def forward(self, queries, keys, values, valid_lens):
# Shape of `queries`, `keys`, or `values`:
# (`batch_size`, no. of queries or key-value pairs, `num_hiddens`)
# Shape of `valid_lens`:
# (`batch_size`,) or (`batch_size`, no. of queries)
# After transposing, shape of output `queries`, `keys`, or `values`:
# (`batch_size` * `num_heads`, no. of queries or key-value pairs,
# `num_hiddens` / `num_heads`)
queries = transpose_qkv(self.W_q(queries), self.num_heads)
keys = transpose_qkv(self.W_k(keys), self.num_heads)
values = transpose_qkv(self.W_v(values), self.num_heads)
if valid_lens is not None:
# On axis 0, copy the first item (scalar or vector) for
# `num_heads` times, then copy the next item, and so on
valid_lens = valid_lens.repeat(self.num_heads, axis=0)
# Shape of `output`: (`batch_size` * `num_heads`, no. of queries,
# `num_hiddens` / `num_heads`)
output = self.attention(queries, keys, values, valid_lens)
# Shape of `output_concat`:
# (`batch_size`, no. of queries, `num_hiddens`)
output_concat = transpose_output(output, self.num_heads)
return self.W_o(output_concat)
```
To allow for parallel computation of multiple heads,
the above `MultiHeadAttention` class uses two transposition functions as defined below.
Specifically,
the `transpose_output` function reverses the operation
of the `transpose_qkv` function.
```
#@save
def transpose_qkv(X, num_heads):
# Shape of input `X`:
# (`batch_size`, no. of queries or key-value pairs, `num_hiddens`).
# Shape of output `X`:
# (`batch_size`, no. of queries or key-value pairs, `num_heads`,
# `num_hiddens` / `num_heads`)
X = X.reshape(X.shape[0], X.shape[1], num_heads, -1)
# Shape of output `X`:
# (`batch_size`, `num_heads`, no. of queries or key-value pairs,
# `num_hiddens` / `num_heads`)
X = X.transpose(0, 2, 1, 3)
# Shape of `output`:
# (`batch_size` * `num_heads`, no. of queries or key-value pairs,
# `num_hiddens` / `num_heads`)
return X.reshape(-1, X.shape[2], X.shape[3])
#@save
def transpose_output(X, num_heads):
"""Reverse the operation of `transpose_qkv`"""
X = X.reshape(-1, num_heads, X.shape[1], X.shape[2])
X = X.transpose(0, 2, 1, 3)
return X.reshape(X.shape[0], X.shape[1], -1)
```
Let us test our implemented `MultiHeadAttention` class
using a toy example where keys and values are the same.
As a result,
the shape of the multi-head attention output
is (`batch_size`, `num_queries`, `num_hiddens`).
```
num_hiddens, num_heads = 100, 5
attention = MultiHeadAttention(num_hiddens, num_heads, 0.5)
attention.initialize()
batch_size, num_queries, num_kvpairs, valid_lens = 2, 4, 6, np.array([3, 2])
X = np.ones((batch_size, num_queries, num_hiddens))
Y = np.ones((batch_size, num_kvpairs, num_hiddens))
attention(X, Y, Y, valid_lens).shape
```
## Summary
* Multi-head attention combines knowledge of the same attention pooling via different representation subspaces of queries, keys, and values.
* To compute multiple heads of multi-head attention in parallel, proper tensor manipulation is needed.
## Exercises
1. Visualize attention weights of multiple heads in this experiment.
1. Suppose that we have a trained model based on multi-head attention and we want to prune least important attention heads to increase the prediction speed. How can we design experiments to measure the importance of an attention head?
[Discussions](https://discuss.d2l.ai/t/1634)
| github_jupyter |
<a href="https://colab.research.google.com/github/Dariush-Mehdiaraghi/bachelor_project/blob/main/ssdlite_mobiledet_transfer_learning.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
**Retrain SSDLite Mobiledet for Coral's EdgeTpu**
This is a slightly modified version of the [notebook](https://github.com/Namburger/edgetpu-ssdlite-mobiledet-retrain) by [Nam Vu ](https://github.com/Namburger)
```
# Import tensorflow 1.x and install tf_slim.
%tensorflow_version 1.x
!pip install tf_slim
!pip show tensorflow
# Install protobuf-compiler and the tensorflow's object detection API.
!apt-get install protobuf-compiler
!git clone https://github.com/tensorflow/models.git
import os
os.environ['PYTHONPATH'] += ':/content/models/research/'
os.environ['PYTHONPATH'] += ':/content/models/research/slim/'
os.environ['PYTHONPATH'] += ':/content/models/research/object_detection/utils/'
os.environ['PYTHONPATH'] += ':/content/models/research/object_detection'
%cd models/research
# Compile all the protobuf dependencies.
!protoc object_detection/protos/*.proto --python_out=.
# Set up and install the object detection API.
!cp object_detection/packages/tf1/setup.py .
!python -m pip install .
# Run a test to make sure setup is correct.
!python object_detection/builders/model_builder_test.py
# Now let's download our training dataset.
#%rm -r /content/dataset
%cd /content
%rm -r /content/dataset
%mkdir /content/dataset
%cd /content/dataset
!curl -L "https://app.roboflow.com/ds/HaWC1lznR0?key=pJugxNUTU4" > roboflow.zip; unzip roboflow.zip; rm roboflow.zip
# IF YOU DONT HAVE TFRECORD FILES Now we can create the tfrecord files.
%cd /content/models/research
!cp object_detection/data/pet_label_map.pbtxt /content/dataset
!python3 object_detection/dataset_tools/create_pet_tf_record.py \
--label_map_path="/content/dataset/pet_label_map.pbtxt" \
--data_dir="/content/dataset" \
--output_dir="/content/dataset"
# Now let's download our ssdlite mobiledet pretrained model from tensorflow's model zoo.
!mkdir /content/pretrained_model
%cd /content/pretrained_model
!wget http://download.tensorflow.org/models/object_detection/ssdlite_mobiledet_edgetpu_320x320_coco_2020_05_19.tar.gz
!tar xvf ssdlite_mobiledet_edgetpu_320x320_coco_2020_05_19.tar.gz
#to save checkpoints to google drive
from google.colab import drive
drive.mount('/content/gdrive')
gdrive_ckpt_dir = "/content/gdrive/MyDrive/colabCheckpoints4SecondRun"
def get_num_classes(pbtxt_fname):
from object_detection.utils import label_map_util
label_map = label_map_util.load_labelmap(pbtxt_fname)
categories = label_map_util.convert_label_map_to_categories(
label_map, max_num_classes=90, use_display_name=True)
category_index = label_map_util.create_category_index(categories)
print(categories)
return len(category_index.keys())
get_num_classes('/content/dataset/train/bottles_label_map.pbtxt')
# Edit Pipeline config to load in our new tfrecord that we just created and add quantization aware training.
import tensorflow as tf
from google.protobuf import text_format
from object_detection.protos import pipeline_pb2
# Hack to find out if you have colab pro or not :)
gpu_info = !nvidia-smi
gpu_info = '\n'.join(gpu_info)
print(gpu_info)
gpu_name = !nvidia-smi --query-gpu=gpu_name --format=csv
# You get Tesla T4 with free colab and Tesla P100-PCIe with colab pro.
colab_pro = False if 'T4' in gpu_name else True
pipeline = pipeline_pb2.TrainEvalPipelineConfig()
config_path = '/content/models/research/object_detection/samples/configs/ssdlite_mobiledet_edgetpu_320x320_coco_sync_4x4.config'
with tf.gfile.GFile(config_path, "r") as f:
proto_str = f.read()
text_format.Merge(proto_str, pipeline)
pipeline.train_input_reader.tf_record_input_reader.input_path[:] = ['/content/dataset/train/bottles.tfrecord']
pipeline.train_input_reader.label_map_path = '/content/dataset/train/bottles_label_map.pbtxt'
pipeline.eval_input_reader[0].tf_record_input_reader.input_path[:] = ['/content/dataset/valid/bottles.tfrecord']
pipeline.eval_input_reader[0].label_map_path = '/content/dataset/valid/bottles_label_map.pbtxt'
#pipeline.train_config.fine_tune_checkpoint = '/content/pretrained_model/ssdlite_mobiledet_edgetpu_320x320_coco_2020_05_19/fp32/model.ckpt'
pipeline.train_config.fine_tune_checkpoint = '/content/gdrive/MyDrive/colabCheckpoints3/model.ckpt-25000'
pipeline.train_config.batch_size = 64 if colab_pro else 32 # Smaller batch size on free gpu to avoid OOM Killer normaly 64
pipeline.train_config.num_steps = 25000 if colab_pro else 10000 # Less steps with free gpu but 10k should be good enough
#CHANGE NUM OF CLASSES
pipeline.model.ssd.num_classes = get_num_classes('/content/dataset/train/bottles_label_map.pbtxt')
# Enable ssdlite, this should already be enabled in the config we downloaded, but this is just to make sure.
pipeline.model.ssd.box_predictor.convolutional_box_predictor.kernel_size = 3
pipeline.model.ssd.box_predictor.convolutional_box_predictor.use_depthwise = True
pipeline.model.ssd.feature_extractor.use_depthwise = True
# Quantization Aware Training
pipeline.graph_rewriter.quantization.delay = 0
pipeline.graph_rewriter.quantization.weight_bits = 8
pipeline.graph_rewriter.quantization.activation_bits = 8
config_text = text_format.MessageToString(pipeline)
with tf.gfile.Open(config_path, "wb") as f:
f.write(config_text)
# This is out config after modifying.
!cat /content/models/research/object_detection/samples/configs/ssdlite_mobiledet_edgetpu_320x320_coco_sync_4x4.config
# Before we start training, let's start tensorboard so we can track the progress.
# More info on tensorflow can be found here: https://www.tensorflow.org/tutorials
%cd /content
!wget https://bin.equinox.io/c/4VmDzA7iaHb/ngrok-stable-linux-amd64.zip
!unzip -o ngrok-stable-linux-amd64.zip
# Starts tensorboard, so we can monitor the training process.
get_ipython().system_raw(
'tensorboard --logdir {} --host 0.0.0.0 --port 6006 &'
.format(gdrive_ckpt_dir + '/eval_0')
)
get_ipython().system_raw('./ngrok http 6006 &')
print('Click on link below to track progress:')
import time
time.sleep(1)
!curl -s http://localhost:4040/api/tunnels | python3 -c \
"import sys, json; print(json.load(sys.stdin)['tunnels'][0]['public_url'])"
# Let's begin training, expects to take a few hours, time for a good stretch :)
%cd /content/models/research/
!python3 object_detection/model_main.py \
--logtostderr=true \
--model_dir={gdrive_ckpt_dir} \
--pipeline_config_path=/content/models/research/object_detection/samples/configs/ssdlite_mobiledet_edgetpu_320x320_coco_sync_4x4.config
# Make inference graph.
best_ckpt_path = gdrive_ckpt_dir + "/model.ckpt-24082"# Make sure to change this checkpoint to the corresponding num step you set from above.
print("best checkpoint is:", best_ckpt_path)
!python3 /content/models/research/object_detection/export_inference_graph.py \
--input_type=image_tensor \
--pipeline_config_path=/content/models/research/object_detection/samples/configs/ssdlite_mobiledet_edgetpu_320x320_coco_sync_4x4.config \
--output_directory=/content/inference_graph \
--trained_checkpoint_prefix={best_ckpt_path}
# Let's download some test data from flickr.
!mkdir /content/test
!cd /content/test
#!wget https://live.staticflickr.com/7921/46683787864_86c9501c24_c_d.jpg -O /content/test/image1.jpg
#!wget https://live.staticflickr.com/4/8451898_8bedb2ae53_c_d.jpg -O /content/test/image2.jpg
# Do a Quick Evaluation on the inference graph model.
import numpy as np
import os
import sys
import tensorflow as tf
from collections import defaultdict
from matplotlib import pyplot as plt
from PIL import Image
from object_detection.utils import ops as utils_ops
from object_detection.utils import label_map_util
from object_detection.utils import visualization_utils as vis_util
%matplotlib inline
# Initialize tf.Graph()
detection_graph = tf.Graph()
with detection_graph.as_default():
od_graph_def = tf.GraphDef()
with tf.gfile.GFile('/content/inference_graph/frozen_inference_graph.pb', 'rb') as fid:
serialized_graph = fid.read()
od_graph_def.ParseFromString(serialized_graph)
tf.import_graph_def(od_graph_def, name='')
# Loads labels
label_map = label_map_util.load_labelmap('/content/dataset/train/bottles_label_map.pbtxt')
categories = label_map_util.convert_label_map_to_categories(label_map, max_num_classes=8, use_display_name=True)
category_index = label_map_util.create_category_index(categories)
# Run Inference and populates results in a dict.
def run_inference(graph, image):
with graph.as_default():
with tf.Session() as sess:
ops = tf.get_default_graph().get_operations()
all_tensor_names = [output.name for op in ops for output in op.outputs]
tensor_dict = {}
tensor_keys = ['num_detections', 'detection_boxes', 'detection_scores', 'detection_classes']
for key in tensor_keys:
tensor_name = key + ':0'
if tensor_name in all_tensor_names:
tensor_dict[key] = tf.get_default_graph().get_tensor_by_name(tensor_name)
# Actual inference.
image_tensor = tf.get_default_graph().get_tensor_by_name('image_tensor:0')
output_dict = sess.run(tensor_dict, feed_dict={image_tensor: np.expand_dims(image, 0)})
output_dict['num_detections'] = int(output_dict['num_detections'][0])
output_dict['detection_classes'] = output_dict['detection_classes'][0].astype(np.uint8)
output_dict['detection_boxes'] = output_dict['detection_boxes'][0]
output_dict['detection_scores'] = output_dict['detection_scores'][0]
return output_dict
test_image_path = [os.path.join('/content/test', 'image{}.jpg'.format(i)) for i in range(1, 6)]
for image_path in test_image_path:
print('Evaluating:', image_path)
image = Image.open(image_path)
img_width, img_height = image.size
image_np = np.array(image.getdata()).reshape((img_height, img_width, 3)).astype(np.uint8)
# Run inference.
output_dict = run_inference(detection_graph, image_np)
# Visualization of the results of a detection.
vis_util.visualize_boxes_and_labels_on_image_array(
image_np,
output_dict['detection_boxes'],
output_dict['detection_classes'],
output_dict['detection_scores'],
category_index,
use_normalized_coordinates=True,
line_thickness=8)
plt.figure(figsize=(48, 32))
plt.imshow(image_np)
# Now we export this model to tflite_graph format.
%cd /content/models/research
!mkdir /content/output_model
!python3 object_detection/export_tflite_ssd_graph.py \
--pipeline_config_path=/content/models/research/object_detection/samples/configs/ssdlite_mobiledet_edgetpu_320x320_coco_sync_4x4.config \
--trained_checkpoint_prefix={best_ckpt_path} \
--output_directory=/content/output_model \
--add_postprocessing_op=true
# Make sure to change the model-ckpt-# to match the checkpoint number you used.
# Now we can convert this custom trained model to a CPU tflite model
!tflite_convert \
--output_file="/content/output_model/ssdlite_mobiledet_own.tflite" \
--graph_def_file="/content/output_model/tflite_graph.pb" \
--inference_type=QUANTIZED_UINT8 \
--input_arrays="normalized_input_image_tensor" \
--output_arrays="TFLite_Detection_PostProcess,TFLite_Detection_PostProcess:1,TFLite_Detection_PostProcess:2,TFLite_Detection_PostProcess:3" \
--mean_values=128 \
--std_dev_values=128 \
--input_shapes=1,320,320,3 \
--allow_custom_ops
# Install tflite_runtime package to evaluate the model.
!pip3 install https://github.com/google-coral/pycoral/releases/download/release-frogfish/tflite_runtime-2.5.0-cp36-cp36m-linux_x86_64.whl
# Now we do evaluation on the tflite model.
import os
import numpy as np
from tflite_runtime.interpreter import Interpreter
from tflite_runtime.interpreter import load_delegate
from PIL import Image
from PIL import ImageDraw
%matplotlib inline
# Creates tflite interpreter
interpreter = Interpreter('/content/output_model/ssdlite_mobiledet_own.tflite')
# This exact code can be used to run inference on the edgetpu by simply creating
# the instantialize the interpreter with libedgetpu delegates:
# interpreter = Interpreter(args.model, experimental_delegates=[load_delegate('libedgetpu.so.1.0')])
interpreter.allocate_tensors()
interpreter.invoke() # warmup
input_details = interpreter.get_input_details()
output_details = interpreter.get_output_details()
width = input_details[0]['shape'][2]
height = input_details[0]['shape'][1]
def run_inference(interpreter, image):
interpreter.set_tensor(input_details[0]['index'], image)
interpreter.invoke()
boxes = interpreter.get_tensor(output_details[0]['index'])[0]
classes = interpreter.get_tensor(output_details[1]['index'])[0]
scores = interpreter.get_tensor(output_details[2]['index'])[0]
# num_detections = interpreter.get_tensor(output_details[3]['index'])[0]
return boxes, classes, scores
test_image_paths = [os.path.join('/content/test', 'image{}.jpg'.format(i)) for i in range(1, 6)]
for image_path in test_image_paths:
print('Evaluating:', image_path)
image = Image.open(image_path)
image_width, image_height = image.size
draw = ImageDraw.Draw(image)
resized_image = image.resize((width, height))
np_image = np.asarray(resized_image)
input_tensor = np.expand_dims(np_image, axis=0)
# Run inference
boxes, classes, scores = run_inference(interpreter, input_tensor)
# Draw results on image
colors = {0:(128, 255, 102), 1:(102, 255, 255)}
labels = {0:'abyssian cat', 1:'american bulldog'}
for i in range(len(boxes)):
if scores[i] > .7:
ymin = int(max(1, (boxes[i][0] * image_height)))
xmin = int(max(1, (boxes[i][1] * image_width)))
ymax = int(min(image_height, (boxes[i][2] * image_height)))
xmax = int(min(image_width, (boxes[i][3] * image_width)))
draw.rectangle((xmin, ymin, xmax, ymax), width=7, outline=colors[int(classes[i])])
draw.rectangle((xmin, ymin, xmax, ymin-10), fill=colors[int(classes[i])])
text = labels[int(classes[i])] + ' ' + str(scores[i]*100) + '%'
draw.text((xmin+2, ymin-10), text, fill=(0,0,0), width=2)
display(image)
# Install the edgetpu compiler.
!curl https://packages.cloud.google.com/apt/doc/apt-key.gpg | apt-key add -
!echo "deb https://packages.cloud.google.com/apt coral-edgetpu-stable main" | sudo tee /etc/apt/sources.list.d/coral-edgetpu.list
!sudo apt-get update
!sudo apt-get install edgetpu-compiler
output_model_dir = "/content/output_model/"
# Compile our model and make a tarball of the finished trained model.
%cd {output_model_dir}
!edgetpu_compiler -s ssdlite_mobiledet_own.tflite
#%cd /content/
# Copy the checkpoints, inference graph, pipeline config, and the tflite models.
#!cp -r /content/gdrive/MyDrive/colabCheckpoints2* {output_model_dir}
!cp -r /content/inference_graph/* {output_model_dir}
!tar cvf ssdlite_mobiledet_own.tar.gz {output_model_dir}
!cp ssdlite_mobiledet_own.tar.gz {gdrive_ckpt_dir}
!pip uninstall tensorflow-datasets
!pip install tensorflow-datasets==1.0.1
!pip install tflite-model-maker
from tflite_model_maker import image_classifier
from tflite_model_maker.image_classifier import DataLoader
data = DataLoader.from_folder('content/dataset')
train_data, test_data = data.split(0.8)
model = image_classifier.create(train_data)
model.export('image_classifier.tflite', 'imageLabels.txt')
# Download model and you're done!
from google.colab import files
files.download('/content/ssdlite_mobiledet_dog_vs_cat.tar.gz')
```
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
import matplotlib.pyplot as plt
%matplotlib inline
from ipywidgets import interact, interactive, fixed, interact_manual
import ipywidgets as widgets
import numpy as np
import torch, torch.optim
import torch.nn.functional as F
torch.backends.cudnn.enabled = True
torch.backends.cudnn.benchmark =True
dtype = torch.cuda.FloatTensor
import os, sys
sys.path.append('utils/*')
os.environ["CUDA_DEVICE_ORDER"] = "PCI_BUS_ID"
os.environ['CUDA_VISIBLE_DEVICES'] = '1'
import models as md
import utils.common_utils as cu
import utils.diffuser_utils as df
import utils.utils_hyperspectral as helper
```
# Single-shot Imaging Demo
Load in the PSF, 2D measurement and rolling shutter mask.
```
simulated = True # True: Use a simulated measurement or False: use an experimental measurement
downsampling_factor = 2
meas_np, mask_np, psf_np, gt_np = helper.load_data(simulated = simulated)
plt.figure(figsize=(20,10))
plt.subplot(1,3,1);plt.title('PSF');plt.imshow(psf_np)
plt.subplot(1,3,2);plt.title('Measurement');plt.imshow(meas_np)
plt.subplot(1,3,3);plt.title('Rolling shutter mask');plt.imshow(mask_np[:,:,20])
```
Initialize the lensless forward model
```
DIMS0 = meas_np.shape[0] # Image Dimensions
DIMS1 = meas_np.shape[1] # Image Dimensions
py = int((DIMS0)//2) # Pad size
px = int((DIMS1)//2) # Pad size
def pad(x):
if len(x.shape) == 2:
out = np.pad(x, ([py, py], [px,px]), mode = 'constant')
elif len(x.shape) == 3:
out = np.pad(x, ([py, py], [px,px], [0, 0]), mode = 'constant')
elif len(x.shape) == 4:
out = np.pad(x, ([py, py], [px,px], [0, 0], [0, 0]), mode = 'constant')
return out
#meas_np = pad(meas_np)
psf_pad = pad(psf_np)
h_full = np.fft.fft2(np.fft.ifftshift(psf_pad))
forward = df.Forward_Model_combined(h_full,
shutter = mask_np,
imaging_type = 'spectral')
if simulated == True:
meas_torch = forward(cu.np_to_torch(gt_np.transpose(2,0,1)).type(dtype).unsqueeze(0))
meas_np = cu.torch_to_np(meas_torch)[0]
plt.imshow(meas_np)
```
Set up parameters and network
```
# Define network hyperparameters:
input_depth = 32
INPUT = 'noise'
pad = 'reflection'
LR = 1e-3
tv_weight = 0
reg_noise_std = 0.05
if simulated == True:
num_iter = 100000
net_input = cu.get_noise(input_depth, INPUT, (meas_np.shape[0], meas_np.shape[1])).type(dtype).detach()
else:
num_iter = 4600
input_depth = 1
net_input = cu.get_noise(input_depth, INPUT, (mask_np.shape[-1], meas_np.shape[0], meas_np.shape[1])).type(dtype).detach()
# Initialize network input
net_input_saved = net_input.detach().clone()
noise = net_input.detach().clone()
# reinitialize netowrk and optimizer
if simulated == True:
NET_TYPE = 'skip'
net = md.get_net(input_depth, NET_TYPE, pad, n_channels=32, skip_n33d=128, skip_n33u=128, skip_n11=4, num_scales=5,upsample_mode='bilinear').type(dtype)
else:
print('experimental')
NET_TYPE = 'skip3D'
input_depth = 1
net = md.get_net(input_depth, NET_TYPE, pad, n_channels=1, skip_n33d=128, skip_n33u=128, skip_n11=4, num_scales=4,upsample_mode='trilinear').type(dtype)
#NET_TYPE = 'skip'
#net = md.get_net(input_depth, NET_TYPE, pad, n_channels=32, skip_n33d=128, skip_n33u=128, skip_n11=4, num_scales=5,upsample_mode='bilinear').type(dtype)
p = [x for x in net.parameters()]
optimizer = torch.optim.Adam(p, lr=LR)
# Losses
mse = torch.nn.MSELoss().type(dtype)
def main():
global recons
full_recons = []
meas_ts = cu.np_to_ts(meas_np)
meas_ts = meas_ts.detach().clone().type(dtype).cuda()
for i in range(num_iter):
optimizer.zero_grad()
net_input = net_input_saved + (noise.normal_() * reg_noise_std)
recons = net(net_input)
gen_meas = forward.forward(recons)
gen_meas = F.normalize(gen_meas, dim=[1,2], p=2)
loss = mse(gen_meas, meas_ts)
loss += tv_weight * df.tv_loss(recons)
loss.backward()
print('Iteration %05d, loss %.8f '%(i, loss.item()), '\r', end='')
if i % 100 == 0:
helper.plot(recons)
print('Iteration {}, loss {:.8f}'.format(i, loss.item()))
optimizer.step()
full_recons = helper.preplot(recons)
return full_recons
```
### Run the reconstruction
```
full_recons = main()
full_recons = helper.preplot2(recons)
```
Reconstructed video
```
def plot_slider(x):
plt.title('Reconstruction: frame %d'%(x))
plt.axis('off')
plt.imshow(full_recons[...,x])
return x
interactive(plot_slider,x=(0,full_recons.shape[-1]-1,1))
```
| github_jupyter |
# Renumbering Test
Demonstrate creating a graph with renumbering.
Most cugraph algorithms operate on a CSR representation of a graph. A CSR representation requires an indices array that is as long as the number of edges and an offsets array that is as 1 more than the largest vertex id. This makes the memory utilization entirely dependent on the size of the largest vertex id. For data sets that have a sparse range of vertex ids, the size of the CSR can be unnecessarily large. It is easy to construct an example where the amount of memory required for the offsets array will exceed the amount of memory in the GPU (not to mention the performance cost of having a large number of offsets that are empty but still have to be read to be skipped).
The cugraph renumbering feature allows us to take two columns of any integer type and translate them into a densely packed contiguous array numbered from 0 to (num_unique_values - 1). These renumbered vertices can be used to create a graph much more efficiently.
Another of the features of the renumbering function is that it can take vertex ids that are 64-bit values and map them down into a range that fits into 32-bit integers. The current cugraph algorithms are limited to 32-bit signed integers as vertex ids. and the renumbering feature will allow the caller to translate ids that are 64-bit into a densly packed 32-bit array of ids that can be used in cugraph algorithms. Note that if there are more than 2^31 - 1 unique vertex ids then the renumber method will fail with an error indicating that there are too many vertices to renumber into a 32-bit signed integer.
Note that this version (0.7) is limited to integer types. The intention is to extend the renumbering function to be able to handle strings and other types.
First step is to import the needed libraries
```
import cugraph
import cudf
import socket
import struct
import pandas as pd
import numpy as np
import networkx as nx
```
# Create some test data
This creates a small circle using some ipv4 addresses, storing the columns in a GPU data frame.
The current version of renumbering operates only on integer types, so we translate the ipv4 strings into 64 bit integers.
```
source_list = [ '192.168.1.1', '172.217.5.238', '216.228.121.209', '192.16.31.23' ]
dest_list = [ '172.217.5.238', '216.228.121.209', '192.16.31.23', '192.168.1.1' ]
source_as_int = [ struct.unpack('!L', socket.inet_aton(x))[0] for x in source_list ]
dest_as_int = [ struct.unpack('!L', socket.inet_aton(x))[0] for x in dest_list ]
print("sources came from: " + str([ socket.inet_ntoa(struct.pack('!L', x)) for x in source_as_int ]))
print(" sources as int = " + str(source_as_int))
print("destinations came from: " + str([ socket.inet_ntoa(struct.pack('!L', x)) for x in dest_as_int ]))
print(" destinations as int = " + str(dest_as_int))
```
# Create our GPU data frame
```
df = pd.DataFrame({
'source_list': source_list,
'dest_list': dest_list,
'source_as_int': source_as_int,
'dest_as_int': dest_as_int
})
gdf = cudf.DataFrame.from_pandas(df[['source_as_int', 'dest_as_int']])
gdf.to_pandas()
```
# Run renumbering
The current version of renumbering takes a column of source vertex ids and a column of dest vertex ids. As mentioned above, these must be integer columns.
Output from renumbering is 3 cudf.Series structures representing the renumbered sources, the renumbered destinations and the numbering map which maps the new ids back to the original ids.
In this case,
* gdf['source_as_int'] is a column of type int64
* gdf['dest_as_int'] is a column of type int64
* src_r will be a series of type int32 (we translate back to 32-bit integers)
* dst_r will be a series of type int32
* numbering will be a series of type int64 that translates the elements of src and dst back to their original 64-bit values
Note that because the renumbering translates us to 32-bit integers, if there are more than 2^31 - 1 unique 64-bit values in the source/dest passed into renumbering this would exceed the size of the 32-bit integers so you will get an error from the renumber call.
```
src_r, dst_r, numbering = cugraph.renumber(gdf['source_as_int'], gdf['dest_as_int'])
gdf.add_column("original id", numbering)
gdf.add_column("src_renumbered", src_r)
gdf.add_column("dst_renumbered", dst_r)
gdf.to_pandas()
```
# Data types
Just to confirm, the data types of the renumbered columns should be int32, the original data should be int64, the numbering map needs to be int64 since the values it contains map to the original int64 types.
```
gdf.dtypes
```
# Quick verification
To understand the renumbering, here's a block of verification logic. In the renumbered series we created a new id for each unique value in the original series. The numbering map identifies that mapping. For any vertex id X in the new numbering, numbering[X] should refer to the original value.
```
for i in range(len(src_r)):
print(" " + str(i) +
": (" + str(source_as_int[i]) + "," + str(dest_as_int[i]) +")"
", renumbered: (" + str(src_r[i]) + "," + str(dst_r[i]) +")"
", translate back: (" + str(numbering[src_r[i]]) + "," + str(numbering[dst_r[i]]) +")"
)
```
# Now let's do some graph things...
To start, let's run page rank. Not particularly interesting on our circle, since everything should have an equal rank.
```
G = cugraph.Graph()
G.add_edge_list(src_r, dst_r)
pr = cugraph.pagerank(G)
pr.add_column("original id", numbering)
pr.to_pandas()
```
# Try to run jaccard
Not at all an interesting result, but it demonstrates a more complicated case. Jaccard returns a coefficient for each edge. In order to show the original ids we need to add columns to the data frame for each column that contains one of renumbered vertices. In this case, the columns source and destination contain renumbered vertex ids.
```
jac = cugraph.jaccard(G)
jac.add_column("original_source",
[ socket.inet_ntoa(struct.pack('!L', numbering[x])) for x in jac['source'] ])
jac.add_column("original_destination",
[ socket.inet_ntoa(struct.pack('!L', numbering[x])) for x in jac['destination'] ])
jac.to_pandas()
```
___
Copyright (c) 2019, NVIDIA CORPORATION.
Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0
Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
___
| github_jupyter |
```
# YOU MUST MANUALLY DELETE `results` LIST
# IF YOU WANT TO START OVER WITH AN EMPTY HYPERPARAMETER TABLE
# results = []
try:
assert len(results) > 0
display(df_results.round(3))
except:
results = []
display(results)
# YOU MUST MANUALLY CHANGE RANDOM `seed` OR `test_size`
# IF YOU WANT TO GET NEW MODEL FITS AND `train_test_split`s
hyperparams = dict(test_size=.2, seed=0)
try:
display(pd.Series(hyperparams))
except:
hyperparams = dict(test_size=.2, seed=0)
display(pd.Series(hyperparams))
import os
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from IPython.display import display
np.random.seed(hyperparams['seed'])
sns.set() # seaborn settings that make prettier plots
pd.options.display.max_columns = 500
pd.options.display.max_rows = 40
pd.options.display.max_colwidth = 100
pd.options.display.min_rows = 10
DATA_DIR = ""
DATA_URL = "https://www4.stat.ncsu.edu/~boos/var.select/diabetes.tab.txt"
df = pd.read_csv(DATA_URL, sep='\t')
df.head()
```
The `diabetes.tab.txt` file is a tab-delimitted text file with the original unnormalized units (age in years, blood pressure in mmHg, gender as 1 for female, 2 for male.
The `diabetes.rwrite1.txt` url will load a dataset with standardized values for each feature, but more informative column names. In both data sets, here is what the names mean.
0. age: in years
1. sex: 1=male, 2=female
2. bmi: body mass index >35=obese, >30=overweight, <18.5=underweight
3. bp/map: mean arterial pressure (blood pressure, systolic+diastolic divided by 2)
4. s1/tc: TC level is a measure of a B12 transportation molecule that is not bound to B12 yet. I high ratio of holotranscobalamin (holo TC or TCH) to transcobalamin (TC) indicates healthy availability of B12 for absorption. TCH above 50 pmol/liter is considered good.
5. s2/ldl: Low density lipid (good cholesterol)
6. s3/hdl: High density Lipid cholesterol (bad cholesterol)
7. s4/tch: holo TC level? (B12 bound tot he transport molecule, **t**rans**c**obalamin, to create **h**olo**t**rans**c**obalamin), <35pmol/L=B12deficiency, >50pmol/L=goodB12
8. s5/ltg:
9. s6/glu: glucose level
10. y: a quantitative measure of disease progression one year after baseline
```
column_names = list(pd.read_csv('https://www4.stat.ncsu.edu/~boos/var.select/diabetes.rwrite1.txt', sep=' ').columns)
df.columns = column_names
df.head().round(1)
column_names[3] = 'bp'
column_names[-1] = 'severity'
df.columns = column_names
df.head()
display(df.round(1))
target_names = ['severity']
feature_names = [name for name in df.columns if name not in target_names]
print(f' target_names: {target_names}')
print(f'feature_names: {feature_names}')
fig = sns.pairplot(df, x_vars=feature_names[:4], y_vars=target_names)
fig = sns.pairplot(df, x_vars=feature_names[4:7], y_vars=target_names)
fig = sns.pairplot(df, x_vars=feature_names[7:], y_vars=target_names)
```
Create a training and testset.
The training set is like the question and answer pairs you get to see during a school lesson.
The test set is like the exam question and answer pairs that the teacher grades you on at the end of the course.
Use the training set to show your machine learning model the relationship between your features (age, gender, bmi, blood tests etc) and your target variables (diabetes severity).
You will use the training set to traiin or fit the model.
You'll use the test set to see how well you model will work (it's accuracy, standard error, precision, recall, etc) in the real world.
You'll make predictions for the test set "questions" (`X_test`) and see how closely they match the test set answers (`y_test`).
```
from sklearn.model_selection import train_test_split
### BEGIN SOLUTION
X_train, X_test, y_train, y_test = train_test_split(df[feature_names], df[target_names], test_size=hyperparams['test_size'])
X_train = pd.DataFrame(X_train, columns=feature_names)
X_test = pd.DataFrame(X_test, columns=feature_names)
y_train = pd.DataFrame(y_train, columns=target_names)
y_test = pd.DataFrame(y_test, columns=target_names)
print(f'X_train.shape: {X_train.shape}; y_train.shape: {y_train.shape}')
print(f' X_test.shape: {X_test.shape}; y_test.shape: {y_test.shape}')
### END SOLUTION
display(X_train.describe(include='all'))
display(y_train.describe(include='all'))
from sklearn.neural_network import MLPRegressor
regr = MLPRegressor(hidden_layer_sizes=(5, 2), max_iter=200000, random_state=0)
# scikitlearn Multi-layer perceptrons expect a 1-D array (pd.DataFrame column or pd.Series) for y:
regr.fit(X=X_train, y=y_train['severity'])
display(X_train[features].shape)
for layer in range(len(regr.coefs_)):
print(f'LAYER: {layer}' + (' (INPUT LAYER)' if layer == 0 else ''))
W = regr.coefs_[layer]
if layer > 0:
w0 = regr.intercepts_[layer - 1]
else:
w0 = np.zeros(W.shape[0])
df_params = pd.concat([
pd.DataFrame(W, columns=[f'neuron_{layer+1}_{i+1}' for i in range(W.shape[1])]),
pd.DataFrame(w0, columns=['intercept'])],
axis=1)
if layer is 0:
df_params.index = feature_names
else:
df_params.index = range(1, len(df_params) + 1)
display(df_params)
print(f'LAYER: {layer+1} (OUTPUT LAYER)')
display(regr.intercepts_[-1])
y_train.values
# fig = sns.scatterplot(x=X_train[features], y=y_train[target_names[0]])
y_train_pred = regr.predict(X_train)
df_y_train = pd.DataFrame(y_train_pred, columns=['pred_severity'])
df_y_train['true_severity'] = y_train.values.flatten()
df_y_train['residual'] = y_train_pred - y_train.values.flatten()
# fig = plt.plot(X_train['bmi'].values.flatten(), , color='r', linewidth=2)
# plt.xlabel('BMI')
# plt.ylabel('Diabetes Severity')
# plt.title(f'severity = {regr.coef_.round(2)[0]} * bmi + {regr.intercept_.round(2)}')
# print(f'lr_bmi.intercept_: {lr_bmi.intercept_.round(2)}')
# print(f' lr_bmi.coef_: {lr_bmi.coef_.round(2)}')
df_y_train
df_y_train.plot(kind='scatter', x='true_severity', y='residual')
def mae_rmse(y, y_pred):
e = y_pred - y
mae = np.sum(np.abs(e)) / len(e_test_bmi) # .mean()
rmse = np.sqrt(np.mean(e ** 2))
return mae[0], rmse[0]
y_train_pred = lr_bmi.predict(X_train[features])
mae_train_bmi, rmse_train_bmi = mae_rmse(y_train, y_train_pred)
results[-1]['train_score'] = lr_bmi.score(X_train[features], y_train)
results[-1]['train_rmse'] = rmse_train_bmi
results[-1]['train_mae'] = mae_train_bmi
display(pd.DataFrame(results).round(2))
### BEGIN SOLUTION
y_test_pred = lr_bmi.predict(X_test[features])
# print(f'y_test_pred.shape: {y_test_pred.shape}')
mae_test_bmi, rmse_test_bmi = mae_rmse(y_test, y_test_pred)
results[-1]['test_score'] = lr_bmi.score(X_test[features], y_test)
results[-1]['test_rmse'] = rmse_test_bmi
results[-1]['test_mae'] = mae_test_bmi
display(pd.DataFrame(results).round(2))
### END SOLUTION
features
rmse_overfit_ratio = round((rmse_test_bmi - rmse_train_bmi) / rmse_test_bmi, 3)
rmse_overfit_ratio
assert abs(rmse_overfit_ratio) < 0.2
score_overfit_ratio = round((results[-1]['train_score'] - results[-1]['test_score']) / results[-1]['test_score'], 3)
score_overfit_ratio
lr_multi = LinearRegression()
features = feature_names
lr_multi = lr_multi.fit(X_train, y_train)
print(f'lr_multi.intercept_: {lr_multi.intercept_.round(2)}')
print('lr_multi_coef:')
lr_multi_coef = pd.Series(lr_multi.coef_[0], index=features)
print(lr_multi_coef.round(2))
```
In the sex column a value of 2 indicates female and 1 indicates male.
What does this coefficient list tell you about the affect that sex (gender) has on ones likelihood of developing diabetes?
```
fig = sns.scatterplot(x=X_train['bmi'].values, y=y_train.values.flatten())
print(lr_multi.coef_)
print(lr_multi.predict(X_train).flatten().shape)
fig = sns.scatterplot(X_train['bmi'], lr_multi.predict(X_train).flatten(), color='r', linewidth=2)
plt.xlabel('BMI')
plt.ylabel('Diabetes Severity')
# print(lr_multi_coef['age'])
# print(lr_multi_coef['bmi'])
plt.title(f'severity = ... + {lr_multi_coef["age"].round(2)}*age + {lr_multi_coef["bmi"].round(2)}*bmi + {lr_multi.intercept_.round(2)}')
print(f'lr_multi.intercept_: {lr_multi.intercept_.round(2)}')
print(' lr_multi_coef_:')
print(lr_multi_coef.round(2))
```
Let's create a function that we can use to train a model and measure it's accuracy.
API:
Inputs:
model: untrained sklearn model object (this will allow us to use any model we like)
X_train, y_train: training set of features (X) and target labels (y), defaults to the entire original train_test_split
X_test, y_test: test set to measure accuracy on an unseen dataset
Outputs:
Dictionary of model hyperparameters (class name, features), trained parameters (coefficients, intercept), and performance (RMSE, MAE, pierson correlation score)
```
def fit_evaluate_model(model=LinearRegression(),
X_train=X_train[features], y_train=y_train,
X_test=X_test[features], y_test=y_test,
hyperparams=hyperparams):
features = list(X_train.columns)
hyperparams.update({
'model': model.__class__.__name__,
'num_features': len(features),
'num_samples': len(X_train)
})
model = model.fit(X_train, np.array(y_train).reshape(-1,1))
print(f'model.intercept_: {model.intercept_.round(2)}')
print('model_coef:')
model_coef = pd.Series(model.coef_.flatten(), index=features)
print(model_coef.round(2))
y_train_pred = model.predict(X_train[features]).flatten()
e_train = (np.array(y_train_pred).flatten() - np.array(y_train).flatten())
mae_train = np.sum(np.abs(e_train)) / len(e_train) # .mean()
rmse_train = np.sqrt((e_train ** 2).mean())
hyperparams.update({'train_score': model.score(X_train[features], y_train),
'train_rmse': rmse_train,
'train_mae': mae_train})
y_test_pred = model.predict(X_test[features]).flatten()
e_test = y_test_pred - y_test.values[:,0]
mae_test = np.sum(np.abs(e_test)) / len(e_test) # .mean()
rmse_test = np.sqrt((e_test ** 2).mean())
hyperparams.update({'test_score': model.score(X_test[features], y_test),
'test_rmse': rmse_test,
'test_mae': mae_test})
results_series = pd.concat([pd.Series(hyperparams), model_coef])
return results_series
lr_multi = LinearRegression()
results.append(fit_evaluate_model(model=lr_multi))
e_train = lr_multi.predict(X_train).flatten() - np.array(y_train).flatten()
fig = sns.scatterplot(x=np.array(y_train).flatten(), y=e_train)
ylab = plt.ylabel('Error (pred_severity - true_severity)')
xlab = plt.xlabel('True Severity')
titl = plt.title('Residuals Plot')
lr_en = ElasticNet()
results.append(fit_evaluate_model(model=lr_en))
lr_lasso = Lasso()
results.append(fit_evaluate_model(model=lr_lasso))
# results[-1]['description'] ='lasso on original features'
df_results = pd.DataFrame(results).sort_values(['test_rmse'])
df_results.round(2)
X_train['ldl_x_hdl'] = X_train['ldl'] * X_train['hdl']
X_test['ldl_x_hdl'] = X_test['ldl'] * X_test['hdl']
lr_cholest = LinearRegression()
results.append(fit_evaluate_model(model=lr_cholest, X_train=X_train, X_test=X_test))
df_results = pd.DataFrame(results).sort_values(['test_rmse'])
df_results.round(2)
X_train['ldl_d_hdl'] = X_train['ldl'] / X_train['hdl']
X_test['ldl_d_hdl'] = X_test['ldl'] / X_test['hdl']
lr_cholest = LinearRegression()
results.append(fit_evaluate_model(model=lr_cholest, X_train=X_train, X_test=X_test))
df_results = pd.DataFrame(results).sort_values(['test_rmse'])
df_results.round(2)
from sklearn import preprocessing
x_scaler = preprocessing.MinMaxScaler()
x_scaler.fit(X_train)
X_train_scaled = pd.DataFrame(x_scaler.transform(X_train), columns=X_train.columns, index=X_train.index)
X_test_scaled = pd.DataFrame(x_scaler.transform(X_test), columns=X_test.columns, index=X_test.index)
X_test_scaled.head()
y_scaler = preprocessing.MinMaxScaler()
y_scaler.fit(y_train)
y_train_scaled = pd.DataFrame(y_scaler.transform(y_train), columns=y_train.columns, index=y_train.index)
y_test_scaled = pd.DataFrame(y_scaler.transform(y_test), columns=y_test.columns, index=y_test.index)
y_test_scaled.head()
lr_scaled = LinearRegression()
results.append(fit_evaluate_model(model=lr_scaled,
X_train=X_train_scaled, X_test=X_test_scaled,
y_train=y_train_scaled, y_test=y_test_scaled))
results[-1]['x_scaler'] = x_scaler.__class__.__name__
results[-1]['y_scaler'] = y_scaler.__class__.__name__
df_results = pd.DataFrame(results).sort_values(['test_score'], ascending=False)
df_results.round(2)
```
Check out the residuals plot for the best model we've created so far (model id=1 a LinearRegression with 10 unscaled features).
Do you see any pattern in this error between your predictions and the truth?
This will help you think of ways to engineer some new features.
```
def plot_residuals(y, y_pred):
e = np.array(y_pred).flatten() - np.array(y).flatten()
fig = sns.scatterplot(x=np.array(y).flatten(), y=e)
plt.ylabel('Error (pred_severity - true_severity)')
plt.xlabel('True Severity')
plt.title('Residuals Plot')
return fig
fig = plot_residuals(y_test, lr_multi.predict(X_test[features]))
```
It seems that a LinearRegression overestimates severity for low severity patients.
And it underestimates severity for patients that developer sever diabetes.
You can square or take the exponent of our features to give the LinearRegression model curvature.
```
(X_train ** 2).isnull().sum()
X_train2 = pd.concat([X_train, X_train ** 2], axis=1)
X_train2.columns = list(X_train.columns) + [c + '2' for c in X_train.columns]
X_test2 = pd.concat([X_test, X_test ** 2], axis=1)
X_test2.columns = list(X_test.columns) + [c + '2' for c in X_test.columns]
lr_sqrd = LinearRegression()
results.append(fit_evaluate_model(model=lr_sqrd,
X_train=X_train2, X_test=X_test2,
y_train=y_train, y_test=y_test))
df_results = pd.DataFrame(results).sort_values(['test_score'], ascending=False)
df_results.round(2)
lr_sqrd_lasso = Lasso()
results.append(fit_evaluate_model(model=lr_sqrd_lasso,
X_train=X_train2, X_test=X_test2,
y_train=y_train, y_test=y_test))
df_results = pd.DataFrame(results).sort_values(['test_score'], ascending=False)
df_results.round(2)
lr_sqrd_elast = ElasticNet()
results.append(fit_evaluate_model(model=lr_sqrd_elast,
X_train=X_train2, X_test=X_test2,
y_train=y_train, y_test=y_test))
results[-1]['alpha'] = lr_sqrd_elast.alpha
df_results = pd.DataFrame(results).sort_values(['test_score'], ascending=False)
df_results.round(2)
### BEGIN SOLUTION
lr_sqrd_ridge = Ridge()
results.append(fit_evaluate_model(model=lr_sqrd_ridge,
X_train=X_train2, X_test=X_test2,
y_train=y_train, y_test=y_test))
### END SOLUTION
results[-1]['alpha'] = lr_sqrd_ridge.alpha
df_results = pd.DataFrame(results).sort_values(['test_score'], ascending=False)
df_results.round(2)
### BEGIN SOLUTION
lr_sqrd_ridge = Ridge(alpha=1000)
results.append(fit_evaluate_model(model=lr_sqrd_ridge,
X_train=X_train2, X_test=X_test2,
y_train=y_train, y_test=y_test))
results[-1]['alpha'] = lr_sqrd_ridge.alpha
### END SOLUTION
df_results = pd.DataFrame(results).sort_values(['test_score'], ascending=False)
df_results.round(2)
### BEGIN SOLUTION
lr_sqrd_lasso100 = Lasso(alpha=100)
results.append(fit_evaluate_model(model=lr_sqrd_lasso100,
X_train=X_train2, X_test=X_test2,
y_train=y_train, y_test=y_test))
results[-1]['alpha'] = lr_sqrd_lasso100.alpha
### END SOLUTION
df_results = pd.DataFrame(results).sort_values(['test_score'], ascending=False)
df_results.round(3)
df_results['overfitness'] = (df_results['train_score'] - df_results['test_score']) / df_results['train_score']
df_results.round(3)
```
| github_jupyter |
```
from sklearn.datasets import load_wine
wine_data = load_wine()
dir(wine_data)
print(wine_data.DESCR)
inputs = wine_data.data
output = wine_data.target
inputs.shape
output.shape
wine_data.feature_names
import pandas as pd
df = pd.DataFrame(inputs, columns=wine_data.feature_names)
df = pd.concat([df, pd.DataFrame(output)], axis=1)
df
df.describe()
df.describe().style.format("{:.5f}")
import matplotlib.pyplot as plt
plt.matshow(df.corr())
plt.xticks(range(len(df.columns)), df.columns)
plt.yticks(range(len(df.columns)), df.columns)
plt.colorbar()
plt.show()
```
Chapter Break
```
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(inputs, output, test_size=0.33, random_state=42)
from sklearn.preprocessing import PolynomialFeatures, StandardScaler
from sklearn.pipeline import make_pipeline
pipe = make_pipeline(LinearRegression())
pipe.fit(X_train, y_train)
pipe.score(X_train, y_train)
pipe.score(X_test, y_test)
from sklearn.linear_model import Ridge
# tactic 1: minimize weights, smaller the better, higher penalty on large weights
# = ridge regression
pipe = make_pipeline(StandardScaler(), PolynomialFeatures(degree=3), Ridge())
pipe.fit(X_train, y_train)
pipe.steps[2][1].coef_
pipe.steps[2][1].coef_.max(), pipe.steps[2][1].coef_.min(), pipe.steps[2][1].coef_.std()
pipe.score(X_train, y_train)
pipe.score(X_test, y_test)
from sklearn.linear_model import Lasso
# tactic 2: minimize number of non-zero weights
# = Lasso
pipe = make_pipeline(StandardScaler(), PolynomialFeatures(degree=3), Lasso())
pipe.fit(X_train, y_train)
pipe.score(X_train, y_train)
pipe.score(X_test, y_test)
pipe.steps[2][1].coef_
pipe.steps[2][1].coef_.max(), pipe.steps[2][1].coef_.min(), pipe.steps[2][1].coef_.std()
from sklearn.linear_model import ElasticNet
# tactic 3: mix lasso and ridge!
# = elasticnet
pipe = make_pipeline(StandardScaler(), PolynomialFeatures(degree=3), ElasticNet())
pipe.fit(X_train, y_train)
pipe.score(X_train, y_train)
pipe.score(X_test, y_test)
pipe.steps[2][1].coef_
pipe.steps[2][1].coef_.max(), pipe.steps[2][1].coef_.min(), pipe.steps[2][1].coef_.std()
```
----
# Implementing a model to classify wines
```
from sklearn.datasets import load_wine
wine_data = load_wine()
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(
wine_data.data, wine_data.target, test_size=0.5, random_state=42)
import numpy as np
import pandas as pd
df_x_train = pd.DataFrame(X_train, columns=wine_data.feature_names)
df_x_train.describe()
from sklearn.preprocessing import StandardScaler
from sklearn.pipeline import make_pipeline
from sklearn import linear_model
from sklearn import svm
from sklearn.naive_bayes import GaussianNB
from sklearn.metrics import confusion_matrix
pipe = make_pipeline(GaussianNB())
pipe.fit(X_train, y_train)
pipe.score(X_train, y_train), pipe.score(X_test, y_test)
confusion_matrix(y_test, pipe.predict(X_test))
pipe = make_pipeline(svm.SVC())
pipe.fit(X_train, y_train)
pipe.score(X_train, y_train), pipe.score(X_test, y_test)
confusion_matrix(y_test, pipe.predict(X_test))
pipe = make_pipeline(linear_model.LogisticRegression())
pipe.fit(X_train, y_train)
pipe.score(X_train, y_train), pipe.score(X_test, y_test)
confusion_matrix(y_test, pipe.predict(X_test))
```
| github_jupyter |
```
envname = 'variables/loop_stim10e-16.0et6.0ph1.0pvaryt0.1plNonebp0.5.pkl'
# import stuff
from placerg.funcs import *
from placerg.objects import*
from placerg.funcsrg import *
from scipy.optimize import curve_fit
import matplotlib.pyplot as plt
import matplotlib.cm as cm
import matplotlib
# set up notebook displayt
np.set_printoptions(threshold=5)
alpha=0.4
color='black'
cmap='Greys'
colorline='black'
linethick=3.
colorfit='grey'
plt.style.use('seaborn-paper')
fontsize=20
ticksize=20
fontsizesmall=25
ticksizesmall=20
legendsize=20
alpha=.3
colorfit='gray'
linecolor='black'
palit=['black','firebrick', 'crimson', 'orangered', 'darkorange', 'goldenrod', 'gold', 'khaki']
mycmap = cm.gnuplot
matplotlib.rcParams['mathtext.fontset'] = 'stix'
matplotlib.rcParams['font.family'] = 'STIXGeneral'
# load in objects
allo=load_object(envname)
orderplot(allo)
if allo.labeltype[0]=='eta':
maxx=np.max(np.array(allo.eta).flatten())
minn=np.min(np.array(allo.eta).flatten())
cc=allo.eta
if allo.labeltype[0]=='epsilon':
maxx=np.max(np.array(allo.epsilon).flatten())
minn=np.min(np.array(allo.epsilon).flatten())
cc=allo.epsilon
if allo.labeltype[0]=='time constant':
maxx=np.max(np.array(allo.timeconst).flatten())
minn=np.min(np.array(allo.timeconst).flatten())
cc=np.array(allo.timeconst)[:,0]
if allo.labeltype[0]=='# of stimuli':
maxx=np.max(np.array(allo.stim).flatten())
minn=np.min(np.array(allo.stim).flatten())
cc=allo.stim
if allo.labeltype[0]=='p':
maxx=np.max(np.array(allo.percell).flatten())
minn=np.min(np.array(allo.percell).flatten())
cc=allo.percell
for i in range(len(allo.labeltype)):
allo.labeltype[i]='q'
mrange=maxx-minn
allo.label
"""
Here plot the eigenvalues from each sucessive RG step, averaged over all clusters and
normalized by cluster size.
"""
fig, ax = plt.subplots(2,2, sharex=True, sharey=True, figsize=(10,10))
ylabel= 'eigenvalue'
ax[0,0].set_ylabel(ylabel, fontsize=fontsize)
ax[1,0].set_ylabel(ylabel, fontsize=fontsize)
ax[1,0].set_xlabel('rank$/K$', fontsize=fontsize)
ax[1,1].set_xlabel('rank$/K$', fontsize=fontsize)
c=0
for h in np.array([0,5,10,19]):
n=int(c/2)
l=c-2*n
errplot=allo.eigspecerr[h]
xplot,plot=(allo.eigspecx[h], allo.eigspec[h])
for m in range(len(xplot)):
ax[n,l].errorbar(xplot[m], plot[m], yerr=errplot[m], \
label= r'$K=$'+str(2**(m+4)),\
color=palit[m+2], marker='o', \
markersize=5, linestyle='None', linewidth=2)
popt=allo.mu[h]
ax[n,l].plot(xplot[m],linfunc(xplot[m], \
popt[0], popt[1]), '--', color=colorfit, linewidth=2)
ax[n,l].tick_params(labelsize=ticksize)
ax[n,l].text(.005, .015, r'$\mu$='+ str(np.round(popt[1],3))+r'$\pm$'\
+str(np.round(allo.muerr[h]\
[0], 3)), fontsize=ticksize)
ax[n,l].text(.005, .0055, r'$q=$'+str(np.round(allo.label[h],2)), \
fontsize=ticksize)
ax[n,l].set_yscale('log')
ax[n,l].set_xscale('log')
ax[n,l].set_ylim(top=1)
c+=1
for n in range(2):
for l in range(2):
ax[n,l].set_yticks([.1, .01,.001,.0001])
ax[n,l].tick_params(length=6, width=1, which='major')
ax[n,l].tick_params(length=3, width=1, which='minor')
#a.grid(True, linewidth=1)
ax[0,0].text(.0015,0.7,r'(A)', fontsize=ticksize, weight='bold')
ax[0,1].text(.0015,0.7,r'(B)', fontsize=ticksize, weight='bold')
ax[1,0].text(.0015,0.7,r'(C)', fontsize=ticksize, weight='bold')
ax[1,1].text(.0015,0.7,r'(D)', fontsize=ticksize, weight='bold')
lines_labels = [ax.get_legend_handles_labels() for ax in fig.axes[:1]]
lines, labels = [sum(z, []) for z in zip(*lines_labels)]
fig.legend(lines, labels, fontsize=fontsize-5, loc=(.2,.6))
plt.tight_layout()
name=str(envname)+'eigs.pdf'
plt.savefig(name)
```
# variance of activity at each RG step over clusters
```
"""
plot coarse grained variance vs. cluster size
"""
fig, ax = plt.subplots(2,2, sharex=True, sharey=True, figsize=(10,10))
ylabel= 'activity variance'
ax[0,0].set_ylabel(ylabel, fontsize=fontsize)
ax[1,0].set_ylabel(ylabel, fontsize=fontsize)
ax[1,0].set_xlabel(r'cluster size $K$', fontsize=fontsize)
ax[1,1].set_xlabel(r'cluster size $K$', fontsize=fontsize)
c=0
for h in np.array([0,5,10,19]):
n=int(c/2)
l=c-2*n
ax[n,l].errorbar(allo.varx[h],allo.var[h], allo.varerr[h], \
color='black', marker='o', markersize=5, linewidth=2, linestyle='None')
popt = allo.alpha[h]
ax[n,l].plot(allo.varx[h],linfunc(allo.varx[h], \
popt[0], popt[1]), '--', color=colorfit, linewidth=2)
ax[n,l].plot(allo.varx[h], linfunc(allo.varx[h], popt[0], 1.), \
color=colorfit, linewidth=2, alpha=alpha)
ax[n,l].text(2, 5, r'$q=$'+str(np.round(allo.label[h],2)), fontsize=ticksize)
ax[n,l].tick_params(labelsize=ticksize)
ax[n,l].text(2, 20, r'${\alpha}$='+ str(np.format_float_positional(popt[1],unique=False, precision=3))+r'$\pm$'+\
str(np.format_float_positional(allo.alphaerr[h][0], unique=False, precision=3)), fontsize=fontsize)
ax[n,l].set_yscale('log')
ax[n,l].set_xscale('log')
ax[n,l].set_ylim(top=260, bottom=.01)
c+=1
for n in range(2):
for l in range(2):
#ax[n,l].set_yticks([.1, .01,.001,.0001])
ax[n,l].tick_params(length=6, width=1, which='major')
ax[n,l].tick_params(length=3, width=1, which='minor')
#a.grid(True, linewidth=1)
ax[0,0].text(.35,155,r'(A)', fontsize=ticksize, weight='bold')
ax[0,1].text(.35,155,r'(B)', fontsize=ticksize, weight='bold')
ax[1,0].text(.35,155,r'(C)', fontsize=ticksize, weight='bold')
ax[1,1].text(.35,155,r'(D)', fontsize=ticksize, weight='bold')
plt.tight_layout()
name=str(envname)+'var.pdf'
plt.savefig(name)
"""
Plot log probability of complete cluster silence vs cluster size
"""
fig, ax = plt.subplots(2,2, sharex=True, sharey=True, figsize=(10,10))
ylabel= r'$F$'
ax[0,0].set_ylabel(ylabel, fontsize=fontsize)
ax[1,0].set_ylabel(ylabel, fontsize=fontsize)
ax[1,0].set_xlabel(r'cluster size $K$', fontsize=fontsize)
ax[1,1].set_xlabel(r'cluster size $K$', fontsize=fontsize)
c=0
for h in np.array([0,5,10,19]):
n=int(c/2)
l=c-2*n
x=allo.psilx[h]
y=allo.psil[h]
popt= allo.beta[h]
ax[n,l].errorbar(allo.psilx[h], allo.psil[h],allo.psilerr[h], \
color='black', marker='o', linestyle='None', markersize=5)
ax[n,l].plot(np.arange(np.min(allo.psilx[h]),np.max(allo.psilx[h]), .01),\
(probfunc(np.arange(np.min(allo.psilx[h]),np.max(allo.psilx[h]), .01), \
popt[0], popt[1])), '--', color=colorfit, linewidth=2)
ax[n,l].text(2, -1.0, r'$q=$'+str(np.round(allo.label[h],2)),\
fontsize=ticksize)
ax[n,l].text(2, -.75, r'$\tilde{\beta}=$'+str(np.format_float_positional(popt[1], unique=False, precision=3))+r'$\pm$'+\
str(np.format_float_positional(allo.alphaerr[h][0], unique=False, precision=3)),fontsize=ticksize)
ax[n,l].tick_params(labelsize=ticksize)
ax[n,l].set_xscale('log')
ax[n,l].set_ylim(top=0.4)
c+=1
for n in range(2):
for l in range(2):
#ax[n,l].set_yticks([.1, .01,.001,.0001])
ax[n,l].tick_params(length=6, width=1, which='major')
ax[n,l].tick_params(length=3, width=1, which='minor')
#a.grid(True, linewidth=1)
ax[0,0].text(.4,.25,r'(A)', fontsize=ticksize, weight='bold')
ax[0,1].text(.4,.25,r'(B)', fontsize=ticksize, weight='bold')
ax[1,0].text(.4, .25,r'(C)', fontsize=ticksize, weight='bold')
ax[1,1].text(.4,.25,r'(D)', fontsize=ticksize, weight='bold')
plt.tight_layout()
name=str(envname)+'freeenergy.pdf'
plt.savefig(name)
minnm=16
maxxm=128
mrangem=np.abs(minnm-maxxm)
x=allo.actmomx
plott=allo.actmom
plterr=allo.actmomerr
fig, ax = plt.subplots(2,2, sharex=True, sharey=True, figsize=(10,10))
ylabel= r'density'
ax[0,0].set_ylabel(ylabel, fontsize=fontsize)
ax[1,0].set_ylabel(ylabel, fontsize=fontsize)
ax[1,0].set_xlabel('normalized activity', fontsize=fontsize)
ax[1,1].set_xlabel('normalized activity', fontsize=fontsize)
c=0
for h in np.array([0,5,10,19]):
n=int(c/2)
l=c-2*n
for i in (np.arange(len(allo.actmomx[0]))):
if i==3:
ax[n,l].errorbar(x[h][i],plott[h][i], plterr[h][i], \
label='N/'+str(2**(i+4)), \
color=palit[i+2], linewidth=2, errorevery=3, alpha=.7)
popt, pcov = curve_fit(gaussian,x[h][i], plott[h][i])
ax[n,l].plot(np.arange(-4, 4,.1), \
gaussian(np.arange(-4, 4, .1),\
popt[0], popt[1]), '--', color=colorfit, linewidth=2)
else:
ax[n,l].plot(x[h][i],plott[h][i], \
label='N/'+str(2**(i+4)), \
color=palit[i+2], linewidth=2)
ax[n,l].text(-8, 4, r'$q=$'+str(np.round(allo.label[h],2)), \
fontsize=ticksize)
ax[n,l].tick_params(labelsize=ticksize)
ax[n,l].set_yscale('log')
ax[n,l].set_ylim(bottom=10**-6, top=9)
c+=1
for n in range(2):
for l in range(2):
#ax[n,l].set_yticks([.1, .01,.001,.0001])
ax[n,l].tick_params(length=6, width=1, which='major')
ax[n,l].tick_params(length=3, width=1, which='minor')
#a.grid(True, linewidth=1)
ax[0,0].legend(fontsize=fontsize)
ax[0,0].text(-14,4,r'(A)', fontsize=ticksize, weight='bold')
ax[0,1].text(-14,4,r'(B)', fontsize=ticksize, weight='bold')
ax[1,0].text(-14,4,r'(C)', fontsize=ticksize, weight='bold')
ax[1,1].text(-14,4,r'(D)', fontsize=ticksize, weight='bold')
plt.tight_layout()
name=str(envname)+'momdist.pdf'
plt.savefig(name)
minnm=2
maxxm=256
mrangem=np.abs(minnm-maxxm)
x=allo.autocorrx
plterr=allo.autocorrerr
result=allo.autocorr
fig, ax = plt.subplots(2,2, sharex=True, sharey=True, figsize=(10,10))
ylabel= r'$C(t)$'
ax[0,0].set_ylabel(ylabel, fontsize=fontsize)
ax[1,0].set_ylabel(ylabel, fontsize=fontsize)
ax[1,0].set_xlabel(r'time $t$', fontsize=fontsize)
ax[1,1].set_xlabel(r'time $t$', fontsize=fontsize)
c=0
for h in np.array([0,5,10,19]):
n=int(c/2)
l=c-2*n
for i in range(result[h].shape[0]):
#print(result[l][i, int(result[l].shape[1]/2)-50:int(result[l].shape[1]/2)+50])
ax[n,l].errorbar((x[h][int(result[h].shape[1]/2)-\
20:int(result[h].shape[1]/2)+20]), \
(result[h][i, int(result[h].shape[1]/2)-20:int(result[h].\
shape[1]/2)+20]),\
(plterr[h][i][int(result[h].shape[1]/2)-20:int(result[h]\
.shape[1]/2)+20]), \
label=r'$K$ ='+str(2**(i+2)),color=palit[i],\
linewidth=2)
ax[n,l].text(-10, 1.0, r'$q=$'+str(np.round(allo.label[h],2)), \
fontsize=fontsize)
ax[n,l].tick_params(labelsize=ticksize)
ax[n,l].set_ylim(top=1.15)
ax[n,l].set_xlim(-15,15)
c+=1
for n in range(2):
for l in range(2):
#ax[n,l].set_yticks([.1, .01,.001,.0001])
ax[n,l].tick_params(length=6, width=1, which='major')
ax[n,l].tick_params(length=3, width=1, which='minor')
#a.grid(True, linewidth=1)
ax[0,0].legend(fontsize=fontsize-5)
ax[0,0].text(-19,1.1,r'(A)', fontsize=ticksize, weight='bold')
ax[0,1].text(-19,1.1,r'(B)', fontsize=ticksize, weight='bold')
ax[1,0].text(-19,1.1,r'(C)', fontsize=ticksize, weight='bold')
ax[1,1].text(-19,1.1,r'(D)', fontsize=ticksize, weight='bold')
plt.tight_layout()
name=str(envname)+'dynamic.pdf'
plt.savefig(name)
"""
plot exponents
"""
fig, ax = plt.subplots(2,2, sharex=True, sharey=True, figsize=(10,10))
ylabel= r'$\tau_c$'
ax[0,0].set_ylabel(ylabel, fontsize=fontsize)
ax[1,0].set_ylabel(ylabel, fontsize=fontsize)
ax[1,0].set_xlabel(r'cluster size $K$', fontsize=fontsize)
ax[1,1].set_xlabel(r'cluster size $K$', fontsize=fontsize)
c=0
for h in np.array([0,5,10,19]):
n=int(c/2)
l=c-2*n
ax[n,l].errorbar(2**np.arange(1,8),allo.tau[h],allo.tauerr[h], color=colorline, \
label='taus', marker='o', markersize=5, linestyle='None')
popt= allo.z[h]
ax[n,l].plot(2**np.arange(1,8), linfunc(2**np.arange(1,8), \
popt[0], popt[1]), '--', label='fit', \
color=colorfit, linewidth=2)
ax[n,l].text(2, 3, r'$\tilde{z}=$'+str(np.format_float_positional(popt[1],unique=False, precision=3))+r'$\pm$'+\
str(np.format_float_positional(allo.zerr[h][0], unique=False, precision=3)), fontsize=ticksize)
ax[n,l].set_yscale('log')
ax[n,l].set_xscale('log')
ax[n,l].text(2, 2.5, r'$q=$'+str(np.round(allo.label[h],2)), \
fontsize=fontsize)
ax[n,l].set_ylim(top=3.8, bottom=0.8)
c+=1
for n in range(2):
for l in range(2):
ax[n,l].set_yticks([1,2,3])
ax[n,l].tick_params(length=6, width=1, which='major', labelsize=ticksize)
ax[n,l].tick_params(length=3, width=1, which='minor')
#a.grid(True, linewidth=1)
ax[0,0].text(1,3.5,r'(A)', fontsize=ticksize, weight='bold')
ax[0,1].text(1,3.5,r'(B)', fontsize=ticksize, weight='bold')
ax[1,0].text(1,3.5,r'(C)', fontsize=ticksize, weight='bold')
ax[1,1].text(1,3.5,r'(D)', fontsize=ticksize, weight='bold')
plt.tight_layout()
name=str(envname)+'dynamicexps.pdf'
plt.savefig(name)
minnm=2
maxxm=256
mrangem=np.abs(minnm-maxxm)
x=allo.autocorrx
plterr=allo.autocorrerr
result=allo.autocorr
ylabel= r'$C(t)$'
fig, ax = plt.subplots(2,2, sharex=True, sharey=True, figsize=(10,10))
ax[0,0].set_ylabel(ylabel, fontsize=fontsize)
ax[1,0].set_ylabel(ylabel, fontsize=fontsize)
ax[1,0].set_xlabel(r'time $t/\tau_c$', fontsize=fontsize)
ax[1,1].set_xlabel(r'time $t/\tau_c$', fontsize=fontsize)
c=0
for h in np.array([0,5,10,19]):
n=int(c/2)
l=c-2*n
for i in range(result[h].shape[0]):
#print(result[l][i, int(result[l].shape[1]/2)-50:int(result[l].shape[1]/2)+50])
ax[n,l].errorbar((x[h][int(result[h].shape[1]/2)-\
20:int(result[h].shape[1]/2)+20])/allo.tau[h][i], \
(result[h][i, int(result[h].shape[1]/2)-20:int(result[h].\
shape[1]/2)+20]),\
(plterr[h][i][int(result[h].shape[1]/2)-20:int(result[h]\
.shape[1]/2)+20]), \
label=r'$K$ ='+str(2**(i+2)), color=palit[i],\
linewidth=2)
ax[n,l].text(-10, 1.0, r'$q=$'+str(np.round(allo.label[h],2)), \
fontsize=fontsize)
ax[n,l].tick_params(labelsize=ticksize)
ax[n,l].set_ylim(top=1.15)
ax[n,l].set_xlim(-15,15)
c+=1
for n in range(2):
for l in range(2):
ax[n,l].tick_params(length=6, width=1, which='major', labelsize=ticksize)
ax[n,l].tick_params(length=3, width=1, which='minor')
#a.grid(True, linewidth=1)
ax[0,0].legend(fontsize=fontsize-5)
ax[0,0].text(-19,1.1,r'(A)', fontsize=ticksize, weight='bold')
ax[0,1].text(-19,1.1,r'(B)', fontsize=ticksize, weight='bold')
ax[1,0].text(-19,1.1,r'(C)', fontsize=ticksize, weight='bold')
ax[1,1].text(-19,1.1,r'(D)', fontsize=ticksize, weight='bold')
plt.tight_layout()
name=str(envname)+'dynamicrescale.pdf'
plt.savefig(name)
inds=[5,6,7,8,9,10,11,12,13,14,15,16,17,18,19]
plotexps(allo, 'percell', inds, fontsize, ticksize, 1.89, 1.25, 0.92, 0.775, 0.49, 0.05, -0.52, -0.95, 0.12, 1.86, \
0.12, 0.91, 0.12, 0.47, 0.12, -0.54)
name=str(envname)+'varvspercell.pdf'
plt.savefig(name)
```
| github_jupyter |
```
with open('/mnt/pmldl/paracrawl-release1.en-ru.zipporah0-dedup-clean.en') as f:
eng_lines = f.readlines()
with open('/mnt/pmldl/paracrawl-release1.en-ru.zipporah0-dedup-clean.ru') as f:
ru_lines = f.readlines()
from transformers import AutoTokenizer, AutoModelForSeq2SeqLM
```
# Use pretrained model and tokenizer
```
tokenizer = AutoTokenizer.from_pretrained("Helsinki-NLP/opus-mt-ru-en")
model = AutoModelForSeq2SeqLM.from_pretrained("Helsinki-NLP/opus-mt-ru-en")
```
# Freeze encoder weights
```
for param in model.base_model.parameters():
param.requires_grad = False
```
# Split data
```
from sklearn.model_selection import train_test_split
ru_train, ru_val, eng_train, eng_val = train_test_split(ru_lines, eng_lines, test_size=.1)
ru_train, ru_val, eng_train, eng_val = ru_train[:10000], ru_val[:1000],\
eng_train[:10000], eng_val[:1000]
train_encodings = tokenizer.prepare_seq2seq_batch(ru_train, eng_train,
truncation=True,
padding=True,
max_length=100)
val_encodings = tokenizer.prepare_seq2seq_batch(ru_val, eng_val,
truncation=True,
padding=True,
max_length=100)
import torch
from torch.utils.data import Dataset
class Seq2seqDataset(Dataset):
def __init__(self, encodings):
self.encodings = encodings
def __getitem__(self, idx):
item = {key: torch.tensor(val[idx]) for key, val in self.encodings.items()}
return item
def __len__(self):
return len(self.encodings["labels"])
train_dataset = Seq2seqDataset(train_encodings)
eval_dataset = Seq2seqDataset(val_encodings)
from torch.utils.data import DataLoader
from transformers import DistilBertForSequenceClassification, AdamW
device = torch.device('cuda') if torch.cuda.is_available() else torch.device('cpu')
model.to(device)
model.train()
train_loader = DataLoader(train_dataset, batch_size=64, shuffle=True)
optim = AdamW(model.lm_head.parameters(), lr=5e-5)
import numpy as np
from tqdm.notebook import tqdm
for epoch in range(3):
epoch_loss = []
for batch in tqdm(train_loader):
optim.zero_grad()
input_ids = batch['input_ids'].to(device)
attention_mask = batch['attention_mask'].to(device)
labels = batch['labels'].to(device)
# embeddings = model.base_model(input_ids,\
# decoder_input_ids=labels,\
# attention_mask=attention_mask)/
# .requires_grad(True)
# outputs = model.lm_head(embeddings)
outputs = model(input_ids, attention_mask=attention_mask, labels=labels)
loss = outputs[0]
epoch_loss.append(loss.item())
loss.backward()
optim.step()
print(f"Epoch {epoch} finished; Loss : {np.mean(epoch_loss)}")
model.eval()
import os
experiment_name = "marian_model_3_epochs_10k_samples_no_max_length"
model.save_pretrained(os.path.join("models", experiment_name))
!nvidia-smi
```
| github_jupyter |
# PROJECT-BI Churn
# Customer Churn Analysis
Context
The leading telecom company has a massive market share but one big problem: several rivals that are constantly trying to steal customers. Because this company has been the market leader for so many years, there are not significant opportunities to grow with new customers. Instead, company executives have decided to focus on their churn: the rate at which they lose customers.
They have two teams especially interested in this data: the marketing team and the customer service team. Each team has its own reason for wanting the analysis. The marketing team wants to find out who the most likely people to churn are and create content that suits their interests. The customer service team would like to proactively reach out to customers who are about to churn, and try to encourage them to stay.
They decide to hire you for two tasks:
Help them identify the types of customers who churn
Predict who of their current customers will churn next month
To do this, they offer you a file of 7,000 customers. Each row is a customer. The Churn column will say Yes if the customer churned in the past month. The data also offers demographic data and data on the services that each customer purchases. Finally there is information on the payments those customers make.
Deliverables - What is expected
# Week 1
A presentation explaining churn for the marketing team - with links to technical aspects of your work. Tell a story to the marketing team to help them understand the customers who churn and what the marketing team can do to prevent it. Highlight the information with helpful visualizations.
How much is churn affecting the business? How big is churn compared to the existing customer base?
Explain churn by the below categories. Are there any factors that combine to be especially impactful?
Customer demographics like age and gender
Services used
Billing information
What services are typically purchased by customers who churned? Are any services especially helpful in retaining customers?
Bonus! How long will it take for the company to lose all its customers? Which demographics will they lose first?
# Data
# Data Preprocessing
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.linear_model import LinearRegression
import warnings
warnings.filterwarnings("ignore")
url="Dataset/datasets_13996_18858_WA_Fn-UseC_-Telco-Customer-Churn (2).csv"
df_bi = pd.read_csv(url)
df_bi.shape
df_bi.head()
df_bi.describe()
df_bi.dtypes
df_bi['TotalCharges'] = pd.to_numeric(df_bi['TotalCharges'],errors='coerce')
df_bi.dtypes
df_bi.isnull().sum()
df_bi['TotalCharges'].dropna(inplace=True,axis=0)
df_bi.info()
df_bi.mean()
df_bi.boxplot()
```
# 1.
# The churn of the customer affect the business cause there's 1869 of customers are churn
# Here you can see how many customers who have churned
# We can see that there are about 5174 people who who have not Churn
```
#let's count the number of people churn or not
data_churn = df_bi['Churn']
data_count = data_churn.value_counts()
data_f = data_count.to_frame()
data_f
```
# How big is churn compared to the existing customer base?
# Let's see what percentage of clients are Churn or Not
```
# How much is churn affecting the business
# Data to plot
sizes = df_bi['Churn'].value_counts(sort = True)
colors = ["#BDFCC9","grey"]
explode = (0.1,0.1)
labels= ['No','Yes']
# Plot
plt.pie(sizes,colors=colors,labels=labels,explode=explode,autopct='%1.1f%%',startangle=270,)
plt.title('Percentage of Churn ')
plt.show()
```
# The amount of churn customers is 26.5% and 73.5% are still our customers
# 2. Explain churn by the below categories.
```
age_gender= df_bi.groupby(['SeniorCitizen','gender','Churn']).size().to_frame()
age_gender
```
# Here we can see how many men are churn and we can see the men who churn and those who don't, including the women.
# Here we can see the customers who Churn by age at the next you'll see the chart
```
df_bi[['SeniorCitizen','Churn']].groupby(['Churn','SeniorCitizen']).size()/df_bi.Churn.count()
```
# Customer demographics like age and gender
# Customer who Churn and who don't Churn by Gender
```
ax = sns.countplot(x="gender", hue="Churn", data=df_bi)
ax.set_title("Customer demographics Churn by gender")
sizes = df_bi['gender'].value_counts(sort = True)
colors = ["#BDFCC9","grey"]
explode = (0.1,0.1)
labels= ['No','Yes']
# Plot
plt.pie(sizes,colors=colors,labels=labels,explode=explode,autopct='%1.1f%%',startangle=270,)
plt.title('Percentage of Churn by gender')
plt.show()
```
# Customer who Churn and who don't Churn by Age
```
ax = sns.countplot(x="SeniorCitizen", hue="Churn", data=df_bi)
ax.set_title("Customer demographics Churn by age")
```
# We gouping all our Service
```
services= df_bi.groupby(['PhoneService','MultipleLines','InternetService','OnlineSecurity','DeviceProtection','TechSupport','StreamingTV','StreamingMovies','PaperlessBilling','PaymentMethod','Churn']).size().to_frame()
services
```
# Services used by Customers who's Churn and who's not Churn
# Count all customers who's Churn or not use PhoneService
```
ax = sns.countplot(x="PhoneService", hue="Churn", data=df_bi)
ax.set_title("Customer demographics Churn by PhoneService ")
```
# Count all customers who's Churn or not use InternetService
```
ax = sns.countplot(x="InternetService", hue="Churn", data=df_bi)
ax.set_title("Customer demographics InternetService in relation to Churn or not Churn")
```
# Count all customers who's Churn or not use StreamingTV
```
ax = sns.countplot(x="StreamingTV", hue="Churn", data=df_bi)
ax.set_title("Customer demographics StreamingTV in relation to Churn or not Churn")
```
# Count all customers who's Churn or not use StreamingMovies
```
ax = sns.countplot(x="StreamingMovies", hue="Churn", data=df_bi)
ax.set_title("Customer demographics StreamingMovies in relation to Churn or not Churn")
```
# Count all customers who's Churn or not use OnlineSecurity
```
ax = sns.countplot(x="OnlineSecurity", hue="Churn", data=df_bi)
ax.set_title("Customer demographics OnlineSecurity in relation to Churn or not Churn")
```
# Count all customers who's Churn or not use OnlineBackup
```
ax = sns.countplot(x="OnlineBackup", hue="Churn", data=df_bi)
ax.set_title("Customer demographics OnlineBackup in relation to Churn or not Churn")
```
# Count all customers who's Churn or not use TechSupport
```
ax = sns.countplot(x="TechSupport", hue="Churn", data=df_bi)
ax.set_title("Customer demographics TechSupport in relation to Churn or not Churn")
```
# Count all customers who's Churn or not use MultipleLines
```
ax = sns.countplot(x="MultipleLines", hue="Churn", data=df_bi)
ax.set_title("Customer demographics MultipleLines in relation to Churn or not Churn")
```
# Count all customers who's Churn or not use DeviceProtection
```
ax = sns.countplot(x="DeviceProtection", hue="Churn", data=df_bi)
ax.set_title("Customer demographics DeviceProtection in relation to Churn or not Churn")
```
# Billing information
# The Churn Billing the compagny let's see by the information that we received by the company
```
df_bi['MonthlyCharges'].describe().to_frame()
df_bi['PaymentMethod'].describe().to_frame()
```
# We can see that the customer
```
ax = sns.countplot(x="PaymentMethod", hue="Churn", data=df_bi)
ax.set_title("Customer demographics Churn by PaymentMethod ")
ax = sns.countplot(x="PaperlessBilling", hue="Churn", data=df_bi)
ax.set_title("Customer demographics Churn by PaperlessBilling ")
df_bi['MonthlyCharges'].plot(kind='hist', figsize=(8, 5))
plt.title('Billing information Churn by MonthlyChargegs') # add a title to the histogram
plt.ylabel('Churn') # add y-label
plt.xlabel('MonthlyCharges') # add x-label
plt.show()
df_bi['tenure'].describe().to_frame()
df_bi['TotalCharges'].plot(kind='hist', figsize=(8, 5))
plt.title('Billing information Churn by TotalCharges') # add a title to the histogram
plt.ylabel('Churn') # add y-label
plt.xlabel('TotalCharges') # add x-label
plt.show()
df_bi['tenure'].plot(kind='hist', figsize=(8, 5))
plt.title('Billing information Churn by Tenure') # add a title to the histogram
plt.ylabel('Churn') # add y-label
plt.xlabel('tenure') # add x-label
plt.show()
df_bi['TotalCharges'].describe().to_frame()
df_bi['TotalCharges'].plot(kind='hist', figsize=(8, 5))
plt.title('Billing information Churn by TotalCharges') # add a title to the histogram
plt.ylabel('Churn') # add y-label
plt.xlabel('TotalCharges') # add x-label
plt.show()
```
df_plot = df_bi.groupby('tenure').Churn.mean().reset_index()
plot_data_bi = [
go.Scatter(
x=df_plot['tenure'],
y=df_plot['Churn'],
mode='markers',
name='Low',
marker= dict(size= 7,
line= dict(width=1),
color= 'blue',
opacity= 0.8
))]
plot_layout = go.Layout(
yaxis= {'title': "Churn Rate"},
xaxis= {'title': "Tenure"},
title='Tenure based Churn rate',
plot_bgcolor = "rgb(243,243,243)",
paper_bgcolor = "rgb(243,243,243)")
fig = go.Figure(data=plot_data_bi, layout=plot_layout)
pyoff.iplot(fig)
# 3. Services are typically purchased by customers who churned
```
df_bi_churn=df_bi.Churn.replace(to_replace = {'Yes' == 1, 'No' == 0}, inplace = True)
cols = df_bi.columns
cols = list(cols)
display(cols)
services_1= df_bi.groupby('Churn').mean()
services_1
val = df_bi[df_bi.Churn=='Yes']
val
plt.figure(figsize = (10,3))
sns.heatmap(val.corr(),annot =True, linewidth = 0.5,cmap ='RdBu_r')
plt.show()
services_1= df_bi.groupby('Churn').mean()
services_1
val = df_bi[df_bi.Churn=='Yes']
val
sizes = val['gender'].value_counts(sort = True)
print(sizes)
colors = ["#BDFCC9","grey"]
explode = (0.1,0.1)
labels= ['Female','Male']
# Plot
plt.pie(sizes,colors=colors,labels=labels,explode=explode,autopct='%1.1f%%',startangle=270,)
plt.title('Percentage of Churn by gender')
plt.show()
service= val.groupby(['PhoneService']).size().to_frame()
service
service= val.groupby(['MultipleLines']).size().to_frame()
service
service= val.groupby(['InternetService']).size().to_frame()
service
service= val.groupby(['OnlineSecurity']).size().to_frame()
service
service= val.groupby(['DeviceProtection']).size().to_frame()
service
service= val.groupby(['TechSupport']).size().to_frame()
service
service= val.groupby(['StreamingTV']).size().to_frame()
service
service= pd.pivot_table(val,index ='PhoneService',columns = ['gender','SeniorCitizen'] ,aggfunc ='count',values ='customerID')
service
```
# Let's see who Churn in all service by gender and SeniorCitizen
```
service= pd.pivot_table(val,index ='MultipleLines',columns = ['gender','SeniorCitizen'] ,aggfunc ='count',values ='customerID')
service
service= pd.pivot_table(val,index ='InternetService',columns = ['gender','SeniorCitizen'] ,aggfunc ='count',values ='customerID')
service
service= pd.pivot_table(val,index ='OnlineSecurity',columns = ['gender','SeniorCitizen'] ,aggfunc ='count',values ='customerID')
service
service= pd.pivot_table(val,index ='DeviceProtection',columns = ['gender','SeniorCitizen'] ,aggfunc ='count',values ='customerID')
service
service= pd.pivot_table(val,index ='TechSupport',columns = ['gender','SeniorCitizen'] ,aggfunc ='count',values ='customerID')
service
service= pd.pivot_table(val,index ='StreamingMovies',columns = ['gender','SeniorCitizen'] ,aggfunc ='count',values ='customerID')
service
service= pd.pivot_table(val,index ='StreamingTV',columns = ['gender','SeniorCitizen'] ,aggfunc ='count',values ='customerID')
service
val1 = df_bi[df_bi.Churn=='No']
val1
y_df = df_bi['TotalCharges']
y_df
# percent of customer churn used phone service
sizes = val['PhoneService'].value_counts(sort = True)
colors = ["#BDFCC9","grey"]
explode = (0.1,0.1)
labels= ['No','Yes']
# Plot
plt.pie(sizes,colors=colors,labels=labels,explode=explode,autopct='%1.1f%%',startangle=270,)
plt.title('Percentage of PhoneService ')
plt.show()
#gender who's use phone service
ax = sns.countplot(x="PhoneService",data=val)
ax.set_title("Customer demographics PhoneService ")
service1= val.groupby(['MultipleLines','gender']).size().to_frame()
service1
#gender who's used MultipleLines and left after
# leu yo gn service tel yo moins desabonne ke leu yo genyen
ax = sns.countplot(x="MultipleLines",data=val)
ax.set_title("Customer demographics MultipleLines use it and left ")
ax = sns.countplot(x="InternetService",data=val)
ax.set_title("Customer demographics InternetService by Customer who Churn only")
ax = sns.countplot(x="OnlineSecurity",data=val)
ax.set_title("Customer demographics InternetService use by Customer who churn more ")
ax = sns.countplot(x="DeviceProtection",data=val)
ax.set_title("Customer demographics InternetService use it and left ")
ax = sns.countplot(x="TechSupport",data=val)
ax.set_title("Customer demographics InternetService use it and left ")
ax = sns.countplot(x="StreamingTV",data=val)
ax.set_title("Customer demographics InternetService use it and left ")
ax = sns.countplot(x="StreamingMovies",data=val)
ax.set_title("Customer demographics InternetService use it and left ")
```
# 4 BONUS !!!
```
round(df_bi['tenure'].mean(),2)
```
| github_jupyter |
## Import Package
```
import pandas as pd
from sklearn.naive_bayes import GaussianNB, MultinomialNB
from naive_bayes_functions import train_test_split, naive_bayes_param, predict, calculate_accuracy, str_convert_float
```
## Iris Data Set (Continuous Features)
### 1 Data Preparation
```
df = pd.read_csv('data/Iris.csv', index_col=0)
train_data, test_data = train_test_split(df, 0.2)
label_column = test_data.columns[-1]
test_labels = test_data[label_column]
test_data = test_data.drop(label_column, axis=1)
train_data.head()
```
### 2 Implementation and Test of Naive Bayes
```
model = naive_bayes_param(train_data)
predict_labels = predict(model, test_data)
print(f'Accuracy of My Naive Bayes: {calculate_accuracy(predict_labels, test_labels)}')
pd.crosstab(test_labels, predict_labels, rownames=[label_column], colnames=["prediction"])
```
### 3 Compare With Sklearn Naive Bayes
```
gnb = GaussianNB()
gnb.fit(train_data.drop(label_column, axis=1), train_data[label_column])
predict_labels = gnb.predict(test_data)
print(f'Accuracy of Sklearn Naive Bayes: {calculate_accuracy(predict_labels, test_labels)}')
pd.crosstab(test_labels, predict_labels, rownames=[label_column], colnames=["prediction"])
```
## Titanic Data Set (Combination of Continuous and Discrete Features)
### 1 Data Preparation
```
df = pd.read_csv('data/Titanic.csv')
df_labels = df.Survived
label_column = 'Survived'
df = df.drop(['PassengerId', 'Survived', 'Name', 'Ticket', 'Cabin'], axis=1)
df[label_column] = df_labels
# Handling missing values
median_age = df.Age.median()
mode_embarked = df.Embarked.mode()[0]
df = df.fillna({'Age': median_age, 'Embarked': mode_embarked})
df.head()
```
### 2 Split Data Set
```
train_data, test_data = train_test_split(df, 0.1)
test_labels = test_data[label_column]
test_data = test_data.drop(label_column, axis=1)
```
### 3 Implementation and Test of Naive Bayes
```
model = naive_bayes_param(train_data)
predict_labels = predict(model, test_data)
print(f'Accuracy of My Naive Bayes: {calculate_accuracy(predict_labels, test_labels)}')
pd.crosstab(test_labels, predict_labels, rownames=[label_column], colnames=["prediction"])
```
### 4 Compare With Sklearn Naive Bayes
```
# Since sklearn doesn't seem to support mixed features
# I need to convert the str feature to number
str_convert_float(train_data)
str_convert_float(test_data)
mnb = MultinomialNB()
mnb.fit(train_data.drop(label_column, axis=1), train_data[label_column])
predict_labels = mnb.predict(test_data)
print(f'Accuracy of Sklearn Naive Bayes: {calculate_accuracy(predict_labels, test_labels)}')
pd.crosstab(test_labels, predict_labels, rownames=[label_column], colnames=["prediction"])
```
| github_jupyter |
```
import pandas as pd
import numpy as np
# Read in feature sets and corresponding outputs
# Some values of a_max were too large for a 64-bit number,
# so a 128-bit float had to be specified in order for the
# column to be parsed correctly (otherwise Pandas defaulted
# to parsing them as strings)
X1 = pd.read_csv("features_idl.csv")
X2 = pd.read_csv("features_lia.csv",dtype={'a_max':np.float128})
y1 = pd.read_csv("best_solver_idl.csv")
y2 = pd.read_csv("best_solver_lia.csv")
# Convert output values to 0 for cvc4, 1 for z3, or 2 for sat
y1 = y1.values
y1 = pd.DataFrame(np.where(y1 == "sat", 2, np.where(y1 == "z3", 1, np.where(y1 == "cvc4", 0, -1))))
y2 = y2.values
y2 = pd.DataFrame(np.where(y2 == "sat", 2, np.where(y2 == "z3", 1, np.where(y2 == "cvc4", 0, -1))))
# Verifies that there were no values in the data other than "cvc4", "z3", or "sat"
assert(not -1 in y1.values)
assert(not -1 in y2.values)
# Combine data from IDL and LIA datasets
X = pd.concat([X1,X2])
y = pd.concat([y1,y2])
from sklearn.model_selection import train_test_split
# Split datasets into
# training (60%)
# validation (20%)
# testing (20%)
X_tv, X_test, y_tv, y_test = train_test_split(X, y, test_size=0.2, random_state=1)
X_train, X_val, y_train, y_val = train_test_split(X_tv, y_tv, test_size=0.25, random_state=1)
# Combine output and features
train = pd.concat([y_train, X_train], axis=1)
val = pd.concat([y_val, X_val], axis=1)
test = pd.concat([y_test, X_test], axis=1)
train.to_csv('train.csv', index=False, header=False)
val.to_csv('validation.csv', index=False, header=False)
test.to_csv('test.csv', index=False, header=False)
import sagemaker, boto3, os
bucket = sagemaker.Session().default_bucket()
prefix = "smt-eager-vs-lazy"
# Upload datasets to S3
boto3.Session().resource('s3').Bucket(bucket).Object(
os.path.join(prefix, 'data/train.csv')).upload_file('train.csv')
boto3.Session().resource('s3').Bucket(bucket).Object(
os.path.join(prefix, 'data/validation.csv')).upload_file('validation.csv')
boto3.Session().resource('s3').Bucket(bucket).Object(
os.path.join(prefix, 'data/test.csv')).upload_file('test.csv')
region = sagemaker.Session().boto_region_name
role = sagemaker.get_execution_role()
from sagemaker.debugger import Rule, rule_configs
from sagemaker.session import TrainingInput
# Configure model
s3_output_location='s3://{}/{}/{}'.format(bucket, prefix, 'xgboost_model')
container=sagemaker.image_uris.retrieve("xgboost", region, "1.2-1")
print(container)
xgb_model=sagemaker.estimator.Estimator(
image_uri=container,
role=role,
instance_count=1,
instance_type='ml.m4.xlarge',
volume_size=5,
output_path=s3_output_location,
sagemaker_session=sagemaker.Session(),
rules=[Rule.sagemaker(rule_configs.create_xgboost_report())]
)
xgb_model.set_hyperparameters(
objective = 'multi:softprob',
num_class = 3,
num_round = 100,
subsample = 0.7,
colsample_bytree = 0.8
)
from sagemaker.session import TrainingInput
train_input = TrainingInput(
"s3://{}/{}/{}".format(bucket, prefix, "data/train.csv"), content_type="csv"
)
validation_input = TrainingInput(
"s3://{}/{}/{}".format(bucket, prefix, "data/validation.csv"), content_type="csv"
)
# Run the training job to fit the model
xgb_model.fit({"train": train_input, "validation": validation_input}, wait=True)
# Get the auto-generated analytics
rule_output_path = xgb_model.output_path + "/" + xgb_model.latest_training_job.name + "/rule-output"
! aws s3 ls {rule_output_path} --recursive
# Download the auto-generated analytics
! aws s3 cp {rule_output_path} ./ --recursive
# When done training/tuning the model, deploy an endpoint to SageMaker
import sagemaker
from sagemaker.serializers import CSVSerializer
xgb_predictor=xgb_model.deploy(
initial_instance_count=1,
instance_type='ml.t2.medium',
serializer=CSVSerializer()
)
import numpy as np
# This function calls the endpoint to get predictions
# from the model and processes the returned data
def predict_multi_class(data, num_class, rows=1000):
assert(num_class >= 2)
num_examples = data.shape[0]
split_array = np.array_split(data, int(num_examples / float(rows) + 1))
predictions = ''
for array in split_array:
predictions = ','.join([predictions, xgb_predictor.predict(array).decode('utf-8')])
# For binary classifiers, predict() returns a single float:
# the probability of a positive outcome
# formally, this means the model returns 1
if num_class == 2:
return np.fromstring(predictions[1:], sep=',')
# Convert string version of 2D array to Python list of strings
pred_list = predictions[1:].replace('[','').replace(']','').strip().split(',')
try:
assert(len(pred_list) == num_examples * num_class)
except AssertionError:
print("Something went wrong. Verify that the value of num_class is correct.")
exit()
# Convert Python list to Numpy array of floats, and reshape to 2D
return np.array(pred_list, dtype=float).reshape([num_examples,num_class])
import sklearn
# Output the accuracy of the model on the test set
log_predictions = predict_multi_class(test.to_numpy()[:,1:], 3)
predictions = np.argmax(log_predictions, axis=1)
sklearn.metrics.accuracy_score(test.iloc[:,0], predictions)
# Output the confusion matrix for the test set
cm = sklearn.metrics.confusion_matrix(test.iloc[:,0], predictions)
cm
np.count_nonzero(np.where(y_train == 0))
np.count_nonzero(np.where(y_train == 1))
np.count_nonzero(np.where(y_train == 2))
```
| github_jupyter |
# Building a Trie in Python
Before we start let us reiterate the key components of a Trie or Prefix Tree. A trie is a tree-like data structure that stores a dynamic set of strings. Tries are commonly used to facilitate operations like predictive text or autocomplete features on mobile phones or web search.
Before we move into the autocomplete function we need to create a working trie for storing strings. We will create two classes:
* A `Trie` class that contains the root node (empty string)
* A `TrieNode` class that exposes the general functionality of the Trie, like inserting a word or finding the node which represents a prefix.
Give it a try by implementing the `TrieNode` and `Trie` classes below!
```
## Represents a single node in the Trie
class TrieNode:
def __init__(self, char):
## Initialize this node in the Trie
self.char = char
self.children = {}
self.is_end = False
def insert(self, char):
## Add a child node in this Trie
if char not in self.children:
self.children[char] = TrieNode(char)
return self.children[char]
## The Trie itself containing the root node and insert/find functions
class Trie:
def __init__(self):
## Initialize this Trie (add a root node)
self.root = TrieNode("")
def insert(self, word):
## Add a word to the Trie
curr = self.root
for char in word:
curr = curr.insert(char)
curr.is_end = True
def find(self, prefix):
## Find the Trie node that represents this prefix
curr = self.root
for char in prefix:
curr = curr.children.get(char, None)
if curr is None:
return curr
return curr
```
# Finding Suffixes
Now that we have a functioning Trie, we need to add the ability to list suffixes to implement our autocomplete feature. To do that, we need to implement a new function on the `TrieNode` object that will return all complete word suffixes that exist below it in the trie. For example, if our Trie contains the words `["fun", "function", "factory"]` and we ask for suffixes from the `f` node, we would expect to receive `["un", "unction", "actory"]` back from `node.suffixes()`.
Using the code you wrote for the `TrieNode` above, try to add the suffixes function below. (Hint: recurse down the trie, collecting suffixes as you go.)
```
class TrieNode:
def __init__(self, char):
## Initialize this node in the Trie
self.char = char
self.children = {}
self.is_end = False
def insert(self, char):
## Add a child node in this Trie
if char not in self.children:
self.children[char] = TrieNode(char)
return self.children[char]
def suffixes(self, suffix = ''):
## Recursive function that collects the suffix for
## all complete words below this point
results = []
if self.is_end and suffix != '':
results.extend([suffix])
for char, node in self.children.items():
results.extend(node.suffixes(suffix + char))
return results
```
# Testing it all out
Run the following code to add some words to your trie and then use the interactive search box to see what your code returns.
```
MyTrie = Trie()
wordList = [
"ant", "anthology", "antagonist", "antonym",
"fun", "function", "factory",
"trie", "trigger", "trigonometry", "tripod"
]
for word in wordList:
MyTrie.insert(word)
from ipywidgets import widgets
from IPython.display import display
from ipywidgets import interact
def f(prefix):
if prefix != '':
prefixNode = MyTrie.find(prefix)
if prefixNode:
print('\n'.join(prefixNode.suffixes()))
else:
print(prefix + " not found")
else:
print('')
interact(f,prefix='');
```
| github_jupyter |
```
''' get the "ground truth" set of ENSPs that should be produced by funco/
cereb for a given variant, randomly extracted from a raw vcf file '''
import vcfpy
import pandas as pd
pd.set_option('display.max_rows', 999)
def trim_funco_vcf(read_path, write_path):
''' gets rid of all the nonesense cerebra doesnt pick up '''
reader = vcfpy.Reader.from_path(read_path)
writer = vcfpy.Writer.from_path(write_path, reader.header)
unwanted_class = ['COULD_NOT_DETERMINE', 'INTRON', 'FIVE_PRIME_UTR',
'THREE_PRIME_UTR', 'IGR', 'FIVE_PRIME_FLANK', 'THREE_PRIME_FLANK', 'LINCRNA']
for record in reader:
funco = record.INFO.get('FUNCOTATION')[0]
keep = True
for elm in unwanted_class:
if elm in funco:
keep = False
if keep:
writer.write_record(record)
def get_indicies(pos, sorted_gtf):
''' get overlapping indicies from the gtf file '''
keep_index_list = []
for idx, row in sorted_gtf.iterrows():
if row.start <= pos and row.end >= pos:
keep_index_list.append(idx)
return(keep_index_list)
def get_ensp_ids(indicies, sorted_gtf):
''' get the ENSP ids from the overlapping indicies in the gtf file '''
sorted_gtf_trim = sorted_gtf.iloc[indicies]
sorted_gtf_trim = sorted_gtf_trim.reset_index(drop=True)
pids = []
for idx, row in sorted_gtf_trim.iterrows():
attr = row.attribute
if 'protein_id' in attr:
pid = attr.split('protein_id')[1].split(';')[0] # this is janky
pid_strip = pid.split('"')[1]
pids.append(pid_strip)
pids_split = [x.split('.')[0] for x in pids]
ret = set(pids_split)
return(ret)
''' MAIN STARTS HERE '''
# create trimmed funco vcf file
trim_funco_vcf('../tmp/G10_1001000340_benchmark.vcf', '../tmp/G10_1001000340_funco_trimmed.vcf')
# pull out a random line
! gshuf -n 1 ../tmp/G10_1001000340_funco_trimmed.vcf
# make sure its in the raw vcf
! grep '43162920' ../tmp/G10_1001000340.vcf
# subset the gtf file by gene name
! grep 'TSPO' /Users/lincoln.harris/code/cerebra/cerebra/tmp/ref/gencode.v27.chr_patch_hapl_scaff.basic.annotation.gtf > ../tmp/gene_sub.gtf
# get PIDs for the randomized position
chrom = 22
pos = 43162920
sub_gtf = pd.read_csv('../tmp/gene_sub.gtf', sep='\t',
names=['chr', 'source', 'feature', 'start', 'end', 'score', 'strand', 'frame', 'attribute'])
sub_gtf_sorted = sub_gtf.sort_values('start')
sub_gtf_sorted = sub_gtf_sorted.reset_index(drop=True)
index_l = get_indicies(pos, sub_gtf_sorted)
ensp_l = get_ensp_ids(index_l, sub_gtf_sorted)
ensp_l = set(ensp_l)
print(ensp_l)
#///////////////////////////////////////////////////////////////////////////////////
#///////////////////////////////////////////////////////////////////////////////////
#///////////////////////////////////////////////////////////////////////////////////
# now lets find out what cereb found
# cerebra missing like a lot of these...(looking at PIDs in ensp_l)
! grep 'ENSP00000338004' ../tmp/cereb_G10_1001000340_sub.csv
# ok what does funco actually have?
# these are ENSP -> ENST converted IDs from BioMart
# ok so funco actually missing quite a few of these as well
enst_convert_list = ['ENST00000268893', 'ENST00000329563', 'ENST00000337554', 'ENST00000343736',
'ENST00000373158', 'ENST00000373161', 'ENST00000396265', 'ENST00000470917',
'ENST00000583777']
! grep 'ENST00000583777' ../tmp/G10_1001000340_benchmark.vcf
```
| github_jupyter |
# Imports
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.dates as mdates
import seaborn as sns
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.dates as mdates
import seaborn as sns
from glob import glob
from hypnospy import Wearable
from hypnospy.data import ActiwatchSleepData
from hypnospy.analysis import SleepWakeAnalysis
from hypnospy.analysis import NonWearingDetector
from hypnospy.analysis import SleepBoudaryDetector
from hypnospy.analysis import CircadianAnalysis
from hypnospy.analysis import PhysicalActivity
from hypnospy import Experiment
from hypnospy.analysis import Viewer
from tqdm import trange, tqdm
import pickle
```
# Read dataset - HCHS
```
# Configure an Experiment
exp = Experiment()
file_path = "HypnosPy-master/data/small_collection_hchs/*"
# Iterates over a set of files in a directory.
# Unfortunately, we have to do it manually with RawProcessing because we are modifying the annotations
for file in glob(file_path):
pp = ActiwatchSleepData(file, col_for_datetime="time", col_for_pid="pid")
w = Wearable(pp) # Creates a wearable from a pp object
exp.add_wearable(w)
print(w.pid)
freq = 30
exp.set_freq_in_secs(freq)
# tsp = NonWearingDetector(exp)
# tsp.fill_no_activity(-0.0001)
# tsp.detect_non_wear(strategy="choi")
# tsp.detect_non_wear(strategy="choi2011")
# tsp.check_consecutive_days(5)
# # print("Valid days:", tsp.get_valid_days())
# # print("Invalid days:", tsp.get_invalid_days())
# # strategy: "annotation", "hr", "angle"
# # sbd = SleepBoudaryDetector(exp)
# # sbd.detect_sleep_boundaries(strategy="annotation",
# # annotation_hour_to_start_search=0,
# # annotation_col='hyp_annotation',
# # output_col='hyp_sleep_period',
# # annotation_only_largest_sleep_period=False)
# tsp.invalidate_day_if_no_sleep(sleep_period_col='hyp_sleep_period')
# print("Valid days:", tsp.get_valid_days())
# tsp.check_valid_days(max_non_wear_minutes_per_day=180, min_activity_threshold=0)
# print("Valid days:", tsp.get_valid_days())
# print("Invalid days:", tsp.get_invalid_days())
```
# Use PhysicalActivity
```
pa = PhysicalActivity(exp, cutoffs=[399, 1404], names=['lpa', 'mvpa', 'vpa'])
pa.generate_pa_columns(based_on='activity')
print('cutoffs:', exp.get_all_wearables()[0].pa_cutoffs)
print('cutoffs region names:', exp.get_all_wearables()[0].pa_names)
```
# Find count of bouts per day per wearable
```
pa.get_bouts(pa_col='mvpa', length_in_minutes=10, decomposite_bouts=True)
```
# Find stats of activity per hour per day per wearable
```
pa.get_stats_pa_representation()
```
# draw physical activity within using Viewer module
```
Viewer(exp).view_signals(signal_categories=['activity'], signal_as_area=['mvpa'],
colors ={"area": ["orange"]}, alphas ={"area": 0.8},
resample_to='5T')
```
## draw MVPA boxplot
Each row represents the wearable MVPA boxplot
<br/>X-axis = 24 hour
<br/>Y-axis = MVPA box plot for 7 days
```
data = pa.get_binned_pa_representation()
data = data.reset_index()
# data = data[data['pid'] == '29881087']
g = sns.catplot(x="hyp_time_col", y="MVPA", data=data,
kind="box", color='white',
row='pid',
height=4, aspect=2.5)
```
| github_jupyter |
<table> <tr>
<td style="background-color:#ffffff;">
<a href="http://qworld.lu.lv" target="_blank"><img src="../images/qworld.jpg" width="25%" align="left"> </a></td>
<td style="background-color:#ffffff;vertical-align:bottom;text-align:right;">
prepared by <a href="http://abu.lu.lv" target="_blank">Abuzer Yakaryilmaz</a> (<a href="http://qworld.lu.lv/index.php/qlatvia/" target="_blank">QLatvia</a>)
</td>
</tr></table>
<table width="100%"><tr><td style="color:#bbbbbb;background-color:#ffffff;font-size:11px;font-style:italic;text-align:right;">This cell contains some macros. If there is a problem with displaying mathematical formulas, please run this cell to load these macros. </td></tr></table>
$ \newcommand{\bra}[1]{\langle #1|} $
$ \newcommand{\ket}[1]{|#1\rangle} $
$ \newcommand{\braket}[2]{\langle #1|#2\rangle} $
$ \newcommand{\dot}[2]{ #1 \cdot #2} $
$ \newcommand{\biginner}[2]{\left\langle #1,#2\right\rangle} $
$ \newcommand{\mymatrix}[2]{\left( \begin{array}{#1} #2\end{array} \right)} $
$ \newcommand{\myvector}[1]{\mymatrix{c}{#1}} $
$ \newcommand{\myrvector}[1]{\mymatrix{r}{#1}} $
$ \newcommand{\mypar}[1]{\left( #1 \right)} $
$ \newcommand{\mybigpar}[1]{ \Big( #1 \Big)} $
$ \newcommand{\sqrttwo}{\frac{1}{\sqrt{2}}} $
$ \newcommand{\dsqrttwo}{\dfrac{1}{\sqrt{2}}} $
$ \newcommand{\onehalf}{\frac{1}{2}} $
$ \newcommand{\donehalf}{\dfrac{1}{2}} $
$ \newcommand{\hadamard}{ \mymatrix{rr}{ \sqrttwo & \sqrttwo \\ \sqrttwo & -\sqrttwo }} $
$ \newcommand{\vzero}{\myvector{1\\0}} $
$ \newcommand{\vone}{\myvector{0\\1}} $
$ \newcommand{\stateplus}{\myvector{ \sqrttwo \\ \sqrttwo } } $
$ \newcommand{\stateminus}{ \myrvector{ \sqrttwo \\ -\sqrttwo } } $
$ \newcommand{\myarray}[2]{ \begin{array}{#1}#2\end{array}} $
$ \newcommand{\X}{ \mymatrix{cc}{0 & 1 \\ 1 & 0} } $
$ \newcommand{\I}{ \mymatrix{rr}{1 & 0 \\ 0 & 1} } $
$ \newcommand{\Z}{ \mymatrix{rr}{1 & 0 \\ 0 & -1} } $
$ \newcommand{\Htwo}{ \mymatrix{rrrr}{ \frac{1}{2} & \frac{1}{2} & \frac{1}{2} & \frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & \frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} & \frac{1}{2} } } $
$ \newcommand{\CNOT}{ \mymatrix{cccc}{1 & 0 & 0 & 0 \\ 0 & 1 & 0 & 0 \\ 0 & 0 & 0 & 1 \\ 0 & 0 & 1 & 0} } $
$ \newcommand{\norm}[1]{ \left\lVert #1 \right\rVert } $
$ \newcommand{\pstate}[1]{ \lceil \mspace{-1mu} #1 \mspace{-1.5mu} \rfloor } $
<h2> Probabilistic States </h2>
[Watch Lecture](https://youtu.be/tJjrF7WgT1g)
Suppose that Asja tosses a fair coin secretly.
Because we do not see the result, our information about the outcome will be probabilistic:
$\rightarrow$ The outcome is heads with probability $0.5$ and the outcome will be tails with probability $0.5$.
If the coin has a bias $ \dfrac{Pr(Head)}{Pr(Tail)} = \dfrac{3}{1}$, then our information about the outcome will be as follows:
$\rightarrow$ The outcome will be heads with probability $ 0.75 $ and the outcome will be tails with probability $ 0.25 $.
<i><u>Explanation</u>: The probability of getting heads is three times of the probability of getting tails.
<ul>
<li>The total probability is 1. </li>
<li> We divide 1 into four parts (three parts are for heads and one part is for tail),
<li> one part is $ \dfrac{1}{4} = 0.25$,</li>
<li> and then give three parts for heads ($0.75$) and one part for tails ($0.25$).</li>
</ul></i>
<h3> Listing probabilities as a column </h3>
We have two different outcomes: heads (0) and tails (1).
Then, we can use a column of size 2 to show the probabilities of getting heads and getting tails.
For the fair coin, our information after the coin-flip will be $ \myvector{0.5 \\ 0.5} $.
For the biased coin, it will be $ \myvector{0.75 \\ 0.25} $.
The first entry shows the probability of getting heads, and the second entry shows the probability of getting tails.
$ \myvector{0.5 \\ 0.5} $ and $ \myvector{0.75 \\ 0.25} $ are two examples of 2-dimensional (column) vectors.
<h3> Task 1 </h3>
Suppose that Balvis secretly flips a coin having the bias $ \dfrac{Pr(Heads)}{Pr(Tails)} = \dfrac{1}{4}$.
Represent your information about the outcome as a column vector.
<h3> Task 2 </h3>
Suppose that Fyodor secretly rolls a loaded (tricky) dice with the bias
$$ Pr(1):Pr(2):Pr(3):Pr(4):Pr(5):Pr(6) = 7:5:4:2:6:1 . $$
Represent your information about the result as a column vector. Remark that the size of your column should be 6.
You may use python for your calculations.
```
#
# your code is here
#
```
<a href="B12_Probabilistic_States_Solutions.ipynb#task2">click for our solution</a>
<h3> Vector representation </h3>
Suppose that we have a system with 4 distiguishable states: $ s_1 $, $s_2 $, $s_3$, and $s_4$.
We expect the system to be in one of them at any moment.
By speaking with probabilities, we say that the system is in one of the states with probability 1, and in any other state with probabilty 0.
Then, by using our column representation, we can show each state as a column vector (by using the vectors in standard basis of $ \mathbb{R}^4 $):
$$
e_1 = \myvector{1\\ 0 \\ 0 \\ 0}, e_2 = \myvector{0 \\ 1 \\ 0 \\ 0}, e_3 = \myvector{0 \\ 0 \\ 1 \\ 0},
\mbox{ and } e_4 = \myvector{0 \\ 0 \\ 0 \\ 1}.
$$
This representation helps us to represent our knowledge on a system when it is in more than one state with certain probabilities.
Remember the case in which the coins are tossed secretly.
For example, suppose that the system is in states $ s_1 $, $ s_2 $, $ s_3 $, and $ s_4 $ with probabilities $ 0.20 $, $ 0.25 $, $ 0.40 $, and $ 0.15 $, respectively.
(<i>The total probability should be 1, i.e., $ 0.20+0.25+0.40+0.15 = 1.00 $</i>)
Then, we can say that the system is in the following probabilistic state:
$$ 0.20 \cdot e_1 + 0.25 \cdot e2 + 0.40 \cdot e_3 + 0.15 \cdot e4 $$
$$ = 0.20 \cdot \myvector{1\\ 0 \\ 0 \\ 0} + 0.25 \cdot \myvector{0\\ 1 \\ 0 \\ 0} + 0.40 \cdot \myvector{0\\ 0 \\ 1 \\ 0} + 0.15 \cdot \myvector{0\\ 0 \\ 0 \\ 1}
$$
$$ = \myvector{0.20\\ 0 \\ 0 \\ 0} + \myvector{0\\ 0.25 \\ 0 \\ 0} + \myvector{0\\ 0 \\0.40 \\ 0} + \myvector{0\\ 0 \\ 0 \\ 0.15 } = \myvector{ 0.20 \\ 0.25 \\ 0.40 \\ 0.15 }, $$
where the summation of entries must be 1.
<h3> Probabilistic state </h3>
A probabilistic state is a linear combination of the vectors in the standard basis.
Here coefficients (scalars) must satisfy certain properties:
<ol>
<li> Each coefficient is non-negative </li>
<li> The summation of coefficients is 1 </li>
</ol>
Alternatively, we can say that a probabilistic state is a probability distribution over deterministic states.
We can show all information as a single mathematical object, which is called as a stochastic vector.
<i> Remark that the state of any linear system is a linear combination of the vectors in the basis. </i>
<h3> Task 3 </h3>
For a system with 4 states, randomly create a probabilistic state, and print its entries, e.g., $ 0.16~~0.17~~0.02~~0.65 $.
<i>Hint: You may pick your random numbers between 0 and 100 (or 1000), and then normalize each value by dividing the summation of all numbers.</i>
```
#
# your solution is here
#
```
<a href="B12_Probabilistic_States_Solutions.ipynb#task3">click for our solution</a>
<h3> Task 4 [extra] </h3>
As given in the hint for Task 3, you may pick your random numbers between 0 and $ 10^k $. For better precision, you may take bigger values of $ k $.
Write a function that randomly creates a probabilisitic state of size $ n $ with a precision up to $ k $ digits.
Test your function.
```
#
# your solution is here
#
```
| github_jupyter |
**[Deep Learning Course Home Page](https://www.kaggle.com/learn/deep-learning)**
---
# Exercise Introduction
We will return to the automatic rotation problem you worked on in the previous exercise. But we'll add data augmentation to improve your model.
The model specification and compilation steps don't change when you start using data augmentation. The code you've already worked with for specifying and compiling a model is in the cell below. Run it so you'll be ready to work on data augmentation.
```
from tensorflow.keras.applications import ResNet50
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Flatten, GlobalAveragePooling2D
num_classes = 2
resnet_weights_path = '../input/resnet50/resnet50_weights_tf_dim_ordering_tf_kernels_notop.h5'
my_new_model = Sequential()
my_new_model.add(ResNet50(include_top=False, pooling='avg', weights=resnet_weights_path))
my_new_model.add(Dense(num_classes, activation='softmax'))
my_new_model.layers[0].trainable = False
my_new_model.compile(optimizer='sgd', loss='categorical_crossentropy', metrics=['accuracy'])
# Set up code checking
from learntools.core import binder
binder.bind(globals())
from learntools.deep_learning.exercise_5 import *
print("Setup Complete")
```
# 1) Fit the Model Using Data Augmentation
Here is some code to set up some ImageDataGenerators. Run it, and then answer the questions below about it.
```
from tensorflow.keras.applications.resnet50 import preprocess_input
from tensorflow.keras.preprocessing.image import ImageDataGenerator
image_size = 224
# Specify the values for all arguments to data_generator_with_aug.
data_generator_with_aug = ImageDataGenerator(preprocessing_function=preprocess_input,
horizontal_flip = True,
width_shift_range = 0.1,
height_shift_range = 0.1)
data_generator_no_aug = ImageDataGenerator(preprocessing_function=preprocess_input)
```
Why do we need both a generator with augmentation and a generator without augmentation? After thinking about it, check out the solution below.
```
q_1.solution()
```
# 2) Choosing Augmentation Types
ImageDataGenerator offers many types of data augmentation. For example, one argument is `rotation_range`. This rotates each image by a random amount that can be up to whatever value you specify.
Would it be sensible to use automatic rotation for this problem? Why or why not?
```
q_2.solution()
```
# 3) Code
Fill in the missing pieces in the following code. We've supplied some boilerplate. You need to think about what ImageDataGenerator is used for each data source.
```
# Specify which type of ImageDataGenerator above is to load in training data
train_generator = data_generator_with_aug.flow_from_directory(
directory = '../input/dogs-gone-sideways/images/train',
target_size=(image_size, image_size),
batch_size=12,
class_mode='categorical')
# Specify which type of ImageDataGenerator above is to load in validation data
validation_generator = data_generator_no_aug.flow_from_directory(
directory = '../input/dogs-gone-sideways/images/val',
target_size=(image_size, image_size),
class_mode='categorical')
my_new_model.fit_generator(
train_generator, # if you don't know what argument goes first, try the hint
epochs = 3,
steps_per_epoch=19,
validation_data=validation_generator)
q_3.check()
# q_3.hint()
# q_3.solution()
```
# 4) Did Data Augmentation Help?
How could you test whether data augmentation improved your model accuracy?
```
q_4.solution()
```
# Keep Going
You are ready for **[a deeper understanding of deep learning](https://www.kaggle.com/dansbecker/a-deeper-understanding-of-deep-learning/)**.
---
**[Deep Learning Course Home Page](https://www.kaggle.com/learn/deep-learning)**
| github_jupyter |
# Interactive Plotting with Jupyter
There are several ways to interactively plot. In this tutorial I will show how to interact with 2D and 1D data. There are other ways to interact with large tables of data using either [Bokeh](https://docs.bokeh.org/en/latest/index.html) (shown the Skyfit notebook) or [Glue](http://docs.glueviz.org/en/stable). A non-python based solution that also works with large tables of data is Topcat.
Most of the methods here will work on the command line. In order to make this work within Jupyter you will need the following modules.
```
conda install -c conda-forge ipympl
conda install -c conda-forge ipywidgets
```
https://ipywidgets.readthedocs.io/
```
import sys
import astropy
import astroquery
import ipywidgets
import matplotlib
print('\n Python version: ', sys.version)
print('\n Astropy version: ', astropy.__version__)
print('\n Matplotlib version: ', matplotlib.__version__)
print('\n Astroquery version: ', astroquery.__version__)
print('\n ipywidgets version: ', ipywidgets.__version__)
import glob,os,sys
import numpy as np
import matplotlib.pyplot as plt
import astropy.io.fits as pyfits
import astropy.units as u
from astroquery.skyview import SkyView
import ipywidgets as widgets
```
Here we need an image to play with, we can either download it via SkyView or load one from our machine.
```
ext = 0
# download an image
pflist = SkyView.get_images(position='M82', survey=['SDSSr'], radius=10 * u.arcmin)
pf = pflist[0] # first element of the list, might need a loop if multiple images
# or load an image
#pf = pyfits.open('m82.fits')
image = pf[ext].data
```
Next we need to turn on the interactive plotting.
```
# turn-on interactive plots
%matplotlib widget
```
# Display an image (2D data)
We plot a 2D image using imshow, we can set the scale of the image as well as the colormap.
```
#plt.ioff()
fig = plt.figure(figsize=[6,6])
plt.ion()
p = fig.add_subplot(111)
p.imshow(image, interpolation='Nearest', origin='lower', vmin=-10, vmax=20, cmap='viridis')
plt.show()
```
# Add an event to the display
There are several types of matplotlib events that you can use to interact with a figure.
A few useful events are the following:
`button_press_event`
`button_release_event`
`key_press_event`
`key_release_event`
For more information on event handling and examples check out the following website:
https://matplotlib.org/stable/users/event_handling.html
Here we add a python function linking to link to the `key_press_event`. The function checks for the which key being pressed and if the condition is met runs its code, in this case plotting a red point on the image. We can easily add more keys adding more functionaly to our interactive figure.
```
#plt.ioff()
fig = plt.figure(figsize=[6,6])
plt.ion()
p = fig.add_subplot(111)
p.imshow(image, interpolation='Nearest', origin='lower', vmin=-10, vmax=20, cmap='viridis')
def on_key_press(event):
xc, yc = event.xdata, event.ydata
if event.key == 'm':
p.plot(xc,yc,'ro', markersize=5)
fig.canvas.draw_idle
fig.canvas.mpl_connect('key_press_event', on_key_press)
plt.show()
```
# Add output to the display with the event
If we want to display the coordinate of the points we mark, we need to use the Output widget.
```
#plt.ioff()
fig = plt.figure(figsize=[6,6])
plt.ion()
p = fig.add_subplot(111)
p.imshow(image, interpolation='Nearest', origin='lower', vmin=-10, vmax=20, cmap='viridis')
out = widgets.Output()
@out.capture()
def on_key_press(event):
xc, yc = event.xdata, event.ydata
if event.key == 'm':
p.plot(xc,yc,'ro', markersize=5)
fig.canvas.draw_idle
print("[%.1f, %.1f] = %.4f" % (xc, yc, image[int(yc),int(xc)]))
fig.canvas.mpl_connect('key_press_event', on_key_press)
display(out)
```
We can also write a Python class, this makes it more convient for dealing with multiple interactive events (i.e. keypress, mouse clicking, dragging, etc).
```
class GUI_inter:
def __init__(self,fig,img):
self.fig = fig
self.p = self.fig.gca()
self.img = img
self.display()
def display(self,sigma=20.0):
plt.clf()
self.v0 = np.mean(self.img) - sigma * np.std(self.img)
self.v1 = np.mean(self.img) + sigma * np.std(self.img)
self.p = self.fig.add_subplot(111)
self.p.imshow(self.img, interpolation='Nearest', origin='lower',
vmin=self.v0, vmax=self.v1, cmap='viridis')
plt.draw()
def on_key_press(self, event):
xc, yc = event.xdata, event.ydata
if event.key == 'm':
self.p.plot(xc,yc,'ro', markersize=5)
fig.canvas.draw_idle
print("[%.2f, %.2f]" % (xc,yc))
fig = plt.figure(figsize=[6,6])
G = GUI_inter(fig, image)
fig.canvas.mpl_connect('key_press_event', G.on_key_press)
#display(fig)
```
# Interactive 1D data
```
slice = image[150,:]
fig = plt.figure(figsize=[6,6])
p = fig.add_subplot(111)
p.plot(slice)
plt.show()
zl,xl = image.shape
fig = plt.figure(figsize=[6,6])
p = fig.add_subplot(111)
#p.set_yscale('log')
slice = image[150,:]
line, = p.plot(slice)
def update(change):
line.set_ydata(image[change.new,:])
fig.canvas.draw()
int_slider = widgets.IntSlider(
value=150,
min=0,
max=zl,
step=1,
description='Z-axis:',
continuous_update=False
)
int_slider.observe(update, 'value')
int_slider
from astroquery.sdss import SDSS
from astropy import coordinates
ra, dec = 148.969687, 69.679383
co = coordinates.SkyCoord(ra=ra, dec=dec,unit=(u.deg, u.deg), frame='fk5')
xid = SDSS.query_region(co, radius=20 * u.arcmin, spectro=True)
sp = SDSS.get_spectra(matches=xid)
print("N =",len(sp))
pf = sp[0]
ext = 1
pf[ext].header
tab = pf[ext].data
spec = tab['flux']
wave = 10**tab['loglam']
fig = plt.figure(figsize=(8,8))
ax = fig.add_subplot(111)
ax.plot(wave,spec)
ax.set_xlabel('Wavelength [Angstroms]')
ax.set_ylabel('Flux')
ext = 1
n_max = len(sp)-1 # total number of spectra - 1
pf = sp[0]
pf[ext].header
tab = pf[ext].data
spec = tab['flux']
wave = 10**tab['loglam']
fig = plt.figure(figsize=(6,6))
ax = fig.add_subplot(111)
line, = ax.plot(wave,spec)
ax.set_xlabel('Wavelength [Angstroms]')
ax.set_ylabel('Flux')
def new_spec(change):
pf = sp[change.new]
pf[ext].header
tab = pf[ext].data
spec = tab['flux']
wave = 10**tab['loglam']
line.set_xdata(wave)
line.set_ydata(spec)
fig.canvas.draw()
int_slider = widgets.IntSlider(
value=0,
min=0,
max=n_max,
step=1,
description='Spectrum:',
continuous_update=False
)
int_slider.observe(new_spec, 'value')
int_slider
ext = 1
n_max = len(sp)-1 # total number of spectra - 1
pf = sp[0]
pf[ext].header
tab = pf[ext].data
spec = tab['flux']
wave = 10**tab['loglam']
fig = plt.figure(figsize=(6,6))
ax = fig.add_subplot(111)
line, = ax.plot(wave,spec)
ax.set_xlabel('Wavelength [Angstroms]')
ax.set_ylabel('Flux')
line2, = ax.plot([6563,6563],[0,20],"--",c="r")
line2.set_visible(False)
def new_spec(change):
pf = sp[change.new]
pf[ext].header
tab = pf[ext].data
spec = tab['flux']
wave = 10**tab['loglam']
line.set_xdata(wave)
line.set_ydata(spec)
fig.canvas.draw()
def display_lines(change):
if change.new: line2.set_visible(True)
else: line2.set_visible(False)
fig.canvas.draw()
int_slider = widgets.IntSlider(
value=0,
min=0,
max=n_max,
step=1,
description='Spectrum:',
continuous_update=False
)
int_slider.observe(new_spec, 'value')
display(int_slider)
chk_box = widgets.Checkbox(
value=False,
description='Line list',
)
chk_box.observe(display_lines, 'value')
display(chk_box)
# turn-off interactive plots
%matplotlib inline
```
# Resources
https://ipywidgets.readthedocs.io/
https://ipywidgets.readthedocs.io/en/stable/examples/Widget%20List.html
https://ipywidgets.readthedocs.io/en/latest/examples/Output%20Widget.html
https://kapernikov.com/ipywidgets-with-matplotlib/
https://matplotlib.org/stable/users/event_handling.html
https://docs.bokeh.org/en/latest/index.html
http://docs.glueviz.org/en/stable
| github_jupyter |
# Titanic dataset classification, from Kaggle
Seemed like an interesting way to learn a little about classification. One thing that's nice about a competition is that you can actually look at how well your algorithm performs relative to others. How else do you findout if you're making a strong model?
```
## imports the essentials
import pandas as pd
import numpy as np
## open the dataset and explore
df = pd.read_csv('train.csv')
df.head()
print(df.shape)
## We need to impute some stuff.
print(df.isna().sum())
## Cabin
df['Cabin'].unique()
## This is weird... the passenger fares are to very high precision.
#print(df['Fare'].unique())
df['Age'].describe()
## This is weird... the passenger fares are to very high precision.
## For families, they are sums.
#print(df['Fare'].unique())
df['Age'].describe()
df[['Pclass','Fare']][df['Pclass'] == 1] ##Though correlated, these are not the same thing!
pd.Series(df['Fare'].unique()).describe()
df.sort_values(by='Ticket').head(10)
df['Ticket'].value_counts().loc['347082']
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import StandardScaler
def feature_engineer(df):
'''
All feature engineering to the dataframe is done here and returned, as a copy.
'''
working_data = df.copy()
ages = SimpleImputer(strategy='mean', add_indicator=True).fit_transform(df[['Age',]])
working_data['Age'] = ages[:,0]
working_data['Age Unknown'] = ages[:,1]
working_data['Sex'] = df['Sex'].apply(lambda x: x == 'female')
working_data['Cabin'] = df['Cabin'].isna()
working_data['Family Size'] = df['SibSp'] + df['Parch']
## The things below didn't improve the performance of my ensembles.
#working_data['Embarked'] = df['Embarked'].map({'S':0,'C':1,'Q':2}).astype('category')
#df['cabin group'] = df['Cabin'].apply(lambda x: x[0])
#working_data['Age Group'] = pd.cut(working_data['Age'], bins = [0,15,30,45,60,75,120], labels=False)
#working_data['is child'] = working_data['Age'].apply(lambda x: x < 18)
## Data exploration suggests that many passengers buy one ticket for multiple persons.
## We divid each fare by the number of people who have a matching ticket number and fill NaN with the mean.
working_data['Fare'] = working_data.apply(lambda x: x['Fare']/working_data['Ticket'].value_counts().loc[x['Ticket']], axis = 1)
working_data['Fare'].fillna(working_data['Fare'].mean(),inplace=True)
## Cut the fare data into quartitles
working_data['Fare Group'] = pd.qcut(working_data['Fare'], 4, labels=False, )
## Select the data you want to use. Based on some experimentation below.
working_data = working_data.drop(['Name','Ticket','Embarked','Family Size','Cabin','Age Unknown','Fare'],axis=1)
return working_data
working_data = feature_engineer(df)
## by playing with the matching below, we can look to see if any variable stands out
## as relevant to different groups.
temp = working_data
#temp = temp[(temp['Sex']==0)]
#temp = temp[temp['SibSp'] == 0]
print(temp.shape)
temp.corr()
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier
from sklearn.tree import DecisionTreeClassifier
X_train, X_test, y_train, y_test = train_test_split(working_data.drop(['PassengerId','Survived'], axis=1),
working_data['Survived'],
train_size = .666)
X_train.shape
X_test.shape
```
## Let's establish a baseline
A decision tree is quick and lets you know if what you've got.
```
tree_clf = DecisionTreeClassifier(max_depth = 1, min_samples_leaf=3)
simple = ['Sex']## use the feature that correlate most obviously with survival.
tree_clf.fit(X_train[simple],y_train)
print(tree_clf.score(X_train[simple],y_train))
print(tree_clf.score(X_test[simple],y_test))
forest_clf = RandomForestClassifier(n_estimators=30, max_depth = 6)
#forest_clf = RandomForestClassifier()
def clf_eval_summary(clf,X=X_test,y=y_test, X_=X_train, y_=y_train):
print("Training score: {}".format(clf.score(X_,y_)))
print("Test score: {}".format(clf.score(X,y)))
print("Confusion:")
print(confusion_matrix(y,clf.predict(X)))
## The following check whether adding a feature improves model performance.
from sklearn.metrics import confusion_matrix
diffs = []
times = 5
for i in range(times):
simple = ['Sex','Fare Group','Age']
forest_clf.fit(X_train[simple], y_train)
simpler = forest_clf.score(X_test[simple], y_test)
forest_clf.fit(X_train,y_train)
diffs.append(forest_clf.score(X_test, y_test) - simpler)
#print(diffs[-1], simpler)
#
forest_clf.fit(X_train[simple], y_train)
clf_eval_summary(forest_clf, X=X_test[simple], X_=X_train[simple])
forest_clf.fit(X_train, y_train)
clf_eval_summary(forest_clf)
print("Model improvement over a simple model: {:.4f}".format(sum(diffs)/times))
```
## Some validation and tuning of the Random Forest
```
##Cross validate the model to see how it looks.
from sklearn.model_selection import cross_val_score
cross_val_score(forest_clf,X_train,y_train).mean()
from sklearn.model_selection import GridSearchCV
parameters = {'n_estimators':[50,100],'max_depth' : [5,7,9]}
forest_search = GridSearchCV(forest_clf, param_grid=parameters, return_train_score=True)
forest_search.fit(X_train,y_train)
forest_search.cv_results_
## It's worth noting that these estimators are mostly within a standard deviation or so
## from each other in test performance. Without a bigger data set, it's hard to validate these parameters
## against each other.
best = forest_search.best_estimator_
best
```
## Other ensembles?
I tried other models. Sometimes they outperformed the Random Forest, but they never improved the kaggle test set performance.
```
from sklearn.ensemble import GradientBoostingClassifier
gradient_clf = GradientBoostingClassifier(n_estimators=100, max_depth=2)
gradient_clf.fit(X_train,y_train)
clf_eval_summary(gradient_clf)
cross_val_score(gradient_clf,X_train,y_train).mean()
from sklearn.svm import SVC
from sklearn.pipeline import Pipeline
svc = Pipeline(steps=[('scaler',StandardScaler()),('svc',SVC(C=1))])
svc.fit(X_train,y_train)
cross_val_score(svc, X_train, y_train).mean()
parameters = {'svc__C':[1,10,100],'svc__kernel':['rbf','poly']}
svc_search = GridSearchCV(svc, param_grid=parameters, return_train_score=True)
svc_search.fit(X_train,y_train)
svc_search.cv_results_
```
## This was a moment
I will let the cat out of the bag and say that a NN was the strongest model I made with only a little tuning. As you can see from the comments below, I was skeptical that a NN could outperform the other models I tried with a relatively small data set. In fact, I was so skeptical that I spent quite awhile feature engineering and tuning other models before I tried these.
```
## I'm getting annoyed that my best classifier seems to top out around .785 on Kaggle and that improvements
## on my dev set aren't translating to the test set.
## I really don't think there's enough data to support a neural net.
from sklearn.neural_network import MLPClassifier
## The parameters selected here are the results of the tuning that I did a few cells down.
mlp_clf = Pipeline(steps=[('scaler',StandardScaler()),('NN', MLPClassifier(alpha=0.01, hidden_layer_sizes=15,
learning_rate_init=0.005))])
mlp_clf.fit(X_train,y_train)
parameters = {'NN__learning_rate_init':[.005,.01], 'NN__hidden_layer_sizes': [10, 15], 'NN__alpha':[.003,.01,.03]}
mlp_grid = GridSearchCV(mlp_clf, param_grid=parameters, return_train_score=True)
mlp_grid.fit(X_train,y_train)
mlp_grid.cv_results_
mlp_grid.best_estimator_
df1 = df.copy()
results = feature_engineer(pd.read_csv('test.csv'))
print(results.isna().sum()) ## double check NaN values in the output.
mlp_clf.fit(X_train,y_train) ## makes sure the model is properly fitted.
results['Survived'] = mlp_clf.predict(results.drop('PassengerId', axis=1))
old_results = pd.read_csv('final_results.csv')
results[['PassengerId','Survived']].set_index('PassengerId').to_csv('final_results.csv')
results
np.abs((results['Survived'] - old_results['Survived'])).sum() ## check to see if the new results are much different from the old.
```
## The MLP scored .79904
As a number of people have mentioned in the discussion of the data, it's unfortunate that the leaderboard hasn't been reset. The actual Titanic data is publicly available. This means that there are lots of people with perfect scores. _But the data isn't really that powerful._
Anyway, we see below that this score places my in the 96th percentile of scores, once we get rid of obviously bad models and too good to be true ones.
```
leaders = pd.read_csv('titanic-publicleaderboard.csv')
leaders = leaders[(leaders['Score'] < .95) & (leaders['Score'] > .7)].sort_values(by='Score')
leaders['Rank'] = leaders['Score'].rank(method='first', ascending=False)
leaders['Percentile'] = (leaders.shape[0]-leaders['Rank'])/leaders.shape[0]
leaders.head()
leaders[leaders['TeamName'] == 'sjlenhart']
```
| github_jupyter |
# A Brief Introduction to NumPy
### "...the fundamental package for scientific computing with Python." - numpy.org
In this notebook, we will cover the basics of NumPy, a package that is the basis for many other libraries in the data science ecosystem. Let's get started.
```
import numpy as np
from IPython.display import Image
import time
from sys import getsizeof
import matplotlib.pyplot
```
# 1. NumPy Arrays
The array data structure is the backbone of the NumPy library. They can be single-dimensional (vectors), two-dimensional (matrices), or multi-dimensional for more complex tasks.
In many ways, they are similar to Python lists.
```
a = ['a', 'b', 'c', 'd', 'e', 'f']
b = np.array(['a', 'b', 'c', 'd', 'e', 'f'])
# Accessible by index
print(b[0])
# Sliceable
print(b[1:3])
# Iterable
for letter in b:
print(letter)
```
So why use NumPy arrays at all? One word: performance! Generally speaking, Python lists take up more space and require more computation than NumPy arrays. Let's take a look at the size differences.
```
n_elements = 1_000_000
# Create using list comprehension
python_list = [x for x in range(n_elements)]
print(getsizeof(python_list))
# Create with existing python list
np_arr = np.array(python_list)
print(getsizeof(np_arr))
```
Now let's look at the speed differences.
```
start = time.process_time()
# Add 100 to every element in the Python list
python_list_mod = [x + 100 for x in python_list]
python_time = time.process_time() - start
print(python_time)
# Add 100 to every element in the Numpy array
start = time.process_time()
np_arr_mod = np_arr + 100
np_time = time.process_time() - start
print(np_time)
Image('python_memory1.png')
```
If NumPy arrays are more efficient computationally and in regards to space, why not use them all the time? There are some constraints, most notably, all of their items must be of the same type. [NumPy Array](https://numpy.org/doc/stable/reference/generated/numpy.array.html#numpy.array)
```
python_list = [1, 'a', 0.222, 'hello from inside the list!']
np_arr = np.array(python_list)
print(python_list)
print(np_arr)
```
# 1.1 Creating
NumPy arrays are created with existing data (standard python lists or lists of lists) or by using a collection of built-in methods.
## 1.1.1 Existing Data
### .array()
Use python lists (or lists of lists) as input.
```
std_list = [1, 2, 3, 4, 5]
np.array(std_list)
std_matrix = [[1, 2, 3], [4, 5, 6], [7, 8, 9]]
np.array(std_matrix)
```
Note: the .array() method is a convenience function for constructing objects of the class ndarray. While it is possible to call .ndarray() directly, it is specifically regarded as an anti-pattern by the NumPy documentation.
## 1.1.2 Fixed Values
### .zeros(), .ones()
Return a new array of given shape and type, filled with zeros or ones.
```
np.zeros(3)
np.ones(17)
# Notice the shape represented as a tuple
np.zeros((5,5))
```
Note: the numbers have periods after them to indicate that these are floating point numbers.
### .full()
Return a new array of given shape and type, filled with fill_value.
```
np.full((4,4), 72)
```
## 1.1.3 Range
### .arange()
Return evenly spaced values within a given interval. Notice that the output is inclusive of the first number parameter and exclusive of the second.
```
np.arange(0,10)
np.arange(0,10,2)
np.arange(0,20,5)
```
### .linspace()
Return evenly spaced numbers over a specified interval. Notice that the output is inclusive of both the first and second number parameters.
```
np.linspace(0,20,5)
np.linspace(0,1, 20)
```
Note: the main difference between .linspace() and .arange() is that with .linspace() you have precise control over the end value, whereas with .arange() you can specify the increments explicitly.
### .logspace()
Return numbers spaced evenly on a log scale.
```
np.logspace(2.0, 3.0, num=4)
np.logspace(2.0, 3.0, num=4, base=2.0)
```
## 1.1.4 Random
### .rand()
Create an array of the given shape and populate it with random samples from a uniform distribution over 0,1.
```
np.random.rand()
np.random.rand(4)
np.random.rand(7,3)
```
### .randn()
Return a sample (or samples) from the “standard normal” distribution.
```
np.random.randn()
np.random.randn(5)
```
Note: .rand() is from a uniform distribution, whereas .randn() is from the standard **normal** distribution.
```
np.random.randn(3, 5)
```
### .randint()
Return random integers from low (inclusive) to high (exclusive).
```
np.random.randint(0,10)
np.random.randint(0,10,size=7)
np.random.randint(0,10,size=(3,2))
```
## 1.2 Attributes and Methods
```
arr = np.random.randint(0,10,size=(2,7))
arr
```
### .shape
Tuple of array dimensions. Note: this is an attribute NOT a method.
```
arr.shape
```
### .reshape()
Gives a new shape to an array without changing its data. Note: this happens 'in place' (does not return new values).
```
arr.reshape(7,2)
arr.reshape(12,4)
```
### .newaxis
Alternate syntax.
```
arr[np.newaxis, :]
```
### .dtype
The type of data in the array.
```
arr.dtype
```
### .astype()
Casts values to a specified type.
```
arr.astype('int8')
# Complex numbers are the combination of a real and an imaginary number.
arr.astype('complex')
```
Note: in mathematics i is used to denote imaginary numbers, but in Python (and many other languages) j is used because i tends to indicate the current value in a system.
```
arr2 = np.full((4,4), 199)
arr2.astype('bool')
```
## 1.3 Indexing
### 1.3.1 One-dimensional
```
arr_1d = np.arange(0,21, 2)
arr_1d
# Get the value at index 5 (the sixth element)
arr_1d[5]
# Get a slice of the array from index 1 (inclusive) to index 5 (exclusive)
arr_1d[1:5]
# Get a slice of the array from index 4 to the end
arr_1d[4:]
# Get the last element in the array
arr_1d[-1]
# Reverse the array
arr_1d[::-1]
```
### 1.3.2 Two-dimensional
```
arr_2d = np.arange(12).reshape((3, 4))
arr_2d
# Get the first row
arr_2d[0]
# Get the second element of the second row
arr_2d[1][1]
# Alternative syntax
arr_2d[1,1]
# Get first and second rows
arr_2d[:2]
# Get first element of both first and second rows
arr_2d[:2,0]
# Maintain shape
arr_2d[:2,0:1]
```
### 1.3.3 Fancy Indexing
```
[arr_1d[2], arr_1d[3], arr_1d[7], arr_1d[8]]
arr_1d[[2,3,7,8]]
# Output of fancy indexing
ind = np.array([[2, 3],
[7, 8]])
arr_1d[ind]
row = [0, 1]
col = [2, 3]
arr_2d[(row, col)]
# Notice if second row value is not provided, NumPy compensates
row = [0]
col = [2, 3]
arr_2d[(row, col)]
```
## 1.4 Selection
```
arr = np.arange(1,5)
arr
bool_arr = np.array([False, False, True, True])
bool_arr
arr[bool_arr]
arr_long = np.arange(0,20)
arr_long
bool_arr_long = arr_long > 11
arr_long[bool_arr_long]
arr_long[arr_long < 5]
## TODO add exercise
```
# 2. Operations
One of the most powerful features of NumPy arrays is that operations are vectorized.
## 2.1 Arithmetic
Arithmetic operations work on NumPy arrays.
### 2.1.1 One-dimensional
```
arr_1d = np.arange(0,11)
arr_1d
arr_1d + 5
arr_1d - 12
arr_1d * 2
arr_1d / 4
arr_1d // 4
arr_1d ** 3
```
### 2.1.2 Two-dimensional
```
arr_2d = np.arange(15).reshape((3,5))
arr_2d
arr_2d * 2
arr_2d ** 2
```
### 2.1.3 Multiple values
```
arr_mult = np.array([1,2,3,4,5])
arr_mult
arr_2d * arr_mult
arr_mult_2 = np.array([1,2,3])
```
## 2.2 Ufuncs
Universal functions. For more information, visit: https://docs.scipy.org/doc/numpy/reference/ufuncs.html
```
arr = np.arange(1,11)
arr
```
### .sum()
Sum of array elements over a given axis.
```
np.sum(arr)
```
### .sqrt()
Return the non-negative square-root of an array, element-wise.
```
np.sqrt(9)
np.sqrt(arr)
```
### .power()
First array elements raised to powers from second array, element-wise.
```
np.power(3,2)
np.power(arr, 2)
```
### .min(), .max()
```
arr = np.random.randint(0,10, 10)
arr
np.min(arr)
arr.min()
np.max(arr)
arr.max()
```
## 2.3 Broadcasting
The term broadcasting describes how numpy treats arrays with different shapes during arithmetic operations. Subject to certain constraints, the smaller array is “broadcast” across the larger array so that they have compatible shapes. (NumPy documentation)
```
arr_one = np.arange(0,20).reshape(4,5)
arr_one
arr_one + 10
arr_one + np.array([10])
arr_one + np.array([10, 20])
arr_two = np.array([10,20,30,40,50])
arr_one + arr_two
arr_one.shape
arr_two.shape
arr_two.reshape(5,1)
arr_one + arr_two
```
Array comparison begins with the trailing dimensions and subsequently works its way foward. Two array dimensions are compatible when:
- they are equal, or
- one of them is 1
(NumPy documentation)
https://jakevdp.github.io/PythonDataScienceHandbook/02.05-computation-on-arrays-broadcasting.html
```
# Plot sin and cos on the same graph, using matplotlib
# Compute the x and y coordinates for points on sine and cosine curves
x = np.arange(0, 5 * np.pi, 0.1)
y_sin = np.sin(x)
y_cos = np.cos(x)
# Set up a subplot grid that has height 2 and width 1,
# and set the first such subplot as active.
plt.subplot(2, 1, 1)
# Make the first plot
plt.plot(x, y_sin)
plt.title('Sine')
# Set the second subplot as active, and make the second plot.
plt.subplot(2, 1, 2)
plt.plot(x, y_cos)
plt.title('Cosine')
# Ensure tight layout
plt.tight_layout()
# Show the figure.
plt.show()
```
| github_jupyter |
# LetsGrowMore
## ***Virtual Internship Program***
***Data Science Tasks***
### ***Author: SARAVANAVEL***
# ***ADVANCED LEVEL TASK***
### Task 9 -Handwritten equation solver using CNN
Simple Mathematical equation solver using character and symbol regonition using image processing and CNN
## 1. Import Libraries/Packages
```
import numpy as np
import pandas as pd
import cv2
import tensorflow as tf
import matplotlib.pyplot as plt
from imutils.contours import sort_contours
import imutils
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Conv2D
from tensorflow.keras.layers import MaxPooling2D
from tensorflow.keras.layers import Dense
from tensorflow.keras.layers import Flatten
from tensorflow.keras.optimizers import SGD
from tensorflow.keras.preprocessing.image import ImageDataGenerator
import os
```
## Data preprocessing
```
print(os.listdir("./input")) #without extracting the data.rar file
```
## Data Augementation
```
train_datagen = ImageDataGenerator(
rescale = 1./255,
shear_range = 0.2,
zoom_range = 0.2,
validation_split = 0.25
)
data_path='./input/extracted_images'
train_set = train_datagen.flow_from_directory(
data_path,
target_size = (40, 40),
color_mode = 'grayscale',
batch_size = 32,
class_mode = 'categorical',
shuffle = True,
subset='training',
seed = 123
)
valid_set = train_datagen.flow_from_directory(
data_path,
target_size = (40, 40),
color_mode = 'grayscale',
batch_size = 32,
class_mode = 'categorical',
shuffle = True,
subset='validation',
seed = 123
)
```
## Model Building
```
model = Sequential()
model.add(Conv2D(32, (3, 3), activation='relu', input_shape=(40, 40, 1)))
model.add(MaxPooling2D((2, 2)))
model.add(Conv2D(64, (3, 3), activation='relu'))
model.add(MaxPooling2D((2, 2)))
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(Dense(18, activation='softmax'))
# compile model
adam = tf.keras.optimizers.Adam(learning_rate = 5e-4)
model.compile(optimizer=adam, loss='categorical_crossentropy', metrics=['accuracy'])
```
## Model Training
```
history=model.fit(train_set,
validation_data=valid_set,
epochs=1,
verbose=1)
```
## Model evaluation
```
val_loss, val_accuracy = model.evaluate(valid_set)
print(val_loss,val_accuracy)
train_set.class_indices
```
print('\n',train_set.class_indices, sep = "\n")
```
label_map = (train_set.class_indices)
label_map
def prediction(img):
#img = cv2.imread(img, cv2.IMREAD_GRAYSCALE)
plt.imshow(img, cmap = 'gray')
img = cv2.resize(img,(40, 40))
norm_image = cv2.normalize(img, None, alpha = 0, beta = 1, norm_type = cv2.NORM_MINMAX, dtype = cv2.CV_32F)
#norm_image=img/255
norm_image = norm_image.reshape((norm_image.shape[0], norm_image.shape[1], 1))
case = np.asarray([norm_image])
pred = (model.predict_classes([case]))
return ([i for i in train_set.class_indices if train_set.class_indices[i]==(pred[0])][0],pred)
image = cv2.imread('./input/data-eqns/test_image1.jpg')
gray = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)
blurred = cv2.GaussianBlur(gray, (5, 5), 0)
# perform edge detection, find contours in the edge map, and sort the
# resulting contours from left-to-right
edged = cv2.Canny(blurred, 30, 150)
cnts = cv2.findContours(edged.copy(), cv2.RETR_EXTERNAL,cv2.CHAIN_APPROX_SIMPLE)
cnts = imutils.grab_contours(cnts)
cnts = sort_contours(cnts, method="left-to-right")[0]
chars=[]
for c in cnts:
# compute the bounding box of the contour
(x, y, w, h) = cv2.boundingRect(c)
# filter out bounding boxes, ensuring they are neither too small
# nor too large
if w*h>1200:
# extract the character and threshold it to make the character
# appear as *white* (foreground) on a *black* background, then
# grab the width and height of the thresholded image
roi = gray[y:y + h, x:x + w]
chars.append(prediction(roi))
cv2.rectangle(image, (x, y), (x + w, y + h), (0, 255, 0), 2)
plt.figure(figsize=(20,20))
plt.imshow(image)
chars
labels=[i for i in train_set.class_indices]
print(labels)
eq=[]
pos=[]
for i in ((chars)):
if len(eq)==0 and i[0][0] in labels[3:]:
eq.append(i[0][0])
elif len(eq)>0 and i[0][0] in labels[4:14]:
eq.append(i[0][0])
elif len(eq)>0 and i[0][0] in labels[:4]:
eq.append(i[0][0])
pos.append(len(eq))
else:
pass
for i in pos:
if eq[i-1]=='+':
print(int(''.join(eq[:pos[0]-1]))+int(''.join(eq[pos[0]:])))
elif eq[i-1]=='%':
print(int(''.join(eq[:pos[0]-1]))/int(''.join(eq[pos[0]:])))
elif eq[i-1]=='*':
print(int(''.join(eq[:pos[0]-1]))*int(''.join(eq[pos[0]:])))
else:
print(int(''.join(eq[:pos[0]-1]))-int(''.join(eq[pos[0]:])))
image = cv2.imread('./input/data-eqns/test0.png')
gray = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)
blurred = cv2.GaussianBlur(gray, (5, 5), 0)
# perform edge detection, find contours in the edge map, and sort the
# resulting contours from left-to-right
edged = cv2.Canny(blurred, 30, 150)
cnts = cv2.findContours(edged.copy(), cv2.RETR_EXTERNAL,cv2.CHAIN_APPROX_SIMPLE)
cnts = imutils.grab_contours(cnts)
cnts = sort_contours(cnts, method="left-to-right")[0]
chars=[]
for c in cnts:
# compute the bounding box of the contour
(x, y, w, h) = cv2.boundingRect(c)
# filter out bounding boxes, ensuring they are neither too small
# nor too large
if w*h>1200:
# extract the character and threshold it to make the character
# appear as *white* (foreground) on a *black* background, then
# grab the width and height of the thresholded image
roi = gray[y:y + h, x:x + w]
chars.append(prediction(roi))
cv2.rectangle(image, (x, y), (x + w, y + h), (0, 255, 0), 2)
plt.figure(figsize=(20,20))
plt.imshow(image)
chars
labels=[i for i in train_set.class_indices]
print(labels)
eq=[]
pos=[]
for i in ((chars)):
if len(eq)==0 and i[0][0] in labels[3:]:
eq.append(i[0][0])
elif len(eq)>0 and i[0][0] in labels[4:14]:
eq.append(i[0][0])
elif len(eq)>0 and i[0][0] in labels[:4]:
eq.append(i[0][0])
pos.append(len(eq))
else:
pass
for i in pos:
if eq[i-1]=='+':
print(int(''.join(eq[:pos[0]-1]))+int(''.join(eq[pos[0]:])))
elif eq[i-1]=='%':
print(int(''.join(eq[:pos[0]-1]))/int(''.join(eq[pos[0]:])))
elif eq[i-1]=='*':
print(int(''.join(eq[:pos[0]-1]))*int(''.join(eq[pos[0]:])))
else:
print(int(''.join(eq[:pos[0]-1]))-int(''.join(eq[pos[0]:])))
```
# THANK YOU!!
| github_jupyter |
<i>Copyright (c) Microsoft Corporation. All rights reserved.<br>
Licensed under the MIT License.</i>
<br>
# Recommender Hyperparameter Tuning w/ AzureML
This notebook shows how to auto-tune hyperparameters of a recommender model by utilizing **Azure Machine Learning service** ([AzureML](https://azure.microsoft.com/en-us/services/machine-learning-service/))<sup><a href="#azureml-search">a</a>, <a href="#azure-subscription">b</a></sup>.
We present an overall process of utilizing AzureML, specifically [**Hyperdrive**](https://docs.microsoft.com/en-us/python/api/azureml-train-core/azureml.train.hyperdrive?view=azure-ml-py) component, for the hyperparameter tuning by demonstrating key steps:
1. Configure AzureML Workspace
2. Create Remote Compute Target (GPU cluster)
3. Prepare Data
4. Prepare Training Scripts
5. Setup and Run Hyperdrive Experiment
6. Model Import, Re-train and Test
In this notebook, we use [**Wide-and-Deep model**](https://ai.googleblog.com/2016/06/wide-deep-learning-better-together-with.html) from **TensorFlow high-level Estimator API (v1.12)** on the movie recommendation scenario. Wide-and-Deep learning jointly trains wide linear model and deep neural networks (DNN) to combine the benefits of memorization and generalization for recommender systems.
For more details about the **Wide-and-Deep** model:
* [Wide-Deep Quickstart notebook](../00_quick_start/wide_deep_movielens.ipynb)
* [Original paper](https://arxiv.org/abs/1606.07792)
* [TensorFlow API doc](https://www.tensorflow.org/api_docs/python/tf/estimator/DNNLinearCombinedRegressor)
Regarding **AuzreML**, please refer:
* [Quickstart notebook](https://docs.microsoft.com/en-us/azure/machine-learning/service/quickstart-create-workspace-with-python)
* [Hyperdrive](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-tune-hyperparameters)
* [Tensorflow model tuning with Hyperdrive](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-train-tensorflow)
---
<sub><span id="azureml-search">a. To use AzureML, you will need an Azure subscription.</span><br>
<span id="azure-subscription">b. When you web-search "Azure Machine Learning", you will most likely to see mixed results of Azure Machine Learning (AzureML) and Azure Machine Learning **Studio**. Please note they are different services where AzureML's focuses are on ML model management, tracking and hyperparameter tuning, while the [ML Studio](https://studio.azureml.net/)'s is to provide a high-level tool for 'easy-to-use' experience of ML designing and experimentation based on GUI.</span></sub>
```
import sys
sys.path.append("../../")
import itertools
import os
import shutil
from tempfile import TemporaryDirectory
import time
from IPython.display import clear_output
import numpy as np
import papermill as pm
import pandas as pd
import sklearn.preprocessing
import tensorflow as tf
import azureml as aml
import azureml.widgets as widgets
import azureml.train.hyperdrive as hd
from reco_utils.dataset.pandas_df_utils import user_item_pairs
from reco_utils.dataset import movielens
from reco_utils.dataset.python_splitters import python_random_split
import reco_utils.evaluation.python_evaluation
print("Azure ML SDK Version:", aml.core.VERSION)
print("Tensorflow Version:", tf.__version__)
tmp_dir = TemporaryDirectory()
```
### 1. Configure AzureML Workspace
**AzureML workspace** is a foundational block in the cloud that you use to experiment, train, and deploy machine learning models via AzureML service. In this notebook, we 1) create a workspace from [**Azure portal**](https://portal.azure.com) and 2) configure from this notebook.
You can find more details about the setup and configure processes from the following links:
* [Quickstart with Azure portal](https://docs.microsoft.com/en-us/azure/machine-learning/service/quickstart-get-started)
* [Quickstart with Python SDK](https://docs.microsoft.com/en-us/azure/machine-learning/service/quickstart-create-workspace-with-python)
#### 1.1 Create a workspace
1. Sign in to the [Azure portal](https://portal.azure.com) by using the credentials for the Azure subscription you use.
2. Select **Create a resource** menu, search for **Machine Learning service workspace** select **Create** button.
3. In the **ML service workspace** pane, configure your workspace with entering the *workspace name* and *resource group* (or **create new** resource group if you don't have one already), and select **Create**. It can take a few moments to create the workspace.
#### 1.2 Configure
To configure this notebook to communicate with the workspace, type in your Azure subscription id, the resource group name and workspace name to `<subscription-id>`, `<resource-group>`, `<workspace-name>` in the above notebook cell. Alternatively, you can create a *.\aml_config\config.json* file with the following contents:
```
{
"subscription_id": "<subscription-id>",
"resource_group": "<resource-group>",
"workspace_name": "<workspace-name>"
}
```
```
# AzureML workspace info. Note, will look up "aml_config\config.json" first, then fall back to use this
SUBSCRIPTION_ID = '<subscription-id>'
RESOURCE_GROUP = '<resource-group>'
WORKSPACE_NAME = '<workspace-name>'
# Remote compute (cluster) configuration. If you want to save the cost more, set these to small.
VM_SIZE = 'STANDARD_NC6'
VM_PRIORITY = 'lowpriority'
# Cluster nodes
MIN_NODES = 4
MAX_NODES = 8
# Hyperdrive experimentation configuration
MAX_TOTAL_RUNS = 100 # Number of runs (training-and-evaluation) to search the best hyperparameters.
MAX_CONCURRENT_RUNS = 8
# Recommend top k items
TOP_K = 10
# Select MovieLens data size: 100k, 1m, 10m, or 20m
MOVIELENS_DATA_SIZE = '100k'
EPOCHS = 50
# Metrics to track
RANKING_METRICS = ['ndcg_at_k', 'precision_at_k']
RATING_METRICS = ['rmse', 'mae']
PRIMARY_METRIC = 'rmse'
# Data column names
USER_COL = 'UserId'
ITEM_COL = 'MovieId'
RATING_COL = 'Rating'
ITEM_FEAT_COL = 'Genres'
```
Now let's see if everything is ready!
```
# Connect to a workspace
try:
ws = aml.core.Workspace.from_config()
except aml.exceptions.UserErrorException:
try:
ws = aml.core.Workspace(
subscription_id=SUBSCRIPTION_ID,
resource_group=RESOURCE_GROUP,
workspace_name=WORKSPACE_NAME
)
ws.write_config()
except aml.exceptions.AuthenticationException:
ws = None
if ws is None:
raise ValueError(
"""Cannot access the AzureML workspace w/ the config info provided.
Please check if you entered the correct id, group name and workspace name"""
)
else:
print("AzureML workspace name: ", ws.name)
clear_output() # Comment out this if you want to see your workspace info.
```
### 2. Create Remote Compute Target
We create a GPU cluster as our **remote compute target**. If a cluster with the same name is already exist in your workspace, the script will load it instead. You can see [this document](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-set-up-training-targets) to learn more about setting up a compute target on different locations.
This notebook selects **STANDARD_NC6** virtual machine (VM) and sets it's priority as *lowpriority* to save the cost.
Size | vCPU | Memory (GiB) | Temp storage (SSD, GiB) | GPU | GPU memory (GiB) | Max data disks | Max NICs
---|---|---|---|---|---|---|---
Standard_NC6 | <div align="center">6</div> | <div align="center">56</div> | <div align="center">340</div> | <div align="center">1</div> | <div align="center">8</div> | <div align="center">24</div> | <div align="center">1</div>
For more information about Azure virtual machine sizes, see [here](https://docs.microsoft.com/en-us/azure/virtual-machines/windows/sizes-gpu).
```
CLUSTER_NAME = 'gpu-cluster-nc6'
try:
compute_target = aml.core.compute.ComputeTarget(workspace=ws, name=CLUSTER_NAME)
print("Found existing compute target")
except aml.core.compute_target.ComputeTargetException:
print("Creating a new compute target...")
compute_config = aml.core.compute.AmlCompute.provisioning_configuration(
vm_size=VM_SIZE,
vm_priority=VM_PRIORITY,
min_nodes=MIN_NODES,
max_nodes=MAX_NODES
)
# create the cluster
compute_target = aml.core.compute.ComputeTarget.create(ws, CLUSTER_NAME, compute_config)
compute_target.wait_for_completion(show_output=True, min_node_count=None, timeout_in_minutes=20)
# Use the 'status' property to get a detailed status for the current cluster.
print(compute_target.status.serialize())
```
### 3. Prepare Data
For demonstration purpose, we use 100k MovieLens dataset. First, download the data and convert the format (multi-hot encode *genres*) to make it work for our model. More details about this step is described in our [Wide-Deep Quickstart notebook](../00_quick_start/wide_deep_movielens.ipynb).
```
data = movielens.load_pandas_df(
size=MOVIELENS_DATA_SIZE,
header=[USER_COL, ITEM_COL, RATING_COL],
genres_col='Genres_string'
)
# Encode 'genres' into int array (multi-hot representation) to use as item features
genres_encoder = sklearn.preprocessing.MultiLabelBinarizer()
data[ITEM_FEAT_COL] = genres_encoder.fit_transform(
data['Genres_string'].apply(lambda s: s.split("|"))
).tolist()
data.drop('Genres_string', axis=1, inplace=True)
data.head()
```
The dataset is split into train, validation, and test sets. The train and validation sets will be used for hyperparameter tuning, and the test set will be used for the final evaluation of the model after we import the best model from AzureML workspace.
Here, we don't use multiple-split directly by passing `ratio=[0.56, 0.19, 0.25]`. Instead, we first split the data into train and test sets with the same `seed` we've been using in other notebooks to make the train set identical across them. Then, we further split the train set into train and validation sets.
```
# Use the same seed to make the train and test sets identical across other notebooks in the repo.
train, test = python_random_split(data, ratio=0.75, seed=42)
# Further split the train set into train and validation set.
train, valid = python_random_split(train)
print(len(train), len(valid), len(test))
```
Now, upload the train and validation sets to the AzureML workspace. Our Hyperdrivce experiment will use them.
```
DATA_DIR = os.path.join(tmp_dir.name, 'aml_data')
os.makedirs(DATA_DIR, exist_ok=True)
TRAIN_FILE_NAME = "movielens_" + MOVIELENS_DATA_SIZE + "_train.pkl"
train.to_pickle(os.path.join(DATA_DIR, TRAIN_FILE_NAME))
VALID_FILE_NAME = "movielens_" + MOVIELENS_DATA_SIZE + "_valid.pkl"
valid.to_pickle(os.path.join(DATA_DIR, VALID_FILE_NAME))
# Note, all the files under DATA_DIR will be uploaded to the data store
ds = ws.get_default_datastore()
ds.upload(
src_dir=DATA_DIR,
target_path='data',
overwrite=True,
show_progress=True
)
```
### 4. Prepare Training Scripts
Next step is to prepare scripts that AzureML Hyperdrive will use to train and evaluate models with selected hyperparameters. We re-use our [Wide-Deep Quickstart notebook](../00_quick_start/wide_deep_movielens.ipynb) for that. To run the model notebook from the Hyperdrive Run, all we need is to prepare an [entry script](../../reco_utils/azureml/wide_deep.py) which parses the hyperparameter arguments, passes them to the notebook, and records the results of the notebook to AzureML Run logs by using `papermill`. Hyperdrive uses the logs to track the performance of each hyperparameter-set and finds the best performed one.
Here is a code snippet from the entry script:
```
...
from azureml.core import Run
run = Run.get_context()
...
NOTEBOOK_NAME = os.path.join(
"notebooks",
"00_quick_start",
"wide_deep_movielens.ipynb"
)
...
parser = argparse.ArgumentParser()
...
parser.add_argument('--dnn-optimizer', type=str, dest='dnn_optimizer', ...
parser.add_argument('--dnn-optimizer-lr', type=float, dest='dnn_optimizer_lr', ...
...
pm.execute_notebook(
NOTEBOOK_NAME,
OUTPUT_NOTEBOOK,
parameters=params,
kernel_name='python3',
)
...
```
```
# Prepare all the necessary scripts which will be loaded to our Hyperdrive Experiment Run
SCRIPT_DIR = os.path.join(tmp_dir.name, 'aml_script')
# Copy scripts to SCRIPT_DIR temporarly
shutil.copytree(os.path.join('..', '..', 'reco_utils'), os.path.join(SCRIPT_DIR, 'reco_utils'))
# We re-use our model notebook for training and testing models.
model_notebook_dir = os.path.join('notebooks', '00_quick_start')
dest_model_notebook_dir = os.path.join(SCRIPT_DIR, model_notebook_dir)
os.makedirs(dest_model_notebook_dir , exist_ok=True)
shutil.copy(
os.path.join('..', '..', model_notebook_dir, 'wide_deep_movielens.ipynb'),
dest_model_notebook_dir
)
# This is our entry script for Hyperdrive Run
ENTRY_SCRIPT_NAME = 'reco_utils/azureml/wide_deep.py'
```
### 5. Setup and Run Hyperdrive Experiment
[Hyperdrive](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-tune-hyperparameters) create a machine learning Experiment [Run](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.run?view=azure-ml-py) on the workspace and utilizes child-runs to search the best set of hyperparameters.
#### 5.1 Create Experiment
[Experiment](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.experiment(class)?view=azure-ml-py) is the main entry point into experimenting with AzureML. To create new Experiment or get the existing one, we pass our experimentation name.
```
# Create an experiment to track the runs in the workspace
EXP_NAME = "movielens_" + MOVIELENS_DATA_SIZE + "_wide_deep_model"
exp = aml.core.Experiment(workspace=ws, name=EXP_NAME)
```
#### 5.2 Define Search Space
Now we define the search space of hyperparameters. For example, if you want to test different batch sizes of {64, 128, 256}, you can use `azureml.train.hyperdrive.choice(64, 128, 256)`. To search from a continuous space, use `uniform(start, end)`. For more options, see [Hyperdrive parameter expressions](https://docs.microsoft.com/en-us/python/api/azureml-train-core/azureml.train.hyperdrive.parameter_expressions?view=azure-ml-py).
In this notebook, we fix model type as `wide_deep` and the number of epochs to 50.
In the search space, we set different linear and DNN optimizers, structures, learning rates and regularization rates. Details about the hyperparameters can be found from our [Wide-Deep Quickstart notebook](../00_quick_start/wide_deep_movielens.ipynb).
Hyperdrive provides three different parameter sampling methods: `RandomParameterSampling`, `GridParameterSampling`, and `BayesianParameterSampling`. Details about each method can be found from [Azure doc](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-tune-hyperparameters). Here, we use the Bayesian sampling.
```
# Fixed parameters
script_params = {
'--datastore': ds.as_mount(),
'--train-datapath': "data/" + TRAIN_FILE_NAME,
'--test-datapath': "data/" + VALID_FILE_NAME,
'--top-k': TOP_K,
'--user-col': USER_COL,
'--item-col': ITEM_COL,
'--item-feat-col': ITEM_FEAT_COL,
'--rating-col': RATING_COL,
'--ranking-metrics': RANKING_METRICS,
'--rating-metrics': RATING_METRICS,
'--epochs': EPOCHS,
'--model-type': 'wide_deep'
}
# Hyperparameter search space
params = {
'--batch-size': hd.choice(64, 128, 256),
# Linear model hyperparameters
'--linear-optimizer': hd.choice('Ftrl'), # 'SGD' and 'Momentum' easily got exploded loss in regression problems.
'--linear-optimizer-lr': hd.uniform(0.0001, 0.1),
'--linear-l1-reg': hd.uniform(0.0, 0.1),
# Deep model hyperparameters
'--dnn-optimizer': hd.choice('Adagrad', 'Adam'),
'--dnn-optimizer-lr': hd.uniform(0.0001, 0.1),
'--dnn-user-embedding-dim': hd.choice(4, 8, 16, 32, 64),
'--dnn-item-embedding-dim': hd.choice(4, 8, 16, 32, 64),
'--dnn-hidden-layer-1': hd.choice(0, 32, 64, 128, 256, 512, 1024), # 0: not using this layer
'--dnn-hidden-layer-2': hd.choice(0, 32, 64, 128, 256, 512, 1024),
'--dnn-hidden-layer-3': hd.choice(0, 32, 64, 128, 256, 512, 1024),
'--dnn-hidden-layer-4': hd.choice(32, 64, 128, 256, 512, 1024),
'--dnn-batch-norm': hd.choice(0, 1),
'--dnn-dropout': hd.choice(0.0, 0.1, 0.2, 0.3, 0.4)
}
```
**AzureML Estimator** is the building block for training. An Estimator encapsulates the training code and parameters, the compute resources and runtime environment for a particular training scenario (Note, this is not TensorFlow's Estimator)
We create one for our experimentation with the dependencies our model requires as follows:
```
conda_packages=['pandas', 'scikit-learn'],
pip_packages=['ipykernel', 'papermill', 'tensorflow-gpu==1.12']
```
To the Hyperdrive Run Config, we set our primary metric name and the goal (our hyperparameter search criteria), hyperparameter sampling method, and number of total child-runs. The bigger the search space, the more number of runs we will need for better results.
```
est = aml.train.estimator.Estimator(
source_directory=SCRIPT_DIR,
entry_script=ENTRY_SCRIPT_NAME,
script_params=script_params,
compute_target=compute_target,
use_gpu=True,
conda_packages=['pandas', 'scikit-learn'],
pip_packages=['ipykernel', 'papermill', 'tensorflow-gpu==1.12']
)
hd_run_config = hd.HyperDriveRunConfig(
estimator=est,
hyperparameter_sampling=hd.BayesianParameterSampling(params),
primary_metric_name=PRIMARY_METRIC,
primary_metric_goal=hd.PrimaryMetricGoal.MINIMIZE,
max_total_runs=MAX_TOTAL_RUNS,
max_concurrent_runs=MAX_CONCURRENT_RUNS
)
```
#### 5.3 Run Experiment
Now we submit the Run to our experiment. You can see the experiment progress from this notebook by using `azureml.widgets.RunDetails(hd_run).show()` or check from the Azure portal with the url link you can get by running `hd_run.get_portal_url()`.
<img src="https://recodatasets.blob.core.windows.net/images/aml_0.png?sanitize=true" width="600"/>
<img src="https://recodatasets.blob.core.windows.net/images/aml_1.png?sanitize=true" width="600"/>
<center><i>AzureML Hyperdrive Widget</i></center>
To load an existing Hyperdrive Run instead of start new one, use `hd_run = hd.HyperDriveRun(exp, <user-run-id>, hyperdrive_run_config=hd_run_config)`. You also can cancel the Run with `hd_run.cancel()`.
```
hd_run = exp.submit(config=hd_run_config)
widgets.RunDetails(hd_run).show()
```
Once all the child-runs are finished, we can get the best run and the metrics.
> Note, if you run Hyperdrive experiment again, you will see the best metrics and corresponding hyperparameters are not the same. It is because of 1) the random initialization of the model and 2) Hyperdrive sampling (when you use RandomSampling). You will get different results as well if you use different training and validation sets.
```
# Get best run and printout metrics
best_run = hd_run.get_best_run_by_primary_metric()
best_run_metrics = best_run.get_metrics()
print("* Best Run Id:", best_run.id)
print("\n* Best hyperparameters:")
print("Model type =", best_run_metrics['MODEL_TYPE'])
print("Batch size =", best_run_metrics['BATCH_SIZE'])
print("Linear optimizer =", best_run_metrics['LINEAR_OPTIMIZER'])
print("\tLearning rate = {0:.4f}".format(best_run_metrics['LINEAR_OPTIMIZER_LR']))
print("\tL1 regularization = {0:.4f}".format(best_run_metrics['LINEAR_L1_REG']))
print("DNN optimizer =", best_run_metrics['DNN_OPTIMIZER'])
print("\tUser embedding dimension =", best_run_metrics['DNN_USER_DIM'])
print("\tItem embedding dimension =", best_run_metrics['DNN_ITEM_DIM'])
hidden_units = []
for i in range(1, 5):
hidden_nodes = best_run_metrics['DNN_HIDDEN_LAYER_{}'.format(i)]
if hidden_nodes > 0:
hidden_units.append(hidden_nodes)
print("\tHidden units =", hidden_units)
print("\tLearning rate = {0:.4f}".format(best_run_metrics['DNN_OPTIMIZER_LR']))
print("\tDropout rate = {0:.4f}".format(best_run_metrics['DNN_DROPOUT']))
print("\tBatch normalization =", best_run_metrics['DNN_BATCH_NORM'])
# Metrics evaluated on validation set
print("\n* Performance metrics:")
print("Top", TOP_K)
for m in RANKING_METRICS:
print("\t{0} = {1:.4f}".format(m, best_run_metrics[m]))
for m in RATING_METRICS:
print("\t{0} = {1:.4f}".format(m, best_run_metrics[m]))
```
### 6. Model Import and Test
[Wide-Deep Quickstart notebook](../00_quick_start/wide_deep_movielens.ipynb), which we've used in our Hyperdrive Experiment, exports the trained model to the output folder (the output path is recorded at `best_run_metrics['saved_model_dir']`). We can download a model from the best run and test it.
```
MODEL_DIR = os.path.join(tmp_dir.name, 'aml_model')
os.makedirs(MODEL_DIR, exist_ok=True)
model_file_dir = os.path.normpath(best_run_metrics['saved_model_dir'][2:-1]) + '/'
print(model_file_dir)
for f in best_run.get_file_names():
if f.startswith(model_file_dir):
output_file_path = os.path.join(MODEL_DIR, f[len(model_file_dir):])
print("Downloading {}..".format(f))
best_run.download_file(name=f, output_file_path=output_file_path)
saved_model = tf.contrib.estimator.SavedModelEstimator(MODEL_DIR)
cols = {
'col_user': USER_COL,
'col_item': ITEM_COL,
'col_rating': RATING_COL,
'col_prediction': 'prediction'
}
tf.logging.set_verbosity(tf.logging.ERROR)
# Prediction input function for TensorFlow SavedModel
def predict_input_fn(df):
def input_fn():
examples = [None] * len(df)
for index, test_sample in df.iterrows():
example = tf.train.Example()
example.features.feature[USER_COL].int64_list.value.extend([test_sample[USER_COL]])
example.features.feature[ITEM_COL].int64_list.value.extend([test_sample[ITEM_COL]])
example.features.feature[ITEM_FEAT_COL].float_list.value.extend(test_sample[ITEM_FEAT_COL])
examples[index] = example.SerializeToString()
return {'inputs': tf.constant(examples)}
return input_fn
# Rating prediction set
X_test = test.drop(RATING_COL, axis=1)
X_test.reset_index(drop=True, inplace=True)
# Rating prediction
predictions = list(itertools.islice(
saved_model.predict(predict_input_fn(X_test)),
len(X_test)
))
prediction_df = X_test.copy()
prediction_df['prediction'] = [p['outputs'][0] for p in predictions]
print(prediction_df['prediction'].describe(), "\n")
for m in RATING_METRICS:
fn = getattr(reco_utils.evaluation.python_evaluation, m)
result = fn(test, prediction_df, **cols)
print(m, "=", result)
# Unique items
if ITEM_FEAT_COL is None:
items = data.drop_duplicates(ITEM_COL)[[ITEM_COL]].reset_index(drop=True)
else:
items = data.drop_duplicates(ITEM_COL)[[ITEM_COL, ITEM_FEAT_COL]].reset_index(drop=True)
# Unique users
users = data.drop_duplicates(USER_COL)[[USER_COL]].reset_index(drop=True)
# Ranking prediction set
ranking_pool = user_item_pairs(
user_df=users,
item_df=items,
user_col=USER_COL,
item_col=ITEM_COL,
user_item_filter_df=pd.concat([train, valid]), # remove seen items
shuffle=True
)
predictions = []
# To prevent creating a tensor proto whose content is larger than 2GB (which will raise an error),
# divide ranking_pool into 10 chunks, predict each, and concat back.
for pool in np.array_split(ranking_pool, 10):
pool.reset_index(drop=True, inplace=True)
# Rating prediction
pred = list(itertools.islice(
saved_model.predict(predict_input_fn(pool)),
len(pool)
))
predictions.extend([p['outputs'][0] for p in pred])
ranking_pool['prediction'] = predictions
for m in RANKING_METRICS:
fn = getattr(reco_utils.evaluation.python_evaluation, m)
result = fn(test, ranking_pool, **{**cols, 'k': TOP_K})
print(m, "=", result)
```
#### Wide-and-Deep Baseline Comparison
To see if Hyperdrive found good hyperparameters, we simply compare with the model with known hyperparameters from [TensorFlow's wide-deep learning example](https://github.com/tensorflow/models/blob/master/official/wide_deep/movielens_main.py) which uses only the DNN part from the wide-and-deep model for MovieLens data.
> Note, this is not 'apples to apples' comparison. For example, TensorFlow's movielens example uses *rating-timestamp* as a numeric feature, but we did not use that here because we think the timestamps are not relevant to the movies' ratings. This comparison is more like to show how Hyperdrive can help to find comparable hyperparameters without requiring exhaustive efforts in going over a huge search-space.
```
OUTPUT_NOTEBOOK = os.path.join(tmp_dir.name, "output.ipynb")
OUTPUT_MODEL_DIR = os.path.join(tmp_dir.name, "known_hyperparam_model_checkpoints")
params = {
'MOVIELENS_DATA_SIZE': MOVIELENS_DATA_SIZE,
'TOP_K': TOP_K,
'MODEL_TYPE': 'deep',
'EPOCHS': EPOCHS,
'BATCH_SIZE': 256,
'DNN_OPTIMIZER': 'Adam',
'DNN_OPTIMIZER_LR': 0.001,
'DNN_HIDDEN_LAYER_1': 256,
'DNN_HIDDEN_LAYER_2': 256,
'DNN_HIDDEN_LAYER_3': 256,
'DNN_HIDDEN_LAYER_4': 128,
'DNN_USER_DIM': 16,
'DNN_ITEM_DIM': 64,
'DNN_DROPOUT': 0.3,
'DNN_BATCH_NORM': 0,
'MODEL_DIR': OUTPUT_MODEL_DIR,
'EVALUATE_WHILE_TRAINING': False,
'EXPORT_DIR_BASE': OUTPUT_MODEL_DIR,
'RANKING_METRICS': RANKING_METRICS,
'RATING_METRICS': RATING_METRICS,
}
start_time = time.time()
pm.execute_notebook(
"../00_quick_start/wide_deep_movielens.ipynb",
OUTPUT_NOTEBOOK,
parameters=params,
kernel_name='python3'
)
end_time = time.time()
print("Training and evaluation of Wide-and-Deep model took", end_time-start_time, "secs.")
nb = pm.read_notebook(OUTPUT_NOTEBOOK)
for m in RANKING_METRICS:
print(m, "=", nb.data[m])
for m in RATING_METRICS:
print(m, "=", nb.data[m])
```
### Concluding Remark
We showed how to tune hyperparameters by utilizing Azure Machine Learning service. Complex and powerful models like Wide-and-Deep model often have many number of hyperparameters that affect on the recommendation accuracy, and it is not practical to tune the model without using a GPU cluster. For example, a training and evaluation of a model took around 3 minutes on 100k MovieLens data on a single *Standard NC6* VM as we tested from the [above cell](#Wide-and-Deep-Baseline-Comparison). When we used 1M MovieLens, it took about 47 minutes. If we want to investigate through 100 different combinations of hyperparameters **manually**, it will take **78 hours** on the VM and we may still wonder if we had tested good candidates of hyperparameters. With AzureML, as we shown in this notebook, we can easily setup different size of GPU cluster fits to our problem and utilize Bayesian sampling to navigate through the huge search space efficiently, and tweak the experiment with different criteria and algorithms for further research.
#### Cleanup
```
tmp_dir.cleanup()
```
| github_jupyter |
Check the performance of different classifiers by applying the following metrics:
* Confusion Matrix
* Accuracy (how many of the predicted results are similar to the test set results?
* Precision (measuring exactness; when it predicts yes, how often is it correct?)
* Recall (measuring completeness; when it's actually yes, how often does it predict yes?)
* F1 Score (compromise between Precision and Recall)
* Save the results within a dataframe and export it to a csv
```
import numpy as np
import pandas as pd
import re
import nltk
nltk.download('stopwords')
from nltk.corpus import stopwords
from nltk.stem.porter import PorterStemmer
dataset = pd.read_csv('Restaurant_Reviews.tsv', delimiter='\t', quoting = 3)
corpus = []
for i in range(0, 1000):
review = re.sub('[^a-zA-Z]', ' ', dataset['Review'][i])
review = review.lower()
review = review.split()
ps = PorterStemmer()
all_stopwords = stopwords.words('english')
all_stopwords.remove('not')
review = [ps.stem(word) for word in review if not word in set(all_stopwords)]
review = ' '.join(review)
corpus.append(review)
from sklearn.feature_extraction.text import CountVectorizer
cv = CountVectorizer(max_features = 1500)
X = cv.fit_transform(corpus).toarray()
y = dataset.iloc[:, -1].values
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2, random_state = 0)
#Naive Bayes Classifier
from sklearn.naive_bayes import GaussianNB
classifier = GaussianNB()
classifier.fit(X_train, y_train)
y_pred = classifier.predict(X_test)
from sklearn.metrics import confusion_matrix, accuracy_score
def model_performance(y_pred):
value_list = []
cm = confusion_matrix(y_test, y_pred)
TN = cm[0][0]
value_list.append(cm[0][0])
TP = cm[1][1]
value_list.append(cm[1][1])
FP = cm[0][1]
value_list.append(cm[0][1])
FN = cm[1][0]
value_list.append(cm[1][0])
Accuracy = (TP + TN) / (TP + TN + FP + FN)
value_list.append(Accuracy)
Precision = TP / (TP + FP)
value_list.append(round(Precision, 3))
Recall = TP / (TP + FN)
value_list.append(round(Recall, 3))
F1 = 2 * Precision * Recall / (Precision + Recall)
value_list.append(round(F1, 3))
return print((cm),'\n'
'True Negatives:', cm[0][0],'\n'
'True Positives:', cm[1][1],'\n'
'False Positives:', cm[0][1],'\n'
'False Negatives:', cm[1][0],'\n'
'Accurary:', Accuracy,'\n'
'Precision:', round(Precision, 3),'\n'
'Recall:', round(Recall, 3),'\n'
'F1 Score:', round(F1, 3)), value_list
list_nb = model_performance(y_pred)[1]
list_nb
#Random Forest Classifier
from sklearn.ensemble import RandomForestClassifier
class_rf = RandomForestClassifier(n_estimators = 30, criterion = 'entropy', random_state = 0)
class_rf.fit(X_train, y_train)
y_pred = class_rf.predict(X_test)
list_rf = model_performance(y_pred)[1]
list_rf
#Support Vector Machine Classifier
from sklearn.svm import SVC
class_svm = SVC(kernel = 'linear', random_state = 0)
class_svm.fit(X_train, y_train)
y_pred = class_svm.predict(X_test)
list_svm = model_performance(y_pred)[1]
list_svm
#Kernel SVM Classifier
class_kern = SVC(kernel = 'rbf', random_state = 0)
class_kern.fit(X_train, y_train)
y_pred = class_kern.predict(X_test)
list_kern = model_performance(y_pred)[1]
list_kern
#Decision Tree Classifier
from sklearn.tree import DecisionTreeClassifier
class_tree = DecisionTreeClassifier(criterion = 'entropy', random_state=0)
class_tree.fit(X_train, y_train)
y_pred = class_tree.predict(X_test)
list_tree = model_performance(y_pred)[1]
list_tree
from sklearn.neighbors import KNeighborsClassifier
class_knn = KNeighborsClassifier(n_neighbors = 10, metric = 'minkowski', p = 2)
class_knn.fit(X_train, y_train)
y_pred = class_knn.predict(X_test)
list_knn = model_performance(y_pred)[1]
list_knn
from sklearn.linear_model import LogisticRegression
class_log = LogisticRegression(random_state = 0)
class_log.fit(X_train, y_train)
y_pred = class_log.predict(X_test)
list_log = model_performance(y_pred)[1]
list_log
#artificial neural network
#feature scaling
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
X_train_scaled = sc.fit_transform(X_train)
X_test_scaled = sc.transform(X_test)
#initialize artificial neural network
import tensorflow as tf
ann = tf.keras.models.Sequential()
#adding input layer and first hidden layer
ann.add(tf.keras.layers.Dense(units=6, activation='relu'))
#adding second hidden layer
ann.add(tf.keras.layers.Dense(units=6, activation='relu'))
#adding output layer
ann.add(tf.keras.layers.Dense(units=1, activation='sigmoid'))
#compiling the ANN
ann.compile(optimizer='adam', loss='binary_crossentropy', metrics=['accuracy'])
#training ANN on training set
ann.fit(X_train_scaled, y_train, batch_size=32, epochs=100)
#predict test set results
y_pred = ann.predict(X_test_scaled)
#change to boolean
y_pred = (y_pred > 0.5)
#check performance of ANN
list_ann = model_performance(y_pred)[1]
list_ann
df = pd.DataFrame(zip(list_nb, list_rf, list_svm, list_kern, list_tree, list_knn, list_log, list_ann), index = ['True Neg', 'True Pos', 'False Pos',
'False Neg', 'Accuracy', 'Precision',
'Recall', 'F1 Score'], columns = ['Naive Bayes', 'Random Forest', 'Linear SVM',
'Kernel SVM', 'Decision Tree', 'K-NN', 'Log Reg.', 'ANN'])
df
df.to_csv('model_selection.csv', index=True, header=True)
```
| github_jupyter |
# Show iterative steps of preprocessing
```
import data_utils
import numpy as np
import matplotlib.pyplot as plt
from preprocessing import binarize_per_slice, all_slice_analysis, fill_hole, two_lung_only, process_mask
# Show iterative steps of computing lung mask
first_patient_pixels, spacing, _ = data_utils.load_dicom_slices("../../data/LIDC-IDRI-DCM/LIDC-IDRI-0001/01-01-2000-30178/3000566-03192/")
print(first_patient_pixels.shape)
import matplotlib.pyplot as plt
plt.hist(first_patient_pixels.flatten(), bins=80, color='c')
plt.xlabel("Hounsfield Units (HU)")
plt.ylabel("Frequency")
plt.show()
# Show some slice in the middle
h = 80
plt.imshow(first_patient_pixels[h], cmap=plt.cm.gray)
plt.show()
bw = binarize_per_slice(first_patient_pixels, spacing)
plt.imshow(bw[h], cmap=plt.cm.gray)
plt.show()
```
Parallélisé mais très long sur Power (de l'ordre de 2 minutes).
```
flag = 0
cut_num = 0
while flag == 0:
bw, flag = all_slice_analysis(bw, spacing, cut_num=cut_num)
cut_num = cut_num + 1
plt.imshow(bw[h], cmap=plt.cm.gray)
plt.show()
```
Pas de multiprocessing.
```
bw = fill_hole(bw)
plt.imshow(bw[h], cmap=plt.cm.gray)
plt.show()
```
Pas de multiprocessing.
```
bw1, bw2, bw = two_lung_only(bw, spacing)
plt.imshow(bw[h], cmap=plt.cm.gray)
plt.show()
```
Pas de multiprocessing. Plutôt long.
```
plt.imshow(bw1[h], cmap=plt.cm.gray)
plt.show()
plt.imshow(bw2[h], cmap=plt.cm.gray)
plt.show()
dm1 = process_mask(bw1)
dm2 = process_mask(bw2)
plt.imshow(dm1[h]+dm2[h], cmap=plt.cm.gray)
plt.show()
dm = process_mask(bw)
plt.imshow(dm[h], cmap=plt.cm.gray)
plt.show()
x = np.load("/wmlce/data/projects/lung_segmentation/output/preprocessing/2_128_256_256/456.npy")
plt.imshow(x[h], cmap=plt.cm.gray)
plt.show()
x_mask = np.load("/wmlce/data/projects/lung_segmentation/output/preprocessing/2_128_256_256/456_mask.npy")
plt.imshow(x_mask[h], cmap=plt.cm.gray)
plt.show()
```
# Using U-Net Lungs Segmentation
```
import os
import sys
import time
import torch
import mlflow
import mlflow.pytorch
import numpy as np
import SimpleITK as sitk
from pathlib import Path
import matplotlib.pyplot as plt
os.environ['MDT_DATASETS_DIR'] = '/wmlce/data/medical-datasets'
from preprocessing import binarize_per_slice, all_slice_analysis, fill_hole, two_lung_only, process_mask, resample_array, lumTrans
LS_PATH = os.path.join('.', 'lung-segmentation')
sys.path.append(LS_PATH)
import predict
from data import utils as data_utils
start_time = time.time()
pid = 'LIDC-IDRI-0489'
path = f'/wmlce/data/medical-datasets/MDT-LIDC-IDRI/NRRDs/{pid}'
target_spacing = (0.7, 0.7, 1.25)
remote_server_uri = "http://mlflow.10.7.13.202.nip.io/"
mlflow.set_tracking_uri(remote_server_uri)
h = 150
# Load scan
img = sitk.ReadImage(os.path.join(path, '{}_CT.nrrd'.format(pid)))
original_spacing = np.array(img.GetSpacing())
img_arr = sitk.GetArrayFromImage(img)
ls_img_arr = np.copy(img_arr)
load_time = time.time()
print(f'{pid}: loaded in {load_time - start_time} s')
# Resample and Normalize
img_arr = resample_array(img_arr, img.GetSpacing(), target_spacing)
lum_img_arr = np.copy(img_arr)
img_arr = np.clip(img_arr, -1200, 600)
img_arr = img_arr.astype(np.float32)
img_arr = (img_arr - np.mean(img_arr)) / np.std(img_arr).astype(np.float16)
norm_time = time.time()
print(f'{pid}: Resampled in {norm_time - load_time} s')
print(f'{pid}: {img_arr.shape}, {target_spacing}')
plt.imshow(img_arr[h], cmap=plt.cm.gray)
plt.show()
# Compute lungs mask
model_name = "2-lungs-segmentation"
unet = mlflow.pytorch.load_model("models:/{}/production".format(model_name))
print(ls_img_arr.shape, original_spacing)
ls_img_arr, spacing = data_utils.prep_img_arr(ls_img_arr, original_spacing)
print(ls_img_arr.shape, spacing)
mask = predict.predict(ls_img_arr, 1, unet, threshold=True, erosion=True)
print(mask.shape, spacing)
mask, spacing = data_utils.prep_img_arr(mask[0][0], spacing, target_shape=img_arr.shape)
mask = mask[0]
mask[mask>0.5] = 1
mask[mask!=1] = 0
print(mask.shape, target_spacing)
ls_time = time.time()
print(f'{pid}: Lung segmentation took {ls_time - norm_time} s')
plt.imshow(mask[h], cmap=plt.cm.gray)
plt.show()
dilatedMask = process_mask(mask)
Mask = mask
extramask = dilatedMask.astype(np.uint8) - Mask.astype(np.uint8)
bone_thresh = 210
pad_value = 1 #170
img_arr[np.isnan(img_arr)]=-2000
sliceim = lumTrans(lum_img_arr)
#sliceim = sliceim*dilatedMask+pad_value*(1-dilatedMask).astype('uint8')
bones = sliceim*extramask>bone_thresh
#sliceim[bones] = pad_value
img_arr = img_arr*dilatedMask+pad_value*(1-dilatedMask).astype('uint8')
img_arr[bones] = pad_value
bones_mask = np.zeros(sliceim.shape)
bones_mask[bones] = 1
print(f'{pid}: Cleaning took {time.time() - ls_time} s')
print(f'{pid}: Ellapsed {time.time() - start_time} s')
# Plot image
plt.subplot(2, 3, 1).imshow(img_arr[h], cmap=plt.cm.gray)
plt.subplot(2, 3, 2).imshow(Mask[h], cmap=plt.cm.gray)
plt.subplot(2, 3, 3).imshow(dilatedMask[h], cmap=plt.cm.gray)
plt.subplot(2, 3, 4).imshow(img_arr[h], cmap=plt.cm.gray)
plt.subplot(2, 3, 5).imshow(bones_mask[h], cmap=plt.cm.gray)
plt.subplot(2, 3, 6).imshow(extramask[h], cmap=plt.cm.gray)
plt.show()
dir_path = "/wmlce/data/medical-datasets/MDT-PP"
imgs = glob.glob(os.path.join(dir_path, "*_rois.npy"))
print(np.min(img_arr), np.max(img_arr))
plt.imshow(img_arr[h], cmap=plt.cm.gray)
plt.show()
```
## Load some images generated by such preprocessing
```
import os, glob
import numpy as np
import matplotlib.pyplot as plt
dir_path = "/wmlce/data/medical-datasets/MDT-PP"
imgs = glob.glob(os.path.join(dir_path, "*_rois.npy"))
h = 150
n = 10
for ix, img in enumerate(np.random.choice(imgs, n), 1):
img_arr = np.load(img.replace("_rois", "_img")).astype(np.float32)
rois_arr = np.load(img)
print(f"Image {os.path.splitext(os.path.basename(img))[0]} {img_arr.shape}, rois {rois_arr.shape}")
plt.subplot(2, n/2, ix).imshow(img_arr[h], cmap=plt.cm.gray)
plt.show()
dir_path = "/wmlce/data/medical-datasets/MDT-PP"
img = "LIDC-IDRI-0338_img.npy"
img = "LIDC-IDRI-0479_img.npy"
img = "LIDC-IDRI-0489_img.npy"
img = "LIDC-IDRI-0015_img.npy"
img = "LIDC-IDRI-0509_img.npy" # This image seems to have been swapped (axes issues / flipped ?)
img_arr = np.load(os.path.join(dir_path, img))
print(img_arr.shape, img_arr.dtype)
plt.imshow(img_arr[:,250,:], cmap=plt.cm.gray)
plt.show()
```
# Crap image analysis
### Resample to original size and save to nrrd
```
from preprocessing import resample_array_to_shape
itkimg = sitk.ReadImage("/wmlce/data/medical-datasets/LIDC-IDRI/LIDC-IDRI-0015/1.3.6.1.4.1.14519.5.2.1.6279.6001.231462296937187240061810311146/1.3.6.1.4.1.14519.5.2.1.6279.6001.227962600322799211676960828223/LIDC-IDRI-0015_CT.nrrd")
seg_mask, seg_spacing = resample_array_to_shape(img_arr, [0.7,0.7,1.25], target_shape=sitk.GetArrayFromImage(itkimg).shape)
new_itk = sitk.GetImageFromArray(seg_mask)
new_itk.SetOrigin(itkimg.GetOrigin())
new_itk.SetSpacing(itkimg.GetSpacing())
sitk.WriteImage(new_itk, 'test.nrrd')
```
# Get list of all images with high spacing / flipped images
```
import nrrd
import os, glob
import numpy as np
import SimpleITK as sitk
import matplotlib.pyplot as plt
def load_itk(filename):
with open(filename) as f:
contents = f.readlines()
line = [k for k in contents if k.startswith('TransformMatrix')][0]
transformM = np.array(line.split(' = ')[1].split(' ')).astype('float')
transformM = np.round(transformM)
if np.any( transformM!=np.array([1,0,0, 0, 1, 0, 0, 0, 1])):
isflip = True
else:
isflip = False
itkimage = sitk.ReadImage(filename)
numpyImage = sitk.GetArrayFromImage(itkimage)
numpyOrigin = np.array(list(reversed(itkimage.GetOrigin())))
numpySpacing = np.array(list(reversed(itkimage.GetSpacing())))
return numpyImage, numpyOrigin, numpySpacing, isflip
list_path = glob.glob("/wmlce/data/medical-datasets/LIDC-IDRI/LIDC-IDRI-*/*/*/*_CT.nrrd")
spaces = set([])
kinds = set([])
dimensions = set([])
high_spacing = set([])
for path in list_path:
h = nrrd.read_header(path)
spaces.add(h['space'])
for k in h['kinds']:
kinds.add(k)
dimensions.add(h['dimension'])
if np.max(h['space directions']) > 2.5:
high_spacing.add(path)
print(spaces)
print(kinds)
print(dimensions)
print(len(high_spacing))
```
# Check scans manually
```
import nrrd
import os, glob
import numpy as np
import SimpleITK as sitk
import matplotlib.pyplot as plt
from ipywidgets import interact, interact_manual
from IPython.display import display
import matplotlib.pyplot as plt
import ipywidgets as widgets
ix = 0
#list_path = glob.glob("/wmlce/data/medical-datasets/LIDC-IDRI/LIDC-IDRI-*/*/*/*_CT.nrrd")
list_path = glob.glob("/wmlce/data/medical-datasets/MDT-PP/*_img.npy")
import csv
#csv_path = 'scans_checkup.csv'
csv_path = 'pp_scans_checkup.csv'
header = ['pid', 'flip', 'crap', 'poor', 'warn', 'note']
if not os.path.exists(csv_path):
f = open(csv_path, 'w+')
writer = csv.DictWriter(f, fieldnames=header)
writer.writeheader()
f.close()
def f(flip, crap, poor, warn, note):
global ix, list_path, csv_path, header
if ix >= 1 and (flip or crap or poor or warn):
#pid = os.path.splitext(os.path.basename(list_path[ix-1]))[0].split('_CT')[0]
pid = os.path.splitext(os.path.basename(list_path[ix-1]))[0].split('_img')[0]
print(f'Adding abnormal img {pid} to csv')
f = open(csv_path, 'a')
writer = csv.DictWriter(f, fieldnames=header)
writer.writerow({header[0]: pid, header[1]: int(flip), header[2]: int(crap), header[3]: int(poor), header[4]: int(warn), header[5]: note})
f.close()
if ix >= 0 and ix < len(list_path):
print(f'Showing img: {ix}: {list_path[ix]}')
## Show ITK
#itkimage = sitk.ReadImage(list_path[ix])
#numpyImage = sitk.GetArrayFromImage(itkimage)
numpyImage = np.load(list_path[ix])
plt.imshow(numpyImage[int(len(numpyImage)/2)].astype(np.uint8), cmap=plt.cm.gray)
plt.show()
elif ix >= 0:
print('Done')
ix = ix + 1
_ = interact_manual(f, flip=False, crap=False, poor=False, warn=False, note='')
```
# Check ROI labels
```
import os, glob
import numpy as np
list_paths = glob.glob("/wmlce/data/medical-datasets/MDT-PP/*_rois.npy")
for path in list_paths[:10]:
arr = np.load(path)
pid = os.path.splitext(os.path.basename(path))[0].split('_')[0]
print(pid, np.unique(arr))
```
## Upsample ROIs to original scan size for visualization
```
os.environ["MDT_DATASETS_DIR"] = "/wmlce/data/medical-datasets"
from preprocessing import resample_array_to_shape
import numpy as np
import os, glob
itkimg = sitk.ReadImage(glob.glob("/wmlce/data/medical-datasets/LIDC-IDRI/LIDC-IDRI-0806/*/*/*_CT.nrrd")[0])
rois_path = "/wmlce/data/medical-datasets/MDT-PP/LIDC-IDRI-0806_rois.npy"
pid = os.path.splitext(os.path.basename(rois_path))[0].split('_')[0]
rois_arr = np.load(rois_path)
rois_arr[rois_arr != 0] = 1
seg_mask, seg_spacing = resample_array_to_shape(rois_arr, [0.7,0.7,1.25], target_shape=sitk.GetArrayFromImage(itkimg).shape)
seg_mask[seg_mask >= 0.5] = 1
seg_mask[seg_mask < 0.5] = 0
seg_mask = seg_mask.astype(np.uint8)
new_itk = sitk.GetImageFromArray(seg_mask)
new_itk.SetOrigin(itkimg.GetOrigin())
new_itk.SetSpacing(itkimg.GetSpacing())
sitk.WriteImage(new_itk, f'{pid}_rois.nrrd')
```
## GAN generated scans
```
import os, glob
import numpy as np
import SimpleITK as sitk
import matplotlib.pyplot as plt
from ipywidgets import interact, interact_manual
from IPython.display import display
import matplotlib.pyplot as plt
import ipywidgets as widgets
list_path = glob.glob("/wmlce/data/medical-datasets/MDT-PP/*-AUG_img.npy")
for path in list_path:
pid = os.path.basename(path).replace("_img.npy", "")
n_nods = len(glob.glob(f"/wmlce/data/medical-datasets/MDT-LIDC-IDRI/NRRDs/{pid}/*nod*"))
print(pid, n_nods, np.unique(np.load(path.replace("_img", "_rois"))))
import csv
#csv_path = 'scans_checkup.csv'
csv_path = 'pp_aug_scans_checkup.csv'
header = ['pid', 'flip', 'crap', 'poor', 'warn', 'note']
if not os.path.exists(csv_path):
f = open(csv_path, 'w+')
writer = csv.DictWriter(f, fieldnames=header)
writer.writeheader()
f.close()
def f(flip, crap, poor, warn, note):
global ix, list_path, csv_path, header
if ix >= 1 and (flip or crap or poor or warn):
#pid = os.path.splitext(os.path.basename(list_path[ix-1]))[0].split('_CT')[0]
pid = os.path.splitext(os.path.basename(list_path[ix-1]))[0].split('_img')[0]
print(f'Adding abnormal img {pid} to csv')
f = open(csv_path, 'a')
writer = csv.DictWriter(f, fieldnames=header)
writer.writerow({header[0]: pid, header[1]: int(flip), header[2]: int(crap), header[3]: int(poor), header[4]: int(warn), header[5]: note})
f.close()
if ix >= 0 and ix < len(list_path):
print(f'Showing img: {ix}: {list_path[ix]}')
## Show ITK
#itkimage = sitk.ReadImage(list_path[ix])
#numpyImage = sitk.GetArrayFromImage(itkimage)
numpyImage = np.load(list_path[ix])
plt.imshow(numpyImage[int(len(numpyImage)/2)].astype(np.uint8), cmap=plt.cm.gray)
plt.show()
elif ix >= 0:
print('Done')
ix = ix + 1
_ = interact_manual(f, flip=False, crap=False, poor=False, warn=False, note='')
```
| github_jupyter |
```
import os
import tensorflow as tf
import keras
from keras.layers import Add,Multiply,Softmax,Input,TimeDistributed,Dense,Average,GlobalAveragePooling1D,Concatenate,Lambda,RepeatVector, Conv2D,ConvLSTM2D, MaxPooling2D,BatchNormalization,Flatten,Reshape,UpSampling2D
from keras.models import Model, load_model
from keras.optimizers import Adam
from keras.utils import plot_model
import numpy as np
import matplotlib.pyplot as plt
import math
import time
import pylab as pl
from IPython import display
from IPython.core.display import HTML
from IPython.core.display import display as html_width
html_width(HTML("<style>.container { width:90% !important; }</style>"))
import tensorflow_probability as tfp
import matplotlib.image as mpimg
from matplotlib.gridspec import GridSpec
import imageio
from tqdm import tqdm
from keras.utils.vis_utils import plot_model
from keras.backend.tensorflow_backend import set_session
#physical_devices = tf.config.experimental.list_physical_devices('GPU')
#tf.config.experimental.set_memory_growth(physical_devices[0], True)
config = tf.ConfigProto()
config.gpu_options.allow_growth = True # dynamically grow the memory used on the GPU
config.log_device_placement = True # to log device placement (on which device the operation ran)
sess = tf.Session(config=config)
set_session(sess)
from tensorflow.python.client import device_lib
def get_available_devices():
local_device_protos = device_lib.list_local_devices()
return [x.name for x in local_device_protos]
print(get_available_devices())
print(tf.__version__)
obs_max = 5
train_N = 40
train_p = np.random.permutation(40)
```
# Deep Modality Blending Networks
```
def get_train_sample(action_type = -1, coef = -1):
n = np.random.randint(0,obs_max)+1
d = train_p[np.random.randint(0, train_N)]
if action_type == -1:
action_type = np.random.randint(0,2)
action_type=0
if action_type == 0:
action_type = 'move'
else:
action_type = 'grasp'
if coef == -1:
coef = np.random.rand()
img_coef = np.ones((1,128)) * coef
pose_coef = np.ones((1,128)) * (1-coef)
observation = np.zeros((1,n,128,128,4))
observation_pose = np.zeros((1,n,8))
target_X = np.zeros((1,1))
target_Y = np.zeros((1,128,128,6))
target_Y_pose = np.zeros((1,14))
pose = np.loadtxt('../data/data2020/%d/%s/joint_%d.txt'%(d,action_type,d))
pose[:,-1] *= 10
time_len = pose.shape[0]
times = np.linspace(0,1,time_len)
perm = np.random.permutation(time_len)
for i in range(n):
observation[0,i,:,:,0] = np.ones((128,128))*times[perm[i]]
observation[0,i,:,:,1:] = mpimg.imread('../data/data2020/%d/%s/%d.jpeg'%(d,action_type,perm[i]))/255.
observation_pose[0,i,0] = times[perm[i]]
observation_pose[0,i,1:] = pose[perm[i]]
target_X[0,0] = times[perm[n]]
target_Y[0,:,:,:3] = mpimg.imread('../data/data2020/%d/%s/%d.jpeg'%(d,action_type,perm[n]))/255.
target_Y_pose[0,:7] = pose[perm[n]]
return [observation, observation_pose, target_X, img_coef, pose_coef], [target_Y, target_Y_pose], d, perm[n]
def custom_loss(y_true, y_predicted):
mean, log_sigma = tf.split(y_predicted, 2, axis=-1)
y_true_value, temp =tf.split(y_true,2,axis=-1)
sigma = tf.nn.softplus(log_sigma)
dist = tfp.distributions.MultivariateNormalDiag(loc=mean, scale_diag=sigma)
loss = -tf.reduce_mean(dist.log_prob(y_true_value))
return loss
image_layer = Input(shape=(None,128,128,4), name="image_observation")
joint_layer = Input(shape=(None,8), name="joint_observation")
target_X_layer = Input(shape=(1,), name = 'target_X')
img_coef_layer = Input(shape=(128,), name = 'image_coef')
pose_coef_layer = Input(shape=(128,), name = 'pose_coef')
encoder_joint_sizes = [64,64,128,128,256]
joint_encoder = TimeDistributed(Dense(32, activation = 'relu'))(joint_layer)
for channel_size in encoder_joint_sizes:
joint_encoder = TimeDistributed(Dense(channel_size, activation = 'relu'))(joint_encoder)
joint_representations = TimeDistributed(Dense(128, activation='relu'))(joint_encoder) #128
joint_representation = GlobalAveragePooling1D()(joint_representations)
multiplied_joint = Multiply()([joint_representation,pose_coef_layer])
###################################################
encoder_img_sizes = [64,64,128,128,256]
image_encoder = TimeDistributed(Conv2D(32,(3,3),padding='same',activation='relu'))(image_layer)
image_encoder = TimeDistributed(MaxPooling2D((2,2)))(image_encoder)
for channel_size in encoder_img_sizes:
image_encoder = TimeDistributed(Conv2D(channel_size,(3,3),padding='same',activation='relu'))(image_encoder)
image_encoder = TimeDistributed(MaxPooling2D((2,2)))(image_encoder)
image_flatten = TimeDistributed(Flatten())(image_encoder)
img_representations = TimeDistributed(Dense(128, activation='relu'))(image_flatten)
img_representation = GlobalAveragePooling1D()(img_representations)
multiplied_img = Multiply()([img_representation,img_coef_layer])
general_representation = Add()([multiplied_joint,multiplied_img])
merged_layer = Concatenate(axis=-1, name='merged')([general_representation,target_X_layer])
####################################################
decoder_representation = Dense(1024, activation='relu') (merged_layer)
" =============== Image Decoder =============== "
decoder_img = Reshape([2,2,256])(decoder_representation)
decoder_img_sizes = [256,128,128,64,64,32]
for channel_size in decoder_img_sizes:
decoder_img = Conv2D(channel_size, (3,3), padding='same', activation='relu')(decoder_img)
decoder_img = UpSampling2D((2, 2))(decoder_img)
img_output = Conv2D(16, (3,3), padding='same', activation='relu')(decoder_img)
img_output = Conv2D(8, (3,3), padding='same', activation='relu')(img_output)
img_output = Conv2D(6, (3,3), padding='same', activation='sigmoid')(img_output)
" =============== Image Decoder =============== "
" =============== Joint Decoder =============== "
decoder_joint = Dense(512, activation='relu')(decoder_representation)
decoder_joint = Dense(216, activation='relu')(decoder_joint)
decoder_joint = Dense(128, activation='relu')(decoder_joint)
decoder_joint = Dense(32, activation='relu')(decoder_joint)
joint_output = Dense(14)(decoder_joint)
" =============== Joint Decoder =============== "
model = Model([image_layer, joint_layer, target_X_layer, img_coef_layer, pose_coef_layer],[img_output,joint_output])
latent_model = Model([image_layer, joint_layer, target_X_layer, img_coef_layer, pose_coef_layer],general_representation)
model.compile(optimizer = Adam(lr = 1e-4),loss=custom_loss, loss_weights=[1,0.01])
model.summary()
#plot_model(model)
loss_checkpoint = 1000
plot_checkpoint = 1000
validation_checkpoint = 1000
validation_error = 9999999
validation_step = -1
max_training_step = 1000000
dataset = ['image','joint']
float_formatter = "{:.4f}".format
np.set_printoptions(formatter={'float_kind':float_formatter})
for step in range(max_training_step):
inp, out, _, _ = get_train_sample()
callback = model.fit(inp,out)
'''
if step % validation_checkpoint == 0:
pass
'''
if step % plot_checkpoint == 0:
#clearing output cell
display.clear_output(wait=True)
display.display(pl.gcf())
print(step)
#plotting on-train examples by user given observations
inp, out, d_id, target_t = get_train_sample()
plt.imshow(out[0][0,:,:,:3])
plt.show()
plt.imshow(model.predict(inp)[0][0,:,:,:3])
plt.show()
print(out[1][0,:7])
print(model.predict(inp)[1][0,:7])
inp, out, d_id, target_t = get_train_sample()
plt.imshow(out[0][0,:,:,:3])
plt.show()
plt.imshow(model.predict(inp)[0][0,:,:,:3])
plt.show()
print(out[1][0,:7])
print(model.predict(inp)[1][0,:7])
```
| github_jupyter |
<a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/master/tutorials/W2D3_BiologicalNeuronModels/student/W2D3_Intro.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Intro
**Our 2021 Sponsors, including Presenting Sponsor Facebook Reality Labs**
<p align='center'><img src='https://github.com/NeuromatchAcademy/widgets/blob/master/sponsors.png?raw=True'/></p>
## Overview
Today you will learn about a few interesting properties of biological neurons and synapses. In his intro lecture Upi Bhalla will start with an overview of the complexity of the neurons and synapses in the brain. He will also introduce a mathematical description of action potential generation and propagation by which neurons communicate with each other. Then, in a series of short tutorials Richard Naud will introduce simple neuron and synapse models. These tutorials will give you insights about how neurons may generate irregular spike patterns and synchronize their activity. In the first tutorial you will learn about the input-output transfer function of the leaky integrate and fire neuron model. In the second tutorial you will use this model to understand how statistics of inputs affects transfer of synchrony. In the third tutorial you will explore the short-term dynamics of synapses which means that synaptic weight is dependent on the recent history of spiking activity of the pre-synaptic neurons. In the fourth tutorial, you will learn about spike timing dependent plasticity and explore how synchrony in the input may shape the synaptic weight distribution. Finally, in the outro lecture Yiota Poirazi will explain how the simplified description of neurons can be expanded to include more biological complexity. She will provide evidence of how dendritic morphology may expand the computational repertoire of individual neurons.
The models we use in today’s lecture fall in the category of how models (W1D1). You will use several concepts from linear systems (W2D2). The insights developed in these tutorials will be useful to understand the dynamics of neural networks (W3D4). Moreover, you will learn about the origin of statistics of neuronal activity which will be useful for several tutorials. For example, the understanding of synchrony will be very useful in appreciating the problem of causality (W3D5).
Neuron and synapse models are essential building blocks of mechanistic models of brain function and dysfunction. One of the common questions in neuroscience is to identify the causes of changes in the statistics of spiking activity patterns. Whether these changes are caused by changes in neuron/synapse properties or by a change in the input or by a combination of both? With the contents of this tutorial, you should have a framework to think about which changes in spike patterns are due to neuron/synapse or input changes.
## Video
```
# @markdown
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV18A411v7Yy", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"MAOOPv3whZ0", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
```
## Slides
```
# @markdown
from IPython.display import IFrame
IFrame(src=f"https://mfr.ca-1.osf.io/render?url=https://osf.io/gyfr2/?direct%26mode=render%26action=download%26mode=render", width=854, height=480)
```
| github_jupyter |
### Heroes Of Pymoli Data Analysis
* Of the 1163 active players, the vast majority are male (84%). There also exists, a smaller, but notable proportion of female players (14%).
* Our peak age demographic falls between 20-24 (44.8%) with secondary groups falling between 15-19 (18.60%) and 25-29 (13.4%).
-----
### Note
* Instructions have been included for each segment. You do not have to follow them exactly, but they are included to help you think through the steps.
```
# Dependencies and Setup
import pandas as pd
# File to Load (Remember to Change These)
file_to_load = "Resources/purchase_data.csv"
# Read Purchasing File and store into Pandas data frame
purchase_data = pd.read_csv(file_to_load)
purchase_data
```
## Player Count
* Display the total number of players
```
Player_count = len(purchase_data["SN"].value_counts())
Player_count
```
## Purchasing Analysis (Total)
* Run basic calculations to obtain number of unique items, average price, etc.
* Create a summary data frame to hold the results
* Optional: give the displayed data cleaner formatting
* Display the summary data frame
```
# calculate unique items, average price, number of purchases, total revenue
Average_price=purchase_data["Price"].mean()
Number_of_Unique_Items=purchase_data["Item Name"].nunique()
Number_of_Purchases=purchase_data["Purchase ID"].count()
Total_Revenue=purchase_data["Price"].sum()
# create a summary data frame
Purchasing_Analysis=pd.DataFrame({"Number of Unique Items":[Number_of_Unique_Items],
"Average Price":[Average_price],
"Number of Purchases": [Number_of_Purchases],
"Total Revenue": [Total_Revenue]})
# formatting the data
Purchasing_Analysis["Average Price"] = Purchasing_Analysis["Average Price"].map("${:.2f}".format)
Purchasing_Analysis["Total Revenue"] = Purchasing_Analysis["Total Revenue"].map("${:.2f}".format)
# display the data
Purchasing_Analysis
```
## Gender Demographics
* Percentage and Count of Male Players
* Percentage and Count of Female Players
* Percentage and Count of Other / Non-Disclosed
```
# group purchase data by Gender
Gender_data= purchase_data.groupby(["Gender"])
count = purchase_data["SN"].value_counts()
# count number of male, female, and other/non-disclosed players
Gender_count=Gender_data.nunique()["SN"]
# calculate percentage of male, female, and other/non-disclosed players
percentage_player= Gender_count / Player_count * 100
# put all data in a dataframe
Gender_Demographics=pd.DataFrame({"Total Count": Gender_count, "Percentage of Players": percentage_player})
# formatting the data
Gender_Demographics["Percentage of Players"] = Gender_Demographics["Percentage of Players"].map("{:.2f}%".format)
#show dataframe
Gender_Demographics
```
## Purchasing Analysis (Gender)
* Run basic calculations to obtain purchase count, avg. purchase price, avg. purchase total per person etc. by gender
* Create a summary data frame to hold the results
* Optional: give the displayed data cleaner formatting
* Display the summary data frame
```
# count purchase count
purchase_count=Gender_data["Purchase ID"].count()
# avg. purchase price
average_purchase=Gender_data["Price"].mean()
# total purchase
total_purchase=Gender_data["Price"].sum()
# avg. purchase total per person
average_purchase_per_person= total_purchase/Gender_count
# create data frame
Purchasing_Analysis_Gender=pd.DataFrame({"Purchase Count": purchase_count,
"Average Purchase Price": average_purchase,
"Total Purchase Value":total_purchase,
"Avg Total Purchase per Person": average_purchase_per_person})
# format dataframe
Purchasing_Analysis_Gender["Average Purchase Price"] = Purchasing_Analysis_Gender["Average Purchase Price"].map("${:.2f}".format)
Purchasing_Analysis_Gender["Total Purchase Value"] = Purchasing_Analysis_Gender["Total Purchase Value"].map("${:.2f}".format)
Purchasing_Analysis_Gender["Avg Total Purchase per Person"] = Purchasing_Analysis_Gender["Avg Total Purchase per Person"].map("${:.2f}".format)
Purchasing_Analysis_Gender
```
## Age Demographics
* Establish bins for ages
* Categorize the existing players using the age bins. Hint: use pd.cut()
* Calculate the numbers and percentages by age group
* Create a summary data frame to hold the results
* Optional: round the percentage column to two decimal points
* Display Age Demographics Table
```
# Create the bins in which Data will be held
bins=[0, 9.9, 14, 19, 24, 29, 34, 39, 150]
# create labels for the bins
group_names = ["<10", "10-14", "15-19", "20-24", "25-29", "30-34", "35-39", "40+"]
# divide purchase data based on the bin and add a new column to the data frame
purchase_data["Age Group"]=pd.cut(purchase_data["Age"],bins, labels=group_names)
# creat new data frame group by age
age_grouped=purchase_data.groupby("Age Group")
# count total plaers by age
total_count_age = age_grouped["SN"].nunique()
# count player percentage based on age
percentage_by_age = (total_count_age/Player_count) * 100
# creat a data frame
Age_Demographics=pd.DataFrame({"Total Count": total_count_age, "Percentage of Players": percentage_by_age})
#format dataframe
Age_Demographics["Percentage of Players"] = Age_Demographics["Percentage of Players"].map("{:.2f}%".format)
Age_Demographics
```
## Purchasing Analysis (Age)
* Bin the purchase_data data frame by age
* Run basic calculations to obtain purchase count, avg. purchase price, avg. purchase total per person etc. in the table below
* Create a summary data frame to hold the results
* Optional: give the displayed data cleaner formatting
* Display the summary data frame
```
# purchase count based on age group
purchase_count_age=age_grouped["Purchase ID"].count()
# average purchase price by age group
average_purchase_price_age=age_grouped["Price"].mean()
# total purchase value by age group
total_purchase_value=age_grouped["Price"].sum()
# average purchase per person by age group
avg_purchase_per_person_age = total_purchase_value/total_count_age
# create data frame
purchase_analysis_age=pd.DataFrame({"Purchase Count": purchase_count_age,
"Average Purchase Price": average_purchase_price_age,
"Total Purchase Value":total_purchase_value,
"Avg Total Purchase per Person": avg_purchase_per_person_age})
# format dataframe
purchase_analysis_age["Average Purchase Price"] = purchase_analysis_age["Average Purchase Price"].map("${:.2f}".format)
purchase_analysis_age["Total Purchase Value"] =purchase_analysis_age["Total Purchase Value"].map("${:.2f}".format)
purchase_analysis_age["Avg Total Purchase per Person"] = purchase_analysis_age["Avg Total Purchase per Person"].map("${:.2f}".format)
purchase_analysis_age
```
## Top Spenders
* Run basic calculations to obtain the results in the table below
* Create a summary data frame to hold the results
* Sort the total purchase value column in descending order
* Optional: give the displayed data cleaner formatting
* Display a preview of the summary data frame
```
spender_groupby= purchase_data.groupby("SN")
spender_groupby
# Group purchase data by SN
spender_groupby= purchase_data.groupby("SN")
# Count the total purchases by SN
purchase_count_spender = spender_groupby["SN"].count()
# Calculate the average purchase price by SN
average_purchase_price_spender = spender_groupby["Price"].mean()
# Calculate total purchase
purchase_total_spender = spender_groupby["Price"].sum()
# Create data frame for the analysis data
spenders_analysis = pd.DataFrame({"Purchase Count": purchase_count_spender,
"Average Purchase Price": average_purchase_price_spender,
"Total Purchase Value":purchase_total_spender})
top_spenders = spenders_analysis.sort_values(["Total Purchase Value"], ascending=False)
# format dataframe
top_spenders["Average Purchase Price"] = top_spenders["Average Purchase Price"].map("${:.2f}".format)
top_spenders["Total Purchase Value"] =top_spenders["Total Purchase Value"].map("${:.2f}".format)
top_spenders.head()
```
## Most Popular Items
* Retrieve the Item ID, Item Name, and Item Price columns
* Group by Item ID and Item Name. Perform calculations to obtain purchase count, item price, and total purchase value
* Create a summary data frame to hold the results
* Sort the purchase count column in descending order
* Optional: give the displayed data cleaner formatting
* Display a preview of the summary data frame
```
Items_data=purchase_data("Item ID", "Item Name", "Price")
Items_data
#Retrieve the Item ID, Item Name, and Item Price columns
Items_data=purchase_data[["Item ID", "Item Name", "Price"]]
#Group by Item ID and Item Name.
Items_analysis=Items_data.groupby(["Item ID","Item Name"])
#Perform calculations to obtain purchase count, item price, and total purchase value
purchase_count_item = Items_analysis["Price"].count()
total_purchase_value = Items_analysis["Price"].sum()
item_price=total_purchase_value /purchase_count_item
# creat a data frame
Popular_items=pd.DataFrame({"Purchase Count": purchase_count_item,
"Item Price": item_price,
"Total Purchase Value":total_purchase_value})
# sort the items
most_poupular_items= Popular_items.sort_values(["Purchase Count"], ascending=False)
# format dataframe
most_poupular_items["Item Price"] = most_poupular_items["Item Price"].map("${:.2f}".format)
most_poupular_items["Total Purchase Value"] =most_poupular_items["Total Purchase Value"].map("${:.2f}".format)
most_poupular_items.head()
```
## Most Profitable Items
* Sort the above table by total purchase value in descending order
* Optional: give the displayed data cleaner formatting
* Display a preview of the data frame
```
#sort the items
most_profitable_items= Popular_items.sort_values(["Total Purchase Value"], ascending=False)
# format dataframe
most_profitable_items["Item Price"] = most_profitable_items["Item Price"].map("${:.2f}".format)
most_profitable_items["Total Purchase Value"] =most_profitable_items["Total Purchase Value"].map("${:.2f}".format)
most_profitable_items.head()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/bereml/iap/blob/master/libretas/1f_fashion_fcn.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Clasificación de Fashion-MNIST con una red densa
Curso: [Introducción al Aprendizaje Profundo](http://turing.iimas.unam.mx/~ricardoml/course/iap/). Profesores: [Bere](https://turing.iimas.unam.mx/~bereml/) y [Ricardo](https://turing.iimas.unam.mx/~ricardoml/) Montalvo Lezama.
---
---
En esta libreta debes entrenar dos clasificadores para el conjunto Fashion-MNIST.
1. El primero usando la misma arquitectura e hiperparámetros que en el ejemplo de MNIST.
2. En un segundo clasificador modifica la arquitectura intentando obtener un mejor desempeño.
Para resolver este ejercicio emplea la clase [`FashionMNIST`](https://pytorch.org/vision/0.8/datasets.html#fashion-mnist) proporcionada por PyTorch.
[Fashion-MNIST](https://github.com/zalandoresearch/fashion-mnist) es un conjunto para remplzar MNIST. Fue recolectado con la intención de proveer un conjunto un poco más dificil que MNIST.
<img src="https://miro.medium.com/max/800/1*RNBs0OsymwAzDyYMk3_0Aw.jpeg" width="600"/>
Conjunto Fashion-MNIST. Imagen tomada de https://medium.com/@sankarchanna2k18/fashion-mnist-data-image-classification-in-tensorflow-bd22f9e680bc.
## 1 Preparación
```
# biblioteca para inspeccionar arquitecturas
# https://github.com/tyleryep/torchinfo
!pip install torchinfo
```
### 1.1 Bibliotecas
```
# funciones aleatorias
import random
# tomar n elementos de una secuencia
from itertools import islice as take
# gráficas
import matplotlib.pyplot as plt
# arreglos multidimensionales
import numpy as np
# redes neuronales
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
import torchvision.transforms as T
# procesamiento de imágenes
from skimage import io
# redes neuronales
from torch.utils.data import DataLoader
from torchvision.datasets import FashionMNIST
# inspección de arquitectura
from torchinfo import summary
# barras de progreso
from tqdm import trange
```
### 1.2 Auxiliares
```
# directorio de datos
DATA_DIR = '../data'
# tamaño del lote
BATCH_SIZE = 32
# filas y columnas de la regilla de imágenes
ROWS, COLS = 4, 8
# Fashion-MNIST classes
CLASSES = {
0: "T-shirt/top",
1: "Trouser",
2: "Pullover",
3: "Dress",
4: "Coat",
5: "Sandal",
6: "Shirt",
7: "Sneaker",
8: "Bag",
9: "Ankle boot",
}
def display_grid(xs, titles, rows, cols, figsize=(12, 6)):
"""Displays examples in a grid."""
fig, ax = plt.subplots(rows, cols, figsize=figsize)
i = 0
for r in range(rows):
for c in range(cols):
ax[r, c].imshow(xs[i], cmap='gray')
ax[r, c].set_title(titles[i])
ax[r, c].set_xticklabels([])
ax[r, c].set_yticklabels([])
i += 1
fig.tight_layout()
plt.show()
def display_batch(x, titles, rows, cols, figsize=(12, 6)):
"""Displays a batch of processed examples in a grid."""
# denormalizamos [0, 1] => [0, 255]
x *= 255
# rotamos canales (C x H x W) => (H x W x C)
x = x.permute(0, 2, 3, 1)
# convertimos a entero
x = (x.numpy()).astype(np.uint8)
# aplanamos canal
x = x.reshape(*x.shape[:3])
# desplegamos
display_grid(x, titles, rows, cols, figsize)
def set_seed(seed=0):
"""Initializes pseudo-random number generators."""
random.seed(seed)
np.random.seed(seed)
torch.manual_seed(seed)
# reproducibilidad
set_seed()
```
## 2 Datos
### 2.1 Tuberias de datos con PyTorch
<img src="https://raw.githubusercontent.com/bereml/iap/master/fig/mnist_pipeline.png"/>
Tuberia de datos para MNIST.
### 2.2 Exploración
### 2.3 Cargadores de datos
<img src="https://raw.githubusercontent.com/bereml/iap/master/fig/eval_trn_tst.svg" width="450"/>
Entrenamiento con una partición de entrenamiento y prueba.
#### Entrenamiento
#### Prueba
## 3 Modelo
<img src="https://raw.githubusercontent.com/bereml/iap/master/fig/fcn_arch.png"/>
Arquitectura de la red completamente conectada.
### 3.1 Definición de la arquitectura
### 3.2 Instancia de la arquitectura
### 3.3 Inspección de la arquitectura
## 4 Entrenamiento
<img src="https://raw.githubusercontent.com/bereml/iap/master/fig/supervisado.svg" width="700"/>
Ciclo de entrenamiento supervisado.
### 4.1 Ciclo de entrenamiento
Entrenamos un modelo:
### 4.2 Gráficas de pérdidas y exactitud
## 5 Evaluación
<img src="https://raw.githubusercontent.com/bereml/iap/master/fig/eval_trn_tst.svg" width="450"/>
Entrenamiento con una partición de entrenamiento y prueba.
### 5.1 Evaluación final
### 5.2 Inferencia
| github_jupyter |
```
from collections import defaultdict, OrderedDict
import warnings
import gffutils
import pybedtools
import pandas as pd
import copy
import os
import re
from gffutils.pybedtools_integration import tsses
from copy import deepcopy
from collections import OrderedDict, Callable
import errno
def mkdir_p(path):
try:
os.makedirs(path)
except OSError as exc: # Python >2.5
if exc.errno == errno.EEXIST and os.path.isdir(path):
pass
else:
raise
class DefaultOrderedDict(OrderedDict):
# Source: http://stackoverflow.com/a/6190500/562769
def __init__(self, default_factory=None, *a, **kw):
if (default_factory is not None and
not isinstance(default_factory, Callable)):
raise TypeError('first argument must be callable')
OrderedDict.__init__(self, *a, **kw)
self.default_factory = default_factory
def __getitem__(self, key):
try:
return OrderedDict.__getitem__(self, key)
except KeyError:
return self.__missing__(key)
def __missing__(self, key):
if self.default_factory is None:
raise KeyError(key)
self[key] = value = self.default_factory()
return value
def __reduce__(self):
if self.default_factory is None:
args = tuple()
else:
args = self.default_factory,
return type(self), args, None, None, self.items()
def copy(self):
return self.__copy__()
def __copy__(self):
return type(self)(self.default_factory, self)
def __deepcopy__(self, memo):
import copy
return type(self)(self.default_factory,
copy.deepcopy(self.items()))
def __repr__(self):
return 'OrderedDefaultDict(%s, %s)' % (self.default_factory,
OrderedDict.__repr__(self))
gtf = '/panfs/qcb-panasas/skchoudh/genomes/hg38/annotation/Homo_sapiens.GRCh38.96.gtf'
gtf_db = '/panfs/qcb-panasas/skchoudh/genomes/hg38/annotation/Homo_sapiens.GRCh38.96.gtf.db'
prefix = '/panfs/qcb-panasas/skchoudh/github_projects/riboraptor/riboraptor/annotation/hg38/v96/'
chrsizes = '/panfs/qcb-panasas/skchoudh/genomes/hg38/fasta/hg38.chrom.sizes'
mkdir_p(prefix)
def create_gene_dict(db):
'''
Store each feature line db.all_features() as a dict of dicts
'''
gene_dict = DefaultOrderedDict(lambda: DefaultOrderedDict(lambda: DefaultOrderedDict(list)))
for line_no, feature in enumerate(db.all_features()):
gene_ids = feature.attributes['gene_id']
feature_type = feature.featuretype
if feature_type == 'gene':
if len(gene_ids)!=1:
logging.warning('Found multiple gene_ids on line {} in gtf'.format(line_no))
break
else:
gene_id = gene_ids[0]
gene_dict[gene_id]['gene'] = feature
else:
transcript_ids = feature.attributes['transcript_id']
for gene_id in gene_ids:
for transcript_id in transcript_ids:
gene_dict[gene_id][transcript_id][feature_type].append(feature)
return gene_dict
db = gffutils.create_db(gtf, dbfn=gtf_db,
merge_strategy='merge',
force=True,
disable_infer_transcripts=True,
disable_infer_genes=True)
db = gffutils.FeatureDB(gtf_db, keep_order=True)
gene_dict = create_gene_dict(db)
def get_gene_list(gene_dict):
return list(set(gene_dict.keys()))
def get_UTR_regions(utrs, cds):
if len(cds)==0:
return [], []
utr5_regions = []
utr3_regions = []
cds_sorted = sorted(list(cds), key=lambda x: x.start)
first_cds = cds_sorted[0]
last_cds = cds_sorted[-1]
for orig_utr in utrs:
utr = deepcopy(orig_utr)
## Push all cds at once
## Sort later to remove duplicates
strand = utr.strand
if utr.start < first_cds.start:
if utr.stop >= first_cds.start:
utr.stop = first_cds.start - 1
if strand == '+':
utr5_regions.append(utr)
else:
utr3_regions.append(utr)
elif utr.stop > last_cds.stop:
if utr.start <= last_cds.stop:
utr.start = last_cds.stop + 1
if strand == '+':
utr3_regions.append(utr)
else:
utr5_regions.append(utr)
return utr5_regions, utr3_regions
def create_bed(regions, bedtype='0'):
'''Create bed from list of regions
bedtype: 0 or 1
0-Based or 1-based coordinate of the BED
'''
bedstr = ''
for region in regions:
assert len(region.attributes['gene_id']) == 1
## GTF start is 1-based, so shift by one while writing
## to 0-based BED format
if bedtype == '0':
start = region.start - 1
else:
start = region.start
bedstr += '{}\t{}\t{}\t{}\t{}\t{}\n'.format(region.chrom,
start,
region.stop,
re.sub('\.\d+', '', region.attributes['gene_id'][0]),
'.',
region.strand)
return bedstr
def rename_regions(regions, gene_id):
regions = list(regions)
if len(regions) == 0:
return []
for region in regions:
region.attributes['gene_id'] = gene_id
return regions
def merge_regions(db, regions):
if len(regions) == 0:
return []
merged = db.merge(sorted(list(regions), key=lambda x: x.start))
return merged
def merge_regions_nostrand(db, regions):
if len(regions) == 0:
return []
merged = db.merge(sorted(list(regions), key=lambda x: x.start), ignore_strand=True)
return merged
utr5_bed = ''
utr3_bed = ''
gene_bed = ''
exon_bed = ''
intron_bed = ''
start_codon_bed = ''
stop_codon_bed = ''
cds_bed = ''
gene_list = []
for gene_id in get_gene_list(gene_dict):
gene_list.append(gene_dict[gene_id]['gene'])
utr5_regions, utr3_regions = [], []
exon_regions, intron_regions = [], []
star_codon_regions, stop_codon_regions = [], []
cds_regions = []
utr_regions = []
for feature in gene_dict[gene_id].keys():
if feature == 'gene':
continue
cds = list(gene_dict[gene_id][feature]['CDS'])
exons = list(gene_dict[gene_id][feature]['exon'])
utrs = list(gene_dict[gene_id][feature]['UTR'])
cds = sorted(list(cds), key=lambda x: x.start)
exons = sorted(list(exons), key=lambda x: x.start)
utrs = sorted(list(utrs), key=lambda x: x.start)
merged_exons = merge_regions(db, exons)
introns = db.interfeatures(merged_exons)
exon_regions += exons
intron_regions += introns
cds_regions += cds
utr_regions += utrs
cds_regions = sorted(list(cds_regions), key=lambda x: x.start)
utr_regions = sorted(list(utr_regions), key=lambda x: x.start)
utr5_regions, utr3_regions = get_UTR_regions(utr_regions, cds_regions)
merged_utr5 = merge_regions(db, utr5_regions)
renamed_utr5 = rename_regions(merged_utr5, gene_id)
merged_utr3 = merge_regions(db, utr3_regions)
renamed_utr3 = rename_regions(merged_utr3, gene_id)
merged_exons = merge_regions(db, exon_regions)
renamed_exons = rename_regions(merged_exons, gene_id)
merged_introns = merge_regions(db, intron_regions)
renamed_introns = rename_regions(merged_introns, gene_id)
merged_cds = merge_regions(db, cds_regions)
renamed_cds = rename_regions(merged_cds, gene_id)
utr3_bed += create_bed(renamed_utr3)
utr5_bed += create_bed(renamed_utr5)
exon_bed += create_bed(renamed_exons)
intron_bed += create_bed(renamed_introns)
cds_bed += create_bed(renamed_cds)
gene_bed = create_bed(gene_list)
gene_bedtool = pybedtools.BedTool(gene_bed, from_string=True)
utr5_bedtool = pybedtools.BedTool(utr5_bed, from_string=True)
utr3_bedtool = pybedtools.BedTool(utr3_bed, from_string=True)
exon_bedtool = pybedtools.BedTool(exon_bed, from_string=True)
intron_bedtool = pybedtools.BedTool(intron_bed, from_string=True)
cds_bedtool = pybedtools.BedTool(cds_bed, from_string=True)
utr5_cds_subtracted = utr5_bedtool.subtract(cds_bedtool)
utr3_cds_subtracted = utr3_bedtool.subtract(cds_bedtool)
utr5_cds_subtracted.remove_invalid().sort().saveas(os.path.join(prefix, 'utr5.bed.gz'))
utr3_cds_subtracted.remove_invalid().sort().saveas(os.path.join(prefix, 'utr3.bed.gz'))
gene_bedtool.remove_invalid().sort().saveas(os.path.join(prefix, 'gene.bed.gz'))
exon_bedtool.remove_invalid().sort().saveas(os.path.join(prefix, 'exon.bed.gz'))
intron_bedtool.remove_invalid().sort().saveas(os.path.join(prefix, 'intron.bed.gz'))
cds_bedtool.remove_invalid().sort().saveas(os.path.join(prefix, 'cds.bed.gz'))
for gene_id in get_gene_list(gene_dict):
start_codons = []
stop_codons = []
for start_codon in db.children(gene_id, featuretype='start_codon'):
## 1 -based stop
## 0-based start handled while converting to bed
start_codon.stop = start_codon.start
start_codons.append(start_codon)
for stop_codon in db.children(gene_id, featuretype='stop_codon'):
stop_codon.start = stop_codon.stop
stop_codon.stop = stop_codon.stop+1
stop_codons.append(stop_codon)
merged_start_codons = merge_regions(db, start_codons)
renamed_start_codons = rename_regions(merged_start_codons, gene_id)
merged_stop_codons = merge_regions(db, stop_codons)
renamed_stop_codons = rename_regions(merged_stop_codons, gene_id)
start_codon_bed += create_bed(renamed_start_codons)
stop_codon_bed += create_bed(renamed_stop_codons)
start_codon_bedtool = pybedtools.BedTool(start_codon_bed, from_string=True)
stop_codon_bedtool = pybedtools.BedTool(stop_codon_bed, from_string=True)
start_codon_bedtool.remove_invalid().sort().saveas(os.path.join(prefix, 'start_codon.bed.gz'))
stop_codon_bedtool.remove_invalid().sort().saveas(os.path.join(prefix, 'stop_codon.bed.gz'))
stop_codon_bedtool
tss = tsses(db, as_bed6=True, merge_overlapping=True)
tss.remove_invalid().sort().saveas(os.path.join(prefix, 'tss.bed'))
promoter = tss.slop(l=1500, r=1500, s=True, g=chrsizes)
promoter.remove_invalid().sort().saveas(os.path.join(prefix, 'promoter.1500.bed.gz'))
promoter.to_dataframe().head()
```
| github_jupyter |
```
######### IMPORT ###########
from collections import deque
import random
import pandas as pd # package for some uses and maybe for visualization
import numpy as np
import heapq
from matplotlib.pylab import plot, show, bar
import matplotlib.pyplot as plt
plt.style.use("ggplot")
df = pd.read_csv('arrival_rates.csv')
df
floors_num = 25
arrival_rates_by_floor_section = [
[18, 10.8, 40.8], [50, 6.8, 12.8], [11, 4.8, 7.8]]
arrival_rates = [
[[], []],
[[], [], []],
[[], [], []]
]
k = -1
for i in range(3):
for j in range(len(arrival_rates[i])):
k += 1
if k == 8:
break
opc = [df.loc[k, "other"], df.loc[k, "15--18"], df.loc[k, "7--10"]]
arrival_rates[i][j].extend(opc)
arrival_rates[0].insert(0, None)
# Creating a dictionary with the range of each floor
# the range ressembles the location in the rate array
lower_floor_range = {floor: 1 for floor in range(1, 16)}
upper_floor_range = {floor: 2 for floor in range(16, 26)}
lower_floor_range.update(upper_floor_range)
floor_range = lower_floor_range
floor_range[0] = 0 # ground floor
######### CLASS ###########
class Event():
def __init__(self, time, eventType, passenger=-1, elevator=-1):
self.time = time # event time
self.eventType = eventType # type of the event
self.passenger = passenger
self.elevator = elevator
heapq.heappush(P, self) # add the event to the events list
def __lt__(self, event2):
return self.time < event2.time
########################
#### Passenger class ###
########################
### VARIABLES AND FUNCTIONS TO REMOVE
#in_journey, is_top_elevator, min_floor,top_floor ,move function,release_passengers function, L_up[0]
class Passenger(object):
def __init__(self, start, end, arrival_time):
self.id = passenger_count # unique identifier
self.start = start
self.end = end
self.arrival_time = arrival_time # time of first arrival
# self.current_floor = floor
#self.in_journey = (start != 0 and end != 0) and ((
#start > 15 and end <= 15) or (start <= 15 and end > 15))
# self.journey_started = False # describes the current part of the journey. Either 1 or 2
self.left = False
self.started_using_sys = False
#if self.in_journey:
# in journey
# always going down firt to floor 0
#self.direction = -1
# not in journey
if end > start:
self.direction = 1
else:
self.direction = -1
class Elevator(object):
def __init__(self, id, starting_floor, direction, top_floor):
self.id = id
self.curr_floor = starting_floor
self.prev_floor = starting_floor
self.stop_time = curr_time # last stop time
self.passengers = []
self.max_capacity = 15
self.remaining_space = self.max_capacity - len(self.passengers)
self.direction = direction # up / down
#self.is_top_elevator = top_floor == 25
self.top_floor = top_floor
#if top_floor == 15:
#self.min_floor = 0 new addition maybe needed
#self.min_floor = 1 # lowest floor in section
#else:
#self.min_floor = 16 # lowest floor in section
self.is_broken = False # elevator starts un-broken
self.last_broken_time = -1 # time of last break
# Gets the elevator destination. Checks if it was broken until arriving
def update_space(self):
self.remaining_space = self.max_capacity - len(self.passengers)
def move(self):
self.prev_floor = self.curr_floor
#if self.is_top_elevator:
#if self.curr_floor == 16 and self.direction == -1:
#self.curr_floor -= 15 # move the elevator past sector #1
#elif self.curr_floor == 0 and self.direction == 1:
#self.curr_floor += 15 # move the elevator past sector #1
self.curr_floor += self.direction
#print("prev_floor: ",self.prev_floor,"curr_floor: ",self.curr_floor)
if (self.curr_floor == 0 and self.direction == -1) or (self.curr_floor == 25 and self.direction == 1):
# if reached end of section, flip direction
self.direction *= -1
def did_break(self):
if self.is_broken:
# it's now fixed after being broken
self.is_broken = False
return False
else:
# elevator has been working until now
rnd = np.random.random()
if rnd < 0.0005:
# and now it broke
self.is_broken = True
self.last_broken_time = curr_time
return True
else:
# still not broken
return False
def switch_direction(self):
self.direction *= -1
# removes leaving passengers and returns journey passengers
def release_passengers(self):
# count the passengers just before they leave
elevator_usage[self.id -
1][len(self.passengers)] += curr_time - self.stop_time
elevator_usage_by_day[i][self.id -
1][len(self.passengers)] += curr_time - self.stop_time
leaving_passengers = list(
[p for p in self.passengers if self.curr_floor == p.end]) # for visualization
for p in leaving_passengers:
service_time.append(curr_time - p.arrival_time)
service_time_by_day[i].append(curr_time - p.arrival_time)
#journey_passengers = []
#if self.curr_floor == 0:
# journey passengers are saved in the system
# journey_passengers = list(
# [p for p in self.passengers if p.in_journey])
#self.passengers = list(
# [p for p in self.passengers if self.curr_floor != p.end and not p.in_journey])
#else:
# create new passengers list with all the passengers that stay
self.passengers = list(
[p for p in self.passengers if self.curr_floor != p.end])
#return journey_passengers
# removes passengers in the broken elevator to the other elevator
def release_when_broken(self, direction):
# count the passengers before they leave
elevator_usage[self.id -
1][len(self.passengers)] += curr_time - self.stop_time
elevator_usage_by_day[i][self.id -
1][len(self.passengers)] += curr_time - self.stop_time
# release only passengers that go to the same direction as the rescue elevator
released = list(
[p for p in self.passengers if self.direction == direction])
self.passengers = list(
[p for p in self.passengers if self.curr_floor != p.end and not self.direction == direction])
return released
######### FUNCTION ###########
## returns current rate ##
##########################
def get_current_rate_by_floor(start_floor, end_floor):
# get current arrival rate according to start floor,
# destination, (or section) and curr_time(which would be determined
# at the start of each While iteration)
start_floor_section = floor_range[start_floor]
end_floor_section = floor_range[end_floor]
if curr_time >= 60*60 and curr_time < 4*60*60:
# 07:00-10:00
hour_range = 2
elif curr_time >= 9*60*60 and curr_time < 12*60*60:
# 15:00-18:00
hour_range = 1
else:
# any other time
hour_range = 0
rate = arrival_rates[start_floor_section][end_floor_section][hour_range]
# Convert to exponential rate
return (60*60)/rate
def get_travel_time(floors):
start = 2
stop = 2
wait = 5
pass_by = 1
return floors * pass_by + start + stop + wait
#### Performance Measures ####
n = 5 # intervals
service_time = [] # for visualization - service time for each passenger
service_time_by_day = i = [ [] for _ in range(n) ] # for visualization - service time for each passenger
avg_service = [] # for visualization - avg service duration
avg_out_of_patience = [0]*n # out of patience per day
elevator_usage = [[0]*16, [0]*16, [0]*16, [0]*16]
elevator_usage_by_day = [ [[0]*16, [0]*16, [0]*16, [0]*16] for _ in range(n)]
##############################
for i in range(n):
seed_1 = np.random.RandomState(i)
seed_2 = np.random.RandomState(i+100)
######### INITIATE ###########
# initialize simulation
curr_time = 0 # current time
SIM_TIME = 14*60*60 # simulation time in minutes
P = [] # heap
L_up = [[] for _ in range(26)] # going up line in every floor
# at floor zero we have 2 seperate lines for the 2 pairs of elevator
#L_up[0] = [[], []]
L_down = [[] for _ in range(26)] # going down line in every floor
passenger_count = 0
# create list of elevators
elevators = [Elevator(id=1, starting_floor=0, direction=1, top_floor=25), Elevator(id=2, starting_floor=7, direction=1, top_floor=25), Elevator(
id=3, starting_floor=14, direction=-1, top_floor=25), Elevator(id=4, starting_floor=25, direction=-1, top_floor=25)]
for start in range(0, 26):
for end in range(0, 26):
if start == end:
continue
time = seed_1.exponential(get_current_rate_by_floor(start, end))
new_passenger = Passenger(start, end, time)
passenger_count += 1
Event(time, "arriving", passenger=new_passenger)
for elevator in elevators:
# init elevators
Event(curr_time + 5, "elevator_close", elevator=elevator)
######### LOOP ###########
while curr_time < SIM_TIME: # loop until sim time ends
event = heapq.heappop(P) # get next event
curr_time = event.time # current event's time
## arriving ##
if event.eventType == "arriving":
passenger = event.passenger
# insert passenger to floor line
if passenger.direction == 1:
#if passenger.start == 0:###########################CHANGE HERE
# passenger is going up on floor zero
#heapq.heappush(
#L_up[0][1 if passenger.end > 15 else 0], (passenger.arrival_time, id(passenger), passenger))
#else:
heapq.heappush(
L_up[passenger.start], (passenger.arrival_time, id(passenger), passenger))
else:
heapq.heappush(L_down[passenger.start],
(passenger.arrival_time, id(passenger), passenger))
# check if he will be out of patience
Event(curr_time + 15*60, "out_of_patience", passenger)
# generate next passenger
rate = get_current_rate_by_floor(passenger.start, passenger.end)
new_time = curr_time + seed_1.exponential(rate)
new_passenger = Passenger(passenger.start, passenger.end, new_time)
passenger_count += 1
Event(new_time, "arriving", passenger=new_passenger)
##############
## elevator_close ##
elif event.eventType == "elevator_close":
elevator = event.elevator
is_broken = elevator.did_break()
# passengers with this destination are free to go, returns journey passengers in floor 0
elevator.release_passengers() ###########################CHANGE HERE
#if elevator.curr_floor == 0:
# if in floor 0, add journey passengers to queue
#for j_passenger in journey_passengers:
# add journey passengers back to the line
# they always go up if in middle of journey
#j_passenger.in_journey = False
#j_passenger.direction = 1
#heapq.heappush(L_up[0][1 if j_passenger.end > 15 else 0],
#(curr_time - 5, id(j_passenger), j_passenger))
# pull passengers from broken elevator
for other_elevator in elevators:
if other_elevator != elevator:
if other_elevator.is_broken and other_elevator.curr_floor == elevator.curr_floor and other_elevator.direction == elevator.direction:
# other elevator is stuck right now on same floor and with same direction
# and also the current elevator didnt break
# therefore we will take any passenger we can with us
released_from_broken = other_elevator.release_when_broken(
elevator.direction)
# add another counting point for passengers leaving to another elevator
other_elevator.stop_time = curr_time
for p in released_from_broken:
elevator.update_space()
if elevator.remaining_space > 0:
# both are going in the same direction
# passenger enters the elevator
# appends randomly
elevator.passengers.append(p)
else:
# return these passengers back to home elevator
other_elevator.passengers.append(p)
if is_broken:
# handle elevator broken
fix_time = curr_time + seed_2.uniform(5, 15)*60
Event(fix_time, "elevator_close", elevator=elevator)
# if elevator is stuck, it won't be moving
else: # not stuck
#print(elevator.curr_floor)
if elevator.direction == 1: # list of customers in line
#if elevator.curr_floor == 0: # if floor is zero - we have 2 seperate up lines ###########################CHANGE HERE
#waiting_passengers = L_up[0][1 if elevator.is_top_elevator else 0]
#else:
waiting_passengers = L_up[elevator.curr_floor]
else:
waiting_passengers = L_down[elevator.curr_floor]
#print(elevator.curr_floor)
while waiting_passengers:
elevator.update_space()
if elevator.remaining_space > 0:
# both are going in the same direction
# passenger enters the elevator
# returns the passenger from tuple
next_in_line = heapq.heappop(waiting_passengers)[2]
# passengers started using the elevators and wont run out of patience
next_in_line.started_using_sys = True
elevator.passengers.append(next_in_line)
else:
break # stop inserting passengers to elevator, no room or no more passengers
#if (elevator.curr_floor == 16 and elevator.direction == -1) or (elevator.curr_floor == 0 and elevator.direction == 1 and elevator.is_top_elevator): ###########################CHANGE HERE
#next_time = curr_time + get_travel_time(floors=16)
#else:
next_time = curr_time + get_travel_time(floors=1)
# if not broken, move as scheduled to next floor
Event(next_time, "elevator_close", elevator=elevator)
elevator.stop_time = curr_time
elevator.move() # moves elevator to next floor + open + close doors
## out_of_patience ##
elif event.eventType == "out_of_patience":
passenger = event.passenger
if passenger.left is False and passenger.started_using_sys is False:
passenger.left = True
# will always be next passenger
if passenger.direction == 1:
#if passenger.start == 0: ###########################CHANGE HERE
#for p_tuple in L_up[0][1 if passenger.end > 15 else 0]:
#if p_tuple[2] == passenger:
#L_up[0][1 if passenger.end >
#15 else 0].remove(p_tuple)
#heapq.heapify(L_up[passenger.start][1 if passenger.end >
#15 else 0])
for p_tuple in L_up[passenger.start]:
if p_tuple[2] == passenger:
L_up[passenger.start].remove(p_tuple)
heapq.heapify(L_up[passenger.start])
else:
for p_tuple in L_down[passenger.start]:
if p_tuple[2] == passenger:
L_down[passenger.start].remove(p_tuple)
heapq.heapify(L_down[passenger.start])
avg_out_of_patience[i] += 1 # for visualization
service_time.append(curr_time - passenger.arrival_time)
service_time_by_day[i].append(curr_time - passenger.arrival_time)
##############
cummulative_day = []
for day in service_time_by_day:
cummulative_day.append(sum(day) / len(day))
print(cummulative_day)
lypo = pd.Series(service_time)
fig = lypo.hist(bins = 100,edgecolor='k',figsize=(12,6)).set_ylabel("service time distribution")
### Average of out of patience passengers ###
needed_average = sum(avg_out_of_patience)/len(avg_out_of_patience)
print("Average of out of patience passengers: ",needed_average)
mat1 = []
mat2 = []
mat3 = []
mat4 = []
for i in range(16):
mat1.append([i,elevator_usage[0][i]])
for i in range(16):
mat2.append([i,elevator_usage[1][i]])
for i in range(16):
mat3.append([i,elevator_usage[2][i]])
for i in range(16):
mat4.append([i,elevator_usage[3][i]])
df1 = pd.DataFrame(mat1, columns=["num_of_people", "time"]) # hold the data in a pandas Data Frame object
title='elevator1'
df1.plot.bar(x="num_of_people", y="time",figsize=(12,3), title=title); #bar plot
print("__________________")
print(avg_out_of_patience)
df2 = pd.DataFrame(mat2, columns=["num_of_people", "time"]) # hold the data in a pandas Data Frame object
title='elevator2'
df2.plot.bar(x="num_of_people", y="time",figsize=(12,3), title=title); #bar plot
df3 = pd.DataFrame(mat3, columns=["num_of_people", "time"]) # hold the data in a pandas Data Frame object
title='elevator3'
df3.plot.bar(x="num_of_people", y="time",figsize=(12,3), title=title); #bar plot
df4 = pd.DataFrame(mat4, columns=["num_of_people", "time"]) # hold the data in a pandas Data Frame object
title='elevator4'
df4.plot.bar(x="num_of_people", y="time",figsize=(12,3), title=title); #bar plo
```
| github_jupyter |
# Variance Component Analysis
This notebook illustrates variance components analysis for two-level
nested and crossed designs.
```
import numpy as np
import statsmodels.api as sm
from statsmodels.regression.mixed_linear_model import VCSpec
import pandas as pd
```
Make the notebook reproducible
```
np.random.seed(3123)
```
## Nested analysis
In our discussion below, "Group 2" is nested within "Group 1". As a
concrete example, "Group 1" might be school districts, with "Group
2" being individual schools. The function below generates data from
such a population. In a nested analysis, the group 2 labels that
are nested within different group 1 labels are treated as
independent groups, even if they have the same label. For example,
two schools labeled "school 1" that are in two different school
districts are treated as independent schools, even though they have
the same label.
```
def generate_nested(n_group1=200, n_group2=20, n_rep=10, group1_sd=2,
group2_sd=3, unexplained_sd=4):
# Group 1 indicators
group1 = np.kron(np.arange(n_group1), np.ones(n_group2 * n_rep))
# Group 1 effects
u = group1_sd * np.random.normal(size=n_group1)
effects1 = np.kron(u, np.ones(n_group2 * n_rep))
# Group 2 indicators
group2 = np.kron(np.ones(n_group1), np.kron(np.arange(n_group2), np.ones(n_rep)))
# Group 2 effects
u = group2_sd * np.random.normal(size=n_group1*n_group2)
effects2 = np.kron(u, np.ones(n_rep))
e = unexplained_sd * np.random.normal(size=n_group1 * n_group2 * n_rep)
y = effects1 + effects2 + e
df = pd.DataFrame({"y":y, "group1": group1, "group2": group2})
return df
```
Generate a data set to analyze.
```
df = generate_nested()
```
Using all the default arguments for `generate_nested`, the population
values of "group 1 Var" and "group 2 Var" are 2^2=4 and 3^2=9,
respectively. The unexplained variance, listed as "scale" at the
top of the summary table, has population value 4^2=16.
```
model1 = sm.MixedLM.from_formula("y ~ 1", re_formula="1", vc_formula={"group2": "0 + C(group2)"},
groups="group1", data=df)
result1 = model1.fit()
print(result1.summary())
```
If we wish to avoid the formula interface, we can fit the same model
by building the design matrices manually.
```
def f(x):
n = x.shape[0]
g2 = x.group2
u = g2.unique()
u.sort()
uv = {v: k for k, v in enumerate(u)}
mat = np.zeros((n, len(u)))
for i in range(n):
mat[i, uv[g2[i]]] = 1
colnames = ["%d" % z for z in u]
return mat, colnames
```
Then we set up the variance components using the VCSpec class.
```
vcm = df.groupby("group1").apply(f).to_list()
mats = [x[0] for x in vcm]
colnames = [x[1] for x in vcm]
names = ["group2"]
vcs = VCSpec(names, [colnames], [mats])
```
Finally we fit the model. It can be seen that the results of the
two fits are identical.
```
oo = np.ones(df.shape[0])
model2 = sm.MixedLM(df.y, oo, exog_re=oo, groups=df.group1, exog_vc=vcs)
result2 = model2.fit()
print(result2.summary())
```
## Crossed analysis
In a crossed analysis, the levels of one group can occur in any
combination with the levels of the another group. The groups in
Statsmodels MixedLM are always nested, but it is possible to fit a
crossed model by having only one group, and specifying all random
effects as variance components. Many, but not all crossed models
can be fit in this way. The function below generates a crossed data
set with two levels of random structure.
```
def generate_crossed(n_group1=100, n_group2=100, n_rep=4, group1_sd=2,
group2_sd=3, unexplained_sd=4):
# Group 1 indicators
group1 = np.kron(np.arange(n_group1, dtype=np.int),
np.ones(n_group2 * n_rep, dtype=np.int))
group1 = group1[np.random.permutation(len(group1))]
# Group 1 effects
u = group1_sd * np.random.normal(size=n_group1)
effects1 = u[group1]
# Group 2 indicators
group2 = np.kron(np.arange(n_group2, dtype=np.int),
np.ones(n_group2 * n_rep, dtype=np.int))
group2 = group2[np.random.permutation(len(group2))]
# Group 2 effects
u = group2_sd * np.random.normal(size=n_group2)
effects2 = u[group2]
e = unexplained_sd * np.random.normal(size=n_group1 * n_group2 * n_rep)
y = effects1 + effects2 + e
df = pd.DataFrame({"y":y, "group1": group1, "group2": group2})
return df
```
Generate a data set to analyze.
```
df = generate_crossed()
```
Next we fit the model, note that the `groups` vector is constant.
Using the default parameters for `generate_crossed`, the level 1
variance should be 2^2=4, the level 2 variance should be 3^2=9, and
the unexplained variance should be 4^2=16.
```
vc = {"g1": "0 + C(group1)", "g2": "0 + C(group2)"}
oo = np.ones(df.shape[0])
model3 = sm.MixedLM.from_formula("y ~ 1", groups=oo, vc_formula=vc, data=df)
result3 = model3.fit()
print(result3.summary())
```
If we wish to avoid the formula interface, we can fit the same model
by building the design matrices manually.
```
def f(g):
n = len(g)
u = g.unique()
u.sort()
uv = {v: k for k, v in enumerate(u)}
mat = np.zeros((n, len(u)))
for i in range(n):
mat[i, uv[g[i]]] = 1
colnames = ["%d" % z for z in u]
return [mat], [colnames]
vcm = [f(df.group1), f(df.group2)]
mats = [x[0] for x in vcm]
colnames = [x[1] for x in vcm]
names = ["group1", "group2"]
vcs = VCSpec(names, colnames, mats)
```
Here we fit the model without using formulas, it is simple to check
that the results for models 3 and 4 are identical.
```
oo = np.ones(df.shape[0])
model4 = sm.MixedLM(df.y, oo[:, None], exog_re=None, groups=oo, exog_vc=vcs)
result4 = model4.fit()
print(result4.summary())
```
| github_jupyter |
### Import libraries and modify notebook settings
```
# Import libraries
import os
import sys
import h5py
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from IPython.display import clear_output
from keras.models import Sequential
from keras.layers import Dense, Dropout, Activation, Flatten
from keras.layers import Convolution2D, MaxPooling2D
from keras.layers import BatchNormalization
from keras.utils import np_utils
from keras.utils import HDF5Matrix
from keras.callbacks import Callback
from keras.callbacks import ModelCheckpoint
from keras import regularizers
# Modify notebook settings
%matplotlib inline
```
### Create paths to data folders and files
```
# Create a variable for the project root directory
proj_root = os.path.join(os.pardir)
# Save the path to the folder that contains
# the interim data sets for modeling:
# /data/interim
interim_data_dir = os.path.join(proj_root,
"data",
"interim")
# Save path to the folder for the
# spectrogram arrays
spectrogram_arrays_path = os.path.join(interim_data_dir,
"spectrogram_arrays")
# Full path for test_hdf5_path
test_hdf5_path = os.path.join(spectrogram_arrays_path,
"spectrogram_arrays_test.hdf5")
# Full path for train_hdf5_path
train_hdf5_path = os.path.join(spectrogram_arrays_path,
"spectrogram_arrays_train.hdf5")
# Save the path to the models folder
models_dir = os.path.join(proj_root,
"models")
# Full path for my_model.hdf5
model_path = os.path.join(models_dir,
"my_model.hdf5")
# Save the path to the models/log folder
models_log_dir = os.path.join(models_dir,
"log")
# log_figure file_name
fig_file_name = "log_figure"
# Save the path to the log_figure
log_fig_path = os.path.join(models_log_dir,
fig_file_name)
# log_dataframe.csv file_name
log_file_name = "log_dataframe.csv"
# Save the path to the log_figure
log_file_path = os.path.join(models_log_dir,
log_file_name)
# Save the path to the models/checkpoints folder
models_checkpoints_dir = os.path.join(models_dir,
"checkpoints")
# add the 'src' directory as one where we can import modules
src_dir = os.path.join(proj_root, "src")
sys.path.append(src_dir)
```
## Create objects for X_train, y_train, X_test, & y_test
```
X_train = HDF5Matrix(train_hdf5_path,
'spectrogram_arrays_X_train')
y_train = HDF5Matrix(train_hdf5_path,
'spectrogram_arrays_y_train')
X_test = HDF5Matrix(test_hdf5_path,
'spectrogram_arrays_X_test')
y_test = HDF5Matrix(test_hdf5_path,
'spectrogram_arrays_y_test')
# 6. Preprocess class labels
Y_train = np_utils.to_categorical(y_train)
Y_test = np_utils.to_categorical(y_test)
print(X_train.shape)
print(X_test.shape)
print(y_train.shape)
print(y_test.shape)
print(Y_train.shape)
print(Y_test.shape)
```
# ...
```
with h5py.File(train_hdf5_path, "r") as f:
dset = f['spectrogram_arrays_X_train']
min_val = np.min(dset[0,:,:,:])
max_val = np.max(dset[0,:,:,:])
for i in range(dset.len()):
min_val_new = np.min(dset[i,:,:,:])
min_val_old = min_val
min_val = np.minimum(min_val_new, min_val_old)
max_val_new = np.max(dset[i,:,:,:])
max_val_old = max_val
max_val = np.maximum(max_val_new, max_val_old)
min_val
max_val
```
# Model...
```
#log_fig_path = log_fig_path
class User_Defined_Callback(Callback):
def on_train_begin(self, logs={}):
self.i = 1
self.x = []
self.accuracy = []
self.val_accuracy = []
self.losses = []
self.val_losses = []
self.fig = plt.figure()
self.logs = []
self.df_log = pd.DataFrame()
self.df_log.to_csv(log_file_path)
def on_epoch_end(self, epoch, logs={}):
# Update lists
self.logs.append(logs)
self.x.append(self.i)
self.accuracy.append(logs.get('acc'))
self.val_accuracy.append(logs.get('val_acc'))
self.losses.append(logs.get('loss'))
self.val_losses.append(logs.get('val_loss'))
self.i += 1
# Create log dataframe
self.df_log = pd.DataFrame({'epoch_x': self.x,
'accuracy' : self.accuracy,
'val_accuracy' : self.val_accuracy,
'losses' : self.losses,
'val_losses' : self.val_losses,
'logs' : self.logs})
# Reorder dataframe columns
self.df_log = self.df_log[['epoch_x', 'accuracy', 'val_accuracy', 'losses', 'val_losses', 'logs']]
# Save log dataframe to csv
self.df_log.to_csv(log_file_path)
# Create summary plots of Loss vs Epoch and Accuracy vs Epoch
clear_output(wait=True)
fig, (ax1, ax2) = plt.subplots(ncols=2, figsize=(8,4))
ax1.plot(self.x, self.losses, label="loss")
ax1.plot(self.x, self.val_losses, label="val_loss")
ax1.axes.set_xlabel('Epoch')
ax1.axes.set_ylabel('Loss Function')
ax1.axes.set_title('Loss vs Epoch')
ax1.legend()
ax2.plot(self.x, self.accuracy, label="acc")
ax2.plot(self.x, self.val_accuracy, label="val_acc")
ax2.axes.set_xlabel('Epoch')
ax2.axes.set_ylabel('Accuracy')
ax2.axes.set_title('Accuracy vs Epoch')
ax2.legend()
plt.tight_layout()
plt.savefig(log_fig_path, dpi=300)
plt.show()
user_defined_callback = User_Defined_Callback()
# Checkpoint file names
checkpoint_file_name="weights-improvement-{epoch:02d}-{val_acc:.2f}.hdf5"
# Save the path to the models/checkpoints folder
models_checkpoints_path = os.path.join(models_checkpoints_dir,
checkpoint_file_name)
checkpoint = ModelCheckpoint(models_checkpoints_path,
monitor='val_acc', verbose=0,
save_best_only=True, mode='max')
# Define the callbacks_list
callbacks_list = [checkpoint, user_defined_callback]
# For reproducibility
np.random.seed(42)
```
# Add BatchNormalization() layers
```
def minmax_norm(x, min_val, max_val):
numerator = np.add(x, (-min_val))
divisor = max_val - min_val
return np.divide(numerator, divisor)
# return (x - min_val) / (max_val - min_val)
from keras.layers import Lambda
# Define model architecture
model = Sequential()
# Input Layer
model.add(Activation(None, input_shape=(96, 173, 1)))
#Min_max_scale
model.add(Lambda(minmax_norm, arguments={"min_val": min_val, "max_val": max_val}))
model.add(BatchNormalization())
# Input Layer
#model.add(Activation(None, input_shape=(96, 173, 1)))
#model.add(BatchNormalization())
# Convolution Layer 1
model.add(Convolution2D(24, (5, 5), activation='relu',
input_shape=(96, 173, 1)))
model.add(MaxPooling2D(pool_size=(4,2)))
model.add(BatchNormalization())
# Convolution Layer 2
model.add(Convolution2D(48, (5, 5), activation='relu'))
model.add(MaxPooling2D(pool_size=(4,2)))
model.add(BatchNormalization())
# Convolution Layer 3
model.add(Convolution2D(48, (5, 5),
padding='same',
activation='relu'))
model.add(BatchNormalization())
# Dense Layer
model.add(Flatten())
model.add(Dense(64, activation='relu',
kernel_regularizer=regularizers.l2(0.001),
activity_regularizer=regularizers.l2(0.001)))
# Softmax Layer
#model.add(Dropout(0.5))
model.add(Dense(10, activation='softmax',
kernel_regularizer=regularizers.l2(0.001),
activity_regularizer=regularizers.l2(0.001)))
# Compile model
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
# Fit model on training data
model.fit(X_train, Y_train,
batch_size=100,
epochs=50,
verbose=1,
callbacks=callbacks_list,
validation_data=(X_test, Y_test),
shuffle="batch")
# Evaluate model on test data
score = model.evaluate(X_test, Y_test, verbose=0)
#...
print('test score:', score[1])
model.save(model_path) # creates a HDF5 file 'my_model.hdf5'
```
from keras.layers import Convolution1D
# Define model architecture
model = Sequential()
# Input Layer
model.add(Activation(None, input_shape=(96, 173, 1)))
#Min_max_scale
model.add(Lambda(minmax_norm, arguments={"min_val": min_val, "max_val": max_val}))
model.add(BatchNormalization())
# Input Layer
#model.add(Activation(None, input_shape=(96, 173, 1)))
#model.add(BatchNormalization())
# Convolution Layer 1
model.add(Convolution1D(24, 173, activation='relu',
input_shape=(96, 173, 1)))
#model.add(MaxPooling2D(pool_size=(4,2)))
model.add(BatchNormalization())
# # Convolution Layer 2
# model.add(Convolution2D(48, (5, 5), activation='relu'))
# model.add(MaxPooling2D(pool_size=(4,2)))
# model.add(BatchNormalization())
# # Convolution Layer 3
# model.add(Convolution2D(48, (5, 5),
# padding='same',
# activation='relu'))
# model.add(BatchNormalization())
# Dense Layer
model.add(Flatten())
model.add(Dense(64, activation='relu',
kernel_regularizer=regularizers.l2(0.001),
activity_regularizer=regularizers.l2(0.001)))
# Softmax Layer
#model.add(Dropout(0.5))
model.add(Dense(10, activation='softmax',
kernel_regularizer=regularizers.l2(0.001),
activity_regularizer=regularizers.l2(0.001)))
# Compile model
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
# Fit model on training data
model.fit(X_train, Y_train,
batch_size=100,
epochs=50,
verbose=1,
callbacks=callbacks_list,
validation_data=(X_test, Y_test),
shuffle="batch")
# Evaluate model on test data
score = model.evaluate(X_test, Y_test, verbose=0)
#...
print('test score:', score[1])
| github_jupyter |
```
import pandas as pd
medicare = pd.read_csv("/netapp2/home/se197/RPDR/Josh Lin/3_EHR_V2/CMS/Data/final_medicare.csv")
medicare = medicare[(medicare.Co_CAD_R0 == 1) | (medicare.Co_Diabetes_R0 == 1) | (medicare.Co_CAD_R0 == 1) |
(medicare.Co_Embolism_R0 == 1) | (medicare.Co_DVT_R0 == 1) | (medicare.Co_PE_R0 == 1) |
(medicare.Co_AFib_R0 == 1) | (medicare.Co_HF_R0 == 1) | (medicare.Co_HemoStroke_R0 == 1) |
(medicare.Co_IscheStroke_R0 == 1) | (medicare.Co_OthStroke_R0 == 1) |(medicare.Co_TIA_R0 == 1)
| (medicare.Co_OldMI_R0 == 1) | (medicare.Co_AcuteMI_R0 == 1) | (medicare.Co_PVD_R0 == 1)]
medicare.shape
train_set = medicare[medicare.Hospital != 'BWH'] # MGH; n = 204014
validation_set = medicare[medicare.Hospital == 'BWH'] # BWH and Neither; n = 115726
import numpy as np
fifty_perc_EHR_cont = np.percentile(medicare['Cal_MPEC_R0'],50)
train_set_high = train_set[train_set.Cal_MPEC_R0 >= fifty_perc_EHR_cont]
train_set_low= train_set[train_set.Cal_MPEC_R0 < fifty_perc_EHR_cont]
validation_set_high = validation_set[validation_set.Cal_MPEC_R0 >= fifty_perc_EHR_cont]
validation_set_low = validation_set[validation_set.Cal_MPEC_R0 < fifty_perc_EHR_cont]
predictor_variable = [
'Co_CAD_R0', 'Co_Embolism_R0', 'Co_DVT_R0', 'Co_PE_R0', 'Co_AFib_R0',
'Co_Hypertension_R0', 'Co_Hyperlipidemia_R0', 'Co_Atherosclerosis_R0',
'Co_HF_R0', 'Co_HemoStroke_R0', 'Co_IscheStroke_R0', 'Co_OthStroke_R0',
'Co_TIA_R0', 'Co_COPD_R0', 'Co_Asthma_R0', 'Co_Pneumonia_R0', 'Co_Alcoholabuse_R0',
'Co_Drugabuse_R0', 'Co_Epilepsy_R0', 'Co_Cancer_R0', 'Co_MorbidObesity_R0',
'Co_Dementia_R0', 'Co_Depression_R0', 'Co_Bipolar_R0', 'Co_Psychosis_R0',
'Co_Personalitydisorder_R0', 'Co_Adjustmentdisorder_R0', 'Co_Anxiety_R0',
'Co_Generalizedanxiety_R0', 'Co_OldMI_R0', 'Co_AcuteMI_R0', 'Co_PUD_R0',
'Co_UpperGIbleed_R0', 'Co_LowerGIbleed_R0', 'Co_Urogenitalbleed_R0',
'Co_Othbleed_R0', 'Co_PVD_R0', 'Co_LiverDisease_R0', 'Co_MRI_R0',
'Co_ESRD_R0', 'Co_Obesity_R0', 'Co_Sepsis_R0', 'Co_Osteoarthritis_R0',
'Co_RA_R0', 'Co_NeuroPain_R0', 'Co_NeckPain_R0', 'Co_OthArthritis_R0',
'Co_Osteoporosis_R0', 'Co_Fibromyalgia_R0', 'Co_Migraine_R0', 'Co_Headache_R0',
'Co_OthPain_R0', 'Co_GeneralizedPain_R0', 'Co_PainDisorder_R0',
'Co_Falls_R0', 'Co_CoagulationDisorder_R0', 'Co_WhiteBloodCell_R0', 'Co_Parkinson_R0',
'Co_Anemia_R0', 'Co_UrinaryIncontinence_R0', 'Co_DecubitusUlcer_R0',
'Co_Oxygen_R0', 'Co_Mammography_R0', 'Co_PapTest_R0', 'Co_PSATest_R0',
'Co_Colonoscopy_R0', 'Co_FecalOccultTest_R0', 'Co_FluShot_R0', 'Co_PneumococcalVaccine_R0', 'Co_RenalDysfunction_R0', 'Co_Valvular_R0', 'Co_Hosp_Prior30Days_R0',
'Co_RX_Antibiotic_R0', 'Co_RX_Corticosteroid_R0', 'Co_RX_Aspirin_R0', 'Co_RX_Dipyridamole_R0',
'Co_RX_Clopidogrel_R0', 'Co_RX_Prasugrel_R0', 'Co_RX_Cilostazol_R0', 'Co_RX_Ticlopidine_R0',
'Co_RX_Ticagrelor_R0', 'Co_RX_OthAntiplatelet_R0', 'Co_RX_NSAIDs_R0',
'Co_RX_Opioid_R0', 'Co_RX_Antidepressant_R0', 'Co_RX_AAntipsychotic_R0', 'Co_RX_TAntipsychotic_R0',
'Co_RX_Anticonvulsant_R0', 'Co_RX_PPI_R0', 'Co_RX_H2Receptor_R0', 'Co_RX_OthGastro_R0',
'Co_RX_ACE_R0', 'Co_RX_ARB_R0', 'Co_RX_BBlocker_R0', 'Co_RX_CCB_R0', 'Co_RX_Thiazide_R0',
'Co_RX_Loop_R0', 'Co_RX_Potassium_R0', 'Co_RX_Nitrates_R0', 'Co_RX_Aliskiren_R0',
'Co_RX_OthAntihypertensive_R0', 'Co_RX_Antiarrhythmic_R0', 'Co_RX_OthAnticoagulant_R0',
'Co_RX_Insulin_R0', 'Co_RX_Noninsulin_R0', 'Co_RX_Digoxin_R0', 'Co_RX_Statin_R0',
'Co_RX_Lipid_R0', 'Co_RX_Lithium_R0', 'Co_RX_Benzo_R0', 'Co_RX_ZDrugs_R0',
'Co_RX_OthAnxiolytic_R0', 'Co_RX_Dementia_R0', 'Co_RX_Hormone_R0',
'Co_RX_Osteoporosis_R0', 'Co_N_Drugs_R0', 'Co_N_Hosp_R0', 'Co_Total_HospLOS_R0',
'Co_N_MDVisit_R0', 'Co_RX_AnyAspirin_R0', 'Co_RX_AspirinMono_R0', 'Co_RX_ClopidogrelMono_R0',
'Co_RX_AspirinClopidogrel_R0', 'Co_RX_DM_R0', 'Co_RX_Antipsychotic_R0'
]
co_train_gpop = train_set[predictor_variable]
co_train_high = train_set_high[predictor_variable]
co_train_low = train_set_low[predictor_variable]
co_validation_gpop = validation_set[predictor_variable]
co_validation_high = validation_set_high[predictor_variable]
co_validation_low = validation_set_low[predictor_variable]
out_train_death_gpop = train_set['ehr_claims_death']
out_train_death_high = train_set_high['ehr_claims_death']
out_train_death_low = train_set_low['ehr_claims_death']
out_validation_death_gpop = validation_set['ehr_claims_death']
out_validation_death_high = validation_set_high['ehr_claims_death']
out_validation_death_low = validation_set_low['ehr_claims_death']
```
# Template LR
```
def lr(X_train, y_train):
from sklearn.linear_model import Lasso
from sklearn.decomposition import PCA
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import GridSearchCV
from imblearn.over_sampling import SMOTE
from sklearn.preprocessing import StandardScaler
model = LogisticRegression()
param_grid = [
{'C' : np.logspace(-4, 4, 20)}
]
clf = GridSearchCV(model, param_grid, cv = 5, verbose = True, n_jobs = -1)
best_clf = clf.fit(X_train, y_train)
return best_clf
def scores(X_train,y_train):
from sklearn.metrics import accuracy_score
from sklearn.metrics import f1_score
from sklearn.metrics import fbeta_score
from sklearn.metrics import roc_auc_score
from sklearn.metrics import log_loss
pred = best_clf.predict(X_train)
actual = y_train
print(accuracy_score(actual,pred))
print(f1_score(actual,pred))
print(fbeta_score(actual,pred, average = 'macro', beta = 2))
print(roc_auc_score(actual, best_clf.predict_proba(X_train)[:,1]))
print(log_loss(actual,best_clf.predict_proba(X_train)[:,1]))
def cross_val(X,y):
from sklearn.model_selection import KFold
from sklearn.model_selection import cross_validate
from sklearn.metrics import log_loss
from sklearn.metrics import roc_auc_score
from sklearn.metrics import fbeta_score
import sklearn
import numpy as np
cv = KFold(n_splits=5, random_state=1, shuffle=True)
log_loss = []
auc = []
accuracy = []
f1 = []
f2 = []
for train_index, test_index in cv.split(X):
X_train, X_test, y_train, y_test = X.iloc[train_index], X.iloc[test_index], y.iloc[train_index], y.iloc[test_index]
model = lr(X_train, y_train)
prob = model.predict_proba(X_test)[:,1] # prob is a vector of probabilities
print(prob)
pred = np.round(prob) # pred is the rounded predictions
log_loss.append(sklearn.metrics.log_loss(y_test, prob))
auc.append(sklearn.metrics.roc_auc_score(y_test, prob))
accuracy.append(sklearn.metrics.accuracy_score(y_test, pred))
f1.append(sklearn.metrics.f1_score(y_test, pred, average = 'macro'))
f2.append(fbeta_score(y_test,pred, average = 'macro', beta = 2))
print(np.mean(accuracy))
print(np.mean(f1))
print(np.mean(f2))
print(np.mean(auc))
print(np.mean(log_loss))
```
# FAMD Transformation
```
from prince import FAMD
famd = FAMD(n_components = 15, n_iter = 3, random_state = 101)
for (colName, colData) in co_train_gpop.iteritems():
if (colName != 'Co_N_Drugs_R0' and colName!= 'Co_N_Hosp_R0' and colName != 'Co_Total_HospLOS_R0' and colName != 'Co_N_MDVisit_R0'):
co_train_gpop[colName].replace((1,0) ,('yes','no'), inplace = True)
co_train_low[colName].replace((1,0) ,('yes','no'), inplace = True)
co_train_high[colName].replace((1,0) ,('yes','no'), inplace = True)
co_validation_gpop[colName].replace((1,0), ('yes','no'), inplace = True)
co_validation_high[colName].replace((1,0), ('yes','no'), inplace = True)
co_validation_low[colName].replace((1,0), ('yes','no'), inplace = True)
famd.fit(co_train_gpop)
co_train_gpop_FAMD = famd.transform(co_train_gpop)
famd.fit(co_train_high)
co_train_high_FAMD = famd.transform(co_train_high)
famd.fit(co_train_low)
co_train_low_FAMD = famd.transform(co_train_low)
famd.fit(co_validation_gpop)
co_validation_gpop_FAMD = famd.transform(co_validation_gpop)
famd.fit(co_validation_high)
co_validation_high_FAMD = famd.transform(co_validation_high)
famd.fit(co_validation_low)
co_validation_low_FAMD = famd.transform(co_validation_low)
```
# General Population
```
best_clf = lr(co_train_gpop_FAMD, out_train_death_gpop)
cross_val(co_train_gpop_FAMD, out_train_death_gpop)
print()
scores(co_validation_gpop_FAMD, out_validation_death_gpop)
comb = []
for i in range(len(predictor_variable)):
comb.append(predictor_variable[i] + str(best_clf.best_estimator_.coef_[:,i:i+1]))
comb
```
# High Continuity
```
best_clf = lr(co_train_high_FAMD, out_train_death_high)
cross_val(co_train_high_FAMD, out_train_death_high)
print()
scores(co_validation_high_FAMD, out_validation_death_high)
comb = []
for i in range(len(predictor_variable)):
comb.append(predictor_variable[i] + str(best_clf.best_estimator_.coef_[:,i:i+1]))
comb
```
# Low Continuity
```
best_clf = lr(co_train_low_FAMD, out_train_death_low)
cross_val(co_train_low_FAMD, out_train_death_low)
print()
scores(co_validation_low_FAMD, out_validation_death_low)
comb = []
for i in range(len(predictor_variable)):
comb.append(predictor_variable[i] + str(best_clf.best_estimator_.coef_[:,i:i+1]))
comb
```
| github_jupyter |
# Moment Based Model
This model is better estimating parameters in biological systems mainly affected by intrinsic noise. It computes the second order moment differential equations of the system to approximate some population statistics as mean and variance of the gene expression. In this way, it is possible to estimate deterministically a system which was simulated in a stochastic way. This model is applied together to a KLD minimization as a loss function.
```
#required libraries
import simsysbio as s2b
import numpy as np
import matplotlib.pyplot as plt
from scipy.optimize import minimize
```
**Loads data**. The data is loaded to save time computing a new whole population.
```
uObs = np.load("CMEmean.npy")
sdObs = np.load("CMEsd.npy")
tog = np.load("tog.npy")
hog = np.load("hog.npy")
```
**Plots data used to perform estimation**
```
plt.figure()
plt.plot(tog, uObs)
plt.plot(tog, uObs+sdObs, 'b')
plt.plot(tog, uObs-sdObs, 'b')
plt.xlabel('Time (min)')
plt.ylabel('Concentration')
plt.grid()
plt.show()
#molecular species
especies = ['mRNA', 'Protein']
#Stoichiometric matrices
reactivos = np.array([[0, 1, 1, 0],[0, 0, 0, 1]])
productos = np.array([[1, 0, 1, 0],[0, 0, 1, 0]])
#kinetic parameters names
parametros = ['c1', 'c2', 'c3', 'c4']
#system input
entrada = 'u'
idxR = 0
#computes simbolic system of differential equations
ecuaciones, variables = s2b.simbMoments(especies, reactivos, productos, parametros,
inputN=entrada)
#creates .py file containing the system of equation upon second moment order
s2b.model2MDefiner(variables["nameVar"][1:], ecuaciones, variables["pars"])
#kinetic parameters
parsValues = [4.0, 0.010, 1.0, 0.006]
#initial concentrations
sp0 = np.zeros(len(variables["nameVar"]) - 1)
#initial noise measurement guess
noise = np.array([0,0])
#regressor variables
regressor = {
"ODEs": ecuaciones,
"matrizR": reactivos,
"matrizP": productos,
"vPars": parsValues,
"idxR": idxR,
"inpU": hog,
"Vtime": tog,
"species0":sp0,
"noise":noise,
"uObs":uObs,
"sdObs":sdObs
}
regressor.update(variables)
```
**Inferring process**
```
#initial parameter guesses
beta0 = np.array([3.0, 0.02, 0.9, 0.01, 10.0, 0.1])
```
#finds moments
ODE2M, vars2M = s2b.simbMoments(especies, reactivos, productos, parametros,
inputN=entrada, indexU=idxR)
sp0 = np.concatenate((regressor["species0"], np.zeros(len(ODE2M) - len(regressor["species"]))))
regressor2 = {
"ODEs": ODE2M,
"regressor":regressor,
"inpU": hog,
"Vtime": tog,
"species0":sp0,
"meanCell":uObs,
"sdCell":sdObs
}
regressor2.update(vars2M)
idx2M = list(map(str, regressor2["species"]))
idx2M = idx2M.index(especies[-1] + '**2')
regressor2["idx2M"] = idx2M
regressor2["errPars"] = errAB
```
#defines estimation function
def KLDmeasure(uObs, sdObs, uM, sdM):
kld = np.log(sdM/sdObs) + ((sdObs**2 + (uObs-uM)**2)/(2*sdM**2)) - 1/2
return np.mean(kld)
def KLDmomentsWrap(Allpars, regressor):
uM, sdM = s2b.solve2M(Allpars[:-2], Allpars[-2:], regressor)
mcKLD = KLDmeasure(regressor["uObs"], regressor["sdObs"], uM, sdM)
print(mcKLD)
return mcKLD
#performs estimation
options = {"maxiter":150} #100 iterations for better estimation
minimum = minimize(KLDmomentsWrap, beta0, args=(regressor,), options=options)
betacal = minimum.x
print("Infered kinetic parameters", betacal[:-2])
print("Infered noise parameters", betacal[-2:])
#simulates output with infered parameters
uMif, sdMif = s2b.solve2M(betacal[:-2], betacal[-2:], regressor)
#plot population curves
plt.figure()
plt.plot(tog, uObs, label="Observed")
plt.plot(tog, uObs+sdObs, 'b')
plt.plot(tog, uObs-sdObs, 'b')
plt.plot(tog, uMif, label="Infered")
plt.plot(tog, uMif+sdMif, 'y')
plt.plot(tog, uMif-sdMif, 'y')
plt.xlabel('Time (min)')
plt.ylabel('Concentration')
plt.legend(loc="best")
plt.grid()
plt.show()
```
| github_jupyter |
# ism3d.uvhelper: visibility imaging
## Setup
We first import essential API functions / modules from `ism3d` and other libraries
**Used ISM3D Functions:**
* `im3d.logger.logger_config`
* `im3d.logger.logger_status`
```
nb_dir=_dh[0]
os.chdir(nb_dir+'/../output/mockup')
sys.path.append(nb_dir)
from notebook_setup import *
%matplotlib inline
%config InlineBackend.figure_format = "png" # ‘png’, ‘retina’, ‘jpeg’, ‘svg’, ‘pdf’.
%reload_ext wurlitzer
%reload_ext memory_profiler
%reload_ext line_profiler
ism3d.logger_config(logfile='ism3d.log',loglevel='INFO',logfilelevel='INFO',log2term=False)
print(''+ism3d.__version__)
print('working dir: {}\n'.format(os.getcwd()))
```
## Data Import
We import the visibility data from CASA measurement sets into the internal `uvdata` variable (essential a simple nested dict, not class yet) and also save them into compressed HDF5 for compact storage and easy retrieve.
Here three ALMA/VLA datasets are used:
* mockup1: based on the VLA GN20 observation 1-channel
* mockup2: based on the ALMA G09 observation 1-channel
* mockup3: based on the ALMA G09 observation 240-channel, only 2mins integration
* mockup3: based on the ALMA G09 observation 240-channel, all on-source data
**Used ISM3D Functions:**
* `im3d.uvhelper.io.to_hdf5`
* `ism3d.uvhelper.ms.rmPointing`
* `ism3d.uvhelper.ms.read_ms`
```
os.system('rm -rf '+'mockup1_basis.ms')
mstransform(vis='../data/gn20/vla/AC974.100409.ms',outputvis='mockup1_basis.ms',
spw='0:60',datacolumn='data')
os.system('rm -rf '+'mockup2_basis.ms')
mstransform(vis='../data/g09/alma/bb4.ms',outputvis='mockup2_basis.ms',
spw='*:60',datacolumn='data')
rmPointing('mockup2_basis.ms',verbose=False)
os.system('rm -rf '+'mockup3_basis.ms')
mstransform(vis='../../data/g09/alma/bb4.ms',outputvis='mockup3_basis.ms',
spw='',timerange='06:08:00~06:10:00',datacolumn='data')
rmPointing('mockup3_basis.ms',verbose=False)
os.system('rm -rf '+'mockup4_basis.ms')
os.system('ln -s ../../data/g09/alma/bb4.ms '+'mockup4_basis.ms')
for model_name in ['mockup1','mockup2','mockup3','mockup4']:
# option 1
#uvdata={}
#read_ms(vis=model_name+'_basis.ms',dataset=uvdata,keyrule='basename')
# option 2
uvdata=read_ms(vis=model_name+'_basis.ms')
# save to / reterive from .h5
to_hdf5(uvdata,outname=model_name+'_basis.h5',checkname=False,compression='gzip')
```
## Imaging
We image the visibility using two different approched implemented in `ism3d`:
* `ism3d.uvhelper.invert`: a function wrapping around casa.tclean etc. to create dirty maps in an organized fahsion
* `ism3d.uvhelper.invert_ft`: the same purpose as above, but based on FINUFFT
**Used ISM3D Functions:**
* `ism3d.uvhelper.imager.invert`
* `ism3d.uvhelper.io.from_hdf5`
* `ism3d.uvhelper.invert`
* `ism3d.uvhelper.invert_ft`
* `ism3d.uvhelper.make_psf`
* `ism3d.uvhelper.ft.advise_header`
* `ism3d.xyhelper.cube.hextract`
```
model_name='mockup4'
uvdata=from_hdf5(model_name+'_basis.h5')
header=advise_header(uvdata['uvw'],
uvdata['phasecenter'],
uvdata['chanfreq'],
uvdata['chanwidth'],
antsize=12*u.m,sortbyfreq=True)
cell=header['CDELT2']<<u.deg
imsize=header['NAXIS1']
print(imsize,cell.to(u.arcsec))
tic= time.time()
invert(vis=model_name+'_basis.ms',
imagename=model_name+'_basis.images/casa',
weighting='natural',specmode='cubedata',width='',start='',nchan=-1, # width=-1,start=239,nchan=-1,
cell=cell.to_value(u.arcsec),imsize=[imsize,imsize],onlydm=False,dropstokes=True)
toc= time.time()
print("Elapsed Time: {:>8.2f} seconds # {} \n".format(toc-tic,'ism3d.uvhelper.imager.invert'))
tic= time.time()
%memit cube=invert_ft(uvdata=uvdata,header=header,sortbyfreq=True).astype(np.float32)
%memit psf=(make_psf(uvdata=uvdata,header=header,sortbyfreq=True)).astype(np.float32)
toc= time.time()
print("Elapsed Time: {:>8.2f} seconds # {} \n".format(toc-tic,'ism3d.uvhelper.ft.invert_ft/.make_psf'))
fits.writeto(model_name+'_basis.images/nufft.image.fits',cube.T,header,overwrite=True)
fits.writeto(model_name+'_basis.images/nufft.psf.fits',psf.T,header,overwrite=True)
for version in ['image','psf']:
tcube,thdr=fits.getdata(model_name+'_basis.images/casa.'+version+'.fits',header=True)
cube,hdr=fits.getdata(model_name+'_basis.images/nufft.'+version+'.fits',header=True)
cube_diff=cube-tcube
fits.writeto(model_name+'_basis.images/diff.'+version+'.fits',cube_diff,thdr,overwrite=True)
if model_name=='mockup4' or model_name=='mockup3':
# get ride of the first plan of mockup4 as it's partially flagged with different PSF.
for version in ['nufft.image','nufft.psf','casa.image','casa.psf','casa.pb','diff.image','diff.psf']:
data,header=fits.getdata(model_name+'_basis.images/'+version+'.fits',header=True)
data_sub,header_sub=hextract(data, header, np.s_[1:,:,:])
fits.writeto(model_name+'_basis.images/'+version+'.fits',data_sub,header_sub,overwrite=True)
```
## Visualize
Here we demostrate the visulization capabiliy of ism3d. Specifically, we plot the results from two imaging appraoching and compare their precisions.
**Used ISM3D Functions:**
* `ism3d.visualize.nb.make_gif`
* `ism3d.visualize.nb.show_gif`
* `ism3d.visualize.plts.im_grid`
```
units=[] ; images=[] ; titles=[]; vmaxs=[]; vmins=[]
for version in ['casa.image','casa.psf','casa.pb','nufft.image','nufft.psf',None,'diff.image','diff.psf']:
if version is not None:
data,hdr=fits.getdata(model_name+'_basis.images/'+version+'.fits',header=True)
titles.append(version)
if 'psf' in titles[-1]:
images.append(data); units.append("Jy/beam")
else:
images.append(data*1e3); units.append("mJy/beam")
vmaxs.append(np.nanmax(images[-1]))
vmins.append(np.nanmin(images[-1]))
else:
titles.append(None); images.append(None); units.append(None); vmaxs.append(None); vmins.append(None)
w = WCS(hdr).celestial
coord = SkyCoord(hdr['CRVAL1'], hdr['CRVAL2'], unit="deg")
offset_w=linear_offset_coords(w,coord)
nchan=hdr['NAXIS3']
stepchan= int(np.maximum(np.floor(int(nchan/5)),1))
fignames=[]
for ichan in range(0,nchan,stepchan):
#clear_output(wait=True)
figname=model_name+'_basis.images/chmap/ch{:03d}'.format(ichan)+'.pdf'
images0=[None if image is None else image[ichan,:,:] for image in images]
titles0=[None if title is None else title+'['+'{}'.format(ichan)+']' for title in titles ]
im_grid(images0,offset_w,units=units,titles=titles0,nxy=(3,3),figsize=(9,9),figname=figname,vmins=vmins,vmaxs=vmaxs) ;
fignames.append(figname)
make_gif(fignames,model_name+'_basis.images/chmap.gif')
show_gif(model_name+'_basis.images/chmap.gif')
```
| github_jupyter |
<h1 style="font-size:42px; text-align:center; margin-bottom:30px;"><span style="color:SteelBlue">Lesson 1:</span> Exploratory Analysis</h1>
<hr>
Welcome to the workbook for <span style="color:royalblue">Lesson 1: Exploratory Analysis</span>!
Our goal with this step is to "get to know" the data.
* Think of it like a "first date."
* Try to learn as much about the data as possible, but don't invest too much time and get stuck on this step.
* You'll probably need to do ad-hoc data exploration later anyway, so you don't need to be 100% comprehensive right now.
<br><hr id="toc">
### In this lesson...
In this lesson, we'll go through the essential exploratory analysis steps:
1. [Basic information](#basic)
2. [Distributions of numeric features](#numeric)
3. [Distributions of categorical features](#categorical)
4. [Segmentations](#segmentations)
5. [Correlations](#correlations)
Along the way, we'll note the practical implications of our findings.
<br>
**(Reminder) To run a code cell...**
1. Click anywhere in the cell to select it.
2. Press <code style="color:steelblue">Shift + Enter</code> to run the code.
3. You can insert new code cells using the **Insert** dropdown in the toolbar at the top.
<br><hr>
### First, let's import libraries and load the dataset.
In general, it's good practice to keep all of your library imports at the top of your notebook or program.
Let's import the libraries we'll need for this lesson.
```
# NumPy for numerical computing
import numpy as np
# Pandas for DataFrames
import pandas as pd
pd.set_option('display.max_columns', 100)
# Matplotlib for visualization
from matplotlib import pyplot as plt
# display plots in the notebook
%matplotlib inline
# Seaborn for easier visualization
import seaborn as sns
```
Next, let's import the dataset.
* Pandas has a <code style="color:steelblue">pd.read_csv()</code> function for importing CSV files into a Pandas DataFrame.
* You can name the DataFrame variable anything, but we prefer the simple name: <code style="color:steelblue">df</code> (short for DataFrame).
```
# Load real estate data from CSV
df = pd.read_csv('project_files/real_estate_data.csv')
```
<br id="basic">
# 1. Basic information
First, always look at basic information about the dataset.
#### Display the dimensions of the dataset.
```
# Dataframe dimensions
df.shape
```
#### Next, display the data types of our features.
```
# Column datatypes
df.dtypes
```
#### Display the first 5 rows to see example observations.
```
# Display first 5 rows of df
df[0:5]
```
<br><hr style="border-color:royalblue;background-color:royalblue;height:1px;">
## <span style="color:RoyalBlue">Exercise 1.1</span>
Before moving on, let's dig a bit deeper into some of these functionalities. Getting some extra practice right now will set you up for smoother success as you continue through the project.
<br>
**First, try to filter <code style="color:steelblue">df.dtypes</code> to only categorical variables.**
* **Tip:** Remember our boolean masks from the Python Crash Course?
* **Tip:** A Series can be filtered by boolean masks the same way that a DataFrame can be.
```
# Filter and display only df.dtypes that are 'object'
df.dtypes[df.dtypes == 'object']
```
#### Iterate through the categorical feature names and print each name.
```
# Loop through categorical feature names and print each one
for features in df.dtypes[df.dtypes == 'object'].index:
print(features)
```
As you'll see later, the ability to select feature names based on some condition (instead of manually typing out each one) will be quite useful.
<br>
**Next, look at a few more examples by displaying the first 10 rows of data, instead of just the first 5.**
* **Tip:** <code style="color:steelblue">df.head()</code> returns the first 5 rows by default, but you can pass in an integer argument for the number of rows you want.
```
# Display the first 10 rows of data
df.head(10)
```
Finally, it's also helpful to look at the last 5 rows of data.
* Sometimes datasets will have **corrupted data** hiding at the very end (depending on the data source).
* It never hurts to double-check.
<br>
**Display the last 5 rows of data.**
* **Hint:** You've seen the "head," but what about the "tail?"
```
# Display last 5 rows of data
df.tail()
```
<hr style="border-color:royalblue;background-color:royalblue;height:1px;">
<div style="text-align:center; margin: 40px 0 40px 0; font-weight:bold">
[Back to Contents](#toc)
</div>
<br id="numeric">
# 2. Distributions of numeric features
One of the most enlightening data exploration tasks is plotting the distributions of your features.
<br><hr style="border-color:royalblue;background-color:royalblue;height:1px;">
## <span style="color:RoyalBlue">Exercise 1.2</span>
**Plot the histogram grid, but make it larger, and rotate the x-axis labels clockwise by 45 degrees.**
* <code style="color:steelblue">df.hist()</code> has a <code style="color:steelblue">figsize=</code> argument takes a tuple for figure size.
* Try making the figure size 14 x 14
* <code style="color:steelblue">df.hist()</code> has a <code style="color:steelblue">xrot=</code> argument rotates x-axis labels **counter-clockwise**.
* The [documentation](http://pandas.pydata.org/pandas-docs/version/0.17.0/generated/pandas.DataFrame.hist.html) is useful for learning more about the arguments to the <code style="color:steelblue">.hist()</code> function.
* **Tip:** It's ok to arrive at the answer through **trial and error** (this is often easier than memorizing the various arguments).
```
# Plot histogram grid
df.hist(xrot=-45, figsize=(14, 14))
# Clear the text "residue"
plt.show()
```
#### Display summary statistics for the numerical features.
```
# Summarize numerical features
df.describe()
```
<hr style="border-color:royalblue;background-color:royalblue;height:1px;">
<div style="text-align:center; margin: 40px 0 40px 0; font-weight:bold">
[Back to Contents](#toc)
</div>
<br id="categorical">
# 3. Distributions of categorical features
Next, let's take a look at the distributions of our categorical features.
<br>
Display summary statistics for categorical features.
```
# Summarize categorical features
df.describe(include=["object"])
```
Plot bar plot for the <code style="color:steelblue">'exterior_walls'</code> feature.
```
# Bar plot for 'exterior_walls'
sns.countplot(y="exterior_walls", data=df)
```
<br><hr style="border-color:royalblue;background-color:royalblue;height:1px;">
## <span style="color:RoyalBlue">Exercise 1.3</span>
**Write a <code style="color:steelblue">for</code> loop to plot bar plots of each of the categorical features.**
* Write the loop to be able to handle any number of categorical features (borrow from your answer to <span style="color:royalblue">Exercise 1.1</span>).
* Invoke <code style="color:steelblue">plt.show()</code> after each bar plot to display all 3 plots in one output.
* Which features suffer from sparse classes?
```
# Plot bar plot for each categorical feature
for features in df.dtypes[df.dtypes == "object"].index:
sns.countplot(y=features, data=df)
plt.show()
```
<hr style="border-color:royalblue;background-color:royalblue;height:1px;">
<div style="text-align:center; margin: 40px 0 40px 0; font-weight:bold">
[Back to Contents](#toc)
</div>
<br id="segmentations">
# 4. Segmentations
Next, let's create some segmentations. Segmentations are powerful ways to cut the data to observe the relationship between **categorical features** and **numeric features**.
<br>
Segment <code style="color:steelblue">'tx_price'</code> by <code style="color:steelblue">'property_type'</code> and plot the resulting distributions
```
# Segment tx_price by property_type and plot distributions
sns.boxplot(y="property_type", x="tx_price", data=df)
```
Segment by <code style="color:steelblue">'property_type'</code> and calculate the average value of each feature within each class:
```
# Segment by property_type and display the means within each class
df.groupby("property_type").mean()
```
<br><hr style="border-color:royalblue;background-color:royalblue;height:1px;">
## <span style="color:RoyalBlue">Exercise 1.4</span>
On average, it looks like single family homes are more expensive.
How else do the different property types differ? Let's see:
<br>
**First, segment <code style="color:steelblue">'sqft'</code> by <code style="color:steelblue">'property_type'</code> and plot the boxplots.**
```
# Segment sqft by sqft and property_type distributions
sns.boxplot(y="property_type", x="sqft", data=df)
```
<br>
**After producing the plot, consider these questions:**
* Which type of property is larger, on average?
* Which type of property sees greater variance in sizes?
* Does the difference in distributions between classes make intuitive sense?
<br>
**Next, display the standard deviations of each feature alongside their means after performing a groupby.**
* This will give you a better idea of the variation within in feature, by class.
* **Tip:** Pass a list of metrics into the <code style="color:steelblue">.agg()</code> function, after performing your groupby.
* Check out the [documentation](http://pandas.pydata.org/pandas-docs/stable/groupby.html#applying-multiple-functions-at-once) for more help.
```
# Segment by property_type and display the means and standard deviations within each class
df.groupby("property_type").agg([np.mean, np.std])
```
<hr style="border-color:royalblue;background-color:royalblue;height:1px;">
<div style="text-align:center; margin: 40px 0 40px 0; font-weight:bold">
[Back to Contents](#toc)
</div>
<br id="correlations">
# 5. Correlations
Finally, let's take a look at the relationships between **numeric features** and **other numeric features**.
<br>
Create a <code style="color:steelblue">correlations</code> dataframe from <code style="color:steelblue">df</code>.
```
# Calculate correlations between numeric features
correlations = df.corr()
# Save correlations to csv to view in Orange
correlations.to_csv("./project_files/correlations.csv")
```
#### Visualize the correlation grid with a heatmap to make it easier to digest.
```
# Make the figsize 7 x 6
plt.figure(figsize=(7,6))
# Plot heatmap of correlations
sns.heatmap(correlations)
```
<br><hr style="border-color:royalblue;background-color:royalblue;height:1px;">
## <span style="color:RoyalBlue">Exercise 1.5</span>
When plotting a heatmap of correlations, it's often helpful to do four things:
1. Change the background to white. This way, 0 correlation will show as white
2. Annotate the cell with their correlations values
3. Mask the top triangle (less visual noise)
4. Drop the legend (colorbar on the side)
<br>
**First, change the background to white.**
* Seaborn has several different **themes**. The default theme is called <code style="color:crimson">'darkgrid'</code>.
* You can change the theme with <code style="color:steelblue">sns.set_style()</code>.
* You only need to run this once, and the theme will persist until you change it again.
* Change the theme to <code style="color:crimson">'white'</code>
* Make the figure size 10 x 8
```
# Change color scheme
sns.set_style("white")
# Make the figsize 10 x 8
plt.figure(figsize=(10,8))
# Plot heatmap of correlations
sns.heatmap(correlations)
```
See how the cells for <code style="color:steelblue">'basement'</code> are now white? That's what we want because they were not able to be calculated.
<br>
**Next, display the correlation values in each cell.**
* The <code style="color:steelblue">annot=</code> argument controls whether to annotate each cell with its value. By default, it's <code style="color:crimson">False</code>.
* To make the chart cleaner, multiply the <code style="color:steelblue">correlations</code> DataFrame by 100 before passing it to the heatmap function.
* Pass in the argument <code style="color:steelblue">fmt=<span style="color:crimson">'.0f'</span></code> to format the annotations to a whole number.
```
# Make the figsize 10 x 8
plt.figure(figsize=(10, 8))
# Plot heatmap of annotated correlations
correlations = correlations * 100
sns.heatmap(correlations, annot=True, fmt='.0f')
```
#### Next, we'll generate a mask for the top triangle. Run this code:
```
# Generate a mask for the upper triangle
mask = np.zeros_like(correlations, dtype=np.bool)
mask[np.triu_indices_from(mask)] = True
```
<br>
**Plot the heatmap again, this time using that mask.**
* <code style="color:steelblue">sns.heatmap()</code> has a <code style="color:steelblue">mask=</code> argument.
* Keep all of the other styling changes you've made up to now.
```
# Make the figsize 10 x 8
plt.figure(figsize=(10, 8))
# Plot heatmap of correlations
sns.heatmap(correlations, annot=True, fmt='.0f', mask=mask)
```
<br>
**Finally, remove the colorbar on the side.**
* <code style="color:steelblue">sns.heatmap()</code> has a <code style="color:steelblue">cbar=</code> argument. By default, it's <code style="color:crimson">True</code>.
* Keep all of the other styling changes you've made up to now.
* But change the figure size to 9 x 8 (since we're removing the sidebar, this will help us keep nice proportions)
```
# Make the figsize 9 x 8
plt.figure(figsize=(9, 8))
# Plot heatmap of correlations
sns.heatmap(correlations, annot=True, fmt='.0f', mask=mask, cbar=False)
```
<hr style="border-color:royalblue;background-color:royalblue;height:1px;">
<div style="text-align:center; margin: 40px 0 40px 0; font-weight:bold">
[Back to Contents](#toc)
</div>
<br>
## Next Steps
Congratulations for making it through Exploratory Analysis!
Before powering on to the next lesson, we recommend going back and reviewing the charts you made. This time, since you've already created them, you can move through more quickly and really start to understand the **story** behind the data.
As a reminder, here are a few things you did in this lesson:
* You explored basic information about your dataset.
* You plotted distributions of numeric and categorical features.
* You segmented your dataset by <code style="color:steelblue">'property_type'</code>.
* And you visualized a heatmap of feature correlations.
As you work through the next lesson, <span style="color:royalblue">Lesson 2: Data Cleaning</span>, we recommend keeping this workbook open in a separate tab. It might be helpful to have your charts in front of you as you clean the data.
<div style="text-align:center; margin: 40px 0 40px 0; font-weight:bold">
[Back to Contents](#toc)
</div>
| github_jupyter |
# Relevancy Analysis
<div class="alert alert-info">
This tutorial is available as an IPython notebook at [Malaya/example/relevancy](https://github.com/huseinzol05/Malaya/tree/master/example/relevancy).
</div>
<div class="alert alert-warning">
This module only trained on standard language structure, so it is not save to use it for local language structure.
</div>
```
%%time
import malaya
```
### Models accuracy
We use `sklearn.metrics.classification_report` for accuracy reporting, check at https://malaya.readthedocs.io/en/latest/models-accuracy.html#relevancy-analysis
### labels supported
Default labels for relevancy module.
```
malaya.relevancy.label
```
### Explanation
Positive relevancy: The article or piece of text is relevant, tendency is high to become not a fake news. Can be a positive or negative sentiment.
Negative relevancy: The article or piece of text is not relevant, tendency is high to become a fake news. Can be a positive or negative sentiment.
**Right now relevancy module only support deep learning model**.
```
negative_text = 'Roti Massimo Mengandungi DNA Babi. Roti produk Massimo keluaran Syarikat The Italian Baker mengandungi DNA babi. Para pengguna dinasihatkan supaya tidak memakan produk massimo. Terdapat pelbagai produk roti keluaran syarikat lain yang boleh dimakan dan halal. Mari kita sebarkan berita ini supaya semua rakyat Malaysia sedar dengan apa yang mereka makna setiap hari. Roti tidak halal ada DNA babi jangan makan ok.'
positive_text = 'Jabatan Kemajuan Islam Malaysia memperjelaskan dakwaan sebuah mesej yang dikitar semula, yang mendakwa kononnya kod E dikaitkan dengan kandungan lemak babi sepertimana yang tular di media sosial. . Tular: November 2017 . Tular: Mei 2014 JAKIM ingin memaklumkan kepada masyarakat berhubung maklumat yang telah disebarkan secara meluas khasnya melalui media sosial berhubung kod E yang dikaitkan mempunyai lemak babi. Untuk makluman, KOD E ialah kod untuk bahan tambah (aditif) dan ianya selalu digunakan pada label makanan di negara Kesatuan Eropah. Menurut JAKIM, tidak semua nombor E yang digunakan untuk membuat sesuatu produk makanan berasaskan dari sumber yang haram. Sehubungan itu, sekiranya sesuatu produk merupakan produk tempatan dan mendapat sijil Pengesahan Halal Malaysia, maka ia boleh digunakan tanpa was-was sekalipun mempunyai kod E-kod. Tetapi sekiranya produk tersebut bukan produk tempatan serta tidak mendapat sijil pengesahan halal Malaysia walaupun menggunakan e-kod yang sama, pengguna dinasihatkan agar berhati-hati dalam memilih produk tersebut.'
```
### List available Transformer models
```
malaya.relevancy.available_transformer()
```
### Load Transformer model
```python
def transformer(model: str = 'xlnet', quantized: bool = False, **kwargs):
"""
Load Transformer relevancy model.
Parameters
----------
model : str, optional (default='bert')
Model architecture supported. Allowed values:
* ``'bert'`` - Google BERT BASE parameters.
* ``'tiny-bert'`` - Google BERT TINY parameters.
* ``'albert'`` - Google ALBERT BASE parameters.
* ``'tiny-albert'`` - Google ALBERT TINY parameters.
* ``'xlnet'`` - Google XLNET BASE parameters.
* ``'alxlnet'`` - Malaya ALXLNET BASE parameters.
* ``'bigbird'`` - Google BigBird BASE parameters.
* ``'tiny-bigbird'`` - Malaya BigBird BASE parameters.
* ``'fastformer'`` - FastFormer BASE parameters.
* ``'tiny-fastformer'`` - FastFormer TINY parameters.
quantized : bool, optional (default=False)
if True, will load 8-bit quantized model.
Quantized model not necessary faster, totally depends on the machine.
Returns
-------
result: model
List of model classes:
* if `bert` in model, will return `malaya.model.bert.MulticlassBERT`.
* if `xlnet` in model, will return `malaya.model.xlnet.MulticlassXLNET`.
* if `bigbird` in model, will return `malaya.model.xlnet.MulticlassBigBird`.
* if `fastformer` in model, will return `malaya.model.fastformer.MulticlassFastFormer`.
"""
```
```
model = malaya.relevancy.transformer(model = 'tiny-bigbird')
```
### Load Quantized model
To load 8-bit quantized model, simply pass `quantized = True`, default is `False`.
We can expect slightly accuracy drop from quantized model, and not necessary faster than normal 32-bit float model, totally depends on machine.
```
quantized_model = malaya.relevancy.transformer(model = 'alxlnet', quantized = True)
```
#### Predict batch of strings
```python
def predict(self, strings: List[str]):
"""
classify list of strings.
Parameters
----------
strings: List[str]
Returns
-------
result: List[str]
"""
```
```
%%time
model.predict([negative_text, positive_text])
%%time
quantized_model.predict([negative_text, positive_text])
```
#### Predict batch of strings with probability
```python
def predict_proba(self, strings: List[str]):
"""
classify list of strings and return probability.
Parameters
----------
strings : List[str]
Returns
-------
result: List[dict[str, float]]
"""
```
```
%%time
model.predict_proba([negative_text, positive_text])
%%time
quantized_model.predict_proba([negative_text, positive_text])
```
#### Open relevancy visualization dashboard
Default when you call `predict_words` it will open a browser with visualization dashboard, you can disable by `visualization=False`.
```python
def predict_words(
self,
string: str,
method: str = 'last',
bins_size: float = 0.05,
visualization: bool = True,
):
"""
classify words.
Parameters
----------
string : str
method : str, optional (default='last')
Attention layer supported. Allowed values:
* ``'last'`` - attention from last layer.
* ``'first'`` - attention from first layer.
* ``'mean'`` - average attentions from all layers.
bins_size: float, optional (default=0.05)
default bins size for word distribution histogram.
visualization: bool, optional (default=True)
If True, it will open the visualization dashboard.
Returns
-------
dictionary: results
"""
```
**This method not available for BigBird models**.
```
quantized_model.predict_words(negative_text)
```
### Vectorize
Let say you want to visualize sentence / word level in lower dimension, you can use `model.vectorize`,
```python
def vectorize(self, strings: List[str], method: str = 'first'):
"""
vectorize list of strings.
Parameters
----------
strings: List[str]
method : str, optional (default='first')
Vectorization layer supported. Allowed values:
* ``'last'`` - vector from last sequence.
* ``'first'`` - vector from first sequence.
* ``'mean'`` - average vectors from all sequences.
* ``'word'`` - average vectors based on tokens.
Returns
-------
result: np.array
"""
```
#### Sentence level
```
texts = [negative_text, positive_text]
r = model.vectorize(texts, method = 'first')
from sklearn.manifold import TSNE
import matplotlib.pyplot as plt
tsne = TSNE().fit_transform(r)
tsne.shape
plt.figure(figsize = (7, 7))
plt.scatter(tsne[:, 0], tsne[:, 1])
labels = texts
for label, x, y in zip(
labels, tsne[:, 0], tsne[:, 1]
):
label = (
'%s, %.3f' % (label[0], label[1])
if isinstance(label, list)
else label
)
plt.annotate(
label,
xy = (x, y),
xytext = (0, 0),
textcoords = 'offset points',
)
```
#### Word level
```
r = quantized_model.vectorize(texts, method = 'word')
x, y = [], []
for row in r:
x.extend([i[0] for i in row])
y.extend([i[1] for i in row])
tsne = TSNE().fit_transform(y)
tsne.shape
plt.figure(figsize = (7, 7))
plt.scatter(tsne[:, 0], tsne[:, 1])
labels = x
for label, x, y in zip(
labels, tsne[:, 0], tsne[:, 1]
):
label = (
'%s, %.3f' % (label[0], label[1])
if isinstance(label, list)
else label
)
plt.annotate(
label,
xy = (x, y),
xytext = (0, 0),
textcoords = 'offset points',
)
```
Pretty good, the model able to know cluster bottom left as positive relevancy.
### Stacking models
More information, you can read at [https://malaya.readthedocs.io/en/latest/Stack.html](https://malaya.readthedocs.io/en/latest/Stack.html)
```
albert = malaya.relevancy.transformer(model = 'albert')
malaya.stack.predict_stack([albert, model], [positive_text, negative_text])
```
| github_jupyter |
# VacationPy
----
#### Note
* Keep an eye on your API usage. Use https://developers.google.com/maps/reporting/gmp-reporting as reference for how to monitor your usage and billing.
* Instructions have been included for each segment. You do not have to follow them exactly, but they are included to help you think through the steps.
```
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import requests
import gmaps
import os
# Import API key
from config import g_key
```
### Store Part I results into DataFrame
* Load the csv exported in Part I to a DataFrame
```
# Read cities file and store into Pandas data frame
file_to_load = "../WeatherPy/output_data/cities.csv"
cities_df = pd.read_csv(file_to_load)
cities_df.head()
```
### Humidity Heatmap
* Configure gmaps.
* Use the Lat and Lng as locations and Humidity as the weight.
* Add Heatmap layer to map.
```
#Configure gmaps
gmaps.configure(api_key=g_key)
#Use the lat and Lng as locations and humidity as the weight
geolocations = cities_df[["Lat", "Lng"]].astype(float)
humidity = cities_df["Humidity"].astype(float)
#Add Heatmap layer to map
fig = gmaps.figure(center=(20,0), zoom_level=1.5)
heat_layer = gmaps.heatmap_layer(geolocations, weights=humidity,
dissipating=False, max_intensity=500,
point_radius = 4)
fig.add_layer(heat_layer)
fig
```
### Create new DataFrame fitting weather criteria
* Narrow down the cities to fit weather conditions.
* Drop any rows will null values.
```
# Narrow down the DataFrame to find your ideal weather condition.
# A max temperature lower than 80 degrees but higher than 70.
# Wind speed less than 10 mph.
# Zero cloudiness.
# Drop any rows that don't contain all three conditions. You want to be sure the weather is ideal.
narrow_df = cities_df.loc[(cities_df["Max Temp"] > 70) & (cities_df["Max Temp"] < 80)
& (cities_df["Wind Speed"] < 10)
& (cities_df["Cloudiness"] == 0 )]
narrow_df.dropna()
narrow_df
#Total 8 rows returned, reasonable count for api hits
```
### Hotel Map
* Store into variable named `hotel_df`.
* Add a "Hotel Name" column to the DataFrame.
* Set parameters to search for hotels with 5000 meters.
* Hit the Google Places API for each city's coordinates.
* Store the first Hotel result into the DataFrame.
* Plot markers on top of the heatmap.
```
#Store filtered and narrow data frame from above to a new data frame that will include hotel information
#Note: received a SettingWithCopyWarning when simply using the = operator. Using loc and copy per documentation
hotel_df = narrow_df[0:len(narrow_df)].copy()
hotel_df["Hotel Name"] = ""
hotel_df["Hotel Address"] = ""
#Set parameters to search for hotels within 5000 meters
params = {
"radius" : 5000,
"keyword" : "hotel",
"key" : g_key
}
for index, row in hotel_df.iterrows():
# get lat, lng from df
lat = row["Lat"]
lng = row["Lng"]
# change location each iteration while leaving original params in place
params["location"] = f"{lat},{lng}"
# Use the lat/lng and the rest of the params as set earlier
base_url = "https://maps.googleapis.com/maps/api/place/nearbysearch/json"
hotels = requests.get(base_url, params=params)
hotels = hotels.json()
try:
hotel_df.loc[index, "Hotel Name"] = hotels["results"][0]["name"]
hotel_df.loc[index, "Hotel Address"] = hotels["results"][0]["vicinity"]
print(f"Hotel found")
except (KeyError, IndexError):
print("Missing field/result... skipping.")
hotel_df
# NOTE: Do not change any of the code in this cell
# Using the template add the hotel marks to the heatmap
info_box_template = """
<dl>
<dt>Name</dt><dd>{Hotel Name}</dd>
<dt>City</dt><dd>{City}</dd>
<dt>Country</dt><dd>{Country}</dd>
</dl>
"""
# Store the DataFrame Row
# NOTE: be sure to update with your DataFrame name
hotel_info = [info_box_template.format(**row) for index, row in hotel_df.iterrows()]
locations = hotel_df[["Lat", "Lng"]]
# Add marker layer ontop of heat map
markers = gmaps.marker_layer(locations, info_box_content=hotel_info)
fig.add_layer(markers)
# Display figure
fig
#Please note, screenshot of the final image included within "output_data/map.png"
```
| github_jupyter |
```
%matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
# plt.style.use('ggplot')
import pysdaza as sd
```
# Reentry Chile Response Rate
*Sebastian Daza*
In this report we describe response rates by wave. We follow these general criteria:
- The estimation by wave only uses cases far beyond the wave-specific observation window. For instance, for the wave 2-months, we only consider those women who have been in the study for 4 months.
- This estimation is based on the administrative records of the study.
Some important definitions:
- Names of waves: baseline, week, two_months, six_months, twelve_months
- Variables names: `d` means date, `c` means clean, and `cd` clean date
- Deadline corresponds to th time threshold to compute a given response rate
We display key date variables to compute the rates and highlight inconsistencies or problems.
```
# getting paths
import os
myfile = os.getcwd() + '/data/Registro general de terreno_todas las olas.xlsx'
```
## Release
There are two date of release variables
in the register data. *Definitive* has a bunch of missing records. **We should define a validated and unique date for release**.
```
import pandas as pd
import numpy as np
import re
# read excel file
df = pd.read_excel(myfile, skiprows=[0,1])
df.columns = df.columns.str.strip()
variables = set(df.columns.tolist())
# extract columns
cols = ['enc','folio','fegresoefectiva', 'lb_fechaentrevista', 'lb_participa',
'primsem_fechaentrevista', 'primsem_participa', 'dosmeses_fechaentrevista',
'dosmeses_participa', 'seismeses_fechaentrevista', 'seismeses_participa',
'docemeses_fechaentrevista', 'docemeses_participa']
len(cols)
df = df.loc[:,cols]
cols = ['enc','folio','fegresoefectiva', 'lb_fechaentrevista', 'lb_participa',
'primsem_fechaentrevista', 'primsem_participa', 'dosmeses_fechaentrevista',
'dosmeses_participa', 'seismeses_fechaentrevista', 'seismeses_participa',
'docemeses_fechaentrevista', 'docemeses_participa']
# rename columns
names = ["int","id", "release_f","dbaseline","r_baseline",
"dweek","r_week","d2months","r_2months",
"d6months","r_6months","d12months","r_12months"]
df.rename(columns=dict(zip(df.columns, names)), inplace=True)
null = df.int.notnull()
df = df.loc[null, names]
df.shape
# remove cases?
remove_cases = [10011] # only case that's not valid
# df.id in remove_cases
df = df.loc[~df.id.isin(remove_cases),:]
df.shape
# # save data
# import dill
# dill.dump(df, open('data/registro.pkl', 'wb'))
# # load the data
# df = dill.load(open('data/registro.pkl', 'rb'))
# no valid records for release_f?
s = df.release_f.isnull()
s.describe()
df['release_year'] = pd.DatetimeIndex(df['release_f']).year
df['release_month'] = pd.DatetimeIndex(df['release_f']).month
# release plot
t = df.reset_index().set_index('release_f')
t = t.resample('M')
ax = t.id.count().plot(title='Egresos por Mes', style='.-', ms=13);
ax.set_ylabel("Número de mujeres");
ax.set_xlabel("Mes de egreso");
```
## Baseline
Let's explore the baseline information. Below a plot with the distribution of the difference between the date of the baseline and release. All values should be **negative**.
To check cases below where we have issues: missing data or positive value (i.e., the baseline was applied after release).
```
# correct some dates
df.loc[df.dbaseline.isnull(),['int','id', 'dbaseline', 'r_baseline']]
# function to clean dates
def clean_dates(text):
if (isinstance(text, str)):
text = str(text)
pp = re.compile("-|\.")
r = re.sub(pp, "/", text)
pp = re.compile("([0-9]+/[0-9]+/[0-9]+)|([0-9]+-[0-9]+-[0-9]+)")
r = re.search(pp, r)
if (r is not None):
r = r.group()
else:
r = ''
else:
r = text
return pd.to_datetime(r, dayfirst=True, errors = 'coerce')
df['cdbaseline'] = [clean_dates(i) for i in df['dbaseline']]
df = df.assign(release_baseline = (df.cdbaseline - df.release_f).dt.days) # it has to be negative
s = ((df.release_baseline > 0) | (df.cdbaseline.isnull()))
df.loc[s, ['int', 'id', 'r_baseline', 'release_f', 'cdbaseline', 'dbaseline', 'release_baseline']].sort_values('id')
```
Descriptives of the difference between release date and baseline:
```
df.release_baseline.describe()
```
## First week
Below a plot with the distribution of the difference between the date of the first week interview and release. All values should be **positive**.
```
df['r_week'] = df['r_week'].str.contains("s|S").astype(float)
df['r_week'] = df.r_week.fillna(0)
df.r_week.value_counts()
df['cdweek'] = [clean_dates(i) for i in df['dweek']]
m = df.cdweek.isnull() & df.dweek.notnull()
df = df.assign(release_week = (df.cdweek - df.release_f).dt.days)
(df.release_week > 7).value_counts()
```
Plot of the difference between date of release and first week interview.
```
ax = df.release_week.hist(bins = 20, alpha = 0.8);
ax.set_xlim(-25,25)
ax.set_xlabel('Día de entrevista desde egreso');
df['week_deadline'] = pd.DatetimeIndex( df['release_f'] ) + pd.DateOffset(weeks = 5) # five weeks!
```
Cases with issues:
```
import datetime
today = datetime.datetime.now()
s = ((df.r_week == 1) & (today > df.week_deadline)) & ( (df.release_week <= 0) | (df.cdweek.isnull()) )
df.loc[np.abs(df.release_week) > 30,['int', 'id', 'r_week', 'release_f', 'cdweek', 'release_week']]
```
Interviews before the first 7 days. The only weird case is 10053!
```
df.loc[df.release_week < 7,['int', 'id', 'r_week', 'release_f', 'cdweek', 'release_week']]
```
Response rate:
```
tab = df.loc[ (today > df.week_deadline) | (df.r_week == 1), 'r_week'].value_counts()
rr_week = [i / sum(list(tab)) for i in tab][0]
print(rr_week)
```
Descriptives of the difference between date of release and interview first week.
```
df.loc[df.r_week == 1, 'release_week'].describe()
```
# Two Months
```
df['r_2months'] = df['r_2months'].str.contains("s|S").astype(float)
df['r_2months'] = df.r_2months.fillna(0)
df['cd2months'] = [clean_dates(i) for i in df['d2months']]
```
Some cases with issues:
```
# replace errors in data entry
# errors = ['27/2']
# new_values = ['02/27/2017']
# for i in range(0,len(errors)):
# df.loc[df['d2months'] == errors[i], 'cd2months'] = pd.to_datetime(new_values[i])
s = (df.r_2months == 1)
#df.loc[s, 'cd2months'].isnull().value_counts()
df = df.assign(release_2months = (df.cd2months - df.release_f).dt.days)
#df.release_2months.describe()
```
Plot of the difference between release and date of two-months interview.
```
ax = df.release_2months.hist(bins = 60);
ax.set_xlabel('Días desde la salida en que la encuesta de 2 meses fue aplicada');
```
Some issues:
```
df['two_months_deadline'] = pd.DatetimeIndex( df['release_f'] ) + pd.DateOffset(months = 5)
s = ((df.r_2months == 1) & (today > df.two_months_deadline)) & ( (df.release_2months <= 0) | (df.cd2months.isnull()))
df.loc[s,['int', 'id', 'r_2months', 'release_f', 'cd2months', 'two_months_deadline', 'release_2months']].sort_values('id')
tab = df.loc[ (today > df.two_months_deadline) | (df.r_2months == 1), 'r_2months'].value_counts()
# print(tab)
rr_2months = [i / sum(list(tab)) for i in tab][0]
# print(rr_2months)
df.loc[np.abs(df.release_2months) > 100,['int', 'id', 'r_2months', 'release_f',
'cd2months', 'two_months_deadline',
'release_2months']].sort_values('release_2months')
df.loc[np.abs(df.release_2months) < 10 ,['int', 'id',
'r_2months', 'release_f', 'cd2months',
'two_months_deadline',
'release_2months']].sort_values('release_2months')
```
Descriptives:
```
df.loc[df.r_2months == 1, 'release_2months'].describe()
```
## Six Months
```
df.r_6months.value_counts()
df['r_6months'] = df['r_6months'].str.contains("s|S").astype(float)
df['r_6months'] = df.r_6months.fillna(0)
df['cd6months'] = [clean_dates(i) for i in df['d6months']]
m = df.cd6months.isnull() & df.d6months.notnull()
m.value_counts()
s = (df.r_6months == 1)
```
There is some issues here:
```
df.loc[df.id == 50209, 'cd6months']
df = df.assign(release_6months = (df.cd6months - df.release_f).dt.days)
sns.distplot(df.loc[df.r_6months==1, 'release_6months'], bins=25, kde=False);
plt.axvline(x=(30.5*6), color='r', ls='dotted', alpha=0.6, lw=1)
plt.text(187, 31, '6 meses', alpha=0.8)
plt.xlabel('Días desde la salida en que la encuesta de 6 meses fue aplicada');
plt.savefig('plots/dias_6meses.pdf', dpi = 600)
df['six_months_deadline'] = pd.DatetimeIndex( df['release_f'] ) + pd.DateOffset(months = 7)
s = ((df.r_6months == 1) & (today > df.six_months_deadline)) & (
(df.release_6months <= 0) | (df.cd6months.isnull()))
df.loc[s,['int', 'id', 'r_6months', 'release_f', 'd6months',
'cd6months', 'six_months_deadline', 'release_6months']].sort_values('id')
```
### Cases higher than 240 days since release
r6_months (if respondent took part of wave 6 months) is not always consistent!
```
df.loc[np.abs(df.release_6months) > 240, ['int', 'id', 'r_6months', 'release_f', 'cd6months',
'six_months_deadline',
'release_6months']].sort_values('release_6months')
```
30247 is fine?|
### Cases lower than 150 days since release
Same issue with r_6months
```
df.loc[np.abs(df.release_6months) < 150, ['int', 'id', 'r_6months', 'release_f', 'cd6months',
'six_months_deadline',
'release_6months']].sort_values('release_6months')
tab = df.loc[(today > df.six_months_deadline) | (df.r_6months == 1), 'r_6months'].value_counts()
# print(tab)
rr_6months = [i / sum(list(tab)) for i in tab][0]
print(rr_6months)
```
Some descriptives:
```
df.loc[df.r_6months == 1, 'release_6months'].describe()
```
# Year
```
df.r_12months.value_counts()
df['r_12months'] = df['r_12months'].str.contains("s|S").astype(float)
df['r_12months'] = df.r_12months.fillna(0)
df.r_12months.value_counts()
df['cd12months'] = [clean_dates(i) for i in df['d12months']]
m = df.cd12months.isnull() & df.d12months.notnull()
m.value_counts()
s = (df.r_12months == 1)
s.value_counts()
df = df.assign(release_12months = (df.cd12months - df.release_f).dt.days)
ax = df.release_12months.hist(bins = 20);
ax.set_xlabel('Días desde la salida en que la encuesta de 12 meses fue aplicada')
plt.savefig('plots/dias_12meses.pdf', dpi = 600);
df['twelve_months_deadline'] = pd.DatetimeIndex( df['release_f'] ) + pd.DateOffset(months = 12)
s = ((df.r_12months == 1) & (today > df.twelve_months_deadline)) & (
(df.release_12months <= 0) | (df.cd12months.isnull()))
df.loc[s,['int', 'id', 'r_12months', 'release_f', 'd12months',
'cd12months', 'twelve_months_deadline', 'release_12months']].sort_values('id')
tab = df.loc[(today > df.twelve_months_deadline) | (df.r_12months == 1), 'r_12months'].value_counts()
# print(tab)
rr_12months = [i / sum(list(tab)) for i in tab][0]
print(rr_12months)
df.loc[np.abs(df.release_12months) > 380, ['int', 'id', 'r_12months', 'release_f', 'cd12months',
'twelve_months_deadline',
'release_12months']].sort_values('release_12months')
df.loc[df.r_12months == 1, 'release_12months'].describe()
```
# Plot of interview days by wave
```
ax = df.loc[:, ['release_baseline', 'release_week', 'release_2months',
'release_6months', 'release_12months']].plot.hist(
# title='Distribución Día de Entrevista según Ola',
alpha=0.6, bins = 300
)
ax.set_xlim(-40,500)
ax.set_ylabel('Frecuencia')
ax.legend(['Línea base', 'Primera semana', 'Dos meses', 'Seis meses', 'Doce meses'])
ax.axvline(7, color = 'gray', linestyle ='--', linewidth=1)
ax.axvline(30.5*2, color = 'gray', linestyle ='--', linewidth=1)
ax.axvline(30.5*6, color = 'gray', linestyle ='--', linewidth=1)
ax.axvline(365.0, color = 'gray', linestyle ='--', linewidth=1)
ax.set_xlabel('Días desde egreso');
plt.savefig('plots/dia_entrevista_ola.pdf', dpi = 600)
```
## Summary rates
```
rr = pd.DataFrame( {"rate" : [rr_week, rr_2months, rr_6months, rr_12months]},
index = ['week' ,'2 months', '6 months', '12 months'])
rr
```
| github_jupyter |
```
import pandas as pd
import pickle
from tqdm import tqdm_notebook
```
## Load Datasets
## Exploring Train
```
with open(r"C:\Users\am21907\Documents\Projects\Hackathon Projects\RwD\neural-hawkes-particle-smoothing\logs\pilotnhp7\dim-16_batch-50_seed-12345_lr-0.001_3464\bt-add-dim-16-batch-50-npn-1-np-1-ga-1.0-lr-0.001-seed-12345_18612\dev.results.pkl","rb") as f:
pkl_results = pickle.load(f)
pkl_results.keys()
pkl_results['MAP'].keys()
pd.DataFrame.from_dict(pkl_results['MAP']['nhps'][5000]).T
pkl_results['MBR']['nhpf']
pkl_results['LogProposal']['nhps']#[5000]
with open('data/pilottaxi/train.pkl', 'rb') as f:
pkl_train = pickle.load(f)
pkl_train.keys()
with open('data/pilottaxi/test.pkl', 'rb') as f:
pkl_test = pickle.load(f)
pkl_train.keys()
len(pkl_train['seqs'])
len(pkl_dev['seqs'])
!
pkl_train.k
pkl_train.
pkl_train.keys()
pkl_train['seqs'][0][:5]
pkl_train['seqs_obs'][0]
temp = pd.DataFrame(pkl_train['seqs'][0])
print(temp.shape)
temp.head()
temp = pd.DataFrame(pkl_train['seqs_obs'][0])
print(temp.shape)
temp.head()
pkl_train['total_num']
```
### seq_obs
```
## convert dict to df
seq_obs_df = pd.DataFrame()
for i in tqdm_notebook(range(0,len(pkl_train['seqs_obs']))):
temp_df = (pd.DataFrame(pkl_train['seqs_obs'][i]))
temp_df['id'] = i
seq_obs_df = pd.concat([seq_obs_df,temp_df])
seq_obs_df.head()
# check all event type occurances
seq_obs_df['type_event'].value_counts(dropna=False)
# check all event idx occurances
seq_obs_df['idx_event'].value_counts(dropna=False)
```
### seq
```
## convert dict to df
seqs_df = pd.DataFrame()
for i in tqdm_notebook(range(0,len(pkl_train['seqs']))):
temp_df = (pd.DataFrame(pkl_train['seqs'][i]))
temp_df['id'] = i
seqs_df = pd.concat([seqs_df,temp_df])
seqs_df.head()
# check all event type occurances
seqs_df['type_event'].value_counts(dropna=False)
```
#unique_event_types matches the number of events provided in censor.config
```
# check all event idx occurances
seqs_df['idx_event'].value_counts(dropna=False)
```
## Mimic training process
### nhps
```
from copy import deepcopy
data = deepcopy(pkl_train['seqs'])
args = {"MaxEpoch":2,"TrackPeriod":5000}
max_episode = args['MaxEpoch'] * len(data)
report_gap = args['TrackPeriod']
time_sample = 0.0
time_train_only = 0.0
time_dev_only = 0.0
input = []
input_particles = []
for episode in range(max_episode):
idx_seq = episode % len(data)
idx_epoch = episode // len(data)
one_seq = data[ idx_seq ]
break
one_seq
pkl_train.keys()
file = "dev.pkl"
with open('data/pilottaxi/'+file, 'rb') as f:
pkl_train = pickle.load(f)
temp = pkl_train.pop('seqs_obs')
with open('data/pilottaxi/'+file, 'wb') as f:
pickle.dump(pkl_train,f)
```
| github_jupyter |
# Graph From Bounding Box
Create a graph from OSM within some bounding box.
```
# OSMnx: New Methods for Acquiring, Constructing, Analyzing, and Visualizing Complex Street Networks
import osmnx as ox
ox.config(use_cache=True, log_console=False)
ox.__version__
```
## all_private
```
north, south, east, west = 37.79, 37.78, -122.41, -122.43 # San Francisco
network_type = 'all_private' # "all_private", "all", "bike", "drive", "drive_service", "walk"
# Create a graph from OSM within some bounding box.
G = ox.graph_from_bbox(
north, south, east, west,
network_type=network_type,
simplify=True,
retain_all=False,
truncate_by_edge=False,
clean_periphery=True,
custom_filter=None)
# Plot a graph.
fig, ax = ox.plot_graph(G)
```
## all
```
north, south, east, west = 37.79, 37.78, -122.41, -122.43 # San Francisco
network_type = 'all' # "all_private", "all", "bike", "drive", "drive_service", "walk"
# Create a graph from OSM within some bounding box.
G2 = ox.graph_from_bbox(
north, south, east, west,
network_type=network_type,
simplify=True,
retain_all=False,
truncate_by_edge=False,
clean_periphery=True,
custom_filter=None)
# Plot a graph.
fig, ax = ox.plot_graph(G2)
```
## bike
```
north, south, east, west = 37.79, 37.78, -122.41, -122.43 # San Francisco
network_type = 'bike' # "all_private", "all", "bike", "drive", "drive_service", "walk"
# Create a graph from OSM within some bounding box.
G3 = ox.graph_from_bbox(
north, south, east, west,
network_type=network_type,
simplify=True,
retain_all=False,
truncate_by_edge=False,
clean_periphery=True,
custom_filter=None)
# Plot a graph.
fig, ax = ox.plot_graph(G3)
```
## drive
```
north, south, east, west = 37.79, 37.78, -122.41, -122.43 # San Francisco
network_type = 'drive' # "all_private", "all", "bike", "drive", "drive_service", "walk"
# Create a graph from OSM within some bounding box.
G4 = ox.graph_from_bbox(
north, south, east, west,
network_type=network_type,
simplify=True,
retain_all=False,
truncate_by_edge=False,
clean_periphery=True,
custom_filter=None)
# Plot a graph.
fig, ax = ox.plot_graph(G4)
```
## drive_service
```
north, south, east, west = 37.79, 37.78, -122.41, -122.43 # San Francisco
network_type = 'drive_service' # "all_private", "all", "bike", "drive", "drive_service", "walk"
# Create a graph from OSM within some bounding box.
G5 = ox.graph_from_bbox(
north, south, east, west,
network_type=network_type,
simplify=True,
retain_all=False,
truncate_by_edge=False,
clean_periphery=True,
custom_filter=None)
# Plot a graph.
fig, ax = ox.plot_graph(G5)
```
## walk
```
north, south, east, west = 37.79, 37.78, -122.41, -122.43 # San Francisco
network_type = 'walk' # "all_private", "all", "bike", "drive", "drive_service", "walk"
# Create a graph from OSM within some bounding box.
G6 = ox.graph_from_bbox(
north, south, east, west,
network_type=network_type,
simplify=True,
retain_all=False,
truncate_by_edge=False,
clean_periphery=True,
custom_filter=None)
# Plot a graph.
fig, ax = ox.plot_graph(G6)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/oferbaharav/DS-Unit-1-Sprint-1-Dealing-With-Data/blob/master/module4-sequence-your-narrative/Copy_of_LS_DS_124_Sequence_your_narrative_Assignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
_Lambda School Data Science_
# Sequence Your Narrative - Assignment
Today we will create a sequence of visualizations inspired by [Hans Rosling's 200 Countries, 200 Years, 4 Minutes](https://www.youtube.com/watch?v=jbkSRLYSojo).
Using this [data from Gapminder](https://github.com/open-numbers/ddf--gapminder--systema_globalis/):
- [Income Per Person (GDP Per Capital, Inflation Adjusted) by Geo & Time](https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--datapoints--income_per_person_gdppercapita_ppp_inflation_adjusted--by--geo--time.csv)
- [Life Expectancy (in Years) by Geo & Time](https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--datapoints--life_expectancy_years--by--geo--time.csv)
- [Population Totals, by Geo & Time](https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--datapoints--population_total--by--geo--time.csv)
- [Entities](https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--entities--geo--country.csv)
- [Concepts](https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--concepts.csv)
Objectives
- sequence multiple visualizations
- combine qualitative anecdotes with quantitative aggregates
Links
- [Hans Rosling’s TED talks](https://www.ted.com/speakers/hans_rosling)
- [Spiralling global temperatures from 1850-2016](https://twitter.com/ed_hawkins/status/729753441459945474)
- "[The Pudding](https://pudding.cool/) explains ideas debated in culture with visual essays."
- [A Data Point Walks Into a Bar](https://lisacharlotterost.github.io/2016/12/27/datapoint-in-bar/): a thoughtful blog post about emotion and empathy in data storytelling
# ASSIGNMENT
1. Replicate the Lesson Code
2. Take it further by using the same gapminder dataset to create a sequence of visualizations that combined tell a story of your choosing.
Get creative! Use text annotations to call out specific countries, maybe: change how the points are colored, change the opacity of the points, change their sized, pick a specific time window. Maybe only work with a subset of countries, change fonts, change background colors, etc. make it your own!
```
# TODO
import pandas as pd
import matplotlib.pyplot as plt
df = pd.read_csv('https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--datapoints--gdp_total_yearly_growth--by--geo--time.csv')
df.head(10)
import seaborn as sns
sns.set(style="ticks")
g= sns.relplot(x="gdp_total_yearly_growth", y="time", hue="geo", data=df, legend = False)
entities = pd.read_csv('https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--entities--geo--country.csv')
entities.head()
new_df = pd.merge(df, entities[['world_6region', 'country']], left_on = 'geo', right_on = 'country')
new_df.head()
import seaborn as sns
sns.set(style="ticks")
g= sns.relplot(x="gdp_total_yearly_growth", y="time", hue="world_6region", data=new_df)
```
# STRETCH OPTIONS
## 1. Animate!
- [How to Create Animated Graphs in Python](https://towardsdatascience.com/how-to-create-animated-graphs-in-python-bb619cc2dec1)
- Try using [Plotly](https://plot.ly/python/animations/)!
- [The Ultimate Day of Chicago Bikeshare](https://chrisluedtke.github.io/divvy-data.html) (Lambda School Data Science student)
- [Using Phoebe for animations in Google Colab](https://colab.research.google.com/github/phoebe-project/phoebe2-docs/blob/2.1/tutorials/animations.ipynb)
## 2. Study for the Sprint Challenge
- Concatenate DataFrames
- Merge DataFrames
- Reshape data with `pivot_table()` and `.melt()`
- Be able to reproduce a FiveThirtyEight graph using Matplotlib or Seaborn.
## 3. Work on anything related to your portfolio site / Data Storytelling Project
```
# TODO
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib
import matplotlib.pyplot as plt
import matplotlib.animation as animation
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import datetime
import seaborn as sns
import matplotlib.pyplot as plt
from gs_quant.markets.portfolio import Portfolio
from gs_quant.risk import MarketDataShockBasedScenario, MarketDataPattern, MarketDataShock, MarketDataShockType, PnlExplain
from gs_quant.markets import PricingContext
plt.rcParams['figure.figsize'] = (16, 8)
from gs_quant.session import GsSession
# external users should substitute their client id and secret; please skip this step if using internal jupyterhub
GsSession.use(client_id=None, client_secret=None, scopes=('run_analytics', 'read_product_data'))
```
In this notebook, we'll load a portfolio and run analytics to understand it's risk and pnl. We'll also run a number of vol and spot scenarios which we'll use to compute VaR.
The content of this notebook is split into the following parts:
* [1: First, the portfolio](#1:-First,-the-portfolio)
* [2: Run risk and PnL explain](#2:-Run-risk-and-PnL-explain)
* [3: Scenarios grid and VaR](#3:-Scenarios-grid-and-VaR)
### 1: First, the portfolio
Let’s first load a portfolio from csv, mapping each column and look at it as a dataframe. Internal users can load this books directly using `from book`.
```
mappers = {
'pair': lambda row: row['foreign ccy'] + 'USD',
'notional_amount': 'notional',
'expiration_date': 'expiry',
'option_type': lambda row: 'Call' if row['C/P'] == 'C' else 'Put',
'strike_price': 'strike',
'premium': lambda row: 0
}
portfolio = Portfolio.from_csv(r'FXBook.csv', mappings=mappers)
portfolio.resolve()
frame = portfolio.to_frame()
frame.index = frame.index.droplevel(0)
frame.head(1).transpose()
# for internal users:
# portfolio = Portfolio.from_book('my_book_id')
```
### 2: Run risk and PnL explain
With the portfolio in hand, let’s use gs-quant to understand risk and pnl over the last business day.
```
from gs_quant.datetime.date import business_day_offset
from gs_quant.markets import CloseMarket, PricingContext, close_market_date
from gs_quant.risk import Price, DollarPrice, PnlExplain, Theta, FXDelta, FXGamma, FXVega
to_date = business_day_offset(close_market_date(), -1)
# Previous business day
from_date = business_day_offset(to_date, -1)
# A risk measure for calculating PnlExplain from that date
explain = PnlExplain(CloseMarket(date=to_date))
# Calculate PnlExplain and dollar price from 1 day ago
with PricingContext(pricing_date=from_date):
result = portfolio.calc((FXDelta, FXGamma, FXVega, DollarPrice, Theta, explain))
# Calculate dollar price with today's market
with PricingContext(pricing_date=to_date):
target_price = portfolio.calc((DollarPrice))
with PricingContext(pricing_date=from_date, market=CloseMarket(date=to_date)):
target_price_ex_theta = portfolio.calc((DollarPrice))
print('Risk and 1day Pnl as of '+str(from_date))
print(f'Book PV (in mUSD): {target_price[DollarPrice].aggregate()/1e6:.1f}')
print(f'Book Delta (in mUSD): {result[FXDelta].aggregate().value.sum()/1e6:.0f}')
print(f'Book Vega (in kUSD): {result[FXVega].aggregate().value.sum():.0f}')
print(f'Dollar price day on day change (in kUSD): {target_price[DollarPrice].aggregate()/1e3 - result[DollarPrice].aggregate()/1e3:.0f}')
print(f'Dollar price day on day change(ex theta) (in kUSD): {target_price_ex_theta[DollarPrice].aggregate()/1e3 - result[DollarPrice].aggregate()/1e3:.0f}')
print(f'Pnl explain total (in kUSD): {result[explain].aggregate().value.sum()/1e3:.0f}')
print(f'Theta total (in kUSD): {result[Theta].aggregate().value.sum()/1e3:.0f}')
print(f'Theta + Pnl explain total (in kUSD): {result[Theta].aggregate().value.sum()/1e3 + result[explain].aggregate().value.sum()/1e3:.0f}')
# Show the PnlExplain breakdown
explain_all = result[explain].aggregate()
explain_all[explain_all.value.abs() > 1.0].round(0)
```
### 3: Scenarios grid and VaR
We can also better understand risk by applying a number of market shocks – in this case we’ll look at a grid of vol and spot shocks, we are can also use to calculate VaR by looking at the 95% percentile price change.
```
shocks_fx = [-10, -7.5, -5, -2,-1, 0, 1, 2, 5, 7.5, 10]
shocks_vols = [-5, -2, -1, -0.5, 0, 0.5, 1, 2, 5, 7.5, 10]
shocked_prices = pd.DataFrame(index=shocks_vols, columns=shocks_fx)
cross = explain_all[explain_all['mkt_type'] == 'FX']['mkt_asset'][0]
with PricingContext(is_async=True):
for fx in shocks_fx:
for vol in shocks_vols:
with MarketDataShockBasedScenario({
MarketDataPattern('FX', cross): MarketDataShock(MarketDataShockType.Proportional, fx / 1e2),
MarketDataPattern('FX Vol', cross, 'ATM Vol'): MarketDataShock(MarketDataShockType.Absolute, vol / 1e2),
}):
shocked_prices[fx][vol] = portfolio.calc((DollarPrice))
# Aggregate and compute pnl by substracting 0 shock PV
shocked_prices_res = pd.DataFrame(index=shocks_vols, columns=shocks_fx, dtype='float')
for fx in shocks_fx:
for vol in shocks_vols:
shocked_prices_res[fx][vol] = shocked_prices[fx][vol].aggregate()
shocked_prices_res -= shocked_prices[0][0].aggregate()
shocked_prices_res /= 1e3
ax = sns.heatmap(shocked_prices_res, cmap='coolwarm', annot=True, fmt='.1f')
ax.set(ylabel='absolute vol point moves', xlabel='% spot change', title='PV changes ($k)')
ax.xaxis.tick_top()
ax.xaxis.set_label_position('top')
```
Compute VaR
```
p = np.percentile(shocked_prices_res, 5) # return 95th percentile
print('Portfolio base price ($m): {:,.1f}'.format(portfolio.price().aggregate()/1e6))
print('Scenario Based VaR with spot/vol grid ($m): {:,.1f}'.format(p/1e3))
```
### Disclaimers
Scenarios/predictions: Simulated results are for illustrative purposes only. GS provides no assurance or guarantee that the strategy will operate or would have operated in the past in a manner consistent with the above analysis. Past performance figures are not a reliable indicator of future results.
Indicative Terms/Pricing Levels: This material may contain indicative terms only, including but not limited to pricing levels. There is no representation that any transaction can or could have been effected at such terms or prices. Proposed terms and conditions are for discussion purposes only. Finalized terms and conditions are subject to further discussion and negotiation.
www.goldmansachs.com/disclaimer/sales-and-trading-invest-rec-disclosures.html If you are not accessing this material via Marquee ContentStream, a list of the author's investment recommendations disseminated during the preceding 12 months and the proportion of the author's recommendations that are 'buy', 'hold', 'sell' or other over the previous 12 months is available by logging into Marquee ContentStream using the link below. Alternatively, if you do not have access to Marquee ContentStream, please contact your usual GS representative who will be able to provide this information to you.
Backtesting, Simulated Results, Sensitivity/Scenario Analysis or Spreadsheet Calculator or Model: There may be data presented herein that is solely for illustrative purposes and which may include among other things back testing, simulated results and scenario analyses. The information is based upon certain factors, assumptions and historical information that Goldman Sachs may in its discretion have considered appropriate, however, Goldman Sachs provides no assurance or guarantee that this product will operate or would have operated in the past in a manner consistent with these assumptions. In the event any of the assumptions used do not prove to be true, results are likely to vary materially from the examples shown herein. Additionally, the results may not reflect material economic and market factors, such as liquidity, transaction costs and other expenses which could reduce potential return.
OTC Derivatives Risk Disclosures:
Terms of the Transaction: To understand clearly the terms and conditions of any OTC derivative transaction you may enter into, you should carefully review the Master Agreement, including any related schedules, credit support documents, addenda and exhibits. You should not enter into OTC derivative transactions unless you understand the terms of the transaction you are entering into as well as the nature and extent of your risk exposure. You should also be satisfied that the OTC derivative transaction is appropriate for you in light of your circumstances and financial condition. You may be requested to post margin or collateral to support written OTC derivatives at levels consistent with the internal policies of Goldman Sachs.
Liquidity Risk: There is no public market for OTC derivative transactions and, therefore, it may be difficult or impossible to liquidate an existing position on favorable terms. Transfer Restrictions: OTC derivative transactions entered into with one or more affiliates of The Goldman Sachs Group, Inc. (Goldman Sachs) cannot be assigned or otherwise transferred without its prior written consent and, therefore, it may be impossible for you to transfer any OTC derivative transaction to a third party.
Conflict of Interests: Goldman Sachs may from time to time be an active participant on both sides of the market for the underlying securities, commodities, futures, options or any other derivative or instrument identical or related to those mentioned herein (together, "the Product"). Goldman Sachs at any time may have long or short positions in, or buy and sell Products (on a principal basis or otherwise) identical or related to those mentioned herein. Goldman Sachs hedging and trading activities may affect the value of the Products.
Counterparty Credit Risk: Because Goldman Sachs, may be obligated to make substantial payments to you as a condition of an OTC derivative transaction, you must evaluate the credit risk of doing business with Goldman Sachs or its affiliates.
Pricing and Valuation: The price of each OTC derivative transaction is individually negotiated between Goldman Sachs and each counterparty and Goldman Sachs does not represent or warrant that the prices for which it offers OTC derivative transactions are the best prices available, possibly making it difficult for you to establish what is a fair price for a particular OTC derivative transaction; The value or quoted price of the Product at any time, however, will reflect many factors and cannot be predicted. If Goldman Sachs makes a market in the offered Product, the price quoted by Goldman Sachs would reflect any changes in market conditions and other relevant factors, and the quoted price (and the value of the Product that Goldman Sachs will use for account statements or otherwise) could be higher or lower than the original price, and may be higher or lower than the value of the Product as determined by reference to pricing models used by Goldman Sachs. If at any time a third party dealer quotes a price to purchase the Product or otherwise values the Product, that price may be significantly different (higher or lower) than any price quoted by Goldman Sachs. Furthermore, if you sell the Product, you will likely be charged a commission for secondary market transactions, or the price will likely reflect a dealer discount. Goldman Sachs may conduct market making activities in the Product. To the extent Goldman Sachs makes a market, any price quoted for the OTC derivative transactions, Goldman Sachs may differ significantly from (i) their value determined by reference to Goldman Sachs pricing models and (ii) any price quoted by a third party. The market price of the OTC derivative transaction may be influenced by many unpredictable factors, including economic conditions, the creditworthiness of Goldman Sachs, the value of any underlyers, and certain actions taken by Goldman Sachs.
Market Making, Investing and Lending: Goldman Sachs engages in market making, investing and lending businesses for its own account and the accounts of its affiliates in the same or similar instruments underlying OTC derivative transactions (including such trading as Goldman Sachs deems appropriate in its sole discretion to hedge its market risk in any OTC derivative transaction whether between Goldman Sachs and you or with third parties) and such trading may affect the value of an OTC derivative transaction.
Early Termination Payments: The provisions of an OTC Derivative Transaction may allow for early termination and, in such cases, either you or Goldman Sachs may be required to make a potentially significant termination payment depending upon whether the OTC Derivative Transaction is in-the-money to Goldman Sachs or you at the time of termination. Indexes: Goldman Sachs does not warrant, and takes no responsibility for, the structure, method of computation or publication of any currency exchange rates, interest rates, indexes of such rates, or credit, equity or other indexes, unless Goldman Sachs specifically advises you otherwise.
Risk Disclosure Regarding futures, options, equity swaps, and other derivatives as well as non-investment-grade securities and ADRs: Please ensure that you have read and understood the current options, futures and security futures disclosure document before entering into any such transactions. Current United States listed options, futures and security futures disclosure documents are available from our sales representatives or at http://www.theocc.com/components/docs/riskstoc.pdf, http://www.goldmansachs.com/disclosures/risk-disclosure-for-futures.pdf and https://www.nfa.futures.org/investors/investor-resources/files/security-futures-disclosure.pdf, respectively. Certain transactions - including those involving futures, options, equity swaps, and other derivatives as well as non-investment-grade securities - give rise to substantial risk and are not available to nor suitable for all investors. If you have any questions about whether you are eligible to enter into these transactions with Goldman Sachs, please contact your sales representative. Foreign-currency-denominated securities are subject to fluctuations in exchange rates that could have an adverse effect on the value or price of, or income derived from, the investment. In addition, investors in securities such as ADRs, the values of which are influenced by foreign currencies, effectively assume currency risk.
Options Risk Disclosures: Options may trade at a value other than that which may be inferred from the current levels of interest rates, dividends (if applicable) and the underlier due to other factors including, but not limited to, expectations of future levels of interest rates, future levels of dividends and the volatility of the underlier at any time prior to maturity. Note: Options involve risk and are not suitable for all investors. Please ensure that you have read and understood the current options disclosure document before entering into any standardized options transactions. United States listed options disclosure documents are available from our sales representatives or at http://theocc.com/publications/risks/riskstoc.pdf. A secondary market may not be available for all options. Transaction costs may be a significant factor in option strategies calling for multiple purchases and sales of options, such as spreads. When purchasing long options an investor may lose their entire investment and when selling uncovered options the risk is potentially unlimited. Supporting documentation for any comparisons, recommendations, statistics, technical data, or other similar information will be supplied upon request.
This material is for the private information of the recipient only. This material is not sponsored, endorsed, sold or promoted by any sponsor or provider of an index referred herein (each, an "Index Provider"). GS does not have any affiliation with or control over the Index Providers or any control over the computation, composition or dissemination of the indices. While GS will obtain information from publicly available sources it believes reliable, it will not independently verify this information. Accordingly, GS shall have no liability, contingent or otherwise, to the user or to third parties, for the quality, accuracy, timeliness, continued availability or completeness of the data nor for any special, indirect, incidental or consequential damages which may be incurred or experienced because of the use of the data made available herein, even if GS has been advised of the possibility of such damages.
Standard & Poor's ® and S&P ® are registered trademarks of The McGraw-Hill Companies, Inc. and S&P GSCI™ is a trademark of The McGraw-Hill Companies, Inc. and have been licensed for use by the Issuer. This Product (the "Product") is not sponsored, endorsed, sold or promoted by S&P and S&P makes no representation, warranty or condition regarding the advisability of investing in the Product.
Notice to Brazilian Investors
Marquee is not meant for the general public in Brazil. The services or products provided by or through Marquee, at any time, may not be offered or sold to the general public in Brazil. You have received a password granting access to Marquee exclusively due to your existing relationship with a GS business located in Brazil. The selection and engagement with any of the offered services or products through Marquee, at any time, will be carried out directly by you. Before acting to implement any chosen service or products, provided by or through Marquee you should consider, at your sole discretion, whether it is suitable for your particular circumstances and, if necessary, seek professional advice. Any steps necessary in order to implement the chosen service or product, including but not limited to remittance of funds, shall be carried out at your discretion. Accordingly, such services and products have not been and will not be publicly issued, placed, distributed, offered or negotiated in the Brazilian capital markets and, as a result, they have not been and will not be registered with the Brazilian Securities and Exchange Commission (Comissão de Valores Mobiliários), nor have they been submitted to the foregoing agency for approval. Documents relating to such services or products, as well as the information contained therein, may not be supplied to the general public in Brazil, as the offering of such services or products is not a public offering in Brazil, nor used in connection with any offer for subscription or sale of securities to the general public in Brazil.
The offer of any securities mentioned in this message may not be made to the general public in Brazil. Accordingly, any such securities have not been nor will they be registered with the Brazilian Securities and Exchange Commission (Comissão de Valores Mobiliários) nor has any offer been submitted to the foregoing agency for approval. Documents relating to the offer, as well as the information contained therein, may not be supplied to the public in Brazil, as the offer is not a public offering of securities in Brazil. These terms will apply on every access to Marquee.
Ouvidoria Goldman Sachs Brasil: 0800 727 5764 e/ou ouvidoriagoldmansachs@gs.com
Horário de funcionamento: segunda-feira à sexta-feira (exceto feriados), das 9hs às 18hs.
Ombudsman Goldman Sachs Brazil: 0800 727 5764 and / or ouvidoriagoldmansachs@gs.com
Available Weekdays (except holidays), from 9 am to 6 pm.
| github_jupyter |
# Implementing the Gradient Descent Algorithm
In this lab, we'll implement the basic functions of the Gradient Descent algorithm to find the boundary in a small dataset. First, we'll start with some functions that will help us plot and visualize the data.
```
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
#Some helper functions for plotting and drawing lines
def plot_points(X, y):
admitted = X[np.argwhere(y==1)]
rejected = X[np.argwhere(y==0)]
plt.scatter([s[0][0] for s in rejected], [s[0][1] for s in rejected], s = 25, color = 'blue', edgecolor = 'k')
plt.scatter([s[0][0] for s in admitted], [s[0][1] for s in admitted], s = 25, color = 'red', edgecolor = 'k')
def display(m, b, color='g--'):
plt.xlim(-0.05,1.05)
plt.ylim(-0.05,1.05)
x = np.arange(-10, 10, 0.1)
plt.plot(x, m*x+b, color)
```
## Reading and plotting the data
```
data = pd.read_csv('data.csv', header=None)
X = np.array(data[[0,1]])
y = np.array(data[2])
plot_points(X,y)
plt.show()
```
## TODO: Implementing the basic functions
Here is your turn to shine. Implement the following formulas, as explained in the text.
- Sigmoid activation function
$$\sigma(x) = \frac{1}{1+e^{-x}}$$
- Output (prediction) formula
$$\hat{y} = \sigma(w_1 x_1 + w_2 x_2 + b)$$
- Error function
$$Error(y, \hat{y}) = - y \log(\hat{y}) - (1-y) \log(1-\hat{y})$$
- The function that updates the weights
$$ w_i \longrightarrow w_i + \alpha (y - \hat{y}) x_i$$
$$ b \longrightarrow b + \alpha (y - \hat{y})$$
```
# Implement the following functions
# Activation (sigmoid) function
def sigmoid(x):
return 1/(1 + np.exp(-x))
# Output (prediction) formula
def output_formula(features, weights, bias):
#print(features.shape)
#print(weights.shape)
z = np.matmul(weights, features.T) + bias
return sigmoid(z)
# Error (log-loss) formula
def error_formula(y, output):
return -y*np.log(output) - (1-y)*np.log(1 - output)
# Gradient descent step
def update_weights(x, y, weights, bias, learnrate):
y_hat = output_formula(x, weights, bias)
weights += learnrate*(y - y_hat)*x
bias += learnrate*(y - y_hat)
return weights, bias
```
## Training function
This function will help us iterate the gradient descent algorithm through all the data, for a number of epochs. It will also plot the data, and some of the boundary lines obtained as we run the algorithm.
```
np.random.seed(44)
epochs = 100
learnrate = 0.01
def train(features, targets, epochs, learnrate, graph_lines=False):
errors = []
n_records, n_features = features.shape
last_loss = None
weights = np.random.normal(scale=1 / n_features**.5, size=n_features)
bias = 0
for e in range(epochs):
del_w = np.zeros(weights.shape)
for x, y in zip(features, targets):
output = output_formula(x, weights, bias)
error = error_formula(y, output)
weights, bias = update_weights(x, y, weights, bias, learnrate)
# Printing out the log-loss error on the training set
out = output_formula(features, weights, bias)
loss = np.mean(error_formula(targets, out))
errors.append(loss)
if e % (epochs / 10) == 0:
print("\n========== Epoch", e,"==========")
if last_loss and last_loss < loss:
print("Train loss: ", loss, " WARNING - Loss Increasing")
else:
print("Train loss: ", loss)
last_loss = loss
predictions = out > 0.5
accuracy = np.mean(predictions == targets)
print("Accuracy: ", accuracy)
if graph_lines and e % (epochs / 100) == 0:
display(-weights[0]/weights[1], -bias/weights[1])
# Plotting the solution boundary
plt.title("Solution boundary")
display(-weights[0]/weights[1], -bias/weights[1], 'black')
# Plotting the data
plot_points(features, targets)
plt.show()
# Plotting the error
plt.title("Error Plot")
plt.xlabel('Number of epochs')
plt.ylabel('Error')
plt.plot(errors)
plt.show()
```
## Time to train the algorithm!
When we run the function, we'll obtain the following:
- 10 updates with the current training loss and accuracy
- A plot of the data and some of the boundary lines obtained. The final one is in black. Notice how the lines get closer and closer to the best fit, as we go through more epochs.
- A plot of the error function. Notice how it decreases as we go through more epochs.
```
train(X, y, epochs, learnrate, True)
```
| github_jupyter |
```
from google.colab import drive
drive.mount('/content/drive')
%cd /content/drive/MyDrive/AGGLIO/github_upload
import numpy as np
from sklearn.model_selection import ShuffleSplit
from sklearn.model_selection import GridSearchCV
from agglio_lib import *
n = 1000
w_radius = 10
dim_list=[10, 20, 30, 40, 50]
val_ind=np.random.randint(n, size=int(0.20*n))
#AGGLIO-GD
l2_agd=[]
l2_agsgd=[]
for d in dim_list:
wAst = np.random.randn(d,1)
X = getData(0, 1, n, d)/np.sqrt(d)
w0 =w_radius*np.random.randn(d,1)/np.sqrt(d)
y = sigmoid(np.matmul(X, wAst))
#AGGLIO-GD
alpha_range = np.linspace(start=1, stop=d*5, num=10).tolist()
B_init_range= np.power(10.0, [-1, -2, -3, -4]).tolist()
B_step_range =np.linspace(start=1.01, stop=2, num=5).tolist()
parameters = dict(alpha = alpha_range, B_init=B_init_range, B_step=B_step_range )
cv = ShuffleSplit( n_splits = 1, test_size = 0.3, random_state = 42 )
grid = GridSearchCV( AG_GD( ), param_grid=parameters, refit = False, cv=cv) # verbose=3
grid.fit( X, y.ravel(), w_init=w0.ravel(), w_star=wAst.ravel())
best = grid.best_params_
print("The best parameters for AGGILIO_GD are %s with a score of %0.2f" % (grid.best_params_, grid.best_score_))
agd = AG_GD(alpha= best["alpha"], B_init=best['B_init'], B_step=best['B_step'] )
agd.fit( X, y.ravel(), w_init = w0.ravel(), w_star = wAst.ravel(), max_iter=600 )
l2_agd.append(agd.distVals[-1])
#AGGLIO-SGD
alpha_range = np.linspace(start=1, stop=d*5, num=5).tolist()
B_init_range= np.power(10.0, [-1, -2, -3, -4]).tolist()
B_step_range =np.linspace(start=1.01, stop=2, num=5).tolist()
parameters = dict(alpha = alpha_range, B_init=B_init_range, B_step=B_step_range )
cv = ShuffleSplit( n_splits = 1, test_size = 0.3, random_state = 42 )
grid = GridSearchCV( AG_SGD( ), param_grid=parameters, refit = False, cv=cv) # verbose=3
grid.fit( X, y.ravel(), w_init=w0.ravel(), w_star=wAst.ravel(), minibatch_size=200)
best = grid.best_params_
print("The best parameters for AGGILIO_SGD are %s with a score of %0.2f" % (grid.best_params_, grid.best_score_))
agsgd = AG_SGD(alpha= best["alpha"], B_init=best['B_init'], B_step=best['B_step'] )
agsgd.fit( X, y.ravel(), w_init = w0.ravel(), w_star = wAst.ravel(), max_iter=600, minibatch_size=200 )
l2_agsgd.append(agsgd.distVals[-1])
import matplotlib.ticker as ticker
import matplotlib.pyplot as plt
y_fmt = ticker.FormatStrFormatter('%2.0e')
fig, ax = plt.subplots()
ax.yaxis.set_major_formatter(y_fmt)
plt.plot(dim_list, l2_agd, label='AGGLIO-GD', color='#1b9e77', linewidth=3)
plt.plot(dim_list, l2_agsgd, label='AGGLIO-SGD', color='#5e3c99', linewidth=3)
plt.legend()
plt.ylabel("$||w^t-w^*||_2$",fontsize=12)
plt.xlabel("dimension",fontsize=12)
plt.grid()
plt.title(r"n=1000" )
plt.savefig('dimension_abliation.pdf', dpi=300, bbox_inches = 'tight')
plt.show()
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.preprocessing import MinMaxScaler
from sklearn.preprocessing import LabelEncoder
from sklearn.naive_bayes import GaussianNB
from sklearn.tree import DecisionTreeClassifier
data = pd.read_csv("healthcare-dataset-stroke-data.csv")
data.head().T
print(f"Data shape: {data.shape}")
# missing values
missing_values = data.isnull().sum()
missing_values
#dropping missing value rows
train_data = data.dropna(axis=0, how="any")
print('Train Data Shape: {}'.format(train_data.shape))
train_data['stroke'].unique()
train_data['smoking_status'].unique()
train_data['ever_married'].unique()
train_data['stroke'].value_counts()
sns.countplot(x=train_data['stroke'])
plt.title('Number of patients affected by stroke',fontsize = 12)
train_data['gender'].value_counts()
train_data.groupby(['gender'])['stroke'].value_counts()
sns.countplot(x=train_data['gender'], hue = train_data['stroke'])
plt.title('Gender Stroke Rate',fontsize = 15)
plt.show()
drop_unknown = train_data[train_data['smoking_status'] == "Unknown"].index
train_data.drop(drop_unknown, inplace = True, axis = 0)
# rows and columns after dropped the unknown values
# from smoking status column
print(f"Data shape after dropped Unknown: {train_data.shape}")
train_data['smoking_status'].value_counts()
train_data.groupby(['smoking_status'])['stroke'].value_counts()
sns.countplot(x = train_data['smoking_status'], hue = train_data['stroke'])
plt.show()
# object datas and numeric data seperation
str_data = train_data.select_dtypes(include=['object'])
int_data = train_data.select_dtypes(include = ['integer','float'])
label = LabelEncoder()
features = str_data.apply(label.fit_transform)
features = features.join(int_data)
features.head()
# dropping 'id' and 'stroke' column to make x_train model
xtrain = features.drop(['stroke','id'], axis = 1)
xtrain.head()
# taking last column for y_train model
ytrain = features['stroke']
ytrain.head()
# Making training and testing models
from sklearn.model_selection import train_test_split
x_train,x_test, y_train, y_test = train_test_split(xtrain,ytrain)
# x_train model as a data frame
x_train.head().T
# GaussianNB model making
model = GaussianNB()
model.fit(x_train, y_train)
test_score = model.score(x_test, y_test)
print("NBtest_score:", test_score)
train_score = model.score(x_train, y_train)
print("NBtrain_score:",train_score)
# Desicion tree model making
desicion_tree_mod = DecisionTreeClassifier(criterion = 'entropy', max_depth = 8)
desicion_tree_mod.fit(xtrain, ytrain)
desicion_tree_test_score = desicion_tree_mod.score(x_test, y_test)
print("Desicion Tree test_score:", desicion_tree_test_score)
desicion_tree_train_score = desicion_tree_mod.score(x_train, y_train)
print("Desicion tree train score: ", desicion_tree_train_score)
y_pred = desicion_tree_mod.predict(x_test)
y_pred
# Support Vector Classifier Algorithm
from sklearn.svm import SVC
svc = SVC(kernel = 'linear', random_state = 42)
svc.fit(x_train, y_train)
y_pred_svc = svc.predict(x_test)
y_pred_svc
svc_test_score = svc.score(x_test, y_test)
print("svc test score: ",svc_test_score)
svc_train_score = svc.score(x_train, y_train)
print("svc train score: ", svc_train_score)
from sklearn.metrics import accuracy_score
accuracy_decision_tree = accuracy_score(y_test, y_pred)
print("Decision tree Accuracy: " + str(accuracy_decision_tree * 100))
sc = MinMaxScaler(feature_range=(0, 1))
dataset_scaled = sc.fit_transform(xtrain.values)
y = ytrain.values
y_pred = desicion_tree_mod.predict(x_test)
y_pred
import joblib as jb
jb.dump(desicion_tree_mod, 'stroke.joblib')
model = jb.load('stroke.joblib')
print(model.predict(sc.transform(np.array([[1.0, 0, 2.0, 1.0, 0, 25.0, 0, 0, 79.20, 38.5]]))))
```
| github_jupyter |
# Generalization vs. OOD detection
In this notebook, we investigate the trade-off between generalization and OOD detection.
We are not interested in so-called OOD generalization. Instead, we consider i.i.d. training locations $x \sim p(x)$, and want to bound the true risk measured via $p(x)$. To provide such generalization bound, we consider PAC-Bayes bounds.
```
from argparse import Namespace
from datetime import datetime
from hypnettorch.utils.sim_utils import setup_environment
import matplotlib.pyplot as plt
import numpy as np
import os
from sklearn.gaussian_process import GaussianProcessRegressor
from sklearn.gaussian_process.kernels import RBF, ExpSineSquared
from scipy import optimize
import sys
from time import time
import torch
# Pretend that notebook is located in base directory of this repo.
curr_dir = os.path.basename(os.path.abspath(os.curdir))
base_dir = os.path.abspath('../..')
if curr_dir == 'pac' and base_dir != sys.path[0]:
sys.path.insert(0, base_dir)
from data.regression_utils import generate_1d_dataset
from utils.plotting import plot_predictive_distributions_1dr
from IPython.display import display, Markdown, Latex
#display(Markdown('*some markdown* $\phi$'))
%matplotlib inline
%load_ext autoreload
%autoreload 2
# Create output directory and other helpers.
config = Namespace()
config.out_dir = './out/pac/run_' + \
datetime.now().strftime('%Y-%m-%d_%H-%M-%S')
config.loglevel_info = False
config.random_seed = 42
config.use_cuda = False # CPU usually faster for simple problems.
config.deterministic_run = True
# Delete file from previous tutorial session:
if os.path.exists(config.out_dir):
shutil.rmtree(config.out_dir)
device, writer, logger = setup_environment(config, logger_name='pac_logger')
```
## Binary KL
```
def bin_kl(q, p):
if q == p:
return 0
if q > 0:
if p == 0:
return np.inf
term1 = q * np.log(q / p)
if q < 1:
if p == 1:
return np.inf
term2 = (1-q) * np.log((1-q)/(1-p))
if q == 0:
return term2
elif q == 1:
return term1
else:
return term1 + term2
```
The inverse binary KL $\text{kl}^{-1}$ is defined as:
$$\text{kl}^{-1}(q, \epsilon) = \max \{ p \in [0, 1]: \text{kl}(q, p) \leq \epsilon \}$$
which according to [Reeb et al.](https://arxiv.org/abs/1810.12263) equals to the unique $p \in [q, 1]$ satisfying $kl(q, p) = \epsilon$.
```
def bin_kl_inv(q, eps):
if q == 1 or np.isinf(eps):
return 1
f = lambda p: bin_kl(q, p) - eps
return optimize.bisect(f, q, 1)
```
Reproducing Fig. 4 in [Reeb et al.](https://arxiv.org/abs/1810.12263).
```
q_vals = np.linspace(0, 1, 20)
eps_vals = np.array([0, 0.1, 0.2, 0.5, 1, 2, 5])
for eps in eps_vals:
p_vals = []
for q in q_vals:
p_vals.append(bin_kl_inv(q, eps))
p_vals = np.array(p_vals)
plt.plot(q_vals, p_vals)
plt.title('Inverse binary KL')
plt.show()
```
## Setup Example GP
```
pd_plot_kwargs = {
'sigma_ll': None, #likelihood_var,
'show_title': False,
'xticks': [0, 5, 10, 15, 20],
'yticks': [-2, 0, 2],
'xlim': [-1, 21],
'ylim': [-3, 3],
'show_legend': False,
'out_dir': config.out_dir,
'ts': 30,
'lw': 15,
'ms': 5,
'figsize': (6, 4),
'show_plot': True,
'vlines': [12.5],
}
data = generate_1d_dataset(task_id=7, num_train=35,
num_val=50, num_test=100, rseed=42)
data.plot_dataset()
X_train_np = data.get_train_inputs()
Y_train_np = data.get_train_outputs()
X_test_np = data.get_test_inputs()
Y_test_np = data.get_test_outputs()
X_val_np = data.get_val_inputs()
Y_val_np = data.get_val_outputs()
# Translate grid to PyTorch tensor.
X_train = torch.from_numpy(X_train_np.astype(np.float32)).to(device)
Y_train = torch.from_numpy(Y_train_np.astype(np.float32)).to(device)
X_test = torch.from_numpy(X_test_np.astype(np.float32)).to(device)
Y_test = torch.from_numpy(Y_test_np.astype(np.float32)).to(device)
X_val = torch.from_numpy(X_val_np.astype(np.float32)).to(device)
Y_val = torch.from_numpy(Y_val_np.astype(np.float32)).to(device)
likelihood_var = .3 # Ground-truth
gp_priors_struct = [
{'type': 'rbf',
'kernel_params': {'length_scale': 1., 'length_scale_bounds': 'fixed'}},
{'type': 'ess',
'kernel_params': {'length_scale': 1.0, 'length_scale_bounds': 'fixed',
'periodicity': 6.28, 'periodicity_bounds': 'fixed'}},
]
for pdict in gp_priors_struct:
if pdict['type'] == 'rbf':
kernel = RBF(**pdict['kernel_params'])
elif pdict['type'] == 'ess':
kernel = ExpSineSquared(**pdict['kernel_params'])
pdict['kernel'] = kernel
gpr = GaussianProcessRegressor(kernel=kernel, alpha=likelihood_var,
random_state=42).fit(X_train_np, Y_train_np)
pdict['gpr_instance'] = gpr
grid_mean, grid_std = gpr.predict(X_test_np, return_std=True)
grid_samples = gpr.sample_y(X_test_np, n_samples=10).squeeze(1)
plot_predictive_distributions_1dr(data, X_test, pd_samples=None,
pd_mean=grid_mean, pd_std=grid_std,
title='GP posterior over functions',
filename='regression1d_gp_%s' % (pdict['type']), **pd_plot_kwargs)
```
## Define loss function and average empirical risk
As a loss function we use:
$$l(y, \hat{y}) = 1 - \exp \big\{ - \frac{(y - \hat{y})^2}{\epsilon} \big\}$$
Furthermore, let $p$ be a posterior over hypothesis. The average empirical risk over randomized estimators is defined as
$$R_\text{emp,avg}(p) = \frac{1}{N} \sum_{n=1}^N \mathbb{E}_{h \sim p} [l(y^{(n)}, h(x^{(n)}))]$$
Similarly, the corresponding empirical Bayes risk is defined as
$$R_\text{emp,Bayes}(p) = \frac{1}{N} \sum_{n=1}^N l(y^{(n)}, \mathbb{E}_{h \sim p} [h(x^{(n)})])$$
```
def loss(y1, y2, eps=1):
return 1 - np.exp(-(y1-y2)**2 / eps)
# Plot loss
y1_vals = np.linspace(-3, 3, 3)
y2_vals = np.linspace(-5, 5, 100)
for y1 in y1_vals:
l_vals = loss(y1, y2_vals)
plt.plot(y2_vals, l_vals, label='y1=%f' % y1)
plt.xlabel('y2')
plt.ylabel('loss')
plt.title('Loss function')
plt.legend()
plt.show()
# We need to estimate the expected value over hypotheses via MC sampling.
num_risk_mc = 100
for pdict in gp_priors_struct:
gpr = pdict['gpr_instance']
train_mean, _ = gpr.predict(X_train_np, return_std=True)
train_samples = gpr.sample_y(X_train_np, n_samples=num_risk_mc).squeeze(1)
loss_bayes = loss(Y_train_np, train_mean)
pdict['r_emp_bayes'] = loss_bayes.mean()
loss_avg = loss(Y_train_np, train_samples).mean(axis=1)
pdict['r_emp_avg'] = loss_avg.mean()
```
## Compute PAC bound
Here, we compute the PAC bound according to theorem 1 in [Reeb et al.](https://arxiv.org/abs/1810.12263). Assume a prior $\pi$ and some $\delta \in (0, 1]$:
$$\text{kl}^{-1} \big( R_\text{emp,avg}(p) , \frac{\text{KL}(p,\pi) + \log \frac{2 \sqrt{N}}{\delta}}{N} \big)$$
Note, that the $\text{KL}(p,\pi)$ between Gaussian process prior and posterior can be computed as the KL between the corresponding multivariate Gaussian distributions determining the distribution over function values at training locations.
```
def kl_mvg(mu1, cov1, mu2, cov2):
"""KL between multivariate Gaussian distributions."""
k = mu1.size
# Check whether covariance matrices are positive definite.
if not np.all(np.linalg.eigvals(cov1) > 0):
display(Markdown('**WARN** Covariance matrix 1 is not positive definite.'))
cov1 += 1e-5 * np.eye(k)
if not np.all(np.linalg.eigvals(cov2) > 0):
display(Markdown('**WARN** Covariance matrix 2 is not positive definite.'))
cov2 += 1e-5 * np.eye(k)
cov1_det = np.linalg.det(cov1)
cov2_det = np.linalg.det(cov2)
with np.errstate(divide='ignore'):
log_cov1_det = np.log(cov1_det)
log_cov2_det = np.log(cov2_det)
# This should not occur if we properky convert the
if np.isnan(log_cov1_det) or np.isinf(log_cov1_det):
display(Markdown('**WARN** Determinant 1 too small for log.'))
log_cov1_det = np.log(cov1_det + 1e-5)
if np.isnan(log_cov2_det) or np.isinf(log_cov2_det):
display(Markdown('**WARN** Determinant 2 too small for log.'))
log_cov2_det = np.log(cov2_det + 1e-5)
#cov1_inv = np.linalg.inv(cov1)
cov2_inv = np.linalg.inv(cov2)
return .5 * (np.trace(cov2_inv @ cov1) + \
((mu2-mu1).T @ cov2_inv @ (mu2-mu1)).squeeze() - \
k + log_cov2_det - log_cov1_det)
delta = .05
betas = [1.5, 1.75, 2., 2.25, 2.5] # Required for bound from Germain et al. Corresponds to Alquier's lambda.
for pdict in gp_priors_struct:
display(Markdown('Kernel: **%s**' % (pdict['type'])))
kernel = pdict['kernel']
gpr = pdict['gpr_instance']
N = X_train_np.shape[0]
prior_mu = np.zeros((N, 1))
prior_cov = kernel(X_train_np, X_train_np)
post_mu, post_cov = gpr.predict(X_train_np, return_cov=True)
kl = kl_mvg(post_mu, post_cov, prior_mu, prior_cov)
assert kl >= 0 # Might fail due to numerical issues.
### Compute PAC-bound according to Eq. 4 in Reeb et al. ###
pac_bound_reeb = bin_kl_inv(pdict['r_emp_avg'], \
(kl + np.log(2*np.sqrt(N)/delta)) / N)
### Compute PAC-bound according to Eq. 1 in Germain et al. ###
# https://arxiv.org/abs/1605.08636
pac_bounds_germain = []
for beta in betas:
pac_bounds_germain.append(1 / (1-np.exp(-beta)) * (1 - np.exp( \
-beta * pdict['r_emp_avg'] - 1/N * (kl + np.log(1/delta)))))
### Compute PAC-bound according to Theorem 2.1 in Alquier. ###
# https://arxiv.org/abs/2110.11216
#pac_bound_alquier = pdict['r_emp_avg'] + beta*1**2 / (8*N) + \
# (kl + np.log(1/delta)) / beta
### Compute MSE on validation set ###
val_mean = gpr.predict(X_val_np)
val_mse = np.mean((val_mean - Y_val_np)**2)
print('Empirical avg. risk: %f' % pdict['r_emp_avg'])
print('Empirical Bayes risk: %f' % pdict['r_emp_bayes'])
print('KL-divergence: %f' % kl)
print('PAC-bound (Reeb et al.) for delta=%f: %f' % (delta, pac_bound_reeb))
for i, beta in enumerate(betas):
print('PAC-bound (Germain et al.) for delta=%f, beta=%f: %f' % \
(delta, beta, pac_bounds_germain[i]))
#print('PAC-bound (Alquier) for delta=%f, lambda=%f: %f' % (delta, beta, pac_bound_alquier))
print('Log-marginal likelihood: %f' % (gpr.log_marginal_likelihood()))
print('MSE on validation set: %f' % (val_mse))
```
## Sanity-Check: KL divergence between GP prior and posterior
Let's reconsider the prior $\pi$ and posterior $p$. According to [Reeb et al.](https://arxiv.org/abs/1810.12263), the [KL in function space](https://arxiv.org/abs/2011.09421) $\text{KL}(p,\pi)$ can (for this special space) be computed as $\text{KL}(p(f_N),\pi(f_N))$, where $f_N$ denotes function values at training locations.
$$\text{KL}(p,\pi) = \text{KL}(p(f_N) p(f \mid f_N),\pi(f_N) \pi(f \mid f_N)) = \text{KL}(p(f_N),\pi(f_N)) + \mathbb{E}_{p(f_N)} [ \text{KL}(p(f \mid f_N),\pi(f \mid f_N)) ]$$
Here, we just show numerically that the last term is actually zero. Note, the KL is non-negative. Thus, if we consistently see that $\text{KL}(p(f \mid f_N),\pi(f \mid f_N)) = 0$ for $f_N \sim p(f_N)$ then we are done.
```
# Use new dataset with less training points due to numerical stability.
data2 = generate_1d_dataset(task_id=7, num_train=5,
num_val=50, num_test=100)
data2.plot_dataset()
X_train_np2 = data2.get_train_inputs()
Y_train_np2 = data2.get_train_outputs()
# Get a random validation batch.
n_val = 10
n_train = data2.num_train_samples
bval_X, bval_Y = data2.next_val_batch(n_val)
# Concat with training set.
X_all = np.concatenate([bval_X, X_train_np2], axis=0)
for pdict in gp_priors_struct:
display(Markdown('Kernel: **%s**' % (pdict['type'])))
if pdict['type'] == 'rbf':
kernel = RBF(**pdict['kernel_params'])
elif pdict['type'] == 'ess':
kernel = ExpSineSquared(**pdict['kernel_params'])
gpr = GaussianProcessRegressor(kernel=kernel, alpha=likelihood_var,
random_state=42).fit(X_train_np2, Y_train_np2)
# Compute joint prior and posterior.
K_all = kernel(X_all)
pmu_all, pcov_all = gpr.predict(X_all, return_cov=True)
# Sample random function values f_N.
f_N = gpr.sample_y(X_train_np2, n_samples=1).squeeze(1)
# Compute conditional distributions (see Sec. 8.1.3 in matrix cook book).
K_val = K_all[:n_val, :n_val]
K_train = K_all[n_val:, n_val:]
K_cross = K_all[:n_val, n_val:]
m_val = pmu_all[:n_val]
m_train = pmu_all[n_val:]
C_val = pcov_all[:n_val, :n_val]
C_train = pcov_all[n_val:, n_val:]
C_cross = pcov_all[:n_val, n_val:]
prior_cond_mu = K_cross @ np.linalg.inv(K_train) @ f_N
post_cond_mu = m_val + C_cross @ np.linalg.inv(C_train) @ (f_N - m_train)
print('Distance between means: %f' % np.sum((prior_cond_mu - post_cond_mu)**2))
prior_cond_cov = K_val - K_cross @ np.linalg.inv(K_train) @ K_cross.T
post_cond_cov = C_val - C_cross @ np.linalg.inv(C_train) @ C_cross.T
kl_cond = kl_mvg(post_cond_mu, post_cond_cov, prior_cond_mu, prior_cond_cov)
print('KL between conditional distributions: %f' % (kl_cond))
```
| github_jupyter |
<a href="https://colab.research.google.com/github/Avery1493/DS-Unit-2-Applied-Modeling/blob/master/module3-permutation-boosting/Quinn_233_LS_DS_233_assignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
Lambda School Data Science
*Unit 2, Sprint 3, Module 3*
---
# Permutation & Boosting
You will use your portfolio project dataset for all assignments this sprint.
## Assignment
Complete these tasks for your project, and document your work.
- [ ] If you haven't completed assignment #1, please do so first.
- [ ] Continue to clean and explore your data. Make exploratory visualizations.
- [ ] Fit a model. Does it beat your baseline?
- [ ] Try xgboost.
- [ ] Get your model's permutation importances.
You should try to complete an initial model today, because the rest of the week, we're making model interpretation visualizations.
But, if you aren't ready to try xgboost and permutation importances with your dataset today, that's okay. You can practice with another dataset instead. You may choose any dataset you've worked with previously.
The data subdirectory includes the Titanic dataset for classification and the NYC apartments dataset for regression. You may want to choose one of these datasets, because example solutions will be available for each.
## Reading
Top recommendations in _**bold italic:**_
#### Permutation Importances
- _**[Kaggle / Dan Becker: Machine Learning Explainability](https://www.kaggle.com/dansbecker/permutation-importance)**_
- [Christoph Molnar: Interpretable Machine Learning](https://christophm.github.io/interpretable-ml-book/feature-importance.html)
#### (Default) Feature Importances
- [Ando Saabas: Selecting good features, Part 3, Random Forests](https://blog.datadive.net/selecting-good-features-part-iii-random-forests/)
- [Terence Parr, et al: Beware Default Random Forest Importances](https://explained.ai/rf-importance/index.html)
#### Gradient Boosting
- [A Gentle Introduction to the Gradient Boosting Algorithm for Machine Learning](https://machinelearningmastery.com/gentle-introduction-gradient-boosting-algorithm-machine-learning/)
- _**[A Kaggle Master Explains Gradient Boosting](http://blog.kaggle.com/2017/01/23/a-kaggle-master-explains-gradient-boosting/)**_
- [_An Introduction to Statistical Learning_](http://www-bcf.usc.edu/~gareth/ISL/ISLR%20Seventh%20Printing.pdf) Chapter 8
- [Gradient Boosting Explained](http://arogozhnikov.github.io/2016/06/24/gradient_boosting_explained.html)
- _**[Boosting](https://www.youtube.com/watch?v=GM3CDQfQ4sw) (2.5 minute video)**_
```
#Imports
import numpy as np
import pandas as pd
pd.options.display.max_columns=200
pd.options.display.max_rows=200
#Loading Data
Q3 = pd.read_csv('https://raw.githubusercontent.com/Avery1493/Lending-Club/master/Data/LC2019Q3_Paid_or_Charged_Off')
Q4 = pd.read_csv('https://raw.githubusercontent.com/Avery1493/Lending-Club/master/Data/LC2019Q4_Paid_or_Charged_Off')
#Combining Data Sets
df = pd.concat([Q3,Q4])
print(df.shape)
#Stripping
df['term'] = df['term'].str.strip(' months')
df['int_rate'] = df['int_rate'].str.strip('%')
df['revol_util'] = df['revol_util'].str.strip('%')
#Combining
df['FICO'] = df[['fico_range_low','fico_range_high']].mean(axis=1)
df['Last_FICO'] = df[['last_fico_range_low','last_fico_range_high']].mean(axis=1)
#installment
#Dropping
df = df.drop(['member_id','url','desc','zip_code', 'fico_range_low',
'fico_range_high','hardship_flag','hardship_type',
'hardship_reason','hardship_status','deferral_term',
'hardship_amount','hardship_start_date','hardship_end_date',
'payment_plan_start_date','hardship_length','hardship_dpd',
'hardship_loan_status','next_pymnt_d',
'orig_projected_additional_accrued_interest',
'hardship_payoff_balance_amount',
'hardship_last_payment_amount','funded_amnt','funded_amnt_inv',
'total_rec_prncp','total_rec_int','total_pymnt_inv','total_pymnt',
'pymnt_plan','recoveries','delinq_amnt','tax_liens',
'pub_rec_bankruptcies','deferral_term','settlement_status',
'settlement_date','settlement_amount','settlement_percentage',
'settlement_term','il_util','bc_util','sec_app_revol_util',
'last_fico_range_low','last_fico_range_high',
'last_credit_pull_d','out_prncp','out_prncp_inv',
'collection_recovery_fee','acc_now_delinq',
'num_tl_120dpd_2m','num_tl_30dpd'], axis = 1)
#Missing
df = df.fillna(np.NaN)
#Datetime
df['issue_d'] = pd.to_datetime(df['issue_d'])
df['earliest_cr_line'] = pd.to_datetime(df['earliest_cr_line'])
print(df.shape)
df.sample(3)
train = df[df['issue_d'].dt.month <= 8]
val = df[(df['issue_d'].dt.month >8) & (df['issue_d'].dt.month <11)]
test = df[df['issue_d'].dt.month >= 11]
train.shape, val.shape, test.shape
```
# Exploritory Visualizations
```
import seaborn as sns
import matplotlib.pyplot as plt
#Separtaing train by class
paid = train[train['loan_status'] == 'Fully Paid']
charged_off = train[train['loan_status'] != 'Fully Paid']
paid.shape, charged_off.shape
paid.describe()
charged_off.describe()
fig, (ax1,ax2) = plt.subplots(1,2, sharey=False)
fig.suptitle('Distribution of Loan Funded Amount')
ax1.hist(paid['loan_amnt'], edgecolor='black',color='blue',
bins=int(20));
ax2.hist(charged_off['loan_amnt'], edgecolor='black',color='orange',
bins=int(20));
mean = train['loan_amnt'].mean()
median = train['loan_amnt'].median()
fig, ax = plt.subplots()
fig.suptitle('Distribution of Loan Funded Amount')
ax.hist(train['loan_amnt'], edgecolor='black',color='blue',
bins=int(20))
ax.vlines(mean,0,1150, color='black')
ax.vlines(median,0,1150, color='red')
plt.show();
data = [paid['loan_amnt'],charged_off['loan_amnt']]
fig, ax = plt.subplots()
ax.set_title('Loan Funded Amount by Class')
plt.boxplot(data,notch=False, vert=True);
sns.violinplot(x=train['loan_status'], y=train['Last_FICO'])
```
# XGBoost
```
target = 'loan_status'
features = train.columns.drop([target, 'id','issue_d','earliest_cr_line'])
X_train = train[features]
y_train = train[target]
X_val = val[features]
y_val = val[target]
features.value_counts().sum()
!pip install category_encoders
!pip install eli5
import category_encoders as ce
from sklearn.impute import SimpleImputer
from sklearn.pipeline import make_pipeline
from sklearn.ensemble import RandomForestClassifier
from xgboost import XGBClassifier
pipeline = make_pipeline(
ce.OrdinalEncoder(),
SimpleImputer(strategy='median'),
XGBClassifier(n_estimaters=100, random_state=42, n_jobs=-1)
)
pipeline.fit(X_train, y_train)
from sklearn.metrics import balanced_accuracy_score
y_pred = pipeline.predict(X_val)
print('Validation accuracy:', balanced_accuracy_score(y_val,y_pred))
```
# Permutation
```
transformers = make_pipeline(
ce.OrdinalEncoder(),
SimpleImputer(strategy='median')
)
X_trained_transformed = transformers.fit_transform(X_train)
X_val_transformed = transformers.fit_transform(X_val)
model = RandomForestClassifier(n_estimators=100,
random_state=42, n_jobs=-1)
model.fit(X_trained_transformed,y_train)
import eli5
from eli5.sklearn import PermutationImportance
#Calculate importances
permuter = PermutationImportance(
model,
scoring = 'balanced_accuracy',
n_iter=5,
random_state=42
)
permuter.fit(X_val_transformed,y_val)
feature_names = X_val.columns.tolist()
pd.Series(permuter.feature_importances_,feature_names).sort_values(ascending=False)
#Display Permutation importances
eli5.show_weights(
permuter,
top=None, #shows all features
feature_names = feature_names
)
```
| github_jupyter |
# Load data
```
import os, glob
import cv2
import numpy as np
import matplotlib.pyplot as plt
from multiprocessing import Pool
import functools
%matplotlib inline
```
## video2npy
```
import skvideo.io
import skvideo.datasets
vid_root = '/data/dataset/UCF/'
vid_ls = glob.glob(vid_root+"v_BabyCrawling**.avi")
vid_ls.sort()
print(len(vid_ls))
print(vid_ls[:5])
videogen = skvideo.io.vreader(vid_ls[55])
for i, frame in enumerate(videogen):
#print(i, frame.shape)
if i == 4:
print(type(frame))
print(frame[:, 40:280, :].shape)
plt.imshow(frame)
def get_frames_npy(vid_path, out_path, length = 7, skip = 2, pre = 1):
if not os.path.exists(out_path):
os.mkdir(out_path)
vid_name = os.path.basename(vid_path).split('.')[0]
videogen = skvideo.io.vreader(vid_path)
frames_lst = [cv2.resize(frame[:, 40:280, :], (256, 256))
for i, frame in enumerate(videogen) if i%skip == 0]
n = 0
for i in range(int(len(frames_lst)/(length*2))):
A = np.asarray(frames_lst[i:i+length])
B = np.asarray(frames_lst[i+length-pre:i+length*2-pre])
v = np.asarray([A,B])
v = np.transpose(v , (0,4,1,2,3))
np.save('{}{}_{}.npy'.format(out_path, vid_name, i),v)
n += 1
return n
```
### 超参数输出
```
npy_path = '/data/dataset/ucf-npy/'
vid_root = '/data/dataset/UCF/'
vid_ls = glob.glob(vid_root+"v_BabyCrawling**.avi")
vid_ls.sort()
total_npy = 0
print('共有视频:{}'.format(len(vid_ls)))
## Hyperparameter
N = 76 # how many videos would be used
length = 7
skip = 2
pre = 1
if N == '': N = len(vid_ls)
if N > len(vid_ls): raise ValueError("处理视频数大于视频总数")
with Pool(12) as p:
p.map(functools.partial(get_frames_npy,
out_path = npy_path, length = length, skip = skip, pre = 1),
vid_ls[:N])
print(len(os.listdir(npy_path)))
```
## 输出检验
```
os.listdir(npy_path)
f_t = npy_path+os.listdir(npy_path)[3]
a= np.load(f_t)
plt.figure(1)
plt.subplot(121)
plt.imshow(np.transpose(a[0], (1,2,3,0))[0])
plt.subplot(122)
plt.imshow(np.transpose(a[1], (1,2,3,0))[0])
rm -rf /data/dataset/ucf-npy/
```
## images2npy
```
def get_one_clip(lst,index ,skip ,length ):
return lst[index:index+length*skip:skip]
def get_clips(img_lst,skip ,length):
clips =[get_one_clip(img_lst,i,skip,length) for i in range(len(img_lst)) if len(get_one_clip(img_lst,i,skip,length)) == length ]
return clips
def get_pair(img_lst,pre ,skip ,length):
A = get_clips(img_lst[:][:-pre],skip ,length)
B = get_clips(img_lst[pre:],skip ,length)
return A,B
read_ = lambda x : cv2.resize(cv2.imread(x)[:, int(1242/2-375/2):int(1242/2+375/2),::-1],(256,256))
def gen_np(c):
# print(c)
img1 = [ read_(i) for i in c[0]]
img2 = [ read_(i) for i in c[1]]
v = np.asarray([img1,img2])
v = np.transpose(v , (0,4,1,2,3))
# v.transpose(0,4,1,2,3)
return v
def dump(img_lst,dirpath='data' ,start =0 ,skip = 2 ,length =7 ,pre =2):
a,b = get_pair(img_lst,pre ,skip ,length)
task = [ (i,j) for i ,j in zip(a,b)]
for i ,j in enumerate(task):
v = gen_np(j)
np.save('{}{}.npy'.format(dirpath,i+start),v)
# return
def f(data_path):
start = 0
img_lst = glob.glob(data_path+"**.png")
img_lst.sort()
dump(img_lst,'video/{}{}'.format(*data_path.split('/')[-3:-1]),start,2 ,7 ,2)
f_lst = glob.glob('/data/dataset/depthdata/vkitti_1.3.1_rgb/**/**/')
with Pool(12) as p:
p.map(f,f_lst)
fi = 'video/000115-deg-left0.npy'
a= np.load(fi)
plt.imshow(np.transpose(a[0],(1,2,3,0))[2])
```
---
| github_jupyter |
```
class IsolationPlayer:
"""Base class for minimax and alphabeta agents -- this class is never
constructed or tested directly.
******************** DO NOT MODIFY THIS CLASS ********************
Parameters
----------
search_depth : int (optional)
A strictly positive integer (i.e., 1, 2, 3,...) for the number of
layers in the game tree to explore for fixed-depth search. (i.e., a
depth of one (1) would only explore the immediate sucessors of the
current state.)
score_fn : callable (optional)
A function to use for heuristic evaluation of game states.
timeout : float (optional)
Time remaining (in milliseconds) when search is aborted. Should be a
positive value large enough to allow the function to return before the
timer expires.
"""
def __init__(self, search_depth=3, timeout=10.):
self.search_depth = search_depth
self.score = score_fn
self.time_left = None
self.TIMER_THRESHOLD = timeout
class MinimaxPlayer(IsolationPlayer):
"""Game-playing agent that chooses a move using depth-limited minimax
search. You must finish and test this player to make sure it properly uses
minimax to return a good move before the search time limit expires.
"""
def get_move(self, game, time_left):
"""Search for the best move from the available legal moves and return a
result before the time limit expires.
************** YOU DO NOT NEED TO MODIFY THIS FUNCTION *************
For fixed-depth search, this function simply wraps the call to the
minimax method, but this method provides a common interface for all
Isolation agents, and you will replace it in the AlphaBetaPlayer with
iterative deepening search.
Parameters
----------
game : `isolation.Board`
An instance of `isolation.Board` encoding the current state of the
game (e.g., player locations and blocked cells).
time_left : callable
A function that returns the number of milliseconds left in the
current turn. Returning with any less than 0 ms remaining forfeits
the game.
Returns
-------
(int, int)
Board coordinates corresponding to a legal move; may return
(-1, -1) if there are no available legal moves.
"""
self.time_left = time_left
# Initialize the best move so that this function returns something
# in case the search fails due to timeout
best_move = (-1, -1)
try:
# The try/except block will automatically catch the exception
# raised when the timer is about to expire.
return self.minimax(game, self.search_depth)
except SearchTimeout:
pass # Handle any actions required after timeout as needed
# Return the best move from the last completed search iteration
return best_move
def terminal_test(game):
g = game.get_legal_moves()
if g == [(-1,-1)]:
return True
else:
return False
def minimax(self, game, depth):
"""Implement depth-limited minimax search algorithm as described in
the lectures.
This should be a modified version of MINIMAX-DECISION in the AIMA text.
https://github.com/aimacode/aima-pseudocode/blob/master/md/Minimax-Decision.md
**********************************************************************
You MAY add additional methods to this class, or define helper
functions to implement the required functionality.
**********************************************************************
Parameters
----------
game : isolation.Board
An instance of the Isolation game `Board` class representing the
current game state
depth : int
Depth is an integer representing the maximum number of plies to
search in the game tree before aborting
Returns
-------
(int, int)
The board coordinates of the best move found in the current search;
(-1, -1) if there are no legal moves
Notes
-----
(1) You MUST use the `self.score()` method for board evaluation
to pass the project tests; you cannot call any other evaluation
function directly.
(2) If you use any helper functions (e.g., as shown in the AIMA
pseudocode) then you must copy the timer check into the top of
each helper function or else your agent will timeout during
testing.
"""
if self.time_left() < self.TIMER_THRESHOLD:
raise SearchTimeout()
# TODO: finish this function!
best_score = float("-inf")
best_move = None
for m in game.get_legal_moves():
v = min_value(game.forecast_move(m), depth - 1)
if v > best_score:
best_score = v
best_move = m
return best_move
def min_value(game,depth):
if self.time_left() < self.TIMER_THRESHOLD:
raise SearchTimeout()
if terminal_test(game):
return 1 # by Assumption 2
while depth <0:
v = float("inf")
for m in game.get_legal_moves():
v = min(v, max_value(game.forecast_move(m),depth-1))
depth -= 1
return v
else: return 0
def max_value(game,depth):
if self.time_left() < self.TIMER_THRESHOLD:
raise SearchTimeout()
if terminal_test(game):
return -1 # by assumption 2
while depth <0:
v = float("-inf")
for m in game.get_legal_moves():
v = max(v, min_value(game.forecast_move(m),depth-1))
depth -= 1
return v
else: return 0
```
| github_jupyter |
# Working with highly imbalanced data
```
import numpy as np
import matplotlib.pyplot as plt
% matplotlib inline
plt.rcParams["figure.dpi"] = 200
np.set_printoptions(precision=3, suppress=True)
import pandas as pd
from sklearn.model_selection import train_test_split, cross_val_score
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import scale, StandardScaler
rng = np.random.RandomState(0)
n_samples_1 = 1000
n_samples_2 = 100
X_syn = np.r_[1.5 * rng.randn(n_samples_1, 2),
0.5 * rng.randn(n_samples_2, 2) + [2, 2]]
y_syn = np.array([0] * (n_samples_1) + [1] * (n_samples_2))
X_syn_train, X_syn_test, y_syn_train, y_syn_test = train_test_split(X_syn, y_syn)
import openml
# mammography dataset https://www.openml.org/d/310
data = openml.datasets.get_dataset(310)
X, y = data.get_data(target=data.default_target_attribute)
X.shape
np.bincount(y)
df = pd.DataFrame(X)
df.hist(bins='auto')
pd.scatter_matrix(df, c=y, alpha=.2);
X_train, X_test, y_train, y_test = train_test_split(X, y, stratify=y, random_state=0)
from sklearn.decomposition import PCA
pca = PCA()
X_train_pca = pca.fit_transform(X_train)
plt.plot(pca.explained_variance_ratio_)
sorting = np.argsort(y_train)
fig, axes = plt.subplots(1, 2)
axes[0].scatter(X_train_pca[:, 0], X_train_pca[:, 1], c=y_train[:], alpha=.3)
axes[1].scatter(X_train_pca[sorting, 0], X_train_pca[sorting, 1], c=y_train[sorting], alpha=.3)
from sklearn.preprocessing import RobustScaler
rs = RobustScaler().fit(X_train)
X_train_scaled = rs.transform(X_train)
pca_scaled = PCA()
X_train_pca_scaled = pca_scaled.fit_transform(X_train_scaled)
plt.plot(pca_scaled.explained_variance_ratio_)
fig, axes = plt.subplots(1, 2)
axes[0].scatter(X_train_pca_scaled[:, 0], X_train_pca_scaled[:, 1], c=y_train[:], alpha=.3)
axes[1].scatter(X_train_pca_scaled[sorting, 0], X_train_pca_scaled[sorting, 1], c=y_train[sorting], alpha=.3)
sorting = np.argsort(y_train)
fig, axes = plt.subplots(1, 2, figsize=(8, 4))
axes[0].scatter(X_train[:, 3], X_train[:, 4], c=y_train[:], alpha=.3)
axes[1].scatter(X_train[sorting, 3], X_train[sorting, 4], c=y_train[sorting], alpha=.3)
axes[0].set_title("Feature 3 vs 4 random order")
axes[1].set_title("Feature 3 vs 4 sorted")
from sklearn.linear_model import LogisticRegression
scores = cross_val_score(LogisticRegression(),
X_train, y_train, cv=10, scoring='roc_auc')
print(scores.mean())
from sklearn.linear_model import LogisticRegressionCV
scores = cross_val_score(LogisticRegressionCV(scoring='roc_auc'), X_train, y_train, cv=10, scoring='roc_auc')
print(scores.mean())
from imblearn.under_sampling import RandomUnderSampler
rus = RandomUnderSampler(replacement=False)
X_train_subsample, y_train_subsample = rus.fit_sample(X_train, y_train)
print(X_train.shape)
print(X_train_subsample.shape)
print(np.bincount(y_train_subsample))
from imblearn.pipeline import make_pipeline as make_imb_pipeline
undersample_pipe = make_imb_pipeline(RandomUnderSampler(), LogisticRegressionCV())
scores = cross_val_score(undersample_pipe, X_train, y_train, cv=10, scoring='roc_auc')
print(np.mean(scores))
from imblearn.over_sampling import RandomOverSampler
ros = RandomOverSampler()
X_train_oversample, y_train_oversample = ros.fit_sample(X_train, y_train)
print(X_train.shape)
print(X_train_oversample.shape)
print(np.bincount(y_train_oversample))
oversample_pipe = make_imb_pipeline(RandomOverSampler(), LogisticRegression())
scores = cross_val_score(oversample_pipe, X_train, y_train, cv=10, scoring='roc_auc')
print(np.mean(scores))
from sklearn.metrics import roc_curve
oversample_pipe.fit(X_train, y_train)
props_oversample = oversample_pipe.predict_proba(X_test)[:, 1]
fpr_over, tpr_over, _ = roc_curve(y_test, props_oversample)
undersample_pipe.fit(X_train, y_train)
props_undersample = undersample_pipe.predict_proba(X_test)[:, 1]
fpr_under, tpr_under, _ = roc_curve(y_test, props_undersample)
lr = LogisticRegression().fit(X_train, y_train)
props_original = lr.predict_proba(X_test)[:, 1]
fpr_org, tpr_org, _ = roc_curve(y_test, props_original)
plt.plot(fpr_org, tpr_org, label="original")
plt.plot(fpr_over, tpr_over, label="oversample")
plt.plot(fpr_under, tpr_under, label="undersample")
plt.legend()
plt.xlabel("FPR")
plt.ylabel("TPR")
from sklearn.ensemble import RandomForestClassifier
scores = cross_val_score(RandomForestClassifier(n_estimators=100),
X_train, y_train, cv=10, scoring='roc_auc')
print(np.mean(scores))
undersample_pipe_rf = make_imb_pipeline(RandomUnderSampler(), RandomForestClassifier())
scores = cross_val_score(undersample_pipe_rf, X_train, y_train, cv=10, scoring='roc_auc')
print(np.mean(scores))
oversample_pipe_rf = make_imb_pipeline(RandomOverSampler(), RandomForestClassifier())
scores = cross_val_score(oversample_pipe_rf, X_train, y_train, cv=10, scoring='roc_auc')
print(np.mean(scores))
from sklearn.metrics import roc_curve
oversample_pipe_rf.fit(X_train, y_train)
props_oversample = oversample_pipe_rf.predict_proba(X_test)[:, 1]
fpr_over, tpr_over, _ = roc_curve(y_test, props_oversample)
undersample_pipe_rf.fit(X_train, y_train)
props_undersample = undersample_pipe_rf.predict_proba(X_test)[:, 1]
fpr_under, tpr_under, _ = roc_curve(y_test, props_undersample)
rf = RandomForestClassifier(n_estimators=100).fit(X_train, y_train)
props_original = rf.predict_proba(X_test)[:, 1]
fpr_org, tpr_org, _ = roc_curve(y_test, props_original)
plt.plot(fpr_org, tpr_org, label="original")
plt.plot(fpr_over, tpr_over, label="oversample")
plt.plot(fpr_under, tpr_under, label="undersample")
plt.legend()
plt.xlabel("FPR")
plt.ylabel("TPR")
plt.title("RF comparison")
```
# Class Weights
```
from sklearn.linear_model import LogisticRegression
scores = cross_val_score(LogisticRegression(class_weight='balanced'),
X_train, y_train, cv=10, scoring='roc_auc')
print(scores.mean())
from sklearn.ensemble import RandomForestClassifier
scores = cross_val_score(RandomForestClassifier(n_estimators=100, class_weight='balanced'),
X_train, y_train, cv=10, scoring='roc_auc')
print(np.mean(scores))
```
# Resampled Ensembles
```
from sklearn.ensemble import VotingClassifier
from sklearn.tree import DecisionTreeClassifier
def make_resample_tree(random_state=0):
tree = make_imb_pipeline(RandomUnderSampler(random_state=random_state, replacement=True),
DecisionTreeClassifier(max_features='auto', random_state=random_state))
return "tree_{}".format(random_state), tree
classifiers = [make_resample_tree(i) for i in range(100)]
resampled_rf = VotingClassifier(classifiers, voting='soft')
scores = cross_val_score(resampled_rf, X_train, y_train, cv=10, scoring='roc_auc')
print(np.mean(scores))
from sklearn.base import clone
def make_resampled_ensemble(estimator, n_estimators=100):
estimators = []
for i in range(n_estimators):
est = clone(estimator)
if hasattr(est, "random_state"):
est.random_state = i
pipe = make_imb_pipeline(RandomUnderSampler(random_state=i, replacement=True),
est)
estimators.append(("est_{}".format(i), pipe))
return VotingClassifier(estimators, voting="soft")
resampled_tree_test = make_resampled_ensemble(DecisionTreeClassifier(max_features='auto'))
scores = cross_val_score(resampled_tree_test, X_train, y_train, cv=10, scoring='roc_auc')
print(np.mean(scores))
resampled_lr = make_resampled_ensemble(LogisticRegression())
scores = cross_val_score(resampled_lr, X_train, y_train, cv=10, scoring='roc_auc')
print(np.mean(scores))
```
# SMOTE
```
from imblearn.over_sampling import SMOTE
smote = SMOTE()
X_train_smote, y_train_smote = smote.fit_sample(X_train, y_train)
print(X_train_smote.shape)
print(np.bincount(y_train_smote))
X_resampled, y_resampled = SMOTE().fit_sample(X_syn, y_syn)
fig, axes = plt.subplots(1, 2)
axes[0].scatter(X_syn[:, 0], X_syn[:, 1], c=plt.cm.Vega10(y_syn), alpha=.3)
axes[1].scatter(X_resampled[:, 0], X_resampled[:, 1], c=plt.cm.Vega10(y_resampled), alpha=.3)
fig, axes = plt.subplots(1, 2, figsize=(8, 4))
sorting = np.argsort(y_train)
axes[0].scatter(X_train[sorting, 3], X_train[sorting, 4], c=y_train[sorting], alpha=.3)
axes[1].scatter(X_train_smote[:, 3], X_train_smote[:, 4], c=y_train_smote, alpha=.3)
from sklearn.utils import shuffle
fig, axes = plt.subplots(1, 2, figsize=(8, 4))
X_smote_sh, y_smote_sh = shuffle(X_train_smote, y_train_smote)
axes[0].scatter(X_train[:, 3], X_train[:, 4], c=y_train, alpha=.3)
axes[1].scatter(X_smote_sh[:, 3], X_smote_sh[:, 4], c=y_smote_sh, alpha=.3)
smote_pipe = make_imb_pipeline(SMOTE(), LogisticRegression())
scores = cross_val_score(smote_pipe, X_train, y_train, cv=10, scoring='roc_auc')
print(np.mean(scores))
smote_pipe_rf = make_imb_pipeline(SMOTE(), RandomForestClassifier(n_estimators=100))
scores = cross_val_score(smote_pipe_rf, X_train, y_train, cv=10, scoring='roc_auc')
print(np.mean(scores))
from sklearn.model_selection import GridSearchCV
param_grid = {'smote__k_neighbors': [3, 5, 7, 9, 11, 15, 31]}
search = GridSearchCV(smote_pipe_rf, param_grid, cv=10, scoring="roc_auc")
search.fit(X_train, y_train)
search.best_score_
results = pd.DataFrame(search.cv_results_)
results.plot("param_smote__k_neighbors", ["mean_test_score", "mean_train_score"])
smote_pipe_rf = make_imb_pipeline(SMOTE(k_neighbors=11), RandomForestClassifier(n_estimators=100))
scores = cross_val_score(smote_pipe_rf, X_train, y_train, cv=10, scoring='roc_auc')
print(np.mean(scores))
from imblearn.over_sampling import SMOTE
smote = SMOTE(k_neighbors=11)
X_train_smote11, y_train_smote11 = smote.fit_sample(X_train, y_train)
fig, axes = plt.subplots(1, 2, figsize=(8, 4))
X_smote_sh11, y_smote_sh11 = shuffle(X_train_smote11, y_train_smote11)
axes[0].scatter(X_smote_sh[:, 3], X_smote_sh[:, 4], c=y_smote_sh, alpha=.3)
axes[1].scatter(X_smote_sh11[:, 3], X_smote_sh11[:, 4], c=y_smote_sh11, alpha=.3)
axes[0].set_title("SMOTE k_neighbors=5")
axes[1].set_title("SMOTE k_neighbors=11")
from imblearn.combine import SMOTEENN, SMOTETomek
smoteenn_pipe_rf = make_imb_pipeline(SMOTEENN(smote=SMOTE(k_neighbors=11)), RandomForestClassifier(n_estimators=100))
scores = cross_val_score(smoteenn_pipe_rf, X_train, y_train, cv=10, scoring='roc_auc')
print(np.mean(scores))
smoteenn = SMOTEENN(smote=SMOTE(k_neighbors=11))
X_train_smoteenn, y_train_smoteenn = smoteenn.fit_sample(X_train, y_train)
fig, axes = plt.subplots(1, 2, figsize=(8, 4))
X_smote_shenn, y_smote_shenn = shuffle(X_train_smoteenn, y_train_smoteenn)
axes[0].scatter(X_smote_sh11[:, 3], X_smote_sh11[:, 4], c=y_smote_sh11, alpha=.3)
axes[1].scatter(X_smote_shenn[:, 3], X_smote_shenn[:, 4], c=y_smote_shenn, alpha=.3)
axes[0].set_title("SMOTE")
axes[1].set_title("SMOTE ENN")
np.bincount(y_train_smote11)
np.bincount(y_train_smoteenn)
from imblearn.under_sampling import CondensedNearestNeighbour
X_resampled, y_resampled = SMOTE().fit_sample(X_syn, y_syn)
X_resampled_enn, y_resampled_enn = SMOTEENN().fit_sample(X_syn, y_syn)
X_resampled_cnn, y_resampled_cnn = make_imb_pipeline(SMOTE(), CondensedNearestNeighbour()).fit_sample(X_syn, y_syn)
fig, axes = plt.subplots(1, 3)
axes[0].scatter(X_resampled[:, 0], X_resampled[:, 1], c=plt.cm.Vega10(y_resampled), alpha=.3)
axes[0].set_title("SMOTE")
axes[1].scatter(X_resampled_enn[:, 0], X_resampled_enn[:, 1], c=plt.cm.Vega10(y_resampled_enn), alpha=.3)
axes[1].set_title("SMOTE ENN")
axes[2].scatter(X_resampled_cnn[:, 0], X_resampled_cnn[:, 1], c=plt.cm.Vega10(y_resampled_cnn), alpha=.3)
axes[2].set_title("SMOTE CNN")
```
| github_jupyter |
# Question 1
What are the top 5 most common valid procedure codes?
How many patients are associated with at least one of those procedures? Please do not use the result values from 1a - provide code that will find the answer without specifying explicitly those code values.
```
# we ingest the csv files
import pandas as pd
from os import listdir, getcwd
from os.path import isfile, join
path = getcwd()
filelist = [f for f in listdir(path) if isfile(join(path, f))]
filelist
import chardet
for file in filelist:
with open(file, 'rb') as f:
result = chardet.detect(f.read(10000))
print(result)
#inspect the datatypes
df_def = pd.read_csv(filelist[5], encoding="us-ascii").convert_dtypes()
df_cpt = pd.read_csv(filelist[3], encoding="us-ascii").convert_dtypes()
df_icd = pd.read_csv(filelist[4], encoding="us-ascii").convert_dtypes()
df_claims = pd.read_csv(filelist[2], encoding="us-ascii").convert_dtypes()
print("procedure types:\n",df_cpt.dtypes)
print("diagnosis types:\n",df_icd.dtypes)
print("claims types:\n",df_claims.dtypes)
#change datatype so it matches column to join
df_cpt['code'] = df_cpt['code'].astype("string")
#join to get valid codes, then count and rank codes
top_procedures= (df_claims.join(df_cpt.set_index("code"), how="inner", on="procedure_code")
.value_counts("procedure_code").head())
top_procedures
```
# Question 2
What are the top 5 most common valid diagnosis codes?
```
import re
#extract codes
df_claims_diag_split = (df_claims["diagnosis_codes"].str.extractall(r"([A-Z]{1}\d{2}\.?\d*)")
.reset_index())
#remove periods
df_claims_diag_split[0] = df_claims_diag_split[0].apply(lambda x: x.replace(".",""))
df_claims_diag_split.rename(columns={'level_0':'sample_row',"match":"entry",0:"code"}, inplace=True)
#join to get valid codes
df_claims_dc = df_claims_diag_split.join(df_icd.set_index("code"), how="inner", on="code").copy()
#count and rank codes
top_diagnosis = df_claims_dc.value_counts("code").head()
top_diagnosis
```
# Question 3
We need to review this file for overall data quality and highlight any potential problems so that they can be discussed with the data provider. Write a series of tests that will identify any errors in the file and provide a list of all errors found. Do not consider any medical details of the treatments, this question does not require any healthcare knowledge.
These quality checks should include, but are not limited to
Sample data matches the table definition
All standardized codes are valid based on given reference material
Date values are logical and chronological trending is consistent
```
#check if procedure code in cpt table (refers to index row not claim id)
valid_proc_code = df_claims["procedure_code"].isin(df_cpt['code'])
#list of not valid
df_claims[~valid_proc_code]
#check if claim_id exists
claim_id_exists = df_claims["claim_id"].notna()
#list of not valid
df_claims[~claim_id_exists]
#check if there diagnostic codes are valid
df_claims_diag_split["is_valid"]= df_claims_diag_split["code"].isin(df_icd["code"])
valid_diag_code = df_claims_diag_split[["sample_row","is_valid"]]
#list of not valid
valid_diag_code[valid_diag_code["is_valid"]==False]
#check if row has valid diagnostic code format
codeformat = r'[A-Z]{1}\d{2}\.?\d*'
has_valid_diag_code_format = (df_claims["diagnosis_codes"].apply(lambda x: bool(re.search(codeformat, str(x))))
.rename("validformat_diagcode"))
#list of not valid
df_claims[~has_valid_diag_code_format]
#check if date is in right format
dateformat = r'^(19|20)\d{2}-(0[1-9]|1[0-2])-(0[1-9]|[12][0-9]|3[01])$'
valid_date_service_format = (df_claims["date_service"].apply(lambda x: bool(re.search(dateformat, str(x))))
.rename("validformat_serviced"))
#list of not valid
df_claims[~valid_date_service_format]
#check if date is in right format
valid_date_received_format = (df_claims["date_received"].apply(lambda x: bool(re.search(dateformat, str(x))))
.rename("validformat_received"))
df_claims[~valid_date_received_format]
#create dates dataframe
import datetime as dt
claims_dates = (df_claims[["date_service","date_received"]].reset_index().rename(columns={"index":"sample_row"})
.join(valid_date_service_format).join(valid_date_received_format))
claims_dates
#check if date_service is before present
claims_valid_date_service = (claims_dates[claims_dates["validformat_serviced"]==True].copy()
.filter(items=("sample_row","date_service","validformat_serviced")))
claims_valid_date_service["date_service"]= pd.to_datetime(claims_valid_date_service['date_service'])
claims_valid_date_service["before_present_s"] = claims_valid_date_service["date_service"] < dt.datetime.now()
#list of not valid
claims_valid_date_service[claims_valid_date_service["before_present_s"] == False]
#check if date_received is before present
claims_valid_date_received = (claims_dates[claims_dates["validformat_received"]==True].copy()
.filter(items=("sample_row","date_received","validformat_received")))
claims_valid_date_received["date_received"]= pd.to_datetime(claims_valid_date_received['date_received'])
claims_valid_date_received["before_present_r"] = claims_valid_date_received["date_received"] < dt.datetime.now()
#list of not valid
claims_valid_date_received[claims_valid_date_received["before_present_r"] == False]
#check if date_service is less or equal to date_received
all_dates_valid = (claims_valid_date_service
.join(claims_valid_date_received.set_index("sample_row"), how="inner",on="sample_row"))
all_dates_valid["valid_date_order"] = all_dates_valid["date_service"] <= all_dates_valid["date_received"]
#list of not valid
all_dates_valid[all_dates_valid["valid_date_order"]==False]
```
| github_jupyter |
```
%matplotlib inline
import numpy as np
import pandas as pd
import math
from scipy import stats
import pickle
from causality.analysis.dataframe import CausalDataFrame
from sklearn.linear_model import LinearRegression
import matplotlib.pyplot as plt
import matplotlib.font_manager as fm
import plotly
import plotly.graph_objs as go
from plotly.offline import download_plotlyjs, init_notebook_mode, plot, iplot
init_notebook_mode(connected=True)
```
Open the data from past notebooks and correct them to only include years that are common between the data structures (>1999).
```
with open('VariableData/money_data.pickle', 'rb') as f:
income_data, housing_data, rent_data = pickle.load(f)
with open('VariableData/demographic_data.pickle', 'rb') as f:
demographic_data = pickle.load(f)
with open('VariableData/endowment.pickle', 'rb') as f:
endowment = pickle.load(f)
with open('VariableData/expander.pickle', 'rb') as f:
expander = pickle.load(f)
endowment = endowment[endowment['FY'] > 1997].reset_index()
endowment.drop('index', axis=1, inplace=True)
demographic_data = demographic_data[demographic_data['year'] > 1999].reset_index()
demographic_data.drop('index', axis=1, inplace=True)
income_data = income_data[income_data['year'] > 1999].reset_index()
income_data.drop('index', axis=1, inplace=True)
housing_data = housing_data[housing_data['year'] > 1999].reset_index()
housing_data.drop('index', axis=1, inplace=True)
rent_data = rent_data[rent_data['year'] > 1999].reset_index()
rent_data.drop('index', axis=1, inplace=True)
```
Define a function to graph (and perform linear regression on) a given set of data.
```
def grapher(x, y, city, title, ytitle, xtitle, filename):
slope, intercept, r_value, p_value, std_err = stats.linregress(x, y)
fit = slope * x + intercept
trace0 = go.Scatter(
x = x,
y = y,
mode = 'markers',
name=city,
marker=go.Marker(color='#D2232A')
)
fit0 = go.Scatter(
x = x,
y = fit,
mode='lines',
marker=go.Marker(color='#AC1D23'),
name='Linear Fit'
)
data = [trace0, fit0]
layout = go.Layout(
title = title,
font = dict(family='Gotham', size=12),
yaxis=dict(
title=ytitle
),
xaxis=dict(
title=xtitle)
)
fig = go.Figure(data=data, layout=layout)
return iplot(fig, filename=filename)
```
Investigate the connection between the endowment's value and the Black population in Cambridge, controlling for rent and housing prices.
```
x = pd.to_numeric(endowment['Value ($B)']).as_matrix()
y = pd.to_numeric(demographic_data['c_black']).as_matrix()
z1 = pd.to_numeric(rent_data['cambridge']).as_matrix()
z2 = pd.to_numeric(housing_data['cambridge']).as_matrix()
X = CausalDataFrame({'x': x, 'y': y, 'z1': z1, 'z2': z2})
plt.rcParams['font.size'] = 10
gotham_black = fm.FontProperties(fname='/Users/hakeemangulu/Library/Fonts/Gotham Black Regular.ttf')
gotham_book = fm.FontProperties(fname='/Users/hakeemangulu/Library/Fonts/Gotham Book Regular.otf')
endow_black = grapher(x, y, "Cambridge", "The Correlation Between Endowment and Black Population", "Black Population of Cambridge", "Endowment ($B)", "endow_black")
causal_endow_black = X.zplot(x='x', y='y', z=['z1', 'z2'], z_types={'z1': 'c', 'z2': 'c'}, kind='line', color="#D2232A")
fig = causal_endow_black.get_figure()
fig.set_size_inches(9, 5.5)
ax = plt.gca()
ax.set_frame_on(False)
ax.get_yaxis().set_visible(False)
ax.legend_.remove()
ax.set_title("The Controlled Correlation Between Endowment (Billions of Dollars) and Black Population", fontproperties=gotham_black, size=10, color="#595959")
ax.set_xlabel("Endowment", fontproperties=gotham_book, fontsize=10, color="#595959")
for tick in ax.get_xticklabels():
tick.set_fontproperties(gotham_book)
tick.set_fontsize(10)
tick.set_color("#595959")
fig.savefig('images/black_endow.svg', format='svg', dpi=1200, bbox_inches='tight')
```
Investigate the connection between the endowment's value and the housing prices in Cambridge, controlling for growth of the population.
```
x = pd.to_numeric(endowment['Value ($B)']).as_matrix()
y = pd.to_numeric(housing_data['cambridge']).as_matrix()
z1 = pd.to_numeric(demographic_data['c_white']).as_matrix()
z2 = pd.to_numeric(demographic_data['c_poc']).as_matrix()
X = CausalDataFrame({'x': x, 'y': y, 'z1': z1, 'z2': z2})
endow_housing = grapher(x, y, "Cambridge", "The Correlation Between Endowment and Housing Prices", "Housing Prices in Cambridge", "Endowment ($B)", "endow_housing")
causal_endow_housing = X.zplot(x='x', y='y', z=['z1', 'z2'], z_types={'z1': 'c', 'z2': 'c'}, kind='line', color="#D2232A")
fig = causal_endow_housing.get_figure()
fig.set_size_inches(9, 5.5)
ax = plt.gca()
ax.set_frame_on(False)
ax.get_yaxis().set_visible(False)
ax.legend_.remove()
ax.set_title("The Controlled Correlation Between Endowment (Billions of Dollars) and Housing Prices", fontproperties=gotham_black, size=10, color="#595959")
ax.set_xlabel("Endowment", fontproperties=gotham_book, fontsize=10, color="#595959")
for tick in ax.get_xticklabels():
tick.set_fontproperties(gotham_book)
tick.set_fontsize(10)
tick.set_color("#595959")
fig.savefig('images/housing_endow.svg', format='svg', dpi=1200, bbox_inches='tight')
```
Investigate the connection between the endowment's value and the rent prices in Cambridge, controlling for growth of the population.
```
x = pd.to_numeric(endowment['Value ($B)']).as_matrix()
y = pd.to_numeric(rent_data['cambridge']).as_matrix()
z1 = pd.to_numeric(demographic_data['c_white']).as_matrix()
z2 = pd.to_numeric(demographic_data['c_poc']).as_matrix()
X = CausalDataFrame({'x': x, 'y': y, 'z1': z1, 'z2': z2})
endow_rent = grapher(x, y, "Cambridge", "The Correlation Between Endowment and Rent", "Rent in Cambridge", "Endowment ($B)", "endow_rent")
causal_endow_rent = X.zplot(x='x', y='y', z=['z1', 'z2'], z_types={'z1': 'c', 'z2': 'c'}, kind='line', title='The Controlled Correlation Between Endowment and Rent')
fig = causal_endow_rent.get_figure()
fig.set_size_inches(9, 5.5)
ax = plt.gca()
ax.set_frame_on(False)
ax.get_yaxis().set_visible(False)
ax.legend_.remove()
ax.set_title("The Controlled Correlation Between Endowment (Billions of Dollars) and Housing Prices", fontproperties=gotham_black, size=10, color="#595959")
ax.set_xlabel("Endowment", fontproperties=gotham_book, fontsize=10, color="#595959")
for tick in ax.get_xticklabels():
tick.set_fontproperties(gotham_book)
tick.set_fontsize(10)
tick.set_color("#595959")
fig.savefig('images/rent_endow.svg', format='svg', dpi=1200, bbox_inches='tight')
```
Investigate the connection between the amount Harvard pays the city of Cambridge per year (PILOT) and the rent prices in Cambridge, controlling for growth of the population.
```
x = pd.to_numeric(expander['Payments to City']).as_matrix()
y = pd.to_numeric(rent_data['cambridge']).as_matrix()
# Remove the last two elements of the other arrays – PILOT data is not sufficient otherwise.
y = y[:-2].copy()
z1 = pd.to_numeric(demographic_data['c_white']).as_matrix()
z1 = z1[:-2].copy()
z2 = pd.to_numeric(demographic_data['c_poc']).as_matrix()
z2 = z2[:-2].copy()
X = CausalDataFrame({'x': x, 'y': y, 'z1': z1, 'z2': z2})
pilot_rent = grapher(x, y, "Cambridge", "The Correlation Between Harvard's PILOT and Rent", "Rent in Cambridge", "PILOT ($)", "pilot_rent")
causal_endow_rent = X.zplot(x='x', y='y', z=['z1', 'z2'], z_types={'z1': 'c', 'z2': 'c'}, kind='line')
```
| github_jupyter |
# Deploy to KFserving

In this introduction we will:
* [Describe the project structure](#Project-Structure)
* [Train some models](#Train-Models)
* [Create Tempo artifacts](#Create-Tempo-Artifacts)
* [Run unit tests](#Unit-Tests)
* [Save python environment for our classifier](#Save-Classifier-Environment)
* [Test Locally on Docker](#Test-Locally-on-Docker)
* [Production on Kubernetes via Tempo](#Production-Option-1-(Deploy-to-Kubernetes-with-Tempo))
* [Prodiuction on Kuebrnetes via GitOps](#Production-Option-2-(Gitops))
## Prerequisites
This notebooks needs to be run in the `tempo-examples` conda environment defined below. Create from project root folder:
```bash
conda env create --name tempo-examples --file conda/tempo-examples.yaml
```
## Project Structure
```
!tree -P "*.py" -I "__init__.py|__pycache__" -L 2
```
## Train Models
* This section is where as a data scientist you do your work of training models and creating artfacts.
* For this example we train sklearn and xgboost classification models for the iris dataset.
```
import os
from tempo.utils import logger
import logging
import numpy as np
logger.setLevel(logging.ERROR)
logging.basicConfig(level=logging.ERROR)
ARTIFACTS_FOLDER = os.getcwd()+"/artifacts"
# %load src/train.py
from typing import Tuple
import joblib
import numpy as np
from sklearn import datasets
from sklearn.linear_model import LogisticRegression
from xgboost import XGBClassifier
SKLearnFolder = "sklearn"
XGBoostFolder = "xgboost"
def load_iris() -> Tuple[np.ndarray, np.ndarray]:
iris = datasets.load_iris()
X = iris.data # we only take the first two features.
y = iris.target
return (X, y)
def train_sklearn(X: np.ndarray, y: np.ndarray, artifacts_folder: str):
logreg = LogisticRegression(C=1e5)
logreg.fit(X, y)
logreg.predict_proba(X[0:1])
with open(f"{artifacts_folder}/{SKLearnFolder}/model.joblib", "wb") as f:
joblib.dump(logreg, f)
def train_xgboost(X: np.ndarray, y: np.ndarray, artifacts_folder: str):
clf = XGBClassifier()
clf.fit(X, y)
clf.save_model(f"{artifacts_folder}/{XGBoostFolder}/model.bst")
from src.data import IrisData
from src.train import train_lr, train_xgb
data = IrisData()
train_lr(ARTIFACTS_FOLDER, data)
train_xgb(ARTIFACTS_FOLDER, data)
```
## Create Tempo Artifacts
```
from src.tempo import get_tempo_artifacts
classifier, sklearn_model, xgboost_model = get_tempo_artifacts(ARTIFACTS_FOLDER)
# %load src/tempo.py
from typing import Tuple
import numpy as np
from src.constants import SKLearnFolder, XGBFolder, SKLearnTag, XGBoostTag
from tempo.serve.metadata import ModelFramework
from tempo.serve.model import Model
from tempo.serve.pipeline import Pipeline, PipelineModels
from tempo.serve.utils import pipeline
def get_tempo_artifacts(artifacts_folder: str) -> Tuple[Pipeline, Model, Model]:
sklearn_model = Model(
name="test-iris-sklearn",
platform=ModelFramework.SKLearn,
local_folder=f"{artifacts_folder}/{SKLearnFolder}",
uri="s3://tempo/basic/sklearn",
description="SKLearn Iris classification model",
)
xgboost_model = Model(
name="test-iris-xgboost",
platform=ModelFramework.XGBoost,
local_folder=f"{artifacts_folder}/{XGBFolder}",
uri="s3://tempo/basic/xgboost",
description="XGBoost Iris classification model",
)
@pipeline(
name="classifier",
uri="s3://tempo/basic/pipeline",
local_folder=f"{artifacts_folder}/classifier",
models=PipelineModels(sklearn=sklearn_model, xgboost=xgboost_model),
description="A pipeline to use either an sklearn or xgboost model for Iris classification",
)
def classifier(payload: np.ndarray) -> Tuple[np.ndarray, str]:
res1 = classifier.models.sklearn(input=payload)
print(res1)
if res1[0] == 1:
return res1, SKLearnTag
else:
return classifier.models.xgboost(input=payload), XGBoostTag
return classifier, sklearn_model, xgboost_model
```
## Unit Tests
* Here we run our unit tests to ensure the orchestration works before running on the actual models.
```
# %load tests/test_deploy.py
import numpy as np
from src.tempo import get_tempo_artifacts
from src.constants import SKLearnTag, XGBoostTag
def test_sklearn_model_used():
classifier, _, _ = get_tempo_artifacts("")
classifier.models.sklearn = lambda input: np.array([[1]])
res, tag = classifier(np.array([[1, 2, 3, 4]]))
assert res[0][0] == 1
assert tag == SKLearnTag
def test_xgboost_model_used():
classifier, _, _ = get_tempo_artifacts("")
classifier.models.sklearn = lambda input: np.array([[0.2]])
classifier.models.xgboost = lambda input: np.array([[0.1]])
res, tag = classifier(np.array([[1, 2, 3, 4]]))
assert res[0][0] == 0.1
assert tag == XGBoostTag
!python -m pytest tests/
```
## Save Classifier Environment
* In preparation for running our models we save the Python environment needed for the orchestration to run as defined by a `conda.yaml` in our project.
```
!cat artifacts/classifier/conda.yaml
from tempo.serve.loader import save
save(classifier)
```
## Test Locally on Docker
* Here we test our models using production images but running locally on Docker. This allows us to ensure the final production deployed model will behave as expected when deployed.
```
from tempo.seldon.docker import SeldonDockerRuntime
docker_runtime = SeldonDockerRuntime()
docker_runtime.deploy(classifier)
docker_runtime.wait_ready(classifier)
classifier(np.array([[1, 2, 3, 4]]))
print(classifier.remote(np.array([[0, 0, 0,0]])))
print(classifier.remote(np.array([[5.964,4.006,2.081,1.031]])))
docker_runtime.undeploy(classifier)
```
## Production Option 1 (Deploy to Kubernetes with Tempo)
* Here we illustrate how to run the final models in "production" on Kubernetes by using Tempo to deploy
### Prerequisites
Create a Kind Kubernetes cluster with Minio and KFserving installed using Ansible from the Tempo project Ansible playbook.
```
ansible-playbook ansible/playbooks/kfserving.yaml
```
```
!kubectl create ns production
!kubectl apply -f k8s/rbac -n production
from tempo.examples.minio import create_minio_rclone
import os
create_minio_rclone(os.getcwd()+"/rclone.conf")
from tempo.serve.loader import upload
upload(sklearn_model)
upload(xgboost_model)
upload(classifier)
from tempo.serve.metadata import RuntimeOptions, KubernetesOptions
runtime_options=RuntimeOptions(
k8s_options=KubernetesOptions(
defaultRuntime="tempo.kfserving.KFServingKubernetesRuntime",
namespace="production",
serviceAccountName="kf-tempo"
)
)
from tempo.kfserving.k8s import KFServingKubernetesRuntime
k8s_runtime = KFServingKubernetesRuntime(runtime_options)
k8s_runtime.deploy(classifier)
k8s_runtime.wait_ready(classifier)
print(classifier.remote(payload=np.array([[0, 0, 0, 0]])))
print(classifier.remote(payload=np.array([[1, 2, 3, 4]])))
```
### Illustrate client using model remotely
With the Kubernetes runtime one can list running models on the Kubernetes cluster and instantiate a RemoteModel to call the Tempo model.
```
models = k8s_runtime.list_models(namespace="production")
print("Name\tDescription")
for model in models:
details = model.get_tempo().model_spec.model_details
print(f"{details.name}\t{details.description}")
models[0].remote(payload=np.array([[1, 2, 3, 4]]))
k8s_runtime.undeploy(classifier)
```
## Production Option 2 (Gitops)
* We create yaml to provide to our DevOps team to deploy to a production cluster
* We add Kustomize patches to modify the base Kubernetes yaml created by Tempo
```
from tempo.kfserving.k8s import KFServingKubernetesRuntime
k8s_runtime = KFServingKubernetesRuntime(runtime_options)
yaml_str = k8s_runtime.to_k8s_yaml(classifier)
with open(os.getcwd()+"/k8s/tempo.yaml","w") as f:
f.write(yaml_str)
!kustomize build k8s
```
| github_jupyter |
# Data Mining, Preparation and Understanding
Today we'll go through Data Mining, Preparation & Understanding which is a really fun one (and important).
In this notebook we'll try out some important libs to understand & also learn how to parse Twitter with some help from `Twint`. All in all we'll go through `pandas`, `twint` and some more - let's start by installing them.
```
%%capture
!pip install twint
!pip install wordcloud
import twint
import pandas as pd
import tqdm
import nltk
nltk.download('stopwords')
```
## Tonights theme: ÅF Pöyry (and perhaps some AFRY)
To be a Data Miner we need something to mine.

In this case it won't be Doge Coin but rather ÅF, ÅF Pöyry & AFRY.
To be honest, it's not the best theme (pretty generic names ones you go ASCII which we'll do to simplify our lifes.
### What is Twint
`Twint` is a really helpful library to scrape Twitter, it uses the search (i.e. not the API) and simplifies the whole process for us as users.
The other way to do this would be to use either the API yourself (time-consuming to learn and also limited in calls) or to use BS4 (Beatiful Soup) which is a great python-lib to scrape websites. But I'd dare say that it is better for static content sites such as Wikipedia, Aftonbladet etc rather than Twitter etc.
This all together led to the choice of `Twint` _even_ though it has a **huge** disadvantage - it does not support UTF8 from what I can find.
### What is pandas
Pandas is a library to parse, understand and work with data. It's really fast using the `DataFrame` they supply.
Using this `DataFrame` we can manipulate the data in different ways. It has all the functions you can imagine from both SQL and Excel, a great tool all in all.
### Bringing it all together
Let's take a look at how we can use this all together!
First a quick look at the Twint config.
```
"""
Twint Config:
Variable Type Description
--------------------------------------------
Retweets (bool) - Display replies to a subject.
Search (string) - Search terms
Store_csv (bool) - Set to True to write as a csv file.
Pandas (bool) - Enable Pandas integration.
Store_pandas (bool) - Save Tweets in a DataFrame (Pandas) file.
Get_replies (bool) - All replies to the tweet.
Lang (string) - Compatible language codes: https://github.com/twintproject/twint/wiki/Langauge-codes (sv, fi & en supported)
Format (string) - Custom terminal output formatting.
Hide_output (bool) - Hide output.
Rest of config: https://github.com/twintproject/twint/wiki/Configuration
"""
c = twint.Config()
c.Query
c.Search = " ÅF "
c.Format = "Username: {username} | Tweet: {tweet}"
c.Pandas = True
c.Store_pandas = True
c.Pandas_clean = True
c.Show_hashtags = True
c.Limit = 10
twint.run.Search(c)
```
**What do we see?**
No Swedish, what so ever. This is not interesting for our usecase as all the tweets are about something else really.
Let's try ÅF Pöyry instead
```
c.Search = "ÅF AFRY Pöyry"
twint.run.Search(c)
```
Looking at this we have a much better result. This really shows the power of Ngrams (bigram).
Let's play around some in the next box trying `@AFkarriar` as keyword and also to include `Replies` and some other fields.
```
c.Replies = True
twint.run.Search(c)
# Play around with params, do whatever!
```
### Results
Ok, so we have tried out a few different things we can use in `Twint`. For me `@AFkarriar` worked out best - **what was your favorite?**
Let's analyze some more.
```
FILENAME = "afpoyry.csv"
c = twint.Config()
c.Query
c.Show_hashtags = True
c.Search = "ÅF"
c.Lang = "sv"
#c.Get_replies = True
c.Store_csv = True
c.Hide_output = True
c.Output = FILENAME
twint.run.Search(c)
data = pd.read_csv(FILENAME)
print(data.shape)
print(data.dtypes)
```
### Cleaning
We can most likely clean some titles from here, just to make it simpler for us
```
data_less = data.filter(["tweet", "username"])
data_less.head()
data_less["tweet"].head()
from wordcloud import WordCloud
from IPython.display import Image
t = '\n'.join([x.tweet for i, x in data_less.iterrows()])
WordCloud().generate(t).to_file('cloud.png')
Image('cloud.png')
```
**Stop Words** - Anyone remember? Let's remove them!
NLTK is a great toolkit for just about everything in NLP, we can find a list of stopwords for most languages here, including Swedish.
```
from nltk.corpus import stopwords
swe_stop = set(stopwords.words('swedish'))
list(swe_stop)[:5]
```
**Stemming** - Anyone remember? Let's do it!
NLTK is _the_ lib to use when you want at least _some_ swedish. But I think I've squeezed all the swedish out of NLTK that I can find right now...
```
from nltk.stem import SnowballStemmer
stemmer = SnowballStemmer("swedish")
stemmer.stem("hoppade")
```
**Cleaning** - Anyone remember? Let's do it!

To have a "better" word cloud we need to reduce the dimensions and keep more important words.
```
%%capture
!pip install regex
from string import punctuation
import regex as re
# bad_words = re.compile("https|http|pic|www|och|med|att|åf|pöyry|läs")
http_re = re.compile("https?.*?(\w+)\.\w+(\/\s)?")
whitespace_re = re.compile("\s+")
punc_set = set(punctuation)
def clean_punct(tweet):
return ''.join([c for c in tweet if c not in punc_set])
def remove_stopwords(tweet):
return " ".join([t for t in tweet.split(" ") if t not in swe_stop])
# Example of cleaning: remove punct, lowercase, https and stemming/lemmatizing
# (we want to reduce the space/dimensions)
def clean_text(tweet):
tweet = tweet.lower()
tweet = ' '.join([word for word in tweet.split() if not word.startswith('pic.')])
tweet = http_re.sub(r'\1', tweet)
tweet = tweet.lower()
tweet = remove_stopwords(clean_punct(tweet)).strip()
tweet = whitespace_re.sub(' ', tweet)
return tweet
clean_text("hej där borta. hur mår du? vem vet.. Jag vet inte. http:/google.com pic.twitterlol")
#data_less["tweet"] = data_less["tweet"].apply(lambda x: clean_text(x))
data_less["tweet"]
from wordcloud import WordCloud
from IPython.display import Image
t = '\n'.join([x.tweet for i, x in data_less.iterrows()])
WordCloud().generate(t).to_file('cloud_clean.png')
Image('cloud_clean.png')
from collections import Counter
def print_most_common(wcount, n=5):
for (name, count) in wcount.most_common(n):
print(f"{name}: {count}")
t_hash = ' '.join([x for x in t.split() if x.startswith("#")])
hash_count = Counter(t_hash.split())
WordCloud().generate(t_hash).to_file('cloud_#.png')
print_most_common(hash_count, 10)
t_at = ' '.join([x for x in t.split() if x.startswith("@")])
at_count = Counter(t_at.split())
WordCloud().generate(t_at).to_file('cloud_@.png')
print_most_common(at_count, 10)
```
### WordClouds!
Let's take a look at what we've got.
```
Image('cloud_clean.png')
Image('cloud_no_stop.png')
Image('cloud_@.png')
Image('cloud_#.png')
```
### What to do?
A big problem with Swedish is that there's very few models which we can do some fun with, and our time is very limited.
Further on we can do the following:
1. Look at Ngram see if we can see common patterns
2. ...
```
"""
1. Try perhaps some type of Ngrams
4. Find different shit
4. Try to find connections
5. Move over to spark (?)
https://towardsdatascience.com/nlp-for-beginners-cleaning-preprocessing-text-data-ae8e306bef0f
https://medium.com/@kennycontreras/natural-language-processing-using-spark-and-pandas-f60a5eb1cfc6
"""
```
### AFRY
Let's create a wordcloud & everything for AFRY. This is for you to implement fully!
```
FILENAME2 = "afry.csv"
c = twint.Config()
c.Query
c.Show_hashtags = True
c.Search = "afry"
c.Lang = "sv"
c.Get_replies = True
c.Store_csv = True
c.Hide_output = True
c.Output = FILENAME2
twint.run.Search(c)
data_afry = pd.read_csv(FILENAME2)
t_afry = '\n'.join([x.tweet for i, x in data_afry.iterrows()])
WordCloud().generate(t_afry).to_file('cloud_afry.png')
Image('cloud_afry.png')
```
### Jonas Sjöstedt (jsjostedt) vs Jimmy Åkesson (jimmieakesson)
Implementation as follows:
1. Get data for both (tip: use `c.Username` or `c.User_id` and don't forget formatting output in terminal if used)
2. Clean data
3. ?? (Perhaps wordclouds etc)
4. TfIdf
5. Join ds & shuffle, train clf
6. Testing!
## Jimmie Åkesson
```
FILENAME = "jimmie2.csv"
c = twint.Config()
c.Query
c.Show_hashtags = True
#c.Search = "ÅF"
c.Username = "jimmieakesson"
#c.Get_replies = True
c.Store_csv = True
c.Output = FILENAME
twint.run.Search(c)
data_jimmie = pd.read_csv(FILENAME)
print(data_jimmie.shape)
data_less_jimmie = data_jimmie.filter(["tweet", "username"])
data_less_jimmie.head()
data_less_jimmie["tweet"] = data_less_jimmie["tweet"].apply(lambda x: clean_text(x))
data_less_jimmie.head()
from wordcloud import WordCloud
from IPython.display import Image
t = '\n'.join([x.tweet for i, x in data_less_jimmie.iterrows()])
WordCloud().generate(t).to_file('cloud_clean_jimmie.png')
Image('cloud_clean_jimmie.png')
```
## Jonas Sjöstedt
```
FILENAME_J = "jonas.csv"
c = twint.Config()
c.Query
c.Show_hashtags = True
#c.Search = "ÅF"
c.Username = "jsjostedt"
#c.Get_replies = True
c.Store_csv = True
c.Hide_output = True
c.Output = FILENAME_J
twint.run.Search(c)
data_jonas = pd.read_csv(FILENAME_J)
print(data_jonas.shape)
data_less_jonas = data_jonas.filter(["tweet", "username"])
data_less_jonas.head()
data_less_jonas["tweet"] = data_less_jonas["tweet"].apply(lambda x: clean_text(x))
data_less_jonas.head()
t = '\n'.join([x.tweet for i, x in data_less_jonas.iterrows()])
WordCloud().generate(t).to_file('cloud_clean_jonas.png')
Image('cloud_clean_jonas.png')
```
# TfIdf
```
from sklearn.feature_extraction.text import TfidfVectorizer
cv=TfidfVectorizer(ngram_range=(1,1))
word_count_vector_jonas = cv.fit_transform(data_less_jonas["tweet"])
feature_names = cv.get_feature_names()
#get tfidf vector for first document
first_document_vector=word_count_vector_jonas[0]
#print the scores
df = pd.DataFrame(first_document_vector.T.todense(), index=feature_names, columns=["tfidf"])
df.sort_values(by=["tfidf"],ascending=False)
word_count_vector_jimmie = cv.fit_transform(data_less_jimmie["tweet"])
feature_names = cv.get_feature_names()
#get tfidf vector for first document
first_document_vector=word_count_vector_jimmie[2]
#print the scores
df = pd.DataFrame(first_document_vector.T.todense(), index=feature_names, columns=["tfidf"])
df.sort_values(by=["tfidf"],ascending=False)
```
# Join dfs & shuffle, train clf
```
print(data_jimmie.shape)
print(data_jonas.shape)
from sklearn.utils import shuffle
tfidf = TfidfVectorizer(ngram_range=(1,2))
data_less_jonas = data_less_jonas.head(2581)
print(data_less_jonas.shape)
combined = pd.concat([data_less_jimmie,data_less_jonas])
combined = shuffle(combined)
print(combined.shape)
combined.head()
from sklearn.model_selection import train_test_split
tweet_tfidf = tfidf.fit_transform(combined["tweet"])
X_train, X_test, y_train, y_test = train_test_split(tweet_tfidf, combined["username"], test_size=0.1, random_state=42)
X_train[:3]
from sklearn.svm import LinearSVC
clf = LinearSVC()
model = clf.fit(X_train, y_train)
from sklearn.metrics import classification_report
y_pred = clf.predict(X_test)
print(classification_report(y_test, y_pred))
```
# Testing!
```
def testClassifier(tweet):
vector = tfidf.transform([clean_text(tweet)])
print(model.predict(vector))
testClassifier("")
testClassifier("Arbetsmarknaden är inte fri svenska kollektivavtal privatisering arbetslösa kommun")
```
# Going forward
I see 4 options:
1. Find stuffs that can help people in the office (@AFRY)
2. Create models for Swedish and perhaps Open Source
3. Make "interesting"/"fun" stuffs (such as applying Text Generation on something like Cards Against Humanity etc)
4. Try something new (perhaps Image Recognition?)
Focusing on Swedish is only possible in 1 & 2.
Some concrete options:
* Explore SparkNLP
* Ask around at AFRY for things to automate
* Apply text-generation with SOTA to generate either something like Cards Against Humanity or some persons Tweet etc.
* Create datasets to create Swedish models on (might need a mech-turk; this will be pretty time-consuming before we see any type of results).
* Something completely different.
```
```
| github_jupyter |
```
%matplotlib inline
from matplotlib import style
style.use('fivethirtyeight')
import matplotlib.pyplot as plt
from datetime import timedelta
import numpy as np
import pandas as pd
import datetime as dt
```
## Reflect Tables into SQLALchemy ORM
```
# Python SQL toolkit and Object Relational Mapper
import sqlalchemy
from sqlalchemy.ext.automap import automap_base
from sqlalchemy.orm import Session
from sqlalchemy import create_engine, func
# create engine to hawaii.sqlite
engine = create_engine("sqlite:///Resources/hawaii.sqlite")
# reflect an existing database into a new model
Base = automap_base()
# reflect the tables
Base.prepare(engine, reflect=True)
# View all of the classes that automap found
Base.classes.keys()
# Save references to each table
Station = Base.classes.station
Measurement = Base.classes.measurement
# Create our session (link) from Python to the DB
session = Session(engine)
last_date = last_date=session.query(Measurement.date).order_by(Measurement.date.desc()).first().date
last_date =dt.datetime.strptime(last_date, "%Y-%m-%d")
first_date = last_date - timedelta(days=365)
```
## Bonus Challenge Assignment: Temperature Analysis II
```
# This function called `calc_temps` will accept start date and end date in the format '%Y-%m-%d'
# and return the minimum, maximum, and average temperatures for that range of dates
def calc_temps(start_date, end_date):
"""TMIN, TAVG, and TMAX for a list of dates.
Args:
start_date (string): A date string in the format %Y-%m-%d
end_date (string): A date string in the format %Y-%m-%d
Returns:
TMIN, TAVE, and TMAX
"""
return session.query(func.min(Measurement.tobs), func.avg(Measurement.tobs), func.max(Measurement.tobs)).\
filter(Measurement.date >= start_date).filter(Measurement.date <= end_date).all()
# For example
print(calc_temps('2012-02-28', '2012-03-05'))
# Use the function `calc_temps` to calculate the tmin, tavg, and tmax
# for a year in the data set
temps = calc_temps(first_date,last_date)
temps
# Plot the results from your previous query as a bar chart.
# Use "Trip Avg Temp" as your Title
# Use the average temperature for bar height (y value)
# Use the peak-to-peak (tmax-tmin) value as the y error bar (yerr)
lower = temps[0][1] - temps[0][0]
upper = temps[0][2] - temps[0][1]
plt.figure(figsize=(3,6))
plt.bar(0, temps[0][1], yerr=[upper-lower], color='salmon', alpha=0.5)
plt.title('Trip Avg Temp')
plt.xticks([])
plt.ylabel('Temp (F)')
plt.ylim()
plt.show()
```
### Daily Rainfall Average
```
# Calculate the total amount of rainfall per weather station for your trip dates using the previous year's
# matching dates.
# Sort this in descending order by precipitation amount and list the station, name, latitude, longitude, and elevation
query_columns = [Station.station, Station.name, Station.latitude,
Station.longitude, Station.elevation, func.sum(Measurement.prcp)]
results = session.query(*query_columns).\
filter(Measurement.station == Station.station).\
filter(Measurement.date >= first_date).\
filter(Measurement.date <= last_date).\
group_by(Station.name).order_by(func.sum(Measurement.prcp).desc()).all()
print(results)
# Use this function to calculate the daily normals
# (i.e. the averages for tmin, tmax, and tavg for all historic data matching a specific month and day)
def daily_normals(date):
"""Daily Normals.
Args:
date (str): A date string in the format '%m-%d'
Returns:
A list of tuples containing the daily normals, tmin, tavg, and tmax
"""
sel = [func.min(Measurement.tobs), func.avg(Measurement.tobs), func.max(Measurement.tobs)]
return session.query(*sel).filter(func.strftime("%m-%d", Measurement.date) == date).all()
# For example
daily_normals("01-01")
# calculate the daily normals for your trip
# push each tuple of calculations into a list called `normals`
# Set the start and end date of the trip
start_date = '2017-08-01'
end_date = '2017-08-07'
# Use the start and end date to create a range of dates
# Strip off the year and save a list of strings in the format %m-%d
dtobj = dt.datetime.strptime(start_date, '%Y-%m-%d')
enddtobj = dt.datetime.strptime(end_date, '%Y-%m-%d')
tripdates = []
normals =[]
while (dtobj <= enddtobj):
tripdates.append(dt.datetime.strftime(dtobj,'%Y-%m-%d'))
datestr = dt.datetime.strftime(dtobj,'%m-%d')
normals.append(list(np.ravel(daily_normals(datestr))))
dtobj = dtobj + dt.timedelta(days = 1)
normals
# Use the `daily_normals` function to calculate the normals for each date string
# and append the results to a list called `normals`.
# Load the previous query results into a Pandas DataFrame and add the `trip_dates` range as the `date` index
thistory = pd.DataFrame(normals, columns=['tmin','tavg','tmax'])
thistory['Date'] = tripdates
thistory = thistory.set_index("Date")
thistory
# Plot the daily normals as an area plot with `stacked=False`
thistory.plot.area(stacked=False)
plt.xticks(rotation=45)
```
## Close Session
| github_jupyter |
```
# ['nigga', 'hate', 'love','ass','hell','better']
# #accuracy over all cross-validation folds: [0.6317343173431734, 0.618450184501845, 0.6140221402214022, 0.622140221402214, 0.6137370753323486]
# mean=0.62 std=0.01
# ['nigga', 'hate', 'love','ass','hell','better','bitch','fuck','dick']
# accuracy over all cross-validation folds: [0.6287822878228783, 0.6199261992619927, 0.6154981549815498, 0.6236162361623616, 0.6159527326440177]
# mean=0.62 std=0.00
# ['nigga', 'hate', 'love','ass','hell','better','bitch','fuck','dick','hey','shit','sexy','awesome']
# accuracy over all cross-validation folds: [0.6501845018450184, 0.6789667896678967, 0.6516605166051661, 0.6494464944649446, 0.6676514032496307]
# mean=0.66 std=0.01
# Having all individual words as features
# accuracy over all cross-validation folds: [0.7011070110701108, 0.6856088560885609, 0.6915129151291513, 0.7166051660516605, 0.7119645494830132]
# mean=0.70 std=0.01
#Same as above but remove all words appearing in less than 1% of all tweets, plus 2 and 3 grams
# accuracy over all cross-validation folds: [0.692250922509225, 0.6789667896678967, 0.6907749077490775, 0.7202952029520295, 0.7171344165435746]
# mean=0.70 std=0.02
#Same as above but remove all words appearing in less than 5% of all tweets, plus 2 and 3 grams
# accuracy over all cross-validation folds: [0.6464944649446495, 0.6686346863468635, 0.6575645756457564, 0.6553505535055351, 0.6506646971935007]
# mean=0.66 std=0.01
#Same as above but remove all words appearing in less than 0.5% of all tweets, plus 2 and 3 grams
# accuracy over all cross-validation folds: [0.6981549815498155, 0.6952029520295203, 0.6929889298892989, 0.7062730627306273, 0.7104874446085672]
# mean=0.70 std=0.01
#if I don`t clean the token which appears time too little, the word table of coefficients for nonhostile will be strange.(even have some japanese)
from collections import Counter
import numpy as np
import pandas as pd
import re
from sklearn.linear_model import LogisticRegression
from sklearn.feature_extraction import DictVectorizer
def load_data(datafile):
"""
Read your data into a single pandas dataframe where
- each row is an instance to be classified
(this could be a tweet, user, or news article, depending on your project)
- there is a column called `label` which stores the class label (e.g., the true
category for this row)
"""
df = pd.read_csv("\\Users\\11977\\elevate-osna-harassment\\osna\\data.csv")[['text', 'hostile']]
df.columns = ['text', 'label']
df['label'] = ['hostile' if i==1 else 'nonhostile' for i in df.label]
# df['directed'] = ['directed' if i==1 else 'nondirected' for i in df.directed]
return df
df = load_data('~/Dropbox/elevate/harassment/training_data/data.csv.gz')
df.label.value_counts()
# def make_features(df):
# vec = DictVectorizer()
# feature_dicts = []
# words_to_track = ['nigga', 'hate', 'love','ass','hell','better','bitch','fuck','dick','hey','shit','sexy','awesome']
# # will get different model for different features.
# #words_to_track = ['you']
# for i,row in df.iterrows():
# features = {}
# token_counts = Counter(re.sub('\W+', ' ', row['text'].lower()).split())
# for w in words_to_track:
# features[w] = token_counts[w]
# feature_dicts.append(features)
# X = vec.fit_transform(feature_dicts)
# return X, vec
#X, vec = make_features(df)
from sklearn.feature_extraction.text import CountVectorizer
vec = CountVectorizer(min_df=0.005,ngram_range=(1,3))
X = vec.fit_transform(t for t in df['text'].values)
X
X.shape
vec.vocabulary_
X[0,5]
X[:,3].sum()
for word, idx in vec.vocabulary_.items():
print('%20s\t%d' % (word, X[:,idx].sum()))
vec.get_feature_names()
y = np.array(df.label)
Counter(y)
y[[0,5,12]]
class_names = set(df.label)
for word, idx in vec.vocabulary_.items():
for class_name in class_names:
class_idx = np.where(y==class_name)[0]
print('%20s\t%20s\t%d' % (word, class_name, X[class_idx, idx].sum()))
clf = LogisticRegression(multi_class='auto')
clf.fit(X, y)
clf.coef_
coef = [-clf.coef_[0], clf.coef_[0]]
print(coef)
clf.classes_
for ci, class_name in enumerate(clf.classes_):
print('coefficients for %s' % class_name)
display(pd.DataFrame([coef[ci]], columns=vec.get_feature_names()))
features = vec.get_feature_names()
for ci, class_name in enumerate(clf.classes_):
print('top features for class %s' % class_name)
for fi in coef[ci].argsort()[::-1][:10]: # descending order.
print('%20s\t%.2f' % (features[fi], coef[ci][fi]))
from sklearn.model_selection import KFold
from sklearn.metrics import accuracy_score
kf = KFold(n_splits=5, shuffle=True, random_state=42)
accuracies = []
for train, test in kf.split(X):
clf.fit(X[train], y[train])
pred = clf.predict(X[test])
accuracies.append(accuracy_score(y[test], pred))
print('accuracy over all cross-validation folds: %s' % str(accuracies))
print('mean=%.2f std=%.2f' % (np.mean(accuracies), np.std(accuracies)))
```
| github_jupyter |
```
import pandas as pd
import numpy as np
from os.path import join as join_path
import os
from PIL import Image
from matplotlib.pyplot import imshow
import matplotlib.pyplot as plt
import time
%matplotlib inline
INPUT_PATH = '../data/'
TRAIN_IMG_PATH = join_path(INPUT_PATH,'raw/train/')
TEST_IMG_PATH = join_path(INPUT_PATH,'raw/test/')
TRAIN_MASKS_PATH = join_path(INPUT_PATH,'raw/train_masks')
TRAIN_MASKS_CSV_PATH = join_path(INPUT_PATH, 'train_masks.csv')
SAMPLE_SUB_CSV_PATH = join_path(INPUT_PATH, 'sample_submission.csv')
OUTPUT_PATH = '../predictions/'
def read_mask_image(car_code, angle_code):
"""
Read image mask, encoding to 0-black 1-white
car_code: code of the car
angle_code: code of the angle
"""
mask_img_path = join_path(TRAIN_MASKS_PATH, car_code + '_' + angle_code + '_mask.gif')
mask_img = np.array(Image.open(mask_img_path))
return mask_img
def show_mask_image(car_code, angle_code):
"""
Show the image mask
"""
mask_img = read_mask_image(car_code, angle_code)
plt.imshow(mask_img, cmap = 'Greys_r')
plt.show()
def rle_encode(mask_image):
"""
Encode the image mask.
mask_image: numpy array, 1 - mask, 0 - background
Returns all pairs of values that contain a start position and a run length
"""
pixels = mask_image.flatten()
pixels[0] = 0
pixels[-1] = 0
runs = np.where(pixels[1:] != pixels[:-1])[0] + 2
runs[1::2] = runs[1::2] - runs[:-1:2]
return runs
def rle_to_string(runs):
return ' '.join(str(x) for x in runs)
def create_submission(encodes):
"""
Create a submission for a given encondes
"""
print('Create submission...')
t = pd.read_csv(SAMPLE_SUB_CSV_PATH)
t['rle_mask'] = encodes
t.to_csv(join_path(OUTPUT_PATH, 'subm_{}.gz'.format(avg_score)), index=False, compression='gzip')
car_code = '00087a6bd4dc'
angle_code = '04'
mask_img_path = join_path(TRAIN_MASKS_PATH, car_code + '_' + angle_code + '_mask.gif')
mask_img = Image.open(mask_img_path)
plt.imshow(mask_img)
resize_img = mask_img.resize((128, 128), Image.ANTIALIAS)
plt.imshow(resize_img)
num = np.array(resize_img).astype(np.uint8)
img_back = Image.fromarray(num)
img_back = img_back.resize((1918,1280), Image.ANTIALIAS)
plt.imshow(img_back)
c = np.array(img_back)
c.shape
img = Image.open(join_path(TRAIN_IMG_PATH, '00087a6bd4dc_04.jpg')).convert('L')
go = np.array(img)
img
go.shape
import torch
import os
from torch.utils.data import Dataset, DataLoader
from torchvision.transforms import ToTensor
from PIL import Image
class CarvanaDataset(Dataset):
def __init__(self, im_dir, mask_dir=None, transforms=None):
```
| github_jupyter |
# Finite Difference Method
This note book illustrates the finite different method for a Boundary Value Problem.
### Example Boudary Value Problem
$$ \frac{d^2 y}{dx^2} = 4y$$
### Boundary Condition
$$ y(0)=1.1752, y(1)=10.0179 $$
```
import numpy as np
import math
import matplotlib.pyplot as plt
import warnings
warnings.filterwarnings("ignore")
class ListTable(list):
""" Overridden list class which takes a 2-dimensional list of
the form [[1,2,3],[4,5,6]], and renders an HTML Table in
IPython Notebook. """
from IPython.core.display import HTML
def _repr_html_(self):
html = ["<table>"]
for row in self:
html.append("<tr>")
for col in row:
html.append("<td>{0}</td>".format(col))
html.append("</tr>")
html.append("</table>")
return ''.join(html)
from IPython.display import HTML
HTML('''<script>
code_show=true;
function code_toggle() {
if (code_show){
$('div.input').hide();
} else {
$('div.input').show();
}
code_show = !code_show
}
$( document ).ready(code_toggle);
</script>
<form action="javascript:code_toggle()"><input type="submit" value="Click here to toggle on/off the raw code."></form>''')
```
## Discrete Axis
The stepsize is defined as
$$h=\frac{b-a}{N}$$
here it is
$$h=\frac{1-0}{10}$$
giving
$$x_i=0+0.1 i$$
for $i=0,1,...10.$
```
## BVP
N=10
h=1/N
x=np.linspace(0,1,N+1)
fig = plt.figure(figsize=(10,4))
plt.plot(x,0*x,'o:',color='red')
plt.xlim((0,1))
plt.xlabel('x',fontsize=16)
plt.title('Illustration of discrete time points for h=%s'%(h),fontsize=32)
plt.show()
```
## The Difference Equation
The gerenal difference equation is
$$ \frac{1}{h^2}\left(y_{i-1}-2y_i+y_{i+1}\right)=4y_i \ \ \ i=1,..,N-1. $$
Rearranging the equation we have the system of N-1 equations
$$i=1: \frac{1}{0.1^2}\color{green}{y_{0}} -\left(\frac{2}{0.1^2}+4\right)y_1 +\frac{1}{0.1^2} y_{2}=0$$
$$i=2: \frac{1}{0.1^2}y_{1} -\left(\frac{2}{0.1^2}+4\right)y_2 +\frac{1}{0.1^2} y_{3}=0$$
$$ ...$$
$$i=8: \frac{1}{0.1^2}y_{7} -\left(\frac{2}{0.1^2}+4\right)y_8 +\frac{1}{0.1^2} y_{9}=0$$
$$i=9: \frac{1}{0.1^2}y_{8} -\left(\frac{2}{0.1^2}+4\right)y_9 +\frac{1}{0.1^2} \color{green}{y_{10}}=0$$
where the green terms are the known boundary conditions.
Rearranging the equation we have the system of 9 equations
$$i=1: -\left(\frac{2}{0.1^2}+4\right)y_1 +\frac{1}{0.1^2} y_{2}=0$$
$$i=2: \frac{1}{0.1^2}y_{1} -\left(\frac{2}{0.1^2}+4\right)y_2 +\frac{1}{0.1^2} y_{3}=-\frac{1}{0.1^2}\color{green}{y_{0}}$$
$$ ...$$
$$i=8: \frac{1}{0.1^2}y_{7} -\left(\frac{2}{0.1^2}+4\right)y_8 +\frac{1}{0.1^2} y_{9}=0$$
$$i=9: \frac{1}{0.1^2}y_{8} -\left(\frac{2}{0.1^2}+4\right)y_9 =0$$
where the green terms are the known boundary conditions.
Putting this into matrix form gives a $9\times 9 $ matrix
$$
A=\left(\begin{array}{ccc ccc ccc}
-204&100&0& 0&0&0& 0&0&0\\
100&-204&100 &0&0&0& 0&0&0\\
0&100&-204& 100&0&0& 0&0&0\\
.&.&.& .&.&.& .&.&.\\
.&.&.& .&.&.& .&.&.\\
0&0&0& 0&0&0& 100&-204&100\\
0&0&0& 0&0&0& 0&100&-204
\end{array}\right)
$$
an unknown vector
$$
\color{red}{\mathbf{y}}=\color{red}{
\left(\begin{array}{c} y_1\\
y_2\\
y_3\\
.\\
.\\
y_8\\
y_9
\end{array}\right)}
$$
```
y=np.zeros((N+1))
# Boundary Condition
y[0]=1.1752
y[N]=10.0179
```
and the known right hand side is a known $9\times 1$ vector with the boundary conditions
$$
\mathbf{b}=\left(\begin{array}{c}-117.52\\
0\\
0\\
.\\
.\\
0\\
-1001.79 \end{array}\right)
$$
$$ A\mathbf{y}=\mathbf{b}$$
The plot below is a graphical representation of the matrix A.
```
b=np.zeros(N-1)
# Boundary Condition
b[0]=-y[0]/(h*h)
b[N-2]=-y[N]/(h*h)
A=np.zeros((N-1,N-1))
# Diagonal
for i in range (0,N-1):
A[i,i]=-(2/(h*h)+4)
for i in range (0,N-2):
A[i+1,i]=1/(h*h)
A[i,i+1]=1/(h*h)
plt.imshow(A)
plt.xlabel('i',fontsize=16)
plt.ylabel('j',fontsize=16)
plt.yticks(np.arange(N-1), np.arange(1,N-0.9,1))
plt.xticks(np.arange(N-1), np.arange(1,N-0.9,1))
clb=plt.colorbar()
clb.set_label('Matrix value')
plt.title('Matrix A',fontsize=32)
plt.tight_layout()
plt.subplots_adjust()
plt.show()
```
## Solving the system
To solve invert the matrix $A$ such that
$$A^{-1}Ay=A^{-1}b$$
$$y=A^{-1}b$$
The plot below shows the graphical representation of $A^{-1}$.
```
invA=np.linalg.inv(A)
plt.imshow(invA)
plt.xlabel('i',fontsize=16)
plt.ylabel('j',fontsize=16)
plt.yticks(np.arange(N-1), np.arange(1,N-0.9,1))
plt.xticks(np.arange(N-1), np.arange(1,N-0.9,1))
clb=plt.colorbar()
clb.set_label('Matrix value')
plt.title(r'Matrix $A^{-1}$',fontsize=32)
plt.tight_layout()
plt.subplots_adjust()
plt.show()
y[1:N]=np.dot(invA,b)
```
## Result
The plot below shows the approximate solution of the Boundary Value Problem (blue v) and the exact solution (black dashed line).
```
fig = plt.figure(figsize=(8,4))
plt.plot(x,y,'v',label='Finite Difference')
plt.plot(x,np.sinh(2*x+1),'k:',label='exact')
plt.xlabel('x')
plt.ylabel('y')
plt.legend(loc='best')
plt.show()
```
| github_jupyter |
### Package installs
If you are using jupyter lab online, all packages will be available. If you are running this on your local computer, you may need to install some packages. Run the cell below if using jupyter lab locally.
```
!pip install numpy
!pip install scipy
!pip install pandas
!pip install scikit-learn
!pip install seaborn
```
### Importing data
To begin, we need to understand the data.
The ribosome genes are available in a .fasta file called 'ribosome_genes.fasta'. You can have a look if you like.
These genes will be imported as classes (RibosomeGene).
Each RibosomeGene object has a name, accession, sequence and length.
You can access these properties using '.' syntax. (see below).
Try to think of each gene as something physical, rather than code.
In real life, each gene has a .length, its organism has a .name, and it has a .sequence. We can write code in this way too.
we will import these into **ribosome_genes**, which is a list of our genes.
```
import warnings
warnings.filterwarnings('ignore')
from utilities import import_16s_sequences
ribosome_genes = import_16s_sequences()
print('{:<20}{:<30}{:<15}{:<10}'.format('gene.accession', 'gene.name', 'gene.length', 'gene.sequence'))
for gene in ribosome_genes:
print('{:<20}{:<30}{:<15}{:<10}'.format(gene.accession, gene.name[:27], gene.length, gene.sequence[:8] + '...'))
```
### SECTION 1: PAIRWISE DISTANCES
To be able to compare organisms via their sequences, we need a way to measure their difference as a distance.
**K-mer distance**<br>
The kmer distance between two sequences is defined here as the total number of k-mers that are unique to either sequence.<br>
eg: If seq1 has 3 unique kmers not found in seq2 (copy number difference also matters), and seq2 has 2 unique kmers, the kmer distance is 5.
```
def create_kmer_dictionary(seq, k):
kmer_dict = {}
num = len(seq) - k + 1
for i in range(num):
kmer = seq[i:i+k]
if kmer not in kmer_dict:
kmer_dict[kmer] = 0
kmer_dict[kmer] += 1
return kmer_dict
create_kmer_dictionary('CCUUCGGG', 2)
def calculate_total_unique_kmers(kmers1, kmers2):
unique_kmers = 0
c1 = c2 = c3 = c4 = c5 = 0
for k in kmers1:
if k not in kmers2:
c1 += kmers1[k]
elif k in kmers1 and k in kmers2:
c2 = c2+ abs(kmers1[k] - kmers2[k])
for k2 in kmers2:
if k2 not in kmers1:
c3 += kmers2[k2]
unique_kmers = c1+c2+c3
return unique_kmers
kmers1 = {'CCUUCGGG':1}
kmers2 = {'CCUUUUUG':2}
calculate_total_unique_kmers(kmers1, kmers2)
def kmer_distance(seq1, seq2, k):
kmers1 = create_kmer_dictionary(seq1,k)
kmers2 = create_kmer_dictionary(seq2,k)
distance = calculate_total_unique_kmers(kmers1, kmers2)
return distance
```
Let's check our function. We can use the first two entries in the 'ribosome_genes' list.
If implemented correctly, the following should return 24
```
distance = kmer_distance(ribosome_genes[1].sequence, ribosome_genes[3].sequence, 8)
print(distance)
```
**smith-waterman alignment**<br>
Another way to compare the similarity of two sequences is through alignment.
The alignment score of two sequences will be high when they are similar, and low when they are distinct.
Keep in mind the matrix must be 1 element larger than the sequence lengths. Consider whether indel scores for the first row and column need to be filled in.
```
import numpy as np
def init_scoregrid(seq1, seq2, indel_score=-4):
rs = len(seq1) +1
cs = len(seq2) +1
scoregrid = np.zeros((rs, cs), np.int)
return scoregrid
```
Let's do a sanity check that the grid has been initialised properly. <br>
The following should print the initialised scoregrid
```
print(init_scoregrid('hello', 'kittycat'))
```
Write a function that calculates the initialised scoregrid. It accepts two sequences, a scoregrid and match/mismatch and indel scores.
```
import itertools
def calculate_scoregrid(seq1, seq2, scoregrid, match_score=1, mismatch_score=-4, indel_score=-4):
for i, j in itertools.product(range(1, scoregrid.shape[0]), range(1, scoregrid.shape[1])):
match = scoregrid[i - 1, j - 1] + (match_score if seq1[i - 1] == seq2[j - 1] else + mismatch_score)
delete = scoregrid[i - 1, j] + indel_score
insert = scoregrid[i, j - 1] + indel_score
scoregrid[i, j] = max(match, delete, insert, 0)
return scoregrid
```
Let's do another sanity check. <br>
The following should print a calculated scoregrid, with the these numbers in the bottom right corner: <br>
2 0 <br>
0 3
```
scoregrid = init_scoregrid('hello', 'helllo')
print(calculate_scoregrid('hello', 'helllo', scoregrid))
def report_alignment_score(scoregrid):
# given a completed scoregrid, return the smith-waterman alignment score.
sw_alignment_score = scoregrid.max()
return sw_alignment_score
```
Final sanity check. Should return 4.
```
scoregrid = init_scoregrid('hello', 'helllo')
calculated_scoregrid = calculate_scoregrid('hello', 'helllo', scoregrid)
print(report_alignment_score(calculated_scoregrid))
```
Ok! now we're ready to put it all together. <br>
Fill in the function below with the three functions you wrote to calculate the alignment score of two sequences
```
def smith_waterman(seq1, seq2):
matrix = init_scoregrid(seq1, seq2, indel_score=-4)
element_scores = calculate_scoregrid(seq1, seq2, scoregrid, match_score=1, mismatch_score=-4, indel_score=-4)
alignment_score = report_alignment_score(scoregrid)
return alignment_score
```
The following should print 4
```
print(smith_waterman('hello', 'helllo'))
```
**pairwise distances**
We have now written two functions which can calculate the distance of two sequences.
We can calculate the k-mer distance, and the smith-waterman alignment score.
lets use these two methods to calculate the pairwise distance of our genes.
```
import numpy as np
def init_distance_matrix(genes):
values=[]
for gene in genes:
s=gene.accession
values.append(s)
values.append(0)
distance_matrix = {}
for gene in ribosome_genes:
key = gene.accession
distance_matrix[key]={values[i]: values[i + 1] for i in range(0, len(values), 2)}
return distance_matrix
```
Let's print the distance matrix to make sure it worked.
```
from utilities import print_distance_matrix
distance_matrix = init_distance_matrix(ribosome_genes)
print_distance_matrix(distance_matrix)
```
Time to fill in the matrix with distances. <br>
Write a function which calculates the pairwise distance of genes using kmer distance.
you will need to call the 'kmer_distance' function you have written above.
```
def calculate_kmer_distance_matrix(genes, matrix, k):
for gene1 in genes:
key1=gene1.accession
for gene2 in genes:
key2=gene2.accession
matrix[key1][key2]=kmer_distance(gene1.sequence,gene2.sequence,k)
return matrix
```
Let's do the same as above, but this time use the 'smith_waterman' alignment distance function you wrote.
```
def calculate_sw_alignment_distance_matrix(genes, matrix):
for gene1 in genes:
key1=gene1.accession
for gene2 in genes:
key2=gene2.accession
matrix[key1][key2]=smith_waterman(gene1.sequence,gene2.sequence)
return matrix
```
Let's test them out. The two cells below will use your calculate_kmer_distance_matrix, and calculate_sw_alignment_distance_matrix functions to add distances to the matrix. <br>
**NOTE:** the smith-waterman distance calculations can take time. Give it a minute.
```
distance_matrix = init_distance_matrix(ribosome_genes)
kmer_distance_matrix = calculate_kmer_distance_matrix(ribosome_genes, distance_matrix, 8)
print('\nkmer distance matrix')
print_distance_matrix(kmer_distance_matrix)
distance_matrix = init_distance_matrix(ribosome_genes)
sw_alignment_distance_matrix = calculate_sw_alignment_distance_matrix(ribosome_genes, distance_matrix)
print('\nsmith waterman alignment score distance matrix')
print_distance_matrix(sw_alignment_distance_matrix)
```
Let's visualise those in a better manner for human eyes. The cell below will plot heatmaps instead of raw numbers.
```
from utilities import heatmap
heatmap(kmer_distance_matrix, sw_alignment_distance_matrix)
```
### SECTION 2: CLUSTERING
From the heatmaps, it seems like there are a few clusters in the data. <br>
First, lets convert the pairwise distances to 2D coordinates.
This is possible using Multidimensional scaling (MDS).
After we have transformed the distance matrix to 2D coordinates, we can plot it to see if any clusters are evident.
```
from utilities import mds_scatterplot, distance_matrix_to_coordinates_MDS
kmer_distances_xy = distance_matrix_to_coordinates_MDS(kmer_distance_matrix)
sw_distances_xy = distance_matrix_to_coordinates_MDS(sw_alignment_distance_matrix)
mds_scatterplot(kmer_distances_xy)
mds_scatterplot(sw_distances_xy)
```
Seems like there is some clustering happening. <br>
Let's use some clustering algorithms to define the clusters.
in this manner, we can have an objective way to talk about the patterns in the data.
Let's implement the k-means algorithm.
```
from utilities import initialise_centroids, average_point, assign_points, plot_kmeans, points_equal, euclidean_distance
def calculate_mean_centroids(data, assignments, k):
centroids = []
for cluster in range(k):
points = [point for point, assignment in zip(data, assignments) if assignment == cluster]
centroids.append(average_point(points))
return centroids
```
Place calculate_mean_centroids() in the kmeans function below to complete kmeans
```
def kmeans(data, k):
centroids=initialise_centroids(data,k)
cluster_assignments=assign_points(centroids,data)
centroids=calculate_mean_centroids(data,cluster_assignments,k)
return centroids, cluster_assignments
```
You can check your implementation using the cell below:
```
centroids, cluster_assignments = kmeans(kmer_distances_xy, 3)
plot_kmeans(kmer_distances_xy, centroids, cluster_assignments, 3)
```
Let's also implement k-medoids while we're at it. <br>
The only difference between k-means and k-medoids is the calculate_mean_centroids() step, which will instead be calculate_median_centroids()
the median can be taken here as the point in the cluster which has smallest cumulative distance to the other points in the cluster
You can use the provided euclidean_distance() function to calculate distances between points
write a function which calculates new centroid locations (using the median)
```
def calculate_median_centroids(data, assignments, k):
centroids = []
for cluster in range(k):
points = [point for point, assignment in zip(data, assignments) if assignment == cluster]
centroids.append(tuple(np.median(np.array(points), axis=0)))
return centroids
```
Place calculate_median_centroids() in the kmedoids function below to complete kmedoids
```
def kmedoids(data, k):
centroids=initialise_centroids(data,k)
cluster_assignments=assign_points(centroids,data)
centroids=calculate_median_centroids(data,cluster_assignments,k)
return centroids, cluster_assignments
```
Here is another check cell, for kmedoids this time:
```
centroids,cluster_assignments = kmedoids(kmer_distances_xy, 3)
plot_kmeans(kmer_distances_xy, centroids, cluster_assignments, 3)
```
| github_jupyter |
# EFP Fanfic Metadata Scraper
This notebook scrapes metadata from the Italian fanfic site [EFP Fanfic](https://efpfanfic.net/). To make it work, put the URL for a particular fandom page (everything up to `pagina=`) in as the *ScraperStem* value below, and set the range to be (1,some-number), where some-number is the final page of the paginated results for that fandom.
```
#Import libraries
import requests
import xml.etree.ElementTree
import csv
import urllib.parse
from random import randint
import time
from time import sleep
import re
import regex
from bs4 import BeautifulSoup
import pandas as pd
#Create a Pandas dataframe with the metadata fields
italianfanfic = pd.DataFrame(columns=["Title", "Storylink", "Color", "LastChap", "AuthName", "AuthID", "Published", "Updated", "Genre", "Chapters", "Status", "Shiptype", "Note", "Warning", "Characters", "Ships", "Contests", "Reviews", "Blurb"])
```
Here's the cell you should modify with the fandom base URL and the range.
```
headers = {'user-agent': 'Mozilla/5.0 (Macintosh; Intel Mac OS X x.y; rv:10.0) Gecko/20100101 Firefox/10.0 Chrome/74.0.3729.131'}
#URL base, which is the page for a particular fandom, up to the place that indicates the page
ScraperStem = "https://efpfanfic.net/categories.php?catid=47&parentcatid=47&offset=15&pagina="
#For each page in a particular range
#(We know the total range by looking at the first page for the fandom, and seeing what # the last page of results is)
for i in range(1172,3766):
#The full URL combines the base with the page number
ScraperURL = ScraperStem + str(i)
#Print the page
print(ScraperURL)
#Retrieve the page
page = requests.get(ScraperURL, headers=headers)
c = page.content
#Parse the page contents with Beautiful Soup
soup = BeautifulSoup(c, 'html.parser')
#Identify the container with the fics
fics = soup.find_all("div", {"class": "storybloc"})
#For each fic
for fic in fics:
#Find the div with the title
title = fic.find('div', {'class': 'titlestoria'})
#Get the link around the title
storylink = title.a['href']
#Get the title text
titlename = title.text
#Find the title bar that has the color/rating
titlebar = fic.find('div', {'class': 'titlebloc'})
#Get the color/rating (ID value of the div below)
color = titlebar.find('div')
color = color.get('id')
#Look for the div that would indicate it's the last chapter
lastchap = titlebar.find('div', {'class': 'ultimochap'})
#If the last chapter div is not empty
if lastchap.text is not None:
#Then give lastchap a value
lastchap = 'lastchapter'
#Otherwise
else:
#Last chapter should be blank
lastchap = ''
#Find the blurb
blurb = fic.find('div', {'class': 'introbloc'}).text
#Find the metadata div
metadata = fic.find('div', {'class', 'notebloc'})
#If metadata isn't empty
if metadata is not None:
#Author ID has uid = [some number]
authid = re.findall(r'uid=([0-9]*)', str(metadata))
#Author name is the text inside the author ID link
authname = metadata.find('a').text
#Publication date comes after 'Pubblicata:'
published = re.search(r'Pubblicata: ((\d\/*)+) ', str(metadata)).group(1)
#Updated date comes after 'Aggiornata'
updated = re.search(r'Aggiornata: ((\d\/*)+) ', str(metadata)).group(1)
#Genre comes after 'Genere'
genre = re.search(r'Genere: (.*?\|)', str(metadata))
#If genre isn't empty
if genre is not None:
#Capture genre value from regex
genre = genre.group(1)
#Chapters come after 'Capitoli'
chapters = re.search(r'Capitoli: (.*?\<)', str(metadata))
#If chapters are not empty
if chapters is not None:
#Capture value of chapters
chapters = chapters.group(1)
chapters = re.search(r'(\d*)', chapters).group(0)
#Capture the text after 'Capitoli' which should also include the status
status = re.search(r'Capitoli: (.*?\<)', str(metadata))
#Refine the text to capture the actual status
if status is not None:
status = status.group(1)
status = re.search(r'(\| )(.*)(<)$', status).group(2)
#Ship type comes after 'Tipo di coppia'
shiptype = re.search(r'Tipo di coppia: ((.*?)\|)', str(metadata))
#Capture value
if shiptype is not None:
shiptype = shiptype.group(2)
#Note comes after text 'Note'
note = re.search(r'Note: ((.*?)\|)', str(metadata))
#Capture note value
if note is not None:
note = note.group(2)
#Text warning comes after 'Avvertimenti'
textwarning = re.search(r'Avvertimenti: (.*)', str(metadata))
#Capture text warning value
if textwarning is not None:
textwarning = textwarning.group(1)
else:
#Or otherwise set it to blank
textwarning =''
#Characters come after Personaggi'
characters = re.search(r'Personaggi: (.*)', str(metadata))
#Capture character value
if characters is not None:
characters = characters.group(1)
#Ships come after 'Coppie'
ships = re.search(r'Coppie: (.*)', str(metadata))
#Capture ship value
if ships is not None:
ships = ships.group(1)
#Contest info comes after 'Contesto'
contest = re.search(r'Contesto: ((.*?)\|)', str(metadata))
#Capture contest info
if contest is not None:
contest = contest.group(2)
#Reviews comes at the end before 'recension'
reviews = re.search(r'>(\d+)</a> recension', str(metadata))
#Capture reviews value
if reviews is not None:
reviews = reviews.group(1)
#Create new item with the data that's been scraped
newitem = {"Title": title, "Storylink": storylink, "Color": color, "LastChap": lastchap, "AuthName": authname, "AuthID": authid, "Published": published, "Updated": updated, "Genre": genre, "Chapters": chapters, "Status": status, "Shiptype": shiptype, "Note": note, "Warning": textwarning, "Characters": characters, "Ships": ships, "Contests": contest, "Reviews": reviews, "Blurb": blurb}
#Append new item to the Pandas dataframe
italianfanfic = italianfanfic.append(newitem, ignore_index=True)
#Sleep 4-10 seconds before going to the next page
sleep(randint(4,10))
#Display the data
italianfanfic
#Remove newlines and tabs, then display the data again
cleanitalianfanfic = italianfanfic.replace(to_replace=[r"\\t|\\n|\\r", "\t|\n|\r"], value=[" "," "], regex=True, inplace=False)
cleanitalianfanfic
#Save the results to a TSV file
cleanitalianfanfic.to_csv('/Users/qad/Documents/italianfanfic2021-2.tsv', index=False, sep="\t")
```
| github_jupyter |
ERROR: type should be string, got "https://github.com/wwrechard/pydlm\n\nhttps://pydlm.github.io/\n\nhttp://ftp.stat.duke.edu/WorkingPapers/11-19.pdf\n\nDynamic Linear Models (DLMs) or state space models define a very general class of non-stationary time series\nmodels. DLMs may include terms to model trends, seasonality, covariates and autoregressive components. The main goals are short-term forecasting, intervention\nanalysis and monitoring.\n\n```\nimport matplotlib.pyplot as plt\n%matplotlib inline\nimport numpy as np\nimport pydlm.plot.dlmPlot as dlmPlot\nfrom pydlm import dlm, trend, seasonality, dynamic, autoReg, longSeason, modelTuner\nfrom pydlm import dynamic\n```\n\nThis package implementes the Bayesian dynamic linear model (Harrison and West, 1999) for time series data analysis. Modeling and fitting is simple and easy with pydlm. Complex models can be constructed via simple operations:\n\n```\ndata = [0] * 100 + [3] * 100\n# construct the base\nmyDLM = dlm(data)\n# adding model components\n# add a first-order trend (linear trending) with prior covariance 1.0\nmyDLM = myDLM + trend(1, name='lineTrend', w=1.0)\n# add a 7 day seasonality with prior covariance 1.0\nmyDLM = myDLM + seasonality(7, name='7day', w=1.0)\n# add a 3 step auto regression\nmyDLM = myDLM + autoReg(degree=3, data=data, name='ar3', w=1.0)\n# show the added components\nmyDLM.ls()\n# delete unwanted component\nmyDLM.delete('7day')\nmyDLM.ls()\n```\n\nUsers can then analyze the data with the constructed model:\n\n```\n# fit forward filter\nmyDLM.fitForwardFilter()\n# fit backward smoother\nmyDLM.fitBackwardSmoother()\n# plot the results\nmyDLM.plot()\n# plot only the filtered results\nmyDLM.turnOff('smoothed plot')\nmyDLM.plot()\n# plot in one figure\nmyDLM.turnOff('multiple plots')\nmyDLM.plot()\n```\n\nUser can also plot the mean of a component (the time series value that attributed to this component):\n\n```\n# plot the component mean of 'ar3'\nmyDLM.turnOn('smoothed plot')\nmyDLM.turnOff('predict')\nmyDLM.plot(name='ar3')\n```\n\nand also the latent states for a given component:\n\n```\n# plot the latent states of the 'ar3'\nmyDLM.plotCoef(name='ar3')\n```\n\nIt also includes the discounting factor, which can be used to control how rapidly the model should adapt to the new data:\n\n```\ndata = [0] * 100 + [3] * 100\nmyDLM = dlm(data) + trend(1, discount=1.0, w=1.0)\nmyDLM.fit()\nmyDLM.plot()\nmyDLM.delete('trend')\nmyDLM = myDLM + trend(1, discount=0.8, w=1.0)\nmyDLM.fit()\nmyDLM.plot()\n```\n\nThe discounting factor can be auto-tuned by the modelTuner provided by the package:\n\n```\nmyTuner = modelTuner(method='gradient_descent', loss='mse')\ntunedDLM = myTuner.tune(myDLM, maxit=100)\ntunedDLM.fit()\ntunedDLM.plot()\n```\n\n## Real data example\n\nhttps://pydlm.github.io/example1.html\n\n```\nls\ndata_file = open(\"data.csv\", 'r')\n```\n\nPlot the raw data, which is the weekly counts of initial claims for unemployment during 2004 - 2012. We can see strong annual pattern and some local trend from the data.\n\n```\nvariables = data_file.readline().strip().split(',')\ndata_map = {}\nfor var in variables:\n data_map[var] = []\n\nfor line in data_file:\n for i, data_piece in enumerate(line.strip().split(',')):\n data_map[variables[i]].append(float(data_piece))\n\n# Extract and store the data.\ntime_series = data_map[variables[0]]\nfeatures = [[data_map[variables[j]][i] for j in range(1, len(variables)) ]\n for i in range(len(time_series))]\n\n# Plot the raw data\n\ndlmPlot.plotData(range(len(time_series)),\n time_series,\n showDataPoint=False,\n label='raw_data')\nplt.legend(loc='best', shadow=True)\nplt.show()\n```\n\nwe first build a simple model with only local linear trend and seasonality component:\n\n```\n# Build a simple model\n\n# A linear trend\nlinear_trend = trend(degree=1, discount=0.95, name='linear_trend', w=10)\n# A seasonality\nseasonal52 = seasonality(period=52, discount=0.99, name='seasonal52', w=10)\n\nsimple_dlm = dlm(time_series) + linear_trend + seasonal52\nsimple_dlm.fit()\n\n# Plot the fitted results\nsimple_dlm.turnOff('data points')\nsimple_dlm.plot()\n# Plot each com\n```\n\nThe blue curve is the forward filtering result, the green curve is the one-day ahead prediction and the red curve is the backward smoothed result. The light-colored ribbon around the curve is the confidence interval (you might need to zoom-in to see it). The one-day ahead prediction shows this simple model captures the time series somewhat good but loses accuracy around the peak crisis at Week 280 (which is between year 2008 - 2009). The one-day-ahead mean squared prediction error is 0.173 which can be obtaied by calling:\n\nWe can decompose the time series into each of its components:\n\n```\n# Plot each component (attribution)\nsimple_dlm.turnOff('predict plot')\nsimple_dlm.turnOff('filtered plot')\nsimple_dlm.plot('linear_trend')\nsimple_dlm.plot('seasonal52')\n```\n\nMost of the time series shape is attributed to the local linear trend and the strong seasonality pattern is easily seen. To further verify the performance, we use this simple model for long-term forecasting. In particular, we use the previous 351 week ‘s data to forecast the next 200 weeks and the previous 251 week ‘s data to forecast the next 200 weeks. We lay the predicted results on top of the real data:\n\n```\n# Plot the prediction give the first 350 weeks and forcast the next 200 weeks.\nsimple_dlm.plotPredictN(N=200, date=350)\n# Plot the prediction give the first 250 weeks and forcast the next 200 weeks.\nsimple_dlm.plotPredictN(N=200, date=250)\n\n# Build a dynamic regression model\nregressor10 = dynamic(features=features, discount=1.0, name='regressor10', w=10)\ndrm = dlm(time_series) + linear_trend + seasonal52 + regressor10\ndrm.fit()\n\n# Plot the fitted results\ndrm.turnOff('data points')\ndrm.plot()\n# Plot each component (attribution)\ndrm.turnOff('predict plot')\ndrm.turnOff('filtered plot')\ndrm.plot('linear_trend')\ndrm.plot('seasonal52')\ndrm.plot('regressor10')\n# Plot the prediction give the first 300 weeks and forcast the next 150 weeks.\ndrm.plotPredictN(N=150, date=300)\n# Plot the prediction give the first 250 weeks and forcast the next 200 weeks.\ndrm.plotPredictN(N=200, date=250)\n```\n\n" | github_jupyter |
```
import pandas as pd
import numpy as np
df_diabetes = pd.read_csv('diabetes.csv')
print(df_diabetes.shape)
df_diabetes.describe().transpose()
target_column = ['Diabetes']
predictors = list(set(list(df_diabetes.columns))-set(target_column))
print(predictors)
from sklearn.preprocessing import StandardScaler
transformer = StandardScaler()
df_std = df_diabetes.copy()
df_std[predictors] = transformer.fit_transform(df_diabetes[predictors])
df_std.describe().transpose()
from sklearn.model_selection import train_test_split
X_std = df_std[predictors].values
y_std = df_std[target_column].values
X_train_std, X_test_std, y_train_std, y_test_std = train_test_split(X_std, y_std, test_size=0.30, random_state=12)
print(X_train_std.shape)
print(X_test_std.shape)
#Específico do Keras
from tensorflow import keras
from tensorflow.keras import Sequential
from tensorflow.keras.layers import Dense
from tensorflow.keras.utils import to_categorical
#Métricas
from sklearn.metrics import classification_report,confusion_matrix
import matplotlib.pyplot as plt
from sklearn.metrics import confusion_matrix
%matplotlib inline
# one hot encode outputs
y_train_std_cat = to_categorical(y_train_std)
y_test_std_cat = to_categorical(y_test_std)
mlp_std = Sequential()
mlp_std.add(Dense(500, activation='relu', input_dim=8))
mlp_std.add(Dense(100, activation='relu'))
mlp_std.add(Dense(50, activation='relu'))
mlp_std.add(Dense(2, activation='softmax'))
mlp_std.compile(optimizer='adam',
loss='categorical_crossentropy',
metrics=['accuracy'])
mlp_std.fit(X_train_std, y_train_std_cat, epochs=20, verbose=1)
predict_test_std = mlp_std.predict_classes(X_test_std)
print()
print(classification_report(y_test_std,predict_test_std))
print()
cnf_mtx = confusion_matrix(y_test_std,predict_test_std)
fig = plt.figure()
ax = fig.add_subplot(111)
cax = ax.matshow(cnf_mtx, cmap=plt.cm.Blues)
plt.title('Confusion matrix of the classifier')
fig.colorbar(cax)
for (i, j), z in np.ndenumerate(cnf_mtx):
ax.text(j, i, '{:0.1f}'.format(z), ha='center', va='center', bbox=dict(facecolor='white'))
ax.set_xticklabels([''] + [0,1])
ax.set_yticklabels([''] + [0,1])
plt.xlabel('Predicted')
plt.ylabel('True')
plt.show()
from sklearn.preprocessing import Normalizer
transformer = Normalizer()
df_norm = df_diabetes.copy()
df_norm[predictors] = transformer.fit_transform(df_diabetes[predictors])
df_norm.describe().transpose()
from sklearn.model_selection import train_test_split
X_norm = df_norm[predictors].values
y_norm = df_norm[target_column].values
X_train_norm, X_test_norm, y_train_norm, y_test_norm = train_test_split(X_norm, y_norm, test_size=0.30, random_state=12)
print(X_train_norm.shape)
print(X_test_norm.shape)
# one hot encode outputs
y_train_norm_cat = to_categorical(y_train_norm)
y_test_norm_cat = to_categorical(y_test_norm)
mlp_norm = Sequential()
mlp_norm.add(Dense(500, activation='relu', input_dim=8))
mlp_norm.add(Dense(100, activation='relu'))
mlp_norm.add(Dense(50, activation='relu'))
mlp_norm.add(Dense(2, activation='softmax'))
mlp_norm.compile(optimizer='adam',
loss='categorical_crossentropy',
metrics=['accuracy'])
mlp_norm.fit(X_train_norm, y_train_norm_cat, epochs=20, verbose=0)
predict_test_norm = mlp_norm.predict_classes(X_test_norm)
print()
print(classification_report(y_test_norm, predict_test_norm))
print()
cnf_mtx = confusion_matrix(y_test_norm, predict_test_norm)
fig = plt.figure()
ax = fig.add_subplot(111)
cax = ax.matshow(cnf_mtx, cmap=plt.cm.Blues)
plt.title('Confusion matrix of the classifier')
fig.colorbar(cax)
for (i, j), z in np.ndenumerate(cnf_mtx):
ax.text(j, i, '{:0.1f}'.format(z), ha='center', va='center', bbox=dict(facecolor='white'))
ax.set_xticklabels([''] + [0,1])
ax.set_yticklabels([''] + [0,1])
plt.xlabel('Predicted')
plt.ylabel('True')
plt.show()
import tensorflow as tf
tf.__version__
```
| github_jupyter |
```
import tensorflow as tf
import numpy as np
from copy import deepcopy
epoch = 20
batch_size = 64
size_layer = 64
dropout_rate = 0.5
n_hops = 2
class BaseDataLoader():
def __init__(self):
self.data = {
'size': None,
'val':{
'inputs': None,
'questions': None,
'answers': None,},
'len':{
'inputs_len': None,
'inputs_sent_len': None,
'questions_len': None,
'answers_len': None}
}
self.vocab = {
'size': None,
'word2idx': None,
'idx2word': None,
}
self.params = {
'vocab_size': None,
'<start>': None,
'<end>': None,
'max_input_len': None,
'max_sent_len': None,
'max_quest_len': None,
'max_answer_len': None,
}
class DataLoader(BaseDataLoader):
def __init__(self, path, is_training, vocab=None, params=None):
super().__init__()
data, lens = self.load_data(path)
if is_training:
self.build_vocab(data)
else:
self.demo = data
self.vocab = vocab
self.params = deepcopy(params)
self.is_training = is_training
self.padding(data, lens)
def load_data(self, path):
data, lens = bAbI_data_load(path)
self.data['size'] = len(data[0])
return data, lens
def build_vocab(self, data):
signals = ['<pad>', '<unk>', '<start>', '<end>']
inputs, questions, answers = data
i_words = [w for facts in inputs for fact in facts for w in fact if w != '<end>']
q_words = [w for question in questions for w in question]
a_words = [w for answer in answers for w in answer if w != '<end>']
words = list(set(i_words + q_words + a_words))
self.params['vocab_size'] = len(words) + 4
self.params['<start>'] = 2
self.params['<end>'] = 3
self.vocab['word2idx'] = {word: idx for idx, word in enumerate(signals + words)}
self.vocab['idx2word'] = {idx: word for word, idx in self.vocab['word2idx'].items()}
def padding(self, data, lens):
inputs_len, inputs_sent_len, questions_len, answers_len = lens
self.params['max_input_len'] = max(inputs_len)
self.params['max_sent_len'] = max([fact_len for batch in inputs_sent_len for fact_len in batch])
self.params['max_quest_len'] = max(questions_len)
self.params['max_answer_len'] = max(answers_len)
self.data['len']['inputs_len'] = np.array(inputs_len)
for batch in inputs_sent_len:
batch += [0] * (self.params['max_input_len'] - len(batch))
self.data['len']['inputs_sent_len'] = np.array(inputs_sent_len)
self.data['len']['questions_len'] = np.array(questions_len)
self.data['len']['answers_len'] = np.array(answers_len)
inputs, questions, answers = deepcopy(data)
for facts in inputs:
for sentence in facts:
for i in range(len(sentence)):
sentence[i] = self.vocab['word2idx'].get(sentence[i], self.vocab['word2idx']['<unk>'])
sentence += [0] * (self.params['max_sent_len'] - len(sentence))
paddings = [0] * self.params['max_sent_len']
facts += [paddings] * (self.params['max_input_len'] - len(facts))
for question in questions:
for i in range(len(question)):
question[i] = self.vocab['word2idx'].get(question[i], self.vocab['word2idx']['<unk>'])
question += [0] * (self.params['max_quest_len'] - len(question))
for answer in answers:
for i in range(len(answer)):
answer[i] = self.vocab['word2idx'].get(answer[i], self.vocab['word2idx']['<unk>'])
self.data['val']['inputs'] = np.array(inputs)
self.data['val']['questions'] = np.array(questions)
self.data['val']['answers'] = np.array(answers)
def bAbI_data_load(path, END=['<end>']):
inputs = []
questions = []
answers = []
inputs_len = []
inputs_sent_len = []
questions_len = []
answers_len = []
for d in open(path):
index = d.split(' ')[0]
if index == '1':
fact = []
if '?' in d:
temp = d.split('\t')
q = temp[0].strip().replace('?', '').split(' ')[1:] + ['?']
a = temp[1].split() + END
fact_copied = deepcopy(fact)
inputs.append(fact_copied)
questions.append(q)
answers.append(a)
inputs_len.append(len(fact_copied))
inputs_sent_len.append([len(s) for s in fact_copied])
questions_len.append(len(q))
answers_len.append(len(a))
else:
tokens = d.replace('.', '').replace('\n', '').split(' ')[1:] + END
fact.append(tokens)
return [inputs, questions, answers], [inputs_len, inputs_sent_len, questions_len, answers_len]
train_data = DataLoader(path='qa5_three-arg-relations_train.txt',is_training=True)
test_data = DataLoader(path='qa5_three-arg-relations_test.txt',is_training=False,
vocab=train_data.vocab, params=train_data.params)
START = train_data.params['<start>']
END = train_data.params['<end>']
def hop_forward(question, memory_o, memory_i, response_proj,
inputs_len, questions_len, is_training):
match = tf.matmul(question, memory_i, transpose_b=True)
match = pre_softmax_masking(match, inputs_len)
match = tf.nn.softmax(match)
match = post_softmax_masking(match, questions_len)
response = tf.matmul(match, memory_o)
return response_proj(tf.concat([response, question], -1))
def pre_softmax_masking(x, seq_len):
paddings = tf.fill(tf.shape(x), float('-inf'))
T = tf.shape(x)[1]
max_seq_len = tf.shape(x)[2]
masks = tf.sequence_mask(seq_len, max_seq_len, dtype=tf.float32)
masks = tf.tile(tf.expand_dims(masks, 1), [1, T, 1])
return tf.where(tf.equal(masks, 0), paddings, x)
def post_softmax_masking(x, seq_len):
T = tf.shape(x)[2]
max_seq_len = tf.shape(x)[1]
masks = tf.sequence_mask(seq_len, max_seq_len, dtype=tf.float32)
masks = tf.tile(tf.expand_dims(masks, -1), [1, 1, T])
return (x * masks)
def shift_right(x):
batch_size = tf.shape(x)[0]
start = tf.to_int32(tf.fill([batch_size, 1], START))
return tf.concat([start, x[:, :-1]], 1)
def embed_seq(x, vocab_size, zero_pad=True):
lookup_table = tf.get_variable('lookup_table', [vocab_size, size_layer], tf.float32)
if zero_pad:
lookup_table = tf.concat((tf.zeros([1, size_layer]), lookup_table[1:, :]), axis=0)
return tf.nn.embedding_lookup(lookup_table, x)
def position_encoding(sentence_size, embedding_size):
encoding = np.ones((embedding_size, sentence_size), dtype=np.float32)
ls = sentence_size + 1
le = embedding_size + 1
for i in range(1, le):
for j in range(1, ls):
encoding[i-1, j-1] = (i - (le-1)/2) * (j - (ls-1)/2)
encoding = 1 + 4 * encoding / embedding_size / sentence_size
return tf.convert_to_tensor(np.transpose(encoding))
def input_mem(x, vocab_size, max_sent_len, is_training):
x = embed_seq(x, vocab_size)
x = tf.layers.dropout(x, dropout_rate, training=is_training)
pos = position_encoding(max_sent_len, size_layer)
x = tf.reduce_sum(x * pos, 2)
return x
def quest_mem(x, vocab_size, max_quest_len, is_training):
x = embed_seq(x, vocab_size)
x = tf.layers.dropout(x, dropout_rate, training=is_training)
pos = position_encoding(max_quest_len, size_layer)
return (x * pos)
class QA:
def __init__(self, vocab_size):
self.questions = tf.placeholder(tf.int32,[None,None])
self.inputs = tf.placeholder(tf.int32,[None,None,None])
self.questions_len = tf.placeholder(tf.int32,[None])
self.inputs_len = tf.placeholder(tf.int32,[None])
self.answers_len = tf.placeholder(tf.int32,[None])
self.answers = tf.placeholder(tf.int32,[None,None])
self.training = tf.placeholder(tf.bool)
max_sent_len = train_data.params['max_sent_len']
max_quest_len = train_data.params['max_quest_len']
max_answer_len = train_data.params['max_answer_len']
lookup_table = tf.get_variable('lookup_table', [vocab_size, size_layer], tf.float32)
lookup_table = tf.concat((tf.zeros([1, size_layer]), lookup_table[1:, :]), axis=0)
with tf.variable_scope('questions'):
question = quest_mem(self.questions, vocab_size, max_quest_len, self.training)
with tf.variable_scope('memory_o'):
memory_o = input_mem(self.inputs, vocab_size, max_sent_len, self.training)
with tf.variable_scope('memory_i'):
memory_i = input_mem(self.inputs, vocab_size, max_sent_len, self.training)
with tf.variable_scope('interaction'):
response_proj = tf.layers.Dense(size_layer)
for _ in range(n_hops):
answer = hop_forward(question,
memory_o,
memory_i,
response_proj,
self.inputs_len,
self.questions_len,
self.training)
question = answer
with tf.variable_scope('memory_o', reuse=True):
embedding = tf.get_variable('lookup_table')
cell = tf.nn.rnn_cell.LSTMCell(size_layer)
vocab_proj = tf.layers.Dense(vocab_size)
state_proj = tf.layers.Dense(size_layer)
init_state = state_proj(tf.layers.flatten(answer))
init_state = tf.layers.dropout(init_state, dropout_rate, training=self.training)
helper = tf.contrib.seq2seq.TrainingHelper(
inputs = tf.nn.embedding_lookup(embedding, shift_right(self.answers)),
sequence_length = tf.to_int32(self.answers_len))
encoder_state = tf.nn.rnn_cell.LSTMStateTuple(c=init_state, h=init_state)
decoder = tf.contrib.seq2seq.BasicDecoder(cell = cell,
helper = helper,
initial_state = encoder_state,
output_layer = vocab_proj)
decoder_output, _, _ = tf.contrib.seq2seq.dynamic_decode(decoder = decoder,
maximum_iterations = tf.shape(self.inputs)[1])
self.outputs = decoder_output.rnn_output
helper = tf.contrib.seq2seq.GreedyEmbeddingHelper(embedding = embedding,
start_tokens = tf.tile(
tf.constant([START],
dtype=tf.int32),
[tf.shape(self.inputs)[0]]),
end_token = END)
decoder = tf.contrib.seq2seq.BasicDecoder(
cell = cell,
helper = helper,
initial_state = encoder_state,
output_layer = vocab_proj)
decoder_output, _, _ = tf.contrib.seq2seq.dynamic_decode(
decoder = decoder,
maximum_iterations = max_answer_len)
self.logits = decoder_output.sample_id
correct_pred = tf.equal(self.logits, self.answers)
self.accuracy = tf.reduce_mean(tf.cast(correct_pred, tf.float32))
self.cost = tf.reduce_mean(tf.contrib.seq2seq.sequence_loss(logits = self.outputs,
targets = self.answers,
weights = tf.ones_like(self.answers, tf.float32)))
self.optimizer = tf.train.AdamOptimizer().minimize(self.cost)
tf.reset_default_graph()
sess = tf.InteractiveSession()
model = QA(train_data.params['vocab_size'])
sess.run(tf.global_variables_initializer())
batching = (train_data.data['val']['inputs'].shape[0] // batch_size) * batch_size
for i in range(epoch):
total_cost, total_acc = 0, 0
for k in range(0, batching, batch_size):
batch_questions = train_data.data['val']['questions'][k:k+batch_size]
batch_inputs = train_data.data['val']['inputs'][k:k+batch_size]
batch_inputs_len = train_data.data['len']['inputs_len'][k:k+batch_size]
batch_questions_len = train_data.data['len']['questions_len'][k:k+batch_size]
batch_answers_len = train_data.data['len']['answers_len'][k:k+batch_size]
batch_answers = train_data.data['val']['answers'][k:k+batch_size]
acc, cost, _ = sess.run([model.accuracy,model.cost,model.optimizer],
feed_dict={model.questions:batch_questions,
model.inputs:batch_inputs,
model.inputs_len:batch_inputs_len,
model.questions_len:batch_questions_len,
model.answers_len:batch_answers_len,
model.answers:batch_answers,
model.training:True})
total_cost += cost
total_acc += acc
total_cost /= (train_data.data['val']['inputs'].shape[0] // batch_size)
total_acc /= (train_data.data['val']['inputs'].shape[0] // batch_size)
print('epoch %d, avg cost %f, avg acc %f'%(i+1,total_cost,total_acc))
testing_size = 32
batch_questions = test_data.data['val']['questions'][:testing_size]
batch_inputs = test_data.data['val']['inputs'][:testing_size]
batch_inputs_len = test_data.data['len']['inputs_len'][:testing_size]
batch_questions_len = test_data.data['len']['questions_len'][:testing_size]
batch_answers_len = test_data.data['len']['answers_len'][:testing_size]
batch_answers = test_data.data['val']['answers'][:testing_size]
logits = sess.run(model.logits,
feed_dict={model.questions:batch_questions,
model.inputs:batch_inputs,
model.inputs_len:batch_inputs_len,
model.questions_len:batch_questions_len,
model.answers_len:batch_answers_len,
model.training:False})
for i in range(testing_size):
print('QUESTION:',' '.join([train_data.vocab['idx2word'][k] for k in batch_questions[i]]))
print('REAL:',train_data.vocab['idx2word'][batch_answers[i,0]])
print('PREDICT:',train_data.vocab['idx2word'][logits[i,0]],'\n')
```
| github_jupyter |
<a href="https://colab.research.google.com/github/AIWintermuteAI/aXeleRate/blob/master/resources/aXeleRate_test_classifier.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
## Classification model Training and Inference
In this notebook we will use axelerate Keras-based framework for AI on the edge to quickly setup model training and then after training session is completed convert it to .tflite and .kmodel formats.
First, let's take care of some administrative details.
1) Before we do anything, make sure you have choosen GPU as Runtime type (in Runtime - > Change Runtime type).
2) We need to mount Google Drive for saving our model checkpoints and final converted model(s). Press on Mount Google Drive button in Files tab on your left.
In the next cell we clone axelerate Github repository and import it.
**It is possible to use pip install or python setup.py install, but in that case you will need to restart the enironment.** Since I'm trying to make the process as streamlined as possibile I'm using sys.path.append for import.
```
%tensorflow_version 1.x
!git clone https://github.com/AIWintermuteAI/aXeleRate.git
import sys
sys.path.append('/content/aXeleRate')
from axelerate import setup_training,setup_inference
```
At this step you typically need to get the dataset. You can use !wget command to download it from somewhere on the Internet or !cp to copy from My Drive as in this example
```
!cp -r /content/drive/'My Drive'/pascal_20_segmentation.zip .
!unzip --qq pascal_20_segmentation.zip
```
For this notebook small test dataset is already in axelerate/sample_datasets folder, so no need to download anything.
Let's visualize our classification test dataset. There are two images per class and class label is the name of the folder with images belonging to that class.
```
%matplotlib inline
import matplotlib.image as mpimg
import matplotlib.pyplot as plt
import glob
def show_image(filename):
print(filename)
image = mpimg.imread(filename)
plt.figure()
plt.imshow(image)
plt.show()
image_files_list = glob.glob('aXeleRate/sample_datasets/classifier/imgs' + '/**/*.jpg', recursive=True)
for filename in image_files_list:
show_image(filename)
```
Next step is defining a config dictionary. Most lines are self-explanatory.
Type is model frontend - Classifier, Detector or Segnet
Architecture is model backend (feature extractor)
- Full Yolo
- Tiny Yolo
- MobileNet1_0
- MobileNet7_5
- MobileNet5_0
- MobileNet2_5
- SqueezeNet
- VGG16
- ResNet50
Fully_connected is number of neurons in classification layers as list.
Dropout value is dropout in classification layers.
```
config = {
"model" : {
"type": "Classifier",
"architecture": "MobileNet7_5",
"input_size": 224,
"fully-connected": [100,50],
"labels": [],
"dropout" : 0.5
},
"weights" : {
"full": "",
"backend": "imagenet",
"save_bottleneck": False
},
"train" : {
"actual_epoch": 5,
"train_image_folder": "aXeleRate/sample_datasets/classifier/imgs",
"train_times": 4,
"valid_image_folder": "aXeleRate/sample_datasets/classifier/imgs_validation",
"valid_times": 4,
"valid_metric": "val_accuracy",
"batch_size": 4,
"learning_rate": 1e-3,
"saved_folder": "classifier",
"first_trainable_layer": "",
"augumentation": True
},
"converter" : {
"type": ["k210","tflite"]
}
}
```
Let's check what GPU we have been assigned in this Colab session, if any.
```
from tensorflow.python.client import device_lib
device_lib.list_local_devices()
```
Finally we start the training by passing config dictionary we have defined earlier to setup_training function. The function will start the training with Checkpoint, Reduce Learning Rate on Plateu and Early Stopping callbacks. After the training has stopped, it will convert the best model into the format you have specified in config and save it to the project folder.
```
model_path = setup_training(config_dict=config)
```
After training it is good to check the actual perfomance of your model by doing inference on your validation dataset and visualizing results. This is exactly what next block does. Obviously since our model has only trained on a few images the results are far from stellar, but if you have a good dataset, you'll have better results.
```
from keras import backend as K
K.clear_session()
setup_inference(config, model_path)
```
Good luck and happy training! Have a look at these articles, that would allow you to get the most of Google Colab or connect to local runtime if there are no GPUs available;
https://medium.com/@oribarel/getting-the-most-out-of-your-google-colab-2b0585f82403
https://research.google.com/colaboratory/local-runtimes.html
| github_jupyter |
# Deep Learning Toolkit for Splunk - Notebook for STL - Seasonality and Trend Decomposition
This notebook contains a barebone example workflow how to work on custom containerized code that seamlessly interfaces with the Deep Learning Toolkit for Splunk.
Note: By default every time you save this notebook the cells are exported into a python module which is then invoked by Splunk MLTK commands like <code> | fit ... | apply ... | summary </code>. Please read the Model Development Guide in the Deep Learning Toolkit app for more information.
## Stage 0 - import libraries
At stage 0 we define all imports necessary to run our subsequent code depending on various libraries.
```
# this definition exposes all python module imports that should be available in all subsequent commands
import json
import numpy as np
import pandas as pd
from statsmodels.tsa.seasonal import STL
import statsmodels as sm
# ...
# global constants
MODEL_DIRECTORY = "/srv/app/model/data/"
# THIS CELL IS NOT EXPORTED - free notebook cell for testing or development purposes
print("numpy version: " + np.__version__)
print("pandas version: " + pd.__version__)
print("statsmodels version: " + sm.__version__)
```
## Stage 1 - get a data sample from Splunk
In Splunk run a search to pipe a dataset into your notebook environment. Note: mode=stage is used in the | fit command to do this.
| inputlookup cyclical_business_process.csv<br>
| fit MLTKContainer mode=stage algo=seasonality_and_trend_decomposition _time logons
After you run this search your data set sample is available as a csv inside the container to develop your model. The name is taken from the into keyword ("barebone_model" in the example above) or set to "default" if no into keyword is present. This step is intended to work with a subset of your data to create your custom model.
```
# this cell is not executed from MLTK and should only be used for staging data into the notebook environment
def stage(name):
with open("data/"+name+".csv", 'r') as f:
df = pd.read_csv(f)
with open("data/"+name+".json", 'r') as f:
param = json.load(f)
return df, param
# THIS CELL IS NOT EXPORTED - free notebook cell for testing or development purposes
df, param = stage("default")
print(df.describe())
print(param)
```
## Stage 2 - create and initialize a model
```
# initialize your model
# available inputs: data and parameters
# returns the model object which will be used as a reference to call fit, apply and summary subsequently
def init(df,param):
model = {}
return model
# THIS CELL IS NOT EXPORTED - free notebook cell for testing or development purposes
print(init(df,param))
model=init(df,param)
```
## Stage 3 - fit the model
```
# train your model
# returns a fit info json object and may modify the model object
def fit(model,df,param):
return "info"
# THIS CELL IS NOT EXPORTED - free notebook cell for testing or development purposes
print(fit(model,df,param))
```
## Stage 4 - apply the model
```
# apply your model
# returns the calculated results
def apply(model,df,param):
data=df
data['_time']=pd.to_datetime(data['_time'])
data = data.set_index('_time') # Set the index to datetime object.
data=data.asfreq('H')
res=STL(data).fit()
results=pd.DataFrame({"seasonality": res.seasonal, "trend": res.trend, "residual": res.resid})
results.reset_index(level=0, inplace=True)
return results
# THIS CELL IS NOT EXPORTED - free notebook cell for testing or development purposes
apply(model,df,param)
```
## Stage 5 - save the model
```
# save model to name in expected convention "<algo_name>_<model_name>"
def save(model,name):
return model
```
## Stage 6 - load the model
```
# load model from name in expected convention "<algo_name>_<model_name>"
def load(name):
return model
```
## Stage 7 - provide a summary of the model
```
# return a model summary
def summary(model=None):
returns = {"version": {"numpy": np.__version__, "pandas": pd.__version__} }
return returns
```
## End of Stages
All subsequent cells are not tagged and can be used for further freeform code
| github_jupyter |
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import os.path
from datetime import datetime
import seaborn as sns
from statsmodels.tsa.seasonal import seasonal_decompose
# this package is extern from Jupyter Notebook
# it is used to create maps. It's so cool. Try it!
#import folium
list_tripFiles = ['dataFiles/data-sample_data-nyctaxi-trips-2009-json_corrigido.json',
'dataFiles/data-sample_data-nyctaxi-trips-2010-json_corrigido.json',
'dataFiles/data-sample_data-nyctaxi-trips-2011-json_corrigido.json',
'dataFiles/data-sample_data-nyctaxi-trips-2012-json_corrigido.json']
dfDataTrips10 = pd.read_json(list_tripFiles[1])
dfDataTrips12 = pd.read_json(list_tripFiles[3])
```
<h2>Vamos responder a Primeira pergunta!</h2>
<h1>1. Qual a distância média percorrida por viagens com no máximo 2 passageiros;</h1>
```
# vou usar apenas um arquivo para ter mais velocidade
dfPassengers = dfDataTrips10.query('passenger_count <= 2')
display(dfPassengers)
```
<h3>Interessante. São 17 colunas. Os dados que precisamos estão nas colunas 'passenger_count' e 'trip_distance'.
Vamos agora tentar gerar a média da distância.</h3>
```
dfMeanTotal = np.mean(dfDataTrips10['trip_distance']) # media total do arquivo
dfMeanPass = np.mean(dfPassengers['trip_distance']) # media das corridas com no máximo dois passageiros
print(dfMeanTotal)
print(dfMeanPass)
```
<h3>Muito Bem. Já temos as médias, agora é só passar para o gráfico na próxima etapa.<br>
Intessante, não existe uma grande diferença relacional entre a distância média e a quantidade de passageiros.<br>
Vamos para a Segunda Pergunta;</h3>
<h1> Quais os 3 maiores vendors em quantidade total de dinheiro arrecadado;</h1>
```
vendorsFile = 'dataFiles/data-vendor_lookup-csv.csv'
dfDataVendors = pd.read_csv(vendorsFile)
display(dfDataVendors)
```
<h3>Um arquivo pequeno. Vamos precisar dos dados da coluna 'vendor_id' e 'name'.</h3>
```
#vamos testar com apenas uma companhia
vendor = dfDataVendors["vendor_id"][0] # pega o id da empresa
#filtra no arquivos das corridas somente as realizadas por essa companhia
dfVendor = dfDataTrips10.query(f'vendor_id == "{vendor}"')
display(dfVendor)
```
<h3>Aí está, somente os dados da companhia com o id 'CMT'.
<br>Vamos precisar das últimas cinco colunas.
<br>'fare_amount' = Valor somente da tarifa
<br>'surcharge' = Valor da taxa extra (bandeira 2, 3)
<br>'tip_amount' = Valor da gorgeta
<br>'tolls_amount' = Valor dos pedágios
<br>'total_amount' = Valor total da corrida
</h3>
```
# soma o valor total da corrida
dfSumVendor = np.sum(dfVendor['total_amount']).round(2)
# soma somente a tarifa e sobre taxas
dfSumFares = np.sum(dfVendor['fare_amount'] + dfVendor['surcharge']).round(2)
print(dfSumVendor)
print(dfSumFares)
```
<h3>Agora temos os valores da companhia CMT-Creative Mobile Technologies, LLC.
<br>Basta colocar isso dentro de um loop e teremos de todas.
<br>Em seguida, depois que coletarmos todo o faturamento de cada companhia, precisamos pegar apenas as três maiores. E isso é simples ao usar o pandas.
<br>É só usar a função "pd.nlargest", e passar os seguintes parâmetros(3, 'Amount') </h3>
Veremos:
```
total_vendors = len(dfDataVendors)
list_amount = []
list_fare_amount = []
df_big_vendors = pd.DataFrame()
df_big_vendors['Alias'] = dfDataVendors['vendor_id']
df_big_vendors['Name'] = dfDataVendors['name']
i = 0
while i < total_vendors:
# get vendor_id and uses it to filter only trips of the respective vendor
vendor = dfDataVendors["vendor_id"][i]
dfVendor = dfDataTrips10.query(f'vendor_id == "{vendor}"')
# so, sum the amount of all trips, in the two ways mentioned above
dfSumVendor = np.sum(dfVendor['total_amount']).round(2)
list_amount.append(int(dfSumVendor))
dfSumFares = np.sum(dfVendor['fare_amount'] + dfVendor['surcharge']).round(2)
list_fare_amount.append(int(dfSumFares))
i += 1
# add sums lists in DataFrame, each in different column
df_big_vendors['Amount'] = list_amount
df_big_vendors['Fare_Amount'] = list_fare_amount
display(df_big_vendors)
df_big_vendors = df_big_vendors.nlargest(3, 'Amount') # get only three biggest vendors
display(df_big_vendors)
```
<h4>Vamos para a próxima pergunta;</h4>
<h2>3 - Faça um histograma da distribuição mensal, nos 4 anos, de corridas pagas em dinheiro</h1>
<h4>Se envolve um histograma de distribuição mensal, antes precisamos converter a coluna com datas ('pickup_datetime')</h4>
```
dfDataTrips10['pickup_datetime'] = pd.to_datetime(dfDataTrips10['pickup_datetime'])
# já vamos separar apenas as corridas pagas em dinheiro
dfCashTrips = dfDataTrips10.query('payment_type == "CASH" | payment_type == "Cash"')
display(dfCashTrips)
```
<h3>Vamos contar os números de corridas por mês.</h3>
```
month = 1
list_count_trips = []
while month <= 12:
# filtra somente as corridas do mês informado na iteração do loop
trips = dfCashTrips.query('pickup_datetime.dt.month == @month')
count_trips = np.count_nonzero(trips['pickup_datetime']) # conta o total de corridas
list_count_trips.append(count_trips)
month += 1
print(list_count_trips)
```
<h3>Que bom. Esses são os números de corridas pagas em dinheiro separadas por mês.
<br>Agora é só colocar isso dentro de outro loop (para separar por ano) e usar uma função de histograma para gerar o gráfico</h3>
<h4>Vamos para a próxima pergunta;</h4>
<h2>4. Faça um gráfico de série temporal contando a quantidade de gorjetas de cada dia, nos últimos 3 meses de 2012.</h1>
```
# Vamos converter a coluna DateTime do arquivo de 2012
dfDataTrips12['pickup_datetime'] = pd.to_datetime(dfDataTrips12['pickup_datetime'])
# E agora vamos pegar apenas os dados das corridas dos últimos 3 meses do ano em que houveram gorgetas.
df_tips = dfDataTrips12.query("pickup_datetime.dt.month > 9 & tip_amount > 0")
display(df_tips)
```
<h3>Legal, agora temos os dados que precisamos, <b>16.152</b> corridas com gorgetas!
<br>Agora precisamos separar por mês, e por dia. Vamos ver!</h3>
```
month = 10 # variável de controle para cada mês
dict_diary_tips = {}
list_days = []
list_tips = []
# o loop externo, que irá verificar os meses
while month <= 12:
day = 1
# o loop interno, que irá verificar os dias
while day <= 31:
list_days.append((str(month)+"-"+str(day)))
# Novembro não possui 31 dias, então precisamos inserir 0 nesse campo
if month == 11 and day == 31:
list_tips.append(0)
else:
# e agora conta a quantidade de corrida do dia respectivo
diary_tips = df_tips.query(f"pickup_datetime.dt.month == {month} & pickup_datetime.dt.day == {day}")
list_tips.append(len(diary_tips.index))
day += 1
# Eu coloquei os dados em um dicionário, por ser mais fácil ir inserindo novos elementos nele
# Em seguida eu adiciono o dicionário no DataFrame
dict_diary_tips["Dias"] = list_days
dict_diary_tips["Gorgetas"] = list_tips
month += 1
df_diary_tips = pd.DataFrame(dict_diary_tips)
#df_diary_tips = df_diary_tips.set_index('Dias')
display(df_diary_tips)
<h3>Hum. Que estranho, não existem dados nos últimos 5 registros.
Vamos tentar novamente!</h3>
display(df_diary_tips.loc[60:])
<h3>Como imaginei, não existem dados do mês de Dezembro.
<br>Vamos tentar novamente, agora com Novembro!</h3>
display(df_diary_tips.loc[31:60])
<h3>E o mesmo ocorre com Novembro.
<br>Vamos tentar com Outubro!</h3>
display(df_diary_tips.loc[:31])
```
<h3>Parece haver um corte no dia 27 de Outubro. Provavelmente para limitar o tamanho do arquivo
<br>Vamos tentar novamente, usando outro período!
<br>De Julho a Setembro</h3>
```
# primeiro vamos pegar novos dados
df_tips = dfDataTrips12.query("pickup_datetime.dt.month > 6 & pickup_datetime.dt.month <= 9 & tip_amount > 0")
month = 7 # variável de controle para cada mês
dict_diary_tips = {}
list_days = []
list_tips = []
# o loop externo, que irá verificar os meses
while month <= 9:
day = 1
# o loop interno, que irá verificar os dias
while day <= 31:
list_days.append((str(month)+"-"+str(day)))
# Setembro não possui 31 dias, então precisamos inserir 0 nesse campo
if month == 9 and day == 31:
list_tips.append(0)
else:
# e agora conta a quantidade de corrida do dia respectivo
diary_tips = df_tips.query(f"pickup_datetime.dt.month == {month} & pickup_datetime.dt.day == {day}")
list_tips.append(len(diary_tips.index))
day += 1
# Eu coloquei os dados em um dicionário, por ser mais fácil ir inserindo novos elementos nele
# Em seguida eu adiciono o dicionário no DataFrame
dict_diary_tips["Dias"] = list_days
dict_diary_tips["Gorgetas"] = list_tips
month += 1
df_diary_tips = pd.DataFrame(dict_diary_tips)
#df_diary_tips = df_diary_tips.set_index('Dias')
display(df_diary_tips)
```
<h3>Agora sim, os dados parecem estar completos.
<br>É só passar esses dados para uma função gerar os gráficos.</h3>
<p><h3>Vamos para a próxima pergunta;</h3>
<h2>5. Qual o tempo médio das corridas nos dias de sábado e domingo.</h2>
<h3>Para isso, vamos precisar converter para datetime também a coluna de desembarque.
<br>Em seguida, vamos armazenar separadamente em 3 dataFrame as corridas da seguinte forma;
<br>-- Dias úteis
<br>-- Sábados
<br>-- Domingos </h3>
```
dfDataTrips10['dropoff_datetime'] = pd.to_datetime(dfDataTrips10['dropoff_datetime'])
# cria os dataframes com as corridas de cada dia
# para isso usei a função dayofweek, do pandas. Ela retorna um inteiro de 0 a 6 para cada dia, sendo 0 Segunda-Feira.
dfSundayTrips = dfDataTrips10.query("pickup_datetime.dt.dayofweek == 6")
dfSaturdayTrips = dfDataTrips10.query("pickup_datetime.dt.dayofweek == 5")
dfWorkdayTrips = dfDataTrips10.query("pickup_datetime.dt.dayofweek <= 4")
display(dfSundayTrips.head())
display(dfSaturdayTrips.head())
display(dfWorkdayTrips.head())
# cria um dicionário para armazenar somente o tempo médio da corrida. Não preciso das outras colunas
# para tirar a média, usei novamente a função mean do numpy
# Fiz a média resultante da subtração entre o desembarque e o embarque.
dict_timeTrips = {}
dict_timeTrips['work_time_trip'] = np.mean(dfWorkdayTrips['dropoff_datetime'] - dfWorkdayTrips['pickup_datetime'])
dict_timeTrips['sat_time_trip'] = np.mean(dfSaturdayTrips['dropoff_datetime'] - dfSaturdayTrips['pickup_datetime'])
dict_timeTrips['sun_time_trip'] = np.mean(dfSundayTrips['dropoff_datetime'] - dfSundayTrips['pickup_datetime'])
# Exibe os resultados.
print("Média dias úteis: ", dict_timeTrips['work_time_trip'])
print("Média sábados: ", dict_timeTrips['sat_time_trip'])
print("Média domingos: ", dict_timeTrips['sun_time_trip'])
```
<h3>Números muito semelhantes. Parece não haver muita diferença no tempo do trajeto das pessoas que andam de taxi durante a semana e aos sábados, com uma quase imperceptível queda aos domingos.</h3>
| github_jupyter |
# Visualizing the Johns Hopkins COVID-19 time series data
**This is a work in progress.** It doesn't work yet in [Binder](https://mybinder.org/v2/gh/dotnet/interactive/master?urlpath=lab) because it relies on HTTP communication between the kernel and the Jupyter frontend.
Also, due to travel restrictions, you should run this at home on isolated compute.
*And don't forget to wash your hands.*
Since Johns Hopkins has put COVID-19 time series data on [GitHub](https://github.com/CSSEGISandData/COVID-19), let's take a look at it. We can download it using PowerShell:
```
#!pwsh
Invoke-WebRequest -Uri "https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_time_series/time_series_19-covid-Confirmed.csv" -OutFile "./Confirmed.csv"
Invoke-WebRequest -Uri "https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_time_series/time_series_19-covid-Deaths.csv" -OutFile "./Deaths.csv"
Invoke-WebRequest -Uri "https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_time_series/time_series_19-covid-Recovered.csv" -OutFile "./Recovered.csv"
```
It needs a little cleaning up:
```
using System.IO;
using System.Text.RegularExpressions;
Clean("Confirmed.csv");
Clean("Deaths.csv");
Clean("Recovered.csv");
void Clean(string filePath)
{
var raw = File.ReadAllText(filePath);
var regex = new Regex("\\\"(.*?)\\\"");
var cleaned = regex.Replace(raw, m => m.Value.Replace(",", " in "));
File.WriteAllText(filePath, cleaned);
}
```
Next, let's load it into a data frame.
```
#r "nuget:Microsoft.Data.Analysis,0.2.0"
using Microsoft.Data.Analysis;
var deaths = DataFrame.LoadCsv("./Deaths.csv");
var confirmed = DataFrame.LoadCsv("./Confirmed.csv");
var recovered = DataFrame.LoadCsv("./Recovered.csv");
var offset = 4;
var date = deaths.Columns[offset].Name;
var deathFiltered = deaths[deaths.Columns[offset].ElementwiseNotEquals(0)];
var confirmedFiltered = confirmed[confirmed.Columns[offset].ElementwiseNotEquals(0)];
var recoveredFiltered = recovered[recovered.Columns[offset].ElementwiseNotEquals(0)];
var current = confirmed.Columns[offset] - (deaths.Columns[offset] + recovered.Columns[offset]);
var currentLocFiltered = confirmed[current.ElementwiseNotEquals(0)];
var currentDataFiltered = current.Clone(current.ElementwiseNotEquals(0));
var deathsSeries = new {
latitude = deathFiltered["Lat"],
longitude = deathFiltered["Long"],
data = deathFiltered.Columns[offset]
};
var confirmedSeries = new {
latitude = confirmedFiltered["Lat"],
longitude = confirmedFiltered["Long"],
data = confirmedFiltered.Columns[offset]
};
var recoveredSeries = new {
latitude = recoveredFiltered["Lat"],
longitude = recoveredFiltered["Long"],
data = recoveredFiltered.Columns[offset]
};
var currentSeries = new {
latitude = currentLocFiltered["Lat"],
longitude = currentLocFiltered["Long"],
data = currentDataFiltered
};
"Ready."
```
Because we've stored our data in top-level variables (`deathsSeries`, `confirmedSeries`, `recoveredSeries`, etc.) in the C# kernel, they're accessible from JavaScript by calling `interactive.csharp.getVariable`. The data will be returned as JSON and we can plot it using the library of our choice, pulled in using [RequireJS](https://requirejs.org/).
We'll use [Plotly](https://plot.ly/).
```
#!javascript
var plot = function() {
plotlyJs_covid_require = require.config({
context: "COVID",
paths: {
plotly: "https://cdn.plot.ly/plotly-latest.min"
}
});
plotlyJs_covid_require(["plotly"], (Plotly) => {
function updateCovidPlot(){
Promise.all([
interactive.csharp.getVariable("deathsSeries"),
interactive.csharp.getVariable("confirmedSeries"),
interactive.csharp.getVariable("recoveredSeries"),
interactive.csharp.getVariable("currentSeries"),
interactive.csharp.getVariable("date")])
.then(data => {
Plotly.animate("plotlyChartCovid", {
data:[
{
lat: data[2].latitude,
lon: data[2].longitude,
text: data[2].data
},
{
lat: data[0].latitude,
lon: data[0].longitude,
text: data[0].data
},
{
lat: data[1].latitude,
lon: data[1].longitude,
text: data[1].data
},
{
lat: data[3].latitude,
lon: data[3].longitude,
text: data[3].data
}],
layout:{
title: "COVID-19 "+ data[4]
}
});
});
}
Promise.all([
interactive.csharp.getVariable("deathsSeries"),
interactive.csharp.getVariable("confirmedSeries"),
interactive.csharp.getVariable("recoveredSeries"),
interactive.csharp.getVariable("currentSeries"),
interactive.csharp.getVariable("date")])
.then(data => {
var recovered = {
name: "recovered",
type: "scattergeo",
mode: "markers",
geo: "geo1",
lat: data[2].latitude,
lon: data[2].longitude,
text: data[2].data,
marker: {
symbol: "square",
colorscale: "Viridis",
}
};
var deaths = {
name: "death",
type: "scattergeo",
geo: "geo2",
mode: "markers",
lat: data[0].latitude,
lon: data[0].longitude,
text: data[0].data,
marker: {
symbol: "circle",
colorscale: "Viridis",
}
};
var confirmed = {
name: "confirmed",
type: "scattergeo",
geo: "geo3",
mode: "markers",
lat: data[1].latitude,
lon: data[1].longitude,
text: data[1].data,
marker: {
symbol: "diamond",
colorscale: "Viridis",
}
};
var current = {
name: "current",
type: "scattergeo",
geo: "geo4",
mode: "markers",
lat: data[3].latitude,
lon: data[3].longitude,
text: data[3].data,
marker: {
symbol: "triangle",
colorscale: "Viridis",
}
};
var traces = [recovered, deaths, confirmed, current];
var layout = {
title: "COVID-19 "+ data[4],
grid: { columns: 4, rows: 1 },
geo1: {
scope: "world",
showland: true,
landcolor: "rgb(250,250,250)",
domain: {
row: 0,
column: 0
}
},
geo2: {
scope: "world",
showland: true,
landcolor: "rgb(250,250,250)",
domain: {
row: 0,
column: 1
}
},
geo3: {
scope: "world",
showland: true,
landcolor: "rgb(250,250,250)",
domain: {
row: 0,
column: 2
}
},
geo4: {
scope: "world",
showland: true,
landcolor: "rgb(250,250,250)",
domain: {
row: 0,
column: 3
}
}
};
Plotly.newPlot("plotlyChartCovid", traces, layout);
setInterval(() => updateCovidPlot(), 100);
});
});
};
```
Notice the `setInterval` call near the end of the previous cell. This rechecks the data in the kernel and updates the plot.
Back on the kernel, we can now update the data so that the kernel can see it.
Yes, this is a contrived example, and we're planning to support true streaming data, but it's a start.
```
#!html
<div id="plotlyChartCovid"></div>
#!js
plot();
#!csharp
for(var i = offset; i < deaths.Columns.Count; i++){
await Task.Delay(100);
date = deaths.Columns[i].Name;
deathFiltered = deaths[deaths.Columns[i].ElementwiseNotEquals(0)];
confirmedFiltered = confirmed[confirmed.Columns[i].ElementwiseNotEquals(0)];
recoveredFiltered = recovered[recovered.Columns[i].ElementwiseNotEquals(0)];
current = confirmed.Columns[i] - (deaths.Columns[i] + recovered.Columns[i]);
currentLocFiltered = confirmed[current.ElementwiseNotEquals(0)];
currentDataFiltered = current.Clone(current.ElementwiseNotEquals(0));
deathsSeries = new {
latitude = deathFiltered["Lat"],
longitude = deathFiltered["Long"],
data = deathFiltered.Columns[i]
};
confirmedSeries = new {
latitude = confirmedFiltered["Lat"],
longitude = confirmedFiltered["Long"],
data = confirmedFiltered.Columns[i]
};
recoveredSeries = new {
latitude = recoveredFiltered["Lat"],
longitude = recoveredFiltered["Long"],
data = recoveredFiltered.Columns[i]
};
currentSeries = new {
latitude = currentLocFiltered["Lat"],
longitude = currentLocFiltered["Long"],
data = currentDataFiltered
};
}
```
| github_jupyter |
# Intro
[PyTorch](https://pytorch.org/) is a very powerful machine learning framework. Central to PyTorch are [tensors](https://pytorch.org/docs/stable/tensors.html), a generalization of matrices to higher ranks. One intuitive example of a tensor is an image with three color channels: A 3-channel (red, green, blue) image which is 64 pixels wide and 64 pixels tall is a $3\times64\times64$ tensor. You can access the PyTorch framework by writing `import torch` near the top of your code, along with all of your other import statements.
This guide will help introduce you to the functionality of PyTorch, but don't worry too much about memorizing it: the assignments will link to relevant documentation where necessary.
```
import torch
```
# Why PyTorch?
One important question worth asking is, why is PyTorch being used for this course? There is a great breakdown by [the Gradient](https://thegradient.pub/state-of-ml-frameworks-2019-pytorch-dominates-research-tensorflow-dominates-industry/) looking at the state of machine learning frameworks today. In part, as highlighted by the article, PyTorch is generally more pythonic than alternative frameworks, easier to debug, and is the most-used language in machine learning research by a large and growing margin. While PyTorch's primary alternative, Tensorflow, has attempted to integrate many of PyTorch's features, Tensorflow's implementations come with some inherent limitations highlighted in the article.
Notably, while PyTorch's industry usage has grown, Tensorflow is still (for now) a slight favorite in industry. In practice, the features that make PyTorch attractive for research also make it attractive for education, and the general trend of machine learning research and practice to PyTorch makes it the more proactive choice.
# Tensor Properties
One way to create tensors from a list or an array is to use `torch.Tensor`. It'll be used to set up examples in this notebook, but you'll never need to use it in the course - in fact, if you find yourself needing it, that's probably not the correct answer.
```
example_tensor = torch.Tensor(
[
[[1, 2], [3, 4]],
[[5, 6], [7, 8]],
[[9, 0], [1, 2]]
]
)
```
You can view the tensor in the notebook by simple printing it out (though some larger tensors will be cut off)
```
example_tensor
```
## Tensor Properties: Device
One important property is the device of the tensor - throughout this notebook you'll be sticking to tensors which are on the CPU. However, throughout the course you'll also be using tensors on GPU (that is, a graphics card which will be provided for you to use for the course). To view the device of the tensor, all you need to write is `example_tensor.device`. To move a tensor to a new device, you can write `new_tensor = example_tensor.to(device)` where device will be either `cpu` or `cuda`.
```
example_tensor.device
```
## Tensor Properties: Shape
And you can get the number of elements in each dimension by printing out the tensor's shape, using `example_tensor.shape`, something you're likely familiar with if you've used numpy. For example, this tensor is a $3\times2\times2$ tensor, since it has 3 elements, each of which are $2\times2$.
```
example_tensor.shape
```
You can also get the size of a particular dimension $n$ using `example_tensor.shape[n]` or equivalently `example_tensor.size(n)`
```
print("shape[0] =", example_tensor.shape[0])
print("size(1) =", example_tensor.size(1))
```
Finally, it is sometimes useful to get the number of dimensions (rank) or the number of elements, which you can do as follows
```
print("Rank =", len(example_tensor.shape))
print("Number of elements =", example_tensor.numel())
```
# Indexing Tensors
As with numpy, you can access specific elements or subsets of elements of a tensor. To access the $n$-th element, you can simply write `example_tensor[n]` - as with Python in general, these dimensions are 0-indexed.
```
example_tensor[1]
```
In addition, if you want to access the $j$-th dimension of the $i$-th example, you can write `example_tensor[i, j]`
```
example_tensor[1, 1, 0]
```
Note that if you'd like to get a Python scalar value from a tensor, you can use `example_scalar.item()`
```
example_scalar = example_tensor[1, 1, 0]
example_scalar.item()
```
In addition, you can index into the ith element of a column by using `x[:, i]`. For example, if you want the top-left element of each element in `example_tensor`, which is the `0, 0` element of each matrix, you can write:
```
example_tensor[:, 0, 0]
```
# Initializing Tensors
There are many ways to create new tensors in PyTorch, but in this course, the most important ones are:
[`torch.ones_like`](https://pytorch.org/docs/master/generated/torch.ones_like.html): creates a tensor of all ones with the same shape and device as `example_tensor`.
```
torch.ones_like(example_tensor)
```
[`torch.zeros_like`](https://pytorch.org/docs/master/generated/torch.zeros_like.html): creates a tensor of all zeros with the same shape and device as `example_tensor`
```
torch.zeros_like(example_tensor)
```
[`torch.randn_like`](https://pytorch.org/docs/stable/generated/torch.randn_like.html): creates a tensor with every element sampled from a [Normal (or Gaussian) distribution](https://en.wikipedia.org/wiki/Normal_distribution) with the same shape and device as `example_tensor`
```
torch.randn_like(example_tensor)
```
Sometimes (though less often than you'd expect), you might need to initialize a tensor knowing only the shape and device, without a tensor for reference for `ones_like` or `randn_like`. In this case, you can create a $2x2$ tensor as follows:
```
torch.randn(2, 2, device='cpu') # Alternatively, for a GPU tensor, you'd use device='cuda'
```
# Basic Functions
There are a number of basic functions that you should know to use PyTorch - if you're familiar with numpy, all commonly-used functions exist in PyTorch, usually with the same name. You can perform element-wise multiplication / division by a scalar $c$ by simply writing `c * example_tensor`, and element-wise addition / subtraction by a scalar by writing `example_tensor + c`
Note that most operations are not in-place in PyTorch, which means that they don't change the original variable's data (However, you can reassign the same variable name to the changed data if you'd like, such as `example_tensor = example_tensor + 1`)
```
(example_tensor - 5) * 2
```
You can calculate the mean or standard deviation of a tensor using [`example_tensor.mean()`](https://pytorch.org/docs/stable/generated/torch.mean.html) or [`example_tensor.std()`](https://pytorch.org/docs/stable/generated/torch.std.html).
```
print("Mean:", example_tensor.mean())
print("Stdev:", example_tensor.std())
```
You might also want to find the mean or standard deviation along a particular dimension. To do this you can simple pass the number corresponding to that dimension to the function. For example, if you want to get the average $2\times2$ matrix of the $3\times2\times2$ `example_tensor` you can write:
```
example_tensor.mean(0)
# Equivalently, you could also write:
# example_tensor.mean(dim=0)
# example_tensor.mean(axis=0)
# torch.mean(example_tensor, 0)
# torch.mean(example_tensor, dim=0)
# torch.mean(example_tensor, axis=0)
```
PyTorch has many other powerful functions but these should be all of PyTorch functions you need for this course outside of its neural network module (`torch.nn`).
# PyTorch Neural Network Module (`torch.nn`)
PyTorch has a lot of powerful classes in its `torch.nn` module (Usually, imported as simply `nn`). These classes allow you to create a new function which transforms a tensor in specific way, often retaining information when called multiple times.
```
import torch.nn as nn
```
## `nn.Linear`
To create a linear layer, you need to pass it the number of input dimensions and the number of output dimensions. The linear object initialized as `nn.Linear(10, 2)` will take in a $n\times10$ matrix and return an $n\times2$ matrix, where all $n$ elements have had the same linear transformation performed. For example, you can initialize a linear layer which performs the operation $Ax + b$, where $A$ and $b$ are initialized randomly when you generate the [`nn.Linear()`](https://pytorch.org/docs/stable/generated/torch.nn.Linear.html) object.
```
linear = nn.Linear(10, 2)
example_input = torch.randn(3, 10)
example_output = linear(example_input)
example_output
```
## `nn.ReLU`
[`nn.ReLU()`](https://pytorch.org/docs/stable/generated/torch.nn.ReLU.html) will create an object that, when receiving a tensor, will perform a ReLU activation function. This will be reviewed further in lecture, but in essence, a ReLU non-linearity sets all negative numbers in a tensor to zero. In general, the simplest neural networks are composed of series of linear transformations, each followed by activation functions.
```
relu = nn.ReLU()
relu_output = relu(example_output)
relu_output
```
## `nn.BatchNorm1d`
[`nn.BatchNorm1d`](https://pytorch.org/docs/stable/generated/torch.nn.BatchNorm1d.html) is a normalization technique that will rescale a batch of $n$ inputs to have a consistent mean and standard deviation between batches.
As indicated by the `1d` in its name, this is for situations where you expects a set of inputs, where each of them is a flat list of numbers. In other words, each input is a vector, not a matrix or higher-dimensional tensor. For a set of images, each of which is a higher-dimensional tensor, you'd use [`nn.BatchNorm2d`](https://pytorch.org/docs/stable/generated/torch.nn.BatchNorm2d.html), discussed later on this page.
`nn.BatchNorm1d` takes an argument of the number of input dimensions of each object in the batch (the size of each example vector).
```
batchnorm = nn.BatchNorm1d(2)
batchnorm_output = batchnorm(relu_output)
batchnorm_output
```
## `nn.Sequential`
[`nn.Sequential`](https://pytorch.org/docs/stable/generated/torch.nn.Sequential.html) creates a single operation that performs a sequence of operations. For example, you can write a neural network layer with a batch normalization as
```
mlp_layer = nn.Sequential(
nn.Linear(5, 2),
nn.BatchNorm1d(2),
nn.ReLU()
)
test_example = torch.randn(5,5) + 1
print("input: ")
print(test_example)
print("output: ")
print(mlp_layer(test_example))
```
# Optimization
One of the most important aspects of essentially any machine learning framework is its automatic differentiation library.
## Optimizers
To create an optimizer in PyTorch, you'll need to use the `torch.optim` module, often imported as `optim`. [`optim.Adam`](https://pytorch.org/docs/stable/optim.html#torch.optim.Adam) corresponds to the Adam optimizer. To create an optimizer object, you'll need to pass it the parameters to be optimized and the learning rate, `lr`, as well as any other parameters specific to the optimizer.
For all `nn` objects, you can access their parameters as a list using their `parameters()` method, as follows:
```
import torch.optim as optim
adam_opt = optim.Adam(mlp_layer.parameters(), lr=1e-1)
```
## Training Loop
A (basic) training step in PyTorch consists of four basic parts:
1. Set all of the gradients to zero using `opt.zero_grad()`
2. Calculate the loss, `loss`
3. Calculate the gradients with respect to the loss using `loss.backward()`
4. Update the parameters being optimized using `opt.step()`
That might look like the following code (and you'll notice that if you run it several times, the loss goes down):
```
train_example = torch.randn(100,5) + 1
adam_opt.zero_grad()
# We'll use a simple loss function of mean distance from 1
# torch.abs takes the absolute value of a tensor
cur_loss = torch.abs(1 - mlp_layer(train_example)).mean()
cur_loss.backward()
adam_opt.step()
print(cur_loss)
```
## `requires_grad_()`
You can also tell PyTorch that it needs to calculate the gradient with respect to a tensor that you created by saying `example_tensor.requires_grad_()`, which will change it in-place. This means that even if PyTorch wouldn't normally store a grad for that particular tensor, it will for that specified tensor.
## `with torch.no_grad():`
PyTorch will usually calculate the gradients as it proceeds through a set of operations on tensors. This can often take up unnecessary computations and memory, especially if you're performing an evaluation. However, you can wrap a piece of code with `with torch.no_grad()` to prevent the gradients from being calculated in a piece of code.
## `detach():`
Sometimes, you want to calculate and use a tensor's value without calculating its gradients. For example, if you have two models, A and B, and you want to directly optimize the parameters of A with respect to the output of B, without calculating the gradients through B, then you could feed the detached output of B to A. There are many reasons you might want to do this, including efficiency or cyclical dependencies (i.e. A depends on B depends on A).
# New `nn` Classes
You can also create new classes which extend the `nn` module. For these classes, all class attributes, as in `self.layer` or `self.param` will automatically treated as parameters if they are themselves `nn` objects or if they are tensors wrapped in `nn.Parameter` which are initialized with the class.
The `__init__` function defines what will happen when the object is created. The first line of the init function of a class, for example, `WellNamedClass`, needs to be `super(WellNamedClass, self).__init__()`.
The `forward` function defines what runs if you create that object `model` and pass it a tensor `x`, as in `model(x)`. If you choose the function signature, `(self, x)`, then each call of the forward function, gets two pieces of information: `self`, which is a reference to the object with which you can access all of its parameters, and `x`, which is the current tensor for which you'd like to return `y`.
One class might look like the following:
```
class ExampleModule(nn.Module):
def __init__(self, input_dims, output_dims):
super(ExampleModule, self).__init__()
self.linear = nn.Linear(input_dims, output_dims)
self.exponent = nn.Parameter(torch.tensor(1.))
def forward(self, x):
x = self.linear(x)
# This is the notation for element-wise exponentiation,
# which matches python in general
x = x ** self.exponent
return x
```
And you can view its parameters as follows
```
example_model = ExampleModule(10, 2)
list(example_model.parameters())
```
And you can print out their names too, as follows:
```
list(example_model.named_parameters())
```
And here's an example of the class in action:
```
input = torch.randn(2, 10)
example_model(input)
```
# 2D Operations
You won't need these for the first lesson, and the theory behind each of these will be reviewed more in later lectures, but here is a quick reference:
* 2D convolutions: [`nn.Conv2d`](https://pytorch.org/docs/master/generated/torch.nn.Conv2d.html) requires the number of input and output channels, as well as the kernel size.
* 2D transposed convolutions (aka deconvolutions): [`nn.ConvTranspose2d`](https://pytorch.org/docs/master/generated/torch.nn.ConvTranspose2d.html) also requires the number of input and output channels, as well as the kernel size
* 2D batch normalization: [`nn.BatchNorm2d`](https://pytorch.org/docs/stable/generated/torch.nn.BatchNorm2d.html) requires the number of input dimensions
* Resizing images: [`nn.Upsample`](https://pytorch.org/docs/master/generated/torch.nn.Upsample.html) requires the final size or a scale factor. Alternatively, [`nn.functional.interpolate`](https://pytorch.org/docs/stable/nn.functional.html#torch.nn.functional.interpolate) takes the same arguments.
| github_jupyter |
```
import sounddevice as sd
import numpy as np
import scipy.signal as ss
import matplotlib.pyplot as plt
from scipy.io.wavfile import write, read
from numpy.fft import fft, ifft
from tqdm import tqdm
import time
def firfilter(sig, fs, low=None, high=None):
if low is None:
low = 0
if high is None:
high = fs/2
assert(low < high)
assert(high <= fs/2)
assert(sig.ndim == 1)
N = sig.size
f = np.arange(N) * fs / N
sigfft = fft(sig)
mask = ((f>=low) & (f<=high)) | ((f>=(fs-high)) & (f<=(fs-low)))
filtfft = np.zeros_like(sigfft)
filtfft[mask] = sigfft[mask]
return np.real(ifft(filtfft))
fs = 48000
sd.default.samplerate = fs
sd.default.channels = 1
```
## Microphone Recording
```
mic_duration = 10
# sd.play(chirp, fschirp)
samples = sd.rec(int(mic_duration*fs))
for i in tqdm(range(int(mic_duration*10))):
time.sleep(0.1)
sd.wait()
rxsig = np.squeeze(samples)
sd.play(rxsig, fs)
sd.stop()
```
## Saving Recording
```
np.save('probe_test', rxsig)
```
## Signal Preview
```
plt.plot(np.arange(3000)/fs, rxsig[:3000])
plt.show()
```
## Chirps
```
txfs, txsig = read('./audio/chirp.wav')
assert(txfs == fs)
txsig = txsig[:len(rxsig)]
txsig = txsig[:,0] # grab left channel only
nperseg = fs//50
noverlap = int(nperseg*0.9)
nfft = nperseg*10
print(nperseg, noverlap, nfft)
f, t, X = ss.spectrogram(txsig, fs, 'hann', nperseg=nperseg, noverlap=noverlap, nfft=nfft)
fwin = (f > 20e3) & (f < 23e3)
twin = (t > 0) & (t < 1)
plt.figure(figsize=(15,3))
plt.pcolormesh(t[twin], f[fwin], np.log(X[fwin][:,twin]))
plt.tight_layout()
plt.title("Tx Signal")
plt.show()
nperseg = fs//50
noverlap = int(nperseg*0.9)
nfft = nperseg*10
f, t, Y = ss.spectrogram(rxsig, fs, 'hann', nperseg=nperseg, noverlap=noverlap)
fwin = (f > 0e3) & (f < 23e3)
twin = (t > 0) & (t < 1)
plt.figure(figsize=(15,3))
plt.pcolormesh(t[twin], f[fwin], np.log(Y[fwin][:,twin]))
plt.tight_layout()
plt.title("Rx Signal")
plt.show()
nperseg = fs//50
noverlap = int(nperseg*0.9)
nfft = nperseg*10
z = np.roll(txsig, 4800//4) * rxsig
f, t, Z = ss.spectrogram(z, fs, 'hann', nperseg=nperseg, noverlap=noverlap, nfft=nfft)
fwin = (f > 19e3) & (f < 24e3)
twin = (t > 0) & (t < 10)
plt.figure(figsize=(15,3))
plt.pcolormesh(t[twin], f[fwin], np.log(Z[fwin][:,twin]))
plt.tight_layout()
plt.title("Tx Rx Product")
plt.show()
```
## Probes
```
txsig = np.load('probe_tx.npy')
nperseg = fs//100
noverlap = int(nperseg*0.9)
nfft = nperseg*10
print(nperseg, noverlap, nfft)
f, t, X = ss.spectrogram(txsig, fs, 'hann', nperseg=nperseg, noverlap=noverlap, nfft=nfft)
fwin = (f > 0e3) & (f < 28e3)
twin = (t > 0) & (t < 0.2)
plt.figure(figsize=(15,3))
plt.pcolormesh(t[twin], f[fwin], np.log(X[fwin][:,twin]+1e-10))
plt.tight_layout()
plt.title("Tx Signal")
plt.show()
nperseg = fs//100
noverlap = int(nperseg*0.9)
nfft = nperseg*10
f, t, Y = ss.spectrogram(rxsig, fs, 'hann', nperseg=nperseg, noverlap=noverlap)
fwin = (f > 0e3) & (f < 28e3)
twin = (t > 0) & (t < 0.2)
plt.figure(figsize=(15,3))
plt.pcolormesh(t[twin], f[fwin], np.log(Y[fwin][:,twin]))
plt.tight_layout()
plt.title("Rx Signal")
plt.show()
probe = np.load('probe.npy')
txcorr = ss.correlate(txsig, probe, mode='full')
rxcorr = ss.correlate(rxsig, probe, mode='full')
pk_tx, _ = ss.find_peaks(np.abs(txcorr), distance = fs/82)
pk_rx, _ = ss.find_peaks(np.abs(rxcorr), distance = fs/82)
offset = pk_rx[0] - pk_tx[0]
txcorr = np.roll(txcorr, offset)
txcorr = txcorr[:rxcorr.size]
txp, _ = ss.find_peaks(np.abs(txcorr), distance = fs/82)
rxp, _ = ss.find_peaks(np.abs(rxcorr), distance = fs/82)
t = np.arange(txcorr.size) / fs
plt.figure(figsize=(15,2.5))
plt.plot(t, txcorr)
plt.scatter(t[txp], txcorr[txp], c='orange')
plt.xlim(4, 4.05)
# plt.xlim(0.00625, 0.01865)
plt.tight_layout()
plt.title("Tx Probe Correlation")
plt.show()
plt.figure(figsize=(15,2.5))
plt.plot(t, rxcorr)
plt.scatter(t[rxp], rxcorr[rxp], c='orange')
plt.xlim(4, 4.05)
plt.ylim(-0.0015, 0.0015)
# plt.xlim(0.00625, 0.01865)
plt.tight_layout()
plt.title("Rx Probe Correlation")
plt.show()
hrx = ss.hilbert(rxcorr)
hmag = np.abs(hrx)
hphase = np.unwrap(np.angle(hrx))
hfreq = np.diff(hphase, prepend=hphase[1]-hphase[0]) / (2 * np.pi) * fs
plt.figure(figsize=(15,3))
plt.plot(t, rx)
plt.scatter(t[rxp], rx[rxp], c='orange')
plt.plot(t, hmag, c='green')
plt.xlim(1, 1+1/80)
plt.tight_layout()
plt.title("Rx Probe Correlation")
plt.show()
plt.figure(figsize=(15,3))
plt.plot(t, hfreq, c='purple')
plt.xlim(1, 1+1/80)
plt.ylim(18e3, 24e3)
plt.tight_layout()
plt.title("Instantaneous Frequency")
plt.show()
```
| github_jupyter |
# Spatially Assign Work
In this example, assignments will be assigned to specific workers based on the city district that it falls in. A layer in ArcGIS Online representing the city districts in Palm Springs will be used.
* Note: This example requires having Arcpy or Shapely installed in the Python environment.
### Import ArcGIS API for Python
Import the `arcgis` library and some modules within it.
```
from datetime import datetime
from arcgis.gis import GIS
from arcgis.geometry import Geometry
from arcgis.mapping import WebMap
from arcgis.apps import workforce
from datetime import datetime
```
### Connect to organization and Get the Project
Let's connect to ArcGIS Online and get the Project with assignments.
```
gis = GIS("https://arcgis.com", "workforce_scripts")
item = gis.content.get("c765482bd0b9479b9104368da54df90d")
project = workforce.Project(item)
```
### Get Layer of City Districts
Let's get the layer representing city districts and display it.
```
districts_layer = gis.content.get("8a79535e0dc04410b5564c0e45428a2c").layers[0]
districts_map = gis.map("Palm Springs, CA", zoomlevel=10)
districts_map.add_layer(districts_layer)
districts_map
```
### Add Assignments to the Map
```
districts_map.add_layer(project.assignments_layer)
```
### Create a spatially enabled dataframe of the districts
```
districts_df = districts_layer.query(as_df=True)
```
### Get all of the unassigned assignments
```
assignments = project.assignments.search("status=0")
```
### Assign Assignments Based on Which District They Intersect¶
Let's fetch the districts layer and query to get all of the districts. Then, for each unassigned assignment intersect the assignment with all districts to determine which district it falls in. Assignments in district 10 should be assigned to James. Assignments in district 9 should be assigned to Aaron. Finally update all of the assignments using "batch_update".
```
aaron = project.workers.get(user_id="aaron_nitro")
james = project.workers.get(user_id="james_Nitro")
for assignment in assignments:
contains = districts_df["SHAPE"].geom.contains(Geometry(assignment.geometry))
containers = districts_df[contains]
if not containers.empty:
district = containers['ID'].iloc[0]
if district == 10:
assignment.worker = james
assignment.status = "assigned"
assignment.assigned_date = datetime.utcnow()
elif district == 9:
assignment.worker = aaron
assignment.status = "assigned"
assignment.assigned_date = datetime.utcnow()
assignments = project.assignments.batch_update(assignments)
```
### Verify Assignments are Assigned
```
webmap = gis.map("Palm Springs", zoomlevel=11)
webmap.add_layer(project.assignments_layer)
webmap
```
| github_jupyter |
# Quantum Generative Adversarial Networks
## Introduction
Generative [adversarial](gloss:adversarial) networks (GANs) [[1]](https://arxiv.org/abs/1406.2661) have swiftly risen to prominence as one of the most widely-adopted methods for unsupervised learning, with showcased abilities in photo-realistic image generation. Given the success of classical GANs, a natural question is whether this success will translate into a quantum computing GAN. In this page, we explore the theory behind quantum generative adversarial networks (QGANs), as well as the practice of implementing one in Qiskit to learn a [Gaussian distribution](gloss:gaussian-distribution). Lastly, we end off with a discussion around the potential use cases of quantum generative adversarial networks and link to relevant research for those who want to read further.
## Classical generative models (theory)
### Generative models
Until recently, the success of supervised learning models has completely overshadowed their generative counterparts. So much so, that the popularity of these supervised models might make it difficult to even conceptualize another approach to machine learning.
The supervised approach, which feels intuitive to us by now, tries to make accurate predictions on new data, demonstrating that it has learned some underlying relations of the dataset. Generative models are different. Instead of focusing on key relationships between input data and labels, they learn to model the underlying distribution holistically, allowing it to generate new data samples. It's the difference between telling apart cats and dogs, and generating completely new images of cats and dogs. The latter is a richer, but also more difficult task.
Why is it more difficult? Adequately discriminating between given data can often be achieved through picking up on a few tell-tale features (like whiskers don't belong on eyes) which help form the strong decision boundaries in the high dimensional space. Consequently, machine learning researchers take great interest in generative models as these learning tasks seem to stab at a deeper notion of learning—trying to reproduce the underlying *creator* function.
So given a pool of training data, the goal of a generative model is to learn/reproduce the probability distribution that generated them. A particularly eye-catching application of GANs is generating [high-resolution visuals](https://thispersondoesnotexist.com/) or [composing music](https://magenta.tensorflow.org/gansynth). Below is a generated image of a fake cat.

<!-- ::: q-block.exercise -->
### Quick quiz
<!-- ::: q-quiz(goal="qml-qgan-0") -->
<!-- ::: .question -->
What would be the most appropriate learning task for a generative model?
<!-- ::: -->
<!-- ::: .option(correct) -->
1. Producing images of handwritten digits
<!-- ::: -->
<!-- ::: .option -->
2. Classifying incoming emails as 'spam' or 'not spam'
<!-- ::: -->
<!-- ::: .option -->
3. Predicting stock prices
<!-- ::: -->
<!-- ::: .option -->
4. Recommending optimal movies
<!-- ::: -->
<!-- ::: -->
<!-- ::: -->
### Generative adversarial networks
A particular class of generative models—generative adversarial networks (GANs)—have witnessed a boom in popularity since they were first proposed in 2014 by Goodfellow I., *et al.* [[1]](https://arxiv.org/abs/1406.2661). To understand the quantum analogue, we first briefly discuss the concept behind classical generative adversarial networks.
Briefly put, GANs use a pair of neural networks pitted against each other—the generator and the discriminator.
### The generator
The generator's primary aim is to create fake data samples that are convincing enough for the discriminator to label them as real. With each training step, the generator improves at this, until it has near complete overlap to the fixed distribution of real data.
To allow the generator to explore a rich space of output non-deterministically, a random noise vector drawn from a [latent space](gloss:latent-space) is fed into the generator as input (usually sampled from a Gaussian distribution). The generator succeeds once it learns to map most points in the latent space (Gaussian noise) onto convincing fake data samples fitting the real distribution.
*At the start of training, the latent space is a meaningless n-dimensional Gaussian distribution. But, as the generator evolves, the generator learns to map the noise vectors in the latent space to valid data in the objective dataset.*
### The discriminator
The discriminator receives data samples from both the generator and the real distribution (not knowing which is which), and its task is to correctly classify the input data samples as fake or real.
Note how the discriminator's objective is directly opposed to its' counterpart. While the discriminator tries to minimize the probability of misclassifying the fake data as real, the generator tries to maximize it.

### Convergence
The GAN finishes training once the generator consistently generates convincing data samples indistinguishable to the real data distribution, leaving the discriminator unable to reasonably decipher between the two. Formally, this point is referred to as the Nash equilibrium (from game theory), at which the generator produces data that corresponds to the real probability distribution, and the *trained* discriminator resorts to guessing between fake or real (50% accuracy).
A common analogy between GANs and art theft brings the concept into frame. The generator is often seen as a fake artist trying to produce paintings identical to those found in the museum. The art expert's objective (discriminator) is to tell apart the generator's fake art from the real art.
Applied to this analogy, the discriminator assesses the paintings' authenticity while the generator creates the best fakes to fool it. The zero-sum game pushes the two networks to constantly one-up each other. Each improvement of the generator in creating convincing data, is bested by the discriminator's update in improved classification, and vice versa.

<!-- ::: q-block.exercise -->
### Quick quiz
<!-- ::: q-quiz(goal="qml-qgan-1") -->
<!-- ::: .question -->
Once the GAN reaches Nash Equilibrium...
<!-- ::: -->
<!-- ::: .option(correct) -->
1. The discriminator randomly guesses fake/real with equal probability
<!-- ::: -->
<!-- ::: .option -->
2. The generator returns to producing noise
<!-- ::: -->
<!-- ::: .option -->
3. The GAN reaches a common failure mode, and the training process must be restarted
<!-- ::: -->
<!-- ::: .option -->
4. The discriminator guesses that all samples are real
<!-- ::: -->
<!-- ::: -->
<!-- ::: -->
## Quantum Generative Adversarial Networks (theory)
In 2018, two companion papers (Ref. [[2]](https://journals.aps.org/pra/abstract/10.1103/PhysRevA.98.012324), [[3]](https://journals.aps.org/prl/abstract/10.1103/PhysRevLett.121.040502)) brought the idea of GANs to the quantum domain. On a high level, quantum generative adversarial networks (QGANs) equip either the discriminator, the generator, or both with [parameterized quantum circuits](./paramterized-quantum-circuits) with the goal of learning some quantum or classical data. In this chapter, we discuss the fully quantum version of QGANs (quantum generator, quantum discriminator), keeping in mind that the broader principles apply across other types of QGANs.

Image from Ref. [5](https://arxiv.org/abs/1901.00848)
There are many analogous concepts, specifically with the adversarial training, between GANs and QGANs. Most importantly, the training structure of GANs largely persists to QGANs. We alternately train the generator & discriminator circuit parameters, while freezing the other's parameters. Through this, the quantum generator learns the target quantum state by proxy of the discriminator's signals, similar to GANs. It's proven that the [stable equilibrium](https://en.wikipedia.org/wiki/Nash_equilibrium) of the quantum adversarial game also occurs when the generator produces data identical to a target distribution [[3]](https://journals.aps.org/prl/abstract/10.1103/PhysRevLett.121.040502).
The objective with a fully quantum QGAN is for the quantum generator to reproduce a desired state $|\psi\rangle$, using of course an adversarial training strategy. Similar to other variational quantum algorithms, the quantum generator moves towards this through an iterative update to its parameters directed by a classical optimizer. However, in the case of a QGAN, the generator's cost function landscape itself evolves and becomes better as the discriminator improves at recognizing real samples. Let's look at the general circuit schematics of what we will be building.


It is worth noting that although we only consider one register each for both the generator and discriminator, we could also add an [auxiliary](gloss:auxiliary) "workspace" register to both the generator and discriminator.
The two circuits illustrated above show the generator concatenated with the discriminator and the unitary loading the target data concatenated with the discriminator.
The discriminator acts on the output from the generator, respectively, the target data unitary, as well as an additional qubit $|0\rangle$. Finally, the discriminator classification takes place by measuring the last qubit. If the outcome corresponds to $|0\rangle$ respectively $|1\rangle$ the input data is being classified as real or fake respectively.
Looking at the first circuit diagram shows us how real data samples are fed into the discriminator. Since we are dealing with a fully quantum QGAN, we need to somehow encode this data of ours into a quantum state to feed into the discriminator. To that end, we prepare the real data through a unitary $\class{u-sub-r}{U_R}$ such that we define:
$|\text{Data}_R\rangle= U_R|0\rangle^{\otimes n}$
which is then fed to the parameterized discriminator $\class{u-sub-d}{U}_{D(\class{theta-d}{\vec{\theta_D}})}$ (possibly containing an auxiliary register), and then measured to arrive at the discriminator's score on real data.
It is worth noting that $U_{D(\class{theta-d}{\vec{\theta_D}})}$ contains several 2-qubit entangling gates to "transmit" relevant features of the real data to the discriminator's register (workspace). Formalized, the unitary evolution is:
$\class{u-sub-d}{U}_{D(\class{theta-d}{\vec{\theta_D}})}(|\text{Data}_R\rangle \otimes |0\rangle)$
where $\class{theta-d}{\vec{\theta_D}}$ is the parameter vector that is updated through a classical optimizer to minimize the [expectation value](gloss:expectation-value) of the last qubit (equivalent to maximizing the probability of the discriminator classifying real data as $|\text{real}\rangle$).
In the second circuit, a generated [wave function](gloss:wave-function) aimed to mimic the real one is fed into the discriminator. In other words, the fake quantum state prepared by $U_{G(\class{theta-g}{\vec{\theta_G}})}$, parameterized by $\class{theta-g}{\vec{\theta_G}}$, is applied on the initial state $|0^{\otimes n}\rangle$, giving us:
$$|\text{Data}_G\rangle = U_{G(\class{theta-g}{\vec{\theta_G}})}|0^{\otimes n}\rangle$$
$|\text{Data}_G\rangle$ is then fed to the discriminator parameterized by $\class{theta-d}{\vec{\theta_D}}$. So taking the expectation value of the observable $I^{\otimes n}Z$ on $U_{D(\class{theta-d}{\vec{\theta_D}})}(|\text{Data}_G\rangle \otimes |0\rangle)$ gives us the discriminator's score on fake data.
It is worth reiterating that $\langle \text{fake} | Z | \text{fake} \rangle = 1$, meaning the discriminator "believes" a given sample to be wholly fake, the expectation value $Z$ with repect to the last qubit will be equal to 1. It then follows naturally that the discriminator would want to correctly "assign" $|0\rangle$ to fake samples and $|1\rangle$ to real samples. The inverse is true for the generator. For it, the optimal scenario would be if the discriminator was completely convinced that its generated quantum state was real, thereby assigning it a $|1\rangle$. We can formalize these adversarial incentives into the following [minimax](gloss:minimax) decision rule (adapted from [reference 2](https://journals.aps.org/pra/abstract/10.1103/PhysRevA.98.012324)):
$\underset{\class{theta-g}{\vec{\theta_G}}}{\min}\underset{\class{theta-d}{\vec{\theta_D}}}{\max}
\hspace{3pt} \Bigg(\class{pr}{\text{Pr}}\bigg(\class{d-disc}{D}(\class{theta-d}{\vec{\theta_D}}, R) = |\text{real}\rangle\bigg) + \hspace{2pt} \class{pr}{\text{Pr}}\bigg(\class{d-disc}{D}(\class{theta-d}{\vec{\theta_D}}, G(\class{theta-g}{\vec{\theta_G}})) = |\text{fake}\rangle\bigg)\Bigg)$
<!-- ::: q-block.exercise -->
### Quick quiz
<!-- ::: q-quiz(goal="qml-qgan-2") -->
<!-- ::: .question -->
Quiz question: How do we obtain the probability of a given data sample being real, as assigned by the discriminator? Let $\langle Z \rangle$ be the expectation value of $Z$ with respect to the last qubit.
<!-- ::: -->
<!-- ::: .option -->
1. $\langle Z\rangle + 1$
<!-- ::: -->
<!-- ::: .option -->
2. $\langle Z\rangle + 1/2$
<!-- ::: -->
<!-- ::: .option(correct) -->
3. $\frac{\langle Z \rangle + 1}{2}$
<!-- ::: -->
<!-- ::: .option -->
4. $2^{\langle Z \rangle}$
<!-- ::: -->
<!-- ::: -->
Hint: the Z-expectation value is bounded between $[-1, 1]$
<!-- ::: -->
## Full implementation I
### Learning a 2 qubit Bell state
Equipped with the adequate theoretical foundation, we can now build an actual QGAN to learn the 2 qubit Bell state through Qiskit! First, we import the standard libraries.
```
import numpy as np
from qiskit import QuantumCircuit, Aer, execute
from qiskit.visualization import plot_histogram
```
### Defining the real distribution
The 2 qubit Bell state is a maximally entangled quantum state, the specific statevector we're interested to reproduce is
$|\psi\rangle = \frac{1}{\sqrt{2}}(|00\rangle + |11\rangle) $
Which can be constructed by applying a CNOT gate followed by a Hadamard.
```
# Number of qubits needed to model real distribution
REAL_DIST_NQUBITS = 2
real_circuit = QuantumCircuit(REAL_DIST_NQUBITS)
real_circuit.h(0)
real_circuit.cx(0, 1);
```
<!--- TODO: Widget: mini composer Have them compose Bell state of interest defined above. Stops when constructs properly --->
### Defining the variational quantum generator and discriminator
We now define the generator ansatz.
Given the primal nature of existing research into QGANs, the community has not yet settled into any optimal generator or discriminator ansatzes. On that note, most of the [hyperparameters](gloss:hyperparameter) chosen in quantum machine learning are still driven by loose [heuristics](gloss:heuristics), and there's a lot still to be explored.
It's worth mentioning that whichever ansatz we choose for the generator, it must have enough [capacity](gloss:capacity) and be [expressible](gloss:expressible) enough to fully reproduce the real quantum state $|\psi\rangle$ defined earlier. So, although the ansatz used here is a little arbitrary, we are confident that it is plenty expressive for the Bell state we are trying to model.
```
# Importing qiskit machine learning parameters
from qiskit.circuit import Parameter, ParameterVector
```
Here, we use the `TwoLocal` class to create an ansatz for the variational quantum generator with single qubit gates $RY$ and $RZ$, paired with the engtangling gate $CZ$.
```
from qiskit.circuit.library import TwoLocal
generator = TwoLocal(REAL_DIST_NQUBITS,
# Parameterized single qubit rotations
['ry', 'rz'],
'cz', # Entangling gate
'full', # Entanglement structure: all to all
reps=2, # Number of layers
parameter_prefix='θ_g',
name='Generator')
generator = generator.decompose() # decompose into standard gates
generator.draw()
```
### Variational quantum discriminator
We now define the ansatz for the discriminator. In this case, instead of using [`TwoLocal`](https://qiskit.org/documentation/stubs/qiskit.circuit.library.TwoLocal.html#qiskit.circuit.library.TwoLocal), we create a custom ansatz with a [`ParameterVector`](https://qiskit.org/documentation/stubs/qiskit.circuit.ParameterVector.html).
```
disc_weights = ParameterVector('θ_d', 12)
discriminator = QuantumCircuit(3, name="Discriminator")
discriminator.barrier()
discriminator.h(0)
discriminator.rx(disc_weights[0], 0)
discriminator.ry(disc_weights[1], 0)
discriminator.rz(disc_weights[2], 0)
discriminator.rx(disc_weights[3], 1)
discriminator.ry(disc_weights[4], 1)
discriminator.rz(disc_weights[5], 1)
discriminator.rx(disc_weights[6], 2)
discriminator.ry(disc_weights[7], 2)
discriminator.rz(disc_weights[8], 2)
discriminator.cx(0, 2)
discriminator.cx(1, 2)
discriminator.rx(disc_weights[9], 2)
discriminator.ry(disc_weights[10], 2)
discriminator.rz(disc_weights[11], 2)
discriminator.draw()
```
### Compiling the QGAN
With all of our components in place, we now construct the two circuits forming the QGAN. The first feeds generated quantum state into the discriminator and the second is comprised of the discriminator applied on the real state. *It is easy to see how this circuit fulfills the general schematic we outlined earlier.*
```
N_GPARAMS = generator.num_parameters
N_DPARAMS = discriminator.num_parameters
# Need extra qubit for the discriminator
gen_disc_circuit = QuantumCircuit(REAL_DIST_NQUBITS+1)
gen_disc_circuit.compose(generator, inplace=True)
gen_disc_circuit.compose(discriminator, inplace=True)
gen_disc_circuit.draw()
```
A very natural question to ask at this point is: why isn't there any noise fed into the generator? As you may recall, in the classical GAN, the latent space was an essential ingredient. If there was no noise for the classical GAN, then it would be impossible for the generator to represent a complete distribution since with each update to its parameters, it would be restricted to output one sample given its deterministic nature. But consider, that in the quantum case, since we are feeding the whole 'fake' wave function directly to the discriminator, the role that noise would play is much less obvious. With or without noise, the variational quantum generator is capable of directly modelling the wave function of interest, so long as the ansatz is of adequate capacity.
With that said, there may still be benefits to equipping the variational quantum generator with a latent space of its own, in fact [reference 5](https://arxiv.org/abs/1901.00848) presents a method to allow the quantum generator to model continuous distributions using a latent space as input. But to keep it simple, we will still omit feeding noise into the varational quantum generator.
Below, we define the parameterized circuit linking the target distribution with the variational discriminator.
```
real_disc_circuit = QuantumCircuit(REAL_DIST_NQUBITS+1)
real_disc_circuit.compose(real_circuit, inplace=True)
real_disc_circuit.compose(discriminator, inplace=True)
real_disc_circuit.draw()
```
### Constructing the cost function
Remember the minimax decision rule we formulated earlier,
$\underset{\class{theta-g}{\vec{\theta_G}}}{\min}\underset{\class{theta-d}{\vec{\theta_D}}}{\max}
\hspace{3pt} \Bigg(\class{pr}{\text{Pr}}\bigg(\class{d-theta}{D(\class{theta-d}{\vec{\theta_D}}, R)} = |\text{real}\rangle\bigg) + \hspace{2pt} \class{pr}{\text{Pr}}\bigg(D(\class{theta-d}{\vec{\theta_D}}, G(\class{theta-g}{\vec{\theta_G}})) = |\text{fake}\rangle\bigg)\Bigg)$
Constructing a loss function for both the discriminator and generator is now trivial. Starting with the discriminator, we have
$\text{Cost}_D = \class{pr}{\text{Pr}}\bigg(D(\class{theta-d}{\vec{\theta_D}}, G(\class{theta-g}{\vec{\theta_G}})) = |\text{real}\rangle\bigg) - \class{pr}{\text{Pr}}\bigg(D(\class{theta-d}{\vec{\theta_D}}, R) = |\text{real}\rangle\bigg)$.
Minimizing this entails maximizing the probability of correctly classifying real data while minimizing the probability of mistakenly classifying fake data.
As a hallmark of vanilla GANs, the generator’s cost function will simply be the negation of the discriminator’s cost, where the optimal strategy is to maximize the probability of the discriminator misclassifying fake data. We omit the term concerning the real quantum state since the generator's weights leave no effect on it.
$\text{Cost}_G = - \class{pr}{\text{Pr}}\bigg(D(\class{theta-d}{\vec{\theta_D}}, G(\class{theta-g}{\vec{\theta_G}})) = |\text{real}\rangle\bigg)$
We now implement the above cost functions. It is worth noticing that after accessing the respective probabilities of each basis state, we arrive at the total probability of a given sample being classified as $|\text{real}\rangle = |1\rangle$ by summing over each basis state that satisfies $|XX1\rangle$ (any state with last qubit measured as $|1\rangle$). However, do note the reverse ordering given Qiskit's [endian](gloss:endian) resulting in $|1XX\rangle$.
```
# We'll use Statevector to retrieve statevector of given circuit
from qiskit.quantum_info import Statevector
import tensorflow as tf
def generator_cost(gen_params: tf.Tensor) -> float:
# .numpy() method extracts numpy array from TF tensor
curr_params = np.append(disc_params.numpy(),
gen_params.numpy())
state_probs = Statevector(gen_disc_circuit
.bind_parameters(curr_params)
).probabilities()
# Get total prob of measuring |1> on q2
prob_fake_true = np.sum(state_probs[0b100:])
cost = -prob_fake_true
return cost
def discriminator_cost(disc_params: tf.Tensor) -> float:
# .numpy() method extracts numpy array from TF tensor
curr_params = np.append(disc_params.numpy(),
gen_params.numpy())
gendisc_probs = Statevector(gen_disc_circuit
.bind_parameters(curr_params)
).probabilities()
realdisc_probs = Statevector(real_disc_circuit.
bind_parameters(disc_params.numpy())
).probabilities()
# Get total prob of measuring |1> on q2
prob_fake_true = np.sum(gendisc_probs[0b100:])
# Get total prob of measuring |1> on q2
prob_real_true = np.sum(realdisc_probs[0b100:])
cost = prob_fake_true - prob_real_true
return cost
```
We now define a helper function to calculate the [Kullback-Leibler divergence](https://en.wikipedia.org/wiki/Kullback%E2%80%93Leibler_divergence) between the model and target distribution. This is a common metric used to track the generator's progress while training since it effectively measures the distance between two distributions. A lower KL divergence indicates that the two distributions are similar, with a KL of 0 implying equivalence.
```
def calculate_KL(model_distribution: dict, target_distribution: dict):
"""Gauge model performance using Kullback Leibler Divergence"""
KL = 0
for bitstring, p_data in target_distribution.items():
if np.isclose(p_data, 0, atol=1e-8):
continue
if bitstring in model_distribution.keys():
KL += (p_data * np.log(p_data)
- p_data * np.log(model_distribution[bitstring]))
else:
KL += p_data * np.log(p_data) - p_data * np.log(1e-6)
return KL
```
### CircuitQNN
For simplicity, we use the [`CircuitQNN`](https://qiskit.org/documentation/machine-learning/stubs/qiskit_machine_learning.neural_networks.CircuitQNN.html) that compiles the parameterized circuit and handles calculation of the gradient recipes. Calling the `forward()` method also directly outputs the probability state vectors of the circuit.
```
from qiskit.utils import QuantumInstance
from qiskit_machine_learning.neural_networks import CircuitQNN
# define quantum instances (statevector and sample based)
qi_sv = QuantumInstance(Aer.get_backend('aer_simulator_statevector'))
# specify QNN to update generator weights
gen_qnn = CircuitQNN(gen_disc_circuit, # parameterized circuit
# frozen input arguements (discriminator weights)
gen_disc_circuit.parameters[:N_DPARAMS],
# differentiable weights (generator weights)
gen_disc_circuit.parameters[N_DPARAMS:],
sparse=True, # returns sparse probability vector
quantum_instance=qi_sv)
# specify QNNs to update discriminator weights
disc_fake_qnn = CircuitQNN(gen_disc_circuit, # parameterized circuit
# frozen input arguments (generator weights)
gen_disc_circuit.parameters[N_DPARAMS:],
# differentiable weights (discrim. weights)
gen_disc_circuit.parameters[:N_DPARAMS],
sparse=True, # get sparse probability vector
quantum_instance=qi_sv)
disc_real_qnn = CircuitQNN(real_disc_circuit, # parameterized circuit
[], # no input parameters
# differentiable weights (discrim. weights)
gen_disc_circuit.parameters[:N_DPARAMS],
sparse=True, # get sparse probability vector
quantum_instance=qi_sv)
```
Here, we use [TensorFlow Keras](gloss:tensorflow-keras) to create an ADAM optimizer instance for both the generator and the discriminator. The ADAM optimizer is a widely-used optimizer in classical machine learning that uses momentum-based gradient updates. It is known to far outperform vanilla gradient descent.
To us the Keras optimizer, we must store the weights as TF variables, which can be easily done through the `tf.Variable` method. We converting back into a `np.ndarray` using the `.numpy()` instance method on the `tf.Variable`.
```
import tensorflow as tf
import pickle # to serialize and deserialize variables
# Initialize parameters
init_gen_params = np.random.uniform(low=-np.pi,
high=np.pi,
size=(N_GPARAMS,))
init_disc_params = np.random.uniform(low=-np.pi,
high=np.pi,
size=(N_DPARAMS,))
gen_params = tf.Variable(init_gen_params)
disc_params = tf.Variable(init_disc_params)
```
Let's look at our starting point for the generator created from random weights.
```
init_gen_circuit = generator.bind_parameters(init_gen_params)
init_prob_dict = Statevector(init_gen_circuit).probabilities_dict()
import matplotlib.pyplot as plt
fig, ax1 = plt.subplots(1, 1, sharey=True)
ax1.set_title("Initial generator distribution")
plot_histogram(init_prob_dict, ax=ax1)
# Initialize Adam optimizer from Keras
generator_optimizer = tf.keras.optimizers.Adam(learning_rate=0.02)
discriminator_optimizer = tf.keras.optimizers.Adam(learning_rate=0.02)
```
### Training
```
# Initialize variables to track metrics while training
best_gen_params = tf.Variable(init_gen_params)
gloss = []
dloss = []
kl_div = []
```
There are a few important points on the following training logic:
1. The discriminator's weights are updated fivefold for each generator update. When dealing with classical GANs, it's also not uncommon to see an unbalanced number of training steps between the two networks. In this case, we arrive at a 5:1 ratio as a best practice through trial and error.
2. The `backward()` method of `CircuitQNN` returns the gradients with respect to each weight for each basis state for each batch. In other words, the return shape of `CircuitQNN.backward(...)[1].todense()` is `(num_batches, num_basis_states, num_weights)`. So to arrive at the gradient for $\text{Cost}_D$, we first sum over all the gradients for each basis state satisfying $|1XX\rangle$, and then subtract them according to the $\text{Cost}_D$ function. Recall that the linearity the derivative allows us to distribute it as implemented below.
3. Due to the instability of GAN training, we store the best generator parameters.
```
table_headers = "Epoch / Generator cost / Discriminator cost / KL Div"
print(table_headers)
for epoch in range(100):
"""Quantum discriminator parameter updates"""
d_steps = 5 # N discriminator updates per generator update
for disc_train_step in range(d_steps):
# Partial derivatives wrt θ_D
d_fake = disc_fake_qnn.backward(gen_params, disc_params
)[1].todense()[0, 0b100:]
d_fake = np.sum(d_fake, axis=0)
d_real = disc_real_qnn.backward([], disc_params
)[1].todense()[0, 0b100:]
d_real = np.sum(d_real, axis=0)
# Recall Cost_D structure
grad_dcost = [d_fake[i] - d_real[i] for i in range(N_DPARAMS)]
grad_dcost = tf.convert_to_tensor(grad_dcost)
# Update disc params with gradient
discriminator_optimizer.apply_gradients(zip([grad_dcost],
[disc_params]))
# Track discriminator loss
if disc_train_step % d_steps == 0:
dloss.append(discriminator_cost(disc_params))
"""Quantum generator parameter updates"""
for gen_train_step in range(1):
# Compute partial derivatives of prob(fake|true) wrt each
# generator weight
grads = gen_qnn.backward(disc_params, gen_params)
grads = grads[1].todense()[0][0b100:]
# Recall Cost_G structure and the linearity of
# the derivative operation
grads = -np.sum(grads, axis=0)
grads = tf.convert_to_tensor(grads)
# Update gen params with gradient
generator_optimizer.apply_gradients(zip([grads], [gen_params]))
gloss.append(generator_cost(gen_params))
"""Tracking KL and saving best performing generator weights"""
# Create test circuit with updated gen parameters
gen_checkpoint_circuit = generator.bind_parameters(gen_params.numpy())
# Retrieve probability distribution of current generator
gen_prob_dict = Statevector(gen_checkpoint_circuit
).probabilities_dict()
# Constant real probability distribution
real_prob_dict = Statevector(real_circuit).probabilities_dict()
current_kl = calculate_KL(gen_prob_dict, real_prob_dict)
kl_div.append(current_kl)
if np.min(kl_div) == current_kl:
# New best
# serialize+deserialize to simply ensure zero links
best_gen_params = pickle.loads(pickle.dumps(gen_params))
if epoch % 10 == 0:
# print table for every 10 epochs
for header, val in zip(table_headers.split('/'),
(epoch, gloss[-1], dloss[-1], kl_div[-1])):
print(f"{val:.3g} ".rjust(len(header)+1), end="")
print()
```
### Results visualized
We plot the collected metrics to examine how the QGAN learned.
```
fig, (loss, kl) = plt.subplots(2, sharex=True,
gridspec_kw={'height_ratios': [0.75, 1]},
figsize=(6,4))
fig.suptitle('QGAN training stats')
fig.supxlabel('Training step')
loss.plot(range(len(gloss)), gloss, label="Generator loss")
loss.plot(range(len(dloss)), dloss, label="Discriminator loss",
color="C3")
loss.legend()
loss.set(ylabel='Loss')
kl.plot(range(len(kl_div)), kl_div, label="KL Divergence (zero is best)",
color="C1")
kl.set(ylabel='KL Divergence')
kl.legend()
fig.tight_layout();
# Create test circuit with new parameters
gen_checkpoint_circuit = generator.bind_parameters(
best_gen_params.numpy())
gen_prob_dict = Statevector(gen_checkpoint_circuit).probabilities_dict()
real_prob_dict = Statevector(real_circuit).probabilities_dict() # constant
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(8,3))
plot_histogram(gen_prob_dict, ax=ax1)
ax1.set_title("Trained generator distribution")
plot_histogram(real_prob_dict, ax=ax2)
ax2.set_title("Real distribution")
fig.tight_layout()
```
With just 100 [epoch](gloss:epoch)s, we see that the generator has approximated the Bell state $|\psi\rangle$ quite well!
Note that reproducing results from above may take a few runs. Due to the fragile nature of training two competing models at once, generative adversarial networks as a whole are notorious for failing to converge—and it is only amplified for QGANs given the lack of best practices. GANs (QGANs inclusive) often suffer from vanishing gradients, often caused by a discriminator that is too good.
Another drawback of the adversarial training structure—albeit less prevalant with fully quantum QGANs—is mode collapse, the failure mode of GANs when the generator and discriminator get caught in a cat and mouse chase. The generator spams a certain sample that has a tendency to fool the discriminator, to which the discriminator adapts to over time, but the generator swiftly follows up yet again with another sample. [Learn more on remedies to GAN failure modes](https://developers.google.com/machine-learning/gan/problems).
## Full implementation II
### Learning a normal distribution with OpFlowQNN
In the following section, we build a QGAN to learn a 3 qubit normal distribution, while changing a few previous methods along the way.
1. Learning a more complex distribution
2. Using [OpFlowQNN](gloss:opflowqnn) to retrieve expectation values directly
3. Amalgamating the discriminator output qubit to the generator's register
### Defining the real distribution
Here we define the real distribution using the `qiskit_finance` module and the generator ansatz that'll be used to model the [Gaussian](gloss:gaussian-distribution).
```
from qiskit_finance.circuit.library import NormalDistribution
REAL_DIST_NQUBITS = 3
real_circuit = NormalDistribution(REAL_DIST_NQUBITS, mu=0, sigma=0.15)
real_circuit = real_circuit.decompose().decompose().decompose()
real_circuit.draw()
```
### Defining the variational quantum discriminator and generator
```
generator = TwoLocal(REAL_DIST_NQUBITS,
# Parameterized single qubit rotations
['ry', 'rz'],
'cz', # Entangling gate
'full', # Entanglement structure - all to all
reps=2, # Number of layers
parameter_prefix='θ_g',
name='Generator')
generator = generator.decompose()
generator.draw()
```
Now, we define a similar ansatz as before for the discriminator, just with the output qubit defined on `q2` instead of `q3` as one might have expected from the previous example. It's important to note that the qubit used to measure $\class{}{\langle Z\rangle_{\text{out}}}$ is largely irrelevant since the discriminator ansatz can universally transform any given quantum state to a desired one. Previously, we defined the output qubit to be one a separate register to make it more intuitive but there exists no true justification with respect to QGAN performance.
```
disc_weights = ParameterVector('θ_d', 12)
discriminator = QuantumCircuit(REAL_DIST_NQUBITS, name="Discriminator")
discriminator.barrier()
discriminator.h(0)
discriminator.rx(disc_weights[0], 0)
discriminator.ry(disc_weights[1], 0)
discriminator.rz(disc_weights[2], 0)
discriminator.h(1)
discriminator.rx(disc_weights[3], 1)
discriminator.ry(disc_weights[4], 1)
discriminator.rz(disc_weights[5], 1)
discriminator.h(2)
discriminator.rx(disc_weights[6], 2)
discriminator.ry(disc_weights[7], 2)
discriminator.rz(disc_weights[8], 2)
discriminator.cx(1,2)
discriminator.cx(0,2)
discriminator.rx(disc_weights[9], 2)
discriminator.ry(disc_weights[10], 2)
discriminator.rz(disc_weights[11], 2)
discriminator.draw()
```
Then we construct the complete circuits.
```
N_GPARAMS = generator.num_parameters
N_DPARAMS = discriminator.num_parameters
```
### Compiling the QGAN
```
gen_disc_circuit = QuantumCircuit(REAL_DIST_NQUBITS)
gen_disc_circuit.compose(generator, inplace=True)
gen_disc_circuit.compose(discriminator, inplace=True)
gen_disc_circuit.draw()
real_disc_circuit = QuantumCircuit(REAL_DIST_NQUBITS)
real_disc_circuit.compose(real_circuit, inplace=True)
real_disc_circuit.compose(discriminator, inplace=True)
real_disc_circuit.draw()
```
### OpflowQNN
We employ the [`OpflowQNN`](https://qiskit.org/documentation/machine-learning/stubs/qiskit_machine_learning.neural_networks.OpflowQNN.html) from Qiskit which takes a (parameterized) operator and leverages Qiskit's gradient framework to complete the backward passes. The operator defined here is equivalent to the expectation value of $Z$ with respect to the last qubit.
```
from qiskit.providers.aer import QasmSimulator
from qiskit.opflow import (StateFn, PauliSumOp, ListOp,
Gradient, AerPauliExpectation)
from qiskit_machine_learning.neural_networks import OpflowQNN
# set method to calculcate expected values
expval = AerPauliExpectation()
# define gradient method
gradient = Gradient()
# define quantum instances (statevector)
qi_sv = QuantumInstance(Aer.get_backend('aer_simulator_statevector'))
# Circuit wave function
gen_disc_sfn = StateFn(gen_disc_circuit)
real_disc_sfn = StateFn(real_disc_circuit)
# construct operator to retrieve Pauli Z expval of the last qubit
H1 = StateFn(PauliSumOp.from_list([('ZII', 1.0)]))
# combine operator and circuit to objective function
gendisc_op = ~H1 @ gen_disc_sfn
realdisc_op = ~H1 @ real_disc_sfn
# construct OpflowQNN with the two operators, the input parameters,
# the weight parameters, the expected value, and quantum instance.
"""|fake> => |0> => 1 ; |real> => |1> => -1"""
gen_opqnn = OpflowQNN(gendisc_op,
# input parameters (discriminator weights)
gen_disc_circuit.parameters[:N_DPARAMS],
# differentiable weights (generator weights)
gen_disc_circuit.parameters[N_DPARAMS:],
expval,
gradient,
qi_sv) # gen wants to to minimize this expval
disc_fake_opqnn = OpflowQNN(gendisc_op,
# input parameters (generator weights)
gen_disc_circuit.parameters[N_DPARAMS:],
# differentiable weights (discrim. weights)
gen_disc_circuit.parameters[:N_DPARAMS],
expval,
gradient,
qi_sv) # disc wants to maximize this expval
disc_real_opqnn = OpflowQNN(realdisc_op,
[],
# differentiable weights (discrim. weights)
gen_disc_circuit.parameters[:N_DPARAMS],
expval,
gradient,
qi_sv) # disc wants to minimize this expval
```
First we initialize the training parameters and define the optimizer
```
### START
init_gen_params = tf.Variable(np.random.uniform(low=-np.pi,
high=np.pi,
size=(N_GPARAMS)))
init_disc_params = tf.Variable(np.random.uniform(low=-np.pi,
high=np.pi,
size=(N_DPARAMS)))
gen_params = init_gen_params
disc_params = init_disc_params
generator_optimizer = tf.keras.optimizers.Adam(learning_rate=0.02)
discriminator_optimizer = tf.keras.optimizers.Adam(learning_rate=0.02)
```
### Reconstructing the cost function
Now we construct the training logic. There are a few key differences to the cost function here that impacts the gradient rule. Since a forward pass now returns the direct expectation value and not a probability statevector, it's important to remind ourselves that $\langle \text{real} |Z| \text{real} \rangle = -1$ and $\langle \text{fake} |Z| \text{fake} \rangle = 1$.
Applying similar logic to before, we arrive at the intuition that the discriminator would want to maximize $\langle \text{fake} |Z| \text{fake} \rangle $ when fed fake data and minimize $\langle \text{real} |Z| \text{real}\rangle$ when receiving the real quantum state. In contrast to that, the generator wishes to minimize $\langle \text{fake} |Z| \text{fake}\rangle $, which is akin to the maximizing the probability of the discriminator classifying fake samples as $|\text{real}\rangle = |1\rangle$
We now cement these ideas into the following minimax decision rule defined by the proper expectation values. Let $\rho^{DR}$ and $\rho^{GR}$ be the density matrix representations of $\bigg(U_{D(\class{theta-d}{\vec{\theta_D}})}U_R|0\rangle^{\otimes n+1}\bigg)$ and $\bigg(U_{D(\class{theta-d}{\vec{\theta_D}})} U_{G(\class{theta-g}{\vec{\theta_G}})}|0\rangle^{\otimes n+1}\bigg)$, respectively. Also recall that the expectation value of $\class{sigma-p}{\sigma^P}$ with respect to an arbitrary density matrix $\rho$ is defined as $\text{tr}(\rho \sigma^P)$ (relevant [chapter](/course/quantum-hardware/density-matrix)). While remembering the linearity of the trace operation, we arrive at
$\underset{\class{theta-g}{\vec{\theta_G}}}{\min} \hspace{2pt} \underset{\class{theta-d}{\vec{\theta_D}}}{\max} \hspace{3pt} \text{tr}\bigg(\big(\rho^{DG}(\class{theta-d}{\vec{\theta_D}}, \class{theta-g}{\vec{\theta_G}}) - \rho^{DR}(\class{theta-d}{\vec{\theta_D}})\Big) Z\bigg)$
Which leads us to the following cost functions (optimum is minimum),
$\text{Cost}_D(\class{theta-d}{\vec{\theta_D}}, \class{theta-g}{\vec{\theta_G}}) = \text{tr}\bigg(Z\rho^{DR}(\class{theta-d}{\vec{\theta_D}}) \bigg) - \text{tr}\bigg(Z\rho^{DG}(\class{theta-d}{\vec{\theta_D}}, \class{theta-g}{\vec{\theta_G}})\bigg)$
$\text{Cost}_G(\class{theta-d}{\vec{\theta_D}}, \class{theta-g}{\vec{\theta_G}}) = \text{tr}\bigg(Z \rho^{DG}(\class{theta-d}{\vec{\theta_D}}, \class{theta-g}{\vec{\theta_G}}) \bigg)$
Meaning that the gradients are,
$\nabla _ {\class{theta-d}{\vec{\theta_D}}}\ \text{Cost}_D(\class{theta-d}{\vec{\theta_D}}, \class{theta-g}{\vec{\theta_G}}) = \nabla _ {\class{theta-d}{\vec{\theta_D}}}\ \text{tr}\bigg(Z\rho^{DR}(\class{theta-d}{\vec{\theta_D}}) \bigg) - \nabla _ {\class{theta-d}{\vec{\theta_D}}}\ \text{tr}\bigg(Z\rho^{DG}(\class{theta-d}{\vec{\theta_D}}, \class{theta-g}{\vec{\theta_G}})\bigg)$
$\nabla _ {\class{theta-g}{\vec{\theta_G}}} \ \text{Cost}_G(\class{theta-d}{\vec{\theta_D}}, \class{theta-g}{\vec{\theta_G}}) = \nabla _ {\class{theta-g}{\vec{\theta_G}}}\ \text{tr}\bigg(Z \rho^{DG}(\class{theta-d}{\vec{\theta_D}}, \class{theta-g}{\vec{\theta_G}}) \bigg)$
and we're complete! We now have all the information needed to implement it since the `OpFlowQNN.backward()` method computes the constituent gradients for us. Let's implement this.
*Keep in mind that the above formulations rely upon our initial definition that $|\text{real}\rangle = |1\rangle$ and $|\text{fake}\rangle = |0\rangle$.*
### Training
```
best_gen_params = init_gen_params
gloss = []
dloss = []
kl_div = []
table_headers = "Epoch / Gen. cost / Discrim. cost / KL Div / New best?"
print(table_headers)
for epoch in range(300):
d_steps = 5
"""Quantum discriminator parameter update"""
for disc_train_step in range(d_steps):
grad_dcost_fake = disc_fake_opqnn.backward(gen_params,
disc_params)[1][0,0]
grad_dcost_real = disc_real_opqnn.backward([],
disc_params)[1][0,0]
grad_dcost = grad_dcost_real - grad_dcost_fake # as formulated above
grad_dcost = tf.convert_to_tensor(grad_dcost)
# update disc_params
discriminator_optimizer.apply_gradients(zip([grad_dcost],
[disc_params]))
if disc_train_step % d_steps == 0:
dloss.append(discriminator_cost(disc_params))
"""Quantum generator parameter update"""
for gen_train_step in range(1):
# as formulated above
grad_gcost = gen_opqnn.backward(disc_params,
gen_params)[1][0,0]
grad_gcost = tf.convert_to_tensor(grad_gcost)
# update gen_params
generator_optimizer.apply_gradients(zip([grad_gcost],
[gen_params]))
gloss.append(generator_cost(gen_params))
"""Tracking KL and saving best performing generator weights"""
# Create test circuit with updated gen parameters
gen_checkpoint_circuit = generator.bind_parameters(gen_params.numpy())
# Retrieve probability distribution of current generator
gen_prob_dict = Statevector(gen_checkpoint_circuit
).probabilities_dict()
# Constant real probability distribution
real_prob_dict = Statevector(real_circuit).probabilities_dict()
current_kl = calculate_KL(gen_prob_dict, real_prob_dict)
kl_div.append(current_kl)
new_best = (np.min(kl_div) == current_kl)
if new_best:
# Store new best generator weights
# serialize+deserialize to just zero links
best_gen_params = pickle.loads(pickle.dumps(gen_params))
if epoch % 30 == 0:
# print table
for header, val in zip(table_headers.split('/'),
(epoch, gloss[-1], dloss[-1],
kl_div[-1], new_best)):
print(f"{val:.3g} ".rjust(len(header)+1), end="")
print()
```
### Results visualized
```
import matplotlib.pyplot as plt
fig, (loss, kl) = plt.subplots(2, sharex=True,
gridspec_kw={'height_ratios': [0.75, 1]},
figsize=(6,4))
fig.suptitle('QGAN training stats')
fig.supxlabel('Training step')
loss.plot(range(len(gloss)), gloss, label="Generator loss")
loss.plot(range(len(dloss)), dloss, label="Discriminator loss",
color="C3")
loss.legend()
loss.set(ylabel='Loss')
kl.plot(range(len(kl_div)), kl_div, label="KL Divergence (zero is best)",
color="C1")
kl.set(ylabel='KL Divergence')
kl.legend()
fig.tight_layout();
# Create test circuit with new parameters
gen_checkpoint_circuit = generator.bind_parameters(best_gen_params.numpy())
gen_prob_dict = Statevector(gen_checkpoint_circuit).probabilities_dict()
real_prob_dict = Statevector(real_circuit).probabilities_dict() # constant
fig, (ax1, ax2) = plt.subplots(1, 2, sharey=True, figsize=(9,3))
plot_histogram(gen_prob_dict, ax=ax1)
ax1.set_title("Trained generator distribution")
plot_histogram(real_prob_dict, ax=ax2)
ax2.set_title("Real distribution")
ax2.set_ylim([0,.5])
fig.tight_layout()
```
Impressive!
<!-- ::: q-block.exercise -->
### Quick quiz
Drag and drop the lines of pseudocode into their correct order to complete the program.
q-drag-and-drop-code
.line For n:=0 to N_EPOCHS do:
.line For d:=0 to num_disc_steps do:
.line Compute Z expectation value of d_out qubit when fed fake and real data
.line.md Update $\vec{\theta}_{D}$ according to $\nabla_{\vec{\theta}_{D}}\text{Cost}_D(\class{theta-d}{\vec{\theta_D}},\class{theta-g}{\vec{\theta_G}})$ using ADAM optimizer
.line For g:=0 to num_gen_steps do
.line Compute Z-expectation value of d_out qubit when fed fake data
.line.md Update $\vec{\theta}_{G}$ according to $\nabla_{\vec{\theta}_G}\text{Cost}_G(\class{theta-d}{\vec{\theta_D}},\class{theta-g}{\vec{\theta_G}})$ using ADAM optimizer
.line Compute KL divergence between G and R
.line If current KL divergence is the lowest yet do
.line Save current generator weights
<!-- ::: -->
## Potential applications
The development of QGANs is still emerging, so there remains much more research to be done on potential applications. However, there is hope that QGANs will enable sampling and manipulation of classically intractable probability distributions (difficult to sample from classically).
One particularly interesting application of QGANs is efficient, approximate data loading. In order to see a quantum advantage in data processing - using, e.g., quantum amplitude estimation [6] — we need to load the input data onto a quantum state. However, loading classical data into a quantum circuit is often expensive, and generally even exponentially expensive [7, 8]. Therefore, the data loading complexity can easily impair any potential quantum advantage.
As shown in reference [4], QGANs offer an interesting approach to efficiently learn and load approximations of generic probability distributions, as demonstrated in this [tutorial](https://qiskit.org/documentation/machine-learning/tutorials/04_qgans_for_loading_random_distributions.html). Once the probability distribution is loaded into a quantum state, a quantum algorithm such as quantum amplitude estimation can process the data. As shown in this [tutorial](https://qiskit.org/documentation/tutorials/finance/10_qgan_option_pricing.html), this workflow may then be used to, e.g., price options with a potential quadratic quantum speedup.
Additionally, in quantum chemistry, quantum computers are believed to have an intrinsic advantage in being able to represent and manipulate correlated [fermionic](gloss:fermionic) states (molecules). A natural question one could ask is: given an adequate ansatz, could QGANs be used to generate new types of molecules that fit the mould of an inputted set of materials/molecules possibly obtained through VQE? That would involve extending QGANs into the conditional realm (inputting a conditional label to both the generator and discriminator, see [conditional GANs](https://arxiv.org/abs/1411.1784) but as of now, it remains an open question.
<!-- ::: q-block.exercise -->
### Try it
To extend the ideas you've just learned, create a QGAN to learn a 3 qubit normal distribution but with a classical discriminator. [Reference 4](https://arxiv.org/abs/1904.00043) will be helpful. You may use the same generator ansatz we've used above, but ensure the discriminator's neural network is adequately sized to match the quantum generator's power.
[Try in IBM Quantum Lab](https://quantum-computing.ibm.com/lab)
<!-- ::: -->
## References
1. I. J. Goodfellow, J. Pouget-Abadie, M. Mirza, B. Xu, D.Warde-Farley, S. Ozair, A. Courville, and Y. Bengio, in *Proceedings of the 27th International Conference on Neural Information Processing Systems* (MIT Press, Cambridge, MA, 2014), Vol. 2, pp. 2672–2680, [arXiv:1406.2661](https://arxiv.org/abs/1406.2661).
2. P.-L. Dallaire-Demers, & N. Killoran, *Quantum generative adversarial networks,* Phys. Rev. A 98, 012324 (2018), [doi.org:10.1103/PhysRevA.98.012324](https://doi.org/10.1103/PhysRevA.98.012324), [arXiv:1804.08641](https://arxiv.org/abs/1804.08641)
3. S. Lloyd, & C. Weedbrook, *Quantum generative adversarial learning*. Phys. Rev. Lett. 121, 040502 (2018), [doi.org:10.1103/PhysRevLett.121.040502](https://doi.org/10.1103/PhysRevLett.121.040502), [arXiv:1804.09139](https://arxiv.org/abs/1804.09139)
4. C. Zoufal, A. Lucchi, and S. Woerner, *Quantum generative adversarial networks for learning and loading random distributions,* npj Quantum Information, 5, Article number: 103 (2019), [doi.org/10.1038/s41534-019-0223-2](https://doi.org/10.1038/s41534-019-0223-2), [arXiv:1904.00043](https://arxiv.org/abs/1904.00043)
5. J. Romero, A. Aspuru-Guzik, *Variational quantum generators: Generative adversarial quantum machine learning for continuous distributions* (2019), [arxiv.org:1901.00848](https://arxiv.org/abs/1901.00848)
6. Brassard, G., Hoyer, P., Mosca, M. & Tapp, A. *Quantum amplitude amplification and estimation*. Contemp. Math. 305, 53–74 (2002), [doi.org/10.1090/conm/305/05215](http://www.ams.org/books/conm/305/), [arXiv:quant-ph/0005055](https://arxiv.org/abs/quant-ph/0005055)
7. L. K. Grover. *Synthesis of quantum superpositions by quantum computation*. Phys. Rev. Lett., 85, (2000), [doi.org/10.1103/PhysRevLett.85.1334](https://link.aps.org/doi/10.1103/PhysRevLett.85.1334)
8. M. Plesch and ˇC. Brukner. *Quantum-state preparation with universal gate decompositions*. Phys. Rev. A, 83, (2010), [doi.org/10.1103/PhysRevA.83.032302](https://doi.org/10.1103/PhysRevA.83.032302)
```
import qiskit.tools.jupyter
%qiskit_version_table
```
| github_jupyter |
## Working with filter pipelines
This Jupyter notebook explains the workflow of setting up and configuring a ground point filtering pipeline. This is an advanced workflow for users that want to define their own filtering workflows. For basic use, preconfigured pipelines are (or rather: will be) provided by `adaptivefiltering`. As always, we first need to import our library:
```
import adaptivefiltering
```
Also, we need to load at least one data set which we will use to interactively preview our filter settings. Note that for a good interactive experience with no downtimes, you should restrict your datasets to a reasonable size (see the [Working with datasets](datasets.ipynb) notebook for how to do it).
```
dataset = adaptivefiltering.DataSet(filename="500k_NZ20_Westport.laz")
```
### Filtering backends
`adaptivefiltering` does not implement its own ground point filtering algorithms. Instead, algorithms from existing packages are accessible through a common interface. Currently, the following backends are available:
* [PDAL](https://pdal.io/): The Point Data Abstraction Library is an open source library for point cloud processing.
* [OPALS](https://opals.geo.tuwien.ac.at/html/stable/index.html) is a proprietary library for processing Lidar data. It can be tested freely for datasets <1M points.
* [LASTools](https://rapidlasso.com/) has a proprietary tool called `lasground_new` that can be used for ground point filtering.
PDAL is always available when using `adaptivefiltering` and is used internally for many tasks that are not directly related to ground point filtering. In order to enable the OPALS backend, `adaptivefiltering` needs to be given the information where your OPALS installation (potentially including your license key) is located. This can either be done by setting the environment variable `OPALS_DIR` or by setting the path at runtime:
```
adaptivefiltering.set_opals_directory("/path/to/opals")
```
Similarly, you can set the path to your installation of LASTools either through the environment variable `LASTOOLS_DIR` or at runtime:
```
adaptivefiltering.set_lastools_directory("/path/to/lastools")
```
Please note that LASTools only ships Windows binaries. Therefore, you will need [Wine](https://www.winehq.org/) installed on your system to successfully use the LASTools backend.
### Configuring a filter pipeline
The main pipeline configuration is done by calling the `pipeline_tuning` function with your dataset as the parameter. This will open the interactive user interface which waits for your user input until you hit the *Finalize* button. The configured filter is then accessible as the Python object `pipeline`:
```
pipeline = adaptivefiltering.pipeline_tuning(dataset)
```
If you want to inspect multiple data sets in parallel while tuning a pipeline, you can do so by passing a list of datasets to the `pipeline_tuning` function. Note that `adaptivefiltering` does currently not parallelize the execution of filter pipeline execution which may have a negative impact on wait times while tuning with multiple parameters.
```
pipeline2 = adaptivefiltering.pipeline_tuning(datasets=[dataset, dataset])
```
### Storing and reloading filter pipelines
Pipeline objects can be stored on disk with the `save_filter` command from `adaptivefiltering`. We typically use the extension `json` for filters. It stands for *JavaScript Object Notation* and is a widely used format for storing custom data structures:
```
adaptivefiltering.save_filter(pipeline, "myfilter.json")
```
The appropriate counterpart is `load_filter`, which restores the pipeline object from a file:
```
old_pipeline = adaptivefiltering.load_filter("myfilter.json")
```
A filter pipeline loaded from a file can be edited using the `pipeline_tuning` command by passing it to the function. As always, the pipeline object returned by `pipeline_tuning` will be a new object - no implicit changes of the loaded pipeline object will occur:
```
edited_pipeline = adaptivefiltering.pipeline_tuning(dataset, pipeline=old_pipeline)
```
### Applying filter pipelines to data
Pipeline objects can also be used to transform data sets by applying the ground point filtering algorithms. This is one of the core tasks of the `adaptivefiltering` library, but this will rarely be done in this manual fashion, as we will provide additional interfaces for (locally adaptive) application of filter pipelines:
```
filtered = pipeline.execute(dataset)
```
| github_jupyter |
# Launch Angle
We're going to analyze 2019 Statcast data with a specific focus on launch angle. We'll be loading it from .csv but previous notebooks have demonstrated use of *pybaseball* module to pull the data directly from *baseball savant*.
1. First we load the data and python libraries
2. We're going to look at the batted results broken down by launch angle in a series of simple bar charts
3. Statistical distribution of launch angle in 2019
4. Relationship of launch angle and wOBA and/or home runs
This is all to produce supporting information for my blog article which can be viewed at cgutwein.github.io
```
## libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
% matplotlib inline
## import data
statcast_data = pd.read_csv('../../blog_posts/statcast_pitches_2019.csv')
from matplotlib.patches import Circle
```
## Ted Williams Reference
The article mentions Ted Williams approximately of 10 degrees launch angle. Here we'll filter the data for:
* batted results
* launch angle <= 10 degrees
We'll get a second look with the data above 10 degrees as well. For information pertaining to the Statcast data, this link will be helpful: https://baseballsavant.mlb.com/csv-docs
```
event_list = list(statcast_data['events'].unique())
batted_list = ['field_out',
'grounded_into_double_play',
'home_run',
'single',
'double',
'force_out',
'field_error',
'double_play',
'sac_fly',
'triple',
'fielders_choice_out',
'fielders_choice',
'sac_fly_double_play',
'triple_play']
# Create the dictionary
batted_dictionary ={'field_out': 'out',
'grounded_into_double_play': 'out',
'single':'single',
'double':'double',
'force_out':'out',
'field_error':'out',
'double_play':'out',
'sac_fly':'out',
'triple':'triple',
'home_run': 'home_run',
'fielders_choice_out':'out',
'fielders_choice':'out',
'sac_fly_double_play':'out',
'triple_play':'out'}
# filter data
batted_statcast_data = statcast_data[statcast_data['events'].isin(batted_list)].copy()
# put all outs and errors in one category
batted_statcast_data['events'] = batted_statcast_data['events'].map(batted_dictionary)
batted_statcast_data['angle10'] = batted_statcast_data['launch_angle'] > 10
# sort data to look better for chart
mapping = {'out': 0,
'single': 1,
'double': 2,
'triple': 3,
'home_run': 4}
key = batted_statcast_data['events'].map(mapping)
# Bar chart - launch angle by result
# Setting FiveThirtyEight style
plt.style.use('fivethirtyeight')
# Setting size of our plot
fig, ax = plt.subplots(figsize=(8,6))
ax1 = sns.countplot(x="events", hue="angle10", data=batted_statcast_data.iloc[key.argsort()])
# Y axis past 0 & above 100 -- grid line will pass 0 & 100 marker
plt.ylim(-5,50000)
# Bolded horizontal line at y=0
#ax1.axhline(y=0, color='#414141', linewidth=1.5, alpha=.5)
# Y-labels to only these
ax.set_yticks([0, 10000,20000, 30000, 40000])
ax.set_yticklabels(labels=['0', '10000', '20000', '30000', '40000'], fontsize=14, color='#414141')
# X-labels and changing label names
ax.set_xticklabels(['Out', 'Single', 'Double', 'Triple','Homer'],fontsize=14, color='#414141')
# Title text
ax.text(x=-1, y=49000, s="Is 10 the magic number?", fontsize=18.5, fontweight='semibold', color='#414141')
# Subtitle text
ax.text(x=-1, y=46500, s='Batted ball results above and below 10 degree launch angle', fontsize=16.5, color='#414141')
ax.set_ylabel('')
ax.set_xlabel('')
# legend
ax.legend(["< 10 degree launch angle", "> 10 degree launch angle"], loc=7)
# Line at bottom for signature line
ax1.text(x = -1, y = -7000,
s = ' ©Chet Gutwein Source: Baseball Savant ',
fontsize = 14, color = '#f0f0f0', backgroundcolor = '#414141');
plt.savefig('ted.png')
```
## Launch vs. Exit Velo
The next chart presents launch angle vs. exit velocity. We somewhat want to re-create the charts of past articles by the Washington Post, etc. that show the "sweet spot" for home runs that exists within this relationship. For our version, we'll want to have home runs and outs presented in different colors.
```
# Scatter chart - launch angle vs exit velo
# new variable - is homer?
batted_statcast_data['is_homer'] = batted_statcast_data['events'] == 'home_run'
# Setting FiveThirtyEight style
plt.style.use('fivethirtyeight')
# Setting size of our plot
fig, ax = plt.subplots(figsize=(12,9))
ax1 = plt.scatter(x=batted_statcast_data["launch_angle"], y=batted_statcast_data["launch_speed"], c=batted_statcast_data['is_homer'], alpha=0.02, cmap='rainbow')
# Y axis past 0 & above 100 -- grid line will pass 0 & 100 marker
plt.ylim(-5,150)
# Bolded horizontal line at y=0
#ax1.axhline(y=0, color='#414141', linewidth=1.5, alpha=.5)
# Y-labels to only these
ax.set_yticks([0, 25, 50, 75, 100, 125])
ax.set_yticklabels(labels=['0', '25', '50', '75', '100', '125'], fontsize=14, color='#414141')
# X-labels and changing label names
#ax.set_xticklabels(['Out', 'Single', 'Double', 'Triple','Homer'],fontsize=14, color='#414141')
# Title text
ax.text(x=-99, y=140, s="Launch angle vs. exit velocity", fontsize=18.5, fontweight='semibold', color='#414141')
# Subtitle text
ax.text(x=-99, y=135, s='Where are the homers being it?', fontsize=16.5, color='#414141')
ax.set_ylabel('')
ax.set_xlabel('Launch Angle')
# legend
#ax.legend(["< 10 degree launch angle", "> 10 degree launch angle"], loc=7)
# Line at bottom for signature line
ax.text(x = -110, y = -25,
s = ' ©Chet Gutwein Source: Baseball Savant ',
fontsize = 14, color = '#f0f0f0', backgroundcolor = '#414141');
plt.savefig('purple_cloud.png')
```
Further examination of launch angle vs. wOBA, we need to group by position player ID. We'll then plot the mean launch angle vs. the mean wOBA. It'd be nice to get a fit line and also annotate a player or two, which might become useful later.
```
batted_grouped = batted_statcast_data.groupby(by='batter').mean()
batted_counts = batted_statcast_data.groupby(by='batter').count()
valid_batter_list = batted_counts[batted_counts['events'] >= 50].index
batted_grouped_trimmed = batted_grouped[batted_grouped.index.isin(valid_batter_list)]
annotated_list = [545361, 593934, 608336, 446359]
batted_grouped_trimmed['is_annotated'] = batted_grouped_trimmed.index.isin(annotated_list)
print(len(batted_grouped))
print(len(batted_grouped_trimmed))
print(sum(batted_grouped_trimmed['is_annotated']))
# Setting FiveThirtyEight style
plt.style.use('fivethirtyeight')
# Setting size of our plot
fig, ax = plt.subplots(figsize=(8,6))
ax1 = plt.scatter(x=batted_grouped_trimmed["launch_angle"], y=batted_grouped_trimmed["woba_value"], alpha=0.3, c=batted_grouped_trimmed['is_annotated'], cmap='rainbow')#, c=batted_grouped_trimmed["woba_value"], cmap='rainbow')
# Y axis past 0 & above 100 -- grid line will pass 0 & 100 marker
plt.ylim(-0.1,0.8)
# Bolded horizontal line at y=0
#ax1.axhline(y=0, color='#414141', linewidth=1.5, alpha=.5)
# Y-labels to only these
ax.set_yticks([0, 0.2, 0.4, 0.6])
ax.set_yticklabels(labels=['0', '0.2', '0.4', '0.6'], fontsize=14, color='#414141')
# X-labels and changing label names
ax.set_xticks([-10, 0, 10, 20,30])
ax.set_xticklabels(['-10', '0', '10', '20', '30'],fontsize=14, color='#414141')
# Title text
ax.text(x=-15, y=0.9, s="Launch angle vs. wOBA", fontsize=18.5, fontweight='semibold', color='#414141')
# Subtitle text
ax.text(x=-15, y=0.85, s='Batters average wOBA on batted balls', fontsize=16.5, color='#414141')
ax.set_ylabel('wOBA')
ax.set_xlabel('Player Mean Launch Angle on batted balls')
# Annotations
ax.annotate('Joey Gallo', xy=(batted_grouped_trimmed.loc[608336]['launch_angle'],batted_grouped_trimmed.loc[608336]['woba_value']),
xytext=((batted_grouped_trimmed.loc[608336]['launch_angle']-6,batted_grouped_trimmed.loc[608336]['woba_value'])))
#ax.add_artist(Circle(((batted_grouped_trimmed.loc[656541]['launch_angle'],batted_grouped_trimmed.loc[656541]['woba_value'])), 0.01, edgecolor='red',fill=False))
#Migel Sano
ax.annotate('Miguel Sano', xy=(batted_grouped_trimmed.loc[593934]['launch_angle'],batted_grouped_trimmed.loc[593934]['woba_value']),
xytext=((batted_grouped_trimmed.loc[593934]['launch_angle']-7,batted_grouped_trimmed.loc[593934]['woba_value'])))
#ax.add_artist(Circle(((batted_grouped_trimmed.loc[593934]['launch_angle'],batted_grouped_trimmed.loc[593934]['woba_value'])), 0.01, edgecolor='red',fill=False))
#Mike Trout
ax.annotate('Mike Trout', xy=(batted_grouped_trimmed.loc[545361]['launch_angle'],batted_grouped_trimmed.loc[545361]['woba_value']),
xytext=((batted_grouped_trimmed.loc[545361]['launch_angle']+0.5,batted_grouped_trimmed.loc[545361]['woba_value'])))
#ax.add_artist(Circle(((batted_grouped_trimmed.loc[446263]['launch_angle'],batted_grouped_trimmed.loc[446263]['woba_value'])), 0.01, edgecolor='red',fill=False))
#Zach Cozart
ax.annotate('Zac Cozart', xy=(batted_grouped_trimmed.loc[446359]['launch_angle'],batted_grouped_trimmed.loc[446359]['woba_value']),
xytext=((batted_grouped_trimmed.loc[446359]['launch_angle']+0.5,batted_grouped_trimmed.loc[446359]['woba_value'])))
# legend
#ax.legend(["< 10 degree launch angle", "> 10 degree launch angle"], loc=7)
# Line at bottom for signature line
ax.text(x = -15, y = -0.3,
s = ' ©Chet Gutwein Source: Baseball Savant ',
fontsize = 14, color = '#f0f0f0', backgroundcolor = '#414141');
plt.savefig('batter_mean.png')
batted_grouped_trimmed[(batted_grouped_trimmed['woba_value'] < 0.2) & (batted_grouped_trimmed['launch_angle'] > 20)]
#id_map = pd.read_csv('../mlb/data/PLAYERIDMAP.csv')
id_map[id_map['MLBID'] == 446359]
#id_map[id_map['PLAYERNAME'] == 'Joey Gallo']['MLBID']
batted_grouped_trimmed.loc[656541]['woba_value']
```
For the final chart, we're going to plot batted balls by Mike Trout in Red and Zach Cozart in blue using the same launch angle vs. exit velocity. This time, for the homer's we'll use a shape or outline the circles as indication.
```
# Setting FiveThirtyEight style
plt.style.use('fivethirtyeight')
# Setting size of our plot
fig, ax = plt.subplots(figsize=(12,9))
x1 = batted_statcast_data[(batted_statcast_data["batter"] == 545361) & (batted_statcast_data["is_homer"] == False)]["launch_angle"]
y1 = batted_statcast_data[(batted_statcast_data["batter"] == 545361) & (batted_statcast_data["is_homer"] == False)]["launch_speed"]
ax1 = plt.scatter(x=x1, y=y1, c='blue', alpha=0.2)
x1b = batted_statcast_data[(batted_statcast_data["batter"] == 545361) & (batted_statcast_data["is_homer"] == True)]["launch_angle"]
y1b = batted_statcast_data[(batted_statcast_data["batter"] == 545361) & (batted_statcast_data["is_homer"] == True)]["launch_speed"]
ax1b = plt.scatter(x=x1b, y=y1b, c='blue', alpha=1, marker="*", edgecolor='pink')
x2 = batted_statcast_data[(batted_statcast_data["batter"] == 446359) & (batted_statcast_data["is_homer"] == False)]["launch_angle"]
y2 = batted_statcast_data[(batted_statcast_data["batter"] == 446359) & (batted_statcast_data["is_homer"] == False)]["launch_speed"]
ax2 = plt.scatter(x=x2, y=y2, c='red', alpha=0.2)
x2b = batted_statcast_data[(batted_statcast_data["batter"] == 446359) & (batted_statcast_data["is_homer"] == True)]["launch_angle"]
y2b = batted_statcast_data[(batted_statcast_data["batter"] == 446359) & (batted_statcast_data["is_homer"] == True)]["launch_speed"]
ax2b = plt.scatter(x=x2b, y=y2b, c='red', alpha=1, marker="*", edgecolor='pink')
# Y axis past 0 & above 100 -- grid line will pass 0 & 100 marker
plt.ylim(-5,150)
# Bolded horizontal line at y=0
#ax1.axhline(y=0, color='#414141', linewidth=1.5, alpha=.5)
# Y-labels to only these
ax.set_yticks([0, 25, 50, 75, 100, 125])
ax.set_yticklabels(labels=['0', '25', '50', '75', '100', '125'], fontsize=14, color='#414141')
# X-labels and changing label names
#ax.set_xticklabels(['Out', 'Single', 'Double', 'Triple','Homer'],fontsize=14, color='#414141')
# Title text
ax.text(x=-75, y=140, s="Launch angle vs. exit velocity", fontsize=18.5, fontweight='semibold', color='#414141')
# Subtitle text
ax.text(x=-75, y=135, s='High launch angle Angels, with different results', fontsize=16.5, color='#414141')
ax.set_ylabel('')
ax.set_xlabel('Launch Angle')
# legend
ax.legend(["Mike Trout", "Mike Trout home runs","Zach Cozart", "Zach Cozart home runs"], loc=3)
# Line at bottom for signature line
ax.text(x = -84, y = -25,
s = ' ©Chet Gutwein Source: Baseball Savant ',
fontsize = 14, color = '#f0f0f0', backgroundcolor = '#414141');
plt.savefig('trout_cozart.png')
sum(batted_grouped_trimmed['launch_angle'] > 20)
```
| github_jupyter |
Instructions: click restart and run all above. Figures will show once the entire notebook has finished running (will take a few minutes)
```
import sys
sys.path.append('..')
import numpy as np
import matplotlib.pyplot as plt
import glob
import ipywidgets as widgets
from tqdm import tqdm
%matplotlib notebook
from membranequant import ImageQuant, direcslist, load_image, af_subtraction, save_img
```
# Assessing inter-embryo reference profile variation
A key assumption of the model is that common reference profiles can be used for all embryos, which we believe to be valid because embryo geometry and imaging conditions are conserved. However, I have previously only performed reference profile calibration on single embryos or multiple embryos in batch, which doesn't give any information about possible inter-embryo variation. Here, I perform calibration on multiple single embryos to test for this
## Cytoplasmic reference profile inter-embryo variation
Performing cytoplasmic reference calibration on a few single embryos to see how variable it is between embryos. Using PKC-3 in a par-3 mutant as before.
### Import data
```
path = '../test_datasets/dataset3_pkc3_par3mutant'
paths = direcslist(path)
images = [load_image(p + '/af_corrected.tif') for p in paths]
rois = [np.loadtxt(p + '/ROI.txt') for p in paths]
```
### Run analysis
```
from scipy.special import erf
sigma = 2
thickness = 50
cytbg = (1 + erf((np.arange(thickness) - thickness / 2) / sigma)) / 2
cytbgs = []
for img, roi in zip(images, rois):
iq = ImageQuant(img, roi=roi, cyt_only=True, uni_cyt=True, iterations=2, adaptive_cytbg=True, thickness=thickness, cytbg=cytbg, descent_steps=200, nfits=100, freedom=10)
iq.run()
cytbgs.append(iq.cytbg)
cytbgs_norm = [x / np.max(x) for x in cytbgs]
```
### Figures
As we can see, the shape of the profile is highly conserved between embryos
(excluding last position as there are sometimes artefacts at this position - not sure why)
```
fig, ax = plt.subplots()
for c in cytbgs_norm:
ax.plot(c[:-1])
cytbg_arr = np.array(cytbgs_norm)
fig, ax = plt.subplots()
mean = np.mean(cytbg_arr, axis=0)[:-1]
std = np.std(cytbg_arr, axis=0)[:-1]
ax.plot(mean)
ax.fill_between(np.arange(len(mean)), mean-std, mean+std, alpha=0.2)
```
## Membrane reference profile inter-embryo variation
Performing membrane reference calibration on a few single embryos to see how variable it is between embryos. Using polarised PAR-2 Neon as before, and the cytoplasmic profile generated [here](./5_intro_to_imagequant.ipynb).
### Import data
```
path2 = '../test_datasets/dataset2_par2_neon'
paths2 = direcslist(path2)
images2 = [load_image(p + '/af_corrected.tif') for p in paths2]
rois2 = [np.loadtxt(p + '/ROI.txt') for p in paths2]
```
### Run analysis
```
sigma = 2
thickness = 50
membg = np.exp(-((np.arange(thickness) - thickness / 2) ** 2) / (2 * sigma ** 2))
cytbg = np.loadtxt('saved_cyt_ref_profile.txt')
membgs = []
for img, roi in zip(images2, rois2):
iq = ImageQuant(img, roi=roi, iterations=2, adaptive_membg=True, adaptive_cytbg=False, thickness=50, cytbg=cytbg, membg=membg, descent_steps=200, uni_cyt=True, nfits=100, freedom=10)
iq.run()
membgs.append(iq.membg)
membgs_norm = [x / np.max(x) for x in membgs]
```
### Figures
As we can see, the shape of the profile is highly conserved between embryos:
```
fig, ax = plt.subplots()
for m in membgs_norm:
ax.plot(m[:-1])
membg_arr = np.array(membgs_norm)
fig, ax = plt.subplots()
mean = np.mean(membg_arr, axis=0)[:-1]
std = np.std(membg_arr, axis=0)[:-1]
ax.plot(mean)
ax.fill_between(np.arange(len(mean)), mean-std, mean+std, alpha=0.2)
```
| github_jupyter |
## Observations and Insights
```
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
import scipy.stats as st
# Study data files
mouse_metadata_path = "data/Mouse_metadata.csv"
study_results_path = "data/Study_results.csv"
# Read the mouse data and the study results
mouse_metadata = pd.read_csv(mouse_metadata_path)
study_results = pd.read_csv(study_results_path)
# Combine the data into a single dataset
# Display the data table for preview
# Checking the number of mice.
# Getting the duplicate mice by ID number that shows up for Mouse ID and Timepoint.
# Optional: Get all the data for the duplicate mouse ID.
# Create a clean DataFrame by dropping the duplicate mouse by its ID.
# Checking the number of mice in the clean DataFrame.
```
## Summary Statistics
```
# Generate a summary statistics table of mean, median, variance, standard deviation, and SEM of the tumor volume for each regimen
# Use groupby and summary statistical methods to calculate the following properties of each drug regimen:
# mean, median, variance, standard deviation, and SEM of the tumor volume.
# Assemble the resulting series into a single summary dataframe.
# Generate a summary statistics table of mean, median, variance, standard deviation, and SEM of the tumor volume for each regimen
# Using the aggregation method, produce the same summary statistics in a single line
```
## Bar and Pie Charts
```
# Generate a bar plot showing the total number of measurements taken on each drug regimen using pandas.
# Generate a bar plot showing the total number of measurements taken on each drug regimen using pyplot.
# Generate a pie plot showing the distribution of female versus male mice using pandas
# Generate a pie plot showing the distribution of female versus male mice using pyplot
```
## Quartiles, Outliers and Boxplots
```
# Calculate the final tumor volume of each mouse across four of the treatment regimens:
# Capomulin, Ramicane, Infubinol, and Ceftamin
# Start by getting the last (greatest) timepoint for each mouse
# Merge this group df with the original dataframe to get the tumor volume at the last timepoint
# Put treatments into a list for for loop (and later for plot labels)
# Create empty list to fill with tumor vol data (for plotting)
# Calculate the IQR and quantitatively determine if there are any potential outliers.
# Locate the rows which contain mice on each drug and get the tumor volumes
# add subset
# Determine outliers using upper and lower bounds
# Generate a box plot of the final tumor volume of each mouse across four regimens of interest
```
## Line and Scatter Plots
```
# Generate a line plot of tumor volume vs. time point for a mouse treated with Capomulin
# Generate a scatter plot of average tumor volume vs. mouse weight for the Capomulin regimen
```
## Correlation and Regression
```
# Calculate the correlation coefficient and linear regression model
# for mouse weight and average tumor volume for the Capomulin regimen
```
| github_jupyter |
```
%matplotlib inline
```
(beta) Building a Convolution/Batch Norm fuser in FX
*******************************************************
**Author**: `Horace He <https://github.com/chillee>`_
In this tutorial, we are going to use FX, a toolkit for composable function
transformations of PyTorch, to do the following:
1) Find patterns of conv/batch norm in the data dependencies.
2) For the patterns found in 1), fold the batch norm statistics into the convolution weights.
Note that this optimization only works for models in inference mode (i.e. `mode.eval()`)
We will be building the fuser that exists here:
https://github.com/pytorch/pytorch/blob/orig/release/1.8/torch/fx/experimental/fuser.py
First, let's get some imports out of the way (we will be using all
of these later in the code).
```
from typing import Type, Dict, Any, Tuple, Iterable
import copy
import torch.fx as fx
import torch
import torch.nn as nn
```
For this tutorial, we are going to create a model consisting of convolutions
and batch norms. Note that this model has some tricky components - some of
the conv/batch norm patterns are hidden within Sequentials and one of the
BatchNorms is wrapped in another Module.
```
class WrappedBatchNorm(nn.Module):
def __init__(self):
super().__init__()
self.mod = nn.BatchNorm2d(1)
def forward(self, x):
return self.mod(x)
class M(nn.Module):
def __init__(self):
super().__init__()
self.conv1 = nn.Conv2d(1, 1, 1)
self.bn1 = nn.BatchNorm2d(1)
self.conv2 = nn.Conv2d(1, 1, 1)
self.nested = nn.Sequential(
nn.BatchNorm2d(1),
nn.Conv2d(1, 1, 1),
)
self.wrapped = WrappedBatchNorm()
def forward(self, x):
x = self.conv1(x)
x = self.bn1(x)
x = self.conv2(x)
x = self.nested(x)
x = self.wrapped(x)
return x
model = M()
model.eval()
```
Fusing Convolution with Batch Norm
-----------------------------------------
One of the primary challenges with trying to automatically fuse convolution
and batch norm in PyTorch is that PyTorch does not provide an easy way of
accessing the computational graph. FX resolves this problem by symbolically
tracing the actual operations called, so that we can track the computations
through the `forward` call, nested within Sequential modules, or wrapped in
an user-defined module.
```
traced_model = torch.fx.symbolic_trace(model)
print(traced_model.graph)
```
This gives us a graph representation of our model. Note that both the modules
hidden within the sequential as well as the wrapped Module have been inlined
into the graph. This is the default level of abstraction, but it can be
configured by the pass writer. More information can be found at the FX
overview https://pytorch.org/docs/master/fx.html#module-torch.fx
Fusing Convolution with Batch Norm
----------------------------------
Unlike some other fusions, fusion of convolution with batch norm does not
require any new operators. Instead, as batch norm during inference
consists of a pointwise add and multiply, these operations can be "baked"
into the preceding convolution's weights. This allows us to remove the batch
norm entirely from our model! Read
https://nenadmarkus.com/p/fusing-batchnorm-and-conv/ for further details. The
code here is copied from
https://github.com/pytorch/pytorch/blob/orig/release/1.8/torch/nn/utils/fusion.py
clarity purposes.
```
def fuse_conv_bn_eval(conv, bn):
"""
Given a conv Module `A` and an batch_norm module `B`, returns a conv
module `C` such that C(x) == B(A(x)) in inference mode.
"""
assert(not (conv.training or bn.training)), "Fusion only for eval!"
fused_conv = copy.deepcopy(conv)
fused_conv.weight, fused_conv.bias = \
fuse_conv_bn_weights(fused_conv.weight, fused_conv.bias,
bn.running_mean, bn.running_var, bn.eps, bn.weight, bn.bias)
return fused_conv
def fuse_conv_bn_weights(conv_w, conv_b, bn_rm, bn_rv, bn_eps, bn_w, bn_b):
if conv_b is None:
conv_b = torch.zeros_like(bn_rm)
if bn_w is None:
bn_w = torch.ones_like(bn_rm)
if bn_b is None:
bn_b = torch.zeros_like(bn_rm)
bn_var_rsqrt = torch.rsqrt(bn_rv + bn_eps)
conv_w = conv_w * (bn_w * bn_var_rsqrt).reshape([-1] + [1] * (len(conv_w.shape) - 1))
conv_b = (conv_b - bn_rm) * bn_var_rsqrt * bn_w + bn_b
return torch.nn.Parameter(conv_w), torch.nn.Parameter(conv_b)
```
FX Fusion Pass
----------------------------------
Now that we have our computational graph as well as a method for fusing
convolution and batch norm, all that remains is to iterate over the FX graph
and apply the desired fusions.
```
def _parent_name(target : str) -> Tuple[str, str]:
"""
Splits a qualname into parent path and last atom.
For example, `foo.bar.baz` -> (`foo.bar`, `baz`)
"""
*parent, name = target.rsplit('.', 1)
return parent[0] if parent else '', name
def replace_node_module(node: fx.Node, modules: Dict[str, Any], new_module: torch.nn.Module):
assert(isinstance(node.target, str))
parent_name, name = _parent_name(node.target)
setattr(modules[parent_name], name, new_module)
def fuse(model: torch.nn.Module) -> torch.nn.Module:
model = copy.deepcopy(model)
# The first step of most FX passes is to symbolically trace our model to
# obtain a `GraphModule`. This is a representation of our original model
# that is functionally identical to our original model, except that we now
# also have a graph representation of our forward pass.
fx_model: fx.GraphModule = fx.symbolic_trace(model)
modules = dict(fx_model.named_modules())
# The primary representation for working with FX are the `Graph` and the
# `Node`. Each `GraphModule` has a `Graph` associated with it - this
# `Graph` is also what generates `GraphModule.code`.
# The `Graph` itself is represented as a list of `Node` objects. Thus, to
# iterate through all of the operations in our graph, we iterate over each
# `Node` in our `Graph`.
for node in fx_model.graph.nodes:
# The FX IR contains several types of nodes, which generally represent
# call sites to modules, functions, or methods. The type of node is
# determined by `Node.op`.
if node.op != 'call_module': # If our current node isn't calling a Module then we can ignore it.
continue
# For call sites, `Node.target` represents the module/function/method
# that's being called. Here, we check `Node.target` to see if it's a
# batch norm module, and then check `Node.args[0].target` to see if the
# input `Node` is a convolution.
if type(modules[node.target]) is nn.BatchNorm2d and type(modules[node.args[0].target]) is nn.Conv2d:
if len(node.args[0].users) > 1: # Output of conv is used by other nodes
continue
conv = modules[node.args[0].target]
bn = modules[node.target]
fused_conv = fuse_conv_bn_eval(conv, bn)
replace_node_module(node.args[0], modules, fused_conv)
# As we've folded the batch nor into the conv, we need to replace all uses
# of the batch norm with the conv.
node.replace_all_uses_with(node.args[0])
# Now that all uses of the batch norm have been replaced, we can
# safely remove the batch norm.
fx_model.graph.erase_node(node)
fx_model.graph.lint()
# After we've modified our graph, we need to recompile our graph in order
# to keep the generated code in sync.
fx_model.recompile()
return fx_model
```
<div class="alert alert-info"><h4>Note</h4><p>We make some simplifications here for demonstration purposes, such as only
matching 2D convolutions. View
https://github.com/pytorch/pytorch/blob/master/torch/fx/experimental/fuser.py
for a more usable pass.</p></div>
Testing out our Fusion Pass
-----------------------------------------
We can now run this fusion pass on our initial toy model and verify that our
results are identical. In addition, we can print out the code for our fused
model and verify that there are no more batch norms.
```
fused_model = fuse(model)
print(fused_model.code)
inp = torch.randn(5, 1, 1, 1)
torch.testing.assert_allclose(fused_model(inp), model(inp))
```
Benchmarking our Fusion on ResNet18
----------
We can test our fusion pass on a larger model like ResNet18 and see how much
this pass improves inference performance.
```
import torchvision.models as models
import time
rn18 = models.resnet18()
rn18.eval()
inp = torch.randn(10, 3, 224, 224)
output = rn18(inp)
def benchmark(model, iters=20):
for _ in range(10):
model(inp)
begin = time.time()
for _ in range(iters):
model(inp)
return str(time.time()-begin)
fused_rn18 = fuse(rn18)
print("Unfused time: ", benchmark(rn18))
print("Fused time: ", benchmark(fused_rn18))
```
As we previously saw, the output of our FX transformation is
(Torchscriptable) PyTorch code, we can easily `jit.script` the output to try
and increase our performance even more. In this way, our FX model
transformation composes with Torchscript with no issues.
```
jit_rn18 = torch.jit.script(fused_rn18)
print("jit time: ", benchmark(jit_rn18))
############
# Conclusion
# ----------
# As we can see, using FX we can easily write static graph transformations on
# PyTorch code.
#
# Since FX is still in beta, we would be happy to hear any
# feedback you have about using it. Please feel free to use the
# PyTorch Forums (https://discuss.pytorch.org/) and the issue tracker
# (https://github.com/pytorch/pytorch/issues) to provide any feedback
# you might have.
```
| github_jupyter |
```
import torch
import torch.nn as nn
import torch.optim as optim
import torchtext
import torchtext.data as data
import torch.nn.functional as F
import matplotlib.pyplot as plt
import os
import random
%matplotlib inline
%config Completer.use_jedi = False
```
```bash
bash ./preprocess.sh dump-tokenized
cat ~/data/tokenized/wiki_ko_mecab.txt ~/data/tokenized/ratings_mecab.txt ~/data/tokenized/korquad_mecab.txt > ~/data/tokenized/corpus_mecab.txt
```
<h1>2-1. Neural Probabilistic Language Model<span class="tocSkip"></span></h1>
<div class="toc"><ul class="toc-item"><li><span><a href="#Create-dataset" data-toc-modified-id="Create-dataset-1"><span class="toc-item-num">1 </span>Create dataset</a></span></li><li><span><a href="#Build-the-model" data-toc-modified-id="Build-the-model-2"><span class="toc-item-num">2 </span>Build the model</a></span></li><li><span><a href="#Train-the-model" data-toc-modified-id="Train-the-model-3"><span class="toc-item-num">3 </span>Train the model</a></span></li><li><span><a href="#Embedding-result" data-toc-modified-id="Embedding-result-4"><span class="toc-item-num">4 </span>Embedding result</a></span></li></ul></div>
## Create dataset
* write each sample as a CSV file
```
n_gram = 3
corpus_path = os.path.join(os.getenv("HOME"), "data/tokenized/corpus_mecab.txt")
csv_path = os.path.join(os.getenv("HOME"), "data/csv/")
ngram_path = os.path.join(csv_path, f"{n_gram}gram_corpus_mecab.csv")
```
```python
import tqdm
import csv
with open(corpus_path) as r:
if not os.path.isdir(csv_path):
os.mkdir(csv_path)
with open(ngram_path, "w", encoding='utf-8') as w:
fieldnames = ['text', 'label']
writer = csv.DictWriter(w, fieldnames=fieldnames)
#writer.writeheader()
for idx, sample in tqdm.tqdm(enumerate(r)):
sentence = sample.split(" ")
for i in range(len(sentence)-n_gram):
text = " ".join(sentence[i:i+n_gram])
target = sentence[i+n_gram]
writer.writerow({"text": text, "label": target})
```
```bash
# `gsplit` if on macOS
split -d -l 1000000 ~/data/csv/3gram_corpus_mecab.csv ~/data/csv/3gram_corpus_mecab_ --additional-suffix=.csv
```
* build custom torchtext dataset with `torchtext.data.TabularDataset`.
```
TEXT = data.Field()
LABEL = data.Field()
datafiles = [fname for fname in os.listdir(csv_path) if fname.startswith("3gram_corpus_mecab_")]
dataset = data.TabularDataset(
os.path.join(csv_path, datafiles[0]),
format="csv",
skip_header=True,
fields=[
("text", TEXT),
("label", LABEL)
]
)
```
* train only on small portion (for speed)
```
trainset, testset = dataset.split(0.1, random_state=random.seed(0))
trainset, validationset = trainset.split(0.9, random_state=random.seed(0))
```
* Build vocabulary
```
MAX_VOCAB_SIZE = 100_000
TEXT.build_vocab(trainset, max_size=MAX_VOCAB_SIZE)
LABEL.build_vocab(trainset)
```
* Now we build a `BucketIterator` for our model.
```
BATCH_SIZE = 64
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
device
trainiter, validationiter, testiter = torchtext.data.BucketIterator.splits(
(trainset, validationset, testset),
batch_size=BATCH_SIZE,
device=device
)
```
## Build the model
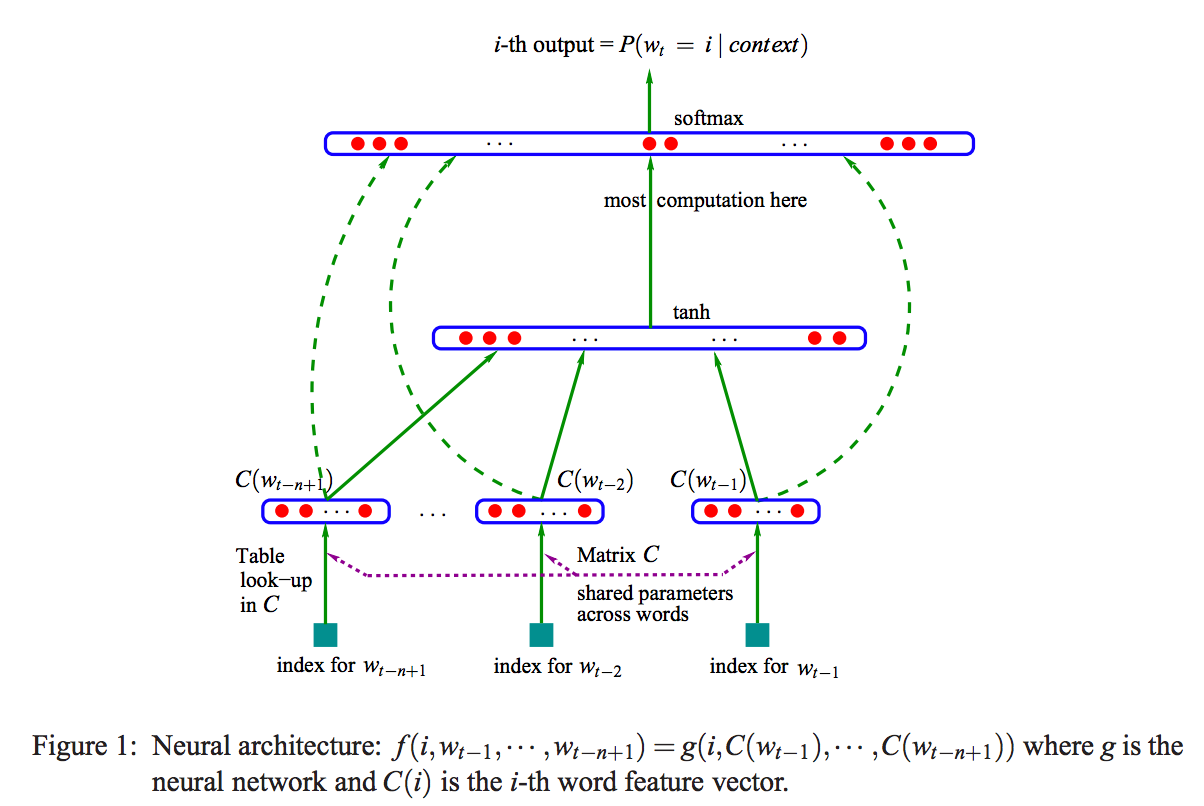
```
VOCAB_SIZE = len(TEXT.vocab)
N_GRAM = 3
EMBEDDING_DIM = 128
HIDDEN_DIM = 128
class NPLM(nn.Module):
def __init__(self, vocab_size, n_gram, embedding_dim, hidden_dim):
super(NPLM, self).__init__()
self.vocab_size = vocab_size
self.n_gram = n_gram
self.embedding_dim = embedding_dim
self.hidden_dim = hidden_dim
# embedding
self.embedding = nn.Embedding(vocab_size, embedding_dim)
# affine layers for tanh
self.linear1 = nn.Linear(n_gram * embedding_dim, hidden_dim)
self.linear2 = nn.Linear(hidden_dim, vocab_size, bias=False)
# affine layer for residual connection
self.linear3 = nn.Linear(n_gram * embedding_dim, vocab_size)
def forward(self, x):
x = self.embedding(x)
#print(x.shape)
x = x.view(-1, self.embedding_dim * self.n_gram)
#print(x.shape)
x1 = torch.tanh(self.linear1(x))
x1 = self.linear2(x1)
x2 = self.linear3(x)
x = x1 + x2
#print(x.shape)
return x
```
## Train the model
* train the model
```
model = NPLM(VOCAB_SIZE, N_GRAM, EMBEDDING_DIM, HIDDEN_DIM)
criterion = nn.CrossEntropyLoss()
optimizer = optim.SGD(model.parameters(), lr=0.01, momentum=0.9)
model = model.to(device)
criterion = criterion.to(device)
def accuracy(pred, target):
pred = torch.argmax(torch.softmax(pred, dim=1), dim=1)
correct = (pred == target).float()
#print(pred.shape, correct.shape)
return correct.sum() / len(correct)
def train(model, iterator, criterion, optimizer):
loss_epoch = 0.
acc_epoch = 0.
for batch in trainiter:
model.zero_grad()
out = model(batch.text)
#print("output:", out.shape)
out = out.squeeze(0)
#print("output (squeezed):", out.shape)
target = batch.label.squeeze(0)
#print("target:", target.shape)
loss = criterion(out, target)
loss.backward()
optimizer.step()
loss_epoch += loss.item()
acc_epoch += accuracy(out, target).item()
return loss_epoch, acc_epoch
N_EPOCH = 200
losses = []
accs = []
for i in range(1, N_EPOCH+1):
loss_epoch, acc_epoch = train(model, trainiter, criterion, optimizer)
losses.append(loss_epoch)
accs.append(acc_epoch)
if i % 5 == 0:
print(f"epoch: {i:03}, loss: {loss_epoch/len(trainiter): .3f}, acc: {acc_epoch/len(trainiter): .4f}")
```
* save the intermediate model
```
torch.save(model.state_dict(), "./NPLM_SGD_lr0.01_momentum0.9_epoch200.pth")
```
## Embedding result
* load the pre-trained final model
```
model = NPLM(VOCAB_SIZE, N_GRAM, EMBEDDING_DIM, HIDDEN_DIM)
model.load_state_dict(torch.load("./NPLM_SGD_lr0.01+0.005+0.001_momentum0.9_epoch200+115+50.pth"))
model.eval()
embedding = model.embedding
```
* plot the embeddings
```
from sklearn.manifold import TSNE
test_words = TEXT.vocab.freqs.most_common(1000)
test_words_raw = [w for w, _ in test_words if len(w) > 1]
test_words = [TEXT.vocab.stoi[w] for w in test_words_raw]
with torch.no_grad():
embed_xy = embedding(torch.tensor(test_words)).detach().numpy()
embed_xy = TSNE(n_components=2).fit_transform(embed_xy)
embed_x, embed_y = list(zip(*embed_xy))
plt.figure(figsize=(16,16))
for xy, word in zip(embed_xy, test_words_raw):
plt.annotate(word, xy, clip_on=True)
plt.title("Word Embedding")
plt.scatter(embed_x, embed_y, alpha=.3)
plt.axhline([0], ls=":", c="grey")
plt.axvline([0], ls=":", c="grey")
```
| github_jupyter |
---
_You are currently looking at **version 1.0** of this notebook. To download notebooks and datafiles, as well as get help on Jupyter notebooks in the Coursera platform, visit the [Jupyter Notebook FAQ](https://www.coursera.org/learn/python-machine-learning/resources/bANLa) course resource._
---
# Applied Machine Learning: Module 3 (Evaluation)
## Evaluation for Classification
### Preamble
```
%matplotlib notebook
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.datasets import load_digits
dataset = load_digits()
X, y = dataset.data, dataset.target
for class_name, class_count in zip(dataset.target_names, np.bincount(dataset.target)):
print(class_name,class_count)
# Creating a dataset with imbalanced binary classes:
# Negative class (0) is 'not digit 1'
# Positive class (1) is 'digit 1'
y_binary_imbalanced = y.copy()
y_binary_imbalanced[y_binary_imbalanced != 1] = 0
print('Original labels:\t', y[1:30])
print('New binary labels:\t', y_binary_imbalanced[1:30])
np.bincount(y_binary_imbalanced) # Negative class (0) is the most frequent class
X_train, X_test, y_train, y_test = train_test_split(X, y_binary_imbalanced, random_state=0)
# Accuracy of Support Vector Machine classifier
from sklearn.svm import SVC
svm = SVC(kernel='rbf', C=1).fit(X_train, y_train)
svm.score(X_test, y_test)
```
### Dummy Classifiers
DummyClassifier is a classifier that makes predictions using simple rules, which can be useful as a baseline for comparison against actual classifiers, especially with imbalanced classes.
```
from sklearn.dummy import DummyClassifier
# Negative class (0) is most frequent
dummy_majority = DummyClassifier(strategy = 'most_frequent').fit(X_train, y_train)
# Therefore the dummy 'most_frequent' classifier always predicts class 0
y_dummy_predictions = dummy_majority.predict(X_test)
y_dummy_predictions
dummy_majority.score(X_test, y_test)
svm = SVC(kernel='linear', C=1).fit(X_train, y_train)
svm.score(X_test, y_test)
```
### Confusion matrices
#### Binary (two-class) confusion matrix
```
from sklearn.metrics import confusion_matrix
# Negative class (0) is most frequent
dummy_majority = DummyClassifier(strategy = 'most_frequent').fit(X_train, y_train)
y_majority_predicted = dummy_majority.predict(X_test)
confusion = confusion_matrix(y_test, y_majority_predicted)
print('Most frequent class (dummy classifier)\n', confusion)
# produces random predictions w/ same class proportion as training set
dummy_classprop = DummyClassifier(strategy='stratified').fit(X_train, y_train)
y_classprop_predicted = dummy_classprop.predict(X_test)
confusion = confusion_matrix(y_test, y_classprop_predicted)
print('Random class-proportional prediction (dummy classifier)\n', confusion)
svm = SVC(kernel='linear', C=1).fit(X_train, y_train)
svm_predicted = svm.predict(X_test)
confusion = confusion_matrix(y_test, svm_predicted)
print('Support vector machine classifier (linear kernel, C=1)\n', confusion)
from sklearn.linear_model import LogisticRegression
lr = LogisticRegression().fit(X_train, y_train)
lr_predicted = lr.predict(X_test)
confusion = confusion_matrix(y_test, lr_predicted)
print('Logistic regression classifier (default settings)\n', confusion)
from sklearn.tree import DecisionTreeClassifier
dt = DecisionTreeClassifier(max_depth=2).fit(X_train, y_train)
tree_predicted = dt.predict(X_test)
confusion = confusion_matrix(y_test, tree_predicted)
print('Decision tree classifier (max_depth = 2)\n', confusion)
```
### Evaluation metrics for binary classification
```
from sklearn.metrics import accuracy_score, precision_score, recall_score, f1_score
# Accuracy = TP + TN / (TP + TN + FP + FN)
# Precision = TP / (TP + FP)
# Recall = TP / (TP + FN) Also known as sensitivity, or True Positive Rate
# F1 = 2 * Precision * Recall / (Precision + Recall)
print('Accuracy: {:.2f}'.format(accuracy_score(y_test, tree_predicted)))
print('Precision: {:.2f}'.format(precision_score(y_test, tree_predicted)))
print('Recall: {:.2f}'.format(recall_score(y_test, tree_predicted)))
print('F1: {:.2f}'.format(f1_score(y_test, tree_predicted)))
# Combined report with all above metrics
from sklearn.metrics import classification_report
print(classification_report(y_test, tree_predicted, target_names=['not 1', '1']))
print('Random class-proportional (dummy)\n',
classification_report(y_test, y_classprop_predicted, target_names=['not 1', '1']))
print('SVM\n',
classification_report(y_test, svm_predicted, target_names = ['not 1', '1']))
print('Logistic regression\n',
classification_report(y_test, lr_predicted, target_names = ['not 1', '1']))
print('Decision tree\n',
classification_report(y_test, tree_predicted, target_names = ['not 1', '1']))
```
### Decision functions
```
X_train, X_test, y_train, y_test = train_test_split(X, y_binary_imbalanced, random_state=0)
y_scores_lr = lr.fit(X_train, y_train).decision_function(X_test)
y_score_list = list(zip(y_test[0:20], y_scores_lr[0:20]))
# show the decision_function scores for first 20 instances
y_score_list
X_train, X_test, y_train, y_test = train_test_split(X, y_binary_imbalanced, random_state=0)
y_proba_lr = lr.fit(X_train, y_train).predict_proba(X_test)
y_proba_list = list(zip(y_test[0:20], y_proba_lr[0:20,1]))
# show the probability of positive class for first 20 instances
y_proba_list
```
### Precision-recall curves
```
from sklearn.metrics import precision_recall_curve
precision, recall, thresholds = precision_recall_curve(y_test, y_scores_lr)
closest_zero = np.argmin(np.abs(thresholds))
closest_zero_p = precision[closest_zero]
closest_zero_r = recall[closest_zero]
plt.figure()
plt.xlim([0.0, 1.01])
plt.ylim([0.0, 1.01])
plt.plot(precision, recall, label='Precision-Recall Curve')
plt.plot(closest_zero_p, closest_zero_r, 'o', markersize = 12, fillstyle = 'none', c='r', mew=3)
plt.xlabel('Precision', fontsize=16)
plt.ylabel('Recall', fontsize=16)
plt.axes().set_aspect('equal')
plt.show()
```
### ROC curves, Area-Under-Curve (AUC)
```
from sklearn.metrics import roc_curve, auc
X_train, X_test, y_train, y_test = train_test_split(X, y_binary_imbalanced, random_state=0)
y_score_lr = lr.fit(X_train, y_train).decision_function(X_test)
fpr_lr, tpr_lr, _ = roc_curve(y_test, y_score_lr)
roc_auc_lr = auc(fpr_lr, tpr_lr)
plt.figure()
plt.xlim([-0.01, 1.00])
plt.ylim([-0.01, 1.01])
plt.plot(fpr_lr, tpr_lr, lw=3, label='LogRegr ROC curve (area = {:0.2f})'.format(roc_auc_lr))
plt.xlabel('False Positive Rate', fontsize=16)
plt.ylabel('True Positive Rate', fontsize=16)
plt.title('ROC curve (1-of-10 digits classifier)', fontsize=16)
plt.legend(loc='lower right', fontsize=13)
plt.plot([0, 1], [0, 1], color='navy', lw=3, linestyle='--')
plt.axes().set_aspect('equal')
plt.show()
from matplotlib import cm
X_train, X_test, y_train, y_test = train_test_split(X, y_binary_imbalanced, random_state=0)
plt.figure()
plt.xlim([-0.01, 1.00])
plt.ylim([-0.01, 1.01])
for g in [0.01, 0.1, 0.20, 1]:
svm = SVC(gamma=g).fit(X_train, y_train)
y_score_svm = svm.decision_function(X_test)
fpr_svm, tpr_svm, _ = roc_curve(y_test, y_score_svm)
roc_auc_svm = auc(fpr_svm, tpr_svm)
accuracy_svm = svm.score(X_test, y_test)
print("gamma = {:.2f} accuracy = {:.2f} AUC = {:.2f}".format(g, accuracy_svm,
roc_auc_svm))
plt.plot(fpr_svm, tpr_svm, lw=3, alpha=0.7,
label='SVM (gamma = {:0.2f}, area = {:0.2f})'.format(g, roc_auc_svm))
plt.xlabel('False Positive Rate', fontsize=16)
plt.ylabel('True Positive Rate (Recall)', fontsize=16)
plt.plot([0, 1], [0, 1], color='k', lw=0.5, linestyle='--')
plt.legend(loc="lower right", fontsize=11)
plt.title('ROC curve: (1-of-10 digits classifier)', fontsize=16)
plt.axes().set_aspect('equal')
plt.show()
```
### Evaluation measures for multi-class classification
#### Multi-class confusion matrix
```
dataset = load_digits()
X, y = dataset.data, dataset.target
X_train_mc, X_test_mc, y_train_mc, y_test_mc = train_test_split(X, y, random_state=0)
svm = SVC(kernel = 'linear').fit(X_train_mc, y_train_mc)
svm_predicted_mc = svm.predict(X_test_mc)
confusion_mc = confusion_matrix(y_test_mc, svm_predicted_mc)
df_cm = pd.DataFrame(confusion_mc,
index = [i for i in range(0,10)], columns = [i for i in range(0,10)])
plt.figure(figsize=(5.5,4))
sns.heatmap(df_cm, annot=True)
plt.title('SVM Linear Kernel \nAccuracy:{0:.3f}'.format(accuracy_score(y_test_mc,
svm_predicted_mc)))
plt.ylabel('True label')
plt.xlabel('Predicted label')
svm = SVC(kernel = 'rbf').fit(X_train_mc, y_train_mc)
svm_predicted_mc = svm.predict(X_test_mc)
confusion_mc = confusion_matrix(y_test_mc, svm_predicted_mc)
df_cm = pd.DataFrame(confusion_mc, index = [i for i in range(0,10)],
columns = [i for i in range(0,10)])
plt.figure(figsize = (5.5,4))
sns.heatmap(df_cm, annot=True)
plt.title('SVM RBF Kernel \nAccuracy:{0:.3f}'.format(accuracy_score(y_test_mc,
svm_predicted_mc)))
plt.ylabel('True label')
plt.xlabel('Predicted label');
```
#### Multi-class classification report
```
print(classification_report(y_test_mc, svm_predicted_mc))
```
#### Micro- vs. macro-averaged metrics
```
print('Micro-averaged precision = {:.2f} (treat instances equally)'
.format(precision_score(y_test_mc, svm_predicted_mc, average = 'micro')))
print('Macro-averaged precision = {:.2f} (treat classes equally)'
.format(precision_score(y_test_mc, svm_predicted_mc, average = 'macro')))
print('Micro-averaged f1 = {:.2f} (treat instances equally)'
.format(f1_score(y_test_mc, svm_predicted_mc, average = 'micro')))
print('Macro-averaged f1 = {:.2f} (treat classes equally)'
.format(f1_score(y_test_mc, svm_predicted_mc, average = 'macro')))
```
### Regression evaluation metrics
```
%matplotlib notebook
import matplotlib.pyplot as plt
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn import datasets
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_squared_error, r2_score
from sklearn.dummy import DummyRegressor
diabetes = datasets.load_diabetes()
X = diabetes.data[:, None, 6]
y = diabetes.target
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0)
lm = LinearRegression().fit(X_train, y_train)
lm_dummy_mean = DummyRegressor(strategy = 'mean').fit(X_train, y_train)
y_predict = lm.predict(X_test)
y_predict_dummy_mean = lm_dummy_mean.predict(X_test)
print('Linear model, coefficients: ', lm.coef_)
print("Mean squared error (dummy): {:.2f}".format(mean_squared_error(y_test,
y_predict_dummy_mean)))
print("Mean squared error (linear model): {:.2f}".format(mean_squared_error(y_test, y_predict)))
print("r2_score (dummy): {:.2f}".format(r2_score(y_test, y_predict_dummy_mean)))
print("r2_score (linear model): {:.2f}".format(r2_score(y_test, y_predict)))
# Plot outputs
plt.scatter(X_test, y_test, color='black')
plt.plot(X_test, y_predict, color='green', linewidth=2)
plt.plot(X_test, y_predict_dummy_mean, color='red', linestyle = 'dashed',
linewidth=2, label = 'dummy')
plt.show()
```
### Model selection using evaluation metrics
#### Cross-validation example
```
from sklearn.model_selection import cross_val_score
from sklearn.svm import SVC
dataset = load_digits()
# again, making this a binary problem with 'digit 1' as positive class
# and 'not 1' as negative class
X, y = dataset.data, dataset.target == 1
clf = SVC(kernel='linear', C=1)
# accuracy is the default scoring metric
print('Cross-validation (accuracy)', cross_val_score(clf, X, y, cv=5))
# use AUC as scoring metric
print('Cross-validation (AUC)', cross_val_score(clf, X, y, cv=5, scoring = 'roc_auc'))
# use recall as scoring metric
print('Cross-validation (recall)', cross_val_score(clf, X, y, cv=5, scoring = 'recall'))
```
#### Grid search example
```
from sklearn.svm import SVC
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import roc_auc_score
dataset = load_digits()
X, y = dataset.data, dataset.target == 1
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0)
clf = SVC(kernel='rbf')
grid_values = {'gamma': [0.001, 0.01, 0.05, 0.1, 1, 10, 100]}
# default metric to optimize over grid parameters: accuracy
grid_clf_acc = GridSearchCV(clf, param_grid = grid_values)
grid_clf_acc.fit(X_train, y_train)
y_decision_fn_scores_acc = grid_clf_acc.decision_function(X_test)
print('Grid best parameter (max. accuracy): ', grid_clf_acc.best_params_)
print('Grid best score (accuracy): ', grid_clf_acc.best_score_)
# alternative metric to optimize over grid parameters: AUC
grid_clf_auc = GridSearchCV(clf, param_grid = grid_values, scoring = 'roc_auc')
grid_clf_auc.fit(X_train, y_train)
y_decision_fn_scores_auc = grid_clf_auc.decision_function(X_test)
print('Test set AUC: ', roc_auc_score(y_test, y_decision_fn_scores_auc))
print('Grid best parameter (max. AUC): ', grid_clf_auc.best_params_)
print('Grid best score (AUC): ', grid_clf_auc.best_score_)
```
#### Evaluation metrics supported for model selection
```
from sklearn.metrics.scorer import SCORERS
print(sorted(list(SCORERS.keys())))
```
### Two-feature classification example using the digits dataset
#### Optimizing a classifier using different evaluation metrics
```
from sklearn.datasets import load_digits
from sklearn.model_selection import train_test_split
from adspy_shared_utilities import plot_class_regions_for_classifier_subplot
from sklearn.svm import SVC
from sklearn.model_selection import GridSearchCV
dataset = load_digits()
X, y = dataset.data, dataset.target == 1
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0)
# Create a two-feature input vector matching the example plot above
# We jitter the points (add a small amount of random noise) in case there are areas
# in feature space where many instances have the same features.
jitter_delta = 0.25
X_twovar_train = X_train[:,[20,59]]+ np.random.rand(X_train.shape[0], 2) - jitter_delta
X_twovar_test = X_test[:,[20,59]] + np.random.rand(X_test.shape[0], 2) - jitter_delta
clf = SVC(kernel = 'linear').fit(X_twovar_train, y_train)
grid_values = {'class_weight':['balanced', {1:2},{1:3},{1:4},{1:5},{1:10},{1:20},{1:50}]}
plt.figure(figsize=(9,6))
for i, eval_metric in enumerate(('precision','recall', 'f1','roc_auc')):
grid_clf_custom = GridSearchCV(clf, param_grid=grid_values, scoring=eval_metric)
grid_clf_custom.fit(X_twovar_train, y_train)
print('Grid best parameter (max. {0}): {1}'
.format(eval_metric, grid_clf_custom.best_params_))
print('Grid best score ({0}): {1}'
.format(eval_metric, grid_clf_custom.best_score_))
plt.subplots_adjust(wspace=0.3, hspace=0.3)
plot_class_regions_for_classifier_subplot(grid_clf_custom, X_twovar_test, y_test, None,
None, None, plt.subplot(2, 2, i+1))
plt.title(eval_metric+'-oriented SVC')
plt.tight_layout()
plt.show()
```
#### Precision-recall curve for the default SVC classifier (with balanced class weights)
```
from sklearn.model_selection import train_test_split
from sklearn.metrics import precision_recall_curve
from adspy_shared_utilities import plot_class_regions_for_classifier
from sklearn.svm import SVC
dataset = load_digits()
X, y = dataset.data, dataset.target == 1
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0)
# create a two-feature input vector matching the example plot above
jitter_delta = 0.25
X_twovar_train = X_train[:,[20,59]]+ np.random.rand(X_train.shape[0], 2) - jitter_delta
X_twovar_test = X_test[:,[20,59]] + np.random.rand(X_test.shape[0], 2) - jitter_delta
clf = SVC(kernel='linear', class_weight='balanced').fit(X_twovar_train, y_train)
y_scores = clf.decision_function(X_twovar_test)
precision, recall, thresholds = precision_recall_curve(y_test, y_scores)
closest_zero = np.argmin(np.abs(thresholds))
closest_zero_p = precision[closest_zero]
closest_zero_r = recall[closest_zero]
plot_class_regions_for_classifier(clf, X_twovar_test, y_test)
plt.title("SVC, class_weight = 'balanced', optimized for accuracy")
plt.show()
plt.figure()
plt.xlim([0.0, 1.01])
plt.ylim([0.0, 1.01])
plt.title ("Precision-recall curve: SVC, class_weight = 'balanced'")
plt.plot(precision, recall, label = 'Precision-Recall Curve')
plt.plot(closest_zero_p, closest_zero_r, 'o', markersize=12, fillstyle='none', c='r', mew=3)
plt.xlabel('Precision', fontsize=16)
plt.ylabel('Recall', fontsize=16)
plt.axes().set_aspect('equal')
plt.show()
print('At zero threshold, precision: {:.2f}, recall: {:.2f}'
.format(closest_zero_p, closest_zero_r))
```
| github_jupyter |
```
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the "../input/" directory.
# For example, running this (by clicking run or pressing Shift+Enter) will list the files in the input directory
import os
import string
from string import digits
import matplotlib.pyplot as plt
%matplotlib inline
import re
import seaborn as sns
from sklearn.utils import shuffle
from sklearn.model_selection import train_test_split
from keras.layers import Input, LSTM, Embedding, Dense
from keras.models import Model
print(os.listdir("../input"))
pd.set_option('display.max_rows', 500)
pd.set_option('display.max_columns', 500)
pd.set_option('display.width', 1000)
pd.set_option('display.max_colwidth', -1)
# Any results you write to the current directory are saved as output.
lines=pd.read_csv("/kaggle/input/hindi-english-truncated-corpus/Hindi_English_Truncated_Corpus.csv",encoding='utf-8')
lines['source'].value_counts()
lines=lines[lines['source']=='ted']
lines.head(10)
pd.isnull(lines).sum()
lines=lines[~pd.isnull(lines['english_sentence'])]
lines.drop_duplicates(inplace=True)
lines=lines.sample(n=25000,random_state=42)
lines.shape
# Lowercase all characters
lines['english_sentence']=lines['english_sentence'].apply(lambda x: x.lower())
lines['hindi_sentence']=lines['hindi_sentence'].apply(lambda x: x.lower())
# Remove quotes
lines['english_sentence']=lines['english_sentence'].apply(lambda x: re.sub("'", '', x))
lines['hindi_sentence']=lines['hindi_sentence'].apply(lambda x: re.sub("'", '', x))
exclude = set(string.punctuation) # Set of all special characters
# Remove all the special characters
lines['english_sentence']=lines['english_sentence'].apply(lambda x: ''.join(ch for ch in x if ch not in exclude))
lines['hindi_sentence']=lines['hindi_sentence'].apply(lambda x: ''.join(ch for ch in x if ch not in exclude))
# Remove all numbers from text
remove_digits = str.maketrans('', '', digits)
lines['english_sentence']=lines['english_sentence'].apply(lambda x: x.translate(remove_digits))
lines['hindi_sentence']=lines['hindi_sentence'].apply(lambda x: x.translate(remove_digits))
lines['hindi_sentence'] = lines['hindi_sentence'].apply(lambda x: re.sub("[२३०८१५७९४६]", "", x))
# Remove extra spaces
lines['english_sentence']=lines['english_sentence'].apply(lambda x: x.strip())
lines['hindi_sentence']=lines['hindi_sentence'].apply(lambda x: x.strip())
lines['english_sentence']=lines['english_sentence'].apply(lambda x: re.sub(" +", " ", x))
lines['hindi_sentence']=lines['hindi_sentence'].apply(lambda x: re.sub(" +", " ", x))
# Add start and end tokens to target sequences
lines['hindi_sentence'] = lines['hindi_sentence'].apply(lambda x : 'START_ '+ x + ' _END')
lines.head(10)
### Get English and Hindi Vocabulary
all_eng_words=set()
for eng in lines['english_sentence']:
for word in eng.split():
if word not in all_eng_words:
all_eng_words.add(word)
all_hindi_words=set()
for hin in lines['hindi_sentence']:
for word in hin.split():
if word not in all_hindi_words:
all_hindi_words.add(word)
len(all_eng_words)
len(all_hindi_words)
lines['length_eng_sentence']=lines['english_sentence'].apply(lambda x:len(x.split(" ")))
lines['length_hin_sentence']=lines['hindi_sentence'].apply(lambda x:len(x.split(" ")))
lines.head(10)
lines.shape
print("maximum length of Hindi Sentence ",max(lines['length_hin_sentence']))
print("maximum length of English Sentence ",max(lines['length_eng_sentence']))
max_length_src=max(lines['length_hin_sentence'])
max_length_tar=max(lines['length_eng_sentence'])
input_words = sorted(list(all_eng_words))
target_words = sorted(list(all_hindi_words))
num_encoder_tokens = len(all_eng_words)
num_decoder_tokens = len(all_hindi_words)
num_encoder_tokens, num_decoder_tokens
num_decoder_tokens += 1 #for zero padding
input_token_index = dict([(word, i+1) for i, word in enumerate(input_words)])
target_token_index = dict([(word, i+1) for i, word in enumerate(target_words)])
reverse_input_char_index = dict((i, word) for word, i in input_token_index.items())
reverse_target_char_index = dict((i, word) for word, i in target_token_index.items())
lines = shuffle(lines)
lines.head(10)
X, y = lines['english_sentence'], lines['hindi_sentence']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2,random_state=42)
X_train.shape, X_test.shape
# we save the data then
X_train.to_pickle('X_train.pkl')
X_test.to_pickle('X_test.pkl')
def generate_batch(X = X_train, y = y_train, batch_size = 128):
''' Generate a batch of data '''
while True:
for j in range(0, len(X), batch_size):
encoder_input_data = np.zeros((batch_size, max_length_src),dtype='float32')
decoder_input_data = np.zeros((batch_size, max_length_tar),dtype='float32')
decoder_target_data = np.zeros((batch_size, max_length_tar, num_decoder_tokens),dtype='float32')
for i, (input_text, target_text) in enumerate(zip(X[j:j+batch_size], y[j:j+batch_size])):
for t, word in enumerate(input_text.split()):
encoder_input_data[i, t] = input_token_index[word] # encoder input seq
for t, word in enumerate(target_text.split()):
if t<len(target_text.split())-1:
decoder_input_data[i, t] = target_token_index[word] # decoder input seq
if t>0:
# decoder target sequence (one hot encoded)
# does not include the START_ token
# Offset by one timestep
decoder_target_data[i, t - 1, target_token_index[word]] = 1.
yield([encoder_input_data, decoder_input_data], decoder_target_data)
latent_dim=300
# Encoder
encoder_inputs = Input(shape=(None,))
enc_emb = Embedding(num_encoder_tokens, latent_dim, mask_zero = True)(encoder_inputs)
encoder_lstm = LSTM(latent_dim, return_state=True)
encoder_outputs, state_h, state_c = encoder_lstm(enc_emb)
# We discard `encoder_outputs` and only keep the states.
encoder_states = [state_h, state_c]
# Set up the decoder, using `encoder_states` as initial state.
decoder_inputs = Input(shape=(None,))
dec_emb_layer = Embedding(num_decoder_tokens, latent_dim, mask_zero = True)
dec_emb = dec_emb_layer(decoder_inputs)
# We set up our decoder to return full output sequences,
# and to return internal states as well. We don't use the
# return states in the training model, but we will use them in inference.
decoder_lstm = LSTM(latent_dim, return_sequences=True, return_state=True)
decoder_outputs, _, _ = decoder_lstm(dec_emb,
initial_state=encoder_states)
decoder_dense = Dense(num_decoder_tokens, activation='softmax')
decoder_outputs = decoder_dense(decoder_outputs)
# Define the model that will turn
# `encoder_input_data` & `decoder_input_data` into `decoder_target_data`
model = Model([encoder_inputs, decoder_inputs], decoder_outputs)
model.compile(optimizer='rmsprop', loss='categorical_crossentropy')
model.summary()
train_samples = len(X_train)
val_samples = len(X_test)
batch_size = 128
epochs = 100
model.fit_generator(generator = generate_batch(X_train, y_train, batch_size = batch_size),
steps_per_epoch = train_samples//batch_size,
epochs=epochs,
validation_data = generate_batch(X_test, y_test, batch_size = batch_size),
validation_steps = val_samples//batch_size)
model.save_weights('nmt_weights.h5')
# Encode the input sequence to get the "thought vectors"
encoder_model = Model(encoder_inputs, encoder_states)
# Decoder setup
# Below tensors will hold the states of the previous time step
decoder_state_input_h = Input(shape=(latent_dim,))
decoder_state_input_c = Input(shape=(latent_dim,))
decoder_states_inputs = [decoder_state_input_h, decoder_state_input_c]
dec_emb2= dec_emb_layer(decoder_inputs) # Get the embeddings of the decoder sequence
# To predict the next word in the sequence, set the initial states to the states from the previous time step
decoder_outputs2, state_h2, state_c2 = decoder_lstm(dec_emb2, initial_state=decoder_states_inputs)
decoder_states2 = [state_h2, state_c2]
decoder_outputs2 = decoder_dense(decoder_outputs2) # A dense softmax layer to generate prob dist. over the target vocabulary
# Final decoder model
decoder_model = Model(
[decoder_inputs] + decoder_states_inputs,
[decoder_outputs2] + decoder_states2)
def decode_sequence(input_seq):
# Encode the input as state vectors.
states_value = encoder_model.predict(input_seq)
# Generate empty target sequence of length 1.
target_seq = np.zeros((1,1))
# Populate the first character of target sequence with the start character.
target_seq[0, 0] = target_token_index['START_']
# Sampling loop for a batch of sequences
# (to simplify, here we assume a batch of size 1).
stop_condition = False
decoded_sentence = ''
while not stop_condition:
output_tokens, h, c = decoder_model.predict([target_seq] + states_value)
# Sample a token
sampled_token_index = np.argmax(output_tokens[0, -1, :])
sampled_char = reverse_target_char_index[sampled_token_index]
decoded_sentence += ' '+sampled_char
# Exit condition: either hit max length
# or find stop character.
if (sampled_char == '_END' or
len(decoded_sentence) > 50):
stop_condition = True
# Update the target sequence (of length 1).
target_seq = np.zeros((1,1))
target_seq[0, 0] = sampled_token_index
# Update states
states_value = [h, c]
return decoded_sentence
train_gen = generate_batch(X_train, y_train, batch_size = 1)
k=-1
k+=1
(input_seq, actual_output), _ = next(train_gen)
decoded_sentence = decode_sequence(input_seq)
print('Input English sentence:', X_train[k:k+1].values[0])
print('Actual Hindi Translation:', y_train[k:k+1].values[0][6:-4])
print('Predicted Hindi Translation:', decoded_sentence[:-4])
k+=1
(input_seq, actual_output), _ = next(train_gen)
decoded_sentence = decode_sequence(input_seq)
print('Input English sentence:', X_train[k:k+1].values[0])
print('Actual Hindi Translation:', y_train[k:k+1].values[0][6:-4])
print('Predicted Hindi Translation:', decoded_sentence[:-4])
k+=1
(input_seq, actual_output), _ = next(train_gen)
decoded_sentence = decode_sequence(input_seq)
print('Input English sentence:', X_train[k:k+1].values[0])
print('Actual Hindi Translation:', y_train[k:k+1].values[0][6:-4])
print('Predicted Hindi Translation:', decoded_sentence[:-4])
k+=1
(input_seq, actual_output), _ = next(train_gen)
decoded_sentence = decode_sequence(input_seq)
print('Input English sentence:', X_train[k:k+1].values[0])
print('Actual Hindi Translation:', y_train[k:k+1].values[0][6:-4])
print('Predicted Hindi Translation:', decoded_sentence[:-4])
k+=1
(input_seq, actual_output), _ = next(train_gen)
decoded_sentence = decode_sequence(input_seq)
print('Input English sentence:', X_train[k:k+1].values[0])
print('Actual Hindi Translation:', y_train[k:k+1].values[0][6:-4])
print('Predicted Hindi Translation:', decoded_sentence[:-4])
a = y_train[k:k+1].values[0][6:-4]
b = decoded_sentence[:-4]
from nltk.translate.bleu_score import sentence_bleu
score = sentence_bleu( a, b)
print('Bleu score:', '%3f'%score)
```
| github_jupyter |
```
print('Hello World')
```
# Python Basics
## Overview
1. Basic Data Types
2. Everything is an Object
3. Primary Data Structures
4. Scientific Data Structures
## Basic Data Types
```
2
type(2)
type(2.4)
type(True)
type('hello')
# note, there is no 'char' type, it is just a string of length 1
type('c')
type(None)
```
### Operators
We can perform mathematical calculations in Python using the basic operators +, -, /, *, %:
```python
>>> 2 + 2
4
>>> 6 * 7
42
>>> 2 ** 16 # power
65536
>>> 13 % 5 # modulo
3
```
We can also use comparison and logic operators: <, >, ==, !=, <=, >= etc. and, or, not
```python
>>> 3 > 4
False
>>> True and True
True
>>> True or False
True
>>> 3 == 3
True
```
### Assignment is done using =
```
text = 'ERDC Python Tutorial'
number = 42
```
## Python is dynamically but strongly typed.
The fact that the two halves of that statement fit together can confuse those who come from a static language type background. In Python it is perfectly legal to do this :
```
pi = 3.1415
type(pi)
pi = 'The Ratio of the circumference of a circle to its diameter'
type(pi)
```
**WARNING:**
**The fact that things can be redefined on the fly gives Python enormous flexibility but
also means you need to be careful about certain things. Never use built in function
names as variable names. i.e. Don't do this ...**
```python
>>> print = 5
>>> print('hello world')
---------------------------------------------------------------------------
TypeError Traceback (most recent call last)
<ipython-input-11-def08996fe0c> in <module>()
----> 1 print('hello world')
TypeError: 'int' object is not callable
```
## Duck Typing
**If it looks like a duck, swims like a duck, and quacks like a duck, then it's a duck.**
```
3.1415 + 2
3.1415 + 'hello'
5 + '5'
# I can cast the types to make it work
5 + int('5')
def my_add(a, b):
return a + b
def my_mult(a, b):
return a*b
my_add(3, 4)
my_add('super', 'man')
my_mult(3, 4)
my_mult(3.1, 4)
my_mult('super', 'man')
my_mult('super', 3)
```
** The principle of duck typing says that you shouldn't care what type of object you have, just whether or not you can do the required action with your object. **
## Everything in Python is an Object and hence has methods
```
print(pi)
# type pi. and hit the tab key, this will bring up a list of methods on the string object pi
# choose the count object and lets see how we can use it to count the number of time the word 'the'
# occurs in pi
pi.
```
### Exercise 1.1
The count in the above should have been two, can you figure out why? Explore the other
methods available and pick one that can help. Apply it first before applying the .count() method
```
# explore methods on pi, type pi. and hit tab, select one, type a ? and hit shift-enter
pi.
# fill in the blank
new_pi = pi._____
count = new_pi.count('the')
print('The word "the" occurs %s times in pi' % count)
# solution
%load 'solutions/01-1.py'
```
## Python Sequences: Lists - Tuples - Strings
```
# lets define 3 lists
# lists are defined using square brackets
fruits = ['apple', 'orange', 'pear', 'watermelon']
scores = [96, 73, 55, 33, 88]
numbers = ['one', 'two', 'three', 'four', 'five', 'six', 'seven', 'eight', 'nine', 'ten']
mixed = ['one', 2, True, None, 3.145, pi] # each element of the list can be any python object
print(mixed)
len(fruits)
'apple' in fruits
'mango' in fruits
fruits.append('mango')
print(fruits)
```
### Exercise 1.2
Get rid of the number 96 from list called scores, add the number 68 and print the list
```
%load 'solutions/01-2.py'
```
## Indexing in Python
**indexing in python is zero based and uses square brackets**
```
numbers
numbers[0]
numbers[3]
numbers[10]
```
### The last element can be accessed with -1
```
numbers
numbers[-1]
numbers[-3]
```
### Slicing
** syntax -> [start:end:increment]**
```
# what will this print
print(numbers)
print(numbers[2:6])
# what will this print
print(numbers)
print(numbers[2:8:2])
# what will this print
print(numbers)
print(numbers[:5])
# what will this print
print(numbers)
print(numbers[5:])
# what will this print
print(numbers)
print(numbers[::2])
```
### Exercise 1.3
Make a list the contains the numbers 40 through 50 and use indexing to print
the last 8 numbers
```
mynumbers = [____]
print(mynumbers[____])
%load solutions/01-3.py
```
## Changing a value in a list
```
numbers
numbers[4]
# Who thinks this will work
numbers[4] = 5
numbers
numbers[:2]
numbers[:2] = [1, 'TWO']
numbers
```
## Tuples
Just like lists, except they are immutable, i.e. read only
tuples are defined with a () or just by comma separating python objects
```
simple_tuple = (1, 2, 3)
numbers_tuple = 'one', 'two', 'three', 'four', 'five', 'six', 'seven', 'eight', 'nine', 'ten'
mixed_tuple = (1, 2, 3, 'four', 'five', 'six', False)
len(numbers_tuple)
numbers_tuple[5:]
numbers_tuple.append(11)
numbers_tuple[5] = 5
## Strings are immutable sequences
pi = 'The Ratio of the circumference of a circle to its diameter'
pi[0]
pi[-1]
pi[10:20]
pi[-8:]
pi[5] = 'x'
```
### Remember lists & tuples are sequences of *any* python object, including themselves
lists of lists, lists of tuples, tuples of lists aaargh!!!!!!
```
coordinates = [(-98.2, 28.3), [-97.4, 28.9], [-96.4, 26.5]]
print(coordinates)
coordinates[1]
coordinates[1][0]
# how can I access the 8?
nested = [1, 2, [4, 5], [(6, [7, 8])], 9, 10]
# print(nested[?])
```
## Loops
In Python you can loop through any kind of sequence
or in technical terms any *iterable*
```
for char in 'Hello':
print('The current character is: ', char)
for n in numbers:
print('The current number is: ', n)
coordinates
# What do you think this prints?
for coord in coordinates:
print('The current coord is:', coord)
for lon, lat in coordinates:
print('The current longitude is:', lon)
print('The current latitude is:', lat)
print('-------------------------------')
# swap
a = 5
b = 10
print('a=%s, b=%s' % (a,b))
a, b = b, a
print('a=%s, b=%s' % (a,b))
coordinates_3d = [[-98.2, 28.3], [-97.4, 28.9, 500], [-96.4, 26.5]]
for lon, lat in coordinates_3d:
print('The current longitude is:', lon)
print('The current latitude is:', lat)
print('-------------------------------')
```
### Exercise 1.4
1. Create a list of coordinates & altitudes in feet:
-98.4, 28.2, 500
-93.2, 28.8, 524
-95.3, 29.6, 552
2. Loop through the list print the altitudes in meters (1 ft=0.3048 m)
```
coordinates = ________
for _____ in _______:
print(______)
# load solution
%load solutions/01-4.py
```
## Conditionals
```
a = 5
if a>10:
print('a is greater than ten')
if a>10:
print('a is greater than ten')
else:
print('a is not greater than ten')
if a>10:
print('a is greater than ten')
elif a<5:
print('a is less than five')
else:
print('a is between five and ten')
if a < 10 and a >= 5:
print('Down with conditionals!')
```
** NOTE: Blocks of code in python are delimited by indentation **
```python
print('Start!') # <- main program
for number in [1,2,3,4,5]: # <- start of the loop block
print(number) # <- part of the loop
square = number * number # <- part of the loop
if square > 9: # <- part of the loop, start of the 'if' block
print('square is greater than 9') # <- part of the loop, part of the 'if' block
square = 9 # <- part of the loop, part of the 'if' block
print(square) # <- part of the loop
print('Done!') # <- main program
```
** This is why you must use a python 'aware' editor **
### Exercise 1.5
Given the string: 'It was the Best of Times, It was the Worst of Times'
Count how many words begin with a capital letter and also add those words to
a list called capwords.
Hints:
- There is a string method that can convert the string to a list of words. Try and find it.
- There is also a string method to check in a word starts with a capital letter OR you can check if the string is all lowercase.
- You can create an empty list using [] or list()
Bonus: only store unique words in the capwords
```
text = 'It was the Best of Times It was the Worst of Times'
words = text.____
count = 0
capwords = []
for word in words:
_______
_______
print('Number of Capitalalized Words =', count)
print('Capitalized Word List = ', capwords)
# load solution
%load solutions/01-5.py
```
## Dictionaries
Dictionaries are defined using curly brackets {}
```
students = ['Lucy', 'Tom', 'Mary', 'Jack', 'Sarah', 'Peter',]
scores = [97, 82, 68, 79, 92, 89]
# What is Jack's Grade?
# Did Tom do better than Sarah?
print('Jacks Score =', scores[3])
print('Tom did better than Sarah=', scores[1]>scores[4])
scores = {'Lucy': 97, 'Tom': 82, 'Mary': 68, 'Jack': 79, 'Sarah': 92, 'Peter': 89}
print('Jacks Score =', scores['Jack'])
print('Tom did better Sarah =', scores['Tom']>scores['Sarah'])
scores.keys()
scores.values()
scores.items()
list(scores.keys())
```
** NOTE: dictionaries don't have an inherent order **
** You can use any immutable (i.e. read only) python object as a dictionary key **
```
mixed_dict = {1: 'hello', 'two': (3,4,5), (3, 'A'): 3.1415}
for key, value in mixed_dict.items():
print('key->', key, ', value->', value)
# Using a Dictionary as a lookup table
# very contrived example.
data = [(5, 'ft'), (10,'m'), (9, 'in'), (30, 'cm')]
conv_factor = {'ft': 0.3048, 'm': 1, 'cm': 0.01, 'in': 0.0254}
converted_data = []
for value, unit in data:
print('value=', value, 'unit=', unit, 'conversion factor=', conv_factor[unit])
converted_data.append((value*conv_factor[unit], 'm'))
print(converted_data)
```
## Functions
```
def my_add(a, b):
c = a + b
return c
my_add(4, 5.0)
def my_sub(a, b):
return a - b
my_sub(10, 5)
my_sub(b=10, a=5)
# functions need not take in any values or for that matter return anything
def five():
return 5
def alert():
print('Red Alert!!!')
a = five()
print('a=', a)
b = alert()
print('b=', b)
# you can return multiple things
def stuff(mystr):
return mystr.upper(), mystr.lower(), len(mystr)
a = stuff('Hello')
print(a)
a, b, c = stuff('Goodbye')
print(a)
print(b)
print(c)
```
### positional vs keyword arguments
```
def my_add(a, b=2, display=False):
result = a + b
if display:
print('a + b =', result)
return result
my_add(4, 5)
my_add(4)
my_add(4, display=True)
my_add(4, 5, display=True)
```
### Exercise 1.6
Write a function that takes in two strings and an optional flag called
reverse that defaults to False.
The function should return the longer string unless reverse is True in
which case it should return the shorter string
```
def get_longest(____):
____
return ____
assert get_longest('hello', 'goodbye') == 'goodbye'
assert get_longest('hello', 'goodbye', reverse=True) == 'hello'
%load solutions/01-6.py
```
## city temperature data
```
import numpy
numpy.loadtxt('data/city_temperatures_2015.csv', delimiter=',')
temperatures = numpy.loadtxt('data/city_temperatures_2015.csv', delimiter=',')
import matplotlib.pyplot as plt
plt.imshow(temperatures)
plt.show()
temperatures.min()
plt.plot(temperatures[0,:])
plt.show()
plt.bar([0, 1, 2, 3, 4, 5, 6, 7, 8, 9], temperatures.min(axis=1))
plt.show()
```
| github_jupyter |
<h1>Table of Contents<span class="tocSkip"></span></h1>
<div class="toc"><ul class="toc-item"><li><span><a href="#create-a-dataset-with-two-stars-at-the-end-of-both-entities" data-toc-modified-id="create-a-dataset-with-two-stars-at-the-end-of-both-entities-1"><span class="toc-item-num">1 </span>create a dataset with two stars at the end of both entities</a></span></li><li><span><a href="#Create-a-dataset-with-a-star-at-the-end-of-first-entity" data-toc-modified-id="Create-a-dataset-with-a-star-at-the-end-of-first-entity-2"><span class="toc-item-num">2 </span>Create a dataset with a star at the end of first entity</a></span></li><li><span><a href="#Read-translated-datasets-in-German" data-toc-modified-id="Read-translated-datasets-in-German-3"><span class="toc-item-num">3 </span>Read translated datasets in German</a></span></li><li><span><a href="#Process-translated-german-sentences" data-toc-modified-id="Process-translated-german-sentences-4"><span class="toc-item-num">4 </span>Process translated german sentences</a></span></li><li><span><a href="#create-dictionary-of-label-pairs" data-toc-modified-id="create-dictionary-of-label-pairs-5"><span class="toc-item-num">5 </span>create dictionary of label pairs</a></span></li><li><span><a href="#Filter" data-toc-modified-id="Filter-6"><span class="toc-item-num">6 </span>Filter</a></span></li></ul></div>
```
import pandas as pd
import numpy as np
import re
with open('data/semeval/test_file_full.txt') as f:
eng_small = f.readlines()
sentences_raw = eng_small[::4] # get strings with sentences
labels = eng_small[1::4] # get labels
sentences_ent = [re.findall(r'(?<=\t")(.*)(?="\n)', string)[0] for string in sentences_raw] # extract sentences
def extract_entities(lst_with_tags):
e1_de = [re.findall(r'(?<=<e1>)(.*)(?=</e1>)', string) for string in lst_with_tags] # get entity 1
e2_de = [re.findall(r'(?<=<e2>)(.*)(?=</e2>)', string) for string in lst_with_tags] # get entity 2
return e1_de, e2_de
# preprocess English dataset
sentences_ent = [string.rstrip() for string in sentences_ent] # extract sentences
e1, e2 = extract_entities(sentences_ent)
```
### create a dataset with two stars at the end of both entities
```
# replace entity markers with stars for correct translation
sentences_ = [re.sub(r'<e1>',"",st) for st in sentences_ent]
sentences_ = [re.sub(r'<e2>',"",st) for st in sentences_]
sentences_ = [re.sub(r'</e1>',"*",st) for st in sentences_]
sentences_ = [re.sub(r'</e2>','*',st) for st in sentences_]
# save redacted sentences
with open('data/semeval_de_new/test_sentences_en_stars.txt', 'w') as f:
f.writelines(line + '\n' for line in sentences_)
```
### Create a dataset with a star at the end of first entity
```
# replace entity markers with stars for correct translation
sentences_1 = [re.sub(r'<e1>',"",st) for st in sentences_ent]
sentences_1 = [re.sub(r'<e2>',"",st) for st in sentences_1]
sentences_1 = [re.sub(r'</e1>',"*",st) for st in sentences_1]
sentences_1 = [re.sub(r'</e2>','',st) for st in sentences_1]
# save redacted sentences
with open('data/semeval_de_new/test_sentences_en_star1.txt', 'w') as f:
f.writelines(line + '\n' for line in sentences_1)
```
### Read translated datasets in German
**Note**: This part should be replaced with working API for DeepL translations. However, the API is not freely available, therefore this step is currently done manually.
```
# read translated sentences in German (2stars)
with open('data/semeval_de_new/test_sentences_de_stars.txt', 'r', encoding = 'utf-8') as f:
de_small = f.readlines()
# read translated sentences in German (1star)
with open('data/semeval_de_new/test_sentences_de_star1.txt', 'r', encoding = 'utf-8') as f:
de_small1 = f.readlines()
```
### Process translated german sentences
```
# replace * with entity end tags
def return_entity_tags(lst_with_stars):
sentences_de = []
for i, sent_de in enumerate(lst_with_stars):
sent = sent_de.rstrip()
# delete special quotation marks
sent = re.sub(r'\"', "", sent)
# add a space at the beginning of each sentence
sent = re.sub(r'^', ' ', sent)
#replace e1
sent = re.sub(r'\*',"</e1>",sent, 1)
sent = re.sub(r'[ ](?=[a-zA-Z\u00C0-\u017F\'-\/0-9\&]+<\/e1>)', ' <e1>', sent)
#replace e2
sent = re.sub(r'\*',"</e2>",sent)
sent = re.sub(r'[ ](?=[a-zA-Z\u00C0-\u017F\'-\/0-9\&]+<\/e2>)', ' <e2>', sent)
sentences_de.append(sent)
return sentences_de
sentences_de = return_entity_tags(de_small) # with stars at the end of both entities
sentences_de1 = return_entity_tags(de_small1) # with one star for 1st entity
#test
for i, sent in enumerate(sentences_de):
print(i,sent)
e1_de, e2_de = extract_entities(sentences_de)
e1_de1, e2_de1 = extract_entities(sentences_de1)
e1_missing = []
for i, word in enumerate(e1_de):
if len(word) == 0:
e1_missing.append(i)
e2_missing = []
for i, word in enumerate(e2_de):
if len(word) == 0:
e2_missing.append(i)
len(e1_missing), len(e2_missing)
#compare e2 from the first list with e1 from first to identify changed labels
changed_labels = [i for i in range(len(e2_de)) if (e2_de[i] == e1_de1[i])&(labels[i]!='Other\n')]
len(changed_labels)
# test
for i in changed_labels:
print(i,
labels[i],
sentences_ent[i],
de_small[i]
)
labels[191]
```
### create dictionary of label pairs
```
set(labels)
opp_dir_rel = {'Cause-Effect(e1,e2)\n': 'Cause-Effect(e2,e1)\n',
'Cause-Effect(e2,e1)\n': 'Cause-Effect(e1,e2)\n',
'Component-Whole(e1,e2)\n': 'Component-Whole(e2,e1)\n',
'Component-Whole(e2,e1)\n': 'Component-Whole(e1,e2)\n',
'Content-Container(e1,e2)\n': 'Content-Container(e2,e1)\n',
'Content-Container(e2,e1)\n': 'Content-Container(e1,e2)\n',
'Entity-Destination(e1,e2)\n': 'Entity-Destination(e2,e1)\n',
'Entity-Destination(e2,e1)\n': 'Entity-Destination(e1,e2)\n',
'Entity-Origin(e1,e2)\n': 'Entity-Origin(e2,e1)\n',
'Entity-Origin(e2,e1)\n': 'Entity-Origin(e1,e2)\n',
'Instrument-Agency(e1,e2)\n': 'Instrument-Agency(e2,e1)\n',
'Instrument-Agency(e2,e1)\n': 'Instrument-Agency(e1,e2)\n',
'Member-Collection(e1,e2)\n': 'Member-Collection(e2,e1)\n',
'Member-Collection(e2,e1)\n': 'Member-Collection(e1,e2)\n',
'Message-Topic(e1,e2)\n': 'Message-Topic(e2,e1)\n',
'Message-Topic(e2,e1)\n': 'Message-Topic(e1,e2)\n',
'Product-Producer(e1,e2)\n': 'Product-Producer(e2,e1)\n',
'Product-Producer(e2,e1)\n': 'Product-Producer(e1,e2)\n',
'Other\n': 'Other\n'
}
labels_new = labels
for i in changed_labels:
labels_new[i] = opp_dir_rel[labels_new[i]]
```
### Filter
```
sentences_de_final = [sentences_de[i] for i in range(len(sentences_de)) if i not in e2_missing]
labels_de_final = [labels_new[i] for i in range(len(sentences_de)) if i not in e2_missing]
len(sentences_de_final), len(labels_de_final)
combined = [str(str(i)+' "'+sentences_de_final[i]+'"'+'\n'+labels_de_final[i]+'Comment: \n'+'\n') for i in range(len(sentences_de_final))]
# save redacted sentences
with open('data/semeval_de_new/final/test_file_de_final.txt', 'w', encoding = 'utf-8') as f:
f.writelines(line for line in combined)
# for review save also initial german with english version
sentences_en_final = [sentences_ent[i] for i in range(len(sentences_ent)) if i not in e2_missing]
labels_en_final = [labels[i] for i in range(len(sentences_ent)) if i not in e2_missing]
comb_review = [str(str(i)
+' "'
+sentences_de_final[i]
+'"'+'\n'
+labels_de_final[i]
+'"'
+sentences_en_final[i]
+'"'
+'\n'
+'Old Label: '
+labels_en_final[i]
+'Comment:'
+'\n\n')
for i in range(len(sentences_de_final))]
# save redacted sentences
with open('data/semeval_de_new/review/test_for_review.txt', 'w', encoding = 'utf-8') as f:
f.writelines(line for line in comb_review)
```
| github_jupyter |
# Training a better model
```
from theano.sandbox import cuda
%matplotlib inline
import utils; reload(utils)
from utils import *
from __future__ import division, print_function
#path = "data/dogscats/sample/"
path = "data/dogscats/"
model_path = path + 'models/'
if not os.path.exists(model_path): os.mkdir(model_path)
batch_size=64
```
## Are we underfitting?
Our validation accuracy so far has generally been higher than our training accuracy. That leads to two obvious questions:
1. How is this possible?
2. Is this desirable?
The answer to (1) is that this is happening because of *dropout*. Dropout refers to a layer that randomly deletes (i.e. sets to zero) each activation in the previous layer with probability *p* (generally 0.5). This only happens during training, not when calculating the accuracy on the validation set, which is why the validation set can show higher accuracy than the training set.
The purpose of dropout is to avoid overfitting. By deleting parts of the neural network at random during training, it ensures that no one part of the network can overfit to one part of the training set. The creation of dropout was one of the key developments in deep learning, and has allowed us to create rich models without overfitting. However, it can also result in underfitting if overused, and this is something we should be careful of with our model.
So the answer to (2) is: this is probably not desirable. It is likely that we can get better validation set results with less (or no) dropout, if we're seeing that validation accuracy is higher than training accuracy - a strong sign of underfitting. So let's try removing dropout entirely, and see what happens!
(We had dropout in this model already because the VGG authors found it necessary for the imagenet competition. But that doesn't mean it's necessary for dogs v cats, so we will do our own analysis of regularization approaches from scratch.)
## Removing dropout
Our high level approach here will be to start with our fine-tuned cats vs dogs model (with dropout), then fine-tune all the dense layers, after removing dropout from them. The steps we will take are:
- Re-create and load our modified VGG model with binary dependent (i.e. dogs v cats)
- Split the model between the convolutional (*conv*) layers and the dense layers
- Pre-calculate the output of the conv layers, so that we don't have to redundently re-calculate them on every epoch
- Create a new model with just the dense layers, and dropout p set to zero
- Train this new model using the output of the conv layers as training data.
As before we need to start with a working model, so let's bring in our working VGG 16 model and change it to predict our binary dependent...
```
model = vgg_ft(2)
```
...and load our fine-tuned weights.
```
model.load_weights(model_path+'finetune3.h5')
```
We're going to be training a number of iterations without dropout, so it would be best for us to pre-calculate the input to the fully connected layers - i.e. the *Flatten()* layer. We'll start by finding this layer in our model, and creating a new model that contains just the layers up to and including this layer:
```
layers = model.layers
last_conv_idx = [index for index,layer in enumerate(layers)
if type(layer) is Convolution2D][-1]
last_conv_idx
layers[last_conv_idx]
conv_layers = layers[:last_conv_idx+1]
conv_model = Sequential(conv_layers)
# Dense layers - also known as fully connected or 'FC' layers
fc_layers = layers[last_conv_idx+1:]
```
Now we can use the exact same approach to creating features as we used when we created the linear model from the imagenet predictions in the last lesson - it's only the model that has changed. As you're seeing, there's a fairly small number of "recipes" that can get us a long way!
```
batches = get_batches(path+'train', shuffle=False, batch_size=batch_size)
val_batches = get_batches(path+'valid', shuffle=False, batch_size=batch_size)
val_classes = val_batches.classes
trn_classes = batches.classes
val_labels = onehot(val_classes)
trn_labels = onehot(trn_classes)
val_features = conv_model.predict_generator(val_batches, val_batches.nb_sample)
trn_features = conv_model.predict_generator(batches, batches.nb_sample)
save_array(model_path + 'train_convlayer_features.bc', trn_features)
save_array(model_path + 'valid_convlayer_features.bc', val_features)
trn_features = load_array(model_path+'train_convlayer_features.bc')
val_features = load_array(model_path+'valid_convlayer_features.bc')
trn_features.shape
```
For our new fully connected model, we'll create it using the exact same architecture as the last layers of VGG 16, so that we can conveniently copy pre-trained weights over from that model. However, we'll set the dropout layer's p values to zero, so as to effectively remove dropout.
```
# Copy the weights from the pre-trained model.
# NB: Since we're removing dropout, we want to half the weights
def proc_wgts(layer): return [o/2 for o in layer.get_weights()]
# Such a finely tuned model needs to be updated very slowly!
opt = RMSprop(lr=0.00001, rho=0.7)
def get_fc_model():
model = Sequential([
MaxPooling2D(input_shape=conv_layers[-1].output_shape[1:]),
Flatten(),
Dense(4096, activation='relu'),
Dropout(0.),
Dense(4096, activation='relu'),
Dropout(0.),
Dense(2, activation='softmax')
])
for l1,l2 in zip(model.layers, fc_layers): l1.set_weights(proc_wgts(l2))
model.compile(optimizer=opt, loss='categorical_crossentropy', metrics=['accuracy'])
return model
fc_model = get_fc_model()
```
And fit the model in the usual way:
```
fc_model.fit(trn_features, trn_labels, nb_epoch=8,
batch_size=batch_size, validation_data=(val_features, val_labels))
fc_model.save_weights(model_path+'no_dropout.h5')
fc_model.load_weights(model_path+'no_dropout.h5')
```
# Reducing overfitting
Now that we've gotten the model to overfit, we can take a number of steps to reduce this.
## Approaches to reducing overfitting
We do not necessarily need to rely on dropout or other regularization approaches to reduce overfitting. There are other techniques we should try first, since regularlization, by definition, biases our model towards simplicity - which we only want to do if we know that's necessary. This is the order that we recommend using for reducing overfitting (more details about each in a moment):
1. Add more data
2. Use data augmentation
3. Use architectures that generalize well
4. Add regularization
5. Reduce architecture complexity.
We'll assume that you've already collected as much data as you can, so step (1) isn't relevant (this is true for most Kaggle competitions, for instance). So the next step (2) is data augmentation. This refers to creating additional synthetic data, based on reasonable modifications of your input data. For images, this is likely to involve one or more of: flipping, rotation, zooming, cropping, panning, minor color changes.
Which types of augmentation are appropriate depends on your data. For regular photos, for instance, you'll want to use horizontal flipping, but not vertical flipping (since an upside down car is much less common than a car the right way up, for instance!)
We recommend *always* using at least some light data augmentation, unless you have so much data that your model will never see the same input twice.
## About data augmentation
Keras comes with very convenient features for automating data augmentation. You simply define what types and maximum amounts of augmentation you want, and keras ensures that every item of every batch randomly is changed according to these settings. Here's how to define a generator that includes data augmentation:
```
# dim_ordering='tf' uses tensorflow dimension ordering,
# which is the same order as matplotlib uses for display.
# Therefore when just using for display purposes, this is more convenient
gen = image.ImageDataGenerator(rotation_range=10, width_shift_range=0.1,
height_shift_range=0.1, width_zoom_range=0.2, shear_range=0.15, zoom_range=0.1,
channel_shift_range=10., horizontal_flip=True, dim_ordering='tf')
```
Let's take a look at how this generator changes a single image (the details of this code don't matter much, but feel free to read the comments and keras docs to understand the details if you're interested).
```
# Create a 'batch' of a single image
img = np.expand_dims(ndimage.imread('data/dogscats/test/7.jpg'),0)
# Request the generator to create batches from this image
aug_iter = gen.flow(img)
# Get eight examples of these augmented images
aug_imgs = [next(aug_iter)[0].astype(np.uint8) for i in range(8)]
# The original
plt.imshow(img[0])
```
As you can see below, there's no magic to data augmentation - it's a very intuitive approach to generating richer input data. Generally speaking, your intuition should be a good guide to appropriate data augmentation, although it's a good idea to test your intuition by checking the results of different augmentation approaches.
```
# Augmented data
plots(aug_imgs, (20,7), 2)
# Ensure that we return to theano dimension ordering
K.set_image_dim_ordering('th')
```
## Adding data augmentation
Let's try adding a small amount of data augmentation, and see if we reduce overfitting as a result. The approach will be identical to the method we used to finetune the dense layers in lesson 2, except that we will use a generator with augmentation configured. Here's how we set up the generator, and create batches from it:
```
gen = image.ImageDataGenerator(rotation_range=15, width_shift_range=0.1,
height_shift_range=0.1, zoom_range=0.1, horizontal_flip=True)
batches = get_batches(path+'train', gen, batch_size=batch_size)
# NB: We don't want to augment or shuffle the validation set
val_batches = get_batches(path+'valid', shuffle=False, batch_size=batch_size)
```
When using data augmentation, we can't pre-compute our convolutional layer features, since randomized changes are being made to every input image. That is, even if the training process sees the same image multiple times, each time it will have undergone different data augmentation, so the results of the convolutional layers will be different.
Therefore, in order to allow data to flow through all the conv layers and our new dense layers, we attach our fully connected model to the convolutional model--after ensuring that the convolutional layers are not trainable:
```
fc_model = get_fc_model()
for layer in conv_model.layers: layer.trainable = False
# Look how easy it is to connect two models together!
conv_model.add(fc_model)
```
Now we can compile, train, and save our model as usual - note that we use *fit_generator()* since we want to pull random images from the directories on every batch.
```
conv_model.compile(optimizer=opt, loss='categorical_crossentropy', metrics=['accuracy'])
conv_model.fit_generator(batches, samples_per_epoch=batches.nb_sample, nb_epoch=8,
validation_data=val_batches, nb_val_samples=val_batches.nb_sample)
conv_model.fit_generator(batches, samples_per_epoch=batches.nb_sample, nb_epoch=3,
validation_data=val_batches, nb_val_samples=val_batches.nb_sample)
conv_model.save_weights(model_path + 'aug1.h5')
conv_model.load_weights(model_path + 'aug1.h5')
```
## Batch normalization
### About batch normalization
Batch normalization (*batchnorm*) is a way to ensure that activations don't become too high or too low at any point in the model. Adjusting activations so they are of similar scales is called *normalization*. Normalization is very helpful for fast training - if some activations are very high, they will saturate the model and create very large gradients, causing training to fail; if very low, they will cause training to proceed very slowly. Furthermore, large or small activations in one layer will tend to result in even larger or smaller activations in later layers, since the activations get multiplied repeatedly across the layers.
Prior to the development of batchnorm in 2015, only the inputs to a model could be effectively normalized - by simply subtracting their mean and dividing by their standard deviation. However, weights in intermediate layers could easily become poorly scaled, due to problems in weight initialization, or a high learning rate combined with random fluctuations in weights.
Batchnorm resolves this problem by normalizing each intermediate layer as well. The details of how it works are not terribly important (although I will outline them in a moment) - the important takeaway is that **all modern networks should use batchnorm, or something equivalent**. There are two reasons for this:
1. Adding batchnorm to a model can result in **10x or more improvements in training speed**
2. Because normalization greatly reduces the ability of a small number of outlying inputs to over-influence the training, it also tends to **reduce overfitting**.
As promised, here's a brief outline of how batchnorm works. As a first step, it normalizes intermediate layers in the same way as input layers can be normalized. But this on its own would not be enough, since the model would then just push the weights up or down indefinitely to try to undo this normalization. Therefore, batchnorm takes two additional steps:
1. Add two more trainable parameters to each layer - one to multiply all activations to set an arbitrary standard deviation, and one to add to all activations to set an arbitary mean
2. Incorporate both the normalization, and the learnt multiply/add parameters, into the gradient calculations during backprop.
This ensures that the weights don't tend to push very high or very low (since the normalization is included in the gradient calculations, so the updates are aware of the normalization). But it also ensures that if a layer does need to change the overall mean or standard deviation in order to match the output scale, it can do so.
### Adding batchnorm to the model
We can use nearly the same approach as before - but this time we'll add batchnorm layers (and dropout layers):
```
conv_layers[-1].output_shape[1:]
def get_bn_layers(p):
return [
MaxPooling2D(input_shape=conv_layers[-1].output_shape[1:]),
Flatten(),
Dense(4096, activation='relu'),
BatchNormalization(),
Dropout(p),
Dense(4096, activation='relu'),
BatchNormalization(),
Dropout(p),
Dense(1000, activation='softmax')
]
def load_fc_weights_from_vgg16bn(model):
"Load weights for model from the dense layers of the Vgg16BN model."
# See imagenet_batchnorm.ipynb for info on how the weights for
# Vgg16BN can be generated from the standard Vgg16 weights.
from vgg16bn import Vgg16BN
vgg16_bn = Vgg16BN()
_, fc_layers = split_at(vgg16_bn.model, Convolution2D)
copy_weights(fc_layers, model.layers)
p=0.6
bn_model = Sequential(get_bn_layers(0.6))
load_fc_weights_from_vgg16bn(bn_model)
def proc_wgts(layer, prev_p, new_p):
scal = (1-prev_p)/(1-new_p)
return [o*scal for o in layer.get_weights()]
for l in bn_model.layers:
if type(l)==Dense: l.set_weights(proc_wgts(l, 0.5, 0.6))
bn_model.pop()
for layer in bn_model.layers: layer.trainable=False
bn_model.add(Dense(2,activation='softmax'))
bn_model.compile(Adam(), 'categorical_crossentropy', metrics=['accuracy'])
bn_model.fit(trn_features, trn_labels, nb_epoch=8,
validation_data=(val_features, val_labels))
bn_model.save_weights(model_path+'bn.h5')
bn_model.load_weights(model_path+'bn.h5')
bn_layers = get_bn_layers(0.6)
bn_layers.pop()
bn_layers.append(Dense(2,activation='softmax'))
final_model = Sequential(conv_layers)
for layer in final_model.layers: layer.trainable = False
for layer in bn_layers: final_model.add(layer)
for l1,l2 in zip(bn_model.layers, bn_layers):
l2.set_weights(l1.get_weights())
final_model.compile(optimizer=Adam(),
loss='categorical_crossentropy', metrics=['accuracy'])
final_model.fit_generator(batches, samples_per_epoch=batches.nb_sample, nb_epoch=1,
validation_data=val_batches, nb_val_samples=val_batches.nb_sample)
final_model.save_weights(model_path + 'final1.h5')
final_model.fit_generator(batches, samples_per_epoch=batches.nb_sample, nb_epoch=4,
validation_data=val_batches, nb_val_samples=val_batches.nb_sample)
final_model.save_weights(model_path + 'final2.h5')
final_model.optimizer.lr=0.001
final_model.fit_generator(batches, samples_per_epoch=batches.nb_sample, nb_epoch=4,
validation_data=val_batches, nb_val_samples=val_batches.nb_sample)
bn_model.save_weights(model_path + 'final3.h5')
for
```
| github_jupyter |
<a href="https://colab.research.google.com/github/kalz2q/mycolabnotebooks/blob/master/numpyexercises.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# 100 numpy exercises
<https://github.com/rougier/numpy-100>
にあったもののコピー。
とりあえず使い方がわからない。まあいいか。やってみよう。
(1) numpyをインポートする。
```
import numpy as np
```
# (2) numpy のバージョンと configuration を print する。
```
print (np.__version__)
# np.show_config()
# 実験: numpy がどんな関数やメソッドを持っているかは dir(np) とすればわかる。version も、show_config もある。
# dir(np)
```
(3) サイズ $10$ の null ベクトルを作る。
```
import numpy as np
Z = np.zeros(10)
print(Z)
# 実験
print (type(Z))
print(Z == [0,0,0,0,0,0,0,0,0,0])
print(len(Z))
print(len([0,0,0,0,0,0,0,0,0,0]))
print(list(Z) == [0,0,0,0,0,0,0,0,0,0])
print(np.array([1,2,3]) == np.array([1,2,3]))
print(np.array((1,2,3)))
```
#### (4) 配列 array のメモリーサイズを調べる方法。
```
Z = np.zeros((10,10))
print("%d bytes" % (Z.size * Z.itemsize))
# 実験
print (Z.size)
print(Z.itemsize)
```
#### (5) numpy の add 関数のドキュメントを得る。
次のコメントアウトした %run `python "import numpy; numpy.info(numpy.add)"` は使えなかった。
help(np.add)
でいいと思う。
```
# %run `python "import numpy; numpy.info(numpy.add)"`
# 実験
# np.info(np.add)
help(np.add)
```
#### (6) 5番目のアイテムの値を 1 にする。
```
Z[4] = 1
print(Z)
```
#### (7) 値が10から49のベクトルを作る。
```
Z = np.arange(10,50)
print(Z)
# 実験
print(range(10,50))
print(list(range(10,50)))
# print(np.ndarray(list(range(10,50))))
print(np.array(list(range(10,50))))
print(type(np.array(list(range(10,50)))))
```
#### (8) ベクトルを逆順にする。
```
Z = np.arange(10)
Z = Z[::-1]
print(Z)
# 実験
Z = list(range(10))
Z.reverse()
print(Z)
Y = np.array([3,2,1,0,1,1,5])
print(Y)
Y = Y[::-1]
print(Y)
```
#### (9) 3x3 の行列を作る。
```
import numpy as np
Z = np.arange(9).reshape(3, 3)
Z
np.array([0,1,2,3,4,5,6,7,8]).reshape(3,3)
# 実験
arr = np.array([[0, 1, 2], [3, 4, 5], [6, 7, 8]])
print(arr)
mat = np.matrix([[0, 1, 2], [3, 4, 5], [6, 7, 8]])
print(mat)
print(type(mat))
```
#### (10) ゼロでない要素のインデクスを見つける。 [1,2,0,0,4,0]
```
import numpy as np
nz = np.nonzero([1,2,0,0,4,0])
print(nz)
# 実験
print(list(enumerate([1,2,0,0,4,0])))
print([x for (x,y) in list(enumerate([1,2,0,0,4,0])) if y>0])
print([x for (x,y) in list(enumerate([1,2,0,0,-4,0])) if y!=0])
```
#### (11) 3x3 の単位行列を作る。
```
Z = np.eye(3)
print(Z)
# 実験
print(np.identity(3))
```
#### (12) 要素がランダムな 3x3x3 の行列を作る。
```
Z = np.random.random((3,3,3))
print(Z)
# 実験
help(np.random.random)
# 実験
print([int(x*10) for x in np.random.random(9)])
np.array([int(x*10) for x in np.random.random(9)]).reshape(3,3)
```
#### (13) 10x10 のランダムな値の配列を作って、その最大値と最小値を調べる。
```
Z = np.random.random((10,10))
Zmin, Zmax = Z.min(), Z.max()
print(Zmin, Zmax)
```
#### (14) ランダムな値のサイズ 30 のベクトルを作って平均を調べる。
```
Z = np.random.random(30)
m = Z.mean()
print(m)
# 実験
sum(list(Z)) / 30
# 実験
import numpy as np
print(type(np.random.random()))
help(np.random.random)
```
#### (15) 上下左右の端が 1 で、中が 0 の配列を作る。
```
Z = np.ones((10,10))
Z[1:-1,1:-1] = 0
print(Z)
# 実験
z = np.array([1 for x in range(100)]).reshape(10,10)
z.reshape(10,10)
z[1:-1,1:-1] = 0
print(z)
```
#### (16) すでにある配列に 0 で縁をつける。
```
Z = np.ones((5,5))
Z = np.pad(Z, pad_width=1, mode='constant', constant_values=0)
print(Z)
# # Using fancy indexing
# Z = np.ones((5,5))
# Z[:, [0, -1]] = 0
# Z[[0, -1], :] = 0
# print(Z)
# 実験
Z = np.ones((5,5))
Z=[[0]+list(l)+[0] for l in Z]
Z= [[0]*len(Z[0])]+Z+[[0]*len(Z[0])]
Z
```
#### (17) np.nan について調べる。
np.nan は値を持たない float。
欠損値。
NaN = Not a Number
```
import numpy as np
print(0*np.nan) #=> nan
print(np.nan==np.nan) #=> False
print(np.inf > np.nan) #=> False
print(np.nan - np.nan) #=> nan
print(np.nan in set([np.nan])) #=> True
print(0.3 == 3 * 0.1) #=> False
# 実験
print()
print(np.nan) #=> nan
print(type(np.nan)) #=> <class 'float'>
print(True if np.nan else False) #=> True
print(None) #=> None
print(type(None)) #=> <class 'NoneType'>
print(True if None else False) #=> False
print()
print(np.isnan((np.nan)))
%%writefile height.csv
name,height
Ahmad,175.2
Eli,
Kobe,180.8
%ls
# 実験
import pandas as pd
import numpy as np
df = pd.read_csv('height.csv')
print(df)
print()
for idx in df.index:
print(np.isnan(df.loc[idx, 'height']))
```
#### (18) 5x5 の行列で、対角線の1つ下に 1,2,3,4 の値を入れる。
```
Z = np.diag(1+np.arange(4),k=-1)
print(Z)
# 実験
print(np.diag(np.array([1,2,3,4]), k=-1))
print(np.arange(4))
print(1+np.arange(4))
print(np.array([1+x for x in range(4)]))
```
#### (19) 8x8 のチェッカーボードパターンの行列を作る。
```
Z = np.zeros((8,8),dtype=int)
Z[1::2,::2] = 1
Z[::2,1::2] = 1
print(Z)
# 実験
Z = np.zeros((8,8),dtype=int)
Z[1::2,0::3] = 1
print(Z)
```
#### (20) 6x7x8 の形の配列の 100 番目の座標 x,y,z を求める。
```
print(np.unravel_index(99,(6,7,8)))
# 実験
print(type(np.unravel_index(99,(6,7,8))))
print(np.where(np.array(range(6*7*8)).reshape(6,7,8) == 99))
tuple([int(x) for x in np.where(np.array(range(6*7*8)).reshape(6,7,8) == 99)])
```
#### (21) tile 関数を使って 8x8 の行列を作る。
```
Z = np.tile( np.array([[0,1],[1,0]]), (4,4))
print(Z)
# 実験
Z = np.tile( np.array([[0,8],[8,8,8]]), (2,4))
print(Z)
print()
Z = np.tile( np.array([[0,8],[8,8]]), (2,4))
print(Z)
```
#### (22) 5x5 のランダムな値の行列を正規化 normalize する。
```
Z = np.random.random((5,5))
Z = (Z - np.mean (Z)) / (np.std (Z))
print(Z)
# 実験
Z = np.random.randint(10, size=(5,5))
print(Z)
print()
Z = (Z - np.mean (Z))
print(Z)
```
#### (23) カスタム dtype を作り、4つの unsigned bytes (RGBA) で色を表現する。
```
# color = np.dtype([("r", np.ubyte, 1),
# ("g", np.ubyte, 1),
# ("b", np.ubyte, 1),
# ("a", np.ubyte, 1)])
color = np.dtype([("r", np.ubyte),
("g", np.ubyte),
("b", np.ubyte),
("a", np.ubyte)])
print(color)
# 実験
a = np.array([1,2,3])
print(a)
print(a.dtype)
a = np.array([1,2.0,3.5])
print(a.dtype)
a = np.array([1,2,3], dtype='int32')
print(a.dtype)
a = np.array([1,2,3], dtype='float')
print(a)
print(a.dtype)
a = np.array([1,2.0,3.5], dtype='int')
print(a)
print(a.dtype)
f = np.array([0, 3, 0, -1], dtype = 'bool')
print(f)
```
#### (24) 5x3 の行列と 3x2 の行列の積を求める。
```
Z = np.dot(np.ones((5,3)), np.ones((3,2)))
print(Z)
# Alternative solution, in Python 3.5 and above
Z = np.ones((5,3)) @ np.ones((3,2))
print(Z)
```
#### (25) 1次元の配列で、3 と 8 の間の値の要素をマイナスにする。
```
Z = np.arange(11)
Z[(3 < Z) & (Z < 8)] *= -1
print(Z)
# 実験
[(-x if (x>3)&(x<8) else x) for x in range(11)]
```
#### (26) 次のプログラムを実行して考察する。
```python
# Author: Jake VanderPlas
print(sum(range(5),-1)) #=> 9
from numpy import *
print(sum(range(5),-1)) #=> 10
```
```
# 実験
print(sum(range(5),-1))
import numpy as np
print(np.sum(range(5),-1))
arr = np.array([[0, 1, 2], [3, 4, 5], [6, 7, 8]])
print(arr)
print(np.sum(arr))
print(np.sum(arr, -1))
print(np.sum(arr, -2))
```
#### (27) 整数のベクトル Z について、次の演算は文法的に可能かどうか。
```python
Z**Z #=> [1 4 27]
2 << Z >> 2 #=> [1 2 4]
Z <- Z #=> [False False Fale]
1j*Z #=> [0.+1.j 0.+2.j 0.+3.j]
Z/1/1 #=> [1. 2. 3.]
Z<Z>Z #=> Error
```
```
import numpy as np
Z = np.array([1,2,3])
print(Z)
print(Z**Z)
print(2 << Z >> 2)
print(Z <- Z)
print(1j*Z)
print(Z/1/1)
# Z<Z>Z #=> Error
# 実験
print(2 << 3) #=> 16
print(16 >> 2) #=> 4
print(3 < 3) #=> False
print(3 <- 3) #=> False
print(1/1) #=> 1.1
print(all(Z<(Z + 1))) #=> True
```
#### (28) 次の式を実行して考察する。
```python
np.array(0) / np.array(0) #=> Error
np.array(0) // np.array(0) #=> Error
np.array([np.nan]).astype(int).astype(float) #=> [-9.22337204e+18]
```
```
# np.array(0) / np.array(0) #=> Error
# np.array(0) // np.array(0) #=> Error
print(np.array([np.nan]).astype(int).astype(float))
print(np.array([np.nan]).astype(int))
print(np.array([np.nan]))
print(np.array([1,2,3]).astype(int).astype(float))
print(type(np.array([np.nan])))
```
#### (29) float の配列を絶対値で切り上げる方法 (ゼロから遠くする)
```
# Author: Charles R Harris
import numpy as np
Z = np.random.uniform(-10,+10,10)
print(np.copysign(np.ceil(np.abs(Z)), Z))
# More readable but less efficient
print(np.where(Z>0, np.ceil(Z), np.floor(Z)))
# 実験
print(np.array([np.ceil(x) if x > 0 else np.floor(x) for x in Z]))
# 実験
print(Z)
Z= np.random.random(10)*20-10
print(Z)
```
#### (30) 2つの入れるの共通の値をみつける。
How to find common values between two arrays? (★☆☆)
```
Z1 = np.random.randint(0,10,10)
Z2 = np.random.randint(0,10,10)
print(np.intersect1d(Z1,Z2))
# 実験
print(np.unique([1,2,3,2,3]))
s1 = np.array([1,2,3,2,3,4])
s2 = np.array([2,3,4,5,6,7,])
print(np.union1d(s1,s2))
print(np.in1d([3,7], s1))
print(np.intersect1d(s1,s2))
print(np.setdiff1d(s2,s1))
print(np.setxor1d(s1,s2))
```
#### (31) numpy の警告を無視する方法。
How to ignore all numpy warnings (not recommended)? (★☆☆)
```
# Suicide mode on
defaults = np.seterr(all="ignore")
Z = np.ones(1) / 0
# Back to sanity
_ = np.seterr(**defaults)
# Equivalently with a context manager
with np.errstate(all="ignore"):
np.arange(3) / 0
# 実験
print(np.ones((3,3)))
print(np.arange((9)).reshape(3,3))
```
#### (32) 次の式の真偽を試して考察する。
Is the following expressions true? (★☆☆)
```python
np.sqrt(-1) == np.emath.sqrt(-1) #=> False なぜなら np.sqrt(-1) は 複素数を表さないが、np.emath.sqrt(-1) は複素数だから。
```
```
# 実験
print(np.emath.sqrt(-1) == 0+1j)
print(np.emath.sqrt(-1) == 1j)
print((1j)**2)
print(np.sqrt(-1))
```
#### (33) 昨日、今日、明日の日付を表示する。
```
yesterday = np.datetime64('today') - np.timedelta64(1)
today = np.datetime64('today')
tomorrow = np.datetime64('today') + np.timedelta64(1)
print(yesterday)
print(today)
print(tomorrow)
```
#### (34) 2016年7月の日付をすべて列挙する。
```
Z = np.arange('2016-07', '2016-08', dtype='datetime64[D]')
print(Z)
```
#### (35) 行列演算 ((A+B)*(-A/2)) を in place で実行する。in place とはコピーを作らずに計算すること。
```
A = np.ones(3)*1
B = np.ones(3)*2
C = np.ones(3)*3
print(np.add(A,B,out=B))
print(np.divide(A,2,out=A))
print(np.negative(A,out=A))
print(np.multiply(A,B,out=A))
# 実験
A = np.ones(3)*1
B = np.ones(3)*2
C = np.ones(3)*3
print((A+B)*(-A/2))
```
#### (36) ランダムな正の数の配列から、整数部分をだけにする方法を4通り示す。
```
Z = np.random.uniform(0,10,10)
print(Z - Z%1)
print(Z // 1)
print(np.floor(Z))
print(Z.astype(int))
print(np.trunc(Z))
```
#### (37) 5x5 でそれぞれの行が 0 1 2 3 4 である行列を作る。
```
Z = np.zeros((5,5))
Z += np.arange(5)
print(Z)
# 実験
import numpy as np
Z = np.zeros((5,5))
Z += range(5,10)
print(Z)
print(np.transpose(Z))
```
#### (38) 10個の整数を作る generator 関数を作り、配列を作るのに使ってみる。
```
def generate():
for x in range(10):
yield x
Z = np.fromiter(generate(),dtype=float,count=-1)
print(Z)
# 実験
def generate():
for x in range(20):
yield x*x
Z = np.fromiter(generate(),dtype=int,count=5)
print(Z)
```
#### (39) 0 から 1 の値のサイズ 10 のベクトルを作る。0 と 1 は含まれないものとする。
```
Z = np.linspace(0,1,11,endpoint=False)[1:]
print(Z)
# 実験
Z = np.linspace(0,1,11)
print(Z)
Z = np.linspace(0,1,11)[1:-1]
print(Z)
Z = np.linspace(0,1,12)[1:-1]
print(Z)
```
#### (40) ランダムな値のサイズ 10 のベクトルを作り、ソートする。
```
import numpy as np
Z = np.random.random(10)
Z.sort()
print(Z)
```
#### (41) 小さな配列について、np.sum より速い方法。
```
# Author: Evgeni Burovski
Z = np.arange(10)
np.add.reduce(Z)
# 実験
import numpy as np
from functools import reduce
Z = np.arange(10)
print(reduce(np.add, Z))
print(reduce(lambda a, b: a+b, Z))
```
#### (42) A と B の2つの配列が同等 equal かどうかを判別する。
```
import numpy as np
A = np.random.randint(0,2,5)
B = np.random.randint(0,2,5)
# Assuming identical shape of the arrays and a tolerance for the comparison of values
equal = np.allclose(A,B)
print(equal)
# Checking both the shape and the element values, no tolerance (values have to be exactly equal)
equal = np.array_equal(A,B)
print(equal)
# 実験
C = np.array([0,0,0,1,1])
D = np.array([0,0,0,1,1.000000001])
equal = np.allclose(C, D)
print(equal)
equal = np.array_equal(C, D)
print(equal)
```
#### (43) 配列をイミュータブル(変更不可)にする。
```
Z = np.zeros(10)
Z.flags.writeable = False
# Z[0] = 1 #=> ValueError: assignment destination is read-only
print(Z)
```
#### (44) デカルト座標を表す 10x2 の行列を極座標に変換する。
```
Z = np.random.random((10,2))
print(Z)
X,Y = Z[:,0], Z[:,1]
R = np.sqrt(X**2+Y**2)
T = np.arctan2(Y,X)
print(R)
print(T)
# 実験
print(np.rad2deg(np.arctan2(1, 1)))
print(np.rad2deg(np.arctan(1)))
print(np.array(list (map (np.array, list(zip(R, T))))))
```
#### (45) ランダムなサイズ 10 のベクトルを作り、最大値を 0 に置き換える。
```
import numpy as np
Z = np.random.random(10)
Z[Z.argmax()] = 0
print(Z)
# 実験
z = np.array([3, 2, 1, 1, 3])
print(z)
print(z.argmax())
print(z.argmin())
y=[3, 2, 1, 1, 3]
print(y)
print(y.index(max(y)))
y[y.index(max(y))] = 0
print(y)
```
#### (46) x 座標と y 座標を持つ構造化配列 structured array を作り、[0,1]x[0,1] の範囲をカバーする。
```
import numpy as np
Z = np.zeros((5,5), [('x',float),('y',float)])
Z['x'], Z['y'] = np.meshgrid(np.linspace(0,1,5),
np.linspace(0,1,5))
print(Z)
np.linspace(0,1,5)
# 実験
dtype = [('x',float),('y',float)]
Y = np.zeros((5,5), dtype=dtype)
Y['x'] = np.linspace(0,1,5)
np.transpose(Y)['y'] = np.linspace(0,1,5)
print(Y)
```
#### (47) 行列 X と Y からコーシー Cauchy 行列 C (Cij =1/(xi - yj)) を作る。
```
# Author: Evgeni Burovski
import numpy as np
X = np.arange(8)
Y = X + 0.5
C = 1.0 / np.subtract.outer(X, Y)
print(np.linalg.det(C))
# 実験
X = np.arange(3)
Y = X + 0.5
print(X)
print(Y)
C = 1.0 / np.subtract.outer(X, Y)
print(C.size)
print(C)
print(np.linalg.det(C))
```
#### (48) numpy のすべてのスカラー型について、最大値、最小値を表示する。
```
for dtype in [np.int8, np.int32, np.int64]:
print(np.iinfo(dtype).min)
print(np.iinfo(dtype).max)
for dtype in [np.float32, np.float64]:
print(np.finfo(dtype).min)
print(np.finfo(dtype).max)
print(np.finfo(dtype).eps) # machine epsilon?
# 実験
print(np.iinfo(np.int8))
print(np.finfo(np.float32))
```
#### (49) 配列のすべての値を表示する。
```
import numpy as np
np.set_printoptions(threshold=float("inf"))
Z = np.zeros((16,16))
print(Z)
# 実験
import numpy as np
np.set_printoptions(threshold=float("10"))
Z = np.zeros((16,16))
print(Z)
```
#### (50) 与えられた値に一番近い値をベクトルの中で探して表示する。
How to find the closest value (to a given scalar) in a vector? (★★☆)
```
Z = np.arange(100)
v = np.random.uniform(0,100)
index = (np.abs(Z-v)).argmin()
print(Z[index])
# 実験
print(Z)
print(v)
print(index)
```
#### (51) 構造化配列で、場所 position (x,y) と色 color (r,g,b) を表現する。
```
Z = np.zeros(10, [ ('position', [ ('x', float),
('y', float)]),
('color', [ ('r', float),
('g', float),
('b', float)])])
print(Z)
```
#### (52) 座標を表すランダムな (10,2) の形のベクトルを考え、点と点の間の距離を計算する。
```
Z = np.random.random((10,2))
X,Y = np.atleast_2d(Z[:,0], Z[:,1])
D = np.sqrt( (X-X.T)**2 + (Y-Y.T)**2)
print(D)
# Much faster with scipy
import scipy
# Thanks Gavin Heverly-Coulson (#issue 1)
import scipy.spatial
# Z = np.random.random((10,2))
D = scipy.spatial.distance.cdist(Z,Z)
print(D)
# 実験
# Z = np.random.random((10,2))
X,Y = np.atleast_2d(Z[:,0], Z[:,1])
X = np.array([Z[:,0]])
Y = np.array([Z[:,1]])
D = np.sqrt( (X-X.T)**2 + (Y-Y.T)**2)
print(D)
```
#### (53) 32 ビット float を 32 ビット integer に in place で変換する。
```
# Thanks Vikas (https://stackoverflow.com/a/10622758/5989906)
# & unutbu (https://stackoverflow.com/a/4396247/5989906)
Z = (np.random.rand(10)*100).astype(np.float32)
Y = Z.view(np.int32)
Y[:] = Z
print(Y)
# 実験
Z=(np.random.rand(10)*100).astype(np.float32)
print(Z)
# Y=Z.view(np.int32)
Y=np.zeros(10).astype(np.int32)
print(Y)
Y[:] = Z
print(Y)
```
#### (54) 次のようなファイルをどう読み込むか。
```
1, 2, 3, 4, 5
6, , , 7, 8
, , 9,10,11
```
```
from io import StringIO
# Fake file
s = StringIO('''1, 2, 3, 4, 5
6, , , 7, 8
, , 9,10,11
''')
Z = np.genfromtxt(s, delimiter=",", dtype=np.int)
print(Z)
# 実験
import numpy as np
import re
def read_data(data):
lines = []
for line in data.split('\n'):
items = []
for item in re.split(',| ,\s',line):
if item == '' or item == ' ':
item = -1
items.append(item)
line=items
lines.append(line)
return lines
data = '''1, 2, 3, 4, 5
6, , , 7, 8
, , 9,10,11'''
print(read_data(data))
```
#### (55) numpy の配列で python の enumerate に相当するものは何か。
```
Z = np.arange(9).reshape(3,3)
for index, value in np.ndenumerate(Z):
print(index, value)
for index in np.ndindex(Z.shape):
print(index, Z[index])
# 実験
print(Z)
print(list((np.ndenumerate(Z))))
print(np.array(list((np.ndenumerate(Z)))).reshape(3,3,2))
```
#### (56) 一般的な2Dガウス配列を作る。
```
import numpy as np
X, Y = np.meshgrid(np.linspace(-1,1,10), np.linspace(-1,1,10))
D = np.sqrt(X*X+Y*Y)
sigma, mu = 1.0, 0.0
G = np.exp(-( (D-mu)**2 / ( 2.0 * sigma**2 ) ) )
print(G)
```
上の解答の意味がわからない。 問題のガウス配列 Gaussian array というのは正規分布のことで、2D というので普通の正規分布のグラフのことを言っているのかと思うのだが、解答 meshgrid を使っているので 3D グラフを描こうとしているのではないか。
ガウス関数は
$$
f(x) = \frac{1}{\sqrt{2\pi\sigma}}\mathrm{exp} \left(- \frac{(x - \mu)^2}{2 \sigma^2}\right)\quad (x \in \mathbb{R})
$$
特に $\mu=0,\, \sigma^2 = 1$ のとき標準正規分布と呼び、
$$
f(x) = \frac{1}{\sqrt{2\pi\sigma}}\mathrm{exp} \left(- \frac{x^2}{2}\right)\quad (x \in \mathbb{R})
$$
とのこと。
この通りに作ればいいだけだよね。
```
# 実験
import numpy as np
X, Y = np.meshgrid(np.linspace(-1,1,100), np.linspace(-1,1,100))
D = np.sqrt(X*X+Y*Y)
sigma, mu = 1.0, 0.0
G = np.exp(- 10 * ( (D-mu)**2 / ( 2.0 * sigma**2 ) ) )
# print(G)
import matplotlib.pyplot as plt
from matplotlib import cm
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
ax.plot_surface(X,Y,G, cmap=cm.jet)
plt.show()
# 実験
import matplotlib.pyplot as plt
import numpy as np
import math
def f(x):
return (math.exp(-x**2/2)) / math.sqrt(2*math.pi)
n = np.linspace(-5.0, 5.0, 50)
p = []
for i in range(len(n)):
p.append(f(n[i]))
# グラフに表示
plt.plot(n, p)
plt.show()
```
#### (57) 2次元の配列で、p 個の要素をランダムに置く。
```
# Author: Divakar
n = 10
p = 3
Z = np.zeros((n,n))
np.put(Z, np.random.choice(range(n*n), p, replace=False),1)
print(Z)
# 実験
n = 5
p = 5
Z = np.full((n,n),'□')
# np.put(Z, np.random.choice(range(n*n), p, replace=False),1)
np.put(Z, np.random.choice(25, p, replace=False),'■')
print(Z)
np.put(Z, np.random.choice(25, p, replace=False),'■')
print(Z)
np.put(Z, np.random.choice(25, p, replace=False),'■')
np.put(Z, np.random.choice(25, p, replace=False),'■')
np.put(Z, np.random.choice(25, p, replace=False),'■')
print(Z)
```
#### (58) 行列の各行の平均を引く。
```
# Author: Warren Weckesser
X = np.random.rand(5, 10)
# Recent versions of numpy
Y = X - X.mean(axis=1, keepdims=True)
# Older versions of numpy
# Y = X - X.mean(axis=1).reshape(-1, 1)
print(Y)
# 実験
print(X[0] - np.array([np.mean(row) for row in X])[0])
Y = X - X.mean(axis=1, keepdims=True)
print(Y[0])
print([[X[n] - np.array([np.mean(row) for row in X])[n]] for n in range(5)])
```
#### (59) n 番目の列で配列をソートする。
```
# Author: Steve Tjoa
Z = np.random.randint(0,10,(3,3))
print(Z)
print(Z[Z[:,1].argsort()])
# 実験
print(Z)
a=Z[:,2].argsort()
print(a)
print(Z[a])
```
#### (60) 2次元配列について、null 列があるかどうかを判定する。
```
# Author: Warren Weckesser
Z = np.random.randint(0,3,(3,10))
print((~Z.any(axis=0)).any())
# 実験
Z=np.array([0,0,1,0]).reshape(2,2)
print(Z)
print(Z.any(axis=0))
print(~Z.any(axis=0))
print(~np.array([True, False]))
print(np.array(list(map (lambda x: not(x), Z.any(axis=0)))))
print(~np.array(list(map (lambda x: not(x), Z.any(axis=0)))))
print(np.array(list(map (lambda x: not(x), Z.any(axis=0)))).any())
```
#### (61) 与えられた値に最も近い値を配列の中からみつける。
Find the nearest value from a given value in an array (★★☆)
```
Z = np.random.uniform(0,1,10)
z = 0.5
m = Z.flat[np.abs(Z - z).argmin()]
print(m)
# 実験
Z = np.random.uniform(0,1,9).reshape(3,3)
z = 0.5
m = Z.flat[np.abs(Z - z).argmin()]
print(Z)
print(np.abs(Z - z).argmin())
print(m)
```
#### (62) 配列のシェープが (1,3) のものと (3,1) のものがあるとして、iterator を使って合計を計算する。
```
import numpy as np
A = np.arange(3).reshape(3,1)
B = np.arange(3).reshape(1,3)
it = np.nditer([A,B,None])
for x,y,z in it: z[...] = x + y
print(it.operands[2])
# 実験
a = [0,1,2]
b = [0,1,2]
np.array([x+y for x in a for y in b]).reshape(3,3)
```
#### (63) name 属性を持つ配列クラスを作る。
Create an array class that has a name attribute (★★☆)
```
import numpy as np
class NamedArray(np.ndarray):
def __new__(cls, array, name="no name"):
obj = np.asarray(array).view(cls)
obj.__class__.name = name
return obj
def __array_finalize__(self, obj):
if obj is None: return
self.info = getattr(obj, 'name', "noo name")
Z = NamedArray(np.arange(10), "range_10")
print(Z.__class__.name)
print (Z.name)
# 実験
Z = NamedArray(np.arange(10))
Z.name = "range_10"
print(Z.__class__.name)
print(Z.name)
Z = NamedArray(None)
Z.name = "range_10"
print(Z.__class__.name)
print(Z.name)
Z = NamedArray("yes", name="no")
Z.name = "range_10"
print(Z.__class__.name)
print(Z.name)
type (Z)
```
通常クラスのインスタンスは何もしないでも、name 属性がある。
```
# 実験
class MyClass:
pass
m = MyClass()
m.name = "myname"
print(m.name)
```
設問の意味はクラスに name 属性をつけたい、ということか。 でも、解答例の
print(Z.name)
は、インスタンスの name を調べているだけで、クラスの name を調べているわけではない。
プログラムは、`__init__()` ではなく、`__new__()` で初期化することにより、self ではなく cls に属性をつける風に読める。
``` python
help(np.ndarray.view) #=> New view of array with the same data.
```
なので、view() はなにもしていない。
```
__array_finalize__(self, obj):
if obj is None: return
self.info = getattr(obj, 'name', "noo name")
```
は何をしているのか?
```
%%script false
help(getattr)
```
#### (64) 与えられたベクトルについて、インデクスのベクトルで指定された要素に 1 を加える。
Consider a given vector, how to add 1 to each element indexed by a second vector (be careful with repeated indices)? (★★★)
```
import numpy as np
# Author: Brett Olsen
Z = np.ones(10)
I = np.random.randint(0,len(Z),20)
Z += np.bincount(I, minlength=len(Z))
print(Z)
# Another solution
# Author: Bartosz Telenczuk
np.add.at(Z, I, 1)
print(Z)
# 実験
Z = np.ones(10)
print(Z)
I = np.random.randint(0,len(Z),3)
print(I)
Z += np.bincount(I, minlength=len(Z))
print(np.bincount(I))
print(Z)
print()
print(Z)
print(I)
np.add.at(Z, I, 1)
print(Z)
```
#### (65) インデクスリスト (I) に基づいて、ベクトル (X) の要素を足しこんで、配列 (F) を作る。
```
# Author: Alan G Isaac
X = [1,2,3,4,5,6]
I = [1,3,9,3,4,1]
F = np.bincount(I,X)
print(F)
# 実験
X = [1,2,3,4,5,6]
I = [1,3,9,3,4,1]
print(np.bincount(I))
F = np.bincount(I,X)
print(F)
```
#### (66) (w, h, 3) の配列の形で表される色配列がある。 ユニークな色の数を求める。 (dtype=ubyte) とする
Considering a (w,h,3) image of (dtype=ubyte), compute the number of unique colors (★★★)
```
# Author: Nadav Horesh
import numpy as np
w,h = 16,16
I = np.random.randint(0,2,(h,w,3)).astype(np.ubyte)
F = I[...,0]*256*256 + I[...,1]*256 +I[...,2]
n = len(np.unique(F))
print(np.unique(I))
```
この模範解答では、実行結果が [0 1] となってしまうので、解答になっていないと思う。
模範解答では w と h を 16 と置いているので、$16 \times 16 \times 3 = 768$ の場所に、多分仮に、だろうが 0 と 1 を置いている。 ということは、2価なので、例えば赤 1,緑 1, 青 0 とすると、多分黄色になる。 順列組み合わせで
0 0 0
0 0 1
0 1 0
0 1 1
1 0 0
1 0 1
1 1 0
1 1 1
で最大 8 種類、もしくはそれ以下の結果になれば正解だと思う。
まず、
`F = I[...,0]*256*256 + I[...,1]*256 +I[...,2]`
が機能していない。 やりたいことは、$16 \times 16 \times 3$ の配列について、0 行目に 256 × 256、1 行目に 256 を掛けて、3 行を足すことによって、色の表現を配列から単なるフラットな数字に変換しようとしているのだろう。
しかし、ubyte、である数字に大きな数を掛けて ubyte の範囲を超えると現在の仕様では 0 になってしまう。
その証拠に print(n) とすると、 [0 0], [0 1], [1 0], [0 0] の4つが取り出されている。
```
print(n)
```
ので、この方法は使えない。 とりあえず 256 を 10 にすると 大丈夫そう。
```
# 実験
I[...,0]*3
# 実験
I[...,0]*256*256
# 実験
I[...,0]*10*10
# 実験
import numpy as np
w,h = 16,16
I = np.random.randint(0,2,(h,w,3)).astype(np.ubyte)
F = I[...,0]*10*10 + I[...,1]*10 +I[...,2]
n = len(np.unique(F))
print(np.unique(I))
print(n)
```
上記の結果で print(n) の値が 8 になるのは 8 が最大値で試験回数が多いからなので、仮に w,h = 4.4 として実行すると8も出るし、6とか7も出る。
`print(np.unique(F)) `
の値が [0 1] になってしまうのは、np.unique の仕様が、配列をフラットにして unique をとる。
```
# 実験
import numpy as np
print (np.unique(np.array([1,2,3,2,1])))
print(np.unique(np.array([[1,2],[1,2],[2,3]])))
print(np.unique(np.array([[1,2],[1,2],[2,3]]), axis=0))
```
下記の例では、一旦 reshape して、axis を指定して unique をとった。
```
# 実験
import numpy as np
w,h = 4,4
I = np.random.randint(0,2,(h,w,3)).astype(np.ubyte)
F = I[...,0]*10*10 + I[...,1]*10 +I[...,2]
n = len(np.unique(F))
print(n)
print(np.unique(F))
np.unique(np.reshape(I, (w*h, 3)), axis=0)
```
#### (67) 4次元の配列を考え、最後の 2 軸の合計をとる。
Considering a four dimensions array, how to get sum over the last two axis at once? (★★★)
```
import numpy as np
A = np.random.randint(0,10,(3,4,3,4))
# solution by passing a tuple of axes (introduced in numpy 1.7.0)
sum = A.sum(axis=(-2,-1))
print(sum)
# solution by flattening the last two dimensions into one
# (useful for functions that don't accept tuples for axis argument)
sum = A.reshape(A.shape[:-2] + (-1,)).sum(axis=-1)
print(sum)
```
4次元で数字が多いとイメージできないので、少なくして実験してみる。
```
# 実験
# 3次元にして、それぞれの要素を2にしてみる。
A = np.random.randint(0,10,(2,2,2))
print(A)
print()
sum = A.sum(axis=(-2,-1)) # これが正解
print(sum) # axis を指定しないとフラットな合計になってしまい、今回の目的に合わない
print(A.sum())
print(A.sum(axis=(0,1))) # 頭から足して行くとこうなる
print(np.sum([[1,2],[3,4]], axis=(0)))
print(np.sum([[1,2],[3,4]], axis=(-1)))
print()
sum = A.reshape(A.shape[:-2] + (-1,)).sum(axis=-1)
print(sum)
```
`sum = A.reshape(A.shape[:-2] + (-1,)).sum(axis=-1)`
の reshape と shape がわからないので以下に実験してみる。
```
# 実験
import numpy as np
A = np.random.randint(0,10,(3,4,5,6,7,8))
print(A.shape[:-2]+(-1,))
print(type(A.shape[:-2]))
print((1,)+(2,)+(3,))
print()
A = np.random.randint(0,10,(3,4,5))
print(A.reshape((6,10)))
print()
A.reshape(3,-1)
# 実験
A = np.random.randint(0,10,(2,2,2))
print(A)
print()
A=A.reshape(A.shape[:-2] + (-1,))
print(A)
sum = A.sum(axis=-1)
print(sum)
```
# いまここ
#### (68) 1次元のベクトルDを考え、インデックスのベクトルSを使用して、Dのサブセットの平均を計算する。
Considering a one-dimensional vector D, how to compute means of subsets of D using a vector S of same size describing subset indices? (★★★)
```
# Author: Jaime Fernández del Río
import numpy as np
D = np.random.uniform(0,1,100)
S = np.random.randint(0,10,100)
D_sums = np.bincount(S, weights=D)
D_counts = np.bincount(S)
D_means = D_sums / D_counts
print(D_means)
# Pandas solution as a reference due to more intuitive code
import pandas as pd
print(pd.Series(D).groupby(S).mean())
np.random.uniform(-10,10,10)
```
#### (69) How to get the diagonal of a dot product? (★★★)
```
# Author: Mathieu Blondel
A = np.random.uniform(0,1,(5,5))
B = np.random.uniform(0,1,(5,5))
# Slow version
np.diag(np.dot(A, B))
# Fast version
np.sum(A * B.T, axis=1)
# Faster version
np.einsum("ij,ji->i", A, B)
```
#### (70) Consider the vector [1, 2, 3, 4, 5], how to build a new vector with 3 consecutive zeros interleaved between each value? (★★★)
```
# Author: Warren Weckesser
Z = np.array([1,2,3,4,5])
nz = 3
Z0 = np.zeros(len(Z) + (len(Z)-1)*(nz))
Z0[::nz+1] = Z
print(Z0)
```
#### (71) Consider an array of dimension (5,5,3), how to mulitply it by an array with dimensions (5,5)? (★★★)
```
A = np.ones((5,5,3))
B = 2*np.ones((5,5))
print(A * B[:,:,None])
```
#### (72) How to swap two rows of an array? (★★★)
```
# Author: Eelco Hoogendoorn
A = np.arange(25).reshape(5,5)
A[[0,1]] = A[[1,0]]
print(A)
```
#### (73) Consider a set of 10 triplets describing 10 triangles (with shared vertices), find the set of unique line segments composing all the triangles (★★★)
```
# Author: Nicolas P. Rougier
faces = np.random.randint(0,100,(10,3))
F = np.roll(faces.repeat(2,axis=1),-1,axis=1)
F = F.reshape(len(F)*3,2)
F = np.sort(F,axis=1)
G = F.view( dtype=[('p0',F.dtype),('p1',F.dtype)] )
G = np.unique(G)
print(G)
```
#### (74) Given an array C that is a bincount, how to produce an array A such that np.bincount(A) == C? (★★★)
```
# Author: Jaime Fernández del Río
C = np.bincount([1,1,2,3,4,4,6])
A = np.repeat(np.arange(len(C)), C)
print(A)
```
#### (75) How to compute averages using a sliding window over an array? (★★★)
```
# Author: Jaime Fernández del Río
def moving_average(a, n=3) :
ret = np.cumsum(a, dtype=float)
ret[n:] = ret[n:] - ret[:-n]
return ret[n - 1:] / n
Z = np.arange(20)
print(moving_average(Z, n=3))
```
#### (76) Consider a one-dimensional array Z, build a two-dimensional array whose first row is (Z[0],Z[1],Z[2]) and each subsequent row is shifted by 1 (last row should be (Z[-3],Z[-2],Z[-1]) (★★★)
```
# Author: Joe Kington / Erik Rigtorp
from numpy.lib import stride_tricks
def rolling(a, window):
shape = (a.size - window + 1, window)
strides = (a.itemsize, a.itemsize)
return stride_tricks.as_strided(a, shape=shape, strides=strides)
Z = rolling(np.arange(10), 3)
print(Z)
```
#### (77) How to negate a boolean, or to change the sign of a float inplace? (★★★)
```
# Author: Nathaniel J. Smith
Z = np.random.randint(0,2,100)
np.logical_not(Z, out=Z)
Z = np.random.uniform(-1.0,1.0,100)
np.negative(Z, out=Z)
```
#### (78) Consider 2 sets of points P0,P1 describing lines (2d) and a point p, how to compute distance from p to each line i (P0[i],P1[i])? (★★★)
```
def distance(P0, P1, p):
T = P1 - P0
L = (T**2).sum(axis=1)
U = -((P0[:,0]-p[...,0])*T[:,0] + (P0[:,1]-p[...,1])*T[:,1]) / L
U = U.reshape(len(U),1)
D = P0 + U*T - p
return np.sqrt((D**2).sum(axis=1))
P0 = np.random.uniform(-10,10,(10,2))
P1 = np.random.uniform(-10,10,(10,2))
p = np.random.uniform(-10,10,( 1,2))
print(distance(P0, P1, p))
```
#### (79) Consider 2 sets of points P0,P1 describing lines (2d) and a set of points P, how to compute distance from each point j (P[j]) to each line i (P0[i],P1[i])? (★★★)
```
# Author: Italmassov Kuanysh
# based on distance function from previous question
P0 = np.random.uniform(-10, 10, (10,2))
P1 = np.random.uniform(-10,10,(10,2))
p = np.random.uniform(-10, 10, (10,2))
print(np.array([distance(P0,P1,p_i) for p_i in p]))
```
#### (80) Consider an arbitrary array, write a function that extract a subpart with a fixed shape and centered on a given element (pad with a `fill` value when necessary) (★★★)
```
# Author: Nicolas Rougier
Z = np.random.randint(0,10,(10,10))
shape = (5,5)
fill = 0
position = (1,1)
R = np.ones(shape, dtype=Z.dtype)*fill
P = np.array(list(position)).astype(int)
Rs = np.array(list(R.shape)).astype(int)
Zs = np.array(list(Z.shape)).astype(int)
R_start = np.zeros((len(shape),)).astype(int)
R_stop = np.array(list(shape)).astype(int)
Z_start = (P-Rs//2)
Z_stop = (P+Rs//2)+Rs%2
R_start = (R_start - np.minimum(Z_start,0)).tolist()
Z_start = (np.maximum(Z_start,0)).tolist()
R_stop = np.maximum(R_start, (R_stop - np.maximum(Z_stop-Zs,0))).tolist()
Z_stop = (np.minimum(Z_stop,Zs)).tolist()
r = [slice(start,stop) for start,stop in zip(R_start,R_stop)]
z = [slice(start,stop) for start,stop in zip(Z_start,Z_stop)]
R[r] = Z[z]
print(Z)
print(R)
```
#### (81) Consider an array Z = [1,2,3,4,5,6,7,8,9,10,11,12,13,14], how to generate an array R = [[1,2,3,4], [2,3,4,5], [3,4,5,6], ..., [11,12,13,14]]? (★★★)
```
# Author: Stefan van der Walt
Z = np.arange(1,15,dtype=np.uint32)
R = stride_tricks.as_strided(Z,(11,4),(4,4))
print(R)
```
#### (82) Compute a matrix rank (★★★)
```
# Author: Stefan van der Walt
Z = np.random.uniform(0,1,(10,10))
U, S, V = np.linalg.svd(Z) # Singular Value Decomposition
rank = np.sum(S > 1e-10)
print(rank)
```
#### (83) How to find the most frequent value in an array?
```
Z = np.random.randint(0,10,50)
print(np.bincount(Z).argmax())
```
#### (84) Extract all the contiguous 3x3 blocks from a random 10x10 matrix (★★★)
```
# Author: Chris Barker
Z = np.random.randint(0,5,(10,10))
n = 3
i = 1 + (Z.shape[0]-3)
j = 1 + (Z.shape[1]-3)
C = stride_tricks.as_strided(Z, shape=(i, j, n, n), strides=Z.strides + Z.strides)
print(C)
```
#### (85) Create a 2D array subclass such that Z[i,j] == Z[j,i] (★★★)
```
# Author: Eric O. Lebigot
# Note: only works for 2d array and value setting using indices
class Symetric(np.ndarray):
def __setitem__(self, index, value):
i,j = index
super(Symetric, self).__setitem__((i,j), value)
super(Symetric, self).__setitem__((j,i), value)
def symetric(Z):
return np.asarray(Z + Z.T - np.diag(Z.diagonal())).view(Symetric)
S = symetric(np.random.randint(0,10,(5,5)))
S[2,3] = 42
print(S)
```
#### (86) Consider a set of p matrices wich shape (n,n) and a set of p vectors with shape (n,1). How to compute the sum of of the p matrix products at once? (result has shape (n,1)) (★★★)
```
# Author: Stefan van der Walt
p, n = 10, 20
M = np.ones((p,n,n))
V = np.ones((p,n,1))
S = np.tensordot(M, V, axes=[[0, 2], [0, 1]])
print(S)
# It works, because:
# M is (p,n,n)
# V is (p,n,1)
# Thus, summing over the paired axes 0 and 0 (of M
```
#### (87) Consider a 16x16 array, how to get the block-sum (block size is 4x4)? (★★★)
```
# Author: Robert Kern
Z = np.ones((16,16))
k = 4
S = np.add.reduceat(np.add.reduceat(Z, np.arange(0, Z.shape[0], k), axis=0),
np.arange(0, Z.shape[1], k), axis=1)
print(S)
```
#### (88) How to implement the Game of Life using numpy arrays? (★★★)
```
# Author: Nicolas Rougier
def iterate(Z):
# Count neighbours
N = (Z[0:-2,0:-2] + Z[0:-2,1:-1] + Z[0:-2,2:] +
Z[1:-1,0:-2] + Z[1:-1,2:] +
Z[2: ,0:-2] + Z[2: ,1:-1] + Z[2: ,2:])
# Apply rules
birth = (N==3) & (Z[1:-1,1:-1]==0)
survive = ((N==2) | (N==3)) & (Z[1:-1,1:-1]==1)
Z[...] = 0
Z[1:-1,1:-1][birth | survive] = 1
return Z
Z = np.random.randint(0,2,(50,50))
for i in range(100): Z = iterate(Z)
print(Z)
```
#### (89) How to get the n largest values of an array (★★★)
```
Z = np.arange(10000)
np.random.shuffle(Z)
n = 5
# Slow
print (Z[np.argsort(Z)[-n:]])
# Fast
print (Z[np.argpartition(-Z,n)[:n]])
```
#### (90) Given an arbitrary number of vectors, build the cartesian product (every combinations of every item) (★★★)
```
# Author: Stefan Van der Walt
def cartesian(arrays):
arrays = [np.asarray(a) for a in arrays]
shape = (len(x) for x in arrays)
ix = np.indices(shape, dtype=int)
ix = ix.reshape(len(arrays), -1).T
for n, arr in enumerate(arrays):
ix[:, n] = arrays[n][ix[:, n]]
return ix
print (cartesian(([1, 2, 3], [4, 5], [6, 7])))
```
#### (91) How to create a record array from a regular array? (★★★)
```
Z = np.array([("Hello", 2.5, 3),
("World", 3.6, 2)])
R = np.core.records.fromarrays(Z.T,
names='col1, col2, col3',
formats = 'S8, f8, i8')
print(R)
```
#### (92) Consider a large vector Z, compute Z to the power of 3 using 3 different methods (★★★)
```
# Author: Ryan G.
x = np.random.rand(int(5e7))
%timeit np.power(x,3)
%timeit x*x*x
%timeit np.einsum('i,i,i->i',x,x,x)
```
#### (93) Consider two arrays A and B of shape (8,3) and (2,2). How to find rows of A that contain elements of each row of B regardless of the order of the elements in B? (★★★)
```
# Author: Gabe Schwartz
A = np.random.randint(0,5,(8,3))
B = np.random.randint(0,5,(2,2))
C = (A[..., np.newaxis, np.newaxis] == B)
rows = np.where(C.any((3,1)).all(1))[0]
print(rows)
```
#### (94) Considering a 10x3 matrix, extract rows with unequal values (e.g. [2,2,3]) (★★★)
```
# Author: Robert Kern
Z = np.random.randint(0,5,(10,3))
print(Z)
# solution for arrays of all dtypes (including string arrays and record arrays)
E = np.all(Z[:,1:] == Z[:,:-1], axis=1)
U = Z[~E]
print(U)
# soluiton for numerical arrays only, will work for any number of columns in Z
U = Z[Z.max(axis=1) != Z.min(axis=1),:]
print(U)
```
#### (95) Convert a vector of ints into a matrix binary representation (★★★)
```
# Author: Warren Weckesser
I = np.array([0, 1, 2, 3, 15, 16, 32, 64, 128])
B = ((I.reshape(-1,1) & (2**np.arange(8))) != 0).astype(int)
print(B[:,::-1])
# Author: Daniel T. McDonald
I = np.array([0, 1, 2, 3, 15, 16, 32, 64, 128], dtype=np.uint8)
print(np.unpackbits(I[:, np.newaxis], axis=1))
```
#### (96) Given a two dimensional array, how to extract unique rows? (★★★)
```
# Author: Jaime Fernández del Río
Z = np.random.randint(0,2,(6,3))
T = np.ascontiguousarray(Z).view(np.dtype((np.void, Z.dtype.itemsize * Z.shape[1])))
_, idx = np.unique(T, return_index=True)
uZ = Z[idx]
print(uZ)
# Author: Andreas Kouzelis
# NumPy >= 1.13
uZ = np.unique(Z, axis=0)
print(uZ)
```
#### (97) Considering 2 vectors A & B, write the einsum equivalent of inner, outer, sum, and mul function (★★★)
```
# Author: Alex Riley
# Make sure to read: http://ajcr.net/Basic-guide-to-einsum/
A = np.random.uniform(0,1,10)
B = np.random.uniform(0,1,10)
np.einsum('i->', A) # np.sum(A)
np.einsum('i,i->i', A, B) # A * B
np.einsum('i,i', A, B) # np.inner(A, B)
np.einsum('i,j->ij', A, B) # np.outer(A, B)
```
#### (98) Considering a path described by two vectors (X,Y), how to sample it using equidistant samples (★★★)?
```
# Author: Bas Swinckels
phi = np.arange(0, 10*np.pi, 0.1)
a = 1
x = a*phi*np.cos(phi)
y = a*phi*np.sin(phi)
dr = (np.diff(x)**2 + np.diff(y)**2)**.5 # segment lengths
r = np.zeros_like(x)
r[1:] = np.cumsum(dr) # integrate path
r_int = np.linspace(0, r.max(), 200) # regular spaced path
x_int = np.interp(r_int, r, x) # integrate path
y_int = np.interp(r_int, r, y)
```
#### (99) Given an integer n and a 2D array X, select from X the rows which can be interpreted as draws from a multinomial distribution with n degrees, i.e., the rows which only contain integers and which sum to n. (★★★)
```
# Author: Evgeni Burovski
X = np.asarray([[1.0, 0.0, 3.0, 8.0],
[2.0, 0.0, 1.0, 1.0],
[1.5, 2.5, 1.0, 0.0]])
n = 4
M = np.logical_and.reduce(np.mod(X, 1) == 0, axis=-1)
M &= (X.sum(axis=-1) == n)
print(X[M])
```
#### (100) Compute bootstrapped 95% confidence intervals for the mean of a 1D array X (i.e., resample the elements of an array with replacement N times, compute the mean of each sample, and then compute percentiles over the means). (★★★)
```
# Author: Jessica B. Hamrick
X = np.random.randn(100) # random 1D array
N = 1000 # number of bootstrap samples
idx = np.random.randint(0, X.size, (N, X.size))
means = X[idx].mean(axis=1)
confint = np.percentile(means, [2.5, 97.5])
print(confint)
```
https://github.com/rougier/numpy-100
| github_jupyter |
```
import numpy as np
import pandas as pd
from pylab import plt,mpl
plt.style.use('seaborn')
mpl.rcParams['font.family']='serif'
%matplotlib inline
data=pd.read_csv('tr_eikon_eod_data.csv',index_col=0,parse_dates=True)
data.head(10)
data=pd.DataFrame(data['.SPX'])
data.dropna(inplace=True)
data['rtes']=np.log(data/data.shift(1))
data['vol']=data['rtes'].rolling(252).std()*np.sqrt(252)
data[['.SPX', 'vol']].plot(subplots=True, figsize=(10, 6))
import math
loops=2500000
a=range(1,loops)
def f(x):
return 3*math.log(x)+math.cos(x)**2
%timeit r=[f(x) for x in a]
import numpy as np
a=np.arange(1,loops)
%timeit r=3*np.log(a)+np.cos(a)**2
import numexpr as ne
ne.set_num_threads(1)
f='3*log(a)+cos(a)**2'
%timeit r=ne.evaluate(f)
ne.set_num_threads(8)
%timeit r=ne.evaluate(f)
from tsfresh.examples.robot_execution_failures import download_robot_execution_failures, \
load_robot_execution_failures
download_robot_execution_failures()
timeseries, y = load_robot_execution_failures()
print(timeseries.head())
import matplotlib.pyplot as plt
%matplotlib inline
timeseries[timeseries['id'] == 3].plot(subplots=True, sharex=True, figsize=(10,10))
from tsfresh import extract_features
extracted_features = extract_features(timeseries, column_id="id", column_sort="time")
from tsfresh import select_features
from tsfresh.utilities.dataframe_functions import impute
impute(extracted_features)
features_filtered = select_features(extracted_features, y)
from tsfresh import extract_relevant_features
features_filtered_direct = extract_relevant_features(timeseries, y,
column_id='id', column_sort='time')
features_filtered_direct.dropna(inplace=True)
y.value_counts()
param = {'num_leaves': 31, 'objective': 'binary'}
param['metric'] = 'binary_logloss'
import numpy as np
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test=train_test_split(features_filtered_direct,y, test_size=0.33, random_state=42)
lgb_train=lgb.Dataset(X_train,y_train)
lgb_eval=lgb.Dataset(X_test,y_test,reference=lgb_train)
import lightgbm as lgb
gbm=lgb.train(param,train_set=lgb_train,num_boost_round=600,
valid_sets=lgb_eval,early_stopping_rounds=5)
ygbm=gbm.predict(features_filtered_direct)
ygbm=np.array(ygbm)
results=np.where(ygbm>=0.5,1,0)
from sklearn.metrics import accuracy_score
accuracy=accuracy_score(results,y)
accuracy
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(y, results)
cm
from sn_random_numbers import *
import sn
```
| github_jupyter |
```
from matplotlib import pyplot as plt
from sklearn.preprocessing import MinMaxScaler
import tensorflow as tf
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import GRU, Dense
from tensorflow.keras.layers import LSTM
from tensorflow.keras import callbacks
from tensorflow.keras import optimizers
import pandas as pd
import numpy as np
df = pd.read_csv('data/international-airline-passengers.csv', index_col='Month')
print(df.head())
df.plot()
```
# Data cleansing
```
columns_to_keep = ['Passengers']
df = df[columns_to_keep]
df['Passengers'] = df['Passengers'].apply(lambda x: x*1000)
df.index.names = ['Month']
df.sort_index(inplace=True)
print('Total rows: {}'.format(len(df)))
df.head()
df.describe()
df.plot()
```
Null values?
```
df.isnull().sum()
null_columns=df.columns[df.isnull().any()]
df[null_columns].isnull().sum()
print(df[df.isnull().any(axis=1)][null_columns].head())
df.dropna(inplace=True)
df.isnull().sum()
df.hist(bins=10)
len(df[df['Passengers'] == 0])
```
# Scaled data
```
print('Min', np.min(df))
print('Max', np.max(df))
```
We can then extract the NumPy array from the dataframe and convert the integer values to floating point values, which are more suitable for modeling with a neural network.
```
dataset = df.astype('float32')
```
LSTMs are sensitive to the scale of the input data, specifically when the sigmoid (default) or tanh activation functions are used. It can be a good practice to rescale the data to the range of 0-to-1, also called normalizing. We can easily normalize the dataset using the MinMaxScaler preprocessing class from the scikit-learn library.
```
scaler = MinMaxScaler(feature_range=(0, 1))
scaled = scaler.fit_transform(dataset)
print('Min', np.min(scaled))
print('Max', np.max(scaled))
print(scaled[:10])
```
# Create the RNN
A simple method that we can use is to split the ordered dataset into train and test datasets. The code below calculates the index of the split point and separates the data into the training datasets with 70% of the observations that we can use to train our model, leaving the remaining 30% for testing the model.
Split into train and test sets
```
train_size = int(len(scaled) * 0.70)
test_size = len(scaled - train_size)
train, test = scaled[0:train_size, :], scaled[train_size: len(scaled), :]
print('train: {}\ntest: {}'.format(len(train), len(test)))
```
convert an array of values into a dataset matrix
The function takes two arguments: the dataset, which is a NumPy array that we want to convert into a dataset, and the look_back, which is the number of previous time steps to use as input variables to predict the next time period — in this case defaulted to 1.
This default will create a dataset where X is the energy quantity at a given time (t) and Y is the qty of energy at the next time (t + 1).
```
def create_dataset(dataset, look_back=1):
print(len(dataset), look_back)
dataX, dataY = [], []
for i in range(len(dataset)-look_back-1):
a = dataset[i:(i+look_back), 0]
print(i)
print('X {} to {}'.format(i, i+look_back))
print(a)
print('Y {}'.format(i + look_back))
print(dataset[i + look_back, 0])
dataset[i + look_back, 0]
dataX.append(a)
dataY.append(dataset[i + look_back, 0])
return np.array(dataX), np.array(dataY)
```
reshape into X=t and Y=t+1
```
look_back = 1
X_train, y_train = create_dataset(train, look_back)
X_test, y_test = create_dataset(test, look_back)
```
The LSTM network expects the input data (X) to be provided with a specific array structure in the form of: [samples, time steps, features].
reshape input to be [samples, time steps, features]
```
X_train = np.reshape(X_train, (X_train.shape[0], X_train.shape[1], 1))
X_test = np.reshape(X_test, (X_test.shape[0], X_test.shape[1], 1))
print(X_train.shape)
print(X_test.shape)
```
The network has a visible layer with 1 input, a hidden layer with 4 LSTM blocks or neurons, and an output layer that makes a single value prediction. The default sigmoid activation function is used for the LSTM blocks. The network is trained for 100 epochs and a batch size of 1 is used.
create and fit the LSTM network
```
batch_size = 1
model = Sequential()
model.add(LSTM(4, batch_input_shape=(batch_size, look_back, 1), stateful=True))
model.add(Dense(1))
model.compile(loss='mean_squared_error', optimizer='adam')
model.fit(X_train, y_train, epochs=100, batch_size=batch_size, verbose=2, shuffle=True)
```
Note that we invert the predictions before calculating error scores to ensure that performance is reported in the same units as the original data (thousands of passengers per month).
Make preditions
```
import math
from sklearn.metrics import mean_squared_error
trainPredict = model.predict(X_train, batch_size=batch_size)
model.reset_states()
testPredict = model.predict(X_test, batch_size=batch_size)
# invert predictions
trainPredict = scaler.inverse_transform(trainPredict)
y_train = scaler.inverse_transform([y_train])
testPredict = scaler.inverse_transform(testPredict)
y_test = scaler.inverse_transform([y_test])
# calculate root mean squared error
trainScore = math.sqrt(mean_squared_error(y_train[0], trainPredict[:,0]))
print('Train Score: %.2f RMSE' % (trainScore))
testScore = math.sqrt(mean_squared_error(y_test[0], testPredict[:,0]))
print('Test Score: %.2f RMSE' % (testScore))
```
Because of how the dataset was prepared, we must shift the predictions so that they align on the x-axis with the original dataset. Once prepared, the data is plotted, showing the original dataset in blue, the predictions for the training dataset in green, and the predictions on the unseen test dataset in red.
shift train predictions for plotting
```
trainPredictPlot = np.empty_like(scaled)
trainPredictPlot[:, :] = np.nan
trainPredictPlot[look_back:len(trainPredict)+look_back, :] = trainPredict
# shift test predictions for plotting
testPredictPlot = np.empty_like(scaled)
testPredictPlot[:, :] = np.nan
testPredictPlot[len(trainPredict)+(look_back*2)+1:len(scaled)-1, :] = testPredict
# plot baseline and predictions
plt.figure(figsize=(20,10))
plt.plot(scaler.inverse_transform(scaled))
plt.plot(trainPredictPlot)
plt.plot(testPredictPlot)
plt.show()
```
| github_jupyter |
# make regional weights files
```
import rhg_compute_tools.kubernetes as rhgk
import dask.distributed as dd
import dask.dataframe as ddf
import geopandas as gpd
import pandas as pd
import xarray as xr
import numpy as np
import cartopy.crs as ccrs
import cartopy.feature as cfeature
%matplotlib inline
import os
CRS_SUPPORT_BUCKET = os.environ['CRS_SUPPORT_BUCKET']
admin1 = gpd.read_file('https://naciscdn.org/naturalearth/10m/cultural/ne_10m_admin_1_states_provinces.zip')
admin0 = gpd.read_file('https://naciscdn.org/naturalearth/10m/cultural/ne_10m_admin_0_countries.zip')
admin1.plot()
admin0.plot()
from distutils.version import LooseVersion
def get_containing_region(df, shapes, feature, x_col='x', y_col='y', crs='epsg:4326', predicate='intersects'):
"""
Return a feature corresponding to the first polygon to contain each (x, y) point in a dataframe
Parameters
----------
df : pd.DataFrame
Dataframe with x/y data to assign to geometries
shapes : gpd.GeoDataFrame
geopandas dataframe with polygon geometries
feature : str
column name in ``shapes`` to return for each matched feature
x_col : str, optional
column name in ``df`` to use for x coordinate of points (default 'x')
y_col : str, optional
column name in ``df`` to use for y coordinate of points (default 'y')
crs : str, optional
Coordinate reference system code for the projection of the (x, y) points in
``df``. Must be interpretable by ``pyproj``. Default ``'epsg:4326'`` (WGS 84).
predicate : str, optional
name of the binary predicate to use on the join. Must be a shapely binary
predicate. See the geopandas docs on
`sjoin <https://geopandas.org/en/stable/docs/reference/api/geopandas.sjoin.html>`_.
Default (``'intersects'``) means the first polygon in ``shapes`` to intersect
the point (either contain or touch with boundary) will be returned.
"""
points = gpd.GeoDataFrame(
geometry=gpd.points_from_xy(df[x_col], df[y_col], crs=crs),
index=df.index,
)
if LooseVersion(gpd.__version__) >= LooseVersion('0.10.0'):
pred = {'predicate': predicate}
else:
pred = {'op': predicate}
return gpd.sjoin(points, shapes[[feature, 'geometry']], how='left', **pred)[feature]
client, cluster = rhgk.get_cluster()
cluster.scale(20)
cluster
gpwv4_unwpp_2020 = ddf.read_parquet(
f'gs://{CRS_SUPPORT_BUCKET}/public_datasets/spatial/exposure/GLOBAL/population/'
'gpw-v4-population-count-adjusted-to-2015-unwpp-country-totals-rev11_2020_30_sec_tif/'
'derived_datasets/reformatted/'
'gpw-v4-population-count-adjusted-to-2015-unwpp-country-totals-rev11_2020_30_sec.parquet',
chunksize='100MB',
)
POP_RASTER_WITH_REGION_ASSIGNMENTS = (
'gs://downscaled-288ec5ac/diagnostics/rasters/'
'gpw-v4-population-count-adjusted-to-2015-unwpp-country-totals-rev11_2020_30_sec/'
'naturalearth_v5.0.0_adm0_and_adm1_assignments.parquet'
)
gpwv4_unwpp_2020
assignments_adm0 = gpwv4_unwpp_2020.map_partitions(
get_containing_region,
shapes=admin0[['ADM0_A3', 'geometry']],
feature='ADM0_A3',
meta='str',
)
assignments_adm0 = assignments_adm0.persist()
dd.progress(assignments_adm0, notebook=False)
dd.wait(assignments_adm0)
assert not any([f.status == 'error' for f in assignments_adm0.dask.values()]), assignments_adm0.compute()
gpwv4_unwpp_2020['ADM0_A3'] = assignments_adm0
assignments_adm1 = gpwv4_unwpp_2020.map_partitions(
get_containing_region,
shapes=admin1[['adm1_code', 'geometry']],
feature='adm1_code',
meta='str',
)
assignments_adm1 = assignments_adm1.persist()
dd.progress(assignments_adm1, notebook=False)
dd.wait(assignments_adm1)
assert not any([f.status == 'error' for f in assignments_adm1.dask.values()]), assignments_adm1.compute()
gpwv4_unwpp_2020['adm1_code'] = assignments_adm1
gpwv4_unwpp_2020 = gpwv4_unwpp_2020.dropna(how='all').compute()
gpwv4_unwpp_2020.repartition(npartitions=20).to_parquet(POP_RASTER_WITH_REGION_ASSIGNMENTS)
client.restart()
cluster.scale(0)
client.close()
cluster.close()
```
| github_jupyter |
Create a reduced basis for a simple sinusoid model
```
%matplotlib inline
import numpy as np
from misc import *
import matplotlib.pyplot as plt
```
Create the signal model:
$h(t) = A\sin{(2\pi(f_0 t + 0.5\dot{f} t^2))}$
```
def signalmodel(A, f0, f1, t):
"""
A time domain sinusoid with an amplitude A, frequency f0,
spin-down f1, evaluated at a set of time stamps t.
"""
return A*np.sin(2*np.pi*(f0*t + 0.5*f1*t**2))
```
Initialise the model time series
```
# a time series
t0 = 0
tend = 86400.*10
N = (tend-t0)/60.
ts = np.linspace(t0, tend, N)
dt = ts[1]-ts[0]
```
Create a training set of 4000 waveforms with random frequency and frequency derivatives within a narrow range.
```
# number of training waveforms
TS_size = 4000
#f0s = np.random.rand(TS_size)*0.0002-0.0001
# set f0s using Chebyshev-Gauss-Lobatto nodes
f0s, df0s = chebyshev_gauss_lobatto_nodes_and_weights(-0.0001, 0.0001, TS_size)
f1s = np.random.rand(TS_size)*(2e-10)-1e-10
# allocate memory and create training set
TS = np.zeros(TS_size*len(ts)).reshape(TS_size, len(ts)) # store training space in TS_size X len(ts) array
A = 1.
for i in range(TS_size):
TS[i] = signalmodel(A, f0s[i], f1s[i], ts)
# normalize
TS[i] /= np.sqrt(abs(dot_product(dt, TS[i], TS[i])))
```
Allocate memory for reduced basis vectors.
```
# Allocate storage for projection coefficients of training space waveforms onto the reduced basis elements
proj_coefficients = np.zeros(TS_size*TS_size).reshape(TS_size, TS_size)
# Allocate matrix to store the projection of training space waveforms onto the reduced basis
projections = np.zeros(TS_size*len(ts)).reshape(TS_size, len(ts))
rb_errors = []
#### Begin greedy: see Field et al. arXiv:1308.3565v2 ####
tolerance = 10e-12 # set maximum RB projection error
sigma = 1 # (2) of Algorithm 1. (projection error at 0th iteration)
rb_errors.append(sigma)
```
Run greedy algorithm for creating the reduced basis
```
RB_matrix = [TS[0]] # (3) of Algorithm 1. (seed greedy algorithm (arbitrary))
iter = 0
while sigma >= tolerance: # (5) of Algorithm 1.
# project the whole training set onto the reduced basis set
projections = project_onto_basis(dt, RB_matrix, TS, projections, proj_coefficients, iter)
residual = TS - projections
# Find projection errors
projection_errors = [dot_product(dt, residual[i], residual[i]) for i in range(len(residual))]
sigma = abs(max(projection_errors)) # (7) of Algorithm 1. (Find largest projection error)
# break out if sigma is less than tolerance, so another basis is not added to the set
# (this can be required is the waveform only requires a couple of basis vectors, and it
# stops a further basis containing large amounts of numerical noise being added)
if sigma < tolerance:
break
print sigma, iter
index = np.argmax(projection_errors) # Find Training-space index of waveform with largest proj. error
rb_errors.append(sigma)
#Gram-Schmidt to get the next basis and normalize
next_basis = TS[index] - projections[index] # (9) of Algorithm 1. (Gram-Schmidt)
next_basis /= np.sqrt(abs(dot_product(dt, next_basis, next_basis))) #(10) of Alg 1. (normalize)
RB_matrix.append(next_basis) # (11) of Algorithm 1. (append reduced basis set)
iter += 1
```
Check that this basis does give the expected residuals for a new set of random waveforms generated from the same parameter range.
```
#### Error check ####
TS_rand_size = 2000
TS_rand = np.zeros(TS_rand_size*len(ts)).reshape(TS_rand_size, len(ts)) # Allocate random training space
f0s_rand = np.random.rand(TS_rand_size)*0.0002-0.0001
f1s_rand = np.random.rand(TS_rand_size)*(2e-10)-1e-10
for i in range(TS_rand_size):
TS_rand[i] = signalmodel(A, f0s_rand[i], f1s_rand[i], ts)
# normalize
TS_rand[i] /= np.sqrt(abs(dot_product(dt, TS_rand[i], TS_rand[i])))
### find projection errors ###
iter = 0
proj_rand = np.zeros(len(ts))
proj_error = []
for h in TS_rand:
while iter < len(RB_matrix):
proj_coefficients_rand = dot_product(dt, RB_matrix[iter], h)
proj_rand += proj_coefficients_rand*RB_matrix[iter]
iter += 1
residual = h - proj_rand
projection_errors = abs(dot_product(dt, residual, residual))
proj_error.append(projection_errors)
proj_rand = np.zeros(len(ts))
iter = 0
plt.scatter(np.linspace(0, len(proj_error), len(proj_error)), np.log10(proj_error))
plt.ylabel('log10 projection error')
plt.show()
```
Now let's try and find an empirical interpolant and set of time stamps.
```
# put basis into complex form
e = np.array(RB_matrix)
indices = []
ts_nodes = []
V = np.zeros((len(e), len(e)))
```
Find emipirical interpolant
```
from scipy.linalg import inv
# seed EIM algorithm
indices.append( int(np.argmax( np.abs(e[0]) )) ) # (2) of Algorithm 2
ts_nodes.append(ts[indices[0]]) # (3) of Algorithm 2
for i in range(1, len(e)): #(4) of Algorithm 2
#build empirical interpolant for e_iter
for j in range(len(indices)): # Part of (5) of Algorithm 2: making V_{ij}
for k in range(len(indices)): # Part of (5) of Algorithm 2: making V_{ij}
V[k][j] = e[j][indices[k]] # Part of (5) of Algorithm 2: making V_{ij}
invV = inv(V[0:len(indices), 0:len(indices)]) # Part of (5) of Algorithm 2: making V_{ij}
B = B_matrix(invV, e) # Part of (5) of Algorithm 2: making B_j(f)
interpolant = emp_interp(B, e[i], indices) # Part of (5) of Algorithm 2: making the empirical interpolant of e
res = interpolant - e[i] # 6 of Algorithm 2
index = int(np.argmax(np.abs(res))) # 7 of Algorithm 2
print "ts{%i} = %f"%(i, ts[index])
indices.append(index) # 8 of Algorithm 2
ts_nodes.append( ts[index] ) # 9 of Algorithm 2
# make B matrix with all the indices
for j in range(len(indices)):
for k in range(len(indices)):
V[k][j] = e[j][indices[k]]
invV = inv(V[0:len(indices), 0:len(indices)])
B = B_matrix(invV, e)
```
Let's compare a random waveform and it's equivalent interpolant for comparison.
```
h_for_comparison = signalmodel(A, f0s_rand[10], f1s_rand[10], ts)
interpolant_for_comparison = np.inner(B.T, h_for_comparison[indices])
plt.plot(ts, h_for_comparison-interpolant_for_comparison, 'b')
plt.xlabel('time (s)')
plt.show()
```
Now let's create a set of random waveforms from within the parameter ranges and further compare errors between the full waveform and interpolant.
```
H_size = 2000
H = np.zeros(H_size*len(ts)).reshape(H_size, len(ts)) # Allocate random training space
f0s_rand = np.random.rand(H_size)*0.0002-0.0001
f1s_rand = np.random.rand(H_size)*(2e-10)-1e-10
# create set of test waveforms
for i in range(H_size):
H[i] = signalmodel(A, f0s_rand[i], f1s_rand[i], ts)
# find errors between full waveform and interpolants
list_of_errors = []
for i in range(H_size):
interpolant = np.inner(B.T, H[i][indices])
interpolant /= np.sqrt(np.vdot(interpolant, interpolant)) #normalize
H[i] /= np.sqrt(np.vdot(H[i], H[i]) ) #normalize
error = abs(np.vdot(H[i] - interpolant, H[i] - interpolant ))
list_of_errors.append(error)
print error
plt.scatter(np.linspace(0, H_size, H_size), np.log10(list_of_errors))
plt.ylabel('log10 interpolation error')
plt.show()
```
Create weights for data-model terms and model-model terms
```
data = np.random.randn(len(ts))
# create weights for data-model terms of interpolant
w = np.inner(B, data.T)
# create weights model-model terms of interpolant
w2 = np.zeros((B.shape[0], B.shape[0]))
for i in range(B.shape[0]):
for j in range(B.shape[0]):
w2[i,j] = np.sum(B[i]*B[j])
```
Now do some accuracy tests
```
sigfull = signalmodel(A, f0s_rand[23], f1s_rand[23], ts)
tsred = np.array(ts_nodes)
sigred = signalmodel(A, f0s_rand[23], f1s_rand[23], tsred)
d_dot_h = np.dot(data, sigfull)
ROQdh = np.dot(w, sigred)
print "regular inner product = %.15e"%d_dot_h
print "ROQ = %.15e"%ROQdh
h_dot_h = np.dot(sigfull, sigfull)
ROQh = np.dot(np.dot(sigred, w2), sigred)
print "regular inner product = %.15e"%h_dot_h
print "ROQ = %.15e"%ROQh
```
Now do a speed test
```
import time
t1 = time.time()
for i in range(50000):
sigfullnew = signalmodel(A, f0s_rand[0], f1s_rand[0], ts)
dms = data-sigfullnew
np.dot(dms, dms) # regular inner product
e1 = time.time()
t2 = time.time()
for i in range(50000):
sigrednew = signalmodel(A, f0s_rand[0], f1s_rand[0], tsred)
np.dot(w, sigred)
np.dot(np.dot(sigrednew, w2), sigrednew) # ROQ inner product
e2 = time.time()
print "regular inner product took %f s"%((e1-t1)/50000.)
print "ROQ took %f s"%((e2-t2)/50000.)
print "speedup = %f"%((e1-t1) / (e2-t2))
print float(len(ts))/float(len(tsred))
```
| github_jupyter |
# MADDPG for Multi-Agent Collaboration and Competition Task
In this project we have trained a multi agent policy gradient reinforcement learning agent to solve a [Multi-Agent Actor-Critic for Mixed
Cooperative-Competitive Environments, MADDPG](https://papers.nips.cc/paper/2017/file/68a9750337a418a86fe06c1991a1d64c-Paper.pdf).
In details, the environment built in 3D space with two tennis rocket learning to keep to ball in the game as long as possible.
There are 2 action available for controlling the each rocket in a continuous space for jumping and move back and forth.
The measured stated contains position, velocity of the ball and two tennis rocket.
The task designed episodically and each episode last maximum of until the ball fall down.
Our solution uses a multi agent policy gradient based reinforcement learning algorithm called
[Multi-Agent Actor-Critic for Mixed Cooperative-Competitive Environments, MADDPG](https://papers.nips.cc/paper/2017/file/68a9750337a418a86fe06c1991a1d64c-Paper.pdf).
The optimal policy should maximize its expected discounted reward $\mu^*(s)=\max_\pi \mathbb{E}[\sum_i \gamma^i r_i]$ by keeping the ball from falling down.
In addition to the vanilla MADDPG algorithm I applied a prioritized reply buffer to solve the environment.
In the following I will describe each step of our method in details and present the results.
In this report the reference to the ideas and publications provide via hyper-link.
#### Solved Environment for collaborative and competitive tennis game.

## Learning Algorithm
* **Reinforcement Learning**: is a type of machine learning method which tries to learn an appropriate closed-loop controller by simply interacting with the process and incrementally improving the control behavior. The goal of reinforcement learning algorithms is to maximize a numerical reward signal by discovering which control commands i.e. actions yield the most reward. Using reinforcement learning algorithms, a controller can be learned with only a small amount of prior knowledge of the process. Reinforcement learning aims at learning control policies for a system in situations where the training
information is basically provided in terms of judging success or failure of the observed system behavior.
* **Markov Decision Process**: The type of control problems we are trying to learn in this work are discrete time control problems and can be formulated as a Markov decision process(MDP). An MDP has four components: a set $S$ of states, a set $A$ of actions, a stochastic transition probability function $p(s, a, s' )$ describing system behavior, and an immediate reward or cost function $c : S × A → R$. The state of the system at time $t$, characterizes the current situation of the agent in the world, denoted by $s(t)$. The chosen action by agent at time step $t$ is denoted by $a(t)$. The immediate reward or cost is the consequence of the taken action and function of state and action. Since the rewards for the taken action can be formulated as cost, the goal of the control agent would be to find an optimal policy $π ∗ : S → A$ that minimizes the cumulated cost for all states. Basically, in reinforcement learning we
try to choose actions over time to minimize/maximize the expected value of the total cost/reward.
* **Q-Learning**: In many real-world problems the state transition probabilities and the reward functions are not given explicitly. But, only a set of states $S$ and a set of actions $A$ are known and we have to learn the dynamic system behaviors by interacting with it. Methods of temporal differences such as Q-Learning were invented to perform learning and optimization in exactly these circumstances. The basic idea in Q-learning is to iteratively learn the value function, Q-function, that maps state-action pairs to expected optimal path costs. The goal of a Q-learning is to find optimal policy which returns highest expected reward give the action-value function. In order to find the optimal action in each state we use Bellman optimality principal. By definition that means at each time step the optimal value of action in a particular state is equal sum its action-value and discounted reward collected after that time stamp onward. The update equation for the action-value function is : $Q(S_t , A_t )\leftarrow Q(S_t , A_t ) + \alpha[R_{t+1} + \gamma \max_a' Q(S_{t+1} , a') - Q(S_t , A_t)]$.
* **On-policy** methods attempt to evaluate or improve the policy that is used to make decisions, whereas **off-policy** methods evaluate or improve a policy different from that used to generate the data. In **on-policy** control methods the policy is generally soft, meaning that $\pi(a|s) > 0$ for all $s \in S$ and all $a \in A(s)$, but gradually shifted closer and closer to a deterministic optimal policy. The on-policy approach in the preceding section is actually a compromise—it learns action values not for the optimal policy, but for a near-optimal policy that still explores.
* **Batch Reinforcement Learning**: At each time point $t$ it observes the environment state $s_t$ , takes an action $a_t$ , and receives feedbacks from the environment including next state $s_{t+1}$ and the instantaneous reward $r_t$. The sole information that we assume available to learn the problem is the one obtained from the observation of a certain number of one-step system transitions (from $t$ to $t + 1$). The agent interacts with the control system in the environment and gathers state transitions in a set of four-tuples $(s_t , a_t , r_t , s_{t+1 })$. Except for very special conditions, it is not possible to exactly determine an optimal control policy from a finites et of transition samples. Batch reinforcement learning aims at computing an approximation of such optimal policy $\pi^∗$, from a set of four-tuples: $D=[(s^l_{t}, a^l_{t} , r^l_{t} , s^l_{t+1}), l = 1,..., \#D]$. This set could be generated by gathering samples corresponding to one single trajectory (or episode) as well as by considering several independently generated trajectories or multi-step episodes. Training algorithms with growing batch have two major benefits. First, from the interaction perspective, it is very similar to the ’pure’ online approach. Second, from the learning point of view, it is similar to an
off-line approach that all the trajectory samples are used for training the algorithm. The main idea in growing batch is to alternate between phases of exploration, where a set of training examples is grown by interacting with the system, and phases of learning, where the whole batch of observations is used. The distribution of the state transitions in the provided batch must resemble the ’true’ transition probabilities of the system in order to allow the derivation of good policies. In practice, exploration cultivates the quality of learned policies by providing more variety in the distribution of the trajectory samples.
* **Policy Gradient Methods** learn a parameterized policy that can select actions without consulting a value function. A value function may still be used to learn the policy parameter, but is not required for action selection. We use the notation $\theta \in \mathbb R^{d'}$ for the policy’s parameter vector. Thus we write $\pi(a|s, \theta) = Pr\{A_t = a | S_t = s, \theta_t =\theta\}$ for the probability that action a is taken at time $t$ given that the environment is in state s at time $t$ with parameter $\theta$. If a method uses a learned value function as well, then the value function’s weight vector is denoted $w \in \mathbb R^d$ as usual, as in $\hat{v}(s,w)$. We use the gradient of some scaler value $J(\theta)$ with respect to policy parameters to learn the policy. The method aim to maximize the performance therefore the parameter updates uses the gradient ascent: $ \theta_{t+1}=\theta_t + \alpha\widehat{\nabla J(\theta_t)}$, where $\widehat{\nabla J(\theta_t)}\in \mathbb R^d$ is a stochastic estimate whose expectation approximates the gradient of the performance measure with respect to its argument $\theta_t$ . All methods that follow this general schema we call **policy gradient methods**, whether or not they also learn an approximate value function. Methods that learn approximations to both policy and value functions are often called **actor–critic methods**, where **‘actor’** is a reference to the learned policy, and **‘critic’** refers to the learned value function, usually a state-value function.
* One advantage of parameterizing policies according to the soft-max in action preferences is that the approximate policy can approach a deterministic policy, whereas with $\epsilon-greedy$ action selection over action values there is always an $\epsilon$ probability of selecting a random action.
* A second advantage of parameterizing policies according to the soft-max in action preferences is that it enables the selection of actions with arbitrary probabilities. In problems with significant function approximation, the best approximate policy may be stochastic.
* **Actor-Critic Method** we use state-value function to critic the actor function. Basically we use the state-value function to bootstrapping (updating the value estimate for a state from the estimated values of subsequent states). This is a useful distinction, for only through bootstrapping do we introduce bias and an asymptotic dependence on the quality of the function approximation. The bias introduced through bootstrapping and reliance on the state representation is often beneficial because it reduces variance and accelerates learning. Actor-critic methods are TD methods that have a separate memory structure to explicitly represent the policy independent of the value function. The policy structure is known as the *actor*, because it is used to select actions, and the estimated value function is known as the *critic*, because it criticizes the actions made by the actor. Learning is always on-policy: the critic must learn about and critique whatever policy is currently being followed by the actor. The critique takes the form of a TD error. This scalar signal is the sole output of the critic and drives all learning in both actor and critic.

* **CONTINUOUS CONTROL WITH DEEP REINFORCEMENT LEARNING(DDGP)** while DQN solves problems with high-dimensional observation spaces, it can only handle discrete and low-dimensional action spaces. Many tasks of interest, most notably physical control tasks, have continuous (real valued) and high dimensional action spaces. DQN cannot be straightforwardly applied to continuous domains since it relies on a finding the action that maximizes the action-value function, which in the continuous valued case requires an iterative optimization process at every step. An obvious approach to adapting deep reinforcement learning methods such as DQN to continuous domains is to to simply discretize the action space. However, this has many limitations, most notably the curse of dimensionality: the number of actions increases exponentially with the number of degrees of freedom. Such large action spaces are difficult to explore efficiently, and thus successfully training DQN-like networks in this context is likely intractable. Additionally, naive discretization of action spaces needlessly throws away information about the structure of the action domain, which may be essential for solving many problems. Here we combine the actor-critic approach with insights from the recent success of Deep Q Network (DQN). Prior to DQN, it was generally believed that learning value functions using large, non-linear function approximators was difficult and unstable. DQN is able to learn value functions using such function approximators in a stable and robust way due to two innovations: 1. the network is trained off-policy with samples from a replay buffer to minimize correlations between samples; 2. the network is trained with a target Q network to give consistent targets during temporal difference backups. It is not possible to straightforwardly apply Q-learning to continuous action spaces, because in continuous spaces finding the greedy policy requires an optimization of at at every timestep; this optimization is too slow to be practical with large, unconstrained function approximators and nontrivial action space. One challenge when using neural networks for reinforcement learning is that most optimization algorithms assume that the samples are independently and identically distributed. Obviously, when the samples are generated from exploring sequentially in an environment this assumption no longer holds. Additionally, to make efficient use of hardware optimizations, it is essential to learn in minibatches, rather than online. As in DQN, we used a replay buffer to address these issues. At each timestep the actor and critic are updated by sampling a minibatch uniformly from the buffer. Because DDPG is an off-policy algorithm, the replay buffer can be large, allowing the algorithm to benefit from learning across a set of uncorrelated transitions.

[source](https://arxiv.org/pdf/1509.02971.pdf)
* **Improvements To DDPG**:
* **batch normalization**: it is a method of adaptive re-parametrization, motivated by the difficulty of training very deep models. One of the key motivations for the development of BatchNorm was the reduction of so-called *internal covariate shift* (ICS). It is a mechanism that aims to stabilize the distribution (over a mini-batch) of inputs to a given network layer during training. It provides an elegant way of re-parameterizing almost any deep network. The re-parametrization significantly reduces the problem of coordinating updates across many layers. To do that, we apply normalization using $\mu$ and $\sigma$ on activation of each hidden layer $H$ before applying nonlinear function: $H' = \frac{H-\mu}{\sigma}$ where $\mu = \frac{1}{m}\sum_i H_i$, and $\sigma = \sqrt{\epsilon + \frac{1}{m}\sum_i(H_i-\mu)^2}$, The major innovation of the batch normalization is that, it prevents the gradient from increasing the standard deviation or mean of $h_i$; the normalization operations remove the effect of such an action and zero out its component in the gradient.
* **Gradient Clipping** One difficulty that arises with optimization of deep neural networks is that large parameter gradients can lead an SGD optimizer to update the parameters strongly into a region where the loss function is much greater, effectively undoing much of the work that was needed to get to the current solution. On the face of an extremely steep cliff structure, the gradient update step can move the parameters extremely far, usually jumping off of the cliff structure altogether **Gradient Clipping** clips the size of the gradients to ensure optimization performs more reasonably near sharp areas of the loss surface. It can be performed in a number of ways. The basic idea is to recall that the gradient does not specify the optimal step size, but only the optimal direction within an infinitesimal region. When the traditional gradient descent algorithm proposes to make a very large step, the gradient clipping heuristic intervenes to reduce the step size to be small enough that it is less likely to go outside the region where the gradient indicates the direction of approximately steepest descent. One option is to simply clip the parameter gradient element-wise before a parameter update. Another option is to clip the norm $||g||$ of the gradient g before a parameter update: $ {if}~||g||>v~then~g←\frac{g^v}{||g||}$ where $v$ is a norm threshold.
* **Less aggressive policy update** in order to minimize the error and stabilize the policy the update the actor-critic network with less frequency, 10 times after each 20 step moves in environment.
* **Markov Games** In this work, we consider a multi-agent extension of Markov decision processes (MDPs) called partially observable Markov games. Given $Q(s,a)$ an agent can maximize its reward using the “greedy” strategy of always choosing the action with the highest $Q-value$. This strategy is greedy because it treats $Q(s,a)$ as a surrogate for immediate reward and then acts to maximize its immediate gain. It is optimal because the $Q-function$ is an accurate summary of future rewards. A similar observation can be used for Markov games once we redefine $V(s)$; to be the expected reward for the optimal policy starting from state $s$ and $Q(s,a,o)$as the expected reward for taking action $a$ when the opponent chooses $o$ from state $s$ and continuing optimally thereafter. We can then treat the $Q(s,a,o)$ values as immediate payoffs in an unrelated sequence of matrix games (one for each state, $s$). The value of a state $s\in S$ in Markov game is $V(s)= \max_{\pi\in PD(A)}\min_{o\in O}\sum_{a\in A}Q(s,a,o)\pi_a$ where the agent policy is its probability distribution over actions $\pi\in PD(A)$ and the quality of the action $a$ against action $o$ in the state $s$ is $Q(s,a,o)= R(s,a,o)+ \gamma\sum_{s'}T(s,a,o,s')V(s')$, where $T$ is state transition probability.
* **Multi-Agent Actor-Critic for Mixed Cooperative-Competitive Environments** is a multi-agent policy gradient algorithm where agents learn a centralized critic
based on the observations and actions of all agents. This could be accomplish by adopting the framework of centralized training with decentralized execution.
Such framework allows the policies to use extra information to ease training, so long as this information is not used at test time.
It is unnatural to do this with Q-learning, as the Q function generally cannot contain different information at training and test time.
Therefore the method proposes a simple extension of actor-critic policy gradient methods where the critic is augmented with extra information about the policies of other agents.
Native policy gradient algorithms perform poorly in simple multi-agent settings. The followings are the constraints which the **MADDPG** is implemented on:
* the learned policies can only use local information (i.e. their own observations) at execution time
* there isn't a differentiable model of the environment dynamics
* there isn't any particular structure on the communication method between agents (no differentiable communication channel)

[source](https://papers.nips.cc/paper/2017/file/68a9750337a418a86fe06c1991a1d64c-Paper.pdf)
### The Experiment Setup :
* Each player agent hits the ball over the net, it receives a reward of +0.1. If an agent lets a ball hit the ground or hits the ball out of bounds, it receives a reward of -0.01.
* Each episode last as long ad the ball in the play.
* The training stopped when agent has collected $0.5$ reward points averaged over past $100$ episodes.
* The hyper parameters are equal to the original MADDPG paper, the followings are changed(similar for all the variation of the algorithm we have tried):
* gamma : 0.99
* actor_learning_rate : 0.0002
* critic_learning_rate: 0.0002
* update_every : 1 (for calculating loss)
* $\tau$: 0.01 (soft update factor)
* replay memory size: 100000
* gradient clip norm: 1.0 (gradient clipping applied to all the algorithm variations)
* batch size: 128
* Critic model architecture describe sequentially:
* Fully Connected Layer: [Input size=24 (state-size), Output=128]
* Normalization: BatchNorm1(128)
* Activation: ReLU
* Fully Connected Layer: [Input size=130, Output=128], the action are concatenated with output of fist fully connected layer
* Activation: ReLU
* Fully Connected Layer: [Input size=128, Output=1], the final output is state-action Q-value
* Actor model architecture describe sequentially:
* Fully Connected Layer: [Input size=24 (state-size), Output=128]
* Normalization: BatchNorm1(128)
* Activation: ReLU
* Fully Connected Layer: [Input size=128 (state-size), Output=128]
* Normalization: BatchNorm1(128)
* Activation: ReLU
* Fully Connected Layer: [Input size=128 (state-size), Output=2 (action-size)]
* Activation: Tanh, the final output is vector of action which could be applied directly
* Last layer of both actor and critic model weight initialized to uniform distribution in range $(-0.003,0.003)$
* Hidden layer of both actor and critic model weight initialized to uniform distribution in range $(-1/sqrt(fanin),1/sqrt(fanin))$
* Before applying each action generated by actor model we added noise using [Ornstein-Uhlenbeck process](https://en.wikipedia.org/wiki/Ornstein%E2%80%93Uhlenbeck_process) to generate temporally correlated
exploration for exploration efficiency in physical control problems with inertia.
* At the beginning of each episode the action noise hare being reset and decayed by factor 0.999 during each episode.
* Action values are clipped between [-1,1]
* The MADDPG agent creates 2 DDPG agents which trained in collaboration and competition with each other
* The Replay Buffer collects samples from both players.
* During the training loop we sample one batch on experiences and train both DDPG agent on the same batch
* For Prioritized Replay Buffer we sample two batch of experience, separate for each DDPG agent, due to updating experience priorities independently
### In the following I have trained the model and visualized the results
```
#!pip -q install ./python
import numpy as np
import time
import sys
import os
from collections import deque
#cd ../maddpgcollaboration
import numpy as np
import time
import sys
from collections import deque
from agent.maddpg import MADDPG
from agent_examples import maddpg_agent_config_default, maddpg_agent_config_per
from utils import (
log_training_info,
save_checkpoint,
get_training_env,
load_agent_parameters,
)
def maddpg_runner(
env,
brain_name="Reacher",
n_episodes=100,
agent_config={},
target_episodes=100,
target_score=0.5,
num_agents=20,
print_log=False,
run_mode="train",
saved_checkpoint_path="model/checkpoint.pth",
):
"""
Deep Deterministic Policy Gradients (DDPG).
Parameters
----------
env :
brain_name :
n_episodes :
agent_config :
target_episodes :
target_score :
num_agents :
print_log :
run_mode:
saved_checkpoint_path:
Returns
-------
"""
saved_good = False
scores = [] # episodic scores
training_info_log = [] # training time and meta data logger
moving_avg = deque(maxlen=100) # last 100 scores
agent = MADDPG(**agent_config)
if run_mode == "test":
load_agent_parameters(agent, saved_checkpoint_path)
for a in agent.agents:
a.actor_local.eval()
a.critic_local.eval()
## Perform n_episodes of training
training_start_time = time.time()
BETA_START = 0.5
for i_episode in range(1, n_episodes + 1):
beta = min(1.0, BETA_START + i_episode * (1.0 - BETA_START) / 1e6)
states = env.reset(train_mode=run_mode == "train")[
brain_name
].vector_observations
scores_episode = np.zeros(num_agents) # rewards per episode for each agent
episode_start_time = time.time()
agent.reset()
while True:
# Perform a step: S;A;R;S'
actions = agent.act(states) # select the next action for each agent
env_info = env.step(actions)[
brain_name
] # send the actions to the environment
rewards = env_info.rewards # get the rewards
next_states = env_info.vector_observations # get the next states
dones = env_info.local_done # see if episode has finished
# Send the results to the Agent
if run_mode == "train":
agent.step(states, actions, rewards, next_states, dones, beta)
# Update the variables for the next iteration
states = next_states
scores_episode += rewards
# break if any agents are done
if np.any(dones):
break
if run_mode == "test":
continue
# Store the rewards and calculate the moving average
score = np.max(scores_episode)
scores.append(score)
moving_avg.append(score)
training_info_log.append(
log_training_info(
i_episode,
training_start_time,
episode_start_time,
scores,
moving_avg,
print_log=print_log,
)
)
## Check if the environment has been solved
# if np.mean(moving_avg) >= target_score and i_episode >= target_episodes:
if np.mean(moving_avg) >= target_score and not saved_good:
# save_checkpoint(
# i_episode,
# scores,
# moving_avg,
# training_info_log,
# agent,
# agent_config,
# training_start_time,
# target_episodes,
# )
#saved_good = True
#print("\n checkpoint at episode {} saved.".format(i_episode))
break
return scores, moving_avg, agent, training_info_log
env_file_path = "../../Tennis_Linux_NoVis/Tennis.x86_64"
env, num_agents, brain_name, state_size, action_size = get_training_env(
env_file_path, test_mode="train"
)
maddpg_agent_config_per["action_size"] = action_size
maddpg_agent_config_per["state_size"] = state_size
scores, moving_avg, agent, training_info_log = maddpg_runner(
env=env,
brain_name=brain_name,
n_episodes=1000,
agent_config=maddpg_agent_config_default,
num_agents=num_agents,
print_log=True,
run_mode="train",
saved_checkpoint_path="",
target_score=0.6
)
from matplotlib import pyplot as plt
import numpy as np
import torch
fig, ax = plt.subplots(1, 1, figsize=(10, 5))
x = range(0, len(scores), 100)
y_mean = np.asarray(
[np.mean(scores[i : i + 100]) for i in range(0, len(scores), 100)]
)
y_std = np.asarray(
[np.std(scores[i : i + 100]) for i in range(0, len(scores), 100)]
)
ax.plot(
x,
y_mean,
color="r",
)
ax.fill_between(x, y_mean - y_std, y_mean + y_std)
ax.set_title("scores's mean and std of trained model")
ax.set_ylabel("Mean & STD scores")
ax.set_xlabel("episode number")
fig, ax = plt.subplots(1, 1, figsize=(10, 5))
ax.plot(scores)
ax.set_title("scores of trained model")
```
### Discussion and Future work:
* We managed to learn from multiple agents running in parallel and collecting samples to improve the policy. Clipping policy results, action, in the certain range could help agent to reach and stay in the target area. Usually applying large actions cause overshot and missing the target.
* As we could expect the adding batch normalization layer to the sequential feed forward neural network improves the performance due to regularization effect. During the training phase each batch of samples has its own mean and standard deviation, which effect distribution of the activation values. Batch normalization set mean of all activation to zero.
* We could not get better performance by using prioritized experience replay buffer in comparison to regular experience buffer.
* The idea for PER-buffer in [this work](https://ieeexplore.ieee.org/stamp/stamp.jsp?tp=&arnumber=9169650) should investigated further and requires hyper parameter tuning.
* In the future in order to achieve high performance on each algorithm we could use [bayesian hyperparameter optimization](https://github.com/fmfn/BayesianOptimization) and search for some of the key parameters which increase the quality of the results.
* Training and [Alpha-Zero algorithm](https://arxiv.org/abs/1712.01815) to learn this game would the very interesting alternative
* To take one step further we could learn from raw pixels instead of the retrieved features.
* One remaining important challenge is to build model which is robust to random initialization and could could converge consistently in short number of episodes.
| github_jupyter |
<a href="https://colab.research.google.com/github/wesleybeckner/technology_fundamentals/blob/main/C2%20Statistics%20and%20Model%20Creation/Tech_Fun_C2_S1_Regression_and_Descriptive_Statistics.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Technology Fundamentals Course 2, Session 1: Regression and Descriptive Statistics
**Instructor**: Wesley Beckner
**Contact**: wesleybeckner@gmail.com
**Teaching Assitants**: Varsha Bang, Harsha Vardhan
**Contact**: vbang@uw.edu, harshav@uw.edu
<br>
## Schedule for this week
<p align="center">
<img src="https://raw.githubusercontent.com/wesleybeckner/technology_fundamentals/main/assets/week2.png" width=800></img>
</p>
---
<br>
In this session we will look at fitting data to a curve using **regression**. We will also look at using regression to make **predictions** for new data points by dividing our data into a training and a testing set. Finally we will examine how much error we make in our fit and then in our predictions by computing the mean squared error.
<br>
---
<a name='x.0'></a>
## 1.0 Preparing Environment and Importing Data
[back to top](#top)
<a name='x.0.1'></a>
### 1.0.1 Import Packages
[back to top](#top)
```
# Import pandas, pyplot, ipywidgets
import pandas as pd
from matplotlib import pyplot as plt
from ipywidgets import interact
# Import Scikit-Learn library for the regression models
import sklearn
from sklearn import linear_model
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error, r2_score
# for enrichment topics
import seaborn as sns
import numpy as np
```
### 1.0.2 Load Dataset
[back to top](#top)
For our discussion on regression and descriptive statistics today we will use a well known dataset of different wines and their quality ratings
```
df = pd.read_csv("https://raw.githubusercontent.com/wesleybeckner/"\
"ds_for_engineers/main/data/wine_quality/winequalityN.csv")
df.shape
df.describe()
```
## 1.1 What is regression?
It is the process of finding a relationship between **_dependent_** and **_independent_** variables to find trends in data. This abstract definition means that you have one variable (the dependent variable) which depends on one or more variables (the independent variables). One of the reasons for which we want to regress data is to understand whether there is a trend between two variables.
**Housing Prices Example**
We can imagine this scenario with housing prices. Envision a **_mixed_** dataset of **_continuous_** and **_discrete_** independent variables. Some features could be continuous, floating point values like location ranking and housing condition. Others could be descrete like the number of rooms or bathrooms. We could take these features and use them to predict a house value. This would be a **_regression_** model.
<p align=center>
<img src="https://raw.githubusercontent.com/wesleybeckner/technology_explorers/main/assets/machine_learning/ML3.png" width=1000px></img>
## 1.2 Linear regression fitting with scikit-learn
#### Exercise 1: rudimentary EDA
What does the data look like? Remember how to visualize data in a pandas dataframe (Sessions 3 and 4)
<ol>
<li> for every column calculate the
* skew: `df.skew()`
* kurtosis: `df.kurtosis()`
* pearsons correlation with the dependent variable: `df.corr()`
* number of missing entries `df.isnull()`
and organize this into a new dataframe
_note:_ pearsons is just one type of correlation, another available to us **_spearman_** which differs from pearsons in that it depends on ranked values rather than their direct quantities, you can read more [here](https://support.minitab.com/en-us/minitab-express/1/help-and-how-to/modeling-statistics/regression/supporting-topics/basics/a-comparison-of-the-pearson-and-spearman-correlation-methods/)
```
df.isnull().sum()
# Cell for Exercise 1
# part A
# using df.<method> define the following four variables with the results from
# skew(), kurtosis(), corr() (and selecting for quality), and isnull()
# for isnull() you'll notice the return is a dataframe of booleans. we would
# like to simply know the number of null values for each column. change the
# return of isnull() using the sum() method
# skew =
# kurt =
# pear =
# null =
# part B
# on line 13, put these results in a list using square brackets and call
# pandas.DataFrame on the list to make your new DataFrame! store it under the
# variable name dff
# part C
# take the transpose of this DataFrame using dff.T. reassign dff to this copy
# part D
# set the column names to 'skew', 'kurtosis', 'pearsons _quality', and
# 'null count' using dff.columns
# Now return dff to the output to view your hand work
# dff
```
I have gone ahead and repeated this exercise with the red vs white wine types:
```
red = df.loc[df['type'] == 'red']
wht = df.loc[df['type'] == 'white']
def get_summary(df):
skew = df.skew()
kurt = df.kurtosis()
pear = df.corr()['quality']
null = df.isnull().sum()
med = df.median()
men = df.mean()
dff = pd.DataFrame([skew, kurt, pear, null, med, men])
dff = dff.T
dff.columns = ['skew', 'kurtosis', 'pearsons _quality', 'null count', 'median',
'mean']
return dff
dffr = get_summary(red)
dffw = get_summary(wht)
desc = pd.concat([dffr, dffw], keys=['red', 'white'])
desc
def my_fig(metric=desc.columns):
fig, ax = plt.subplots(1, 1, figsize=(10,10))
pd.DataFrame(desc[metric]).unstack()[metric].T.plot(kind='barh', ax=ax)
interact(my_fig)
```
#### Question 1: Discussion Around EDA Plot
What do we think of this plot?
> `metric = mean`, the cholrides values <br>
`metric = kurtosis`, residual sugar <br>
`metric = pearsons _quality`, _magnitudes_ and _directions_ <br>
How to improve the plot, what other plots would we like to see?
```
df['chlorides'].describe()
fig, ax = plt.subplots(1,1,figsize=(10,10))
df['chlorides'].plot(kind='kde',ax=ax)
ax.set_xlim(0,.61)
df['chlorides'].sort_values(ascending=False)[:50]
```
### 1.2.2 Visualizing the data set - motivating regression analysis
We can create a scatter plot of fixed acidity vs density of red wine in the dataset using `df.plot()` and see that there appears to be a general trend between the two features:
```
fig, ax = plt.subplots(1, 1, figsize=(5,5))
df.loc[df['type'] == 'red'].plot(x='fixed acidity', y='density', ax=ax,
ls='', marker='.')
```
### 1.2.3 Estimating the regression coefficients
It looks like density increases with fixed acidity following a line, maybe something like
$$y(x)= m \cdot x + b \;\;\;\;\;\;\;\;\sf{eq. 1}$$
with $y=\sf density$, $x=\sf fixed acidity$, and $m$ the slope and $b$ the intercept.
To solve the problem, we need to find the values of $b$ and $m$ in equation 1 to best fit the data. This is called **linear regression**.
In linear regression our goal is to minimize the error between computed values of positions $y^{\sf calc}(x_i)\equiv y^{\sf calc}_i$ and known values $y^{\sf exact}(x_i)\equiv y^{\sf exact}_i$, i.e. find $b$ and $m$ which lead to lowest value of
$$\epsilon (m,b) =SS_{\sf res}=\sum_{i=1}^{N}\left(y^{\sf exact}_i - y^{\sf calc}_i\right)^2 = \sum_{i=1}^{N}\left(y^{\sf exact}_i - m\cdot x_i - b \right)^2\;\;\;\;\;\;\;\;\;\;\;\sf{eq. 2}$$
To find out more see e.g. https://en.wikipedia.org/wiki/Simple_linear_regression
#### Question 2: linear regression loss function
> Do we always want *m* and *b* to be large positive numbers so as to minimize eq. 2?
Luckily [scikit-learn](https://scikit-learn.org/stable/) contains many functions related to regression including [linear regression](https://scikit-learn.org/stable/modules/linear_model.html).
The function we will use is called <code> LinearRegression() </code>.
```
# Create linear regression object
model = linear_model.LinearRegression()
# Use model to fit to the data, the x values are densities and the y values are fixed acidity
# Note that we need to reshape the vectors to be of the shape x - (n_samples, n_features) and y (n_samples, n_targets)
x = red['density'].values.reshape(-1, 1)
y = red['fixed acidity'].values.reshape(-1, 1)
```
```
# Create linear regression object
model = linear_model.LinearRegression()
# Use model to fit to the data, the x values are densities and the y values are fixed acidity
# Note that we need to reshape the vectors to be of the shape x - (n_samples, n_features) and y (n_samples, n_targets)
x = red['density'].values.reshape(-1, 1)
y = red['fixed acidity'].values.reshape(-1, 1)
```
```
print(red['density'].values.shape, red['fixed acidity'].values.shape)
print(x.shape, y.shape)
```
```
print(red['density'].values.shape, red['fixed acidity'].values.shape)
print(x.shape, y.shape)
```
```
# Fit to the data
model.fit(x, y)
# Extract the values of interest
m = model.coef_[0][0]
b = model.intercept_[0]
# Print the slope m and intercept b
print('Scikit learn - Slope: ', m , 'Intercept: ', b )
```
What happens when we try to fit the data as is?
```
# Fit to the data
# model.fit(x, y)
```
#### Exercise 2: drop Null Values (and practice pandas operations)
Let's look back at our dataset description dataframe above, what do we notice, what contains null values?
There are several strategies for dealing with null values. For now let's take the simplest case, and drop rows in our dataframe that contain null
```
# Cell for Exercise 2
# For this templated exercise you are going to complete everything in one line
# of code, but we are going to break it up into steps. So for each part (A, B,
# etc.) paste your answer from the previous part to begin
# step A
# select the 'density' and 'fixed acidity' columns of red. make sure the return
# is a dataframe
# step B
# now use the dropna() method on axis 0 (the rows) to drop any null values
# step B
# select column 'density'
# step C
# select the values
# step D
# reshape the result with an empty second dimension using .reshape() and store
# the result under variable x
# repeat the same process with 'fixed acidity' and variable y
```
Now that we have our x and y arrays we can fit using ScikitLearn
```
x = red[['density', 'fixed acidity']].dropna(axis=0)['density'].values.reshape(-1,1)
y = red[['density', 'fixed acidity']].dropna(axis=0)['fixed acidity'].values.reshape(-1,1)
# Fit to the data
model.fit(x, y)
# Extract the values of interest
m = model.coef_[0][0]
b = model.intercept_[0]
# Print the slope m and intercept b
print('Scikit learn - Slope: ', m , 'Intercept: ', b )
```
#### Exercise 3: calculating y_pred
Estimate the values of $y$ by using your fitted parameters. Hint: Use your <code>model.coef_</code> and <code>model.intercept_</code> parameters to estimate y_pred following equation 1
```
# define y_pred in terms of m, x, and b
# y_pred =
fig, ax = plt.subplots(1,1, figsize=(10,10))
ax.plot(x, y_pred, ls='', marker='*')
ax.plot(x, y, ls='', marker='.')
```
We can also return predictions directly with the model object using the predict() method
```
# Another way to get this is using the model.predict function
y_pred = model.predict(x)
fig, ax = plt.subplots(1,1, figsize=(10,10))
ax.plot(x, y_pred, ls='', marker='*')
ax.plot(x, y, ls='', marker='.')
```
## 1.3 Error and topics of model fitting (assessing model accuracy)
### 1.3.1 Measuring the quality of fit
#### 1.3.1.1 Mean Squared Error
The plot in Section 1.2.3 looks good, but numerically what is our error? What is the mean value of $\epsilon$, i.e. the **Mean Squared Error (MSE)**?
$${\sf MSE}=\epsilon_{\sf ave} = \frac{\sum_{i=1}^{N_{\sf times}}\left(y^{\sf exact}_i - m\cdot t_i - b \right)^2}{N_{\sf times}}\;\;\;\;\;\sf eq. 3$$
```
# The mean squared error
print('Mean squared error: %.2f' % mean_squared_error(y, y_pred))
```
```
# The mean squared error
print('Mean squared error: %.2f' % mean_squared_error(y, y_pred))
```
#### 1.3.1.2 R-square
Another way to measure error is the regression score, $R^2$. $R^2$ is generally defined as the ratio of the total sum of squares $SS_{\sf tot} $ to the residual sum of squares $SS_{\sf res} $:
$$SS_{\sf tot}=\sum_{i=1}^{N} \left(y^{\sf exact}_i-\bar{y}\right)^2\;\;\;\;\; \sf eq. 4$$
$$SS_{\sf res}=\sum_{i=1}^{N} \left(y^{\sf exact}_i - y^{\sf calc}_i\right)^2\;\;\;\;\; \sf eq. 5$$
$$R^2 = 1 - {SS_{\sf res}\over SS_{\sf tot}} \;\;\;\;\;\; \sf eq. 6$$
In eq. 4, $\bar{y}=\sum_i y^{\sf exact}_i/N$ is the average value of y for $N$ points. The best value of $R^2$ is 1 but it can also take a negative value if the error is large.
See all the different regression metrics [here](https://scikit-learn.org/stable/modules/model_evaluation.html).
#### Question 3
> Do we need a large value of $SS_{\sf tot}$ to minimize $R^2$ - is this something which we have the power to control?
```
# Print the coefficient of determination - 1 is perfect prediction
print('Coefficient of determination: %.2f' % r2_score(y, y_pred))
```
```
# Print the coefficient of determination - 1 is perfect prediction
print('Coefficient of determination: %.2f' % r2_score(y, y_pred))
```
### 1.3.2 Corollaries with classification models
For classification tasks, we typically assess accuracy vs MSE or R-square, since we are dealing with categorical rather than numerical predictions.
What is accuracy? It is defined as the ratio of True assignments to all assignments. For a binary positive/negative classification task this can be written as the following:
$ Acc = \frac{T_p + T_n}{F_p + F_n + T_p + T_n} $
Where $T$ is True, $F$ is false, $p$ is positive, $n$ is negative
Just as a quick example, we can perform this type of task on our wine dataset by predicting on quality, which is a discrete 3-9 quality score:
```
y_train = df['type'].values.reshape(-1,1)
x_train = df['quality'].values.reshape(-1,1)
# train a logistic regression model on the training set
from sklearn.linear_model import LogisticRegression
# instantiate model
logreg = LogisticRegression()
# fit model
logreg.fit(x_train, y_train)
# make class predictions for the testing set
y_pred_class = logreg.predict(x_train)
# calculate accuracy
from sklearn import metrics
print(metrics.accuracy_score(y_train, y_pred_class))
```
### 1.3.3 Beyond a single input feature
(_also: quick appreciative beat for folding in domain area expertise into our models and features_)
The **acidity** of the wine (the dependent variable v) could depend on:
* potassium from the soil (increases alkalinity)
* unripe grapes (increases acidity)
* grapes grown in colder climates or reduced sunshine create less sugar (increases acidity)
* preprocessing such as adding tartaric acid to the grape juice before fermentation (increases acidity)
* malolactic fermentation (reduces acidity)
* \+ others
So in our lab today we will look at folding in additional variables in our dataset into the model
<hr style="border:1px solid grey"> </hr>
## 1.4 Multivariate regression
Let's now turn our attention to wine quality.
The value we aim to predict or evaluate is the quality of each wine in our dataset. This is our dependent variable. We will look at how this is related to the 12 other independent variables, also known as *input features*. We're going to do this is just the red wine data
```
red.head()
```
### 1.4.1 Linear regression with all input fields
For this example, notice we have a categorical data variable in the 'type' column. We will ignore this for now, and only work with our red wines. In the future we will discuss how to deal with categorical variable such as this in a mathematical representation.
```
# this is a list of all our features or independent variables
features = list(red.columns[1:])
# we're going to remove our target or dependent variable, density from this
# list
features.remove('density')
# now we define X and y according to these lists of names
X = red.dropna(axis=0)[features].values
y = red.dropna(axis=0)['density'].values
red.isnull().sum(axis=0) # we are getting rid of some nasty nulls!
```
```
# Create linear regression object - note that we are using all the input features
model = linear_model.LinearRegression()
model.fit(X, y)
y_calc = model.predict(X)
```
```
# Create linear regression object - note that we are using all the input features
model = linear_model.LinearRegression()
model.fit(X, y)
y_calc = model.predict(X)
```
Let's see what the coefficients look like ...
```
print("Fit coefficients: \n", model.coef_, "\nNumber of coefficients:", len(model.coef_))
```
```
print("Fit coefficients: \n", model.coef_, "\nNumber of coefficients:", len(model.coef_))
```
We have 11 !!! That's because we are regressing respect to all **11 independent variables**!!!
So now, $$y_{\sf calc}= m_1x_1 +\, m_2x_2 \,+ \,m_3x_3 \,+\,... \,+ \,b =\sum_{i=1}^{13}m_i x_i + b\;\;\;\;\; \sf eq. 7$$
```
print("We have 13 slopes / weights:\n\n", model.coef_)
print("\nAnd one intercept: ", model.intercept_)
```
```
print("We have 11 slopes / weights:\n\n", model.coef_)
print("\nAnd one intercept: ", model.intercept_)
```
```
# This size should match the number of columns in X
if len(X[0]) == len(model.coef_):
print("All good! The number of coefficients matches the number of input features.")
else:
print("Hmm .. something strange is going on.")
```
```
# This size should match the number of columns in X
if len(X[0]) == len(model.coef_):
print("All good! The number of coefficients matches the number of input features.")
else:
print("Hmm .. something strange is going on.")
```
### 1.4.2 Exercise 4: evaluate the error
Let's **evaluate the error** by computing the MSE and $R^2$ metrics (see eq. 3 and 6).
```
# The mean squared error
# part A
# calculate the MSE using mean_squared_error()
# mse =
# part B
# calculate the R square using r2_score()
# r2 =
print('Mean squared error: {:.2f}'.format(mse)
print('Coefficient of determination: {:.2f}'.format(r2)
```
```
# The mean squared error
# part A
# calculate the MSE using mean_squared_error()
# mse =
# part B
# calculate the R square using r2_score()
# r2 =
print('Mean squared error: {:.2f}'.format(mse)
print('Coefficient of determination: {:.2f}'.format(r2)
```
### 1.4.3 Exercise 5: make a plot of y actual vs y predicted
We can also look at how well the computed values match the true values graphically by generating a scatterplot.
```
# generate a plot of y predicted vs y actual using plt.plot()
# remember you must set ls to an empty string and marker to some marker style
# plt.plot()
plt.title("Linear regression - computed values on entire data set", fontsize=16)
plt.xlabel("y$^{\sf calc}$")
plt.ylabel("y$^{\sf true}$")
plt.show()
```
```
# generate a plot of y predicted vs y actual using plt.plot()
# remember you must set ls to an empty string and marker to some marker style
# plt.plot()
plt.title("Linear regression - computed values on entire data set", fontsize=16)
plt.xlabel("y$^{\sf calc}$")
plt.ylabel("y$^{\sf true}$")
plt.show()
```
### 1.4.2 **Enrichment**: Splitting into train and test sets
To see whether we can predict, we will carry out our regression only on a part, 80%, of the full data set. This part is called the **training** data. We will then test the trained model to predict the rest of the data, 20% - the **test** data. The function which fits won't see the test data until it has to predict it.
**We will motivate the use of train/test sets more explicitly in Course 2 Session 1**
We start by splitting out data using scikit-learn's <code> train_test_split() </code> function:
```
X_train, X_test, y_train, y_test = train_test_split(X, y,
test_size=0.20,
random_state=42)
```
```
X_train, X_test, y_train, y_test = train_test_split(X, y,
test_size=0.20,
random_state=42)
```
Now we check the size of <code> y_train </code> and <code> y_test </code>, the sum should be the size of y! If this works then we move on and carry out regression but we only use the training data!
```
if len(y_test)+len(y_train) == len(y):
print('All good, ready to to go and regress!\n')
# Carry out linear regression
print('Running linear regression algorithm on the training set\n')
model = linear_model.LinearRegression()
model.fit(X_train, y_train)
print('Fit coefficients and intercept:\n\n', model.coef_, '\n\n', model.intercept_ )
# Predict on the test set
y_pred_test = model.predict(X_test)
```
```
if len(y_test)+len(y_train) == len(y):
print('All good, ready to to go and regress!\n')
# Carry out linear regression
print('Running linear regression algorithm on the training set\n')
model = linear_model.LinearRegression()
model.fit(X_train, y_train)
print('Fit coefficients and intercept:\n\n', model.coef_, '\n\n', model.intercept_ )
# Predict on the test set
y_pred_test = model.predict(X_test)
```
Now we can plot our predicted values to see how accurate we are in predicting. We will generate a scatterplot and computing the MSE and $R^2$ metrics of error.
```
sns.scatterplot(x=y_pred_test, y=y_test, color="mediumvioletred", s=50)
plt.title("Linear regression - predict test set", fontsize=16)
plt.xlabel("y$^{\sf calc}$")
plt.ylabel("y$^{\sf true}$")
plt.show()
print('Mean squared error: %.2f' % mean_squared_error(y_test, y_pred_test))
print('Coefficient of determination: %.2f' % r2_score(y_test, y_pred_test))
```
```
sns.scatterplot(x=y_pred_test, y=y_test, color="mediumvioletred", s=50)
plt.title("Linear regression - predict test set", fontsize=16)
plt.xlabel("y$^{\sf calc}$")
plt.ylabel("y$^{\sf true}$")
plt.show()
print('Mean squared error: %.2f' % mean_squared_error(y_test, y_pred_test))
print('Coefficient of determination: %.2f' % r2_score(y_test, y_pred_test))
```
#### 1.4.2.1 Other data considerations
* Do we need all the independent variables?
* Topics of interential statistics covered in a couple sessions
* Can we output integer quality scores?
* Topics of non-binary classification tasks covered in week 4
### 1.4.3 **Enrichment**: Other regression algorithms
There are many other regression algorithms the two we want to highlight here are Ridge, LASSO, and Elastic Net. They differ by an added term to the loss function. Let's review. Eq. 2 expanded to multivariate form yields:
$$\sum_{i=1}^{N}(y_i - \sum_{j=1}^{P}x_{ij}\beta_{j})^2$$
for Ridge regression, we add a **_regularization_** term known as **_L2_** regularization:
$$\sum_{i=1}^{N}(y_i - \sum_{j=1}^{P}x_{ij}\beta_{j})^2 + \lambda \sum_{j=1}^{P}\beta_{j}^2$$
for **_LASSO_** (Least Absolute Shrinkage and Selection Operator) we add **_L1_** regularization:
$$\sum_{i=1}^{N}(y_i - \sum_{j=1}^{P}x_{ij}\beta_{j})^2 + \lambda \sum_{j=1}^{P}|\beta_{j}|$$
The key difference here is that LASSO will allow coefficients to shrink to 0 while Ridge regression will not. **_Elastic Net_** is a combination of these two regularization methods.
```
model = linear_model.Ridge()
model.fit(X_train, y_train)
print('Fit coefficients and intercept:\n\n', model.coef_, '\n\n', model.intercept_ )
# Predict on the test set
y_calc_test = model.predict(X_test)
```
```
model = linear_model.Ridge()
model.fit(X_train, y_train)
print('Fit coefficients and intercept:\n\n', model.coef_, '\n\n', model.intercept_ )
# Predict on the test set
y_calc_test = model.predict(X_test)
```
```
sns.scatterplot(x=y_calc_test, y=y_test, color="lightseagreen", s=50)
plt.title("Ridge regression - predict test set",fontsize=16)
plt.xlabel("y$^{\sf calc}$")
plt.ylabel("y$^{\sf true}$")
plt.show()
print('Mean squared error: %.2f' % mean_squared_error(y_test, y_calc_test))
print('Coefficient of determination: %.2f' % r2_score(y_test, y_calc_test))
```
```
sns.scatterplot(x=y_calc_test, y=y_test, color="lightseagreen", s=50)
plt.title("Ridge regression - predict test set",fontsize=16)
plt.xlabel("y$^{\sf calc}$")
plt.ylabel("y$^{\sf true}$")
plt.show()
print('Mean squared error: %.2f' % mean_squared_error(y_test, y_calc_test))
print('Coefficient of determination: %.2f' % r2_score(y_test, y_calc_test))
```
#### Exercise 6: Tune Hyperparameter for Ridge Regression
Use the docstring to peak into the hyperparameters for Ridge Regression. What is the optimal value of lambda?
Plot the $\beta$ values vs $\lambda$ from the results of your analysis
```
# cell for exercise 3
out_lambdas = []
out_coefs = []
out_scores = []
for i in range(10):
lambdas = []
coefs = []
scores = []
X_train, X_test, y_train, y_test = train_test_split(X, y,
test_size=0.20)
for lamb in range(1,110):
model = linear_model.Ridge(alpha=lamb/50, normalize=True)
model.fit(X_train, y_train)
lambdas.append(lamb)
coefs.append(model.coef_)
scores.append(r2_score(y_test, model.predict(X_test)))
# print('MSE: %.4f' % mean_squared_error(y_test, model.predict(X_test)))
# print('R2: %.4f' % r2_score(y_test, model.predict(X_test)))
out_lambdas.append(lambdas)
out_coefs.append(coefs)
out_scores.append(scores)
coef_means = np.array(out_coefs).mean(axis=0)
coef_stds = np.array(out_coefs).std(axis=0)
results_means = pd.DataFrame(coef_means,columns=features)
results_stds = pd.DataFrame(coef_stds,columns=features)
results_means['lambda'] = [i/50 for i in lambdas]
fig, ax = plt.subplots(1,1,figsize=(10,10))
for feat in features:
ax.errorbar([i/50 for i in lambdas], results_means[feat], yerr=results_stds[feat], label=feat)
# results.plot('lambda', 'scores', ax=ax[1])
ax.legend()
results = pd.DataFrame(coefs,columns=features)
results['lambda'] = [i/50 for i in lambdas]
results['scores'] = scores
fig, ax = plt.subplots(1,2,figsize=(10,5))
for feat in features:
results.plot('lambda', feat, ax=ax[0])
results.plot('lambda', 'scores', ax=ax[1])
```
## 1.5 **Enrichment**: Additional Regression Exercises
### Problem 1) Number and choice of input features
* Load the red wine dataset and evaluate how the linear regression predictions changes as you change the **number and choice of input features**. The total number of columns in X is 11 and each column represent a specific input feature.
* Estimate the MSE
```
print(X_train.shape)
```
```
print(X_train.shape)
```
If you want to use the first 5 features you could proceed as following:
```
X_train_five = X_train[:,0:5]
X_test_five = X_test[:,0:5]
```
```
X_train_five = X_train[:,0:5]
X_test_five = X_test[:,0:5]
```
Check that the new variables have the shape your expect
```
print(X_train_five.shape)
print(X_test_five.shape)
```
```
print(X_train_five.shape)
print(X_test_five.shape)
```
Now you can use these to train your linear regression model and repeat for different numbers or sets of input features! Note that you do not need to change the output feature! It's size is independent from the number of input features, yet recall that its length is the same as the number of values per input feature.
Questions to think about while you work on this problem
- How many input feature variables does one need? Is there a maximum or minimum number?
- Could one input feature variable be better than the rest?
- What if values are missing for one of the input feature variables - is it still worth using it?
- Can you use **_L1_** or **_L2_** to determine these optimum features more quickly?
### Problem 2) Type of regression algorithm
Try using other types of linear regression methods on the wine dataset: the LASSO model and the Elastic net model which are described by the
<code > sklearn.linear_model.ElasticNet() </code> <br>
<code > sklearn.linear_model.Lasso() </code>
scikit-learn functions.
For more detail see [ElasticNet](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.ElasticNet.html#sklearn.linear_model.ElasticNet) and [Lasso]( https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.Lasso.html#sklearn.linear_model.Lasso).
Questions to think about while you work on this problem
- How does the error change with each model?
- Which model seems to perform best?
- How can you optimize the hyperparameter, $\lambda$
- Does one model do better than the other at determining which input features are more important?
- How about non linear regression / what if the data does not follow a line?
- How do the bias and variance change for each model
```
from sklearn.linear_model import ElasticNet
from sklearn.linear_model import Lasso
from sklearn.linear_model import Ridge
from sklearn.linear_model import LinearRegression
for model in [ElasticNet, Lasso, Ridge, LinearRegression]:
model = model()
model.fit(X_train, y_train)
print('Mean squared error: %.2f' % mean_squared_error(y_test, model.predict(X_test)))
print('Coefficient of determination: %.2f' % r2_score(y_test, model.predict(X_test)))
```
<hr style="border:1px solid grey"> </hr>
# References
* **Linear Regression**
To find out more see https://en.wikipedia.org/wiki/Simple_linear_regression
* **scikit-learn**
* Scikit-learn: https://scikit-learn.org/stable/
* Linear regression in scikit-learn: https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LinearRegression.html
* Metrics of error: https://scikit-learn.org/stable/modules/model_evaluation.html
* The Boston dataset: https://scikit-learn.org/stable/datasets/index.html#boston-dataset
* **Pearson correlation**
To find out more see https://en.wikipedia.org/wiki/Pearson_correlation_coefficient
* **Irreducible error, bias and variance**
* Great Coursera videos here: https://www.coursera.org/lecture/ml-regression/irreducible-error-and-bias-qlMrZ
and here: https://www.coursera.org/lecture/ml-regression/variance-and-the-bias-variance-tradeoff-ZvP40
| github_jupyter |

[](https://colab.research.google.com/github/JohnSnowLabs/spark-nlp-workshop/blob/master/tutorials/Certification_Trainings/Healthcare/8.Generic_Classifier.ipynb)
# Generic Classifier
```
import json
with open('workshop_license_keys_Aug2020.json') as f:
license_keys = json.load(f)
license_keys.keys()
license_keys['JSL_VERSION']
# template for license_key.json
{'JSL_VERSION':'jjj',
'PUBLIC_VERSION':'vvv',
'SECRET':"xxx",
'SPARK_NLP_LICENSE': 'aaa',
'JSL_OCR_LICENSE': 'bbb',
'AWS_ACCESS_KEY_ID':"ccc",
'AWS_SECRET_ACCESS_KEY':"ddd",
'JSL_OCR_SECRET':"eee"}
import os
# Install java
! apt-get update -qq
! apt-get install -y openjdk-8-jdk-headless -qq > /dev/null
os.environ["JAVA_HOME"] = "/usr/lib/jvm/java-8-openjdk-amd64"
os.environ["PATH"] = os.environ["JAVA_HOME"] + "/bin:" + os.environ["PATH"]
! java -version
secret = license_keys['SECRET']
os.environ['SPARK_NLP_LICENSE'] = license_keys['SPARK_NLP_LICENSE']
os.environ['SPARK_OCR_LICENSE'] = license_keys['SPARK_OCR_LICENSE']
os.environ['AWS_ACCESS_KEY_ID']= license_keys['AWS_ACCESS_KEY_ID']
os.environ['AWS_SECRET_ACCESS_KEY'] = license_keys['AWS_SECRET_ACCESS_KEY']
jsl_version = license_keys['JSL_VERSION']
version = license_keys['PUBLIC_VERSION']
! pip install --ignore-installed -q pyspark==2.4.4
! python -m pip install --upgrade spark-nlp-jsl==$jsl_version --extra-index-url https://pypi.johnsnowlabs.com/$secret
! pip install --ignore-installed -q spark-nlp==$version
import sparknlp
print (sparknlp.version())
import json
import os
from pyspark.ml import Pipeline
from pyspark.sql import SparkSession
from sparknlp.annotator import *
from sparknlp_jsl.annotator import *
from sparknlp.base import *
import sparknlp_jsl
spark = sparknlp_jsl.start(secret)
# if you want to start the session with custom params as in start function above
def start(secret):
builder = SparkSession.builder \
.appName("Spark NLP Licensed") \
.master("local[*]") \
.config("spark.driver.memory", "16G") \
.config("spark.serializer", "org.apache.spark.serializer.KryoSerializer") \
.config("spark.kryoserializer.buffer.max", "2000M") \
.config("spark.jars.packages", "com.johnsnowlabs.nlp:spark-nlp_2.11:"+version) \
.config("spark.jars", "https://pypi.johnsnowlabs.com/"+secret+"/spark-nlp-jsl-"+jsl_version+".jar")
return builder.getOrCreate()
#spark = start(secret)
spark
```
## load dataset
```
!wget -q https://raw.githubusercontent.com/JohnSnowLabs/spark-nlp-workshop/master/tutorials/Certification_Trainings/Healthcare/data/petfinder-mini.csv
import pandas as pd
dataframe = pd.read_csv('petfinder-mini.csv')
# In the original dataset "4" indicates the pet was not adopted.
import numpy as np
dataframe['target'] = np.where(dataframe['AdoptionSpeed']==4, 0, 1)
dataframe = dataframe.drop(['AdoptionSpeed'], axis=1)
dataframe.head()
dataframe.columns
dataframe.info()
dataframe.target.value_counts()
dataframe.Description = dataframe.Description.fillna('- no description -')
```
## Featurize with Sklearn Column Transformer
```
from sklearn.compose import make_column_transformer
from sklearn.compose import ColumnTransformer
from sklearn.feature_extraction.text import CountVectorizer, TfidfVectorizer
from sklearn.preprocessing import OneHotEncoder, StandardScaler
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
column_trans = make_column_transformer(
(OneHotEncoder(), ['Type', 'Breed1', 'Gender', 'Color1', 'Color2', 'MaturitySize',
'FurLength', 'Vaccinated', 'Sterilized', 'Health']),
(TfidfVectorizer(max_features=1000, norm='l2', ngram_range=(1, 3)), 'Description'),
remainder=StandardScaler())
X = column_trans.fit_transform(dataframe.drop(['target'], axis=1))
y = dataframe.target
X.shape
df = pd.DataFrame.sparse.from_spmatrix(X)
df.columns = ['col_{}'.format(i) for i in range(X.shape[1])]
df['target']= y
df.head()
```
## Train with Spark NLP Generic Classifier
**Building a pipeline**
The FeaturesAssembler is used to collect features from different columns. It can collect features from single value columns (anything which can be cast to a float, if casts fails then the value is set to 0), array columns or SparkNLP annotations (if the annotation is an embedding, it takes the embedding, otherwise tries to cast the 'result' field). The output of the transformer is a FEATURE_VECTOR annotation (the numeric vector is in the 'embeddings' field).
The GenericClassifierApproach takes FEATURE_VECTOR annotations as input, classifies them and outputs CATEGORY annotations. The operation of the classifier is controled by the following methods:
*setEpochsNumber(int)* - Determines how many epochs the model is trained.
*setBatchSize(int)* - Sets the batch size during training.
*setLearningRate(float)* - Sets the learning rate.
*setValidationSplit(float)* - Sets the proportion of examples in the training set used for validation.
*setModelFile(string)* - Loads a model from the specified location and uses it instead of the default model.
*setFixImbalance(boolean)* - If set to true, it tries to balance the training set by weighting the classes according to the inverse of the examples they have.
*setFeatureScaling(string)* - Normalizes the feature factors using the specified method ("zscore", "minmax" or empty for no normalization).
*setOutputLogsPath(string)* - Sets the path to a folder where logs of training progress will be saved. No logs are generated if no path is specified.
```
spark_df = spark.createDataFrame(df)
spark_df.show(2)
(training_data, test_data) = spark_df.randomSplit([0.8, 0.2], seed = 100)
print("Training Dataset Count: " + str(training_data.count()))
print("Test Dataset Count: " + str(test_data.count()))
!mkdir gc_graph
!wget -q https://raw.githubusercontent.com/JohnSnowLabs/spark-nlp-workshop/master/tutorials/Certification_Trainings/Healthcare/generic_classifier_graph/pet.in1202D.out2.pb -P /content/gc_graph
from sparknlp_jsl.base import *
features_asm = FeaturesAssembler()\
.setInputCols(['col_{}'.format(i) for i in range(X.shape[1])])\
.setOutputCol("features")
gen_clf = GenericClassifierApproach()\
.setLabelColumn("target")\
.setInputCols(["features"])\
.setOutputCol("prediction")\
.setModelFile('/content/gc_graph/pet.in1202D.out2.pb')\
.setEpochsNumber(50)\
.setBatchSize(100)\
.setFeatureScaling("zscore")\
.setFixImbalance(True)\
.setLearningRate(0.001)\
.setOutputLogsPath("logs")\
.setValidationSplit(0.2) # keep 20% of the data for validation purposes
clf_Pipeline = Pipeline(stages=[
features_asm,
gen_clf])
clf_model = clf_Pipeline.fit(training_data)
! cd logs && ls
!cat logs/GenericClassifierApproach_76b7941dd049.log
pred_df = clf_model.transform(test_data)
pred_df.select('target','prediction.result').show()
preds_df = pred_df.select('target','prediction.result').toPandas()
# Let's explode the array and get the item(s) inside of result column out
preds_df['result'] = preds_df['result'].apply(lambda x : int(x[0]))
preds_df
# We are going to use sklearn to evalute the results on test dataset
from sklearn.metrics import classification_report, accuracy_score
print (classification_report(preds_df['result'], preds_df['target'], digits=4))
print (accuracy_score(preds_df['result'], preds_df['target']))
```
## get prediction for random input
```
pd.DataFrame([dataframe.loc[5191].to_dict()])
input_X = column_trans.transform(pd.DataFrame([dataframe.loc[0].to_dict()]).drop(['target'], axis=1))
input_y = dataframe.target[0]
input_df = pd.DataFrame.sparse.from_spmatrix(input_X)
input_df.columns = ['col_{}'.format(i) for i in range(input_X.shape[1])]
input_df['target']= input_y
input_df.head()
input_spark_df = spark.createDataFrame(input_df)
input_spark_df.show(2)
clf_model.transform(input_spark_df).select('target','prediction.result').show()
```
# GC TF Graph
```
!wget -q https://raw.githubusercontent.com/JohnSnowLabs/spark-nlp-workshop/master/tutorials/Certification_Trainings/Healthcare/generic_classifier_graph/basetfmodel.py
!wget -q https://raw.githubusercontent.com/JohnSnowLabs/spark-nlp-workshop/master/tutorials/Certification_Trainings/Healthcare/generic_classifier_graph/build_model.py
!wget -q https://raw.githubusercontent.com/JohnSnowLabs/spark-nlp-workshop/master/tutorials/Certification_Trainings/Healthcare/generic_classifier_graph/generic_classifier_model.py
!wget -q https://raw.githubusercontent.com/JohnSnowLabs/spark-nlp-workshop/master/tutorials/Certification_Trainings/Healthcare/generic_classifier_graph/progresstracker.py
!wget -q https://raw.githubusercontent.com/JohnSnowLabs/spark-nlp-workshop/master/tutorials/Certification_Trainings/Healthcare/generic_classifier_graph/settings.py
from generic_classifier_model import GenericClassifierModel
from settings import Settings
build_params = {
"hidden_layers": [400, 200, 100, 50],
#"hidden_act_l2": 0.0001, ##l2 activation regularization of hidden layers
#"hidden_weights_l2": 0.0001, ##l2 weight regularization of hidden layers
"batch_norm": True,
"hidden_act": "relu",
"input_dim": 1202,
"output_dim": 2,
}
model = GenericClassifierModel("pet")
model.build(build_params)
model.export_graph(
model_dir='/content/gc_graph',
log_dir='/content/gc_graph')
```
| github_jupyter |
## Hosting a Pretrained Model on SageMaker
Amazon SageMaker is a service to accelerate the entire machine learning lifecycle. It includes components for building, training and deploying machine learning models. Each SageMaker component is modular, so you're welcome to only use the features needed for your use case. One of the most popular features of SageMaker is [model hosting](https://docs.aws.amazon.com/sagemaker/latest/dg/how-it-works-deployment.html). Using SageMaker Hosting you can deploy your model as a scalable, highly available, multi-process API endpoint with a few lines of code. In this notebook, we will demonstrate how to host a pretrained model (GPT-2) in Amazon SageMaker.
SageMaker provides prebuilt containers that can be used for training, hosting, or data processing. The inference containers include a web serving stack, so you don't need to install and configure one. We will be using the SageMaker [PyTorch container](https://github.com/aws/deep-learning-containers), but you may use the [TensorFlow container](https://github.com/aws/deep-learning-containers/blob/master/available_images.md), or bring your own container if needed.
This notebook will walk you through how to deploy a pretrained Hugging Face model as a scalable, highly available, production ready API in under 15 minutes.
## Retrieve Model Artifacts
First we will download the model artifacts for the pretrained [GPT-2](https://d4mucfpksywv.cloudfront.net/better-language-models/language_models_are_unsupervised_multitask_learners.pdf) model. GPT-2 is a popular text generation model that was developed by OpenAI. Given a text prompt it can generate synthetic text that may follow.
```
!pip install transformers==3.3.1 sagemaker==2.15.0 --quiet
import os
from transformers import GPT2Tokenizer, GPT2LMHeadModel
tokenizer = GPT2Tokenizer.from_pretrained('gpt2')
model = GPT2LMHeadModel.from_pretrained('gpt2')
model_path = 'model/'
code_path = 'code/'
if not os.path.exists(model_path):
os.mkdir(model_path)
model.save_pretrained(save_directory=model_path)
tokenizer.save_vocabulary(save_directory=model_path)
```
## Write the Inference Script
Since we are bringing a model to SageMaker, we must create an inference script. The script will run inside our PyTorch container. Our script should include a function for model loading, and optionally functions generating predicitions, and input/output processing. The PyTorch container provides default implementations for generating a prediction and input/output processing. By including these functions in your script you are overriding the default functions. You can find additional [details here](https://sagemaker.readthedocs.io/en/stable/frameworks/pytorch/using_pytorch.html#serve-a-pytorch-model).
In the next cell we'll see our inference script. You will notice that it uses the [transformers library from Hugging Face](https://huggingface.co/transformers/). This Python library is not installed in the container by default, so we will have to add that in the next section.
```
!pygmentize code/inference_code.py
```
## Package Model
For hosting, SageMaker requires that the deployment package be structed in a compatible format. It expects all files to be packaged in a tar archive named "model.tar.gz" with gzip compression. To install additional libraries at container startup, we can add a [requirements.txt](https://sagemaker.readthedocs.io/en/stable/frameworks/pytorch/using_pytorch.html#using-third-party-libraries) text file that specifies the libraries to be installed using [pip](https://pypi.org/project/pip/). Within the archive, the PyTorch container expects all inference code and requirements.txt file to be inside the code/ directory. See the [guide here](https://sagemaker.readthedocs.io/en/stable/frameworks/pytorch/using_pytorch.html#for-versions-1-2-and-higher) for a thorough explanation of the required directory structure.
```
import tarfile
zipped_model_path = os.path.join(model_path, "model.tar.gz")
with tarfile.open(zipped_model_path, "w:gz") as tar:
tar.add(model_path)
tar.add(code_path)
```
## Deploy Model
Now that we have our deployment package, we can use the [SageMaker SDK](https://sagemaker.readthedocs.io/en/stable/index.html) to deploy our API endpoint with two lines of code. We need to specify an IAM role for the SageMaker endpoint to use. Minimally, it will need read access to the default SageMaker bucket (usually named sagemaker-{region}-{your account number}) so it can read the deployment package. When we call deploy(), the SDK will save our deployment archive to S3 for the SageMaker endpoint to use. We will use the helper function [get_execution_role](https://sagemaker.readthedocs.io/en/stable/api/utility/session.html?highlight=get_execution_role#sagemaker.session.get_execution_role) to retrieve our current IAM role so we can pass it to the SageMaker endpoint. You may specify another IAM role here. Minimally it will require read access to the model artifacts in S3 and the [ECR repository](https://github.com/aws/deep-learning-containers/blob/master/available_images.md) where the container image is stored by AWS.
You may notice that we specify our PyTorch version and Python version when creating the PyTorchModel object. The SageMaker SDK uses these parameters to determine which PyTorch container to use.
The full size [GPT-2 model has 1.2 billion parameters](https://d4mucfpksywv.cloudfront.net/better-language-models/language_models_are_unsupervised_multitask_learners.pdf). Even though we are using the small version of the model, our endpoint will need to fit millions of parameters in to memory. We'll choose an m5 instance for our endpoint to ensure we have sufficient memory to serve our model.
```
from sagemaker.pytorch import PyTorchModel
from sagemaker import get_execution_role
endpoint_name = 'GPT2'
model = PyTorchModel(entry_point='inference_code.py',
model_data=zipped_model_path,
role=get_execution_role(),
framework_version='1.5',
py_version='py3')
predictor = model.deploy(initial_instance_count=1,
instance_type='ml.m5.xlarge',
endpoint_name=endpoint_name)
```
## Get Predictions
Now that our RESTful API endpoint is deployed, we can send it text to get predictions from our GPT-2 model. You can use the SageMaker Python SDK or the [SageMaker Runtime API](https://docs.aws.amazon.com/sagemaker/latest/APIReference/API_runtime_InvokeEndpoint.html) to invoke the endpoint.
```
import boto3
import json
sm = boto3.client('sagemaker-runtime')
prompt = "Working with SageMaker makes machine learning "
response = sm.invoke_endpoint(EndpointName=endpoint_name,
Body=json.dumps(prompt),
ContentType='text/csv')
response['Body'].read().decode('utf-8')
```
## Conclusion
You have successfully created a scalable, high available, RESTful API that is backed by a GPT-2 model! If you are still interested in learning more, check out some of the more advanced features of SageMaker Hosting, like [model monitoring](https://docs.aws.amazon.com/sagemaker/latest/dg/model-monitor.html) to detect concept drift, [autoscaling](https://docs.aws.amazon.com/sagemaker/latest/dg/endpoint-auto-scaling.html) to dynamically adjust the number of instances, or [VPC config](https://docs.aws.amazon.com/sagemaker/latest/dg/host-vpc.html) to control network access to/from your endpoint.
You can also look in to the [ezsmdeploy SDK](https://aws.amazon.com/blogs/opensource/deploy-machine-learning-models-to-amazon-sagemaker-using-the-ezsmdeploy-python-package-and-a-few-lines-of-code/) that automates most of this process.
| github_jupyter |
# Chapter 4 - Evaluation and Optimization
```
%pylab inline
import pandas as pandas
```
We generate two inputs:
* features – a matrix of input features
* target – an array of target variables corresponding to those features
```
features = rand(100,5)
target = rand(100) > 0.5
```
### The holdout method
We divide into a randomized training and test set:
```
int(floor(0.7*100))
N = features.shape[0]
N_train = int(floor(0.7 * N))
# Randomize index
# Note: sometimes you want to retain the order in the dataset and skip this step
# E.g. in the case of time-based datasets where you want to test on 'later' instances
idx = random.permutation(N)
# Split index
idx_train = idx[:N_train]
idx_test = idx[N_train:]
# Break your data into training and testing subsets
features_train = features[idx_train,:]
target_train = target[idx_train]
features_test = features[idx_test,:]
target_test = target[idx_test]
# Build, predict, evaluate (to be filled out)
# model = train(features_train, target_train)
# preds_test = predict(model, features_test)
# accuracy = evaluate_acc(preds_test, target_test)
print(features_train.shape)
print(features_test.shape)
print(target_train.shape)
print(target_test.shape)
```
### K-fold cross-validation
```
N = features.shape[0]
K = 10 # number of folds
preds_kfold = np.empty(N)
folds = np.random.randint(0, K, size=N)
print(folds)
for idx in np.arange(K):
# For each fold, break your data into training and testing subsets
features_train = features[folds != idx,:]
target_train = target[folds != idx]
features_test = features[folds == idx,:]
# Print the indices in each fold, for inspection
print("Positions of "+str(idx)+" in fold array: ", end="")
print(nonzero(folds == idx)[0])
# Build and predict for CV fold (to be filled out)
# model = train(features_train, target_train)
# preds_kfold[folds == idx] = predict(model, features_test)
# accuracy = evaluate_acc(preds_kfold, target)
```
### The ROC curve
```
def roc_curve(true_labels, predicted_probs, n_points=100, pos_class=1):
thr = linspace(0,1,n_points)
tpr = zeros(n_points)
fpr = zeros(n_points)
pos = true_labels == pos_class
neg = logical_not(pos)
n_pos = count_nonzero(pos)
n_neg = count_nonzero(neg)
for i,t in enumerate(thr):
tpr[i] = count_nonzero(logical_and(predicted_probs >= t, pos)) / n_pos
fpr[i] = count_nonzero(logical_and(predicted_probs >= t, neg)) / n_neg
return fpr, tpr, thr
# Randomly generated predictions should give us a diagonal ROC curve
preds = rand(len(target))
fpr, tpr, thr = roc_curve(target, preds, pos_class=True)
plot(fpr, tpr)
```
### The area under the ROC curve
```
def auc(true_labels, predicted_labels, pos_class=1):
fpr, tpr, thr = roc_curve(true_labels, predicted_labels,
pos_class=pos_class)
area = -trapz(tpr, x=fpr)
return area
auc(target, preds, pos_class=True)
```
### Multi-class classification
```
d = pandas.read_csv("data/mnist_small.csv")
d_train = d[:int(0.8*len(d))]
d_test = d[int(0.8*len(d)):]
from sklearn.ensemble import RandomForestClassifier
rf = RandomForestClassifier()
rf.fit(d_train.drop('label', axis=1), d_train['label'])
from sklearn.metrics import confusion_matrix
preds = rf.predict(d_test.drop('label', axis=1))
cm = confusion_matrix(d_test['label'], preds)
matshow(cm, cmap='Greys')
colorbar()
savefig("figures/figure-4.19.eps", format='eps')
```
### The root-mean-square error
```
def rmse(true_values, predicted_values):
n = len(true_values)
residuals = 0
for i in range(n):
residuals += (true_values[i] - predicted_values[i])**2.
return np.sqrt(residuals/n)
rmse(rand(10), rand(10))
```
### The R-squared error
```
def r2(true_values, predicted_values):
n = len(true_values)
mean = np.mean(true_values)
residuals = 0
total = 0
for i in range(n):
residuals += (true_values[i] - predicted_values[i])**2.
total += (true_values[i] - mean)**2.
return 1.0 - residuals/total
r2(arange(10)+rand(), arange(10)+rand(10))
```
### Grid search with kernel-SVM model
Importing modules:
```
from sklearn.metrics import roc_auc_score
from sklearn.svm import SVC
```
Loading data and performang poor-mans feature engineering:
```
d = pandas.read_csv("data/titanic.csv")
# Target
y = d["Survived"]
# Features
X = d.drop(["Survived", "PassengerId", "Cabin","Ticket","Name", "Fare"], axis=1)
X['Sex'] = list(map(lambda x: 1 if x=="male" else 0, X['Sex']))
X['Embarked-Q'] = list(map(lambda x: 1 if x=="Q" else 0, X['Embarked']))
X['Embarked-C'] = list(map(lambda x: 1 if x=="C" else 0, X['Embarked']))
X['Embarked-S'] = list(map(lambda x: 1 if x=="S" else 0, X['Embarked']))
X = X.drop(["Embarked"], axis=1)
X = X.fillna(-1)
gam_vec, cost_vec = np.meshgrid(np.logspace(0.01, 0.1, 11),
np.linspace(1, 5, 10))
print(gam_vec)
print(cost_vec)
```
Performing grid-search to find the optimal hyper-parameters:
```
# grid of (gamma, C) values to try
gam_vec, cost_vec = np.meshgrid(np.logspace(0.01, 0.1, 11),
np.linspace(1, 5, 10))
AUC_all = [] # initialize empty array to store AUC results
# set up cross-validation folds
N = len(y)
K = 10 # number of cross-validation folds
folds = np.random.randint(0, K, size=N)
# search over every value of the grid
for param_ind in np.arange(len(gam_vec.ravel())):
# initialize cross-validation predictions
y_cv_pred = np.empty(N)
# loop through the cross-validation folds
for ii in np.arange(K):
# break your data into training and testing subsets
# X_train = X.ix[folds != ii,:]
# y_train = y.ix[folds != ii]
# X_test = X.ix[folds == ii,:]
X_train = X.iloc[folds != ii,:]
y_train = y.iloc[folds != ii]
X_test = X.iloc[folds == ii,:]
#X_train = X.iloc[folds, :]
#X_train = X_train.drop(ii)
#y_train = y.iloc[folds]
#y_train = y.drop(ii)
#X_test = X.iloc[folds, :]
#X_test = X_test[folds == ii]
# build a model on the training set
model = SVC(gamma=gam_vec.ravel()[param_ind], C=cost_vec.ravel()[param_ind])
model.fit(X_train, y_train)
# generate and store model predictions on the testing set
y_cv_pred[folds == ii] = model.predict(X_test)
# evaluate the AUC of the predictions
AUC_all.append(roc_auc_score(y, y_cv_pred))
indmax = np.argmax(AUC_all)
print("Maximum = %.3f" % (np.max(AUC_all)))
print("Tuning Parameters: (gamma = %.2f, C = %.2f)" % (gam_vec.ravel()[indmax], cost_vec.ravel()[indmax]))
ix=2
print(folds)
# Train subset taking all rows except the ones with index == to the positions of ix in the folds array
X_train = X.iloc[folds!=ix,:]
print(X_train.head(20))
X_test = X.iloc[folds==ix,:]
print(X_test.head(20))
```
Plotting the contours of the parameter performance:
```
AUC_grid = np.array(AUC_all).reshape(gam_vec.shape)
contourf(gam_vec, cost_vec, AUC_grid, 20, cmap='Greys')
xlabel("kernel coefficient, gamma")
ylabel("penalty parameter, C")
colorbar()
savefig("figures/figure-4.25.eps", format='eps')
```
| github_jupyter |
```
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import sklearn.preprocessing
import sklearn.neural_network
import warnings
warnings.simplefilter(action='ignore', category=FutureWarning)
import h5py
warnings.resetwarnings()
warnings.simplefilter(action='ignore', category=ImportWarning)
warnings.simplefilter(action='ignore', category=RuntimeWarning)
warnings.simplefilter(action='ignore', category=DeprecationWarning)
warnings.simplefilter(action='ignore', category=ResourceWarning)
import theano
import theano.tensor as T
import pickle, gzip
f = gzip.open('mnist.pkl.gz','rb')
train_set, valid_set, test_set = pickle.load(f,encoding='latin-1')
f.close()
train_set[0][train_set[0]>=0.5] = 1
train_set[0][train_set[0]<0.5] = 0
test_set[0][test_set[0]>=0.5] = 1
test_set[0][test_set[0]<0.5] = 0
x_train=theano.function([], T.concatenate([train_set[0], valid_set[0]]))()
x_test = theano.function([], theano.shared(test_set[0]))()
n_hidden = 100 # the size of hidden layers in MLP
n_latent = 3 # the dimension of z
n_input = x_train.shape[1] # the dimension of x's feature space
batch_size = 100
n_epochs = 10000
def init_w(shape):
x = np.random.randn(*shape)
float_x = np.asarray(x * 0.01, dtype=theano.config.floatX)
return theano.shared(float_x)
# Parameters
# Gaussian MLP weights and biases (encoder)
#initialize \phi
b3 = init_w((n_hidden, ))
b2 = init_w((n_latent, ))
b1 = init_w((n_latent, ))
W3 = init_w((n_input, n_hidden))
W2 = init_w((n_hidden, n_latent))
W1 = init_w((n_hidden, n_latent))
# Gaussian Encoder
x = T.matrix("x")
h_encoder = T.tanh(T.dot(x, W3) + b3)
mu = T.dot(h_encoder, W1) + b1
log_sig2 = T.dot(h_encoder, W2) + b2
# This expression is simple (not an expectation) because we're using normal priors and posteriors
DKL = (1.0 + log_sig2 - mu**2 - T.exp(log_sig2)).sum(axis = 1)/2.0
# Bernoulli MLP weights and biases (decoder)
bernoulli_b1 = init_w((n_hidden, ))
bernoulli_b2 = init_w((n_input, ))
bernoulli_W1 = init_w((n_latent, n_hidden))
bernoulli_W2 = init_w((n_hidden, n_input))
# Bernoulli Decoder
std_normal = T.matrix("std_normal")
z = mu + T.sqrt(T.exp(log_sig2))*std_normal
h_decoder = T.tanh(T.dot(z, bernoulli_W1) + bernoulli_b1)
y = T.nnet.sigmoid(T.dot(h_decoder, bernoulli_W2) + bernoulli_b2)
log_likelihood = -T.nnet.binary_crossentropy(y, x).sum(axis = 1)
# Only the weight matrices W will be regularized (weight decay)
W = [W3, W1, W2, bernoulli_W1, bernoulli_W2]
b = [b3, b1, b2, bernoulli_b1, bernoulli_b2]
params = W + b
# Our lb and cost
lower_bound = (DKL + log_likelihood).mean()
cost = -lower_bound
def adagrad(cost, params, lr=0.001, epsilon=1e-6):
grads = T.grad(cost=cost, wrt=params)
updates = []
for p, g in zip(params, grads):
acc = theano.shared(p.get_value() * 0.)
acc_new = acc + g ** 2
gradient_scaling = T.sqrt(acc_new + epsilon)
g = g / gradient_scaling
updates.append((acc, acc_new))
updates.append((p, p - lr * g))
return updates
updates = adagrad(cost, params, lr=0.02)
train_model = theano.function(inputs=[x, std_normal],
outputs=cost,
updates=updates,
mode='FAST_RUN',
allow_input_downcast=True)
eval_model = theano.function(inputs=[x, std_normal], outputs=lower_bound,
mode='FAST_RUN',
allow_input_downcast=True)
training = []
#validation = []
for i in range(n_epochs):
minibatch_train = [ x_train[j] for j in np.random.randint(0,x_train.shape[0],batch_size) ]
#val_cost = eval_model(x_test, np.random.normal(size = (len(x_test), n_latent)))
train_cost = train_model(minibatch_train, np.random.normal(size = (batch_size, n_latent)))
training.append(train_cost)
#validation.append(val_cost)
plt.ylabel("-Lower bound")
plt.xlabel("Minibatch (" + str(batch_size) + " samples)")
plt.plot(-np.array(training), label = "Train")
plt.show()
# Extract estimated parameters (W, b)
est_W3 = updates[1][0].get_value()
est_W1 = updates[3][0].get_value()
est_W2 = updates[5][0].get_value()
est_b_W1 = updates[7][0].get_value()
est_b_W2 = updates[9][0].get_value()
est_b3 = updates[11][0].get_value()
est_b1 = updates[13][0].get_value()
est_b2 = updates[15][0].get_value()
est_b_b1 = updates[17][0].get_value()
est_b_b2 = updates[19][0].get_value()
est_b3
import numba
from numba import jit
@jit
def sigmoid(x):
return 1 / (1 + np.exp(-x))
est_h_encoder = np.tanh(np.dot(x_test, est_W3) + est_b3)
phi_mu = np.dot(est_h_encoder, est_W1) + est_b1
phi_log_sig2 = np.dot(est_h_encoder, est_W2) + est_b2
a, b =phi_log_sig2.shape
a, b
z_ = phi_mu + np.sqrt(np.exp(phi_log_sig2))*np.random.randn(a, b)
est_h_decoder = np.tanh(np.dot(z_, est_b_W1) + est_b_b1)
c = np.dot(est_h_decoder, est_b_W2)
y_ = sigmoid(c + est_b_b2)
y_.shape
# Regenerate data for visualization
p = y_.mean(axis =0)
y_.shape
x_new = np.random.binomial(1,y_,[10000,784])
x_new = x_new.reshape([10000, 28, 28])
for i in range(10):
plt.subplot(2,5,i+1)
plt.imshow(x_new[i], cmap='gray')
plt.xticks([])
plt.yticks([])
plt.tight_layout()
x_test = x_test.reshape([10000, 28, 28])
for i in range(10):
plt.subplot(2,5,i+1)
plt.imshow(x_test[i], cmap='gray')
plt.xticks([])
plt.yticks([])
plt.tight_layout()
t = T.vector()
h = T.tanh(T.dot(t, bernoulli_W1) + bernoulli_b1)
yt = T.nnet.sigmoid(T.dot(h, bernoulli_W2) + bernoulli_b2)
visualize = theano.function([t], yt,
mode='FAST_RUN',
allow_input_downcast=True)
'''gaussian decoder and freyface starts next:'''
from scipy.io import loadmat
ff = loadmat('frey_rawface.mat', squeeze_me=True, struct_as_record=False)
ff1 = ff["ff"].T
f_train=ff1[:1500,]
f_test=ff1[1501:,]
x=f_train
x.shape[1]
std_normal = T.matrix("std_normal")
z = mu + T.sqrt(T.exp(log_sig2))*std_normal
h_decoder = T.tanh(T.dot(z, W6) + b6)
mu_prime = T.dot(h_decoder, W4) + b4
log_sig2_prime = T.dot(h_decoder, W5) + b5
T.sum(log_sig2_prime)
log_sig2_prime.sum(axis=1)
# our logpxz
-0.5*(x.shape[1]*np.log(2*np.pi))-0.5*(log_sig2_prime.sum(axis=1))-0.5*(x-mu_prime)**2/(np.exp(log_sig2_prime)).sum(axis=1)
#from VAE github:y0st
# had to change axis=2 to 1, SOOO unreliable this guy!! I would use ours!
logpxz = (-(0.5 * np.log(2 * np.pi) + 0.5 * log_sig2_prime) - 0.5 * ((x - mu_prime)**2 / T.exp(log_sig2_prime))).sum(axis=1).mean(axis=0)
logpxz
# Parameters
# Gaussian MLP weights and biases (decoder)
#initialize \theta
b6 = init_w((n_hidden, ))
b5 = init_w((n_input, ))
b4 = init_w((n_input, ))
W6 = init_w((n_latent, n_hidden))
W5 = init_w((n_hidden, n_input))
W4 = init_w((n_hidden, n_input))
# Gaussian Decoder --- ours!!!
std_normal = T.matrix("std_normal")
z = mu + T.sqrt(T.exp(log_sig2))*std_normal
h_decoder = T.tanh(T.dot(z, W6) + b6)
mu_prime = T.dot(h_decoder, W4) + b4
log_sig2_prime = T.dot(h_decoder, W5) + b5
log_likelihood_gaus=-0.5*(x.shape[1]*np.log(2*np.pi))-0.5*(log_sig2_prime.sum(axis=1))-0.5*(x-mu_prime)**2/(np.exp(log_sig2_prime)).sum(axis=1)
```
| github_jupyter |
# Navigation
---
You are welcome to use this coding environment to train your agent for the project. Follow the instructions below to get started!
### 1. Start the Environment
Run the next code cell to install a few packages. This line will take a few minutes to run!
```
!pip install numpy --upgrade
!pip -q install ./python
```
The environment is already saved in the Workspace and can be accessed at the file path provided below. Please run the next code cell without making any changes.
```
from unityagents import UnityEnvironment
from dqn_agent import Agent
import torch
import numpy as np
import random
from collections import deque
import matplotlib.pyplot as plt
%matplotlib inline
# please do not modify the line below
env = UnityEnvironment(file_name="/data/Banana_Linux_NoVis/Banana.x86_64")
```
Environments contain **_brains_** which are responsible for deciding the actions of their associated agents. Here we check for the first brain available, and set it as the default brain we will be controlling from Python.
```
# get the default brain
brain_name = env.brain_names[0]
brain = env.brains[brain_name]
```
### 2. Examine the State and Action Spaces
Run the code cell below to print some information about the environment.
```
# reset the environment
env_info = env.reset(train_mode=True)[brain_name]
# number of agents in the environment
print('Number of agents:', len(env_info.agents))
# number of actions
action_size = brain.vector_action_space_size
print('Number of actions:', action_size)
# examine the state space
state = env_info.vector_observations[0]
print('States look like:', state)
state_size = len(state)
print('States have length:', state_size)
```
### 3. Take Random Actions in the Environment
In the next code cell, you will learn how to use the Python API to control the agent and receive feedback from the environment.
Note that **in this coding environment, you will not be able to watch the agent while it is training**, and you should set `train_mode=True` to restart the environment.
```
#Here we are taking random steps
env_info = env.reset(train_mode=True)[brain_name] # reset the environment
state = env_info.vector_observations[0] # get the current state
score = 0 # initialize the score
while True:
action = np.random.randint(action_size) # select an action
env_info = env.step(action)[brain_name] # send the action to the environment
next_state = env_info.vector_observations[0] # get the next state
reward = env_info.rewards[0] # get the reward
done = env_info.local_done[0] # see if episode has finished
score += reward # update the score
state = next_state # roll over the state to next time step
if done: # exit loop if episode finished
break
print("Score: {}".format(score))
```
When finished, you can close the environment.
### 4. Train your agent
A few **important notes**:
- When training the environment, set `train_mode=True`, so that the line for resetting the environment looks like the following:
```python
env_info = env.reset(train_mode=True)[brain_name]
```
- In this coding environment, you will not be able to watch the agent while it is training. However, **_after training the agent_**, you can download the saved model weights to watch the agent on your own machine!
##### 4.1 Learning Algorithm - DQN
Deep Q-Networks(DQN) was proposed by Mnih et al. (2015). It takes agent's state as input and outputs Q action values. It uses experience replay and target network to stabilize the model training.
<figure>
<img src="images/pseudocode-dqn.png" width="400" height="400">
<br>
<figcaption style = "text-align:center; font-style:italic">Taken from Human-level control through deep reinforcement learning(Mnih et al. (2015))</figcaption>
</figure>
##### 4.2 Model Architecture
The model is made of three fully connected layers. The number of neurons in first two layers is 64 and in the last layer it's equal to action size. Each layer's output except the last layer is transformed using the RelU activation function.
##### 4.3 Hyperparameters
* BUFFER_SIZE = int(1e5) # replay buffer size
* BATCH_SIZE = 64 # minibatch size
* GAMMA = 0.99 # discount factor
* TAU = 1e-3 # for soft update of target parameters
* LR = 5e-4 # learning rate
* n_episodes = 2000 # maximum number of training episodes
* max_t = 1000 # maximum number of time steps per episode
* eps_start = 1.0 # starting value of epsilon, for epsilon-greedy action selection
* eps_end = 0.01 # minimum value of epsilon
* eps_decay = 0.995 # multiplicative factor (per episode) for decreasing epsilon
```
agent = Agent(state_size=37, action_size=4, seed=0)
def dqn(n_episodes = 2000, max_t = 1000, eps_start = 1.0, eps_end = 0.01, eps_decay = 0.995):
"""Deep Q-Learning.
Params
======
n_episodes (int): maximum number of training episodes
max_t (int): maximum number of time steps per episode
eps_start (float): starting value of epsilon, for epsilon-greedy action selection
eps_end (float) : minimum value of epsilon
eps_decay (float) : multiplicative factor (per episode) for decreasing epsilon
"""
scores = [] #list containing scores from each episode
scores_window = deque(maxlen = 100) # last 100 scores
eps = eps_start #Initialize epsilon
for i_episode in range(1, n_episodes+1):
env_info = env.reset(train_mode=True)[brain_name]
state = env_info.vector_observations[0]
score = 0
for t in range(max_t):
action = agent.act(state, eps)
env_info = env.step(action)[brain_name]
next_state = env_info.vector_observations[0]
reward = env_info.rewards[0]
done = env_info.local_done[0]
agent.step(state, action, reward, next_state, done)
state = next_state
score += reward
if done:
break
scores_window.append(score) # save most recent scores
scores.append(score) # save most recent score
eps = max(eps_end, eps_decay*eps) # decrease epsilon
print("\rEpisode {}\tAverage Score: {:.2f}".format(i_episode, np.mean(scores_window)), end ="")
if i_episode % 100 == 0:
print("\rEpisode {}\tAverage Score: {:.2f}".format(i_episode, np.mean(scores_window)))
if np.mean(scores_window) >= 13.0:
print("\nEnvironment solved in {:d} episodes!\t Average score: {:.2f}".format(i_episode, np.mean(scores_window)))
torch.save(agent.qnetwork_local.state_dict(), 'checkpoint.pth')
break
return scores
scores = dqn()
fig = plt.figure()
ax = fig.add_subplot(111)
plt.plot(np.arange(len(scores)), scores)
plt.ylabel("Score")
plt.xlabel("No of Episodes")
plt.show()
#Load a trained agent
agent.qnetwork_local.load_state_dict(torch.load("checkpoint.pth"))
for i in range(3):
env_info = env.reset(train_mode=False)[brain_name]
state = env_info.vector_observations[0]
score = 0
for j in range(1000):
action = agent.act(state)
env_info = env.step(action)[brain_name]
next_state = env_info.vector_observations[0]
reward = env_info.rewards[0]
done = env_info.local_done[0]
agent.step(state, action, reward, next_state, done)
state = next_state
score += reward
if done:
break
print("\rEpisode {}\t Score: {:.2f}".format(i+1, score))
env.close()
```
### 5. Future ideas to improve the agent's performance
More experiments can be done to increase the performance of agent by applying different extensions of DQN:
* Double DQN (DDQN)
* Prioritized experience replay
* Dueling DQN
* A3C
* Distributional DQN
* Noisy DQN
We can also apply all the abobe extensions together. This was done by Deepmind's researchers and they have termed it Rainbow. This algorithm has outperformed each of the extension achieved SOTA results on Atari 2600.
<figure>
<img src="images/rainbow.png" width="400" height="400">
<br>
<figcaption style = "text-align:center; font-style:italic">Taken from Rainbow: Combining Improvements in Deep Reinforcement Learning(Hessel et al. (2017))</figcaption>
</figure>
| github_jupyter |
```
!rm -rf output-*/
```
## Test 1: discretize = False
```
!mkdir -p output-1
!docker run -it \
--mount type='bind',src="$(pwd)",target='/datadir' \
fiddle-v020 \
python -m FIDDLE.run \
--data_fname='/datadir/input/data.csv' \
--population_fname='/datadir/input/pop.csv' \
--config_fname='/datadir/input/config-1.yaml' \
--output_dir='/datadir/output-1/' \
--T=4 --dt=1.0 \
--theta_1=0.001 --theta_2=0.001 --theta_freq=1 \
--stats_functions 'min' 'max' 'mean' \
--no_prefilter --no_postfilter
import numpy as np
import pandas as pd
import json
import sparse
S = sparse.load_npz('output-1/S_all.npz')
S_names = json.load(open('output-1/S_all.feature_names.json', 'r'))
S_index = pd.read_csv('output-1/S.ID.csv').set_index(['ID'])
df_S = pd.DataFrame(S.todense(), columns=S_names, index=S_index.index)
X = sparse.load_npz('output-1/X_all.npz')
X_names = json.load(open('output-1/X_all.feature_names.json', 'r'))
X_index = pd.read_csv('output-1/X.ID,t_range.csv').set_index(['ID', 't_range'])
df_X = pd.DataFrame(X.todense().reshape(-1, X.shape[-1]), columns=X_names, index=X_index.index)
display(df_S)
display(df_X)
```
## Test 2: discretize = True, use_ordinal_encoding = False
```
!mkdir -p output-2
!docker run -it \
--mount type='bind',src="$(pwd)",target='/datadir' \
fiddle-v020 \
python -m FIDDLE.run \
--data_fname='/datadir/input/data.csv' \
--population_fname='/datadir/input/pop.csv' \
--config_fname='/datadir/input/config-2.yaml' \
--output_dir='/datadir/output-2/' \
--T=4 --dt=1.0 \
--theta_1=0.001 --theta_2=0.001 --theta_freq=1 \
--stats_functions 'min' 'max' 'mean' \
--no_prefilter --no_postfilter
import numpy as np
import pandas as pd
import json
import sparse
S = sparse.load_npz('output-2/S_all.npz')
S_names = json.load(open('output-2/S_all.feature_names.json', 'r'))
S_index = pd.read_csv('output-2/S.ID.csv').set_index(['ID'])
df_S = pd.DataFrame(S.todense(), columns=S_names, index=S_index.index)
X = sparse.load_npz('output-2/X_all.npz')
X_names = json.load(open('output-2/X_all.feature_names.json', 'r'))
X_index = pd.read_csv('output-2/X.ID,t_range.csv').set_index(['ID', 't_range'])
df_X = pd.DataFrame(X.todense().reshape(-1, X.shape[-1]), columns=X_names, index=X_index.index)
display(df_S)
display(df_X)
```
## Test 3: discretize = True, use_ordinal_encoding = True
```
!mkdir -p output-3
!docker run -it \
--mount type='bind',src="$(pwd)",target='/datadir' \
fiddle-v020 \
python -m FIDDLE.run \
--data_fname='/datadir/input/data.csv' \
--population_fname='/datadir/input/pop.csv' \
--config_fname='/datadir/input/config-3.yaml' \
--output_dir='/datadir/output-3/' \
--T=4 --dt=1.0 \
--theta_1=0.001 --theta_2=0.001 --theta_freq=1 \
--stats_functions 'min' 'max' 'mean' \
--no_prefilter --no_postfilter
import numpy as np
import pandas as pd
import json
import sparse
S = sparse.load_npz('output-3/S_all.npz')
S_names = json.load(open('output-3/S_all.feature_names.json', 'r'))
S_index = pd.read_csv('output-3/S.ID.csv').set_index(['ID'])
df_S = pd.DataFrame(S.todense(), columns=S_names, index=S_index.index)
X = sparse.load_npz('output-3/X_all.npz')
X_names = json.load(open('output-3/X_all.feature_names.json', 'r'))
X_index = pd.read_csv('output-3/X.ID,t_range.csv').set_index(['ID', 't_range'])
df_X = pd.DataFrame(X.todense().reshape(-1, X.shape[-1]), columns=X_names, index=X_index.index)
display(df_S)
display(df_X)
```
| github_jupyter |
## CORDEX ESGF submission form
#### General Information
Data to be submitted for ESGF data publication must follow the rules outlined in the Cordex Archive Design Document <br /> (https://verc.enes.org/data/projects/documents/cordex-archive-design)
Thus file names have to follow the pattern:<br />
> *VariableName_Domain_GCMModelName_CMIP5ExperimentName_CMIP5EnsembleMember_RCMModelName_RCMVersionID_Frequency[_StartTime-EndTime].nc* <br />
**Example:** *tas_AFR-44_MPI-M-MPI-ESM-LR_rcp26_r1i1p1_MPI-CSC-REMO2009_v1_mon_yyyymm-yyyymm.nc*
The directory structure in which these files are stored follow the pattern:<br />
> *activity/product/Domain/Institution/
> GCMModelName/CMIP5ExperimentName/CMIP5EnsembleMember/
> RCMModelName/RCMVersionID/Frequency/VariableName* <br />
**Example:** *CORDEX/output/AFR-44/MPI-CSC/MPI-M-MPI-ESM-LR/rcp26/r1i1p1/MPI-CSC-REMO2009/v1/mon/tas/tas_AFR-44_MPI-M-MPI-ESM-LR_rcp26_r1i1p1_MPI-CSC-REMO2009_v1_mon_yyyymm-yyyymm.nc*
**Notice:** If your model is not yet registered, please contact contact **cordex-registration@cordex.org**
specifying: Full institution name, Short institution name (acronym), Contact person and
e-mail, RCM Name (acronym), Terms of Use (unrestricted or non-commercial only) and the CORDEX domains in which you are interested.
At some CORDEX ESGF data centers a 'data submission form' is in use in order to improve initial information exchange between data providers and the data center. The form has to be filled before the publication process can be started. In case you have questions pleas contact the individual data centers:
o at DKRZ: cordex@dkrz.de
o at SMHI: rossby.cordex@smhi.se
---
```
from dkrz_forms import form_widgets
form_widgets.show_status('form-submission')
```
# Start submission procedure
The submission is based on this interactive document consisting of "cells" you can modify and then evaluate
evaluation of cells is done by selecting the cell and then press the keys "Shift" + "Enter"
<br /> please evaluate the following cell to initialize your form
```
MY_LAST_NAME = "...." # e.gl MY_LAST_NAME = "schulz"
#-------------------------------------------------
from dkrz_forms import form_handler, form_widgets, checks
form_info = form_widgets.check_pwd(MY_LAST_NAME)
sfg = form_handler.init_form(form_info)
sf = sf.sub.entity_out.form_info
```
please provide information on the contact person for this CORDEX data submission request
#### Type of submission
please specify the type of this data submission:
- "initial_version" for first submission of data
- "new _version" for a re-submission of previousliy submitted data
- "retract" for the request to retract previously submitted data
```
sf.submission_type = "..." # example: sf.submission_type = "initial_version"
```
## Requested general information
## Please provide model and institution info as well as an example of a file name
#### institution
The value of this field has to equal the value of the optional NetCDF attribute 'institution'
(long version) in the data files if the latter is used.
```
sf.institution = "..." # example: sf.institution = "Alfred Wegener Institute"
```
##### institute_id
The value of this field has to equal the value of the global NetCDF attribute 'institute_id'
in the data files and must equal the 4th directory level. It is needed before the publication
process is started in order that the value can be added to the relevant CORDEX list of CV1
if not yet there. Note that 'institute_id' has to be the first part of 'model_id'
```
sf.institute_id = "..." # example: sf.institute_id = "AWI"
```
##### model_id
The value of this field has to be the value of the global NetCDF attribute 'model_id'
in the data files. It is needed before the publication process is started in order that
the value can be added to the relevant CORDEX list of CV1 if not yet there.
Note that it must be composed by the 'institute_id' follwed by the RCM CORDEX model name,
separated by a dash. It is part of the file name and the directory structure.
```
sf.model_id = "..." # example: sf.model_id = "AWI-HIRHAM5"
```
#### experiment_id and time_period
Experiment has to equal the value of the global NetCDF attribute 'experiment_id'
in the data files. Time_period gives the period of data for which the publication
request is submitted. If you intend to submit data from multiple experiments you may
add one line for each additional experiment or send in additional publication request sheets.
```
sf.experiment_id = "..." # example: sf.experiment_id = "evaluation"
# ["value_a","value_b"] in case of multiple experiments
sf.time_period = "..." # example: sf.time_period = "197901-201412"
# ["time_period_a","time_period_b"] in case of multiple values
```
#### Example file name
Please provide an example file name of a file in your data collection,
this name will be used to derive the other
```
sf.example_file_name = "..." # example: sf.example_file_name = "tas_AFR-44_MPI-M-MPI-ESM-LR_rcp26_r1i1p1_MPI-CSC-REMO2009_v1_mon_yyyymm-yyyymm.nc"
# Please run this cell as it is to check your example file name structure
# to_do: implement submission_form_check_file function - output result (attributes + check_result)
form_handler.cordex_file_info(sf,sf.example_file_name)
```
#### information on the grid_mapping
the NetCDF/CF name of the data grid ('rotated_latitude_longitude', 'lambert_conformal_conic', etc.),
i.e. either that of the native model grid, or 'latitude_longitude' for the regular -XXi grids
```
sf.grid_mapping_name = "..." # example: sf.grid_mapping_name = "rotated_latitude_longitude"
```
Does the grid configuration exactly follow the specifications in ADD2 (Table 1)
in case the native grid is 'rotated_pole'? If not, comment on the differences; otherwise write 'yes' or 'N/A'. If the data is not delivered on the computational grid it has to be noted here as well.
```
sf.grid_as_specified_if_rotated_pole = "..." # example: sf.grid_as_specified_if_rotated_pole = "yes"
```
### Please provide information on quality check performed on the data you plan to submit
Please answer 'no', 'QC1', 'QC2-all', 'QC2-CORDEX', or 'other'.
'QC1' refers to the compliancy checker that can be downloaded at http://cordex.dmi.dk.
'QC2' refers to the quality checker developed at DKRZ.
If your answer is 'other' give some informations.
```
sf.data_qc_status = "..." # example: sf.data_qc_status = "QC2-CORDEX"
sf.data_qc_comment = "..." # any comment of quality status of the files
```
### Terms of use
Please give the terms of use that shall be asigned to the data.
The options are 'unrestricted' and 'non-commercial only'.
For the full text 'Terms of Use' of CORDEX data refer to
http://cordex.dmi.dk/joomla/images/CORDEX/cordex_terms_of_use.pdf
```
sf.terms_of_use = "..." # example: sf.terms_of_use = "unrestricted"
```
## Information on directory structure and data access path
(and other information needed for data transport and data publication)
If there is any directory structure deviation from the CORDEX standard please specify here.
Otherwise enter 'compliant'. Please note that deviations MAY imply that data can not be accepted.
```
sf.directory_structure = "..." # example: sf.directory_structure = "compliant"
```
Give the path where the data reside, for example:
blizzard.dkrz.de:/scratch/b/b364034/. If not applicable write N/A and give data access information in the data_information string
```
sf.data_path = "..." # example: sf.data_path = "mistral.dkrz.de:/mnt/lustre01/work/bm0021/k204016/CORDEX/archive/"
sf.data_information = "..." # ...any info where data can be accessed and transfered to the data center ... "
```
#### Exclude variable list
In each CORDEX file there may be only one variable which shall be published and searchable at the ESGF portal (target variable). In order to facilitate publication, all non-target variables are included in a list used by the publisher to avoid publication. A list of known non-target variables is [time, time_bnds, lon, lat, rlon ,rlat ,x ,y ,z ,height, plev, Lambert_Conformal, rotated_pole]. Please enter other variables into the left field if applicable (e.g. grid description variables), otherwise write 'N/A'.
```
sf.exclude_variables_list = "..." # example: sf.exclude_variables_list=["bnds", "vertices"]
```
#### Uniqueness of tracking_id and creation_date
In case any of your files is replacing a file already published, it must not have the same tracking_id nor
the same creation_date as the file it replaces.
Did you make sure that that this is not the case ?
Reply 'yes'; otherwise adapt the new file versions.
```
sf.uniqueness_of_tracking_id = "..." # example: sf.uniqueness_of_tracking_id = "yes"
```
## Variable list
list of variables submitted -- please remove the ones you do not provide:
```
sf.variable_list_day = [
"clh","clivi","cll","clm","clt","clwvi",
"evspsbl","evspsblpot",
"hfls","hfss","hurs","huss","hus850",
"mrfso","mrro","mrros","mrso",
"pr","prc","prhmax","prsn","prw","ps","psl",
"rlds","rlus","rlut","rsds","rsdt","rsus","rsut",
"sfcWind","sfcWindmax","sic","snc","snd","snm","snw","sund",
"tas","tasmax","tasmin","tauu","tauv","ta200","ta500","ta850","ts",
"uas","ua200","ua500","ua850",
"vas","va200","va500","va850","wsgsmax",
"zg200","zg500","zmla"
]
sf.variable_list_mon = [
"clt",
"evspsbl",
"hfls","hfss","hurs","huss","hus850",
"mrfso","mrro","mrros","mrso",
"pr","psl",
"rlds","rlus","rlut","rsds","rsdt","rsus","rsut",
"sfcWind","sfcWindmax","sic","snc","snd","snm","snw","sund",
"tas","tasmax","tasmin","ta200",
"ta500","ta850",
"uas","ua200","ua500","ua850",
"vas","va200","va500","va850",
"zg200","zg500"
]
sf.variable_list_sem = [
"clt",
"evspsbl",
"hfls","hfss","hurs","huss","hus850",
"mrfso","mrro","mrros","mrso",
"pr","psl",
"rlds","rlus","rlut","rsds","rsdt","rsus","rsut",
"sfcWind","sfcWindmax","sic","snc","snd","snm","snw","sund",
"tas","tasmax","tasmin","ta200","ta500","ta850",
"uas","ua200","ua500","ua850",
"vas","va200","va500","va850",
"zg200","zg500"
]
sf.variable_list_fx = [
"areacella",
"mrsofc",
"orog",
"rootd",
"sftgif","sftlf"
]
```
## Check your submission form
Please evaluate the following cell to check your submission form.
In case of errors, please go up to the corresponden information cells and update your information accordingly.
```
# simple consistency check report for your submission form
res = form_handler.check_submission(sf)
sf.sub.valid_submission = res['valid_submission']
form_handler.DictTable(res)
```
# Save your form
your form will be stored (the form name consists of your last name plut your keyword)
```
form_handler.save_form(sf,"..my comment..") # edit my comment info
#evaluate this cell if you want a reference to the saved form emailed to you
# (only available if you access this form via the DKRZ form hosting service)
form_handler.email_form_info()
# evaluate this cell if you want a reference (provided by email)
# (only available if you access this form via the DKRZ hosting service)
form_handler.email_form_info(sf)
```
# officially submit your form
the form will be submitted to the DKRZ team to process
you also receive a confirmation email with a reference to your online form for future modifications
```
form_handler.email_form_info(sf)
form_handler.form_submission(sf)
```
| github_jupyter |
```
import numpy as np
import torch
import gym
import pybullet_envs
import os
import utils
import TD3
import OurDDPG
import DDPG
# Runs policy for X episodes and returns average reward
# A fixed seed is used for the eval environment
def eval_policy(policy, env_name, seed, eval_episodes=10):
eval_env = gym.make(env_name)
eval_env.seed(seed + 100)
avg_reward = 0.
for _ in range(eval_episodes):
state, done = eval_env.reset(), False
while not done:
action = policy.select_action(np.array(state))
state, reward, done, _ = eval_env.step(action)
avg_reward += reward
avg_reward /= eval_episodes
print("---------------------------------------")
print(f"Evaluation over {eval_episodes} episodes: {avg_reward:.3f}")
print("---------------------------------------")
return avg_reward
def main():
args = {
"policy" : "TD3", # Policy name (TD3, DDPG or OurDDPG)
"env" : "AntBulletEnv-v0", # OpenAI gym environment name
"seed" : 0, # Sets Gym, PyTorch and Numpy seeds
"start_timesteps" : 25e3, # Time steps initial random policy is used
"eval_freq" : 5e3, # How often (time steps) we evaluate
"max_timesteps" : 2e6, # Max time steps to run environment
"expl_noise" : 0.1, # Std of Gaussian exploration noise
"batch_size" : 256, # Batch size for both actor and critic
"discount" : 0.99, # Discount factor
"tau" : 0.005, # Target network update rate
"policy_noise" : 0.2, # Noise added to target policy during critic update
"noise_clip" : 0.5, # Range to clip target policy noise
"policy_freq" : 2, # Frequency of delayed policy updates
"save_model" : "store_true", # Save model and optimizer parameters
"load_model" : "", # Model load file name, "" doesn't load, "default" uses file_name
}
file_name = f"{args['policy']}_{args['env']}_{args['seed']}"
print("---------------------------------------")
print(f"Policy: {args['policy']}, Env: {args['env']}, Seed: {args['seed']}")
print("---------------------------------------")
if not os.path.exists("./results"):
os.makedirs("./results")
if args['save_model'] and not os.path.exists("./models"):
os.makedirs("./models")
env = gym.make(args['env'])
# Set seeds
env.seed(args['seed'])
env.action_space.seed(args['seed'])
torch.manual_seed(args['seed'])
np.random.seed(args['seed'])
state_dim = env.observation_space.shape[0]
action_dim = env.action_space.shape[0]
max_action = float(env.action_space.high[0])
kwargs = {
"state_dim": state_dim,
"action_dim": action_dim,
"max_action": max_action,
"discount": args['discount'],
"tau": args['tau'],
}
# Initialize policy
if args['policy'] == "TD3":
# Target policy smoothing is scaled wrt the action scale
kwargs["policy_noise"] = args['policy_noise'] * max_action
kwargs["noise_clip"] = args['noise_clip'] * max_action
kwargs["policy_freq"] = args['policy_freq']
policy = TD3.TD3(**kwargs)
elif args['policy'] == "OurDDPG":
policy = OurDDPG.DDPG(**kwargs)
elif args['policy'] == "DDPG":
policy = DDPG.DDPG(**kwargs)
if args['load_model'] != "":
policy_file = file_name if args['load_model'] == "default" else args['load_model']
policy.load(f"./models/{policy_file}")
replay_buffer = utils.ReplayBuffer(state_dim, action_dim)
# Evaluate untrained policy
evaluations = [eval_policy(policy, args['env'], args['seed'])]
state, done = env.reset(), False
episode_reward = 0
episode_timesteps = 0
episode_num = 0
for t in range(int(args['max_timesteps'])):
episode_timesteps += 1
# Select action randomly or according to policy
if t < args['start_timesteps']:
action = env.action_space.sample()
else:
action = (
policy.select_action(np.array(state))
+ np.random.normal(0, max_action * args['expl_noise'], size=action_dim)
).clip(-max_action, max_action)
# Perform action
next_state, reward, done, _ = env.step(action)
done_bool = float(done) if episode_timesteps < env._max_episode_steps else 0
# Store data in replay buffer
replay_buffer.add(state, action, next_state, reward, done_bool)
state = next_state
episode_reward += reward
# Train agent after collecting sufficient data
if t >= args['start_timesteps']:
policy.train(replay_buffer, args['batch_size'])
if done:
# +1 to account for 0 indexing. +0 on ep_timesteps since it will increment +1 even if done=True
print(f"Total T: {t+1} Episode Num: {episode_num+1} Episode T: {episode_timesteps} Reward: {episode_reward:.3f}")
# Reset environment
state, done = env.reset(), False
episode_reward = 0
episode_timesteps = 0
episode_num += 1
# Evaluate episode
if (t + 1) % args['eval_freq'] == 0:
evaluations.append(eval_policy(policy, args['env'], args['seed']))
np.save(f"./results/{file_name}", evaluations)
if args['save_model']: policy.save(f"./models/{file_name}")
main()
```
| github_jupyter |
## Dependencies
```
import json, warnings, shutil
from tweet_utility_scripts import *
from tweet_utility_preprocess_roberta_scripts import *
from transformers import TFRobertaModel, RobertaConfig
from tokenizers import ByteLevelBPETokenizer
from tensorflow.keras.models import Model
from tensorflow.keras import optimizers, metrics, losses, layers
from tensorflow.keras.callbacks import EarlyStopping, TensorBoard, ModelCheckpoint
SEED = 0
seed_everything(SEED)
warnings.filterwarnings("ignore")
```
# Load data
```
database_base_path = '/kaggle/input/tweet-dataset-split-roberta-base-96/'
k_fold = pd.read_csv(database_base_path + '5-fold.csv')
display(k_fold.head())
# Unzip files
!tar -xvf /kaggle/input/tweet-dataset-split-roberta-base-96/fold_1.tar.gz
!tar -xvf /kaggle/input/tweet-dataset-split-roberta-base-96/fold_2.tar.gz
!tar -xvf /kaggle/input/tweet-dataset-split-roberta-base-96/fold_3.tar.gz
# !tar -xvf /kaggle/input/tweet-dataset-split-roberta-base-96/fold_4.tar.gz
# !tar -xvf /kaggle/input/tweet-dataset-split-roberta-base-96/fold_5.tar.gz
```
# Model parameters
```
vocab_path = database_base_path + 'vocab.json'
merges_path = database_base_path + 'merges.txt'
base_path = '/kaggle/input/qa-transformers/roberta/'
config = {
"MAX_LEN": 96,
"BATCH_SIZE": 32,
"EPOCHS": 5,
"LEARNING_RATE": 3e-5,
"ES_PATIENCE": 1,
"question_size": 4,
"N_FOLDS": 1,
"base_model_path": base_path + 'roberta-base-tf_model.h5',
"config_path": base_path + 'roberta-base-config.json'
}
with open('config.json', 'w') as json_file:
json.dump(json.loads(json.dumps(config)), json_file)
```
# Model
```
module_config = RobertaConfig.from_pretrained(config['config_path'], output_hidden_states=False)
def model_fn(MAX_LEN):
input_ids = layers.Input(shape=(MAX_LEN,), dtype=tf.int32, name='input_ids')
attention_mask = layers.Input(shape=(MAX_LEN,), dtype=tf.int32, name='attention_mask')
base_model = TFRobertaModel.from_pretrained(config['base_model_path'], config=module_config, name="base_model")
sequence_output = base_model({'input_ids': input_ids, 'attention_mask': attention_mask})
last_state = sequence_output[0]
x_start = layers.Dropout(0.1)(last_state)
x_start = layers.Conv1D(1, 1)(x_start)
x_start = layers.Flatten()(x_start)
y_start = layers.Activation('softmax', name='y_start')(x_start)
x_end = layers.Dropout(0.1)(last_state)
x_end = layers.Conv1D(1, 1)(x_end)
x_end = layers.Flatten()(x_end)
y_end = layers.Activation('softmax', name='y_end')(x_end)
model = Model(inputs=[input_ids, attention_mask], outputs=[y_start, y_end])
model.compile(optimizers.Adam(lr=config['LEARNING_RATE']),
loss=losses.CategoricalCrossentropy(),
metrics=[metrics.CategoricalAccuracy()])
return model
```
# Tokenizer
```
tokenizer = ByteLevelBPETokenizer(vocab_file=vocab_path, merges_file=merges_path, lowercase=True, add_prefix_space=True)
tokenizer.save('./')
```
# Train
```
history_list = []
AUTO = tf.data.experimental.AUTOTUNE
for n_fold in range(config['N_FOLDS']):
n_fold +=1
print('\nFOLD: %d' % (n_fold))
# Load data
base_data_path = 'fold_%d/' % (n_fold)
x_train = np.load(base_data_path + 'x_train.npy')
y_train = np.load(base_data_path + 'y_train.npy')
x_valid = np.load(base_data_path + 'x_valid.npy')
y_valid = np.load(base_data_path + 'y_valid.npy')
### Delete data dir
shutil.rmtree(base_data_path)
# Train model
model_path = 'model_fold_%d.h5' % (n_fold)
model = model_fn(config['MAX_LEN'])
es = EarlyStopping(monitor='val_loss', mode='min', patience=config['ES_PATIENCE'],
restore_best_weights=True, verbose=1)
checkpoint = ModelCheckpoint(model_path, monitor='val_loss', mode='min',
save_best_only=True, save_weights_only=True)
history = model.fit(list(x_train), list(y_train),
validation_data=(list(x_valid), list(y_valid)),
batch_size=config['BATCH_SIZE'],
callbacks=[checkpoint, es],
epochs=config['EPOCHS'],
verbose=2).history
history_list.append(history)
# Make predictions
train_preds = model.predict(list(x_train))
valid_preds = model.predict(list(x_valid))
k_fold.loc[k_fold['fold_%d' % (n_fold)] == 'train', 'start_fold_%d' % (n_fold)] = train_preds[0].argmax(axis=-1)
k_fold.loc[k_fold['fold_%d' % (n_fold)] == 'train', 'end_fold_%d' % (n_fold)] = train_preds[1].argmax(axis=-1)
k_fold.loc[k_fold['fold_%d' % (n_fold)] == 'validation', 'start_fold_%d' % (n_fold)] = valid_preds[0].argmax(axis=-1)
k_fold.loc[k_fold['fold_%d' % (n_fold)] == 'validation', 'end_fold_%d' % (n_fold)] = valid_preds[1].argmax(axis=-1)
k_fold['end_fold_%d' % (n_fold)] = k_fold['end_fold_%d' % (n_fold)].astype(int)
k_fold['start_fold_%d' % (n_fold)] = k_fold['start_fold_%d' % (n_fold)].astype(int)
k_fold['end_fold_%d' % (n_fold)].clip(0, k_fold['text_len'], inplace=True)
k_fold['start_fold_%d' % (n_fold)].clip(0, k_fold['end_fold_%d' % (n_fold)], inplace=True)
k_fold['prediction_fold_%d' % (n_fold)] = k_fold.apply(lambda x: decode(x['start_fold_%d' % (n_fold)], x['end_fold_%d' % (n_fold)], x['text'], config['question_size'], tokenizer), axis=1)
k_fold['prediction_fold_%d' % (n_fold)].fillna('', inplace=True)
k_fold['jaccard_fold_%d' % (n_fold)] = k_fold.apply(lambda x: jaccard(x['text'], x['prediction_fold_%d' % (n_fold)]), axis=1)
```
# Model loss graph
```
sns.set(style="whitegrid")
for n_fold in range(config['N_FOLDS']):
print('Fold: %d' % (n_fold+1))
plot_metrics(history_list[n_fold])
```
# Model evaluation
```
display(evaluate_model_kfold(k_fold, config['N_FOLDS']).style.applymap(color_map))
```
# Visualize predictions
```
display(k_fold[[c for c in k_fold.columns if not (c.startswith('textID') or
c.startswith('text_len') or
c.startswith('selected_text_len') or
c.startswith('text_wordCnt') or
c.startswith('selected_text_wordCnt') or
c.startswith('fold_') or
c.startswith('start_fold_') or
c.startswith('end_fold_'))]].head(15))
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.